code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Detecting Cultural Violence using Natural Language processing.
#
# We seek to detect cultural violence in natural language through measuring the self-other gradient. Cultural violence is a theory proposed by <NAME> which seeks to explain how aspects of culture - religion, ideology, language, art along with formal and empirical science - are used to legitimise violence. This notebook is focusses on the representation of religion and ideology in language. Galtung explores how each aspect can be used as a mode of influence to create a ‘self-other gradient’ between ‘Chosen People’ - referred to as an ingroup - and others deemed ‘lower down the scale of worthiness’ - referred to as an outgroup. A general thesis follows whereby the steeper the gradient, the more legitimate violence becomes. To measure the self-other gradient, therefore, means establishing a schema for measuring how aspects of culture are used to elevate the self as an ingroup and debase the other as an outgroup.
#
# The main contributions of this paper are as follows
# 1. Propose cultural violence as a guiding theory for detecting harmful content in natural language.
# 2. Propose a novel methodology for detecting harmful content in natural language.
# 3. Identifying where current general purpose NLP technologies fall short in the specific task of detecting cultural violence
# 4. A new nlp pipeline workflow for annotating the ingroup andoutgroups in natural language texts.
#
# In having proposed cultural violence as a guiding theory, we proposed the following three step methodology.
# 1. Detect the ingroup and outgroup of an orator's text
# 2. Detect how each mode of influence is used for elevating the ingroup and debase the outgroup to create a gradient between each
# 3. Devise a schema for measuring the gradient
#
# For the detectiong of cultural violence in natural langugage we test existing Natural Language Processing (NLP) technologies for each step of the methodology from which new pipeline components have been devised. NLP is a branch of artificial intelligence which seeks to process natural language to derive meaning. In general terms there are two fields of NLP, theoretical and applied. Theoretical NLP is concerned with the technical aspects of processing language and Applied NLP is concerned with the application of such technologies. This research sits within applied NLP and in seeking to qualify sociological theory sits within the field of the Digital Humanities.
#
# The datasets we are using for these test are speech transcripts of George Bush and Osama bin Laden in which they advocated violence, therefore, these datasets contain cultural violence.
#
# As a reference dataset we use speeches made by <NAME> who uses many of the same aspects of culture used by Bush and bin Laden, but he does not advocate for violence. The difference between Luther King's and the other speeches, therefore, may reveal the defining features of culturally violent language.
#
# The tests are as follows:
# 1. A test of named entity recognition in the spaCy language models against the dataset.
# 2. A test of sentiment analysis technologies for detecting the ingroup and outgroup of a text.
# 3. Detailed testing of the Watson API
#
# From these tests, the following pipeline components have been created:
# 1. Supplemtary Named Entity Recognition - contains corrections for named entities either not identified or incorrectly identified by the model.
# 2. A typology of cultural violence based on Mike Martin's 'Why We Fight' and Social Identity Theory for detecting aspects of culture.
# 3. Named Concept Recognition - Using the cultural violence typology, and seed words from each speech, a component for detecting concepts related to each aspect of culture.
#
# This notebook now begins with Identifying the ingroup and outgroup of a text.
# ## Methodology Step 1: Identifying the ingroup and outgroup of a text
#
# The purpose of this notebook is to test the feasibility of detecting the ingroup and outgroup of a text.
#
# To detect each of these groups is the foundation for measuring the self-other gradient of cultural violence.
#
# Through various techniques, sentiment analysis seeks to quantify the sentiment of a text.
#
# This test uses the spaCy, which is a free, open-source library for advanced Natural Language Processing (NLP) in Python.
#
# For sentiment analysis we test TextBlob, which uses word scoring, and IBM's Watson.
#
# We begin with the first cell which imports the relevant libraries
# +
import sys
import platform
import json
import pandas as pd
import datetime
import tqdm
import spacy
from spacy.tokens import Span
from spacy.pipeline import merge_entities
from spacy.matcher import Matcher
from spacy import displacy
from VPipeLibrary.custpipe import EntityMatcher
pd.set_option('display.max_colwidth', -1)
pd.set_option("display.max_columns", 2000)
pd.set_option("display.max_rows", 2000)
spacy.info()
print('============================== Info about python ==============================')
print('python version: ', platform.sys.version)
print(f'completed at: {datetime.datetime.now().strftime("%b %d %Y %H:%M:%S")}')
# -
# ## The spaCy language model
#
# The spacy module is based on langugage models which provide a reference dataset for predicting linguistic annotations. spaCy v2.0 features neural models for tagging, parsing and entity recognition.
#
# For english there are 8 models listed as follows
# 1. en_core_web_sm: English multi-task CNN trained on OntoNotes. Assigns context-specific token vectors, POS tags, dependency parse and named entities.
#
# 2. en_core_web_md: English multi-task CNN trained on OntoNotes, with GloVe vectors trained on Common Crawl. Assigns word vectors, context-specific token vectors, POS tags, dependency parse and named entities
#
# 3. en_core_web_lg: English multi-task CNN trained on OntoNotes, with GloVe vectors trained on Common Crawl. Assigns word vectors, context-specific token vectors, POS tags, dependency parse and named entities.
#
# 4. en_vectors_web_lg
#
# 5. en_trf_bertbaseuncased_lg: Provides weights and configuration for the pretrained transformer model bert-base-uncased, published by Google Research. The package uses HuggingFace's transformers implementation of the model. Pretrained transformer models assign detailed contextual word representations, using knowledge drawn from a large corpus of unlabelled text. You can use the contextual word representations as features in a variety of pipeline components that can be trained on your own data.
#
# 6. en_trf_robertabase_lg: Provides weights and configuration for the pretrained transformer model roberta-base, published by Facebook. The package uses HuggingFace's transformers implementation of the model. Pretrained transformer models assign detailed contextual word representations, using knowledge drawn from a large corpus of unlabelled text. You can use the contextual word representations as features in a variety of pipeline components that can be trained on your own data.
#
# 7. en_trf_distilbertbaseuncased_lg: Provides weights and configuration for the pretrained transformer model distilbert-base-uncased, published by Hugging Face. The package uses HuggingFace's transformers implementation of the model. Pretrained transformer models assign detailed contextual word representations, using knowledge drawn from a large corpus of unlabelled text. You can use the contextual word representations as features in a variety of pipeline components that can be trained on your own data.
#
# 8. en_trf_xlnetbasecased_lg: Provides weights and configuration for the pretrained transformer model xlnet-base-cased, published by CMU and Google Brain. The package uses HuggingFace's transformers implementation of the model. Pretrained transformer models assign detailed contextual word representations, using knowledge drawn from a large corpus of unlabelled text. You can use the contextual word representations as features in a variety of pipeline components that can be trained on your own data.
#
# The accuracy of an nlp tasks very much depends on the model.
#
# This test uses en_core_web_md.
# +
print('=========================== Loading Language Models ===========================')
model = 'en_core_web_md'
print('loading', model)
nlp = spacy.load(model)
print('loaded', model)
print(f'completed at: {datetime.datetime.now().strftime("%b %d %Y %H:%M:%S")}')
# +
{
"tags": [
"hide_input",
]
}
from IPython.core.display import Image, display
image_filepath = r"C:/Users/Steve/Documents/CulturalViolence/spacyPipeline.png"
display(Image(image_filepath, width=1900, unconfined=False))
# -
# image_weblink = 'https://spacy.io/pipeline-7a14d4edd18f3edfee8f34393bff2992.svg'
# 
# ## The spaCy pipeline
#
# The spacy pipeline, shown in the image above is based on a series of components shown in the table below.
#
# The processing pipeline always depends on the statistical model and its capabilities. For example, a pipeline can only include an entity recognizer component if the model includes data to make predictions of entity labels
# +
{
"tags": [
"hide_input",
]
}
Col1 = "Component"
Col2 = "Creates Objects"
Col3 = "Description"
pipeline = {
"Tokenizer" : {Col1 : "Tokenizer", Col2 : "Doc", Col3 : "Segment text into tokens."},
"Tagger" : {Col1 : "Tagger", Col2 : "Doc[i].tag", Col3 : "Assign part-of-speech tags."},
"Parser" : {Col1 : "Dependency Parser", Col2 : "Doc[i].head, Doc[i].dep, Doc.sents, Doc.noun_chunks", Col3 : "Assign dependency labels."},
"ner" : {Col1 : "Entity Recognizer", Col2 : "Doc.ents, Doc[i].ent_iob, Doc[i].ent_type",Col3 : "Detect and label named entities."},
"textcat" : {Col1 : "Text Categorizer", Col2 : "Doc.cats", Col3 : "Assign document labels."},
"..." : {Col1 : "Custom Components", Col2 : "Doc._.xxx, Token._.xxx, Span._.xxx", Col3 : "Assign custom attributes, methods or properties."}
}
display(pd.DataFrame(pipeline).T)
# -
# ## Named Concept Matcher
#
# Since the models have been generated for general purpose, they require tuning for a specifc context.
#
# The tuning in this test is based on a typology of cultural violence developed for this research. The typology is based on seven modes of influence with associated seed words for each category. The modes of influence are as follow:
# 1. Social: everyday terms using the social environment
# 2. Medical: terms related to parts of the body and health
# 3. Religion: terms related to a religion
# 4. Geopolitics: terms related to geopolitical entities
# 5. Economics: terms related to financial economics
# 6. Justice: terms related to security and justice including the law
# 7. Military: terms related to a military environment.
#
# Each mode of influence is modeled using the 5 group problems propsed by Dr <NAME> in 'Why We Fight'. For each category and subcategory seed terms have been identified thereby employing named concept recognition.
#
# - Identity: how the group is identified
# -- Group categories identifying the ingroup and outgroup
# -- Entities identifying the mode of influence as a context
# - Hierarchy: how is the group organised
# -- Terms related to a rank, title within each context
# - Trade: the ideologies defining group interactions determined to be good or bad?
# -- ideologies defining what are good terms of trade
# -- ideologies defining what are bad terms of trade
# - Disease:does the group control disease
# -- separated out with a distinct category
# - Punishment: how the group determines right and wrong)
# -- Actions and concepts determined to be right
# -- Actions and concepts determined to be wrong
#
# +
class NamedConcept(object):
def __init__(self, nlp):
self.matcher = Matcher(nlp.vocab)
with open(r'C:\Users\Steve\Documents\CulturalViolence\KnowledgeBases\group_typology.json', 'r') as fp:
self.named_concept_rules = json.load(fp)
for entry in self.named_concept_rules.values():
for pattern in entry.values():
for subcat, terms in pattern.items():
self.matcher.add(subcat, None, [{"LEMMA" : {"IN" : terms}}])
def __call__(self, doc):
matches = self.matcher(doc)
spans = [] # keep the spans for later so we can merge them afterwards
for match_id, start, end in matches:
## gather up noun phrases
concept = Span(doc, start, end, label=doc.vocab.strings[match_id])
if "the" in [word.lower_ for word in list(doc[start].lefts)]:
Span(doc, start - doc[start].n_lefts, end + doc[start].n_rights, label = doc.vocab.strings[match_id])
#print(concept, '=>', concept.label_)
# elif doc[start].dep_ in ["poss", "compound"] and end != len(doc):
# try:
# concept = Span(doc, start, list(doc[start + 1].rights)[-1].i + 1, label=doc.vocab.strings[match_id])
# print(concept, '=>', concept.label_)
# except:
# continue
elif doc[start - 1].dep_ in ["amod", "compound"] and start != 0:
if doc[start -1].ent_type_:
concept = Span(doc, start - 1, end, label=doc[start -1].ent_type_)
else:
concept = Span(doc, start - 1, end, label=doc.vocab.strings[match_id])
#print(concept, '=>', concept.label_)
elif doc[start - 1].pos_ in ["NOUN", "PROPN"] and start != 0:
concept = Span(doc, start - 1, end, label=doc.vocab.strings[match_id])
#print(concept, '=>', concept.label_)
elif doc[start + 1].pos_ in ["NOUN", "PROPN"] and end != len(doc):
concept = Span(doc, start, end + 1, label=doc.vocab.strings[match_id])
#print(concept, '=>', concept.label_)
# elif doc[start - 2].dep_ in ["nsubj", "amod"] and doc[start].dep_ in ["pobj"] and start != 0:
# concept = Span(doc, start - 2, end, label=doc.vocab.strings[match_id])
# #print(concept, '=>', concept.label_)
elif doc[start].dep_ in ["nsubj", "csubj", "pobj"] and end != len(doc):
if doc[start + 1].dep_ in ["prep"]:
try:
concept = Span(doc, start, list(doc[start + 1].rights)[-1].i + 1, label=doc.vocab.strings[match_id])
#print(concept, '=>', concept.label_)
except:
continue
doc.ents = spacy.util.filter_spans(list(doc.ents) + [concept])
spans.append(concept)
with doc.retokenize() as retokenizer:
# Iterate over all spans and merge them into one token. This is done
# after setting the entities – otherwise, it would cause mismatched
# indices!
for span in spacy.util.filter_spans(doc.ents):
retokenizer.merge(span)
return doc # don't forget to return the Doc!
from spacy.pipeline import EntityRuler
# remove all pipeline components
for pipe in nlp.pipe_names:
if pipe not in ['tagger', "parser", "ner"]:
nlp.remove_pipe(pipe)
#add entity ruler to the pipe
with open(r'C:\Users\Steve\Documents\CulturalViolence\KnowledgeBases\group_typology.json', 'r') as fp:
group_typology = json.load(fp)
ruler = EntityRuler(nlp)
patterns = []
for entry in group_typology.values():
for pattern in entry.values():
for subcat, terms in pattern.items():
patterns.append({"label" : subcat, "pattern" : [{"LEMMA" : {"IN" : terms}}]})
# add new pipe components
ruler.add_patterns(patterns)
#nlp.add_pipe(ruler)
nlp.add_pipe(EntityMatcher(nlp), before = "ner")
nlp.add_pipe(NamedConcept(nlp), after = "ner")
nlp.add_pipe(merge_entities)
print(f'completed at: {datetime.datetime.now().strftime("%b %d %Y %H:%M:%S")}')
# -
# ## IBM Watson
#
# (insert information about IBM Watson)
# +
import json
from ibm_watson import NaturalLanguageUnderstandingV1
from ibm_cloud_sdk_core.authenticators import IAMAuthenticator
from ibm_watson.natural_language_understanding_v1 import Features, EntitiesOptions, KeywordsOptions, SentimentOptions
apikey = '<KEY>'
url = 'https://api.eu-gb.natural-language-understanding.watson.cloud.ibm.com/instances/204e6ba7-952c-41ae-99e9-fe4e8208bfde'
authenticator = IAMAuthenticator(apikey)
service = NaturalLanguageUnderstandingV1(version='2019-07-12', authenticator=authenticator)
service.set_service_url(url)
# response = service.analyze(
# text="The evidence we have gathered all points to a collection of loosely affiliated terrorist organizations known as al-Qa\'eda.",
# features=Features(sentiment=SentimentOptions()
# )).get_result()
# #entities=EntitiesOptions(emotion=True, sentiment=True, limit=2)
# #keywords=KeywordsOptions(emotion=True, sentiment=True, limit=2)
# print(json.dumps(response, indent=2))
print(f'completed at: {datetime.datetime.now().strftime("%b %d %Y %H:%M:%S")}')
# +
with open(r'C:\Users\Steve\Documents\CulturalViolence\KnowledgeBases\bush_filelist.json', 'r') as fp:
bush_filelist = json.load(fp)
bushpath = 'C:/Users/Steve/Documents/CulturalViolence/George Bush/'
text = ''
for file in bush_filelist[3:]:
with open(bushpath + file[1], 'r') as fp:
text = text + fp.read()
#doc = nlp(text)
targets = list({ent.lower_ for ent in doc.ents if ent.label_ in named_entities})
# response = service.analyze(text=text, features=Features( \
# sentiment=SentimentOptions(targets=targets), \
# entities=EntitiesOptions(sentiment=True), \
# keywords=KeywordsOptions(sentiment=True,emotion=True)
# )).get_result()
with open(r"C:\Users\Steve\Documents\CulturalViolence\KnowledgeBases\Bush_Analysis_v2.json", "wb") as f:
f.write(json.dumps(response).encode("utf-8"))
print(f'completed at: {datetime.datetime.now().strftime("%b %d %Y %H:%M:%S")}')
# -
# ## TextBlob
#
# (insert information about TextBlob)
from textblob import TextBlob
from textblob.np_extractors import ConllExtractor
from textblob.sentiments import NaiveBayesAnalyzer
extractor = ConllExtractor()
# +
with open(r'C:\Users\Steve\Documents\CulturalViolence\George Bush\20010120-Address to Joint Session of Congress Following 911 Attacks.txt', 'r') as fp:
doc = nlp(fp.read())
for sentence in doc.sents:
displacy.render(sentence, style = "ent")
# -
# ## Test 1: using IBM Watson and Text Blob for detecting ingroup and outgroup
#
# This test is based on the following steps
#
# 1. take sentences containing named entity and mode of influence
# 2. sentiment analysis using some library
# 3. if the sentence is positive entity is tagged as ingroup
# 4. if the sentence is negative entity is tagged as outgroup
#
# +
#list of named entities
named_entities = set()
named_entities = {"NORP", "GPE", "NORP", "PERSON", "ORG"}
#list of labels for each mode of influence
labels = set()
for value in group_typology.values():
for subcat in value.values():
for term in list(subcat.keys()):
labels.add(term)
# a dictionary object of ingroup and outgroup sentences determined by sentiment score
groups = {"ingroup" : [], "outgroup" : []}
for sentence in tqdm.tqdm(doc.sents, total = len(list(doc.sents))):
#get the tokens.ent_types_ for each token in the sentence
ent_types = {token.ent_type_ for token in sentence if token.ent_type_}
# if the sentence ent_types contain both a named entities and mode of influence
if not ent_types.isdisjoint(named_entities) and not ent_types.isdisjoint(labels):
# get the sentiment score for TextBlob
textblob_score = TextBlob(sentence.text).sentiment.polarity
# get the score for IBM Watson
#response = service.analyze(text=sentence.text,features=Features(sentiment=SentimentOptions())).get_result()
#watson_score = response['sentiment']['document']['score']
watson_score = 0
# result for the sentence
result = (sentence.start, sentence.end, textblob_score, watson_score)
## append to ingroup category if either have a positive score and vice versa
if watson_score > 0 or textblob_score > 0:
groups["ingroup"].append(result)
elif textblob_score < 0 or watson_score < 0:
groups["outgroup"].append(result)
with open(r"C:\Users\Steve\Documents\CulturalViolence\KnowledgeBases\group_sentiment_ruler.json", "wb") as f:
f.write(json.dumps(groups).encode("utf-8"))
print(f'completed at: {datetime.datetime.now().strftime("%b %d %Y %H:%M:%S")}')
# -
# ## Test Results
#
# We can see that neither is able to detect the sentences defining the ingroup and outgroup of a text.
#
# Despite Watson's sophistication, it provides no more value than TextBlob, which is a much simpler technology.
#
# Through observation, we know the following are outgroup named entities from Bush's speech:
# 1. al Qaeda
# 2. Egyptian Islamic Jihad
# 3. Islamic Movement of Uzbekistan
# 4. the Taliban
#
# A successful test with return these named entities as an outgroup.
# +
with open(r'C:\Users\Steve\Documents\CulturalViolence\KnowledgeBases\group_sentiment_namedconcept.json', 'r') as fp:
groups = json.load(fp)
group = "ingroup"
for index in groups[group]:
print('-----')
print(f'these are the result for {group}')
sentence = doc[index[0]:index[1]]
token = []
pos = []
ent_type = []
sentiment = []
dep = []
sent_table = {
"token" : [token.text for token in sentence],
"dep" : [token.dep_ for token in sentence],
"ent_type" : [token.ent_type_ for token in sentence],
"pos" : [token.pos_ for token in sentence],
"sentiment" : [TextBlob(token.text).sentiment.polarity for token in sentence]
}
print(f'{group} sentence sentiment score for TextBlob: {index[2]}')
print(f'{group} sentence sentiment score for Watson: {index[3]}')
display(pd.DataFrame.from_dict(sent_table, orient = "index"))
# -
# ## Test 2. Sentiment related to Named Entities
#
# The following test uses Watson for detecting sentiment related to target terms.
#
# The target terms are user defined and here based on the spaCy NER model with the supplementary component.
#
# The technique works by sending a text and list of target terms to the API and the scores related to each are returned.
#
# These results show this technique negatively scores a number of entities which would reasonably be expected to be positive.
# +
with open(r'C:\Users\Steve\Documents\CulturalViolence\KnowledgeBases\Bush_Analysis_v2.json', 'r') as fp:
response = json.load(fp)
table = response["sentiment"]["targets"]
keys = "sentiment"
frames = []
print(f'document sentiment score:{response["sentiment"]["document"]["score"]}')
print(f'document sentiment: {response["sentiment"]["document"]["label"]}')
sentiment = "negative"
frames = []
for entry in table:
objs = {"sentiment" : entry["score"], "label" : entry["label"]}
if objs["label"] == sentiment:
df = pd.DataFrame(objs, index = [entry["text"]], columns = list(objs.keys())).fillna("")
frames.append(df)
display(pd.concat(frames, sort = False).sort_values("sentiment", ascending = True).style.background_gradient(cmap=cmp))
# -
# ## Test 3. Sentiment Scores for feature words of the Text
#
# This next test shows how Watson scores feature terms of the text.
#
# The feature terms are assigned by the Watson API.
#
# This test develops upon a simple score and provides some degree of explanation as to why a feature is scored negatively.
#
# For example:
# - Al Qaeda is scored negatively along with the emotion of fear.
# - The Taliban Regime is scored negatively with the emotion of disgust.
# - Terrorists is scored negatively with the emotion of anger.
# - American people is also scored negatively but with the emotion of sadness.
#
# While these results seem plausible, however, the API does not seem to provide much explanatory value for the other features.
#
# Moreover, there is no explanation as to why these decisions have been made.
# +
table = response["keywords"]
frames = []
print(f'document sentiment score:{response["sentiment"]["document"]["score"]}')
print(f'document sentiment: {response["sentiment"]["document"]["label"]}')
#text => sentiment => relvance => emotion => count
sentiment = "negative"
for entry in table:
objs = {"count" : entry["count"], "sentiment" : entry["sentiment"]["score"], "label" : entry["sentiment"]["label"], \
"sadness" : entry["emotion"]["sadness"], "joy" : entry["emotion"]["joy"], "fear" : entry["emotion"]["fear"], \
"disgust": entry["emotion"]["disgust"], "anger" : entry["emotion"]["anger"]}
if objs["label"] == sentiment:
df = pd.DataFrame(objs, index = [entry["text"]], columns = list(objs.keys())).fillna("")
frames.append(df)
print(len(table))
cmp = "Reds"
### don't work on this one
display(pd.concat(frames, sort = False).sort_values("sentiment", ascending = True).style.background_gradient(cmap=cmp))
# -
# ## Test 4. Sentiment Scores for Watson Defined Entities
#
# This final test looks at the scores asigned to the named entities identified by Watson as opposed to those which are user defined.
#
# In the first instance, while Al Qaeda and the Taliban are identified, the Egyptian Islamic Jihad" and "Islamic Movement of Uzbekistan" - both identified by Bush as adversaries - have not been identified by Watson.
#
# Much like the other Watson components, entities which would reasonably be expected to be scored positively are scored negatively.
# +
table = response["entities"]
frames = []
print(f'document sentiment score:{response["sentiment"]["document"]["score"]}')
print(f'document sentiment: {response["sentiment"]["document"]["label"]}')
#text => sentiment => relvance => emotion => count
#print([e["text"] for e in table])
sentiment = "negative"
print(len(table))
for entry in table:
objs = {'count' : entry["count"], 'type' : entry["type"], 'sentiment': entry["sentiment"]["score"], 'label' : entry["sentiment"]["label"]}
if objs["label"] == sentiment:
df = pd.DataFrame(objs, index = [entry["text"]], columns = list(objs.keys())).fillna("")
frames.append(df)
cmp = "Reds"
### don't work on this one
display(pd.concat(frames, sort = False).sort_values(["sentiment"], ascending = False).style.background_gradient(cmap=cmp))
| Quantitative Analysis/.ipynb_checkpoints/1. Group Identifier-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="klGNgWREsvQv"
# # "Control of a Cart-Pole Dynamic System with Ray RLlib"
#
# > Reinforcement Learning (RL) to control the balancing of a pole on a moving cart
# - toc: true
# - branch: master
# - badges: false
# - comments: true
# - hide: false
# - search_exclude: true
# - metadata_key1: metadata_value1
# - metadata_key2: metadata_value2
# - image: images/RLlib-CartPole.png
# - categories: [Control, Reinforcement_Learning, Ray_RLlib]
# - show_tags: true
# + id="4E8ujfcND6pJ" colab={"base_uri": "https://localhost:8080/"} outputId="623946a3-8f15-4cac-d702-84bb76aa06a2"
#hide
from google.colab import drive
drive.mount('/content/gdrive', force_remount=True)
root_dir = "/content/gdrive/My Drive/"
base_dir = root_dir + 'RLlib/'
# base_dir = ""
# + [markdown] id="lsaQlK8fFQqH"
# ## 1. Introduction
#
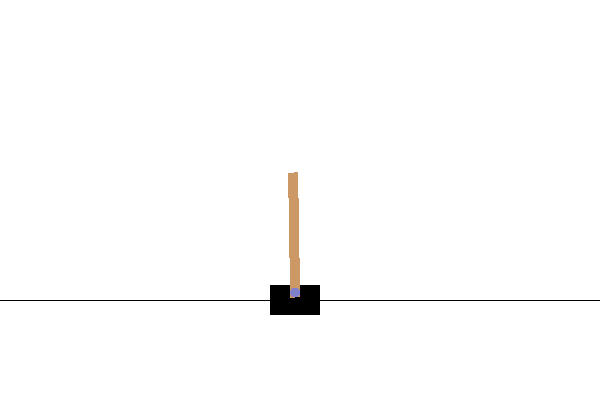
# The cart-pole problem can be considered as the "Hello World" problem of Reinforcement Learning (RL). It was described by [Barto (1983)](http://www.derongliu.org/adp/adp-cdrom/Barto1983.pdf). The physics of the system is as follows:
#
# * All motion happens in a vertical plane
# * A hinged pole is attached to a cart
# * The cart slides horizontally on a track in an effort to balance the pole vertically
# * The system has four state variables:
#
# $x$: displacement of the cart
#
# $\theta$: vertical angle on the pole
#
# $\dot{x}$: velocity of the cart
#
# $\dot{\theta}$: angular velocity of the pole
#
# + [markdown] id="cKOCZlhUgXVK"
# Here is a graphical representation of the system:
#
# 
# + [markdown] id="S0-ZNxUXoMR6"
# ## 2. Purpose
#
# The purpose of our activity in this blog post is to construct and train an entity, let's call it a *controller*, that can manage the horizontal motions of the cart so that the pole remains as close to vertical as possible. The controlled entity is, of course, the *cart and pole* system.
# + [markdown] id="1u9QVVsShC9X"
# ## 3. RLlib Setup
#
# We will use the Ray RLlib framework. In addition, this notebook will be run in Google Collab.
# + id="KEHR2Ui-lo8O" colab={"base_uri": "https://localhost:8080/"} outputId="9e43ab4a-794b-49e5-e228-425214225366"
# !pip install ray[rllib]
# + id="nz7IE2gQbBA5"
import ray
import ray.rllib.agents.ppo as ppo
import pandas as pd
import json
import os
import shutil
import sys
# + colab={"base_uri": "https://localhost:8080/"} id="1K7_uLc_nrCs" outputId="c0c73e74-538d-428a-ad13-2989f6bb43f5"
# !pip list | grep ^ray
# + id="_4y2uUcYdZH2"
#
# setup a folder for checkpoints
CHECKPOINT_ROOT = base_dir+"checkpoints/ppo/cart"
# + id="tPpp1fsidZDg"
# hide
# clean output from previous run
# checkpoints: (may want to KEEP)
# shutil.rmtree(CHECKPOINT_ROOT, ignore_errors=True, onerror=None)
# Tensorboard:
# ray_results = f'{os.getenv("HOME")}/ray_results/'
# shutil.rmtree(ray_results, ignore_errors=True, onerror=None)
# + colab={"base_uri": "https://localhost:8080/"} id="JTAaiM03dY6v" outputId="a3b88a41-0e17-47bb-f401-d63502285c81"
# start Ray
ray.init(ignore_reinit_error=True)
# + [markdown] id="y1s77CKrv1m_"
# ## 4. Hyperparameters
# Here we specify all the hyperparameters for the problem:
# + id="psVAtI4Vv4zL"
N_ITERATIONS = 10 #number of training runs
config = ppo.DEFAULT_CONFIG.copy()
config["log_level"] = "WARN"
config["num_workers"] = 1 #use > 1 for using more CPU cores, including over a cluster
config["num_sgd_iter"] = 10 #number of SGD (stochastic gradient descent) iterations per training minibatch
config["sgd_minibatch_size"] = 250
config["model"]["fcnet_hiddens"] = [100, 50]
config["num_cpus_per_worker"] = 0 #avoids running out of resources in the notebook environment when this cell is re-executed
# + [markdown] id="VMsJC3DEgI0x"
# ## 5. Environment
#
# Let's start with the controlled entity. In Reinforcement Learning, the controlled entity is known as an **environment**. We make use of an environment provided by the OpenAI Gym framework, known as "CartPole-v1".
# + id="pYEz-S9gEv2-"
import gym
env = gym.make("CartPole-v1")
# + [markdown] id="NSmAGfjnRzUE"
# ### Input to Environment
#
# Actions to the environment come from an action space with a size of 2.
# + colab={"base_uri": "https://localhost:8080/"} id="xBBaR-jTSJ3B" outputId="563747d5-9197-4a57-c189-50ca892f2384"
env.action_space
# + [markdown] id="wsQ8Qb5gSXSP"
# We will use the convention that the `action` on the cart is as follows:
#
# * `0` means LEFT
# * `1` means RIGHT
# + [markdown] id="r_fatIpdUMMg"
# ### Evolution of the Environment
#
# The arrival of an `action` at the input of the environment leads to the update of its state. This is how the environment evolves. To advance the state of the environment, the `environment.step` method takes an input `action` and applies it to the environment.
#
# The next fragment of code drives the environment through 30 steps by applying random actions:
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="cBGwVeokuWoW" outputId="a7ae6f16-4e5d-4b3d-e5f3-7ad04d2cfd35"
env.reset()
for i in range(30):
observation, reward, done, info = env.step(env.action_space.sample())
print("step", i, observation, reward, done, info)
env.close()
# + colab={"base_uri": "https://localhost:8080/"} id="2TpHYDa2uWjW" outputId="56778ce1-e317-41e8-ccf2-cf0a3d0cd222"
#
# install dependencies needed for recording videos
# !apt-get install -y xvfb x11-utils
# !pip install pyvirtualdisplay==0.2.*
# + id="A9ETYHlCvMjs"
from pyvirtualdisplay import Display
display = Display(visible=False, size=(1400, 900))
_ = display.start()
# + colab={"base_uri": "https://localhost:8080/"} id="oJNYgOQEvMeR" outputId="1a990949-4601-4069-993b-8ca21ee2e649"
from gym.wrappers.monitoring.video_recorder import VideoRecorder
before_training = "before_training.mp4"
video = VideoRecorder(env, before_training)
# returns an initial observation
env.reset()
for i in range(200):
env.render()
video.capture_frame()
observation, reward, done, info = env.step(env.action_space.sample())
video.close()
env.close()
# + id="lJXqoUXRvMWG"
from base64 import b64encode
def render_mp4(videopath: str) -> str:
"""
Gets a string containing a b4-encoded version of the MP4 video
at the specified path.
"""
mp4 = open(videopath, 'rb').read()
base64_encoded_mp4 = b64encode(mp4).decode()
return f'<video width=400 controls><source src="data:video/mp4;' \
f'base64,{base64_encoded_mp4}" type="video/mp4"></video>'
# + colab={"base_uri": "https://localhost:8080/", "height": 288} id="D3q_AKujvMLa" outputId="f666d510-f968-4182-bc46-ffc658604233"
from IPython.display import HTML
html = render_mp4(before_training)
HTML(html)
# + [markdown] id="4MoIFnVtVJuA"
# ### Output from Environment
#
# The output from the environment returns a tuple containing:
#
# * the next observation of the environment
# * the reward
# * a flag indicating whether the episode is done
# * some other information
# + [markdown] id="E9lW_OZYFR8A"
# ## 6. Agent
#
# The controller in our problem is the algorithm used to solve the problem. In RL parlance the controller is known as an `Agent`. RLlib provides implementations of a variety of `Agents`.
#
# For our problem we will use the PPO agent.
#
# The fundamental problem for an Agent is how to find the next best action to submit to the environment.
# + [markdown] id="C58G9mygALn9"
# ## 7. Train the agent
# + id="jPKVnmvHAKV4"
ENV = "CartPole-v1" #OpenAI Gym environment for Cart Pole
# + colab={"base_uri": "https://localhost:8080/"} id="MxjioUCe0Tdt" outputId="a3f81b2d-76dc-4d48-fdfe-54cf973500c5"
agent = ppo.PPOTrainer(config, env=ENV)
results = []
episode_data = []
episode_json = []
for n in range(N_ITERATIONS):
result = agent.train()
results.append(result)
episode = {'n': n,
'episode_reward_min': result['episode_reward_min'],
'episode_reward_mean': result['episode_reward_mean'],
'episode_reward_max': result['episode_reward_max'],
'episode_len_mean': result['episode_len_mean']}
episode_data.append(episode)
episode_json.append(json.dumps(episode))
file_name = agent.save(CHECKPOINT_ROOT)
print(f'{n:3d}: Min/Mean/Max reward: {result["episode_reward_min"]:8.4f}/{result["episode_reward_mean"]:8.4f}/{result["episode_reward_max"]:8.4f}. Checkpoint saved to {file_name}')
# + colab={"base_uri": "https://localhost:8080/", "height": 359} id="ldRMixTj0TEh" outputId="df55e459-fca6-4c4b-9097-1a323e7880ea"
df = pd.DataFrame(data=episode_data)
df
# + colab={"base_uri": "https://localhost:8080/", "height": 279} id="_Ot6Y5O31FEK" outputId="b6e6fc4d-8218-4e0e-accb-07bac7ae463b"
df.plot(x="n", y=["episode_reward_mean", "episode_reward_min", "episode_reward_max"], secondary_y=True);
# + colab={"base_uri": "https://localhost:8080/"} id="vtP_pjfV1E_c" outputId="0f71afa6-c6e2-4eb4-b47f-c56833f1561c"
import pprint
policy = agent.get_policy()
model = policy.model
pprint.pprint(model.variables())
pprint.pprint(model.value_function())
print(model.base_model.summary())
# + id="0xgrKTeT1Ewu"
# hide
# # tuned training
# # %%time
# stop = {"episode_reward_mean": 195}
# analysis = ray.tune.run(
# "PPO",
# config=config,
# stop=stop,
# checkpoint_at_end=True,
# )
# + [markdown] id="68jNcA_TiJDq"
# ### Visualization after training
#
# + colab={"base_uri": "https://localhost:8080/", "height": 305} id="Fj-jv3IqA4RZ" outputId="a04a11d3-a185-46df-ab50-a2f46467304f"
after_training = "after_training.mp4"
after_video = VideoRecorder(env, after_training)
observation = env.reset()
done = False
while not done:
env.render()
after_video.capture_frame()
action = agent.compute_action(observation)
observation, reward, done, info = env.step(action)
after_video.close()
env.close()
# You should get a video similar to the one below.
html = render_mp4(after_training)
HTML(html)
# + id="Biyz2Wo8A4PV"
# + id="aMVZXsfkA4Lx"
# + id="i4GDOBrHA4IV"
# + id="YFOnCObIA4FB"
# + id="64IHqzVHA37h"
| _notebooks/2021-11-29-RLlib_CartPole.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import random
# 返回一个[0, 1.0)之间的浮点随机数
random.random()
# 返回一个随机浮点数N,当a <= b时,a <= N <= b,否则b <= N <= a。
print random.uniform(1, 11)
# 返回一个随机整数N,并且a <= N <= b。
random.randint(1, 100)
# 随机从根据输入参数生成的序列中选取一个数字返回,并不实际生成序列。
random.randrange(10, 100, 2)
# 从非空序列中随机选择一个元素返回。若序列非空,产生IndexError。
random.choice(range(10))
# 打乱序列的顺序
l = range(10)
random.shuffle(l)
print l
# 从序列中返回k个采样元素,返回元素存储在列表中。
random.sample([1, 1, 1, 2], 2)
import os
print os.urandom(5)
# # 参考文献
# 1. https://docs.python.org/2/library/random.html 官方文档
# 2. http://fulerbakesi.iteye.com/blog/1589097 简单说明
| Python random.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
from astropy.table import Table
import matplotlib.pyplot as plt
import numpy as np
table = Table.read('data/2M19505021+4804508.csv')
# +
from astropy.time import Time
times = Time(table['hjd'], format='jd')
# fluxes = 500 + np.ones(len(times)) + np.random.randn(len(times)) #table['flux (mJy)'].data
# errors = np.ones(len(times)) #5 * table['flux err'].data
fluxes = table['flux (mJy)'].data
errors = table['flux err'].data
#mask = np.ones(len(times)).astype(bool) #(fluxes < 700) & (fluxes > 400)
mask = (fluxes < 700) & (fluxes > 400)
sort = np.argsort(times[mask])
times = times[mask][sort]
fluxes = fluxes[mask][sort]
errors = errors[mask][sort]
# -
np.load('data/spitzer.npy')
spitzer_time, spitzer_flux, spitzer_error = np.load('data/spitzer.npy').T
# +
plt.figure(figsize=(7, 3))
plt.errorbar(times.decimalyear, fluxes, errors, fmt='.', color='k', ecolor='silver')
plt.ylabel('Flux [mJy]')
plt.xlabel('Year')
# plt.ylim([400, 700])
ax = plt.gca()
for s in ['right', 'top']:
ax.spines[s].set_visible(False)
ax.grid(ls='--')
plt.savefig('plots/asas-sn.pdf', bbox_inches='tight')
# +
# def v_vector(theta):
# """
# Hogg+ 2010, Eqn 29.
# """
# return [[-np.sin(theta)], [np.cos(theta)]]
# def lnprior(p, max_theta=1.55, min_theta=1.5, min_lnf=0):
# theta, b, lnf = p
# if not ((min_theta < theta < max_theta) and (-0.5 < b < 0.5) and
# (lnf > min_lnf)):
# return -np.inf
# else:
# return 0
# def ln_likelihood(p, x, y, x_err, y_err):
# """
# Hogg+ 2010, Eqn 30., with an additional parameter that scales up the
# uncertainty in the x dimension, ``x_err``, by a constant factor.
# The likelihood has been written assuming x and y uncertainties are
# uncorrelated.
# """
# # theta, b, lnf, V = p
# theta, b, lnf = p
# # Assert prior:
# # lnf < min_lnf or V < 0
# # if (theta < min_theta or theta > max_theta or b < -0.5 or b > 0.5
# # or lnf < min_lnf):
# v = v_vector(theta)
# f = np.exp(lnf)
# lnp = lnprior(p)
# if not np.isfinite(lnp):
# return lnp
# delta = v[0][0] * x + v[1][0] * y - b * np.cos(theta)
# sigma_sq = v[0][0]**2 * (f * x_err)**2 + v[1][0]**2 * y_err**2
# # sigma_sq = v[0][0]**2 * x_err**2 + v[1][0]**2 * y_err**2
# ln_like = np.sum(-0.5 * (delta**2 / sigma_sq + np.log(sigma_sq) +
# np.log(2*np.pi)))
# return ln_like
# +
import emcee
def model(p, x):
m, b, lnf = p
return m * x + b
def lnprior(p):
# m, b, lnf = p
return 0
def ln_likelihood(p, x, y, y_err):
m, b, lnf = p
f = np.exp(lnf)
return -0.5 * np.sum((model(p, x) - y)**2 / (f * y_err)**2 + np.log(2 * (f * y_err)**2))
from multiprocessing import Pool
nwalkers = 10
ndim = 3
n_steps_burnin = 1000
n_steps_postburnin = 10000
p0 = []
while len(p0) < nwalkers:
trial = np.array([-30, 500, 1]) + 1 * np.random.randn(ndim)
if lnprior(trial) == 0:
p0.append(trial)
args = (times.decimalyear - times.decimalyear.mean(), fluxes, errors)
with Pool() as pool:
sampler = emcee.EnsembleSampler(nwalkers, ndim, ln_likelihood, args=args,
pool=pool)
# Burn in for this many steps:
p1 = sampler.run_mcmc(p0, n_steps_burnin)
sampler.reset()
p2 = sampler.run_mcmc(p1, n_steps_burnin)
sampler.reset()
# Now run for this many more steps:
sampler.run_mcmc(p2, n_steps_postburnin)
samples = sampler.flatchain #sampler.chain[:, :, :].reshape((-1, ndim))
# +
params, resid, rank, singvals = np.linalg.lstsq(np.vander(times.decimalyear - times.decimalyear.mean(), 2), fluxes)
m_init, b_init = params
print(m_init, b_init)
# +
from corner import corner
corner(samples, labels=[r'$m$', '$b$', '$\ln f$'])
plt.show()
# -
spitzer_time
# +
slope = samples[:, 0]
intercept = samples[:, 1]
f = np.exp(samples[:, 2])
# plt.errorbar(times.decimalyear, fluxes, f.mean() * errors, fmt='.', ecolor='silver')
plt.figure(figsize=(7, 3))
plt.errorbar(times.decimalyear, fluxes, f.mean() * errors, fmt='.', color='k', ecolor='silver', label='ASAS-SN')
plt.ylabel('Flux [mJy]')
plt.xlabel('Year')
plt.grid(ls='--')
skip = 1000
for m, b in zip(slope[::skip], intercept[::skip]):
plt.plot(times.decimalyear, m*(times.decimalyear - times.decimalyear.mean()) + b, alpha=0.1, color='r', zorder=-10)
# plt.ylim([400, 700])
plt.legend(loc='upper left')
ax2 = plt.gca().twinx()
ax2.errorbar(spitzer_time, spitzer_flux, spitzer_error, fmt='s', color='DodgerBlue', ecolor='silver', label='Spitzer')
ax2.set_ylabel('Flux [DN/s]')
plt.legend(loc='upper right')
plt.savefig('plots/asas-sn.pdf', bbox_inches='tight')
# +
from astropy.coordinates import SkyCoord
hat11 = SkyCoord.from_name('HAT-P-11')
c1 = SkyCoord.from_name('GAIA DR2 2086521642419240960')
c2 = SkyCoord.from_name('GAIA DR2 2086521298821848832')
# -
control1 = Table.read('data/64348.csv') # GAIA DR2 2086521642419240960
control2 = Table.read('data/64655.csv') # GAIA DR2 2086521298821848832
# +
plt.figure(figsize=(4, 3))
plt.plot(Time(control1['hjd'], format='jd').decimalyear, control1['flux (mJy)'], '.',
label="{0:.1f}".format(c1.separation(hat11).to(u.arcmin)))
plt.plot(Time(control2['hjd'], format='jd').decimalyear, control2['flux (mJy)'], '.',
label="{0:.1f}".format(c2.separation(hat11).to(u.arcmin)))
plt.legend()
ax = plt.gca()
for s in ['right', 'top']:
ax.spines[s].set_visible(False)
plt.ylabel('Flux [mJy]')
plt.xlabel('Year')
ax.grid(ls='--')
plt.ylim([30, 85])
plt.savefig('plots/control.pdf', bbox_inches='tight')
# -
| asas-sn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
os.chdir('../')
from rsna_retro.imports import *
from rsna_retro.metadata import *
from rsna_retro.preprocess import *
from rsna_retro.train import *
from rsna_retro.train3d import *
torch.cuda.set_device(1)
path_appfeat512 = path/'appian_features_512'
path_appfeat512_tst = path/'appian_tst_features_512'
dls_feat = get_3d_dls_feat(Meta.df_comb, path=path_appfeat512, bs=32, meta=True)
type(dls_feat.train.dataset.)
xb,yb = dls_feat.one_batch()
# ## Model
# +
class NeuralNet(nn.Module):
def __init__(self, n_classes=6, embed_size=1024*4+1, LSTM_UNITS=1024*4+1, DO = 0.3):
super(NeuralNet, self).__init__()
# self.embedding_dropout = SpatialDropout(0.0) #DO)
self.flat = nn.Sequential(AdaptiveConcatPool2d(), Flatten())
self.hook = ReshapeBodyHook(self.flat)
self.lstm1 = nn.LSTM(embed_size, LSTM_UNITS, bidirectional=True, batch_first=True)
self.lstm2 = nn.LSTM(LSTM_UNITS * 2, LSTM_UNITS, bidirectional=True, batch_first=True)
self.linear1 = nn.Linear(LSTM_UNITS*2, LSTM_UNITS*2)
self.linear2 = nn.Linear(LSTM_UNITS*2, LSTM_UNITS*2)
self.linear = nn.Linear(LSTM_UNITS*2, n_classes)
def forward(self, x):
# x = torch.cat(x, axis=-1)
x,pos = x
x = x.view(*x.shape[:2], -1, 4, 4)
h_embedding = torch.cat([self.flat(x), pos], axis=-1)
# print(h_embedding.shape)
h_embadd = torch.cat((h_embedding, h_embedding), -1)
h_lstm1, _ = self.lstm1(h_embedding)
h_lstm2, _ = self.lstm2(h_lstm1)
h_conc_linear1 = F.relu(self.linear1(h_lstm1))
h_conc_linear2 = F.relu(self.linear2(h_lstm2))
# print([x.shape for x in [h_lstm1, h_lstm2, h_conc_linear1, h_conc_linear2, h_embadd]])
hidden = h_lstm1 + h_lstm2 + h_conc_linear1 + h_conc_linear2 + h_embadd
output = self.linear(hidden)
return output
# -
m = NeuralNet()
name = 'train3d_adj_feat_lstm_2ndplace_meta'
learn = get_learner(dls_feat, m, name=name)
learn.add_cb(DePadLoss())
# +
# learn.summary()
# -
# ## Training
learn.lr_find()
do_fit(learn, 10, 1e-3)
learn.save(f'runs/{name}-1')
learn.load(f'runs/{name}-1')
do_fit(learn, 4, 1e-4)
learn.save(f'runs/{name}-2')
# ## Testing
learn.dls = get_3d_dls_feat(Meta.df_tst, path=path_appfeat512_tst, bs=32, test=True, meta=True)
sub_fn = f'subm/{name}'
learn.load(f'runs/{name}-1')
preds,targs = learn.get_preds()
preds.shape, preds.min(), preds.max()
pred_csv = submission(Meta.df_tst, preds, fn=sub_fn)
api.competition_submit(f'{sub_fn}.csv', name, 'rsna-intracranial-hemorrhage-detection')
api.competitions_submissions_list('rsna-intracranial-hemorrhage-detection')[0]
| 03_train3d_experiments/03_train3d_04_train_appian_lstm_head_2ndPlace_meta.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Interacting with Python
#
# __Content modified under Creative Commons Attribution license CC-BY 4.0,
# code under BSD 3-Clause License © 2020 <NAME>__
# These notebooks are a combination of original work and modified notebooks from [Engineers Code](https://github.com/engineersCode/EngComp.git) learning modules. The learning modules are covered under a Creative Commons License, so we can modify and publish *and give credit to <NAME> and <NAME>*.
#
# Our first goal is to interact with Python and handle data in Python.
# But let's also learn a little bit of background.
# + tags=["hide-cell"]
import numpy as np
import matplotlib.pyplot as plt
plt.style.use('fivethirtyeight')
# -
# ## What is Python?
#
# Python was born in the late 1980s. Its creator, [Guido van Rossum](https://en.wikipedia.org/wiki/Guido_van_Rossum), named it after the British comedy "Monty Python's Flying Circus." His goal was to create "an easy and intuitive language just as powerful as major competitors," producing computer code "that is as understandable as plain English."
#
# We say that Python is a _general-purpose_ language, which means that you can use it for anything: organizing data, scraping the web, creating websites, analyzing sounds, creating games, and of course _engineering computations_.
#
# Python is an _interpreted_ language. This means that you can write Python commands and the computer can execute those instructions directly. Other programming languages—like C, C++ and Fortran—require a previous _compilation_ step: translating the commands into machine language.
# A neat ability of Python is to be used _interactively_. [<NAME>](https://en.wikipedia.org/wiki/Fernando_Pérez_(software_developer) famously created **IPython** as a side-project during his PhD. The "I" in IPython stands for interactive: a style of computing that is very powerful for developing ideas and solutions incrementally, thinking with the computer as a kind of collaborator.
# ## Why Python?
#
#
# _Because it's fun!_ With Python, the more you learn, the more you _want_ to learn.
# You can find lots of resources online and, since Python is an open-source project, you'll also find a friendly community of people sharing their knowledge.
# _And it's free!_
#
# Python is known as a _high-productivity language_. As a programmer, you'll need less time to develop a solution with Python than with most other languages.
# This is important to always bring up whenever someone complains that "Python is slow."
# Your time is more valuable than a machine's!
# (See the Recommended Readings section at the end of this lesson.)
# And if we really need to speed up our program, we can re-write the slow parts in a compiled language afterwards.
# Because Python plays well with other languages :–)
#
# The top technology companies use Python: Google, Facebook, Dropbox, Wikipedia, Yahoo!, YouTube… Python took the No. 1 spot in the interactive list of [The 2017 Top Programming Languages](http://spectrum.ieee.org/computing/software/the-2017-top-programming-languages), by _IEEE Spectrum_ ([IEEE](http://www.ieee.org/about/index.html) is the world's largest technical professional society).
# #### _Python is a versatile language, you can analyze data, build websites (e.g., Instagram, Mozilla, Pinterest), make art or music, etc. Because it is a versatile language, employers love Python: if you know Python they will want to hire you._ —<NAME>, ex Director of the Python Software Foundation, in a [2014 tutorial](https://youtu.be/rkx5_MRAV3A).
# ## Let's get started
#
# You could follow this first lesson using IPython. If you have it installed in the computer you're using, you enter the program by typing `ipython` on the command-line interface (the **Terminal** app on Mac OSX, and on Windows the **PowerShell** or a similar app).
# A free service to try IPython online, right from your browser, is [Python Anywhere](https://www.pythonanywhere.com/try-ipython/). You can execute all the examples of this lesson in IPython using this service.
#
# You can also use Jupyter: an environment that combines programming with other content, like text and images, to form a "computational narrative." All of these lessons are written in Jupyter notebooks.
#
# For this lesson, we will assume you have been guided to open a blank Jupyter notebook, or are working interactively with this lesson.
# On a blank Jupyter notebook, you should have in front of you the input line counter:
#
# `In[1]:`
#
# That input line is ready to receive any Python code to be executed interactively. The output of the code will be shown to you next to `Out[1]`, and so on for successive input/output lines in IPython, or code cells in Jupyter.
# ### Your first program
#
# In every programming class ever, your first program consists of printing a _"Hello"_ message. In Python, you use the `print()` function, with your message inside quotation marks.
print("Hello world!!")
# Easy peasy!! You just wrote your first program and you learned how to use the `print()` function. Yes, `print()` is a function: we pass the _argument_ we want the function to act on, inside the parentheses. In the case above, we passed a _string_, which is a series of characters between quotation marks. Don't worry, we will come back to what strings are later on in this lesson.
#
# ##### Key concept: function
#
# A function is a compact collection of code that executes some action on its _arguments_. Every Python function has a _name_, used to call it, and takes its arguments inside round brackets. Some arguments may be optional (which means they have a default value defined inside the function), others are required. For example, the `print()` function has one required argument: the string of characters it should print out for you.
#
# Python comes with many _built-in_ functions, but you can also build your own. Chunking blocks of code into functions is one of the best strategies to deal with complex programs. It makes you more efficient, because you can reuse the code that you wrote into a function. Modularity and reuse are every programmer's friends.
# ### Python as a calculator
#
# Try any arithmetic operation in IPython or a Jupyter code cell. The symbols are what you would expect, except for the "raise-to-the-power-of" operator, which you obtain with two asterisks: `**`. Try all of these:
#
# ```python
# + - * / ** % //
# ```
#
# The `%` symbol is the _modulo_ operator (divide and return the remainder), and the double-slash is _floor division_.
2 + 2
1.25 + 3.65
5 - 3
2 * 4
7 / 2
2**3
# Let's see an interesting case:
9**1/2
# ##### Discuss with your neighbor:
# _What happened?_ Isn't $9^{1/2} = 3$? (Raising to the power $1/2$ is the same as taking the square root.) Did Python get this wrong?
#
# Compare with this:
9**(1/2)
# Yes! The order of operations matters!
#
# If you don't remember what we are talking about, review the [Arithmetics/Order of operations](https://en.wikibooks.org/wiki/Arithmetic/Order_of_Operations). A frequent situation that exposes this is the following:
3 + 3 / 2
(3 + 3) / 2
# In the first case, we are adding $3$ plus the number resulting of the operation $3/2$. If we want the division to apply to the result of $3+3$, we need the parentheses.
# ##### Exercises:
# Use Python (as a calculator) to solve the following two problems:
#
# 1. The volume of a sphere with radius $r$ is $\frac{4}{3}\pi r^3$. What is the volume of a sphere with diameter 6.65 cm?
#
# For the value of $\pi$ use 3.14159 (for now). Compare your answer with the solution up to 4 decimal numbers.
#
# Hint: 523.5983 is wrong and 615.9184 is also wrong.
#
# 2. Suppose the cover price of a book is $\$ 24.95$, but bookstores get a $40\%$ discount. Shipping costs $\$3$ for the first copy and $75$ cents for each additional copy. What is the total wholesale cost for $60$ copies? Compare your answer with the solution up to 2 decimal numbers.
#
# To reveal the answers, highlight the following line of text using the mouse:
# Answer exercise 1: <span style="color:white"> 153.9796 </span> Answer exercise 2: <span style="color:white"> 945.45 </span>
# ### Variables and their type
#
# Variables consist of two parts: a **name** and a **value**. When we want to give a variable its name and value, we use the equal sign: `name = value`. This is called an _assignment_. The name of the variable goes on the left and the value on the right.
#
# The first thing to get used to is that the equal sign in a variable assignment has a different meaning than it has in Algebra! Think of it as an arrow pointing from `name` to `value`.
#
#
# <img src="./images/variables.png" style="width: 400px;"/>
#
# We have many possibilities for variable names: they can be made up of upper and lowercase letters, underscores and digits… although digits cannot go on the front of the name. For example, valid variable names are:
#
# ```python
# x
# x1
# X_2
# name_3
# NameLastname
# ```
# Keep in mind, there are reserved words that you can't use; they are the special Python [keywords](https://docs.python.org/3/reference/lexical_analysis.html#keywords).
#
# OK. Let's assign some values to variables and do some operations with them:
x = 3
y = 4.5
# ##### Exercise:
# Print the values of the variables `x` and `y`.
# Let's do some arithmetic operations with our new variables:
x + y
2**x
y - 3
# And now, let's check the values of `x` and `y`. Are they still the same as they were when you assigned them?
#
print(x)
print(y)
# ### String variables
#
# In addition to name and value, Python variables have a _type_: the type of the value it refers to. For example, an integer value has type `int`, and a real number has type `float`. A string is a variable consisting of a sequence of characters marked by two quotes, and it has type `str`.
#
z = 'this is a string'
w = '1'
# What if you try to "add" two strings?
z + w
# The operation above is called _concatenation_: chaining two strings together into one. Insteresting, eh? But look at this:
#
# `>> x + w`
#
# _Error!_ Why? Let's inspect what Python has to say and explore what is happening.
#
# Python is a _dynamic language_, which means that you don't _need_ to specify a type to invoke an existing object. The humorous nickname for this is "duck typing":
#
# #### "If it looks like a duck, and quacks like a duck, then it's probably a duck."
#
# In other words, a variable has a type, but we don't need to specify it. It will just behave like it's supposed to when we operate with it (it'll quack and walk like nature intended it to).
#
# But sometimes you need to make sure you know the type of a variable. Thankfully, Python offers a function to find out the type of a variable: `type()`.
type(x)
type(w)
type(y)
# ### More assignments
#
# Here we assign a new variable to the result of an operation that involves other variables.
sum_xy = x + y
diff_xy = x - y
print('The sum of x and y is:', sum_xy)
print('The difference between x and y is:', diff_xy)
# Notice what we did above: we used the `print()` function with a string message, followed by a variable, and Python printed a useful combination of the message and the variable value. This is a pro tip! You want to print for humans. Let's now check the type of the new variables we just created above:
type(sum_xy)
type(diff_xy)
# ## Reflection point
# When we created `sum_xy` and `diff_xy` two new variables were created that depended upon previously created variables `x` and `y`. How else can we accomplish this? Could we make a function? Could we combine the commands in one block as a script?
# ### Special variables
#
# Python has special variables that are built into the language. These are:
# `True`, `False`, `None` and `NotImplemented`.
# For now, we will look at just the first three of these.
#
# **Boolean variables** are used to represent truth values, and they can take one of two possible values: `True` and `False`.
# _Logical expressions_ return a boolean. Here is the simplest logical expression, using the keyword `not`:
#
# ```Python
# not True
# ```
#
# It returns… you guessed it… `False`.
#
# The Python function `bool()` returns a truth value assigned to any argument. Any number other than zero has a truth value of `True`, as well as any nonempty string or list. The number zero and any empty string or list will have a truth value of `False`. Explore the `bool()` function with various arguments.
#
bool(0)
bool('Do we need oxygen?')
bool('We do need oxygen')
# **None is not Zero**: `None` is a special variable indicating that no value was assigned or that a behavior is undefined. It is different than the value zero, an empty string, or some other nil value.
#
# You can check that it is not zero by trying to add it to a number. Let's see what happens when we try that:
# +
a = None
b = 3
# -
# ```python
# >>> a + b
#
# ---------------------------------------------------------------------------
# TypeError Traceback (most recent call last)
# <ipython-input-71-ca730b97bf8a> in <module>
# ----> 1 a+b
#
# TypeError: unsupported operand type(s) for +: 'NoneType' and 'int'
# ```
#
# ### Logical and comparison operators
#
# The Python comparison operators are: `<`, `<=`, `>`, `>=`, `==`, `!=`. They compare two objects and return either `True` or `False`: smaller than, smaller or equal, greater than, greater or equal, equal, not equal. Try it!
x = 3
y = 5
x > y
# We can assign the truth value of a comparison operation to a new variable name:
z = x > y
z
type(z)
# Logical operators are the following: `and`, `or`, and `not`. They work just like English (with the added bonus of being always consistent, not like English speakers!). A logical expression with `and` is `True` if both operands are true, and one with `or` is `True` when either operand is true. And the keyword `not` always negates the expression that follows.
#
# Let's do some examples:
a = 5
b = 3
c = 10
a > b and b > c
# Remember that the logical operator `and` is `True` only when both operands are `True`. In the case above the first operand is `True` but the second one is `False`.
#
# If we try the `or` operation using the same operands we should get a `True`.
a > b or b > c
# And the negation of the second operand results in …
not b > c
# What if we negate the second operand in the `and` operation above?
#
# ##### Note:
#
# Be careful with the order of logical operations. The order of precedence in logic is:
#
# 1. Negation
# 2. And
# 3. Or
#
# If you don't rememeber this, make sure to use parentheses to indicate the order you want.
# ##### Exercise:
#
# What is happening in the case below? Play around with logical operators and try some examples.
a > b and not b > c
# ## What we've learned
#
# * Using the `print()` function. The concept of _function_.
# * Using Python as a calculator.
# * Concepts of variable, type, assignment.
# * Special variables: `True`, `False`, `None`.
# * Supported operations, logical operations.
# * Reading error messages.
# ## References
#
# Throughout this course module, we will be drawing from the following references:
#
# 1. _Effective Computation in Physics: Field Guide to Research with Python_ (2015). <NAME> & <NAME>. O'Reilly Media, Inc.
# 2. _Python for Everybody: Exploring Data Using Python 3_ (2016). <NAME>. [PDF available](http://do1.dr-chuck.com/pythonlearn/EN_us/pythonlearn.pdf)
# 3. _Think Python: How to Think Like a Computer Scientist_ (2012). <NAME>. Green Tea Press. [PDF available](http://greenteapress.com/thinkpython/thinkpython.pdf)
# ## Problems
#
# __1.__ Calculate some properties of a rectangular box that is 12.5"$\times$11"$\times$14" and weighs 31 lbs
#
# a. What is the volume of the box?
#
# b. What is the average density of the box?
#
# c. What is the result of the following logical operation, `volume>1000` (in inches^3)
#
#
#
# __2.__ Use the variables given below, `str1` and `str2`, and check the following
#
# a. `str1<str2`
#
# b. `str1==str2`
#
# c. `str1>str2`
#
# d. How could you force (b) to be true? [Hint](https://docs.python.org/3/library/stdtypes.html?highlight=str.lower#str.lower) or [Hint](https://docs.python.org/3/library/stdtypes.html?highlight=str.lower#str.upper)
str1 = 'Python'
str2 = 'python'
# __3.__ The following code has an error, fix the error so that the correct result is returned:
#
# ```y is 20 and x is less than y```
#
# ```python jupyter={"source_hidden": true}
# x = '1'
# y = 20
#
# if x<y and y==20:
# print('y is 20 and x is less than y')
# else:
# print('x is not less than y')
# ```
# __4.__ Write a script using `if-else` that returns `g=9.81` if `unit = 'kgms'` and `g=32.2` if `unit = 'lbfts'`:
unit = 'kgms'
| getting-started/01_interactions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # AI4M Course 1 week 3 lecture notebook
# # Extract a sub-section
#
# In the assignment you will be extracting sub-sections of the MRI data to train your network. The reason for this is that training on a full MRI scan would be too memory intensive to be practical. To extract a sub-section in the assignment, you will need to write a function to isolate a small "cube" of the data for training. This example is meant to show you how to do such an extraction for 1D arrays. In the assignment you will apply the same logic in 3D.
import numpy as np
import keras
import pandas as pd
# Define a simple one dimensional "image" to extract from
image = np.array([10,11,12,13,14,15])
image
# Compute the dimensions of your "image"
image_length = image.shape[0]
image_length
# ### Sub-sections
# In the assignment, you will define a "patch size" in three dimensions, that will be the size of the sub-section you want to extract. For this exercise, you only need to define a patch size in one dimension.
# Define a patch length, which will be the size of your extracted sub-section
patch_length = 3
# To extract a patch of length `patch_length` you will first define an index at which to start the patch.
#
# Run the next cell to define your start index
# Define your start index
start_i = 0
# At the end of the next cell you are adding 1 to the start index. Run cell a few times to extract some one dimensional sub-sections from your "image"
#
# What happens when you run into the edge of the image (when `start_index` is > 3)?
# +
# Define an end index given your start index and patch size
print(f"start index {start_i}")
end_i = start_i + patch_length
print(f"end index {end_i}")
# Extract a sub-section from your "image"
sub_section = image[start_i: end_i]
print("output patch length: ", len(sub_section))
print("output patch array: ", sub_section)
# Add one to your start index
start_i +=1
# -
# You'll notice when you run the above multiple times, that eventually the sub-section returned is no longer of length `patch_length`.
#
# In the assignment, your neural network will be expecting a particular sub-section size and will not accept inputs of other dimensions. For the start indices, you will be randomly choosing values and you need to ensure that your random number generator is set up to avoid the edges of your image object.
#
# The next few code cells include a demonstration of how you could determine the constraints on your start index for the simple one dimensional example.
# Set your start index to 3 to extract a valid patch
start_i = 3
print(f"start index {start_i}")
end_i = start_i + patch_length
print(f"end index {end_i}")
sub_section = image[start_i: end_i]
print("output patch array: ", sub_section)
# Compute and print the largest valid value for start index
print(f"The largest start index for which "
f"a sub section is still valid is "
f"{image_length - patch_length}")
# +
# Compute and print the range of valid start indices
print(f"The range of valid start indices is:")
# Compute valid start indices, note the range() function excludes the upper bound
valid_start_i = [i for i in range(image_length - patch_length + 1)]
print(valid_start_i)
# -
# ### Random selection of start indices
# In the assignment, you will need to randomly select a valid integer for the start index in each of three dimensions. The way to do this is by following the logic above to identify valid start indices and then selecting randomly from that range of valid numbers.
#
# Run the next cell to select a valid start index for the one dimensional example
# Choose a random start index, note the np.random.randint() function excludes the upper bound.
start_i = np.random.randint(image_length - patch_length + 1)
print(f"randomly selected start index {start_i}")
# Randomly select multiple start indices in a loop
for _ in range(10):
start_i = np.random.randint(image_length - patch_length + 1)
print(f"randomly selected start index {start_i}")
# ### Background Ratio
#
# Another thing you will be doing in the assignment is to compute the ratio of background to edema and tumorous regions. You will be provided with a file containing labels with these categories:
#
# * 0: background
# * 1: edema
# * 2: non-enhancing tumor
# * 3: enhancing tumor
#
# Let's try to demonstrate this in 1-D to get some intuition on how to implement it in 3D later in the assignment.
# +
# We first simulate input data by defining a random patch of length 16. This will contain labels
# with the categories (0 to 3) as defined above.
patch_labels = np.random.randint(0, 4, (16))
print(patch_labels)
# +
# A straightforward approach to get the background ratio is
# to count the number of 0's and divide by the patch length
bgrd_ratio = np.count_nonzero(patch_labels == 0) / len(patch_labels)
print("using np.count_nonzero(): ", bgrd_ratio)
bgrd_ratio = len(np.where(patch_labels == 0)[0]) / len(patch_labels)
print("using np.where(): ", bgrd_ratio)
# +
# However, take note that we'll use our label array to train a neural network
# so we can opt to compute the ratio a bit later after we do some preprocessing.
# First, we convert the label's categories into one-hot format so it can be used to train the model
patch_labels_one_hot = keras.utils.to_categorical(patch_labels, num_classes=4)
print(patch_labels_one_hot)
# -
# **Note**: We hardcoded the number of classes to 4 in our simple example above.
# In the assignment, you should take into account that the label file can have
# a different number of categories
# +
# Let's convert the output to a dataframe just so we can see the labels more clearly
pd.DataFrame(patch_labels_one_hot, columns=['background', 'edema', 'non-enhancing tumor', 'enhancing tumor'])
# +
# What we're interested in is the first column because that
# indicates if the element is part of the background
# In this case, 1 = background, 0 = non-background
print("background column: ", patch_labels_one_hot[:,0])
# +
# we can compute the background ratio by counting the number of 1's
# in the said column divided by the length of the patch
bgrd_ratio = np.sum(patch_labels_one_hot[:,0])/ len(patch_labels)
print("using one-hot column: ", bgrd_ratio)
# -
# #### That's all for this lab, now you have the basic tools you need for sub-section extraction in this week's graded assignment!
| AI for Medical Diagnosis/Week 3/AI4M_C1_W3_lecture_ex_02.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="y77NH3fZc7ZX"
# # Kaggle
# + colab={"base_uri": "https://localhost:8080/"} id="L9KKRFAcCi7q" outputId="fd8b52f6-5ecb-4d9f-d660-da0957165230"
# ! pip install --upgrade --force-reinstall --no-deps kaggle
# + colab={"base_uri": "https://localhost:8080/", "height": 39, "resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY>", "headers": [["content-type", "application/javascript"]], "ok": true, "status": 200, "status_text": "OK"}}} id="dPdtNDuSDCZ-" outputId="50652024-0070-4adb-b8f8-27f2d8980df6"
from google.colab import files
files.upload()
# + id="_xdoXQzcDDp4"
# ! mkdir ~/.kaggle
# ! cp kaggle.json ~/.kaggle/
# + id="W-Pe-Cp8DlBM"
# ! chmod 600 ~/.kaggle/kaggle.json
# + colab={"base_uri": "https://localhost:8080/"} id="09JoO7QSDo6G" outputId="57b11b96-7ac3-4f86-fe4f-ab570132f605"
# ! kaggle datasets download -d tongpython/cat-and-dog
# + colab={"base_uri": "https://localhost:8080/"} id="ftMSfyI4DuLt" outputId="f39beba2-3e56-4b3e-fee7-5af006ee326b"
# ! unzip cat-and-dog.zip -d dataset
# + id="SiaO3SMHJiav"
# + [markdown] id="c7774n6jCk28"
# # Transfer Learning
# + id="frYItZ93CnE2"
import tensorflow as tf
import matplotlib.pyplot as plt
import numpy as np
# + id="HZM-Z8F1DDb_"
training_path = "/content/dataset/training_set/training_set"
test_path = "/content/dataset/test_set/test_set"
# + colab={"base_uri": "https://localhost:8080/"} id="f9OkUYlFCx7N" outputId="94b426f7-7569-44b5-d0af-a6b8bf71cdc8"
#Define augmentation
dataGen = tf.keras.preprocessing.image.ImageDataGenerator(
rescale = 1./255,
rotation_range=5,
horizontal_flip=True,
vertical_flip=True,
)
train_dataset_aug = dataGen.flow_from_directory(
training_path,
target_size = (224,224),
batch_size = 32,
shuffle=True,
seed = 865
)
test_dataset_aug = dataGen.flow_from_directory(
test_path,
target_size = (224,224),
batch_size = 32,
shuffle=True,
seed = 865
)
# + colab={"base_uri": "https://localhost:8080/"} id="UT36AgLBETBT" outputId="d242331b-2a0e-4717-9045-fa8c2d9b3d83"
# pretrained_model = tf.keras.applications.MobileNetV2(input_shape=(224,224,3),
# include_top=False,
# weights='imagenet')
pretrained_model = tf.keras.applications.ResNet50(
include_top=False,
weights="imagenet",
input_shape=(224,224,3),
)
pre_model = tf.keras.applications.InceptionResNetV2(
include_top=False, weights='imagenet',
input_shape=(224,224,3),
# + id="WOa6HHLWEkGY"
pretrained_model.trainable = False
# + id="PUufrqbbEu1J"
pretrained_model.summary()
# + id="h8yLCAgeFgAF"
model = tf.keras.models.Sequential()
# + id="qQYkvEprGtH2"
model.add(pretrained_model)
# + id="vj14UZvyG0SS"
model.add(tf.keras.layers.GlobalAveragePooling2D())
# + id="HMza7fVPG52f"
model.add(tf.keras.layers.Dense(1024, activation='relu'))
model.add(tf.keras.layers.Dense(512, activation='relu'))
model.add(tf.keras.layers.Dense(2, activation='softmax'))
# + id="YwirSWZ_II9C"
model.compile(optimizer = 'adam',
loss='categorical_crossentropy',metrics = ['accuracy'])
# + colab={"background_save": true, "base_uri": "https://localhost:8080/"} id="Ig-IUQZpIOZK" outputId="4badffdd-a45d-4b86-8973-f6da94f17d15"
history = model.fit(train_dataset_aug, validation_data=test_dataset_aug, epochs = 15)
# + colab={"base_uri": "https://localhost:8080/", "height": 300} id="1kSZuwqfISOQ" outputId="cb5c08e8-055c-42fb-9d4d-d178f773eef1"
plt.plot(history.history['accuracy'], label='accuracy')
plt.plot(history.history['val_accuracy'], label = 'val_accuracy')
plt.xlabel('Epoch')
plt.ylabel('Accuracy')
plt.ylim([0.9, 1])
plt.legend(loc='lower right')
# + id="-COUSELRI5nq"
| resources/PESU-IO Slot 11 Deep Learning Course material/Week3_I_O.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Daniel-ASG/Aulas_de_cursos/blob/main/Clustering_B%C3%A1sico_k_means%2C_DBSCAN_e_mean_shift.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="LhnjAriCJ1V8"
# # Aula 1 - Análise exploratória
# + [markdown] id="Ifh4YGaFJ5GV"
# Imagine que você está trabalhando como cientista de dados em uma dessas empresas grandes de logística. Nesse caso, seu trabalho é organizar dados!
#
# Seu chefe então te mandou um conjunto de vinhos e pediu para que você os organizasse.
#
#
# + id="Ir22bVhnJhAx"
import plotly.graph_objects as go
import plotly.express as px
import pandas as pd
import numpy as np
# + colab={"base_uri": "https://localhost:8080/", "height": 255} id="R9gwBjTaKmR7" outputId="8e4af58a-dcc9-4f05-b7a5-88eb7125b778"
df = pd.read_csv('https://raw.githubusercontent.com/alura-cursos/Clustering/Aula-1.-An%C3%A1lise-Explorat%C3%B3ria/Wine.csv')
print(f'O dataframe possui {df.shape[0]} amostras e {df.shape[1]} atributos.\n\n')
display(df.head())
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="sgdL_ey_LXMM" outputId="3c16edd8-9126-4ab9-b9d4-4e1c18b8dc78"
df.rename(columns={'Alcohol': 'Alcool',
'Ash': 'Po',
'Ash_Alcanity': 'Alcalinidade_po',
'Magnesium': 'Magnesio',
'Color_Intensity': 'Intensidade_de_cor'
}, inplace=True)
df.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 297} id="fg1qGsiZMSQg" outputId="5fad58a6-db09-441b-833a-2cf7c5dd718c"
df.describe()
# + [markdown] id="5c7XGVfdTMoH"
# Vamos saber a relação que existe entre os atributos do dataframe, para saber quais são realmente fundamentais para o seu DataFrame e quais não são.
# + id="2_pvMrqPMnHc" colab={"base_uri": "https://localhost:8080/"} outputId="318bda97-cbc7-4c7d-de2b-43ada6c42b15"
pip install biokit
# + id="CWO05L-RTZxf" colab={"base_uri": "https://localhost:8080/"} outputId="738887cd-a645-48e6-d9a9-5a68293e2292"
from biokit.viz import corrplot
import matplotlib.pyplot as plt
# + colab={"base_uri": "https://localhost:8080/", "height": 483} id="RPw2o5b-UqiC" outputId="121931d0-b369-4711-f452-b348db0e6622"
matriz_corr = df.corr()
matriz_corr
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="zpa6j1WqTsMQ" outputId="ed0c30fc-a64c-4b93-a4b5-29de91b23541"
corr_graf = corrplot.Corrplot(matriz_corr)
corr_graf.plot(upper='ellipse', fontsize='x-large')
fig = plt.gcf()
fig.set_size_inches(20,8);
# + [markdown] id="u5iunKs4VlwV"
# Chegamos a algumas conclusões:
# * “Total_Fenois” é bastante correlacionado com “Flavanoids”
# * A correlação entre “Hue” e “Malic_Acid” é uma correlação negativa
#
# Isso é muito bom quando nós queremos remover alguns atributos de informação redundante. Nós poderíamos remover um deles e deixar o nosso DataFrame menor, com isso nós necessitaríamos de menos espaço para armazenar os nossos dados e também poderíamos utilizar modelos de inteligência artificial menor.
# + [markdown] id="oPqodtbuWMdn"
# Como nós podemos perceber, os atributos dos nossos dados variam diferentemente.
#
# A maioria dos modelos de inteligência artificial e clusterização não trabalham bem com dados que variam de maneira diferente. Para nós resolvermos esse problema vai ser necessário colocarmos os nossos atributos variando na mesma faixa, ou seja, de 0 a 1 e de -1 a 1 através de um processo chamado normalização.
#
# Nós iremos utilizar uma forma de normalização chamado “MinMaxScaler”, ou seja, normalização pelo mínimo e pelo máximo.
# + id="ZeY20oyuUQkm"
from sklearn import preprocessing
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="4IL-0gQAXcfJ" outputId="45bf8f82-3cd1-4a90-8ab4-23c45c976a31"
min_max_scaler = preprocessing.MinMaxScaler()
d = min_max_scaler.fit_transform(df)
df_normalized = pd.DataFrame(d, columns=df.columns)
df_normalized.head()
# + [markdown] id="h9XtbAg6aP6M"
# Como nós podemos perceber agora, todos os atributos das nossas amostras estão variando de 0 a 1.
# + colab={"base_uri": "https://localhost:8080/", "height": 297} id="-tprJufyXqGH" outputId="63062b19-6685-4480-c1de-eaac04ef4b83"
df_normalized.describe()
# + [markdown] id="m09kA1xgcljP"
# Podemos fazer a transformação inversa e obter o dataframe original a partir do normalizado.
# + colab={"base_uri": "https://localhost:8080/", "height": 535} id="UalKl7pwaS7X" outputId="2e900389-2381-46ef-9cc4-6be8eff39d4f"
d = min_max_scaler.inverse_transform(df_normalized)
df = pd.DataFrame(d, columns=df_normalized.columns)
display(df.head())
print('\n\n')
display(df.describe())
# + [markdown] id="TSQCBfJMdM7L"
# # Aula 2 - K-means
#
# O método de clusterização K-means classifica os objetos dentro de múltiplos grupos, de forma que a variação intra-cluster seja minimizada pela soma dos quadrados das distâncias Euclidianas entre os itens e seus centroides.
#
# Os pontos de dados dentro de um cluster são homogêneas e heterogêneas para grupos de pares.
#
# Como K-means faz os clusters:
#
# 1. K-means pega número k de pontos para cada grupo conhecido como centróide.
# 2. Cada ponto de dados forma um cluster com o centróide mais próximo, i.e. k aglomerados.
# 3. Encontra-se o centróide de cada cluster com base em membros do cluster existente. Aqui temos novos centróides.
# 4. Em tendo novos centróides, repita o passo 2 e 3. Encontre a distância mais próxima para cada ponto de dados a partir de novos centróides e os associe com as novas-k clusters. Repita este processo até que a convergência ocorra, isto é, os centróides não mudem.
# + id="cVEltMj3cMAW"
from sklearn.cluster import KMeans
# + colab={"base_uri": "https://localhost:8080/"} id="xqOOBEawdiBV" outputId="9e3bd392-0f5c-48f7-98b3-6c8e71157fe7"
agrupador = KMeans(n_clusters=4)
agrupador.fit(df)
# + colab={"base_uri": "https://localhost:8080/"} id="p-hExf22dvBd" outputId="868cc388-56ca-4d8f-be70-8e98078bcd05"
agrupador.fit(df)
labels = agrupador.labels_
print(labels)
# + colab={"base_uri": "https://localhost:8080/"} id="oxszDOxPd3TM" outputId="e7c88226-0585-4813-befe-c8e6c4996312"
agrupador.fit(df)
labels = agrupador.labels_
print(labels)
# + [markdown] id="hwAZUTKZaP4d"
# A visualização dos clusters na forma de lista não ajuda muito a perceber a distribuição dos dados.
#
# Para facilitar isso iremos visualizar esses clusters num gráfico 2D.
# + colab={"base_uri": "https://localhost:8080/"} id="Dk-tX0UdYQwr" outputId="1ff4d2fe-49de-4c19-af41-2d6069ff36c3"
df_normalized['labels'] = labels
df_normalized.head()
# + colab={"base_uri": "https://localhost:8080/"} id="M1T40iMYVNOg" outputId="4a98d1a9-9b78-408a-a002-c87c416ad922"
fig = px.scatter(df_normalized, x='Intensidade_de_cor', y='Alcool', color='labels', width=800, height=800)
fig.show()
# + [markdown] id="X8UajEV9cqsu"
# Agora podemos perceber cada grupo representado por sua respectiva cor.
#
# Vamos testar a clusterização com outra quantidade de clusters.
# + colab={"base_uri": "https://localhost:8080/"} id="JiqRBAwxZTLi" outputId="2adc53a4-b475-4bcb-f196-31c3136bb051"
df_normalized.drop('labels', axis=1, inplace=True)
agrupador = KMeans(n_clusters=3)
agrupador.fit(df_normalized)
labels = agrupador.labels_
df_normalized['labels'] = labels
print(labels)
# + colab={"base_uri": "https://localhost:8080/"} id="sVWDarT5dml2" outputId="c4c36a7b-6cbd-4e2c-9cc3-7c63aaf32b8a"
fig = px.scatter(df_normalized, x='Intensidade_de_cor', y='Alcool', color='labels', width=800, height=800)
fig.show()
# + [markdown] id="c_MpCTIgel8_"
# Como nós podemos perceber ficou melhor organizado, os grupos ficaram mais bem definidos.
#
# Vamos agora passar para o gráfico 3D.
# + colab={"base_uri": "https://localhost:8080/"} id="IWnJhrmLeH6F" outputId="23296105-372f-4fe8-ffaf-504240acb4e2"
fig = px.scatter_3d(df_normalized, x='Intensidade_de_cor', y='Alcool', z='Proline', color='labels', width=800, height=800)
fig.show()
# + [markdown] id="LOV7iRyjtc6e"
# Já temos o gráfico 3D, mas não sabemos onde estão os centróides. Vamos adicioná-los.
# + colab={"base_uri": "https://localhost:8080/"} id="k2Wp34cxfQah" outputId="e584e0cb-a570-47b7-a51f-e7c13b25a932"
df_normalized.drop('labels', axis=1, inplace=True)
centros = pd.DataFrame(agrupador.cluster_centers_)
centros.columns = df_normalized.columns
centros.head()
# + colab={"base_uri": "https://localhost:8080/"} id="QgSRI9eJhp_D" outputId="ace3a091-346b-49cc-f5a4-d080e0a2b817"
df_normalized['labels'] = labels
fig.add_trace(go.Scatter3d(x=centros.Intensidade_de_cor,
y=centros.Alcool,
z=centros.Proline,
mode='markers',
marker=dict(color='cyan'),
text=[0,1,2]))
df_normalized.drop('labels', axis=1, inplace=True)
fig.show()
# + [markdown] id="TL2hfgtxuc_o"
# # Aula 3 - DBSCAN
#
# DBSCAN significa “Density-Based Spatial Clustering of Applications with Noise”, ou seja, “Agrupamento Espacial Baseado em Densidade para Aplicações com Ruído”.
#
# Por que utilizar? O K-means não funciona bem quando os nossos clusters não têm simetria radial.
#
# O DBSCAN define uma distância mínima para que os pontos sejam considerados vizinhos, essa distância é chamada de “eps”. Ele inicializa em uma amostra aleatória, verifica todos os vizinhos dessa amostra e vai fazendo isso até que ele encontre uma amostra que não tenha mais vizinhos, ou seja, até ele não possa mais expandir.
#
# O objetivo do algoritmo é fazer com que cada grupo tenha pelo menos um número mínimo ou a densidade mínima (min_samples) de pontos vizinhos, considerando como condição de vizinhança uma distancia menor ou igual a “eps”, que é o “epsilon”.
#
# Vantagens do DBSCAN:
# * Ele é robusto a “outliers”, ou seja, ele é robusto a ruído.
# * Ele pode detectar qualquer tipo de forma de cluster.
# * O custo computacional dele não é tão alto.
# * Ele é o segundo algoritmo de clusterização mais utilizado, ou seja, ele é muito popular.
#
# Desvantagens:
# * Ele não funciona bem com dados de alta dimensionalidade, assim como no KMeans, porque as métricas de distância acabam perdendo um pouco de sentido.
# * Ele não consegue identificar clusters de várias densidades porque o “min_samples” é fixo e o “eps” também. Se eles fossem variáveis, de alguma maneira talvez ele conseguisse fazer isso.
# * A parametrização de “eps” e “min_samples” nem sempre são fáceis.
#
# + colab={"base_uri": "https://localhost:8080/"} id="S0Un7lkkqYTD" outputId="c181a83f-6a7e-4a43-889e-3150cc5dc337"
from sklearn.cluster import DBSCAN
agrupador = DBSCAN(eps=1.31, min_samples=15, metric='manhattan')
agrupador.fit(df_normalized)
# + colab={"base_uri": "https://localhost:8080/"} id="JK1DV4p-wSIr" outputId="6531f138-af1e-4a51-d0c0-f3cb49aa99f7"
agrupador.labels_
# + colab={"base_uri": "https://localhost:8080/"} id="qAH_iybjwjcM" outputId="017f4e5f-6786-43d4-d909-30f0a32ae563"
np.unique(agrupador.labels_)
# + [markdown] id="nU1CpbP16eZ6"
# Vamos diminuir o eps e ver como o DBSCAN responde.
# + colab={"base_uri": "https://localhost:8080/"} id="ZNDyIFvzprzL" outputId="71e872d9-65f9-46ee-d04d-7fc8fa3b2cd0"
labels = agrupador.labels_
df_normalized['labels'] = labels
fig = px.scatter_3d(df_normalized, x='Intensidade_de_cor', y='Alcool', z='Proline', color='labels', width=800, height=800)
df_normalized.drop('labels', axis=1, inplace=True)
fig.show()
# + colab={"base_uri": "https://localhost:8080/"} id="Fl_r00ZRySk7" outputId="fe4d46b9-feca-4989-a378-b8baf8a7a738"
agrupador = DBSCAN(eps=0.01, min_samples=15, metric='manhattan')
agrupador.fit(df_normalized)
agrupador.labels_
# + [markdown] id="eSe_hsN16xLB"
# Todos os nossos dados ou todas as nossas amostras foram consideradas como ruído.
#
# Quando nós utilizamos um valor muito pequeno de “eps” nós falamos que os vizinhos eram só as amostras que estivessem muito pertos e provavelmente nós tínhamos poucas amostras desse jeito.
#
# Agora vamos verificar eps=10
# + colab={"base_uri": "https://localhost:8080/"} id="ewNgTanz6qk8" outputId="591cd89a-7f69-449d-f117-4fa77125c8ac"
agrupador = DBSCAN(eps=10, min_samples=15, metric='manhattan')
agrupador.fit(df_normalized)
agrupador.labels_
# + [markdown] id="VjWYDj3k7VGc"
# Todas as nossas amostras foram consideradas no mesmo cluster.
#
# Vamos experimentar eps=1.5 e min_samples=30
# + colab={"base_uri": "https://localhost:8080/"} id="pFb8nb-B7T8T" outputId="0566794b-25c9-40cc-8eb5-b9f16ed89c39"
agrupador = DBSCAN(eps=1.5, min_samples=30, metric='manhattan')
agrupador.fit(df_normalized)
agrupador.labels_
# + colab={"base_uri": "https://localhost:8080/"} id="tKnwc4OFrPed" outputId="7468f47c-2efb-4b73-b3ca-0d8bc97298b5"
labels = agrupador.labels_
df_normalized['labels'] = labels
fig = px.scatter_3d(df_normalized, x='Intensidade_de_cor', y='Alcool', z='Proline', color='labels', width=800, height=800)
df_normalized.drop('labels', axis=1, inplace=True)
fig.show()
# + [markdown] id="7z1mUsU57xgr"
# Ficamos com um único cluster e o resto das amostras como ruído.
#
# Se aumentarmos o min_samples para 80, precisaremos de 80 amostras para formar um cluster. Assim, todas amostram serão ruído.
# + colab={"base_uri": "https://localhost:8080/"} id="SItf1O4_8Jjg" outputId="41735210-eedd-43b8-9cc5-094564dca1d7"
agrupador = DBSCAN(eps=1.5, min_samples=80, metric='manhattan')
agrupador.fit(df_normalized)
agrupador.labels_
# + [markdown] id="tag4XSbh8b12"
# Agora, vamos alterar o parâmetro metric para 'euclidian'.
# + colab={"base_uri": "https://localhost:8080/"} id="VfPIr2YS8LA3" outputId="818b0810-3e04-4af8-d5cb-05590552827b"
agrupador = DBSCAN(eps=1.5, min_samples=15, metric='euclidean')
agrupador.fit(df_normalized)
agrupador.labels_
# + [markdown] id="212L7rWbjVVp"
# Vamos seguir com uma sequência de testes alterando os parâmetros e observando os resultados:
# + colab={"base_uri": "https://localhost:8080/"} id="DAfbkRNM8or4" outputId="6661cab2-aade-4be0-e263-f0f773f694dc"
agrupador = DBSCAN(eps=0.7, min_samples=15, metric='euclidean')
agrupador.fit(df_normalized)
agrupador.labels_
# + colab={"base_uri": "https://localhost:8080/"} id="ITQIQc8xrTKm" outputId="f9884df2-bb76-4cf2-bc38-b87c04fc49a2"
labels = agrupador.labels_
df_normalized['labels'] = labels
fig = px.scatter_3d(df_normalized, x='Intensidade_de_cor', y='Alcool', z='Proline', color='labels', width=800, height=800)
df_normalized.drop('labels', axis=1, inplace=True)
fig.show()
# + colab={"base_uri": "https://localhost:8080/"} id="qTOOOuA-j3EO" outputId="f5ca2a94-bc95-4035-aba9-af686465ed2c"
agrupador = DBSCAN(eps=0.3, min_samples=15, metric='euclidean')
agrupador.fit(df_normalized)
agrupador.labels_
# + colab={"base_uri": "https://localhost:8080/"} id="g1QUtrblkC3e" outputId="7329135d-81c0-4c2c-c7b7-51ccb9ee83f5"
agrupador = DBSCAN(eps=0.5, min_samples=15, metric='euclidean')
agrupador.fit(df_normalized)
agrupador.labels_
# + colab={"base_uri": "https://localhost:8080/"} id="C9LUDUdsrm4F" outputId="e1da9cae-bcac-4646-9af4-9a96d1a2a988"
labels = agrupador.labels_
df_normalized['labels'] = labels
fig = px.scatter_3d(df_normalized, x='Intensidade_de_cor', y='Alcool', z='Proline', color='labels', width=800, height=800)
df_normalized.drop('labels', axis=1, inplace=True)
fig.show()
# + [markdown] id="FxNhwex8kSSr"
# O “eps” e o “min_samples” variam de maneira inversamente proporcional, ou seja, o efeito causado pelo aumento do “eps” é o mesmo efeito causado pela diminuição do “min_samples”. O efeito causado pelo aumento do “min_samples” é o mesmo efeito causado pela diminuição do “eps”.
#
# A métrica de distância influencia bastante, porque ela influencia na forma como eu calculo o “eps”.
#
# Existem outras métricas de distância, além das apresentadas aqui. Para alterar no algoritmo, basta modificar o parâmetro metric. As métricas de distância permitidas são:
# * Do scikit-learn:
# * ‘cityblock’
# * ‘cosine’
# * ‘euclidean’
# * ‘l1’
# * ‘l2’
# * ‘manhattan’]
# * Do scipy.spatial.distance:
# * ‘braycurtis’
# * ‘canberra’
# * ‘chebyshev’
# * ‘correlation’
# * ‘dice’
# * ‘hamming’
# * ‘jaccard’
# * ‘kulsinski’
# * ‘mahalanobis’
# * ‘minkowski’
# * ‘rogerstanimoto’
# * ‘russellrao’
# * ‘seuclidean’
# * ‘sokalmichener’
# * ‘sokalsneath’
# * ‘sqeuclidean’
# * ‘yule’
# + [markdown] id="wiP_3Ocmm-yE"
# # Aula 4 - Mean shift
#
# Vamos agora utilizar o método de clusterizaçaõ cahamdo Mean Shift, o qual não tem a necessidade de definirmos parâmetros.
# + id="TeY71FzukG0d"
from sklearn.cluster import MeanShift
# + colab={"base_uri": "https://localhost:8080/"} id="tXFLGzAEr-9-" outputId="983f7c8b-c282-422d-a185-f6f4e70f5cc6"
agrupador = MeanShift()
agrupador.fit(df)
agrupador.labels_
# + colab={"base_uri": "https://localhost:8080/"} id="SNvUiLwfun_m" outputId="0b870c2f-ac2d-484c-cd44-9b50005ea5e8"
labels = agrupador.labels_
df['labels'] = labels
fig = px.scatter_3d(df, x='Intensidade_de_cor', y='Alcool', z='Proline', color='labels', width=800, height=800)
df.drop('labels', axis=1, inplace=True)
fig.show()
# + [markdown] id="pn4ZkmaJxY2l"
# O objetivo do algoritmo é agrupar os pontos de acordo com as regiões de alta densidade.
#
# * Vantagens:
# * K-means não costuma apresentar um bom resultado quando há ruído e grupos de simetria não radial
# * Ele tem menos parâmetros para ser ajustado que o DBSCAN. O único parâmetro que nós mexemos é a largura de banda
# * Ele é robusto aos “outliers”
# * Ele pode detectar qualquer tipo de forma:
# * se você colocar uma largura de banda menor, você acaba detectando formas mais estranhas
# * larguras de banda maiores podemos detectar formas com simetria mais radial
# * Desvantagens:
# * Não funciona bem com dados de alta dimensionalidade
# * A parametrização da largura de banda não é muito simples
# * É computacionalmente caro, porque todo ponto no início é tratado como um cluster
# + id="dKdagEjPupD2"
from sklearn.cluster import estimate_bandwidth
# + colab={"base_uri": "https://localhost:8080/"} id="MCx87FhOzWAu" outputId="d2c667fa-40ae-4863-e6a4-1adda2751acd"
BW = estimate_bandwidth(df, quantile=0.1)
agrupador = MeanShift(BW)
agrupador.fit(df)
agrupador.labels_
# + colab={"base_uri": "https://localhost:8080/"} id="2rWuS1L_z5Ze" outputId="e24f02b9-9787-402a-d45d-f6bddb954f26"
labels = agrupador.labels_
df['labels'] = labels
fig = px.scatter_3d(df, x='Intensidade_de_cor', y='Alcool', z='Proline', color='labels', width=800, height=800)
df.drop('labels', axis=1, inplace=True)
fig.show()
# + [markdown] id="JazM-tsF1CA1"
# Com o valor do quantil pequeno (quantile=0.1) nossa região de busca dentro do MeanShift, ou seja, nossa região de cálculo da média e de deslocamento da média ficou muito pequena, acabamos gerando vários clusters, porque o centro dos nossos clusters quase não caminharam então eles, não se agruparam muito.
#
# Se colocarmos um valor grande no quantil, esperamos que aconteça o contrário, que tenhamos dois clusters ou um.
# + colab={"base_uri": "https://localhost:8080/"} id="WoY_erRK0Z7g" outputId="c4186327-739f-4661-dd0f-a305a5546444"
BW = estimate_bandwidth(df, quantile=0.8)
agrupador = MeanShift(BW)
agrupador.fit(df)
agrupador.labels_
# + colab={"base_uri": "https://localhost:8080/"} id="clWwY6o20pYw" outputId="916ce1f0-7374-49cc-fbfa-517aa791aec7"
labels = agrupador.labels_
df['labels'] = labels
fig = px.scatter_3d(df, x='Intensidade_de_cor', y='Alcool', z='Proline', color='labels', width=800, height=800)
df.drop('labels', axis=1, inplace=True)
fig.show()
# + [markdown] id="Ji0PKzqJ4J4Q"
# Vamos buscar um valor de BW que produza 3 clusters.
# + colab={"base_uri": "https://localhost:8080/"} id="ZBznvCRC0tDa" outputId="1830f475-0779-4bfa-d56d-4346ff723151"
BW = estimate_bandwidth(df, quantile=0.3)
agrupador = MeanShift(BW)
agrupador.fit(df)
agrupador.labels_
# + colab={"base_uri": "https://localhost:8080/"} id="evRFijXi1xbl" outputId="7414a3c1-bbc5-4892-f4d7-0a5941077ebe"
labels = agrupador.labels_
df['labels'] = labels
fig = px.scatter_3d(df, x='Intensidade_de_cor', y='Alcool', z='Proline', color='labels', width=800, height=800)
df.drop('labels', axis=1, inplace=True)
fig.show()
# + [markdown] id="8nrHy40N4Yxw"
# # Aula 5 - Coeficiente de silhueta para avaliação de clusterizações
#
# + colab={"base_uri": "https://localhost:8080/"} id="4ThlOeHR2nmQ" outputId="878c0cf1-c3ab-4a4c-cfc9-cb9f4d74834c"
agrupador = KMeans(n_clusters=5)
agrupador.fit(df)
agrupador.labels_
# + colab={"base_uri": "https://localhost:8080/", "height": 817} id="uooEaYf747Go" outputId="1540c8c4-bd34-438f-bd7b-ba2211dc3e5a"
labels = agrupador.labels_
df['labels'] = labels
fig = px.scatter_3d(df, x='Intensidade_de_cor', y='Alcool', z='Proline', color='labels', width=800, height=800)
df.drop('labels', axis=1, inplace=True)
fig.show()
# + [markdown] id="AB05NtSH5b-D"
# Depois de realizar todos esses tipos de clusterização, K-means, DBSCAN e MeanShift, gostaríamos de saber se existe alguma métrica que indicasse que a clusterização está boa.
#
# Qual é a maneira de nós avaliarmos? Para isso, podemos utilizar uma métrica de coeficiente de silhueta. Ela basicamente analisa o quanto os pontos do cluster estão próximos dos demais pontos do mesmo cluster e afastado dos pontos do cluster vizinho.
#
# Ele vai avaliar uma coisa chamada **coesão**. O quanto eu estou próximo aos elementos do meu grupo. Quanto maior a coesão, melhor o coeficiente de silhueta e de **similaridade**, o quanto eu estou afastado dos elementos do outro bairro. Quanto mais afastado eu estiver deles, melhor.
#
# Quanto maior o valor do coeficiente de silhueta, ou seja, quanto **maior a coesão** e **maior a similaridade**, melhor e mais bem organizado, mais bem agrupado - segundo o critério de distância - estão os meus pontos.
# + id="867QWG_84_4N"
from sklearn.metrics import silhouette_score
# + colab={"base_uri": "https://localhost:8080/"} id="p5YMmp_J7GHc" outputId="ef333b51-3b03-4636-da40-133c270e128d"
faixa_n_clusters = [i for i in range(2,10)]
print(faixa_n_clusters)
# + id="LnVA_Tgj7Q3b"
valores_silhueta = []
for k in faixa_n_clusters:
agrupador = KMeans(n_clusters=k)
labels = agrupador.fit_predict(df_normalized)
media_silhueta = silhouette_score(df_normalized, labels)
valores_silhueta.append(media_silhueta)
# + colab={"base_uri": "https://localhost:8080/", "height": 418} id="LOdhaS7d75Ey" outputId="e0f6b9d2-1700-484c-9b6c-61ea1636f32b"
fig = px.scatter(x=faixa_n_clusters, y=valores_silhueta,width=600, height=400)
fig.update_layout(title = 'Valores de silhueta médios',
xaxis_title = 'Número de Clusters',
yaxis_title = 'Valor médio de silhueta')
fig.show()
# + [markdown] id="e06wsfTy-_Ua"
# Como nós podemos perceber, o maior valor de coeficiente de silhueta que nós temos é quando o nosso valor de “K”, que é o número de clusters, é igual a “3”.
#
# Vamos testar agora para o MeanShift.
# + id="wZ07Q1Xt81hG" colab={"base_uri": "https://localhost:8080/"} outputId="1d77d1e1-4fac-4a24-f090-7c6704c9b9f4"
faixa_quantil = np.linspace(0.02, 0.5, 15)
faixa_quantil
# + id="Gq6e3LEBKsRV"
valores_silhueta = []
quantil_plot = []
for quantil in faixa_quantil:
bandwidth = estimate_bandwidth(df_normalized, quantile=quantil)
agrupador = MeanShift(bandwidth)
labels = agrupador.fit_predict(df_normalized)
if(len(np.unique(labels)) < len(df_normalized) and len(np.unique(labels)) > 1):
media_silhueta = silhouette_score(df_normalized, labels)
valores_silhueta.append(media_silhueta)
quantil_plot.append(quantil)
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="nYUq1OcjLTPl" outputId="4dd07a87-45db-4b37-dfd1-1fa7dc400ea3"
fig = go.Figure()
fig.add_trace(go.Scatter(x=quantil_plot,
y=valores_silhueta))
fig.update_layout(dict(xaxis_title='Quantil',
yaxis_title='Média Silhueta'))
fig.show()
# + colab={"base_uri": "https://localhost:8080/"} id="8jucjIdayGfe" outputId="2e8a767b-71ca-47ea-db9e-9a50268141e8"
faixa_quantil = np.linspace(0.02, 0.5, 30)
faixa_quantil
# + id="BJt2YWiUOBs7"
valores_silhueta = []
quantil_plot = []
for quantil in faixa_quantil:
bandwidth = estimate_bandwidth(df_normalized, quantile=quantil)
agrupador = MeanShift(bandwidth)
labels = agrupador.fit_predict(df_normalized)
if(len(np.unique(labels)) < len(df_normalized) and len(np.unique(labels)) > 1):
media_silhueta = silhouette_score(df_normalized, labels)
valores_silhueta.append(media_silhueta)
quantil_plot.append(quantil)
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="5uZr2bQAxDJr" outputId="bacfa0e7-1a9f-4cab-e3c3-e7df3ea25ad7"
fig = go.Figure()
fig.add_trace(go.Scatter(x=quantil_plot,
y=valores_silhueta))
fig.update_layout(dict(xaxis_title='Quantil',
yaxis_title='Média Silhueta'))
fig.show()
# + [markdown] id="HkNmtlbTysdH"
# Comparando Mean shift, DBSCAN e K-means
# + colab={"base_uri": "https://localhost:8080/"} id="-OA-WWH5xExr" outputId="cf41ad99-f79b-4fee-81d6-54a73a8d5503"
agrupador_kmeans = KMeans(n_clusters=3)
agrupador_DBSCAN = DBSCAN(eps=2.1, min_samples=56, metric='manhattan')
BW = estimate_bandwidth(df_normalized, quantile=0.28)
agrupador_meanshift = MeanShift(bandwidth=BW)
labels_kmeans = agrupador_kmeans.fit_predict(df_normalized)
labels_dbscan = agrupador_DBSCAN.fit_predict(df_normalized)
labels_meanshift = agrupador_meanshift.fit_predict(df_normalized)
print(f'Labels K-means: {labels_kmeans}\n')
print(f'Labels DBSCAN: {labels_dbscan}\n')
print(f'Labels MeanShift: {labels_meanshift}\n')
# + colab={"base_uri": "https://localhost:8080/"} id="HvsK52iT0b4w" outputId="33cc3cbf-74b3-41ba-a3cd-e08165e01983"
print(f'O coeficiente de silhueta do K-means é: {silhouette_score(df_normalized, labels_kmeans)}\n')
print(f'O coeficiente de silhueta do DBSCAN é: {silhouette_score(df_normalized, labels_dbscan)}\n')
print(f'O coeficiente de silhueta do MeanShift é: {silhouette_score(df_normalized, labels_meanshift)}\n')
# + [markdown] id="UL1KfxX31SXp"
# Com o resultado acima percebemos que o melhor método para este dataset é o K-means com 3 Clusters.
#
# Isso se deve provavelmente ao fato de que a dimensionalidade do dataset ser grande e os métodos DBSCAN e Mean Shift não performam bem nesta situação. Esses dois métodos não conseguiram dividir os dados em clusters muito bem definidos, por isso que eles deram um coeficiente de silhueta não tão bom.
#
# Sempre que nós formos aplicar alguma coisa relacionada clusterização, nós não devemos utilizar apenas uma técnica, mas várias e depois avaliarmos qual delas produziu o melhor resultado segundo algum tipo de análise, que no nosso caso foi o coeficiente de silhueta. Uma vez analisado isso, você define a melhor técnica e utiliza essa para agrupar os seus dados.
# + id="1SldbzIH1HZM"
| Clustering_Básico_k_means,_DBSCAN_e_mean_shift.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1.前言
#
# 总是聊并发的话题,聊到大家都免疫了,所以这次串讲下个话题——数据库(欢迎纠正补充)
#
# 看完问自己一个问题来自我检测:**NoSQL我到底该怎么选?**
#
# 数据库排行:https://db-engines.com/en/ranking
#
# ## 1.1.分类
#
# 主要有这么三大类:(再老的数据库就不说了)
#
# ### 1.传统数据库(SQL):
#
# - 关系数据库:SQLite、MySQL、SQLServer、PostgreSQL、Oracle...
#
# ### 2.高并发产物(NoSQL):
#
# 1. **键值数据库**:**Redis**、MemCached...
# - PS:现在`LevelDB`和`RocksDB`也是比较高效的解决方案
# - PS:360开发的`pika`(redis的补充)可以解决Redis容量过大的问题
# 2. **文档数据库**:**MongoDB**、ArangoDB、CouchBase、CouchDB、RavenDB...
# - PS:Python有一款基于json存储的轻量级数据库:`tinydb`
# 3. 列式数据库:**Cassandra**、**HBase**、BigTable...
# - PS:小米的**Pegasus**计划取代HBase
# 4. **搜索引擎系**:**Elasticsearch**、Solr、Sphinx...
# - PS:这几年用Rust编写的轻量级搜索引擎`sonic`很火
# 5. 图形数据库:**Neo4J**、**Flockdb**、**ArangoDB**、OrientDB、Infinite Graph、InfoGrid...
# - PS:基于Redis有一款图形数据库用的也挺多:`RedisGraph`
# - PS:随着Go的兴起,这些图形数据库很火:**Cayley**、`Dgraph`、`Beam`
#
# PS:项目中最常用的其实就是`Redis`、`Elasticsearch`、`MongoDB`
# > PS:`ArangoDB`是一个原生的多模型数据库,具有文档,图形和键值的灵活数据库
#
# ### 3.新时代产物(TSDB):
#
# - 时序数据库:**InfluxDB**、**LogDevice**、Graphite、、OpenTSDB...
#
# 来看个权威的图:(红色的是推荐NoSQL,灰色是传统SQL)
# 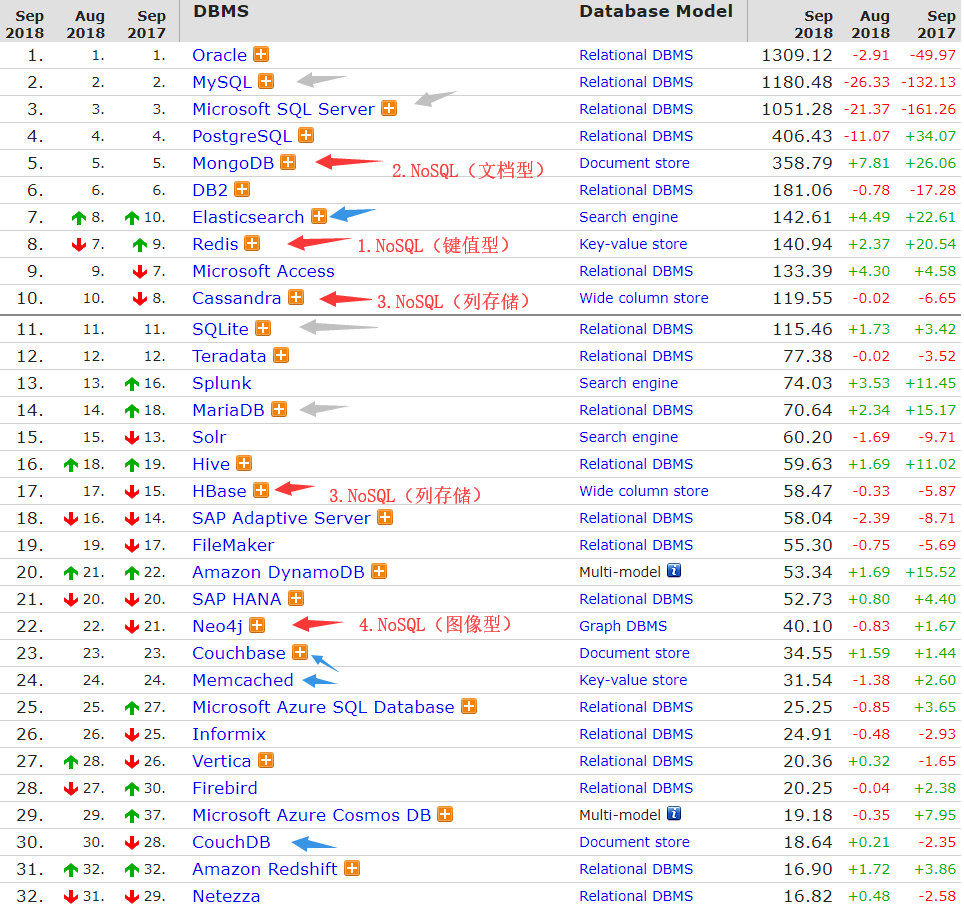
#
# ## 1.2.概念
#
# 先说下NoSQL不是不要使用传统SQL了,而是不仅仅是传统的SQL(not only sql)
#
# ### 1.关系型数据库优劣
#
# 先看看传统数据库的好处:
# 1. 通过事务保持数据一致
# 2. 可以Join等复杂查询
# 3. 社区完善(遇到问题简单搜下就ok了)
#
# 当然了也有不足的地方:
# 1. 数据量大了的时候修改表结构。eg:加个字段,如果再把这个字段设置成索引那是卡到爆,完全不敢在工作时间搞啊
# 2. 列不固定就更蛋疼了,一般设计数据库不可能那么完善,都是后期越来越完善,就算自己预留了`保留字段`也不人性化啊
# 3. 大数据写入处理比较麻烦,eg:
# 1. 数据量不大还好,批量写入即可。
# 2. 可是本身数据量就挺大的,进行了`主从复制`,读数据在`Salver`进行到没啥事,但是大量写数据库怼到`Master`上去就吃不消了,必须得加主数据库了。
# 3. 加完又出问题了:虽然把主数据库一分为二,但是容易发生`数据不一致`(同样数据在两个主数据库更新成不一样的值),这时候得结合分库分表,把表分散在不同的主数据库中。
# 4. 完了吗?NoNoNo,想一想表之间的Join咋办?岂不是要跨数据库和跨服务器join了?简直就是拆东墙补西墙的节奏啊,所以各种中间件就孕育而生了【SQLServer这方面扩展的挺不错的,列存储也自带,也跨平台了(建议在Docker中运行)(<a href="https://www.cnblogs.com/dunitian/p/6041323.html" target="_blank">点我查看几年前写的一篇文章</a>)】
# 5. 欢迎补充~(说句良心话,中小型公司`SQLServer`绝对是最佳选择,能省去很多时间)
#
# ---
#
# ### 2.NoSQL
#
# 现在说说NoSQL了:(其实你可以理解为:**NoSQL就是对原来SQL的扩展补充**)
# 1. 分表分库的时候一般把关联的表放在同一台服务器上,这样便于join操作。而NoSQL不支持join,反而不用这么局限了,数据更容易分散存储
# 2. 大量数据处理这块,读方面传统SQL并没有太多劣势,NoSQL主要是进行缓存处理,批量写数据方面测试往往远高于传统SQL,而且NoSQL在扩展方面方便太多了
# 3. 多场景类型的NoSQL(键值,文档、列、图形)
#
# ---
#
# 如果还是不清楚到底怎么选择NoSQL,那就再详细说说每个类型的特点:
#
# 1. 键值数据库:这个大家很熟悉,主要就是`键值存储`,代表=>**Redis**(支持持久化和数据恢复,后面我们会详谈)
# 2. 文档数据库:代表=>**MongoDB**(优酷的在线评论就是用基于MongoDB的)
# 1. 一般都不具备事务(`MongoDB 4.0`开始支持`ACID`事务了)
# 2. 不支持Join(Value是一个可变的`类JSON格式`,表结构修改比较方便)
# 3. 列式数据库:代表:**Cassandra**、`HBase`
# 1. 对大量行少量列进行修改更新(新增一字段,批量做个啥操作的不要太方便啊~针对列为单位读写)
# 2. 扩展性高,数据增加也不会降低对应的处理速度(尤其是写)
# 4. 搜索引擎系:代表:**Elasticsearch**,太经典了不用说了(传统模糊搜索只能like太low,所以有了这个)
# 5. 图形数据库:代表:**Neo4J**、**Flockdb**、**ArangoDB**(数据模型是图结构的,主要用于 **关系比较复杂** 的设计,比如绘制一个QQ群关系的可视化图、或者绘制一个微博粉丝关系图等)
#
# 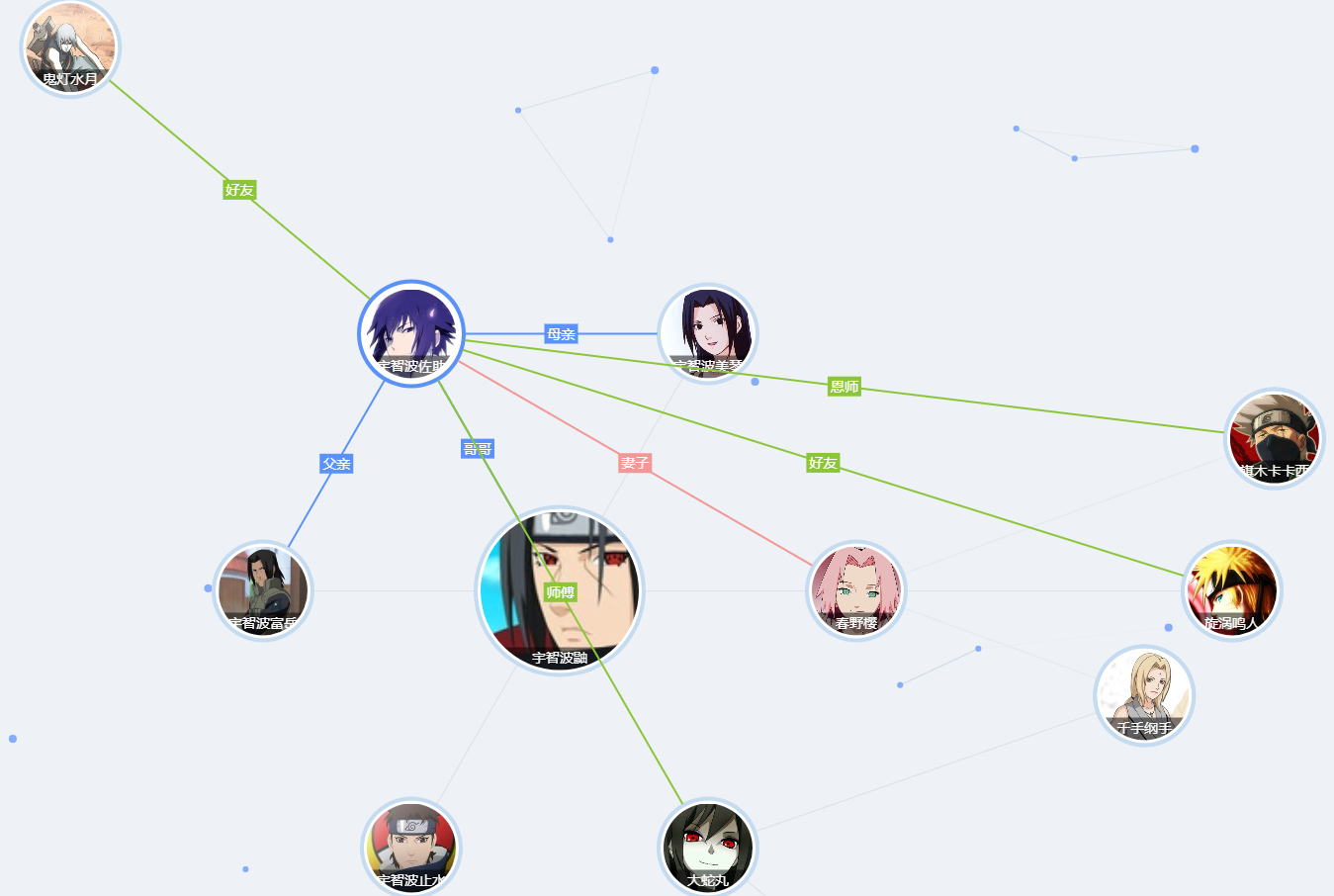
#
# ---
#
# 回头还是要把并发剩余几个专题深入的,认真看的同志会发现不管什么语言底层实现都是差不多的。
#
# 比如说进程,其底层就是用到了我们第一讲说的`OS.fork`。再说进(线)程通信,讲的`PIPE、FIFO、Lock、Semaphore`等很少用吧?但是`Queue`底层就是这些实现的,不清楚的话怎么读源码?
#
# 还记得当时`引入Queue篇`提到Java里的`CountDownLatch`吗?要是不了解`Condition`怎么自己快速模拟一个Python里面没有的功能呢?
#
# 知其然不知其所以然是万万不可取的。等后面讲MQ的时候又得用到`Queue`的知识了,可谓一环套一环~
#
# 既然不是公司的萌妹子,所以呢~技术的提升还是得靠自己了^_^,先到这吧,最后贴个常用解决方案:
#
# Python、NetCore常用解决方案(持续更新)<https://github.com/LessChina>
#
# ---
#
# # 2.概念
#
# 上篇提到了`ACID`这次准备说说,然后再说说`CAP`和数据一致性
#
# ## 2.1.ACID事务
#
# 以小明和小张转账的例子继续说说:
# 1. A:原子性(Atomic)
# - 小明转账1000给小张:小明-=1000 => 小张+=1000,这个 **(事务)是一个不可分割的整体**,如果小明-1000后出现问题,那么1000得退给小明
# 2. C:一致性(Consistent)
# - 小明转账1000给小张,必须保证小明+小张的总额不变(假设不受其他转账(事务)影响)
# 3. I:隔离性(Isolated)
# - 小明转账给小张的同时,小潘也转钱给了小张,需要保证他们相互不影响(主要是并发情况下的隔离)
# 4. D:持久性(Durable)
# - 小明转账给小张银行要有记录,即使以后扯皮也可以拉流水账【事务执行成功后进行的持久化(就算数据库之后挂了也能通过Log恢复)】
#
# ## 2.2.CAP概念
#
# <a href="https://baike.baidu.com/item/CAP原则" target="_blank">CAP</a>是分布式系统需要考虑的三个指标,数据共享只能满足两个而不可兼得:
# 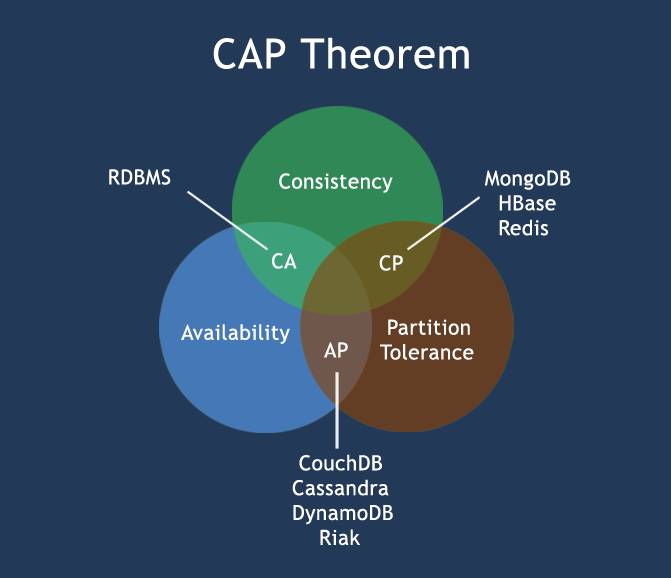
#
# 1. C:一致性(**C**onsistency)
# - 所有节点访问同一份最新的数据副本(在分布式系统中的所有数据备份,在同一时刻是否同样的值)
# - eg:分布式系统里更新后,某个用户都应该读取最新值
# 2. A:可用性(**A**vailability)
# - 在集群中一部分节点故障后,集群整体是否还能响应客户端的读写请求。(对数据更新具备高可用性)
# - eg:分布式系统里每个操作总能在一定时间内返回结果(超时不算【网购下单后一直等算啥?机房挂几个服务器也不影响】)
# 3. P:分区容错性(**P**artition Toleranc)
# - 以实际效果而言,分区相当于对通信的时限要求。系统如果不能在时限内达成数据一致性,就意味着发生了分区的情况,必须就当前操作在C和A之间做出选择。
# - eg:分布式系统里,存在网络延迟(分区)的情况下依旧可以接受满足一致性和可用性的请求
#
# ### CA
#
# 代表:**`传统关系型数据库`**
#
# 如果想避免分区容错性问题的发生,一种做法是将所有的数据(与事务相关的)都放在一台机器上。虽然无法100%保证系统不会出错,但不会碰到由分区带来的负面效果(会严重的影响系统的扩展性)
#
# 作为一个分布式系统,放弃P,即相当于放弃了分布式,一旦并发性很高,单机服务根本不能承受压力。像很多银行服务,确确实实就是舍弃了P,只用高性能的单台小型机保证服务可用性。(**所有NoSQL数据库都是假设P是存在的**)
#
# ### CP
#
# 代表:**`Zookeeper`、`Redis`**(分布式数据库、分布式锁)
#
# 相对于放弃“分区容错性“来说,其反面就是放弃可用性。`一旦遇到分区容错故障,那么受到影响的服务需要等待数据一致`(**等待数据一致性期间系统无法对外提供服务**)
#
# ### AP
#
# 代表:**`DNS数据库`**(`IP和域名相互映射的分布式数据库`,联想修改IP后为什么`TTL`需要10分钟左右保证所有解析生效)
#
# 
#
# 反DNS查询:<a href="https://www.cnblogs.com/dunitian/p/5074773.html" target="_blank">https://www.cnblogs.com/dunitian/p/5074773.html</a>
#
# 放弃强一致,保证最终一致性。所有的`NoSQL`数据库都是介于`CP`和`AP`之间,尽量往`AP`靠,(**传统关系型数据库注重数据一致性,而对海量数据的分布式处理来说可用性和分区容错性优先级高于数据一致性**)eg:
#
# **不同数据对一致性要求是不一样的**,eg:
# 1. 用户评论、弹幕这些对一致性是不敏感的,很长时间不一致性都不影响用户体验
# 2. 像商品价格等等你敢来个看看?对一致性是很高要求的,容忍度铁定低于10s,就算使用了缓存在订单里面价格也是最新的(平时注意下JD商品下面的缓存说明,JD尚且如此,其他的就不用说了)
#
# 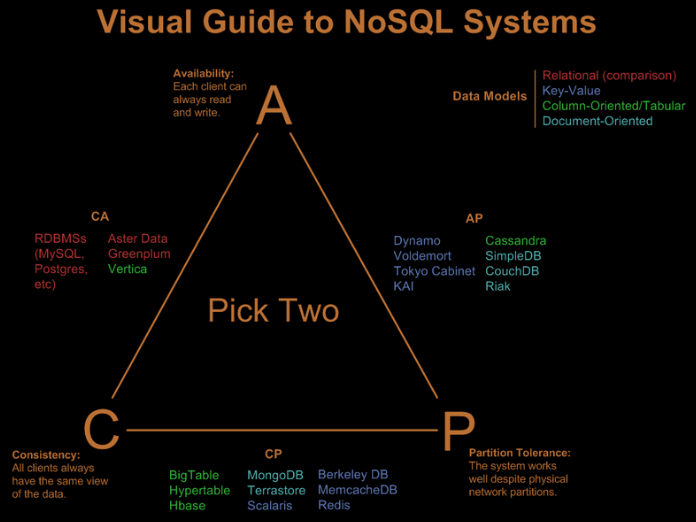
#
# ## 2.3.数据一致性
#
# 传统关系型数据库一般都是使用悲观锁的方式,但是例如秒杀这类的场景是hou不动,这时候往往就使用乐观锁了(CAS机制,之前讲并发和锁的时候提过),上面也稍微提到了不同业务需求对一致性有不同要求而CAP不能同时满足,这边说说主要就两种:
# 1. **强一致性**:无论更新在哪个副本上,之后对操作都要能够获取最新数据。多副本数据就需要**`分布式事物来保证数据一致性`**了(这就是问什么项目里面经常提到的原因)
# 2. **最终一致性**:在这种约束下保证用户最终能读取到最新数据。举几个例子:
# 1. 因果一致性:A、B、C三个独立进程,A修改了数据并通知了B,这时候B得到的是最新数据。因为A没通知C,所以C不是最新数据
# 2. **会话一致性**:用户自己提交更新,他可以在会话结束前获取更新数据,会话结束后(其他用户)可能不是最新的数据(提交后JQ修改下本地值,不能保证数据最新)
# 3. **读自写一致性**:和上面差不多,只是不局限在会话中了。用户更新数据后他自己获取最新数据,其他用户可能不是最新数据(一定延迟)
# 4. 单调读一致性:用户读取某个数值,后续操作不会读取到比这个数据还早的版本(新的程度>=读取的值)
# 5. **单调写一致性**(时间轴一致性):所有数据库的所有副本按照相同顺序执行所有更新操作(有点像`Redis`的`AOF`)
#
# ## 2.4.一致性实现方法
#
# ### `Quorum`系统`NRW`策略(常用)
#
# Quorum是集合A,A是全集U的子集,A中任意取集合B、C,他们两者都存在交集。
#
# **NRW算法**:
# 1. N:表示数据所具有的副本数。
# 2. R:表示完成读操作所需要读取的最小副本数(一次读操作所需参与的最小节点数)
# 3. W:表示完成写操作所需要写入的最小副本数(一次写操作所需要参与的最小节点数)
# 4. **只需要保证`R + W > N`就可以保证强一致性**(读取数据的节点和被同步写入的节点是有重叠的)比如:N=3,W=2,R=2(有一个节点是读+写)
#
# **扩展**:
# 1. 关系型数据库中,如果N=2,可以设置W=2,R=1(写耗性能一点),这时候系统需要两个节点上数据都完成更新才能确认结果并返回给用户
# 2. 如果`R + W <= N`,这时候读写不会在一个节点上同时出现,系统只能保证最终一致性。副本达到一致性的时间依赖于系统**异步更新的方式**,不一致性时间=从更新节点~所有节点都异步更新完毕的耗时
# 3. R和W设置直接影响系统的性能、扩展和一致性:
# 1. 如果W设置为1,那么一个副本更新完就返回给用户,然后通过异步机制更新剩余的N-W个节点
# 2. 如果R设置为1,只要有一个副本被读取就可以完成读操作,R和W的值如果较小会影响一致性,较大会影响性能
# - 当W=1,R=N==>系统对写要求高,但读操作会比较慢(N个节点里面有1个挂了,读就完成不了了)
# - 当R=1,W=N==>系统对读操作有高要求,但写性能就低了(N个节点里面有1个挂了,写就完成不了了)
# 3. **常用方法**:一般设置**`R = W = N/2 + 1`**,这样性价比高,eg:N=3,W=2,R=2(`3个节点==>1写,1读,1读写`)
#
# 参考文章:
# ```
# http://book.51cto.com/art/201303/386868.htm
# https://blog.csdn.net/jeffsmish/article/details/54171812
# ```
#
# ### 时间轴策略(常用)
#
# 1. 主要是关系型数据库的日记==>记录事物操作,方便数据恢复
# 2. 还有就是并行数据存储的时候,由于数据是分散存储在不同节点的,对于同一节点来说只要关心`数据更新+消息通信`(数据同步):
# - **保证较晚发生的更新时间>较早发生的更新时间**
# - **消息接收时间 > 消息发送时刻的时间**(要考虑服务器时间差的问题~时间同步服务器)
#
# ### 其他策略
#
# 其实还有很多策略来保证,这些概念的对象逆天不是很熟~比如:**向量时钟策略**
#
# `推荐几篇文章`:
#
# ```
# https://www.cnblogs.com/yanghuahui/p/3767365.html
# http://blog.chinaunix.net/uid-27105712-id-5612512.html
# https://blog.csdn.net/dellme99/article/details/16845991
# https://blog.csdn.net/blakeFez/article/details/48321323
# ```
#
| database/notebook/2.NoSQL.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="I08sFJYCxR0Z"
# 
# + [markdown] id="FwJ-P56kq6FU"
# [](https://colab.research.google.com/github/JohnSnowLabs/spark-nlp-workshop/blob/master/tutorials/Certification_Trainings/Healthcare/1.5.Resume_MedicalNer_Model_Training.ipynb)
# + [markdown] id="7ztGkR5mb6zN"
# # 1.5 Resume MedicalNer Model Training
#
# Steps:
# - Train a new model for a few epochs.
# - Load the same model and train for more epochs on the same taxnonomy, and check stats.
# - Train a model already trained on a different data
# + [markdown] id="-68NMdHxJIco"
# ## Colab Setup
# + id="h9Mn1PNTNJTq"
import json
from google.colab import files
license_keys = files.upload()
with open(list(license_keys.keys())[0]) as f:
license_keys = json.load(f)
# Defining license key-value pairs as local variables
locals().update(license_keys)
# Adding license key-value pairs to environment variables
import os
os.environ.update(license_keys)
# + id="yvHraBK7b-LU"
# Installing pyspark and spark-nlp
# ! pip install --upgrade -q pyspark==3.1.2 spark-nlp==$PUBLIC_VERSION
# Installing Spark NLP Healthcare
# ! pip install --upgrade -q spark-nlp-jsl==$JSL_VERSION --extra-index-url https://pypi.johnsnowlabs.com/$SECRET
# Installing Spark NLP Display Library for visualization
# ! pip install -q spark-nlp-display
# + id="wIp1yLCWNWsn"
# if you want to start the session with custom params as in start function above
from pyspark.sql import SparkSession
def start(SECRET):
builder = SparkSession.builder \
.appName("Spark NLP Licensed") \
.master("local[*]") \
.config("spark.driver.memory", "16G") \
.config("spark.serializer", "org.apache.spark.serializer.KryoSerializer") \
.config("spark.kryoserializer.buffer.max", "2000M") \
.config("spark.jars.packages", "com.johnsnowlabs.nlp:spark-nlp_2.12:"+PUBLIC_VERSION) \
.config("spark.jars", "https://pypi.johnsnowlabs.com/"+SECRET+"/spark-nlp-jsl-"+JSL_VERSION+".jar")
return builder.getOrCreate()
#spark = start(SECRET)
# + colab={"base_uri": "https://localhost:8080/"} id="1t5Kp93GcH7z" outputId="063ee555-6026-464f-8912-beeba71e335e"
import json
import os
from pyspark.ml import Pipeline,PipelineModel
from pyspark.sql import SparkSession
from sparknlp.annotator import *
from sparknlp_jsl.annotator import *
from sparknlp.base import *
import sparknlp_jsl
import sparknlp
import warnings
warnings.filterwarnings('ignore')
params = {"spark.driver.memory":"16G", # Amount of memory to use for the driver process, i.e. where SparkContext is initialized
"spark.kryoserializer.buffer.max":"2000M", # Maximum allowable size of Kryo serialization buffer, in MiB unless otherwise specified.
"spark.driver.maxResultSize":"2000M"} # Limit of total size of serialized results of all partitions for each Spark action (e.g. collect) in bytes.
# Should be at least 1M, or 0 for unlimited.
spark = sparknlp_jsl.start(license_keys['SECRET'],params=params)
print ("Spark NLP Version :", sparknlp.version())
print ("Spark NLP_JSL Version :", sparknlp_jsl.version())
# + [markdown] id="PeWhvSsHRXsf"
# ## Download Clinical Word Embeddings for training
# + colab={"base_uri": "https://localhost:8080/"} id="Ua5dHNeyRWQc" outputId="19f91b60-fb4f-4d8f-e5c9-306a8e41d292"
clinical_embeddings = WordEmbeddingsModel.pretrained('embeddings_clinical', "en", "clinical/models")\
.setInputCols(["sentence", "token"])\
.setOutputCol("embeddings")
# + [markdown] id="bfT1AaKvOG4J"
# ## Download Data for Training (NCBI Disease Dataset)
# + id="KMOycTPUcty7"
# !wget -q https://raw.githubusercontent.com/JohnSnowLabs/spark-nlp-workshop/master/tutorials/Certification_Trainings/Healthcare/data/NCBI_disease_official_test.conll
# !wget -q https://raw.githubusercontent.com/JohnSnowLabs/spark-nlp-workshop/master/tutorials/Certification_Trainings/Healthcare/data/NCBI_disease_official_train_dev.conll
# + colab={"base_uri": "https://localhost:8080/"} id="FetZIfagc-eh" outputId="66230226-4d74-405d-f8ca-15f56933b976"
from sparknlp.training import CoNLL
training_data = CoNLL().readDataset(spark, 'NCBI_disease_official_train_dev.conll')
training_data.show(3)
# + colab={"base_uri": "https://localhost:8080/"} id="Fm1bfPJidC9X" outputId="a246fbd1-6552-4fd2-eb5e-addfbc318990"
from sparknlp.training import CoNLL
test_data = CoNLL().readDataset(spark, 'NCBI_disease_official_test.conll')
test_data.show(3)
# + [markdown] id="AaTp5atqc_C-"
# ## Split the test data into two parts:
# - We Keep the first part separate and use it for training the model further, as it will be totally unseen data from the same taxonomy.
#
# - The second part will be used to testing and evaluating
# + id="D_gInTNZRhnC"
(test_data_1, test_data_2) = test_data.randomSplit([0.5, 0.5], seed = 100)
# save the test data as parquet for easy testing
clinical_embeddings.transform(test_data_1).write.parquet('test_1.parquet')
clinical_embeddings.transform(test_data_2).write.parquet('test_2.parquet')
# + [markdown] id="WX2jO2FPRo6Q"
# ## Train a new model, pause, and resume training on the same dataset.
# + [markdown] id="tc2Uoh_YR3dA"
# ### Create a graph
# + colab={"base_uri": "https://localhost:8080/"} id="WFzmo-JzRs3x" outputId="98be816b-cafe-40ee-b9c6-eca1bda8a9db"
from sparknlp_jsl.training import tf_graph
# %tensorflow_version 1.x
tf_graph.print_model_params("ner_dl")
tf_graph.build("ner_dl", build_params={"embeddings_dim": 200, "nchars": 128, "ntags": 12, "is_medical": 1}, model_location="./medical_ner_graphs", model_filename="auto")
# + [markdown] id="PT5pF7V7R69G"
# ### Train for 2 epochs
# + id="X7J4j069R9ty"
nerTagger = MedicalNerApproach()\
.setInputCols(["sentence", "token", "embeddings"])\
.setLabelColumn("label")\
.setOutputCol("ner")\
.setMaxEpochs(2)\
.setLr(0.003)\
.setBatchSize(8)\
.setRandomSeed(0)\
.setVerbose(1)\
.setEvaluationLogExtended(True) \
.setEnableOutputLogs(True)\
.setIncludeConfidence(True)\
.setTestDataset('./test_2.parquet')\
.setGraphFolder('./medical_ner_graphs')\
.setOutputLogsPath('./ner_logs')
ner_pipeline = Pipeline(stages=[
clinical_embeddings,
nerTagger
])
# + colab={"base_uri": "https://localhost:8080/"} id="u5Phv3rFSBR_" outputId="1495b5a5-98fd-4052-aac3-339941431635"
# %%time
ner_model = ner_pipeline.fit(training_data)
# + colab={"base_uri": "https://localhost:8080/"} id="sPZ2tmHFSBUy" outputId="53c9b45b-0a47-40f9-ccab-3b7acdda1029"
# Training Logs
# ! cat ner_logs/MedicalNerApproach_a6231a3fe051.log
# + id="p2onzNGuP_QH"
# Logs of 4 consecutive epochs to compare with 2+2 epochs on separate datasets from same taxonomy
# #!cat ner_logs/MedicalNerApproach_4d3d69967c3f.log
# + [markdown] id="qLKJpTfKWLwC"
# ### Evaluate
# + colab={"base_uri": "https://localhost:8080/"} id="Ec5aux5DSBXE" outputId="1187fdc8-3efe-45f0-f6dc-b938ae76cd08"
from sparknlp_jsl.eval import NerDLMetrics
import pyspark.sql.functions as F
pred_df = ner_model.stages[1].transform(clinical_embeddings.transform(test_data_2))
evaler = NerDLMetrics(mode="full_chunk", dropO=True)
eval_result = evaler.computeMetricsFromDF(pred_df.select("label","ner"), prediction_col="ner", label_col="label").cache()
eval_result.withColumn("precision", F.round(eval_result["precision"],4))\
.withColumn("recall", F.round(eval_result["recall"],4))\
.withColumn("f1", F.round(eval_result["f1"],4)).show(100)
print(eval_result.selectExpr("avg(f1) as macro").show())
print (eval_result.selectExpr("sum(f1*total) as sumprod","sum(total) as sumtotal").selectExpr("sumprod/sumtotal as micro").show())
# + [markdown] id="77HQ3yYyT-RZ"
# ### Save the model to disk
# + id="_KEUa1yaSBZW"
ner_model.stages[1].write().overwrite().save('models/NCBI_NER_2_epoch')
# + [markdown] id="y_U4mQcXUTg7"
# ### Train using the saved model on unseen dataset
# #### We use unseen data from same taxonomy
# + id="dgFviuf5SBei"
nerTagger = MedicalNerApproach()\
.setInputCols(["sentence", "token", "embeddings"])\
.setLabelColumn("label")\
.setOutputCol("ner")\
.setMaxEpochs(2)\
.setLr(0.003)\
.setBatchSize(8)\
.setRandomSeed(0)\
.setVerbose(1)\
.setEvaluationLogExtended(True) \
.setEnableOutputLogs(True)\
.setIncludeConfidence(True)\
.setTestDataset('/content/test_2.parquet')\
.setOutputLogsPath('ner_logs')\
.setGraphFolder('medical_ner_graphs')\
.setPretrainedModelPath("models/NCBI_NER_2_epoch") ## load exisitng model
ner_pipeline = Pipeline(stages=[
clinical_embeddings,
nerTagger
])
# + colab={"base_uri": "https://localhost:8080/"} id="xVZGKEsKUSrS" outputId="f3a7aa6b-2c67-42e1-e14f-1d490305816f"
# %%time
ner_model_retrained = ner_pipeline.fit(test_data_1)
# + colab={"base_uri": "https://localhost:8080/"} id="zVTfOVnIgwpP" outputId="ed866f20-b0fa-49da-ff9b-5ca70a9086f0"
# !cat ./ner_logs/MedicalNerApproach_f7726480b5ef.log
# + colab={"base_uri": "https://localhost:8080/"} id="K_vg2Ajse6aZ" outputId="52d09ee5-6c55-48ea-b150-804b3bfdfdf9"
from sparknlp_jsl.eval import NerDLMetrics
import pyspark.sql.functions as F
pred_df = ner_model_retrained.stages[1].transform(clinical_embeddings.transform(test_data_2))
evaler = NerDLMetrics(mode="full_chunk", dropO=True)
eval_result = evaler.computeMetricsFromDF(pred_df.select("label","ner"), prediction_col="ner", label_col="label").cache()
eval_result.withColumn("precision", F.round(eval_result["precision"],4))\
.withColumn("recall", F.round(eval_result["recall"],4))\
.withColumn("f1", F.round(eval_result["f1"],4)).show(100)
print(eval_result.selectExpr("avg(f1) as macro").show())
print (eval_result.selectExpr("sum(f1*total) as sumprod","sum(total) as sumtotal").selectExpr("sumprod/sumtotal as micro").show())
# + [markdown] id="rw0jTK2RbMAi"
# ## Now let's take a model trained on a different dataset and train on this dataset
# + colab={"base_uri": "https://localhost:8080/"} id="U2eVAVhfUSt0" outputId="80567602-414f-4b88-8341-54a311ce2308"
jsl_ner = MedicalNerModel.pretrained('ner_jsl','en','clinical/models')
jsl_ner.getClasses()
# + [markdown] id="EL0NQ88ofMur"
# ### Now train a model using this model as base
# + id="f-zM49XJbb2j"
nerTagger = MedicalNerApproach()\
.setInputCols(["sentence", "token", "embeddings"])\
.setLabelColumn("label")\
.setOutputCol("ner")\
.setMaxEpochs(2)\
.setLr(0.003)\
.setBatchSize(8)\
.setRandomSeed(0)\
.setVerbose(1)\
.setEvaluationLogExtended(True) \
.setEnableOutputLogs(True)\
.setIncludeConfidence(True)\
.setTestDataset('/content/test_2.parquet')\
.setOutputLogsPath('ner_logs')\
.setGraphFolder('medical_ner_graphs')\
.setPretrainedModelPath("/root/cache_pretrained/ner_jsl_en_3.1.0_2.4_1624566960534")\
.setOverrideExistingTags(True) # since the tags do not align, set this flag to true
# do hyperparameter by tuning the params above (max epoch, LR, dropout etc.) to get better results
ner_pipeline = Pipeline(stages=[
clinical_embeddings,
nerTagger
])
# + colab={"base_uri": "https://localhost:8080/"} id="dXKmO-MJbb5N" outputId="54020a1c-c360-4174-a513-e8b8b9a599d5"
# %%time
ner_jsl_retrained = ner_pipeline.fit(training_data)
# + colab={"base_uri": "https://localhost:8080/"} id="ra_btnoXgvnF" outputId="d29e6f4e-1448-490d-ea6f-243e49c47677"
# !cat ./ner_logs/MedicalNerApproach_880049ba07d7.log
# + colab={"base_uri": "https://localhost:8080/"} id="yKeO-Kqcbb7i" outputId="927ad5a4-f1bb-431b-c97b-7b31ce0e64bb"
from sparknlp_jsl.eval import NerDLMetrics
import pyspark.sql.functions as F
pred_df = ner_jsl_retrained.stages[1].transform(clinical_embeddings.transform(test_data_2))
evaler = NerDLMetrics(mode="full_chunk", dropO=True)
eval_result = evaler.computeMetricsFromDF(pred_df.select("label","ner"), prediction_col="ner", label_col="label").cache()
eval_result.withColumn("precision", F.round(eval_result["precision"],4))\
.withColumn("recall", F.round(eval_result["recall"],4))\
.withColumn("f1", F.round(eval_result["f1"],4)).show(100)
print(eval_result.selectExpr("avg(f1) as macro").show())
print (eval_result.selectExpr("sum(f1*total) as sumprod","sum(total) as sumtotal").selectExpr("sumprod/sumtotal as micro").show())
| tutorials/Certification_Trainings/Healthcare/1.5.Resume_MedicalNer_Model_Training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Homework #2
#
# Name: <NAME>
# ## Question 1:
# Define a variable that will be a natural number, greater than 0, that you can change. Create a for-loop that prints out the sum of the natural numbers up to this variable. Make sure you check this a few times with a sum "by-hand". For example, if you're input is 10 then your output should be: 0+1+2+3+4+5+6+7+8+9+10 = 55
#
# Note: "natural number" means no fractions/decimals.
# +
variable = 10 # your user can change this number
mySum = 0 # start off with a zero sum, and then add to this with your for-loop
# +
# for loop code goes here
# -
# ## Question 2:
#
# Define a variable that is a temperature in either Celcius or Fahrenheit as a string that includes the numerical value of the temperature and either a "C" or an "F".
#
# If the temperature is in Fahrenheit, convert it to Celcius. If it isn't either, print out a warning message.
#
# You will probably want to do an explicit type conversion at some point between your sliced input string and a number:
float('66.7')
# +
temp = '15C'
# your code here
# -
# Some examples you can try are:
#
# ```python
# temp = '15C'
# ```
# should print out something like:
#
# `the temperature is 15C`
#
# and
#
# ```python
# temp = '15F'
# ```
# should print out
#
# `the temperature is -9.44444C`
#
# and
#
# ```python
# temp = '15B'
# ```
# should print out
#
# `ERROR: not Celcius or Fahrenheit!`
#
# Note, you can choose a bit how word what gets printed out!
# ## Question 3:
# Building upon question 2 for a list of temperatures to convert and print out each of their conversions.
# +
temperatureList = ['15C', '14F', '66F']
# your conversion code goes here
# -
# For example:
# ```python
# temperatureList = ['15C', '14F', '66F']
# ```
# should print out something like:
#
# `the input temperature of 15C is 15C`
#
# `the input temperature of 14F is -10C`
#
# `the input temperature of 66F is 18.8889C`
#
# Don't forget to check for temperature formatting errors like in question 2!
#
# ## Question 4:
#
# This question will require you to look up indexing in NumPy arrays. What will you search for (Google/other search engine) to find what you need? You can also ask the instructor for suggestions -- this is a totally fine way to solve this problem!
#
#
# Loop through the 3rd dimension of the "y" array below and plot it with respect to x:
# +
import numpy as np
import matplotlib.pyplot as plt
# how many x-values do we want?
numX = 10
x = np.arange(numX) # this makes an array of ints from 0-10
# how many lines do we want in our plot?
numLines = 5
y = np.random.randint(0,100,size=[numX,numLines]) # pulls ints in range 0-100
# -
| lesson02/.ipynb_checkpoints/example_hw_lesson02-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="QFWpPc5bbpzK"
# Install libraries
# %%capture
# ! pip install git+https://github.com/microsoft/FLAML.git#egg=flaml[catboost]
# ! pip install optuna
# ! pip install ray[tune]
# + id="8o_tuhj3AnHt"
# Download the dataset
# %%capture
# ! rm -rf *
# ! wget http://172.16.31.10/hotels.zip
# ! unzip hotels.zip
# ! rm hotels.zip
# + id="RgH_i_Ka5DZZ"
# Import libraries
# %%capture
from tqdm.notebook import tqdm
import pandas as pd
import matplotlib
import numpy as np
from pathlib import Path
import pickle
import gc
import datetime
# Machine learning libraries
from flaml import AutoML
from sklearn.metrics import roc_auc_score
# Pandas settings to show more columns are rows in the jupyter notebook
pd.set_option('display.max_rows', 500)
pd.set_option('display.max_columns', 50000)
# + id="JA5tj-IOdTXl"
# Set variables
data_dir = Path("hotels")
train_file = data_dir/"train.csv"
test_file = data_dir/"test.csv"
scoring = "roc_auc"
target_name = "is_booking"
base_date = datetime.datetime(2020,10,1)
# + colab={"base_uri": "https://localhost:8080/"} id="gmCGuZg49pps" executionInfo={"status": "ok", "timestamp": 1637656263551, "user_tz": -210, "elapsed": 78763, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhjlqmzHBsiQ0120I9A35ROA3i9Us5XI8V7QDuYtQ=s64", "userId": "05742314090925954056"}} outputId="c10935ce-0ca6-4469-c7d8-7d96daf90a86"
# Read the raw training data
raw_df = pd.read_csv(train_file, parse_dates=["search_date","checkIn_date","checkOut_date"],
dtype={'is_booking':bool, "is_package": bool, "is_mobile": bool,
'n_adults':'int8','n_children':'int8','n_rooms':'int8'
})
# column types are:
raw_df.dtypes
# + colab={"base_uri": "https://localhost:8080/"} id="nlN3IYgZBnMB" executionInfo={"status": "ok", "timestamp": 1637656273006, "user_tz": -210, "elapsed": 9467, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhjlqmzHBsiQ0120I9A35ROA3i9Us5XI8V7QDuYtQ=s64", "userId": "05742314090925954056"}} outputId="c04c7e0f-e913-4c59-afe7-212e25fcf236"
# Select only subset of it (last two months)
df = raw_df[raw_df.search_date>=base_date]
del raw_df
gc.collect()
# Remove user ids
df.drop(columns=["user"], inplace=True)
# Change string to int
for col in ["channel","destination","hotel_category"]:
df[col] = df[col].apply(lambda x: x[1:])
for col in ["destination"]:
df[col] = df[col].astype('uint32')
for col in ["channel","hotel_category"]:
df[col] = df[col].astype('uint8')
# Column types are:
df.dtypes
# + colab={"base_uri": "https://localhost:8080/", "height": 345} id="2cjLyBN_R0Kk" executionInfo={"status": "ok", "timestamp": 1637656273008, "user_tz": -210, "elapsed": 103, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhjlqmzHBsiQ0120I9A35ROA3i9Us5XI8V7QDuYtQ=s64", "userId": "05742314090925954056"}} outputId="78f02b7f-e44c-42ca-f23e-719bc4ab85ea"
# Print some statistics
print(f"Number of rows is : {df.shape[0]}")
print(f"Booking percentage is : {round(100*df[df.is_booking==True].shape[0]/df.shape[0],2)}%")
df.head()
# + colab={"base_uri": "https://localhost:8080/"} id="ibul2fX1B4SB" executionInfo={"status": "ok", "timestamp": 1637656273009, "user_tz": -210, "elapsed": 56, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhjlqmzHBsiQ0120I9A35ROA3i9Us5XI8V7QDuYtQ=s64", "userId": "05742314090925954056"}} outputId="4bec91df-43f9-477b-c8df-841553c375e5"
# Check for missing values
pd.isnull(df).sum()
# + colab={"base_uri": "https://localhost:8080/"} id="17N4brAjRri8" executionInfo={"status": "ok", "timestamp": 1637656273524, "user_tz": -210, "elapsed": 563, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhjlqmzHBsiQ0120I9A35ROA3i9Us5XI8V7QDuYtQ=s64", "userId": "05742314090925954056"}} outputId="90a72bfa-95c7-4646-9cbb-b956df9a9fb1"
# Drop missing values, as they are a few of them (compared to the dataset size)
df.dropna(subset=['checkIn_date', 'checkOut_date'], inplace=True)
pd.isnull(df).sum()
# + colab={"base_uri": "https://localhost:8080/"} id="glxmXCFy6WNw" executionInfo={"status": "ok", "timestamp": 1637656274035, "user_tz": -210, "elapsed": 525, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhjlqmzHBsiQ0120I9A35ROA3i9Us5XI8V7QDuYtQ=s64", "userId": "05742314090925954056"}} outputId="619e864b-f9c4-4660-b457-791623153e1b"
# Split the training and validation dataset
df_len = df.shape[0]
data_train = df[:int(df_len*0.8)]
data_valid = df[int(df_len*0.8):]
print(f"Train booking percentage is : {round(100*data_train[data_train.is_booking==True].shape[0]/data_train.shape[0],2)}%")
print(f"Validation Booking percentage is : {round(100*data_valid[data_valid.is_booking==True].shape[0]/data_valid.shape[0],2)}%")
del df
y = data_train[target_name]
xs = data_train.drop(columns=target_name)
del data_train
valid_y = data_valid[target_name]
valid_xs = data_valid.drop(columns=target_name)
del data_valid
# + [markdown] id="cB3l-fMptKCD"
# We did not do any feature engineering, but we could have done:
#
# * Days to go (checkIn - searchDate)
# * Number of days to stay (checkOut - checkIn)
# * Number of people (number of adults + number of children)
# * Person per room (Number of people divide by Number of rooms)
# * combination of is_mobile and is_package columns
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 143, "referenced_widgets": ["195693a49a3b473fb60da89ac22f22fc", "8528f103b1ed454a9a987f1a563dfc14", "c8db3715b3ff4f9a924105210c9c50e7", "a558ab6dd2054463a7e63c7531a0c9ab", "d13dec52d0b24af9ae6e2abb2cce3af6", "ac14e94dfe3d4df4abc7a5a882117d96", "41bd2ff3c06c4c8586e75bc1058ca95e", "93ca07cb07c84c00a37fb3998bf44492", "<KEY>", "26277ad96961483b838294c21e70924b", "c01fa1086aae4c2ea27e8a8a4e9a9011"]} id="rgncGUXV7sDw" executionInfo={"status": "ok", "timestamp": 1637660174058, "user_tz": -210, "elapsed": 184083, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhjlqmzHBsiQ0120I9A35ROA3i9Us5XI8V7QDuYtQ=s64", "userId": "05742314090925954056"}} outputId="1969b119-7be1-4bbc-cc67-f4a8fbe51f29"
# Evalaution metric function
def calc_auc_roc(y, prob_pred):
return roc_auc_score(y, prob_pred)
# Initialize an AutoML instance
gc.collect()
performance_list = []
model_names = {"lgbm":"Light Gradient Boosting", "catboost": "Cat Boost"}
for key in tqdm(model_names):
model = AutoML()
# Specify automl goal and constraint
settings = {
"time_budget": 1800,
"metric": scoring,
"task": 'classification',
"verbose": 0,
"estimator_list": [key],
"n_jobs": -1,
"eval_method": "cv",
"n_splits": 5,
"mem_thres": 11294967296,
}
model.fit(xs, y, **settings)
# Save the model
with open(f'{key}.pkl', 'wb') as f:
pickle.dump(model, f, pickle.HIGHEST_PROTOCOL)
# Log the performance
performance = {}
performance["model"] = model_names[key]
train_pred_proba = model.predict_proba(xs)[:,1]
performance["training (auc_roc)"] = calc_auc_roc(y, train_pred_proba)
validation_pred_proba = model.predict_proba(valid_xs)[:,1]
performance["validation (auc_roc)"] = calc_auc_roc(valid_y, validation_pred_proba)
performance_list.append(performance)
gc.collect()
performance_df = pd.DataFrame(performance_list).round(3)
display(performance_df)
# + id="J1jaX2jH-xg0" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1637660302752, "user_tz": -210, "elapsed": 487, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhjlqmzHBsiQ0120I9A35ROA3i9Us5XI8V7QDuYtQ=s64", "userId": "05742314090925954056"}} outputId="94cf9e51-e26b-493d-8dbf-2d36e6532169"
best_model_name = "lgbm"
with open(f'{best_model_name}.pkl', 'rb') as f:
selected_model = pickle.load(f)
print(f"selected model is {best_model_name}.\n")
print("Its parameters are:")
selected_model.model.get_params()
# + id="fN1KUyJlYroS" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1637660332274, "user_tz": -210, "elapsed": 1053, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhjlqmzHBsiQ0120I9A35ROA3i9Us5XI8V7QDuYtQ=s64", "userId": "05742314090925954056"}} outputId="cea4be34-3a43-49e3-9091-4d9aab0c1527"
test_df = pd.read_csv(test_file, parse_dates=["search_date","checkIn_date","checkOut_date"],
dtype={'is_booking':bool, "is_package": bool, "is_mobile": bool,
'n_adults':'int8','n_children':'int8','n_rooms':'int8'
})
# Remove user ids
test_df.drop(columns=["user"], inplace=True)
# Change string to int
for col in ["channel", "destination", "hotel_category"]:
test_df[col] = test_df[col].apply(lambda x: x[1:])
for col in ["destination"]:
test_df[col] = test_df[col].astype('uint32')
for col in ["channel", "hotel_category"]:
test_df[col] = test_df[col].astype('uint8')
# Column types are:
test_df.dtypes
# + id="md85TC3OXIPo" colab={"base_uri": "https://localhost:8080/", "height": 204} executionInfo={"status": "ok", "timestamp": 1637660879403, "user_tz": -210, "elapsed": 4911, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhjlqmzHBsiQ0120I9A35ROA3i9Us5XI8V7QDuYtQ=s64", "userId": "05742314090925954056"}} outputId="9840d96a-5899-4dfc-beb6-66f4a9946179"
submission_df = pd.DataFrame()
submission_df["prediction"] = selected_model.predict_proba(test_df)[:,1]
submission_df.to_csv("output.csv", index=False)
submission_df.head()
| contests/codecup6_machine_learning/webinar/tabular.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from scipy import stats
import scipy
import matplotlib.pyplot as plt
import pandas as pd
import math
# +
""" Precificação utilizando Black and Scholes.
cp: +1 -> call; -1 put
s: valor da ação
k: strike
t: tempo em dias até expirar a opção
v: volatilidade
rf: taxa de juros neutra risco
"""
def black_scholes (cp, s, k, t, rf, v, div = 0):
d1 = (math.log(s/k)+(rf+0.5*math.pow(v,2))*t)/(v*math.sqrt(t))
d2 = d1 - v*math.sqrt(t)
optprice = (cp*s*math.exp(-div*t)*stats.norm.cdf(cp*d1)) - (cp*k*math.exp(-rf*t)*stats.norm.cdf(cp*d2))
return optprice
""" Calculo da volatilidade implicita
cp: +1 -> call; -1 put
s: valor da ação
k: strike
t: tempo em dias até expirar a opção
rf: taxa de juros neutra risco
price: cotação da opção pelo mercado
Função H(vol): Seja B(vol) o valor calculado por B&S dada volatilidade, e P a cotação da opção, H(vol) = B(vol) - P
É a função a ser usada na bisseção
"""
def volat_impl(cp, s, k, t, rf, price):
def h(vol):
return black_scholes(cp, s, k, t, rf, vol) - price
return scipy.optimize.bisect(h,1e-6,5,xtol=1e-16)
# -
data = pd.read_csv('BRFOODS.csv')
# +
'''
Setting CONSTANTS
'''
sigla_acao = 'BRFS3'
empresa = 'BRFoods S.A.'
preco_acao = 27.00
dias = [6,28]
puts = data['Tipo'] == 'PUT'
calls = data['Tipo'] == 'CALL'
dia1706 = data['TF'] == '17-06-2019'
dia1507 = data['TF'] == '15-07-2019'
# +
## PUT Com 6 dias a vencer
## Buscando as informações no DataFrame
df_k = data[puts & dia1706].iloc[0:,2:3]
df_s = data[puts & dia1706].iloc[0:,3:4]
ks_put_6 = df_k.values.flatten()
Ss_put_6 = df_s.values.flatten()
## Setando o array com as volatilidades a serem plotadas
vs_put_6 = []
for (k,s) in zip(ks_put_6,Ss_put_6):
vs_put_6.append(volat_impl(-1,preco_acao,k,dias[0]/365,0.065,s))
## Plot do gráfico
plt.figure(figsize=(10,6))
plt.plot(ks_put_6,vs_put_6, marker='o', linestyle='--', color='g', markerfacecolor='r')
plt.xlabel('Strikes - em R$')
plt.ylabel('Volatilidade')
plt.title('Gráfico Smile - Volatilidade x Strike - PUT Com 6 dias a vencer')
plt.grid()
plt.show()
# +
## CALL Com 6 dias a vencer
## Buscando as informações no DataFrame
df_k = data[calls & dia1706].iloc[0:,2:3]
df_s = data[calls & dia1706].iloc[0:,3:4]
ks_call_6 = df_k.values.flatten()
Ss_call_6 = df_s.values.flatten()
## Setando o array com as volatilidades a serem plotadas
vs_call_6 = []
for (k,s) in zip(ks_call_6,Ss_call_6):
vs_call_6.append(volat_impl(1,preco_acao,k,dias[0]/365,0.065,s))
## Plot do gráfico
plt.figure(figsize=(10,6))
plt.plot(ks_call_6,vs_call_6, marker='o', linestyle='--', color='g', markerfacecolor='r')
plt.xlabel('Strikes - em R$')
plt.ylabel('Volatilidade')
plt.title('Gráfico Smile - Volatilidade x Strike - CALL Com 6 dias a vencer')
plt.grid()
plt.show()
# +
## PUT Com 28 dias a vencer
## Buscando as informações no DataFrame
df_k = data[puts & dia1507].iloc[0:,2:3]
df_s = data[puts & dia1507].iloc[0:,3:4]
ks_put_28 = df_k.values.flatten()
Ss_put_28 = df_s.values.flatten()
## Setando o array com as volatilidades a serem plotadas
vs_put_28 = []
for (k,s) in zip(ks_put_28,Ss_put_28):
vs_put_28.append(volat_impl(-1,preco_acao,k,dias[1]/365,0.065,s))
## Plot do gráfico
plt.figure(figsize=(10,6))
plt.plot(ks_put_28,vs_put_28, marker='o', linestyle='--', color='g', markerfacecolor='r')
plt.xlabel('Strikes - em R$')
plt.ylabel('Volatilidade')
plt.title('Gráfico Smile - Volatilidade x Strike - PUT com 28 dias a vencer')
plt.grid()
plt.show()
# +
## CALL Com 28 dias a vencer
## Buscando as informações no DataFrame
df_k = data[calls & dia1507].iloc[0:,2:3]
df_s = data[calls & dia1507].iloc[0:,3:4]
ks_call_28 = df_k.values.flatten()
Ss_call_28 = df_s.values.flatten()
## Setando o array com as volatilidades a serem plotadas
vs_call_28 = []
for (k,s) in zip(ks_call_28,Ss_call_28):
vs_call_28.append(volat_impl(1,preco_acao,k,dias[1]/365,0.065,s))
## Plot do gráfico
plt.figure(figsize=(10,6))
plt.plot(ks_call_28,vs_call_28, marker='o', linestyle='--', color='g', markerfacecolor='r')
plt.xlabel('Strikes - em R$')
plt.ylabel('Volatilidade')
plt.title('Gráfico Smile - Volatilidade x Strike - CALL com 28 dias a vencer')
plt.grid()
plt.show()
# -
# ---
# ---
| mod-mat-financas-I-2019-1/Project_3/Project3_almost.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # Pooling
#
# As we process images (or other data sources) we will eventually want to reduce the resolution of the images. After all, we typically want to output an estimate that does not depend on the dimensionality of the original image. Secondly, when detecting lower-level features, such as edge detection (we covered this in the section on [convolutional layers](conv-layer.md)), we often want to have some degree of invariance to translation. For instance, if we take the image `X` with a sharp delineation between black and white and if we shift it by one pixel to the right, i.e. `Z[i,j] = X[i,j+1]`, then the output for for the new image `Z` will be vastly different. The edge will have shifted by one pixel and with it all the activations. In reality objects hardly ever occur exactly at the same place. In fact, even with a tripod and a stationary object, vibration of the camera due to the movement of the shutter might shift things by a pixel or so (this is why high end cameras have a special option to fix this). Given that, we need a mathematical device to address the problem.
#
# This section introduces pooling layers, which were proposed to alleviate the excessive sensitivity of the convolutional layer to location and to reduce the resolution of images through the processing pipeline.
#
# ## Maximum Pooling and Average Pooling
#
# Like convolutions, pooling computes the output for each element in a fixed-shape window (also known as a pooling window) of input data. Different from the cross-correlation computation of the inputs and kernels in the convolutional layer, the pooling layer directly calculates the maximum or average value of the elements in the pooling window. These operations are called maximum pooling or average pooling respectively. In maximum pooling, the pooling window starts from the top left of the input array, and slides in the input array from left to right and top to bottom. When the pooling window slides to a certain position, the maximum value of the input subarray in the window is the element at the corresponding location in the output array.
#
# 
#
# The output array in the figure above has a height of 2 and a width of 2. The four elements are derived from the maximum value of $\text{max}$:
#
# $$
# \max(0,1,3,4)=4,\\
# \max(1,2,4,5)=5,\\
# \max(3,4,6,7)=7,\\
# \max(4,5,7,8)=8.\\
# $$
#
# Average pooling works like maximum pooling, only with the maximum operator replaced by the average operator. The pooling layer with a pooling window shape of $p \times q$ is called the $p \times q$ pooling layer. The pooling operation is called $p \times q$ pooling.
#
# Let us return to the object edge detection example mentioned at the beginning of this section. Now we will use the output of the convolutional layer as the input for $2\times 2$ maximum pooling. Set the convolutional layer input as `X` and the pooling layer output as `Y`. Whether or not the values of `X[i, j]` and `X[i, j+1]` are different, or `X[i, j+1]` and `X[i, j+2]` are different, the pooling layer outputs all include `Y[i, j]=1`. That is to say, using the $2\times 2$ maximum pooling layer, we can still detect if the pattern recognized by the convolutional layer moves no more than one element in height and width.
#
# As shown below, we implement the forward computation of the pooling layer in the `pool2d` function. This function is very similar to the `corr2d` function in the section on [convolutions](conv-layer.md). The only difference lies in the computation of the output `Y`.
# + attributes={"classes": [], "id": "", "n": "11"}
from mxnet import nd
from mxnet.gluon import nn
def pool2d(X, pool_size, mode='max'):
p_h, p_w = pool_size
Y = nd.zeros((X.shape[0] - p_h + 1, X.shape[1] - p_w + 1))
for i in range(Y.shape[0]):
for j in range(Y.shape[1]):
if mode == 'max':
Y[i, j] = X[i: i + p_h, j: j + p_w].max()
elif mode == 'avg':
Y[i, j] = X[i: i + p_h, j: j + p_w].mean()
return Y
# -
# We can construct the input array `X` in the above diagram to validate the output of the two-dimensional maximum pooling layer.
# + attributes={"classes": [], "id": "", "n": "13"}
X = nd.array([[0, 1, 2], [3, 4, 5], [6, 7, 8]])
pool2d(X, (2, 2))
# -
# At the same time, we experiment with the average pooling layer.
# + attributes={"classes": [], "id": "", "n": "14"}
pool2d(X, (2, 2), 'avg')
# -
# ## Padding and Stride
#
# Like the convolutional layer, the pooling layer can also change the output shape by padding the two sides of the input height and width and adjusting the window stride. The pooling layer works in the same way as the convolutional layer in terms of padding and strides. We will demonstrate the use of padding and stride in the pooling layer through the two-dimensional maximum pooling layer MaxPool2D in the `nn` module. We first construct an input data of shape `(1, 1, 4, 4)`, where the first two dimensions are batch and channel.
# + attributes={"classes": [], "id": "", "n": "15"}
X = nd.arange(16).reshape((1, 1, 4, 4))
X
# -
# By default, the stride in the `MaxPool2D` class has the same shape as the pooling window. Below, we use a pooling window of shape `(3, 3)`, so we get a stride shape of `(3, 3)` by default.
# + attributes={"classes": [], "id": "", "n": "16"}
pool2d = nn.MaxPool2D(3)
# Because there are no model parameters in the pooling layer, we do not need
# to call the parameter initialization function
pool2d(X)
# -
# The stride and padding can be manually specified.
# + attributes={"classes": [], "id": "", "n": "7"}
pool2d = nn.MaxPool2D(3, padding=1, strides=2)
pool2d(X)
# -
# Of course, we can specify an arbitrary rectangular pooling window and specify the padding and stride for height and width, respectively.
# + attributes={"classes": [], "id": "", "n": "8"}
pool2d = nn.MaxPool2D((2, 3), padding=(1, 2), strides=(2, 3))
pool2d(X)
# -
# ## Multiple Channels
#
# When processing multi-channel input data, the pooling layer pools each input channel separately, rather than adding the inputs of each channel by channel as in a convolutional layer. This means that the number of output channels for the pooling layer is the same as the number of input channels. Below, we will concatenate arrays `X` and `X+1` on the channel dimension to construct an input with 2 channels.
# + attributes={"classes": [], "id": "", "n": "9"}
X = nd.concat(X, X + 1, dim=1)
X
# -
# As we can see, the number of output channels is still 2 after pooling.
# + attributes={"classes": [], "id": "", "n": "10"}
pool2d = nn.MaxPool2D(3, padding=1, strides=2)
pool2d(X)
# -
# ## Summary
#
# * Taking the input elements in the pooling window, the maximum pooling operation assigns the maximum value as the output and the average pooling operation assigns the average value as the output.
# * One of the major functions of a pooling layer is to alleviate the excessive sensitivity of the convolutional layer to location.
# * We can specify the padding and stride for the pooling layer.
# * Maximum pooling, combined with a stride larger than 1 can be used to reduce the resolution.
# * The pooling layer's number of output channels is the same as the number of input channels.
#
#
# ## Exercises
#
# 1. Implement average pooling as a convolution.
# 1. What is the computational cost of the pooling layer? Assume that the input to the pooling layer is of size $c\times h\times w$, the pooling window has a shape of $p_h\times p_w$ with a padding of $(p_h, p_w)$ and a stride of $(s_h, s_w)$.
# 1. Why do you expect maximum pooling and average pooling to work differently?
# 1. Do we need a separate minimum pooling layer? Can you replace it with another operation?
# 1. Is there another operation between average and maximum pooling that you could consider (hint - recall the softmax)? Why might it not be so popular?
#
# ## Scan the QR Code to [Discuss](https://discuss.mxnet.io/t/2352)
#
# 
| book-d2l-en/chapter_convolutional-neural-networks/pooling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.10 64-bit (''CoinbaseBot_Env'': venv)'
# name: python3
# ---
# # Evolution of trade
#
# ### The evolution of trades corresponds to the criteria estimating the possible profites of having a trade at a certain time.
# ### It corresponds to the choice of buying or selling a trade
# ## Loading Database
# +
# Load of database
import numpy as np
from database import Historic_coinbase_dtb
df = Historic_coinbase_dtb.load()
print(df.columns) # All cryptos studied by the bot
# -
# ## Initialization Evolution
# 1rst proposed Method
from algorithms.evolution.evolution_trade import Evolution_Trade_Median
evolution_method = Evolution_Trade_Median(start_check_future=60, end_check_future=120)
# ## Display Data
# +
# Generate data
crypto_name_test = 'ALGO-USD'
trade_value = df[crypto_name_test]
## Convert array to be conform with method
trade_adapt = trade_value.to_numpy()
trade_adapt = trade_adapt.reshape((trade_adapt.shape[0], -1))
evolution = evolution_method.get_evolution(trade_adapt)
# +
# Display
# %matplotlib notebook
# # %matplotlib inline
import matplotlib.pyplot as plt
fig, ax = plt.subplots(2,1,sharex=True)
ax[0].plot(trade_value)
ax[1].plot(df.index, evolution)
# plt.show()
# -
# WARNING: Interactive Dipslay is not available on VSCode
| Python_Bot/algorithms/evolution/note_evolution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
# +
# Pip install selenium and beautifulsoup4. Then download
# ChromeDriver https://sites.google.com/a/chromium.org/chromedriver/downloads
# +
from bs4 import BeautifulSoup
from selenium import webdriver
# replace this with the path of where you downloaded chromedriver
chromedriver_path = "/Users/alexcombs/Downloads/chromedriver"
browser = webdriver.Chrome(chromedriver_path)
# -
sats = 'https://www.google.com/flights/?f=0#f=0&flt=/m/02_286.r/m/02j9z.2018-12-01*r/m/02j9z./m/02_286.2018-12-08;c:USD;e:1;s:0*1;sd:1;t:e'
browser.get(sats)
browser.title
browser.save_screenshot('/Users/alexcombs/Desktop/test_flights.png')
from IPython.core.display import HTML
HTML(browser.page_source)
from bs4 import BeautifulSoup
soup = BeautifulSoup(browser.page_source, "html5lib")
cards = soup.select('div[class*=info-container]')
cards[0]
for card in cards:
print(card.select('h3')[0].text)
print(card.select('span[class*=price]')[0].text)
print('\n')
# +
# we'll get fares for all saturday to saturday flights to europe for the next 10 weeks
# we'll then create a time series and look for outliers
# -
from datetime import date, timedelta
from time import sleep
# +
start_sat = '2018-12-01'
end_sat = '2018-12-08'
start_sat_date = datetime.strptime(start_sat, '%Y-%m-%d')
end_sat_date = datetime.strptime(end_sat, '%Y-%m-%d')
fare_dict = {}
for i in range(26):
sat_start = str(start_sat_date).split()[0]
sat_end = str(end_sat_date).split()[0]
fare_dict.update({sat_start: {}})
sats = "https://www.google.com/flights/?f=0#f=0&flt=/m/02_286.r/m/02j9z." + \
sat_start + "*r/m/02j9z./m/02_286." + \
sat_end + ";c:USD;e:1;s:0*1;sd:1;t:e"
sleep(np.random.randint(3,7))
browser.get(sats)
soup = BeautifulSoup(browser.page_source, "html5lib")
cards = soup.select('div[class*=info-container]')
for card in cards:
city = card.select('h3')[0].text
fare = card.select('span[class*=price]')[0].text
fare_dict[sat_start] = {**fare_dict[sat_start], **{city: fare}}
start_sat_date = start_sat_date + timedelta(days=7)
end_sat_date = end_sat_date + timedelta(days=7)
# -
fare_dict
city_key = 'Milan'
for key in fare_dict:
print(key, fare_dict[key][city_key])
city_dict = {}
for k,v in fare_dict.items():
city_dict.update({k:int(v[city_key].replace(',','').split('$')[1])})
city_dict
prices = [int(x) for x in city_dict.values()]
dates = city_dict.keys()
fig,ax = plt.subplots(figsize=(10,6))
plt.scatter(dates, prices, color='black', s=50)
ax.set_xticklabels(dates, rotation=-70);
from PyAstronomy import pyasl
# +
r = pyasl.generalizedESD(prices, 3, 0.025, fullOutput=True)
print('Total Outliers:', r[0])
out_dates = {}
for i in sorted(r[1]):
out_dates.update({list(dates)[i]: list(prices)[i]})
print('Outlier Dates', out_dates.keys(), '\n')
print(' R Lambda')
for i in range(len(r[2])):
print('%2d %8.5f %8.5f' % ((i+1), r[2][i], r[3][i]))
fig, ax = plt.subplots(figsize=(10,6))
plt.scatter(dates, prices, color='black', s=50)
ax.set_xticklabels(dates, rotation=-70);
for i in range(r[0]):
plt.plot(r[1][i], prices[r[1][i]], 'rp')
# -
city_mean = np.mean(list(city_dict.values()))
for k,v in out_dates.items():
if v < city_mean:
print('Alert for', city_key + '!')
print('Fare: $' + str(v), 'on', k)
print('\n')
from scipy import stats
fix, ax = plt.subplots(figsize=(10,6))
stats.probplot(list(city_dict.values()), plot=plt)
plt.show()
| Chapter03/PMLB SE - Chapter 3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Denoising Autoencoders And Where To Find Them
#
# Today we're going to train deep autoencoders and deploy them to faces and search for similar images.
#
# Our new test subjects are human faces from the [lfw dataset](http://vis-www.cs.umass.edu/lfw/).
# +
# !wget https://raw.githubusercontent.com/yandexdataschool/Practical_DL/fall19/week08_autoencoders/lfw_dataset.py -O lfw_dataset.py
# !pip install Image Pillow==5.1.0 scipy==1.1.0
import torch
import numpy as np
from torch.utils.data import Dataset, DataLoader
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
EPOCHS = 100
BATCH_SIZE = 32
LEARNING_RATE = 1e-3
LATENT_DIMENSION = 4
device = torch.device("cpu") # or you print here you favorite gpu card
#torch.set_default_tensor_type(torch.cuda.FloatTensor)
# uses cuda floats by default
print(device)
# -
BATCH_SIZE = 32
# +
import numpy as np
from lfw_dataset import fetch_lfw_dataset
from sklearn.model_selection import train_test_split
X, attr = fetch_lfw_dataset(use_raw=True,dimx=38,dimy=38)
X = X.transpose([0,3,1,2]).astype('float32') / 256.0
img_shape = X.shape[1:]
X_train, X_test = train_test_split(X, test_size=0.1,random_state=42)
# -
X_train_tensor = torch.from_numpy(X_train).type(torch.DoubleTensor)
X_test_tensor = torch.Tensor(X_test).type(torch.DoubleTensor)
img_shape
# +
# %matplotlib inline
import matplotlib.pyplot as plt
plt.title('sample image')
for i in range(6):
plt.subplot(2,3,i+1)
plt.imshow(X[i].transpose([1,2,0]))
print("X shape:",X.shape)
print("attr shape:",attr.shape)
# -
# ### Autoencoder architecture
#
# Let's design autoencoder as a single lasagne network, going from input image through bottleneck into the reconstructed image.
#
# <img src="http://nghiaho.com/wp-content/uploads/2012/12/autoencoder_network1.png" width=640px>
#
#
# ## First step: PCA
#
# Principial Component Analysis is a popular dimensionality reduction method.
#
# Under the hood, PCA attempts to decompose object-feature matrix $X$ into two smaller matrices: $W$ and $\hat W$ minimizing _mean squared error_:
#
# $$\|(X W) \hat{W} - X\|^2_2 \to_{W, \hat{W}} \min$$
# - $X \in \mathbb{R}^{n \times m}$ - object matrix (**centered**);
# - $W \in \mathbb{R}^{m \times d}$ - matrix of direct transformation;
# - $\hat{W} \in \mathbb{R}^{d \times m}$ - matrix of reverse transformation;
# - $n$ samples, $m$ original dimensions and $d$ target dimensions;
#
# In geometric terms, we want to find d axes along which most of variance occurs. The "natural" axes, if you wish.
#
# 
#
#
# PCA can also be seen as a special case of an autoencoder.
#
# * __Encoder__: X -> Dense(d units) -> code
# * __Decoder__: code -> Dense(m units) -> X
#
# Where Dense is a fully-connected layer with linear activaton: $f(X) = W \cdot X + \vec b $
#
#
# Note: the bias term in those layers is responsible for "centering" the matrix i.e. substracting mean.
# +
# this class corresponds to view-function and may be used as a reshape layer
class View(nn.Module):
def __init__(self, *shape):
super(View, self).__init__()
self.shape = shape
def forward(self, input):
return input.view(*self.shape)
# -
class pca_autoencoder(nn.Module):
"""
Here we define a simple linear autoencoder as described above.
We also flatten and un-flatten data to be compatible with image shapes
"""
def __init__(self, code_size=32):
super(pca_autoencoder, self).__init__()
self.enc = nn.Sequential(View(-1, np.prod(img_shape)), nn.Linear(np.prod(img_shape), code_size))
self.dec = nn.Sequential(nn.Linear(code_size, np.prod(img_shape)), View(-1, img_shape[0], img_shape[1], img_shape[2]))
def batch_loss(self, batch, reference):
reconstruction = #<Your code: define reconstruction object>
return torch.mean((reference - reconstruction)**2)
# ### Train the model
#
# As usual, iterate minibatches of data and call train_step, then evaluate loss on validation data.
#
# __Note to py2 users:__ you can safely drop `flush=True` from any code below.
from tqdm import tqdm
def train(model, dataset, dataset_test, num_epoch=32, gd=None, noise_function=None, noise_function_params=None):
model.double()
model.to(device)
if gd is None:
gd = optim.Adamax(model.parameters(), lr=0.002)
if noise_function_params is None:
noise_function_params = {}
dataloader = DataLoader(dataset, batch_size=BATCH_SIZE, shuffle=True)
losses = []
dataloader_test = DataLoader(dataset_test, batch_size=BATCH_SIZE, shuffle=True)
scores = []
for epoch in range(num_epoch):
model.train(True)
for i, (batch) in tqdm(enumerate(dataloader)):
gd.zero_grad()
if noise_function is not None:
batch_noised = noise_function(batch, **noise_function_params).to(device=device)
loss = model.batch_loss(batch_noised, batch.to(device=device))
else:
batch = batch.to(device=device)
loss = model.batch_loss(batch, batch)
(loss).backward()
losses.append(loss.data.cpu().numpy())
gd.step()
gd.zero_grad()
train_mse = np.mean(losses[-(i+1):])
model.train(False)
for i, (batch) in enumerate(dataloader_test):
batch = batch.to(device=device)
scores.append(model.batch_loss(batch, batch).data.cpu().numpy())
test_mse = np.mean(scores[-(i+1):])
print(f"{epoch+1}, Train loss: {train_mse}, Test loss: {test_mse}")
def visualize(img, model):
"""Draws original, encoded and decoded images"""
model.train(False)
code = model.enc(img[None].cuda(device = device))
reco = model.dec(code)
plt.subplot(1,3,1)
plt.title("Original")
plt.imshow(img.cpu().numpy().transpose([1, 2, 0]).clip(0, 1))
plt.subplot(1,3,2)
plt.title("Code")
plt.imshow(code.cpu().detach().numpy().reshape([code.shape[-1] // 2, -1]))
plt.subplot(1,3,3)
plt.title("Reconstructed")
plt.imshow(reco[0].cpu().detach().numpy().transpose([1, 2, 0]).clip(0, 1))
plt.show()
aenc = pca_autoencoder()
train(aenc, X_train_tensor, X_test_tensor, 40)
dataloader_test = DataLoader(X_test_tensor, batch_size=BATCH_SIZE, shuffle=True)
scores = []
for i, (batch) in enumerate(dataloader_test):
batch = batch.to(device=device)
scores.append(aenc.batch_loss(batch, batch).data.cpu().numpy())
print (np.mean(scores))
for i in range(5):
img = X_test_tensor[i]
visualize(img,aenc)
# ### Going deeper
#
# PCA is neat but surely we can do better. This time we want you to build a deep autoencoder by... stacking more layers.
#
# In particular, your encoder and decoder should be at least 3 layers deep each. You can use any nonlinearity you want and any number of hidden units in non-bottleneck layers provided you can actually afford training it.
#
# 
#
# A few sanity checks:
# * There shouldn't be any hidden layer smaller than bottleneck (encoder output).
# * Don't forget to insert nonlinearities between intermediate dense layers.
# * Convolutional layers are allowed but not required. To undo convolution use L.Deconv2D, pooling - L.UpSampling2D.
# * Adding activation after bottleneck is allowed, but not strictly necessary.
class pca_autoencoder_deep(nn.Module):
def __init__(self, code_size=32):
super(pca_autoencoder_deep, self).__init__()
self.enc = #<Your code: define encoder as per instructions above>
self.dec = #<Your code: define decoder as per instructions above>
def batch_loss(self, batch, reference):
a = self.enc(batch)
reconstruction = self.dec(a)
return torch.mean((reference - reconstruction)**2)
#Check autoencoder shapes along different code_sizes
get_dim = lambda layer: np.prod(layer.output_shape[1:])
for code_size in [1,8,32,128,512,1024]:
help_tensor = next(iter(DataLoader(X_train_tensor, batch_size=BATCH_SIZE)))
model = pca_autoencoder_deep(code_size).double().to(device)
encoder_out = model.enc(help_tensor.cuda(device))
decoder_out = model.dec(encoder_out)
print("Testing code size %i" % code_size)
assert encoder_out.shape[1:]==torch.Size([code_size]),"encoder must output a code of required size"
assert decoder_out.shape[1:]==img_shape, "decoder must output an image of valid shape"
assert (sum([1 + len(list(c1level.children())) for c1level in model.dec.children()]) >= 6), "decoder must contain at least 3 dense layers"
del encoder_out
del decoder_out
torch.cuda.empty_cache()
print("All tests passed!")
# __Hint:__ if you're getting "Encoder layer is smaller than bottleneck" error, use code_size when defining intermediate layers.
#
# For example, such layer may have code_size*2 units.
aenc_deep = pca_autoencoder_deep()
train(aenc_deep, X_train_tensor, X_test_tensor, 50)
# Training may take long, it's okay.
# +
aenc_deep.train(False)
dataloader_test = DataLoader(X_test_tensor, batch_size=BATCH_SIZE, shuffle=True)
scores = []
for i, (batch) in enumerate(dataloader_test):
batch = batch.to(device=device)
scores.append(aenc_deep.batch_loss(batch, batch).data.cpu().numpy())
encoder_out = aenc_deep.enc(batch)
reconstruction_mse = np.mean(scores)
assert reconstruction_mse <= 0.0055, "Compression is too lossy. See tips below."
assert len(encoder_out.shape)==2 and encoder_out.shape[1]==32, "Make sure encoder has code_size units"
print("Final MSE:", reconstruction_mse)
for i in range(5):
img = X_test_tensor[i]
visualize(img,aenc_deep)
# -
# __Tips:__ If you keep getting "Compression to lossy" error, there's a few things you might try:
#
# * Make sure it converged. Some architectures need way more than 32 epochs to converge. They may fluctuate a lot, but eventually they're going to get good enough to pass. You may train your network for as long as you want.
#
# * Complexity. If you already have, like, 152 layers and still not passing threshold, you may wish to start from something simpler instead and go in small incremental steps.
#
# * Architecture. You can use any combination of layers (including convolutions, normalization, etc) as long as __encoder output only stores 32 numbers per training object__.
#
# A cunning learner can circumvent this last limitation by using some manual encoding strategy, but he is strongly recommended to avoid that.
# ## Denoising AutoEncoder
#
# Let's now make our model into a denoising autoencoder.
#
# We'll keep your model architecture, but change the way it trains. In particular, we'll corrupt it's input data randomly before each epoch.
#
# There are many strategies to apply noise. We'll implement two popular one: adding gaussian noise and using dropout.
def apply_gaussian_noise(X,sigma=0.1):
"""
adds noise from normal distribution with standard deviation sigma
:param X: image tensor of shape [batch,height,width,3]
"""
<Your code: define noise>
return X + noise
#noise tests
theoretical_std = (X[:100].std()**2 + 0.5**2)**.5
our_std = apply_gaussian_noise(torch.from_numpy(X[:100]),sigma=0.5).std()
assert abs(theoretical_std - our_std) < 0.01, "Standard deviation does not match it's required value. Make sure you use sigma as std."
assert abs(apply_gaussian_noise(torch.from_numpy(X[:100]),sigma=0.5).mean() - torch.from_numpy(X[:100]).mean()) < 0.01, "Mean has changed. Please add zero-mean noise"
plt.subplot(1,4,1)
plt.imshow(X[0].transpose([1,2,0]))
plt.subplot(1,4,2)
plt.imshow(apply_gaussian_noise(torch.from_numpy(X[:1]),sigma=0.01).data.numpy()[0].transpose([1,2,0]).clip(0, 1))
plt.subplot(1,4,3)
plt.imshow(apply_gaussian_noise(torch.from_numpy(X[:1]),sigma=0.1).data.numpy()[0].transpose([1,2,0]).clip(0, 1))
plt.subplot(1,4,4)
plt.imshow(apply_gaussian_noise(torch.from_numpy(X[:1]),sigma=0.5).data.numpy()[0].transpose([1,2,0]).clip(0, 1))
aenc = pca_autoencoder()
train(aenc, X_train_tensor, X_test_tensor, 50, noise_function=apply_gaussian_noise)
# __Note:__ if it hasn't yet converged, increase the number of iterations.
#
# __Bonus:__ replace gaussian noise with masking random rectangles on image.
# +
dataloader_test = DataLoader(X_test_tensor, batch_size=BATCH_SIZE, shuffle=True)
scores = []
for i, (batch) in enumerate(dataloader_test):
batch_noised = apply_gaussian_noise(batch).to(device=device)
scores.append(aenc.batch_loss(batch_noised, batch.cuda(device = device)).data.cpu().numpy())
encoder_out = aenc.enc(batch_noised)
reconstruction_mse = np.mean(scores)
print("Final MSE:", reconstruction_mse)
for i in range(5):
img = apply_gaussian_noise(X_test_tensor[i])
visualize(img,aenc)
# -
# ### Image retrieval with autoencoders
#
# So we've just trained a network that converts image into itself imperfectly. This task is not that useful in and of itself, but it has a number of awesome side-effects. Let's see it in action.
#
# First thing we can do is image retrieval aka image search. We we give it an image and find similar images in latent space.
#
# To speed up retrieval process, we shall use Locality-Sensitive Hashing on top of encoded vectors. We'll use scikit-learn's implementation for simplicity. In practical scenario, you may want to use [specialized libraries](https://erikbern.com/2015/07/04/benchmark-of-approximate-nearest-neighbor-libraries.html) for better performance and customization.
# +
#encodes batch of images into a codes
codes = <encode all images in X_train_tensor>
# -
assert codes.shape[0] == X_train_tensor.shape[0]
from sklearn.neighbors import LSHForest
lshf = LSHForest(n_estimators=50).fit(codes.detach().cpu().numpy())
images = torch.from_numpy(X_train).type(torch.DoubleTensor)
def get_similar(image, n_neighbors=5):
assert len(image.shape)==3,"image must be [batch,height,width,3]"
code = aenc.enc(image.cuda(device)).detach().cpu().numpy()
(distances,),(idx,) = lshf.kneighbors(code,n_neighbors=n_neighbors)
return distances,images[idx]
def show_similar(image):
distances,neighbors = get_similar(image,n_neighbors=11)
plt.figure(figsize=[8,6])
plt.subplot(3,4,1)
plt.imshow(image.cpu().numpy().transpose([1,2,0]))
plt.title("Original image")
for i in range(11):
plt.subplot(3,4,i+2)
plt.imshow(neighbors[i].cpu().numpy().transpose([1,2,0]))
plt.title("Dist=%.3f"%distances[i])
plt.show()
#smiles
show_similar(X_test_tensor[2])
#ethnicity
show_similar(X_test_tensor[500])
#glasses
show_similar(X_test_tensor[66])
# ## Bonus: cheap image morphing
#
image1,image2 = X_test[np.random.randint(0,len(X_test),size=2)]
a = np.stack([image1,image2])
a.shape
# +
for _ in range(5):
image1,image2 = X_test_tensor[np.random.randint(0,len(X_test),size=2)]
code1, code2 = aenc.enc(torch.cat((image1,image2), 0).cuda(device))
plt.figure(figsize=[10,4])
for i,a in enumerate(np.linspace(0,1,num=7)):
output_code = code1*(1-a) + code2*(a)
output_image = aenc.dec(output_code[None])[0]
plt.subplot(1,7,i+1)
plt.imshow(output_image.cpu().detach().numpy())
plt.title("a=%.2f"%a)
plt.show()
# -
# Of course there's a lot more you can do with autoencoders.
#
# If you want to generate images from scratch, however, we recommend you our honor track seminar about generative adversarial networks.
| week08_autoencoders/autoencoders_torch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Copy makes it easy to make duplicates of existing objects. Provides functions for making shallow and deep copies of an object.
# ### Shallow copy - A new container populated with references to the contents of the original object
# ### Deep copy - A new container populated with copies of the contents of the original object
# +
import copy
original_list = [int(x) for x in range(10)]
print(original_list, end = ' ')
# -
# ### Shallow copy
# +
# the reference in shallow_list is to the same object in original_list
shallow_list = copy.copy(original_list)
print(shallow_list, end = ' ')
# -
print('original_list:', original_list)
print('shallow_list:', shallow_list)
print('shallow_list is original_list:', (shallow_list is original_list))
print('shallow_list == original_list:', (shallow_list == original_list))
print('shallow_list[0] is original_list[0]:', (shallow_list[0] is original_list[0]))
print('shallow_list[0] == original_list[0]:', (shallow_list[0] == original_list[0]))
# ### Deepcopy
deep_list = copy.deepcopy(original_list)
print(deep_list, end = ' ')
print('original_list:', original_list)
print('deep_list:', deep_list)
print('deep_list is original_list:', (deep_list is original_list))
print('deep_list == original_list:', (deep_list == original_list))
print('deep_list[0] is original_list[0]:', (deep_list[0] is original_list[0])) # A class instance gives this false!
print('deep_list[0] == original_list[0]:', (deep_list[0] == original_list[0]))
# ### Let's try this with classes!!!
# +
import functools
@functools.total_ordering
class someClass:
def __init__(self, name):
self.name = name
def __eq__(self, other):
return self.name == other.name
def __gt__(self, other):
return self.name > other.name
# +
# Creating shallow copy
ins = someClass('first')
class_list =[ins]
sh_copy = copy.copy(class_list)
dp_copy = copy.deepcopy(class_list)
# -
# ### shallow copy
print('class_list:', class_list)
print('sh_copy:', sh_copy)
print('sh_copy is class_list:', (sh_copy is class_list))
print('sh_copy == class_list:', (sh_copy == class_list))
print('sh_copy[0] is class_list[0]:', (sh_copy[0] is class_list[0]))
print('sh_copy[0] == class_list[0]:', (sh_copy[0] == class_list[0]))
print('class_list:', class_list)
print('dp_copy:', dp_copy)
print('dp_copy is class_list:', (dp_copy is class_list))
print('dp_copy == class_list:', (dp_copy == class_list))
print('dp_copy[0] is class_list[0]:', (dp_copy[0] is class_list[0]))
print('dp_copy[0] == class_list[0]:', (dp_copy[0] == class_list[0]))
| data structures/copy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # In this note book the following steps are taken:
# 1. Remove highly correlated attributes
# 2. Find the best hyper parameters for estimator
# 3. Find the most important features by tunned random forest
# 4. Find f1 score of the tunned full model
# 5. Find best hyper parameter of model with selected features
# 6. Find f1 score of the tuned seleccted model
# 7. Compare the two f1 scores
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn.feature_selection import RFECV,RFE
from sklearn.model_selection import train_test_split, GridSearchCV, KFold,RandomizedSearchCV
from sklearn.preprocessing import StandardScaler
from sklearn.pipeline import Pipeline
from sklearn import metrics
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import accuracy_score,f1_score
import numpy as np
from sklearn.metrics import make_scorer
f1_score = make_scorer(f1_score)
#import data
Data=pd.read_csv("Saskatoon-Transfomed-Data-BS-NoBreak - Copy.csv")
X = Data.iloc[:,:-1]
y = Data.iloc[:,-1]
#split test and training set.
np.random.seed(60)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.2,
random_state = 1000)
#Define estimator and model
classifiers = {}
classifiers.update({"Random Forest": RandomForestClassifier(random_state=1000)})
# +
#Define range of hyperparameters for estimator
np.random.seed(60)
parameters = {}
parameters.update({"Random Forest": { "classifier__n_estimators": [100,105,110,115,120,125,130,135,140,145,150,155,160,170,180,190,200],
# "classifier__n_estimators": [2,4,5,6,7,8,9,10,20,30,40,50,60,70,80,90,100,110,120,130,140,150,160,170,180,190,200],
#"classifier__class_weight": [None, "balanced"],
"classifier__max_features": ["auto", "sqrt", "log2"],
"classifier__max_depth" : [4,6,8,10,11,12,13,14,15,16,17,18,19,20,22],
#"classifier__max_depth" : [1,2,3,4,5,6,7,8,9,10,11,12,13,14,15,16,17,18,19,20],
"classifier__criterion" :["gini", "entropy"]
}})
# +
# Make correlation matrix
corr_matrix = X_train.corr(method = "spearman").abs()
# Draw the heatmap
sns.set(font_scale = 1.0)
f, ax = plt.subplots(figsize=(11, 9))
sns.heatmap(corr_matrix, cmap= "YlGnBu", square=True, ax = ax)
f.tight_layout()
plt.savefig("correlation_matrix.png", dpi = 1080)
# Select upper triangle of matrix
upper = corr_matrix.where(np.triu(np.ones(corr_matrix.shape), k = 1).astype(np.bool))
# Find index of feature columns with correlation greater than 0.8
to_drop = [column for column in upper.columns if any(upper[column] > 0.8)]
# Drop features
X_train = X_train.drop(to_drop, axis = 1)
X_test = X_test.drop(to_drop, axis = 1)
# -
X_train
FEATURE_IMPORTANCE = {"Random Forest"}
selected_classifier = "Random Forest"
classifier = classifiers[selected_classifier]
scaler = StandardScaler()
steps = [("scaler", scaler), ("classifier", classifier)]
pipeline = Pipeline(steps = steps)
#Define parameters that we want to use in gridsearch cv
param_grid = parameters[selected_classifier]
# Initialize GridSearch object for estimator
gscv = RandomizedSearchCV(pipeline, param_grid, cv = 3, n_jobs= -1, verbose = 1, scoring = f1_score, n_iter=30)
# Fit gscv (Tunes estimator)
print(f"Now tuning {selected_classifier}. Go grab a beer or something.")
gscv.fit(X_train, np.ravel(y_train))
#Getting the best hyperparameters
best_params = gscv.best_params_
best_params
#Getting the best score of model
best_score = gscv.best_score_
best_score
# +
#Check overfitting of the estimator
from sklearn.model_selection import cross_val_score
mod = RandomForestClassifier(#class_weight= None,
criterion= 'entropy',
max_depth= 17,
max_features= 'auto',
n_estimators= 115 ,random_state=10000)
scores_test = cross_val_score(mod, X_test, y_test, scoring='f1', cv=5)
scores_test
# -
tuned_params = {item[12:]: best_params[item] for item in best_params}
classifier.set_params(**tuned_params)
#Find f1 score of the model with all features (Model is tuned for all features)
results={}
model=classifier.set_params(criterion= 'entropy',
max_depth= 17,
max_features= 'auto',
n_estimators= 115 ,random_state=10000)
model.fit(X_train,y_train)
y_pred = model.predict(X_test)
F1 = metrics.f1_score(y_test, y_pred)
results = {"classifier": model,
"Best Parameters": best_params,
"Training f1": best_score*100,
"Test f1": F1*100}
results
# Select Features using RFECV
class PipelineRFE(Pipeline):
# Source: https://ramhiser.com/post/2018-03-25-feature-selection-with-scikit-learn-pipeline/
def fit(self, X, y=None, **fit_params):
super(PipelineRFE, self).fit(X, y, **fit_params)
self.feature_importances_ = self.steps[-1][-1].feature_importances_
return self
# +
steps = [("scaler", scaler), ("classifier", classifier)]
pipe = PipelineRFE(steps = steps)
np.random.seed(60)
# Initialize RFECV object
feature_selector = RFECV(pipe, cv = 5, step = 1, verbose = 1)
# Fit RFECV
feature_selector.fit(X_train, np.ravel(y_train))
# Get selected features
feature_names = X_train.columns
selected_features = feature_names[feature_selector.support_].tolist()
# +
performance_curve = {"Number of Features": list(range(1, len(feature_names) + 1)),
"F1": feature_selector.grid_scores_}
performance_curve = pd.DataFrame(performance_curve)
# Performance vs Number of Features
# Set graph style
sns.set(font_scale = 1.75)
sns.set_style({"axes.facecolor": "1.0", "axes.edgecolor": "0.85", "grid.color": "0.85",
"grid.linestyle": "-", 'axes.labelcolor': '0.4', "xtick.color": "0.4",
'ytick.color': '0.4'})
colors = sns.color_palette("RdYlGn", 20)
line_color = colors[3]
marker_colors = colors[-1]
# Plot
f, ax = plt.subplots(figsize=(13, 6.5))
sns.lineplot(x = "Number of Features", y = "F1", data = performance_curve,
color = line_color, lw = 4, ax = ax)
sns.regplot(x = performance_curve["Number of Features"], y = performance_curve["F1"],
color = marker_colors, fit_reg = False, scatter_kws = {"s": 200}, ax = ax)
# Axes limits
plt.xlim(0.5, len(feature_names)+0.5)
plt.ylim(0.60, 1)
# Generate a bolded horizontal line at y = 0
ax.axhline(y = 0.625, color = 'black', linewidth = 1.3, alpha = .7)
# Turn frame off
ax.set_frame_on(False)
# Tight layout
plt.tight_layout()
# -
#Define new training and test set based based on selected features by RFECV
X_train_rfecv = X_train[selected_features]
X_test_rfecv= X_test[selected_features]
np.random.seed(60)
classifier.fit(X_train_rfecv, np.ravel(y_train))
#Finding important features
np.random.seed(60)
feature_importance = pd.DataFrame(selected_features, columns = ["Feature Label"])
feature_importance["Feature Importance"] = classifier.feature_importances_
feature_importance = feature_importance.sort_values(by="Feature Importance", ascending=False)
feature_importance
# Initialize GridSearch object for model with selected features
np.random.seed(60)
gscv = RandomizedSearchCV(pipeline, param_grid, cv = 3, n_jobs= -1, verbose = 1, scoring = f1_score, n_iter=30)
#Tuning random forest classifier with selected features
np.random.seed(60)
gscv.fit(X_train_rfecv,y_train)
#Getting the best parameters of model with selected features
best_params = gscv.best_params_
best_params
#Getting the score of model with selected features
best_score = gscv.best_score_
best_score
# +
#Check overfitting of the tuned model with selected features
from sklearn.model_selection import cross_val_score
mod = RandomForestClassifier(#class_weight= None,
criterion= 'entropy',
max_depth= 18,
max_features= 'sqrt',
n_estimators= 105 ,random_state=10000)
scores_test = cross_val_score(mod, X_test_rfecv, y_test, scoring='f1', cv=5)
scores_test
# -
results={}
model=classifier.set_params(criterion= 'entropy',
max_depth= 18,
max_features= 'sqrt',
n_estimators= 105 ,random_state=10000)
scores_test = cross_val_score(mod, X_test_rfecv, y_test, scoring='f1', cv=5)
model.fit(X_train_rfecv,y_train)
y_pred = model.predict(X_test_rfecv)
F1 = metrics.f1_score(y_test, y_pred)
results = {"classifier": model,
"Best Parameters": best_params,
"Training f1": best_score*100,
"Test f1": F1*100}
results
| RandomForest-RFECV-BreakStatus-Saskatoon-RandomizedSearch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
# # Toma de contacto con Jupyter y SQL
# Esta hoja muestra cómo acceder a bases de datos SQL y también a conectar la salida con Jupyter. Las partes en SQL también se pueden realizar directamente en MySQL ejecutando el programa `mysql` del contenedor.
#
# Igual que en la práctica de introducción, los contenedores se pueden lanzar automáticamente usando `docker-compose` desde el directorio `bdge/sql` una vez bajado el repositorio Git de las prácticas:
#
# $ git clone https://github.com/dsevilla/bdge.git
# $ cd bdge/sql
# $ docker-compose up
#
# Dentro del _Notebook_, la base de datos está disponible en el host con nombre `mysql`.
# Instalación de los paquetes Python necesarios:
# !pip install pymysql ipython-sql
# %load_ext sql
# #%config SqlMagic.feedback = False # Evitar que muestre el número de filas
# +
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib
# %matplotlib inline
matplotlib.style.use('ggplot')
# + language="sql"
# mysql+pymysql://root:root@mysql/?charset=utf8mb4&local_infile=1
# -
# - Format: 7zipped
# - Files:
# - **badges**.xml
# - UserId, e.g.: "420"
# - Name, e.g.: "Teacher"
# - Date, e.g.: "2008-09-15T08:55:03.923"
# - **comments**.xml
# - Id
# - PostId
# - Score
# - Text, e.g.: "@Stu Thompson: Seems possible to me - why not try it?"
# - CreationDate, e.g.:"2008-09-06T08:07:10.730"
# - UserId
# - **posts**.xml
# - Id
# - PostTypeId
# - 1: Question
# - 2: Answer
# - ParentID (only present if PostTypeId is 2)
# - AcceptedAnswerId (only present if PostTypeId is 1)
# - CreationDate
# - Score
# - ViewCount
# - Body
# - OwnerUserId
# - LastEditorUserId
# - LastEditorDisplayName="<NAME>"
# - LastEditDate="2009-03-05T22:28:34.823"
# - LastActivityDate="2009-03-11T12:51:01.480"
# - CommunityOwnedDate="2009-03-11T12:51:01.480"
# - ClosedDate="2009-03-11T12:51:01.480"
# - Title=
# - Tags=
# - AnswerCount
# - CommentCount
# - FavoriteCount
# - **posthistory**.xml
# - Id
# - PostHistoryTypeId
# - 1: Initial Title - The first title a question is asked with.
# - 2: Initial Body - The first raw body text a post is submitted with.
# - 3: Initial Tags - The first tags a question is asked with.
# - 4: Edit Title - A question's title has been changed.
# - 5: Edit Body - A post's body has been changed, the raw text is stored here as markdown.
# - 6: Edit Tags - A question's tags have been changed.
# - 7: Rollback Title - A question's title has reverted to a previous version.
# - 8: Rollback Body - A post's body has reverted to a previous version - the raw text is stored here.
# - 9: Rollback Tags - A question's tags have reverted to a previous version.
# - 10: Post Closed - A post was voted to be closed.
# - 11: Post Reopened - A post was voted to be reopened.
# - 12: Post Deleted - A post was voted to be removed.
# - 13: Post Undeleted - A post was voted to be restored.
# - 14: Post Locked - A post was locked by a moderator.
# - 15: Post Unlocked - A post was unlocked by a moderator.
# - 16: Community Owned - A post has become community owned.
# - 17: Post Migrated - A post was migrated.
# - 18: Question Merged - A question has had another, deleted question merged into itself.
# - 19: Question Protected - A question was protected by a moderator
# - 20: Question Unprotected - A question was unprotected by a moderator
# - 21: Post Disassociated - An admin removes the OwnerUserId from a post.
# - 22: Question Unmerged - A previously merged question has had its answers and votes restored.
# - PostId
# - RevisionGUID: At times more than one type of history record can be recorded by a single action. All of these will be grouped using the same RevisionGUID
# - CreationDate: "2009-03-05T22:28:34.823"
# - UserId
# - UserDisplayName: populated if a user has been removed and no longer referenced by user Id
# - Comment: This field will contain the comment made by the user who edited a post
# - Text: A raw version of the new value for a given revision
# - If PostHistoryTypeId = 10, 11, 12, 13, 14, or 15 this column will contain a JSON encoded string with all users who have voted for the PostHistoryTypeId
# - If PostHistoryTypeId = 17 this column will contain migration details of either "from <url>" or "to <url>"
# - CloseReasonId
# - 1: Exact Duplicate - This question covers exactly the same ground as earlier questions on this topic; its answers may be merged with another identical question.
# - 2: off-topic
# - 3: subjective
# - 4: not a real question
# - 7: too localized
# - **postlinks**.xml
# - Id
# - CreationDate
# - PostId
# - RelatedPostId
# - PostLinkTypeId
# - 1: Linked
# - 3: Duplicate
# - **users**.xml
# - Id
# - Reputation
# - CreationDate
# - DisplayName
# - EmailHash
# - LastAccessDate
# - WebsiteUrl
# - Location
# - Age
# - AboutMe
# - Views
# - UpVotes
# - DownVotes
# - **votes**.xml
# - Id
# - PostId
# - VoteTypeId
# - ` 1`: AcceptedByOriginator
# - ` 2`: UpMod
# - ` 3`: DownMod
# - ` 4`: Offensive
# - ` 5`: Favorite - if VoteTypeId = 5 UserId will be populated
# - ` 6`: Close
# - ` 7`: Reopen
# - ` 8`: BountyStart
# - ` 9`: BountyClose
# - `10`: Deletion
# - `11`: Undeletion
# - `12`: Spam
# - `13`: InformModerator
# - CreationDate
# - UserId (only for VoteTypeId 5)
# - BountyAmount (only for VoteTypeId 9)
# + language="sql"
# DROP SCHEMA IF EXISTS stackoverflow;
# CREATE SCHEMA stackoverflow CHARACTER SET utf8mb4 COLLATE utf8mb4_unicode_ci;
# + language="sql"
# USE stackoverflow;
# -
# Se tiene que habilitar esto para que se permita importar CSVs.
# + language="sql"
# SET GLOBAL local_infile = true;
# + language="sql"
# DROP TABLE IF EXISTS Posts;
# CREATE TABLE Posts (
# Id INT,
# AcceptedAnswerId INT NULL DEFAULT NULL,
# AnswerCount INT DEFAULT 0,
# Body TEXT,
# ClosedDate TIMESTAMP NULL DEFAULT NULL,
# CommentCount INT DEFAULT 0,
# CommunityOwnedDate TIMESTAMP NULL DEFAULT NULL,
# CreationDate TIMESTAMP NULL DEFAULT NULL,
# FavoriteCount INT DEFAULT 0,
# LastActivityDate TIMESTAMP NULL DEFAULT NULL,
# LastEditDate TIMESTAMP NULL DEFAULT NULL,
# LastEditorDisplayName TEXT,
# LastEditorUserId INT NULL DEFAULT NULL,
# OwnerDisplayName TEXT,
# OwnerUserId INT NULL DEFAULT NULL,
# ParentId INT NULL DEFAULT NULL,
# PostTypeId INT, -- 1 = Question, 2 = Answer
# Score INT DEFAULT 0,
# Tags TEXT,
# Title TEXT,
# ViewCount INT DEFAULT 0,
# PRIMARY KEY(Id)
# )
# CHARACTER SET utf8mb4 COLLATE utf8mb4_unicode_ci;
# + language="bash"
# file=../Posts.csv
# test -e $file || wget http://neuromancer.inf.um.es:8080/es.stackoverflow/`basename ${file}`.gz -O - 2>/dev/null | gunzip > $file
# + language="sql"
# LOAD DATA LOCAL INFILE "../Posts.csv" INTO TABLE Posts
# CHARACTER SET utf8mb4
# COLUMNS TERMINATED BY ','
# OPTIONALLY ENCLOSED BY '"'
# ESCAPED BY '"'
# LINES TERMINATED BY '\r\n'
# IGNORE 1 LINES
# (Id,
# @AcceptedAnswerId,
# @AnswerCount,
# Body,
# @ClosedDate,
# @CommentCount,
# @CommunityOwnedDate,
# CreationDate,
# @FavoriteCount,
# @LastActivityDate,
# @LastEditDate,
# LastEditorDisplayName,
# @LastEditorUserId,
# OwnerDisplayName,
# @OwnerUserId,
# @ParentId,
# PostTypeId,
# Score,
# Tags,
# Title,
# @ViewCount)
# SET ParentId = nullif (@ParentId, ''),
# ClosedDate = nullif(@ClosedDate, ''),
# LastEditorUserId = nullif(@OLastEditorUserId, ''),
# LastActivityDate = nullif(@LastActivityDate, ''),
# LastEditDate = nullif(@LastEditDate, ''),
# AcceptedAnswerId = nullif (@AcceptedAnswerId, ''),
# OwnerUserId = nullif(@OwnerUserId, ''),
# LastEditorUserId = nullif(@LastEditorUserId, ''),
# CommunityOwnedDate = nullif(@CommunityOwnedDate, ''),
# FavoriteCount = if(@FavoriteCount = '',0,@FavoriteCount),
# CommentCount = if(@CommentCount = '',0,@CommentCount),
# ViewCount = if(@ViewCount = '',0,@ViewCount),
# AnswerCount = if(@AnswerCount = '',0,@AnswerCount)
# ;
# + language="sql"
# select count(*) from Posts;
# + language="sql"
# select Id,Title from Posts LIMIT 2;
# + language="bash"
# file=../Users.csv
# test -e $file || wget http://neuromancer.inf.um.es:8080/es.stackoverflow/`basename ${file}`.gz -O - 2>/dev/null | gunzip > $file
# + language="sql"
# DROP TABLE IF EXISTS Users;
# CREATE TABLE Users (
# Id INT,
# AboutMe TEXT,
# AccountId INT,
# Age INT NULL DEFAULT NULL,
# CreationDate TIMESTAMP NULL DEFAULT NULL,
# DisplayName TEXT,
# DownVotes INT DEFAULT 0,
# LastAccessDate TIMESTAMP NULL DEFAULT NULL,
# Location TEXT,
# ProfileImageUrl TEXT,
# Reputation INT DEFAULT 0,
# UpVotes INT DEFAULT 0,
# Views INT DEFAULT 0,
# WebsiteUrl TEXT,
# PRIMARY KEY(Id)
# )
# CHARACTER SET utf8mb4 COLLATE utf8mb4_unicode_ci;
# + language="sql"
# LOAD DATA LOCAL INFILE "../Users.csv" INTO TABLE Users
# CHARACTER SET utf8mb4
# COLUMNS TERMINATED BY ','
# OPTIONALLY ENCLOSED BY '"'
# ESCAPED BY '"'
# LINES TERMINATED BY '\r\n'
# IGNORE 1 LINES
# (Id,AboutMe,@AccountId,@Age,@CreationDate,DisplayName,DownVotes,LastAccessDate,Location,ProfileImageUrl,
# Reputation,UpVotes,Views,WebsiteUrl)
# SET LastAccessDate = nullif(@LastAccessDate,''),
# Age = nullif(@Age, ''),
# CreationDate = nullif(@CreationDate,''),
# AccountId = nullif(@AccountId, '')
# ;
# + language="sql"
# select count(*) from Users;
# + language="bash"
# file=../Tags.csv
# test -e $file || wget http://neuromancer.inf.um.es:8080/es.stackoverflow/`basename ${file}`.gz -O - 2>/dev/null | gunzip > $file
# + language="sql"
# DROP TABLE IF EXISTS Tags;
# CREATE TABLE Tags (
# Id INT,
# Count INT DEFAULT 0,
# ExcerptPostId INT NULL DEFAULT NULL,
# TagName TEXT,
# WikiPostId INT NULL DEFAULT NULL,
# PRIMARY KEY(Id)
# )
# CHARACTER SET utf8mb4 COLLATE utf8mb4_unicode_ci;
# + language="sql"
# LOAD DATA LOCAL INFILE "../Tags.csv" INTO TABLE Tags
# CHARACTER SET utf8mb4
# COLUMNS TERMINATED BY ','
# OPTIONALLY ENCLOSED BY '"'
# ESCAPED BY '"'
# LINES TERMINATED BY '\r\n'
# IGNORE 1 LINES
# (Id,Count,@ExcerptPostId,TagName,@WikiPostId)
# SET WikiPostId = nullif(@WikiPostId, ''),
# ExcerptPostId = nullif(@ExcerptPostId, '')
# ;
# + language="bash"
# file=../Comments.csv
# test -e $file || wget http://neuromancer.inf.um.es:8080/es.stackoverflow/`basename ${file}`.gz -O - 2>/dev/null | gunzip > $file
# + language="sql"
# DROP TABLE IF EXISTS Comments;
# CREATE TABLE Comments (
# Id INT,
# CreationDate TIMESTAMP NULL DEFAULT NULL,
# PostId INT NULL DEFAULT NULL,
# Score INT DEFAULT 0,
# Text TEXT,
# UserDisplayName TEXT,
# UserId INT NULL DEFAULT NULL,
# PRIMARY KEY(Id)
# )
# CHARACTER SET utf8mb4 COLLATE utf8mb4_unicode_ci;
# + language="sql"
# LOAD DATA LOCAL INFILE "../Comments.csv" INTO TABLE Comments
# CHARACTER SET utf8mb4
# COLUMNS TERMINATED BY ','
# OPTIONALLY ENCLOSED BY '"'
# ESCAPED BY '"'
# LINES TERMINATED BY '\r\n'
# IGNORE 1 LINES
# (Id,@CreationDate,@PostId,Score,Text,@UserDisplayName,@UserId)
# SET UserId = nullif(@UserId, ''),
# PostId = nullif(@PostId, ''),
# CreationDate = nullif(@CreationDate,''),
# UserDisplayName = nullif(@UserDisplayName,'')
# ;
# + language="sql"
# SELECT Count(*) FROM Comments;
# + language="sql"
# DROP TABLE IF EXISTS Votes;
# CREATE TABLE Votes (
# Id INT,
# BountyAmount INT DEFAULT 0,
# CreationDate TIMESTAMP NULL DEFAULT NULL,
# PostId INT NULL DEFAULT NULL,
# UserId INT NULL DEFAULT NULL,
# VoteTypeId INT,
# PRIMARY KEY(Id)
# )
# CHARACTER SET utf8mb4 COLLATE utf8mb4_unicode_ci;
# + language="bash"
# file=../Votes.csv
# test -e $file || wget http://neuromancer.inf.um.es:8080/es.stackoverflow/`basename ${file}`.gz -O - 2>/dev/null | gunzip > $file
# + language="sql"
# LOAD DATA LOCAL INFILE "../Votes.csv" INTO TABLE Votes
# CHARACTER SET utf8mb4
# COLUMNS TERMINATED BY ','
# OPTIONALLY ENCLOSED BY '"'
# ESCAPED BY '"'
# LINES TERMINATED BY '\r\n'
# IGNORE 1 LINES
# (Id,@BountyAmount,@CreationDate,@PostId,@UserId,VoteTypeId)
# SET UserId = nullif(@UserId, ''),
# PostId = nullif(@PostId, ''),
# BountyAmount = if(@BountyAmount = '',0,@BountyAmount),
# CreationDate = nullif(@CreationDate, '')
# ;
# -
# ## Añadimos las claves ajenas para que todas las tablas estén referenciadas correctamente
#
# Usaremos los comandos `alter table`.
# + language="sql"
#
# ALTER TABLE Posts ADD FOREIGN KEY (ParentId) REFERENCES Posts(Id);
# ALTER TABLE Posts ADD FOREIGN KEY (OwnerUserId) REFERENCES Users(Id);
# ALTER TABLE Posts ADD FOREIGN KEY (LastEditorUserId) REFERENCES Users(Id);
# ALTER TABLE Posts ADD FOREIGN KEY (AcceptedAnswerId) REFERENCES Posts(Id);
# + language="sql"
#
# ALTER TABLE Tags ADD FOREIGN KEY (WikiPostId) REFERENCES Posts(Id);
# ALTER TABLE Tags ADD FOREIGN KEY (ExcerptPostId) REFERENCES Posts(Id);
# + language="sql"
#
# ALTER TABLE Comments ADD FOREIGN KEY (PostId) REFERENCES Posts(Id);
# ALTER TABLE Comments ADD FOREIGN KEY (UserId) REFERENCES Users(Id);
# + language="sql"
#
# ALTER TABLE Votes ADD FOREIGN KEY (PostId) REFERENCES Posts(Id);
# ALTER TABLE Votes ADD FOREIGN KEY (UserId) REFERENCES Users(Id);
# + language="sql"
# EXPLAIN
# SELECT Y.PostId,Y.Present
# FROM (SELECT v.PostId AS PostId, COALESCE(p.Id,CONCAT('No: ', v.PostId)) AS Present
# FROM Votes v LEFT JOIN Posts p ON v.PostId = p.Id) AS Y
# WHERE Y.Present LIKE 'No%';
# + language="sql"
# EXPLAIN SELECT PostId from Votes WHERE PostId NOT IN (select Id from Posts);
# + language="sql"
# select * from Votes LIMIT 20;
# + language="sql"
# SELECT Y.Id, Y.PostId, Y.Present
# FROM (SELECT v.PostId AS PostId, v.Id AS Id, p.Id AS Pid, COALESCE(p.Id, CONCAT('No: ', v.PostId)) AS Present
# FROM Votes v LEFT JOIN Posts p ON v.PostId = p.Id) AS Y
# WHERE Y.Pid IS NULL
# LIMIT 1000
# -
# ## EJERCICIO: Eliminar de `Votes` las entradas que se refieran a Posts inexistentes
# + language="sql"
# -- DELETE FROM Votes WHERE ...;
# + language="sql"
# -- Y ahora sí
# ALTER TABLE Votes ADD FOREIGN KEY (PostId) REFERENCES Posts(Id);
# ALTER TABLE Votes ADD FOREIGN KEY (UserId) REFERENCES Users(Id);
# -
# %sql use stackoverflow
# + language="sql"
# SHOW TABLES;
# + language="sql"
# DESCRIBE Posts;
# -
# top_tags = %sql SELECT Id, TagName, Count FROM Tags ORDER BY Count DESC LIMIT 40;
# ¡¡Los resultados de `%sql` se pueden convertir a un `DataFrame`!!
top_tags_df = top_tags.DataFrame()
# invert_y_axis() hace que el más usado aparezca primero. Por defecto es al revés.
top_tags_df.plot(kind='barh',x='TagName', y='Count', figsize=(14,14*2/3)).invert_yaxis()
top_tags
# + language="sql"
# select Id,TagName,Count from Tags WHERE Count > 5 ORDER BY Count ASC LIMIT 40;
# -
# ### Para comparación con HBase
#
# Voy a hacer unas consultas para comparar la eficiencia con HBase. Calcularé el tamaño medio del texto de los comentarios de un post en particular (he seleccionado el 7251, que es el que más tiene comentarios, 32). Hago el cálculo en local porque aunque existe la función `AVG` de SQL, es posible que la función que tuviéramos que calcular no la tuviera la base de datos, con lo que tenemos que obtener todos los datos y calcularla en local. Eso también nos dará una idea de la eficiencia de recuperación de la base de datos.
# + language="sql"
# SELECT p.Id, MAX(p.CommentCount) AS c FROM Posts p GROUP BY p.Id ORDER BY c DESC LIMIT 1;
# -
# %sql SELECT AVG(CHAR_LENGTH(Text)) from Comments WHERE PostId = 7251;
# +
from functools import reduce
def doit():
# q = %sql select Text from Comments WHERE PostId = 7251;
(s,n) = reduce(lambda res, e: (res[0]+len(e[0]), res[1]+1), q, (0,0))
return (s/n)
# %timeit doit()
# -
# ## EJERCICIO: Calcular las preguntas con más respuestas
#
# En la casilla siguiente:
# + language="sql"
# -- Preguntas con más respuestas (20 primeras)
#
# + language="sql"
# select Title from Posts where Id = 5;
# -
# ### Código de suma de posts de cada Tag
# +
# Calcular la suma de posts cada Tag de manera eficiente
import re
# Obtener los datos iniciales de los Tags
# results = %sql SELECT Id, Tags FROM Posts where Tags IS NOT NULL;
tagcount = {}
for result in results:
# Inserta las tags en la tabla Tag
tags = re.findall('<(.*?)>', result[1])
for tag in tags:
tagcount[tag] = tagcount.get(tag,0) + 1;
# Comprobar que son iguales las cuentas
for k in tagcount:
# res = %sql select TagName,SUM(Count) from Tags WHERE TagName = :k GROUP BY TagName;
if tagcount[k] != res[0][1]:
print("Tag %s NO coincide (%d)!!" % (k, res[0][1]))
# -
tagcount
df = pd.DataFrame({'count' : pd.Series(list(tagcount.values()),
index=list(tagcount.keys()))})
df
sort_df = df.sort_values(by='count',ascending=False)
sort_df
sort_df[:100].plot(kind='bar',figsize=(20,20*2/3))
sort_df[-100:].plot(kind='bar',figsize=(20,20*2/3))
# ## EJERCICIO: Crear una tabla "PostTags" que relaciona cada Tag con su Post
#
# Tendrá cuatro entradas, Id, PostId (referencia a Posts.Id), TagId (referencia a Tags.Id) y TagName (nombre del tag copiado de Tags)
# ## EJERCICIO: ¿Cómo se podría encontrar lo más rápido posible todos los Posts de un Tag en particular (dando el TagName)?
#
# Se pueden dar varias alternativas comparando la eficiencia de cada una. Se pueden hacer pruebas de eficiencia.
| sql/sesion1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Integrating eccentric Comets with MERCURIUS
# In this example, we study highly eccentric comets which interact with a Neptune mass planet.
#
# MERCURIUS ia a hybrid integration scheme which combines the WHFAST and IAS15 algorithms. It smoothly transitions between the two integrators, similar to what the hybrid integrator in the MERCURY package is doing.
import rebound
import numpy as np
# First let's choose the basic properties required for the MERCURIUS integrator to run correctly. In particular, we are:
# * adding comets as *semi-active* bodies, which means they can influence/be influenced by other active bodies, but are invisible to each other. This is done by setting `testparticle_type = 1`. Setting `testparticle_type = 0` would indicate that we are adding comets as *test* bodies.
# * merging bodies when a collision is triggered, conserving momentum and mass.
# * removing particles that leave our pre-defined box.
# * tracking the energy lost due to ejections or collisions.
# +
sim = rebound.Simulation()
np.random.seed(42)
#integrator options
sim.integrator = "mercurius"
sim.dt = 1
sim.testparticle_type = 1
#collision and boundary options
sim.collision = "direct"
sim.collision_resolve = "merge"
sim.collision_resolve_keep_sorted = 1
sim.boundary = "open"
boxsize = 200.
sim.configure_box(boxsize)
sim.track_energy_offset = 1
#simulation time
tmax = 1e4
# -
# Now that the preliminary setup is complete, it's time to add some particles to the system! When using the MERCURIUS integrator it is important to add active bodies first and semi-active bodies later. The `sim.N_active` variable distinguishes massive bodies from semi-active/test bodies.
# +
#massive bodies
sim.add(m=1., r=0.005) # Sun
a_neptune = 30.05
sim.add(m=5e-5,r=2e-4,a=a_neptune,e=0.01) # Neptune
sim.N_active = sim.N
# -
# Now, let's create some comets! For this simple example we are assuming that all comets have the same mass and radius.
# +
# semi-active bodies
n_comets = 100
a = np.random.random(n_comets)*10 + a_neptune
e = np.random.random(n_comets)*0.009 + 0.99
inc = np.random.random(n_comets)*np.pi/2.
m = 1e-10
r = 1e-7
for i in xrange(0,n_comets):
rand = np.random.random()*2*np.pi
sim.add(m=m, r=r, a=a[i], e=e[i], inc=inc[i], Omega=0, omega=rand, f=rand)
# -
# We need to move to the COM frame to avoid drifting out of our simulation box. Also it is always good practice to monitor the change in energy over the course of a simulation, which requires us to calculate it before and after the simulation.
sim.move_to_com()
E0 = sim.calculate_energy()
# We can visualize our setup using `rebound.OrbitPlot`
# %matplotlib inline
fig = rebound.OrbitPlot(sim,Narc=300)
# Alternatively, we can also use the WebGL Widget to get an interactive visualization of the simulation.
sim.getWidget(size=(500,300),scale=1.8*a_neptune)
# Finally, let's simulate our system for and check that our final relative energy error is small. The energy error is a key measure of whether the integration was performed accurately or not.
sim.integrate(tmax)
dE = abs((sim.calculate_energy() - E0)/E0)
print(dE)
| rebound/rebound_official_examples/EccentricComets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
# -
#export
from exp.nb_07a import *
# # Image ItemList
# ## Get images
URLs.IMAGENETTE_160
path = untar_data(URLs.IMAGENETTE_160)
path
#export
import PIL, os, mimetypes
Path.ls = lambda x: list(x.iterdir())
path.ls()
(path/'val').ls()
path_tench = (path/'val'/'n01440764')
img_fn = path_tench.ls()[0]
img_fn
img = PIL.Image.open(img_fn)
img
plt.imshow(img);
import numpy
imga = numpy.array(img)
imga.shape
imga[:5,:5,0]
# +
# mimetypes.types_map.items()
# -
#export
image_extensions = set(k for k,v in mimetypes.types_map.items() if v.startswith('image/'))
' '.join(image_extensions)
#export
def setify(o): return o if isinstance(o, set) else set(listify(o))
test_eq(setify('aa'), {'aa'})
test_eq(setify(['aa',1]), {'aa',1})
test_eq(setify(None), set())
test_eq(setify(1), {1})
test_eq(setify({1}), {1})
#export
def _get_files(path, files, extensions=None):
p = Path(path)
res = [p/f for f in files if not f.startswith('.') and ((not extensions) or f'.{f.split(".")[-1].lower()}' in extensions)]
return res
t = [o.name for o in os.scandir(path_tench)]
t[:3]
type(t[0])
t_path = _get_files(path_tench, t, image_extensions)
t_path[:3]
next(os.scandir(path_tench)).name
#export
def get_files(path, extensions=None, recurse=False, include=None):
path = Path(path)
extensions = setify(extensions)
extensions = {e.lower() for e in extensions}
if recurse:
res = []
for i, (p,d,f) in enumerate(os.walk(path)): # returns (dirpath, dirnames, filesnames)
if include is not None and i == 0: #
d[:] = [o for o in d if o in include] #
else: #
d[:] = [o for o in d if not o.startswith('.')] # the if else statement changes which directories are looked into; now only the hidden directories are skipped
res += _get_files(p, f, extensions)
return res
else:
f = [o.name for o in os.scandir(path) if o.is_file()]
return _get_files(path, f, extensions)
get_files(path_tench, image_extensions)[:3]
get_files(path, image_extensions, recurse=True)[:3]
get_files(path, image_extensions, recurse=True)[:3]
all_fns = get_files(path, image_extensions, recurse=True)
len(all_fns)
# %timeit -n 10 get_files(path, image_extensions, recurse=True)
# # Prepare for modeling
# ## Get files
#export
def compose(x, funcs, *args, order_key='_order', **kwargs):
key = lambda o: getattr(o, order_key, 0)
for f in sorted(listify(funcs), key=key): x = f(x, **kwargs)
return x
# +
# ListContainer??
# +
#export
class ItemList(ListContainer):
def __init__(self, items, path='.', tfms=None):
super().__init__(items)
self.path, self.tfms = Path(path), tfms
def __repr__(self): return f'{super().__repr__()}\nPath: {self.path}'
def new(self, items, cls=None):
if cls is None: cls = self.__class__
return cls(items, self.path, self.tfms)
def get(self, i): return i
def _get(self, i): return compose(self.get(i), self.tfms)
def __getitem__(self, idx):
res = super().__getitem__(idx)
if isinstance(res, list): return [self._get(o) for o in res]
return self._get(res)
class ImageList(ItemList):
@classmethod
def from_files(cls, path, extensions=None, recurse=True, include=None, **kwargs):
if extensions is None: extensions = image_extensions
return cls(get_files(path, extensions, recurse, include), path, **kwargs)
def get(self, fn): return PIL.Image.open(fn)
# +
#export
class Transform(): _order = 0
class MakeRGB(Transform):
def __call__(self, item): return item.convert('RGB')
def make_rgb(item): return item.convert('RGB')
# -
il = ImageList.from_files(path, tfms=make_rgb)
il
img = il[0]; img
il[:1]
# ## Split Validation set
fn = il.items[0]; fn
fn.parent.parent.name
#export
def grandparent_splitter(fn, train_name='train', valid_name='valid'):
gp = fn.parent.parent.name
return True if gp == valid_name else False if gp == train_name else None
#export
def split_by_func(items, f):
mask = [f(o) for o in items]
# 'None' values will be filtered out
t = [o for o,m in zip(items, mask) if m == False] # items in training folder
v = [o for o,m in zip(items, mask) if m == True] # items in val folder
return t,v
splitter = partial(grandparent_splitter, valid_name='val')
# %time train, valid = split_by_func(il, splitter)
len(train), len(valid)
#export
class SplitData():
def __init__(self, train, valid): self.train, self.valid = train, valid
def __getattr__(self, k): return getattr(self.train, k)
# This is needed if we want to picle SplitData and be able to load it back without recursion erros
def __setstate__(self, data:Any): self.__dict__.update(data)
@classmethod
def split_by_func(cls, il, f):
# lists = map(il.new, split_by_func(il.items, f))
lists = map(il.new, split_by_func(il, f))
return cls(*lists)
def __repr__(self): return f'{self.__class__.__name__}\nTrain: {self.train}\nValid: {self.valid}\n'
sd = SplitData.split_by_func(il, splitter); sd
sd.train
# ## Labeling
# +
#export
from collections import OrderedDict
def uniqueify(x, sort=False):
res = list(OrderedDict.fromkeys(x).keys())
if sort: res.sort()
return res
# +
#export
class Processor():
def process(self, items): return items
class CategoryProcessor(Processor):
def __init__(self): self.vocab = None
def __call__(self, items):
# The vocab is defined on the first use
if self.vocab is None:
self.vocab = uniqueify(items)
self.otoi = {v:k for k,v in enumerate(self.vocab)}
return [self.proc1(o) for o in items]
def proc1(self, item): return self.otoi[item]
def deprocess(self, idxs):
assert self.vocab is not None
return [self.deproc1(idx) for idx in idxs]
def deproc1(self, idx): return self.vocab[idx]
# +
#export
def parent_labeler(fn): return fn.parent.name
def _label_by_func(il, f, cls=ItemList): return cls([f(o) for o in il.items], path=il.path)
# -
#export
class LabeledData():
def process(self, il, proc): return il.new(compose(il, proc)) # maybe il.items needed?
def __init__(self, x, y, proc_x=None, proc_y=None):
self.x, self.y = self.process(x, proc_x), self.process(y, proc_y)
self.proc_x, self.proc_y = proc_x, proc_y
def __repr__(self): return f'{self.__class__.__name__}\nx: {self.x}\ny: {self.y}\n'
def __getitem__(self, idx): return self.x[idx], self.y[idx]
def __len__(self): return len(self.x)
def x_obj(self, idx): return self.obj(self.x, idx, self.proc_x)
def y_obj(self, idx): return self.obj(self.y, idx, self.proc_y)
def obj(self, items, idx, procs):
isint = isinstance(idx, int) or (isinstance(idx, torch.LongTensor) and not idx.ndim)
item = items[idx]
for proc in reversed(listify(procs)):
item = proc.deproc1(item) if isint else proc.deprocess(item)
return item
@classmethod
def label_by_func(cls, il, f, proc_x=None, proc_y=None):
return cls(il, _label_by_func(il, f), proc_x=proc_x, proc_y=proc_y)
#export
def label_by_func(sd, f, proc_x=None, proc_y=None):
train = LabeledData.label_by_func(sd.train, f, proc_x=proc_x, proc_y=proc_y)
valid = LabeledData.label_by_func(sd.valid, f, proc_x=proc_x, proc_y=proc_y)
return SplitData(train, valid)
ll = label_by_func(sd, parent_labeler, proc_y=CategoryProcessor())
assert ll.train.proc_y is ll.valid.proc_y
ll.train.y
ll.train.y.items[0]
ll.train.y_obj(0)
ll.train.y_obj(slice(2))
ll
# ## Transform to tensor
ll.train[0]
ll.train[0][0]
ll.train[0][0].resize((128,128))
# +
#export
class ResizeFixed(Transform):
_order = 10
def __init__(self, size):
if isinstance(size, int): size = (size,size)
self.size = size
def __call__(self, item): return item.resize(self.size, PIL.Image.BILINEAR)
def to_byte_tensor(item):
res = torch.ByteTensor(torch.ByteStorage.from_buffer(item.tobytes()))
w, h = item.size
return res.view(h,w,-1).permute(2,0,1)
# return res.view(h, w, -1)
to_byte_tensor._order = 20
def to_float_tensor(item): return item.float().div_(255.)
to_float_tensor._order = 30
# +
tfms = [make_rgb, ResizeFixed(128), to_byte_tensor, to_float_tensor]
il = ImageList.from_files(path, tfms=tfms)
sd = SplitData.split_by_func(il, splitter)
ll = label_by_func(sd, parent_labeler, proc_y=CategoryProcessor())
# -
#export
def show_image(im, figsize=(3,3)):
plt.figure(figsize=figsize)
plt.axis('off')
plt.imshow(im.permute(1,2,0))
# plt.imshow(im)
x, y = ll.train[0]
x.shape
show_image(x)
# # Modeling
# ## DataBunch
bs = 64
train_dl, valid_dl = get_dls(ll.train, ll.valid, bs, num_workers=4)
x, y = next(iter(train_dl))
x.shape
show_image(x[0])
ll.train.proc_y.vocab[y[0]]
y
#export
class DataBunch():
def __init__(self, train_dl, valid_dl, c_in=None, c_out=None):
self.train_dl, self.valid_dl, self.c_in, self.c_out = train_dl, valid_dl, c_in, c_out
@property
def train_ds(self): return self.train_dl.dataset
@property
def valid_ds(self): return self.valid_dl.dataset
# +
#export
def databunchify(sd, bs, c_in=None, c_out=None, **kwargs):
dls = get_dls(sd.train, sd.valid, bs, **kwargs)
return DataBunch(*dls, c_in=c_in, c_out=c_out)
SplitData.to_databunch = databunchify
# +
path = untar_data(URLs.IMAGENETTE_160)
tfms = [make_rgb, ResizeFixed(128), to_byte_tensor, to_float_tensor]
il = ImageList.from_files(path, tfms=tfms)
sd = SplitData.split_by_func(il, partial(grandparent_splitter, valid_name='val'))
ll = label_by_func(sd, parent_labeler, proc_y=CategoryProcessor())
data = ll.to_databunch(bs, c_in=3, c_out=10, num_workers=4)
# -
# ## Model
cbfs = [partial(AvgStatsCallback, accuracy),
CudaCallback]
m, s = x.mean((0,2,3)).cuda(), x.std((0,2,3)).cuda()
m, s
# +
#export
def normalize_chan(x, mean, std):
return (x - mean[...,None,None] / std[...,None,None])
_m = tensor([0.47, 0.48, 0.45])
_s = tensor([0.29, 0.28, 0.30])
norm_imagenette = partial(normalize_chan, mean=_m.cuda(), std=_s.cuda())
# -
cbfs.append(partial(BatchTransformXCallback, norm_imagenette))
nfs = [64, 64, 128, 256]
#export
import math
def prev_pow_2(x): return 2**math.floor(math.log2(x))
#export
def get_learn_run(nfs, data, lr, layer, cbs=None, opt_func=None, **kwargs):
model = get_cnn_model(data, nfs, layer, **kwargs)
init_cnn(model)
return get_runner(model, data, lr=lr, cbs=cbs, opt_func=opt_func)
#export
def get_cnn_model(data, nfs, layer, **kwargs):
return nn.Sequential(*get_cnn_layers(data, nfs, layer, **kwargs))
#export
def get_cnn_layers(data, nfs, layer, **kwargs):
def f(ni, nf, stride=2): return layer(ni, nf, 3, stride=stride, **kwargs)
l1 = data.c_in
l2 = prev_pow_2(l1*3*3) #prev_pow_2(27) = 2^4 = 16
layers = [f(l1, l2, stride=1),
f(l2, l2*2, stride=2),
f(l2*2, l2*4, stride=2)]
nfs = [l2*4] + nfs
layers += [f(nfs[i], nfs[i+1]) for i in range(len(nfs)-1)]
layers += [nn.AdaptiveAvgPool2d(1), Lambda(flatten), nn.Linear(nfs[-1], data.c_out)]
return layers
sched = combine_scheds([0.3, 0.7], cos_1cycle_anneal(0.1, 0.3, 0.05))
learn, run = get_learn_run(nfs, data, 0.2, conv_layer, cbs=cbfs + [partial(ParamScheduler, 'lr', sched)])
#export
def model_summary(run, learn, data, find_all=False):
xb, yb = get_batch(data.valid_dl, run)
device = next(learn.model.parameters()).device
xb, yb = xb.to(device), yb.to(device)
mods = find_modules(learn.model, is_lin_layer) if find_all else learn.model.children()
f = lambda hook,mod,inp,out: print(f"{mod}\n{out.shape}\n")
with Hooks(mods, f) as hooks: learn.model(xb)
model_summary(run, learn, data)
# %time run.fit(5, learn)
# !python notebook2script.py 08_data_block.ipynb
| nbs/dl2/selfmade/08_data_block.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ## Homework 5: Fitting (cont)
# Please complete this homework assignment in code cells in the iPython notebook. Include comments in your code when necessary. Please rename the notebook as SIS ID_HW05.ipynb (your student ID number) and save the notebook once you have executed it as a PDF (note, that when saving as PDF you don't want to use the option with latex because it crashes, but rather the one to save it directly as a PDF).
#
# **The homework should be submitted on bCourses under the Assignments tab (both the .ipynb and .pdf files). Please label it by your student ID number (SIS ID)**
# ## Problem 1: Optical Pumping experiment
#
# One of the experiments in the 111B (111-ADV) lab is the study of the optical pumping of atomic rubidium. In that experiment, we measure the resonant frequency of a Zeeman transition as a function of the applied current (local magnetic field). Consider a mock data set:
# <table border="1" align="center">
#
# <tr>
# <td>Current <i>I</i> (Amps)
# </td><td>0.0 </td><td> 0.2 </td><td> 0.4 </td><td> 0.6 </td><td> 0.8 </td><td> 1.0 </td><td> 1.2 </td><td> 1.4 </td><td> 1.6 </td><td> 1.8 </td><td> 2.0 </td><td> 2.2
# </td></tr>
# <tr>
# <td>Frequency <i>f</i> (MHz)
# </td><td> 0.14 </td><td> 0.60 </td><td> 1.21 </td><td> 1.74 </td><td> 2.47 </td><td> 3.07 </td><td> 3.83 </td><td> 4.16 </td><td> 4.68 </td><td> 5.60 </td><td> 6.31 </td><td> 6.78
# </td></tr></table>
#
# 1. Plot a graph of the pairs of values. Assuming a linear relationship between $I$ and $f$, determine the slope and the intercept of the best-fit line using the least-squares method with equal weights, and draw the best-fit line through the data points in the graph.
# 1. From what s/he knows about the equipment used to measure the resonant frequency, your lab partner hastily estimates the uncertainty in the measurement of $f$ to be $\sigma(f) = 0.01$ MHz. Estimate the probability that the straight line you found is an adequate description of the observed data if it is distributed with the uncertainty guessed by your lab partner. (Hint: use scipy.stats.chi2 class to compute the quantile of the chi2 distribution). What can you conclude from these results?
# 1. Repeat the analysis assuming your partner estimated the uncertainty to be $\sigma(f) = 1$ MHz. What can you conclude from these results?
# 1. Assume that the best-fit line found in Part 1 is a good fit to the data. Estimate the uncertainty in measurement of $y$ from the scatter of the observed data about this line. Again, assume that all the data points have equal weight. Use this to estimate the uncertainty in both the slope and the intercept of the best-fit line. This is the technique you will use in the Optical Pumping lab to determine the uncertainties in the fit parameters.
# 1. Now assume that the uncertainty in each value of $f$ grows with $f$: $\sigma(f) = 0.03 + 0.03 * f$ (MHz). Determine the slope and the intercept of the best-fit line using the least-squares method with unequal weights (weighted least-squares fit)
#
# +
import numpy as np
import matplotlib.pyplot as plt
from numpy.linalg import *
import scipy.stats
import scipy.optimize as fitter
# Use current as the x-variable in your plots/fitting
current = np.arange(0, 2.3, .2) # Amps
frequency = np.array([.14, .6, 1.21, 1.74, 2.47, 3.07, 3.83, 4.16, 4.68, 5.6, 6.31, 6.78]) # MHz
# +
def linear_model(x, slope, intercept):
'''Model function to use with curve_fit();
it should take the form of a line'''
# Use fitter.curve_fit() to get the line of best fit
# Plot this line, along with the data points -- remember to label
# -
# The rest is pretty short, but the statistics might be a bit complicated. Ask questions if you need advice or help. Next, the problem is basically asking you to compute the $\chi^2$ for the above fit twice, once with $0.01$ as the error for each point (in the 'denominator' of the $\chi^2$ formula) and once with $1$.
#
# These values can then be compared to a "range of acceptable $\chi^2$ values", found with `scipy.stats.chi2.ppf()` -- which takes two inputs. The second input should be the number of degrees of freedom used during fitting (# data points minus the 2 free parameters). The first input should be something like $0.05$ and $0.95$ (one function call of `scipy.stats.chi2.ppf()` for each endpoint fo the acceptable range). If the calculated $\chi^2$ statistic falls within this range, then the assumed uncertainty is reasonable.
# Now, estimate the uncertainty in the frequency measurements, and use this to find the uncertainty in the best-fit parameters. [This document](https://pages.mtu.edu/~fmorriso/cm3215/UncertaintySlopeInterceptOfLeastSquaresFit.pdf) is a good resource for learning to propagate errors in the context of linear fitting.
#
# Finally, repeat the fitting with the weighted errors (from the $\sigma(f)$ uncertainty formula) given to `scipy.optimize.curve_fit()`
| Week09/HW05/Homework05.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="CuRCRaMYIB4g"
# ## Deep Learning for Computer Vision
#
# ### Logo Detection
# ### Bocconi University
#
#
#
# + [markdown] id="3R0gkI3Wa9Qi"
# ### 1. Configure environment / download Object Detection API github
# + id="fUEUJ46NmUC8" colab={"base_uri": "https://localhost:8080/"} outputId="071ed515-8b1f-44ce-f4e7-f8ca1e7d596b"
import os
import pathlib
# Clone the tensorflow models repository
if "models" in pathlib.Path.cwd().parts:
while "models" in pathlib.Path.cwd().parts:
os.chdir('..')
elif not pathlib.Path('models').exists():
# !git clone --depth 1 https://github.com/tensorflow/models
# + colab={"base_uri": "https://localhost:8080/"} id="44k5-8JC4PG_" outputId="5ae85c5f-7885-4a0c-cf29-a6a9aeb95a07"
# %%shell
python -m pip install --upgrade pip
sudo apt install -y protobuf-compiler
# cd models/research/
protoc object_detection/protos/*.proto --python_out=.
# cp object_detection/packages/tf2/setup.py .
python -m pip install .
# + [markdown] id="MmniDjcbI90b"
# #2. Importing libraries and define functions
# + id="qIi0IGDvmg_F"
import io
import os
import scipy.misc
import numpy as np
import six
import time
import pathlib
import pandas as pd
from os import listdir
from os.path import isfile, join
import cv2
from six import BytesIO
import matplotlib
import matplotlib.pyplot as plt
from PIL import Image, ImageDraw, ImageFont
import tensorflow as tf
from object_detection.utils import visualization_utils as viz_utils
from PIL import Image
# %matplotlib inline
import warnings
warnings.filterwarnings('ignore')
# + id="-y9R0Xllefec"
# Function img to np
def load_image_into_numpy_array(path):
img_data = tf.io.gfile.GFile(path, 'rb').read()
image = Image.open(BytesIO(img_data))
(im_width, im_height) = image.size
return np.array(image.getdata()).reshape(
(im_height, im_width, 3)).astype(np.uint8)
#Function to compute IOU
#From https://www.pyimagesearch.com/2016/11/07/intersection-over-union-iou-for-object-detection/
def bb_intersection_over_union(boxA, boxB):
# determine the (x, y) coordinates of the intersection rectangle
xA = max(boxA[0], boxB[0])
yA = max(boxA[1], boxB[1])
xB = min(boxA[2], boxB[2])
yB = min(boxA[3], boxB[3])
# compute the area of intersection rectangle
interArea = max(0, xB - xA + 1) * max(0, yB - yA + 1)
# compute the area of both the prediction and ground-truth rectangles
boxAArea = (boxA[2] - boxA[0] + 1) * (boxA[3] - boxA[1] + 1)
boxBArea = (boxB[2] - boxB[0] + 1) * (boxB[3] - boxB[1] + 1)
iou = interArea / float(boxAArea + boxBArea - interArea)
return iou
# Func int to text class for 14 logos:
def class_text_to_int(row_label):
if row_label == 'Nike':
return 1
elif row_label == 'Adidas':
return 2
elif row_label == 'Starbucks':
return 3
elif row_label == 'Apple Inc.':
return 4
elif row_label == 'NFL':
return 5
elif row_label == 'Mercedes-Benz':
return 6
elif row_label == 'Under Armour':
return 7
elif row_label == 'Coca-Cola':
return 8
elif row_label == 'Hard Rock Cafe':
return 9
elif row_label == 'Puma':
return 10
elif row_label == 'The North Face':
return 11
elif row_label == 'Toyota':
return 12
elif row_label == 'Chanel':
return 13
elif row_label == 'Pepsi':
return 14
else:
None
# + [markdown] id="PrYxHU87Jldu"
# #3. Load a pretrained model and evaluate it on the test data
# + colab={"base_uri": "https://localhost:8080/"} id="evN8bXZz6lhA" outputId="11e95bca-a0a1-4a64-a403-35f0a3c88c8d"
# Mount the drive
# Make sure to creat a shorcut of DLCV_group folder to Google Drive
from google.colab import drive
drive.mount('/content/drive')
# + colab={"base_uri": "https://localhost:8080/"} id="i_TEiTqpmhD6" outputId="3492472f-0cad-4f90-addd-25fd7088e3a5"
# Load a saved model
tf.keras.backend.clear_session()
detect_fn = tf.saved_model.load('/content/drive/MyDrive/DLCV_group/model_centernet_14logos/saved_model')
# + colab={"base_uri": "https://localhost:8080/"} id="WnxTjSZJmhGe" outputId="c8d36e9d-0397-4445-d9f6-dbc8804b7198"
image_dir = '/content/drive/MyDrive/DLCV_group/data/test'
image_path_list = [f for f in listdir(image_dir) if isfile(join(image_dir, f))]
prediction = pd.DataFrame(index= range(len(image_path_list)),columns=['filename', 'yminB', 'xminB', 'ymaxB','xmaxB','classB','score'])
#it takes up to 10h in colab, we run it on the virtual machine
for i in range(len(image_path_list)):
image_path = os.path.join(image_dir, image_path_list[i])
image_np = load_image_into_numpy_array(image_path)
input_tensor = np.expand_dims(image_np, 0)
detections = detect_fn(input_tensor)
prediction['filename'][i]=image_path_list[i]
prediction['yminB'][i]=detections['detection_boxes'][0][0].numpy()[0]
prediction['xminB'][i]=detections['detection_boxes'][0][0].numpy()[1]
prediction['ymaxB'][i]=detections['detection_boxes'][0][0].numpy()[2]
prediction['xmaxB'][i]=detections['detection_boxes'][0][0].numpy()[3]
prediction['classB'][i]=detections['detection_classes'][0][0].numpy().astype(np.int32)
prediction['score'][i]=detections['detection_scores'][0][0]
if i % 100 == 0:
print('{} Pictures processed'.format(i))
prediction.to_csv('temp_result.csv', index=False)
# + [markdown] id="c1tck1XrmbHQ"
# We noticed that some image's sizes were wrong, therefore we rewrite the height and width of annotations. Then apply our model on the test data and calculate the IOU.
#
# + id="G4S1xvyZmhJ6"
# Upload temp_result.csv
prediction = pd.read_csv('temp_result.csv')
prediction['score'] = prediction['score'].apply(lambda x: float(x[10:12]))
prediction['classB'] = prediction['classB'].apply(class_int_to_text)
prediction = prediction.loc[:,['filename','xminB','yminB','xmaxB','ymaxB','classB','score']]
# Upload test set and normalizing the box dimension
test_set = pd.read_csv('/content/drive/MyDrive/DLCV_group/data/test.csv')
test_set = test_set.rename(columns={'photo_filename':'filename'})
# Get width and height of every image
image_dir = '/content/drive/MyDrive/DLCV_groupdata/test'
for i in range(len(test_set)):
image_path = os.path.join(image_dir, test_set['filename'][i])
im = Image.open(image_path)
w, h = im.size
test_set['width'][i]=w
test_set['height'][i]=h
# Merging
test_prediction = pd.merge(test_set,prediction,on='filename')
# Get the full box prediction
test_prediction['xminB'] = test_prediction['xminB']*test_prediction['width']
test_prediction['xmaxB'] = test_prediction['xmaxB']*test_prediction['width']
test_prediction['yminB'] = test_prediction['yminB']*test_prediction['height']
test_prediction['ymaxB'] = test_prediction['ymaxB']*test_prediction['height']
# Calculating IOU
test_prediction['IOU'] = np.nan
for i in range(len(test_prediction)):
boxA = [test_prediction['xmin'][i],test_prediction['ymin'][i],test_prediction['xmax'][i],test_prediction['ymax'][i]]
boxB = [test_prediction['xminB'][i],test_prediction['yminB'][i],test_prediction['xmaxB'][i],test_prediction['ymaxB'][i]]
test_prediction['IOU'][i] = bb_intersection_over_union(boxA, boxB)
test_prediction.to_csv('final_result.csv',index=False)
# + [markdown] id="bsR3ktpWmuYa"
# #4. Results
# Average perfomances of our model for each brand
# + id="P6eQGXwVmhMH" colab={"base_uri": "https://localhost:8080/", "height": 488} outputId="6d465041-63c7-49af-b42a-8a17740ffd19"
# Result mean IOU by class
test_prediction= pd.read_csv('/content/drive/MyDrive/DLCV_group/final_result.csv')
result_iou = test_prediction.groupby('class').mean()[['IOU']].reset_index().sort_values(by=['IOU'],ascending=False).reset_index(drop=True)
result_iou['IOU'] = np.around(result_iou['IOU'], decimals=3)
result_iou.to_csv('final_results_agg_class.csv',index=False)
result_iou
| colab_tutorials/3_Results.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import getpass
import time
import mechanicalsoup
import random
from bs4 import BeautifulSoup
import unicodedata
def credentials():
''' Prompt user for login credentials. '''
USERNAME = input("nbgallery ID: ")
PASSWORD = getpass.getpass("nbgallery pwd:")
return USERNAME, PASSWORD
def getSession(USERNAME,PASSWORD):
''' Login to nbgallery using provided credentials. '''
LOGIN_URL='http://localhost:3000/'
FORM_ID='#sign_in_user'
browser = mechanicalsoup.StatefulBrowser( )
browser.open(LOGIN_URL)
browser.select_form(FORM_ID) #in form: #loginForm
#The following are the form name attributes
browser["user[email]"] = USERNAME
browser["user[password]"] = PASSWORD
resp = browser.submit_selected()
return browser
def _get_page(s, url):
''' Retrieve a page given its URL. '''
#Play nice
time.sleep(random.uniform(0.1, 0.3))
r=s.open(url)
return r
def _search_by_code(s,code, structured=False):
''' Search for VLE sites associated with a particular module code. '''
surl='https://learn2.open.ac.uk/course/search.php?search={}'.format(code)
search = _get_page(s, surl)
soup = BeautifulSoup(search.content, "lxml")
links = [h3.find_next('a') for h3 in soup.find_all("h3",{'class':'coursename'})]
module_links=[]
for link in links:
if link.get('href'):
#https://stackoverflow.com/a/22497114/454773
for link_txt in link.findAll('span'):
link_txt.unwrap()
module_links.append( (link.text,link.get('href')) )
if not structured: return module_links
return [{'title':t,'url':u,
'codepres':t.split()[0],
'code':t.split()[0].split('-')[0]} for t,u in module_links]
def _search_focus(s):
''' Prompter for searching for a VLE module by module code or module-presentation code. '''
code = input("Module code (e.g. TM351 or TM129-17J): ")
results = _search_by_code(s,code)
if not len(results):
print('Nothing found for "{}"'.format(code))
elif len(results)>1:
print("Please be more specific:\n\t{}\n".format('\n\t'.join([r[0].split(' ')[0] for r in results])))
else: return results[0]
return None
# -
u,p=credentials()
br = getSession(u,p)
# +
#br = ms.StatefulBrowser(raise_on_404=True)
#br.open('http://www.csm-testcenter.org/test?do=show&subdo=common&test=file_upload')
#form = br.select_form()
#form['file_upload'] = '/path/to/some/file'
#br.submit_selected()
# -
url='http://localhost:3000/'
br.open(url)
form = br.select_form('#uploadFileForm')
form.set_checkbox({'agree':True})
form['file'] = '/Users/tonyhirst/notebooks/Untitled.ipynb'
r = br.submit_selected()
form=br.select_form('#stageForm')
form.set("title", "dummy notebook")
form.set("description", "dummy desc")
form.set("tags", "dummytag1,dummytag2")
form.set_checkbox({'agree':True})
br.submit_selected()
# +
#br.form.add_file(open(filename), 'text/plain', filename)
# +
#br.form.add_file(open(filename), 'text/plain', filename)
# +
wp=s.get_current_page()
#Click upload button
#uploadModalButton
# -
wp.find("form", { "id":"uploadFileForm"})#.find('table')
wp.select_form('#uploadFileForm') #in form: #loginForm
#The following are the form name attributes
#s["user[email]"] = USERNAME
wp.find_control(type="checkbox")#.selected =True
#r
#resp = browser.submit_selected()
# +
#br.form.add_file(open(filename), 'text/plain', filename)
resp = browser.submit_selected()
# -
# Multistage form:
#
# - stages
# - preprocess
# - tags
# - notebook
#
#
# ## PART 1
#
# <center><h4 class="modal-title">Upload Notebook - Part 1 of 2</h4></center></div><div class="modal-body"><form id="uploadFileForm" enctype="multipart/form-data" data-toggle="validator" role="form" action="/stages" accept-charset="UTF-8" method="post"><input name="utf8" type="hidden" value="✓" /><input type="hidden" name="authenticity_token" value="<KEY> /><div class="alert alert-danger text-center" hidden="true" id="uploadErrorWarning"></div><div class="form-group has-feedback"><div class="input-group"><span class="input-group-addon upload-addon">Notebook</span><input accept=".ipynb" class="form-control" id="uploadFile" name="file" required="" type="file" /></div><span aria-hidden="true" class="glyphicon form-control-feedback"></span></div><div class="form-group"><div class="alert alert-info agree"><div class="checkbox"><label><input name="agree" required="" type="checkbox" value="yes" /><p>I acknowledge that I have all intellectual property rights and approvals (if applicable)
# <br>
# for the content contained within this notebook.
# </p><p>Also I acknowledge that the nbgallery will remove all output from my notebook and will only save the code and markup sections</p></label></div></div></div><div class="modal-footer"><div class="form-group"><div style="float:right"><button class="btn btn-primary" id="uploadFileSubmit" type="submit">Next</button>
# ## PART 2
#
# <center><h4 class="modal-title">Upload Notebook - Part 2 of 2</h4></center></div><div class="modal-body"><form id="stageForm" enctype="multipart/form-data" data-toggle="validator" role="form" action="/notebooks" accept-charset="UTF-8" method="post"><input name="utf8" type="hidden" value="✓" /><input type="hidden" name="authenticity_token" value="<KEY> /><div class="alert alert-danger text-center" hidden="true" id="stageErrorWarning"></div><div class="form-group has-feedback"><div class="input-group"><span class="input-group-addon upload-addon">Notebook</span><input class="form-control" id="stagedName" name="staged" readonly="" type="text" value="" /></div><span aria-hidden="true" class="glyphicon form-control-feedback"></span></div><div class="form-group has-feedback"><div class="input-group"><span class="input-group-addon upload-addon">Title</span><input class="form-control" id="stageTitle" name="title" placeholder="Enter a title for your notebook" required="" type="text" /></div><div class="help-block with-errors"></div><span aria-hidden="true" class="glyphicon form-control-feedback"></span></div><div class="form-group" hidden="true" id="stageOverwrite"><div class="alert alert-danger overwrite"><div class="checkbox"><label><input name="overwrite" type="checkbox" value="true" /><b>Overwrite this notebook</b></label></div></div></div><div class="form-group has-feedback"><div class="input-group"><span class="input-group-addon upload-addon">Description</span><textarea class="form-control" id="stageDescription" name="description" placeholder="Enter a description of this notebook" required=""></textarea></div><div class="help-block with-errors"></div><span aria-hidden="true" class="glyphicon form-control-feedback"></span></div><div class="form-group"><div class="input-group"><span class="input-group-addon upload-addon">Ownership</span><select class="form-control" id="stageOwnership" name="owner"><option value="self">Myself</option></select></div></div><div class="form-group"><div class="input-group" data-toggle="tooltip" title="Enter tags here"><span class="input-group-addon upload-addon">Tags</span><input class="form-control" id="stageTags" name="tags" type="text" /></div></div><div class="form-group"><div class="checkbox"><label><input id="stagePrivate" name="private" type="checkbox" value="true" />This notebook is private (optional)</label></div></div><input id="stagingId" name="staging_id" type="hidden" value="" /><div class="form-group"><div class="alert alert-info agree"><div class="checkbox"><label><input name="agree" required="" type="checkbox" value="yes" /><p>I acknowledge that I have all intellectual property rights and approvals (if applicable)
# <br>
# for the content contained within this notebook.
# </p><p>Also I acknowledge that the nbgallery will remove all output from my notebook and will only save the code and markup sections</p></label></div></div></div><div class="modal-footer"><div class="form-group"><div style="float:right"><button class="btn btn-primary" id="stageSubmit" type="submit">Submit</button></div></div><div id="stageFeedbackProgressBar"></div></div></form>
# + active=""
# curl 'http://localhost:3000/notebooks/4-sfs' -H 'Connection: keep-alive' -H 'Upgrade-Insecure-Requests: 1' -H 'DNT: 1' -H 'User-Agent: Mozilla/5.0 (Macintosh; Intel Mac OS X 10_13_6) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/70.0.3538.110 Safari/537.36' -H 'Accept: text/html,application/xhtml+xml,application/xml;q=0.9,image/webp,image/apng,*/*;q=0.8' -H 'Referer: http://localhost:3000/notebooks/3-tesd' -H 'Accept-Encoding: gzip, deflate, br' -H 'Accept-Language: en-US,en;q=0.9' -H 'Cookie: _xsrf=2|d3a53387|f4cb6517674e957b0f0bdf072d676208|1547211073; username-localhost-8888="2|1:0|10:1547211134|23:username-localhost-8888|44:Nzc2ODg5ZDQyYjU2NDdhMTk5M2VlMmYwMTliNTI4MDU=|73e07fb8e5374796fd205de243015c055e6686255ed1bb8868b850af28e7dc32"; host=.butterfly; home_viewed=1547669814; _jupyter-gallery_session=RTVRbzFJWjRzc0EyZUxYeVpyRkRSdG9ucElFM3lvSE5ERGl3V21jd2JXU1dFM0E4VmFwdm1RbXdKb3NzRnE3WXRjVTF1UXRsY2xwbXAvUUQwclZacUFLWStScmpNRTRnYjV2eEsvc013WElkNzZLem54c3R5M3F6RFdtMm8weGVlRzhLWmVWdGtBSmFVN3NwbkJ2aXNzU3BTdm1ndFJ2cTZKRnZxWnZYSkFjREFDOXcrVEJrOW1QSENTcnNBZ3BPR2RnS3pLY0hLMVVlSFhiaVhGNFNPL245ZER5RmlFTDlNUklWb3Z3R1Y5RXBBeng3elB3TUNvZXFDUUdBdkxqZXdvSnZWRFJrY3pjWko1K2JBaWVLbmJXak5tamFDVFRvb21sTkJkZUhwNVkxWldPVGpxam1MaHdPc3F6eGx4NWstLVVIa3BBdFJtRWx3ZVZldnhOYko0THc9PQ%3D%3D--cf9f049229c03ed5baf699ca612417046f3e2c8a' --compressed ;
#
#
#
# curl 'http://localhost:3000/stages' -X POST -H 'Cookie: _xsrf=2|d3a53387|f4cb6517674e957b0f0bdf072d676208|1547211073; username-localhost-8888="2|1:0|10:1547211134|23:username-localhost-8888|44:Nzc2ODg5ZDQyYjU2NDdhMTk5M2VlMmYwMTliNTI4MDU=|73e07fb8e5374796fd205de243015c055e6686255ed1bb8868b850af28e7dc32"; host=.butterfly; home_viewed=1547669814; _jupyter-gallery_session=cHNOcFJiTHNkZGdjdGVzV0xLanFvOG5XNW41SThyamh6dWhPZEZNMlBoMkVwK05kU1d0cmZvUE9LTWxNNlNkbWcxNWltdGo1Y0hpaHVXb0JMWHlhUVIvMEVoSUphWlVzUUd1ak9jOWdwWWg2Z1lQZUkxbjU3N0tKMXVEYm9iMzRqSWNVZ0RzRlRhZE85SUlha053TmRjWkY5eXd4TDRwcjVZWVJ5dDlaRHFRaGhncUtnTlI4MWtrdVJSclVJbkhvdC9xazNzQjR2bHFVV0xKaFJHRVZjVjNvcUk0TWxDSHZOUE5qQTA0OHFvVUFocFYyM0xSRC8zSWQ2MkJ1VWo0VjZid3JzN2VTdzdWK25aeWVFMldHbStMaWNjYzA1QUlBREJXUlVMSDNER1h5N2FjSkw3RDQwdGRiaFdPY3AyQVAtLStyaDVQTU5Gb0xLS3hZVmJqOFRETUE9PQ%3D%3D--3940fc4cfef7b7b75f95e414fcf01a06e577e912' -H 'Origin: http://localhost:3000' -H 'Accept-Encoding: gzip, deflate, br' -H 'X-CSRF-Token: Id0UKstNSDZx+bkGSOkDqpHRpLU+RJKeSUTwNBiZNxAnLtHwBmcQ7IfkiRaQsIazD8yBPNfWnEXPKlnFsgB8qQ==' -H 'Accept-Language: en-US,en;q=0.9' -H 'User-Agent: Mozilla/5.0 (Macintosh; Intel Mac OS X 10_13_6) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/70.0.3538.110 Safari/537.36' -H 'Content-Type: multipart/form-data; boundary=----WebKitFormBoundaryz389yaDTcGKyBqQm' -H 'Accept: */*' -H 'Referer: http://localhost:3000/notebooks/3-tesd' -H 'X-Requested-With: XMLHttpRequest' -H 'Connection: keep-alive' -H 'Content-Length: 586538' -H 'DNT: 1' --compressed ;
#
#
#
# curl 'http://localhost:3000/stages/fb0ca8f6-3ff9-41b6-88c8-edb0e0d2b9a7/preprocess' -H 'Cookie: _xsrf=2|d3a53387|f4cb6517674e957b0f0bdf072d676208|1547211073; username-localhost-8888="2|1:0|10:1547211134|23:username-localhost-8888|44:Nzc2ODg5ZDQyYjU2NDdhMTk5M2VlMmYwMTliNTI4MDU=|73e07fb8e5374796fd205de243015c055e6686255ed1bb8868b850af28e7dc32"; host=.butterfly; home_viewed=1547669814; _jupyter-gallery_session=SlA4d1dUQUV3K0UwTFVlN1FTN1RSdm1obis5Wng2cmNhNkNzMVFZWWlDQWd1SHV5dktXaWhxd21sV1BsNW5tZlpRSkVBZEY0OFY0MWliS2VhdWFpaFJ1UUNIYlRiUzhZWkdQRzdZYmc5VUVpK1hKY2pnT2N5d0M3SkV0Qk1hOUVNQksyM1BRQWt5OWJTTHR6OGVmNTNiS1Y5bTBiWWZKUmhvUmRBenZMdnQxZ1RYbnhRTlRzTFhoN0pvWHJJR1o0YzNucy9CQ24vK0YyTVRoNVFHMlZCc1hYdzRWUTZVNXUzWFAvejFCRVdrWTJZVHJyU1NFejFxdU9tWjhkc0pjTHk5SEJRdVA3alFRU0FiZTU1VytKN2J1TDNKRmxRUjhQY1lTOUJwMS81MXNTNXpaUUhNbTJJdXV2WUd0aGJabGwtLTZjWkRkNmtPbTltdlhJcnJoNGVvaGc9PQ%3D%3D--d91feb64721a2ea09b9d535ac5416fdb66b4e8ea' -H 'DNT: 1' -H 'Accept-Encoding: gzip, deflate, br' -H 'X-CSRF-Token: <KEY> -H 'Accept-Language: en-US,en;q=0.9' -H 'User-Agent: Mozilla/5.0 (Macintosh; Intel Mac OS X 10_13_6) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/70.0.3538.110 Safari/537.36' -H 'Accept: application/json' -H 'Referer: http://localhost:3000/notebooks/3-tesd' -H 'X-Requested-With: XMLHttpRequest' -H 'Connection: keep-alive' --compressed ;
#
#
#
# curl 'http://localhost:3000/tags' -H 'Cookie: _xsrf=2|d3a53387|f4cb6517674e957b0f0bdf072d676208|1547211073; username-localhost-8888="2|1:0|10:1547211134|23:username-localhost-8888|44:Nzc2ODg5ZDQyYjU2NDdhMTk5M2VlMmYwMTliNTI4MDU=|73e07fb8e5374796fd205de243015c055e6686255ed1bb8868b850af28e7dc32"; host=.butterfly; home_viewed=1547669814; _jupyter-gallery_session=M2ZlMDM4ejl5V1poMDBkd0sxWmJJRDh0Mit2UzlsbU5SOTZTeXBGNTJNNW5UUEVKa2Y3MEMvUms0bW5UNHl0OWoxQ1pVRUtIWTN2ZkVPaVpmRUxhL1lPbW03aTJobUdQQnNiNGVEMXpzMWQzVVd1eUxUNFErWmZ4ZElSUkg1aEZ5eUY5SytEYSsyWUZtSXU5L2x2b0p0TDR4a3h3N0xPb0Q5d1lTN0RyOG9raDVoa2gyc3UweHN4OGU0NHB5amRLOXpKMUlpUUNnSUh2REFTTkRRK3RCZGt3SWNZV3JDZS92Ly82U2I3bSsyR1c2VlJ2L3Q1bG9xWW0wM091R05TRUtkckpvUmVweTJlTi9QZEY2blQ0aHE0UUZyeXlZOVR0T2Y5bEVEMXdpM2FROXlQeE5jc0l2RzlTckhDbzdiWDAtLVJ6R1luM0pDcEYvVm5IcUtMNkVyT1E9PQ%3D%3D--6d662299581e0f99ebc2aa8bd41eb08f8ce08a93' -H 'DNT: 1' -H 'Accept-Encoding: gzip, deflate, br' -H 'X-CSRF-Token: <KEY> -H 'Accept-Language: en-US,en;q=0.9' -H 'User-Agent: Mozilla/5.0 (Macintosh; Intel Mac OS X 10_13_6) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/70.0.3538.110 Safari/537.36' -H 'Accept: application/json' -H 'Referer: http://localhost:3000/notebooks/3-tesd' -H 'X-Requested-With: XMLHttpRequest' -H 'If-None-Match: W/"d751713988987e9331980363e24189ce"' -H 'Connection: keep-alive' --compressed ;
#
#
#
# curl 'http://localhost:3000/assets/images/ui-bg_highlight-soft_100_eeeeee_1x100.png' -H 'DNT: 1' -H 'Accept-Encoding: gzip, deflate, br' -H 'Accept-Language: en-US,en;q=0.9' -H 'User-Agent: Mozilla/5.0 (Macintosh; Intel Mac OS X 10_13_6) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/70.0.3538.110 Safari/537.36' -H 'Accept: image/webp,image/apng,image/*,*/*;q=0.8' -H 'Referer: http://localhost:3000/assets/application-ffa3df68632b1ad85efd4685b2c250772406e5fd588d8c278041469ee99a0526.css' -H 'Cookie: _xsrf=2|d3a53387|f4cb6517674e957b0f0bdf072d676208|1547211073; username-localhost-8888="2|1:0|10:1547211134|23:username-localhost-8888|44:Nzc2ODg5ZDQyYjU2NDdhMTk5M2VlMmYwMTliNTI4MDU=|73e07fb8e5374796fd205de243015c055e6686255ed1bb8868b850af28e7dc32"; host=.butterfly; home_viewed=1547669814; _jupyter-gallery_session=TUJCWHdIeWpYRUxZZGNtTkRKcUc4dVMwVzZRcXpwY0lPUmxld21sOGN1Y1d6RVppQ0k1S0txdjFyZVJNeHhwbk5xR2xiY0tNcWRmU2trdWRDb0VWeFFDNTlKR0Y5b01IYVBFTWxBMnFxOWxXaktIRkNoK3lOVUtyd2dUckNUbVlSVHh2aTU1VWJwYlU1T1p3cXdRYXZJRkplL3lqS3M1MHdkcW9CdWhRKzBTL0xXbXhWRXBCTVk2Vy9Lb3BER0lxWE82UWtlRDM2eW9RcVhBU2JnRndHUDdFWnJLSXI3czlUb0xFeG82MEF5S0F1ZDFwWWF2NE5NeFFZSDJjSWZEdkNKTjU0UGVYME1vaDBKUHBNeEpPSER2UG5LdUpQRDdqK3k1RVIyc0hkb2Zzd3BVMnhJRVQ0TVg2TFBuNGdkY1MtLUgzeVJYTVl4TWo2b0NtZE9RSytLanc9PQ%3D%3D--80a4ec63738c87cca4cf90e4c55b31dc5ff0304a' -H 'Connection: keep-alive' --compressed ;
#
#
#
# curl 'http://localhost:3000/notebooks' -H 'Cookie: _xsrf=2|d3a53387|f4cb6517674e957b0f0bdf072d676208|1547211073; username-localhost-8888="2|1:0|10:1547211134|23:username-localhost-8888|44:Nzc2ODg5ZDQyYjU2NDdhMTk5M2VlMmYwMTliNTI4MDU=|73e07fb8e5374796fd205de243015c055e6686255ed1bb8868b850af28e7dc32"; host=.butterfly; home_viewed=1547669814; _jupyter-gallery_session=TUJCWHdIeWpYRUxZZGNtTkRKcUc4dVMwVzZRcXpwY0lPUmxld21sOGN1Y1d6RVppQ0k1S0txdjFyZVJNeHhwbk5xR2xiY0tNcWRmU2trdWRDb0VWeFFDNTlKR0Y5b01IYVBFTWxBMnFxOWxXaktIRkNoK3lOVUtyd2dUckNUbVlSVHh2aTU1VWJwYlU1T1p3cXdRYXZJRkplL3lqS3M1MHdkcW9CdWhRKzBTL0xXbXhWRXBCTVk2Vy9Lb3BER0lxWE82UWtlRDM2eW9RcVhBU2JnRndHUDdFWnJLSXI3czlUb0xFeG82MEF5S0F1ZDFwWWF2NE5NeFFZSDJjSWZEdkNKTjU0UGVYME1vaDBKUHBNeEpPSER2UG5LdUpQRDdqK3k1RVIyc0hkb2Zzd3BVMnhJRVQ0TVg2TFBuNGdkY1MtLUgzeVJYTVl4TWo2b0NtZE9RSytLanc9PQ%3D%3D--80a4ec63738c87cca4cf90e4c55b31dc5ff0304a' -H 'Origin: http://localhost:3000' -H 'Accept-Encoding: gzip, deflate, br' -H 'X-CSRF-Token: <KEY> -H 'Accept-Language: en-US,en;q=0.9' -H 'User-Agent: Mozilla/5.0 (Macintosh; Intel Mac OS X 10_13_6) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/70.0.3538.110 Safari/537.36' -H 'Content-Type: multipart/form-data; boundary=----WebKitFormBoundaryCw6eZsEA0AIxw8vG' -H 'Accept: */*' -H 'Referer: http://localhost:3000/notebooks/3-tesd' -H 'X-Requested-With: XMLHttpRequest' -H 'Connection: keep-alive' -H 'DNT: 1' --data-binary $'------WebKitFormBoundaryCw6eZsEA0AIxw8vG\r\nContent-Disposition: form-data; name="utf8"\r\n\r\n✓\r\n------WebKitFormBoundaryCw6eZsEA0AIxw8vG\r\nContent-Disposition: form-data; name="authenticity_token"\r\n\r\nId0UKstNSDZx+bkGSOkDqpHRpLU+RJKeSUTwNBiZNxAnLtHwBmcQ7IfkiRaQsIazD8yBPNfWnEXPKlnFsgB8qQ==\r\n------WebKitFormBoundaryCw6eZsEA0AIxw8vG\r\nContent-Disposition: form-data; name="staged"\r\n\r\nfb0ca8f6-3ff9-41b6-88c8-edb0e0d2b9a7\r\n------WebKitFormBoundaryCw6eZsEA0AIxw8vG\r\nContent-Disposition: form-data; name="title"\r\n\r\nsfs\r\n------WebKitFormBoundaryCw6eZsEA0AIxw8vG\r\nContent-Disposition: form-data; name="description"\r\n\r\ndesc\r\n------WebKitFormBoundaryCw6eZsEA0AIxw8vG\r\nContent-Disposition: form-data; name="owner"\r\n\r\nself\r\n------WebKitFormBoundaryCw6eZsEA0AIxw8vG\r\nContent-Disposition: form-data; name="tags"\r\n\r\n\r\n------WebKitFormBoundaryCw6eZsEA0AIxw8vG\r\nContent-Disposition: form-data; name="staging_id"\r\n\r\nfb0ca8f6-3ff9-41b6-88c8-edb0e0d2b9a7\r\n------WebKitFormBoundaryCw6eZsEA0AIxw8vG\r\nContent-Disposition: form-data; name="agree"\r\n\r\nyes\r\n------WebKitFormBoundaryCw6eZsEA0AIxw8vG--\r\n' --compressed ;
#
# -
FORM DATA:
utf8: ✓
authenticity_token: Id0UKstNSDZx bkGSOkDqpHRpLU RJKeSUTwNBiZNxAnLtHwBmcQ7IfkiRaQsIazD8yBPNfWnEXPKlnFsgB8qQ==
staged: fb0ca8f6-3ff9-41b6-88c8-edb0e0d2b9a7
title: sfs
description: desc
owner: self
tags:
staging_id: fb0ca8f6-3ff9-41b6-88c8-edb0e0d2b9a7
agree: yes
| broken/nbgallery scraper.ipynb |
# ---
# jupyter:
# jupytext:
# formats: ipynb,py:light
# notebook_metadata_filter: nbsphinx
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# nbsphinx:
# execute: never
# ---
# Evaluating ideal ratio mask on WHAM!
# =====================================
#
# This recipe evaluates an oracle ideal ratio mask on the `mix_clean`
# and `min` subset in the WHAM dataset. This recipe is annotated
# as a notebook for documentation but can be run directly
# as a script in `docs/recipes/ideal_ratio_mask.py`.
#
# We evaluate three approaches to constructing the ideal ratio mask:
#
# - Magnitude spectrum approximation
# - Phase sensitive spectrum approximation
# - Truncated phase sensitive spectrum approximation
#
# Imports
# ----------
# +
from nussl import datasets, separation, evaluation
import os
import multiprocessing
from concurrent.futures import ThreadPoolExecutor
import logging
import json
import tqdm
import glob
import numpy as np
import termtables
# set up logging
logger = logging.getLogger()
logger.setLevel(logging.INFO)
# -
# Setting up
# ----------
#
# Make sure to point `WHAM_ROOT` where you've actually
# built and saved the WHAM dataset. There's a few different
# ways to use ideal ratio masks, so we're going to set those
# up in a dictionary.
# +
WHAM_ROOT = '/home/data/wham/'
NUM_WORKERS = multiprocessing.cpu_count() // 4
OUTPUT_DIR = os.path.expanduser('~/.nussl/recipes/ideal_ratio_mask/')
APPROACHES = {
'Phase-sensitive spectrum approx.': {
'kwargs': {
'range_min': -np.inf, 'range_max':np.inf
},
'approach': 'psa',
'dir': 'psa'
},
'Truncated phase-sensitive approx.': {
'kwargs': {
'range_min': 0.0, 'range_max': 1.0
},
'approach': 'psa',
'dir': 'tpsa'
},
'Magnitude spectrum approximation': {
'kwargs': {},
'approach': 'msa',
'dir': 'msa'
}
}
RESULTS_DIR = os.path.join(OUTPUT_DIR, 'results')
for key, val in APPROACHES.items():
_dir = os.path.join(RESULTS_DIR, val['dir'])
os.makedirs(_dir, exist_ok=True)
# -
# Evaluation
# ----------
# +
test_dataset = datasets.WHAM(WHAM_ROOT, sample_rate=8000, split='tt')
for key, val in APPROACHES.items():
def separate_and_evaluate(item):
output_path = os.path.join(
RESULTS_DIR, val['dir'], f"{item['mix'].file_name}.json")
separator = separation.benchmark.IdealRatioMask(
item['mix'], item['sources'], approach=val['approach'],
mask_type='soft', **val['kwargs'])
estimates = separator()
evaluator = evaluation.BSSEvalScale(
list(item['sources'].values()), estimates, compute_permutation=True)
scores = evaluator.evaluate()
with open(output_path, 'w') as f:
json.dump(scores, f)
pool = ThreadPoolExecutor(max_workers=NUM_WORKERS)
for i, item in enumerate(tqdm.tqdm(test_dataset)):
if i == 0:
separate_and_evaluate(item)
else:
pool.submit(separate_and_evaluate, item)
pool.shutdown(wait=True)
json_files = glob.glob(f"{RESULTS_DIR}/{val['dir']}/*.json")
df = evaluation.aggregate_score_files(json_files)
overall = df.mean()
print(''.join(['-' for i in range(len(key))]))
print(key.upper())
print(''.join(['-' for i in range(len(key))]))
headers = ["", f"OVERALL (N = {df.shape[0]})", ""]
metrics = ["SAR", "SDR", "SIR"]
data = np.array(df.mean()).T
data = [metrics, data]
termtables.print(data, header=headers, padding=(0, 1), alignment="ccc")
| docs/recipes/wham/ideal_ratio_mask.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Installation
# ---
# ## TLDR
pip install --upgrade aiqc
# If that doesn't work, read the rest of this notebook (e.g. supported Python versions). As always, restart kernel if you are using a computational notebook.
import aiqc
aiqc.setup()
# `setup()` only needs to be ran once. Going forward, just `import aiqc`. Don't use `as` when importing.
# > Technically, the backend files created by `setup()` could be created automatically upon import, but we don't want to get in the habit of running system commands without explicit user consent.
# The database is simply a SQLite **file**, and AIQC serves as an ORM/ API for that SQL database.
#
# > So you ***do not*** have to worry about anything like installing a database server, database client, database users, configuring ports, configuring passwords/ secrets/ environment variables, or starting and restopping the database. Shoutout to the [ORM, peewee](http://docs.peewee-orm.com/en/latest/index.html). Glad we found this fantastic and simple alternative to SQLAlchemy.
# Finally, be sure to read the plotting section below.
# ---
# ## Environment
# ### Python Version
#
# Requires Python 3+ (check your deep learning library's Python requirements). AIQC was developed on Python 3.7.12 in order to ensure compatibility with Google Colab.
#
# We highly recommend `pyenv` for managing Python installations and virtualenvs.
#
# Additionally, check the Python version required by the machine learning libraries that you intend to use. For example, at the time this was written, Tensorflow/ Keras required Python 3.5–3.8. If you need more information about dependencies, the PyPI `setup.py` is in the root of the github.com/aiqc/aiqc repository.
import sys
sys.version
# #### Pickle Disclaimer
#
# AIQC, much like PyTorch, relies heavily on [Pickle](https://docs.python.org/3/library/pickle.html) for saving Python objects in its database. Therefore, as a caveat of Pickle, if you create objects in your `aiqc.sqlite` file using one version of Python and try to interact with it on a newer version of Python, then you may find that pickle is no longer able to deserialize the object. For this reason, `sys.version` and other helpful info about your OS/ Python version is stored in the `config.json` file at the time of creation.
# ### Operating System
# AIQC was designed to be OS-agnostic. It has been tested on the following operating systems:
#
# - macOS 10.15 and 11.6.1
# - Linux (Ubuntu, Alpine, RHEL).
# - Windows 10 (and WSL).
#
# > If you run into trouble with the installation process on your OS, please submit a GitHub issue so that we can attempt to resolve, document, and release a fix as quickly as possible.
# ### Optional - JupyterLab IDE
# AIQC runs anywhere Python runs. We just like Jupyter for interactive visualization and data transformation. FYI, *jupyterlab* is not an official dependency of the AIQC package.
pip install jupyterlab
# JupyterLab requires Node.js >= 10. Once all extensions switch to JupyterLab 3.0 prebuilding, this will no longer be necessary.
# !node -v
# ### Plotting
# AIQC uses Plotly for interactive charts.
pip install --upgrade plotly
# + [markdown] tags=[]
# #### Optional - Plotly in JupyterLab
# This requires the following dependencies:
# -
# * Python packages: [`ipywidgets`, `plotly`]
# * JupyterLab Extension: [`jupyterlab-plotly`].
#
# > https://github.com/plotly/plotly.py#jupyterlab-support
pip install ipywidgets
# ##### New: simplified, pre-built extension for plotly>=5.0.0
# The plotly JupyterLab Extension is now pre-built and included in the Plotly Python package. If you install plotly while you have a JupyterLab server running, then do a hard restart (not kernel reset) of the server for it to take effect.
# ##### *Deprecated:* manually building extension process for plotly<=5.0.0
# +
# Deprecated
# #!jupyter labextension install jupyterlab-plotly
# #!jupyter labextension check jupyterlab-plotly
# #!jupyter lab build
#After the build completes (typically takes a few minutes) restart your Jupyter server by interrupting the command line process (not restarting kernel).
# -
# ### Optional - Swap Space for Failover Memory
# On local machines, it is good practice to configure “swap space” so that if you run out of memory/ RAM it will simply spill over onto the swap partition (dynamic is possible) of your hard drive as opposed to risking out-of-memory crashes. For GB sized datasets, spinning media HDDs (5,400/ 7,200 RPM) may be too slow for using swap in production, but you can get by with NVMe/ SSD.
# ---
# ## AIQC Package
pip install --upgrade aiqc
# Troubleshooting:
#
# * Be sure to include `--upgrade` when running the command above.
# * The command above must often be ran twice (PyPI bug).
# * Upgrade `pip` itself: `pip install --upgrade pip`
# * Install`wheel` is recommended: `pip install --upgrade wheel`
import aiqc
# ---
# ## AIQC Config & Database
# ### a) One-Line Install
# The `setup()` method just runs the following methods in one go: `create_folder()`, `create_config()`, and `create_db()`.
aiqc.setup()
# ### b) Or Line-by-Line Configuration
# If you need to troubleshoot your installation, then run the installation steps one-by-one.
# #### Create Config File
# Enter the following commands one-by-one and follow any instructions returned by the command prompt to resolve errors should they arise.
aiqc.create_folder()
aiqc.create_config()
# If you run the `.create_*()` commands in the future, don't worry, they won't overwrite your existing data. They will detect the presence of the data and skip creation.
# #### Create Database File
# This creates a SQLite database file that AIQC uses to reproducibly record experiments.
# Only now that the configuration exists can we create the database tables. After creating the config, the AIQC module should reload itself automatically. If you encounter errors, reference the Troubleshooting section below.
aiqc.create_db()
# ### Config File
# The configuration file contains low level information about:
# * Where AIQC should persist data.
# * Runtime (Python, OS) environment for reproducibility and troubleshooting.
aiqc.get_config()
# ---
# ## Location of AIQC Files
# AIQC makes use of the Python package, `appdirs`, for an operating system (OS) agnostic location to store configuration and database files. This not only keeps your `$HOME` directory clean, but also helps prevent careless users from deleting your database.
#
# > The installation process checks not only that the corresponding appdirs folder exists on your system but also that you have the permissions neceessary to read from and write to that location. If these conditions are not met, then you will be provided instructions during the installation about how to create the folder and/ or grant yourself the appropriate permissions.
#
# > We have attempted to support both Windows (`icacls` permissions and backslashes `C:\\`) as well as POSIX including Mac and Linux including containers & Google Colab (`chmod letters` permissions and slashes `/`). Note: due to variations in the ordering of appdirs author and app directories in different OS', we do not make use of the appdirs `appauthor` directory, only the `appname` directory.
#
# ### Location Based on OS
#
# Test it for yourself: <br/>
# `import appdirs; appdirs.user_data_dir('aiqc');`
#
# * Mac: <br />`/Users/Username/Library/Application Support/aiqc`
#
# * Linux - Alpine and Ubuntu: <br />`/root/.local/share/aiqc`
#
# * Windows: <br />`C:\Users\Username\AppData\Local\aiqc`
# ---
# ## Optional - Deleting the Database
# If, for whatever reason, you find that you need to destroy your SQLite database file and start from scratch, then you can do so without having to manually find and `rm` the database file. In order to reduce the chance of an accident, `confirm:bool=False` by default.
# > Bear in mind that if you are on either a server or shared OS, then this database may contain more than just your data.
# ### a) One-Liner
# Both `confirm:bool=False` and `rebuild:bool=False`, so it only does what you command it to do.
aiqc.destroy_db(confirm=True, rebuild=True)
# ### b) Or Line-by-Line
aiqc.destroy_db(confirm=True)
aiqc.create_db()
# ---
# ## Troubleshooting
# ### Reloading the Package
# After CRUD'ing the config files, AIQC needs the be reimported in order to detect those changes. This can be done in one of three ways:
#
# * If everything goes smoothly, it should automatically happen behind the scenes: `reload(sys.modules['aiqc'])`.
# * Manually by the user: `from importlib import reload; reload(aiqc)`.
# * Manually restarting your Python kernel/ session and `import aiqc`.
| docs/notebooks/installation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Generate Multivariate Dataset
#
# Expression for a chaotic time series:
#
# $\quad X[n + 1] = X[n]^2 + \epsilon[n] + b, \quad \epsilon \sim \mathcal{N} (0, \sigma_n^2)$
#
# for a chaotic time series $b = -1.9$ and $X[1] = 0.5$.
#
# Generate correlated time series:
#
# $\quad X[n + 1]_1 = X[n]_1^2 + \epsilon[n]_1 + b$
#
# $\quad\vdots$
#
# $\quad X[n + 1]_t = X[n + 1]_{t - 1} - a \cdot ( X[n + 1]_{t - 1} - 1) + \epsilon[n]_t$
#
# $\quad\vdots$
#
# $\quad X[n + 1]_T = X[n + 1]_{T - 1} - a \cdot ( X[n + 1]_{T - 1} - 1) + \epsilon[n]_T$
#
#
# the task index is $t = \{ 1, \dots, T \}$, T is tasks number, $a$ is any correlation constant, In this experiment is $a = \frac{3\cdot t}{2}$. And if there is noise, it is independet for each taks, so $\epsilon_t \sim \mathcal{N} (0, \sigma_n^2)$.
# +
import numpy as np
import matplotlib.pyplot as plt
# Generate Correlated Time Series Noiseless or with white Noise added
def generate_multivariate_predictiors_serie(X, T = 2, mu = 0, s2 = 0., a = 3/2):
N = X.shape[0]
Y = np.empty((N, T))
E = np.random.normal(mu, s2, N)
Y[..., 0] = X + E
for t in range(1, T):
E = np.random.normal(mu, s2, N)
Y[..., t] = Y[..., t - 1] - a*t*(Y[..., t - 1] - 1.) + E
return Y
# Generate Chaotic System
def generate_chaotic_time_series(N, c = -1.9, x_n = .5):
X = [x_n]
for i in range(N):
x_n_1 = X[-1]**2 + c
x_n = x_n_1
X.append(x_n)
return np.asarray(X)
# Number of sample on dataset
N = 300
# Dataset Chaotic Parameters
X = generate_chaotic_time_series(N, c = -1.9, x_n = .5)
print(X.shape)
X = generate_multivariate_predictiors_serie(X, T = 4, mu = 0., s2 = 0.25, a = 3./2.)
print(X.shape)
# -
plt.figure(figsize = (15, 5))
plt.plot(X[:, 0])
plt.plot(X[:, 1])
plt.plot(X[:, 2])
plt.plot(X[:, 3])
plt.show()
| Example - Correlated Multivariate Dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 2. Linear Algebra_3. Matrix
# Matrix는 Systems of Linear Equation을 compact하게 표현해주는 것 뿐만 아니라, Linear functions(Linear mapping)의 역할도 수행하는 중요한 개념이다.
#
# Matrix는 연산 시, element-wise 연산을 기본적으로 수행한다. vector와 matrix간 연산도 마찬가지이다.
#
# 단, 연산 시 행과 열의 숫자에 주의해야 한다
A = np.array([[1,2,3],[3,4,5],[5,6,7]])
v = np.array([[1],[2],[3]])
B = np.array([[10,20,30],[30,40,50],[50,60,70]])
A@B
B@A
A@v
# +
# 연산 시, 행과 열의 숫자가 맞지 않아 오류가 난 경우이다.
# v = (3,1), A = (3,3) 이므로 v@A는 연산수행이 불가능하다.
v@A
# -
# Matrix는 Associativity(결합법칙), Distributivity(분배법칙)이 성립한다. 단, **교환법칙은 성립하지 않는다.**
# + [markdown] school_cell_uuid="9e6d469db699456a97a81171ce73befe"
# 행렬의 곱셈은 곱하는 행렬의 순서를 바꾸는 교환 법칙이 성립하지 않는다. 그러나 덧셈에 대한 분배 법칙은 성립한다.
#
# $$
# \begin{align}
# AB \neq BA
# \tag{2.2.53}
# \end{align}
# $$
#
# $$
# \begin{align}
# A(B + C) = AB + AC
# \tag{2.2.54}
# \end{align}
# $$
#
# $$
# \begin{align}
# (A + B)C = AC + BC
# \tag{2.2.55}
# \end{align}
# $$
#
#
# $A$, $B$, $C$가 다음과 같을 때 위 법칙을 넘파이로 살펴보자.
#
# $$
# \begin{align}
# A = \begin{bmatrix} 1 & 2 \\ 3 & 4 \end{bmatrix}
# \tag{2.2.56}
# \end{align}
# $$
#
# $$
# \begin{align}
# B = \begin{bmatrix} 5 & 6 \\ 7 & 8 \end{bmatrix}
# \tag{2.2.57}
# \end{align}
# $$
#
# $$
# \begin{align}
# C = \begin{bmatrix} 9 & 8 \\ 7 & 6 \end{bmatrix}
# \tag{2.2.58}
# \end{align}
# $$
# -
A = np.array([[1, 2], [3, 4]])
B = np.array([[5, 6], [7, 8]])
C = np.array([[9, 8], [7, 6]])
# + [markdown] school_cell_uuid="8199e602364f46c2855d5776c470e3f1"
# $AB$ 와 $BA$의 값은 다음처럼 다른 값이 나오므로 교환법칙이 성립하지 않음을 알 수 있다.
# -
A@B
B@A
A@B == B@A
# + [markdown] school_cell_uuid="c99ddcd80a574a178a0f5b7d97acbdab"
# 분배법칙은 다음과 같이 성립한다.
# -
A @ (B + C) == A@B + A@C
# Identity Matrix(항등행렬)도 존재한다. Identity Matrix는 "N x N matrix which contains '1' on the diagonal and '0' everywhere else" 로 정의된다.
#
# Numpy의 eye 메소드를 통해 구현 가능하다.
np.eye(3)
# 이러한 identity matrix는 역행렬을 정의하는 데 활용된다.
#
# $$ AB = BA = I, B = A^{-1}$$
A
A_inverse = np.linalg.inv(A)
A_inverse
A@A_inverse
# 이 밖에도 대칭행렬 등 다양한 행렬이 존재하며, 이후 행렬연산을 통한 Linear Equation problem 해결 및 problem solving 을 수행할 수 있다.
| 1.Study/2. with computer/1.Math_code/5. Mathematics for Machine Learning/2. Linear Algebra_3. Matrix.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: IRIS Python
# language: python
# name: irispython
# ---
# # IRIS Running process from Python
#
#
# +
import pandas as pd
pd.options.display.max_rows = 9
import iris
statement = iris.sql.exec('SELECT * FROM %SYS.ProcessQuery')
df = statement.dataframe()
jsn = df.to_json
print(jsn)
#df
df.dtypes
print(df.columns)
print(df.size)
#pd.DataFrame(index=[0, 1, 2, 3])
#df = pd.read_csv('/opt/irisapp/misc/titanic.csv')
#print(df.to_string())
#df
#print(df.head(10))
#print(df.info())
#df
#df.duplicated()
#df.corr()
# -
| src/Notebooks/IRISPyRunningProcesses.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:py3-tf2]
# language: python
# name: conda-env-py3-tf2-py
# ---
import pandas as pd
import spacy
nlp = spacy.load('en_core_web_sm', disable = ['ner', 'parser'])
# ## Use USPTO-2M dataset
df = pd.read_csv('../USPTO-2M/uspto_2m.tsv', sep='\t')
df.head()
# +
#only keep digital patent numbers
#df['No'] = df['No'].map(lambda x: x[3:])
# -
df = df.dropna().reset_index(drop=True)
len(df)
df.columns
abst_list = list(df['Abstract'])
title_list = list(df['Title'])
all_contents = abst_list + title_list
len(all_contents)
all_contents[0:2]
# ## Preprocessing
import re
import time
from nltk.tokenize import word_tokenize
from nltk.tokenize import sent_tokenize
#remove non-alphabetic characters
temp_all_contents = [re.sub('[^a-zA-Z]', ' ', str(item)).lower() for item in all_contents]
len(temp_all_contents)
# +
#use Spacy for tokenization
start_time = time.time()
test_sentence = temp_all_contents[0:10000]
all_words_list = []
for sent in nlp.pipe(test_sentence, batch_size=50, n_threads=4):
tmp_word_list = [token.text for token in sent]
all_words_list.append(tmp_word_list)
print("--- %s seconds ---" % (time.time() - start_time))
# -
#use NLTK for tokenization - faster
start_time = time.time()
test_sentence = temp_all_contents[0:100000]
all_words = [word_tokenize(sent) for sent in test_sentence]
print("--- %s seconds ---" % (time.time() - start_time))
start_time = time.time()
all_words_list = [word_tokenize(sent) for sent in temp_all_contents]
print("--- %s seconds ---" % (time.time() - start_time))
# ## Training the model - gensim word2vec
train_set = all_words_list[0:1998373]
# +
from gensim.models import Word2Vec
import logging
import multiprocessing
cores = multiprocessing.cpu_count()
logging.basicConfig(format='%(asctime)s : %(levelname)s : %(message)s', level=logging.INFO)
start_time = time.time()
model = Word2Vec(train_set,
size = 100,
sg = 1,
window = 5,
min_alpha = 0.001,
alpha = 0.05,
workers=cores-1,
iter = 5
)
print("--- %s seconds ---" % (time.time() - start_time))
# -
#save the model
model.wv.save_word2vec_format('uspto_2m_abstract_word2vec.bin', binary=True)
model.wv.similarity('computer','program')
#analogy
model.wv.most_similar(positive=['bottle', 'chip'], negative=['computer'], topn=1)
# ## Visualization
# ### 1. PCA
from sklearn.decomposition import PCA
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
len(model.wv.vocab)
selected_words = ['physics', 'chemistry', 'vehicle', 'plastics', 'hygiene', 'photography', 'electric']
words = selected_words.copy()
similar_words = {}
for key in selected_words:
similar_words[key] = [item[0] for item in model.wv.most_similar(key, topn=5)]
similar_words['physics']
# +
for key,value in similar_words.items():
words = words + value
#get vectors for all the words
sample_wv = model.wv[words]
# -
fig = plt.figure(1, figsize=(20, 15))
pca = PCA(n_components=2)
result = pca.fit_transform(sample_wv)
plt.scatter(result[:, 0], result[:, 1])
for i, word in enumerate(words):
plt.annotate(word, xy=(result[i, 0], result[i, 1]))
# ### 2. t-SNE
from sklearn.manifold import TSNE
import numpy as np
# +
tsne = TSNE(n_components=2, random_state=0, n_iter=10000, perplexity=15)
np.set_printoptions(suppress=True)
result = tsne.fit_transform(sample_wv)
labels = words
plt.figure(figsize=(15, 15))
plt.scatter(result[:, 0], result[:, 1], c='orange', edgecolors='r')
for label, x, y in zip(labels, result[:, 0], result[:, 1]):
plt.annotate(label, xy=(x+1, y+1), xytext=(0, 0), textcoords='offset points')
# -
# Future work for phrases:
# 1) use doc2vec
# 2) average all word vectors in the phrases
# 3) create phrases during tokenization
#
# In the experiment, the stopwords have not been removed, becaused it might remove the dependency information of the words from the sentences.
| wordEmbedding_word2vec.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from queue import Queue
def reverse(queue):
while True:
n = int(input("Enter number: "))
if n == -1:
break
queue.put(n)
stack = []
while not queue.empty():
stack.append(queue.queue[0])
queue.get()
while len(stack) != 0:
queue.put(stack[-1])
stack.pop()
def Print(queue):
while not queue.empty():
print(queue.queue[0], end = ",")
queue.get()
print("\n")
if __name__=="__main__":
queue = Queue()
while True:
print("1. Reverse")
print("2. display")
ch = int(input("Enter your choice: "))
if ch == 1:
reverse(queue)
elif ch == 2:
Print(queue)
else:
break
# -
| Queue/Reverse and Sorting Queue.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# formats: ipynb,jl:hydrogen
# text_representation:
# extension: .jl
# format_name: hydrogen
# format_version: '1.3'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.8.0-DEV
# language: julia
# name: julia-1.8
# ---
# %%
using Plots
using TaylorSeries
n = 15
x = range(-4π, 4π; length=1000)
plot(; legend=:outertopright)
plot!(x, sin.(x); label="sin(x)")
for k in 3:2:2n+1
f(x) = evaluate(sin(Taylor1(k)), x)
plot!(x, f.(x); label="k = $k", ls=:auto)
end
plot!(; ylim=(-10, 10), size=(600, 300))
# %%
using Plots
using QuadGK
function fourier_coeffs(f, N)
a, b = eps(), 1-eps()
c = Vector{Float64}(undef, 2N+1)
c[1] = quadgk(f, a, b)[1]
for k in 1:N
c[k+1] = quadgk(x -> cospi(2k*x)*f(x), a, b)[1]
c[k+N+1] = quadgk(x -> sinpi(2k*x)*f(x), a, b)[1]
end
c
end
function fourier_sum(x, c, N)
L = (length(c) - 1) ÷ 2
c[1] + 2sum(c[k+1]*cospi(2k*x) + c[k+L+1]*sinpi(2k*x) for k in 1:N)
end
function fourier_plot(f, N=20)
c = fourier_coeffs(f, N)
x = range(-0.6, 1.6; length=1101)
plot(; legend=:outertopright)
f_mod1(x) = f(mod(x, 1))
plot!(x, f_mod1.(x); label="f(mod(x, 1))")
for k in (N, N÷5)
f_k(x) = fourier_sum(x, c, k)
plot!(x, f_k.(x); label="k = $k")
end
plot!(; size=(600, 300))
end
# %%
f(x) = x
fourier_plot(f)
# %%
f(x) = 0 ≤ x < 0.5 ? 1.0 : 0.0
fourier_plot(f)
# %%
f(x) = (x - 0.5)^2
fourier_plot(f)
# %%
| 0005/Taylor and Fourier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# <img src="images/continuum_analytics_logo.png"
# alt="Continuum Logo",
# align="right",
# width="30%">,
#
# Introduction to Blaze
# =====================
#
# In this tutorial we'll learn how to use Blaze to discover, migrate, and query data living in other databases. Generally this tutorial will have the following format
#
# 1. `odo` - Move data to database
# 2. `blaze` - Query data in database
#
#
# Install
# -------
#
# This tutorial uses many different libraries that are all available with the [Anaconda Distribution](http://continuum.io/downloads). Once you have Anaconda install, please run these commands from a terminal:
#
# ```
# $ conda install -y blaze
# $ conda install -y bokeh
# $ conda install -y odo
# ```
#
# nbviewer: http://nbviewer.ipython.org/github/ContinuumIO/pydata-apps/blob/master/Section-1_blaze.ipynb
#
# github: https://github.com/ContinuumIO/pydata-apps
#
# <hr/>
#
#
# Goal: Accessible, Interactive, Analytic Queries
# -----------------------------------------------
#
# NumPy and Pandas provide accessible, interactive, analytic queries; this is valuable.
import pandas as pd
df = pd.read_csv('iris.csv')
df.head()
df.groupby(df.Species).PetalLength.mean() # Average petal length per species
# <hr/>
#
# But as data grows and systems become more complex, moving data and querying data become more difficult. Python already has excellent tools for data that fits in memory, but we want to hook up to data that is inconvenient.
#
# From now on, we're going to assume one of the following:
#
# 1. You have an inconvenient amount of data
# 2. That data should live someplace other than your computer
#
# <hr/>
# Databases and Python
# --------------------
#
# When in-memory arrays/dataframes cease to be an option, we turn to databases. These live outside of the Python process and so might be less convenient. The open source Python ecosystem includes libraries to interact with these databases and with foreign data in general.
#
# Examples:
#
# * SQL - [`sqlalchemy`](http://sqlalchemy.org)
# * Hive/Cassandra - [`pyhive`](https://github.com/dropbox/PyHive)
# * Impala - [`impyla`](https://github.com/cloudera/impyla)
# * RedShift - [`redshift-sqlalchemy`](https://pypi.python.org/pypi/redshift-sqlalchemy)
# * ...
# * MongoDB - [`pymongo`](http://api.mongodb.org/python/current/)
# * HBase - [`happybase`](http://happybase.readthedocs.org/en/latest/)
# * Spark - [`pyspark`](http://spark.apache.org/docs/latest/api/python/)
# * SSH - [`paramiko`](http://www.paramiko.org/)
# * HDFS - [`pywebhdfs`](https://pypi.python.org/pypi/pywebhdfs)
# * Amazon S3 - [`boto`](https://boto.readthedocs.org/en/latest/)
#
# Today we're going to use some of these indirectly with `odo` (was `into`) and Blaze. We'll try to point out these libraries as we automate them so that, if you'd like, you can use them independently.
#
# <hr />
# <img src="images/continuum_analytics_logo.png"
# alt="Continuum Logo",
# align="right",
# width="30%">,
#
# `odo` (formerly `into`)
# =======================
#
# Odo migrates data between formats and locations.
#
# Before we can use a database we need to move data into it. The `odo` project provides a single consistent interface to move data between formats and between locations.
#
# We'll start with local data and eventually move out to remote data.
#
# [*odo docs*](http://odo.readthedocs.org/en/latest/index.html)
#
#
# <hr/>
# ### Examples
#
#
# Odo moves data into a target from a source
#
# ```python
# >>> odo(source, target)
# ```
#
# The target and source can be either a Python object or a string URI. The following are all valid calls to `into`
#
# ```python
# >>> odo('iris.csv', pd.DataFrame) # Load CSV file into new DataFrame
# >>> odo(my_df, 'iris.json') # Write DataFrame into JSON file
# >>> odo('iris.csv', 'iris.json') # Migrate data from CSV to JSON
# ```
#
# <hr/>
# ### Exercise
#
# Use `odo` to load the `iris.csv` file into a Python `list`, a `np.ndarray`, and a `pd.DataFrame`
from odo import odo
import numpy as np
import pandas as pd
odo("iris.csv", pd.DataFrame)
# <hr/>
#
#
# URI Strings
# -----------
#
# Odo refers to foreign data either with a Python object like a `sqlalchemy.Table` object for a SQL table, or with a string URI, like `postgresql://hostname::tablename`.
#
# URI's often take on the following form
#
# protocol://path-to-resource::path-within-resource
#
# Where `path-to-resource` might point to a file, a database hostname, etc. while `path-within-resource` might refer to a datapath or table name. Note the two main separators
#
# * `://` separates the protocol on the left (`sqlite`, `mongodb`, `ssh`, `hdfs`, `hive`, ...)
# * `::` separates the path within the database on the right (e.g. tablename)
#
# [*odo docs on uri strings*](http://odo.readthedocs.org/en/latest/uri.html)
#
# <hr/>
# ### Examples
#
# Here are some example URIs
#
# ```
# myfile.json
# myfiles.*.csv'
# postgresql://hostname::tablename
# mongodb://hostname/db::collection
# ssh://user@host:/path/to/myfile.csv
# hdfs://user@host:/path/to/*.csv
# ```
#
# <hr />
# ### Exercise
#
# Migrate your CSV file into a table named `iris` in a new SQLite database at `sqlite:///my.db`. Remember to use the `::` separator and to separate your database name from your table name.
#
# [*odo docs on SQL*](http://odo.readthedocs.org/en/latest/sql.html)
odo("iris.csv", "sqlite:///my.db::iris")
# What kind of object did you get receive as output? Call `type` on your result.
type(_)
# <hr/>
#
# How it works
# ------------
#
# Odo is a network of fast pairwise conversions between pairs of formats. We when we migrate between two formats we traverse a path of pairwise conversions.
#
# We visualize that network below:
#
# 
#
# Each node represents a data format. Each directed edge represents a function to transform data between two formats. A single call to into may traverse multiple edges and multiple intermediate formats. Red nodes support larger-than-memory data.
#
# A single call to into may traverse several intermediate formats calling on several conversion functions. For example, we when migrate a CSV file to a Mongo database we might take the following route:
#
# * Load in to a `DataFrame` (`pandas.read_csv`)
# * Convert to `np.recarray` (`DataFrame.to_records`)
# * Then to a Python `Iterator` (`np.ndarray.tolist`)
# * Finally to Mongo (`pymongo.Collection.insert`)
#
# Alternatively we could write a special function that uses MongoDB's native CSV
# loader and shortcut this entire process with a direct edge `CSV -> Mongo`.
#
# These functions are chosen because they are fast, often far faster than converting through a central serialization format.
#
# This picture is actually from an older version of `odo`, when the graph was still small enough to visualize pleasantly. See [*odo docs*](http://odo.readthedocs.org/en/latest/overview.html) for a more updated version.
#
# <hr/>
# Remote Data
# -----------
#
# We can interact with remote data in three locations
#
# 1. On Amazon's S3 (this will be quick)
# 2. On a remote machine via `ssh`
# 3. On the Hadoop File System (HDFS)
#
# For most of this we'll wait until we've seen Blaze, briefly we'll use S3.
#
# ### S3
#
# For now, we quickly grab a file from Amazon's `S3`.
#
# This example depends on [`boto`](https://boto.readthedocs.org/en/latest/) to interact with S3.
#
# conda install boto
#
# [*odo docs on aws*](http://odo.readthedocs.org/en/latest/aws.html)
odo('s3://nyqpug/tips.csv', pd.DataFrame)
# <hr/>
#
# <img src="images/continuum_analytics_logo.png"
# alt="Continuum Logo",
# align="right",
# width="30%">,
#
# Blaze
# =====
#
# Blaze translates a subset of numpy/pandas syntax into database queries. It hides away the database.
#
# On simple datasets, like CSV files, Blaze acts like Pandas with slightly different syntax. In this case Blaze is just using Pandas.
# <hr/>
#
# ### Pandas example
# +
import pandas as pd
df = pd.read_csv('iris.csv')
df.head(5)
# -
df.Species.unique()
df.Species.drop_duplicates()
# <hr/>
#
# ### Blaze example
# +
import blaze as bz
d = bz.Data('iris.csv')
d.head(5)
# -
d.Species.distinct()
# <hr/>
#
# Foreign Data
# ------------
#
# Blaze does different things under-the-hood on different kinds of data
#
# * CSV files: Pandas DataFrames (or iterators of DataFrames)
# * SQL tables: [SQLAlchemy](http://sqlalchemy.org).
# * Mongo collections: [PyMongo](http://api.mongodb.org/python/current/)
# * ...
#
# SQL
# ---
#
# We'll play with SQL a lot during this tutorial. Blaze translates your query to SQLAlchemy. SQLAlchemy then translates to the SQL dialect of your database, your database then executes that query intelligently.
#
# * Blaze $\rightarrow$ SQLAlchemy $\rightarrow$ SQL $\rightarrow$ Database computation
#
# This translation process lets analysts interact with a familiar interface while leveraging a potentially powerful database.
#
# To keep things local we'll use SQLite, but this works with any database with a SQLAlchemy dialect. Examples in this section use the iris dataset. Exercises use the Lahman Baseball statistics database, year 2013.
#
# If you have not downloaded this dataset you could do so here - https://github.com/jknecht/baseball-archive-sqlite/raw/master/lahman2013.sqlite.
#
# <hr/>
# !ls
# ### Examples
#
# Lets dive into Blaze Syntax. For simple queries it looks and feels similar to Pandas
db = bz.Data('sqlite:///my.db')
#db.iris
#db.iris.head()
db.iris.Species.distinct()
db.iris[db.iris.Species == 'versicolor'][['Species', 'SepalLength']]
# <hr />
#
# ### Work happens on the database
#
# If we were using pandas we would read the table into pandas, then use pandas' fast in-memory algorithms for computation. Here we translate your query into SQL and then send that query to the database to do the work.
#
# * Pandas $\leftarrow_\textrm{data}$ SQL, then Pandas computes
# * Blaze $\rightarrow_\textrm{query}$ SQL, then database computes
#
# If we want to dive into the internal API we can inspect the query that Blaze transmits.
#
# <hr />
# Inspect SQL query
query = db.iris[db.iris.Species == 'versicolor'][['Species', 'SepalLength']]
print bz.compute(query)
query = bz.by(db.iris.Species, longest=db.iris.PetalLength.max(),
shortest=db.iris.PetalLength.min())
print bz.compute(query)
odo(query, list)
# <hr />
#
# ### Exercises
#
# Now we load the Lahman baseball database and perform similar queries
# db = bz.Data('postgresql://postgres:postgres@ec2-54-159-160-163.compute-1.amazonaws.com') # Use Postgres if you don't have the sqlite file
db = bz.Data('sqlite:///lahman2013.sqlite')
db.dshape
# View the Salaries table
# What are the distinct teamIDs in the Salaries table?
# What is the minimum and maximum yearID in the Sarlaries table?
# For the Oakland Athletics (teamID OAK), pick out the playerID, salary, and yearID columns
# Sort that result by salary.
# Use the ascending=False keyword argument to the sort function to find the highest paid players
# <hr />
#
# ### Example: Split-apply-combine
#
# In Pandas we perform computations on a *per-group* basis with the `groupby` operator. In Blaze our syntax is slightly different, using instead the `by` function.
import pandas as pd
iris = pd.read_csv('iris.csv')
iris.groupby('Species').PetalLength.min()
iris = bz.Data('sqlite:///my.db::iris')
bz.by(iris.Species, largest=iris.PetalLength.max(),
smallest=iris.PetalLength.min())
print(_)
# <hr/>
#
# Store Results
# -------------
#
# By default Blaze only shows us the first ten lines of a result. This provides a more interactive feel and stops us from accidentally crushing our system. Sometimes we do want to compute all of the results and store them someplace.
#
# Blaze expressions are valid sources for `odo`. So we can store our results in any format.
# +
iris = bz.Data('sqlite:///my.db::iris')
query = bz.by(iris.Species, largest=iris.PetalLength.max(), # A lazily evaluated result
smallest=iris.PetalLength.min())
odo(query, list) # A concrete result
# -
# <hr/>
#
# ### Exercise: Storage
#
# The solution to the first split-apply-combine problem is below. Store that result in a list, a CSV file, and in a new SQL table in our database (use a uri like `sqlite://...` to specify the SQL table.)
result = bz.by(db.Salaries.teamID, avg=db.Salaries.salary.mean(),
max=db.Salaries.salary.max(),
ratio=db.Salaries.salary.max() / db.Salaries.salary.min()
).sort('ratio', ascending=False)
odo(result, list)[:10]
| Section_1_blaze.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Statistical terms
#
#
# **Probability** - the expected relative frequency of a particular outcome
#
# P(heads) = 0.5 --> discrete distribution
# P(height>69) = 0.3 --> contious distribution
#
# **Random variable** (RV) - variable determined by random experiment
#
# P(H>h) --> probability that height is larger than some observed height h
#
# **Probability distribution, f(h)**
# - describes the distribution of a random variable
# - area under pdf fives probability
#
#
# What distribution do we use?
# * typically assume normal (Gaussian)
# * functions of normals give other popular distributions
# - Chisquare is the square of a normal
# - T involves a normal and a chisquares
# - F is the ratio of two chisquares
#
# **Expected value** - the mean of a random variable (E[Y])
#
# **Variance** - how the values of the RV are dispersed about the mean
# **Covariance** - how much 2RV's vary together
# - $Cov[X,Y] = E[(X - E[X])(Y - E[Y])]$
# - if two RC's are independent, $Cov[x, Y] = 0$ BUT the oposite is not true
#
# **Bias** and **variance** can be used to assess an estimator
# - **Estimator** - equation that estimates a parameter for you
# - **Bias**: on average, the estimate is correct
# - **Variance**: the reliability of the estimate
#
# How do we quantify the quality of an estimator?
# ## Simple linear regression
#
# Introduction without matrix algebra.
#
# Note:
# - I'll stick with the terminology of *association* when describing regression results.
# - *causality* is definitely a different type of approach
# - Some will debate whether linear regression results can be interpreted as *predictive*
#
# Models helps us tell stories! (correlation of age with reaction time)
#
# For the ith observation unit
#
# $Y_{i} = \beta_{0} + \beta_{1} X_{i} + \epsilon_{i}$
#
# - $Y_{i}$: the dependent (random) variable
# - $X_{i}$: independent variable (not random)
# - $\beta_{0}, \beta_{1}$: model parameters
# - $\epsilon_{i}$: Random error, how the observation deviates from the population mean
#
# **Fixed**: $\beta_{0} + \beta_{1} X_{i}$
# - Mean of $Y_{i}, (E[Y_{i}])$
#
# **Random**: $\epsilon_{i}$ --> this is where the variance is desctibed
# - Variability of $Y_{i}$
# - $E(\epsilon_{i}) = 0$ --> variance has a mean of 0
# - $Var(\epsilon_{i} = \sigma^{2}$ --> variance is the same for each subject
# - $Cov(\epsilon_{i}, \epsilon_{j})=0$ --> subjects are not correlated with each other
# - it folows that the variance of $Y_i$ is $\sigma^2$
#
# ## How do we fit the model?
#
# Which line fits the data best?
#
# - Minimize the distance between the data and the line (error)
# - Absolute distance? squared distance?
# - $\epsilon_{i} = Y_i - (\beta_0+\beta_{1}X_{1})$
#
#
# ### Least Squares
# - minimize squared differences
# - minimize $\sum^{N}_{i=1}\epsilon_{i}^2 = \sum^{N}_{i=1}(Y_{i} - Y_{esti}^2)$
# - works out nicely distribution-wise
# - you cna use calculus to get the estimates
#
#
# ### Property of least squares
#
# Ga<NAME>
# 1. Assumptions
# - error has mean 0
# - things aren't correlated (homoscedasticity)
# - variance is the same for all observations
#
# 2. *Unbiased* and have *lowest variance* among all unbiased estimators
# 3. Best Linear Unbiased Estimator (BLUE)
#
# We don't need normality --> we need it only when estimating p-values
#
#
# ### What about the variance?
# - we also need an estimate for $\sigma^2$
# - start with the sim of squared error
# - $SSE = \sum(Y_i - Y_esti)^2 = \sum e_{i}^2$
# - divide by the appropriate degrees of freedom: # of independent pieces of information - # parameters in the model
#
# $\sigma^2 = \frac{\sum e_{i}^2}{N-2}$
#
# ### Multiple Linear Regression
# - Add more parameters to the model
#
# $Y_{i} = \beta_{0} + \beta_{1} X_{i} + \beta_{2} X_{i} + \beta_{3} X_{i} + \epsilon_{i}$
#
# - Time for linear algebra!
#
#
# ## Important stuff:
#
# - What parameters in the regression are considered "fixed"?
# - What parameters are considered "random"?
# - How do we define the "best" line?
# - How many parameters do you end up estimating in a simple linear regression
# - What is residual?
| 04-general_linear_model/linear_regression_notes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#import important libraries
import pandas as pd
import numpy as np
import seaborn as sns
from sklearn.metrics import classification_report
from sklearn.metrics import roc_auc_score as roc
from sklearn.model_selection import train_test_split as tts
from sklearn.linear_model import LogisticRegression
from sklearn.preprocessing import StandardScaler
#import the dataset and visualize the dataset
dataset = pd.read_csv('./dataset/creditcard.csv')
dataset.head()
#visualize the target class 'Class'
sns.countplot(x='Class', data=dataset)
#seperating positive and negative classes
positiveDataset = dataset.loc[dataset['Class'] == 1]
negativeDataset = dataset.loc[dataset['Class'] == 0]
#creating training and testing set with negative class split 1:1 and positive class split 4:1, also keeping random_state constant so that all splits are same
positiveTrain, positiveTest = tts(positiveDataset, test_size=0.2, random_state=21)
negativeTrain, negativeTest = tts(negativeDataset, test_size=0.5)
trainDataset = positiveTrain.append(negativeTrain)
testDataset = positiveTest.append(negativeTest)
#create Regression object and scale the dataset
classifier = LogisticRegression(random_state=21)
yTrain = trainDataset['Class']
yTest = testDataset['Class']
xTrain = trainDataset.drop(columns=['Class'])
xTest = testDataset.drop(columns=['Class'])
scaler = StandardScaler()
xTrain = scaler.fit_transform(xTrain)
xTest = scaler.fit_transform(xTest)
#fit the dataset to the train values
classifier.fit(xTrain, yTrain)
#predict the model on the train values and check results
predTrain = classifier.predict(xTrain)
print(classification_report(yTrain, predTrain))
print('ROC AUC Score: ',roc(yTrain, predTrain))
#predict test values and check results
predTest = classifier.predict(xTest)
print(classification_report(yTest, predTest))
print('ROC AUC Score: ',roc(yTest, predTest))
| Logistic Regression Classifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + deletable=true editable=true
import tensorflow as tf
config = tf.ConfigProto()
config.gpu_options.allow_growth = True
import scipy
from corpus import Corpus
import numpy as np
# + deletable=true editable=true
corp_path='/home/velkey/corp/webkorpusz.wpl'
corp=Corpus(corpus_path=corp_path,language="Hun",size=1000,encoding_len=10)
# + deletable=true editable=true
all_features=corp.featurize_data_charlevel_onehot(corp.hun_lower)
train=all_features[0:int(len(all_features)*0.8)]
test=all_features[int(len(all_features)*0.8):len(all_features)]
# + deletable=true editable=true
x_train = train.reshape((len(train), np.prod(train.shape[1:])))
x_test = test.reshape((len(test), np.prod(test.shape[1:])))
print(x_train.shape)
# + deletable=true editable=true
import random
import matplotlib.pyplot as plt
class Experiment:
def __init__(self,x_train,x_test,y_train,y_test,layer_intervals,encoder_index,optimizer,lossmethod,step_size=0):
self.layernum=len(layer_intervals)
self.layer_intervals=layer_intervals
self.encoder_index=encoder_index
self.optimizer=optimizer
self.lossmethod=loss
self.tried_list=[]
self.train_losses=[]
self.test_losses=[]
self.x_train=x_train
self.y_train=y_train
self.train_len=len(x_train)
self.test_len=len(x_test)
self.x_test=x_test
self.y_test=y_test
self.data_dim=x_train[0].shape[0]*x_train[0].shape[1]
def gen_model(self,layer_data,type):
"""
@layer_data: [[size,activation],[size,activation]] with the last layer
"""
def run(self):
"""
"""
def show_words(predict_base,num=30):
encoded_text=encoder.predict(predict_base)
decoded_text = decoder.predict(encoded_text)
for i in range(num):
x=random.randint(0,len(predict_base)-1)
print("original:\t",corp.defeaturize_data_charlevel_onehot([predict_base[x].reshape(10,36)]),\
"\tdecoded:\t",corp.defeaturize_data_charlevel_onehot([decoded_text[x].reshape(10,36)]))
def plot_words_as_img():
encoded_imgs=encoder.predict(x_train)
decoded_imgs = decoder.predict(encoded_imgs)
n = 6 # how many digits we will display
plt.figure(figsize=(21, 4))
for i in range(n):
# display original
ax = plt.subplot(2, n, i + 1)
plt.imshow(x_test[i].reshape(10, 36))
plt.gray()
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
# display reconstruction
ax = plt.subplot(2, n, i + 1 + n)
plt.imshow(decoded_imgs[i].reshape(10,36))
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
plt.show()
# + deletable=true editable=true
def xavier_init(fan_in, fan_out, constant = 1):
low = -constant * np.sqrt(6.0 / (fan_in + fan_out))
high = constant * np.sqrt(6.0 / (fan_in + fan_out))
return tf.random_uniform((fan_in, fan_out),
minval = low, maxval = high,
dtype = tf.float32)
# + deletable=true editable=true
class Autoencoder_ffnn():
def __init__(self, featurelen,length,layerlist,encode_index,optimizer = tf.train.AdamOptimizer()):
"""
"""
self.layerlist=layerlist
self.layernum=len(layerlist)
self.n_input = featurelen*length
self.encode_index=encode_index
network_weights = self._initialize_weights()
self.weights = network_weights
self._create_layers()
# cost
self.cost = 0.5*tf.reduce_sum(tf.pow(tf.subtract(self.reconstruction, self.x), 2.0))
self.optimizer = optimizer.minimize(self.cost)
init = tf.global_variables_initializer()
self.sess = tf.Session(config=config)
self.sess.run(init)
def _initialize_weights(self):
all_weights = dict()
all_weights['w'+str(1)]=tf.Variable(xavier_init(self.n_input, self.layerlist[0][0]))
all_weights['b'+str(1)] = tf.Variable(tf.random_normal([self.layerlist[0][0]], dtype=tf.float32))
for i in range(1,self.layernum):
all_weights['w'+str(i+1)]=tf.Variable(xavier_init(self.layerlist[i-1][0], self.layerlist[i][0]))
all_weights['b'+str(i+1)] = tf.Variable(tf.random_normal([self.layerlist[i][0]], dtype=tf.float32))
return all_weights
def _create_layers(self):
"""
"""
self.x = tf.placeholder(tf.float32, [None, self.n_input])
layer=(self.layerlist[0][1])(tf.add(tf.matmul(self.x, self.weights['w1']), self.weights['b1']))
for i in range(1,self.layernum):
layer=(self.layerlist[i][1])(tf.add(tf.matmul(layer, self.weights['w'+str(i+1)]), self.weights['b'+str(i+1)]))
if i==self.encode_index:
print("enc")
self.encoded=layer
self.reconstruction=layer
def partial_fit(self, X):
cost, opt = self.sess.run((self.cost, self.optimizer), feed_dict={self.x: X})
return cost
def calc_total_cost(self, X):
return self.sess.run(self.cost, feed_dict = {self.x: X})
def encode(self, X):
return self.sess.run(self.encoded, feed_dict={self.x: X})
def decode(self, encoded = None):
if encoded is None:
encoded = np.random.normal(size=self.weights["b1"])
return self.sess.run(self.reconstruction, feed_dict={self.encoded: encoded})
def reconstruct(self, X):
return self.sess.run(self.reconstruction, feed_dict={self.x: X})
def train(self,X_train,X_test,batch_size,max_epochs):
for epoch in range(training_epochs):
avg_cost = 0.
total_batch = int(len(X_train) / batch_size)
# Loop over all batches
for i in range(total_batch):
batch_xs = self.get_random_block_from_data(X_train, batch_size)
cost = autoencoder.partial_fit(batch_xs)
avg_cost += cost / batch_size
# Display logs per epoch step
if epoch % display_step == 0:
print ("Epoch:", '%04d' % (epoch + 1), \
"cost=", "{:.9f}".format(avg_cost))
def get_random_block_from_data(self,data, batch_size):
start_index = np.random.randint(0, len(data) - batch_size)
return data[start_index:(start_index + batch_size)]
# + deletable=true editable=true
training_epochs = 40
batch_size = 1280
def ekv(e):
return e
display_step = 1
a=[[360,tf.nn.softplus],[360,ekv]]
autoencoder = Autoencoder_ffnn(10,36,
layerlist=a,
encode_index=1,
optimizer = tf.train.AdamOptimizer(learning_rate = 0.001))
autoencoder.train(x_train,x_test,512,10)
print ("Total cost: " + str(autoencoder.calc_total_cost(x_test)))
# + deletable=true editable=true
# + deletable=true editable=true
# + deletable=true editable=true
| old/autoencoder_experiment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lecture 6.2: Naive Bayes & Decision Trees
#
# This lecture, we are going to train and compare a Gaussian naive bayes model and a decision tree on a real dataset.
#
# **Learning goals:**
# - train a Gaussian naive bayes model
# - train a decision tree classifier
# - visualize and compare the model decision boundaries
# - analyse the effect of regularization parameters `max_depth` & `min_samples_leaf`
# ## 1. Introduction
#
# Let's try to improve our fake banknote detector from lecture 6.1. 🕵️♀️ We'll use the same [banknote authentication dataset](https://archive.ics.uci.edu/ml/datasets/banknote+authentication), and try to solve the fake/genuine classification task.
# ## 2. Classification
#
# ### 2.1 Data Munging
#
# Let's load our `.csv` into a pandas `DataFrame`, and have a look at the dataset:
# +
import pandas as pd
df = pd.read_csv('bank_note.csv')
df.head()
# -
df.describe()
# Recall that we are dealing with 4 features, and one binary label. The features are standardized, so no further preprocessing is necessary.
#
# We can create our feature matrix, `X`, and our label vector, `y`:
X = df[['feature_2', 'feature_4']].values
y = df['is_fake'].values
# And we can visualize the dataset to remember the complexity of the classification task:
# +
import matplotlib.pyplot as plt
fig = plt.figure(figsize=(5,5), dpi=120)
ax = fig.add_subplot()
scatter = ax.scatter(X[:, 0], X[:, 1], c=y, cmap=plt.cm.coolwarm, s=20, edgecolors='k', alpha=0.5)
ax.set_xlabel('feature_2')
ax.set_ylabel('feature_4')
ax.set_title('Banknote Classification')
handles, labels = scatter.legend_elements()
ax.legend(handles=handles, labels=['genuine', 'fake']);
# -
# The data is _not separable_ , and the relationship between `feature_2` and `feature_4` is _non-linear_. This should make a good challenge for our decision trees! 🌳
# ### 2.2 Training
#
# #### 2.2.1 Gaussian Naive Bayes
#
# We can use our favourite sklearn model api with `.fit` and `.predict()`. The class for Gaussian Naive Bayes is ... `GaussianNB` 😏
from sklearn.naive_bayes import GaussianNB
nb_clf = GaussianNB()
nb_clf = nb_clf.fit(X, y)
# 🧠 Can you list all the steps that sklearn had to go through to train this Gaussian Naive Bayes model with the function `.fit()`?
#
# Let's investigate our fitted _model parameters_.
# $ P(y|X) = \frac{P(y) \prod_{i} P(x_{i}|y)}{P(X)}$
#
# where:
#
# $P(x_{i}|y) \sim \mathcal{N}(\mu_{y},\,\sigma_{y}^{2})\,$,
#
# so that:
#
# $P(x_i \mid y) = \frac{1}{\sqrt{2\pi\sigma^2_y}} \exp\left(-\frac{(x_i - \mu_y)^2}{2\sigma^2_y}\right)$
# Remember that our Gaussian Naive Bayes model has learned a $\mu$ and a $\sigma$ independently for each feature, and for each class! In this case, we expect $2 x 2$ values.
#
# ⚠️ Sklearn makes things a little confusing (which is out of character), and names $\mu$ -> `theta_`
nb_clf.theta_
nb_clf.sigma_
# We can also access the class priors, $P(y)$:
nb_clf.class_prior_
# We stated earlier that the class priors are just the relative frequencies of the classes in the dataset... we can check this directly:
y.mean()
# Indeed the mean of `1` values in the dataset matches the $P(y=1)$!
#
# 🧠 These are all the model parameters that the Gaussian Naive Bayes model needed to learn from the dataset to be able to estimate $P(y|X)$. Make sure you understand how that works!
#
# Since we'll be visualizing a lot of classifications in 2D throughout this notebook, let's write some helper functions (code from the [sklearn documentation](https://scikit-learn.org/0.18/auto_examples/svm/plot_iris.html)).
#
# Just like last lecture, the function `.plot_classification()` plots both the dataset and the decision boundary for a given feature matrix `X`, label vector `y`, and a classifier `clf`:
# +
import numpy as np
from matplotlib.lines import Line2D
def make_meshgrid(x, y, h=.02):
"""Create a mesh of points to plot in
Parameters
----------
x: data to base x-axis meshgrid on
y: data to base y-axis meshgrid on
h: stepsize for meshgrid, optional
Returns
-------
xx, yy : ndarray
"""
x_min, x_max = x.min() - 1, x.max() + 1
y_min, y_max = y.min() - 1, y.max() + 1
xx, yy = np.meshgrid(np.arange(x_min, x_max, h),
np.arange(y_min, y_max, h))
return xx, yy
def plot_decision_boundary(ax, clf, xx, yy, **params):
"""Plot the decision boundaries for a classifier.
Parameters
----------
ax: matplotlib axes object
clf: a classifier
xx: meshgrid ndarray
yy: meshgrid ndarray
params: dictionary of params to pass to contourf, optional
"""
Z = clf.predict(np.c_[xx.ravel(), yy.ravel()])
Z = Z.reshape(xx.shape)
out = ax.contourf(xx, yy, Z, **params)
return out
def plot_contours(ax, clf, xx, yy, **params):
"""Plot the decision boundaries for a classifier.
Parameters
----------
ax: matplotlib axes object
clf: a classifier
xx: meshgrid ndarray
yy: meshgrid ndarray
params: dictionary of params to pass to contourf, optional
"""
plot_decision_boundary(ax, clf, xx, yy, **params)
def plot_classification(ax, X, y, clf):
X0, X1 = X[:, 0], X[:, 1]
xx, yy = make_meshgrid(X0, X1)
plot_contours(ax, clf, xx, yy,
cmap=plt.cm.coolwarm, alpha=0.8)
scatter = ax.scatter(X0, X1, c=y, cmap=plt.cm.coolwarm, s=20, edgecolors='k', alpha=1.0)
ax.set_xlim(xx.min(), xx.max())
ax.set_ylim(yy.min(), yy.max())
ax.set_xlabel('x1')
ax.set_ylabel('x2')
ax.set_title('Bank Notes Classification')
handles, labels = scatter.legend_elements()
ax.legend(handles=handles, labels=['genuine', 'fake'])
# -
# We can now easily plot our Gaussian Naive Bayes model's predictions 🎨:
fig = plt.figure(figsize=(5,5), dpi=120)
ax = fig.add_subplot()
plot_classification(ax, X, y, nb_clf)
# Notice the non-linear (quadratic) decision boundary. Does it help classification here? Can you visualise where the Gaussian Distribution means are located?
# ### 2.2 Training
#
# #### 2.2.1 Decision Trees
#
# We can use our favourite sklearn model api with `.fit` and `.predict()`. The class for decision tree classifiers is ... `DecisionTreeClassifier` 😏
from sklearn.tree import DecisionTreeClassifier
tree_clf = DecisionTreeClassifier(random_state=0)
tree_clf = tree_clf.fit(X, y)
# 🧠 Can you list all the steps that sklearn had to go through to train this decision tree with the function `.fit()`?
#
# We usually try to investigate our model parameters after fitting the model. However, decision trees don't have a vector $\theta$ or support vectors, they are _non-parametric models_. In the lecture slides, we introduced decision trees as nested if-statements. So instead, we can investigate their _decision node splits_.
#
# Let's check the _depth_ of our decision tree. Remember this is the maximal length of a decision _branch_. We can also output the total number of leaf nodes:
print(f'This decision tree has depth {tree_clf.get_depth()}, and contains {tree_clf.get_n_leaves()} leaves')
# 27 decision levels, that's quite a big tree we've grown here! 🌳
#
# Recall that decision trees make predictions by stepping through their decision nodes. By visualizing our tree's nodes, we can interpret its predictions. Decision trees are sometimes called white box models, because we can examine their inner workings.
#
# We'll use sklearn's `.plot_tree()` method to visualize the decision nodes. We must _truncate_ the tree with a `max_depth=2`, or else the visualization will be too big to fit on the screen.
# +
from sklearn.tree import plot_tree
fig = plt.figure(dpi=200)
ax = fig.add_subplot(111)
plot_tree(ax=ax, decision_tree=tree_clf, filled=True, max_depth=2, feature_names=['x1', 'x2'], rounded=True, precision=2);
# -
# This visualization is packed with information 🤤
#
# - The first line of each node defines its _split_. i.e The feature and the value which partitions incoming data into its children nodes.
#
# - The second line indicates the _gini impurity_ metric associated with each split on the training dataset. Remember, low gini impurity implies homogeneity and is therefore a good thing!
#
# - The third line displays the number of training examples belonging to that node.
#
# - The fourth line shows the _class split_ of this node. i.e How many genuine vs fake bills were present in this node during training. We expect this gap to get larger as we go deeper down the branches.
#
# - The node color represents the same information as the fourth line: bluer nodes contain more genuine bills, redder nodes contain more fake bills. We also expect these hues to get more pronounced closer to the leafs nodes.
#
# Note that all lines except the first tell information specific to _training_. All that is needed for prediction is the feature values of the splits, and the class attribution of each leaf node.
#
# 🧠 Take your time to understand this graph and how it was "greedily" built during training.
#
# These splits (if-statements) shape the model's decision boundary. We'd like to visualize this along with the dataset.
# We can now easily plot our decision tree classifier's predictions 🎨:
fig = plt.figure(figsize=(5,5), dpi=120)
ax = fig.add_subplot()
plot_classification(ax, X, y, tree_clf)
# That's a funky decision boundary! 😳 It is made exclusively of vertical and horizontal lines because predictions are made from a succession of if-statements on single features. The very thin rectangular areas are typical of _overfit_ decision trees. They try to fit single data points instead of the underlying patterns of the data.
# #### 2.2.3 Comparison
#
# Let's compare these two models with another type of classifier: a RBF kernel SVM (see lecture 6.1).
#
# Just like last lecture, we can write a small loop to visualize these models next to eachother. We'll then train a non-linear SVM to compare with the decision tree and the random forest:
# +
def compare_classification(X, y, clfs, titles):
fig = plt.figure(figsize=(14, 4), dpi=100)
for i, clf in enumerate(clfs):
ax = fig.add_subplot(1, len(clfs), i+1)
plot_classification(ax, X, y, clf)
ax.set_title(titles[i])
from sklearn.svm import SVC
svm_rbf = SVC(kernel='rbf', random_state=0)
svm_rbf = svm_rbf.fit(X, y)
compare_classification(X, y, [svm_rbf, nb_clf, tree_clf], ['RBF SVM', 'Naive Bayes', 'Decision Tree'])
# -
# All three models exhibit non-linear decision boundaries, but in very different ways.
# - the SVM is busy maximizing the margin shaped by its distance features, creating smooth "blobs" that follow the edges of the data
# - the naive bayes fits its quadratice decision boundary
# - the decision tree tries its best to split the dataset with single feature thresholds, splitting the data into nested rectangles
#
#
# 🧠🧠 None of these models seem to be able to correctly predict the fake examples near $[1, -1]$. Why is that?
#
# 🧠🧠 In your opinion, which is the algorithm most adapted to this dataset and classification task? Why?
# ### 2.3 Prediction
# Let's test our models by asking them to predict a banknote in the small `genuine` cluster on the left hand side of the graphs above. We'll use $feature\_1 = -1; feature\_2 = 0$:
# +
x_predict = np.array([-1, 0]).reshape(1, 2)
print(f'Features: {x_predict}')
svm_rbf_prediction = svm_rbf.predict(x_predict)
print(f'RBF SVM prediction: {svm_rbf_prediction}')
tree_clf_prediction = nb_clf.predict(x_predict)
print(f'Naive Bayes prediction: {tree_clf_prediction}')
forest_clf_prediction = tree_clf.predict(x_predict)
print(f'Decision Tree prediction: {forest_clf_prediction}')
# -
# ### 2.4 Analysis
#
# #### 2.4.1 Regularization: max depth
#
# We have successfully trained decision trees, but we haven't played with a different regularization hyperparameter: `max_depth`.
#
# `max_depth` "cuts" branches which are too long. i.e During training, nodes deeper than `max_depth` automatically become leaf nodes.
#
# Let's directly visualize the effect of this hyperparameter on the models's classification by plotting decision boundaries for different values of `max_depth`. We'll be using the handy `**kwargs` as arguments, here's a great [blog post](https://realpython.com/python-kwargs-and-args/) if you haven't heard about them.
# +
def train_tree(X, y, **kwargs):
tree_clf = DecisionTreeClassifier(random_state=0, **kwargs)
return tree_clf.fit(X, y)
max_depth_values = [2, 5, 20]
trees = [train_tree(X, y, max_depth=m) for m in max_depth_values]
titles = [f'max_depth={max_depth}' for max_depth in max_depth_values]
compare_classification(X, y, trees, titles)
# -
# We can clearly see the regularizing effect of "cutting off" trees branches after a certain depth. `max_depth` has for effect of reducting the number of nested if-statements, and therefore limiting the number of "angles" in the decision boundary.
# ## 4. Summary
#
# Today we learned about two new supervised learning models: **naive bayes** and **decision trees**. We started by defining how naive bayes uses Baye's theorem to reframe classification problems as the estimation of a conditional probability. We described the model's tricks, namely: assuming the independence of the features, and assuming that they are Gaussian distributed. We explained how this allows to create fast and simple classifiers only by estimating Gaussian parameters and class priors. We then described the **decision tree** algorithm, and showed how it makes non-linear predictions with nested if-statements. We then went through its training procedure which uses **homogeneity metrics** or **variance reduction** to optimally split decision nodes. After noting these models' tendency to **overfit**, we introduced a **regularization** procedures: changing the trees' maximum depth.
# Finally, we applied naive bayes and decision trees to the banknote classification dataset. We trained and visualized the models, as well as analysing the effect of regularization parameters.
#
# # Resources
#
# ## Core Resources
#
#
# - [sklearn naive bayes](https://scikit-learn.org/stable/modules/naive_bayes.html)
# - [sklearn decision trees](https://scikit-learn.org/stable/modules/tree.html)
# Official documentation about the tree package, handy breakdown of tree models in sklearn
#
#
# ### Additional Resources
#
# - [Understanding gini impurity](https://victorzhou.com/blog/gini-impurity/)
# Same blog from <NAME> going into the mathematics of gini impurity
# - [args and kwargs demystified](https://realpython.com/python-kwargs-and-args/)
# blog post about \*\*kwargs in python
#
| week_6/6.2_naive_bayes_decision_trees/naive_bayes_decision_trees.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Analyzing Quantum Volume Errors
# This notebook analyzes the error rates required for achieving Quantum Volume at a particular depths. For a given m = depth = number of qubits, plot the HOG for np.logspace outputs to view when it crosses the 2/3rds probability threshold.
# +
# Configuration parameters. Feel free to mess with these!
import cirq
num_repetitions = 10
depth = 4
num_samplers = 50
device = cirq.google.Bristlecone
compiler = lambda circuit: cirq.google.optimized_for_xmon(
circuit=circuit,
new_device=device)
print(f"Configuration: depth {depth} with "
f"{num_repetitions} runs of {num_samplers} samplers")
# +
# Run the Quantum Volume algorithm over the above parameters.
import numpy as np
from cirq.contrib import quantum_volume
errors = np.logspace(-1, -4, num=num_samplers)
samplers = [
cirq.DensityMatrixSimulator(noise=cirq.ConstantQubitNoiseModel(
qubit_noise_gate=cirq.DepolarizingChannel(p=error)))
for error in errors]
result = quantum_volume.calculate_quantum_volume(num_repetitions=num_repetitions,
depth=depth,
num_qubits=depth,
device=device,
samplers=samplers,
compiler=compiler,
seed=None)
# +
# Create a chart that plots the HOG rate relative to the simulated error ratio.
from matplotlib import pyplot as plt
fig, axs = plt.subplots()
sampler_results = [res.sampler_result for res in result]
axs.plot(errors,
[sum(res) / len(res) for res in np.transpose(sampler_results)])
#[sum(res) / len(res) for res in result.sampler_results.values()])
# Line markers for asymptotic ideal heavy output probability and the ideal Heavy
# Output Generation threshold.
axs.axhline((1 + np.log(2)) / 2,
color='tab:green',
label='Asymptotic ideal',
linestyle='dashed')
axs.axhline(2 / 3, label='HOG threshold', color='k', linestyle='dotted')
plt.xscale('log')
axs.set_ybound(0.4, 1)
axs.set_xlabel("error rate")
axs.set_ylabel("est. heavy output probability")
fig.suptitle(f'HOG probability by simulated error rate for d={depth}')
| examples/advanced/quantum_volume_errors.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Expectation Reflection + Least Absolute Deviations
#
# In the following, we demonstrate how to apply Least Absolute Deviations (LAD) for classification task such as medical diagnosis.
#
# We import the necessary packages to the Jupyter notebook:
# +
import numpy as np
import pandas as pd
from sklearn.model_selection import train_test_split,KFold
from sklearn.utils import shuffle
from sklearn.metrics import confusion_matrix,accuracy_score,precision_score,\
recall_score,roc_curve,auc
import expectation_reflection as ER
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.preprocessing import MinMaxScaler
from function import split_train_test,make_data_balance
# -
np.random.seed(1)
# First of all, the processed data are imported.
data_list = np.loadtxt('data_list.txt',dtype='str')
#data_list = ['29parkinson','30paradox2','31renal','32patientcare','33svr','34newt','35pcos']
print(data_list)
def read_data(data_id):
data_name = data_list[data_id]
print('data_name:',data_name)
Xy = np.loadtxt('../data/%s/data_processed.dat'%data_name)
X = Xy[:,:-1]
y = Xy[:,-1]
#print(np.unique(y,return_counts=True))
X,y = make_data_balance(X,y)
print(np.unique(y,return_counts=True))
X, y = shuffle(X, y, random_state=1)
X_train,X_test,y_train,y_test = train_test_split(X,y,test_size=0.5,random_state = 1)
sc = MinMaxScaler()
X_train = sc.fit_transform(X_train)
X_test = sc.transform(X_test)
return X_train,X_test,y_train,y_test
def measure_performance(X_train,X_test,y_train,y_test):
n = X_train.shape[1]
l2 = [0.0001,0.001,0.01,0.1,1.,10.,100.]
#l2 = [0.0001,0.001,0.01,0.1,1.,10.]
nl2 = len(l2)
# cross validation
kf = 4
kfold = KFold(n_splits=kf,shuffle=False,random_state=1)
h01 = np.zeros(kf)
w1 = np.zeros((kf,n))
cost1 = np.zeros(kf)
h0 = np.zeros(nl2)
w = np.zeros((nl2,n))
cost = np.zeros(nl2)
for il2 in range(len(l2)):
for i,(train_index,val_index) in enumerate(kfold.split(y_train)):
X_train1, X_val = X_train[train_index], X_train[val_index]
y_train1, y_val = y_train[train_index], y_train[val_index]
#h01[i],w1[i,:] = ER.fit(X_train1,y_train1,niter_max=100,l2=l2[il2])
h01[i],w1[i,:] = ER.fit_LAD(X_train1,y_train1,niter_max=100,l2=l2[il2])
y_val_pred,p_val_pred = ER.predict(X_val,h01[i],w1[i])
cost1[i] = ((p_val_pred - y_val)**2).mean()
h0[il2] = h01.mean(axis=0)
w[il2,:] = w1.mean(axis=0)
cost[il2] = cost1.mean()
# optimal value of l2:
il2_opt = np.argmin(cost)
print('optimal l2:',l2[il2_opt])
# performance:
y_test_pred,p_test_pred = ER.predict(X_test,h0[il2_opt],w[il2_opt,:])
fp,tp,thresholds = roc_curve(y_test, p_test_pred, drop_intermediate=False)
roc_auc = auc(fp,tp)
#print('AUC:', roc_auc)
acc = accuracy_score(y_test,y_test_pred)
#print('Accuracy:', acc)
precision = precision_score(y_test,y_test_pred)
#print('Precision:',precision)
recall = recall_score(y_test,y_test_pred)
#print('Recall:',recall)
return acc,roc_auc,precision,recall
n_data = len(data_list)
roc_auc = np.zeros(n_data) ; acc = np.zeros(n_data)
precision = np.zeros(n_data) ; recall = np.zeros(n_data)
for data_id in range(n_data):
X_train,X_test,y_train,y_test = read_data(data_id)
acc[data_id],roc_auc[data_id],precision[data_id],recall[data_id] =\
measure_performance(X_train,X_test,y_train,y_test)
print(data_id,acc[data_id],roc_auc[data_id])
# +
print('acc_mean:',acc.mean())
print('roc_mean:',roc_auc.mean())
print('precision:',precision.mean())
print('recall:',recall.mean())
# -
np.savetxt('ER_LAD_result_30sets.dat',(roc_auc,acc,precision,recall),fmt='%f')
| 20.01.1400_GridSearchCV/ER_LAD_30sets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# > Texto fornecido sob a Creative Commons Attribution license, CC-BY. Todo o código está disponível sob a FSF-approved BSD-3 license.<br>
# > (c) Original por <NAME>, <NAME> em 2017, traduzido por <NAME> em 2020.<br>
# > [@LorenaABarba](https://twitter.com/LorenaABarba) - [@fschuch](https://twitter.com/fschuch)
# 12 passos para Navier-Stokes
# ======
# ***
# Você deter ter completado os Passos [1](./01_Passo_1.ipynb) e [2](./02_Passo_2.ipynb) antes de continuar. Esse Jupyter notebook é uma continuação dos **12 passos para Navier-Stokes**, um módulo prático aplicado como um curso interativo de Dinâmica dos Fluidos Computacional (CFD, do Inglês *Computational Fluid Dynamics*), por [Prof. <NAME>](http://lorenabarba.com). Adaptado e traduzido para português por [<NAME>](https://fschuch.github.io/).
# Passo 3: Difusão Unidimensional
# -----
# ***
# O equação de difusão unidimensional é escrita como:
#
# $$\frac{\partial u(x,t)}{\partial t}= \nu \frac{\partial^2 u(x,t)}{\partial x^2}$$
#
# A primeira coisa que você deve notar é que, diferente das duas equações anteriores que estudamos, essa equação tem uma derivada de segunda ordem. Nós primeiro precisamos ver o que fazemos com isso!
# ### Discretizando $\frac{\partial ^2 u}{\partial x^2}$
# A derivada de segunda ordem pode ser representada geometricamente como a linha que tangencia a curva dada pela derivada primeira. Vamos discretizar a derivada de segunda ordem com um esquema de diferença centrada: uma combinação entre a diferença para frente e a diferença para trás. Considerando a expansão em séries de Taylor de $u_{i+1}$ e $u_{i-1}$ em torno de $u_i$:
#
# $$u_{i+1} = u_i + \Delta x \frac{\partial u}{\partial x}\bigg|_i + \frac{\Delta x^2}{2} \frac{\partial ^2 u}{\partial x^2}\bigg|_i + \frac{\Delta x^3}{3!} \frac{\partial ^3 u}{\partial x^3}\bigg|_i + O(\Delta x^4)$$
#
# $$u_{i-1} = u_i - \Delta x \frac{\partial u}{\partial x}\bigg|_i + \frac{\Delta x^2}{2} \frac{\partial ^2 u}{\partial x^2}\bigg|_i - \frac{\Delta x^3}{3!} \frac{\partial ^3 u}{\partial x^3}\bigg|_i + O(\Delta x^4)$$
#
# Se adicionarmos as duas expansões, você pode ver que os termos de derivada ímpar cancelam uns aos outros. Se desprezarmos qualquer termos de ordem $O(\Delta x^4)$ ou maior (e realmente, eles são muito pequenos), então podemos rearranjar a soma dessas duas expansões e isolar a derivada de segunda ordem.
#
# $$u_{i+1} + u_{i-1} = 2u_i+\Delta x^2 \frac{\partial ^2 u}{\partial x^2}\bigg|_i + O(\Delta x^4)$$
#
# Então rearranjamos e isolamos $\frac{\partial ^2 u}{\partial x^2}\bigg|_i$, resultando em:
#
# $$\frac{\partial ^2 u_i}{\partial x^2}=\frac{u_{i+1}-2u_{i}+u_{i-1}}{\Delta x^2} + O(\Delta x^2)$$
#
# ### De volta ao Passo 3
# Podemos escrever a versão discretizada da equação de difusão unidimensional como:
#
# $$\frac{u_{i}^{n+1}-u_{i}^{n}}{\Delta t}=\nu\frac{u_{i+1}^{n}-2u_{i}^{n}+u_{i-1}^{n}}{\Delta x^2}$$
#
# Assim como antes, perceba que temos uma condição inicial e que a única incógnita é $u_{i}^{n+1}$, então, podemos rearranjar a equação para isolar nossa incógnita:
#
# $$u_{i}^{n+1}=u_{i}^{n}+\frac{\nu\Delta t}{\Delta x^2}(u_{i+1}^{n}-2u_{i}^{n}+u_{i-1}^{n})$$
#
# A equação discretizada acima nos permite escrever o programa para avançar a solução no tempo. Mas precisamos de uma condição inicial (CI). Vamos continuar com a nossa favorita: a função chapéu (*hat function*). Então, usando $u_0 = 2$ onde $ 0,5 \leq x \leq 1 $, senão $u = 1$, no intervalo $0 \le x \le 2$. Estamos prontos para prosseguir aos números.
# + jupyter={"outputs_hidden": false}
import numpy #Carregamos nossa biblioteca preferida
from matplotlib import pyplot #E nos preparamos para produzir gráficos
# %matplotlib inline
x = numpy.linspace(0., 2., num = 41) #Coordenada espacial
nt = 20 #Número de passos de tempo que queremos calcular
nu = 0.3 #O valor para viscosidade
sigma = .2 #Sigma é um parâmetro, veremos mais detalhes em breve
nx = x.size
dx = x[1] - x[0]
dt = sigma * dx**2 / nu #dt é calculado em função de sigma, já saberemos o porquê
u = numpy.ones(nx) #Um arranjo numpy com nx elementos e todos iguais a 1
u[(0.5<=x) & (x<=1)] = 2 #Então definimos u = 2 entre 0,5 e 1, nossa CI
un = numpy.ones(nx) #Inicializar o arranjo temporário, para manter a solução no passo de tempo
for n in range(nt): #Laço temporal
un = u.copy() ##Cópia dos valores de u para un
for i in range(1, nx - 1):
u[i] = un[i] + nu * dt / dx**2 * (un[i+1] - 2 * un[i] + un[i-1])
pyplot.plot(x, u);
# -
# Pós experimentar mudar os parâmetros no código acima, consulte o material extra ou vá direto para o [Passo 4](./05_Passo_4.ipynb).
# + [markdown] colab_type="text" id="5ikOedw5wjl2"
# Material Complementar
# -----
# ***
# -
# Para uma explicação passo à passo sobre a discretização da equação de convecção linear com diferenças finitas (e também os passos seguintes, até o Passo 4), assista **Video Lesson 4** por Prof. Barba no YouTube.
# + jupyter={"outputs_hidden": false}
from IPython.display import YouTubeVideo
YouTubeVideo('y2WaK7_iMRI')
# + jupyter={"outputs_hidden": false}
from IPython.core.display import HTML
def css_styling():
styles = open("../styles/custom.css", "r").read()
return HTML(styles)
css_styling()
# -
# > A célula acima executa o estilo para esse notebook. Nós modificamos o estilo encontrado no GitHub de [CamDavidsonPilon](https://github.com/CamDavidsonPilon), [@Cmrn_DP](https://twitter.com/cmrn_dp).
| tarefas/04_Passo_3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import tensorflow as tf
# If there are multiple GPUs and we only want to use one/some, set the number in the visible device list.
os.environ["CUDA_DEVICE_ORDER"]="PCI_BUS_ID"
os.environ["CUDA_VISIBLE_DEVICES"]="0"
# This sets the GPU to allocate memory only as needed
physical_devices = tf.config.experimental.list_physical_devices('GPU')
if len(physical_devices) != 0:
tf.config.experimental.set_memory_growth(physical_devices[0], True)
# -
# ### **1. Loading the dataset**
# This assignment will focus on the CIFAR10 dataset. This is a collection of small images in 10 classes such as cars, cats, birds, etc. You can find more information here: https://www.cs.toronto.edu/~kriz/cifar.html. We start by loading and examining the data.
# +
import numpy as np
from tensorflow.keras.datasets import cifar10
(X_train, y_train), (X_test, y_test) = cifar10.load_data()
print("Shape of training data:")
print(X_train.shape)
print(y_train.shape)
print("Shape of test data:")
print(X_test.shape)
print(y_test.shape)
# -
# #### **<span style="color:red">Question 1:</span>**
# The shape of X_train and X_test has 4 values. What do each of these represent?
#
# Answer: The 5000 represents the number of pictures in the dataset (batchsize)
# 32, 32 is the dimensions of the picture with 32x32 pixels
# 3 is the number of color channels that each picture has, so red, green and blue color channels
# ##### **Plotting some images**
# This plots a random selection of images from each class. Rerun the cell to see a different selection.
# +
from Custom import PlotRandomFromEachClass
cifar_labels = ['airplane', 'automobile', 'bird', 'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck']
PlotRandomFromEachClass(X_train, y_train, 3, labels=cifar_labels)
# -
# ##### **Preparing the dataset**
# Just like the MNIST dataset we normalize the images to [0,1] and transform the class indices to one-hot encoded vectors.
# +
from tensorflow.keras.utils import to_categorical
# Transform label indices to one-hot encoded vectors
y_train_c = to_categorical(y_train, num_classes=10)
y_test_c = to_categorical(y_test , num_classes=10)
# Normalization of pixel values (to [0-1] range)
X_train = X_train.astype('float32') / 255
X_test = X_test.astype('float32') / 255
# -
# ### **2. Fully connected classifier**
# We will start by creating a fully connected classifier using the ```Dense``` layer. We give you the first layer that flattens the image features to a single vector. Add the remaining layers to the network.
#
# Consider what the size of the output must be and what activation function you should use in the output layer.
# +
from tensorflow.keras.optimizers import SGD
from tensorflow.keras.models import Model
from tensorflow.keras.layers import Input, Dense, Flatten
x_in = Input(shape=X_train.shape[1:])
x = Flatten()(x_in)
# === Add your code here ===
x = Dense(128, activation = 'relu')(x)
x = Dense(64, activation = 'relu')(x)
x = Dense(10, activation = 'softmax')(x)
# ==========================
model = Model(inputs=x_in, outputs=x)
# Now we build the model using Stochastic Gradient Descent with Nesterov momentum. We use accuracy as the metric.
sgd = SGD(lr=0.01, decay=1e-6, momentum=0.9, nesterov=True)
model.compile(optimizer=sgd, loss='categorical_crossentropy', metrics=['accuracy'])
model.summary(100)
# -
# ##### **Training the model**
# In order to show the differences between models in the first parts of the assignment, we will restrict the training to the following command using 15 epochs, batch size 32, and 20% validation data. From section 5 and forward you can change this as you please to increase the accuracy, but for now stick with this command.
history = model.fit(X_train,y_train_c, epochs=15, batch_size=32, verbose=1, validation_split=0.2)
# ##### **Evaluating the model**
# We use ```model.evaluate``` to get the loss and metric scores on the test data. To plot the results we give you a custom function that does the work for you.
# +
score = model.evaluate(X_test, y_test_c, batch_size=128, verbose=0)
for i in range(len(score)):
print("Test " + model.metrics_names[i] + " = %.3f" % score[i])
# +
from Custom import PlotModelEval
# Custom function for evaluating the model and plotting training history
PlotModelEval(model, history, X_test, y_test, cifar_labels)
# -
# #### **<span style="color:red">Question 2:</span>**
# Train a model that achieves above 45% accuracy on the test data. In the report, provide a (short) description of your model and show the evaluation image.
# #### **<span style="color:red">Question 3:</span>**
# Compare this model to the one you used for the MNIST dataset in the first assignment, in terms of size and test accuracy. Why do you think this dataset is much harder to classify than the MNIST handwritten digits?
# ### **3. CNN classifier**
# We will now move on to a network architecture that is more suited for this problem, the convolutional neural network. The new layers you will use are ```Conv2D``` and ```MaxPooling2D```, which you can find the documentation of here https://www.tensorflow.org/api_docs/python/tf/keras/layers/Conv2D and here https://www.tensorflow.org/api_docs/python/tf/keras/layers/MaxPool2D.
# ##### **Creating the CNN model**
#
# A common way to build convolutional neural networks is to create blocks of layers of the form **[convolution - activation - pooling]**, and then stack several of these block to create the full convolution stack. This is often followed by a fully connected network to create the output classes. Use this recipe to build a CNN that acheives at least 62% accuracy on the test data.
#
# *Side note. Although this is a common way to build CNNs, it is be no means the only or even best way. It is a good starting point, but later in part 5 you might want to explore other architectures to acheive even better performance.*
# +
from tensorflow.keras.layers import Conv2D, MaxPooling2D
x_in = Input(shape=X_train.shape[1:])
# === Add your code here ===
#conv1
conv2D_1=Conv2D(filters = 32,kernel_size = (3,3), padding = 'same', activation = 'relu')(x_in)
MaxPool2D_1 = MaxPooling2D(pool_size=(2,2), strides=(1,1), padding='same')(conv2D_1)
#conv2
conv2D_2=Conv2D(filters = 64,kernel_size = (3,3), padding = 'same', activation = 'relu')(MaxPool2D_1)
MaxPool2D_L1 = MaxPooling2D(pool_size=(2,2), strides=(1,1), padding='same')(conv2D_2)
Conv2D_3=Conv2D(filters = 128,kernel_size = (3,3), padding = 'same', activation = 'relu')(MaxPool2D_L1)
MaxP_3 = MaxPooling2D(pool_size=(2,2), strides=(1,1), padding='same')(Conv2D_3)
# copied from prev q
x = Flatten()(MaxP_3)
# === dd your code here ===
x = Dense(128, activation = 'relu')(x)
x = Dense(64, activation = 'relu')(x)
x = Dense(10, activation = 'softmax')(x)
# ==========================
model = Model(inputs=x_in, outputs=x)
sgd = SGD(lr=0.01, decay=1e-6, momentum=0.9, nesterov=True)
model.compile(loss='categorical_crossentropy', metrics=['accuracy'], optimizer=sgd)
model.summary(100)
# -
# ##### **Training the CNN**
history = model.fit(X_train, y_train_c, batch_size=32, epochs=15, verbose=1, validation_split=0.2)
# ##### **Evaluating the CNN**
# +
score = model.evaluate(X_test, y_test_c, batch_size=128, verbose=0)
for i in range(len(score)):
print("Test " + model.metrics_names[i] + " = %.3f" % score[i])
# -
PlotModelEval(model, history, X_test, y_test, cifar_labels)
# #### **<span style="color:red">Question 4:</span>**
# Train a model that achieves at least 62% test accuracy. In the report, provide a (short) description of your model and show the evaluation image.
#
# ANSWER:
#
# The neural network contains three convolutional layers with the activation function relu, the kernel size used for convolution is a (3x3) matrix. Each convoultion layer is followed by a pooling function where the pooling is set to a "window" of (2x2) matrix. Where it will take the highest pixel-value in each "window". The strides or stepsize of which the window moves is (1x1). The filters of each of the convultions goes from simple to more complex with the values: 32,64, 128.
#
# Then the output is flattened into a single vector and used as input into the fully connected classifier with two layer network with relu as activation and 64 and 32 neurons each. The output layer uses softmax activation and has 10 neurons which corresponds to the number of classes.
# #### **<span style="color:red">Question 5:</span>**
# Compare this model with the previous fully connected model. You should find that this one is much more efficient, i.e. achieves higher accuracy with fewer parameters. Explain in your own words how this is possible.
#
# The combination of convolution - activation - pooling is can be seen as the feature learning phase of the neural network. Where network tries to learn the primitive image features as edges and lines of the dataset which will provide as kind of preprocessing for the neural network to improve the input into the fully connected network. The convoulution layer detects particular features depending on the kernel used. These multiple features are pooled together in the maxpooling phase which combines the maximum pixel values in the defined pooling window (in our case, window is a 2X2 matrix). Because of this preprocessing arrangement we get more abstract features as input to the fully connected classifier, hence we see a better result.
# ### **4. Regularization**
# #### **4.1 Dropout**
# You have probably seen that your CNN model overfits the training data. One way to prevent this is to add ```Dropout``` layers to the model, that randomly "drops" hidden nodes each training-iteration by setting their output to zero. Thus the model cannot rely on a small set of very good hidden features, but must instead learns to use different sets of hidden features each time. Dropout layers are usually added after the pooling layers in the convolution part of the model, or after activations in the fully connected part of the model.
#
# *Side note. In the next assignment you will work with Ensemble models, a way to use the output from several individual models to achieve higher performance than each model can achieve on its own. One way to interpret Dropout is that each random selection of nodes is a separate model that is trained only on the current iteration. The final output is then the average of outputs from all the individual models. In other words, Dropout can be seen as a way to build ensembling directly into the network, without having to train several models explicitly.*
#
# Extend your previous model with the Dropout layer and test the new performance.
# +
from tensorflow.keras.layers import Dropout
x_in = Input(shape=X_train.shape[1:])
# === Add your code here ===
Conv2D_1=Conv2D(filters = 32,kernel_size = (3,3), padding = 'same', activation = 'relu')(x_in)
MaxP_1 = MaxPooling2D(pool_size=(2,2), strides=(1,1), padding='same')(Conv2D_1)
Drop_1=Dropout(0.3)(MaxP_1)
#conv2
Conv2D_2=Conv2D(filters = 64,kernel_size = (3,3), padding = 'same', activation = 'relu')(Drop_1)
MaxP_2 = MaxPooling2D(pool_size=(2,2), strides=(1,1), padding='same')(Conv2D_2)
Drop_2=Dropout(0.3)(MaxP_2)
Conv2D_3=Conv2D(filters = 128,kernel_size = (3,3), padding = 'same', activation = 'relu')(Drop_2)
MaxP_3 = MaxPooling2D(pool_size=(2,2), strides=(1,1), padding='same')(Conv2D_2)
Drop_3=Dropout(0.3)(MaxP_3)
# copied from prev q
x = Flatten()(Drop_3)
# === dd your code here ===
x = Dense(128, activation = 'relu')(x)
x=Dropout(0.3)(x)
x = Dense(64, activation = 'relu')(x)
x=Dropout(0.3)(x)
x = Dense(10, activation = 'softmax')(x)
# ==========================
model = Model(inputs=x_in, outputs=x)
# Compile model
sgd = SGD(lr=0.01, decay=1e-6, momentum=0.9, nesterov=True)
model.compile(loss='categorical_crossentropy', metrics=['accuracy'], optimizer=sgd)
model.summary(100)
# -
history = model.fit(X_train, y_train_c, batch_size=32, epochs=15, verbose=1, validation_split=0.2)
# +
score = model.evaluate(X_test, y_test_c, batch_size=128, verbose=0)
for i in range(len(score)):
print("Test " + model.metrics_names[i] + " = %.3f" % score[i])
# -
PlotModelEval(model, history, X_test, y_test, cifar_labels)
# #### **<span style="color:red">Question 6:</span>**
# Train the modified CNN-model. Save the evaluation image for the report.
# #### **<span style="color:red">Question 7:</span>**
# Compare this model and the previous in terms of the training accuracy, validation accuracy, and test accuracy. Explain the similarities and differences (remember that the only difference between the models should be the addition of Dropout layers).
#
# Hint: what does the dropout layer do at test time?
#
# | | training | validation | test |
# | -------- | -------- | ---------- | ----- |
# | CNN | 0.9388 | 0.6414 | 0.632 |
# | CNN-drop | 0.6087 | 0.5933 | 0.592 |
#
#
# The training accuracy is much higher in the CNN without drop compared to the one with. Using drop training validation and test have a similar accuracy. The difference in training to validation and test accuracy for the CNN without drop is because the model overfits the data for the training because a lack of regularization. By introducing the drop functionality the model generalizes better as and becomes less sensitive to the data, hence does not overfit.
# #### **4.2 Batch normalization**
# The final layer we will explore is ```BatchNormalization```. As the name suggests, this layer normalizes the data in each batch to have a specific mean and standard deviation, which is learned during training. The reason for this is quite complicated (and still debated among the experts), but suffice to say that it helps the optimization converge faster which means we get higher performance in fewer epochs. The normalization is done separatly for each feature, i.e. the statistics are calculated accross the batch dimension of the input data. The equations for batch-normalizing one feature are the following, where $N$ is the batch size, $x$ the input features, and $y$ the normalized output features:
#
# $$ \mu = \frac{1}{N} \sum_{i=0}^{N}x_i,\;\;\;\; \sigma^2 = \frac{1}{N} \sum_{i=0}^{N}(x_i - \mu)^2 $$
#
# $$ \hat{x}_i = \frac{x_i - \mu}{\sqrt{\sigma^2 + \epsilon}} $$
#
# $$ y_i = \gamma \hat{x}_i + \beta $$
#
# At first glance this might look intimidating, but all it means is that we begin by scaling and shifting the data to have mean $\mu=0$ and standard deviation $\sigma=1$. After this we use the learnable parameters $\gamma$ and $\beta$ to decide the width and center of the final distribution. $\epsilon$ is a small constant value that prevents the denominator from being zero.
#
# In addition to learning the parameters $\gamma$ and $\beta$ by gradient decent just like the weights, Batch Normalization also keeps track of the running average of minibatch statistics $\mu$ and $\sigma$. These averages are used to normalize the test data. We can tune the rate at which the running averages are updated with the *momentum* parameter of the BatchNormalization layer. A large momentum means that the statistics converge more slowly and therefore requires more updates before it represents the data. A low momentum, on the other hand, adapts to the data more quickly but might lead to unstable behaviour if the latest minibatches are not representative of the whole dataset. For this test we recommend a momentum of 0.75, but you probably want to change this when you design a larger network in Section 5.
#
# The batch normalization layer should be added after the hidden layer linear transformation, but before the nonlinear activation. This means that we cannot specify the activation funciton in the ```Conv2D``` or ```Dense``` if we want to batch-normalize the output. We therefore need to use the ```Activation``` layer to add a separate activation to the network stack after batch normalization. For example, the convolution block will now look like **[conv - batchnorm - activation - pooling]**.
#
# Extend your previous model with batch normalization, both in the convolution and fully connected part of the model.
# +
from tensorflow.keras.layers import BatchNormalization, Activation
x_in = Input(shape=X_train.shape[1:])
# === Add your code here ===
# ==========================
model = Model(inputs=x_in, outputs=x)
sgd = SGD(lr=0.01, decay=1e-6, momentum=0.9, nesterov=True)
model.compile(loss='categorical_crossentropy', metrics=['accuracy'], optimizer=sgd)
model.summary(100)
# -
history = model.fit(X_train, y_train_c, batch_size=32, epochs=25, verbose=1, validation_split=0.2)
# +
score = model.evaluate(X_test, y_test_c, batch_size=128, verbose=0)
for i in range(len(score)):
print("Test " + model.metrics_names[i] + " = %.3f" % score[i])
# -
PlotModelEval(model, history, X_test, y_test, cifar_labels)
# #### **<span style="color:red">Question 8:</span>**
# Train the model and save the evaluation image for the report.
# #### **<span style="color:red">Question 9:</span>**
# When using BatchNorm one must take care to select a good minibatch size. Describe what problems might arise if the wrong minibatch size is used.
#
# You can reason about this given the description of BatchNorm above, or you can search for the information in other sources. Do not forget to provide links to the sources if you do!
#
# Batch Normalization keeps track of the running average of minibatch statistics $\mu$ and $\sigma$ later these averages are used to normalize the test data. If we choose a small minbatch size, there is a high chance that the sample statistics calculated using the data samples within that batch might not represent the overall statistics. For example: If we select the minbatch size as 1
# In addition, if we use a large batchsize then there would small number of batches that could be used to learn the statistics for batchnormalization.
#
# In batch normalization, the parameters $\mu$ and standard deviation $\sigma$ are calculated from the input batch. These values are updated based on the momentum parameter we set. The final mean and standard deviations are calculated from the last batch and values are stored in the model. When we use the model to run over the test data, these parameters are reused from the training phase and not learnt over the test data.
#
# Now, if the batch size is very small, then the $\mu$ and $\sigma$ calculated are not sufficient statistics hence the results might not be stable when we use a large batch/no batch in the testing phase. Because, our $\mu$ value would be very small and there would not be efficient normalization.
#
# When using BatchNorm one must take care to select a good minibatch size. Describe what problems might arise if the wrong minibatch size is used.
#
# You can reason about this given the description of BatchNorm above, or you can search for the information in other sources. Do not forget to provide links to the sources if you do!
#
# As its mentioned in the description of Batch Normalization above, the batch size we select should contain the data points that represent the whole dataset, there by to calculate optimal statistics. In addition, from equations provided above, if the batch size = 1, then the normalized output will be zero. Hence the batch size should be always >1. Now if batch sizes are smaller, chances that the samples representing whole dataset is very less hence algorithm might update the noisy parameters, which is good in a way as we can have better regularization. There is another possibility of frequent skipping of unfavourable local minima if the batch sizes are smaller because of noisy updates. On the other hand, for the large batch size, the parameter updates are closer to the actual gradient and less regularization.(Ref:https://kth.diva-portal.org/smash/get/diva2:955562/FULLTEXT01.pdf)
# (1 liked)
# ### **5. Putting it all together**
# We now want you to create your own model based on what you have learned. We want you to experiment and see what works and what doesn't, so don't go crazy with the number of epochs until you think you have something that works.
#
# To pass this assignment, we want you to acheive **75%** accuracy on the test data in no more than **25 epochs**. This is possible using the layers and techniques we have explored in this notebook, but you are free to use any other methods that we didn't cover. (You are obviously not allowed to cheat, for example by training on the test data.)
# +
from tensorflow.keras.utils import plot_model
x_in = Input(shape=X_train.shape[1:])
# === Add your code here ===
# ==========================
model = Model(inputs=x_in, outputs=x)
sgd = SGD(lr=0.01, decay=1e-6, momentum=0.9, nesterov=True)
model.compile(loss='categorical_crossentropy', metrics=['accuracy'], optimizer=sgd)
model.summary(100)
plot_model(model, show_shapes=True, show_layer_names=False)
# -
history = model.fit(X_train, y_train_c, batch_size=32, epochs=15, verbose=1, validation_split=0.2)
# +
score = model.evaluate(X_test, y_test_c, batch_size=128, verbose=0)
for i in range(len(score)):
print("Test " + model.metrics_names[i] + " = %.3f" % score[i])
# -
PlotModelEval(model, history, X_test, y_test, cifar_labels)
# #### **<span style="color:red">Question 10:</span>**
# Design and train a model that achieves at least 75% test accuracy in at most 25 epochs. Save the evaluation image for the report. Also, in the report you should explain your model and motivate the design choices you have made.
#
#
#
# The design of the model is an extension of the previous model from CNN now with 4 convuloutional layers with 32,64,128 and 256 filters each. The chosen paramaters are as for previous models. After each convolutional layer batchnormalization is performed with 32 as the min-batchsize. Then activation with relu is used followed by pooling as the same as before. After each pass through the steps of convolution - batchnormalization - activation -pooling there is a drop function called with the probability of 30% that the results calculated will not be used and previous results will be sent forward to the next part of the network.
#
# For the fully connected part the neural network layers contained 128, 64 neurons. After each layer batchnormalization is applied and the drop function is called. As previous the output layer uses softmax as activation with 10 neurons.
#
# This model achived in 20 epochs:
#
# Test loss = 0.741
# Test accuracy = 0.756
#
# The addition of batch normalization is to increase the speed of convergence of the network and the drop function is included for regularization such that the model will avoid overfitting on the data. From trial and error we found adding another learning layer and increasing the number of nerons in the fully connected neural network increased both the training and test accuracy. Addding another feature learning layer increased the abstraction of the image features and provide better input for the fully connected part. Implementing batchNormalization in the network after each convolution increases the speed of convergence of the network and the drop function is included for regularization such that the model will avoid overfitting on the data. Increasing the number of neurons in the fully connected part to the model increases the number of non-linear operations which improves the classification.
# ### **Want some extra challenge?**
# For those of you that feel the competitive spark right now, we will hold an **optional** competition where you can submit your trained model-file for evaluation. To make this fair, you are not allowed to train for more than **50 epochs**, but other than that we want you to get creative. The competition is simple, we will evaluate all submitted models and the model with highest test accuracy wins. The prize is nothing less than eternal glory.
#
# Here are some things to look into, but note that we don't have the answers here. Any of these might improve the performance, or might not, or it might only work in combination with each other. This is up to you to figure out. This is how deep learning research often happens, trying things in a smart way to see what works best.
# * Tweak or change the optimizer or training parameters.
# * Tweak the filter parameters, such as numbers and sizes of filters.
# * Use other activation functions.
# * Add L1/L2 regularization (see https://www.tensorflow.org/api_docs/python/tf/keras/regularizers)
# * Include layers that we did not cover here (see https://www.tensorflow.org/api_docs/python/tf/keras/layers). For example, our best model uses the global pooling layers.
# * Take inspiration from some well-known architectures, such as ResNet or VGG16. (But don't just copy-paste those architectures. For one, what's the fun in that? Also, they take a long time to train, you will not have time.)
# * Use explicit model ensembing (training multiple models that vote on or average the outputs - this will also take a lot of time.)
# * Use data augmentation to create a larger training set (see https://www.tensorflow.org/api_docs/python/tf/keras/preprocessing/image/ImageDataGenerator).
#
# Write your competition model here. This way you can try different things without deleting the model you created above. Also set the GroupName variable to your LiU IDs or some unique name; that way our scripts can be a lot easier, thanks and good luck :)
# +
GroupName = "Test"
x_in = Input(shape=X_train.shape[1:])
# Your code here
model = Model(inputs=x_in, outputs=x, name=GroupName)
# You can also change this if you want
sgd = SGD(lr=0.01, decay=1e-6, momentum=0.9, nesterov=True)
model.compile(loss='categorical_crossentropy', metrics=['accuracy'], optimizer=sgd)
# Print the summary and model image
model.summary(100)
plot_model(model, show_shapes=True, show_layer_names=False)
# -
history = model.fit(X_train, y_train_c, batch_size=32, epochs=5, verbose=1, validation_split=0.2)
PlotModelEval(model, history, X_test, y_test, cifar_labels)
# **Don't forget to save your model!**
model.save("CompetionModel_" + GroupName + ".h5")
| A2_DeepLearning/CIFAR10-Lab.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import sys
sys.path.append('../')
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from src.data.data_load_lib import get_from_STEAD, get_from_INSTANCE, get_from_DiTing, get_instance_for_training,train_instance_plot
# ## Check STEAD
STEAD_csv_file_name = '/mnt/GPT_disk/DL_datasets/STEAD/metadata_eq.csv'
STEADDatasetPath = '/mnt/GPT_disk/DL_datasets/STEAD/waveforms.hdf5'
STEAD_csv = pd.read_csv(STEAD_csv_file_name)
STEAD_csv.iloc[:5]
keys = list(STEAD_csv['trace_name'])
choice = np.random.choice(keys)
print(choice)
data, p_t, s_t = get_from_STEAD(key = choice, h5file_path=STEADDatasetPath)
print(np.shape(data))
plt.figure(figsize=(12,6))
# normalize data for plot
for ch_dx in range(np.shape(data)[-1]):
data[:,ch_dx] -= np.mean(data[:,ch_dx])
data[:,ch_dx] /= np.max(data[:,ch_dx])
plt.plot(data[:,ch_dx]+ ch_dx*2,color='k')
plt.plot([p_t,p_t],[-1,5],'b')
plt.plot([s_t,s_t],[-1,5],'r')
#plt.savefig('/home/zhaoming/xzw/DiTIngModelZoo_V2/Debug_imgs/get_from_STEAD.png',dpi=300)
plt.show()
plt.close()
# +
keys = list(STEAD_csv['trace_name'])
choice_key = np.random.choice(keys)
temp_data_X, temp_data_Y = get_instance_for_training(dataset='STEAD',
dataset_path=STEADDatasetPath,
data_length = 1200,
data_channel_num = 1,
key = choice_key,
wave_type='P',
shift_max = 400)
check_id = np.random.randint(16)
temp_data_X = temp_data_X[:,:]
temp_data_Y = temp_data_Y[:,:]
train_instance_plot(temp_data_X, temp_data_Y, './imgs/STEAD_P_vis.png')
# +
keys = list(STEAD_csv['trace_name'])
choice_key = np.random.choice(keys)
temp_data_X, temp_data_Y = get_instance_for_training(dataset='STEAD',
dataset_path=STEADDatasetPath,
data_length = 1600,
data_channel_num = 2,
key = choice_key,
wave_type='S',
shift_max = 400)
check_id = np.random.randint(16)
temp_data_X = temp_data_X[:,:]
temp_data_Y = temp_data_Y[:,:]
train_instance_plot(temp_data_X, temp_data_Y, './imgs/STEAD_S_vis.png')
# -
# ## Check INSTANCE
INSTANCEDatasetPath = '/mnt/GPT_disk/DL_datasets/INSTANCE/Instance_events_counts.hdf5'
INSTANCE_csv_file_name = '/mnt/GPT_disk/DL_datasets/INSTANCE/metadata_Instance_events_both_p_s.csv'
INSTANCE_csv = pd.read_csv(INSTANCE_csv_file_name)
# +
total_lines = len(INSTANCE_csv)
choice_id = np.random.randint(total_lines)
choice_line = INSTANCE_csv.iloc[choice_id]
key = choice_line['trace_name']
print(key)
p_t = choice_line['trace_P_arrival_sample']
s_t = choice_line['trace_S_arrival_sample']
data = get_from_INSTANCE(key=key, h5file_path=INSTANCEDatasetPath)
data = data.T
print(np.shape(data))
plt.figure(figsize=(12,6))
# normalize data for plot
for ch_dx in range(np.shape(data)[-1]):
data[:,ch_dx] -= np.mean(data[:,ch_dx])
data[:,ch_dx] /= np.max(data[:,ch_dx])
plt.plot(data[:,ch_dx]+ ch_dx*2,color='k')
plt.plot([p_t,p_t],[-1,5],'b')
plt.plot([s_t,s_t],[-1,5],'r')
#plt.savefig('/home/zhaoming/xzw/DiTIngModelZoo_V2/Debug_imgs/get_from_STEAD.png',dpi=300)
plt.show()
plt.close()
# +
total_lines = len(INSTANCE_csv)
choice_id = np.random.choice(total_lines)
choice_line = INSTANCE_csv.iloc[choice_id]
key = choice_line['trace_name']
p_t = choice_line['trace_P_arrival_sample']
s_t = choice_line['trace_S_arrival_sample']
temp_data_X, temp_data_Y = get_instance_for_training(dataset='INSTANCE',
dataset_path=INSTANCEDatasetPath,
data_length = 1200,
data_channel_num = 1,
key = key,
wave_type = 'P',
shift_max = 400,
p_t = p_t,
s_t = s_t)
check_id = np.random.randint(16)
temp_data_X = temp_data_X[:,:]
temp_data_Y = temp_data_Y[:,:]
train_instance_plot(temp_data_X, temp_data_Y, './imgs/INSTANCE_P_vis.png')
# +
total_lines = len(INSTANCE_csv)
choice_id = np.random.choice(total_lines)
choice_line = INSTANCE_csv.iloc[choice_id]
key = choice_line['trace_name']
p_t = choice_line['trace_P_arrival_sample']
s_t = choice_line['trace_S_arrival_sample']
temp_data_X, temp_data_Y = get_instance_for_training(dataset='INSTANCE',
dataset_path=INSTANCEDatasetPath,
data_length = 1600,
data_channel_num = 2,
key = key,
wave_type = 'S',
shift_max = 1,
p_t = p_t,
s_t = s_t)
check_id = np.random.randint(16)
temp_data_X = temp_data_X[:,:]
temp_data_Y = temp_data_Y[:,:]
train_instance_plot(temp_data_X, temp_data_Y, './imgs/INSTANCE_S_vis.png')
# -
# # Check DiTing Dataset
DiTingDatasetPath = '/mnt/GPT_disk/DL_datasets/DiTing330km_publish/'
DiTing_csv_file_name = '/mnt/GPT_disk/DL_datasets/DiTing330km_publish/merge.csv'
DiTing_csv = pd.read_csv(DiTing_csv_file_name, dtype = {'key': str})
# +
total_lines = len(DiTing_csv)
choice_id = np.random.randint(total_lines)
choice_line = DiTing_csv.iloc[choice_id]
part = choice_line['part']
key = choice_line['key']
key_correct = key.split('.')
key = key_correct[0].rjust(6,'0') + '.' + key_correct[1].ljust(4,'0')
p_t = int(choice_line['p_pick']*100 + 3000)
s_t = int(choice_line['s_pick']*100 + 3000)
data = get_from_DiTing(part=part, key=key, h5file_path=DiTingDatasetPath)
print(np.shape(data))
plt.figure(figsize=(12,6))
# normalize data for plot
for ch_dx in range(np.shape(data)[-1]):
data[:,ch_dx] -= np.mean(data[:,ch_dx])
data[:,ch_dx] /= np.max(data[:,ch_dx])
plt.plot(data[:,ch_dx]+ ch_dx*2,color='k')
plt.plot([p_t,p_t],[-1,5],'b')
plt.plot([s_t,s_t],[-1,5],'r')
#plt.savefig('/home/zhaoming/xzw/DiTIngModelZoo_V2/Debug_imgs/get_from_STEAD.png',dpi=300)
plt.show()
plt.close()
# +
total_lines = len(DiTing_csv)
choice_id = np.random.choice(total_lines)
choice_line = DiTing_csv.iloc[choice_id]
part = choice_line['part']
key = choice_line['key']
key_correct = key.split('.')
key = key_correct[0].rjust(6,'0') + '.' + key_correct[1].ljust(4,'0')
p_t = choice_line['p_pick']
s_t = choice_line['s_pick']
temp_data_X, temp_data_Y = get_instance_for_training(dataset='DiTing',
dataset_path=DiTingDatasetPath,
data_length = 1200,
data_channel_num = 1,
part = part,
key = key,
wave_type = 'P',
shift_max = 400,
p_t = p_t,
s_t = s_t)
check_id = np.random.randint(16)
temp_data_X = temp_data_X[:,:]
temp_data_Y = temp_data_Y[:,:]
train_instance_plot(temp_data_X, temp_data_Y, './imgs/DiTing_P_vis.png')
# +
total_lines = len(DiTing_csv)
choice_id = np.random.choice(total_lines)
choice_line = DiTing_csv.iloc[choice_id]
part = choice_line['part']
key = choice_line['key']
key_correct = key.split('.')
key = key_correct[0].rjust(6,'0') + '.' + key_correct[1].ljust(4,'0')
p_t = choice_line['p_pick']
s_t = choice_line['s_pick']
temp_data_X, temp_data_Y = get_instance_for_training(dataset='DiTing',
dataset_path=DiTingDatasetPath,
data_length = 1600,
data_channel_num = 2,
part = part,
key = key,
wave_type = 'S',
shift_max = 500,
p_t = p_t,
s_t = s_t)
temp_data_X = temp_data_X[:,:]
temp_data_Y = temp_data_Y[:,:]
train_instance_plot(temp_data_X, temp_data_Y, './imgs/DiTing_S_vis.png')
# -
| notebooks/Inspect STEAD, INSTANCE and DiTing data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Jupyter Notebook
#
# * See the notebook intro from week 1 [here](https://github.com/raspstephan/ESS-Python-Tutorial/blob/master/materials/week1/jupyter-intro.ipynb).
# * You can shut a notebook down by closing browser windows and, from the terminal shell, typing ```ctrl``` + ```c``` twice.
#
# # Quick Python introduction
#
# The [official Python tutorial](https://docs.python.org/3/tutorial/) is very informative, and most of the info below follows it, more or less...
2+2
(70 - (5*6)) / 4
# ## Floats versus integers
#
# Python is a high-level language, and there are **no variable declarations** (Python does this implicitly).
#
# * Numbers like ```2``` and ```4``` are interpreted as type **```int```**, and numbers with decimals (```62.5```, ```10.0```) are type **```float```**.
# * Division **``` / ```** always returns a floating point number
# * Get the type of a number or Python object using ```type()```
2/2
type(2/2)
2/1
# ## Powers in Python
#
# To raise something to a power, use a double asterisk: __```**```__
#
# To ensure your numbers are interpreted as **```float```** types, good practice is to always use decimals in at least one number:
5**2
5**2.0
5.**2
# ## Assigning and printing
#
# * Assign variables using the ```=``` sign
#
# * Print variables using the ```print()``` function
width = 5.0
height = 7.0
area = width*height
print(area)
# Note that Python supports very compact, __in-place__ variable manipulation, allowing for ```+=```, ```-=```, ```*=```, and ```/=```
# ## ==========> NOW YOU TRY <==========
# * Switch between lines 6 and 7 below and mess around with the compact addition, subtraction, multiplication, and division notation
# * __NOTE:__ Python commends are preceded by the ```#``` sign, or you can enclose text ```'''between three quotes'''```
# +
# comment
""" also a comment """
''' also ALSO a comment '''
width = 5.0
width = width+5
#width += 5
print(width)
# -
# ## Strings
#
# Variables can also be assigned as strings when you enclose them in quotes. __You can use single OR double quotes.__
string1 = 'pants'
print(string1)
# Multiplying and adding strings repeats them...
print(string1*3)
print(string1+string1+string1)
# * Strings can also be indexed
# * __NOTE: Python has zero indexing__
string1[0]
string1[1]
# You can also index backwards in Python: index ```-1``` is the last one:
string1[-1]
string1[-3]
# ## Lists
#
# * This is a versatile way of creating compound data types
# * Indexing works in a similar way as above
# * Create a list by enclosing comma-separated items in square brackets
empty_list = []
list1 = [0,1,2,3,4,5,6,7,8,9]
print(empty_list)
print(list1)
print(list1[0])
print(list1[-3])
# ### Slicing lists
#
# Slicing operations return new lists.
list1[3:] # note index 3 is the FOURTH element, since indexing starts at zero
# ### Appending to lists
#
# Add to the end of a list by using the ```append()``` method.
#
# __Note ```list1``` is an "object" in python and has methods that can be accessed via the ```.``` syntax__
list1.append(234)
print(list1)
# ### Assigning and replacing list values
#
# * You can also assign specific values, or do this to slices
# * Lists can contain mixed data types
list2 = ['a', 'b', 'c', 'd', 'e', 'f', 'g']
print(list2)
list2[0:2] = 17,18
print(list2)
# ## ==========> NOW YOU TRY <==========
# * Replace the zeroth position of ```list1``` with your first name (as a ```'string'```)
# * Append your last name to the end of ```list1```
# * You can "pop" an element out of a list, as well, by specifying its index
# * Now delete your first name from the list by using ```list1.pop(index)```
# ### Getting the length of a list
#
# Use the ```len()``` function to get the length of a list.
len(list1)
len(list2)
# ### The ```range()``` and ```list()``` functions
#
# * **```range()```** is a function that creates an *internal* list (i.e., an [iterator](https://wiki.python.org/moin/Iterator) of numbers)
#
# * ```range(N)``` will go FROM ```0``` TO ```N-1``` and will have exactly ```N``` elements
print(range(5)) # goes from 0 to 4 inclusive
print(type(range(5)))
# * Note it returns a range iterator object
# * You can convert this into a Python list by using the **```list()```** function:
list(range(5))
# The **```range()```** function takes at most 3 arguments: start, stop, and interval.
list(range(2,5)) # start, stop
# ## ==========> Question: Why does the output below not include 50? <==========
# +
list(range(0,50,5)) # start, stop, interval
#list(range(0,51,5))
# -
# ## For loops
#
# * Python does not require ```end``` statements or semicolons. Instead, the use of spacing in the loops implicitly tells python where they begin and end. Manually insert (4) spaces (or use a ```tab``` on the keyboard) to struture a loop
#
# * All output in Python is suppressed automatically, unless you choose to ```print()``` it. This means you don't need semicolons at the end of lines, and you'll get an error if you do.
#
# * Loops can begin with **```for```** and **```while```**. Other statements, such as **```else```**, **```if```**, and **```elif```** also exist. Note **```elif```** and **```else if```** are equivalent and both acceptable.
#
# * Note **```range()```** can be used in combination with the **```len()```** or other functions to make looping more streamlined
#
# * Information on all statements useful in loops can be found there, including
# **```if```**, **```for```**, **```break```**, **```continue```**, **```pass```**, and more
weekdays = ['Monday','Tuesday','Wednesday','Thursday','Friday','Saturday','Sunday']
for i in range(7):
print(weekdays[i])
# * Logicals are straightforward: ```>```, ```>=```, ```<```, ```<=```, ```==```, ```!=```
i = 0
while i < 7:
if i==4:
print(" w00t it's Friday")
else:
print(weekdays[i])
i += 1
# * Use ```len(list)``` for looping over indices:
for i in range(len(weekdays)):
print(i, weekdays[i])
# * You can also simply loop over the list itself:
for i in weekdays:
print(i)
# ## Defining functions (an example with the Fibonacci series)
#
# *image from [Wikipedia](https://en.wikipedia.org/wiki/Fibonacci_number)*
#
# 
a,b = 0,1 # note multiple assignments in one line
while b<10:
print(b)
a,b = b,a+b
def fib(n):
"""
Write a Fibonacci series up to AND INCLUDING integer n.
Note this text is called a docstring.
This is how python functions are documented.
""" # <----- this is documentation for the function
a,b = 0,1
while a<n+1:
print(a)
a,b = b, a+b
fib(500)
# ?fib
# ## ==========> NOW YOU TRY <==========
#
# Can you write a function that _returns_ the Fibonacci series in a LIST? The answer is at the bottom of this notebook
# * More advanced information on defining functions can be found online at the [tutorial link](https://docs.python.org/3/tutorial/controlflow.html)
#
# ## The ```lambda``` keyword
#
# Finally, the **```lambda```** keyword is very useful when defining functions. __There's not enough time in this tutorial to go through all the uses of ```lambda```, but it's something you will likely come across when doing more advanced statistical function fitting.__
#
# * In short, the **```lambda```** keyword will help you create anonymous functions (i.e., functions not bound to a name)
#
# * Note the use of **```return```** here, as well.
def raise_to_power(n):
""" This creates a polynomial that raises a number x to the nth power """
return lambda x: x**n
f = raise_to_power(5)
f(0)
f(1)
f(2)
# Also note the string after the **```def()```** statement above. This is a *function annotation*, and anything enclosed in three quotes (**double** or **single**) will not be printed but is useful for multi-line documentation.
# # Common data types in Python
#
# See [Data Structures](https://docs.python.org/3/tutorial/datastructures.html) in the official Python documentation for more information
#
# * integer ```a=1```
#
# * float ```b=1.``` or ```b=1.0```
#
# * list ```c=[1,2,3,4]```
#
# * tuple ```d=(1,2)```
# * Like a list, but can't be changed once it's created
#
# * dictionary ```dict1 = {'a':1, 'b':1.0, 'c':[1,2,3,4], 'd':(1,2) }```
# * Can store mixed data types
# * Unordered set of ```key:value``` pairs
dict1 = {'a':1, 'b':1.0, 'c':[1,2,3,4], 2:(1,2)}
print(dict1.keys())
print(dict1['c'])
print(dict1[2])
# * You can also create an empty dictionary with ```dict1 = {}``` (overwriting the original):
dict1 = {}
# * And then fill it with whatever key you want
dict1['a'] = 1e30
dict1[5] = 'pants'
dict1.keys()
# #### [One possible] answer to Fibonacci series question
# +
# def fib(n):
# fib_list = []
# a,b = 0,1
# fib_list.append(a)
# while a<n+1:
# a,b = b, a+b
# fib_list.append(a)
# return(fib_list)
| materials/week2/1_python_introduction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Results for evaluation with bias
# +
import os
import numpy as np
import random
import pandas as pd
from sklearn import metrics
from operator import itemgetter
import json
map_dict = {'Male': 0,
'Female': 1,
'Sex':2,
'0-19':3,
'20-29':4,
'30-39':5,
'40-49':6,
'50-59':7,
'60-69':8,
'70-79':9,
'80-89':10,
'90-':11,
'Age': 12,
'ecig':13,
'ex': 14,
'never':15,
'ltOnce':16,
'1to10':17,
'21+':18,
'Smoke':19,
'it':20,
'en':21,
'es':22,
'de':23,
'Others':24
}
#Uid;Age;Sex;Medhistory;Smoking;Language;Date;Folder Name;Symptoms;Covid-Tested;Hospitalized;Location;Voice filename;Cough filename;Breath filename
def get_covid(temp):
cot, sym, med, smo = temp[9], temp[8], temp[3], temp[4]
#print(cot, sym, med, smo)
sym_dict = {'drycough':0.0, 'smelltasteloss':0.0, 'headache':0.0,'sorethroat':0.0,
'muscleache':0.0,'wetcough':0.0,'shortbreath':0.0,'tightness':0.0,
'fever':0.0,'dizziness':0.0,'chills':0.0,'runnyblockednose':0.0, 'None': 0.0}
syms = sym.split(',')
for s in syms:
if s == 'tighness':
s = 'tightness'
if s == 'drycoough':
s = 'drycough'
if s == 'runny':
s = 'runnyblockednose'
if s == 'none' or s == '':
s = 'None'
if s in sym_dict:
sym_dict[s] = 1
sym_feature = [sym_dict[s] for s in sorted(sym_dict)]
if cot == 'last14' or cot == 'yes' or cot == 'positiveLast14': #or cot == 'positiveOver14' or cot == 'over14' :
if sym in ['None','','none']: #'pnts'
label = 'covidnosym'
else:
label = 'covidsym'
elif cot == 'negativeNever':
if sym in ['None','','none']:
label = 'healthnosym'
else:
label = 'healthsym'
else:
label = 'negativeLast14_over14'
return label,sym_feature
def get_demo(temp):
dis = [0]*25
uid, age, sex, smo, lan = temp[0], temp[1], temp[2], temp[4], temp[5]
#print(uid, age, sex, smo, lan)
if age in map_dict:
dis[map_dict[age]] = 1.0
else:
dis[map_dict['Age']] = 1.0
if sex in map_dict:
dis[map_dict[sex]] = 1.0
else:
dis[map_dict['Sex']] = 1.0
if smo in map_dict:
dis[map_dict[smo]] = 1.0
else:
dis[map_dict['Smoke']] = 1.0
#Android 10,IOS 12
if lan != 'en': #LOCALE
if len(uid) == 12:
if lan not in ['en','it']:
lan = 'en'
if len(uid) == 10:
if lan not in ['en','it','es','de','pt','el','fr','ru','ro','zh','hi']:
lan = 'en'
#Web users label manaually
if uid in ['2020-04-07-09_13_39_351468',
'2020-04-07-18_06_25_599640',
'2020-04-09-16_06_44_367357',
'2020-04-10-22_47_24_172784',
'2020-04-11-06_26_23_961941',
'2020-04-11-22_05_49_593805',
'2020-04-18-15_48_05_138883',
'2020-06-10-05_39_49_549021',
'2020-09-23-07_01_51_181332',
'2020-11-01-11_42_45_049808',
'2020-11-01-12_58_38_378997',
'2020-11-03-23_31_39_409588',
'2020-11-04-15_18_07_232635',
'2020-11-09-23_11_09_661625',
'2020-11-13-17_08_23_362377',
'2020-11-16-06_11_16_109602',
'2020-11-22-11_06_00_777194',
'2020-11-26-01_59_23_375517',
'2020-12-03-22_29_21_357264',
'2020-12-05-17_17_27_367716',
'2020-12-05-17_39_04_713242',
'2020-12-07-15_23_57_415845',
'2020-12-22-18_31_12_168304',
'2020-12-25-17_26_41_854217',
'2020-12-29-05_57_24_070615',
'2020-12-29-07_16_08_949536',
'2020-12-29-19_20_29_055459',
'2021-01-05-15_13_37_511023',
'2021-01-18-13_11_43_410199',
'2021-01-30-09_11_50_857102',
'2021-03-04-08_23_04_002407',
'2021-03-09-14_44_14_876702',
'2021-03-09-15_11_42_488832',
'2021-03-10-09_53_26_214430',
'2021-03-11-16_22_49_221496',
'2021-03-20-09_00_53_503347',
'']:
lan = 'en'
if uid in ['2020-04-07-18_52_05_802580',
'2020-04-08-19_50_32_255545',
'2020-04-10-09_01_06_580390',
'2020-04-13-18_37_12_935759',
'2020-04-14-10_12_04_529979',
'2020-04-15-07_01_54_598919',
'2020-04-15-13_21_10_153301',
'2020-04-15-17_14_07_448851',
'2020-04-16-11_38_40_431566',
'2020-04-17-19_23_25_139694',
'2020-04-21-11_36_49_766752',
'2020-04-21-12_31_25_239458',
'2020-04-26-16_15_50_150762',
'2020-04-26-16_20_02_616687',
'2020-12-31-08_24_54_055012',
'2021-01-13-20_19_47_216023',
'2021-02-03-17_24_30_080652',
'2021-02-18-18_00_36_137539',
'2021-04-22-20_07_08_880185']:
lan = 'it'
if uid in ['2020-09-08-14_21_40_763674',
'2020-11-11-10_24_29_994922',
'2020-11-19-07_35_21_245792',
'2020-12-10-23_13_10_824970'
]:
lan = 'de'
if uid in ['2020-04-26-10_36_54_607302',
'2020-04-26-19_33_46_515975',
'2020-05-01-17_53_43_125632',
'2020-05-21-08_45_42_105928',
'2020-11-04-14_30_24_061857',
'2021-02-08-15_57_50_219590',
'2021-02-16-21_46_55_000456'
]:
lan = 'es'
if lan in map_dict:
dis[map_dict[lan]] = 1.0
else:
dis[map_dict['Others']] = 1.0
return dis
demo_dict = {}
with open('../COVID19_prediction/data/preprocess/results_all_raw_0426.csv') as f:
for index, line in enumerate(f):
if index>0:
temp = line.strip().split(';')
demo_dict[temp[0] + '/' + temp[7]] = temp
demo_label = sorted(map_dict.items(),key = lambda x:x[1],reverse = False)
def get_metrics(probs,label):
predicted = []
for i in range(len(probs)):
if probs[i]> 0.5:
predicted.append(1)
else:
predicted.append(0)
pre = metrics.precision_score(label, predicted)
acc = metrics.accuracy_score(label, predicted)
auc = metrics.roc_auc_score(label, probs)
precision, recall, _ = metrics.precision_recall_curve(label, probs)
rec = metrics.recall_score(label, predicted)
TN, FP, FN, TP = metrics.confusion_matrix(label,predicted).ravel()
# Sensitivity, hit rate, recall, or true positive rate
TPR = TP*1.0/(TP+FN)
# Specificity or true negative rate
TNR = TN*1.0/(TN+FP)
return auc, TPR, TNR, 0
def get_CI(data, AUC, Sen, Spe):
AUCs = []
TPRs = []
TNRs = []
for s in range(1000):
np.random.seed(s) #Para2
sample = np.random.choice(range(len(data)), len(data), replace=True)
samples = [data[i] for i in sample]
sample_pro = [x[0] for x in samples]
sample_label = [x[1] for x in samples]
try:
get_metrics(sample_pro,sample_label)
except ValueError:
np.random.seed(1001) #Para2
sample = np.random.choice(range(len(data)), len(data), replace=True)
samples = [data[i] for i in sample]
sample_pro = [x[0] for x in samples]
sample_label = [x[1] for x in samples]
else:
auc, TPR, TNR, _ = get_metrics(sample_pro,sample_label)
AUCs.append(auc)
TPRs.append(TPR)
TNRs.append(TNR)
q_0 = pd.DataFrame(np.array(AUCs)).quantile(0.025)[0] #2.5% percentile
q_1 = pd.DataFrame(np.array(AUCs)).quantile(0.975)[0] #97.5% percentile
q_2 = pd.DataFrame(np.array(TPRs)).quantile(0.025)[0] #2.5% percentile
q_3 = pd.DataFrame(np.array(TPRs)).quantile(0.975)[0] #97.5% percentile
q_4 = pd.DataFrame(np.array(TNRs)).quantile(0.025)[0] #2.5% percentile
q_5 = pd.DataFrame(np.array(TNRs)).quantile(0.975)[0] #97.5% percentile
return('&' + str(AUC.round(2)) + '(' + str(q_0.round(2)) + '-' + str(q_1.round(2)) + ')'
+ '&' + str(Sen.round(2)) + '(' + str(q_2.round(2)) + '-' + str(q_3.round(2)) + ')'
'&' + str(Spe.round(2)) + '(' + str(q_4.round(2)) + '-' + str(q_5.round(2)) + ')' )
# -
# ## Supplementary Table 2: User Split
# +
File = 'output/main_userSplitS10_2.txt'
#main results
user = []
probs = []
labels = []
data = []
negative_user = []
with open(File) as f:
for line in f:
uid, pro, label = line.split()
UID, date = uid.split('/')
temp = demo_dict[uid]
demo = get_demo(temp)
if True:
user.append(UID)
pro = float(pro)
label = float(label)
label = 1 if label > 0 else 0
probs.append(pro)
data.append([pro,label])
labels.append(label)
if label == 0:
negative_user.append(UID)
auc, TPR, TNR, _ = get_metrics(probs,labels)
ss = get_CI(data, auc, TPR, TNR)
print('user(samplse)&auc&sensitivity&specificity')
print('&'+str(len(set(user))-len(set(negative_user))) + '(' + str(len((user))-len((negative_user))) + ')'
'/' + str(len(set(negative_user))) + '(' + str(len((negative_user))) + ')'+ ss )
# demographic groups
for d in [0,1]:
print(demo_label[d],'-------------------------------------------------------')
user = []
probs = []
labels = []
data = []
negative_user = []
with open(File) as f:
for line in f:
uid, pro, label = line.split()
UID, date = uid.split('/')
temp = demo_dict[uid]
demo = get_demo(temp)
if demo[d] == 1:
user.append(UID)
pro = float(pro)
label = float(label)
label = 1 if label > 0 else 0
probs.append(pro)
data.append([pro,label])
labels.append(label)
if label == 0:
negative_user.append(UID)
auc, TPR, TNR,_ = get_metrics(probs,labels)
ss = get_CI(data,auc, TPR, TNR)
print('&'+str(len(set(user))-len(set(negative_user))) + '(' + str(len((user))-len((negative_user))) + ')'
'/' + str(len(set(negative_user))) + '(' + str(len((negative_user))) + ')'
+ ss + '//')
for d in [9]:
print('16-39','-------------------------------------------------------')
user = []
probs = []
labels = []
data = []
negative_user = []
with open(File) as f:
for line in f:
uid, pro, label = line.split()
UID, date = uid.split('/')
temp = demo_dict[uid]
demo = get_demo(temp)
if sum(demo[3:6]) >= 1:
user.append(UID)
pro = float(pro)
label = float(label)
label = 1 if label > 0 else 0
probs.append(pro)
data.append([pro,label])
labels.append(label)
if label == 0:
negative_user.append(UID)
auc, TPR, TNR,_ = get_metrics(probs,labels)
ss = get_CI(data,auc, TPR, TNR)
print('&'+str(len(set(user))-len(set(negative_user))) + '(' + str(len((user))-len((negative_user))) + ')'
'/' + str(len(set(negative_user))) + '(' + str(len((negative_user))) + ')'
+ ss + '//')
for d in [9]:
print('40-59','-------------------------------------------------------')
user = []
probs = []
labels = []
data = []
negative_user = []
with open(File) as f:
for line in f:
uid, pro, label = line.split()
UID, date = uid.split('/')
temp = demo_dict[uid]
demo = get_demo(temp)
if sum(demo[6:8]) >= 1:
user.append(UID)
pro = float(pro)
label = float(label)
label = 1 if label > 0 else 0
probs.append(pro)
data.append([pro,label])
labels.append(label)
if label == 0:
negative_user.append(UID)
auc, TPR, TNR,_ = get_metrics(probs,labels)
ss = get_CI(data,auc, TPR, TNR)
print('&'+str(len(set(user))-len(set(negative_user))) + '(' + str(len((user))-len((negative_user))) + ')'
'/' + str(len(set(negative_user))) + '(' + str(len((negative_user))) + ')'
+ ss + '//')
for d in [7]:
print('60-','-------------------------------------------------------')
user = []
probs = []
labels = []
data = []
negative_user = []
with open(File) as f:
for line in f:
uid, pro, label = line.split()
UID, date = uid.split('/')
temp = demo_dict[uid]
demo = get_demo(temp)
if sum(demo[8:12]) >= 1:
user.append(UID)
pro = float(pro)
label = float(label)
label = 1 if label > 0 else 0
probs.append(pro)
data.append([pro,label])
labels.append(label)
if label == 0:
negative_user.append(UID)
auc, TPR, TNR,_ = get_metrics(probs,labels)
ss = get_CI(data,auc, TPR, TNR)
print('&'+str(len(set(user))-len(set(negative_user))) + '(' + str(len((user))-len((negative_user))) + ')'
'/' + str(len(set(negative_user))) + '(' + str(len((negative_user))) + ')'
+ ss + '//')
import pandas as pd
df = pd.read_csv('../COVID19_prediction/data/preprocess/data_0426_en_split.csv', delimiter=',')
fold = df['fold'].tolist()
random.seed(10) #Para1
random.shuffle(fold)
df['fold_new'] = fold
df_train = df[(df['fold_new']<2) & (df['label']==1)]
df_test = df[(df['fold_new']==2) & (df['label']==1)]
df_train_users = df_train['uid'].tolist()
df_test_users = df_test['uid'].tolist()
train_user = [temp.split('/')[0] for temp in df_train_users]
test_user = [temp.split('/')[0] for temp in df_test_users]
#print(len(set(test_user)),len(test_user),len(set(train_user)), len(train_user))
unseen = []
for u in test_user:
if u not in train_user:
unseen.append(u)
#print(len(set(unseen)),len(unseen))
df_train = df[(df['fold_new']<2) & (df['label']==0)]
df_test = df[(df['fold_new']==2) & (df['label']==0)]
df_train_users = df_train['uid'].tolist()
df_test_users = df_test['uid'].tolist()
train_user = [temp.split('/')[0] for temp in df_train_users]
test_user = [temp.split('/')[0] for temp in df_test_users]
#print(len(set(test_user)),len(test_user),len(set(train_user)), len(train_user))
unseen = []
for u in test_user:
if u not in train_user:
unseen.append(u)
#print(len(set(unseen)),len(unseen))
df_train = df[(df['fold_new']<2)]
df_test = df[(df['fold_new']==2)]
df_train_users = df_train['uid'].tolist()
df_test_users = df_test['uid'].tolist()
train_user = [temp.split('/')[0] for temp in df_train_users]
test_user = [temp.split('/')[0] for temp in df_test_users]
# sympotom
for d in [' ']:
user = []
probs = []
labels = []
data = []
negative_user = []
with open(File) as f:
for line in f:
uid, pro, label = line.split()
UID, date = uid.split('/')
temp = demo_dict[uid]
demo = get_demo(temp)
covid,sym = get_covid(temp)
if UID in train_user:
pro = float(pro)
label = float(label)
label = 1 if label > 0 else 0
user.append(UID)
probs.append(pro)
data.append([pro,label])
labels.append(label)
if label == 0:
negative_user.append(UID)
auc, TPR, TNR,_ = get_metrics(probs,labels)
print('Seen:')
ss = get_CI(data,auc, TPR, TNR)
print('&'+str(len(set(user))-len(set(negative_user))) + '(' + str(len((user))-len((negative_user))) + ')'
'/' + str(len(set(negative_user))) + '(' + str(len((negative_user))) + ')'
+ ss + '//')
for d in [' ']:
user = []
probs = []
labels = []
data = []
negative_user = []
with open(File) as f:
for line in f:
uid, pro, label = line.split()
UID, date = uid.split('/')
temp = demo_dict[uid]
demo = get_demo(temp)
covid,sym = get_covid(temp)
if UID not in train_user:
pro = float(pro)
label = float(label)
label = 1 if label > 0 else 0
user.append(UID)
probs.append(pro)
data.append([pro,label])
labels.append(label)
#pre = 1 if pro>0.5 else 0
#if label != pre:
# print(uid, pro, label)
if label == 0:
negative_user.append(UID)
auc, TPR, TNR,_ = get_metrics(probs,labels)
print('Unseen:')
ss = get_CI(data,auc, TPR, TNR)
print('&'+str(len(set(user))-len(set(negative_user))) + '(' + str(len((user))-len((negative_user))) + ')'
'/' + str(len(set(negative_user))) + '(' + str(len((negative_user))) + ')'
+ ss + '//')
# -
# ## Supplementary Table 3: Gender bias
# +
File = 'output/main_gender_bias.txt'
#main results
user = []
probs = []
labels = []
data = []
negative_user = []
with open(File) as f:
for line in f:
uid, pro, label = line.split()
UID, date = uid.split('/')
temp = demo_dict[uid]
demo = get_demo(temp)
if True:
user.append(UID)
pro = float(pro)
label = float(label)
label = 1 if label > 0 else 0
probs.append(pro)
data.append([pro,label])
labels.append(label)
if label == 0:
negative_user.append(UID)
auc, TPR, TNR, _ = get_metrics(probs,labels)
ss = get_CI(data, auc, TPR, TNR)
print('user(samplse)&auc&sensitivity&specificity')
print('&'+str(len(set(user))-len(set(negative_user))) + '(' + str(len((user))-len((negative_user))) + ')'
'/' + str(len(set(negative_user))) + '(' + str(len((negative_user))) + ')'+ ss )
# demographic groups
for d in [0,1]:
print(demo_label[d],'-------------------------------------------------------')
user = []
probs = []
labels = []
data = []
negative_user = []
with open(File) as f:
for line in f:
uid, pro, label = line.split()
UID, date = uid.split('/')
temp = demo_dict[uid]
demo = get_demo(temp)
if demo[d] == 1:
user.append(UID)
pro = float(pro)
label = float(label)
label = 1 if label > 0 else 0
probs.append(pro)
data.append([pro,label])
labels.append(label)
if label == 0:
negative_user.append(UID)
auc, TPR, TNR,_ = get_metrics(probs,labels)
ss = get_CI(data,auc, TPR, TNR)
print('&'+str(len(set(user))-len(set(negative_user))) + '(' + str(len((user))-len((negative_user))) + ')'
'/' + str(len(set(negative_user))) + '(' + str(len((negative_user))) + ')'
+ ss + '//')
for d in [9]:
print('16-39','-------------------------------------------------------')
user = []
probs = []
labels = []
data = []
negative_user = []
with open(File) as f:
for line in f:
uid, pro, label = line.split()
UID, date = uid.split('/')
temp = demo_dict[uid]
demo = get_demo(temp)
if sum(demo[3:6]) >= 1:
user.append(UID)
pro = float(pro)
label = float(label)
label = 1 if label > 0 else 0
probs.append(pro)
data.append([pro,label])
labels.append(label)
if label == 0:
negative_user.append(UID)
auc, TPR, TNR,_ = get_metrics(probs,labels)
ss = get_CI(data,auc, TPR, TNR)
print('&'+str(len(set(user))-len(set(negative_user))) + '(' + str(len((user))-len((negative_user))) + ')'
'/' + str(len(set(negative_user))) + '(' + str(len((negative_user))) + ')'
+ ss + '//')
for d in [9]:
print('40-59','-------------------------------------------------------')
user = []
probs = []
labels = []
data = []
negative_user = []
with open(File) as f:
for line in f:
uid, pro, label = line.split()
UID, date = uid.split('/')
temp = demo_dict[uid]
demo = get_demo(temp)
if sum(demo[6:8]) >= 1:
user.append(UID)
pro = float(pro)
label = float(label)
label = 1 if label > 0 else 0
probs.append(pro)
data.append([pro,label])
labels.append(label)
if label == 0:
negative_user.append(UID)
auc, TPR, TNR,_ = get_metrics(probs,labels)
ss = get_CI(data,auc, TPR, TNR)
print('&'+str(len(set(user))-len(set(negative_user))) + '(' + str(len((user))-len((negative_user))) + ')'
'/' + str(len(set(negative_user))) + '(' + str(len((negative_user))) + ')'
+ ss + '//')
for d in [7]:
print('60-','-------------------------------------------------------')
user = []
probs = []
labels = []
data = []
negative_user = []
with open(File) as f:
for line in f:
uid, pro, label = line.split()
UID, date = uid.split('/')
temp = demo_dict[uid]
demo = get_demo(temp)
if sum(demo[8:12]) >= 1:
user.append(UID)
pro = float(pro)
label = float(label)
label = 1 if label > 0 else 0
probs.append(pro)
data.append([pro,label])
labels.append(label)
if label == 0:
negative_user.append(UID)
auc, TPR, TNR,_ = get_metrics(probs,labels)
ss = get_CI(data,auc, TPR, TNR)
print('&'+str(len(set(user))-len(set(negative_user))) + '(' + str(len((user))-len((negative_user))) + ')'
'/' + str(len(set(negative_user))) + '(' + str(len((negative_user))) + ')'
+ ss + '//')
# -
# ## Supplementary Table 4 and 5: Age bias
# +
#File = 'output/main_age_bias.txt'
File = 'output/main_age2_bias.txt'
#main results
user = []
probs = []
labels = []
data = []
negative_user = []
with open(File) as f:
for line in f:
uid, pro, label = line.split()
UID, date = uid.split('/')
temp = demo_dict[uid]
demo = get_demo(temp)
if True:
user.append(UID)
pro = float(pro)
label = float(label)
label = 1 if label > 0 else 0
probs.append(pro)
data.append([pro,label])
labels.append(label)
if label == 0:
negative_user.append(UID)
auc, TPR, TNR, _ = get_metrics(probs,labels)
ss = get_CI(data, auc, TPR, TNR)
print('user(samplse)&auc&sensitivity&specificity')
print('&'+str(len(set(user))-len(set(negative_user))) + '(' + str(len((user))-len((negative_user))) + ')'
'/' + str(len(set(negative_user))) + '(' + str(len((negative_user))) + ')'+ ss )
# demographic groups
for d in [0,1]:
print(demo_label[d],'-------------------------------------------------------')
user = []
probs = []
labels = []
data = []
negative_user = []
with open(File) as f:
for line in f:
uid, pro, label = line.split()
UID, date = uid.split('/')
temp = demo_dict[uid]
demo = get_demo(temp)
if demo[d] == 1:
user.append(UID)
pro = float(pro)
label = float(label)
label = 1 if label > 0 else 0
probs.append(pro)
data.append([pro,label])
labels.append(label)
if label == 0:
negative_user.append(UID)
auc, TPR, TNR,_ = get_metrics(probs,labels)
ss = get_CI(data,auc, TPR, TNR)
print('&'+str(len(set(user))-len(set(negative_user))) + '(' + str(len((user))-len((negative_user))) + ')'
'/' + str(len(set(negative_user))) + '(' + str(len((negative_user))) + ')'
+ ss + '//')
'''
for d in [9]:
print('16-39','-------------------------------------------------------')
user = []
probs = []
labels = []
data = []
negative_user = []
with open(File) as f:
for line in f:
uid, pro, label = line.split()
UID, date = uid.split('/')
temp = demo_dict[uid]
demo = get_demo(temp)
if sum(demo[3:6]) >= 1:
user.append(UID)
pro = float(pro)
label = float(label)
label = 1 if label > 0 else 0
probs.append(pro)
data.append([pro,label])
labels.append(label)
if label == 0:
negative_user.append(UID)
auc, TPR, TNR,_ = get_metrics(probs,labels)
ss = get_CI(data,auc, TPR, TNR)
print('&'+str(len(set(user))-len(set(negative_user))) + '(' + str(len((user))-len((negative_user))) + ')'
'/' + str(len(set(negative_user))) + '(' + str(len((negative_user))) + ')'
+ ss + '//')
'''
for d in [9]:
print('40-59','-------------------------------------------------------')
user = []
probs = []
labels = []
data = []
negative_user = []
with open(File) as f:
for line in f:
uid, pro, label = line.split()
UID, date = uid.split('/')
temp = demo_dict[uid]
demo = get_demo(temp)
if sum(demo[6:8]) >= 1:
user.append(UID)
pro = float(pro)
label = float(label)
label = 1 if label > 0 else 0
probs.append(pro)
data.append([pro,label])
labels.append(label)
if label == 0:
negative_user.append(UID)
auc, TPR, TNR,_ = get_metrics(probs,labels)
ss = get_CI(data,auc, TPR, TNR)
print('&'+str(len(set(user))-len(set(negative_user))) + '(' + str(len((user))-len((negative_user))) + ')'
'/' + str(len(set(negative_user))) + '(' + str(len((negative_user))) + ')'
+ ss + '//')
for d in [7]:
print('60-','-------------------------------------------------------')
user = []
probs = []
labels = []
data = []
negative_user = []
with open(File) as f:
for line in f:
uid, pro, label = line.split()
UID, date = uid.split('/')
temp = demo_dict[uid]
demo = get_demo(temp)
if sum(demo[8:12]) >= 1:
user.append(UID)
pro = float(pro)
label = float(label)
label = 1 if label > 0 else 0
probs.append(pro)
data.append([pro,label])
labels.append(label)
if label == 0:
negative_user.append(UID)
auc, TPR, TNR,_ = get_metrics(probs,labels)
ss = get_CI(data,auc, TPR, TNR)
print('&'+str(len(set(user))-len(set(negative_user))) + '(' + str(len((user))-len((negative_user))) + ')'
'/' + str(len(set(negative_user))) + '(' + str(len((negative_user))) + ')'
+ ss + '//')
# -
# ## Supplementary Table 6: Language bias
# +
File = 'output/main_lan_bias.txt'
# change file to main_lan_B.txt, main_lan_C.txt, main_lan_V.txt
print('user(samplse)&auc&sensitivity&specificity')
#main results
print('Three modality:')
user = []
probs = []
labels = []
data = []
negative_user = []
with open(File) as f:
for line in f:
uid, pro, label = line.split()
UID, date = uid.split('/')
temp = demo_dict[uid]
demo = get_demo(temp)
if True:
user.append(UID)
pro = float(pro)
label = float(label)
label = 1 if label > 0 else 0
probs.append(pro)
data.append([pro,label])
labels.append(label)
if label == 0:
negative_user.append(UID)
auc, TPR, TNR, _ = get_metrics(probs,labels)
ss = get_CI(data, auc, TPR, TNR)
print('&'+str(len(set(user))-len(set(negative_user))) + '(' + str(len((user))-len((negative_user))) + ')'
'/' + str(len(set(negative_user))) + '(' + str(len((negative_user))) + ')'+ ss )
user = []
probs = []
labels = []
data = []
negative_user = []
with open(File) as f:
for line in f:
uid, pro, label = line.split()
UID, date = uid.split('/')
temp = demo_dict[uid]
demo = get_demo(temp)
if demo[20]==1:
user.append(UID)
pro = float(pro)
label = float(label)
label = 1 if label > 0 else 0
probs.append(pro)
data.append([pro,label])
labels.append(label)
if label == 0:
negative_user.append(UID)
auc, TPR, TNR, _ = get_metrics(probs,labels)
ss = get_CI(data, auc, TPR, TNR)
print('Italy')
print('&'+str(len(set(user))-len(set(negative_user))) + '(' + str(len((user))-len((negative_user))) + ')'
'/' + str(len(set(negative_user))) + '(' + str(len((negative_user))) + ')'+ ss )
user = []
probs = []
labels = []
data = []
negative_user = []
with open(File) as f:
for line in f:
uid, pro, label = line.split()
UID, date = uid.split('/')
temp = demo_dict[uid]
demo = get_demo(temp)
if demo[20]!=1:
user.append(UID)
pro = float(pro)
label = float(label)
label = 1 if label > 0 else 0
probs.append(pro)
data.append([pro,label])
labels.append(label)
if label == 0:
negative_user.append(UID)
auc, TPR, TNR, _ = get_metrics(probs,labels)
ss = get_CI(data, auc, TPR, TNR)
print('English')
print('&'+str(len(set(user))-len(set(negative_user))) + '(' + str(len((user))-len((negative_user))) + ')'
'/' + str(len(set(negative_user))) + '(' + str(len((negative_user))) + ')'+ ss )
File = 'output/main_lan_B.txt'
#main results
print('Breathing modality:')
user = []
probs = []
labels = []
data = []
negative_user = []
with open(File) as f:
for line in f:
uid, pro, label = line.split()
UID, date = uid.split('/')
temp = demo_dict[uid]
demo = get_demo(temp)
if True:
user.append(UID)
pro = float(pro)
label = float(label)
label = 1 if label > 0 else 0
probs.append(pro)
data.append([pro,label])
labels.append(label)
if label == 0:
negative_user.append(UID)
auc, TPR, TNR, _ = get_metrics(probs,labels)
ss = get_CI(data, auc, TPR, TNR)
print('&'+str(len(set(user))-len(set(negative_user))) + '(' + str(len((user))-len((negative_user))) + ')'
'/' + str(len(set(negative_user))) + '(' + str(len((negative_user))) + ')'+ ss )
user = []
probs = []
labels = []
data = []
negative_user = []
with open(File) as f:
for line in f:
uid, pro, label = line.split()
UID, date = uid.split('/')
temp = demo_dict[uid]
demo = get_demo(temp)
if demo[20]==1:
user.append(UID)
pro = float(pro)
label = float(label)
label = 1 if label > 0 else 0
probs.append(pro)
data.append([pro,label])
labels.append(label)
if label == 0:
negative_user.append(UID)
auc, TPR, TNR, _ = get_metrics(probs,labels)
ss = get_CI(data, auc, TPR, TNR)
print('Italy')
print('&'+str(len(set(user))-len(set(negative_user))) + '(' + str(len((user))-len((negative_user))) + ')'
'/' + str(len(set(negative_user))) + '(' + str(len((negative_user))) + ')'+ ss )
user = []
probs = []
labels = []
data = []
negative_user = []
with open(File) as f:
for line in f:
uid, pro, label = line.split()
UID, date = uid.split('/')
temp = demo_dict[uid]
demo = get_demo(temp)
if demo[20]!=1:
user.append(UID)
pro = float(pro)
label = float(label)
label = 1 if label > 0 else 0
probs.append(pro)
data.append([pro,label])
labels.append(label)
if label == 0:
negative_user.append(UID)
auc, TPR, TNR, _ = get_metrics(probs,labels)
ss = get_CI(data, auc, TPR, TNR)
print('English')
print('&'+str(len(set(user))-len(set(negative_user))) + '(' + str(len((user))-len((negative_user))) + ')'
'/' + str(len(set(negative_user))) + '(' + str(len((negative_user))) + ')'+ ss )
File = 'output/main_lan_C.txt'
#main results
print('Cough modality:')
user = []
probs = []
labels = []
data = []
negative_user = []
with open(File) as f:
for line in f:
uid, pro, label = line.split()
UID, date = uid.split('/')
temp = demo_dict[uid]
demo = get_demo(temp)
if True:
user.append(UID)
pro = float(pro)
label = float(label)
label = 1 if label > 0 else 0
probs.append(pro)
data.append([pro,label])
labels.append(label)
if label == 0:
negative_user.append(UID)
auc, TPR, TNR, _ = get_metrics(probs,labels)
ss = get_CI(data, auc, TPR, TNR)
print('&'+str(len(set(user))-len(set(negative_user))) + '(' + str(len((user))-len((negative_user))) + ')'
'/' + str(len(set(negative_user))) + '(' + str(len((negative_user))) + ')'+ ss )
user = []
probs = []
labels = []
data = []
negative_user = []
with open(File) as f:
for line in f:
uid, pro, label = line.split()
UID, date = uid.split('/')
temp = demo_dict[uid]
demo = get_demo(temp)
if demo[20]==1:
user.append(UID)
pro = float(pro)
label = float(label)
label = 1 if label > 0 else 0
probs.append(pro)
data.append([pro,label])
labels.append(label)
if label == 0:
negative_user.append(UID)
auc, TPR, TNR, _ = get_metrics(probs,labels)
ss = get_CI(data, auc, TPR, TNR)
print('Italy')
print('&'+str(len(set(user))-len(set(negative_user))) + '(' + str(len((user))-len((negative_user))) + ')'
'/' + str(len(set(negative_user))) + '(' + str(len((negative_user))) + ')'+ ss )
user = []
probs = []
labels = []
data = []
negative_user = []
with open(File) as f:
for line in f:
uid, pro, label = line.split()
UID, date = uid.split('/')
temp = demo_dict[uid]
demo = get_demo(temp)
if demo[20]!=1:
user.append(UID)
pro = float(pro)
label = float(label)
label = 1 if label > 0 else 0
probs.append(pro)
data.append([pro,label])
labels.append(label)
if label == 0:
negative_user.append(UID)
auc, TPR, TNR, _ = get_metrics(probs,labels)
ss = get_CI(data, auc, TPR, TNR)
print('English')
print('&'+str(len(set(user))-len(set(negative_user))) + '(' + str(len((user))-len((negative_user))) + ')'
'/' + str(len(set(negative_user))) + '(' + str(len((negative_user))) + ')'+ ss )
File = 'output/main_lan_V.txt'
#main results
print('Voice modality:')
user = []
probs = []
labels = []
data = []
negative_user = []
with open(File) as f:
for line in f:
uid, pro, label = line.split()
UID, date = uid.split('/')
temp = demo_dict[uid]
demo = get_demo(temp)
if True:
user.append(UID)
pro = float(pro)
label = float(label)
label = 1 if label > 0 else 0
probs.append(pro)
data.append([pro,label])
labels.append(label)
if label == 0:
negative_user.append(UID)
auc, TPR, TNR, _ = get_metrics(probs,labels)
ss = get_CI(data, auc, TPR, TNR)
print('&'+str(len(set(user))-len(set(negative_user))) + '(' + str(len((user))-len((negative_user))) + ')'
'/' + str(len(set(negative_user))) + '(' + str(len((negative_user))) + ')'+ ss )
user = []
probs = []
labels = []
data = []
negative_user = []
with open(File) as f:
for line in f:
uid, pro, label = line.split()
UID, date = uid.split('/')
temp = demo_dict[uid]
demo = get_demo(temp)
if demo[20]==1:
user.append(UID)
pro = float(pro)
label = float(label)
label = 1 if label > 0 else 0
probs.append(pro)
data.append([pro,label])
labels.append(label)
if label == 0:
negative_user.append(UID)
auc, TPR, TNR, _ = get_metrics(probs,labels)
ss = get_CI(data, auc, TPR, TNR)
print('Italy')
print('&'+str(len(set(user))-len(set(negative_user))) + '(' + str(len((user))-len((negative_user))) + ')'
'/' + str(len(set(negative_user))) + '(' + str(len((negative_user))) + ')'+ ss )
user = []
probs = []
labels = []
data = []
negative_user = []
with open(File) as f:
for line in f:
uid, pro, label = line.split()
UID, date = uid.split('/')
temp = demo_dict[uid]
demo = get_demo(temp)
if demo[20]!=1:
user.append(UID)
pro = float(pro)
label = float(label)
label = 1 if label > 0 else 0
probs.append(pro)
data.append([pro,label])
labels.append(label)
if label == 0:
negative_user.append(UID)
auc, TPR, TNR, _ = get_metrics(probs,labels)
ss = get_CI(data, auc, TPR, TNR)
print('English')
print('&'+str(len(set(user))-len(set(negative_user))) + '(' + str(len((user))-len((negative_user))) + ')'
'/' + str(len(set(negative_user))) + '(' + str(len((negative_user))) + ')'+ ss )
# -
| results/Experiment_with_bias_table.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # nteract - open issue and open pr counts
# Send a graphql query to GitHub
# and work with files for reports.
#
# Supports Python 3.6+
# +
import csv
import json
import os
import pprint
import requests
# -
# get api token and set authorization
api_token = os.environ['GITHUB_API_TOKEN']
headers = {'Authorization': f'token {api_token}'}
# set url to a graphql endpoint
url = 'https://api.github.com/graphql'
# add a json query
query = """
{
organization(login: "nteract") {
repositories(first: 80) {
nodes {
name
url
issues(states: OPEN) {
totalCount
}
pullRequests(states: OPEN) {
totalCount
}
}
}
}
}
"""
# ### Make request and create json and csv files
# submit the request
r = requests.post(url=url, json={'query': query}, headers=headers)
# create a json file from response
with open('data.json', 'w') as f:
json.dump(r.json(), f)
# +
# unpack the layers of json
nodes = r.json()['data']['organization']['repositories']['nodes']
unpacked = []
for node in nodes:
unpacked.append(node)
# +
headers = ['name', 'url', 'issues', 'prs']
rows = []
for obj in unpacked:
new_dict = {'name':obj['name'], 'url':obj['url'], 'issues':obj['issues']['totalCount'], 'prs':obj['pullRequests']['totalCount']}
rows.append(new_dict)
# -
with open('mydata.csv', 'w') as f:
f_csv = csv.DictWriter(f, headers)
f_csv.writeheader()
f_csv.writerows(rows)
# Check file
# +
# #%%bash
#less mydata.csv
# -
# ## Bring into pandas
import pandas as pd
df = pd.read_csv('mydata.csv')
df.columns
# +
# df.head()
# -
# Generate basic report of total open issues
# +
# df.dtypes
# +
# df.index
# +
# df.values
# -
# ### Reports
# By repo name
df.sort_values(by=['name'])
# By open issue count
df.sort_values(by=['issues'], ascending=False).head(10)
# by open pr count
df.sort_values(by=['prs'], ascending=False)
# +
# output data to a csv
# df.to_csv('issue_report.csv')
# -
| nteract-open-issues-pr-counts-by-repo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="cedtXySEYb28"
# <div class="alert alert-block alert-info"><b></b>
# <h1><center> <font color='black'> Homework 04 </font></center></h1>
# <h2><center> <font color='black'> Cross-sell / Up-sell using Recommendations </font></center></h2>
# <h2><center> <font color='black'> Due date : 26 April 23:59 </font></center></h2>
# <h2><center> <font color='black'> BDA - University of Tartu - Spring 2020</font></center></h3>
# </div>
# + [markdown] colab_type="text" id="dHyOkasBYb3D"
# # Homework instructions
#
# + [markdown] colab_type="text" id="B-pvZUeIYb3G"
# - Insert your team member names and student IDs in the field "Team mates" below. If you are not working in a team please insert only your name, surname and student ID
#
# - The accepted submission formats are Colab links or .ipynb files. If you are submitting Colab links please make sure that the privacy settings for the file is public so we can access your code.
#
# - The submission will automatically close at 12:00 am, so please make sure you have enough time to submit the homework.
#
# - Only one of the teammates should submit the homework. We will grade and give points to both of you!
#
# - You do not necessarily need to work on Colab. Especially as the size and the complexity of datasets will increase through the course, you can install jupyter notebooks locally and work from there.
#
# - If you do not understand what a question is asking for, please ask in Moodle.
#
# + [markdown] colab_type="text" id="9OWlFadiYb3I"
# **<h2><font color='red'>Team mates:</font></h2>**
#
#
# <font color='red'>Name Surname: Enlik -</font>  <font color='red'>Student ID: B96323</font>
#
#
# <font color='red'>Name Surname: </font>  <font color='red'>Student ID: </font>
# + [markdown] colab_type="text" id="boFT1CkoYb3K"
# # 1. Market Basket Analysis (2 points)
# + [markdown] colab_type="text" id="a3hBebgbYb3M"
# **1.1 Consider the following businesses and think about one case of cross selling and one case of up selling techniques they could use. This question is not restricted to only traditional, standard examples. If you wish you can provide something that you would like these businesses to do. (1 points)**
# + [markdown] colab_type="text" id="HxMUA01DYb3P"
# a. An OnlineTravel Agency like Booking.com or AirBnB
# + [markdown] colab_type="text" id="RODzp7BPYb3T"
# <font color='red'> **Cross selling:**</font> recommend some tour packages made by locals
#
# <font color='red'> **Up selling:**</font> get the booking cancellation for free by adding extra 10% of the booking price
#
# + [markdown] colab_type="text" id="Qbw_w9p1Yb3U"
# b. A software company which produces products related to cyber security like Norton, Kaspersky, Avast and similar ones.
# + [markdown] colab_type="text" id="j0SyXnB6Yb3W"
# <font color='red'> **Cross selling:**</font> get 50% discount for the mobile version (Android/iOS) when purchase our subscription plan in Windows/Mac
#
# <font color='red'> **Up selling:**</font> get the best value (30% cheaper compare to monthly plan) when subscribe our yearly plan
# + [markdown] colab_type="text" id="7EUCv8TtYb3X"
# c. A company that sells cell phones
# + [markdown] colab_type="text" id="NFHO-dI6Yb3Y"
# <font color='red'> **Cross selling:**</font> give an offer to purchase high-quality screen protector and phone case that suits to the phone that you will buy.
#
# <font color='red'> **Up selling:**</font> get additional 128GB internal storage just by adding 50EUR to current price
# + [markdown] colab_type="text" id="_wnH4-lrYb3a"
# d. A supermarket like Konsum, Rimi, Maxima etc.
# + [markdown] colab_type="text" id="I4CNtNYBYb3b"
# <font color='red'> **Cross selling:**</font> group the similar products together for a specific occasion, for example **Christmas** season, when customer can buy decoration, gift wrap, chocolate, and other christmas gifts in one place
#
# <font color='red'> **Up selling:**</font> give an incentive (for example: extra 50% loyalty reward point) when customer spending more than 50EUR in one-time shopping
# + [markdown] colab_type="text" id="DLp7o0cdYb3c"
# **1.2 One of the techniques which we discussed in the theory lecture for recommendater systems is Market Basket Analysis. The aim is to study the products bought frequently together and to recommend product in bunndles. Let's suppose that our client is a retail company that has an online shop. They have given to us the OnlineRetail.csv dataset (we have previously used this dataset in our practice sessions 03). It contains data about the online sales of several products. The client or wants to know which product bundles to promote. Let us find the 5 association rules with the highest lift.**
# + colab={} colab_type="code" id="b7HLlQ30Yb3e"
import pandas as pd
df = pd.read_csv('OnlineRetailPurchase.csv', header=0 )
# + colab={} colab_type="code" id="VWBRFwuUYb3l" outputId="56e2ce1f-6d98-4ed1-aa07-2c6c9c268b11"
df.head()
# + [markdown] colab_type="text" id="kcjIimkHYb35"
# **1.2 Use describe function from pandas to get statistical information about the values in the dataframe. Do you notice something which might not be correct? If so please perform the necessary operations. (Hint: Remember what we did in the practice session 03)(0.25 points)**
# + colab={} colab_type="code" id="RakInjZBY4Wu"
df.describe()
# -
print(sum(df['Quantity'] < 0))
# Remove all rows with quantity less than zero
df = df[df['Quantity'] >= 0]
df.describe()
df['Description'] = df['Description'].str.strip()
df.dropna(axis=0, subset=['InvoiceNo'], inplace=True)
df['InvoiceNo'] = df['InvoiceNo'].astype('str')
df = df[~df['InvoiceNo'].str.contains('C')]
df.head()
# + [markdown] colab_type="text" id="J5a0X9dtYb4K"
# **1.3 Create a dataframe name as "Basket", where each row has an distintive value of InvoiceNo and each column has a distinctive Description. The cells in the table contain the count of each item (Description) mentioned in one invoice. For example basket.loc['536365','WHITE HANGING HEART T-LIGHT HOLDER'] has a value of 1 because the product with WHITE HANGING HEART T-LIGHT HOLDER was entered only once in the invoice 536365. Hint: Remember the function you used in Homework 1 for a similar task or in practice session no.07 (0.25 points)**
# + colab={} colab_type="code" id="D4lUPlKAYb4L"
# Reference from Lab 07 - Reommendations System
## get number of ratings given by every user
# df_users_cnt = pd.DataFrame(df_ratings_drop_movies.groupby('userId').size(), columns=['count'])
# df_users_cnt.head()
# basket = pd.DataFrame(df.groupby(['InvoiceNo', 'Description']).sum().unstack().reset_index().set_index('InvoiceNo'))
# basket.rename_axis(None, inplace=True)
# basket = pd.DataFrame(df.groupby(['InvoiceNo', 'Description']).size(), columns=['Count'])
# Reference:
# https://pbpython.com/market-basket-analysis.html
# basket = (df.groupby(['InvoiceNo', 'Description'])['Quantity']
# .sum().unstack().reset_index().fillna(0)
# .set_index('InvoiceNo'))
basket = (df.groupby(['InvoiceNo', 'Description'])
.size().unstack().reset_index().fillna(0).set_index('InvoiceNo'))
basket.head()
# -
basket.loc['536365','WHITE HANGING HEART T-LIGHT HOLDER']
basket.loc['537224','WHITE BELL HONEYCOMB PAPER']
basket.loc['536412','12 DAISY PEGS IN WOOD BOX']
# +
# basket.describe()
# + [markdown] colab_type="text" id="3rwKSVg3Yb4d"
# **1.4 Some products are mentioned more than once in one invoice. You can check the maximum number for each column to
# verify. Modify your dataframe such that every cell which has a value higher than one will be replaced with 1. If the cell has the value 0 it will remain the same. (0.25 points)**
# +
# fucntion for "one-hot encoding" of the data
def encode_units(x):
if x <= 0:
return 0
if x >= 1:
return 1
basket_sets = basket.applymap(encode_units)
#remove "POSTAGE" column because it's not relevant in our association exploration
basket_sets.drop('POSTAGE', inplace=True, axis=1)
# -
basket_sets
# + [markdown] colab_type="text" id="KfWgocGTYb4k"
# **1.5 We do not need to spend time on calculating the association rules by ourselves as there already exists a package for python to do so, called mlxtend. We are going to use the mlxtend package to find frequent items bought together and then create some rules on what to recomend to a user based on what he/she/they have bought. We have given you the first part of the code which calculates the frequent items bought together.**
# + colab={} colab_type="code" id="rCw4ii7tYb4l"
# #!pip install mlxtend
from mlxtend.frequent_patterns import apriori
from mlxtend.frequent_patterns import association_rules
import mlxtend as ml
import math
# + colab={} colab_type="code" id="nQBjILk5Yb4p"
# Mlxtend has implemented Apriori, a popular algorithm for extracting frequent itemsets
# We can change the value of minimum support but it will
# But as well we get less useful results for the next step.
# Setting use_colnames=True to convert the returned integer indices into item names
frequent_itemsets = apriori(basket_sets, min_support=0.03, use_colnames=True)
frequent_itemsets
# + [markdown] colab_type="text" id="GcF5RyYRYb4y"
# **Please read the documentation of the associaton rules function in mlextend [here](http://rasbt.github.io/mlxtend/user_guide/frequent_patterns/association_rules/) and then complete the code so we get the 5 rules with the highest lift. Print those rules. In the output antecedents represent if .... clause and consequents represent else... clause. For example if user bought product basket A then the algorithm recommends product basket B. (0.25 points)**
# -
rules = association_rules(frequent_itemsets, metric="lift")
rules.head()
# + [markdown] colab_type="text" id="kRqo0ek4Yb47"
# # 2. Collaborative filtering (3.5 points )
# + [markdown] colab_type="text" id="_U1OvsCJYb48"
# We are going to use Books.csv dataset which contains ratings from Amazon website and the data has the following features:
#
# UserID: The ID of the users who read the books
#
# BookTitle: The title of the book
#
# Book-Rating: A rating given to the book in a scale from 0 to 10
#
# Below we are going to perform the same steps we did with movies dataset in the practice session
# + [markdown] colab_type="text" id="_-wOm7yLYb49"
# **2.0 Load the dataset and take a look at the books titles. Please pick one of them which you like (or think that you would like) the most.(0.1 points)**
# + colab={} colab_type="code" id="Z_2CgjU6Yb4-"
books = pd.read_csv("Books.csv", header=0)
books.head()
# -
# pick my favorite book = 'Harry Potter and the Order of the Phoenix (Book 5)'
books[books['BookTitle'] == 'H<NAME>ter and the Order of the Phoenix (Book 5)'].head()
books.iloc[[1818]]
# + [markdown] colab_type="text" id="Q_f2ywpLYb5J"
# **2.1 Our next step will be to perform user based collaborative filtering using KNN algorithm. As KNN algorithm does not accept strings, use a Label Encoder for BookTitle column.After that reshape the books matrix using pivot so every column will be a UserID and every row a BookTitle. (0.45 points)**
# + colab={} colab_type="code" id="-Gs_CAGKYb5K"
# Reference
# https://stackoverflow.com/questions/24458645/label-encoding-across-multiple-columns-in-scikit-learn
from sklearn.preprocessing import LabelEncoder
books_encoded = books.apply(LabelEncoder().fit_transform)
books_encoded = books_encoded.rename(columns={"BookTitle": "BookID_Encoded"})
books_encoded
# -
books_encoded_no_dup = books_encoded.drop_duplicates(['UserID', 'BookID_Encoded'])
books_encoded_no_dup
# +
# Reference
# https://datascienceplus.com/building-a-book-recommender-system-the-basics-knn-and-matrix-factorization/
# Practice Lab 07
books_matrix = books_encoded_no_dup.pivot(index = 'BookID_Encoded', columns = 'UserID', values = 'Book-Rating').fillna(0)
print(books_matrix.shape)
books_matrix.head()
# + [markdown] colab_type="text" id="4RwLx90KYb5R"
# **2.2 Build a sparse matrix for books data and show it. (0.45 points)**
# + colab={} colab_type="code" id="uwVtesasYb5U"
# Reference - Practice Lab 07
# transform matrix to scipy sparse matrix
from scipy.sparse import csr_matrix
books_matrix_sparse = csr_matrix(books_matrix.values)
print(f"Sparse matrix:\n{books_matrix_sparse}")
# + [markdown] colab_type="text" id="PrKKbiRJYb5g"
# **2.3 Build and train two different KNN models (use cosine metric for similarity for both) but with different n_neighbours, that is 2 and 10. Recommend top 5 books based on your favourite one from 2.0 in both cases (1 points)**
# + colab={} colab_type="code" id="bHN1hcjOYb5h"
from sklearn.neighbors import NearestNeighbors
model_knn_1 = NearestNeighbors(metric = 'cosine', algorithm = 'brute', n_neighbors=2)
model_knn_2 = NearestNeighbors(metric = 'cosine', algorithm = 'brute', n_neighbors=10)
# -
model_knn_1.fit(books_matrix_sparse)
model_knn_2.fit(books_matrix_sparse)
# Create new dataframe for
# Reference
# https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.concat.html
# https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.drop.html
books_encoded_title = pd.concat([books_encoded, books], join="outer", axis = 1)
books_encoded_title = books_encoded_title[['BookID_Encoded', 'BookTitle']]
books_encoded_title = books_encoded_title.drop_duplicates()
books_encoded_title = books_encoded_title.reset_index()
books_encoded_title = books_encoded_title.drop(columns = ['index'])
books_encoded_title.head()
# create mapper from book title to index
# book: index
book_to_idx = {
book: i for i, book in enumerate(list(books_encoded_title.set_index('BookID_Encoded').loc[books_matrix.index].BookTitle))
}
book_to_idx
# +
# utils import
from fuzzywuzzy import fuzz
def fuzzy_matching(mapper, fav_book, verbose=True):
"""
return the closest match via fuzzy ratio. If no match found, return None
Parameters
----------
mapper: dict, map book title name to index of the book in data
fav_book: str, name of user input book
verbose: bool, print log if True
Return
------
index of the closest match
"""
match_tuple = []
# get match
for title, idx in mapper.items():
ratio = fuzz.ratio(title.lower(), fav_book.lower())
if ratio >= 60:
match_tuple.append((title, idx, ratio))
# sort
match_tuple = sorted(match_tuple, key=lambda x: x[2])[::-1]
if not match_tuple:
print('Oops! No match is found')
return
if verbose:
print('Found possible matches in our database: {0}\n'.format([x[0] for x in match_tuple]))
return match_tuple[0][1]
def make_recommendation(model_knn, data, mapper, fav_book, n_recommendations):
"""
return top n similar book recommendations based on user's input book
Parameters
----------
model_knn: sklearn model, knn model
data: book-user matrix
mapper: dict, map book title name to index of the book in data
fav_book: str, name of user input book
n_recommendations: int, top n recommendations
Return
------
list of top n similar book recommendations
"""
# fit
model_knn.fit(data)
# get input book index
print('You have input book:', fav_book)
idx = fuzzy_matching(mapper, fav_book, verbose=True)
# inference
print('Recommendation system start to make inference')
print('......\n')
distances, indices = model_knn.kneighbors(data[idx], n_neighbors=n_recommendations+1)
# get list of raw idx of recommendations
raw_recommends = \
sorted(list(zip(indices.squeeze().tolist(), distances.squeeze().tolist())), key=lambda x: x[1])[:0:-1]
# get reverse mapper
reverse_mapper = {v: k for k, v in mapper.items()}
# print recommendations
print('Recommendations for {}:'.format(fav_book))
for i, (idx, dist) in reversed(list(enumerate(raw_recommends))):
#j =i
print('{0}: {1}, with distance of {2}'.format(n_recommendations-i, reverse_mapper[idx], dist))
# -
my_favorite = '<NAME> and the Order of the Phoenix (Book 5)'
# my_favorite = '<NAME> and the Prisoner of Azkaban'
# my_favorite = 'How To Win Friends And Influence People'
# my_favorite = 'Sushi for Beginners : A Novel (Keyes, Marian)'
# +
#TODO - check why the result with different model is exactly same
# Recommendation based on KNN model 1
make_recommendation(
model_knn=model_knn_1, # trained model (model)
data=books_matrix_sparse, # sparse matrix (data)
fav_book=my_favorite, # fav_book
mapper=book_to_idx, # {book: index} (mapper)
n_recommendations=5)
# -
# Recommendation based on KNN model 2
make_recommendation(
model_knn=model_knn_2, # trained model (model)
data=books_matrix_sparse, # sparse matrix (data)
fav_book=my_favorite, # fav_book
mapper=book_to_idx, # {book: index} (mapper)
n_recommendations=5)
# + [markdown] colab_type="text" id="WCJz_Do9Yb5q"
# **2.4 Discuss the results you received in both cases. Would you like to read some of the recommended books? Out of 2 or 10 neighbors, which one worked better? (There is no right or wrong answer in this question) (0.25 points)**
# + [markdown] colab_type="text" id="2CdPc75QYb5r"
# <font color='red'> **Answer:**</font>
#
# - I would like to read The Da Vinci Code and Quidditch Through the Ages, but not the other books
# - It give the exact same result with same distance value, so both of models works in same way
# + [markdown] colab_type="text" id="G6T3K3VFYb5s"
# **2.5 Add a new user (with user “UserID” = 6293) in your data. Using the two trained models in task 2.3 suggest which books should this user read if his ratings are:**
#
# French Cuisine for All: 4
#
#
# <NAME> and the Sorcerer's Stone Movie Poster Book: 5
#
#
# El Perfume: Historia De UN Asesino/Perfume : The Story of a Murderer: 1
#
# **(1. 25 points)**
#
#
# + colab={} colab_type="code" id="o-EJOEy1Yb5t"
books_encoded_title[books_encoded_title['BookTitle'] == 'French Cuisine for All']
# -
books_encoded_title[books_encoded_title['BookTitle'] == "Harry Potter and the Sorcerer's Stone Movie Poster Book"]
books_encoded_title[books_encoded_title['BookTitle'] == "El Perfume: Historia De UN Asesino/Perfume : The Story of a Murderer"]
new_df = pd.DataFrame({"UserID":[6293, 6293, 6293],
"Book-Rating":[4, 5, 1],
"BookID_Encoded":[128, 145, 116]})
new_df
books_encoded_no_dup_new = books_encoded_no_dup.append(new_df)
books_encoded_no_dup_new = books_encoded_no_dup_new.reset_index()
books_encoded_no_dup_new = books_encoded_no_dup_new.drop(columns = ['index'])
books_encoded_no_dup_new.tail()
books_matrix_new = books_encoded_no_dup_new.pivot(index = 'BookID_Encoded', columns = 'UserID', values = 'Book-Rating').fillna(0)
books_matrix_sparse_new = csr_matrix(books_matrix_new.values)
# print(f"Sparse matrix:\n{books_matrix_sparse_new}")
my_favorite = "Harry Potter and the Sorcerer's Stone Movie Poster Book"
# Recommendation based on KNN model 1
make_recommendation(
model_knn=model_knn_1, # trained model (model)
data=books_matrix_sparse_new, # sparse matrix (data)
fav_book=my_favorite, # fav_book
mapper=book_to_idx, # {book: index} (mapper)
n_recommendations=5)
# Recommendation based on KNN model 2
make_recommendation(
model_knn=model_knn_2, # trained model (model)
data=books_matrix_sparse_new, # sparse matrix (data)
fav_book=my_favorite, # fav_book
mapper=book_to_idx, # {book: index} (mapper)
n_recommendations=5)
# + [markdown] colab_type="text" id="VMyW4UlbYb5x"
# # 3. Recommender systems evaluation (1.5 points)
# + [markdown] colab_type="text" id="EINSDAbXYb5y"
# We are going to compare different methods of recommender systems by their RMSE score. One useful package that has several recommender algorithms for Python is [Surprise](https://surprise.readthedocs.io/en/stable/getting_started.html). Below we have split the books dataset into training and test and used the KNNBasic algorithm to predict the ratings for the test set using surprise.
# -
books_encoded_no_dup_new
# + colab={} colab_type="code" id="OoLm-EC1Yb5z"
from surprise import accuracy
from surprise.model_selection import train_test_split
from surprise import Reader
from surprise import Dataset
from surprise import SVD
from surprise import NormalPredictor
from surprise import KNNBasic
# The reader is necessary for surprise to interpret the ratings
reader = Reader(rating_scale=(0, 10))
# This function loads data from a pandas dataframe into surprise dataset structure
# The columns should always be ordered like this
# data = Dataset.load_from_df(df[['UserID', 'BookTitle', 'Book-Rating']], reader)
data = Dataset.load_from_df(books[['UserID', 'BookTitle', 'Book-Rating']], reader)
# Split in trainset and testset
# No need to define the label y because for surprise the last column is always the rating
trainset, testset = train_test_split(data, test_size=.25, random_state=0 )
knn = KNNBasic()
knn.fit(trainset)
predictions = knn.test(testset)
print('KNN RMSE', accuracy.rmse(predictions))
# + [markdown] colab_type="text" id="sdIaAghiYb53"
# **3.1 After taking a look at surprise documentation and the code above, follow the same steps as with KNN, and predict the ratings in test set using the NormalPredictor which predicts a random rating based on the distribution of the training set. Do the same for SVD which is a matrix factorization technique. For both of them report RMSE. (We already have imported the functions for you)**
# -
from surprise.model_selection import cross_validate
# + colab={} colab_type="code" id="VWcalcl4Yb56"
# np_result = cross_validate(NormalPredictor(), data, measures=['RMSE'], cv=2, verbose=True)
npred = NormalPredictor()
npred.fit(trainset)
pred_npred = npred.test(testset)
accuracy.rmse(pred_npred)
# -
svd = SVD()
svd.fit(trainset)
pred_SVD = svd.test(testset)
accuracy.rmse(pred_SVD)
# +
# Do benchmarking between KNNBasic, NormalPredictor, and SVD
# Reference:
# https://towardsdatascience.com/building-and-testing-recommender-systems-with-surprise-step-by-step-d4ba702ef80b
benchmark = []
# Iterate over all algorithms
for algorithm in [KNNBasic(), NormalPredictor(), SVD(), ]:
# Perform cross validation
results = cross_validate(algorithm, data, measures=['RMSE'], cv=3, verbose=False)
# Get results & append algorithm name
tmp = pd.DataFrame.from_dict(results).mean(axis=0)
tmp = tmp.append(pd.Series([str(algorithm).split(' ')[0].split('.')[-1]], index=['Algorithm']))
benchmark.append(tmp)
pd.DataFrame(benchmark).set_index('Algorithm').sort_values('test_rmse')
# + [markdown] colab_type="text" id="OjJgAOSRYb6A"
# # 4. Neural Networks (3 Points)
# + [markdown] colab_type="text" id="h5TF1ePBYb6L"
# **4.1 We are now going to build a recommender system using Neural Networks. Being this dataset is really small in terms of features you might not see great improvements but it is a good starting point to learn. Please build exactly the same neural network as we did in practice session part 3, which had the following layers:**
# - 2 Embedding
# - 2 Reshape
# - 1 Dense
#
# **Use the Neural Network you built to learn from the train data of part 3 of this homework. The column UserID should be used as input to your NN for the user embedding layer. For the books embedding layer we will use BookTitle column. Lastly, the ratings will be your target variable. Regarding the evaluation metric for the training phase use RMSE. To make your training fast you can use a batch size of 200 or above. (1.75 points)**
# + colab={} colab_type="code" id="PbuvaC1eYb6Q"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import os
import warnings
from keras.layers import Input, Embedding, Flatten, Dot, Dense, Multiply, Concatenate, Dropout, Reshape
from keras.models import Model, Sequential
from sklearn.model_selection import train_test_split
from keras.optimizers import Adam
from keras.regularizers import l2
# -
print(books_encoded_no_dup_new['UserID'].nunique())
# +
# user_enc = LabelEncoder()
# books_encoded_no_dup_new['UserID'] = user_enc.fit_transform(books_encoded_no_dup_new['UserID'].values)
# n_users = books_encoded_no_dup_new['UserID'].nunique()
# item_enc = LabelEncoder()
# books_encoded_no_dup_new['BookID_Encoded'] = item_enc.fit_transform(books_encoded_no_dup_new['BookID_Encoded'].values)
# n_books = books_encoded_no_dup_new['BookID_Encoded'].nunique()
# books_encoded_no_dup_new['Book-Rating'] = books_encoded_no_dup_new['Book-Rating'].values.astype(np.float32)
# min_rating = min(books_encoded_no_dup_new['Book-Rating'])
# max_rating = max(books_encoded_no_dup_new['Book-Rating'])
# n_users, n_books, min_rating, max_rating
n_users = books_encoded_no_dup_new['UserID'].nunique()
n_books = books_encoded_no_dup_new['BookID_Encoded'].nunique()
min_rating = min(books_encoded_no_dup_new['Book-Rating'])
max_rating = max(books_encoded_no_dup_new['Book-Rating'])
n_users, n_books, min_rating, max_rating
# +
X = books_encoded_no_dup_new[['UserID', 'BookID_Encoded']].values
y = books_encoded_no_dup_new['Book-Rating'].values
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.25, random_state=0)
X_train.shape, X_test.shape, y_train.shape, y_test.shape
# # The reader is necessary for surprise to interpret the ratings
# reader = Reader(rating_scale=(0, 10))
# # This function loads data from a pandas dataframe into surprise dataset structure
# # The columns should always be ordered like this
# # data = Dataset.load_from_df(df[['UserID', 'BookTitle', 'Book-Rating']], reader)
# data = Dataset.load_from_df(books[['UserID', 'BookTitle', 'Book-Rating']], reader)
# # Split in trainset and testset
# # No need to define the label y because for surprise the last column is always the rating
# X_train, X_test, y_train, y_test = train_test_split(data, test_size=.25, random_state=0 )
# -
n_factors = 50
X_train_array = [X_train[:, 0], X_train[:, 1]]
X_test_array = [X_test[:, 0], X_test[:, 1]]
# using only embeddings, same as previous lab session
def RecommenderV1(n_users, n_books, n_factors):
user = Input(shape=(1,))
## n_users should be added + 1 to resolve InvalidArgumentError: indices[x,x] = xxxx is not in [0, xxxx)
## Reference: https://stackoverflow.com/questions/54880279/how-to-handle-invalid-argument-error-in-keras
u = Embedding(n_users+1, n_factors, embeddings_initializer='he_normal',
embeddings_regularizer=l2(1e-6))(user)
u = Reshape((n_factors,))(u)
book = Input(shape=(1,))
m = Embedding(n_books, n_factors, embeddings_initializer='he_normal',
embeddings_regularizer=l2(1e-6))(book)
m = Reshape((n_factors,))(m)
x = Dot(axes=1)([u, m])
model = Model(inputs=[user, book], outputs=x)
opt = Adam(lr=0.001)
model.compile(loss='mean_squared_error', optimizer=opt)
return model
# +
from keras.layers import Concatenate, Dense, Dropout
from keras.layers import Add, Activation, Lambda
class EmbeddingLayer:
def __init__(self, n_items, n_factors):
self.n_items = n_items
self.n_factors = n_factors
def __call__(self, x):
x = Embedding(self.n_items, self.n_factors, embeddings_initializer='he_normal',
embeddings_regularizer=l2(1e-6))(x)
x = Reshape((self.n_factors,))(x)
return x
def RecommenderV2(n_users, n_books, n_factors, min_rating, max_rating):
user = Input(shape=(1,))
u = EmbeddingLayer(n_users+1, n_factors)(user)
ub = EmbeddingLayer(n_users+1, 1)(user) # 1st improvement
book = Input(shape=(1,))
m = EmbeddingLayer(n_books, n_factors)(book)
mb = EmbeddingLayer(n_books, 1)(book) # 1snd improvement
x = Dot(axes=1)([u, m])
x = Add()([x, ub, mb])
x = Activation('sigmoid')(x) # 2nd improvement
x = Lambda(lambda x: x * (max_rating - min_rating) + min_rating)(x)
model = Model(inputs=[user, book], outputs=x)
opt = Adam(lr=0.001)
model.compile(loss='mean_squared_error', optimizer=opt)
return model
def RecommenderNet(n_users, n_books, n_factors, min_rating, max_rating):
user = Input(shape=(1,))
u = EmbeddingLayer(n_users+1, n_factors)(user)
book = Input(shape=(1,))
m = EmbeddingLayer(n_books, n_factors)(book)
x = Concatenate()([u, m])
x = Dropout(0.05)(x)
x = Dense(10, kernel_initializer='he_normal')(x)
x = Activation('relu')(x)
x = Dropout(0.5)(x)
x = Dense(1, kernel_initializer='he_normal')(x)
x = Activation('sigmoid')(x)
x = Lambda(lambda x: x * (max_rating - min_rating) + min_rating)(x)
model = Model(inputs=[user, book], outputs=x)
opt = Adam(lr=0.001)
model.compile(loss='mean_squared_error', optimizer=opt)
return model
# +
def compile_fit_plot(model,num):
model.summary()
model.compile(loss='mse', optimizer='adam', metrics=['mse'])
# Set batch_size to 200 or more, to make training process faster
history = model.fit(x=X_train_array, y=y_train, batch_size=250, epochs=10,
verbose=1, validation_data=(X_test_array, y_test))
# Plot training & validation accuracy values
plt.plot(history.history['mse'])
plt.plot(history.history['val_mse'])
plt.title('Model MSE')
plt.ylabel('mean_squared_error')
plt.xlabel('Epoch')
plt.legend(['Train', 'Test'], loc='upper left')
plt.show()
# Plot training & validation loss values
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('Model loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.legend(['Train', 'Test'], loc='upper left')
plt.show()
import math
# Show the best validation RMSE
min_val_loss, idx = min((val, idx) for (idx, val) in enumerate(history.history['val_loss']))
print(f"\nModel: {num},\nMinimum RMSE at epoch: {idx+1} = {math.sqrt(min_val_loss)}")
#print ('\nModel: {:d}\n'.format(num),'Minimum RMSE at epoch', '{:d}'.format(idx+1), '=', '{:.4f}'.format(math.sqrt(min_val_loss)))
def predict_recommend(model,test_user ):
# Function to predict the ratings given User ID and Book ID
def predict_rating1(user_id, item_id):
return model.predict([np.array([user_id-1]), np.array([item_id-1])])[0][0]
# Function to predict the ratings given User ID and Book ID
def predict_rating2(user_id, item_id):
if item_id<=336:
prediction = model.predict([np.array([user_id-1]), np.array([item_id])])[0][0]
return prediction
TEST_USER = test_user
user_ratings = books_encoded_no_dup_new[books_encoded_no_dup_new['UserID'] == TEST_USER][['UserID', 'BookID_Encoded', 'Book-Rating']]
user_ratings['prediction'] = user_ratings.apply(lambda x: predict_rating1(TEST_USER, x['BookID_Encoded']), axis=1)
user_ratings.sort_values(by='Book-Rating',
ascending=False).merge(books_encoded_title,
on='BookID_Encoded',
how='inner',
suffixes=['_u', '_m']).head(20)
recommendations = books_encoded_no_dup_new[books_encoded_no_dup_new['BookID_Encoded'].isin(user_ratings['BookID_Encoded']) == False][['BookID_Encoded']].drop_duplicates()
recommendations['prediction'] = recommendations.apply(lambda x: predict_rating2(TEST_USER, x['BookID_Encoded']), axis=1)
recommendations.sort_values(by='prediction',
ascending=False).merge(books_encoded_title,
on='BookID_Encoded',
how='inner',
suffixes=['_u', '_m']).head(20)
return user_ratings, recommendations
# +
# Define constants
#K_FACTORS = 100 # The number of dimensional embeddings for books and users
# TEST_USER = 6293
model1 = RecommenderV1(n_users, n_books, n_factors)
model2 = RecommenderV2(n_users, n_books, n_factors, min_rating, max_rating)
model3 = RecommenderNet(n_users, n_books, n_factors, min_rating, max_rating)
# compile_fit_plot(model1, 1)
# user_ratings_1, recommendations_1 = predict_recommend(model1,TEST_USER)
# + [markdown] colab_type="text" id="HsmPj7Wq1cyS"
# **4.2 Plot the RMSE values during the training phase, as well as the model loss. Report the best RMSE. Is it better than the RMSE from the models we built in Section 2 and 3 ? (0.75 points)**
# + colab={} colab_type="code" id="JCJFqfDm1-HA"
compile_fit_plot(model1, 1)
# -
compile_fit_plot(model2, 2)
compile_fit_plot(model3, 3)
# Our `model3` using Keras model gives better RMSE value compare to previous model using KNNBasic() and SVD()
# +
# intialise data of lists.
data = {'Model':['Keras', 'SVD', 'KNNBasic'], 'RMSE value':[3.84, 3.85, 4.13]}
# Create DataFrame
rmse_comparison = pd.DataFrame(data)
# Print the output.
print("Comparison Table \n", rmse_comparison)
# + [markdown] colab_type="text" id="LVLaC5K11-fN"
# **4.3 Use your trained model to recommend books for user with ID 6293. (0.5 points)**
# -
# We'll using `model3` for our model prediction, because it gives the best value of RMSE
user_ratings_3, recommendations_3 = predict_recommend(model3,6293)
user_ratings_3
# recommendations_3
result = recommendations_3.sort_values(by=['prediction'], ascending = False).head(5)
result
# +
# get reverse mapper
mapper = book_to_idx
reverse_mapper = {v: k for k, v in mapper.items()}
# print recommendations
print('Book recommendations for user 6293:\n')
for i in result['BookID_Encoded']:
print("- ", reverse_mapper[i])
# + [markdown] colab_type="text" id="zwpOi51caTUp"
# ## How long did it take you to solve the homework?
#
# * Please answer as precisely as you can. It does not affect your points or grade in any way. It is okay, if it took 0.5 hours or 24 hours. The collected information will be used to improve future homeworks.
#
# <font color='red'> **Answer:**</font>
#
# **<font color='red'>(please change X in the next cell into your estimate)</font>**
#
# 15 hours
#
# ## What is the level of difficulty for this homework?
# you can put only number between $0:10$ ($0:$ easy, $10:$ difficult)
#
# <font color='red'> **Answer:**</font>
# 7
# -
| HW4/Homework_04_Enlik.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.6 64-bit
# language: python
# name: python3
# ---
import torch, torchvision
model = torchvision.models.resnet18(pretrained=True)
data = torch.rand(1, 3, 64, 64)
labels = torch.rand(1, 1000)
prediction = model(data) # forward pass
loss = (prediction - labels).sum()
loss.backward() # backward pass
optim = torch.optim.SGD(model.parameters(), lr=1e-2, momentum=0.9)
optim.step()
a = torch.tensor([2., 3.], requires_grad=True)
b = torch.tensor([6., 4.], requires_grad=True)
Q = 3*a**3 - b**2
external_grad = torch.tensor([1., 1.])
Q.backward(gradient=external_grad)
print(9*a**2 == a.grad)
print(-2*b == b.grad)
print(Q)
| 2-autograd.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="HZiF5lbumA7j"
# ##### Copyright 2020 The TensorFlow Authors.
# + cellView="form" id="KsOkK8O69PyT"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="eNj0_BTFk479"
# # 既製の TF Lattice モデル
# + [markdown] id="T3qE8F5toE28"
# <table class="tfo-notebook-buttons" align="left">
# <td><a target="_blank" href="https://www.tensorflow.org/lattice/tutorials/premade_models"><img src="https://www.tensorflow.org/images/tf_logo_32px.png"> TensorFlow.orgで表示</a></td>
# <td><a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/ja/lattice/tutorials/premade_models.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png"> Google Colab で実行</a></td>
# <td><a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/ja/lattice/tutorials/premade_models.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png">GitHub でソースを表示{</a></td>
# <td><a href="https://storage.googleapis.com/tensorflow_docs/docs-l10n/site/ja/lattice/tutorials/premade_models.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png">ノートブックをダウンロード/a0}</a></td>
# </table>
# + [markdown] id="HEuRMAUOlFZa"
# ## 概要
#
# 既製のモデルは、典型的な使用事例向けの TFL `tf.keras.model` インスタンスを素早く簡単位構築する方法です。このガイドでは、既製の TFL モデルを構築し、トレーニングやテストを行うために必要な手順を説明します。
# + [markdown] id="f2--Yq21lhRe"
# ## セットアップ
#
# TF Lattice パッケージをインストールします。
# + id="XizqBCyXky4y"
#@test {"skip": true}
# !pip install tensorflow-lattice pydot
# + [markdown] id="2oKJPy5tloOB"
# 必要なパッケージをインポートします。
# + id="9wZWJJggk4al"
import tensorflow as tf
import copy
import logging
import numpy as np
import pandas as pd
import sys
import tensorflow_lattice as tfl
logging.disable(sys.maxsize)
# + [markdown] id="kpJJSS7YmLbG"
# UCI Statlog(心臓)データセットをダウンロードします。
# + id="AYTcybljmQJm"
csv_file = tf.keras.utils.get_file(
'heart.csv', 'http://storage.googleapis.com/applied-dl/heart.csv')
df = pd.read_csv(csv_file)
train_size = int(len(df) * 0.8)
train_dataframe = df[:train_size]
test_dataframe = df[train_size:]
df.head()
# + [markdown] id="ODe0oavWmtAi"
# 特徴量とラベルを抽出して、テンソルに変換します。
# + id="3ae-Mx-PnGGL"
# Features:
# - age
# - sex
# - cp chest pain type (4 values)
# - trestbps resting blood pressure
# - chol serum cholestoral in mg/dl
# - fbs fasting blood sugar > 120 mg/dl
# - restecg resting electrocardiographic results (values 0,1,2)
# - thalach maximum heart rate achieved
# - exang exercise induced angina
# - oldpeak ST depression induced by exercise relative to rest
# - slope the slope of the peak exercise ST segment
# - ca number of major vessels (0-3) colored by flourosopy
# - thal 3 = normal; 6 = fixed defect; 7 = reversable defect
#
# This ordering of feature names will be the exact same order that we construct
# our model to expect.
feature_names = [
'age', 'sex', 'cp', 'chol', 'fbs', 'trestbps', 'thalach', 'restecg',
'exang', 'oldpeak', 'slope', 'ca', 'thal'
]
feature_name_indices = {name: index for index, name in enumerate(feature_names)}
# This is the vocab list and mapping we will use for the 'thal' categorical
# feature.
thal_vocab_list = ['normal', 'fixed', 'reversible']
thal_map = {category: i for i, category in enumerate(thal_vocab_list)}
# + id="x5p3OgC-m4TW"
# Custom function for converting thal categories to buckets
def convert_thal_features(thal_features):
# Note that two examples in the test set are already converted.
return np.array([
thal_map[feature] if feature in thal_vocab_list else feature
for feature in thal_features
])
# Custom function for extracting each feature.
def extract_features(dataframe,
label_name='target',
feature_names=feature_names):
features = []
for feature_name in feature_names:
if feature_name == 'thal':
features.append(
convert_thal_features(dataframe[feature_name].values).astype(float))
else:
features.append(dataframe[feature_name].values.astype(float))
labels = dataframe[label_name].values.astype(float)
return features, labels
# + id="7DgoAgkIm8tr"
train_xs, train_ys = extract_features(train_dataframe)
test_xs, test_ys = extract_features(test_dataframe)
# + id="qcguGFRcFgCQ"
# Let's define our label minimum and maximum.
min_label, max_label = float(np.min(train_ys)), float(np.max(train_ys))
# Our lattice models may have predictions above 1.0 due to numerical errors.
# We can subtract this small epsilon value from our output_max to make sure we
# do not predict values outside of our label bound.
numerical_error_epsilon = 1e-5
# + [markdown] id="oyOrtol7mW9r"
# このガイドのトレーニングに使用されるデフォルト値を設定します。
# + id="ns8pH2AnmgAC"
LEARNING_RATE = 0.01
BATCH_SIZE = 128
NUM_EPOCHS = 500
PREFITTING_NUM_EPOCHS = 10
# + [markdown] id="Ix2elMrGmiWX"
# ## 特徴量の構成
#
# 特徴量の較正と特徴量あたりの構成は <a>tfl.configs.FeatureConfig</a> によって設定します。特徴量の構成には、単調性制約、特徴量あたりの正則化(<a>tfl.configs.RegularizerConfig</a> を参照)、および格子モデルの格子のサイズが含まれます。
#
# モデルが認識する必要のあるすべての特徴量に対し、完全な特徴量の構成を指定する必要があります。指定されていない場合、モデルは特徴量の存在を認識できません。
# + [markdown] id="WLSfZ5G7-YT_"
# ### 分位数を計算する
#
# `tfl.configs.FeatureConfig` の `pwl_calibration_input_keypoints` のデフォルト設定は 'quantiles' ですが、既製のモデルについては、入力キーポイントを手動で定義する必要があります。これを行うには、まず、分位数を計算するためのヘルパー関数を独自に定義します。
# + id="-LqqEp3k-06d"
def compute_quantiles(features,
num_keypoints=10,
clip_min=None,
clip_max=None,
missing_value=None):
# Clip min and max if desired.
if clip_min is not None:
features = np.maximum(features, clip_min)
features = np.append(features, clip_min)
if clip_max is not None:
features = np.minimum(features, clip_max)
features = np.append(features, clip_max)
# Make features unique.
unique_features = np.unique(features)
# Remove missing values if specified.
if missing_value is not None:
unique_features = np.delete(unique_features,
np.where(unique_features == missing_value))
# Compute and return quantiles over unique non-missing feature values.
return np.quantile(
unique_features,
np.linspace(0., 1., num=num_keypoints),
interpolation='nearest').astype(float)
# + [markdown] id="<KEY>"
# ### 特徴量の構成を定義する
#
# 分位数を計算できるようになったので、モデルが入力として使用する各特徴量に対する特徴量の構成を定義します。
# + id="8y27RmHIrSBn"
# Feature configs are used to specify how each feature is calibrated and used.
feature_configs = [
tfl.configs.FeatureConfig(
name='age',
lattice_size=3,
monotonicity='increasing',
# We must set the keypoints manually.
pwl_calibration_num_keypoints=5,
pwl_calibration_input_keypoints=compute_quantiles(
train_xs[feature_name_indices['age']],
num_keypoints=5,
clip_max=100),
# Per feature regularization.
regularizer_configs=[
tfl.configs.RegularizerConfig(name='calib_wrinkle', l2=0.1),
],
),
tfl.configs.FeatureConfig(
name='sex',
num_buckets=2,
),
tfl.configs.FeatureConfig(
name='cp',
monotonicity='increasing',
# Keypoints that are uniformly spaced.
pwl_calibration_num_keypoints=4,
pwl_calibration_input_keypoints=np.linspace(
np.min(train_xs[feature_name_indices['cp']]),
np.max(train_xs[feature_name_indices['cp']]),
num=4),
),
tfl.configs.FeatureConfig(
name='chol',
monotonicity='increasing',
# Explicit input keypoints initialization.
pwl_calibration_input_keypoints=[126.0, 210.0, 247.0, 286.0, 564.0],
# Calibration can be forced to span the full output range by clamping.
pwl_calibration_clamp_min=True,
pwl_calibration_clamp_max=True,
# Per feature regularization.
regularizer_configs=[
tfl.configs.RegularizerConfig(name='calib_hessian', l2=1e-4),
],
),
tfl.configs.FeatureConfig(
name='fbs',
# Partial monotonicity: output(0) <= output(1)
monotonicity=[(0, 1)],
num_buckets=2,
),
tfl.configs.FeatureConfig(
name='trestbps',
monotonicity='decreasing',
pwl_calibration_num_keypoints=5,
pwl_calibration_input_keypoints=compute_quantiles(
train_xs[feature_name_indices['trestbps']], num_keypoints=5),
),
tfl.configs.FeatureConfig(
name='thalach',
monotonicity='decreasing',
pwl_calibration_num_keypoints=5,
pwl_calibration_input_keypoints=compute_quantiles(
train_xs[feature_name_indices['thalach']], num_keypoints=5),
),
tfl.configs.FeatureConfig(
name='restecg',
# Partial monotonicity: output(0) <= output(1), output(0) <= output(2)
monotonicity=[(0, 1), (0, 2)],
num_buckets=3,
),
tfl.configs.FeatureConfig(
name='exang',
# Partial monotonicity: output(0) <= output(1)
monotonicity=[(0, 1)],
num_buckets=2,
),
tfl.configs.FeatureConfig(
name='oldpeak',
monotonicity='increasing',
pwl_calibration_num_keypoints=5,
pwl_calibration_input_keypoints=compute_quantiles(
train_xs[feature_name_indices['oldpeak']], num_keypoints=5),
),
tfl.configs.FeatureConfig(
name='slope',
# Partial monotonicity: output(0) <= output(1), output(1) <= output(2)
monotonicity=[(0, 1), (1, 2)],
num_buckets=3,
),
tfl.configs.FeatureConfig(
name='ca',
monotonicity='increasing',
pwl_calibration_num_keypoints=4,
pwl_calibration_input_keypoints=compute_quantiles(
train_xs[feature_name_indices['ca']], num_keypoints=4),
),
tfl.configs.FeatureConfig(
name='thal',
# Partial monotonicity:
# output(normal) <= output(fixed)
# output(normal) <= output(reversible)
monotonicity=[('normal', 'fixed'), ('normal', 'reversible')],
num_buckets=3,
# We must specify the vocabulary list in order to later set the
# monotonicities since we used names and not indices.
vocabulary_list=thal_vocab_list,
),
]
# + [markdown] id="-XuAnP_-vyK6"
# 次に、カスタム語彙(上記の 'thal' など)を使用した特徴量に単調性を適切に設定していることを確認する必要があります。
# + id="ZIn2-EfGv--m"
tfl.premade_lib.set_categorical_monotonicities(feature_configs)
# + [markdown] id="Mx50YgWMcxC4"
# ## 較正済みの線形モデル
#
# 既製の TFL モデルを構築するには、<a>tfl.configs</a> からモデル構成を構築します。較正された線形モデルは、<a>tfl.configs.CalibratedLinearConfig</a> を使用して構築されます。これは、ピースごとに線形で分類上の較正を入力特徴量に適用し、その後で、線形の組み合わせとオプションの出力ピース単位線形較正を適用します。出力較正を使用する場合、または出力境界が指定されている場合、線形レイヤーは加重平均を較正済みの入力に適用します。
#
# この例では、最初の 5 つの特徴量で較正済みの線形モデルを作成します。
# + id="UvMDJKqTc1vC"
# Model config defines the model structure for the premade model.
linear_model_config = tfl.configs.CalibratedLinearConfig(
feature_configs=feature_configs[:5],
use_bias=True,
# We must set the output min and max to that of the label.
output_min=min_label,
output_max=max_label,
output_calibration=True,
output_calibration_num_keypoints=10,
output_initialization=np.linspace(min_label, max_label, num=10),
regularizer_configs=[
# Regularizer for the output calibrator.
tfl.configs.RegularizerConfig(name='output_calib_hessian', l2=1e-4),
])
# A CalibratedLinear premade model constructed from the given model config.
linear_model = tfl.premade.CalibratedLinear(linear_model_config)
# Let's plot our model.
tf.keras.utils.plot_model(linear_model, show_layer_names=False, rankdir='LR')
# + [markdown] id="3MC3-AyX00-A"
# ここで、ほかの [tf.keras.Model](https://www.tensorflow.org/api_docs/python/tf/keras/Model) と同様に、モデルをコンパイルしてデータに適合させます。
# + id="hPlEK-yG1B-U"
linear_model.compile(
loss=tf.keras.losses.BinaryCrossentropy(),
metrics=[tf.keras.metrics.AUC()],
optimizer=tf.keras.optimizers.Adam(LEARNING_RATE))
linear_model.fit(
train_xs[:5],
train_ys,
epochs=NUM_EPOCHS,
batch_size=BATCH_SIZE,
verbose=False)
# + [markdown] id="OG2ua0MGAkoi"
# モデルをトレーニングしたら、テストデータを使ってモデルを評価することができます。
# + id="HybGTvXxAoxV"
print('Test Set Evaluation...')
print(linear_model.evaluate(test_xs, test_ys))
# + [markdown] id="jAAJK-wlc15S"
# ## 較正済みの格子モデル
#
# 較正済みの格子モデルは、<a>tfl.configs.CalibratedLatticeConfig</a> を使って構築されます。較正済みの格子モデルは、入力特徴量にピース単位の線形と分類上の較正を適用し、その後で、格子モデルとオプションの出力ピース単位線形較正を適用します。
#
# この例では、最初の 5 つの特徴量で較正済みの格子モデルを作成します。
# + id="u7gNcrMtc4Lp"
# This is a calibrated lattice model: inputs are calibrated, then combined
# non-linearly using a lattice layer.
lattice_model_config = tfl.configs.CalibratedLatticeConfig(
feature_configs=feature_configs[:5],
output_min=min_label,
output_max=max_label - numerical_error_epsilon,
output_initialization=[min_label, max_label],
regularizer_configs=[
# Torsion regularizer applied to the lattice to make it more linear.
tfl.configs.RegularizerConfig(name='torsion', l2=1e-2),
# Globally defined calibration regularizer is applied to all features.
tfl.configs.RegularizerConfig(name='calib_hessian', l2=1e-2),
])
# A CalibratedLattice premade model constructed from the given model config.
lattice_model = tfl.premade.CalibratedLattice(lattice_model_config)
# Let's plot our model.
tf.keras.utils.plot_model(lattice_model, show_layer_names=False, rankdir='LR')
# + [markdown] id="nmc3TUIIGGoH"
# 前と同じように、モデルをコンパイルし、適合して評価します。
# + id="vIjOQGD2Gp_Z"
lattice_model.compile(
loss=tf.keras.losses.BinaryCrossentropy(),
metrics=[tf.keras.metrics.AUC()],
optimizer=tf.keras.optimizers.Adam(LEARNING_RATE))
lattice_model.fit(
train_xs[:5],
train_ys,
epochs=NUM_EPOCHS,
batch_size=BATCH_SIZE,
verbose=False)
print('Test Set Evaluation...')
print(lattice_model.evaluate(test_xs[:5], test_ys))
# + [markdown] id="bx74CD4Cc4T3"
# ## 較正済みの格子アンサンブルモデル
#
# 特徴量の数が大きい場合、アンサンブルモデルを使用できます。アンサンブルモデルは、単一の大型の格子を作成する代わりに、特徴量のサブセットにより小さな格子を作成し、その出力を平均化します。アンサンブル格子モデルは、<a>tfl.configs.CalibratedLatticeEnsembleConfig</a> を使って作成します。較正済み格子アンサンブルモデルは、特徴量にピース単位の線形および分類上の較正を適用し、その後で、格子モデルのアンサンブルとオプションの出力ピース単位線形較正を適用します。
# + [markdown] id="mbg4lsKqnEkV"
# ### 明示的な格子アンサンブルの初期化
#
# 格子にどの特徴量サブセットをフィードするのかがわかっている場合は、特徴量の名前を使用して格子を明示的に設定することができます。この例では、5 つの格子と格子当たり 3 つの特徴量を使用して、較正済みの格子アンサンブルモデルを作成します。
# + id="yu8Twg8mdJ18"
# This is a calibrated lattice ensemble model: inputs are calibrated, then
# combined non-linearly and averaged using multiple lattice layers.
explicit_ensemble_model_config = tfl.configs.CalibratedLatticeEnsembleConfig(
feature_configs=feature_configs,
lattices=[['trestbps', 'chol', 'ca'], ['fbs', 'restecg', 'thal'],
['fbs', 'cp', 'oldpeak'], ['exang', 'slope', 'thalach'],
['restecg', 'age', 'sex']],
num_lattices=5,
lattice_rank=3,
output_min=min_label,
output_max=max_label - numerical_error_epsilon,
output_initialization=[min_label, max_label])
# A CalibratedLatticeEnsemble premade model constructed from the given
# model config.
explicit_ensemble_model = tfl.premade.CalibratedLatticeEnsemble(
explicit_ensemble_model_config)
# Let's plot our model.
tf.keras.utils.plot_model(
explicit_ensemble_model, show_layer_names=False, rankdir='LR')
# + [markdown] id="PJYR0i6MMDyh"
# 前と同じように、モデルをコンパイルし、適合して評価します。
# + id="capt98IOMHEm"
explicit_ensemble_model.compile(
loss=tf.keras.losses.BinaryCrossentropy(),
metrics=[tf.keras.metrics.AUC()],
optimizer=tf.keras.optimizers.Adam(LEARNING_RATE))
explicit_ensemble_model.fit(
train_xs, train_ys, epochs=NUM_EPOCHS, batch_size=BATCH_SIZE, verbose=False)
print('Test Set Evaluation...')
print(explicit_ensemble_model.evaluate(test_xs, test_ys))
# + [markdown] id="VnI70C9gdKQw"
# ### ランダムな格子アンサンブル
#
# 格子にどの特徴量サブセットをフィードしていいのか不明な場合は、別のオプションとして、各格子にランダムな特徴量サブセットを使用する方法があります。この例では、5 つの格子と格子当たり 3 つの特徴量を使用して、較正済みの格子アンサンブルモデルを作成します。
# + id="7EhWrQaPIXj8"
# This is a calibrated lattice ensemble model: inputs are calibrated, then
# combined non-linearly and averaged using multiple lattice layers.
random_ensemble_model_config = tfl.configs.CalibratedLatticeEnsembleConfig(
feature_configs=feature_configs,
lattices='random',
num_lattices=5,
lattice_rank=3,
output_min=min_label,
output_max=max_label - numerical_error_epsilon,
output_initialization=[min_label, max_label],
random_seed=42)
# Now we must set the random lattice structure and construct the model.
tfl.premade_lib.set_random_lattice_ensemble(random_ensemble_model_config)
# A CalibratedLatticeEnsemble premade model constructed from the given
# model config.
random_ensemble_model = tfl.premade.CalibratedLatticeEnsemble(
random_ensemble_model_config)
# Let's plot our model.
tf.keras.utils.plot_model(
random_ensemble_model, show_layer_names=False, rankdir='LR')
# + [markdown] id="sbxcIF0PJUDc"
# 前と同じように、モデルをコンパイルし、適合して評価します。
# + id="w0YdCDyGJY1G"
random_ensemble_model.compile(
loss=tf.keras.losses.BinaryCrossentropy(),
metrics=[tf.keras.metrics.AUC()],
optimizer=tf.keras.optimizers.Adam(LEARNING_RATE))
random_ensemble_model.fit(
train_xs, train_ys, epochs=NUM_EPOCHS, batch_size=BATCH_SIZE, verbose=False)
print('Test Set Evaluation...')
print(random_ensemble_model.evaluate(test_xs, test_ys))
# + [markdown] id="ZhJWe7fZIs4-"
# ### RTL レイヤーのランダムな格子アンサンブル
#
# ランダムな格子アンサンブルを使用する場合、モデルが単一の `tfl.layers.RTL` レイヤーを使用するように設定することができます。`tfl.layers.RTL` は単調性制約のみをサポートしており、すべての特徴量と特徴量ごとの正則化において同じ格子サイズが必要です。`tfl.layers.RTL` レイヤーを使用すると、別の `tfl.layers.Lattice` インスタンスを使用する場合よりもさらに大きなアンサンブルにスケーリングすることができます。
#
# この例では、5 つの格子と格子当たり 3 つの特徴量を使用して、較正済みの格子アンサンブルモデルを作成します。
# + id="0PC9oRFYJMF_"
# Make sure our feature configs have the same lattice size, no per-feature
# regularization, and only monotonicity constraints.
rtl_layer_feature_configs = copy.deepcopy(feature_configs)
for feature_config in rtl_layer_feature_configs:
feature_config.lattice_size = 2
feature_config.unimodality = 'none'
feature_config.reflects_trust_in = None
feature_config.dominates = None
feature_config.regularizer_configs = None
# This is a calibrated lattice ensemble model: inputs are calibrated, then
# combined non-linearly and averaged using multiple lattice layers.
rtl_layer_ensemble_model_config = tfl.configs.CalibratedLatticeEnsembleConfig(
feature_configs=rtl_layer_feature_configs,
lattices='rtl_layer',
num_lattices=5,
lattice_rank=3,
output_min=min_label,
output_max=max_label - numerical_error_epsilon,
output_initialization=[min_label, max_label],
random_seed=42)
# A CalibratedLatticeEnsemble premade model constructed from the given
# model config. Note that we do not have to specify the lattices by calling
# a helper function (like before with random) because the RTL Layer will take
# care of that for us.
rtl_layer_ensemble_model = tfl.premade.CalibratedLatticeEnsemble(
rtl_layer_ensemble_model_config)
# Let's plot our model.
tf.keras.utils.plot_model(
rtl_layer_ensemble_model, show_layer_names=False, rankdir='LR')
# + [markdown] id="yWdxZpS0JWag"
# 前と同じように、モデルをコンパイルし、適合して評価します。
# + id="HQdkkWwqJW8p"
rtl_layer_ensemble_model.compile(
loss=tf.keras.losses.BinaryCrossentropy(),
metrics=[tf.keras.metrics.AUC()],
optimizer=tf.keras.optimizers.Adam(LEARNING_RATE))
rtl_layer_ensemble_model.fit(
train_xs, train_ys, epochs=NUM_EPOCHS, batch_size=BATCH_SIZE, verbose=False)
print('Test Set Evaluation...')
print(rtl_layer_ensemble_model.evaluate(test_xs, test_ys))
# + [markdown] id="A61VpAl8uOiT"
# ### Crystals 格子アンサンブル
#
# 既製は、[Crystals](https://papers.nips.cc/paper/6377-fast-and-flexible-monotonic-functions-with-ensembles-of-lattices) と呼ばれるヒューリスティックな特徴量配置アルゴリズムも提供しています。Crystals アルゴリズムはまず、ペアでの特徴量の相互作用を推定する事前適合モデルをトレーニングします。次に、同じ格子内により多くの非線形相互作用を持つ特徴量が存在するように、最終的なアンサンブルを配置します。
#
# 既製のライブラリは、事前適合モデルの構成を構築し、Crystals 構成を抽出するためのヘルパー関数を提供しています。事前適合モデルは完全にトレーニングされている必要はないため、数エポックのみで十分だといえます。
#
# この例では、5 つの格子と格子当たり 3 つの特徴量を使用して、較正済みの格子アンサンブルモデルを作成します。
# + id="yT5eiknCu9sj"
# This is a calibrated lattice ensemble model: inputs are calibrated, then
# combines non-linearly and averaged using multiple lattice layers.
crystals_ensemble_model_config = tfl.configs.CalibratedLatticeEnsembleConfig(
feature_configs=feature_configs,
lattices='crystals',
num_lattices=5,
lattice_rank=3,
output_min=min_label,
output_max=max_label - numerical_error_epsilon,
output_initialization=[min_label, max_label],
random_seed=42)
# Now that we have our model config, we can construct a prefitting model config.
prefitting_model_config = tfl.premade_lib.construct_prefitting_model_config(
crystals_ensemble_model_config)
# A CalibratedLatticeEnsemble premade model constructed from the given
# prefitting model config.
prefitting_model = tfl.premade.CalibratedLatticeEnsemble(
prefitting_model_config)
# We can compile and train our prefitting model as we like.
prefitting_model.compile(
loss=tf.keras.losses.BinaryCrossentropy(),
optimizer=tf.keras.optimizers.Adam(LEARNING_RATE))
prefitting_model.fit(
train_xs,
train_ys,
epochs=PREFITTING_NUM_EPOCHS,
batch_size=BATCH_SIZE,
verbose=False)
# Now that we have our trained prefitting model, we can extract the crystals.
tfl.premade_lib.set_crystals_lattice_ensemble(crystals_ensemble_model_config,
prefitting_model_config,
prefitting_model)
# A CalibratedLatticeEnsemble premade model constructed from the given
# model config.
crystals_ensemble_model = tfl.premade.CalibratedLatticeEnsemble(
crystals_ensemble_model_config)
# Let's plot our model.
tf.keras.utils.plot_model(
crystals_ensemble_model, show_layer_names=False, rankdir='LR')
# + [markdown] id="PRLU1z-216h8"
# As before, we compile, fit, and evaluate our model.
# + id="U73On3v91-Qq"
crystals_ensemble_model.compile(
loss=tf.keras.losses.BinaryCrossentropy(),
metrics=[tf.keras.metrics.AUC()],
optimizer=tf.keras.optimizers.Adam(LEARNING_RATE))
crystals_ensemble_model.fit(
train_xs, train_ys, epochs=NUM_EPOCHS, batch_size=BATCH_SIZE, verbose=False)
print('Test Set Evaluation...')
print(crystals_ensemble_model.evaluate(test_xs, test_ys))
| site/ja/lattice/tutorials/premade_models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.5 64-bit (''myenv'': conda)'
# name: python395jvsc74a57bd003dd13d48678367c8c9c8d2bc4e4058efaffff37d2a70d9886c86b6aa2328a71
# ---
import torch
from torch.cuda import is_available
from torch.optim import Adam
from torch.nn import MSELoss, BCELoss
import numpy as np
import matplotlib.pyplot as plt
import cv2
from torch.utils.data import Dataset, DataLoader
from utils import ims, imshow
from conv3d import Conv3D, Check, PEI
# ### The outputs after each layer:
# `dim = (batch_size, channels, frames, height, width)`
# * Conv3d layer 1: `(1, 16, 1, 160, 120)`
#
# * Conv3d layer 2: `(1, 32, 1, 160, 120)`
#
# * Conv3d layer 3: `(1, 16, 1, 160, 120)`
#
# * Conv3d layer 4: `(1, 1, 1, 160, 120)`
#
torch.cuda.is_available()
model = Conv3D()
device = 'cuda:0'
print(device)
model.to(device)
from pytictoc import TicToc
t = TicToc()
t.tic()
dataset = PEI(3, angle=0, keyposes=[4,5,6])
t.toc()
len(dataset)
train_dl = DataLoader(dataset,batch_size=1,shuffle=True)
#Loss
criterion = MSELoss()
#Optimizer
optimizer = Adam(model.parameters(), lr=0.001)
# ## Training
# +
# from tqdm import tqdm
# +
#Epochs
n_epochs = 5
for epoch in range(1, n_epochs+1):
# monitor training loss
train_loss = 0.0
#Training
for images,y in train_dl:
images,y = images.to(device), y.to(device)
# print('images.shape',images.shape)
optimizer.zero_grad()
out = model(images)
# print(out.view(out.shape[3],out.shape[4]).shape,y.view)
loss = criterion(out.view(out.shape[3],out.shape[4]), y.view(y.shape[1],y.shape[2]))
loss.backward()
optimizer.step()
train_loss += loss.item()
train_loss = train_loss/len(train_dl)
print('Epoch: {} \tTraining Loss: {:.6f}'.format(epoch, train_loss))
# -
# ## Check
# `240 images (80 samples from 15 experiments, 3 frames each)`
def ims(model_out,actual):
fig, axes = plt.subplots(nrows=1, ncols=2, sharex=True, sharey=True, figsize=(12,4))
axes[0].imshow(model_out,cmap='gray',vmin=0,vmax=1)
axes[0].set_title('Model')
axes[1].imshow(actual,cmap='gray',vmin=0,vmax=1)
axes[1].set_title('Average')
plt.show()
# + tags=[]
for i,data in enumerate(train_dl):
if i==5:
break
output = model(data[0].to(device=device))
data[0] = data[0].reshape(data[0].shape[2],160,120)
output = output.view(output.shape[2],160, 120)
# print(output.shape)
output = output.cpu().detach().numpy()
output = np.mean(output,axis=0)
ims(output,data[1].reshape(160,120))
# -
# ### Check one sample
# # TEST DATA: ANGLE 90, KEYPOSE 6
test_ds = PEI(1,angle=90,keyposes=[6])
test_dl = DataLoader(test_ds,batch_size=1)
# +
dataiter = iter(test_dl)
images, y = dataiter.next()
# images = images.to('cpu',dtype=torch.float)
# print(type(images),images.shape)
#Sample outputs
output = model(images.to(device=device))
images = images.cpu().numpy().reshape(images.shape[2],160,120)
output = output.view(output.shape[2],160, 120)
output = output.cpu().detach().numpy()
output = np.mean(output,axis=0)
ims(output,y.view(160,120))
# -
fig,ax = plt.subplots(nrows=1,ncols=3)
ax[0].imshow(images[0],cmap='gray')
ax[1].imshow(images[1],cmap='gray')
ax[2].imshow(images[2],cmap='gray')
fig.show()
images.shape
| .ipynb_checkpoints/test-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] pycharm={"is_executing": false, "name": "#%% md\n"}
# ## Exploratory data analysis
#
# This notebook provides info about the distribution of the data collected as well as how it has been gathered.
# -
# ### Raw data collection
#
# The task to be performed requires a big corpus of Java source code and the choice made is to clone and compile a number
# of popular open source projects to perform static analysis and label source files as vulnerable or not.
#
# The projects that have been analyzed are mostly from the Apache Software Foundation:
#
# * [Atlas (1.2.0-rc3)](https://github.com/apache/atlas/releases/tag/release-1.2.0-rc3)
# * [Calcite (1.18.0)](https://github.com/apache/calcite/releases/tag/calcite-1.18.0)
# * [Camel (2.20.4)](https://github.com/apache/camel/tree/220d36e64b669bc1464a41a613f512e6bae23edb)
# * [Dubbo (2.7.3)](https://github.com/apache/dubbo/releases/tag/dubbo-2.7.3)
# * [Flink (1.9.0)](https://github.com/apache/flink/releases/tag/release-1.9.0)
# * [Hadoop (3.1.1)](https://github.com/apache/hadoop/tree/2b9a8c1d3a2caf1e733d57f346af3ff0d5ba529c)
# * [Hbase (2.1.6)](https://github.com/apache/hbase/releases/tag/rel%2F2.1.6)
# * [Ignite (2.07.05)](https://github.com/apache/ignite/releases/tag/2.7.5)
# * [Knox (0.3.0)](https://github.com/apache/knox/releases/tag/v0.3.0-final)
# * [Maven (3.5.0)](https://github.com/apache/maven/tree/ff8f5e7444045639af65f6095c62210b5713f426)
# * [Nifi (0.4.0)](https://github.com/apache/nifi/releases/tag/nifi-0.4.0)
# * [Pulsar (2.4.1)](https://github.com/apache/pulsar/releases/tag/v2.4.1)
# * [Ranger (2.0.0)](https://github.com/apache/ranger/releases/tag/release-ranger-2.0.0)
# * [Rocketmq (4.3.0)](https://github.com/apache/rocketmq/releases/tag/rocketmq-all-4.3.0)
# * [Storm (0.9.3)](https://github.com/apache/storm/releases/tag/v0.9.3)
# * [Syncope (1.2.8)](https://github.com/apache/syncope/releases/tag/syncope-1.2.8)
# * [Tika (1.0)](https://github.com/apache/tika/releases/tag/1.0)
# * [Tomcat (8.05.01)](https://github.com/apache/tomcat/releases/tag/8.5.1)
#
# Also intentionally vulnerable projects were taken into account:
#
# * [bodgeit (1.4.0)](https://github.com/psiinon/bodgeit)
# * [Webbank](https://github.com/pentestingforfunandprofit/webbank)
# * [Webgoat](https://github.com/WebGoat/WebGoat)
# ### Source files labeling
#
# Artifacts of all the projects were used to perform static analysis with SpotBugs and its plugin Find Security Bugs. As
# the task is to work at file-level granularity, files are marked vulnerable if they presented at least a bug belonging to
# correctness, multi-threaded correctness, malicious code, bad practice and security categories.
#
# The choice of having the output of a static analyzer as ground truth (and so to avoid using public databases like the
# NVD) derives from the results shown in previous works that highlight how models trained on synthetic source code perform
# worse than those trained on SASTs output.
#
# Although SpotBugs has been reported to output a relevant amount of false-positives (this behavior has been improved with
# the latest releases), the results of this study are to be considered as a lower bound of the performance that a similar
# model would achieve if trained on a hand-crafted dataset made by security experts.
# ### Dataset distribution
#
# The data collected turned out not to be split equally between safe and vulnerable files.
# + pycharm={"name": "#%%\n", "is_executing": false}
import src.preparation.stats_extractor as stats_extractor
stats_extractor.__extract_stats()
# + [markdown] pycharm={"name": "#%% md\n"}
# The stats for every project analyzed are reported in the table below.
# + pycharm={"name": "#%%\n", "is_executing": false}
projects_stats = stats_extractor.count_project_files()
total = 0
weak = 0
print("{:12s} {:14s} {:15s}".format('Project','Total files','Weak files'))
for project in projects_stats.keys():
total += projects_stats[project]["total_files"]
weak += projects_stats[project]["weak_files"]
print("{:12s} {:>11d} {:>13d}".format(project, projects_stats[project]["total_files"],
projects_stats[project]["weak_files"]))
print("\n{:12s} {:>11d} {:>13d}".format("Total", total, weak))
# + [markdown] pycharm={"name": "#%% md\n"}
# Overall the stats reported show the high unbalance between vulnerable and non-vulnerable files and this probably
# suggests the need for undersampling when it comes to modeling.
#
# More detailed info are reported by the pie charts that follow.
# + pycharm={"name": "#%%\n", "is_executing": false}
import matplotlib.pyplot as plt
# %matplotlib inline
files_count = stats_extractor.count_files()
lines_count = stats_extractor.count_lines()
tokens_count = stats_extractor.count_tokens()
fig, ax = plt.subplots(nrows=1, ncols=3, figsize=(15,7), constrained_layout=True)
wedges, texts, autotexts = ax[0].pie([files_count["safe"], files_count["weak"]],
autopct='%.2f%%', textprops=dict(color="w"))
ax[0].axis("equal")
ax[0].legend(["Safe files", "Weak files"], loc="upper center", fontsize=16)
plt.setp(autotexts, size=16, weight="bold")
ax[0].set_title("Files distibution", fontsize=24)
wedges, texts, autotexts = ax[1].pie([lines_count["safe"], lines_count["weak"]],
autopct='%.2f%%', textprops=dict(color="w"))
ax[1].axis("equal")
ax[1].legend(["Safe lines", "Weak lines"], loc="upper center", fontsize=16)
plt.setp(autotexts, size=16, weight="bold")
ax[1].set_title("Lines distribution", fontsize=24)
wedges, texts, autotexts = ax[2].pie([tokens_count["safe"], tokens_count["weak"]],
autopct='%.2f%%', textprops=dict(color="w"))
ax[2].axis("equal")
ax[2].legend(["Safe tokens", "Weak tokens"], loc="upper center", fontsize=16)
plt.setp(autotexts, size=16, weight="bold")
ax[2].set_title("Tokens distribution", fontsize=24)
plt.show()
| notebook/eda/dataset_distribution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Regression Challenge
#
# Predicting the selling price of a residential property depends on a number of factors, including the property age, availability of local amenities, and location.
#
# In this challenge, you will use a dataset of real estate sales transactions to predict the price-per-unit of a property based on its features. The price-per-unit in this data is based on a unit measurement of 3.3 square meters.
#
# > **Citation**: The data used in this exercise originates from the following study:
# >
# > *<NAME>., & <NAME>. (2018). Building real estate valuation models with comparative approach through case-based reasoning. Applied Soft Computing, 65, 260-271.*
# >
# > It was obtained from the UCI dataset repository (<NAME>. and <NAME>. (2019). [UCI Machine Learning Repository]([http://archive.ics.uci.edu/ml). Irvine, CA: University of California, School of Information and Computer Science).
#
# ## Review the data
#
# Run the following cell to load the data and view the first few rows.
# +
import pandas as pd
# load the training dataset
data = pd.read_csv('data/real_estate.csv')
data.head()
# -
# The data consists of the following variables:
#
# - **transaction_date** - the transaction date (for example, 2013.250=2013 March, 2013.500=2013 June, etc.)
# - **house_age** - the house age (in years)
# - **transit_distance** - the distance to the nearest light rail station (in meters)
# - **local_convenience_stores** - the number of convenience stores within walking distance
# - **latitude** - the geographic coordinate, latitude
# - **longitude** - the geographic coordinate, longitude
# - **price_per_unit** house price of unit area (3.3 square meters)
#
# ## Train a Regression Model
#
# Your challenge is to explore and prepare the data, identify predictive features that will help predict the **price_per_unit** label, and train a regression model that achieves the lowest Root Mean Square Error (RMSE) you can achieve (which must be less than **7**) when evaluated against a test subset of data.
#
# Add markdown and code cells as required to create your solution.
#
# > **Note**: There is no single "correct" solution. A sample solution is provided in [02 - Real Estate Regression Solution.ipynb](02%20-%20Real%20Estate%20Regression%20Solution.ipynb).
# Your code to explore data and train a regression model
numeric_features = ['house_age', 'transit_distance', 'latitude', 'longitude']
data[numeric_features + ['price_per_unit']].describe()
import numpy as np
import pandas as pd
data['month'], data['year'] = np.modf(data['transaction_date'])
data['month'] = data['month'] * 12
data['year'].round()
data.month = data.month.astype(int)+1
data.year = data.year.astype(int)
data.describe
# +
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
label = data['price_per_unit']
fig, ax = plt.subplots(2, 1, figsize = (9,12))
ax[0].hist(label, bins=100)
ax[0].set_ylabel('Frequency')
ax[0].axvline(label.mean(), color='magenta', linestyle='dashed', linewidth=2)
ax[0].axvline(label.median(), color='cyan', linestyle='dashed', linewidth=2)
ax[1].boxplot(label, vert=False)
ax[1].set_xlabel('price_per_unit')
fig.suptitle('Price per unit Distribution')
fig.show()
# -
# remove outliers
data = data[data['price_per_unit']<70]
for col in numeric_features:
fig = plt.figure(figsize=(9, 6))
ax = fig.gca()
feature = data[col]
feature.hist(bins=100, ax = ax)
ax.axvline(feature.mean(), color='magenta', linestyle='dashed', linewidth=2)
ax.axvline(feature.median(), color='cyan', linestyle='dashed', linewidth=2)
ax.set_title(col)
plt.show()
categorical_features = ['local_convenience_stores','month','year']
for col in categorical_features:
counts = data[col].value_counts().sort_index()
fig = plt.figure(figsize=(9, 6))
ax = fig.gca()
counts.plot.bar(ax = ax, color='steelblue')
ax.set_title(col + ' counts')
ax.set_xlabel(col)
ax.set_ylabel("Frequesncy")
plt.show()
for col in numeric_features:
fig = plt.figure(figsize=(9, 6))
ax = fig.gca()
feature = data[col]
label = data['price_per_unit']
correlation = feature.corr(label)
plt.scatter(x=feature, y=label)
plt.xlabel(col)
plt.ylabel('Price per unit')
ax.set_title('price vs '+ col + '- correlation: ' + str(correlation))
plt.show()
for col in categorical_features:
fig = plt.figure(figsize=(9, 6))
ax = fig.gca()
data.boxplot(column = 'price_per_unit', by = col, ax = ax)
ax.set_title('Label by ' + col)
ax.set_ylabel("Prices")
plt.show()
X, y = data[['house_age', 'transit_distance', 'local_convenience_stores', 'latitude', 'longitude', 'month', 'year']].values, data['price_per_unit'].values
print('Features:', X[:10], '\nLabels:', y[:10], sep='\n')
# +
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=0)
print('X_train shape: ',X_train.shape,'\nX_test shape: ',X_test.shape)
# +
from sklearn.linear_model import LinearRegression
model = LinearRegression()
model.fit(X_train, y_train)
# +
import numpy as np
predictions = model.predict(X_test)
np.set_printoptions(suppress=True)
print('Predicted labels: ', np.round(predictions)[:10])
print('Actual labels: ', y_test[:10])
# +
import matplotlib.pyplot as plt
# %matplotlib inline
plt.scatter(y_test, predictions)
plt.xlabel('Actual Labels')
plt.ylabel('Predicted Labels')
plt.title('Predictions vs actual labels')
z = np.polyfit(y_test, predictions, 1)
p = np.poly1d(z)
plt.plot(y_test, p(y_test), color='magenta')
plt.show()
# +
from sklearn.metrics import mean_squared_error, r2_score
mse = mean_squared_error(y_test, predictions)
print('MSE:', mse)
rmse = np.sqrt(mse)
print('RMSE:', rmse)
r2 = r2_score(y_test, predictions)
print('R2:', r2)
# +
# Try Lasso
from sklearn.linear_model import Lasso
model = Lasso()
model.fit(X_train, y_train)
predictions = model.predict(X_test)
mse = mean_squared_error(y_test, predictions)
print('MSE:', mse)
rmse = np.sqrt(mse)
print('RMSE:', rmse)
r2 = r2_score(y_test, predictions)
print('R2:', r2)
plt.scatter(y_test, predictions)
plt.xlabel('Actuel Labels')
plt.ylabel('Predicted Labels')
plt.title('Predictions vs actual labels')
z = np.polyfit(y_test, predictions, 1)
p = np.poly1d(z)
plt.plot(y_test, p(y_test), color='magenta')
plt.show
# +
# Try DecisionTreeRegressor
from sklearn.tree import DecisionTreeRegressor
model = DecisionTreeRegressor()
model.fit(X_train, y_train)
predictions = model.predict(X_test)
predictions[:10]
# +
mse = mean_squared_error(y_test, predictions)
print('MSE:', mse)
rmse = np.sqrt(mse)
print('RMSE:', rmse)
r2 = r2_score(y_test, predictions)
print('R2:', r2)
plt.scatter(y_test, predictions)
plt.xlabel('Actual Labels')
plt.ylabel('Predictions')
plt.title('Predictions vs labels')
z = np.polyfit(y_test, predictions, 1)
p = np.poly1d(z)
plt.plot(y_test, p(y_test), color='magenta')
plt.show
# +
# Lasso is the best one for now
# let's try another algorithm
# let's try GradientBoostingRegressor
# +
from sklearn.ensemble import GradientBoostingRegressor
model = GradientBoostingRegressor().fit(X_train, y_train)
predictions = model.predict(X_test)
mse = mean_squared_error(y_test, predictions)
print('MSE:', mse)
rmse = np.sqrt(mse)
print('RMSE:', rmse)
r2 = r2_score(y_test, predictions)
print('R2:', r2)
plt.scatter(y_test, predictions)
plt.xlabel('Actual Labes')
plt.ylabel('Predicted Labes')
plt.title('Predictions vs labels')
z = np.polyfit(y_test, predictions, 1)
p = np.poly1d(z)
plt.plot(y_test, p(y_test), color='magenta')
plt.show()
# +
# GradientBoostingRegressor is better than Lasso
# let's try to optimize hyperparameters of GradientBoostingRegressor
# +
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import make_scorer, r2_score
alg = GradientBoostingRegressor()
params = {
'learning_rate': [0.1, 0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.9, 1.0],
'n_estimators': [50, 100, 150]
}
score = make_scorer(r2_score)
gridsearch = GridSearchCV(alg, params, scoring=score, cv=3, return_train_score=True)
gridsearch.fit(X_train, y_train)
print('Best parameters: ', gridsearch.best_params_, '\n')
model = gridsearch.best_estimator_
print(model, '\n')
# +
predictions = model.predict(X_test)
mse = mean_squared_error(y_test, predictions)
print('MSE:', mse)
rmse = np.sqrt(mse)
print('RMSE:', rmse)
r2 = r2_score(y_test, predictions)
print('R2:', r2)
plt.scatter(y_test, predictions)
plt.xlabel('actual labels')
plt.ylabel('predictions')
plt.title('preds. vs labels')
z = np.polyfit(y_test, predictions, 1)
p = np.poly1d(z)
plt.plot(y_test, p(y_test), color='magenta')
plt.show()
# +
# Preprocess the data
from sklearn.compose import ColumnTransformer
from sklearn.pipeline import Pipeline
from sklearn.impute import SimpleImputer
from sklearn.preprocessing import StandardScaler, OneHotEncoder
from sklearn.linear_model import LinearRegression
import numpy as np
numeric_features = [0,1,3,4]
numeric_transformer = Pipeline(steps=[
('scaler', StandardScaler())
])
categorical_features = [2,5,6]
categorical_transformer = Pipeline(steps=[
('onehot', OneHotEncoder(handle_unknown='ignore'))
])
preprocessor = ColumnTransformer(
transformers=[
('num', numeric_transformer, numeric_features),
('cat', categorical_transformer, categorical_features)
])
pipeline = Pipeline(steps=[('preprocessor', preprocessor),
('regressor', GradientBoostingRegressor())])
model = pipeline.fit(X_train, (y_train))
print(model)
# +
predictions = model.predict(X_test)
mse = mean_squared_error(y_test, predictions)
print('MSE:', mse)
rmse = np.sqrt(mse)
print('RMSE:', rmse)
r2 = r2_score(y_test, predictions)
print('R2:', r2)
plt.scatter(y_test, predictions)
plt.xlabel('actual labels')
plt.ylabel('predictions')
plt.title('preds. vs labels')
z = np.polyfit(y_test, predictions, 1)
p = np.poly1d(z)
plt.plot(y_test, p(y_test), color='magenta')
plt.show()
# +
# try pipeline with RandomForestRegressor
from sklearn.ensemble import RandomForestRegressor
pipeline = Pipeline(steps=[('preprocessor', preprocessor),
('regressor', RandomForestRegressor())])
model = pipeline.fit(X_train, (y_train))
print (model)
# +
predictions = model.predict(X_test)
mse = mean_squared_error(y_test, predictions)
print('MSE:', mse)
rmse = np.sqrt(mse)
print('RMSE:', rmse)
r2 = r2_score(y_test, predictions)
print('R2:', r2)
plt.scatter(y_test, predictions)
plt.xlabel('actual labels')
plt.ylabel('predictions')
plt.title('preds. vs labels')
z = np.polyfit(y_test, predictions, 1)
p = np.poly1d(z)
plt.plot(y_test, p(y_test), color='magenta')
plt.show()
# +
# conclusion
Best parameters: {'learning_rate': 0.1, 'n_estimators': 50}
GradientBoostingRegressor(n_estimators=50)
# -
# ## Use the Trained Model
#
# Save your trained model, and then use it to predict the price-per-unit for the following real estate transactions:
#
# | transaction_date | house_age | transit_distance | local_convenience_stores | latitude | longitude |
# | ---------------- | --------- | ---------------- | ------------------------ | -------- | --------- |
# |2013.167|16.2|289.3248|5|24.98203|121.54348|
# |2013.000|13.6|4082.015|0|24.94155|121.50381|
# +
# Your code to use the trained model
| challenges/02 - Real Estate Regression Challenge.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import os
import seaborn as sns
import matplotlib.pyplot as plt
# build data directory
data_dir = '../../data/stage1'
os.listdir(data_dir)
# Load user profiles
churn_od_df = pd.read_csv(
os.path.join(data_dir, 'Churn_OD.txt'),
delimiter = "|",
encoding = "ISO-8859-1"
)
churn_od_df.sort_values(by=['Demand_weight'], ascending = False).head(10)
# +
#churn_od_df.groupby(['County_of_Origin']).agg({'Demand_weight':'sum'})
# -
# ## What is the most popular route from each county?
df_max_demand_from = churn_od_df.groupby(['County_of_Origin']).agg({'Demand_weight':'max'})
df_max_demand_from = pd.merge(churn_od_df, df_max_demand_from, on=['County_of_Origin', 'Demand_weight'], how = 'inner')
df_max_demand_from[['County_of_Origin','County_of_Public_Transportation','Demand_weight']].sort_values(by=['Demand_weight'], ascending = False)
# ## Routes per County
churn_od_df['County_of_Origin'] = churn_od_df['County_of_Origin'].str.upper()
(churn_od_df.groupby('County_of_Public_Transportation')
.agg({"County_of_Public_Transportation": 'count'})
.rename(columns={'County_of_Public_Transportation': 'number_of_routes'})
.sort_values(by=['number_of_routes'], ascending = False).head(10))
## most routes end at Lisboa followed by Porto
## numbers are high as there are multiple perish codes per county. Dont have perish codes for origin
churn_od_df[churn_od_df['County_of_Public_Transportation'] == 'LISBOA']
# #### Most routes end in Lisboa and Porto
#
# ## Internal routes
# +
churn_od_df['County_of_Origin'] = churn_od_df['County_of_Origin'].str.upper()
routes_count = (churn_od_df.groupby('County_of_Origin')
.agg({"County_of_Origin": 'count'})
.rename(columns={'County_of_Origin': 'number_of_routes'})
.sort_values(by=['number_of_routes'], ascending = False))
churn_od_df_internal = churn_od_df[churn_od_df['County_of_Public_Transportation'] == churn_od_df['County_of_Origin']]
churn_od_df_internal_agg = (churn_od_df_internal.groupby('County_of_Origin')
.agg({'County_of_Origin': 'count'})
.rename(columns={'County_of_Origin': 'number_of_routes_internal'}))
routes_count_final = pd.merge(routes_count, churn_od_df_internal_agg, left_index=True, right_index=True, how = 'inner')
routes_count_final['percentage_of_routes_internal'] = routes_count_final['number_of_routes_internal']/ routes_count_final['number_of_routes']
routes_count_final['number_of_routes_external'] = routes_count_final['number_of_routes'] - routes_count_final['number_of_routes_internal']
routes_count_final['percentage_of_routes_external'] = 1- routes_count_final['percentage_of_routes_internal']
routes_count_final
# +
# can filter out return trips?
# -
pd.crosstab(churn_od_df.County_of_Origin , churn_od_df.County_of_Public_Transportation )
| notebooks/stage1/Churn_OD_exploration_vinay.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: cmap
# language: python
# name: cmap
# ---
# +
import os
import numpy as np
import pandas as pd
import pkg_resources
from cmapPy.pandasGEXpress.parse import parse
import matplotlib.pyplot as plt
import seaborn as sns
# notebook settings
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
# Print version of cmapPy being used in current conda environment
pkg_resources.get_distribution("cmapPy").version
# -
def clean_cell_names(pd_series):
# remove trailing strings
clean = pd_series.apply(lambda x: x.split()[0].strip() if x else None).replace("\([a-zA-Z0-9\-]+", "", regex=True)
# remove punctuation & capitalize
clean = clean.replace("[^\w\s]+", "", regex=True).str.upper().values
return clean
# +
def getTCGA(disease):
path = "/srv/nas/mk2/projects/pan-cancer/TCGA_CCLE_GCP/TCGA/TCGA_{}_counts.tsv.gz"
files = [path.format(d) for d in disease]
return files
def readGCP(files, biotype='protein_coding', mean=True):
"""
Paths to count matrices.
"""
data_dict = {}
for f in files:
key = os.path.basename(f).split("_")[1]
data = pd.read_csv(f, sep='\t', index_col=0)
# transcript metadata
meta = pd.DataFrame([row[:-1] for row in data.index.str.split("|")],
columns=['ENST', 'ENSG', 'OTTHUMG', 'OTTHUMT', 'GENE-NUM', 'GENE', 'BP', 'BIOTYPE'])
#meta = pd.MultiIndex.from_frame(meta)
data.index = meta
# subset transcripts
data = data.xs(key=biotype, level='BIOTYPE')
data = data.droplevel(['ENST', 'ENSG', 'OTTHUMG', 'OTTHUMT', 'GENE-NUM', 'BP'])
# average gene expression of splice variants
data = data.T
if mean:
data = data.groupby(by=data.columns, axis=1).mean()
data_dict[key] = data
return data_dict
# -
# ## TCGA Gene list
f = getTCGA(disease=['PRAD'])
tcga = pd.read_csv(f[0], sep='\t', index_col=0)
# transcript metadata
meta = pd.DataFrame([row[:-1] for row in tcga.index.str.split("|")],
columns=['ENST', 'ENSG', 'OTTHUMG', 'OTTHUMT', 'GENE-NUM', 'GENE', 'BP', 'BIOTYPE'])
tcga_genes = meta[meta['BIOTYPE']=='protein_coding']['GENE'].unique()
# ### Shared Compound Data
base = "/srv/nas/mk2/projects/pan-cancer/LINCS/GSE70138/"
# compound (perturbation) meta data
sig_info = pd.read_csv(os.path.join(base, "GSE70138_Broad_LINCS_sig_info_2017-03-06.txt.gz"), sep="\t")
sig_info['clean_name'] = clean_cell_names(sig_info['cell_id'])
sig_info['clean_name'].nunique()
sig_info.head()
ccle_meta = pd.read_csv('/srv/nas/mk2/projects/pan-cancer/TCGA_CCLE_GCP/CCLE/CCLE_GDC_Metadata.tsv.gz', sep='\t')
uq = sig_info[sig_info['clean_name'].isin(ccle_meta['stripped_cell_line_name'])]['clean_name'].unique()
uq
dm_ccle_meta = pd.read_csv('/srv/nas/mk2/projects/pan-cancer/DepMap/CCLE/sample_info_v2.csv', sep=",")
dm_ccle_meta[dm_ccle_meta['stripped_cell_line_name'].isin(uq)]
# extract sig_ids from pert_id and cell_id of interest
sig_meta = sig_info[sig_info['clean_name'].isin(ccle_meta['stripped_cell_line_name'])]
sig_meta = sig_meta.set_index("sig_id")
len(sig_meta.index)
# genetic metadata
gene_info = pd.read_csv("ftp://ftp.ncbi.nlm.nih.gov/geo/series/GSE70nnn/GSE70138/suppl/GSE70138_Broad_LINCS_gene_info_2017-03-06.txt.gz", sep="\t", dtype=str)
gene_info.head()
# get landmark genes only
landmark_gene_meta = gene_info[gene_info["pr_is_lm"] == "1"]
landmark_gene_meta = landmark_gene_meta.set_index('pr_gene_id')
len(landmark_gene_meta.index)
landmark_gene_meta = landmark_gene_meta[landmark_gene_meta['pr_gene_symbol'].isin(tcga_genes)]
len(landmark_gene_meta)
# # get perturbations info
# pert_meta = pd.read_csv("ftp://ftp.ncbi.nlm.nih.gov/geo/series/GSE70nnn/GSE70138/suppl/GSE70138_Broad_LINCS_pert_info_2017-03-06.txt.gz", sep="\t")
# pert_meta.head()
# ### Subset GCToo Object
gctoo = parse(os.path.join(base, "GSE70138_Broad_LINCS_Level2_GEX_n345976x978_2017-03-06.gctx"))
sig_meta['id_list'] = sig_meta['distil_id'].str.split("|")
sig_dict = {}
for sid,sig_rep in zip(sig_meta.index, sig_meta['id_list']):
for i in sig_rep:
sig_dict[i] = sid
gctoo = parse(os.path.join(base, "GSE70138_Broad_LINCS_Level2_GEX_n345976x978_2017-03-06.gctx"),
rid=landmark_gene_meta.index, convert_neg_666=True)
gctoo.row_metadata_df = landmark_gene_meta
gctoo.data_df.shape
gctoo.data_df.head()
gctoo.data_df.index = gctoo.row_metadata_df['pr_gene_symbol']
# Group replicate experiments and take the average
gctoo_grp = gctoo.data_df.groupby(sig_dict, axis=1).mean()
gctoo.data_df.loc['AARS'].hist()
gctoo_grp.loc['AARS'].hist()
base
gctoo_grp.T.to_pickle(os.path.join(base, "processed/phase2_mean_rep.pkl"))
sig_meta.to_pickle(os.path.join(base, "processed/phase2_mean_row.pkl"))
landmark_gene_meta.to_pickle(os.path.join(base, "processed/phase2_mean_col.pkl"))
| notebook/2020.03.31_cmappy_2.7.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
import neurolab as nl
# %matplotlib inline
text = np.loadtxt('data_perceptron.txt')
data = text[:, :2]
labels = text[:, 2]
plt.figure()
plt.scatter(data[labels==0, 0], data[labels==0,1], marker='o')
plt.scatter(data[labels==1, 0], data[labels==1,1], marker='x')
plt.xlabel('Dimension 1')
plt.ylabel('Dimension 2')
plt.title('Input data')
plt.show()
# +
dim1_min, dim1_max, dim2_min, dim2_max = 0, 1, 0, 1
dim1 = [dim1_min, dim1_max]
dim2 = [dim2_min, dim2_max]
num_output = 1
perceptron = nl.net.newp([dim1, dim2], num_output)
error_progress = perceptron.train(data, labels.reshape(-1,1), epochs=100, show=20, lr=0.03)
plt.figure()
plt.plot(error_progress)
plt.xlabel('Number of epochs')
plt.ylabel('Training error')
plt.title('Training error progress')
plt.grid()
plt.show()
# +
xy = np.random.rand(100, 2)
out = perceptron.sim(xy).ravel()
plt.figure()
plt.scatter(xy[out==0,0], xy[out==0,1], marker='o')
plt.scatter(xy[out==1,0], xy[out==1,1], marker='x')
plt.xlabel('Dimension 1')
plt.ylabel('Dimension 2')
plt.show()
# -
| artificial-intelligence-with-python-ja-master/Chapter 14/perceptron_classifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Optimizing Code: Holiday Gifts
# In the last example, you learned that using vectorized operations and more efficient data structures can optimize your code. Let's use these tips for one more example.
#
# Say your online gift store has one million users that each listed a gift on a wish list. You have the prices for each of these gifts stored in `gift_costs.txt`. For the holidays, you're going to give each customer their wish list gift for free if it is under 25 dollars. Now, you want to calculate the total cost of all gifts under 25 dollars to see how much you'd spend on free gifts. Here's one way you could've done it.
import time
import numpy as np
# +
with open('gift_costs.txt') as f:
gift_costs = f.read().split('\n')
gift_costs = np.array(gift_costs).astype(int) # convert string to int
# +
start = time.time()
total_price = 0
for cost in gift_costs:
if cost < 25:
total_price += cost * 1.08 # add cost after tax
print(total_price)
print('Duration: {} seconds'.format(time.time() - start))
# -
# Here you iterate through each cost in the list, and check if it's less than 25. If so, you add the cost to the total price after tax. This works, but there is a much faster way to do this. Can you refactor this to run under half a second?
# ## Refactor Code
# **Hint:** Using numpy makes it very easy to select all the elements in an array that meet a certain condition, and then perform operations on them together all at once. You can them find the sum of what those values end up being.
# +
start = time.time()
total_price = gift_costs[gift_costs < 25].sum() # TODO: compute the total price
print(total_price)
print('Duration: {} seconds'.format(time.time() - start))
# -
| lessons/SoftwarePractices/OptimizeAndRefactor/OptimizeHolidayGifts/optimizing_code_holiday_gifts.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # TPOT training
# I have realized that TPOT takes a really long time to run, days. This notebook contains the TPOT runs I have tried. Another notebook contains the inspection of the results, as I can do it while running using checkpoints.
# ## Setup
import pickle
import sys
sys.path.append('..')
from utils import *
x_train, y_train, eras_train = train_data(weras=True)
x_val, y_val = val_data()
bl = baseline(x_train, y_train, x_val, y_val)
bl
# ## TPOT
from tpot import TPOTClassifier
# ### TPOT light
model = TPOTClassifier(
config_dict='TPOT light',
scoring='neg_log_loss',
periodic_checkpoint_folder='tpot',
n_jobs=8,
verbosity=3,
)
model.fit(x_train, y_train, groups=eras_train)
y_val_pred = model.predict_proba(x_val)
validate(y_val, y_val_pred, bl, True)
model.export('tpot.py')
with open('tpot.pkl', 'wb') as auto_sklearn_file:
pickle.dump(model, auto_sklearn_file)
# ## Auto-sklearn
from autosklearn.classification import AutoSklearnClassifier
from autosklearn.metrics import log_loss
model = AutoSklearnClassifier(
time_left_for_this_task=4*60*60, # 4h
ml_memory_limit=6144, # 6GB
)
# %time model.fit(x_train, y_train, metric=log_loss)
model.show_models()
model.get_models_with_weights()
y_val_pred = model.predict_proba(x_val)
validate(y_val, y_val_pred, bl, True)
with open('auto_sklearn.pkl', 'wb') as auto_sklearn_file:
pickle.dump(model, auto_sklearn_file)
| 2-iter/tpot_train.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Suma de polinomios 1
#
# m+n+s; -m-n+s =
#
# n+s; -n+s =
#
# s+s =
#
# 2s
# # Suma de polinomios 2
#
# -7x-4y+6z; 10x-20y-8z; -5x+24y+2z =
#
# -2x
# # Resta de polinomios 1
#
# ## De x^3 - y^2 restar - y^2 + 2x^3 - 2xy
#
# x^3 - y^2 - (-y^2 + 2x^3 - 2xy) =
#
# -1x^3 + 2xy
# # Resta de polinomios 1
#
# # De x^5 - x^2 y^3 +6xy^4 + 25y^5 restar -3xy^4 -8x^3 y^2 + 19y^5+18
#
# x^5 - x^2.y^3 +6xy^4 + 25y^5 - (-3xy^4 -8x^3.y^2 + 19y^5+18) =
#
# x^5 + 8x^3.y^2 - x^2.y^3 + 9xy^4 + 6y^5 - 18
| 01_algebra/modulo_II_polinomios/ejercicios/00-suma-resta.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import random
import sys, os
import pandas as pd
from pathlib import Path
import matplotlib.pyplot as plt
from PIL import Image, ImageDraw
sys.path.append(os.path.abspath("./"))
from utils import *
import ipywidgets as widgets
from IPython.display import display as disp
from ipywidgets import interact, interactive, SelectionRangeSlider
# +
classes_path = '../model_data/coco_classes.txt'
detection_file_path = '../data/output/mhca/detection_output.txt'
classes = get_classes(classes_path)
id2class = {str(i): classes[i] for i in range(len(classes))}
img2bbs = get_predictions(detection_file_path)
img2bb = explode_dict(img2bbs)
non_classified_image_paths = [path for path, bbs in img2bbs.items() if len(bbs) == 0]
# -
# ## Boxer
# +
img = get_sample_from_list(non_classified_image_paths)[0]
def print_box2(vertical, horizontal):
bottom, top = vertical
left, right = horizontal
bb = BoundingBox(f'{top},{left},{right},{bottom},{0}')
disp(print_coords(img, bb))
print(f'{bb.top}, {bb.left}, {bb.right}, {bb.bottom}')
return (top, left, right, bottom)
v_slider = widgets.SelectionRangeSlider(
orientation='vertical',
options=[i for i in range(img.size[1], -1, -1)],
index=(0, img.size[1]-1),
description='Top - Down',
disabled=False
)
h_slider = widgets.SelectionRangeSlider(
orientation='horizontal',
options=[i for i in range(0, img.size[0])],
index=(0, img.size[0]-1),
description='Left - Right',
disabled=False
)
out = widgets.interactive_output(
print_box2,
{'vertical': v_slider, 'horizontal':h_slider})
ui_top = widgets.HBox([h_slider])
ui_bottom = widgets.HBox([v_slider, out])
display(ui_top, ui_bottom)
# +
def get_coord(v_slider, h_slider, path='/path/to/image/', label='label'):
return f'{path} {v_slider.value[1]},{h_slider.value[0]},{h_slider.value[1]},{v_slider.value[0]},{label}'
get_coord(v_slider, h_slider)
| notebook/X.3.boxer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="GznIXM97XwBY"
# # CAP379 - Topics in Spatial Science
#
# **Course project - Group 1**
#
# _<small>Last update: 2019-09-01</small>_
#
# ## Objectives - Part 1
#
# * Reproduce and generalize the PythonSQL (AstroML) for SDCC datasets, using Jupyter Notebook
# * Observatory Sloan Digital Sky Survey (SDSS)
# * Algorithm Fetch SDSS Galaxy Colors
# * Dataset CDM
#
# ## Summary
# 
# + [markdown] colab_type="text" id="uHrHybiqaW25"
# ## The telescope Sloan Digital Sky Survey (SDSS)
#
# 
# *Telescope SDSS, New Mexico, USA. Source: https://classic.sdss.org*
#
# The Sloan Digital Sky Survey or SDSS is a major multi-spectral imaging and spectroscopic redshift survey using a dedicated 2.5-m wide-angle optical telescope at Apache Point Observatory in New Mexico, United States. The project was named after the Alfred P. Sloan Foundation, which contributed significant funding.
#
# Data collection began in 2000; the final imaging data release (DR9) covers over 35% of the sky, with photometric observations of around nearly 1 billion objects, while the survey continues to acquire spectra, having so far taken spectra of over 4 million objects. The main galaxy sample has a median redshift of z = 0.1; there are redshifts for luminous red galaxies as far as z = 0.7, and for quasars as far as z = 5; and the imaging survey has been involved in the detection of quasars beyond a redshift z = 6.
#
# Data release 8 (DR8), released in January 2011, includes all photometric observations taken with the SDSS imaging camera, covering 14,555 square degrees on the sky (just over 35% of the full sky). Data release 9 (DR9), released to the public on 31 July 2012, includes the first results from the Baryon Oscillation Spectroscopic Survey (BOSS) spectrograph, including over 800,000 new spectra. Over 500,000 of the new spectra are of objects in the Universe 7 billion years ago (roughly half the age of the universe).Data release 10 (DR10), released to the public on 31 July 2013, includes all data from previous releases, plus the first results from the APO Galactic Evolution Experiment (APOGEE) spectrograph, including over 57,000 high-resolution infrared spectra of stars in the Milky Way. DR10 also includes over 670,000 new BOSS spectra of galaxies and quasars in the distant universe. The publicly available images from the survey were made between 1998 and 2009.
#
# **Data access**
#
# The survey makes the data releases available over the Internet. The SkyServer provides a range of interfaces to an underlying Microsoft SQL Server. Both spectra and images are available in this way, and interfaces are made very easy to use so that, for example, a full color image of any region of the sky covered by an SDSS data release can be obtained just by providing the coordinates. The data are available for non-commercial use only, without written permission. The SkyServer also provides a range of tutorials aimed at everyone from schoolchildren up to professional astronomers. The tenth major data release, DR10, released in July 2013, provides images, imaging catalogs, spectra, and redshifts via a variety of search interfaces.
#
# The raw data (from before being processed into databases of objects) are also available through another Internet server, and first experienced as a 'fly-through' via the NASA World Wind program.
#
# Sky in Google Earth includes data from the SDSS, for those regions where such data are available. There are also KML plugins for SDSS photometry and spectroscopy layers, allowing direct access to SkyServer data from within Google Sky.
#
# The data is also available on Hayden Planetarium with a 3D visualizer.
#
# There is also the ever-growing list of data for the Stripe 82 region of the SDSS.
#
# Following from Technical Fellow <NAME>'s contribution on behalf of Microsoft Research with the SkyServer project, Microsoft's WorldWide Telescope makes use of SDSS and other data sources.
#
# MilkyWay@home also used SDSS's data for creating a highly accurate three dimensional model of the Milky Way galaxy.
#
# *Source: https://pt.wikipedia.org/wiki/Sloan_Digital_Sky_Survey (adapted)*
# + [markdown] colab_type="text" id="GznIXM97XwBY"
# ## AstroML
#
# AstroML (Machine Learning and Data Mining for Astronomy) is a Python module for machine learning and data mining built on numpy, scipy, scikit-learn, matplotlib, and astropy. The goal is to provide a community repository for fast Python implementations of common tools and routines used for statistical data analysis in astronomy and astrophysics, to provide a uniform and easy-to-use interface to freely available astronomical datasets.
#
# The astroML project was started in 2012 to accompany the book "Statistics, Data Mining, and Machine Learning in Astronomy" by <NAME>, <NAME>, <NAME>, and <NAME>, published by Princeton University Press (https://press.princeton.edu/titles/10159.html)
#
# 
# *Source: www.astroml.org*
#
# ## Data Sets
#
# https://www.astroml.org/user_guide/datasets.html
#
# One of the major components of astroML is its tools for downloading and working with astronomical data sets. The available routines are available in the module astroML.datasets.
#
# ## SDSS Data
#
# Much of the data made available by astroML comes from the **Sloan Digital Sky Survey (SDSS)**, a decade-plus photometric and spectroscopic survey at the Apache Point Observatory in New Mexico. The survey obtained photometry for hundreds of millions of stars, quasars, and galaxies, and spectra for several million of these objects. In addition, the second phase of the survey performed repeated imaging over a small portion of the sky, called Stripe 82, enabling the study of the time-variation of many objects.
#
# SDSS photometric data are observed through five filters, u, g, r, i, and z. A visualization of the range of these filters is shown below:
#
# 
# *SDSS photometric data. Source: https://www.astroml.org*
#
# ## List of datasets
#
# https://www.astroml.org/user_guide/datasets.html
# + [markdown] colab_type="text" id="EmtEfL7vl245"
# ## Dataset - Python SQL - SDSS Photometry
#
# The photometric data can be accessed directly using the *SQL interface* (http://cas.sdss.org/) to the SDSS Catalog Archive Server (CAS). AstroML contains a function which accesses this data directly using a **Python SQL** query tool. The function is called **fetch_sdss_galaxy_colors()** and can be used as a template for making custom data sets available with a simple Python command.
# -
# ## http://cas.sdss.org/
#
# SkyServer is currently hosting the SDSS's Data Release 15 (DR15).
#
# The BestDR15 database can be downloaded as a multi-file SQL Server backup. It is about 6.5 TB in size (compressed) and the uncompressed size of the database is 12+ TB.
#
# When complete, the survey data will occupy about 40 terabytes (TB) of image data, and about 3 TB of processed data.
#
# 
# 
# 
# 
# FITS is the most commonly used digital file format in astronomy.
#
# Finding the meaning of columns. Example:
#
# ```sql
# SELECT TOP 5000
# p.u, p.g, p.r, p.i, p.z, s.class, s.z, s.zerr
# FROM PhotoObj AS p
# JOIN SpecObj AS s ON s.bestobjid = p.objid
# WHERE
# p.u BETWEEN 0 AND 19.6
# AND p.g BETWEEN 0 AND 20
# AND s.class <> 'UNKNOWN'
# AND s.class <> 'STAR'
# AND s.class <> 'SKY'
# AND s.class <> 'STAR_LATE'
# ```
#
# As example, the PhotoObj database:
# 
# ## Data Set DR15 HARDWARE
#
# Hardware needed to support a SDSS Catalog Archive Server (CAS) mirror site:
#
# - Separate boxes for Web server and database (DB) server
# - The DR12 DB servers as well as MyDB servers at JHU (Johns Hopkins University) have the following hardware configuration:
# - Intel Xeon 24-core CPU ES-2630v2 (2x12) @2.60GHz 64GB RAM
# - Network Adapter: Broadcom bcm5709c netxtreme ii gige
# - RAID controllers: Intel C600 SATA
# - Disks configured to RAID 5 with 2 logical volumes of 8.2 TB each and 2 dedicated TempDB volumes. Each hard drive is 700GB @10000 RPM.
# - All MyDB databases are hosted on 2 database servers, one of them being online at any given time.
# - For our production cluster we have 3-4 db servers per release, so that the public and collab users as well as CasJobs users can be adequately supported, and skyServer, ImgCutout and CasJobs queries can be load-balanced on different boxes. We dedicate one server to ImgCutout queries for multiple releases, since these queries can be quite large in number and intensive.
# - At any given time, we have 2 or more DB servers in production per release, and the rest are used for testing and data loading purposes. Quick and long queries are pointed to two separate servers. One of the two MyDB servers will be in production and the other one is used for warm backup (MyDBs are bakced up daily to the backup machine).
# 
# *DR12 Hardware configuration at JHU for SDSS Servers*
#
# *Source: http://www.skyserver.org/mirrors/hardware.aspx*
# + [markdown] colab_type="text" id="jVHzKacQrJ18"
# ## Loading of Datasets: astroML.datasets
#
# Astronomy Datasets
#
# - fetch_sdss_galaxy_colors : Loader for SDSS galaxy colors
#
# Loader for SDSS galaxy colors.
#
# This function directly queries the sdss SQL database at http://cas.sdss.org/
#
# Parameters
#
# - **data_homeoptional, default=None** : specify another download and cache folder for the datasets. By default all scikit learn data is stored in ‘~/astroML_data’ subfolders.
#
# - **download_if_missingoptional, default=True** : if False, raise a IOError if the data is not locally available instead of trying to download the data from the source site.
#
# Returns
#
# - **datarecarray, shape = (10000,)** : record array containing magnitudes and redshift for each galaxy.
#
# ## Source code for fetch_sdss_galaxy_colors
#
# https://www.astroml.org/_modules/astroML/datasets/sdss_galaxy_colors.html
#
# + colab={"base_uri": "https://localhost:8080/", "height": 381} colab_type="code" id="WUGXmse9rH-R" outputId="9fb71f74-e6f3-4913-b04c-f76f8b69b3a1"
# print_function: make print a function
# division: change the division operator, // for floor division
from __future__ import print_function, division
import os
import sys
import numpy as np
import astroML as am
from astroML.datasets import get_data_home
from astroML.datasets.tools import sql_query
NOBJECTS = 50000
GAL_COLORS_NAMES = ['u', 'g', 'r', 'i', 'z', 'specClass', 'redshift', 'redshift_err']
ARCHIVE_FILE = 'sdss_galaxy_colors.npy' # local file generated
def fetch_sdss_galaxy_colors(data_home=None, download_if_missing=True):
"""Loader for SDSS galaxy colors.
This function directly queries the sdss SQL database at
http://cas.sdss.org/
Parameters
----------
data_home : optional, default=None
Specify another download and cache folder for the datasets. By default
all scikit learn data is stored in '~/astroML_data' subfolders.
download_if_missing : optional, default=True
If False, raise a IOError if the data is not locally available
instead of trying to download the data from the source site.
Returns
-------
data : recarray, shape = (10000,)
record array containing magnitudes and redshift for each galaxy
"""
data_home = get_data_home(data_home)
archive_file = os.path.join(data_home, ARCHIVE_FILE)
query_text = ('\n'.join(("SELECT TOP %i" % NOBJECTS,
" p.u, p.g, p.r, p.i, p.z, s.class, s.z, s.zerr",
"FROM PhotoObj AS p",
" JOIN SpecObj AS s ON s.bestobjid = p.objid",
"WHERE ",
" p.u BETWEEN 0 AND 19.6",
" AND p.g BETWEEN 0 AND 20",
" AND s.class <> 'UNKNOWN'",
" AND s.class <> 'STAR'",
" AND s.class <> 'SKY'",
" AND s.class <> 'STAR_LATE'")))
if not os.path.exists(archive_file):
if not download_if_missing:
raise IOError('data not present on disk. '
'set download_if_missing=True to download')
print("querying for %i objects" % NOBJECTS)
print(query_text)
output = sql_query(query_text)
print("finished.")
kwargs = {'delimiter': ',', 'skip_header': 2,
'names': GAL_COLORS_NAMES, 'dtype': None}
if sys.version_info[0] >= 3:
kwargs['encoding'] = 'ascii'
data = np.genfromtxt(output, **kwargs)
np.save(archive_file, data)
else:
data = np.load(archive_file)
return data
# -
# ## Example - SDSS Galaxy Colors
#
# https://www.astroml.org/examples/datasets/plot_sdss_galaxy_colors.html
#
# The function **fetch_sdss_galaxy_colors()** used below actually queries the SDSS CASjobs server for the colors of the 50,000 galaxies. Below we extract the u - g and g - r colors for 5000 objects, and scatter-plot the results.
#
# Notes:
#
# - Quasars are also commonly referred to as **QSO**s (Quasi-Stellar Objects). <small> *Source: https://github.com/astroML/sklearn_tutorial/blob/master/doc/classification.rst* </small>
# +
# Author: <NAME> <<EMAIL>>
# License: BSD
# The figure is an example from astroML: see http://astroML.github.com
from matplotlib import pyplot as plt
from astroML.datasets import fetch_sdss_galaxy_colors
#------------------------------------------------------------
# Download data
data = fetch_sdss_galaxy_colors() #############
data = data[::10] # truncate for plotting
data
# +
# Extract colors and spectral class
ug = data['u'] - data['g']
gr = data['g'] - data['r']
spec_class = data['specClass']
spec_class
# +
galaxies = (spec_class == 'GALAXY')
galaxies
# +
qsos = (spec_class == 'QSO')
qsos
# +
#------------------------------------------------------------
# Prepare plot
fig = plt.figure(figsize=(6,8))
ax = fig.add_subplot(111)
ax.set_xlim(-0.5, 2.5)
ax.set_ylim(-0.5, 1.5)
ax.plot(ug[galaxies], gr[galaxies], '.', ms=4, c='b', label='galaxies')
ax.plot(ug[qsos], gr[qsos], '.', ms=4, c='r', label='qsos')
ax.legend(loc=2)
ax.set_xlabel('$u-g$')
ax.set_ylabel('$g-r$')
plt.show()
# + [markdown] colab_type="text" id="vrESwpcSsOel"
# ## Lambda CDM
#
# "The ΛCDM (Lambda cold dark matter) or Lambda-CDM model is a parametrization of the Big Bang cosmological model in which the universe contains three major components:
# - a cosmological constant denoted by Lambda (Greek Λ) and associated with dark energy;
# - the postulated cold dark matter (abbreviated CDM);
# - ordinary matter.
#
# It is frequently referred to as the standard model of Big Bang cosmology because it is the simplest model that provides a reasonably good account of the properties of the cosmos". <small> *Source: https://en.wikipedia.org/wiki/Lambda-CDM_model (adapted).* </small>
#
# The _Flat Lambda CDM model_ is used in _astroML.datasets.generate_mu_z_ that generate a **dataset of distance modulus vs redshift**:
# - https://www.astroml.org/modules/generated/astroML.datasets.generate_mu_z.html
#
# Generate a dataset of distance modulus vs redshift.
#
# Parameters
#
# - size : int or tuple = size of generated data
# - z0 : float = parameter in redshift distribution: p(z) ~ (z / z0)^2 exp[-1.5 (z / z0)]
# - dmu_0, dmu_1 : float = specify the error in mu, dmu = dmu_0 + dmu_1 * mu
# - random_state: None, int, or np.random.RandomState instance = random seed or random number generator
# - cosmo : **astropy**.cosmology instance specifying cosmology = to use when generating the sample. If not provided, a Flat Lambda CDM model (**astropy**) with H0=71, Om0=0.27, Tcmb=0 is used.
#
# Returns
#
# - z, mu, dmu : ndarrays = arrays of shape 'size'
#
# ## Astropy.cosmology
#
# https://docs.astropy.org/en/stable/cosmology/
#
# Astropy are utilities to extend or substitute existing astronomical data analysis tools on a modern, object-oriented platform. Among the first projects were a replacement of the command language for the Image Reduction and Analysis Facility (IRAF) with a Python frontend, and the PyFITS interface to the Flexible Image Transport System.
#
# IRAF (Image Reduction and Analysis Facility) is a collection of software written at the National Optical Astronomy Observatory (NOAO) geared towards the reduction of astronomical images in pixel array form. This is primarily data taken from imaging array detectors such as CCDs.
# -
# ## Example - Lambda CDM
#
# Figure 8.11 - Cosmology Regression Example
#
# A Gaussian process regression analysis of the simulated supernova sample. Uses a squared-exponential covariance model, with bandwidth learned through cross-validation.
#
# https://www.astroml.org/book_figures/chapter8/fig_gp_mu_z.html
#
# Two datasets are used:
#
# - fetch_sdss_galaxy_colors: dataset of colors of the 50,000 galaxies
# - generate_mu_z: dataset of distance modulus vs redshift (uses Astropy)
# +
# Author: <NAME>
# License: BSD
# The figure produced by this code is published in the textbook
# "Statistics, Data Mining, and Machine Learning in Astronomy" (2013)
# For more information, see http://astroML.github.com
# To report a bug or issue, use the following forum:
# https://groups.google.com/forum/#!forum/astroml-general
from __future__ import print_function, division
import numpy as np
from matplotlib import pyplot as plt
from sklearn.gaussian_process.kernels import ConstantKernel, RBF
from sklearn.gaussian_process import GaussianProcessRegressor
from astropy.cosmology import LambdaCDM
import astroML as am
from astroML.datasets import fetch_sdss_galaxy_colors
from astroML.datasets import generate_mu_z
# ----------------------------------------------------------------------
# This function adjusts matplotlib settings for a uniform feel in the textbook.
# Note that with usetex=True, fonts are rendered with LaTeX. This may
# result in an error if LaTeX is not installed on your system. In that case,
# you can set usetex to False.
#if "setup_text_plots" not in globals():
# from astroML.plotting import setup_text_plots
#setup_text_plots(fontsize=8, usetex=True)
# ------------------------------------------------------------
# Generate data
cosmo = LambdaCDM(H0=71, Om0=0.27, Ode0=0.73, Tcmb0=0) # Astropy
z_sample, mu_sample, dmu = generate_mu_z(100, random_state=0, cosmo=cosmo)
z = np.linspace(0.01, 2, 1000)
mu_true = cosmo.distmod(z)
# ------------------------------------------------------------
# fit the data
# Mesh the input space for evaluations of the real function,
# the prediction and its MSE
z_fit = np.linspace(0, 2, 1000)
kernel = ConstantKernel(1.0, (1e-3, 1e3)) * RBF(10, (1e-2, 1e2))
gp = GaussianProcessRegressor(kernel=kernel, alpha=dmu ** 2)
gp.fit(z_sample[:, None], mu_sample)
y_pred, sigma = gp.predict(z_fit[:, None], return_std=True)
# ------------------------------------------------------------
# Plot the gaussian process
# gaussian process allows computation of the error at each point
# so we will show this as a shaded region
fig = plt.figure(figsize=(6, 6))
fig.subplots_adjust(left=0.1, right=0.95, bottom=0.1, top=0.95)
ax = fig.add_subplot(111)
ax.plot(z, mu_true, '--k')
ax.errorbar(z_sample, mu_sample, dmu, fmt='.k', ecolor='gray', markersize=6)
ax.plot(z_fit, y_pred, '-k')
ax.fill_between(z_fit, y_pred - 1.96 * sigma, y_pred + 1.96 * sigma,
alpha=0.2, color='b', label='95% confidence interval')
ax.set_xlabel('$z$')
ax.set_ylabel(r'$\mu$')
ax.set_xlim(0, 2)
ax.set_ylim(36, 48)
plt.show()
# + [markdown] colab_type="text" id="OTpS_b58hnYJ"
# ## References
#
# - http://www.astroml.org
# - https://www.sdss.org
# - https://en.wikipedia.org/wiki/Lambda-CDM_model
# - https://www.astropy.org/
# - http://www.sdss.jhu.edu/
| CAP379_Topics_in_Spatial_Science.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
from plotnine import *
from plotnine.data import *
# %matplotlib inline
# -
# ### Density Plot
mpg.head()
# The defaults are not exactly beautiful, but still quite clear.
(ggplot(mpg, aes(x='cty'))
+ geom_density()
)
# Plotting multiple groups is straightforward, but as each group is plotted as an independent PDF summing to 1, the relative size of each group will be normalized.
(ggplot(mpg, aes(x='cty', color='drv', fill='drv'))
+ geom_density(alpha=0.1)
)
# To plot multiple groups and scale them by their relative size, you can map the `y` aesthetic to `'count'` (calculated by `stat_density`).
(ggplot(mpg, aes(x='cty', color='drv', fill='drv'))
+ geom_density(aes(y=after_stat('count')), alpha=0.1)
)
# ### Density Plot + Histogram
#
# To overlay a histogram onto the density, the `y` aesthetic of the density should be mapped to the `'count'` scaled by the `binwidth` of the histograms.
#
# **Why?**
#
# The `count` calculated by `stat_density` is $count = density * n$ where `n` is the number of points . The `density` curves have an area of 1 and have no information about the absolute frequency of the values along curve; only the relative frequencies. The `count` curve reveals the absolute frequencies. The scale of this `count` corresponds to the `count` calculated by the `stat_bin` for the histogram when the bins are 1 unit wide i.e. `binwidth=1`. The `count * binwidth` curve matches the scale of `count`s for the histogram for a give `binwidth`.
# +
binwidth = 2 # The same for geom_density and geom_histogram
(ggplot(mpg, aes(x='cty', color='drv', fill='drv'))
+ geom_density(aes(y=after_stat('count*binwidth')), alpha=0.1)
+ geom_histogram(aes(fill='drv', y=after_stat('count')), binwidth=binwidth, color='none', alpha=0.5)
# It is the histogram that gives us the meaningful y axis label
# i.e. 'count' and not 'count*2'
+ labs(y='count')
)
| demo_plot/plotnine-examples/examples/geom_density.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="LikIUYuS_DyQ"
# #### GoogLeNet
#
# This notebook is an implement of [___Going deeper with convolutions___](https://arxiv.org/pdf/1409.4842.pdf) by Szedegy et al. The original model was trained for ImageNet dataset, but in this notebook we fine-tuned it for Cifar 10 dataset, which is a relatively smaller dataset and thus takes less space to store on clusters and less time for training.
# + [markdown] id="MfJAV5slGQq_"
# We first need to install and import all the dependent libraries in the session.
# + colab={"base_uri": "https://localhost:8080/"} id="aVhDJW4GGVUT" outputId="66856870-e9da-4eab-d752-29b82882efed"
import matplotlib.pyplot as plt
import numpy as np
import tensorflow as tf
from tensorflow.compat.v1 import ConfigProto
from tensorflow.compat.v1 import InteractiveSession
from tensorflow.keras.layers import *
from tensorflow.keras.regularizers import l2
from tensorflow.keras import Model
import tensorflow_datasets as tfds
from sklearn.model_selection import train_test_split
config = ConfigProto()
config.gpu_options.allow_growth = True
session = InteractiveSession(config=config)
# + [markdown] id="NLaTochAFfm8"
# In this part of the program, we get the Cifar 10 dataset using tensorflow dataset and separate it into training set and test set.
# + colab={"base_uri": "https://localhost:8080/", "height": 418, "referenced_widgets": ["ef34eedb880e47f0aafadb3120aea380", "5b03611d6e4f413880d2f78e8af48660", "86dcb6a2cce648308ce633ce58c823bf", "<KEY>", "1f09dfc329ee4495ae7c6e02c5ac4f20", "754dcdf1f0014f9e8ff92d079300a4b6", "<KEY>", "f02b2aed8dee418c970acd806f6b44e3", "<KEY>", "<KEY>", "584d88dea75345b6ae9a7d77bc9f21b0", "<KEY>", "8a306137fd9a47d5b2714e40c3437921", "<KEY>", "<KEY>", "454a4447a407462ca644bc81137ed5f6", "bb44a87e7b0645188471745eb88a96b3", "1fd992ac91ea4405af96725a54cf67d8", "91b0df07c4aa48fd96490e89989c32a4", "<KEY>", "<KEY>", "f3c09692f9c74c589f7f0efe7d85f037", "<KEY>", "bedd3e4bdc974d588cd706f7aa5c822e", "<KEY>", "0506b1f3ffe045af8df7624b765e4058", "388ce343e4414db794323d3d1ed8284a", "8803ce07a20340f587fbc19e3255c139", "<KEY>", "f2d4da8079be4ac9b4e843fa60f4e64a", "db765227a4a74fd0ab29bed7b5f9a9f9", "4800278ddb4746aebaefedaed79f3579", "918cc0a5f0e14470b6ee30fd91ee6f01", "4b3d25b04a2242f3aea9f8b5cc789e62", "0528db7a4e6c4b7e9eceeefc7ae5b10e", "b20ec0d20c74431eaff3354b7de34266", "<KEY>", "<KEY>", "d3a53533ffab4a76ac9ada9169bfff28", "<KEY>", "81e8f533e9134a1b8075f123ff1b7358", "2e27bbd74a24414b86e1da4a0ef640c8", "<KEY>", "c1c992efae044df39517b5c4973cef27", "<KEY>", "<KEY>", "b1eef36a9ffc41b6baf4e1e694ed69ac", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "1ae399483646413eb54d43c50ead2be8", "edca3d397f2344a6b17ce9ae8d57ab34", "2762f59788ad4aed92a7dffebec14ac2", "e150ae3eeddc42498e5a5d792093eecd", "<KEY>", "75bbb328dbef4f8f8e106663ff6dd1c7", "<KEY>", "0ffd2946869e4ba59d0e9c1c4c4e94eb", "70194de5ff26462c923dfd8ed271e114", "97b4ab535dc84e2391b567c17250a566", "<KEY>", "1b67eaff6ad144db93eab03e3fe4e679", "<KEY>", "<KEY>", "5fb22e29634c4e2a9d7b2688536effef", "7a52b439688a44cba259f55fe3b7aa45", "fdc273dbf7744e18b54eea536fada0f7", "8fb6c52e547c40ac9066ef0c21507777", "00547a582ab94234a28584c5447e5ce7", "<KEY>", "<KEY>", "<KEY>", "7bad3800395d4abebcc3df2c2ac9a9f5"]} id="B_ecmr8NFtIM" outputId="18bcff8d-d0d5-4a6c-c4a8-053ae4a8a5de"
def get_data():
train, test = tfds.as_numpy(tfds.load("cifar10", split = ["train", "test"]
, shuffle_files = True, as_supervised = True, batch_size = -1))
return train[0], test[0], train[1], test[1]
train_x, test_x, train_y, test_y = get_data()
train_x = train_x / 255.0
test_x = test_x / 255.0
# + [markdown] id="WK2DZExoAPVk"
# This is function that constructs a GoogLeNet model. In ```inception``` function we constructs an inception block while in ```createGoogLeNet``` function we build a fine-tuned GoogLeNet model. The structure of the model is almost same with the original paper, but the kernel size and strides are adjusted to fit the smaller pictures of Cifar 10. The most important change is that we only keep four inception layers because the images are very small and deep network would only cause overfitting instead of increasing performance. We define ```weight_decay``` as the hyperparameters of the model for kernel regularization.
# + id="71DVAiu4KFjz"
def inception(input, f1, f3r, f3, f5r, f5, pp, weight_decay, output = False, output_name = None):
x1 = Conv2D(kernel_size = 1, filters = f1, activation = "relu", padding = "same", kernel_regularizer = l2(weight_decay))(input)
x2 = Conv2D(kernel_size = 1, filters = f3r, activation = "relu", padding = "same", kernel_regularizer = l2(weight_decay))(input)
x2 = BatchNormalization()(x2)
x2 = Conv2D(kernel_size = 3, filters = f3, activation = "relu", padding = "same", kernel_regularizer = l2(weight_decay))(x2)
x3 = Conv2D(kernel_size = 1, filters = f5r, activation = "relu", padding = "same", kernel_regularizer = l2(weight_decay))(input)
x3 = BatchNormalization()(x3)
x3 = Conv2D(kernel_size = 5, filters = f5, activation = "relu", padding = "same", kernel_regularizer = l2(weight_decay))(x3)
x4 = MaxPooling2D(pool_size = 3, padding = "same", strides = 1)(input)
x4 = Conv2D(kernel_size = 1, filters = pp, activation = "relu", padding = "same", kernel_regularizer = l2(weight_decay))(x4)
x5 = None
if output == True:
x5 = GlobalAveragePooling2D()(x)
x5 = Conv2D(kernel_size = 1, padding = "same", filters = 128, activation = "relu", kernel_regularizer = l2(weight_decay))(x5)
x5 = BatchNormalization()(x5)
x5 = Flatten()(x5)
x5 = Dense(1024, activation = "relu", kernel_regularizer = l2(weight_decay))(x5)
x5 = Dropout(rate = .7)(x5)
x5 = Dense(10, activation = "softmax", name = output_name)(x5)
x = Concatenate()([x1, x2, x3, x4])
x = BatchNormalization()(x)
x = Dropout(.3)(x)
return x, x5
def createGoogLeNet(weight_decay):
input = Input(shape = (32, 32, 3), name = "image")
x, _ = inception(input, 64, 96, 128, 16, 32, 32, weight_decay)
x, _ = inception(x, 128, 128, 192, 32, 96, 64, weight_decay)
x, _ = inception(x, 192, 96, 208, 16, 48, 64, weight_decay)
x, _ = inception(x, 160, 112, 224, 24, 64, 64, weight_decay)
x = GlobalAveragePooling2D()(x)
x = Flatten()(x)
x = Dropout(rate = .7)(x)
output = Dense(10, activation = "softmax", name = "out")(x)
model = tf.keras.Model(inputs = [input], outputs = [output], name = "GoogLeNet")
return model
# + [markdown] id="L7dpVq72RSei"
# This part trains the GoogLeNet model on Cifar 10 dataset. We tested several sets of hyperparameters and adopted one with the best validation loss. We then store the best weights of each training epochs on server so that we can continue training even if the session disconnects. We also add learning rate scheduler to slow down learning rate when validation loss stops increasing. We show the result of the training process with a graph about the training and validation accuracy for each epoch.
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="FaLmu9n5ROzk" outputId="a80fc0d9-c1e7-4a5e-a5ab-3705804797b9"
# Set a checkpoint to save weights
cp = tf.keras.callbacks.ModelCheckpoint("weights", monitor = "val_loss", verbose = 1, save_best_only = True, mode = "auto")
lr = tf.keras.callbacks.ReduceLROnPlateau(monitor='val_loss', factor=0.1, patience=5, verbose=0, mode='auto', min_delta=0.0001, cooldown=0, min_lr=0)
model = createGoogLeNet(.1)
model.compile(optimizer = tf.keras.optimizers.Adam(learning_rate = 5e-5),
loss = tf.keras.losses.SparseCategoricalCrossentropy(),
metrics = ["accuracy"])
# We can use the existing data if the training process has started
# model.load_weights("weights")
history = model.fit({"image": train_x}, {"out": train_y}, epochs = 50, validation_split = .2, callbacks = [cp, lr], batch_size = 64)
plt.plot(history.history['accuracy'])
plt.plot(history.history['val_accuracy'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'val'], loc = 'upper left')
plt.show()
# + [markdown] id="nUhdLp-cB1_G"
# Here we test our model on test set and show how GoogLeNet predicts on sample images in the test set.
# + colab={"base_uri": "https://localhost:8080/", "height": 409} id="ok-a_tSrKW8w" outputId="d6cfd62e-84d6-4efc-f979-b43c1c9c557c"
labels = ["airplane", "automobile", "bird", "cat", "deer", "dog", "frog", "horse", "ship", "truck"]
pred = np.argmax(model.predict(test_x), axis = 1)
print("Test Accuracy: {:.2%}".format(np.sum(pred == test_y) / len(test_y)))
sample_data = test_x[: 9]
sample_label = test_y[: 9]
fig = plt.figure(figsize = (10, 40))
for i in range(len(sample_data)):
ax = fig.add_subplot(911 + i)
ax.imshow(test_x[i])
ax.set_title("Labelled as " + labels[int(sample_label[i])] + ", classified as " + labels[int(pred[i])])
| cv/GoogLeNet/GoogLeNet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.10 64-bit
# name: python3
# ---
# # comparison
#
# > AquaCrop-OSP | AquaCrop-OS | AquaCrop | Test Comparison
# > aquacrop python is tested against windows aquacrop and matlab to ensure
#
# ## Excersise 7: Wheat production in Tunis
#
# ### From Chapter 7 of <a href="../pdfs/AquaCrop_TrainingHandookB.pdf">AquaCrop Training Handbook</a> (pg. 64)
# ### Description
#
# This set of exercises will assess the average winter wheat production that can be expected in
# Tunis (Tunisia) under different environmental conditions and agronomic practices (local crop
# varieties, soils, management). Both rainfed and irrigated cropping systems will be evaluated.
#
#
# The capital Tunis is located in north Tunisia (36°48’N, 10°10’E) and has a semi-arid climate
# (Fig. 7.1). Rainfed crops can be cultivated between October and March, when most rainfall
# events take place and ET 0 is relatively low. In addition, irrigated crops can be cultivated at other
# times in the year, even in the summer months when no rainfall occurs.
# <img src="images/tunis_weather_pic.png" width="500" height="250"/>
# ## Exercise 7.1: Assessing crop yield for local soils
#
# Determine the average winter wheat yield that can be expected in the region of Tunis when soil
# fertility is non-limiting. Run therefore a series of 23 years of historical weather data (1979-
# 2002). According to local practices, farmers sow wheat at the beginning of the rainy season,
# assume on 15 October. Assess the yield on a sandy loam soil and on a local soil.
# Assume that on the day of sowing the soil is wetted at the top, but still dry in deeper soil layers.
# ### Imports
# +
# # !pip install aquacrop
# from google.colab import output
# output.clear()
# +
#export
# import sys
# _=[sys.path.append(i) for i in ['.', '..']]
# +
# #hide
# # %load_ext autoreload
# # %autoreload 2
# -
#export
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from aquacrop.core import *
from aquacrop.classes import *
import seaborn as sns
from aquacrop.crops.crop_params import crop_params
# ### Run comparison
# +
#model.Outputs.Final
# -
#export
def run_comparison(model,name):
"""
Function to run a comparison between python matlab and windows.
Plots yields and prints mean and mean absolute error between them
*Arguments:*
`name`: `str` : name of directory containing input files
*Returns:*
None
"""
Outputs = model.Outputs
py = Outputs.Final.round(3)
py.columns = ["Season","CropType","HarvestDate","Harvest Date (Step)","Yield","Seasonal irrigation (mm)"]
matlab = pd.read_csv(get_filepath(name+'_matlab.txt'),delim_whitespace=True,header=None)
matlab.columns = ["season","crop","plantdate","stepplant","harvestdate","stepharvest","Yield","tirr"]
windows_names = (' RunNr Day1 Month1 Year1 Rain ETo GD CO2 Irri Infilt Runoff Drain Upflow E E/Ex Tr TrW Tr/Trx SaltIn SaltOut SaltUp SaltProf Cycle SaltStr FertStr WeedStr TempStr ExpStr StoStr BioMass Brelative HI Yield WPet DayN MonthN YearN'.split())
windows = pd.read_csv(get_filepath(name+'_windows.OUT'),skiprows=5,delim_whitespace=True,names=windows_names,encoding="ISO-8859-1")
combined = pd.DataFrame([py.Yield,windows.Yield,matlab.Yield]).T
combined.columns = ["py", "windows","matlab"]
mae = np.round(np.abs(combined.py - combined.windows).mean(),2)
pymean = combined.mean().py.round(2)
print(f'python seasonal mean: {pymean} kg/ha\nMAE from windows: {mae} kg/ha')
mae_mat = np.round(np.abs(combined.py - combined.matlab).mean(),3)
print(f'MAE from matlab: {mae_mat} kg/ha')
plt.style.use('seaborn')
fig, ax = plt.subplots(2,1,sharex=True,figsize=(11,8))
ax[0].plot(py.Yield,label='Python')
ax[0].plot(matlab.Yield,label='Matlab')
ax[0].plot(windows.Yield,'--',label='Windows')
ax[0].legend(fontsize = 18)
ax[0].set_ylabel('Yield',fontsize=18)
#sns.jointplot(np.arange(len(py)), py.Yield - windows.Yield,
# kind="resid",color="m",ratio=10)
ax[1].scatter(np.arange(len(py)),py.Yield - windows.Yield,label='Python')
ax[1].scatter(np.arange(len(py)),matlab.Yield - windows.Yield,label='Matlab')
ax[1].plot([0,len(py)],[0,0],'--',color='black')
ax[1].set_xlabel('Season',fontsize=18)
ax[1].set_ylabel('Residuals',fontsize=18)
ax[1].legend(fontsize = 18)
plt.show()
return Outputs,windows
# +
wdf = prepare_weather(get_filepath('tunis_climate.txt'))
wdf.Date.min(),wdf.Date.max()
soil = SoilClass('ac_TunisLocal')
crop = CropClass('WheatGDD',PlantingDate= '10/15')
iwc = InitWCClass('Num','Depth',[0.3,0.9],[0.3,0.15])
model = AquaCropModel('1979/10/15','2002/05/31',wdf,soil,crop,InitWC=iwc)
model.initialize()
# %time model.step(till_termination=True)
# -
res = run_comparison(model,'tunis_test_1')
# +
#hide
#(model.Outputs.Flux/model.Outputs.Flux.max()).plot()
# res[0].Flux.groupby('SeasonCounter')["P"].sum().plot()
# res[1].Rain.plot()
#model.Outputs.Final
# res[0].Flux.columns
# step=res[0].Final.Step.values
# var = "B"; df = res[0].Growth[res[0].Growth.TimeStepCounter.isin(step)]
# df=df[var]
# plt.plot(df.values)
# (100*res[1]["Tr"]).plot()
# res[0].Flux.groupby('SeasonCounter').sum()["DeepPerc"].plot()
# (1*res[1]["Irri"]).plot()
# res[1].columns
# res[0].Growth.columns
# res[0].Growth.Zroot
# res[1][['ExpStr', 'StoStr']].plot()
# res[0].Growth.groupby('SeasonCounter').max()["CC"].plot()
# res[0].Growth.groupby('SeasonCounter').max()["CC_NS"].plot()
# -
# ### Sandy loam soil
sandy_loam = SoilClass('SandyLoam')
model = AquaCropModel('1979/01/01','2002/05/31',wdf,sandy_loam,crop,InitWC=iwc)
model.initialize()
model.step(till_termination=True)
res = run_comparison(model,'tunis_test_1_SandyLoam')
# ## Excercise 7.2 Local Wheat variety
local_wheat = CropClass('WheatGDD',PlantingDate= '10/15',
Emergence=289,
MaxRooting = 1322,
Senescence = 2835,
Maturity = 3390,
HIstart = 2252,
Flowering = 264,
YldForm = 1073,
PlantPop=3_500_000,
CCx=0.9,
CDC=0.003888,
CGC=0.002734)
model = AquaCropModel('1979/01/01','2002/05/31',wdf,sandy_loam,local_wheat,InitWC=iwc)
model.initialize()
model.step(till_termination=True)
_ = run_comparison(model,'tunis_test_2_long')
# ## Excercise 7.3 Different initial conditions
iwc30taw = InitWCClass('Pct','Layer',[1],[30])
model = AquaCropModel('1979/01/01','2002/05/31',wdf,sandy_loam,
crop,InitWC=iwc30taw)
model.initialize()
model.step(till_termination=True)
_ = run_comparison(model,'tunis_test_3_30taw')
# ## Excercise 7.6 Net irrigation requirement
net_irr = IrrMngtClass(IrrMethod=4,NetIrrSMT=78.26)
wp = InitWCClass(value=['WP'])
wheat_dec = CropClass('WheatGDD',PlantingDate='12/01',HarvestDate='07/30')
model = AquaCropModel('1979/08/15','2001/07/30',wdf,sandy_loam,
wheat_dec,InitWC=wp,IrrMngt=net_irr)
model.initialize()
model.step(till_termination=True)
res = run_comparison(model,'tunis_test_6')
# ## Excersise 8 : Hyderabad : Chapter 8 of <a href="../pdfs/AquaCrop_TrainingHandookB.pdf">AquaCrop Training Handbook</a>
#
# ## Excersise 8.1
# +
wdf = prepare_weather(get_filepath('hyderabad_climate.txt'))
wdf.Date.min(),wdf.Date.max()
# -
rice = CropClass('localpaddy',PlantingDate= '08/01',)
paddy = SoilClass('Paddy')
iwc_paddy = InitWCClass(depth_layer=[1,2],value=['FC','FC'])
fm = FieldMngtClass(Bunds=True,zBund=0.2)
model = AquaCropModel('2000/01/01','2010/12/31',wdf,paddy,
rice,InitWC=iwc_paddy,FieldMngt=fm,
FallowFieldMngt=fm)
model.initialize()
model.step(till_termination=True)
_ = run_comparison(model,'paddyrice_hyderabad')
# ## Excersise 9 : Brussels : Chapter 9 of <a href="../pdfs/AquaCrop_TrainingHandookB.pdf">AquaCrop Training Handbook</a>
# ### Excersise 9.1
# +
wdf = prepare_weather(get_filepath('brussels_climate.txt'))
wdf.Date.min(),wdf.Date.max()
# -
potato = CropClass('PotatoLocal',PlantingDate= '04/25')
loam = SoilClass('Loam')
model = AquaCropModel('1976/01/01','2005/12/31',wdf,loam,potato,InitWCClass())
model.initialize()
model.step(till_termination=True)
_ = run_comparison(model,'potato')
| docs/notebooks/05_comparison.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # December 2016: Advent of Code Solutions
#
# ## <NAME>
#
# From Dec. 1 to Dec. 25, [I](http://norvig.com) will be solving the puzzles that appear each day at *[Advent of Code](http://adventofcode.com/)*. The two-part puzzles are released at midnight EST (9:00PM PST); points are awarded to the first 100 people to solve the day's puzzles. The code shown here basically represents what I did to solve the problem, but slightly cleaned up:
# - On days when I start at 9:00PM and am competing against the clock, I take shortcuts. I use shorter names, because I'm not a fast typist. I run test cases in the Jupyter Notebook, but don't make them into `assert` statements. Even then, I'm not really competitive with the fastest solvers.
# - On days when I didn't get a chance to start until after all the points are gone, what you see here is pretty much exactly what I did, or at least what I ended up with after correcting typos and other errors.
#
# To understand the problems completely, you will have to read the full description in the **"[Day 1](http://adventofcode.com/2016/day/1):"** link in each day's section header.
#
# # Day 0: Getting Ready
#
# On November 30th, I spent some time preparing:
#
# - I'll import my favorite modules and functions, so I don't have to do it each day.
#
# - From looking at [last year's](http://adventofcode.com/2015) puzzles, I knew that there would be a data file on many days, so I defined the function `Input` to open the file (and for those using this notebook on a remote machine, to fetch the file from the web). My data files are at [http://norvig.com/ipython/advent2016/](http://norvig.com/ipython/advent2016/).
#
# - From working on another puzzle site, [Project Euler](https://projecteuler.net/), I had built up a collection of utility functions, shown below:
# +
# Python 3.x
import re
import numpy as np
import math
import urllib.request
from collections import Counter, defaultdict, namedtuple, deque
from functools import lru_cache
from itertools import permutations, combinations, chain, cycle, product, islice
from heapq import heappop, heappush
def Input(day):
"Open this day's input file."
filename = 'advent2016/input{}.txt'.format(day)
try:
return open(filename)
except FileNotFoundError:
return urllib.request.urlopen("http://norvig.com/ipython/" + filename)
def transpose(matrix): return zip(*matrix)
def first(iterable): return next(iter(iterable))
def nth(iterable, n, default=None):
"Returns the nth item of iterable, or a default value"
return next(islice(iterable, n, None), default)
cat = ''.join
Ø = frozenset() # Empty set
inf = float('inf')
BIG = 10 ** 999
def grep(pattern, lines):
"Print lines that match pattern."
for line in lines:
if re.search(pattern, line):
print(line)
def groupby(iterable, key=lambda it: it):
"Return a dic whose keys are key(it) and whose values are all the elements of iterable with that key."
dic = defaultdict(list)
for it in iterable:
dic[key(it)].append(it)
return dic
def powerset(iterable):
"Yield all subsets of items."
items = list(iterable)
for r in range(len(items)+1):
for c in combinations(items, r):
yield c
# 2-D points implemented using (x, y) tuples
def X(point): return point[0]
def Y(point): return point[1]
def neighbors4(point):
"The four neighbors (without diagonals)."
x, y = point
return ((x+1, y), (x-1, y), (x, y+1), (x, y-1))
def neighbors8(point):
"The eight neighbors (with diagonals)."
x, y = point
return ((x+1, y), (x-1, y), (x, y+1), (x, y-1),
(x+1, y+1), (x-1, y-1), (x+1, y-1), (x-1, y+1))
def cityblock_distance(p, q=(0, 0)):
"City block distance between two points."
return abs(X(p) - X(q)) + abs(Y(p) - Y(q))
def euclidean_distance(p, q=(0, 0)):
"Euclidean (hypotenuse) distance between two points."
return math.hypot(X(p) - X(q), Y(p) - Y(q))
def trace1(f):
"Print a trace of the input and output of a function on one line."
def traced_f(*args):
result = f(*args)
print('{}({}) = {}'.format(f.__name__, ', '.join(map(str, args)), result))
return result
return traced_f
def astar_search(start, h_func, moves_func):
"Find a shortest sequence of states from start to a goal state (a state s with h_func(s) == 0)."
frontier = [(h_func(start), start)] # A priority queue, ordered by path length, f = g + h
previous = {start: None} # start state has no previous state; other states will
path_cost = {start: 0} # The cost of the best path to a state.
while frontier:
(f, s) = heappop(frontier)
if h_func(s) == 0:
return Path(previous, s)
for s2 in moves_func(s):
new_cost = path_cost[s] + 1
if s2 not in path_cost or new_cost < path_cost[s2]:
heappush(frontier, (new_cost + h_func(s2), s2))
path_cost[s2] = new_cost
previous[s2] = s
return dict(fail=True, front=len(frontier), prev=len(previous))
def Path(previous, s):
"Return a list of states that lead to state s, according to the previous dict."
return ([] if (s is None) else Path(previous, previous[s]) + [s])
# -
# Some tests/examples for these:
assert tuple(transpose(((1, 2, 3), (4, 5, 6)))) == ((1, 4), (2, 5), (3, 6))
assert first('abc') == first(['a', 'b', 'c']) == 'a'
assert cat(['a', 'b', 'c']) == 'abc'
assert (groupby(['test', 'one', 'two', 'three', 'four'], key=len)
== {3: ['one', 'two'], 4: ['test', 'four'], 5: ['three']})
# # [Day 1](http://adventofcode.com/2016/day/1): No Time for a Taxicab
#
# Given a sequence of moves, such as `"R2, L3"`, which means turn 90° to the right and go forward 2 blocks, then turn 90° left and go 3 blocks, how many blocks do we end up away from the start? I make the following choices:
# * **Intersection Points** in the city grid will be represented as points on the complex plane.
# * **Headings and turns** can be represented by unit vectors in the complex plane: if you are heading east (along the positive real axis), then a left turn means you head north, and a right turn means you head south, and [in general](https://betterexplained.com/articles/understanding-why-complex-multiplication-works/) a left or right turn is a multiplication of your current heading by the `North` or `South` unit vectors, respectively.
# * **Moves** of the form `"R53"` will be parsed into a `(turn, distance)` pair, e.g. `(South, 53)`.
#
# To solve the puzzle with the function `how_far(moves)`, I initialize the starting location as the origin and the starting heading as North, and follow the list of moves, updating the heading and location on each step, before returning the distance from the final location to the origin.
# +
Point = complex
N, S, E, W = 1j, -1j, 1, -1 # Unit vectors for headings
def distance(point):
"City block distance between point and the origin."
return abs(point.real) + abs(point.imag)
def how_far(moves):
"After following moves, how far away from the origin do we end up?"
loc, heading = 0, N # Begin at origin, heading North
for (turn, dist) in parse(moves):
heading *= turn
loc += heading * dist
return distance(loc)
def parse(text):
"Return a list of (turn, distance) pairs from text of form 'R2, L42, ...'"
turns = dict(L=N, R=S)
return [(turns[RL], int(d))
for (RL, d) in re.findall(r'(R|L)(\d+)', text)]
assert distance(Point(3, 4)) == 7 # City block distance; Euclidean distance would be 5
assert parse('R2, L42') == [(S, 2), (N, 42)]
assert how_far("R2, L3") == 5
assert how_far("R2, R2, R2") == 2
assert how_far("R5, L5, R5, R3") == 12
how_far(Input(1).read())
# -
# In **part two** of this puzzle, I have to find the first point that is visited twice. To support that, I keep track of the set of visited points. My first submission was wrong, because I didn't consider that the first point visited twice might be in the middle of a move, not the end, so I added the "`for i`" loop to iterate over the path of a move, one point at a time.
# +
def visited_twice(text):
"Following moves in text, find the first location we visit twice, and return the distance to it."
loc, heading = 0, N # Begin at origin, heading North
visited = {loc}
for (turn, dist) in parse(text):
heading *= turn
for i in range(dist):
loc += heading
if loc in visited:
return distance(loc)
visited.add(loc)
assert visited_twice("R8, R4, R4, R8") == 4
assert visited_twice("R8, R4, R4, L8") == None
assert visited_twice("R8, R0, R1") == 7
visited_twice(Input(1).read())
# -
# # [Day 2](http://adventofcode.com/2016/day/2): Bathroom Security
#
# Given instructions in the form of a sequence of Up/Down/Right/Left moves, such as `'ULL'`, output the keys on the bathroom lock keypad that the instructions correspond to. Start at the 5 key. Representation choices:
# * **Keypad**: a keypad is an array of strings: `keypad[y][x]` is a key. The character `'.'` indicates a location that is `off` the keypad; by surrounding the keys with a border of `off` characters, I avoid having to write code that checks to see if we hit the edge.
# * **Key**: A key is a character other than `'.'`.
# * **Instructions**: A sequence of lines of `"UDRL"` characters, where each line leads to the output of one key.
# +
Keypad = str.split
keypad = Keypad("""
.....
.123.
.456.
.789.
.....
""")
assert keypad[2][2] == '5'
off = '.'
def decode(instructions, x=2, y=2):
"""Follow instructions, keeping track of x, y position, and
yielding the key at the end of each line of instructions."""
for line in instructions:
for C in line:
x, y = move(C, x, y)
yield keypad[y][x]
def move(C, x, y):
"Make the move corresponding to this character (L/R/U/D)"
if C == 'L' and keypad[y][x-1] is not off: x -= 1
elif C == 'R' and keypad[y][x+1] is not off: x += 1
elif C == 'U' and keypad[y-1][x] is not off: y -= 1
elif C == 'D' and keypad[y+1][x] is not off: y += 1
return x, y
assert move('U', 2, 2) == (2, 1)
assert move('U', 2, 1) == (2, 1)
assert cat(decode("ULL RRDDD LURDL UUUUD".split())) == '1985'
cat(decode(Input(2)))
# -
# In **part two**, we have to deal with a different keypad. I won't need any new functions, but I will need to redefine the global variable `keypad`, and provide `decode` with the new `x` and `y` coordinates of the `5` key:
# +
keypad = Keypad("""
.......
...1...
..234..
.56789.
..ABC..
...D...
.......
""")
assert keypad[3][1] == '5'
cat(decode(Input(2), x=1, y=3))
# -
# # [Day 3](http://adventofcode.com/2016/day/3): Squares With Three Sides
#
# From a file of numbers, three to a line, count the number that represent valid triangles; that is, numbers that satisfy the [triangle inequality](https://en.wikipedia.org/wiki/Triangle_inequality).
# +
def is_triangle(sides):
"Do these side lengths form a valid triangle?"
x, y, z = sorted(sides)
return z < x + y
def parse_ints(text):
"All the integers anywhere in text."
return [int(x) for x in re.findall(r'\d+', text)]
triangles = [parse_ints(line) for line in Input(3)]
sum(map(is_triangle, triangles))
# -
# In **part two**, the triangles are denoted not by three sides in the same line, but by three sides in the same column. For example, given the text:
#
# 101 301 501
# 102 302 502
# 103 303 503
# 201 401 601
# 202 402 602
# 203 403 603
#
# The triangles are:
#
# [101, 102, 103]
# [301, 302, 303]
# [501, 502, 503]
# [201, 202, 203]
# [401, 402, 403]
# [601, 602, 603]
#
# The task is still to count the number of valid triangles.
# +
def invert(triangles):
"Take each 3 lines and transpose them."
for i in range(0, len(triangles), 3):
yield from transpose(triangles[i:i+3])
sum(map(is_triangle, invert(triangles)))
# -
# # [Day 4](http://adventofcode.com/2016/day/4): Security Through Obscurity
#
# Given a list of room names like `"aaaaa-bbb-z-y-x-123[abxyz]"`, consisting of an encrypted name followed by a dash, a sector ID, and a checksum in square brackets, compute the sum of the sectors of the valid rooms. A room is valid if the checksum is the five most common characters, in order (ties listed in alphabetical order).
# +
def parse(line):
"Return (name, sector, checksum)."
return re.match(r"(.+)-(\d+)\[([a-z]+)\]", line).groups()
def sector(line):
"Return the sector number if valid, or 0 if not."
name, sector, checksum = parse(line)
return int(sector) if valid(name, checksum) else 0
def valid(name, checksum):
"Determine if name is valid according to checksum."
counts = Counter(name.replace('-', ''))
# Note: counts.most_common(5) doesn't work because it breaks ties arbitrarily.
letters = sorted(counts, key=lambda L: (-counts[L], L))
return checksum == cat(letters[:5])
assert parse('aaaaa-bbb-z-y-x-123[abxyz]') == ('aaaaa-bbb-z-y-x', '123', 'abxyz')
assert sector('aaaaa-bbb-z-y-x-123[abxyz]') == 123
assert valid('aaaaa-bbb-z-y-x', 'abxyz')
sum(map(sector, Input(4)))
# -
# Initially I had a bug: I forgot the `name.replace('-', '')` to make sure that we don't count hyphens.
#
# In **part two**, we are asked *"What is the sector ID of the room where North Pole objects are stored?"* We are told that names are to be decrypted by a shift cipher, shifting each letter forward in the alphabet by the sector number.
# +
def decrypt(line):
"Decrypt the line (shift the name by sector; discard checksum)."
name, sector, _ = parse(line)
return shift(name, int(sector)) + ' ' + sector
def shift(text, N, alphabet='abcdefghijklmnopqrstuvwxyz'):
"Shift cipher: letters in text rotate forward in alphabet by N places."
N = N % len(alphabet)
tr = str.maketrans(alphabet, alphabet[N:] + alphabet[:N])
return text.translate(tr)
assert shift('hal', 1) == 'ibm'
assert shift('qzmt-zixmtkozy-ivhz', 343) == 'very-encrypted-name'
grep("north", map(decrypt, Input(4)))
# -
# # [Day 5](http://adventofcode.com/2016/day/5): How About a Nice Game of Chess?
#
# This puzzle involves md5 hashes and byte encodings; it took me a while to look up how to do that. What I have to do, for integers starting at 0, is concatenate my door ID string with the integer, get the md5 hex hash, and if the first five digits of the hash are 0, collect the sixth digit; repeat until I have collected eight digits:
# +
import hashlib
door = "ffykfhsq"
def find_password(door):
"First 8 sixth digits of md5 hashes of door+i that begin with '00000'."
password = ''
for i in range(BIG):
x = hashlib.md5(bytes(door + str(i), 'utf-8')).hexdigest()
if x.startswith('00000'):
password += x[5]
print(i, x, password) # Just to see something happen
if len(password) == 8:
return password
find_password(door)
# -
# In **part two**, the sixth digit of the hash that starts with `'00000'` is to be treated as an index that tells where in the password to place the *seventh* digit of the hash. For example, if the sixth digit is `2` and the seventh digit is `a`, then place `a` as the second digit of the final password. Do nothing if the sixth digit is not less than 8, or if a digit has already been placed at that index location.
# +
def find_tougher_password(door):
"For md5 hashes that begin with '00000', the seventh digit goes in the sixth-digit slot of the password."
password = [off] * 8
for i in range(BIG):
x = hashlib.md5(bytes(door + str(i), 'utf-8')).hexdigest()
if x.startswith('00000'):
index = int(x[5], 16)
if index < 8 and password[index] is off:
password[index] = x[6]
print(i, x, cat(password)) # Just to see something happen
if off not in password:
return cat(password)
# %time find_tougher_password(door)
# -
# # [Day 6](http://adventofcode.com/2016/day/6): Signals and Noise
#
# Given a file, where each line is a string of letters and every line has the same length, find the most common letter in each column. We can easily do this with the help of `Counter.most_common`. (Note I use `Input(6).read().split()` instead of just `Input(6)` so that I don't get the `'\n'` at the end of each line.)
counts = [Counter(col) for col in transpose(Input(6).read().split())]
cat(c.most_common(1)[0][0] for c in counts)
# Just to make it clear, here's how we ask for the most common character (and its count) in the first column:
c = counts[0]
c.most_common(1)
# And here is how we pick out the `'t'` character:
c.most_common(1)[0][0]
# In **part two**, we ask for the *least* common character in each column. Easy-peasy:
cat(c.most_common()[-1][0] for c in counts)
# # [Day 7](http://adventofcode.com/2016/day/7): Internet Protocol Version 7
#
# Given input lines of the form `'abcd[1234]fghi[56789]zz[0]z'`, count the number of lines that are a *TLS*, meaning they have an *ABBA* outside of square brackets, but no *ABBA* inside brackets. An *ABBA* is a 4-character subsequence where the first two letters are the same as the last two, but not all four are the same.
#
# I assume brackets are in proper pairs, and are never nested. Then if I do a `re.split` on brackets, the even-indexed pieces of the split will be outside the brackets, and the odd-indexed will be inside. For example:
# - Given the line `'abcd[1234]fghi[56789]zz'`
# - Split on brackets to get `['abcd', '1234', 'fghi', '56789', 'zz']`
# - Outsides of brackets are `'abcd, fghi, zz'` at indexes 0, 2, 4.
# - Insides of brackets are `'1234, 56789'` at indexes 1, 3.
# +
def abba(text): return any(a == d != b == c for (a, b, c, d) in subsequences(text, 4))
def subsequences(seq, n): return [seq[i:i+n] for i in range(len(seq) + 1 - n)]
def segment(line): return re.split(r'\[|\]', line)
def outsides(segments): return ', '.join(segments[0::2])
def insides(segments): return ', '.join(segments[1::2])
def tls(segments): return abba(outsides(segments)) and not abba(insides(segments))
sum(tls(segment(line)) for line in Input(7))
# -
# Here are some tests:
assert abba('abba') and not abba('aaaa') and not abba('abbc')
assert subsequences('abcdefg', 4) == ['abcd', 'bcde', 'cdef', 'defg']
assert segment('abcd[1234]fghi[56789]zz') == ['abcd', '1234', 'fghi', '56789', 'zz']
assert outsides(['abcd', '1234', 'fghi', '56789', 'zz']) == 'abcd, fghi, zz'
assert insides(['abcd', '1234', 'fghi', '56789', 'zz']) == '1234, 56789'
assert tls(['abba', '123'])
assert not tls(['bookkeeper', '123']) and not tls(['abba', 'xxyyx'])
# In **part two**, we are asked to count the number of *SSL* lines: an *SSL* is when there is an *ABA* outside brackets, and the corresponding *BAB* inside brackets. An *ABA* is a three-character sequence with first and third (but not all three) the same. The corresponding *BAB* has the first character of the *ABA* surrounded by two copies of the second character.
# +
alphabet = 'abcdefghijklmnopqrstuvwxyz'
def ssl(segments):
"Is there an ABA outside brackets, and the corresponding BAB inside?"
outs, ins = outsides(segments), insides(segments)
return any(a+b+a in outs and b+a+b in ins
for a in alphabet for b in alphabet if a != b)
sum(ssl(segment(line)) for line in Input(7))
# -
# # [Day 8](http://adventofcode.com/2016/day/8): Two-Factor Authentication
#
# Given an array of pixels on a screen, follow commands that can:
# - Turn on a sub-rectangle of pixels in the upper left corner: `rect 3x2`
# - Rotate a row of pixels: `rotate row y=0 by 4`
# - Rotate a column of pixels: `rotate column x=1 by 1`
#
# Then count the total number of `1` pixels in the screen.
#
# I will use `numpy` two-dimensional arrays, mostly because of the `screen[:, A]` notation for getting at a column.
# +
def interpret(cmd, screen):
"Interpret this command to mutate screen."
A, B = map(int, re.findall(r'(\d+)', cmd)) # There should be 2 numbers on every command line
if cmd.startswith('rect'):
screen[:B, :A] = 1
elif cmd.startswith('rotate row'):
screen[A, :] = rotate(screen[A, :], B)
elif cmd.startswith('rotate col'):
screen[:, A] = rotate(screen[:, A], B)
def rotate(items, n): return np.append(items[-n:], items[:-n])
def Screen(): return np.zeros((6, 50), dtype=np.int)
def run(commands, screen):
"Do all the commands and return the final pixel array."
for cmd in Input(8):
interpret(cmd, screen)
return screen
screen = run(Input(8), Screen())
np.sum(screen)
# -
# In **part two**, we are asked what message is on the screen. I won't try to do OCR; I'll just print the screen and look at the output:
for row in screen:
print(cat(' @'[pixel] for pixel in row))
# My answer is `EOARGPHYAO`.
# # [Day 9](http://adventofcode.com/2016/day/9): Explosives in Cyberspace
#
# In this puzzle we are asked to decompress text of the form `'A(2x5)BCD'`, where the `'(2x5)'` means to make 5 copies of the next 2 characters, yielding `'ABCBCBCBCBCD'`. We'll go through the input text, a character at a time, and if a `re` matcher detects a `'(CxR)'` pattern, process it; otherwise just collect the character. Note that the `C` characters that are to be repeated `R` times are taken literally; even if they contain an embedded `'(1x5)'`.
# +
matcher = re.compile(r'[(](\d+)x(\d+)[)]').match # e.g. matches "(2x5)" as ('2', '5')
def decompress(s):
"Decompress string s by interpreting '(2x5)' as making 5 copies of the next 2 characters."
s = re.sub(r'\s', '', s) # "whitespace is ignored"
result = []
i = 0
while i < len(s):
m = matcher(s, i)
if m:
i = m.end() # Advance to end of '(CxR)' match
C, R = map(int, m.groups())
result.append(s[i:i+C] * R) # Collect the C characters, repeated R times
i += C # Advance past the C characters
else:
result.append(s[i]) # Collect 1 regular character
i += 1 # Advance past it
return cat(result)
len(decompress(Input(9).read()))
# -
# In **part two**, the copied characters *are* recursively decompressed. So, given `'(8x2)(3x3)ABC'`, the `(8x2)` directive picks out the 8 characters `'(3x3)ABC'`, which would then be decompressed to get `'ABCABCABC'` and then the `'x2'` is applied to get `'ABCABCABCABCABCABC'`. However, for this part, we are not asked to actually build up the decompressed string, just to compute its length:
# +
def decompress_length(s):
"""Decompress string s by interpreting '(2x5)' as making 5 copies of the next 2 characters.
Recursively decompress these next 5 characters. Return the length of the decompressed string."""
s = re.sub(r'\s', '', s) # "whitespace is ignored"
length = 0
i = 0
while i < len(s):
m = matcher(s, i)
if m:
C, R = map(int, m.groups())
i = m.end(0) # Advance to end of '(CxR)'
length += R * decompress_length(s[i:i+C]) # Decompress C chars and add to length
i += C # Advance past the C characters
else:
length += 1 # Add 1 regular character to length
i += 1 # Advance past it
return length
decompress_length(Input(9).read())
# -
# Here are some tests:
# +
assert decompress('A(2x5)BCD') == 'ABCBCBCBCBCD'
assert decompress('ADVENT') == 'ADVENT'
assert decompress('(3x3)XYZ') == 'XYZXYZXYZ'
assert decompress('(5x4)(3x2)') == '(3x2)(3x2)(3x2)(3x2)'
assert decompress('X(8x2)(3x3)ABCY') == 'X(3x3)ABC(3x3)ABCY'
assert decompress_length('(8x2)(3x3)ABC') == 18
assert decompress_length('(25x3)(3x3)ABC(2x3)XY(5x2)PQRSTX(18x9)(3x2)TWO(5x7)SEVEN') == 445
assert decompress_length('(9x999)(2x999)xx') == 999 * 999 * 2 == 1996002
# -
# # [Day 10](http://adventofcode.com/2016/day/10): Balance Bots
#
# In this puzzle, a fleet of robots exchange some chips from input bins,
# passing them among themselves, and eventually putting them in output bins. We are given instructions like this:
#
# value 5 goes to bot 2
# bot 2 gives low to bot 1 and high to bot 0
# value 3 goes to bot 1
# bot 1 gives low to output 1 and high to bot 0
# bot 0 gives low to output 2 and high to output 0
# value 2 goes to bot 2
#
# At first I thought I just had to interpret these instructions sequentially, but then I realized this is actually a *data flow* problem: *whenever* a bot acquires two chips, it passes the low number chip to one destination and the high number to another. So my representation choices are:
# - Bots and bins are represented as strings: `'bot 1'` and `'output 2'`.
# - Chips are represented by ints. (Not strings, because we want 9 to be less than 10).
# - Keep track of which bot currently has which chip(s) with a dict: `has['bot 2'] = {5}`
# - Keep track of what a bot does when it gets 2 chips with a dict: `gives['bot 1'] = ('output 1', 'bot 0')`
# - Pull this information from instructions with `re.findall`. The order of instructions is not important.
# - A function, `give`, moves a chip to a recipient, and if the recipient now has two chips, that triggers two more `give` calls.
# +
def bots(instructions, goal={17, 61}):
"Follow the data flow instructions, and if a bot gets the goal, print it."
def give(giver, chip, recip):
"Pass the chip from giver to recipient."
has[giver].discard(chip)
has[recip].add(chip)
chips = has[recip]
if chips == goal:
print(recip, 'has', goal)
if len(chips) == 2:
give(recip, min(chips), gives[recip][0])
give(recip, max(chips), gives[recip][1])
has = defaultdict(set) # who has what
gives = {giver: (dest1, dest2) # who will give what
for (giver, dest1, dest2)
in re.findall(r'(bot \d+) gives low to (\w+ \d+) and high to (\w+ \d+)', instructions)}
for (chip, recip) in re.findall(r'value (\d+) goes to (\w+ \d+)', instructions):
give('input bin', int(chip), recip)
return has
has = bots(Input(10).read())
# -
# In **part two**, we are asked for the product of three output bins:
# +
def out(i): return has['output ' + str(i)].pop()
out(0) * out(1) * out(2)
# -
# # [Day 11](http://adventofcode.com/2016/day/11): Radioisotope Thermoelectric Generators
#
# I *knew* my `astar_search` function would come in handy! To search for the shortest path to the goal, I need to provide an initial state of the world, a heuristic function (which estimates how many moves away from the goal a state is), and a function that says what states can be reached by moving the elevator up or down, and carrying some stuff. I will make these choices:
# * The state of the world is represented by the `State` type, which says what floor the elevator is on, and for a tuple of floors, the set of objects that are on the floor. We use frozensets so that a state will be hashable.
# * To figure out what moves can be made, consider both directions for the elevator (up or down); find all combinations of one or two items on each floor, and keep all of those moves, as long as they don't violate the constraint that we can't have a chip on the same floor as an RTG, unless the chip's own RTG is there.
# * To calculate the heuristic, add up the number of floors away each item is from the top floor and divide by two (since a move might carry two items).
#
# Here is my input:
#
# * The first floor contains a thulium generator, a thulium-compatible microchip, a plutonium generator, and a strontium generator.
# * The second floor contains a plutonium-compatible microchip and a strontium-compatible microchip.
# * The third floor contains a promethium generator, a promethium-compatible microchip, a ruthenium generator, and a ruthenium-compatible microchip.
# * The fourth floor contains nothing relevant.
# +
State = namedtuple('State', 'elevator, floors')
def fs(*items): return frozenset(items)
legal_floors = {0, 1, 2, 3}
def combos(things):
"All subsets of 1 or 2 things."
for s in chain(combinations(things, 1), combinations(things, 2)):
yield fs(*s)
def moves(state):
"All legal states that can be reached in one move from this state"
L, floors = state
for L2 in {L + 1, L - 1} & legal_floors:
for stuff in combos(floors[L]):
newfloors = tuple((s | stuff if i == L2 else
s - stuff if i == state.elevator else
s)
for (i, s) in enumerate(state.floors))
if legal_floor(newfloors[L]) and legal_floor(newfloors[L2]):
yield State(L2, newfloors)
def legal_floor(floor):
"Floor is legal if no RTG, or every chip has its corresponding RTG."
rtgs = any(r.endswith('G') for r in floor)
chips = [c for c in floor if c.endswith('M')]
return not rtgs or all(generator_for(c) in floor for c in chips)
def generator_for(chip): return chip[0] + 'G'
def h_to_top(state):
"An estimate of the number of moves needed to move everything to top."
total = sum(len(floor) * i for (i, floor) in enumerate(reversed(state.floors)))
return math.ceil(total / 2) # Can move two items in one move.
# -
# Let's try it out on an easy sample problem:
# +
easy = State(0, (fs('RG'), Ø, fs('RM'), Ø))
astar_search(easy, h_to_top, moves)
# -
# Now to solve the real problem. The answer we need is the number of elevator moves, which is one less than the path length (because the path includes the initial state).
# +
part1 = State(0, (fs('TG', 'TM', 'PG', 'SG'), fs('PM', 'SM'), fs('pM', 'pG', 'RM', 'RG'), Ø))
# %time path = astar_search(part1, h_to_top, moves)
len(path) - 1
# -
# In **part two**, we add four more items. Now, in part one there were 10 items, each of which could be on any of the 4 floors, so that's 4<sup>10</sup> ≈ 1 million states. Adding 4 more items yields ≈ 260 million states. We won't visit every state, but the run time could be around 100 times longer. I think I'll start this running, take the dog for a walk, and come back to see if it worked.
# +
part2 = State(0, (fs('TG', 'TM', 'PG', 'SG', 'EG', 'EM', 'DG', 'DM'),
fs('PM', 'SM'), fs('pM', 'pG', 'RM', 'RG'), Ø))
# %time path = astar_search(part2, h_to_top, moves)
len(path) - 1
# -
# It worked. And it took about 60 times longer. If I wanted to make it more efficient, I would focus on *symmetry*: when there are two symmetric moves, we only need to consider one. For example, in terms of finding the shortest path, it is the same to move, say `{'TG', 'TM'}` or `{'EG', 'EM'}` or `{'DG', 'DM'}` when they are all on the ground floor.
# # [Day 12](http://adventofcode.com/2016/day/12): Leonardo's Monorail
#
# This one looks pretty easy: an interpreter for an assembly language with 4 op codes and 4 registers. We start by parsing a line like `"cpy 1 a"` into a tuple, `('cpy', 1, 'a')`. Then to `interpret` the code, we set the program counter, `pc`, to 0, and interpret the instruction at `code[0]`, and continue until the `pc` is past the end of the code:
# +
def interpret(code, regs):
"Execute instructions until pc goes off the end."
def val(x): return (regs[x] if x in regs else x)
pc = 0
while pc < len(code):
inst = code[pc]
op, x, y = inst[0], inst[1], inst[-1]
pc += 1
if op == 'cpy': regs[y] = val(x)
elif op == 'inc': regs[x] += 1
elif op == 'dec': regs[x] -= 1
elif op == 'jnz' and val(x): pc += y - 1
return regs
def parse(line):
"Split line into words, and convert to int where appropriate."
return tuple((x if x.isalpha() else int(x))
for x in line.split())
code = [parse(line) for line in Input(12)]
interpret(code, dict(a=0, b=0, c=0, d=0))
# -
# I had a bug initially: in the `jnz` instruction, I had `pc += y`, to do the relative jump, but I forgot the `-1` to offset the previous `pc += 1`.
#
# In **part two** all we have to do is initialize register `c` to 1, not 0:
interpret(code, dict(a=0, b=0, c=1, d=0))
# # [Day 13](http://adventofcode.com/2016/day/13): A Maze of Twisty Little Cubicles
#
# This is a maze-solving puzzle, where the maze is infinite in the non-negative (x, y) quarter-plane. Each space in that infinite grid is open or closed according to this computation:
# > Find `x*x + 3*x + 2*x*y + y + y*y`.
# Add the office designer's favorite number (your puzzle input).
# Find the binary representation of that sum; count the number of bits that are 1.
# If the number of bits that are 1 is even, it's an open space (denoted `'.'`).
# If the number of bits that are 1 is odd, it's a wall (denoted `'#'`).
#
# The problem is to find the length of the shortest path to the goal location, (31, 39). So I'll be using `astar_search` again.
# +
favorite = 1362
goal = (31, 39)
def is_open(location):
"Is this an open location?"
x, y = location
num = x*x + 3*x + 2*x*y + y + y*y + favorite
return x >= 0 and y >= 0 and bin(num).count('1') % 2 == 0
def open_neighbors(location): return filter(is_open, neighbors4(location))
path = astar_search((1, 1), lambda p: cityblock_distance(p, goal), open_neighbors)
len(path) - 1
# -
# Here we see a portion of the maze:
for y in range(30):
print(cat(('.' if is_open((x, y)) else '#') for x in range(90)))
# In **part two**, we're asked how many locations we can reach in 50 moves or less. I'll grab the `breadth_first` search function from [aima-python](https://github.com/aimacode/aima-python/blob/master/search-4e.ipynb) and modify it to find all the states within N steps:
# +
def count_locations_within(start, N, neighbors):
"Find how many locations are within N steps from start."
frontier = deque([start]) # A queue of states
distance = {start: 0} # distance to start; also tracks all states seen
while frontier:
s = frontier.popleft()
if distance[s] < N:
for s2 in neighbors(s):
if s2 not in distance:
frontier.append(s2)
distance[s2] = distance[s] + 1
return len(distance)
count_locations_within((1, 1), 50, open_neighbors)
# -
# # [Day 14](http://adventofcode.com/2016/day/14): One-Time Pad
#
# For this problem I again have to take the md5 hash of a string with increasing integers appended. The puzzle is to find the integer that yields the 64th key, where a hash is a key if:
# - It contains three of the same character in a row, like 777. Only consider the first such triplet in a hash.
# - One of the next 1000 hashes in the stream contains that same character five times in a row, like 77777.
#
# I'll use `lru_cache` to avoid repeating the hashing of the next 1000.
# +
salt = '<PASSWORD>'
@lru_cache(1001)
def hashval(i): return hashlib.md5(bytes(salt + str(i), 'utf-8')).hexdigest()
def is_key(i):
"A key has a triple like '777', and then '77777' in one of the next thousand hashval(i)."
three = re.search(r'(.)\1\1', hashval(i))
if three:
five = three.group(1) * 5
return any(five in hashval(i+delta) for delta in range(1, 1001))
def nth_key(N): return nth(filter(is_key, range(BIG)), N)
nth_key(63)
# -
# In **part two**, we do *key stretching*, hashing an additional 2016 times. Eeverything else is the same:
# +
@lru_cache(1001)
def hashval(i, stretch=2016):
h = hashlib.md5(bytes(salt + str(i), 'utf-8')).hexdigest()
for i in range(stretch):
h = hashlib.md5(bytes(h, 'utf-8')).hexdigest()
return h
# %time nth_key(63)
# -
# This was my highest-scoring day, finishing #20 on part two.
# # [Day 15](http://adventofcode.com/2016/day/15): Timing is Everything
#
# In this puzzle rotating discs with a slot in position 0 spin around. We are asked at what time will all the slots be lined up for a capsule to fall through the slots. The capsule takes one time unit to fall through each disc (not clear why it doesn't accelerate as it falls) and the discs spin one position per time unit.
# +
def parse(inputs):
"Parse an input string into (disc#, positions, pos) triples."
return [tuple(map(int, triple))
for triple in re.findall(r'#(\d+).* (\d+) positions.* (\d+)[.]', inputs)]
discs = parse('''
Disc #1 has 13 positions; at time=0, it is at position 1.
Disc #2 has 19 positions; at time=0, it is at position 10.
Disc #3 has 3 positions; at time=0, it is at position 2.
Disc #4 has 7 positions; at time=0, it is at position 1.
Disc #5 has 5 positions; at time=0, it is at position 3.
Disc #6 has 17 positions; at time=0, it is at position 5.
''')
def falls(t, discs):
"If we drop the capsule at time t, does it fall through all slots?"
return all((pos + t + d) % positions == 0
for (d, positions, pos) in discs)
first(t for t in range(BIG) if falls(t, discs))
# -
assert discs == [(1, 13, 1), (2, 19, 10), (3, 3, 2), (4, 7, 1), (5, 5, 3), (6, 17, 5)]
assert falls(5, [(1, 5, 4), (2, 2, 1)])
# For **part two**, we add a 7th disc, with 11 positions, at position 0 at time 0. I coud go through all the possible times, just as in part one, but I see a way to get a 19-fold speedup: Disc #2 is the largest, with 19 positions, so we only need consider every 19 values of `t`. But what is the first one to consider? Disc @2 starts at position 10, so to get back to position 0, we have to release at `t=7`, because 10 + 2 + 7 = 19 = 0 mod 19. So we will iterate `t` over `range(7, BIG, 19)`:
# +
discs.append((7, 11, 0))
first(t for t in range(7, BIG, 19) if falls(t, discs))
# -
# # [Day 16](http://adventofcode.com/2016/day/16) Dragon Checksum
#
# Given a bit string of the form `'01...'`, expand it until it fills N bits, then report the checksum.
#
# The rules for expanding:
# - Call the data you have at this point "a".
# - Make a copy of "a"; call this copy "b".
# - Reverse the order of the characters in "b".
# - In "b", replace all instances of 0 with 1 and all 1s with 0.
# - The resulting data is "a", then a single 0, then "b".
# - If this gives N or more bits, take the first N; otherwise repeat the process.
#
# The rules for the checksum:
# - Assume the string is 110010110100
# - Consider each pair: 11, 00, 10, 11, 01, 00.
# - These are same, same, different, same, different, same, producing 110101.
# - The resulting string has length 6, which is even, so we repeat the process.
# - The pairs are 11 (same), 01 (different), 01 (different).
# - This produces the checksum 100, which has an odd length, so we stop.
# +
def expand(a, N):
"Expand seed `a` until it has length N."
while len(a) < N:
b = flip(a[::-1])
a = a + '0' + b
return a[:N]
def flip(text, table=str.maketrans('10', '01')): return text.translate(table)
def checksum(a):
"Compute the checksum of `a` by comparing pairs until len is odd."
while len(a) % 2 == 0:
a = cat(('1' if a[i] == a[i+1] else '0')
for i in range(0, len(a), 2))
return a
seed = '10010000000110000'
checksum(expand(seed, 272))
# -
# In **part two**, we take the same seed, but expand it to fill 35Mb of space:
# %time checksum(expand(seed, 35651584))
# # [Day 17](http://adventofcode.com/2016/day/17) Two Steps Forward
#
# In this puzzle, we move through a 4x4 grid/maze, starting at position (0, 0) and trying to reach (3, 3), but the door from one position to the next is open or not depending on the hash of the path to get there (and my passcode), so doors open and lock themselves as you move around. I'll represent a state as a tuple of `(position, path)` and use `astar_search` to find the shortest path to the goal:
#
# +
passcode = '<PASSWORD>'
openchars = 'bcdef'
grid = set((x, y) for x in range(4) for y in range(4))
start, goal = (0, 0), (3, 3)
def to_goal(state):
"City block distance between state's position and goal."
pos, path = state
return cityblock_distance(pos, goal)
directions = [(0, 'U', (0, -1)), (1, 'D', (0, 1)), (2, 'L', (-1, 0)), (3, 'R', (1, 0))]
def moves(state):
"All states reachable from this state."
(x, y), path = state
hashx = hashlib.md5(bytes(passcode + path, 'utf-8')).hexdigest()
for (i, p, (dx, dy)) in directions:
pos2 = (x+dx, y+dy)
if hashx[i] in openchars and pos2 in grid:
yield (pos2, path+p)
astar_search((start, ''), to_goal, moves)
# -
# In **part two**, we're asked for the longest path to the goal. We have to stop when we reach the goal, but we can make repeated visits to positions along the way, as long as the doors are open. I'll use a depth-first search, and keep track of the longest path length:
# +
def longest_search(state, goal, moves):
"Find the longest path to goal by depth-first search."
longest = 0
frontier = [state]
while frontier:
state = (pos, path) = frontier.pop()
if pos == goal:
longest = max(longest, len(path))
else:
frontier.extend(moves(state))
return longest
longest_search((start, ''), goal, moves)
# -
# # [Day 18](http://adventofcode.com/2016/day/18) Like a Rogue
#
# Here we have a cellular automaton, where a cell is a "trap" iff the 3 tiles in the row above, (one to the left above, directly above, and one to the right above) are one of the set `{'^^.', '.^^', '^..', '..^'}`; in other words if the first of the three is different from the last of the three. Given an initial row, we're asked for the count of all the safe tiles in the first 40 rows:
#
# +
safe, trap = '.', '^'
initial = '.^^.^^^..^.^..^.^^.^^^^.^^.^^...^..^...^^^..^^...^..^^^^^^..^.^^^..^.^^^^.^^^.^...^^^.^^.^^^.^.^^.^.'
def rows(n, row=initial):
"The first n rows of tiles (given the initial row)."
result = [row]
for i in range(n-1):
previous = safe + result[-1] + safe
result.append(cat((trap if previous[i-1] != previous[i+1] else safe)
for i in range(1, len(previous) - 1)))
return result
cat(rows(40)).count(safe)
# -
# Here I reproduce the simple example from the puzzle page:
rows(10, '.^^.^.^^^^')
# In **part two**, we just have to run longer (but only a few seconds):
# %time cat(rows(400000)).count(safe)
# # [Day 19](http://adventofcode.com/2016/day/19) An Elephant Named Joseph
#
# Elves numbered 1 to *N* sit in a circle. Each Elf brings a present. Then, starting with the first Elf, they take turns stealing all the presents from the Elf to their left. An Elf with no presents is removed from the circle and does not take turns. So, if *N* = 5, then:
#
# Elf 1 takes Elf 2's present.
# Elf 2 has no presents and is skipped.
# Elf 3 takes Elf 4's present.
# Elf 4 has no presents and is also skipped.
# Elf 5 takes Elf 1's two presents.
# Neither Elf 1 nor Elf 2 have any presents, so both are skipped.
# Elf 3 takes Elf 5's three presents, ending the game.
#
# Who ends up with all the presents for general case of *N*?
# First, I note that I only need to keep track of the Elf number of the remaining elves,
# I don't need to count how many presents each one has. I see two representation choices:
# - Represent the circle of elves as a list of elf numbers, and everytime an Elf's presents are taken, delete the elf from the list. But this is O(*N*<sup>2</sup>), where *N* = 3 million, so this will be slow.
# - Represent the elves by a range, and instead of deleting elf-by-elf, instead limit the range round-by-round.
# If there is an even number of elves, then the elf in position 0 takes from position 1; position 2 takes from position 3, and so on, leaving only the even positions, which we denote `elves[0::2]`. If there is an odd number of elves, then it is the same, except that the last elf takes from the one in position 0, leaving `elves[2::2]`. Here's the code:
# +
def Elves(N=3018458): return range(1, N+1)
def winner(elves): return (elves[0] if (len(elves) == 1) else winner(one_round(elves)))
def one_round(elves): return (elves[0::2] if (len(elves) % 2 == 0) else elves[2::2])
assert winner(Elves(5)) == 3
winner(Elves())
# -
# Here is a cool thing about representing the elves with a range: the total storage is O(1), not O(*N*).
# We never need to make a list of 3 million elements.
# Here we see a trace of the calls to `one_round`:
#
one_round = trace1(one_round)
winner(Elves())
# In **part two** the rules have changed, and each elf now takes from the elf *across* the circle. If there is an even number of elves, take from the elf directly across. Fo example, with 12 elves in a circle (like a clock face), Elf 1 takes from Elf 7. With an odd number of elves, directly across the circle falls between two elves, so choose the one that is earlier in the circle. For example, with 11 elves, Elf 2 takes from the Elf at position 7. Now who ends up with the presents?
#
# This is tougher. I can't think of a simple `range` expression to describe who gets eliminated in a round. But I can represent the circle as a list and write a loop to eliminate elves one at a time. Again, if I did that with a `del` statement for each elf, it would be O(*N*<sup>2</sup>). But if instead I do one round at a time, replacing each eliminated elf with `None` in the list, and then filtering out the `None` values, then each round is only O(*N*), and sine there will be log(*N*) rounds, the whole thing is only O(*N* log(*N*)). That should be reasonably fast.
#
# It is still tricky to know which elf to eliminate. If there are *N* elves, then the elf at position *i* should elminate the one at position *i* + *N* // 2. But we have to skip over the already-eliminated spaces; we can do that by keeping track of the number of eliminated elves in the variable `eliminated`. We also need to keep track of the current value of `N`, since it will change. And, since I don't want to deal with the headaches of wrapping around the circletconn, I will only deal with the first third of the elves: the first third all eliminate elves in the other two-thirds; if we went more than 1/3 of the way through, we would have to worry about wrapping around. (I had a bug here: at first I just iterated through `N // 3`. But when `N` is 2, that does no iteration at all, which is wrong; with two elves, the first should eliminate the other. It turns out it is safe to iterate through `ceil(N /3)` on each round.)
#
# I will change the `Elves` function to return a `list`, not a `range`. The function `winner` stays the same. The `one_round` function is where the work goes:
# +
def Elves(N=3018458): return list(range(1, N+1))
def one_round(elves):
"The first third of elves eliminate ones across the circle from them; who is left?"
N = len(elves)
eliminated = 0
for i in range(int(math.ceil(N / 3))):
across = i + eliminated + (N // 2)
elves[across] = None
N -= 1
eliminated += 1
return list(filter(None, elves[i+1:] + elves[:i+1]))
assert winner(Elves(5)) == 2
assert one_round(Elves(5)) == [4, 1, 2]
# %time winner(Elves())
# -
# I was worried that this solution might take over a minute to run, but it turns out to only take about a second.
# # [Day 20](http://adventofcode.com/2016/day/20) Firewall Rules
#
# We are given a list of blocked IP addresses, in the form `"2365712272-2390766206"`, indicating the low and high numbers that are blocked by the firewall. I will parse the numbers into `(low, high)` pairs, and sort them by the low number first (and peek at the first 5 to see if I got it right):
# +
pairs = sorted(map(parse_ints, Input(20)))
pairs[:5]
# -
# We are asked what is the lowest non-negative integer that is not blocked. I will generate all the unblocked numbers, and just ask for the first one. (Why do it that way? Because it feels like `unblocked` is the fundamental issue of the problem, and we already have a function to compute `first`; there's no need for a `first_unblocked` function that conflates two ideas.) To find unblocked numbers, start a counter, `i` at zero, and increment it past the high value of each range, after yielding any numbers from `i` to the low value of the range:
# +
def unblocked(pairs):
"Find the lowest unblocked integer, given the sorted pairs of blocked numbers."
i = 0
for (low, high) in pairs:
yield from range(i, low)
i = max(i, high + 1)
first(unblocked(pairs))
# -
# In **part two** we are asked how many numbers are unblocked:
len(list(unblocked(pairs)))
# # [Day 21](http://adventofcode.com/2016/day/21) Scrambled Letters and Hash
#
# In this puzzle we are asked to take a password string, scramble it according to a list of instructions, and output the result, which will be a permutation of the original password. This is tedious because there are seven different instructions, but each one is pretty straightforward. I make the following choices:
# - I'll transform `password` (a `str`) into `pw` (a `list`), because lists are mutable and easier to manipulate. At the end I'll turn it back into a `str`.
# - I'll define functions `rot` and `swap` because they get used multiple times by different instructions.
# - I use the variables `A, B` to denote the first two integers anywhere in a line. If there is only one integer (or none), then `B` (and `A`) get as a default value `0`. I accept ill-formed instructions, such as `"move 1 to 4"` instead of requiring `"move position 1 to position 4"`.
#
# +
def scramble(password, instructions=list(Input(21)), verbose=False):
"Scramble the password according to the instructions."
pw = list(password)
def rot(N): pw[:] = pw[-N:] + pw[:-N]
def swap(A, B): pw[A], pw[B] = pw[B], pw[A]
for line in instructions:
words = line.split()
A, B, = parse_ints(line + ' 0 0')[:2]
cmd = line.startswith
if cmd('swap position'): swap(A, B)
elif cmd('swap letter'): swap(pw.index(words[2]), pw.index(words[5]))
elif cmd('rotate right'): rot(A)
elif cmd('rotate left'): rot(-A)
elif cmd('reverse'): pw[A:B+1] = pw[A:B+1][::-1]
elif cmd('move'): pw[A:A+1], pw[B:B] = [], pw[A:A+1]
elif cmd('rotate based'):
i = pw.index(words[6])
rot((i + 1 + (i >= 4)) % len(pw))
if verbose:
print(line + ': ' + cat(pw))
return cat(pw)
scramble('abcdefgh')
# -
# When I ran the first version of this code the answer I got was incorrect, and I couldn't see where I went wrong, so I implemented the test case from the problem description and inspected the results line by line.
# +
test = '''swap position 4 with position 0
swap letter d with letter b
reverse positions 0 through 4
rotate left 1 step
move position 1 to position 4
move position 3 to position 0
rotate based on position of letter b
rotate based on position of letter d'''.splitlines()
scramble('abcde', test, verbose=True)
# -
# That was enough to show me that I had two bugs (which are fixed above):
# - For `"reverse"`, I thought `"positions 0 through 4"` meant `[0:4]`, when actually it means `[0:5]`.
# - For `"rotate based"`, in the case where the rotation is longer than the password, I need to take the modulo of the password length.
#
# For **part two**, the task is to find the password that, when scrambled, yields `'fbgdceah'`. I think the puzzle designer was trying to tempt solvers into implementing an `unscramble` function, which would be another 20 or 30 lines of code. Fortunately, I was too lazy to go down that path. I realized there are only 40 thousand permutations of an 8-character password, so we can just brute force them all (which would be infeasible with a 20-character password):
{cat(p) for p in permutations('fbgdceah')
if scramble(p) == 'fbgdceah'}
# # [Day 22](http://adventofcode.com/2016/day/22) Grid Computing
#
# We are given a description of files across a grid computing cluster, like this:
#
# root@ebhq-gridcenter# df -h
# Filesystem Size Used Avail Use%
# /dev/grid/node-x0-y0 92T 70T 22T 76%
# /dev/grid/node-x0-y1 86T 65T 21T 75%
#
# For part one, we are asked how many pairs of nodes can viably make a transfer of data. The pair (A, B) is viable if
# - Node A is not empty (its Used is not zero).
# - Nodes A and B are not the same node.
# - The data on node A (its Used) would fit on node B (its Avail).
#
# I'll represent a node as a `namedtuple` of six integers; the rest is easy:
# +
Node = namedtuple('Node', 'x, y, size, used, avail, pct')
nodes = [Node(*parse_ints(line)) for line in Input(22) if line.startswith('/dev')]
def viable(A, B): return A != B and 0 < A.used <= B.avail
sum(viable(A, B) for A in nodes for B in nodes)
# -
# That worked, but let's make sure the nodes look reasonable:
nodes[:5]
# In **part two**, we are asked to move data from the node in the upper right (the one with maximum `x` value and `y=0`) to the upper left (x=0, y=0). At first I worried about all sorts of complications: could we split the data into two or more pieces, copying different pieces into different nodes, and then recombining them? I spent many minutes thinking about these complications. Eventually, after a more careful reading of the rules, I decided such moves were not allowed, and the answer had to just involve moving the empty square around. So to proceed, we need to find the initial position of the empty node, and the maximum x value, so we know where the data is:
# +
empty = first(node for node in nodes if node.used == 0)
maxx = max(node.x for node in nodes)
empty, maxx
# -
# I will also define the `grid` as a dict of `{(x, y): node}` entries (which will enable me to find neighbors of a node):
grid = {(node.x, node.y): node
for node in nodes}
# An `astar_search` seems appropriate. Each state of the search keeps track of the position of the data we are trying to get, and the position of the currently empty node. The heuristic is the city block distance of the data to the origin:
# +
State = namedtuple('State', 'datapos, emptypos')
def distance(state): return cityblock_distance(state.datapos)
def moves(state):
"Try moving any neighbor we can into the empty position."
for pos in neighbors4(state.emptypos):
if pos in grid:
# Try to move contents of `node` at pos into `empty` at emptypos
node, empty = grid[pos], grid[state.emptypos]
if node.used <= empty.size:
newdatapos = (state.emptypos if pos == state.datapos else state.datapos)
yield State(newdatapos, pos)
path = astar_search(State((maxx, 0), (empty.x, empty.y)), distance, moves)
len(path) - 1
# -
# # [Day 23](http://adventofcode.com/2016/day/23) Safe Cracking
#
# This day's puzzle is just like Day 12, except there is one more instruction, `tgl`. I made four mistakes in the process of coding this up:
# - At first I didn't read the part that says register `'a'` should initially be 7.
# - I wasn't sure exactly what constitutes an invalid instruction; it took me a few tries to get that right:
# the only thing that is invalid is a `cpy` that does not copy into a register.
# - I forgot to subtract one from the `pc` (again!) in the `tgl` instruction.
# - I forgot that I had `parse` return instructions as immutable tuples; I had to change that to mutable lists.
# +
def interpret(code, regs):
"Execute instructions until pc goes off the end."
def val(x): return (regs[x] if x in regs else x)
pc = 0
while 0 <= pc < len(code):
inst = code[pc]
op, x, y = inst[0], inst[1], inst[-1]
pc += 1
if op == 'cpy' and y in regs: regs[y] = val(x)
elif op == 'inc': regs[x] += 1
elif op == 'dec': regs[x] -= 1
elif op == 'jnz' and val(x): pc += val(y) - 1
elif op == 'tgl': toggle(code, pc - 1 + val(x))
return regs
def toggle(code, i):
"Toggle the instruction at location i."
if 0 <= i < len(code):
inst = code[i]
inst[0] = ('dec' if inst[0] == 'inc' else
'inc' if len(inst) == 2 else
'cpy' if inst[0] == 'jnz' else
'jnz')
def parse(line):
"Split line into words, and convert to int where appropriate."
return [(x if x.isalpha() else int(x))
for x in line.split()]
# +
text = '''
cpy a b
dec b
cpy a d
cpy 0 a
cpy b c
inc a
dec c
jnz c -2
dec d
jnz d -5
dec b
cpy b c
cpy c d
dec d
inc c
jnz d -2
tgl c
cpy -16 c
jnz 1 c
cpy 84 c
jnz 75 d
inc a
inc d
jnz d -2
inc c
jnz c -5
'''.strip()
code = [parse(line) for line in text.splitlines()]
regs = dict(a=7, b=0, c=0, d=0)
interpret(code, regs)
# -
# In **part two**, we are told to run the same computation, but with register `a` set to 12. We are also warned that this will take a long time, and we might consider implementing a multiply instruction, but I was too lazy to make sense of the assembly code, and just let my interpreter run to completion, even if it takes a while.
# +
code = [parse(line) for line in text.splitlines()]
regs = dict(a=12, b=0, c=0, d=0)
# %time interpret(code, regs)
# -
# Well, it completed, and gave me the right answer. But I feel like the intent of *Advent of Code* is like *Project Euler*: all code should run in about a minute or less. So I don't think this counts as a "real" solution.
# # [Day 24](http://adventofcode.com/2016/day/24) Air Duct Spelunking
#
# This is another maze-solving problem; it should be easy for my `astar_search`. First the maze:
maze = tuple(Input(24))
# The tricky part is that we have to visit all the digits in the maze, starting at `0`, and not necessarily going in order. How many digits are there?
set(cat(maze))
# OK, there are 8 digits. What is the start square (the square that currently holds a `'0'`)?
zero = first((x, y) for y, row in enumerate(maze) for x, c in enumerate(row) if c == '0')
zero
# Now I'm ready to go. The state of the search will include the x, y position, and also the digits visited so far, which I can represent as a sorted string (a `frozenset` would also work):
# +
def h(state):
"Heuristic: the number of digits not yet visited."
_, visited = state
return 8 - len(visited) # Note: 8 == len('01234567')
def moves(state):
"Move to any neighboring square that is not a wall. Track the digits visited."
pos, visited = state
for x1, y1 in neighbors4(pos):
c = maze[y1][x1]
if c != '#':
visited1 = (visited if c in visited or c == '.' else cat(sorted(visited + c)))
yield (x1, y1), visited1
path = astar_search((zero, '0'), h, moves)
len(path) - 1
# -
# In **part two** we need to get the robot back to the start square. I'll do that by creating a new heuristic function that still requires us to collect all the digits, and also measures the distance back to the start (`zero`) square.
# +
def h2(state):
"Heuristic: the number of digits not yet visited, plus the distance back to start."
pos, visited = state
return 8 - len(visited) + cityblock_distance(pos, zero)
path2 = astar_search((zero, '0'), h2, moves)
len(path2) - 1
# -
# # [Day 25](http://adventofcode.com/2016/day/25) Clock Signal
#
# This is another assembly language interpreter puzzle. This time there is one more instruction, `out`, which transmits a signal. We are asked to find the lowest positive integer value for register `a` that causes the program to output an infinite series of `0, 1, 0, 1, 0, 1, ...` signals. Dealing with infinity is difficult, so I'll approximate that by asking: what is the lowest value for register `a` that causes the program to output at least 100 elements in the `0, 1, 0, 1, 0, 1, ...` series, within the first million instructions executed?
#
# To do that, I'll change `interpret` to be a generator that yields signals, and change it to take an argument saying the number of steps to execute before halting:
def interpret(code, regs, steps=BIG):
"Execute instructions until pc goes off the end, or until we execute the given number of steps."
def val(x): return (regs[x] if x in regs else x)
pc = 0
for _ in range(steps):
if not (0 <= pc < len(code)):
return
inst = code[pc]
op, x, y = inst[0], inst[1], inst[-1]
pc += 1
if op == 'cpy' and y in regs: regs[y] = val(x)
elif op == 'inc': regs[x] += 1
elif op == 'dec': regs[x] -= 1
elif op == 'jnz' and val(x): pc += val(y) - 1
elif op == 'tgl': toggle(code, pc - 1 + val(x))
elif op == 'out': yield val(x)
# Here is my program, and the function `repeats`, which returns True if the code repeats with a given value of the register `a`. Then all we need to do is iterate through integer values for register `a` until we find one that repeats:
# +
text = '''
cpy a d
cpy 4 c
cpy 633 b
inc d
dec b
jnz b -2
dec c
jnz c -5
cpy d a
jnz 0 0
cpy a b
cpy 0 a
cpy 2 c
jnz b 2
jnz 1 6
dec b
dec c
jnz c -4
inc a
jnz 1 -7
cpy 2 b
jnz c 2
jnz 1 4
dec b
dec c
jnz 1 -4
jnz 0 0
out b
jnz a -19
jnz 1 -21
'''.strip()
code = [parse(line) for line in text.splitlines()]
def repeats(a, code, steps=10**6, minsignals=100):
"Does this value for register a cause code to repeat `out` signals of 0, 1, 0, 1, ...?"
signals = interpret(code, dict(a=a, b=0, c=0, d=0), steps)
expecteds = cycle((0, 1))
for (i, (signal, expected)) in enumerate(zip(signals, expecteds)):
if signal != expected:
return False
# We'll say "yes" if the code outputs at least a minimum number of 0, 1, ... signals, and nothing else.
return i >= minsignals
first(a for a in range(1, BIG) if repeats(a, code))
# -
# That's all folks! Thank you [<NAME>](http://was.tl/), that was fun!
| ipynb/Advent of Code.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Chart analysis
#
# In this tutorial, you will learn about:
#
# * What are the different types of charts for plotting stock data
# * What are their purposes
#
# <hr size="5"/>
# Requirements:
# * [pandas](https://pypi.org/project/pandas/)
# * [matplotlib](https://matplotlib.org/)
# * [numpy](https://numpy.org/)
# +
# import libraries
import pandas as pd
import numpy as np
import datetime
import matplotlib
import matplotlib.pyplot as plt
import matplotlib.mlab as mlab
import mpl_finance
from mpl_finance import volume_overlay3
from matplotlib.dates import num2date
from matplotlib.dates import date2num
# -
# ### 1. Line chart
# +
# load aapl csv file
df = pd.read_csv('../../database/nasdaq_ticks_day/nasdaq_AAPL.csv', header=0, index_col='Date', parse_dates=True)
# We select only the data from Jan to Dec 2019
df = df.loc[pd.Timestamp('2019-01-01'):pd.Timestamp('2019-12-31')]
df.head()
# +
plt.style.use('ggplot')
# Initialise the plot figure
fig = plt.figure()
fig.set_size_inches(18.5, 10.5)
ax1 = plt.subplot2grid((6,1), (0,0), rowspan=5, colspan=1)
ax2 = plt.subplot2grid((6,1), (5,0), rowspan=1, colspan=1, sharex=ax1)
df['50ma'] = df['Close'].rolling(window=50, min_periods=0).mean()
df.dropna(inplace=True)
ax1.plot(df.index, df['Close'])
ax1.plot(df.index, df['50ma'])
ax2.bar(df.index, df['Volume'])
fig.savefig('./figures/00-line-chart.png', dpi=200)
plt.show()
# -
# ### 2. Candlesticks chart
# +
# load aapl csv file
df = pd.read_csv('../../database/hkex_ticks_day/hkex_0005.csv', header=0, index_col='Date', parse_dates=True)
# We select only the data from Nov to Dec 2019
df = df.loc[pd.Timestamp('2019-11-01'):pd.Timestamp('2019-12-31')]
df.head()
# -
df.index = mdates.date2num(df.index)
data = df.reset_index().values # Convert dataframe into 2-D list
print(data[:5,:])
# +
# import libaries
import matplotlib.dates as mdates
from mpl_finance import candlestick_ochl as candlestick
from matplotlib.dates import MO, TU, WE, TH, FR, SA, SU # for locator
fig = plt.figure()
fig.set_size_inches(18.5, 10.5)
ax1 = plt.subplot2grid((6,1), (0,0), rowspan=5, colspan=1)
ax2 = plt.subplot2grid((6,1), (5,0), rowspan=1, colspan=1, sharex=ax1)
# plot candlesticks
mpl_finance.candlestick_ohlc(ax1, data, width=0.7, colorup='g', colordown='r')
ax.grid() # show grids
############# x-axis locater settings #################
locator = mdates.AutoDateLocator() # interval automically set
#locator = mdates.DayLocator(bymonthday=None, interval=2) # day as unit
#locator = mdates.WeekdayLocator(byweekday=(MO, TH)) # every mon and thur
#locator = mdates.WeekdayLocator(byweekday=(MO)) # every mon
#locator = mdates.MonthLocator() # monthly
#locator = mdates.YearLocator() # yearly
ax1.xaxis.set_major_locator(locator) # as as interval in a-axis
ax1.xaxis.set_minor_locator(mdates.DayLocator())
############# x-axis locater settings #################
ax1.xaxis.set_major_formatter(mdates.AutoDateFormatter(locator)) # set x-axis label as date format
fig.autofmt_xdate() # rotate date labels on x-axis
pos = df['Open'] - df['Close'] < 0
neg = df['Open'] - df['Close'] > 0
ax2.bar(df.index[pos],df['Volume'][pos],color='green',width=1,align='center')
ax2.bar(df.index[neg],df['Volume'][neg],color='red',width=1,align='center')
#ax2.bar(df.index, df['Volume'])
plt.savefig('./figures/00-candlestick-chart.png', dpi=200)
plt.show()
# -
# #### 2.3 Bar chart
# + tags=[]
from matplotlib.lines import Line2D
def westerncandlestick(ax, quotes, width=0.2, colorup='k', colordown='r',
ochl=True, linewidth=0.5):
"""
Plot the time, open, high, low, close as a vertical line ranging
from low to high. Use a rectangular bar to represent the
open-close span. If close >= open, use colorup to color the bar,
otherwise use colordown
Parameters
----------
ax : `Axes`
an Axes instance to plot to
quotes : sequence of quote sequences
data to plot. time must be in float date format - see date2num
(time, open, high, low, close, ...) vs
(time, open, close, high, low, ...)
set by `ochl`
width : float
fraction of a day for the open and close lines
colorup : color
the color of the lines close >= open
colordown : color
the color of the lines where close < open
ochl: bool
argument to select between ochl and ohlc ordering of quotes
linewidth: float
linewidth of lines
Returns
-------
ret : tuple
returns (lines, openlines, closelines) where lines is a list of lines
added
"""
OFFSET = width / 2.0
lines = []
openlines = []
closelines = []
for q in quotes:
if ochl:
t, open, close, high, low = q[:5]
else:
t, open, high, low, close = q[:5]
if close >= open:
color = colorup
else:
color = colordown
vline = Line2D( xdata=(t, t), ydata=(low, high),
color=color, linewidth=linewidth, antialiased=True)
lines.append(vline)
openline = Line2D(xdata=(t - OFFSET, t), ydata=(open,open),
color=color, linewidth=linewidth, antialiased=True)
openlines.append(openline)
closeline = Line2D(xdata=(t , t+OFFSET), ydata=(close,close),
color=color, linewidth=linewidth, antialiased=True)
closelines.append(closeline)
ax.add_line(vline)
ax.add_line(openline)
ax.add_line(closeline)
ax.autoscale_view()
return lines, openlines, closelines
# +
fig = plt.figure()
fig.set_size_inches(18.5, 10.5)
ax1 = plt.subplot2grid((6,1), (0,0), rowspan=5, colspan=1)
ax2 = plt.subplot2grid((6,1), (5,0), rowspan=1, colspan=1, sharex=ax1)
# plot westrencandlestick
westerncandlestick(ax1, data, width=0.6, linewidth=1.44, ochl=False)
ax.grid() # show grids
############# x-axis locater settings #################
locator = mdates.AutoDateLocator() # interval automically set
#locator = mdates.DayLocator(bymonthday=None, interval=2) # day as unit
#locator = mdates.WeekdayLocator(byweekday=(MO, TH)) # every mon and thur
#locator = mdates.WeekdayLocator(byweekday=(MO)) # every mon
#locator = mdates.MonthLocator() # monthly
#locator = mdates.YearLocator() # yearly
ax1.xaxis.set_major_locator(locator) # as as interval in a-axis
ax1.xaxis.set_minor_locator(mdates.DayLocator())
############# x-axis locater settings #################
ax1.xaxis.set_major_formatter(mdates.AutoDateFormatter(locator)) # set x-axis label as date format
fig.autofmt_xdate() # rotate date labels on x-axis
pos = df['Open'] - df['Close'] < 0
neg = df['Open'] - df['Close'] > 0
ax2.bar(df.index[pos], df['Volume'][pos], color='green', width=1, align='center')
ax2.bar(df.index[neg], df['Volume'][neg], color='red', width=1, align='center')
#ax2.bar(df.index, df['Volume'])
plt.savefig('./figures/00-western-candlestick-chart.png', dpi=200)
plt.show()
# -
# #### Plotting OHLC data with mplfinance
#
# You can read the documentation [here](https://github.com/matplotlib/mplfinance).
# ! pip install --upgrade mplfinance
# +
import mplfinance as mpf
# load aapl csv file
df = pd.read_csv('../../database/nasdaq_ticks_day/nasdaq_AAPL.csv', header=0, index_col='Date', parse_dates=True)
# We select only the data from Jan to Dec 2019
df = df.loc[pd.Timestamp('2019-01-01'):pd.Timestamp('2019-03-31')]
plt.savefig('./figures/00-mplfinance-ohlc-chart.png', dpi=200)
mpf.plot(df, style='yahoo', title="AAPL - OHLC chart")
# -
# try a different style
mpf.plot(df, type='candle', style='brasil', title="AAPL - Candlesticks chart")
plt.savefig('./figures/00-mplfinance-candlestick-chart.png', dpi=200)
# #### 2.4 Renko chart
# +
# load aapl csv file
df = pd.read_csv('../../database/nyse_ticks_day/nyse_IBM.csv', header=0, index_col='Date', parse_dates=True)
# We select only the data from Jan to Dec 2019
df = df.loc[pd.Timestamp('2019-06-01'):pd.Timestamp('2020-01-01')]
mpf.plot(df, type='renko', returnfig=True, title='IBM - Renko chart')
plt.savefig('./figures/00-mplfinance-renko-chart.png', dpi=200)
# -
# #### 2.5 Point and figure chart
# +
# load aapl csv file
df = pd.read_csv('../../database/nyse_ticks_day/nyse_IBM.csv', header=0, index_col='Date', parse_dates=True)
# We select only the data from Jan to Dec 2019
df = df.loc[pd.Timestamp('2019-06-01'):pd.Timestamp('2019-12-31')]
mpf.plot(df, type='pnf', title="IBM - Point and Finger chart")
plt.savefig('./figures/00-mplfinance-pnf-chart.png', dpi=200)
# -
# #### 2.6 Arithmetic scaling & Semi-logarithmic scaling charts
# There are two types of scales for plotting charts - arithmetic or semi-logarithmic. As most of us who have studied science/mathematics should know, examples of logarithmic scales include growth of microbes, mortality rate due to epidemics and so on. The difference in scale can completely alter the shape of the chart even though it is plotted using the same set of data. Semi-logarithmic charts are sometimes more preferrable in order to overcome the weaknesses inherent in arithmetic charts.
# ##### 2.6.1 Arithmetic scaling
#
# In arithmetic or linear charts, both x and y axes scales are plotted at an equal distance.
#
# <b>Key points</b>
# * On a linear scale, as the distance in the axis increases the corresponding value also increases linearly.
# * When the values of data fluctuate between extremely small values and very large values – the linear scale will miss out the smaller values thus conveying a wrong picture of the underlying phenomenon.
#
# ##### 2.6.2 Semi-logarithmic scaling
#
# A semi-log plot is a graph where the data in one axis is on logarithmic scale (either x axis or y axis), and data in the other axis is on normal scale (i.e. linear scale).
#
# <b>Key points</b>
# * On a logarithmic scale, as the distance in the axis increases the corresponding value increases exponentially.
# * With logarithmic scale, both smaller valued data and bigger valued data can be captured in the plot more accurately to provide a holistic view.
#
# Therefore, semi-logarithmic charts can be of immense help especially when plotting long-term charts, or when the price points show significant volatility even in short-term charts. The underlying chart patterns will be revealed more clearly in semi-logarithmic scale charts.
# +
# load data
df = pd.read_csv('../../database/hkex_ticks_day/hkex_0005.csv', header=0, index_col='Date', parse_dates=True)
df = df.loc[pd.Timestamp('2013-01-01'):pd.Timestamp('2019-12-31')]
plt.style.use('ggplot')
fig, (ax1, ax2) = plt.subplots(1, 2)
fig.set_size_inches(18.5, 7.0)
### Subplot 1 - Semi-logarithmic ###
plt.subplot(121)
plt.grid(True, which="both")
# Linear X axis, Logarithmic Y axis
plt.semilogy(df.index, df['Close'], 'r')
plt.ylim([10,500])
plt.xlabel("Date")
plt.title('Semi-logarithmic scale')
fig.autofmt_xdate()
### Subplot 2 - Arithmetic ###
plt.subplot(122)
plt.plot(df.index, df['Close'], 'b')
plt.xlabel("Date")
plt.title('Arithmetic scale')
fig.autofmt_xdate()
# show plot
plt.savefig('./figures/00-semilog-vs-arithmetic-chart.png', dpi=200)
plt.show()
# -
# ### References
# * <NAME>., Point & Figure Charting, Wiley, 1995.
# * [Investopedia - Technical Analysis](https://www.investopedia.com/terms/t/technicalanalysis.asp)
# * [Investopedia - Candlestick Charts](https://www.investopedia.com/trading/candlestick-charting-what-is-it/)
# * [Drawing a semilog plot with matplotlib](https://pythontic.com/visualization/charts/semilog)
| code/technical-analysis_basics/chart-analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: auto-sklearn
# language: python
# name: auto-sklearn
# ---
# ## Introduction to Multi-Fidelity Optimization
# 在上一个教程中,我们学习了如何实现一个简单的AutoML系统。但这个AutoML系统的核心:贝叶斯优化算法往往需要大量对采样的评价才能获得比较好的结果。
#
# 然而,在自动机器学习(Automatic Machine Learning, AutoML)任务中评价往往通过 k 折交叉验证获得,在大数据集的机器学习任务上,`获得一个评价的时间代价巨大`。这也影响了优化算法在自动机器学习问题上的效果。所以一些`减少评价代价`的方法被提出来,其中**多保真度优化**(Multi-Fidelity Optimization)[<sup>[1]</sup>](#refer-anchor-1)就是其中的一种。而**多臂老虎机算法**(Multi-armed Bandit Algorithm, MBA)[<sup>[2]</sup>](#refer-anchor-2)是多保真度算法的一种。在此基础上,有两种主流的`bandit-based`优化策略:
#
# - Successive Halving (SH) [<sup>[3]</sup>](#refer-anchor-3)
# - Hyperband (HB) [<sup>[4]</sup>](#refer-anchor-4)
#
#
# 首先我们介绍连续减半(Successive Halving ,SH)。在连续减半策略中, 我们将`评价代价`参数化为一个变量`budget`,即预算。根据BOHB论文[<sup>[5]</sup>](#refer-anchor-5)的阐述,我们可以根据不同的场景定义不同的budget,举例如下:
#
# 1. 迭代算法的迭代数(如:神经网络的epoch、随机森林,GBDT的树的个数)
# 2. 机器学习算法所使用的样本数
# 3. 贝叶斯神经网络[<sup>[6]</sup>](#refer-anchor-6)中MCMC链的长度
# 4. 深度强化学习中的尝试数
#
# 举例说明,我们定义$budget_{max}=1$, $budget_{min}=\frac{1}{8}$, $\eta=2$ (`eta` = 2) 。在这里`budget`的语义表示使用$100\times budget$%的样本。
#
# 1. 首先我们从配置空间(或称为超参空间)**随机采样**8个配置,实例化为8个机器学习模型。
# 2. 然后用$\frac{1}{8}$的训练样本训练这8个模型并在验证集得到相应的损失值。
# 3. 保留这8个模型中loss最低的前4个模型,其余的舍弃。
# 4. 依次类推,最后仅保留一个模型,并且其`budget=1`(可以用全部的样本进行训练)
# 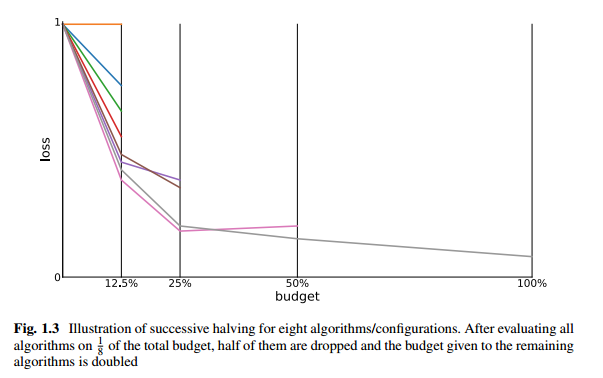
# 上图描述了例子中的迭代过程(图片来自[<sup>[1]</sup>](#refer-anchor-1)) 。我们可以用`ultraopt.multi_fidelity`中的`SuccessiveHalvingIterGenerator`来实例化这一过程:
from ultraopt.multi_fidelity import SuccessiveHalvingIterGenerator, HyperBandIterGenerator
SH = SuccessiveHalvingIterGenerator(min_budget=1/8, max_budget=1, eta=2)
SH.get_table()
# 接下来我们介绍HyperBand(HB)的策略。
SH = HyperBandIterGenerator(min_budget=1/8, max_budget=1, eta=2)
SH.get_table()
# ### Combine Multi-Fidelity Optimization and Bayesian Optimization in UltraOpt
# 我们注意到,上文描述的SH和HB策略在采样时都是**随机采样**,而`UltraOpt`将**优化器**和**多保真迭代生成器**这两个部分解耦和了,您可以将任意的**贝叶斯优化算法**和**多保真优化算法**进行组合。
#
# 这样的组合其实就是BOHB(Bayesian Optimization Hyperband)算法[<sup>[5]</sup>](#refer-anchor-5)。UltraOpt在很多代码上借鉴和直接使用了HpBandSter[<sup>[7]</sup>](#refer-anchor-7)这个开源项目,我们感谢他们优秀的工作。
# 如果您需要采用多保真优化策略,您的评价函数需要增加一个`float`类型的`budget`参数:
#
# ```python
# def evaluate(config: dict, budget:float) -> float :
# pass
# ```
# 为了测试, 我们采用`ultraopt.tests.mock`中自带的一个含有`budget`的评价函数,以及相应的配置空间:
from ultraopt.tests.mock import evaluate, config_space
from ultraopt import fmin
from ultraopt.multi_fidelity import HyperBandIterGenerator
# 在调用`ultraopt.fmin`函数时,采用多保真策略时需要做以下修改:
#
# 1. 需要指定`multi_fidelity_iter_generator`(多保真迭代生成器)
# 2. `n_iterations`参数与普通模式不同,不再代表评价函数的调用次数,而代表`iter_generator`的迭代次数,需要酌情设置
# 3. `parallel_strategy`需要设置为`AsyncComm`,不改变默认值就没事
# 首先我们实例化一个`iter_generator`(多保真迭代生成器),并根据`get_table()`函数的可视化结果设置`n_iterations`。
#
# 因为测试函数的`max_budget = 100`, 我们按照`25, 50, 100`来递增`budget`:
iter_generator = HyperBandIterGenerator(min_budget=25, max_budget=100, eta=2)
iter_generator.get_table()
result = fmin(evaluate, config_space, n_iterations=50, multi_fidelity_iter_generator=iter_generator, n_jobs=3)
result
# 按budget分组的随时间变化拟合曲线:
result.plot_convergence_over_time(yscale="log");
# `low_budget`推荐得到的优势配置会保留到`high_budget`,从而可以根据`loss-pairs`计算不同`budget`之间的相关性:
result.plot_correlation_across_budgets();
# ### Multi-Fidelity in AutoML Scenarios
# 虽然`ultraopt.tests.mock`中提供的合成函数可以测试结合多保真策略的优化,但这毕竟不是真实场景。
#
# 现在,我们就通过修改教程`05. Implement a Simple AutoML System`中的AutoML评价器,将其改造为一个支持多保真优化的评价器,并进行相应的测试。
# +
from sklearn.svm import LinearSVC
from sklearn.ensemble import RandomForestClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.datasets import load_digits
import seaborn as sns
import numpy as np
import warnings
from ultraopt.hdl import layering_config
from sklearn.model_selection import StratifiedKFold # 采用分层抽样
warnings.filterwarnings("ignore")
X, y = load_digits(return_X_y=True)
cv = StratifiedKFold(n_splits=3, shuffle=True, random_state=0)
def evaluate(config: dict, budget: float) -> float:
layered_dict = layering_config(config)
AS_HP = layered_dict['classifier'].copy()
AS, HP = AS_HP.popitem()
ML_model = eval(AS)(**HP)
# 注释掉采用原版的采用所有数据进行训练的方法(相当于budget=1)
# scores = cross_val_score(ML_model, X, y, cv=cv, scoring=metric)
# -------------------------------------------------------------
# 采用在对【 5折交叉验证中的训练集 】进行采样的方法,采样率为 budget
sample_ratio = budget
scores = []
for i, (train_ix, valid_ix) in enumerate(cv.split(X, y)):
rng = np.random.RandomState(i)
size = int(train_ix.size * sample_ratio)
train_ix = rng.choice(train_ix, size, replace=False)
X_train = X[train_ix, :]
y_train = y[train_ix]
X_valid = X[valid_ix, :]
y_valid = y[valid_ix]
ML_model.fit(X_train, y_train)
scores.append(ML_model.score(X_valid, y_valid))
# -------------------------------------------------------------
score = np.mean(scores)
return 1 - score
# -
config = {'classifier:__choice__': 'LinearSVC',
'classifier:LinearSVC:C': 1.0,
'classifier:LinearSVC:dual': 'True:bool',
'classifier:LinearSVC:loss': 'squared_hinge',
'classifier:LinearSVC:max_iter': 600,
'classifier:LinearSVC:multi_class': 'ovr',
'classifier:LinearSVC:penalty': 'l2',
'classifier:LinearSVC:random_state': '42:int'}
evaluate(config, 0.125)
evaluate(config, 0.5)
evaluate(config, 1)
# 可以看到我们已经成功定义了一个结合多保真策略的AutoML评价器,并且按照一般规律:budget越大,评价代价也越大,模型表现也越好,loss越小。
# 我们将上述代码整合到`05. Implement a Simple AutoML System.py`脚本中,形成`06. Combine Multi-Fidelity Optimization.py`脚本:
# +
# #!/usr/bin/env python
# -*- coding: utf-8 -*-
# @Author : <NAME>
# @Date : 2020-12-28
# @Contact : <EMAIL>.cn
import warnings
from sklearn.svm import LinearSVC
from sklearn.ensemble import RandomForestClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.datasets import load_digits
from sklearn.model_selection import StratifiedKFold # 采用分层抽样
from sklearn.model_selection import cross_val_score
import sklearn.metrics
import numpy as np
from ultraopt import fmin
from ultraopt.hdl import hdl2cs, plot_hdl, layering_config
from ultraopt.multi_fidelity import HyperBandIterGenerator
warnings.filterwarnings("ignore")
HDL = {
'classifier(choice)':{
"LinearSVC": {
"max_iter": {"_type": "int_quniform","_value": [300, 3000, 100], "_default": 600},
"penalty": {"_type": "choice", "_value": ["l1", "l2"],"_default": "l2"},
"dual": {"_type": "choice", "_value": [True, False],"_default": False},
"loss": {"_type": "choice", "_value": ["hinge", "squared_hinge"],"_default": "squared_hinge"},
"C": {"_type": "loguniform", "_value": [0.01, 10000],"_default": 1.0},
"multi_class": "ovr",
"random_state": 42,
"__forbidden": [
{"penalty": "l1","loss": "hinge"},
{"penalty": "l2","dual": False,"loss": "hinge"},
{"penalty": "l1","dual": False},
{"penalty": "l1","dual": True,"loss": "squared_hinge"},
]
},
"RandomForestClassifier": {
"n_estimators": {"_type": "int_quniform","_value": [10, 200, 10], "_default": 100},
"criterion": {"_type": "choice","_value": ["gini", "entropy"],"_default": "gini"},
"max_features": {"_type": "choice","_value": ["sqrt","log2"],"_default": "sqrt"},
"min_samples_split": {"_type": "int_uniform", "_value": [2, 20],"_default": 2},
"min_samples_leaf": {"_type": "int_uniform", "_value": [1, 20],"_default": 1},
"bootstrap": {"_type": "choice","_value": [True, False],"_default": True},
"random_state": 42
},
"KNeighborsClassifier": {
"n_neighbors": {"_type": "int_loguniform", "_value": [1,100],"_default": 3},
"weights" : {"_type": "choice", "_value": ["uniform", "distance"],"_default": "uniform"},
"p": {"_type": "choice", "_value": [1, 2],"_default": 2},
},
}
}
CS = hdl2cs(HDL)
g = plot_hdl(HDL)
default_cv = StratifiedKFold(n_splits=3, shuffle=True, random_state=0)
X, y = load_digits(return_X_y=True)
class Evaluator():
def __init__(self,
X, y,
metric="accuracy",
cv=default_cv):
# 初始化
self.X = X
self.y = y
self.metric = metric
self.cv = cv
def __call__(self, config: dict, budget: float) -> float:
layered_dict = layering_config(config)
AS_HP = layered_dict['classifier'].copy()
AS, HP = AS_HP.popitem()
ML_model = eval(AS)(**HP)
# scores = cross_val_score(ML_model, self.X, self.y, cv=self.cv, scoring=self.metric)
# -------------------------------------------------------------
# 采用在对【 5折交叉验证中的训练集 】进行采样的方法,采样率为 budget
sample_ratio = budget
scores = []
for i, (train_ix, valid_ix) in enumerate(self.cv.split(X, y)):
rng = np.random.RandomState(i)
size = int(train_ix.size * sample_ratio)
train_ix = rng.choice(train_ix, size, replace=False)
X_train = X[train_ix, :]
y_train = y[train_ix]
X_valid = X[valid_ix, :]
y_valid = y[valid_ix]
ML_model.fit(X_train, y_train)
y_pred = ML_model.predict(X_valid)
score = eval(f"sklearn.metrics.{self.metric}_score")(y_valid, y_pred)
scores.append(score)
# -------------------------------------------------------------
score = np.mean(scores)
return 1 - score
evaluator = Evaluator(X, y)
iter_generator = HyperBandIterGenerator(min_budget=1/4, max_budget=1, eta=2)
result = fmin(evaluator, HDL, optimizer="ETPE", n_iterations=30, multi_fidelity_iter_generator=iter_generator, n_jobs=3)
print(result)
# -
# 我们可以对结合多保真策略得到的优化结果进行数据分析:
import pylab as plt
plt.rcParams['figure.figsize'] = (16, 12)
plt.subplot(2, 2, 1)
result.plot_convergence_over_time();
plt.subplot(2, 2, 2)
result.plot_concurrent_over_time(num_points=200);
plt.subplot(2, 2, 3)
result.plot_finished_over_time();
plt.subplot(2, 2, 4)
result.plot_correlation_across_budgets();
# 图1的`budget分组拟合曲线`和图4`多budget间相关性图`我们在之前已经介绍过了,图2和图3分别阐述了`随时间的并行数`和`随时间的完成情况`。
# ---
#
# **参考文献**
#
#
#
# <div id="refer-anchor-1"></div>
#
# - [1] [<NAME>., <NAME>. (2019) Hyperparameter Optimization. In: <NAME>., <NAME>., <NAME>. (eds) Automated Machine Learning. The Springer Series on Challenges in Machine Learning. Springer, Cham.](https://link.springer.com/chapter/10.1007/978-3-030-05318-5_1#citeas)
#
# <div id="refer-anchor-2"></div>
#
# - [2] [<NAME> and <NAME>. “A Survey on Practical Applications of Multi-Armed and Contextual Bandits.” ArXiv abs/1904.10040 (2019): n. pag.](https://arxiv.org/abs/1904.10040)
#
# <div id="refer-anchor-3"></div>
#
# - [3] [<NAME>. and <NAME>. “Non-stochastic Best Arm Identification and Hyperparameter Optimization.” AISTATS (2016).](https://arxiv.org/abs/1502.07943)
#
# <div id="refer-anchor-4"></div>
#
# - [4] [<NAME>. et al. “Hyperband: A Novel Bandit-Based Approach to Hyperparameter Optimization.” J. Mach. Learn. Res. 18 (2017): 185:1-185:52.](https://arxiv.org/abs/1603.06560)
#
# <div id="refer-anchor-5"></div>
#
# - [5] [<NAME> et al. “BOHB: Robust and Efficient Hyperparameter Optimization at Scale.” ICML (2018).](https://arxiv.org/abs/1807.01774)
#
# <div id="refer-anchor-6"></div>
#
# - [6] [https://github.com/automl/pybnn](https://github.com/automl/pybnn)
#
#
# <div id="refer-anchor-7"></div>
#
# - [7] [https://github.com/automl/HpBandSter](https://github.com/automl/HpBandSter)
| tutorials/06. Combine Multi-Fidelity Optimization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Example usage
# Here we will demonstrate how to use `pycounts_rl` to count the words in a text file and plot the top 5 results.
# # Imports
from pycounts_rl.pycounts_rl import count_words
from pycounts_rl.plotting import plot_words
# # Create a text file
#
# We'll first create a text file to work with using a famous quote from Einstein:
quote = """Insanity is doing the same thing
over and over and expecting different results."""
with open ("einstein.txt","w") as file:
file.write(quote)
# # Count words
#
# We can count the words in our text file using the `count_words()` function. Note that this function removes punctuation and makes all words lowercase before counting.
counts = count_words("einstein.txt")
print(counts)
# # Plot words
# We can now plot the results using the `plot_words()` function:
fig = plot_words(counts, n=5)
| docs/example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/KhushbooSingh17/18CSE140/blob/main/Assignment2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="A-HxM_pV49Uv"
#Indian_food dataset comma separated value
path = "https://raw.githubusercontent.com/KhushbooSingh17/18CSE140/main/indian_food.csv"
# + id="qG3QQP-G5C3L"
import pandas as pd
# + id="X3_huhEj5Nt2"
data = pd.read_csv(path)
# + colab={"base_uri": "https://localhost:8080/", "height": 416} id="7M7Vr74o5RPh" outputId="796a22c6-b02b-43fd-9e41-0ddd32f4bee0"
data
# + id="nWQYTRPp5XZq"
data = pd.read_csv("https://raw.githubusercontent.com/KhushbooSingh17/18CSE140/main/indian_food.csv")
# + colab={"base_uri": "https://localhost:8080/", "height": 416} id="Ti2eLkhF5dW8" outputId="4bc8623f-a441-4537-a5fd-3a846d3f5c31"
data
# + colab={"base_uri": "https://localhost:8080/"} id="pWEKXvIa5jKf" outputId="9af42809-e192-47b2-a570-9172684f6458"
type(data)
# + colab={"base_uri": "https://localhost:8080/"} id="redCFfdm5tM_" outputId="05dfcbec-93f6-4548-df94-48f49cf7386a"
data.shape
# + colab={"base_uri": "https://localhost:8080/"} id="eiWgli6O5wrS" outputId="d4bfa1b2-969a-45d6-83e7-337dace677da"
data.info()
# + colab={"base_uri": "https://localhost:8080/"} id="WAIXXDlO6Cdt" outputId="50275c89-f627-468d-d61e-aa1e24f29158"
data.index
# + colab={"base_uri": "https://localhost:8080/"} id="w52vt5b06EBF" outputId="52f4641e-d7ff-41e2-e713-3f1e94008b07"
data.columns
# + colab={"base_uri": "https://localhost:8080/", "height": 202} id="vMH1oneQ6NlV" outputId="62cf8e74-4ed1-4d14-9907-fa27288fb1ee"
data.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 202} id="nEv_vIY-6UiR" outputId="88760ceb-fba1-4663-900f-0f1e15082506"
data.tail()
# + colab={"base_uri": "https://localhost:8080/", "height": 141} id="SvDastl86dG7" outputId="2367b75e-95ba-49a7-83de-fd37ebc0b8c3"
data.head(3)
# + colab={"base_uri": "https://localhost:8080/", "height": 356} id="tvmxByqo6h-L" outputId="53eed10d-accf-4a09-f222-f9d815a028ca"
data[['ingredients',"diet"]].head(10)
# + colab={"base_uri": "https://localhost:8080/"} id="SCcXhKfz6mF5" outputId="a9b4ef13-d73f-4e00-8642-cbe1f881f2b9"
### Data Wrangling (Working With Null Values)
data.isnull().sum()
# + colab={"base_uri": "https://localhost:8080/"} id="ncbulUDX6vWB" outputId="4145dd29-551f-435b-ff35-23fbd89a0ba9"
data.dropna(inplace=True) # removed the null values 1st method remove rows when large data we having
data.isnull().sum()
# + colab={"base_uri": "https://localhost:8080/"} id="Ia6Cj8n663OM" outputId="1fbe0556-a996-4593-dc7f-ed2aac55b918"
data.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 356} id="8H5gPXRD7Apc" outputId="3b8891ce-8ec7-4e53-97fb-d530dd6cf099"
data.head(10)
# + colab={"base_uri": "https://localhost:8080/"} id="Lb8__k5u7Ijr" outputId="6243277b-d6c7-4e8a-9cf0-4a2b7be0e2a7"
#2nd method handling missing values
data['prep_time'].mean()
# + colab={"base_uri": "https://localhost:8080/"} id="vcmg1m9S7SOR" outputId="6b7569d8-279e-46dd-b729-780aea4c2f1b"
data['prep_time'].head()
# + id="aVyeRlvq7XLl"
import numpy as np
# + colab={"base_uri": "https://localhost:8080/"} id="xnlxOFtc7aUS" outputId="d145054c-7fa6-4174-d481-31d20de4cbaa"
data['prep_time'].replace(np.NaN,data['prep_time'].mean()).head()
| Assignment2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from csbdeep.models import Config, CARE
import numpy as np
from csbdeep.utils import plot_some, plot_history
from csbdeep.utils.n2v_utils import manipulate_val_data
from matplotlib import pyplot as plt
import urllib
import os
import zipfile
os.environ["CUDA_VISIBLE_DEVICES"]="0"
# +
# We need to normalize the data before we feed it into our network, and denormalize it afterwards.
def normalize(img, mean, std):
zero_mean = img - mean
return zero_mean/std
def denormalize(x, mean, std):
return x*std + mean
# -
model = CARE(None, 'n2v_model', basedir='/home/prakash/Desktop/falcon/outdata/n2v_CTC_n10/train_100/')
path_train = '/home/prakash/Desktop/falcon/train_data/TrainVal10.npz'
train_val = np.load(path_train)
X_train = train_val['X_train']
X_val = train_val['X_val']
Y_train = train_val['Y_train']
Y_val = train_val['Y_val']
path_train_N2V = '/home/prakash/Desktop/falcon/train_data/TrainValN2V10.npz'
train_val_N2V = np.load(path_train_N2V)
X_train_N2V = train_val_N2V['X_train']
mean, std = np.mean(X_train_N2V), np.std(X_train_N2V)
path_test = '/home/prakash/Desktop/falcon/test_data/Test10.npz'
test = np.load(path_test)
X_test = test['X_test']
Y_test = test['Y_test']
X_train = normalize(X_train, mean, std)
X_val = normalize(X_val, mean, std)
X_test = normalize(X_test, mean, std)
predictions = []
# Denoise all images
for i in range(X_train.shape[0]):
predictions.append(denormalize(model.predict(X_train[i], axes='YX',normalizer=None ), mean, std))
X_train_d = np.array(predictions)
predictions = []
# Denoise all images
for i in range(X_val.shape[0]):
predictions.append(denormalize(model.predict(X_val[i], axes='YX',normalizer=None ), mean, std))
X_val_d = np.array(predictions)
predictions = []
# Denoise all images
for i in range(X_test.shape[0]):
predictions.append(denormalize(model.predict(X_test[i], axes='YX',normalizer=None ), mean, std))
X_test_d = np.array(predictions)
np.savez_compressed("/home/prakash/Desktop/falcon/train_data/TrainVal10_denoised_incomplete.npz", X_train=X_train_d, X_val=X_val_d,Y_train=Y_train, Y_val=Y_val)
np.savez_compressed("/home/prakash/Desktop/falcon/test_data/Test10_denoised_incomplete.npz", X_test=X_test_d, Y_test=Y_test)
| datasets/CTC/sequential/TrainValDenoising_and_saveNPZ10.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python3
# name: python3
# ---
# # 1. Numerical Differentiation
# Often in machine learning, we will need to calculate the derivative of a function programmatically. Now, in general we will use a library to accomplish this-however, understanding how the implementation works under the hood is most definitely a valuable skill.
#
# So, based on our knowledge of the derivative, we know that for a small magnitude $\epsilon$, the value of the derivative of a function $g(\omega)$ is approximately:
#
# $$\frac{d}{d\omega}g(\omega) \approx \frac{g(\omega + \epsilon) - g(\omega)}{\epsilon}$$
#
# So, in order to make a program that estimates the derivative of some function at a point, we can simply choose a small positive value $\epsilon$ and approximate all derivatives we come across as:
#
# $$\frac{g(\omega + \epsilon) - g(\omega)}{\epsilon}$$
class NumericalDerivative:
"""A function for computing the numerical derivative of an arbitrary
input function and user-chosen epsilon"""
def __init__(self, g):
# load in function to differentiate and user epsilon
self.g = g; self.epsilon = 10*-5
def __call__(self, w, **kwargs):
# make local copies
g, epsilon = self.g, self.epsilon
print(kwargs)
# set epsilon to desired value or use default
if 'epsilon' in kwargs:
epsilon = kwargs['epsilon']
# Compute derivative approximation and return
approx = (g(w + epsilon) - g(w))/epsilon
return approx
| Mathematics/archived/Derivate Calculators.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Desafio 4
#
# Neste desafio, vamos praticar um pouco sobre testes de hipóteses. Utilizaremos o _data set_ [2016 Olympics in Rio de Janeiro](https://www.kaggle.com/rio2016/olympic-games/), que contém dados sobre os atletas das Olimpíadas de 2016 no Rio de Janeiro.
#
# Esse _data set_ conta com informações gerais sobre 11538 atletas como nome, nacionalidade, altura, peso e esporte praticado. Estaremos especialmente interessados nas variáveis numéricas altura (`height`) e peso (`weight`). As análises feitas aqui são parte de uma Análise Exploratória de Dados (EDA).
#
# > Obs.: Por favor, não modifique o nome das funções de resposta.
# ## _Setup_ geral
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import scipy.stats as sct
import seaborn as sns
import statsmodels.api as sm
"""
%matplotlib inline
from IPython.core.pylabtools import figsize
figsize(12, 8)
sns.set()
"""
athletes = pd.read_csv("athletes.csv")
def get_sample(df, col_name, n=100, seed=42):
"""Get a sample from a column of a dataframe.
It drops any numpy.nan entries before sampling. The sampling
is performed without replacement.
Example of numpydoc for those who haven't seen yet.
Parameters
----------
df : pandas.DataFrame
Source dataframe.
col_name : str
Name of the column to be sampled.
n : int
Sample size. Default is 100.
seed : int
Random seed. Default is 42.
Returns
-------
pandas.Series
Sample of size n from dataframe's column.
"""
np.random.seed(seed)
random_idx = np.random.choice(df[col_name].dropna().index, size=n, replace=False)
return df.loc[random_idx, col_name]
# ## Inicia sua análise a partir daqui
# +
# Sua análise começa aqui.
linhaBra, linhaUsa, linhaCan = [], [], []
for i in range(athletes.shape[0]):
if athletes["nationality"][i] == "BRA":
linhaBra.append(i)
elif athletes["nationality"][i] == "USA":
linhaUsa.append(i)
elif athletes["nationality"][i] == "CAN":
linhaCan.append(i)
bra = athletes.loc[linhaBra]
usa = athletes.loc[linhaUsa]
can = athletes.loc[linhaCan]
# -
# ## Questão 1
#
# Considerando uma amostra de tamanho 3000 da coluna `height` obtida com a função `get_sample()`, execute o teste de normalidade de Shapiro-Wilk com a função `scipy.stats.shapiro()`. Podemos afirmar que as alturas são normalmente distribuídas com base nesse teste (ao nível de significância de 5%)? Responda com um boolean (`True` ou `False`).
def q1():
# Retorne aqui o resultado da questão 1.
amostraHeight3000 = get_sample(athletes, "height", 3000)
testeNormalidadeShapiro = sct.shapiro(amostraHeight3000)
plt.hist(amostraHeight3000, bins=25)
#plt.show()
sm.qqplot(amostraHeight3000)
#plt.show()
print(testeNormalidadeShapiro)
return False
pass
# __Para refletir__:
#
# * Plote o histograma dessa variável (com, por exemplo, `bins=25`). A forma do gráfico e o resultado do teste são condizentes? Por que?
# * Plote o qq-plot para essa variável e a analise.
# * Existe algum nível de significância razoável que nos dê outro resultado no teste? (Não faça isso na prática. Isso é chamado _p-value hacking_, e não é legal).
# ## Questão 2
#
# Repita o mesmo procedimento acima, mas agora utilizando o teste de normalidade de Jarque-Bera através da função `scipy.stats.jarque_bera()`. Agora podemos afirmar que as alturas são normalmente distribuídas (ao nível de significância de 5%)? Responda com um boolean (`True` ou `False`).
def q2():
# Retorne aqui o resultado da questão 2.
amostraHeight3000 = get_sample(athletes, "height", 3000)
testeNormalidadeBera = sct.jarque_bera(amostraHeight3000)
print(testeNormalidadeBera)
return False
pass
# __Para refletir__:
#
# * Esse resultado faz sentido?
# ## Questão 3
#
# Considerando agora uma amostra de tamanho 3000 da coluna `weight` obtida com a função `get_sample()`. Faça o teste de normalidade de D'Agostino-Pearson utilizando a função `scipy.stats.normaltest()`. Podemos afirmar que os pesos vêm de uma distribuição normal ao nível de significância de 5%? Responda com um boolean (`True` ou `False`).
def q3():
# Retorne aqui o resultado da questão 3.
amostraWeight3000 = get_sample(athletes, "weight", 3000)
testeNormalidadePearson = sct.normaltest(amostraWeight3000)
sm.qqplot(amostraWeight3000, fit=True, line="45")
#plt.show()
print(testeNormalidadePearson)
return False
pass
# __Para refletir__:
#
# * Plote o histograma dessa variável (com, por exemplo, `bins=25`). A forma do gráfico e o resultado do teste são condizentes? Por que?
# * Um _box plot_ também poderia ajudar a entender a resposta.
# ## Questão 4
#
# Realize uma transformação logarítmica em na amostra de `weight` da questão 3 e repita o mesmo procedimento. Podemos afirmar a normalidade da variável transformada ao nível de significância de 5%? Responda com um boolean (`True` ou `False`).
def q4():
# Retorne aqui o resultado da questão 4.
athletes["weight"] = np.log(athletes["weight"])
amostraWeight3000 = get_sample(athletes, "weight", 3000)
testeNormalidadePearson = sct.normaltest(amostraWeight3000)
sm.qqplot(amostraWeight3000, fit=True, line="45")
#plt.show()
print(testeNormalidadePearson)
return False
pass
# __Para refletir__:
#
# * Plote o histograma dessa variável (com, por exemplo, `bins=25`). A forma do gráfico e o resultado do teste são condizentes? Por que?
# * Você esperava um resultado diferente agora?
# > __Para as questão 5 6 e 7 a seguir considere todos testes efetuados ao nível de significância de 5%__.
# ## Questão 5
#
# Obtenha todos atletas brasileiros, norte-americanos e canadenses em `DataFrame`s chamados `bra`, `usa` e `can`,respectivamente. Realize um teste de hipóteses para comparação das médias das alturas (`height`) para amostras independentes e variâncias diferentes com a função `scipy.stats.ttest_ind()` entre `bra` e `usa`. Podemos afirmar que as médias são estatisticamente iguais? Responda com um boolean (`True` ou `False`).
def q5():
# Retorne aqui o resultado da questão 5.
testeHipotese = sct.ttest_ind(bra.dropna()["height"], usa.dropna()["height"])
print(testeHipotese)
return False
pass
# ## Questão 6
#
# Repita o procedimento da questão 5, mas agora entre as alturas de `bra` e `can`. Podemos afimar agora que as médias são estatisticamente iguais? Reponda com um boolean (`True` ou `False`).
def q6():
# Retorne aqui o resultado da questão 6.
testeHipotese = sct.ttest_ind(bra.dropna()["height"], can.dropna()["height"])
print(testeHipotese)
return True
pass
# ## Questão 7
#
# Repita o procedimento da questão 6, mas agora entre as alturas de `usa` e `can`. Qual o valor do p-valor retornado? Responda como um único escalar arredondado para oito casas decimais.
def q7():
# Retorne aqui o resultado da questão 7.
testeHipotese = sct.ttest_ind(usa.dropna()["height"], can.dropna()["height"])
return testeHipotese[1].round(8)
pass
# __Para refletir__:
#
# * O resultado faz sentido?
# * Você consegue interpretar esse p-valor?
# * Você consegue chegar a esse valor de p-valor a partir da variável de estatística?
| Modulo 5/main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Query and explore data included in WALIS
# This notebook contains scripts that allow querying and extracting data from the "World Atlas of Last Interglacial Shorelines" (WALIS) database. The notebook calls scripts contained in the /scripts folder. After downloading the database (internet connection required), field headers are renamed, and field values are substituted, following 1:n or n:n relationships. The tables composing the database are then saved in CSV, XLSS (multi-sheet), and geoJSON formats. The notebook also contains some plotting functions.
# ## Dependencies and packages
# This notebook calls various scripts that are included in the \scripts folder. The following is a list of the python libraries needed to run this notebook.
# +
#Main packages
import pandas as pd
import pandas.io.sql as psql
import geopandas
import pygeos
import numpy as np
import mysql.connector
from datetime import date
import xlsxwriter as writer
import math
from scipy import optimize
from scipy import stats
#Plots
import seaborn as sns
import matplotlib.pyplot as plt
from mpl_toolkits.axes_grid1 import make_axes_locatable
#Jupyter data display
import tqdm
from tqdm.notebook import tqdm_notebook
from IPython.display import *
import ipywidgets as widgets
from ipywidgets import *
#Geographic
from shapely.geometry import Point
from shapely.geometry import box
import cartopy as ccrs
import cartopy.feature as cfeature
#System
import os
import glob
import shutil
#pandas options for debugging
pd.set_option('display.max_columns', None)
pd.set_option('display.max_rows', None)
#Set a date string for exported file names
date=date.today()
dt_string = date.strftime("_%d_%m_%Y")
# Ignore warnings
import warnings
warnings.simplefilter(action='ignore', category=FutureWarning)
warnings.filterwarnings('ignore')
# -
# ## Import database
# Connect to the online MySQL database containing WALIS data and download data into a series of pandas data frames.
# + hide_input=false
## Connect to the WALIS database server
# %run -i scripts/connection.py
## Import data tables and show progress bar
with tqdm_notebook(total=len(SQLtables),desc='Importing tables from WALIS') as pbar:
for i in range(len(SQLtables)):
query = "SELECT * FROM {}".format(SQLtables[i])
walis_dict[i] = psql.read_sql(query, con=db)
query2 = "SHOW FULL COLUMNS FROM {}".format(SQLtables[i])
walis_cols[i] = psql.read_sql(query2, con=db)
pbar.update(1)
# %run -i scripts/create_outfolder.py
# -
# ## Query the database
# Now, the data is ready to be queried according to a user input. There are two ways to extact data of interest from WALIS. Run either one and proceed.
#
# 1. [Select by author](#Query-option-1---Select-by-author)
# 2. [Select by geographic coordinates](#Query-option-2---Select-by-geographic-extent)
# ### Query option 1 - Select by author
#
# This option compiles data from multiple users who collaborated to create regional datasets for the WALIS Special Issue in ESSD. Select "WALIS Admin" in the dropdown menu if you want to extract the entire database.
#
# **NOTE: If you want to change users, just re-run this cell and select a different set of values**
# %run -i scripts/select_user.py
multiUsr
# Once the selection is done, run the following cell to query the database and extract only the data inserted by the selected user(s)
# %run -i scripts/multi_author_query.py
# ### Query option 2 - Select by geographic extent
# This option allows the download of data by geographic extent, defined as maximum-minimum bounds on Latitude and Longitude. Use this website to quickly find bounding coordinates: http://bboxfinder.com.
# +
# bounding box coordinates in decimal degrees (x=Lon, y=Lat)
xmin=-69.292145
xmax=-68.616486
ymin=12.009771
ymax=12.435235
# Curacao: -69.292145,12.009771,-68.616486,12.435235
#2.103882,39.219487,3.630981,39.993956
# -
# From the dictionary in connection.py, extract the dataframes
# %run -i scripts/geoextent_query.py
# ## Substitute data codes
# The following code makes joins between the data, substituting numerical or comma-separated codes with the corresponding text values.
#
# **WARNING - MODIFICATIONS TO THE ORIGINAL DATA**
#
# <u>The following adjustments to the data are made:</u>
# 1. If there is an age in ka, but the uncertainty field is empty, the age uncertainty is set to 30%
# 2. If the "timing constraint" is missing, the "MIS limit" is taken. If still empty, it is set to "Equal to"
# %run -i scripts/substitutions.py
# %run -i scripts/make_summary.py
# ## Write output
# The following scripts save the data in Xlsx, CSV, and geoJSON format (for use in GIS software).
# %run -i scripts/write_spreadsheets.py
# %run -i scripts/write_geojson.py
print ('Done!')
# ## Explore queried data through graphs
# The following scrips produce a series of images representing different aspects of the data included in the database. Each graph is saved in the "Output/Images" folder in svg format.
#
# The following graphs can be plotted:
# 1. [Monthly data insertion/update](#Monthly-data-insertion/update)
# 2. [References by year of publication](#References-by-year-of-publication)
# 3. [Elevation errors](#Elevation-errors)
# 4. [Sea level index points](#Sea-level-index-points)
# 5. [Elevation and positioning histograms](#Elevation-and-positioning-histograms)
# 6. [Quality plots](#Quality-plots)
# 7. [Maps](#Maps)
# 8. [Radiometric ages distribution](#Radiometric-ages-distribution)
# ### Monthly data insertion/update
# This graph explores the timeline of data insertion or update in WALIS since its inception. Peaks in this graph correspond to data updated in bulk by the admin.
# %run -i scripts/Database_contributions.py
# ### References by year of publication
# This graph shows the year of publication of the manuscripts included in the WALIS "References" table. Note that these might not all be used in further data compilations.
References_query=References_query[References_query['Year'] != 0] #to eliminate works that are marked as "in prep" from the graph
# %run -i scripts/References_hist.py
# ### Elevation errors
# These two graphs show the measured elevation errors (plotted as Kernel Density Estimate) reported for sea-level data within WALIS. These include "RSL from statigraphy" data points and single coral or speleothems indicating former RSL positions. The difference in the two plots resides in the treatment of outliers. Points having elevation uncertainties higher than 3.5 times the median absolute deviation are excluded from the graph in the left. All points are considered on the graph on the right side.
#
# The outlier exclusion is bases on this reference:
# ><NAME> and <NAME> (1993), "Volume 16: How to Detect and Handle Outliers", The ASQC Basic References in Quality Control: Statistical Techniques, <NAME>, Ph.D., Editor.
#
# And was derived from this link: https://stackoverflow.com/questions/11882393/matplotlib-disregard-outliers-when-plotting
# %run -i scripts/Elevation_error.py
# ### Sea level index points
# This graph shows the frequency of sea-level indicators within the query, including the grouping in indicator types.
# %run -i scripts/SL_Ind_Hist.py
# ### Elevation and positioning histograms
# These graphs show the distributions of the elevation metadata (Elevation measurement technique and sea-level datum) used to describe sea-level datapoints in WALIS.
# %run -i scripts/Vrt_meas_hist.py
# %run -i scripts/SL_datum_hist.py
# ### Quality plots
# The RSL datapoints from stratigraphy contain two "data quality" fields, one for age and one for RSL information. Database compilers scored each site following standard guidelines (as per database documentation). This plot shows these quality scores plotted against each other. As the quality scores of one area can be better appreciated by comparison with other areas, tools to compare two nations or two regions are given.
# #### Overall quality of selected area
# %run -i scripts/Quality_plot.py
# #### Compare two nations
# %run -i scripts/select_nation_quality.py
box
# %run -i scripts/Quality_nations.py
# #### Compare two regions
# %run -i scripts/select_region_quality.py
box
# %run -i scripts/Quality_regions.py
# ## Maps
# In this section, the data is organized in a series of maps. Some styling choices are available.
# %run -i scripts/select_map_options.py
# %run -i scripts/Static_maps.py
# %run -i scripts/global_maps.py
# ## Radiometric ages distribution
# The code below plots the age distribution of radiometric ages within the query. The data is run through a Monte-Carlo sampling of the gaussian distribution of each radiometric age, and Kernel density estimate (KDE) plots are derived.
#Insert age limits to be plotted
min_age=0
max_age=300
# %run -i scripts/age_kde.py
# # Create ZIP archive
# Create a ZIP archive of the entire "Output" folder.
shutil.make_archive('Output', 'zip', Output_path)
# # Suggested acknowledgments
# WALIS is the result of the work of several people, within different projects. For this reason, we kindly ask you to follow these simple rules to properly acknowledge those who worked on it:
#
# 1. Cite the original authors - Please maintain the original citations for each datapoint, to give proper credit to those who worked to collect the original data in the field or in the lab.
# 2. Acknowledge the database contributor - The name of each contributor is listed in all public datapoints. This is the data creator, who spent time to make sure the data is standardized and (as much as possible) free of errors.
# 3. Acknowledge the database structure and interface creators - The database template used in this study was developed by the ERC Starting Grant "WARMCOASTS" (ERC-StG-802414) and is a community effort under the PALSEA (PAGES / INQUA) working group.
#
# Example of acknowledgments: The data used in this study were *[extracted from / compiled in]* WALIS, a sea-level database interface developed by the ERC Starting Grant "WARMCOASTS" (ERC-StG-802414), in collaboration with PALSEA (PAGES / INQUA) working group. The database structure was designed by <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME> and <NAME>. The data points used in this study were contributed to WALIS by *[list names of contributors here]*.
| Code/.ipynb_checkpoints/Query_and_Explore_data-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## 歌曲序列建模
# by [@寒小阳](http://blog.csdn.net/han_xiaoyang)
# ### NLP场景下 - One Shot 词编码
# 我是中国人 => 我 是 中国 人 # 先分词<br>
# 我 => [1,0,0,0] # 再编码 1*500<br>
# 是 => [0,1,0,0] 1乘以500的向量<br>
# ...<br>
# 1*4 vector
# one-hot encoding
#
# 将word 映射成向量=> vector
#
# distance similarity
#
# #### 判断周边的环境 - google word2vec 可以在CPU上执行 - 窗口大小
# ### Online: C++
# ### Offline: Gensim -> python 的Word2Vec
# 
# ```python
# # word2vec 获得习近平 周边的信息
# result = model.most_similar(u"习近平")
# for e in result:
# print e[0], e[1]
# ```
# ```
# 胡锦涛 0.809472680092
# 江泽民 0.754633367062
# 李克强 0.739740967751
# 贾庆林 0.737033963203
# 曾庆红 0.732847094536
# 吴邦国 0.726941585541
# 总书记 0.719057679176
# 李瑞环 0.716384887695
# 温家宝 0.711952567101
# 王岐山 0.703570842743
# ```
# 说起来word2vec,其实就是把词映射成一定维度的稠密向量,同时保持住词和词之间的关联性,主要体现在(欧式)距离的远近上。
#
# 那么问题来了,word2vec为什么能够学习到这样的结果?
#
# 因为我们相信“物以类聚,人以群分” “一个人的层次与他身边最近的一些人是差不多的”
#
# 同样的考虑,我们是不是可以认为,一个歌单里的歌曲,相互之间都有一定的关联性呢?就像句子中的词一样。答案是,是的!
#
# 咱们来写个程序跑一把。
# ### 从word2vec到song2vec
# 我们把歌曲的id序列取出来,类比于分完词后的句子,送到word2vec中去学习一下,看看会有什么效果。
# +
#coding: utf-8
# 当做模型:用于序列建模
import multiprocessing
import gensim # NLP 中的主要model,涵盖word2vec
import sys
from random import shuffle
# 解析歌单,将序列存在plylist
def parse_playlist_get_sequence(in_line, playlist_sequence): # playlist_sequence is output file
song_sequence = []
#print "*** in_line is*** ", in_line
contents = in_line.strip().split("\t") # 空格分割
#print "contents >>>>> is ",contents
#print "contents[1:] >>>>> is ",contents[1:]
# 解析歌单序列
for song in contents[1:]:
try:
song_id, song_name, artist, popularity = song.split(":::")
song_sequence.append(artist) # song_id
except:
print "song format error"
print song+"\n"
for i in range(len(song_sequence)): #word2vec 窗口为7
shuffle(song_sequence)
playlist_sequence.append(song_sequence)
def train_song2vec(in_file, out_file):
#所有歌单序列
playlist_sequence = []
#遍历所有歌单
for line in open(in_file): # line 指的是从in_file 读取的每一 “行” 的文本数据
#print "line is >>>", line
parse_playlist_get_sequence(line, playlist_sequence)
#print ">>> playlist_sequence is ",playlist_sequence[1:10]
#使用word2vec训练
cores = multiprocessing.cpu_count()
print "using all "+str(cores)+" cores"
print "Training word2vec model..."
# min_count = 3,最少的出现次数
# window = 7 窗口长度为7
# workers=cores 双核处理
model = gensim.models.Word2Vec(sentences=playlist_sequence, size=150, min_count=3, window=7, workers=cores)
print "Saving model..."
model.save(out_file)
# -
# ### 整个训练的过程实际是针对某首歌曲,查找到“最近的”歌曲 (向量距离最近的歌曲)
# +
song_sequence_file = "./RawData/popular.playlist" # 在歌单playlist 里面 取出一部分华语流行的部分
model_file = "./Model/song2vec_artist.model"
# 下面是用于训练的过程
# %time train_song2vec(song_sequence_file, model_file)
# -
# 模型已经训练完了,咱们来试一把预测,看看效果
#
# 这个预测的过程,实际上就是对某首歌曲,查找“最近”的歌曲(向量距离最近的歌曲)
import cPickle as pickle # 歌曲Id到歌曲的映射pickle
song_dic = pickle.load(open("./InternalData/7_popular_song.pkl","rb"))
model_str = "./Model/song2vec_artist.model"
model = gensim.models.Word2Vec.load(model_str)
for song in song_dic.keys()[:10]:
print song, song_dic[song]
song_id_list = song_dic.keys()[1000:1500:50]
for song_id in song_id_list:
result_song_list = model.most_similar(song_id)
print song_id, song_dic[song_id]
print "\n相似歌曲 和 相似度 分别为:"
for song in result_song_list:
print "\t", song_dic[song[0]], song[1]
print "\n"
# ### 进一步思考
# 所以我们用word2vec学会了哪些歌曲和哪些歌曲最接近。
#
# 我们来思考一些很现实同时又很难解决的问题。比如:
# #### 1)冷启动问题
# 我们经常会遇到冷启动的问题,比如没有任何信息的歌曲,我们如何对它做推荐呢?
# * 如果是歌手发行的新歌曲,我们怎么进行推荐呢?
# * 如果我听完(并收藏)了一首很冷门的歌,怎么进行推荐呢?
#
# 我们知道新歌(或者小众的歌)是非常难和其他的歌关联上的,我们有的信息太少了(很少有用户在它上面发生行为)。
#
# 1.1 一种解决办法当然是推荐热门的歌曲,但是其实没从个人兴趣出发,我们知道这并不是最好的办法,并没有太大的卵用。
#
# 1.2 我们把问题的粒度放粗一点,用同样的思路,比如一个可考虑的解决方案是,我们把**歌曲**的粒度上升到对应的**歌手**,把刚才的song_list替换成artist_list,重新用word2vec建模,这样我们可以得到和一个歌手最相关(接近)的歌手,再推荐这个歌手最热门的歌曲,相对1.1的方法针对性强一些。
#
# 将商品 上升到=> 品类<br>
# 品类list => 送到word2vec里面去学习<br>
# [上衣,上衣,上衣,牛仔裤,牛仔裤,连衣裙...]<br>
#
# #### 2)用户兴趣预测问题
# 我们刚才完成的功能,类似酷狗音乐和网易音乐里针对一首歌的**“相似音乐”**,那么问题又来了,如果我们现在要对一个user用这套song2vec的方式推荐,我们怎么做呢?
#
# * 每个人的兴趣都是有时效性的,这意味着说,3年前我喜欢王菲的歌,去年我喜欢五月天的歌,而今年我可能就改摇滚路线,喜欢汪峰的歌了。
# * 每一首歌的热度也是不一样的,有一些热门的歌,如果用户能喜欢,当然是首选
#
# 那么,我们来做一个粗暴一点点的处理,把这2个维度拉进来,一起来针对一个用户做推荐。
#
# **把每个用户喜欢(收藏)过的歌,沿着时间轴排好,同时由近到远给不同的衰减因子(比如最近一首歌是1,前一首是0.98,再前一首是0.98^2,以此类推...),同时我们针对不同的歌曲热度,给定不同的推荐因子(比如热度100的是1,热度80的是0.9...),每一首歌都可以拿回一个song2vec的推荐列表和对应的相似度,对相似度以时间衰减因子和热度权重进行加权,最后的结果排序后,展示给用户。
| RecommendSystem/.ipynb_checkpoints/3.sequence_modelling-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # CSV file with JSON columns
#
# This notebook will take a csv file with one or more columns in json format and expand those json columns. I use this notebook to process some O365 Security and Compliance log files in the absence of the higher-tiered licensing that provide in-place analysis.
#
# #### Requirements: The CSV file should have headers. Each cell within a JSON column should contain the same fields/names. Non-integer row indices are okay. Row indices are preserved.
import pandas as pd
import json
# ### The following cell identifies the CSV file to be processed
#
# You can statically specify your file in the leading section or else enter it when prompted.
# +
####################
#ENTER YOUR DATA HERE: Path to csv file
#If file path is not statically specified here, the user will be prompted on execution of cell.
csv_file_path = ""
####################
file_found = False
while file_found == False:
#Prompt user for file path and name if not entered above statically
if csv_file_path == "":
print("Please specify the path and filename of your csv file:")
csv_file_path = input()
#Convert csv file into a dataframe
try:
csv_df = pd.read_csv(csv_file_path)
file_found = True
break
except FileNotFoundError:
print("File not found: Please check your csv file path and name and rerun this cell.")
except pd.errors.EmptyDataError:
print("Data is Empty: Please check the contents of your file and rerun this cell.")
except pd.errors.ParserError:
print("Parsing Error: Please check the format of your file contents and rerun this cell.")
csv_file_path = ""
if csv_df.shape[0] < 1:
print("Zero rows found. Please ensure your csv file has data in its rows.")
# -
#Identify columns and preview dataframe
columns = csv_df.columns
print(columns)
csv_df
# ### The following cell determines which columns contain JSON and should be expanded
#
# You can statically specify the columns, allow the code to autodect (based on the contents of the first row), or indicate column by column which to classify as json columns.
#
# If you leave json_columns as an empty array, you will be presented with the columns that were autodetected as json and you can press enter (submit an empty response to the input request) to accept that default and continue. If you submit a non-empty response, you will be prompted to identify each column. You may also consider stopping the cell after the columns have been autodetected so you can copy and paste the columns into the leading section if you have only a small tweak to make.
# +
####################
#ENTER YOUR DATA HERE: Specify json columns as a list of column names (strings).
#Columns are printed from the previous cell for review. User will be prompted if not specified here.
json_columns = []
####################
if len(json_columns) == 0:
for col in columns:
if type(csv_df.iloc[0][col]) == str:
try:
json.loads(csv_df.iloc[0][col])
json_columns.append(col)
except json.JSONDecodeError:
continue
print("The following have been identified as json columns. Please rerun this cell if not correct.")
print(json_columns)
print("Press Enter to accept, otherwise enter any character to manually choose columns instead.")
manual_select = input()
if manual_select != "":
json_columns = []
#Prompt user to identify json columns if not identified statically above
print("For each column, enter 1 to identify a json column and 0 otherwise.")
for col in columns:
print(col,end=": ")
response = input()
if response == '1':
json_columns.append(col)
print("The following have been identified as json columns. Please rerun this cell if not correct.")
print(json_columns)
# +
#Construct a new column list with extra columns to accomodate the json expansions
new_columns = []
for col in columns:
if col in json_columns:
json_string = csv_df.iloc[0][col]
example = json.loads(json_string)
sub_columns = list(example.keys())
for sub_col in sub_columns:
#Prefix columns derived from json expansion with original column name
new_columns.append(col+"_"+sub_col)
else:
new_columns.append(col)
#Construct an empty dataframe with new column names
json_expanded_df = pd.DataFrame(columns = new_columns)
json_expanded_df
# +
new_row = []
for index, row in csv_df.iterrows():
for col, value in csv_df.loc[index].items():
if col in json_columns:
new_row.extend(list(json.loads(value).values()))
else:
new_row.append(value)
json_expanded_df.loc[index] = new_row
new_row = []
json_expanded_df
# +
# Append _json_expanded to original filename
if len(csv_file_path) > 4 and csv_file_path[len(csv_file_path)-4:] == ".csv":
csv_json_expanded_path = csv_file_path[:len(csv_file_path)-4]+'_json_expanded.csv'
else:
print('There was a problem modifying the file path. The csv file will be saved in the root directory for jupyter notebooks as csv_json_expanded.csv instead.')
csv_json_expanded_path = 'csv_json_expanded.csv'
json_expanded_df.to_csv(csv_json_expanded_path)
# -
| csv-json-expansion.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # A protocol for identifying problems in continuous movement data
#
# This notebook provides an open-source implementation of the protocol presented in the paper:
#
# <NAME>. (2021) An exploratory data analysis protocol for identifying problems in continuous movement data. *Journal of Location Based Services.* http://dx.doi.org/10.1080/17489725.2021.1900612.
#
# The individual protocol steps are demonstrated using a dataset of vessel tracking data (AIS) published by the Danish Maritime Authority. The demo data covers two days (July, 1st 2017 and January, 1st 2018). Since the datasets are too large for Github, they have been made available via Figshare: https://doi.org/10.6084/m9.figshare.11577543
#
#
# ## Content
#
# - [Setup](#Setup)
# - [A. Missing data](#A.-Missing-data)
# - [B. Precision problems](#B.-Precision-problems)
# - [C. Consistency problems](#C.-Consistency-problems)
# - [D. Accuracy problems](#D.-Accuracy-problems)
# ## Setup
#
# Before running this notebook, make sure to [download](https://doi.org/10.6084/m9.figshare.11577543) *dk_csv_20170701.7z* and *dk_csv_20180101.7z* and unzip the files into the data directory:
input_files = [
'./data/aisdk_20170701.csv',
'./data/aisdk_20180101.csv'
]
# +
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:100% !important; }</style>"))
import warnings
warnings.filterwarnings('ignore')
# -
FIGSIZE = (600,400)
SMSIZE = 300
COLOR = 'darkblue'
COLOR_HIGHLIGHT = 'red'
COLOR_BASE = 'grey'
# +
from math import sin, cos, atan2, radians, degrees, sqrt, pi
from datetime import datetime, date
import numpy as np
import pandas as pd
import geopandas as gpd
import movingpandas as mpd
import datashader as ds
import holoviews as hv
from shapely.geometry import Point, LineString
from holoviews.operation.datashader import datashade, spread
from holoviews.element import tiles
from holoviews import opts, dim
import hvplot
import movingpandas as mp
from shapely.geometry import Point
R_EARTH = 6371000 # radius of earth in meters
C_EARTH = 2 * R_EARTH * pi # circumference
BG_TILES = tiles.CartoLight()
pd.set_option('use_inf_as_na', True)
# -
def plot_single_mover(df, mover_id, the_date):
tmp = df[(df.id==mover_id) & (df.index.date==the_date)]
gdf = gpd.GeoDataFrame(tmp.drop(['x', 'y'], axis=1), crs={'init': 'epsg:3857'}, geometry=[Point(xy) for xy in zip(tmp.x, tmp.y)])
plot = mp.Trajectory(gdf, 1).hvplot(title=f'Mover {mover_id} ({the_date})', c='speed_m/s', cmap='RdYlBu', colorbar=True, clim=(0,15),
line_width=5, width=FIGSIZE[0], height=FIGSIZE[1], tiles='CartoLight')
return plot
df = pd.read_csv(input_files[0], nrows=100)
df.head()
df['SOG'].hist(bins=100, figsize=(15,3))
# +
df = None
for input_file in input_files[:2]:
a = pd.read_csv(input_file, usecols=['# Timestamp', 'MMSI', 'Latitude', 'Longitude', 'SOG', 'Type of mobile', 'Ship type', 'Navigational status'])
a = a[(a['Type of mobile'] == 'Class A') & (a.SOG>0)]
a.drop(columns=['Type of mobile', 'SOG'], inplace=True)
if df is None:
df = a
else:
df = df.append(a)
df.rename(columns={'# Timestamp':'time', 'MMSI':'id', 'Latitude':'lat', 'Longitude':'lon', 'Ship type':'shiptype', 'Navigational status':'navstat'}, inplace=True)
df['time'] = pd.to_datetime(df['time'], format='%d/%m/%Y %H:%M:%S')
# +
df.loc[:, 'x'], df.loc[:, 'y'] = ds.utils.lnglat_to_meters(df.lon, df.lat)
df.set_index('time', inplace=True)
df['navstat'] = df['navstat'].astype('category')
df['shiptype'] = df['shiptype'].astype('category')
# -
df.head()
print('Number of records: {} million'.format(round(len(df)/1000000)))
# ## A. Missing data
#
# Checking for missing data is a common starting point for exploring new movement datasets. Missing data may indicate issues with the data collection process or the data export used to generate the analysis dataset. At this early stage, we usually start with raw location records that have yet to be aggregated into trajectories. Therefore, initial analyses look at elementary position records. The following protocol steps target issues of missing data with respect to movement data's spatial, temporal, and attribute dimensions.
#
#
# ### A-1. Spatial gaps & outliers
#
# To gain an overview, the visual analysis should start from the whole time span before drilling down. Spatial context (usually in the form of base maps) is essential when assessing spatial extent and gaps because context influences movement.
# #### Spatial spread / extent & outliers
#
# This step addresses the question if the dataset covers the expected spatial extent. This can be as simple as checking the minimum and maximum coordinate values of the raw records. However, it is not uncommon to encounter spurious location records or outliers that are not representative of the actual covered extent. These outliers may be truly erroneous positions but can also be correct positions that happen to be located outside the usual extent. Looking at elementary position records only, it is usually not possible to distinguish these two cases. It is therefore necessary to take note of these outliers and investigate further in later steps.
#
# Unexpected spatial extent can have different consequences, depending on whether the extent is too small, too large or covering a wrong area. If the area of interest to the analyst's work is not covered, it may be necessary to go back to the data collection phase. If the extent is larger than expected, it can cause excessive processing run times. For example, if the dataset is to be rasterized with a fixed target raster cell size, an unexpected large extent will result in a larger than expected raster. Additionally, any outliers that exceed the valid coordinate ranges, for example, for latitude and longitude values, should inform the analyst to proceed with caution and stay alert regarding other potential data quality issues.
#
# Classic scatter plots (or point maps) are helpful at this step. Point density maps (often called heat maps) on their default settings tend to hide outliers and are therefore not recommended. Interactive map visualizations should be preferred to static maps since interaction capabilities (such as zooming and panning) enable a quicker assessment of the situation
print(f'Spatial extent: x_min={df.lon.min()}, x_max={df.lon.max()}, y_min={df.lat.min()}, y_max={df.lat.max()}')
def plot_basic_scatter(df, color='darkblue', title='', width=FIGSIZE[0], height=FIGSIZE[1], size=2):
opts.defaults(opts.Overlay(active_tools=['wheel_zoom']))
pts = df.hvplot.scatter(x='x', y='y', datashade=True, cmap=[color, color], frame_width=width, frame_height=height, title=str(title))
return BG_TILES * spread(pts, px=size)
plot_basic_scatter(df, title='Spatial extent & outliers')
# **Optional cropping of outliers**
df = df[(df.lon>-90) & (df.lon<90) & (df.lat>0) & (df.lat<80)]
cropped_df = df[(df.lon>0) & (df.lon<20) & (df.lat>52) & (df.lat<60)]
cropped_df['navstat'] = cropped_df['navstat'].astype('category')
cropped_df['shiptype'] = cropped_df['shiptype'].astype('category')
plot_basic_scatter(cropped_df, title='Cropped dataset')
# #### Spatial gaps (selected areas / all movers / whole time span)
#
# This step addresses the question if there are spatial gaps in the data coverage. Depending on the type of movers, gaps in certain spatial contexts are to be expected. For example, we wouldn't expect taxi locations in lakes. Therefore, it is essential to evaluate these gaps in their spatial context using base maps showing relevant geographic features, such as the road network for vehicle data or navigation markers for vessel data.
#
# Unexpected spatial gaps are particularly problematic if they affect the area of interest to the analyst's work. However, even if the area of interest is not affected, spatial gaps may indicate systematic issues with the data collection process, such as observation gaps, that require addressing to achieve continuous coverage.
#
# Point density maps are helpful since they make it easy to identify areas with low densities, ignoring occasional outliers. The visualization scale influences which size of gaps can be discovered. However, there are of course practical limitations to exploring ever more detailed scales and resulting continuously growing numbers of gaps.
def plot_point_density(df, title='', width=FIGSIZE[0], height=FIGSIZE[1]):
opts.defaults(opts.Overlay(active_tools=['wheel_zoom']))
pts = df.hvplot.scatter(x='x', y='y', title=str(title), datashade=True, frame_width=width, frame_height=height)
return BG_TILES * pts
plot_point_density(df, title='Spatial gaps')
# ### A-2. Temporal gaps & outliers
#
# #### Temporal extent & outliers (whole territory / all movers / whole time span)
#
# This step addresses the question if the dataset covers the expected temporal extent. Similar to exploring the spatial extent, the obvious step is to determine the minimum and maximum timestamps first. Since GPS tracking requires accurate clocks to function, time information on the tracker is usually reliable. However, it is not guaranteed that these timestamps make it through the whole data collection and (pre)processing chain leading up to the exploratory analysis. For example, in some cases, tracker (or sender) time is replaced by receiver or storage time. Thus clock errors on the receiving or storage devices can result in unexpected timestamps.
#
# Undiscovered timestamp issues can affect the derived temporal extent, as well as all other temporal and spatiotemporal analyses. A typical problem (especially when working with CSV data sources) is erroneous parsing of dates, for example, switching the digits for day and month when they cannot be inferred unambiguously. This leads to wrong temporal assignments, such as records from Feb, 3rd being assigned to March, 2nd. In other datasets, time information may be provided in the form of an offset value from a certain starting time. In this case, the record timestamps have to be reconstructed in order to put the movement data in context with other spatiotemporal data.
#
# Temporal charts, particularly record counts over time, are helpful to gain a first impression of the overall temporal extent and whether it is continuous or split into multiple time frames with little or no data in between.
#
print(f'Temporal extent: {df.index.min()} to {df.index.max()}')
# +
TIME_SAMPLE = '15min'
df['id'].resample(TIME_SAMPLE).count()\
.hvplot(title=f'Number of records per {TIME_SAMPLE}', width=FIGSIZE[0])
# -
# #### Temporal gaps in linear sequence & temporal cycles (whole territory / all movers / time spans)
#
# This step addresses the question if there are temporal gaps in the dataset. Temporal gaps can be due to scheduled breaks in data collection, deliberate choices during data export, as well as unintended issues during data collection or (pre)processing. Similar to exploring spatial gaps, the temporal scale influences which size of gaps can be discovered. Temporal gaps can be one-time events or exhibit reoccurring patterns. For example, daily and weekly cycles are typical for human movement data.
#
# Undiscovered one-time, as well as reoccurring temporal gaps can be benign if they reflect mover behavior. However, if they are caused by systematic errors in the data collection or (pre)processing workflows, they can affect the validity of analysis and model results. For example, a data-driven model may predict low mover density based on its erroneous training data even though the real movement situation may be different.
#
# Two-dimensional time histograms are particularly helpful to discover reoccurring temporal gaps. Typical temporal cycles that may be worth exploring include daily, weekly, monthly, and yearly cycles. As already discussed with regards to spatial gaps, the discovery of temporal gaps is also affected by scale, that is, the temporal binning influences which size of gaps can be discovered.
#
counts_df = df['id'].groupby([df.index.hour, pd.Grouper(freq='d')]).count().to_frame(name='n')
counts_df.rename_axis(['hour', 'day'], inplace=True)
counts_df.hvplot.heatmap(title='Record count', x='hour', y='day', C='n', width=FIGSIZE[0])
# ### A-3. Spatiotemporal changes / gaps
#
# While the previous two steps looked at spatial gaps over the whole time span or temporal gaps for the whole territory, this step aims to explore spatiotemporal changes and gaps.
#
# #### Changing extent
#
# This step addresses the question whether there are changes in spatial extent over time. Changing spatial extent may be due to planned extensions or reductions of the data collection / observation area. Similarly, the extent is also expected to shift if the movers collectively change their location, as is the case, for example, with tracks of migrating birds.
#
# Unexpected changing or shifting extents mean that data may not be available for the full temporal extent for the whole area. This can limit the area suitable for spatiotemporal analyses, such as trend detection or training predictive models. Alternatively, analysts need to chose analysis methods and models that can handle missing data. In this case, care has to be taken when interpreting results and comparing performance in different regions with different data availability.
#
# Small multiples provide a quick way to compare extents between different time spans. To make sure that outliers can be easily spotted, classical point maps should be preferred to density maps.
#
# +
def plot_multiple_by_day(df, day, **kwargs):
return plot_basic_scatter(df[df.index.date==day], title=day, width=SMSIZE, height=SMSIZE, **kwargs)
def plot_multiples_by_day(df, **kwargs):
days = df.index.to_period('D').unique()
a = None
for a_day in days:
a_day = a_day.to_timestamp().date()
plot = plot_multiple_by_day(df, a_day, **kwargs)
if a is None: a = plot
else: a = a + plot
return a
# -
plot_multiples_by_day(df).cols(2)
plot_multiples_by_day(cropped_df).cols(2)
# +
def plot_multiple_by_hour_of_day(df, hour, fun):
return fun(df[df.index.hour==hour], title=hour, width=SMSIZE, height=SMSIZE)
def plot_multiples_by_hour_of_day(df, hours=range(0,24), fun=plot_basic_scatter):
a = None
for hour in hours:
plot = plot_multiple_by_hour_of_day(df, hour, fun)
if a is None: a = plot
else: a = a + plot
return a
# -
plot_multiples_by_hour_of_day(df, hours=[6,7,8,9]).cols(2)
# #### Temporary gaps
#
# This step addresses the question whether there are temporary gaps in the overall spatial coverage. These local gaps may be one-time or reoccurring issues. Like temporary changes in the overall extent, temporary gaps can be due to mover behavior, as well as planned and unplanned changes of the data collection or (pre)processing workflows.
#
# Unexpected temporary local gaps can have similar consequences as temporal gaps. However, due to their localize nature they may be harder to spot and can therefore remain hidden for longer. Resulting delays in discovering data issues can be costly, for example, because time-intensive model training has to be repeated.
#
# Besides small multiples of density maps animated density maps can be helpful at this step. Care should be taken to ensure that the color map configuration is consistent between time frames, that is, that the minimum and maximum values do not change since, otherwise, the density maps are not comparable.
#
plot_multiples_by_hour_of_day(cropped_df, hours=[0,6,12,18], fun=plot_point_density).cols(2)
# ### A-4. Attribute gaps
#
# Some attributes may only be available during certain time spans / or in certain areas.
#
# #### Spatial attribute gaps
#
# This step addresses the question if there are areas with missing attribute data. Locally missing attribute data can be due to heterogeneous data collection setups.
#
# Unexpected changes in attribute coverage can severely limit the usefulness of affected records. If a certain attribute is essential for further analysis or modeling but is not available in all regions and cannot be inferred by any other means, the consequence of these attribute gaps can be as severe as spatial gaps with completely missing records.
#
# The methods used to explore spatial extent and gaps can be adopted to missing attribute data. Small multiples can be used to compare the spatial distribution of records with and without certain attribute values.
#
CATEGORY = 'shiptype'
cats = df[CATEGORY].unique()
cmap = {}
for cat in cats:
cmap[cat] = COLOR_BASE
cmap['Unknown value'] = COLOR_HIGHLIGHT
cmap['Undefined'] = COLOR_HIGHLIGHT
def plot_categorized_scatter(df, cat, title='', width=SMSIZE, height=SMSIZE, cmap=cmap):
opts.defaults(opts.Overlay(active_tools=['wheel_zoom']))
pts = df.hvplot.scatter(x='x', y='y', datashade=True, by=cat, colormap=cmap, legend=True, frame_width=width, frame_height=height, title=str(title))
return BG_TILES * pts
# +
unknown = df[(df[CATEGORY]=='Unknown value') | (df[CATEGORY]=='Undefined')]
known = df[(df[CATEGORY]!='Unknown value') & (df[CATEGORY]!='Undefined')]
( plot_categorized_scatter(df, CATEGORY, title='Categorized', width=SMSIZE, height=SMSIZE, cmap=cmap) +
plot_basic_scatter(unknown, COLOR_HIGHLIGHT, title=f'Unknown {CATEGORY} only', width=SMSIZE, height=SMSIZE, size=1) +
plot_basic_scatter(known, COLOR_BASE, title=f'Known {CATEGORY} only', width=SMSIZE, height=SMSIZE, size=1)
)
# -
# #### Temporal attribute gaps
#
# This step addresses the question if there are temporary gaps in attribute data. Changes to the data collection or (pre)processing workflow can affect which attributes are available during certain time spans. Temporary attribute gaps may be one-time or reoccurring issues.
#
# Temporary attribute gaps can have similar consequences as spatial attribute gaps. If the attribute gap was caused by a change in the (pre)processing workflow, the damage may be reversible if the original data is still available.
#
# The methods used to explore temporal extent and gaps can be adopted to missing attribute data. Temporal charts, as well as two-dimensional time histograms may be used to find attribute gaps in linear sequence or in temporal cycles.
#
plot_multiples_by_day(unknown, color='red').cols(2)
DATE = date(2017,7,1)
unknown['id'].where(unknown.index.date==DATE).dropna().resample(TIME_SAMPLE).count().hvplot(
title=f'Records per {TIME_SAMPLE} on {DATE}', frame_width=SMSIZE, color='red', frame_height=SMSIZE, ylim=(0,82000), label='unknown'
) * known['id'].where(known.index.date==DATE).dropna().resample(TIME_SAMPLE).count().hvplot(
color='gray', label='known'
)
DATE = date(2018,1,1)
unknown['id'].where(unknown.index.date==DATE).dropna().resample(TIME_SAMPLE).count().hvplot(
title=f'Records per {TIME_SAMPLE} on {DATE}', frame_width=SMSIZE, color='red', frame_height=SMSIZE, ylim=(0,82000), label='unknown'
) * known['id'].where(known.index.date==DATE).dropna().resample(TIME_SAMPLE).count().hvplot(
color='gray', label='known'
)
# ### DATA PREPARATION: Computing segment information
# +
def time_difference(row):
t1 = row['prev_t']
t2 = row['t']
return (t2-t1).total_seconds()
def speed_difference(row):
return row['speed_m/s'] - row['prev_speed']
def acceleration(row):
if row['diff_t_s'] == 0:
return None
return row['diff_speed'] / row['diff_t_s']
def spherical_distance(lon1, lat1, lon2, lat2):
delta_lat = radians(lat2 - lat1)
delta_lon = radians(lon2 - lon1)
a = sin(delta_lat/2) * sin(delta_lat/2) + cos(radians(lat1)) * cos(radians(lat2)) * sin(delta_lon/2) * sin(delta_lon/2)
c = 2 * atan2(sqrt(a), sqrt(1 - a))
dist = R_EARTH * c
return dist
def distance_to_prev(row):
return spherical_distance(row['prev_lon'], row['prev_lat'], row['lon'], row['lat'])
def distance_to_next(row):
return spherical_distance(row['next_lon'], row['next_lat'], row['lon'], row['lat'])
def direction(row):
lon1, lat1, lon2, lat2 = row['prev_lon'], row['prev_lat'], row['lon'], row['lat']
lat1 = radians(lat1)
lat2 = radians(lat2)
delta_lon = radians(lon2 - lon1)
x = sin(delta_lon) * cos(lat2)
y = cos(lat1) * sin(lat2) - (sin(lat1) * cos(lat2) * cos(delta_lon))
initial_bearing = atan2(x, y)
initial_bearing = degrees(initial_bearing)
compass_bearing = (initial_bearing + 360) % 360
return compass_bearing
def angular_difference(row):
diff = abs(row['prev_dir'] - row['dir'])
if diff > 180:
diff = abs(diff - 360)
return diff
def compute_segment_info(df):
df = df.copy()
df['t'] = df.index
df['prev_t'] = df.groupby('id')['t'].shift()
df['diff_t_s'] = df.apply(time_difference, axis=1)
df['prev_lon'] = df.groupby('id')['lon'].shift()
df['prev_lat'] = df.groupby('id')['lat'].shift()
df['prev_x'] = df.groupby('id')['x'].shift()
df['prev_y'] = df.groupby('id')['y'].shift()
df['diff_x'] = df['x'] - df['prev_x']
df['diff_y'] = df['y'] - df['prev_y']
df['next_lon'] = df.groupby('id')['lon'].shift(-1)
df['next_lat'] = df.groupby('id')['lat'].shift(-1)
df['dist_prev_m'] = df.apply(distance_to_prev, axis=1)
df['dist_next_m'] = df.apply(distance_to_next, axis=1)
df['speed_m/s'] = df['dist_prev_m']/df['diff_t_s']
df['prev_speed'] = df.groupby('id')['speed_m/s'].shift()
df['diff_speed'] = df.apply(speed_difference, axis=1)
df['acceleration'] = df.apply(acceleration, axis=1)
df['dir'] = df.apply(direction, axis=1)
df['prev_dir'] = df.groupby('id')['dir'].shift()
df['diff_dir'] = df.apply(angular_difference, axis=1)
df = df.drop(columns=['prev_x', 'prev_y', 'next_lon', 'next_lat', 'prev_speed', 'prev_dir'])
return df
# +
# %%time
try:
segment_df = pd.read_pickle('./segments.pkl')
except:
segment_df = compute_segment_info(cropped_df)
segment_df.to_pickle("./segments.pkl")
# -
easteregg = cropped_df[(cropped_df.id==636092484) | (cropped_df.id==636092478)]
easteregg['id'] = 1
segment_df = segment_df.append(compute_segment_info(easteregg))
# ### A-5. Gaps in trajectories
#
#
# Gaps in movement tracks can be due to technical failure of the tracking device, the mover leaving the observable area, deliberate deactivation of the tracking device, or (pre)processing issues.
#
# Unexpected gaps in tracks can affect derived trajectory measures, particularly by underestimating total length and derived measures, such as speed. Additionally, some regions may be more prone to gaps, for example due to unreliable local coverage. Consequently, there may be a lack of reliable data for these regions.
#
# Line density maps can be used to explore the distribution of gaps in tracks. The key is to only plot long segments, that is connections between consecutive records that exceed a certain length.
#
# +
GAP_MIN = 10000
GAP_MAX = 100000
segment_df['is_gap'] = ( (segment_df['dist_prev_m']>GAP_MIN) & (segment_df['dist_prev_m']<GAP_MAX) ) | ( (segment_df['dist_next_m']>GAP_MIN) & (segment_df['dist_next_m']<GAP_MAX) )
segment_df['id_by_gap'] = segment_df.groupby("id")['is_gap'].transform(lambda x: x.ne(x.shift()).cumsum())
# -
grouped = [df[['x','y']] for name, df in segment_df[segment_df.is_gap].groupby(['id', 'id_by_gap']) ]
path = hv.Path(grouped, kdims=['x','y'])
plot = datashade(path, cmap=COLOR_HIGHLIGHT).opts(frame_height=FIGSIZE[1], frame_width=FIGSIZE[0])
BG_TILES * plot
# ## B. Precision problems
#
# Precision issues in movement data may affect both spatial coordinates as well as timestamps of records. The following protocol steps therefore target issues of excessively truncated coordinates and timestamps.
#
# ### B-1. Coordinate imprecision
#
# This step addresses the question if the coordinates have been truncated excessively. Due to the limited accuracy of conventional GPS, one may argue that here is little benefit to more than five decimal places if coordinates are reported as latitude and longitude. However, coordinate precision should be evaluated irrespective of the number of digits that are provided in the dataset.
#
# Imprecise or truncated coordinates can lead to stair-shaped trajectories, particularly in densely sampled datasets. Similar to GPS noise (which will be discussed in step D-2) these stair-shaped patterns affect derived segment measures, such as length, speed, direction, and acceleration, as well as trajectory measures that aggregate these segment measures.
#
# Histograms of direction values or polar histograms are useful in revealing excessively truncated coordinates which would result in an over-representation of direction values at 45 degree intervals.
#
segment_df['dir'][segment_df.dist_prev_m>0].hvplot.hist(bins=72, title='Histogram of directions')
# ### B-2. Timestamp imprecision
#
# This step addresses the question if timestamps have been truncated excessively. Imprecise timestamps are the result of undue truncation or rounding in the date collection or (pre)processing workflow. Truncation can result in duplicate/multiple position records of the same mover referring to the same time.
#
# Truncated timestamps can result in consecutive records with identical timestamps but different positions. This will result in zero-length time deltas between affected records and thus to division-by-zero errors when computing speeds. If positions are sparsely sampled, moderate truncation (for example, of milliseconds) will not result in multiple records with identical timestamps. However, derived speed values may still suffer from higher noise due to the imprecise representation of time between locations.
#
# Counts of records per timestamp and mover ID can help identify cases of excessively truncated timestamps. Histograms of the number of duplicate timestamps per mover ID show whether this issue affects all movers equally or if there are certain movers where this issue appears more frequently. Additional visualizations than can shed light on the issue of duplicate records include spatial plots (such as point and density maps) as well as temporal plots (such as two-dimensional time histograms) and spatiotemporal space-time cubes.
#
# +
non_zero_movement = segment_df[segment_df.dist_prev_m>0]
n_per_id_t = non_zero_movement[['id', 't', 'x']].groupby(['id', 't']).count().reset_index()
n_per_id_t['x'].plot.hist(title='Counts of records per timestamp and mover ID', log=True)
#n_per_id_t.groupby('x').count().hvplot(title='Counts of records per timestamp and mover ID', y='id', logy=True) # line plot not ideal
#n_per_id_t['x'].hvplot.hist(title='Counts of records per timestamp and mover ID', logy=True) # upstream bug in log scale
# -
duplicates_per_id = n_per_id_t[n_per_id_t.x>1].drop(columns=['t']).groupby(['id']).count().rename(columns={'x':'n'})
duplicates_per_id['n'].plot.hist(title='Count of duplicate timestamps per mover ID', log=True)
# ## C. Consistency problems
#
# Datasets may not be as consistent with regards to collection parameters and type of movers as analysts expect. These problems usually cannot be detected from elementary position records. Therefore, intermediate segments or overall trajectories are needed. The following protocol steps target issues of heterogeneous sampling intervals as well as unexpected heterogeneous mover types and tracker types.
#
# ### C-1. Sampling heterogeneity
#
# This step addresses the question whether the sampling frequency is stable. Some tracking systems provide records at regular time intervals. Other systems have rule-based sampling strategies. For example, in the Automatic Identification System (AIS), updates are more frequent when objects move quickly than when they stand still. In other contexts, GPS trackers may skip positions during straight-line movement. Other systems work on a best-effort base with a target sampling interval that may be exceeded if the system is busy.
#
# Heterogeneous sampling intervals make datasets harder to analyze. Existing methods may expect regularly-sampled input data. Depending on the extent of the heterogeneity, it may be possible to resample the data. Failing that, the methods have to be adjusted to support irregular or mixed sampling intervals.
#
# Histograms of sampling intervals help determine whether sampling intervals are stable and, if yes, what the typical sampling interval is. If not, they show the range of observed sampling intervals.
#
segment_df.diff_t_s.hvplot.hist(title='Histogram of intervals between consecutive records (in seconds)', bins=100)
segment_df[segment_df.diff_t_s<=120].diff_t_s.hvplot.hist(title='Histogram of intervals between consecutive records (in seconds)', bins=60)
# Coordinate change plots can reveal previously mentioned resampling strategies. For example, if the resampling strategy is based on a certain minimum distance between records, there will be a hole in the center of the plot.
segment_df.hvplot.scatter(title='Coordinate change plot', x='diff_x', y='diff_y', datashade=True,
xlim=(-1000,1000), ylim=(-1000,1000), frame_width=FIGSIZE[1], frame_height=FIGSIZE[1])
# ### C-2. Mover heterogeneity
#
# This step addresses the question whether the dataset contains heterogeneous types of movers. Datasets of human movement are expected to contain a mix of different transport modes. Other datasets, such as floating car data (FCD), are expected to be more heterogeneous, for example, to only contain car movements. However, errors in the collection process can invalidate this assumption. For example, if mobile (as opposed to built-in) trackers are used, they may be removed from vehicles and carried around by other means of transport. Other sources of heterogeneity are not due to errors but may still surprise the analysts. For example, AIS datasets also contain track from search and rescue vessels which include helicopters.
#
# An unexpected mix of mover types can affect the validity of analysis results and the performance of derived models. Certain movement statistics may be over or underestimated due to the presence of unexpected mover types. For example, the mean speed along a road may be underestimated if pedestrian tracks are mixed with vehicle tracks. Consequently, a travel time prediction algorithm may predict exaggerated travel times. Therefore, certain movers may have to be removed to ensure the validity of analysis and model results.
#
# Scatter plots of different combinations of trajectory characteristics, such as total length, mean speed, mean direction change, and typical acceleration can help gain a better understanding of how heterogeneous the movers in a dataset are.
#
#
# +
non_zero_speed = segment_df[(segment_df['speed_m/s']>0.1)]
daily = non_zero_speed.groupby(['id', pd.Grouper(freq='d')]).agg({'dist_prev_m':'sum', 'speed_m/s':'median'})
daily.hvplot.scatter(title='Daily travelled distance over median speed (m/s)', x='dist_prev_m', y='speed_m/s',
hover_cols=['id','time'], frame_width=FIGSIZE[1], frame_height=FIGSIZE[1], alpha=0.3,
xlim=(-100000,1500000), ylim=(-10,100))
# -
def plot_paths(original_df, title='', add_bg=True, height=FIGSIZE[1], width=FIGSIZE[0]):
grouped = [df[['x','y']] for name, df in original_df.groupby(['id']) ]
path = hv.Path(grouped, kdims=['x','y'])
plot = datashade(path, cmap=COLOR_HIGHLIGHT).opts(title=title, frame_height=height, frame_width=width)
if add_bg:
return BG_TILES * plot
else:
return plot
speedsters = daily[daily['speed_m/s']>20].reset_index().id.unique()
speedsters = segment_df[segment_df.id.isin(speedsters)]
plot_paths(speedsters, title='Speedsters')
daily.hvplot.scatter(
title='Daily distance over median speed (m/s)', x='dist_prev_m', y='speed_m/s',
hover_cols=['id','time'], frame_width=SMSIZE, frame_height=SMSIZE, alpha=0.3, xlim=(-200000,4500000), ylim=(-10,100)
) + plot_paths(
speedsters, title='Speedsters', height=SMSIZE, width=SMSIZE
)
longdist = daily[daily['dist_prev_m']>800000].reset_index().id.unique()
longdist = segment_df[segment_df.id.isin(longdist)]
plot_paths(longdist, title='Long distance travelers')
# ### DATA PREPARATION: Computing trajectory information
# +
MINIMUM_NUMBER_OF_RECORDS = 100
MINIMUM_SPEED_MS = 1
def reset_values_at_daybreaks(tmp, columns):
tmp['ix'] = tmp.index
tmp['zero'] = 0
ix_first = tmp.groupby(['id', pd.Grouper(freq='d')]).first()['ix']
for col in columns:
tmp[col] = tmp['zero'].where(tmp['ix'].isin(ix_first), tmp[col])
tmp = tmp.drop(columns=['zero', 'ix'])
return tmp
def compute_traj_info(segment_df):
tmp = segment_df.copy()
tmp['acceleration_abs'] = np.abs(tmp['acceleration'])
tmp['diff_speed_abs'] = np.abs(tmp['diff_speed'])
tmp = tmp.replace([np.inf, -np.inf], np.nan)
tmp = reset_values_at_daybreaks(tmp, ['diff_t_s','dist_prev_m','diff_speed_abs','acceleration_abs'])
traj_df = tmp.groupby(['id', pd.Grouper(freq='d')]) \
.agg({'diff_t_s':['median', 'sum'],
'speed_m/s':['median','std'],
'diff_dir':['median','std'],
'dist_prev_m':['median', 'sum'],
'diff_speed_abs':['max'],
'acceleration_abs':['median','max','mean','std'],
't':['min','count'],
'shiptype':lambda x:x.value_counts().index[0]})
traj_df.columns = ["_".join(x) for x in traj_df.columns.ravel()]
traj_df = traj_df.rename(columns={'t_count':'n', 'shiptype_<lambda>':'shiptype',
'diff_t_s_sum':'duration_s', 'dist_prev_m_sum':'length_m'})
traj_df['length_km'] = traj_df['length_m'] / 1000
traj_df['duration_h'] = traj_df['duration_s'] / 3600
traj_df['t_min_h'] = traj_df['t_min'].dt.hour + traj_df['t_min'].dt.minute / 60
traj_df = traj_df[traj_df.n>=MINIMUM_NUMBER_OF_RECORDS]
traj_df = traj_df[traj_df['speed_m/s_median']>=MINIMUM_SPEED_MS]
return traj_df
# +
# %%time
try:
traj_df = pd.read_pickle('./traj.pkl')
except:
traj_df = compute_traj_info(segment_df)
traj_df.to_pickle("./traj.pkl")
traj_df
# -
hvplot.scatter_matrix(
traj_df[['length_km', 'speed_m/s_median', 'duration_h', 'acceleration_abs_mean', 'diff_dir_median']]
)
# ### C-3. Tracker heterogeneity
#
#
# This step addresses the question whether the dataset contains records from devices with different tracking characteristics. Devices with GPS tracking capabilities vary widely in performance. For example, when data is collected using smartphone apps, coordinates may have passed through a variety of (not always fully transparent) preprocessing steps that depend on the operating system version and hardware manufacturer.
#
# Unexpected heterogeneity of tracker characteristics can affect the performance of analysis tasks. For example, if a pattern matcher was trained using high-frequency data with regular sampling intervals, it may not perform as expected when inputs change to irregular sampling intervals or lower frequency. If some trackers exhibit higher GPS noise than other trackers in the dataset, the higher-noise trajectories will suffer an overestimation of distance and derived measures (for more details on noise see step D-2).
#
# Effects of heterogeneous trackers can be hard to distinguish from effects of heterogeneous movers. Tracker heterogeneity may result in sampling rates and/or spatial accuracy that differ between movers. Furthermore, differences in the availability of additional attribute data within movement records may point towards tracker heterogeneity.
#
#
traj_df[(traj_df['diff_t_s_median']<=120) & (traj_df['speed_m/s_median']>0)] \
.hvplot.scatter(
title='Median sampling interval over median speed', alpha=0.3,
x='diff_t_s_median', y='speed_m/s_median', hover_cols=['id','time'],
frame_width=FIGSIZE[1], frame_height=FIGSIZE[1], ylim=(-10,100))
# ## D. Accuracy problems
#
# Incorrect mover identities, coordinates, and timestamps can affect movement data analyses in a variety of ways. These problems usually cannot be detected from elementary position records. Therefore, intermediate segments or overall trajectories are needed. The following protocol steps target issues of mover identity, as well as spatial and temporal inaccuracy.
#
#
# ### D-1. Mover identity issues
#
# Reliable mover identifiers are needed to identify which movement data records belong to the same mover. Identity issues occur when IDs are not unique, i.e. if multiple movers are assigned the same identifier. A single mover may also be referred to by multiple different identifiers, either at the same time or due to changes over time. This can happen because the data collection system or (pre)processing workflow reassigns identifiers based on business rules or in regular time intervals.
#
# #### Non-unique IDs
#
# This step addresses the question whether the dataset contains cases of non-unique identifiers. Due to misconfiguration of trackers or (pre)processing errors, the same identifier may be assigned to multiple movers simultaneously (or to different movers over time which is covered by the unstable IDs step).
#
# Simultaneous non-unique IDs result in trajectories that connect location records by multiple movers traveling on their distinct paths. The resulting trajectory therefore jumps between locations along these different paths. Consequently, the trajectory assumes a zigzag shape and speeds derived from consecutive location records assume unrealistic values. Non-unique IDs can make it impossible to reliably distinguish affected movers. In some settings, it may be possible to salvage records if they include sufficient other information that can be used to infer identity, for example, mover names or mover properties (such as size, type or color).
#
# Scatter plots of trajectory length and direction change are useful to identify cases of non-unique IDs. Assuming that movers with identical IDs do not typically travel in close vicinity, potential candidates for non-unique IDs are characterized by long trajectories with high direction change values.
#
traj_df.hvplot.scatter(
title='Trajectory length over direction difference (median)', alpha=0.3,
x='length_km', y='diff_dir_median', hover_cols=['id','time'],
frame_width=FIGSIZE[1], frame_height=250#,
) + traj_df.sort_values(by='length_km', ascending=False)[:10][['length_km', 'speed_m/s_median', 'diff_dir_median']].hvplot.table(
title='Top 10 trajectories - length', frame_width=FIGSIZE[1])
plot_single_mover(segment_df, 1, date(2017,7,1))
traj_df = traj_df.drop(1, level='id')
# #### Unstable IDs
#
# This step addresses the question whether mover identifiers are stable and for how long they remain stable. Some data sources do not provide permanently stable identifiers. Systems may reassign identifiers based on business rules or in regular time intervals. For example, taxi floating car systems may not include stable vehicle IDs, instead relying on trip IDs that are reassigned whenever a taxi finishes a trip.
#
# Unstable IDs can limit the analysis potential of the dataset. For example, if trip IDs keep changing and there is no stable mover ID, it becomes difficult to reliably determine mover statistics, such as the daily number of trips or the total distance moved.
#
# Scatter matrixes of trajectory duration versus start time (a combination of scatter plots and histograms) are useful to find out how often IDs change and whether they tend to change at the same time. For example, if IDs change daily and at the same time, the histograms will exhibit spikes at the 24 hour duration and corresponding time of day. In contrast, if IDs are stable over the whole observation period, the histograms will reflect the time spans during which individual movers were tracked.
#
hvplot.scatter_matrix(traj_df[['t_min_h', 'duration_h']])
# ### D-2. Spatial inaccuracy
#
# Coordinate errors range from basic noise due to the inherent inaccuracy of GPS to large jumps caused by technical errors or deliberate action.
#
#
# #### Outliers with unrealistic jumps
#
#
# This step addresses the question if trajectories contain large erroneous jumps that result in unrealistic derived speed values and require data cleaning. The limit for being unrealistic depends on the use case. For example, for ground-based transport, Fillekes et al. (2019) set the limit at 330 km/h based on the maximum speed of German high-speed trains.
#
# Large jumps affect derived segment measures, including direction, length, speed, and acceleration. Consequently they also affect trajectory measures which are aggregates of these segment measures. However, single jumps may not be immediately recognizable when looking at trajectory measures only.
#
# Histograms of derived speed between consecutive location records are useful to see if there is a long tail of high speed values.
#
segment_df['speed_m/s'].hvplot.hist(
title='Histogram of speed between consecutive records', bins=100, frame_width=FIGSIZE[1], frame_height=250
) + segment_df.sort_values(by='speed_m/s', ascending=False)[:10][['id', 'speed_m/s']].hvplot.table(
title='Top 10 records - speed', frame_width=FIGSIZE[1])
plot_single_mover(segment_df, 218057000, date(2018,1,1))
plot_single_mover(segment_df, 219348000, date(2017,7,1))
# #### Jitter / noise
#
# This step addresses the question of how noisy the trajectories are. GPS tracks are inherently noisy. However, in some cases, what appears as excessive noise can reflect real movement patterns. For example, vessel routes may have a zig-zag shape in case of adverse weather conditions. Noise also affects trajectories of movers that are standing still, appearing as fake jittery movement.
#
# Jitter or noise causes a systematic "overestimation of distance" when the sampling frequency is high. On the other hand, distances are underestimated when the sampling frequency is low. Without evaluating the sampling frequency, distance and derived speed values therefore are insufficient to understand noise.
#
# Scatter plots of direction change that compare median change and the observed standard deviation of change values can provide insights into the presence of excessive jitter or noise. However, high median values (approaching 180°) indicate out-of-sequence rather than jitter issues and will be discussed in the next step.
#
traj_df.hvplot.scatter(
title='Direction difference median over standard deviation', alpha=0.3,
x='diff_dir_median', y='diff_dir_std', hover_cols=['id','time'], #datashade=True,
frame_width=FIGSIZE[1], frame_height=250, ylim=(-10,100)
) + traj_df.sort_values(by='diff_dir_median', ascending=False)[:10][['diff_dir_median','diff_dir_std']].hvplot.table(
title='Top 10 trajectories - direction difference', frame_width=FIGSIZE[1])
plot_single_mover(segment_df, 244063000, date(2018,1,1))
plot_single_mover(segment_df, 220614000, date(2018,1,1))
# ### D-3. Temporal inaccuracy
#
# Timestamp errors potentially affect the synchronization between trajectories as well as the order of records within individual trajectories.
#
# #### Time zone and daylight saving issues
#
# This step addresses the question how time zones and daylight saving affect the dataset. In some datasets, time zone information may be included with each time stamp. However usually, this is not the case and analysts have to resort to metadata or documentation which are not always comprehensive or reliable. Time zone issues can be hard to detect, particularly if the dataset contains tracks from multiple time zones but the zone information got lost along the way. These issues may be discovered due to unexpected derived movement patterns, such as, for example, significant numbers of people leaving their homes in the middle of the night or excessive movement of nocturnal animals during the day. However, small shifts, such as missing daylight savings information, can be hard to distinguish from the normal variation.
#
# Unresolved time zone and daylight saving issues can have diverse side effects. For example, two tracks that appear to be moving together (collocated movement) may actually represent movement that happened at different points in time. Depending on the analysis area, a mix of local time and UTC (Coordinated Universal Time) could result in spikes of activity at unexpected times of the day.
#
# Temporal charts of record counts are helpful to detect gaps or double counting at the date and time when daylight saving goes into and out of effect. Two-dimensional time histograms of movement properties, such as speed, can help recognize time zone issues by revealing unusual temporal patterns.
#
tmp = segment_df[segment_df['speed_m/s']>1]
hourly = tmp['id'].groupby([tmp.index.hour, pd.Grouper(freq='d')]).count().to_frame(name='n')
hourly.rename_axis(['hour', 'day'], inplace=True)
hourly.hvplot.heatmap(title='Count of records with speed > 1m/s', x='hour', y='day', C='n', width=FIGSIZE[0])
# #### Out-of-sequence positions
#
# This step addresses the question if records belonging to a trajectory appear out of sequence. A closely related problem is when a mover appears at two different locations at the same time. These problems can happen in systems that do not provide tracker timestamps and instead use receiver or storage time. For example, the Automatic Identification System (AIS) protocol does not transmit tracker timestamps and instead provides only offsets (in second) from the previously transmitted message which is insufficient to establish temporal order "since positional updates from a single vessel may come from a series of base stations (those within range of its antenna along the route)" (Patroumpas et al. 2017).
#
# Out-of-sequence positions affect many derived trajectory measures, including direction, length, speed, and acceleration. In affected trajectories, these measures will be severely overestimated.
#
# Scatter plots of direction change and speed are helpful to detect out-of-sequence problems. The sudden reversals of movement direction result in high direction change values (approaching 180°) and high speeds.
#
traj_df.hvplot.scatter(
title='Direction difference (median) over speed (median)', alpha=0.3,
x='diff_dir_median', y='speed_m/s_median', hover_cols=['id','time'], #datashade=True,
frame_width=FIGSIZE[1], frame_height=250#, ylim=(-10,100)
) + traj_df.sort_values(by='diff_dir_median', ascending=False)[:10][['diff_dir_median','diff_dir_std','speed_m/s_median']].hvplot.table(
title='Top 10 trajectories - direction difference', frame_width=FIGSIZE[1])
plot_single_mover(segment_df, 308322000, date(2017,7,1))
plot_single_mover(segment_df, 265615040, date(2017,7,1))
| protocol.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Aula 06
# <NAME>
# 816118386
# 1-Defina a função soma_nat que recebe como argumento um número natural n e devolve a soma de todos os números naturais até n.
#
# Ex: soma_nat(5) = 15
def soma_nat (n):
return 1 if n ==1 else n + soma_nat(n-1)
print(soma_nat(2), soma_nat(1), soma_nat(5))
# 2- Defina a função div que recebe como argumentos dois números naturais m e n e devolve o resultado da divisão inteira de m por n. Neste exercício você não pode recorrer às operações aritmétcas de multplicação, divisão e resto da divisão inteira.
#
# Ex: div(7,2) = 3
def div(dividendo, divisor):
return 0 if dividendo < divisor else 1 + div(dividendo - divisor, divisor)
print(div(5,2))
print(div(2,3))
# 3-Defina a função prim_alg que recebe como argumento um número natural e devolve o primeiro algarismo (o mais signifcatvo) na representação decimal de n.
#
# Ex: prim_alg(5649) = 5 Ex: prim_alg(7) = 7
# +
def prim_alg(numero):
if numero < 0:
return None
else:
return str(numero)[:1]
assert prim_alg(5565), 'Informe um número natural >= a zero'
print(prim_alg(5565))
# -
#
# 4-Defina a função prod_lista que recebe como argumento uma lista de inteiros e devolve o produto dos seus elementos.
# +
def prod_lista(lista):
return reduce (lambda x, y: x * y, lista)
assert prod_lista([1,2,3,4,5,6]), 'Algo deu errado'
print(prod_lista([1,2,3,4,5,6]))
# -
#
# 5-Defina a função contem_parQ que recebe como argumento uma lista de números inteiros $w$ e devolve True se $w$ contém um número par e False em caso contrário.
# +
def contem_parQ(w):
if w[0] % 2 == 0:
return True
else:
if len(w) > 1:
contem_parQ (w + 1)
return False
assert contem_parQ([2,3,1,2,3,4]), 'Não há números pares na lista '
print(contem_parQ([2,3,1,2,3,4]))
# -
# 6- Defina a função todos_imparesQ que recebe como argumento uma lista de números inteiros w e devolve True se w contém apenas números ímpares e False em caso contrário.
# +
def todos_imparesQ(w):
return True if len(w) == 0 or (w[len(w) - 1] % 2 != 0 and todos_imparesQ(w[:-1])) else False
assert(todos_imparesQ([1,1,3,7,11]) == True)
assert(todos_imparesQ([1,2,3,4,5]) == False)
assert(todos_imparesQ([2,4,6,8,10]) == False)
print(todos_imparesQ([1,1,3,7,11]), todos_imparesQ([1,2,3,4,5]), todos_imparesQ([2,4,6,8,10]))
# -
#
# 7-Defina a função pertenceQ que recebe como argumentos uma lista de números inteiros w e um número inteiro n e devolve True se n ocorre em w e False em caso contrário.
def pertenceQ(w,n):
return True if len(w)> 0 and (n in w or pertenceQ(w[:-1],n)) else False
print('Teste da função:')
testes = [0,1,2,3,5,8,13]
print ('Lista: ' , testes)
for teste in range(8):
if(pertenceQ(testes,teste) == False):
print(teste, ' não ocorre na lista')
else:
print(teste, ' ocorre na lista')
# 8-Defina a função junta que recebe como argumentos duas listas de números inteiros w1 e w2 e devolve a concatenação de w1 com w2 .
# Ex: junta([1,2,3],[4,5,6]) = [1, 2, 3, 4, 5, 6] Ex: junta([],[4,5,6]) = [4, 5, 6] Ex: junta([1,2,3],[]) = [1, 2, 3]
# 9-Defina a função temPrimoQ que recebe como argumento uma lista de listas de números inteiros w e devolve True se alguma das sublistas w tem um número primo e False em caso contrário
# 1- Defina a função inverteLista que recebe como argumento uma lista w e devolve a mesma lista mas invertda.
# Ex: inverteLista([1,2,3,4,5]) = [5, 4, 3, 2, 1] Ex: inverteLista([])
| Exercicio06.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#Python Basics
#Objects and Classes
#In this ttutorial, several examples to demonstrate objects and classes in python have been presented
#Python has many different kinds of DATA TYPEs
#Data types: int: 3,5, 6, 9, float: 0.089, 12.5496, 2.008, string: Behdad, Mohammad, list: ["BMJ", 189, 2003, 0.00675]
#tuples: (12422, "Python", "Classes"), dictionary: {"key1": val1, "key2": val2, "key3": val3}
# boolean: True, False
#In python each of the mentioned type of data is an object
#Each object has: Type, an internal data representation (a blueprint), a sets interacting with the data (methods)
#An object is an instance (SHEIY) of a particular type
#We can find the type of an object by using the command" type ()"
#A "class" or "method" are functions that every instance of that class or types provides
#example
#Sorting is an example of a method that interacts with the data in the object
Rating=[12, 67, 92, 46, 18, 9, 0, 3, 232, 11]
print("Rating, before method:", Rating)
#here rating is an object which is connected to method, namely sort with ()
Rating.sort()
print("Rating, after method:", Rating)
Rating
#example
#Reversing is an example of a method that interacts with the data in the object
INV=[12, 67, 92, 46, 18, 9, 0, 3, 232, 11]
print("INV, after method:", INV)
#here INV is an object which is connected to method, namely sort with ()
INV.reverse()
print("INV, after method:", INV)
INV
#So, Object is a type of data and class or method is an operator can create change on object : "object.method()"
# +
#You can creat your own type or class (method) in Python
#Defining Classes
#The class has data attributes. The class has methods
#We then create an instances or instances of that class or objects
#We define our class. We then initialize each instance of the class
# First definition: class NameOfClass(object or class parent):
class Circle2(object):
#Initialization of class:
#def __init__(Attribute1,Attribute2, Attribute3, ...)
def __init__(self, radius1, color):
self.radius1=radius1;
self.color=color;
#the function "__init__" is a constructor
#The ‘radius’ and ‘color’ are parameters (Attributes)
#The ’self’ parameter refers to the newly created instance of the class
#We can set the value of the radius and color data attributes
#to the values passed to the constructor method by self.radius(); self.color();
#SELF:
#It is helpful to think of self as a box
#that this box contains all the data attributes of the object
#Similarly, we can define the class rectangle
class Rectangle(object):
def __init__(self, hight, width ,color):
self.hight=hight;
self.width=width;
self.color=color;
#After we have created the class, in order to create an object of class circle
#we introduce a variable; this will be the name of the object
# We create the object by using the object constructor
#When we create a Circle object we call the code like a function
C1=Circle2(20, "Green")
#C1: the name of object
#Circle: Name of the class
#(10, "red): Data Attributes (Parameters)
#Circle(10, "red"): Object Constructor
#Typing the object's name followed by a dot and the data attribute name
# gives us the data
C1.radius1
#or
print(C1.radius1)
C2=Circle2(45, "Pink")
C2.color
#or
print(C2.color)
# +
#exaple 1
class CLASS1(object):
#Initialization of class:
#def __init__(Attribute1,Attribute2, Attribute3, ...)
def __init__(self, obj1, objSt2, obj3, obj4):
self.obj1=(obj1*2)+10+obj4;
self.objSt2=(objSt2+" AI"+"ANNs")*obj4;
self.obj3=obj3;
self.obj4=obj4;
#the function "__init__" is a constructor
v1=CLASS1(5,"Methods:", "GOOD", 15)
print(v1.obj1)
print(v1.objSt2)
print(v1.obj3)
print(v1.obj4)
#example 2
class RobotControl(object):
#Initialization of class:
#def __init__(Attribute1,Attribute2, Attribute3, ...)
def __init__(self, obj1, obj2, obj3, obj4):
aws=obj1/obj4+19.5
sw=256/obj2
self.obj1=(obj1*2)+90+obj4+aws;
self.obj2=(obj2+sw)*obj4;
self.obj3=obj3;
self.obj4=obj4;
#the function "__init__" is a constructor
v1=RobotControl(5,11,67, 15)
print(v1.obj1)
print(v1.obj2)
print(v1.obj3)
print(v1.obj4)
#example 2
class PrintingStorage(object):
#Initialization of class:
#def __init__(Attribute1,Attribute2, Attribute3, ...)
def __init__(self, obj1):
m1="It is full"
m2="It needs to be recounted"
m3="It is ok"
if obj1>12:
print(m1)
self.obj1=obj1/2;
else:
self.obj1=obj1*2;
print(m3)
#the function "__init__" is a constructor
v1=PrintingStorage(10)
print(v1.obj1)
# +
#Setting or Changing the data attribute
#In Python, we can also set or change the data attribute directly
class PrintingStorage(object):
def __init__(self, obj1):
m1="It is full"
m2="It needs to be recounted"
m3="It is ok"
if (obj1>20):
print(m1)
self.obj1=obj1/2;
print("self.obj1 is: ", self.obj1)
elif (obj1<=220)and(obj1>10):
print(m2)
self.obj1=obj1;
print("self.obj1 is: ", self.obj1)
else:
self.obj1=obj1*1.5;
print(m3)
print("self.obj1 is: ", self.obj1)
#Typing the object's name, followed by a dot and the data attribute name,
#and set it equal to the corresponding value
PS1=PrintingStorage(30)
PS1.obj1=50
print(PS1.obj1)
# +
#Methods
#Methods are functions that interact and change the data attributes, changing or using the data attributes of the object
class Rectangle(object):
#We can add default values to the parameters
#If you indicate any value, that value is on priority than the default one
def __init__(self, hight=14, width=7 ,color="green"):
self.hight=hight;
self.width=width;
self.color=color;
#Let's say we would like to change the size of a rectangle
#this involves changing the hight, width attribute
#We add a method "add Dimensions" to the class Rectangle
#IT IS SO IMPORTANT
#IT IS SO IMPORTANT
#When you use a object from the class you should write ((self.object)) in the method
def Add_Dim(self,h, w):
#When you use a object from the class you should write ((self.object)) in the method
self.hight=self.hight+h
#When you use a object from the class you should write ((self.object)) in the method
self.width=self.width+w
return(self.hight, self.width)
#As before, we create an object with the object constructor
Rec=Rectangle(12,25,"Yellow")
print(Rec.hight)
Rec.Add_Dim(7, 3)
# +
#More examples
class Car(object):
def __init__(self,make,model,color):
self.make=make;
self.model=model;
self.color=color;
self.owner_number=0
def car_info(self):
print("make: ",self.make)
print("model:", self.model)
print("color:",self.color)
print("number of owners:",self.owner_number)
def sell(self):
self.owner_number=self.owner_number+1
BMW1=Car("BMW", "X6", "Blue")
print(BMW1.car_info())
#print(BMW1.sell())
| PythonBasics-Objects&Classes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Session 4: Learning Curves, Regularization and Cross Validation
#
# ## House sale-value prediction using the Boston housing dataset
#
# ------------------------------------------------------
# *Introduction to Data Science & Machine Learning*
#
# *<NAME> <EMAIL>*
#
# ------------------------------------------------------
# ## Importing Packages
# +
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
# %matplotlib inline
# -
# Today, we will continue with the example we used for session 2: Predicting house values using the average number of rooms in the [Boston housing dataset](https://archive.ics.uci.edu/ml/machine-learning-databases/housing/housing.names).
#
# Attribute Information can be found [here](https://www.kaggle.com/c/boston-housing).
#
# ### Loading Data
#
# We will manage the database using [Panda's library and Dataframes](http://pandas.pydata.org/pandas-docs/stable/tutorials.html)
# +
housing_data=pd.read_csv('./Boston_train.csv')
housing_data.head(10)
# -
# We divide the whole data set in **80% training** and **20% test**:
# +
N = len(housing_data)
N_train = np.round(0.8 * N).astype(np.int32)
np.random.seed(seed=10) #To fix the random seed
mask = np.random.permutation(len(housing_data))
regression_data_frame = housing_data[['rm','medv']].iloc[list(mask[0:N_train])]
X_0 = np.array(regression_data_frame['rm'])
Y = np.array(regression_data_frame['medv'])
regression_data_frame_test = housing_data[['rm','medv']].iloc[list(mask[N_train:-1])]
X_0_test = np.array(regression_data_frame_test['rm'])
Y_test = np.array(regression_data_frame_test['medv'])
# -
# # Learning curves
#
# Learning curves are a very powerfull and informative methodology to understand if your regression model is exposed to **overfitting** or is too simple, i.e., it is **too biased**.
#
# To plot the learning curves for a regression model with $N$ training points and $Ntest$ test points, we plot the error and test averagre MSE for different **subset sizes of training points**. More precisely, for $N_0\leq N$, we compute:
# $\boldsymbol{\theta}^*$
# $$J_{train}(\boldsymbol{\theta}^*,N_0) = \frac{1}{N_0} \sum_{i=1}^{N_0} (y^{(i)}-(\boldsymbol{\theta}^*)^T\overline{\mathbf{x}}^{(i)})^2$$
# and
# $$J_{test}(\boldsymbol{\theta}^*) = \frac{1}{N_{Test}} \sum_{i=1}^{N_{Test}} (y^{(i)}-(\boldsymbol{\theta}^*)^T\tilde{\mathbf{x}}^{(i)})^2$$
# where $\boldsymbol{\theta}^*$ is the LS solution for a subset of training points of size $N_0\leq N$. In general, both quantities above **must** be averaged for different subsets.
#
# Lets plot the learning curves for different regression models to understand what they mean.
# +
# A function that normalizes data with pre-scecified mean and std.
def normalize(X,mu=0,std=1,flag_train=True):
if(flag_train):
mu = np.mean(X,0)
std = np.std(X,0)
X -= mu
X /= std
return X,mu,std
# A function to add the all-ones column
def add_interfit(X):
col_ones = np.ones([X.shape[0],1])
return np.concatenate([col_ones,X],1)
# A function to create the normalized function with polynomial features up to degree deg_max
def create_feature_matrix(X_0,deg_max,mu=0,std=1,flag_train=True):
X = np.zeros([X_0.shape[0],deg_max])
deg=1
while deg<=deg_max:
X[:,deg-1] = X_0**deg
deg += 1
X,train_mean,train_std = normalize(X,mu,std,flag_train)
X = add_interfit(X)
return X,train_mean,train_std
# A function to compute the LS solution
def LS_solution(X,Y):
return (np.linalg.pinv(X) @ Y)
# A function to evaluate the LS solution
def LS_evaluate(X,T):
return (X @ T.transpose())
def J_error(Y,Y_est):
return np.mean((Y-Y_est)**2)
# A function to compute the test, training data for given data inputos and for a list of polynomial degrees
def eval_J(X_train,X_test,deg_list,Y_train,Y_test,N):
# Lets compute the normalized feature matrices F_train, F_test
F_train,train_mean,train_std = create_feature_matrix(X_train,np.max(deg_list),0,1,flag_train=True)
F_test,_,_ = create_feature_matrix(X_test,np.max(deg_list),train_mean,train_std,flag_train=False)
J_train = []
T = []
J_test = []
for d in deg_list:
#We train with N random points (this is useful to plot learning curves)
mask = np.random.permutation(X_train.shape[0])
T.append(LS_solution(F_train[mask[:N],0:d+1],Y_train[mask[:N]]))
J_train.append(J_error(Y_train[mask[:N]],LS_evaluate(F_train[mask[:N],0:d+1],T[-1])))
J_test.append(J_error(Y_test,LS_evaluate(F_test[:,0:d+1],T[-1])))
return J_train,T,J_test
# A function to evaluate the learning curves for a given polynomial degree
def eval_learning_curves(X_train,X_test,degree,Y_train,Y_test,N_list,num_subsets):
J_train = []
J_test = []
for n in N_list:
aux = np.zeros([num_subsets,])
aux_t = np.zeros([num_subsets,])
for p in range(num_subsets):
J,_,Jt = eval_J(X_train,X_test,[degree],Y_train,Y_test,n)
#Note that eval_J returns lists
aux[p] = J[0]
aux_t[p] = Jt[0]
J_train.append(np.mean(aux))
J_test.append(np.mean(aux_t))
return J_train,J_test
# -
# ## Model 1: An example of bias
#
# Imagine we use a polynomial of degree 1, i.e., a straight line to estimate the house value from the number of rooms. Lets compute and plot the learning curves.
# +
N_list = np.arange(5,N_train,20)
num_subsets = 20 #With how many random subset we estimate J_train, J_test
degree = 1
J_train_1,J_test_1 = eval_learning_curves(X_0,X_0_test,degree,Y,Y_test,N_list,num_subsets)
# +
plt.semilogy(N_list,J_train_1,'b-o',ms=10,label='J_train')
plt.semilogy(N_list,J_test_1,'b--<',ms=10,label='J_test')
plt.xlabel('Training points')
plt.ylabel('Mean Squared Error (MSE)')
plt.title('Model polynomial order regression = %d' %(degree))
plt.legend()
plt.rcParams["figure.figsize"] = [8,8]
# -
# What do you conclude from the plot above? How does the model improve as we increase the number of training points?
# ## Model 2: An example of mild overfitting
#
# Imagine now we use a polynomial of degree 5 to estimate the house value from the number of rooms. Lets compute and plot the learning curves.
# +
degree = 5
J_train_5,J_test_5 = eval_learning_curves(X_0,X_0_test,degree,Y,Y_test,N_list,num_subsets)
# +
plt.semilogy(N_list,J_train_5,'r-o',ms=10,label='J_train' + ' Pol. 5')
plt.semilogy(N_list,J_test_5,'r--<',ms=10,label='J_test' + ' Pol. 5')
plt.semilogy(N_list,J_train_1,'m-o',ms=10,label='J_train' + ' Pol. 1')
plt.semilogy(N_list,J_test_1,'m--<',ms=10,label='J_test' + ' Pol. 1')
plt.xlabel('Training points')
plt.ylabel('Mean Squared Error (MSE)')
plt.title('Model polynomial order regression = %d' %(degree))
plt.legend()
plt.ylim([1e0,1e03])
# -
# What do you conclude from the plot above? How does the model improve as we increase the number of training points?
# ## Model 3: An example of extreme overfitting
#
# Imagine now we use a polynomial of degree 15 to estimate the house value from the number of rooms. Lets compute and plot the learning curves.
degree = 10
J_train_10,J_test_10 = eval_learning_curves(X_0,X_0_test,degree,Y,Y_test,N_list,num_subsets)
# +
plt.semilogy(N_list,J_train_10,'g-o',ms=10,label='J_train' + ' Pol. 15')
plt.semilogy(N_list,J_test_10,'g--<',ms=10,label='J_test' + ' Pol. 15')
plt.semilogy(N_list,J_train_5,'r-o',ms=10,label='J_train' + ' Pol. 5')
plt.semilogy(N_list,J_test_5,'r--<',ms=10,label='J_test' + ' Pol. 5')
plt.semilogy(N_list,J_train_1,'m-o',ms=10,label='J_train' + ' Pol. 1')
plt.semilogy(N_list,J_test_1,'m--<',ms=10,label='J_test' + ' Pol. 1')
plt.xlabel('Training points')
plt.ylabel('Mean Squared Error (MSE)')
plt.title('Model polynomial order regression = %d' %(degree))
plt.legend()
plt.ylim([1e0,200])
# -
# # Model regularization
#
# We have seen that complex models tend to **overfit** the data **unless we are able to increase the number of training points**, which is many times not an option.
#
# As we show in Lesson 2, one of the properties of overfitted linear regression models is that they tend to show regression coefficients with very large absolute values.
#
# In the next plot we show the regression coefficients for different polynomial interpolation degrees in our running example:
deg_list = [3,5,10]
_,T_list,_ = eval_J(X_0,X_0_test,deg_list,Y,Y_test,N_train)
# +
f, axes = plt.subplots(1, len(deg_list))
for i,t in enumerate(T_list):
axes[i].stem(t)
axes[i].set_title("Degree = %d" %(deg_list[i]))
plt.rcParams["figure.figsize"] = [20,8]
# -
# In model regularization, we combine the flexibility of a **complex model** with a **penalization function** that prevents the model to converge to solutions with very large coefficients.
#
# The easiest regularization example is the $L_2$ penalization or ** Ridge regression**:
#
# $$\boldsymbol{\theta}_\lambda = \arg \min_{\theta} \frac{1}{N} \left[\sum_{i=1}^{N} (y^{(i)}-\boldsymbol{\theta}^T\mathbf{x}^{(i)})^2 + \lambda \sum_{j=1}^{D+1} \theta_j^2\right],$$
#
# where note we do not penalize large values of the intercept $\theta_0$, since it essentially captures the mean of $y$. Also, $\lambda$ is called the **regularization parameter** and it is another parameter that **we have to learn from our data**, looking for right balance between **explaining our training set** and **the penalization term**.
#
# Fortunately, the $L2$ penalization norm does not change the convexity of the loss function and, indeed, we still have a **closed-form** solution
#
# $$\boldsymbol{\theta}_\lambda = (\mathbf{X}^T\mathbf{X} + \mathbf{D}_\lambda)^{-1}\mathbf{X}^T\mathbf{y},$$
#
# where
#
# $$ \mathbf{D} = \left[ \begin{array}{cc} 0 & 0 \\ 0 & \lambda \mathbf{I}_{D\times D}\end{array}\right] $$
#
# Note that the diagonal matrix $\mathbf{D}_\lambda$ ensures in general that the matrix $\mathbf{X}^T\mathbf{X} + \lambda \mathbf{I}$ is invertible, leading to much better condition solutions.
#
# ___
# ## Simple Cross Validation
#
# In order to fit $\lambda$, we will split the training set once more, to create the **validation set**:
#
# <img src="validation_set.png">
#
# The validation set will be used to choose the best $\lambda$ among the same model with different $\lambda$ values, all trained using the training set. This process is known as **cross validation**.
#
# -----
#
# ** Cross Validation Steps **:
#
# 1) Select a regression model and construct the normalized feature matrix for both the training, validation and test sets.
#
# 2) Set a grid of $\lambda$ values to test (i.e. $0.001, 0.003,0.01,0.03,0.1,0.3,1,3,10,....100$)
#
# 3) Compute the Ridge regression solution $\boldsymbol{\theta}_\lambda$ for each $\lambda$ value **using only the training set**
#
#
# 4) Compute the training and validation MSE for each solution (**without the penalization**):
#
# $$J_{train}(\boldsymbol{\theta}_\lambda) = \frac{1}{N} \sum_{i=1}^{N} (y^{(i)}-(\boldsymbol{\theta}_\lambda)^T\overline{\mathbf{x}}^{(i)})^2$$
#
# $$J_{train}(\boldsymbol{\theta}_\lambda) = \frac{1}{N_{val}} \sum_{i=1}^{N_{val}} (y^{(i)}-(\boldsymbol{\theta}_\lambda)^T\hat{\mathbf{x}}^{(i)})^2$$
#
# where $\hat{\mathbf{x}}^{(i)})$ is the $i$-th normalized feature verctor of the validation set. Recall, both the validation and test sets are normalized using the train set statistics.
#
# 5) Select $\lambda^*$ that minimizes the validation error (or refine the search for a narrower set of $\lambda$ values)
#
# 6) Retrain the model for $\lambda^*$ using **both the training and validation sets**
#
# 7) Compute the train and test MSE:
#
# $$J_{train}(\boldsymbol{\theta}_{\lambda^*}) = \frac{1}{N+N_{val}} \sum_{i=1}^{N+N_{val}} (y^{(i)}-(\boldsymbol{\theta}_{\lambda^*})^T\overline{\mathbf{x}}^{(i)})^2$$
#
# $$J_{test}(\boldsymbol{\theta}_{\lambda^*}) = \frac{1}{N_{test}} \sum_{i=1}^{N_{test}} (y^{(i)}-(\boldsymbol{\theta}_{\lambda^*})^T\hat{\mathbf{x}}^{(i)})^2$$
#
# Compare different regression models (i.e. different polynomials) using the test MSE.
#
# ---
#
# Unless the dataset is large enough (and so does the validation set), the robustnetss of the above procedure is improved by averaging the training and validation MSE step 4) over multiple random partitions of the validation/training set (**Repeated random sub-sampling validation**) or splits the training set in K-folds and averages the results using one of the folds as validation set at a time (**K-fold cross validation**). See the [Wikipedia entry](https://en.wikipedia.org/wiki/Cross-validation) about CV for more details.
#
#
# ### Goals for today
#
# Using the functions provided in the beginning of the notebook as guide, create new functions to implement ridge regression and cross validation. Try to observe the effect of regularization in a complex model that is very exposed to overfitting.
# +
# A function to compute the LS solution
def Ridge_solution(X,Y,l):
A = l*np.eye(X.shape[1])
A[0,0] = 0
A += X.transpose() @ X
return (np.linalg.inv(A) @ X.transpose() @ Y)
# A function to compute the test, training data for given data inputos and for a list of polynomial degrees
def eval_J_Ridge(X_train,Xvt,deg_list,Y_train,Yvt,N,lambda_list):
# Xvt,Yvt --> We use this function to evaluate either validation error or test error
# Lets compute the normalized feature matrices F_train, F_test
F_train,train_mean,train_std = create_feature_matrix(X_train,np.max(deg_list),0,1,flag_train=True)
F_vt,_,_ = create_feature_matrix(Xvt,np.max(deg_list),train_mean,train_std,flag_train=False)
J_train = []
T = []
J_vt = []
for d in deg_list:
J_train_d = []
T_d = []
J_vt_d = []
#We train with N random points (this is useful to plot learning curves)
mask = np.random.permutation(X_train.shape[0])
for l in lambda_list:
T_d.append(Ridge_solution(F_train[mask[:N],0:d+1],Y_train[mask[:N]],l))
J_train_d.append(J_error(Y_train[mask[:N]],LS_evaluate(F_train[mask[:N],0:d+1],T_d[-1])))
J_vt_d.append(J_error(Yvt,LS_evaluate(F_vt[:,0:d+1],T_d[-1])))
J_train.append(J_train_d)
T.append(T_d)
J_vt.append(J_vt_d)
return J_train,T,J_vt
# A function to randomly split a data set
def split_set(X_0,Y_0,fraction):
N = X_0.shape[0]
N_split = np.round(fraction * X_0.shape[0]).astype(np.int32)
mask = np.random.permutation(N)
X_1 = X_0[mask[N_split:-1]]
Y_1 = Y_0[mask[N_split:-1]]
X_0 = X_0[mask[:N_split]]
Y_0 = Y_0[mask[:N_split]]
return X_0,X_1,Y_0,Y_1
def random_splitting_eval_J_Ridge(X_0,Y,deg_list,lambda_list,num_splits,fraction):
J_train = np.zeros([len(deg_list),len(lambda_list)])
J_val = np.zeros([len(deg_list),len(lambda_list)])
for k in range(num_splits):
# Split training set in training and validation
X_train,X_val,Y_train,Y_val = split_set(X_0,Y,fraction)
# This only evaluates MSE! Not including the penalization! For validation purposes ...
J_t,_,J_v = eval_J_Ridge(X_train,X_val,deg_list,Y_train,Y_val,X_train.shape[0],lambda_list)
J_train += np.array(J_t)
J_val += np.array(J_v)
J_train /= (num_splits+0.0)
J_val /= (num_splits+0.0)
lambda_opt = np.zeros([len(deg_list),1])
for i,d in enumerate(deg_list):
lambda_opt[i] = lambda_list[np.argmin(J_val[i,:])]
return J_train,J_val,lambda_opt
# +
deg_list = [5,10,15]
lambda_list = list(np.logspace(-4,2,20))
J_train,J_val,lambda_opt = random_splitting_eval_J_Ridge(X_0,Y,deg_list,lambda_list,200,0.8)
# +
# Lets visualize the train and validation MSE
f, axes = plt.subplots(1, len(deg_list))
for i,t in enumerate(deg_list):
axes[i].semilogx(lambda_list,J_train[i,:],'b-o',label='Train MSE')
axes[i].semilogx(lambda_list,J_val[i,:],'r--<',label='Validation MSE')
axes[i].set_title("Degree = %d, lambda* = %f" %(deg_list[i],lambda_opt[i]))
axes[i].set_xlabel("$\lambda$")
axes[i].legend()
plt.rcParams["figure.figsize"] = [20,8]
# +
# And visualize the solution with and without regularization
index_model = 2
deg = deg_list[index_model]
l_opt = lambda_opt[index_model]
# This is not data! Just to visualize the polynomial
values = np.sort(np.append(np.arange(4,10,0.01),X_0))
X_plot = values = np.sort(np.append(np.arange(4,10,0.01),X_0))
F_train,train_mean,train_std = create_feature_matrix(X_0,deg,mu=0,std=1,flag_train=True)
F_plot,_,_ = create_feature_matrix(X_plot,deg,mu=0,std=1,flag_train=True)
T_lambda_0 = Ridge_solution(F_train,Y,0)
T_lambda_opt = Ridge_solution(F_train,Y,l_opt)
Y_est_0 = LS_evaluate(F_plot,T_lambda_0)
Y_est_opt = LS_evaluate(F_plot,T_lambda_opt)
plt.plot(X_0,Y,'s', label='Real Values (Train)')
plt.plot(X_0_test,Y_test,'m*', label='Real Values (Test)')
plt.plot(values,Y_est_0,'r-',ms=20,label='lambda = 0')
plt.plot(values,Y_est_opt,'b-',ms=20,label='lamda = %r' %(l_opt[0]))
plt.legend()
plt.rcParams["figure.figsize"] = [8,8]
| Notebooks/Session 4 Regularization/S4-Learning Curves, Regularization and Cross Validation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/mahfuz978/TECH-I.S.---DATA-PROCESSING/blob/main/Mahfuzur_Rahman_Step_1_4e_Pandas_Data_Visualization.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="qQbdAHLm9HbI"
import pandas as pd
import seaborn as sns
import numpy as np
# + colab={"base_uri": "https://localhost:8080/"} id="ECLKrftH9hgT" outputId="ac6aed0b-e8bf-49e3-acc8-bcbdeea695c6"
arr1 = np.random.rand(10,4)
arr1
# + colab={"base_uri": "https://localhost:8080/", "height": 358} id="D_YZf7e49qtT" outputId="931a75d0-17dd-4247-db8d-6a4b15f57bad"
df = pd.DataFrame(arr1, columns=['a','b','c','d'])
df
# + colab={"base_uri": "https://localhost:8080/", "height": 316} id="_YWh2utK97Tn" outputId="13eb76e1-f827-4a31-9d09-7e17107ed3d5"
df.plot.bar(figsize = (8,5));
# + colab={"base_uri": "https://localhost:8080/", "height": 316} id="uY3WBvpl-HC7" outputId="93557cb5-25fe-4ceb-f7b6-af516e3bd472"
df.plot.bar(stacked = True, figsize = (8,5));
# + colab={"base_uri": "https://localhost:8080/", "height": 320} id="gRGLPWpG-eWZ" outputId="470cf8f3-9996-4986-ecc7-2bf09667c52a"
df.plot.barh(figsize = (8,5), stacked = True);
# + id="jElgPXaz-2P9"
# set the color palette
sns.set_palette('magma')
# + colab={"base_uri": "https://localhost:8080/", "height": 316} id="u5NR9jCS_VN1" outputId="e6e9fee8-bb36-4196-edbf-1cbc7f948b08"
df.plot.bar(figsize = (8,5), stacked = True);
# + colab={"base_uri": "https://localhost:8080/", "height": 428} id="GPp5bB9e_YJg" outputId="1f19badd-74e9-434c-9a69-88612e5caae7"
sns.set_palette('muted')
df.plot.area(figsize = (10,7));
# + colab={"base_uri": "https://localhost:8080/", "height": 428} id="wyREu6ZW_xTn" outputId="b5a21a22-90a2-4183-ca0d-dc5b41392b98"
df.plot.area(figsize = (10,7),stacked = False);
# + colab={"base_uri": "https://localhost:8080/", "height": 358} id="tzRaesVSAKeE" outputId="28e0dafb-2d2c-43b2-ab67-a0b03f9706ca"
df.diff()
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="fD9_-kUcAtgi" outputId="0250b063-02e8-4a7c-acd9-f441380ec984"
df.diff().plot.box(vert = False,
color={'medians':'lightblue',
'boxes':'blue',
'caps':'darkblue',});
# + id="ApdS_T5BBAjQ"
arr = np.random.rand(100,1)
# + colab={"base_uri": "https://localhost:8080/"} id="4NSMAkQgBgJK" outputId="d933e688-fdb6-4cf5-a5ca-00a02d04e319"
arr[:5]
# + colab={"base_uri": "https://localhost:8080/", "height": 417} id="xVKGx-FcBhVL" outputId="aeb61bd9-4aa1-4a29-bcb6-868445789b2a"
df = pd.DataFrame(arr, columns=["val"]).reset_index()
df
# + colab={"base_uri": "https://localhost:8080/", "height": 428} id="I6gTy6gdB8JO" outputId="99df42d0-e729-4b46-9228-5b681facf2d9"
df['val'].plot(figsize = (10,7));
# + colab={"base_uri": "https://localhost:8080/", "height": 428} id="ddP-wJ1ACo8T" outputId="9bd894f9-4835-4205-f463-42c445f508e7"
df.val.rolling(10).mean().plot(figsize = (10,7));
# + colab={"base_uri": "https://localhost:8080/", "height": 268} id="unZ0agKxDC71" outputId="317badd4-5046-42c5-fd53-5b798e378dd1"
df.val.plot.kde();
# + colab={"base_uri": "https://localhost:8080/", "height": 497} id="W75KbZsgF_q-" outputId="3834fbc6-8437-4edb-d815-c8945265d6a5"
df.plot.scatter(x='index',y='val',
c='C', #color of data points
s=df.val*200,figsize=(10,7));
# + colab={"base_uri": "https://localhost:8080/", "height": 421} id="TGaUYumQD33q" outputId="0a1d8c22-79fd-4b17-c5f7-50feb23d85d1"
df.plot.hexbin(x = 'index', y= 'val', gridsize = 20, figsize = (10,7));
# + id="VMcZgN9WEYgW"
arr = np.random.rand(5,2)
# + id="AKqjfzXQJG8V"
df = pd.DataFrame(arr,index=list("ABCDE"), columns=list("XY"))
# + colab={"base_uri": "https://localhost:8080/", "height": 203} id="Ce5Ij9MFJPLK" outputId="062b7f79-59ff-40aa-e753-e1ca23b4a45a"
df
# + colab={"base_uri": "https://localhost:8080/", "height": 285} id="ui-sE_xlJQiC" outputId="0cdfcffb-80de-4dfd-cffb-b7a9dbfd3972"
df.plot.pie(subplots = True, figsize = (10,7));
# + colab={"base_uri": "https://localhost:8080/", "height": 203} id="71l1GGE5JXwy" outputId="f81054f5-f6b5-4626-fa29-6a30052020ec"
arr = np.random.rand(100, 4)
df = pd.DataFrame(arr, columns=list('ABCD'))
df.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 400} id="KpHu5TXfJ3WG" outputId="534d24ca-97ce-4e02-c033-b3c12a8c3279"
df.plot(subplots = True, figsize = (10,7));
# + colab={"base_uri": "https://localhost:8080/", "height": 399} id="dXr37yAZK_Hi" outputId="8afadf23-c9a0-4a0e-ebfc-969e7d25a8b4"
df.plot(subplots = True, layout = (2,2), figsize = (10,7));
# + id="n-ZnO-TTMs_u"
| Mahfuzur_Rahman_Step_1_4e_Pandas_Data_Visualization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Array
# An array is a data structure that stores values of same data type. In Python, this is the main difference between
# arrays and lists.
#
# While python lists can contain values corresponding to different data types, arrays in python can only contain
# values corresponding to same data type. In this tutorial, we will understand the Python arrays with few examples.
# If you are new to Python, get started with the Python Introduction article.
#
# To use arrays in python language, you need to import the standard array module. This is because array is not a
# fundamental data type like strings, integer etc. Here is how you can import array module in python :
# from array import *
# Once you have imported the array module, you can declare an array. Here is how you do it:
# arrayIdentifierName = array(typecode, [Initializers])
# +
my_array = array('i', [1,2,3,4,5])
print(my_array[1])
print(my_array[2])
print(my_array[0])
# -
from array import *
my_array = array('i', [1,2,3,4,5])
for i in my_array:
print(i)
# * Append any value using append() method
my_array = array('i', [1,2,3,4,5])
my_array.append(6)
print(my_array)
# * Insert value using insert() method
my_array = array('i', [1,2,3,4,5])
my_array.insert(0,0)
print(my_array)
# * Extend value using extend() method
my_array = array('i', [1,2,3,4,5])
my_extnd_array = array('i', [7,8,9,10])
my_array.extend(my_extnd_array)
print(my_array)
# * Add items from list into array using fromlist()
my_array = array('i', [1,2,3,4,5])
c=[11,12,13]
my_array.fromlist(c)
print(my_array)
# * Remove any array element using remove() method
my_array = array('i', [1,2,3,4,5])
my_array.remove(4)
print(my_array)
# * Remove last array element using pop() method
my_array = array('i', [1,2,3,4,5])
my_array.pop()
print(my_array)
# * Fetch any element through its index using index()
# method
# +
my_array = array('i', [1,2,3,4,5])
print(my_array.index(5))
my_array = array('i', [1,2,3,3,5])
print(my_array.index(3))
# -
# * Reverse a python array using reverse() method
my_array = array('i', [1,2,3,4,5])
my_array.reverse()
my_array
# * Get array buer information through
# buer_info() method
my_array = array('i', [1,2,3,4,5])
my_array.buffer_info()
# * Check for number of occurrences of an element
# using count() method
my_array = array('i', [1,2,3,3,5])
my_array.count(3)
# * Convert array to a python list with same
# elements using tolist() method
my_array = array('i', [1,2,3,4,5])
c = my_array.tolist()
c
# * Append a string to char array using fromstring()
# method
# # Let's make change
| Allinone py/Array.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This notebook was prepared by [<NAME>](http://donnemartin.com). Source and license info is on [GitHub](https://github.com/donnemartin/interactive-coding-challenges).
# # Challenge Notebook
# ## Problem: Compress a string such that 'AAABCCDDDD' becomes 'A3BC2D4'. Only compress the string if it saves space.
#
# * [Constraints](#Constraints)
# * [Test Cases](#Test-Cases)
# * [Algorithm](#Algorithm)
# * [Code](#Code)
# * [Unit Test](#Unit-Test)
# * [Solution Notebook](#Solution-Notebook)
# ## Constraints
#
# * Can we assume the string is ASCII?
# * Yes
# * Note: Unicode strings could require special handling depending on your language
# * Is this case sensitive?
# * Yes
# * Can we use additional data structures?
# * Yes
# * Can we assume this fits in memory?
# * Yes
# ## Test Cases
#
# * None -> None
# * '' -> ''
# * 'AABBCC' -> 'AABBCC'
# * 'AAABCCDDDD' -> 'A3BC2D4'
# ## Algorithm
#
# Refer to the [Solution Notebook](http://nbviewer.ipython.org/github/donnemartin/interactive-coding-challenges/blob/master/arrays_strings/compress/compress_solution.ipynb). If you are stuck and need a hint, the solution notebook's algorithm discussion might be a good place to start.
# ## Code
class CompressString(object):
def compress(self, string):
# TODO: Implement me
if string is None or not string:
return string
out, prev_char, n = "", string[0], 0
for char in string:
if char == prev_char:
n += 1
else:
out += prev_char + (str(int(n)) if n > 1 else "")
n = 1
prev_char = char
out += prev_char + (str(int(n)) if n > 1 else "")
print(out, string)
return out if len(out) < len(string) else string
# ## Unit Test
#
#
# **The following unit test is expected to fail until you solve the challenge.**
# +
# # %load test_compress.py
from nose.tools import assert_equal
class TestCompress(object):
def test_compress(self, func):
assert_equal(func(None), None)
assert_equal(func(''), '')
assert_equal(func('AABBCC'), 'AABBCC')
assert_equal(func('AAABCCDDDDE'), 'A3BC2D4E')
assert_equal(func('BAAACCDDDD'), 'BA3C2D4')
assert_equal(func('AAABAACCDDDD'), 'A3BA2C2D4')
print('Success: test_compress')
def main():
test = TestCompress()
compress_string = CompressString()
test.test_compress(compress_string.compress)
if __name__ == '__main__':
main()
# -
# ## Solution Notebook
#
# Review the [Solution Notebook](http://nbviewer.ipython.org/github/donnemartin/interactive-coding-challenges/blob/master/arrays_strings/compress/compress_solution.ipynb) for a discussion on algorithms and code solutions.
| arrays_strings/compress/compress_challenge.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SageMath 8.0
# language: ''
# name: sagemath
# ---
# <h1><NAME> - 17/03/2018</h1>
# <h1>EJERCICIO 1</h1>
# <b> Apartado 1</b>
# Programamos una funcion que escoge un numero al azar entre 0 y 1. Después si x es menor que <b>prob0</b> (la probabilidad que le hemos asignado al cero) la funcion devuelve cero, y uno en el caso contrario. Por lo tanto hay una probabilidad <b>prob0</b> de que salga <b>0</b> y <b>1 - prob0</b> de que sea <b>1</b>
def moneda_trucada(prob0):
x = random()
if x <= prob0:
return 0
else:
return 1
([moneda_trucada(1/3) for n in srange(10**5)]).count(1)
# Se puede ver que de 10000 lanzamientos se han obtenido 66520 unos, que es prácticamente 2/3 de 10000, como cabía esperar porque le hemos asignado una probabilidad de 1/3 al cero.
# A continuacion programamos una funcion que recibe el número de lanzamientos a realizar (número de veces que se llama a la función anterior) y la probabilidad de que el lanzamiento de como resultado 0 (y 1-prob0 como resultado 1).
def lanzamientos(nLanza, prob0):
count = 0;
for i in xsrange(0, nLanza):
if moneda_trucada(prob0)==0:
count += 1
return count
L = [lanzamientos(1000, 1/10) for i in xsrange(1000)]
media = sum(L)/len(L)
print media.n()
# Hemos realizado 1000 lanzamientos 1000 veces, con 1/10 de posibilidades de que saliera el cero (el cero y el uno lo podemos considerar cara o cruz independientemente). El resultado es una media de 99.88 veces que ha salido el cero, lo cual era lo esperado.
def function(nVeces, nLanza):
LT = list()
for k in xsrange(1, 10):
L = [lanzamientos(nLanza, k*1/10) for i in xsrange(nVeces)]
media = sum(L)/len(L)
LT.append((k*1/10, media.n()))
return LT
function(1000, 1000)
# <b>Apartado 2</b>
# A continuación se muestra una función que devuelve números aleatorios entre a y b, ambos inclusive.
def aleatorio(a, b):
return (a+(b-a)*random())
Q = [aleatorio(5, 10) for i in xsrange(10)]
print Q
# <b>Apartado 3</b>
# La continuación que sigue tiene el objetivo de contar el número de lanzamientos que se deben realizar para que un jugador con <b>euros</b> Euros se arruine, considerando que el casino tiene dinero infinito. Es obvio que el jugador tarde o temprano se arruina, ya que es el que tiene dinero finito.
def cuantoTardaEnArruinarse(euros):
count = 0
while(1):
x = moneda_trucada(1/2)
count += 1
if x == 0:
euros -= 1
if euros==0:
return count
else:
euros += 1
W = [cuantoTardaEnArruinarse(100) for i in xsrange(10)]
print W
# Suponiendo que el jugador decide retirarse cuando le quede la mitad o cuando alcance el doble de su dinero inicial, calcularemos la probabilidad de "ganar", que es duplicar su dinero.
# La siguiente función simula un "Juego", es decir, recibe la cantidad inicial del jugador y realiza lanzamientos, dandole un euro si el jugador acierta o quitándoselo en caso contrario, hasta que el jugador gana (duplica su dinero) devolviendo 1 o pierde (alcanza la mitad de su dinero inicial) devolviendo 0.
def juego(euros):
ini = euros
while(1):
x = moneda_trucada(1/2)
if x==0:
euros -= 1
if euros == (ini//2):
return 0
else:
euros += 1
if euros == 2*ini:
return 1
# Por ultimo, la siguiente función realiza <b>N</b> juegos con un dinero inicial de <b>euros</b> y devuelve la suma de las veces que el jugador ganó, dividido entre el número de partidas.
def probJuego(N, euros):
count = 0
for i in xsrange(N):
if juego(euros)==1:
count += 1
return (count/N).n()
# %time probJuego(10000, 100)
# Se puede ver que gana al rededor de 1/3 de las partidas, lo cual es claramente NO rentable.
| 2_Curso/Laboratorio/SAGE-noteb/IPYNB/IPYNB-mios/Ejercicios semana 12 de marzo - Alejandro Santorum.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1. Read the Dataset¶
#
# # 1.1 File Info.
# * 문서 앱내 사용시 발생하는 클라이언 로그 데이터 (after parsed)
# * 서버로그와 달리, 유저의 행동이 발생할 경우 로그 수집 (유저 행동 패턴 파악에 용이)
# * 클라이언트(앱) 로그
# * 서버 로그_ really purchase
# +
# %matplotlib inline
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import warnings
warnings.filterwarnings('ignore')
plt.style.use('seaborn-paper')
# -
import pandas as pd
df= pd.read_csv('df_funnel.csv',index_col=0)
df.head(5)
df.tail(5)
# +
# key infor => data en entries, missing values, type
df.info()
# -
#
# # 2. Preprocessing
# * 데이터 타입 변환
# * 데이터 값 변경
# * 결측치 처리
# * 신규 session id 부여
# * 대소문자 처리
# # 2.1 날짜를 pandas datetime 형태로 변환하기
#
# pandas.to_datetime()
#
# +
# if datetime is string? no sort
df.groupby ('datetime').size().head(15)
# -
## case 1. if data type is string,
str_date =['2018/01/01','2018.01.02','2018-01-03','2018-01-04','2018/01/05']#string type
str_date
pd.to_datetime(str_date)# parsed
pd.Series(pd.to_datetime(str_date))# to pd:series
## case 2. 날짜가 timestamp인 경우
ts_date =[1349720105, 1349806505, 1349892905, 1349979305, 1350065705]
ts_date
pd.Series(pd.to_datetime([1530837876, 1530751476, 1530665076, 1530578676, 1530492276], unit='s')).dt.date
# * timestamp converter: https://www.epochconverter.com/
# +
## case 3. in case of changing the type of column
df.info()
# -
df.head() # object =string
# option1. string to datetime
df['datetime'].astype('datetime64[ns]').head()
# option2
pd.to_datetime(df['datetime']).head()
df['datetime']=pd.to_datetime(df['datetime'])
df.info()
df.head()
df.groupby('datetime').size()#.plot()
# day to year
df['datetime'].dt.year[:10]
# day to month
df['datetime'].dt.month[:10]
# day to day
df['datetime'].dt.day[:10]
# # 2.2 Missing value 확인
df.info()
df.isnull().sum()
# * 2.3 결측치 처리
# Drop
# 경우에 따라 결측치 처리 방법이 달라진다.
# 샘플수가 많다면 missing values 를 포함하는 행을 모두 삭제하는 것이 가능하다
# * 결측치가 하나라도 있으면 버리는 코드 예제
# df.dropna()
#
# * 모든 값이 Null일 경우만 버리는 코드 예제
# df.dropna(how='all')
#
# * 결측치가 하나 이상 있는 Case만 선택하는 코드 예제
# df[df.isnull().any(axis=1)]
# * Imputation
# * 만약 샘플수가 충분하지 않을 경우, Pandas의 fillna() 명령어로 Null 값을 채우는 것이 가능하다.
# * 연속형인 경우 Mean이나 Median을 이용하고 명목형인 경우 Mode(최빈치)나 예측 모형을 통해 Null 값을 대체할 수 있다.
# * Null 값을 median으로 대체하는 코드 예제
# df.fillna(df.mean())
# Reference
# https://machinelearningmastery.com/handle-missing-data-python/
df_by_screen = df.groupby(['datetime','screen'])['sessionid'].nunique().unstack()
df_by_screen = df.groupby(["datetime", "screen"])['sessionid'].nunique().unstack()
df_by_screen[:10]
df_by_screen.isnull().sum()
pd.get_dummies(x)
| DSES_2nd_review/19-05-11_lecturenote.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <center><img src="http://i.imgur.com/sSaOozN.png" width="500"></center>
# ## Course: Computational Thinking for Governance Analytics
#
# ### Prof. <NAME>, PhD
# * Visiting Professor of Computational Policy at Evans School of Public Policy and Governance, and eScience Institute Senior Data Science Fellow, University of Washington.
# * Professor of Government and Political Methodology, Pontificia Universidad Católica del Perú.
#
# _____
#
# # Session 1: Programming Fundamentals
# ## Part C: Building Functions in Python
# We build functions to make the code more readable. Functions, plus the data structures and control of execution capabilities you saw before, will give you the basic tools you need to develop programs.
#
# A function is a three-step process: Input, Transformation, Output. For example, if you need to convert a numeric value from Fahrenheit into Celsius , the input is the value in Fahrenheit, the transformation is the formula, and the output the result of the formula (a value in Celsius).
def converterToCelsius(valueInFarenheit): #input
#transformation
resultInCelsius= (valueInFarenheit-32)*5/9
#output
return resultInCelsius
# From above, creating functions in Python requires the use of **def** followed by the name of the function; the function arguments continue between parenthesis. The process comes after the _colon_, notice that _identation_ is needed. The command **return** serves to give the output. For Python, there is a new function available:
converterToCelsius(100)
# <a id='beginning'></a>
# This session will be organized on the following topics:
#
# 1. [Input components.](#part1)
# 2. [Output organization.](#part2)
# 3. Applying functions
# + [to simple structures.](#part3)
# + [to composite structures.](#part4)
#
#
# ____
#
# <a id='part1'></a>
#
# ## The function input
# We control the amount of input in a function:
# this function requires TWO inputs:
def XsumY(valueX,valueY):
###
resultSum=valueX+valueY
###
return resultSum
# The code above receives two values and outputs their sum. You can see how it works this way:
XsumY(3,10)
# The next function uses two inputs and one of them has a *default* value:
def riseToPower(base,exponent=2): # two argument names!!!, with a exponent default equals to 2
###
result=1
if exponent > 0:
for times in range(1,exponent+1): # use 'exponent + 1'...!
result=result*base
###
return(result)
# Since you have a default value in the input arguments, you decide if you give that input or not. Let’s see how it works:
riseToPower(9)
riseToPower(9,3)
riseToPower(9,0)
# for sure you can use the arguments name:
riseToPower(base=9,exponent=0)
# using arguments names does not require order:
riseToPower(exponent=0,base=9)
# ### Homework:
#
# Change the above function to create the function **riseToPowerPlus**, which gives a good answer even when the power is negative.
#
# ____
def riseToPowerPlus(base,exponent): # two argument names
###
result=1
if exponent > 0:
for times in range(1,exponent+1): # use 'exponent + 1'...!
result=result*base
elif exponent < 0:
for lalala in range(exponent,0):
result=result*(1/base)
###
return(result)
#
riseToPowerPlus(base=2,exponent=-2)
# Functions need argument names in the input definition, but if you have many arguments, you need to keep the order. However, Python offers two additional ways to input **several arguments**. First, let me know what happens when we divide by zero:
3/0
# Then
def divRounded(numerator,denominator,precision=2):
try:
result = numerator/denominator
return round(result, precision)
except ZeroDivisionError:
print('You can not use 0 as the denominator')
# testing:
n=13
d=12
p=5
divRounded(n,d,p)
# A different approach would be to use a list or tuple with the arguments, the function requires ONE '*':
inputArgs=[13,12,5] # order matters, keep it.
divRounded(*inputArgs)
# A dict can be very useful, just use TWO '*':
inputArgs={'numerator':13, 'precision':5,'denominator':12} # order does not matter
divRounded(**inputArgs)
# [Go to page beginning](#beginning)
# ____
#
# <a id='part2'></a>
#
# ## The function output
# Our output has been a single value, but it can be several ones; however, you need the right structure.
# one input, and several output in simple data structure:
def factors(number):
factorsList=[] # empty list that will collect output
for i in range(1, number + 1):
#if the remainder of 'number'/'i' equals zero...
if number % i == 0:
# ...add 'i' to the list of factors!
factorsList.append(i)
return factorsList # returning values in a list.
factors(20)
# ### Homework:
#
# Change the function ’factors’to reduce the amount of iterations in the for loop and still get the factors shown above.
# In this next case, you can have several input, and get an output organized in a more complex structure (a data frame):
# several input, a composite data structure:
def powerDF(aList,power=2):
import pandas as pd
# list comprehension
powerList=[val**power for val in aList]
# both lists into a dict:
answerAsDicts={'number':aList,'power'+str(power):powerList}
# data frame is created, and that is returned:
return pd.DataFrame(answerAsDicts)
powerDF(factors(10),3)
# of course, this works:
valsDict={'aList':factors(10), 'power':3}
powerDF(**valsDict)
# ### Homework:
# Make a function that reads two lists and returns a data frame with those lists and extra columns with their sum, difference, multiplication and division.
# [Go to page beginning](#beginning)
#
# ____
# <a id='part3'></a>
#
# ## Applying functions to simple structures
# Imaging you have created a function that converts a value like:
def double(x):
return 2*x
# and you have this list:
myList=[1,2,3]
# What can you get here?
double(myList)
# I bet you wanted something like this:
map(double,myList)
# You just see an strange result!...Well Python did do what you need, but you can't see it because it returned an **iterator**. Do this then:
list(map(double,myList))
# With **map** you can apply the function to every element of the list.
# Easy functions can be written using **lambda** notation:
double2=lambda x: 2*x
list(map(double2,myList))
# You can use these functions to create filters:
drinkingAge= lambda x: x >= 21
agesList=[12,34,56,19,24,13]
list(filter(drinkingAge,agesList))
# In the last line above, you filtered the original vector agesVals by combining **filter** and _drinkingAge_, the filtering works by selecting the values that have TRUE in the output of drinkingAge.
# [Go to page beginning](#beginning)
#
# ____
# <a id='part4'></a>
#
# ## Applying functions to composite structures
# We will be using data frames often. This is a particular structure that has its **own** mechanism to apply functions:
#Creating data frame
import pandas as pd
data={'numberA':[10,20,30,4,5],'numberB':[6,7,8,9,10]}
dataDF=pd.DataFrame(data)
dataDF
# Now applying function _double_ to it:
double(dataDF)
# The function at the element level worked well, that is because the columns (which came from a list) are now arrays.
# However, often you need to put more effort to make functions work in pandas. The function **apply** is very important to use a function in a data frame in pandas:
# this will double each element column-wise
dataDF.apply(double,axis=0)
# this will double each element row-wise
dataDF.apply(double,axis=1)
# The axis argument tells in what direction the function should be applied. Double works at the level of cells, so it made no difference.
#
# Our function made no difference, but compare for _sum_:
# the sum of the colums
dataDF.apply(sum,axis=0)
# the sum of the rows
dataDF.apply(sum,axis=1)
# Compare for min:
dataDF.apply(min) # axis=0 is the default, I can omit it.
dataDF.apply(min,axis=1)
# Pandas has the function **applymap** to especifically apply a function to every cell of the data frame:
dataDF.applymap(double)
# You can have functions that operate at the cell level, or at the column (_Series_) level; _apply_ will work at both levels, in the particular axis of interest. _applymap_ works at the cell level for data frames as a whole, but not at the _Series_ level. Sometimes the difference is not obvious.
# Just make sure what you have:
# This is a Series
dataDF.numberA
# This is a Series
dataDF['numberA']
# This is a data frame:
dataDF[['numberA']]
# This is a Series
dataDF.loc[:,'numberA']
# This is a data frame:
dataDF.loc[:,['numberA']]
# This is a Series
dataDF.iloc[:,0]
# This is a data frame:
dataDF.iloc[:,[0]]
# ____
#
# Solve the homework in a new Jupyter notebook, and then upload it to GitHub. Name the notebook as 'hw_functions'.
# _____
#
# * [Go to page beginning](#beginning)
# * [Go to REPO in Github](https://github.com/EvansDataScience/ComputationalThinking_Gov_1)
# * [Go to Course schedule](https://evansdatascience.github.io/GovernanceAnalytics/)
| .ipynb_checkpoints/S1_C_Py_progfunctions-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Use PyTorch to recognize hand-written digits with `ibm-watson-machine-learning`
# This notebook contains steps and code to demonstrate support of Deep Learning model training and scoring in Watson Machine Learning service. It introduces commands for getting data, training_definition persistance to Watson Machine Learning repository, model training, model persistance, model deployment and scoring.
#
# Some familiarity with Python is helpful. This notebook uses Python 3.8.
#
#
# ## Learning goals
#
# The learning goals of this notebook are:
#
# - Working with Watson Machine Learning service.
# - Training Deep Learning models (TensorFlow).
# - Saving trained models in Watson Machine Learning repository.
# - Online deployment and scoring of trained model.
#
#
# ## Contents
#
# This notebook contains the following parts:
#
# 1. [Setup](#setup)
# 2. [Create model definition](#model_def)
# 3. [Train model](#training)
# 4. [Persist trained model](#persist)
# 5. [Deploy and Score](#deploy)
# 6. [Clean up](#clean)
# 7. [Summary and next steps](#summary)
# <a id="setup"></a>
# ## 1. Set up the environment
#
# Before you use the sample code in this notebook, you must perform the following setup tasks:
#
# - Contact with your Cloud Pack for Data administrator and ask him for your account credentials
# ### Connection to WML
#
# Authenticate the Watson Machine Learning service on IBM Cloud Pack for Data. You need to provide platform `url`, your `username` and `api_key`.
username = 'PASTE YOUR USERNAME HERE'
api_key = 'PASTE YOUR API_KEY HERE'
url = 'PASTE THE PLATFORM URL HERE'
wml_credentials = {
"username": username,
"apikey": api_key,
"url": url,
"instance_id": 'openshift',
"version": '4.0'
}
# Alternatively you can use `username` and `password` to authenticate WML services.
#
# ```
# wml_credentials = {
# "username": ***,
# "password": ***,
# "url": ***,
# "instance_id": 'openshift',
# "version": '4.0'
# }
#
# ```
# ### Install and import the `ibm-watson-machine-learning` package
# **Note:** `ibm-watson-machine-learning` documentation can be found <a href="http://ibm-wml-api-pyclient.mybluemix.net/" target="_blank" rel="noopener no referrer">here</a>.
# !pip install -U ibm-watson-machine-learning
# +
from ibm_watson_machine_learning import APIClient
client = APIClient(wml_credentials)
# -
# ### Working with spaces
#
# First of all, you need to create a space that will be used for your work. If you do not have space already created, you can use `{PLATFORM_URL}/ml-runtime/spaces?context=icp4data` to create one.
#
# - Click New Deployment Space
# - Create an empty space
# - Go to space `Settings` tab
# - Copy `space_id` and paste it below
#
# **Tip**: You can also use SDK to prepare the space for your work. More information can be found [here](https://github.com/IBM/watson-machine-learning-samples/blob/master/cpd4.0/notebooks/python_sdk/instance-management/Space%20management.ipynb).
#
# **Action**: Assign space ID below
space_id = 'PASTE YOUR SPACE ID HERE'
# You can use `list` method to print all existing spaces.
client.spaces.list(limit=10)
# To be able to interact with all resources available in Watson Machine Learning, you need to set **space** which you will be using.
client.set.default_space(space_id)
# <a id="model_def"></a>
# # 2. Create model definition
# ### 2.1 Prepare model definition metadata
model_definition_metadata = {
client.model_definitions.ConfigurationMetaNames.NAME: "PyTorch Hand-written Digit Recognition",
client.model_definitions.ConfigurationMetaNames.DESCRIPTION: "PyTorch Hand-written Digit Recognition",
client.model_definitions.ConfigurationMetaNames.COMMAND: "torch_mnist.py --epochs 1",
client.model_definitions.ConfigurationMetaNames.PLATFORM: {"name": "python", "versions": ["3.8"]},
client.model_definitions.ConfigurationMetaNames.VERSION: "2.0",
client.model_definitions.ConfigurationMetaNames.SPACE_UID: space_id
}
# ### 2.2 Get sample model definition content file from git
# +
import wget, os
filename='pytorch-model.zip'
if not os.path.isfile(filename):
filename = wget.download('https://github.com/IBM/watson-machine-learning-samples/raw/master/cpd4.0/definitions/pytorch/mnist/pytorch-model.zip')
# -
# **Tip**: Convert below cell to code and run it to see model deinition's code.
# + active=""
# !unzip -oqd . pytorch-model.zip && cat torch_mnist.py
# -
# ### 2.3 Publish model definition
definition_details = client.model_definitions.store(filename, model_definition_metadata)
model_definition_id = client.model_definitions.get_id(definition_details)
print(model_definition_id)
# #### List models definitions
client.model_definitions.list(limit=5)
# <a id="training"></a>
# # 3. Train model
# #### **Warning**: Before executing deep learning experiment make sure that [training data](https://github.com/IBM/watson-machine-learning-samples/tree/master/cpd4.0/data/mnist/raw) is saved in a folder where Watson Machine Learning Accelerator is installed.
# ### 3.1 Prepare training metadata
training_metadata = training_metadata = {
client.training.ConfigurationMetaNames.NAME: "PyTorch hand-written Digit Recognition",
client.training.ConfigurationMetaNames.DESCRIPTION: "PyTorch hand-written Digit Recognition",
client.training.ConfigurationMetaNames.TRAINING_RESULTS_REFERENCE: {
"name":"MNIST results",
"connection":{
},
"location":{
"path":f"spaces/{space_id}/assets/experiment"
},
"type":"fs"
},
client.training.ConfigurationMetaNames.MODEL_DEFINITION:{
"id": model_definition_id,
"hardware_spec": {
"name": "K80",
"nodes": 1
},
"software_spec": {
"name": "pytorch-onnx_1.7-py3.8"
}
},
client.training.ConfigurationMetaNames.TRAINING_DATA_REFERENCES: [
{
"name":"training_input_data",
"type":"fs",
"connection":{
},
"location":{
"path": "pytorch-mnist"
},
"schema":{
"id":"idmlp_schema",
"fields":[
{
"name":"text",
"type":"string"
}
]
}
}
]
}
# ### 3.2 Train the model in the background
training = client.training.run(training_metadata)
# ### 3.3 Get training id and status
training_id = client.training.get_id(training)
print(training_id)
client.training.get_status(training_id)['state']
# ### 3.4 Get training details
# +
import json
training_details = client.training.get_details(training_id)
print(json.dumps(training_details, indent=2))
# -
# #### List trainings
client.training.list(limit=5)
# #### Cancel training
# You can cancel the training run by calling the method below.
# **Tip**: If you want to delete train runs and results add `hard_delete=True` as a parameter.
# + active=""
# client.training.cancel(training_id)
# -
# <a id="persist"></a>
# # 4. Persist trained model
# ### 4.1 Publish model
software_spec_uid = client.software_specifications.get_uid_by_name('pytorch-onnx_1.7-py3.8')
# +
model_meta_props = {client.repository.ModelMetaNames.NAME: "PyTorch Mnist Model",
client.repository.ModelMetaNames.TYPE: "pytorch-onnx_1.7",
client.repository.ModelMetaNames.SOFTWARE_SPEC_UID: software_spec_uid
}
published_model_details = client.repository.store_model(training_id, meta_props=model_meta_props)
model_uid = client.repository.get_model_uid(published_model_details)
# -
# ### 4.2 Get model details
model_details = client.repository.get_details(model_uid)
print(json.dumps(model_details, indent=2))
# #### List stored models
client.repository.list_models(limit=5)
# <a id="deploy"></a>
# # 5. Deploy and score
# ### 5.1 Crate online deployment for published model
# You can deploy the stored model as a web service (online) by running code in the following cell.
# +
deployment = client.deployments.create(model_uid, meta_props={
client.deployments.ConfigurationMetaNames.NAME:"PyCharm Mnist deployment",
client.deployments.ConfigurationMetaNames.ONLINE:{}})
scoring_url = client.deployments.get_scoring_href(deployment)
deployment_uid = client.deployments.get_id(deployment)
# -
# ### 5.2 Get deployments details
deployments_details = client.deployments.get_details(deployment_uid)
print(json.dumps(deployments_details, indent=2))
# ### 5.3 Score deployed model
# Prepare sample scoring data to score deployed model.
# **Hint:** You may need to install wget using following command `!pip install wget`
# +
import wget
dataset_filename='mnist.npz'
if not os.path.isfile(dataset_filename):
dataset_filename = wget.download('https://github.com/IBM/watson-machine-learning-samples/raw/master/cpd4.0/data/mnist/mnist.npz')
# +
import numpy as np
dataset_filename='mnist.npz'
mnist_dataset = np.load(dataset_filename)
x_test = mnist_dataset['x_test']
# -
image_1 = [x_test[0].tolist()]
image_2 = [x_test[1].tolist()]
# %matplotlib inline
import matplotlib.pyplot as plt
for i, image in enumerate([x_test[0], x_test[1]]):
plt.subplot(2, 2, i + 1)
plt.axis('off')
plt.imshow(image, cmap=plt.cm.gray_r, interpolation='nearest')
# Build scoring dictionary consisting of two digits and send it to deployed model to get predictions.
scoring_payload = {
client.deployments.ScoringMetaNames.INPUT_DATA : [
{'values': [image_1, image_2]}
]
}
scores = client.deployments.score(deployment_uid, meta_props=scoring_payload)
print("Scoring result:\n" + json.dumps(scores, indent=2))
# #### List deployments
client.deployments.list(limit=5)
# <a id="clean"></a>
# # 6. Clean up
# If you want to clean up all created assets:
# - experiments
# - trainings
# - pipelines
# - model definitions
# - models
# - functions
# - deployments
#
# please follow up this sample [notebook](https://github.com/IBM/watson-machine-learning-samples/blob/master/cpd4.0/notebooks/python_sdk/instance-management/Machine%20Learning%20artifacts%20management.ipynb).
# <a id="summary"></a>
# # 7. Summary and next steps
# You successfully completed this notebook! You learned how to use `ibm-watson-machine-learning-client` to train and score PyTorch models.
# Check out our [Online Documentation](https://dataplatform.cloud.ibm.com/docs/content/analyze-data/wml-setup.html) for more samples, tutorials, documentation, how-tos, and blog posts.
# ### Author
#
# **<NAME>**, Intern in Watson Machine Learning.
# Copyright © 2020, 2021 IBM. This notebook and its source code are released under the terms of the MIT License.
| cpd4.0/notebooks/python_sdk/experiments/deep_learning/Use PyTorch to recognize hand-written digits.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/abdelkadergelany/Intrusion-Detection-in-IoT-Based-Network/blob/main/Testing_under_sampling.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="SRFv6XrqPxVM"
# data cleaning and plots
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
plt.style.use('ggplot')
# %matplotlib inline
# sklearn: data preprocessing
from sklearn.compose import ColumnTransformer
from sklearn.preprocessing import StandardScaler, LabelEncoder, OneHotEncoder ,MinMaxScaler
# sklearn: train model
from sklearn.model_selection import train_test_split
from sklearn.model_selection import cross_val_score, cross_validate, StratifiedKFold
from sklearn.metrics import precision_recall_curve, precision_score, recall_score, f1_score, accuracy_score
from sklearn.metrics import roc_curve, auc, roc_auc_score, confusion_matrix, classification_report
# sklearn classifiers
from sklearn.linear_model import LogisticRegression
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier,ExtraTreesClassifier
from sklearn.neural_network import MLPClassifier
#Feature selection
from sklearn.feature_selection import chi2
from sklearn.feature_selection import SelectKBest
import xgboost, lightgbm
# + [markdown] id="F30WDG00PxVN"
# # Data Processing
#
# ## Load Data
#
# **UNSW-NB15: a comprehensive data set for network intrusion detection systems**
#
#
# attack_cat: This dataset has nine types of attacks, namely, Fuzzers, Analysis, Backdoors, DoS, Exploits, Generic, Reconnaissance, Shellcode and Worms.
#
# Label: 0 for normal and 1 for attack records
#
# + id="aMvHjORZPxVO"
# Load data
train = pd.read_csv('https://raw.githubusercontent.com/Nir-J/ML-Projects/master/UNSW-Network_Packet_Classification/UNSW_NB15_training-set.csv')
test = pd.read_csv('https://raw.githubusercontent.com/Nir-J/ML-Projects/master/UNSW-Network_Packet_Classification/UNSW_NB15_testing-set.csv')
combined_data = pd.concat([train, test]).drop(['id'],axis=1)
# + id="GQlJuki7PxVP" colab={"base_uri": "https://localhost:8080/"} outputId="df2d91ea-da9c-4b0e-b78d-f71f605a1c22"
# Look at the first 5 rows
print(combined_data)
# + id="fJv9WFwoPxVQ" colab={"base_uri": "https://localhost:8080/"} outputId="18697d90-7ecf-43ba-d498-85a72c7e4ab0"
# information of the data:
combined_data.info()
# + [markdown] id="TwPGNv4BPxVR"
#
# ### Check missing values
# First, we should check if there are missing values in the dataset.
# + id="OEpul595PxVS" colab={"base_uri": "https://localhost:8080/"} outputId="367f2015-adbe-44e9-aa9b-68c096938c63"
# check if there are Null values
combined_data.isnull().sum()
# + [markdown] id="qo9HocAXPxVT"
# A basic strategy to use incomplete datasets is to discard entire rows and/or columns containing missing values. Actually, there exists some strategies to impute missing values (see [here](https://scikit-learn.org/stable/modules/impute.html)).
# + id="6uS7cLmFPxVT" colab={"base_uri": "https://localhost:8080/"} outputId="29479f85-1d4c-44f9-f9ad-53c175ec0c73"
# Discard the rows with missing values
data_to_use = combined_data.dropna()
# Shape of the data: we could see that the number of rows remains the same as no null values were reported
data_to_use.shape
# + [markdown] id="GgJ6oSU_as6l"
# <h1>Contamination mean pollution (outliers) in data</h1>
# + colab={"base_uri": "https://localhost:8080/"} id="ck1K_7ecapc-" outputId="7bfbf356-43ed-47ef-b43b-355ad4c54d2f"
# Contaminsation mean pollution (outliers) in data
tmp = train.where(train['attack_cat'] == "Normal").dropna()
contamination = round(1 - len(tmp)/len(train), 2)
print("train contamination ", contamination)
tmp = test.where(test['attack_cat'] == "Normal").dropna()
print("test contamination ", round(1 - len(tmp)/len(test),2),'\n')
if contamination > 0.5:
print(f'contamination is {contamination}, which is greater than 0.5. Fixing...')
contamination = round(1-contamination,2)
print(f'contamination is now {contamination}')
# + id="QMw1Oynscjm4"
# + id="bEaX2MbxcPwQ" outputId="45e42ff2-d7a1-44d5-889e-cc5de9bd2a8a" colab={"base_uri": "https://localhost:8080/"}
tmp.shape
# + [markdown] id="ynNesImwPxVT"
# ### Check imbalanced issue on y
#
# First, we get the `X` and `y1` and `y2` .
# + id="vFAeAuusPxVU"
# X = data_to_use.drop(axis=1, columns=['attack_cat']) # X is a dataframe
# X = X.drop(axis=1, columns=['label'])
# y1 = data_to_use['attack_cat'].values # y is an array
# y2 = data_to_use['label'].values
# #We will convert the orginal training data to the datframes called X_train, y1_train, y2_train
# X_train = X
# y1_train = y1
# y2_train = y2
# + [markdown] id="xBzbTxYKdqd5"
# <h1>Encoding non numeric Values</h1>
# + colab={"base_uri": "https://localhost:8080/"} id="j5HlZ5VhdkJ-" outputId="cb344c65-c5f3-42bb-bd1d-d29805107cc8"
le1 = LabelEncoder()
le = LabelEncoder()
vector = combined_data['attack_cat']
print("attack cat:", set(list(vector))) # use print to make it print on single line
combined_data['attack_cat'] = le1.fit_transform(vector)
combined_data['proto'] = le.fit_transform(combined_data['proto'])
combined_data['service'] = le.fit_transform(combined_data['service'])
combined_data['state'] = le.fit_transform(combined_data['state'])
vector = combined_data['attack_cat']
print('\nDescribing attack_type: ')
print("min", vector.min())
print("max", vector.max())
print("mode",vector.mode(), "Which is,", le1.inverse_transform(vector.mode()))
print("mode", len(np.where(vector.values==6)[0])/len(vector),"%")
# + [markdown] id="9YOOek9Le3lw"
# <h1>Spliting the dataset</h1>
# + id="9yCFUo94e2z0"
data_x = combined_data.drop(['attack_cat','label'], axis=1) # droped label
data_y = combined_data.loc[:,['label']]
X_train, X_test, y_train, y_test = train_test_split(data_x, data_y, test_size=.20, random_state=42)
# + id="7wWdFLn_lnsI" outputId="173d3c76-535c-460d-c14f-25482b741027" colab={"base_uri": "https://localhost:8080/", "height": 439}
combined_data
# + id="ZCaz1tAfGM6r" outputId="9d3576ee-0f91-468c-e503-16d17f0beddd" colab={"base_uri": "https://localhost:8080/", "height": 142}
y_train.head(3)
# + [markdown] id="TCTJsAJIeXG9"
# <h1>Feature selection with CHi2</h1>
# + [markdown] id="NLFGxfFgOzth"
#
# + id="_4coPg1_ecw3"
#Feature selection
chi2_selector = SelectKBest(chi2, k=14)
X_train_kbest = chi2_selector.fit_transform(X_train, y_train)
X_test_kbest = chi2_selector.fit_transform(X_test, y_test)
#print(X_kbest.shape)
# X_train_transform = X_train_kbest
# X_test_transform = X_test_kbest
X_train = X_train_kbest
X_test = X_test_kbest
# + id="RODqVVNvPxVU"
# Calculate Y2 ratio
def data_ratio(y2):
'''
Calculate Y2's ratio
'''
unique, count = np.unique(y2, return_counts=True)
ratio = round(count[0]/count[1], 1)
return f'{ratio}:1 ({count[0]}/{count[1]})'
# + id="iHdZivEPqpxy"
combined_data = pd.concat([train, test]).drop(['id'],axis=1)
attacks_type = combined_data['attack_cat'].values
class_type = combined_data['label'].values
# + id="KuibhvM5q5yQ"
# + id="ku2dx9RyPxVU" colab={"base_uri": "https://localhost:8080/", "height": 690} outputId="ee1d1257-758b-414a-d1b4-09aa4575ad83"
print('The class ratio for the original data:', data_ratio(attacks_type))
plt.figure(figsize=(13,5))
sns.countplot(attacks_type,label="Sum")
plt.show()
print('The class ratio for the original data:', data_ratio(class_type))
sns.countplot(class_type,label="Sum")
plt.show()
# + id="dNCY5xYljoI7"
normal = combined_data.where(combined_data['attack_cat'] == 6).dropna()
Backdoor = combined_data.where(combined_data['attack_cat'] == 5).dropna()
Analysis = combined_data.where(combined_data['attack_cat'] == 4).dropna()
Fuzzers = combined_data.where(combined_data['attack_cat'] == 3).dropna()
Shellcode = combined_data.where(combined_data['attack_cat'] == 2).dropna()
Reconnaissance = combined_data.where(combined_data['attack_cat'] == 1).dropna()
Exploits = combined_data.where(combined_data['attack_cat'] != 0).dropna()
Dos = combined_data.where(combined_data['attack_cat'] == 7).dropna()
Worms = combined_data.where(combined_data['attack_cat'] == 8).dropna()
Generic = combined_data.where(combined_data['attack_cat'] == 9).dropna()
# + id="iBVL1GCkkl3x" outputId="44a8afe1-6fc7-4b34-a597-0085fba3c768" colab={"base_uri": "https://localhost:8080/"}
normal.shape
# + id="BeBJDWf0pMQV"
normal = normal.append(Generic.head(850), ignore_index = True)
normal = normal.append(Worms.head(850), ignore_index = True)
normal = normal.append(Dos.head(850), ignore_index = True)
normal = normal.append(Exploits.head(850), ignore_index = True)
normal = normal.append(Reconnaissance.head(850), ignore_index = True)
normal = normal.append(Shellcode.head(850), ignore_index = True)
normal = normal.append(Fuzzers.head(850), ignore_index = True)
normal = normal.append(Analysis.head(850), ignore_index = True)
normal = normal.append(Backdoor.head(850), ignore_index = True)
# + id="Y4PnEqAYrBY7" outputId="71b54c1a-8e95-4f07-82e6-96f8e987b452" colab={"base_uri": "https://localhost:8080/", "height": 690}
#combined_data = pd.concat([train, test]).drop(['id'],axis=1)
attacks_type = normal['attack_cat'].values
class_type = normal['label'].values
print('The class ratio for the original data:', data_ratio(attacks_type))
plt.figure(figsize=(13,5))
sns.countplot(attacks_type,label="Sum")
plt.show()
print('The class ratio for the original data:', data_ratio(class_type))
sns.countplot(class_type,label="Sum")
plt.show()
# + id="55WtluXPpgu9" outputId="e9552f2b-2091-445f-f129-ebc41711d790" colab={"base_uri": "https://localhost:8080/"}
normal.shape
# + id="s6nelPTTrdTs"
data_x = normal.drop(['attack_cat','label'], axis=1) # droped label
data_y = normal.loc[:,['label']]
X_train, X_test, y_train, y_test = train_test_split(data_x, data_y, test_size=.20, random_state=42)
# + id="KCDIYQnIrn8j"
y_train
# + [markdown] id="gYGDhWScPxVV"
# We could see that the dataset is not perfectly balanced. There are some sampling techniques to deal with this issue. Here, we ignore this issue because we are aimed to implement several ML models to compare their performance.
# + [markdown] id="chI822OUPxVf"
# # Train ML Models
#
# We will train several machine learning models for the training set and evaluate their performance on both training and testing set.This will helps us to choose the best Supervised algorithm.
#
#
#
# + id="2m4pxcBMhBn7"
# y_train = y_train['label'].values
# y_test = y_test['label'].values
# + id="4szgxJw3g0Dd"
# ===== Step 1: cross-validation ========
# define a Logistic Regression classifier
clf = LogisticRegression(solver='lbfgs', random_state=123, max_iter = 4000)
# define Stratified 5-fold cross-validator, it provides train/validate indices to split data in train/validate sets.
cv = StratifiedKFold(n_splits=5, shuffle=True, random_state=123)
# define metrics for evaluating
scoring = ['accuracy', 'precision', 'recall', 'f1', 'roc_auc']
# perform the 5-fold CV and get the metrics results
cv_results = cross_validate(estimator=clf,
X=X_train,
y=y_train,
scoring=scoring,
cv=cv,
return_train_score=False)
# + id="wxfDDIbRhS5O" colab={"base_uri": "https://localhost:8080/"} outputId="22f3c8ea-0328-4de7-947d-4c9457f7a63f"
cv_results['test_accuracy'].mean()
# + [markdown] id="RbRkyQNiPxVi"
# ## Several ML Models
#
# We will implement several ML models through the above steps. The only difference part is to change `clf = LogisticRegression()` as other model functions, for example, `clf = DecisionTreeClassifier()`.
#
# The followings are ML models functions:
#
# - `LogisticRegression()`
# - `DecisionTreeClassifier()`
# - `RandomForestClassifier()`
# - `MLPClassifier()`
#
# Note that in `MLPClassifier()` we set the solver as `lbfgs` which has better performance for small size of data. Also, we set the maximum iterations as 5000 to ensure convergence. `random_state` is used to ensure reproducible results.
# + id="9Rf6MMmvPxVj"
# Define four models
models = [('LogisticRegression', LogisticRegression(random_state=123, max_iter=5000)),
('DecisionTree', DecisionTreeClassifier(random_state=123)),
('RandomForest', RandomForestClassifier(random_state=123)),
('MultiLayerPerceptron', MLPClassifier(random_state=123, solver='adam', max_iter=8000)),
('ExtraTreesClassifier', ExtraTreesClassifier(n_estimators=200, random_state=42, n_jobs=-1))
]
# + [markdown] id="sl6KRIKiPxVj"
# We could check the hyperparameters values in these models:
# + id="H7lBSS5wPxVj"
for model_name, clf in models:
print(clf)
# + id="y8EoC-DF4Fwm"
y_train_copy = y_train['label'].values
y_test_copy = y_test['label'].values
# + [markdown] id="5HbJR1v3PxVk"
# **Finally, we write the code to perform the above four ML models and store their cross-validation results and evaluation results on testing data.**
# + [markdown] id="xjTdnJlAGOQq"
# <h1>#Test our candidate algorithms
# <h1>
# + colab={"base_uri": "https://localhost:8080/"} id="6uutqufPGH-T" outputId="9a6982bc-efb6-4f2a-f8d9-f12cdcfd0fc1"
#Test our candidates algorithm
# DTC = DecisionTreeClassifier()
RFC = RandomForestClassifier(n_estimators=150, random_state=42, n_jobs=-1)
ETC = ExtraTreesClassifier(n_estimators=200, random_state=42, n_jobs=-1)
XGB = xgboost.XGBClassifier(n_estimators=150, n_jobs=-1)
GBM = lightgbm.LGBMClassifier(objective='binary', n_estimators= 500) # multiclass
list_of_CLFs_names = []
list_of_CLFs = [ RFC, ETC, XGB, GBM]
ranking = []
for clf in list_of_CLFs:
_ = clf.fit(X_train,y_train_copy)
pred = clf.score(X_test,y_test_copy)
name = str(type(clf)).split(".")[-1][:-2]
print("Acc: %0.5f for the %s" % (pred, name))
ranking.append(pred)
list_of_CLFs_names.append(name)
# + id="JsVrj5vnPxVk"
# define several lists and dataframe to store the CV results and evaluation results on testing data
model_names_list = []
cv_fit_time_mean_list = []
cv_accuracy_mean_list = []
cv_precision_mean_list = []
cv_recall_mean_list = []
cv_f1_mean_list = []
cv_roc_auc_mean_list = []
test_accuracy_list = []
test_precision_list = []
test_recall_list = []
test_f1_list = []
test_roc_auc_list = []
test_roc_curve_df = pd.DataFrame()
for model_name, clf in models:
# ==== Step 1: Cross-validation =====
# define Stratified 5-fold cross-validator
cv = StratifiedKFold(n_splits=5, shuffle=True, random_state=123)
# define metrics for evaluating
scoring = ['accuracy', 'precision', 'recall', 'f1', 'roc_auc']
# perform the 5-fold CV and get the metrics results
cv_results = cross_validate(estimator=clf,
X=X_train,
y=y_train,
scoring=scoring,
cv=cv,
return_train_score=False) # prevent to show the train scores on cv splits.
# calculate the mean values of those scores
cv_fit_time_mean = cv_results['fit_time'].mean()
cv_accuracy_mean = cv_results['test_accuracy'].mean()
cv_precision_mean = cv_results['test_precision'].mean()
cv_recall_mean = cv_results['test_recall'].mean()
cv_f1_mean = cv_results['test_f1'].mean()
cv_roc_auc_mean = cv_results['test_roc_auc'].mean()
# store CV results into those lists
model_names_list.append(model_name)
cv_fit_time_mean_list.append(cv_fit_time_mean)
cv_accuracy_mean_list.append(cv_accuracy_mean)
cv_precision_mean_list.append(cv_precision_mean)
cv_recall_mean_list.append(cv_recall_mean)
cv_f1_mean_list.append(cv_f1_mean)
cv_roc_auc_mean_list.append(cv_roc_auc_mean)
# ==== Step 2: Evaluation on Testing data =====
# fit model
clf.fit(X=X_train, y=y_train)
# predition on testing data
# predicted label or class
y_pred_class = clf.predict(X=X_test)
# predicted probability of the label 1
y_pred_score = clf.predict_proba(X=X_test)[:, 1]
# accuracy
accuracy_ontest = accuracy_score(y_true=y_test, y_pred=y_pred_class)
# auc of ROC
auc_ontest = roc_auc_score(y_true=y_test, y_score=y_pred_score)
# precision score
precision_ontest = precision_score(y_true=y_test, y_pred=y_pred_class)
# recall score
recall_ontest = recall_score(y_true=y_test, y_pred=y_pred_class)
# F1 score
f1_ontest = f1_score(y_true=y_test, y_pred=y_pred_class)
# roc curve dataframe
fpr, tpr, threshold_roc = roc_curve(y_true=y_test, y_score=y_pred_score)
roc_df = pd.DataFrame(list(zip(fpr, tpr, threshold_roc)),
columns=['False Positive Rate', 'True Positive Rate', 'Threshold'])
roc_df['Model'] = '{} (AUC = {:.3f})'.format(model_name, auc_ontest)
# store the above values
test_accuracy_list.append(accuracy_ontest)
test_roc_auc_list.append(auc_ontest)
test_precision_list.append(precision_ontest)
test_recall_list.append(recall_ontest)
test_f1_list.append(f1_ontest)
test_roc_curve_df = pd.concat([test_roc_curve_df, roc_df],
ignore_index=True)
# + [markdown] id="Pt50AFpEPxVl"
# ### Model Comparison
#
# We've stored CV results and evaluation results of testing data for the four ML models. Then, we could create a dataframe to view them.
# + id="X82EJ0JtPxVm"
results_dict = {'Model Name': model_names_list,
'CV Fit Time': cv_fit_time_mean_list,
'CV Accuracy mean': cv_accuracy_mean_list,
'CV Precision mean': cv_precision_mean_list,
'CV Recall mean': cv_recall_mean_list,
'CV F1 mean': cv_f1_mean_list,
'CV AUC mean': cv_roc_auc_mean_list,
'Test Accuracy': test_accuracy_list,
'Test Precision': test_precision_list,
'Test Recall': test_recall_list,
'Test F1': test_f1_list,
'Test AUC': test_roc_auc_list
}
results_df = pd.DataFrame(results_dict)
# sort the results according to F1 score on testing data
results_df.sort_values(by='Test F1', ascending=False)
# + [markdown] id="C1JhJnrXNdc5"
# <h1>Choice of the Best algorithm</h1>
# + colab={"base_uri": "https://localhost:8080/"} id="RxTcnYgdNbrb" outputId="1c17f8ac-33e7-4a1e-ec3b-173c341a36db"
RFC = RandomForestClassifier(n_estimators=150, random_state=42, n_jobs=-1)
RFC.fit(X_train,y_train_copy)
# + [markdown] id="chXSRiY4NwXH"
# <h1>Testing the performance of the Best algorithm on test dataset</h1>
# + id="IdwwWpW5N66h"
X_test
# + colab={"base_uri": "https://localhost:8080/"} id="Gndf6YQqNs5o" outputId="708d7dd0-b587-4023-cf2b-1f2a503dc38f"
#predict the class label with the selected algorithm
New_X_test = X_test
New_X_test['sign_deciscion']= RFC.predict(New_X_test)
New_X_test.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="BvZql08iPaIB" outputId="a7880782-6b86-4832-b98d-fe6822fc22d6"
#store the predicted lable
deciscion = pd.DataFrame(New_X_test['sign_deciscion'])
deciscion
# + id="U50FCsCmQvwg" colab={"base_uri": "https://localhost:8080/", "height": 224} outputId="21346680-7c14-47af-8080-0c61cf1fc6e9"
#Remove the added column
X_test = X_test.drop(['sign_deciscion'], axis=1)
X_test.head()
# + [markdown] id="f0CzwHRexdin"
# <h1>Under sampling for Anomaly detection</h1>
# + id="RlCFfE26xvW0" outputId="013e51aa-5699-4449-8dc5-49b079c51f01" colab={"base_uri": "https://localhost:8080/"}
# !pip install imbalanced-learn
# + id="TPjitoA0x027" outputId="256360a1-4090-4080-c52c-8b0f887a47d8" colab={"base_uri": "https://localhost:8080/"}
import imblearn
print(imblearn.__version__)
# + id="EaGkyKOFxcyN" outputId="ce8ea813-6ade-44dc-8da3-fc3481d2ddb4" colab={"base_uri": "https://localhost:8080/"}
# example of random oversampling to balance the class distribution
from collections import Counter
from sklearn.datasets import make_classification
from imblearn.over_sampling import RandomOverSampler
from imblearn.under_sampling import RandomUnderSampler
# define dataset
X, y = make_classification(n_samples=10000, weights=[0.99], flip_y=0)
# summarize class distribution
print(Counter(y))
# + id="Xv2uesBWFvb7"
# from imblearn.combine import SMOTETomek
# smt = SMOTETomek(ratio='auto')
# X_smt, y_smt = smt.fit_sample(X, y)
# + id="Dxufk8rnF4xF" outputId="ef16d1d5-7281-4934-df4a-0d9214048ee4" colab={"base_uri": "https://localhost:8080/"}
print(Counter(y_smt))
# + id="LVDzYerK09Lq" outputId="2088f9fd-573e-418a-a251-f67b20c1150f" colab={"base_uri": "https://localhost:8080/"}
y.shape
# + id="d-YO70wZ03vL" outputId="920b81e1-5b08-4e6e-e326-13b33ccf3858" colab={"base_uri": "https://localhost:8080/"}
# define oversampling strategy
undersample = RandomUnderSampler(sampling_strategy=0.97)
# fit and apply the transform
X_over, y_over = undersample.fit_resample(X_train, y_train)
# summarize class distribution
#print(Counter(y_over))
# X= X_over
type(y_over)
# + id="NycpLS93Hdmm"
# define oversampling strategy
undersample = RandomUnderSampler(sampling_strategy=0.75)
# fit and apply the transform
X_over, y_over = undersample.fit_resample(X_over, y_over)
# summarize class distribution
#print(Counter(y_over))
# X= X_over
type(y_over)
# + id="fLwKOP5aA8-a"
import numpy
# + id="DZXrZpfZA0Cu" outputId="893d6910-091a-4d45-89b1-f0d2c374470d" colab={"base_uri": "https://localhost:8080/"}
unique, counts = numpy.unique(y_over, return_counts=True)
dict(zip(unique, counts))
# + id="SEVMHXPm7XYh"
X_over
# + id="nU5JDgdi2kHi"
# define oversampling strategy
over = RandomOverSampler(sampling_strategy='minority')
# fit and apply the transform
X_over, y_over = over.fit_resample(X, y)
# summarize class distribution
print(Counter(y))
# + [markdown] id="RC9jjcYUS8cP"
# <h1>Unsupervied ML (Anomaly)</h1>
# + id="6whmKwOzVZD4"
# %%capture
# !pip install pyod
# !pip install hdbscan
# !pip install combo
# !pip install SOM...... # https://github.com/AICoE/log-anomaly-detector/blob/master/Notebooks/SOM_retrain_notebook/SOM_UserFeedbackPOC.ipynb
# !pip install somtf
# + colab={"base_uri": "https://localhost:8080/"} id="tO9a5CgdVcDv" outputId="67e933d4-f512-435d-a6dd-d23c26e83faa"
# !pip install rrcf
# + id="gtZJY7hzVqDj"
import gc, os, pickle
from datetime import datetime
import numpy as np
import pandas as pd
from collections import Counter
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
import warnings
warnings.simplefilter(action='ignore', category=FutureWarning)
import pyod
from hdbscan import HDBSCAN
import rrcf
import sklearn
from sklearn.model_selection import train_test_split
from sklearn.decomposition import *
from sklearn.preprocessing import *
from sklearn.svm import *
from sklearn.metrics import precision_recall_curve, precision_score, recall_score, f1_score, accuracy_score
from sklearn.metrics import roc_curve, auc, roc_auc_score, confusion_matrix, classification_report
# + id="qwKMr_tQVtLQ"
from pyod.models import lof, cblof, cof, pca, iforest, knn, mcd, ocsvm, sod, abod, hbos #, auto_encoder, vae
contamination = 0.4
threshold = 0.75
# + [markdown] id="aTD47EFqVxqL"
# <table class="docutils align-default">
# <colgroup>
# <col style="width: 8%">
# <col style="width: 6%">
# <col style="width: 41%">
# <col style="width: 2%">
# <col style="width: 21%">
# <col style="width: 22%">
# </colgroup>
# <thead>
# <tr class="row-odd"><th class="head"><p>Type</p></th>
# <th class="head"><p>Abbr</p></th>
# <th class="head"><p>Algorithm</p></th>
# <th class="head"><p>Year</p></th>
# <th class="head"><p>Class</p></th>
# <th class="head"><p>Ref</p></th>
# </tr>
# </thead>
# <tbody>
# <tr class="row-even"><td><p>Linear Model</p></td>
# <td><p>PCA</p></td>
# <td><p>Principal Component Analysis (the sum of weighted projected distances to the eigenvector hyperplanes)</p></td>
# <td><p>2003</p></td>
# <td><p><a class="reference internal" href="pyod.models.html#pyod.models.pca.PCA" title="pyod.models.pca.PCA"><code class="xref py py-class docutils literal notranslate"><span class="pre">pyod.models.pca.PCA</span></code></a></p></td>
# <td><p><span id="id3">[<a class="reference internal" href="#id53"><span>ASCSC03</span></a>]</span></p></td>
# </tr>
# <tr class="row-odd"><td><p>Linear Model</p></td>
# <td><p>MCD</p></td>
# <td><p>Minimum Covariance Determinant (use the mahalanobis distances as the outlier scores)</p></td>
# <td><p>1999</p></td>
# <td><p><a class="reference internal" href="pyod.models.html#pyod.models.mcd.MCD" title="pyod.models.mcd.MCD"><code class="xref py py-class docutils literal notranslate"><span class="pre">pyod.models.mcd.MCD</span></code></a></p></td>
# <td><p><span id="id4">[<a class="reference internal" href="#id57"><span>ARD99</span></a>,<a class="reference internal" href="#id58"><span>AHR04</span></a>]</span></p></td>
# </tr>
# <tr class="row-even"><td><p>Linear Model</p></td>
# <td><p>OCSVM</p></td>
# <td><p>One-Class Support Vector Machines</p></td>
# <td><p>2001</p></td>
# <td><p><a class="reference internal" href="pyod.models.html#pyod.models.ocsvm.OCSVM" title="pyod.models.ocsvm.OCSVM"><code class="xref py py-class docutils literal notranslate"><span class="pre">pyod.models.ocsvm.OCSVM</span></code></a></p></td>
# <td><p><span id="id5">[<a class="reference internal" href="#id68"><span>AScholkopfPST+01</span></a>]</span></p></td>
# </tr>
# <tr class="row-odd"><td><p>Linear Model</p></td>
# <td><p>LMDD</p></td>
# <td><p>Deviation-based Outlier Detection (LMDD)</p></td>
# <td><p>1996</p></td>
# <td><p><a class="reference internal" href="pyod.models.html#pyod.models.lmdd.LMDD" title="pyod.models.lmdd.LMDD"><code class="xref py py-class docutils literal notranslate"><span class="pre">pyod.models.lmdd.LMDD</span></code></a></p></td>
# <td><p><span id="id6">[<a class="reference internal" href="#id75"><span>AAAR96</span></a>]</span></p></td>
# </tr>
# <tr class="row-even"><td><p>Proximity-Based</p></td>
# <td><p>LOF</p></td>
# <td><p>Local Outlier Factor</p></td>
# <td><p>2000</p></td>
# <td><p><a class="reference internal" href="pyod.models.html#pyod.models.lof.LOF" title="pyod.models.lof.LOF"><code class="xref py py-class docutils literal notranslate"><span class="pre">pyod.models.lof.LOF</span></code></a></p></td>
# <td><p><span id="id7">[<a class="reference internal" href="#id55"><span>ABKNS00</span></a>]</span></p></td>
# </tr>
# <tr class="row-odd"><td><p>Proximity-Based</p></td>
# <td><p>COF</p></td>
# <td><p>Connectivity-Based Outlier Factor</p></td>
# <td><p>2002</p></td>
# <td><p><a class="reference internal" href="pyod.models.html#pyod.models.cof.COF" title="pyod.models.cof.COF"><code class="xref py py-class docutils literal notranslate"><span class="pre">pyod.models.cof.COF</span></code></a></p></td>
# <td><p><span id="id8">[<a class="reference internal" href="#id69"><span>ATCFC02</span></a>]</span></p></td>
# </tr>
# <tr class="row-even"><td><p>Proximity-Based</p></td>
# <td><p>CBLOF</p></td>
# <td><p>Clustering-Based Local Outlier Factor</p></td>
# <td><p>2003</p></td>
# <td><p><a class="reference internal" href="pyod.models.html#pyod.models.cblof.CBLOF" title="pyod.models.cblof.CBLOF"><code class="xref py py-class docutils literal notranslate"><span class="pre">pyod.models.cblof.CBLOF</span></code></a></p></td>
# <td><p><span id="id9">[<a class="reference internal" href="#id59"><span>AHXD03</span></a>]</span></p></td>
# </tr>
# <tr class="row-odd"><td><p>Proximity-Based</p></td>
# <td><p>LOCI</p></td>
# <td><p>LOCI: Fast outlier detection using the local correlation integral</p></td>
# <td><p>2003</p></td>
# <td><p><a class="reference internal" href="pyod.models.html#pyod.models.loci.LOCI" title="pyod.models.loci.LOCI"><code class="xref py py-class docutils literal notranslate"><span class="pre">pyod.models.loci.LOCI</span></code></a></p></td>
# <td><p><span id="id10">[<a class="reference internal" href="#id62"><span>APKGF03</span></a>]</span></p></td>
# </tr>
# <tr class="row-even"><td><p>Proximity-Based</p></td>
# <td><p>HBOS</p></td>
# <td><p>Histogram-based Outlier Score</p></td>
# <td><p>2012</p></td>
# <td><p><a class="reference internal" href="pyod.models.html#pyod.models.hbos.HBOS" title="pyod.models.hbos.HBOS"><code class="xref py py-class docutils literal notranslate"><span class="pre">pyod.models.hbos.HBOS</span></code></a></p></td>
# <td><p><span id="id11">[<a class="reference internal" href="#id52"><span>AGD12</span></a>]</span></p></td>
# </tr>
# <tr class="row-odd"><td><p>Proximity-Based</p></td>
# <td><p>kNN</p></td>
# <td><p>k Nearest Neighbors (use the distance to the kth nearest neighbor as the outlier score</p></td>
# <td><p>2000</p></td>
# <td><p><a class="reference internal" href="pyod.models.html#pyod.models.knn.KNN" title="pyod.models.knn.KNN"><code class="xref py py-class docutils literal notranslate"><span class="pre">pyod.models.knn.KNN</span></code></a></p></td>
# <td><p><span id="id12">[<a class="reference internal" href="#id48"><span>ARRS00</span></a>,<a class="reference internal" href="#id49"><span>AAP02</span></a>]</span></p></td>
# </tr>
# <tr class="row-even"><td><p>Proximity-Based</p></td>
# <td><p>AvgKNN</p></td>
# <td><p>Average kNN (use the average distance to k nearest neighbors as the outlier score)</p></td>
# <td><p>2002</p></td>
# <td><p><a class="reference internal" href="pyod.models.html#pyod.models.knn.KNN" title="pyod.models.knn.KNN"><code class="xref py py-class docutils literal notranslate"><span class="pre">pyod.models.knn.KNN</span></code></a></p></td>
# <td><p><span id="id13">[<a class="reference internal" href="#id48"><span>ARRS00</span></a>,<a class="reference internal" href="#id49"><span>AAP02</span></a>]</span></p></td>
# </tr>
# <tr class="row-odd"><td><p>Proximity-Based</p></td>
# <td><p>MedKNN</p></td>
# <td><p>Median kNN (use the median distance to k nearest neighbors as the outlier score)</p></td>
# <td><p>2002</p></td>
# <td><p><a class="reference internal" href="pyod.models.html#pyod.models.knn.KNN" title="pyod.models.knn.KNN"><code class="xref py py-class docutils literal notranslate"><span class="pre">pyod.models.knn.KNN</span></code></a></p></td>
# <td><p><span id="id14">[<a class="reference internal" href="#id48"><span>ARRS00</span></a>,<a class="reference internal" href="#id49"><span>AAP02</span></a>]</span></p></td>
# </tr>
# <tr class="row-even"><td><p>Proximity-Based</p></td>
# <td><p>SOD</p></td>
# <td><p>Subspace Outlier Detection</p></td>
# <td><p>2009</p></td>
# <td><p><a class="reference internal" href="pyod.models.html#pyod.models.sod.SOD" title="pyod.models.sod.SOD"><code class="xref py py-class docutils literal notranslate"><span class="pre">pyod.models.sod.SOD</span></code></a></p></td>
# <td><p><span id="id15">[<a class="reference internal" href="#id71"><span>AKKrogerSZ09</span></a>]</span></p></td>
# </tr>
# <tr class="row-odd"><td><p>Proximity-Based</p></td>
# <td><p>ROD</p></td>
# <td><p>Rotation-based Outlier Detection</p></td>
# <td><p>2020</p></td>
# <td><p><a class="reference internal" href="pyod.models.html#pyod.models.rod.ROD" title="pyod.models.rod.ROD"><code class="xref py py-class docutils literal notranslate"><span class="pre">pyod.models.rod.ROD</span></code></a></p></td>
# <td><p><span id="id16">[<a class="reference internal" href="#id81"><span>AABC20</span></a>]</span></p></td>
# </tr>
# <tr class="row-even"><td><p>Probabilistic</p></td>
# <td><p>ABOD</p></td>
# <td><p>Angle-Based Outlier Detection</p></td>
# <td><p>2008</p></td>
# <td><p><a class="reference internal" href="pyod.models.html#pyod.models.abod.ABOD" title="pyod.models.abod.ABOD"><code class="xref py py-class docutils literal notranslate"><span class="pre">pyod.models.abod.ABOD</span></code></a></p></td>
# <td><p><span id="id17">[<a class="reference internal" href="#id50"><span>AKZ+08</span></a>]</span></p></td>
# </tr>
# <tr class="row-odd"><td><p>Probabilistic</p></td>
# <td><p>FastABOD</p></td>
# <td><p>Fast Angle-Based Outlier Detection using approximation</p></td>
# <td><p>2008</p></td>
# <td><p><a class="reference internal" href="pyod.models.html#pyod.models.abod.ABOD" title="pyod.models.abod.ABOD"><code class="xref py py-class docutils literal notranslate"><span class="pre">pyod.models.abod.ABOD</span></code></a></p></td>
# <td><p><span id="id18">[<a class="reference internal" href="#id50"><span>AKZ+08</span></a>]</span></p></td>
# </tr>
# <tr class="row-even"><td><p>Probabilistic</p></td>
# <td><p>COPOD</p></td>
# <td><p>COPOD: Copula-Based Outlier Detection</p></td>
# <td><p>2020</p></td>
# <td><p><a class="reference internal" href="pyod.models.html#pyod.models.copod.COPOD" title="pyod.models.copod.COPOD"><code class="xref py py-class docutils literal notranslate"><span class="pre">pyod.models.copod.COPOD</span></code></a></p></td>
# <td><p><span id="id19">[<a class="reference internal" href="#id80"><span>ALZB+20</span></a>]</span></p></td>
# </tr>
# <tr class="row-odd"><td><p>Probabilistic</p></td>
# <td><p>MAD</p></td>
# <td><p>Median Absolute Deviation (MAD)</p></td>
# <td><p>1993</p></td>
# <td><p><a class="reference internal" href="pyod.models.html#pyod.models.mad.MAD" title="pyod.models.mad.MAD"><code class="xref py py-class docutils literal notranslate"><span class="pre">pyod.models.mad.MAD</span></code></a></p></td>
# <td><p><span id="id20">[<a class="reference internal" href="#id79"><span>AIH93</span></a>]</span></p></td>
# </tr>
# <tr class="row-even"><td><p>Probabilistic</p></td>
# <td><p>SOS</p></td>
# <td><p>Stochastic Outlier Selection</p></td>
# <td><p>2012</p></td>
# <td><p><a class="reference internal" href="pyod.models.html#pyod.models.sos.SOS" title="pyod.models.sos.SOS"><code class="xref py py-class docutils literal notranslate"><span class="pre">pyod.models.sos.SOS</span></code></a></p></td>
# <td><p><span id="id21">[<a class="reference internal" href="#id61"><span>AJHuszarPvdH12</span></a>]</span></p></td>
# </tr>
# <tr class="row-odd"><td><p>Outlier Ensembles</p></td>
# <td><p>IForest</p></td>
# <td><p>Isolation Forest</p></td>
# <td><p>2008</p></td>
# <td><p><a class="reference internal" href="pyod.models.html#pyod.models.iforest.IForest" title="pyod.models.iforest.IForest"><code class="xref py py-class docutils literal notranslate"><span class="pre">pyod.models.iforest.IForest</span></code></a></p></td>
# <td><p><span id="id22">[<a class="reference internal" href="#id44"><span>ALTZ08</span></a>,<a class="reference internal" href="#id45"><span>ALTZ12</span></a>]</span></p></td>
# </tr>
# <tr class="row-even"><td><p>Outlier Ensembles</p></td>
# <td></td>
# <td><p>Feature Bagging</p></td>
# <td><p>2005</p></td>
# <td><p><a class="reference internal" href="pyod.models.html#pyod.models.feature_bagging.FeatureBagging" title="pyod.models.feature_bagging.FeatureBagging"><code class="xref py py-class docutils literal notranslate"><span class="pre">pyod.models.feature_bagging.FeatureBagging</span></code></a></p></td>
# <td><p><span id="id23">[<a class="reference internal" href="#id51"><span>ALK05</span></a>]</span></p></td>
# </tr>
# <tr class="row-odd"><td><p>Outlier Ensembles</p></td>
# <td><p>LSCP</p></td>
# <td><p>LSCP: Locally Selective Combination of Parallel Outlier Ensembles</p></td>
# <td><p>2019</p></td>
# <td><p><a class="reference internal" href="pyod.models.html#pyod.models.lscp.LSCP" title="pyod.models.lscp.LSCP"><code class="xref py py-class docutils literal notranslate"><span class="pre">pyod.models.lscp.LSCP</span></code></a></p></td>
# <td><p><span id="id24">[<a class="reference internal" href="#id63"><span>AZNHL19</span></a>]</span></p></td>
# </tr>
# <tr class="row-even"><td><p>Outlier Ensembles</p></td>
# <td><p>XGBOD</p></td>
# <td><p>Extreme Boosting Based Outlier Detection <strong>(Supervised)</strong></p></td>
# <td><p>2018</p></td>
# <td><p><a class="reference internal" href="pyod.models.html#pyod.models.xgbod.XGBOD" title="pyod.models.xgbod.XGBOD"><code class="xref py py-class docutils literal notranslate"><span class="pre">pyod.models.xgbod.XGBOD</span></code></a></p></td>
# <td><p><span id="id25">[<a class="reference internal" href="#id56"><span>AZH18</span></a>]</span></p></td>
# </tr>
# <tr class="row-odd"><td><p>Outlier Ensembles</p></td>
# <td><p>LODA</p></td>
# <td><p>Lightweight On-line Detector of Anomalies</p></td>
# <td><p>2016</p></td>
# <td><p><a class="reference internal" href="pyod.models.html#pyod.models.loda.LODA" title="pyod.models.loda.LODA"><code class="xref py py-class docutils literal notranslate"><span class="pre">pyod.models.loda.LODA</span></code></a></p></td>
# <td><p><span id="id26">[<a class="reference internal" href="#id77"><span>APevny16</span></a>]</span></p></td>
# </tr>
# <tr class="row-even"><td><p>Neural Networks</p></td>
# <td><p>AutoEncoder</p></td>
# <td><p>Fully connected AutoEncoder (use reconstruction error as the outlier score)</p></td>
# <td><p>2015</p></td>
# <td><p><a class="reference internal" href="pyod.models.html#pyod.models.auto_encoder.AutoEncoder" title="pyod.models.auto_encoder.AutoEncoder"><code class="xref py py-class docutils literal notranslate"><span class="pre">pyod.models.auto_encoder.AutoEncoder</span></code></a></p></td>
# <td><p><span id="id27">[<a class="reference internal" href="#id54"><span>AAgg15</span></a>]</span></p></td>
# </tr>
# <tr class="row-odd"><td><p>Neural Networks</p></td>
# <td><p>VAE</p></td>
# <td><p>Variational AutoEncoder (use reconstruction error as the outlier score)</p></td>
# <td><p>2013</p></td>
# <td><p><a class="reference internal" href="pyod.models.html#pyod.models.vae.VAE" title="pyod.models.vae.VAE"><code class="xref py py-class docutils literal notranslate"><span class="pre">pyod.models.vae.VAE</span></code></a></p></td>
# <td><p><span id="id28">[<a class="reference internal" href="#id76"><span>AKW13</span></a>]</span></p></td>
# </tr>
# <tr class="row-even"><td><p>Neural Networks</p></td>
# <td><p>Beta-VAE</p></td>
# <td><p>Variational AutoEncoder (all customized loss term by varying gamma and capacity)</p></td>
# <td><p>2018</p></td>
# <td><p><a class="reference internal" href="pyod.models.html#pyod.models.vae.VAE" title="pyod.models.vae.VAE"><code class="xref py py-class docutils literal notranslate"><span class="pre">pyod.models.vae.VAE</span></code></a></p></td>
# <td><p><span id="id29">[<a class="reference internal" href="#id78"><span>ABHP+18</span></a>]</span></p></td>
# </tr>
# <tr class="row-odd"><td><p>Neural Networks</p></td>
# <td><p>SO_GAAL</p></td>
# <td><p>Single-Objective Generative Adversarial Active Learning</p></td>
# <td><p>2019</p></td>
# <td><p><a class="reference internal" href="pyod.models.html#pyod.models.so_gaal.SO_GAAL" title="pyod.models.so_gaal.SO_GAAL"><code class="xref py py-class docutils literal notranslate"><span class="pre">pyod.models.so_gaal.SO_GAAL</span></code></a></p></td>
# <td><p><span id="id30">[<a class="reference internal" href="#id64"><span>ALLZ+19</span></a>]</span></p></td>
# </tr>
# <tr class="row-even"><td><p>Neural Networks</p></td>
# <td><p>MO_GAAL</p></td>
# <td><p>Multiple-Objective Generative Adversarial Active Learning</p></td>
# <td><p>2019</p></td>
# <td><p><a class="reference internal" href="pyod.models.html#pyod.models.mo_gaal.MO_GAAL" title="pyod.models.mo_gaal.MO_GAAL"><code class="xref py py-class docutils literal notranslate"><span class="pre">pyod.models.mo_gaal.MO_GAAL</span></code></a></p></td>
# <td><p><span id="id31">[<a class="reference internal" href="#id64"><span>ALLZ+19</span></a>]</span></p></td>
# </tr>
# </tbody>
# </table>
# + [markdown] id="uflPBRzzWA44"
# ## Uses PYOD
# + [markdown] id="t4v_RIyKWGD8"
# <h1><b>Printing Score metrics</b></h1>
# + id="XkogpL4SV6SO"
def PrintScoreMetrics(algorithm,X_test,y_test):
y_pred_class = algorithm.predict(X_test)
y_pred_score = algorithm.predict_proba(X=X_test)[:, 1]
# AUC of ROC
auc_ontest = roc_auc_score(y_true=y_test, y_score=y_pred_score)
# confusion matrix
cm_ontest = confusion_matrix(y_true=y_test, y_pred=y_pred_class)
# precision score
precision_ontest = precision_score(y_true=y_test, y_pred=y_pred_class)
# recall score
recall_ontest = recall_score(y_true=y_test, y_pred=y_pred_class)
# classifition report
cls_report_ontest = classification_report(y_true=y_test, y_pred=y_pred_class)
# print the above results
print('The model scores {:1.5f} ROC AUC on the test set.'.format(auc_ontest))
print('The precision score on the test set: {:1.5f}'.format(precision_ontest))
print('The recall score on the test set: {:1.5f}'.format(recall_ontest))
print('Confusion Matrix:\n', cm_ontest)
# Print classification report:
print('Classification Report:\n', cls_report_ontest)
# + [markdown] id="rZv1cX_6V5fo"
#
# + [markdown] id="WdnDSH_1mDHr"
# <h1><b>the LOF model</b><h1>
# + id="XpSg_uGZjR9J"
#===============TRAIN THE LOF MODEL ======================
lof_clf = lof.LOF(contamination=contamination, n_jobs=-1)
_ = lof_clf.fit(X_train)
# + id="sEC0BtSsM76E" colab={"base_uri": "https://localhost:8080/"} outputId="b3919116-5b29-4ce6-ee6f-6a0e4079d64d"
#======= EVALUATE LOF MODEL =================
PrintScoreMetrics(lof_clf,X_test,y_test)
# + [markdown] id="WWw2wUPmlcva"
# <h1><b>the CBLOF model</b><h1>
# + id="9HMfAuoOjSAq"
#========== TRAIN CBLOF MODEL ==========
cblof_clf = cblof.CBLOF(contamination=contamination, n_jobs=-1, n_clusters=45)
_ = cblof_clf.fit(X_train)
# + id="OMIRzSwffHFB" colab={"base_uri": "https://localhost:8080/"} outputId="04cdc8d5-3311-48a3-c889-3f521c98c856"
#======= EVALUATE CBLOF MODEL =================
PrintScoreMetrics(cblof_clf,X_test,y_test)
# + [markdown] id="0KtZja8qohBq"
# <h1><b>the Connectivity-Based Outlier Factor (cof)</b><h1>
# + id="o3yKHghIjSEV"
#========== TRAIN COF MODEL ==========
cof_clf = cof.COF(contamination=contamination)
_ = cof_clf.fit(X_train[:5000])
# + id="GV3nL4qzoeIL"
#======= EVALUATE COF MODEL =================
PrintScoreMetrics(cof_clf,X_test,y_test)
# + [markdown] id="c5tr85k4vq0F"
# <h1><b>The PCA MODEL</b><h1>
# + id="ixQ7uR00jSIB"
# https://pyod.readthedocs.io/en/latest/_modules/pyod/models/pca.html
#=============TRAINING PCA MODEL=========================
pca_clf = pca.PCA(contamination=contamination)
_ = pca_clf.fit(X_train)
# + id="0GamMfdVwPXC"
#======= EVALUATE PCA MODEL =================
PrintScoreMetrics(pca_clf,X_test,y_test)
# + id="k5lR1t6u-aut"
pca_clf = pca.PCA(2, contamination=contamination)
_ = pca_clf.fit(X_train)
PrintScoreMetrics(pca_clf,X_test,y_test)
# + [markdown] id="A6AEdW_MxT9t"
# <h1><b>The IFOREST MODEL</b><h1>
# + id="4OHfbx1qjSLS"
iforest_clf = iforest.IForest(contamination=contamination, n_estimators=300, max_samples= 1028, n_jobs=-1)
_ = iforest_clf.fit(X_train)
# + id="jRVZrEo81SDY"
X_test.head(3)
# + id="a1xa5_vVyPsm"
#======= EVALUATE IFOREST MODEL =================
PrintScoreMetrics(iforest_clf,X_test,y_test)
# + id="GPkBQtQ6RTst"
Anom_X_test = X_test
#Anom_X_test['scores']=iforest_clf.decision_function(X_test)
Anom_X_test['anom_deciscion']=lof_clf.predict(X_test)
Anom_X_test.head(5)
# + id="LxL0MDPZpxQe"
deciscion['anom_decision'] = pd.DataFrame(Anom_X_test['anom_deciscion'])
# + id="0hq96QqcjwHd"
Final_decision = pd.DataFrame(columns=['Output'])
# + id="QLrGPgekq4QC"
Final_decision.head(3)
# + id="_M8gkmtUo9rE"
deciscion.head(20)
# + [markdown] id="tsPXPt4i9Zok"
# <h1>Find the accuracy of our model</h1>
# + id="vz8zsvgk9e1t"
for index, row in deciscion.iterrows():
if((row['anom_decision']==1) and (row['sign_deciscion']== 1)):
Final_decision.loc[index] = 1
elif((row['anom_decision']==0) and (row['sign_deciscion']== 0)):
Final_decision.loc[index] = 0
elif((row['anom_decision']==1) and (row['sign_deciscion']== 0)):
Final_decision.loc[index] = 1
else:
Final_decision.loc[index] = -1
# + id="qEZmijyu90wF"
#Final_decision.head(20)
pred_attack = Final_decision.where(Final_decision['Output'] == 1).dropna()
pred_attack.shape
# + id="IPOgONURwR-S"
true_attack = y_test.where(y_test['label'] == 1).dropna()
true_attack.shape
# + id="_IxdgCPA1XLL"
accuracy = len(pred_attack)/len(true_attack)
print(100* accuracy)
# + [markdown] id="ayEzNKEozXja"
# <h1><b>The KNN MODEL</b><h1>
# + id="twvrr6d5jSOq"
#========== TRAIN KNN MODEL ==========
knn_clf = knn.KNN(contamination=contamination, radius=1.5, n_neighbors=20, n_jobs=-1) #TODO radius
_ = knn_clf.fit(X_train)
# predictions = knn_clf.predict(X_train)
# print(f'Acc of train: {accuracy_score(y_train, predictions)}')
# predictions = knn_clf.predict(X_test)
# print(f'Acc of test: {accuracy_score(y_test, predictions)}')
# + id="qg2ZR9hN0BSC"
#======= EVALUATE KNN MODEL =================
PrintScoreMetrics(knn_clf,X_test,y_test)
# + [markdown] id="7qgYsFJR1Ndc"
# <h1><b>The OCSVM MODEL</b><h1>
# + id="I8gsbiYtjSRk"
#================== TRAIN OCSVM ===============================
ocsvm_clf = ocsvm.OCSVM(contamination=contamination)
_ = ocsvm_clf.fit(X_train[:1000])
# + id="2u3FZihu1eIO"
#======= EVALUATE OCSVM MODEL =================
PrintScoreMetrics(ocsvm_clf,X_test,y_test)
# + [markdown] id="FVMMaa4H2LGi"
# <h1><b>The Angle-Based Outlier Detection (ABOD) MODEL</b><h1>
#
#
# + id="1T_uPrDSjRop"
#================ TRAIN ABOD MODEL =====================
X_temp = X_train.astype(np.float)
abod_clf = abod.ABOD(contamination=contamination, n_neighbors=10, )
_ = abod_clf.fit(X_temp)
# + id="eYYW2Fy-2m-i"
#======= EVALUATE ABOD MODEL =================
PrintScoreMetrics(abod_clf,X_test,y_test)
# + [markdown] id="RBDFquUePxVn"
# #### ROC Curve Comparison
#
# A receiver operating characteristic curve, or ROC curve, is a graphical plot that illustrates the diagnostic ability of a binary classifier system as its discrimination threshold is varied. The ROC curve is created by plotting the true positive rate (TPR) against the false positive rate (FPR) at various threshold settings.
#
# The AUC score or AUC ROC (Area Under the Receiver Operating Characteristic Curve) score represents degree or measure of separability. **Higher the AUC, better the model is at predicting 0s as 0s and 1s as 1s. By analogy, Higher the AUC, better the model is at distinguishing between patients with disease and no disease.**
#
# Here, we show the ROC curves with AUC scores for the four models. We will use a plot package `plotly` to show an interactive figure.
#
# In the above code, we have computed TPR, FPR and threshold values for plotting ROC curves, which have been stored into `test_roc_curve_df` with four columns `False Positive Rate`, `True Positive Rate`, `Threshold`, and `Model`.
#
# + id="pcgcbeioPxVo"
# # !pip install plotly
# # !pip install cufflinks
# + id="as5L1aYtPxVp"
# plotly imports
# import plotly.express as px
# import plotly.graph_objects as go
# + id="O5xFWgq2PxVp"
# ROC_fig = px.line(test_roc_curve_df,
# x='False Positive Rate',
# y='True Positive Rate',
# color='Model',
# hover_data=['Threshold'])
# ROC_fig.update_layout(
# legend=go.layout.Legend(
# x=0.5,
# y=0.1,
# traceorder="normal",
# font=dict(
# # family="sans-serif",
# size=9,
# color="black"
# ),
# bgcolor="LightSteelBlue",
# bordercolor="Black",
# borderwidth=2
# ),
# title=go.layout.Title(text="ROC Curve on Hold-out Testing Dataset",
# xref="paper",
# x=0
# ),
# xaxis=go.layout.XAxis(
# title=go.layout.xaxis.Title(
# text="False Positive Rate"
# )
# ),
# yaxis=go.layout.YAxis(
# title=go.layout.yaxis.Title(
# text="True Positive Rate"
# )
# )
# )
# ROC_fig.show()
# + [markdown] id="k6EZft49PxWW"
# # Model Inspection
#
# Predictive performance is often the main goal of developing machine learning models. Yet summarising performance with an evaluation metric is often insufficient: it assumes that the evaluation metric and test dataset perfectly reflect the target domain, which is rarely true.
#
# In certain domains, a model needs a certain level of interpretability before it can be deployed. A model that is exhibiting performance issues needs to be debugged for one to understand the model’s underlying issue.
#
# The `sklearn.inspection` module provides tools to help understand the predictions from a model and what affects them. This can be used to evaluate assumptions and biases of a model, design a better model, or to diagnose issues with model performance.
# + [markdown] id="YmfoVhRnPxWf"
# ## Partial dependence plots (PDPs)
#
# Partial dependence plots (PDPs) show the dependence between the target response $y$ and a set of ‘target’ features $X$, marginalizing over the values of all other features (the ‘complement’ features). Intuitively, we can interpret the partial dependence as the expected target response as a function of the ‘target’ features.
#
# **In general, PDPs show how a feature affects predictions.** If you are familiar with linear regression models, PDPs can be interpreted similarly to the coefficients in those models. Though, PDPs for sophisticated models can capture more complex patterns than coefficients from simple models. We will show a couple examples, explain the interpretation of these plots, and then review the code to create these plots.
#
# ### Example: PDP for LogisticRegression
#
# Let's plot a PDP for LogisticRegression that we have trained before. Assume that we are interested in the effect of all the features on the response `attack_cat`
# + id="wVfqlCkJPxWf"
# import the plot function
from sklearn.inspection import plot_partial_dependence
# + [markdown] id="4YugjvwSPxWg"
# `plot_partial_dependence` function has five important arguments to specify which features are plotted.
#
# - `estimator`: your fitted model/classifier.
# - `X`: your training data
# - `features` : the indexes of your interested features
# - `feature_names` : the list of the features names in your training data
# - `target` : In a multi-class setting, specifies the class for which the PDPs should be computed.
# Note that for binary classification, the positive class (index 1) is always used.
#
# Therefore, we first need to figure out the feature indexes in our training data. Remember that we have transformed our training data such that the orginal features names and orders have been changed.
#
# Let's look at what the current features names and orders in `X_train_transform`. Again, we should call `get_column_names_from_ColumnTransformer` on `col_trans` to get those new names.
# + id="a2n8I41kPxWg"
from sklearn.pipeline import Pipeline
def get_column_names_from_ColumnTransformer(column_transformer):
col_name = []
for transformer_in_columns in column_transformer.transformers_:
raw_col_name = transformer_in_columns[2]
if isinstance(transformer_in_columns[1], Pipeline):
transformer = transformer_in_columns[1].steps[-1][1]
else:
transformer = transformer_in_columns[1]
try:
names = transformer.get_feature_names(input_features=raw_col_name)
except AttributeError: # if no 'get_feature_names' function, use raw column name
if transformer_in_columns[1] == "passthrough":
names = column_transformer._feature_names_in[raw_col_name]
else:
names = raw_col_name
if isinstance(names,np.ndarray):
col_name += names.tolist()
elif isinstance(names,pd.Index):
col_name += names.tolist()
elif isinstance(names,list):
col_name += names
elif isinstance(names,str):
col_name.append(names)
return col_name
# + id="hMVPrLArPxWh" colab={"base_uri": "https://localhost:8080/"} outputId="4a68c7b1-2dc3-4d2a-fe2d-3c8ed337d368"
new_cols = get_column_names_from_ColumnTransformer(col_trans)
new_cols
# + id="DmaP3x6XPxWh"
models[0]
# + id="-bhx5UuxPxWh" colab={"base_uri": "https://localhost:8080/", "height": 737} outputId="002f3cb3-4cf2-4337-a44d-afcac51c6394"
clf_name = models[0][0] # the classifier name
clf = models[0][1] # the classifier itself
features = [0,1,2,3,4,5,6,7,8,9,10,11]
plot_partial_dependence(estimator=clf, X=X_train_transform,
features=features, feature_names=new_cols,
target=1)
fig = plt.gcf() # get current figure (gcf)
fig.suptitle("Partial dependence of Attack or Normal on all features\n"
"with {}".format(clf_name))
fig.set_size_inches(20, 10)
fig.subplots_adjust(hspace=.4, wspace=.25, top=.9)
# + id="ZwPf-AcVPxWj" colab={"base_uri": "https://localhost:8080/", "height": 449} outputId="db3900f6-7cc0-47d8-cfd2-c55ba1ac7da0"
clf_name = models[0][0] # the classifier name
clf = models[0][1] # the classifier itself
features = [12,13,14,15,16,17,18,19,20]
plot_partial_dependence(estimator=clf, X=X_train_transform,
features=features, feature_names=new_cols,
target=1)
fig = plt.gcf() # get current figure (gcf)
fig.suptitle("Partial dependence of Attack or Normal on all features\n"
"with {}".format(clf_name))
fig.set_size_inches(20, 10)
fig.subplots_adjust(hspace=.4, wspace=.5, top=.9)
# + id="yYUcdpyTPxWk" colab={"base_uri": "https://localhost:8080/", "height": 449} outputId="1ad63676-f6e7-4b33-fad4-a57c6a068540"
clf_name = models[0][0] # the classifier name
clf = models[0][1] # the classifier itself
features = [21,22,23,24,25,26,27,28,29]
plot_partial_dependence(estimator=clf, X=X_train_transform,
features=features, feature_names=new_cols,
target=1)
fig = plt.gcf() # get current figure (gcf)
fig.suptitle("Partial dependence of Attack or Normal on all features\n"
"with {}".format(clf_name))
fig.set_size_inches(20, 10)
fig.subplots_adjust(hspace=.4, wspace=.25, top=.9)
# + id="nTpWH4kHPxWk" colab={"base_uri": "https://localhost:8080/", "height": 449} outputId="289b371b-d937-4a13-ddd8-d8ebf53994bd"
clf_name = models[0][0] # the classifier name
clf = models[0][1] # the classifier itself
features = [30,31,32,33,34,35,36,37,38]
plot_partial_dependence(estimator=clf, X=X_train_transform,
features=features, feature_names=new_cols,
target=1)
fig = plt.gcf() # get current figure (gcf)
fig.suptitle("Partial dependence of Attack or Normal on all features\n"
"with {}".format(clf_name))
fig.set_size_inches(20, 10)
fig.subplots_adjust(hspace=.4, wspace=.35, top=0.9)
# + [markdown] id="vw7oLPjLPxWm"
# We could also plot a 2-way PDP, for example, visualizing the effect of the joint of `swin` and `ct_src_dport_itm` on the response by specifying a `tuple` that contains the indexes of them.
# + id="p4vcXX4PPxWm" outputId="9dced4c6-f6bd-4c85-95bc-bd32f1eb7e0f"
features = [(7, 16)]
plot_partial_dependence(estimator=clf, X=X_train_transform,
features=features, feature_names=new_cols,
target=1)
fig = plt.gcf() # get current figure (gcf)
fig.suptitle("Partial dependence of Attack or Normal \n"
"with {}".format(clf_name))
fig.set_size_inches(7.5, 6.5)
fig.subplots_adjust(hspace=0.3)
#We can do whatever we want in this code, cause our data features are too many, here just show one instance of top two features of permutation plots
# + [markdown] id="rLMbzDplPxWn"
# We could also plot the one-way and two-way PDP together.
# + id="GMgp6pVjPxWn" outputId="21ce7a7b-42d0-4e2d-e711-512f9c6de4e4"
features = [16, 7,(16, 7)]
plot_partial_dependence(estimator=clf, X=X_train_transform,
features=features, feature_names=new_cols,
target=1)
fig = plt.gcf() # get current figure (gcf)
fig.suptitle("Partial dependence of Attack or Normal (one-way and two-way)\n"
"with {}".format(clf_name))
fig.set_size_inches(16.5, 6.5)
fig.subplots_adjust(wspace = 0.3)
# + id="t25mazgGPxWp" outputId="b86d5f15-1e40-4fe6-e687-1274b1ad855c"
features = [37, 38, (37, 38)]
plot_partial_dependence(estimator=clf, X=X_train_transform,
features=features, feature_names=new_cols,
target=1)
fig = plt.gcf() # get current figure (gcf)
fig.suptitle("Partial dependence of Attack or Normal (one-way and two-way)\n"
"with {}".format(clf_name))
fig.set_size_inches(16.5, 6.5)
fig.subplots_adjust(wspace = 0.3)
# + id="obZeCE3HPxWq" outputId="379b2d56-ed7c-46e3-d933-8c0b4bfbc3e7"
features = [7, 37, (7,37)]
plot_partial_dependence(estimator=clf, X=X_train_transform,
features=features, feature_names=new_cols,
target=1)
fig = plt.gcf() # get current figure (gcf)
fig.suptitle("Partial dependence of Attack or Normal (one-way and two-way)\n"
"with {}".format(clf_name))
fig.set_size_inches(16.5, 6.5)
fig.subplots_adjust(wspace = 0.3)
# + id="cJ0-RXiUPxWq" outputId="6129bc35-8c8c-43fa-a47a-4c3637c9bcf9"
features = [7, 38, (7,38)]
plot_partial_dependence(estimator=clf, X=X_train_transform,
features=features, feature_names=new_cols,
target=1)
fig = plt.gcf() # get current figure (gcf)
fig.suptitle("Partial dependence of Attack or Normal (one-way and two-way)\n"
"with {}".format(clf_name))
fig.set_size_inches(16.5, 6.5)
fig.subplots_adjust(wspace = 0.3)
# + id="UPqF5LenPxWr" outputId="6bb67ea7-d5ae-424b-f6a1-6976da89bfae"
features = [16, 37, (16,37)]
plot_partial_dependence(estimator=clf, X=X_train_transform,
features=features, feature_names=new_cols,
target=1)
fig = plt.gcf() # get current figure (gcf)
fig.suptitle("Partial dependence of Attack or Normal (one-way and two-way)\n"
"with {}".format(clf_name))
fig.set_size_inches(16.5, 6.5)
fig.subplots_adjust(wspace = 0.3)
# + id="o7aUubhxPxWs" outputId="8b9ac26a-8a27-4073-dd94-bbbe989f1cdb"
features = [16, 38, (16,38)]
plot_partial_dependence(estimator=clf, X=X_train_transform,
features=features, feature_names=new_cols,
target=1)
fig = plt.gcf() # get current figure (gcf)
fig.suptitle("Partial dependence of Attack or Normal (one-way and two-way)\n"
"with {}".format(clf_name))
fig.set_size_inches(16.5, 6.5)
fig.subplots_adjust(wspace = 0.3)
# + [markdown] id="GRgHqW8VPxWs"
# ### Example: PDP for RandomTree
# Let's plot the effect (one-way and two-way) for `RandomForest`.
# + id="_SGLrcnTPxWs" outputId="00f4408f-560e-42a3-ff09-67ec31f9299b"
models[2]
# + id="-YbztYyEPxXH" outputId="7e861467-93e4-4cf2-862f-2c5e35573425"
clf_name = models[2][0]
clf = models[2][1]
features = [(16, 7)]
plot_partial_dependence(estimator=clf, X=X_train_transform,
features=features, feature_names=new_cols,
target=1)
fig = plt.gcf() # get current figure (gcf)
fig.suptitle("Partial dependence of Attack Type\n"
"with {}".format(clf_name))
fig.set_size_inches(7.5, 6.5)
fig.subplots_adjust(hspace=0.3)
# + [markdown] id="-Mwid9-PPxXI"
# ## Permutation Feature Importance
#
# Previous section shows that Partial Dependence Plots could present how a feature affects predictions. In this section, we focus on another basic question: What features have the biggest impact on predictions?
#
# This concept is called **feature importance**.
#
# There are multiple ways to measure feature importance. Some approaches answer subtly different versions of the question above. Other approaches have documented shortcomings.
#
# We'll focus on **Permutation Feature Importance**. Compared to most other approaches, Permutation Feature Importance is:
#
# - fast to calculate,
# - widely used and understood, and
# - consistent with properties we would want a feature importance measure to have.
#
# The Permutation Feature Importance is defined to be the **decrease in a model score** when a single feature value is randomly shuffled. This procedure breaks the relationship between the feature and the target, thus the drop in the model score is indicative of how much the model depends on the feature.
#
# The above definition might be confused. Here is a clear example to show you how it works: https://www.kaggle.com/dansbecker/permutation-importance
# + id="MrH547kDPxXJ"
# import permutation function
from sklearn.inspection import permutation_importance
# + [markdown] id="XaHjOXdZPxXK"
# ### Example: Permutation Feature Importance for Logistic Regression
#
# #### Permutation Feature Importance on Training Data
# Let's first plot the Permutation Feature Importance on **training data** with the trained Logistic Regression.
# + id="SYgD-XGGPxXK" outputId="99079e70-7f45-4f41-a44b-a60b2e9065b5"
clf_name = models[0][0]
clf = models[0][1]
result = permutation_importance(estimator=clf,
X=X_train_transform,
y=y2_train_transform,
scoring="accuracy",
n_repeats=50,
random_state=123)
sorted_idx = result.importances_mean.argsort()
fig, ax = plt.subplots()
ax.boxplot(result.importances[sorted_idx].T,
vert=False, labels=np.array(new_cols)[sorted_idx])
ax.set_title("Permutation Importances (training set)\n"
"with {}".format(clf_name))
fig.set_size_inches(15, 13)
plt.show()
# + [markdown] id="fSNrTaYOPxXL"
# **Interpretation**
#
# - The values towards the top are the most important features, and those towards the bottom matter least.
#
# - The values show how much model performance decreased with a random shuffling (in this case, using `accuracy` as the performance metric).
#
# The above plot shows that `ct_src_dport_itm` is the most important feature in the Logistic Regression model because once we shuffle the `ct_src_dport_itm`` column of the training data, leaving the target and all other columns in place, the **decrease** of the `accuracy` score of predictions is around 0.24. This is significant finding.
# + [markdown] id="leIIuyYQPxXM"
# #### Permutation Feature Importance on Hold-out Testing Data
#
# Permutation importances can either be computed on the training set or an held-out testing or validation set.
# - Using a held-out set makes it possible to highlight which features contribute the most to the generalization power of the inspected model.
#
# - Features that are important on the training set but not on the held-out set might cause the model to overfit.
#
# Let's plot the Permutation Feature Importance on **hold-out testing data**
# + id="-ECxsS02PxXN" outputId="46dd9aa9-b1f1-4a7c-f26d-5dbd765f15c9"
result = permutation_importance(estimator=clf,
X=X_test_transform,
y=y2_test_transform,
scoring="accuracy",
n_repeats=50,
random_state=123)
sorted_idx = result.importances_mean.argsort()
fig, ax = plt.subplots()
ax.boxplot(result.importances[sorted_idx].T,
vert=False, labels=np.array(new_cols)[sorted_idx])
ax.set_title("Permutation Importances (hold-out testing set)\n"
"with {}".format(clf_name))
fig.set_size_inches(15, 13)
plt.show()
# + id="Y75CfTyTPxXO"
| Testing_under_sampling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from sqlalchemy import create_engine
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
from config import DB_USER, DB_PASS
engine = create_engine(f"postgresql://{DB_USER}:{DB_PASS}@localhost/employee_db")
conn = engine.connect()
salaries = pd.read_sql("SELECT * FROM salaries", conn)
salaries.head()
titles = pd.read_sql("SELECT * FROM titles", conn)
titles.head()
merged = pd.merge(salaries, titles, on="emp_no", how="inner")
merged.head()
grouped = merged.groupby("title").mean()
grouped
title_salary_df = grouped.drop(columns = "emp_no")
title_salary_df
title_salary_df = title_salary_df.reset_index()
title_salary_df
# +
x_axis = title_salary_df["title"]
ticks = np.arange(len(x_axis))
y_axis = title_salary_df["salary"]
plt.bar(x_axis, y_axis, align="center", alpha=1.0, color=["pink", "b", "r", "orange", "y", "b", "g"])
plt.xticks(ticks, x_axis, rotation="vertical")
plt.ylabel("Salaries ($)")
plt.xlabel("Employee Title")
plt.title("Average Employee Salary by Title")
plt.savefig("../Images/avg_salary_by_title.png")
plt.show()
# -
| employee data analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import os
os.environ['MPLCONFIGDIR'] = "/home/mventura/latent_ode_expts"
import sys
import matplotlib
matplotlib.use('Agg')
import matplotlib.pyplot
import matplotlib.pyplot as plt
import time
import datetime
import argparse
import numpy as np
import pandas as pd
from random import SystemRandom
from sklearn import model_selection
import torch
import torch.nn as nn
from torch.nn.functional import relu
import torch.optim as optim
import lib.utils as utils
from lib.plotting import *
from lib.rnn_baselines import *
from lib.ode_rnn import *
from lib.create_latent_ode_model import create_LatentODE_model
from lib.parse_datasets import parse_datasets
from lib.ode_func import ODEFunc, ODEFunc_w_Poisson
from lib.diffeq_solver import DiffeqSolver
from mujoco_physics import HopperPhysics
from lib.utils import compute_loss_all_batches
# %load_ext autoreload
# %autoreload 2
# -
# # LatentODE(10)
# +
# # !python3 run_models.py --niters 50 -n 1000 -s 1 -l 10 --dataset periodic --latent-ode --noise-weight 0 --viz
# +
# # !python3 run_models.py --niters 50 -n 1000 -s 0.5 -l 10 --dataset periodic --latent-ode --noise-weight 0 --viz
# -
# latent dim = 10, missing = 0, noise = 0
# !python3 run_models.py --niters 50 -n 1000 -s 1 -l 10 --dataset periodic_d1 --latent-ode --noise-weight 0 --viz
# latent dim = 10, missing = 50, noise = 0
# !python3 run_models.py --niters 50 -n 1000 -s 0.5 -l 10 --dataset periodic_d1 --latent-ode --noise-weight 0 --viz
# # LatentODE(5)
# +
# # !python3 run_models.py --niters 50 -n 1000 -s 1 -l 5 --dataset periodic --latent-ode --noise-weight 0 --viz
# +
# # !python3 run_models.py --niters 50 -n 1000 -s 0.5 -l 5 --dataset periodic --latent-ode --noise-weight 0 --viz
# -
# latent dim = 5, missing = 0, noise = 0
# !python3 run_models.py --niters 50 -n 1000 -s 1 -l 5 --dataset periodic_d1 --latent-ode --noise-weight 0 --viz
# latent dim = 5, missing = 50, noise = 0
# !python3 run_models.py --niters 50 -n 1000 -s 0.5 -l 5 --dataset periodic_d1 --latent-ode --noise-weight 0 --viz
| Experiment 2 - Adding Discontinuity 1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# TensorFlow Wide & Deep Learning Tutorial
#python wide_n_deep_tutorial.py --model_type=wide_n_deep
# -
wide_columns = [
gender, native_country, education, occupation, workclass, relationship, age_buckets,
tf.contrib.layers.crossed_column([education, occupation], hash_bucket_size=int(1e4)),
tf.contrib.layers.crossed_column([native_country, occupation], hash_bucket_size=int(1e4)),
tf.contrib.layers.crossed_column([age_buckets, education, occupation], hash_bucket_size=int(1e6))]
deep_columns = [
tf.contrib.layers.embedding_column(workclass, dimension=8),
tf.contrib.layers.embedding_column(education, dimension=8),
tf.contrib.layers.embedding_column(gender, dimension=8),
tf.contrib.layers.embedding_column(relationship, dimension=8),
tf.contrib.layers.embedding_column(native_country, dimension=8),
tf.contrib.layers.embedding_column(occupation, dimension=8),
age, education_num, capital_gain, capital_loss, hours_per_week]
import tempfile
model_dir = tempfile.mkdtemp()
m = tf.contrib.learn.DNNLinearCombinedClassifier(
model_dir=model_dir,
linear_feature_columns=wide_columns,
dnn_feature_columns=deep_columns,
dnn_hidden_units=[100, 50])
# +
import pandas as pd
import urllib
# Define the column names for the data sets.
COLUMNS = ["age", "workclass", "fnlwgt", "education", "education_num",
"marital_status", "occupation", "relationship", "race", "gender",
"capital_gain", "capital_loss", "hours_per_week", "native_country", "income_bracket"]
LABEL_COLUMN = 'label'
CATEGORICAL_COLUMNS = ["workclass", "education", "marital_status", "occupation",
"relationship", "race", "gender", "native_country"]
CONTINUOUS_COLUMNS = ["age", "education_num", "capital_gain", "capital_loss",
"hours_per_week"]
# Download the training and test data to temporary files.
# Alternatively, you can download them yourself and change train_file and
# test_file to your own paths.
train_file = tempfile.NamedTemporaryFile()
test_file = tempfile.NamedTemporaryFile()
urllib.urlretrieve("http://mlr.cs.umass.edu/ml/machine-learning-databases/adult/adult.data", train_file.name)
urllib.urlretrieve("http://mlr.cs.umass.edu/ml/machine-learning-databases/adult/adult.test", test_file.name)
# Read the training and test data sets into Pandas dataframe.
df_train = pd.read_csv(train_file, names=COLUMNS, skipinitialspace=True)
df_test = pd.read_csv(test_file, names=COLUMNS, skipinitialspace=True, skiprows=1)
df_train[LABEL_COLUMN] = (df_train['income_bracket'].apply(lambda x: '>50K' in x)).astype(int)
df_test[LABEL_COLUMN] = (df_test['income_bracket'].apply(lambda x: '>50K' in x)).astype(int)
def input_fn(df):
# Creates a dictionary mapping from each continuous feature column name (k) to
# the values of that column stored in a constant Tensor.
continuous_cols = {k: tf.constant(df[k].values)
for k in CONTINUOUS_COLUMNS}
# Creates a dictionary mapping from each categorical feature column name (k)
# to the values of that column stored in a tf.SparseTensor.
categorical_cols = {k: tf.SparseTensor(
indices=[[i, 0] for i in range(df[k].size)],
values=df[k].values,
dense_shape=[df[k].size, 1])
for k in CATEGORICAL_COLUMNS}
# Merges the two dictionaries into one.
feature_cols = dict(continuous_cols.items() + categorical_cols.items())
# Converts the label column into a constant Tensor.
label = tf.constant(df[LABEL_COLUMN].values)
# Returns the feature columns and the label.
return feature_cols, label
def train_input_fn():
return input_fn(df_train)
def eval_input_fn():
return input_fn(df_test)
# -
m.fit(input_fn=train_input_fn, steps=200)
results = m.evaluate(input_fn=eval_input_fn, steps=1)
for key in sorted(results):
print("%s: %s" % (key, results[key]))
| crackingcode/day13/cc_tf_day13_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.7.1
# language: julia
# name: julia-1.7
# ---
# + [markdown] hideCode=false hidePrompt=false slideshow={"slide_type": "slide"}
# # 2022-01-19 Conditioning
#
# * Office hours: Monday 9-10pm, Tuesday 2-3pm, Thursday 2-3pm
# * More on floating point
# * Discuss Taylor Series activity
# * Condition numbers
# * Forward and backward error
# * Computing volume of a polygon
# + slideshow={"slide_type": "skip"}
using Plots
default(linewidth=4)
# + [markdown] slideshow={"slide_type": "slide"}
# # Floating point representation is **relative**
#
# <img src="https://upload.wikimedia.org/wikipedia/commons/a/a9/IEEE_754_Double_Floating_Point_Format.svg" width="80%" />
# + [markdown] cell_style="split" slideshow={"slide_type": ""}
# Let $\operatorname{fl}$ round to the nearest floating point number.
#
# $$ \operatorname{fl}(x) = x (1 + \epsilon), \quad \text{where} |\epsilon| \le \epsilon_{\text{machine}} $$
#
# This also means that the relative error in representing $x$ is small:
#
# $$ \frac{|\operatorname{fl}(x) - x|}{|x|} \le \epsilon_{\text{machine}} $$
# + cell_style="split"
plot(x -> (1 + x) - 1, xlims=(-1e-15, 1e-15))
plot!(x -> x)
# + [markdown] slideshow={"slide_type": "slide"}
# # Exact arithmetic, correctly rounded
# + [markdown] cell_style="split" slideshow={"slide_type": ""}
# Take an elementary math operation $*$ (addition, subtraction, multiplication, division), and the discrete operation that our computers perform, $\circledast$. Then
#
# $$x \circledast y := \operatorname{fl}(x * y)$$
#
# with a relative accuracy $\epsilon_{\text{machine}}$,
#
# $$ \frac{|(x \circledast y) - (x * y)|}{|x * y|} \le \epsilon_{\text{machine}} . $$
# + [markdown] cell_style="split" slideshow={"slide_type": "fragment"}
# ## Seems easy, how do operations compose?
#
# Is this true?
#
# $$ \frac{\Big\lvert \big((x \circledast y) \circledast z\big) - \big((x * y) * z\big) \Big\rvert}{|(x * y) * z|} \le^? \epsilon_{\text{machine}} $$
# + slideshow={"slide_type": "fragment"}
f(x; y=1, z=-1) = (x+y)+z # The best arbitrary numbers are 0, 1, and -1
plot(x -> abs(f(x) - x)/abs(x), xlims=(-1e-15, 1e-15))
# + [markdown] slideshow={"slide_type": "slide"}
# # Which operation caused the error?
#
# 1. $\texttt{tmp} = \operatorname{fl}(x + 1)$
# 2. $\operatorname{fl}(\texttt{tmp} - 1)$
#
# Use Julia's [`BigFloat`](https://docs.julialang.org/en/v1/base/numbers/#BigFloats-and-BigInts)
# -
@show typeof(big(.1))
@show big(.1) # Or BigFloat(.1); parsed as Float64, then promoted
@show BigFloat(".1"); # Parse directly to BigFloat
tmp = 1e-15 + 1
tmp_big = big(1e-15) + 1 # Parse as Float64, then promote
abs(tmp - tmp_big) / abs(tmp_big)
r = tmp - 1
r_big = big(tmp) - 1
abs(r - r_big) / abs(r_big)
# + [markdown] slideshow={"slide_type": "slide"}
# # Activity: [2022-01-12-taylor-series](https://classroom.github.com/a/VkPvGOgu)
#
# * Use Julia, Jupyter, Git
# * Look at how fast series converge when taking only finitely many terms
# * Explore instability, as is occuring for large negative `x` above, but not for the standard library `expm1`
# + slideshow={"slide_type": "skip"}
function myexp(x, k)
sum = 0
term = 1
n = 1
# modify so at most k terms are summed
while sum + term != sum
sum += term
term *= x / n
# YOUR CODE HERE
if n == k
break
end
n += 1
end
sum
end
rel_error(x, k) = abs(myexp(x, k) - exp(x)) / exp(x)
ks = 2 .^ (0:10) # [1, 2, 4, ..., 1024];
# + cell_style="split"
plot(ks, k -> rel_error(-20, k), xscale=:log10, yscale=:log10)
# + cell_style="split"
plot(x -> rel_error(x, 1000) + 1e-17, xlims=(-20, 20), yscale=:log10)
# + [markdown] slideshow={"slide_type": "slide"}
# # What happened?
# Let's look at the terms for positive and negative $x$
# + cell_style="split"
function expterms(x, k=50)
term = 1.
terms = [term]
for n in 1:k
term *= x / n
push!(terms, term)
end
terms
end
x = -10
@show sum(expterms(x)) - exp(x)
@show (sum(expterms(x)) - exp(x)) / exp(x)
expterms(x)
# + cell_style="split" slideshow={"slide_type": ""}
@show exp(-10)
bar(expterms(-10))
# + [markdown] slideshow={"slide_type": "slide"}
# # Conditioning
#
# > What sort of functions cause small errors to become big?
#
# Consider a function $f: X \to Y$ and define the **absolute condition number**
# $$ \hat\kappa = \lim_{\delta \to 0} \max_{|\delta x| < \delta} \frac{|f(x + \delta x) - f(x)|}{|\delta x|} = \max_{\delta x} \frac{|\delta f|}{|\delta x|}. $$
# If $f$ is differentiable, then $\hat\kappa = |f'(x)|$.
#
# Floating point offers relative accuracy, so it's more useful to discuss **relative condition number**,
# $$ \kappa = \max_{\delta x} \frac{|\delta f|/|f|}{|\delta x|/|x|}
# = \max_{\delta x} \Big[ \frac{|\delta f|/|\delta x|}{|f| / |x|} \Big] $$
# or, if $f$ is differentiable,
# $$ \kappa = |f'(x)| \frac{|x|}{|f|} . $$
# + [markdown] slideshow={"slide_type": "slide"}
# # Condition numbers
#
# > $$ \kappa = |f'(x)| \frac{|x|}{|f|} $$
# -
f(x) = x - 1; fp(x) = 1
plot(x -> abs(fp(x)) * abs(x) / abs(f(x)), xlims=(0, 2))
# + [markdown] slideshow={"slide_type": "slide"}
# # Back to $f(x) = e^x - 1$
# + cell_style="split"
f(x) = exp(x) - 1
fp(x) = exp(x)
plot(x -> abs(fp(x)) * abs(x) / abs(f(x)))
# + [markdown] cell_style="split" slideshow={"slide_type": "fragment"}
# ## What does it mean?
#
# * The function $f(x) = e^x - 1$ is well-conditioned
# * The function $f_1(x) = e^x$ is well-conditioned
# * The function $f_2(x) = x - 1$ is ill-conditioned for $x \approx 1$
#
# ## The **algorithm** is unstable
#
# * `f(x) = exp(x) - 1` is unstable
# * Algorithms are made from elementary operations
# * Unstable algorithms do something ill-conditioned
# + [markdown] slideshow={"slide_type": "slide"}
# # A stable algorithm
# + [markdown] cell_style="split" slideshow={"slide_type": ""}
# We used the series expansion previously.
# * accurate for small $x$
# * less accurate for negative $x$ (see activity)
# * we could use symmetry to fix
# * inefficient because we have to sum lots of terms
#
# Standard math libraries define a more efficient stable variant, $\texttt{expm1}(x) = e^x - 1$.
# + cell_style="split"
plot([x -> exp(x) - 1,
x -> expm1(x)],
xlims = (-1e-15, 1e-15))
# + [markdown] slideshow={"slide_type": "slide"}
# # Another example $\log(1 + x)$
#
# What is the condition number of $f(x) = \log(1 + x)$ for $x \approx 0$?
# + cell_style="split" slideshow={"slide_type": "fragment"}
plot([x -> log(1 + x),
x -> log1p(x)],
xlims = (-1e-15, 1e-15))
# + cell_style="split"
cond1(x) = abs(1/(1+x) * x / log1p(x))
cond2(x) = abs(1/x * x / log(x))
plot([cond1 cond2], xlims=(-1, 2), ylims=(0, 100))
# + [markdown] slideshow={"slide_type": "slide"}
# # Reliable = well-conditioned and stable
#
# ## Mathematical functions $f(x)$ can be ill-conditioned (big $\kappa$)
# * Modeling is how we turn an abstract question into a mathematical function
# * We want well-conditioned models (small $\kappa$)
# * Some systems are intrinsically sensitive: fracture, chaotic systems, combustion
#
# ## Algorithms `f(x)` can be unstable
# * Unreliable, though sometimes practical
# * Unstable algorithms are constructed from ill-conditioned parts
# + [markdown] slideshow={"slide_type": "slide"}
# # An ill-conditioned problem from <NAME>
#
# From [Surely You're Joking, Mr. Feynman](https://sistemas.fciencias.unam.mx/%7Ecompcuantica/RICHARD%20P.%20FEYNMAN-SURELY%20YOU%27RE%20JOKING%20MR.%20FEYNMAN.PDF) (page 113)
#
# > So Paul is walking past the lunch place and these guys are all excited. "Hey,
# Paul!" they call out. "Feynman's terrific! We give him a problem that can be stated in ten
# seconds, and in a minute he gets the answer to 10 percent. Why don't you give him one?"
# Without hardly stopping, he says, "The tangent of 10 to the 100th."
# I was sunk: you have to divide by pi to 100 decimal places! It was hopeless.
# + [markdown] cell_style="split"
# What's the condition number?
#
# $$ \kappa = |f'(x)| \frac{|x|}{|f|} $$
#
# * $f(x) = \tan x$
# * $f'(x) = 1 + \tan^2 x$
#
# $$ \kappa = \lvert x \rvert \Bigl( \lvert \tan x \rvert + \bigl\lvert \frac{1}{\tan x} \bigr\rvert \Bigr)$$
# + cell_style="split"
tan(1e100)
# + cell_style="split"
tan(BigFloat("1e100", precision=500))
# + [markdown] slideshow={"slide_type": "slide"}
# # Go find some functions...
#
# * Find a function $f(x)$ that models something you're interested in
# * Plot its condition number $\kappa$ as a function of $x$
# * Plot the relative error (using single or double precision; compare using Julia's `big`)
# * Is the relative error ever much bigger than $\kappa \epsilon_{\text{machine}}$?
# * Can you find what caused the instability?
# * Share on Zulip
#
# ## Further reading: [FNC Introduction](https://fncbook.github.io/fnc/intro/overview.html)
| slides/2022-01-19-conditioning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Disclaimer
# #### This is my personal repository, this is not Google approved code
# #### Most of the code was taken from the documentation of the modules used
# + active=""
# %pip install pdfminer.six
# %pip install spacy
# %pip install textacy
# %pip install pyenchant
# + active=""
# ## get the location of the python exe and use that to install the spacy language module
# # I'm getting the large model which supports more words but is more memory intesive
# import sys
# !{sys.executable} -m spacy download en_core_web_lg
#
# # map the core module to the generic 'en' alias for textacy
# !{sys.executable} -m spacy link en_core_web_lg en
#
# !sudo apt-get update -y & sudo apt-get install -y enchant
# +
from pprint import pprint as prt
import pandas as pd
pd.set_option('display.max_rows', 100)
from pprint import pprint as prt
import collections
from io import StringIO, BytesIO
import warnings
warnings.filterwarnings('ignore')
import spacy
nlp = spacy.load('en_core_web_lg')
import textacy
# -
# # How to build a spelling dictionary with industry terms
# ### Download the top 10k most common used words
# #### This is derived from Google's Trillion Word Corpus
# +
import urllib
top_10k_american_english = 'https://raw.githubusercontent.com/first20hours/google-10000-english/master/google-10000-english-usa.txt'
webUrl = urllib.request.urlopen(top_10k_american_english)
top_10k_ae = webUrl.read()
top_10k_ae = top_10k_ae.decode('utf-8').upper().split('\n')
top_10k_ae[:10]
# -
# ### Let's mine a PDF for industry specific words
# #### You'll want to use a corpus of documents to mine for words & terms
# #### Be sure to pick source documents that have a high likelyhood of spelling accuracy
pdf_url = 'https://www.toyota.com/t3Portal/document/om-s/OM60R53U/pdf/OM60R53U.pdf'
webUrl = urllib.request.urlopen(pdf_url)
pdf_data = webUrl.read()
# +
from pdfminer.converter import TextConverter
from pdfminer.layout import LAParams
from pdfminer.pdfdocument import PDFDocument
from pdfminer.pdfinterp import PDFResourceManager, PDFPageInterpreter
from pdfminer.pdfpage import PDFPage
from pdfminer.pdfparser import PDFParser
output_string = StringIO()
parser = PDFParser(BytesIO(pdf_data))
doc = PDFDocument(parser)
rsrcmgr = PDFResourceManager()
device = TextConverter(rsrcmgr, output_string, laparams=LAParams())
interpreter = PDFPageInterpreter(rsrcmgr, device)
for page in PDFPage.create_pages(doc):
interpreter.process_page(page)
pdf_text = output_string.getvalue()
# -
# ### Extract and normalize words
# #### This example does not extract words that have numbers or special characters embedded
# #### Words are uppercased to standardize for matching
spacy_doc = nlp(pdf_text)
word_list_raw = [i.text.upper() for i in spacy_doc if i.string.isalpha() == True] #isalnum
word_count = len(word_list_raw)
print(f'{word_count} words were found')
# #### Get word frequency counts
# #### If you use a large enough corpus you should see a delinator for misspelled words
# #### An example would be removing any words that have less then 3 occurrences
# #### When you look through the words you should see examples of singular & plural words
# #### we will convert the plural words to singular using lemmatization
word_list_raw_df = pd.DataFrame(word_list_raw, columns=['word'])
word_list_raw_df['count'] = 1
word_list_raw_counts = word_list_raw_df.groupby('word').count().reset_index()
word_list_raw_counts.sort_values('count', ascending=False, inplace=True )
# remove words that are to short
word_list_raw_counts = word_list_raw_counts[ word_list_raw_counts['word'].str.len() > 2 ]
word_list_raw_counts.head(20)
word_list_raw_counts.tail(20)
# #### Check if word is plural and build a hash lookup
# +
replacements = {}
for token in spacy_doc:
if len(token.text.replace('\n', '')) > 1:
word = token.text.upper()
singular = token.lemma_.upper()
if word != singular:
replacements[word] = singular
dict(list(replacements.items())[0:10])
# +
auto_dictionary = set(word_list_raw_counts['word'])
for k,v in replacements.items():
if v != '-PRON-': # Remove PRON as it will throw off modeling
try:
auto_dictionary.remove(k)
auto_dictionary.add(v)
except:
pass
list(auto_dictionary)[:10]
# -
# #### Join the 10k & auto industry lists
spelling_list = list(set(top_10k_ae + list(auto_dictionary)))
print('10k\t', len(top_10k_ae), '\nAuto\t', len(list(auto_dictionary)), '\ntotal\t', len(spelling_list))
# #### Create a custom spell checker
# +
### Reddit comments to check with spelling list
# +
# %%bigquery words_raw
WITH comments AS (SELECT SPLIT(REPLACE(body, '\n', ' '), ' ') AS words
FROM `fh-bigquery.reddit_comments.20*`
WHERE UPPER(subreddit) LIKE '%AUTOMOTIVE%'
OR UPPER(subreddit) LIKE '%MECHANIC%'),
word_list AS (SELECT UPPER(words) AS word
FROM comments,
UNNEST(words) as words
WHERE REGEXP_CONTAINS(words, '[^\\w]') = False)
SELECT word, count(*) as freq
FROM word_list
WHERE ABS(MOD(FARM_FINGERPRINT(word), 10)) < 3
GROUP BY 1
ORDER BY freq DESC
# -
words_raw.tail()
dictionary_df = pd.DataFrame(words_series, columns=['word'])
dictionary_df['match'] = True
dictionary_df.sort_values('word')
words_raw['word'] = words_raw['word'].apply(lambda x: x.strip())
dictionary_df['word'] = dictionary_df['word'].apply(lambda x: x.strip())
check = pd.merge(words_raw, dictionary_df, on='word', how='left')
check
check[(check['match'] != True) & (check['freq'] > 100)]
# +
import enchant
words_series = pd.Series(list(spelling_list))
words_series.to_csv('auto_words_custom_dic.txt', index=False)
auto_spell_checker = enchant.PyPWL('auto_words_custom_dic.txt')
print('Check word in list', auto_spell_checker.check('HOMELINK'))
print('Check word not in list', auto_spell_checker.check('HOMLUNK'))
print('\nSuggestions', auto_spell_checker.suggest('HOMLUNK'))
# -
| building_custom_dictionaries.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: venv-datascience
# language: python
# name: venv-datascience
# ---
import pandas as pd
# +
airports = pd.DataFrame([
['Seatte-Tacoma', 'Seattle', 'USA'],
['Dulles', 'Washington', 'USA'],
['Heathrow', 'London', 'United Kingdom'],
['Schiphol', 'Amsterdam', 'Netherlands'],
['Changi', 'Singapore', 'Singapore'],
['Pearson', 'Toronto', 'Canada'],
['Narita', 'Tokyo', 'Japan']
],
columns = ['Name', 'City', 'Country']
)
airports
# -
# ### one column
airports['City']
# ### multiple columns
airports[["City", "Country"]]
# ## iloc for specific rows and columns
airports.iloc[0,0]
airports.iloc[2,2]
# ### Range
airports.iloc[:,:]
airports.iloc[0:2,:]
airports.iloc[5:, :]
airports.iloc[3:5, 1:]
# ### Individual columns
airports.iloc[:, [0,2]]
# ### Using loc by names
airports.loc[:, ["Name", "Country"]]
| More Python Data Tools - Microsoft/02.Query a pandas DataFrame.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="Ee5lf3Kk71-C" colab_type="code" colab={}
# Execute this command to check GPU model
# In Colab the best one is Tesla P100
# !nvidia-smi
# + id="NlhNcXef439t" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1600506705110, "user_tz": -120, "elapsed": 10335, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13422679531871080513"}}
import sys
sys.path.append('/content/drive/My Drive/DeepNILM')
import model_testing
import metrics
# + [markdown] id="H4ZAcz3z5Yj1" colab_type="text"
# ### Dishwasher test
# + id="Gdv18hxR4bdC" colab_type="code" colab={}
ground_truth, predicted_values = model_testing.test_model(appliance_name='dishwasher',
main_path='path/to/main_test.csv',
appliance_path='path/to/dishwasher_test.csv',
model_path='path/to/dishwasher_model',
window_size=600,
batch_size=512,
rescaling='normalize',
appliance_min_power=0.0,
appliance_max_power=2570.6,
main_min_power=73.48100000000002,
main_max_power=6048.699999999999)
# + id="uUcIGeVn-gUk" colab_type="code" colab={}
f1 = metrics.compute_F1_score(predicted_values, ground_truth)
print('Energy based F1 score on test set: {}'.format(f1))
# + [markdown] id="2BRWIlGn5c-2" colab_type="text"
# ### Fridge test
# + id="lGHyy0TK5egw" colab_type="code" colab={}
ground_truth, predicted_values = model_testing.test_model(appliance_name='fridge',
main_path='path/to/main_test.csv',
appliance_path='path/to/fridge_test.csv',
model_path='path/to/fridge_model',
window_size=600,
batch_size=512,
rescaling='standardize',
appliance_mean_power=37.23710644724372,
appliance_std_power=46.9886959530205,
main_mean_power=370.91555422946004,
main_std_power=549.1880538356259)
# + id="w8BqfYGm-lDo" colab_type="code" colab={}
f1 = metrics.compute_F1_score(predicted_values, ground_truth)
print('Energy based F1 score on test set: {}'.format(f1))
| TestingNotebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Outliers and Their Influence on the Final Model - tutorial
#
# ## Table of Contents:
#
# 1. Read point data and take 10% of it as a sample for the further analysis (dataset A),
# 2. Check if outliers are present in a data and create additional dataset without outliers (dataset B),
# 3. Create the Variogram Point Cloud model for the datasets A and B,
# 4. Remove outliers from the datasets A and B,
# 5. Create four Ordinary Kriging models and compare their performance.
#
# ## Level: Intermediate
#
# ## Changelog
#
# | Date | Change description | Author |
# |------|--------------------|--------|
# | 2021-12-14 | Sill selection was upgraded: now optimal sill is derived from the grid search within `TheoreticalSemivariogram` class | @SimonMolinsky |
# | 2021-12-13 | Changed behavior of `select_values_in_range()` function | @SimonMolinsky |
# | 2021-12-11 | Behavior of `prepare_kriging_data()` function has benn changed | @SimonMolinsky |
# | 2021-10-13 | Refactored TheoreticalSemivariogram (name change of class attribute) and refactored `calc_semivariance_from_pt_cloud()` function to protect calculations from `NaN's`. | @ethmtrgt & @SimonMolinsky |
# | 2021-08-22 | Initial release | @SimonMolinsky |
#
# ## Introduction
#
# Outliers may affect our analysis and the final interpolation results. In this tutorial we learn about their influence on the final model and we compare interpolation error for different scenarios where data is treated in a different ways.
#
# We are able to remove too high or too low values at the preprocessing stage (check part 2 of the tutorial) or we can remove outliers directly from the variogram point cloud (part 4). Results from each type of preprocessing (and a raw dataset analysis) are different and we are going to compare them.
#
# We use:
#
# - DEM data which is stored in a file `sample_data/point_data/poland_dem_gorzow_wielkopolski`.
# ## Import packages
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from pyinterpolate.distance import calc_point_to_point_distance
from pyinterpolate.io_ops import read_point_data
from pyinterpolate.kriging import Krige
from pyinterpolate.semivariance import build_variogram_point_cloud, show_variogram_cloud, remove_outliers
from pyinterpolate.semivariance import calc_semivariance_from_pt_cloud
from pyinterpolate.semivariance import TheoreticalSemivariogram
# -
# ## 1) Read point data and divide it into training and test set
# +
# Read data from file
dem = read_point_data('../sample_data/point_data/poland_dem_gorzow_wielkopolski', data_type='txt')
# +
# Divide data into training and test set
def create_train_test(data, training_fraction):
idxs = np.arange(0, len(data))
number_of_training_samples = int(len(data) * training_fraction)
training_idxs = np.random.choice(idxs, size=number_of_training_samples, replace=False)
test_idxs = [i for i in idxs if i not in training_idxs]
training_set = data[training_idxs, :]
test_set = data[test_idxs, :]
return training_set, test_set
# -
train, test = create_train_test(dem, 0.1)
train
# ## 2) Check outliers: analyze distribution of the values
#
# To find if our dataset contains outliers we are going to inspect all values in the `train` set. At the beginning we plot data distribution with the `violinplot`.
# +
# Distribution plot
plt.figure(figsize=(10, 6))
plt.violinplot(train[:, -1])
plt.show()
# -
# > **NOTE:** Your plot may be different than presented in the tutorial. Why is that? Because we take a random sample of 10% of values and after each iteration the algorithm takes different points for the analysis.
#
# **Clarification:**
#
# Investigation of the plot tells us that our data is:
#
# - grouped around the lowest values, and most of the values are below 50 meters,
# - has (probably) three different distributions mixed together, it can be a sign that Digital Elevation Model covers three different types of the elevation. One is grouped around 20 meters, next around 50 meters and the faintest is visible around 70 meters.
#
# **Violinplot** is good for the distribution analysis. Especially if we are looking for the complex patterns in the dataset. But reading outliers from it may be challenging and we should change a plot type to understand if outliers exist in a dataset. The good choice is the `boxplot`:
# +
# Boxplot
plt.figure(figsize=(10, 6))
plt.boxplot(train[:, -1])
plt.show()
# -
# **Boxplot** is a special and very useful tool for the data visualization and analysis. Let's analyze this plot from the bottom up to the top.
#
# > **NOTE:** Boxplot represents values sorted in the ascending order and their statistical properties: quartiles, median and outliers.
#
# - The bottom whisker (horizontal line) represents the lower range of values in our dataset,
# - The box lower line is the first quartile of a data or, in other words, 25% of values of our dataset are below this point. We name it Q1.
# - The middle line is a median of our dataset. We name it Q2 or median.
# - The upper line is the third quartile of a data or, in other words, 75% of values are below this point,
# - The top whisker represents the upper range of values in our dataset. We name it Q3.
# - Individual points (if they are exist then we see them as a points below the bottom whisker or above the top whisker) are considered as outliers. They could be outliers in the upper range as well as lower range of our data. Long distance between Q1 and the bottom whisker and/or between Q3 and the top whisker is an indicator of potential outliers. Package **matplotlib** calculates potential outliers based on the absolute distance from the Q1 or Q3 to the whiskers. Points below or above this distance are treated as outliers. The outlier distance is calculated as the $weight * (Q3 - Q1)$ where we can set `weight` but other parameters are read directly from the data.
#
# We use this knowledge to remove outliers from the dataset with the assumption that _outliers are rather anomalies than unbiased readings_. We will perform the outlier removal from the data with a more *aggresive* assumption than it is done in **matplotlib** and we set weight to the `1.0`.
# +
# Create training set without outliers
q1 = np.quantile(train[:, -1], 0.25)
q3 = np.quantile(train[:, -1], 0.75)
top_limit = q3 + (q3 - q1)
train_without_outliers = train[train[:, -1] < top_limit]
# -
print('Length of the full training set is {} records'.format(len(train)))
print('Length of the pre-processed training set is {} records'.format(len(train_without_outliers)))
fig, ax = plt.subplots(nrows=1, ncols=2, figsize=(16, 6))
ax[0].violinplot(train_without_outliers[:, -1])
ax[0].set_title('Distribution of the training set without outliers')
ax[1].boxplot(train_without_outliers[:, -1])
ax[1].set_title('Statistics of the training set without outliers')
plt.show()
# **Clarification**: with our data processing we have cut some records from the baseline training dataset. Distribution plot (`violinplot`) has shorter tail and ends more abruptly; and a `boxplot` of the new data doesn't have any outliers. The one important thing to notice is that the observations are still skewed but this is not a problem for this concrete tutorial.
#
# > **NOTE**: if you are eager to know how to deal with the skewed datasets we recommend article [**Transforming Skewed Data**](https://anatomisebiostats.com/biostatistics-blog/transforming-skewed-data/).
# ## 3) Create the Variogram Point Cloud model for datasets A and B
#
# Now we are making one step further and we will transform both datasets with- and without- outliers and calculate variogram point clouds from these. Then we compare both variogram point clouds.
def get_variogram_point_cloud(dataset, max_range, number_of_lags=16):
distances = calc_point_to_point_distance(dem[:, :-1])
step_size = max_range / number_of_lags
cloud = build_variogram_point_cloud(dataset, step_size, max_range)
return cloud
# +
full_distance = np.max(calc_point_to_point_distance(train[:, :-1]))
cloud_full = get_variogram_point_cloud(train, full_distance)
cloud_processed = get_variogram_point_cloud(train_without_outliers, full_distance)
# +
# Show variogram cloud: initial training dataset
show_variogram_cloud(cloud_full, plot_type='boxplot')
# +
# Show variogram cloud: pre-processed training dataset
show_variogram_cloud(cloud_processed, plot_type='boxplot')
# -
# **Clarification:** a quick look into the results shows that each lag is full of outliers in the top part of the semivariances values. Processed dataset has lowest absolute semivariance than the raw readings. Both variograms have a similar shape. Dispersion of semivariances seems to be very high in both cases. It is especially alarming when we consider the shortest distances where abrupt changes in the elevation are not so likely.
#
# In the next cell we will check a standard deviation of the lag variances.
for k, v in cloud_full.items():
print('Lag {:.2f}'.format(k))
v_raw = int(np.std(v))
v_pro = int(np.std(cloud_processed[k]))
v_smape = 100 * (np.abs(v_raw - v_pro) / (0.5 * (v_raw + v_pro)))
print('Standard Deviation raw dataset:', v_raw)
print('Standard Deviation processed dataset:', v_pro)
print('Symmetric Mean Absolute Percentage Error of Variances: {:.2f}'.format(v_smape))
print('')
# **Clarification:** The differences (sMAPE) per lag vary a lot. We can see that the preprocessing of raw values introduces the information lost. It is especially painful for the closest neighbors. It doesn't mean that the preprocessing of raw observations is not recommended but it is a good idea to include the **spatial component** in the outliers detection process.
#
# Not everything is wrong. Data cleaning has lowered the semivariances dispersion for the middle lags (where we have the largest number of point pairs for the analysis).
#
# At this point we are not able to judge which dataset is better for the modeling. Instead we are going to remove outliers from the both **variograms** (instead of the **raw data**).
# ## 4) Remove outliers from the variograms
#
# In this step we are going to use **pyinterpolate's** function `remove_outliers()` to build additional two variogram point clouds from the raw and processed datasets. We delete the top part outliers of the **semivariance values** rather than the raw readings.
raw_without_outliers = remove_outliers(cloud_full, exclude_part='top', weight=1.25)
prep_without_outliers = remove_outliers(cloud_processed, exclude_part='top', weight=1.25)
data_raw = [x for x in cloud_full.values()]
data_raw_not_out = [x for x in raw_without_outliers.values()]
data_prep = [x for x in cloud_processed.values()]
data_prep_not_out = [x for x in prep_without_outliers.values()]
# +
fig, ax = plt.subplots(nrows=2, ncols=2, figsize=(16, 14))
ax[0, 0].boxplot(data_raw)
ax[0, 0].set_title('Semi-variances Distribution in the Raw Dataset')
ax[0, 0].set_xlabel('Lag number')
ax[0, 0].set_ylabel('Semivariance value')
ax[0, 1].boxplot(data_raw_not_out)
ax[0, 1].set_title('Raw Dataset after the Outliers Detection and Removal')
ax[0, 1].set_xlabel('Lag number')
ax[0, 1].set_ylabel('Semivariance value')
ax[1, 0].boxplot(data_prep)
ax[1, 0].set_title('Semi-variances Distribution in the Pre-processed Dataset')
ax[1, 0].set_xlabel('Lag number')
ax[1, 0].set_ylabel('Semivariance value')
ax[1, 1].boxplot(data_prep_not_out)
ax[1, 1].set_title('Pre-processed Dataset after the Outliers Detection and Removal')
ax[1, 1].set_xlabel('Lag number')
ax[1, 1].set_ylabel('Semivariance value')
plt.show()
# -
# **Clarification:** Comparison of multiple variogram clouds could be hard. We see that the largest semivariances are present in the raw data. A heavily processed data has the lowest number of outliers. The medians in each dataset are distributed over a similar pattern. How is it similar? We can check it if we transform the variogram point cloud into the experimental semivariogram. Pyinterpolate has function for it: `calc_semivariance_from_pt_cloud()`. We use it and compare four plots of semivariances to gain more insight into the transformations.
raw_semivar = calc_semivariance_from_pt_cloud(cloud_full)
raw_semivar_not_out = calc_semivariance_from_pt_cloud(raw_without_outliers)
prep_semivar = calc_semivariance_from_pt_cloud(cloud_processed)
prep_semivar_not_out = calc_semivariance_from_pt_cloud(prep_without_outliers)
plt.figure(figsize=(14, 6))
plt.plot(raw_semivar[:, 1])
plt.plot(raw_semivar_not_out[:, 1])
plt.plot(prep_semivar[:, 1])
plt.plot(prep_semivar_not_out[:, 1])
plt.title('Comparison of experimental semivariograms created with the different data preprocessing techniques')
plt.ylabel('Semivariance')
plt.xlabel('Lag number')
plt.legend(['Raw', 'Raw - remove_outliers()', 'Pre-processed', 'Pre-processed - remove_outliers()'])
plt.show()
# **Clarification:** An understanding of those plots is not an easy task. Let's divide reasoning into multiple points:
#
# - Raw dataset and preprocessed raw dataset show a similar pattern, the differences are more pronounced for the distant lags than for the closest point pairs,
# - Datasets with the cleaned variograms are different from the raw data. The absolute semivariance values per lag are smaller and the semivariogram pattern is slightly different. What is interesting is that the possible two distributions within the dataset are more visible in the case of cleaned variograms (one distribution with a peak around 6th lag and other with a peak around 13th lag).
# - The differences between semivariograms are mostly visible at larger distances. For the closest point pairs differences are smaller.
#
# Semivariograms visual inspection does not infor us of the modeling performance but we can assume that models will be slightly different. Let's test this assumption!
# ## 5) Create Four Ordinary Kriging models based on the four Variogram Point Clouds and compare their performance
# +
number_of_ranges = 32
# Fit different semivariogram models into prepared datasets and variograms
# Raw
raw_theo = TheoreticalSemivariogram(points_array=train,
empirical_semivariance=raw_semivar)
_ = raw_theo.find_optimal_model(weighted=False, number_of_ranges=number_of_ranges)
# Raw with cleaned variogram
raw_theo_no_out = TheoreticalSemivariogram(points_array=train,
empirical_semivariance=raw_semivar_not_out)
_ = raw_theo_no_out.find_optimal_model(weighted=False, number_of_ranges=number_of_ranges)
# Preprocessed
prep_theo = TheoreticalSemivariogram(points_array=train_without_outliers,
empirical_semivariance=prep_semivar)
_ = prep_theo.find_optimal_model(weighted=False, number_of_ranges=number_of_ranges)
# Preprocessed with cleaned variogram
prep_theo_no_out = TheoreticalSemivariogram(points_array=train_without_outliers,
empirical_semivariance=prep_semivar_not_out)
_ = prep_theo_no_out.find_optimal_model(weighted=False, number_of_ranges=number_of_ranges)
# +
# Set Kriging models
# Raw
raw_model = Krige(semivariogram_model=raw_theo, known_points=train)
# Raw & cleaned
c_raw_model = Krige(semivariogram_model=raw_theo_no_out, known_points=train)
# Preprocessed
prep_model = Krige(semivariogram_model=prep_theo, known_points=train_without_outliers)
# Preprocessed & cleaned
c_prep_model = Krige(semivariogram_model=prep_theo_no_out, known_points=train_without_outliers)
# +
# Build test function
def test_kriging_model(model, test_set, max_nn=32):
"""
Function tests performance of a given kriging model.
INPUT:
:param model: (Krige) Kriging model,
:param test_set: (array),
:param max_nn: (int) default=256, maximum number of neighbors.
OUTPUT:
:returns: (list) root mean squared errors of prediction
"""
rmses = []
for pt in test_set:
coordinates = pt[:-1]
value = pt[-1]
predicted = model.ordinary_kriging(coordinates, max_no_neighbors=max_nn)[0]
error = np.sqrt((value - predicted)**2)
rmses.append(error)
return rmses
# -
r_test = test_kriging_model(raw_model, test)
cr_test = test_kriging_model(c_raw_model, test)
p_test = test_kriging_model(prep_model, test)
cp_test = test_kriging_model(c_prep_model, test)
df = pd.DataFrame(data=np.array([r_test, cr_test, p_test, cp_test]).transpose(),
columns=['Raw', 'Raw-cleaned', 'Preprocessed', 'Preprocessed-cleaned'])
fig, ax = plt.subplots(nrows=1, ncols=1, figsize=(12, 7))
ax.boxplot(df)
ax.set_xticklabels(['Raw',
'Raw-cleaned',
'Preprocessed',
'Preprocessed-cleaned'])
plt.show()
# It is very hard to distinguish any differences in the figure but we can use `.describe()` method of **pandas** to get the columns statistic:
df.describe()
# **Clarification:** In this particular case the final statistics goes hand in hand with the initial assumptions that:
#
# **a)** Raw dataset has lowest error of prediction. The rationale behind it is that if we throw away the observations at the preprocessing step we risk an information lost. It could damage our model.
#
# **b)** Raw dataset with the **cleaned variogram** is the best one. We have removed the point pairs with the largest error. In reality we got rid off the potentially wrong measurements where at one point elevation is small and it's neighbour is very high.
#
# > **NOTE:** Example in this tutorial is related to the Digital Elevation Model which was preprocessed by the data provider (*Copernicus Land Monitoring Services*). You shouldn't get impression that the raw data preprocessing and filtering is not required for the analysis. There are cases where the sensor may produce unreliable and biased results, as example a saturated pixel from the satellite camera. It is better to remove it with the specific noise-filtering algorithm before the variogram point cloud development.
# ---
| docs/build/doctrees/nbsphinx/tutorials/Outliers and Their Influence on the Final Model (Intermediate).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # Logging and Messages
#
# `Parameterized` objects provide methods for displaying messages, warnings, and other output in a way that can be controlled and redirected globally using the standard Python [logging](https://docs.python.org/3/library/logging.html) module (see the [logging cookbook](https://docs.python.org/3/howto/logging.html)). Compared to using a Python logger directly, using these methods inside your Parameterized class helps users by making the messages consistent, each prepending with information about the instance where the call was made.
#
# By default, a Python logger named `param` will be instantiated to do the logging, but another logger can be supplied by setting `param.parameterized.logger` to it after importing `param.parameterized`.
#
#
# ## Writing log messages
#
# Each logging message has an associated logging `level` that indicates how severe a condition is being described (DEBUG, VERBOSE, INFO (aka "message"), WARNING, ERROR, or CRITICAL). These levels are as defined by the logging module, except for the addition of VERBOSE as a level intermediate between DEBUG (internal debugging information) and INFO (user-relevant non-error messages).
#
# The typical way to print a message is to call one of the following methods, each accepting the same arguments:
#
# - `.param.debug()`: Detailed debugging information, not displayed onscreen by default.
# - `.param.verbose()`: Additional sometimes-useful information, not displayed onscreen by default.
# - `.param.message()`: Informative message, displayed onscreen by default.
# - `.param.warning()`: Warning of an unexpected or inappropriate but non-fatal condition, displayed onscreen by default.
#
# The arguments accepted are the same as those of [logging.debug()](https://docs.python.org/3/library/logging.html#logging.debug). Specifically, each call is like `.param.debug(msg,*args,**kw)`, where `msg` is an [old-style ('%') format string](https://wiki.python.org/moin/StringFormatting) and the `args` and `kwargs` will be merged with that format string. E.g.:
# +
import param
desired = 1
actual = 5
param.main.param.message("Welcome!")
param.main.param.verbose("Local variables: %s", locals())
param.main.param.warning("Value %02d is not %d", actual, desired)
# -
# Here we've used the default global Parameterized object `param.main`, useful for generic module-level messages, but more often you would make a call like `self.param.warning()` _inside_ a Parameterized class instead. You can see that the messages are each prefixed by the logging level, `param` (the name of the default logger), and the name of this object (`main` in this case). You can also see that, by default, verbose messages are not actually printed.
#
# You may wonder (a) why the formatting string is "old style" , and (b) why the formatting values "actual, desired" are not combined directly with the formatting string. I.e., why not just use a Python3 f-string, like:
param.main.param.warning(f"Value {actual:02} is not {desired}") # Discouraged!
# The answer is that particularly for `debug` and `verbose` messages that could occur inside frequently executed code, we want logging to be "lazy", in that we do not want to render a string representation for `actual`, `desired`, etc. unless we are actually printing the message. If we use an f-string or any other locally formatted string, the string formatting will be done _whether_ _or_ _not_ the message will be displayed, potentially causing drastic slowdowns in your code. For instance, in the code above, the entire `locals()` dictionary would be iterated over, printed to strings. Of course, since the message isn't being printed in that case, the entire format string would then be discarded, greatly slowing down the code without producing any output. So, even though it is more awkward, it is highly recommended to use this old-style, lazy string formatting support.
# ## Controlling the logging level
#
# You can use the `param.parameterized.logging_level` context manager to temporarily reduce or elevate the logging level while you execute code:
with param.parameterized.logging_level('CRITICAL'):
param.main.param.message("Message 1")
param.main.param.verbose("Verbose 1")
with param.parameterized.logging_level('DEBUG'):
param.main.param.message("Message 2")
param.main.param.verbose("Verbose 2")
# You can also set the value more globally (and permanently) on the logger object:
param.parameterized.get_logger().setLevel(param.parameterized.DEBUG)
param.main.param.message("Message 2")
param.main.param.verbose("Verbose 2")
# For continuous integration (CI) or other specific applications, you can also set `param.parameterized.warnings_as_exceptions = True`, which will cause your program to raise an exception the first time it encounters a warning.
# # Controlling the formatting of log messages
#
# The Python logging module provides many options for configuring how the log messages are generated. For complete control, you can instantiate your own logger and set `param.parameterized.logger` to it after importing `param.parameterized`.
#
# A hook is provided for the relatively common case when you want to prefix each message with the time of day, a progress indication, a simulator time, or some other indication of a global state. Specifically, you can set `param.parameterized.dbprint_prefix` to a callable object returning a string. The object will be called when constructing each message:
# +
from datetime import datetime
param.parameterized.dbprint_prefix=lambda: str(datetime.now())
param.main.param.warning("Message 4")
param.main.param.warning("Message 5")
# -
# ## Counting warnings
#
# Typically, a program will abort if an error is encountered, making such a condition hard to miss, but warning messages might be lost in a sea of informational, verbose, or debug messages. Param keeps track of how many times `.param.warning()` has been called, and it is often useful to print out that value at the end of a program run:
print(f"Run completed. {param.parameterized.warning_count} warnings were encountered.")
| examples/user_guide/Logging_and_Messages.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Intro
#
# At the end of this lesson, you will be able to write TensorFlow and Keras code to use one of the best models in computer vision.
#
# # Lesson
#
# + _kg_hide-input=true
from IPython.display import YouTubeVideo
YouTubeVideo('sDG5tPtsbSA', width=800, height=450)
# -
# # Sample Code
#
# ### Choose Images to Work With
# +
from os.path import join
image_dir = '../input/dog-breed-identification/train/'
img_paths = [join(image_dir, filename) for filename in
['0c8fe33bd89646b678f6b2891df8a1c6.jpg',
'0c3b282ecbed1ca9eb17de4cb1b6e326.jpg',
'04fb4d719e9fe2b6ffe32d9ae7be8a22.jpg',
'0e79be614f12deb4f7cae18614b7391b.jpg']]
# -
# ### Function to Read and Prep Images for Modeling
# +
import numpy as np
from tensorflow.python.keras.applications.resnet50 import preprocess_input
from tensorflow.python.keras.preprocessing.image import load_img, img_to_array
image_size = 224
def read_and_prep_images(img_paths, img_height=image_size, img_width=image_size):
imgs = [load_img(img_path, target_size=(img_height, img_width)) for img_path in img_paths]
img_array = np.array([img_to_array(img) for img in imgs])
output = preprocess_input(img_array)
return(output)
# -
# ### Create Model with Pre-Trained Weights File. Make Predictions
# +
from tensorflow.python.keras.applications import ResNet50
my_model = ResNet50(weights='../input/resnet50/resnet50_weights_tf_dim_ordering_tf_kernels.h5')
test_data = read_and_prep_images(img_paths)
preds = my_model.predict(test_data)
# -
# ### Visualize Predictions
# +
from learntools.deep_learning.decode_predictions import decode_predictions
from IPython.display import Image, display
most_likely_labels = decode_predictions(preds, top=3, class_list_path='../input/resnet50/imagenet_class_index.json')
for i, img_path in enumerate(img_paths):
display(Image(img_path))
print(most_likely_labels[i])
# -
# # Exercise
# Now you are ready to **[use a powerful TensorFlow model](#$EXERCISE_FORKING_URL$)** yourself.
| notebooks/deep_learning/raw/tut3_programming_tf_and_keras.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pickle
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
import os
from scipy.stats import pearsonr
# # Load Data
# test
order = pickle.load(open('C:/Users/Vanda/PycharmProjects/dense/results/nppmi/order/glove.6B.400k.300d.txt_f_conceptnet56_top50000_base_order.p', 'rb'))
map_c = pickle.load(open('C:/Users/Vanda/PycharmProjects/dense/results/evaluation/cummulated/glove.6B.400k.300d.txt_f_conceptnet56_top50000_map_cummulated.p', 'rb'))
order
class EvalData(object):
def __init__(self, order, mp_c, map_c, mrr_c, tsa_base_c, tsa_concept_c, name):
self.order = order
self.mp = mp_c
self.map = map_c
self.mrr = mrr_c
self.tsa_base = tsa_base_c
self.tsa_concept = tsa_concept_c
self.name = name
def load_data(embedding_name, dense=False):
if dense:
folder = "dense"
else:
folder = "all"
order_dir = 'C:/Users/Vanda/PycharmProjects/' + folder + '/results/nppmi/order/'
cummulated_dir = 'C:/Users/Vanda/PycharmProjects/' + folder + '/results/evaluation/cummulated/'
order = pickle.load(open(order_dir + embedding_name + '_base_order.p', 'rb'))
mp_c = pickle.load(open(cummulated_dir + embedding_name + '_mp_cummulated.p', 'rb'))
map_c = pickle.load(open(cummulated_dir + embedding_name + '_map_cummulated.p', 'rb'))
mrr_c = pickle.load(open(cummulated_dir + embedding_name + '_mrr_r_cummulated.p', 'rb'))
tsa_base_c = pickle.load(open(cummulated_dir + embedding_name + '_tsa_base_k_acc_cummulated.p', 'rb'))
tsa_concept_c = pickle.load(open(cummulated_dir + embedding_name + '_tsa_concept_k_acc_cummulated.p', 'rb'))
cd = EvalData(order, mp_c, map_c, mrr_c, tsa_base_c, tsa_concept_c, embedding_name)
return cd
dl_400000_cNet = load_data('glove300d_l_0.5_DL_top400000.emb.gz_f_conceptnet56_top50000')
gs_400000_cNet = load_data('glove300d_l_0.5_GS_top400000.emb.gz_f_conceptnet56_top50000')
kmeans_400000_cNet = load_data('glove300d_l_0.5_kmeans_top400000.emb.gz_f_conceptnet56_top50000')
dense = load_data('glove.6B.400k.300d.txt_f_conceptnet56_top50000', True)
# # Cummulate Data
class CummulatedData(object):
def __init__(self, data, point):
self.data = data
self.point = point
def cummulate(ev_metric_c, order):
ev_cummulated = []
found = False
point = len(order)
for i in range(len(ev_metric_c)):
size = i
base_id = order[i][0]
ppmi_val = order[i][1]
if size == 0:
ev_cummulated.append(ev_metric_c[base_id])
else:
prev = ev_cummulated[size-1]
ev_cummulated.append(prev+ev_metric_c[base_id])
if not found and ppmi_val <= 0:
found = True
point = i
if point != len(ev_metric_c)-1:
point = point-1
for i in range(len(ev_cummulated)):
size = i+1
ev_cummulated[i] = ev_cummulated[i]/size
return CummulatedData(ev_cummulated, point)
def cummulate2(ev_metric_c, order):
ev_cummulated1 = []
ev_cummulated2 = []
found = False
point = len(ev_metric_c)-1
for i in range(len(ev_metric_c)):
size = i
base_id = order[i][0]
ppmi_val = order[i][1]
values = ev_metric_c[base_id]
if size == 0:
ev_cummulated1.append(values[0])
ev_cummulated2.append(values[1])
else:
prev1 = ev_cummulated1[size-1]
prev2 = ev_cummulated2[size-1]
ev_cummulated1.append(prev1+values[0])
ev_cummulated2.append(prev2+values[1])
if not found and ppmi_val <= 0:
found = True
point = i
if point != len(ev_metric_c)-1:
point = point-1
for i in range(len(ev_cummulated1)):
size = i+1
ev_cummulated1[i] = ev_cummulated1[i]/size
ev_cummulated2[i] = ev_cummulated2[i]/size
return CummulatedData(ev_cummulated1, point), CummulatedData(ev_cummulated2, point)
def cummulate_concepts(ev_metric):
ev_1 = sorted([v1 for v1, v2 in ev_metric], reverse=True)
ev_2 = sorted([v2 for v1, v2 in ev_metric], reverse=True)
ev_cummulated1 = []
ev_cummulated2 = []
point = -1
for i in range(len(ev_1)):
size = i
if size == 0:
ev_cummulated1.append(ev_1[i])
ev_cummulated2.append(ev_2[i])
else:
prev1 = ev_cummulated1[size-1]
prev2 = ev_cummulated2[size-1]
ev_cummulated1.append(prev1+ev_1[i])
ev_cummulated2.append(prev2+ev_2[i])
for i in range(len(ev_cummulated1)):
size = i+1
ev_cummulated1[i] = ev_cummulated1[i]/size
ev_cummulated2[i] = ev_cummulated2[i]/size
return CummulatedData(ev_cummulated1, point), CummulatedData(ev_cummulated2, point)
# # Plot Data
def gather_metrics(data):
mp = cummulate(data.mp, data.order)
mapp = cummulate(data.map, data.order)
mrr, mr = cummulate2(data.mrr, data.order)
tsa_base, tsa_base_k = cummulate2(data.tsa_base, data.order)
tsa_concept, tsa_concept_k = cummulate_concepts(data.tsa_concept)
return mp, mapp, mrr, mr, tsa_base, tsa_base_k, tsa_concept, tsa_concept_k
# +
def plot_metric_by_dictionary(metric1, metric2, metric3, metric4, metric_name, title, names):
markersize = 10
markeredgewidth = 3
zero1 = metric1.data[0:metric1.point]
zero2 = metric2.data[0:metric2.point]
zero3 = metric3.data[0:metric3.point]
zero4 = metric4.data[0:metric4.point]
# print(len(zero1), len(zero2), len(zero3), print(zero4))
plt.plot(metric1.data, 'r')
plt.plot(metric3.data, 'g')
plt.plot(metric2.data, 'b')
plt.plot(metric4.data, 'c')
plt.plot(zero1, 'r')
plt.plot(zero3, 'g')
plt.plot(zero2, 'b')
plt.plot(zero4, 'c')
if metric1.point != -1:
plt.plot(metric1.point, metric1.data[metric1.point], 'rx', ms=markersize, mew=markeredgewidth)
plt.plot(metric2.point, metric2.data[metric2.point], 'bx', ms=markersize, mew=markeredgewidth)
plt.plot(metric3.point, metric3.data[metric3.point], 'gx', ms=markersize, mew=markeredgewidth)
plt.plot(metric4.point, metric4.data[metric4.point], 'cx', ms=markersize, mew=markeredgewidth)
# plt.ylabel(metric_name, fontsize=20)
if metric_name.find('concept')!=-1:
plt.xlabel("#concepts", fontsize=20)
else:
plt.xlabel("#bases", fontsize=20)
plt.title(title, fontsize=20)
plt.xticks(fontsize=20)
plt.yticks(fontsize=20)
if metric_name == 'MP' or metric_name == 'TAB' or metric_name == 'TAC':
plt.legend(names, fontsize=15)
out_dir = '../../../dense/results/evaluation/cummulated/plots/'
if not os.path.exists(out_dir):
os.makedirs(out_dir)
out_name = out_dir + metric_name.replace(' ', '_') + title.replace(' ', '_')
plt.tight_layout()
plt.savefig((out_name+"_by_dict_0.pdf"))
plt.show()
# -
def tac_mean_dev(data):
tac = data.tsa_concept
tac_1 = [v1 for v1, v2 in tac]
print('mean: ', np.mean(tac_1))
print('std dev: ',np.std(tac_1))
tac_mean_dev(dl_400000_cNet)
tac_mean_dev(gs_400000_cNet)
tac_mean_dev(kmeans_400000_cNet)
tac_mean_dev(dense)
def plot_by_dictionary(data1, data2, data3, data4, title = " ", names=['DLSC', 'kmeans', 'GMPO']):
mp1, mapp1, mrr1, mr1, tsa_base1, tsa_base_k1, tsa_concept1, tsa_concept_k1 = gather_metrics(data1)
mp2, mapp2, mrr2, mr2, tsa_base2, tsa_base_k2, tsa_concept2, tsa_concept_k2 = gather_metrics(data2)
mp3, mapp3, mrr3, mr3, tsa_base3, tsa_base_k3, tsa_concept3, tsa_concept_k3 = gather_metrics(data3)
mp4, mapp4, mrr4, mr4, tsa_base4, tsa_base_k4, tsa_concept4, tsa_concept_k4 = gather_metrics(data4)
plot_metric_by_dictionary(mp1, mp2, mp3, mp4, "MP", title, names)
plot_metric_by_dictionary(mapp1, mapp2, mapp3, mapp4, "MAP", title, names)
plot_metric_by_dictionary(mrr1, mrr2, mrr3, mrr4, "MRR", title, names)
plot_metric_by_dictionary(tsa_base1, tsa_base2, tsa_base3, tsa_base4, "TAB", title, names)
plot_metric_by_dictionary(tsa_base_k1, tsa_base_k2, tsa_base_k3, tsa_base_k4, "TAB_k", title,names)
plot_metric_by_dictionary(tsa_concept1, tsa_concept2, tsa_concept3, tsa_concept4, "TAC", title, names)
plot_metric_by_dictionary(tsa_concept_k1, tsa_concept_k2, tsa_concept_k3, tsa_concept_k4, "TAC_k", title, names)
plot_by_dictionary(dense, dl_400000_cNet, gs_400000_cNet, kmeans_400000_cNet, names=['dense', 'DL', 'GS', 'kmeans'])
def plot_reg_coeffs(metric1, metric2, metric_name, title):
markersize = 10
plt.plot(metric1.data, 'r')
plt.plot(metric2.data, 'b')
print(metric1.point, metric2.point)
if metric1.point != -1:
plt.plot(metric1.point, metric1.data[metric1.point-1], 'rx', ms=markersize)
plt.plot(metric2.point, metric2.data[metric2.point-1], 'bx', ms=markersize)
# plt.ylabel(metric_name, fontsize=20)
if metric_name.find('concept')!=-1:
plt.xlabel("#concepts", fontsize=20)
else:
plt.xlabel("#bases", fontsize=20)
plt.title(title, fontsize=20)
plt.legend(['0.1', '0.5'], fontsize=18)
plt.xticks(fontsize=18)
plt.yticks(fontsize=18)
plt.tight_layout()
out_name = metric_name.replace(' ', '_') + '_' + title.replace(' ', '_')
plt.savefig((out_name+"_reg_coeffs.pdf"))
plt.show()
def compare_reg_coeffs(data1, data2, title):
mp1, mapp1, mrr1, mr1, tsa_base1, tsa_base_k1, tsa_concept1, tsa_concept_k1 = gather_metrics(data1)
mp2, mapp2, mrr2, mr2, tsa_base2, tsa_base_k2, tsa_concept2, tsa_concept_k2 = gather_metrics(data2)
plot_reg_coeffs(mp1, mp2, 'MP', ('Regularization coefficients, ' + title))
plot_reg_coeffs(mapp1, mapp2, 'MAP', ('Regularization coefficients, ' + title))
plot_reg_coeffs(mrr1, mrr2, 'MRR', ('Regularization coefficients, ' + title))
plot_reg_coeffs(tsa_base1, tsa_base2, 'TAB', ('Regularization coefficients, ' + title))
plot_reg_coeffs(tsa_concept1, tsa_concept2, 'TAC', ('Regularization coefficients, ' + title))
dl_400000_cNet_01 = load_data('glove300d_l_0.1_DL_top400000.emb.gz_f_conceptnet56_top50000')
dl_400000_cNet_05 = load_data('glove300d_l_0.5_DL_top400000.emb.gz_f_conceptnet56_top50000')
gs_400000_cNet_01 = load_data('glove300d_l_0.1_GS_top400000.emb.gz_f_conceptnet56_top50000')
gs_400000_cNet_05 = load_data('glove300d_l_0.5_GS_top400000.emb.gz_f_conceptnet56_top50000')
kmeans_400000_cNet_01 = load_data('glove300d_l_0.1_kmeans_top400000.emb.gz_f_conceptnet56_top50000')
kmeans_400000_cNet_05 = load_data('glove300d_l_0.5_kmeans_top400000.emb.gz_f_conceptnet56_top50000')
# kmeans_400000_cNet_01 = load_data('glove300d_l_0.1_kmeans_top400000.emb.gz_f_conceptnet56_top50000')
compare_reg_coeffs(dl_400000_cNet_01, dl_400000_cNet_05, 'DL')
compare_reg_coeffs(gs_400000_cNet_01, gs_400000_cNet_05, 'GS')
compare_reg_coeffs(kmeans_400000_cNet_01, kmeans_400000_cNet_05, 'kmeans')
# # F-score
dl_400000_cNet = load_data('glove300d_l_0.5_DL_top400000.emb.gz_f_conceptnet56_top50000')
gs_400000_cNet = load_data('glove300d_l_0.5_GS_top400000.emb.gz_f_conceptnet56_top50000')
kmeans_400000_cNet = load_data('glove300d_l_0.5_kmeans_top400000.emb.gz_f_conceptnet56_top50000')
dense = load_data('glove.6B.400k.300d.txt_f_conceptnet56_top50000', True)
def fscore(metric1=dl_400000_cNet.map, metric2=dl_400000_cNet.tsa_base):
precision = np.mean(metric1)
recall = np.mean(metric2)
fscore = 2*precision*recall/(precision+recall)
return fscore
print('DL:',fscore(dl_400000_cNet.map, dl_400000_cNet.tsa_base))
print('GS:',fscore(gs_400000_cNet.map, gs_400000_cNet.tsa_base))
print('kmeans:',fscore(kmeans_400000_cNet.map, kmeans_400000_cNet.tsa_base))
print('dense:',fscore(dense.map, dense.tsa_base))
# # ROC curve
def plot_roc_curve(fpr, tpr):
plt.plot(fpr, tpr, color='orange', label='ROC')
plt.plot([0, 1], [0, 1], color='darkblue', linestyle='--')
plt.xlabel('False Positive Rate', fontsize=15)
plt.ylabel('True Positive Rate', fontsize=15)
plt.title('ROC Curve', fontsize=15)
plt.legend(fontsize=13)
out_name = 'ROC_curve.pdf'
plt.savefig(out_name)
plt.show()
def get_recall(data):
return np.mean(data.tsa_base)
# true positive rate = recall, false positive rate = fp/(fp+tn)
# order: dl, gs, kmeans, dense
fpr = [0.0, 0.16539197170583445, 0.1744726213493775, 0.23957202606644293, 0.7367252791828194, 1.0]
tpr = [0.0, get_recall(dl_400000_cNet), get_recall(gs_400000_cNet), get_recall(kmeans_400000_cNet), get_recall(dense), 1.0]
tpr
plot_roc_curve(fpr, tpr)
| src/sparse_alignments/plot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Load text
import tensorflow as tf
import tensorflow_datasets as tfds
import os
physical_devices = tf.config.list_physical_devices('GPU')
try:
tf.config.experimental.set_memory_growth(physical_devices[0], True)
except:
print('Invalid device or cannot modify virtual devices once initialized.')
# +
DIRECTORY_URL = 'https://storage.googleapis.com/download.tensorflow.org/data/illiad/'
FILE_NAMES = ['cowper.txt', 'derby.txt', 'butler.txt']
for name in FILE_NAMES:
text_dir = tf.keras.utils.get_file(name, origin=DIRECTORY_URL+name)
parent_dir = os.path.dirname(text_dir)
parent_dir
# -
# ## Load text into datasets
# +
# This will iterate over every example in the dataset, returning (example, label) pairs.
def labeler(example, index):
return example, tf.cast(index, tf.int64)
labeled_data_sets = []
for i, file_name in enumerate(FILE_NAMES):
lines_dataset = tf.data.TextLineDataset(os.path.join(parent_dir, file_name))
labeled_dataset = lines_dataset.map(lambda ex: labeler(ex, i))
labeled_data_sets.append(labeled_dataset)
# +
# Combine these labeled datasets into a single dataset, and shuffle it.
BUFFER_SIZE = 50000
BATCH_SIZE = 64
TAKE_SIZE = 5000
all_labeled_data = labeled_data_sets[0]
for labeled_dataset in labeled_data_sets[1:]:
all_labeled_data = all_labeled_data.concatenate(labeled_dataset)
all_labeled_data = all_labeled_data.shuffle(
BUFFER_SIZE, reshuffle_each_iteration=False)
# -
for ex in all_labeled_data.take(7):
print(ex[0].numpy().decode())
# ## Encode text lines as numbers
# ### Build vocabulary
# +
tokenizer = tfds.features.text.Tokenizer()
vocabulary_set = set()
for text_tensor, _ in all_labeled_data:
some_tokens = tokenizer.tokenize(text_tensor.numpy())
vocabulary_set.update(some_tokens)
vocab_size = len(vocabulary_set)
vocab_size
# -
# ### Encode examples
encoder = tfds.features.text.TokenTextEncoder(vocabulary_set)
example_text = next(iter(all_labeled_data))[0].numpy()
print(example_text.decode())
encoded_example = encoder.encode(example_text)
print(encoded_example)
def encode(text_tensor, label):
encoded_text = encoder.encode(text_tensor.numpy())
return encoded_text, label
# +
def encode_map_fn(text, label):
# py_func doesn't set the shape of the returned tensors.
encoded_text, label = tf.py_function(encode,
inp=[text, label],
Tout=(tf.int64, tf.int64))
# `tf.data.Datasets` work best if all components have a shape set
# so set the shapes manually:
encoded_text.set_shape([None])
label.set_shape([])
return encoded_text, label
all_encoded_data = all_labeled_data.map(encode_map_fn)
# -
# ## Split the dataset into test and train batches
# +
train_data = all_encoded_data.skip(TAKE_SIZE).shuffle(BUFFER_SIZE)
train_data = train_data.padded_batch(BATCH_SIZE, padded_shapes=([None],[]))
test_data = all_encoded_data.take(TAKE_SIZE)
test_data = test_data.padded_batch(BATCH_SIZE, padded_shapes=([None],[]))
# +
sample_text, sample_labels = next(iter(test_data))
print(sample_text[0].numpy(), sample_labels[0].numpy())
# -
vocab_size += 1
# ## Build the model
model = tf.keras.Sequential()
model.add(tf.keras.layers.Embedding(vocab_size, 64))
#model.add(tf.keras.layers.Bidirectional(tf.keras.layers.RNN(tf.keras.layers.LSTMCell(64))))
model.add(tf.keras.layers.Bidirectional(tf.keras.layers.LSTM(64)))
# +
# One or more dense layers.
# Edit the list in the `for` line to experiment with layer sizes.
for units in [64, 64]:
model.add(tf.keras.layers.Dense(units, activation='relu'))
# Output layer. The first argument is the number of labels.
model.add(tf.keras.layers.Dense(3))
# -
model.compile(optimizer='adam',
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy'])
# ## Train the model
model.fit(train_data, epochs=3, validation_data=test_data)
# +
eval_loss, eval_acc = model.evaluate(test_data)
print('\nEval loss: {:.3f}, Eval accuracy: {:.3f}'.format(eval_loss, eval_acc))
# -
| Load text.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/pranjalrawat007/Econometrics/blob/main/OLS.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="9pXuANWr2U36" outputId="d1ad9295-77f1-49fc-e729-49e7974e23bf" colab={"base_uri": "https://localhost:8080/", "height": 279}
from numpy.random import normal, seed
from numpy import dot, shape, identity, round, sqrt, var, delete, diagonal, where, zeros, array, var
from numpy.linalg import inv, matrix_rank
from scipy import stats
# Generate Data
seed(1)
n, k = 100, 10
β = normal(10, 1, (k, 1))
X = normal(0, 1, (n, k))
ε = normal(0, 1, (n, 1))
y = dot(X, β) + ε
X[:, 0:3] = normal(0, 1, (n, 3)) # noise
print(X.shape, y.shape)
# Random Estimates
n, k = X.shape[0], X.shape[1]
b = normal(0, 1, (k, 1))
ŷ = dot(X, b)
e = y - ŷ
RSS = dot(e.T,e)
print(RSS)
# Least Squares
n, k = X.shape[0], X.shape[1]
A = inv(dot(X.T, X))
b = dot(A, dot(X.T, y))
ŷ = dot(X, b)
e = y - ŷ
RSS = dot(e.T, e)
P = dot(dot(X, dot(X.T, X)), X.T) #Projection
M = identity(X.shape[0]) - P # Annihilation
TSS = dot(y.T,y) # Total Sum of Squares
ESS = dot(ŷ.T, ŷ) # Explained Sum of Squares
s = sqrt(RSS/(n-k)) # Estimate of SE of Unobserved
b_V = (s ** 2) * inv(dot(X.T, X))
b_se = diagonal(s * sqrt(where(A<0,0,A))).reshape(-1,1) # Estimate of SE of b
t = b/b_se.reshape(-1,1)
R2_UC = ESS/TSS # done with no intercept, to avoid less than zero R2.
R2 = 1 - RSS/(var(y)*n) # done when intercept is included, good!
# Influence of a single row
ia = normal(0,1,(n,))
for i in range(n):
x_ = X[i, :]
ia[i] = dot(dot(x.T, A), x)
# Hypothesis testing under Normality assumption
# Individual tTests on Coefficients
b_test = zeros((10,1)) # our guess of true Beta i.e Null Hyp
df = n-k # degree of freedom
tstat = (b - b_test)/b_se # t-statistic for Null Hyp
α = 0.05 # level of signifiance
c = stats.t.ppf(1-α/2, df) # t-critical values
where(abs(tstat)>c,1,0) # confirm or reject
lower_conf = b - b_se * c # Confidence intervals
upper_conf = b + b_se * c
# P-values - give true null, the prob of obtaining a more extreme t than currently obtained
cdf_bel = stats.t.cdf(abs(tstat), df) # prob of less extreme t
cdf_abv = 1 - cdf_bel # prob of more extreme t
p = cdf_abv*2 # as this is a two tailed test
round(p, 2)
# Wald F-Test for systemic hypothesis "Ho: dot(R,β) = r"
# our Ho: all coeffs are 0
R = identity(10)
r = zeros((1,10)).T
print(R)
print(r)
t1 = dot(R,b) - r
t2 = inv(dot(R, dot(b_varcov, R.T)))
Fstat = dot(t1.T, dot(t2,t1))/(matrix_rank(R))
cdf_bel = stats.f.cdf(Fstat, matrix_rank(R), n - k)
cdf_abv = 1 - cdf_bel # one tailed test
p = cdf_abv*2
print(p, Fstat)
# GLS
X_V = cov(X)
# Check
from statsmodels.api import OLS
model = sm.OLS(y, X)
result = model.fit()
result.summary()
R2
# + id="nGbaiNjS2Wbl"
| OLS.ipynb |