code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Boston Housing Data
#
# In order to gain a better understanding of the metrics used in regression settings, we will be looking at the Boston Housing dataset.
#
# First use the cell below to read in the dataset and set up the training and testing data that will be used for the rest of this problem.
# +
from sklearn.datasets import load_boston
from sklearn.model_selection import train_test_split
import numpy as np
import tests2 as t
boston = load_boston()
y = boston.target
X = boston.data
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.33, random_state=42)
# -
# > **Step 1:** Before we get too far, let's do a quick check of the models that you can use in this situation given that you are working on a regression problem. Use the dictionary and corresponding letters below to provide all the possible models you might choose to use.
# +
# When can you use the model - use each option as many times as necessary
a = 'regression'
b = 'classification'
c = 'both regression and classification'
models = {
'decision trees': c,
'random forest': c,
'adaptive boosting': c,
'logistic regression': b,
'linear regression': a,
}
#checks your answer, no need to change this code
t.q1_check(models)
# -
# > **Step 2:** Now for each of the models you found in the previous question that can be used for regression problems, import them using sklearn.
# +
# Import models from sklearn - notice you will want to use
# the regressor version (not classifier) - googling to find
# each of these is what we all do!
from sklearn.ensemble import RandomForestRegressor, AdaBoostRegressor
from sklearn.linear_model import LinearRegression
from sklearn.tree import DecisionTreeRegressor
# -
# > **Step 3:** Now that you have imported the 4 models that can be used for regression problems, instantate each below.
# +
# Instantiate each of the models you imported
# For now use the defaults for all the hyperparameters
tree_mod = DecisionTreeRegressor()
rf_mod = RandomForestRegressor()
ada_mod = AdaBoostRegressor()
reg_mod = LinearRegression()
# -
# > **Step 4:** Fit each of your instantiated models on the training data.
# Fit each of your models using the training data
tree_mod.fit(X_train, y_train)
rf_mod.fit(X_train, y_train)
ada_mod.fit(X_train, y_train)
reg_mod.fit(X_train, y_train)
# > **Step 5:** Use each of your models to predict on the test data.
# +
# Predict on the test values for each model
preds_tree = tree_mod.predict(X_test)
preds_rf = rf_mod.predict(X_test)
preds_ada = ada_mod.predict(X_test)
preds_reg = reg_mod.predict(X_test)
# -
# > **Step 6:** Now for the information related to this lesson. Use the dictionary to match the metrics that are used for regression and those that are for classification.
# +
# potential model options
a = 'regression'
b = 'classification'
c = 'both regression and classification'
#
metrics = {
'precision': b,
'recall': b,
'accuracy': b,
'r2_score': a,
'mean_squared_error': a,
'area_under_curve': b,
'mean_absolute_area': a
}
#checks your answer, no need to change this code
t.q6_check(metrics)
# -
# > **Step 6:** Now that you have identified the metrics that can be used in for regression problems, use sklearn to import them.
# Import the metrics from sklearn
from sklearn.metrics import r2_score, mean_squared_error, mean_absolute_error
# > **Step 7:** Similar to what you did with classification models, let's make sure you are comfortable with how exactly each of these metrics is being calculated. We can then match the value to what sklearn provides.
# +
def r2(actual, preds):
'''
INPUT:
actual - numpy array or pd series of actual y values
preds - numpy array or pd series of predicted y values
OUTPUT:
returns the r-squared score as a float
'''
sse = np.sum((actual-preds)**2)
sst = np.sum((actual-np.mean(actual))**2)
return 1 - sse/sst
# Check solution matches sklearn
print(r2(y_test, preds_tree))
print(r2_score(y_test, preds_tree))
print("Since the above match, we can see that we have correctly calculated the r2 value.")
# -
# > **Step 8:** Your turn fill in the function below and see if your result matches the built in for mean_squared_error.
# +
def mse(actual, preds):
'''
INPUT:
actual - numpy array or pd series of actual y values
preds - numpy array or pd series of predicted y values
OUTPUT:
returns the mean squared error as a float
'''
return np.sum((actual-preds)**2)/len(actual)
# Check your solution matches sklearn
print(mse(y_test, preds_tree))
print(mean_squared_error(y_test, preds_tree))
print("If the above match, you are all set!")
# -
# > **Step 9:** Now one last time - complete the function related to mean absolute error. Then check your function against the sklearn metric to assure they match.
# +
def mae(actual, preds):
'''
INPUT:
actual - numpy array or pd series of actual y values
preds - numpy array or pd series of predicted y values
OUTPUT:
returns the mean absolute error as a float
'''
return np.sum(np.abs(actual-preds))/len(actual)
# Check your solution matches sklearn
print(mae(y_test, preds_tree))
print(mean_absolute_error(y_test, preds_tree))
print("If the above match, you are all set!")
# -
# > **Step 10:** Which model performed the best in terms of each of the metrics? Note that r2 and mse will always match, but the mae may give a different best model. Use the dictionary and space below to match the best model via each metric.
# +
#match each metric to the model that performed best on it
a = 'decision tree'
b = 'random forest'
c = 'adaptive boosting'
d = 'linear regression'
best_fit = {
'mse': b,
'r2': b,
'mae': b
}
#Tests your answer - don't change this code
t.check_ten(best_fit)
# +
# cells for work
# -
def print_metrics(y_true, preds, model_name=None):
'''
INPUT:
y_true - the y values that are actually true in the dataset (numpy array or pandas series)
preds - the predictions for those values from some model (numpy array or pandas series)
model_name - (str - optional) a name associated with the model if you would like to add it to the print statements
OUTPUT:
None - prints the mse, mae, r2
'''
if model_name == None:
print('Mean Squared Error: ', format(mean_squared_error(y_true, preds)))
print('Mean Absolute Error: ', format(mean_absolute_error(y_true, preds)))
print('R2 Score: ', format(r2_score(y_true, preds)))
print('\n\n')
else:
print('Mean Squared Error ' + model_name + ' :' , format(mean_squared_error(y_true, preds)))
print('Mean Absolute Error ' + model_name + ' :', format(mean_absolute_error(y_true, preds)))
print('R2 Score ' + model_name + ' :', format(r2_score(y_true, preds)))
print('\n\n')
# +
# Print Decision Tree scores
print_metrics(y_test, preds_tree, 'tree')
# Print Random Forest scores
print_metrics(y_test, preds_rf, 'random forest')
# Print AdaBoost scores
print_metrics(y_test, preds_ada, 'adaboost')
# Linear Regression scores
print_metrics(y_test, preds_reg, 'linear reg')
# -
| 01_supervised_learning/4_ModelEvaluationMetrics/.ipynb_checkpoints/Regression Metrics Solution-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from os import listdir
from time import time
import matplotlib.pyplot as plt
import numpy as np
from tensorflow import math
from tensorflow.keras.callbacks import EarlyStopping, LearningRateScheduler
from tensorflow.keras.models import load_model
from tensorflow.keras.optimizers import Adam
from utils import auc, load_dataset, plot_roc, train
from einops import rearrange
# -
def schedule(epoch, lr) -> float:
if epoch >= 200 and epoch % 25 == 0:
lr = lr * math.exp(-0.1)
return lr
# +
scheduler = LearningRateScheduler(schedule)
es = EarlyStopping(monitor="loss", patience=5)
optimizer = Adam(lr=1e-4)
epochs = 1500
validation_freq = 2
# boundaries = [100, 300, 500, 1000, 2500, 5000]
fname = "model_2.h5"
# + active=""
# %%time
# for n in [1, 2, 3, 5, 10]:
# print(f'-------------- Starting {fname.replace(".h5", "")} on boundary {n}000 --------------')
# model = load_model(f'../model_saves/eval_models/{fname}', compile=False)
# X_train, y_train, X_test, y_test = load_dataset(f'MA0035_{n}000', labels="binlabels")
# X_train = rearrange(X_train, "w h c -> w c h")
# X_test = rearrange(X_test, "w h c -> w c h")
#
# model.compile(optimizer=optimizer, loss="binary_crossentropy", metrics=["acc"])
#
# model = train(dataset=(X_train, y_train, X_test, y_test),
# model=model,
# epochs=epochs,
# verbose=1,
# validation_freq=validation_freq,
# callbacks=[scheduler, es],
# )
#
# plot_roc(y_test, model.predict(X_test), [0.5])
# for boundary in boundaries:
# print(f'Boundary: {boundary}\tAUC: {auc(y_test, model.predict(X_test), boundary):.5f}')
#
# model.save(f'../model_saves/evals/{fname.replace(".h5", "")}-{n}.h5')
# -
model = load_model('../model_saves/evals/danq_small-2000.h5')
# +
X_train, y_train, X_test, y_test = load_dataset("FOX")
X_train = rearrange(X_train, "w h c -> w c h")
X_test = rearrange(X_test, "w h c -> w c h")
y_train = rearrange(np.where(y_train < 5000, 0, 1), "w h -> h w")
y_test = rearrange(np.where(y_test < 5000, 0, 1), "w h -> h w")
# -
for i, labels in enumerate(y_test):
if np.all(labels == labels[0]):
print("continue")
continue
print(i, auc(labels, model.predict(X_test)))
# print(f"{i} accuracy: {model.evaluate(X_test, labels, verbose=0)[1]}")
| src/Evaluate models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 第 3 章 字典和集合
# 正是因为字典至关重要,Python 对它的实现做了高度优化,而散列表则是字典类型性能 出众的根本原因。
# 集合(set)的实现其实也依赖于散列表。
# ## 3.1 泛映射类型
# collections.abc 模块中有 Mapping 和 MutableMapping 这两个抽象基类,它们的作用是为 dict 和其他类似的类型定义形式接口。
# 这些抽象基类的主要作用是作为形式化的文档,它们定义了构建一个映射类型所需要的最基本的接口。
# 然后它们还可以跟 isinstance 一起被用来判定某个数据是不是广义上的映射类型:
from collections import abc
my_dict = {}
isinstance(my_dict, abc.Mapping)
# 如果一个对象是可散列的,那么在这个对象的生命周期中,它的散列值是不变的,而且这个对象需要实现 __hash__() 方法。另外可散列对象还要有 __qe__() 方法,这样才能跟其他键做比较。如果两个可散列对象是相等的, 那么它们的散列值一定是一样的。
tt = (1, 2, (30, 40))
hash(tt)
tl = (1, 2, [30, 40])
hash(tl)
tf = (1, 2, frozenset([30, 40]))
hash(tf)
# 创建字典的不同方式
# +
a = dict(one=1, two=2, three=3)
b = {'one': 1, 'two': 2, 'three': 3}
c = dict(zip(['one', 'two', 'three'], [1, 2, 3]))
d = dict([('two', 2), ('one', 1), ('three', 3)])
e = dict({'three': 3, 'one': 1, 'two': 2})
a == b == c == d == e
# -
# ## 3.2 字典推导
# 字典推导(dictcomp)可以从任何以键值对作为元素的可迭代对象中构建出字典。
DIAL_CODES = [
(86, 'China'),
(91, 'India'),
(1, 'United States'),
(62, 'Indonesia'),
(55, 'Brazil'),
(92, 'Pakistan'),
(880, 'Bangladesh'),
(234, 'Nigeria'),
(7, 'Russia'),
(81, 'Japan'),
]
country_code = {country : code for code, country in DIAL_CODES} # for 后面为内容,前面为 dict 的 key 和 value
country_code
{code : country.upper() for country, code in country_code.items() if code < 66}
# + [markdown] slideshow={"slide_type": "slide"} tags=[]
# ## 3.3 常见的映射方法 —— 用setdefault处理找不到的键
# + tags=[]
"""创建一个从文件中找出单词到其出现位置的映射
返回值为 单词 [(行数, 单词首字母在行内的位置)]"""
import sys
import re
WORD_RE = re.compile(r'\w+') # 将正则表达式的样式编译为一个正则表达式对象(正则对象),可以用于匹配
# \w = [A-Za-z0-9_]
index = {}
with open('file_for_exp3_2.txt', encoding='utf-8') as fp:
for line_no, line in enumerate(fp, start = 1):
for match in WORD_RE.finditer(line):
word = match.group() # 将 Match类对象转化为 str 对象
column_no = match.start()+1
location = (line_no, column_no)
# 这其实是一种很不好的实现,这样写只是为了证明论点
occurrences = index.get(word, []) # 去 index 字典中获取 word 对应的位置的列表,如果找不到,则返回一个空列表
occurrences.append(location) # 将本次循环中找到的单词位置加入列表中
index[word] = occurrences # 将新获得的位置信息赋值给 index[word]
# 以字母顺序打印出结果
for word in sorted(index, key=str.upper):
print(word, index[word])
# +
import sys
import re
WORD_RE = re.compile(r'\w+')
index = {}
with open('file_for_exp3_2.txt', encoding='utf-8') as fp:
for line_no, line in enumerate(fp, 1):
for match in WORD_RE.finditer(line):
word = match.group()
column_no = match.start()+1
location = (line_no, column_no)
index.setdefault(word, []).append(location)
# 以字母顺序打印出结果
for word in sorted(index, key=str.upper):
print(word, index[word])
# -
# setdefault 函数的作用:
my_dict = {}
key = 'abc'
new_value = 5
# +
my_dict.setdefault(key, []).append(new_value)
my_dict
# -
# 等价于
# +
if key not in my_dict:
my_dict[key] = []
my_dict[key].append(new_value)
my_dict
# -
# 个人见解:setdefault这个函数的作用在于,当字典内没有所要的key时,将key和默认值组成的键值对放入字典中,并返回默认值。
# note: enumerate(iterable, start=0)
# 返回一个枚举对象。iterable 必须是一个序列,或 iterator,或其他支持迭代的对象。
# enumerate() 返回的迭代器的 \_\_next\_\_() 方法返回一个元组,里面包含一个计数值(从 start 开始,默认为 0)和通过迭代 iterable 获得的值。
seasons = ['Spring', 'Summer', 'Fall', 'Winter']
list(enumerate(seasons))
list(enumerate(seasons, start=1))
# note: re.finditer(pattern, string, flags=0)
# pattern 在 string 里所有的非重复匹配,返回为一个迭代器 iterator 保存了匹配对象。
# string 从左到右扫描,匹配按顺序排列。空匹配也包含在结果里。
# +
import re
string = 'a123bc1cd22d23d'
pattern = '[a-z]([0-9]*)[a-z]'
for match in re.finditer(pattern, string):
#print(type(match))
s = match.start()
e = match.end()
print('String match "%s" at %d:%d' % (string[s:e], s, e))
# -
re_obj = re.compile(pattern)
for match in re_obj.finditer(string):
s = match.start()
e = match.end()
print('String match "%s" at %d:%d' % (string[s:e], s, e))
# note: Match.group([group1, ...])
# 返回一个或者多个匹配的子组。
# 如果只有一个参数,结果就是一个字符串,如果有多个参数,结果就是一个元组(每个参数对应一个项),如果没有参数,组1默认到0(整个匹配都被返回)。
m = re.match(r"(\w+) (\w+)", "<NAME>, physicist, Hello hhh")
m.group()
m.group(0) # The entire match
m.group(1) # The first parenthesized subgroup.
m.group(2) # The second parenthesized subgroup.
m.group(1, 2) # Multiple arguments give us a tuple.
for match in re_obj.finditer(string):
word = match.group()
print(word)
print("word class : ",type(word))
print("match class : ", type(match))
| 03-dict-set/fpln_st3_1ed3_3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import tensorflow as tf
import tensorflow.keras
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Reshape, Conv1D, MaxPooling1D, Flatten, Dense, Dropout
from tensorflow.keras.utils import to_categorical
import numpy as np
import matplotlib.pyplot as plt
import os
import random
import pandas as pd
import ast
from spectra_ml.io_ import load_spectra_metadata
spectrum_len = 500 # automate this
parent_dir = os.environ['PWD']
stddata_path = os.path.join(os.environ['DATA_DIR'], "StdData-" + str(spectrum_len))
os.chdir(os.path.join(parent_dir, "lab-notebook", "smunukutla"))
# +
metadata = load_spectra_metadata(os.path.join(stddata_path,"spectra-metadata.csv"))
metadata = metadata[metadata['value_type'] == "reflectance"]
metadata = metadata[~metadata['spectrometer_purity_code'].str.contains("NIC4")]
metadata = metadata[metadata['raw_data_path'].str.contains("ChapterM")] # add in ChapterS Soils and Mixtures later
# -
metadata.sort_values('material',inplace=True)
print(metadata.to_string())
# +
names = []
num = []
frame = pd.DataFrame(columns=['material', 'count'])
series = metadata['material']
series = series.apply(lambda x: x.split(" ")[0])
series = series.value_counts()
# series = series.to_frame()
frame['count'] = series.values
frame['material'] = series.index
# frame = frame[frame['count'] >= 12]
# series.columns = ['count']
# series['material'] = series.index
# series.reset_index([], inplace=True)
# # series.columns = ['material', 'count']
# # for i in range(counts.size):
# # print(counts.index[i] + " " + str(counts[i]))
# series
frame.iloc[:, 0].tolist()
print(frame.to_string())
# -
dictionary = {frame.iloc[:, 0].tolist()[i] : i for i in range(len(frame.iloc[:, 0].tolist()))}
dictionary
| lab-notebook/smunukutla/2019-09-21-SAM - Mineral Naming.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
# ## Functions
#
# Rather than writing out everything, every time you can bundle a set of code into a function and then call the function. A function is defined using the `def` keyword.
#
# If we wanted to have a function to calculate the mean for a list we could simply define this:
def calculate_mean(some_list):
total = sum(some_list) # We could do this manually, but it's easier to use the sum function
count = len(some_list)
return total / count
# We can then use the function to carry out what we need
c = calculate_mean([1,2,3,4,5])
print(c)
# Functions return a value with the `return` statement. If a function doesn't have a `return` statement, the function returns `None`.
#
# Functions usually pass by *reference*, this means the object is changed in the function.
# +
a = dict(value=2.0)
def double_it(d):
d["double"] = d["value"] * 2
print(a)
double_it(a)
print(a)
# -
# Now an exercise for you:
# * Create a function that calculates the root mean squared error for a predicted vs an actual
#
# +
predicted = [1, 3, 6, 9, 12, 15]
actual = [0, 4, 10, 15, 20, 25]
def root_mean_squared_error(predicted_values, actual_values):
# You need to check and see whether the lists are the same length
# Calculate the deltas
# Square the values
# Calculate the mean value
# Take the square root of it
# return the value
pass
# -
# ### The importance of documentation
#
# It's always important to make use of the expressiveness of Python.
#
# Here are some general recommendations:
# * Use snake_case, rather than camelCase or PascalCase for variables
# * if you build classes use `PascalCase`
# * check out [PEP-8](https://www.python.org/dev/peps/pep-0008/#naming-conventions) for the official guidance; most editors will include some syntax checkers
# * Use **good** names; `a` is less comprehensible than `sum_of_terms`
# * This applies to functions as well as variables
# * You can use a triple quoted codeblock in the start of a function to document the function, input variables and output - editors can take advantage of this for code completion, type checking and other conveniences
#
# ```python
# def camel_to_snake(name):
# """
# Convert a CamelCase name into a snake case string
# :param str name: camel case name
# :rtype: str
# :return: the transformed name
# """
# a = re.compile(r'((?<=[a-z0-9])[A-Z]|(?!^)[A-Z](?=[a-z]))')
# return a.sub(r'_\1', name).lower()
#
# ```
# ## Python Modules
#
# As you create more and more functions it makes sense to package them up into `modules`.
#
# A python module is a package of functionality that you can use by importing it into your program when you need it. Python has a set of modules included in the core distribution - these are called the stdlib (or standard lib). They cover a superset of functionality that any programmer might need to build applications. You can check out the stdlib documentation at [Python Standard Library documentation](https://docs.python.org/3/library/).
#
# In Jupyter each cell you run updates the current environment; if you don't run the cell with the import statement then the module won't be available.
#
# As an example, we've worked with functions and loops to calculate the mean of a list of integer values; we can instead use the built in `statistics` module to calculate the mean (and some other representative statistics).
# +
from statistics import mean, mode, median, stdev
from random import randint
length = 60
a = [randint(1,35) for x in range(1, length)]
print("Random values:",a)
print("Mean:", mean(a))
print("Median:", median(a))
print("Standard Deviation:", stdev(a))
print("Mode:", mode(a))
# -
# Much simpler!
#
# We used the `randint` function from the `random` module to generate a pseudo-random integer value (between the values of 1 and 35 in this case).
#
# If you want to know more about what a module contains you can use the builtin function `dir` - this will list all the elements the module exports. In python there is no real 'data encapsulation' - no components are really private. Elements that are not expected to be used by external calls are named with underscores (eg in the list shown below
import random
dir(random)
# You can use the `help` built in function to present the documentation for a given module:
help(random.randint)
# The stdlib is in general good enough for most of what you might want to do; in cases where extensions are warranted then people create libraries and make them available through the Python Packaging Initiative. You can search for packages on [PyPI](https://pypi.org/)
#
# Python uses the module name as a namespace for the functions therein - we imported the `random` module above, but we have no way of accessing the functions directly
#
# this will give us a NameError as it has no way of looking up the function
print(randint(0, 100))
# We specify where to look for the function through the namespace
print(random.randint(0, 100))
# ## Import Syntax
#
# When you use a module, you import it into your current python environment.
#
# You can import a module:
# ```python
# import statistics
#
# statistics.mean([0,1,2,3,4,5])
# ```
# or, you can import one or more functions from a module
# ```python
# from statistics import mean
#
# mean([0,1,2,3,4,5])
# ```
# or you can import all the functions from a module
# ```python
# from statistics import *
#
# mean([0,1,2,3,4,5])
# ```
# **NOTE** - don't import all functions from a module, it loads everything into memory
#
#
# +
# Example import module and then reference
import random
print(random.randint(0, 100))
# +
from random import randint
# I can now use randint directly (ie without the module namespace)
print(randint(0, 100))
# -
# There is also syntax to allow you to import all the components of a module, although this is generally frowned upon (why load things into memory that you're never going to use).
# Don't do this!
from random import *
print(gauss(1.2, 0.2))
# If you're interested in what this function is
# help(random.gauss)
# You can also *alias* the module you import, to cut down the number of characters you need to type.
import pandas as pd
# ## Project Setup
#
# When you create a project you need to specify what dependencies the project has; the convention for doing this in Python is by use of a `requirements.txt` file. If you look in the project folder for this file you can see the following contents
#
# ```
# jupyter
# pandas
# numpy
# requests
# ```
#
# This tells the user what dependencies this project has - in this case we need jupyter to provide the notebooks we are using now; we will cover `pandas` and `numpy` in the next module (they make data engineering and data science **much** easier) and we will cover `requests` in the final module.
#
# When you share the project you should ensure that your dependencies are up to date. List what modules you have installed using the `pip` tool
# !pip list
# Note that the output of the previous command does not include only the 4 modules we listed above, the reason is that each of our dependencies will also have dependencies (the `pip list` will list all the installed modules).
#
# Now, update the `requirements.txt` to add `matplotlib` to the end of the file
# !pip install -r requirements.txt
# ## Simple Input and Output
# One of the most common activities is opening, reading and writing a file. There are a couple of libraries in the stdlib that make this simple. Firstly, we are going to use the `os` module to handle cases such as ensuring the file we are looking for exists in a platform independent way.
# +
import os
# gets the current directory
print(os.getcwd())
# get the parent directory
print(os.path.dirname(os.getcwd()))
# establish that the requirements.txt file exists
print(os.path.exists(os.path.join(os.getcwd(), "requirements.txt")))
# establish that the fruitbowl.txt file does not exist
print(os.path.exists(os.path.join(os.getcwd(), "fruitbowl.txt")))
# -
# Notice we used the `os.path.join` function above - this will join a path together in an OS independent way; on a Windows Machine it will use the `\` character and on a Linux/OSX machine it will use the `/` character.
#
# **NOTE**: Always write your code with no base assumption about where it's going to be run!
# Now we're going to open a file for reading; in this case it is a dump of conditions from the FAERS dataset and exported as a CSV
# +
# we open the file with a context manager, this with automatically close the file for us
with open("condition.csv", "r") as fh:
contents = fh.read()
# print the first 100 characters
print(contents[:100])
# -
# So, we opened the file and we can see the content. We want to be able to do some useful work with that however so we need to be able to treat the data correctly.
#
# As a first step, let's break up the file by lines
lines = contents.split('\n')
print("There are",len(lines) - 2,"conditions")
# And then split the lines into condition and count
# +
frequency = []
for line in lines[1:]:
if line:
frequency.append(line.split(','))
print(len(frequency))
print(frequency[100])
# -
# So we've loaded the contents and the parsed them out and got a list of lists; lets dig a little deeper. How many total instances of conditions are there? We can use the sum function here:
#
total = sum([int(x[1]) for x in frequency])
print("There were", total, "records")
# Ouch, that didn't work! It looks like splitting the lines based on newline characters and commas won't take into account cases where the condition includes a comma. It's time to use a module called `csv`
# +
import csv
# reset the contents
contents = []
# open the file (read-only)
with open("condition.csv", "r") as fh:
# use a DictReader, which reads in the file to a list of dicts predicated on the column headers
dr = csv.DictReader(fh)
for line in dr:
contents.append(line)
print("There are",len(contents),"conditions")
# -
# Now, let's get our count
total = sum([int(x.get('COUNT')) for x in contents])
print("There were", total, "records")
# Now, an exercise for you! Find the most commonly reported ADR from the dataset in the `condition.csv` file
# +
# define our references
max_count_value = 0
max_count_condition = None
def most_common_condition(contents):
"""
Take a list of dicts and extract the key and value for the maximum value
"""
pass
print("Condition ", max_count_condition, "had", max_count_value, "records")
# -
# ## Next
#
# Next up, we're going to briefly look at the two superstar modules for the Data Scientist of discernment, numpy and pandas. Click [here](05_numpy_pandas.ipynb) to continue
#
# <table><tr>
# <td><img src="author-geoff%20low%20small.png"></td>
# <td><img src="author-sam-hume-small.png"></td>
# </tr></table>
# <img src="Logo%20standard.png" alt="PHUSE Education" style="width: 400px;"/>
| 04_functions_and_libraries.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
diab = pd.read_csv('pima-indians-diabetes.csv')
diab.head()
diab.shape
cols = diab.columns
cols
cols_to_norm = ['Number_pregnant', 'Glucose_concentration', 'Blood_pressure', 'Triceps',
'Insulin', 'BMI', 'Pedigree']
diab[cols_to_norm] = diab[cols_to_norm].apply(lambda x:(x-x.min()) / (x.max()-x.min() ))
diab.head()
diab.columns
import tensorflow as tf
num_preg = tf.feature_column.numeric_column('Number_pregnant')
plasma_gluc = tf.feature_column.numeric_column('Glucose_concentration')
diab_press = tf.feature_column.numeric_column('Blood_pressure')
tricep = tf.feature_column.numeric_column('Triceps')
insulin = tf.feature_column.numeric_column('Insulin')
bmi = tf.feature_column.numeric_column('BMI')
diab_pedigree = tf.feature_column.numeric_column('Pedigree')
age = tf.feature_column.numeric_column('Age')
group = tf.feature_column.categorical_column_with_vocabulary_list('Group',{'A','B','C','D'})
assigned_group = tf.feature_column.categorical_column_with_hash_bucket('Group',hash_bucket_size=10)
import matplotlib.pyplot as plt
# %matplotlib inline
diab['Age'].hist(bins=20)
age_bucket = tf.feature_column.bucketized_column(age,boundaries=[20,30,40,50,60,70,80])
feat_cols = [num_preg,plasma_gluc,diab_press,tricep,insulin,bmi,diab_pedigree,assigned_group,age_bucket]
x_data = diab.drop('Class',axis=1)
labels = diab['Class']
x_data.head()
from sklearn.model_selection import train_test_split
x_train,x_test,y_train,y_test = train_test_split(x_data,labels,test_size=0.3,random_state=101)
x_train.shape
x_test.shape
input_func = tf.estimator.inputs.pandas_input_fn(x=x_train,y=y_train,num_epochs=1000,shuffle=True)
model = tf.estimator.LinearClassifier(feature_columns=feat_cols,n_classes=2)
model.train(input_fn=input_func,steps=1000)
eval_fn =tf.estimator.inputs.pandas_input_fn(x=x_test,y = y_test,batch_size=10,
num_epochs=1,shuffle=False)
result = model.evaluate(eval_fn)
result
# +
pred_func = tf.estimator.inputs.pandas_input_fn(x=x_test,y=y_test,num_epochs=1,shuffle=False)
# -
predictions = model.predict(pred_func)
preds = list(predictions)
preds
# ## DNN Model
dnn_model = tf.estimator.DNNClassifier(hidden_units=[10,20,20,20,10],feature_columns=feat_cols,
n_classes=2)
embedded_group_col = tf.feature_column.embedding_column(assigned_group,
dimension=4)
feat_cols = [num_preg,plasma_gluc,diab_press,tricep,insulin,bmi,diab_pedigree,embedded_group_col,age_bucket]
input_func = tf.estimator.inputs.pandas_input_fn(x_train,y_train,
batch_size=10,num_epochs=1000
,shuffle=True)
dnn_model = tf.estimator.DNNClassifier(hidden_units=[10,10,10],feature_columns=feat_cols,
n_classes=2)
dnn_model.train(input_func,steps=1000)
eval_input_fn = tf.estimator.inputs.pandas_input_fn(x=x_test,y=y_test
,batch_size=1,shuffle=False)
dnn_model.evaluate(eval_input_fn)
| Classification_ex.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="V9zNGvape2-I"
# # **pix2pix**
#
# ---
#
# <font size = 4>pix2pix is a deep-learning method allowing image-to-image translation from one image domain type to another image domain type. It was first published by [Isola *et al.* in 2016](https://arxiv.org/abs/1611.07004). The image transformation requires paired images for training (supervised learning) and is made possible here by using a conditional Generative Adversarial Network (GAN) architecture to use information from the input image and obtain the equivalent translated image.
#
# <font size = 4> **This particular notebook enables image-to-image translation learned from paired dataset. If you are interested in performing unpaired image-to-image translation, you should consider using the CycleGAN notebook instead.**
#
# ---
#
# <font size = 4>*Disclaimer*:
#
# <font size = 4>This notebook is part of the *Zero-Cost Deep-Learning to Enhance Microscopy* project (https://github.com/HenriquesLab/DeepLearning_Collab/wiki). Jointly developed by the Jacquemet (link to https://cellmig.org/) and Henriques (https://henriqueslab.github.io/) laboratories.
#
# <font size = 4>This notebook is based on the following paper:
#
# <font size = 4> **Image-to-Image Translation with Conditional Adversarial Networks** by Isola *et al.* on arXiv in 2016 (https://arxiv.org/abs/1611.07004)
#
# <font size = 4>The source code of the PyTorch implementation of pix2pix can be found here: https://github.com/junyanz/pytorch-CycleGAN-and-pix2pix
#
# <font size = 4>**Please also cite this original paper when using or developing this notebook.**
# + [markdown] id="N3azwKB9O0oW"
# # **License**
#
# ---
# + id="ByW6Vqdn9sYV" cellView="form"
#@markdown ##Double click to see the license information
#------------------------- LICENSE FOR ZeroCostDL4Mic------------------------------------
#This ZeroCostDL4Mic notebook is distributed under the MIT licence
#------------------------- LICENSE FOR CycleGAN ------------------------------------
#Copyright (c) 2017, <NAME> and <NAME>
#All rights reserved.
#Redistribution and use in source and binary forms, with or without
#modification, are permitted provided that the following conditions are met:
#* Redistributions of source code must retain the above copyright notice, this
# list of conditions and the following disclaimer.
#* Redistributions in binary form must reproduce the above copyright notice,
# this list of conditions and the following disclaimer in the documentation
# and/or other materials provided with the distribution.
#THIS SOFTWARE IS PROVIDED BY THE COPYRIGHT HOLDERS AND CONTRIBUTORS "AS IS"
#AND ANY EXPRESS OR IMPLIED WARRANTIES, INCLUDING, BUT NOT LIMITED TO, THE
#IMPLIED WARRANTIES OF MERCHANTABILITY AND FITNESS FOR A PARTICULAR PURPOSE ARE
#DISCLAIMED. IN NO EVENT SHALL THE COPYRIGHT HOLDER OR CONTRIBUTORS BE LIABLE
#FOR ANY DIRECT, INDIRECT, INCIDENTAL, SPECIAL, EXEMPLARY, OR CONSEQUENTIAL
#DAMAGES (INCLUDING, BUT NOT LIMITED TO, PROCUREMENT OF SUBSTITUTE GOODS OR
#SERVICES; LOSS OF USE, DATA, OR PROFITS; OR BUSINESS INTERRUPTION) HOWEVER
#CAUSED AND ON ANY THEORY OF LIABILITY, WHETHER IN CONTRACT, STRICT LIABILITY,
#OR TORT (INCLUDING NEGLIGENCE OR OTHERWISE) ARISING IN ANY WAY OUT OF THE USE
#OF THIS SOFTWARE, EVEN IF ADVISED OF THE POSSIBILITY OF SUCH DAMAGE.
#--------------------------- LICENSE FOR pix2pix --------------------------------
#BSD License
#For pix2pix software
#Copyright (c) 2016, <NAME> and <NAME>
#All rights reserved.
#Redistribution and use in source and binary forms, with or without
#modification, are permitted provided that the following conditions are met:
#* Redistributions of source code must retain the above copyright notice, this
# list of conditions and the following disclaimer.
#* Redistributions in binary form must reproduce the above copyright notice,
# this list of conditions and the following disclaimer in the documentation
# and/or other materials provided with the distribution.
#----------------------------- LICENSE FOR DCGAN --------------------------------
#BSD License
#For dcgan.torch software
#Copyright (c) 2015, Facebook, Inc. All rights reserved.
#Redistribution and use in source and binary forms, with or without modification, are permitted provided that the following conditions are met:
#Redistributions of source code must retain the above copyright notice, this list of conditions and the following disclaimer.
#Redistributions in binary form must reproduce the above copyright notice, this list of conditions and the following disclaimer in the documentation and/or other materials provided with the distribution.
#Neither the name Facebook nor the names of its contributors may be used to endorse or promote products derived from this software without specific prior written permission.
#THIS SOFTWARE IS PROVIDED BY THE COPYRIGHT HOLDERS AND CONTRIBUTORS "AS IS" AND ANY EXPRESS OR IMPLIED WARRANTIES, INCLUDING, BUT NOT LIMITED TO, THE IMPLIED WARRANTIES OF MERCHANTABILITY AND FITNESS FOR A PARTICULAR PURPOSE ARE DISCLAIMED. IN NO EVENT SHALL THE COPYRIGHT HOLDER OR CONTRIBUTORS BE LIABLE FOR ANY DIRECT, INDIRECT, INCIDENTAL, SPECIAL, EXEMPLARY, OR CONSEQUENTIAL DAMAGES (INCLUDING, BUT NOT LIMITED TO, PROCUREMENT OF SUBSTITUTE GOODS OR SERVICES; LOSS OF USE, DATA, OR PROFITS; OR BUSINESS INTERRUPTION) HOWEVER CAUSED AND ON ANY THEORY OF LIABILITY, WHETHER IN CONTRACT, STRICT LIABILITY, OR TORT (INCLUDING NEGLIGENCE OR OTHERWISE) ARISING IN ANY WAY OUT OF THE USE OF THIS SOFTWARE, EVEN IF ADVISED OF THE POSSIBILITY OF SUCH DAMAGE.
# + [markdown] id="jWAz2i7RdxUV"
# # **How to use this notebook?**
#
# ---
#
# <font size = 4>Video describing how to use our notebooks are available on youtube:
# - [**Video 1**](https://www.youtube.com/watch?v=GzD2gamVNHI&feature=youtu.be): Full run through of the workflow to obtain the notebooks and the provided test datasets as well as a common use of the notebook
# - [**Video 2**](https://www.youtube.com/watch?v=PUuQfP5SsqM&feature=youtu.be): Detailed description of the different sections of the notebook
#
#
# ---
# ###**Structure of a notebook**
#
# <font size = 4>The notebook contains two types of cell:
#
# <font size = 4>**Text cells** provide information and can be modified by douple-clicking the cell. You are currently reading the text cell. You can create a new text by clicking `+ Text`.
#
# <font size = 4>**Code cells** contain code and the code can be modfied by selecting the cell. To execute the cell, move your cursor on the `[ ]`-mark on the left side of the cell (play button appears). Click to execute the cell. After execution is done the animation of play button stops. You can create a new coding cell by clicking `+ Code`.
#
# ---
# ###**Table of contents, Code snippets** and **Files**
#
# <font size = 4>On the top left side of the notebook you find three tabs which contain from top to bottom:
#
# <font size = 4>*Table of contents* = contains structure of the notebook. Click the content to move quickly between sections.
#
# <font size = 4>*Code snippets* = contain examples how to code certain tasks. You can ignore this when using this notebook.
#
# <font size = 4>*Files* = contain all available files. After mounting your google drive (see section 1.) you will find your files and folders here.
#
# <font size = 4>**Remember that all uploaded files are purged after changing the runtime.** All files saved in Google Drive will remain. You do not need to use the Mount Drive-button; your Google Drive is connected in section 1.2.
#
# <font size = 4>**Note:** The "sample data" in "Files" contains default files. Do not upload anything in here!
#
# ---
# ###**Making changes to the notebook**
#
# <font size = 4>**You can make a copy** of the notebook and save it to your Google Drive. To do this click file -> save a copy in drive.
#
# <font size = 4>To **edit a cell**, double click on the text. This will show you either the source code (in code cells) or the source text (in text cells).
# You can use the `#`-mark in code cells to comment out parts of the code. This allows you to keep the original code piece in the cell as a comment.
# + [markdown] id="vNMDQHm0Ah-Z"
# #**0. Before getting started**
# ---
# <font size = 4> For pix2pix to train, **it needs to have access to a paired training dataset**. This means that the same image needs to be acquired in the two conditions and provided with indication of correspondence.
#
# <font size = 4> Therefore, the data structure is important. It is necessary that all the input data are in the same folder and that all the output data is in a separate folder. The provided training dataset is already split in two folders called Training_source and Training_target. Information on how to generate a training dataset is available in our Wiki page: https://github.com/HenriquesLab/ZeroCostDL4Mic/wiki
#
# <font size = 4>**We strongly recommend that you generate extra paired images. These images can be used to assess the quality of your trained model (Quality control dataset)**. The quality control assessment can be done directly in this notebook.
#
# <font size = 4> **Additionally, the corresponding input and output files need to have the same name**.
#
# <font size = 4> Please note that you currently can **only use .PNG files!**
#
#
# <font size = 4>Here's a common data structure that can work:
# * Experiment A
# - **Training dataset**
# - Training_source
# - img_1.png, img_2.png, ...
# - Training_target
# - img_1.png, img_2.png, ...
# - **Quality control dataset**
# - Training_source
# - img_1.png, img_2.png
# - Training_target
# - img_1.png, img_2.png
# - **Data to be predicted**
# - **Results**
#
# ---
# <font size = 4>**Important note**
#
# <font size = 4>- If you wish to **Train a network from scratch** using your own dataset (and we encourage everyone to do that), you will need to run **sections 1 - 4**, then use **section 5** to assess the quality of your model and **section 6** to run predictions using the model that you trained.
#
# <font size = 4>- If you wish to **Evaluate your model** using a model previously generated and saved on your Google Drive, you will only need to run **sections 1 and 2** to set up the notebook, then use **section 5** to assess the quality of your model.
#
# <font size = 4>- If you only wish to **run predictions** using a model previously generated and saved on your Google Drive, you will only need to run **sections 1 and 2** to set up the notebook, then use **section 6** to run the predictions on the desired model.
# ---
# + [markdown] id="DMNHVZfHmbKb"
# # **1. Initialise the Colab session**
# ---
#
#
#
#
#
# + [markdown] id="BCPhV-pe-syw"
#
# ## **1.1. Check for GPU access**
# ---
#
# By default, the session should be using Python 3 and GPU acceleration, but it is possible to ensure that these are set properly by doing the following:
#
# <font size = 4>Go to **Runtime -> Change the Runtime type**
#
# <font size = 4>**Runtime type: Python 3** *(Python 3 is programming language in which this program is written)*
#
# <font size = 4>**Accelator: GPU** *(Graphics processing unit)*
#
# + id="VNZetvLiS1qV" cellView="form"
#@markdown ##Run this cell to check if you have GPU access
import tensorflow as tf
if tf.test.gpu_device_name()=='':
print('You do not have GPU access.')
print('Did you change your runtime ?')
print('If the runtime setting is correct then Google did not allocate a GPU for your session')
print('Expect slow performance. To access GPU try reconnecting later')
else:
print('You have GPU access')
# !nvidia-smi
# + [markdown] id="UBrnApIUBgxv"
# ## **1.2. Mount your Google Drive**
# ---
# <font size = 4> To use this notebook on the data present in your Google Drive, you need to mount your Google Drive to this notebook.
#
# <font size = 4> Play the cell below to mount your Google Drive and follow the link. In the new browser window, select your drive and select 'Allow', copy the code, paste into the cell and press enter. This will give Colab access to the data on the drive.
#
# <font size = 4> Once this is done, your data are available in the **Files** tab on the top left of notebook.
# + id="01Djr8v-5pPk" cellView="form"
#@markdown ##Run this cell to connect your Google Drive to Colab
#@markdown * Click on the URL.
#@markdown * Sign in your Google Account.
#@markdown * Copy the authorization code.
#@markdown * Enter the authorization code.
#@markdown * Click on "Files" site on the right. Refresh the site. Your Google Drive folder should now be available here as "drive".
#mounts user's Google Drive to Google Colab.
from google.colab import drive
drive.mount('/content/gdrive')
# + [markdown] id="n4yWFoJNnoin"
# # **2. Install pix2pix and dependencies**
# ---
#
# + id="3u2mXn3XsWzd" cellView="form"
Notebook_version = ['1.11']
#@markdown ##Install pix2pix and dependencies
#Here, we install libraries which are not already included in Colab.
# !git clone https://github.com/junyanz/pytorch-CycleGAN-and-pix2pix
import os
os.chdir('pytorch-CycleGAN-and-pix2pix/')
# !pip install -r requirements.txt
# !pip install fpdf
import imageio
from skimage import data
from skimage import exposure
from skimage.exposure import match_histograms
import glob
import os.path
# ------- Common variable to all ZeroCostDL4Mic notebooks -------
import numpy as np
from matplotlib import pyplot as plt
import urllib
import os, random
import shutil
import zipfile
from tifffile import imread, imsave
import time
import sys
from pathlib import Path
import pandas as pd
import csv
from glob import glob
from scipy import signal
from scipy import ndimage
from skimage import io
from sklearn.linear_model import LinearRegression
from skimage.util import img_as_uint
import matplotlib as mpl
from skimage.metrics import structural_similarity
from skimage.metrics import peak_signal_noise_ratio as psnr
from astropy.visualization import simple_norm
from skimage import img_as_float32
from skimage.util import img_as_ubyte
from tqdm import tqdm
from fpdf import FPDF, HTMLMixin
from datetime import datetime
import subprocess
from pip._internal.operations.freeze import freeze
# Colors for the warning messages
class bcolors:
WARNING = '\033[31m'
#Disable some of the tensorflow warnings
import warnings
warnings.filterwarnings("ignore")
print('----------------------------')
print("Libraries installed")
# Check if this is the latest version of the notebook
Latest_notebook_version = pd.read_csv("https://raw.githubusercontent.com/HenriquesLab/ZeroCostDL4Mic/master/Colab_notebooks/Latest_ZeroCostDL4Mic_Release.csv")
if Notebook_version == list(Latest_notebook_version.columns):
print("This notebook is up-to-date.")
if not Notebook_version == list(Latest_notebook_version.columns):
print(bcolors.WARNING +"A new version of this notebook has been released. We recommend that you download it at https://github.com/HenriquesLab/ZeroCostDL4Mic/wiki")
# !pip freeze > requirements.txt
# + [markdown] id="Fw0kkTU6CsU4"
# # **3. Select your parameters and paths**
#
# ---
#
# + [markdown] id="BLmBseWbRvxL"
# ## **3.1. Setting main training parameters**
# ---
# <font size = 4>
#
#
#
# + [markdown] id="CB6acvUFtWqd"
# <font size = 5> **Paths for training, predictions and results**
#
# <font size = 4>**`Training_source:`, `Training_target`:** These are the paths to your folders containing the Training_source and Training_target training data respecively. To find the paths of the folders containing the respective datasets, go to your Files on the left of the notebook, navigate to the folder containing your files and copy the path by right-clicking on the folder, **Copy path** and pasting it into the right box below.
#
# <font size = 4>**`model_name`:** Use only my_model -style, not my-model (Use "_" not "-"). Do not use spaces in the name. Avoid using the name of an existing model (saved in the same folder) as it will be overwritten.
#
# <font size = 4>**`model_path`**: Enter the path where your model will be saved once trained (for instance your result folder).
#
# <font size = 5>**Training parameters**
#
# <font size = 4>**`number_of_epochs`:**Input how many epochs (rounds) the network will be trained. Preliminary results can already be observed after a few (10) epochs, but a full training should run for 200 epochs or more. Evaluate the performance after training (see 5). **Default value: 200**
#
# <font size = 5>**Advanced Parameters - experienced users only**
#
# <font size = 4>**`patch_size`:** pix2pix divides the image into patches for training. Input the size of the patches (length of a side). The value should be smaller than the dimensions of the image and divisible by 8. **Default value: 512**
#
# <font size = 4>**When choosing the patch_size, the value should be i) large enough that it will enclose many instances, ii) small enough that the resulting patches fit into the RAM.**<font size = 4>
#
# <font size =4>**`batch_size:`** This parameter defines the number of patches seen in each training step. Reducing or increasing the **batch size** may slow or speed up your training, respectively, and can influence network performance. **Default value: 1**
#
# <font size = 4>**`initial_learning_rate`:** Input the initial value to be used as learning rate. **Default value: 0.0002**
# + id="pIrTwJjzwV-D" cellView="form"
#@markdown ###Path to training images:
Training_source = "" #@param {type:"string"}
InputFile = Training_source+"/*.png"
Training_target = "" #@param {type:"string"}
OutputFile = Training_target+"/*.png"
#Define where the patch file will be saved
base = "/content"
# model name and path
#@markdown ###Name of the model and path to model folder:
model_name = "" #@param {type:"string"}
model_path = "" #@param {type:"string"}
# other parameters for training.
#@markdown ###Training Parameters
#@markdown Number of epochs:
number_of_epochs = 200#@param {type:"number"}
#@markdown ###Advanced Parameters
Use_Default_Advanced_Parameters = True #@param {type:"boolean"}
#@markdown ###If not, please input:
patch_size = 512#@param {type:"number"} # in pixels
batch_size = 1#@param {type:"number"}
initial_learning_rate = 0.0002 #@param {type:"number"}
if (Use_Default_Advanced_Parameters):
print("Default advanced parameters enabled")
batch_size = 1
patch_size = 512
initial_learning_rate = 0.0002
#here we check that no model with the same name already exist, if so delete
if os.path.exists(model_path+'/'+model_name):
print(bcolors.WARNING +"!! WARNING: "+model_name+" already exists and will be deleted in the following cell !!")
print(bcolors.WARNING +"To continue training "+model_name+", choose a new model_name here, and load "+model_name+" in section 3.3")
#To use pix2pix we need to organise the data in a way the network can understand
Saving_path= "/content/"+model_name
#Saving_path= model_path+"/"+model_name
if os.path.exists(Saving_path):
shutil.rmtree(Saving_path)
os.makedirs(Saving_path)
imageA_folder = Saving_path+"/A"
os.makedirs(imageA_folder)
imageB_folder = Saving_path+"/B"
os.makedirs(imageB_folder)
imageAB_folder = Saving_path+"/AB"
os.makedirs(imageAB_folder)
TrainA_Folder = Saving_path+"/A/train"
os.makedirs(TrainA_Folder)
TrainB_Folder = Saving_path+"/B/train"
os.makedirs(TrainB_Folder)
# Here we disable pre-trained model by default (in case the cell is not ran)
Use_pretrained_model = False
# Here we disable data augmentation by default (in case the cell is not ran)
Use_Data_augmentation = False
# This will display a randomly chosen dataset input and output
random_choice = random.choice(os.listdir(Training_source))
x = imageio.imread(Training_source+"/"+random_choice)
#Find image XY dimension
Image_Y = x.shape[0]
Image_X = x.shape[1]
Image_min_dim = min(Image_Y, Image_X)
#Hyperparameters failsafes
if patch_size > min(Image_Y, Image_X):
patch_size = min(Image_Y, Image_X)
print (bcolors.WARNING + " Your chosen patch_size is bigger than the xy dimension of your image; therefore the patch_size chosen is now:",patch_size)
# Here we check that patch_size is divisible by 4
if not patch_size % 4 == 0:
patch_size = ((int(patch_size / 4)-1) * 4)
print (bcolors.WARNING + " Your chosen patch_size is not divisible by 4; therefore the patch_size chosen is now:",patch_size)
# Here we check that patch_size is at least bigger than 256
if patch_size < 256:
patch_size = 256
print (bcolors.WARNING + " Your chosen patch_size is too small; therefore the patch_size chosen is now:",patch_size)
y = imageio.imread(Training_target+"/"+random_choice)
f=plt.figure(figsize=(16,8))
plt.subplot(1,2,1)
plt.imshow(x, interpolation='nearest')
plt.title('Training source')
plt.axis('off');
plt.subplot(1,2,2)
plt.imshow(y, interpolation='nearest')
plt.title('Training target')
plt.axis('off');
plt.savefig('/content/TrainingDataExample_pix2pix.png',bbox_inches='tight',pad_inches=0)
# + [markdown] id="5LEowmfAWqPs"
# ## **3.2. Data augmentation**
# ---
# <font size = 4>
#
#
#
#
# + [markdown] id="Flz3qoQrWv0v"
# <font size = 4>Data augmentation can improve training progress by amplifying differences in the dataset. This can be useful if the available dataset is small since, in this case, it is possible that a network could quickly learn every example in the dataset (overfitting), without augmentation. Augmentation is not necessary for training and if your training dataset is large you should disable it.
#
# <font size = 4>Data augmentation is performed here by [Augmentor.](https://github.com/mdbloice/Augmentor)
#
# <font size = 4>[Augmentor](https://github.com/mdbloice/Augmentor) was described in the following article:
#
# <font size = 4><NAME>, <NAME>, <NAME>, Biomedical image augmentation using Augmentor, Bioinformatics, https://doi.org/10.1093/bioinformatics/btz259
#
# <font size = 4>**Please also cite this original paper when publishing results obtained using this notebook with augmentation enabled.**
# + id="OsIBK-sywkfy" cellView="form"
#Data augmentation
Use_Data_augmentation = False #@param {type:"boolean"}
if Use_Data_augmentation:
# !pip install Augmentor
import Augmentor
#@markdown ####Choose a factor by which you want to multiply your original dataset
Multiply_dataset_by = 2 #@param {type:"slider", min:1, max:30, step:1}
Save_augmented_images = False #@param {type:"boolean"}
Saving_path = "" #@param {type:"string"}
Use_Default_Augmentation_Parameters = True #@param {type:"boolean"}
#@markdown ###If not, please choose the probability of the following image manipulations to be used to augment your dataset (1 = always used; 0 = disabled ):
#@markdown ####Mirror and rotate images
rotate_90_degrees = 0 #@param {type:"slider", min:0, max:1, step:0.1}
rotate_270_degrees = 0 #@param {type:"slider", min:0, max:1, step:0.1}
flip_left_right = 0 #@param {type:"slider", min:0, max:1, step:0.1}
flip_top_bottom = 0 #@param {type:"slider", min:0, max:1, step:0.1}
#@markdown ####Random image Zoom
random_zoom = 0 #@param {type:"slider", min:0, max:1, step:0.1}
random_zoom_magnification = 0 #@param {type:"slider", min:0, max:1, step:0.1}
#@markdown ####Random image distortion
random_distortion = 0 #@param {type:"slider", min:0, max:1, step:0.1}
#@markdown ####Image shearing and skewing
image_shear = 0 #@param {type:"slider", min:0, max:1, step:0.1}
max_image_shear = 10 #@param {type:"slider", min:1, max:25, step:1}
skew_image = 0 #@param {type:"slider", min:0, max:1, step:0.1}
skew_image_magnitude = 0 #@param {type:"slider", min:0, max:1, step:0.1}
if Use_Default_Augmentation_Parameters:
rotate_90_degrees = 0.5
rotate_270_degrees = 0.5
flip_left_right = 0.5
flip_top_bottom = 0.5
if not Multiply_dataset_by >5:
random_zoom = 0
random_zoom_magnification = 0.9
random_distortion = 0
image_shear = 0
max_image_shear = 10
skew_image = 0
skew_image_magnitude = 0
if Multiply_dataset_by >5:
random_zoom = 0.1
random_zoom_magnification = 0.9
random_distortion = 0.5
image_shear = 0.2
max_image_shear = 5
skew_image = 0.2
skew_image_magnitude = 0.4
if Multiply_dataset_by >25:
random_zoom = 0.5
random_zoom_magnification = 0.8
random_distortion = 0.5
image_shear = 0.5
max_image_shear = 20
skew_image = 0.5
skew_image_magnitude = 0.6
list_files = os.listdir(Training_source)
Nb_files = len(list_files)
Nb_augmented_files = (Nb_files * Multiply_dataset_by)
if Use_Data_augmentation:
print("Data augmentation enabled")
# Here we set the path for the various folder were the augmented images will be loaded
# All images are first saved into the augmented folder
#Augmented_folder = "/content/Augmented_Folder"
if not Save_augmented_images:
Saving_path= "/content"
Augmented_folder = Saving_path+"/Augmented_Folder"
if os.path.exists(Augmented_folder):
shutil.rmtree(Augmented_folder)
os.makedirs(Augmented_folder)
#Training_source_augmented = "/content/Training_source_augmented"
Training_source_augmented = Saving_path+"/Training_source_augmented"
if os.path.exists(Training_source_augmented):
shutil.rmtree(Training_source_augmented)
os.makedirs(Training_source_augmented)
#Training_target_augmented = "/content/Training_target_augmented"
Training_target_augmented = Saving_path+"/Training_target_augmented"
if os.path.exists(Training_target_augmented):
shutil.rmtree(Training_target_augmented)
os.makedirs(Training_target_augmented)
# Here we generate the augmented images
#Load the images
p = Augmentor.Pipeline(Training_source, Augmented_folder)
#Define the matching images
p.ground_truth(Training_target)
#Define the augmentation possibilities
if not rotate_90_degrees == 0:
p.rotate90(probability=rotate_90_degrees)
if not rotate_270_degrees == 0:
p.rotate270(probability=rotate_270_degrees)
if not flip_left_right == 0:
p.flip_left_right(probability=flip_left_right)
if not flip_top_bottom == 0:
p.flip_top_bottom(probability=flip_top_bottom)
if not random_zoom == 0:
p.zoom_random(probability=random_zoom, percentage_area=random_zoom_magnification)
if not random_distortion == 0:
p.random_distortion(probability=random_distortion, grid_width=4, grid_height=4, magnitude=8)
if not image_shear == 0:
p.shear(probability=image_shear,max_shear_left=20,max_shear_right=20)
if not skew_image == 0:
p.skew(probability=skew_image,magnitude=skew_image_magnitude)
p.sample(int(Nb_augmented_files))
print(int(Nb_augmented_files),"matching images generated")
# Here we sort through the images and move them back to augmented trainning source and targets folders
augmented_files = os.listdir(Augmented_folder)
for f in augmented_files:
if (f.startswith("_groundtruth_(1)_")):
shortname_noprefix = f[17:]
shutil.copyfile(Augmented_folder+"/"+f, Training_target_augmented+"/"+shortname_noprefix)
if not (f.startswith("_groundtruth_(1)_")):
shutil.copyfile(Augmented_folder+"/"+f, Training_source_augmented+"/"+f)
for filename in os.listdir(Training_source_augmented):
os.chdir(Training_source_augmented)
os.rename(filename, filename.replace('_original', ''))
#Here we clean up the extra files
shutil.rmtree(Augmented_folder)
if not Use_Data_augmentation:
print(bcolors.WARNING+"Data augmentation disabled")
# + [markdown] id="v-leE8pEWRkn"
#
# ## **3.3. Using weights from a pre-trained model as initial weights**
# ---
# <font size = 4> Here, you can set the the path to a pre-trained model from which the weights can be extracted and used as a starting point for this training session. **This pre-trained model needs to be a pix2pix model**.
#
# <font size = 4> This option allows you to perform training over multiple Colab runtimes or to do transfer learning using models trained outside of ZeroCostDL4Mic. **You do not need to run this section if you want to train a network from scratch**.
#
# + id="CbOcS3wiWV9w" cellView="form"
# @markdown ##Loading weights from a pre-trained network
Use_pretrained_model = False #@param {type:"boolean"}
#@markdown ###If not, please provide the path to the model folder:
pretrained_model_path = "" #@param {type:"string"}
# --------------------- Check if we load a previously trained model ------------------------
if Use_pretrained_model:
h5_file_path = os.path.join(pretrained_model_path, "latest_net_G.pth")
# --------------------- Check the model exist ------------------------
if not os.path.exists(h5_file_path):
print(bcolors.WARNING+'WARNING: Pretrained model does not exist')
Use_pretrained_model = False
print(bcolors.WARNING+'No pretrained network will be used.')
if os.path.exists(h5_file_path):
print("Pretrained model "+os.path.basename(pretrained_model_path)+" was found and will be loaded prior to training.")
else:
print(bcolors.WARNING+'No pretrained network will be used.')
# + [markdown] id="rQndJj70FzfL"
# # **4. Train the network**
# ---
# + [markdown] id="-A4ipz8gs3Ew"
# ## **4.1. Prepare the training data for training**
# ---
# <font size = 4>Here, we use the information from Section 3 to prepare the training data into a suitable format for training. **Your data will be copied in the google Colab "content" folder which may take some time depending on the size of your dataset.**
#
#
#
#
# + id="_V2ujGB60gDv" cellView="form"
#@markdown ##Prepare the data for training
# --------------------- Here we load the augmented data or the raw data ------------------------
if Use_Data_augmentation:
Training_source_dir = Training_source_augmented
Training_target_dir = Training_target_augmented
if not Use_Data_augmentation:
Training_source_dir = Training_source
Training_target_dir = Training_target
# --------------------- ------------------------------------------------
print("Data preparation in progress")
if os.path.exists(model_path+'/'+model_name):
shutil.rmtree(model_path+'/'+model_name)
os.makedirs(model_path+'/'+model_name)
#--------------- Here we move the files to trainA and train B ---------
print('Copying training source data...')
for f in tqdm(os.listdir(Training_source_dir)):
shutil.copyfile(Training_source_dir+"/"+f, TrainA_Folder+"/"+f)
print('Copying training target data...')
for f in tqdm(os.listdir(Training_target_dir)):
shutil.copyfile(Training_target_dir+"/"+f, TrainB_Folder+"/"+f)
#---------------------------------------------------------------------
#--------------- Here we combined A and B images---------
os.chdir("/content")
# !python pytorch-CycleGAN-and-pix2pix/datasets/combine_A_and_B.py --fold_A "$imageA_folder" --fold_B "$imageB_folder" --fold_AB "$imageAB_folder"
# pix2pix uses EPOCH without lr decay and EPOCH with lr decay, here we automatically choose half and half
number_of_epochs_lr_stable = int(number_of_epochs/2)
number_of_epochs_lr_decay = int(number_of_epochs/2)
if Use_pretrained_model :
for f in os.listdir(pretrained_model_path):
if (f.startswith("latest_net_")):
shutil.copyfile(pretrained_model_path+"/"+f, model_path+'/'+model_name+"/"+f)
print('------------------------')
print("Data ready for training")
# + [markdown] id="wQPz0F6JlvJR"
# ## **4.2. Start Training**
# ---
# <font size = 4>When playing the cell below you should see updates after each epoch (round). Network training can take some time.
#
# <font size = 4>* **CRITICAL NOTE:** Google Colab has a time limit for processing (to prevent using GPU power for datamining). Training time must be less than 12 hours! If training takes longer than 12 hours, please decrease the number of epochs or number of patches or continue the training in a second Colab session. **Pix2pix will save model checkpoints every 5 epochs.**
# + id="eBD50tAgv5qf" cellView="form"
#@markdown ##Start training
start = time.time()
os.chdir("/content")
#--------------------------------- Command line inputs to change pix2pix paramaters------------
# basic parameters
#('--dataroot', required=True, help='path to images (should have subfolders trainA, trainB, valA, valB, etc)')
#('--name', type=str, default='experiment_name', help='name of the experiment. It decides where to store samples and models')
#('--gpu_ids', type=str, default='0', help='gpu ids: e.g. 0 0,1,2, 0,2. use -1 for CPU')
#('--checkpoints_dir', type=str, default='./checkpoints', help='models are saved here')
# model parameters
#('--model', type=str, default='cycle_gan', help='chooses which model to use. [cycle_gan | pix2pix | test | colorization]')
#('--input_nc', type=int, default=3, help='# of input image channels: 3 for RGB and 1 for grayscale')
#('--output_nc', type=int, default=3, help='# of output image channels: 3 for RGB and 1 for grayscale')
#('--ngf', type=int, default=64, help='# of gen filters in the last conv layer')
#('--ndf', type=int, default=64, help='# of discrim filters in the first conv layer')
#('--netD', type=str, default='basic', help='specify discriminator architecture [basic | n_layers | pixel]. The basic model is a 70x70 PatchGAN. n_layers allows you to specify the layers in the discriminator')
#('--netG', type=str, default='resnet_9blocks', help='specify generator architecture [resnet_9blocks | resnet_6blocks | unet_256 | unet_128]')
#('--n_layers_D', type=int, default=3, help='only used if netD==n_layers')
#('--norm', type=str, default='instance', help='instance normalization or batch normalization [instance | batch | none]')
#('--init_type', type=str, default='normal', help='network initialization [normal | xavier | kaiming | orthogonal]')
#('--init_gain', type=float, default=0.02, help='scaling factor for normal, xavier and orthogonal.')
#('--no_dropout', action='store_true', help='no dropout for the generator')
# dataset parameters
#('--dataset_mode', type=str, default='unaligned', help='chooses how datasets are loaded. [unaligned | aligned | single | colorization]')
#('--direction', type=str, default='AtoB', help='AtoB or BtoA')
#('--serial_batches', action='store_true', help='if true, takes images in order to make batches, otherwise takes them randomly')
#('--num_threads', default=4, type=int, help='# threads for loading data')
#('--batch_size', type=int, default=1, help='input batch size')
#('--load_size', type=int, default=286, help='scale images to this size')
#('--crop_size', type=int, default=256, help='then crop to this size')
#('--max_dataset_size', type=int, default=float("inf"), help='Maximum number of samples allowed per dataset. If the dataset directory contains more than max_dataset_size, only a subset is loaded.')
#('--preprocess', type=str, default='resize_and_crop', help='scaling and cropping of images at load time [resize_and_crop | crop | scale_width | scale_width_and_crop | none]')
#('--no_flip', action='store_true', help='if specified, do not flip the images for data augmentation')
#('--display_winsize', type=int, default=256, help='display window size for both visdom and HTML')
# additional parameters
#('--epoch', type=str, default='latest', help='which epoch to load? set to latest to use latest cached model')
#('--load_iter', type=int, default='0', help='which iteration to load? if load_iter > 0, the code will load models by iter_[load_iter]; otherwise, the code will load models by [epoch]')
#('--verbose', action='store_true', help='if specified, print more debugging information')
#('--suffix', default='', type=str, help='customized suffix: opt.name = opt.name + suffix: e.g., {model}_{netG}_size{load_size}')
# visdom and HTML visualization parameters
#('--display_freq', type=int, default=400, help='frequency of showing training results on screen')
#('--display_ncols', type=int, default=4, help='if positive, display all images in a single visdom web panel with certain number of images per row.')
#('--display_id', type=int, default=1, help='window id of the web display')
#('--display_server', type=str, default="http://localhost", help='visdom server of the web display')
#('--display_env', type=str, default='main', help='visdom display environment name (default is "main")')
#('--display_port', type=int, default=8097, help='visdom port of the web display')
#('--update_html_freq', type=int, default=1000, help='frequency of saving training results to html')
#('--print_freq', type=int, default=100, help='frequency of showing training results on console')
#('--no_html', action='store_true', help='do not save intermediate training results to [opt.checkpoints_dir]/[opt.name]/web/')
# network saving and loading parameters
#('--save_latest_freq', type=int, default=5000, help='frequency of saving the latest results')
#('--save_epoch_freq', type=int, default=5, help='frequency of saving checkpoints at the end of epochs')
#('--save_by_iter', action='store_true', help='whether saves model by iteration')
#('--continue_train', action='store_true', help='continue training: load the latest model')
#('--epoch_count', type=int, default=1, help='the starting epoch count, we save the model by <epoch_count>, <epoch_count>+<save_latest_freq>, ...')
#('--phase', type=str, default='train', help='train, val, test, etc')
# training parameters
#('--n_epochs', type=int, default=100, help='number of epochs with the initial learning rate')
#('--n_epochs_decay', type=int, default=100, help='number of epochs to linearly decay learning rate to zero')
#('--beta1', type=float, default=0.5, help='momentum term of adam')
#('--lr', type=float, default=0.0002, help='initial learning rate for adam')
#('--gan_mode', type=str, default='lsgan', help='the type of GAN objective. [vanilla| lsgan | wgangp]. vanilla GAN loss is the cross-entropy objective used in the original GAN paper.')
#('--pool_size', type=int, default=50, help='the size of image buffer that stores previously generated images')
#('--lr_policy', type=str, default='linear', help='learning rate policy. [linear | step | plateau | cosine]')
#('--lr_decay_iters', type=int, default=50, help='multiply by a gamma every lr_decay_iters iterations'
#---------------------------------------------------------
#----- Start the training ------------------------------------
if not Use_pretrained_model:
# !python pytorch-CycleGAN-and-pix2pix/train.py --dataroot "$imageAB_folder" --name $model_name --model pix2pix --batch_size $batch_size --preprocess scale_width_and_crop --load_size $Image_min_dim --crop_size $patch_size --checkpoints_dir "$model_path" --no_html --n_epochs $number_of_epochs_lr_stable --n_epochs_decay $number_of_epochs_lr_decay --lr $initial_learning_rate --display_id 0 --save_epoch_freq 5
if Use_pretrained_model:
# !python pytorch-CycleGAN-and-pix2pix/train.py --dataroot "$imageAB_folder" --name $model_name --model pix2pix --batch_size $batch_size --preprocess scale_width_and_crop --load_size $Image_min_dim --crop_size $patch_size --checkpoints_dir "$model_path" --no_html --n_epochs $number_of_epochs_lr_stable --n_epochs_decay $number_of_epochs_lr_decay --lr $initial_learning_rate --display_id 0 --save_epoch_freq 5 --continue_train
#---------------------------------------------------------
print("Training, done.")
# Displaying the time elapsed for training
dt = time.time() - start
mins, sec = divmod(dt, 60)
hour, mins = divmod(mins, 60)
print("Time elapsed:",hour, "hour(s)",mins,"min(s)",round(sec),"sec(s)")
# save FPDF() class into a
# variable pdf
from datetime import datetime
class MyFPDF(FPDF, HTMLMixin):
pass
pdf = MyFPDF()
pdf.add_page()
pdf.set_right_margin(-1)
pdf.set_font("Arial", size = 11, style='B')
Network = 'pix2pix'
day = datetime.now()
datetime_str = str(day)[0:10]
Header = 'Training report for '+Network+' model ('+model_name+')\nDate: '+datetime_str
pdf.multi_cell(180, 5, txt = Header, align = 'L')
# add another cell
training_time = "Training time: "+str(hour)+ "hour(s) "+str(mins)+"min(s) "+str(round(sec))+"sec(s)"
pdf.cell(190, 5, txt = training_time, ln = 1, align='L')
pdf.ln(1)
Header_2 = 'Information for your materials and method:'
pdf.cell(190, 5, txt=Header_2, ln=1, align='L')
all_packages = ''
for requirement in freeze(local_only=True):
all_packages = all_packages+requirement+', '
#print(all_packages)
#Main Packages
main_packages = ''
version_numbers = []
for name in ['tensorflow','numpy','torch']:
find_name=all_packages.find(name)
main_packages = main_packages+all_packages[find_name:all_packages.find(',',find_name)]+', '
#Version numbers only here:
version_numbers.append(all_packages[find_name+len(name)+2:all_packages.find(',',find_name)])
cuda_version = subprocess.run('nvcc --version',stdout=subprocess.PIPE, shell=True)
cuda_version = cuda_version.stdout.decode('utf-8')
cuda_version = cuda_version[cuda_version.find(', V')+3:-1]
gpu_name = subprocess.run('nvidia-smi',stdout=subprocess.PIPE, shell=True)
gpu_name = gpu_name.stdout.decode('utf-8')
gpu_name = gpu_name[gpu_name.find('Tesla'):gpu_name.find('Tesla')+10]
#print(cuda_version[cuda_version.find(', V')+3:-1])
#print(gpu_name)
shape = io.imread(Training_source+'/'+os.listdir(Training_source)[1]).shape
dataset_size = len(os.listdir(Training_source))
text = 'The '+Network+' model was trained from scratch for '+str(number_of_epochs)+' epochs on '+str(dataset_size)+' paired image patches (image dimensions: '+str(shape)+', patch size: ('+str(patch_size)+','+str(patch_size)+')) with a batch size of '+str(batch_size)+' and a vanilla GAN loss function, using the '+Network+' ZeroCostDL4Mic notebook (v '+Notebook_version[0]+') (von Chamier & Laine et al., 2020). Key python packages used include tensorflow (v '+version_numbers[0]+'), numpy (v '+version_numbers[1]+'), torch (v '+version_numbers[2]+'), cuda (v '+cuda_version+'). The training was accelerated using a '+gpu_name+'GPU.'
if Use_pretrained_model:
text = 'The '+Network+' model was trained for '+str(number_of_epochs)+' epochs on '+str(dataset_size)+' paired image patches (image dimensions: '+str(shape)+', patch size: ('+str(patch_size)+','+str(patch_size)+')) with a batch size of '+str(batch_size)+' and a vanilla GAN loss function, using the '+Network+' ZeroCostDL4Mic notebook (v '+Notebook_version[0]+') (von Chamier & Laine et al., 2020). The model was retrained from a pretrained model. Key python packages used include tensorflow (v '+version_numbers[0]+'), numpy (v '+version_numbers[1]+'), torch (v '+version_numbers[2]+'), cuda (v '+cuda_version+'). The training was accelerated using a '+gpu_name+'GPU.'
pdf.set_font('')
pdf.set_font_size(10.)
pdf.multi_cell(190, 5, txt = text, align='L')
pdf.set_font('')
pdf.set_font('Arial', size = 10, style = 'B')
pdf.ln(1)
pdf.cell(28, 5, txt='Augmentation: ', ln=0)
pdf.set_font('')
if Use_Data_augmentation:
aug_text = 'The dataset was augmented by a factor of '+str(Multiply_dataset_by)+' by'
if rotate_270_degrees != 0 or rotate_90_degrees != 0:
aug_text = aug_text+'\n- rotation'
if flip_left_right != 0 or flip_top_bottom != 0:
aug_text = aug_text+'\n- flipping'
if random_zoom_magnification != 0:
aug_text = aug_text+'\n- random zoom magnification'
if random_distortion != 0:
aug_text = aug_text+'\n- random distortion'
if image_shear != 0:
aug_text = aug_text+'\n- image shearing'
if skew_image != 0:
aug_text = aug_text+'\n- image skewing'
else:
aug_text = 'No augmentation was used for training.'
pdf.multi_cell(190, 5, txt=aug_text, align='L')
pdf.set_font('Arial', size = 11, style = 'B')
pdf.ln(1)
pdf.cell(180, 5, txt = 'Parameters', align='L', ln=1)
pdf.set_font('')
pdf.set_font_size(10.)
if Use_Default_Advanced_Parameters:
pdf.cell(200, 5, txt='Default Advanced Parameters were enabled')
pdf.cell(200, 5, txt='The following parameters were used for training:')
pdf.ln(1)
html = """
<table width=40% style="margin-left:0px;">
<tr>
<th width = 50% align="left">Parameter</th>
<th width = 50% align="left">Value</th>
</tr>
<tr>
<td width = 50%>number_of_epochs</td>
<td width = 50%>{0}</td>
</tr>
<tr>
<td width = 50%>patch_size</td>
<td width = 50%>{1}</td>
</tr>
<tr>
<td width = 50%>batch_size</td>
<td width = 50%>{2}</td>
</tr>
<tr>
<td width = 50%>initial_learning_rate</td>
<td width = 50%>{3}</td>
</tr>
</table>
""".format(number_of_epochs,str(patch_size)+'x'+str(patch_size),batch_size,initial_learning_rate)
pdf.write_html(html)
#pdf.multi_cell(190, 5, txt = text_2, align='L')
pdf.set_font("Arial", size = 11, style='B')
pdf.ln(1)
pdf.cell(190, 5, txt = 'Training Dataset', align='L', ln=1)
pdf.set_font('')
pdf.set_font('Arial', size = 10, style = 'B')
pdf.cell(30, 5, txt= 'Training_source:', align = 'L', ln=0)
pdf.set_font('')
pdf.multi_cell(170, 5, txt = Training_source, align = 'L')
pdf.set_font('')
pdf.set_font('Arial', size = 10, style = 'B')
pdf.cell(29, 5, txt= 'Training_target:', align = 'L', ln=0)
pdf.set_font('')
pdf.multi_cell(170, 5, txt = Training_target, align = 'L')
#pdf.cell(190, 5, txt=aug_text, align='L', ln=1)
pdf.ln(1)
pdf.set_font('')
pdf.set_font('Arial', size = 10, style = 'B')
pdf.cell(22, 5, txt= 'Model Path:', align = 'L', ln=0)
pdf.set_font('')
pdf.multi_cell(170, 5, txt = model_path+'/'+model_name, align = 'L')
pdf.ln(1)
pdf.cell(60, 5, txt = 'Example Training pair', ln=1)
pdf.ln(1)
exp_size = io.imread('/content/TrainingDataExample_pix2pix.png').shape
pdf.image('/content/TrainingDataExample_pix2pix.png', x = 11, y = None, w = round(exp_size[1]/8), h = round(exp_size[0]/8))
pdf.ln(1)
ref_1 = 'References:\n - ZeroCostDL4Mic: <NAME>, <NAME>, Romain, et al. "ZeroCostDL4Mic: an open platform to simplify access and use of Deep-Learning in Microscopy." BioRxiv (2020).'
pdf.multi_cell(190, 5, txt = ref_1, align='L')
ref_2 = '- pix2pix: Isola, Phillip, et al. "Image-to-image translation with conditional adversarial networks." Proceedings of the IEEE conference on computer vision and pattern recognition. 2017.'
pdf.multi_cell(190, 5, txt = ref_2, align='L')
if Use_Data_augmentation:
ref_3 = '- Augmentor: Bloice, <NAME>., <NAME>, and <NAME>. "Augmentor: an image augmentation library for machine learning." arXiv preprint arXiv:1708.04680 (2017).'
pdf.multi_cell(190, 5, txt = ref_3, align='L')
pdf.ln(3)
reminder = 'Important:\nRemember to perform the quality control step on all newly trained models\nPlease consider depositing your training dataset on Zenodo'
pdf.set_font('Arial', size = 11, style='B')
pdf.multi_cell(190, 5, txt=reminder, align='C')
pdf.output(model_path+'/'+model_name+'/'+model_name+"_training_report.pdf")
# + [markdown] id="XQjQb_J_Qyku"
# ##**4.3. Download your model(s) from Google Drive**
#
#
# ---
# <font size = 4>Once training is complete, the trained model is automatically saved on your Google Drive, in the **model_path** folder that was selected in Section 3. It is however wise to download the folder as all data can be erased at the next training if using the same folder.
# + [markdown] id="2HbZd7rFqAad"
# # **5. Evaluate your model**
# ---
#
# <font size = 4>This section allows the user to perform important quality checks on the validity and generalisability of the trained model.
#
# <font size = 4>**We highly recommend to perform quality control on all newly trained models.**
#
#
#
#
#
#
#
# + [markdown] id="NEBRRG8QyEDG"
# ## **5.1. Choose the model you want to assess**
# + id="EdcnkCr9Nbl8" cellView="form"
# model name and path
#@markdown ###Do you want to assess the model you just trained ?
Use_the_current_trained_model = True #@param {type:"boolean"}
#@markdown ###If not, please provide the path to the model folder:
QC_model_folder = "" #@param {type:"string"}
#Here we define the loaded model name and path
QC_model_name = os.path.basename(QC_model_folder)
QC_model_path = os.path.dirname(QC_model_folder)
if (Use_the_current_trained_model):
QC_model_name = model_name
QC_model_path = model_path
full_QC_model_path = QC_model_path+'/'+QC_model_name+'/'
if os.path.exists(full_QC_model_path):
print("The "+QC_model_name+" network will be evaluated")
else:
W = '\033[0m' # white (normal)
R = '\033[31m' # red
print(R+'!! WARNING: The chosen model does not exist !!'+W)
print('Please make sure you provide a valid model path and model name before proceeding further.')
# + [markdown] id="ry9qN2tlydXq"
# ## **5.2. Identify the best checkpoint to use to make predictions**
# + [markdown] id="1yauWCc78HKD"
# <font size = 4> Pix2pix save model checkpoints every five epochs. Due to the stochastic nature of GAN networks, the last checkpoint is not always the best one to use. As a consequence, it can be challenging to choose the most suitable checkpoint to use to make predictions.
#
# <font size = 4>This section allows you to perform predictions using all the saved checkpoints and to estimate the quality of these predictions by comparing them to the provided ground truths images. Metric used include:
#
# <font size = 4>**1. The SSIM (structural similarity) map**
#
# <font size = 4>The SSIM metric is used to evaluate whether two images contain the same structures. It is a normalized metric and an SSIM of 1 indicates a perfect similarity between two images. Therefore for SSIM, the closer to 1, the better. The SSIM maps are constructed by calculating the SSIM metric in each pixel by considering the surrounding structural similarity in the neighbourhood of that pixel (currently defined as window of 11 pixels and with Gaussian weighting of 1.5 pixel standard deviation, see our Wiki for more info).
#
# <font size=4>**mSSIM** is the SSIM value calculated across the entire window of both images.
#
# <font size=4>**The output below shows the SSIM maps with the mSSIM**
#
# <font size = 4>**2. The RSE (Root Squared Error) map**
#
# <font size = 4>This is a display of the root of the squared difference between the normalized predicted and target or the source and the target. In this case, a smaller RSE is better. A perfect agreement between target and prediction will lead to an RSE map showing zeros everywhere (dark).
#
#
# <font size =4>**NRMSE (normalised root mean squared error)** gives the average difference between all pixels in the images compared to each other. Good agreement yields low NRMSE scores.
#
# <font size = 4>**PSNR (Peak signal-to-noise ratio)** is a metric that gives the difference between the ground truth and prediction (or source input) in decibels, using the peak pixel values of the prediction and the MSE between the images. The higher the score the better the agreement.
#
# <font size=4>**The output below shows the RSE maps with the NRMSE and PSNR values.**
#
#
# + id="2nBPucJdK3KS" cellView="form"
#@markdown ##Choose the folders that contain your Quality Control dataset
import glob
import os.path
Source_QC_folder = "" #@param{type:"string"}
Target_QC_folder = "" #@param{type:"string"}
Image_type = "Grayscale" #@param ["Grayscale", "RGB"]
# average function
def Average(lst):
return sum(lst) / len(lst)
# Create a quality control folder
if os.path.exists(QC_model_path+"/"+QC_model_name+"/Quality Control"):
shutil.rmtree(QC_model_path+"/"+QC_model_name+"/Quality Control")
os.makedirs(QC_model_path+"/"+QC_model_name+"/Quality Control")
# Create a quality control/Prediction Folder
QC_prediction_results = QC_model_path+"/"+QC_model_name+"/Quality Control/Prediction"
if os.path.exists(QC_prediction_results):
shutil.rmtree(QC_prediction_results)
os.makedirs(QC_prediction_results)
# Here we count how many images are in our folder to be predicted and we had a few
Nb_files_Data_folder = len(os.listdir(Source_QC_folder)) +10
# List images in Source_QC_folder
# This will find the image dimension of a randomly choosen image in Source_QC_folder
random_choice = random.choice(os.listdir(Source_QC_folder))
x = imageio.imread(Source_QC_folder+"/"+random_choice)
#Find image XY dimension
Image_Y = x.shape[0]
Image_X = x.shape[1]
Image_min_dim = min(Image_Y, Image_X)
# Here we need to move the data to be analysed so that pix2pix can find them
Saving_path_QC= "/content/"+QC_model_name+"_images"
if os.path.exists(Saving_path_QC):
shutil.rmtree(Saving_path_QC)
os.makedirs(Saving_path_QC)
Saving_path_QC_folder = Saving_path_QC+"/QC"
if os.path.exists(Saving_path_QC_folder):
shutil.rmtree(Saving_path_QC_folder)
os.makedirs(Saving_path_QC_folder)
imageA_folder = Saving_path_QC_folder+"/A"
os.makedirs(imageA_folder)
imageB_folder = Saving_path_QC_folder+"/B"
os.makedirs(imageB_folder)
imageAB_folder = Saving_path_QC_folder+"/AB"
os.makedirs(imageAB_folder)
testAB_folder = Saving_path_QC_folder+"/AB/test"
os.makedirs(testAB_folder)
testA_Folder = Saving_path_QC_folder+"/A/test"
os.makedirs(testA_Folder)
testB_Folder = Saving_path_QC_folder+"/B/test"
os.makedirs(testB_Folder)
QC_checkpoint_folders = "/content/"+QC_model_name
if os.path.exists(QC_checkpoint_folders):
shutil.rmtree(QC_checkpoint_folders)
os.makedirs(QC_checkpoint_folders)
for files in os.listdir(Source_QC_folder):
shutil.copyfile(Source_QC_folder+"/"+files, testA_Folder+"/"+files)
for files in os.listdir(Target_QC_folder):
shutil.copyfile(Target_QC_folder+"/"+files, testB_Folder+"/"+files)
#Here we create a merged folder containing only imageA
os.chdir("/content")
# !python pytorch-CycleGAN-and-pix2pix/datasets/combine_A_and_B.py --fold_A "$imageA_folder" --fold_B "$imageB_folder" --fold_AB "$imageAB_folder"
# This will find the image dimension of a randomly choosen image in Source_QC_folder
random_choice = random.choice(os.listdir(Source_QC_folder))
x = imageio.imread(Source_QC_folder+"/"+random_choice)
#Find image XY dimension
Image_Y = x.shape[0]
Image_X = x.shape[1]
Image_min_dim = int(min(Image_Y, Image_X))
patch_size_QC = Image_min_dim
if not patch_size_QC % 256 == 0:
patch_size_QC = ((int(patch_size_QC / 256)) * 256)
print (" Your image dimensions are not divisible by 256; therefore your images have now been resized to:",patch_size_QC)
if patch_size_QC < 256:
patch_size_QC = 256
Nb_Checkpoint = len(glob.glob(os.path.join(full_QC_model_path, '*G.pth')))
print(Nb_Checkpoint)
## Initiate list
Checkpoint_list = []
Average_ssim_score_list = []
for j in range(1, len(glob.glob(os.path.join(full_QC_model_path, '*G.pth')))+1):
checkpoints = j*5
if checkpoints == Nb_Checkpoint*5:
checkpoints = "latest"
print("The checkpoint currently analysed is ="+str(checkpoints))
Checkpoint_list.append(checkpoints)
# Create a quality control/Prediction Folder
QC_prediction_results = QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints)
if os.path.exists(QC_prediction_results):
shutil.rmtree(QC_prediction_results)
os.makedirs(QC_prediction_results)
# Create a quality control/Prediction Folder
QC_prediction_results = QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints)
if os.path.exists(QC_prediction_results):
shutil.rmtree(QC_prediction_results)
os.makedirs(QC_prediction_results)
#---------------------------- Predictions are performed here ----------------------
os.chdir("/content")
# !python pytorch-CycleGAN-and-pix2pix/test.py --dataroot "$imageAB_folder" --name "$QC_model_name" --model pix2pix --epoch $checkpoints --no_dropout --preprocess scale_width --load_size $patch_size_QC --crop_size $patch_size_QC --results_dir "$QC_prediction_results" --checkpoints_dir "$QC_model_path" --direction AtoB --num_test $Nb_files_Data_folder
#-----------------------------------------------------------------------------------
#Here we need to move the data again and remove all the unnecessary folders
Checkpoint_name = "test_"+str(checkpoints)
QC_results_images = QC_prediction_results+"/"+QC_model_name+"/"+Checkpoint_name+"/images"
QC_results_images_files = os.listdir(QC_results_images)
for f in QC_results_images_files:
shutil.copyfile(QC_results_images+"/"+f, QC_prediction_results+"/"+f)
os.chdir("/content")
#Here we clean up the extra files
shutil.rmtree(QC_prediction_results+"/"+QC_model_name)
#-------------------------------- QC for RGB ------------------------------------
if Image_type == "RGB":
# List images in Source_QC_folder
# This will find the image dimension of a randomly choosen image in Source_QC_folder
random_choice = random.choice(os.listdir(Source_QC_folder))
x = imageio.imread(Source_QC_folder+"/"+random_choice)
def ssim(img1, img2):
return structural_similarity(img1,img2,data_range=1.,full=True, multichannel=True)
# Open and create the csv file that will contain all the QC metrics
with open(QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints)+"/"+"QC_metrics_"+QC_model_name+str(checkpoints)+".csv", "w", newline='') as file:
writer = csv.writer(file)
# Write the header in the csv file
writer.writerow(["image #","Prediction v. GT mSSIM","Input v. GT mSSIM"])
# Initiate list
ssim_score_list = []
# Let's loop through the provided dataset in the QC folders
for i in os.listdir(Source_QC_folder):
if not os.path.isdir(os.path.join(Source_QC_folder,i)):
print('Running QC on: '+i)
shortname_no_PNG = i[:-4]
# -------------------------------- Target test data (Ground truth) --------------------------------
test_GT = imageio.imread(os.path.join(QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints), shortname_no_PNG+"_real_B.png"))
# -------------------------------- Source test data --------------------------------
test_source = imageio.imread(os.path.join(QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints),shortname_no_PNG+"_real_A.png"))
# -------------------------------- Prediction --------------------------------
test_prediction = imageio.imread(os.path.join(QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints),shortname_no_PNG+"_fake_B.png"))
#--------------------------- Here we normalise using histograms matching--------------------------------
test_prediction_matched = match_histograms(test_prediction, test_GT, multichannel=True)
test_source_matched = match_histograms(test_source, test_GT, multichannel=True)
# -------------------------------- Calculate the metric maps and save them --------------------------------
# Calculate the SSIM maps
index_SSIM_GTvsPrediction, img_SSIM_GTvsPrediction = ssim(test_GT, test_prediction_matched)
index_SSIM_GTvsSource, img_SSIM_GTvsSource = ssim(test_GT, test_source_matched)
ssim_score_list.append(index_SSIM_GTvsPrediction)
#Save ssim_maps
img_SSIM_GTvsPrediction_8bit = (img_SSIM_GTvsPrediction* 255).astype("uint8")
io.imsave(QC_model_path+'/'+QC_model_name+"/Quality Control/"+str(checkpoints)+"/SSIM_GTvsPrediction_"+shortname_no_PNG+'.tif',img_SSIM_GTvsPrediction_8bit)
img_SSIM_GTvsSource_8bit = (img_SSIM_GTvsSource* 255).astype("uint8")
io.imsave(QC_model_path+'/'+QC_model_name+"/Quality Control/"+str(checkpoints)+"/SSIM_GTvsSource_"+shortname_no_PNG+'.tif',img_SSIM_GTvsSource_8bit)
writer.writerow([i,str(index_SSIM_GTvsPrediction),str(index_SSIM_GTvsSource)])
#Here we calculate the ssim average for each image in each checkpoints
Average_SSIM_checkpoint = Average(ssim_score_list)
Average_ssim_score_list.append(Average_SSIM_checkpoint)
#------------------------------------------- QC for Grayscale ----------------------------------------------
if Image_type == "Grayscale":
def ssim(img1, img2):
return structural_similarity(img1,img2,data_range=1.,full=True, gaussian_weights=True, use_sample_covariance=False, sigma=1.5)
def normalize(x, pmin=3, pmax=99.8, axis=None, clip=False, eps=1e-20, dtype=np.float32):
mi = np.percentile(x,pmin,axis=axis,keepdims=True)
ma = np.percentile(x,pmax,axis=axis,keepdims=True)
return normalize_mi_ma(x, mi, ma, clip=clip, eps=eps, dtype=dtype)
def normalize_mi_ma(x, mi, ma, clip=False, eps=1e-20, dtype=np.float32):#dtype=np.float32
if dtype is not None:
x = x.astype(dtype,copy=False)
mi = dtype(mi) if np.isscalar(mi) else mi.astype(dtype,copy=False)
ma = dtype(ma) if np.isscalar(ma) else ma.astype(dtype,copy=False)
eps = dtype(eps)
try:
import numexpr
x = numexpr.evaluate("(x - mi) / ( ma - mi + eps )")
except ImportError:
x = (x - mi) / ( ma - mi + eps )
if clip:
x = np.clip(x,0,1)
return x
def norm_minmse(gt, x, normalize_gt=True):
if normalize_gt:
gt = normalize(gt, 0.1, 99.9, clip=False).astype(np.float32, copy = False)
x = x.astype(np.float32, copy=False) - np.mean(x)
#x = x - np.mean(x)
gt = gt.astype(np.float32, copy=False) - np.mean(gt)
#gt = gt - np.mean(gt)
scale = np.cov(x.flatten(), gt.flatten())[0, 1] / np.var(x.flatten())
return gt, scale * x
# Open and create the csv file that will contain all the QC metrics
with open(QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints)+"/"+"QC_metrics_"+QC_model_name+str(checkpoints)+".csv", "w", newline='') as file:
writer = csv.writer(file)
# Write the header in the csv file
writer.writerow(["image #","Prediction v. GT mSSIM","Input v. GT mSSIM", "Prediction v. GT NRMSE", "Input v. GT NRMSE", "Prediction v. GT PSNR", "Input v. GT PSNR"])
# Let's loop through the provided dataset in the QC folders
for i in os.listdir(Source_QC_folder):
if not os.path.isdir(os.path.join(Source_QC_folder,i)):
print('Running QC on: '+i)
ssim_score_list = []
shortname_no_PNG = i[:-4]
# -------------------------------- Target test data (Ground truth) --------------------------------
test_GT_raw = imageio.imread(os.path.join(QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints), shortname_no_PNG+"_real_B.png"))
test_GT = test_GT_raw[:,:,2]
# -------------------------------- Source test data --------------------------------
test_source_raw = imageio.imread(os.path.join(QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints),shortname_no_PNG+"_real_A.png"))
test_source = test_source_raw[:,:,2]
# Normalize the images wrt each other by minimizing the MSE between GT and Source image
test_GT_norm,test_source_norm = norm_minmse(test_GT, test_source, normalize_gt=True)
# -------------------------------- Prediction --------------------------------
test_prediction_raw = imageio.imread(os.path.join(QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints),shortname_no_PNG+"_fake_B.png"))
test_prediction = test_prediction_raw[:,:,2]
# Normalize the images wrt each other by minimizing the MSE between GT and prediction
test_GT_norm,test_prediction_norm = norm_minmse(test_GT, test_prediction, normalize_gt=True)
# -------------------------------- Calculate the metric maps and save them --------------------------------
# Calculate the SSIM maps
index_SSIM_GTvsPrediction, img_SSIM_GTvsPrediction = ssim(test_GT_norm, test_prediction_norm)
index_SSIM_GTvsSource, img_SSIM_GTvsSource = ssim(test_GT_norm, test_source_norm)
ssim_score_list.append(index_SSIM_GTvsPrediction)
#Save ssim_maps
img_SSIM_GTvsPrediction_8bit = (img_SSIM_GTvsPrediction* 255).astype("uint8")
io.imsave(QC_model_path+'/'+QC_model_name+"/Quality Control/"+str(checkpoints)+"/SSIM_GTvsPrediction_"+shortname_no_PNG+'.tif',img_SSIM_GTvsPrediction_8bit)
img_SSIM_GTvsSource_8bit = (img_SSIM_GTvsSource* 255).astype("uint8")
io.imsave(QC_model_path+'/'+QC_model_name+"/Quality Control/"+str(checkpoints)+"/SSIM_GTvsSource_"+shortname_no_PNG+'.tif',img_SSIM_GTvsSource_8bit)
# Calculate the Root Squared Error (RSE) maps
img_RSE_GTvsPrediction = np.sqrt(np.square(test_GT_norm - test_prediction_norm))
img_RSE_GTvsSource = np.sqrt(np.square(test_GT_norm - test_source_norm))
# Save SE maps
img_RSE_GTvsPrediction_8bit = (img_RSE_GTvsPrediction* 255).astype("uint8")
io.imsave(QC_model_path+'/'+QC_model_name+"/Quality Control/"+str(checkpoints)+"/RSE_GTvsPrediction_"+shortname_no_PNG+'.tif',img_RSE_GTvsPrediction_8bit)
img_RSE_GTvsSource_8bit = (img_RSE_GTvsSource* 255).astype("uint8")
io.imsave(QC_model_path+'/'+QC_model_name+"/Quality Control/"+str(checkpoints)+"/RSE_GTvsSource_"+shortname_no_PNG+'.tif',img_RSE_GTvsSource_8bit)
# -------------------------------- Calculate the RSE metrics and save them --------------------------------
# Normalised Root Mean Squared Error (here it's valid to take the mean of the image)
NRMSE_GTvsPrediction = np.sqrt(np.mean(img_RSE_GTvsPrediction))
NRMSE_GTvsSource = np.sqrt(np.mean(img_RSE_GTvsSource))
# We can also measure the peak signal to noise ratio between the images
PSNR_GTvsPrediction = psnr(test_GT_norm,test_prediction_norm,data_range=1.0)
PSNR_GTvsSource = psnr(test_GT_norm,test_source_norm,data_range=1.0)
writer.writerow([i,str(index_SSIM_GTvsPrediction),str(index_SSIM_GTvsSource),str(NRMSE_GTvsPrediction),str(NRMSE_GTvsSource),str(PSNR_GTvsPrediction),str(PSNR_GTvsSource)])
#Here we calculate the ssim average for each image in each checkpoints
Average_SSIM_checkpoint = Average(ssim_score_list)
Average_ssim_score_list.append(Average_SSIM_checkpoint)
# All data is now processed saved
# -------------------------------- Display --------------------------------
# Display the IoV vs Threshold plot
plt.figure(figsize=(20,5))
plt.plot(Checkpoint_list, Average_ssim_score_list, label="SSIM")
plt.title('Checkpoints vs. SSIM')
plt.ylabel('SSIM')
plt.xlabel('Checkpoints')
plt.legend()
plt.savefig(full_QC_model_path+'/Quality Control/SSIMvsCheckpoint_data.png',bbox_inches='tight',pad_inches=0)
plt.show()
# -------------------------------- Display RGB --------------------------------
from ipywidgets import interact
import ipywidgets as widgets
if Image_type == "RGB":
random_choice_shortname_no_PNG = shortname_no_PNG
@interact
def show_results(file=os.listdir(Source_QC_folder), checkpoints=Checkpoint_list):
random_choice_shortname_no_PNG = file[:-4]
df1 = pd.read_csv(QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints)+"/"+"QC_metrics_"+QC_model_name+str(checkpoints)+".csv", header=0)
df2 = df1.set_index("image #", drop = False)
index_SSIM_GTvsPrediction = df2.loc[file, "Prediction v. GT mSSIM"]
index_SSIM_GTvsSource = df2.loc[file, "Input v. GT mSSIM"]
#Setting up colours
cmap = None
plt.figure(figsize=(15,15))
# Target (Ground-truth)
plt.subplot(3,3,1)
plt.axis('off')
img_GT = imageio.imread(os.path.join(QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints), random_choice_shortname_no_PNG+"_real_B.png"), as_gray=False, pilmode="RGB")
plt.imshow(img_GT, cmap = cmap)
plt.title('Target',fontsize=15)
# Source
plt.subplot(3,3,2)
plt.axis('off')
img_Source = imageio.imread(os.path.join(QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints), random_choice_shortname_no_PNG+"_real_A.png"), as_gray=False, pilmode="RGB")
plt.imshow(img_Source, cmap = cmap)
plt.title('Source',fontsize=15)
#Prediction
plt.subplot(3,3,3)
plt.axis('off')
img_Prediction = io.imread(os.path.join(QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints), random_choice_shortname_no_PNG+"_fake_B.png"))
plt.imshow(img_Prediction, cmap = cmap)
plt.title('Prediction',fontsize=15)
#SSIM between GT and Source
plt.subplot(3,3,5)
#plt.axis('off')
plt.tick_params(
axis='both', # changes apply to the x-axis and y-axis
which='both', # both major and minor ticks are affected
bottom=False, # ticks along the bottom edge are off
top=False, # ticks along the top edge are off
left=False, # ticks along the left edge are off
right=False, # ticks along the right edge are off
labelbottom=False,
labelleft=False)
img_SSIM_GTvsSource = imageio.imread(os.path.join(QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints), "SSIM_GTvsSource_"+random_choice_shortname_no_PNG+".tif"))
imSSIM_GTvsSource = plt.imshow(img_SSIM_GTvsSource, cmap = cmap, vmin=0, vmax=1)
#plt.colorbar(imSSIM_GTvsSource,fraction=0.046, pad=0.04)
plt.title('Target vs. Source',fontsize=15)
plt.xlabel('mSSIM: '+str(round(index_SSIM_GTvsSource,3)),fontsize=14)
plt.ylabel('SSIM maps',fontsize=20, rotation=0, labelpad=75)
#SSIM between GT and Prediction
plt.subplot(3,3,6)
#plt.axis('off')
plt.tick_params(
axis='both', # changes apply to the x-axis and y-axis
which='both', # both major and minor ticks are affected
bottom=False, # ticks along the bottom edge are off
top=False, # ticks along the top edge are off
left=False, # ticks along the left edge are off
right=False, # ticks along the right edge are off
labelbottom=False,
labelleft=False)
img_SSIM_GTvsPrediction = imageio.imread(os.path.join(QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints), "SSIM_GTvsPrediction_"+random_choice_shortname_no_PNG+".tif"))
imSSIM_GTvsPrediction = plt.imshow(img_SSIM_GTvsPrediction, cmap = cmap, vmin=0,vmax=1)
#plt.colorbar(imSSIM_GTvsPrediction,fraction=0.046, pad=0.04)
plt.title('Target vs. Prediction',fontsize=15)
plt.xlabel('mSSIM: '+str(round(index_SSIM_GTvsPrediction,3)),fontsize=14)
plt.savefig(full_QC_model_path+'/Quality Control/QC_example_data.png',bbox_inches='tight',pad_inches=0)
# -------------------------------- Display Grayscale --------------------------------
if Image_type == "Grayscale":
random_choice_shortname_no_PNG = shortname_no_PNG
@interact
def show_results(file=os.listdir(Source_QC_folder), checkpoints=Checkpoint_list):
random_choice_shortname_no_PNG = file[:-4]
df1 = pd.read_csv(QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints)+"/"+"QC_metrics_"+QC_model_name+str(checkpoints)+".csv", header=0)
df2 = df1.set_index("image #", drop = False)
index_SSIM_GTvsPrediction = df2.loc[file, "Prediction v. GT mSSIM"]
index_SSIM_GTvsSource = df2.loc[file, "Input v. GT mSSIM"]
NRMSE_GTvsPrediction = df2.loc[file, "Prediction v. GT NRMSE"]
NRMSE_GTvsSource = df2.loc[file, "Input v. GT NRMSE"]
PSNR_GTvsSource = df2.loc[file, "Input v. GT PSNR"]
PSNR_GTvsPrediction = df2.loc[file, "Prediction v. GT PSNR"]
plt.figure(figsize=(20,20))
# Currently only displays the last computed set, from memory
# Target (Ground-truth)
plt.subplot(3,3,1)
plt.axis('off')
img_GT = imageio.imread(os.path.join(QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints), random_choice_shortname_no_PNG+"_real_B.png"))
plt.imshow(img_GT, norm=simple_norm(img_GT, percent = 99))
plt.title('Target',fontsize=15)
# Source
plt.subplot(3,3,2)
plt.axis('off')
img_Source = imageio.imread(os.path.join(QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints), random_choice_shortname_no_PNG+"_real_A.png"))
plt.imshow(img_Source, norm=simple_norm(img_Source, percent = 99))
plt.title('Source',fontsize=15)
#Prediction
plt.subplot(3,3,3)
plt.axis('off')
img_Prediction = io.imread(os.path.join(QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints), random_choice_shortname_no_PNG+"_fake_B.png"))
plt.imshow(img_Prediction, norm=simple_norm(img_Prediction, percent = 99))
plt.title('Prediction',fontsize=15)
#Setting up colours
cmap = plt.cm.CMRmap
#SSIM between GT and Source
plt.subplot(3,3,5)
#plt.axis('off')
plt.tick_params(
axis='both', # changes apply to the x-axis and y-axis
which='both', # both major and minor ticks are affected
bottom=False, # ticks along the bottom edge are off
top=False, # ticks along the top edge are off
left=False, # ticks along the left edge are off
right=False, # ticks along the right edge are off
labelbottom=False,
labelleft=False)
img_SSIM_GTvsSource = imageio.imread(os.path.join(QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints), "SSIM_GTvsSource_"+random_choice_shortname_no_PNG+".tif"))
img_SSIM_GTvsSource = img_SSIM_GTvsSource / 255
imSSIM_GTvsSource = plt.imshow(img_SSIM_GTvsSource, cmap = cmap, vmin=0, vmax=1)
plt.colorbar(imSSIM_GTvsSource,fraction=0.046, pad=0.04)
plt.title('Target vs. Source',fontsize=15)
plt.xlabel('mSSIM: '+str(round(index_SSIM_GTvsSource,3)),fontsize=14)
plt.ylabel('SSIM maps',fontsize=20, rotation=0, labelpad=75)
#SSIM between GT and Prediction
plt.subplot(3,3,6)
#plt.axis('off')
plt.tick_params(
axis='both', # changes apply to the x-axis and y-axis
which='both', # both major and minor ticks are affected
bottom=False, # ticks along the bottom edge are off
top=False, # ticks along the top edge are off
left=False, # ticks along the left edge are off
right=False, # ticks along the right edge are off
labelbottom=False,
labelleft=False)
img_SSIM_GTvsPrediction = imageio.imread(os.path.join(QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints), "SSIM_GTvsPrediction_"+random_choice_shortname_no_PNG+".tif"))
img_SSIM_GTvsPrediction = img_SSIM_GTvsPrediction / 255
imSSIM_GTvsPrediction = plt.imshow(img_SSIM_GTvsPrediction, cmap = cmap, vmin=0,vmax=1)
plt.colorbar(imSSIM_GTvsPrediction,fraction=0.046, pad=0.04)
plt.title('Target vs. Prediction',fontsize=15)
plt.xlabel('mSSIM: '+str(round(index_SSIM_GTvsPrediction,3)),fontsize=14)
#Root Squared Error between GT and Source
plt.subplot(3,3,8)
#plt.axis('off')
plt.tick_params(
axis='both', # changes apply to the x-axis and y-axis
which='both', # both major and minor ticks are affected
bottom=False, # ticks along the bottom edge are off
top=False, # ticks along the top edge are off
left=False, # ticks along the left edge are off
right=False, # ticks along the right edge are off
labelbottom=False,
labelleft=False)
img_RSE_GTvsSource = imageio.imread(os.path.join(QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints), "RSE_GTvsSource_"+random_choice_shortname_no_PNG+".tif"))
img_RSE_GTvsSource = img_RSE_GTvsSource / 255
imRSE_GTvsSource = plt.imshow(img_RSE_GTvsSource, cmap = cmap, vmin=0, vmax = 1)
plt.colorbar(imRSE_GTvsSource,fraction=0.046,pad=0.04)
plt.title('Target vs. Source',fontsize=15)
plt.xlabel('NRMSE: '+str(round(NRMSE_GTvsSource,3))+', PSNR: '+str(round(PSNR_GTvsSource,3)),fontsize=14)
#plt.title('Target vs. Source PSNR: '+str(round(PSNR_GTvsSource,3)))
plt.ylabel('RSE maps',fontsize=20, rotation=0, labelpad=75)
#Root Squared Error between GT and Prediction
plt.subplot(3,3,9)
#plt.axis('off')
plt.tick_params(
axis='both', # changes apply to the x-axis and y-axis
which='both', # both major and minor ticks are affected
bottom=False, # ticks along the bottom edge are off
top=False, # ticks along the top edge are off
left=False, # ticks along the left edge are off
right=False, # ticks along the right edge are off
labelbottom=False,
labelleft=False)
img_RSE_GTvsPrediction = imageio.imread(os.path.join(QC_model_path+"/"+QC_model_name+"/Quality Control/"+str(checkpoints), "RSE_GTvsPrediction_"+random_choice_shortname_no_PNG+".tif"))
img_RSE_GTvsPrediction = img_RSE_GTvsPrediction / 255
imRSE_GTvsPrediction = plt.imshow(img_RSE_GTvsPrediction, cmap = cmap, vmin=0, vmax=1)
plt.colorbar(imRSE_GTvsPrediction,fraction=0.046,pad=0.04)
plt.title('Target vs. Prediction',fontsize=15)
plt.xlabel('NRMSE: '+str(round(NRMSE_GTvsPrediction,3))+', PSNR: '+str(round(PSNR_GTvsPrediction,3)),fontsize=14)
plt.savefig(full_QC_model_path+'/Quality Control/QC_example_data.png',bbox_inches='tight',pad_inches=0)
#Make a pdf summary of the QC results
from datetime import datetime
class MyFPDF(FPDF, HTMLMixin):
pass
pdf = MyFPDF()
pdf.add_page()
pdf.set_right_margin(-1)
pdf.set_font("Arial", size = 11, style='B')
Network = 'pix2pix'
day = datetime.now()
datetime_str = str(day)[0:10]
Header = 'Quality Control report for '+Network+' model ('+QC_model_name+')\nDate: '+datetime_str
pdf.multi_cell(180, 5, txt = Header, align = 'L')
all_packages = ''
for requirement in freeze(local_only=True):
all_packages = all_packages+requirement+', '
pdf.set_font('')
pdf.set_font('Arial', size = 11, style = 'B')
pdf.ln(2)
pdf.cell(190, 5, txt = 'Development of Training Losses', ln=1, align='L')
pdf.ln(1)
exp_size = io.imread(full_QC_model_path+'/Quality Control/SSIMvsCheckpoint_data.png').shape
pdf.image(full_QC_model_path+'/Quality Control/SSIMvsCheckpoint_data.png', x = 11, y = None, w = round(exp_size[1]/8), h = round(exp_size[0]/8))
pdf.ln(2)
pdf.set_font('')
pdf.set_font('Arial', size = 10, style = 'B')
pdf.ln(3)
pdf.cell(80, 5, txt = 'Example Quality Control Visualisation', ln=1)
pdf.ln(1)
exp_size = io.imread(full_QC_model_path+'/Quality Control/QC_example_data.png').shape
if Image_type == 'RGB':
pdf.image(full_QC_model_path+'/Quality Control/QC_example_data.png', x = 16, y = None, w = round(exp_size[1]/5), h = round(exp_size[0]/5))
if Image_type == 'Grayscale':
pdf.image(full_QC_model_path+'/Quality Control/QC_example_data.png', x = 16, y = None, w = round(exp_size[1]/8), h = round(exp_size[0]/8))
pdf.ln(1)
pdf.set_font('')
pdf.set_font('Arial', size = 11, style = 'B')
pdf.ln(1)
pdf.cell(180, 5, txt = 'Quality Control Metrics', align='L', ln=1)
pdf.set_font('')
pdf.set_font_size(10.)
pdf.ln(1)
for checkpoint in os.listdir(full_QC_model_path+'/Quality Control'):
if os.path.isdir(os.path.join(full_QC_model_path,'Quality Control',checkpoint)) and checkpoint != 'Prediction':
pdf.set_font('')
pdf.set_font('Arial', size = 10, style = 'B')
pdf.cell(70, 5, txt = 'Metrics for checkpoint: '+ str(checkpoint), align='L', ln=1)
html = """
<body>
<font size="8" face="Courier New" >
<table width=95% style="margin-left:0px;">"""
with open(full_QC_model_path+'/Quality Control/'+str(checkpoint)+'/QC_metrics_'+QC_model_name+str(checkpoint)+'.csv', 'r') as csvfile:
metrics = csv.reader(csvfile)
header = next(metrics)
image = header[0]
mSSIM_PvsGT = header[1]
mSSIM_SvsGT = header[2]
header = """
<tr>
<th width = 60% align="left">{0}</th>
<th width = 20% align="center">{1}</th>
<th width = 20% align="center">{2}</th>
</tr>""".format(image,mSSIM_PvsGT,mSSIM_SvsGT)
html = html+header
for row in metrics:
image = row[0]
mSSIM_PvsGT = row[1]
mSSIM_SvsGT = row[2]
cells = """
<tr>
<td width = 60% align="left">{0}</td>
<td width = 20% align="center">{1}</td>
<td width = 20% align="center">{2}</td>
</tr>""".format(image,str(round(float(mSSIM_PvsGT),3)),str(round(float(mSSIM_SvsGT),3)))
html = html+cells
html = html+"""</body></table>"""
pdf.write_html(html)
pdf.ln(2)
else:
continue
pdf.ln(1)
pdf.set_font('')
pdf.set_font_size(10.)
ref_1 = 'References:\n - ZeroCostDL4Mic: <NAME>, Lucas & Laine, Romain, et al. "ZeroCostDL4Mic: an open platform to simplify access and use of Deep-Learning in Microscopy." BioRxiv (2020).'
pdf.multi_cell(190, 5, txt = ref_1, align='L')
ref_2 = '- pix2pix: <NAME>, et al. "Image-to-image translation with conditional adversarial networks." Proceedings of the IEEE conference on computer vision and pattern recognition. 2017.'
pdf.multi_cell(190, 5, txt = ref_2, align='L')
pdf.ln(3)
reminder = 'To find the parameters and other information about how this model was trained, go to the training_report.pdf of this model which should be in the folder of the same name.'
pdf.set_font('Arial', size = 11, style='B')
pdf.multi_cell(190, 5, txt=reminder, align='C')
pdf.output(full_QC_model_path+'/Quality Control/'+QC_model_name+'_QC_report.pdf')
# + [markdown] id="Esqnbew8uznk"
# # **6. Using the trained model**
#
# ---
#
# <font size = 4>In this section the unseen data is processed using the trained model (in section 4). First, your unseen images are uploaded and prepared for prediction. After that your trained model from section 4 is activated and finally saved into your Google Drive.
# + [markdown] id="d8wuQGjoq6eN"
# ## **6.1. Generate prediction(s) from unseen dataset**
# ---
#
# <font size = 4>The current trained model (from section 4.2) can now be used to process images. If you want to use an older model, untick the **Use_the_current_trained_model** box and enter the name and path of the model to use. Predicted output images are saved in your **Result_folder** folder as PNG images.
#
# <font size = 4>**`Data_folder`:** This folder should contain the images that you want to use your trained network on for processing.
#
# <font size = 4>**`Result_folder`:** This folder will contain the predicted output images.
#
# <font size = 4>**`checkpoint`:** Choose the checkpoint number you would like to use to perform predictions. To use the "latest" checkpoint, input "latest".
#
# + id="yb3suNkfpNA9" cellView="form"
#@markdown ### Provide the path to your dataset and to the folder where the predictions are saved, then play the cell to predict outputs from your unseen images.
import glob
import os.path
latest = "latest"
Data_folder = "" #@param {type:"string"}
Result_folder = "" #@param {type:"string"}
# model name and path
#@markdown ###Do you want to use the current trained model?
Use_the_current_trained_model = False #@param {type:"boolean"}
#@markdown ###If not, please provide the path to the model folder:
Prediction_model_folder = "" #@param {type:"string"}
#@markdown ###What model checkpoint would you like to use?
checkpoint = latest#@param {type:"raw"}
#Here we find the loaded model name and parent path
Prediction_model_name = os.path.basename(Prediction_model_folder)
Prediction_model_path = os.path.dirname(Prediction_model_folder)
#here we check if we use the newly trained network or not
if (Use_the_current_trained_model):
print("Using current trained network")
Prediction_model_name = model_name
Prediction_model_path = model_path
#here we check if the model exists
full_Prediction_model_path = Prediction_model_path+'/'+Prediction_model_name+'/'
if os.path.exists(full_Prediction_model_path):
print("The "+Prediction_model_name+" network will be used.")
else:
W = '\033[0m' # white (normal)
R = '\033[31m' # red
print(R+'!! WARNING: The chosen model does not exist !!'+W)
print('Please make sure you provide a valid model path and model name before proceeding further.')
Nb_Checkpoint = len(glob.glob(os.path.join(full_Prediction_model_path, '*G.pth')))+1
if not checkpoint == "latest":
if checkpoint < 10:
checkpoint = 5
if not checkpoint % 5 == 0:
checkpoint = ((int(checkpoint / 5)-1) * 5)
print (bcolors.WARNING + " Your chosen checkpoints is not divisible by 5; therefore the checkpoints chosen is now:",checkpoints)
if checkpoint == Nb_Checkpoint*5:
checkpoint = "latest"
if checkpoint > Nb_Checkpoint*5:
checkpoint = "latest"
# Here we need to move the data to be analysed so that pix2pix can find them
Saving_path_prediction= "/content/"+Prediction_model_name
if os.path.exists(Saving_path_prediction):
shutil.rmtree(Saving_path_prediction)
os.makedirs(Saving_path_prediction)
imageA_folder = Saving_path_prediction+"/A"
os.makedirs(imageA_folder)
imageB_folder = Saving_path_prediction+"/B"
os.makedirs(imageB_folder)
imageAB_folder = Saving_path_prediction+"/AB"
os.makedirs(imageAB_folder)
testAB_Folder = Saving_path_prediction+"/AB/test"
os.makedirs(testAB_Folder)
testA_Folder = Saving_path_prediction+"/A/test"
os.makedirs(testA_Folder)
testB_Folder = Saving_path_prediction+"/B/test"
os.makedirs(testB_Folder)
for files in os.listdir(Data_folder):
shutil.copyfile(Data_folder+"/"+files, testA_Folder+"/"+files)
shutil.copyfile(Data_folder+"/"+files, testB_Folder+"/"+files)
# Here we create a merged A / A image for the prediction
os.chdir("/content")
# !python pytorch-CycleGAN-and-pix2pix/datasets/combine_A_and_B.py --fold_A "$imageA_folder" --fold_B "$imageB_folder" --fold_AB "$imageAB_folder"
# Here we count how many images are in our folder to be predicted and we had a few
Nb_files_Data_folder = len(os.listdir(Data_folder)) +10
# This will find the image dimension of a randomly choosen image in Data_folder
random_choice = random.choice(os.listdir(Data_folder))
x = imageio.imread(Data_folder+"/"+random_choice)
#Find image XY dimension
Image_Y = x.shape[0]
Image_X = x.shape[1]
Image_min_dim = min(Image_Y, Image_X)
#-------------------------------- Perform predictions -----------------------------
#-------------------------------- Options that can be used to perform predictions -----------------------------
# basic parameters
#('--dataroot', required=True, help='path to images (should have subfolders trainA, trainB, valA, valB, etc)')
#('--name', type=str, default='experiment_name', help='name of the experiment. It decides where to store samples and models')
#('--gpu_ids', type=str, default='0', help='gpu ids: e.g. 0 0,1,2, 0,2. use -1 for CPU')
#('--checkpoints_dir', type=str, default='./checkpoints', help='models are saved here')
# model parameters
#('--model', type=str, default='cycle_gan', help='chooses which model to use. [cycle_gan | pix2pix | test | colorization]')
#('--input_nc', type=int, default=3, help='# of input image channels: 3 for RGB and 1 for grayscale')
#('--output_nc', type=int, default=3, help='# of output image channels: 3 for RGB and 1 for grayscale')
#('--ngf', type=int, default=64, help='# of gen filters in the last conv layer')
#('--ndf', type=int, default=64, help='# of discrim filters in the first conv layer')
#('--netD', type=str, default='basic', help='specify discriminator architecture [basic | n_layers | pixel]. The basic model is a 70x70 PatchGAN. n_layers allows you to specify the layers in the discriminator')
#('--netG', type=str, default='resnet_9blocks', help='specify generator architecture [resnet_9blocks | resnet_6blocks | unet_256 | unet_128]')
#('--n_layers_D', type=int, default=3, help='only used if netD==n_layers')
#('--norm', type=str, default='instance', help='instance normalization or batch normalization [instance | batch | none]')
#('--init_type', type=str, default='normal', help='network initialization [normal | xavier | kaiming | orthogonal]')
#('--init_gain', type=float, default=0.02, help='scaling factor for normal, xavier and orthogonal.')
#('--no_dropout', action='store_true', help='no dropout for the generator')
# dataset parameters
#('--dataset_mode', type=str, default='unaligned', help='chooses how datasets are loaded. [unaligned | aligned | single | colorization]')
#('--direction', type=str, default='AtoB', help='AtoB or BtoA')
#('--serial_batches', action='store_true', help='if true, takes images in order to make batches, otherwise takes them randomly')
#('--num_threads', default=4, type=int, help='# threads for loading data')
#('--batch_size', type=int, default=1, help='input batch size')
#('--load_size', type=int, default=286, help='scale images to this size')
#('--crop_size', type=int, default=256, help='then crop to this size')
#('--max_dataset_size', type=int, default=float("inf"), help='Maximum number of samples allowed per dataset. If the dataset directory contains more than max_dataset_size, only a subset is loaded.')
#('--preprocess', type=str, default='resize_and_crop', help='scaling and cropping of images at load time [resize_and_crop | crop | scale_width | scale_width_and_crop | none]')
#('--no_flip', action='store_true', help='if specified, do not flip the images for data augmentation')
#('--display_winsize', type=int, default=256, help='display window size for both visdom and HTML')
# additional parameters
#('--epoch', type=str, default='latest', help='which epoch to load? set to latest to use latest cached model')
#('--load_iter', type=int, default='0', help='which iteration to load? if load_iter > 0, the code will load models by iter_[load_iter]; otherwise, the code will load models by [epoch]')
#('--verbose', action='store_true', help='if specified, print more debugging information')
#('--suffix', default='', type=str, help='customized suffix: opt.name = opt.name + suffix: e.g., {model}_{netG}_size{load_size}')
#('--ntest', type=int, default=float("inf"), help='# of test examples.')
#('--results_dir', type=str, default='./results/', help='saves results here.')
#('--aspect_ratio', type=float, default=1.0, help='aspect ratio of result images')
#('--phase', type=str, default='test', help='train, val, test, etc')
# Dropout and Batchnorm has different behavioir during training and test.
#('--eval', action='store_true', help='use eval mode during test time.')
#('--num_test', type=int, default=50, help='how many test images to run')
# rewrite devalue values
# To avoid cropping, the load_size should be the same as crop_size
#parser.set_defaults(load_size=parser.get_default('crop_size'))
#------------------------------------------------------------------------
#---------------------------- Predictions are performed here ----------------------
os.chdir("/content")
# !python pytorch-CycleGAN-and-pix2pix/test.py --dataroot "$imageAB_folder" --name "$Prediction_model_name" --model pix2pix --no_dropout --preprocess scale_width --load_size $Image_min_dim --crop_size $Image_min_dim --results_dir "$Result_folder" --checkpoints_dir "$Prediction_model_path" --num_test $Nb_files_Data_folder --epoch $checkpoint
#-----------------------------------------------------------------------------------
Checkpoint_name = "test_"+str(checkpoint)
Prediction_results_folder = Result_folder+"/"+Prediction_model_name+"/"+Checkpoint_name+"/images"
Prediction_results_images = os.listdir(Prediction_results_folder)
for f in Prediction_results_images:
if (f.endswith("_real_B.png")):
os.remove(Prediction_results_folder+"/"+f)
# + [markdown] id="EIe3CRD7XUxa"
# ## **6.2. Inspect the predicted output**
# ---
#
#
# + id="LmDP8xiwXTTL" cellView="form"
# @markdown ##Run this cell to display a randomly chosen input and its corresponding predicted output.
import os
# This will display a randomly chosen dataset input and predicted output
random_choice = random.choice(os.listdir(Data_folder))
random_choice_no_extension = os.path.splitext(random_choice)
x = imageio.imread(Result_folder+"/"+Prediction_model_name+"/test_"+str(checkpoint)+"/images/"+random_choice_no_extension[0]+"_real_A.png")
y = imageio.imread(Result_folder+"/"+Prediction_model_name+"/test_"+str(checkpoint)+"/images/"+random_choice_no_extension[0]+"_fake_B.png")
f=plt.figure(figsize=(16,8))
plt.subplot(1,2,1)
plt.imshow(x, interpolation='nearest')
plt.title('Input')
plt.axis('off');
plt.subplot(1,2,2)
plt.imshow(y, interpolation='nearest')
plt.title('Prediction')
plt.axis('off');
# + [markdown] id="hvkd66PldsXB"
# ## **6.3. Download your predictions**
# ---
#
# <font size = 4>**Store your data** and ALL its results elsewhere by downloading it from Google Drive and after that clean the original folder tree (datasets, results, trained model etc.) if you plan to train or use new networks. Please note that the notebook will otherwise **OVERWRITE** all files which have the same name.
# + [markdown] id="Rn9zpWpo0xNw"
#
# #**Thank you for using pix2pix!**
| Colab_notebooks/pix2pix_ZeroCostDL4Mic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Stress Test
#
# The idea of this code is to see how the production Endpoint will behave when a **bunch** of requests arrive it.
# Let's simulate several users doing predictions at the same time
# +
import threading
import boto3
import numpy as np
import time
import math
from multiprocessing.pool import ThreadPool
from sklearn import datasets
# +
sm = boto3.client("sagemaker-runtime")
endpoint_name_mask='iris-model-%s'
iris = datasets.load_iris()
dataset = np.insert(iris.data, 0, iris.target,axis=1)
# +
from sagemaker.serializers import CSVSerializer
def predict(payload):
csv_serializer = CSVSerializer()
payload = payload
X = payload[1:]
y = payload[0]
elapsed_time = time.time()
resp = sm.invoke_endpoint(
EndpointName=endpoint_name_mask % env,
ContentType='text/csv',
Accept='text/csv',
Body=csv_serializer.serialize(X)
)
elapsed_time = time.time() - elapsed_time
resp = float(resp['Body'].read().decode('utf-8').strip())
return (resp == y, elapsed_time)
# -
def run_test(max_threads, max_requests):
num_batches = math.ceil(max_requests / len(dataset))
requests = []
for i in range(num_batches):
batch = dataset.copy()
np.random.shuffle(batch)
requests += batch.tolist()
len(requests)
pool = ThreadPool(max_threads)
result = pool.map(predict, requests)
pool.close()
pool.join()
correct_random_forest=0
elapsedtime_random_forest=0
for i in result:
correct_random_forest += i[0]
elapsedtime_random_forest += i[1]
print("Score classifier: {}".format(correct_random_forest/len(result)))
print("Elapsed time: {}s".format(elapsedtime_random_forest))
env='production'
# %%time
print("Starting test 1")
run_test(10, 1000)
# %%time
print("Starting test 2")
run_test(100, 10000)
# %%time
print("Starting test 3")
run_test(150, 100000000)
# > While this test is running, go to the **AWS Console** -> **Sagemaker**, then click on the **Endpoint** and then click on the **CloudWatch** monitoring logs to see the Endpoint Behavior
# ## In CloudWatch, mark the following three checkboxes
# 
# ## Then, change the following config, marked in RED
#
# 
# ## Now, while your stress test is still running, you will see the Auto Scaling Alarm like this, after 3 datapoints above 750 Invocations Per Instance
#
# 
#
# When this happens, the Endpoint Autoscaling will start adding more instances to your cluster. You can observe in the Graph from the previous image that, after new instances are added to the cluster, the **Invocations** metrics grows.
# ## Well done!
| lab/03_TestingHacking/.ipynb_checkpoints/01_Stress Test-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Checking the data
# <NAME>, February 2022
# +
# %matplotlib inline
import sys
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
sys.path.append('..')
from utils import gsearch, read_json, str_extract, int_extract
sns.set_context('talk')
sns.set_style('white')
# -
# ## Data wrangling
# Load data from main trials
data_files = gsearch('inputs/student_data*.json')
data_files.sort()
print('Found %i batches of data' % len(data_files))
print(*data_files, sep='\n')
# +
student_list = []
time_list = []
processed_workers = []
for f in data_files:
batch = str_extract('[0-9]{4}-[0-9]{2}-[0-9]{2}-[0-9]{4}', f)
print('Processing batch: %s' % batch)
student_json = read_json(f)
print('%i participants found' % len(student_json))
print(student_json[0].keys())
for stud in student_json:
worker_id = stud['worker']
if worker_id not in processed_workers:
order = stud['cond']
data = pd.DataFrame(stud['data'])
data['worker'] = worker_id
data['order'] = order
data['batch'] = batch
time = data.iloc[-1]['time_elapsed']
student_list.append(data)
time_list.append((batch, worker_id, time))
processed_workers.append(worker_id)
all_data = pd.concat(student_list).drop(columns=['trial_index', 'internal_node_id'])
student_df = all_data[all_data.trial_type == 'student-bet']
student_df = student_df.dropna(how='all', axis=1).drop(columns=['trial_type'])
student_df.to_csv('outputs/student_behavior.csv', index=False)
print(student_df.shape)
student_df.head()
# -
# Sanity check: Do all students belong to exactly one batch?
batches_per_student = student_df.groupby(['worker']).agg({'batch': 'nunique'})
print(np.all(batches_per_student.batch == 1))
# Sanity check: How many students per batch?
student_df.groupby('batch').agg({'worker': 'nunique'}).reset_index().worker
# ## Calculate bonuses
# Calculate total time:
time_elapsed = pd.DataFrame(time_list, columns=['batch', 'worker', 'time_elapsed'])
time_elapsed['t_minutes'] = time_elapsed['time_elapsed']/1000/60
time_elapsed['t_minutes'].agg(['min', 'max', 'median'])
sns.displot(time_elapsed.t_minutes)
# Calculate bonuses:
latest_batch = student_df.batch.unique()[-1]# process only for latest batch
bonuses = (
student_df
[student_df.batch == latest_batch]
.groupby('worker')
.agg({'bonus': 'sum'})
.reset_index()
)
bonuses['bonus'] = np.round(bonuses.bonus, 2)
print(bonuses.shape)
bonuses.head()
# Calculate pay per hour:
pay_per_hour = time_elapsed.merge(bonuses)
pay_per_hour['total_pay'] = pay_per_hour['bonus']+2
pay_per_hour['hourly_rate'] = pay_per_hour['total_pay']/(pay_per_hour['t_minutes']/60)
pay_stats = pay_per_hour.hourly_rate.agg(['min', 'max', 'median'])
pay_stats
# Adjust bonus payments to increase hourly rate:
bonus_file = 'outputs/student_bonuses_%s.csv' % latest_batch
print('Saving bonuses to: %s' % bonus_file)
bonuses.to_csv(bonus_file, header=False, index=False)
# ## Check assignments
# Load teacher data, for comparison:
# +
# Exclude wiggly participants
excluded = np.loadtxt('../1_preprocessing/outputs/excluded_participants.txt', dtype=str)
excluded = [int_extract('(?<=sub-)[0-9]+', s) for s in excluded]
# Clean up behavioral data
teacher_df = pd.read_csv('../2_behavioral/outputs/teaching_behavior.csv')
teacher_df = teacher_df[~teacher_df.subject.isin(excluded)]
teacher_df = teacher_df.dropna(subset=['example'], axis=0)
# Find teacher-problem pairings that were included in student task
n_hints = teacher_df.groupby(['subject', 'problem'])['example'].agg('count').reset_index()
valid_hints = (
n_hints
[n_hints.example > 1]
[['subject', 'problem']]
.rename(columns={'subject': 'teacher'})
)
valid_hints
# -
# Was each participant assigned to unique order?
orders = student_df.order.unique()
orders.sort()
orders
assignments = student_df.groupby('order').agg({'worker': 'nunique'}).reset_index()
np.all(assignments.worker == 1)
np.setdiff1d(np.arange(139), orders)
# How many ratings has each teacher received so far?
# +
# Count # of students who have seen each teacher-problem pairing
n_ratings = (
student_df
.groupby(['teacher', 'problem'])
.agg({'worker': 'nunique'})
.reset_index()
.rename(columns={'worker': 'students'})
)
n_ratings['teacher'] = n_ratings.teacher.apply(lambda s: int_extract('(?<=sub-)[0-9]+', s))
n_ratings['problem'] = n_ratings.problem.astype(int)
# Expand valid hints into a teacher x problem matrix
hint_mtx = valid_hints.merge(n_ratings, how='left').fillna(0)
hint_mtx = hint_mtx.pivot(index='problem', columns='teacher')
# -
# Plot distribution
fig,ax=plt.subplots(figsize=(16,16))
sns.heatmap(hint_mtx, square=True, cbar=False, annot=True, cmap='viridis', ax=ax)
xlabels = [item.get_text() for item in ax.get_xticklabels()]
xlabels = [int_extract('(?<=students-)[0-9]+', x) for x in xlabels]
ax.set(xlabel='Teacher', ylabel='Problem', xticklabels=xlabels)
ax.xaxis.tick_top()
ax.xaxis.set_label_position('top')
# How many unique teachers did each student see?
# +
unique_teachers = student_df.groupby('worker').agg({'teacher': 'nunique'}).reset_index()
g = sns.displot(unique_teachers.teacher, aspect=1.5)
unique_teachers['teacher'].agg(['min', 'max', 'median'])
g.axes[0,0].set(xlabel='# Unique teachers per student')
# -
# Check: How many problem-teacher combinations did we drop from the study?
dropped_hints = valid_hints.groupby('teacher').agg({'problem': 'nunique'}).reset_index()
dropped_hints['dropped'] = 40 - dropped_hints.problem
dropped_hints.groupby('dropped').agg({'teacher': 'nunique'})
# ## Check survey responses
# Task descriptions:
# +
task_descriptions = (
all_data
[all_data.trial_type == 'survey-text']
[['worker', 'response']]
)
task_descriptions['response'] = task_descriptions.response.apply(lambda r: r['Q0'])
for _, row in task_descriptions.iterrows():
print('%s: "%s"\n' % (row.worker, row.response))
# -
# Worker feedback:
posttest = (
all_data
[all_data.trial_type == 'post-survey']
.dropna(axis=1, how='all')
)
posttest.head()
| 3_student/2_check_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ---
# layout: global
# title: Clustering
# displayTitle: 聚类
# ---
#
# 本章节涵盖MLlib中的聚类算法。
# [RDD接口的聚类指南](mllib-clustering.html)也包含这些算法的相关信息。
#
# ## K均值
#
# [k-means](http://en.wikipedia.org/wiki/K-means_clustering)是一个常用的聚类算法,它将数据点按预定的簇数进行聚集。
#
# K-means算法的基本思想是:以空间中k个点为中心进行聚类,对最靠近他们的对象归类。
# 通过迭代的方法,逐次更新各聚类中心的值,直至得到最好的聚类结果。
#
# 假设要把样本集分为c个类别,算法描述如下:
#
# (1)适当选择c个类的初始中心;
#
# (2)在第k次迭代中,对任意一个样本,求其到c个中心的距离,将该样本归到距离最短的中心所在的类;
#
# (3)利用均值等方法更新该类的中心值;
#
# (4)对于所有的c个聚类中心,如果利用(2)(3)的迭代法更新后,值保持不变,则迭代结束,否则继续迭代。
#
# MLlib实现了一个[k-means++](http://en.wikipedia.org/wiki/K-means%2B%2B)方法的并行版本,叫做[kmeans||](http://theory.stanford.edu/~sergei/papers/vldb12-kmpar.pdf)。
#
# `KMeans`作为一个`Estimator`来实现,产生一个`KMeansModel`作为作为基础模型。
#
# ### 输入列
#
# <table class="table">
# <thead>
# <tr>
# <th align="left">参数名</th>
# <th align="left">类型</th>
# <th align="left">默认值</th>
# <th align="left">描述</th>
# </tr>
# </thead>
# <tbody>
# <tr>
# <td>featuresCol</td>
# <td>Vector</td>
# <td>"features"</td>
# <td>特征向量</td>
# </tr>
# </tbody>
# </table>
#
# ### 输出列
#
# <table class="table">
# <thead>
# <tr>
# <th align="left">参数名</th>
# <th align="left">类型</th>
# <th align="left">默认值</th>
# <th align="left">描述</th>
# </tr>
# </thead>
# <tbody>
# <tr>
# <td>predictionCol</td>
# <td>Int</td>
# <td>"prediction"</td>
# <td>预测的簇中心点索引</td>
# </tr>
# </tbody>
# </table>
#
# **样例**
#
# 请参考[Python API文档](api/python/pyspark.ml.html#pyspark.ml.clustering.KMeans)了解更多细节。
#
# 完整样例代码可以在[Spark仓库](https://github.com/apache/spark)中的"examples/src/main/python/ml/kmeans_example.py"找到
# +
from pyspark.ml.clustering import KMeans
from pyspark.ml.evaluation import ClusteringEvaluator
# Loads data.
dataset = spark.read.format("libsvm").load("data/mllib/sample_kmeans_data.txt")
# Trains a k-means model.
kmeans = KMeans().setK(2).setSeed(1)
model = kmeans.fit(dataset)
# Make predictions
predictions = model.transform(dataset)
# Evaluate clustering by computing Silhouette score
evaluator = ClusteringEvaluator()
silhouette = evaluator.evaluate(predictions)
print("Silhouette with squared euclidean distance = " + str(silhouette))
# Shows the result.
centers = model.clusterCenters()
print("Cluster Centers: ")
for center in centers:
print(center)
# -
# ## 隐Dirichlet分配(LDA)
#
# LDA(Latent Dirichlet Allocation)是一种文档主题生成模型,也称为一个三层贝叶斯概率模型,包含词、主题和文档三层结构。
# 所谓生成模型,就是说,我们认为一篇文章的每个词都是通过“以一定概率选择了某个主题,并从这个主题中以一定概率选择某个词语”这样一个过程得到。
# 文档到主题服从多项式分布,主题到词服从多项式分布。
#
# LDA是一种非监督机器学习技术,可以用来识别大规模文档集(document collection)或语料库(corpus)中潜藏的主题信息。它采用了词袋(bag of words)的方法,这种方法将每一篇文档视为一个词频向量,从而将文本信息转化为了易于建模的数字信息。但是词袋方法没有考虑词与词之间的顺序,这简化了问题的复杂性,同时也为模型的改进提供了契机。每一篇文档代表了一些主题所构成的一个概率分布,而每一个主题又代表了很多单词所构成的一个概率分布。
#
# `LDA`作为一个`Estimator`来实现,支持`EMLDAOptimizer`和`OnlineLDAOptimizer`,产生一个`LDAModel`作为作为基础模型。
# 高级用户需要时可以将`EMLDAOptimizer`生成的`LDAModel`转换成`DistributedLDAModel`。
#
# **样例**
#
# 请参考[Python API文档](api/python/pyspark.ml.html#pyspark.ml.clustering.LDA)了解更多细节。
#
# 完整样例代码可以在[Spark仓库](https://github.com/apache/spark)中的"examples/src/main/python/ml/lda_example.py"找到
# +
from pyspark.ml.clustering import LDA
# Loads data.
dataset = spark.read.format("libsvm").load("data/mllib/sample_lda_libsvm_data.txt")
# Trains a LDA model.
lda = LDA(k=10, maxIter=10)
model = lda.fit(dataset)
ll = model.logLikelihood(dataset)
lp = model.logPerplexity(dataset)
print("The lower bound on the log likelihood of the entire corpus: " + str(ll))
print("The upper bound on perplexity: " + str(lp))
# Describe topics.
topics = model.describeTopics(3)
print("The topics described by their top-weighted terms:")
topics.show(truncate=False)
# Shows the result
transformed = model.transform(dataset)
transformed.show(truncate=False)
# -
# ## 二分K均值
#
# 二分K均值(Bisecting k-means)是一种[层次聚类](https://en.wikipedia.org/wiki/Hierarchical_clustering)算法,使用分割(自顶向下)的逼近:所有的观察值开始是一个簇,递归地向下一个层级分裂。
#
# 分裂依据为选择能最大程度降低聚类代价函数(也就是误差平方和)的簇划分为两个簇。
# 以此进行下去,直到簇的数目等于用户给定的数目k为止。
#
# 二分K均值常常比传统K均值算法有更快的计算速度,但产生的簇群与传统K均值算法往往也是不同的。
#
# `BisectingKMeans`作为一个`Estimator`来实现,产生一个`BisectingKMeansModel`作为作为基础模型。
#
# **样例**
#
# 请参考[Python API文档](api/python/pyspark.ml.html#pyspark.ml.clustering.BisectingKMeans)了解更多细节。
#
# 完整样例代码可以在[Spark仓库](https://github.com/apache/spark)中的"examples/src/main/python/ml/bisecting_k_means_example.py"找到
# +
from pyspark.ml.clustering import BisectingKMeans
# Loads data.
dataset = spark.read.format("libsvm").load("data/mllib/sample_kmeans_data.txt")
# Trains a bisecting k-means model.
bkm = BisectingKMeans().setK(2).setSeed(1)
model = bkm.fit(dataset)
# Evaluate clustering.
cost = model.computeCost(dataset)
print("Within Set Sum of Squared Errors = " + str(cost))
# Shows the result.
print("Cluster Centers: ")
centers = model.clusterCenters()
for center in centers:
print(center)
# -
# ## 高斯混合模型(GMM)
#
# 一个高斯混合模型[Gaussian Mixture Model](http://en.wikipedia.org/wiki/Mixture_model#Multivariate_Gaussian_mixture_model)
# 表示为数据点服从*k*个子高斯分布的混合分布,每个子分布都有独立的概率。
# `spark.ml`实现使用[expectation-maximization](http://en.wikipedia.org/wiki/Expectation%E2%80%93maximization_algorithm)算法来推导给定样本下的最大似然。
#
# `GaussianMixture`作为一个`Estimator`来实现,产生一个`GaussianMixtureModel`作为作为基础模型。
#
# ### 输入列
#
# <table class="table">
# <thead>
# <tr>
# <th align="left">参数名</th>
# <th align="left">类型</th>
# <th align="left">默认值</th>
# <th align="left">描述</th>
# </tr>
# </thead>
# <tbody>
# <tr>
# <td>featuresCol</td>
# <td>Vector</td>
# <td>"features"</td>
# <td>特征向量</td>
# </tr>
# </tbody>
# </table>
#
# ### 输出列
#
# <table class="table">
# <thead>
# <tr>
# <th align="left">参数名</th>
# <th align="left">类型</th>
# <th align="left">默认值</th>
# <th align="left">描述</th>
# </tr>
# </thead>
# <tbody>
# <tr>
# <td>predictionCol</td>
# <td>Int</td>
# <td>"prediction"</td>
# <td>预测的簇中心点索引</td>
# </tr>
# <tr>
# <td>probabilityCol</td>
# <td>Vector</td>
# <td>"probability"</td>
# <td>属于每个簇的概率</td>
# </tr>
# </tbody>
# </table>
#
# **样例**
#
# 请参考[Python API文档](api/python/pyspark.ml.html#pyspark.ml.clustering.GaussianMixture)了解更多细节。
#
# 完整样例代码可以在[Spark仓库](https://github.com/apache/spark)中的"examples/src/main/python/ml/gaussian_mixture_example.py"找到
# +
from pyspark.ml.clustering import GaussianMixture
# loads data
dataset = spark.read.format("libsvm").load("data/mllib/sample_kmeans_data.txt")
gmm = GaussianMixture().setK(2).setSeed(538009335)
model = gmm.fit(dataset)
print("Gaussians shown as a DataFrame: ")
model.gaussiansDF.show(truncate=False)
| ml-clustering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="61zGMMdJNK8z" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 170} executionInfo={"status": "ok", "timestamp": 1597220323055, "user_tz": -120, "elapsed": 12080, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjrJTLi8lKnRzgdGJy_n7ioL_sYr6iEm7zO_5dG=s64", "userId": "17506803442388757883"}} outputId="0b10a476-7ef9-4960-fac0-f00986a61319"
# !apt-get install -y xvfb python-opengl x11-utils > /dev/null 2>&1
# !pip install gym pyvirtualdisplay scikit-video > /dev/null 2>&1
# %tensorflow_version 2.x
import tensorflow as tf
import numpy as np
import base64, io, time, gym
import IPython, functools
import matplotlib.pyplot as plt
from tqdm import tqdm
# !pip install mitdeeplearning
import mitdeeplearning as mdl
# + [markdown] id="nQjMzuA5NexF" colab_type="text"
# Reinforcement learning problems in general:
# 1. Initialize our environment and our agent: here we will describe the different observations and actions the agent can make in the environment.
# 2. Define our agent's memory: this will enable the agent to remember its past actions, observations, and rewards.
# 3. Define a reward function: describes the reward associated with an action or sequence of actions.
# 4. Define the learning algorithm: this will be used to reinforce the agent's good behaviors and discourage bad behaviors.
# + [markdown] id="HZpzfV-QOBIv" colab_type="text"
# # Cartpole
# + id="PHjZ9yppNK86" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1597220325103, "user_tz": -120, "elapsed": 585, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjrJTLi8lKnRzgdGJy_n7ioL_sYr6iEm7zO_5dG=s64", "userId": "17506803442388757883"}} outputId="44708f2b-2cf4-48c4-dad1-05c377002191"
env = gym.make("CartPole-v0")
env.seed(1) # initialize a same random env every time
# + [markdown] id="34g3GW8jO_32" colab_type="text"
# First, let's consider the observation space. In this Cartpole environment our observations are:
# 1. Cart position
# 2. Cart velocity
# 3. Pole angle
# 4. Pole rotation rate
# + id="RaC0oAX5Oy35" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1597220327727, "user_tz": -120, "elapsed": 553, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjrJTLi8lKnRzgdGJy_n7ioL_sYr6iEm7zO_5dG=s64", "userId": "17506803442388757883"}} outputId="f8938a4d-653e-43cf-99eb-69e413ad5aad"
# confirm by querying the environment
n_observations = env.observation_space
print("Environment has observation space =", n_observations)
# + id="OUjviw8NPdL8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1597220329303, "user_tz": -120, "elapsed": 552, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjrJTLi8lKnRzgdGJy_n7ioL_sYr6iEm7zO_5dG=s64", "userId": "17506803442388757883"}} outputId="03ab19cd-18e2-4f2c-dbe3-c8c894eb760c"
n_actions = env.action_space.n
print("Number of possible actions that the agent can choose from =", n_actions)
# + id="d7VZu5VDP9Ib" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1597220330147, "user_tz": -120, "elapsed": 501, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjrJTLi8lKnRzgdGJy_n7ioL_sYr6iEm7zO_5dG=s64", "userId": "17506803442388757883"}}
# define the agent
def create_cartpole_model():
model = tf.keras.models.Sequential([
tf.keras.layers.Dense(units=32, activation='relu'),
tf.keras.layers.Dense(units=n_actions, activation=None)
])
return model
cartpole_model = create_cartpole_model()
# + id="-w85hO9ZRXFF" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1597220332055, "user_tz": -120, "elapsed": 579, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjrJTLi8lKnRzgdGJy_n7ioL_sYr6iEm7zO_5dG=s64", "userId": "17506803442388757883"}}
# define the agent action function
def choose_action(model, observation):
observation = np.expand_dims(observation, axis=0)
logits = model.predict(observation)
prob_weights = tf.nn.softmax(logits).numpy()
# print("Check dimensions: ", observation.shape, logits.shape, prob_weights.shape)
action = np.random.choice(n_actions, size=1, p=prob_weights.flatten())[0]
# print("What action did it choose?", action)
return action
# + id="WOwthOfqUN53" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1597220333559, "user_tz": -120, "elapsed": 490, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjrJTLi8lKnRzgdGJy_n7ioL_sYr6iEm7zO_5dG=s64", "userId": "17506803442388757883"}}
# agent memory
class Memory:
def __init__(self):
self.clear()
def clear(self):
# reset memory buffer
self.observations = []
self.actions = []
self.rewards = []
def add_to_memory(self, new_observation, new_action, new_reward):
self.observations.append(new_observation)
self.actions.append(new_action)
self.rewards.append(new_reward)
memory = Memory()
# + id="vZmneN2CVHtk" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1597220338496, "user_tz": -120, "elapsed": 519, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjrJTLi8lKnRzgdGJy_n7ioL_sYr6iEm7zO_5dG=s64", "userId": "17506803442388757883"}}
# reward function
def normalize(x):
x -= np.mean(x)
x /= np.std(x)
return x.astype(np.float32)
def discount_rewards(rewards, gamma=0.95):
discounted_rewards = np.zeros_like(rewards)
R = 0
for t in reversed(range(0, len(rewards))): # prefer to get reward now instead of long term
R = R * gamma + rewards[t]
discounted_rewards[t] = R
return normalize(discounted_rewards)
# + [markdown] id="XoSO-tNyW78R" colab_type="text"
# Now define the learning steps
# + id="ptduEAn1W0PE" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1597220336369, "user_tz": -120, "elapsed": 664, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjrJTLi8lKnRzgdGJy_n7ioL_sYr6iEm7zO_5dG=s64", "userId": "17506803442388757883"}}
# loss
def compute_loss(logits, actions, rewards):
neg_logprob = tf.nn.sparse_softmax_cross_entropy_with_logits(logits=logits, labels=actions)
loss = tf.reduce_mean(neg_logprob * rewards)
return loss
# + id="BAQDf6KMYMe-" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1597220345121, "user_tz": -120, "elapsed": 497, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjrJTLi8lKnRzgdGJy_n7ioL_sYr6iEm7zO_5dG=s64", "userId": "17506803442388757883"}}
# training step
def train_step(model, optimizer, observations, actions, discounted_rewards):
with tf.GradientTape() as tape:
logits = model(observations)
loss = compute_loss(logits, actions, discounted_rewards)
grads = tape.gradient(loss, model.trainable_variables)
optimizer.apply_gradients(zip(grads, model.trainable_variables))
# + id="mlYX3y6pZIx3" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 541} executionInfo={"status": "ok", "timestamp": 1597219830506, "user_tz": -120, "elapsed": 524094, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjrJTLi8lKnRzgdGJy_n7ioL_sYr6iEm7zO_5dG=s64", "userId": "17506803442388757883"}} outputId="2f3efa23-d6d8-40ae-8c3a-edd37ef55a73"
# start training!
learning_rate = 1e-3
optimizer = tf.keras.optimizers.Adam(learning_rate)
cartpole_model = create_cartpole_model()
smoothed_reward = mdl.util.LossHistory(smoothing_factor=0.9)
plotter = mdl.util.PeriodicPlotter(sec=2, xlabel="Iterations", ylabel='Rewards')
if hasattr(tqdm, '_instances'): tqdm._instances.clear() # clear if it exists
for i_episode in range(500):
plotter.plot(smoothed_reward.get())
observation = env.reset()
memory.clear()
while True:
action = choose_action(cartpole_model, observation)
next_observation, reward, done, info = env.step(action)
memory.add_to_memory(next_observation, action, reward)
if done:
total_reward = sum(memory.rewards)
smoothed_reward.append(total_reward)
train_step(
cartpole_model,
optimizer,
observations = np.vstack(memory.observations),
actions = np.array(memory.actions),
discounted_rewards = discount_rewards(memory.rewards)
)
memory.clear()
break
observation = next_observation
# + id="5Mdpo2bSpc9Y" colab_type="code" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1597219832484, "user_tz": -120, "elapsed": 525822, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjrJTLi8lKnRzgdGJy_n7ioL_sYr6iEm7zO_5dG=s64", "userId": "17506803442388757883"}} outputId="304f74f7-bd67-4eca-b51b-0b277ff71fd2"
# display the learning result
saved_cartpole = mdl.lab3.save_video_of_model(cartpole_model, "CartPole-v0")
mdl.lab3.play_video(saved_cartpole)
# + [markdown] id="TgG9pYSZqBCQ" colab_type="text"
# # Pong
# + id="rLtOsg6bpshg" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} executionInfo={"status": "ok", "timestamp": 1597220352209, "user_tz": -120, "elapsed": 927, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjrJTLi8lKnRzgdGJy_n7ioL_sYr6iEm7zO_5dG=s64", "userId": "17506803442388757883"}} outputId="7143f198-1146-429c-e272-e03e34b1842f"
env = gym.make("Pong-v0", frameskip=5)
env.seed(1)
print("Environment has observation space=", env.observation_space)
n_actions = env.action_space.n
print("Number of actions that agent can choose from =", n_actions)
# + id="OE25Ko5rqgmi" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1597220353454, "user_tz": -120, "elapsed": 540, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjrJTLi8lKnRzgdGJy_n7ioL_sYr6iEm7zO_5dG=s64", "userId": "17506803442388757883"}}
# pong agent
Conv2D = functools.partial(tf.keras.layers.Conv2D, padding='same', activation='relu')
Flatten = tf.keras.layers.Flatten
Dense = tf.keras.layers.Dense
def create_pong_model():
model = tf.keras.models.Sequential([
Conv2D(filters=16, kernel_size=7, strides=4),
Conv2D(filters=32, kernel_size=5, strides=2),
Conv2D(filters=48, kernel_size=3, strides=2),
Flatten(),
Dense(units=64, activation='relu'),
Dense(units=n_actions, activation=None)
])
return model
pong_model = create_pong_model()
# + id="fJIOXXilsKQk" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1597220355327, "user_tz": -120, "elapsed": 497, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjrJTLi8lKnRzgdGJy_n7ioL_sYr6iEm7zO_5dG=s64", "userId": "17506803442388757883"}}
# Pong reward function
def discount_rewards(rewards, gamma=0.99):
discounted_rewards = np.zeros_like(rewards)
R = 0
for t in reversed(range(0, len(rewards))):
# reset the rewards to 0 in two cases: win (1), lose (-1)
if rewards[t] != 0:
R = 0
R = R * gamma + rewards[t]
discounted_rewards[t] = R
return normalize(discounted_rewards)
# + id="f5GXBWW_tk3m" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 244} executionInfo={"status": "ok", "timestamp": 1597220358713, "user_tz": -120, "elapsed": 1143, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjrJTLi8lKnRzgdGJy_n7ioL_sYr6iEm7zO_5dG=s64", "userId": "17506803442388757883"}} outputId="3f9dea98-9c1b-4b93-8fdb-7689ee9d526f"
observation = env.reset()
for i in range(30):
observation, _, _, _ = env.step(0)
observation_pp = mdl.lab3.preprocess_pong(observation)
f = plt.figure(figsize=(10, 3))
ax = f.add_subplot(121)
ax2 = f.add_subplot(122)
ax.imshow(observation)
ax.grid(False)
ax2.imshow(np.squeeze(observation_pp))
ax2.grid(False)
plt.title("Preprocessed Observation")
# + id="_mFQ9sbxuxJM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 279} outputId="32d6dc75-d5c8-4679-8c7c-3638b73e13d8"
# training pong
learning_rate = 1e-4
MAX_ITER = 10000
pong_model = create_pong_model()
optimizer = tf.keras.optimizers.Adam(learning_rate)
# plotting
smoothed_reward = mdl.util.LossHistory(smoothing_factor=0.9)
plotter = mdl.util.PeriodicPlotter(sec=5, xlabel='Iterations', ylabel='Rewards')
memory = Memory()
for i_episode in range(MAX_ITER):
plotter.plot(smoothed_reward.get())
observation = env.reset()
previous_frame = mdl.lab3.preprocess_pong(observation)
while True:
current_frame = mdl.lab3.preprocess_pong(observation)
obs_change = current_frame - previous_frame
action = choose_action(pong_model, obs_change)
next_observation, reward, done, info = env.step(action)
memory.add_to_memory(obs_change, action, reward)
if done:
total_reward = sum(memory.rewards)
smoothed_reward.append(total_reward)
train_step(
pong_model,
optimizer,
np.stack(memory.observations, 0),
np.array(memory.actions),
discount_rewards(memory.rewards)
)
memory.clear()
break
observation = next_observation
previous_frame = current_frame
# + id="O38Tbp_3xNH_" colab_type="code" colab={}
saved_pong = mdl.lab3.save_video_of_model(
pong_model, "Pong-v0", obs_diff=True,
pp_fn=mdl.lab3.preprocess_pong)
mdl.lab3.play_video(saved_pong)
| TensorFlow_MIT/Reinforcement Learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Demo1
#
#
# [AMath 585, Winter Quarter 2020](http://staff.washington.edu/rjl/classes/am585w2020/) at the University of Washington. Developed by <NAME> and distributed under the [BSD license](https://github.com/rjleveque/amath585w2020/blob/master/LICENSE). You are free to modify and use as you please, with attribution.
#
# These notebooks are all [available on Github](https://github.com/rjleveque/amath585w2020/).
#
# This notebook just illustrates a few things to make simple plots.
#
# ### Markdown
#
# Note that some cells are executable code cells, others are Markdown cell, which display text and in which you can use latex for mathematics. The notebooks use [Github flavored markdown](https://help.github.com/en/github/writing-on-github/getting-started-with-writing-and-formatting-on-github). Double click on a markdown cell to edit it, then execute it to render.
# ### Make plots appear inline:
#
# The following notebook "magic" command `%matplotlib inline` makes plots show up in the notebook rather than opening a new window.
#
# Instead you can specify `%matplotlib notebook` so they show up in a manner that you can interact with them, e.g. zooming in. If you do this then when you give plot commands in more than one cell you should close earlier figures before plotting new ones (or your plot will be sent to the previous figure), or start each such cell with a `figure` command. If you have too many figures open at once in this mode, you may get an error message telling you to close some. Closing a figure will leave the figure in the notebook, but it's no longer interactive.
# %matplotlib inline
# ### Import a bunch of useful functions
#
# The next line imports many things from [numpy](http://www.numpy.org/) and [matplotlib](http://matplotlib.org/), including things like `pi`, `sin`, `linspace`, and `plot` that are used below.
from pylab import *
x = linspace(0, 4*pi, 100)
y = x*x + x**3*sin(x)
# ### Make a plot
#
# Here is a plot of $f(x) = x^2 + x^3\sin(x)$.
#
figure(figsize=(6,4))
plot(x,y,'r')
# Make another version that has different colored points and line.
#
# Also save the result as a `png` file that can be downloaded.
# +
x = linspace(0, 4*pi, 50)
y = x*x + x**3*sin(x)
figure(figsize=(6,4))
plot(x, y, 'r-')
plot(x, y, 'bo')
title('My plot')
grid(True)
savefig('myplot.png')
# -
# We can also display this png file (or any other image) inline:
from IPython.display import Image
Image('myplot.png')
# ### For help with plotting:
#
# See the [matplotlib gallery](http://matplotlib.org/gallery.html) for many examples.
| notebooks/Demo1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (pytorch)
# language: python
# name: pytorch
# ---
# # Preparing Dataset
# json
# incl
# solvent
# peak = [ppm,intensity]
#
# Yppm batch X 100
# Yintensity batch X 100
InChi = ['1S/CH4/h1H4', '1S/H2O/h1H2', '1S/CH3.Li/h1H3', '1S/H4Si/h1H4', '1S/C3H4/c1-2-3-1/h1-2H,3H2', '1S/C3H5NO/c1-2-3(4)5/h2H,1H2,(H2,4,5)', '1S/C3H4O2/c1-2-3(4)5/h2H,1H2,(H,4,5)', '1S/C2H7NS/c3-1-2-4/h4H,1-3H2']
solvent = ['CCl4','CDCl3', 'benzene-d6', 'cyclohexane', 'neat', 'D2O', 'CDCl3', 'D2O']
frequency = [300, 399.65, 399.65, 300, 300, 89.56, 89.56]
peak = [[[69.60, 0.232, 1000]],
[[631.05, 1.579, 1000]],
[[-526.24, -1.316, 1000],
[-534.29, -1.336, 29],
[-534.90, -1.338, 25]],
[[960.00, 3.200, 1000]],
[[2118.9, 7.063, 995],
[2117.19, 7.057, 997],
[279.85, 0.933, 1000],
[278.21, 0.927, 998]],
[[583.06,6.511,25],
[566.06,6.321,410],
[563,6.287,444],
[558.56,6.237,1000],
[546,6.097,70],
[540.25,6.033,46],
[527.5,5.89,339],
[523.06,5.841,242],
[520.19,5.809,217],
[515.75,5.759,194]],
[[1073.93, 11.992, 489],
[595.92,6.654,287],
[592.50, 6.616, 300],
[580.29, 6.480, 665],
[576.87, 6.442, 813],
[571.06, 6.377, 85],
[563.02, 6.287, 516],
[553.03, 6.175, 1000],
[550.89, 6.152, 130],
[539.18, 6.021, 829],
[537.30, 6.000, 686],
[535.77, 5.983, 846],
[534.23, 5.966, 87],
[529.19, 5.909, 356],
[525.77, 5.871, 136]],
[[445.88, 4.979, 970],
[282.19, 3.151, 30],
[278.19, 3.107, 321],
[276.38, 3.086, 322],
[271.88, 3.036, 474],
[269.75, 3.012, 1000],
[267.38, 2.986, 199],
[264.19, 2.950, 846],
[263.38, 2.941, 860],
[259.63, 2.899, 124],
[259.06, 2.893, 119],
[258.00, 2.881, 103],
[257.06, 2.871, 95],
[256.81, 2.868 , 95],
[256.31, 2.862, 92],
[256.00, 2.859, 88],
[255.31, 2.851, 90],
[255.00, 2.848, 91],
[254.75, 2.845, 91],
[248.00, 2.770, 689],
[241.63, 2.698, 747],
[235.13, 2.626, 268],
[233.25, 2.605, 234]]]
# # Data Preprocessing
import numpy as np
from rdkit import Chem, DataStructs
from rdkit.Chem import AllChem, Draw
from rdkit.Chem.Crippen import MolLogP
from rdkit.Chem.rdMolDescriptors import CalcTPSA
from oddt import toolkits
from oddt.toolkits.extras import rdkit
import matplotlib.pyplot
from rdkit.Chem.Draw import IPythonConsole
IPythonConsole.ipython_useSVG = True
def atom_feature(atom):
return np.array(one_of_k_encoding_unk(atom.GetSymbol(),
['C', 'N', 'O', 'S', 'F', 'H', 'Si', 'P', 'Cl', 'Br',
'Li', 'Na', 'K', 'Mg', 'Ca', 'Fe', 'As', 'Al', 'I', 'B',
'V', 'Tl', 'Sb', 'Sn', 'Ag', 'Pd', 'Co', 'Se', 'Ti', 'Zn',
'Ge', 'Cu', 'Au', 'Ni', 'Cd', 'Mn', 'Cr', 'Pt', 'Hg', 'Pb']) +
one_of_k_encoding(atom.GetDegree(), [0, 1, 2, 3, 4, 5]) +
one_of_k_encoding_unk(atom.GetTotalNumHs(), [0, 1, 2, 3, 4]) +
one_of_k_encoding_unk(atom.GetImplicitValence(), [0, 1, 2, 3, 4, 5]) +
[atom.GetIsAromatic()]) # (40, 6, 5, 6, 1)
# +
def one_of_k_encoding(x, allowable_set):
if x not in allowable_set:
raise Exception("input {0} not in allowable set{1}:".format(x, allowable_set))
#print list((map(lambda s: x == s, allowable_set)))
return list(map(lambda s: x == s, allowable_set))
def one_of_k_encoding_unk(x, allowable_set):
"""Maps inputs not in the allowable set to the last element."""
if x not in allowable_set:
x = allowable_set[-1]
return list(map(lambda s: x == s, allowable_set))
# +
def mol_with_atom_index(mol):
n_atoms = mol.GetNumAtoms()
for idx in range(n_atoms):
mol.GetAtomWithIdx(idx).SetProp('molAtomMapNumber', str(mol.GetAtomWithIdx(idx).GetIdx()))
return mol
def assign_peak_to_H(mol, allowable_set):
partial_charge = []
n_atoms = mol.GetNumAtoms()
for i, atom in enumerate(mol.GetAtoms()):
if atom.GetSymbol() == 'H':
pc = [int(i), float(atom.GetProp("_GasteigerCharge"))]
partial_charge.append(pc)
else:
partial_charge.append([int(i), 0])
partial_charge.sort(key = lambda x: x[1])
partial_charge.reverse()
#for i in range(len(partial_charge)):
# print ("{0}".format(partial_charge[i]))
'''now list partial_charge is oriented with the largest value to lowest
value of the partial charge assigned with its atom number'''
'''Allowable set is NMR peak data which is sorted from highest ppm to
lowest ppm, we have to assign each ppm to **several** numbers of protons'''
nmr_peak = dict()
allowable_set_pad = np.zeros(n_atoms)
allowable_set_pad[0:len(allowable_set)] = allowable_set
#print(allowable_set_pad)
nmr_peak['{0}'.format(partial_charge[0][0])] = allowable_set_pad[0]
#partial_charge[i][1] 값을 비교 -> 전꺼와 같으면 같은 값, 다르면 그 다음 값
j = 0
for i in range(1,len(partial_charge)):
if partial_charge[i-1][1] == partial_charge[i][1]:
nmr_peak['{0}'.format(partial_charge[i][0])] = allowable_set_pad[j]
elif partial_charge[i-1][1] > partial_charge[i][1]:
nmr_peak['{0}'.format(partial_charge[i][0])] = allowable_set_pad[j+1]
j += 1
return nmr_peak
# -
def convert_to_graph(InChi_list, peak_list):
adj = []
adj_norm = []
features = []
maxNumAtoms = 50
for i in InChi_list:
#Mol
#iMol = Chem.MolFromSmiles(i.strip())
iMol = Chem.inchi.MolFromInchi(i.strip())
iMol = Chem.rdmolops.AddHs(iMol)
#Adj
iAdjTmp = Chem.rdmolops.GetAdjacencyMatrix(iMol)
iMol = mol_with_atom_index(iMol)
AllChem.ComputeGasteigerCharges(iMol)
# Feature
if(iAdjTmp.shape[0] <= maxNumAtoms):
# Feature-preprocessing
iFeature = np.zeros((maxNumAtoms, 58))
iFeatureTmp = []
nmr_dict = assign_peak_to_H(iMol, peak_list)
nmr_peak = []
for atom in iMol.GetAtoms():
#iFeatureTmp.append(atom_feature(atom)) ### atom features only
arr = atom_feature(atom)
#print(arr)
#print(arr[0], arr[1], arr[2], arr[3], arr[4], arr[5])
nmr_peak.append(np.array(nmr_dict[str(atom.GetAtomMapNum())]))
iFeatureTmp.append(arr)
#print(arr)
iFeature[0:len(iFeatureTmp), 0:58] = iFeatureTmp ### 0 padding for feature-set
nmrpeak = np.zeros(maxNumAtoms)
nmrpeak[0:len(nmr_peak)] = nmr_peak
for i in range(len(iFeature)):
iFeature[i][5] = nmrpeak[i]
features.append(iFeature)
# Adj-preprocessing
iAdj = np.zeros((maxNumAtoms, maxNumAtoms))
iAdj[0:len(iFeatureTmp), 0:len(iFeatureTmp)] = iAdjTmp + np.eye(len(iFeatureTmp))
adj.append(np.asarray(iAdj))
features = np.asarray(features)
return features, adj
features, adj = convert_to_graph(['InChI=1S/C5H10O/c1-3-4-5(2)6/h3-6H,1-2H3/b4-3+'],[5.64, 5.53, 4.23, 1.92, 1.681, 1.240])
for i in range(50):
print(features[0][i][5])
inchi = 'InChI=1S/C5H10O/c1-3-4-5(2)6/h3-6H,1-2H3/b4-3+'
[['A', 5,64],
['B', 5.53],
['C', 4.23],
['D', 1.92],
['E', 1.681],
['F', 1.240]]
sets = [5.64, 5.53, 4.23, 1.92, 1.681, 1.240]
m = Chem.inchi.MolFromInchi(inchi)
m = Chem.rdmolops.AddHs(m)
m = mol_with_atom_index(m)
Chem.Draw.MolToMPL(m)
AllChem.ComputeGasteigerCharges(m)
nmr = assign_peak_to_H(m, sets)
print(nmr)
# +
def mol_with_atom_index(mol):
n_atoms = mol.GetNumAtoms()
for idx in range(n_atoms):
mol.GetAtomWithIdx(idx).SetProp('molAtomMapNumber', str(mol.GetAtomWithIdx(idx).GetIdx()))
return mol
def assign_peak_to_H(mol, allowable_set):
partial_charge = []
n_atoms = mol.GetNumAtoms()
for i, atom in enumerate(mol.GetAtoms()):
pc = [int(i), float(atom.GetProp("_GasteigerCharge"))]
partial_charge.append(pc)
partial_charge.sort(key = lambda x: x[1])
partial_charge.reverse()
#for i in range(len(partial_charge)):
# print ("{0}".format(partial_charge[i]))
'''now list partial_charge is oriented with the largest value to lowest
value of the partial charge assigned with its atom number'''
'''Allowable set is NMR peak data which is sorted from highest ppm to
lowest ppm, we have to assign each ppm to **several** numbers of protons'''
nmr_peak = []
allowable_set_pad = np.zeros(n_atoms)
allowable_set_pad[0:len(allowable_set)] = allowable_set
#print(allowable_set_pad)
nmr_peak.append([partial_charge[0][0],allowable_set_pad[0]])
#partial_charge[i][1] 값을 비교 -> 전꺼와 같으면 같은 값, 다르면 그 다음 값
j = 0
for i in range(1,len(partial_charge)):
if partial_charge[i-1][1] == partial_charge[i][1]:
nmr_peak.append([partial_charge[i][0], allowable_set_pad[j]])
elif partial_charge[i-1][1] > partial_charge[i][1]:
nmr_peak.append([partial_charge[i][0], allowable_set_pad[j+1]])
j += 1
return nmr_peak
# -
inchi = 'InChI=1S/C5H10O/c1-3-4-5(2)6/h3-6H,1-2H3/b4-3+'
[['A', 5,64],
['B', 5.53],
['C', 4.23],
['D', 1.92],
['E', 1.681],
['F', 1.240]]
sets = [5.64, 5.53, 4.23, 1.92, 1.681, 1.240]
m = Chem.inchi.MolFromInchi(inchi)
m = Chem.rdmolops.AddHs(m)
m = mol_with_atom_index(m)
Chem.Draw.MolToMPL(m)
AllChem.ComputeGasteigerCharges(m)
nmr = assign_peak_to_H(m, sets)
print(nmr)
| Term Project/Partial_Charge.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import psycopg2
import sqlalchemy
# %load_ext sql
# Format %sql dialect+driver://username:password@host:port/database
# Example format
# %sql postgresql://postgres:password@localhost/postgres
# + language="sql"
#
# Create table If Not Exists Employees (id int, salary int);
# Truncate table Employees;
# insert into Employees (id, salary) values ('1', '100');
# insert into Employees (id, salary) values ('2', '200');
# insert into Employees (id, salary) values ('3', '300');
#
#
# + language="sql"
#
# select * from Employees
# + language="sql"
#
# CREATE FUNCTION getNthHighestSalary(N INT) RETURNS INT
# BEGIN
# RETURN (
# select salary as getnth from Employee where id = N
# );
# END
# + language="sql"
#
# select case (when 0 > (select count(*) from Employees) then null
# else salary end) as nthsal from
# (select *,dense_rank() over (order by salary) as salrank from Employees)
# Emp2 where salrank = 2
# + language="sql"
# CREATE FUNCTION getNthHighestSalary(N INT) RETURNS INT
# BEGIN
# RETURN (
# select *,dense_rank() over (order by salary) as salrank from Employees
# );
# END
# -
| SQL/SQL_GetN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Robust Scaler - Experimento
#
# Este é um componante que dimensiona atributos usando estatísticas robustas para outliers. Este Scaler remove a mediana e dimensiona os dados de acordo com o intervalo quantil (o padrão é Amplitude interquartil). Amplitude interquartil é o intervalo entre o 1º quartil (25º quantil) e o 3º quartil (75º quantil). Faz uso da implementação do [Scikit-learn](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.RobustScaler.html). <br>
# Scikit-learn é uma biblioteca open source de machine learning que suporta apredizado supervisionado e não supervisionado. Também provê várias ferramentas para montagem de modelo, pré-processamento de dados, seleção e avaliação de modelos, e muitos outros utilitários.
# ## Declaração de parâmetros e hiperparâmetros
#
# Declare parâmetros com o botão <img src="data:image/png;base64,iVBORw0KGgoAAAANSUhEUgAAABQAAAAUCAYAAACNiR0NAAABhWlDQ1BJQ0MgcHJvZmlsZQAAKJF9kT1Iw0AcxV9TtaIVBzuIOASpThb8QhylikWwUNoKrTqYXPohNGlIUlwcBdeCgx+LVQcXZ10dXAVB8APEydFJ0UVK/F9SaBHjwXE/3t173L0DhFqJqWbbGKBqlpGMRcVMdkUMvKID3QhiCOMSM/V4aiENz/F1Dx9f7yI8y/vcn6NHyZkM8InEs0w3LOJ14ulNS+e8TxxiRUkhPiceNeiCxI9cl11+41xwWOCZISOdnCMOEYuFFpZbmBUNlXiKOKyoGuULGZcVzluc1VKFNe7JXxjMacsprtMcRAyLiCMBETIq2EAJFiK0aqSYSNJ+1MM/4PgT5JLJtQFGjnmUoUJy/OB/8LtbMz854SYFo0D7i21/DAOBXaBete3vY9uunwD+Z+BKa/rLNWDmk/RqUwsfAb3bwMV1U5P3gMsdoP9JlwzJkfw0hXweeD+jb8oCfbdA16rbW2Mfpw9AmrpaugEODoGRAmWveby7s7W3f880+vsBocZyukMJsmwAAAAGYktHRAD/AP8A/6C9p5MAAAAJcEhZcwAADdcAAA3XAUIom3gAAAAHdElNRQfkBgsMIwnXL7c0AAACDUlEQVQ4y92UP4gTQRTGf29zJxhJZ2NxbMBKziYWlmJ/ile44Nlkd+dIYWFzItiNgoIEtFaTzF5Ac/inE/urtLWxsMqmUOwCEpt1Zmw2xxKi53XitPO9H9978+aDf/3IUQvSNG0450Yi0jXG7C/eB0cFeu9viciGiDyNoqh2KFBrHSilWstgnU7nFLBTgl+ur6/7PwK11kGe5z3n3Hul1MaiuCgKDZwALHA7z/Oe1jpYCtRaB+PxuA8kQM1aW68Kt7e3zwBp6a5b1ibj8bhfhQYVZwMRiQHrvW9nWfaqCrTWPgRWvPdvsiy7IyLXgEJE4slk8nw+T5nDgDbwE9gyxryuwpRSF5xz+0BhrT07HA4/AyRJchUYASvAbhiGaRVWLIMBYq3tAojIszkMoNRulbXtPM8HwV/sXSQi54HvQRDcO0wfhGGYArvAKjAq2wAgiqJj3vsHpbtur9f7Vi2utLx60LLW2hljEuBJOYu9OI6vAzQajRvAaeBLURSPlsBelA+VhWGYaq3dwaZvbm6+m06noYicE5ErrVbrK3AXqHvvd4bD4Ye5No7jSERGwKr3Pms2m0pr7Rb30DWbTQWYcnFvAieBT7PZbFB1V6vVfpQaU4UtDQetdTCZTC557/eA48BlY8zbRZ1SqrW2tvaxCvtt2iRJ0i9/xb4x5uJRwmNlaaaJ3AfqIvKY/+78Av++6uiSZhYMAAAAAElFTkSuQmCC" /> na barra de ferramentas.<br>
# A variável `dataset` possui o caminho para leitura do arquivos importados na tarefa de "Upload de dados".<br>
# Você também pode importar arquivos com o botão <img src="data:image/png;base64,iVBORw0KGgoAAAANSUhEUgAAABQAAAAUCAYAAACNiR0NAAABhWlDQ1BJQ0MgcHJvZmlsZQAAKJF9kT1Iw0AcxV9TtaIVBzuIOASpThb8QhylikWwUNoKrTqYXPohNGlIUlwcBdeCgx+LVQcXZ10dXAVB8APEydFJ0UVK/F9SaBHjwXE/3t173L0DhFqJqWbbGKBqlpGMRcVMdkUMvKID3QhiCOMSM/V4aiENz/F1Dx9f7yI8y/vcn6NHyZkM8InEs0w3LOJ14ulNS+e8TxxiRUkhPiceNeiCxI9cl11+41xwWOCZISOdnCMOEYuFFpZbmBUNlXiKOKyoGuULGZcVzluc1VKFNe7JXxjMacsprtMcRAyLiCMBETIq2EAJFiK0aqSYSNJ+1MM/4PgT5JLJtQFGjnmUoUJy/OB/8LtbMz854SYFo0D7i21/DAOBXaBete3vY9uunwD+Z+BKa/rLNWDmk/RqUwsfAb3bwMV1U5P3gMsdoP9JlwzJkfw0hXweeD+jb8oCfbdA16rbW2Mfpw9AmrpaugEODoGRAmWveby7s7W3f880+vsBocZyukMJsmwAAAAGYktHRAD/AP8A/6C9p5MAAAAJcEhZcwAADdcAAA3XAUIom3gAAAAHdElNRQfkBgsOBy6ASTeXAAAC/0lEQVQ4y5WUT2gcdRTHP29m99B23Uiq6dZisgoWCxVJW0oL9dqLfyhCvGWY2YUBI95MsXgwFISirQcLhS5hfgk5CF3wJIhFI7aHNsL2VFZFik1jS1qkiZKdTTKZ3/MyDWuz0fQLc/m99/vMvDfv+4RMlUrlkKqeAAaBAWAP8DSgwJ/AXRG5rao/WWsvTU5O3qKLBMD3fSMiPluXFZEPoyj67PGAMzw83PeEMABHVT/oGpiamnoAmCcEWhH5tFsgF4bh9oWFhfeKxeJ5a+0JVT0oImWgBPQCKfAQuAvcBq67rltX1b+6ApMkKRcKhe9V9QLwbavV+qRer692Sx4ZGSnEcXw0TdP3gSrQswGYz+d/S5IkVtXTwOlCoZAGQXAfmAdagAvsAErtdnuXiDy6+023l7qNRsMODg5+CawBzwB9wFPA7mx8ns/KL2Tl3xCRz5eWlkabzebahrHxPG+v4zgnc7ncufHx8Z+Hhoa29fT0lNM03Q30ikiqqg+ttX/EcTy3WTvWgdVqtddaOw/kgXvADHBHROZVNRaRvKruUNU+EdkPfGWM+WJTYOaSt1T1LPDS/4zLWWPMaLVaPWytrYvIaBRFl/4F9H2/JCKvGmMu+76/X0QOqGoZKDmOs1NV28AicMsYc97zvFdc1/0hG6kEeNsY83UnsCwivwM3VfU7YEZE7lhr74tIK8tbnJiYWPY8b6/ruleAXR0ftQy8boyZXi85CIIICDYpc2ZgYODY3NzcHmvt1eyvP64lETkeRdE1yZyixWLx5U2c8q4x5mIQBE1g33/0d3FlZeXFR06ZttZesNZejuO4q1NE5CPgWVV9E3ij47wB1IDlJEn+ljAM86urq7+KyAtZTgqsO0VV247jnOnv7/9xbGzMViqVMVX9uANYj6LonfVtU6vVkjRNj6jqGeCXzGrPAQeA10TkuKpOz87ONrayhnIA2Qo7BZwKw3B7kiRloKSqO13Xja21C47jPNgysFO1Wi0GmtmzQap6DWgD24A1Vb3SGf8Hfstmz1CuXEIAAAAASUVORK5CYII=" /> na barra de ferramentas.
# + tags=["parameters"]
# parâmetros
dataset = "/tmp/data/iris.csv" #@param {type:"string"}
target = None #@param {type:"feature", label:"Atributo alvo", description: "Esse valor será utilizado para garantir que o alvo não seja removido."}
with_centering = True #@param {type:"boolean", label:"Centralização", description:"Centralizar os dados antes de dimensionar. Ocorre exceção quando usado com matrizes esparsas"}
with_scaling = True #@param {type:"boolean", label:"Dimensionamento", description:"Dimensionar os dados para um intervalo interquartil"}
# -
# ## Acesso ao conjunto de dados
#
# O conjunto de dados utilizado nesta etapa será o mesmo carregado através da plataforma.<br>
# O tipo da variável retornada depende do arquivo de origem:
# - [pandas.DataFrame](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.html) para CSV e compressed CSV: .csv .csv.zip .csv.gz .csv.bz2 .csv.xz
# - [Binary IO stream](https://docs.python.org/3/library/io.html#binary-i-o) para outros tipos de arquivo: .jpg .wav .zip .h5 .parquet etc
# +
import pandas as pd
df = pd.read_csv(dataset)
# -
has_target = True if target is not None and target in df.columns else False
# +
X = df.copy()
if has_target:
X = df.drop(target, axis=1)
y = df[target]
# -
# ## Acesso aos metadados do conjunto de dados
#
# Utiliza a função `stat_dataset` do [SDK da PlatIAgro](https://platiagro.github.io/sdk/) para carregar metadados. <br>
# Por exemplo, arquivos CSV possuem `metadata['featuretypes']` para cada coluna no conjunto de dados (ex: categorical, numerical, or datetime).
# +
import numpy as np
from platiagro import stat_dataset
metadata = stat_dataset(name=dataset)
featuretypes = metadata["featuretypes"]
columns = df.columns.to_numpy()
featuretypes = np.array(featuretypes)
if has_target:
target_index = np.argwhere(columns == target)
columns = np.delete(columns, target_index)
featuretypes = np.delete(featuretypes, target_index)
# -
# ## Configuração dos atributos
# +
from platiagro.featuretypes import NUMERICAL
# Selects the indexes of numerical
numerical_indexes = np.where(featuretypes == NUMERICAL)[0]
non_numerical_indexes = np.where(~(featuretypes == NUMERICAL))[0]
# After the step of the make_column_transformer,
# numerical features are grouped in the beggining of the array
numerical_indexes_after_first_step = np.arange(len(numerical_indexes))
# -
# ## Treina um modelo usando sklearn.preprocessing.RobustScaler
# +
from sklearn.compose import make_column_transformer
from sklearn.impute import SimpleImputer
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import RobustScaler
pipeline = Pipeline(
steps=[
(
"imputer",
make_column_transformer(
(SimpleImputer(), numerical_indexes), remainder="passthrough"
),
),
(
"robust_scaler",
make_column_transformer(
(
RobustScaler(
with_centering=with_centering, with_scaling=with_scaling
),
numerical_indexes_after_first_step,
),
remainder="passthrough",
),
),
]
)
# Train model and transform dataset
X = pipeline.fit_transform(X)
# Put numerical features in the lowest indexes
features_after_pipeline = np.concatenate(
(columns[numerical_indexes], columns[non_numerical_indexes])
)
# +
# Put data back in a pandas.DataFrame
df = pd.DataFrame(data=X, columns=features_after_pipeline)
if has_target:
df[target] = y
# -
# ## Cria visualização do resultado
#
# Cria visualização do resultado como uma planilha.
# +
import matplotlib.pyplot as plt
from platiagro.plotting import plot_data_table
ax = plot_data_table(df)
plt.show()
# -
# ## Salva alterações no conjunto de dados
#
# O conjunto de dados será salvo (e sobrescrito com as respectivas mudanças) localmente, no container da experimentação, utilizando a função `pandas.DataFrame.to_csv`.<br>
# save dataset changes
df.to_csv(dataset, index=False)
# ## Salva resultados da tarefa
#
# A plataforma guarda o conteúdo de `/tmp/data/` para as tarefas subsequentes.
# +
from joblib import dump
artifacts = {
"pipeline": pipeline,
"columns": columns,
"features_after_pipeline": features_after_pipeline,
}
dump(artifacts, "/tmp/data/robust-scaler.joblib")
| tasks/robust-scaler/Experiment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Import
# +
import sys
from pathlib import Path
import matplotlib.pyplot as plt
import tensorflow as tf
# -
sys.path.append('../../')
from test_fast_inference.data import *
from test_fast_inference.tensorflow_tensorrt_conversion import *
# # Train
# +
# Config
batch_size = 28
num_batches = 20
num_epochs = 280000//(batch_size*num_batches)
lr = 1e-3
# Loss
loss = tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True)
# Metrics
metrics = [
tf.keras.metrics.SparseCategoricalCrossentropy(),
tf.keras.metrics.SparseCategoricalAccuracy()
]
# Optimizer
optimizer = tf.keras.optimizers.Adam(learning_rate=lr)
# Model
in_channels = 1
out_channels = 5
layout_encoder = [[[16,5,2]],
[[16,3,1],[16,3,2]],
[[16,3,1],[16,3,2]],
[[16,3,1],[16,3,2]]]
model = UNET_TRT(get_encoder(in_channels, layout_encoder), out_channels)
# -
# Training dataset is large so won't be cacheable in memory
ds_trn = (
tf.data.TFRecordDataset(['/data/fiducial_detect/trn/tfrecord/trn.tfrecord'])
.map(preprocess_tfrecord)
.batch(batch_size)
.prefetch(buffer_size=tf.data.experimental.AUTOTUNE)
)
# Validation dataset is small so cache it in memory
ds_val = (
tf.data.Dataset.list_files('/data/fiducial_detect/val/*_mask.png', shuffle=False)
.map(preprocess_file)
.batch(batch_size)
.cache()
.prefetch(buffer_size=tf.data.experimental.AUTOTUNE)
)
# Train
callback_checkpoint = tf.keras.callbacks.ModelCheckpoint(
'checkpoints/best.tf',
monitor='val_sparse_categorical_accuracy',
save_best_only=True,
mode='auto',
save_freq='epoch'
)
model.compile(optimizer, loss, metrics)
model_history = model.fit(ds_trn,
epochs=num_epochs,
steps_per_epoch=num_batches,
validation_data=ds_val,
callbacks=callback_checkpoint)
# # Test
model.load_weights('checkpoints/best.tf')
for X,y in ds_val.take(1):
pass
y_hat = model.predict(X)
idx = 0
plt.figure(figsize=(15,10))
plt.imshow(X[idx], cmap='gray', vmin=-1, vmax=1)
plt.figure(figsize=(15,10))
plt.imshow(tf.argmax(y_hat, axis=3)[idx])
plt.figure(figsize=(15,10))
plt.imshow(y[idx])
plt.figure(figsize=(15,10))
plt.imshow(tf.math.not_equal(tf.argmax(y_hat, axis=3)[idx], tf.squeeze(y[idx])))
| training/attempt_4/train.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Back to the main [Index](../index.ipynb)
# ## Simple Example
#
# This shows how to set up a simple model in Tellurium and solve it as an ODE. Tellurium uses a human-readable representation of SBML models called Antimony. The Antimony code for this example contains a single reaction with associated kinetics. After creating the Antimony string, use the `loada` function to load it into the simulator.
import tellurium as te
r = te.loada('S1 -> S2; k1*S1; k1 = 0.1; S1 = 10')
r.simulate(0, 50, 100)
r.plot();
# ## More Complex Example
#
# Tellurium can also handle stochastic models. This example shows how to select Tellurium's stochastic solver. The underlying simulation engine used by Tellurium implements a Gibson direct method for simulating this model.
# +
import tellurium as te
import numpy as np
r = te.loada('''
J1: S1 -> S2; k1*S1;
J2: S2 -> S3; k2*S2 - k3*S3
# J2_1: S2 -> S3; k2*S2
# J2_2: S3 -> S2; k3*S3;
J3: S3 -> S4; k4*S3;
k1 = 0.1; k2 = 0.5; k3 = 0.5; k4 = 0.5;
S1 = 100;
''')
# use a stochastic solver
r.integrator = 'gillespie'
r.integrator.seed = 1234
# selections specifies the output variables in a simulation
selections = ['time'] + r.getBoundarySpeciesIds() + r.getFloatingSpeciesIds()
r.integrator.variable_step_size = False
# run repeated simulation
Ncol = len(r.selections)
Nsim = 30
points = 101
s_sum = np.zeros(shape=[points, Ncol])
#s_sum = r.simulate(0, 50, points, selections=selections)
for k in range(Nsim):
r.resetToOrigin()
s = r.simulate(0, 50, points, selections=selections)
s_sum += s
# no legend, do not show
r.plot(s, alpha=0.5, show=False)
#r.show()
# add mean curve, legend, show everything and set labels, titels, ...
te.plot(s[:,0], s_sum[:,1:]/Nsim, colnames=selections, title="Stochastic simulation", xtitle="time", ytitle="concentration")
| examples/notebooks/core/introduction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#default_exp core
# -
#export
from local.test import *
from local.imports import *
from local.notebook.showdoc import *
# # Core
#
# > Basic functions used in the fastai library
# export
defaults = SimpleNamespace()
# ## Metaclasses
# See this [blog post](https://realpython.com/python-metaclasses/) for more information about metaclasses.
# - `PrePostInitMeta` ensures that the classes defined with it run `__pre_init__` and `__post_init__` (without having to write `self.__pre_init__()` and `self.__post_init__()` in the actual `init`
# - `NewChkMeta` gives the `PrePostInitMeta` functionality and ensures classes defined with it don't re-create an object of their type whenever it's passed to the constructor
# - `BypassNewMeta` ensures classes defined with it can easily be casted form objects they subclass.
#export
class FixSigMeta(type):
"A metaclass that fixes the signature on classes that override __new__"
def __new__(cls, name, bases, dict):
res = super().__new__(cls, name, bases, dict)
if res.__init__ is not object.__init__: res.__signature__ = inspect.signature(res.__init__)
return res
#export
class PrePostInitMeta(FixSigMeta):
"A metaclass that calls optional `__pre_init__` and `__post_init__` methods"
def __call__(cls, *args, **kwargs):
res = cls.__new__(cls)
if type(res)==cls:
if hasattr(res,'__pre_init__'): res.__pre_init__(*args,**kwargs)
res.__init__(*args,**kwargs)
if hasattr(res,'__post_init__'): res.__post_init__(*args,**kwargs)
return res
show_doc(PrePostInitMeta, title_level=3)
# +
class _T(metaclass=PrePostInitMeta):
def __pre_init__(self): self.a = 0; assert self.a==0
def __init__(self,b=0): self.a += 1; assert self.a==1
def __post_init__(self): self.a += 1; assert self.a==2
t = _T()
test_eq(t.a, 2)
# -
#export
class NewChkMeta(FixSigMeta):
"Metaclass to avoid recreating object passed to constructor"
def __call__(cls, x=None, *args, **kwargs):
if not args and not kwargs and x is not None and isinstance(x,cls):
x._newchk = 1
return x
res = super().__call__(*((x,) + args), **kwargs)
res._newchk = 0
return res
class _T(metaclass=NewChkMeta):
"Testing"
def __init__(self, o=None, b=1):
self.foo = getattr(o,'foo',0) + 1
self.b = b
# +
class _T2():
def __init__(self, o): self.foo = getattr(o,'foo',0) + 1
t = _T(1)
test_eq(t.foo,1)
t2 = _T(t)
test_eq(t2.foo,1)
test_is(t,t2)
t3 = _T(t, b=2)
test_eq(t3.b, 2)
assert not t3 is t
t = _T2(1)
test_eq(t.foo,1)
t2 = _T2(t)
test_eq(t2.foo,2)
test_eq(_T.__doc__, "Testing")
# TODO: this shouldn't have "self, "
test_eq(str(inspect.signature(_T)), '(self, o=None, b=1)')
# -
#export
class BypassNewMeta(FixSigMeta):
"Metaclass: casts `x` to this class if it's of type `cls._bypass_type`, initializing with `_new_meta` if available"
def __call__(cls, x=None, *args, **kwargs):
if hasattr(cls, '_new_meta'): x = cls._new_meta(x, *args, **kwargs)
elif not isinstance(x,getattr(cls,'_bypass_type',object)) or len(args) or len(kwargs):
x = super().__call__(*((x,)+args), **kwargs)
if cls!=x.__class__: x.__class__ = cls
return x
# +
class T0: pass
class _T(T0, metaclass=BypassNewMeta):
_bypass_type=T0
def __init__(self,x): self.x=x
t = T0()
t.a = 1
t2 = _T(t)
test_eq(type(t2), _T)
test_eq(t2.a,1)
test_is(t2,t)
t = _T(2)
t.x = 2
# -
# ## Foundational functions
#export
def copy_func(f):
"Copy a non-builtin function (NB `copy.copy` does not work for this)"
if not isinstance(f,types.FunctionType): return copy(f)
fn = types.FunctionType(f.__code__, f.__globals__, f.__name__, f.__defaults__, f.__closure__)
fn.__dict__.update(f.__dict__)
return fn
#export
def patch_to(cls, as_prop=False):
"Decorator: add `f` to `cls`"
if not isinstance(cls, (tuple,list)): cls=(cls,)
def _inner(f):
for c_ in cls:
nf = copy_func(f)
# `functools.update_wrapper` when passing patched function to `Pipeline`, so we do it manually
for o in functools.WRAPPER_ASSIGNMENTS: setattr(nf, o, getattr(f,o))
nf.__qualname__ = f"{c_.__name__}.{f.__name__}"
setattr(c_, f.__name__, property(nf) if as_prop else nf)
return f
return _inner
# +
class _T3(int): pass
@patch_to(_T3)
def func1(x, a): return x+a
t = _T3(1)
test_eq(t.func1(2), 3)
# -
# If `cls` is a tuple, `f` is added to all types in the tuple.
# +
class _T4(int): pass
@patch_to((_T3,_T4))
def func2(x, a): return x+2*a
t = _T3(1)
test_eq(t.func2(1), 3)
t = _T4(1)
test_eq(t.func2(1), 3)
# -
#export
def patch(f):
"Decorator: add `f` to the first parameter's class (based on f's type annotations)"
cls = next(iter(f.__annotations__.values()))
return patch_to(cls)(f)
# +
@patch
def func(x:_T3, a):
"test"
return x+2
t = _T3(1)
test_eq(t.func(2), 3)
test_eq(t.func.__qualname__, '_T3.func')
# -
# If annotation is a tuple, the function is added to all types in the tuple.
# +
@patch
def func3(x:(_T3,_T4), a):
"test"
return x+2*a
t = _T3(1)
test_eq(t.func3(2), 5)
test_eq(t.func3.__qualname__, '_T3.func3')
t = _T4(1)
test_eq(t.func3(2), 5)
test_eq(t.func3.__qualname__, '_T4.func3')
# -
#export
def patch_property(f):
"Decorator: add `f` as a property to the first parameter's class (based on f's type annotations)"
cls = next(iter(f.__annotations__.values()))
return patch_to(cls, as_prop=True)(f)
# +
@patch_property
def prop(x:_T3): return x+1
t = _T3(1)
test_eq(t.prop, 2)
# -
#export
def _mk_param(n,d=None): return inspect.Parameter(n, inspect.Parameter.KEYWORD_ONLY, default=d)
def test_sig(f, b): test_eq(str(inspect.signature(f)), b)
#export
def use_kwargs(names, keep=False):
"Decorator: replace `**kwargs` in signature with `names` params"
def _f(f):
sig = inspect.signature(f)
sigd = dict(sig.parameters)
k = sigd.pop('kwargs')
s2 = {n:_mk_param(n) for n in names if n not in sigd}
sigd.update(s2)
if keep: sigd['kwargs'] = k
f.__signature__ = sig.replace(parameters=sigd.values())
return f
return _f
# +
@use_kwargs(['y', 'z'])
def foo(a, b=1, **kwargs): pass
test_sig(foo, '(a, b=1, *, y=None, z=None)')
@use_kwargs(['y', 'z'], keep=True)
def foo(a, *args, b=1, **kwargs): pass
test_sig(foo, '(a, *args, b=1, y=None, z=None, **kwargs)')
# -
#export
def delegates(to=None, keep=False):
"Decorator: replace `**kwargs` in signature with params from `to`"
def _f(f):
if to is None: to_f,from_f = f.__base__.__init__,f.__init__
else: to_f,from_f = to,f
from_f = getattr(from_f,'__func__',from_f)
if hasattr(from_f,'__delwrap__'): return f
sig = inspect.signature(from_f)
sigd = dict(sig.parameters)
k = sigd.pop('kwargs')
s2 = {k:v for k,v in inspect.signature(to_f).parameters.items()
if v.default != inspect.Parameter.empty and k not in sigd}
sigd.update(s2)
if keep: sigd['kwargs'] = k
from_f.__signature__ = sig.replace(parameters=sigd.values())
from_f.__delwrap__ = to_f
return f
return _f
# +
def basefoo(e, c=2): pass
@delegates(basefoo)
def foo(a, b=1, **kwargs): pass
test_sig(foo, '(a, b=1, c=2)')
@delegates(basefoo, keep=True)
def foo(a, b=1, **kwargs): pass
test_sig(foo, '(a, b=1, c=2, **kwargs)')
# +
class BaseFoo:
def __init__(self, e, c=2): pass
@delegates()
class Foo(BaseFoo):
def __init__(self, a, b=1, **kwargs): super().__init__(**kwargs)
test_sig(Foo, '(a, b=1, c=2)')
# -
#export
def funcs_kwargs(cls):
"Replace methods in `self._methods` with those from `kwargs`"
old_init = cls.__init__
def _init(self, *args, **kwargs):
for k in cls._methods:
arg = kwargs.pop(k,None)
if arg is not None:
if isinstance(arg,types.MethodType): arg = types.MethodType(arg.__func__, self)
setattr(self, k, arg)
old_init(self, *args, **kwargs)
functools.update_wrapper(_init, old_init)
cls.__init__ = use_kwargs(cls._methods)(_init)
return cls
#export
def method(f):
"Mark `f` as a method"
# `1` is a dummy instance since Py3 doesn't allow `None` any more
return types.MethodType(f, 1)
# +
@funcs_kwargs
class T:
_methods=['b']
def __init__(self, f=1, **kwargs): assert not kwargs
def a(self): return 1
def b(self): return 2
t = T()
test_eq(t.a(), 1)
test_eq(t.b(), 2)
t = T(b = lambda:3)
test_eq(t.b(), 3)
test_sig(T, '(f=1, *, b=None)')
test_fail(lambda: T(a = lambda:3))
@method
def _f(self,a=1): return a+1
t = T(b = _f)
test_eq(t.b(2), 3)
class T2(T):
def __init__(self,a):
super().__init__(b = lambda:3)
self.a=a
t = T2(a=1)
test_eq(t.b(), 3)
test_sig(T2, '(a)')
def _g(a=1): return a+1
class T3(T): b = staticmethod(_g)
t = T3()
test_eq(t.b(2), 3)
# -
# Runtime type checking is handy, so let's make it easy!
@contextmanager
def working_directory(path):
"Change working directory to `path` and return to previous on exit."
prev_cwd = Path.cwd()
os.chdir(path)
try: yield
finally: os.chdir(prev_cwd)
# +
#def is_listy(x): return isinstance(x,(list,tuple,Generator))
# -
#export core
def add_docs(cls, cls_doc=None, **docs):
"Copy values from `docs` to `cls` docstrings, and confirm all public methods are documented"
if cls_doc is not None: cls.__doc__ = cls_doc
for k,v in docs.items():
f = getattr(cls,k)
if hasattr(f,'__func__'): f = f.__func__ # required for class methods
f.__doc__ = v
# List of public callables without docstring
nodoc = [c for n,c in vars(cls).items() if callable(c)
and not n.startswith('_') and c.__doc__ is None]
assert not nodoc, f"Missing docs: {nodoc}"
assert cls.__doc__ is not None, f"Missing class docs: {cls}"
#export core
def docs(cls):
"Decorator version of `add_docs`, using `_docs` dict"
add_docs(cls, **cls._docs)
return cls
# +
class _T:
def f(self): pass
@classmethod
def g(cls): pass
add_docs(_T, "a", f="f", g="g")
test_eq(_T.__doc__, "a")
test_eq(_T.f.__doc__, "f")
test_eq(_T.g.__doc__, "g")
# -
#export
def custom_dir(c, add:list):
"Implement custom `__dir__`, adding `add` to `cls`"
return dir(type(c)) + list(c.__dict__.keys()) + add
show_doc(is_iter)
assert is_iter([1])
assert not is_iter(array(1))
assert is_iter(array([1,2]))
assert (o for o in range(3))
#export
class _Arg:
def __init__(self,i): self.i = i
arg0 = _Arg(0)
arg1 = _Arg(1)
arg2 = _Arg(2)
arg3 = _Arg(3)
arg4 = _Arg(4)
#export
class bind:
"Same as `partial`, except you can use `arg0` `arg1` etc param placeholders"
def __init__(self, fn, *pargs, **pkwargs):
self.fn,self.pargs,self.pkwargs = fn,pargs,pkwargs
self.maxi = max((x.i for x in pargs if isinstance(x, _Arg)), default=-1)
def __call__(self, *args, **kwargs):
args = list(args)
kwargs = {**self.pkwargs,**kwargs}
for k,v in kwargs.items():
if isinstance(v,_Arg): kwargs[k] = args.pop(v.i)
fargs = [args[x.i] if isinstance(x, _Arg) else x for x in self.pargs] + args[self.maxi+1:]
return self.fn(*fargs, **kwargs)
def myfn(a,b,c,d=1,e=2): return(a,b,c,d,e)
test_eq(bind(myfn, arg1, 17, arg0, e=3)(19,14), (14,17,19,1,3))
test_eq(bind(myfn, 17, arg0, e=3)(19,14), (17,19,14,1,3))
test_eq(bind(myfn, 17, e=3)(19,14), (17,19,14,1,3))
test_eq(bind(myfn)(17,19,14), (17,19,14,1,2))
test_eq(bind(myfn, 17,19,14,e=arg0)(3), (17,19,14,1,3))
# ## GetAttr -
#export
class GetAttr:
"Inherit from this to have all attr accesses in `self._xtra` passed down to `self.default`"
_default='default'
@property
def _xtra(self): return [o for o in dir(getattr(self,self._default)) if not o.startswith('_')]
def __getattr__(self,k):
if k not in ('_xtra',self._default) and (self._xtra is None or k in self._xtra): return getattr(getattr(self,self._default), k)
raise AttributeError(k)
def __dir__(self): return custom_dir(self, self._xtra)
def __setstate__(self,data): self.__dict__.update(data)
# +
class _C(GetAttr):
_xtra = ['lower']
def __init__(self,a): self.default = a
def foo(self): noop
t = _C('Hi')
test_eq(t.lower(), 'hi')
test_fail(lambda: t.upper())
assert 'lower' in dir(t)
# -
#export
def delegate_attr(self, k, to):
"Use in `__getattr__` to delegate to attr `to` without inheriting from `GetAttr`"
if k.startswith('_') or k==to: raise AttributeError(k)
try: return getattr(getattr(self,to), k)
except AttributeError: raise AttributeError(k) from None
# +
class _C:
f = 'Hi'
def __getattr__(self, k): return delegate_attr(self, k, 'f')
t = _C()
test_eq(t.lower(), 'hi')
# -
# ## L -
# +
#export
def _is_array(x): return hasattr(x,'__array__') or hasattr(x,'iloc')
def _listify(o):
if o is None: return []
if isinstance(o, list): return o
if isinstance(o, str) or _is_array(o): return [o]
if is_iter(o): return list(o)
return [o]
# -
# export
def coll_repr(c, max_n=10):
"String repr of up to `max_n` items of (possibly lazy) collection `c`"
return f'(#{len(c)}) [' + ','.join(itertools.islice(map(str,c), max_n)) + (
'...' if len(c)>10 else '') + ']'
test_eq(coll_repr(range(1000), 5), '(#1000) [0,1,2,3,4...]')
# export
def mask2idxs(mask):
"Convert bool mask or index list to index `L`"
if isinstance(mask,slice): return mask
mask = list(mask)
if len(mask)==0: return []
it = mask[0]
if hasattr(it,'item'): it = it.item()
if isinstance(it,(bool,NoneType,np.bool_)): return [i for i,m in enumerate(mask) if m]
return [int(i) for i in mask]
# just for tests
import torch
test_eq(mask2idxs([False,True,False,True]), [1,3])
test_eq(mask2idxs(array([False,True,False,True])), [1,3])
test_eq(mask2idxs(torch.tensor([False,True,False,True])), [1,3])
test_eq(mask2idxs(array([1,2,3])), [1,2,3])
#export
listable_types = typing.Collection,Generator,map,filter,zip
#export
class CollBase:
"Base class for composing a list of `items`"
def __init__(self, items): self.items = items
def __len__(self): return len(self.items)
def __getitem__(self, k): return self.items[k]
def __setitem__(self, k, v): self.items[list(k) if isinstance(k,CollBase) else k] = v
def __delitem__(self, i): del(self.items[i])
def __repr__(self): return self.items.__repr__()
def __iter__(self): return self.items.__iter__()
#export
def cycle(o):
"Like `itertools.cycle` except creates list of `None`s if `o` is empty"
o = _listify(o)
return itertools.cycle(o) if o is not None and len(o) > 0 else itertools.cycle([None])
test_eq(itertools.islice(cycle([1,2,3]),5), [1,2,3,1,2])
test_eq(itertools.islice(cycle([]),3), [None]*3)
test_eq(itertools.islice(cycle(None),3), [None]*3)
test_eq(itertools.islice(cycle(1),3), [1,1,1])
#export
def zip_cycle(x, *args):
"Like `itertools.zip_longest` but `cycle`s through elements of all but first argument"
return zip(x, *map(cycle,args))
test_eq(zip_cycle([1,2,3,4],list('abc')), [(1, 'a'), (2, 'b'), (3, 'c'), (4, 'a')])
#export
def is_indexer(idx):
"Test whether `idx` will index a single item in a list"
return isinstance(idx,int) or not getattr(idx,'ndim',1)
#export
def negate_func(f):
"Create new function that negates result of `f`"
def _f(*args, **kwargs): return not f(*args, **kwargs)
return _f
def f(a): return a>0
test_eq(f(1),True)
test_eq(negate_func(f)(1),False)
test_eq(negate_func(f)(a=-1),True)
#export
class L(CollBase, GetAttr, metaclass=NewChkMeta):
"Behaves like a list of `items` but can also index with list of indices or masks"
_default='items'
def __init__(self, items=None, *rest, use_list=False, match=None):
if rest: items = (items,)+rest
if items is None: items = []
if (use_list is not None) or not _is_array(items):
items = list(items) if use_list else _listify(items)
if match is not None:
if is_coll(match): match = len(match)
if len(items)==1: items = items*match
else: assert len(items)==match, 'Match length mismatch'
super().__init__(items)
def _new(self, items, *args, **kwargs): return type(self)(items, *args, use_list=None, **kwargs)
def __getitem__(self, idx): return self._get(idx) if is_indexer(idx) else L(self._get(idx), use_list=None)
def _get(self, i):
if is_indexer(i) or isinstance(i,slice): return getattr(self.items,'iloc',self.items)[i]
i = mask2idxs(i)
return (self.items.iloc[list(i)] if hasattr(self.items,'iloc')
else self.items.__array__()[(i,)] if hasattr(self.items,'__array__')
else [self.items[i_] for i_ in i])
def __setitem__(self, idx, o):
"Set `idx` (can be list of indices, or mask, or int) items to `o` (which is broadcast if not iterable)"
idx = idx if isinstance(idx,L) else _listify(idx)
if not is_iter(o): o = [o]*len(idx)
for i,o_ in zip(idx,o): self.items[i] = o_
def __iter__(self): return iter(self.items.itertuples() if hasattr(self.items,'iloc') else self.items)
def __contains__(self,b): return b in self.items
def __invert__(self): return self._new(not i for i in self)
def __eq__(self,b): return False if isinstance(b, (str,dict,set)) else all_equal(b,self)
def __repr__(self): return repr(self.items) if _is_array(self.items) else coll_repr(self)
def __mul__ (a,b): return a._new(a.items*b)
def __add__ (a,b): return a._new(a.items+_listify(b))
def __radd__(a,b): return a._new(b)+a
def __addi__(a,b):
a.items += list(b)
return a
def sorted(self, key=None, reverse=False):
if isinstance(key,str): k=lambda o:getattr(o,key,0)
elif isinstance(key,int): k=itemgetter(key)
else: k=key
return self._new(sorted(self.items, key=k, reverse=reverse))
@classmethod
def split(cls, s, sep=None, maxsplit=-1): return cls(s.split(sep,maxsplit))
@classmethod
def range(cls, a, b=None, step=None):
if is_coll(a): a = len(a)
return cls(range(a,b,step) if step is not None else range(a,b) if b is not None else range(a))
def map(self, f, *args, **kwargs):
g = (bind(f,*args,**kwargs) if callable(f)
else f.format if isinstance(f,str)
else f.__getitem__)
return self._new(map(g, self))
def filter(self, f, negate=False, **kwargs):
if kwargs: f = partial(f,**kwargs)
if negate: f = negate_func(f)
return self._new(filter(f, self))
def unique(self): return L(dict.fromkeys(self).keys())
def enumerate(self): return L(enumerate(self))
def val2idx(self): return {v:k for k,v in self.enumerate()}
def itemgot(self, idx): return self.map(itemgetter(idx))
def attrgot(self, k, default=None): return self.map(lambda o:getattr(o,k,default))
def cycle(self): return cycle(self)
def map_dict(self, f=noop, *args, **kwargs): return {k:f(k, *args,**kwargs) for k in self}
def starmap(self, f, *args, **kwargs): return self._new(itertools.starmap(partial(f,*args,**kwargs), self))
def zip(self, cycled=False): return self._new((zip_cycle if cycled else zip)(*self))
def zipwith(self, *rest, cycled=False): return self._new([self, *rest]).zip(cycled=cycled)
def map_zip(self, f, *args, cycled=False, **kwargs): return self.zip(cycled=cycled).starmap(f, *args, **kwargs)
def map_zipwith(self, f, *rest, cycled=False, **kwargs): return self.zipwith(*rest, cycled=cycled).starmap(f, **kwargs)
def concat(self): return self._new(itertools.chain.from_iterable(self.map(L)))
def shuffle(self):
it = copy(self.items)
random.shuffle(it)
return self._new(it)
#export
add_docs(L,
__getitem__="Retrieve `idx` (can be list of indices, or mask, or int) items",
range="Same as builtin `range`, but returns an `L`. Can pass a collection for `a`, to use `len(a)`",
split="Same as builtin `str.split`, but returns an `L`",
sorted="New `L` sorted by `key`. If key is str then use `attrgetter`. If key is int then use `itemgetter`",
unique="Unique items, in stable order",
val2idx="Dict from value to index",
filter="Create new `L` filtered by predicate `f`, passing `args` and `kwargs` to `f`",
map="Create new `L` with `f` applied to all `items`, passing `args` and `kwargs` to `f`",
map_dict="Like `map`, but creates a dict from `items` to function results",
starmap="Like `map`, but use `itertools.starmap`",
itemgot="Create new `L` with item `idx` of all `items`",
attrgot="Create new `L` with attr `k` of all `items`",
cycle="Same as `itertools.cycle`",
enumerate="Same as `enumerate`",
zip="Create new `L` with `zip(*items)`",
zipwith="Create new `L` with `self` zip with each of `*rest`",
map_zip="Combine `zip` and `starmap`",
map_zipwith="Combine `zipwith` and `starmap`",
concat="Concatenate all elements of list",
shuffle="Same as `random.shuffle`, but not inplace")
# You can create an `L` from an existing iterable (e.g. a list, range, etc) and access or modify it with an int list/tuple index, mask, int, or slice. All `list` methods can also be used with `L`.
t = L(range(12))
test_eq(t, list(range(12)))
test_ne(t, list(range(11)))
t.reverse()
test_eq(t[0], 11)
t[3] = "h"
test_eq(t[3], "h")
t[3,5] = ("j","k")
test_eq(t[3,5], ["j","k"])
test_eq(t, L(t))
test_eq(L(L(1,2),[3,4]), ([1,2],[3,4]))
t
# There are optimized indexers for arrays, tensors, and DataFrames.
# +
arr = np.arange(9).reshape(3,3)
t = L(arr, use_list=None)
test_eq(t[1,2], arr[[1,2]])
arr = np.arange(9).reshape(3,3)
t = L(arr, use_list=None)
test_eq(t[1,2], arr[[1,2]])
df = pd.DataFrame({'a':[1,2,3]})
t = L(df, use_list=None)
test_eq(t[1,2], L(pd.DataFrame({'a':[2,3]}, index=[1,2]), use_list=None))
# -
# You can also modify an `L` with `append`, `+`, and `*`.
t = L()
test_eq(t, [])
t.append(1)
test_eq(t, [1])
t += [3,2]
test_eq(t, [1,3,2])
t = t + [4]
test_eq(t, [1,3,2,4])
t = 5 + t
test_eq(t, [5,1,3,2,4])
test_eq(L(1,2,3), [1,2,3])
test_eq(L(1,2,3), L(1,2,3))
t = L(1)*5
t = t.map(operator.neg)
test_eq(t,[-1]*5)
test_eq(~L([True,False,False]), L([False,True,True]))
t = L(range(4))
test_eq(zip(t, L(1).cycle()), zip(range(4),(1,1,1,1)))
t = L.range(100)
test_shuffled(t,t.shuffle())
def _f(x,a=0): return x+a
t = L(1)*5
test_eq(t.map(_f), t)
test_eq(t.map(_f,1), [2]*5)
test_eq(t.map(_f,a=2), [3]*5)
# An `L` can be constructed from anything iterable, although tensors and arrays will not be iterated over on construction, unless you pass `use_list` to the constructor.
test_eq(L([1,2,3]),[1,2,3])
test_eq(L(L([1,2,3])),[1,2,3])
test_ne(L([1,2,3]),[1,2,])
test_eq(L('abc'),['abc'])
test_eq(L(range(0,3)),[0,1,2])
test_eq(L(o for o in range(0,3)),[0,1,2])
test_eq(L(array(0)),[array(0)])
test_eq(L([array(0),array(1)]),[array(0),array(1)])
test_eq(L(array([0.,1.1]))[0],array([0.,1.1]))
test_eq(L(array([0.,1.1]), use_list=True), [array(0.),array(1.1)]) # `use_list=True` to unwrap arrays/arrays
# If `match` is not `None` then the created list is same len as `match`, either by:
#
# - If `len(items)==1` then `items` is replicated,
# - Otherwise an error is raised if `match` and `items` are not already the same size.
test_eq(L(1,match=[1,2,3]),[1,1,1])
test_eq(L([1,2],match=[2,3]),[1,2])
test_fail(lambda: L([1,2],match=[1,2,3]))
# If you create an `L` from an existing `L` then you'll get back the original object (since `L` uses the `NewChkMeta` metaclass).
test_is(L(t), t)
# An `L` is considred equal to a list if they have the same elements. It's never considered equal to a `str` a `set` or a `dict` even if they have the same elements/keys.
test_eq(L(['a', 'b']), ['a', 'b'])
test_ne(L(['a', 'b']), 'ab')
test_ne(L(['a', 'b']), {'a', 'b'})
test_ne(L(['a', 'b']), {'a':1, 'b':2})
# ### Methods
show_doc(L.__getitem__)
t = L(range(12))
test_eq(t[1,2], [1,2]) # implicit tuple
test_eq(t[[1,2]], [1,2]) # list
test_eq(t[:3], [0,1,2]) # slice
test_eq(t[[False]*11 + [True]], [11]) # mask
test_eq(t[array(3)], 3)
show_doc(L.__setitem__)
t[4,6] = 0
test_eq(t[4,6], [0,0])
t[4,6] = [1,2]
test_eq(t[4,6], [1,2])
show_doc(L.unique)
test_eq(L(1,2,3,4,4).unique(), [1,2,3,4])
show_doc(L.val2idx)
test_eq(L(1,2,3).val2idx(), {3:2,1:0,2:1})
show_doc(L.filter)
list(t)
test_eq(t.filter(lambda o:o<5), [0,1,2,3,1,2])
test_eq(t.filter(lambda o:o<5, negate=True), [5,7,8,9,10,11])
show_doc(L.map)
test_eq(L.range(4).map(operator.neg), [0,-1,-2,-3])
# If `f` is a string then it is treated as a format string to create the mapping:
test_eq(L.range(4).map('#{}#'), ['#0#','#1#','#2#','#3#'])
# If `f` is a dictionary (or anything supporting `__getitem__`) then it is indexed to create the mapping:
test_eq(L.range(4).map(list('abcd')), list('abcd'))
# If the special argument `_arg` is passed, then that is the kwarg used in the map.
# +
#What is this? TODO Jeremy: fix
#L.range(4).map(f, b=arg0)
# -
def f(a=None,b=None): return b
test_eq(L.range(4).map(f, b=arg0), range(4))
show_doc(L.map_dict)
test_eq(L(range(1,5)).map_dict(), {1:1, 2:2, 3:3, 4:4})
test_eq(L(range(1,5)).map_dict(operator.neg), {1:-1, 2:-2, 3:-3, 4:-4})
show_doc(L.zip)
t = L([[1,2,3],'abc'])
test_eq(t.zip(), [(1, 'a'),(2, 'b'),(3, 'c')])
t = L([[1,2,3,4],['a','b','c']])
test_eq(t.zip(cycled=True ), [(1, 'a'),(2, 'b'),(3, 'c'),(4, 'a')])
test_eq(t.zip(cycled=False), [(1, 'a'),(2, 'b'),(3, 'c')])
show_doc(L.map_zip)
t = L([1,2,3],[2,3,4])
test_eq(t.map_zip(operator.mul), [2,6,12])
show_doc(L.zipwith)
b = [[0],[1],[2,2]]
t = L([1,2,3]).zipwith(b)
test_eq(t, [(1,[0]), (2,[1]), (3,[2,2])])
show_doc(L.map_zipwith)
test_eq(L(1,2,3).map_zipwith(operator.mul, [2,3,4]), [2,6,12])
show_doc(L.itemgot)
test_eq(t.itemgot(1), b)
show_doc(L.attrgot)
a = [SimpleNamespace(a=3,b=4),SimpleNamespace(a=1,b=2)]
test_eq(L(a).attrgot('b'), [4,2])
show_doc(L.sorted)
test_eq(L(a).sorted('a').attrgot('b'), [2,4])
show_doc(L.split)
test_eq(L.split('a b c'), list('abc'))
show_doc(L.range)
test_eq_type(L.range([1,1,1]), L(range(3)))
test_eq_type(L.range(5,2,2), L(range(5,2,2)))
show_doc(L.concat)
test_eq(L([0,1,2,3],4,L(5,6)).concat(), range(7))
# # Export -
#hide
from local.notebook.export import notebook2script
notebook2script(all_fs=True)
| dev/01_core.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 这只是一个普通的pipeline,意义不大。具体请看4.3节。
import re
import torch
import torch.nn as nn
from transformers import BertForTokenClassification, BertTokenizer
from transformers import AdamW
from torch.utils.data import TensorDataset, DataLoader, RandomSampler, SequentialSampler
from sklearn.model_selection import train_test_split
import numpy as np
from tqdm import tqdm, trange
file = "biaoji.txt"
all_ner_data = []
with open(file, encoding="utf-8") as f:
for s in f.readlines():
s = s.strip('\n')
ner_data = []
result_1 = re.finditer(r'\[\@', s)
result_2 = re.finditer(r'\*\]', s)
begin = []
end = []
for each in result_1:
begin.append(each.start())
for each in result_2:
end.append(each.end())
assert len(begin) == len(end)
i = 0
j = 0
while i < len(s):
if i not in begin:
ner_data.append([s[i], 'O'])
i = i + 1
else:
ann = s[i + 2:end[j] - 2]
entity, ner = ann.rsplit('#')
if (len(entity) == 1):
ner_data.append([entity, 'S-' + ner])
else:
if (len(entity) == 2):
ner_data.append([entity[0], 'B-' + ner])
ner_data.append([entity[1], 'E-' + ner])
else:
ner_data.append([entity[0], 'B-' + ner])
for n in range(1, len(entity) - 1):
ner_data.append([entity[n], 'I-' + ner])
ner_data.append([entity[-1], 'E-' + ner])
i = end[j]
j = j + 1
all_ner_data.append(ner_data)
f.close()
all_ner_data_list = []
for seq_list in all_ner_data:
zi = []
mark = []
for zi_mark in seq_list:
zi.append(zi_mark[0])
mark.append(zi_mark[1])
seq_tuple = (zi, mark)
all_ner_data_list.append(seq_tuple)
all_ner_data_list[0]
all_ner_data_list[1]
all_ner_data_list[2]
# ### 输入处理
#
# 由于BERT模型的特殊性,需要再处理一下输入:
#
# - input_ids(padding)
# - attention_masks
# - labels
from keras.preprocessing.sequence import pad_sequences # padding
print("Is CUDA available: ", torch.cuda.is_available())
n_gpu = torch.cuda.device_count()
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
print("GPU numbers: ", n_gpu)
print("device_name: ", torch.cuda.get_device_name(0))
# +
# 构建 tag 到 索引 的字典
tag_to_ix = {"B-Location": 0,
"I-Location": 1,
"E-Location": 2,
"O": 3,
"[CLS]":4,
"[SEP]":5,
"[PAD]":6}
ix_to_tag = {0:"B-Location",
1:"I-Location",
2:"E-Location",
3:"O",
4:"[CLS]",
5:"[SEP]",
6:"[PAD]"}
# -
# +
all_sentences = [] # 句子
all_labels = [] # labels
for seq_pair in all_ner_data_list:
sentence = "".join(seq_pair[0])
labels = [tag_to_ix[t] for t in seq_pair[1]]
all_sentences.append(sentence)
all_labels.append(labels)
print(all_sentences)
print(all_labels)
# -
# padding
tokenizer = BertTokenizer.from_pretrained('./bert-chinese/', do_lower_case=True)
tokenized_texts = [tokenizer.encode(sent, add_special_tokens=True) for sent in all_sentences]
tokenized_texts[0]
# +
# 句子padding
# 句子最长长度
MAX_LEN = 32
# 输入padding
# 此函数在keras里面
input_ids = pad_sequences([txt for txt in tokenized_texts],
maxlen=MAX_LEN,
dtype="long",
truncating="post",
padding="post")
# -
print(len(input_ids[0]))
print(input_ids[0])
# [3] 代表 O 实体
for label in all_labels:
label.insert(len(label), 5) # [SEP]
label.insert(0, 4) # [CLS]
if MAX_LEN > len(label) -1:
for i in range(MAX_LEN - len(label)):
label.append(3) # [PAD]
print(len(all_labels[0]))
print(all_labels[0])
# +
# 创建attention masks
attention_masks = []
# Create a mask of 1s for each token followed by 0s for padding
for seq in input_ids:
seq_mask = [float(i > 0) for i in seq]
attention_masks.append(seq_mask)
# -
# 第一句话的 attention_masks
print(np.array(attention_masks[0]))
print(len(np.array(attention_masks[0])))
# ### 训练集和验证集分开
train_inputs, validation_inputs, train_labels, validation_labels = train_test_split(input_ids,
all_labels,
random_state=2019,
test_size=0.1)
train_masks, validation_masks, _, _ = train_test_split(attention_masks,
input_ids,
random_state=2019,
test_size=0.1)
# +
print(len(train_inputs))
print(len(validation_inputs))
print(train_inputs[0])
print(validation_inputs[0])
# -
# tensor化
train_inputs = torch.tensor(train_inputs)
validation_inputs = torch.tensor(validation_inputs)
train_labels = torch.tensor(train_labels)
validation_labels = torch.tensor(validation_labels)
train_masks = torch.tensor(train_masks)
validation_masks = torch.tensor(validation_masks)
train_inputs
train_labels
train_masks
# ### 创建迭代器
# +
# batch size
batch_size = 16
# 形成训练数据集
train_data = TensorDataset(train_inputs, train_masks, train_labels)
# 随机采样
train_sampler = RandomSampler(train_data)
# 读取数据
train_dataloader = DataLoader(train_data, sampler=train_sampler, batch_size=batch_size)
# 形成验证数据集
validation_data = TensorDataset(validation_inputs, validation_masks, validation_labels)
# 随机采样
validation_sampler = SequentialSampler(validation_data)
# 读取数据
validation_dataloader = DataLoader(validation_data, sampler=validation_sampler, batch_size=batch_size)
# -
# ### BERT的微调
model = BertForTokenClassification.from_pretrained("./bert-chinese/", num_labels=7)
model.cuda()
# +
# BERT fine-tuning parameters
param_optimizer = list(model.named_parameters())
no_decay = ['bias', 'LayerNorm.weight']
# 权重衰减
optimizer_grouped_parameters = [
{'params': [p for n, p in param_optimizer if not any(nd in n for nd in no_decay)],
'weight_decay': 0.01},
{'params': [p for n, p in param_optimizer if any(nd in n for nd in no_decay)],
'weight_decay': 0.0}]
# -
# 优化器
optimizer = AdamW(optimizer_grouped_parameters,
lr=5e-5)
# 保存loss
train_loss_set = []
# epochs
epochs = 100
# BERT training loop
for _ in trange(epochs):
## 训练
print(f"当前epoch: {_}")
# 开启训练模式
model.train()
tr_loss = 0 # train loss
nb_tr_examples, nb_tr_steps = 0, 0
# Train the data for one epoch
for step, batch in enumerate(train_dataloader):
# 把batch放入GPU
batch = tuple(t.to(device) for t in batch)
# 解包batch
b_input_ids, b_input_mask, b_labels = batch
# 梯度归零
optimizer.zero_grad()
# 前向传播loss计算
output = model(input_ids=b_input_ids,
attention_mask=b_input_mask,
labels=b_labels)
loss = output[0]
# print(loss)
# 反向传播
loss.backward()
# Update parameters and take a step using the computed gradient
# 更新模型参数
optimizer.step()
# Update tracking variables
tr_loss += loss.item()
nb_tr_examples += b_input_ids.size(0)
nb_tr_steps += 1
print(f"当前 epoch 的 Train loss: {tr_loss/nb_tr_steps}")
# ## 验证
# +
# 验证状态
model.eval()
# 建立变量
eval_loss, eval_accuracy = 0, 0
nb_eval_steps, nb_eval_examples = 0, 0
# Evaluate data for one epoch
# -
# 验证集的读取也要batch
for batch in tqdm(validation_dataloader):
# 元组打包放进GPU
batch = tuple(t.to(device) for t in batch)
# 解开元组
b_input_ids, b_input_mask, b_labels = batch
# 预测
with torch.no_grad():
# segment embeddings,如果没有就是全0,表示单句
# position embeddings,[0,句子长度-1]
outputs = model(input_ids=b_input_ids,
attention_mask=b_input_mask,
token_type_ids=None,
position_ids=None)
# print(logits[0])
# Move logits and labels to CPU
scores = outputs[0].detach().cpu().numpy() # 每个字的标签的概率
pred_flat = np.argmax(scores[0], axis=1).flatten()
label_ids = b_labels.to('cpu').numpy() # 真实labels
# print(logits, label_ids)
pred_flat # 预测值
label_ids # 真实值
# 这句话
test_tokens = b_input_ids[0].cpu().numpy()
tokenizer.decode(test_tokens)
outputs
| 04其他模型教程/4.02 BERT-NER.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.3 64-bit (''base'': conda)'
# language: python
# name: python3
# ---
# default_exp special
from nbdev import *
from nbdev.showdoc import *
# # Context free grammar
#
# > A collection of somewhat handy functions to make your AoC puzzle life solving a bit easier
# ## Context free grammar
# Takes a grammar, converts it into Chomsky Normal Form (CNF)
#exporti
from collections import defaultdict
from itertools import product
# +
#export
class CFG():
""""Takes a grammer as dict with tuple of options as values. Terminal values should not be in a tuple but as a string
Usage:
cfg = CFG(grammar_dict)
reverse as optional parameter when k,v are reversed
converts the grammar to Chomsky Normal form by taking care of options, unit productions and triplets
cfg.solve(messages_list) returns dict of substrings with possible rules to make them
"""
def __init__(self, grammar = None, reverse = True):
self.outcomes = defaultdict(set)
if grammar:
# convert grammar to CNF and add terminals to outcomes
self.grammar = self.grammar_to_cnf(grammar, reverse)
self.outcomes.update({k:v for k,v in self.grammar.items() if isinstance(k, str)})
def grammar_to_cnf(self, grammar, reverse):
grammar = self.to_cnf_remove_options(grammar, reverse)
grammar = self.to_cnf_remove_triplets(grammar)
return self.to_cnf_remove_unit_productions(grammar)
def to_cnf_remove_options(self, grammar, reverse):
# if reverse change from X : AB to AB : {X}
# if there are options, these are given a separate entry, e.g.
# X : (AB, CD) --> X: AB and X: CD
new_grammar = defaultdict(set)
if reverse:
for k,v in grammar.items():
for option in v:
new_grammar[option].add(k)
else:
for k,v in grammar.items():
for option in k:
new_grammar[option].add(v)
return new_grammar
def to_cnf_remove_triplets(self, grammar):
# reduces triplets or larger to pairs
# changes X : ABC to
# X: AY, Y = BC
new_grammar = defaultdict(set)
for k,v in grammar.items():
if len(k) > 2:
for i, t in enumerate(k[0:-2]):
newvar = str(v) + '_' + str(i)
oldvar = str(v) + '_' + str(i-1)
if i == 0:
new_grammar[t,newvar] = v
else:
new_grammar[t,newvar] = {oldvar}
new_grammar[k[-2:]].add(newvar)
else:
new_grammar[k] |= v
return new_grammar
def to_cnf_remove_unit_productions(self,grammar):
# step to get to Chomsky Normal Form
# if X : A, duplicate all A : Y with X : Y
singulars = {k[0]:next(iter(v)) for k,v in grammar.items() if len(k)!=2 and not isinstance(k,str)}
for k,v in singulars.items():
for j in grammar.values():
if k in j:
j.add(v)
return grammar
def pieces(self, test,l):
# gets all possibilities of len l out of a string
assert isinstance(test, str)
return {test[i:i+l] for i in range(len(test)-l+1) if test[i:i+l] not in self.outcomes}
def splitter(self,option):
# splits string into all options of two substrings
assert isinstance(option, str)
return {(option[:i], option[i:]) for i in range(1,len(option))}
def check_possible_option(self, option):
first = self.outcomes[option[0]]
second = self.outcomes[option[1]]
res = set()
for potential in product(first,second):
if potential in self.grammar: res |= self.grammar[potential]
return res
def solve(self, messages):
# takes a list of messages and returns all possibilities for the substrings of m
for num, m in enumerate(messages):
if num % 100 == 0: print(num*10, 'messages done')
for i in range(2,len(m)+1):
for j in self.pieces(m, i):
for k in self.splitter(j):
res = self.check_possible_option(k)
if res:
self.outcomes[j] |= res
print('finished all messages, returning dict')
return self.outcomes
# -
cfg = CFG()
assert cfg.pieces('abcde',3) == {'abc', 'bcd', 'cde'}
assert cfg.splitter('abcd') == {('a', 'bcd'), ('ab', 'cd'), ('abc', 'd')}
# +
grammar = {'0': (('4', '1', '5'),),
'1': (('2', '3'), ('3', '2')),
'2': (('4', '4'), ('5', '5')),
'3': (('4', '5'), ('5', '4')),
'4': ('a',),
'5': ('b',)}
messages = ['ababbb', 'bababa', 'abbbab', 'aaabbb', 'aaaabbb']
cfg = CFG(grammar)
out = cfg.solve(messages)
assert sum([1 for m in messages if (m in out) and ('0' in out[m])]) == 2
# -
cfg.grammar.values()
# +
rules, messages = open('test.txt').read().split('\n\n')
grammar = {}
for rule in rules.split('\n'):
num, makefrom = rule.split(': ')
makefrom = makefrom.split(' | ')
if len(makefrom) > 1:
grammar[num] = tuple([tuple(option.split()) for option in makefrom])
else:
grammar[num] = tuple(makefrom[0].split())
for k, v in grammar.items():
if 'a' in v[0]:
grammar[k] = 'a',
if 'b' in v[0]:
grammar[k] = 'b',
messages = messages.split('\n')
cfg = CFG(grammar)
res = cfg.solve(messages)
# -
| 06_context_free_grammar.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.cm as cm
from astropy.table import Table
import emcee
# -
import matplotlib
matplotlib.rcParams.update({'font.size':18})
matplotlib.rcParams.update({'font.family':'serif'})
# +
# TESS Luminosity
LUMIN = 31.061790465873248 # from "padova_lum" notebook
LUMIN_err = 0.039444830755122
# Kepler Luminosity
E_POINT = 30.67828435767249 # from "padova_lum" notebook
E_POINT_err = 0.04457627419541614
gj1243_dir = '../gj1243_10yrs/'
file = gj1243_dir + 'gj1243_tess.dat.fbeye'
colnames = ('Event_ID', 'Start_INDX', 'Stop_INDX', 't_peak', 't_start', 't_stop', 't_rise', 't_decay',
'Flux_peak', 'Equiv_Dur', 'S/N', 'CPLX_flg', 'MLTPK_flg', 'MLTPK_num', 't_MLTPK', 'L_mltpk',
'MLTPK_INDX', 'quies','x')
fbeye = pd.read_table(file, sep='\s+', comment='#', names=colnames)[1:] # skip the first junk row
file2 = gj1243_dir + 'gj1243_tess15.dat.fbeye'
fbeye2 = pd.read_table(file2, sep='\s+', comment='#', names=colnames)[1:] # skip the first junk row
file = gj1243_dir + 'tess2019198215352-s0014-0000000273589987-0150-s_lc.fits'
tbl = Table.read(file, format='fits')
q = tbl['QUALITY'] == 0
file2 = gj1243_dir + 'tess2019226182529-s0015-0000000273589987-0151-s_lc.fits'
tbl2 = Table.read(file2, format='fits')
q2 = tbl2['QUALITY'] == 0
dt = np.nanmedian(np.diff(tbl['TIME'][q]))
ED = np.sort(np.concatenate((fbeye['Equiv_Dur'].values, fbeye2['Equiv_Dur'].values)))[::-1]
TOTDUR = dt * (sum(q) + sum(q2)) # days
ffd_x = np.log10(ED) + LUMIN
ffd_y = np.log10(np.arange(1, len(ED)+1) / TOTDUR)
plt.figure(figsize=(9,7))
plt.plot(ffd_x, ffd_y, '-o', color='C0', alpha=0.3)
plt.xlabel('log E (erg)')
plt.ylabel(r'log $\nu$ (day$^{-1}$)')
plt.title('GJ 1243')
# +
# gj1243 : G-J = 11.55090 - 8.586
# YZCMi : G-J = 9.6807 - 6.58
print(11.55090 - 8.586, 9.6807 - 6.58)
# +
# LUMINY = 31.00192752813018 # from updated notebook (gj1243_10yrs/spectra/padova_lum)
# LUMINY_err = 0.22985294989763078
LUMINY = 30.976368864140518 # from updated notebook (gj1243_10yrs/spectra/padova_lum)
LUMINY_err = 0.1148749988137026 # but using i-J instead of g-J b/c larger errors in g!
file = 'tess2019006130736-s0007-0000000266744225-0131-s_lc.fits.dat.fbeye'
fbeyeY = pd.read_table(file, sep='\s+', comment='#', names=colnames)[1:] # skip the first junk row
file = 'tess2019006130736-s0007-0000000266744225-0131-s_lc.fits'
tblY = Table.read(file, format='fits')
qY = tblY['QUALITY'] == 0
dt = np.nanmedian(np.diff(tblY['TIME'][qY]))
####
EDY = np.sort(fbeyeY['Equiv_Dur'].values)[::-1]
TOTDURY = dt * sum(qY) # days
ffd_xY = np.log10(EDY) + LUMINY
ffd_yY = np.log10(np.arange(1, len(EDY)+1) / TOTDURY)
plt.figure(figsize=(9,7))
plt.plot(ffd_xY, ffd_yY, '-o', color='C1', alpha=0.7, label='YZ CMi, TESS S007')
plt.plot(ffd_x, ffd_y, '-o', color='C0', alpha=0.7, label='GJ 1243, TESS S014-S015')
plt.legend(fontsize=13)
plt.xlabel('log E (erg)')
plt.ylabel(r'log $\nu$ (day$^{-1}$)')
# plt.title('GJ 1243')
print(fbeyeY.shape)
# +
# plt.plot(tblY['TIME'][qY], tblY['PDCSAP_FLUX'][qY])
import exoplanet as xo
lnp = xo.lomb_scargle_estimator(tblY['TIME'][qY].data, tblY['PDCSAP_FLUX'][qY].data, min_period=0.2, max_period=15)
freq, pwr = lnp['periodogram']
plt.plot(1./freq, pwr)
plt.xscale('log')
print(lnp['peaks'][0]['period'])
# -
# +
import sys
sys.path
sys.path.append('/Users/james/python/FFD/')
from FFD import FFD, FlareKernel
#gj1243
ffd_x,ffd_ylog,ffd_xerr,ffd_yerrlog = FFD(np.concatenate((fbeye['Equiv_Dur'].values,
fbeye2['Equiv_Dur'].values)), TOTEXP=TOTDUR, Lum=LUMIN,
dur=np.concatenate((fbeye['t_stop'].values-fbeye['t_start'].values,
fbeye2['t_stop'].values-fbeye2['t_start'].values)),
fluxerr=np.nanmedian(tbl['PDCSAP_FLUX_ERR'][q] / np.nanmedian(tbl['PDCSAP_FLUX'][q])),logY=True)
ffd_x,ffd_y,ffd_xerr,ffd_yerr = FFD(np.concatenate((fbeye['Equiv_Dur'].values,
fbeye2['Equiv_Dur'].values)), TOTEXP=TOTDUR, Lum=LUMIN,
dur=np.concatenate((fbeye['t_stop'].values-fbeye['t_start'].values,
fbeye2['t_stop'].values-fbeye2['t_start'].values)),
fluxerr=np.nanmedian(tbl['PDCSAP_FLUX_ERR'][q] / np.nanmedian(tbl['PDCSAP_FLUX'][q])),logY=False)
#yzcmi
ffd_xY,ffd_yYlog,ffd_xerrY,ffd_yerrYlog = FFD(fbeyeY['Equiv_Dur'].values, TOTEXP=TOTDURY, Lum=LUMINY,
dur=fbeyeY['t_stop'].values-fbeyeY['t_start'].values,
fluxerr=np.nanmedian(tblY['PDCSAP_FLUX_ERR'][qY] / np.nanmedian(tblY['PDCSAP_FLUX'][qY])),logY=True)
ffd_xY,ffd_yY,ffd_xerrY,ffd_yerrY = FFD(fbeyeY['Equiv_Dur'].values, TOTEXP=TOTDURY, Lum=LUMINY,
dur=fbeyeY['t_stop'].values-fbeyeY['t_start'].values,
fluxerr=np.nanmedian(tblY['PDCSAP_FLUX_ERR'][qY] / np.nanmedian(tblY['PDCSAP_FLUX'][qY])),logY=False )
# +
plt.figure(figsize=(9,7))
plt.errorbar(ffd_x, ffd_y, xerr=np.sqrt(ffd_xerr**2+LUMIN_err**2),
yerr=ffd_yerr,
color='C0', alpha=0.25, linestyle='none', lw=4, marker='o')
plt.errorbar(ffd_xY, ffd_yY, xerr=np.sqrt(ffd_xerrY**2+LUMINY_err**2),
yerr=ffd_yerrY,
color='C1', alpha=0.25, linestyle='none', lw=4, marker='o')
plt.yscale('log')
plt.xlabel('log Energy (erg)')
plt.ylabel(r'Cumulative Flare Rate (day$^{-1}$)')
# plt.title('GJ 1243 (M4, P$_{rot}$=0.59d)', fontsize=14)
# plt.text(31.5, 5, 'Kepler',color='k')
plt.text(30.5, 0.7,'GJ 1243', color='C0')
plt.text(32.5, 0.7,'YZ CMi', color='C1')
plt.xlim(28.8,34)
plt.savefig('YZCMI_vs_GJ1243.png', dpi=150, bbox_inches='tight', pad_inches=0.25)
# -
# ## fitting the FFD
# there's lots of ways to fit a FFD, perhaps none of the good
from scipy.odr import ODR, Model, Data, RealData
def f(B, x):
if B[0] > 0:
return np.inf
# B[0] = -1 # fix slope? no
return 10**(B[0]*x + B[1])
def quickfit(x,y,xe,ye):
b00, b10 = -1., 30
linear = Model(f)
ok = np.where(np.isfinite(x) & np.isfinite(y) & np.isfinite(xe) & np.isfinite(ye))[0]
mydata = RealData(x[ok], y[ok], sx=xe[ok], sy=ye[ok])
myodr = ODR(mydata, linear, beta0=[b00, b10])
myoutput = myodr.run()
m, b = myoutput.beta[0], myoutput.beta[1]
m_err, b_err = myoutput.sd_beta[0], myoutput.sd_beta[1]
return m,b,m_err,b_err
# +
plt.figure(figsize=(9,7))
plt.errorbar(ffd_x, ffd_y, xerr=np.sqrt(ffd_xerr**2+LUMIN_err**2),
yerr=ffd_yerr,
color='C0', alpha=0.25, linestyle='none', lw=1, marker='.')
plt.errorbar(ffd_xY, ffd_yY, xerr=np.sqrt(ffd_xerrY**2+LUMINY_err**2),
yerr=ffd_yerrY,
color='C1', alpha=0.25, linestyle='none', lw=1, marker='.')
m,b,m_err,b_err = quickfit(ffd_x, ffd_y,
np.sqrt(ffd_xerr**2+LUMIN_err**2), ffd_yerr)
mY,bY,m_errY,b_errY = quickfit(ffd_xY, ffd_yY,
np.sqrt(ffd_xerrY**2+LUMINY_err**2), ffd_yerrY)
plt.plot(ffd_x, 10**(m*ffd_x + b))
plt.plot(ffd_xY, 10**(mY*ffd_xY + bY))
plt.yscale('log')
plt.xlabel('log Energy (erg)')
plt.ylabel(r'Cumulative Flare Rate (day$^{-1}$)')
# plt.title('GJ 1243 (M4, P$_{rot}$=0.59d)', fontsize=14)
# plt.text(31.5, 5, 'Kepler',color='k')
plt.text(30.5, 0.7,'GJ 1243', color='C0')
plt.text(32.5, 0.7,'YZ CMi', color='C1')
plt.xlim(28.8,34)
plt.title('Fit: ODR')
# plt.savefig('YZCMI_vs_GJ1243.png', dpi=150, bbox_inches='tight', pad_inches=0.25)
print(m,b,m_err,b_err)
print(mY,bY,m_errY,b_errY)
# -
def therng(thething, rng=[15,50,84]):
pcent = np.percentile(thething, rng)
q = np.diff(pcent)
return pcent[1], q[0], q[1]
def FFD_mcmc(ffd_x, ffd_y, ffd_xerr, ffd_yerr, Nmin=10, Nsteps=2000):
def _lnProb(theta, x, y, xerr, yerr):
m, b, xmin = theta
model = 10**(m * x + b)
xmodel = (np.log10(y) - b)/m
xok = x > xmin
# I hate that Python lets variables creep into scope like this, but OK!
if (sum(xok) >= Nmin) and (-5 < m < -0) and (0 < b < 100) and ( 0 < xmin < np.max(x)):
# my 1st attempt at reduced chi-sq, trying to approximate X and Y errors
# chisq = (np.sum(((y[xok] - model[xok])/yerr[xok] )** 2) +
# np.sum(((x[xok] - xmodel[xok])/xerr[xok] )** 2)) / np.float(np.sum(xok))
# my 2nd attempt, better i think
chisq = np.sum(((y[xok] - model[xok])/yerr[xok])** 2 *
((x[xok] - xmodel[xok])/xerr[xok])** 2) / np.float(np.sum(xok))
BIC = chisq + np.size(theta) * np.log(len(x)+(np.float(len(x) - np.sum(xok))))
return -0.5 * BIC
# return -0.5 * chisq
return -np.inf
# initialization
xok = (ffd_x >= np.mean(ffd_x))
fit, cov = np.polyfit(ffd_x[xok], np.log10(ffd_y[xok]), 1,
w=1./((ffd_yerr[xok]**2 + (-1 * ffd_xerr[xok])**2)), cov=True)
pos = np.array([fit[0], fit[1], np.mean(ffd_x)]) + 1e-4 * np.random.randn(32, 3)
nwalkers, ndim = pos.shape
sampler = emcee.EnsembleSampler(nwalkers, ndim, _lnProb,
args=(ffd_x, ffd_y, ffd_xerr, ffd_yerr))
sampler.run_mcmc(pos, Nsteps, progress=True);
samples = sampler.get_chain()
flat_samples = sampler.get_chain(discard=int(Nsteps*0.1), thin=15, flat=True)
return flat_samples
flat_samplesY = FFD_mcmc(ffd_xY, ffd_yY, ffd_xerrY, ffd_yerrY, Nsteps=1000)
# flat_samplesY = FFD_mcmc(ffd_xY, ffd_yY,
# np.sqrt(ffd_xerrY**2+(LUMINY_err)**2), ffd_yerrY, Nsteps=1000)
print(np.median(flat_samplesY, axis=0))
# +
plt.figure(figsize=(9,7))
inds = np.random.randint(len(flat_samplesY), size=100)
for ind in inds:
sample = flat_samplesY[ind]
plt.plot([sample[2], max(ffd_xY)], 10**(sample[0]*np.array([sample[2], max(ffd_xY)]) + sample[1]),
"k", alpha=0.1)
plt.errorbar(ffd_xY, ffd_yY, xerr=np.sqrt(ffd_xerrY**2+LUMINY_err**2),
yerr=ffd_yerrY,
color='C1', alpha=0.25, linestyle='none', lw=1, marker='.')
plt.plot(ffd_xY[ffd_xY >= np.median(flat_samplesY[:,2])],
10**(np.median(flat_samplesY[:,0]) * ffd_xY[ffd_xY >= np.median(flat_samplesY[:,2])] + np.median(flat_samplesY[:,1])),
c='r', lw=3)
plt.yscale('log')
plt.xlim(29.5,33.5)
plt.ylim(3e-2, 1e1)
plt.xlabel('log Energy (erg)')
plt.ylabel(r'Cumulative Flare Rate (day$^{-1}$)')
print(therng(flat_samplesY[:,0]))
print(therng(flat_samplesY[:,1]))
print(therng(flat_samplesY[:,2]))
# -
# flat_samples = FFD_mcmc(ffd_x, ffd_y, np.sqrt(ffd_xerr**2+LUMIN_err**2), ffd_yerr, Nsteps=1000)
flat_samples = FFD_mcmc(ffd_x, ffd_y, ffd_xerr, ffd_yerr, Nsteps=1000)
print(np.median(flat_samples, axis=0))
# +
plt.figure(figsize=(9,7))
inds = np.random.randint(len(flat_samples), size=100)
for ind in inds:
sample = flat_samples[ind]
plt.plot([sample[2], max(ffd_x)], 10**(sample[0]*np.array([sample[2], max(ffd_x)]) + sample[1]), "k", alpha=0.1)
plt.errorbar(ffd_x, ffd_y, xerr=np.sqrt(ffd_xerr**2+LUMIN_err**2),
yerr=ffd_yerr,
color='C0', alpha=0.25, linestyle='none', lw=1, marker='.')
plt.plot(ffd_x[ffd_x >= np.median(flat_samples[:,2])],
10**(np.median(flat_samples[:,0]) * ffd_x[ffd_x >= np.median(flat_samples[:,2])] + np.median(flat_samples[:,1])),
c='r', lw=3)
plt.yscale('log')
plt.xlim(29.5,33.5)
plt.ylim(3e-2, 1e1)
print(therng(flat_samples[:,0]))
print(therng(flat_samples[:,1]))
print(therng(flat_samples[:,2]))
# -
# +
plt.figure(figsize=(9,7))
plt.errorbar(ffd_x, ffd_y, xerr=ffd_xerr,#np.sqrt(ffd_xerr**2+LUMIN_err**2),
yerr=ffd_yerr,
color='C0', alpha=0.25, linestyle='none', lw=1, marker='.')
plt.errorbar(ffd_xY, ffd_yY, xerr=ffd_xerrY,#np.sqrt(ffd_xerrY**2+LUMINY_err**2),
yerr=ffd_yerrY,
color='C1', alpha=0.25, linestyle='none', lw=1, marker='.')
plt.plot(ffd_x[ffd_x >= np.median(flat_samples[:,2])],
10**(np.median(flat_samples[:,0]) * ffd_x[ffd_x >= np.median(flat_samples[:,2])] + np.median(flat_samples[:,1])),
c='r', lw=3)
plt.plot(ffd_xY[ffd_xY >= np.median(flat_samplesY[:,2])],
10**(np.median(flat_samplesY[:,0]) * ffd_xY[ffd_xY >= np.median(flat_samplesY[:,2])] + np.median(flat_samplesY[:,1])),
c='r', lw=3)
plt.yscale('log')
plt.xlim(29.5,33.5)
plt.ylim(3e-2, 1e1)
# -
def nrpt(x3, y3, m, b):
''' find the nearest point on a line '''
# https://stackoverflow.com/a/47198877
x1 = 0.
x2 = x3
y1 = x1*m + b
y2 = x2*m + b
dx, dy = x2-x1, y2-y1
det = dx*dx + dy*dy
a = (dy*(y3-y1) + dx*(x3-x1)) / det
return x1+a*dx, y1+a*dy
# +
xok = (ffd_x >= 32)
fit = np.polyfit(ffd_x[xok], ffd_ylog[xok], 1)
print(fit)
# fit = [-1, -1]
plt.errorbar(ffd_x[xok], ffd_ylog[xok], xerr=ffd_xerr[xok], yerr=ffd_yerrlog[xok])
# plt.scatter(ffd_x, ffd_ylog, c='C1')
plt.plot(ffd_x[xok], np.polyval(fit, ffd_x[xok]), c='C2')
xnew, ynew = nrpt(ffd_x, ffd_ylog, *fit)
plt.scatter(xnew[xok], ynew[xok], c='C3')
normalchisq = np.sum( ((ffd_ylog[xok] - np.polyval(fit, ffd_x[xok]))/ffd_yerrlog[xok])**2 ) / np.float(np.sum(xok))
dist = np.sqrt((ffd_x[xok]-xnew[xok])**2 + (ffd_ylog[xok]-ynew[xok])**2)
errs = np.sqrt((ffd_xerr[xok])**2 + (ffd_yerrlog[xok])**2)
newchisq = np.sum( (dist/errs)**2 ) / np.float(np.sum(xok))
print(normalchisq, newchisq)
# +
def _lnProb(theta, x, y, xerr, yerr):
Nmin = 10
m, b, xmin = theta
xok = x > xmin
# implement some strict "priors" here
if ((sum(xok) >= Nmin) and (-2 < m < -0.5) and (20 < b < 40) and
(np.min(x) < xmin < np.max(x))):
# orthogonal distance of points to the line
xnew, ynew = nrpt(x, y, m, b)
# the York2004 version of the chisq
# chisq = np.sum((y[xok] - model[xok])**2 /
# (yerr[xok]**2 + (m * xerr[xok])**2)) / np.float(np.sum(xok))
# chisq = np.sum(((x[xok]-xnew[xok])/xerr[xok])**2 *
# ((y[xok]-ynew[xok])/yerr[xok])**2 ) / np.float(np.sum(xok))
# orthog distance chisq
dist = np.sqrt((x[xok]-xnew[xok])**2 + (y[xok]-ynew[xok])**2)
errs = np.sqrt((xerr[xok])**2 + (yerr[xok])**2)
chisq = np.sum( (dist/errs)**2 ) / np.float(np.sum(xok))
BIC = chisq + np.size(theta) * np.log(len(x)+(np.float(len(x) - np.sum(xok))))
return -0.5 * BIC
return -np.inf
# -
def FFD_odr(ffd_x, ffd_y, ffd_xerr, ffd_yerr, Nsteps=2000):
# initialization
xok = (ffd_x >= np.mean(ffd_x))
fit = np.polyfit(ffd_x[xok], ffd_y[xok], 1)
pos = np.array([fit[0], fit[1], np.min(ffd_x)])
pos = pos + 1e-4 * np.random.randn(2**5, pos.size)
nwalkers, ndim = pos.shape
sampler = emcee.EnsembleSampler(nwalkers, ndim, _lnProb,
args=(ffd_x, ffd_y, ffd_xerr, ffd_yerr))
sampler.run_mcmc(pos, Nsteps, progress=True);
samples = sampler.get_chain()
flat_samples = sampler.get_chain(discard=int(Nsteps*0.1), thin=15, flat=True)
return flat_samples
# +
# Our errors seem too small (chisq << 0 near the best model)
# so this actually seems to work oddly well by down-weighting the errors. Huh.
fudge = 7.
flat_samplesY2 = FFD_odr(ffd_xY, ffd_yYlog, ffd_xerrY/fudge, ffd_yerrYlog/fudge, Nsteps=1000)
# +
plt.figure(figsize=(9,7))
inds = np.random.randint(len(flat_samplesY2), size=100)
for ind in inds:
sample = flat_samplesY2[ind]
plt.plot([sample[2], max(ffd_xY)], (sample[0]*np.array([sample[2], max(ffd_xY)]) + sample[1]),
"k", alpha=0.1)
plt.errorbar(ffd_xY, ffd_yYlog, xerr=ffd_xerrY,#np.sqrt(ffd_xerrY**2+LUMINY_err**2),
yerr=ffd_yerrYlog,
color='C1', alpha=0.25, linestyle='none', lw=1, marker='.')
plt.plot(ffd_xY[ffd_xY >= np.median(flat_samplesY2[:,2])],
(np.median(flat_samplesY2[:,0]) * ffd_xY[ffd_xY >= np.median(flat_samplesY2[:,2])] +
np.median(flat_samplesY2[:,1])),
c='r', lw=3)
plt.xlim(29.5,33.5)
plt.ylim(-2, 1)
plt.xlabel('log Energy (erg)')
plt.ylabel(r'log Cumulative Flare Rate (day$^{-1}$)')
print(therng(flat_samplesY2[:,0]))
print(therng(flat_samplesY2[:,1]))
print(therng(flat_samplesY2[:,2]))
# -
flat_samples2 = FFD_odr(ffd_x[:-30], ffd_ylog[:-30], ffd_xerr[:-30]/fudge, ffd_yerrlog[:-30]/fudge, Nsteps=1000)
# +
plt.figure(figsize=(9,7))
inds = np.random.randint(len(flat_samples2), size=100)
for ind in inds:
sample = flat_samples2[ind]
plt.plot([sample[2], max(ffd_x)], (sample[0]*np.array([sample[2], max(ffd_x)]) + sample[1]),
"k", alpha=0.1)
plt.errorbar(ffd_x, ffd_ylog, xerr=ffd_xerr,
yerr=ffd_yerrlog,
color='C0', alpha=0.25, linestyle='none', lw=1, marker='.')
plt.plot(ffd_x[ffd_x >= np.median(flat_samples2[:,2])],
(np.median(flat_samples2[:,0]) * ffd_x[ffd_x >= np.median(flat_samples2[:,2])] +
np.median(flat_samples2[:,1])),
c='r', lw=3)
plt.xlim(29.5,33.5)
plt.ylim(-2, 1)
plt.xlabel('log Energy (erg)')
plt.ylabel(r'log Cumulative Flare Rate (day$^{-1}$)')
print(therng(flat_samples2[:,0]))
print(therng(flat_samples2[:,1]))
print(therng(flat_samples2[:,2]))
# +
Ndraws = 50
plt.figure(figsize=(9,7))
inds = np.random.randint(len(flat_samples2), size=Ndraws)
for ind in inds:
sample = flat_samples2[ind]
plt.plot([sample[2], max(ffd_x)], (sample[0]*np.array([sample[2], max(ffd_x)]) + sample[1]),
"k", alpha=0.2)
plt.errorbar(ffd_x, ffd_ylog, xerr=ffd_xerr, yerr=ffd_yerrlog,
color='DodgerBlue', alpha=0.25, linestyle='none', lw=1, marker='.')
plt.plot(ffd_x[ffd_x >= np.median(flat_samples2[:,2])],
(np.median(flat_samples2[:,0]) * ffd_x[ffd_x >= np.median(flat_samples2[:,2])] +
np.median(flat_samples2[:,1])),
c='cyan', lw=3)
inds = np.random.randint(len(flat_samplesY2), size=Ndraws)
for ind in inds:
sample = flat_samplesY2[ind]
plt.plot([sample[2], max(ffd_xY)], (sample[0]*np.array([sample[2], max(ffd_xY)]) + sample[1]),
"k", alpha=0.2)
plt.errorbar(ffd_xY, ffd_yYlog, xerr=ffd_xerrY, yerr=ffd_yerrYlog,
color='Firebrick', alpha=0.25, linestyle='none', lw=1, marker='.')
plt.plot(ffd_xY[ffd_xY >= np.median(flat_samplesY2[:,2])],
(np.median(flat_samplesY2[:,0]) * ffd_xY[ffd_xY >= np.median(flat_samplesY2[:,2])] +
np.median(flat_samplesY2[:,1])),
c='r', lw=3)
plt.xlim(30.1,33.1)
plt.ylim(-1.7, .9)
plt.xlabel('log Energy (erg)')
plt.ylabel(r'log Cumulative Flare Rate Per Day')
plt.savefig('YZCMI_vs_GJ1243_mcmc.png', dpi=150, bbox_inches='tight', pad_inches=0.25)
# -
| yzcmi_vs_gj1243.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Now, every face will be cropped and saved
# In this cell, some libraries were imported
import cv2
import sys
import os
from PIL import Image, ImageDraw
import pylab
import time
# Face Detection Function
def detectFaces(image_name):
print ("Face Detection Start.")
# Read the image and convert to gray to reduce the data
img = cv2.imread(image_name)
gray = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)#Color => Gray
# The haarcascades classifier is used to train data
#face_cascade = cv2.CascadeClassifier("/usr/share/opencv/haarcascades/haarcascade_frontalface_default.xml")
faces = face_cascade.detectMultiScale(gray, 1.2, 5)#1.3 and 5are the min and max windows of the treatures
result = []
for (x,y,width,height) in faces:
result.append((x,y,x+width,y+height))
print ("Face Detection Complete.")
return result
#Crop faces and save them in the same directory
filepath ="/home/xilinx/jupyter_notebooks/Final_presents/images/"
dir_path ="/home/xilinx/jupyter_notebooks/Final_presents/"
filecount = len(os.listdir(filepath))-1
image_count = 1#count is the number of images
face_cascade = cv2.CascadeClassifier("/usr/share/opencv/haarcascades/haarcascade_frontalface_default.xml")
for fn in os.listdir(filepath): #fn 表示的是文件名
start = time.time()
if image_count <= filecount:
image_name = str(image_count) + '.JPG'
image_path = filepath + image_name
image_new = dir_path + image_name
#print (image_path)
#print (image_new)
os.system('cp '+(image_path)+ (' /home/xilinx/jupyter_notebooks/OpenCV/Face_Detection/'))
faces = detectFaces(image_name)
if not faces:
print ("Error to detect face")
if faces:
#All croped face images will be saved in a subdirectory
face_name = image_name.split('.')[0]
#os.mkdir(save_dir)
count = 0
for (x1,y1,x2,y2) in faces:
file_name = os.path.join(dir_path,face_name+str(count)+".jpg")
Image.open(image_name).crop((x1,y1,x2,y2)).save(file_name)
#os.system('rm -rf '+(image_path)+' /home/xilinx/jupyter_notebooks/OpenCV/Face_Detection/')
count+=1
os.system('rm -rf '+(image_new))
print("The " + str(image_count) +" image were done.")
print("Congratulation! The total of the " + str(count) + " faces in the " +str(image_count) + " image.")
end = time.time()
TimeSpan = end - start
if image_count <= filecount:
print ("The time of " + str(image_count) + " image is " +str(TimeSpan) + " s.")
image_count = image_count + 1
# Initiate ORB detector
orb = cv2.ORB_create()
img1 = cv2.imread('20.jpg',cv2.COLOR_BGR2GRAY)
img2 = cv2.imread('31.jpg',cv2.COLOR_BGR2GRAY)
# +
# find the keypoints and descriptors with SIFT for face 1
kp1, des1 = orb.detectAndCompute(img1,None)
imgOut1 = cv2.drawKeypoints(img1,kp,None, color=(255,0,0), flags=0)
plt.imshow(imgOut1),plt.show()
# find the keypoints and descriptors with SIFT for face 2
kp2, des2 = orb.detectAndCompute(img2,None)
imgOut2 = cv2.drawKeypoints(img2,kp,None, color=(255,0,0), flags=0)
plt.imshow(imgOut2),plt.show()
# +
# create BFMatcher object
bf = cv2.BFMatcher(cv2.NORM_HAMMING, crossCheck=True)
# Match descriptors.
matches = bf.match(des1,des2)
# Sort them in the order of their distance.
matches = sorted(matches, key = lambda x:x.distance)
# -
def drawMatches(img1, kp1, img2, kp2, matches):
"""
My own implementation of cv2.drawMatches as OpenCV 2.4.9
does not have this function available but it's supported in
OpenCV 3.0.0
This function takes in two images with their associated
keypoints, as well as a list of DMatch data structure (matches)
that contains which keypoints matched in which images.
An image will be produced where a montage is shown with
the first image followed by the second image beside it.
Keypoints are delineated with circles, while lines are connected
between matching keypoints.
img1,img2 - Grayscale images
kp1,kp2 - Detected list of keypoints through any of the OpenCV keypoint
detection algorithms
matches - A list of matches of corresponding keypoints through any
OpenCV keypoint matching algorithm
"""
# Create a new output image that concatenates the two images together
# (a.k.a) a montage
rows1 = img1.shape[0]
cols1 = img1.shape[1]
rows2 = img2.shape[0]
cols2 = img2.shape[1]
out = np.zeros((max([rows1,rows2]),cols1+cols2,3), dtype='uint8')
# Place the first image to the left
out[:rows1,:cols1] = np.dstack([img1, img1, img1])
# Place the next image to the right of it
out[:rows2,cols1:] = np.dstack([img2, img2, img2])
# For each pair of points we have between both images
# draw circles, then connect a line between them
for mat in matches:
# Get the matching keypoints for each of the images
img1_idx = mat.queryIdx
img2_idx = mat.trainIdx
# x - columns
# y - rows
(x1,y1) = kp1[img1_idx].pt
(x2,y2) = kp2[img2_idx].pt
# Draw a small circle at both co-ordinates
# radius 4
# colour blue
# thickness = 1
cv2.circle(out, (int(x1),int(y1)), 4, (255, 0, 0), 1)
cv2.circle(out, (int(x2)+cols1,int(y2)), 4, (255, 0, 0), 1)
# Draw a line in between the two points
# thickness = 1
# colour blue
cv2.line(out, (int(x1),int(y1)), (int(x2)+cols1,int(y2)), (255, 0, 0), 1)
return out
# +
# Draw first 10 matches.
print (len(matches))
img3 = drawMatches(img1,kp1,img2,kp2,matches[:10])
plt.imshow(img3),plt.show()
# -
| teams/team_lynx/code/.ipynb_checkpoints/CU-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# <center><img src="img/open_access.png"><center>
#
# # <center>Data<center>
# + [markdown] slideshow={"slide_type": "slide"}
# # Open data is
#
# - Available
# - the data should be
# - in whole,
# - downloadable from the Internet
# - with no costs apart from reproduction fees
# - Accessible
# - the data should be provided in a convenient form that can be modified
# - Reusable:
# - this should be expressed under terms provided with the data
# - Redistributable
# - the data can be combined with data from other research
# - Unrestricted
# - everyone can use, modify, and share the data, regardless of how they use the data (e.g. for commercial, non-commercial, or educational purposes)
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Benefits of sharing Open Access Data
# - Ease of Replication
# - Most research is funded by the public
# - Therefore the public should not have to pay to access the data they have already paid for
# - Better meta-analyses when full data are available
# - Scientific transparancy and credibility
# - If you hold a CIHR, this is part of the mandate
# - Publisher requirements
# - Increase collaborations
# - Enable future discoveries
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Open data can take many forms...
#
# <img src="img/opendatatable.png">
# + [markdown] slideshow={"slide_type": "slide"}
# # You already benefit tremendously from open data
# - GPS
# - Medical research
# - Stats Canada
# - IMDB
# - Rate My Prof
# + [markdown] slideshow={"slide_type": "slide"}
# # Ethics and open access data
#
# - Make sure you apply for ethics approval to post data online in open access repositories
# - You can always write an amendment
# - It is important to follow your ethics guidelines and anonymize data before it is posted online
# + [markdown] slideshow={"slide_type": "slide"}
# ## Where to make data available
#
# - <a href = "https://learn.scholarsportal.info/all-guides/dataverse/" target="_blank">Mac Dataverse</a>
# - <a href="https://osf.io/" target="_blank">Open Science Framework</a>
# - <a href="http://oad.simmons.edu/oadwiki/Data_repositories" target="_blank">Simmons list of open access Data Repositories</a>
# - <a href="https://data.mendeley.com/" target="_blank">Mendelay (owned by Elsevier)</a>
# - <a href="http://www.science.gc.ca/eic/site/063.nsf/eng/h_94D49094.html" target="_blank">Government of Canada Annex</a>
# + [markdown] slideshow={"slide_type": "slide"}
# # <a href="https://learn.scholarsportal.info/all-guides/dataverse/" target="_blank">McMaster Dataverse</a>
# + [markdown] slideshow={"slide_type": "fragment"}
#
# ## Dataverse is in consortium with many universities globally
# - Deposit data
# - Create metadata
# - Release and share data openly or privately
# - Visualize and explore data
# - Search for data
# - Get credit with a DOI
#
# <a href="https://dataverse.scholarsportal.info/dataset.xhtml?ownerId=41126" target="_blank">What it looks like to deposit data...</a>
#
# + [markdown] slideshow={"slide_type": "slide"}
# # <a href="https://learn.scholarsportal.info/all-guides/dataverse/" target="_blank">McMaster Dataverse</a>
# + [markdown] slideshow={"slide_type": "fragment"}
# ## What can you deposit?
# - research data
# - supplementary tables and documentation
# - publications associated with data
# - presentations
# - stimuli
#
# - Any file type
#
# - Data Explorer feature
# - e.g. Stata, SPSS, R, Excel (xlsx) and CSV
# + [markdown] slideshow={"slide_type": "slide"}
# # Best practices in making data available
# <table>
# <td style="text-align:left">
# <ul>
# <li><font size="+1">Good</li>
# <ul>
# <li><font size="+1">Post final data set</li>
# </ul>
# <li><font size="+1">Better</li>
# <ul>
# <li><font size="+1">Post raw data</li>
# </ul>
# <li>Best </li>
# <ul>
# <li><font size="+1">Post raw data</li>
# <li><font size="+1">Show how you get from raw data to analysis</li>
# <li><font size="+1">Include a codebook</li>
# <li><font size="+1">Comment all code extensively</li>
# <li><font size="+1">Post data with preprint</li>
# </ul>
# </ul>
# </td>
# <td><img src="img/good_better_best.jpg" width=70%>
# </td>
# </table>
#
# + [markdown] slideshow={"slide_type": "subslide"}
# # Try not to use proprietary software or file formats
#
# - Proprietary formats
# - can disappear when comanies decide
# - are not accessible to many in non-wealthy countries
# - do not facilitate open source practices
#
# + [markdown] slideshow={"slide_type": "subslide"}
# # Proprietary formats and alternatives
# <img src="img/formats.png">
# + [markdown] slideshow={"slide_type": "slide"}
# # Tools to facilitate open access data
#
# - <a href="https://jupyter.org/" target="_blank">Anaconds's Jupyter Notebooks</a>
# - <a href="https://www.rstudio.com/" target="_blank">RStudio Markdom (Rmd)</a>
# - <a href="https://cran.r-project.org/web/packages/codebook/index.html" target="_blank">Codebook</a>
# - <a href="https://github.com/" target="_blank">Github</a>
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Anaconda
#
# <table>
# <td style="text-align:left">
# <ul>
# <li><font size="+2"><a href="https://www.anaconda.com" target="_blank">Anaconda</a> is a scientific Python distribution</font></li>
# <li><font size="+2"><a href="https://jupyter.org/ target="_blank"">Jupyter notebooks</a> run codeblocks and markdown to create detailed explanations of data processing and analysis</font></li>
# <ul>
# <li><font size="+2">Support for JUlia, PYThon, R (JUPYTR) and other languages</font></li>
# <li><font size="+2">This presentation was written in Jupyter notebook</font></li>
# </ul>
# <li><font size="+2">Spyder is an IDE modelled after MatLab</font></li>
# <li><font size="+2">Most people prefer <a href="https://www.jetbrains.com/pycharm/" target="_blank">PyCharm</a></font></li>
# <ul>
# <li><font size="+2">Pycharm professional (free to academics) has Science Mode</font></li>
# </ul>
# </ul>
# </td>
# <td>
# <img src="img/jupyter-demo.png">
# </td>
# </table>
# + [markdown] slideshow={"slide_type": "slide"}
# <table>
# <td style="text-align:left">
# <ul>
# <li><a href="https://www.rstudio.com/" target="_blank"><font size="+2">RStudio Markdom (Rmd)</font></a></li>
# <li><font size="+2">Support for R, Python, and other languages</font></li>
# <li><font size="+2">Use codeblocks to create detailed explanations of data processing and analysis</font></li>
# </ul>
# </td>
# <td>
# <img src="img/notebook-demo.png">
# </td>
# </table>
# + [markdown] slideshow={"slide_type": "slide"}
# <h1><a href="https://rubenarslan.github.io/codebook/articles/codebook.html" target="_blank">Codebook</a></h1>
#
#
# - automate the following tasks to describe data frames:
# - Summarise the distributions, and labelled missings of variables graphically and using descriptive statistics
# - For surveys, compute and summarise reliabilities (internal consistencies, retest, multilevel) for psychological scales.
# - Combine this information with metadata (such as item labels and labelled values) that is derived from R attributes.
# - Generate HTML, PDF, and Word documents
#
# + [markdown] slideshow={"slide_type": "slide"}
# <center><a href="https://github.com/"><img src="img/github.jpg" width=25%></a></center>
# <ul>
# <li> A version tracking open access repository for code </li>
# <li> You can post datasets on Github</li>
# <li> You can host websites that explain your data and analysis:</li>
# <ul>
# <li>- e.g. <a href="https://rubenarslan.github.io/ovulatory_shifts/#table-of-contents" target="_blank">Ruben Arslan's Ovulagory Shifts site</a>
# </li>
# </ul>
# <li>You can also host open access slides and presentations, like this <a href= "https://drfeinberg.github.io/PNB-Open-Science-Summer-Workshop/05-Open-Access-Data.slides.html" target="_blank">one</a>
# </li>
# <li><a href="https://github.com/collections/open-data" target="_blank">GitHub open data collection</a></li>
# </ul>
#
# GitHub is now owned by Microsoft. The open source community has mixed feelings about this.
#
# + [markdown] slideshow={"slide_type": "slide"}
# # You don't need to be a programmer to have open access data
#
# - You can use excel
# - You can use SPSS and have it record your button presses and translate them into a script
# - You can do whatever you like
# - Any data sharing is better than none
# - Do what you are comfortable with
# - Each to their own abilities
# + [markdown] slideshow={"slide_type": "slide"}
# # Questions and discussion
#
# ---------
# Some materials from
# https://libraries.mit.edu/data-management/files/2014/05/DataSharingStorage_20170125.pdf
| 05-Open-Access-Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Comprobar velocidad de extrusión
# Vamos a comprobar que la velocidad de extrusión es constante
# Hora de inicio: 11:36
# Hora de fin: 12:06
#
# Pellet usado, 100% reciclado
# %pylab inline
#Importamos las librerías utilizadas
import numpy as np
import pandas as pd
import seaborn as sns
#Mostramos las versiones usadas de cada librerías
print ("Numpy v{}".format(np.__version__))
print ("Pandas v{}".format(pd.__version__))
print ("Seaborn v{}".format(sns.__version__))
#Abrimos el fichero csv con los datos de la muestra
datos = pd.read_csv('7211533.CSV')
#Mostramos un resumen de los datos obtenidoss
datos.describe()
# +
#Mostramos en varias gráficas la información obtenida tras el ensayo
plt.figure(figsize=(20,10))
plt.plot(datos['RPM TRAC'], label=('f(x)'))
plt.ylim(0.9,2.1)
plt.xlim(0,1200)
plt.title('Velocidad de extrusión')
plt.xlabel('Tiempo')
plt.ylabel('Velocidad tractora RPM')
# -
| medidas/21072015/modelado.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
revIncrease = int(row[1]) - prevRevenue
change = change + revIncrease
prevRevenue = int(row[1])
average_change = round(change/total_months,2)
# -
| PyBank/.ipynb_checkpoints/Untitled-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: test-env
# language: python
# name: test-env
# ---
# # Modeling and Simulation in Python
#
# Chapter 1
#
# Copyright 2020 <NAME>
#
# License: [Creative Commons Attribution 4.0 International](https://creativecommons.org/licenses/by/4.0)
# ## Jupyter
#
# Welcome to *Modeling and Simulation*, welcome to Python, and welcome to Jupyter.
#
# This is a Jupyter notebook, which is a development environment where you can write and run Python code. Each notebook is divided into cells. Each cell contains either text (like this cell) or Python code.
#
# ### Selecting and running cells
#
# To select a cell, click in the left margin next to the cell. You should see a blue frame surrounding the selected cell.
#
# To edit a code cell, click inside the cell. You should see a green frame around the selected cell, and you should see a cursor inside the cell.
#
# To edit a text cell, double-click inside the cell. Again, you should see a green frame around the selected cell, and you should see a cursor inside the cell.
#
# To run a cell, hold down SHIFT and press ENTER.
#
# * If you run a text cell, Jupyter formats the text and displays the result.
#
# * If you run a code cell, Jupyter runs the Python code in the cell and displays the result, if any.
#
# To try it out, edit this cell, change some of the text, and then press SHIFT-ENTER to format it.
# ### Adding and removing cells
#
# You can add and remove cells from a notebook using the buttons in the toolbar and the items in the menu, both of which you should see at the top of this notebook.
#
# Try the following exercises:
#
# 1. From the Insert menu select "Insert cell below" to add a cell below this one. By default, you get a code cell, as you can see in the pulldown menu that says "Code".
#
# 2. In the new cell, add a print statement like `print('Hello')`, and run it.
#
# 3. Add another cell, select the new cell, and then click on the pulldown menu that says "Code" and select "Markdown". This makes the new cell a text cell.
#
# 4. In the new cell, type some text, and then run it.
#
# 5. Use the arrow buttons in the toolbar to move cells up and down.
#
# 6. Use the cut, copy, and paste buttons to delete, add, and move cells.
#
# 7. As you make changes, Jupyter saves your notebook automatically, but if you want to make sure, you can press the save button, which looks like a floppy disk from the 1990s.
#
# 8. Finally, when you are done with a notebook, select "Close and Halt" from the File menu.
# ### Using the notebooks
#
# The notebooks for each chapter contain the code from the chapter along with additional examples, explanatory text, and exercises. I recommend you
#
# 1. Read the chapter first to understand the concepts and vocabulary,
# 2. Run the notebook to review what you learned and see it in action, and then
# 3. Attempt the exercises.
#
# If you try to work through the notebooks without reading the book, you're gonna have a bad time. The notebooks contain some explanatory text, but it is probably not enough to make sense if you have not read the book. If you are working through a notebook and you get stuck, you might want to re-read (or read!) the corresponding section of the book.
# ### Installing modules
#
# These notebooks use standard Python modules like NumPy and SciPy. I assume you already have them installed in your environment.
#
# They also use two less common modules: Pint, which provides units, and modsim, which contains code I wrote specifically for this book.
#
# The following cells check whether you have these modules already and tries to install them if you don't.
# +
# try:
# import pint
# except ImportError:
# # !pip install pint
# import pint
import pint
# +
# try:
# from modsim import *
# except ImportError:
# # !pip install modsimpy
# from modsim import *
from modsim import *
# -
# The first time you run this on a new installation of Python, it might produce a warning message in pink. That's probably ok, but if you get a message that says `modsim.py depends on Python 3.7 features`, that means you have an older version of Python, and some features in `modsim.py` won't work correctly.
#
# If you need a newer version of Python, I recommend installing Anaconda. You'll find more information in the preface of the book.
#
# You can find out what version of Python and Jupyter you have by running the following cells.
# !python --version
# !jupyter-notebook --version
# ### Configuring Jupyter
#
# The following cell:
#
# 1. Uses a Jupyter "magic command" to specify whether figures should appear in the notebook, or pop up in a new window.
#
# 2. Configures Jupyter to display some values that would otherwise be invisible.
#
# Select the following cell and press SHIFT-ENTER to run it.
# +
# Configure Jupyter so figures appear in the notebook
# %matplotlib inline
# Configure Jupyter to display the assigned value after an assignment
# %config InteractiveShell.ast_node_interactivity='last_expr_or_assign'
# -
# ## The penny myth
#
# The following cells contain code from the beginning of Chapter 1.
#
# `modsim` defines `UNITS`, which contains variables representing pretty much every unit you've ever heard of. It uses [Pint](https://pint.readthedocs.io/en/latest/), which is a Python library that provides tools for computing with units.
#
# The following lines create new variables named `meter` and `second`.
meter = UNITS.meter
second = UNITS.second
# To find out what other units are defined, type `UNITS.` (including the period) in the next cell and then press TAB. You should see a pop-up menu with a list of units.
# Create a variable named `a` and give it the value of acceleration due to gravity.
a = 9.8 * meter / second**2
# Create `t` and give it the value 4 seconds.
t = 4 * second
# Compute the distance a penny would fall after `t` seconds with constant acceleration `a`. Notice that the units of the result are correct.
a * t**2 / 2
# **Exercise**: Compute the velocity of the penny after `t` seconds. Check that the units of the result are correct.
a*t
# **Exercise**: Why would it be nonsensical to add `a` and `t`? What happens if you try?
# The error messages you get from Python are big and scary, but if you read them carefully, they contain a lot of useful information.
#
# 1. Start from the bottom and read up.
# 2. The last line usually tells you what type of error happened, and sometimes additional information.
# 3. The previous lines are a "traceback" of what was happening when the error occurred. The first section of the traceback shows the code you wrote. The following sections are often from Python libraries.
#
# In this example, you should get a `DimensionalityError`, which is defined by Pint to indicate that you have violated a rules of dimensional analysis: you cannot add quantities with different dimensions.
#
# Before you go on, you might want to delete the erroneous code so the notebook can run without errors.
# ## Falling pennies
#
# Now let's solve the falling penny problem.
#
# Set `h` to the height of the Empire State Building:
h = 381 * meter
# Compute the time it would take a penny to fall, assuming constant acceleration.
#
# $ a t^2 / 2 = h $
#
# $ t = \sqrt{2 h / a}$
t = sqrt(2 * h / a)
# Given `t`, we can compute the velocity of the penny when it lands.
#
# $v = a t$
v = a * t
# We can convert from one set of units to another like this:
mile = UNITS.mile
hour = UNITS.hour
v.to(mile/hour)
# **Exercise:** Suppose you bring a 10 foot pole to the top of the Empire State Building and use it to drop the penny from `h` plus 10 feet.
#
# Define a variable named `foot` that contains the unit `foot` provided by `UNITS`. Define a variable named `pole_height` and give it the value 10 feet.
#
# What happens if you add `h`, which is in units of meters, to `pole_height`, which is in units of feet? What happens if you write the addition the other way around?
foot = UNITS.foot
pole_height = 10*foot
pole_height + h, h + pole_height
# **Exercise:** In reality, air resistance limits the velocity of the penny. At about 18 m/s, the force of air resistance equals the force of gravity and the penny stops accelerating.
#
# As a simplification, let's assume that the acceleration of the penny is `a` until the penny reaches 18 m/s, and then 0 afterwards. What is the total time for the penny to fall 381 m?
#
# You can break this question into three parts:
#
# 1. How long until the penny reaches 18 m/s with constant acceleration `a`.
# 2. How far would the penny fall during that time?
# 3. How long to fall the remaining distance with constant velocity 18 m/s?
#
# Suggestion: Assign each intermediate result to a variable with a meaningful name. And assign units to all quantities!
mps = UNITS.meter/UNITS.second
v = 18*mps
t = v/a
d = a * t**2 / 2
d
remaining_d = 381*UNITS.meter - d
remaining_time = remaining_d/v
# ### Restart and run all
#
# When you change the contents of a cell, you have to run it again for those changes to have an effect. If you forget to do that, the results can be confusing, because the code you are looking at is not the code you ran.
#
# If you ever lose track of which cells have run, and in what order, you should go to the Kernel menu and select "Restart & Run All". Restarting the kernel means that all of your variables get deleted, and running all the cells means all of your code will run again, in the right order.
#
# **Exercise:** Select "Restart & Run All" now and confirm that it does what you want.
| notebooks/chap01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## UAP Generate & Evaluate (whitebox)
#
# attacks
# - sgd-uap (untargeted) eps = 2-12
#
# dataset
# - cifar10
# - svhn
#
# model_name
# - resnet18
# - [soft-pruning]
# - resnet18_sfP
# - resnet18_sfP-mixup
# - resnet18_sfP-cutout
# - [post-training pruning]
# - resnet18_PX_0.Y
# - X = 2, 3, 4
# - Y = 3, 6, 9
# +
import numpy as np
import pandas as pd
import os
import torch
import json
from utils_attack import uap_batch
from utils_model import *
dir_results = './results/'
dir_uap = './uaps/'
# -
nb_epoch = 10
atk_name = 'sgd-uap'
y_target = -1
model_names = ['resnet18', 'resnet18_sfP', 'resnet18_sfP-mixup', 'resnet18_sfP-cutout']
for layer in [2, 3, 4]:
for prune_pct in ['0.3', '0.6', '0.9']:
model_names.append('resnet18_P%i-%s' % (layer, prune_pct))
model_names.pop(6)
model_names.pop(8)
model_names
atk_sources = model_names.copy()
for Q_param in [2, 3, 4]:
atk_sources.append('resnet20_Q%i' % Q_param)
atk_sources
def eval_uap(uap):
top, top_probs, top1acc, top5acc, outputs, labels = evaluate(model, testloader, uap = uap)
print('Top 1 accuracy', sum(top1acc) / len(labels))
print('Top 5 accuracy', sum(top5acc) / len(labels))
dict_entry = {}
dict_entry['model_name'] = model_name
dict_entry['dataset'] = dataset
dict_entry['atk_source'] = atk_source
dict_entry['atk_name'] = atk_name
dict_entry['atk_param'] = y_target
dict_entry['eps'] = eps
if atk_name[:3] == 'sgd': dict_entry['nb_epoch'] = nb_epoch
dict_entry['top1acc'] = sum(top1acc) / len(labels)
dict_entry['top5acc'] = sum(top5acc) / len(labels)
dict_entry['UER'] = 1 - sum(top1acc) / len(labels)
print('UER ', dict_entry['UER'])
print('\n')
if y_target >= 0: dict_entry['tgt_success'] = sum(outputs == y_target) / len(labels)
# Output distribution
for i in range(10):
dict_entry['label_dist%i' % i] = sum(outputs == i) / len(labels)
all_results.append(dict_entry)
# + active=""
# for dataset in ['cifar10', 'svhn']:
# all_results = []
#
# testloader = get_testdata(dataset)
#
# for model_name in model_names:
# model = get_model(model_name, dataset)
# model.eval()
#
# for atk_source in atk_sources:
# uap_load_path = dir_uap + dataset + '/' + atk_source
#
# for eps in range(2, 13, 2):
# uap_path = uap_load_path + '/sgd-eps%i.pth' % eps
# print(uap_path)
# uap = torch.load(uap_path)
# eval_uap(uap)
#
# pd.DataFrame(all_results).to_csv(dir_results + 'eval_untgt-transfer-%s.csv' % dataset)
#
# pd.DataFrame(all_results).to_csv(dir_results + 'eval_untgt-transfer-%s.csv' % dataset)
# -
Q_param = 3
for dataset in ['cifar10', 'svhn']:
all_results = []
testloader = get_testdata(dataset)
for model_name in ['resnet20_Q%i' % Q_param]:
model = get_model(model_name, dataset)
model.eval()
for atk_source in atk_sources:
uap_load_path = dir_uap + dataset + '/' + atk_source
for eps in range(2, 13, 2):
uap_path = uap_load_path + '/sgd-eps%i.pth' % eps
print(uap_path)
uap = torch.load(uap_path)
eval_uap(uap)
pd.DataFrame(all_results).to_csv(dir_results + 'evalQ%i_untgt-transfer-%s.csv' % (Q_param, dataset))
pd.DataFrame(all_results).to_csv(dir_results + 'evalQ%i_untgt-transfer-%s.csv' % (Q_param, dataset))
| 11-03 eval_untgt-transfer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Bell and CHSH inequalities
# $\newcommand{\bra}[1]{\left\langle{#1}\right|}$
# $\newcommand{\ket}[1]{\left|{#1}\right\rangle}$
#
# The purpose of this notebook is to simulate the CHSH experiment
# described in the IBM Quantum Experience tutorial in the section
# entitled
#
# >Multiple Qubits, Gates, and Entangled States/Entanglement and Bell Tests
# First change your working directory to the qubiter directory in your computer, and add its path to the path environment variable.
import os
import sys
print(os.getcwd())
os.chdir('../../')
print(os.getcwd())
sys.path.insert(0,os.getcwd())
from qubiter.SEO_writer import *
from qubiter.SEO_simulator import *
from qubiter.StateVec import *
import numpy as np
# First, we define matrices $S, T, H, \sigma_X, \sigma_Y, \sigma_Z, I_2$.
# We will denote the Pauli matrices by $\sigma_X, \sigma_Y, \sigma_Z$
#
# Recal that
#
# $S = diag(1, i) = e^{i \frac{\pi}{4}} e^{-i \frac{\pi}{4} \sigma_Z}$
#
# $T = diag(1, e^{i \frac{\pi}{4}}) = e^{i \frac{\pi}{8}} e^{-i \frac{\pi}{8} \sigma_Z}$
#
# $H = \frac{1}{\sqrt{2}}(\sigma_Z + \sigma_X)$
#
# $H^2 = 1$
#
# $H\sigma_X H = \sigma_Z$
#
smat = np.matrix([[1, 0], [0, 1j]])
tmat = np.matrix([[1, 0], [0, np.exp(1j*np.pi/4)]])
had = np.matrix([[1, 1], [1, -1]])/np.sqrt(2)
sigx = np.matrix([[0, 1], [1, 0]])
sigy = np.matrix([[0, -1j], [1j, 0]])
sigz = np.matrix([[1, 0], [0, -1]])
id2 = np.matrix([[1, 0], [0, 1]])
# Define $\sigma_{n} = \hat{n}\cdot\vec{\sigma}$
# for any 3dim unit vector $\hat{n}$.
#
# Recall that
#
# $\sigma_Z\ket{0_Z} = \ket{0_Z}$
#
# $\sigma_Z\ket{1_Z} = -\ket{1_Z}$,
#
# or, more succinctly,
#
# $\sigma_Z\ket{b_Z} = (-1)^b\ket{b_Z}$
#
# for $b=0,1$.
#
# Likewise,
#
# $\sigma_n\ket{b_n} = (-1)^b\ket{b_n}$
#
# for any 3dim unit vector $\hat{n}$ and $b=0, 1$.
#
# One can show by Taylor expansion that
#
# $e^{i\theta \sigma_n} = \cos(\theta) + i\sigma_n \sin(\theta)$
def exp_mat2(theta, vec4):
# vec4 is 4 dimensional np.array. Zero component not used.
unit_vec = np.array([0, vec4[1], vec4[2], vec4[3]])
unit_vec = unit_vec/np.linalg.norm(unit_vec)
mat = unit_vec[1]*sigx + unit_vec[2]*sigy + unit_vec[3]*sigz
return np.cos(theta)*id2 + 1j*mat*np.sin(theta)
# Define
#
# $roty = e^{i \frac{\pi}{8}\sigma_Y}$
#
# $\hat{w} = \frac{1}{\sqrt{2}}(\hat{x} + \hat{z})$
#
# $\hat{v} = \frac{1}{\sqrt{2}}(-\hat{x} + \hat{z})$
#
# $sigw = \sigma_W = \frac{1}{\sqrt{2}}(\sigma_X + \sigma_Z)$
#
# $sigv = \sigma_V = \frac{1}{\sqrt{2}}(-\sigma_X + \sigma_Z)$
#
roty = exp_mat2(np.pi/8, np.array([0, 0, 1, 0]))
sigw = (sigx + sigz)/np.sqrt(2)
sigv = (-sigx + sigz)/np.sqrt(2)
# Check that
#
# $\sigma_W = e^{-i \frac{\pi}{8}\sigma_Y}\sigma_Z e^{i \frac{\pi}{8}\sigma_Y}$
#
# $\sigma_V = e^{i \frac{\pi}{8}\sigma_Y}\sigma_Z e^{-i \frac{\pi}{8}\sigma_Y}$
print(np.linalg.norm(sigw - roty.getH()*sigz*roty))
print(np.linalg.norm(sigv - roty*sigz*roty.getH()))
# Check that
#
# $ e^{i \frac{\pi}{8}\sigma_Y} = e^{-i \frac{\pi}{8}} S^\dagger H T H S$
roty1 = np.exp(-1j*np.pi/8)*smat.getH()*had*tmat*had*smat
print(np.linalg.norm(roty - roty1))
# Therefore, (Note that $S$ and $\sigma_Z$ are both diagonal so they commute)
#
# $\sigma_W = (S^\dagger H T^\dagger H S) \sigma_Z (S^\dagger H T H S)=
# (S^\dagger H T^\dagger H ) \sigma_Z ( H T H S)$
#
# $\sigma_V =
# (S^\dagger H T H ) \sigma_Z ( H T^\dagger H S)$
#
# Note that
#
# $\sigma_Z =\ket{0_Z}\bra{0_Z} - \ket{1_Z}\bra{1_Z} $
#
# so the same is true if we replace the $Z$ by $W$ or $V$ or any 3dim unit vector.
#
# Therefore,
# a W measurement $\bra{b_W}$ is related to a Z measurment $\bra{ b_Z}$ by
#
# $\bra{ b_W} = \bra{ b_Z} H T H S$
#
# Likewise,
#
# $\bra{ b_V} = \bra{ b_Z} H T^\dagger H S$
#
# for $b= 0, 1$
# Note that
#
# $\bra{\psi} \sigma_A(0) \sigma_B(1)\ket{\psi}
# =\bra{\psi}
# \begin{array}{c}
# (\ket{0_A}\bra{0_A} - \ket{1_A}\bra{1_A} )(0)
# \\
# (\ket{0_B}\bra{0_B} - \ket{1_B}\bra{1_B})(1)
# \end{array}
# \ket{\psi}$
#
# so
#
# $\bra{\psi} \sigma_A(0) \sigma_B(1)\ket{\psi}
# = Prob(0, 0) + Prob(1, 1) - Prob(0, 1) - Prob(1, 0)$
def write_bell_plus(file_prefix, bell_only=True, extra_had=False, t_herm=False):
num_qbits = 2
z_axis = 3
emb = CktEmbedder(num_qbits, num_qbits)
print('-------------------', file_prefix)
wr = SEO_writer(file_prefix, emb)
wr.write_one_qbit_gate(0, OneQubitGate.had2)
control_pos = 0
target_pos = 1
trols = Controls.new_single_trol(num_qbits, control_pos, kind=True)
wr.write_controlled_one_qbit_gate(
target_pos, trols, OneQubitGate.sigx)
if not bell_only:
if extra_had:
wr.write_one_qbit_gate(0, OneQubitGate.had2) # H(0)
wr.write_one_qbit_gate(1, OneQubitGate.rot_ax, [-np.pi/4, z_axis]) # S(1)
wr.write_one_qbit_gate(1, OneQubitGate.had2) # H(1)
if t_herm:
pm_one = -1
else:
pm_one = 1
wr.write_one_qbit_gate(1, OneQubitGate.rot_ax,
[-pm_one*np.pi/8, z_axis]) # T(1) if pm_one=1
wr.write_one_qbit_gate(1, OneQubitGate.had2) # H(1)
wr.close_files()
wr.print_pic_file(jup=True)
init_st_vec = StateVec.get_standard_basis_st_vec([0, 0])
sim = SEO_simulator(file_prefix, num_qbits, init_st_vec)
StateVec.describe_st_vec_dict(sim.cur_st_vec_dict, print_st_vec=True, do_pp=True,
omit_zero_amps=True, show_pp_probs=True)
fin_st_vec = sim.cur_st_vec_dict["pure"]
print('Prob(bit0=j, bit1=k) for j,k=0,1:')
prob_arr = np.abs(fin_st_vec.arr)**2
print(prob_arr)
mean = prob_arr[0, 0] \
+ prob_arr[1, 1] \
- prob_arr[0, 1] \
- prob_arr[1, 0]
print('mean=', mean)
return mean
# sigz(0)sigz(1) measurement
file_prefix = 'bell_zz_meas'
mean_zz = write_bell_plus(file_prefix, bell_only=True)
# sigz(0)sigw(1) measurement
file_prefix = 'bell_zw_meas'
mean_zw = write_bell_plus(file_prefix, bell_only=False, extra_had=False, t_herm=False)
# sigz(0)sigv(1) measurement
file_prefix = 'bell_zv_meas'
mean_zv = write_bell_plus(file_prefix, bell_only=False, extra_had=False, t_herm=True)
# sigx(0)sigw(1) measurement
file_prefix = 'bell_xw_meas'
mean_xw = write_bell_plus(file_prefix, bell_only=False, extra_had=True, t_herm=False)
# sigx(0)sigv(1) measurement
file_prefix = 'bell_xv_meas'
mean_xv = write_bell_plus(file_prefix, bell_only=False, extra_had=True, t_herm=True)
# Let
#
# $mean\_ab = \bra{\psi} \sigma_A(0) \sigma_B(1)\ket{\psi}$
#
# where
#
# $\ket{\psi} = \frac{1}{\sqrt{2}}(\ket{00}+ \ket{11})$
#
# Ckeck that
#
# $C = mean\_zw + mean\_zv + mean\_xw - mean\_xv = 2\sqrt{2}$
#
# The classical analogue of $C$ satisfies $|C| \leq 2$
print('-----------------------')
print('mean_zw + mean_zv + mean_xw - mean_xv - 2*np.sqrt(2)=',
mean_zw + mean_zv + mean_xw - mean_xv - 2*np.sqrt(2))
| qubiter/jupyter_notebooks/Bell_and_CHSH_inequalities.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Circuitos magnéticamente acoplados
# __UNLZ - Facultad de Ingeniería__
# __Electrotecnia__
# __Alumno:__ <NAME>
# <a href="https://colab.research.google.com/github/daniel-lorenzo/Electrotecnia/blob/master/Circuitos_magneticamente_acoplados.ipynb"><img align="left" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab" title="Open and Execute in Google Colaboratory"></a>
# 1. Se dice que dos bobinas están acopladas mutuamente si el flujo magnético $\phi$ que emana de una de ellas pasa por la otra. La inductancia mutua entre las dos bobinas está dada por
# $$ M = k \sqrt{L_1 L_2} $$
# donde $k$ es el coeficiente de acoplamiento $0<k<1$.
# <div class="alert-success">La <strong>inductancia mutua</strong> es la capacidad de un inductor de inducir una tensión en un inductor cercano, medida en henrys (H).
# </div>
# 2. Si $v_1$ e $i_1$ son la tensión y la corriente en la bobina 1, mientras que $v_2$ e $i_2$ son la tensión y la corriente en la bobina 2, entonces
# $$ v_1 = L_1 \frac{di_1}{dt} + M \frac{di_2}{dt} \qquad \mbox{y} \qquad v_2 = L_2 \frac{di_2}{dt} + M \frac{di_1}{dt} $$
# Así, la tensión inducida en una bobina acoplada consta de la tensión autoinducida y la tensión mutua.
# 3. La polaridad de la tensión inducida mutuamente se expresa en diagramas mediante la convención de las marcas de polaridad
# 4. La energía almacenada en las dos bobinas acopladas es
# $$ \frac{1}{2} L_1 i_1^2 + \frac{1}{2} L_2 i_2^2 \pm Mi_1i_2 $$
# 5. Un transformador es un dispositivo de cuatro terminales que contiene dos o más bobinas acopladas magnéticamente. Se emplea para modificar el nivel de corriente, tensión o impedancia en un circuito.
# 6. Las bobinas de un transformador lineal (o acoplado con holgura) están devanadas magéticamente en un material lineal. Este transformador puede reemplazarse por una red T o $\Pi$ equivalente para efectos de análisis.
# 7. Un transformador ideal (o con núcleo de hierro) es un transformador sin pérdidas $(R_1=R_2=R_3)=0$ con coeficiente de acoplamiento unitario $(k=1)$ e inductancias infinitas $(L_1, \, L_2, \,M \rightarrow \infty)$.
# 8. En un transformador ideal
# $$ V_2 = n V_1 \qquad I_2 = \frac{I_1}{n} \qquad S_1 = S_2 \qquad Z_R = \frac{Z_L}{n^2} $$
# donde $n=N_2/N_1$, es la relación de vueltas. $N_1$ es el número de vueltas del devanado primario y $N_2$ el número de vueltas del devanado secundario. El transformador aumenta la tensión primaria cuando $n>1$, la reduce cuando $n<1$ o sirve como dispositivo acoplador cuando $n=1$.
# 9. Un autotransformador es un transformador con un mismo devanado común a los circuitos primario y secundario.
# 10. Los transformadores son necesarios en todas las etapas de los sistemas de distribución de potencia. Las tensiones trifásicas pueden aumentarse o reducirse mediante transformadores trifásicos.
# 11. Usos importantes de los transformadores en aplicaciones electrónicas son como dispositivos de aislamiento eléctrico y como dispositivos de acoplamiento de impedancias.
# ## Ejemplo 13.1
# Calcule las corrientes fasoriales $I_1$ e $I_2$ del circuito de la figura
# <img src="img/ej13-1.png">
# ### Solución
# En relación con la bobina 1, la LTK da como resultado
# $$ -12 + (-j4 + j5)I_1 - j3 I_2 = 0 $$
# o sea
# $$ jI_1 - j3I_2 = 12 \tag{1} $$
# En la bobina 2, la LTK da por resultado
# $$ -j3I_1 + (12 + j6)I_2 = 0 $$
# o sea
# $$ I_1 = \frac{(12 + j6)I_2}{j3} = (2 - j4)I_2 \tag{2} $$
# Al sustituir (2) en (1) se obtiene
# $$ j(2 - j4)I_2 - j3I_2 = 12 $$
# $$ (j2 + 4)I_2 - j3I_2 = 12 $$
# $$ I_2(4 - j) = 12 $$
# $$ I_2 = \frac{12}{4-j} = 2,91 \angle 14,04^\circ \, \mathrm{A} \tag{3} $$
# Con base en las ecuaciones (2) y (3)
# $$ I_1 = (2-j4)I_2 = (4,472 \angle -63,43^\circ) \cdot (2,91 \angle 14,04^\circ) $$
# $$ I_1 = 13,02 \angle -49,40^\circ \, \mathrm{A} $$
# ### Usando sympy
import sympy as sp
import math, cmath
I1, I2 = sp.symbols('I1 I2')
Eq1 = sp.Eq(-12 + (-4j + 5j)*I1 - 3j*I2 , 0)
Eq2 = sp.Eq(-3j*I1 + (12 + 6j)*I2 , 0)
Sol = sp.solve([Eq1,Eq2],[I1,I2])
# + jupyter={"source_hidden": true}
print('I1 = (%.2f < %.2f°) A'%( abs(Sol[I1]) , math.degrees( cmath.phase(Sol[I1] ) ) ) )
print('I2 = (%.2f < %.2f°) A'%( abs(Sol[I2]) , math.degrees( cmath.phase(Sol[I2] ) ) ) )
# -
# ### Por regla de Cramer, (usando numpy)
# $$\left[
# \begin{array}{cc}
# 1j & -3j \\
# -3j & 12+6j
# \end{array}
# \right]
# \left[
# \begin{array}{c}
# I_1 \\
# I_2
# \end{array}
# \right]
# =
# \left[
# \begin{array}{c}
# 12 \\
# 0
# \end{array}
# \right]$$
import numpy as np
M = np.array([[1j , -3j],[-3j , 12+6j] ])
M1 = np.array([[12 , -3j],[0 , 12+6j] ])
M2 = np.array([[1j , 12],[-3j , 0] ])
Delta = np.linalg.det(M)
Delta1 = np.linalg.det(M1)
Delta2 = np.linalg.det(M2)
I1 = Delta1/Delta
I2 = Delta2/Delta
# $$\Delta = \left|
# \begin{array}{cc}
# 1j & -3j \\
# -3j & 12+6j
# \end{array}
# \right| $$
print('Delta = {:.2f}'.format(Delta))
# $$\Delta_1 = \left|
# \begin{array}{cc}
# 12 & -3j \\
# 0 & 12+6j
# \end{array}
# \right| $$
print('Delta1 = {:.2f}'.format(Delta1))
# $$\Delta_2 = \left|
# \begin{array}{cc}
# 1j & 12 \\
# -3j & 0
# \end{array}
# \right| $$
print('Delta2 = {:.2f}'.format(Delta2))
# $$ I_1 = \frac{\Delta_1}{\Delta} \qquad ; \qquad I_2 = \frac{\Delta_2}{\Delta} $$
# + jupyter={"source_hidden": true}
print('I1 = (%.2f < %.2f°) A'%(abs(I1) , math.degrees( cmath.phase(I1) )))
print('I2 = (%.2f < %.2f°) A'%(abs(I2) , math.degrees( cmath.phase(I2) )))
# -
# ### Otro método
# $$\left[
# \begin{array}{cc}
# 1j & -3j \\
# -3j & 12+6j
# \end{array}
# \right]
# \left[
# \begin{array}{c}
# I_1 \\
# I_2
# \end{array}
# \right]
# =
# \left[
# \begin{array}{c}
# 12 \\
# 0
# \end{array}
# \right]$$
# $$ A I = B $$
# Entonces
# $$ I = A^{-1} B $$
Ap = np.linalg.inv(M)
B = np.array([[12] , [0]])
I = np.dot(Ap , B)
# + jupyter={"source_hidden": true}
print('I1 = (%.2f < %.2f) A'%(abs(I[0]) , math.degrees( cmath.phase(I[0]) ) ))
print('I2 = (%.2f < %.2f) A'%(abs(I[1]) , math.degrees( cmath.phase(I[1]) ) ))
# -
# %reset -s -f
# ## Problema de práctica 13.1
# Determine la tensión $V_0$ en el circuito de la figura.
# <img src="img/ejp13-1.png">
# ### Solución
import numpy as np
import math, cmath
# Datos:
Vs = cmath.rect(100 , np.deg2rad(45) ) # V
A = np.array([ [4+8j , -1j],[-1j , 10+5j] ])
B = np.array([ [Vs],[0] ])
I = np.dot(np.linalg.inv(A) , B)
# + jupyter={"source_hidden": true}
print('I1 = (%.2f < %.2f) A'%(abs(I[0]) , np.rad2deg( cmath.phase(I[0]) ) ))
print('I2 = (%.2f < %.2f) A'%(abs(I[1]) , np.rad2deg( cmath.phase(I[1]) ) ))
# -
# $$ V = IR $$
I1 = I[0] ; I2 = I[1]
R2 = 10 # Ohm
Vo = -(I2*R2) # (caida de tensión)
# + jupyter={"source_hidden": true}
print('Vo = (%.2f < %.2f° V)'%(abs(Vo) , np.rad2deg( cmath.phase(Vo) ) ))
# -
A1 = np.array([ [Vs , -1j],[0 , 10+5j] ])
A2 = np.array([ [4+8j , Vs],[-1j , 0] ])
Delta = np.linalg.det(A)
Delta1 = np.linalg.det(A1)
Delta2 = np.linalg.det(A2)
I1 = Delta1/Delta
I2 = Delta2/Delta
# + jupyter={"source_hidden": true}
print('I1 = (%.2f < %.2f) A'%(abs(I1) , np.rad2deg( cmath.phase(I1) ) ))
print('I2 = (%.2f < %.2f) A'%(abs(I2) , np.rad2deg( cmath.phase(I2) ) ))
# -
Vo = -(I2*R2) # (caida de tensión)
# + jupyter={"source_hidden": true}
print('Vo = (%.2f < %.2f° V)'%(abs(Vo) , np.rad2deg( cmath.phase(Vo) ) ))
# -
# %reset -s -f
# ## Ejemplo 13.2
# Calcule las corrientes de malla en el circuito de la figura.
# <img src="img/ej13-2.png">
# ### Solución
# La clave para analizar un circuito magnéticamente acoplado es conocer la polaridad de la tensión mutua. Se debe aplicar la regla del punto. En la figura, supóngase que la bobina 1 es aquella cuya reactancia es de $6 \, \Omega$, la bobina 2 aquella cuya reactancia es de $8 \, \Omega$. Para deducir la polaridad de la tensión mutua en la bobina 1 debida a la corriente $I_2$, se observa que $I_2$ sale de la terminal marcada de la bobina 2. Puesto que se está aplicando la LTK en el sentido de las manecillas del reloj, esto implica que la tensión mutua es negativa, es decir $-j2I_2$.
# Así, en cuanto al lazo 1 de la figura, la LTK da como resultado
# $$ -100 + I_1 (4 - j3 + j6) - j6I_2 - j2 I_2 = 0 $$
# o
# $$ (4+j3)I_1 - j8I_2 = 100 \tag{1} $$
# En consecuencia, en relación con la malla 2 de la figura, la LTK produce
# $$ -2jI_1 -j6I_1 + (j6 + j8 + j2 \times 2 + 5)I_2 = 0 $$
# o
# $$ -j8I_1 + (5 + j18)I_2 = 0 \tag{2} $$
# Al colocar las ecuaciones (1) y (2) en forma matricial se obtiene
# $$\left[
# \begin{array}{cc}
# 4+j3 & -j8 \\
# -j8 & 5+j18
# \end{array}
# \right]
# \left[
# \begin{array}{c}
# I_1 \\
# I_2
# \end{array}
# \right]
# =
# \left[
# \begin{array}{c}
# 100 \\
# 0
# \end{array}
# \right]$$
# Alternativamnete, podría ser mejor deducir la tensión mutua redibujando la porción pertinente del circuito, como se muestra en la fig., donde resulta claro que la tensión mutua es $V_1 = -2jI_2$.
# <img src="img/ej13-2-2.png">
# Los determinantes son
# $$\begin{align}
# \Delta &= \left|
# \begin{array}{cc}
# 4+j3 & -j8 \\
# -j8 & 5+j18
# \end{array}
# \right| = 30+j87 \\
# \Delta_1 &= \left|
# \begin{array}{cc}
# 100 & -j8 \\
# 0 & 5+j18
# \end{array}
# \right| = 500+j1800 \\
# \Delta_2 &= \left|
# \begin{array}{cc}
# 4+j3 & 100 \\
# -j8 & 0
# \end{array}
# \right| = 0+j800
# \end{align}$$
import numpy as np
# +
M = np.array([ [4+3j , -8j],[-8j , 5+18j] ])
M1 = np.array([ [100 , -8j],[0 , 5+18j] ])
M2 = np.array([ [4+3j , 100],[-8j , 0] ])
Delta = np.linalg.det(M)
Delta1 = np.linalg.det(M1)
Delta2 = np.linalg.det(M2)
# -
print('Delta = {:.0f}'.format(Delta))
print('Delta1 = {:.0f}'.format(Delta1))
print('Delta2 = {:.0f}'.format(Delta2))
# $$ I_1 = \frac{\Delta_1}{\Delta} = 20,30 \angle 3,50^\circ \, \mathrm{A} $$
# $$ I_2 = \frac{\Delta_2}{\Delta} = 8,693 \angle 19,03^\circ \, \mathrm{A} $$
I1 = Delta1/Delta
I2 = Delta2/Delta
# + jupyter={"source_hidden": true}
print('I1 = (%.2f < %.2f°) A'%(abs(I1) , np.rad2deg( np.angle(I1) ) ))
print('I2 = (%.3f < %.2f°) A'%(abs(I2) , np.rad2deg( np.angle(I2) ) ))
# -
# %reset -s -f
# ## Ejemplo 13.3
# Considere el circuito de la figura. Determine el coeficiente de acoplamiento. Calcule la energía almacenada en los inductores acoplados en el momento $t=1 \, s$ si $v = 60 \cos (4t + 30^\circ) \, \mathrm{V}$
# <img src="img/ej13-3.png">
# ### Solución
# El coeficiente de acoplamiento es
# $$ k = \frac{M}{\sqrt{L_1 L_2}} = \frac{2,5}{\sqrt{5 \cdot 4}} = 0,559 $$
# lo que indica que los inductores están acoplados estrechamente. Para hallar la energía almacenada, se debe calcular la corriente. Para encontrar la corriente, debe obtenerse el equivalente del circuito en el dominio de la frecuencia.
# $$\begin{array}{rcl}
# 60 \cos (4t + 30^\circ) & \Rightarrow & 60 \angle 30^\circ, \quad \omega = 4 \, \mathrm{rad/s} \\
# 5 \, \mathrm{H} & \Rightarrow & j \omega L_1 = j20 \, \Omega \\
# 2,5 \, \mathrm{H} & \Rightarrow & j \omega M = j10 \, \Omega \\
# 4 \, \mathrm{H} & \Rightarrow & j \omega L_2 = j16 \, \Omega \\
# \displaystyle \frac{1}{16} \, \mathrm{F} & \Rightarrow & \displaystyle \frac{1}{j \omega C} = -j4 \, \Omega
# \end{array}$$
# Ahora se aplica el análisis de mallas. En cuanto al lazo 1,
# $$ (10 + j20) I_1 + j10 I_2 = 60 \angle 30^\circ $$
# En cuanto al lazo 2
# $$ j10 I_1 + (j16 - j4)I_2 = 0 $$
# o sea
# $$ j10I_1 + j12 I_2 = 0 $$
import cmath
import numpy as np
# Datos
w = 4 # rad/s
R1 = 10 # Ohm
L1 = 5 # H
L2 = 4 # H
M = 2.5 # H
C1 = 1/16 # F
Vs = cmath.rect( 60 , np.deg2rad(30) )
XL1 = complex(0 , w*L1)
XL2 = complex(0 , w*L2)
XC1 = complex(0 , -1/(w*C1) )
Xm = complex(0 , w*M)
A = np.array([[R1+XL1 , Xm],[Xm , XL2 + XC1]])
B = np.array([ [Vs] , [0] ])
I = np.dot( np.linalg.inv(A) , B )
print('I1 = (%.3f < %.2f°) A'%(abs(I[0]) , np.rad2deg( np.angle(I[0]) ) ) )
print('I2 = (%.3f < %.2f°) A'%(abs(I[1]) , np.rad2deg( np.angle(I[1]) ) ) )
# __Simulación en qucs:__
#
# <img src="img/ej13-3-2.png">
#
# <center><a href="qucs/ej13-3.sch">Descargar archivo qucs</a></center>
# En el dominio temporal
# $\begin{array}{l}
# i_1 = 3,905 \cos (4t - 19,4^\circ) \\
# i_2 = 3,254 \cos (4t + 160,6^\circ)
# \end{array}$
# En el momento $t = 1 \, s \quad \rightarrow \quad 4t = 4 \, \mathrm{rad} = 229,2^\circ$
# +
def i1(t):
return abs(I[0])*np.cos(4*t + np.angle(I[0]))
def i2(t):
return abs(I[1])*np.cos(4*t + np.angle(I[1]))
# -
time = 1 # s
print('i1 = %.3f A'%i1(time))
print('i2 = %.3f A'%i2(time))
# La energía total almacenada en los dos inductores acoplados es
# $$ \omega = \frac{1}{2} L_1 i_1^2 + \frac{1}{2} L_2 i_2^2 + M i_1 i_2 $$
W = 1/2 * L1 * i1(time)**2 + 1/2 * L2 * i2(time)**2 + M * i1(time) * i2(time)
print('W = %.2f J'%W)
# %reset -s -f
# ## Problema de práctica 13.3
# En referencia al circuito de la figura, determine el coeficiente de acoplamiento y la energía almacenada en los inductores acoplados en $t = 1,5 \, \mathrm{s}$.
# <img src="img/ejp13-3.png">
# ### Solución
# El coeficiente de acoplamiento es
# $$ k = \frac{M}{\sqrt{L_1 L_2}} = \frac{1}{\sqrt{2 \cdot 1}} = 0,707 $$
import cmath
import numpy as np
k = 1/np.sqrt(2*1)
print('k = %.4f'%k)
# En este caso $k>0,5$, entonces se dice que las bobinas están acopladas estrechamente.
# Para encontrar la corriente, debe obtenerse el equivalente del circuito en el dominio de la frecuencia.
# $$\begin{array}{rcl}
# 40 \cos 2t & \Rightarrow & 40 \angle 0^\circ , \quad \omega = 2 \, \mathrm{rad/s} \\
# 2 \, \mathrm{H} & \Rightarrow & j \omega L_1 = j4 \, \Omega \\
# 1 \, \mathrm{H} & \Rightarrow & j \omega M = j2 \, \Omega\\
# 1 \, \mathrm{H} & \Rightarrow & j \omega L_2 = j2 \, \Omega\\
# \displaystyle \frac{1}{8} \, \mathrm{F} & \Rightarrow & \displaystyle \frac{1}{j \omega C} = -j4 \, \Omega
# \end{array}$$
# Datos:
Vs = 40 # V
w = 2 # rad/s
R1 = 4 # Ohm
R2 = 2 # Ohm
L1 = 2 # H
L2 = 1 # H
M = 1 # H
C1 = 1/8 # F
XL1 = complex(0 , w*L1)
XL2 = complex(0 , w*L2)
XM = complex(0 , w*M)
XC1 = complex(0 , -1/(w*C1))
# + jupyter={"source_hidden": true}
print('XL1 = {:.1f} Ohm'.format(XL1))
print('XL2 = {:.1f} Ohm'.format(XL2))
print('XM = {:.1f} Ohm'.format(XM))
print('XC1 = {:.1f} Ohm'.format(XC1))
# -
A = np.array([ [R1 + XC1 + XL1 , XM], [XM , XL2 + R2 ] ])
B = np.array([ [Vs],[0] ])
I = np.dot(np.linalg.inv(A) , B)
# + jupyter={"source_hidden": true}
print('I1 = (%.3f < %.2f°) A'%(abs(I[0]) , np.rad2deg( cmath.phase(I[0]) ) ))
print('I2 = (%.3f < %.2f°) A'%(abs(I[1]) , np.rad2deg( cmath.phase(I[1]) ) ))
# -
# __Simulación en qucs:__
#
# <img src="img/ejp13-3-2.png">
#
#
# <center><a href="qucs/ejp13-3.sch">Descargar archivo qucs</a></center>
# En el dominio temporal
#
# $\begin{array}{l}
# i_1 = 7,845 \cos (2t + 11,31^\circ) \\
# i_2 = 5,547 \cos (2t - 123,69^\circ)
# \end{array}$
# +
def i1(t):
return abs(I[0])*np.cos(w*t + np.angle(I[0]))
def i2(t):
return abs(I[1])*np.cos(w*t + np.angle(I[1]))
# -
# En el momento $t = 1,5 \, s$
time = 1.5 # s
print('i1(1.5s) = %.3f A'%i1(time))
print('i2(1.5s) = %.3f A'%i2(time))
# La energía total almacenada en los dos inductores acoplados es
# $$ W = \frac{1}{2} L_1 i_1^2 + \frac{1}{2} L_2^2 i_2^2 + M i_1 i_2 $$
W = (1/2)*L1*i1(time)**2 + (1/2)*L2*i2(time)**2 + M*i1(time) * i2(time)
print('W = %.2f J'%W)
# %reset -s -f
# <a href="https://colab.research.google.com/github/daniel-lorenzo/Electrotecnia/blob/master/Circuitos_magneticamente_acoplados.ipynb"><img align="left" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab" title="Open and Execute in Google Colaboratory"></a>
| Circuitos_magneticamente_acoplados.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a id='8'></a>
# # Описание проекта "Разведывательный анализ данных".
#
# ## Цель проекта.
#
# Целью проекта является проведение разведывательного анализа данных предложенного набора данных для его подготовки к обучению и тестированию ML-модели.
#
# ## Задачи проекта.
#
# Для достижения поставленной цели необходимым видится решение следующих задач:
#
# <a href='#1'>1. Первичный осмотр данных.
#
# <a href='#2'>2. Оценка распределения значений для количественных признаков, поиск и устранение ошибок/аномальных значений.
#
# <a href='#3'>3. Корреляционный анализ количественных признаков.
#
# <a href='#4'>4. Визуальный анализ номинативных признаков, оценка количества уникальных значений.
#
# <a href='#5'>5. t-test номинативных и смешанных переменных.
#
# <a href='#6'>6. Заполнение пропусков.
#
# <a href='#7'>7. Итоговый вывод.
# ### Импорт библиотек и опцианальные настройки
# +
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from itertools import combinations
from scipy.stats import ttest_ind
import numpy as np
import warnings
import random
warnings.simplefilter('ignore')
pd.set_option('display.max_rows', 50) # показывать больше строк
pd.set_option('display.max_columns', 50) # показывать больше колонок
# Добавим метод чтобы красиво отобржать текст
from IPython.display import Markdown, display
def printmd(string):
display(Markdown(string))
# -
# ### Загрузка датасета
stud_df = pd.read_csv('stud_math.csv')
# ### Определяем полезные функции
# +
def first_look(df: pd.DataFrame, column: object, outlier_bounds: dict, descriptions: dict):
'''
Функция для осмотра переменных
'''
printmd('### ' + str(column))
if column in list(descriptions.keys()):
printmd(descriptions[column])
if df.loc[:, col].dtypes == np.dtype('O'): # Проверка на строковый тип данных
fig, ax = plt.subplots(figsize=(6, 5))
sns.countplot(df.loc[:, column], ax=ax)
plt.show()
# Print misses percent
misses = (1 - (df[column].count() / df.shape[0]))
print('Процент пропусков: ', round(misses*100, 2), "%")
else:
# Prepare plot data frame
plot_df = pd.DataFrame({'Распределение': df[column]})
if search_mistakes(df, column, outlier_bounds):
low, high = IQR_outlier(df, column, verbose=False)
plot_df['Распределение в границах выбросов'] = df.loc[df.loc[:,column].between(low, high)][column]
# Draw plot
fig, ax = plt.subplots(figsize=(7 * plot_df.shape[1], 4))
plot_df.hist(ax=ax)
plt.show()
# Print misses percent
misses = (1 - (df[column].count() / df.shape[0]))
print('Процент пропусков: ', round(misses*100, 2), "%")
def search_mistakes(df: pd.DataFrame, column: object, outlier_bounds: dict) -> bool:
'''
Функция проверяет переменные на наличие ошибок в данных
'''
if column not in list(outlier_bounds.keys()):
low, high = IQR_outlier(df, column, verbose=False)
if df[column].min() < low or df[column].max() > high:
out_df = df.loc[(~df.loc[:, column].between(
low,high)) & pd.notnull(df.loc[:, column])]
print('Найдены ошибки в количестве: ', out_df.shape[0])
if out_df.shape[0]>10:
print('\nПервые 10:')
display(out_df[:10])
return True
else:
print('Выбросы не обнаружены')
else:
low, high = outlier_bounds[column][0], outlier_bounds[column][1]
if df[column].min() < low or df[column].max() > high:
out_df = df.loc[(~df.loc[:, column].between(
low,high)) & pd.notnull(df.loc[:, column])]
print('Найдены выбросы по IQR в количестве: ', out_df.shape[0])
if out_df.shape[0]>10:
print('\nПервые 10:')
display(out_df[:10])
return True
else:
print('Ошибки не обнаружены')
def IQR_outlier(df: pd.DataFrame, column: object, verbose: bool=True) -> tuple:
'''
Функция для отображения границ межквартильного размаха
'''
perc25 = df[column].quantile(0.25)
perc75 = df[column].quantile(0.75)
IQR = perc75 - perc25
low = perc25 - 1.5*IQR
high = perc75 + 1.5*IQR
if verbose:
print('25-й перцентиль: {},'.format(perc25)[:-1], '75-й перцентиль: {},'.format(perc75),
"IQR: {}, ".format(IQR), "Границы выбросов: [{f}, {l}].".format(f=low, l=high))
return (low, high)
def fill_na(df: pd.DataFrame, column: object, method='auto') -> None:
'''
Функция для заполнения пропущенных значений
Варианты аргумента method:
mode - заполнение наиболее частовстречаемым значением
median - заполнение медианой
density - случайное заполнение с учетом вероятности появления
auto - для численных переменных медиана, для номинативных и смешанных density
'''
if method=='auto':
if (df[column].dtypes == np.dtype('O')) and (df[column].nunique()<10):
prob = df[column].value_counts(normalize=True)
ind = df[df[column].isna()==True].index
fill_list = pd.Series(random.choices(prob.index, prob, k=df[column].isna().sum()))
fill_list.index = ind
df[column] = df[column].fillna(fill_list)
else:
median = df[column].median()
df[column] = df[column].fillna(median)
elif method=='mode':
mode = df[column].mode()[0]
df[column] = df[column].fillna(mode)
elif method=='median':
median = df[column].median()
df[column] = df[column].fillna(median)
elif method=='density':
prob = df[column].value_counts(normalize=True)
ind = df[df[column].isna()==True].index
fill_list = pd.Series(random.choices(prob.index, prob, k=df[column].isna().sum()))
fill_list.index = ind
df[column] = df[column].fillna(fill_list)
else:
raise NameError('method может принимать следующие значения: mode, median, density, auto')
def get_boxplot(df: pd.DataFrame, column: object, ax_y='score') -> None:
'''
Функция для отображения boxplot-графиков для номинативных и смешанных переменных
'''
fig, ax = plt.subplots(figsize = (8, 4))
box_plot = sns.boxplot(x=column, y=ax_y, data=df,ax=ax)
cnt = df[column].value_counts()
medians = stud_df.groupby([column])[ax_y].median()
vertical_offset = stud_df.groupby([column])[ax_y].median() * 0.12
x=stud_df[column].unique()
x = list(filter(lambda v: v==v, x))
if df.loc[:, col].dtypes != np.dtype('O'):
x = sorted(x)
for xtick, ytick in zip(list(box_plot.get_xticks()), x):
box_plot.text(xtick, medians.loc[ytick]-vertical_offset[ytick], cnt.loc[ytick],
horizontalalignment='center',size=15, color='w',weight='semibold')
plt.xticks(rotation=45)
ax.set_title('Boxplot for ' + column)
plt.show()
def get_stat_dif(df: pd.DataFrame, column: object, target='score') -> bool:
'''
Функция для проведения теста Стьюдента для номинативных и смешанных переменных
'''
cols = df.loc[:, column].value_counts()
cols = cols[cols>15].index
combinations_all = list(combinations(cols, 2))
for comb in combinations_all:
if ttest_ind(df.loc[df.loc[:, column] == comb[0], target].dropna(),
df.loc[df.loc[:, column] == comb[1], target].dropna()).pvalue \
<= 0.075/len(combinations_all): # Учли поправку Бонферони
print('Найдены статистически значимые различия для колонки', column)
return True
break
# -
# <a id='1'></a>
# ## Первичный осмотр данных
stud_df.head()
stud_df.info()
# В датасете содержится информация о 395 учениках. Все столбцы, кроме первых 3-х (shool, sex, age) имеют в значениях пропуски. Данные представлены в 30 столбцах, из них 17 столбцов имеют строковый тип, остальные 13 - числовой тип.
#
# Однако в числовые столбцы попали упорядоченные категориальные переменные, у которых значения закодированы числовым диапазоном.
#
# Поэтому по факту у нас есть только 3 истинно количественных переменных: age (непрерывная количиственная), absences и score (дискретные количественные переменные), при этом score - целевая переменная.
#Проверим датасет на наличие дубликатов:
if len(stud_df)>len(stud_df.drop_duplicates()):
print('Дубликаты есть')
display(stud_df[stud_df.duplicated()])
else:
print('Дубликатов нет')
# Посмотрим как распеделены пропуски по датасету:
plt.figure(figsize=(10, 7))# размер тепловой карты
sns.heatmap(stud_df.isnull(), cmap="YlGnBu", annot=True);
# Как видим, пропуски распределены равномерно по всему датасету. Какой-то концентрации в строках не видно. Значит удаление в качестве решения - не наш вариант.
# Проверим какое максимальное количество пропусков в строке:
Count_rows = stud_df.apply(lambda x: sum(x.isnull()), axis=1).value_counts()
percent_nans=round(pd.Series(Count_rows.index/stud_df.shape[1]*100)).sort_values(ascending=False).astype(str)+' %'
misses = max(stud_df.apply(lambda x: sum(x.isnull()), axis=1))/stud_df.shape[1]
print('Максимум незаполненных строк в датафрейме:', round(misses*100, 2), "%")
pd.DataFrame({'Количество случаев строке': Count_rows,
'Количество пропусков в строке': Count_rows.index,
'Процент незаполненных значений в строке': percent_nans}).sort_values('Количество пропусков в строке', ascending=False).reset_index().drop('index',axis=1)
# Как видим процент не достаточно высок (>=17% только в 7 случаях), чтобы принять решение об удалении отдельных проблемных строк.
# Считаем процент пропусков
per_nans = pd.DataFrame(1-stud_df.count()/stud_df.shape[0]).reset_index().rename({0:'percent'}, \
axis=1).sort_values('percent', ascending=False).query('percent>0')
# Строим график относительного количества пропущенных значений:
fig, ax = plt.subplots(figsize = (10, 8))
bar_plot = sns.barplot(x='percent', y='index', data=per_nans, alpha=0.8, ci=None)
for ytick in bar_plot.get_yticks():
bar_plot.text(per_nans.iloc[ytick][1]+0.005, ytick+0.2, str(round(per_nans.iloc[ytick][1]*100,2))+' %',
horizontalalignment='center',size=10, color='black',weight='semibold')
plt.title('Относительное количество пропусков')
plt.ylabel('')
plt.xlabel('Percent', fontsize=12)
plt.show()
# <a id='2'></a>
# ## Оценка распределения значений для количественных признаков, поиск и устранение ошибок/аномальных значений.
# Объявим словарь описаний полей, для дальнейшего использования при анализе
descriptions = {
"school": "аббревиатура школы, в которой учится ученик",
"sex": "пол ученика ('F' - женский, 'M' - мужской)",
"age": "возраст ученика (от 15 до 22)",
"address": "тип адреса ученика ('U' - городской, 'R' - за городом)",
"famsize": "размер семьи('LE3' <= 3, 'GT3' >3)",
"Pstatus": "статус совместного жилья родителей ('T' - живут вместе 'A' - раздельно)",
"Medu": "образование матери (0 - нет, 1 - 4 класса, 2 - 5-9 классы, 3 - среднее специальное или 11 классов, 4 - высшее)",
"Fedu": "образование отца (0 - нет, 1 - 4 класса, 2 - 5-9 классы, 3 - среднее специальное или 11 классов, 4 - высшее)",
"Mjob": "работа матери ('teacher' - учитель, 'health' - сфера здравоохранения, 'services' - гос служба, 'at_home' - не работает, 'other' - другое)",
"Fjob": "работа отца ('teacher' - учитель, 'health' - сфера здравоохранения, 'services' - гос служба, 'at_home' - не работает, 'other' - другое)",
"reason": "причина выбора школы ('home' - близость к дому, 'reputation' - репутация школы, 'course' - образовательная программа, 'other' - другое)",
"guardian": "опекун ('mother' - мать, 'father' - отец, 'other' - другое)",
"traveltime": "время в пути до школы (1 - <15 мин., 2 - 15-30 мин., 3 - 30-60 мин., 4 - >60 мин.)",
"studytime": "время на учёбу помимо школы в неделю (1 - <2 часов, 2 - 2-5 часов, 3 - 5-10 часов, 4 - >10 часов)",
"failures": "количество внеучебных неудач (n, если 1<=n<=3, иначе 0)",
"schoolsup": "дополнительная образовательная поддержка (yes или no)",
"famsup": "семейная образовательная поддержка (yes или no)",
"paid": "дополнительные платные занятия по математике (yes или no)",
"activities": "дополнительные внеучебные занятия (yes или no)",
"nursery": "посещал детский сад (yes или no)",
"higher": "хочет получить высшее образование (yes или no)",
"internet": "наличие интернета дома (yes или no)",
"romantic": "в романтических отношениях (yes или no)",
"famrel": "семейные отношения (от 1 - очень плохо до 5 - очень хорошо)",
"freetime": "свободное время после школы (от 1 - очень мало до 5 - очень мого)",
"goout": "проведение времени с друзьями (от 1 - очень мало до 5 - очень много)",
"health": "текущее состояние здоровья (от 1 - очень плохо до 5 - очень хорошо)",
"absences": "количество пропущенных занятий",
"score": "баллы по госэкзамену по математике"
}
# Определим словарь с границами данными по условию:
outline_bounds = {
"age": [15, 22],
"Medu": [0, 4],
"Fedu": [0, 4],
"traveltime": [1, 4],
"studytime": [1, 4],
"failures": [0, 3],
"famrel": [1, 5],
"freetime": [1, 5],
"goout": [1, 5],
"health": [1, 5],
'score': [0,100]
}
for col in stud_df.columns:
first_look(stud_df, col, outline_bounds, descriptions)
# ### Промежуточный выводы:
# Данные достаточно чистые, однако есть некоторые проблемы, а именно:
# * в некоторых переменных (higher, Pstatus, school, famrel) наблюдается дисбаланс классов;
# * на основе age можно создать доп. переменную, объединяющую редких великовозрастных школьников;
# * на основе признаков поддержки (schoolsup, famsup) можно создать признак наличия поддержки в общем виде;
# * в переменных Fedu, famrel обнаружены ошибки: значения недопустимы по условию задания;
# * в переменной Absences, "studytime,granular" в наличии выбросы;
# * целевая переменная score содержит 1,5%: пропусков;
# * распределение неописанной в задании studytime, granular напоминает распределение studytime.
#Отдельно посмотрим на Absences
plt.figure(figsize=(8, 4))
sns.boxplot(data=stud_df['absences'], orient="h",
palette="Set2", whis=1.5);
# Как видим, только 2 значения сильно выбиваются из общей массы. Их сложно объяснить логически и они, скорее всего, лишь запутают будущую модель. Так как мы работаем в условиях весьма ограниченного объема данных, то вместо удаления заменим их на типичное значение - медиану.
#Код предобработки по результатам анализа:
stud_df.Fedu = stud_df.Fedu.replace(40,4)
stud_df.famrel = stud_df.famrel.replace(-1,1)
stud_df.absences = stud_df.absences.apply(lambda x: stud_df.absences.median() if x>100 else x)
stud_df['age_cat'] = stud_df.age.apply(lambda x: 'young' if x<18 else 'old')
stud_df['is_sup'] = np.where((stud_df.schoolsup == 'yes') | (stud_df.famsup == 'yes'), 'yes', 'no')
stud_df.dropna(subset=['score'], inplace=True)
# В переменной "studytime,granular" также в наличии выбросы, но пока с ними ничего делать не будем.
# Рассмотрим подробнее нашу целевую переменную score
# +
# Смотрим графики
fig, axes = plt.subplots(ncols=2, figsize=(6, 4))
sns.boxplot(data=stud_df['score'], orient="h",
palette="Set2", whis=1.5, ax=axes[0]);
stud_df.score.hist(bins=20, ax=axes[1])
fig.tight_layout(rect=[-1, 0.03, 1, 0.95])
plt.show()
search_mistakes(stud_df, 'score',outline_bounds)
# -
# #### Обращаем внимание:
#
# * более 30 учеников, получивших 0 баллов
# * 'яма' в диапазоне от 0 до 20 после чего начинается нормальное распределение
# * выбросов и ошибок нет
# Провал, на наш взгляд, можно объяснить только наличием "проходного балла", т.е., если школьник не набрал "пороговое" значение 20, то ему проставлялся 0. Вообще, большое количество 0 оценок выглядит подозрительно. Но на этапе разведывательного анализа считаем, что правильнее эти значения оставить, и посмотреть сможет ли будущая модель предсказывать склонных к провалу экзамену учеников.
# <a id='3'></a>
# ## Корреляционный анализ количественных признаков.
# Выясним какие столбцы коррелируют с оценкой на госэкзамене по математике. Это поможет понять какие признаки стоит оставить для модели, а какие нужно будет исключить из анализа
#
# - **по количественным переменным**:
stud_num = ['age', 'absences', 'score']
sns.pairplot(stud_df[stud_num], kind='reg');
stud_df[stud_num].corr()
# Как видим линейная связь absences со score очень незначительна. Проверим наличие нелинейной связи.
plt.figure(figsize=(14, 7))
sns.lineplot(data=stud_df, x='absences', y='score');
# Делаем вывод о низкой значимости предиктора для будущей модели
# - **по количественным и смешанным переменным**:
# Используем тепловую карту для удобства:
quantitative_features = stud_df.select_dtypes(include='number').columns
corr_matrix = stud_df[quantitative_features].corr().round(2)
plt.rcParams['figure.figsize'] = (10, 9)
matrix = np.triu(corr_matrix)
x = sns.heatmap(corr_matrix, mask=matrix, annot=True)
# Как видим, не описанная в условии переменная "studytime, granular" имеет полную обратную корреляцию с studytime. Соответственно, она не несет доп. информации и ее следует удалить. Также довольна сильная линейная зависимость между переменными Fedu, Medu. Это можно использовать для:
# * восстановления пропущенных значений
# * генерации дополнительного признака на основе двух исходных.
# +
# Сделаем отдельный датафрейм для кореллирующих значений
score_correlation_df = pd.DataFrame(stud_df.corr()["score"].values,
index=stud_df.corr()["score"].index,
columns=['correlation'])
score_correlation_df = score_correlation_df.drop("score")
score_correlation_df['corellation_amount'] = abs(
score_correlation_df.correlation)
printmd("#### Кореллирующие значения в порядке убывания")
score_correlation_df.sort_values(by='corellation_amount', ascending=False)
# -
# Из этой таблицы можно сделать вывод, что больше всего в обучении мешают проблемы вне учебных заведений, а также юный возраст и активное общение с друзьями. А позитивно на результатах сказывается образование родителей и самостоятельное обучение. Как ни странно, количество прогулов и свободного времени после занятий не оказывает заметного влияния на результаты экзамена.
#
# В целях очистки датасета, удалим стобцы с значением корелляции ниже 0.1, однако для будущей модели, возможно, их тоже можно было бы принять к рассмотрению, если останется время на эксперименты.
#Код предобработки на основе корреляционного анализа:
stud_df.drop('studytime, granular', axis=1, inplace=True)
stud_df.drop(score_correlation_df[score_correlation_df.corellation_amount < 0.1].index, axis=1, inplace=True)
stud_df['P_edu'] = stud_df['Fedu']+stud_df['Medu']
# <a id='4'></a>
#
# ## Визуальный анализ номинативных признаков, оценка количества уникальных значений.
nom_cols = list(set(stud_df.columns) - set(['age', 'absences', 'score']))
for col in nom_cols:
get_boxplot(stud_df, col)
stud_df['is_dad_teacher'] = stud_df.Fjob.apply(lambda x: 1 if x=='teacher' else 0)
# #### Промежуточный вывод:
# После осмотра "ящиков с усами" перспективными для моделирования предикторами представляются:
# * schoolsup
# * Fedu
# * P_edu
# * failures
# * Mjob
# * Medu
# * is_dad_teacher
# * higher
# * age_cat
# * goout
# * school
# * address
# * studytime
# <a id='5'></a>
# ## t-test номинативных и смешанных переменных
# С помощью теста Стьюдента проверим есть ли статистическая разница в распределении оценок по номинативным признакам, проверив нулевую гипотезу о том, что распределение оценок по госэкзамену в зависимости от уровней категорий неразличимы
nom_cols = list(set(stud_df.columns) - set(['age', 'absences', 'score']))
lst = []
for col in nom_cols:
if get_stat_dif(stud_df, col):
lst.append(col)
# Тест Стьюдента и анализ номинативных переменных с помощью boxplot-графиков взаимодополняющие методы, а не взаимоисключающие. Поэтому на этапе EDA следует оставлять признаки, которые показались информативными и в первом и во втором случае.
# Итоговый датафрейм:
list_columns = set(['schoolsup', 'Fedu', 'P_edu', 'failures', 'Mjob', 'Medu', \
'is_dad_teacher', 'higher', 'age_cat', 'goout', 'school', 'address', 'studytime', 'age', 'score'] + lst)
stud_df = stud_df[list_columns]
stud_df.head()
# <a id='6'></a>
# ## Заполнение пропусков
# Заполнение пропусков обширная и тонкая тема, и там иногда применяются весьма сложные техники. В нашей таблице не так уж и велико отношение количества пропусков к количеству всех элементов (не более 12%!), поэтому в данном случае имплементацией можно пренебречь.
#
# Однако, да бы избежать с ошибками при подаче данных в ML алгоритм, как вариант, использовать простые способы заполнения пропусков. Кроме заполнения модой, номинативные признаки можно заполнять случайным значением переменной, с учетом вероятности появления уровня категории в заполненной переменной. Так, мы сохраним исходное распределение.
# +
# Восстанавливаем пропущенные значения, используя сильную линейную зависимость:
stud_df.Fedu = np.where(stud_df.Fedu.isna(), stud_df.Medu, stud_df.Fedu)
stud_df.Medu = np.where(stud_df.Medu.isna(), stud_df.Fedu, stud_df.Medu)
#Заполняем числовые пропуски медианой, остальные - на основе вероятности появления
for col in stud_df.columns:
fill_na(stud_df, col, method='auto')
# -
# <a id='7'></a>
# ## Итоговый вывод.
# **В результате проведенного EDA можно сделать следующие заключения относильно датасета:**
# 1. Данные достаточно чистые:
# * количество пропущенных значений варьируется от 1% до 11%. Есть три переменные, в которых данные 100% заполнены;
# * ошибки обнаружены в переменных Fedu, famrel и заменены на основе некоторых предположений исходя из здравого смысла;
# * переменная Absences содержала 2 аномальных значения (>200), которые были заменены на медиану с целью сохранения информации, содержащейся в других предикторах.
# 2. После подробного осмотра распределений, было решено:
# * создать признак наличия поддержки в общем виде - is.sup, который оказался незначимым в итоге;
# * создать создать доп. переменную - age_cat, объединяющую редких великовозрастных школьников в одну группу;
# * удалить пропуски из целевой переменной за ненадобностью;
# * оставить 0 значения в score для выяснения возможности моделирования этих случаев.
# 3. В результате корреляционного анализа:
# * обнаружена сильная обратная корреляция между studytume и studytume_granular, поэтому один из них был удален за ненадобностью;
# * обнаружена линейная зависимость между Fedu, Medu, которая была использована для создания нового значимого признака и взаимного восстановления пропусков;
# * экспертно исключены пременные с коэффициентом корреляции менее 0.1 по модулю как самые бесперспективные на этапе EDA.
# 4. Анализ номинативных и смешанных переменных с помощью boxplot и t-testa позволил выделить следующие значимые признаки:
# age_cat, goout, sex, paid, is_dad_teacher, Mjob, failures, Medu, Fedu, address, romantic, schoolsup, school, studytime, higher, P_edu
# **Итоговый ответ:**
# Для дальнейшего моделирования рекомендуется использовать параметры: sex, address, Mjob, schoolsup, paid, higher, romantic, age, Medu, Fedu, studytime, failures, goout - как наиболее перспективные.
| module_2/Module_suppl_notebooks/Project 2 - EDA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/FernandoMartinezHernandez/daa_2021_1/blob/master/7_Octubre.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="Rrxzbp49Sik3"
# #Busqueda Lineal
# Dado un conjunto de datos no ordenada, la busqueda lineal consste en recorrer el conjunto de datos desde el inicio al final, moviendose de uno en uno hasta encontrar el elemento o llegar al final del conjunto.
#
# datos = [4.18,47,2,34,14,78,12,48,21,31,19,1,3,5]
#
# #Busqueda Binaria
# Funciona sobre un conjun to de datos ordenado.
# Consiste en dividir el conjunto en mitades y buscar en esa mitad. Si el elemento buscado no esta en la mitad, preguntas si el elemento esta a la derecha o a la izquierda.
# Haces la lista igual a lamitad correspondiente y repites el proceso
#
# L= [1, 2, 3, 4.18, 5, 12, 14, 19, 21, 31, 34, 47, 48, 78]
#
# Der = longitud(L)-1
#
# Izq =0
#
# Mid apuntara a la mitad del segmento de busqueda
#
# buscado: es el valor a buscar
#
# 1. Hacer Der = longitu(L)-1
# 2. Hacer Izq =0
# 3. Si Izq > Der significa que el arreglo no esta ordenado
# 4. Calcular Mid = int((Izq + Der)/2)
# 5. Mientras L[MD] != buscado hacer
# 6. - Preguntar L[MID] > buscado
# -hacer Der = MID
# - de lo contrario
# -Hacer Izq = mid
# - preguntar (der - izq) % 2
# -MID = (Izq + ((Der - Izq)/2))+1
# - de lo contrario
# -Mid = Izq + ((Der - Izq)/2)
# 7. return Mid
# + id="moDH6m-XShcI" outputId="4a5625bb-3dbd-44e1-8ab7-dfc2c798c018" colab={"base_uri": "https://localhost:8080/", "height": 109}
"""
Busqueda lineal
regresa la posicion del elemento "Buscado" si se encuentra dentro de la lista
regresa -1 si el elemento no existe dentro de la lista
"""
def busq_lineal (L, buscado):
indice = -1
contador =0
for idx in range(len(L)):
contador +=1
if L[idx] == buscado:
indice = idx
break
#print(f"Numero de comparaciones{contador}")
return indice +1
"""
Busqueda Binaria
"""
def busq_bin( L, buscado):
indice =-1
Izq =0
Der = len(L)-1
Mid= 0
while not (Izq>Der):
Mid =(Izq +Der)//2
if L[Mid] < buscado:
Izq = Mid+1
else:
Der = Mid-1
print(f"Comparar buscado{buscado} con {L[Mid]}")
if L[Mid] == buscado:
indice = Mid
break
return indice
def busq_bin_recur(L,buscado,Izq,Der):
Mid = (Izq +Der)//2
if Izq >= Der:
return -1
if L[Mid]== buscado:
return Mid
elif L[Mid]<buscado:
return busq_bin_recur(L,buscado,Mid+1,Der)
else:
return busq_bin_recur(L,buscado,Izq,Mid)
def main():
datos = [4.18,47,2,34,14,78,12,48,21,31,19,1,3,5]
dato = int(input("¿Que valor desea buscar?"))
resultado = busq_lineal(datos, dato)
#print("Resultado: ", resultado)
#print("Busqueda lineal en una lista ordenada")
datos.sort()
print(datos)
resultado = busq_lineal(datos, dato)
#
print("Resultado: ", resultado)
print("Busqueda Binaria")
#posicion = busq_bin(datos, dato)
#print(f"El elemento {dato} esta en la posicion { posicion+1 } de la lista")
posicion = busq_bin_recur(datos,dato,0,len(datos))
print(f"El elemento {dato} esta en la posicion { posicion+1 } de la lista")
main()
| 7_Octubre.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# Root Finding
# ===========
#
# [Download from](https://github.com/kpenev/REUpython/raw/master/Roots%2C%20Integration%20and%20ODEs.ipynb)
# + slideshow={"slide_type": "skip"}
from IPython.core.display import display, HTML
from matplotlib import pyplot, rcParams
display(HTML("<style>.container { width:108ch !important; }</style>"))
# + slideshow={"slide_type": "slide"}
import scipy
class DemonstrationFunction:
"""
A callable defining a polynomial used in the subsequent demonstrations.
We use a class instead of a function so that we can spy on the
solving process.
Attributes:
coefficients (scipy.array(dtype=float)): The coefficients of the
polynomial. The n-th entry is the coefficient in front of x^n.
evaluations (dict('x'=[], 'y'=[]): Stores the points where the
function was evaluated and the result of the evaluation.
"""
def __init__(self, coefficients):
"""Create a polynomial with the specified coefficients."""
self.coefficients = coefficients
self.evaluations = dict(x=[], y=[])
def __call__(self, x):
"""Evaluate the polynomial at the given abscissa."""
x_to_n = 1.0
result = 0.0
for coefficient in self.coefficients:
result += coefficient * x_to_n
x_to_n *= x
self.evaluations['x'].append(x)
self.evaluations['y'].append(result)
return result
def derivative(self, x, deriv_order=1):
"""Return the derivative of the specified order at x."""
deriv_factor = scipy.math.factorial(deriv_order)
result = 0.0
x_to_n = 1.0
for x_power, coefficient in enumerate(
self.coefficients[deriv_order:]
):
result += deriv_factor * coefficient * x_to_n
deriv_factor *= (x_power + deriv_order + 1) / (x_power + 1)
x_to_n *= x
return result
def reset_evaluations(self):
"""Clear the current function evaluation history."""
self.evaluations = dict(x=[], y=[])
# + [markdown] slideshow={"slide_type": "slide"}
# **Plotting the function**
# + slideshow={"slide_type": "fragment"}
import matplotlib
from matplotlib import pyplot
def setup_readable_plots():
"""Configure matplotlib to create readable plots."""
matplotlib.rc('figure', figsize=(15,6))
matplotlib.rc('font', size=24)
def plot(function, derivative, plot_range, tangent_x=-0.5):
"""Plot the given function in the given range."""
plot_x = scipy.linspace(*plot_range, 1000)
tangent_line = lambda x: (function(tangent_x)
+
derivative(tangent_x) * (x - tangent_x))
pyplot.axhline(0.0, color='black', linewidth=3)
pyplot.plot(plot_x,
function(plot_x),
color='red',
linewidth=3,
label='f(x)')
pyplot.plot(plot_x,
tangent_line(plot_x),
color='blue',
linewidth=2,
label='tangent')
pyplot.xlabel('x')
pyplot.ylabel('y')
pyplot.legend()
# + hideCode=false slideshow={"slide_type": "slide"}
if __name__ == '__main__':
coefficients = scipy.linspace(-3, 2, 6)
coefficients[0] += 2.5
polynomial = DemonstrationFunction(coefficients)
setup_readable_plots()
pyplot.subplot(1, 2, 1)
plot(polynomial, polynomial.derivative, (-1.1, 1.1), tangent_x=-0.619)
pyplot.title('Function')
pyplot.subplot(1, 2, 2)
plot(polynomial.derivative,
lambda x: polynomial.derivative(x, 2),
(-1.1, 1.1))
pyplot.title('First Derivative')
# + [markdown] slideshow={"slide_type": "slide"}
# Finding Roots
# ===========
#
# [Here is the documentation](https://docs.scipy.org/doc/scipy/reference/optimize.html#root-finding)
# + [markdown] hideCode=true slideshow={"slide_type": "slide"}
# Let's display what the solver is doing
# ----------------------------------------------------
# + slideshow={"slide_type": "fragment"}
def plot_solver_iterations(function, evaluations, plot_range=None):
"""Create a plot demonstrating the root finding iterations."""
if plot_range is None:
plot_range = min(evaluations['x']), max(evaluations['x'])
plot_x = scipy.linspace(*plot_range, 1000)
pyplot.plot(plot_x,
function(plot_x),
color='black',
linewidth=2,
label='f(x)')
pyplot.axhline(0.0, color='blue', linewidth=2)
offsets = {var: evaluations[var][1:] - evaluations[var][:-1]
for var in ['x', 'y']}
pyplot.quiver(evaluations['x'][:-1],
evaluations['y'][:-1],
offsets['x'],
offsets['y'],
color='red',
width=5e-3,
angles='xy',
scale_units='xy',
scale=1,
label='iterations')
pyplot.xlabel('x')
pyplot.ylabel('y')
pyplot.legend()
# + [markdown] slideshow={"slide_type": "slide"}
# Newton's method
# -------------------------
#
# [Here is the documentation](https://docs.scipy.org/doc/scipy/reference/generated/scipy.optimize.newton.html#scipy.optimize.newton)
# + slideshow={"slide_type": "fragment"}
polynomial.reset_evaluations()
solution = scipy.optimize.newton(func=polynomial,
x0=-0.622,
fprime=polynomial.derivative)
solver_evaluations = {var:scipy.array(polynomial.evaluations[var])
for var in ['x', 'y']}
plot_solver_iterations(polynomial, solver_evaluations)
pyplot.suptitle("Newton's method");
# + [markdown] slideshow={"slide_type": "slide"}
# Secant method
# ----------------------
#
# + slideshow={"slide_type": "fragment"}
polynomial.reset_evaluations()
solution = scipy.optimize.newton(func=polynomial,
x0=-0.6)
solver_evaluations = {var:scipy.array(polynomial.evaluations[var])
for var in ['x', 'y']}
plot_solver_iterations(polynomial, solver_evaluations)
pyplot.suptitle('Secant method')
# + [markdown] slideshow={"slide_type": "slide"}
# Numerical Integration
# ==================
#
# [See the documentation](https://docs.scipy.org/doc/scipy/reference/tutorial/integrate.html)
# + slideshow={"slide_type": "fragment"}
from scipy import integrate
quad_integral = lambda x: (integrate.quad(polynomial.derivative,
0.0,
x)[0]
+
polynomial(0.0))
plot_x = scipy.linspace(-1.0, 1.0, 100)
pyplot.plot(plot_x, polynomial(plot_x), color='black', linewidth=7)
pyplot.plot(plot_x,
[quad_integral(x) for x in plot_x],
color='yellow',
linewidth=4)
# + [markdown] slideshow={"slide_type": "slide"}
# Ordinary Differential Equations
# =========================
#
# [See the documentation](https://docs.scipy.org/doc/scipy/reference/tutorial/integrate.html#ordinary-differential-equations-odeint)
# + [markdown] slideshow={"slide_type": "slide"}
# In one dimension
# ------------------------
# + slideshow={"slide_type": "fragment"}
integrated = integrate.odeint(lambda y, x: polynomial.derivative(x),
polynomial(-1.0),
plot_x)
pyplot.plot(plot_x, polynomial(plot_x), color='black', linewidth=7)
pyplot.plot(plot_x, integrated, color='yellow', linewidth=4);
# + [markdown] slideshow={"slide_type": "slide"}
# Damped Harmonic Oscillator
# -----------------------------------------
#
# Standard way to express ODE:
#
# $ \vec{y} = \left(\begin{array}{c} x \\ \frac{dx}{dt}\end{array}\right)$
#
# Then:
#
# $ \frac{d\vec{y}}{dt} = \left(\begin{array}{c} y_1 \\ -k y_0 - f y_1\end{array}\right)$
# + slideshow={"slide_type": "fragment"}
def damped_oscillator_eom(offset_velocity,
time,
restoring_constant,
damping_coefficient):
"""
Equation of motion for damped harmonic oscillator.
Args:
offset_speed (2-element array): The current offset and velocity
of the oscillator.
time: The time wheth the equation of motion is being evaluated
(ignored).
restoring_consant (float): The constant defining the restoring
force.
damping_coefficient (float): The coefficient defining the damping
force.
Returns:
2-element array:
The time derivative of the displacement and speed.
"""
offset, velocity = offset_velocity
return [velocity,
-restoring_constant * offset - damping_coefficient * velocity]
# + slideshow={"slide_type": "slide"}
plot_t = scipy.linspace(0, 100, 1000)
solution = integrate.odeint(damped_oscillator_eom,
[1.0, 0.0],
plot_t,
args=(1.0, 0.1))
pyplot.subplot(1, 2, 1)
pyplot.plot(plot_t, solution[:, 0], linewidth=3, color='black')
pyplot.xlabel('time')
pyplot.title('offset')
pyplot.subplot(1, 2, 2)
pyplot.plot(plot_t, solution[:, 1], linewidth=3, color='black')
pyplot.xlabel('time')
pyplot.title('velocity');
| Roots, Integration and ODEs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # BagOfWords
import nltk
nltk.download()
paragraph = '''In a country like India with a galloping population, unfortunately nobody is paying attention to the issue
of population. Political parties are feeling shy, politicians are feeling shy, Parliament also does not adequately discuss
about the issue,” said Naidu while addressing the 58th convocation of Indian Agricultural Research Institute (IARI).
He said, “You know how population is growing, creating problems. See the problems in Delhi, traffic, more human beings,
more vehicles, more tension, less attention. If you have tension you cannot pay attention.”
Emphasising on the need to increase food production to meet demand of growing population, Naidu said,
“In future if population increases like this, and you are not able to adequately match it with increase in production,
there will be problem'''
import re
from nltk.corpus import stopwords
from nltk.stem.porter import PorterStemmer
from nltk.stem import WordNetLemmatizer
ps = PorterStemmer()
wordnet = WordNetLemmatizer()
sentences = nltk.sent_tokenize(paragraph)
sentences
len(sentences)
# STEPS: Emplt list -- LOOP -- REGEX (re.sub) -- LOWER() -- SPLIT() -- LISTCOMPREHENSION -- REGEX (join) -- APPND LIST
#list, loop, re.sub, lower, split, listcomprehension, join, append
corpus = []
for i in range(len(sentences)):
review = re.sub('[^a-zA-Z]', ' ', sentences[i])
review = review.lower()
review = review.split()
review = [wordnet.lemmatize(word) for word in review if word not in set(stopwords.words('english'))]
review = ' '.join(review)
corpus.append(review)
corpus
# +
# Creating BagOfWords / Document Matrix
# -
from sklearn.feature_extraction.text import CountVectorizer
cv = CountVectorizer()
X = cv.fit_transform(corpus).toarray()
X.shape
X
X.view()
# +
# BagOfWords / DocumentMatrix / Text Visualization
# +
# STEPS
# 1. Tokenization (paragraph to sentences or words).
# 2. Histogram (Frequency of word count present in the sentences or words. Words Vs Count).
# 3. Sort the Histogram in Descending order.
# 4. Filter the words (10 or 10K most frequent words depending on data size)
# 5. Create the matrix / Bag of words
# (Words/Documents: Sentence or index of items, Horizontal word list, Target: 0 or 1/Y or N/+ve or -ve for sentimenatl analysis)
# -
import nltk
nltk.download()
paragraph = '''In a country like India with a galloping population, unfortunately nobody is paying attention to the issue
of population. Political parties are feeling shy, politicians are feeling shy, Parliament also does not adequately discuss
about the issue,” said Naidu while addressing the 58th convocation of Indian Agricultural Research Institute (IARI).
He said, “You know how population is growing, creating problems. See the problems in Delhi, traffic, more human beings,
more vehicles, more tension, less attention. If you have tension you cannot pay attention.”
Emphasising on the need to increase food production to meet demand of growing population, Naidu said,
“In future if population increases like this, and you are not able to adequately match it with increase in production,
there will be problem'''
# +
# paragraph
# +
# Cleaning the text
# -
import re
from nltk.corpus import stopwords
from nltk.stem.porter import PorterStemmer
from nltk.stem import WordNetLemmatizer
ps = PorterStemmer()
wordnet = WordNetLemmatizer()
sentences = nltk.sent_tokenize(paragraph)
len(sentences)
corpus = []
# Steps: Loop--Regex (re.sub)--lower()--split()--ListComprehension--RegEx(join)--Append list
for i in range(len(sentences)):
review = re.sub("[^a-zA-Z]", " ", sentences[i])
review = review.lower()
review = review.split()
review = [wordnet.lemmatize(word) for word in review if word not in set(stopwords.words('english'))]
review = ' '.join(review)
corpus.append(review)
# +
# Creating the BagOfWords / DocumentMatrix
# -
from sklearn.feature_extraction.text import CountVectorizer # Create Histogram, count, sort, filter & Creating matrix or BOW
cv = CountVectorizer()
X = cv.fit_transform(corpus).toarray()
X
# +
# X.view()
# -
X.shape
| 004-vk_NLP - BagOfWords or DocumentMatrix.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import nltk
nltk.download_shell()
messages = [line.rstrip() for line in open('smsspamcollection/SMSSpamCollection')]
print(len(messages))
messages[50]
for mess, message in enumerate(messages[:10]):
print(mess, message,'\n')
import pandas as pd
messages = pd.read_csv('smsspamcollection/SMSSpamCollection', sep='\t',
names=['label','message'])
messages.head()
messages.describe()
messages.groupby('label').describe()
messages['length'] = messages['message'].apply(len)
messages.head()
import matplotlib.pyplot as plt
import seaborn as sns
messages['length'].plot.hist(bins=150)
messages['length'].describe()
messages[messages['length']==910]['message'].iloc[0]
messages.hist(column='length', by='label', bins=60, figsize=(12,5))
import string
from nltk.corpus import stopwords
stopwords.words('english')
def text_process(mess):
"""
1. remove punc
2. remove stopwords
3. return list of clean text words
"""
nopunc = [char for char in mess if char not in string.punctuation]
nopunc = ''.join(nopunc)
return [word for word in nopunc.split() if word.lower() not in stopwords.words('english')]
messages['message'].head().apply(text_process)
from sklearn.feature_extraction.text import CountVectorizer
bow_transformer = CountVectorizer(analyzer=text_process).fit(messages['message'])
print(len(bow_transformer.vocabulary_))
message_bow = bow_transformer.transform(messages['message'])
print('Shape of Sparse Matrix: ', message_bow.shape)
message_bow.nnz
from sklearn.feature_extraction.text import TfidfTransformer
tfidf_transformer = TfidfTransformer().fit(message_bow)
tfidf_transformer.idf_[bow_transformer.vocabulary_['university']]
message_tfidf = tfidf_transformer.transform(message_bow)
from sklearn.naive_bayes import MultinomialNB
spam_detect = MultinomialNB().fit(message_tfidf, messages['label'])
all_pred = spam_detect.predict(message_tfidf)
all_pred
from sklearn.model_selection import train_test_split
msg_train, msg_test, label_train, label_test = train_test_split(messages['message'],
messages['label'],test_size=0.3)
from sklearn.pipeline import Pipeline
pipeline = Pipeline([
('bow', CountVectorizer(analyzer=text_process)),
('tfidf', TfidfTransformer()),
('classifier', MultinomialNB())
])
pipeline.fit(msg_train, label_train)
predicitons = pipeline.predict(msg_test)
from sklearn.metrics import classification_report, confusion_matrix
print(classification_report(label_test, predicitons))
print(confusion_matrix(label_test, predicitons))
| Natural Language Processing/NLP.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="HrJkjB4HMB46" colab_type="text"
# Getting HP stock historical data from Alpha Vantage using API_KEY
# + id="et_uRvwoIDQi" colab_type="code" colab={}
import urllib
import urllib3
# + id="dzH5cxQnMKTu" colab_type="code" outputId="d4207a48-ee85-42cd-bb7f-caf8e3681d81" executionInfo={"status": "ok", "timestamp": 1590129294873, "user_tz": -480, "elapsed": 659, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgCYv1_ND6g-5ubWbHtx1AyMq0u9HksnsZXwto-KA=s64", "userId": "17032662719666247644"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
api_key = '<KEY>'
print(api_key)
# + id="q5kS6JFoGO0R" colab_type="code" colab={}
ticker_2 = "HP"
# + id="DLOOZKBsGC_1" colab_type="code" colab={}
url_string = "https://www.alphavantage.co/query?function=
TIME_SERIES_DAILY&symbol=%s&outputsize=full&apikey=%s"%(ticker_2,api_key)
# + [markdown] id="CjOuR1RgNoXr" colab_type="text"
# a varaible is defiend to create a .csv file with the specified stock ticker at the end.
#
# + id="64c2tCuSNy46" colab_type="code" colab={}
HP__historical__OHLC = 'stock_market_data-%s.csv'%ticker_2
# + [markdown] id="i3TJzTRV_O0X" colab_type="text"
# displaying the csv file.
# + id="OLdvygrE_UDD" colab_type="code" outputId="e37ac63c-ac97-4295-efb8-f37cd70e690a" executionInfo={"status": "ok", "timestamp": 1590129444674, "user_tz": -480, "elapsed": 821, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgCYv1_ND6g-5ubWbHtx1AyMq0u9HksnsZXwto-KA=s64", "userId": "17032662719666247644"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
HP__historical__OHLC
# + id="QdzhvQN2IN9d" colab_type="code" outputId="74f6225e-49e6-41a7-95d6-a350c673b590" executionInfo={"status": "error", "timestamp": 1587630149175, "user_tz": -480, "elapsed": 805, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgCYv1_ND6g-5ubWbHtx1AyMq0u9HksnsZXwto-KA=s64", "userId": "17032662719666247644"}} colab={"base_uri": "https://localhost:8080/", "height": 129}
with urllib.request.urlopen() as url:
data = json.loads(url.read().decode())
# extract stock market data
data = data['Time Series (Daily)']
df = pd.DataFrame(columns=['Date','Low','High','Close','Open'])
for k,v in data.items():
date = dt.datetime.strptime(k, '%Y-%m-%d')
data_row = [date.date(),float(v['3. low']),float(v['2. high']),
float(v['4. close']),float(v['1. open'])]
df.loc[-1,:] = data_row
df.index = df.index + 1
print('Data saved to : %s'%file_to_save)
df.to_csv(file_to_save)
# + id="VeUxTnwAIorb" colab_type="code" colab={}
import urllib
with urllib.request.urlopen(ticker__path__string) as stock__url:
data = json.loads(stock__url.read().decode())
data = data['Time Series (Daily)']
df = pd.DataFrame(columns=['Date','Low','High','Close','Open'])
for k,v in data.items():
date = dt.datetime.strptime(k, '%Y-%m-%d')
data_row = [date.date(),float(v['3. low']),float(v['2. high']),
float(v['4. close']),float(v['1. open'])]
df.loc[-1,:] = data_row
df.index = df.index + 1
print('Data saved to : %s'%HP__historical__OHLC)
df.to_csv(HP__historical__OHLC)
| SourceCodes/methodology__data_provider/P2_Methodology__chap3___Data_Provider.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Drop random values from a pandas dataframe column
# This notebook demonstrates how to drop random values from pandas dataframe column, replacing them with numpy.nan.
# ## Imports
import numpy as np
import pandas as pd
import datetime
import random
# ## Generate Data
# First, let's create a date index for our dataframe.
rows_count = 20
dates = pd.date_range('20130101', periods=rows_count); dates
# Next, we'll populate a `value` clumn with random numerical data.
df = pd.DataFrame(np.random.randn(rows_count, 1), index=dates, columns=["value"]);df
indexes = [i for i in range(df.shape[0])]; indexes
df.shape
percent = 0.5
df = df.mask(np.random.random(df.shape) < percent)
df
| notebooks/drop-random-values-from-pandas-dataframe.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from datetime import date
from gs_quant.common import PayReceive
from gs_quant.instrument import IRSwap, IRSwaption
from gs_quant.markets.portfolio import Portfolio
from gs_quant.session import Environment, GsSession
from gs_quant.common import SwapSettlement
# -
# external users should substitute their client id and secret; please skip this step if using internal jupyterhub
GsSession.use(Environment.PROD, client_id=None,
client_secret=None, scopes=('run_analytics',))
# +
swaptions = Portfolio()
# you don't need to specify any parameters to get a valid trade. All properties have defaults
swaptions.append(IRSwaption())
# -
# get list of properties of an interest rate swaption
# Many of these properties overlap with the IRSwap properties (outlined in example '010001_swap_trade_construction')
IRSwaption.properties()
# pay_or_receive can be a string of 'pay', 'receive', 'straddle' (an option strategy
# where you enter into both a payer and receiver) or the PayReceive enum
# relates to whether you expect to pay/receive fixed for the underlying. default is 'straddle'
swaptions.append(IRSwaption(pay_or_receive=PayReceive.Receive))
swaptions.append(IRSwaption(pay_or_receive='Receive'))
# expiration_date is the date the option expires and may be a tenor relative
# to the active PricingContext.pricing_date or a datetime.date, default is '10y'
swaptions.append(IRSwaption(expiration_date='6m'))
swaptions.append(IRSwaption(expiration_date=date(2022, 2, 12)))
# strike is the rate at which the option can be exercised
# It also represents the interest rate on the fixed leg of the swap if the swaption expires ITM. Defaults to Par Rate (ATM).
# Can be expressed as 'ATM', 'ATM+25' for 25bp above par, a-100 for 100bp below par, 0.01 for 1%
swaptions.append(IRSwaption(strike='ATM'))
swaptions.append(IRSwaption(strike='ATM+50'))
swaptions.append(IRSwaption(strike='a-100'))
swaptions.append(IRSwaption(strike=.02))
# effective_date is the start date of the underlying swap and may be a tenor relative
# to the expiration_date or a datetime.date. Default is spot dates from expiration
# For example, for a swaption w/ notional_currency as GBP, spot date is T+0, so effective_date = expiration_date.
# for a swaption w/ notional_currency USD, spot is T+2 days and the effective_date is 2b after expiration_date
swaptions.append(IRSwaption(effective_date='5b'))
swaptions.append(IRSwaption(effective_date=date(2031, 2, 12)))
# An IRSwaption's strike will resolve to an IRSwap's fixed_rate if the swaps' paramaters match and
# the swaption's effective_date is equivalent to the swap's effective_date
s = IRSwap(notional_currency='GBP', effective_date='10y')
swp = IRSwaption(notional_currency='GBP', expiration_date = '10y', effective_date='0b')
print(s.fixed_rate*100)
print(swp.strike*100)
# settlement is the settlement convention of the swaption and can be a string of:
# 'Phys.CLEARED' (enter into cleared swap), 'Cash.PYU' (PYU - Par Yield Unadjusted,
# cash payment calculated with PYU), 'Physical' (enter into a uncleared swap),
# 'Cash.CollatCash' (collateralized, cash settled at expiry) or the SwapSettlement enum
swaptions.append(IRSwaption(settlement=SwapSettlement.Phys_CLEARED))
swaptions.append(IRSwaption(settlement='Cash.PYU'))
# premium is the amount to be exchanged for the option contract. A positive premium will have a
# negative impact on the PV. premium is a default is 0.
swaptions.append(IRSwaption(premium=1e4))
# premium_payment_date is the date when premium is exchanged.
# premium_payment_date can be a datetime.date or a tenor. defaulted to spot dates from the
# PricingContext.pricing_date
swaptions.append(IRSwaption(premium_payment_date='5d'))
swaptions.append(IRSwaption(premium_payment_date=date(2020, 2, 13)))
# in some markets, the convention is for premium to be exchanged at expiration
# this can be expressed by changing the premium_payment_date to the swaption's expiration_date
swaptions.append(IRSwaption(premium_payment_date='5y', expiration_date='5y'))
swaptions.price()
| gs_quant/documentation/02_pricing_and_risk/00_instruments_and_measures/examples/01_rates/000108_swaption_trade_construction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import simtk.openmm.app as app
import simtk.openmm as mm
import simtk.unit as u
import mdtraj
import numpy as np
import time
# -
# ## System
# +
prmtop = app.AmberPrmtopFile('complex.top')
pdb = app.PDBFile('complex.pdb')
system = prmtop.createSystem(nonbondedMethod=app.PME,
constraints=app.HBonds,
nonbondedCutoff=12*u.angstroms,
switchDistance=10*u.angstroms)
topology = mdtraj.Topology.from_openmm(prmtop.topology)
total_steps = 100000 # Reducing for testing purposes 3000000
# -
# ## Integrator, forces and restraints
integrator = mm.LangevinIntegrator(50*u.kelvin, 5/u.picosecond, 0.002*u.picoseconds)
barostat = mm.MonteCarloBarostat(1.0*u.bar, 300*u.kelvin)
# ## Init simulation and positions
# +
simulation = app.Simulation(prmtop.topology, system, integrator)
simulation.context.setPositions(pdb.positions)
simulation.context.setVelocitiesToTemperature(50*u.kelvin)
# -
# ## Minimize energy
# 100 steps while gradually releasing the constraints then
# 1000 steps with no constraints
simulation.minimizeEnergy(maxIterations=1000)
# ## Heating
# +
for temperature in np.linspace(50, 300, 251)*u.kelvin:
integrator.setTemperature(temperature)
simulation.step(100)
simulation.step(5000)
# -
# ## Equilibrate
simulation.system.addForce(barostat)
# +
simulation.step(int(total_steps*1/3))
simulation.saveState('equilibrated.xml')
# -
# ## Production
simulation.reporters.append(app.CheckpointReporter('checkpnt.chk', int(total_steps/10)))
simulation.reporters.append(app.DCDReporter('simulation.dcd', int(total_steps/400)))
simulation.reporters.append(app.StateDataReporter('simulation.log', int(total_steps/200) ,
step=True,
time=True,
totalEnergy=True))
simulation.step(int(total_steps*2/3))
print()
# +
print('Benchmarking...')
initial_time = time.time()
final_time = time.time()
elapsed_time = (final_time - initial_time) * u.seconds
simulated_time = total_steps * integrator.getStepSize()
performance = (simulated_time / elapsed_time)
print('completed %8d steps in %8.3f s : performance is %8.3f ns/day' % (total_steps, elapsed_time / u.seconds, performance / (u.nanoseconds/u.day)))
# -
print(time.time())
| examples/openmm_esmacs/openmm_esmacs_no_constraint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#if else
#check even and odd numbers
Number_to_be_checked=int(input("Enter a natural number to be checked"))
if Number_to_be_checked % 2 == 0:
print(Number_to_be_checked, "is an even number")
else:
print(Number_to_be_checked, "is an odd number")
# +
#if elif else
#check number lesser than 1 and greater than 100
Number_to_be_checked=float(input("Enter a number to be checked"))
if Number_to_be_checked < 10 :
print(Number_to_be_checked, "is lesser than 10")
elif Number_to_be_checked > 100 :
print(Number_to_be_checked, "is greater than 100")
else:
print(Number_to_be_checked, "is between 10 and 100")
# +
#And
#check number below than 5 and positive number
Number_to_be_checked=float(input("Enter a number to be checked"))
if Number_to_be_checked < 5 and Number_to_be_checked > 0 :
print(Number_to_be_checked, "is a positive number below 5")
else:
print(Number_to_be_checked, "NOT APPLICABLE")
# +
#Or
Number_to_be_checked=float(input("Enter a number to be checked :"))
if Number_to_be_checked <1 or Number_to_be_checked <100:
print(Number_to_be_checked,"is satisfying one of the above condition")
else:
print(Number_to_be_checked,"is an invalid input")
# +
Number_to_be_checked=float(input("Enter a number to be checked :"))
if Number_to_be_checked %2 == 0 and Number_to_be_checked <100:
print(Number_to_be_checked,"is an even number below 100")
else:
print(Number_to_be_checked,"is an invalid input")
# +
#for loop
list=[1,23,24-87,34,-2056,35,100,-1]
for i in list:
print(i)
# +
#for and else if
list=[1,23,24-87,34,-2056,35,100,-1]
for i in list:
if i>0:
print(i,"is a positive number")
else:
print(i,"is a negative number")
# +
list1=[1,23,24-87,34,-2056,35,100,6.5,7.2,-1]
pos_list=[]
neg_list=[]
decimal_list=[]
for i in list1:
if i>0:
pos_list.append(i)
elif i<0:
neg_list.append(i)
else:
decimal_list.append(i)
print(pos_list)
print(neg_list)
print(decimal_list)
#how to separate float from integers
# -
list1=[1,23,24,-87,34,-2056,35,100,6.5,7.2,-1]
pos_list=[]
neg_list=[]
for i in list1:
if i>0:
pos_list.append(i)
else:
neg_list.append(i)
print(pos_list)
print(neg_list)
list1=[1,23,24,-87,34,-2056,35,100,6.5,7.2,-1]
pos_list=[]
neg_list=[]
for i in list1:
if i>0:
pos_list.append(i)
else:
neg_list.append(i)
print(pos_list)
print(neg_list)
#if any number % 1 -----0 its an integer and !=0 then its a float
#not equals to !=
print(29%1)
print(2.9%1)
# +
#for and if loop, make a separate list for floats
list1=[24.1,23,43,1.2]
floatlist=[]
for i in list1:
if i%1!=0:
floatlist.append(i)
floatlist
# +
#while loop
count=0
while (count<10):
print("I am:",count)
count=count+1
print("Thanks")
# +
#Range function (start,end, interval)
list1=[range(0,15,3)]
list1
# -
list1=[*range(0,15,3)]
list1
# +
list1=[*range(0,10)]
for i in list1:
print("I am:",i)
print("Thanks")
print(list1)
# +
#Guessing game
import random
number=random.randint(1,20)
guess=0
count=0
while guess!=number:
guess=input("guess the number: ")
if guess=="exit":
break
guess=int(guess)
count=count+1
if guess < number:
print("you guessed too low")
elif guess > number:
print("You guessed too high")
else:
print("you guessed the right number that is",number,"in",count, "tries")
# +
#palindrome
string=input("Enter a word to check: ")
string_lower=string.lower()
string1=string_lower[::-1]
if string1==string_lower:
print("It's a palindrome")
else:
print("It's not a palindrome")
# +
string="I am akshata"
string=string.reverse()
print(string)
# -
list=[1,2,3,4,5,6,7]
list.reverse()
print(list)
# +
print("Rules: ")
print("1) Rock wins over scissor")
print("2) Scissor wins over paper")
print("3) Paper wins over rock")
print("-"*100)
print("Enter R for rock")
print("Enter S for scissor")
print("Enter P for paper")
print("Enter exit to quit")
print("-"*100)
while True:
player_1=input("Player_1: ")
player_2=input("Player_2: ")
if player_1=="R" and player_2=="S":
print("Player 1 wins!")
elif player_1=="R" and player_2=="P":
print("Player 2 wins!")
elif player_1=="R" and player_2=="R":
print("It is a tie!")
elif player_1=="S" and player_2=="R":
print("Player 2 wins!")
elif player_1=="S" and player_2=="P":
print("player 1 wins!")
elif player_1=="S" and player_2=="S":
print("its a tie!")
elif player_1=="P" and player_2=="R":
print("player 1 wins!")
elif player_1=="P" and player_2=="S":
print("player 2 wins!")
elif player_1=="P" and player_2=="P":
print("Its a tie!")
elif player_1=="exit" or player_2=="exit":
print("Hope you enjoyed the game!")
break
else:
print("Invalid!")
# -
print("hello kitty")
# +
#Bank program
print("WELCOME TO MANIPAL BANK")
print("-"*100)
Bank_customers={"Akshata":10000, "Ayush":12000, "Eva":8000}
Bank_pins={"Akshata":"7788", "Ayush":1920, "Eva":8286}
User_name=input("Enter your name: ")
User_pin=input("Enter your pin: ")
print("-"*100)
if User_name=="Akshata" and User_pin=="7788":
print("Press 1 to Deposit amount")
print("Press 2 to Withdraw amount")
print("Press 3 to check the balance")
print("Press 4 to transfer the amount")
print("Press 5 to create a new account")
print("Press 6 to delete the current account")
else:
print("Enter valid details!")
print("-"*100)
User=input("Select the option: ")
print("-"*100)
if User=="1":
print("Your current balance is",Bank_customers["Akshata"])
Amount_to_deposit=float(input("Enter the amount to be deposited: "))
print("-"*100)
print("The amount",Amount_to_deposit,"has been deposited")
print("Your current balance now is",Bank_customers["Akshata"]+Amount_to_deposit)
Bank_customers["Akshata"]=10000+Amount_to_deposit
print(Bank_customers)
print("-"*100)
print("Thank You")
elif User=="2":
print("Your current balance is",Bank_customers["Akshata"])
Amount_to_withdraw=float(input("Enter the amount to be withdrawn: "))
print("-"*100)
print("The amount",Amount_to_withdraw,"has been withdrawn")
print("Your current balance now is",Bank_customers["Akshata"]-Amount_to_withdraw)
Bank_customers["Akshata"]=10000-Amount_to_withdraw
print(Bank_customers)
print("-"*100)
print("Thank You")
elif User=="3":
print("Your current balance is",Bank_customers["Akshata"])
print("-"*100)
print("Thank You")
elif User=="4":
print("Your current balance is",Bank_customers["Akshata"])
Amount_transfer_to=input("Transfer amount to: ")
Amount_to_transfer=float(input("Amount to be transferred: "))
if Amount_transfer_to=="Ayush":
Bank_customers["Ayush"]=12000+Amount_to_transfer
print("Rs.",Amount_to_transfer,"has been transferred")
print("Your current balance is",Bank_customers["Akshata"]-Amount_to_transfer)
print("-"*100)
print("Thank You")
elif Amount_transfer_to=="Eva" :
Bank_customers["Eva"]=8000+Amount_to_transfer
print("Rs.",Amount_to_transfer,"has been transferred")
print("Your current balance is",Bank_customers["Akshata"]-Amount_to_transfer)
print("Rs.",Amount_to_transfer,"has been transferred")
print("Your current balance is",Bank_customers["Akshata"]-Amount_to_transfer)
print("-"*100)
print("Thank You")
else:
print("TRY AGAIN")
Bank_customers["Akshata"]=10000-Amount_to_transfer
print(Bank_customers)
elif User=="5":
print("WELCOME TO MANIPAL BANK")
print("-"*100)
Enter_your_name=input("Enter your name: ")
Enter_the_pin=input("Enter the pin: ")
Enter_the_amount=input("Enter the amount to deposit: ")
print("Your new account by the name",Enter_your_name,"has been created with the balance of",Enter_the_amount)
print("Thank you!!")
Bank_customers[Enter_your_name]=Enter_the_amount
Bank_pins[Enter_the_pin]=Enter_the_pin
print(Bank_customers)
print(Bank_pins)
elif User=="6":
Confirm=input("Are you sure? ")
if Confirm=="Yes":
print("Your account has been deleted")
print(Bank_customers)
elif Confirm=="No":
print("Your account is not deleted")
else:
print("TRY AGAIN")
#How to delete an item from dictionary
print("-"*100)
print("-"*100)
print("-"*100)
# +
# RUN THIS ONE
print("WELCOME TO MANIPAL BANK")
print("-"*100)
Bank_customers={"Akshata":10000, "Ayush":12000, "Eva":8000}
Bank_pins={"Akshata":7788, "Ayush":1920, "Eva":8286}
New_account=input("Do you want to create a new account: ")
if New_account=="yes":
print("Welcome to Manipal bank!")
New_user=input("Enter your name: ")
New_pin=input("Enter the pin")
New_deposit=input("How much amount do you want to deposit? ")
print("Your account has been created, hope you enjoy our service!")
Bank_customers[New_user]=New_deposit
Bank_pins[ New_user]=New_pin
print(Bank_customers)
print(Bank_pins)
elif New_account=="No":
print("Thank you!")
User_name=input("Enter your name: ")
User_pin=input("Enter your pin: ")
print("-"*100)
if User_name=="Akshata" and User_pin=="7788":
print("Press 1 to Deposit amount")
print("Press 2 to Withdraw amount")
print("Press 3 to check the balance")
print("Press 4 to transfer the amount")
print("Press 5 to create a new account")
print("Press 6 to delete the current account")
elif User_name=="Ayush" and User_pin=="1920":
print("Press 1 to Deposit amount")
print("Press 2 to Withdraw amount")
print("Press 3 to check the balance")
print("Press 4 to transfer the amount")
print("Press 5 to create a new account")
print("Press 6 to delete the current account")
elif User_name=="Eva" and User_pin=="8286":
print("Press 1 to Deposit amount")
print("Press 2 to Withdraw amount")
print("Press 3 to check the balance")
print("Press 4 to transfer the amount")
print("Press 5 to create a new account")
print("Press 6 to delete the current account")
print("-"*100)
Select_the_option=input("Select the option: ")
print("-"*100)
if Select_the_option=="1" and User_name=="Akshata":
print("Your current balance is",Bank_customers["Akshata"])
Amount_to_deposit=float(input("Enter the amount to be deposited: "))
print("-"*100)
print("The amount",Amount_to_deposit,"has been deposited")
print("Your current balance now is",Bank_customers["Akshata"]+Amount_to_deposit)
Bank_customers["Akshata"]=10000+Amount_to_deposit
print(Bank_customers)
print("-"*100)
print("Thank You")
elif Select_the_option=="1" and User_name=="Eva":
print("Your current balance is",Bank_customers["Eva"])
Amount_to_deposit=float(input("Enter the amount to be deposited: "))
print("-"*100)
print("The amount",Amount_to_deposit,"has been deposited")
print("Your current balance now is",Bank_customers["Eva"]+Amount_to_deposit)
Bank_customers["Eva"]=8000+Amount_to_deposit
print(Bank_customers)
print("-"*100)
print("Thank You")
elif Select_the_option=="1" and User_name=="Ayush":
print("Your current balance is",Bank_customers["Ayush"])
Amount_to_deposit=float(input("Enter the amount to be deposited: "))
print("-"*100)
print("The amount",Amount_to_deposit,"has been deposited")
print("Your current balance now is",Bank_customers["Ayush"]+Amount_to_deposit)
Bank_customers["Eva"]=12000+Amount_to_deposit
print(Bank_customers)
print("-"*100)
print("Thank You")
if Select_the_option=="2" and User_name=="Akshata":
print("Your current balance is",Bank_customers["Akshata"])
Amount_to_withdraw=float(input("Enter the amount to be withdrawn: "))
print("-"*100)
print("The amount",Amount_to_withdraw,"has been withdrawn")
print("Your current balance now is",Bank_customers["Akshata"]-Amount_to_withdraw)
Bank_customers["Akshata"]=10000-Amount_to_withdraw
print(Bank_customers)
print("-"*100)
print("Thank You")
elif Select_the_option=="2" and User_name=="Eva":
print("Your current balance is",Bank_customers["Eva"])
Amount_to_withdraw=float(input("Enter the amount to be withdrawn: "))
print("-"*100)
print("The amount",Amount_to_withdraw,"has been withdrawn")
print("Your current balance now is",Bank_customers["Eva"]-Amount_to_withdraw)
Bank_customers["Eva"]=8000-Amount_to_withdraw
print(Bank_customers)
print("-"*100)
print("Thank You")
elif Select_the_option=="2" and User_name=="Ayush":
print("Your current balance is",Bank_customers["Ayush"])
Amount_to_withdraw=float(input("Enter the amount to be withdrawn: "))
print("-"*100)
print("The amount",Amount_to_withdraw,"has been withdrawn")
print("Your current balance now is",Bank_customers["Ayush"]-Amount_to_withdraw)
Bank_customers["Ayush"]=12000-Amount_to_withdraw
print(Bank_customers)
print("-"*100)
print("Thank You")
if Select_the_option=="3" and User_name=="Akshata":
print("Your current balance is",Bank_customers["Akshata"])
print("-"*100)
print("Thank You")
elif Select_the_option=="3" and User_name=="Ayush":
print("Your current balance is",Bank_customers["Ayush"])
print("-"*100)
print("Thank You")
elif Select_the_option=="3" and User_name=="Eva":
print("Your current balance is",Bank_customers["Eva"])
print("-"*100)
print("Thank You")
if Select_the_option=="4" and User_name=="Akshata":
print("Your current balance is",Bank_customers["Akshata"])
Amount_transfer_to=input("Transfer amount to: ")
Amount_to_transfer=float(input("Amount to be transferred: "))
if Amount_transfer_to=="Ayush":
Bank_customers["Ayush"]=12000+Amount_to_transfer
print("Rs.",Amount_to_transfer,"has been transferred")
print("Your current balance is",Bank_customers["Akshata"]-Amount_to_transfer)
print("-"*100)
print("Thank You")
elif Amount_transfer_to=="Eva" :
Bank_customers["Eva"]=8000+Amount_to_transfer
print("Rs.",Amount_to_transfer,"has been transferred")
print("Your current balance is",Bank_customers["Akshata"]-Amount_to_transfer)
print("-"*100)
print("Thank You")
else:
print("TRY AGAIN")
Bank_customers["Akshata"]=10000-Amount_to_transfer
print(Bank_customers)
elif Select_the_option=="4" and User_name=="Ayush":
print("Your current balance is",Bank_customers["Ayush"])
Amount_transfer_to=input("Transfer amount to: ")
Amount_to_transfer=float(input("Amount to be transferred: "))
if Amount_transfer_to=="Akshata":
Bank_customers["Akshata"]=10000+Amount_to_transfer
print("Rs.",Amount_to_transfer,"has been transferred")
print("Your current balance is",Bank_customers["Ayush"]-Amount_to_transfer)
print("-"*100)
print("Thank You")
elif Amount_transfer_to=="Eva" :
Bank_customers["Eva"]=8000+Amount_to_transfer
print("Rs.",Amount_to_transfer,"has been transferred")
print("Your current balance is",Bank_customers["Ayush"]-Amount_to_transfer)
print("-"*100)
print("Thank You")
if Select_the_option=="4" and User_name=="Eva":
print("Your current balance is",Bank_customers["Eva"])
Amount_transfer_to=input("Transfer amount to: ")
Amount_to_transfer=float(input("Amount to be transferred: "))
if Amount_transfer_to=="Ayush":
Bank_customers["Ayush"]=12000+Amount_to_transfer
print("Rs.",Amount_to_transfer,"has been transferred")
print("Your current balance is",Bank_customers["Eva"]-Amount_to_transfer)
print("-"*100)
print("Thank You")
elif Amount_transfer_to=="Akshata" :
Bank_customers["Akshata"]=10000+Amount_to_transfer
print("Rs.",Amount_to_transfer,"has been transferred")
print("Your current balance is",Bank_customers["Eva"]-Amount_to_transfer)
print("-"*100)
print("Thank You")
else:
print("TRY AGAIN")
Bank_customers["Akshata"]=10000-Amount_to_transfer
Bank_customers["Eva"]=8000-Amount_to_transfer
Bank_customers["Ayush"]=12000-Amount_to_transfer
print(Bank_customers)
if Select_the_option=="5" and User_name=="Akshata":
print("WELCOME TO MANIPAL BANK")
print("-"*100)
Enter_your_name=input("Enter your name: ")
Enter_the_pin=input("Enter the pin: ")
Enter_the_amount=input("Enter the amount to deposit: ")
print("Your new account by the name",Enter_your_name,"has been created with the balance of",Enter_the_amount)
print("Thank you!!")
Bank_customers[Enter_your_name]=Enter_the_amount
Bank_pins[Enter_the_pin]=Enter_the_pin
print(Bank_customers)
print(Bank_pins)
elif Select_the_option=="5" and User_name=="Eva":
print("WELCOME TO MANIPAL BANK")
print("-"*100)
Enter_your_name=input("Enter your name: ")
Enter_the_pin=input("Enter the pin: ")
Enter_the_amount=input("Enter the amount to deposit: ")
print("Your new account by the name",Enter_your_name,"has been created with the balance of",Enter_the_amount)
print("Thank you!!")
Bank_customers[Enter_your_name]=Enter_the_amount
Bank_pins[Enter_the_pin]=Enter_the_pin
print(Bank_customers)
print(Bank_pins)
elif Select_the_option=="5" and User_name=="Ayush":
print("WELCOME TO MANIPAL BANK")
print("-"*100)
Enter_your_name=input("Enter your name: ")
Enter_the_pin=input("Enter the pin: ")
Enter_the_amount=input("Enter the amount to deposit: ")
print("Your new account by the name",Enter_your_name,"has been created with the balance of",Enter_the_amount)
print("Thank you!!")
Bank_customers[Enter_your_name]=Enter_the_amount
Bank_pins[Enter_the_pin]=Enter_the_pin
print(Bank_customers)
print(Bank_pins)
if Select_the_option=="6" and User_name=="Akshata":
Confirm=input("Are you sure? ")
if Confirm=="yes":
print("Your account has been deleted")
print(Bank_customers)
if "Akshata" in Bank_customers:
del Bank_customers["Akshata"]
print(Bank_customers)
elif Confirm=="no":
print("Your account is not deleted")
elif Select_the_option=="6" and User_name=="Ayush":
Confirm=input("Are you sure? ")
if Confirm=="yes":
print("Your account has been deleted")
print(Bank_customers)
if "Ayush" in Bank_customers:
del Bank_customers["Ayush"]
print(Bank_customers)
elif Confirm=="no":
print("Your account is not deleted")
if Select_the_option=="6" and User_name=="Eva":
Confirm=input("Are you sure? ")
if Confirm=="yes":
print("Your account has been deleted")
print(Bank_customers)
if "Eva" in Bank_customers:
del Bank_customers["Eva"]
print(Bank_customers)
elif Confirm=="no":
print("Your account is not deleted")
else:
print("TRY AGAIN")
#How to delete an item from dictionary
print("-"*100)
# +
#Run this one 2
print("WELCOME TO MANIPAL BANK")
print("-"*100)
Bank_customers={"Akshata":10000, "Ayush":12000, "Eva":8000}
Bank_pins={"Akshata":7788, "Ayush":1920, "Eva":8286}
while True:
New_account=input("Do you want to create a new account: ")
if New_account=="yes":
print("Welcome to Manipal bank!")
New_user=input("Enter your name: ")
New_pin=input("Enter the pin")
New_deposit=input("How much amount do you want to deposit? ")
print("Your account has been created, hope you enjoy our service!")
Bank_customers[New_user]=New_deposit
Bank_pins[ New_user]=New_pin
print(Bank_customers)
print(Bank_pins)
elif New_account=="No":
print("Thank you!")
User_name=input("Enter your name: ")
User_pin=input("Enter your pin: ")
print("-"*100)
if User_name=="Akshata" and User_pin=="7788":
print("Press 1 to Deposit amount")
print("Press 2 to Withdraw amount")
print("Press 3 to check the balance")
print("Press 4 to transfer the amount")
print("Press 5 to create a new account")
print("Press 6 to delete the current account")
elif User_name=="Ayush" and User_pin=="1920":
print("Press 1 to Deposit amount")
print("Press 2 to Withdraw amount")
print("Press 3 to check the balance")
print("Press 4 to transfer the amount")
print("Press 5 to create a new account")
print("Press 6 to delete the current account")
elif User_name=="Eva" and User_pin=="8286":
print("Press 1 to Deposit amount")
print("Press 2 to Withdraw amount")
print("Press 3 to check the balance")
print("Press 4 to transfer the amount")
print("Press 5 to create a new account")
print("Press 6 to delete the current account")
print("-"*100)
Select_the_option=input("Select the option: ")
print("-"*100)
if Select_the_option=="1" and User_name=="Akshata":
print("Your current balance is",Bank_customers["Akshata"])
Amount_to_deposit=float(input("Enter the amount to be deposited: "))
print("-"*100)
print("The amount",Amount_to_deposit,"has been deposited")
print("Your current balance now is",Bank_customers["Akshata"]+Amount_to_deposit)
Bank_customers["Akshata"]=10000+Amount_to_deposit
print(Bank_customers)
print("-"*100)
print("Thank You")
elif Select_the_option=="1" and User_name=="Eva":
print("Your current balance is",Bank_customers["Eva"])
Amount_to_deposit=float(input("Enter the amount to be deposited: "))
print("-"*100)
print("The amount",Amount_to_deposit,"has been deposited")
print("Your current balance now is",Bank_customers["Eva"]+Amount_to_deposit)
Bank_customers["Eva"]=8000+Amount_to_deposit
print(Bank_customers)
print("-"*100)
print("Thank You")
elif Select_the_option=="1" and User_name=="Ayush":
print("Your current balance is",Bank_customers["Ayush"])
Amount_to_deposit=float(input("Enter the amount to be deposited: "))
print("-"*100)
print("The amount",Amount_to_deposit,"has been deposited")
print("Your current balance now is",Bank_customers["Ayush"]+Amount_to_deposit)
Bank_customers["Eva"]=12000+Amount_to_deposit
print(Bank_customers)
print("-"*100)
print("Thank You")
if Select_the_option=="2" and User_name=="Akshata":
print("Your current balance is",Bank_customers["Akshata"])
Amount_to_withdraw=float(input("Enter the amount to be withdrawn: "))
print("-"*100)
print("The amount",Amount_to_withdraw,"has been withdrawn")
print("Your current balance now is",Bank_customers["Akshata"]-Amount_to_withdraw)
Bank_customers["Akshata"]=10000-Amount_to_withdraw
print(Bank_customers)
print("-"*100)
print("Thank You")
elif Select_the_option=="2" and User_name=="Eva":
print("Your current balance is",Bank_customers["Eva"])
Amount_to_withdraw=float(input("Enter the amount to be withdrawn: "))
print("-"*100)
print("The amount",Amount_to_withdraw,"has been withdrawn")
print("Your current balance now is",Bank_customers["Eva"]-Amount_to_withdraw)
Bank_customers["Eva"]=8000-Amount_to_withdraw
print(Bank_customers)
print("-"*100)
print("Thank You")
elif Select_the_option=="2" and User_name=="Ayush":
print("Your current balance is",Bank_customers["Ayush"])
Amount_to_withdraw=float(input("Enter the amount to be withdrawn: "))
print("-"*100)
print("The amount",Amount_to_withdraw,"has been withdrawn")
print("Your current balance now is",Bank_customers["Ayush"]-Amount_to_withdraw)
Bank_customers["Ayush"]=12000-Amount_to_withdraw
print(Bank_customers)
print("-"*100)
print("Thank You")
if Select_the_option=="3" and User_name=="Akshata":
print("Your current balance is",Bank_customers["Akshata"])
print("-"*100)
print("Thank You")
elif Select_the_option=="3" and User_name=="Ayush":
print("Your current balance is",Bank_customers["Ayush"])
print("-"*100)
print("Thank You")
elif Select_the_option=="3" and User_name=="Eva":
print("Your current balance is",Bank_customers["Eva"])
print("-"*100)
print("Thank You")
if Select_the_option=="4" and User_name=="Akshata":
print("Your current balance is",Bank_customers["Akshata"])
Amount_transfer_to=input("Transfer amount to: ")
Amount_to_transfer=float(input("Amount to be transferred: "))
if Amount_transfer_to=="Ayush":
Bank_customers["Ayush"]=12000+Amount_to_transfer
print("Rs.",Amount_to_transfer,"has been transferred")
print("Your current balance is",Bank_customers["Akshata"]-Amount_to_transfer)
print("-"*100)
print("Thank You")
elif Amount_transfer_to=="Eva" :
Bank_customers["Eva"]=8000+Amount_to_transfer
print("Rs.",Amount_to_transfer,"has been transferred")
print("Your current balance is",Bank_customers["Akshata"]-Amount_to_transfer)
print("-"*100)
print("Thank You")
else:
print("TRY AGAIN")
Bank_customers["Akshata"]=10000-Amount_to_transfer
print(Bank_customers)
elif Select_the_option=="4" and User_name=="Ayush":
print("Your current balance is",Bank_customers["Ayush"])
Amount_transfer_to=input("Transfer amount to: ")
Amount_to_transfer=float(input("Amount to be transferred: "))
if Amount_transfer_to=="Akshata":
Bank_customers["Akshata"]=10000+Amount_to_transfer
print("Rs.",Amount_to_transfer,"has been transferred")
print("Your current balance is",Bank_customers["Ayush"]-Amount_to_transfer)
print("-"*100)
print("Thank You")
elif Amount_transfer_to=="Eva" :
Bank_customers["Eva"]=8000+Amount_to_transfer
print("Rs.",Amount_to_transfer,"has been transferred")
print("Your current balance is",Bank_customers["Ayush"]-Amount_to_transfer)
print("-"*100)
print("Thank You")
if Select_the_option=="4" and User_name=="Eva":
print("Your current balance is",Bank_customers["Eva"])
Amount_transfer_to=input("Transfer amount to: ")
Amount_to_transfer=float(input("Amount to be transferred: "))
if Amount_transfer_to=="Ayush":
Bank_customers["Ayush"]=12000+Amount_to_transfer
print("Rs.",Amount_to_transfer,"has been transferred")
print("Your current balance is",Bank_customers["Eva"]-Amount_to_transfer)
print("-"*100)
print("Thank You")
elif Amount_transfer_to=="Akshata" :
Bank_customers["Akshata"]=10000+Amount_to_transfer
print("Rs.",Amount_to_transfer,"has been transferred")
print("Your current balance is",Bank_customers["Eva"]-Amount_to_transfer)
print("-"*100)
print("Thank You")
else:
print("TRY AGAIN")
Bank_customers["Akshata"]=10000-Amount_to_transfer
Bank_customers["Eva"]=8000-Amount_to_transfer
Bank_customers["Ayush"]=12000-Amount_to_transfer
print(Bank_customers)
if Select_the_option=="5" and User_name=="Akshata":
print("WELCOME TO MANIPAL BANK")
print("-"*100)
Enter_your_name=input("Enter your name: ")
Enter_the_pin=input("Enter the pin: ")
Enter_the_amount=input("Enter the amount to deposit: ")
print("Your new account by the name",Enter_your_name,"has been created with the balance of",Enter_the_amount)
print("Thank you!!")
Bank_customers[Enter_your_name]=Enter_the_amount
Bank_pins[Enter_the_pin]=Enter_the_pin
print(Bank_customers)
print(Bank_pins)
elif Select_the_option=="5" and User_name=="Eva":
print("WELCOME TO MANIPAL BANK")
print("-"*100)
Enter_your_name=input("Enter your name: ")
Enter_the_pin=input("Enter the pin: ")
Enter_the_amount=input("Enter the amount to deposit: ")
print("Your new account by the name",Enter_your_name,"has been created with the balance of",Enter_the_amount)
print("Thank you!!")
Bank_customers[Enter_your_name]=Enter_the_amount
Bank_pins[Enter_the_pin]=Enter_the_pin
print(Bank_customers)
print(Bank_pins)
elif Select_the_option=="5" and User_name=="Ayush":
print("WELCOME TO MANIPAL BANK")
print("-"*100)
Enter_your_name=input("Enter your name: ")
Enter_the_pin=input("Enter the pin: ")
Enter_the_amount=input("Enter the amount to deposit: ")
print("Your new account by the name",Enter_your_name,"has been created with the balance of",Enter_the_amount)
print("Thank you!!")
Bank_customers[Enter_your_name]=Enter_the_amount
Bank_pins[Enter_the_pin]=Enter_the_pin
print(Bank_customers)
print(Bank_pins)
if Select_the_option=="6" and User_name=="Akshata":
Confirm=input("Are you sure? ")
if Confirm=="yes":
print("Your account has been deleted")
print(Bank_customers)
if "Akshata" in Bank_customers:
del Bank_customers["Akshata"]
print(Bank_customers)
elif Confirm=="no":
print("Your account is not deleted")
elif Select_the_option=="6" and User_name=="Ayush":
Confirm=input("Are you sure? ")
if Confirm=="yes":
print("Your account has been deleted")
print(Bank_customers)
if "Ayush" in Bank_customers:
del Bank_customers["Ayush"]
print(Bank_customers)
elif Confirm=="no":
print("Your account is not deleted")
if Select_the_option=="6" and User_name=="Eva":
Confirm=input("Are you sure? ")
if Confirm=="yes":
print("Your account has been deleted")
print(Bank_customers)
if "Eva" in Bank_customers:
del Bank_customers["Eva"]
print(Bank_customers)
elif Confirm=="no":
print("Your account is not deleted")
break
else:
print("TRY AGAIN")
#How to delete an item from dictionary
print("-"*100)
# -
name="my name is ayush"
name
| bank_program.ipynb |
# ---
# jupyter:
# jupytext:
# notebook_metadata_filter: all,-language_info
# split_at_heading: true
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Permutation and the t-test
#
# In [the idea of permutation]({{ site.baseurl }}/chapters/05/permutation_idea),
# we use permutation to compare a difference between two groups of numbers.
#
# In our case, each number corresponded to one person in the study. The number
# for each subject was the number of mosquitoes flying towards them. The subjects
# were from two groups: people who had just drunk beer, and people who had just
# drunk water. There were 25 subjects who had drunk beer, and therefore, 25
# numbers of mosquitoes corresponding to the "beer" group. There were 18
# subjects who had drunk water, and 18 numbers corresponding to the "water" group.
#
# Here we repeat the permutation test, as a reminder.
#
# As before, you can download the data from [mosquito_beer.csv]({{ site.baseurl
# }}/data/mosquito_beer.csv).
#
# See [this
# page](https://github.com/matthew-brett/datasets/tree/master/mosquito_beer) for
# more details on the dataset, and [the data license page]({{ site.baseurl
# }}/data/license).
#
# +
# Import Numpy library, rename as "np"
import numpy as np
# Import Pandas library, rename as "pd"
import pandas as pd
# Set up plotting
import matplotlib.pyplot as plt
# %matplotlib inline
plt.style.use('fivethirtyeight')
# -
# Read in the data, get the numbers of mosquitoes flying towards the beer
# drinkers, and towards the water drinkers, after they had drunk their beer or
# water. See the [the idea of permutation]({{ site.baseurl
# }}/chapters/05/permutation_idea),
# Read in the data, select beer and water values.
mosquitoes = pd.read_csv('mosquito_beer.csv')
after_rows = mosquitoes[mosquitoes['test'] == 'after']
beer_rows = after_rows[after_rows['group'] == 'beer']
beer_activated = np.array(beer_rows['activated'])
water_rows = after_rows[after_rows['group'] == 'water']
water_activated = np.array(water_rows['activated'])
# There are 25 values in the beer group, and 18 in the water group:
print('Number in beer group:', len(beer_activated))
print('Number in water group:', len(water_activated))
# We are interested in the difference between the means of these numbers:
observed_difference = np.mean(beer_activated) - np.mean(water_activated)
observed_difference
# In the permutation test we simulate a ideal (null) world in which there is no
# average difference between the numbers in the two groups. We do this by
# pooling the beer and water numbers, shuffling them, and then making fake beer
# and water groups when we know, from the shuffling, that the average difference
# will, in the long run, be zero. By doing this shuffle, sample step many times
# we build up the distribution of the average difference. This is the *sampling
# distribution* of the mean difference:
pooled = np.append(beer_activated, water_activated)
n_iters = 10000
fake_differences = np.zeros(n_iters)
for i in np.arange(n_iters):
np.random.shuffle(pooled)
fake_differences[i] = np.mean(pooled[:25]) - np.mean(pooled[25:])
plt.hist(fake_differences)
plt.title('Sampling difference of means');
# We can work out the proportion of the sampling distribution that is greater
# than or equal to the observed value, to get an estimate of the probability of
# the observed value, if we are in fact in the null (ideal) world:
permutation_p = np.count_nonzero(
fake_differences >= observed_difference)/ n_iters
permutation_p
# Remember that the *standard deviation* is a measure of the spread of
# a distribution.
sampling_sd = np.std(fake_differences)
sampling_sd
# We can use the standard deviation as unit of distance in the distribution.
#
# A way of getting an idea of how extreme the observed value is, is to ask how
# many standard deviations the observed value is from the center of the
# distribution, which is zero.
like_t = observed_difference / sampling_sd
like_t
# Notice the variable name `like_t`. This number is rather like the famous [t
# statistic](https://en.wikipedia.org/wiki/T-statistic).
#
# The difference between this `like_t` value and the *t statistic* is that the t
# statistic is the observed difference divided by another *estimate* of the
# standard deviation of the sampling distribution. Specifically it is an
# estimate that relies on the assumption that the `beer_activated` and
# `water_activated` numbers come from a simple bell-shaped [normal
# distribution](https://en.wikipedia.org/wiki/Normal_distribution).
#
# The specific calculation relies on calculating the *prediction errors* when we
# use the mean from each group as the prediction for the values in the group.
beer_errors = beer_activated - np.mean(beer_activated)
water_errors = water_activated - np.mean(water_activated)
all_errors = np.append(beer_errors, water_errors)
# The estimate for the standard deviation of the sampling distribution follows
# this formula. The derivation of the formula is well outside the scope of the
# class.
# The t-statistic estimate.
n1 = len(beer_activated)
n2 = len(water_activated)
est_error_sd = np.sqrt(np.sum(all_errors ** 2) / (n1 + n2 - 2))
sampling_sd_estimate = est_error_sd * np.sqrt(1 / n1 + 1 / n2)
sampling_sd_estimate
# Notice that this is rather similar to the estimate we got directly from the
# permutation distribution:
sampling_sd
# The t statistic is the observed mean difference divided by the estimate of the
# standard deviation of the sampling distribution.
t_statistic = observed_difference / sampling_sd_estimate
t_statistic
# This is the same t statistic value calculated by the *independent sample t
# test* routine from Scipy:
from scipy.stats import ttest_ind
t_result = ttest_ind(beer_activated, water_activated)
t_result.statistic
# The equivalent probability from a t test is also outside the scope of the
# course, but, if the data we put into the t test is more or less compatible with
# a normal distribution, then the matching p value is similar to that of the
# permutation test.
# The "one-tailed" probability from the t-test.
t_result.pvalue / 2
# The permutation p value is very similar.
permutation_p
# The permutation test is more general than the t test, because the t test relies
# on the assumption that the numbers come from a normal distribution, but the
# permutation test does not.
| notebooks/05/permutation_and_t_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import pandas as pd
import numpy as np
hr_data = pd.read_csv('data/hr.csv', header=0)
hr_data.head()
hr_data = hr_data.dropna()
data_trnsf = pd.get_dummies(hr_data, columns =['salary', 'sales'])
data_trnsf.columns
# -
X= data_trnsf.drop('left', axis=1)
X.columns
Y = data_trnsf.left# feature extraction
# +
#Variance Threshold
from sklearn.feature_selection import VarianceThreshold
# Set threshold to 0.1
select_features = VarianceThreshold(threshold = 0.2)
select_features.fit_transform(X)
# Subset features
X_subset = select_features.transform(X)
print('Number of features:', X.shape[1])
print('Reduced number of features:',X_subset.shape[1])
# +
#Chi2 Selector
from sklearn.feature_selection import SelectKBest
from sklearn.feature_selection import chi2
chi2_model = SelectKBest(score_func=chi2, k=4)
X_best_feat = chi2_model.fit_transform(X, Y)
# selected features
print('Number of features:', X.shape[1])
print('Reduced number of features:',X_best_feat.shape[1])
# +
#Recursive Feature Elimination
from sklearn.feature_selection import RFE
from sklearn.linear_model import LogisticRegression
# create a base classifier used to evaluate a subset of attributes
logistic_model = LogisticRegression()
# create the RFE model and select 4 attributes
rfe = RFE(logistic_model, 4)
rfe = rfe.fit(X, Y)
# Ranking of the attributes
print(sorted(zip(map(lambda x: round(x, 4), rfe.ranking_),X)))
# -
# Feature Importance
from sklearn.ensemble import RandomForestClassifier
# fit a RandomForest model to the data
model = RandomForestClassifier()
model.fit(X, Y)
# display the relative importance of each attribute
print(sorted(zip(map(lambda x: round(x, 4), model.feature_importances_),X)))
| Chapter 03/Feature_Selection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
"""Convert files of a selected directory in jpg format"""
import converter
# # !pip install easyocr
import easyocr
#download the model
reader = easyocr.Reader(['en'], gpu = True)
# show an image
import PIL
from PIL import ImageDraw
from PIL import Image
import cv2
import PIL
from PIL import ImageDraw
from PIL import Image
import cv2
import pandas as pd
from pandas import DataFrame
import pandas as pd
import json
import glob
# import xlrd
import csv
# import os
# print(os.getcwd())
# # path = '/Users/neerajyadav/Documents/pycv/PICK-pytorch/'
# path = '/Volumes/Extreme SSD/MLWork/DocAI/PICK-pytorch'
# os.chdir(path)
# import argparse
# import torch
# from tqdm import tqdm
# from pathlib import Path
# from torch.utils.data.dataloader import DataLoader
# from allennlp.data.dataset_readers.dataset_utils.span_utils import bio_tags_to_spans
# from parse_config import ConfigParser
# import model.pick as pick_arch_module
# from data_utils.pick_dataset import PICKDataset
# from data_utils.pick_dataset import BatchCollateFn
# from utils.util import iob_index_to_str, text_index_to_str
# import converter
import shutil, os
### convert image into transcript file
"""Select jpg files and convert into transcript files"""
filenames = glob.glob("/Volumes/Extreme SSD/MLWork/DocAI/TestImage/*.jpg")
filenamesj = glob.glob("/Volumes/Extreme SSD/MLWork/DocAI/TestImage/*.jpeg")
filenames = filenames + filenamesj
filenames.sort()
def draw_boxes(image, bounds, color='green', width=1):
draw = ImageDraw.Draw(image)
for bound in bounds:
p0, p1, p2, p3 = bound[0]
draw.line([*p0, *p1, *p2, *p3, *p0], fill=color , width=width)
# if bound[1] == "ToTAL" or bound[1] =="TOTAL" or bound[1]=="TOTAL" or bound[1] =="Total Payable;" or bound[1] =="Total Payable:" or bound[1] =="Total Payable:" or bound[1]=='Total' or bound[1]=='TOTAL' or bound[1]=="Totz' Ingi, 0f GST" or bound[1]=="Total Sales (Inclusive of GST)" or bound[1]=="Net Total (MYR)":
# draw.line([*p0, *p1, *p2, *p3, *p0], fill=color, width=width)
# print(bound[0])
# image.save("temp.jpg")
return image
# draw_boxes(im, bounds)
def concatenate_list_data(list):
result= ''
for element in list:
result = result +str(element)
return result
for s in filenames:
# s = "Invoice0.jpg"
filen = s.split(".")[0]
print(filen)
im = PIL.Image.open(s).convert('RGB')
# Doing OCR. Get bounding boxes.
bounds = reader.readtext(s)
im = PIL.Image.open(s).convert('RGB')
df = pd.DataFrame()
CoordinatesValue = []
for i in bounds:
Coordinates =[]
CoordinatesValue=[]
temp_df = pd.DataFrame()
Coordinates.append(concatenate_list_data(i[0]).replace("][",",").replace("[","").replace("]","").replace(" ",""))
# print(i[1])
CoordinatesValue.append(i[1])
temp_df = DataFrame(zip(Coordinates,CoordinatesValue),columns = ['Coordinates', 'Value'])
# print(temp_df)
df = df.append(temp_df)
# print(item[0])
combine_lambda = lambda x: '{},{}'.format(x.Coordinates, x.Value)
df['Result'] = df.apply(combine_lambda, axis = 1)
dfnew= df['Result']
dfnew = dfnew[0].str.split(',', expand=True)
dfnew.insert(0,'name_of_column','')
dfnew['name_of_column'] = 1
# dfnew.to_csv(str(filen)+".tsv", sep = ',',index=False ,header=False )
dfnew.to_csv(str(filen)+".tsv",sep = ',',index=False,header=False, quotechar='',escapechar='\\',quoting=csv.QUOTE_NONE, )
# ### copy file from source folder to destination folder ###
for f in filenames:
shutil.copy(f, 'test_img/')
filetsv = glob.glob("/Volumes/Extreme SSD/MLWork/DocAI/TestImage/*.tsv")
for f in filetsv:
shutil.copy(f, 'test_boxes_and_transcripts/')
# -
import os
print(os.getcwd())
# path = '/Users/neerajyadav/Documents/pycv/PICK-pytorch/'
path = '/Volumes/Extreme SSD/MLWork/DocAI/PICK-pytorch'
os.chdir(path)
import argparse
import torch
from tqdm import tqdm
from pathlib import Path
from torch.utils.data.dataloader import DataLoader
from allennlp.data.dataset_readers.dataset_utils.span_utils import bio_tags_to_spans
from parse_config import ConfigParser
import model.pick as pick_arch_module
from data_utils.pick_dataset import PICKDataset
from data_utils.pick_dataset import BatchCollateFn
from utils.util import iob_index_to_str, text_index_to_str
import converter
| inferenceNotebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Programming Exercise 6:
# # Support Vector Machines
# ## Introduction
# In this exercise, you will be using support vector machines (SVMs) to build a spam classifier. Before starting on the programming exercise, we strongly recommend watching the video lectures and completing the review questions for the associated topics.
# +
# used for manipulating directory paths
import os
# Scientific and vector computation for python
import numpy as np
# Import regular expressions to process emails
import re
# Plotting library
from matplotlib import pyplot
# Optimization module in scipy
from scipy import optimize
# will be used to load MATLAB mat datafile format
from scipy.io import loadmat
# tells matplotlib to embed plots within the notebook
# %matplotlib inline
# -
# ## 1 Support Vector Machines
# In the first half of this exercise, you will be using support vector machines (SVMs) with various example 2D datasets. Experimenting with these datasets will help you gain an intuition of how SVMs work and how to use a Gaussian kernel with SVMs. In the next half of the exercise, you will be using support vector machines to build a spam classifier.
# ### 1.1 Example Dataset 1
# In this dataset, the positions of the positive examples (indicated with x) and the negative examples (indicated with o) suggest a natural separation indicated by the gap. However, notice that there is an outlier positive example x on the far left at about (0.1, 4.1). As part of this exercise, you will also see how this outlier affects the SVM decision boundary.
def plotData(X, y, grid=False):
"""
Plots the data points X and y into a new figure. Uses `+` for positive examples, and `o` for
negative examples. `X` is assumed to be a Mx2 matrix
Parameters
----------
X : numpy ndarray
X is assumed to be a Mx2 matrix.
y : numpy ndarray
The data labels.
grid : bool (Optional)
Specify whether or not to show the grid in the plot. It is False by default.
Notes
-----
This was slightly modified such that it expects y=1 or y=0.
"""
# Find Indices of Positive and Negative Examples
pos = y == 1
neg = y == 0
# Plot Examples
pyplot.plot(X[pos, 0], X[pos, 1], 'X', mew=1, ms=10, mec='k')
pyplot.plot(X[neg, 0], X[neg, 1], 'o', mew=1, mfc='y', ms=10, mec='k')
pyplot.grid(grid)
# +
# Load from ex6data1
# You will have X, y as keys in the dict data
data = loadmat(os.path.join('Data', 'ex6data1.mat'))
X, y = data['X'], data['y'][:, 0]
# Plot training data
plotData(X, y)
# -
# In this part of the exercise, you will try using different values of the $C$ parameter with SVMs. Informally, the $C$ parameter is a positive value that controls the penalty for misclassified training examples. A large $C$ parameter tells the SVM to try to classify all the examples correctly. $C$ plays a role similar to $1/\lambda$, where $\lambda$ is the regularization parameter that we were using previously for logistic regression.
#
# The following cell will run the SVM training (with $C=1$) using SVM software that we have included with the starter code (function svmTrain of this exercise). When $C=1$, you should find that the SVM puts the decision boundary in the gap between the two datasets and misclassifies the data point on the far left, as shown in the figure below.
def svmTrain(X, Y, C, kernelFunction, tol=1e-3, max_passes=5, args=()):
"""
Trains an SVM classifier using a simplified version of the SMO algorithm.
Parameters
---------
X : numpy ndarray
(m x n) Matrix of training examples. Each row is a training example, and the
jth column holds the jth feature.
Y : numpy ndarray
(m, ) A vector (1-D numpy array) containing 1 for positive examples and 0 for negative examples.
C : float
The standard SVM regularization parameter.
kernelFunction : func
A function handle which computes the kernel. The function should accept two vectors as
inputs, and returns a scalar as output.
tol : float, optional
Tolerance value used for determining equality of floating point numbers.
max_passes : int, optional
Controls the number of iterations over the dataset (without changes to alpha)
before the algorithm quits.
args : tuple
Extra arguments required for the kernel function, such as the sigma parameter for a
Gaussian kernel.
Returns
-------
model :
The trained SVM model.
Notes
-----
This is a simplified version of the SMO algorithm for training SVMs. In practice, if
you want to train an SVM classifier, we recommend using an optimized package such as:
- LIBSVM (http://www.csie.ntu.edu.tw/~cjlin/libsvm/)
- SVMLight (http://svmlight.joachims.org/)
- scikit-learn (http://scikit-learn.org/stable/modules/svm.html) which contains python wrappers
for the LIBSVM library.
"""
# make sure data is signed int
Y = Y.astype(int)
# Dataset size parameters
m, n = X.shape
passes = 0
E = np.zeros(m)
alphas = np.zeros(m)
b = 0
# Map 0 to -1
Y[Y == 0] = -1
# Pre-compute the Kernel Matrix since our dataset is small
# (in practice, optimized SVM packages that handle large datasets
# gracefully will **not** do this)
# We have implemented the optimized vectorized version of the Kernels here so
# that the SVM training will run faster
if kernelFunction.__name__ == 'linearKernel':
# Vectorized computation for the linear kernel
# This is equivalent to computing the kernel on every pair of examples
K = np.dot(X, X.T)
elif kernelFunction.__name__ == 'gaussianKernel':
# vectorized RBF Kernel
# This is equivalent to computing the kernel on every pair of examples
X2 = np.sum(X**2, axis=1)
K = X2 + X2[:, None] - 2 * np.dot(X, X.T)
if len(args) > 0:
K /= 2*args[0]**2
K = np.exp(-K)
else:
K = np.zeros((m, m))
for i in range(m):
for j in range(i, m):
K[i, j] = kernelFunction(X[i, :], X[j, :])
K[j, i] = K[i, j]
while passes < max_passes:
num_changed_alphas = 0
for i in range(m):
E[i] = b + np.sum(alphas * Y * K[:, i]) - Y[i]
if (Y[i]*E[i] < -tol and alphas[i] < C) or (Y[i]*E[i] > tol and alphas[i] > 0):
# select the alpha_j randomly
j = np.random.choice(list(range(i)) + list(range(i+1, m)), size=1)[0]
E[j] = b + np.sum(alphas * Y * K[:, j]) - Y[j]
alpha_i_old = alphas[i]
alpha_j_old = alphas[j]
if Y[i] == Y[j]:
L = max(0, alphas[j] + alphas[i] - C)
H = min(C, alphas[j] + alphas[i])
else:
L = max(0, alphas[j] - alphas[i])
H = min(C, C + alphas[j] - alphas[i])
if L == H:
continue
eta = 2 * K[i, j] - K[i, i] - K[j, j]
# objective function positive definite, there will be a minimum along the direction
# of linear equality constrain, and eta will be greater than zero
# we are actually computing -eta here (so we skip of eta >= 0)
if eta >= 0:
continue
alphas[j] -= Y[j] * (E[i] - E[j])/eta
alphas[j] = max(L, min(H, alphas[j]))
if abs(alphas[j] - alpha_j_old) < tol:
alphas[j] = alpha_j_old
continue
alphas[i] += Y[i]*Y[j]*(alpha_j_old - alphas[j])
b1 = b - E[i] - Y[i]*(alphas[i] - alpha_i_old) * K[i, j] \
- Y[j] * (alphas[j] - alpha_j_old) * K[i, j]
b2 = b - E[j] - Y[i]*(alphas[i] - alpha_i_old) * K[i, j] \
- Y[j] * (alphas[j] - alpha_j_old) * K[j, j]
if 0 < alphas[i] < C:
b = b1
elif 0 < alphas[j] < C:
b = b2
else:
b = (b1 + b2)/2
num_changed_alphas += 1
if num_changed_alphas == 0:
passes += 1
else:
passes = 0
idx = alphas > 0
model = {'X': X[idx, :],
'y': Y[idx],
'kernelFunction': kernelFunction,
'b': b,
'args': args,
'alphas': alphas[idx],
'w': np.dot(alphas * Y, X)}
return model
def svmPredict(model, X):
"""
Returns a vector of predictions using a trained SVM model.
Parameters
----------
model : dict
The parameters of the trained svm model, as returned by the function svmTrain
X : array_like
A (m x n) matrix where each example is a row.
Returns
-------
pred : array_like
A (m,) sized vector of predictions {0, 1} values.
"""
# check if we are getting a vector. If so, then assume we only need to do predictions
# for a single example
if X.ndim == 1:
X = X[np.newaxis, :]
m = X.shape[0]
p = np.zeros(m)
pred = np.zeros(m)
if model['kernelFunction'].__name__ == 'linearKernel':
# we can use the weights and bias directly if working with the linear kernel
p = np.dot(X, model['w']) + model['b']
elif model['kernelFunction'].__name__ == 'gaussianKernel':
# vectorized RBF Kernel
# This is equivalent to computing the kernel on every pair of examples
X1 = np.sum(X**2, 1)
X2 = np.sum(model['X']**2, 1)
K = X2 + X1[:, None] - 2 * np.dot(X, model['X'].T)
if len(model['args']) > 0:
K /= 2*model['args'][0]**2
K = np.exp(-K)
p = np.dot(K, model['alphas']*model['y']) + model['b']
else:
# other non-linear kernel
for i in range(m):
predictions = 0
for j in range(model['X'].shape[0]):
predictions += model['alphas'][j] * model['y'][j] \
* model['kernelFunction'](X[i, :], model['X'][j, :])
p[i] = predictions
pred[p >= 0] = 1
return pred
def linearKernel(x1, x2):
"""
Returns a linear kernel between x1 and x2.
Parameters
----------
x1 : numpy ndarray
A 1-D vector.
x2 : numpy ndarray
A 1-D vector of same size as x1.
Returns
-------
: float
The scalar amplitude.
"""
return np.dot(x1, x2)
def visualizeBoundaryLinear(X, y, model):
"""
Plots a linear decision boundary learned by the SVM.
Parameters
----------
X : array_like
(m x 2) The training data with two features (to plot in a 2-D plane).
y : array_like
(m, ) The data labels.
model : dict
Dictionary of model variables learned by SVM.
"""
w, b = model['w'], model['b']
xp = np.linspace(min(X[:, 0]), max(X[:, 0]), 100)
yp = -(w[0] * xp + b)/w[1]
plotData(X, y)
pyplot.plot(xp, yp, '-b')
# +
C = 1
model = svmTrain(X, y, C, linearKernel, 1e-3, 20)
visualizeBoundaryLinear(X, y, model)
# -
# Your task is to try different values of $C$ on this dataset. Specifically, you should change the value of $C$ in the next cell to $C = 100$ and run the SVM training again. When $C = 100$, you should find that the SVM now classifies every single example correctly, but has a decision boundary that does not appear to be a natural fit for the data.
# +
C = 100
model = svmTrain(X, y, C, linearKernel, 1e-3, 20)
visualizeBoundaryLinear(X, y, model)
# -
# ### 1.2 SVM with Gaussian Kernels
# In this part of the exercise, you will be using SVMs to do non-linear classification. In particular, you will be using SVMs with Gaussian kernels on datasets that are not linearly separable.
# #### 1.2.1 Gaussian Kernel
# To find non-linear decision boundaries with the SVM, we need to first implement a Gaussian kernel. You can think of the Gaussian kernel as a similarity function that measures the “distance” between a pair of examples, ($x^{(i)}$, $x^{(j)}$). The Gaussian kernel is also parameterized by a bandwidth parameter, $\sigma$, which determines how fast the similarity metric decreases (to 0) as the examples are further apart. You should now complete the code in gaussianKernel to compute the Gaussian kernel between two examples, ($x^{(i)}$, $x^{(j)}$). The Gaussian kernel function is defined as:
#
# $$ K_{\text{gaussian}} \left( x^{(i)}, x^{(j)} \right) = \exp \left( - \frac{\left\lvert\left\lvert x^{(i)} - x^{(j)}\right\lvert\right\lvert^2}{2\sigma^2} \right) = \exp \left( -\frac{\sum_{k=1}^n \left( x_k^{(i)} - x_k^{(j)}\right)^2}{2\sigma^2} \right)$$
def gaussianKernel(x1, x2, sigma):
"""
Computes the radial basis function
Returns a radial basis function kernel between x1 and x2.
Parameters
----------
x1 : numpy ndarray
A vector of size (n, ), representing the first datapoint.
x2 : numpy ndarray
A vector of size (n, ), representing the second datapoint.
sigma : float
The bandwidth parameter for the Gaussian kernel.
Returns
-------
sim : float
The computed RBF between the two provided data points.
Instructions
------------
Fill in this function to return the similarity between `x1` and `x2`
computed using a Gaussian kernel with bandwidth `sigma`.
"""
sim = 0
# ====================== YOUR CODE HERE ======================
sim = np.exp(-np.sum((x1 - x2)**2) / (2 * sigma**2))
# =============================================================
return sim
# Once you have completed the function gaussianKernel the following cell will test your kernel function on two provided examples and you should expect to see a value of 0.324652.
# +
x1 = np.array([1, 2, 1])
x2 = np.array([0, 4, -1])
sigma = 2
sim = gaussianKernel(x1, x2, sigma)
print('Gaussian Kernel between x1 = [1, 2, 1], x2 = [0, 4, -1], sigma = %0.2f:'
'\n\t%f\n(for sigma = 2, this value should be about 0.324652)\n' % (sigma, sim))
# -
# #### 1.2.2 Example Dataset 2
# The next part in this notebook will load and plot dataset 2, as shown in the figure below.
# +
# Load from ex6data2
# You will have X, y as keys in the dict data
data = loadmat(os.path.join('Data', 'ex6data2.mat'))
X, y = data['X'], data['y'][:, 0]
# Plot training data
plotData(X, y)
# -
# From the figure, you can obserse that there is no linear decision boundary that separates the positive and negative examples for this dataset. However, by using the Gaussian kernel with the SVM, you will be able to learn a non-linear decision boundary that can perform reasonably well for the dataset. If you have correctly implemented the Gaussian kernel function, the following cell will proceed to train the SVM with the Gaussian kernel on this dataset.
#
# You should get a decision boundary as shown in the figure below, as computed by the SVM with a Gaussian kernel. The decision boundary is able to separate most of the positive and negative examples correctly and follows the contours of the dataset well.
def visualizeBoundary(X, y, model):
"""
Plots a non-linear decision boundary learned by the SVM and overlays the data on it.
Parameters
----------
X : array_like
(m x 2) The training data with two features (to plot in a 2-D plane).
y : array_like
(m, ) The data labels.
model : dict
Dictionary of model variables learned by SVM.
"""
plotData(X, y)
# make classification predictions over a grid of values
x1plot = np.linspace(min(X[:, 0]), max(X[:, 0]), 100)
x2plot = np.linspace(min(X[:, 1]), max(X[:, 1]), 100)
X1, X2 = np.meshgrid(x1plot, x2plot)
vals = np.zeros(X1.shape)
for i in range(X1.shape[1]):
this_X = np.stack((X1[:, i], X2[:, i]), axis=1)
vals[:, i] = svmPredict(model, this_X)
pyplot.contour(X1, X2, vals, colors='y', linewidths=2)
pyplot.pcolormesh(X1, X2, vals, cmap='YlGnBu', alpha=0.25, edgecolors='None', lw=0)
pyplot.grid(False)
# +
# SVM Parameters
C = 1
sigma = 0.1
model= svmTrain(X, y, C, gaussianKernel, args=(sigma,))
visualizeBoundary(X, y, model)
# -
# #### 1.2.3 Example Dataset 3
# In this part of the exercise, you will gain more practical skills on how to use a SVM with a Gaussian kernel. The next cell will load and display a third dataset, which should look like the figure below.
#
# You will be using the SVM with the Gaussian kernel with this dataset. In the provided dataset, ex6data3.mat, you are given the variables X, y, Xval, yval
# +
# Load from ex6data3
# You will have X, y, Xval, yval as keys in the dict data
data = loadmat(os.path.join('Data', 'ex6data3.mat'))
X, y, Xval, yval = data['X'], data['y'][:, 0], data['Xval'], data['yval'][:, 0]
# Plot training data
plotData(X, y)
# -
# Your task is to use the cross validation set Xval, yval to determine the best $C$ and $\sigma$ parameter to use. You should write any additional code necessary to help you search over the parameters $C$ and $\sigma$. For both $C$ and $\sigma$, we suggest trying values in multiplicative steps (e.g., 0.01, 0.03, 0.1, 0.3, 1, 3, 10, 30). Note that you should try all possible pairs of values for $C$ and $\sigma$ (e.g., $C = 0.3$ and $\sigma = 0.1$). For example, if you try each of the 8 values listed above for $C$ and for $\sigma^2$, you would end up training and evaluating (on the cross validation set) a total of $8^2 = 64$ different models. After you have determined the best $C$ and $\sigma$ parameters to use, you should modify the code in dataset3Params, filling in the best parameters you found. For our best parameters, the SVM returned a decision boundary shown in the figure below.
def dataset3Params(X, y, Xval, yval):
"""
Returns your choice of C and sigma for Part 3 of the exercise
where you select the optimal (C, sigma) learning parameters to use for SVM
with RBF kernel.
Parameters
----------
X : array_like
(m x n) matrix of training data where m is number of training examples, and
n is the number of features.
y : array_like
(m, ) vector of labels for ther training data.
Xval : array_like
(mv x n) matrix of validation data where mv is the number of validation examples
and n is the number of features
yval : array_like
(mv, ) vector of labels for the validation data.
Returns
-------
C, sigma : float, float
The best performing values for the regularization parameter C and
RBF parameter sigma.
Instructions
------------
Fill in this function to return the optimal C and sigma learning
parameters found using the cross validation set.
You can use `svmPredict` to predict the labels on the cross
validation set. For example,
predictions = svmPredict(model, Xval)
will return the predictions on the cross validation set.
Note
----
You can compute the prediction error using
np.mean(predictions != yval)
"""
# You need to return the following variables correctly.
C = 1
sigma = 0.3
# ====================== YOUR CODE HERE ======================
vals = [0.01, 0.03, 0.1, 0.3, 1, 3, 10, 30]
result = []
for C in(vals):
for sigma in(vals):
model = svmTrain(X, y, C, gaussianKernel, args=(sigma,))
predictions = svmPredict(model, Xval)
error = np.mean(predictions != yval)
result.append([C, sigma, error])
# ============================================================
return min(result, key=lambda x: x[2])
# The provided code in the next cell trains the SVM classifier using the training set $(X, y)$ using parameters loaded from dataset3Params. Note that this might take a few minutes to execute.
# +
# Try different SVM Parameters here
result = dataset3Params(X, y, Xval, yval)
C, sigma = result[0], result[1]
# Train the SVM
# model = utils.svmTrain(X, y, C, lambda x1, x2: gaussianKernel(x1, x2, sigma))
model = svmTrain(X, y, C, gaussianKernel, args=(sigma,))
visualizeBoundary(X, y, model)
print(C, sigma)
# -
| exercise6/exercise6-SVM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/raviakasapu/MachineLearning/blob/main/ML_Random_test_sample_generators_sklearn.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="bDAYFrhf0h2y"
# # Generate Test data using Pandas & Numpy
# + colab={"base_uri": "https://localhost:8080/"} id="xcrFLfWa0osv" outputId="50c602ae-38aa-4866-e3a3-ded7d0c2bf35"
import numpy as np
import pandas as pd
#create a list of cities
cities = ['Delhi', 'Bangalore','Hyderabad','Berlin','NewYork','London','Tokyo']
n = len(cities)
n
# + colab={"base_uri": "https://localhost:8080/"} id="kAK0i85x72c4" outputId="75c2ec33-7fb2-44eb-c383-b22f82479c1f"
data = {'Temparature':np.random.normal(24,3,n),
'Humidity': np.random.normal(78,2.5,n),
'Wind': np.random.normal(15,4,n)
}
data['Wind']
# + colab={"base_uri": "https://localhost:8080/", "height": 266} id="-5rut7328b0q" outputId="734ff456-57b4-4e66-bb6a-8d62f28c559a"
df = pd.DataFrame(data= data, index=cities)
df
# + [markdown] id="3SGLxu8h-2lh"
# ## Test data for Classificiation and Clustering
# + [markdown] id="Jf9noarRCFD0"
# make_blobs will create a set of clusters to be used in ML
# + id="xNg7TMnQ-1sL"
import numpy as np
import matplotlib.pyplot as plt
from sklearn.datasets import make_blobs
# + id="qr7q5f5tCOZN"
no_classes = 5
data, labels = make_blobs(n_samples = 1000,
centers = no_classes,
random_state = 10)
# + colab={"base_uri": "https://localhost:8080/", "height": 312} id="mwmE-ceKCvTm" outputId="88164cd4-6a29-45aa-f144-7fae86def2ec"
fig, ax = plt.subplots()
colours = ['green','orange','blue','pink','red']
for label in range(no_classes):
ax.scatter(x=data[labels==label, 0],
y=data[labels==label, 1],
c=colours[label],
s=40,
label=label
)
ax.set(xlabel='X',
ylabel='Y',
title = "Random Blobs for Clustering")
ax.legend(loc='upper right')
# + [markdown] id="Qs--NPipGgv6"
# make_blobs with predefined centers
# + id="A3uK1y0jGb-q"
centers = [[12,3],[4,17],[9,10],[13,9],[3,6]]
data, labels = make_blobs(n_samples = 1000,
centers = np.array(centers),
random_state = 1)
# + colab={"base_uri": "https://localhost:8080/", "height": 312} id="6IvajVrqG8eN" outputId="49aa0053-eb44-4aa7-ab8c-54e4c679ac2f"
fig, ax = plt.subplots()
colours = ['green','orange','blue','pink','red']
for label in range(len(centers)):
ax.scatter(x=data[labels==label, 0],
y=data[labels==label, 1],
c=colours[label],
s=40,
label=label
)
ax.set(xlabel='X',
ylabel='Y',
title = "Random Blobs for Clustering")
ax.legend(loc='upper right')
# + colab={"base_uri": "https://localhost:8080/"} id="BMoYE2ooI7s4" outputId="8307f795-0ea2-475d-e9b5-fd5b3fe15762"
labels = labels.reshape((labels.shape[0],1))
xp_data = np.concatenate((data,labels), axis=1)
xp_data[:10]
# + id="blW-Af5bJeXD"
np.savetxt("final_data.txt",
xp_data)
# + [markdown] id="OWetOQdEMQSL"
# make_moons will create a set of clusters to be used in ML
# + colab={"base_uri": "https://localhost:8080/", "height": 312} id="kALlij_aMXea" outputId="fedaa9bd-b713-48ff-a191-dc5b6226a985"
import sklearn.datasets as ds
data, labels = ds.make_circles(n_samples=1000,
noise=0.1,
random_state=1)
fig, ax = plt.subplots()
colours = ['green','orange','blue','pink','red']
for label in range(no_classes):
ax.scatter(x=data[labels==label, 0],
y=data[labels==label, 1],
c=colours[label],
s=40,
label=label
)
ax.set(xlabel='X',
ylabel='Y',
title = "Make Swiss Roll")
ax.legend(loc='upper right')
| ML_Random_test_sample_generators_sklearn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: bentoml-dev-py36
# language: python
# name: bentoml-dev-py36
# ---
# + [markdown] colab_type="text" id="jYysdyb-CaWM"
# # Basic classification: Classify images of clothing
# A tensorflow serving style service example using BentoML
#
#
# 
# +
# %reload_ext autoreload
# %autoreload 2
# %matplotlib inline
# add venv PATH to shell command PATH
import sys, os
if sys.base_prefix not in os.environ['PATH']:
os.environ['PATH'] = f"{sys.base_prefix}/bin:{os.environ['PATH']}"
# + colab={} colab_type="code" id="dzLKpmZICaWN"
from __future__ import absolute_import, division, print_function, unicode_literals
import io
# TensorFlow
import tensorflow as tf
# Helper libraries
import numpy as np
import matplotlib.pyplot as plt
print(tf.__version__)
# + colab={} colab_type="code" id="7MqDQO0KCaWS"
fashion_mnist = tf.keras.datasets.fashion_mnist
(_train_images, train_labels), (_test_images, test_labels) = fashion_mnist.load_data()
class_names = ['T-shirt/top', 'Trouser', 'Pullover', 'Dress', 'Coat',
'Sandal', 'Shirt', 'Sneaker', 'Bag', 'Ankle boot']
train_images = _train_images / 255.0
test_images = _test_images / 255.0
# -
class FashionMnist(tf.keras.Model):
def __init__(self):
super(FashionMnist, self).__init__()
self.cnn = tf.keras.Sequential([
tf.keras.layers.Flatten(input_shape=(28, 28)),
tf.keras.layers.Dense(128, activation='relu'),
tf.keras.layers.Dense(10, activation='softmax')
])
@staticmethod
def image_bytes2tensor(inputs):
inputs = tf.map_fn(lambda i: tf.io.decode_png(i, channels=1), inputs, dtype=tf.uint8)
inputs = tf.cast(inputs, tf.float32)
inputs = (255.0 - inputs) / 255.0
inputs = tf.reshape(inputs, [-1, 28, 28])
return inputs
@tf.function(input_signature=[tf.TensorSpec(shape=(None,), dtype=tf.string)])
def predict_image(self, inputs):
inputs = self.image_bytes2tensor(inputs)
return self(inputs)
def call(self, inputs):
return self.cnn(inputs)
# ## test the image preprocessing
# +
# pick up a test image
d_test_img = _test_images[0]
print(class_names[test_labels[0]])
plt.imshow(255.0 - d_test_img, cmap='gray')
plt.imsave("test.png", 255.0 - d_test_img, cmap='gray')
# read bytes
with open("test.png", "rb") as f:
img_bytes = f.read()
# + [markdown] colab={} colab_type="code" id="9ODch-OFCaW4"
# ## train the model
#
# + colab={} colab_type="code" id="Lhan11blCaW7"
model = FashionMnist()
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
model.fit(train_images, train_labels, epochs=1)
# -
# ## test the model
predict = model.predict_image(tf.constant([img_bytes] * 3))
klass = tf.argmax(predict, axis=1)
[class_names[k] for k in klass]
# + [markdown] colab_type="text" id="YFc2HbEVCaXd"
# And the model predicts a label as expected.
# -
# # Define & save BentoService
# +
# %%writefile tensorflow_fashion_mnist.py
import base64
import bentoml
import tensorflow as tf
import numpy as np
from PIL import Image
from bentoml.artifact import (
TensorflowSavedModelArtifact,
)
from bentoml.handlers import TensorflowTensorHandler, ClipperStringsHandler
FASHION_MNIST_CLASSES = ['T-shirt/top', 'Trouser', 'Pullover', 'Dress', 'Coat',
'Sandal', 'Shirt', 'Sneaker', 'Bag', 'Ankle boot']
@bentoml.env(pip_dependencies=['tensorflow', 'numpy', 'pillow'])
@bentoml.artifacts([TensorflowSavedModelArtifact('model')])
class FashionMnistTensorflow(bentoml.BentoService):
@bentoml.api(TensorflowTensorHandler)
def predict(self, inputs):
outputs = self.artifacts.model.predict_image(inputs)
output_classes = tf.math.argmax(outputs, axis=1)
return [FASHION_MNIST_CLASSES[o] for o in output_classes]
@bentoml.api(ClipperStringsHandler)
def predict_clipper(self, strings):
_bytes = [base64.b64decode(i) for i in strings]
inputs = tf.constant(_bytes, dtype=tf.string)
outputs = self.artifacts.model.predict_image(inputs)
output_classes = tf.math.argmax(outputs, axis=1)
return [FASHION_MNIST_CLASSES[o] for o in output_classes]
# +
from tensorflow_fashion_mnist import FashionMnistTensorflow
bento_svc = FashionMnistTensorflow()
bento_svc.pack("model", model)
saved_path = bento_svc.save()
# -
# # Build & Run Clipper Service in Docker
from clipper_admin import ClipperConnection, DockerContainerManager
from bentoml.clipper import deploy_bentoml
cl = ClipperConnection(DockerContainerManager())
try:
cl.start_clipper()
except:
cl.connect()
NAME = saved_path.split('/')[-1].lower()
cl.register_application(NAME, 'strings', 'default_pred', 300000)
# replace PIP_INDEX_URL with your prefer pypi mirror
clipper_model_name, clipper_model_version = deploy_bentoml(
cl, saved_path, 'predict_clipper',
build_envs=dict(
PIP_INDEX_URL="http://192.168.138.2/simple",
PIP_TRUSTED_HOST="192.168.138.2",
)
)
cl.link_model_to_app(NAME, clipper_model_name)
addr = cl.get_query_addr()
clipper_url = f"http://{addr}/{NAME}/predict"
# # Test with requests
# +
import base64
import json
import requests
with open("test.png", "rb") as f:
img_bytes = f.read()
img_b64 = base64.b64encode(img_bytes).decode()
headers = {"content-type": "application/json"}
data = json.dumps(
{"input": img_b64}
)
json_response = requests.post(clipper_url, data=data, headers=headers)
print(json_response)
print(json_response.text)
# -
# # Benchmark with locust
# +
# %%writefile benchmark_clipper.py
from locust import HttpLocust, TaskSet, task, constant
from functools import lru_cache
import math
import random
import numpy as np
import pandas as pd
import json
import base64
import requests
@lru_cache(maxsize=1)
def data_producer():
with open("test.png", "rb") as f:
img_bytes = f.read()
img_b64 = base64.b64encode(img_bytes).decode()
def _gen_data(size=3):
headers = {"content-type": "application/json"}
data = json.dumps(
{"input": img_b64}
)
return headers, data
return _gen_data
class WebsiteTasks(TaskSet):
@staticmethod
def get_data():
headers, data = data_producer()(3)
return headers, data
@task
def index(self):
headers, data = self.get_data()
self.client.post("/predict", data, headers=headers)
class WebsiteUser(HttpLocust):
task_set = WebsiteTasks
wait_time = constant(1)
# -
# !locust -f benchmark_clipper.py -H {clipper_url} -c 10 --step-clients 10
| benchmark/tensorflow_2_fashion_mnist/tensorflow_2_fashion_mnist-clipper.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
sys.path.append('../scripts/')
from ideal_robot import *
from scipy.stats import expon, norm
class Robot(IdealRobot): ###add_stuck### noise, biasメソッドは省略で
def __init__(self, pose, agent=None, sensor=None, color="black", \
noise_per_meter=5, noise_std=math.pi/60,\
bias_rate_stds=(0.1,0.1),\
expected_stuck_time = 1e100, expected_escape_time = 1e-100): #追加
super().__init__(pose, agent, sensor, color)
self.noise_pdf = expon(scale=1.0/(1e-100 + noise_per_meter))
self.distance_until_noise = self.noise_pdf.rvs()
self.theta_noise = norm(scale=noise_std)
self.bias_rate_nu = norm.rvs(loc=1.0, scale=bias_rate_stds[0])
self.bias_rate_omega = norm.rvs(loc=1.0, scale=bias_rate_stds[1])
self.stuck_pdf = expon(scale=expected_stuck_time) #以下追加
self.escape_pdf = expon(scale=expected_escape_time)
self.time_until_stuck = self.stuck_pdf.rvs()
self.time_until_escape = self.escape_pdf.rvs()
self.is_stuck = False
def noise(self, pose, nu, omega, time_interval):
self.distance_until_noise -= abs(nu)*time_interval + self.r*omega*time_interval
if self.distance_until_noise <= 0.0:
self.distance_until_noise += self.noise_pdf.rvs()
pose[2] += self.theta_noise.rvs()
return pose
def bias(self, nu, omega):
return nu*self.bias_rate_nu, omega*self.bias_rate_omega
def stuck(self, nu, omega, time_interval): #追加
if self.is_stuck:
self.time_until_escape -= time_interval
if self.time_until_escape <= 0.0:
self.time_until_escape += self.escape_pdf.rvs()
self.is_stuck = False
else:
self.time_until_stuck -= time_interval
if self.time_until_stuck <= 0.0:
self.time_until_stuck += self.stuck_pdf.rvs()
self.is_stuck = True
return nu*(not self.is_stuck), omega*(not self.is_stuck)
def one_step(self, time_interval):
if not self.agent: return
obs =self.sensor.data(self.pose) if self.sensor else None
nu, omega = self.agent.decision(obs)
nu, omega = self.bias(nu, omega)
nu, omega = self.stuck(nu, omega, time_interval) #追加
self.pose = self.state_transition(nu, omega, time_interval, self.pose)
self.pose = self.noise(self.pose, nu, omega, time_interval)
if self.sensor: self.sensor.data(self.pose)
# +
world = World(30, 0.1) ###stuck_simulation###
circling = Agent(0.2, 10.0/180*math.pi)
for i in range(100):
r = Robot( np.array([0, 0, 0]).T, sensor=None, agent=circling, color="gray", \
noise_per_meter=0, bias_rate_stds=(0.0,0.0), \
expected_stuck_time=60.0, expected_escape_time=60.0)
world.append(r)
r = IdealRobot( np.array([0, 0, 0]).T, sensor=None, agent=circling, color="red" )
world.append(r)
world.draw()
# -
| section_uncertainty/noise_simulation4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import string
import zeyrek
#Easier reading
pd.set_option('display.max_colwidth',None)
#Twitter dataset (one month)
#Getting input as the name of the file.
month_name = input("Please enter filename (e.g:jan20.csv): ")
twit_jan20_DF = pd.read_csv(month_name, sep=";", header=None)
#Transpose
twit_jan20_DFT = twit_jan20_DF.T
#twit_jan20_DFT.describe()
#ANEW Turkish dataset
anew_turkish = pd.read_excel("ANEW_Turkish.xlsx",sheet_name="Sheet1")
#column addition
anew_turkish.columns = ["isimler", "ing-karsiligi", "valence", "arousal", "dominance", "etiketler"]
#lemmatization object
analyzer = zeyrek.MorphAnalyzer()
#Good ol' mean, but does not cry when it is empty
def get_mean(alist):
if len(alist) == 0:
return 0
return sum(alist)/len(alist)
#Ignores the zeros in the list
def get_mean_twit(alist):
modded_length = 0
for i in range(len(alist)):
if alist[i] != 0:
modded_length += 1
if modded_length == 0:
modded_length = 1
return sum(alist)/modded_length
#This function takes a string (a Twit), lemmatize it, then assign statistical value of each word if it exists in ANEW dictionary.
def nlp_operations_VAD(a_twit):
#Getting input twit
string1 = str(a_twit)
#Lowercase it
string1 = string1.lower()
#Split it (or call it tokenizing)
split1 = [i for i in string1.split(' ')]
t1 = []
#This lemmatizes the words in the string
for i in range(len(split1)):
t1.append(analyzer.lemmatize(split1[i]))
#creating lists for valence, arousal and dominance values' mean, so that row of twit can have these values
t1_V, t1_A, t1_D = [], [], []
#iterate over list that holds the lemmatized words
for i in range(len(t1)):
#iterate over lemmatization tuples
for words in t1[i][0]:
#Lemmas can be more than one so getting mean of interpretations could be useful.
word_sum_V, word_sum_A, word_sum_D = [], [], []
#Lemmtization products are in a list, take it by itself
if type(words) == list:
for i in words:
if (anew_turkish["isimler"] == i).any():
word_sum_V.append(anew_turkish.loc[anew_turkish["isimler"] == i]["valence"].iloc[0])
word_sum_A.append(anew_turkish.loc[anew_turkish["isimler"] == i]["arousal"].iloc[0])
word_sum_D.append(anew_turkish.loc[anew_turkish["isimler"] == i]["dominance"].iloc[0])
#Just string, which is the word without lemmazitaion
elif type(words) == str:
if (anew_turkish["isimler"] == i).any():
word_sum_V.append(anew_turkish.loc[anew_turkish["isimler"] == i]["valence"].iloc[0])
word_sum_A.append(anew_turkish.loc[anew_turkish["isimler"] == i]["arousal"].iloc[0])
word_sum_D.append(anew_turkish.loc[anew_turkish["isimler"] == i]["dominance"].iloc[0])
#Bunch up the words in mean to twit part.
t1_V.append(get_mean(word_sum_V))
t1_A.append(get_mean(word_sum_A))
t1_D.append(get_mean(word_sum_D))
return [round(get_mean_twit(t1_V), 2), round(get_mean_twit(t1_A), 2), round(get_mean_twit(t1_D), 2)]
#Some textual preprocessing
df1 = twit_jan20_DFT
#Add Twits column name
twit_jan20_DFT.columns = ["Twits"]
df1['Twits'] = df1['Twits'].astype("string")
df1['Twits'] = df1['Twits'].str.replace('@[A-Za-z0-9]+','')
df1['Twits'] = df1['Twits'].str.replace('http\S+','')
df1['Twits'] = df1['Twits'].str.replace('[^\w\s]+','')
df1['Twits'] = df1['Twits'].str.replace(' +', ' ')
df1["Twits"] = df1["Twits"].str.strip()
df1["Valence"] = None
df1["Arousal"] = None
df1["Dominance"] = None
#Going over the dataset and putting out the V, A, D columns
for idx in range(df1.shape[0]):
#temp_VAD has 3 float numbers in it, returned from nlp_operations_VAD
temp_VAD = nlp_operations_VAD(df1.iloc[idx, 0])
#assigning Valence, Arousal, Dominance
df1.iloc[idx, 1] = temp_VAD[0]
df1.iloc[idx, 2] = temp_VAD[1]
df1.iloc[idx, 3] = temp_VAD[2]
#Output file name operations
x_ = month_name.split(".")
month_p_name = x_[0]+"_p."+x_[1]
df1.to_csv(month_p_name, header=True, sep=';', decimal='.')
print(month_p_name, "is created.")
# -
#df1["Twits"] = df1['Twits'].str.replace(r"\\r|\\n|\\t", "", regex=True)
# +
# +
# -
# +
"""
twit_jan20_DFT["Arousal"]=None
twit_jan20_DFT["Dominance"] = None
twit_jan20_DFT.head(5)
test1df.drop(test1df.columns[-1], axis=1, inplace=True)
#A piece of string
string1 = "abi selam naber"
#Lowercase it
string1 = string1.lower()
#Split it (or call it tokenizing)
split1 = [i for i in string1.split(' ')]
t1 = []
#This lemmatizes the words in the string
for i in range(len(split1)):
t1.append(analyzer.lemmatize(split1[i]))
t1_V, t1_A, t1_D = [], [], []
#iterate over list that holds the lemmatized words
for i in range(len(t1)): #len(t1)
#creating lists for valence, arousal and dominance values' mean, so that row of twit can have these values
#iterate over lemmatization tuples
for words in t1[i][0]:
#Lemmas can be more than one so getting mean of interpretations could be useful.
word_sum_V, word_sum_A, word_sum_D = [], [], []
#Lemmtization products are in a list, take it by itself
if type(words) == list:
for i in words:
if (anew_turkish["isimler"] == i).any():
word_sum_V.append(anew_turkish.loc[anew_turkish["isimler"] == i]["valence"].iloc[0])
word_sum_A.append(anew_turkish.loc[anew_turkish["isimler"] == i]["arousal"].iloc[0])
word_sum_D.append(anew_turkish.loc[anew_turkish["isimler"] == i]["dominance"].iloc[0])
#Just string, which is the word without lemmazitaion
elif type(words) == str:
if (anew_turkish["isimler"] == i).any():
word_sum_V.append(anew_turkish.loc[anew_turkish["isimler"] == i]["valence"].iloc[0])
word_sum_A.append(anew_turkish.loc[anew_turkish["isimler"] == i]["arousal"].iloc[0])
word_sum_D.append(anew_turkish.loc[anew_turkish["isimler"] == i]["dominance"].iloc[0])
#Bunch up the words in mean to twit part.
t1_V.append(get_mean(word_sum_V))
t1_A.append(get_mean(word_sum_A))
t1_D.append(get_mean(word_sum_D))
print(get_mean_twit(t1_V))
print(get_mean_twit(t1_A))
print(get_mean_twit(t1_D))
#print([get_mean_twit(t1_V), get_mean_twit(t1_A), get_mean_twit(t1_D)])
"""
#Removing stuff testing
"""
#Handles
abe = twit_jan20_DFT.iloc[3].astype("string")
print(abe)
abe = abe.str.replace('@[A-Za-z0-9]+', '')
print(abe)
#punctuation testing
abe = twit_jan20_DFT.iloc[3].astype("string")
print(abe)
abe = abe.str.replace('[^\w\s]+','')
print(abe)
#URLs
abe = twit_jan20_DFT.iloc[1].astype("string")
print(abe)
abe = abe.str.replace('http\S+','')
print(abe)
"""
"""
#Adding other columns
twit_jan20_DFT["Valence"] = None
twit_jan20_DFT["Arousal"]=None
twit_jan20_DFT["Dominance"] = None
"""
| codes/senior1-automated.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="FibDZlR2iQD8"
# # BOARD OF IMMIGRATION APPEALS DOCUMENT SCRAPER
#
# • This scraper uses the spaCy library in order to gather information about the case. There is no way to ensure it is 100% accurate, except to scrape a document and compare the information gathered directly with the original PDF.
#
# • This code currently lives inside a Jupyter Notebook for ease of testing and iteration, but will ultimately need to graduate into a standard .py file.
#
# • Scraping is extremely difficult task, as in most cases just searching the document for a keyword isn't enough. It must live in the correct context. No douct the code in this notebook can constantly be improved for a very long time. Before delving into this code, I recommend reading a good chunk of case file PDFs in order to get aquianted with their structre and information.
# + [markdown] id="xS3b78vODefG"
# #### IMPORTS:
# + id="jEsm4q0_S7mn"
# %%capture
# !pip install spacy
# !pip install bs4
# !pip install geonamescache
# + colab={"base_uri": "https://localhost:8080/"} id="m21I3dxQSwhd" outputId="6240ac11-8af2-4024-adba-5688634b9b23"
from typing import List, Tuple, Union, Callable, Dict, Iterator
from collections import defaultdict
from pprint import pprint
from difflib import SequenceMatcher
from datetime import datetime
import bs4
from bs4 import BeautifulSoup, element
import geonamescache
import requests
import pandas as pd
import numpy as np
from pathlib import Path
import re
import os
import spacy
from spacy.tokens.doc import Doc
from spacy.tokens.span import Span
from spacy.tokens.token import Token
spacy.cli.download("en_core_web_lg")
# I currently interface with the text files through my google drive.
# The relevant documents should be able upon request through Slack,
# and ideally should at some point be stored and obtained through a DB
from google.colab import drive
drive.mount('/content/drive')
# + [markdown] id="H_Dy3MWnDjEm"
# #### FILES ARE MOUNTED FROM GOOGLE DRIVE:
# + id="c2cTaSDPb9yK"
ASYLUM_DIR: Path
ASYLUM_DIR = Path("./drive/MyDrive/scraped_asylum_cases")
text_files: List[str]
text_files = os.listdir(ASYLUM_DIR)
def get_text_from(text_file: str, dir: str = ASYLUM_DIR):
with open(dir / text_file) as f:
text = f.read()
return text
ex_text = get_text_from(text_files[1])
# + id="a6vRaYDB0Mib"
# loading in the spaCy model
nlp = spacy.load('en_core_web_lg')
# + [markdown] id="b5MsPRGADDVH"
# #### GATHER BIA JUDGES:
# • The judges that compromise the Board of Immigration Appeals were scraped off of the BIA's wikipedia, which will only show the current judges. This means that in any older PDF files, a judge that is no longer in the BIA is unaccounted for. I could not find a list of previous BIA judges, and focused on these only, for the time being.
# + colab={"base_uri": "https://localhost:8080/", "height": 942} id="Ian7APwuirJU" outputId="9882d3e8-9e18-4ccd-e430-2e9f9b432d03"
# Extracting current Appellate Immigration Judges
# From Wikipedia, using Beautiful Soup. Code is mostly biolerplate
judges_url: str
judges_url = 'https://en.wikipedia.org/wiki/Board_of_Immigration_Appeals'
html: str
html = requests.get(judges_url).text
soup: BeautifulSoup
soup = BeautifulSoup(html, 'html.parser')
table: element.Tag
table = soup.findAll(lambda tag: tag.name =='table')[1]
rows: element.ResultSet
rows = table.findAll(lambda tag: tag.name == 'tr')
column_names: List[str]
column_names = [
col.get_text().strip().lower().replace(' ', '_')
for col in rows.pop(0) if col.name == 'th'
]
rows: List[List[str]]
rows = [
[
cell.get_text().strip().replace(',', '')
for cell in row if cell.name == 'td'
]
for row in rows
]
as_dict: List[Dict[str, str]]
as_dict = [
dict(zip(column_names, row)) for row in rows
]
judges_df: pd.DataFrame
judges_df = pd.DataFrame.from_dict(as_dict)
judges_df
# + [markdown] id="WeN1jtnRkBgs"
# #### UTILITIES:
#
# • The Optical Character Recognition (OCR) used through pytesseract to convert PDFs to text ISN'T perfect. Often artifacts show up, such as the character 'l' shown up as '!', or 'nn' as 'm'.
#
# • The difflib library comes in handy here in order to compare strings by inexact values. The more two strings are alike, the sequence matcher ratio will approach 1, the less they are alike, it will approach 0.
#
# • I recently found a library that could offer a better implementation. It's called FuzzyWuzzy, future teams should look into it.
# + id="bepD85LnBm9x"
# I would like to later reimplement these with the python
# library FuzzyWuzzy, look it up!
def similar(a: str, return_b: str, min_score: float) -> str:
'''
• Returns 2nd string if similarity score is above supplied
minimum score. Else, returns None.
'''
return return_b \
if SequenceMatcher(None, a, return_b).ratio() >= min_score \
else None
# this function implements the similar function, but on a list
# it uses a closure in order to return a function
def similar_in_list(lst: List[str]) -> Callable:
'''
• Uses a closure on supplied list to return a function that iterates over
the list in order to search for the first similar term. It's used widely
in the scraper.
'''
def impl(item: str, min_score: float) -> Union[str, None]:
for s in lst:
s = similar(item, s, min_score)
if s:
return s
return None
return impl
# + [markdown] id="yzJKypoKCleI"
# • In the judge name matching algorithm below, I made a last minute change. Originally is was checking very strictly (First, Last, and any intials must match), but for example, I realized it wasn't picking up the judge Cassidy. I changed the algorithm just to check if the last name is present, which resolved the problem.
# + id="m052dd5O_Dvi"
def get_if_judge(name: str) -> Union[str, None]:
'''
• Returns the judge's name if a match is found. Currently, the match
is very strictly defined by the current judge's names found through
Wikipedia. It will 100% stop any false positives, but some leniency should
be introduced in order to prevent any false negatives.
'''
clean_name: Callable[[str], str]
clean_name = lambda s: s.lower() \
.replace(',', '') \
.replace('.', '')
judges_names = judges_df['name'] \
.apply(lambda s: clean_name(s).split()[-1])
name = clean_name(name).split()[-1]
for i, jn in enumerate(judges_names):
if jn in name:
return judges_df['name'].iloc[i]
return None
# # Tuple of split, sorted judge name, and original judge name
# judges_names: List[Tuple[List[str], str]]
# judges_names = [
# (sorted(clean_name(jn).split()), jn)
# for jn in judges_df['name']
# ]
# name: List[str]
# name = sorted(map(clean_name, name.split()))
# for jn_low, jn in judges_names:
# is_judge: bool
# is_judge = all([
# similar(n, j, 0.8)
# for n, j in zip(name, jn_low)
# if len(n) != 1
# ])
# if is_judge:
# return jn
return None
# + id="wG0fYt4vGgxh"
# Use geonamescache library in order to get current list of all countries
gc = geonamescache.GeonamesCache()
COUNTRIES: Iterator[str]
COUNTRIES = gc.get_countries_by_names().keys()
# + [markdown] id="NnakkL3bEEWq"
# #### BIA SCRAPER CLASS:
# • A smarter implementation of this in a notebook would have been separating these methods out into individual functions in different cells. I preemtively set it up this way imagining it would improve optimization, but with the small size of the documents we are handling, optimization should not be a huge concern. Experimentation and validation should be the #1 priority.
# + id="4AXvG3fFeJaz"
class BIACase:
def __init__(self, text: str):
'''
• Input will be text from a BIA case pdf file, after the pdf has
been converted from PDF to text.
• Scraping works utilizing spaCy, tokenizing the text, and iterating
token by token searching for matching keywords.
'''
self.doc: Doc
self.doc = nlp(text)
self.ents: Tuple[Span]
self.ents = self.doc.ents
def get_ents(self, labels: List[str] = None) -> Iterator[Span]:
'''
• Retrieves entitiess of a specified label(s) in the document,
if no label is specified, returns all entities
'''
return (ent for ent in self.ents if ent.label_ in labels) \
if labels \
else self.ents
def get_country_of_origin(self) -> str:
'''
• Returns the country of origin of the applicant. Currently just checks
the document for a country that is NOT the United States.
'''
locations: Iterator[str]
locations = map(lambda ent: ent.text,
self.get_ents(['GPE']))
similar_country: Callable[[str, float], Union[str, None]]
similar_country = similar_in_list(COUNTRIES)
for loc in locations:
if not similar(loc, 'United States', 0.9):
origin: Union[str, None]
origin = similar_country(loc, 0.9)
if origin: return origin
else: continue
return None
def get_date(self) -> Union[str, None]:
'''
• Returns date of the document. Easy to validate by the PDF filename,
whether its hosted on scribd or somewhere else.
'''
clean_date: Callable[[str], str]
clean_date = lambda s: ''.join([
char for char in s
if char.isalnum()
or char.isspace()
])
dates: Iterator[str]
dates = map(lambda ent: clean_date(ent.text),
self.get_ents(['DATE']))
for date in dates:
try:
# SHOULD return list of length 3,
# Such as ['Sept', '2', '2019']
d: List[str]
d = date.split()
if len(d) != 3:
continue
else:
# Ex. Jan, Feb, ..., Sep, Oct, Dec
month: str
month = d[0][:3].title()
# Ex. 01, 02, 03, ..., 29, 30, 31
day: str
day = '0' + d[1] \
if len(d[1]) == 1 else d[1]
# Ex. 1991, 1992, ..., 2020, 2021
year: str
year = d[2]
# Ex. Jan 09 2014
parsed_date: str
parsed_date = ' '.join([month, day, year])
# datetime obj, Ex Repr: 2020-09-24 00:00:00
dt: datetime
dt = datetime.strptime(parsed_date, '%b %d %Y')
# strip time of hours/min/sec, save as str
dt: str
dt = str(dt).split()[0]
return dt
except:
continue
return None
def get_panel(self) -> Union[List[str], None]:
'''
• Returns the panel members of case in document.
TODO: Check judges names less strictly - I've seen a document
that named the Judge Monsky differently than how she regularly
appears.
'''
panel_members = List[str]
panel_members = []
possible_members: Iterator[Span]
possible_members = map(lambda ent: ent.text,
self.get_ents(['PERSON', 'ORG']))
for member in possible_members:
judge: Union[str, None]
judge = get_if_judge(member)
if judge:
panel_members.append(judge)
return list(set(panel_members)) \
if panel_members \
else None
def get_surrounding_sents(self, token: Token) -> Span:
'''
• This function will return the two sentences surrounding the token,
including the sentence holding the token.
'''
start: int
start = token.sent.start
end: int
end = token.sent.end
try:
sent_before_start: int
sent_before_start = self.doc[start-1].sent.start
sent_after_end: int
sent_after_end = self.doc[end+1].sent.end
except:
return token.sent
surrounding: Span
surrounding = self.doc[sent_before_start:sent_after_end+1]
return surrounding
def get_protected_grounds(self) -> Union[List[str], None]:
'''
• This will return the protected ground(s) of the applicant. Special
checks are needed. Checking for keywords is not enough, as sometimes
documents label laws that describe each protected ground. Examples
are 'Purely Political Offense' and 'Real Id Act'.
'''
protected_grounds: List[str]
protected_grounds = [
'race',
'religion',
'nationality',
'social',
'political',
]
pgs = []
similar_pg: Callable[[str, float], Union[str, None]]
similar_pg = similar_in_list(protected_grounds)
for token in self.doc:
sent: str
sent = token.sent.text.lower()
s: Union[str, None]
s = similar_pg(token.text.lower(), 0.9)
if s == 'social':
next_word = self.doc[token.i+1].text.lower()
if not similar(next_word, 'group', 0.95):
continue
elif s == 'political':
next_word = self.doc[token.i+1].text.lower()
if similar(next_word, 'offense', 0.95):
continue
elif s == 'nationality':
next_word = self.doc[token.i+1].text.lower()
if similar(next_word, 'act', 1):
continue
if s:
surrounding: Span
surrounding = self.get_surrounding_sents(token)
if 'real id' in sent:
continue
elif 'grounds specified' in surrounding.text.lower():
continue
elif 'no claim' in surrounding.text.lower():
continue
pgs.append(s)
return list(set(pgs)) if pgs else None
def get_application(self) -> Dict[str, bool]:
'''
• This will return the seeker's application, found after 'APPLICATION'.
Because HRF is only interested in Asylum, Withholding of Removal,
and Convention Against Torture applications, the others should be
ignored and not included in the dataset.
'''
relevant_applications: List[str]
relevant_applications = [
'asylum',
'withholding',
'torture'
]
similar_app: Callable[[str, float], Union[str, None]]
similar_app = similar_in_list(relevant_applications)
app: Dict[str, bool]
application = {
'asylum': False,
'withholding_of_removal': False,
'CAT': False
}
for token in self.doc:
if similar(token.text, 'APPLICATION', .86):
for i in range(1,30):
word: str
word = self.doc[i + token.i].text.lower()
app: Union[str, None]
app = similar_app(word, 0.9)
if app == 'asylum':
application['asylum'] = True
elif app == 'withholding':
application['withholding_of_removal'] = True
elif app == 'torture':
application['CAT'] = True
return application
def get_outcome(self) -> Union[str, None]:
'''
• Returns the outcome of the case. This will appear after 'ORDER'
at the end of the document.
'''
outcomes: List[str]
outcomes = [
'remanded',
'reversal',
'dismissed',
'sustained',
'terminated',
'granted',
'denied',
'returned'
]
outcomes: Iterator[str]
outcomes_lemma = map(lambda s: nlp(s)[0].lemma_, outcomes)
similar_outcome: Callable[[str, float], Union[str, None]]
similar_outcome = similar_in_list(outcomes)
similar_outcome_l: Callable[[str, float], Union[str, None]]
similar_outcome_l = similar_in_list(outcomes)
dlen: int
dlen = len(self.doc)
# iterating token by token through document in reverse
# improves efficiency only slightly
for i in reversed(range(dlen-1)):
token: Token
token = self.doc[i]
if similar(token.text, 'ORDER', 0.9):
for ii in range(i+1, dlen):
o: Union[str, None]
o = similar_outcome(self.doc[ii].text, 0.9)
o = o if o else similar_outcome_l(self.doc[ii].text, 0.92)
if o:
return nlp(o)[0].lemma_
return None
def get_based_violence(self) -> Union[Dict[str, List[str]], None]:
'''
• Returns a dictionary where the keys are:
Family-based violence,
Gender-based violence,
Gang-based violence
• If a key is in the dict, it means the based_violence is present
in the document, and the relevant sentence(s) where the information is
contained in the key's value
'''
violent_terms: List[str]
violent_terms = [
'hurt',
'kill',
'rape',
'assassinate',
'abuse',
'threat',
'murder',
'torture',
'assault',
'shoot',
'suffer',
'abduct',
'kidnap',
'harm',
'persecute',
'scare',
'fear'
]
sg_family: List[str]
sg_family = [
'family',
'woman',
'partner',
'husband',
'wife',
'son',
'daughter',
'child',
'ethnicity',
'parent'
]
sg_gender: List[str]
sg_gender = [
'sex'
'gender',
'sexuality',
'woman',
'transgender',
'lgbt',
'lgbtq',
'lgbtqia',
'homosexual',
'homosexuality',
'gay',
'lesbian',
'queer',
]
similar_vterm: Callable[[str, float], Union[str, None]]
similar_vterm = similar_in_list(violent_terms)
similar_sg_family: Callable[[str, float], Union[str, None]]
similar_sg_family = similar_in_list(sg_family)
similar_sg_gender: Callable[[str, float], Union[str, None]]
similar_sg_gender = similar_in_list(sg_gender)
based_v = defaultdict(lambda: [])
for token in self.doc:
if similar_sg_family(token.lemma_.lower(), 0.9):
sent: Span
sent = token.sent
for w in sent:
vterm = similar_vterm(w.lemma_.lower(), 0.86)
if vterm and 'statute' not in token.sent.text:
based_v['family-based'] += [token.lemma_.lower()]
elif similar_sg_gender(token.text.lower(), 0.86):
sent: Span
sent = self.get_surrounding_sents(token)
for w in sent:
vterm = similar_vterm(w.lemma_.lower(), 0.86)
if vterm and 'statute' not in token.sent.text:
based_v['gender-based'] += [token.lemma_.lower()]
elif similar(token.text.lower(), 'gang', 0.9):
sent = token.sent
based_v['gang-based'] += [token.lemma_.lower()]
if based_v:
based_v: Dict[str, List[str]]
based_v = {k:list(set(v)) for k, v in based_v.items()}
return based_v if based_v else None
def references_AB27_216(self) -> bool:
'''
• Returns True if the case file mentions
Matter of AB, 27 I&N Dec. 316 (A.G. 2018)
'''
for token in self.doc:
if token.text == 'I&N':
sent = token.sent.text
if '316' in sent and '27' in sent:
return True
return False
def references_LEA27_581(self) -> bool:
'''
• Returns True if the case file mentions
Matter of L-E-A-, 27 I&N Dec. 581 (A.G. 2019)
'''
for sent in self.doc.sents:
if 'L-E-A-' in sent.text:
if '27' in sent.text:
return True
return False
def get_seeker_sex(self) -> str:
'''
• This field needs to be validated. Currently, it assumes the
sex of the seeker by the number of instances of pronouns in the
document.
'''
male: int
male = 0
female: int
female = 0
for token in self.doc:
if similar(token.text, 'he', 1) \
or similar(token.text, 'him', 1) \
or similar(token.text, 'his', 1):
male += 1
elif similar(token.text, 'she', 1) \
or similar(token.text, 'her', 1):
female += 1
return 'male' if male > female \
else 'female' if female > male \
else 'unkown'
# + [markdown] id="KtNqYRO0EyI8"
# #### ASSEMBLING DATA:
# • I assembled data here in order to fit the structure of a csv, but any implementation could have been made. CSV has the pro of being easily manipulated by the Pandas library, but web people particularly like their JSON. The only downside to CSV is that most of these fields are categorical, and can contain 0 to n "tags" that would be better represented in a DB schema. You can see specifically how I manipulated this CSV in the streamlit repository.
# + id="KN87zvLWi1rB"
text_files = os.listdir(ASYLUM_DIR)
data = []
for i, f in enumerate(text_files):
case = BIACase(get_text_from(f))
case_data = {}
remove_pw: Callable[[str], str]
remove_pw = lambda s: s[:s.find('?secret')]
case_data['filename'] = remove_pw(f) if 'password' in f else f[:-4]
app = case.get_application()
app = [ap for ap, b in app.items() if b]
case_data['application'] = '; '.join(app) if app else None
case_data['date'] = case.get_date()
case_data['country_of_origin'] = case.get_country_of_origin()
panel = case.get_panel()
case_data['panel_members'] = '; '.join(panel) if panel else None
case_data['outcome'] = case.get_outcome()
pgs = case.get_protected_grounds()
case_data['protected_grounds'] = '; '.join(pgs) if pgs else None
based_violence = case.get_based_violence()
violence = '; '.join([k for k, v in based_violence.items() if v]) \
if based_violence \
else None
keywords = '; '.join(['; '.join(v) for v in based_violence.values()]) \
if based_violence \
else None
case_data['based_violence'] = violence
case_data['keywords'] = keywords
references = [
'Matter of AB, 27 I&N Dec. 316 (A.G. 2018)'
if case.references_AB27_216() else None,
'Matter of L-E-A-, 27 I&N Dec. 581 (A.G. 2019)'
if case.references_LEA27_581() else None
]
case_data['references'] = '; '.join([r for r in references if r])
case_data['sex_of_applicant'] = case.get_seeker_sex()
data.append(case_data)
# + id="6oWg3WeMoOel"
df = pd.DataFrame(data)
df = df.fillna(value=np.nan)
df.to_csv('asylum_cases.csv', index=False)
# + colab={"base_uri": "https://localhost:8080/"} id="-dccBfiXlpbx" outputId="a5021e12-811c-4551-c4b2-99aa81eb8193"
print('spaCy',spacy.__version__)
print('bs4', bs4.__version__)
print('geonamescache', geonamescache.__version__)
# + [markdown] id="1Fz8QG-CFyWZ"
# # CLOSING NOTES:
#
# • There are numerous ways text mining can be done. These documents are fairly structured and consistent generally, but there are many 'little things' that need to be accounted for and can only be discovered through the process of getting aquanted with the documents and experimenting with the scraper. Learning the domain is the hardest part, we're not lawyers.
#
# • Something that I avoided doing was relying on newline breaks that typically structure these documents. That's 100% an alternative to explore, and the text that gets converted from PDFs generally preverse these newline separations.
#
# • Finally, and good framework to apply when gathering data from these documents is accounting for false negatives and false positives and how you can minimize the two. Again, the only way to validate it to test a batch of documents, see if the data acquired is accurate to what the acutal document says, and go from there. It can be a slow and tedious process somtimes, but it's a fascinating one if you're interested in practicing/learning NLP and text mining.
| notebooks/BIA_SCRAPER.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import pandas as pd
import numpy as np
from sklearn.metrics.pairwise import pairwise_distances
from sklearn.cluster import KMeans
# %autoreload
pwd
# cd ../seattle/
sea = pd.read_csv('analysis_sea.csv', index_col=0)
sea = sea.reset_index(drop=True)
sea_meta = pd.read_csv('meta_sea.csv', index_col=0)
# cd ../san_fran/
sf = pd.read_csv('analysis_sf.csv', index_col=0)
sf = sf.reset_index(drop=True)
sf_meta = pd.read_csv('meta_sf.csv', index_col=0)
s = sea_meta.head(10)
def make_url(row):
url = "http://www.zillow.com/{0}-{1}-{2}/zpid_".format(row[1], row[2], row[3])
print (url + str(row[4])).replace(' ', '-')
for row in s.iterrows():
make_url(row[1])
# +
# km = KMeans(n_clusters=4, n_jobs=-1)
# km.fit(sf)
# predictions = km.predict(df_sf)
# len(predictions)
# -
# cd ../../code
from learn_preferences import LearnPreferences
from metrics import space_distance, walk_distance
df_sf_ref
lp = LearnPreferences(df_sf, df_sea, [space_distance, walk_distance], 1, 30)
# +
# for i in range(8):
# lp.get_user_choice(df_sf, df_sea)
# lp.guess_preferences()
# lp.recommendation_history
# -
# cd ../data/seattle/
s = pd.read_csv('trx_sea.csv')
s.columns
| src/ab_testing/code/.ipynb_checkpoints/similarity-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
import scipy.stats as stats
maxwell = stats.maxwell
uniform = stats.uniform
norm = stats.norm
# # 2.1)
#
# **\*Nota: Tener cuidado con los nombres de los archivos que se cargan.**
#
# **\*Nota 2: El número de colisiones no es exactamente el que se dice, es aproximado.**
#
# ### Distribución uniforme
# **Distribución uniforme 20 colisiones y temperatura más baja**
# +
v = np.loadtxt("output20uni.txt")
_, bins, _ = plt.hist(v, 20, density=1, alpha=0.5)
maxwell = stats.maxwell
param1,param2 = maxwell.fit(v, floc=0)
best_fit_line = maxwell.pdf(bins, param1, param2)
plt.plot(bins, best_fit_line)
# -
# **Distribución uniforme 800 colisiones y temperatura más baja**
# +
v1 = np.loadtxt("output800uni.txt")
_, bins, _ = plt.hist(v1, 20, density=1, alpha=0.5)
maxwell = stats.maxwell
param1,param2 = maxwell.fit(v1, floc=0)
best_fit_line = maxwell.pdf(bins, param1, param2)
plt.plot(bins, best_fit_line)
# -
# **Distribución uniforme con 1500 colisiones y temperatura más baja (Opcional)**
# +
v2 = np.loadtxt("output1500uni.txt")
_, bins, _ = plt.hist(v2, 20, density=1, alpha=0.5)
maxwell = stats.maxwell
param1,param2 = maxwell.fit(v2, floc=0)
best_fit_line = maxwell.pdf(bins, param1, param2)
plt.plot(bins, best_fit_line)
# -
# --------------
# # 2.2)
# **Distribución uniforme 50 colisiones y temperatura media**
# +
v3 = np.loadtxt("output50unimedia.txt")
_, bins, _ = plt.hist(v3, 20, density=1, alpha=0.5)
maxwell = stats.maxwell
param1,param2 = maxwell.fit(v3, floc=0)
best_fit_line = maxwell.pdf(bins, param1, param2)
plt.plot(bins, best_fit_line)
# -
# **Distribución uniforme 800 colisiones y temperatura media**
# +
v4 = np.loadtxt("output800unimedia.txt")
_, bins, _ = plt.hist(v4, 20, density=1, alpha=0.5)
maxwell = stats.maxwell
param1,param2 = maxwell.fit(v4, floc=0)
best_fit_line = maxwell.pdf(bins, param1, param2)
plt.plot(bins, best_fit_line)
# -
# **Distribución uniforme 1600 colisiones y temperatura media (opcional)**
# +
v5 = np.loadtxt("output1600unimedia.txt")
_, bins, _ = plt.hist(v5, 20, density=1, alpha=0.5)
maxwell = stats.maxwell
param1,param2 = maxwell.fit(v5, floc=0)
best_fit_line = maxwell.pdf(bins, param1, param2)
plt.plot(bins, best_fit_line)
# -
# # 2.3)
# **Distribución uniforme con 100 colisiones y temperatura máxima**
# +
v6 = np.loadtxt("output100unialta.txt")
_, bins, _ = plt.hist(v6, 20, density=1, alpha=0.5)
maxwell = stats.maxwell
param1,param2 = maxwell.fit(v6, floc=0)
best_fit_line = maxwell.pdf(bins, param1, param2)
plt.plot(bins, best_fit_line)
# -
# **Distribución uniforme con 800 colisiones y temperatura máxima**
# +
v7 = np.loadtxt("output800unialta.txt")
_, bins, _ = plt.hist(v7, 20, density=1, alpha=0.5)
maxwell = stats.maxwell
param1,param2 = maxwell.fit(v7, floc=0)
best_fit_line = maxwell.pdf(bins, param1, param2)
plt.plot(bins, best_fit_line)
# -
# **Distribución uniforme con 1600 colisiones y temperatura máxima (opcional)**
# +
v8 = np.loadtxt("output1600unialta.txt")
_, bins, _ = plt.hist(v8, 20, density=1, alpha=0.5)
maxwell = stats.maxwell
param1,param2 = maxwell.fit(v8, floc=0)
best_fit_line = maxwell.pdf(bins, param1, param2)
plt.plot(bins, best_fit_line)
# -
# ---------------
# # 2.4)
# **Distribución Normal con 20 colisiones y temperatura mínima**
# +
v9 = np.loadtxt("output20norm.txt")
_, bins, _ = plt.hist(v9, 20, density=1, alpha=0.5)
maxwell = stats.maxwell
param1,param2 = maxwell.fit(v9, floc=0)
best_fit_line = maxwell.pdf(bins, param1, param2)
plt.plot(bins, best_fit_line)
# -
# **Distribución Normal con 800 colisiones y temperatura mínima**
# +
v10 = np.loadtxt("output800norm.txt")
_, bins, _ = plt.hist(v10, 20, density=1, alpha=0.5)
maxwell = stats.maxwell
param1,param2 = maxwell.fit(v10, floc=0)
best_fit_line = maxwell.pdf(bins, param1, param2)
plt.plot(bins, best_fit_line)
# -
# **Distribución Normal con 1000 colisiones y temperatura mínima (opcional)**
# +
v11 = np.loadtxt("output1000norm.txt")
_, bins, _ = plt.hist(v11, 20, density=1, alpha=0.5)
maxwell = stats.maxwell
param1,param2 = maxwell.fit(v11, floc=0)
best_fit_line = maxwell.pdf(bins, param1, param2)
plt.plot(bins, best_fit_line)
# -
# # 2.5)
# **Distribución Normal con 50 colisiones y temperatura media**
# +
v12 = np.loadtxt("output50normmedia.txt")
_, bins, _ = plt.hist(v12, 20, density=1, alpha=0.5)
maxwell = stats.maxwell
param1,param2 = maxwell.fit(v12, floc=0)
best_fit_line = maxwell.pdf(bins, param1, param2)
plt.plot(bins, best_fit_line)
# -
# **Distribución Normal con 500 colisiones y temperatura media**
# +
v13 = np.loadtxt("output500normmedia.txt")
_, bins, _ = plt.hist(v13, 20, density=1, alpha=0.5)
maxwell = stats.maxwell
param1,param2 = maxwell.fit(v13, floc=0)
best_fit_line = maxwell.pdf(bins, param1, param2)
plt.plot(bins, best_fit_line)
# -
# **Distribución Normal con 1000 colisiones y temperatura media (opcional)**
# +
v14 = np.loadtxt("output1000normmedia.txt")
_, bins, _ = plt.hist(v14, 20, density=1, alpha=0.5)
maxwell = stats.maxwell
param1,param2 = maxwell.fit(v14, floc=0)
best_fit_line = maxwell.pdf(bins, param1, param2)
plt.plot(bins, best_fit_line)
# -
# # 2.6)
# **Distribución Normal con 100 colisiones y temperatura máxima**
# +
v15 = np.loadtxt("output100normalta.txt")
_, bins, _ = plt.hist(v15, 20, density=1, alpha=0.5)
maxwell = stats.maxwell
param1,param2 = maxwell.fit(v15, floc=0)
best_fit_line = maxwell.pdf(bins, param1, param2)
plt.plot(bins, best_fit_line)
# -
# **Distribución Normal con 500 colisiones y temperatura máxima**
# +
v16 = np.loadtxt("output500normalta.txt")
_, bins, _ = plt.hist(v16, 20, density=1, alpha=0.5)
maxwell = stats.maxwell
param1,param2 = maxwell.fit(v16, floc=0)
best_fit_line = maxwell.pdf(bins, param1, param2)
plt.plot(bins, best_fit_line)
# -
# **Distribución Normal con 1000 colisiones y temperatura máxima (opcional)**
# +
v17 = np.loadtxt("output1000normalta.txt")
_, bins, _ = plt.hist(v17, 20, density=1, alpha=0.5)
maxwell = stats.maxwell
param1,param2 = maxwell.fit(v17, floc=0)
best_fit_line = maxwell.pdf(bins, param1, param2)
plt.plot(bins, best_fit_line)
# -
# -----------------
# # 2.7)
# **Una sola partícula con 200 colisiones y termperatura baja**
# +
v18 = np.loadtxt("output200una.txt")
_, bins, _ = plt.hist(v18, 20, density=1, alpha=0.5)
maxwell = stats.maxwell
param1,param2 = maxwell.fit(v18, floc=0)
best_fit_line = maxwell.pdf(bins, param1, param2)
plt.plot(bins, best_fit_line)
# -
# **Una sola partícula con 400 colisiones y termperatura baja**
# +
v19 = np.loadtxt("output400una.txt")
_, bins, _ = plt.hist(v19, 20, density=1, alpha=0.5)
maxwell = stats.maxwell
param1,param2 = maxwell.fit(v19, floc=0)
best_fit_line = maxwell.pdf(bins, param1, param2)
plt.plot(bins, best_fit_line)
# -
# **Una sola partícula con 600 colisiones y termperatura baja (opcional)**
# +
v20 = np.loadtxt("output600una.txt")
_, bins, _ = plt.hist(v20, 20, density=1, alpha=0.5)
maxwell = stats.maxwell
param1,param2 = maxwell.fit(v20, floc=0)
best_fit_line = maxwell.pdf(bins, param1, param2)
plt.plot(bins, best_fit_line)
# -
# # 2.8)
# **Una sola partícula con 200 colisiones y termperatura media**
# +
v21 = np.loadtxt("output200unamedia.txt")
_, bins, _ = plt.hist(v21, 20, density=1, alpha=0.5)
maxwell = stats.maxwell
param1,param2 = maxwell.fit(v21, floc=0)
best_fit_line = maxwell.pdf(bins, param1, param2)
plt.plot(bins, best_fit_line)
# -
# **Una sola partícula con 400 colisiones y termperatura media**
#
# +
v22 = np.loadtxt("output400unamedia.txt")
_, bins, _ = plt.hist(v22, 20, density=1, alpha=0.5)
maxwell = stats.maxwell
param1,param2 = maxwell.fit(v22, floc=0)
best_fit_line = maxwell.pdf(bins, param1, param2)
plt.plot(bins, best_fit_line)
# -
# **Una sola partícula con 800 colisiones y termperatura media (opcional)**
#
# +
v23 = np.loadtxt("output800unamedia.txt")
_, bins, _ = plt.hist(v23, 20, density=1, alpha=0.5)
maxwell = stats.maxwell
param1,param2 = maxwell.fit(v23, floc=0)
best_fit_line = maxwell.pdf(bins, param1, param2)
plt.plot(bins, best_fit_line)
# -
# # 2.9)
# **Una sola partícula con 200 colisiones y termperatura máxima**
#
# +
v24 = np.loadtxt("output200unaalta.txt")
_, bins, _ = plt.hist(v24, 20, density=1, alpha=0.5)
maxwell = stats.maxwell
param1,param2 = maxwell.fit(v24, floc=0)
best_fit_line = maxwell.pdf(bins, param1, param2)
plt.plot(bins, best_fit_line)
# -
# **Una sola partícula con 400 colisiones y termperatura máxima**
#
# +
v25 = np.loadtxt("output400unaalta.txt")
_, bins, _ = plt.hist(v25, 20, density=1, alpha=0.5)
maxwell = stats.maxwell
param1,param2 = maxwell.fit(v25, floc=0)
best_fit_line = maxwell.pdf(bins, param1, param2)
plt.plot(bins, best_fit_line)
# -
# **Una sola partícula con 800 colisiones y termperatura máxima (opcional)**
#
# +
v26 = np.loadtxt("output800unamedia.txt")
_, bins, _ = plt.hist(v26, 20, density=1, alpha=0.5)
maxwell = stats.maxwell
param1,param2 = maxwell.fit(v26, floc=0)
best_fit_line = maxwell.pdf(bins, param1, param2)
plt.plot(bins, best_fit_line)
# -
| Ajustes_python/fitsSimulacion.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# [](https://www.pythonista.io)
# # Introducción a *Javascript*.
# ## Despliegue en la consola.
# La función ```console.log()```.
#
# ``` javascript
# console.log("Hola, mundo.");
# ```
# ## Definición de variables.
# * ```let```
# * ```const```
# * ```var```
# ```javascript
# let x = 2;
# console.log(x + 2);
# x = x * 3;
# const NUMERO = 12;
# console.log(NUMERO + 2);
# NUMER0 = 25;
# ```
# ## Tipos de datos.
#
# *
#
# https://developer.mozilla.org/es/docs/Web/JavaScript/Data_structures
# ```javascript
# console.log(typeof(1));
# console.log(typeof('hola'));
# console.log(typeof(true));
# console.log(typeof(['hola', 'mundo']));
# console.log(typeof(null));
# console.log(typeof(Infinity));
# console.log(typeof(undefined));
# ```
# ## Arreglos.
# ``` javascript
# let arreglo = ["Hugo", "Paco", "Luis"];
# console.log(arreglo[0])
# ```
# ``` javascript
# for (let i of arreglo) {
# console.log(i);
# }
# ```
# ## Objetos convencionales de *Javascript*.
# ``` javascript
# let objeto = {"nombre": "Juan",
# "valor": 2};
# console.log(typeof(objeto));
# console.log(objeto["nombre"]);
# console.log(objeto["valor"]);
# ```
#
# ``` javascript
#
# for (let i in objeto) {
# console.log(i);
# }
#
# for (let i in objeto) {
# console.log(i, objeto[i]);
# }
# ```
# ## Expresioness y operadores.
# https://developer.mozilla.org/es/docs/Web/JavaScript/Guide/Expressions_and_Operators
# ## Mensajes y formularios.
# ``` javascript
# alert("Hola, mundo.");
# ```
# ``` javascript
# let mensaje = prompt("Ingresa el mensaje.")
# console.log(mensaje);
# console.log(typeof(mensaje);
# ```
# ## Condicionales.
# https://developer.mozilla.org/en-US/docs/Web/JavaScript/Reference/Statements/if...else
# ```javascript
# let valor = 3;
# if (valor % 2 == 0) {
# console.log("Es par.");
# }
# else {
# console.log("Es non.");
# }
# ```
# ## Ciclos.
# let iterador = 0;
# while (iterador < 5) {
# iterador +=1;
# console.log(iterador)
# }
# ``` javascript
# iterador = 0;
# do {
# iterador +=1;
# console.log(iterador)
# } while (iterador < 5)
# ```
# ## Funciones.
# ```javascript
# function saluda(nombre){
# console.log("Hola", nombre);
# }
# console.log(saluda("Juan"));
# ```
# ``` javascript
# function suma(a, b=3){
# return a + b
# }
# console.log(suma(2,2));
# console.log(suma(7));
# ```
# ### Funciones flecha.
# let sumatoria = (a, b) => a + b;
# console.log(sumatoria(2, 5));
# ``` javascript
# let par = numero => numero % 2 == 0;
# console.log(par(2));
# ```
# ``` javascript
# let sumatoria = (a, b) => a + b;
# console.log(sumatoria(2, 5));
# ```
# <p style="text-align: center"><a rel="license" href="http://creativecommons.org/licenses/by/4.0/"><img alt="Licencia Creative Commons" style="border-width:0" src="https://i.creativecommons.org/l/by/4.0/80x15.png" /></a><br />Esta obra está bajo una <a rel="license" href="http://creativecommons.org/licenses/by/4.0/">Licencia Creative Commons Atribución 4.0 Internacional</a>.</p>
# <p style="text-align: center">© <NAME>. 2022.</p>
| 11_introduccion_a_javascript.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/grzegorzkwolek/DS-Unit-1-Sprint-1-Dealing-With-Data/blob/master/module2-loadingdata/GKwolek_2nd_assignment_LS_DS_112_Loading_Data_Assignment.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="MSnsTgZLKO72"
# # Practice Loading Datasets
#
# This assignment is purposely semi-open-ended you will be asked to load datasets both from github and also from CSV files from the [UC Irvine Machine Learning Repository](https://archive.ics.uci.edu/ml/index.php).
#
# Remember that the UCI datasets may not have a file type of `.csv` so it's important that you learn as much as you can about the dataset before you try and load it. See if you can look at the raw text of the file either locally, on github, using the `!curl` shell command, or in some other way before you try and read it in as a dataframe, this will help you catch what would otherwise be unforseen problems.
#
# + [markdown] id="156P6ndeKojO" colab_type="text"
# ## 1) Load a dataset from Github (via its *RAW* URL)
#
# Pick a dataset from the following repository and load it into Google Colab. Make sure that the headers are what you would expect and check to see if missing values have been encoded as NaN values:
#
# <https://github.com/ryanleeallred/datasets>
# + id="NJdISe69ZT7E" colab_type="code" outputId="97cb5cdd-2033-4b39-ed97-4f561fd36e5a" colab={"base_uri": "https://localhost:8080/", "height": 630}
# NFL dataset
import pandas as pd
df = pd.read_csv("https://raw.githubusercontent.com/ryanleeallred/datasets/master/NFL-Plays.csv")
df.head()
# + id="pkC7iwZreoSY" colab_type="code" outputId="122605e9-b7a7-4b47-ee04-a5f22ec1f57b" colab={"base_uri": "https://localhost:8080/", "height": 1000}
df.describe().T
# + id="2SMiBp2-fu10" colab_type="code" outputId="81689a4f-c2d9-4924-b4f9-ec8029fa90a1" colab={"base_uri": "https://localhost:8080/", "height": 1000}
import numpy as np
dfcopy = df
dfcopy.fillna("NaN")
dfcopy.isna()
# not sure if all replacements right
# + id="HnffjVj0gxlo" colab_type="code" outputId="47ad0abf-dc8a-48a7-bb66-fb6e0393fd8b" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# inspecting a line of data
dfcopy.iloc[16]
# + [markdown] id="-gFnZR6iLLPY" colab_type="text"
# ## 2) Load a dataset from your local machine
# Download a dataset from the [UC Irvine Machine Learning Repository](https://archive.ics.uci.edu/ml/index.php) and then upload the file to Google Colab either using the files tab in the left-hand sidebar or by importing `files` from `google.colab` The following link will be a useful resource if you can't remember the syntax: <https://towardsdatascience.com/3-ways-to-load-csv-files-into-colab-7c14fcbdcb92>
#
# While you are free to try and load any dataset from the UCI repository, I strongly suggest starting with one of the most popular datasets like those that are featured on the right-hand side of the home page.
#
# Some datasets on UCI will have challenges associated with importing them far beyond what we have exposed you to in class today, so if you run into a dataset that you don't know how to deal with, struggle with it for a little bit, but ultimately feel free to simply choose a different one.
#
# - Make sure that your file has correct headers, and the same number of rows and columns as is specified on the UCI page. If your dataset doesn't have headers use the parameters of the `read_csv` function to add them. Likewise make sure that missing values are encoded as `NaN`.
# + id="qUmwX-ZoM9cq" colab_type="code" outputId="728cc82e-e2e6-4108-be92-e32b2690755d" colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY> "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 75}
# Abalone dataset fail ("RangeError: Maximum call stack size exceeded"). Loading wine dataset.
from google.colab import files
uploaded = files.upload()
# + id="-8RRPzhlucBv" colab_type="code" colab={}
wines_headers = ["Class",
"Alcohol",
"Malic acid",
"Ash",
"Alcalinity of ash",
"Magnesium",
"Total phenols",
"Flavanoids",
"Nonflavanoid phenols",
"Proanthocyanins",
"Color intensity",
"Hue",
"OD280/OD315 of diluted wines",
"Proline"]
# + id="hZnOSLEQo1_B" colab_type="code" colab={}
wines = pd.read_csv("wine (2).data", header = None, names = wines_headers)
# + id="fntC112EycSy" colab_type="code" outputId="558683df-52e3-43de-93f9-917841717902" colab={"base_uri": "https://localhost:8080/", "height": 278}
wines.head()
# + id="My48S9v1sFwZ" colab_type="code" outputId="19ca5796-bac4-4809-874e-e0639cf82eee" colab={"base_uri": "https://localhost:8080/", "height": 34}
wines.shape
# + id="TyGEREoAyGQP" colab_type="code" outputId="ea9d1827-0c2e-4bfd-f9df-69234ee839cf" colab={"base_uri": "https://localhost:8080/", "height": 255}
wines_headers
# + id="ZFYhtXcyuVwT" colab_type="code" outputId="00cfa557-0fb8-4c0a-a704-40fbfa01e0c9" colab={"base_uri": "https://localhost:8080/", "height": 258}
wines.columns = wines_headers
wines.head()
# + id="rt9Q1rUYImTL" colab_type="code" outputId="43ca1f37-3c9c-493b-8fab-988d608cf5b6" colab={"base_uri": "https://localhost:8080/", "height": 1000}
wines.isna()
#the file seems to have no missing values.
# + [markdown] id="mq_aQjxlM-u5" colab_type="text"
# ## 3) Load a dataset from UCI using `!wget`
#
# "Shell Out" and try loading a file directly into your google colab's memory using the `!wget` command and then read it in with `read_csv`.
#
# With this file we'll do a bit more to it.
#
# - Read it in, fix any problems with the header as make sure missing values are encoded as `NaN`.
# - Use the `.fillna()` method to fill any missing values.
# - [.fillna() documentation](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.fillna.html)
# - Create one of each of the following plots using the Pandas plotting functionality:
# - Scatterplot
# - Histogram
# - Density Plot
#
# + id="rhshR4jLIUWj" colab_type="code" colab={}
# !wget https://archive.ics.uci.edu/ml/machine-learning-databases/abalone/abalone.data
# + id="lU3Bpr9NJFLG" colab_type="code" outputId="323d2835-4ca7-4216-ed68-a5836ae57f92" colab={"base_uri": "https://localhost:8080/", "height": 221}
abalone_headers = ["Sex",
"Length",
"Diameter",
"Height",
"Whole weight",
"Shucked weight",
"Viscera weight",
"Shell weight",
"Age"]
df = pd.read_csv("abalone.data", header = None, names = abalone_headers)
df.head()
#heh, I was unable to load this set through google.colab upload
# + id="Tg45v2LVKeAJ" colab_type="code" outputId="26c8714e-4454-4596-a972-730b7ab3b6f3" colab={"base_uri": "https://localhost:8080/", "height": 1000}
df.isna()
#another file with no missing values...
# + id="zJ2xdCGGLLwG" colab_type="code" outputId="6b0509e7-8a56-4a07-ed6c-b48d818d5037" colab={"base_uri": "https://localhost:8080/", "height": 293}
# !wget https://archive.ics.uci.edu/ml/machine-learning-databases/auto-mpg/auto-mpg.data
# + id="Zd-z2JbuNFaE" colab_type="code" outputId="f5ce17a4-7886-4bd0-a99a-97bf90360a2c" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# !curl https://archive.ics.uci.edu/ml/machine-learning-databases/auto-mpg/auto-mpg.data
# + id="tVYP8M5hLYf3" colab_type="code" outputId="63de2ad7-3cb1-4ff4-ff42-5fa75958c4c6" colab={"base_uri": "https://localhost:8080/", "height": 204}
cars_headers = ["mpg",
"cylinders",
"displacement",
"hp",
"weight",
"acceleration",
"model year",
"origin",
"car name"]
#df = pd.read_csv ("auto-mpg.data", names = cars_headers, delimiter=" ")
df = pd.read_csv ("auto-mpg.data")
df.head()
# + id="gGQK4kQFRJGR" colab_type="code" outputId="a2731379-6ae1-4d12-978c-02035817d007" colab={"base_uri": "https://localhost:8080/", "height": 1000}
help(pd.read_csv)
# + id="ICg13Ji5PTLP" colab_type="code" outputId="b9df78d0-8a16-4432-a9c6-71a1e21f6155" colab={"base_uri": "https://localhost:8080/", "height": 381}
# although the below instrictions fail, I leave them as I would like to get some support (trying to identify uneven space size as separator)
import numpy as np
df2 = np.loadtxt(df)
df2.head()
# + id="z2q9R_c2S-vo" colab_type="code" outputId="dd85519a-8926-479a-bdfa-2402ebddc244" colab={"base_uri": "https://localhost:8080/", "height": 293}
# !wget https://archive.ics.uci.edu/ml/machine-learning-databases/heart-disease/processed.va.data
# + id="L0BdtY_6VF6x" colab_type="code" outputId="18bf63a2-cc31-40e3-dad8-e5786f26d810" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# # !curl https://archive.ics.uci.edu/ml/machine-learning-databases/heart-disease/processed.va.data
# + id="Cp5S-Q_hXUkZ" colab_type="code" outputId="326b4118-23bf-4646-86b8-a2492baff180" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# # !curl https://archive.ics.uci.edu/ml/machine-learning-databases/heart-disease/heart-disease.names
# + id="FlM3QEKXTDkb" colab_type="code" outputId="b419d351-a35b-41eb-f4ee-44c7fec7f398" colab={"base_uri": "https://localhost:8080/", "height": 244}
heart_dis_headers = ["age",
"sex",
"cp",
"trestbps",
"chol",
"fbs",
"restecg",
"thalach",
"exang",
"oldpeak",
"slope",
"ca",
"thal",
"num"]
df3 = pd.read_csv("processed.va.data",
header = None,
names = heart_dis_headers,
na_values = "?")
df3.head()
# + id="UZumyrP2pp-d" colab_type="code" outputId="b2a5674b-2a87-4911-9a83-959456d20d69" colab={"base_uri": "https://localhost:8080/", "height": 1000}
df3.fillna(method = "bfill",
axis = 0)
# + id="LrhcII6BoYCw" colab_type="code" outputId="e79b4937-d979-438b-b96a-6ca2cdcb5e8c" colab={"base_uri": "https://localhost:8080/", "height": 272}
df3.dtypes
# + id="CQltlsCCpUMW" colab_type="code" colab={}
df3["trestbps"] = df3["trestbps"].astype(float)
# + id="qcLbxj33n28-" colab_type="code" outputId="8bbc4efc-3cf0-4828-dc27-30fecd102cf0" colab={"base_uri": "https://localhost:8080/", "height": 286}
df3['age'].plot.density()
# + id="sB7cDJ30n-YX" colab_type="code" outputId="5098b653-8bb2-489a-da6a-157bb1a23040" colab={"base_uri": "https://localhost:8080/", "height": 300}
df3.plot.scatter("age",
"cp")
# + id="tljACmnjoQpM" colab_type="code" outputId="b878f9e0-3e64-466f-c2c1-0acce9d217c5" colab={"base_uri": "https://localhost:8080/", "height": 300}
df3.plot.scatter("age",
"trestbps")
# + id="Knlf2eY4rz_r" colab_type="code" outputId="92f5e5a3-00e0-4b46-990b-cabc693780ba" colab={"base_uri": "https://localhost:8080/", "height": 300}
df3.plot.scatter("trestbps",
"thalach")
# + id="xfd8MOJIrhtG" colab_type="code" outputId="596aefd5-acb6-4a8f-c6ad-5255d6de8464" colab={"base_uri": "https://localhost:8080/", "height": 286}
df3['thalach'].hist(bins = 50)
# + [markdown] id="MZCxTwKuReV9" colab_type="text"
# ## Stretch Goals - Other types and sources of data
#
# Not all data comes in a nice single file - for example, image classification involves handling lots of image files. You still will probably want labels for them, so you may have tabular data in addition to the image blobs - and the images may be reduced in resolution and even fit in a regular csv as a bunch of numbers.
#
# If you're interested in natural language processing and analyzing text, that is another example where, while it can be put in a csv, you may end up loading much larger raw data and generating features that can then be thought of in a more standard tabular fashion.
#
# Overall you will in the course of learning data science deal with loading data in a variety of ways. Another common way to get data is from a database - most modern applications are backed by one or more databases, which you can query to get data to analyze. We'll cover this more in our data engineering unit.
#
# How does data get in the database? Most applications generate logs - text files with lots and lots of records of each use of the application. Databases are often populated based on these files, but in some situations you may directly analyze log files. The usual way to do this is with command line (Unix) tools - command lines are intimidating, so don't expect to learn them all at once, but depending on your interests it can be useful to practice.
#
# One last major source of data is APIs: https://github.com/toddmotto/public-apis
#
# API stands for Application Programming Interface, and while originally meant e.g. the way an application interfaced with the GUI or other aspects of an operating system, now it largely refers to online services that let you query and retrieve data. You can essentially think of most of them as "somebody else's database" - you have (usually limited) access.
#
# *Stretch goal* - research one of the above extended forms of data/data loading. See if you can get a basic example working in a notebook. Image, text, or (public) APIs are probably more tractable - databases are interesting, but there aren't many publicly accessible and they require a great deal of setup.
# + id="f4QP6--JBXNK" colab_type="code" colab={}
| module2-loadingdata/GKwolek_2nd_assignment_LS_DS_112_Loading_Data_Assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Web Crawling using Selenium in Python
# ## Practice 1
# - Naver에 로그인하여 네이버페이에서 구매내역 크롤링하기
# +
# Modules Import
from selenium import webdriver
from bs4 import BeautifulSoup
# Webdriver를 이용하여 브라우저 다루기
# 여기서는 Chrome 브라우저를 이용한다.
driver = webdriver.Chrome('chromedriver.exe')
# 브라우저가 로딩될 때까지 대기한다.(3초)
driver.implicitly_wait(3)
# driver를 통해 해당 페이지에 접속한다.(Naver 로그인 화면)
# 이후 로그인에 필요한 아이디와 비밀번호를 각각 입력시킨다.
# 로그인 버튼을 클릭한다.
# 로그인 버튼의 Tag가 접근하기 어려워 xpath를 이용하여 접근한다.
driver.get('https://nid.naver.com/nidlogin.login?mode=form&url=https%3A%2F%2Fwww.naver.com')
driver.find_element_by_name('id').send_keys('------')
driver.find_element_by_name('pw').send_keys('-----------')
driver.find_element_by_xpath('//*[@id="frmNIDLogin"]/fieldset/input').click()
# 네이버페이에 접속한다.
driver.get('https://order.pay.naver.com/home')
# 구매내역이 많으면 과거 내역들은 생략된다.
# 따라서 '더 보기'버튼을 클릭해줘야 한다.
driver.find_element_by_id('_moreButton').click()
# 해당 사이트의 HTML소스를 불러온다.
# 불러온 HTML소스를 Python으로 읽을 수 있도록 BeautifulSoup을 이용한다.
# 이후 구매내역의 이름이 적혀있는 Tag를 찾아 불러온다.
html = driver.page_source
soup = BeautifulSoup(html, 'html.parser')
pay_list = soup.select('div.p_inr > div.p_info > a > span')
for name in pay_list:
print(name.text.strip())
# -
# ## 오류 해결하기
#
# ### 1. 자동입력방지로 인하여 로그인이 안 되는 오류 해결
# - 아이디와 비밀번호 입력속도를 줄였다.
# - 위 문제로도 해결이 안 될 시 새로고침을 이용하였다.
#
# ### 2. 더 보기 버튼이 적용되지 않은 상태에서 크롤링되는 오류 해결
# - 페이지가 로딩된 후 일정시간 대기
# - 버튼이 적용 된 후 일정시간 대기
#
# ### 3. 구매내역 중 이름에 부가설명(강조)이 포함되어 있는 것 제거(ex.알라딘커뮤니티케이션)
# - strong태그가 포함되어 있으므로 해당 태그 제거
# - decompos() 함수 이용
# +
from selenium import webdriver
from bs4 import BeautifulSoup
import time
driver = webdriver.Chrome('chromedriver.exe')
driver.implicitly_wait(3)
driver.get('https://nid.naver.com/nidlogin.login?mode=form&url=https%3A%2F%2Fwww.naver.com')
# ID와 PW 입력 속도 감소
# ID와 PW 각각을 한 글자씩 천천히 입력한다.(string으로 불러와 한 글자씩 입력)
id_input = driver.find_element_by_name('id')
my_id = '-------'
for word in my_id:
id_input.send_keys(word)
time.sleep(0.3)
pw_input = driver.find_element_by_name('pw')
my_pw = '-----------'
for word in my_pw:
pw_input.send_keys(word)
time.sleep(0.3)
driver.implicitly_wait(10)
driver.find_element_by_xpath('//*[@id="frmNIDLogin"]/fieldset/input').click()
# 새로고침
driver.refresh()
driver.get('https://order.pay.naver.com/home')
# 페이지가 로딩된 후 일정시간 대기
driver.implicitly_wait(10)
driver.find_element_by_id('_moreButton').click()
# 더 보기 버튼이 적용된 후 일정시간 대기
driver.implicitly_wait(100)
html = driver.page_source
soup = BeautifulSoup(html, 'html.parser')
pay_list = soup.select('div.p_inr > div.p_info > a > span')
for name in pay_list:
# 만약 strong태그가 존재하면 제거하라
if name.strong in name:
name.strong.decompose()
print(name.text.strip())
| Web_Crawling_Practice02_Selenium.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: tensorflow_gpuenv
# language: python
# name: tensorflow_gpuenv
# ---
# +
import random
from os import path
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import matplotlib.patches as patches
from coco import COCO
# path to the ground truth annotation and the data
annFile = '/home/test/data/nightowls/nightowls_validation.json'
image_directory = '/home/test/data/nightowls/nightowls_validation'
# load the annotations and the list of images
cocoGt = COCO(annFile)
imgIds = sorted(cocoGt.getImgIds())
print('There are %d images in the training set' % len(imgIds))
annotations = cocoGt.getAnnIds()
print('There are %d annotations in the training set' % len(annotations))
for i in range(20):
# get a random image and its path
im_id = imgIds[random.randint(0, len(imgIds))]
image = cocoGt.loadImgs(ids=im_id)[0]
file_path = path.join(image_directory, image['file_name'])
# get the annotations of the images
anns = cocoGt.getAnnIds([image['id']])
if(len(anns) <1):
continue
print(image)
print('File path: ' + str(file_path))
print('Number of annotations in the image: ' + str(len(anns)))
# draw the annotations on the image
img=mpimg.imread(file_path)
fig,ax = plt.subplots(1)
ax.imshow(img)
for ann_id in anns:
ann = cocoGt.loadAnns(ids=ann_id)[0]
print(ann)
bbox = ann['bbox']
rect = patches.Rectangle((bbox[0],bbox[1]),bbox[2],bbox[3],linewidth=2,edgecolor='g',facecolor='none')
ax.add_patch(rect)
#print
plt.show()
# -
| Code/Preliminary_tests/3-Dataset utils test/My test in loading stuff.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:py35]
# language: python
# name: conda-env-py35-py
# ---
from gensim.models import KeyedVectors
import numpy as np
ft_file = "/Users/i337036/Documents/Data/wiki.en.vec"
word_vecs = KeyedVectors.load_word2vec_format(ft_file, limit=50000)
import pandas as pd
csv_file = "/Users/i337036/Downloads/train.csv"
df = pd.read_csv(csv_file, encoding='utf-8')
len(df)
df.columns.values
df = df[df['is_duplicate']==1]
df = df.drop_duplicates(subset=['question1'], keep='first')
df = df.drop_duplicates(subset=['question2'], keep='first')
len(df)
len((df["question1"] + df["question2"]).unique())
q1_list = df.question1.tolist()
q2_list = df.question2.tolist()
print(len(q1_list), len(q2_list))
def clean_sentence(sentence):
sentence = [word for word in sentence.strip().split() if word in word_vecs.vocab.keys()]
return sentence
def get_sentence_matrix(sentences):
sentence_mat = np.zeros((len(sentences), 300), dtype=float)
for idx in tqdm(range(len(sentences)), desc="Building vectors for %d sentences" % len(sentences)):
sentence = sentences[idx]
for word in sentence:
sentence_mat[idx, :] += word_vecs[word]
return sentence_mat
from tqdm import tqdm
q1_word_list = [clean_sentence(txt1) for txt1 in q1_list]
q2_word_list = [clean_sentence(txt2) for txt2 in q2_list]
q2_mat = get_sentence_matrix(q1_word_list)
q1_mat = get_sentence_matrix(q2_word_list)
# +
from sklearn.metrics.pairwise import cosine_similarity
def find_similar(q1_mat, q2_mat, index, top_n=5):
qd_sims = cosine_similarity(q1_mat[index: index+1], q2_mat).flatten()
qd_indices = qd_sims.argsort()[:-top_n:-1]
return qd_indices
# -
from random import randint
search_idx = randint(0, len(q1_list))
print("Searched for: %s at [%d]" % (q1_list[search_idx], search_idx))
for index in find_similar(q1_mat, q2_mat, search_idx):
print ("%s %d" % (q2_list[index], index))
q1_list[search_idx], q2_list[search_idx]
from tqdm import tqdm
correct_index = 0
for search_idx in tqdm(range(500)):
for index in find_similar(q1_mat, q2_mat, search_idx):
if search_idx==index or q1_list[search_idx]==q2_list[index]:
correct_index += 1
break
print(correct_index)
| Chapter04/3_PreTrainedWordEmbedding_Retrieval.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={} colab_type="code" id="8A5Jw5NR1iEI"
#Declare an int value and store it in a variable.
a=10
#Check the type and print the id of the same.
print(type(a))
print(id(a))
# + colab={} colab_type="code" id="0YU8LFTn1rAX"
#Take one int value between 0 - 256.
#Assign it to two different variables.
#Check the id of both the variables. It should come same. Check why?
a=10
d=10
print(id(a),id(d))
#Because the value of a and d is similar and it is in between the 0 to 256
#object reusability
print()
#Take one int value either less than -5 or greater than 256.
#Assign it to two different variables.
#Check the id of both the variables. It should come different.Check why?
b=-56
c=-56
print(id(b),id(c))
#Because the value of a and b is similar but that are out of ranged from 0 to 256
# + colab={} colab_type="code" id="YzEIG0ZZ1tSK"
#Arithmatic Operations on integers
#Take two different intger values.
#Store them in two different variables.
#Do below operations on them:-
#Find sum of both numbers
#Find differce between them
#Find the product of both numbers.
#Find value after dividing first num with second number
#Find the remainder after dividing first number with second number
#Find the quotient after dividing first number with second number
#Find the result of first num to the power of second number.
a=5
b=3
c=a+b
d=a-b
e=a*b
f=a/b
i=a%b
g=a//b
h=b**a
print(c,d,e,f,i,g,h)
# + colab={} colab_type="code" id="GGM7CdzA1wGn"
#Comparison Operators on integers
#Take two different intger values.
#Store them in two different variables.
#Do below operations on them:-
#Compare se two numbers with below operator:-
#Greater than, '>'
#Smaller than, '<'
#Greater than or equal to, '>='
#Less than or equal to, '<='
#Observe their output(return type should be boolean)
a=4
b=5
c=(a>b)
d=(a<b)
e=(a>=b)
f=(a<=b)
print(c,d,e,f)
# + colab={} colab_type="code" id="9x904sUE1y9t"
#Equality Operator
#Take two different intger values.
#Store them in two different variables.
#Equuate them using equality operator (==, !=)
#Observe the output(return type should be boolean)
a=3
b=4
c=(a==b)
d=(a!=b)
print (c,d)
# + colab={} colab_type="code" id="JmQFHUwc11S-"
#Logical operators
#Observe the output of below code
#Cross check the output manually
print(10 and 20) #----------------------------------------->Output is 20
print(0 and 20) #----------------------------------------->Output is 0
print(20 and 0) #----------------------------------------->Output is 0
print(0 and 0) #----------------------------------------->Output is 0
print(10 or 20) #----------------------------------------->Output is 10
print(0 or 20) #----------------------------------------->Output is 20
print(20 or 0) #----------------------------------------->Output is 20
print(0 or 0) #----------------------------------------->Output is 0
print(not 10) #----------------------------------------->Output is False
print(not 0) #----------------------------------------->Output is True
# ManuallyChecking:
# note: Basically AND fuction searches false, if one of them flase, returns false else returns true .
# 1) 10 is ture and 20 is true then the 20 is printing
# 2) 0 is false and 20 is true then the 0 is printing
# 3) 20 is ture and 0 is false then the 0 is printing
# 4) 0 is false and 0 is false then the 0 is printing
# note: Basically OR fuction searches true only, if one of them true returns ture value else returns flase value
# 5) 10 is ture or 20 is true then the 10 is printing
# 6) 0 is false or 20 is true then the 20 is printing
# 7) 20 is ture or 0 is false then the 20 is printing
# 8) 0 is false or 0 is false then the 0 is printing
# note: not operator returns only True or Flase, if value true returs flase else returns true
# 9) not 10 this true value but the returns flase
# 10)not 0 this false value but the returns true
# + colab={} colab_type="code" id="-0tvoulX14Hi"
#Bitwise Operators
#Do below operations on the values provided below:-
#Bitwise and(&) -----------------------------------------> 10, 20 -------> Output is 0
#Bitwise or(|) -----------------------------------------> 10, 20 -------> Output is 30
#Bitwise(^) -----------------------------------------> 10, 20 -------> Output is 30
#Bitwise negation(~) ------------------------------------> 10 -------> Output is -11
#Bitwise left shift ------------------------------------> 10,2 -------> Output is 40
#Bitwise right shift ------------------------------------> 10,2 -------> Output is 2
#Cross check the output manually
# 10 --------------> 01010
# 20 --------------> 10100
# ----------------------------
# Bitwise and(&)---> 00000 --------------> 0
# Bitwise or(|)----> 11110 -------------->30
# Bitwise(^) -----> 11110 -------------->30
#Bitwise negation(~)~n=~10=-(10+1)------>-11
#left shift (10<<2) = 101000------------> 40
#right shift (10>>2)= 0010--------------> 2
# + colab={} colab_type="code" id="YCcx-Qx016hg"
#What is the output of expression inside print statement. Cross check before running the program.
a = 10
b = 10
print(a is b) #True or False? True
print(a is not b) #True or False? False
a = 1000
b = 1000
print(a is b) #True or False? False
print(a is not b) #True or False? True
# + colab={} colab_type="code" id="Un2To3XN1_Il"
#What is the output of expression inside print statement. Cross check before running the program.
print(10+(10*32)5//2**&20+(~(-10))<<2)
# 1)10*32=320
# 2)2**5=32
# 3)320//32-------=20
# 4)~(-10)=9
# 5)20+9=29
# 6)29<<2---------=116
# 7)20&116-------->20
# + colab={} colab_type="code" id="kGRb5RMd1_1I"
#Membership operation
#in, not in are two membership operators and it returns boolean value
print('2' in 'Python2.7.8') # true
print(10 in [10,10.20,10+20j,'Python']) # true
print(10 in (10,10.20,10+20j,'Python')) # true
print(2 in {1,2,3}) # true
print(3 in {1:100, 2:200, 3:300}) # true
print(10 in range(20)) # true
# + colab={} colab_type="code" id="Es7iSvL92B9W"
#An integer can be represented in binary, octal or hexadecimal form.
a=6
print((bin(a),oct(a),hex(a)))
#Declare one binary, one octal and one hexadecimal value and store them in three different variables.
b=0b1010
c=0o64721
d=0xe345
print((b),(c),(d))
# #Convert 9876 to its binary, octal and hexadecimal equivalent and print their corresponding value.
print()
print(bin(9876),oct(9876),hex(9876))
# -
#What will be the outut of following:-
a = 0b1010000
print(a)
#Decimal value :80
b = 0o7436
print(b)
#Decimal value :3870
c = 0xfade
print(c)
#Decimal value :64222
print(bin(80))
#Binary value :0b1010000
print(oct(3870))
#octal value :0o7436
print(hex(64222))
#hexadecimal value :0xfade
print(bin(0b1010000))
#Binary value :0b1010000
print(bin(0xfade))
#Binary value :0b1111101011011110
print(oct(0xfade))
#octal value :0o175336
print(oct(0o7436))
#octal value :0o7436
print(hex(0b1010000))
#hexadecimal value :0x50
print(hex(0xfade))
#hexadecimal value :0xfade
| Karthi/Assignments/1st term/1. karthi_int_Assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Rolling Regression
#
# Rolling OLS applies OLS across a fixed windows of observations and then rolls
# (moves or slides) the window across the data set. They key parameter is `window`
# which determines the number of observations used in each OLS regression. By
# default, `RollingOLS` drops missing values in the window and so will estimate
# the model using the available data points.
#
# Estimated values are aligned so that models estimated using data points
# $i, i+1, ... i+window$ are stored in location $i+window$.
#
# Start by importing the modules that are used in this notebook.
# + pycharm={"is_executing": false}
import pandas_datareader as pdr
import pandas as pd
import statsmodels.api as sm
from statsmodels.regression.rolling import RollingOLS
import matplotlib.pyplot as plt
import seaborn
seaborn.set_style('darkgrid')
pd.plotting.register_matplotlib_converters()
# %matplotlib inline
# + [markdown] pycharm={"name": "#%% md\n"}
# `pandas-datareader` is used to download data from
# [Ken French's website](https://mba.tuck.dartmouth.edu/pages/faculty/ken.french/data_library.html).
# The two data sets downloaded are the 3 Fama-French factors and the 10 industry portfolios.
# Data is available from 1926.
#
# The data are monthly returns for the factors or industry portfolios.
# + pycharm={"is_executing": false, "name": "#%%\n"}
factors = pdr.get_data_famafrench('F-F_Research_Data_Factors', start='1-1-1926')[0]
print(factors.head())
industries = pdr.get_data_famafrench('10_Industry_Portfolios', start='1-1-1926')[0]
print(industries.head())
# + [markdown] pycharm={"name": "#%% md\n"}
# The first model estimated is a rolling version of the CAPM that regresses
# the excess return of Technology sector firms on the excess return of the market.
#
# The window is 60 months, and so results are available after the first 60 (`window`)
# months. The first 59 (`window - 1`) estimates are all `nan` filled.
# + pycharm={"is_executing": false, "name": "#%%\n"}
endog = industries.HiTec - factors.RF.values
exog = sm.add_constant(factors['Mkt-RF'])
rols = RollingOLS(endog, exog, window=60)
rres = rols.fit()
params = rres.params
print(params.head())
print(params.tail())
# + [markdown] pycharm={"name": "#%% md\n"}
# We next plot the market loading along with a 95% point-wise confidence interval.
# The `alpha=False` omits the constant column, if present.
# + pycharm={"is_executing": false, "name": "#%%\n"}
fig = rres.plot_recursive_coefficient(variables=['Mkt-RF'], figsize=(14,6))
# -
# Next, the model is expanded to include all three factors, the excess market, the size factor
# and the value factor.
# + pycharm={"is_executing": false, "name": "#%%\n"}
exog_vars = ['Mkt-RF', 'SMB', 'HML']
exog = sm.add_constant(factors[exog_vars])
rols = RollingOLS(endog, exog, window=60)
rres = rols.fit()
fig = rres.plot_recursive_coefficient(variables=exog_vars, figsize=(14,18))
# -
# ## Formulas
#
# `RollingOLS` and `RollingWLS` both support model specification using the formula interface. The example below is equivalent to the 3-factor model estimated previously. Note that one variable is renamed to have a valid Python variable name.
joined = pd.concat([factors, industries], axis=1)
joined['Mkt_RF'] = joined['Mkt-RF']
mod = RollingOLS.from_formula('HiTec ~ Mkt_RF + SMB + HML', data=joined, window=60)
rres = mod.fit()
print(rres.params.tail())
# ## `RollingWLS`: Rolling Weighted Least Squares
#
# The `rolling` module also provides `RollingWLS` which takes an optional `weights` input to perform rolling weighted least squares. It produces results that match `WLS` when applied to rolling windows of data.
# ## Fit Options
#
# Fit accepts other optional keywords to set the covariance estimator. Only two estimators are supported, `'nonrobust'` (the classic OLS estimator) and `'HC0'` which is White's heteroskedasticity robust estimator.
#
# You can set `params_only=True` to only estimate the model parameters. This is substantially faster than computing the full set of values required to perform inference.
#
# Finally, the parameter `reset` can be set to a positive integer to control estimation error in very long samples. `RollingOLS` avoids the full matrix product when rolling by only adding the most recent observation and removing the dropped observation as it rolls through the sample. Setting `reset` uses the full inner product every `reset` periods. In most applications this parameter can be omitted.
# + pycharm={"is_executing": false, "name": "#%%\n"}
# %timeit rols.fit()
# %timeit rols.fit(params_only=True)
| v0.12.2/examples/notebooks/generated/rolling_ls.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# [](https://githubtocolab.com/giswqs/leafmap/blob/master/examples/notebooks/01_leafmap_intro.ipynb)
# [](https://gishub.org/leafmap-pangeo)
#
# Uncomment the following line to install [leafmap](https://leafmap.org) if needed.
# +
# # !pip install leafmap
# -
import leafmap
m = leafmap.Map()
m
m = leafmap.Map(center=[50, 19], zoom=4) #center=[lat, lon]
m
m = leafmap.Map(draw_control=False, measure_control=False, fullscreen_control=False, attribution_control=True)
m
m = leafmap.Map(height="450px")
m
| examples/notebooks/01_leafmap_intro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/EngMarceloPaulo/PYTHON/blob/master/PastaDeArquivos/Modulo2Aula05.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="b741HqNoky_C" outputId="caf95c18-d51a-4a1c-87ce-7acfffa4367b"
''' Cópia por valor e cópia por referência.
As melhores práticas em Python
'''
x = [20, 30, 10, -50]
print(x)
# + colab={"base_uri": "https://localhost:8080/"} id="jN8uBjVul0oY" outputId="32473a0c-dc25-4b86-b6f9-701e4e8c4975"
y = x # ERRO! Foi passada a REFERÊNCIA
y[0] = 3 # Ao modificar y armazeno o resultado em x
print(y) # Referência é um vínculo entre as variáveis
print(x)
# + colab={"base_uri": "https://localhost:8080/"} id="XRPg_m4kmxf9" outputId="23f477bc-36ee-4939-cb13-c9f6c705cefc"
z = x.copy() # Foi passado o VALOR
z[0] = 70 # Ao modificar o valor não há modificação em x
print(z) # z representa uma variável local
print(x)
| PastaDeArquivos/Modulo2Aula05.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# +
endpoint = 'http://630317bb-4577-4216-82a1-5e4fe48d92db.southeastasia.azurecontainer.io/score'
key = '<KEY>'
import urllib.request
import urllib.error
import json
import os
data = {
"Inputs": {
"WebServiceInput0": [
{
"id": 2525363215,
"date": "20201013T000000",
"price": 356300,
"bedrooms": 3,
"bathrooms": 1,
"sqft_living": 1180,
"sqft_lot": 5650,
"floors": 1,
"waterfront": 0,
"view": 0,
"condition": 3,
"grade": 7,
"sqft_above": 1180,
"sqft_basement": 0,
"yr_built": 1955,
"yr_renovated": 0,
"zipcode": 98178,
"lat": 475.11199999999997,
"long": -122.257,
"sqft_living15": 1340,
"sqft_lot15": 5650
},
{
"id": 5054545054,
"date": "20211012T000000",
"price": 520231,
"bedrooms": 4,
"bathrooms": 2,
"sqft_living": 1680,
"sqft_lot": 6750,
"floors": 2,
"waterfront": 1,
"view": 1,
"condition": 5,
"grade": 9,
"sqft_above": 1180,
"sqft_basement": 0,
"yr_built": 2000,
"yr_renovated": 2015,
"zipcode": 98177,
"lat": 479.213245451444,
"long": -132.217,
"sqft_living15": 1240,
"sqft_lot15": 5850
},
{
"id": 6561254452,
"date": "20201015T000000",
"price": 600000,
"bedrooms": 4,
"bathrooms": 3,
"sqft_living": 1880,
"sqft_lot": 7850,
"floors": 2,
"waterfront": 1,
"view": 1,
"condition": 6,
"grade": 10,
"sqft_above": 1380,
"sqft_basement": 130,
"yr_built": 2012,
"yr_renovated": 2015,
"zipcode": 98177,
"lat": 485.212545331254,
"long": -142.257,
"sqft_living15": 1640,
"sqft_lot15": 7650
}
]
},
"GlobalParameters": {}
}
body = str.encode(json.dumps(data))
headers = {
'Content-Type':'application/json',
'Authorization':('Bearer '+ key)
}
req = urllib.request.Request(endpoint, body, headers)
try:
response = urllib.request.urlopen(req)
result = response.read()
json_rslt = json.loads(result)
data = json_rslt["Results"]["WebServiceOutput0"]
print(data)
except urllib.error.HTTPError as error :
print("Request gagal dengan kode status: " + str(error.code))
print(error.info())
print(json.loads(error.read().decode("utf8", 'ignore')))
| Testing/Testing2/test-price-prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/mohitmishra786/Machine-Learning-Models/blob/main/Basic_Keras_Model_for_Diabetes_Dataset.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="kiJ9SvUE2aIt"
# first neural network with keras tutorial
import pandas as pd
from keras.models import Sequential
from keras.layers import Dense
from sklearn.model_selection import train_test_split
# + id="Ar1ukgO03OHB"
# load the dataset
dataset = pd.read_csv('/content/diabetes_.csv')
# split into input (X) and output (y) variables
X = dataset.iloc[:,0:8].astype(float)
y = dataset.iloc[:,8]
# Splitting data into train and test split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.25, random_state=42)
# + colab={"base_uri": "https://localhost:8080/"} id="QAR55vYrANnv" outputId="4c858418-03b9-4899-c2ab-e6e54cc19723"
# define the keras model
model = Sequential()
model.add(Dense(12, input_dim=8, activation='relu'))
model.add(Dense(8, activation='relu'))
model.add(Dense(1, activation='sigmoid'))
# compile the keras model
model.compile(loss='binary_crossentropy', optimizer='adam', metrics=['accuracy'])
# fit the keras model on the dataset
model.fit(X_train, y_train, epochs=150, batch_size=10)
# evaluate the keras model
y_pred = model.predict(X_test)
# + colab={"base_uri": "https://localhost:8080/"} id="3mofxJkDANqY" outputId="36f056c8-e8ac-493b-bf94-5149018bed93"
# evaluate the keras model
_, accuracy = model.evaluate(X_test, y_test)
print('Accuracy: %.2f' % (accuracy*100))
# + id="H4HFxHAGANsn"
| Basic_Keras_Model_for_Diabetes_Dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Chapter 7: Sequential Data (TL;DR)
# + [markdown] slideshow={"slide_type": "skip"}
# **Sequences** are an *abstract* concept that summarizes *four* behaviors an object may or may not exhibit. Sequences are
# - **finite** and
# - **ordered**
# - **containers** that we may
# - **loop over**.
#
# Examples are the `list`, `tuple`, but also the `str` types.
#
# Objects that exhibit all behaviors *except* being ordered are referred to as **collections**.
#
# The objects inside a sequence are called its **elements** and may be labeled with a unique **index**, an `int` object in the range $0 \leq \text{index} < \lvert \text{sequence} \rvert$.
#
# `list` objects are **mutable**. That means we can change the references to the other objects it contains, and, in particular, re-assign them.
#
# On the contrary, `tuple` objects are like **immutable** lists: We can use them in place of any `list` object as long as we do *not* need to mutate it. Often, `tuple` objects are also used to model **records** of related **fields**.
| 07_sequences/06_summary.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.8 64-bit (''base'': conda)'
# name: python3
# ---
import yfinance as yf
import pandas as pd
import numpy as np
def download_dados(tickers, interval = '1d', period = 'max'):
df = yf.download(tickers, interval=interval, period=period)['Adj Close']
#retorno, df_ln = coin.calc_ret_ln(df)
return df
ativos = pd.read_csv('ativos.csv', sep=';')
ativos = ativos.iloc[:-40]
ativos.drop([425, 506, 561, 681], inplace=True)
ativos
tickers = ' '
for i in range(len(ativos)):
aux = ativos.iloc[i][1]
tickers = tickers +' '+ aux
tickers
# +
#tickers = "ABEV3.SA AZUL4.SA B3SA3.SA BBAS3.SA BBDC3.SA BBDC4.SA BBSE3.SA BRAP4.SA BRFS3.SA BRKM5.SA BRML3.SA CCRO3.SA CIEL3.SA CMIG4.SA COGN3.SA CRFB3.SA CSAN3.SA CSNA3.SA CVCB3.SA CYRE3.SA ECOR3.SA EGIE3.SA ELET3.SA ELET6.SA EMBR3.SA ENBR3.SA EQTL3.SA FLRY3.SA GGBR4.SA GNDI3.SA GOAU4.SA GOLL4.SA HAPV3.SA HGTX3.SA HYPE3.SA IGTA3.SA IRBR3.SA ITSA4.SA ITUB4.SA JBSS3.SA KLBN11.SA LAME4.SA LREN3.SA MGLU3.SA MRFG3.SA MRVE3.SA MULT3.SA PETR3.SA PETR4.SA QUAL3.SA RADL3.SA RAIL3.SA RENT3.SA SANB11.SA SBSP3.SA SULA11.SA SUZB3.SA TAEE11.SA TOTS3.SA UGPA3.SA USIM5.SA VALE3.SA VIIA3.SA WEGE3.SA YDUQ3.SA"
# +
from mlpairs import OpticsPairs
import pandas as pd
import pickle
with open('close_price.pickle', 'rb') as cp:
stock_prices = pickle.load(cp)
#stock_prices = yf.download(tickers, interval='1d', period='3y')['Adj Close']
#stock_prices = pd.read_csv('stock_prices.csv', index_col='Date')
train = stock_prices[:int(len(stock_prices)/0.7)]
test = stock_prices[int(len(stock_prices)*0.7):]
train.head()
# -
int(len(stock_prices)*0.7)
for i in range(len(train)):
for j in range(len(train.columns)):
try:
if np.isnan(train.iloc[i][j]) == True and np.isnan(train.iloc[i+1][j]) == True:
train.iloc[i][j] = 0
elif np.isnan(train.iloc[i][j]) == True and train.iloc[i+1][j] > 0:
train.iloc[i][j] = (train.iloc[i-1][j]+train.iloc[i+1][j])/2
except:
pass
op = OpticsPairs(train)
op.returns.head()
#op.returns.dropna(inplace=True, axis=1)
import numpy as np
for i in range(len(op.returns)):
for j in range(len(op.returns.columns)):
try:
if np.isnan(op.returns.iloc[i][j]) == True and np.isnan(op.returns.iloc[i-1][j]) == True:
op.returns.iloc[i][j] = 0
except:
pass
if np.isnan(op.returns.iloc[i][j]) == True and op.returns.iloc[i-1][j] != 0:
op.returns.iloc[i][j] = op.returns.iloc[i-1][j]
elif op.returns.iloc[i][j] == -np.inf or op.returns.iloc[i][j] == np.inf:
op.returns.iloc[i][j] = (op.returns.iloc[i-1][j]+op.returns.iloc[i-2][j])/2
op.returns.fillna(0, inplace=True)
op.reduce_PCA()
op.plot_loadings()
op.find_pairs()
op.pairs
op.plot_clusters(n_dimensions=3)
op.calc_eg_norm_spreads()
op.calc_hurst_exponents()
op.calc_half_lives()
op.calc_avg_cross_count()
op.filter_pairs()
op.filtered_pairs
#for i in op.filtered_pairs.index:
#op.plot_pair_price_spread(idx=i)
op.plot_pair_price_spread(idx=511)
op.alpha
op.norm_spreads
# +
buy = 0
sell = 0
par = 26
for i in range(len(op.norm_spreads)):
if buy == 0 and sell == 0:
if op.norm_spreads[par][i] > 1:
print('Entrada', op.alpha, op.beta, op.norm_spreads.index[i])
print('compra', op.filtered_pairs.loc[par]['pair'][1], '\n', 'venda', op.filtered_pairs.loc[par]['pair'][0])
sell = 1
if buy == 0 and sell == 0:
if op.norm_spreads[par][i] < -1:
print('Entrada', op.alpha, op.beta, op.norm_spreads.index[i])
print('compra', op.filtered_pairs.loc[par]['pair'][0], '\n', 'venda', op.filtered_pairs.loc[par]['pair'][1])
buy = 1
if buy == 1 and op.norm_spreads[par][i] > 0:
buy = 0
print('compra finalizada', op.norm_spreads.index[i])
elif sell == 1 and op.norm_spreads[par][i] < 0:
sell = 0
print('venda finalizada', op.norm_spreads.index[i])
# -
op.filtered_pairs.loc[136]['pair'][0]
| pca.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:qiime2]
# language: python
# name: conda-env-qiime2-py
# ---
from gzip import GzipFile
from collections import Counter
import csv
barcodes = Counter()
with GzipFile('../data/mock-6/s_6_1_sequences_barcodes.fastq.gz', 'r') as barcode_file:
for i, barcode in enumerate(barcode_file):
if i % 4 == 1:
barcodes[barcode.decode('utf-8').strip()] += 1
candidates = sorted([list(reversed(b)) for b in barcodes.items()], reverse=True)[:67]
candidates
with open('../processed/mock-6/rev_eng_map.tsv', 'w') as mapping_file:
writer = csv.writer(mapping_file, delimiter='\t')
writer.writerow(['#SampleID', 'BarcodeSequence'])
i = 1
for n, barcode in candidates:
if 'N' in barcode:
break
writer.writerow(['Barcode%02d' % i, barcode])
i += 1
# !cat ../processed/mock-6/rev_eng_map.tsv
| nbk/reverse_engineer_mock6_barcodes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
from azureml.core import Workspace, Experiment
from azureml.core.conda_dependencies import CondaDependencies
from azureml.core.compute import AmlCompute, ComputeTarget
from azureml.data.data_reference import DataReference
from azureml.core.runconfig import RunConfiguration
from azureml.core import ScriptRunConfig
from azureml.widgets import RunDetails
import json
# +
with open('config/config.json', 'r') as f:
config = json.loads(f.read())
subscription_id = config["SUBSCRIPTION_ID"]
resource_group = config["RESOURCE_GROUP"]
workspace_name = config["WORKSPACE_NAME"]
gpu_cluster_name = config["GPU_CLUSTER_NAME"]
ws = Workspace(workspace_name=workspace_name, subscription_id=subscription_id, resource_group=resource_group)
# +
scripts_folder = "scripts"
if gpu_cluster_name in ws.compute_targets:
gpu_cluster = ws.compute_targets[gpu_cluster_name]
if gpu_cluster and type(gpu_cluster) is AmlCompute:
print('Compute target found. Using: ' + gpu_cluster_name)
else:
print("Creating new cluster")
# vm_size parameter below could be modified to one of the RAPIDS-supported VM types
provisioning_config = AmlCompute.provisioning_configuration(vm_size = "Standard_NC6s_v2", min_nodes=1, max_nodes = 1)
# create the cluster
gpu_cluster = ComputeTarget.create(ws, gpu_cluster_name, provisioning_config)
gpu_cluster.wait_for_completion(show_output=True)
# +
file_root = 'unswiot'
ds = ws.get_default_datastore()
# data already uploaded to the datastore
data_ref = DataReference(data_reference_name='data', datastore=ds, path_on_datastore=file_root)
# -
run_config = RunConfiguration()
run_config.framework = 'python'
run_config.environment.python.user_managed_dependencies = True
run_config.environment.python.interpreter_path = '/conda/envs/rapids/bin/python'
run_config.target = gpu_cluster_name
run_config.environment.docker.enabled = True
run_config.environment.docker.gpu_support = True
run_config.environment.docker.base_image = "todrabas/mlads_rapids:latest"
# run_config.environment.docker.base_image = "rapidsai/rapidsai:cuda9.2-runtime-ubuntu18.04"
run_config.environment.spark.precache_packages = False
run_config.data_references={'data':data_ref.to_config()}
# +
src = ScriptRunConfig(source_directory=scripts_folder,
script='3_Rapids_flow_classification.py',
arguments = ['--data_dir', str(data_ref)],
run_config=run_config
)
exp = Experiment(ws, 'rapidstest_iot_flow_gpu')
run = exp.submit(config=src)
run.wait_for_completion(show_output=True)
| notebooks/intro/Mortgage/3_Rapids_flow_classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
# +
import os
import sys
import numpy as np
import pandas as pd
import plotly as pl
# +
sys.path.insert(0, "..")
import ccal
np.random.random(20121020)
pl.offline.init_notebook_mode(connected=True)
# +
grch_directory_path = os.path.expanduser("~/grch")
people_directory_path = os.path.expanduser("~/people")
reference_fasta_gz_file_path = "{}/Homo_sapiens.GRCh38.dna.primary_assembly.fa.gz".format(
grch_directory_path
)
assert os.path.isfile(reference_fasta_gz_file_path)
reference_gff3_gz_file_path = os.path.abspath(
"{}/Homo_sapiens.GRCh38.93.chr.gff3.gz".format(grch_directory_path)
)
assert os.path.isfile(reference_gff3_gz_file_path)
vcf_gz_file_path = os.path.abspath("{}/0/genome.vcf.gz".format(people_directory_path))
assert os.path.isfile(vcf_gz_file_path)
# +
from genome.Genome import is_valid_vcf_gz
is_valid_vcf_gz(vcf_gz_file_path)
# +
from genome.Genome import Genome
genome = Genome(
reference_fasta_gz_file_path,
reference_gff3_gz_file_path,
vcf_gz_file_path,
reset=True,
)
genome
# +
genome_dict = genome.explore_genome_by_variant("rs235")
genome_dict
# +
genome_dict = genome.explore_genome_by_variant("rs88888888")
genome_dict
# +
genome_dict = genome.explore_genome_by_gene("KRAS")
genome_dict
# +
genome_dict = genome.explore_genome_by_gene("Kwat")
genome_dict
# +
genome_dict = genome.explore_genome_by_region("8", 0, 800000)
genome_dict
# +
genome_dict = genome.explore_genome_by_region("88888888", 0, 800000)
genome_dict
| notebook/genome.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# This notebook was prepared by [<NAME>](https://github.com/donnemartin). Source and license info is on [GitHub](https://github.com/donnemartin/interactive-coding-challenges).
# # Solution Notebook
# ## Problem: Create a list for each level of a binary tree.
#
# * [Constraints](#Constraints)
# * [Test Cases](#Test-Cases)
# * [Algorithm](#Algorithm)
# * [Code](#Code)
# * [Unit Test](#Unit-Test)
# ## Constraints
#
# * Is this a binary search tree?
# * Yes
# * Should each level be a list of nodes?
# * Yes
# * Can we assume we already have a Node class with an insert method?
# * Yes
# * Can we assume this fits memory?
# * Yes
# ## Test Cases
#
# * 5, 3, 8, 2, 4, 1, 7, 6, 9, 10, 11 -> [[5], [3, 8], [2, 4, 7, 9], [1, 6, 10], [11]]
#
# Note: Each number in the result is actually a node containing the number
# ## Algorithm
#
# We can use either a depth-first or a breadth-first search. Intuitively, it seems like a breadth-first search might be a better fit as we are creating a linked list for each level.
#
# We can use a modified breadth-first search that keeps track of parents as we build the linked list for the current level.
#
# * Append the root to the current level's linked list `current`
# * While the `current` is not empty:
# * Add `current` to `results`
# * Set `parents` to `current` to prepare to go one level deeper
# * Clear `current` so it can hold the next level
# * For each `parent` in `parents`, add the children to `current`
# * Return the results
#
# Complexity:
# * Time: O(n)
# * Space: O(n)
# ## Code
# %run ../bst/bst.py
class BstLevelLists(Bst):
def create_level_lists(self):
if self.root is None:
return
results = []
current = []
parents = []
current.append(self.root)
while current:
results.append(current)
parents = list(current)
current = []
for parent in parents:
if parent.left is not None:
current.append(parent.left)
if parent.right is not None:
current.append(parent.right)
return results
# ## Unit Test
# %run ../utils/results.py
# +
# %%writefile test_tree_level_lists.py
from nose.tools import assert_equal
class TestTreeLevelLists(object):
def test_tree_level_lists(self):
bst = BstLevelLists(Node(5))
bst.insert(3)
bst.insert(8)
bst.insert(2)
bst.insert(4)
bst.insert(1)
bst.insert(7)
bst.insert(6)
bst.insert(9)
bst.insert(10)
bst.insert(11)
levels = bst.create_level_lists()
results_list = []
for level in levels:
results = Results()
for node in level:
results.add_result(node)
results_list.append(results)
assert_equal(str(results_list[0]), '[5]')
assert_equal(str(results_list[1]), '[3, 8]')
assert_equal(str(results_list[2]), '[2, 4, 7, 9]')
assert_equal(str(results_list[3]), '[1, 6, 10]')
assert_equal(str(results_list[4]), '[11]')
print('Success: test_tree_level_lists')
def main():
test = TestTreeLevelLists()
test.test_tree_level_lists()
if __name__ == '__main__':
main()
# -
# %run -i test_tree_level_lists.py
| graphs_trees/tree_level_lists/tree_level_lists_solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Correlation
import pandas_datareader as pdr
import datetime as dt
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib notebook
# +
tickers = ['SPY', 'TLT']
start = dt.datetime(2008, 1, 1)
end = dt.datetime(2017, 12, 31)
data = pdr.get_data_yahoo(tickers, start, end)
# -
data = data['Adj Close']
log_returns = np.log(data/data.shift())
log_returns.corr()
fig, ax = plt.subplots()
(data/data.iloc[0]).plot(ax=ax)
# +
data_set = data.loc['2008-05':'2011-04']
fig, ax = plt.subplots()
(data_set/data_set.iloc[0]).plot(ax=ax)
# -
# ### Project
# - Calculate the return (CAGR), maximal drawdown and volatility of TLT
# #### Step 1
# - Calculate the return (CAGR) of SPY and TLT
cagr_spy = (data['SPY'].iloc[-1]/data['SPY'].iloc[0])**(1/10) - 1
cagr_tlt = (data['TLT'].iloc[-1]/data['TLT'].iloc[0])**(1/10) - 1
cagr_spy, cagr_tlt
# #### Step 2
# - Calculate the maximum drawdown of SPY and TLT
def max_drawdown(data):
rolling_max = data.cummax()
daily_drawdown = data/rolling_max - 1
max_drawdown = daily_drawdown.cummin().iloc[-1]
return max_drawdown
max_drawdown(data['SPY']), max_drawdown(data['TLT'])
# #### step 3
# - Calculate the volatility
log_returns.std()*(252**0.5)
| Python For Financial Analysis/05 - Correlation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Modeling and Simulation in Python
#
# Starter code for the orbit example
#
# Copyright 2017 <NAME>
#
# License: [Creative Commons Attribution 4.0 International](https://creativecommons.org/licenses/by/4.0)
#
# <NAME>- Studio 2
# +
# Configure Jupyter so figures appear in the notebook
# %matplotlib inline
# Configure Jupyter to display the assigned value after an assignment
# %config InteractiveShell.ast_node_interactivity='last_expr_or_assign'
# import functions from the modsim.py module
from modsim import *
from numpy import *
# +
# units we'll need
s = UNITS.second
N = UNITS.newton
kg = UNITS.kilogram
m = UNITS.meter
# +
# And an initial condition (with everything in SI units)
r_0 = 4436.82e9 * m
v_0 = -6100 * m/s
init = State(x = r_0,
y = 0 * m,
vx = 0 * m / s,
vy = v_0)
# +
# Making a system object
r_pluto = 1187000 * m
r_sun = 695.508e6 * m
system = System(init=init,
G=6.674e-11 * N / kg**2 * m**2,
m1=1.989e30 * kg,
r_final=r_sun + r_pluto,
m2=0.01303e24 * kg,
t_0=0 * s,
t_end=7820000 * s,
slices = 7)
# +
# function that computes the force of gravity
def universal_gravitation(state, system):
"""Computes gravitational force.
state: State object with distance r
system: System object with m1, m2, and G
"""
x, y, vx, vy = state
unpack(system)
r = Vector(x, y)
v = Vector(vx, vy)
force_mag = G * m1 * m2 / r.mag**2
force = r.hat() * force_mag
# force = Vector(G*m1*m2/x**2, G*m1*m2/y**2)
return force
# -
universal_gravitation(init, system)
# +
def slope_func(state, t, system):
"""Compute derivatives of the state.
state: position, velocity
t: time
system: System object containing `g`
returns: derivatives of y and v
"""
x, y, vx, vy = state
unpack(system)
force_x, force_y = universal_gravitation(state, system)
dxdt = vx
dydt = vy
dvxdt = -force_x / m2
dvydt = -force_y / m2
return dxdt, dydt, dvxdt, dvydt
# +
#test the slope function
slope_func(init, 0, system)
# +
# Event function that stops the simulation upon collision
def event_func(state, t, system):
x, y, vx, vy = state
position = Vector(x,y)
return position.mag - system.r_final
# -
events = event_func(init, 0, system)
dt = (system.t_end - system.t_0)/ system.slices
times = (linrange(system.t_0, system.t_end, dt))
# +
# run the simulation
results, details = run_ode_solver(system, slope_func, events=event_func, t_eval = times )
details
# -
results
# +
#conversions
day = results.index/86400
def convert_m(meters):
#meters to kilometers
km = meters/1000
#kilometers to million kilometers
Mkm = km/1e6
return Mkm
# -
#Zero Initial Velocity
plot(day,convert_m(results.x), label = 'X postion')
plot(day,convert_m(results.y), label = 'Y positon')
decorate(title = 'Intitial Velocity = 0m/s',
xlabel='Time (days)',
ylabel='Distance from sun (million km)')
# +
plot(convert_m(results.x),convert_m(results.y), label ='Path')
decorate(title = 'Intitial Velocity = -30,300 m/s',
xlabel='Distance from sun (million km)',
ylabel='Distance from sun (million km)')
# +
S = results.index[0]
E = results.index[1]
a_vec = Vector(results.x[results.index[0]], results.y[results.index[0]])
A = a_vec.mag
b_vec = Vector(results.x[results.index[1]], results.y[results.index[1]])
B = b_vec.mag
C = sqrt((results.x[results.index[1]]-results.x[results.index[0]])**2 +(results.y[results.index[1]]-results.y[results.index[0]])**2 )
SP = (A+B+C)/2
area = sqrt(SP * (SP - A) * (SP - B) * (SP - C))
state= State(S=S, E=E, A=A, B=B, C=C, SP=SP, area=area)
# -
def area_of_sector(results,system,state):
unpack(system)
dt = (t_end-t_0)/slices
t_interval = linrange(t_0,t_end,dt)
frame = TimeFrame(columns = state.index)
frame.row[1] = state
reps = linrange (1, slices)
for t in reps:
frame.row[t+1] = table_row(frame.row[t], t, system, results)
return frame
# +
def table_row(state, t , system, results):
s, e, a, b, c, sp, area = state
unpack(system)
dt = (t_end-t_0)/slices/UNITS.s
news = e
newe = news + dt
a_vec = Vector(results.x[results.index[t-1]], results.y[results.index[t-1]])
newa = a_vec.mag
b_vec = Vector(results.x[results.index[t]], results.y[results.index[t]])
newb = b_vec.mag
newc = sqrt((results.x[results.index[t]]-results.x[results.index[t-1]])**2 +(results.y[results.index[t]]-results.y[results.index[t-1]])**2 )
newsp = (newa + newb + newc)/2
newarea = sqrt(newsp * (newsp - newa) * (newsp - newb) * (newsp - newc))
return State (S=news, E=newe, A=newa, B=newb, C=newc, SP=newsp, area=newarea)
# -
numbs = area_of_sector(results, system, state)
results
plot(numbs.area)
numbs.S[1]-numbs.E[1]
numbs.S[2]-numbs.E[2]
numbs.S[3]-numbs.E[3]
| code/Proj3-ploto.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Dependencies and Setup
import pandas as pd
import numpy as np
import warnings
warnings.filterwarnings('ignore')
warnings.filterwarnings('ignore', category=DeprecationWarning)
# -
# Files to Load
enroll1516_data_to_load = "SCHOOL_ENROLL.xls"
PIpercent1516_data_to_load = "1516_BUILDING_ACHIEVEMENT.xls"
VA1516_data_to_load = "1516_VA_org_DETAILS.xls"
enroll1516_data = pd.read_excel(enroll1516_data_to_load,sheet_name=[0, 1, 2])
PIpercent1516_data = pd.read_excel(PIpercent1516_data_to_load)
VA1516_data = pd.read_excel(VA1516_data_to_load)
#Function that filters data by District
def data_filter(df):
return df[df["District IRN"] == 43786]
enroll1516_df0= data_filter(enroll1516_data[0])
enroll1516_df1= data_filter(enroll1516_data[1])
PIpercent1516_df=data_filter(PIpercent1516_data)
VA1516_df=data_filter(VA1516_data)
#replace NC values with zero
enroll1516_df0.replace('NC', '0', inplace=True)
enroll1516_df1.replace('NC', '0', inplace=True)
VA1516_df.replace('NC', '0', inplace=True)
PIpercent1516_df.replace('NC', '0', inplace=True)
#replace <10 value with zero
enroll1516_df0.replace('^.*<.*$', '0', regex=True, inplace=True)
enroll1516_df1.replace('^.*<.*$', '0', regex=True, inplace=True)
VA1516_df.replace('^.*<.*$', '0', regex=True, inplace=True)
PIpercent1516_df.replace('^.*<.*$', '0', regex=True, inplace=True)
# Function to concatenate all DataFrames from each Excel sheet into one single pandas DataFrame
def concat_function(df1, df2):
df_concat1 = pd.concat([df1, df2])
return pd.concat([df_concat1])
#Run concat function for 2015-2016,2016-2017 data
cmsd_enroll1516 = concat_function(enroll1516_df0, enroll1516_df1)
#Convert values into integers for Enrollment columns only
cmsd_enroll1516= cmsd_enroll1516.astype({'Pre-School Enrollment': 'int', 'Kindergarten Enrollment':'int',
'First Grade Enrollment': 'int', 'Second Grade Enrollment': 'int',
'Third Grade Enrollment': 'int', 'Fourth Grade Enrollment': 'int',
'Fifth Grade Enrollment':'int', 'Sixth Grade Enrollment': 'int',
'Seventh Grade Enrollment':'int', 'Eighth Grade Enrollment':'int',
'Ninth Grade Enrollment ': 'int', 'Tenth Grade Enrollment': 'int',
'Eleventh Grade Enrollment': 'int', 'Twelfth Grade Enrollment': 'int',
'Thirteenth Grade Enrollment': 'int', 'Enrollment Past Twelfth Grade (Students with Disabilities)': 'int'})
#Convert values into integers for Performance Index Percent column
PI1516_df= PIpercent1516_df.astype({'Performance Index Percent 2015-16':'float'})
#create a new column for enrollment total and use iloc to add from last column Preschool enrollment
cmsd_enroll1516['Enrollment 2015-2016'] = cmsd_enroll1516.iloc[:, -17:-1].sum(1)
#Keep necessary columns, drop all others
cmsd_enroll1516 = cmsd_enroll1516[['Building Name','Building IRN','Enrollment 2015-2016']]
VA1516_df= VA1516_df[['Building Name','Building IRN','Overall Value Added Grade']]
PI1516_df= PI1516_df[['Building Name','Building IRN','Performance Index Percent 2015-16']]
# Function to merge three dataframes into a single DataFrame
from functools import reduce
dfl=[cmsd_enroll1516, PI1516_df, VA1516_df]
cmsd_combined = reduce(lambda left,right: pd.merge(left,right,on=['Building IRN'],how='outer'), dfl)
cmsd_combined
#Create function that drops _y suffixes
def drop_y(df):
# list comprehension of the columns that end with _y
to_drop = [x for x in df if x.endswith('_y')]
df.drop(to_drop, axis=1, inplace=True)
#Create function that drops _x suffixes
def drop_x(df):
# list comprehension of the columns that end with _x
to_drop = [x for x in df if x.endswith('_x')]
df.drop(to_drop, axis=1, inplace=True)
#Run drop_y function for 1516 and 1617 data
drop_x(cmsd_combined)
drop_y(cmsd_combined)
cmsd_combined
#Add new column School Year
cmsd_combined_df= cmsd_combined.assign(School_Year = '2015-2016')[['School_Year']+cmsd_combined.columns.tolist()]
cmsd_combined_df
# +
grouped_date_df = cmsd_combined_df.groupby(['School_Year','Building Name', 'Overall Value Added Grade'])
df5=grouped_date_df.mean()
df5.head(20)
# -
df5.to_excel("cmsd1516.xls")
| Data_cleaned.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import torch
import wordfreq
import math
from transformers import XLNetTokenizer, XLNetLMHeadModel
# + tags=[]
model = XLNetLMHeadModel.from_pretrained("xlnet-base-cased")
tokenizer = XLNetTokenizer.from_pretrained("xlnet-base-cased")
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
# -
origStr = "I mean, when you go to a movie and it’s set to start at a certain time, would you not be upset if 7 hours later said movie has not started?"
# "<mask> <mask> <mask> <mask> <mask> <mask> a <mask> and <mask> <mask> <mask> <mask> <mask> <mask> <mask> <mask> <mask> <mask> <mask> <mask> <mask> <mask> <mask> <mask> <mask> <mask> <mask> <mask> <mask> <mask>?"
testStr = "I mean, when you go to a <mask> and it’s set to start at a certain time, would you not be upset if 7 hours later said movie has not started?"
encoded_str = tokenizer.encode(testStr)
tokens_tensor = torch.tensor([encoded_str])
tokenizer.convert_ids_to_tokens(encoded_str)
perm_mask = torch.zeros((1, tokens_tensor.shape[1], tokens_tensor.shape[1]), dtype=torch.float)
perm_mask[:, :, 8] = 1.0 # Previous tokens don't see masked token
target_mapping = torch.zeros((1, 1, tokens_tensor.shape[1]), dtype=torch.float) # Shape [1, 1, seq_length] => let's predict one token
target_mapping[0, 0, 8] = 1.0 # Our first (and only) prediction will be the last token of the sequence (the masked token)
model
model.transformer.word_embedding.weight
model.lm_loss.weight is model.transformer.word_embedding.weight
embeddings = model.transformer.word_embedding.weight
embeddings
orig_embeddings = embeddings.detach().numpy().copy()
embeddings.shape
embeddings.requires_grad
with torch.no_grad():
embeddings.copy_(torch.tensor(orig_embeddings))
orig_std = orig_embeddings.std()
orig_std
# Var[X] = v
# Var[a X] = a^2 v
noise = torch.randn_like(embeddings)
noise *= orig_std * .01
noise.std()
with torch.no_grad():
embeddings += noise
# + tags=[]
with torch.no_grad():
outputs = model(tokens_tensor, perm_mask=perm_mask, target_mapping=target_mapping)
next_token_logits = outputs[0][0, 0, :]
# + tags=[]
print(outputs[0].shape)
# + tags=[]
print([tokenizer.convert_ids_to_tokens(index.item()) for index in next_token_logits.topk(10).indices])
# + tags=[]
print([tokenizer.convert_ids_to_tokens(index.item()) for index in next_token_logits.topk(10).indices])
# -
PADDING_TEXT = """In 1991, the remains of Russian Tsar Nicholas II and his family
(except for Alexei and Maria) are discovered.
The voice of Nicholas's young son, Tsarevich <NAME>, narrates the
remainder of the story. 1883 Western Siberia,
a young Grigori Rasputin is asked by his father and a group of men to perform magic.
Rasputin has a vision and denounces one of the men as a horse thief. Although his
father initially slaps him for making such an accusation, Rasputin watches as the
man is chased outside and beaten. Twenty years later, Rasputin sees a vision of
the Virgin Mary, prompting him to become a priest. Rasputin quickly becomes famous,
with people, even a bishop, begging for his blessing. <eod> </s> <eos>"""
START_INDEX = 166 # TODO: change hard-coded value
# +
def computeLogProb(original_text, index, tokens_tensor, perm_mask, target_mapping):
with torch.no_grad():
outputs = model(tokens_tensor, perm_mask=perm_mask, target_mapping=target_mapping)
next_token_logits = outputs[0][0, 0, :]
preds = [tokenizer.convert_ids_to_tokens(index.item()) for index in next_token_logits.topk(5).indices]
next_token_logprobs = next_token_logits - next_token_logits.logsumexp(0)
logProb = next_token_logprobs[tokenizer.convert_tokens_to_ids(original_text[index])].item()
return (preds, logProb, next_token_logprobs)
def computePredsLogProbs(preds, next_token_logprobs):
predLogProbs = []
for i in preds:
predLogProbs.append(next_token_logprobs[tokenizer.convert_tokens_to_ids(i)].item())
return predLogProbs
def bigContext(tokenized_text, index):
encoded_ids = tokenizer.convert_tokens_to_ids(tokenized_text)
tokens_tensor = torch.tensor([encoded_ids])
perm_mask = torch.zeros((1, tokens_tensor.shape[1], tokens_tensor.shape[1]), dtype=torch.float)
perm_mask[:, :, index] = 1.0
target_mapping = torch.zeros((1, 1, tokens_tensor.shape[1]), dtype=torch.float)
target_mapping[0, 0, index] = 1.0
return computeLogProb(tokenized_text, index, tokens_tensor, perm_mask, target_mapping)
def smallContext(tokenized_text, index):
tokens_tensor = torch.tensor([tokenizer.convert_tokens_to_ids(tokenized_text)])
perm_mask = torch.zeros((1, tokens_tensor.shape[1], tokens_tensor.shape[1]), dtype=torch.float)
for i in range(START_INDEX, len(tokenized_text) - 1):
if i != index - 1 and i != index + 1:
perm_mask[:, :, i] = 1.0
target_mapping = torch.zeros((1, 1, tokens_tensor.shape[1]), dtype=torch.float)
target_mapping[0, 0, index] = 1.0
return computeLogProb(tokenized_text, index, tokens_tensor, perm_mask, target_mapping)
def noContext(word):
if word in '.?,:!;\'\"‘’“”|-/\\':
return -1 # FIXME
freq = wordfreq.word_frequency(word, 'en')
if freq == 0:
print("word not found:", word)
return -100
return math.log(freq)
def compute_scores(input_text):
tokenized_text = tokenizer.tokenize(PADDING_TEXT + " " + input_text + "</s>", add_special_tokens=False, return_tensors='pt')
usedModels = ["bigContext", "smallContext", "noContext"]
results = []
compoundBigPreds = []
compoundSmallPreds = []
compoundBigLogProb = 0
compoundSmallLogProb = 0
currentWord = ""
startID = 0
# For each token not in PADDING_TEXT
for i in range(START_INDEX, len(tokenized_text) - 1):
# Compute the top 5 model predictions, the log probability of the
# correct answer, and the next_token_logprobs
bigPreds, bigLogProb, bigNextLogProbs = bigContext(tokenized_text, i)
smallPreds, smallLogProb, smallNextLogProbs = smallContext(tokenized_text, i)
# Generate the log probabilities of the top 5 small model predictions
# given big context vs. small context
bigPredsLogProbs = computePredsLogProbs(smallPreds, bigNextLogProbs)
smallPredsLogProbs = computePredsLogProbs(smallPreds, smallNextLogProbs)
# if the current token is a start token
if tokenized_text[i].startswith("▁"):
compoundBigLogProb = bigLogProb
compoundSmallLogProb = smallLogProb
compoundBigPreds = bigPreds
compoundSmallPreds = smallPreds
currentWord = tokenized_text[i]
startID = i
# If the current token is a continuation token
else:
compoundBigLogProb += bigLogProb
compoundSmallLogProb += smallLogProb
currentWord += tokenized_text[i]
# if the next token is not a start token or the end of sequence, don't do any more work
# because that means the next token is a continuation token
if not (tokenized_text[i + 1].startswith("▁") or tokenized_text[i + 1] == "</s>"):
continue
currentWord = currentWord.replace("▁", "")
# Compute the no-context log probabilities of the current word and
# the predictions generated by the small context model
noContextLogProb = noContext(currentWord)
noPredsLogProbs = []
for j in smallPreds:
processed_word = j.replace("▁", "")
noPredsLogProbs.append(noContext(processed_word))
results.append(dict(
id = startID,
word=currentWord,
src="original",
model="smallContext",
score=compoundSmallLogProb)
)
results.append(dict(
id = startID,
word=currentWord,
src="original",
model="bigContext",
score=compoundBigLogProb)
)
results.append(dict(
id = startID,
word=currentWord,
src="original",
model="noContext",
score=noContextLogProb)
)
for j in range(0, len(smallPreds)):
results.append(dict(
id = startID,
word=smallPreds[j],
src="smallContext",
model="smallContext",
score=smallPredsLogProbs[j])
)
results.append(dict(
id = startID,
word=smallPreds[j],
src="smallContext",
model="bigContext",
score=bigPredsLogProbs[j])
)
results.append(dict(
id = startID,
word=smallPreds[j],
src="smallContext",
model="noContext",
score=noPredsLogProbs[j])
)
compoundBigLogProb = 0
compoundSmallLogProb = 0
compoundBigPreds = []
compoundSmallPreds = []
currentWord = ""
return (results, usedModels)
# -
compute_scores("Hello. This is a test.")
| Hyechan/ContextVue2/testXLNet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import json
import os
if not os.path.exists("output"):
os.makedirs("output")
# -
with open("../notebooks/clean.ipynb", "r") as r:
clean = json.load(r)
with open("../notebooks/dirty.ipynb", "r") as r:
dirty = json.load(r)
print(json.dumps(clean, indent=1))
print(json.dumps(dirty, indent=1))
# +
def clean_nb(dirty, outputs_to_remove=[]):
cells = dirty.get('cells', [])
to_remove = set(outputs_to_remove)
for cell in cells:
if cell["cell_type"] != "code": continue
cell["execution_count"] = None
if to_remove: # WIP
ix_to_remove = []
for output in cell["outputs"]:
if 'data' in output:
keys = output['data'].keys()
print(keys)
elif 'name' in output:
print(output)
else:
print(output)
else:
cell["outputs"] = []
clean_nb(dirty, outputs_to_remove=["stdout", "text/plain"])
#print(json.dumps(dirty, indent=1))
# -
with open("output/originally_clean.ipynb", "w") as w:
json.dump(clean, w, indent=4)
with open("output/clean.ipynb", "w") as w:
json.dump(dirty, w, indent=4)
| playground/examine_notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ## Ejercicios de Matplotlib
# 1. Importa pyplot, numpy y pandas
import matplotlib as mpl
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
# 2. Activa matplotlib de forma estática
# %matplotlib inline
# 3. Sabemos que podemos pintar gráficas de dos formas: la figura incluye los ejes o teniendo figura y ejes por separado.
# Usando solo una figura, usa numpy para los valores del eje X entre 0 y 5. Pinta dos gráficas en dos cajas distintas, a la izquierda una recta con pendiente positiva de 3 que pase por (0,0) y a la derecha una recta con pendiente negativa de 3 que pase por (0,-5). Elige la precisión en el eje X que desees.
# +
plt.style.use('seaborn-whitegrid')
plt.figure()
x1 = np.linspace(-2,2,1000)
plt.subplot(1,2,1)
plt.xlim(-5,5)
plt.ylim(-5, 5)
plt.plot(x1, x1)
x2 = np.linspace(-2, 2,1000)
y2 = np.linspace(-2, -7,1000)
plt.subplot(1,2,2)
plt.xlim(-2, 3 )
plt.ylim(-8, 1)
plt.plot(x2, y2);
# -
# 4. Fija el eje X entre 0 y 5 y el eje Y entre -15 y 15
# +
x1 = np.linspace(-2,2,1000)
plt.xlim(0,5 )
plt.ylim(-15, 15)
plt.subplot(1,2,1)
plt.plot(x1, x1)
x2 = np.linspace(-2, 2,1000)
y2 = np.linspace(-2, -7,1000)
plt.subplot(1,2,2)
plt.plot(x2, y2);
# -
# 5. Llama al eje X "eje X", al eje Y "eje Y" y pon de títulos "recta sube" y "recta baja". Muestra dos etiquetas de ejeX pero solo una de eje Y.
# +
x1 = np.linspace(-2,2,1000)
plt.xlim(0,5 )
plt.ylim(-15, 15)
plt.subplot(1,2,1)
plt.title("Recta sube")
plt.xlabel("eje X")
plt.ylabel("eje Y");
plt.plot(x1, x1)
x2 = np.linspace(-2, 2,1000)
y2 = np.linspace(-2, -7,1000)
plt.subplot(1,2,2)
plt.title("Recta baja")
plt.xlabel("eje X")
plt.ylabel("eje Y");
plt.plot(x2, y2);
# -
# Vamos a pintar lo mismo pero accediendo directamente a los ejes
# 6. Usando una figura Y EJES POR SEPARADO, usa numpy para los valores del eje X entre 0 y 5. Pinta dos gráficas en dos cajas distintas, a la izquierda una recta con pendiente positiva de 3 que pase por (0,0) y a la derecha una recta con pendiente negativa de 3 que pase por (0,-5). Elige la precisión en el eje X que desees.
# +
# obtén solo la figura y los ejes y mira el resultado
fig, ax = plt.subplots(1,2)
#ax[0].plot(x, np.sin(x))
#ax[1].plot(x, np.cos(x))
x1 = np.linspace(-2,2,1000)
ax[0].set_xlim(0,5 )
ax[0].set_ylim(-15, 15)
#plt.subplot(1,2,1)
ax[0].set_title("Recta sube")
ax[0].set_xlabel("eje X")
ax[0].set_ylabel("eje Y");
ax[0].plot(x1, x1)
x2 = np.linspace(-2, 2,1000)
y2 = np.linspace(-2, -7,1000)
ax[1].set_title("Recta baja")
ax[1].set_xlabel("eje X")
ax[1].set_ylabel("eje Y");
ax[1].plot(x2, y2);
# +
# basándote en el código previo, cambia solo la manipulación de los ejes
# recuerda que hay unos métodos que su nombre cambia un poco
# -
# 7. Existen distintos modelos de gráficas.
# Crea una lista de coordenadas X: 20, 22, 24, 26, 28
# Crea una lista de coordenadas Y: 5, 15, -5, 20, 5
# Usa un gráfico de barras colocadas en X con alturas Y
# +
x = [20, 22, 24, 26, 28]
y = [5, 15, -5, 20, 5]
plt.bar(x,y,align='center')
plt.xlabel('X')
plt.ylabel('Y')
plt.show()
# -
# 8. Crea un gráfico de barras. Pintamos la altura de un grupo de amigos, cada barra representa a una persona. Ana mide 160 cm, Luis mide 180 cm, Pedro mide 175 cm, Sofía mide 190 cm, Carmen mide 170 cm. Las barras serán verdes.
# Consejo: si no vas a tener que manipular especialmente los ejes, es más sencillo dejarlos dentro de la figura.
# +
x = ['Ana', 'Luis', 'Pedro', 'Sofia', 'Carmen']
y = [160, 180, 175, 190, 170]
plt.style.use('seaborn-white')
plt.bar(x,y,align='center', color='g')
plt.xlabel('Amigos')
plt.ylabel('Alturas')
plt.show()
# -
# 9. Basándote en el gráfico anterior, escribe encima de cada barra la altura de cada amigo.
# Pista: usa un bucle que lea cada barra de barplot = plt.bar(x,y)
# bar tiene los métodos get_height(), get_x(), get_width()
# plt.text(x,y,valor, va='bottom')
# +
x = ['Ana', 'Luis', 'Pedro', 'Sofia', 'Carmen']
y = [160, 180, 175, 190, 170]
plt.style.use('seaborn-white')
barplot = plt.bar(x,y,align='center', color='g')
plt.xlabel('Amigos')
plt.ylabel('Alturas')
for bar in barplot:
plt.text(bar.get_x()+0.30,bar.get_height()+7,str(bar.get_height()), va='top')
plt.show()
# -
# 10. Cambia el tamaño de la figura a (3,5)
# +
x = ['Ana', 'Luis', 'Pedro', 'Sofia', 'Carmen']
y = [160, 180, 175, 190, 170]
plt.style.use('seaborn-white')
plt.figure(figsize=(3,5))
plt.xlabel('Amigos')
plt.ylabel('Alturas')
barplot = plt.bar(x,y,align='center', color='g')
for bar in barplot:
plt.text(bar.get_x()+0.30,bar.get_height()+7,str(bar.get_height()), va='top')
plt.show()
# -
# 11. Basándote en el gráfico anterior, borra la escala del eje Y (ya aparece en la altura)
# Pista: cuando plt.yticks() recibe una lista vacía no pinta el eje Y
# +
x = ['Ana', 'Luis', 'Pedro', 'Sofia', 'Carmen']
y = [160, 180, 175, 190, 170]
plt.style.use('seaborn-white')
plt.figure(figsize=(3,5), frameon = False)
plt.xlabel('Amigos')
plt.ylabel('Alturas')
plt.yticks([])
plt.xticks(rotation=90)
plt.box(False)
barplot = plt.bar(x,y,align='center', color='g')
for bar in barplot:
plt.text(bar.get_x(),bar.get_height()+7,str(bar.get_height()), va='top')
plt.show()
# -
# 12. Prueba a hacer el gráfico con las barras en horizontal (no es necesario poner el texto al final de la barra).
# +
x = ['Ana', 'Luis', 'Pedro', 'Sofia', 'Carmen']
y = [160, 180, 175, 190, 170]
plt.style.use('seaborn-white')
plt.figure(figsize=(3,5), frameon = False)
plt.ylabel('Amigos')
plt.xlabel('Alturas')
plt.yticks([])
plt.xticks(rotation=90)
plt.box(False)
barplot = plt.barh(x,y,align='center', color='g')
for i, bar in enumerate(barplot):
plt.text(205,bar.get_y()+0.5,x[i], va='top')
plt.text(205,bar.get_y()+0.3,y[i], va='top')
plt.show()
# -
# 13. Prueba a invertir los ejes con ax.invert_axis()
# Necesitarás tener los ejes disponibles fuera de la figura
# 14. En el último gráfico, cambia el estilo a 'dark_background'
# +
x = ['Ana', 'Luis', 'Pedro', 'Sofia', 'Carmen']
y = [160, 180, 175, 190, 170]
plt.style.use('dark_background')
plt.figure(figsize=(3,5), frameon = False)
plt.ylabel('Amigos')
plt.xlabel('Alturas')
plt.yticks([])
plt.xticks(rotation=90)
plt.box(False)
barplot = plt.barh(x,y,align='center', color='g')
for i, bar in enumerate(barplot):
plt.text(205,bar.get_y()+0.5,x[i], va='top')
plt.text(205,bar.get_y()+0.3,y[i], va='top')
plt.show()
# -
# 15. Vamos a introducir pandas. Crea un DataFrame con las columnas "Year" de valores 2015, 2016, 2017, 2018, 2019 y la columna "Sold_items_A" de valores 1000, 3500, 4000, 5500, 7000
# 16. Ahora pintamos un gráfico de línea con las ventas respecto al año. Pon un título y etiquetas en los ejes.
# Cambia el estilo a 'seaborn-white'. La línea debe ser con rayas y verde.
# 17. Otro departamento B ha vendido en esos años 2000, 3100, 5000, 4000, 6000 unidades. Incluye esa columna en el DataFrame y pinta en la misma gráfica las dos líneas. B es una línea punteada y roja. Muestra una leyenda abajo a la derecha.
# +
# df
# -
# 18. Haz un scatter del departamento A usando __solo el DataFrame__.
# Pista: el propio DataFrame tiene un método plot.
# df.plot('columna X', 'columna Y', 'kind' = 'scatter)
# +
# -
# 19. Prueba a cambiar kind a 'pie'
# 20. Prueba a quitar la leyenda incluyendo legend igual a False, añade labels y quita la etiqueta en Y
# 21. Vamos a pintar un histograma
# +
np.random.seed(1) # cada vez que le pida N números aleatorios, me dará los mismos
mydf = pd.DataFrame({"Altura" : np.random.randint(low=150, high=190, size=300)})
# -
# Pista: https://pandas.pydata.org/pandas-docs/version/0.23.4/generated/pandas.DataFrame.plot.html
# +
# Histogram
# -
# Nota: con ax = df.plot()
#
# se pueden poner las etiquetas con
#
# ax.set(xlabel="Bins")
# 22. Contornos de 3D a 2D. Escribe una función que recibiendo x,y devuelva (1 - x / 2 + x ** 5 + y ** 3) * np.exp(-x ** 2 -y ** 2)
#
# Tablero: Tanto x como y van de -3 a 3 y usaremos 256 puntos.
# Usa contourf, con 8 niveles (cortes), una transparencia de 0.75 y un color map de tipo 'jet'.
# Bonus: pinta las líneas de los contornos también de negro
# +
# ESTE NO
| 2-EDA/3-Matplotlib/practica/FedeRuiz_ejercicios_Matplotlib.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# ## Question 2 [Leave-One-Out Cross-Validation, 15 Marks]
# ### Load Libraries
# libraries
library(reshape2)
library(ggplot2)
# import data
train_1a <- read.csv("./Task1A_train.csv")
test_1a <- read.csv("./Task1A_test.csv")
# Knn function from question 1
# KNN function (distance = manhattan)
knn <- function(train.data, train.label, test.data, K=K, distance = 'manhattan'){
## count number of train samples
train.len <- nrow(train.data)
## count number of test samples
test.len <- nrow(test.data)
## calculate distances between samples
dist <- as.matrix(dist(rbind(test.data, train.data), method= distance))[1:test.len, (test.len+1):(test.len+train.len),
drop = FALSE]
## for each test sample...
for (i in 1:test.len){
### ...find its K nearest neighbours from training sampels...
nn <- as.data.frame(sort(dist[i,], index.return = TRUE))[1:K,2]
###... and calculate the predicted values according to average.
test.label[i]<- sum(train.label[nn])/K
}
## return the predictions as output
return (test.label)
}
# ### 1. Implement a Leave-One-Out cross-validation (CV) function for your KNN regressor: cv(train.data, train.label, K)
# +
# LOOCV function
cv <- function(train.data,train.label, K){
# define cv folds
folds <- nrow(train.data)
# initialize error
SE <- 0
# run cv for each data point in train data
for (i in 1:folds){
# define variables
train_x <- train.data[-i,,drop = FALSE]
train_y <- train.label[-i,]
test_x <- train.data[i,,drop = FALSE]
test_y <- train.label[i,]
# perform knn
knn_predict <- knn(train_x,train_y,test_x, K=K)
# calculate and add up error
SE <- SE + (knn_predict-test_y)^2
}
# calculate average RMSE
RMSE <- sqrt(SE/folds)
return (RMSE)
}
# -
# ### 2. Run LOOCV for k = 1:20 and plot error vs 1/k
# extract x and y values of train data as single column
train_1a_x <- train_1a[,1,drop = FALSE]
train_1a_y <- train_1a[,2,drop = FALSE]
test.label <- data.frame("y" = 0)
# +
# create a table to store RMSE for k = 1:20
RMSE_cv <- data.frame('K'=1:20, 'RMSE'=rep(0,20))
# calcualte rmse for k = 1:20
for (k in 1:20){
RMSE_cv[k,"RMSE"] <- cv(train_1a_x,train_1a_y,k)
}
# plot the errors vs log(1/k) values
ggplot(data=RMSE_cv, aes(x=log(1/K), y=RMSE)) + geom_line() +
scale_color_discrete(guide = guide_legend(title = NULL)) + theme_minimal() +
ggtitle("Root mean square error")
# -
# inspect errors for k 1:20
RMSE_cv
# ### 3. Report the optimum value for K
#
# The plot shows, root mean square errors decrease as k decreases, the lower the k value, the smaller the error. There is one exception around log(1/k) = -0.75, where RMSE increase significantly. Also, RMSE increase significantly when log(1/k) > -1.5. Thus, any point between -1.5 and -0.75 is acceptable. In this case, the optimal k value that produce the smallest RMSE shown on the plot is at k = 2.
| Machine Learning study notes/Linear Models for Classification and regression/src/31436285_assessment_1_q2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="bjOGToW5Yk9A" colab_type="code" colab={}
import numpy as np
import matplotlib.pyplot as plt
from sklearn import linear_model, svm
from scipy.special import expit
# + [markdown] id="5w0t8CNhZdQB" colab_type="text"
# #Generate Data
# + id="hPjxf5lSZrwJ" colab_type="code" colab={}
# General a toy dataset:s it's just a straight line with some Gaussian noise:
xmin, xmax = -5, 5
n_samples = 100
np.random.seed(0)
X = np.random.normal(size=n_samples)
y = (X > 0).astype(np.float)
X[X > 0] *= 4
X += .3 * np.random.normal(size=n_samples)
X = X[:, np.newaxis]
# + [markdown] id="6eerJWjLlSUK" colab_type="text"
# # Fit Different Classifiers
#
#
# * Linear Regression
# * Logistic Regression
# * Linear SVM
#
#
# + id="kgpstIdga3vC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 225} outputId="1e23b4fb-98a9-4e10-8a2f-efe6303ab0b4"
#part of code is taken from https://scikit-learn.org/stable/auto_examples/linear_model/plot_logistic.html#sphx-glr-auto-examples-linear-model-plot-logistic-py
#plot the result
plt.figure(1, figsize=(4, 3))
plt.clf()
plt.scatter(X.ravel(), y, color='green')#, zorder=20)
X_test = np.linspace(-5, 10, 300)
# Logistic Regression
clf = linear_model.LogisticRegression(C=1e5)
clf.fit(X, y)
plt.plot(X_test, clf.coef_[0]*X_test + clf.intercept_, color='red', linewidth=3, label = 'Logistic Regression Model')
# Linear Regression
ols = linear_model.LinearRegression()
ols.fit(X, y)
plt.plot(X_test, ols.coef_ * X_test + ols.intercept_, linewidth=1, label = 'Linear Regression Model')
plt.axhline(.5, color='.5')
# Linear SVM
clf_svm = svm.LinearSVC()
clf_svm.fit(X, y)
plt.plot(X_test, clf_svm.coef_[0] * X_test + clf_svm.intercept_, linewidth=1, label = 'Linear SVM')
plt.axhline(.5, color='.5')
plt.ylabel('y')
plt.xlabel('X')
plt.xticks(range(-5, 10))
plt.yticks([0, 0.5, 1])
plt.ylim(-.25, 1.25)
plt.xlim(-4, 10)
plt.legend(loc="lower right", fontsize='small')
plt.tight_layout()
plt.show()
# + [markdown] id="Lsm-OWgfkL8t" colab_type="text"
# # Accuracy
# + id="C_SglTEAhjZb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="7ba1e248-02cb-4444-fe76-f51266501550"
print(f"Test accuracy Logistic Regression: {clf.score(X_test.reshape(-1,1), Y_test.reshape(-1,1))}")
print(f"Test accuracy Linear Regression: {ols.score(X_test.reshape(-1,1), Y_test.reshape(-1,1))}")
print(f"Test accuracy SVM: {clf_svm.score(X_test.reshape(-1,1), Y_test.reshape(-1,1))}")
# + [markdown] id="v8Bdy_bkmCF8" colab_type="text"
# We can observe from previous plot that SVM and Logistic Regression provides better classfier.
#
# SVM maximises geometric separation between two classes.
# Logistic regression tries to maximise posterior probabilities.
# + id="cMGsWFytmbgK" colab_type="code" colab={}
| SVM_vs_Logistic_regression_vs_Linear_Regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="cac470df-29e7-4148-9bbd-d8b9a32fa570" tags=[]
# # (tip) 아나콘다에서 Jupyter lab 사용하기
# >
#
# - toc:true
# - branch: master
# - badges: true
# - comments: false
# - author: 최서연
# - categories: [anaconda, jupyter lab]
# -
# `conda env list`
# list 보고 싶다.
# `conda remove -n test`
# test라는 환경을 지우겠다
# `conda activate test`
# test 환경에 들어간다.
# `jupyter lab`
# 환경에 들어가서 주피터랩 싱행 후 사용, but, 깔려있어야 하겠지
# `conda deactivate test`
# test라는 환경에서 나온다.
# `conda create -n test`
# test라는 환경을 만든다. 아직 아무것도 없는 상태
# `conda install -c conda-forge jupyterlab`
# conda라는 앱스토어에 들어가서 주피터랩 깔아라, 그런데 conda에서 추천하는 버전으로!!
#
# 중간에 뭐 깔꺼냐고 y/n 선택하라고 물어보는 이유, 적절한 건 있는데 이렇게 설치 정말 할건지!, 그니까 원래 있던 버전 지우고 다른 버전깐다고 물어볼수도..
#
# pip / conda intall 차이점 pip는 최신 패키지만 다운 받음 conda install는 적절한 버전을 다운 받기 가능
# `conda install -c conda forge rise`
# 주피터 노트북으로 발효자 만들기, 슬라이드 쇼 만들 수 있게!
# `jupyter notebook`
# 주피터 노트북으로 들어가자
# 주피터 노트북에서 view에서 slide show~, 슬라이드쇼 하는 중간에 수정도 가능
| _notebooks/2022-05-10-conda-jupyter-lab.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.cm as cm
import pandas.io.sql as sqlio
import psycopg2
from sklearn.cluster import MiniBatchKMeans
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.decomposition import PCA
from sklearn.manifold import TSNE
# -
conn = psycopg2.connect(user="xxxx", password="<PASSWORD>", host="xxxx", database="Ingestion")
query = "select * from itc_obs_0421_savecheck3"
#execute query and save it to a variable
dataset = sqlio.read_sql_query(query,conn)
pd.set_option('display.max_columns', None)
dataset.head()
dataset.shape
# +
df = dataset[['description_of_requirement','level_2_category','level_3_category','co_bus_size_determination',
'business_rule_tier','contract_name']]
df = df.astype({'level_2_category': str})
df = df.astype({'level_3_category': str})
df = df.astype({'co_bus_size_determination': str})
df = df.astype({'contract_name': str})
df = df.astype({'level_3_category': str})
df = df.astype({'description_of_requirement': str})
# -
df["req"] = df[['description_of_requirement','level_2_category','level_3_category','co_bus_size_determination',
'business_rule_tier','contract_name']].apply(lambda x: ' '.join(x), axis = 1)
tfidf = TfidfVectorizer(
min_df = 5,
max_df = 0.95,
max_features = 8000,
stop_words = 'english'
)
tfidf.fit(df.req)
text = tfidf.transform(df.req)
# +
def find_optimal_clusters(df, max_k):
iters = range(2, max_k+1, 2)
sse = []
for k in iters:
sse.append(MiniBatchKMeans(n_clusters=k, init_size=1024, batch_size=2048, random_state=20).fit(df).inertia_)
print('Fit {} clusters'.format(k))
f, ax = plt.subplots(1, 1)
ax.plot(iters, sse, marker='o')
ax.set_xlabel('Cluster Centers')
ax.set_xticks(iters)
ax.set_xticklabels(iters)
ax.set_ylabel('SSE')
ax.set_title('SSE by Cluster Center Plot')
find_optimal_clusters(text, 20)
# -
clusters = MiniBatchKMeans(n_clusters=16, init_size=1024, batch_size=2048, random_state=20).fit_predict(text)
# +
def plot_tsne_pca(data, labels):
max_label = max(labels)
max_items = np.random.choice(range(data.shape[0]), size=3000, replace=False)
pca = PCA(n_components=2).fit_transform(data[max_items,:].todense())
tsne = TSNE().fit_transform(PCA(n_components=50).fit_transform(data[max_items,:].todense()))
idx = np.random.choice(range(pca.shape[0]), size=300, replace=False)
label_subset = labels[max_items]
label_subset = [cm.hsv(i/max_label) for i in label_subset[idx]]
f, ax = plt.subplots(1, 2, figsize=(14, 6))
ax[0].scatter(pca[idx, 0], pca[idx, 1], c=label_subset, label=clusters)
ax[0].set_title('PCA Cluster Plot')
ax[0].legend()
ax[1].scatter(tsne[idx, 0], tsne[idx, 1], c=label_subset, label=clusters)
ax[1].set_title('TSNE Cluster Plot')
ax[1].legend()
plot_tsne_pca(text, clusters)
# +
def plot_tsne_pca(data, labels, sizelist, cmap='tab10'):
max_label = max(labels)
max_items = np.random.choice(range(data.shape[0]), sizelist, replace=False)
pca = PCA(n_components=2).fit_transform(data[max_items, :].todense())
tsne = TSNE().fit_transform(PCA(n_components=1).fit_transform(data[max_items, :].todense()))
idx = np.random.choice(range(pca.shape[0]), sizelist, replace=False)
label_subset = labels[max_items]
#label_subset = [cm.hsv(i / max_label) for i in label_subset[idx]]
f, ax = plt.subplots(1, 2, figsize=(20, 6))
ax[0].scatter(pca[idx, 0], pca[idx, 1], c=label_subset, cmap=cmap)
ax[0].set_title('PCA Cluster Plot')
sc = ax[1].scatter(tsne[idx, 0], tsne[idx, 1], c=label_subset, cmap=cmap)
ax[1].set_title('TSNE Cluster Plot')
ax[1].legend(*sc.legend_elements(), title='clusters')
plot_tsne_pca(text, clusters, sizelist)
# -
# Add clusters label to DF
df['clusters'] = clusters
df.head()
# +
def get_top_keywords(data, clusters, labels, n_terms):
df = pd.DataFrame(data.todense()).groupby(clusters).mean()
for i,r in df.iterrows():
print('\nCluster {}'.format(i))
print(','.join([labels[t] for t in np.argsort(r)[-n_terms:]]))
get_top_keywords(text, clusters, tfidf.get_feature_names(), 10)
| ML/Acquistioners_TSNE_Clustering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # Get the most common word
#
# Now is your turn to do an application, with no _training wheels_.
#
# The exercise is:
#
# > Print the word with most occurrences, alongside its count.
#
# You will, roughly, need to follow the following steps:
#
# - Initialize the `storage` library.
# - Retrieve the persistent `Result` instance.
# - Iterate all the results.
# - Check which word has most occurrences.
# - Print the overall most common word alongside its number of occurrences.
...
| wordcount/03-get-most-common.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import glob
import pathlib
import pickle
import requests
import tarfile
import time
import joblib
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from sklearn import model_selection
import torch
from torch import nn, optim
import torch.nn.functional as F
from torch.utils import data
import torchinfo
import torchmetrics
from torchvision import models, transforms
# -
# # Training Deep Neural Networks using GPUs
# # Data
#
# ## CIFAR-10 Dataset
#
# The original [CIFAR-10](http://www.cs.toronto.edu/~kriz/cifar.html) dataset consists of 60000 32x32 colour images in 10 classes, with 6000 images per class. There are 50000 training images and 10000 test images. The dataset is divided into five training batches and one test batch, each with 10000 images. The test batch contains exactly 1000 randomly-selected images from each class. The training batches contain the remaining images in random order, but some training batches may contain more images from one class than another. Between them, the training batches contain exactly 5000 images from each class.
CLASS_LABELS = {
0: "airplane",
1: "automobile",
2: "bird",
3: "cat",
4: "deer",
5: "dog",
6: "frog",
7: "horse",
8: "ship",
9: "truck"
}
# ### Download and extract the data
# +
DATA_DIR = pathlib.Path("../data/")
RAW_DATA_DIR = DATA_DIR / "cifar-10"
URL = "https://www.cs.toronto.edu/~kriz/cifar-10-python.tar.gz"
RAW_DATA_DIR.mkdir(parents=True, exist_ok=True)
with open(RAW_DATA_DIR / "cifar-10-python.tar.gz", "wb") as f:
response = requests.get(URL)
f.write(response.content)
with tarfile.open(RAW_DATA_DIR / "cifar-10-python.tar.gz", "r:gz") as f:
f.extractall(RAW_DATA_DIR)
# -
# ### Load the data
#
# We will load the data using the [Pandas](https://pandas.pydata.org/) library. Highly recommend the most recent edition of [*Python for Data Analysis*](https://learning.oreilly.com/library/view/python-for-data/9781491957653/) by Pandas creator <NAME> for anyone interested in learning how to use Pandas.
# +
_data = []
_labels = []
filepaths = glob.glob("../data/cifar-10/cifar-10-batches-py/*_batch*")
for filepath in sorted(filepaths):
with open(filepath, "rb") as f:
_batch = pickle.load(f, encoding="latin1")
_data.append(_batch["data"])
_labels.extend(_batch["labels"])
# each image has 3 channels with height and width of 32 pixels
features = pd.DataFrame(
np.vstack(_data),
columns=[f"p{i}" for i in range(3 * 32 * 32)],
dtype="uint8",
)
target = pd.Series(_labels, dtype="uint8", name="labels")
# -
# ### Explore the data
features.info()
features.head()
target.head()
# ### Visualize the data
# +
fig, axes = plt.subplots(10, 10, sharex=True, sharey=True, figsize=(15, 15))
for i in range(10):
for j in range(10):
m, _ = features.shape
k = np.random.randint(m)
img = (features.loc[k, :]
.to_numpy()
.reshape((3, 32, 32))
.transpose(1, 2, 0))
_ = axes[i, j].imshow(img)
_ = axes[i, j].set_title(CLASS_LABELS[target[k]])
fig.suptitle("Random CIFAR-10 images", x=0.5, y=1.0, fontsize=25)
fig.tight_layout()
# -
# # Creating Train, Val, and Test Data
#
# Before we look at the data any further, we need to create a test set, put it aside, and never look at it (until we are ready to test our trainined machine learning model!). Why? We don't want our machine learning model to memorize our dataset (this is called overfitting). Instead we want a model that will generalize well (i.e., make good predictions) for inputs that it didn't see during training. To do this we hold split our dataset into training and testing datasets. The training dataset will be used to train our machine learning model(s) and the testing dataset will be used to make a final evaluation of our machine learning model(s). We also need to create a validation dataset for tuning hyperparameters and deciding when to stop training.
#
# ## If you might refresh data in the future...
#
# ...then you want to use some particular hashing function to compute the hash of a unique identifier for each observation of data and include the observation in the test set if resulting hash value is less than some fixed percentage of the maximum possible hash value for your algorithm. This way even if you fetch more data, your test set will never include data that was previously included in the training data.
# +
import zlib
def in_holdout_data(identifier, test_size):
_hash = zlib.crc32(bytes(identifier))
return _hash & 0xffffffff < test_size * 2**32
def split_data_by_id(data, test_size, id_column):
ids = data[id_column]
in_holdout_set = ids.apply(lambda identifier: in_holdout_data(identifier, test_size))
return data.loc[~in_holdout_set], data.loc[in_holdout_set]
# -
# ## If this is all the data you will ever have...
#
# ...then you can just set a seed for the random number generator and then randomly split the data. Scikit-Learn has a [`model_selection`](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.train_test_split.html) module that contains tools for splitting datasets. First, split the dataset into training and testing datasets. Next split the training dataset into training and validation datasets.
# +
SEED = 42
SEED_GENERATOR = np.random.RandomState(SEED)
def generate_seed():
return SEED_GENERATOR.randint(np.iinfo("uint16").max)
# +
# split the dataset into training and testing data
_seed = generate_seed()
_random_state = np.random.RandomState(_seed)
_train_features, test_features, _train_target, test_target = model_selection.train_test_split(
features,
target,
test_size=1e-1,
random_state=_random_state
)
train_features, val_features, train_target, val_target = model_selection.train_test_split(
_train_features,
_train_target,
test_size=1e-1,
random_state=_random_state
)
# -
train_features.info()
val_features.info()
test_features.info()
# # Training a Neural Network
#
# When working with GPUs we need to tell PyTorch which device to use when training.
device = torch.device("cuda") if torch.cuda.is_available() else torch.device("cpu")
# Next we need to define the components of our training loop that we developed in this morning session.
# +
def accuracy(output, target):
return torchmetrics.functional.accuracy(output, target)
def partial_fit(model_fn, loss_fn, X_batch, y_batch, opt):
# forward pass
loss = loss_fn(model_fn(X_batch), y_batch)
# back propagation
loss.backward()
opt.step()
opt.zero_grad() # don't forget to reset the gradient after each batch!
def validate(model_fn, loss_fn, data_loader):
with torch.no_grad():
batch_accs = []
batch_losses = []
for X, y in data_loader:
batch_accs.append(accuracy(model_fn(X), y))
batch_losses.append(loss_fn(model_fn(X), y))
avg_accuracy = (torch.stack(batch_accs)
.mean())
avg_loss = (torch.stack(batch_losses)
.mean())
return avg_accuracy, avg_loss
def fit(model_fn, loss_fn, train_data_loader, opt, lr_scheduler, val_data_loader=None, number_epochs=2):
for epoch in range(number_epochs):
# train the model
model_fn.train()
for X_batch, y_batch in train_data_loader:
partial_fit(model_fn, loss_fn, X_batch, y_batch, opt)
# compute validation loss after each training epoch
model_fn.eval()
if val_data_loader is not None:
val_acc, val_loss = validate(model_fn, loss_fn, val_data_loader)
print(f"Training epoch: {epoch}, Validation accuracy: {val_acc}, Validation loss: {val_loss}")
# update the learning rate
lr_scheduler.step()
# -
# In this section we introduce a `CustomDataset` to better encapsulate data preprocessing transformations using PyTorch primitives instead of Scikit-Learn. We also reuse the `LambdaLayer` and the `WrappedDataLoader` classes from this morning session. However, instead of using the `WrappedDataLoader` to implement data preprocessing steps, we will instead use the class to send our training data batches from the CPU to the GPU during the training loop.
# +
class CustomDataset(data.Dataset):
def __init__(self, features, target, transforms = None):
self._data = (features.to_numpy()
.reshape(-1, 3, 32, 32)
.transpose(0, 2, 3, 1))
self._target = target.to_numpy()
self._transforms = transforms
def __getitem__(self, index):
X, y = self._data[index], self._target[index]
return (self._transforms(X), y) if self._transforms is not None else (X, y)
def __len__(self):
return len(self._data)
class LambdaLayer(nn.Module):
def __init__(self, f):
super().__init__()
self._f = f
def forward(self, X):
return self._f(X)
class WrappedDataLoader:
def __init__(self, data_loader, f):
self._data_loader = data_loader
self._f = f
def __len__(self):
return len(self._data_loader)
def __iter__(self):
for batch in iter(self._data_loader):
yield self._f(*batch)
# -
# ## Defining the LeNet-5 architecture
model_fn = nn.Sequential(
nn.Conv2d(3, 6, kernel_size=5, stride=1, padding=0),
nn.Tanh(),
nn.AvgPool2d(kernel_size=2, stride=2),
nn.Conv2d(6, 16, kernel_size=5, stride=1, padding=0),
nn.Tanh(),
nn.AvgPool2d(kernel_size=2, stride=2),
LambdaLayer(lambda X: X.view(X.size(0), -1)),
nn.Linear(400, 120),
nn.Tanh(),
nn.Linear(120, 84),
nn.Tanh(),
nn.Linear(84, 10)
)
_ = model_fn.to(device)
torchinfo.summary(model_fn, input_size=(64, 3, 32, 32))
# ## Train the neural network
# +
# use same loss function from last time
loss_fn = F.cross_entropy
# define some preprocessing transforms (done on CPU!)
_transforms = transforms.Compose([
transforms.ToTensor(),
])
# move the tensor from the CPU to the GPU
_to_device = lambda X, y: (X.to(device), y.to(device))
# define the datasets and dataloaders
_train_dataset = CustomDataset(train_features, train_target, _transforms)
_train_data_loader = data.DataLoader(_train_dataset, batch_size=64, shuffle=True)
train_data_loader = WrappedDataLoader(_train_data_loader, _to_device)
_val_dataset = CustomDataset(val_features, val_target, _transforms)
_val_data_loader = data.DataLoader(_val_dataset, batch_size=128, shuffle=False)
val_data_loader = WrappedDataLoader(_val_data_loader, _to_device)
_test_dataset = CustomDataset(test_features, test_target, _transforms)
_test_data_loader = data.DataLoader(_test_dataset, batch_size=128, shuffle=False)
test_data_loader = WrappedDataLoader(_test_data_loader, _to_device)
# define the optimizer and the learning rate scheduler
opt = optim.SGD(model_fn.parameters(), lr=1e-2, momentum=0.9)
lr_scheduler = optim.lr_scheduler.ExponentialLR(opt, gamma=0.9, verbose=True)
# -
fit(model_fn,
loss_fn,
train_data_loader,
opt,
lr_scheduler,
val_data_loader,
number_epochs=10)
average_accuracy, average_loss = validate(model_fn, loss_fn, test_data_loader)
# ### Exercise: Build your own neural network
#
# Modify the LeNet-5 archtiecture as you see fit in order to gain experience building your own neural network.
# +
# insert code here!
# -
# ### Exercise: Experiment with different batch sizes
#
# Train your model for 10 epochs with different batch sizes: 1, 4, 16, 64, 256. Do you notice any patterns?
# +
# insert code here!
# -
# ### Exercise: Experiment with different learning rate schedulers
#
# Train your model for 10 epochs with different batch size of 64 but experiment with different learning rate schedulers. Does one learning rate scheduler outperform the others?
# +
# insert code here!
# -
# ## Experimenting with different architectures
#
# In practice, it is unlikely that you will be designing your own neural network architectures from scratch. Instead you will be starting from some pre-existing neural network architecture. The [torchvision](https://pytorch.org/vision/stable/) project contains a number of neural network architectures that have found widespread use in computer vision applications.
#
# For the remainder of this notebook we will be using the [ResNet-18](https://arxiv.org/pdf/1512.03385.pdf) model which was developed in 2015. The ResNet family of models were designed to be trained on larger images (224 x 224) and a larger number of classes (1000) so we need to make some small modifications in order to adapt this network for our dataset.
models.
model_fn = models.resnet18(num_classes=10)
model_fn.conv1 = nn.Conv2d(3, 64, kernel_size=(3, 3), stride=(1,1), padding=(1,1), bias=False)
_ = model_fn.to(device)
torchinfo.summary(model_fn, input_size=(64, 3, 32, 32))
# +
# use same loss function from last time
loss_fn = F.cross_entropy
# define some preprocessing transforms (done on CPU!)
_transforms = transforms.Compose([
transforms.ToTensor(),
])
# move the tensor from the CPU to the GPU
_to_device = lambda X, y: (X.to(device), y.to(device))
# define the datasets and dataloaders
_train_dataset = CustomDataset(train_features, train_target, _transforms)
_train_data_loader = data.DataLoader(_train_dataset, batch_size=128, shuffle=True)
train_data_loader = WrappedDataLoader(_train_data_loader, _to_device)
_val_dataset = CustomDataset(val_features, val_target, _transforms)
_val_data_loader = data.DataLoader(_val_dataset, batch_size=256, shuffle=False)
val_data_loader = WrappedDataLoader(_val_data_loader, _to_device)
_test_dataset = CustomDataset(test_features, test_target, _transforms)
_test_data_loader = data.DataLoader(_test_dataset, batch_size=256, shuffle=False)
test_data_loader = WrappedDataLoader(_test_data_loader, _to_device)
# define the optimizer and the learning rate scheduler
opt = optim.SGD(model_fn.parameters(), lr=1e-1, momentum=0.9)
lr_scheduler = optim.lr_scheduler.StepLR(opt, step_size=2, gamma=0.1, verbose=True)
# -
fit(model_fn,
loss_fn,
train_data_loader,
opt,
lr_scheduler,
val_data_loader,
number_epochs=20)
average_accuracy, average_loss = validate(model_fn, loss_fn, test_data_loader)
average_accuracy, average_loss
| notebooks/introduction-to-pytorch-part-2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# We implement Bahdanau et al. (2015)'s attention architecture with an encoder-decoder to translate French and English phrases from the Tatoeba Project.
import pandas as pd
import numpy as np
import scipy as sp
import tensorflow as tf
import tensorflow.keras as keras
import unicodedata
import gc
import collections
import os
import time
import re
import pickle
import matplotlib.pyplot as plt
import matplotlib.patches as mpatches
import seaborn as sns
from copy import deepcopy
from tensorflow.keras.models import Model, Sequential, load_model
from tensorflow.keras.layers import LSTM, GRU, Dense, Dropout, Activation, Bidirectional, Embedding, Input
# <h2> STEP 1: Data Preprocessing
eng, frn = collections.deque(), collections.deque()
with open("./data/fra.txt",encoding='utf-8') as f:
for line in f.readlines():
text = line.strip().split("\t")
eng.append(text[0])
frn.append(text[1])
eng, ind = np.unique(eng, return_index=True)
frn = np.array(frn)[ind]
def preprocess(string):
regexp = re.compile(r'\s+', re.UNICODE)
ns = regexp.sub(' ', string)
ns = re.sub("[^a-zA-Z0-9«».,?!\"\']"," ",ns)
ns = re.sub(r'([«».,?!\"\'])', r' \1 ', ns)
return ns.lower()
eng = np.vectorize(preprocess)(eng)
eng = [elem.split() for elem in eng]
frn = np.vectorize(preprocess)(frn)
frn = [elem.split() for elem in frn]
#Enforce a 14-word restriction on the set
mask = np.array([len(elem)<=14 for elem in eng])
mask = mask & np.array([len(elem)<=14 for elem in frn])
eng = [eng[i] for i in range(len(eng)) if mask[i]]
frn = [frn[i] for i in range(len(frn)) if mask[i]]
#Tokenize
def tokenize(sents):
data = np.zeros((len(sents),16), dtype=np.int64)
word_to_index, index_to_word = {"<begin>":1,"<end>":2}, {1:"<begin>", 2:"<end>"}
curindex = 3
for i in range(len(sents)):
data[i,0] = 1
for j in range(len(sents[i])):
if word_to_index.get(sents[i][j], None) is None:
word_to_index[sents[i][j]] = curindex
index_to_word[curindex] = sents[i][j]
curindex+=1
data[i,j+1] = word_to_index[sents[i][j]]
data[i,len(sents[i])+1] = word_to_index["<end>"]
return data, word_to_index, index_to_word
#Complete Tokenization and Create train-test sets
engdata, engword_to_index, engindex_to_word = tokenize(eng)
frndata, frnword_to_index, frnindex_to_word = tokenize(frn)
train_eng, test_eng = engdata[:90000], engdata[90000:]
train_frn, test_frn = frndata[:90000], frndata[90000:]
train_eng.shape, test_eng.shape, train_frn.shape, test_frn.shape
# <h2> STEP 2: Designing Encoder, Decoder, and Attention Systems
def build_encoder():
tf.keras.backend.clear_session()
inp = Input((16,))
embed = Embedding(len(engword_to_index)+1, 256, embeddings_initializer="uniform")
rep = embed(inp)
encoding, hidden_h, hidden_c = LSTM(512, return_sequences=True, return_state=True)(rep)
return Model(inputs=inp, outputs=[encoding, hidden_h, hidden_c], name="Encoder")
build_encoder().summary()
def build_attention():
tf.keras.backend.clear_session()
#Take in inputs from encoder
enc_output = Input((16,512))
hidden_h = Input((512,))
#expand dims to broadcast to the output shape
hidden = tf.expand_dims(hidden_h, axis=1)
#Define the attention layer's sub-layers
dense1 = Dense(units=512, activation=None)
dense2 = Dense(units=512, activation=None)
mid = Activation(activation="tanh")
final = Dense(units=1, activation=None)
#Calculate score and attention matrix
score = final(mid(dense1(enc_output)+dense2(hidden)))
attmatrix = tf.nn.softmax(score, axis=1)
vector = tf.reduce_sum(attmatrix * enc_output, axis=1)
return Model(inputs=[enc_output, hidden_h], outputs=[vector, attmatrix], name="Bahdanau-Attention")
build_attention().summary()
def build_decoder(attlayer):
tf.keras.backend.clear_session()
#Read in Encoder and previous-prediction Decoder input
enc_output = Input((16,512))
hidden_h, hidden_c = Input((512,)), Input((512,))
prevpred = Input((1,))
#Run Bahdanau Attention
vector, attmatrix = attlayer([enc_output, hidden_h])
#Extract the French Embedding
embed = Embedding(len(frnword_to_index)+1, 256, embeddings_initializer="uniform")
rep = embed(prevpred)
rep = tf.concat([tf.expand_dims(vector, axis=1), rep], axis=2)
#Run a Forward LSTM
recur = LSTM(512, return_sequences=True, return_state=True)
pred, newhidden_h, newhidden_c = recur(rep, initial_state=[hidden_h, hidden_c])
pred = tf.squeeze(pred, [1])
#Predict Next Word
pred = Dense(len(frnword_to_index)+1)(pred)
return Model(inputs=[enc_output, hidden_h, hidden_c, prevpred],
outputs=[pred, newhidden_h, newhidden_c, attmatrix], name="Decoder")
build_decoder(build_attention()).summary()
# <h2> STEP 3: Build Training Infrastructure
optimizer = tf.keras.optimizers.Adam()
def loss(true, pred):
ls = tf.keras.losses.SparseCategoricalCrossentropy(
from_logits=True, reduction=tf.keras.losses.Reduction.NONE)
ls = ls(true, pred)
print(ls)
temptrue = tf.cast(true, tf.float32)
ls = tf.where(tf.math.equal(temptrue, 0.0), 0.0, ls)
return tf.reduce_mean(ls)
#Build all model graphs
encoder = build_encoder()
attnlayer = build_attention()
decoder = build_decoder(attnlayer)
@tf.function
def batch_trainer(engsent, frnsent):
batchloss = 0
with tf.GradientTape() as tape:
encoutput, hh, hc = encoder(engsent)
prevpred = tf.expand_dims(frnsent[:,0], axis=1)
for i in range(1,frnsent.shape[1]):
pred, hh, hc, _ = decoder([encoutput, hh, hc, prevpred])
batchloss+=loss(frnsent[:,i], pred)
prevpred = tf.expand_dims(frnsent[:,i], axis=1)
batchloss /= frnsent.shape[1]
grads = tape.gradient(batchloss, encoder.trainable_variables+decoder.trainable_variables)
optimizer.apply_gradients(zip(grads, encoder.trainable_variables+decoder.trainable_variables))
return batchloss
numepochs=100
batchsize = 128
trainedges = np.arange(0, train_eng.shape[0]+batchsize, batchsize)
trainloss = collections.deque()
for epoch in range(numepochs):
eptrain = 0
for i in range(len(trainedges)-1):
eptrain+=batch_trainer(train_eng[trainedges[i]:trainedges[i+1]],
train_frn[trainedges[i]:trainedges[i+1]])
trainloss.append(eptrain/(len(trainedges)-1))
pickle.dump(trainloss, open("./data/trainloss.pkl","wb"))
encoder.save("./data/encoder.h5")
decoder.save("./data/decoder.h5")
# <h2>STEP 4: Visualize Performance
encoder.load_weights("./data/encoder.h5")
decoder.load_weights("./data/decoder.h5")
def evaluator(engsent, frnsent):
#Set up the sentence prediction matrix
predfrnsent = np.zeros(frnsent.shape, dtype=np.int64)
predfrnsent[:,0] = frnsent[:,0]
#Set up the attention matrix
frn_attn_matrix = np.zeros((frnsent.shape[0], engsent.shape[1], frnsent.shape[1]))
encoutput, hc, hh = encoder.predict(engsent)
prevpred = deepcopy(frnsent[:,0]).reshape(-1,1)
for i in range(1,frnsent.shape[1]):
pred, hh, hc, attmatrix = decoder.predict([encoutput, hh, hc, prevpred])
predfrnsent[:,i] = np.argmax(pred, axis=1)
prevpred = predfrnsent[:,i].reshape(-1,1)
frn_attn_matrix[:,:,i] = attmatrix.reshape(-1,16)
return predfrnsent, frn_attn_matrix
def get_sentences(sent, index_to_word):
ret = collections.deque()
for i in range(sent.shape[0]):
phrase = ""
for j in range(sent.shape[1]):
phrase+=index_to_word[sent[i,j]]+" "
if index_to_word[sent[i,j]]=="<end>":
break
ret.append(phrase)
return ret
pred = np.zeros(test_frn.shape, dtype=np.int64)
batchsize = 256
edges = np.arange(0, pred.shape[0]+batchsize, batchsize)
for i in range(len(edges)-1):
pred[edges[i]:edges[i+1]] = evaluator(test_eng[edges[i]:edges[i+1]],
test_frn[edges[i]:edges[i+1]])[0]
predictedsent = get_sentences(pred, frnindex_to_word)
truesent = get_sentences(test_frn, frnindex_to_word)
| french-english-NMT/Machine Translation with Bahdanau Attention.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/IemProg/DataChallengeXINF554/blob/main/Linear_SVR.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="juLi_0abKDze"
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import torch
import os, sys
from collections import Counter
# + colab={"base_uri": "https://localhost:8080/"} id="NpyyIgY3qjef" outputId="9e54f2b2-4ead-40fe-8b9a-53428ed89b39"
# !pip install verstack
# + id="O-jCbaFBKJfd"
from sklearn.metrics import mean_absolute_error
from verstack.stratified_continuous_split import scsplit
from sklearn.svm import LinearSVR
from sklearn.pipeline import make_pipeline
from sklearn import metrics
from sklearn import preprocessing
from sklearn.preprocessing import MinMaxScaler
from sklearn.model_selection import train_test_split
# + colab={"base_uri": "https://localhost:8080/"} id="RT7sY-ogKLa1" outputId="3a22c524-e827-463f-cf5e-eb11b6437506"
from google.colab import drive
drive.mount("/content/drive")
# + id="YdWfAOZyKhUo"
path = "/content/drive/MyDrive/DataSets/"
path_data_train = path + "train.csv"
# + id="OvhlAfYWKjSd"
train_data = pd.read_csv(path_data_train)
# + id="DXun-dR6Kth0"
#Non-relevant features, can not be used for SVMs models
train_data.drop('timestamp', axis=1, inplace=True)
train_data.drop('user_mentions', axis=1, inplace=True)
train_data.drop('urls', axis=1, inplace=True)
train_data.drop('hashtags', axis=1, inplace=True)
train_data.drop('text', axis=1, inplace=True)
train_data.drop('id', axis=1, inplace=True)
train_data.drop('user_verified', axis=1, inplace=True)
# + colab={"base_uri": "https://localhost:8080/"} id="mDfz6FjGtcW7" outputId="e6cf7416-8f5c-4c82-edec-98363457de3b"
nbr = train_data[train_data.retweet_count != 0].shape[0]
nbr1 = train_data[train_data.retweet_count == 0].shape[0]
print("\t Number of rows where retweets != 0: ", nbr)
print("\t Number of rows where retweets == 0: ", nbr1)
# + colab={"base_uri": "https://localhost:8080/"} id="w7WR7SqstKt1" outputId="20fc8fe4-f4eb-4365-c8b4-d9e0c1a3aa78"
# Shuffle the Dataset.
shuffled_train = train_data.sample(frac=1, random_state=4)
# Put all the samples where they dont have zeros re-tweet in a separate dataset.
non_zero_retweet = shuffled_train.loc[shuffled_train['retweet_count'] != 0]
nbr_samples = non_zero_retweet.shape[0] // 1 #We will take only sixth of it
#Randomly select samples observations from the zero re-tweet (majority class)
zero_retweet = shuffled_train.loc[shuffled_train['retweet_count'] == 0].sample(n=nbr_samples, random_state=42)
print("Shape of non_zero_retweet: ", non_zero_retweet[:nbr_samples].shape)
print("Shape of zero_retweet: ", zero_retweet.shape)
# Concatenate both dataframes again
normalized_train = pd.concat([non_zero_retweet[:nbr_samples], zero_retweet])
print("Normalized train dataset: ", normalized_train.shape)
# + colab={"base_uri": "https://localhost:8080/"} id="BfVq__7Mz76c" outputId="603660ff-a8b7-43b6-84a0-8504485bf974"
nbr = normalized_train[normalized_train.retweet_count != 0].shape[0]
nbr1 = normalized_train[normalized_train.retweet_count == 0].shape[0]
print("\t Number of rows in normalized_train where retweets != 0: ", nbr)
print("\t Number of rows in normalized_train where retweets == 0: ", nbr1)
# + colab={"base_uri": "https://localhost:8080/"} id="IsHud9F3vJ-d" outputId="53a6ee81-33a1-4d84-fe48-afea620244c1"
normalized_train.columns
# + id="b4WIqem7tPPA"
min_max_scaler = preprocessing.MinMaxScaler()
X_train_minmax = min_max_scaler.fit_transform(normalized_train)
# + id="xuxzALx_K4mb"
X_train, X_test, y_train, y_test = train_test_split(normalized_train, normalized_train["retweet_count"],
test_size=0.3, random_state=85)
# We remove the actual number of retweets from our features since it is the value that we are trying to predict
X_train = X_train.drop(['retweet_count'], axis=1)
X_test = X_test.drop(['retweet_count'], axis=1)
# + colab={"base_uri": "https://localhost:8080/"} id="TdhViZaG0Tpo" outputId="490b87d5-daf3-41e6-bd99-bd8ef13cc391"
X_train.columns
X_test.columns
# + colab={"base_uri": "https://localhost:8080/"} id="zWE7aF5mLV7k" outputId="ce59d278-d7a8-4a89-99f9-2a2a334f29da"
print("\t Train dataset shape: ", X_train.shape)
print("\t Test dataset shape: ", X_test.shape)
# + [markdown] id="EqPPt1g-1QQq"
# **Comment:**
# > - Storing the kernel matrix requires memory that scales quadratically with the number of data points. Training time for traditional SVM algorithms also scales superlinearly with the number of data points. So, these algorithms aren't feasible for large data sets.
# LinearSVR is Similar to SVR with parameter kernel=’linear’, but implemented in terms of liblinear rather than libsvm, so it has more flexibility in the choice of penalties and loss functions and should scale better to large numbers of samples.
# + id="cjwYi6QC0LE_"
linear_svr = LinearSVR()
# + id="F2SyrFP1seGj" colab={"base_uri": "https://localhost:8080/"} outputId="c4ad7f6c-ad19-4d5b-bb96-311dea7e9779"
linear_svr.fit(X_train, y_train)
# + colab={"base_uri": "https://localhost:8080/"} id="ehG-lYCQLq-z" outputId="f47bdb44-d0b8-43e7-832c-2d527d2fe6d8"
linear_svr_score = linear_svr.score(X_train, y_train)
print("\t Train accuracy: ", linear_svr_score)
# + colab={"base_uri": "https://localhost:8080/"} id="IRrWUULRLuKt" outputId="25378bc2-37bd-4812-9ad4-214755713527"
predict = linear_svr.predict(X_test)
print("\t Test Linear SVM accuracy: ", predict)
print("\t Prediction error using MAE: ", mean_absolute_error(y_true=y_test, y_pred=predict))
# + id="ITugmcFuscWP"
path_data_eval = path + "/evaluation.csv"
evaluation = pd.read_csv(path_data_eval)
# + id="v7aauxYbsp6g"
#Non-relevant features, can not be used for LR models
evaluation.drop('timestamp', axis=1, inplace=True)
evaluation.drop('user_mentions', axis=1, inplace=True)
evaluation.drop('urls', axis=1, inplace=True)
evaluation.drop('hashtags', axis=1, inplace=True)
evaluation.drop('text', axis=1, inplace=True)
evaluation.drop('user_verified', axis=1, inplace=True)
# + id="72XkcmXjss7i"
withoutID = evaluation.copy()
withoutID.drop('id', axis=1, inplace=True)
# + id="sTxVbuU_vOeV"
min_max_scaler = preprocessing.MinMaxScaler()
withoutID_minmax = min_max_scaler.fit_transform(withoutID)
# + id="uq1f7_Btsu5-"
kaggleOut = linear_svr.predict(withoutID_minmax)
# + colab={"base_uri": "https://localhost:8080/"} id="dnOlvF2qre3z" outputId="7aaeb7e4-4d39-45c2-82ad-45d82ec5aa4f"
import csv
f = open("linearsvr.csv", "w+")
with open("linearsvr.csv", 'w') as f:
writer = csv.writer(f)
writer.writerow(["TweetID", "NoRetweets"])
for index, prediction in enumerate(kaggleOut):
print(str(evaluation['id'].iloc[index]) + " ," + str(int(prediction)))
writer.writerow([str(evaluation['id'].iloc[index]) , str(int(prediction))])
# + id="QNiwHAMfqfFh"
| Linear_SVR.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
import pandas as pd
import numpy as np
import datetime
import seaborn as sns
import matplotlib.pyplot as plt
import matplotlib.dates as mdates
# %matplotlib inline
from sklearn.decomposition import PCA
from sklearn.preprocessing import StandardScaler
from sklearn.feature_selection import SelectKBest, mutual_info_regression
from sklearn.model_selection import train_test_split
# -
from sklearn.pipeline import make_pipeline
from sklearn.model_selection import KFold, cross_val_predict
from sklearn.neighbors import KNeighborsRegressor
from sklearn.metrics import mean_squared_error, r2_score
from sklearn.linear_model import LinearRegression
from math import sqrt
path = './dbs/handled/'
df_brazil_final = pd.read_csv(path + 'df_brazilian_states_cases.csv')
df_brazil_rolled = pd.read_csv(path + 'df_brazil_rolled_cases.csv')
df_seird = pd.read_csv(path + 'df_brazil_appllied_seird_model.csv')
df_parameters_seird = pd.read_csv(path + 'df_parameters_seird.csv', parse_dates=['Date'], index_col='Date')
df_parameters_seird
# +
holiday = ['2020-02-26',
'2020-04-10',
'2020-04-21',
'2020-05-01',
'2020-06-11',
'2020-10-12',
'2020-11-02',
'2020-11-15',
'2020-12-25',
'2021-01-01',
'2021-02-16',
'2021-04-02',
'2021-04-21',
'2021-05-01']
canceled_holiday = ['2020-09-07']
elections = ['2020-11-15',
'2020-11-29']
political_interventions = ['2020-04-16', #Mandetta left
'2020-05-15', #Teich left
'2021-03-23'] #Pazuello left
restriction_flexibility = ['2020-08-01'] #Flexibilization in São Paulo
lockdown_ini = ['2020-05-05']#,'2020-11-01']
lockdown_end = ['2020-06-14']#,'2021-01-30']
# -
moving_average=14
# ## Mobility Data
# +
df_mobility = pd.read_csv('./dbs/brasil_df.csv', parse_dates=['date'], index_col='date')
first_day = df_parameters_seird.index.min()
last_day = df_parameters_seird.index.max()
df_mobility = df_mobility.loc[(df_mobility.index >= first_day) & (df_mobility.index <= last_day)]
# -
# replacing accumulated cases by daily cases
df_mobility['cases'] = df_parameters_seird['cases']
df_mobility['deaths'] = df_parameters_seird['deaths']
df_mobility['holiday'] = df_parameters_seird['holiday']
df_mobility
# +
fig, (ax1, ax2) = plt.subplots(2, figsize=(17, 10))
ax1.plot(df_mobility['retail and recreation'].rolling(moving_average, min_periods=1).mean(), label='retail and recreation')
ax1.plot(df_mobility['transit stations'].rolling(moving_average, min_periods=1).mean(), label='transit stations')
ax1.plot(df_mobility['parks'].rolling(moving_average, min_periods=1).mean(), label='parks')
ax1.plot(df_mobility['workplaces'].rolling(moving_average, min_periods=1).mean(), label='workplaces')
ax1.plot(df_mobility['grocery and pharmacy'].rolling(moving_average, min_periods=1).mean(), label='grocery and pharmacy')
ax1.plot(df_mobility['residential'].rolling(moving_average, min_periods=1).mean(), label='residential')
ax1.set_ylabel("Mobility Indice")
ax1.grid(axis='x')
ax1.xaxis.set_major_locator(mdates.DayLocator(interval=14))
ax1.set_xlabel("Biweekly period")
ax1.xaxis.set_label_position('top')
ax1.legend(loc=4)
for i in range(len(holiday)):
ax1.axvspan(datetime.datetime.strptime(holiday[i], '%Y-%m-%d'),
datetime.datetime.strptime(holiday[i], '%Y-%m-%d') + datetime.timedelta(days=14),
label="Brazilian Elections",
color="crimson",
alpha=0.075)
for i in range(len(elections)):
ax1.axvspan(datetime.datetime.strptime(elections[i], '%Y-%m-%d'),
datetime.datetime.strptime(elections[i], '%Y-%m-%d') + datetime.timedelta(days=14),
label="Brazilian Elections",
color="blue",
alpha=0.075)
ax1.xaxis.set_major_locator(mdates.DayLocator(interval=14))
plt.gca().xaxis.set_major_formatter(mdates.DateFormatter('%W\n%y'))
ax1.set_xticklabels([])
ax2.plot(df_parameters_seird['cases'].rolling(moving_average, min_periods=1).mean(),
color=plt.cm.viridis(0))
# ax2.plot(df_parameters_seird['Date'],
# df_parameters_seird['deaths'].rolling(moving_average, min_periods=1).mean(),
# color=plt.cm.viridis(1),
# label='COVID-19 Daily Deaths')
ax2.grid(axis='x')
ax2.set_xlabel("Biweekly period")
ax2.set_ylabel("COVID-19 Daily Cases")
#ax2.set_xticklabels([])
ax2.xaxis.set_ticks_position('top')
ax2.xaxis.set_label_position('bottom')
ax2.xaxis.set_major_locator(mdates.DayLocator(interval=14))
plt.gca().xaxis.set_major_formatter(mdates.DateFormatter('%W\n%y'))
for i in range(len(holiday)):
ax2.axvspan(datetime.datetime.strptime(holiday[i], '%Y-%m-%d'),
datetime.datetime.strptime(holiday[i], '%Y-%m-%d') + datetime.timedelta(days=14),
label="Holiday + 14 days",
color="crimson",
alpha=0.2)
for i in range(len(elections)):
ax2.axvspan(datetime.datetime.strptime(elections[i], '%Y-%m-%d'),
datetime.datetime.strptime(elections[i], '%Y-%m-%d') + datetime.timedelta(days=14),
label="Brazilian Elections",
color="blue",
alpha=0.2)
for i in range(len(canceled_holiday)):
ax2.axvspan(datetime.datetime.strptime(canceled_holiday[i], '%Y-%m-%d'),
datetime.datetime.strptime(canceled_holiday[i], '%Y-%m-%d') + datetime.timedelta(days=14),
label="Holiday + 14 days",
color="crimson",
alpha=0.05)
def remove_duplicate_labels_from_legends(ax):
handles, labels = ax.get_legend_handles_labels()
unique = [(h, l) for i, (h, l) in enumerate(zip(handles, labels)) if l not in labels[:i]]
ax.legend(*zip(*unique))
remove_duplicate_labels_from_legends(ax2)
plt.subplots_adjust(hspace=0.175)
plt.savefig('./images/mobility.pdf', transparent=True)
plt.show()
# -
# ---------------
# # PCA
# +
features = ['retail and recreation',
'grocery and pharmacy',
'parks',
'transit stations',
'workplaces',
'residential']
# Separating out the features
X = df_mobility.loc[:,features].values
X = StandardScaler().fit_transform(X)
# -
pca.explained_variance_ratio_
# +
pca = PCA()
X_reduced = pca.fit_transform(X)
print(pca.explained_variance_ratio_)
print(pca.explained_variance_ratio_.cumsum())
fig = plt.figure(figsize=(6,2))
plt.plot(pca.explained_variance_ratio_.cumsum(), 'ro-')
plt.grid(axis='y')
plt.xticks([0,1, 2,3,4,5],[1, 2,3,4,5,6])
plt.xlabel('PCA Assumption')
plt.ylabel('Data variability')
plt.savefig('./images/mobility_pca_qtty.pdf', transparent=True, bbox_inches='tight', pad_inches=0)
plt.show()
# -
# ### Com 2 PCAs é possível generalizar os dados de mobilidade em 90%
pca_2comp = PCA(n_components=2)
principal_components = pca_2comp.fit_transform(X)
df_principal_components = pd.DataFrame(data = principal_components, columns=['pc1', 'pc2'])
df_principal_components.index = df_mobility.index
# +
fig, ax = plt.subplots(figsize=(17, 5))
ax.plot(df_parameters_seird['R0_ma'], linestyle='-',label="Transmission Rate R0")
ax.axhline(y=1, color='r', linestyle='-')
ax.grid(axis='x')
ax.set_ylabel("Transmission Rate")
ax.set_xlabel("Biweekly period")
ax.legend(loc=2)
ci = 0.95 * np.std(df_parameters_seird['R0_ma'])/np.mean(df_parameters_seird['R0_ma'])
ax.fill_between(df_parameters_seird.index,
(df_parameters_seird['R0_ma']-ci),
(df_parameters_seird['R0_ma']+ci),
color='b',
alpha=.05)
ax1 = ax.twinx()
ax1.plot(df_principal_components['pc1'].rolling(moving_average, min_periods=1).mean(), color='m', linestyle='--', label='PC1')
ax1.plot(df_principal_components['pc2'].rolling(moving_average, min_periods=1).mean(), color='g', linestyle='--', label='PC2')
ax1.set_ylabel("PCA - Mobility Indice")
ax1.legend(loc=4)
for i in range(len(holiday)):
ax.axvspan(datetime.datetime.strptime(holiday[i], '%Y-%m-%d'),
datetime.datetime.strptime(holiday[i], '%Y-%m-%d') + datetime.timedelta(days=14),
label="Brazilian Elections",
color="crimson",
alpha=0.075)
for i in range(len(elections)):
ax.axvspan(datetime.datetime.strptime(elections[i], '%Y-%m-%d'),
datetime.datetime.strptime(elections[i], '%Y-%m-%d') + datetime.timedelta(days=14),
label="Brazilian Elections",
color="blue",
alpha=0.075)
ax.xaxis.set_major_locator(mdates.DayLocator(interval=14))
plt.gca().xaxis.set_major_formatter(mdates.DateFormatter('%W\n%y'))
plt.savefig('./images/transmission_rate_and_mobility_PCAs.pdf', transparent=True)
plt.show()
# -
# #### PCA with flag HOLIDAY - 1 and 0
df_principal_components['holiday'] = df_mobility['holiday']
df_principal_components['deaths'] = df_mobility.deaths
df_principal_components['r0'] = df_parameters_seird['R0_ma']
# +
fig = plt.figure(figsize=(5,5))
ax = fig.add_subplot(1,1,1)
ax.set_xlabel('PC1')
ax.set_ylabel('PC2')
for n, grp in df_principal_components.groupby('holiday'):
ax.scatter(x = "pc1",
y = "pc2",
data=grp,
label=n)
ax.set_title('PCA - Holiday period flag')
ax.legend(title="Holiday flag")
plt.show()
# -
# #### PCA with Ro > 1
df_principal_components_r0 = df_principal_components
df_principal_components_r0['R0>1'] = np.where(df_parameters_seird['R0_ma'] > 1, 1, 0)
# +
fig = plt.figure(figsize=(5,5))
ax = fig.add_subplot(1,1,1)
ax.set_xlabel('PC1')
ax.set_ylabel('PC2')
for n, grp in df_principal_components_r0.groupby('R0>1'):
ax.scatter(x = "pc1",
y = "pc2",
data=grp,
label=n)
ax.set_title('PCA - Holiday period with R0 > 1')
ax.legend(title="R0 > 1")
plt.savefig('./images/pca_clustering_holiday_with_r0.pdf', transparent=True)
plt.show()
# -
# #### PCA with Ro > 1 - only 2020
df_principal_components_r0_2020 = df_principal_components[df_principal_components.index <= '2020-12-31']
# +
fig = plt.figure(figsize=(5,5))
ax = fig.add_subplot(1,1,1)
ax.set_xlabel('PC1')
ax.set_ylabel('PC2')
for n, grp in df_principal_components_r0_2020.groupby('R0>1'):
ax.scatter(x = "pc1",
y = "pc2",
data=grp,
label=n)
ax.set_title('PCA - Holiday period flag with R0 > 1 in 2020')
ax.legend(title="R0 > 1")
plt.savefig('./images/pca_clustering_holiday_with_r0_2020.pdf', transparent=True)
plt.show()
# -
# #### PCA with Ro > 1 - only 2021
df_principal_components_r0_2021 = df_principal_components[df_principal_components.index > '2020-12-31']
# +
fig = plt.figure(figsize=(5,5))
ax = fig.add_subplot(1,1,1)
ax.set_xlabel('PC1')
ax.set_ylabel('PC2')
for n, grp in df_principal_components_r0_2021.groupby('R0>1'):
ax.scatter(x = "pc1",
y = "pc2",
data=grp,
label=n)
ax.set_title('PCA - Holiday period flag with R0 > 1 in 2021')
ax.legend(title="R0 > 1")
plt.savefig('./images/pca_clustering_holiday_with_r0_2021.pdf', transparent=True)
plt.show()
| notebooks/Holiday - passo 2 - Data Analysis SEIRD and Mobility.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Our Mission ##
#
# Spam detection is one of the major applications of Machine Learning in the interwebs today. Pretty much all of the major email service providers have spam detection systems built in and automatically classify such mail as 'Junk Mail'.
#
# In this mission we will be using the Naive Bayes algorithm to create a model that can classify SMS messages as spam or not spam, based on the training we give to the model. It is important to have some level of intuition as to what a spammy text message might look like. Often they have words like 'free', 'win', 'winner', 'cash', 'prize' and the like in them as these texts are designed to catch your eye and in some sense tempt you to open them. Also, spam messages tend to have words written in all capitals and also tend to use a lot of exclamation marks. To the human recipient, it is usually pretty straightforward to identify a spam text and our objective here is to train a model to do that for us!
#
# Being able to identify spam messages is a binary classification problem as messages are classified as either 'Spam' or 'Not Spam' and nothing else. Also, this is a supervised learning problem, as we will be feeding a labelled dataset into the model, that it can learn from, to make future predictions.
#
# ### Step 1.1: Understanding our dataset ###
#
#
# We will be using a dataset originally compiled and posted on the UCI Machine Learning repository which has a very good collection of datasets for experimental research purposes. If you're interested, you can review the [abstract](https://archive.ics.uci.edu/ml/datasets/SMS+Spam+Collection) and the original [compressed data file](https://archive.ics.uci.edu/ml/machine-learning-databases/00228/) on the UCI site. For this exercise, however, we've gone ahead and downloaded the data for you.
#
#
# **Here's a preview of the data:**
#
# <img src="images/dqnb.png" height="1242" width="1242">
#
# The columns in the data set are currently not named and as you can see, there are 2 columns.
#
# The first column takes two values, 'ham' which signifies that the message is not spam, and 'spam' which signifies that the message is spam.
#
# The second column is the text content of the SMS message that is being classified.
# >**Instructions:**
# * Import the dataset into a pandas dataframe using the **read_table** method. The file has already been downloaded, and you can access it using the filepath 'smsspamcollection/SMSSpamCollection'. Because this is a tab separated dataset we will be using '\\t' as the value for the 'sep' argument which specifies this format.
# * Also, rename the column names by specifying a list ['label', 'sms_message'] to the 'names' argument of read_table().
# * Print the first five values of the dataframe with the new column names.
# '!' allows you to run bash commands from jupyter notebook.
print("List all the files in the current directory\n")
# !ls
# The required data table can be found under smsspamcollection/SMSSpamCollection
print("\n List all the files inside the smsspamcollection directory\n")
# !ls smsspamcollection
# +
import pandas as pd
# Dataset available using filepath 'smsspamcollection/SMSSpamCollection'
df = pd.read_table('smsspamcollection/SMSSpamCollection', header=1, names=["label", "message"])#TODO
# Output printing out first 5 rows
df.head()
# -
# ### Step 1.2: Data Preprocessing ###
#
# Now that we have a basic understanding of what our dataset looks like, let's convert our labels to binary variables, 0 to represent 'ham'(i.e. not spam) and 1 to represent 'spam' for ease of computation.
#
# You might be wondering why do we need to do this step? The answer to this lies in how scikit-learn handles inputs. Scikit-learn only deals with numerical values and hence if we were to leave our label values as strings, scikit-learn would do the conversion internally(more specifically, the string labels will be cast to unknown float values).
#
# Our model would still be able to make predictions if we left our labels as strings but we could have issues later when calculating performance metrics, for example when calculating our precision and recall scores. Hence, to avoid unexpected 'gotchas' later, it is good practice to have our categorical values be fed into our model as integers.
# >**Instructions:**
# * Convert the values in the 'label' column to numerical values using map method as follows:
# {'ham':0, 'spam':1} This maps the 'ham' value to 0 and the 'spam' value to 1.
# * Also, to get an idea of the size of the dataset we are dealing with, print out number of rows and columns using
# 'shape'.
'''
Solution
'''
df['label'] = df.label.map({'spam':1,'ham':0})
# ### Bag of Words in scikit-learn ###
#
'''
Solution
'''
from sklearn.feature_extraction.text import CountVectorizer
count_vector =CountVectorizer()
# **Data preprocessing with CountVectorizer()**
#
# In Step 2.2, we implemented a version of the CountVectorizer() method from scratch that entailed cleaning our data first. This cleaning involved converting all of our data to lower case and removing all punctuation marks. CountVectorizer() has certain parameters which take care of these steps for us. They are:
#
# * `lowercase = True`
#
# The `lowercase` parameter has a default value of `True` which converts all of our text to its lower case form.
#
#
# * `token_pattern = (?u)\\b\\<KEY>`
#
# The `token_pattern` parameter has a default regular expression value of `(?u)\\b\\w\\w+\\b` which ignores all punctuation marks and treats them as delimiters, while accepting alphanumeric strings of length greater than or equal to 2, as individual tokens or words.
#
#
# * `stop_words`
#
# The `stop_words` parameter, if set to `english` will remove all words from our document set that match a list of English stop words defined in scikit-learn. Considering the small size of our dataset and the fact that we are dealing with SMS messages and not larger text sources like e-mail, we will not use stop words, and we won't be setting this parameter value.
#
# You can take a look at all the parameter values of your `count_vector` object by simply printing out the object as follows:
'''
Practice node:
Print the 'count_vector' object which is an instance of 'CountVectorizer()'
'''
# No need to revise this code
print(count_vector)
# The `get_feature_names()` method returns our feature names for this dataset, which is the set of words that make up our vocabulary for 'documents'.
# ### Step 3.1: Training and testing sets ###
#
# Now that we understand how to use the Bag of Words approach, we can return to our original, larger UCI dataset and proceed with our analysis. Our first step is to split our dataset into a training set and a testing set so we can first train, and then test our model.
# Now we have a clean representation of the documents in terms of the frequency distribution of the words in them. To make it easier to understand our next step is to convert this array into a dataframe and name the columns appropriately.
#
# >>**Instructions:**
# Split the dataset into a training and testing set using the train_test_split method in sklearn, and print out the number of rows we have in each of our training and testing data. Split the data
# using the following variables:
# * `X_train` is our training data for the 'sms_message' column.
# * `y_train` is our training data for the 'label' column
# * `X_test` is our testing data for the 'sms_message' column.
# * `y_test` is our testing data for the 'label' column.
#
# +
'''
Solution
'''
# split into training and testing sets
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(df['message'],
df['label'],
random_state=1)
print('Number of rows in the total set: {}'.format(df.shape[0]))
print('Number of rows in the training set: {}'.format(X_train.shape[0]))
print('Number of rows in the test set: {}'.format(X_test.shape[0]))
# -
# ### Step 3.2: Applying Bag of Words processing to our dataset. ###
#
# Now that we have split the data, our next objective is to follow the steps from "Step 2: Bag of Words," and convert our data into the desired matrix format. To do this we will be using CountVectorizer() as we did before. There are two steps to consider here:
#
# * First, we have to fit our training data (`X_train`) into `CountVectorizer()` and return the matrix.
# * Secondly, we have to transform our testing data (`X_test`) to return the matrix.
#
# Note that `X_train` is our training data for the 'sms_message' column in our dataset and we will be using this to train our model.
#
# `X_test` is our testing data for the 'sms_message' column and this is the data we will be using (after transformation to a matrix) to make predictions on. We will then compare those predictions with `y_test` in a later step.
#
# For now, we have provided the code that does the matrix transformations for you!
# +
'''
Solution
'''
# Fit the training data and then return the matrix
training_data = count_vector.fit_transform(X_train)
# Transform testing data and return the matrix. Note we are not fitting the testing data into the CountVectorizer()
testing_data = count_vector.transform(X_test)
# -
# ### Naive Bayes implementation using scikit-learn ###
#
# Now let's return to our spam classification context. Thankfully, sklearn has several Naive Bayes implementations that we can use, so we do not have to do the math from scratch. We will be using sklearn's `sklearn.naive_bayes` method to make predictions on our SMS messages dataset.
#
# Specifically, we will be using the multinomial Naive Bayes algorithm. This particular classifier is suitable for classification with discrete features (such as in our case, word counts for text classification). It takes in integer word counts as its input. On the other hand, Gaussian Naive Bayes is better suited for continuous data as it assumes that the input data has a Gaussian (normal) distribution.
'''
Solution
'''
from sklearn.naive_bayes import MultinomialNB
naive_bayes = MultinomialNB()
naive_bayes.fit(training_data, y_train)
'''
Solution
'''
predictions = naive_bayes.predict(testing_data)
# Now that predictions have been made on our test set, we need to check the accuracy of our predictions.
# ### Step 6: Evaluating our model ###
#
# Now that we have made predictions on our test set, our next goal is to evaluate how well our model is doing. There are various mechanisms for doing so, so first let's review them.
#
# **Accuracy** measures how often the classifier makes the correct prediction. It’s the ratio of the number of correct predictions to the total number of predictions (the number of test data points).
#
# **Precision** tells us what proportion of messages we classified as spam, actually were spam.
# It is a ratio of true positives (words classified as spam, and which actually are spam) to all positives (all words classified as spam, regardless of whether that was the correct classification). In other words, precision is the ratio of
#
# `[True Positives/(True Positives + False Positives)]`
#
# **Recall (sensitivity)** tells us what proportion of messages that actually were spam were classified by us as spam.
# It is a ratio of true positives (words classified as spam, and which actually are spam) to all the words that were actually spam. In other words, recall is the ratio of
#
# `[True Positives/(True Positives + False Negatives)]`
#
# For classification problems that are skewed in their classification distributions like in our case - for example if we had 100 text messages and only 2 were spam and the other 98 weren't - accuracy by itself is not a very good metric. We could classify 90 messages as not spam (including the 2 that were spam but we classify them as not spam, hence they would be false negatives) and 10 as spam (all 10 false positives) and still get a reasonably good accuracy score. For such cases, precision and recall come in very handy. These two metrics can be combined to get the **F1 score**, which is the weighted average of the precision and recall scores. This score can range from 0 to 1, with 1 being the best possible F1 score.
# We will be using all 4 of these metrics to make sure our model does well. For all 4 metrics whose values can range from 0 to 1, having a score as close to 1 as possible is a good indicator of how well our model is doing.
'''
Solution
'''
from sklearn.metrics import accuracy_score, precision_score, recall_score, f1_score
print('Accuracy score: ', format(accuracy_score(y_test,predictions)))
print('Precision score: ', format(precision_score(y_test,predictions)))
print('Recall score: ', format(recall_score(y_test,predictions)))
print('F1 score: ', format(f1_score(y_test,predictions)))
# ### Step 7: Conclusion ###
#
# One of the major advantages that Naive Bayes has over other classification algorithms is its ability to handle an extremely large number of features. In our case, each word is treated as a feature and there are thousands of different words. Also, it performs well even with the presence of irrelevant features and is relatively unaffected by them. The other major advantage it has is its relative simplicity. Naive Bayes' works well right out of the box and tuning its parameters is rarely ever necessary, except usually in cases where the distribution of the data is known.
# It rarely ever overfits the data. Another important advantage is that its model training and prediction times are very fast for the amount of data it can handle. All in all, Naive Bayes' really is a gem of an algorithm!
#
# ### Turns Out...
#
# We can see from the scores above that our Naive Bayes model actually does a pretty good job of classifying spam and "ham." However, let's take a look at a few additional models to see if we can't improve anyway.
#
# Specifically in this notebook, we will take a look at the following techniques:
#
# * [BaggingClassifier](http://scikit-learn.org/stable/modules/generated/sklearn.ensemble.BaggingClassifier.html#sklearn.ensemble.BaggingClassifier)
# * [RandomForestClassifier](http://scikit-learn.org/stable/modules/generated/sklearn.ensemble.RandomForestClassifier.html#sklearn.ensemble.RandomForestClassifier)
# * [AdaBoostClassifier](http://scikit-learn.org/stable/modules/generated/sklearn.ensemble.AdaBoostClassifier.html#sklearn.ensemble.AdaBoostClassifier)
#
# Another really useful guide for ensemble methods can be found [in the documentation here](http://scikit-learn.org/stable/modules/ensemble.html).
#
# These ensemble methods use a combination of techniques you have seen throughout this lesson:
#
# * **Bootstrap the data** passed through a learner (bagging).
# * **Subset the features** used for a learner (combined with bagging signifies the two random components of random forests).
# * **Ensemble learners** together in a way that allows those that perform best in certain areas to create the largest impact (boosting).
#
#
# In this notebook, let's get some practice with these methods, which will also help you get comfortable with the process used for performing supervised machine learning in Python in general.
#
# Since you cleaned and vectorized the text in the previous notebook, this notebook can be focused on the fun part - the machine learning part.
#
# ### This Process Looks Familiar...
#
# In general, there is a five step process that can be used each time you want to use a supervised learning method (which you actually used above):
#
# 1. **Import** the model.
# 2. **Instantiate** the model with the hyperparameters of interest.
# 3. **Fit** the model to the training data.
# 4. **Predict** on the test data.
# 5. **Score** the model by comparing the predictions to the actual values.
#
# Follow the steps through this notebook to perform these steps using each of the ensemble methods: **BaggingClassifier**, **RandomForestClassifier**, and **AdaBoostClassifier**.
#
# > **Step 1**: First use the documentation to `import` all three of the models.
# Import the Bagging, RandomForest, and AdaBoost Classifier
from sklearn.ensemble import BaggingClassifier, RandomForestClassifier, AdaBoostClassifier
# > **Step 2:** Now that you have imported each of the classifiers, `instantiate` each with the hyperparameters specified in each comment. In the upcoming lessons, you will see how we can automate the process to finding the best hyperparameters. For now, let's get comfortable with the process and our new algorithms.
# +
# Instantiate a BaggingClassifier with:
# 200 weak learners (n_estimators) and everything else as default values
bag_mod = BaggingClassifier(n_estimators=200)
# Instantiate a RandomForestClassifier with:
# 200 weak learners (n_estimators) and everything else as default values
rf_mod = RandomForestClassifier(n_estimators=200)
# Instantiate an a AdaBoostClassifier with:
# With 300 weak learners (n_estimators) and a learning_rate of 0.2
ada_mod = AdaBoostClassifier(n_estimators=300, learning_rate=0.2)
# -
# > **Step 3:** Now that you have instantiated each of your models, `fit` them using the **training_data** and **y_train**. This may take a bit of time, you are fitting 700 weak learners after all!
# +
# Fit your BaggingClassifier to the training data
bag_mod.fit(training_data, y_train)
# Fit your RandomForestClassifier to the training data
rf_mod.fit(training_data, y_train)
# Fit your AdaBoostClassifier to the training data
ada_mod.fit(training_data, y_train)
# -
# > **Step 4:** Now that you have fit each of your models, you will use each to `predict` on the **testing_data**.
# +
# Predict using BaggingClassifier on the test data
bag_preds = bag_mod.predict(testing_data)
# Predict using RandomForestClassifier on the test data
rf_preds = rf_mod.predict(testing_data)
# Predict using AdaBoostClassifier on the test data
ada_preds = ada_mod.predict(testing_data)
# -
# > **Step 5:** Now that you have made your predictions, compare your predictions to the actual values using the function below for each of your models - this will give you the `score` for how well each of your models is performing. It might also be useful to show the Naive Bayes model again here, so we can compare them all side by side.
def print_metrics(y_true, preds, model_name=None):
'''
INPUT:
y_true - the y values that are actually true in the dataset (numpy array or pandas series)
preds - the predictions for those values from some model (numpy array or pandas series)
model_name - (str - optional) a name associated with the model if you would like to add it to the print statements
OUTPUT:
None - prints the accuracy, precision, recall, and F1 score
'''
if model_name == None:
print('Accuracy score: ', format(accuracy_score(y_true, preds)))
print('Precision score: ', format(precision_score(y_true, preds)))
print('Recall score: ', format(recall_score(y_true, preds)))
print('F1 score: ', format(f1_score(y_true, preds)))
print('\n\n')
else:
print('Accuracy score for ' + model_name + ' :' , format(accuracy_score(y_true, preds)))
print('Precision score ' + model_name + ' :', format(precision_score(y_true, preds)))
print('Recall score ' + model_name + ' :', format(recall_score(y_true, preds)))
print('F1 score ' + model_name + ' :', format(f1_score(y_true, preds)))
print('\n\n')
# +
# Print Bagging scores
print_metrics(y_test, bag_preds, 'bagging')
# Print Random Forest scores
print_metrics(y_test, rf_preds, 'random forest')
# Print AdaBoost scores
print_metrics(y_test, ada_preds, 'adaboost')
# Naive Bayes Classifier scores
print_metrics(y_test, predictions, 'naive bayes')
# -
def build_roc_auc(model, X_train, X_test, y_train, y_test):
'''
INPUT:
model - an sklearn instantiated model
X_train - the training data
y_train - the training response values (must be categorical)
X_test - the test data
y_test - the test response values (must be categorical)
OUTPUT:
auc - returns auc as a float
prints the roc curve
'''
import numpy as np
import matplotlib.pyplot as plt
from itertools import cycle
from sklearn.metrics import roc_curve, auc, roc_auc_score
from scipy import interp
y_preds = model.fit(X_train, y_train).predict_proba(X_test)
# Compute ROC curve and ROC area for each class
fpr = dict()
tpr = dict()
roc_auc = dict()
for i in range(len(y_test)):
fpr[i], tpr[i], _ = roc_curve(y_test, y_preds[:, 1])
roc_auc[i] = auc(fpr[i], tpr[i])
# Compute micro-average ROC curve and ROC area
fpr["micro"], tpr["micro"], _ = roc_curve(y_test.ravel(), y_preds[:, 1].ravel())
roc_auc["micro"] = auc(fpr["micro"], tpr["micro"])
plt.plot(fpr[2], tpr[2], color='darkorange',
lw=2, label='ROC curve (area = %0.2f)' % roc_auc[2])
plt.plot([0, 1], [0, 1], color='navy', lw=2, linestyle='--')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Receiver operating characteristic curve')
plt.show()
return roc_auc_score(y_test, np.round(y_preds[:, 1]))
# +
build_roc_auc(naive_bayes, training_data, testing_data, y_train, y_test)
# -
| Spam Classifier/Spam Classifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Linear regression
# +
""" Starter code for simple linear regression example using placeholders
Created by <NAME> (<EMAIL>)
CS20: "TensorFlow for Deep Learning Research"
cs20.stanford.edu
Lecture 03
"""
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import os
os.environ['TF_CPP_MIN_LOG_LEVEL']='2'
import time
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
sess_config = tf.ConfigProto(gpu_options=tf.GPUOptions(allow_growth=True))
# -
# ## Data reading
def read_birth_life_data(filename):
"""
Read in birth_life_2010.txt and return:
data in the form of NumPy array
n_samples: number of samples
"""
text = open(filename, 'r').readlines()[1:]
data = [line[:-1].split('\t') for line in text]
births = [float(line[1]) for line in data]
lifes = [float(line[2]) for line in data]
data = list(zip(births, lifes))
n_samples = len(data)
data = np.asarray(data, dtype=np.float32)
return data, n_samples
# +
DATA_FILE = '../datasets/birth_life_2010.txt'
# Step 1: read in data from the .txt file
data, n_samples = utils.read_birth_life_data(DATA_FILE)
# -
# ## Phase 1: Build a graph
# Step 2: create placeholders for X (birth rate) and Y (life expectancy)
# Remember both X and Y are scalars with type float
X, Y = None, None
# Step 3: create weight and bias, initialized to 0.0
# Make sure to use tf.get_variable
w, b = None, None
# Step 4: build model to predict Y
# e.g. how would you derive at Y_predicted given X, w, and b
Y_predicted = None
# Step 5: use the square error as the loss function
loss = None
# Step 6: using gradient descent with learning rate of 0.001 to minimize loss
optimizer = tf.train.GradientDescentOptimizer(learning_rate=0.001).minimize(loss)
# ## Phase 2: Train a model using `tf.Session`
# +
start = time.time()
# Create a filewriter to write the model's graph to TensorBoard
#############################
########## TO DO ############
#############################
# Step 7: initialize the necessary variables, in this case, w and b
#############################
########## TO DO ############
#############################
# Step 8: train the model for 100 epochs
for i in range(100):
total_loss = 0
for x, y in data:
# Execute train_op and get the value of loss.
# Don't forget to feed in data for placeholders
_, loss = ########## TO DO ############
total_loss += loss
print('Epoch {0}: {1}'.format(i, total_loss/n_samples))
# close the writer when you're done using it
#############################
########## TO DO ############
#############################
writer.close()
# Step 9: output the values of w and b
w_out, b_out = None, None
#############################
########## TO DO ############
#############################
print('Took: %f seconds' %(time.time() - start))
# -
# ## Plot the result
plt.plot(data[:,0], data[:,1], 'bo', label='Real data')
plt.plot(data[:,0], data[:,0] * w_out + b_out, 'r', label='Predicted data')
plt.legend()
plt.show()
| 03.regression/01-2.linear.regression.eager.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
# # Supervised Learning Part 2 -- Regression Analysis
# In regression we are trying to predict a continuous output variable -- in contrast to the nominal variables we were predicting in the previous classification examples.
#
# Let's start with a simple toy example with one feature dimension (explanatory variable) and one target variable. We will create a dataset out of a sine curve with some noise:
x = np.linspace(-3, 3, 100)
print(x)
rng = np.random.RandomState(42)
y = np.sin(4 * x) + x + rng.uniform(size=len(x))
plt.plot(x, y, 'o');
# Linear Regression
# =================
#
# The first model that we will introduce is the so-called simple linear regression. Here, we want to fit a line to the data, which
#
# One of the simplest models again is a linear one, that simply tries to predict the data as lying on a line. One way to find such a line is `LinearRegression` (also known as [*Ordinary Least Squares (OLS)*](https://en.wikipedia.org/wiki/Ordinary_least_squares) regression).
# The interface for LinearRegression is exactly the same as for the classifiers before, only that ``y`` now contains float values, instead of classes.
# As we remember, the scikit-learn API requires us to provide the target variable (`y`) as a 1-dimensional array; scikit-learn's API expects the samples (`X`) in form a 2-dimensional array -- even though it may only consist of 1 feature. Thus, let us convert the 1-dimensional `x` NumPy array into an `X` array with 2 axes:
#
print('Before: ', x.shape)
X = x[:, np.newaxis]
print('After: ', X.shape)
# Again, we start by splitting our dataset into a training (75%) and a test set (25%):
# +
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.25, random_state=42)
# -
# Next, we use the learning algorithm implemented in `LinearRegression` to **fit a regression model to the training data**:
# +
from sklearn.linear_model import LinearRegression
regressor = LinearRegression()
regressor.fit(X_train, y_train)
# -
# After fitting to the training data, we paramerterized a linear regression model with the following values.
print('Weight coefficients: ', regressor.coef_)
print('y-axis intercept: ', regressor.intercept_)
# Since our regression model is a linear one, the relationship between the target variable (y) and the feature variable (x) is defined as
#
# $$y = weight \times x + \text{intercept}$$.
#
# Plugging in the min and max values into thos equation, we can plot the regression fit to our training data:
# +
min_pt = X.min() * regressor.coef_[0] + regressor.intercept_
max_pt = X.max() * regressor.coef_[0] + regressor.intercept_
plt.plot([X.min(), X.max()], [min_pt, max_pt])
plt.plot(X_train, y_train, 'o');
# -
# Similar to the estimators for classification in the previous notebook, we use the `predict` method to predict the target variable. And we expect these predicted values to fall onto the line that we plotted previously:
y_pred_train = regressor.predict(X_train)
plt.plot(X_train, y_train, 'o', label="data")
plt.plot(X_train, y_pred_train, 'o', label="prediction")
plt.plot([X.min(), X.max()], [min_pt, max_pt], label='fit')
plt.legend(loc='best')
# As we can see in the plot above, the line is able to capture the general slope of the data, but not many details.
# Next, let's try the test set:
y_pred_test = regressor.predict(X_test)
plt.plot(X_test, y_test, 'o', label="data")
plt.plot(X_test, y_pred_test, 'o', label="prediction")
plt.plot([X.min(), X.max()], [min_pt, max_pt], label='fit')
plt.legend(loc='best');
# Again, scikit-learn provides an easy way to evaluate the prediction quantitatively using the ``score`` method. For regression tasks, this is the R<sup>2</sup> score. Another popular way would be the Mean Squared Error (MSE). As its name implies, the MSE is simply the average squared difference over the predicted and actual target values
#
# $$MSE = \frac{1}{n} \sum_{i=1}^{n} (\text{predicted}_i - \text{true}_i)^2$$
regressor.score(X_test, y_test)
# <div class="alert alert-success">
# <b>EXERCISE</b>:
# <ul>
# <li>
# Add a feature containing `sin(4x)` to `X` and redo the fit. Visualize the predictions with this new richer, yet linear, model.
# </li>
# </ul>
# </div>
# +
# # %load solutions/06B_lin_with_sine.py
# -
# KNeighborsRegression
# =======================
# As for classification, we can also use a neighbor based method for regression. We can simply take the output of the nearest point, or we could average several nearest points. This method is less popular for regression than for classification, but still a good baseline.
from sklearn.neighbors import KNeighborsRegressor
kneighbor_regression = KNeighborsRegressor(n_neighbors=1)
kneighbor_regression.fit(X_train, y_train)
# Again, let us look at the behavior on training and test set:
# +
y_pred_train = kneighbor_regression.predict(X_train)
plt.plot(X_train, y_train, 'o', label="data", markersize=10)
plt.plot(X_train, y_pred_train, 's', label="prediction", markersize=4)
plt.legend(loc='best');
# -
# On the training set, we do a perfect job: each point is its own nearest neighbor!
# +
y_pred_test = kneighbor_regression.predict(X_test)
plt.plot(X_test, y_test, 'o', label="data", markersize=8)
plt.plot(X_test, y_pred_test, 's', label="prediction", markersize=4)
plt.legend(loc='best');
# -
# On the test set, we also do a better job of capturing the variation, but our estimates look much messier than before.
# Let us look at the R<sup>2</sup> score:
kneighbor_regression.score(X_test, y_test)
# Much better than before! Here, the linear model was not a good fit for our problem; it was lacking in complexity and thus under-fit our data.
# <div class="alert alert-success">
# <b>EXERCISE</b>:
# <ul>
# <li>
# Compare the KNeighborsRegressor and LinearRegression on the boston housing dataset. You can load the dataset using ``sklearn.datasets.load_boston``. You can learn about the dataset by reading the ``DESCR`` attribute.
# </li>
# </ul>
# </div>
# +
# # %load solutions/06A_knn_vs_linreg.py
| notebooks/06.Supervised_Learning-Regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="3In88vKrq4WB"
# # Causality Tutorial Exercises – Python
#
# Contributors: <NAME>, <NAME>, <NAME>, <NAME>, <NAME>.
# The MIT License applies; copyright is with the authors.
# Some exercises are adapted from "Elements of Causal Inference: Foundations and Learning Algorithms" by <NAME>, <NAME> and <NAME>.
#
# + [markdown] id="KnsIE8yWlVIQ"
# # Exercise 1 – Structural Causal Model
#
# + [markdown] id="FSNemB3GrBIE"
#
# Let's first draw a sample from an SCM
# + id="5Cy58Ut1liKd"
import numpy as np
# set seed
np.random.seed(1)
rnorm = lambda n: np.random.normal(size=n)
n = 200
C = rnorm(n)
A = .8 * rnorm(n)
K = A + .1 * rnorm(n)
X = C - 2 * A + .2 * rnorm(n)
F = 3 * X + .8 * rnorm(n)
D = -2 * X + .5 * rnorm(n)
G = D + .5 * rnorm(n)
Y = 2 * K - D + .2 * rnorm(n)
H = .5 * Y + .1 * rnorm(n)
data = np.c_[C, A, K, X, F, D, G, Y, H]
# + [markdown] id="8PMvvEeIoKFN"
# __a)__
#
# What are the parents and children of $X$ in the above SCM ?
#
# Take a pair of variables and think about whether you expect this pair to be dependent
# (at this stage, you can only guess, later you will have tools to know). Check empirically.
#
# __b)__
#
# Generate a sample of size 300 from the interventional distribution $P_{\mathrm{do}(X=\mathcal{N}(2, 1))}$
# and store the data matrix as `data_int`.
# + id="FtbA6c2Ron5f"
# + [markdown] id="l3wOg_4vozpz"
# __c)__
#
# Do you expect the marginal distribution of $Y$ to be different in both samples?
# + [markdown] id="3paV1bkro6lV"
# Double-click (or enter) to edit
# + [markdown] id="CH9Tt444o-RH"
# __d)__
#
# Do you expect the joint distribution of $(A, Y)$ to be different in both samples?
#
# + [markdown] id="FJz4fZKEpE4-"
# Double-click (or enter) to edit
# + [markdown] id="eZmh_AizpGp-"
# __e)__
#
# Check your answers to c) and d) empirically.
# + id="q2PMSXqKpLpH"
# + [markdown] id="1Idk_ElwrEht"
# # Exercise 2 – Adjusting
# + [markdown] id="il0b9fnVq-bz"
#
# 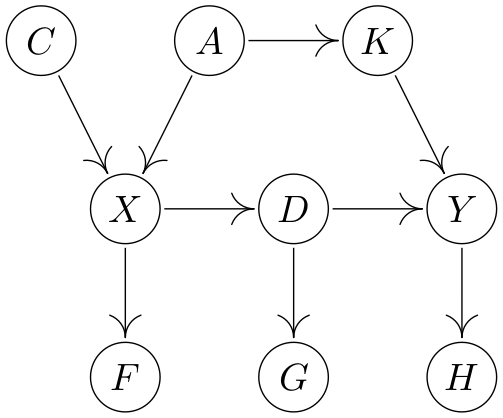
#
# Suppose we are given a fixed DAG (like the one above).
#
# a) What are valid adjustment sets (VAS) used for?
#
# b) Assume we want to find a VAS for the causal effect from $X$ to $Y$.
# What are general recipies (plural 😉) for constructing VASs (no proof)?
# Which sets are VAS in the DAG above?
#
# c) The following code samples from an SCM. Perform linear regressions using different VAS and compare the regression coefficient against the causal effect from $X$ to $Y$.
#
# + id="R3y5ckYKJHiJ"
import numpy as np
# set seed
np.random.seed(1)
rnorm = lambda n: np.random.normal(size=n)
n = 200
C = rnorm(n)
A = .8 * rnorm(n)
K = A + .1 * rnorm(n)
X = C - 2 * A + .2 * rnorm(n)
F = 3 * X + .8 * rnorm(n)
D = -2 * X + .5 * rnorm(n)
G = D + .5 * rnorm(n)
Y = 2 * K - D + .2 * rnorm(n)
H = .5 * Y + .1 * rnorm(n)
data = np.c_[C, A, K, X, F, D, G, Y, H]
# + [markdown] id="UqFFtwP5JQVw"
# d) Why could it be interesting to have several options for choosing a VAS?
#
# e) If you indeed have access to several VASs, what would you do?
# + [markdown] id="LQ7RuuF4rMD6"
# # Exercise 3 – Independence-based Causal Structure Learning
# + [markdown] id="p21N9AFBrB0o"
# __a)__
#
# Assume $P^{X,Y,Z}$ is Markov and faithful wrt. $G$. Assume all (!) conditional independences are
#
# $$
# \newcommand{\indep}{{\,⫫\,}}
# \newcommand{\dep}{\not{}\!\!\indep}
# $$
#
# $$X \dep Z \mid \emptyset$$
#
# (plus symmetric statements). What is $G$?
#
# __b)__
#
# Assume $P^{W,X,Y,Z}$ is Markov and faithful wrt. $G$. Assume all (!) conditional independences are
#
# $$\begin{aligned}
# (Y,Z) &\indep W \mid \emptyset \\
# W &\indep Y \mid (X,Z) \\
# (X,W) &\indep Y | Z
# \end{aligned}
# $$
#
# (plus symmetric statements). What is $G$?
# + [markdown] id="craCADN8rKd3"
# # Exercise 4 – Additive Noise Models
# + [markdown] id="OlFh1Zk50_z7"
# Set-up required packages:
# + id="qk3IE7jvvUxG"
# set up – not needed when run on mybinder
# if needed (colab), change False to True and run cell
if False:
# !mkdir ../data/
# !wget https://raw.githubusercontent.com/sweichwald/causality-tutorial-exercises/main/data/Exercise-ANM.csv -q -O ../data/Exercise-ANM.csv
# !wget https://raw.githubusercontent.com/sweichwald/causality-tutorial-exercises/main/python/kerpy/__init__.py -q -O kerpy.py
# !pip install pygam
# + id="GNsEcFUJ1P4I"
from kerpy import hsic
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from pygam import GAM, s
# + [markdown] id="pmh91goS1DCT"
# Let's load and plot some real data set:
# + id="2hwvlkYX1EPW"
data = pd.read_csv('../data/Exercise-ANM.csv')
plt.scatter(data["X"].values, data["Y"].values, s=2.);
# + [markdown] id="-uDnv5eD2pGd"
# __a)__
#
# Do you believed that $X \to Y$ or that $X \gets Y$? Why?
# + [markdown] id="4owvM1J_2rcM"
# Double-click (or enter) to edit
# + [markdown] id="mYdffpZN2uDc"
# $$
# \newcommand{\indep}{{\,⫫\,}}
# \newcommand{\dep}{\not{}\!\!\indep}
# $$
#
# __b)__
# Let us now try to get a more statistical answer. We have heard that we cannot
# have
# $$Y = f(X) + N_Y,\ N_Y \indep X$$
# and
# $$X = g(Y) + N_X,\ N_X \indep Y$$
# at the same time.
#
# Given a data set over $(X,Y)$,
# we now want to decide for one of the two models.
#
# Come up with a method to do so.
#
# Hints:
# * `GAM(s(0)).fit(A, B).deviance_residuals(A, B)` provides residuals when regressing $B$ on $A$.
# * `hsic(a, b)` can be used as an independence test (here, `a` and `b` are $n \times 1$ numpy arrays).
# + id="llz5Eeck2xz5"
# + [markdown] id="o8SBfEFi6oqH"
# __c)__
#
# Assume that the error terms are Gaussian with zero mean and variances
# $\sigma_X^2$ and $\sigma_Y^2$, respectively.
# The maximum likelihood for DAG G is
# then proportional to
# $-\log(\mathrm{var}(R^G_X)) - \log(\mathrm{var}(R^G_Y))$,
# where $R^G_X$ and $R^G_Y$ are the residuals obtained from regressing $X$ and $Y$ on
# their parents in $G$, respectively (no proof).
#
# Find the maximum likelihood solution.
# + id="pASFG1DC6sQA"
# + [markdown] id="d4JPYnSHrOfW"
# # Exercise 5 – Invariant Causal Prediction
# + [markdown] id="Fb5CwEUEAaOp"
# Set-up required packages and data:
# + id="wudBwtYswFeo"
# set up – not needed when run on mybinder
# if needed (colab), change False to True and run cell
if False:
# !mkdir ../data/
# !wget https://raw.githubusercontent.com/sweichwald/causality-tutorial-exercises/main/data/Exercise-ICP.csv -q -O ../data/Exercise-ICP.csv
# + id="DIosbymbbkhg"
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
import statsmodels.api as sm
# + [markdown] id="gy8eUIaDdmrz"
# __a)__
#
# Generate some observational and interventional data:
# + id="IGBQGfetbYPj"
# Generate n=1000 observations from the observational distribution
na = 1000
Xa = np.random.normal(size=na)
Ya = 1.5*Xa + np.random.normal(size=na)
# Generate n=1000 observations from an interventional distribution
nb = 1000
Xb = np.random.normal(loc=2, scale=1, size=nb)
Yb = 1.5*Xb + np.random.normal(size=nb)
# plot Y vs X1
fig, ax = plt.subplots(figsize=(7,5))
ax.scatter(Xa, Ya, label='observational', marker='o', alpha=0.6)
ax.scatter(Xb, Yb, label='interventional', marker ='^', alpha=0.6)
ax.legend();
# + [markdown] id="uZcSibWjypDR"
# Look at the above plot. Is the predictor $\{X\}$ an invariant set, that is (roughly speaking), does $Y \mid X = x$ have the same distribution in the orange and blue data?
# + [markdown] id="rhnmzEIiyvmt"
# Double-click (or enter) to edit
# + [markdown] id="DnDdgV_QeEFH"
# __b)__
# + [markdown] id="BqcN5gRdeRoi"
# We now consider data over a response and three covariates $X1, X2$, and $X3$
# and try to infer $\mathrm{pa}(Y)$. To do so, we need to find all sets for which this
# invariance is satisfied.
# + id="i4vMv59_wjKG"
# load data
data = pd.read_csv('../data/Exercise-ICP.csv')
data['env'] = np.concatenate([np.repeat('observational', 140), np.repeat('interventional', 80)])
# pairplot
sns.pairplot(data, hue='env', height=2, plot_kws={'alpha':0.6});
# + id="2yF7KhYZe7g9"
# The code below plots the residuals versus fitted values for all sets of
# predictors.
# extract response and predictors
Y = data['Y'].to_numpy()
X = data[['X1','X2','X3']].to_numpy()
# get environment indicator
obs_ind = data[data['env'] == 'observational'].index
int_ind = data[data['env'] == 'interventional'].index
# create all sets
all_sets = [(0,), (1,), (2,), (0,1), (0,2), (1,2), (0,1,2)]
# label each set
set_labels = ['X1', 'X2', 'X3', 'X1,X2', 'X1,X3', 'X2,X3', 'X1,X2,X3']
# fit OLS and store fitted values and residuals for each set
fitted = []
resid = []
for s in all_sets:
model = sm.OLS(Y, X[:, s]).fit()
fitted += [model.fittedvalues]
resid += [model.resid]
# plotting function
def plot_fitted_resid(fv, res, ax, title):
ax.scatter(fv[obs_ind], res[obs_ind], label='observational', marker='o', alpha=0.6)
ax.scatter(fv[int_ind], res[int_ind], label='interventional', marker ='^', alpha=0.6)
ax.legend()
ax.set_xlabel('fitted values')
ax.set_ylabel('residuals')
ax.set_title(title)
# creating plots
fig, axes = plt.subplots(4, 2, figsize=(7,14))
# plot result for the empty set predictor
ax0 = axes[0,0]
ax0.scatter(obs_ind, Y[obs_ind], label='observational', marker='o', alpha=0.6)
ax0.scatter(int_ind, Y[int_ind], label='interventional', marker ='^', alpha=0.6)
ax0.legend()
ax0.set_xlabel('index')
ax0.set_ylabel('Y')
ax0.set_title('empty set')
# plot result for the other sets
for i, ax in enumerate(axes.flatten()[1:]):
plot_fitted_resid(fitted[i], resid[i], ax, set_labels[i])
# make tight layout
plt.tight_layout()
# + [markdown] id="1GfZKCL7zJve"
# Which of the sets are invariant? (There are two plots with four scatter plots each.)
# + [markdown] id="j0sgjfRSzWEt"
# Double-click (or enter) to edit
# + [markdown] id="AO7tZSjLzMr0"
# __c)__
# What is your best guess for $\mathrm{pa}(Y)$?
# + [markdown] id="B6QtA9p9zdD7"
# Double-click (or enter) to edit
# + [markdown] id="AZGGVS8lP0Ly"
# __d) (optional)__
#
# Use the function ICP to check your result.
# + id="1Qi2_GCnQmEG"
# set up – not needed when run on mybinder
# if needed (colab), change False to True and run cell
if False:
# !pip install causalicp
# + id="fqUzMXw5QLva"
import causalicp as icp
| python/Causality_Tutorial_Exercises.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
pd.options.display.max_columns = 500
def fusion_tot(n):
mainpath= "./1_Extraction/WOS/All_databases/savedrecs"
frames =[]
for k in range(n):
file = mainpath + str(k) +".txt"
df = pd.read_table(file, sep = '\t', header = 0, index_col=False)
frames.append(df)
Df = pd.concat(frames, sort=False)
return(Df)
def fusion_core(n):
mainpath= "./1_Extraction/WOS/Core_collection/savedrecs"
frames =[]
for k in range(n):
file = mainpath + str(k) +".txt"
df = pd.read_table(file, sep = '\t', header = 0, index_col=False)
frames.append(df)
Df = pd.concat(frames, sort=False)
return(Df)
df = fusion_tot(8)
print("%d X %d dataframe" % (len(df), len(df.columns) ))
df["DE"] = np.nan
df_key_null = df.groupby(['UT']).first().reset_index()
print("%d X %d dataframe" % (len(df_key_null), len(df_key_null.columns) ))
df1 = fusion_core(8)
print("%d X %d dataframe" % (len(df1), len(df1.columns) ))
df2 = pd.read_table("./1_Extraction/WOS/Current_contents_connect/savedrecs0.txt", sep = '\t', header = 0, index_col=False)
df3 = pd.read_table("./1_Extraction/WOS/KCI-Korean/savedrecs0.txt", sep = '\t', header = 0, index_col=False)
df4 = pd.read_table("./1_Extraction/WOS/MEDLINE/savedrecs0.txt", sep = '\t', header = 0, index_col=False)
df5 = pd.read_table("./1_Extraction/WOS/Russian_Science/savedrecs0.txt", sep = '\t', header = 0, index_col=False)
df6 = pd.read_table("./1_Extraction/WOS/SciELO/savedrecs0.txt", sep = '\t', header = 0, index_col=False)
df.set_index("UT", inplace = True)
df1.set_index("UT", inplace = True)
df2.set_index("UT", inplace = True)
df3.set_index("UT", inplace = True)
df4.set_index("UT", inplace = True)
df5.set_index("UT", inplace = True)
df6.set_index("UT", inplace = True)
df.update(df1)
df.update(df2)
df.update(df3)
df.update(df4)
df.update(df5)
df.update(df6)
df.reset_index(0, inplace =True)
df.head(1)
df.to_excel("./1_Extraction/WOS/WOS_all_data_fusion.xlsx")
df.rename(columns ={'UT': 'WOS_number',
'TI': 'title',
'PT': 'doc_type',
'AU': 'authors',
'SO': 'source',
'DI': 'doi',
'PY': 'publication_year',
'AB': 'abstract',
'DE': 'author_keywords'}, inplace=True)
col=['WOS_number','doc_type','authors','title','source','doi','publication_year','abstract','author_keywords','email']
df = df.reindex(columns=col)
df.head()
print("%d X %d dataframe" % (len(df), len(df.columns) ))
import joblib
joblib.dump((df), "df_WOS.pkl")
| 1_Constitution_database/.ipynb_checkpoints/1.Fusion_WOS-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Transfer Learning on pretrained VGGish model for emotion classification
#
# This notebook shows how we can use a pretrained model that was previously used to create embeddings for Youtube audio ([VGGish](https://github.com/tensorflow/models/tree/master/research/audioset)) to train a classifier to recognise emotions in a much smaller dataset ([RAVDESS](https://smartlaboratory.org/ravdess/)).
#
# ## Setup
#
# Before getting started we should make sure that we have a suitable GPU for the task that is CUDA compatible so that we can run tensorflow. I trained this model on a relatively recent (at time of writing this) Dell Windows laptop with an i7-7700HQ, a GTX 1060-MaxQ, and 16GB of RAM. Running the whole notebook takes under 30 minutes for me (a decent chunk of which is loading and preprocessing the data).
#
# With these requirements met we can start setting the project up. First we will need to fetch a few files and load up the environment:
#
# 1. Download the pretrained VGG model from [here](https://storage.googleapis.com/audioset/vggish_model.ckpt)
# 1. Move the VGG model into `./pretrained_models/`
# 1. Download the RAVDESS speech dataset from [here](https://zenodo.org/record/1188976/files/Audio_Speech_Actors_01-24.zip?download=1)
# 1. Extract the zip into `./data`
# 1. Install [conda](https://conda.io/)
# 1. Install dependencies: `conda env create -f environment.yml`
# 1. Activate environment: `source activate vggec`
# 1. Run jupyter: `jupyter notebook`
# 1. Open the notebook in your browser.
# 1. Hit run!
#
# ## Load labels
#
# First we need to load our label data and get it into a shape we can work with. The labels are given in the `.wav` file names, so we'll have to load the names for processing:
# +
from pathlib import Path
data_path = Path("../data")
data_path
# -
# ### List all WAV files
wav_files = data_path.glob("*.wav")
wav_files = list(wav_files)
wav_files[:5]
# ### Map file names to their classes.
#
# Each emotion is labelled as 01 - 08, so we convert that to labels 0 - 7. However, we will later find out that the model has difficulty keeping some similar emotions apart, so we combine a few and end up with labels 0 - 4.
#
# We also need the speaker information so that we can later split the data into train/val/test based on speaker, to evaluate how well the model performs when it encouters unknown speakers (something it would obviously have to deal with in the real world).
# +
def class_from_file_name(fname):
# neutral + calm
# sad + fearful
# happy
# angry + disgust
# surprised
original_class = int(fname.split('-')[2]) - 1
if original_class in [0, 1]:
return 0
if original_class in [3, 5]:
return 1
if original_class in [2]:
return 2
if original_class in [4, 6]:
return 3
return 4
def speaker_from_file_name(fname):
return int(fname.split('-')[6].split('.')[0]) - 1
labels = {
f.name: {
"class": class_from_file_name(f.name),
"speaker": speaker_from_file_name(f.name)
}
for f in wav_files
}
[(k, v) for k, v in labels.items()][:5]
# -
# ### Extract number of classes
CLASSES = list(set([v["class"] for v in labels.values()]))
NUM_CLASSES = len(CLASSES)
NUM_CLASSES
# ## Load Data
# ### Fix PYTHONPATH
#
# The original VGGish repository includes some files for loading the original trained model as well as preprocessing the audio data into the input shape the model expects. I have copied those files for convenience. We just need to add the path to them to the pythonpath so that we can import the modules:
import os
import sys
nb_dir = Path(os.getcwd())
vgg_dir = nb_dir.parent / 'vgg'
vgg_dir
if str(vgg_dir) not in sys.path:
sys.path.append(str(vgg_dir))
sys.path
# ### Read WAV files
#
# Read in the wav files and convert them into the correct shape for the VGGish model (this is thankfully taken care of already by the example code provided)
from vggish_input import wavfile_to_examples
data = {
f.name: wavfile_to_examples(str(f))
for f in wav_files
}
data['03-01-01-01-01-01-01.wav'].shape
# ### Split dataset
#
# As previously mentioned, we need to split the different sets by speaker - especially since there are multiple files per speaker saying the same thing with slightly different intonation, which would result in some overlap between the sets otherwise!
#
# Split into
#
# * train: 80%
# * val: 10%
# * test: 10%
#
# (This split works a bit better to create even splits since we only have 24 speakers.)
# +
from random import shuffle
speakers = list(set([v["speaker"] for k, v in labels.items()]))
seed = 987234871
shuffle(speakers)
train_speakers_index = int(0.8 * len(speakers))
val_speakers_index = int(0.9 * len(speakers))
train_speakers = speakers[:train_speakers_index]
val_speakers = speakers[train_speakers_index:val_speakers_index]
test_speakers = speakers[val_speakers_index:]
print(f"Training speakers: {train_speakers}")
print(f"Validation speakers: {val_speakers}")
print(f"Test speakers: {test_speakers}")
# -
# Now that we have split the speakers into groups, we can can split the actual data by speaker:
# +
import numpy as np
x_train, y_train = zip(*[(data[key], value["class"]) for key, value in labels.items() if value["speaker"] in train_speakers])
x_val, y_val = zip(*[(data[key], value["class"]) for key, value in labels.items() if value["speaker"] in val_speakers])
x_test, y_test = zip(*[(data[key], value["class"]) for key, value in labels.items() if value["speaker"] in test_speakers])
x_train = np.array(x_train)
x_val = np.array(x_val)
x_test = np.array(x_test)
y_train = np.array(y_train)
y_val = np.array(y_val)
y_test = np.array(y_test)
print(f"Training size: {len(x_train)}")
print(f"Validation size: {len(x_val)}")
print(f"Test size: {len(x_test)}")
# -
# ### Convert labels to one-hot vectors
#
# For multi-class classification using categorical crossentropy we want the labels in one-hot encoded form.
#
# E.g. label `1` becomes `[0, 1, 0, 0 ,0 ,0, 0, 0]`
# +
import numpy as np
def to_one_hot(y, num_classes):
y_one_hot = np.zeros(((y.size, num_classes)))
y_one_hot[np.arange(y.size), y] = 1
return y_one_hot
# -
y_train = to_one_hot(y_train, NUM_CLASSES)
y_val = to_one_hot(y_val, NUM_CLASSES)
y_test = to_one_hot(y_test, NUM_CLASSES)
y_train[:5]
# ### Compute balanced weights
#
# The number of instances for each class isn't balanced, so we need to create weightings for each class to even things out during training
# +
from sklearn.utils import class_weight
class_weights = class_weight.compute_class_weight('balanced',
CLASSES,
list([v["class"] for v in labels.values()]))
class_weights
# -
# ### Create function to get batches from data
from random import shuffle
def get_shuffled_batches(x, y):
assert len(x) == len(y)
indexes = list(range(len(x)))
shuffle(indexes)
for x, y in zip(x[indexes], y[indexes]):
x_batch = np.reshape(x, (1, *x.shape, 1))
y_batch = np.reshape(y, (1, *y.shape))
yield x_batch, y_batch
# ## Convert pretrained TF weights to Keras model checkpoint
# Don't want to deal with the TF library, Keras is much easier to use imo. That means we first need to turn the TF checkpoint into a Keras checkpoint.
#
# It should theoretically be possible to load the TF checkpoint as is, but this way I know it'll work.
# ### Define exact VGGish model in Keras
#
# Code taken from https://github.com/SNeugber/vggish2Keras, which is a clone of https://github.com/antoinemrcr/vggish2Keras
# +
from keras.layers import Input, Dense, Conv2D, MaxPooling2D, Flatten
from keras.models import Model
import vggish_params
def get_vggish_keras():
input_shape = (vggish_params.NUM_FRAMES,vggish_params.NUM_BANDS,1)
img_input = Input( shape=input_shape)
# Block 1
x = Conv2D(64, (3, 3), activation='relu', padding='same', name='conv1')(img_input)
x = MaxPooling2D((2, 2), strides=(2, 2), name='pool1')(x)
# Block 2
x = Conv2D(128, (3, 3), activation='relu', padding='same', name='conv2')(x)
x = MaxPooling2D((2, 2), strides=(2, 2), name='pool2')(x)
# Block 3
x = Conv2D(256, (3, 3), activation='relu', padding='same', name='conv3_1')(x)
x = Conv2D(256, (3, 3), activation='relu', padding='same', name='conv3_2')(x)
x = MaxPooling2D((2, 2), strides=(2, 2), name='pool3')(x)
# Block 4
x = Conv2D(512, (3, 3), activation='relu', padding='same', name='conv4_1')(x)
x = Conv2D(512, (3, 3), activation='relu', padding='same', name='conv4_2')(x)
x = MaxPooling2D((2, 2), strides=(2, 2), name='pool4')(x)
# Block fc
x = Flatten(name='flatten')(x)
x = Dense(4096, activation='relu', name='fc1_1')(x)
x = Dense(4096, activation='relu', name='fc1_2')(x)
x = Dense(vggish_params.EMBEDDING_SIZE, activation='relu', name='fc2')(x)
model = Model(img_input, x, name='vggish')
return model
# -
# ### Load weights
#
# Need to turn TF layer names into Keras layer names
# +
import tensorflow as tf
import vggish_slim
with tf.Graph().as_default(), tf.Session() as sess:
vggish_slim.define_vggish_slim(training=False)
vggish_slim.load_vggish_slim_checkpoint(sess, '../pretrained_models/vggish_model.ckpt')
weights = {}
operations = sess.graph.get_operations()
for op in operations:
name = op.name
if 'read' in name:
name2 = name.replace('vggish/','').replace('/read','').replace('conv3/','').replace('conv4/','').replace('/fc1','')
name2_layer, name2_type = name2.split('/')
if name2_type == 'weights':
weights[name2_layer] = []
weights[name2_layer].append(sess.run(op.values())[0])
for op in operations:
name = op.name
if 'read' in name:
name2 = name.replace('vggish/','').replace('/read','').replace('conv3/','').replace('conv4/','').replace('/fc1','')
name2_layer, name2_type = name2.split('/')
if name2_type == 'biases':
weights[name2_layer].append(sess.run(op.values())[0])
# -
weights.keys()
# ### Save as Keras model
model = get_vggish_keras()
model.summary()
for layer in model.layers:
if layer.name in list(weights.keys()):
layer.set_weights(weights[layer.name])
model.save_weights('../pretrained_models/vgg_model.h5')
# ## Load VGGish model wrapped in TimeDistributed layer
#
# The normal model only works on a single time-frame. We want to train on multiple frames, so we need to wrap everything in Keras' [TimeDistributed](https://keras.io/layers/wrappers/#TimeDistributed)
# +
from keras.layers import TimeDistributed, Dropout
def get_vggish_keras_timedistributed(dropout_between_dense = 0):
input_shape = (None, vggish_params.NUM_FRAMES,vggish_params.NUM_BANDS,1)
img_input = Input(shape=input_shape)
# Block 1
x = TimeDistributed(Conv2D(64, (3, 3), activation='relu', padding='same'), name='conv1')(img_input)
x = TimeDistributed(MaxPooling2D((2, 2), strides=(2, 2)), name='pool1')(x)
# Block 2
x = TimeDistributed(Conv2D(128, (3, 3), activation='relu', padding='same', name='conv2'))(x)
x = TimeDistributed(MaxPooling2D((2, 2), strides=(2, 2)), name='pool2')(x)
# Block 3
x = TimeDistributed(Conv2D(256, (3, 3), activation='relu', padding='same', name='conv3_1'))(x)
x = TimeDistributed(Conv2D(256, (3, 3), activation='relu', padding='same', name='conv3_2'))(x)
x = TimeDistributed(MaxPooling2D((2, 2), strides=(2, 2)), name='pool3')(x)
# Block 4
x = TimeDistributed(Conv2D(512, (3, 3), activation='relu', padding='same', name='conv4_1'))(x)
x = TimeDistributed(Conv2D(512, (3, 3), activation='relu', padding='same', name='conv4_2'))(x)
x = TimeDistributed(MaxPooling2D((2, 2), strides=(2, 2)), name='pool4')(x)
# Block fc
x = TimeDistributed(Flatten(), name='flatten')(x)
x = TimeDistributed(Dense(4096, activation='relu'), name='fc1_1')(x)
x = TimeDistributed(Dropout(dropout_between_dense), name='dropout_1')(x)
x = TimeDistributed(Dense(4096, activation='relu'), name='fc1_2')(x)
x = TimeDistributed(Dropout(dropout_between_dense), name='dropout_2')(x)
x = TimeDistributed(Dense(vggish_params.EMBEDDING_SIZE, activation='relu'), name='fc2')(x)
model = Model(img_input, x, name='vggish_across_time')
return model
# -
model = get_vggish_keras_timedistributed()
model.summary()
model.load_weights('../pretrained_models/vgg_model.h5')
# +
from keras.layers import CuDNNLSTM
from tensorflow import Tensor
x = CuDNNLSTM(127)(model.layers[-1].output)
x = Dense(NUM_CLASSES, activation='softmax', name='out')(x)
final_model = Model(inputs=model.input, outputs=[x])
final_model.summary()
# -
# ## Model Training
#
# Putting the pretrained model to use!
#
# ### Compile model
#
# First we need to compile the model, for which we'll use the same parameters (for now) as used originally:
# * Adam optimizer
# * LR of 1e-4
# * Adam Epsilon of 1e-8
# +
from keras.optimizers import Adam
from keras.metrics import categorical_accuracy
model = final_model
optimizer = Adam(lr=vggish_params.LEARNING_RATE, epsilon=vggish_params.ADAM_EPSILON)
metrics = [categorical_accuracy]
model.compile(optimizer=optimizer, loss='categorical_crossentropy', metrics=metrics)
# -
# ### Manual training
#
# Since we're dealing with var-length data, we can't just call `fit`, but instead need to train on each batch manually. Since a batch needs to be a numpy array with the correct dimensions (instead of a list of var-length numpy arrays) we are restricted to online learning (i.e. batch size == 1).
#
# We therefore need to pass each batch manually into `model.train_on_batch`
def train_epoch(model, x_train, y_train):
train_loss = 0.0
train_batches = 0.0
train_accuracy = 0.0
for x, y in get_shuffled_batches(x_train, y_train):
loss, acc = model.train_on_batch(x, y, class_weight=class_weights)
train_loss += loss
train_accuracy += acc
train_batches += 1
return train_loss / train_batches, train_accuracy / train_batches
# ### Manual validation
#
# Similar to training, we need to evaluate each batch in our validation data manually. For some reason the Keras API demands the class weights on a per-sample basis here, so we need to extract those first.
def sample_weights_from_class_weights(class_weights, labels):
return np.array([class_weights[np.argmax(label)] for label in labels])
sample_weights_from_class_weights(class_weights=class_weights, labels=y_train[[1, 400, 800, 1000]])
# Now we can define the validation function
def validate_epoch(model, x_val, y_val, class_weights):
val_loss = 0.0
val_batches = 0.0
val_accuracy = 0.0
for x, y in get_shuffled_batches(x_val, y_val):
sample_weights = sample_weights_from_class_weights(class_weights, y)
loss, acc = model.evaluate(x, y, batch_size=1, sample_weight=sample_weights, verbose=0)
val_loss += loss
val_accuracy += acc
val_batches += 1
return val_loss / val_batches, val_accuracy / val_batches
# ### Plotting progress
#
# In order to keep track of the training process, we'll create a plot function that plots the training/validation loss & accuracy.
#
# Updating the same plot in a notebook is a bit faffy, so we need to clear the current output every time. (**Note**: this still doesn't work perfectly due to all the plots being shown at the end)
#
# **TODO**: Turn this into a Keras callback
# +
from IPython import display
def plot_losses(epoch, train_loss, train_acc, val_loss, val_acc):
fig = plt.figure()
ax1 = fig.add_subplot(211)
ax2 = ax1.twinx()
ax3 = fig.add_subplot(212)
fig.canvas.draw()
epochs = list(range(epoch + 1))
ax1.set_xlabel("Epoch")
ax1.set_ylabel("Train Loss")
ax2.set_ylabel("Val Loss")
ax1.plot(epochs, train_loss)
ax2.plot(epochs, val_loss)
ax3.set_xlabel("Epoch")
ax3.set_ylabel("Accuracy")
ax3.plot(epochs, train_acc)
ax3.plot(epochs, val_acc)
ax3.legend()
fig.tight_layout()
fig.canvas.draw()
display.clear_output(wait=True)
display.display(fig)
# -
# ### Saving the best model
#
# Keras provides a number of predefined [Callbacks](https://keras.io/callbacks/) which it would automatically invoke before/after every batch/epoch. Even though we do the training manually we should still use the same API, since it makes it pretty obvious where callbacks should be invoked!
#
# Using the `ModelCheckpoint` callback we can make sure we save the best model based on the validation loss. The checkpoint name is just the current timestamp, epoch, and validation loss:
# +
from datetime import datetime
from keras.callbacks import ModelCheckpoint
import os
checkpoint_dir = "../model_checkpoints/"
if not os.path.exists(checkpoint_dir):
os.makedirs(checkpoint_dir)
timestamp = datetime.utcnow().strftime('%Y-%m-%d_%H-%M-%S')
model_checkpoint_filepath = checkpoint_dir + timestamp + "_epoch-{epoch:02d}_val-{val_loss:.4f}.hdf5"
model_checkpointer = ModelCheckpoint(model_checkpoint_filepath,
monitor='val_loss',
verbose=0,
save_best_only=True,
save_weights_only=False,
mode='auto',
period=1)
# -
# ### Training
#
# We now have everything in place so that we can start training. Each epoch the training loop needs to keep track of the losses, update the callback(s), and - obviously - perform the training.
# +
# %matplotlib inline
NUM_EPOCHS = 5
train_losses = []
train_accs = []
val_losses = []
val_accs = []
model_checkpointer.set_model(model)
model_checkpointer.on_train_begin()
for epoch in range(NUM_EPOCHS):
model_checkpointer.on_epoch_begin(epoch)
train_loss, train_acc = train_epoch(model, x_train, y_train)
train_losses.append(train_loss)
train_accs.append(train_acc)
val_loss, val_acc = validate_epoch(model, x_val, y_val, class_weights)
val_losses.append(val_loss)
val_accs.append(val_acc)
model_checkpointer.on_epoch_end(epoch, logs={
"loss": train_loss,
"val_loss": val_loss
})
plot_losses(epoch, train_losses, train_accs, val_losses, val_accs)
display.clear_output(wait=True)
plot_losses(epoch, train_losses, train_accs, val_losses, val_accs)
# -
# As we can see, the model starts overfitting after 3 epochs already (more on that later). Let's test how well the best-found fit does on our test set.
# ## Testing the model
#
# We have held a test-set back in order to verify that the model works on unseen data (and we don't overfit e.g. our hyperparameters to the validation set!).
#
# ### Loading the best model
#
# First we should use our checkpointer callback to load the best model. Since it only stores newer models if the validation loss is lower, we can just load the latest model with the known timestamp (assuming they are returned in descending order):
timestamp = "_".join(os.path.basename(model_checkpointer.filepath).split("_")[:2])
best_model_path = [os.path.join(checkpoint_dir, i) for i in os.listdir(checkpoint_dir)
if os.path.isfile(os.path.join(checkpoint_dir, i))
and timestamp in i][-1]
model.load_weights(best_model_path)
# ### Making predictions
#
# Same as with training/validation, run through each batch and create predictions for it.
predictions = []
true_labels = []
sample_weights = []
for x, y in get_shuffled_batches(x_test, y_test):
prediction = model.predict_on_batch(x)
predictions.append(prediction)
true_labels.append(y)
sample_weights.append(sample_weights_from_class_weights(class_weights, y))
# ### One-hot to categories
#
# The predictions the model returns are one-hot encoded, so we need to turn them back into categories.
#
# We also need to transform the sample weights to a 1D array for creating the balanced accuracy metric later.
#
# First we wrap them as numpy arrays:
# +
predictions = np.array(predictions)
true_labels = np.array(true_labels)
sample_weights = np.array(sample_weights)
print(predictions.shape)
print(true_labels.shape)
print(sample_weights.shape)
# -
# One-hot to categories:
# +
predictions = np.argmax(predictions, axis=2)
true_labels = np.argmax(true_labels, axis=2)
print(predictions.shape)
print(true_labels.shape)
# -
# Remove the 1D array for 1D batches:
# +
predictions = np.reshape(predictions, predictions.shape[0])
true_labels = np.reshape(true_labels, true_labels.shape[0])
sample_weights = np.reshape(sample_weights, sample_weights.shape[0])
print(predictions.shape)
print(true_labels.shape)
print(sample_weights.shape)
# -
# ### Calculate metrics
# +
from sklearn.metrics import precision_score, recall_score, f1_score, accuracy_score, confusion_matrix
metrics = dict(
precision = precision_score(true_labels, predictions, average=None),
recall = recall_score(true_labels, predictions, average=None),
f1 = f1_score(true_labels, predictions, average=None),
accuracy_raw = accuracy_score(true_labels, predictions),
accuracy_weighted = accuracy_score(true_labels, predictions, sample_weight=sample_weights),
confusion = confusion_matrix(true_labels, predictions)
)
metrics
# -
# ### Confusion matrix
#
# I'm pretty sure I copied this code from somewhere ages ago (i.e. the first result on StackOverflow). It just shows the confusion matrix as a heatmap with the relevant prediction results, either raw or normalised.
# +
import itertools
import matplotlib.pylab as plt
def plot_confusion_matrix(cm,
classes,
normalize=False,
title='Confusion matrix',
cmap=plt.cm.Blues,
block=False):
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
fmt = '.2f' if normalize else 'd'
thresh = np.average(cm)
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], fmt),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
if block:
plt.show()
# -
# Raw:
# +
# emotions = ["neutral", "calm", "happy", "sad", "angry", "fearful", "disgust", "surprised"]
emotions = ["neutral", "sad", "happy", "angry", "surprised"]
plot_confusion_matrix(metrics["confusion"], classes=emotions)
# -
# Normalised:
plot_confusion_matrix(metrics["confusion"], classes=emotions, normalize=True)
# ## Results
#
# The classifier isn't doing a half-bad job! It gets a lot of classes right, but clearly struggles with classifying happiness well. In some runs it gets to 80% accuracy overall, which is pretty decent across 5 classes, although the emotions we are most interested in (happy/sad) are also the least accurate. Previous results with all 8 classes got to about 60% accuracy, which resulted in me combining classes that had a lot of prediction overlap to make the problem a bit easier for the classifier.
#
# ## Improvements
#
# The model is overfitting quite quickly so we could look into some regularisation methods. For example, the dataset is only made up of 1441 samples, so scaling that up with data augmentation should be an easy win.
#
# We are currently also training the entire set of layers, which means even the very first few layers are able to specialise to our dataset. We could therefore see if just training e.g. the final few dense layers is sufficient.
#
# If more regularisation is required we can also look into adding some batch normalisation and/or dropout between our layers. Since we're re-training the entire model anyway, we may as well change it around a bit and see if that improves the results.
#
# Whether Adam is the best choice for online learning is also questionable (based on other models trained in the past), so looking into plain SGD + Nesterov momentum could be an option.
# ## Combine code for reuse
#
# Before moving on, let's clean up the code a little to make sure we can reuse it more easily going forward
# +
from keras.layers import CuDNNLSTM
from keras.optimizers import Adam
from keras.metrics import categorical_accuracy
from tensorflow import Tensor
from datetime import datetime
from keras.callbacks import ModelCheckpoint, Callback
import matplotlib as mpl
import os
from sklearn.metrics import precision_score, recall_score, f1_score, accuracy_score, confusion_matrix
from IPython import display
class PlotTrainProgress(Callback):
def __init__(self):
super().__init__()
self.train_loss_color = "C0"
self.val_loss_color = "C1"
mpl.style.use("seaborn-notebook")
self.logs = []
self.fig = None
self.losses = np.array([])
self.val_losses = np.array([])
self.accs = np.array([])
self.val_accs = np.array([])
self.x = []
self.i = 0
self.ax1 = None
self.ax2 = None
self.ax3 = None
self.ax4 = None
def _reset(self):
self.logs = []
self.fig = None
self.losses = np.array([])
self.val_losses = np.array([])
self.accs = np.array([])
self.val_accs = np.array([])
self.x = []
self.i = 0
self.ax1 = None
self.ax2 = None
self.ax3 = None
self.ax4 = None
def on_train_begin(self, logs={}):
self._reset()
self.fig = plt.figure()
self.ax1 = self.fig.add_subplot(211)
self.ax2 = self.ax1.twinx()
self.ax3 = self.fig.add_subplot(212)
self.ax1.tick_params("y", colors=self.train_loss_color)
self.ax2.tick_params("y", colors=self.val_loss_color)
self.fig.canvas.draw()
def on_epoch_end(self, epoch, logs={}):
self.logs.append(logs)
self.x.append(self.i)
self.losses = np.append(self.losses, logs.get('loss'))
self.val_losses = np.append(self.val_losses, logs.get('val_loss'))
self.accs = np.append(self.accs, logs.get('categorical_accuracy'))
self.val_accs = np.append(self.val_accs, logs.get('val_categorical_accuracy'))
self.i += 1
self.ax1.clear()
self.ax2.clear()
self.ax3.clear()
self.ax1.set_xlabel("Epoch")
self.ax1.set_ylabel("Train Loss", color=self.train_loss_color)
self.ax2.set_ylabel("Val Loss", color=self.val_loss_color)
self.ax1.plot(self.x, self.losses, color=self.train_loss_color)
self.ax2.plot(self.x, self.val_losses, color=self.val_loss_color)
self.ax3.set_xlabel("Epoch")
self.ax3.set_ylabel("Accuracy")
self.ax3.plot(self.x, self.accs, color=self.train_loss_color, label="Train Accuracy")
self.ax3.plot(self.x, self.val_accs, color=self.val_loss_color, label="Val Accuracy")
self.ax3.legend()
self.fig.tight_layout()
self.fig.canvas.draw()
display.clear_output(wait=True)
display.display(self.fig)
# def on_train_end(self, logs=None):
# display.clear_output(wait=True)
# display.display(self.fig)
# plt.ioff()
# plt.show()
def pretrained_vgg_with_custom_head():
model = get_vggish_keras_timedistributed()
model.load_weights('../pretrained_models/vgg_model.h5')
x = CuDNNLSTM(127)(model.layers[-1].output)
x = Dense(NUM_CLASSES, activation='softmax', name='out')(x)
model = Model(inputs=model.input, outputs=[x])
optimizer = Adam(lr=vggish_params.LEARNING_RATE, epsilon=vggish_params.ADAM_EPSILON)
metrics = [categorical_accuracy]
model.compile(optimizer=optimizer, loss='categorical_crossentropy', metrics=metrics)
return model
def get_model_checkpointer(checkpoint_dir = "../model_checkpoints/"):
if not os.path.exists(checkpoint_dir):
os.makedirs(checkpoint_dir)
timestamp = datetime.utcnow().strftime('%Y-%m-%d_%H-%M-%S')
model_checkpoint_filepath = checkpoint_dir + timestamp + "_epoch-{epoch:02d}_val-{val_loss:.4f}.hdf5"
model_checkpointer = ModelCheckpoint(model_checkpoint_filepath,
monitor='val_loss',
verbose=0,
save_best_only=True,
save_weights_only=False,
mode='auto',
period=1)
return model_checkpointer
def load_best_weights(model, model_checkpointer, checkpoint_dir = "../model_checkpoints/"):
timestamp = "_".join(os.path.basename(model_checkpointer.filepath).split("_")[:2])
best_model_path = [os.path.join(checkpoint_dir, i) for i in os.listdir(checkpoint_dir)
if os.path.isfile(os.path.join(checkpoint_dir, i))
and timestamp in i][-1]
model.load_weights(best_model_path)
return model
def train(model, x_train, y_train, x_val, y_val, class_weights, callbacks = None, num_epochs = 5):
callbacks = [] if callbacks is None else callbacks
for callback in callbacks:
callback.set_model(model)
callback.on_train_begin()
for epoch in range(num_epochs):
for callback in callbacks:
callback.on_epoch_begin(epoch)
train_loss, train_acc = train_epoch(model, x_train, y_train)
val_loss, val_acc = validate_epoch(model, x_val, y_val, class_weights)
for callback in callbacks:
callback.on_epoch_end(epoch, logs={
"loss": train_loss,
"val_loss": val_loss,
"categorical_accuracy": train_acc,
"val_categorical_accuracy": val_acc
})
for callback in callbacks:
callback.on_train_end()
for callback in callbacks:
if isinstance(callback, ModelCheckpoint):
model = load_best_weights(model, callback)
break
return model
def test(model, x_test, y_test, class_weights):
predictions = []
true_labels = []
sample_weights = []
for x, y in get_shuffled_batches(x_test, y_test):
prediction = model.predict_on_batch(x)
predictions.append(prediction)
true_labels.append(y)
sample_weights.append(sample_weights_from_class_weights(class_weights, y))
predictions = np.array(predictions)
true_labels = np.array(true_labels)
sample_weights = np.array(sample_weights)
predictions = np.argmax(predictions, axis=2)
true_labels = np.argmax(true_labels, axis=2)
predictions = np.reshape(predictions, predictions.shape[0])
true_labels = np.reshape(true_labels, true_labels.shape[0])
sample_weights = np.reshape(sample_weights, sample_weights.shape[0])
return dict(
precision = precision_score(true_labels, predictions, average=None),
recall = recall_score(true_labels, predictions, average=None),
f1 = f1_score(true_labels, predictions, average=None),
accuracy_raw = accuracy_score(true_labels, predictions),
accuracy_weighted = accuracy_score(true_labels, predictions, sample_weight=sample_weights),
confusion = confusion_matrix(true_labels, predictions)
)
# -
# ## Freezing layers
#
# The model quickly overfits to our dataset. We can reduce this by only training the last few layers.
# +
model = get_vggish_keras_timedistributed(dropout_between_dense = 0.5)
model.load_weights('../pretrained_models/vgg_model.h5')
x = CuDNNLSTM(127)(model.layers[-1].output)
x = Dense(NUM_CLASSES, activation='softmax', name='out')(x)
model = Model(inputs=model.input, outputs=[x])
for layer in model.layers[:-5]:
layer.trainable = False
optimizer = Adam(lr=vggish_params.LEARNING_RATE, epsilon=vggish_params.ADAM_EPSILON)
metrics = [categorical_accuracy]
model.compile(optimizer=optimizer, loss='categorical_crossentropy', metrics=metrics)
model.summary()
# +
# %matplotlib inline
callbacks = [
get_model_checkpointer(),
PlotTrainProgress()
]
model = train(model = model, x_train = x_train, y_train = y_train,
x_val = x_val, y_val = y_val, class_weights = class_weights, callbacks = callbacks, num_epochs = 10)
results = test(model = model, x_test = x_test, y_test = y_test, class_weights = class_weights)
# -
results
plot_confusion_matrix(results["confusion"], classes=emotions, normalize=True)
# ### Unlocking more layers with a lower learning rate
# +
import math
import keras.backend as K
model = get_vggish_keras_timedistributed(dropout_between_dense = 0)
model.load_weights('../pretrained_models/vgg_model.h5')
x = CuDNNLSTM(127)(model.layers[-1].output)
x = Dense(NUM_CLASSES, activation='softmax', name='out')(x)
model = Model(inputs=model.input, outputs=[x])
for layer in model.layers[:-5]:
layer.trainable = False
optimizer = Adam(lr=vggish_params.LEARNING_RATE, epsilon=vggish_params.ADAM_EPSILON)
metrics = [categorical_accuracy]
model.compile(optimizer=optimizer, loss='categorical_crossentropy', metrics=metrics)
callbacks = [
get_model_checkpointer(),
PlotTrainProgress()
]
callbacks = [] if callbacks is None else callbacks
for callback in callbacks:
callback.set_model(model)
callback.on_train_begin()
for epoch in range(10):
for callback in callbacks:
callback.on_epoch_begin(epoch)
train_loss, train_acc = train_epoch(model, x_train, y_train)
val_loss, val_acc = validate_epoch(model, x_val, y_val, class_weights)
for callback in callbacks:
callback.on_epoch_end(epoch, logs={
"loss": train_loss,
"val_loss": val_loss,
"categorical_accuracy": train_acc,
"val_categorical_accuracy": val_acc
})
# if epoch == 2:
# for layer in model.layers[-7:]:
# layer.trainable = True
# optimizer = Adam(lr=eval(model.optimizer.lr) / 2, epsilon=vggish_params.ADAM_EPSILON)
# model.compile(optimizer=optimizer, loss='categorical_crossentropy', metrics=metrics)
K.set_value(model.optimizer.lr, K.get_value(model.optimizer.lr) / 2)
# if epoch == 4:
# # for layer in model.layers:
# # layer.trainable = True
# optimizer = Adam(lr=eval(model.optimizer.lr) / 10, epsilon=vggish_params.ADAM_EPSILON)
# model.compile(optimizer=optimizer, loss='categorical_crossentropy', metrics=metrics)
# if epoch == 3:
# optimizer = Adam(lr=eval(model.optimizer.lr) / 10, epsilon=vggish_params.ADAM_EPSILON)
# model.compile(optimizer=optimizer, loss='categorical_crossentropy', metrics=metrics)
for callback in callbacks:
callback.on_train_end()
for callback in callbacks:
if isinstance(callback, ModelCheckpoint):
model = load_best_weights(model, callback)
break
results = test(model = model, x_test = x_test, y_test = y_test, class_weights = class_weights)
# -
results
emotions = ["neutral", "sad", "happy", "angry", "surprised"]
plot_confusion_matrix(results["confusion"], classes=emotions, normalize=True)
import keras.backend as K
K.clear_session()
# ## Data Augmentation
#
# To be continued...
#
# // last update: w/c 2018-11-19, should get this chapter out next week :)
# +
from pydub import AudioSegment
from pydub.playback import play
def read_audiosegment(file_path):
return AudioSegment.from_file(file_path, format="wav")
audio = read_audiosegment(str(wav_files[0]))
audio_datatype = np.array(audio.get_array_of_samples()).dtype
assert audio_datatype == np.int16
play(audio)
# +
def normalise_volume(audio_segment):
return audio_segment.apply_gain(-audio_segment.max_dBFS)
print("Max amplitude pre normalisation: ", max(audio.get_array_of_samples()))
audio = normalise_volume(audio)
print("Max amplitude post normalisation", max(audio.get_array_of_samples()))
play(audio)
# +
def detect_leading_silence(audio_segment, silence_threshold=-50.0, chunk_size=10):
trim_ms = 0 # ms
for pos in range(0, len(audio_segment), chunk_size):
seq = audio_segment[pos:pos + chunk_size]
if seq.dBFS < silence_threshold:
trim_ms += chunk_size
else:
break
return trim_ms
def trim_silence(audio_segment):
trim_start = detect_leading_silence(audio_segment)
trim_end_length = detect_leading_silence(audio_segment.reverse())
trim_end = len(audio_segment) - trim_end_length
return audio_segment[trim_start:trim_end]
print("Audio length before trimming: ", len(audio))
audio = trim_silence(audio)
print("Audio length after trimming: ", len(audio))
play(audio)
# +
# taken from: https://stackoverflow.com/questions/43408833/how-to-increase-decrease-playback-speed-on-wav-file
def change_speed(audio_segment, speed=1.0):
# Manually override the frame_rate. This tells the computer how many
# samples to play per second
altered_frame_rate = audio_segment._spawn(
audio_segment.raw_data,
overrides={
"frame_rate": int(audio_segment.frame_rate * speed)
}
)
# convert the sound with altered frame rate to a standard frame rate
# so that regular playback programs will work right. They often only
# know how to play audio at standard frame rate (like 44.1k)
return altered_frame_rate.set_frame_rate(audio_segment.frame_rate)
print("Audio length before changing speed: ", len(audio))
audio = change_speed(audio, speed=1.1)
print("Audio length sped up by 10%: ", len(audio))
play(audio)
# +
from pydub.generators import WhiteNoise
def add_noise(audio_segment, noise_db):
noise = WhiteNoise(sample_rate=audio_segment.frame_rate)
noise_seg = noise.to_audio_segment(len(audio_segment), volume=noise_db)
return audio_segment.overlay(noise_seg)
audio = add_noise(audio, -30)
play(audio)
# +
from vggish_input import waveform_to_examples
def audiosegment_to_samples(audio_segment):
raw_data = np.array(audio_segment.get_array_of_samples())
raw_data = raw_data / 32768.0 # convert to range [-1, 1]
return waveform_to_examples(raw_data, audio_segment.frame_rate)
input_data = audiosegment_to_samples(audio)
print(input_data.shape)
# -
def generate_augmentations(audio_segment):
for speed in [0.8, 0.9, 1.1, 1.2]:
yield change_speed(audio_segment, speed)
for noise_db in [-30, -40]:
yield add_noise(audio_segment, noise_db)
def wav_files_in_path(path):
data_path = Path(path)
wav_files = data_path.glob("*.wav")
for f in wav_files:
yield f.name, str(f)
wav_files[0]
# +
import types
class Step:
def __init__(self, func, creates_key=False):
self.func = func
self.creates_key = creates_key
def run(self, in_gen):
for elem in in_gen:
if not self.creates_key:
key, elem = elem
output = self.func(elem)
if isinstance(output, types.GeneratorType):
for output_elem in output:
if self.creates_key:
yield output_elem
else:
yield key, output_elem
else:
if self.creates_key:
yield output
else:
yield key, output
pipeline = [
read_audiosegment,
normalise_volume,
trim_silence,
generate_augmentations,
audiosegment_to_samples
]
input_step = Step(wav_files_in_path, creates_key=True)
compiled_pipeline = input_step.run(["../data"])
for func in pipeline:
step = Step(func)
compiled_pipeline = step.run(compiled_pipeline)
# +
from collections import defaultdict
data = defaultdict(list)
for key, output in compiled_pipeline:
data[key].append(output)
# +
import itertools
x_train, y_train = zip(*[(data[key], [value["class"]] * len(data[key])) for key, value in labels.items() if value["speaker"] in train_speakers])
x_val, y_val = zip(*[(data[key], [value["class"]] * len(data[key])) for key, value in labels.items() if value["speaker"] in val_speakers])
x_test, y_test = zip(*[(data[key], [value["class"]] * len(data[key])) for key, value in labels.items() if value["speaker"] in test_speakers])
x_train = list(itertools.chain.from_iterable(x_train))
x_val = list(itertools.chain.from_iterable(x_val))
x_test = list(itertools.chain.from_iterable(x_test))
y_train = list(itertools.chain.from_iterable(y_train))
y_val = list(itertools.chain.from_iterable(y_val))
y_test = list(itertools.chain.from_iterable(y_test))
x_train = np.array(x_train)
x_val = np.array(x_val)
x_test = np.array(x_test)
y_train = np.array(y_train)
y_val = np.array(y_val)
y_test = np.array(y_test)
y_train = to_one_hot(y_train, NUM_CLASSES)
y_val = to_one_hot(y_val, NUM_CLASSES)
y_test = to_one_hot(y_test, NUM_CLASSES)
# -
print(f"Training size: {len(x_train)}")
print(f"Validation size: {len(x_val)}")
print(f"Test size: {len(x_test)}")
# +
# %matplotlib inline
model = pretrained_vgg_with_custom_head()
model = train(model=model, x_train = x_train, y_train = y_train,
x_val = x_val, y_val = y_val, class_weights = class_weights)
results = test(model=model, x_test = x_test, y_test = y_test, class_weights = class_weights)
# +
emotions = ["neutral", "sad", "happy", "angry", "surprised"]
plot_confusion_matrix(results["confusion"], classes=emotions)
# -
plot_confusion_matrix(results["confusion"], classes=emotions, normalize=True)
| experiments/Transfer Learning on pretrained VGGish model for emotion classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Generative QA with "Retrieval-Augmented Generation"
#
# [](https://colab.research.google.com/github/deepset-ai/haystack/blob/master/tutorials/Tutorial7_RAG_Generator.ipynb)
#
# While extractive QA highlights the span of text that answers a query,
# generative QA can return a novel text answer that it has composed.
# In this tutorial, you will learn how to set up a generative system using the
# [RAG model](https://arxiv.org/abs/2005.11401) which conditions the
# answer generator on a set of retrieved documents.
# ### Prepare environment
#
# #### Colab: Enable the GPU runtime
# Make sure you enable the GPU runtime to experience decent speed in this tutorial.
# **Runtime -> Change Runtime type -> Hardware accelerator -> GPU**
#
# <img src="https://raw.githubusercontent.com/deepset-ai/haystack/master/docs/_src/img/colab_gpu_runtime.jpg">
# + pycharm={"name": "#%%\n"}
# Make sure you have a GPU running
# !nvidia-smi
# -
# Here are the packages and imports that we'll need:
# + pycharm={"name": "#%%\n"}
# !pip install grpcio-tools==1.34.1
# !pip install git+https://github.com/deepset-ai/haystack.git
# + pycharm={"name": "#%%\n"}
from typing import List
import requests
import pandas as pd
from haystack import Document
from haystack.document_store.faiss import FAISSDocumentStore
from haystack.generator.transformers import RAGenerator
from haystack.retriever.dense import DensePassageRetriever
# -
# Let's download a csv containing some sample text and preprocess the data.
#
# + pycharm={"name": "#%%\n"}
# Download sample
temp = requests.get("https://raw.githubusercontent.com/deepset-ai/haystack/master/tutorials/small_generator_dataset.csv")
open('small_generator_dataset.csv', 'wb').write(temp.content)
# Create dataframe with columns "title" and "text"
df = pd.read_csv("small_generator_dataset.csv", sep=',')
# Minimal cleaning
df.fillna(value="", inplace=True)
print(df.head())
# -
# We can cast our data into Haystack Document objects.
# Alternatively, we can also just use dictionaries with "text" and "meta" fields
# + pycharm={"name": "#%%\n"}
# Use data to initialize Document objects
titles = list(df["title"].values)
texts = list(df["text"].values)
documents: List[Document] = []
for title, text in zip(titles, texts):
documents.append(
Document(
text=text,
meta={
"name": title or ""
}
)
)
# -
# Here we initialize the FAISSDocumentStore, DensePassageRetriever and RAGenerator.
# FAISS is chosen here since it is optimized vector storage.
# + pycharm={"name": "#%%\n"}
# Initialize FAISS document store.
# Set `return_embedding` to `True`, so generator doesn't have to perform re-embedding
document_store = FAISSDocumentStore(
faiss_index_factory_str="Flat",
return_embedding=True
)
# Initialize DPR Retriever to encode documents, encode question and query documents
retriever = DensePassageRetriever(
document_store=document_store,
query_embedding_model="facebook/dpr-question_encoder-single-nq-base",
passage_embedding_model="facebook/dpr-ctx_encoder-single-nq-base",
use_gpu=True,
embed_title=True,
)
# Initialize RAG Generator
generator = RAGenerator(
model_name_or_path="facebook/rag-token-nq",
use_gpu=True,
top_k=1,
max_length=200,
min_length=2,
embed_title=True,
num_beams=2,
)
# -
# We write documents to the DocumentStore, first by deleting any remaining documents then calling `write_documents()`.
# The `update_embeddings()` method uses the retriever to create an embedding for each document.
#
# + pycharm={"name": "#%%\n"}
# Delete existing documents in documents store
document_store.delete_documents()
# Write documents to document store
document_store.write_documents(documents)
# Add documents embeddings to index
document_store.update_embeddings(
retriever=retriever
)
# -
# Here are our questions:
# + pycharm={"name": "#%%\n"}
QUESTIONS = [
"who got the first nobel prize in physics",
"when is the next deadpool movie being released",
"which mode is used for short wave broadcast service",
"who is the owner of reading football club",
"when is the next scandal episode coming out",
"when is the last time the philadelphia won the superbowl",
"what is the most current adobe flash player version",
"how many episodes are there in dragon ball z",
"what is the first step in the evolution of the eye",
"where is gall bladder situated in human body",
"what is the main mineral in lithium batteries",
"who is the president of usa right now",
"where do the greasers live in the outsiders",
"panda is a national animal of which country",
"what is the name of manchester united stadium",
]
# -
# Now let's run our system!
# The retriever will pick out a small subset of documents that it finds relevant.
# These are used to condition the generator as it generates the answer.
# What it should return then are novel text spans that form and answer to your question!
# + pycharm={"name": "#%%\n"}
# Now generate an answer for each question
for question in QUESTIONS:
# Retrieve related documents from retriever
retriever_results = retriever.retrieve(
query=question
)
# Now generate answer from question and retrieved documents
predicted_result = generator.predict(
query=question,
documents=retriever_results,
top_k=1
)
# Print you answer
answers = predicted_result["answers"]
print(f'Generated answer is \'{answers[0]["answer"]}\' for the question = \'{question}\'')
# + pycharm={"name": "#%%\n"}
# Or alternatively use the Pipeline class
from haystack.pipeline import GenerativeQAPipeline
pipe = GenerativeQAPipeline(generator=generator, retriever=retriever)
for question in QUESTIONS:
res = pipe.run(query=question, top_k_generator=1, top_k_retriever=5)
print(res)
# -
# ## About us
#
# This [Haystack](https://github.com/deepset-ai/haystack/) notebook was made with love by [deepset](https://deepset.ai/) in Berlin, Germany
#
# We bring NLP to the industry via open source!
# Our focus: Industry specific language models & large scale QA systems.
#
# Some of our other work:
# - [German BERT](https://deepset.ai/german-bert)
# - [GermanQuAD and GermanDPR](https://deepset.ai/germanquad)
# - [FARM](https://github.com/deepset-ai/FARM)
#
# Get in touch:
# [Twitter](https://twitter.com/deepset_ai) | [LinkedIn](https://www.linkedin.com/company/deepset-ai/) | [Slack](https://haystack.deepset.ai/community/join) | [GitHub Discussions](https://github.com/deepset-ai/haystack/discussions) | [Website](https://deepset.ai)
#
# By the way: [we're hiring!](https://apply.workable.com/deepset/)
| tutorials/Tutorial7_RAG_Generator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <div class="alert alert-block alert-warning">
# <b>Disclaimer:</b> The objective of this notebook is to introduce Project Jupyter and then give a short introduction about its functionalities.
# </div>
# # What is the Jupyter project?
#
# <img src="../../docs/images/jupyter-logo.svg" alt="JupyterLogo" style="width: 200px;"/>
#
# ## History
# Project *Jupyter* is a non-profit, open-source project, born out of the *IPython* Project in 2014 as it evolved to support interactive data science and scientific computing across many programming languages.
#
# *Jupyter* is an acronym for **Ju**lia, **Pyt**hon, and **R**. These three languages were the first target languages of the *Jupyter* application, but nowadays, the notebook technology also support many other languages (C, C#, C++, Scala, Ruby, Perl, Javascript, Fortran,...).
#
# *Jupyter* allows users to create and share documents that contain live code, equations, visualizations and narrative text.
#
# **References**
# - https://jupyter.org
# - https://www.nature.com/articles/d41586-018-07196-1
# - https://opencredo.com/blogs/writing-a-custom-jupyterhub-spawner/
#
# ## What are Jupyter hub, lab notebook app and notebook document?
#
# ### JupyterHub
# From https://jupyter.org/hub
# > *JupyterHub* brings the power of notebooks to groups of users. It gives them access to computational environments and resources without burdening with installation and maintenance tasks. Users can get their work done in their own workspaces on shared resources which can be managed efficiently by system administrators. *JupyterHub* runs in the cloud or on users' own hardware, and makes it possible to serve a pre-configured data science environment to any user in the world. It is customizable and scalable, and is suitable for small and large teams, academic courses, and large-scale infrastructure.
#
# ### JupyterLab
# <div>
# <img src="../../docs/images/launcher_jupyterlab.png" alt="JupyterLabLauncher" style="width: 500px;"/>
# <div style="text-align:center"> <span style="font-style:italic; font-size:1em;"> JupterLab launcher </span></div>
# </div>
#
# *Jupyterlab* is the next-generation web interface for the project *Jupyter*. It offers all the building blocks of the *Jupyter Notebook*, with additional drag-and-drop functionality, file browsers, data viewers, text editors and a command console. Users can arrange multiple documents and activities side by side in the work area using tabs and splitters. Documents and activities integrate with each other, enabling new workflows for interactive computing. *JupyterLab* will eventually replace the classic *Jupyter Notebook App*.
#
# <div>
# <img src="../../docs/images/running_jupyterlab.png" alt="JupyterLab" style="width: 500px;"/>
# <div style="text-align:center"> <span style="font-style:italic; font-size:1em;"> Example of layout of JupterLab showing a terminal, a running jupyter notebook, a rendered markdown file and the file browser.</span></div>
# </div>
#
# **References:**
# - https://github.com/jupyterlab/jupyterlab
#
# - https://jupyterlab.readthedocs.io/en/latest/
#
# ### Jupyter Notebook App
# A *Jupyter Notebook App* is a web-based interactive development environment for creating notebook documents.
#
# ### Jupyter Notebook document
# A *Jupyter Notebook* document consists of a list of input and output ordered cells that can contain computer code, [Markdown text](https://en.wikipedia.org/wiki/Markdown), mathematical expressions, plots, figures, links,... It is a [JavaScript Object Notation](https://en.wikipedia.org/wiki/JSON) (JSON) document. Its standard extension is `.ipynb`; but it can be converted to different formats such as `html`, slides, `pdf` or `Python` script.
#
# **Reference:** https://jupyter-notebook.readthedocs.io/en/stable/
#
# #### Kernels
# *Jupyter* supports over 40 programming languages including Python, Julia, R, C.
#
# *Notebook kernels* are processes that run interactive code in a particular programming language and return output to the user.
#
# Opening a *Jupyter Notebook* document automatically launches the associated kernel. When executing the notebook either cell-by-cell or in its entirety, this kernel performs the computation and produces the results, which are displayed in the output cells of the *Jupyter Notebook* document.
#
# [TryJupyter](https://jupyter.org/try) to try in your browser without any required installation.
#
# **Reference:** https://jupyter-notebook-beginner-guide.readthedocs.io/en/latest/what_is_jupyter.html
#
# # Short introduction about Jupyter notebook
#
# ## Jupyter dashboard
# The dashboard displays the folders and notebooks in the current directory and allows users to launch or create *Jupyter* notebooks in any subfolders.
#
# <div>
# <img src="../../docs/images/jupyter-dashboard.png" alt="JupyterDashboard" style="width: 500px;"/>
# <div style="text-align:center"> <span style="font-style:italic; font-size:1em;"> Example of Jupyter dashboard.</span></div>
# </div>
#
# ### Exercises
# #### Exercise 1
# - Create a new folder by clicking on `New` (upper right corner)
#
# <img src="../../docs/images/jupyter-dashboard-new.png" alt="CreateNewFromDashboard" style="width: 500px;"/>
#
# - Rename the folder 'MyNewFolder'.
# To do so, click in the checkbox next to the directory name and choose `Rename`. A new window will open in which you can type the new name of the directory. You can use this same process to rename any folder.
# - Remove this newly created folder.
# In this case, make sure that the checkbox next to your folder is still ticked, click on the 'Trash bin' button and confirm the deletion.
#
# #### Exercise 2
# - Create a new notebook by clicking the `New` dropdown (upper right corner) and select `Python 3` as kernel.
#
# <img src="../../docs/images/jupyter-dashboard-new.png" alt="CreateNewFromDashboard" style="width: 500px;"/>
#
# - You should get a new tab in your browser, which looks like the figure below
#
# <img src="../../docs/images/jupyter-notebook-green-code-cell.png" alt="NotebookGreenCodeCell" style="width: 500px;"/>
#
# - Rename it 'MyNewNotebook' by clicking on the 'Untitled' at the top of the page.
#
# <img src="../../docs/images/rename_jupyter_notebook.png" alt="RenameNotebook" style="width: 500px;"/>
#
# - Go back to the dashboard (clicking on the `Home` tab, *i.e.* the previously open tab in your browser) and check that your notebook is in the displayed list with a green icon (showing that it is running).
#
# - To close the notebook, check that the checkbox is ticked and click on `Shutdown` in the top left set of icons as show below
#
# <div>
# <img src="../../docs/images/jupyter-dashboard-shutdown.png" alt="ShutdownNotebook" style="width: 500px;"/>
# <div style="text-align:center"> <span style="font-style:italic; font-size:1em;"> How to close a notebook: select the notebook you want to close by ticking the checkbox next to it and then click on the `Shutdown` button.</span></div>
# </div>
#
# #### Note
# The material used during this half-day training is composed of different notebooks stored in the `python-ikon-course/notebooks` folder.
#
# **References**
# - [Python for Data Science Cheat Sheet - Jupyter notebook](https://datacamp-community-prod.s3.amazonaws.com/48093c40-5303-45f4-bbf9-0c96c0133c40)
#
# - [Jupyter notebook cheat sheet from edureka!](https://www.edureka.co/blog/wp-content/uploads/2018/10/Jupyter_Notebook_CheatSheet_Edureka.pdf)
# ## Jupyter notebook
# ### Menu bar and toolbar
# <div>
# <img src="../../docs/images/jupyter_notebook_toolbar_annotations.png" alt="NotebookToolbarAnnotations" style="width: 600px;"/>
# <div style="text-align:center"> <span style="font-style:italic; font-size:1em;"> Menubar, toolbar and additional buttons and dropdowns related to installed notebook extensions "nbextensions", such as spellchecker,opening the command panel... </span></div>
# </div>
#
# The menu bar presents different options that may be used to manipulate how the notebook functions. For example, from `Kernel`, you can interrupt, change or restart the kernel with the option of clearing the cells' output.
# The toolbar gives a quick access to the most-used operations by clicking on an icon. Among these operations, one can save, add or remove a cell, change between code cell and a markdown cell, or run a cell.
# ### Cells
# #### Types of cells
# The content of the notebook is a linear sequence of cells, where users can write code or text on mutiple lines. The content of each cell can be executed using `<shift>+<enter>` or by clicking on the `Play` button of the toolbar.
#
# #### Modal editor
# Depending on which mode the Notebook is in, typing on the keyboard can have different outcomes. There are two possible modes:
# - **Edit mode**
# The *Edit mode* is indicated by a green cell border and a prompt in the cell allowing you to type in.
#
# <img src="../../docs/images/jupyter-notebook-green-code-cell.png" alt="NotebookGreenCodeCell" style="width: 600px;"/>
#
# To enter the *Edit mode*, press `<enter>` or click on the cell you want to edit.
#
# - **Command mode**
# The *Command mode* is indicated by a grey cell border with a blue left margin
#
# <img src="../../docs/images/jupyter-notebook-blue-code-cell.png" alt="NotebookGreenCodeCell" style="width: 600px;"/>
#
# In the *Command mode*,
# - you can edit the notebook as a whole, but you cannot type into individual cells.
# - the keyboard is mapped to a set of shortcuts that let you perform notebook and cell actions efficiently. For example, if you are in command mode and you press `l`, you will display the line numbers in the current cell - no modifier is needed. The list of defined "Keyboard shortcuts" can be accessed by clicking on the `Help` dropdown of the menu bar.
#
# Note that trying to type into a cell when in the *Command mode* might result in unexpected things to happen.
#
# To enter the *Command mode*, press `<esc>` or click outside a cell’s editor area. Pressing `<control>+<enter>` will run the current cell and also enter the *Command mode*.
#
# #### Running the cells
# To run a cell, use `<shift>+<enter>` or press the `Play` button in the toolbar.
# The output depends on the cell type:
# - a markdown cell is rendered
# - for a code cell, its content is sent to the associated kernel and the result of the computation is displayed below the input cell, which is marked with '[\*]' indicating that the kernel is running and then with '[$n$]', $n$ being a number indicating that this cell is the $n^{th}$ code cell to be run as shown below:
#
# <img src="../../docs/images/jupyter-notebook-executed-cells.png" alt="NotebookExecutedCells" style="width: 600px;"/>
#
# **Exercise**
# - Try typing something like `print("Hello World")` into the cell below (it should be a *Code* cell).
# - To run the code in the cell and see the output, click the Run button (`Play` icon) on the toolbar, or press `<shift>+<enter>`:
# -- YOUR CODE BELOW --
# #### Saving your Notebook
# Once you start editing your Notebook, it is best practice to save it regularly. Pressing `<ctrl> + S` or `<command> + S` will save your notebook by calling the "Save and Checkpoint" command (from the `File` dropdown in the menu bar).
#
# **Note:** What is a checkpoint?
# From https://www.codecademy.com/articles/how-to-use-jupyter-notebooks
# > Every time you create a new notebook, a checkpoint file is created as well as your notebook file; it will be located within a hidden subdirectory of your save location called .ipynb_checkpoints and is also a .ipynb file. By default, *Jupyter* will autosave your notebook every 120 seconds to this checkpoint file without altering your primary notebook file. When you "Save and Checkpoint," both the notebook and checkpoint files are updated. Hence, the checkpoint enables you to recover your unsaved work in the event of an unexpected issue. You can revert to the checkpoint from the menu via "File > Revert to Checkpoint."
#
# #### Closing your Notebook
# As mentioned above, when a notebook is opened, its kernel is automatically started. Closing the notebook browser tab will not shut down the kernel.
#
# To shut down the notebook and its kernel, from the menu bar of the notebook, click on `File` and select `Close and Halt`. Another option is, from the Notebook Dashboard, to select the *Running* notebook you want to close and then click on the `Shutdown` button.
#
# #### Downloading your Notebook
# If you want to run the Notebook you created on your own system or if you want to convert it to another format (`Python` script, `.pdf`...), it can be downloaded and converted using, from the `File` dropdown, `Downloads as`.
# ### Short tutorial about Markdown
# Below are a few examples of what can be done in Markdown.
#
# **References**
# - https://www.markdowntutorial.com/
# - https://guides.github.com/features/mastering-markdown/
#
# #### Fonts
# You can write text in *italic*, **bold**.
#
# Here is a blockquote:
#
# > This is a quote.
# It can be on several lines.
#
# ##### LaTeX equations
# LaTeX equations can be written in Markdown cells, such as
#
# $$\psi_f(s)=\sum_{n=0}^{\infty}\frac{1}{n^s} \qquad \beta(t)=\prod_p\frac{1}{\alpha - p^{-t}} \qquad I=\int_0^{\pi}\sin^2(\omega t)$$
#
# $$\begin{equation*}
# \mathbf{V}_1 \times \mathbf{V}_2 = \begin{vmatrix}
# \mathbf{i} & \mathbf{j} & \mathbf{k} \\
# \frac{\partial X}{\partial u} & \frac{\partial Y}{\partial u} & 0 \\
# \frac{\partial X}{\partial v} & \frac{\partial Y}{\partial v} & 0
# \end{vmatrix}
# \end{equation*} $$
#
# *Maxwell's equations*
#
# $$\begin{align}
# \nabla \times \vec{\mathbf{B}} -\, \frac1c\, \frac{\partial\vec{\mathbf{E}}}{\partial t} & = \frac{4\pi}{c}\vec{\mathbf{j}} \\ \nabla \cdot \vec{\mathbf{E}} & = 4 \pi \rho \\
# \nabla \times \vec{\mathbf{E}}\, +\, \frac1c\, \frac{\partial\vec{\mathbf{B}}}{\partial t} & = \vec{\mathbf{0}} \\
# \nabla \cdot \vec{\mathbf{B}} & = 0
# \end{align} $$
#
# ##### Embedded code
# Code can also be written in markdown using specific languages' highlighting. For example, using the Python syntax:
#
# ```python
# print('bar')
# ```
#
# ##### Colors
# We can write text in different colors, for example, <font color='red'>red</font>, <font color='blue'>blue</font>...
#
# #### Layout
# You can add horizontal rules:
#
# ---
#
# ##### Headings
# Examples of the hierarchy of headings:
#
# ---
#
# # Header 1
# ## Header 2
# ### Header 3
# #### Header 4 ####
# ##### Header 5 #####
# ###### Header 6 ######
#
# ---
#
# #### Lists
# You can build nested itemized or enumerated lists:
#
# * First element of list
# - Sublist item1
# - Subsublist
# - Sublist item2
# - Subsublist item1
# - Subsublist item2
# * Second element of list
# - Sublist
# * Third element of list
# - Sublist
#
# Now another list:
#
# 1. First element of enumerated list
# 1. Example of embedded enumerated list
# 2. Second iem of this enumerated sublist
# 2. Second element of enumerated list
# 3. Third element of enumerated list
#
# #### Links
# You can also add links:
#
# [Jupyter website](https://jupyter.org)
#
# #### Tables
# You can create tables very easily
#
# | Header | in first | row |
# | :--- | :----: | ---: |
# | left | centered | right |
# | aligned | text | aligned |
#
# Not even aligning the borders and you can modify the font style with stars double stars or quotes
#
# Markdown | Less | Pretty
# --- | --- | ---
# *Still* | `renders` | **nicely**
# 1 | 2 | 3
#
# #### Magic commands
# Magic commands are enhancements that `IPython` offers compared to the standard Python shell. These magic commands act as convenient functions where `Python` syntax is not the most natural one. In other words, you can run code in different languages in different cells within your notebook.
#
# ##### Types of magic commands
# - **Line magics**
# They use input on the same line. They start with **%** character.
# - **Cell magics**
# They start with **%%** characters. They can operate on multiline inputs.
#
# *Examples:*
# - `%lsmagic`: list all magic commands
# - `%history`: display previous inputs at once
# - `%run`: run external code
# For example
# ```python
# # the following command will execute and show the output from all code cells of the specified notebook
# # %run ./plotting-with-matplotlib.ipynb
# ```
# - `%who`: list all variables of global scope.
# ```python
# # %who str
# ```
#
# - `%matplotline inline` to show matplotlib plots inline the notebook
# - `%%writefile` magic saves the contents of that cell to an external file.
# - `%pycat` does the opposite, and shows you (in a popup) the syntax highlighted contents of an external file.
#
# *Example:* using the following two cells, you start with writing the content of the first cell in a `Python` script, `simple_python_script.py`, and you execute it in the second cell.
# +
# %%writefile simple_python_script.py
def print_info():
import datetime
now = datetime.datetime.now()
print("This is a test of magic commands run on {}". format(now))
print_info()
# -
# %run simple_python_script.py
# ##### Executing shell commands
# Another useful modifier is the exclamation mark **!**.
# It allows to execute shell commands from the notebook. Below are a few examples:
#
# To check the *Jupyter notebooks* in the same folder as the running document:
# ``` python
# # !ls *.ipynb
# ```
#
# To check packages, you can use
# ``` python
# # !pip install numpy
# # !pip list | grep pandas
# ```
#
# **Note:** Difference between **!** and **%**
#
# From https://stackoverflow.com/questions/45784499/difference-between-and-in-jupyter-notebooks
# > **!** calls out to a shell (in a new process), while **%** affects the notebook itself. Many **%** commands have no shell equivalent.
# `!cd foo`, by itself, has no lasting effect, since the process with the changed directory immediayely terminates. `%cd foo` changes the current directory of the notebook process, which is a lasting effect.
# **References:**
# - [Built-in magic commands](https://ipython.readthedocs.io/en/stable/interactive/magics.html)
#
# - [demonstration of how to use Python, Julia, Fortran and R cooperatively to analyze data, in the same process](https://nbviewer.jupyter.org/gist/fperez/5b49246af4e340c37549265a90894ce6/polyglot-ds.ipynb)
#
# ##### Examples using HTML, bash and perl
# Code cells can be run using a different kernel, for example, `Html`, `LaTeX`, `bash`...
# + language="bash"
# echo $HOME
# -
# %%HTML
<H4>Text entered in code cell, rendered as HTML</H4>
# + language="perl"
# @months = ("Jan", "Feb", "Mar");
# print($months[1])
# -
# # Inline help
help([])
# The `help` function output the documentation for an object.
help(dict())
# Another way to get help with Jupyter and ipython os to use the '?' at the end of an object.
# +
# dict?
# -
# Typing the name of an object on the last line in a cell output information about the object
a = dict(a=3, b=5)
a
# but it has to be on the last line. The following will not create an output
b = range(10)
b
c = 3
# unless we use `print`
b = range(10)
print(b)
c = 3
# The wonderful **`TAB`** key. `TAB` invokes auto-completion.
#
# Try type "l" immediately followed by TAB in the cell below and see what happens
# -- YOUR CODE BELOW --
# It list the ways that the "l" can be completed. If there is only one option, it will just complete your input for you. If you type "li" followed by `TAB` it will list those starting with "li" and so on. Note that you can select from the drop down menu that appears.
#
# Try to type 'from numpy import ' followed by `TAB`.
# -- YOUR CODE BELOW --
# ## Customizing my notebook
# There is a number of ways to add features and customize your jupyter notebooks. For example, with [Jupyter Notebook extensions](https://towardsdatascience.com/jupyter-notebook-extensions-517fa69d2231), one can include:
# - table of contents
# - add a spellchecker
# - scratchpad
# - freeze (read-only cells)
#
# ## Examples in scientific community
# - [neutron imaging community](https://github.com/neutrons/IPythonNotebookTutorial)
# - [icet tutorials](https://gitlab.com/materials-modeling/icet-tutorials): scientific software for constructing atomic scale models of materials
# - [Atomic Simulation Environment](https://github.com/ajjackson/ase-tutorial-symmetry)
# - [qef](https://github.com/jmborr/qef/tree/master/notebooks): lmfit models for fitting quasielastic neutron scattering data
# - [QENS models](https://github.com/QENSlibrary/QENSmodels/tree/master/examples)
# - [McStasScript](https://github.com/PaNOSC-ViNYL/McStasScript/tree/master/examples)
# - [scipp](https://scipp.readthedocs.io/en/latest/visualization/plotting-overview.html): multi-dimensional data arrays with labeled dimensions
# - Google group [Jupyter at Research Facilities](https://groups.google.com/forum/#!forum/jupyter-research-facilities)
# # Additional materials
# ## Uploading my files to JupyterHub
# If you want to use your own datafiles, click on the `Upload` button as shown in the figure below:
#
# <div>
# <img src="../../docs/images/upload-my-file.png" alt="UploadMyFile" style="width: 600px;"/>
# <div style="text-align:center"> <span style="font-style:italic; font-size:1em;"> Screenshot of Jupyter dashboard highlighting the `Upload` button to load user's own file(s) in the dashboard. </span></div>
# </div>
#
# ## Installation (not required for this workshop)
# Although it is possible to run each notebook of this repository using the *JupyterHub* (meaning that they are run on an external compute resources), you may want to try writing your own *Jupyter* notebooks and *Python* code.
#
# If it is the case, you will have to install *Jupyter* notebooks locally (*e.g.* on your own laptop or desktop). This can be achieved by downloading and installing Anaconda. This is available for Windows, macOS and Linux [here](https://www.anaconda.com/distribution/#download-section).
#
# Once installed the *Jupyter* notebooks can be opened, like how it is done during this training session. This creates a new window in your web browser: this is the *Jupyter* file navigator. Using this you can navigate to a specific folder, open an existing notebook or create a new one and save it on your system.
#
# ## ipywidgets
# * [basic widget tutorial](../graphics_and_widgets/ipywidgets.ipynb)
# * [complete collection of jupyter widgets tutorials](https://github.com/jupyter-widgets/tutorial/tree/master/notebooks)
# * [list of widgets](https://ipywidgets.readthedocs.io/en/latest/examples/Widget%20List.html)
#
# ## What to do when things go wrong?
#
# https://jupyter-notebook.readthedocs.io/en/stable/troubleshooting.html
#
# ## Trusting notebooks
# https://jupyter-notebook.readthedocs.io/en/latest/security.html#notebook-security
| notebooks/1_jupyter_basics/jupyter-notebook-intro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# %matplotlib inline
import astropy.io.fits as pyfits
import PythonPhot as pp
import numpy as np
# read in some sample data from SDSS
gimage,rimage = \
pyfits.getdata('Pal5-g-SDSS.fits'),pyfits.getdata('Pal5-r-SDSS.fits')
# convert units back into counts for poisson statistics
ggain = 1/pyfits.getval('Pal5-g-SDSS.fits','NMGY')
rgain = 1/pyfits.getval('Pal5-r-SDSS.fits','NMGY')
gimage *= ggain
rimage *= rgain
# +
# now let's get started with the find.py algorithm
# first the sky background - let's just call the readnoise 0
rskymod, rskysig, rskyskw = pp.mmm.mmm(rimage,readnoise=0,minsky=200)
gskymod, gskysig, gskyskw = pp.mmm.mmm(rimage,readnoise=0,minsky=200)
# This algorithm is not the best, helps to tune hmin a bit
rhmin = rskysig*5
ghmin = gskysig*5
# take a stab at the image FWHM
fwhm = 5
gxstar,gystar,flux,sharp,round = \
pp.find.find(gimage,ghmin,fwhm,[-1.0,1.0],[0.2,1.0])
rxstar,rystar,flux,sharp,round = \
pp.find.find(rimage,rhmin,fwhm,[-1.0,1.0],[0.2,1.0])
# Now let's see how we did
import pylab as plt
plt.ion()
plt.imshow(gimage,vmin=0,vmax=10*gskysig,cmap='Greys_r')
plt.plot(gxstar,gystar,'o',ms=5,mfc='none',lw=2,mec='r')
plt.xlim([400,1350]); plt.ylim([450,1250])
# not bad!
# +
# let's do some aperture photometry and
# see how the color mag diagrams look
# first, draw a box around just the cluster stars
ipal_g = ((gxstar > 400) & (gxstar < 1350) &
(gystar > 450) & (gystar < 1250))
ipal_r = ((rxstar > 400) & (rxstar < 1350) &
(rystar > 450) & (rystar < 1250))
print('detected %i cluster stars'%len(np.where(ipal_g == True)[0]))
# coords are close in both images, but not exactly
# let's centroid them
gxstar,gystar = pp.cntrd.cntrd(gimage,gxstar,gystar,fwhm,verbose=False)
rxstar,rystar = pp.cntrd.cntrd(rimage,rxstar,rystar,fwhm,verbose=False)
gmag,gmagerr,gflux,gfluxerr,gsky,gskyerr,badflag,outstr = \
pp.aper.aper(gimage/ggain,gxstar[ipal_g],gystar[ipal_g],phpadu=ggain,apr=5,zeropoint=25.11,
skyrad=[3*fwhm,5*fwhm],badpix=[-12000,60000],exact=True,setskyval=0)
rmag,rmagerr,rflux,rfluxerr,rsky,rskyerr,badflag,outstr = \
pp.aper.aper(rimage/rgain,rxstar[ipal_r],rystar[ipal_r],phpadu=rgain,apr=5,zeropoint=24.80,
skyrad=[3*fwhm,5*fwhm],badpix=[-12000,60000],exact=True,setskyval=0)
# match up g and r coords
gmag_pal,rmag_pal = np.array([]),np.array([])
for gx,gy,i in zip(gxstar[ipal_g],gystar[ipal_g],range(len(gmag))):
sep = np.sqrt((gx-rxstar[ipal_r])**2.+ (gy-rystar[ipal_r])**2.)
imatch = np.where((sep == min(sep)))
if len(imatch[0]):
gmag_pal,rmag_pal = np.append(gmag_pal,gmag[i]),\
np.append(rmag_pal,rmag[imatch][0])
plt.plot(gmag_pal-rmag_pal,rmag_pal,'.',color='k')
plt.ylim([25,17])
plt.xlim([0.3,1.5])
plt.ylabel('$g$')
plt.xlabel('$g-r$')
# Not the prettiest color mag diagram - I really should
# be matching up WCS coordinates instead of pixel coords
# as these images are not quite WCS-aligned.
# But, it works!
# -
# Let's just look at the brightest stars
# for the next steps, pkfit is a little slow
ibright = np.argsort(gmag)[0:50]
gxstar,gystar,gflux = gxstar[ipal_g][ibright],gystar[ipal_g][ibright],gflux[ibright]
| examples/PythonPhot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Br3ndo0n/OOP-58002/blob/main/Prelim_Exam.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="ZYef5xoRFgRJ"
# #PRELIM EXAM
# + colab={"base_uri": "https://localhost:8080/"} id="73ZoGNEJGYm5" outputId="06a6f76e-d569-418f-b023-a3c868119928"
class Person:
def __init__(self,student,number,age,school,course):
self.student = student
self.number = number
self.age = age
self.school = school
self.course = course
def myFunction(self):
print("I am ",self.student,"my age is",self.age, "my student number is",self.number, "studying in", self.school, "and I am taking", self.course)
p1= Person("Campaña, <NAME>.", 202113816, 18, "Adamson University", "Computer Engineering")
p1.myFunction()
| Prelim_Exam.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# # !wget https://storage.googleapis.com/xlnet/released_models/cased_L-12_H-768_A-12.zip -O xlnet.zip
# # !unzip xlnet.zip
# -
import os
os.environ['CUDA_VISIBLE_DEVICES'] = ''
# +
import sentencepiece as spm
from prepro_utils import preprocess_text, encode_ids
sp_model = spm.SentencePieceProcessor()
sp_model.Load('xlnet_cased_L-12_H-768_A-12/spiece.model')
def tokenize_fn(text):
text = preprocess_text(text, lower= False)
return encode_ids(sp_model, text)
# +
SEG_ID_A = 0
SEG_ID_B = 1
SEG_ID_CLS = 2
SEG_ID_SEP = 3
SEG_ID_PAD = 4
special_symbols = {
"<unk>" : 0,
"<s>" : 1,
"</s>" : 2,
"<cls>" : 3,
"<sep>" : 4,
"<pad>" : 5,
"<mask>" : 6,
"<eod>" : 7,
"<eop>" : 8,
}
VOCAB_SIZE = 32000
UNK_ID = special_symbols["<unk>"]
CLS_ID = special_symbols["<cls>"]
SEP_ID = special_symbols["<sep>"]
MASK_ID = special_symbols["<mask>"]
EOD_ID = special_symbols["<eod>"]
# -
text = 'A politician is a person active in party politics, or a person holding or seeking office in government. Politicians propose, support and create laws or policies that govern the land and, by extension, its people'
text
aug_percent = 0.8
splitted = text.split()
size = len(splitted)
cnt = int(aug_percent * size)
cnt
import json
with open('en.json') as fopen:
stopwords = json.load(fopen)
# +
import random
import string
results = []
samples = random.sample([i for i in range(size)], cnt)
for token_idx, token in enumerate(samples):
if splitted[token] in string.punctuation:
continue
if splitted[token] in stopwords:
continue
results.append(token)
results
# +
import numpy as np
def tokenizer(string, mask_id):
string = string.split()
ids = []
for no, word in enumerate(string):
if no == mask_id:
ids.append(MASK_ID)
ids.extend(tokenize_fn(word))
mask_ind = ids.index(MASK_ID)
segment_id = [SEG_ID_A] * len(ids)
input_mask = [0] * len(ids)
perm_masks = np.zeros((1, len(ids)))
perm_masks[0, mask_ind] = 1.0
target_mappings = np.zeros((1, len(ids)))
target_mappings[0, mask_ind] = 1.0
return ids, segment_id, input_mask, mask_ind, perm_masks, target_mappings
# +
import xlnet
import tensorflow as tf
import model_utils
kwargs = dict(
is_training=True,
use_tpu=False,
use_bfloat16=False,
dropout=0.0,
dropatt=0.0,
init='normal',
init_range=0.1,
init_std=0.05,
clamp_len=-1)
xlnet_parameters = xlnet.RunConfig(**kwargs)
xlnet_config = xlnet.XLNetConfig(json_path='xlnet_cased_L-12_H-768_A-12/xlnet_config.json')
# +
def top_k_logits(logits, k):
if k == 0:
return logits
def _top_k():
values, _ = tf.nn.top_k(logits, k=k)
min_values = values[:, -1, tf.newaxis]
return tf.where(
logits < min_values,
tf.ones_like(logits, dtype=logits.dtype) * -1e10,
logits,
)
return tf.cond(
tf.equal(k, 0),
lambda: logits,
lambda: _top_k(),
)
def top_p_logits(logits, p):
with tf.variable_scope('top_p_logits'):
logits_sort = tf.sort(logits, direction='DESCENDING')
probs_sort = tf.nn.softmax(logits_sort)
probs_sums = tf.cumsum(probs_sort, axis=1, exclusive=True)
logits_masked = tf.where(probs_sums < p, logits_sort, tf.ones_like(
logits_sort)*1000) # [batchsize, vocab]
min_logits = tf.reduce_min(logits_masked, axis=1, keepdims=True) # [batchsize, 1]
return tf.where(
logits < min_logits,
tf.ones_like(logits, dtype=logits.dtype) * -1e10,
logits,
)
class Model:
def __init__(
self,
):
self.X = tf.placeholder(tf.int32, [None, None])
self.segment_ids = tf.placeholder(tf.int32, [None, None])
self.input_masks = tf.placeholder(tf.float32, [None, None])
self.perm_masks = tf.placeholder(tf.float32, [None, None, None])
self.target_mappings = tf.placeholder(tf.float32, [None, None, None])
self.top_p = tf.placeholder(tf.float32, None)
self.top_k = tf.placeholder(tf.int32, None)
self.k = tf.placeholder(tf.int32, None)
self.temperature = tf.placeholder(tf.float32, None)
self.indices = tf.placeholder(tf.int32, [None, None])
xlnet_model = xlnet.XLNetModel(
xlnet_config=xlnet_config,
run_config=xlnet_parameters,
input_ids=self.X,
seg_ids=self.segment_ids,
input_mask=self.input_masks,
perm_mask = self.perm_masks,
target_mapping = self.target_mappings
)
output = xlnet_model.get_sequence_output()
self.output = output
lookup_table = xlnet_model.get_embedding_table()
initializer = xlnet_model.get_initializer()
with tf.variable_scope('model', reuse = tf.AUTO_REUSE):
with tf.variable_scope('lm_loss'):
softmax_w = lookup_table
softmax_b = tf.get_variable(
'bias',
[xlnet_config.n_token],
dtype = output.dtype,
initializer = tf.zeros_initializer(),
)
logits = tf.einsum('ibd,nd->ibn', output, softmax_w) + softmax_b
self.logits = logits
logits = tf.gather_nd(self.logits, self.indices)
logits = logits / self.temperature
def necleus():
return top_p_logits(logits, self.top_p)
def select_k():
return top_k_logits(logits, self.top_k)
logits = tf.cond(self.top_p > 0, necleus, select_k)
self.samples = tf.multinomial(
logits, num_samples=self.k, output_dtype=tf.int32)
# +
tf.reset_default_graph()
sess = tf.InteractiveSession()
model = Model()
sess.run(tf.global_variables_initializer())
# +
import collections
import re
def get_assignment_map_from_checkpoint(tvars, init_checkpoint):
"""Compute the union of the current variables and checkpoint variables."""
assignment_map = {}
initialized_variable_names = {}
name_to_variable = collections.OrderedDict()
for var in tvars:
name = var.name
m = re.match('^(.*):\\d+$', name)
if m is not None:
name = m.group(1)
name_to_variable[name] = var
init_vars = tf.train.list_variables(init_checkpoint)
assignment_map = collections.OrderedDict()
for x in init_vars:
(name, var) = (x[0], x[1])
if name not in name_to_variable:
continue
assignment_map[name] = name_to_variable[name]
initialized_variable_names[name] = 1
initialized_variable_names[name + ':0'] = 1
return (assignment_map, initialized_variable_names)
# -
tvars = tf.trainable_variables()
checkpoint = 'xlnet_cased_L-12_H-768_A-12/xlnet_model.ckpt'
assignment_map, initialized_variable_names = get_assignment_map_from_checkpoint(tvars,
checkpoint)
saver = tf.train.Saver(var_list = assignment_map)
saver.restore(sess, checkpoint)
import numpy as np
from tensorflow.keras.preprocessing.sequence import pad_sequences
tokenized = [tokenizer(text, result) for result in results]
a = list(zip(*tokenized))
len(a)
# +
# ids, segment_id, input_mask, mask_ind, perm_masks, target_mappings
batch_x = pad_sequences(a[0],padding='post')
batch_segment = pad_sequences(a[1],padding='post', value = SEG_ID_PAD)
batch_mask = pad_sequences(a[2],padding='post', value = 1)
perm_masks = pad_sequences(a[4],padding='post')
target_mappings = pad_sequences(a[5],padding='post')
# -
indices = a[3]
batch_indices = np.array([np.arange(len(indices)), indices]).T
batch_indices.shape
batch_mask.shape
# +
# self.segment_ids = tf.placeholder(tf.int32, [None, None])
# self.input_masks = tf.placeholder(tf.float32, [None, None])
# self.perm_masks = tf.placeholder(tf.float32, [None, None, None])
# self.target_mappings = tf.placeholder(tf.float32, [None, None, None])
samples = sess.run(model.samples, feed_dict = {model.X: batch_x,
model.input_masks: batch_mask,
model.segment_ids: batch_segment,
model.perm_masks: perm_masks,
model.target_mappings: target_mappings,
model.top_p: 0.8,
model.top_k: 100,
model.temperature: 0.8,
model.indices: batch_indices,
model.k: 5})
# -
def convert_ids_to_tokens(ids):
return [sp_model.IdToPiece(i) for i in ids]
for i in range(samples.shape[1]):
print('SAMPLE %d'%(i))
sample_i = samples[:, i]
samples_tokens = convert_ids_to_tokens(samples[:, i].tolist())
new_splitted = splitted[:]
for no, index in enumerate(results):
new_splitted[index] = samples_tokens[no]
new = ' '.join(new_splitted)
print('BEFORE:', text)
print('AFTER:', new)
print()
| text-augmentation/8.xlnet-augmentation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (qnlp-ws)
# language: python
# name: qnlp-ws
# ---
# # Dimensionality reduction
from urllib.request import urlretrieve
import urllib.request
import bz2
import os
RETRIEVE_DATA = False
if not os.path.exists("data"):
os.mkdir("data")
if RETRIEVE_DATA:
rslt = urlretrieve("http://wikipedia2vec.s3.amazonaws.com/models/en/2018-04-20/enwiki_20180420_100d.pkl.bz2",
filename="data/enwiki_20180420_100d.pkl.bz2")
with bz2.open("data/enwiki_20180420_100d.pkl.bz2", "rb") as f:
data = f.read()
with open("data/enwiki_20180420_100d.pkl", "wb") as f:
f.write(data)
from wikipedia2vec import Wikipedia2Vec
# download and unpack from http://wikipedia2vec.s3.amazonaws.com/models/en/2018-04-20/enwiki_20180420_100d.pkl.bz2
wiki2vec = Wikipedia2Vec.load("data/enwiki_20180420_100d.pkl")
print(wiki2vec.get_word_vector("account"))
wiki2vec.most_similar(wiki2vec.get_word('nail'),10)
# +
from words import nouns
from pprint import pprint
import numpy as np
vecs = []
for noun in nouns:
if noun == "charge_n":
vecs.append(wiki2vec.get_word_vector("charge").tolist())
else:
vecs.append(wiki2vec.get_word_vector(noun).tolist())
vecs = np.array(vecs)
# -
vecs.shape
from sklearn.decomposition import PCA
pca = PCA(n_components=4)
vecs_new = pca.fit_transform(vecs)
print(pca.explained_variance_ratio_)
vecs_new
sum_of_rows = np.linalg.norm(vecs_new,axis=1)
normalized_array = vecs_new / sum_of_rows[:, np.newaxis]
new_noun_vectors = {}
for i, noun in enumerate(nouns):
new_noun_vectors[noun] = normalized_array[i].tolist()
import json
fiel_new_vectors = "new_noun_vectors.json"
with open(fiel_new_vectors, 'w') as fp:
json.dump(new_noun_vectors, fp)
| dimensionalty_reduction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Rapids Nightly [py37]
# language: python
# name: rapids-nightly
# ---
import dask_cudf
import cudf
import pandas as pd
output_file = 'timestamp.parquet'
output_dir = 'ts.parquet'
# Clean up before the demo here
# !rm {output_dir} -r
# !rm {output_file}
df = pd.DataFrame({'timestamp': pd.datetime(2019, 10, 21)}, index=[1])
df
# Use pandas to write a parquet file
df.to_parquet(output_file)
# Use cudf to read the parquet file. This works fine
df = cudf.read_parquet(output_file)
df
df = dask_cudf.read_parquet(output_file)
df.compute()
# Try and write a parquet file using the cudf dataframe. Get an error because the cudf implementation seems to
# be trying to write a directory with a partition inside of it. I would expect this to try and write a file called
# "timestamp.parquet"
df = cudf.read_parquet(output_file)
df.to_parquet(output_dir)
# Let's now make that directory so that cudf can write
# !mkdir {output_dir}
# Now if we save to the directory, it seems to be outputting correctly
df.to_parquet(output_dir)
# !ls {output_dir}
# Let's write to that directory again
df.to_parquet(output_dir)
# !ls {output_dir}
# +
# Seems as though we're writing a new parquet file out each time we call to_parquet. This is somewhat unintuitive.
# The to_parquet docs don't seem to suggest anything here either.
# df.to_parquet??
# -
# # Read in with cudf
# Seems that you can't read in that directory though
df = cudf.read_parquet(output_dir)
df
# Need to remind myself what output_dir is here
output_dir
# Let's try and make it obviously a directory
cudf.read_parquet(f'{output_dir}/')
# Ah that's the right syntax. Need to tell it to read all files in that dir
cudf.read_parquet(f'{output_dir}/*')
# Though if you do read it in with pandas, things seem to work just fine, but now we have two rows instead of the one
pd.read_parquet(output_dir)
# # Read in with dask_cudf
# Reading in with dask_cudf seems to work if we just pass it in a directory path. no need for the extra /* like with
# cudf, though in this case we get a TypeError because it seems to not understand how to handle a datetime64[ns] dtype
dask_cudf.read_parquet(output_dir)
# we get the same error no matter how we try and read these files in
dask_cudf.read_parquet(f'{output_dir}/')
# we get the same error no matter how we try and read these files in
dask_cudf.read_parquet(f'{output_dir}/*')
df = dask_cudf.read_parquet('tmp.parquet')
df.dtypes
| RAPIDS/notebooks/dask_cudf_timestamps.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.6.10 64-bit (''lab'': conda)'
# name: python3
# ---
# %matplotlib inline
import matplotlib.pyplot as plt
plt.style.use('seaborn-whitegrid')
import numpy as np
# +
x = np.linspace(0, 10, 30)
y = np.sin(x)
plt.figure(figsize = (30,15))
plt.plot(x, y, 'o', color = 'black');
# -
rng = np.random.RandomState(0)
plt.figure(figsize=(30,15))
for marker in ['o', '.', ',', 'x', '+', 'v', '^', '<', '>', 's', 'd']:
plt.plot(rng.rand(5), rng.rand(5), marker,
label="marker='{0}'".format(marker))
plt.legend(numpoints=1)
plt.xlim(0, 1.8);
plt.figure(figsize=(30, 15))
plt.plot(x, y, '-ok');
plt.figure(figsize=(30,15))
plt.plot(x, y, '-p', color='gray',
markersize=15, linewidth=4,
markeredgecolor='gray',
markeredgewidth=2)
plt.ylim(-1.2, 1.2);
plt.figure(figsize=(30,15))
plt.scatter(x, y, marker='o');
# +
rng = np.random.RandomState(0)
x = rng.randn(100)
y = rng.randn(100)
colors = rng.rand(100)
sizes = 1000 * rng.rand(100)
plt.figure(figsize=(30,15))
plt.scatter(x, y, c=colors, s=sizes, alpha=0.3, cmap='viridis')
plt.colorbar();
# +
from sklearn.datasets import load_iris
iris = load_iris()
features = iris.data.T
plt.figure(figsize=(30,15))
plt.scatter(features[0], features[1], alpha=0.2, s=100*features[3], c=iris.target, cmap='viridis')
plt.xlabel(iris.feature_names[0])
plt.ylabel(iris.feature_names[1]);
# -
| Matplotlib_plots/4-Simple_Scatter_Plots.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import collections
from collections import defaultdict
import sys
import json
import random
from jsmin import jsmin
from io import StringIO
import numpy as np
import copy
import importlib
from functools import partial
import math
import os
import argparse
script_n = 'pfs_connectivity_ratio_210306'
sys.path.insert(0, '/n/groups/htem/Segmentation/shared-nondev/cb2_segmentation/analysis_mf_grc')
import my_plot
importlib.reload(my_plot)
from my_plot import MyPlotData
max_dist = 150
threshold = 6
from weight_database import WeightDatabase
weightdb = WeightDatabase()
# weightdb.load_syn_db('/n/groups/htem/Segmentation/shared-nondev/cb2_segmentation/analysis_mf_grc/gen_db/pfs/gen_201224_setup01_syndb_threshold_10_coalesced.gz')
weightdb.load_syn_db('/n/groups/htem/Segmentation/shared-nondev/cb2_segmentation/analysis_mf_grc/gen_db/pfs/gen_210101_setup01_syndb_threshold_10_coalesced_filtered_10.gz')
weightdb.load_touch_db(
'/n/groups/htem/Segmentation/shared-nondev/cb2_segmentation/analysis_mf_grc/purkinje/db_pf_contacts_201224.gz',
max_dist=max_dist
)
# +
import compress_pickle
db_f = '/n/groups/htem/Segmentation/shared-nondev/cb2_segmentation/analysis_mf_grc/pfs_density_db_210306.gz'
pf_db = compress_pickle.load(db_f)
mpd = MyPlotData()
n_pfs = 0
for block in pf_db:
block_pfs = pf_db[block]['pfs']
if len(block_pfs) == 0:
continue
rates = []
for pf in block_pfs:
n_pfs += 1
rate = weightdb.calc_connection_rate(pf)
rates.append(rate)
avg = sum(rates) / len(rates)
print(f'{block}: {avg}')
mpd.add_data_point(
y=pf_db[block]['y_dist'],
rate=avg*100,
)
print(f'n_pfs = {n_pfs}')
# +
importlib.reload(my_plot); my_plot.my_cat_bar_plot(
mpd,
x="y",
y="rate",
# hue="type",
# hue_order=['All', 'Per PC', 'Per pf'],
# hue_order=['All', 'Per PC'],
ylim=[25, 75],
context='paper',
kind='box',
# font_scale=1.4,
# add_swarm=True,
# inner='box',
height=4,
# bw=.25,
# cut=0,
y_axis_label='Average pf-PC connectivity (%)',
x_axis_label='Molecular layer height (µm*1000)',
save_filename=f'{script_n}_max_dist_{max_dist}_threshold_{threshold}.svg',
show=True,
)
# +
importlib.reload(my_plot); my_plot.my_relplot(
mpd,
x="y",
y="rate",
ylim=[0, 100],
context='paper',
# kind='box',
height=2,
width=4,
y_axis_label='pf-PC conn.\nrate (%)',
x_axis_label='Molecular layer height (µm*1000)',
save_filename=f'{script_n}_max_dist_{max_dist}_threshold_{threshold}_line.svg',
show=True,
)
| analysis/pfs_pc_analysis/pfs_connectivity_ratio_210306.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 31009 - Final Project - RNNs Model
# ### Ada, Rohit, Dylan
import numpy as np
import pandas as pd
import re
import nltk
from nltk.corpus import stopwords
from nltk.stem.porter import PorterStemmer
from collections import Counter
import seaborn as sns
import matplotlib.pyplot as plt
from IPython.core.display import display, HTML
import string
import tensorflow as tf
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.preprocessing.sequence import pad_sequences
from tqdm import tqdm
from tensorflow.keras.models import Sequential, load_model
from tensorflow.keras.layers import LSTM, Dense, Dropout
from tensorflow.keras.layers import Embedding,SimpleRNN
from sklearn.model_selection import train_test_split
from keras import optimizers,initializers
##Load Data
train = pd.read_csv("Cleaned_Train.csv")
train_y = train.target
train.head()
# Tokenizer sequence and index words
tokenizer = Tokenizer()
tokenizer.fit_on_texts(train.text)
word_index = tokenizer.word_index
num_words = len(tokenizer.word_index)+1
print('Number of unique words:',len(word_index))
# +
training_sequences = tokenizer.texts_to_sequences(train.text)
# Ading padding at the front of text sequence
training_padded = pad_sequences(training_sequences,
maxlen=50,
padding='pre',
truncating='pre')
# Split data set for further training and validation
X_train, X_test, Y_train, Y_test = train_test_split(training_padded, train_y, test_size=.25,random_state=0)
# +
# Matching words with Glove embedding 6B.300D
embedding_dict={}
with open('glove.6B.300d.txt','r',encoding='utf-8') as f:
for line in f:
values=line.split()
word=values[0]
vectors=np.asarray(values[1:],'float32')
embedding_dict[word]=vectors
f.close()
embedding_dim=300
embedding_matrix = np.zeros((num_words, embedding_dim))
for word, i in tqdm(word_index.items()):
if i < num_words:
embedding_vector = embedding_dict.get(word)
if embedding_vector is not None:
embedding_matrix[i] = embedding_vector
embedding_matrix.shape
# -
# ## RNN without LSTM Layer
# Building the model
model = Sequential()
model.add(Embedding(input_dim=num_words,
output_dim=300,
embeddings_initializer=initializers.Constant(embedding_matrix),
input_length=50,trainable=False))
model.add(Dropout(0.2))
model.add(SimpleRNN(units=64, activation="sigmoid"))
model.add(Dense(units=1, activation="sigmoid"))
model.compile(optimizer="adam", loss="binary_crossentropy", metrics=['accuracy'])
model.summary()
#Fit the mode and evaluate the model
model_1_fit = model.fit(X_train, Y_train, validation_split=.25, epochs=10, batch_size=10)
model.evaluate(X_test, Y_test, batch_size=10)
#Save model file to disk
model_json = model.to_json()
with open("rnnmodel.json", "w") as json_file:
json_file.write(model_json)
# serialize weights to HDF5
model.save_weights("rnnmodel.h5")
print("Saved model to disk")
# ## RNN with LSTM Layer
# +
# Building the model
model2 = Sequential()
model2.add(Embedding(input_dim=num_words,
output_dim=300,
embeddings_initializer=initializers.Constant(embedding_matrix),
input_length=50,trainable=False))
model2.add(Dropout(0.2))
model2.add(LSTM(64,dropout=0.2, recurrent_dropout=0.2))
model2.add(Dense(units=1, activation="sigmoid"))
model2.compile(optimizer="adam", loss="binary_crossentropy", metrics=['accuracy'])
model2.summary()
# -
#Fit the mode and evaluate the model
model_2_fit = model2.fit(X_train, Y_train, validation_split=.25, epochs=10, batch_size=10)
model2.evaluate(X_test, Y_test,batch_size=10)
#Save model file to disk
model2_json = model2.to_json()
with open("rnn2model.json", "w") as json_file:
json_file.write(model2_json)
# serialize weights to HDF5
model.save_weights("rnn2model.h5")
print("Saved model to disk")
| 3.2_RNN_GloVe.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.8 64-bit ('NLP')
# metadata:
# interpreter:
# hash: 2136a9c3637fd160483224d7922e48bf03b650be5dff26724a0c1f8d1279953b
# name: python3
# ---
# # DaCy and Sentiment
# DaCy currently does not include any tools for sentiment extraction, but a couple of good tools already exists. Thus DaCy wraps these in the DaCy framework, but if you use this in a publication or similar be sure to credit the original authors.
# !pip install git+https://github.com/KennethEnevoldsen/DaCy
import dacy
import spacy
# ## BertTone
# ---
#
# BertTone is a model trained by DaNLP, actually it is two. One for classification of polarity (whether a sentence is positive, negative or neutral) and subjectivity (whether a text is subjective or not).
#
# To read more about BertTone as well as its performance matched against other models see DaNLP's [GitHub](https://github.com/alexandrainst/danlp/blob/master/docs/docs/tasks/sentiment_analysis.md).
#
# Here I will show a simple use case of both models. Furthermore if you wish to inspect the TransformerData to see e.g. the used wordpieces you check out the `doc._.berttone_subj_trf_data` or `doc._.berttone_pol_trf_data`
nlp = spacy.blank("da")
nlp = dacy.sentiment.add_berttone_subjectivity(nlp)
# +
texts = ["Analysen viser, at økonomien bliver forfærdelig dårlig",
"Jeg tror alligvel, det bliver godt"]
docs = nlp.pipe(texts)
for doc in docs:
print(doc._.subjectivity)
print(doc._.subjectivity_prop)
# +
nlp = dacy.sentiment.add_berttone_polarity(nlp)
docs = nlp.pipe(texts)
for doc in docs:
print(doc._.polarity)
print(doc._.polarity_prop)
# -
# BertEmotion
# ---
#
# Siliar to BertTone is a BertEmoiton is a model trained by DaNLP, actually it is also two. One for classifying whether a text is emotionally laden or not, and one for emotion classification using. The possible emotions to classify from is:
#
# - "Glæde/Sindsro"
# - "Tillid/Accept"
# - "Forventning/Interrese"
# - "Overasket/Målløs"
# - "Vrede/Irritation"
# - "Foragt/Modvilje"
# - "Sorg/trist"
# - "Frygt/Bekymret"
#
# Their transformerData can be accessed using `bertemotion_laden_trf_data` for the model whether a text is emotionally laden and `bertemotion_emo_trf_data` for the model predicting emotion. Similarly to above you can always use the `*_prop` prefix to extract the probabilities of each label.
nlp = dacy.sentiment.add_bertemotion_laden(nlp) # whether a text is emotionally laden
nlp = dacy.sentiment.add_bertemotion_emo(nlp) # what emotion is portrayed
# +
texts = ['bilen er flot',
'jeg ejer en rød bil og det er en god bil',
'jeg ejer en rød bil men den er gået i stykker',
"Ifølge TV udsendelsen så bliver vejret skidt imorgen",
"Fuck jeg hader bare Hitler. Han er bare så FUCKING træls!",
"Har i set at Tesla har landet en raket på månen? Det er vildt!!",
"Nu må vi altså få ændret noget",
"En sten kan ikke flyve. Morlille kan ikke flyve. Ergo er morlille en sten!"]
docs = nlp.pipe(texts)
for doc in docs:
print(doc._.laden)
print("\t", doc._.emotion)
# -
# Den opmærksomme person, ville med rette undre sig over outputtet altid er emotional. Det gjorde jeg også selv så har forsøgt med en lang række eksempler. Desværre har jeg endnu ikke fundet et som giver et "No emotion"-tagget. Har indberettet dette som en fejl til DaNLP's [GitHub](https://github.com/alexandrainst/danlp/issues/122). Dog som det ser ud pt. vil jeg ikke anbefale at bruge modellen i praktiske applikationer. Det ses særligt tydeligt i det næste output som også printer sandsynlighederne:
docs = nlp.pipe(texts)
for doc in docs:
print(doc._.laden_prop)
# ## DaVader
#
# ---
#
# DaVader is a Danish Sentiment model developing using [Vader](https://github.com/fnielsen/afinn) and the dictionary list of [SentiDa](https://github.com/guscode/sentida) and [AFINN](https://github.com/fnielsen/afinn). This adaption is developed by Center for Humanities Computing Aarhus and by the author of this package. It is a lexicon and rule-based sentiment analysis tool which predict sentiment valence - the degree to which a text is positive or negative - as opposed to BertTone which simply predict whether or not it is.
#
# An additional advantage of it being rule-based is that it is transparent (the entire lexion can be found in the sentiment folder) and very fast compared to transformer-based approaches.
# +
from spacy.tokens import Doc
from dacy.sentiment import da_vader_getter
Doc.set_extension("vader_da", getter=da_vader_getter)
# +
nlp = spacy.load("da_core_news_sm")
texts = ['Jeg er så glad', 'jeg ejer en rød bil og det er en god bil', 'jeg ejer en rød bil men den er gået i stykker']
docs = nlp.pipe(texts)
for doc in docs:
print(doc._.vader_da)
# -
# If you are have never used a VADER model before I suggest you read the ["about the scoring"](https://github.com/cjhutto/vaderSentiment#about-the-scoring) on the website for the original (English) VADER implementation.
| tutorials/dacy-sentiment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# orphan: true
# ---
# + tags=["remove-input", "active-ipynb", "remove-output"]
# try:
# import openmdao.api as om
# except ImportError:
# !python -m pip install openmdao[notebooks]
# import openmdao.api as om
# -
# # Connecting Variables
#
# To cause data to flow between two systems in a model, we must connect at
# least one output variable from one system to at least one input variable
# from the other. If the variables have units defined, then the framework
# will automatically perform the conversion. We can also connect only part
# of an array output to an input by specifying the indices of the entries
# that we want.
#
# To connect two variables within a model, use the `connect` function.
#
# ```{eval-rst}
# .. automethod:: openmdao.core.group.Group.connect
# :noindex:
# ```
#
# ## Usage
#
# 1: Connect an output variable to an input variable, with an automatic unit conversion.
# +
import numpy as np
p = om.Problem()
p.model.set_input_defaults('x', np.ones(5), units='ft')
exec_comp = om.ExecComp('y=sum(x)',
x={'value': np.zeros(5), 'units': 'inch'},
y={'units': 'inch'})
p.model.add_subsystem('comp1', exec_comp, promotes_inputs=['x'])
p.setup()
p.run_model()
print(p.get_val('x', units='ft'))
print(p.get_val('comp1.x'))
print(p.get_val('comp1.y'))
# + tags=["remove-input", "remove-output"]
from openmdao.utils.assert_utils import assert_near_equal
assert_near_equal(p.get_val('x', units='ft'), np.ones(5))
assert_near_equal(p.get_val('comp1.x'), np.ones(5)*12.)
assert_near_equal(p.get_val('comp1.y'), 60.)
# -
# 2: Connect one output to many inputs.
# +
p = om.Problem()
p.model.add_subsystem('C1', om.ExecComp('y=sum(x)*2.0', x=np.zeros(5)), promotes_inputs=['x'])
p.model.add_subsystem('C2', om.ExecComp('y=sum(x)*4.0', x=np.zeros(5)), promotes_inputs=['x'])
p.model.add_subsystem('C3', om.ExecComp('y=sum(x)*6.0', x=np.zeros(5)), promotes_inputs=['x'])
p.setup()
p.set_val('x', np.ones(5))
p.run_model()
print(p.get_val('C1.y'))
print(p.get_val('C2.y'))
print(p.get_val('C3.y'))
# + tags=["remove-input", "remove-output"]
assert_near_equal(p.get_val('C1.y'), 10.)
assert_near_equal(p.get_val('C2.y'), 20.)
assert_near_equal(p.get_val('C3.y'), 30.)
# -
# (connect-with-src-indices)=
# 3: Connect only part of an array output to an input of a smaller size.
# +
p = om.Problem()
p.model.add_subsystem('indep', om.IndepVarComp('x', np.ones(5)))
p.model.add_subsystem('C1', om.ExecComp('y=sum(x)*2.0', x=np.zeros(3)))
p.model.add_subsystem('C2', om.ExecComp('y=sum(x)*4.0', x=np.zeros(2)))
# connect C1.x to the first 3 entries of indep.x
p.model.connect('indep.x', 'C1.x', src_indices=[0, 1, 2])
# connect C2.x to the last 2 entries of indep.x
# use -2 (same as 3 in this case) to show that negative indices work.
p.model.connect('indep.x', 'C2.x', src_indices=[-2, 4])
p.setup()
p.run_model()
print(p['C1.x'])
print(p['C1.y'])
print(p['C2.x'])
print(p['C2.y'])
# + tags=["remove-input", "remove-output"]
assert_near_equal(p['C1.x'], np.ones(3))
assert_near_equal(p['C1.y'], 6.)
assert_near_equal(p['C2.x'], np.ones(2))
assert_near_equal(p['C2.y'], 8.)
# -
# 4: Connect only part of a non-flat array output to a non-flat array input.
# +
p = om.Problem()
p.model.add_subsystem('indep', om.IndepVarComp('x', np.arange(12).reshape((4, 3))))
p.model.add_subsystem('C1', om.ExecComp('y=sum(x)*2.0', x=np.zeros((2, 2))))
# connect C1.x to entries (0,0), (-1,1), (2,1), (1,1) of indep.x
p.model.connect('indep.x', 'C1.x',
src_indices=[[(0, 0), (-1, 1)],
[(2, 1), (1, 1)]], flat_src_indices=False)
p.setup()
p.run_model()
print(p.get_val('indep.x'))
print(p.get_val('C1.x'))
print(p.get_val('C1.y'))
# + tags=["remove-input", "remove-output"]
assert_near_equal(p.get_val('indep.x'), np.array([[0., 1., 2.],
[3., 4., 5.],
[6., 7., 8.],
[9., 10., 11.]]))
assert_near_equal(p.get_val('C1.x'), np.array([[0., 10.],
[7., 4.]]))
assert_near_equal(p.get_val('C1.y'), 42.)
| openmdao/docs/openmdao_book/features/core_features/working_with_groups/connect.ipynb |