code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Reassign ID in AO
# This notebook remap ID in annotation ontology (AO). ID range changes from uint32 to uint16.
#
# - inputs
# - RL_VC.json
# - annotation_100_segmented_combined_gene_fiber_LR.nrrd
# - outputs
# - RL_VC_reID.json
# - LR_reID.nrrd
# - IDnameVC_RL.csv
# +
dir_data = 'data'
# forFigs
fn_input_AO = '1_VC.json'
fn_output_AO = '1_VC_remapID.json'
fn_output_csv = 'remapIDpairs_base.csv'
# -
import os
import pandas as pd
import json
import copy
from collections import OrderedDict
from jsonpath_rw import jsonpath, parse
# # Load data
with open(os.path.join(dir_data, fn_input_AO)) as f:
df_AO = json.load(f, object_pairs_hook=OrderedDict)
# # Reassign ID in AO
def reassign_ID(match_id, match_fullpath):
global df_AOnew
global temp_ID
old_ID = match_id
new_ID = temp_ID
temp_ID +=1
exec("df_AOnew['msg'][0]"+\
str(match_fullpath).replace('.','')\
.replace('children', "['children']").replace('id', "['id']")+\
"= "+ str(new_ID))
exec("voxel_count = df_AOnew['msg'][0]"+\
str(match_fullpath).replace('.','').\
replace('children', "['children']").\
replace('id', "")+"['voxel_count']", globals())
exec("acronym = df_AOnew['msg'][0]"+\
str(match_fullpath).replace('.','').\
replace('children', "['children']").\
replace('id', "")+"['acronym']", globals())
exec("name = df_AOnew['msg'][0]"+\
str(match_fullpath).replace('.','').\
replace('children', "['children']").\
replace('id', "")+"['name']", globals())
exec("color_hex = df_AOnew['msg'][0]"+\
str(match_fullpath).replace('.','').\
replace('children', "['children']").\
replace('id', "")+"['color_hex_triplet']", globals())
parent_id = eval("df_AOnew['msg'][0]"+\
str(match_fullpath)[0:str(match_fullpath).rfind(".children")+1]\
.replace('.','').replace('children',"['children']")+"['id']")
if match_id == 4000000000:# manually assigned root id
parent_id = None
exec("df_AOnew['msg'][0]"+\
str(match_fullpath).replace('.','')\
.replace('children', "['children']").replace('id', "['parent_structure_id']")+\
"= "+ str(parent_id))
return [old_ID, new_ID, acronym, name, voxel_count, color_hex]
# +
jsonpath_expr = parse('$..id')
df_AOnew = copy.deepcopy(df_AO)
temp_ID = 1
oldnewID_list = [reassign_ID(match.value, match.full_path) \
for match in jsonpath_expr.find(df_AO['msg'][0])]
# oldnewID = pd.DataFrame(oldnewID_list, columns = ['old_ID', 'new_ID', 'voxel_count'])
oldnewID = pd.DataFrame(oldnewID_list, columns =\
['old_ID', 'new_ID', 'acronym', 'name', 'voxel_count', 'color'])
# -
# # Save AO with reassigned ID
# +
with open(os.path.join(dir_data, fn_output_AO), mode='w') as fw:
json.dump(df_AOnew, fw, indent=4)
oldnewID.to_csv(os.path.join(dir_data, fn_output_csv), index=False)
# -
# # Check data
oldnewID.info() # 322 non-null
oldnewID
| notebooks/Reassign_ID.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="copyright"
# Copyright 2020 Google LLC
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="title"
# # Vertex client library: Hyperparameter tuning image classification model
#
# <table align="left">
# <td>
# <a href="https://colab.research.google.com/github/GoogleCloudPlatform/vertex-ai-samples/blob/master/notebooks/community/gapic/custom/showcase_hyperparmeter_tuning_image_classification.ipynb">
# <img src="https://cloud.google.com/ml-engine/images/colab-logo-32px.png" alt="Colab logo"> Run in Colab
# </a>
# </td>
# <td>
# <a href="https://github.com/GoogleCloudPlatform/vertex-ai-samples/blob/master/notebooks/community/gapic/custom/showcase_hyperparmeter_tuning_image_classification.ipynb">
# <img src="https://cloud.google.com/ml-engine/images/github-logo-32px.png" alt="GitHub logo">
# View on GitHub
# </a>
# </td>
# </table>
# <br/><br/><br/>
# + [markdown] id="overview:custom,hpt"
# ## Overview
#
# This tutorial demonstrates how to use the Vertex client library for Python to do hyperparameter tuning for a custom image classification model.
# + [markdown] id="dataset:custom,cifar10,icn"
# ### Dataset
#
# The dataset used for this tutorial is the [CIFAR10 dataset](https://www.tensorflow.org/datasets/catalog/cifar10) from [TensorFlow Datasets](https://www.tensorflow.org/datasets/catalog/overview). The version of the dataset you will use is built into TensorFlow. The trained model predicts which type of class an image is from ten classes: airplane, automobile, bird, cat, deer, dog, frog, horse, ship, truck.
# + [markdown] id="objective:custom,hpt"
# ### Objective
#
# In this notebook, you learn how to create a hyperparameter tuning job for a custom image classification model from a Python script in a docker container using the Vertex client library. You can alternatively hyperparameter tune models using the `gcloud` command-line tool or online using the Google Cloud Console.
#
#
# The steps performed include:
#
# - Create an Vertex hyperparameter turning job for training a custom model.
# - Tune the custom model.
# - Evaluate the study results.
# + [markdown] id="costs"
# ### Costs
#
# This tutorial uses billable components of Google Cloud (GCP):
#
# * Vertex AI
# * Cloud Storage
#
# Learn about [Vertex AI
# pricing](https://cloud.google.com/vertex-ai/pricing) and [Cloud Storage
# pricing](https://cloud.google.com/storage/pricing), and use the [Pricing
# Calculator](https://cloud.google.com/products/calculator/)
# to generate a cost estimate based on your projected usage.
# + [markdown] id="install_aip"
# ## Installation
#
# Install the latest version of Vertex client library.
# + id="install_aip"
import os
import sys
# Google Cloud Notebook
if os.path.exists("/opt/deeplearning/metadata/env_version"):
USER_FLAG = "--user"
else:
USER_FLAG = ""
# ! pip3 install -U google-cloud-aiplatform $USER_FLAG
# + [markdown] id="install_storage"
# Install the latest GA version of *google-cloud-storage* library as well.
# + id="install_storage"
# ! pip3 install -U google-cloud-storage $USER_FLAG
# + [markdown] id="restart"
# ### Restart the kernel
#
# Once you've installed the Vertex client library and Google *cloud-storage*, you need to restart the notebook kernel so it can find the packages.
# + id="restart"
if not os.getenv("IS_TESTING"):
# Automatically restart kernel after installs
import IPython
app = IPython.Application.instance()
app.kernel.do_shutdown(True)
# + [markdown] id="before_you_begin"
# ## Before you begin
#
# ### GPU runtime
#
# *Make sure you're running this notebook in a GPU runtime if you have that option. In Colab, select* **Runtime > Change Runtime Type > GPU**
#
# ### Set up your Google Cloud project
#
# **The following steps are required, regardless of your notebook environment.**
#
# 1. [Select or create a Google Cloud project](https://console.cloud.google.com/cloud-resource-manager). When you first create an account, you get a $300 free credit towards your compute/storage costs.
#
# 2. [Make sure that billing is enabled for your project.](https://cloud.google.com/billing/docs/how-to/modify-project)
#
# 3. [Enable the Vertex APIs and Compute Engine APIs.](https://console.cloud.google.com/flows/enableapi?apiid=ml.googleapis.com,compute_component)
#
# 4. [The Google Cloud SDK](https://cloud.google.com/sdk) is already installed in Google Cloud Notebook.
#
# 5. Enter your project ID in the cell below. Then run the cell to make sure the
# Cloud SDK uses the right project for all the commands in this notebook.
#
# **Note**: Jupyter runs lines prefixed with `!` as shell commands, and it interpolates Python variables prefixed with `$` into these commands.
# + id="set_project_id"
PROJECT_ID = "[your-project-id]" # @param {type:"string"}
# + id="autoset_project_id"
if PROJECT_ID == "" or PROJECT_ID is None or PROJECT_ID == "[your-project-id]":
# Get your GCP project id from gcloud
# shell_output = !gcloud config list --format 'value(core.project)' 2>/dev/null
PROJECT_ID = shell_output[0]
print("Project ID:", PROJECT_ID)
# + id="set_gcloud_project_id"
# ! gcloud config set project $PROJECT_ID
# + [markdown] id="region"
# #### Region
#
# You can also change the `REGION` variable, which is used for operations
# throughout the rest of this notebook. Below are regions supported for Vertex. We recommend that you choose the region closest to you.
#
# - Americas: `us-central1`
# - Europe: `europe-west4`
# - Asia Pacific: `asia-east1`
#
# You may not use a multi-regional bucket for training with Vertex. Not all regions provide support for all Vertex services. For the latest support per region, see the [Vertex locations documentation](https://cloud.google.com/vertex-ai/docs/general/locations)
# + id="region"
REGION = "us-central1" # @param {type: "string"}
# + [markdown] id="timestamp"
# #### Timestamp
#
# If you are in a live tutorial session, you might be using a shared test account or project. To avoid name collisions between users on resources created, you create a timestamp for each instance session, and append onto the name of resources which will be created in this tutorial.
# + id="timestamp"
from datetime import datetime
TIMESTAMP = datetime.now().strftime("%Y%m%d%H%M%S")
# + [markdown] id="gcp_authenticate"
# ### Authenticate your Google Cloud account
#
# **If you are using Google Cloud Notebook**, your environment is already authenticated. Skip this step.
#
# **If you are using Colab**, run the cell below and follow the instructions when prompted to authenticate your account via oAuth.
#
# **Otherwise**, follow these steps:
#
# In the Cloud Console, go to the [Create service account key](https://console.cloud.google.com/apis/credentials/serviceaccountkey) page.
#
# **Click Create service account**.
#
# In the **Service account name** field, enter a name, and click **Create**.
#
# In the **Grant this service account access to project** section, click the Role drop-down list. Type "Vertex" into the filter box, and select **Vertex Administrator**. Type "Storage Object Admin" into the filter box, and select **Storage Object Admin**.
#
# Click Create. A JSON file that contains your key downloads to your local environment.
#
# Enter the path to your service account key as the GOOGLE_APPLICATION_CREDENTIALS variable in the cell below and run the cell.
# + id="gcp_authenticate"
# If you are running this notebook in Colab, run this cell and follow the
# instructions to authenticate your GCP account. This provides access to your
# Cloud Storage bucket and lets you submit training jobs and prediction
# requests.
# If on Google Cloud Notebook, then don't execute this code
if not os.path.exists("/opt/deeplearning/metadata/env_version"):
if "google.colab" in sys.modules:
from google.colab import auth as google_auth
google_auth.authenticate_user()
# If you are running this notebook locally, replace the string below with the
# path to your service account key and run this cell to authenticate your GCP
# account.
elif not os.getenv("IS_TESTING"):
# %env GOOGLE_APPLICATION_CREDENTIALS ''
# + [markdown] id="bucket:custom"
# ### Create a Cloud Storage bucket
#
# **The following steps are required, regardless of your notebook environment.**
#
# When you submit a custom training job using the Vertex client library, you upload a Python package
# containing your training code to a Cloud Storage bucket. Vertex runs
# the code from this package. In this tutorial, Vertex also saves the
# trained model that results from your job in the same bucket. You can then
# create an `Endpoint` resource based on this output in order to serve
# online predictions.
#
# Set the name of your Cloud Storage bucket below. Bucket names must be globally unique across all Google Cloud projects, including those outside of your organization.
# + id="bucket"
BUCKET_NAME = "gs://[your-bucket-name]" # @param {type:"string"}
# + id="autoset_bucket"
if BUCKET_NAME == "" or BUCKET_NAME is None or BUCKET_NAME == "gs://[your-bucket-name]":
BUCKET_NAME = "gs://" + PROJECT_ID + "aip-" + TIMESTAMP
# + [markdown] id="create_bucket"
# **Only if your bucket doesn't already exist**: Run the following cell to create your Cloud Storage bucket.
# + id="create_bucket"
# ! gsutil mb -l $REGION $BUCKET_NAME
# + [markdown] id="validate_bucket"
# Finally, validate access to your Cloud Storage bucket by examining its contents:
# + id="validate_bucket"
# ! gsutil ls -al $BUCKET_NAME
# + [markdown] id="setup_vars"
# ### Set up variables
#
# Next, set up some variables used throughout the tutorial.
# ### Import libraries and define constants
# + [markdown] id="import_aip:protobuf"
# #### Import Vertex client library
#
# Import the Vertex client library into our Python environment.
# + id="import_aip:protobuf"
import time
from google.cloud.aiplatform import gapic as aip
from google.protobuf import json_format
from google.protobuf.json_format import MessageToJson, ParseDict
from google.protobuf.struct_pb2 import Struct, Value
# + [markdown] id="aip_constants"
# #### Vertex constants
#
# Setup up the following constants for Vertex:
#
# - `API_ENDPOINT`: The Vertex API service endpoint for dataset, model, job, pipeline and endpoint services.
# - `PARENT`: The Vertex location root path for dataset, model, job, pipeline and endpoint resources.
# + id="aip_constants"
# API service endpoint
API_ENDPOINT = "{}-aiplatform.googleapis.com".format(REGION)
# Vertex location root path for your dataset, model and endpoint resources
PARENT = "projects/" + PROJECT_ID + "/locations/" + REGION
# + id="accelerators:training"
if os.getenv("IS_TESTING_TRAIN_GPU"):
TRAIN_GPU, TRAIN_NGPU = (
aip.AcceleratorType.NVIDIA_TESLA_K80,
int(os.getenv("IS_TESTING_TRAIN_GPU")),
)
else:
TRAIN_GPU, TRAIN_NGPU = (aip.AcceleratorType.NVIDIA_TESLA_K80, 1)
# + [markdown] id="container:training"
# #### Container (Docker) image
#
# Next, we will set the Docker container images for training.
#
# - Set the variable `TF` to the TensorFlow version of the container image. For example, `2-1` would be version 2.1, and `1-15` would be version 1.15. The following list shows some of the pre-built images available:
#
# - TensorFlow 1.15
# - `gcr.io/cloud-aiplatform/training/tf-cpu.1-15:latest`
# - `gcr.io/cloud-aiplatform/training/tf-gpu.1-15:latest`
# - TensorFlow 2.1
# - `gcr.io/cloud-aiplatform/training/tf-cpu.2-1:latest`
# - `gcr.io/cloud-aiplatform/training/tf-gpu.2-1:latest`
# - TensorFlow 2.2
# - `gcr.io/cloud-aiplatform/training/tf-cpu.2-2:latest`
# - `gcr.io/cloud-aiplatform/training/tf-gpu.2-2:latest`
# - TensorFlow 2.3
# - `gcr.io/cloud-aiplatform/training/tf-cpu.2-3:latest`
# - `gcr.io/cloud-aiplatform/training/tf-gpu.2-3:latest`
# - TensorFlow 2.4
# - `gcr.io/cloud-aiplatform/training/tf-cpu.2-4:latest`
# - `gcr.io/cloud-aiplatform/training/tf-gpu.2-4:latest`
# - XGBoost
# - `gcr.io/cloud-aiplatform/training/xgboost-cpu.1-1`
# - Scikit-learn
# - `gcr.io/cloud-aiplatform/training/scikit-learn-cpu.0-23:latest`
# - Pytorch
# - `gcr.io/cloud-aiplatform/training/pytorch-cpu.1-4:latest`
# - `gcr.io/cloud-aiplatform/training/pytorch-cpu.1-5:latest`
# - `gcr.io/cloud-aiplatform/training/pytorch-cpu.1-6:latest`
# - `gcr.io/cloud-aiplatform/training/pytorch-cpu.1-7:latest`
#
# For the latest list, see [Pre-built containers for training](https://cloud.google.com/vertex-ai/docs/training/pre-built-containers).
# + id="container:training"
if os.getenv("IS_TESTING_TF"):
TF = os.getenv("IS_TESTING_TF")
else:
TF = "2-1"
if TF[0] == "2":
if TRAIN_GPU:
TRAIN_VERSION = "tf-gpu.{}".format(TF)
else:
TRAIN_VERSION = "tf-cpu.{}".format(TF)
else:
if TRAIN_GPU:
TRAIN_VERSION = "tf-gpu.{}".format(TF)
else:
TRAIN_VERSION = "tf-cpu.{}".format(TF)
TRAIN_IMAGE = "gcr.io/cloud-aiplatform/training/{}:latest".format(TRAIN_VERSION)
print("Training:", TRAIN_IMAGE, TRAIN_GPU, TRAIN_NGPU)
# + [markdown] id="machine:training"
# #### Machine Type
#
# Next, set the machine type to use for training.
#
# - Set the variable `TRAIN_COMPUTE` to configure the compute resources for the VMs you will use for for training.
# - `machine type`
# - `n1-standard`: 3.75GB of memory per vCPU.
# - `n1-highmem`: 6.5GB of memory per vCPU
# - `n1-highcpu`: 0.9 GB of memory per vCPU
# - `vCPUs`: number of \[2, 4, 8, 16, 32, 64, 96 \]
#
# *Note: The following is not supported for training:*
#
# - `standard`: 2 vCPUs
# - `highcpu`: 2, 4 and 8 vCPUs
#
# *Note: You may also use n2 and e2 machine types for training and deployment, but they do not support GPUs*.
# + id="machine:training"
if os.getenv("IS_TESTING_TRAIN_MACHINE"):
MACHINE_TYPE = os.getenv("IS_TESTING_TRAIN_MACHINE")
else:
MACHINE_TYPE = "n1-standard"
VCPU = "4"
TRAIN_COMPUTE = MACHINE_TYPE + "-" + VCPU
print("Train machine type", TRAIN_COMPUTE)
# + [markdown] id="tutorial_start:custom,hpt"
# # Tutorial
#
# Now you are ready to start creating your own hyperparameter tuning and training of a custom image classification.
# + [markdown] id="clients:custom,hpt"
# ## Set up clients
#
# The Vertex client library works as a client/server model. On your side (the Python script) you will create a client that sends requests and receives responses from the Vertex server.
#
# You will use different clients in this tutorial for different steps in the workflow. So set them all up upfront.
#
# - Model Service for `Model` resources.
# - Job Service for hyperparameter tuning.
# + id="clients:custom,hpt"
# client options same for all services
client_options = {"api_endpoint": API_ENDPOINT}
def create_job_client():
client = aip.JobServiceClient(client_options=client_options)
return client
def create_model_client():
client = aip.ModelServiceClient(client_options=client_options)
return client
clients = {}
clients["job"] = create_job_client()
clients["model"] = create_model_client()
for client in clients.items():
print(client)
# + [markdown] id="tune_custom_model:simple"
# ## Tuning a model - Hello World
#
# There are two ways you can hyperparameter tune and train a custom model using a container image:
#
# - **Use a Google Cloud prebuilt container**. If you use a prebuilt container, you will additionally specify a Python package to install into the container image. This Python package contains your code for hyperparameter tuning and training a custom model.
#
# - **Use your own custom container image**. If you use your own container, the container needs to contain your code for hyperparameter tuning and training a custom model.
# + [markdown] id="train_custom_job_specification:prebuilt_container,hpt"
# ## Prepare your hyperparameter tuning job specification
#
# Now that your clients are ready, your first step is to create a Job Specification for your hyperparameter tuning job. The job specification will consist of the following:
#
# - `trial_job_spec`: The specification for the custom job.
# - `worker_pool_spec` : The specification of the type of machine(s) you will use for hyperparameter tuning and how many (single or distributed)
# - `python_package_spec` : The specification of the Python package to be installed with the pre-built container.
#
# - `study_spec`: The specification for what to tune.
# - `parameters`: This is the specification of the hyperparameters that you will tune for the custom training job. It will contain a list of the
# - `metrics`: This is the specification on how to evaluate the result of each tuning trial.
# + [markdown] id="train_custom_job_machine_specification"
# ### Prepare your machine specification
#
# Now define the machine specification for your custom hyperparameter tuning job. This tells Vertex what type of machine instance to provision for the hyperparameter tuning.
# - `machine_type`: The type of GCP instance to provision -- e.g., n1-standard-8.
# - `accelerator_type`: The type, if any, of hardware accelerator. In this tutorial if you previously set the variable `TRAIN_GPU != None`, you are using a GPU; otherwise you will use a CPU.
# - `accelerator_count`: The number of accelerators.
# + id="train_custom_job_machine_specification"
if TRAIN_GPU:
machine_spec = {
"machine_type": TRAIN_COMPUTE,
"accelerator_type": TRAIN_GPU,
"accelerator_count": TRAIN_NGPU,
}
else:
machine_spec = {"machine_type": TRAIN_COMPUTE, "accelerator_count": 0}
# + [markdown] id="train_custom_job_disk_specification"
# ### Prepare your disk specification
#
# (optional) Now define the disk specification for your custom hyperparameter tuning job. This tells Vertex what type and size of disk to provision in each machine instance for the hyperparameter tuning.
#
# - `boot_disk_type`: Either SSD or Standard. SSD is faster, and Standard is less expensive. Defaults to SSD.
# - `boot_disk_size_gb`: Size of disk in GB.
# + id="train_custom_job_disk_specification"
DISK_TYPE = "pd-ssd" # [ pd-ssd, pd-standard]
DISK_SIZE = 200 # GB
disk_spec = {"boot_disk_type": DISK_TYPE, "boot_disk_size_gb": DISK_SIZE}
# + [markdown] id="train_custom_job_worker_pool_specification:prebuilt_container"
# ### Define the worker pool specification
#
# Next, you define the worker pool specification for your custom hyperparameter tuning job. The worker pool specification will consist of the following:
#
# - `replica_count`: The number of instances to provision of this machine type.
# - `machine_spec`: The hardware specification.
# - `disk_spec` : (optional) The disk storage specification.
#
# - `python_package`: The Python training package to install on the VM instance(s) and which Python module to invoke, along with command line arguments for the Python module.
#
# Let's dive deeper now into the python package specification:
#
# -`executor_image_spec`: This is the docker image which is configured for your custom hyperparameter tuning job.
#
# -`package_uris`: This is a list of the locations (URIs) of your python training packages to install on the provisioned instance. The locations need to be in a Cloud Storage bucket. These can be either individual python files or a zip (archive) of an entire package. In the later case, the job service will unzip (unarchive) the contents into the docker image.
#
# -`python_module`: The Python module (script) to invoke for running the custom hyperparameter tuning job. In this example, you will be invoking `trainer.task.py` -- note that it was not neccessary to append the `.py` suffix.
#
# -`args`: The command line arguments to pass to the corresponding Pythom module. In this example, you will be setting:
# - `"--model-dir=" + MODEL_DIR` : The Cloud Storage location where to store the model artifacts. There are two ways to tell the hyperparameter tuning script where to save the model artifacts:
# - direct: You pass the Cloud Storage location as a command line argument to your training script (set variable `DIRECT = True`), or
# - indirect: The service passes the Cloud Storage location as the environment variable `AIP_MODEL_DIR` to your training script (set variable `DIRECT = False`). In this case, you tell the service the model artifact location in the job specification.
# - `"--epochs=" + EPOCHS`: The number of epochs for training.
# - `"--steps=" + STEPS`: The number of steps (batches) per epoch.
# - `"--distribute=" + TRAIN_STRATEGY"` : The hyperparameter tuning distribution strategy to use for single or distributed hyperparameter tuning.
# - `"single"`: single device.
# - `"mirror"`: all GPU devices on a single compute instance.
# - `"multi"`: all GPU devices on all compute instances.
# + id="train_custom_job_worker_pool_specification:prebuilt_container"
JOB_NAME = "custom_job_" + TIMESTAMP
MODEL_DIR = "{}/{}".format(BUCKET_NAME, JOB_NAME)
if not TRAIN_NGPU or TRAIN_NGPU < 2:
TRAIN_STRATEGY = "single"
else:
TRAIN_STRATEGY = "mirror"
EPOCHS = 20
STEPS = 100
DIRECT = True
if DIRECT:
CMDARGS = [
"--model-dir=" + MODEL_DIR,
"--epochs=" + str(EPOCHS),
"--steps=" + str(STEPS),
"--distribute=" + TRAIN_STRATEGY,
]
else:
CMDARGS = [
"--epochs=" + str(EPOCHS),
"--steps=" + str(STEPS),
"--distribute=" + TRAIN_STRATEGY,
]
worker_pool_spec = [
{
"replica_count": 1,
"machine_spec": machine_spec,
"disk_spec": disk_spec,
"python_package_spec": {
"executor_image_uri": TRAIN_IMAGE,
"package_uris": [BUCKET_NAME + "/trainer_cifar10.tar.gz"],
"python_module": "trainer.task",
"args": CMDARGS,
},
}
]
# + [markdown] id="create_study_spec:simple"
# ### Create a study specification
#
# Let's start with a simple study. You will just use a single parameter -- the *learning rate*. Since its just one parameter, it doesn't make much sense to do a random search. Instead, we will do a grid search over a range of values.
#
# - `metrics`:
# - `metric_id`: In this example, the objective metric to report back is `'val_accuracy'`
# - `goal`: In this example, the hyperparameter tuning service will evaluate trials to maximize the value of the objective metric.
# - `parameters`: The specification for the hyperparameters to tune.
# - `parameter_id`: The name of the hyperparameter that will be passed to the Python package as a command line argument.
# - `scale_type`: The scale type determines the resolution the hyperparameter tuning service uses when searching over the search space.
# - `UNIT_LINEAR_SCALE`: Uses a resolution that is the same everywhere in the search space.
# - `UNIT_LOG_SCALE`: Values close to the bottom of the search space are further away.
# - `UNIT_REVERSE_LOG_SCALE`: Values close to the top of the search space are further away.
# - **search space**: This is where you will specify the search space of values for the hyperparameter to select for tuning.
# - `integer_value_spec`: Specifies an integer range of values between a `min_value` and `max_value`.
# - `double_value_spec`: Specifies a continuous range of values between a `min_value` and `max_value`.
# - `discrete_value_spec`: Specifies a list of values.
# - `algorithm`: The search method for selecting hyperparameter values per trial:
# - `GRID_SEARCH`: Combinatorically search -- which is used in this example.
# - `RANDOM_SEARCH`: Random search.
#
# + id="create_study_spec:simple"
study_spec = {
"metrics": [
{
"metric_id": "val_accuracy",
"goal": aip.StudySpec.MetricSpec.GoalType.MAXIMIZE,
}
],
"parameters": [
{
"parameter_id": "lr",
"discrete_value_spec": {"values": [0.001, 0.01, 0.1]},
"scale_type": aip.StudySpec.ParameterSpec.ScaleType.UNIT_LINEAR_SCALE,
}
],
"algorithm": aip.StudySpec.Algorithm.GRID_SEARCH,
}
# + [markdown] id="assemble_custom_hpt_job_specification"
# ### Assemble a hyperparameter tuning job specification
#
# Now assemble the complete description for the custom hyperparameter tuning specification:
#
# - `display_name`: The human readable name you assign to this custom hyperparameter tuning job.
# - `trial_job_spec`: The specification for the custom hyperparameter tuning job.
# - `study_spec`: The specification for what to tune.
# - `max_trial_count`: The maximum number of tuning trials.
# - `parallel_trial_count`: How many trials to try in parallel; otherwise, they are done sequentially.
# + id="assemble_custom_hpt_job_specification"
hpt_job = {
"display_name": JOB_NAME,
"trial_job_spec": {"worker_pool_specs": worker_pool_spec},
"study_spec": study_spec,
"max_trial_count": 6,
"parallel_trial_count": 1,
}
# + [markdown] id="examine_training_package"
# ### Examine the hyperparameter tuning package
#
# #### Package layout
#
# Before you start the hyperparameter tuning, you will look at how a Python package is assembled for a custom hyperparameter tuning job. When unarchived, the package contains the following directory/file layout.
#
# - PKG-INFO
# - README.md
# - setup.cfg
# - setup.py
# - trainer
# - \_\_init\_\_.py
# - task.py
#
# The files `setup.cfg` and `setup.py` are the instructions for installing the package into the operating environment of the Docker image.
#
# The file `trainer/task.py` is the Python script for executing the custom hyperparameter tuning job. *Note*, when we referred to it in the worker pool specification, we replace the directory slash with a dot (`trainer.task`) and dropped the file suffix (`.py`).
#
# #### Package Assembly
#
# In the following cells, you will assemble the training package.
# + id="examine_training_package"
# Make folder for Python hyperparameter tuning script
# ! rm -rf custom
# ! mkdir custom
# Add package information
# ! touch custom/README.md
setup_cfg = "[egg_info]\n\ntag_build =\n\ntag_date = 0"
# ! echo "$setup_cfg" > custom/setup.cfg
setup_py = "import setuptools\n\nsetuptools.setup(\n\n install_requires=[\n\n 'tensorflow_datasets==1.3.0',\n\n ],\n\n packages=setuptools.find_packages())"
# ! echo "$setup_py" > custom/setup.py
pkg_info = "Metadata-Version: 1.0\n\nName: CIFAR10 image classification\n\nVersion: 0.0.0\n\nSummary: Demostration hyperparameter tuning script\n\nHome-page: www.google.com\n\nAuthor: Google\n\nAuthor-email: <EMAIL>\n\nLicense: Public\n\nDescription: Demo\n\nPlatform: Vertex"
# ! echo "$pkg_info" > custom/PKG-INFO
# Make the training subfolder
# ! mkdir custom/trainer
# ! touch custom/trainer/__init__.py
# + [markdown] id="taskpy_contents:hpt,simple"
# #### Task.py contents
#
# In the next cell, you write the contents of the hyperparameter tuning script task.py. I won't go into detail, it's just there for you to browse. In summary:
#
# - Passes the hyperparameter values for a trial as a command line argument (`parser.add_argument('--lr',...)`)
# - Mimics a training loop, where on each loop (epoch) the variable `accuracy` is set to the loop iteration * the learning rate.
# - Reports back the objective metric `accuracy` back to the hyperparameter tuning service using `report_hyperparameter_tuning_metric()`.
# + id="taskpy_contents:hpt,simple"
# %%writefile custom/trainer/task.py
# HP Tuning hello world example
from __future__ import absolute_import, division, print_function, unicode_literals
import tensorflow_datasets as tfds
import tensorflow as tf
from tensorflow.python.client import device_lib
from hypertune import HyperTune
import argparse
import os
import sys
import time
tfds.disable_progress_bar()
parser = argparse.ArgumentParser()
parser.add_argument('--lr', dest='lr',
default=0.001, type=float,
help='Learning rate.')
parser.add_argument('--epochs', dest='epochs',
default=10, type=int,
help='Number of epochs.')
parser.add_argument('--steps', dest='steps',
default=200, type=int,
help='Number of steps per epoch.')
parser.add_argument('--model-dir',
dest='model_dir',
default='/tmp/saved_model',
type=str,
help='Model dir.')
parser.add_argument('--distribute', dest='distribute', type=str, default='single',
help='distributed training strategy')
args = parser.parse_args()
print('Python Version = {}'.format(sys.version))
print('TensorFlow Version = {}'.format(tf.__version__))
print('TF_CONFIG = {}'.format(os.environ.get('TF_CONFIG', 'Not found')))
print(device_lib.list_local_devices())
# Instantiate the HyperTune reporting object
hpt = HyperTune()
for epoch in range(1, args.epochs+1):
# mimic metric result at the end of an epoch
acc = args.lr * epoch
# save the metric result to communicate back to the HPT service
hpt.report_hyperparameter_tuning_metric(
hyperparameter_metric_tag='val_accuracy',
metric_value=acc,
global_step=epoch)
print('epoch: {}, accuracy: {}'.format(epoch, acc))
time.sleep(1)
# + [markdown] id="tarball_training_script"
# #### Store hyperparameter tuning script on your Cloud Storage bucket
#
# Next, you package the hyperparameter tuning folder into a compressed tar ball, and then store it in your Cloud Storage bucket.
# + id="tarball_training_script"
# ! rm -f custom.tar custom.tar.gz
# ! tar cvf custom.tar custom
# ! gzip custom.tar
# ! gsutil cp custom.tar.gz $BUCKET_NAME/trainer_cifar10.tar.gz
# + [markdown] id="report_hypertune"
# #### Reporting back the result of the trial using hypertune
#
# For each trial, your Python script needs to report back to the hyperparameter tuning service the objective metric for which you specified as the criteria for evaluating the trial.
#
# For this example, you will specify in the study specification that the objective metric will be reported back as `val_accuracy`.
#
# You report back the value of the objective metric using `HyperTune`. This Python module is used to communicate key/value pairs to the hyperparameter tuning service. To setup this reporting in your Python package, you will add code for the following three steps:
#
# 1. Import the HyperTune module: `from hypertune import HyperTune()`.
# 2. At the end of every epoch, write the current value of the objective function to the log as a key/value pair using `hpt.report_hyperparameter_tuning_metric()`. In this example, the parameters are:
# - `hyperparameter_metric_tag`: The name of the objective metric to report back. The name must be identical to the name specified in the study specification.
# - `metric_value`: The value of the objective metric to report back to the hyperparameter service.
# - `global_step`: The epoch iteration, starting at 0.
# + [markdown] id="tune_custom_job"
# ## Hyperparameter Tune the model
#
# Now start the hyperparameter tuning of your custom model on Vertex. Use this helper function `create_hyperparameter_tuning_job`, which takes the following parameter:
#
# -`hpt_job`: The specification for the hyperparameter tuning job.
#
# The helper function calls job client service's `create_hyperparameter_tuning_job` method, with the following parameters:
#
# -`parent`: The Vertex location path to `Dataset`, `Model` and `Endpoint` resources.
# -`hyperparameter_tuning_job`: The specification for the hyperparameter tuning job.
#
# You will display a handful of the fields returned in `response` object, with the two that are of most interest are:
#
# `response.name`: The Vertex fully qualified identifier assigned to this custom hyperparameter tuning job. You save this identifier for using in subsequent steps.
#
# `response.state`: The current state of the custom hyperparameter tuning job.
# + id="tune_custom_job"
def create_hyperparameter_tuning_job(hpt_job):
response = clients["job"].create_hyperparameter_tuning_job(
parent=PARENT, hyperparameter_tuning_job=hpt_job
)
print("name:", response.name)
print("display_name:", response.display_name)
print("state:", response.state)
print("create_time:", response.create_time)
print("update_time:", response.update_time)
return response
response = create_hyperparameter_tuning_job(hpt_job)
# + [markdown] id="hpt_job_id:response"
# Now get the unique identifier for the hyperparameter tuning job you created.
# + id="hpt_job_id:response"
# The full unique ID for the hyperparameter tuning job
hpt_job_id = response.name
# The short numeric ID for the hyperparameter tuning job
hpt_job_short_id = hpt_job_id.split("/")[-1]
print(hpt_job_id)
# + [markdown] id="get_hpt_job"
# ### Get information on a hyperparameter tuning job
#
# Next, use this helper function `get_hyperparameter_tuning_job`, which takes the following parameter:
#
# - `name`: The Vertex fully qualified identifier for the hyperparameter tuning job.
#
# The helper function calls the job client service's `get_hyperparameter_tuning_job` method, with the following parameter:
#
# - `name`: The Vertex fully qualified identifier for the hyperparameter tuning job.
#
# If you recall, you got the Vertex fully qualified identifier for the hyperparameter tuning job in the `response.name` field when you called the `create_hyperparameter_tuning_job` method, and saved the identifier in the variable `hpt_job_id`.
# + id="get_hpt_job"
def get_hyperparameter_tuning_job(name, silent=False):
response = clients["job"].get_hyperparameter_tuning_job(name=name)
if silent:
return response
print("name:", response.name)
print("display_name:", response.display_name)
print("state:", response.state)
print("create_time:", response.create_time)
print("update_time:", response.update_time)
return response
response = get_hyperparameter_tuning_job(hpt_job_id)
# + [markdown] id="wait_tuning_complete"
# ## Wait for tuning to complete
#
# Hyperparameter tuning the above model may take upwards of 20 minutes time.
#
# Once your model is done tuning, you can calculate the actual time it took to tune the model by subtracting `end_time` from `start_time`.
#
# For your model, we will need to know the location of the saved models for each trial, which the Python script saved in your local Cloud Storage bucket at `MODEL_DIR + '/<trial_number>/saved_model.pb'`.
# + id="wait_tuning_complete"
while True:
job_response = get_hyperparameter_tuning_job(hpt_job_id, True)
if job_response.state != aip.JobState.JOB_STATE_SUCCEEDED:
print("Study trials have not completed:", job_response.state)
if job_response.state == aip.JobState.JOB_STATE_FAILED:
break
else:
if not DIRECT:
MODEL_DIR = MODEL_DIR + "/model"
print("Study trials have completed")
break
time.sleep(60)
# + [markdown] id="review_study_results"
# ### Review the results of the study
#
# Now review the results of trials.
# + id="review_study_results"
best = (None, None, None, 0.0)
for trial in job_response.trials:
print(trial)
# Keep track of the best outcome
if float(trial.final_measurement.metrics[0].value) > best[3]:
try:
best = (
trial.id,
float(trial.parameters[0].value),
float(trial.parameters[1].value),
float(trial.final_measurement.metrics[0].value),
)
except:
best = (
trial.id,
float(trial.parameters[0].value),
None,
float(trial.final_measurement.metrics[0].value),
)
# + [markdown] id="best_trial"
# ### Best trial
#
# Now look at which trial was the best:
# + id="best_trial"
print("ID", best[0])
print("Learning Rate", best[1])
print("Decay", best[2])
print("Validation Accuracy", best[3])
# + [markdown] id="get_best_model"
# ## Get the Best Model
#
# If you used the method of having the service tell the tuning script where to save the model artifacts (`DIRECT = False`), then the model artifacts for the best model are saved at:
#
# MODEL_DIR/<best_trial_id>/model
# + id="get_best_model"
BEST_MODEL_DIR = MODEL_DIR + "/" + best[0] + "/model"
# + [markdown] id="tune_custom_model:random"
# ## Tuning a model - CIFAR10
#
# Now that you have seen the overall steps for hyperparameter tuning a custom training job using a Python package that mimics training a model, you will do a new hyperparameter tuning job for a custom training job for a CIFAR10 model.
#
# For this example, you will change two parts:
#
# 1. Specify the CIFAR10 custom hyperparameter tuning Python package.
# 2. Specify a study specification specific to the hyperparameters used in the CIFAR10 custom hyperparameter tuning Python package.
# + [markdown] id="create_study_spec:random"
# ### Create a study specification
#
# In this study, you will tune for two hyperparameters using the random search algorithm:
#
# - **learning rate**: The search space is a set of discrete values.
# - **learning rate decay**: The search space is a continuous range between 1e-6 and 1e-2.
#
# The objective (goal) is to maximize the validation accuracy.
#
# You will run a maximum of six trials.
# + id="create_study_spec:random"
study_spec = {
"metrics": [
{
"metric_id": "val_accuracy",
"goal": aip.StudySpec.MetricSpec.GoalType.MAXIMIZE,
}
],
"parameters": [
{
"parameter_id": "lr",
"discrete_value_spec": {"values": [0.001, 0.01, 0.1]},
"scale_type": aip.StudySpec.ParameterSpec.ScaleType.UNIT_LINEAR_SCALE,
},
{
"parameter_id": "decay",
"double_value_spec": {"min_value": 1e-6, "max_value": 1e-2},
"scale_type": aip.StudySpec.ParameterSpec.ScaleType.UNIT_LINEAR_SCALE,
},
],
"algorithm": aip.StudySpec.Algorithm.RANDOM_SEARCH,
}
# + [markdown] id="assemble_custom_hpt_job_specification"
# ### Assemble a hyperparameter tuning job specification
#
# Now assemble the complete description for the custom hyperparameter tuning specification:
#
# - `display_name`: The human readable name you assign to this custom hyperparameter tuning job.
# - `trial_job_spec`: The specification for the custom hyperparameter tuning job.
# - `study_spec`: The specification for what to tune.
# - `max_trial_count`: The maximum number of tuning trials.
# - `parallel_trial_count`: How many trials to try in parallel; otherwise, they are done sequentially.
# + id="assemble_custom_hpt_job_specification"
hpt_job = {
"display_name": JOB_NAME,
"trial_job_spec": {"worker_pool_specs": worker_pool_spec},
"study_spec": study_spec,
"max_trial_count": 6,
"parallel_trial_count": 1,
}
# + [markdown] id="taskpy_contents:hpt,cifar10"
# #### Task.py contents
#
# In the next cell, you write the contents of the hyperparameter tuning script task.py. I won't go into detail, it's just there for you to browse. In summary:
#
# - Parse the command line arguments for the hyperparameter settings for the current trial.
# - Get the directory where to save the model artifacts from the command line (`--model_dir`), and if not specified, then from the environment variable `AIP_MODEL_DIR`.
# - Download and preprocess the CIFAR10 dataset.
# - Build a CNN model.
# - The learning rate and decay hyperparameter values are used during the compile of the model.
# - A definition of a callback `HPTCallback` which obtains the validation accuracy at the end of each epoch (`on_epoch_end()`) and reports it to the hyperparameter tuning service using `hpt.report_hyperparameter_tuning_metric()`.
# - Train the model with the `fit()` method and specify a callback which will report the validation accuracy back to the hyperparameter tuning service.
# + id="taskpy_contents:hpt,cifar10"
# %%writefile custom/trainer/task.py
# Custom Job for CIFAR10
import tensorflow_datasets as tfds
import tensorflow as tf
from hypertune import HyperTune
import argparse
import os
import sys
# Command Line arguments
parser = argparse.ArgumentParser()
parser.add_argument('--model-dir', dest='model_dir',
default=os.getenv('AIP_MODEL_DIR'), type=str, help='Model dir.')
parser.add_argument('--lr', dest='lr',
default=0.001, type=float,
help='Learning rate.')
parser.add_argument('--decay', dest='decay',
default=0.98, type=float,
help='Decay rate')
parser.add_argument('--epochs', dest='epochs',
default=10, type=int,
help='Number of epochs.')
parser.add_argument('--steps', dest='steps',
default=200, type=int,
help='Number of steps per epoch.')
parser.add_argument('--distribute', dest='distribute', type=str, default='single',
help='distributed training strategy')
args = parser.parse_args()
# Scaling CIFAR-10 data from (0, 255] to (0., 1.]
def scale(image, label):
image = tf.cast(image, tf.float32)
image /= 255.0
return image, label
# Download the dataset
datasets = tfds.load(name='cifar10', as_supervised=True)
# Preparing dataset
BUFFER_SIZE = 10000
BATCH_SIZE = 64
train_dataset = datasets['train'].map(scale).shuffle(BUFFER_SIZE).batch(BATCH_SIZE)
test_dataset = datasets['test'].map(scale).batch(BATCH_SIZE)
# Build the Keras model
def build_and_compile_cnn_model():
model = tf.keras.Sequential([
tf.keras.layers.Conv2D(32, 3, activation='relu', input_shape=(32, 32, 3)),
tf.keras.layers.MaxPooling2D(),
tf.keras.layers.Conv2D(32, 3, activation='relu'),
tf.keras.layers.MaxPooling2D(),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(10, activation='softmax')
])
model.compile(
loss=tf.keras.losses.sparse_categorical_crossentropy,
optimizer=tf.keras.optimizers.SGD(learning_rate=args.lr, decay=args.decay),
metrics=['accuracy'])
return model
model = build_and_compile_cnn_model()
# Instantiate the HyperTune reporting object
hpt = HyperTune()
# Reporting callback
class HPTCallback(tf.keras.callbacks.Callback):
def on_epoch_end(self, epoch, logs=None):
global hpt
hpt.report_hyperparameter_tuning_metric(
hyperparameter_metric_tag='val_accuracy',
metric_value=logs['val_accuracy'],
global_step=epoch)
# Train the model
model.fit(train_dataset, epochs=5, steps_per_epoch=10, validation_data=test_dataset.take(8),
callbacks=[HPTCallback()])
model.save(args.model_dir)
# + [markdown] id="tarball_training_script"
# #### Store hyperparameter tuning script on your Cloud Storage bucket
#
# Next, you package the hyperparameter tuning folder into a compressed tar ball, and then store it in your Cloud Storage bucket.
# + id="tarball_training_script"
# ! rm -f custom.tar custom.tar.gz
# ! tar cvf custom.tar custom
# ! gzip custom.tar
# ! gsutil cp custom.tar.gz $BUCKET_NAME/trainer_cifar10.tar.gz
# + [markdown] id="report_hypertune"
# #### Reporting back the result of the trial using hypertune
#
# For each trial, your Python script needs to report back to the hyperparameter tuning service the objective metric for which you specified as the criteria for evaluating the trial.
#
# For this example, you will specify in the study specification that the objective metric will be reported back as `val_accuracy`.
#
# You report back the value of the objective metric using `HyperTune`. This Python module is used to communicate key/value pairs to the hyperparameter tuning service. To setup this reporting in your Python package, you will add code for the following three steps:
#
# 1. Import the HyperTune module: `from hypertune import HyperTune()`.
# 2. At the end of every epoch, write the current value of the objective function to the log as a key/value pair using `hpt.report_hyperparameter_tuning_metric()`. In this example, the parameters are:
# - `hyperparameter_metric_tag`: The name of the objective metric to report back. The name must be identical to the name specified in the study specification.
# - `metric_value`: The value of the objective metric to report back to the hyperparameter service.
# - `global_step`: The epoch iteration, starting at 0.
# + [markdown] id="tune_custom_job"
# ## Hyperparameter Tune the model
#
# Now start the hyperparameter tuning of your custom model on Vertex. Use this helper function `create_hyperparameter_tuning_job`, which takes the following parameter:
#
# -`hpt_job`: The specification for the hyperparameter tuning job.
#
# The helper function calls job client service's `create_hyperparameter_tuning_job` method, with the following parameters:
#
# -`parent`: The Vertex location path to `Dataset`, `Model` and `Endpoint` resources.
# -`hyperparameter_tuning_job`: The specification for the hyperparameter tuning job.
#
# You will display a handful of the fields returned in `response` object, with the two that are of most interest are:
#
# `response.name`: The Vertex fully qualified identifier assigned to this custom hyperparameter tuning job. You save this identifier for using in subsequent steps.
#
# `response.state`: The current state of the custom hyperparameter tuning job.
# + id="tune_custom_job"
def create_hyperparameter_tuning_job(hpt_job):
response = clients["job"].create_hyperparameter_tuning_job(
parent=PARENT, hyperparameter_tuning_job=hpt_job
)
print("name:", response.name)
print("display_name:", response.display_name)
print("state:", response.state)
print("create_time:", response.create_time)
print("update_time:", response.update_time)
return response
response = create_hyperparameter_tuning_job(hpt_job)
# + [markdown] id="job_id:response"
# Now get the unique identifier for the custom job you created.
# + id="job_id:response"
# The full unique ID for the custom job
job_id = response.name
# The short numeric ID for the custom job
job_short_id = job_id.split("/")[-1]
print(job_id)
# + [markdown] id="get_hpt_job"
# ### Get information on a hyperparameter tuning job
#
# Next, use this helper function `get_hyperparameter_tuning_job`, which takes the following parameter:
#
# - `name`: The Vertex fully qualified identifier for the hyperparameter tuning job.
#
# The helper function calls the job client service's `get_hyperparameter_tuning_job` method, with the following parameter:
#
# - `name`: The Vertex fully qualified identifier for the hyperparameter tuning job.
#
# If you recall, you got the Vertex fully qualified identifier for the hyperparameter tuning job in the `response.name` field when you called the `create_hyperparameter_tuning_job` method, and saved the identifier in the variable `hpt_job_id`.
# + id="get_hpt_job"
def get_hyperparameter_tuning_job(name, silent=False):
response = clients["job"].get_hyperparameter_tuning_job(name=name)
if silent:
return response
print("name:", response.name)
print("display_name:", response.display_name)
print("state:", response.state)
print("create_time:", response.create_time)
print("update_time:", response.update_time)
return response
response = get_hyperparameter_tuning_job(hpt_job_id)
# + [markdown] id="wait_tuning_complete"
# ## Wait for tuning to complete
#
# Hyperparameter tuning the above model may take upwards of 20 minutes time.
#
# Once your model is done tuning, you can calculate the actual time it took to tune the model by subtracting `end_time` from `start_time`.
#
# For your model, we will need to know the location of the saved models for each trial, which the Python script saved in your local Cloud Storage bucket at `MODEL_DIR + '/<trial_number>/saved_model.pb'`.
# + id="wait_tuning_complete"
while True:
job_response = get_hyperparameter_tuning_job(hpt_job_id, True)
if job_response.state != aip.JobState.JOB_STATE_SUCCEEDED:
print("Study trials have not completed:", job_response.state)
if job_response.state == aip.JobState.JOB_STATE_FAILED:
break
else:
if not DIRECT:
MODEL_DIR = MODEL_DIR + "/model"
print("Study trials have completed")
break
time.sleep(60)
# + [markdown] id="review_study_results"
# ### Review the results of the study
#
# Now review the results of trials.
# + id="review_study_results"
best = (None, None, None, 0.0)
for trial in job_response.trials:
print(trial)
# Keep track of the best outcome
if float(trial.final_measurement.metrics[0].value) > best[3]:
try:
best = (
trial.id,
float(trial.parameters[0].value),
float(trial.parameters[1].value),
float(trial.final_measurement.metrics[0].value),
)
except:
best = (
trial.id,
float(trial.parameters[0].value),
None,
float(trial.final_measurement.metrics[0].value),
)
# + [markdown] id="best_trial"
# ### Best trial
#
# Now look at which trial was the best:
# + id="best_trial"
print("ID", best[0])
print("Learning Rate", best[1])
print("Decay", best[2])
print("Validation Accuracy", best[3])
# + [markdown] id="get_best_model"
# ## Get the Best Model
#
# If you used the method of having the service tell the tuning script where to save the model artifacts (`DIRECT = False`), then the model artifacts for the best model are saved at:
#
# MODEL_DIR/<best_trial_id>/model
# + id="get_best_model"
BEST_MODEL_DIR = MODEL_DIR + "/" + best[0] + "/model"
# + [markdown] id="load_saved_model"
# ## Load the saved model
#
# Your model is stored in a TensorFlow SavedModel format in a Cloud Storage bucket. Now load it from the Cloud Storage bucket, and then you can do some things, like evaluate the model, and do a prediction.
#
# To load, you use the TF.Keras `model.load_model()` method passing it the Cloud Storage path where the model is saved -- specified by `MODEL_DIR`.
# + id="load_saved_model"
import tensorflow as tf
model = tf.keras.models.load_model(MODEL_DIR)
# + [markdown] id="evaluate_custom_model:image"
# ## Evaluate the model
#
# Now find out how good the model is.
#
# ### Load evaluation data
#
# You will load the CIFAR10 test (holdout) data from `tf.keras.datasets`, using the method `load_data()`. This will return the dataset as a tuple of two elements. The first element is the training data and the second is the test data. Each element is also a tuple of two elements: the image data, and the corresponding labels.
#
# You don't need the training data, and hence why we loaded it as `(_, _)`.
#
# Before you can run the data through evaluation, you need to preprocess it:
#
# x_test:
# 1. Normalize (rescaling) the pixel data by dividing each pixel by 255. This will replace each single byte integer pixel with a 32-bit floating point number between 0 and 1.
#
# y_test:<br/>
# 2. The labels are currently scalar (sparse). If you look back at the `compile()` step in the `trainer/task.py` script, you will find that it was compiled for sparse labels. So we don't need to do anything more.
# + id="evaluate_custom_model:image,cifar10"
import numpy as np
from tensorflow.keras.datasets import cifar10
(_, _), (x_test, y_test) = cifar10.load_data()
x_test = (x_test / 255.0).astype(np.float32)
print(x_test.shape, y_test.shape)
# + [markdown] id="perform_evaluation_custom"
# ### Perform the model evaluation
#
# Now evaluate how well the model in the custom job did.
# + id="perform_evaluation_custom"
model.evaluate(x_test, y_test)
# + [markdown] id="cleanup"
# # Cleaning up
#
# To clean up all GCP resources used in this project, you can [delete the GCP
# project](https://cloud.google.com/resource-manager/docs/creating-managing-projects#shutting_down_projects) you used for the tutorial.
#
# Otherwise, you can delete the individual resources you created in this tutorial:
#
# - Dataset
# - Pipeline
# - Model
# - Endpoint
# - Batch Job
# - Custom Job
# - Hyperparameter Tuning Job
# - Cloud Storage Bucket
# + id="cleanup"
delete_dataset = True
delete_pipeline = True
delete_model = True
delete_endpoint = True
delete_batchjob = True
delete_customjob = True
delete_hptjob = True
delete_bucket = True
# Delete the dataset using the Vertex fully qualified identifier for the dataset
try:
if delete_dataset and "dataset_id" in globals():
clients["dataset"].delete_dataset(name=dataset_id)
except Exception as e:
print(e)
# Delete the training pipeline using the Vertex fully qualified identifier for the pipeline
try:
if delete_pipeline and "pipeline_id" in globals():
clients["pipeline"].delete_training_pipeline(name=pipeline_id)
except Exception as e:
print(e)
# Delete the model using the Vertex fully qualified identifier for the model
try:
if delete_model and "model_to_deploy_id" in globals():
clients["model"].delete_model(name=model_to_deploy_id)
except Exception as e:
print(e)
# Delete the endpoint using the Vertex fully qualified identifier for the endpoint
try:
if delete_endpoint and "endpoint_id" in globals():
clients["endpoint"].delete_endpoint(name=endpoint_id)
except Exception as e:
print(e)
# Delete the batch job using the Vertex fully qualified identifier for the batch job
try:
if delete_batchjob and "batch_job_id" in globals():
clients["job"].delete_batch_prediction_job(name=batch_job_id)
except Exception as e:
print(e)
# Delete the custom job using the Vertex fully qualified identifier for the custom job
try:
if delete_customjob and "job_id" in globals():
clients["job"].delete_custom_job(name=job_id)
except Exception as e:
print(e)
# Delete the hyperparameter tuning job using the Vertex fully qualified identifier for the hyperparameter tuning job
try:
if delete_hptjob and "hpt_job_id" in globals():
clients["job"].delete_hyperparameter_tuning_job(name=hpt_job_id)
except Exception as e:
print(e)
if delete_bucket and "BUCKET_NAME" in globals():
# ! gsutil rm -r $BUCKET_NAME
| notebooks/community/gapic/custom/showcase_hyperparmeter_tuning_image_classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Debugging
#
# Debugging notebooks can be done in several ways:
#
# * ipdb (or pdb but ipdb is more jupyter friendly)
# * pixie debugger - a GUI debugger that runs inside the notebook
# +
import sys
def printit(x):
print(x)
# -
import ipdb
for i in range(5):
ipdb.set_trace()
printit(i)
import pixiedust
# + pixiedust={"displayParams": {}}
# %%pixie_debugger
for i in range(12):
printit(i)
# -
| python/Debugging.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import glob
from location_test_gui_functions import TestLocationsGUI
import cv2
# +
import cv2
import numpy as np
import pandas as pd
import random
import matplotlib.cm as cm
def rescale_image(image, max_size):
""" Rescale image so that longest dimension equals max size.
Args:
image: 3d numpy array
max_size: number longest output size of image
"""
im_size = np.array(image.shape)
longest_side = np.max(im_size)
scale = max_size / longest_side
new_im_shape = (im_size * scale).astype(int)
new_image = cv2.resize(image,
(new_im_shape[1], new_im_shape[0]),
interpolation=cv2.INTER_NEAREST
)
return new_image, scale
class TestLocationsGUI():
#constructor
def __init__(self, image_files, csv_files, max_size):
"""
Args:
image_files: list of bat images
csv_files: list of csvs with frame's bat info
max_size: largest dimension in pixels of output display
"""
assert len(image_files) == len(csv_files)
self.total_images = len(image_files)
self.test_num = 0
self.focal_tip = 1
self.image_files = image_files
self.csv_files = csv_files
self.max_size = max_size
self.show_points = True
self.load_new_test()
def load_new_test(self):
raw_image = cv2.imread(self.image_files[self.test_num])
self.raw_image, self.image_rescale = rescale_image(raw_image, self.max_size)
self.info = pd.read_csv(self.csv_files[self.test_num])
def refresh_windows(self):
self.image = np.copy(self.raw_image)
def save_validation(self):
""" Overwrite test's wing positions file with current values."""
self.info.to_csv(self.csv_files[self.test_num], index=False)
def change_frame(self, amount):
"""Change test frame forward or backward by amount.
0 is minimum frame ind and number of frames is max (no periodic boundaries)
Args:
amount: number of frames to move positive or negative"""
self.save_validation()
self.test_num += amount
self.test_num = np.max([0, self.test_num])
self.test_num = np.min([self.total_images-1, self.test_num])
self.load_new_test()
def change_focal_wingtip(self):
"""Change focal wingtip to the other."""
self.save_validation()
if self.focal_tip == 1:
self.focal_tip = 2
elif self.focal_tip == 2:
self.focal_tip = 1
def draw_wingtip_positions(self):
""" Draw the location of all wingtip points."""
def _draw_wingtip(wingtip, color, radius, thickness, image_rescale):
if not np.any(np.isnan(wingtip)):
if not np.any(wingtip==-1):
cv2.circle(self.image,
(int(wingtip[0]*image_rescale),
int(wingtip[1]*image_rescale)),
radius=radius,
color=color, thickness=thickness)
point_thickness = -1
circle_thickness = 1
point_radius = int(.006*self.max_size)
circle_radius = int(.018*self.max_size)
color = [255, 255, 255]
if self.info.loc[0, 'hard']:
color = [0, 0, 255]
if self.show_points:
wingtip1 = np.array(
[self.info.loc[0, 'wingtip1_x'], self.info.loc[0, 'wingtip1_y']]
)
_draw_wingtip(wingtip1, color, point_radius, point_thickness, self.image_rescale)
if self.focal_tip == 1:
_draw_wingtip(wingtip1, color, circle_radius, circle_thickness, self.image_rescale)
wingtip2 = np.array(
[self.info.loc[0, 'wingtip2_x'], self.info.loc[0, 'wingtip2_y']]
)
_draw_wingtip(wingtip2, color, point_radius, point_thickness, self.image_rescale)
if self.focal_tip == 2:
_draw_wingtip(wingtip2, color, circle_radius, circle_thickness, self.image_rescale)
def add_frame_info(self):
font = cv2.FONT_HERSHEY_SIMPLEX
bottomLeftCornerOfText = (20, 40)
fontScale = 1
fontColor = (255,255,255)
lineType = 2
cv2.putText(self.image,
f'{self.test_num} / {self.total_images}',
bottomLeftCornerOfText,
font,
fontScale,
fontColor,
lineType
)
def change_wingtip_locations(self, x, y):
"""Record x and y position of click in bat image assosiated with wing focal ind.
Args:
x, y: from mouse click
"""
corrected_x = x / self.image_rescale
corrected_y = y / self.image_rescale
if self.focal_tip == 1 or self.focal_tip == 2:
self.info.loc[0, f'wingtip{self.focal_tip}_x'] = corrected_x
self.info.loc[0, f'wingtip{self.focal_tip}_y'] = corrected_y
self.change_focal_wingtip()
def show_windows(self):
cv2.imshow('bat-image', self.image)
def not_present(self):
# Can't see wings to mark
if self.focal_tip == 1 or self.focal_tip == 2:
self.info.loc[0, f'wingtip{self.focal_tip}_x'] = -2
self.info.loc[0, f'wingtip{self.focal_tip}_y'] = -2
self.change_frame(1)
def mark_hard(self):
# Can't see wings to mark
if self.info.loc[0, 'hard'] != True:
self.info.loc[0, 'hard'] = True
else:
self.info.loc[0, 'hard'] = False
def react_to_keypress(self, key):
""" Process key press
Args:
key: return from cv2 cv2.waitkey
"""
if key == ord('l'):
self.change_frame(1)
elif key == ord('j'):
self.change_frame(-1)
elif key == ord('i'):
self.change_focal_wingtip()
elif key == ord('k'):
self.mark_hard()
elif key == ord('p'):
self.toggle_show_points()
elif key == ord('n'):
self.not_present()
def clicked(self, event, x, y, flags, param):
if event == cv2.EVENT_LBUTTONDOWN:
self.change_wingtip_locations(x, y)
def toggle_show_points(self):
self.show_points = not self.show_points
# -
annotater_name = 'koger'
root_folder = '.../bats-data/wing-validation'
images_folder = os.path.join(root_folder, 'validation-images')
csvs_folder = os.path.join(root_folder, 'validation-csvs')
image_files = sorted(glob.glob(os.path.join(images_folder, '*.png')))
csv_files = sorted(glob.glob(os.path.join(csvs_folder,'*.csv')))
assert len(image_files) == len(csv_files), f"{len(image_files)} {len(csv_files)} every image should have coresponding .csv file"
# +
max_size = 600
gui = TestLocationsGUI(image_files, csv_files, max_size)
# +
cv2.namedWindow('bat-image')
cv2.setMouseCallback('bat-image', gui.clicked)
while True:
gui.refresh_windows()
gui.draw_wingtip_positions()
gui.add_frame_info()
gui.show_windows()
key = cv2.waitKey(500) & 255
gui.react_to_keypress(key)
if key == ord('q'):
gui.save_validation()
print('quitting')
break
cv2.destroyAllWindows()
cv2.waitKey(1)
# -
x = pd.read_csv(csv_files[0])
x.loc[0, 'hard']
| location_test_gui.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Conditional Statements:
# Let's recall the example we have done in class last lecture:
name = input("What's yor name? ")
surname = input("What's your surname? ")
print("That is", name,surname)
# In programming term, that is a sequential structure, that is we have a set of statements that are executed in the given order. However, there are situations were this structure is not feasible, such as when we have to deal with a set of conditional statements. Hence, we have to resort to more complex structures, like the decision-making structure. <br>
# In Python, the if statement is used to implement a decision-making structure. The first line of this statement is called _if clause_: it starts with the _if keyword_ followed by a condition and ends with a colon. And then we have the body which contains a set of statements taht are executed only if the condition is true.
# **Remark**. If the stamente is false, then the body is not executed, and hence nothing happen
name = input('Enter a name of a person you know: ')
if name == "Anna":
print('I know {}!'.format(name))
# We can add an extra level in our conditional statement by using the _else clause_. The **if-else** statement enables to print a double alternative decision-making structure, which implies two possible execution paths (branches) depending on whether the condition is true or false
name = input('Enter a name of a person you know: ')
if name == "Anna":
print('I know {}!'.format(name))
else:
print("I don't know that person")
# We can also nest different conditional statements, as follows:
name = input('Enter a name of a person you know: ')
if name == "Anna":
print('I know {}!'.format(name))
else:
if name == "Mark":
print("I know {}! That is my brother!".format(name))
else:
print("I don't know that person")
# ### Logical Operators (AND - OR)
# Suppose we are working in the marketing department of a national TV broadcaster, and we are interesting in making a marketing campaing that investigate our clients' Sunday habits! We would like to understand whether each of them is at home on Sunday, and if yes, understand the number of hours she watch the TV to propose a new sport offer.<br>
# To do so, we use extend the decision-making structure by using logical operators.
# +
hobby = input("Do you like sports? (y/n) ").lower()
sunday = input("Are you typically at home on sunday? (y/n)").lower()
hours = int(input("How many hours do you watch TV? "))
if ((hobby in ['yes','y'])&(sunday in ['yes','y'])&(hours>6))|\
((hobby in ['yes','y'])&(sunday in ['no','n'])&(hours>14)):
print("You will get an offer from us!")
else:
print("That client is not profittable for us")
| Miscellaneous/ConditionalStatements.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # 3. The Flow of Association and Causation in Graphs
# ## 3.1 Graph Terminology
#
# ## 3.2 Bayesian Networks
#
# ```{admonition} Assumption 3.1 (Local Markov Assumption)
# :class: note
# Given its parents in the DAG, a node $X$ is independent of all its non-descendants.
# ```
#
# ```{admonition} Definition 3.1 (Bayesian Network Factorization)
# :class: warning
# Given a probability distribution $P$ and a DAG $G$, $P$ factorizes according to $P$ if
#
# $$P(x_1,\dots,x_n)=\prod_iP(x_i|pa_i)$$
# ```
#
# ```{admonition} Assumption 3.2 (Minimality Assumption)
# :class: note
# - Given its parents in the DAG, a node $X$ is independent of all its non-descendants (Assumption 3.1).
# - Adjacent nodes in the DAG are dependent.
# ```
#
# 相比于假设3.1,3.2告诉我们不能有额外的独立性存在,例如图3.8,通过假设3.1,可以将$P(x,y)$分解为$P(x)P(y|x)$或者$P(x)P(y)$,但在3.2的假设下,$P(x)P(y)$是不成立的
#
# <img src="images/ch3/3.8.png" width=300px />
# ## 3.3 Causal Graphs
#
# ```{admonition} Definition 3.2 (What is a cause?)
# :class: warning
# A variable $X$ is said to be a cause of a variable $Y$ if $Y$ can change in response to changes in $X$.
# ```
#
# ```{admonition} Assumption 3.3 ((Strict) Causal Edges Assumption)
# :class: note
# In a directed graph, every parent is a direct cause of all its children.
# ```
# ## 3.4 Two-Node Graphs and Graphical Building Blocks
#
# 本节我们将通过最基础的组成模块来理解因果性和相关性在DAG中的是如何传递的。
#
# 最基础的组成模块包括:
#
# 1. chain
# 2. fork
# 3. immorality(collider)
# 4. 两个不相连的节点
# 5. 两个相连的节点
#
# 对于每一种组成模块,我们会给出两个节点(条件)独立或不独立的intuition以及证明。
#
# <img src="images/ch3/3.9.png" width=800px />
#
# 如图3.10,两个不相连的节点,根据[Definition 3.1]()可以得到对应的贝叶斯网络分解式:
#
# $$P(x_1,x_2)=P(x_1)P(x_2)$$
#
# <img src="images/ch3/3.10.png" width=300px />
#
# 如图3.11,根据[Assumption 3.3](),$x_1$是$x_2$的因。
#
# <img src="images/ch3/3.11.png" width=300px />
# ## 3.5 Chains and Forks
#
# ### 3.5.1 相关性(Association)
#
# 要证明$X_1\amalg X_3|X_2$,只需要说明$P(x_1,x_3|x_2)=P(x_1|x_2)P(x_3|x_2)$。
#
# #### Chains
#
# <img src="images/ch3/3.12.png" width=300px />
#
# 对于chains,可以做如下因式分解:
#
# $$P(x_1,x_2,x_3)=P(x_1)P(x_2|x_1)P(x_3|x_2)$$
#
# 通过贝叶斯定理得到
#
# $$\begin{split}P(x_1,x_3|x_2)&=\frac{P(x_1,x_2,x_3)}{P(x_2)}\\&=\frac{P(x_1)P(x_2|x_1)P(x_3|x_2)}{P(x_2)}\\&=\frac{P(x_1,x_2)}{P(x_2)}P(x_3|x_2)\\&=P(x_1|x_2)P(x_3|x_2)\end{split}$$
#
# #### Forks
#
# <img src="images/ch3/3.13.png" width=300px />
#
# $$P(x_1,x_2,x_3)=P(x_2)P(x_1|x_2)P(x_3|x_2)$$
#
# 通过贝叶斯定理得到
#
# $$\begin{split}P(x_1,x_3|x_2)&=\frac{P(x_1,x_2,x_3)}{P(x_2)}\\&=\frac{P(x_2)P(x_1|x_2)P(x_3|x_2)}{P(x_2)}\\&=\frac{P(x_1,x_2)}{P(x_2)}P(x_3|x_2)\\&=P(x_1|x_2)P(x_3|x_2)\end{split}$$
#
# ### 3.5.2 因果性(Causation)
#
# - 相关性是对称双向的,而因果性只在单个方向上传递
# - 因果性只在有向路径上传递
# - 相关性在不包含immorality的任意路径上传递
# ## 3.6 Colliders and their Descendants
#
# ### 3.6.1 immorality(colliders)
#
# <img src="images/ch3/3.16.png" width=300px />
#
# 不同于chains和forks,在immorality(colliders)中$X_1\amalg X_3$,通过贝斯网络因式分解和边缘化$x_2$可以说明
#
# $$\begin{split}P(x_1,x_3)&=\sum_{x_2}P(x_1,x_2,x_3)\\&=\sum_{x_2}P(x_1)P(x_3)P(x_2|x_1,x_3)\\&=P(x_1)P(x_3)\sum_{x_2}P(x_2|x_1,x_3)\\&=P(x_1)P(x_3)\end{split}$$
#
# ### 3.6.2 Good-Looking Men are Jerks
#
# <img src="images/ch3/3.17.png" width=300px />
#
# #### Berkson's paradox
#
# <img src="images/ch3/berkson's_paradox.png" width=800px />
#
# ### 3.6.3 Numerical Example
#
# 考虑如下DGP(data generating process)数据生成过程,$X_1$和$X_3$独立地服从标准正态分布,并用来计算$X_2$:
#
# $$\begin{split}X_1 \sim N(0,1),\ \ X_3\sim N(0,1)\\X_2=X_1+X_3\end{split}$$
#
# 我们已经声明了$X_1$和$X_3$是独立的,但是为了能够方便对比,计算其协方差:
#
# $$\begin{split}Cov(X_1,X_3)&=\mathbb{E}[(X_1-\mathbb{E}[X_1])(X_3-\mathbb{E}[X_3])]\\&=\mathbb{E}[X_1X_3]-\mathbb{E}[X_1]\mathbb{E}[X_3]\\&=\mathbb{E}[X_1,X_3]\ \ \ \ \ \ \ \ \ \ \ \ \ \ \ \ \ \ \ \ \ \ \ \ \ \ \ \ \ \ \ \ \ \ \ \ \ (zero \ mean)\\&=\mathbb{E}[X_1]\mathbb{E}[X_3]\ \ \ \ \ \ \ \ \ \ \ \ \ \ \ \ \ \ \ \ \ \ \ \ \ \ \ \ \ \ \ (independent)\\&=0\end{split}$$
#
# condition on $X_2$时,协方差是非零值,说明$X_1$和$X_3$是相关的:
#
# $$\begin{split}Cov(X_1,X_3|X_2=x)&=\mathbb{E}[X_1X_3|X_2=x]\\&=\mathbb{E}[X_1(x-X_1)]\\&=x\mathbb{E}[X_1]-\mathbb{E}[X_1^2]\\&=-1\end{split}$$
#
# ### 3.6.4 Descendants of Colliders
#
# 因为从collider到它的孩子存在一条有向因果路径,他们是相关的,因此可以把collider的孩子看作其代理人,因此condition on collider的孩子跟它自己是相似的。
#
# ## 3.7 d-separation
#
# ```{admonition} Definition 3.3 (blocked path)
# :class: warning
# A path between nodes $X$ and $Y$ is blocked by a (potentially empty) conditioning set $Z$ if either of the following is true:
#
# 1. Along the path, there is a chain $\cdots \rightarrow W \rightarrow \cdots$ or a fork $\cdots \leftarrow W \rightarrow \cdots$, where $W$ is conditioned on ($W\in Z$).
# 2. There is a collider $W$ on the path that is not conditioned on ($W\notin Z$) and none of its descendants are conditioned on ($de(W)\nsubseteq Z$).
# ```
# ```{admonition} Definition 3.4 (d-separation)
# :class: warning
# Two (sets of) nodes $X$ and $Y$ are d-separated by a set of nodes $Z$ if all of the paths between (any node in) $X$ and (any node in) $Y$ are blocked by $Z$.
# ```
#
# ```{admonition} Theorem 3.1 (global Markov assumption)
# :class: important
# Given that $P$ is *Markov* with respect to $G$ (satisfies the local Markov assumption, Assumption 3.1), if $X$ and $Y$ are d-separated in $G$conditioned on $Z$, then $X$ and $Y$ are independent in $P$ conditioned on $Z$. We can write this succinctly as follows:
#
# $$X\amalg_GY|Z \Longrightarrow X\amalg_PY|Z$$
# ```
#
# 这个定理非常重要,公式称为 **global Markov assumption**
#
# local Markov assumption,Bayesian network factorization 和 global Markov assumption 是等价的(all equivalent)。
# ## 练习
#
# <img src="images/ch3/3.19a.png" width=300px />
#
# [Fgiure 3.19a exercise](https://www.notion.so/daabbc9b8d4943df92eebd6e286d3f24)
#
# <img src="images/ch3/3.19b.png" width=300px />
#
# [Fgiure 3.19b exercise](https://www.notion.so/7dca44fb15954ae18c348fa125049e5c)
# ## 3.8 Flow of Association and Causation
#
# 回顾Section 1.3.2,不仅仅是相关性不是因果性,其实因果性是相关性的子类,这就是为什么相关性和因果性都在有向路径中传递。
#
# <img src="images/ch3/3.20.png" width=300px />
#
# 我们将有向路径中传递的相关性看做因果相关性,一种常见的非因果相关性叫做混淆相关性(confounding association),如图3.20。
#
# 常规的贝叶斯网络是单纯的统计模型,因此我们只能在贝叶斯网络中讨论相关性的传递。相关性在因果图中也有着完全相同的方式。通过基础的模块(chain、fork、collider)我们能知道相关性在普遍DAG图中是如何传递的,也能通过两个节点是否是d-sparated来判断是否相关。
#
# Assumption 3.3赋予了有向路径传递因果性的独有角色,同时表明因果性是不对称的,相关性是对称的。
#
# ### 3.8.1 d-separation说明相关性是因果性
#
# 在把$X$的所有出度(从$X$指向外部的边)删除的途中,如果$X$和$Y$是d-separated,那么他们之间只存在单纯的非因果相关性,因为因果性只存在$X$指向$Y$的边中,已经被删掉了。
#
# <img src="images/ch3/3.21.png" width=900px />
#
# 图3.21通过解释相关性的传递说明了每个假设(assumption)的意义:
#
# 1. [Markov Assumption(Assumption 3.1)]()告诉我们哪些节点是不相关的;
# 2. [Minimality Assumption(Assumption 3.2)]()又告诉我们哪些路径上存在相关性;
# 3. [Causal Edges Assumption(Assumption 3.3)]()让我们知道因果性只在有向路径上传递
#
# 最重要是1和3,因为Minimality Assumption的第一部分就是Markov Assumption,第二部分也被囊括在Causal Edges Assumption中了。
| docs/causal_inference/introduction_to_causal_inference/ch3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# TODO come up wiht good way to compare models!!!!
# TODO Get rid of scaling.
# TODO talk to marco about consumption.
# TODO make average model the baseline.
# TODO make scoring the parent class. Give only fitting and _func as input.
# TODO, build some resistance against unseen actions into the thermal model
import sys
sys.path.append("..")
sys.path.append("../MPC")
sys.path.append('../MPC/ThermalModels')
from ThermalDataManager import ThermalDataManager
from ThermalModel import *
import pandas as pd
import numpy as np
import pickle
import datetime
import yaml
import pytz
import pprint
import datetime
import utils
import matplotlib.pyplot as plt
from xbos import get_client
from sklearn.model_selection import cross_val_score
from sklearn.utils import shuffle
# -
from ThermalModel import ThermalModel
from AverageThermalModel import AverageThermalModel
from AverageThermalModel import AverageMPCThermalModel as AverageMPCThermalModel
from BaselineThermalModel import BaselineThermalModel
# from ActionThermalModel import ThermalModel
# +
# TODO instead of hardcoding input. just give it a dictionary it should append.
def save_thermal_model_data(thermalModel, corresponding_days, params, file_name):
"""saves the whole model to a yaml file.
RECOMMENDED: PYAML should be installed for prettier config file.
param: thermalModel : object, thermal model which stores the types of scoring.
param: corresponding_days: list, the order of data according to the number of days from current day.
param: params, the past parameters"""
config_dict = {}
config_dict["Param Order"] = thermalModel._params_order
config_dict["Params"] = params
# store evaluations and RMSE's.
config_dict["Evaluations"] = {}
config_dict["Evaluations"]["Corresponding_Days"] = corresponding_days
config_dict["Evaluations"]["Baseline"] = thermalModel.baseline_error
config_dict["Evaluations"]["Model"] = thermalModel.model_error
config_dict["Evaluations"]["ActionOrder"] = thermalModel.scoreTypeList
config_dict["Evaluations"]["Better Than Baseline"] = thermalModel.betterThanBaseline
with open("./thermal_model_" + file_name, 'wb') as ymlfile:
# TODO Note import pyaml here to get a pretty config file.
try:
import pyaml
pyaml.dump(config_dict, ymlfile)
except ImportError:
yaml.dump(config_dict, ymlfile)
days = sorted(4 * list(range(1, 101, 5)))
# TODO change file name
# save_thermal_model_data(thermalModel, days, params, "Animal_shelter" + "_average_adding.yml")
# +
# CONSISTANCY CHECKS
def apply_consistency_check_to_model(data=None, thermal_model=None, thermal_model_class=None):
"""
:param data: Only used for fitting the thermal model."""
assert thermal_model is not None or (thermal_model_class is not None and data is not None)
# evaluate actions. On same temperatures, heating should increase, cooling decrease, and no action should be no different
if thermal_model is None:
thermalModel = thermal_model_class()
thermalModel.fit(data, data["t_next"])
else:
thermalModel = thermal_model
def prepare_consistency_test_data(thermal_model, start_temperature=50, end_temperature=100, increments=5, dt=5):
filter_columns = thermal_model._filter_columns
data = []
for temperature in range(start_temperature, end_temperature+increments, increments):
# TODO potentially not hardcode dt
for action in range(0, 6):
datapoint = {"dt": dt, "action": action,
"t_out": temperature, "t_in": temperature}
for col in filter_columns:
if "zone_temperature_" in col:
datapoint[col] = temperature
data.append(datapoint)
return pd.DataFrame(data)
consistancy_test_data = prepare_consistency_test_data(thermalModel)
consistancy_test_data["prediction"] = thermalModel.predict(consistancy_test_data, should_round=False)
def consistency_check(df):
"""Consistency check for a df with 3 entries. The datapoints can only differ in the action to be meaningful.
:param df: pd.df columns as given by ThermalDataManager plus a column with the predctions"""
t_in = df['t_in'].values[0]
dt = df["dt"].values[0]
no_action_temperature = df[(df['action'] == utils.NO_ACTION)]["prediction"].values
heating_temperature = df[df['action'] == utils.HEATING_ACTION]["prediction"].values
cooling_temperature = df[df['action'] == utils.COOLING_ACTION]["prediction"].values
two_stage_heating_temperature = df[df['action'] == utils.TWO_STAGE_HEATING_ACTION]["prediction"].values
two_stage_cooling_temperature = df[df['action'] == utils.TWO_STAGE_COOLING_ACTION]["prediction"].values
consistency_flag = True
# TODO only use this check when t_out and zone temperature are the same as t_in
# Following checks with t_in are only possible when everything has the same temperature
# check if predicted heating temperature is higher than current
if heating_temperature <= t_in:
consistency_flag = False
print("Warning, heating_temperature is lower than t_in.")
if cooling_temperature >= t_in:
consistency_flag = False
print("Warning, cooling_temperature is higher than t_in.")
# check that heating is more than no action and cooling
if heating_temperature <= no_action_temperature or heating_temperature <= cooling_temperature:
consistency_flag = False
print("Warning, heating_temperature is too low compared to other actions.")
# check that two stage heating is more than no action and cooling
if heating_temperature <= no_action_temperature or heating_temperature <= cooling_temperature:
consistency_flag = False
print("Warning, heating_temperature is too low compared to other actions.")
# check cooling is lower than heating and no action
if cooling_temperature >= no_action_temperature or cooling_temperature >= heating_temperature:
consistency_flag = False
print("Warning, cooling_temperature is too high compared to other actions.")
# check if no action is between cooling and heating
if not cooling_temperature < no_action_temperature < heating_temperature:
consistency_flag = False
print("Warning, no action is not inbetween heating temperature and cooling temperature.")
# want to know for what data it didn't work
if not consistency_flag:
print("Inconsistency for following data:")
print(df)
print("")
return consistency_flag
consistentcy_results = consistancy_test_data.groupby(["t_in", "dt"]).apply(lambda df: consistency_check(df))
is_zone_consistent = all(consistentcy_results.values)
if is_zone_consistent:
print("The thermal model is consistent.")
else:
print("The thermal model is inconsistent.")
def check_consistency_building(thermal_data, thermal_model=None, thermal_model_class=None):
assert not (thermal_model is None and thermal_model_class is None)
for zone, data in thermal_data.items():
print("--------- Consistency check for zone %s ---------" % zone)
# TODO fix the consistency check, can't give thermal model to whole building.
# apply_consistency_check_to_data(data, thermal_model=thermal_model, thermal_model_class=thermal_model_class)
# +
# Going through all config files
import os
# Go to folder where all config files are located
config_folder_location = "../Buildings/"
all_dir = os.walk(config_folder_location).next()
for d in all_dir[1]:
print("============================================================================")
print("============================================================================")
end_dir = config_folder_location + d + "/"
files = os.walk(end_dir).next()[2]
print("in dir: ", end_dir)
for f in files:
if ".yml" not in f:
print("%s is not a yaml file. Continue to next file." % f)
continue
print("Getting file %s" % f)
# Loads the configs
with open("./" + end_dir + "/" + f, 'r') as o:
config = yaml.load(o)
# Do whatever check on config
building = config["Building"]
zone_thermal_data = utils.get_data(building=building, days_back=100, force_reload=True)
# print(zone_thermal_data.keys())
# zone = "HVAC_Zone_Eastzone"
# print(sum(zone_thermal_data[zone]["action"] == 5))
# filtered_thermal_data = {}
# for zone, data in zone_thermal_data.items():
# filtered_thermal_data[zone] = data[data["dt"] != 1]
# buidling_thermal_data = concat_zone_data(filtered_thermal_data)
# print("================== Scores for Building: %s ==================" % building)
# get_scores_for_days(thermal_data=thermal_data)
# print("================== Evaluate data for Buidling: %s ==================" % building)
# for zone, zone_data in thermal_data.items():
# print("---------- Evaluating data for zone %s ----------" % zone)
# evaluate_zone_data(zone_data)
print("================== Consistency check for Building: %s ==================" % building)
for zone, data in zone_thermal_data.items():
th = ThermalModel().fit(data, data["t_next"])
# apply_consistency_check_to_model(thermal_model=th)
# print("================== Get averages for Building: %s ==================" % building)
# evaluate_action_impact(thermal_data=thermal_data)
# +
# TODO give each function a thermal model param so we can input the class/module.
def get_zone_scores(zone_data, test_on_training=True, thermal_model=None, thermal_model_class=None):
assert not (thermal_model is None and thermal_model_class is None)
if thermal_model is None:
thermalModel = thermal_model_class()
if not test_on_training:
training_data, test_data = get_training_test_data(zone_data)
else:
training_data = test_data = zone_data
NUM_DATA = zone_data.shape[0]
thermalModel.fit(training_data, training_data['t_next'])
else:
thermalModel = thermal_model
test_data = zone_data
for score_action in range(-1, 3):
thermalModel.score(test_data, test_data['t_next'], scoreType=score_action)
# ==== end ====
pprint.pprint("ScoreType List: ")
pprint.pprint(thermalModel.scoreTypeList)
print("")
pprint.pprint("Better than baseline: ")
pprint.pprint(thermalModel.betterThanBaseline)
print("")
df_model_error = pd.DataFrame(thermalModel.model_error, index=thermalModel.scoreTypeList)
df_baseline_error = pd.DataFrame(thermalModel.baseline_error, index=thermalModel.scoreTypeList)
df_model_baseline_error = pd.concat([df_model_error, df_baseline_error], axis=0, keys=["Model","Baseline"])
print("Model and Baseline Error:")
print(df_model_baseline_error)
print("")
# save_thermal_model_data(thermalModel, [45], thermalModel._params, "avenal-animal-shelter" + "_cross_eval_" + "_average_all.yml")
# print(thermalModel.past_coeff)
def get_building_scores(thermal_data, test_on_training=False, thermal_model=None, thermal_model_class=None):
for zone, zone_data in thermal_data.items():
print("--------- Scoring Zone: %s ---------" % zone)
get_zone_scores(zone_data, test_on_training=test_on_training, thermal_model_class=thermal_model_class, thermal_model=thermal_model)
# Find best number of days to get for good thermalModel performance
# TODO Right now only doing it for 50 days really
# +
def apply_temperature_change(df):
no_change_data = df[df["t_next"] == df["t_in"]]
increase_data = df[df["t_next"] > df["t_in"]]
decrease_data = df[df["t_next"] < df["t_in"]]
to_return = {"total": df.shape[0],
"total_no_change": no_change_data.shape[0],
"percent_no_change": 100 * no_change_data.shape[0]/float(df.shape[0]),
"total_increase": increase_data.shape[0],
"percent_increase": 100 * increase_data.shape[0]/float(df.shape[0]),
"total_decrease": decrease_data.shape[0],
"percent_decrease": 100 * decrease_data.shape[0]/float(df.shape[0])}
return pd.Series(to_return)
def no_temperature_change_interval(df):
def no_temp_change_for_action(a_df):
no_change = a_df[a_df["t_min"] == a_df["t_max"]]
return pd.Series({"total": a_df.shape[0], "percent_no_change": 100 * no_change.shape[0]/float(a_df.shape[0]),
"total_no_change": no_change.shape[0]})
return df.groupby("action").apply(no_temp_change_for_action)
def evaluate_zone_data(data, find_single_actions=False):
"""Does basic data evaluation.
:param data: (pd.df) data for one zone. columns: (t_in, t_next, t_out, a1, a2, zone_temperature+zones* )"""
# Filter for action data
no_action_data = data[(data["a1"]==0) & (data["a2"]==0)]
heating_data = data[data["a1"] == 1]
cooling_data = data[data["a2"] == 1]
# ====== Find the mean increase in temperature for each action for the given zone data ======
print("----- Get avearge change in temperature for each action. -----")
def get_delta_mean(action_data):
# get the mean change of temperature from now to next.
return np.mean((action_data["t_next"] - action_data["t_in"])/action_data["dt"])
mean_cooling_delta = get_delta_mean(cooling_data)
mean_heating_delta = get_delta_mean(heating_data)
mean_no_action_delta = get_delta_mean(no_action_data)
mean_all_action_delta = get_delta_mean(data)
print("For cooling there was an average %s degree change." % str(mean_cooling_delta))
print("For heating there was an average %s degree change." % str(mean_heating_delta))
print("For no action there was an average %s degree change." % str(mean_no_action_delta))
print("For all actions there was an average %s degree change." % str(mean_all_action_delta))
# ====== end ======
# ====== Number and percentage of individual action data ========
print("--------- Number and percentage of individual action data ---------")
num_data = data.shape[0]
num_no_action_data = no_action_data.shape[0]
num_heating_data = heating_data.shape[0]
num_cooling_data = cooling_data.shape[0]
print("We have %f total data points." % num_data)
print("We have %f no action data points, which is %f percent of total." % (num_no_action_data, 100 * float(num_no_action_data)/num_data))
print("We have %f heating data points, which is %f percent of total." % (num_heating_data, 100 * float(num_heating_data)/num_data))
print("We have %f cooling data points, which is %f percent of total." % (num_cooling_data, 100 * float(num_cooling_data)/num_data))
# ========= end ==========
# ========= Find temperature change by action ========
print("--------- Find temperature change by action ---------")
no_action_change = apply_temperature_change(no_action_data)
heating_change = apply_temperature_change(heating_data)
cooling_change = apply_temperature_change(cooling_data)
print("No action temperature changes:")
print(no_action_change)
print("")
print("Heating temperature changes:")
print(heating_change)
print("")
print("Cooling temperature changes:")
print(cooling_change)
print("")
# ======== end =======
# Group all data by dt and find evaluate temperature changes.
print("---------- Group all data by dt and find evaluate temperature changes. --------")
def group_dt_and_action(to_group_data):
"""Group the data by the interval lengths and whether temperature increased, dropped or stayed the same."""
return to_group_data.groupby(by=["dt"]).apply(apply_temperature_change)
no_action_dt_change = group_dt_and_action(no_action_data)
heating_dt_change = group_dt_and_action(heating_data)
cooling_dt_chage = group_dt_and_action(cooling_data)
print("No Action, Action Change data:")
print(no_action_dt_change)
print("")
print("Heating, Action Change data:")
print(heating_dt_change)
print("")
print("Cooling, Action Change data:")
print(cooling_dt_chage)
print("")
# ====== end =======
if find_single_actions:
# ====== Find actions which are not integers and stand by themselves
print("--------- Find non interger actions with no action pre/suceeding ---------")
idx_action = []
for i in range(1,len(data.index)-1):
curr = data["action"][i]
prev = data["action"][i-1]
next_a = data["action"][i+1]
if curr not in [0,1,2] and prev == 0 and next_a == 0:
print("++++++++++++")
print("Action is by itself with action %f for data:" % curr)
print(data.iloc[i-2:i+2])
idx_action.append(i)
print("-------------")
print("There were %f lone standing actions out of %f data points." % (len(idx_action), data.shape[0]))
# ====== end =======
# ======= Find intervals where there was no change in temperature throughout the interval =======
print("--------- Find data where temperature did not change in interval ---------")
if "t_min" in data.columns:
print(data.groupby("dt").apply(no_temperature_change_interval))
else:
print("We don't have evaluation type data. i.e. no [t_min, t_max, etc] fields.")
# -
# buildings = ["ciee", "avenal-animal-shelter", "avenal-veterans-hall", "avenal-movie-theatre", "avenal-public-works-yard", "avenal-recreation-center", "orinda-community-center","north-berkeley-senior-center","south-berkeley-senior-center"]
buildings = ["avenal-veterans-hall"]
for bldg in buildings:
print(bldg)
get_raw_data(building=bldg, days_back=10, force_reload=False)
# +
bldg = 'avenal-veterans-hall'
inside_data, outside_data = get_raw_data(building=bldg)
# print(inside_data.keys()[2])
# print(zone_data[zone_data["t_in"].isna()])
zone_data_inside = inside_data.values()[2]
print(zone_data_inside.iloc[4230:4300])
# c = get_client()
# cfg = get_config(bldg)
# dataManager = ThermalDataManager(cfg, c)
# thermal_data = dataManager._preprocess_thermal_data(inside_data, outside_data, evaluate_preprocess=True)
# +
bldg = 'avenal-veterans-hall'
# bldg = "north-berkeley-senior-center"
zone_thermal_data = get_data(building=bldg, days_back=50, evaluate_preprocess=True, force_reload=False)
# for zone, data in zone_thermal_data.items():
# print("------- %s -------" % zone)
# evaluate_zone_data(data)
zone_data = zone_thermal_data.values()[2]
print(zone_thermal_data.keys()[1])
row_filter = ["zone_temperature_" in col for col in zone_data.columns]
for i in range(zone_data.shape[0]):
row = zone_data.iloc[i]
tout = row["t_out"]
tin = row["t_in"]
tnext = row["t_next"]
zone_temperatures = row[row_filter]
others_less_than_curr_zone = all([temp < tin for temp in zone_temperatures])
others_higher_than_curr_zone = all([temp > tin for temp in zone_temperatures])
action = row["action"]
if action == 0:
continue
if others_higher_than_curr_zone and tout > tin and action != 2 and action != 5:
if tin > tnext:
print("All other temperature were higher than curr temperature, but temperature fell:")
print(row)
print("")
elif others_less_than_curr_zone and tout < tin and action != 1 and action != 3:
if tin < tnext:
print("All other temperature were lower than curr temperature, but temperature rose:")
print(row)
print("")
print("/n")
print("/n")
evaluate_zone_data(zone_data)
# +
# For scoring model.
bldg = "ciee"
zone_thermal_data = get_data(building=bldg, days_back=50, evaluate_preprocess=False)
building_thermal_data = concat_zone_data(zone_thermal_data)
# for zone, data in zone_thermal_data.items():
# print("========= Zone % s =========" % zone)
# evaluate_zone_data(data)
def get_models(zone_thermal_data, ThermalModelClass):
"""Trains the given thermalModel to three instances. Once on all data, once on only 15 min data and once
on data larger or equal to 5 min."""
thermal_models = {}
for zone, data in zone_thermal_data.items():
models = {"all": ThermalModelClass().fit(data, data["t_next"]),
"15min": ThermalModelClass().fit(data[data["dt"] == 15], data[data["dt"] == 15]["t_next"]),
"5up": ThermalModelClass().fit(data[data["dt"] >=5], data[data["dt"] >=5]["t_next"])}
thermal_models[zone] = models
return thermal_models
def score_models(zone_thermal_data, thermal_models):
for zone, data in zone_thermal_data.items():
print("====== Zone %s ======" % zone)
print("------ All data ------")
get_zone_scores(zone_data=data, thermal_model=thermal_models[zone]["all"])
print("------ 15 min Data ------")
get_zone_scores(zone_data=data[data["dt"] == 15], thermal_model=thermal_models[zone]["15min"])
print("------ 5 and up Data ------")
get_zone_scores(zone_data=data[data["dt"] >=5], thermal_model=thermal_models[zone]["5up"])
def get_training_test_data(all_data, percent_training=0.8, should_shuffle=False):
"""Get training and test data for each zone in dictionary
:param data: {zone: pd.df} ."""
training_data = {}
test_data = {}
assert percent_training <= 1
for zone, data in all_data.items():
if should_shuffle:
data = shuffle(data)
no_action_data = data[(data["a1"] == 0) & (data["a2"] == 0)]
heating_data = data[data["a1"] == 1]
cooling_data = data[data["a2"] == 1]
training_data[zone] = pd.concat([no_action_data.iloc[:int(no_action_data.shape[0]*percent_training)],
heating_data.iloc[:int(heating_data.shape[0]*percent_training)],
cooling_data.iloc[:int(cooling_data.shape[0]*percent_training)]])
test_data[zone] = pd.concat([no_action_data.iloc[int(no_action_data.shape[0]*percent_training):],
heating_data.iloc[int(heating_data.shape[0]*percent_training):],
cooling_data.iloc[int(cooling_data.shape[0]*percent_training):]])
return training_data, test_data
training_data, test_data = get_training_test_data(zone_thermal_data)
# evaluate_zone_data(test_data["HVAC_Zone_Centralzone"])
print(training_data["HVAC_Zone_Eastzone"])
avg_models = get_models(training_data, AverageThermalModel)
normal_models = get_models(training_data, ThermalModel)
# for zone, model in normal_models.items():
# print("======== %s =======" % zone)
# apply_consistency_check_to_model(thermal_model=model["15min"])
score_models(test_data, avg_models)
print("============================\n============================")
score_models(test_data, normal_models)
# -
def getBestHorizon(data):
"""Finds the best stretch of data, i.e. with most actions which are either heating or cooling. Gives
preference to cooling.
returns: best data slice"""
horizon = datetime.timedelta(hours=3)
start = datetime.datetime(year=2018, month=3, day=9, hour=8, minute=15)
good_dates = []
delta_minutes = datetime.timedelta(minutes=15)
for row in data.itertuples():
start = row[0]
if sum(data.loc[start:start+horizon]["action"]) > 0:
good_dates.append((start, sum(data.loc[start:start+horizon]["action"])))
good_dates.sort(key=lambda x: x[1])
new_start = good_dates[-1][0]
return data.loc[new_start:new_start+horizon]
trivial_predictions = {zone: np.ones(data.shape[0]) * data["t_in"][0] for zone, data in thermal_data.items()} # we predict the first temperature for the whole horizon
# +
def constantZonePredictions(data, thermal_model):
zone_temp_filter = data.columns[["zone_temperature_" in col for col in data.columns]]
constant_zone_temperatures = data[zone_temp_filter].iloc[0] # get the first row for constant zone temperatures.
constant_zone_predictions = [data.iloc[0]["t_in"]]
for row in data.iterrows():
# constant zone temperature predictions
row_data = row[1] # work with the row data
row_data["t_in"] = constant_zone_predictions[-1] # assign last prediction
row_data[constant_zone_temperatures.index] = constant_zone_temperatures.values
constant_zone_predictions.append(thermal_model.predict(row_data))
return constant_zone_predictions
def perfectZonePredictions(data, thermal_model):
predictions = [data.iloc[0]["t_in"]]
for row in data.iterrows():
row_data = row[1] # work with the row data
# assume we know the zone temperatures perfectly
row_data["t_in"] = perfect_zone_predictions[-1] # assign last prediction
perfect_zone_predictions.append(thermal_model.predict(row_data))
return perfect_zone_predictions
# -
southzone_best_data = getBestHorizon(thermal_data["HVAC_Zone_Southzone"]) # use some interval
start = southzone_best_data.index[0]
end = southzone_best_data.index[-1]
bestData = {zone: data[start:end] for zone, data in thermal_data.items()}
rename_col = {zone: "zone_temperature_" + zone for zone in thermal_data.keys()}
for zone in thermal_data.keys():
for zone1 in thermal_data.keys():
ti = "zone_temperature_" + zone1
if ti not in southzone_best_data.columns and zone != zone1:
bestData[zone] = southzone_best_data.rename({"t_in" : ti, "zone_temperature_"+ zone: "t_in"}, axis='columns')
print(bestData)
# +
# bestData = {zone: southzone_best_data for zone, data in thermal_data.items()}
# iteration evaluation
# start by assuming that we only know current zone temperatures. Find predictions for each zone that way
last_prediction_data = {zone: constantZonePredictions(data, thermal_model_zones[zone]) for zone, data in bestData.items()}
print(last_prediction_data)
for _ in range(1):
temp_prediction_data = {}
for zone, data in bestData.items():
for prediction_zone, prediction in last_prediction_data.items():
if prediction_zone != zone:
data.loc[:,"zone_temperature_" + prediction_zone] = prediction[:-1] # TODO NOTE we predict one more than the data we have.
print(data)
temp_prediction_data[zone] = perfectZonePredictions(data, thermal_model_zones[zone])
last_prediction_data = temp_prediction_data
print(last_prediction_data)
# +
d = getBestHorizon(thermal_data["HVAC_Zone_Southzone"])
print(constantZonePredictions(d, thermal_model_zones["HVAC_Zone_Southzone"]))
print(trivial_prediction)
print(constant_zone_predictions)
print(perfect_zone_predictions)
# +
trivial_error = []
constant_error = []
perfect_error = []
dt = data["dt"]
t_in = data["t_in"]
for i in range(1,len(t_in)):
trivial_error.append(thermal_model._normalizedRMSE_STD(trivial_prediction[:i], t_in[:i], dt[:i]))
constant_error.append(thermal_model._normalizedRMSE_STD(constant_zone_predictions[:i], t_in[:i], dt[:i]))
perfect_error.append(thermal_model._normalizedRMSE_STD(perfect_zone_predictions[:i], t_in[:i], dt[:i]))
# +
diff = constant_zone_predictions[:-1] - t_in[:]
print(np.square(diff))
plt.plot(trivial_error)
plt.show()
print(constant_error)
plt.plot(constant_error)
plt.show()
plt.plot(perfect_error)
plt.show()
# -
np.array(constant_error)[:, 1]
# $$T_{in} + dt * ( a_1*c_1 + a_2*c_2 + (T_{out} - T_{in})*c_3 + c_4 + \sum_{i = 0}^N (T_{zone_i} - T_{in}) * c_{5+i})$$
(15*100)/60.
# TODO
# In preprocessing maybe not take averages ouf outdoor and zone temperatures, because even when we predict we only know starting temperature.
# Add bias terms for temperatures. Since we are in fahrenheit we can never account for 0. If we do that though we can't
# have Tin*action because we will never do anything if Tin is zero. Talk to thanos. There are actually multiple issues
# iwth multiplying by Tin, such as lower temperatures should correspond to a higher temperature increase, but the opposite is the case.
# NEED TO ADD COEFICCIENT BECAUSE THEN action(T_in + coeff) where we can then learn the temperature of the heater/cooler!!1
# Totally messed up with 1min data.
#I added the 1 min in preprocessing. So it is actually 0 min data, so no change makes absolute sense.
# Best thing to do is to keep a past pointer to the last row, so t_next and t_in of contigious samples are the same.
| DP/Server/iPythonNotebooks/Evaluations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# ## Data Import
# +
library(dplyr)
library(arules)
bookings <- read.csv("output.csv")
head(bookings)
# +
# Discretize lead_time and adr to (low, medium, high)
bookings <- discretizeDF(bookings, methods = list(
lead_time = list(method = "frequency", breaks = 3,
labels = c("Low", "Medium", "High")),
adr = list(method = "frequency", breaks = 3,
labels = c("Low", "Medium", "High")),
stays_in_weekend_nights = list(method = "frequency", breaks = 3,
labels = c("Low", "Medium", "High")),
stays_in_week_nights = list(method = "frequency", breaks = 3,
labels = c("Low", "Medium", "High")),
adults = list(method = "frequency", breaks = 3,
labels = c("Low", "Medium", "High"))
),
default = list(method = "none")
)
head(bookings)
# -
index <- 1:ncol(bookings)
bookings[ , index] <- lapply(bookings[ , index], as.factor)
str(bookings)
head(bookings)
# ## Transactions
# +
trans <- as(bookings, "transactions")
image(trans)
summary(trans)
str(bookings)
str(trans)
# -
# ## Apriori
# +
rules <- apriori(trans,parameter=list(minlen=2, support=.005, confidence=.9), control = list(verbose=F),appearance = list(rhs=c("adr=Low","adr=Medium","adr=High"),default="lhs"))
rules_lift <- sort(rules, by="lift")
rules_pruned <- rules_lift[!is.redundant(rules_lift, measure="lift")]
length(rules_pruned)
# -
inspect(rules_pruned,by="lift")
# ## Plots
library(arulesViz)
plot(rules_pruned, method="grouped")
plot(rules_pruned, method="graph")
plot(rules_pruned, method="grouped matrix")
plot(rules_pruned, method="matrix3D")
plot(rules_pruned, method="paracoord")
plot(rules_pruned, measure=c("support", "lift"), shading = "confidence")
top10Rules <- head(rules_pruned, n = 10, by = "confidence")
plot(top10Rules, method = "graph", engine = "htmlwidget")
saveAsGraph(head(rules_pruned, n = 100, by = "lift"), file = "rules.graphml")
# ## SubRules
subrules <- rules_pruned[quality(rules_pruned)$lift > 2.95]
subrules
plot(subrules, method="grouped")
inspect(subrules,by="lift")
| lab3/lab3-Chris-R.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Configurations for Colab
# +
import sys
IN_COLAB = "google.colab" in sys.modules
if IN_COLAB:
# !apt install python-opengl
# !apt install ffmpeg
# !apt install xvfb
# !pip install pyvirtualdisplay
# !pip install gym
from pyvirtualdisplay import Display
# Start virtual display
dis = Display(visible=0, size=(400, 400))
dis.start()
# -
# # 03. Prioritized Experience Replay (PER)
#
# [<NAME> al., "Prioritized Experience Replay." arXiv preprint arXiv:1511.05952, 2015.](https://arxiv.org/pdf/1511.05952.pdf)
#
# Using a replay memory leads to design choices at two levels: which experiences to store, and which experiences to replay (and how to do so). This paper addresses only the latter: making the most effective use of the replay memory for learning, assuming that its contents are outside of our control.
#
# The central component of prioritized replay is the criterion by which the importance of each transition is measured. A reasonable approach is to use the magnitude of a transition’s TD error $\delta$, which indicates how ‘surprising’
# or unexpected the transition is. This algorithm stores the last encountered TD error along with each transition in the replay memory. The transition with the largest absolute TD error is replayed from the memory. A Q-learning update
# is applied to this transition, which updates the weights in proportion to the TD error. One thing to note that new transitions arrive without a known TD-error, so it puts them at maximal priority in order to guarantee that all experience is seen at least once. (see *store* method)
#
# We might use 2 ideas to deal with TD-error: 1. greedy TD-error prioritization, 2. stochastic prioritization. However, greedy TD-error prioritization has a severe drawback. Greedy prioritization focuses on a small subset of the experience: errors shrink slowly, especially when using function approximation, meaning that the initially high error transitions get replayed frequently. This lack of diversity that makes the system prone to over-fitting. To overcome this issue, we will use a stochastic sampling method that interpolates between pure greedy prioritization and uniform random sampling.
#
# $$
# P(i) = \frac{p_i^{\alpha}}{\sum_k p_k^{\alpha}}
# $$
#
# where $p_i > 0$ is the priority of transition $i$. The exponent $\alpha$ determines how much prioritization is used, with $\alpha = 0$ corresponding to the uniform case. In practice, we use additional term $\epsilon$ in order to guarantee all transactions can be possibly sampled: $p_i = |\delta_i| + \epsilon$, where $\epsilon$ is a small positive constant.
#
# One more. Let's recall one of the main ideas of DQN. To remove correlation of observations, it uses uniformly random sampling from the replay buffer. Prioritized replay introduces bias because it doesn't sample experiences uniformly at random due to the sampling proportion correspoding to TD-error. We can correct this bias by using importance-sampling (IS) weights
#
# $$
# w_i = \big( \frac{1}{N} \cdot \frac{1}{P(i)} \big)^\beta
# $$
#
# that fully compensates for the non-uniform probabilities $P(i)$ if $\beta = 1$. These weights can be folded into the Q-learning update by using $w_i\delta_i$ instead of $\delta_i$. In typical reinforcement learning scenarios, the unbiased nature of the updates is most important near convergence at the end of training, We therefore exploit the flexibility of annealing the amount of importance-sampling correction over time, by defining a schedule on the exponent $\beta$ that reaches 1 only at the end of learning.
# +
import os
import random
from typing import Dict, List, Tuple
import gym
import matplotlib.pyplot as plt
import numpy as np
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from IPython.display import clear_output
if IN_COLAB and not os.path.exists("segment_tree.py"):
# download segment tree module
# !wget https://raw.githubusercontent.com/curt-park/rainbow-is-all-you-need/master/segment_tree.py
from segment_tree import MinSegmentTree, SumSegmentTree
# -
# ## Replay buffer
#
# Please see *01.dqn.ipynb* for detailed description.
class ReplayBuffer:
"""A simple numpy replay buffer."""
def __init__(self, obs_dim: int, size: int, batch_size: int = 32):
self.obs_buf = np.zeros([size, obs_dim], dtype=np.float32)
self.next_obs_buf = np.zeros([size, obs_dim], dtype=np.float32)
self.acts_buf = np.zeros([size], dtype=np.float32)
self.rews_buf = np.zeros([size], dtype=np.float32)
self.done_buf = np.zeros(size, dtype=np.float32)
self.max_size, self.batch_size = size, batch_size
self.ptr, self.size, = 0, 0
def store(
self,
obs: np.ndarray,
act: np.ndarray,
rew: float,
next_obs: np.ndarray,
done: bool,
):
self.obs_buf[self.ptr] = obs
self.next_obs_buf[self.ptr] = next_obs
self.acts_buf[self.ptr] = act
self.rews_buf[self.ptr] = rew
self.done_buf[self.ptr] = done
self.ptr = (self.ptr + 1) % self.max_size
self.size = min(self.size + 1, self.max_size)
def sample_batch(self) -> Dict[str, np.ndarray]:
idxs = np.random.choice(self.size, size=self.batch_size, replace=False)
return dict(obs=self.obs_buf[idxs],
next_obs=self.next_obs_buf[idxs],
acts=self.acts_buf[idxs],
rews=self.rews_buf[idxs],
done=self.done_buf[idxs])
def __len__(self) -> int:
return self.size
# ## Prioritized replay Buffer
#
# The key concept of PER's implementation is *Segment Tree*. It efficiently stores and samples transitions while managing the priorities of them. We recommend you understand how it works before you move on. Here are references for you:
#
# - In Korean: https://mrsyee.github.io/rl/2019/01/25/PER-sumtree/
# - In English: https://www.geeksforgeeks.org/segment-tree-set-1-sum-of-given-range/
class PrioritizedReplayBuffer(ReplayBuffer):
"""Prioritized Replay buffer.
Attributes:
max_priority (float): max priority
tree_ptr (int): next index of tree
alpha (float): alpha parameter for prioritized replay buffer
sum_tree (SumSegmentTree): sum tree for prior
min_tree (MinSegmentTree): min tree for min prior to get max weight
"""
def __init__(
self,
obs_dim: int,
size: int,
batch_size: int = 32,
alpha: float = 0.6
):
"""Initialization."""
assert alpha >= 0
super(PrioritizedReplayBuffer, self).__init__(obs_dim, size, batch_size)
self.max_priority, self.tree_ptr = 1.0, 0
self.alpha = alpha
# capacity must be positive and a power of 2.
tree_capacity = 1
while tree_capacity < self.max_size:
tree_capacity *= 2
self.sum_tree = SumSegmentTree(tree_capacity)
self.min_tree = MinSegmentTree(tree_capacity)
def store(
self,
obs: np.ndarray,
act: int,
rew: float,
next_obs: np.ndarray,
done: bool
):
"""Store experience and priority."""
super().store(obs, act, rew, next_obs, done)
self.sum_tree[self.tree_ptr] = self.max_priority ** self.alpha
self.min_tree[self.tree_ptr] = self.max_priority ** self.alpha
self.tree_ptr = (self.tree_ptr + 1) % self.max_size
def sample_batch(self, beta: float = 0.4) -> Dict[str, np.ndarray]:
"""Sample a batch of experiences."""
assert len(self) >= self.batch_size
assert beta > 0
indices = self._sample_proportional()
obs = self.obs_buf[indices]
next_obs = self.next_obs_buf[indices]
acts = self.acts_buf[indices]
rews = self.rews_buf[indices]
done = self.done_buf[indices]
weights = np.array([self._calculate_weight(i, beta) for i in indices])
return dict(
obs=obs,
next_obs=next_obs,
acts=acts,
rews=rews,
done=done,
weights=weights,
indices=indices,
)
def update_priorities(self, indices: List[int], priorities: np.ndarray):
"""Update priorities of sampled transitions."""
assert len(indices) == len(priorities)
for idx, priority in zip(indices, priorities):
assert priority > 0
assert 0 <= idx < len(self)
self.sum_tree[idx] = priority ** self.alpha
self.min_tree[idx] = priority ** self.alpha
self.max_priority = max(self.max_priority, priority)
def _sample_proportional(self) -> List[int]:
"""Sample indices based on proportions."""
indices = []
p_total = self.sum_tree.sum(0, len(self) - 1)
segment = p_total / self.batch_size
for i in range(self.batch_size):
a = segment * i
b = segment * (i + 1)
upperbound = random.uniform(a, b)
idx = self.sum_tree.retrieve(upperbound)
indices.append(idx)
return indices
def _calculate_weight(self, idx: int, beta: float):
"""Calculate the weight of the experience at idx."""
# get max weight
p_min = self.min_tree.min() / self.sum_tree.sum()
max_weight = (p_min * len(self)) ** (-beta)
# calculate weights
p_sample = self.sum_tree[idx] / self.sum_tree.sum()
weight = (p_sample * len(self)) ** (-beta)
weight = weight / max_weight
return weight
# ## Network
#
# We are going to use a simple network architecture with three fully connected layers and two non-linearity functions (ReLU).
class Network(nn.Module):
def __init__(self, in_dim: int, out_dim: int):
"""Initialization."""
super(Network, self).__init__()
self.layers = nn.Sequential(
nn.Linear(in_dim, 128),
nn.ReLU(),
nn.Linear(128, 128),
nn.ReLU(),
nn.Linear(128, out_dim)
)
def forward(self, x: torch.Tensor) -> torch.Tensor:
"""Forward method implementation."""
return self.layers(x)
# ## DQN + PER Agent
#
# Here is a summary of DQNAgent class.
#
# | Method | Note |
# | --- | --- |
# |select_action | select an action from the input state. |
# |step | take an action and return the response of the env. |
# |compute_dqn_loss | return dqn loss. |
# |update_model | update the model by gradient descent. |
# |target_hard_update| hard update from the local model to the target model.|
# |train | train the agent during num_frames. |
# |test | test the agent (1 episode). |
# |plot | plot the training progresses. |
#
#
# All differences from pure DQN are noted with comments - PER.
#
# #### __init__
#
# Here, we use PrioritizedReplayBuffer, instead of ReplayBuffer, and use hold 2 more parameters beta and priority epsilon which are used to calculate weights and new priorities respectively.
#
# #### compute_dqn_loss & update_model
#
# It returns every loss per each sample for importance sampling before average. After updating the nework, it is necessary to update priorities of all sampled experiences.
#
# #### train
#
# beta linearly increases to 1 at every training step.
class DQNAgent:
"""DQN Agent interacting with environment.
Attribute:
env (gym.Env): openAI Gym environment
memory (ReplayBuffer): replay memory to store transitions
batch_size (int): batch size for sampling
epsilon (float): parameter for epsilon greedy policy
epsilon_decay (float): step size to decrease epsilon
max_epsilon (float): max value of epsilon
min_epsilon (float): min value of epsilon
target_update (int): period for target model's hard update
gamma (float): discount factor
dqn (Network): model to train and select actions
dqn_target (Network): target model to update
optimizer (torch.optim): optimizer for training dqn
transition (list): transition information including
state, action, reward, next_state, done
beta (float): determines how much importance sampling is used
prior_eps (float): guarantees every transition can be sampled
"""
def __init__(
self,
env: gym.Env,
memory_size: int,
batch_size: int,
target_update: int,
epsilon_decay: float,
max_epsilon: float = 1.0,
min_epsilon: float = 0.1,
gamma: float = 0.99,
# PER parameters
alpha: float = 0.2,
beta: float = 0.6,
prior_eps: float = 1e-6,
):
"""Initialization.
Args:
env (gym.Env): openAI Gym environment
memory_size (int): length of memory
batch_size (int): batch size for sampling
target_update (int): period for target model's hard update
epsilon_decay (float): step size to decrease epsilon
lr (float): learning rate
max_epsilon (float): max value of epsilon
min_epsilon (float): min value of epsilon
gamma (float): discount factor
alpha (float): determines how much prioritization is used
beta (float): determines how much importance sampling is used
prior_eps (float): guarantees every transition can be sampled
"""
obs_dim = env.observation_space.shape[0]
action_dim = env.action_space.n
self.env = env
self.batch_size = batch_size
self.epsilon = max_epsilon
self.epsilon_decay = epsilon_decay
self.max_epsilon = max_epsilon
self.min_epsilon = min_epsilon
self.target_update = target_update
self.gamma = gamma
# device: cpu / gpu
self.device = torch.device(
"cuda" if torch.cuda.is_available() else "cpu"
)
print(self.device)
# PER
# In DQN, We used "ReplayBuffer(obs_dim, memory_size, batch_size)"
self.beta = beta
self.prior_eps = prior_eps
self.memory = PrioritizedReplayBuffer(
obs_dim, memory_size, batch_size, alpha
)
# networks: dqn, dqn_target
self.dqn = Network(obs_dim, action_dim).to(self.device)
self.dqn_target = Network(obs_dim, action_dim).to(self.device)
self.dqn_target.load_state_dict(self.dqn.state_dict())
self.dqn_target.eval()
# optimizer
self.optimizer = optim.Adam(self.dqn.parameters())
# transition to store in memory
self.transition = list()
# mode: train / test
self.is_test = False
def select_action(self, state: np.ndarray) -> np.ndarray:
"""Select an action from the input state."""
# epsilon greedy policy
if self.epsilon > np.random.random():
selected_action = self.env.action_space.sample()
else:
selected_action = self.dqn(
torch.FloatTensor(state).to(self.device)
).argmax()
selected_action = selected_action.detach().cpu().numpy()
if not self.is_test:
self.transition = [state, selected_action]
return selected_action
def step(self, action: np.ndarray) -> Tuple[np.ndarray, np.float64, bool]:
"""Take an action and return the response of the env."""
next_state, reward, done, _ = self.env.step(action)
if not self.is_test:
self.transition += [reward, next_state, done]
self.memory.store(*self.transition)
return next_state, reward, done
def update_model(self) -> torch.Tensor:
"""Update the model by gradient descent."""
# PER needs beta to calculate weights
samples = self.memory.sample_batch(self.beta)
weights = torch.FloatTensor(
samples["weights"].reshape(-1, 1)
).to(self.device)
indices = samples["indices"]
# PER: importance sampling before average
elementwise_loss = self._compute_dqn_loss(samples)
loss = torch.mean(elementwise_loss * weights)
self.optimizer.zero_grad()
loss.backward()
self.optimizer.step()
# PER: update priorities
loss_for_prior = elementwise_loss.detach().cpu().numpy()
new_priorities = loss_for_prior + self.prior_eps
self.memory.update_priorities(indices, new_priorities)
return loss.item()
def train(self, num_frames: int, plotting_interval: int = 200):
"""Train the agent."""
self.is_test = False
state = self.env.reset()
update_cnt = 0
epsilons = []
losses = []
scores = []
score = 0
for frame_idx in range(1, num_frames + 1):
action = self.select_action(state)
next_state, reward, done = self.step(action)
state = next_state
score += reward
# PER: increase beta
fraction = min(frame_idx / num_frames, 1.0)
self.beta = self.beta + fraction * (1.0 - self.beta)
# if episode ends
if done:
state = self.env.reset()
scores.append(score)
score = 0
# if training is ready
if len(self.memory) >= self.batch_size:
loss = self.update_model()
losses.append(loss)
update_cnt += 1
# linearly decrease epsilon
self.epsilon = max(
self.min_epsilon, self.epsilon - (
self.max_epsilon - self.min_epsilon
) * self.epsilon_decay
)
epsilons.append(self.epsilon)
# if hard update is needed
if update_cnt % self.target_update == 0:
self._target_hard_update()
# plotting
if frame_idx % plotting_interval == 0:
self._plot(frame_idx, scores, losses, epsilons)
self.env.close()
def test(self) -> List[np.ndarray]:
"""Test the agent."""
self.is_test = True
state = self.env.reset()
done = False
score = 0
frames = []
while not done:
frames.append(self.env.render(mode="rgb_array"))
action = self.select_action(state)
next_state, reward, done = self.step(action)
state = next_state
score += reward
print("score: ", score)
self.env.close()
return frames
def _compute_dqn_loss(self, samples: Dict[str, np.ndarray]) -> torch.Tensor:
"""Return dqn loss."""
device = self.device # for shortening the following lines
state = torch.FloatTensor(samples["obs"]).to(device)
next_state = torch.FloatTensor(samples["next_obs"]).to(device)
action = torch.LongTensor(samples["acts"].reshape(-1, 1)).to(device)
reward = torch.FloatTensor(samples["rews"].reshape(-1, 1)).to(device)
done = torch.FloatTensor(samples["done"].reshape(-1, 1)).to(device)
# G_t = r + gamma * v(s_{t+1}) if state != Terminal
# = r otherwise
curr_q_value = self.dqn(state).gather(1, action)
next_q_value = self.dqn_target(
next_state
).max(dim=1, keepdim=True)[0].detach()
mask = 1 - done
target = (reward + self.gamma * next_q_value * mask).to(self.device)
# calculate element-wise dqn loss
elementwise_loss = F.smooth_l1_loss(curr_q_value, target, reduction="none")
return elementwise_loss
def _target_hard_update(self):
"""Hard update: target <- local."""
self.dqn_target.load_state_dict(self.dqn.state_dict())
def _plot(
self,
frame_idx: int,
scores: List[float],
losses: List[float],
epsilons: List[float],
):
"""Plot the training progresses."""
clear_output(True)
plt.figure(figsize=(20, 5))
plt.subplot(131)
plt.title('frame %s. score: %s' % (frame_idx, np.mean(scores[-10:])))
plt.plot(scores)
plt.subplot(132)
plt.title('loss')
plt.plot(losses)
plt.subplot(133)
plt.title('epsilons')
plt.plot(epsilons)
plt.show()
# ## Environment
#
# You can see the [code](https://github.com/openai/gym/blob/master/gym/envs/classic_control/cartpole.py) and [configurations](https://github.com/openai/gym/blob/master/gym/envs/__init__.py#L53) of CartPole-v0 from OpenAI's repository.
# environment
env_id = "CartPole-v0"
env = gym.make(env_id)
if IN_COLAB:
env = gym.wrappers.Monitor(env, "videos", force=True)
# ## Set random seed
# +
seed = 777
def seed_torch(seed):
torch.manual_seed(seed)
if torch.backends.cudnn.enabled:
torch.backends.cudnn.benchmark = False
torch.backends.cudnn.deterministic = True
np.random.seed(seed)
random.seed(seed)
seed_torch(seed)
env.seed(seed)
# -
# ## Initialize
# +
# parameters
num_frames = 20000
memory_size = 2000
batch_size = 32
target_update = 100
epsilon_decay = 1 / 2000
# train
agent = DQNAgent(env, memory_size, batch_size, target_update, epsilon_decay)
# -
# ## Train
agent.train(num_frames)
# ## Test
#
# Run the trained agent (1 episode).
frames = agent.test()
# ## Render
# +
if IN_COLAB: # for colab
import base64
import glob
import io
import os
from IPython.display import HTML, display
def ipython_show_video(path: str) -> None:
"""Show a video at `path` within IPython Notebook."""
if not os.path.isfile(path):
raise NameError("Cannot access: {}".format(path))
video = io.open(path, "r+b").read()
encoded = base64.b64encode(video)
display(HTML(
data="""
<video alt="test" controls>
<source src="data:video/mp4;base64,{0}" type="video/mp4"/>
</video>
""".format(encoded.decode("ascii"))
))
list_of_files = glob.glob("videos/*.mp4")
latest_file = max(list_of_files, key=os.path.getctime)
print(latest_file)
ipython_show_video(latest_file)
else: # for jupyter
from matplotlib import animation
from JSAnimation.IPython_display import display_animation
from IPython.display import display
def display_frames_as_gif(frames: List[np.ndarray]) -> None:
"""Displays a list of frames as a gif, with controls."""
patch = plt.imshow(frames[0])
plt.axis('off')
def animate(i):
patch.set_data(frames[i])
anim = animation.FuncAnimation(
plt.gcf(), animate, frames = len(frames), interval=50
)
display(display_animation(anim, default_mode='loop'))
# display
display_frames_as_gif(frames)
| 03.per.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Install latest version from GitHub
# !pip install -q -U git+https://github.com/jdvelasq/techminer
# # Direct queries over data
# Some queries can be made directly over the dataframe without recurring to special functions or code.
# +
import pandas as pd
from techminer import DataFrame, Plot, heatmap
#
# Data loading
#
df = DataFrame(
pd.read_json(
"https://raw.githubusercontent.com/jdvelasq/techminer/master/data/tutorial/"
+ "cleaned-data.json",
orient="records",
lines=True,
)
)
#
# Columns of the dataframe
#
df.columns
# -
#
# Number of records in the dataframe
#
len(df)
#
# Data coverage
#
df.coverage().head()
#
# Number of terms
#
df.count_report()
#
# Number of terms for individual columns
#
df.count_terms('Author Keywords')
#
# Top N most cited documents
#
df.most_cited_documents().head(10)
#
# Or
#
df.citations_by_term('Title').head(10)
#
# Most cited authors
#
df.most_cited_authors().head()
#
# Top 10 most cited authors
#
df.most_cited_authors().head(10).Authors
#
# Top 10 most frequent authors
#
df.documents_by_term('Authors').head(10).Authors
# ## Record extraction by IDs
#
# IDs for top five documents
#
IDs = df.citations_by_term('Title')['ID'].head(5)
IDs
#
# Selects `Title` and `Authors` by IDs
#
df.get_rows_by_IDs(IDs)[['Title', 'Authors']]
| sphinx/tutorial/13-direct-queries.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # 8. Discrete Probability Distributions
# -
# We use `scipy.stats` to demonstrate the pdf, cdf, and sampling from several distribution families. First we import a few helpful libraries.
# +
from scipy.stats import bernoulli, binom, poisson
import matplotlib.pyplot as plt
import numpy as np
import ipywidgets as widgets
# Properties for plots
plt.style.use([{
"figure.figsize":(12,9), # Figure size
"xtick.labelsize": "large", # Font size of the X-ticks
"ytick.labelsize": "large", # Font size of the Y-ticks
"legend.fontsize": "x-large", # Font size of the legend
"axes.labelsize": "x-large", # Font size of labels
"axes.titlesize": "xx-large", # Font size of title
"axes.spines.top": False,
"axes.spines.right": False,
},'seaborn-poster'])
# -
# ## 8.1 Bernoulli
# Let's explore the basic methods for a Bernoulli random variable.
print(bernoulli.pmf(0,p=.3))
print(bernoulli.pmf(range(3),p=.3))
print(bernoulli.cdf([0,.5,1,1.5],p=.3))
# Next plot the pdf and cdf. For simplicity we are omitting labels and legend, we will see how to plot them in the next subsection.
plt.stem([-0.2,0,1,1.2],bernoulli.pmf([-0.2,0,1,1.2],p=.3))
plt.plot(np.linspace(-0.1,1.1,1200),bernoulli.cdf(np.linspace(-0.1,1.1,1200),p=0.3),'g')
plt.xlim([-0.1,1.1])
plt.ylim([-0.2,1.1])
plt.show()
# Next we generate and then plot Bernoulli samples.
print(bernoulli.rvs(size=10,p=0.3))
plt.hist(bernoulli.rvs(size=10,p=0.3),normed=True)
plt.show()
# ## 8.2 Binomial
# You can calculate the pdf and cdf of the Binomial distribution the same way we did for Bernoulli, just replace `bernoulli.pmf` by `binom.pmf` etc. and including the appropriate arguments. Here is one example.
print(binom.rvs(size=50,n=20, p=0.4))
# + [markdown] slideshow={"slide_type": "slide"}
# The following code plots the probability mass function (PMF) of $B_{p,n}$, the binomial distribution with parameters $p$ and $n$. It contains interactive sliders that you can use to vary $n$ over the interval $[0,30]$ and $p$ over the interval $[0, 1]$.
# -
@widgets.interact(n=(0,30),p=(0.0,1.0),samples=(1,1000), continuous_update=False)
def plot_pmf(n, p,samples=100,histogram=False):
'''
Plot the probability mass function of Binom(n, p)
'''
k = np.arange(0, n + 1)
P_binom = binom.pmf(k, n, p)
# This plots a bar plot
# plt.bar(k, P_binom,color='b')
plt.plot(k, P_binom, '-o', color='r')
if histogram:
height,y = np.histogram(binom.rvs(size=samples,n=n,p=p),range=(0,n),bins=n+1,normed=True)
plt.bar(k,height,color='r')
plt.title('PMF of Bin(%i, %.2f)' % (n, p))
plt.xlabel('k')
plt.ylabel('$B_{20,0.3}(k)$')
plt.show()
# ## 8.3 Poisson
# ### 8.3.1 PMF
# We follow the same procedure to plot the Poisson PMF.
@widgets.interact(n=(0,50),samples=(1,1000),λ=(0.0,30.0))
def f(n, λ, samples=100,histogram=False):
k = np.arange(0, n+1)
P_poisson = poisson.pmf(k, λ)
plt.plot(k, P_poisson, '-o')
if histogram:
height,y = np.histogram(poisson.rvs(size=samples,mu=λ),range=(0,n),bins=n+1,normed=True)
plt.bar(k,height,color='r')
plt.title('PMF of Poisson(%i)' %λ)
plt.xlabel('Number of Events')
plt.ylabel('Probability of Number of Events')
plt.show()
# ### 8.3.2 Poisson Approximation of the Binomial Distribution
# Observe how well Poisson$(np)$ approximates Binomial$(n, p)$ for small values of $p$.
# +
import numpy as np
import matplotlib.pyplot as plt
e = np.e
@widgets.interact(n=(2,1000),p=(0.0,.2,0.001),continuous_update=False)
def f(n, p):
k = np.arange(0, n+1)
x = np.linspace(0, n+1, 1000)
λ = n*p
stddev = λ**0.5
P_poisson = poisson.pmf(k, λ)
P_binom = binom.pmf(k, n, p)
plt.plot(k, P_poisson, 'r', label = "Poisson(%0.2f)" %λ)
plt.plot(k, P_binom, 'b-', label = "Bin(%i, %0.2f)" %(n,p))
plt.title('Poisson Approximation of Binomial')
plt.xlabel('n')
plt.ylabel('y')
plt.legend()
plt.show()
print('|| P_Poisson - P_Binomial ||\u2081 = ',sum(abs(P_poisson-P_binom)))
# -
# ## 8.4 Geometric Distribution
# ### 8.4.1 PMF and CDF
@widgets.interact(n=(1,200),p=(0.0,1.0))
def f(n, p, CDF=False):
x = np.arange(1, n + 1)
y = [((1 - p)**(z - 1)) * p for z in x]
z = [(1 - (1 - p)**zz) for zz in x]
plt.plot(x, y, 'o-', label='PDF')
if CDF == True:
plt.plot(x, z, 'ro-', label='CDF')
if n == 1:
plt.plot([0, 1], [p, p], 'b')
plt.xticks([1])
plt.xlabel('n')
plt.ylabel('y')
plt.title('PMF of Geometric(%0.2f)' % p, fontsize=20)
plt.legend()
plt.show()
| Week 8 _ Distribution Families/Topic8_Lecture.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + init_cell=true
# %logstop
# %logstart -ortq ~/.logs/pw.py append
# %matplotlib inline
import matplotlib
import seaborn as sns
sns.set()
matplotlib.rcParams['figure.dpi'] = 144
# -
from static_grader import grader
# # PW Miniproject
# ## Introduction
#
# The objective of this miniproject is to exercise your ability to use basic Python data structures, define functions, and control program flow. We will be using these concepts to perform some fundamental data wrangling tasks such as joining data sets together, splitting data into groups, and aggregating data into summary statistics.
# **Please do not use `pandas` or `numpy` to answer these questions.**
#
# We will be working with medical data from the British NHS on prescription drugs. Since this is real data, it contains many ambiguities that we will need to confront in our analysis. This is commonplace in data science, and is one of the lessons you will learn in this miniproject.
# ## Downloading the data
#
# We first need to download the data we'll be using from Amazon S3:
# + language="bash"
# mkdir pw-data
# wget http://dataincubator-wqu.s3.amazonaws.com/pwdata/201701scripts_sample.json.gz -nc -P ./pw-data
# wget http://dataincubator-wqu.s3.amazonaws.com/pwdata/practices.json.gz -nc -P ./pw-data
# -
# ## Loading the data
#
# The first step of the project is to read in the data. We will discuss reading and writing various kinds of files later in the course, but the code below should get you started.
import gzip
import simplejson as json
# +
with gzip.open('./pw-data/201701scripts_sample.json.gz', 'rb') as f:
scripts = json.load(f)
with gzip.open('./pw-data/practices.json.gz', 'rb') as f:
practices = json.load(f)
# -
# This data set comes from Britain's National Health Service. The `scripts` variable is a list of prescriptions issued by NHS doctors. Each prescription is represented by a dictionary with various data fields: `'practice'`, `'bnf_code'`, `'bnf_name'`, `'quantity'`, `'items'`, `'nic'`, and `'act_cost'`.
scripts[:2]
# A [glossary of terms](http://webarchive.nationalarchives.gov.uk/20180328130852tf_/http://content.digital.nhs.uk/media/10686/Download-glossary-of-terms-for-GP-prescribing---presentation-level/pdf/PLP_Presentation_Level_Glossary_April_2015.pdf/) and [FAQ](http://webarchive.nationalarchives.gov.uk/20180328130852tf_/http://content.digital.nhs.uk/media/10048/FAQs-Practice-Level-Prescribingpdf/pdf/PLP_FAQs_April_2015.pdf/) is available from the NHS regarding the data. Below we supply a data dictionary briefly describing what these fields mean.
#
# | Data field |Description|
# |:----------:|-----------|
# |`'practice'`|Code designating the medical practice issuing the prescription|
# |`'bnf_code'`|British National Formulary drug code|
# |`'bnf_name'`|British National Formulary drug name|
# |`'quantity'`|Number of capsules/quantity of liquid/grams of powder prescribed|
# | `'items'` |Number of refills (e.g. if `'quantity'` is 30 capsules, 3 `'items'` means 3 bottles of 30 capsules)|
# | `'nic'` |Net ingredient cost|
# |`'act_cost'`|Total cost including containers, fees, and discounts|
# The `practices` variable is a list of member medical practices of the NHS. Each practice is represented by a dictionary containing identifying information for the medical practice. Most of the data fields are self-explanatory. Notice the values in the `'code'` field of `practices` match the values in the `'practice'` field of `scripts`.
practices[:2]
# In the following questions we will ask you to explore this data set. You may need to combine pieces of the data set together in order to answer some questions. Not every element of the data set will be used in answering the questions.
# ## Question 1: summary_statistics
#
# Our beneficiary data (`scripts`) contains quantitative data on the number of items dispensed (`'items'`), the total quantity of item dispensed (`'quantity'`), the net cost of the ingredients (`'nic'`), and the actual cost to the patient (`'act_cost'`). Whenever working with a new data set, it can be useful to calculate summary statistics to develop a feeling for the volume and character of the data. This makes it easier to spot trends and significant features during further stages of analysis.
#
# Calculate the sum, mean, standard deviation, and quartile statistics for each of these quantities. Format your results for each quantity as a list: `[sum, mean, standard deviation, 1st quartile, median, 3rd quartile]`. We'll create a `tuple` with these lists for each quantity as a final result.
# +
def describe(key):
total = 0
avg = 0
s = 0
q25 = 0
med = 0
q75 = 0
return (total, avg, s, q25, med, q75)
# -
summary = [('items', describe('items')),
('quantity', describe('quantity')),
('nic', describe('nic')),
('act_cost', describe('act_cost'))]
grader.score.pw__summary_statistics(summary)
# ## Question 2: most_common_item
#
# Often we are not interested only in how the data is distributed in our entire data set, but within particular groups -- for example, how many items of each drug (i.e. `'bnf_name'`) were prescribed? Calculate the total items prescribed for each `'bnf_name'`. What is the most commonly prescribed `'bnf_name'` in our data?
#
# To calculate this, we first need to split our data set into groups corresponding with the different values of `'bnf_name'`. Then we can sum the number of items dispensed within in each group. Finally we can find the largest sum.
#
# We'll use `'bnf_name'` to construct our groups. You should have *5619* unique values for `'bnf_name'`.
bnf_names = ...
assert(len(bnf_names) == 5619)
# We want to construct "groups" identified by `'bnf_name'`, where each group is a collection of prescriptions (i.e. dictionaries from `scripts`). We'll construct a dictionary called `groups`, using `bnf_names` as the keys. We'll represent a group with a `list`, since we can easily append new members to the group. To split our `scripts` into groups by `'bnf_name'`, we should iterate over `scripts`, appending prescription dictionaries to each group as we encounter them.
groups = {name: [] for name in bnf_names}
for script in scripts:
# INSERT ...
# Now that we've constructed our groups we should sum up `'items'` in each group and find the `'bnf_name'` with the largest sum. The result, `max_item`, should have the form `[(bnf_name, item total)]`, e.g. `[('Foobar', 2000)]`.
max_item = [("", 0)]
# **TIP:** If you are getting an error from the grader below, please make sure your answer conforms to the correct format of `[(bnf_name, item total)]`.
grader.score.pw__most_common_item(max_item)
# **Challenge:** Write a function that constructs groups as we did above. The function should accept a list of dictionaries (e.g. `scripts` or `practices`) and a tuple of fields to `groupby` (e.g. `('bnf_name')` or `('bnf_name', 'post_code')`) and returns a dictionary of groups. The following questions will require you to aggregate data in groups, so this could be a useful function for the rest of the miniproject.
def group_by_field(data, fields):
groups = {}
return groups
# +
groups = group_by_field(scripts, ('bnf_name',))
test_max_item = ...
assert test_max_item == max_item
# -
# ## Question 3: postal_totals
#
# Our data set is broken up among different files. This is typical for tabular data to reduce redundancy. Each table typically contains data about a particular type of event, processes, or physical object. Data on prescriptions and medical practices are in separate files in our case. If we want to find the total items prescribed in each postal code, we will have to _join_ our prescription data (`scripts`) to our clinic data (`practices`).
#
# Find the total items prescribed in each postal code, representing the results as a list of tuples `(post code, total items prescribed)`. Sort your results ascending alphabetically by post code and take only results from the first 100 post codes. Only include post codes if there is at least one prescription from a practice in that post code.
#
# **NOTE:** Some practices have multiple postal codes associated with them. Use the alphabetically first postal code.
# We can join `scripts` and `practices` based on the fact that `'practice'` in `scripts` matches `'code'` in `practices`. However, we must first deal with the repeated values of `'code'` in `practices`. We want the alphabetically first postal codes.
practice_postal = {}
for practice in practices:
if practice['code'] in practice_postal:
practice_postal[practice['code']] = ...
else:
practice_postal[practice['code']] = ...
# **Challenge:** This is an aggregation of the practice data grouped by practice codes. Write an alternative implementation of the above cell using the `group_by_field` function you defined previously.
assert practice_postal['K82019'] == 'HP21 8TR'
# Now we can join `practice_postal` to `scripts`.
joined = scripts[:]
for script in joined:
script['post_code'] = ...
# Finally we'll group the prescription dictionaries in `joined` by `'post_code'` and sum up the items prescribed in each group, as we did in the previous question.
items_by_post = ...
# +
postal_totals = [('B11 4BW', 20673)] * 100
grader.score.pw__postal_totals(postal_totals)
# -
# ## Question 4: items_by_region
#
# Now we'll combine the techniques we've developed to answer a more complex question. Find the most commonly dispensed item in each postal code, representing the results as a list of tuples (`post_code`, `bnf_name`, amount dispensed as proportion of total). Sort your results ascending alphabetically by post code and take only results from the first 100 post codes.
#
# **NOTE:** We'll continue to use the `joined` variable we created before, where we've chosen the alphabetically first postal code for each practice. Additionally, some postal codes will have multiple `'bnf_name'` with the same number of items prescribed for the maximum. In this case, we'll take the alphabetically first `'bnf_name'`.
# Now we need to calculate the total items of each `'bnf_name'` prescribed in each `'post_code'`. Use the techniques we developed in the previous questions to calculate these totals. You should have 141196 `('post_code', 'bnf_name')` groups.
total_items_by_bnf_post = ...
assert len(total_items_by_bnf_post) == 141196
# Let's use `total_items` to find the maximum item total for each postal code. To do this, we will want to regroup `total_items_by_bnf_post` by `'post_code'` only, not by `('post_code', 'bnf_name')`. First let's turn `total_items` into a list of dictionaries (similar to `scripts` or `practices`) and then group it by `'post_code'`. You should have 118 groups in the resulting `total_items_by_post` after grouping `total_items` by `'post_code'`.
total_items = ...
assert len(total_items_by_post) == 118
# Now we will aggregate the groups in `total_by_item_post` to create `max_item_by_post`. Some `'bnf_name'` have the same item total within a given postal code. Therefore, if more than one `'bnf_name'` has the maximum item total in a given postal code, we'll take the alphabetically first `'bnf_name'`. We can do this by [sorting](https://docs.python.org/2.7/howto/sorting.html) each group according to the item total and `'bnf_name'`.
max_item_by_post = ...
# In order to express the item totals as a proportion of the total amount of items prescribed across all `'bnf_name'` in a postal code, we'll need to use the total items prescribed that we previously calculated as `items_by_post`. Calculate the proportions for the most common `'bnf_names'` for each postal code. Format your answer as a list of tuples: `[(post_code, bnf_name, total)]`
items_by_region = [('B11 4BW', 'Salbutamol_Inha 100mcg (200 D) CFF', 0.0341508247)] * 100
grader.score.pw__items_by_region(items_by_region)
# *Copyright © 2019 The Data Incubator. All rights reserved.*
| data-wrangling/miniprojects/.ipynb_checkpoints/pw-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SageMath 7.5.1
# language: ''
# name: sagemath
# ---
def xor(p,q):
return (p or q) and not(p and q)
for p in [0,1]:
for q in [0,1]:
print p,q,xor(p,q)
import jdk2py_MD as my
mitabla=[["p", "q", "r", "p iff r"]]
for p in [0,1]:
for q in [0,1]:
r=xor(p and q, p and not q)
#r=(p and q) and (p and not q)
renglon=[int(p), int(q), int(r), int(my.iff(p,r))]
mitabla.append(renglon)
show(table(mitabla))
mitabla2=[]
for p in [0,1]:
for q in [0,1]:
for r in [0,1]:
s = xor(xor(p,q), r)
t = xor(p, xor(q,r))
renglon = [int(p), int(q), int(r), int(s), int(t), int(my.iff(s,t))]
mitabla2.append(renglon)
show(table(mitabla2))
mitabla3=[["p", "q", "r", "s", "r iff s"]]
for p in [0,1]:
for q in [0,1]:
r=xor(p and not q, xor(p and q, q and not p))
s=p or q
#r=(p and q) and (p and not q)
renglon=[int(p), int(q), int(r), int(s), int(my.iff(r,s))]
mitabla3.append(renglon)
show(table(mitabla3))
| latex/MD_SMC/MD01_Actividad_03.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Computer Vision - P6
#
# ### **Carefully read the file `README.md` as well as the following instructions before start coding.**
# ## Delivery
#
# Up to **1 point out of 10** will be penalized if the following requirements are not fulfilled:
#
# - Implemented code should be commented.
#
# - The questions introduced in the exercises must be answered.
#
# - Add title to the figures to explain what is displayed.
#
# - Comments need to be in **english**.
#
# - The deliverable must be a file named **P6_Student1_Student2.zip** that includes:
# - The notebook P6_Student1_Student2.ipynb completed with the solutions to the exercises and their corresponding comments.
# - All the images used in this notebook.
#
# **Deadline (Campus Virtual): January 2th, 23:59 h**
# ## Haar-like features applied for face detection
#
# Today's exercices will practise the following:
#
# - Integral images and a classical use for fast harr-like feature computation.
# - Use of Adaboost for classification.
# - Decisions based on a user-defined threshold for balancing precision and recall.
# +
# Some useful imports for the exercises
import numpy as np
import os, warnings
warnings.filterwarnings("ignore")
import matplotlib
from PIL import Image
#from multiprocessing import Pool
#from functools import partial
from skimage import io
import time
import timeit
from matplotlib import pyplot as plt
# %matplotlib inline
# -
# **1) Build a function `to_integral_image` that computes the integral image of an input (2D) array.**
#
# The integral image must have an additional row and column full of zeros (first row and first column).
# Make sure that the values of the integral image are correct.
#
# ```python
# def to_integral_image(img_arr):
# row_sum = np.zeros(img_arr.shape)
# integral_image_arr = np.zeros((img_arr.shape[0] + 1, img_arr.shape[1] + 1))
# # Add here code
#
# return integral_image_arr
# ```
#
#
# You can make the following tests:
#
# - `sum(img_array) == ii_img_array[-1,-1]`
# - `img_array[0,:].sum() == ii_img_array[1,-1]`
#
# Plot the output of the integral image for the following array:
#
# ```
# img_array = np.array([[1,2,2,2,1],[1,2,2,2,1],[1,2,2,2,1],[1,2,2,2,1]])
# ```
#
#
#
img_array = np.array([[1,2,2,2,1],[1,2,2,2,1],[1,2,2,2,1],[1,2,2,2,1]])
img_array.shape
plt.imshow(img_array, cmap="gray", vmin=0, vmax= 5)
# +
def to_integral_image(img_arr):
"""
Calculates the integral image based on this instance's original image data.
:param img_arr: Image source data
:type img_arr: numpy.ndarray
:return Integral image for given image
:rtype: numpy.ndarray
"""
# Since the integral image needs extra zeros row and colum an extended matrix is created.
integral_image_arr = np.zeros((img_arr.shape[0] + 1, img_arr.shape[1] + 1))
# Iterative version.
#r, c = img_arr.shape
#integral_image_arr[1:, 1:] = np.array([np.array([np.sum(img_arr[:i+1, :j+1]) for j in range(c)]) for i in range(r)])
# Using numpy's function for cumulative summatory.
tmp = np.copy(img_arr)
np.cumsum(tmp, axis=0, out=tmp)
np.cumsum(tmp, axis=1, out=tmp)
integral_image_arr[1:, 1:] = tmp
return integral_image_arr
# Tests
ii_img_array = to_integral_image(img_array)
assert np.sum(img_array) == ii_img_array[-1,-1]
assert np.sum(img_array[0,:]) == ii_img_array[1,-1]
plt.imshow(ii_img_array, cmap='gray')
# -
# **2) Build a function to compute the sum of the pixel intensities within a rectangle using the integral image. The rectangle will be defined using the top left (x, y) and bottom right (x, y) coordinates.**
#
# Make the function with the following header:
# ```
# def sum_region(integral_img_arr, top_left, bottom_right):
# ```
#
def sum_region(integral_img_arr, top_left, bottom_right):
r1, c1 = top_left[0], top_left[1]
r2, c2 = bottom_right[0], bottom_right[1]
"""
For some weird reason the exercise asks for a thing and the 'correct' result is another completely
different.
The exercise ask for the summatory of the pixels within a specified rectangle, which would be computed
by: np.sum(integral_img_arr[r1:r2, c1:c2]). However the 'correct' output is achieved by summing the
specified corners and subtracting the other two.
"""
return integral_img_arr[r1, c1] + integral_img_arr[r2, c2] - integral_img_arr[r1, c2] - integral_img_arr[r2, c1]
# result you should get (12)
sum_region(ii_img_array, [1,1],[3,4])
# result you should get (32)
sum_region(ii_img_array, [0,0],[-1,-1])
# **3) Compute the integral image for all the following images:**
#
# - training images of faces: save results in **`faces_ii_training`**
# - testing images of face: save the results in **`faces_ii_testing`**
# - training images of non faces: **`non_faces_ii_training`**
# - testing images of non faces: **`non_faces_ii_testing`**
#
# To do so build a function to read all the images inside a given folder:
#
# ```python
# def load_images(path):
# images = []
# for _file in os.listdir(path):
# #### Read image
# #### Remember to scale the image (wih the max pixel intensity value)
#
# return images
# ```
pos_training_path = "trainingdata/faces"
neg_training_path = "trainingdata/nonfaces"
pos_testing_path = "trainingdata/faces/test"
neg_testing_path = "trainingdata/nonfaces/test"
def load_images(path):
# Normalizing function.
normalize = lambda x: x/np.max(x)
# Read, normalize and compute integral image of every ".png" file in the specified path, all in a cool
# and compact list comprehension.
return [to_integral_image(normalize(io.imread(path + "/" + file))) for file in os.listdir(path)
if file.endswith(".png")]
faces_ii_training = load_images(pos_training_path)
faces_ii_testing = load_images(pos_testing_path)
non_faces_ii_training = load_images(neg_training_path)
non_faces_ii_testing = load_images(neg_testing_path)
# **4) Compute the Haar features of an image**
#
# The code given will use the `sum_region` function you have implemented to compute Haar-like features.
# The following code, for example, will compute a vertical Haar-like feature
# ```python
# first = sum_region(int_img,
# self.top_left,
# (self.top_left[0] + self.width, int(self.top_left[1] + self.height / 2)))
# second = sum_region(int_img,
# (self.top_left[0], int(self.top_left[1] + self.height / 2)),
# self.bottom_right)
# score = first - second
# ```
#
# We provide you with `HaarLikeFeature` class that has build in a `get_score` function and a `get_vote` function.
#
# Your job is to
# ```python
# def _create_features(img_height, img_width, min_feature_width, max_feature_width, min_feature_height, max_feature_height):
# print('Creating Haar-like features..')
# t0 = time.time()
# features = []
# for feature in FeatureTypes:
# # FeatureTypes are just tuples
# feature_start_width = max(min_feature_width, feature[0])
# for feature_width in range(feature_start_width, max_feature_width, feature[0]):
# feature_start_height = max(min_feature_height, feature[1])
# for feature_height in range(feature_start_height, max_feature_height, feature[1]):
# # Loop over possible x values and y values
# # - For each (x,y) create the HarrLikeFeature objects.
# # - append the HaarlikeFeatures in the features list.
# # Notice that Haarlike features contain polarity, append features for polarity 1 and -1
# # The threshold can be set to 0 for all of them.
#
#
# print('\t' + str(len(features)) + ' features created.')
# print('\tTime needed for calculating Harr-like features:', time.time()-t0)
# return features
# ```
# +
def enum(**enums):
return type('Enum', (), enums)
FeatureType = enum(TWO_VERTICAL=(1, 2), TWO_HORIZONTAL=(2, 1), THREE_HORIZONTAL=(3, 1), THREE_VERTICAL=(1, 3), FOUR=(2, 2))
FeatureTypes = [FeatureType.TWO_VERTICAL, FeatureType.TWO_HORIZONTAL, FeatureType.THREE_VERTICAL, FeatureType.THREE_HORIZONTAL, FeatureType.FOUR]
class HaarLikeFeature(object):
"""
Class representing a haar-like feature.
"""
def __init__(self, feature_type, position, width, height, threshold, polarity):
"""
Creates a new haar-like feature.
:param feature_type: Type of new feature, see FeatureType enum
:type feature_type: violajonse.HaarLikeFeature.FeatureTypes
:param position: Top left corner where the feature begins (x, y)
:type position: (int, int)
:param width: Width of the feature
:type width: int
:param height: Height of the feature
:type height: int
:param threshold: Feature threshold
:type threshold: float
:param polarity: polarity of the feature -1 or 1
:type polarity: int
"""
self.type = feature_type
self.top_left = position
self.bottom_right = (position[0] + width, position[1] + height)
self.width = width
self.height = height
self.threshold = threshold
self.polarity = polarity
self.weight = 1
def get_score(self, int_img):
"""
Get score for given integral image array.
:param int_img: Integral image array
:type int_img: numpy.ndarray
:return: Score for given feature
:rtype: float
"""
score = 0
if self.type == FeatureType.TWO_VERTICAL:
first = sum_region(int_img, self.top_left, (self.top_left[0] + self.width, int(self.top_left[1] + self.height / 2)))
second = sum_region(int_img, (self.top_left[0], int(self.top_left[1] + self.height / 2)), self.bottom_right)
score = first - second
elif self.type == FeatureType.TWO_HORIZONTAL:
first = sum_region(int_img, self.top_left, (int(self.top_left[0] + self.width / 2), self.top_left[1] + self.height))
second = sum_region(int_img, (int(self.top_left[0] + self.width / 2), self.top_left[1]), self.bottom_right)
score = first - second
elif self.type == FeatureType.THREE_HORIZONTAL:
first = sum_region(int_img, self.top_left, (int(self.top_left[0] + self.width / 3), self.top_left[1] + self.height))
second = sum_region(int_img, (int(self.top_left[0] + self.width / 3), self.top_left[1]), (int(self.top_left[0] + 2 * self.width / 3), self.top_left[1] + self.height))
third = sum_region(int_img, (int(self.top_left[0] + 2 * self.width / 3), self.top_left[1]), self.bottom_right)
score = first - second + third
elif self.type == FeatureType.THREE_VERTICAL:
first = sum_region(int_img, self.top_left, (self.bottom_right[0], int(self.top_left[1] + self.height / 3)))
second = sum_region(int_img, (self.top_left[0], int(self.top_left[1] + self.height / 3)), (self.bottom_right[0], int(self.top_left[1] + 2 * self.height / 3)))
third = sum_region(int_img, (self.top_left[0], int(self.top_left[1] + 2 * self.height / 3)), self.bottom_right)
score = first - second + third
elif self.type == FeatureType.FOUR:
# top left area
first = sum_region(int_img, self.top_left, (int(self.top_left[0] + self.width / 2), int(self.top_left[1] + self.height / 2)))
# top right area
second = sum_region(int_img, (int(self.top_left[0] + self.width / 2), self.top_left[1]), (self.bottom_right[0], int(self.top_left[1] + self.height / 2)))
# bottom left area
third = sum_region(int_img, (self.top_left[0], int(self.top_left[1] + self.height / 2)), (int(self.top_left[0] + self.width / 2), self.bottom_right[1]))
# bottom right area
fourth = sum_region(int_img, (int(self.top_left[0] + self.width / 2), int(self.top_left[1] + self.height / 2)), self.bottom_right)
score = first - second - third + fourth
return score
def get_vote(self, int_img):
"""
Get vote of this feature for given integral image.
:param int_img: Integral image array
:type int_img: numpy.ndarray
:return: 1 iff this feature votes positively, otherwise -1
:rtype: int
"""
score = self.get_score(int_img)
return self.weight * (1 if score < self.polarity * self.threshold else -1)
# -
# > ⚠️ **NOTICE:** Since the practicum can only be implemented in a Linux machine (Windows machine end up in an endless loop), the code meant to be ran with Pool to reduce execution time has been replaced with a nested loop. An attempt was made to replace the usage of the Pool library with Parallel, but it was taking more time than the execution time with nested loops. So there's that.
# > Expect a longer execution time.
# +
def learn(positive_iis, negative_iis, num_classifiers=-1, min_feature_width=1, max_feature_width=-1, min_feature_height=1, max_feature_height=-1):
"""
Selects a set of classifiers. Iteratively takes the best classifiers based
on a weighted error.
:param positive_iis: List of positive integral image examples
:type positive_iis: list[numpy.ndarray]
:param negative_iis: List of negative integral image examples
:type negative_iis: list[numpy.ndarray]
:param num_classifiers: Number of classifiers to select, -1 will use all
classifiers
:type num_classifiers: int
:return: List of selected features
:rtype: list[violajones.HaarLikeFeature.HaarLikeFeature]
"""
num_pos = len(positive_iis)
num_neg = len(negative_iis)
num_imgs = num_pos + num_neg
img_height, img_width = positive_iis[0].shape
# Maximum feature width and height default to image width and height
max_feature_height = img_height if max_feature_height == -1 else max_feature_height
max_feature_width = img_width if max_feature_width == -1 else max_feature_width
# Create initial weights and labels
pos_weights = np.ones(num_pos) * 1. / (2 * num_pos)
neg_weights = np.ones(num_neg) * 1. / (2 * num_neg)
weights = np.hstack((pos_weights, neg_weights))
labels = np.hstack((np.ones(num_pos), np.ones(num_neg) * -1))
images = positive_iis + negative_iis
# Create features for all sizes and locations
features = _create_features(img_height, img_width, min_feature_width, max_feature_width, min_feature_height, max_feature_height)
num_features = len(features)
feature_indexes = list(range(num_features))
num_classifiers = num_features if num_classifiers == -1 else num_classifiers
print('Calculating scores for images..')
t0 = time.time()
votes = np.zeros((num_imgs, num_features))
# Use as many workers as there are CPUs
#pool = Pool(processes=None)
#for i in range(num_imgs):
# votes[i, :] = np.array(list(pool.map(partial(_get_feature_vote, image=images[i]), features)))
for i in range(num_imgs):
votes[i, :] = np.array([_get_feature_vote(features[j], images[i]) for j in range(len(features))])
#for i in range(num_imgs):
# votes[i, :] = Parallel(n_jobs=multiprocessing.cpu_count())(delayed(
# partial(_get_feature_vote))(j, images[j]) for j in features)
print('\tTime needed for calculating scores:', time.time()-t0)
# select classifiers
classifiers = []
t0 = time.time()
print('Selecting classifiers..')
for _ in range(num_classifiers):
classification_errors = np.zeros(len(feature_indexes))
# normalize weights
weights *= 1. / np.sum(weights)
# select best classifier based on the weighted error
for f in range(len(feature_indexes)):
f_idx = feature_indexes[f]
# classifier error is the sum of image weights where the classifier
# is right
error = sum(map(lambda img_idx: weights[img_idx] if labels[img_idx] != votes[img_idx, f_idx] else 0, range(num_imgs)))
classification_errors[f] = error
# get best feature, i.e. with smallest error
min_error_idx = np.argmin(classification_errors)
best_error = classification_errors[min_error_idx]
best_feature_idx = feature_indexes[min_error_idx]
# set feature weight
best_feature = features[best_feature_idx]
feature_weight = 0.5 * np.log((1 - best_error) / best_error)
best_feature.weight = feature_weight
classifiers.append(best_feature)
# update image weights
weights = np.array(list(map(lambda img_idx: weights[img_idx] * np.sqrt((1-best_error)/best_error) if labels[img_idx] != votes[img_idx, best_feature_idx] else weights[img_idx] * np.sqrt(best_error/(1-best_error)), range(num_imgs))))
# remove feature (a feature can't be selected twice)
feature_indexes.remove(best_feature_idx)
print('\tTime needed for Selecting Classifiers:', time.time()-t0,'\n')
return classifiers
def _get_feature_vote(feature, image):
return feature.get_vote(image)
def _create_features(img_height, img_width, min_feature_width, max_feature_width, min_feature_height, max_feature_height):
print('Creating Haar-like features..')
t0 = time.time()
features = []
for feature in FeatureTypes:
# FeatureTypes are just tuples
feature_start_width = max(min_feature_width, feature[0])
for feature_width in range(feature_start_width, max_feature_width, feature[0]):
feature_start_height = max(min_feature_height, feature[1])
for feature_height in range(feature_start_height, max_feature_height, feature[1]):
# Iterate over possible x and y values.
for x in range(img_width - feature_width):
for y in range(img_height - feature_height):
# Two Haar features are created for every position. The first one has positive polarity whereas
# the second one has a negative one.
# self, feature_type, position, width, height, threshold, polarity
h1 = HaarLikeFeature(feature, (x, y), feature_width, feature_height, 0, 1)
h2 = HaarLikeFeature(feature, (x, y), feature_width, feature_height, 0, -1)
# The new features are added at the end of the feature list.
features.extend([h1, h2])
print('\t' + str(len(features)) + ' features created.')
print('\tTime needed for calculating Harr-like features:', time.time()-t0)
return features
# -
# **5)Use the learn method to learn a list of classifiers with the train data**
#
# With the `learn` function you can build a list of classifiers that detect whether an image contains a face or not.
#
# Use the following hyperparameters of the features and `num_classifiers`.
num_classifiers = 2
min_feature_height = 8
max_feature_height = 10
min_feature_width = 8
max_feature_width = 10
# ### Aprox. wall time: 4 minutes
# %%time
"""
faces_ii_training = load_images(pos_training_path)
faces_ii_testing = load_images(pos_testing_path)
non_faces_ii_training = load_images(neg_training_path)
non_faces_ii_testing = load_images(neg_testing_path)
"""
classifiers = learn(faces_ii_training, non_faces_ii_training, num_classifiers=num_classifiers,
min_feature_width=min_feature_width, max_feature_width=max_feature_width,
min_feature_height=min_feature_height, max_feature_height=max_feature_height)
# **6) Make a function for voting with different classifiers**
#
# Build two functions `ensemble_vote` and `ensemble_vote_all`.
#
# - `ensemble_vote(int_img, classifiers)` has to return a 1 if the majority of the votes of the classifiers is positive and a zero otherwise
#
# - `ensemble_vote_all(int_imgs, classifiers)` has to loop over the list `int_imgs` and compute the `ensemble_vote` for each image in the list. It has to return a list containing all the votes for all the images in `int_imgs`.
#
# Use the functions to compute the train and test acurracies for faces and non faces.
#
# Print the results in the following format:
# ```
# train results:
# Correctly identified Faces: 2129/2429 (87.64923836969946%)
# Correctly identified non-Faces: 4276/8548 (50.02339728591484%)
#
# test results:
# Correctly identified Faces: 300/472 (63.559322033898304%)
# Correctly identified non-Faces: 74/128 (57.8125%)
# ```
#
# It is not required to get this exact results but print the information in this format. It facilitates understanding the results.
# +
def ensemble_vote(int_img, classifiers):
# Recover all votes from all classifiers.
votes = np.sum([i.get_vote(int_img) for i in classifiers])
# True if most votes are positive, false elsewhere.
return votes >= 0
def ensemble_vote_all(int_imgs, classifiers, percent=False):
all_votes = [ensemble_vote(i, classifiers) for i in int_imgs]
tot, pos = len(all_votes), np.sum(all_votes)
if percent: return pos, tot, (pos/tot)*100
return pos, tot
# -
def print_results(pos_tr, neg_tr, pos_tst, neg_tst, classifiers):
# Print results in the specified format.
fpos_train, tn_train, fpcent_train = ensemble_vote_all(faces_ii_training, classifiers, percent=True)
nfpos_train, tnf_train, nfpcent_train = ensemble_vote_all(non_faces_ii_training, classifiers, percent=True)
fpos_test, tn_test, fpcent_test = ensemble_vote_all(faces_ii_testing, classifiers, percent=True)
nfpos_test, tnf_test, nfpcent_test = ensemble_vote_all(non_faces_ii_testing, classifiers, percent=True)
print("train results:")
print("Correctly identified Faces: {}/{} ({}%)".format(fpos_train, tn_train, fpcent_train))
print("Correctly identified non-Faces: {}/{} ({}%)\n".format(nfpos_train, tnf_train, nfpcent_train))
print("test results:")
print("Correctly identified Faces: {}/{} ({}%)".format(fpos_test, tn_test, fpcent_test))
print("Correctly identified non-Faces: {}/{} ({}%)\n".format(nfpos_test, tnf_test, nfpcent_test))
print_results(faces_ii_training, non_faces_ii_training, faces_ii_testing, non_faces_ii_training,
classifiers)
# **7) Make another test with 20 classifiers instead of 2**
#
# Inspect the classification results if you use adaboost with 20 classifiers. Use the same hyperameters for the features.
# Print the results as in the previous exercise:
#
# ```
# train results:
# Correctly identified Faces: 2256/2429 (92.87772745986003%)
# Correctly identified non-Faces: 7046/8548 (82.42863827795975%)
#
# test results:
# Correctly identified Faces: 285/472 (60.381355932203384%)
# Correctly identified non-Faces: 104/128 (81.25%)
# ```
#
# - Do the classification results improved in the train data?
#
#
num_classifiers = 20
min_feature_height = 8
max_feature_height = 10
min_feature_width = 8
max_feature_width = 10
# ### Aprox. wall time: 10 minutes
# %%time
"""
faces_ii_training = load_images(pos_training_path)
faces_ii_testing = load_images(pos_testing_path)
non_faces_ii_training = load_images(neg_training_path)
non_faces_ii_testing = load_images(neg_testing_path)
"""
classifiers20 = learn(faces_ii_training, non_faces_ii_training, num_classifiers=num_classifiers,
min_feature_width=min_feature_width, max_feature_width=max_feature_width,
min_feature_height=min_feature_height, max_feature_height=max_feature_height)
print_results(faces_ii_training, non_faces_ii_training, faces_ii_testing, non_faces_ii_training,
classifiers20)
# **8) Change the voting functions so that you can set a threshold for deciding a prediction**
#
# The threshold value indicates the minimum score for assigning a "positive" label (detect a face).
#
# Create the following functions
#
# - `ensemble_vote_t`: returns the final decision of a list of classifiers for a given threshold.
# - `ensemble_vote_all_t`: Iterates over a list of integral images and returns the final decision of a list of classifiers for each of the images (for a given threshold).
#
#
#
# compute the following:
#
# - a) number of correct faces over all faces (in the train data)
# - b) number of correct non faces over all non faces (in the train data)
# - c) number of correct faces over all faces (in the test data)
# - d) number of correct non faces over all non faces (in the test data)
#
# Using the list of 20 classifiers.
#
#
# The quantities have to be computed for each of the following thresholds:
#
# ```
# thresholds = np.array([x for x in range(-5,5,1)])/10.
# ```
#
# - Make a bar bar plot for a) b) c) and d). In the x axis write the threshold value.
#
# - What happens when you increase the threshold value ?
# +
def ensemble_vote_t(int_img, classifiers, threshold):
# Recover all votes from all classifiers.
votes = [i.get_vote(int_img) > threshold for i in classifiers]
# As before, the vote is based on the majority of votes
return np.sum(votes)/len(votes) > 0.5
def ensemble_vote_all_t(int_imgs, classifiers, threshold):
# Votes for all images.
return [ensemble_vote_t(i, classifiers, threshold) for i in int_imgs]
# -
thresholds = np.array([x for x in range(-5,5,1)])/10.
def plot_results(ftrain, nftrain, ftst, nftst, thresholds):
# Plot the results as specified on the exercise.
n_ftrain, n_nftrain, n_ftst, n_nftst = len(ftrain), len(nftrain), len(ftst), len(nftst)
print("Results for thresholds: {}\nLegend:".format(thresholds))
print("\ta) Correct faces over all faces - train")
print("\tb) Correct !faces over all non faces - train")
print("\tc) Correct faces over all faces - test")
print("\td) Correct !faces over all non faces - test")
# Correct faces over all faces - train
a = [np.sum(ftrain[i])/len(ftrain[i]) for i in range(n_nftrain)]
# Correct !faces over all non faces - train
b = [(len(nftrain[i]) - np.sum(nftrain[i]))/len(nftrain[i]) for i in range(n_nftrain)]
# Correct faces over all faces - test
c = [np.sum(ftst[i])/len(ftst[i]) for i in range(n_nftrain)]
# Correct !faces over all non faces - test
d = [(len(nftst[i]) - np.sum(nftst[i]))/len(nftst[i]) for i in range(n_nftrain)]
# Plot values
pa, = plt.plot(thresholds, a, label="a")
pb, = plt.plot(thresholds, b, label="b")
pc, = plt.plot(thresholds, c, label="c")
pd, = plt.plot(thresholds, d, label="d")
# Set title, label for x axis and its values and show the legend.
plt.title("Results")
plt.xlabel("Thresholds")
plt.xticks(thresholds)
plt.legend(handles=[pa, pb, pc, pd], fontsize='small', fancybox=True, loc='best')
plt.show()
correct_faces_train_t = [ensemble_vote_all_t(faces_ii_training, classifiers20, t) for t in thresholds]
correct_non_faces_train_t = [ensemble_vote_all_t(non_faces_ii_training, classifiers20, t) for t in thresholds]
correct_faces_test_t = [ensemble_vote_all_t(faces_ii_testing, classifiers20, t) for t in thresholds]
correct_non_faces_test_t = [ensemble_vote_all_t(non_faces_ii_testing, classifiers20, t) for t in thresholds]
plot_results(correct_faces_train_t, correct_non_faces_train_t, correct_faces_test_t,
correct_non_faces_test_t, thresholds)
# +
th = thresholds*20
correct_faces_train_t2 = [ensemble_vote_all_t(faces_ii_training, classifiers20, t) for t in th]
correct_non_faces_train_t2 = [ensemble_vote_all_t(non_faces_ii_training, classifiers20, t) for t in th]
correct_faces_test_t2 = [ensemble_vote_all_t(faces_ii_testing, classifiers20, t) for t in th]
correct_non_faces_test_t2 = [ensemble_vote_all_t(non_faces_ii_testing, classifiers20, t) for t in th]
plot_results(correct_faces_train_t2, correct_non_faces_train_t2, correct_faces_test_t2,
correct_non_faces_test_t2, th)
# -
# **Answer:** Augmenting the thresholds has resulted in outrageous and not representative results. To achieve better results (better accuracy), low thresholds.
| P6/P6_JoelBolwijn_RodrigoCabezas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import mmh3
import os
from bitarray import bitarray
import math
import socket
import sys
import pickle
from threading import Thread
## method to make a bloom filter , list_param = list of things,
## size = size of filter, no_of_hashes = no of hashes
def make_filter(list_param, size, no_of_hashes):
bit_array = bitarray(size)
bit_array.setall(0)
hash_num = 41
print "building own summary..."
for url in list_param:
for i in range(no_of_hashes):
bit = mmh3.hash(url, hash_num+i) % size
bit_array[bit] = 1
return bit_array
## method to check if thing is present in bloom filter , bit_array = bloom filter,
## size = size of filter(no of bits), no_of_hashes = no of hashes, url = object to check
def check_filter(bit_array, size, no_of_hashes, url):
bit_list = []
hash_num = 41
for i in range(int(no_of_hashes)):
bit_list.append(mmh3.hash(url, hash_num+i) % size)
for bit in bit_list:
if bit_array[int(bit)] == False:
return False
return True
## method to iterate check_filter over all the proxies
def check_filter_list(filter_dict, url):
## list all the proxies that might have the URl
proxy_true_list = []
if len(filter_dict) is 0:
print "empty summary cache..."
return proxy_true_list
print "checking in summary cache dictionary"
for key, value in filter_dict.items():
if check_filter(value[0], value[1], value[2], url):
print "found a hit in: ",key
proxy_true_list.append(key)
return proxy_true_list
##method to calculate size of a bloom filter and no of hashes required given the no of items and FP probability
def size_hash_calc(items,prob):
size = math.ceil((items * math.log(prob)) / math.log(1.0 / (pow(2.0, math.log(2.0)))))
hashes = round(math.log(2.0) * size / items)
print "size and no of hashes calculated"
return size, hashes
## method to fetch all the proxies avilable
## returns a list
def fetch_proxy_list():
file = open("proxy_list.txt", "r")
proxy_list = [line[:-1] for line in file]
file.close()
print "proxy list fetched from proxy_list.txt"
return proxy_list[:-1]
def send_filter_detail_worker(item,index_list_srl):
##connect to the server and send the index_list_srl
sock = socket.socket(socket.AF_INET, socket.SOCK_STREAM)
server_address = (item, 9000)
print ('connecting to %s port %s' % server_address)
try:
sock.connect(server_address)
sock.sendall(index_list_srl.encode('utf-8'))
sock.close()
print "sent own summary to: ",item
except socket.error as msg:
print "proxy offline: ",item
print "reason: ", msg
pass
sock.close()
## send an object to all remote proxies
def send_filter_details(bit_array, size, no_of_hashes):
## fetch list of proxies to send
proxy_list = fetch_proxy_list()
## make a list of filter,size,no_of_hashes
index_list = [bit_array, size, no_of_hashes]
## serialize the object into string using pickle
index_list_srl = pickle.dumps(index_list)
## Connect the socket to the port where the server is listeningusing threads
print "sending own summary to all the proxies online"
for item in proxy_list:
thread_send_filter_details_worker = Thread(target = send_filter_detail_worker, args = (item,index_list_srl))
thread_send_filter_details_worker.start()
| .ipynb_checkpoints/lib-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tpyes of Machine Learning Systems
# ## Instance-Based vs. Model-Based Learning
# model selection
#
# utility/fitness function
#
# cost function
#
# training the model: y = ax + b, find the best fiting a & b.
#
# A hyperparameter is a parameter of a learning algorithm (not of the model).
#
# Tuning hyperparameters is an important part of building a Machine Learning system.
| Chapter 1 - The Machine Learning Landcape.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import matplotlib.backends.backend_tkagg
import matplotlib.pylab as plt
from astropy.io import fits
from astropy import units as units
import astropy.io.fits as pyfits
from astropy.convolution import Gaussian1DKernel, convolve
from extinction import calzetti00, apply, ccm89
from scipy import optimize
import sys
import time
import emcee
import corner
from multiprocessing import Pool,cpu_count
import warnings
import glob, os
import math
warnings.filterwarnings('ignore')
# %matplotlib inline
# -
ncpu = cpu_count()
print("{0} CPUs".format(ncpu))
emcee.__version__
plt.tight_layout()
plt.rc('lines', linewidth=1, markersize=2)
plt.rc('font', size=12, family='serif')
plt.rc('mathtext', fontset='stix')
plt.rc('axes', linewidth=2)
plt.rc('xtick.major', width=1.5, size=4)
plt.rc('ytick.major', width=1.5, size=4)
plt.tick_params(axis='both', which='major', labelsize=18)
plt.tick_params(axis='both', which='minor', labelsize=18)
plt.subplots_adjust(bottom=0.2, left=0.2)
# + code_folding=[2, 14]
tik = time.clock()
df_cat=pd.read_csv('/Volumes/My Passport/uds_3dhst_v4.1.5_catalogs/uds_3dhst.v4.1.5.zbest.rf', delim_whitespace=True,header=None,comment='#',index_col=False)
df_cat.columns=["id", "z_best", "z_type", "z_spec", "DM", "L153", "nfilt153", "L154","nfilt154", "L155", "nfilt155", "L161", "nfilt161", "L162", "nfilt162", \
"L163", "nfilt163", "L156", "nfilt156", "L157", "nfilt157", "L158", "nfilt158", "L159", "nfilt159", "L160", "nfilt160", "L135", "nfilt135", "L136", "nfilt136",\
"L137", "nfilt137", "L138", "nfilt138", "L139", "nfilt139", "L270", "nfilt270", "L271", "nfilt271", "L272", "nfilt272", "L273", "nfilt273", "L274", "nfilt274", "L275", "nfilt275"]
# df = pd.read_csv('/Volumes/My Passport/GV_CMD_fn_table_20180904/matching_galaxies_uds_20180823_GV.csv', sep=',')
# df = pd.read_csv('/Volumes/My Passport/TPAGB/database/matching_galaxies_uds_20200206_PSB.csv', sep=',')
df = pd.read_csv('/Volumes/My Passport/TPAGB/database/matching_galaxies_uds_20200301_PSB.csv', sep=',')
df = pd.read_csv('/Volumes/My Passport/TPAGB/database/matching_galaxies_uds_20200303_PSB.csv', sep=',')
df.columns=['detector','ID','region','filename','chip']
df_photometry=pd.read_csv('/Volumes/My Passport/uds_3dhst.v4.2.cats/Catalog/uds_3dhst.v4.2.cat', delim_whitespace=True,header=None,comment='#',index_col=False)
df_photometry.columns=["id", "x", "y", "ra", "dec", "faper_F160W", "eaper_F160W","faper_F140W", "eaper_F140W", "f_F160W", "e_F160W", "w_F160W", \
"f_u", "e_u", "w_u","f_B", "e_B", "w_B","f_V", "e_V", "w_V", "f_F606W", "e_F606W","w_F606W",\
"f_R", "e_R", "w_R", "f_i", "e_i", "w_i", "f_F814W", "e_F814W", "w_F814W", "f_z", "e_z", "w_z",\
"f_F125W", "e_F125W", "w_F125W","f_J", "e_J", "w_J", "f_F140W", "e_F140W", "w_F140W",\
"f_H", "e_H", "w_H","f_K", "e_K", "w_K", "f_IRAC1", "e_IRAC1", "w_IRAC1", "f_IRAC2", "e_IRAC2", "w_IRAC2",\
"f_IRAC3", "e_IRAC3", "w_IRAC3", "f_IRAC4", "e_IRAC4", "w_IRAC4","tot_cor", "wmin_ground", "wmin_hst","wmin_wfc3",\
"wmin_irac", "z_spec", "star_flag", "kron_radius", "a_image", "b_image", "theta_J2000", "class_star", "flux_radius", "fwhm_image",\
"flags", "IRAC1_contam", "IRAC2_contam", "IRAC3_contam", "IRAC4_contam", "contam_flag","f140w_flag", "use_phot", "near_star", "nexp_f125w", "nexp_f140w", "nexp_f160w"]
df_fast = pd.read_csv('/Volumes/My Passport/uds_3dhst.v4.2.cats/Fast/uds_3dhst.v4.2.fout', delim_whitespace=True,header=None,comment='#',index_col=False)
df_fast.columns = ['id', 'z', 'ltau', 'metal','lage','Av','lmass','lsfr','lssfr','la2t','chi2']
tok = time.clock()
print('Time to read the catalogues:'+str(tok-tik))
df_zfit = pd.read_csv('/Volumes/My Passport/uds_3dhst_v4.1.5_catalogs/uds_3dhst.v4.1.5.zfit.concat.dat',delim_whitespace=True,header=None,comment='#',index_col=False)
df_zfit.columns=['phot_id','grism_id','jh_mag','z_spec','z_peak_phot','z_phot_l95',\
'z_phot_l68','z_phot_u68','z_phot_u95','z_max_grism','z_peak_grism',\
'l95','l68','u68','u95','f_cover','f_flagged','max_contam','int_contam',\
'f_negative','flag1','flag2']
# + code_folding=[0]
# ### Ma05
tik2 = time.clock()
norm_wavelength= 5500.0
df_Ma = pd.read_csv('/Volumes/My Passport/M09_ssp_pickles.sed', delim_whitespace=True, header=None, comment='#', index_col=False)# only solar metallicity is contained in this catalogue
df_Ma.columns = ['Age','ZH','l','Flambda']
age = df_Ma.Age
metallicity = df_Ma.ZH
wavelength = df_Ma.l
Flux = df_Ma.Flambda
age_1Gyr_index = np.where(age==1.0)[0]
age_1Gyr = age[age_1Gyr_index]
metallicity_1Gyr = metallicity[age_1Gyr_index]
wavelength_1Gyr = wavelength[age_1Gyr_index]
Flux_1Gyr = Flux[age_1Gyr_index]
F_5500_1Gyr_index=np.where(wavelength_1Gyr==norm_wavelength)[0]
F_5500_1Gyr = Flux_1Gyr[wavelength_1Gyr==norm_wavelength].values # this is the band to be normalized
df_M13 = pd.read_csv('/Volumes/My Passport/M13_models/sed_M13.ssz002',delim_whitespace=True,header=None,comment='#',index_col=False)
df_M13.columns = ['Age','ZH','l','Flambda']
age_M13 = df_M13.Age
metallicity_M13 = df_M13.ZH
wavelength_M13 = df_M13.l
Flux_M13 = df_M13.Flambda
age_1Gyr_index_M13 = np.where(age_M13==1.0)[0]#[0]
age_1Gyr_M13 = age_M13[age_1Gyr_index_M13]
metallicity_1Gyr_M13 = metallicity_M13[age_1Gyr_index_M13]
wavelength_1Gyr_M13 = wavelength_M13[age_1Gyr_index_M13]
Flux_1Gyr_M13 = Flux_M13[age_1Gyr_index_M13]
F_5500_1Gyr_index_M13=np.where(abs(wavelength_1Gyr_M13-norm_wavelength)<15)[0]
F_5500_1Gyr_M13 = 0.5*(Flux_1Gyr_M13.loc[62271+F_5500_1Gyr_index_M13[0]]+Flux_1Gyr_M13.loc[62271+F_5500_1Gyr_index_M13[1]])
# ### BC03
df_BC = pd.read_csv('/Volumes/My Passport/ssp_900Myr_z02.spec',delim_whitespace=True,header=None,comment='#',index_col=False)
df_BC.columns=['Lambda','Flux']
wavelength_BC = df_BC.Lambda
Flux_BC = df_BC.Flux
F_5500_BC_index=np.where(wavelength_BC==norm_wavelength)[0]
Flux_BC_norm = Flux_BC[F_5500_BC_index]
### Read in the BC03 models High-resolution, with Stelib library, Salpeter IMF, solar metallicity
BC03_fn='/Volumes/My Passport/bc03/models/Stelib_Atlas/Salpeter_IMF/bc2003_hr_stelib_m62_salp_ssp.ised_ASCII'
BC03_file = open(BC03_fn,"r")
BC03_X = []
for line in BC03_file:
BC03_X.append(line)
BC03_SSP_m62 = np.array(BC03_X)
BC03_age_list = np.array(BC03_SSP_m62[0].split()[1:])
BC03_age_list_num = BC03_age_list.astype(np.float)/1.0e9 # unit is Gyr
BC03_wave_list = np.array(BC03_SSP_m62[6].split()[1:])
BC03_wave_list_num = BC03_wave_list.astype(np.float)
BC03_flux_list = np.array(BC03_SSP_m62[7:-12])
BC03_flux_array = np.zeros((221,7178))
for i in range(221):
BC03_flux_array[i,:] = BC03_flux_list[i].split()[1:]
BC03_flux_array[i,:] = BC03_flux_array[i,:]/BC03_flux_array[i,2556]# Normalize the flux
# + code_folding=[0]
## Prepare the M05 models and store in the right place
M05_model = []
M05_model_list=[]
for i in range(30):
age_index = i
age_prior = df_Ma.Age.unique()[age_index]
galaxy_age_string = str(age_prior)
split_galaxy_age_string = str(galaxy_age_string).split('.')
fn1 = '/Volumes/My Passport/SSP_models/new/M05_age_'+'0_'+split_galaxy_age_string[1]+'_Av_00_z002.csv'
M05_model = np.loadtxt(fn1)
M05_model_list.append(M05_model)
fn1 = '/Volumes/My Passport/SSP_models/new/M05_age_1_Av_00_z002.csv'
fn2 = '/Volumes/My Passport/SSP_models/new/M05_age_1_5_Av_00_z002.csv'
M05_model = np.loadtxt(fn1)
M05_model_list.append(M05_model)
M05_model = np.loadtxt(fn2)
M05_model_list.append(M05_model)
for i in range(32,46):
age_index = i
age_prior = df_Ma.Age.unique()[age_index]
galaxy_age_string = str(age_prior)
split_galaxy_age_string = str(galaxy_age_string).split('.')
fn2 = '/Volumes/My Passport/SSP_models/new/M05_age_'+split_galaxy_age_string[0]+'_Av_00_z002.csv'
M05_model = np.loadtxt(fn2)
M05_model_list.append(M05_model)
## Prepare the M13 models and store in the right place
M13_model = []
M13_model_list=[]
fn1 = '/Volumes/My Passport/SSP_models/new/M13_age_1e-06_Av_00_z002.csv'
fn2 = '/Volumes/My Passport/SSP_models/new/M13_age_0_0001_Av_00_z002.csv'
M13_model = np.genfromtxt(fn1)
M13_model_list.append(M13_model)
M13_model = np.genfromtxt(fn2)
M13_model_list.append(M13_model)
for i in range(2,51):
age_index = i
age_prior = df_M13.Age.unique()[age_index]
galaxy_age_string = str(age_prior)
split_galaxy_age_string = str(galaxy_age_string).split('.')
fn1 = '/Volumes/My Passport/SSP_models/new/M13_age_'+'0_'+split_galaxy_age_string[1]+'_Av_00_z002.csv'
M13_model = np.loadtxt(fn1)
M13_model_list.append(M13_model)
fn1 = '/Volumes/My Passport/SSP_models/new/M13_age_1_Av_00_z002.csv'
fn2 = '/Volumes/My Passport/SSP_models/new/M13_age_1_5_Av_00_z002.csv'
M13_model = np.loadtxt(fn1)
M13_model_list.append(M13_model)
M13_model = np.loadtxt(fn2)
M13_model_list.append(M13_model)
for i in range(53,67):
age_index = i
age_prior = df_M13.Age.unique()[age_index]
galaxy_age_string = str(age_prior)
split_galaxy_age_string = str(galaxy_age_string).split('.')
fn2 = '/Volumes/My Passport/SSP_models/new/M13_age_'+split_galaxy_age_string[0]+'_Av_00_z002.csv'
M13_model = np.loadtxt(fn2)
M13_model_list.append(M13_model)
# + code_folding=[0, 19, 53, 61, 169, 177, 277, 378, 478, 578, 688, 799, 899, 998, 1039, 1086, 1129, 1171, 1180, 1182, 1192]
def read_spectra(row):
"""
region: default 1 means the first region mentioned in the area, otherwise, the second region/third region
"""
detector=df.detector[row]
region = df.region[row]
chip = df.chip[row]
ID = df.ID[row]
redshift_1=df_cat.loc[ID-1].z_best
mag = -2.5*np.log10(df_cat.loc[ID-1].L161)+25#+0.02
#print mag
#WFC3 is using the infrared low-resolution grism, and here we are using the z band
if detector == 'WFC3':
filename="/Volumes/My Passport/UDS_WFC3_V4.1.5/uds-"+"{0:02d}".format(region)+"/1D/ASCII/uds-"+"{0:02d}".format(region)+"-G141_"+"{0:05d}".format(ID)+".1D.ascii"
OneD_1 = np.loadtxt(filename,skiprows=1)
if detector =="ACS":
filename="/Volumes/My Passport/UDS_ACS_V4.1.5/acs-uds-"+"{0:02d}".format(region)+"/1D/FITS/"+df.filename[row]
OneD_1 = fits.getdata(filename, ext=1)
return ID, OneD_1,redshift_1, mag
def Lick_index_ratio(wave, flux, band=3):
if band == 3:
blue_min = 1.06e4 # 1.072e4#
blue_max = 1.08e4 # 1.08e4#
red_min = 1.12e4 # 1.097e4#
red_max = 1.14e4 # 1.106e4#
band_min = blue_max
band_max = red_min
# Blue
blue_mask = (wave >= blue_min) & (wave <= blue_max)
blue_wave = wave[blue_mask]
blue_flux = flux[blue_mask]
# Red
red_mask = (wave >= red_min) & (wave <= red_max)
red_wave = wave[red_mask]
red_flux = flux[red_mask]
band_mask = (wave >= band_min) & (wave <= band_max)
band_wave = wave[band_mask]
band_flux = flux[band_mask]
if len(blue_wave) == len(red_wave) and len(blue_wave) != 0:
ratio = np.mean(blue_flux) / np.mean(red_flux)
elif red_wave == []:
ratio = np.mean(blue_flux) / np.mean(red_flux)
elif len(blue_wave) != 0 and len(red_wave) != 0:
ratio = np.mean(blue_flux) / np.mean(red_flux)
# ratio_err = np.sqrt(np.sum(1/red_flux**2*blue_flux_err**2)+np.sum((blue_flux/red_flux**2*red_flux_err)**2))
return ratio # , ratio_err
def binning_spec_keep_shape(wave,flux,bin_size):
wave_binned = wave
flux_binned = np.zeros(len(wave))
# flux_err_binned = np.zeros(len(wave))
for i in range((int(len(wave)/bin_size))):
flux_binned[bin_size*i:bin_size*(i+1)] = np.mean(flux[bin_size*i:bin_size*(i+1)])
#flux_err_binned[bin_size*i:bin_size*(i+1)] = np.mean(flux_err[bin_size*i:bin_size*(i+1)])
return wave_binned, flux_binned#, flux_err_binned
def derive_1D_spectra_Av_corrected(OneD_1,redshift_1,rownumber,wave_list,band_list,photometric_flux,photometric_flux_err,photometric_flux_err_mod,A_v):
"""
OneD_1 is the oneD spectra
redshift_1 is the redshift of the spectra
rownumber is the row number in order to store the spectra
"""
region = df.region[rownumber]
ID = df.ID[rownumber]
n = len(OneD_1)
age=10**(df_fast.loc[ID-1].lage)/1e9 ## in Gyr
metal = df_fast.loc[ID-1].metal
sfr = 10**(df_fast.loc[ID-1].lsfr)
intrinsic_Av = df_fast.loc[ID-1].Av
norm_factor_BC = int((OneD_1[int(n/2+1)][0]-OneD_1[int(n/2)][0])/(1+redshift_1)/1)
norm_limit_BC = int(5930/norm_factor_BC)*norm_factor_BC+400
smooth_wavelength_BC_1 = wavelength_BC[400:norm_limit_BC].values.reshape(-1,norm_factor_BC).mean(axis=1)
smooth_wavelength_BC = np.hstack([smooth_wavelength_BC_1,wavelength_BC[norm_limit_BC:]])
smooth_Flux_BC_1 = Flux_BC[400:norm_limit_BC].values.reshape(-1,norm_factor_BC).mean(axis=1)
smooth_Flux_BC = np.hstack([smooth_Flux_BC_1,Flux_BC[norm_limit_BC:]])/Flux_BC_norm.values[0]
norm_factor_Ma = int((OneD_1[int(n/2+1)][0]-OneD_1[int(n/2)][0])/(1+redshift_1)/5)
norm_limit_Ma = int(4770/norm_factor_Ma)*norm_factor_Ma
smooth_wavelength_Ma = wavelength_1Gyr[:norm_limit_Ma].values.reshape(-1,norm_factor_Ma).mean(axis=1)
smooth_Flux_Ma_1Gyr = Flux_1Gyr[:norm_limit_Ma].values.reshape(-1,norm_factor_Ma).mean(axis=1)/F_5500_1Gyr
if redshift_1<=0.05:
i = 2
temp_norm_wave = wave_list[i]/(1+redshift_1)
index_wave_norm = find_nearest(smooth_wavelength_BC,temp_norm_wave)
norm_band = photometric_flux[i]
#plt.text(5000,0.55,'normalized at V: rest frame '+"{0:.2f}".format(temp_norm_wave),fontsize=16)
#plt.axvline(temp_norm_wave,linewidth=2,color='b')
elif redshift_1<=0.14:
i = 6
temp_norm_wave = wave_list[i]/(1+redshift_1)
index_wave_norm = find_nearest(smooth_wavelength_BC,temp_norm_wave)
norm_band = photometric_flux[i]
#plt.text(5000,0.55,'normalized at F606W: rest frame '+"{0:.2f}".format(temp_norm_wave),fontsize=16)
#plt.axvline(temp_norm_wave,linewidth=2,color='b')
elif redshift_1<=0.26:
i = 3
temp_norm_wave = wave_list[i]/(1+redshift_1)
index_wave_norm = find_nearest(smooth_wavelength_BC,temp_norm_wave)
norm_band = photometric_flux[i]
#plt.text(5000,0.55,'normalized at R: rest frame '+"{0:.2f}".format(temp_norm_wave),fontsize=16)
#plt.axvline(temp_norm_wave,linewidth=2,color='b')
elif redshift_1<=0.42:
i = 4
temp_norm_wave = wave_list[i]/(1+redshift_1)
index_wave_norm = find_nearest(smooth_wavelength_BC,temp_norm_wave)
norm_band = photometric_flux[i]
#plt.text(5000,0.55,'normalized at i: rest frame '+"{0:.2f}".format(temp_norm_wave),fontsize=16)
#plt.axvline(temp_norm_wave,linewidth=2,color='b')
elif redshift_1<=0.54:
i = 7
temp_norm_wave = wave_list[i]/(1+redshift_1)
index_wave_norm = find_nearest(smooth_wavelength_BC,temp_norm_wave)
norm_band = photometric_flux[i]
#plt.text(5000,0.55,'normalized at F814W: rest frame '+"{0:.2f}".format(temp_norm_wave),fontsize=16)
#plt.axvline(temp_norm_wave,linewidth=2,color='b')
else:
i = 5
temp_norm_wave = wave_list[i]/(1+redshift_1)
index_wave_norm = find_nearest(smooth_wavelength_BC,temp_norm_wave)
norm_band = photometric_flux[i]
#plt.text(5000,0.55,'normalized at z: rest frame '+"{0:.2f}".format(temp_norm_wave),fontsize=16)
#plt.axvline(temp_norm_wave,linewidth=2,color='b')
x = np.zeros(n)
y = np.zeros(n)
y_err = np.zeros(n)
sensitivity = np.zeros(n)
for i in range(0,n):
x[i] = OneD_1[i][0]#/(1+redshift_1)
print('wavelength range:',x[0],x[-1])
spectra_extinction = calzetti00(x, A_v, 4.05)
for i in range(n):
spectra_flux_correction = 10**(0.4*spectra_extinction[i])# from obs to obtain the true value: the absolute value
x[i] = x[i]/(1+redshift_1)
y[i] = (OneD_1[i][1]-OneD_1[i][3])/OneD_1[i][6]*spectra_flux_correction#/Flux_0 # (flux-contamination)/sensitivity
y_err[i] = OneD_1[i][2]/OneD_1[i][6]*spectra_flux_correction#/Flux_0
sensitivity[i] = OneD_1[i][6]
# end_index = np.argmin(np.diff(sensitivity[263:282],2)[1:],0)+263
# start_index = np.argmin(np.diff(sensitivity[40:50],2)[1:])+42
start_index = np.argmin(abs(x*(1+redshift_1)-11407.53))
end_index = np.argmin(abs(x*(1+redshift_1)-16428.61))
print('masking region:',x[start_index]*(1+redshift_1),x[end_index]*(1+redshift_1),start_index,end_index)
# plt.plot(x*(1+redshift_1),sensitivity,color='k')
# plt.plot(x[start_index:end_index]*(1+redshift_1),sensitivity[start_index:end_index],color='red')
print('before masking',len(x))
x = x[start_index:end_index]#[int(n*2/10):int(n*8/10)]
y = y[start_index:end_index]*1e-17/norm_band#[int(n*2/10):int(n*8/10)]*1e-17/norm_band
y_err = y_err[start_index:end_index]*1e-17/norm_band#[int(n*2/10):int(n*8/10)]*1e-17/norm_band
print('after masking',len(x))
# mask_non_neg_photo = np.where(photometric_flux>0)
# wave_list = wave_list[mask_non_neg_photo]
# band_list = band_list[mask_non_neg_photo]
# photometric_flux = photometric_flux[mask_non_neg_photo]
# photometric_flux_err_mod = photometric_flux_err_mod[mask_non_neg_photo]
return x, y, y_err, wave_list/(1+redshift_1), band_list/(1+redshift_1), photometric_flux/norm_band, photometric_flux_err/norm_band, photometric_flux_err_mod/norm_band
def binning_spec_keep_shape_x(wave,flux,flux_err,bin_size):
wave_binned = wave
flux_binned = np.zeros(len(wave))
flux_err_binned = np.zeros(len(wave))
for i in range((int(len(wave)/bin_size))+1):
flux_binned[bin_size*i:bin_size*(i+1)] = np.mean(flux[bin_size*i:bin_size*(i+1)])
flux_err_binned[bin_size*i:bin_size*(i+1)] = np.mean(flux_err[bin_size*i:bin_size*(i+1)])
return wave_binned, flux_binned, flux_err_binned
def minimize_age_AV_vector_weighted(X):
galaxy_age= X[0]
intrinsic_Av = X[1]
# print('minimize process age av grid',X)
n=len(x)
age_index = find_nearest(df_Ma.Age.unique(), galaxy_age)
age_prior = df_Ma.Age.unique()[age_index]
AV_string = str(intrinsic_Av)
galaxy_age_string = str(age_prior)
split_galaxy_age_string = str(galaxy_age_string).split('.')
# print(age_prior)
if age_prior < 1:
if galaxy_age < age_prior:
model1 = (M05_model_list[age_index]*(galaxy_age-df_Ma.Age.unique()[age_index-1]) \
+ M05_model_list[age_index-1]*(age_prior-galaxy_age))/(df_Ma.Age.unique()[age_index]-df_Ma.Age.unique()[age_index-1])
elif galaxy_age > age_prior:
model1 = (M05_model_list[age_index]*(df_Ma.Age.unique()[age_index+1]-galaxy_age) \
+ M05_model_list[age_index+1]*(galaxy_age-age_prior))/(df_Ma.Age.unique()[age_index+1]-df_Ma.Age.unique()[age_index])
elif galaxy_age == age_prior:
model1 = M05_model_list[age_index]
elif age_prior == 1.5:
if galaxy_age >=1.25 and galaxy_age <1.5:
model1 = 2.*(1.5-galaxy_age)*M05_model_list[30] + 2.*(galaxy_age-1.0)*M05_model_list[31]
elif galaxy_age >= 1.5 and galaxy_age <= 1.75:
model1 = 2.*(2.0-galaxy_age)*M05_model_list[31] + 2.*(galaxy_age-1.5)*M05_model_list[32]
elif len(split_galaxy_age_string[1])==1:
if galaxy_age >= 1.0 and galaxy_age < 1.25:
model1 = 2.*(1.5-galaxy_age)*M05_model_list[30] + 2.*(galaxy_age-1.0)*M05_model_list[31]
elif galaxy_age >=1.75 and galaxy_age < 2.0:
model1 = 2.*(2.0-galaxy_age)*M05_model_list[31] + 2.*(galaxy_age-1.5)*M05_model_list[32]
elif galaxy_age >= 2.0 and galaxy_age < 3.0:
model1 = (3.0-galaxy_age)*M05_model_list[32] + (galaxy_age-2.0)*M05_model_list[33]
elif galaxy_age >= 3.0 and galaxy_age < 4.0:
model1 = (4.0-galaxy_age)*M05_model_list[33] + (galaxy_age-3.0)*M05_model_list[34]
elif galaxy_age >= 4.0 and galaxy_age < 5.0:
model1 = (5.0-galaxy_age)*M05_model_list[34] + (galaxy_age-4.0)*M05_model_list[35]
elif galaxy_age >= 5.0 and galaxy_age < 6.0:
model1 = (6.0-galaxy_age)*M05_model_list[35] + (galaxy_age-5.0)*M05_model_list[36]
elif galaxy_age >= 6.0 and galaxy_age < 7.0:
model1 = (7.0-galaxy_age)*M05_model_list[36] + (galaxy_age-6.0)*M05_model_list[37]
elif galaxy_age >= 7.0 and galaxy_age < 8.0:
model1 = (8.0-galaxy_age)*M05_model_list[37] + (galaxy_age-7.0)*M05_model_list[38]
elif galaxy_age >= 8.0 and galaxy_age < 9.0:
model1 = (9.0-galaxy_age)*M05_model_list[38] + (galaxy_age-8.0)*M05_model_list[39]
elif galaxy_age >= 9.0 and galaxy_age < 10.0:
model1 = (10.0-galaxy_age)*M05_model_list[39] + (galaxy_age-9.0)*M05_model_list[40]
elif galaxy_age >= 10.0 and galaxy_age < 11.0:
model1 = (11.0-galaxy_age)*M05_model_list[40] + (galaxy_age-10.0)*M05_model_list[41]
elif galaxy_age >= 11.0 and galaxy_age < 12.0:
model1 = (12.0-galaxy_age)*M05_model_list[41] + (galaxy_age-11.0)*M05_model_list[42]
elif galaxy_age >= 12.0 and galaxy_age < 13.0:
model1 = (13.0-galaxy_age)*M05_model_list[42] + (galaxy_age-12.0)*M05_model_list[43]
elif galaxy_age >= 13.0 and galaxy_age < 14.0:
model1 = (14.0-galaxy_age)*M05_model_list[43] + (galaxy_age-13.0)*M05_model_list[44]
elif galaxy_age >= 14.0 and galaxy_age < 15.0:
model1 = (15.0-galaxy_age)*M05_model_list[44] + (galaxy_age-14.0)*M05_model_list[45]
else:
model1 = M05_model_list[age_index]
spectra_extinction = calzetti00(model1[0,:], intrinsic_Av, 4.05)
spectra_flux_correction = 10**(-0.4*spectra_extinction)
M05_flux_center = model1[1,:]*spectra_flux_correction
F_M05_index=700#167
Flux_M05_norm_new = M05_flux_center[F_M05_index]
smooth_Flux_Ma_1Gyr_new = M05_flux_center/Flux_M05_norm_new
binning_index = find_nearest(model1[0,:],np.median(x))
if binning_index == 0:
binning_index = 1
elif binning_index ==len(x):
binning_index = len(x)-1
if (x[int(n/2)]-x[int(n/2)-1]) > (model1[0,binning_index]-model1[0,binning_index-1]):
binning_size = int((x[int(n/2)]-x[int(n/2)-1])/(model1[0,binning_index]-model1[0,binning_index-1]))
model_wave_binned,model_flux_binned = binning_spec_keep_shape(model1[0,:], smooth_Flux_Ma_1Gyr_new,binning_size)
x2 = reduced_chi_square(x, y, y_err, model_wave_binned, model_flux_binned)
if np.isnan(x2):
print('spectra chi2 is nan,binning model',model_flux_binned)
print('spectra model wave', model1[0,:], model1[1,:], intrinsic_Av)
print('model flux before binning', spectra_extinction, spectra_flux_correction, M05_flux_center, Flux_M05_norm_new)
x2_photo = chisquare_photo(model_wave_binned, model_flux_binned, redshift_1, wave_list, band_list, photometric_flux, photometric_flux_err, photometric_flux_err_mod)
# print('binning model, model 1', n, (model1[0,binning_index]-model1[0,binning_index-1]), (x[int(n/2)]-x[int(n/2)-1]), binning_size)
else:
binning_size = int((model1[0,binning_index]-model1[0,binning_index-1])/(x[int(n/2)]-x[int(n/2)-1]))
x_binned,y_binned,y_err_binned=binning_spec_keep_shape_x(x,y,y_err,binning_size)
x2 = reduced_chi_square(x_binned, y_binned, y_err_binned, model1[0,:], smooth_Flux_Ma_1Gyr_new)
# print('binning data, model 1', n, (model1[0,binning_index]-model1[0,binning_index-1]), (x[int(n/2)]-x[int(n/2)-1]), binning_size)
x2_photo = chisquare_photo(model1[0,:], smooth_Flux_Ma_1Gyr_new, redshift_1, wave_list, band_list, photometric_flux, photometric_flux_err, photometric_flux_err_mod)
try:
if 0.01<galaxy_age<13 and 0.0<=intrinsic_Av<=4.0 and not np.isinf(0.5*x2+0.5*x2_photo):
x2_tot = 0.5*weight1*x2+0.5*weight2*x2_photo
else:
x2_tot = np.inf
except ValueError: # NaN value case
x2_tot = np.inf
print('ValueError', x2_tot)
# print('M05 x2 tot:',x2, x2_photo, x2_tot)
return x2_tot
def lg_minimize_age_AV_vector_weighted(X):
galaxy_age= X[0]
intrinsic_Av = X[1]
n=len(x)
age_index = find_nearest(df_Ma.Age.unique(), galaxy_age)
age_prior = df_Ma.Age.unique()[age_index]
AV_string = str(intrinsic_Av)
galaxy_age_string = str(age_prior)
split_galaxy_age_string = str(galaxy_age_string).split('.')
if age_prior < 1:
if galaxy_age < age_prior:
model1 = (M05_model_list[age_index]*(galaxy_age-df_Ma.Age.unique()[age_index-1]) \
+ M05_model_list[age_index-1]*(age_prior-galaxy_age))/(df_Ma.Age.unique()[age_index]-df_Ma.Age.unique()[age_index-1])
elif galaxy_age > age_prior:
model1 = (M05_model_list[age_index]*(df_Ma.Age.unique()[age_index+1]-galaxy_age) \
+ M05_model_list[age_index+1]*(galaxy_age-age_prior))/(df_Ma.Age.unique()[age_index+1]-df_Ma.Age.unique()[age_index])
elif galaxy_age == age_prior:
model1 = M05_model_list[age_index]
elif age_prior == 1.5:
if galaxy_age >=1.25 and galaxy_age <1.5:
model1 = 2.*(1.5-galaxy_age)*M05_model_list[30] + 2.*(galaxy_age-1.0)*M05_model_list[31]
elif galaxy_age >= 1.5 and galaxy_age <= 1.75:
model1 = 2.*(2.0-galaxy_age)*M05_model_list[31] + 2.*(galaxy_age-1.5)*M05_model_list[32]
elif len(split_galaxy_age_string[1])==1:
if galaxy_age >= 1.0 and galaxy_age < 1.25:
model1 = 2.*(1.5-galaxy_age)*M05_model_list[30] + 2.*(galaxy_age-1.0)*M05_model_list[31]
elif galaxy_age >=1.75 and galaxy_age < 2.0:
model1 = 2.*(2.0-galaxy_age)*M05_model_list[31] + 2.*(galaxy_age-1.5)*M05_model_list[32]
elif galaxy_age >= 2.0 and galaxy_age < 3.0:
model1 = (3.0-galaxy_age)*M05_model_list[32] + (galaxy_age-2.0)*M05_model_list[33]
elif galaxy_age >= 3.0 and galaxy_age < 4.0:
model1 = (4.0-galaxy_age)*M05_model_list[33] + (galaxy_age-3.0)*M05_model_list[34]
elif galaxy_age >= 4.0 and galaxy_age < 5.0:
model1 = (5.0-galaxy_age)*M05_model_list[34] + (galaxy_age-4.0)*M05_model_list[35]
elif galaxy_age >= 5.0 and galaxy_age < 6.0:
model1 = (6.0-galaxy_age)*M05_model_list[35] + (galaxy_age-5.0)*M05_model_list[36]
elif galaxy_age >= 6.0 and galaxy_age < 7.0:
model1 = (7.0-galaxy_age)*M05_model_list[36] + (galaxy_age-6.0)*M05_model_list[37]
elif galaxy_age >= 7.0 and galaxy_age < 8.0:
model1 = (8.0-galaxy_age)*M05_model_list[37] + (galaxy_age-7.0)*M05_model_list[38]
elif galaxy_age >= 8.0 and galaxy_age < 9.0:
model1 = (9.0-galaxy_age)*M05_model_list[38] + (galaxy_age-8.0)*M05_model_list[39]
elif galaxy_age >= 9.0 and galaxy_age < 10.0:
model1 = (10.0-galaxy_age)*M05_model_list[39] + (galaxy_age-9.0)*M05_model_list[40]
elif galaxy_age >= 10.0 and galaxy_age < 11.0:
model1 = (11.0-galaxy_age)*M05_model_list[40] + (galaxy_age-10.0)*M05_model_list[41]
elif galaxy_age >= 11.0 and galaxy_age < 12.0:
model1 = (12.0-galaxy_age)*M05_model_list[41] + (galaxy_age-11.0)*M05_model_list[42]
elif galaxy_age >= 12.0 and galaxy_age < 13.0:
model1 = (13.0-galaxy_age)*M05_model_list[42] + (galaxy_age-12.0)*M05_model_list[43]
elif galaxy_age >= 13.0 and galaxy_age < 14.0:
model1 = (14.0-galaxy_age)*M05_model_list[43] + (galaxy_age-13.0)*M05_model_list[44]
elif galaxy_age >= 14.0 and galaxy_age < 15.0:
model1 = (15.0-galaxy_age)*M05_model_list[44] + (galaxy_age-14.0)*M05_model_list[45]
else:
model1 = M05_model_list[age_index]
spectra_extinction = calzetti00(model1[0,:], intrinsic_Av, 4.05)
spectra_flux_correction = 10**(-0.4*spectra_extinction)
M05_flux_center = model1[1,:]*spectra_flux_correction
F_M05_index=700#167
Flux_M05_norm_new = M05_flux_center[F_M05_index]
smooth_Flux_Ma_1Gyr_new = M05_flux_center/Flux_M05_norm_new
binning_index = find_nearest(model1[0,:],np.median(x))
if binning_index == 0:
binning_index = 1
elif binning_index ==len(x):
binning_index = len(x)-1
if (x[int(n/2)]-x[int(n/2)-1]) > (model1[0,binning_index]-model1[0,binning_index-1]):
binning_size = int((x[int(n/2)]-x[int(n/2)-1])/(model1[0,binning_index]-model1[0,binning_index-1]))
model_wave_binned,model_flux_binned = binning_spec_keep_shape(model1[0,:], smooth_Flux_Ma_1Gyr_new,binning_size)
x2 = reduced_chi_square(x, y, y_err, model_wave_binned, model_flux_binned)
# x2_photo = chisquare_photo(model_wave_binned, model_flux_binned, redshift_1,wave_list, band_list, photometric_flux, photometric_flux_err, photometric_flux_err_mod)
x2_photo = chisquare_photo(model_wave_binned, model_flux_binned, redshift_1,wave_list, band_list, photometric_flux, photometric_flux_err, photometric_flux_err_mod)
# print('binning model, model 1', n, (model1[0,binning_index]-model1[0,binning_index-1]), (x[int(n/2)]-x[int(n/2)-1]), binning_size)
else:
binning_size = int((model1[0,binning_index]-model1[0,binning_index-1])/(x[int(n/2)]-x[int(n/2)-1]))
x_binned,y_binned,y_err_binned=binning_spec_keep_shape_x(x,y,y_err,binning_size)
x2 = reduced_chi_square(x_binned, y_binned, y_err_binned, model1[0,:], smooth_Flux_Ma_1Gyr_new)
x2_photo = chisquare_photo(model1[0,:], smooth_Flux_Ma_1Gyr_new,redshift_1,wave_list, band_list, photometric_flux, photometric_flux_err, photometric_flux_err_mod)
# x2_photo = chisquare_photo(model1[0,:], smooth_Flux_Ma_1Gyr_new, redshift_1,wave_list, band_list, photometric_flux, photometric_flux_err, photometric_flux_err_mod)
# print('binning data, model 1', n, (model1[0,binning_index]-model1[0,binning_index-1]), (x[int(n/2)]-x[int(n/2)-1]), binning_size)
# print('binning size, model 1', n, (model1[0,binning_index]-model1[0,binning_index-1]), (x[int(n/2)]-x[int(n/2)-1]), binning_size)
# x2_photo = reduced_chi_square(wave_list, photometric_flux, photometric_flux_err, model1[0,:], smooth_Flux_Ma_1Gyr_new)
try:
if 0.01<galaxy_age<13 and 0.0<=intrinsic_Av<=4.0 and not np.isinf(0.5*x2+0.5*x2_photo):
lnprobval = -0.5*(0.5*x2+0.5*x2_photo)#np.log(np.exp(-0.5*(0.5*weight1*x2+0.5*weight2*x2_photo)))
if np.isnan(lnprobval):
lnprobval = -np.inf
else:
lnprobval = -np.inf
except ValueError: # NaN value case
lnprobval = -np.inf
print('valueError',lnprobval)
if np.isinf(lnprobval):
print('lnprob:',lnprobval, x2, x2_photo,galaxy_age,intrinsic_Av)
return lnprobval
def minimize_age_AV_vector_weighted_return_flux(X):
galaxy_age= X[0]
intrinsic_Av = X[1]
n=len(x)
age_index = find_nearest(df_Ma.Age.unique(), galaxy_age)
age_prior = df_Ma.Age.unique()[age_index]
#print('galaxy age', galaxy_age, 'age prior:', age_prior)
AV_string = str(intrinsic_Av)
#print('intrinsic Av:', intrinsic_Av)
galaxy_age_string = str(age_prior)
split_galaxy_age_string = str(galaxy_age_string).split('.')
if age_prior < 1:
if galaxy_age < age_prior:
model1 = (M05_model_list[age_index]*(galaxy_age-df_Ma.Age.unique()[age_index-1]) \
+ M05_model_list[age_index-1]*(age_prior-galaxy_age))/(df_Ma.Age.unique()[age_index]-df_Ma.Age.unique()[age_index-1])
elif galaxy_age > age_prior:
model1 = (M05_model_list[age_index]*(df_Ma.Age.unique()[age_index+1]-galaxy_age) \
+ M05_model_list[age_index+1]*(galaxy_age-age_prior))/(df_Ma.Age.unique()[age_index+1]-df_Ma.Age.unique()[age_index])
elif galaxy_age == age_prior:
model1 = M05_model_list[age_index]
elif age_prior == 1.5:
if galaxy_age >=1.25 and galaxy_age <1.5:
model1 = 2.*(1.5-galaxy_age)*M05_model_list[30] + 2.*(galaxy_age-1.0)*M05_model_list[31]
elif galaxy_age >= 1.5 and galaxy_age <= 1.75:
model1 = 2.*(2.0-galaxy_age)*M05_model_list[31] + 2.*(galaxy_age-1.5)*M05_model_list[32]
elif len(split_galaxy_age_string[1])==1:
if galaxy_age >= 1.0 and galaxy_age < 1.25:
model1 = 2.*(1.5-galaxy_age)*M05_model_list[30] + 2.*(galaxy_age-1.0)*M05_model_list[31]
elif galaxy_age >=1.75 and galaxy_age < 2.0:
model1 = 2.*(2.0-galaxy_age)*M05_model_list[31] + 2.*(galaxy_age-1.5)*M05_model_list[32]
elif galaxy_age >= 2.0 and galaxy_age < 3.0:
model1 = (3.0-galaxy_age)*M05_model_list[32] + (galaxy_age-2.0)*M05_model_list[33]
elif galaxy_age >= 3.0 and galaxy_age < 4.0:
model1 = (4.0-galaxy_age)*M05_model_list[33] + (galaxy_age-3.0)*M05_model_list[34]
elif galaxy_age >= 4.0 and galaxy_age < 5.0:
model1 = (5.0-galaxy_age)*M05_model_list[34] + (galaxy_age-4.0)*M05_model_list[35]
elif galaxy_age >= 5.0 and galaxy_age < 6.0:
model1 = (6.0-galaxy_age)*M05_model_list[35] + (galaxy_age-5.0)*M05_model_list[36]
elif galaxy_age >= 6.0 and galaxy_age < 7.0:
model1 = (7.0-galaxy_age)*M05_model_list[36] + (galaxy_age-6.0)*M05_model_list[37]
elif galaxy_age >= 7.0 and galaxy_age < 8.0:
model1 = (8.0-galaxy_age)*M05_model_list[37] + (galaxy_age-7.0)*M05_model_list[38]
elif galaxy_age >= 8.0 and galaxy_age < 9.0:
model1 = (9.0-galaxy_age)*M05_model_list[38] + (galaxy_age-8.0)*M05_model_list[39]
elif galaxy_age >= 9.0 and galaxy_age < 10.0:
model1 = (10.0-galaxy_age)*M05_model_list[39] + (galaxy_age-9.0)*M05_model_list[40]
elif galaxy_age >= 10.0 and galaxy_age < 11.0:
model1 = (11.0-galaxy_age)*M05_model_list[40] + (galaxy_age-10.0)*M05_model_list[41]
elif galaxy_age >= 11.0 and galaxy_age < 12.0:
model1 = (12.0-galaxy_age)*M05_model_list[41] + (galaxy_age-11.0)*M05_model_list[42]
elif galaxy_age >= 12.0 and galaxy_age < 13.0:
model1 = (13.0-galaxy_age)*M05_model_list[42] + (galaxy_age-12.0)*M05_model_list[43]
elif galaxy_age >= 13.0 and galaxy_age < 14.0:
model1 = (14.0-galaxy_age)*M05_model_list[43] + (galaxy_age-13.0)*M05_model_list[44]
elif galaxy_age >= 14.0 and galaxy_age < 15.0:
model1 = (15.0-galaxy_age)*M05_model_list[44] + (galaxy_age-14.0)*M05_model_list[45]
else:
model1 = M05_model_list[age_index]
spectra_extinction = calzetti00(model1[0,:], intrinsic_Av, 4.05)
spectra_flux_correction = 10**(-0.4*spectra_extinction)
M05_flux_center = model1[1,:]*spectra_flux_correction
F_M05_index=700#167
Flux_M05_norm_new = M05_flux_center[F_M05_index]
smooth_Flux_Ma_1Gyr_new = M05_flux_center/Flux_M05_norm_new
binning_index = find_nearest(model1[0,:],np.median(x))
if binning_index == 0:
binning_index = 1
elif binning_index ==len(x):
binning_index = len(x)-1
if (x[int(n/2)]-x[int(n/2)-1]) > (model1[0,binning_index]-model1[0,binning_index-1]):
binning_size = int((x[int(n/2)]-x[int(n/2)-1])/(model1[0,binning_index]-model1[0,binning_index-1]))
model_wave_binned,model_flux_binned = binning_spec_keep_shape(model1[0,:], smooth_Flux_Ma_1Gyr_new,binning_size)
x2 = reduced_chi_square(x, y, y_err, model_wave_binned, model_flux_binned)
# x2_photo = chisquare_photo(model_wave_binned, model_flux_binned, redshift_1,wave_list, band_list, photometric_flux, photometric_flux_err, photometric_flux_err_mod)
x2_photo = chisquare_photo(model_wave_binned, model_flux_binned, redshift_1,wave_list, band_list, photometric_flux, photometric_flux_err, photometric_flux_err_mod)
# print('binning model, model 1', n, (model1[0,binning_index]-model1[0,binning_index-1]), (x[int(n/2)]-x[int(n/2)-1]), binning_size)
else:
binning_size = int((model1[0,binning_index]-model1[0,binning_index-1])/(x[int(n/2)]-x[int(n/2)-1]))
x_binned,y_binned,y_err_binned=binning_spec_keep_shape_x(x,y,y_err,binning_size)
x2 = reduced_chi_square(x_binned, y_binned, y_err_binned, model1[0,:], smooth_Flux_Ma_1Gyr_new)
x2_photo = chisquare_photo(model1[0,:], smooth_Flux_Ma_1Gyr_new, redshift_1,wave_list, band_list, photometric_flux, photometric_flux_err, photometric_flux_err_mod)
# x2_photo = chisquare_photo(model1[0,:], smooth_Flux_Ma_1Gyr_new, redshift_1,wave_list, band_list, photometric_flux, photometric_flux_err, photometric_flux_err_mod)
# print('binning data, model 1', n, (model1[0,binning_index]-model1[0,binning_index-1]), (x[int(n/2)]-x[int(n/2)-1]), binning_size)
# x2_photo = reduced_chi_square(wave_list, photometric_flux, photometric_flux_err, model1[0,:], smooth_Flux_Ma_1Gyr_new)
try:
if 0.01<galaxy_age<13 and 0.0<=intrinsic_Av<=4.0 and not np.isinf(0.5*x2+0.5*x2_photo):
x2_tot = 0.5*weight1*x2+0.5*weight2*x2_photo
else:
x2_tot = np.inf
except ValueError: # NaN value case
x2_tot = np.inf
print('valueError', x2_tot)
# print('model wave range', model1[0,0], model1[0,-1])
return x2_tot, model1[0,:], smooth_Flux_Ma_1Gyr_new
def minimize_age_AV_vector_weighted_return_chi2_sep(X):
galaxy_age= X[0]
intrinsic_Av = X[1]
n=len(x)
age_index = find_nearest(df_Ma.Age.unique(), galaxy_age)
age_prior = df_Ma.Age.unique()[age_index]
#print('galaxy age', galaxy_age, 'age prior:', age_prior)
AV_string = str(intrinsic_Av)
#print('intrinsic Av:', intrinsic_Av)
galaxy_age_string = str(age_prior)
split_galaxy_age_string = str(galaxy_age_string).split('.')
if age_prior < 1:
if galaxy_age < age_prior:
model1 = (M05_model_list[age_index]*(galaxy_age-df_Ma.Age.unique()[age_index-1]) \
+ M05_model_list[age_index-1]*(age_prior-galaxy_age))/(df_Ma.Age.unique()[age_index]-df_Ma.Age.unique()[age_index-1])
elif galaxy_age > age_prior:
model1 = (M05_model_list[age_index]*(df_Ma.Age.unique()[age_index+1]-galaxy_age) \
+ M05_model_list[age_index+1]*(galaxy_age-age_prior))/(df_Ma.Age.unique()[age_index+1]-df_Ma.Age.unique()[age_index])
elif galaxy_age == age_prior:
model1 = M05_model_list[age_index]
elif age_prior == 1.5:
if galaxy_age >=1.25 and galaxy_age <1.5:
model1 = 2.*(1.5-galaxy_age)*M05_model_list[30] + 2.*(galaxy_age-1.0)*M05_model_list[31]
elif galaxy_age >= 1.5 and galaxy_age <= 1.75:
model1 = 2.*(2.0-galaxy_age)*M05_model_list[31] + 2.*(galaxy_age-1.5)*M05_model_list[32]
elif len(split_galaxy_age_string[1])==1:
if galaxy_age >= 1.0 and galaxy_age < 1.25:
model1 = 2.*(1.5-galaxy_age)*M05_model_list[30] + 2.*(galaxy_age-1.0)*M05_model_list[31]
elif galaxy_age >=1.75 and galaxy_age < 2.0:
model1 = 2.*(2.0-galaxy_age)*M05_model_list[31] + 2.*(galaxy_age-1.5)*M05_model_list[32]
elif galaxy_age >= 2.0 and galaxy_age < 3.0:
model1 = (3.0-galaxy_age)*M05_model_list[32] + (galaxy_age-2.0)*M05_model_list[33]
elif galaxy_age >= 3.0 and galaxy_age < 4.0:
model1 = (4.0-galaxy_age)*M05_model_list[33] + (galaxy_age-3.0)*M05_model_list[34]
elif galaxy_age >= 4.0 and galaxy_age < 5.0:
model1 = (5.0-galaxy_age)*M05_model_list[34] + (galaxy_age-4.0)*M05_model_list[35]
elif galaxy_age >= 5.0 and galaxy_age < 6.0:
model1 = (6.0-galaxy_age)*M05_model_list[35] + (galaxy_age-5.0)*M05_model_list[36]
elif galaxy_age >= 6.0 and galaxy_age < 7.0:
model1 = (7.0-galaxy_age)*M05_model_list[36] + (galaxy_age-6.0)*M05_model_list[37]
elif galaxy_age >= 7.0 and galaxy_age < 8.0:
model1 = (8.0-galaxy_age)*M05_model_list[37] + (galaxy_age-7.0)*M05_model_list[38]
elif galaxy_age >= 8.0 and galaxy_age < 9.0:
model1 = (9.0-galaxy_age)*M05_model_list[38] + (galaxy_age-8.0)*M05_model_list[39]
elif galaxy_age >= 9.0 and galaxy_age < 10.0:
model1 = (10.0-galaxy_age)*M05_model_list[39] + (galaxy_age-9.0)*M05_model_list[40]
elif galaxy_age >= 10.0 and galaxy_age < 11.0:
model1 = (11.0-galaxy_age)*M05_model_list[40] + (galaxy_age-10.0)*M05_model_list[41]
elif galaxy_age >= 11.0 and galaxy_age < 12.0:
model1 = (12.0-galaxy_age)*M05_model_list[41] + (galaxy_age-11.0)*M05_model_list[42]
elif galaxy_age >= 12.0 and galaxy_age < 13.0:
model1 = (13.0-galaxy_age)*M05_model_list[42] + (galaxy_age-12.0)*M05_model_list[43]
elif galaxy_age >= 13.0 and galaxy_age < 14.0:
model1 = (14.0-galaxy_age)*M05_model_list[43] + (galaxy_age-13.0)*M05_model_list[44]
elif galaxy_age >= 14.0 and galaxy_age < 15.0:
model1 = (15.0-galaxy_age)*M05_model_list[44] + (galaxy_age-14.0)*M05_model_list[45]
else:
model1 = M05_model_list[age_index]
spectra_extinction = calzetti00(model1[0,:], intrinsic_Av, 4.05)
spectra_flux_correction = 10**(-0.4*spectra_extinction)
M05_flux_center = model1[1,:]*spectra_flux_correction
F_M05_index=700#167
Flux_M05_norm_new = M05_flux_center[F_M05_index]
smooth_Flux_Ma_1Gyr_new = M05_flux_center/Flux_M05_norm_new
binning_index = find_nearest(model1[0,:],np.median(x))
if binning_index == 0:
binning_index = 1
elif binning_index ==len(x):
binning_index = len(x)-1
if (x[int(n/2)]-x[int(n/2)-1]) > (model1[0,binning_index]-model1[0,binning_index-1]):
binning_size = int((x[int(n/2)]-x[int(n/2)-1])/(model1[0,binning_index]-model1[0,binning_index-1]))
model_wave_binned,model_flux_binned = binning_spec_keep_shape(model1[0,:], smooth_Flux_Ma_1Gyr_new,binning_size)
x2 = reduced_chi_square(x, y, y_err, model_wave_binned, model_flux_binned)
x2_photo = chisquare_photo(model_wave_binned, model_flux_binned, redshift_1,wave_list, band_list, photometric_flux, photometric_flux_err, photometric_flux_err_mod)
# x2_photo = chisquare_photo(model_wave_binned, model_flux_binned, redshift_1,wave_list, band_list, photometric_flux, photometric_flux_err, photometric_flux_err_mod)
# print('binning model, model 1', n, (model1[0,binning_index]-model1[0,binning_index-1]), (x[int(n/2)]-x[int(n/2)-1]), binning_size)
else:
binning_size = int((model1[0,binning_index]-model1[0,binning_index-1])/(x[int(n/2)]-x[int(n/2)-1]))
x_binned,y_binned,y_err_binned=binning_spec_keep_shape_x(x,y,y_err,binning_size)
x2 = reduced_chi_square(x_binned, y_binned, y_err_binned, model1[0,:], smooth_Flux_Ma_1Gyr_new)
x2_photo = chisquare_photo(model1[0,:], smooth_Flux_Ma_1Gyr_new,redshift_1,wave_list, band_list, photometric_flux, photometric_flux_err, photometric_flux_err_mod)
# x2_photo = chisquare_photo(model1[0,:], smooth_Flux_Ma_1Gyr_new, redshift_1,wave_list, band_list, photometric_flux, photometric_flux_err, photometric_flux_err_mod)
# print('binning data, model 1', n, (model1[0,binning_index]-model1[0,binning_index-1]), (x[int(n/2)]-x[int(n/2)-1]), binning_size)
# x2_photo = reduced_chi_square(wave_list, photometric_flux, photometric_flux_err, model1[0,:], smooth_Flux_Ma_1Gyr_new)
try:
if 0.01<galaxy_age<13 and 0.0<=intrinsic_Av<=4.0 and not np.isinf(0.5*x2+0.5*x2_photo):
pass
else:
x2 = np.inf
x2_photo = np.inf
except ValueError: # NaN value case
x2 = np.inf
x2_photo = np.inf
print('ValueError', x2)
return x2, x2_photo
def minimize_age_AV_vector_weighted_M13(X):
galaxy_age= X[0]
intrinsic_Av = X[1]
# print('minimize process age av grid M13:',X)
n=len(x)
age_index = find_nearest(df_M13.Age.unique(), galaxy_age)
age_prior = df_M13.Age.unique()[age_index]
age_prior = float(age_prior)
AV_string = str(intrinsic_Av)
galaxy_age_string = str(age_prior)
split_galaxy_age_string = str(galaxy_age_string).split('.')
if age_prior < 1e-5:
model2 = M13_model_list[0]
elif age_prior >= 1e-5 and age_prior < 1:
if galaxy_age < age_prior:
model2 = (M13_model_list[age_index]*(galaxy_age-df_M13.Age.unique()[age_index-1]) \
+ M13_model_list[age_index-1]*(age_prior-galaxy_age))/(df_M13.Age.unique()[age_index]-df_M13.Age.unique()[age_index-1])
elif galaxy_age > age_prior:
model2 = (M13_model_list[age_index]*(df_M13.Age.unique()[age_index+1]-galaxy_age) \
+ M13_model_list[age_index+1]*(galaxy_age-age_prior))/(df_M13.Age.unique()[age_index+1]-df_M13.Age.unique()[age_index])
elif galaxy_age == age_prior:
model2 = M13_model_list[age_index]
elif age_prior == 1.5:
if galaxy_age >=1.25 and galaxy_age <1.5:
model2 = 2.*(1.5-galaxy_age)*M13_model_list[51] + 2.*(galaxy_age-1.0)*M13_model_list[52]
elif galaxy_age >= 1.5 and galaxy_age <= 1.75:
model2 = 2.*(2.0-galaxy_age)*M13_model_list[52] + 2.*(galaxy_age-1.5)*M13_model_list[53]
elif len(split_galaxy_age_string[1])==1:
if galaxy_age >= 1.0 and galaxy_age < 1.25:
model2 = 2.*(1.5-galaxy_age)*M13_model_list[51] + 2.*(galaxy_age-1.0)*M13_model_list[52]
elif galaxy_age >=1.75 and galaxy_age < 2.0:
model2 = 2.*(2.0-galaxy_age)*M13_model_list[52] + 2.*(galaxy_age-1.5)*M13_model_list[53]
elif galaxy_age >= 2.0 and galaxy_age < 3.0:
model2 = (3.0-galaxy_age)*M13_model_list[53] + (galaxy_age-2.0)*M13_model_list[54]
elif galaxy_age >= 3.0 and galaxy_age < 4.0:
model2 = (4.0-galaxy_age)*M13_model_list[54] + (galaxy_age-3.0)*M13_model_list[55]
elif galaxy_age >= 4.0 and galaxy_age < 5.0:
model2 = (5.0-galaxy_age)*M13_model_list[55] + (galaxy_age-4.0)*M13_model_list[56]
elif galaxy_age >= 5.0 and galaxy_age < 6.0:
model2 = (6.0-galaxy_age)*M13_model_list[56] + (galaxy_age-5.0)*M13_model_list[57]
elif galaxy_age >= 6.0 and galaxy_age < 7.0:
model2 = (7.0-galaxy_age)*M13_model_list[57] + (galaxy_age-6.0)*M13_model_list[58]
elif galaxy_age >= 7.0 and galaxy_age < 8.0:
model2 = (8.0-galaxy_age)*M13_model_list[58] + (galaxy_age-7.0)*M13_model_list[59]
elif galaxy_age >= 8.0 and galaxy_age < 9.0:
model2 = (9.0-galaxy_age)*M13_model_list[59] + (galaxy_age-8.0)*M13_model_list[60]
elif galaxy_age >= 9.0 and galaxy_age < 10.0:
model2 = (10.0-galaxy_age)*M13_model_list[60] + (galaxy_age-9.0)*M13_model_list[61]
elif galaxy_age >= 10.0 and galaxy_age < 11.0:
model2 = (11.0-galaxy_age)*M13_model_list[61] + (galaxy_age-10.0)*M13_model_list[62]
elif galaxy_age >= 11.0 and galaxy_age < 12.0:
model2 = (12.0-galaxy_age)*M13_model_list[62] + (galaxy_age-11.0)*M13_model_list[63]
elif galaxy_age >= 12.0 and galaxy_age < 13.0:
model2 = (13.0-galaxy_age)*M13_model_list[63] + (galaxy_age-12.0)*M13_model_list[64]
elif galaxy_age >= 13.0 and galaxy_age < 14.0:
model2 = (14.0-galaxy_age)*M13_model_list[64] + (galaxy_age-13.0)*M13_model_list[65]
elif galaxy_age >= 14.0 and galaxy_age < 15.0:
model2 = (15.0-galaxy_age)*M13_model_list[65] + (galaxy_age-14.0)*M13_model_list[66]
else:
model2 = M13_model_list[age_index]
spectra_extinction = calzetti00(model2[0,:], intrinsic_Av, 4.05)
spectra_flux_correction = 10**(-0.4*spectra_extinction)
M13_flux_center = model2[1,:]*spectra_flux_correction
F_M13_index = 326#126##np.where(abs(model2[0,:]-norm_wavelength)<10.5)[0][0]
Flux_M13_norm_new = M13_flux_center[F_M13_index]
smooth_Flux_M13_1Gyr_new = M13_flux_center/Flux_M13_norm_new
binning_index = find_nearest(model2[0,:],np.median(x))
if binning_index == 0:
binning_index = 1
elif binning_index ==len(x):
binning_index = len(x)-1
if (x[int(n/2)]-x[int(n/2)-1]) > (model2[0,binning_index]-model2[0,binning_index-1]):
binning_size = int((x[int(n/2)]-x[int(n/2)-1])/(model2[0,binning_index]-model2[0,binning_index-1]))
model_wave_binned,model_flux_binned = binning_spec_keep_shape(model2[0,:], smooth_Flux_M13_1Gyr_new,binning_size)
x2 = reduced_chi_square(x, y, y_err, model_wave_binned, model_flux_binned)
if np.isnan(x2):
print('spectra chi2 is nan, binning model', model_flux_binned)
print('spectra model wave', model2[0,:],intrinsic_Av)
print('model flux before binning', spectra_extinction, spectra_flux_correction, M13_flux_center, Flux_M13_norm_new)
sys.exit()
x2_photo = chisquare_photo(model_wave_binned, model_flux_binned, redshift_1,wave_list, band_list, photometric_flux, photometric_flux_err, photometric_flux_err_mod)
# print('binning model, model 2', n, (model2[0,binning_index]-model2[0,binning_index-1]), (x[int(n/2)]-x[int(n/2)-1]),binning_size)
else:
binning_size = int((model2[0,binning_index]-model2[0,binning_index-1])/(x[int(n/2)]-x[int(n/2)-1]))
x_binned,y_binned,y_err_binned = binning_spec_keep_shape_x(x,y,y_err,binning_size)
x2 = reduced_chi_square(x_binned, y_binned, y_err_binned, model2[0,:], smooth_Flux_M13_1Gyr_new)
if np.isnan(x2):
print('spectra chi2 is nan,binning data',x_binned)
print('spectra model wave', model2[0,:],intrinsic_Av)
print('model flux before binning', spectra_extinction, spectra_flux_correction, M13_flux_center, Flux_M13_norm_new)
sys.exit()
x2_photo = chisquare_photo(model2[0,:], smooth_Flux_M13_1Gyr_new,redshift_1,wave_list, band_list, photometric_flux, photometric_flux_err, photometric_flux_err_mod)
if np.isnan(x2_photo):
print('model 2 photo nan', x2_photo)
# print('binning data, model 2', n, (model2[0,binning_index]-model2[0,binning_index-1]), (x[int(n/2)]-x[int(n/2)-1]),binning_size)
# x2_photo = reduced_chi_square(wave_list, photometric_flux, photometric_flux_err, model2[0,:], smooth_Flux_M13_1Gyr_new)
# print(x2_photo)
try:
if 0.01<galaxy_age<13 and 0.0<=intrinsic_Av<=4.0 and not np.isinf(0.5*x2+0.5*x2_photo):
x2_tot = 0.5*weight1*x2+0.5*weight2*x2_photo
else:
x2_tot = np.inf
except ValueError: # NaN value case
x2_tot = np.inf
print('ValueError', x2_tot)
return x2_tot
def lg_minimize_age_AV_vector_weighted_M13(X):
tik = time.clock()
galaxy_age= X[0]
intrinsic_Av = X[1]
n=len(x)
age_index = find_nearest(df_M13.Age.unique(), galaxy_age)
age_prior = df_M13.Age.unique()[age_index]
age_prior = float(age_prior)
AV_string = str(intrinsic_Av)
galaxy_age_string = str(age_prior)
split_galaxy_age_string = str(galaxy_age_string).split('.')
model2 = np.zeros((2,762))
if age_prior < 1e-5:
model2 = M13_model_list[0]
elif age_prior >= 1e-5 and age_prior < 1:
if galaxy_age < age_prior:
model2 = (M13_model_list[age_index]*(galaxy_age-df_M13.Age.unique()[age_index-1]) \
+ M13_model_list[age_index-1]*(age_prior-galaxy_age))/(df_M13.Age.unique()[age_index]-df_M13.Age.unique()[age_index-1])
# print('age interval', (galaxy_age-df_M13.Age.unique()[age_index-1]), (age_prior-galaxy_age))
elif galaxy_age > age_prior:
model2 = (M13_model_list[age_index]*(df_M13.Age.unique()[age_index+1]-galaxy_age) \
+ M13_model_list[age_index+1]*(galaxy_age-age_prior))/(df_M13.Age.unique()[age_index+1]-df_M13.Age.unique()[age_index])
elif galaxy_age == age_prior:
model2 = M13_model_list[age_index]
elif age_prior == 1.5:
if galaxy_age >=1.25 and galaxy_age <1.5:
model2 = 2.*(1.5-galaxy_age)*M13_model_list[51] + 2.*(galaxy_age-1.0)*M13_model_list[52]
elif galaxy_age >= 1.5 and galaxy_age <= 1.75:
model2 = 2.*(2.0-galaxy_age)*M13_model_list[52] + 2.*(galaxy_age-1.5)*M13_model_list[53]
elif len(split_galaxy_age_string[1])==1:
if galaxy_age >= 1.0 and galaxy_age < 1.25:
model2 = 2.*(1.5-galaxy_age)*M13_model_list[51] + 2.*(galaxy_age-1.0)*M13_model_list[52]
elif galaxy_age >=1.75 and galaxy_age < 2.0:
model2 = 2.*(2.0-galaxy_age)*M13_model_list[52] + 2.*(galaxy_age-1.5)*M13_model_list[53]
elif galaxy_age >= 2.0 and galaxy_age < 3.0:
model2 = (3.0-galaxy_age)*M13_model_list[53] + (galaxy_age-2.0)*M13_model_list[54]
elif galaxy_age >= 3.0 and galaxy_age < 4.0:
model2 = (4.0-galaxy_age)*M13_model_list[54] + (galaxy_age-3.0)*M13_model_list[55]
elif galaxy_age >= 4.0 and galaxy_age < 5.0:
model2 = (5.0-galaxy_age)*M13_model_list[55] + (galaxy_age-4.0)*M13_model_list[56]
elif galaxy_age >= 5.0 and galaxy_age < 6.0:
model2 = (6.0-galaxy_age)*M13_model_list[56] + (galaxy_age-5.0)*M13_model_list[57]
elif galaxy_age >= 6.0 and galaxy_age < 7.0:
model2 = (7.0-galaxy_age)*M13_model_list[57] + (galaxy_age-6.0)*M13_model_list[58]
elif galaxy_age >= 7.0 and galaxy_age < 8.0:
model2 = (8.0-galaxy_age)*M13_model_list[58] + (galaxy_age-7.0)*M13_model_list[59]
elif galaxy_age >= 8.0 and galaxy_age < 9.0:
model2 = (9.0-galaxy_age)*M13_model_list[59] + (galaxy_age-8.0)*M13_model_list[60]
elif galaxy_age >= 9.0 and galaxy_age < 10.0:
model2 = (10.0-galaxy_age)*M13_model_list[60] + (galaxy_age-9.0)*M13_model_list[61]
elif galaxy_age >= 10.0 and galaxy_age < 11.0:
model2 = (11.0-galaxy_age)*M13_model_list[61] + (galaxy_age-10.0)*M13_model_list[62]
elif galaxy_age >= 11.0 and galaxy_age < 12.0:
model2 = (12.0-galaxy_age)*M13_model_list[62] + (galaxy_age-11.0)*M13_model_list[63]
elif galaxy_age >= 12.0 and galaxy_age < 13.0:
model2 = (13.0-galaxy_age)*M13_model_list[63] + (galaxy_age-12.0)*M13_model_list[64]
elif galaxy_age >= 13.0 and galaxy_age < 14.0:
model2 = (14.0-galaxy_age)*M13_model_list[64] + (galaxy_age-13.0)*M13_model_list[65]
elif galaxy_age >= 14.0 and galaxy_age < 15.0:
model2 = (15.0-galaxy_age)*M13_model_list[65] + (galaxy_age-14.0)*M13_model_list[66]
else:
model2 = M13_model_list[age_index]
spectra_extinction = calzetti00(model2[0,:], intrinsic_Av, 4.05)
spectra_flux_correction = 10**(-0.4*spectra_extinction)
M13_flux_center = model2[1,:]*spectra_flux_correction
F_M13_index = 326#126##np.where(abs(model2[0,:]-norm_wavelength)<10.5)[0][0]
Flux_M13_norm_new = M13_flux_center[F_M13_index]
smooth_Flux_M13_1Gyr_new = M13_flux_center/Flux_M13_norm_new
binning_index = find_nearest(model2[0,:],np.median(x))
if binning_index == 0:
binning_index = 1
elif binning_index == len(model2[0,:]):
binning_index = len(model2[0,:])-1
# print('binning index:',binning_index,len(model2[0,:]),len(x), model2[:,binning_index-2:binning_index])
# print('galaxy age:', galaxy_age, age_prior,age_index)
# print(x, n)
# print(len(model2),galaxy_age, age_prior, age_index, len(x), len(model2), np.median(x), np.min(model2[0,:]),np.max(model2[0,:]), binning_index)
if (x[int(n/2)]-x[int(n/2)-1]) > (model2[0,binning_index]-model2[0,binning_index-1]):
binning_size = int((x[int(n/2)]-x[int(n/2)-1])/(model2[0,binning_index]-model2[0,binning_index-1]))
# print('bin size', model2[0,binning_index],\
# model2[0,binning_index-1],\
# (model2[0,binning_index]-model2[0,binning_index-1]),\
# int((x[int(n/2)]-x[int(n/2)-1])),\
# binning_size)
model_wave_binned,model_flux_binned = binning_spec_keep_shape(model2[0,:], smooth_Flux_M13_1Gyr_new, binning_size)
x2 = reduced_chi_square(x, y, y_err, model_wave_binned, model_flux_binned)
x2_photo = chisquare_photo(model_wave_binned, model_flux_binned, redshift_1,wave_list, band_list, photometric_flux, photometric_flux_err, photometric_flux_err_mod)
else:
binning_size = int((model2[0,binning_index]-model2[0,binning_index-1])/(x[int(n/2)]-x[int(n/2)-1]))
x_binned,y_binned,y_err_binned = binning_spec_keep_shape_x(x,y,y_err,binning_size)
x2 = reduced_chi_square(x_binned, y_binned, y_err_binned, model2[0,:], smooth_Flux_M13_1Gyr_new)
x2_photo = chisquare_photo(model2[0,:], smooth_Flux_M13_1Gyr_new,redshift_1,wave_list, band_list, photometric_flux, photometric_flux_err, photometric_flux_err_mod)
tok = time.clock()
# print('time for lg_minimize',tok-tik)
try:
if 0.01<galaxy_age<13 and 0.0<=intrinsic_Av<=4.0 and not np.isinf(0.5*x2+0.5*x2_photo):
lnprobval = -0.5*(0.5*x2+0.5*x2_photo)#np.log(np.exp(-0.5*(0.5*weight1*x2+0.5*weight2*x2_photo)))
if np.isnan(lnprobval):
lnprobval = -np.inf
else:
lnprobval = -np.inf
except ValueError: # NaN value case
lnprobval = -np.inf
print('valueError',lnprobval,x2, x2_photo)
# print('lnprob:',lnprobval)
return lnprobval
def minimize_age_AV_vector_weighted_M13_return_flux(X):
galaxy_age= X[0]
intrinsic_Av = X[1]
n=len(x)
age_index = find_nearest(df_M13.Age.unique(), galaxy_age)
age_prior = df_M13.Age.unique()[age_index]
age_prior = float(age_prior)
AV_string = str(intrinsic_Av)
galaxy_age_string = str(age_prior)
split_galaxy_age_string = str(galaxy_age_string).split('.')
model2 = np.zeros((2,762))
if age_prior < 1e-5:
model2 = M13_model_list[0]
elif age_prior >= 1e-5 and age_prior < 1:
if galaxy_age < age_prior:
model2 = (M13_model_list[age_index]*(galaxy_age-df_M13.Age.unique()[age_index-1]) \
+ M13_model_list[age_index-1]*(age_prior-galaxy_age))/(df_M13.Age.unique()[age_index]-df_M13.Age.unique()[age_index-1])
elif galaxy_age > age_prior:
model2 = (M13_model_list[age_index]*(df_M13.Age.unique()[age_index+1]-galaxy_age) \
+ M13_model_list[age_index+1]*(galaxy_age-age_prior))/(df_M13.Age.unique()[age_index+1]-df_M13.Age.unique()[age_index])
elif galaxy_age == age_prior:
model2 = M13_model_list[age_index]
elif age_prior == 1.5:
if galaxy_age >=1.25 and galaxy_age <1.5:
model2 = 2.*(1.5-galaxy_age)*M13_model_list[51] + 2.*(galaxy_age-1.0)*M13_model_list[52]
elif galaxy_age >= 1.5 and galaxy_age <= 1.75:
model2 = 2.*(2.0-galaxy_age)*M13_model_list[52] + 2.*(galaxy_age-1.5)*M13_model_list[53]
elif len(split_galaxy_age_string[1])==1:
if galaxy_age >= 1.0 and galaxy_age < 1.25:
model2 = 2.*(1.5-galaxy_age)*M13_model_list[51] + 2.*(galaxy_age-1.0)*M13_model_list[52]
elif galaxy_age >=1.75 and galaxy_age < 2.0:
model2 = 2.*(2.0-galaxy_age)*M13_model_list[52] + 2.*(galaxy_age-1.5)*M13_model_list[53]
elif galaxy_age >= 2.0 and galaxy_age < 3.0:
model2[0,:] = (3.0-galaxy_age)*M13_model_list[53][0,:] + (galaxy_age-2.0)*M13_model_list[54][0,:]
model2[1,:] = (3.0-galaxy_age)*M13_model_list[53][1,:] + (galaxy_age-2.0)*M13_model_list[54][1,:]
elif galaxy_age >= 3.0 and galaxy_age < 4.0:
model2 = (4.0-galaxy_age)*M13_model_list[54] + (galaxy_age-3.0)*M13_model_list[55]
elif galaxy_age >= 4.0 and galaxy_age < 5.0:
model2 = (5.0-galaxy_age)*M13_model_list[55] + (galaxy_age-4.0)*M13_model_list[56]
elif galaxy_age >= 5.0 and galaxy_age < 6.0:
model2 = (6.0-galaxy_age)*M13_model_list[56] + (galaxy_age-5.0)*M13_model_list[57]
elif galaxy_age >= 6.0 and galaxy_age < 7.0:
model2 = (7.0-galaxy_age)*M13_model_list[57] + (galaxy_age-6.0)*M13_model_list[58]
elif galaxy_age >= 7.0 and galaxy_age < 8.0:
model2 = (8.0-galaxy_age)*M13_model_list[58] + (galaxy_age-7.0)*M13_model_list[59]
elif galaxy_age >= 8.0 and galaxy_age < 9.0:
model2 = (9.0-galaxy_age)*M13_model_list[59] + (galaxy_age-8.0)*M13_model_list[60]
elif galaxy_age >= 9.0 and galaxy_age < 10.0:
model2 = (10.0-galaxy_age)*M13_model_list[60] + (galaxy_age-9.0)*M13_model_list[61]
elif galaxy_age >= 10.0 and galaxy_age < 11.0:
model2 = (11.0-galaxy_age)*M13_model_list[61] + (galaxy_age-10.0)*M13_model_list[62]
elif galaxy_age >= 11.0 and galaxy_age < 12.0:
model2 = (12.0-galaxy_age)*M13_model_list[62] + (galaxy_age-11.0)*M13_model_list[63]
elif galaxy_age >= 12.0 and galaxy_age < 13.0:
model2 = (13.0-galaxy_age)*M13_model_list[63] + (galaxy_age-12.0)*M13_model_list[64]
elif galaxy_age >= 13.0 and galaxy_age < 14.0:
model2 = (14.0-galaxy_age)*M13_model_list[64] + (galaxy_age-13.0)*M13_model_list[65]
elif galaxy_age >= 14.0 and galaxy_age < 15.0:
model2 = (15.0-galaxy_age)*M13_model_list[65] + (galaxy_age-14.0)*M13_model_list[66]
else:
model2 = M13_model_list[age_index]
spectra_extinction = calzetti00(model2[0,:], intrinsic_Av, 4.05)
spectra_flux_correction = 10**(-0.4*spectra_extinction)
M13_flux_center = model2[1,:]*spectra_flux_correction
F_M13_index = 326#126##np.where(abs(model2[0,:]-norm_wavelength)<10.5)[0][0]
Flux_M13_norm_new = M13_flux_center[F_M13_index]
smooth_Flux_M13_1Gyr_new = M13_flux_center/Flux_M13_norm_new
binning_index = find_nearest(model2[0,:],np.median(x))
if binning_index == 0:
binning_index = 1
elif binning_index ==len(x):
binning_index = len(x)-1
if (x[int(n/2)]-x[int(n/2)-1]) > (model2[0,binning_index]-model2[0,binning_index-1]):
binning_size = int((x[int(n/2)]-x[int(n/2)-1])/(model2[0,binning_index]-model2[0,binning_index-1]))
model_wave_binned,model_flux_binned = binning_spec_keep_shape(model2[0,:], smooth_Flux_M13_1Gyr_new, binning_size)
x2 = reduced_chi_square(x, y, y_err, model_wave_binned, model_flux_binned)
x2_photo = chisquare_photo(model_wave_binned, model_flux_binned, redshift_1,wave_list, band_list, photometric_flux, photometric_flux_err, photometric_flux_err_mod)
smooth_Flux_M13_1Gyr_new = model_flux_binned
else:
binning_size = int((model2[0,binning_index]-model2[0,binning_index-1])/(x[int(n/2)]-x[int(n/2)-1]))
x_binned,y_binned,y_err_binned = binning_spec_keep_shape_x(x,y,y_err,binning_size)
x2 = reduced_chi_square(x_binned, y_binned, y_err_binned, model2[0,:], smooth_Flux_M13_1Gyr_new)
x2_photo = chisquare_photo(model2[0,:], smooth_Flux_M13_1Gyr_new, redshift_1,wave_list, band_list, photometric_flux, photometric_flux_err, photometric_flux_err_mod)
try:
if 0.01<galaxy_age<13 and 0.0<=intrinsic_Av<=4.0 and not np.isinf(0.5*x2+0.5*x2_photo):
x2_tot = 0.5*weight1*x2+0.5*weight2*x2_photo
else:
x2_tot = np.inf
except ValueError: # NaN value case
x2_tot = np.inf
print('valueError', x2_tot)
# print('model wave range', model2[0,0], model2[0,-1], split_galaxy_age_string )
# print('model wave separately', M13_model_list[53][0,0],M13_model_list[53][0,-1],len(M13_model_list[53][0,:]),len(M13_model_list[54][0,:]),M13_model_list[54][0,0],M13_model_list[53][0,-1])
# print('model test', model_test[0,0], model_test[0,-1])
# print('age',galaxy_age,age_prior)
return x2_tot, model2[0,:], smooth_Flux_M13_1Gyr_new
def minimize_age_AV_vector_weighted_M13_return_chi2_sep(X):
galaxy_age= X[0]
intrinsic_Av = X[1]
n=len(x)
age_index = find_nearest(df_M13.Age.unique(), galaxy_age)
age_prior = df_M13.Age.unique()[age_index]
age_prior = float(age_prior)
AV_string = str(intrinsic_Av)
galaxy_age_string = str(age_prior)
split_galaxy_age_string = str(galaxy_age_string).split('.')
if age_prior < 1e-5:
model2 = M13_model_list[0]
elif age_prior >= 1e-5 and age_prior < 1:
if galaxy_age < age_prior:
model2 = (M13_model_list[age_index]*(galaxy_age-df_M13.Age.unique()[age_index-1]) \
+ M13_model_list[age_index-1]*(age_prior-galaxy_age))/(df_M13.Age.unique()[age_index]-df_M13.Age.unique()[age_index-1])
elif galaxy_age > age_prior:
model2 = (M13_model_list[age_index]*(df_M13.Age.unique()[age_index+1]-galaxy_age) \
+ M13_model_list[age_index+1]*(galaxy_age-age_prior))/(df_M13.Age.unique()[age_index+1]-df_M13.Age.unique()[age_index])
elif galaxy_age == age_prior:
model2 = M13_model_list[age_index]
elif age_prior == 1.5:
if galaxy_age >=1.25 and galaxy_age <1.5:
model2 = 2.*(1.5-galaxy_age)*M13_model_list[51] + 2.*(galaxy_age-1.0)*M13_model_list[52]
elif galaxy_age >= 1.5 and galaxy_age <= 1.75:
model2 = 2.*(2.0-galaxy_age)*M13_model_list[52] + 2.*(galaxy_age-1.5)*M13_model_list[53]
elif len(split_galaxy_age_string[1])==1:
if galaxy_age >= 1.0 and galaxy_age < 1.25:
model2 = 2.*(1.5-galaxy_age)*M13_model_list[51] + 2.*(galaxy_age-1.0)*M13_model_list[52]
elif galaxy_age >=1.75 and galaxy_age < 2.0:
model2 = 2.*(2.0-galaxy_age)*M13_model_list[52] + 2.*(galaxy_age-1.5)*M13_model_list[53]
elif galaxy_age >= 2.0 and galaxy_age < 3.0:
model2 = (3.0-galaxy_age)*M13_model_list[53] + (galaxy_age-2.0)*M13_model_list[54]
elif galaxy_age >= 3.0 and galaxy_age < 4.0:
model2 = (4.0-galaxy_age)*M13_model_list[54] + (galaxy_age-3.0)*M13_model_list[55]
elif galaxy_age >= 4.0 and galaxy_age < 5.0:
model2 = (5.0-galaxy_age)*M13_model_list[55] + (galaxy_age-4.0)*M13_model_list[56]
elif galaxy_age >= 5.0 and galaxy_age < 6.0:
model2 = (6.0-galaxy_age)*M13_model_list[56] + (galaxy_age-5.0)*M13_model_list[57]
elif galaxy_age >= 6.0 and galaxy_age < 7.0:
model2 = (7.0-galaxy_age)*M13_model_list[57] + (galaxy_age-6.0)*M13_model_list[58]
elif galaxy_age >= 7.0 and galaxy_age < 8.0:
model2 = (8.0-galaxy_age)*M13_model_list[58] + (galaxy_age-7.0)*M13_model_list[59]
elif galaxy_age >= 8.0 and galaxy_age < 9.0:
model2 = (9.0-galaxy_age)*M13_model_list[59] + (galaxy_age-8.0)*M13_model_list[60]
elif galaxy_age >= 9.0 and galaxy_age < 10.0:
model2 = (10.0-galaxy_age)*M13_model_list[60] + 2.*(galaxy_age-9.0)*M13_model_list[61]
elif galaxy_age >= 10.0 and galaxy_age < 11.0:
model2 = (11.0-galaxy_age)*M13_model_list[61] + 2.*(galaxy_age-10.0)*M13_model_list[62]
elif galaxy_age >= 11.0 and galaxy_age < 12.0:
model2 = (12.0-galaxy_age)*M13_model_list[62] + 2.*(galaxy_age-11.0)*M13_model_list[63]
elif galaxy_age >= 12.0 and galaxy_age < 13.0:
model2 = (13.0-galaxy_age)*M13_model_list[63] + 2.*(galaxy_age-12.0)*M13_model_list[64]
elif galaxy_age >= 13.0 and galaxy_age < 14.0:
model2 = (14.0-galaxy_age)*M13_model_list[64] + 2.*(galaxy_age-13.0)*M13_model_list[65]
elif galaxy_age >= 14.0 and galaxy_age < 15.0:
model2 = (15.0-galaxy_age)*M13_model_list[65] + 2.*(galaxy_age-14.0)*M13_model_list[66]
else:
model2 = M13_model_list[age_index]
spectra_extinction = calzetti00(model2[0,:], intrinsic_Av, 4.05)
spectra_flux_correction = 10**(-0.4*spectra_extinction)
M13_flux_center = model2[1,:]*spectra_flux_correction
F_M13_index = 326#126##np.where(abs(model2[0,:]-norm_wavelength)<10.5)[0][0]
Flux_M13_norm_new = M13_flux_center[F_M13_index]
smooth_Flux_M13_1Gyr_new = M13_flux_center/Flux_M13_norm_new
binning_index = find_nearest(model2[0,:],np.median(x))
if binning_index == 0:
binning_index = 1
elif binning_index ==len(x):
binning_index = len(x)-1
if (x[int(n/2)]-x[int(n/2)-1]) > (model2[0,binning_index]-model2[0,binning_index-1]):
binning_size = int((x[int(n/2)]-x[int(n/2)-1])/(model2[0,binning_index]-model2[0,binning_index-1]))
model_wave_binned,model_flux_binned = binning_spec_keep_shape(model2[0,:], smooth_Flux_M13_1Gyr_new,binnning_size)
x2 = reduced_chi_square(x, y, y_err, model_wave_binned, model_flux_binned)
x2_photo = chisquare_photo(model_wave_binned, model_flux_binned, redshift_1,wave_list, band_list, photometric_flux, photometric_flux_err, photometric_flux_err_mod)
# print('binning model, model 2', n, (model2[0,binning_index]-model2[0,binning_index-1]), (x[int(n/2)]-x[int(n/2)-1]),binning_size)
else:
binning_size = int((model2[0,binning_index]-model2[0,binning_index-1])/(x[int(n/2)]-x[int(n/2)-1]))
x_binned,y_binned,y_err_binned = binning_spec_keep_shape_x(x,y,y_err,binning_size)
x2 = reduced_chi_square(x_binned, y_binned, y_err_binned, model2[0,:], smooth_Flux_M13_1Gyr_new)
x2_photo = chisquare_photo(model2[0,:], smooth_Flux_M13_1Gyr_new,redshift_1,wave_list, band_list, photometric_flux, photometric_flux_err, photometric_flux_err_mod)
try:
if 0.01<galaxy_age<13 and 0.0<=intrinsic_Av<=4.0 and not np.isinf(0.5*x2+0.5*x2_photo):
pass
else:
x2 = np.inf
x2_photo = np.inf
except ValueError: # NaN value case
x2 = np.inf
x2_photo = np.inf
print('ValueError', x2)
return x2, x2_photo
def minimize_age_AV_vector_weighted_BC03(X):
galaxy_age= X[0]
intrinsic_Av = X[1]
n=len(x)
age_index = find_nearest(BC03_age_list_num, galaxy_age)
age_prior = BC03_age_list_num[age_index]
AV_string = str(intrinsic_Av)
# print(galaxy_age,age_prior)
if galaxy_age == age_prior:
model3_flux = BC03_flux_array[age_index, :7125]
elif galaxy_age < age_prior:
age_interval = BC03_age_list_num[age_index+1] - BC03_age_list_num[age_index]
model3_flux = (BC03_flux_array[age_index, :7125]*(BC03_age_list_num[age_index+1]-galaxy_age) + BC03_flux_array[age_index+1, :7125]*(galaxy_age-BC03_age_list_num[age_index]))*1./age_interval
elif galaxy_age > age_prior:
age_interval = BC03_age_list_num[age_index] - BC03_age_list_num[age_index-1]
model3_flux = (BC03_flux_array[age_index, :7125]*(BC03_age_list_num[age_index]-galaxy_age) + BC03_flux_array[age_index+1, :7125]*(galaxy_age-BC03_age_list_num[age_index-1]))*1./age_interval
spectra_extinction = calzetti00(BC03_wave_list_num, intrinsic_Av, 4.05)
spectra_flux_correction = 10**(-0.4*spectra_extinction)
BC03_flux_attenuated = model3_flux*spectra_flux_correction
BC03_flux_norm = BC03_flux_attenuated[2556]
BC03_flux_attenuated = BC03_flux_attenuated/BC03_flux_norm
binning_index = find_nearest(BC03_wave_list_num, np.median(x))
if (x[int(n/2)]-x[int(n/2)-1]) < (BC03_wave_list_num[binning_index]-BC03_wave_list_num[binning_index-1]):
binning_size = int((BC03_wave_list_num[binning_index]-BC03_wave_list_num[binning_index-1])/(x[int(n/2)]-x[int(n/2)-1]))
x_binned,y_binned,y_err_binned = binning_spec_keep_shape_x(x,y,y_err,binning_size)
x2 = reduced_chi_square(x_binned, y_binned, y_err_binned, BC03_wave_list_num, BC03_flux_attenuated)
x2_photo = chisquare_photo(BC03_wave_list_num, BC03_flux_attenuated, redshift_1,wave_list, band_list, photometric_flux, photometric_flux_err, photometric_flux_err_mod)
# print('bin data', n, binning_size, x2)
else:
binning_size = int((x[int(n/2)]-x[int(n/2)-1])/(BC03_wave_list_num[binning_index]-BC03_wave_list_num[binning_index-1]))
model_wave_binned, model_flux_binned = binning_spec_keep_shape(BC03_wave_list_num, BC03_flux_attenuated,binning_size)
x2 = reduced_chi_square(x, y, y_err, model_wave_binned, model_flux_binned)
x2_photo = chisquare_photo(model_wave_binned, model_flux_binned,redshift_1,wave_list, band_list, photometric_flux, photometric_flux_err, photometric_flux_err_mod)
# print('bin model',binning_size, x2)
# print('binning size, model 3', n, (BC03_wave_list_num[binning_index]-BC03_wave_list_num[binning_index-1]), (x[int(n/2)]-x[int(n/2)-1]), binning_size)
# x2_photo = reduced_chi_square(wave_list, photometric_flux, photometric_flux_err, BC03_wave_list_num, BC03_flux_attenuated)
# print('BC x2_nu',x2,x2_photo,0.5*weight1*x2+0.5*weight2*x2_photo)
return 0.5*weight1*x2+0.5*weight2*x2_photo
def lg_minimize_age_AV_vector_weighted_BC03(X):
galaxy_age= X[0]
intrinsic_Av = X[1]
n=len(x)
age_index = find_nearest(BC03_age_list_num, galaxy_age)
age_prior = BC03_age_list_num[age_index]
AV_string = str(intrinsic_Av)
if galaxy_age == age_prior:
model3_flux = BC03_flux_array[age_index, :7125]
elif galaxy_age < age_prior and galaxy_age <1.97500006e+01:
age_interval = BC03_age_list_num[age_index+1] - BC03_age_list_num[age_index]
model3_flux = (BC03_flux_array[age_index, :7125]*(BC03_age_list_num[age_index+1]-galaxy_age) + BC03_flux_array[age_index+1, :7125]*(galaxy_age-BC03_age_list_num[age_index]))*1./age_interval
elif galaxy_age > age_prior and galaxy_age <1.97500006e+01:
age_interval = BC03_age_list_num[age_index] - BC03_age_list_num[age_index-1]
model3_flux = (BC03_flux_array[age_index, :7125]*(BC03_age_list_num[age_index]-galaxy_age) + BC03_flux_array[age_index+1, :7125]*(galaxy_age-BC03_age_list_num[age_index-1]))*1./age_interval
else:
model3_flux = BC03_flux_array[-1, :7125]
spectra_extinction = calzetti00(BC03_wave_list_num, intrinsic_Av, 4.05)
spectra_flux_correction = 10**(-0.4*spectra_extinction)
BC03_flux_attenuated = model3_flux*spectra_flux_correction
BC03_flux_norm = BC03_flux_attenuated[2556]
BC03_flux_attenuated = BC03_flux_attenuated/BC03_flux_norm
binning_index = find_nearest(BC03_wave_list_num, np.median(x))
if (x[int(n/2)]-x[int(n/2)-1]) < (BC03_wave_list_num[binning_index]-BC03_wave_list_num[binning_index-1]):
binning_size = int((BC03_wave_list_num[binning_index]-BC03_wave_list_num[binning_index-1])/(x[int(n/2)]-x[int(n/2)-1]))
x_binned,y_binned,y_err_binned=binning_spec_keep_shape_x(x,y,y_err,binning_size)
x2 = reduced_chi_square(x_binned, y_binned, y_err_binned, BC03_wave_list_num, BC03_flux_attenuated)
x2_photo = chisquare_photo(BC03_wave_list_num, BC03_flux_attenuated, redshift_1,wave_list, band_list, photometric_flux, photometric_flux_err, photometric_flux_err_mod)
# print('bin data', binning_size, x2)
else:
binning_size = int((x[int(n/2)]-x[int(n/2)-1])/(BC03_wave_list_num[binning_index]-BC03_wave_list_num[binning_index-1]))
model_wave_binned, model_flux_binned = binning_spec_keep_shape(BC03_wave_list_num, BC03_flux_attenuated,binning_size)
x2 = reduced_chi_square(x, y, y_err, model_wave_binned, model_flux_binned)
x2_photo = chisquare_photo(model_wave_binned, model_flux_binned,redshift_1,wave_list, band_list, photometric_flux, photometric_flux_err, photometric_flux_err_mod)
# print('bin model',binning_size, x2)
# print('binning size, model 3', n, (BC03_wave_list_num[binning_index]-BC03_wave_list_num[binning_index-1]), (x[int(n/2)]-x[int(n/2)-1]), binning_size)
# x2_photo = reduced_chi_square(wave_list, photometric_flux, photometric_flux_err, BC03_wave_list_num, BC03_flux_attenuated)
if 0.01<galaxy_age<13 and 0.0<=intrinsic_Av<=4.0 and not np.isinf(0.5*x2+0.5*1e-3*x2_photo):
return np.log(np.exp(-0.5*(0.5*weight1*x2+0.5*weight2*x2_photo)))
else:
return -np.inf
def minimize_age_AV_vector_weighted_BC03_mod_no_weight_return_flux(X):
galaxy_age= X[0]
intrinsic_Av = X[1]
n=len(x)
age_index = find_nearest(BC03_age_list_num, galaxy_age)
age_prior = BC03_age_list_num[age_index]
AV_string = str(intrinsic_Av)
if galaxy_age == age_prior:
model3_flux = BC03_flux_array[age_index, :7125]
elif galaxy_age < age_prior and galaxy_age <1.97500006e+01:
age_interval = BC03_age_list_num[age_index] - BC03_age_list_num[age_index-1]
model3_flux = (BC03_flux_array[age_index-1, :7125]*(BC03_age_list_num[age_index]-galaxy_age)\
+ BC03_flux_array[age_index, :7125]*(galaxy_age-BC03_age_list_num[age_index-1]))*1./age_interval
elif galaxy_age > age_prior and galaxy_age <1.97500006e+01:
age_interval = BC03_age_list_num[age_index+1] - BC03_age_list_num[age_index]
model3_flux = (BC03_flux_array[age_index, :7125]*(BC03_age_list_num[age_index+1]-galaxy_age)\
+ BC03_flux_array[age_index+1, :7125]*(galaxy_age-BC03_age_list_num[age_index]))*1./age_interval
spectra_extinction = calzetti00(BC03_wave_list_num, intrinsic_Av, 4.05)
spectra_flux_correction = 10**(-0.4*spectra_extinction)
BC03_flux_attenuated = model3_flux*spectra_flux_correction
BC03_flux_norm = BC03_flux_attenuated[2556]
BC03_flux_attenuated = BC03_flux_attenuated/BC03_flux_norm
binning_index = find_nearest(BC03_wave_list_num, np.median(x))
if (x[int(n/2)]-x[int(n/2)-1]) < (BC03_wave_list_num[binning_index]-BC03_wave_list_num[binning_index-1]):
binning_size = int((BC03_wave_list_num[binning_index]-BC03_wave_list_num[binning_index-1])/(x[int(n/2)]-x[int(n/2)-1]))
x_binned,y_binned,y_err_binned=binning_spec_keep_shape_x(x,y,y_err,binning_size)
x2 = reduced_chi_square(x_binned, y_binned, y_err_binned, BC03_wave_list_num, BC03_flux_attenuated)
x2_photo = chisquare_photo(BC03_wave_list_num, BC03_flux_attenuated, redshift_1,wave_list, band_list, photometric_flux, photometric_flux_err, photometric_flux_err_mod)
# print('bin data', binning_size, x2)
else:
binning_size = int((x[int(n/2)]-x[int(n/2)-1])/(BC03_wave_list_num[binning_index]-BC03_wave_list_num[binning_index-1]))
model_wave_binned, model_flux_binned = binning_spec_keep_shape(BC03_wave_list_num, BC03_flux_attenuated,binning_size)
x2 = reduced_chi_square(x, y, y_err, model_wave_binned, model_flux_binned)
x2_photo = chisquare_photo(model_wave_binned, model_flux_binned,redshift_1,wave_list, band_list, photometric_flux, photometric_flux_err, photometric_flux_err_mod)
# print('bin model',binning_size, x2)
# print('binning size, model 3', n, (BC03_wave_list_num[binning_index]-BC03_wave_list_num[binning_index-1]), (x[int(n/2)]-x[int(n/2)-1]), binning_size)
# x2_photo = reduced_chi_square(wave_list, photometric_flux, photometric_flux_err, BC03_wave_list_num, BC03_flux_attenuated)
return 0.5*weight1*x2+0.5*weight2*x2_photo,BC03_flux_attenuated
def minimize_age_AV_vector_weighted_BC03_return_chi2_sep(X):
galaxy_age= X[0]
intrinsic_Av = X[1]
n=len(x)
age_index = find_nearest(BC03_age_list_num, galaxy_age)
age_prior = BC03_age_list_num[age_index]
AV_string = str(intrinsic_Av)
if galaxy_age == age_prior:
model3_flux = BC03_flux_array[age_index, :7125]
elif galaxy_age < age_prior:
age_interval = BC03_age_list_num[age_index+1] - BC03_age_list_num[age_index]
model3_flux = (BC03_flux_array[age_index, :7125]*(BC03_age_list_num[age_index+1]-galaxy_age) + BC03_flux_array[age_index+1, :7125]*(galaxy_age-BC03_age_list_num[age_index]))*1./age_interval
elif galaxy_age > age_prior:
age_interval = BC03_age_list_num[age_index] - BC03_age_list_num[age_index-1]
model3_flux = (BC03_flux_array[age_index, :7125]*(BC03_age_list_num[age_index]-galaxy_age) + BC03_flux_array[age_index+1, :7125]*(galaxy_age-BC03_age_list_num[age_index-1]))*1./age_interval
spectra_extinction = calzetti00(BC03_wave_list_num, intrinsic_Av, 4.05)
spectra_flux_correction = 10**(-0.4*spectra_extinction)
BC03_flux_attenuated = model3_flux*spectra_flux_correction
BC03_flux_norm = BC03_flux_attenuated[2556]
BC03_flux_attenuated = BC03_flux_attenuated/BC03_flux_norm
binning_index = find_nearest(BC03_wave_list_num, np.median(x))
if (x[int(n/2)]-x[int(n/2)-1]) < (BC03_wave_list_num[binning_index]-BC03_wave_list_num[binning_index-1]):
binning_size = int((BC03_wave_list_num[binning_index]-BC03_wave_list_num[binning_index-1])/(x[int(n/2)]-x[int(n/2)-1]))
x_binned,y_binned,y_err_binned=binning_spec_keep_shape_x(x,y,y_err,binning_size)
x2 = reduced_chi_square(x_binned, y_binned, y_err_binned, BC03_wave_list_num, BC03_flux_attenuated)
x2_photo = chisquare_photo(BC03_wave_list_num, BC03_flux_attenuated, redshift_1,wave_list, band_list, photometric_flux, photometric_flux_err, photometric_flux_err_mod)
# print('bin data', binning_size, x2)
else:
binning_size = int((x[int(n/2)]-x[int(n/2)-1])/(BC03_wave_list_num[binning_index]-BC03_wave_list_num[binning_index-1]))
model_wave_binned, model_flux_binned = binning_spec_keep_shape(BC03_wave_list_num, BC03_flux_attenuated,binning_size)
x2 = reduced_chi_square(x, y, y_err, model_wave_binned, model_flux_binned)
x2_photo = chisquare_photo(model_wave_binned, model_flux_binned,redshift_1,wave_list, band_list, photometric_flux, photometric_flux_err, photometric_flux_err_mod)
# print('bin model',binning_size, x2)
# print('binning size, model 3', n, (BC03_wave_list_num[binning_index]-BC03_wave_list_num[binning_index-1]), (x[int(n/2)]-x[int(n/2)-1]), binning_size)
# x2_photo = reduced_chi_square(wave_list, photometric_flux, photometric_flux_err, BC03_wave_list_num, BC03_flux_attenuated)
return x2,x2_photo
def find_nearest(array,value):
idx = np.searchsorted(array, value, side="left")
# print('find nearest idx searchsorted:', idx)
if np.isnan(idx):
print('find nearest',idx,value)
if idx > 0 and (idx == len(array) or math.fabs(value - array[idx-1]) < math.fabs(value - array[idx])):
return idx-1#array[idx-1]
else:
return idx#array[idx]
def all_same(items):
return all(x == items[0] for x in items)
def reduced_chi_square(data_wave,data,data_err,model_wave,model):
n=len(data_wave)
chi_square = 0
for i in range(n):
model_flux_interp = np.interp(data_wave[i], model_wave, model)
chi_square += (data[i]-model_flux_interp)**2/(data_err[i]**2)
# print('spectra chisquare processes new',i,chi_square, data_wave[i],model_flux_interp)
dof = n-2
reduced_chi_square = chi_square/dof
return reduced_chi_square
def chisquare_photo(model_wave, model_flux, redshift_1,wave_list, band_list, photometric_flux, photometric_flux_err, photometric_flux_err_mod):
"""
work in the observed frame
"""
tik = time.clock()
model_wave = model_wave*(1+redshift_1)
model_flux = model_flux
filter_array_index= np.arange(1,15)
# SNR Mask
mask_SNR3_photo = np.where(photometric_flux/photometric_flux_err>3.)
photometric_flux = photometric_flux[mask_SNR3_photo]
photometric_flux_err = photometric_flux_err[mask_SNR3_photo]
photometric_flux_err_mod = photometric_flux_err_mod[mask_SNR3_photo]
filter_array_index = filter_array_index[mask_SNR3_photo]
photometry_list = np.zeros(len(photometric_flux))
photometry_list_index = 0
# print('masked filter array index:',filter_array_index)
for i in filter_array_index:
sum_flambda_AB_K = 0
sum_transmission = 0
length = 0
filter_curve = filter_curve_list[i-1]
wave_inter = np.zeros(len(model_wave))
wave_inter[:-1] = np.diff(model_wave)
index = np.where(model_wave<filter_curve[-1,0])[0]#[0]
wave = model_wave[index]
flux = model_flux[index]
wave_inter = wave_inter[index]
index = np.where(wave>filter_curve[0,0])
wave = wave[index]
flux = flux[index]
wave_inter = wave_inter[index]
transmission = np.interp(wave, filter_curve[:,0], filter_curve[:,1])
n = len(flux)
if n!= 0 and n!=1:
for j in range(n):
try:
if all_same(wave_inter):
flambda_AB_K = flux[j]*transmission[j]
sum_flambda_AB_K += flambda_AB_K
sum_transmission += transmission[j]
length = length+1
else:
flambda_AB_K = flux[j]*transmission[j]*wave_inter[j]
sum_flambda_AB_K += flambda_AB_K
sum_transmission += transmission[j]*wave_inter[j]
length = length+1
except:
print('Error',n,transmission_index, j,wave[j],filter_curve[0,0],filter_curve[-1,0])
elif n==1:
flambda_AB_K = flux[0]*transmission[0]
sum_flambda_AB_K += flambda_AB_K*wave_inter
sum_transmission += np.sum(transmission)*wave_inter
length = length+1
if length == 0:
photometry_list[photometry_list_index]=0
else:
photometry_list[photometry_list_index] = sum_flambda_AB_K/sum_transmission
photometry_list_index += 1
chisquare_photo_list = ((photometric_flux-photometry_list)/photometric_flux_err_mod)**2
tok = time.clock()
dof = len(chisquare_photo_list)-2
reduced_chi_square_photo = np.sum(chisquare_photo_list)/dof
return reduced_chi_square_photo
# +
columns = ['ID','region','field',
'M05_age_opt','M05_AV_opt','M13_age_opt','M13_AV_opt','BC_age_opt','BC_AV_opt',\
'x2_spectra_M05_opt','x2_photo_M05_opt','x2_spectra_M13_opt','x2_photo_M13_opt','x2_spectra_BC_opt','x2_photo_BC_opt',\
'M05_age_MCMC50','M05_age_std','M05_AV_MCMC50','M05_AV_std','M13_age_MCMC50','M13_age_std','M13_AV_MCMC50','M13_AV_std','BC_age_MCMC50','BC_age_std','BC_AV_MCMC50','BC_AV_std',\
'x2_spectra_M05_MCMC50','x2_photo_M05_MCMC50','x2_spectra_M13_MCMC50','x2_photo_M13_MCMC50','x2_spectra_BC_MCMC50','x2_photo_BC_MCMC50',\
'x2_M05_opt','x2_M13_opt','x2_BC_opt','x2_M05_MCMC50','x2_M13_MCMC50','x2_BC_MCMC50',\
'model','grism_index','grism_index_AV_corr','age_opt','age_opt_std','AV_opt','AV_opt_std']
chi_square_list = pd.DataFrame(index=df.index,columns=columns)#np.zeros([len(df), 31])
chi_square_list_final = pd.DataFrame(index=df.index,columns=columns)
weight1 = 1./2.575
weight2 = 1./1.153
nsteps=3000
current_dir = '/Volumes/My Passport/TPAGB/'
outcome_dir = 'outcome/'
date='20200328_photo'
plot_dir = 'plot/'+str(date)+'_uds/'
tik = time.time()
filter_fn_list = []
filter_curve_list=[]
filter_curve_fit_list=[]
path = "/Volumes/My Passport/TAPS/filter/uds/"
import glob, os
os.chdir(path)
for i in range(1,15):
for file in glob.glob("f"+str(i)+"_*"):
print(file)
fn = path+file
filter_fn_list.append(fn)
filter_curve = np.loadtxt(fn)
filter_curve_list.append(filter_curve)
filter_f = interpolate.interp1d(filter_curve[:,0], filter_curve[:,1])
filter_curve_fit_list.append(filter_f)
tok = time.time()
print('Time reading the filter curves and without generate filter functions:',tok-tik)
# -
# ### 0 Initializing the parameters
# +
##
row=5
[ID, OneD_1, redshift_1, mag_1] = read_spectra(row)
print(row, ID)
ID_no = ID-1
redshift = df_photometry.loc[ID_no].z_spec
region = df.region[row]
intrinsic_Av = df_fast.loc[ID-1].Av
print('intrinsic Av:'+str(intrinsic_Av))
galaxy_age = 10**(df_fast.loc[ID-1].lage)/1e9
print('Galaxy age:', galaxy_age)
A_v=0.0563
c=3e10
chi_square_list.loc[row,'ID'] = float(ID)
chi_square_list.loc[row,'region'] = region
chi_square_list.loc[row,'field'] = 'uds'
# + code_folding=[21]
# Photometry
#U | CFHT | Almaini/Foucaud in prep.
# CFHT_megacam_u
u_wave = 3.86e3
u_band = 574.8/2.
u = df_photometry.loc[ID_no].f_u/((u_wave)**2)*c*1e8*3.63e-30
u_err = df_photometry.loc[ID_no].e_u/((u_wave)**2)*c*1e8*3.63e-30
# B,V,R,i,z | SXDS | Furusawa et al. (2008)
# B: 450, V: 548, Rc: 650, i’: 768, z’: 889
#use cosmos filter
B_wave = 4.50e3
B_band = 1030.5/2.
B = df_photometry.loc[ID_no].f_B/((B_wave)**2)*c*1e8*3.63e-30
B_err = df_photometry.loc[ID_no].e_B/((B_wave)**2)*c*1e8*3.63e-30
V_wave = 5.48e3
V_band = 1337.9/2.
V = df_photometry.loc[ID_no].f_V/((V_wave)**2)*c*1e8*3.63e-30
V_err = df_photometry.loc[ID_no].e_V/((V_wave)**2)*c*1e8*3.63e-30
R_wave = 6.5e3
R_band = 1143.2/2.
R = df_photometry.loc[ID_no].f_R/((R_wave)**2)*c*1e8*3.63e-30
R_err = df_photometry.loc[ID_no].e_R/((R_wave)**2)*c*1e8*3.63e-30
i_wave = 7.68e3
i_band = 1505.7/2.
i = df_photometry.loc[ID_no].f_i/((i_wave)**2)*c*1e8*3.63e-30
i_err = df_photometry.loc[ID_no].e_i/((i_wave)**2)*c*1e8*3.63e-30
z_wave = 8.89e3
z_band = 1403.5/2.
z = df_photometry.loc[ID_no].f_z/((z_wave)**2)*c*1e8*3.63e-30
z_err = df_photometry.loc[ID_no].e_z/((z_wave)**2)*c*1e8*3.63e-30
# CANDELS | Koekemoer et al. 2011, what wavelength this should take? : the same as above
F606W_wave = 5.98e3
F606W_band = 2324./2.
F606W = df_photometry.loc[ID_no].f_F606W/((F606W_wave)**2)*c*1e8*3.63e-30
F606W_err = df_photometry.loc[ID_no].e_F606W/((F606W_wave)**2)*c*1e8*3.63e-30
F814W_wave = 7.91e3
F814W_band = 1826./2.
F814W = df_photometry.loc[ID_no].f_F814W/((F814W_wave)**2)*c*1e8*3.63e-30
F814W_err = df_photometry.loc[ID_no].e_F814W/((F814W_wave)**2)*c*1e8*3.63e-30
# CANDELS | Grogin et al. 2011, Koekemoer et al. 2011|
F125W_wave = 1.250e4
F125W_band = 3005./2.
F125W = df_photometry.loc[ID_no].f_F125W/((F125W_wave)**2)*c*1e8*3.63e-30
F125W_err = df_photometry.loc[ID_no].e_F125W/((F125W_wave)**2)*c*1e8*3.63e-30
F160W_wave = 1.539e4
F160W_band = 2874./2.
F160W = df_photometry.loc[ID_no].f_F160W/((F160W_wave)**2)*c*1e8*3.63e-30 #http://www.stsci.edu/hst/wfc3/design/documents/handbooks/currentIHB/c07_ir06.html
F160W_err = df_photometry.loc[ID_no].e_F160W/((F160W_wave)**2)*c*1e8*3.63e-30
# 3D-HST | Brammer et al. 2012
F140W_wave = 13635
F140W_band = 3947./2.
F140W = df_photometry.loc[ID_no].f_F140W/((F140W_wave)**2)*c*1e8*3.63e-30 #http://svo2.cab.inta-csic.es/svo/theory/fps3/index.php?id=HST/WFC3_IR.F140W
F140W_err = df_photometry.loc[ID_no].e_F140W/((F140W_wave)**2)*c*1e8*3.63e-30
# J, H, Ks | UKIDSS /WFCAM? | Almaini et al .in prep.
# J: 1251, H:1636, K: 2206
J_wave = 1.251e4
J_band = 1590./2
J = df_photometry.loc[ID_no].f_J/J_wave**2*c*1e8*3.63e-30
J_err = df_photometry.loc[ID_no].e_J/J_wave**2*c*1e8*3.63e-30
H_wave = 1.636e4
H_band = 2920./2.
H = df_photometry.loc[ID_no].f_H/H_wave**2*c*1e8*3.63e-30
H_err = df_photometry.loc[ID_no].e_H/H_wave**2*c*1e8*3.63e-30
K_wave = 2.206e4
K_band = 3510./2.
K = df_photometry.loc[ID_no].f_K/K_wave**2*c*1e8*3.63e-30
K_err = df_photometry.loc[ID_no].e_K/K_wave**2*c*1e8*3.63e-30
wave_list = np.array([u_wave, B_wave, V_wave, R_wave, i_wave, z_wave, F606W_wave, F814W_wave, F125W_wave, F140W_wave, F160W_wave, J_wave, H_wave, K_wave])
band_list = np.array([u_band, B_band, V_band, R_band, i_band, z_band, F606W_band, F814W_band, F125W_band, F140W_band, F160W_band, J_band, H_band, K_band])
photometric_flux = np.array([u, B, V, R, i, z, F606W, F814W, F125W, F140W, F160W,J, H, K])
photometric_flux_err = np.array([u_err, B_err, V_err, R_err, i_err, z_err, F606W_err, F814W_err, F125W_err, F140W_err, F160W_err,J_err, H_err, K_err])
photometric_flux_err_mod = np.array([u_err+0.1*u, B_err+0.1*B, V_err+0.1*V, R_err+0.1*R, i_err+0.1*i, z_err+0.1*z,\
F606W_err+0.03*F606W, F814W_err+0.03*F814W, F125W_err+0.03*F125W, F140W_err+0.03*F140W, F160W_err+0.03*F160W,\
J_err+0.1*J, H_err+0.1*H, K_err+0.1*K])
# + code_folding=[0]
#-------------------------------------------------Initial Reduce the spectra ----------------------------------------------------------
print('-------------------------------------Initial fit ---------------------------------------------------------------------------------------')
[x, y, y_err, wave_list, band_list, photometric_flux, photometric_flux_err, photometric_flux_err_mod ] = \
derive_1D_spectra_Av_corrected(OneD_1, redshift_1, row, wave_list, band_list, photometric_flux, photometric_flux_err, photometric_flux_err_mod, A_v)
if redshift< 0.49:
try:
chi_square_list.loc[row,'grism_index'] = Lick_index_ratio(x,y)
except:
pass
# print(int(len(x)/2))
# print(x)
# print(x)
# print(wave_list)
# print(photometric_flux)
# print(x[int(len(x)/2)]-x[int(len(x)/2)-2])
# global x,y,y_err,wave_list,band_list,photometric_flux,photometric_flux_err
# Testing fitting a line
photo_list_for_scaling = []
photo_err_list_for_scaling = []
grism_flux_list_for_scaling = []
grism_flux_err_list_for_scaling = []
grism_wave_list_for_scaling =[]
for i in range(len(wave_list)):
if wave_list[i]-band_list[i] > x[0] and wave_list[i] + band_list[i] < x[-1]:
print(i)
scale_index = find_nearest(x, wave_list[i])
photo_list_for_scaling.append(photometric_flux[i])
photo_err_list_for_scaling.append(photometric_flux_err[i])
grism_flux_list_for_scaling.append(y[scale_index])
grism_flux_err_list_for_scaling.append(y_err[scale_index])
grism_wave_list_for_scaling.append(x[scale_index])
photo_array_for_scaling = np.array(photo_list_for_scaling)
photo_err_array_for_scaling = np.array(photo_err_list_for_scaling)
grism_flux_array_for_scaling = np.array(grism_flux_list_for_scaling)
grism_flux_err_array_for_scaling = np.array(grism_flux_err_list_for_scaling)
grism_wave_array_for_scaling = np.array(grism_wave_list_for_scaling)
print('Number of photometric points for rescaling:',len(photo_array_for_scaling))
print(np.mean(photo_array_for_scaling/grism_flux_array_for_scaling))
coeff = np.mean(photo_array_for_scaling/grism_flux_array_for_scaling)
y = y*coeff
# -
chisquare_photo(model_wave, model_flux, redshift_1,wave_list, band_list, photometric_flux, photometric_flux_err, photometric_flux_err_mod)
# + code_folding=[0]
## M05
print('____________________M05_________________________ Optimization__________________________')
X = np.array([galaxy_age, intrinsic_Av])
# X = np.array([3.43397335, 0.22173541])
bnds = ((0.01, 13.0), (0.0, 4.0))
sol = optimize.minimize(minimize_age_AV_vector_weighted, X, bounds = bnds, method='TNC')#, options = {'disp': True})
# print('Optimized weighted reduced chisqure result:', sol)
[age_prior_optimized, AV_prior_optimized] = sol.x
X = sol.x
x2_optimized = minimize_age_AV_vector_weighted(X)
x2_spec, x2_phot = minimize_age_AV_vector_weighted_return_chi2_sep(X)
chi_square_list.loc[row,'M05_age_opt'] = X[0]
chi_square_list.loc[row,'M05_AV_opt'] = X[1]
chi_square_list.loc[row,'x2_M05_opt'] = x2_optimized
chi_square_list.loc[row,'x2_spectra_M05_opt'] = x2_spec
chi_square_list.loc[row,'x2_photo_M05_opt'] = x2_phot
# + code_folding=[0]
#--- Plot
X=sol.x
n = len(x)
print(X)
fig1 = plt.figure(figsize=(20,10))
frame1 = fig1.add_axes((.1,.35,.8,.6))
plt.step(x, y, color='r',lw=3)
plt.fill_between(x,(y+y_err),(y-y_err),alpha=0.1)
plt.errorbar(wave_list, photometric_flux, xerr=band_list, yerr=photometric_flux_err_mod, color='r', fmt='o', label='photometric data', markersize='14')
model_wave =minimize_age_AV_vector_weighted_return_flux(X)[1]
model_flux =minimize_age_AV_vector_weighted_return_flux(X)[2]
model1_wave =minimize_age_AV_vector_weighted_return_flux(X)[1]
model1_flux =minimize_age_AV_vector_weighted_return_flux(X)[2]
plt.plot(model_wave, model_flux, color='k',label='TP-AGB heavy',lw=0.5)
plt.xlim([2.5e3,1.9e4])
plt.ylim([0.05, 1.1])#plt.ylim([ymin,ymax])
plt.semilogx()
plt.ylabel(r'$\rm F_{\lambda}/F_{0.55\mu m}$',fontsize=24)
plt.tick_params(axis='both', which='major', labelsize=22)
plt.legend(loc='upper right',fontsize=24)
plt.axvspan(1.06e4,1.08e4, color='gray',alpha=0.1)
plt.axvspan(1.12e4,1.14e4, color='gray',alpha=0.1)
frame2 = fig1.add_axes((.1,.2,.8,.15))
relative_spectra = np.zeros([1,n])
relative_spectra_err = np.zeros([1,n])
relative_sigma = np.zeros([1,n])
index0 = 0
for wave in x:
if y[index0]>0.25 and y[index0]<1.35:
index = find_nearest(model_wave, wave);#print index
relative_spectra[0, index0] = y[index0]/model_flux[index]
relative_spectra_err[0, index0] = y_err[index0]/model_flux[index]
relative_sigma[0, index0] = (y[index0]-model_flux[index])/y_err[index0]
index0 = index0+1
# plt.step(x[:index0], relative_spectra[0,:index0], color='r', linewidth=2)
# plt.fill_between(x[:index0],(relative_spectra[0,:index0]+relative_spectra_err[0,:index0]),\
# (relative_spectra[0,:index0]-relative_spectra_err[0,:index0]),alpha=0.1)
plt.step(x[:index0], relative_sigma[0,:index0], color='r', linewidth=2)
# print(relative_sigma[0,:index0])
index0 = 0
# relative_photo = np.zeros([1,(len(wave_list))])
for i in range(len(wave_list)):
try:
index = find_nearest(model_wave, wave_list[i])
# relative_photo[0, index0] = model_flux[index]/(photometric_flux[i])
except:
pass
plt.errorbar(wave_list[i], (photometric_flux[i]-model_flux[index])/photometric_flux_err_mod[i], xerr=band_list[i], fmt='o', color='r', markersize=12)
# plt.errorbar(wave_list[i], (photometric_flux[i])/model_flux[index], xerr=band_list[i], yerr=photometric_flux_err[i]/model_flux[index], fmt='o', color='r', markersize=16)
index0 = index0+1
plt.xlim([2.5e3,1.9e4])
plt.semilogx()
# plt.axhline(1.0, linestyle='--', linewidth=2, color='k')
# plt.ylim([0.6,1.5])
# plt.ylim([0.9,1.1])
# plt.ylim([0.7,1.45])
plt.axhline(3.0, linestyle='--', linewidth=1, color='k')
plt.axhline(-3.0, linestyle='--', linewidth=1, color='k')
plt.axhline(1.0, linestyle='--', linewidth=0.5, color='k')
plt.axhline(-1.0, linestyle='--', linewidth=0.5, color='k')
plt.ylim([-5,5])
plt.ylabel(r'$\rm (F_{\lambda,\rm data}-F_{\lambda,\rm model})/F_{\lambda,\rm err}$',fontsize=16)
plt.xlabel(r'Wavelength($\rm \AA$)', fontsize=20)
plt.axvspan(1.06e4,1.08e4, color='gray',alpha=0.1)
plt.axvspan(1.12e4,1.14e4, color='gray',alpha=0.1)
plt.tick_params(axis='both', which='major', labelsize=20)
plt.tick_params(axis='both', which='minor', labelsize=20)
# -
# ### 2 M13 model
# + code_folding=[0]
##
print('____________________M13_________________________ Optimization__________________________')
bnds = ((0.0, 13.0), (0.0, 4.0))
X = np.array([galaxy_age, intrinsic_Av])
sol_M13 = optimize.minimize(minimize_age_AV_vector_weighted_M13, X, bounds = bnds, method='TNC')#, options = {'disp': True})
# print('Optimized M13 weighted reduced chisqure result:', sol_M13)
[age_prior_optimized_M13, AV_prior_optimized_M13] = sol_M13.x
X = sol_M13.x
x2_optimized = minimize_age_AV_vector_weighted_M13(X)
x2_spec, x2_phot = minimize_age_AV_vector_weighted_M13_return_chi2_sep(X)
chi_square_list.loc[row,'M13_age_opt'] = X[0]#"{0:.2f}".format(X[0])
chi_square_list.loc[row,'M13_AV_opt'] = X[1]#"{0:.2f}".format(X[1])
chi_square_list.loc[row,'x2_M13_opt'] = x2_optimized
chi_square_list.loc[row,'x2_spectra_M13_opt'] = x2_spec
chi_square_list.loc[row,'x2_photo_M13_opt'] = x2_phot
# + code_folding=[0]
#--- Plot
X = sol_M13.x
n = len(x)
fig1 = plt.figure(figsize=(20,10))
frame1 = fig1.add_axes((.1,.35,.8,.6))
plt.step(x, y, color='r',lw=3)
plt.fill_between(x,(y+y_err),(y-y_err),alpha=0.1)
plt.errorbar(wave_list, photometric_flux, xerr=band_list, yerr=photometric_flux_err_mod, color='r', fmt='o', label='photometric data', markersize='14')
model_wave =minimize_age_AV_vector_weighted_M13_return_flux(X)[1]
model_flux =minimize_age_AV_vector_weighted_M13_return_flux(X)[2]
model2_wave =minimize_age_AV_vector_weighted_M13_return_flux(X)[1]
model2_flux =minimize_age_AV_vector_weighted_M13_return_flux(X)[2]
plt.plot(model_wave, model_flux, color='g',label='TP-AGB mild',lw=0.5)
plt.xlim([2.5e3,1.9e4])
plt.ylim([0.05, 1.1])
plt.semilogx()
plt.ylabel(r'$\rm F_{\lambda}/F_{0.55\mu m}$',fontsize=24)
plt.tick_params(axis='both', which='major', labelsize=22)
plt.legend(loc='upper right',fontsize=24)
plt.axvspan(1.06e4,1.08e4, color='gray',alpha=0.1)
plt.axvspan(1.12e4,1.14e4, color='gray',alpha=0.1)
frame2 = fig1.add_axes((.1,.2,.8,.15))
relative_spectra = np.zeros([1,n])
relative_spectra_err = np.zeros([1,n])
relative_sigma = np.zeros([1,n])
index0 = 0
for wave in x:
if y[index0]>0.25 and y[index0]<1.35:
index = find_nearest(model_wave, wave);#print index
relative_spectra[0, index0] = y[index0]/model_flux[index]
relative_spectra_err[0, index0] = y_err[index0]/model_flux[index]
relative_sigma[0, index0] = (y[index0]-model_flux[index])/y_err[index0]
index0 = index0+1
# plt.step(x[:index0], relative_spectra[0,:index0], color='r', linewidth=2)
# plt.fill_between(x[:index0],(relative_spectra[0,:index0]+relative_spectra_err[0,:index0]),\
# (relative_spectra[0,:index0]-relative_spectra_err[0,:index0]),alpha=0.1)
plt.step(x[:index0], relative_sigma[0,:index0], color='r', linewidth=2)
# print(relative_sigma[0,:index0])
index0 = 0
# relative_photo = np.zeros([1,(len(wave_list))])
for i in range(len(wave_list)):
try:
index = find_nearest(model_wave, wave_list[i])
# relative_photo[0, index0] = model_flux[index]/(photometric_flux[i])
except:
pass
plt.errorbar(wave_list[i], (photometric_flux[i]-model_flux[index])/photometric_flux_err_mod[i], xerr=band_list[i], fmt='o', color='r', markersize=12)
# plt.errorbar(wave_list[i], (photometric_flux[i])/model_flux[index], xerr=band_list[i], yerr=photometric_flux_err[i]/model_flux[index], fmt='o', color='r', markersize=16)
index0 = index0+1
plt.xlim([2.5e3,1.9e4])
plt.semilogx()
# plt.axhline(1.0, linestyle='--', linewidth=2, color='k')
plt.axvspan(1.06e4,1.08e4, color='gray',alpha=0.1)
plt.axvspan(1.12e4,1.14e4, color='gray',alpha=0.1)
# plt.ylim([0.75,1.5])
# plt.ylim([0.9,1.1])
# plt.ylim([0.7,1.45])
plt.axhline(3.0, linestyle='--', linewidth=1, color='k')
plt.axhline(-3.0, linestyle='--', linewidth=1, color='k')
plt.axhline(1.0, linestyle='--', linewidth=0.5, color='k')
plt.axhline(-1.0, linestyle='--', linewidth=0.5, color='k')
plt.ylim([-5,5])
plt.ylabel(r'$\rm (F_{\lambda,\rm data}-F_{\lambda,\rm model})/F_{\lambda,\rm err}$',fontsize=16)
plt.tick_params(axis='both', which='major', labelsize=20)
plt.tick_params(axis='both', which='minor', labelsize=20)
plt.xlabel(r'Wavelength($\rm \AA$)', fontsize=20)
# -
with Pool() as pool:
ndim, nwalkers = 2, 10
tik = time.clock()
p0 = [sol_M13.x + 4.*np.random.rand(ndim) for i in range(nwalkers)]
sampler = emcee.EnsembleSampler(nwalkers, ndim, lg_minimize_age_AV_vector_weighted_M13, pool=pool)
sampler.run_mcmc(p0, nsteps,progress=True)
samples = sampler.chain[:, 500:, :].reshape((-1,ndim))
samples = samples[(samples[:,0] > age_prior_optimized_M13*0.1) & (samples[:,0] < age_prior_optimized_M13*2.0) & (samples[:,1] < AV_prior_optimized_M13*3.0)]
tok = time.clock()
multi_time = tok-tik
print("Multiprocessing took {0:.1f} seconds".format(multi_time))
print('Time to run M13 MCMC:'+str(tok-tik))
# + code_folding=[0]
if samples.size > 1e3 :
value2=np.percentile(samples, 50, axis=0)
[std_age_prior_optimized_M13, std_AV_prior_optimized_M13] = np.std(samples, axis=0)
plt.figure(figsize=(32,32),dpi=100)
fig = corner.corner(samples,
labels=["age(Gyr)", r"$\rm A_V$"],
levels=(1-np.exp(-0.5),),
truths=[age_prior_optimized_M13, AV_prior_optimized_M13],
show_titles=True,title_kwargs={'fontsize':12},
quantiles=(0.16,0.5, 0.84))
axes = np.array(fig.axes).reshape((ndim, ndim))
for i in range(ndim):
ax = axes[i, i]
ax.axvline(X[i], color="g")
# Loop over the histograms
for i in range(ndim):
ax = axes[i, i]
ax.axvline(X[i], color="g")
ax.axvline(value2[i],color='r')
# Loop over the histograms
for yi in range(ndim):
for xi in range(yi):
ax = axes[yi, xi]
ax.axvline(X[xi], color="g")
ax.axvline(value2[xi], color="r")
ax.axhline(X[yi], color="g")
ax.axhline(value2[yi], color="r")
ax.plot(X[xi], X[yi], "sg")
ax.plot(value2[xi],value2[yi],'sr')
plt.rcParams.update({'font.size': 12})
# + code_folding=[0]
#--- Plot
X = np.percentile(samples, 50, axis=0)
x2_optimized = minimize_age_AV_vector_weighted_M13(X)
x2_spec, x2_phot = minimize_age_AV_vector_weighted_M13_return_chi2_sep(X)
chi_square_list.loc[row,'M13_age_MCMC50'] = X[0]#"{0:.2f}".format(X[0])
chi_square_list.loc[row,'M13_AV_MCMC50'] = X[1]#"{0:.2f}".format(X[1])
chi_square_list.loc[row,'x2_M13_MCMC50'] = x2_optimized
chi_square_list.loc[row,'x2_spectra_M13_MCMC50'] = x2_spec
chi_square_list.loc[row,'x2_photo_M13_MCMC50'] = x2_phot
chi_square_list.loc[row,'M13_age_std'] = np.std(samples, axis=0)[0]#"{0:.2f}".format(np.std(samples, axis=0)[0])
chi_square_list.loc[row,'M13_AV_std'] = np.std(samples, axis=0)[1]#"{0:.2f}".format(np.std(samples, axis=0)[1])
n = len(x)
fig1 = plt.figure(figsize=(20,10))
frame1 = fig1.add_axes((.1,.35,.8,.6))
plt.step(x, y, color='r',lw=3)
plt.fill_between(x,(y+y_err),(y-y_err),alpha=0.1)
plt.errorbar(wave_list, photometric_flux, xerr=band_list, yerr=photometric_flux_err_mod, color='r', fmt='o', label='photometric data', markersize='14')
model_wave =minimize_age_AV_vector_weighted_M13_return_flux(X)[1]
model_flux =minimize_age_AV_vector_weighted_M13_return_flux(X)[2]
plt.plot(model_wave, model_flux, color='g',label='TP-AGB mild',lw=0.5)
plt.xlim([2.5e3,1.9e4])
plt.ylim([0.05, 1.1])
plt.semilogx()
plt.ylabel(r'$\rm F_{\lambda}/F_{0.55\mu m}$',fontsize=24)
plt.tick_params(axis='both', which='major', labelsize=22)
plt.legend(loc='upper right',fontsize=24)
plt.axvspan(1.06e4,1.08e4, color='gray',alpha=0.1)
plt.axvspan(1.12e4,1.14e4, color='gray',alpha=0.1)
frame2 = fig1.add_axes((.1,.2,.8,.15))
relative_spectra = np.zeros([1,n])
relative_spectra_err = np.zeros([1,n])
relative_sigma = np.zeros([1,n])
index0 = 0
for wave in x:
if y[index0]>0.25 and y[index0]<1.35:
index = find_nearest(model_wave, wave);#print index
relative_spectra[0, index0] = y[index0]/model_flux[index]
relative_spectra_err[0, index0] = y_err[index0]/model_flux[index]
relative_sigma[0, index0] = (y[index0]-model_flux[index])/y_err[index0]
index0 = index0+1
# plt.step(x[:index0], relative_spectra[0,:index0], color='r', linewidth=2)
# plt.fill_between(x[:index0],(relative_spectra[0,:index0]+relative_spectra_err[0,:index0]),\
# (relative_spectra[0,:index0]-relative_spectra_err[0,:index0]),alpha=0.1)
plt.step(x[:index0], relative_sigma[0,:index0], color='r', linewidth=2)
# print(relative_sigma[0,:index0])
index0 = 0
# relative_photo = np.zeros([1,(len(wave_list))])
for i in range(len(wave_list)):
try:
index = find_nearest(model_wave, wave_list[i])
# relative_photo[0, index0] = model_flux[index]/(photometric_flux[i])
except:
pass
plt.errorbar(wave_list[i], (photometric_flux[i]-model_flux[index])/photometric_flux_err_mod[i], xerr=band_list[i], fmt='o', color='r', markersize=12)
# plt.errorbar(wave_list[i], (photometric_flux[i])/model_flux[index], xerr=band_list[i], yerr=photometric_flux_err[i]/model_flux[index], fmt='o', color='r', markersize=16)
index0 = index0+1
plt.xlim([2.5e3,1.9e4])
plt.semilogx()
# plt.axhline(1.0, linestyle='--', linewidth=2, color='k')
plt.axvspan(1.06e4,1.08e4, color='gray',alpha=0.1)
plt.axvspan(1.12e4,1.14e4, color='gray',alpha=0.1)
# plt.ylim([0.75,1.5])
# plt.ylim([0.9,1.1])
# plt.ylim([0.7,1.45])
plt.axhline(3.0, linestyle='--', linewidth=1, color='k')
plt.axhline(-3.0, linestyle='--', linewidth=1, color='k')
plt.axhline(1.0, linestyle='--', linewidth=0.5, color='k')
plt.axhline(-1.0, linestyle='--', linewidth=0.5, color='k')
plt.ylim([-5,5])
plt.ylabel(r'$\rm (F_{\lambda,\rm data}-F_{\lambda,\rm model})/F_{\lambda,\rm err}$',fontsize=16)
plt.xlabel(r'Wavelength($\rm \AA$)', fontsize=20)
plt.tick_params(axis='both', which='major', labelsize=20)
plt.tick_params(axis='both', which='minor', labelsize=20)
# -
# ### 3 BC03 model
# + code_folding=[0]
##
print('____________________BC03_________________________ Optimization__________________________')
X = np.array([galaxy_age, intrinsic_Av])
bnds = ((0.0, 13.0), (0.0, 4.0))
sol_BC03 = optimize.minimize(minimize_age_AV_vector_weighted_BC03, X, bounds = bnds, method='TNC', options = {'disp': True})
print('Optimized BC03 weighted reduced chisqure result:', sol_BC03)
[age_prior_optimized_BC03, AV_prior_optimized_BC03] = sol_BC03.x
X = sol_BC03.x
x2_optimized = minimize_age_AV_vector_weighted_BC03(X)
x2_spec, x2_phot = minimize_age_AV_vector_weighted_BC03_return_chi2_sep(X)
chi_square_list.loc[row,'BC_age_opt'] = X[0]#"{0:.2f}".format(X[0])
chi_square_list.loc[row,'BC_AV_opt'] = X[1]#"{0:.2f}".format(X[1])
chi_square_list.loc[row,'x2_BC_opt'] = x2_optimized
chi_square_list.loc[row,'x2_spectra_BC_opt'] = x2_spec
chi_square_list.loc[row,'x2_photo_BC_opt'] = x2_phot
# + code_folding=[0]
#--- Plot
X = sol_BC03.x
n = len(x)
fig1 = plt.figure(figsize=(20,10))
frame1 = fig1.add_axes((.1,.35,.8,.6))
plt.step(x, y, color='r',lw=3)
plt.fill_between(x,(y+y_err),(y-y_err),alpha=0.1)
plt.errorbar(wave_list, photometric_flux, xerr=band_list, yerr=photometric_flux_err_mod, color='r', fmt='o', label='photometric data', markersize='14')
BC03_flux_attenuated = minimize_age_AV_vector_weighted_BC03_mod_no_weight_return_flux(X)[1]
plt.plot(BC03_wave_list_num, BC03_flux_attenuated, color='orange',label='TP-AGB light',lw=0.5)
model_wave = BC03_wave_list_num
model_flux = BC03_flux_attenuated
model3_wave = BC03_wave_list_num
model3_flux = BC03_flux_attenuated
plt.xlim([2.5e3,1.9e4])
plt.ylim([0.05, 1.1])
# plt.ylim([0.7,1.45])
plt.semilogx()
plt.ylabel(r'$\rm F_{\lambda}/F_{0.55\mu m}$',fontsize=24)
plt.tick_params(axis='both', which='major', labelsize=22)
plt.legend(loc='upper right',fontsize=24)
plt.axvspan(1.06e4,1.08e4, color='gray',alpha=0.1)
plt.axvspan(1.12e4,1.14e4, color='gray',alpha=0.1)
frame2 = fig1.add_axes((.1,.2,.8,.15))
relative_spectra = np.zeros([1,n])
relative_spectra_err = np.zeros([1,n])
relative_sigma = np.zeros([1,n])
index0 = 0
for wave in x:
if y[index0]>0.25 and y[index0]<1.35:
index = find_nearest(model_wave, wave);#print index
relative_spectra[0, index0] = y[index0]/model_flux[index]
relative_spectra_err[0, index0] = y_err[index0]/model_flux[index]
relative_sigma[0, index0] = (y[index0]-model_flux[index])/y_err[index0]
index0 = index0+1
# plt.step(x[:index0], relative_spectra[0,:index0], color='r', linewidth=2)
# plt.fill_between(x[:index0],(relative_spectra[0,:index0]+relative_spectra_err[0,:index0]),\
# (relative_spectra[0,:index0]-relative_spectra_err[0,:index0]),alpha=0.1)
plt.step(x[:index0], relative_sigma[0,:index0], color='r', linewidth=2)
# print(relative_sigma[0,:index0])
index0 = 0
# relative_photo = np.zeros([1,(len(wave_list))])
for i in range(len(wave_list)):
try:
index = find_nearest(model_wave, wave_list[i])
# relative_photo[0, index0] = model_flux[index]/(photometric_flux[i])
except:
pass
plt.errorbar(wave_list[i], (photometric_flux[i]-model_flux[index])/photometric_flux_err_mod[i], xerr=band_list[i], fmt='o', color='r', markersize=12)
# plt.errorbar(wave_list[i], (photometric_flux[i])/model_flux[index], xerr=band_list[i], yerr=photometric_flux_err[i]/model_flux[index], fmt='o', color='r', markersize=16)
index0 = index0+1
plt.xlim([2.5e3,1.9e4])
plt.semilogx()
# plt.axhline(1.0, linestyle='--', linewidth=2, color='k')
# plt.ylim([0.6,1.5])
# plt.ylim([0.9,1.1])
# plt.ylim([0.7,1.45])
plt.axhline(3.0, linestyle='--', linewidth=1, color='k')
plt.axhline(-3.0, linestyle='--', linewidth=1, color='k')
plt.axhline(1.0, linestyle='--', linewidth=0.5, color='k')
plt.axhline(-1.0, linestyle='--', linewidth=0.5, color='k')
plt.ylim([-5,5])
plt.ylabel(r'$\rm (F_{\lambda,\rm data}-F_{\lambda,\rm model})/F_{\lambda,\rm err}$',fontsize=16)
plt.xlabel(r'Wavelength($\rm \AA$)', fontsize=20)
plt.axvspan(1.06e4,1.08e4, color='gray',alpha=0.1)
plt.axvspan(1.12e4,1.14e4, color='gray',alpha=0.1)
plt.tick_params(axis='both', which='major', labelsize=20)
plt.tick_params(axis='both', which='minor', labelsize=20)
# + code_folding=[0]
with Pool() as pool:
ndim, nwalkers = 2, 10
tik = time.clock()
p0 = [sol_BC03.x + 4.*np.random.rand(ndim) for i in range(nwalkers)]
sampler = emcee.EnsembleSampler(nwalkers, ndim, lg_minimize_age_AV_vector_weighted_BC03, pool=pool)
sampler.run_mcmc(p0, nsteps, progress=True)
samples = sampler.chain[:, 500:, :].reshape((-1,ndim))
samples = samples[(samples[:,0] > age_prior_optimized_BC03*0.1) &
(samples[:,0] < age_prior_optimized_BC03*2.0) &
(samples[:,1] < AV_prior_optimized_BC03*3.0)]
tok = time.clock()
multi_time = tok-tik
print("Multiprocessing took {0:.1f} seconds".format(multi_time))
print('Time to run BC03 MCMC:'+str(tok-tik))
# + code_folding=[0]
if samples.size > 1e3:
value2=np.percentile(samples,50,axis=0)
[std_age_prior_optimized_BC03, std_AV_prior_optimized_BC03] = np.std(samples, axis=0)
plt.figure(figsize=(32,32),dpi=100)
fig = corner.corner(samples,
labels=["age(Gyr)", r"$\rm A_V$"],\
truths=[age_prior_optimized_BC03, AV_prior_optimized_BC03],\
levels = (1-np.exp(-0.5),),\
show_titles=True,title_kwargs={'fontsize':12},
quantiles=(0.16,0.5, 0.84))
axes = np.array(fig.axes).reshape((ndim, ndim))
for i in range(ndim):
ax = axes[i, i]
ax.axvline(X[i], color="g")
ax.axvline(value2[i],color='r')
# Loop over the histograms
for yi in range(ndim):
for xi in range(yi):
ax = axes[yi, xi]
ax.axvline(X[xi], color="g")
ax.axvline(value2[xi], color="r")
ax.axhline(X[yi], color="g")
ax.axhline(value2[yi], color="r")
ax.plot(X[xi], X[yi], "sg")
ax.plot(value2[xi],value2[yi],'sr')
plt.rcParams.update({'font.size': 12})
# + code_folding=[0]
#--- Plot
X = np.percentile(samples, 50, axis=0)
x2_optimized = minimize_age_AV_vector_weighted_BC03(X)
x2_spec, x2_phot = minimize_age_AV_vector_weighted_BC03_return_chi2_sep(X)
chi_square_list.loc[row,'BC_age_MCMC50'] = X[0]#"{0:.2f}".format(X[0])
chi_square_list.loc[row,'BC_AV_MCMC50'] =X[1] #"{0:.2f}".format(X[1])
chi_square_list.loc[row,'x2_BC_MCMC50'] = x2_optimized
chi_square_list.loc[row,'x2_spectra_BC_MCMC50'] = x2_spec
chi_square_list.loc[row,'x2_photo_BC_MCMC50'] = x2_phot
chi_square_list.loc[row,'BC_age_std'] = np.std(samples, axis=0)[0] #"{0:.2f}".format(np.std(samples, axis=0)[0])
chi_square_list.loc[row,'BC_AV_std'] = np.std(samples, axis=0)[1]#"{0:.2f}".format(np.std(samples, axis=0)[1])
n = len(x)
fig1 = plt.figure(figsize=(20,10))
frame1 = fig1.add_axes((.1,.35,.8,.6))
plt.step(x, y, color='r',lw=3)
plt.fill_between(x,(y+y_err),(y-y_err),alpha=0.1)
plt.errorbar(wave_list, photometric_flux, xerr=band_list, yerr=photometric_flux_err_mod, color='r', fmt='o', label='photometric data', markersize='14')
BC03_flux_attenuated = minimize_age_AV_vector_weighted_BC03_mod_no_weight_return_flux(X)[1]
plt.plot(BC03_wave_list_num, BC03_flux_attenuated, color='orange',label='TP-AGB light',lw=0.5)
model_wave = BC03_wave_list_num
model_flux = BC03_flux_attenuated
plt.xlim([2.5e3,1.9e4])
plt.ylim([0.05, 1.1])
plt.semilogx()
plt.ylabel(r'$\rm F_{\lambda}/F_{0.55\mu m}$',fontsize=24)
plt.tick_params(axis='both', which='major', labelsize=22)
plt.legend(loc='upper right',fontsize=24)
plt.axvspan(1.06e4,1.08e4, color='gray',alpha=0.1)
plt.axvspan(1.12e4,1.14e4, color='gray',alpha=0.1)
frame2 = fig1.add_axes((.1,.2,.8,.15))
relative_spectra = np.zeros([1,n])
relative_spectra_err = np.zeros([1,n])
relative_sigma = np.zeros([1,n])
index0 = 0
for wave in x:
if y[index0]>0.25 and y[index0]<1.35:
index = find_nearest(model_wave, wave);#print index
relative_spectra[0, index0] = y[index0]/model_flux[index]
relative_spectra_err[0, index0] = y_err[index0]/model_flux[index]
relative_sigma[0, index0] = (y[index0]-model_flux[index])/y_err[index0]
index0 = index0+1
# plt.step(x[:index0], relative_spectra[0,:index0], color='r', linewidth=2)
# plt.fill_between(x[:index0],(relative_spectra[0,:index0]+relative_spectra_err[0,:index0]),\
# (relative_spectra[0,:index0]-relative_spectra_err[0,:index0]),alpha=0.1)
plt.step(x[:index0], relative_sigma[0,:index0], color='r', linewidth=2)
# print(relative_sigma[0,:index0])
index0 = 0
# relative_photo = np.zeros([1,(len(wave_list))])
for i in range(len(wave_list)):
try:
index = find_nearest(model_wave, wave_list[i])
# relative_photo[0, index0] = model_flux[index]/(photometric_flux[i])
except:
pass
plt.errorbar(wave_list[i], (photometric_flux[i]-model_flux[index])/photometric_flux_err_mod[i], xerr=band_list[i], fmt='o', color='r', markersize=12)
# plt.errorbar(wave_list[i], (photometric_flux[i])/model_flux[index], xerr=band_list[i], yerr=photometric_flux_err[i]/model_flux[index], fmt='o', color='r', markersize=16)
index0 = index0+1
plt.xlim([2.5e3,1.9e4])
plt.semilogx()
# plt.axhline(1.0, linestyle='--', linewidth=2, color='k')
# plt.ylim([0.6,1.5])
# plt.ylim([0.9,1.1])
# plt.ylim([0.7,1.45])
plt.axhline(3.0, linestyle='--', linewidth=1, color='k')
plt.axhline(-3.0, linestyle='--', linewidth=1, color='k')
plt.axhline(1.0, linestyle='--', linewidth=0.5, color='k')
plt.axhline(-1.0, linestyle='--', linewidth=0.5, color='k')
plt.ylim([-5,5])
plt.ylabel(r'$\rm (F_{\lambda,\rm data}-F_{\lambda,\rm model})/F_{\lambda,\rm err}$',fontsize=16)
plt.xlabel(r'Wavelength($\rm \AA$)', fontsize=20)
plt.axvspan(1.06e4,1.08e4, color='gray',alpha=0.1)
plt.axvspan(1.12e4,1.14e4, color='gray',alpha=0.1)
plt.tick_params(axis='both', which='major', labelsize=20)
plt.tick_params(axis='both', which='minor', labelsize=20)
# -
# ### 4 Testing the filter sets
print(redshift_1)
filter_fn_list = []
path = "/Volumes/My Passport/TAPS/filter/uds/"
import glob, os
os.chdir(path)
for i in range(1,15):
for file in glob.glob("f"+str(i)+"_*"):
print(file)
fn = path+file
filter_fn_list.append(fn)
# filter_fn_list[0]
filter_curve = np.loadtxt(fn)
# print(filter_curve.size)#[:,0]
plt.plot(filter_curve[:,0],filter_curve[:,1])
model_wave = model3_wave*(1+redshift_1)
model_flux = model3_flux
def all_same(items):
return all(x == items[0] for x in items)
# + code_folding=[13]
def all_same(items):
return all(x == items[0] for x in items)
plt.figure(figsize=(12,6),dpi=300)
plt.plot(model_wave, model_flux, color='orange',lw=0.5)
photometry_list = np.zeros(len(wave_list))
plt.xlim([3.e3,2.9e4])
plt.ylim([-0.05, 1.1])
plt.semilogx()
plt.step(x*(1+redshift_1), y, color='r',lw=3)
plt.fill_between(x*(1+redshift_1),(y+y_err),(y-y_err),alpha=0.1)
plt.ylabel(r'$\rm F_{\lambda}/F_{0.55\mu m}$',fontsize=24)
for i in range(1,15):
for file in glob.glob("f"+str(i)+"_*"):
print(i,file)
fn = path+file
filter_fn_list.append(fn)
filter_curve = np.loadtxt(fn)
print(filter_curve.size)#[:,0]
sum_flambda_AB_K = 0
sum_transmission = 0
length = 0
for j in range(len(filter_curve)-1):
wave_inter = np.zeros(len(model_wave))
wave_inter[:-1] = np.diff(model_wave)
index = np.where(model_wave<filter_curve[j+1,0])[0]#[0]
wave = model_wave[index]
flux = model_flux[index]
wave_inter = wave_inter[index]
index = np.where(wave>filter_curve[j,0])
wave = wave[index]
flux = flux[index]
wave_inter = wave_inter[index]
n = len(flux)
if n!= 0 and n!=1:
try:
transmission = np.interp(wave, filter_curve[j:j+2,0], filter_curve[j:j+2,1])
except:
print('Error')
# Checking if all spectral elements are the same
if all_same(wave_inter):
flambda_AB_K = np.sum(flux*transmission)
sum_flambda_AB_K += flambda_AB_K
sum_transmission += np.sum(transmission)
length = length+1
else:
flambda_AB_K = np.sum(flux*transmission*wave_inter)
sum_flambda_AB_K += flambda_AB_K
sum_transmission += np.sum(transmission*wave_inter)
length = length+1
elif n==1:
transmission = np.interp(wave, filter_curve[j:j+2,0], filter_curve[j:j+2,1])
flambda_AB_K = flux[0]*transmission[0]
sum_flambda_AB_K += flambda_AB_K*wave_inter
sum_transmission += np.sum(transmission)*wave_inter#/len(transmission)#np.trapz(transmission, x=wave)
length = length+1
if length == 0:
photometry_list[i-1]=0
else:
photometry_list[i-1] = sum_flambda_AB_K/sum_transmission
print(wave_list[i-1]*(1+redshift_1), photometry_list[i-1], sum_flambda_AB_K, sum_transmission,length)#, wave[int(n/2)])
plt.errorbar(wave_list[i-1]*(1+redshift_1),photometry_list[i-1],\
xerr=band_list[i-1], color='g', fmt='o', markersize=14)
plt.errorbar(wave_list[i-1]*(1+redshift_1), photometric_flux[i-1],\
xerr=band_list[i-1], yerr=photometric_flux_err_mod[i-1], color='r', fmt='o', label='photometric data', markersize='14')
chisquare_photo_list = ((photometric_flux-photometry_list)/photometric_flux_err_mod)**2
# -
chisquare_photo_list = ((photometric_flux-photometry_list)/photometric_flux_err_mod)**2
print(chisquare_photo_list)
print(np.sum(chisquare_photo_list))
chi2_M05 = chisquare_photo(model1_wave, model1_flux, redshift_1,wave_list, band_list, photometric_flux, photometric_flux_err, photometric_flux_err_mod)
chi2_M13 = chisquare_photo(model2_wave, model2_flux, redshift_1,wave_list, band_list, photometric_flux, photometric_flux_err, photometric_flux_err_mod)
chi2_BC = chisquare_photo(model3_wave, model3_flux, redshift_1,wave_list, band_list, photometric_flux, photometric_flux_err, photometric_flux_err_mod)
| examples/5. UDS_reduce_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# %matplotlib inline
from __future__ import division
import numpy as np
from sklearn.metrics import confusion_matrix
import matplotlib.pyplot as plt
from utils import draw_in_row
from utils import NormalDistribution
from utils import plot_confusion_matrix
from utils import MixtureGaussians
plt.rcParams['figure.figsize'] = (10,6)
np.random.seed(42)
# -
# # Cautious classification
#
# Performing cautious classification is a method to predict the classes considering
# the option of abstaining to predict any instance, given that the instance seems
# ambiguous. In that case, the missclassification could be worst than abstaining
# giving the possibility of performing a further analysis of the instance.
#
# For example, lets assume that we have two classes with the following description
# +
norm_f1 = NormalDistribution(mu=0.9, sigma=0.2)
norm_f2 = NormalDistribution(mu=1.1, sigma=0.25)
p_f1 = 0.4
p_f2 = 0.6
x_min = np.min([norm_f1.mu-4*norm_f1.sigma, norm_f2.mu-4*norm_f2.sigma])
x_max = np.max([norm_f1.mu+4*norm_f1.sigma, norm_f2.mu+4*norm_f2.sigma])
x_lin = np.linspace(x_min, x_max, 100)
p_x_g_f1 = norm_f1.pdf(x_lin)
p_x_g_f2 = norm_f2.pdf(x_lin)
plt.plot(x_lin, p_x_g_f1, color='yellowgreen', label='$p(x|f_1)$', linewidth=3)
plt.plot(x_lin, p_x_g_f2, color='orange', label='$p(x|f_2)$', linewidth=3)
plt.ylabel('Class densities')
plt.legend()
plt.xlim([x_min, x_max])
plt.grid(True)
# -
# Then we can compute the posterior probabilities for both classes using Bayes' theorem.
# +
p_x = p_x_g_f1*p_f1 + p_x_g_f2*p_f2
p_f1_g_f_x = (p_x_g_f1*p_f1)/p_x
p_f2_g_f_x = (p_x_g_f2*p_f2)/p_x
plt.plot(x_lin, p_f1_g_f_x, color='yellowgreen', label='$p(f_1|f, x)$', linewidth=3)
plt.plot(x_lin, p_f2_g_f_x, color='orange', label='$p(f_2|f, x)$', linewidth=3)
plt.legend()
plt.xlim([x_min, x_max])
plt.grid(True)
# -
# # Chow's rule
#
# Chow's rule states that it is possible to make cautious classifications by
# rejecting the predictions for which the posterior probabilites for the instance
# $x$ are
#
# $$ \forall_i,p(y=i|x) < \theta $$
#
# We can visualize this with the posterior probabilities and we can see that this
# corresponds to reject the regions of the input space where the posterior probabilites
# are really similar.
# +
p_x = p_x_g_f1*p_f1 + p_x_g_f2*p_f2
p_f1_g_f_x = (p_x_g_f1*p_f1)/p_x
p_f2_g_f_x = (p_x_g_f2*p_f2)/p_x
theta = 0.6
def plot_predictions(x, f1, f2, reject):
plt.plot(x, f1, color='yellowgreen', label=r'$p(f_1|x)$', linewidth=3)
plt.plot(x, f2, color='orange', label=r'$p(f_2|x)$', linewidth=3)
plt.plot(x, reject, color='red', label=r'$reject$', linewidth=3)
plt.legend()
plt.xlim([x.min(), x.max()])
plt.grid(True)
plt.title("Prediction is the argmax of the next three")
reject = np.ones_like(x_lin)*theta
plot_predictions(x_lin, p_f1_g_f_x, p_f2_g_f_x, reject)
# -
# # <NAME> and <NAME>
#
# Another option is to have one thershold $\theta$ per class. Then, if the posterior probability
# of one class is lower than its corresponding threshold then the models abstains.
# +
theta_f1 = 0.6
theta_f2 = 0.9
thetas = np.array([theta_f1, theta_f2])
def plot_original_thresholds(x, f1, f2, theta1, theta2):
plt.plot(x, f1, color='yellowgreen', label=r'$p(f_1|x)$', linewidth=3)
plt.plot(x, f2, color='orange', label=r'$p(f_2|x)$', linewidth=3)
plt.plot(x, np.ones_like(x)*theta1, '--', color='yellowgreen', label=r"$\theta_1 = {}$".format(theta1), linewidth=3)
plt.plot(x, np.ones_like(x)*theta2, '--', color='orange', label=r"$\theta_2 = {}$".format(theta2), linewidth=3)
plt.legend()
plt.xlim([x.min(), x.max()])
plt.grid(True)
plt.subplot(2,1,1)
plot_original_thresholds(x_lin, p_f1_g_f_x, p_f2_g_f_x, theta_f1, theta_f2)
c = np.argmax(np.vstack((p_f1_g_f_x, p_f2_g_f_x)), axis=0)
reject = thetas[c]
plt.subplot(2,1,2)
plot_predictions(x_lin, p_f1_g_f_x, p_f2_g_f_x, reject)
# -
# However with imbalanced classes this method can be problematic as some of the classes could have really low probabilities. For that reason the same authors proposed the use of scaled probabilities with respect to their corresponding thresholds.
#
# If $\exists p_i, p_i \ge \theta_i$ then class = argmax$_i(p_i / \theta_i)$
#
# Else class = abstain
#
# ## TODO modify this example
# +
plt.subplot(2,1,1)
plot_original_thresholds(x_lin, p_f1_g_f_x, p_f2_g_f_x, theta_f1, theta_f2)
reject = np.ones_like(x_lin)
plt.subplot(2,1,2)
plot_predictions(x_lin, p_f1_g_f_x/theta_f1, p_f2_g_f_x/theta_f2, reject)
# -
# However, the previous approach is merging two different aspects in the thresholds:
#
# 1. The relevance of each class, given the differences between the thresholds
# 2. The degree of abstention, given by the size of the thresholds
#
# Using the previous method, in order to decrease the abstention all the thresholds need to
# be decreased in the same proportion. In order to avoid that, it is possible to make the
# class biases to sum to one and then separate both aspects.
#
# Define the Class Bias $K = {k_i}$ for $i \in {1,\dots |C|}$, where $\sum_i k_i = 1$ and a window
# size $w$, $0\ge w \ge 1$.
# +
k_f1 = 0.2
k_f2 = 0.8
w = 0.5
theta_f1 = (1-k_f1)*w + k_f1
theta_f2 = (1-k_f2)*w + k_f2
thetas = np.array([theta_f1, theta_f2])
plt.subplot(2,1,1)
plot_original_thresholds(x_lin, p_f1_g_f_x, p_f2_g_f_x, theta_f1, theta_f2)
reject = np.ones_like(x_lin)
plt.subplot(2,1,2)
plot_predictions(x_lin, p_f1_g_f_x/theta_f1, p_f2_g_f_x/theta_f2, reject)
# -
# ## Background Check
#
# It is possible to solve the same task by assuming that there exist a _background_ class with the same distribution as the _foreground_ but with a different prior probability. In this case, regions where the probabilities for all the _foreground_ classes are lower than the _background_ probability is considered as an uncertain region and can be rejected.
#
# We achieve this behaviour by using the affine inductive bias
#
# $$ q_b(x) = (1-q_f(x))\mu(0) + q_f(x)\mu(1)$$
#
# were in this particular case $\mu(0) = 0$ and $\mu(1) = \theta$.
#
# Then
#
# $$ q_b(x) = q_f(x)\theta $$
#
# This means that the ratio between the two distributions is qual to the inverse of the threshold $\theta$
#
# $$ \frac{q_f(x)}{q_b(x)} = \theta^{-1} = r(x) $$
#
# thus
#
# $$
# p(b|x) = \frac{1}{1+\theta^{-1}} \\
# p(f_c|x) = \frac{p(f_c|f,x)\theta^{-1}}{1+\theta^{-1}}
# $$
#
# Because of this parametrization, the density of the _foreground_ and the _background_ are not important as they cancel each other and what it remains is their ratio. For that reason in the next examples we arebitrarly assigned to the _foreground_ a fixed constant distribution with the value 1.
# +
p_f_x = 1
mu0 = 0
mu1 = 0.6
theta = mu1
p_b_x = np.ones_like(x_lin)*1/(1 + theta**-1)
p_f1_g_x = (p_f1_g_f_x*theta**-1)/(1+theta**-1)
p_f2_g_x = (p_f2_g_f_x*theta**-1)/(1+theta**-1)
plot_predictions(x_lin, p_f1_g_x, p_f2_g_x, p_b_x)
plt.ylim([0,1])
# -
# The same can be done with multiple thresholds as explained by <NAME> and <NAME> by setting a __background check__ per class.
#
#
# $$
# q_{bf_1}(x) = q_{f_1}(x)\theta_1 \\
# q_{bf_2}(x) = q_{f_2}(x)\theta_2
# $$
#
# ## TODO create example with various thresholds (one per class)
# +
k_f1 = 0.2
k_f2 = 0.8
w = 0.5
theta_f1 = (1-k_f1)*w + k_f1
theta_f2 = (1-k_f2)*w + k_f2
thetas = np.array([theta_f1, theta_f2])
p_f_x = 1
mu0 = 0
mu1 = theta_f1
p_b_f1_x = np.ones_like(x_lin)*((1-p_f1_g_f_x)*mu0 + p_f1_g_f_x*mu1)
mu1 = theta_f1
p_b_f1_x = np.ones_like(x_lin)*((1-p_f1_g_f_x)*mu0 + p_f1_g_f_x*mu1)
plt.subplot(2,1,1)
plot_original_thresholds(x_lin, p_f1_g_f_x, p_f2_g_f_x, theta_f1, theta_f2)
reject = np.ones_like(x_lin)
plt.subplot(2,1,2)
plot_predictions(x_lin, p_f1_g_f_x/theta_f1, p_f2_g_f_x/theta_f2, reject)
# -
| jupyter/tutorial_03_application_cautios_classification_background_check.ipynb |
# # NbConvert, Python library
# In this Notebook, I will introduce you to the programatic API of nbconvert to show you how to use it in various context.
#
# For this I will use one of [@jakevdp](https://github.com/jakevdp) great [blog post](http://jakevdp.github.io/blog/2013/04/15/code-golf-in-python-sudoku/).
# I've explicitely chosen a post with no javascript tricks as Jake seem to be found of right now, for the reason that the becommings of embeding javascript in nbviewer, which is based on nbconvert is not fully decided yet.
#
#
# This will not focus on using the command line tool to convert file. The attentive reader will point-out that no data are read from, or written to disk during the conversion process. Indeed, nbconvert as been though as much as
# possible to avoid IO operation and work as well in a database, or web-based environement.
# #### Quick overview
# <pre style='background-color:#FDD'> Warning, Do use 1.0 or 1.x branch and not master naming have changed.</pre>
#
# <pre style='background-color:#FDD'> Warning, NbConvert is a Tech-Preview, API will change within the next 6 month.</pre>
# Credit, <NAME> (@jdfreder on github)
#
# <center>
# 
# </center>
# The main principle of nbconvert is to instanciate a `Exporter` that controle
# a pipeline through which each notebook you want to export with go through.
# Let's start by importing what we need from the API, and download @jakevdp's notebook.
import requests
response = requests.get('http://jakevdp.github.com/downloads/notebooks/XKCD_plots.ipynb')
response.content[0:60]+'...'
# If you do not have request install downlad by hand, and read the file as usual.
# We read the response into a slightly more convenient format which represent IPython notebook.
# There are not real advantages for now, except some convenient methods, but with time this structure should be able to
# guarantee that the notebook structure is valid. Note also that the in-memory format and on disk format can be slightly different. In particual, on disk, multiline strings might be spitted into list of string to be more version control friendly.
from IPython.nbformat import current as nbformat
jake_notebook = nbformat.reads_json(response.content)
jake_notebook.worksheets[0].cells[0]
# So we have here Jake's notebook in a convenient form, which is mainly a Super-Powered dict and list nested.
# You don't need to worry about the exact structure.
# The nbconvert API exposes some basic exporter for common format and default options. We will start
# by using one of them. First we import it, instanciate an instance with most of the default parameters and fed it
# the downloaded notebook.
import IPython.nbconvert
# +
from IPython.config import Config
from IPython.nbconvert import HTMLExporter
## I use `basic` here to have less boilerplate and headers in the HTML.
## we'll see later how to pass config to exporters.
exportHtml = HTMLExporter(config=Config({'HTMLExporter':{'default_template':'basic'}}))
# -
(body, resources) = exportHtml.from_notebook_node(jake_notebook)
# The exporter returns a tuple containing the body of the converted notebook, here raw HTML, as well as a resources dict.
# The resource dict contains (among many things) the extracted PNG, JPG [...etc] from the notebook when applicable.
# The basic HTML exporter does keep them as embeded base64 into the notebook, but one can do ask the figures to be extracted. Cf advance use. So for now the resource dict **should** be mostly empty, except for 1 key containing some css, and 2 others whose content will be obvious.
#
# Exporter are stateless, you won't be able to extract any usefull information (except their configuration) from them.
# You can directly re-use the instance to convert another notebook. Each exporter expose for convenience a `from_file` and `from_filename` methods if you need.
print resources.keys()
print resources['metadata']
print resources['output_extension']
# print resources['inlining'] # too lng to be shown
# Part of the body, here the first Heading
start = body.index('<h1 id', )
print body[:400]+'...'
# You can directly write the body into an HTML file if you wish, as you see it does not contains any body tag, or style declaration, but thoses are included in the default HtmlExporter if you do not pass it a config object as I did.
# #### Extracting Figures
# When exporting one might want to extract the base64 encoded figures to separate files, this is by default what does the RstExporter does, let see how to use it.
# +
from IPython.nbconvert import RSTExporter
rst_export = RSTExporter()
(body,resources) = rst_export.from_notebook_node(jake_notebook)
# -
print body[:970]+'...'
print '[.....]'
print body[800:1200]+'...'
# Here we see that base64 images are not embeded, but we get what look like file name. Actually those are (Configurable) keys to get back the binary data from the resources dict we havent inspected earlier.
#
# So when writing a Rst Plugin for any blogengine, Sphinx or anything else, you will be responsible for writing all those data to disk, in the right place.
# Of course to help you in this task all those naming are configurable in the right place.
# let's try to see how to get one of these images
resources['outputs'].keys()
# We have extracted 5 binary figures, here `png`s, but they could have been svg, and then wouldn't appear in the binary sub dict.
# keep in mind that a object having multiple _repr_ will store all it's repr in the notebook.
#
# Hence if you provide `_repr_javascript_`,`_repr_latex_` and `_repr_png_`to an object, you will be able to determine at conversion time which representaition is the more appropriate. You could even decide to show all the representaition of an object, it's up to you. But this will require beeing a little more involve and write a few line of Jinja template. This will probably be the subject of another tutorial.
#
# Back to our images,
#
#
from IPython.display import Image
Image(data=resources['outputs']['output_3_0.png'],format='png')
# Yep, this is indeed the image we were expecting, and I was able to see it without ever writing or reading it from disk. I don't think I'll have to show to you what to do with those data, as if you are here you are most probably familiar with IO.
# ## Extracting figures with HTML Exporter ?
# Use case:
#
# > I write an [awesome blog](http://jakevdp.github.io/) in HTML, and I want all but having base64 embeded images.
# Having one html file with all inside is nice to send to coworker, but I definitively want resources to be cached !
# So I need an HTML exporter, and I want it to extract the figures !
# ### Some theory
# The process of converting a notebook to a another format with the nbconvert Exporters happend in a few steps:
#
# - Get the notebook data and other required files. (you are responsible for that)
# - Feed them to the exporter that will
# - sequentially feed the data to a number of `Transformers`. Transformer only act on the **structure**
# of the notebook, and have access to it all.
# - feed the notebook through the jinja templating engine
# - the use templates are configurable.
# - templates make use of configurable macros called filters.
# - The exporter return the converted notebook as well as other relevant resources as a tuple.
# - Write what you need to disk, or elsewhere. (You are responsible for it)
# Here we'll be interested in the `Transformers`. Each `Transformer` is applied successively and in order on the notebook before going through the conversion process.
#
# We provide some transformer that do some modification on the notebook structure by default.
# One of them, the `ExtractOutputTransformer` is responsible for crawling notebook,
# finding all the figures, and put them into the resources directory, as well as choosing the key
# (`filename_xx_y.extension`) that can replace the figure in the template.
#
#
# The `ExtractOutputTransformer` is special in the fact that it **should** be availlable on all `Exporter`s, but is just inactive by default on some exporter.
# second transformer shoudl be Instance of ExtractFigureTransformer
exportHtml._transformers # 3rd one shouel be <ExtractOutputTransformer>
# To enable it we will use IPython configuration/Traitlets system. If you are have already set some IPython configuration options,
# this will look pretty familiar to you. Configuration option are always of the form:
#
# ClassName.attribute_name = value
#
# A few ways exist to create such config, like reading a config file in your profile, but you can also do it programatically usign a dictionary. Let's create such a config object, and see the difference if we pass it to our `HtmlExporter`
# +
from IPython.config import Config
c = Config({
'ExtractOutputTransformer':{'enabled':True}
})
exportHtml = HTMLExporter()
exportHtml_and_figs = HTMLExporter(config=c)
(_, resources) = exportHtml.from_notebook_node(jake_notebook)
(_, resources_with_fig) = exportHtml_and_figs.from_notebook_node(jake_notebook)
print 'resources without the "figures" key :'
print resources.keys()
print ''
print 'Here we have one more field '
print resources_with_fig.keys()
resources_with_fig['outputs'].keys()
# -
# So now you can loop through the dict and write all those figures to disk in the right place...
# #### Custom transformer
# Of course you can imagine many transformation that you would like to apply to a notebook. This is one of the reason we provide a way to register your own transformers that will be applied to the notebook after the default ones.
#
# To do so you'll have to pass an ordered list of `Transformer`s to the Exporter constructor.
#
# But what is an transformer ? Transformer can be either *decorated function* for dead-simple `Transformer`s that apply
# independently to each cell, for more advance transformation that support configurability You have to inherit from
# `Transformer` and define a `call` method as we'll see below.
#
# All transforers have a magic attribute that allows it to be activated/disactivate from the config dict.
from IPython.nbconvert.transformers import Transformer
import IPython.config
print "Four relevant docstring"
print '============================='
print Transformer.__doc__
print '============================='
print Transformer.call.__doc__
print '============================='
print Transformer.transform_cell.__doc__
print '============================='
# ***
# We don't provide convenient method to be aplied on each worksheet as the **data structure** for worksheet will be removed. (not the worksheet functionnality, which is still on it's way)
# ***
# ### Example
# I'll now demonstrate a specific example [requested](https://github.com/ipython/nbconvert/pull/137#issuecomment-18658235) while nbconvert 2 was beeing developped. The ability to exclude cell from the conversion process based on their index.
#
# I'll let you imagin how to inject cell, if what you just want is to happend static content at the beginning/end of a notebook, plese refer to templating section, it will be much easier and cleaner.
from IPython.utils.traitlets import Integer
class PelicanSubCell(Transformer):
"""A Pelican specific transformer to remove somme of the cells of a notebook"""
# I could also read the cells from nbc.metadata.pelican is someone wrote a JS extension
# But I'll stay with configurable value.
start = Integer(0, config=True, help="first cell of notebook to be converted")
end = Integer(-1, config=True, help="last cell of notebook to be converted")
def call(self, nb, resources):
#nbc = deepcopy(nb)
nbc = nb
# don't print in real transformer !!!
print "I'll keep only cells from ", self.start, "to ", self.end, "\n\n"
for worksheet in nbc.worksheets :
cells = worksheet.cells[:]
worksheet.cells = cells[self.start:self.end]
return nbc, resources
# I create this on the fly, but this could be loaded from a DB, and config object support merging...
c = Config({
'PelicanSubCell':{
'enabled':True,
'start':4,
'end':6,
}
})
# I'm creating a pelican exporter that take `PelicanSubCell` extra transformers and a `config` object as parameter. This might seem redundant, but with configuration system you'll see that one can register an inactive transformer on all exporters and activate it at will form its config files and command line.
pelican = RSTExporter(transformers=[PelicanSubCell], config=c)
print pelican.from_notebook_node(jake_notebook)[0]
# ### Programatic example of extending templates / cutom filters
# +
from IPython.nbconvert.filters.highlight import _pygment_highlight
from pygments.formatters import HtmlFormatter
from IPython.nbconvert.exporters import HTMLExporter
from IPython.config import Config
from IPython.nbformat import current as nbformat
# -
# Here we define a dustom 'highlight' filter that apply a custom class to code in css. We register this filter with a already existing name, so it will replace the default one.
# +
def my_highlight(source, language='ipython'):
formatter = HtmlFormatter(cssclass='highlight-ipynb')
return _pygment_highlight(source, formatter, language)
c = Config({'CSSHtmlHeaderTransformer':
{'enabled':False, 'highlight_class':'highlight-ipynb'}})
exportHtml = HTMLExporter( config=c , filters={'highlight2html': my_highlight} )
(body,resources) = exportHtml.from_notebook_node(jake_notebook)
# -
i = body.index('highlight-ipynb')
body[i-12:i+50]
# ### Programatically make templates
# +
from jinja2 import DictLoader
dl = DictLoader({'html_full.tpl':
"""
{%- extends 'html_basic.tpl' -%}
{% block footer %}
FOOOOOOOOTEEEEER
{% endblock footer %}
"""})
exportHtml = HTMLExporter( config=None , filters={'highlight': my_highlight}, extra_loaders=[dl] )
(body,resources) = exportHtml.from_notebook_node(jake_notebook)
for l in body.split('\n')[-4:]:
print l
# -
# ### Real World Use
# @jakevdp use Pelican and IPython Notebook to blog. Pelican [Will use](https://github.com/getpelican/pelican-plugins/pull/21) nbconvert programatically to generate blog post. Have a look a [Pythonic Preambulations](http://jakevdp.github.io/) for Jake blog post.
# @damianavila Wrote a Nicholas Plugin to [Write blog post as Notebook](http://www.damian.oquanta.info/posts/one-line-deployment-of-your-site-to-gh-pages.html) and is developping a js-extension to publish notebooks in one click from the web app.
# <center>
# <blockquote class="twitter-tweet"><p>As <a href="https://twitter.com/Mbussonn">@Mbussonn</a> requested... easieeeeer! Deploy your Nikola site with just a click in the IPython notebook! <a href="http://t.co/860sJunZvj">http://t.co/860sJunZvj</a> cc <a href="https://twitter.com/ralsina">@ralsina</a></p>— <NAME> (@damian_avila) <a href="https://twitter.com/damian_avila/statuses/370306057828335616">August 21, 2013</a></blockquote>
# </center>
# And finaly, what you just did, is replicate what [nbviewer](http://nbviewer.ipython.org) does. WHich to fetch a notebook from url, convert it and send in back to you as a static html.
# ##### A few gotchas
# Jinja blocks use `{% %}`by default which does not play nicely with $\LaTeX$, hence thoses are replaced by `((* *))` in latex templates.
| examples/Notebook/Using nbconvert as a Library.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Neural Network Example
#
# Build a 2-hidden layers fully connected neural network (a.k.a multilayer perceptron) with TensorFlow.
#
# - Author: <NAME>
# - Project: https://github.com/aymericdamien/TensorFlow-Examples/
# ## Neural Network Overview
#
# <img src="http://cs231n.github.io/assets/nn1/neural_net2.jpeg" alt="nn" style="width: 400px;"/>
#
# ## MNIST Dataset Overview
#
# This example is using MNIST handwritten digits. The dataset contains 60,000 examples for training and 10,000 examples for testing. The digits have been size-normalized and centered in a fixed-size image (28x28 pixels) with values from 0 to 1. For simplicity, each image has been flattened and converted to a 1-D numpy array of 784 features (28*28).
#
# 
#
# More info: http://yann.lecun.com/exdb/mnist/
# +
from __future__ import print_function
# Import MNIST data
from tensorflow.examples.tutorials.mnist import input_data
mnist = input_data.read_data_sets("/tmp/data/", one_hot=True)
import tensorflow as tf
# +
# Parameters
learning_rate = 0.1
num_steps = 500
batch_size = 128
display_step = 100
# Network Parameters
n_hidden_1 = 256 # 1st layer number of neurons
n_hidden_2 = 256 # 2nd layer number of neurons
num_input = 784 # MNIST data input (img shape: 28*28)
num_classes = 10 # MNIST total classes (0-9 digits)
# tf Graph input
X = tf.placeholder("float", [None, num_input])
Y = tf.placeholder("float", [None, num_classes])
# -
# Store layers weight & bias
weights = {
'h1': tf.Variable(tf.random_normal([num_input, n_hidden_1])),
'h2': tf.Variable(tf.random_normal([n_hidden_1, n_hidden_2])),
'out': tf.Variable(tf.random_normal([n_hidden_2, num_classes]))
}
biases = {
'b1': tf.Variable(tf.random_normal([n_hidden_1])),
'b2': tf.Variable(tf.random_normal([n_hidden_2])),
'out': tf.Variable(tf.random_normal([num_classes]))
}
# Create model
def neural_net(x):
# Hidden fully connected layer with 256 neurons
layer_1 = tf.add(tf.matmul(x, weights['h1']), biases['b1'])
# Hidden fully connected layer with 256 neurons
layer_2 = tf.add(tf.matmul(layer_1, weights['h2']), biases['b2'])
# Output fully connected layer with a neuron for each class
out_layer = tf.matmul(layer_2, weights['out']) + biases['out']
return out_layer
# +
# Construct model
logits = neural_net(X)
# Define loss and optimizer
loss_op = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(logits=logits, labels=Y))
optimizer = tf.train.AdamOptimizer(learning_rate=learning_rate)
train_op = optimizer.minimize(loss_op)
# Evaluate model (with test logits, for dropout to be disabled)
correct_pred = tf.equal(tf.argmax(logits, 1), tf.argmax(Y, 1))
accuracy = tf.reduce_mean(tf.cast(correct_pred, tf.float32))
# Initialize the variables (i.e. assign their default value)
init = tf.global_variables_initializer()
# -
# Start training
with tf.Session() as sess:
# Run the initializer
sess.run(init)
for step in range(1, num_steps+1):
batch_x, batch_y = mnist.train.next_batch(batch_size)
# Run optimization op (backprop)
sess.run(train_op, feed_dict={X: batch_x, Y: batch_y})
if step % display_step == 0 or step == 1:
# Calculate batch loss and accuracy
loss, acc = sess.run([loss_op, accuracy], feed_dict={X: batch_x,
Y: batch_y})
print("Step " + str(step) + ", Minibatch Loss= " + \
"{:.4f}".format(loss) + ", Training Accuracy= " + \
"{:.3f}".format(acc))
print("Optimization Finished!")
# Calculate accuracy for MNIST test images
print("Testing Accuracy:", \
sess.run(accuracy, feed_dict={X: mnist.test.images,
Y: mnist.test.labels}))
| notebooks/3_NeuralNetworks/1-neural_network_raw.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import csv
def create_csv(p, head: list):
path = p
with open(path, "w") as f:
csv_write = csv.writer(f)
csv_head = head
csv_write.writerow(csv_head)
def write_csv(p, data: list):
with open(p, "a+") as f:
csv_write = csv.writer(f)
csv_write.writerow(data)
# -
import os
import json
head = ['schoolList','schoolid','schoolName','url','street_address','city','zip',
'lowGrade','highGrade','schoolLevel',"isCharterSchool","isMagnetSchool",
"isVirtualSchool",'isTitleISchool','districtName','countyName','rank',
"rankOf",'rankStars','rankLevel','rankStatewidePercentage','rankMovement',
'numberOfStudents','teachersFulltime','isPrivate'
]
# +
def concatecsv(p,file):
test = open(file)
content = test.readlines()
j = json.loads(content[0])
result= []
for school in j['schoolList']:
result.append(school['schoolid'])
result.append(school['schoolName'])
result.append(school['url'])
result.append(school['address']['street'])
result.append(school['address']['city'])
result.append(school['address']['zip'])
result.append(school['lowGrade'])
result.append(school['highGrade'])
result.append(school['schoolLevel'])
result.append(school['isCharterSchool'])
result.append(school['isMagnetSchool'])
result.append(school['isVirtualSchool'])
result.append(school['isTitleISchool'])
result.append(school['district']['districtName'])
result.append(school['county']['countyName'])
result.append(school['rankHistory'][0]['rank'])
result.append(school['rankHistory'][0]['rankOf'])
result.append(school['rankHistory'][0]['rankStars'])
result.append(school['rankHistory'][0]['rankLevel'])
result.append(school['rankHistory'][0]['rankStatewidePercentage'])
result.append(school['rankHistory'][0]['averageStandardScore'])
result.append(schhol['rankHistory']['rankMovement'])
result.append(schhol['schoolYearlyDetails'][0]['numberOfStudents'])
result.append(schhol['schoolYearlyDetails'][0]['percentFreeDiscLunch'])
result.append(schhol['schoolYearlyDetails'][0]['teachersFulltime'])
result.append(schhol['isPrivate'])
# -
result ={}
files=os.listdir()
j = json.loads(content[0])
j['schoolList']
| Data_raw/school_json/.ipynb_checkpoints/Untitled-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.11 64-bit (''bulk-insert'': conda)'
# name: python3
# ---
import os
import pandas as pd
# +
# pip install python-dotenv
from dotenv import load_dotenv
# %load_ext dotenv
# %dotenv
# +
# fetch dataset from ADX
# pip install azure-kusto-data
from azure.kusto.data import KustoClient, KustoConnectionStringBuilder
from azure.kusto.data.helpers import dataframe_from_result_table
import pandas as pd
AAD_TENANT_ID = os.environ['AAD_TENANT_ID']
KUSTO_CLUSTER = os.environ['KUSTO_CLUSTER']
KUSTO_DATABASE = os.environ['KUSTO_DATABASE']
# -
# Interactive login
KCSB = KustoConnectionStringBuilder.with_aad_device_authentication(KUSTO_CLUSTER)
KCSB.authority_id = AAD_TENANT_ID
# +
# Service principal login
# https://github.com/Azure/azure-kusto-python/tree/master/azure-kusto-data
# https://github.com/Azure/azure-kusto-python/blob/master/azure-kusto-data/tests/sample.py
# client_id = os.environ['CLIENT_ID']
# client_secret = os.environ['CLIENT_SECRET']
# authority_id = os.environ['AAD_TENANT_ID']
# KCSB = KustoConnectionStringBuilder.with_aad_application_key_authentication(KUSTO_CLUSTER, client_id, client_secret, authority_id)
# +
# %%time
KUSTO_CLIENT = KustoClient(KCSB)
KUSTO_QUERY = "malware_table_10000"
# KUSTO_QUERY = "set notruncation; malware_table_1000000"
RESPONSE = KUSTO_CLIENT.execute(KUSTO_DATABASE, KUSTO_QUERY)
# -
# %%time
# convert to data frame or native test
df = dataframe_from_result_table(RESPONSE.primary_results[0])
df.head()
len(df)
# Save dataframe to CSV
df.to_csv(r'data/adx-df.csv', index = False, header=True)
# df.to_csv(r'data/adx-df-1000000.csv', index = False, header=True)
df.memory_usage(index=True).sum()
# 663,999,464
| adx-df-csv.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
from IPython.display import display
df1 = pd.DataFrame({'one':[2,1,1,1],'two':[1,3,2,4],'three':[5,4,3,2]})
df1
# -
df1.sort_values(by='two')
df1[['one', 'two', 'three']].sort_values(by=['one','two'])
df1.sort_values(by=['one','two'])
# +
df = pd.DataFrame(np.random.randn(5, 3),
index=['a', 'c', 'e', 'f', 'h'],
columns=['one', 'two', 'three'])
display(df)
df2=df
#df2=df.copy()
df2.loc[['a','c','h'],['one']]=np.nan
df
# -
df['one'].sum()
df.mean(0)
df.mean(1)
display(df)
df.cumsum()
display(df)
df.groupby('one').mean()
df2= pd.DataFrame(np.random.randn(5, 3),
index=['a', 'c', 'e', 'f', 'h'],
columns=['one', 'two', 'three'])
df2['four'] = 'bar'
df2['five'] = df['one'] > 0
df2['timestamp'] = pd.Timestamp('20120101')
display(df2)
df2.loc[['a','c','h'],['one','timestamp']] = np.nan
df2
df2.fillna(0)
df2.fillna(method='pad')
df2.fillna(method='bfill')
df2.fillna(df2.mean())
df3 = pd.DataFrame({'A': [1, 2.1, np.nan, 4.7, 5.6, 6.8],
'B': [.25, np.nan, np.nan, 4, 12.2, 14.4]})
df3.interpolate()
sample = pd.DataFrame({'PR':[10,100,40] })
sample['PR'] = sample['PR'].mask(sample['PR'] < 90, np.nan)
sample
sample = pd.DataFrame({'PR':[10,100,40] })
sample.loc[sample['PR'] < 90, 'PR'] = np.nan
sample
| learning python/learning pandas/pandas_lec2_02.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import seaborn as sns
from matplotlib import pyplot as plt
import sys, os
sys.path.append(f'/home/{os.environ.get("USER")}/PythonLibrary')
# import EDA
# -
tr = pd.read_feather('../data/train.f')
te = pd.read_feather('../data/test.f')
tr_dt = pd.read_feather('../data/train_datetime.f')
te_dt = pd.read_feather('../data/test_datetime.f')
pri = te[te_dt.AvSigVersion>="2018-10-26"]
pub = te[te_dt.AvSigVersion<"2018-10-26"]
tr.columns
pd.pivot_table(tr, index='Census_OSVersion', columns='ProductName', values='HasDetections', aggfunc='mean')
dir(tr.groupby(['Census_OSVersion', 'ProductName']))
tr.groupby(['Census_OSVersion', 'ProductName']).size()
tr.groupby(['Census_OSVersion', 'ProductName']).size() / tr.groupby(['Census_OSVersion']).size()
| jn/EDA_007 OS.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import re
#create a dictonary to store all words of a given length
en_dict = {}
#import the 10,000 most common english words
with open("words.txt","r") as words_input:
for word in words_input:
word_clean = word[:-1]
en_dict.setdefault(len(word_clean),[]).append(word_clean)
"""yeilds char's in range"""
def char_range(start,end):
for c in range(ord(start),ord(end)+1):
yield chr(c)
"""constructs regex to match words with letters in places, such as __a_b_
and returns bag of letters and length"""
def get_regex(input_word):
length = len(input_word)
expr = ""
ltr_bag = ""
for letter in input_word:
if letter == '_':
expr += "[a-z]"
else:
expr += letter
ltr_bag += letter
return length, expr, ltr_bag
"""function to calculate the letter frequency of a word of a given length
with letters in the position of not filled by _ """
def get_freq(input_word):
length, expr, ltr_bag = get_regex(input_word)
c_expr = re.compile(expr)
freq_letter = {char : 0 for char in char_range('a','z')}
#find all words of a given length
for word in en_dict[length]:
if c_expr.match(word):
print(word)
w_list = list(word)
for letter in ltr_bag:
w_list.remove(letter)
for letter in w_list:
freq_letter[letter] += 1
return freq_letter
# -
"""Displays hangman board corosponding with the current guess"""
steps = [
"""
\
\
\
\
\
\
|--------------|\
""",
"""
_______________
|--------------|
""",
"""
_________
|
|
|
|
|_______________
|--------------|
""",
"""
_________
| |
|
|
|
|_______________
|--------------|
""",
"""
_________
| |
| 0
|
|
|_______________
|--------------|
""",
"""
_________
| |
| 0
| |
|
|_______________
|--------------|
""",
"""
_________
| |
| 0
| |\
|
|_______________\
|--------------|\
""",
"""
_________\
| |
| 0
| /|\
|
|_______________
|--------------|
""",
"""
_________
| |
| 0
| /|\
| /
|_______________
|--------------|
""",
"""
_________
| |
| 0
| /|\
| / \
|_______________
|--------------|
"""]
print(steps[-1])
freq = get_freq("__d")
max(freq,key = freq.get)
get_freq("__a_j_i__z")
for char in char_range('a','z'):
print(char)
| AutoHangman.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#Homework Section 6
# -
import pandas as pd
import os
os.getcwd()
movies = pd.read_csv('Movie-Ratings.csv')
biz = pd.read_csv('moviesimbd.csv', encoding = 'latin1')
biz.info()
print(biz.columns)
# +
#Clean up column headers
biz.columns = ['DayOfWeek', 'Director', 'Genre', 'MovieTitle', 'ReleaseDate',
'Studio', 'AdjustedGross_M', 'Budget_M', 'Gross_M',
'IMDbRating', 'MovieLensRating', 'Overseas_M', 'Overseas_Pct',
'Profit_M', 'Profit_Pct', 'Runtime_min', 'US_M',
'Gross_Pct_US']
# +
#Update datatypes
biz.DayOfWeek = biz.DayOfWeek.astype('category')
biz.Genre = biz.Genre.astype('category')
biz.Studio = biz.Studio.astype('category')
biz.Profit_M = biz.Profit_M.astype('float64')
# -
biz.info()
# +
#import data science libraries
from matplotlib import pyplot as plt
import seaborn as sns
import numpy as np
# %matplotlib inline
import warnings
warnings.filterwarnings('ignore')
# +
figure, axes = plt.subplots(figsize=(20,10))
v1 = sns.boxplot(data=biz2, x='Genre', y='Gross_Pct_US')
# +
figure, axes = plt.subplots(figsize=(20,10))
v1 = sns.boxplot(data=biz[(biz.Genre == 'action') | \
(biz.Genre == 'comedy')| \
(biz.Genre == 'adventure')| \
(biz.Genre == 'animation')| \
(biz.Genre == 'drama')], x='Genre', y='Gross_Pct_US')
#alldata_balance = alldata[(alldata[IBRD] !=0) | (alldata[IMF] !=0)]
# -
biz2.info()
biz2.describe()
biz2.info()
v2 = ax = sns.swarmplot(data=biz[(biz.Genre == 'action') | \
(biz.Genre == 'comedy')| \
(biz.Genre == 'adventure')| \
(biz.Genre == 'animation')| \
(biz.Genre == 'drama')], x='Studio', y='Gross_Pct_US', hue='Studio')
# +
#Filter out genres of interest
filter_genre = (biz.Genre == 'action')\
| (biz.Genre == 'comedy')\
| (biz.Genre == 'adventure')\
| (biz.Genre == 'animation')\
| (biz.Genre == 'drama')
filter_studio = (biz.Studio == 'Buena Vista Studios')\
| (biz.Studio == 'Sony')\
| (biz.Studio == 'Universal')\
| (biz.Studio == 'WB')\
| (biz.Studio == 'Paramount Pictures')\
| (biz.Studio == 'Fox')
# -
subset = pd.DataFrame(biz[filter_genre & filter_studio])
subset2 = subset.dropna(axis=0, how='all')
| section6-movies/.ipynb_checkpoints/Section6Homework-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from __future__ import absolute_import, division, print_function, unicode_literals
import pathlib
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers
# -
dataset = pd.read_csv('data_log.csv')
dataset.tail()
dt2 = pd.DataFrame(
data=dataset,
columns=[
'total_tweets', 'tweet_ratio_0', 'tweet_ratio_1', 'tweet_ratio_2', 'tweet_ratio_3', 'tweet_ratio_4',
'tweet_per_user_mean_0', 'tweet_per_user_mean_1', 'tweet_per_user_mean_2', 'tweet_per_user_mean_3','tweet_per_user_mean_4',
]
)
corr = dt2.corr()
sns.heatmap(corr, cmap='BuPu')
train_dataset = dataset.sample(frac=0.8, random_state=0xe621)
test_dataset = dataset.drop(train_dataset.index)
l = [
'total_tweets', 'tweet_ratio_0', 'tweet_ratio_1', 'tweet_ratio_2', 'tweet_ratio_3', 'tweet_ratio_4'
]
# + jupyter={"outputs_hidden": true}
sns.pairplot(train_dataset[l], diag_kind="kde")
# -
train_labels = train_dataset.pop('total_tweets')
train_db = train_dataset.pop('db')
test_labels = test_dataset.pop('total_tweets')
test_db = test_dataset.pop('db')
# + jupyter={"outputs_hidden": true}
train_stats = train_dataset.describe()
# train_stats.pop("total_tweets")
train_stats = train_stats.transpose()
train_stats
# -
def norm(x):
return (x - train_stats['mean']) / train_stats['std']
normed_train_data = norm(train_dataset)
normed_test_data = norm(test_dataset)
def build_model():
model = keras.Sequential([
layers.Dense(41, activation='relu', input_shape=[len(train_dataset.keys())]),
layers.Dense(82, activation='relu'),
layers.Dense(1)
])
optimizer = tf.keras.optimizers.RMSprop(0.001)
model.compile(loss='mse',
optimizer=optimizer,
metrics=['mae', 'mse'])
return model
model = build_model()
model.summary()
# +
# %%time
# Display training progress by printing a single dot for each completed epoch
class PrintDot(keras.callbacks.Callback):
def on_epoch_end(self, epoch, logs):
if epoch % 100 == 0: print('')
print('.', end='')
EPOCHS = 1000
history = model.fit(
normed_train_data, train_labels,
epochs=EPOCHS, validation_split = 0.2, verbose=0,
callbacks=[PrintDot()])
# -
hist = pd.DataFrame(history.history)
hist['epoch'] = history.epoch
hist.tail()
# +
def plot_history(history):
hist = pd.DataFrame(history.history)
hist['epoch'] = history.epoch
plt.figure()
plt.xlabel('Epoch')
plt.ylabel('Mean Abs Error [MPG]')
plt.plot(hist['epoch'], hist['mae'],
label='Train Error')
plt.plot(hist['epoch'], hist['val_mae'],
label = 'Val Error')
# plt.ylim([0,5])
plt.legend()
plt.figure()
plt.xlabel('Epoch')
plt.ylabel('Mean Square Error [$MPG^2$]')
plt.plot(hist['epoch'], hist['mse'],
label='Train Error')
plt.plot(hist['epoch'], hist['val_mse'],
label = 'Val Error')
# plt.ylim([0,20])
plt.legend()
plt.figure()
plt.xlabel('Epoch')
plt.ylabel('Loss')
plt.plot(hist['epoch'], hist['loss'],
label='Loss')
plt.plot(hist['epoch'], hist['val_loss'],
label='Val Loss')
plt.legend()
plt.show()
plot_history(history)
# +
model = build_model()
# The patience parameter is the amount of epochs to check for improvement
early_stop = keras.callbacks.EarlyStopping(monitor='val_loss', patience=10)
history = model.fit(normed_train_data, train_labels, epochs=EPOCHS,
validation_split = 0.2, verbose=0, callbacks=[early_stop, PrintDot()])
plot_history(history)
# +
loss, mae, mse = model.evaluate(normed_test_data, test_labels, verbose=2)
print("Testing set Mean Abs Error: {:5.2f} MPG".format(mae))
# +
test_predictions = model.predict(normed_train_data).flatten()
plt.scatter(train_labels, test_predictions)
plt.xlabel('True Values [MPG]')
plt.ylabel('Predictions [MPG]')
plt.axis('equal')
plt.axis('square')
plt.xlim([0,plt.xlim()[1]])
plt.ylim([0,plt.ylim()[1]])
_ = plt.plot([-100, 100], [-100, 100])
# plt.savefig('owo.png')
# -
error = test_predictions - train_labels
plt.hist(error, bins = 25)
plt.xlabel("Prediction Error [MPG]")
_ = plt.ylabel("Count")
model.summary()
| nn_t.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Visualizing Google Forms Data with Seaborn
# This is the second part of an article from [Practical Business Python](htp://pbpython.com) describing how to retrieve and analyze data from a Google Form.
#
# Please review [part 1](http://pbpython.com/pandas-google-forms-part1.html) for the details of how to set up authentication and get the data into the pandaqs dataframe.
#
# The full article corresponding to this notebook is [here](http://pbpython.com/pandas-google-forms-part2.html)
# ## Setup
# Bring in our standard imports as well as the authentication libraries we will need to get access to our form.
import gspread
from oauth2client.client import SignedJwtAssertionCredentials
import pandas as pd
import json
# Import Ipython display as well as graphing libraries. For this article, we will be using [seaborn](http://stanford.edu/~mwaskom/software/seaborn/index.html).
from IPython.display import display
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# Setup authentication process to pull in the survey data stored in the Google Sheet.
SCOPE = ["https://spreadsheets.google.com/feeds"]
SECRETS_FILE = "Pbpython-key.json"
SPREADSHEET = "PBPython User Survey"
# Based on docs here - http://gspread.readthedocs.org/en/latest/oauth2.html
# Load in the secret JSON key (must be a service account)
json_key = json.load(open(SECRETS_FILE))
# Authenticate using the signed key
credentials = SignedJwtAssertionCredentials(json_key['client_email'],
json_key['private_key'], SCOPE)
# Now open up the file and read all data in a DataFrame
gc = gspread.authorize(credentials)
# Open up the workbook based on the spreadsheet name
workbook = gc.open(SPREADSHEET)
# Get the first sheet
sheet = workbook.sheet1
# Extract all data into a dataframe
results = pd.DataFrame(sheet.get_all_records())
results.head()
# We need to do some cleanup to make the data easier to analyze.
# Do some minor cleanups on the data
# Rename the columns to make it easier to manipulate
# The data comes in through a dictionary so we can not assume order stays the
# same so must name each column
column_names = {'Timestamp': 'timestamp',
'What version of python would you like to see used for the examples on the site?': 'version',
'How useful is the content on practical business python?': 'useful',
'What suggestions do you have for future content?': 'suggestions',
'How frequently do you use the following tools? [Python]': 'freq-py',
'How frequently do you use the following tools? [SQL]': 'freq-sql',
'How frequently do you use the following tools? [R]': 'freq-r',
'How frequently do you use the following tools? [Javascript]': 'freq-js',
'How frequently do you use the following tools? [VBA]': 'freq-vba',
'How frequently do you use the following tools? [Ruby]': 'freq-ruby',
'Which OS do you use most frequently?': 'os',
'Which python distribution do you primarily use?': 'distro',
'How would you like to be notified about new articles on this site?': 'notify'
}
results.rename(columns=column_names, inplace=True)
results.timestamp = pd.to_datetime(results.timestamp)
results.head()
# There are a small number of free form comments. Let's strip those out and remove them from the results.
suggestions = results[results.suggestions.str.len() > 0]["suggestions"]
# Since there are only a small number of comments, just print them out.
# However, if we had more comments and wanted to do more analysis we certainly good.
for index, row in suggestions.iteritems():
display(row)
# Drop the suggestions. We won't use them any more.
results.drop("suggestions", axis=1, inplace=True)
results.head()
# ## Explore the data
# For Numeric columns, start with describe to see what we have
results.describe()
# Because we only have 1, 2, 3 as options the numeric results aren't telling us that much. I am going to convert the number to more useful descriptions.
results['useful'] = results['useful'].map({1: '1-low', 2: '2-medium', 3: '3-high'})
results.head()
# Value counts give us an easy distribution view into the raw numbers
results["version"].value_counts()
# Use normalize to see it by percentage.
results.os.value_counts(normalize=True)
# While the numbers are useful, wouldn't it be nicer to visually show the results?
#
# Seaborn's [factorplot](http://stanford.edu/~mwaskom/software/seaborn/generated/seaborn.factorplot.html) is helpful for showing this kind of categorical data.
#
# Because factorplot is so powerful, I'll build up step by step to show how it can be used for complex data analysis.
#
# First, look at number of users by OS.
sns.factorplot("os", data=results, palette="BuPu")
# It is easy to order the results using x_order
sns.factorplot("os", x_order=["Linux", "Windows", "Mac"], data=results, palette="BuPu")
# Do a similar plot on python version
sns.factorplot("version", data=results, palette="BuPu")
# This is useful but wouldn't it be better to compare with OS and preferred python version? This is where factorplot starts to show more versatility. The key component is to use hue to automatically slice the data by python version (in this case).
sns.factorplot("os", hue="version", x_order=["Linux", "Windows", "Mac"], data=results, palette="Paired")
# Because seaborn knows how to work with dataframes, we just need to pass in the column names for the various arguments and it will do the analysis and presentation.
#
# How about if we try to see if there is any relationship between how useful the site is and OS/Python choice? We can add the useful column into the plot using col.
sns.factorplot("version", hue="os", data=results, col="useful", palette="Paired")
# If we can add a column, we can also add a row and seaborn takes care of the rest.
#
# In looking at the data, we have two different versions of winpython so clean that up first.
results['distro'] = results['distro'].str.replace('WinPython', 'winpython')
results.head()
# We can also look at the distros. Since there is some overlap with the distros and os, let's only look at a subset of distros. For instance, someone using winpython is not going to be using it on a Mac.
results['distro'].value_counts()
# The most meaningful data would be looking at the Anaconda and Official python.org binaries. Let's filter all of our data only on these two values.
results_distro = results[results["distro"].isin(["Anaconda", "Official python.org binaries"])]
results_distro.head()
# Now do our factorplot with multiple columns and rows using row and col.
sns.factorplot("version", hue="os", data=results_distro, col="useful", row="distro", margin_titles=True, sharex=False)
# ## Responses over time
# We know that we have 55 results now. It would be interesting to see how those results came in over time. Using this method, we can very simply look at this by any time period we want.
#
# The seaborn's [timeseries](http://stanford.edu/~mwaskom/software/seaborn/generated/seaborn.tsplot.html) supports this type of analysis and much more.
#
# For ease of calculating responses over time, add a count colum for each response.
results["count"] = 1
results.head()
# To get totals over time, set our index to the timestamp
total_results = results.set_index('timestamp')
total_results.head()
# Use pandas TimeGrouper to summarize the data by day and do a cumulative sum. We could easily do this for any time period too.
running_results = total_results.groupby(pd.TimeGrouper('D'))["count"].count().cumsum()
running_results
# To label the x-axis we need to define our time range
step = pd.Series(range(0,len(running_results)), name="Days")
sns.tsplot(running_results, value="Total Responses", time=step, color="husl")
# ## Heatmaps and Clustermaps
# The final section of data to analyze is the frequency with which readers are using different technology. I am going to use a [heatmap](http://stanford.edu/~mwaskom/software/seaborn/generated/seaborn.heatmap.html#seaborn.heatmap) to look for any interesting insights.
# Let's look at the data again.
results.head()
results["freq-py"].value_counts()
# What we need to do is construct a single DataFrame with all the value_counts for the specific technology.
# First we will create a list containing each value count.
all_counts = []
for tech in ["freq-py", "freq-sql", "freq-r", "freq-ruby", "freq-js", "freq-vba"]:
all_counts.append(results[tech].value_counts())
display(all_counts)
# Now, concat the lists along axis=1.
#
# Fill in any nan values with 0 too.
tech_usage = pd.concat(all_counts, keys=["Python", "SQL", "R", "Ruby", "javascript", "VBA"], axis=1)
tech_usage = tech_usage.fillna(0)
tech_usage
# We have a nice table but there are a few problems.
#
# First, we have one column with blank values that we don't want.
#
# Secondly, we would like to order from Daily -> Never. Use reindex to accomplish both tasks.
tech_usage = tech_usage.reindex(["Daily", "A couple times a week", "Once a month", "Infrequently", "Never"])
tech_usage
# Now that the data is in the correct table format, we can create a heatmap.
sns.heatmap(tech_usage, annot=True)
# So, what does this tell us?
#
# Not surprisingly, most people use python very frequently.
#
# Additionally, it looks like very few survey takers are using Ruby or VBA.
# A variation of the heatmap is the [clustermap](http://stanford.edu/~mwaskom/software/seaborn/generated/seaborn.clustermap.html#seaborn.clustermap). The main feature it does is that it tries to reorganize the data to more easily see relationships/clusters.
sns.clustermap(tech_usage, annot=True)
# At first glance, it may seem to be a repeat but you'll notice that the order of the axes are different.
#
# For instance, python and SQL are clusterd in the lower right with higher usage and Ruby and VBA have a cluster in the upper left with lower usage.
| notebooks/18_Google-Forms-Data-Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Stochastic Gradient Descent
#
# We now know how to use a neural network to make predictions, and how to train that network with regard to some input data. However, to avoid numerical blowup, we need to learn pretty slowly for each iteration, and if we want to get anywhere, we'll need a lot more iterations.
#
# To do that, we'll randomly subdivide our training data into lots of small pieces, and train on each piece. We'll use all the data equally, but move a little based on each little piece.
#
# Each training step is now very quick, but the extra overhead of subdividing and so on will mean each epoch now takes a fair amount longer. It'll be worth it, I promise.
# ## Subdividing the Data
#
# What we want to do is *randomly* divide the data into lots of little pieces called *mini-batches*. We'll fix the size of the pieces in advance. Typically, larger pieces mean fewer training steps per epoch (bad) but smaller pieces mean more randomness in each training step (also bad). So we'll try to strike a balance, but it turns out not to be too big of a deal, so long as we randomize each epoch to avoid certain kinds of bias, and make our pieces large enough to avoid whole mini-batches of all one class (for example).
#
# This is fairly easy to do with the `numpy.random.permutation` method:
def get_mini_batches(batch_size, X, Y):
scrambled_indices = np.random.permutation(len(X))
batch_edges = list(range(0, len(X), batch_size)) + [len(X)]
num_batches = len(batch_edges)-1
for i in range(0, num_batches):
batch_indices = scrambled_indices[batch_edges[i]:batch_edges[i+1]]
yield X[batch_indices], Y[batch_indices]
# The above code will take in the batch size, as well as the training data (input and answers). It scrambles the indices of the rows, and divides those indices into pieces of length `batch_size`, possibly with a last piece of smaller size if `X` doesn't divide evenly into mini-batches of the specified size. Every row of `X` appears in exactly one mini-batch, and the order is random and different every time.
# ## Using Stochastic Gradient Descent
#
# We can now adapt the code from the simulations before, where for each epoch, we cycle through all the mini-batches and train on them. Let's use the MNIST data from before:
# +
from mnist_import import get_mnist_nice
train_X, train_Y, test_X, test_Y = get_mnist_nice()
n = train_X.shape[1]
k = train_Y.shape[1]
# -
# We'll also use the same architecture as before, for the sake of a fair comparison:
# +
from basic_nn import *
np.random.seed(31)
# Step 1: pick architecture (in prose above)
cost_function = cost_CE # this is a classification problem
learning_rate = 0.125 # picked arbitrarily, seems to work okay
# Step 2: initialize
#100 input neurons, 100 neurons in a hidden layer, so 100+100+10 total
neuron_sizes = [100, 100]
weights, biases = initialize_xavier_sigmoid(n, k, neuron_sizes)
acts = [act_sigmoid for _ in range(0, len(weights))] # all sigmoid
# +
# Step 3: train
import time
t1 = time.time()
num_epochs = 20
batch_size = 50
for epoch in range(0, num_epochs):
# we'll keep track of the cost as we go
total_cost = 0
num_batches = 0
for X_mb, Y_mb in get_mini_batches(batch_size, train_X, train_Y):
x, z, y = forward_prop(weights, biases, acts, X_mb)
bp_grad_w, bp_grad_b = back_prop(weights, biases, acts, cost_function, X_mb, Y_mb, x, y, z)
for i in range(0, len(weights)):
weights[i] -= learning_rate * bp_grad_w[i] / len(X_mb)
biases[i] -= learning_rate * bp_grad_b[i] / len(X_mb)
total_cost += cost_function(y[-1], Y_mb, aggregate=True)
num_batches += 1
cost = total_cost / num_batches # average cost
print("Cost {2:0.7f} through epoch {0}; took {1:0.3f} seconds so far.".format(epoch, time.time()-t1, cost))
# -
# Some notes:
# 1. We now call it the error "through" epoch [n]. This is because we no longer get all the error at once, but get a little error, train a little, get a little more error, train a little more, and so on.
# 2. Epochs take a lot longer; they took about 5 seconds before, and now take about 10 seconds (depending on the computer).
# 3. Each epoch accomplishes a lot lot more.
#
# Let's do a comparison of classification error, as opposed to last time.
# ## Evaluating the Results
#
# Let's look at classification error. It's not as convenient as it was before to save the predictions at each epoch, since we didn't really construct them all at once. But we can use the network we have now and evaluate its classification error.
_, _, y = forward_prop(weights, biases, acts, train_X)
success_rate = classification_success_rate(y[-1], train_Y)
print("After {0} epochs, got {1:0.3f}% classifications correct.".format(num_epochs, 100*success_rate))
# That's not bad, right? We'll do a lot better after we implement more bells and whistles (the state of the art is above 99%) but this is worlds ahead of our previous attempt, which got less than 12% correct.
# ## Why It Works
#
# We are trying to do gradient descent, and fundamentally we are. Unfortunately, computing the whole gradient is extremely time-consuming when the data is large. What we do instead is take a *random sample* from the dataset, and compute the gradient using that small sample. Since it's smaller, we can do it a lot more quickly. However, since it doesn't include all the data, it could be slightly wrong; that is, this process introduces random noise. This is why it's called *stochastic gradient descent*.
#
# So in a nutshell, in the time it used to take to make two gradient steps, it can now do two thousand gradient steps, with a small amount of error at every stage. Since we use all the data eventually, these errors balance each other out (sort of) and we end up going mostly in the right direction.
# ## Choosing the Batch Size
#
# You'll notice we picked a batch size of 50, without any justification. How did we pick that? Should you pick something else? Like the learning rate, this is a *hyperparameter* you need to understand and pick intelligently.
#
# In theory, the lower the batch size, the faster the training. Smaller mini-batches mean smaller matrices, which mean faster training time. Cutting your batch size in half should mean half the rows in each matrix, so half the training time per batch. This isn't completely true, as there is some overhead in the process which has a fixed time, but it's basically true. Phrased another way, your whole epoch takes an approximately fixed amount of time, and you'd like to train it as much as possible.
#
# On the other hand, the lower the batch size, the more random noise you inject. Since you're only taking a random sample of the data, it's not going to represent the total dataset perfectly; it'll have a little *bias* from overrepresenting certain kinds of data. This can actually be a good thing, helping you get off of local minima, but if the mini-batches are too small, you have too much random noise, and you're "training" really fast by bouncing around in random directions.
#
# So what you want is a balance. However, this parameter isn't super important. Don't make it too small, but it turns out to be essentially a "computational" parameter -- if you get it wrong, your network will train slower (higher aggregate errors by epoch) but you won't introduce skew or permanently hurt your models. So Bengio's advice is to tune the rest of your network well, then experiment with a few different batch sizes to see which one trains fastest, and use that one.
| 07 - Stochastic Gradient Descent.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import random
import pickle
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import plotly.express as px
from scipy.stats import pearsonr
from sklearn.metrics.pairwise import cosine_similarity
from evolvemb import load_diachronic_dataset, compute_emb_snapshots, list_new_tokens, list_multiple_meanings_tokens, list_semantic_shift_tokens, plot_emb_over_time
# %load_ext autoreload
# %autoreload 2
# -
def get_emb_snapshots(snapshots, start_date="2019-01-01", local_emb_name="dummy", min_freq=100, n_tokens=10000,
saveemb=True, datapath="data/nytimes_dataset.txt"):
if local_emb_name.startswith("data/"):
# e.g. for a fine-tuned model saved in the data folder
savepath = f"data/snapshot_emb_{local_emb_name.lower()[5:]}_{start_date}_{snapshots[-1]}_{min_freq}.pkl"
else:
savepath = f"data/snapshot_emb_{local_emb_name.lower()}_{start_date}_{snapshots[-1]}_{min_freq}.pkl"
# see if we can just load the embeddings
if os.path.exists(savepath):
try:
snapshot_emb = pickle.load(open(savepath, "rb"))
return snapshot_emb
except Exception as e:
print("could not load embeddings:", e)
# load dataset
sentences, dates = load_diachronic_dataset(datapath, start_date, snapshots[-1])
# compute snapshots
snapshot_emb = compute_emb_snapshots(sentences, dates, snapshots, local_emb_name, min_freq, n_tokens)
# possibly save embeddings
if saveemb:
try:
pickle.dump(snapshot_emb, open(savepath, "wb"), -1)
print(f"successfully saved embeddings at {savepath}")
except Exception as e:
print("error saving embeddings:", e)
return snapshot_emb
# ## Minimal Example (to create embedding snapshots for the Dash App)
# desired snapshot dates: pre- and post-corona outbreak in detail
snapshots = [f"2019-{i:02}-31" for i in range(6, 13)] + [f"2020-{i:02}-31" for i in range(1, 13)]
# compute embedding snapshots with "bert-base-uncased" (can be abbreviated as "bert"; only works for bert and roberta)
snapshot_emb = get_emb_snapshots(snapshots, start_date="2019-04-01", local_emb_name="bert", min_freq=50)
# save embeddings to use with app.py
pickle.dump(snapshot_emb, open("snapshot_emb.pkl", "wb"), -1)
# see which words have changed the most at some point in the time period
tokens = list_new_tokens(snapshot_emb)
print("new tokens:")
print("\n".join([f"{x[0]:15} ({x[1]})" for x in tokens[:25]]))
tokens = list_multiple_meanings_tokens(snapshot_emb)
print("tokens with multiple meanings:")
print("\n".join([f"{x[0]:15} ({x[1]:.4f})" for x in tokens[:25]]))
tokens = list_semantic_shift_tokens(snapshot_emb)
print("tokens with a semantic shift:")
print("\n".join([f"{x[0]:15} ({x[1]:.4f})" for x in tokens[:25]]))
# create interactive plots for word "category"
fig_time, fig_pca = plot_emb_over_time(snapshot_emb, "positive")
fig_time.show()
fig_pca.show()
# ## Full Analysis (to reproduce results from paper)
# #### "Dorian" Plot from the introduction
snapshots = [f"2019-{i:02}-{j}" for i in range(6, 13) for j in [15, 31]]
snapshot_emb = get_emb_snapshots(snapshots, start_date="2019-01-01", local_emb_name="data/nyt_bert", min_freq=35, saveemb=True)
# create interactive plots for word "Dorian" and save
fig_time, fig_pca = plot_emb_over_time(snapshot_emb, "dorian", savefigs="nyt_bert", savestyle=1)
fig_time.show()
fig_pca.show()
# #### Compare different transformer architectures
# +
def test_cosine_sim_knn(snapshot_emb, k=10):
# check the overlap between cosine similarity and knn intersection score (Gonen et al., 2020)
snapshots = sorted(snapshot_emb)
f, l = snapshots[0], snapshots[-1]
token_sim = []
token_knn_score = []
# ignore words that had a zero embedding in the beginning
tokens = [t for t in snapshot_emb[f].input_model.index2token if np.any(snapshot_emb[f][t] != 0)]
for i, t in enumerate(tokens):
if not i%100: print(f"Processing {i+1:6}/{len(tokens)}", end="\r")
token_sim.append(cosine_similarity(snapshot_emb[f][t][None, :], snapshot_emb[l][t][None, :])[0, 0])
knn1 = set(snapshot_emb[f].get_nneighbors(t, k, include_simscore=False))
knn2 = set(snapshot_emb[l].get_nneighbors(t, k, include_simscore=False))
token_knn_score.append(len(knn1.intersection(knn2))/k)
print(f"Processing {len(tokens):6}/{len(tokens)}")
token_sim, token_knn_score = np.array(token_sim), np.array(token_knn_score)
plt.figure()
plt.scatter(token_sim, token_knn_score)
plt.xlabel("cosine similarity")
plt.ylabel(f"intersection of NN @ k={k}")
plt.title(f"correlation: {pearsonr(token_sim, token_knn_score)[0]:.3f}")
return tokens, token_sim, token_knn_score
def compare_most_changed_tokens(tokens1, tokens2, name1, name2, c="#7C0033", new_fig=True):
# compare the similarity scores of the most changed tokens from two models
tokens1, tokens2 = dict(tokens1), dict(tokens2)
tokens = set(tokens1.keys())
tokens.intersection_update(tokens2.keys())
tokens = sorted(tokens)
scores1 = np.array([tokens1[t] for t in tokens])
scores2 = np.array([tokens2[t] for t in tokens])
if new_fig:
plt.figure(figsize=(6, 6))
plt.grid()
plt.scatter(scores1, scores2, s=10, c=c, alpha=0.5)
plt.xlabel(name1, fontsize=14)
plt.ylabel(name2, fontsize=14)
corr = pearsonr(scores1, scores2)[0]
plt.title(f"correlation: {corr:.3f}")
return corr
# +
# desired snapshot dates: pre- and post-corona outbreak in detail
snapshots = [f"2019-{i:02}-31" for i in range(6, 13)] + [f"2020-{i:02}-31" for i in range(1, 13)]
def run_analysis(local_emb_name="dummy", savefigs="", check_knn_score=False):
# generate/load embeddings
snapshot_emb = get_emb_snapshots(snapshots, start_date="2019-04-01", local_emb_name=local_emb_name, min_freq=50, n_tokens=10000, saveemb=True)
# see which words have changed the most at some point in the time period
changed_tokens = list_new_tokens(snapshot_emb)
print("new tokens:")
print("\n".join([f"{x[0]:15} ({x[1]})" for x in changed_tokens[:25]]))
changed_tokens = list_multiple_meanings_tokens(snapshot_emb)
print("tokens with multiple meanings:")
print("\n".join([f"{x[0]:15} ({x[1]:.4f})" for x in changed_tokens[:25]]))
changed_tokens = list_semantic_shift_tokens(snapshot_emb)
print("tokens with a semantic shift:")
print("\n".join([f"{x[0]:15} ({x[1]:.4f})" for x in changed_tokens[:25]]))
if check_knn_score:
# see in how far the cosine similarity and knn intersection score agree
for k in [10, 100, 1000]:
tokens, token_sim, token_knn_score = test_cosine_sim_knn(snapshot_emb, k=k)
# create plots from
fig_time, fig_pca = plot_emb_over_time(snapshot_emb, "category", k=5, savefigs=savefigs)
fig_time.show()
fig_pca.show()
return snapshot_emb, changed_tokens
# -
# run analysis for bert
snapshot_emb, bert_most_changed = run_analysis(local_emb_name="bert", savefigs="bert")
_ = plot_emb_over_time(snapshot_emb, "biden")
# same analysis for roberta
_, roberta_most_changed = run_analysis(local_emb_name="roberta")
# and both finetuned models
snapshot_emb, bert_ft_most_changed = run_analysis(local_emb_name="data/nyt_bert")
_ = plot_emb_over_time(snapshot_emb, "biden")
_, roberta_ft_most_changed = run_analysis(local_emb_name="data/nyt_roberta")
# see in how far the most changed tokens from BERT and RoBERTa agree (before and after fine-tuning)
_ = compare_most_changed_tokens(bert_most_changed, bert_ft_most_changed, "BERT", "BERT (fine-tuned)")
_ = compare_most_changed_tokens(roberta_most_changed, roberta_ft_most_changed, "RoBERTa", "RoBERTa (fine-tuned)")
corr1 = compare_most_changed_tokens(bert_most_changed, roberta_most_changed, "BERT", "RoBERTa")
corr_ft = compare_most_changed_tokens(bert_ft_most_changed, roberta_ft_most_changed, "BERT", "RoBERTa", c="#00537C", new_fig=False)
plt.title("")
plt.legend([f"pre-trained $(r: {corr1:.3f})$", f"fine-tuned $(r: {corr_ft:.3f})$"], fontsize=14)
# #### Evaluate on data with artificial semantic (non-)shifts
# check most changed tokens when sentences are shuffled
# (i.e. determine threshold on cosine similarity to avoid false positives)
savepath = f"data/snapshot_emb_shuffled_2019-04-01_{snapshots[-1]}_50.pkl"
# see if we can just load the embeddings
if os.path.exists(savepath):
snapshot_emb = pickle.load(open(savepath, "rb"))
else:
# load dataset
sentences, dates = load_diachronic_dataset("data/nytimes_dataset.txt", "2019-04-01", snapshots[-1])
# shuffle sentences (but leave dates as they were!)
random.seed(10)
random.shuffle(sentences) # inplace
# compute snapshots as before with shuffled sentences
snapshot_emb = compute_emb_snapshots(sentences, dates, snapshots, "bert", 50)
pickle.dump(snapshot_emb, open(savepath, "wb"), -1)
# see which words have changed the most
tokens = list_new_tokens(snapshot_emb)
print("new tokens:")
print("\n".join([f"{x[0]:15} ({x[1]})" for x in tokens[:25]]))
tokens = list_multiple_meanings_tokens(snapshot_emb)
print("tokens with multiple meanings:")
print("\n".join([f"{x[0]:15} ({x[1]:.4f})" for x in tokens[:25]]))
tokens = list_semantic_shift_tokens(snapshot_emb)
print("tokens with a semantic shift:")
print("\n".join([f"{x[0]:15} ({x[1]:.4f})" for x in tokens[:25]]))
# example plot for our previous most changed token
fig_time, fig_pca = plot_emb_over_time(snapshot_emb, "category")
fig_time.show()
fig_pca.show()
# load original bert checkpoint
savepath = f"data/snapshot_emb_bert_2019-04-01_{snapshots[-1]}_50.pkl"
snapshot_emb = pickle.load(open(savepath, "rb"))
# select two words that occur fairly often and that don't have too much in common
# the input model of the embeddings already contains counts of the tokens, check the 100 most frequent
print(snapshot_emb[snapshots[-1]].input_model.token_counts.most_common(100))
# select two words from which we believe they aren't too similar
word1 = "president"
word2 = "coronavirus"
# check their cosine similarities to be sure they really are not very similar
print(f"cosine similarity between {word1} and {word2}", cosine_similarity(snapshot_emb[snapshots[-1]][word1][None, :], snapshot_emb[snapshots[-1]][word2][None, :]))
# look at plots for both words to check their original nearest neighbors over time
fig_time, _ = plot_emb_over_time(snapshot_emb, word1)
fig_time.show()
fig_time, _ = plot_emb_over_time(snapshot_emb, word2)
fig_time.show()
# load dataset
sentences, dates = load_diachronic_dataset("data/nytimes_dataset.txt", "2019-04-01", snapshots[-1])
# split the original list with sentences into 3 list: those with word1, with word2, and without any of the words
sentences_word1 = []
sentences_word2 = []
sentences_without = []
dates_without = []
# create an artificial new word as a combination of both words
newword = f"{word1}{word2}"
for i, s in enumerate(sentences):
if word1 in s:
# ignore sentences with both words
if word2 in s:
continue
# replace original word with artificial word
sentences_word1.append([newword if w == word1 else w for w in s])
elif word2 in s:
sentences_word2.append([newword if w == word2 else w for w in s])
else:
sentences_without.append(s)
dates_without.append(dates[i])
print(f"number of sentences with {word1}:", len(sentences_word1))
print(f"number of sentences with {word2}:", len(sentences_word2))
print("number of sentences without the words:", len(sentences_without))
# +
# sigmoid function based on which we'll draw the sentences
def sigm(i, n):
return 1/(1+np.exp(-(i-n/2)/(n/10)))
# check that it looks correctly independent of the number of sentences
# for n in [1000, 10000]:
# x = np.arange(n)
# plt.figure()
# plt.plot(x, sigm(x, n));
# -
# shuffle both sets of sentences and take the same number from each
random.seed(23)
random.shuffle(sentences_word1)
random.shuffle(sentences_word2)
min_len = min(len(sentences_word1), len(sentences_word2))
sentences_word1, sentences_word2 = sentences_word1[:min_len], sentences_word2[:min_len]
# combine both lists into a single list where we first have a high priority of choosing sentences from
# the first word and then from the second
sentences_both = []
n = len(sentences_word1)+len(sentences_word2)
for i in range(n):
# add either a sentence with word1 or word2 depending on sigmoid threshold
if (len(sentences_word1) > len(sentences_word2)) or (len(sentences_word1) and random.random() >= sigm(i, n)):
sentences_both.append(sentences_word1.pop())
else:
sentences_both.append(sentences_word2.pop())
# check some sentences at the beginning ... all about word1
print("\n".join([" ".join(s) for s in sentences_both[:10]]))
# ... and some at the end; they are about word2
print("\n".join([" ".join(s) for s in sentences_both[-10:]]))
# interleave the new sentences with the originals
sentences_new = []
dates_new = []
# every r_th sentence should be from our artificial list
r = len(sentences_without) // len(sentences_both)
n = len(sentences_without)
i_both = 0
for i in range(n):
# always add the original sentence
sentences_new.append(sentences_without[i])
dates_new.append(dates_without[i])
# in between add a sentence for the new list
if not i % r and i_both < len(sentences_both):
sentences_new.append(sentences_both[i_both])
i_both += 1
# add the same date again
dates_new.append(dates_without[i])
# possibly add a last new sentence
if i_both < len(sentences_both):
sentences_new.append(sentences_both[i_both])
dates_new.append(dates_without[i])
# save new sentences as a dataset to fine tune bert on
with open(f"data/nytimes_dataset_{newword}.txt", "w") as f:
f.write("\n".join([f"{dates_new[i]}\t{' '.join(sentences_new[i])}" for i in range(len(dates_new))]))
# compute snapshots from our new sentences
savepath = f"data/snapshot_emb_bert_{newword}_2019-04-01_{snapshots[-1]}_50.pkl"
# see if we can just load the embeddings
if os.path.exists(savepath):
snapshot_emb = pickle.load(open(savepath, "rb"))
else:
snapshot_emb = compute_emb_snapshots(sentences_new, dates_new, snapshots, "bert", 50)
pickle.dump(snapshot_emb, open(savepath, "wb"), -1)
# see which words have changed the most at some point in the time period
tokens = list_semantic_shift_tokens(snapshot_emb)
print("tokens with a semantic shift:")
print("\n".join([f"{x[0]:15} ({x[1]:.4f})" for x in tokens[:200]]))
# example plot for our new word
fig_time, fig_pca = plot_emb_over_time(snapshot_emb, newword, k=5, savefigs="bert", savestyle=1)
fig_time.show()
fig_pca.show()
# compute snapshots from our new sentences with the fine-tuned model
savepath = f"data/snapshot_emb_nyt_bert_{newword}_2019-04-01_{snapshots[-1]}_50.pkl"
# see if we can just load the embeddings
if os.path.exists(savepath):
snapshot_emb = pickle.load(open(savepath, "rb"))
else:
snapshot_emb = compute_emb_snapshots(sentences_new, dates_new, snapshots, f"data/nyt_bert_{newword}", 50)
pickle.dump(snapshot_emb, open(savepath, "wb"), -1)
# check which are now the most changed words
tokens = list_semantic_shift_tokens(snapshot_emb)
print("tokens with a semantic shift:")
print("\n".join([f"{x[0]:15} ({x[1]:.4f})" for x in tokens[:60]]))
# example plot for our new word
fig_time, fig_pca = plot_emb_over_time(snapshot_emb, newword, k=5, savefigs="nyt_bert", savestyle=1)
fig_time.show()
fig_pca.show()
# +
# check what our semantic shift score computes for different
def corrplot(sims):
sims = np.array(sims)
diff = sims[:-1] - sims[1:]
plt.figure()
plt.plot(np.arange(len(sims)), sims, "b", label="cosine similarity to last embedding")
plt.plot(np.arange(1, len(sims)), -np.maximum(diff, 0), "g", label="decrease from consecutive scores")
plt.plot([0, len(sims)-1], [0, 0], "--r", linewidth=0.5)
plt.title(f"overall change: {sims[-1] - sims[0]:.2f}, overall decrease: {-np.sum(diff[diff>0]):.2f}, final score: {sims[-1] - sims[0]-np.sum(diff[diff>0]):.2f}")
sims = [0., 0., 0., 0., 0., 0.2, 0.4, 0.6, 0.8, 1., 1., 1., 1., 1., 1.] # steep increase, no wiggles -> 1
corrplot(sims)
sims = [0., 0., 0.05, 0.1, 0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.9, 0.95, 1., 1.] # slow increase, no wiggles -> 1
corrplot(sims)
sims = [0., 0., 0.1, 0.2, 0.3, 0.2, 0.4, 0.3, 0.5, 0.4, 0.6, 0.7, 0.6, 0.8, 1., 1.] # increase with wiggles -> < 1
corrplot(sims)
sims = [0.5, 0.5, 0.5, 0.5, 0.5, 0.6, 0.7, 0.8, 0.9, 1., 1., 1., 1., 1., 1.] # increase, smaller over all change -> < 1
corrplot(sims)
sims = [1., 1., 1., 0.8, 0.4, 0.2, 0.4, 0.6, 0.8, 1., 1., 1., 1., 1., 1.] # periodic, start at 1 --> negative
corrplot(sims)
sims = [1., 0.8, 0.6, 0.8, 0.6, 0.4, 0.2, 0.4, 0.6, 0.8, 0.8, 1., 1., 1., 1.] # periodic, start at 1 --> negative
corrplot(sims)
sims = [0., 0.2, 0.4, 0.6, 0.8, 0.6, 0.4, 0.2, 0., 0.2, 0.4, 0.6, 0.8, 1., 1., 1.] # periodic, start at 0 --> small
corrplot(sims)
# -
| src/nytimes_diachronic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### A問題
N = int(input())
print(N if N%2 == 0 else N*2)
# ### B問題
N = int(input())
A = list(map(int, input().split()))
print(abs(max(A)-min(A)))
# ### C問題
# わからなかったので,解説を読んだ。<br>
#
# #### 解説
# Bi = Ai − i と定義すれば、問題は、自由に整数 b を選び、abs(Bi − b) の総和を最小化する問 題です。ここで、b を Bi の中央値にするのが最適であることがわかります。中央値でない場合は、 中央値に近づけることで損をしないことからこれはわかります。よって、Bi をソートして中央値 を求め、実際に答えを計算すればよいです。ソートがボトルネックとなり、この問題は O(N logN ) で解けます。<br>
#
#
# このことから,中央値を求めればよいので,以下のようなコードを実装する。
# +
N = int(input())
A = list(map(int, input().split()))
for i in range(N):
A[i] -= i+1
A.sort()
median = A[N//2]
for i in range(N):
A[i] -= median
A[i] = abs(A[i])
print(sum(A))
# -
# で,とりあえず解けた。
# 高速に解いているコードをフォローする。
# 最速のコードは,<br>
# https://abc102.contest.atcoder.jp/submissions/2778856<br>
# だったので,解読する。
# +
N = int(input())
An = list(map(int, input().split()))
for n in range(N):
An[n] -= (n+1)
An.sort()
if (N-1) % 2 != 0:
m = N//2 - 1
else:
m = (N-1)//2
# ここのbfは,中央値よりも小さい値の和を出している。
# そのため,合計値の和の負の値とその数☓中央値を出している。
bf = -sum(An[0:m+1:]) + (m+1)*An[m]
# ここのafは,中央値よりも大きな値の和を出している。
# その逆。
af = sum(An[m+1::]) - (N-m-1)*An[m]
ans = bf + af
print(ans)
# -
# ```
# for i in range(N):
# A[i] -= i+1
# A.sort()
# ```
# と
#
# ```
# for i in range(N):
# A[i] -= median
# A[i] = abs(A[i])
# print(sum(A))
# ```
# がブサイクなので方法がないか探していると,<br>
# https://abc102.contest.atcoder.jp/submissions/2783497<br>
# に一行でまとめる方法が書いてあった。
#
# ```
# C = sorted (a - i for a, i in enumerate(A, 1))
# ```
#
# ```
# ans = sum(abs(a-median) for a in A)
# ```
# と書けばよかった。改める。
# +
N = int(input())
A = list(map(int, input().split()))
C = sorted(a - i for i, a in enumerate(A,1))
median = C[N//2]
print(sum(abs(c-median) for c in C))
# -
# これでコードが小さくなった。
# ```
# for i, a in enumerate(A, 1)
# ```
# は,iterateする数字,iterateするlistの順に書く。
# ### D問題
#
# 全探索はしたくない。相変わらず区切る方針が不明なので,解説を読む。<br>
# 予め累積和を求めておき、ある区間の和を O(1) で求められるようにしておきます。
# 切り込みを入れる 3 箇所のうち、真ん中の切れ込みを 1 つ固定したとします。真ん中の切れ込み で切ったあとの 2 つの数列を L,R とします。あとは、L と R をそれぞれ 1 回ずつ切ることを考え ます。<br>
# L の切り方を考えると、切ったあとの 2 つの数列の要素の総和ができるだけ近いほうが良いこと がわかります。よって、真ん中の切れ込みを固定すると、左の切れ込みの位置を決めることが出来 ます。真ん中の切れ込みの位置が、i 番目の要素と i + 1 番目の要素の間の時、最適な左の切れ込 みの位置を F (i) とおきます。このとき、F (i) は i に関して単調増加であり、尺取り方を用ること で F (i) をすべての i に対して O(N ) で求めることが出来ます。<br>
# 右の切れ込みについても、同様にして求めることが出来ます。これで、すべての真ん中の切れ込 みの位置に対して、残りの 2 つの切れ込みの位置を求めることが出来ました。あとは、実際にそれ ぞれの場合に対して答えを計算すれば、答えが求まります。<br>
# よって、O(N) でこの問題は解けました。<br>
#
# ????<br>
# わからない。
# 1. 累積和を求める
# 2. 真ん中の切る位置を決める
# 3. Lの切る位置を決める
# 4. Rの切る位置を決める
#
# この真ん中の切る位置をどう決めようか?<br>
# とりあえず,[最速](https://abc102.contest.atcoder.jp/submissions/2784349)と[二番目](https://abc102.contest.atcoder.jp/submissions/2780320)に早いコードを読むと,
# ```
# from itertools import accumulate
# ```
# していたので,これの意味を読む。(https://docs.python.jp/3/library/itertools.html#itertools.accumulate)
# <br>
#
# このモジュールは累積和のlist objectを返してくれる。
# +
from itertools import accumulate
N = input()
A = list(map(int, input().split()))
print(A)
B = accumulate(A)
# -
B
# このままだと展開できなかったので,
B = list(accumulate(A))
print(B)
# で展開できる。また,[二番目に早いコード](https://abc102.contest.atcoder.jp/submissions/2780320)を見ると
*sum_a, =accumulate(A)
print(sum_a)
# のような書き方をしていた。これは何?前のアスタリスクは,list展開してくれる[もの](https://docs.python.jp/3/tutorial/controlflow.html#unpacking-argument-lists)とか[ここ](http://or1ko.hatenablog.com/entry/20080901/1220268174)とか。カンマはないとエラー返したので必要。
# +
from itertools import accumulate
N = int(input())
A = list(map(int, input().split()))
print(A)
*B, = accumulate(A)
print(B)
# -
# ここまでで累積和まで出せた。次にコードを写経する。
# +
le, ri, ce, diff =0, 2, B[-1], B[-1]
for i in range(1, N-2):
print(i)
print(le, B[i], B[le + 1] +B[le])
print(ri)
print(ri, c + B[i], B[ri + 1] +B[ri])
while B[i] > B[le + 1] +B[le]:
le += 1
while ce + B[i] > B[ri + 1] + B[ri]:
ri += 1
p = B[le]
q = B[i] - p
r = B[ri] - q - p
s = c - B[ri]
diff = min(diff, max(p, q, r, s) - min(p, q, r, s))
print(diff)
# -
# 1. 真ん中の切れ込みを動かす。
# 2. 真ん中の切れ込みに対して,左側の切れ込みが動いていく。このとき,真ん中の切れ込みよりも大きくならないようになっている。次の値がそれまでの累積和に比べて異常に大きいならその位置から動かない。
# 3. 真ん中の切れ込みに対して,右側の切れ込みも動かす。このとき,累積和の最大値に近づくように計算を行っていく。上と似たような感じ。
# 4. p, q, r,s を計算する。
# 5. 前回の結果と,今回の差分の結果の小さい方を保存する
#
# これは自分ではかけなさそうだけど,[しゃくとり法](https://qiita.com/drken/items/ecd1a472d3a0e7db8dce)というアルゴリズムについて学べてよかった。
| ABC102.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.5 32-bit
# metadata:
# interpreter:
# hash: 1a0aef5bc7ca9123676bdf3ec95dad4f6d93924f8ab55cb688f2ea9759e5da3f
# name: python3
# ---
# # Covis-19 Impact on Airport Traffic
#
# # Kaggle's Dataset
#
# ---------------------------------------------------------------------
# ### installing libs
# !pip install plotly
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import plotly.express as px
df = pd.read_csv('covid_impact_on_airport_traffic.csv' ,engine='python')
df.head()
df.info()
df.describe()
print('Maximum Impact: ', df['PercentOfBaseline'].min() , '%')
print('Mean Impact: ', round(df['PercentOfBaseline'].mean()) , '%')
print('Minimum Impact: ', df['PercentOfBaseline'].max() , '%')
print(df['City'].unique())
print('Unique Cities: ',len(df['City'].unique()))
print(df['AirportName'].unique())
print('Unique Airports: ',len(df['AirportName'].unique()))
print(df['State'].unique())
print('Unique States: ',len(df['State'].unique()))
print(df['Country'].unique())
print('Unique Countries: ',len(df['Country'].unique()))
df['lat'] = df['Centroid'].apply(lambda x: x[6: x.find(" ")] )
df['lon'] = df['Centroid'].apply(lambda x: x[x.find(" ") : x.find(")")] )
fig = px.scatter_geo(df,
lat = 'lon',
lon = 'lat',
hover_name = 'AirportName',
color = 'PercentOfBaseline',
title = 'Covid-19 Impact on Airports Traffic',
projection = 'hammer')
fig.show()
| covid19.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # A deep geothermal doublet
#
# In this notebook, we will take a look at **geothermal doublets**, evaluating some results of a numerical doublet simulation in terms of the temporal behaviour as well as some economic calculations.
# This simulation is run with SHEMAT, a numerical code for simulating heat- and mass transfer in a porous medium. The basis is a synthetic model of a sedimentary, hydrothermal system, with implemented production and injection wells, just as in our project work for the shallow systems. The difference is the higher temperatures and flow rates, which allow producing electricity.
#
# ### Introduction
# A geothermal doublet system usually consists of -at least- two boreholes, connected via a surface application installation (such as a powerplant for producing electricity, or an installation for district heating). One geothermal well produces the hot fluid (or steam) from the subsurface. This well is called the producer. The heat of the produced fluid is used accordingly, and the significant cooler fluid is then re-injected in the geothermal reservoir. In the figure to the left, the producing well is marked as a red line, while the injecting well is marked as a blue line, representing the difference in fluid temperature.
#
# If the heat content of the produced "fluid" is large enough, i.e. *dry or wet steam* is produced, a turbine can be directly operated in one circuit. If the temperatures of the fluide are lower and do not really suffice to operate a turbine directly, a binary cycle is often installed. In such a cycle, the produced fluid heats up a secondary fluid in a heat-exchanger. This secondary fluid has a significantly lower boiling point, an can thus be used to operate a turbine instead (more details in the lecture).
#
# Operating such a system over a prolonged time will eventually cause a decrease in production temperature, as the cooling front of the re-injected water reaches the producing borehole. The point in time, where production temperature starts to decrease significantly is called a _thermal breakthrough_. In the results of a simplified doublet simulation, which we study in this notebook, we will also look for a thermal breakthrough.
#
# ## 3D reservoir model
# For assessing this geothermal system, we simulate the heat- and mass transfer in a synthetic, three dimensional model. The model consists of 4 geological units.
#
# In the plot below, we see a vertical cross-section of the model in x-direction (let's say an East-West cross-section). On the left, you can see the geological units with the inclined reseroir (light green), on the right the temperature field and some arrows indicating the flow during the operation of the doublet. Here, we can clearly see the cooler temperatures around the injection, as well as the flow towards the production well.
#
#
#
# 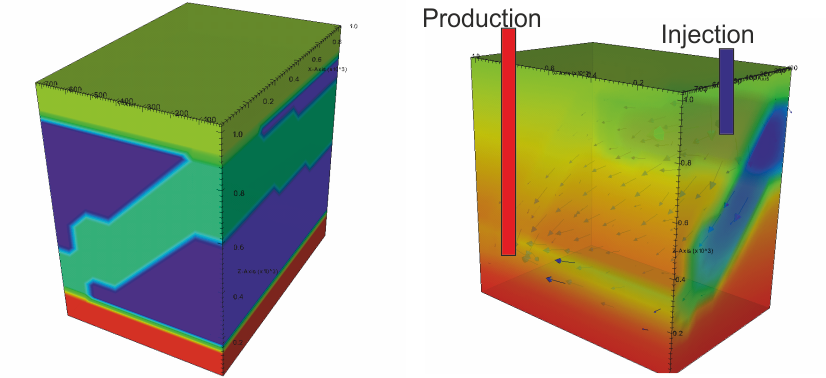
#
#
# First, we need to import the data from the monitoring files of the simulation. The model is run with different **production rates**, the first one has a production rate of 50 l per s, the second one 100 l per sec. We want to assess the dfference of both simulations, so we need to import both data sets (each from a monitoring point at the production well).
# +
# import some libraries
import numpy as np
import pandas as p
import matplotlib.pyplot as plt
# %matplotlib inline
headers = ['Time','TimeStep','Head','Vx','Vy','Vz','Por','HydrCond','Perm','Temp','ThermCond','Redox']
data1 = p.read_csv('data/sedres50.002', skiprows=2, names = headers, sep = '\s+')
data2 = p.read_csv('data/sedres100.002', skiprows=2, names = headers, sep = '\s+')
# plot the first 10 lines to check the data file...
data1.head(10)
# -
# Then, we want to plot the temperature for both production rates for the whole simulation period which is 30 years. Consider the time units wich is originally "days". You might want to convert it into "years". Which production rate seems more sustainable?
# +
T1=data1['Temp']
T2=data2['Temp']
t1=data1['Time']
t2=data2['Time']
#Days to years
d2y=1/(365.25);
ty1=t1*d2y
ty2=t2*d2y
plt.plot(ty1,T1,'-', color='red',label='50 L s$^{-1}$')
plt.plot(ty2,T2,'-', color='blue',label='100 L s$^{-1}$')
plt.legend()
plt.xlabel('Time (years)')
plt.ylabel('Temperature (°C)')
plt.grid(True)
# -
# Now we want to calculate and plot the **electric power** provided by the geothermal power plant, for this we first need to determine the **thermal power** according to
# $$ P= \Delta T\, \rho c_p\,\dot{V}$$
# Also, we need to set some values: use 998 kg m$^{-3}$ for density and 4179 J kg$^{-3}$ K$^{-1}$ for specific heat capacity of the produced fluid. For $\Delta T$, we assume a constant injection temperature of 80°C.
#
# Furthermore, we assume an efficiency of 0.1 and a *parasitic power* of 0.5 MW (used by the systems itself, mainly for the pumps).
#
# +
# production rates
Vdot1=0.05
Vdot2=0.1
# Density
rho= 998
# Specific heat capacity
cp = 4179
# Delta T
dT1=T1-80
dT2=T2-80
#Thermal power produced from wells
p1 = cp * rho * Vdot1 * dT1
p2 = cp * rho * Vdot2 * dT2
#Efficiency
eta = 0.1
#Installation consumption (MW)
ic = 0.5e6
#Electrical power
pel1 = eta * p1 - ic
pel2 = eta * p2 - ic
# plotting the electrical power (in MW)
plt.plot(ty1,pel1/1e6,'-', color='red',label='50 L s$^{-1}$')
plt.plot(ty2,pel2/1e6,'-', color='blue',label='100 L s$^{-1}$')
plt.legend()
plt.xlabel('Time (years)')
plt.ylabel('Electrical power (MW)')
plt.grid(True)
# -
# Finally, we want to calculate the produced energy, since the financial gain is calculated from selling energy at a price of 0.25 € per kWh (German feed-in tariff, EEG) and compare it with the invest of 46 million €.
# For calculating this cumulative energy, we need to know the time step size, which is stored in the monitoring file as well.
# What is the pay-off time?
# +
# Time step size for both cases (originally in days!)
dt1=data1['TimeStep']
dt2=data2['TimeStep']
# Calculating the energy produced in each time step
Eelstep1=pel1*dt1*24*60*60
Eelstep2=pel2*dt2*24*60*60
# Cumulative energy during the 30 years of operation in Joule
Eel1=np.cumsum(Eelstep1)
Eel2=np.cumsum(Eelstep2)
#Converting to kWh:
Eel1kWh=Eel1/3.6e6
Eel2kWh=Eel2/3.6e6
#Payment in Germany - EEG (Euro/kWh)
EEG=0.25
#Investment (Mio. Euro) (for plotting as an array with the same length as pel1)
invest = np.ones(len(pel1))*46;
# Gain in Mio EUR
G1=Eel1kWh*EEG/1e6
G2=Eel2kWh*EEG/1e6
# plotting the gain (in Mio EUR)
plt.plot(ty1,G1,'-', color='red',label='50 L s$^{-1}$')
plt.plot(ty2,G2,'-', color='blue',label='100 L s$^{-1}$')
plt.plot(ty1,invest,'--', color='black',label='Investment')
plt.legend()
plt.xlabel('Time (years)')
plt.ylabel('Gain (Mio. Euro))')
plt.grid(True)
# -
| Deep Geothermal_Doublet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="rSJGV02ugjrE"
# # <NAME>
#
# # TASK-2 :Prediction Using Unsupervised ML
# From the given ‘Iris’ dataset, predict the optimum number of clusters
# and represent it visually.
# + id="_LaplIzdgzf6"
# Importing the libraries
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
from sklearn import datasets
from sklearn.cluster import KMeans
# + colab={"base_uri": "https://localhost:8080/", "height": 203} id="1yRRBIExTdEC" outputId="7c7a5ce0-fee1-4cc3-88fd-2854ee6e6fc3"
# Load and Read the iris dataset
iris = datasets.load_iris()
df = pd.DataFrame(iris.data, columns = iris.feature_names)
df.head()
# + colab={"base_uri": "https://localhost:8080/"} id="n9662G0ItOH0" outputId="1d81786a-7dcd-4aa3-f7bb-0fb9c5c48225"
# To know number of rows and columns
df.shape
# summary of dataset
df.info()
# checking null values in data
df.isnull().sum()
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="gKinVFq7hDZC" outputId="df6a6b15-78f4-454d-c216-ce10bc3a9db7"
# Finding the optimum number of clusters for k-means classification
x = df.iloc[:, [0, 1, 2, 3]].values
wcss = []
for i in range(1, 11):
kmeans = KMeans(n_clusters = i, init = 'k-means++',
max_iter = 300, n_init = 10, random_state = 0)
kmeans.fit(x)
wcss.append(kmeans.inertia_)
# Plotting the elblow curve to find no of clusters
plt.plot(range(1, 11), wcss,color="red",marker="*")
plt.title('The Elbow Method')
plt.xlabel('Number of clusters')
plt.ylabel('WCSS') # Within cluster sum of squares
plt.grid(alpha=0.45)
plt.show()
# + [markdown] id="5AT8Hmj_wZLX"
#
# + [markdown] id="satoifK7hY8u"
# From the above graph, we observe that the optimum clusters is where the elbow occurs. This is when the within cluster sum of squares (WCSS) doesn't decrease significantly with every iteration.From this ,we choose the number of clusters as 3.
#
#
# + id="d_b-IA55hJBC"
# Applying kmeans to the dataset
kmeans = KMeans(n_clusters = 3, init = 'k-means++',
max_iter = 300, n_init = 10, random_state = 0)
y_kmeans = kmeans.fit_predict(x)
# + colab={"base_uri": "https://localhost:8080/", "height": 268} id="BnqSQOWOhMY7" outputId="8f9534b7-00cf-4fbc-f235-1dc5fb8698d9"
# Visualising the clusters
plt.scatter(x[y_kmeans == 0, 0], x[y_kmeans == 0, 1],
s = 100, c = 'green', label = 'Iris-setosa')
plt.scatter(x[y_kmeans == 1, 0], x[y_kmeans == 1, 1],
s = 100, c = 'purple', label = 'Iris-versicolour')
plt.scatter(x[y_kmeans == 2, 0], x[y_kmeans == 2, 1],
s = 100, c = 'red', label = 'Iris-virginica')
# Plotting the centroids of the clusters
plt.scatter(kmeans.cluster_centers_[:, 0], kmeans.cluster_centers_[:,1],
s = 100, c = 'yellow', label = 'Centroids')
plt.legend()
plt.grid( alpha=0.25)
plt.show()
| Task-2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import numpy as np
vector = np.array([1,2,3])
print(vector)
print("")
mat = np.array([[1,2,3],[1,3,3]])
print(mat)
print("")
vec2 = np.arange(0, 20, 4)
vec3 = np.linspace(0, 30, 3)
print(vec2)
print(vec3)
print("")
tempArray = [2, 4, 6, 8]
print(tempArray)
# Convert array to matrtix
print(vec3.reshape(3, 1))
# -
matrixOne = np.array([range(0,5), range(5, 10)])
m = np.matrix([range(0,5), range(5, 10)])
print(m * m.getT())
print(np.arange(0, 12).reshape(4, 3))
print([1, True, "Hi"])
np.cos(np.pi)
np.sqrt(1.44)
np.log(6)
np.exp(4)
print(np.arange(4, 20, 5))
print(np.arange(10, 20)[np.arange(0, 10, 3)])
# +
array1 = np.array([range(0,5), range(5, 10)])
array2 = array1
print(array1)
array2 = array1 * array1
print("")
print(array1)
print("")
print(array2)
# -
matrixOne = np.array([[1, 2, 3], [4, 5, 6], [7, 8, 9]])
print(matrixOne)
print(matrixOne[1, 1])
vec = np.array([4, 7, 8, 9, 10, 6, 1])
print(vec[vec>6])
# +
import pandas as pd
score = [10, 15, 20, 25]
print(pd.Series(data=score, index = ['a','b','c','d']))
# -
df2 = pd.DataFrame(np.array([[1, 2, 3], [4, 5, 6], [7, 8, 9]]),
columns=['a', 'b', 'c'])
newColumn = [2, 2, 2]
df2['c'] = newColumn
print(df2)
print(np.arange(0, 22, 6))
# +
demo_array = np.arange(10,21)
subset_demo_array = demo_array[0:7]
subset_demo_array[:]= 101
print(subset_demo_array)
# +
flowers = pd.Series([2, 3, 5, 4], index=['lily', 'rose', 'daisy', 'lotus'])
print(flowers['daisy'])
print(df2.drop('c', axis=1))
# -
s1 = pd.Series(['a', 'b'])
s2 = pd.Series(['c', 'd'])
print(pd.concat([s1,s2]))
| src/Dynamic Programming/Practice.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (aws)
# language: python
# name: aws
# ---
from pprint import pprint
import json
from dask_cloudprovider import FargateCluster
cluster = FargateCluster(n_workers=1)
cluster.scale(2) # Sets the total number of workers to 2
cluster.dashboard_link
from dask.distributed import Client
client = Client(cluster)
import dask.array as da
arr = da.random.random((2000, 1000, 10000), chunks=(100, 100, 1000))
arr = arr.mean().persist()
arr.compute()
client.close()
cluster.close()
del cluster
| rapids_ecs/Dask and AWS Fargate.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="8iFBj0zfqDHF"
import numpy as np
from docplex.mp.model import Model
from qiskit import BasicAer
from qiskit.aqua import QuantumInstance
from qiskit.aqua import aqua_globals
from qiskit.aqua.algorithms import VQE, NumPyMinimumEigensolver, QAOA, NumPyEigensolver
from qiskit.aqua.components.optimizers import SPSA
from qiskit.circuit.library import RealAmplitudes
from qiskit.optimization import QuadraticProgram
from qiskit.optimization.algorithms import MinimumEigenOptimizer, RecursiveMinimumEigenOptimizer, \
MinimumEigenOptimizationResult
from qiskit.optimization.converters import QuadraticProgramToQubo
from qiskit.optimization.applications.ising import docplex
import matplotlib.pyplot as plt
# + id="6Weyv35il5ac"
machines_times = [3, 6, 2]
machines_costs = [2, 5, 1]
tasks = [6, 12, 18, 24]
paths = [[0, 1, 3], [0, 2, 3]]
d = 14
# + id="FcawJFAUklUP"
def get_time_matrix():
r = []
for i in machines_times:
tmp = []
for j in tasks:
tmp.append(j / i)
r.append(tmp)
return np.array(r)
def get_cost_matrix(tasks_execution_times):
m = []
for i in range(len(tasks_execution_times)):
tmp = []
for j in tasks_execution_times[i]:
tmp.append(machines_costs[i] * j)
m.append(tmp)
return m
# + id="vsguKC4XhGOL"
time_matrix = np.array(get_time_matrix())
cost_matrix = np.array(get_cost_matrix(time_matrix))
# + colab={"base_uri": "https://localhost:8080/"} id="w10-zz2vhdEp" outputId="3bc205e5-73ac-4e1c-cfbd-cc4deb7c305b"
print("Time matrix:\n {}".format(time_matrix))
# + colab={"base_uri": "https://localhost:8080/"} id="_lsxEuMHhZrh" outputId="59473a70-6385-4702-9885-a29798110bb8"
print("Cost matrix:\n {}".format(cost_matrix))
# + [markdown] id="vZRN7hczkixC"
#
# In this specific situation there are:
#
#
# 1. two qubits for each task: **8 qubits**,
# 2. first path minimum time is *1+2+4=7*. *log2(14-7)=log2(7)*: **3 qubits**,
# 3. second path minimum time is *1+3+4=8*. *log2(14-8)=log2(6)*: **3 qubits**.
#
#
# In total, the model has **14 variables**.
# + [markdown] id="cqDR-_9CjMsV"
# Model with:
#
#
# 1. 13 variables needs 4 GiB of RAM
# 2. 14 variables needs 16 GiB of RAM
# 3. 15 variables needs 64 GiB of RAM
#
#
# + id="N-StWiujfbok"
# Correct only for three machines!
correct_machines = ['00', '01', '11']
machine_to_index = {'00': 0, '01': 1, '11': 2}
def get_task_subvector(vector, task_index):
x = len(machines_costs) - 1 # When 3 machines, then subvector for each task has length of 2.
subvector = ''
for i in range(2 * task_index, 2 * task_index + x):
bit_value = str(int(vector[i]))
subvector += bit_value
return subvector
def solution_vector_correct(vector):
for i in range(len(tasks)):
if get_task_subvector(vector, i) not in correct_machines:
return False
return True
def execution_times_not_bigger_than_deadline(vector):
for path in paths:
path_time_sum = 0
for task in path:
task_machine = machine_to_index.get(get_task_subvector(vector, task))
path_time_sum += time_matrix[task_machine, task]
if path_time_sum > d:
return False
return True
def is_solution_correct(vector):
return solution_vector_correct(vector) and execution_times_not_bigger_than_deadline(vector)
# + id="yJjzRI1os2A0"
def print_solutions_with_inreasing_energy(hamiltonian):
eigensolver = NumPyEigensolver(hamiltonian, 100)
eigensolver_result = eigensolver.compute_eigenvalues()
print("Vector\t\t\tEnergy\t\t\tCorrect?")
for eigenstate, eigenvalue in zip(eigensolver_result.eigenstates, eigensolver_result.eigenvalues):
eigenstate, = eigenstate.sample().keys()
eigenstate = eigenstate[::-1]
print("{}\t\t{}\t\t{}".format(eigenstate, eigenvalue, is_solution_correct(eigenstate)))
# + id="NFRfzo4zg70N"
def get_quadratic_problem():
tasks_count = len(tasks)
machines_count = len(machines_costs)
mdl = Model(name='workflow')
x = {(i, j): mdl.binary_var(name='x_{0}_{1}'.format(i, j))
for i in range(0, tasks_count) for j in range(0, machines_count - 1)}
objective = mdl.sum(cost_matrix[2, i] * x[(i, 0)]
+ cost_matrix[1, i] * (x[(i, 1)] - x[(i, 0)]) ** 2
+ cost_matrix[0, i] * (1 - x[(i, 1)]) for i in range(0, tasks_count))
mdl.minimize(objective)
for k in range(0, len(paths)):
mdl.add_constraint(mdl.sum([time_matrix[2, i] * x[(i, 0)]
+ time_matrix[1, i] * (x[(i, 1)] - x[(i, 0)])
+ time_matrix[0, i] * (1 - x[(i, 1)])
for i in paths[k]]) <= d, "deadline_path_{}".format(k))
qp = QuadraticProgram()
qp.from_docplex(mdl)
converter = QuadraticProgramToQubo()
qubo = converter.convert(qp)
H, _ = qubo.to_ising()
print_solutions_with_inreasing_energy(H)
return qp
# + colab={"base_uri": "https://localhost:8080/"} id="2_LOjo1WhKKR" outputId="fa3e7332-1f56-42ca-b2f3-b9b7cd28b72c"
operator = get_quadratic_problem()
# + id="pMRkfXTbhPuH"
aqua_globals.random_seed = 10598
quantum_instance = QuantumInstance(BasicAer.get_backend('statevector_simulator'),
seed_simulator=aqua_globals.random_seed,
seed_transpiler=aqua_globals.random_seed)
reps = 2
max_trials = 1000
entanglement = 'full'
exact_mes = NumPyMinimumEigensolver()
spsa = SPSA(maxiter=max_trials)
ry = RealAmplitudes(operator.get_num_binary_vars(), reps=reps, entanglement=entanglement)
vqe_mes = VQE(quantum_instance=quantum_instance, var_form=ry, optimizer=spsa)
qaoa_mes = QAOA(quantum_instance=quantum_instance, initial_point=[0., 0.])
# + id="niqMoFlQfeTm"
exact = MinimumEigenOptimizer(exact_mes)
vqe = MinimumEigenOptimizer(vqe_mes)
qaoa = MinimumEigenOptimizer(qaoa_mes)
recursive = RecursiveMinimumEigenOptimizer(min_eigen_optimizer=qaoa, min_num_vars=8, min_num_vars_optimizer=exact)
# + id="SSYGdE1bg6Ns"
def perform_optimization(optimization, qubit_op, name):
result = optimization.solve(qubit_op)
print_optimization_output(result, name)
def print_optimization_output(result, solver_name):
print("------ %s ------" % solver_name)
print(result)
print("Is the found vector correct? : {}".format(is_solution_correct(result.x)))
if type(result) is MinimumEigenOptimizationResult:
print("Most probable sample: {}".format(max(result.samples, key=lambda item: item[2])))
print("Lowest energy smple: {}".format(min(result.samples, key=lambda item: item[1])))
print("Samples count: {}\n\n".format(len(result.samples)))
plt.figure(figsize=(20,10))
for vector, energy, _ in result.samples:
color = "green" if is_solution_correct(vector) else "red"
plt.axvline(x=energy, color=color)
plt.xlabel("Energy")
plt.ylabel("")
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 823} id="wtHeG0hKgbcp" outputId="0d32a293-0d8a-4f5e-ebc4-acf50a1c1327"
# %%time
perform_optimization(exact, operator, "Numpy Eigensolver")
# + colab={"base_uri": "https://localhost:8080/", "height": 823} id="G9sOVKr_gbgk" outputId="add785f1-5615-4174-c532-179d0ea4c67c"
# %%time
perform_optimization(vqe, operator, "VQE")
# + id="awwxnWbNgahT" colab={"base_uri": "https://localhost:8080/", "height": 823} outputId="075a8179-72a0-49c4-ee88-bb842953bd41"
# %%time
perform_optimization(qaoa, operator, "QAOA")
# + id="dt64oVdqgZVZ" colab={"base_uri": "https://localhost:8080/"} outputId="aa376e4e-8e95-4296-ae43-095127085dff"
# %%time
perform_optimization(recursive, operator, "Recursive")
| domain-wall-encoding/minimum_eigen_optimizer_with_docplex.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import tensorflow as tf
import numpy as np
import keras
import pandas as pd
x_train = pd.read_csv('trainingfeatures.csv').drop(columns=['Unnamed: 0'])
y_train = pd.read_csv('traininglabels.csv').drop(columns=['Unnamed: 0'])
x_test = pd.read_csv('testingfeatures.csv').drop(columns=['Unnamed: 0'])
y_test = pd.read_csv('testinglabels.csv').drop(columns=['Unnamed: 0'])
temp_x_train=[]
for row in x_train.iterrows():
index, data = row
temp_x_train.append(data.tolist())
temp_y_train=[]
for row in y_train.iterrows():
index, data = row
temp_y_train.append(data.tolist())
temp_x_test=[]
for row in x_test.iterrows():
index, data = row
temp_x_test.append(data.tolist())
temp_y_test=[]
for row in y_test.iterrows():
index, data = row
temp_y_test.append(data.tolist())
# -
# ## Part B Change Activation function
# Activation = **relu** + **tanh** + **tanh**
#
# Loss = cross_entropy
# +
x= np.array(temp_x_train)
y=np.array(keras.utils.to_categorical(y_train))
# +
num_input = 3 # MNIST data input
num_classes = 4 # MNIST total classes (0-3 digits)
EPOCHS = 10000
BATCH_SIZE = 1000
display_step = 500
with tf.name_scope('Inputs_B'):
X = tf.placeholder("float", [None, num_input],name='Features_B')
Y = tf.placeholder("float", [None, num_classes],name='Label_B')
# using two numpy arrays
features, labels = (X, Y)
# make a simple model
def Neuron(x):
with tf.name_scope('layer1_B'):
net = tf.layers.dense(x, 100, activation=tf.nn.relu)
with tf.name_scope('layer2_B'):
net = tf.layers.dense(net, 50, activation=tf.tanh)
with tf.name_scope('layer3_B'):
net = tf.layers.dense(net, 20, activation=tf.tanh)
with tf.name_scope('out_layer_B'):
prediction = tf.layers.dense(net, 4)
return prediction
prediction = Neuron(X)
loss = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(logits=prediction, labels=Y))
tf.summary.scalar('loss_B',loss)
#tf.losses.mean_squared_error(prediction, y) # pass the second value
correct_pred = tf.equal(tf.argmax(prediction, 1), tf.argmax(Y, 1))
accuracy = tf.reduce_mean(tf.cast(correct_pred, tf.float32))
tf.summary.scalar('acuracy_B',accuracy)
#from iter.get_net() as label
train_op = tf.train.AdamOptimizer().minimize(loss)
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
sess.run(tf.local_variables_initializer())
merge_summary= tf.summary.merge_all()
writer = tf.summary.FileWriter('files/B')
writer.add_graph(sess.graph)
for i in range(EPOCHS):
_, loss_value,acc_value = sess.run([train_op, loss,accuracy],feed_dict={X: x, Y: y})
if i% display_step == 0:
print("Iter: {}, Loss: {:.4f}".format(i+1, loss_value))
print("Accurancy: " +str(acc_value))
print(x.shape)
summary=sess.run(merge_summary,feed_dict={X:x, Y:y})
writer.add_summary(summary,i)
correct_pred = tf.equal(tf.argmax(prediction, 1), tf.argmax(Y, 1))
accuracy = tf.reduce_mean(tf.cast(correct_pred, tf.float32))
print("Test accuracy: "+ str(accuracy.eval({X: np.array(temp_x_test), Y: np.array(keras.utils.to_categorical(y_test))})))
# -
# * Change the activation function. How does it effect the accuracy?
#
# **Answer:** The Auccracy gets increased.
#
# * How does it effect how quickly the network plateaus?
#
# **Answer:** The plateaus gets shorter.
| Assignment 5/Assignment 5/partB.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 回顾
#
# - 因为Torch的核心是Tensor,Tensor的数据由Storage管理,所以这两个类的关系搞清楚,就可以使用Tensor了。
#
# ## Tensor的Python构造器定义如下
# ```python
# Tensor.__init__(torch.device device)
# Tensor.__init__(torch.Storage storage)
# Tensor.__init__(Tensor other)
# Tensor.__init__(tuple of ints size, torch.device device)
# Tensor.__init__(object data, torch.device device)
# ```
# ## Storage的Python构造器定义如下
# ```python
# FloatStorage.__init__() no arguments
# FloatStorage.__init__(int size)
# FloatStorage.__init__(Sequence data)
# FloatStorage.__init__(torch.FloatStorage view_source)
# FloatStorage.__init__(torch.FloatStorage view_source, int offset)
# FloatStorage.__init__(torch.FloatStorage view_source, int offset, int size)
# ```
# # 关于Tensor
# ## Tensor与Numpy
# - 实际上按照Python一贯的思路,会提供很多函数来替代构造器的使用,这样做有两个原因:
# - 个性化,方便,简单;
# - 使用工厂模式来创建对象,符合软件的常见设计模式,Python大量采用;
# - 今后不要动不动就说面向对象最好,最方便。最直观,最方便的还是函数,拿来就用,不需要构建对象才能使用。
#
# - Torch号称是GPU版本的Numpy,Numpy有的Tensor都有,所以按照Numpy的思路,在构建好对象后,有三大块功能是需要数理下的,掌握这三大基础功能,后面的内容就容易理解:
# 1. 基本属性
# - 了解对象的内存与数据结构
# 2. 基本操作
# - 数据进出
# 3. 数学运算
# - 构建数据对象的最终目的就是计算;
# - 计算的类别很多,基本数学运算,随机采样,线性代数的矩阵运算,统计计算,......
# - 这里先明白基本的数学运算。
# ## Tensor的官方文档结构
# ```python
# torch
# Tensors
# Creation Ops
# Indexing, Slicing, Joining, Mutating Ops
# Generators
# Random sampling
# In-place random sampling
# Quasi-random sampling
# Serialization
# Parallelism
# Locally disabling gradient computation
# Math operations
# Pointwise Ops
# Reduction Ops
# Comparison Ops
# Spectral Ops
# Other Operations
# BLAS and LAPACK Operations
# Utilities
# ```
# - 这里先搞定Tensor本身的基本属性与操作
# - 基本属性(从C/C++文档对应)
# - 基本操作
# - Indexing(索引访问操作)
# - Slicing (切片访问操作【是索引的批量级升级版本】)
# - Joining(数据组合与合并)
# - Mutating Ops(数据访问:索引与切片的函数版本)
# - 数学运算:
# - Pointwise Ops(元素运算)
# - Reduction Ops(降维运算)
# - Comparison Ops(比较运算)
# - Spectral Ops(谱运算)
# - Other Operations(其他运算)
# - BLAS and LAPACK Operations(线性代数运算)
# 1. BLAS
# - Basic Linear Algebra Subprograms(Fortran语言编写,Fortran史上经典古老的数学计算语言);
# 2. LAPACK
# - Linear Algebra Package,底层使用的也是BLAS;
# 3. ATLAS
# - Automatically Tuned Linear Algebra Software;
# 4. OpenBLAS:
# - 在编译时根据目标硬件进行优化,生成运行效率很高的程序或者库。OpenBLAS的优化是在编译时进行的,所以其运行效率一般比ATLAS要高。因此OpenBLAS对硬件的依赖比较高,换一个硬件平台可能会重新进行编译。
# 5. cuBLAS与ACML:
# - Intel的MKL和AMD的ACML都是在BLAS的基础上,针对自己特定的CPU平台进行针对性的优化加速。以及NVIDIA针对GPU开发的cuBLAS。
# # Tensor的基本属性与属性函数
# - 先构建一个张量(Tensor)使用;
# +
import torch
t_vector = torch.LongTensor(
data= [1, 2, 3, 4, 5]
)
print(t_vector)
t_matrix = torch.LongTensor(
data= [
[1, 2, 3, 4],
[5, 6, 7, 8]
]
)
print(t_matrix)
# -
# ## 属性
# ### 属性-T
#
# - 返回Tensor的转置;
print(t_vector.T) # 向量转置还是本身,不产生转置效果
print(t_matrix.T)
# ### 属性-data
#
# - 返回张量的数据, 返回的也是张量,就是张量本身;
# - 返回不同的id;
# - 共享同一个Stroage;
# - 但是data返回的数据状态改变:require s_grad = False,就是不能求导。
# 地址不同
print(t_vector.data)
print(type(t_vector.data))
print(id(t_vector), id(t_vector.data))
# 数据相互影响
d = t_vector.data
t_vector[2] =88
print(d)
# +
# data与原张量的差异
# -
# ### 属性-dtype
#
# - Tensoor元素类型
print(t_vector.dtype)
# ### 属性-grad,grad_fn,requires_grad
#
# - 导数:
# - 默认是None
# - 调用backward计算导数,导数是累加的。如果每次单独计算,需要清空;
# - 导数的计算需要导数函数grad_fn(没有指定函数的张量无法计算导数)。
# - grad_fn函数自动跟踪,需要设置requires_grad=True
# 1. grad属性
print(t_vector.grad)
t_vector.backward()
# 2. 属性-grad_fn
# - 张量所在的导数函数
# +
t1 = torch.Tensor([1.0])
t2 = torch.Tensor([1.0])
t3 = t1 + t2
print(t3)
print(t3.grad_fn)
# -
# 3. requires_grad属性
t1.requires_grad=True
t2.requires_grad=True
t4 = t1 + t2
print(t4)
print(t4.grad_fn)
print(type(t4.grad_fn))
print(t4.requires_grad)
print(t1.grad_fn)
print(t4.grad)
t4.backward()
print(t4.grad) # 没有导数
print(t1.grad) # t1与t2导数(偏导数)
print(t2.grad)
print(t1.grad_fn)
# ### 属性-is_cuda,device
#
# - 判断是否是cuda计算(GPU计算)
# - device使用专门的类构造;
t1 = torch.Tensor([2.5])
print(t1.is_cuda)
t2 = torch.Tensor([2.5], device=torch.device('cpu:0'))
t2 = torch.Tensor([2.5], device=torch.device('cpu'))
print(t2.is_cuda)
t3 = torch.Tensor([2.5], device=torch.device('cuda'))
print(t3.is_cuda) # 苹果电脑不支持,请在Nvidia的显卡上运算,其他支持GPU运算的电脑上运行
# 在window上还需要安装厂商驱动
# 判定电脑是否之处GPU运算
print(torch.cuda.is_available())
# ### 属性-is_leaf,grad与retain_grad函数
#
# - 这个属性用来判定张量Tensor是否是Leaf Tensor,下面两种情况都应该是Leaf Tensor:
# - 属性requires_grad为False的。
# - 属性requires_grad=True,但是用户构建的Tensor,表示该张量不是计算结果,而是用户构建的初始张量。
#
# - 运行backward后,仅仅只有Leaf Tensor在才会有grad属性。如果非Leaf Tensor需要具有grad属性,需要使用retain_grad函数开启grad属性。
# +
# 演示叶子Tensor与grad,backward的关系
import torch
t1 = torch.Tensor([1.0]) # 用户构建的都是Leaf Tensor
t1.requires_grad=True
t2 = torch.Tensor([2.0])
t2.requires_grad=True
t3 = t1 + t2
t3.backward()
print(t1.is_leaf, t2.is_leaf, t3.is_leaf)
print(t1.grad) # Leaf Tensor的grad属性由backward函数产生。
# +
# 演示Non-Leaf Tensor 与 retain_grad的关系
import torch
t1 = torch.Tensor([1.0]) # 用户构建的都是Leaf Tensor
t1.requires_grad=True
t2 = torch.Tensor([2.0])
t2.requires_grad=True
t3 = t1 + t2
t3.retain_grad() # 调用该函数后,t3才有grad属性,可以注释这个语句体验
t3.backward()
print(t1.is_leaf, t2.is_leaf, t3.is_leaf)
print(t3.grad)
# -
# ### 属性-ndim与dim函数
#
# - Tensor的维度
# +
import torch
t1 = torch.Tensor([1.0, 20]) # 用户构建的都是Leaf Tensor
t2 = torch.Tensor(
[
[2.0, 1.0],
[1.0, 2.0]
]
)
print(t1.ndim) # 1 维
print(t2.ndim) # 2 维
print(t2.dim())
# -
# ### 属性-shape与size函数
#
# - Tensor的形状,与size函数一样
# +
import torch
t1 = torch.Tensor([1.0, 20]) # 用户构建的都是Leaf Tensor
t2 = torch.Tensor(
[
[2.0, 1.0],
[1.0, 2.0]
]
)
print(t2.shape) # 属性shape
print(t2.size()) # 函数size()
# -
# ### 属性-is_sparse
#
# - 是否稀疏张量:
# - 在矩阵中,若数值为0的元素数目远远多于非0元素的数目,并且非0元素分布没有规律时,则称该矩阵为稀疏矩阵;与之相反,若非0元素数目占大多数时,则称该矩阵为稠密矩阵。定义非零元素的总数比上矩阵所有元素的总数为矩阵的稠密度。
#
# - is_sparse该属性是只读,不可写的
# - 稀疏张量提供专门的API产生。
#
#
# - 稀疏张量有自己的构造规则:
# - 稀疏张量被表示为一对致密张量:一维张量和二维张量的索引。可以通过提供这两个张量来构造稀疏张量,以及稀疏张量的大小。
# +
# 默认的张量都是稠密张量
import torch
t1 = torch.Tensor([0, 0]) # 用户构建的都是Leaf Tensor
t2 = torch.Tensor(
[
[1, 0],
[0, 0]
]
)
print(t1.is_sparse) # 属性shape
print(t2.is_sparse) # 函数size()
t3 = torch.Tensor(1000,1000)
t3.fill_(0)
t3[0,0]=1
print(t3.is_sparse)
print(t3)
t3.is_sparse=True # 不能修改该属性,该属性是只读,不可写的。
# -
# 稀疏矩阵
import torch
ts = torch.sparse.FloatTensor(2, 3)
print(ts.is_sparse)
print(ts)
print(ts.to_dense())
# ### 属性-layout
#
# - 张量Tensor使用Storage表示都是一维的,其构成张量只要采用布局计算。这个布局使用layout属性设置
# - 一般都是采用strided
# - 稀疏矩阵的布局使用的是:`torch.sparse_coo`
#
# - 目前常用的就是这两种布局layout。
import torch
t1 = torch.Tensor([0, 0]) # 用户构建的都是Leaf Tensor
t2 = torch.Tensor(
[
[1, 0],
[0, 0]
]
)
print(t1.layout, t2.layout)
# ### 属性-output_nr
#
# - 在反向传播中存放输出。
# - 具体用途先存疑。
import torch
t1 = torch.Tensor([0, 0]) # 用户构建的都是Leaf Tensor
t2 = torch.Tensor(
[
[1, 0],
[0, 0]
]
)
print(t2.output_nr)
# +
# 演示Non-Leaf Tensor 与 retain_grad的关系
import torch
t1 = torch.Tensor([2.0]) # 用户构建的都是Leaf Tensor
t1.requires_grad=True
t2 = torch.Tensor([2.0])
t2.requires_grad=True
t3 = t1.sin()
print(t1.output_nr, t2.output_nr, t2.output_nr)
t3.retain_grad() # 调用该函数后,t3才有grad属性,可以注释这个语句体验
t3.backward()
print(t3)
print(t1.is_leaf, t2.is_leaf, t3.is_leaf)
print(t3.grad)
print(t1.output_nr, t2.output_nr, t2.output_nr)
# -
# ### 属性-其他
#
# - is_mkldnn:intel提供的加速CPU运算的方法,判定是否CPU加速
# - is_quantized:是否被量化(量化指将信号的连续取值近似为有限多个离散值)
# - name:张量名
# - volatile:新版本已经停用;
#
# +
# 演示Non-Leaf Tensor 与 retain_grad的关系
import torch
t1 = torch.Tensor([2.0]) # 用户构建的都是Leaf Tensor
t1.requires_grad=True
t2 = torch.Tensor([2.0])
t2.requires_grad=True
t3 = t1 + t2
print(t1.is_mkldnn)
print(t2.name)
print(t2.is_quantized)
# -
# # 附录:mkldnn的使用
# 1. 下载地址
# - `https://github.com/intel/mkl-dnn`
# 2. 安装
# - cmake安装,直接套路
# 3. 如果torch不支持mkldnn,就需要使用源代码重新安装!
# ----
#
| 东南大学/D02机器学习与深度学习/Torch基础/02Tensor-01Tensor的基本属性.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="9_ZORURKg-fp"
# # Lab 11: Dialogue Act Tagging
#
# Dialogue act (DA) tagging is an important step in the process of developing dialog systems. DA tagging is a problem usually solved by supervised machine learning approaches that all require large amounts of hand labeled data. A wide range of techniques have been investigated for DA tagging. In this lab, we explore two approaches to DA classification. We are using the Switchboard Dialog Act Corpus for training.
# Corpus can be downloaded from http://compprag.christopherpotts.net/swda.html.
#
# + [markdown] colab_type="text" id="ziKyA9R4gyw9"
# The downloaded dataset should be kept in a data folder in the same directory as this file.
# + colab_type="code" id="jmTpKt_uefe5" colab={"base_uri": "https://localhost:8080/", "height": 33} outputId="66041b81-6499-48cc-d161-7adcd5f363a1"
import pandas as pd
import glob
from keras.preprocessing.sequence import pad_sequences
from sklearn.model_selection import train_test_split
import numpy as np
import sklearn.metrics
import tensorflow as tf
import matplotlib.pyplot as plt
from tqdm import tqdm_notebook as tqdm
# + [markdown] id="l7UKWcnjQ3HR" colab_type="text"
# Upload the swda.zip to google colab and then run the next cell
# + id="OoSuNgG7iIZg" colab_type="code" outputId="02c68b48-6d72-414f-b7e4-ee8f51c098a3" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# !unzip swda.zip
# + colab_type="code" id="6E8axaw1hAbM" colab={}
f = glob.glob("swda/sw*/sw*.csv")
frames = []
for i in range(0, len(f)):
frames.append(pd.read_csv(f[i]))
result = pd.concat(frames, ignore_index=True)
# + colab_type="code" id="b7hKGF7EhM4s" outputId="47325045-63f2-4d66-ef8b-2e679f9ba89c" colab={"base_uri": "https://localhost:8080/", "height": 33}
print("Number of converations in the dataset:",len(result))
# + [markdown] colab_type="text" id="0ttyB2lQhc7B"
# The dataset has many different features, we are only using act_tag and text for this training.
#
# + colab_type="code" id="-jUifIdshhD0" colab={}
reduced_df = result[['act_tag','text']]
# + [markdown] colab_type="text" id="-iPmZvysqg2i"
# Reduce down the number of tags to 43 - converting the combined tags to their generic classes:
# + colab_type="code" id="MQuHm0jPt_lz" colab={}
# Imported from "https://github.com/cgpotts/swda"
# Convert the combination tags to the generic 43 tags
import re
def damsl_act_tag(input):
"""
Seeks to duplicate the tag simplification described at the
Coders' Manual: http://www.stanford.edu/~jurafsky/ws97/manual.august1.html
"""
d_tags = []
tags = re.split(r"\s*[,;]\s*", input)
for tag in tags:
if tag in ('qy^d', 'qw^d', 'b^m'): pass
elif tag == 'nn^e': tag = 'ng'
elif tag == 'ny^e': tag = 'na'
else:
tag = re.sub(r'(.)\^.*', r'\1', tag)
tag = re.sub(r'[\(\)@*]', '', tag)
if tag in ('qr', 'qy'): tag = 'qy'
elif tag in ('fe', 'ba'): tag = 'ba'
elif tag in ('oo', 'co', 'cc'): tag = 'oo_co_cc'
elif tag in ('fx', 'sv'): tag = 'sv'
elif tag in ('aap', 'am'): tag = 'aap_am'
elif tag in ('arp', 'nd'): tag = 'arp_nd'
elif tag in ('fo', 'o', 'fw', '"', 'by', 'bc'): tag = 'fo_o_fw_"_by_bc'
d_tags.append(tag)
# <NAME> says (p.c.) that it makes sense to take the first;
# there are only a handful of examples with 2 tags here.
return d_tags[0]
# + colab_type="code" id="S8N_PUCAblq3" outputId="cc55223a-aa98-4719-e71b-6be14b7fbd6f" colab={"base_uri": "https://localhost:8080/", "height": 117}
reduced_df["act_tag"] = reduced_df["act_tag"].apply(lambda x: damsl_act_tag(x))
# + [markdown] colab_type="text" id="0UNy0vvhhqpD"
# There are 43 tags in this dataset. Some of the tags are Yes-No-Question('qy'), Statement-non-opinion('sd') and Statement-opinion('sv'). Tags information can be found here http://compprag.christopherpotts.net/swda.html#tags.
#
# + [markdown] colab_type="text" id="9biiyP8UiGDe"
# To get unique tags:
# + colab_type="code" id="BrhW8gyLfQQK" colab={}
unique_tags = set()
for tag in reduced_df['act_tag']:
unique_tags.add(tag)
# + colab_type="code" id="LMOX5KwgiPmu" colab={}
one_hot_encoding_dic = pd.get_dummies(list(unique_tags))
# + colab_type="code" id="ZPHPCxE3iPby" colab={}
tags_encoding = []
for i in range(0, len(reduced_df)):
tags_encoding.append(one_hot_encoding_dic[reduced_df['act_tag'].iloc[i]])
# + [markdown] colab_type="text" id="LVI8QyVzjqWh"
# The tags are one hot encoded.
# + [markdown] colab_type="text" id="SQJTiffPjUtu"
# To create sentence embeddings:
# + colab_type="code" id="PmkyD1TfjWGO" colab={}
sentences = []
for i in range(0, len(reduced_df)):
sentences.append(reduced_df['text'].iloc[i].split(" "))
# + colab_type="code" id="MlD6L6e3jV-7" colab={}
wordvectors = {}
index = 1
for s in sentences:
for w in s:
if w not in wordvectors:
wordvectors[w] = index
index += 1
# + colab_type="code" id="e7_cjDHrjV1c" colab={}
# Max length of 137
MAX_LENGTH = len(max(sentences, key=len))
# + colab_type="code" id="LX6DidEvjVWs" colab={}
sentence_embeddings = []
for s in sentences:
sentence_emb = []
for w in s:
sentence_emb.append(wordvectors[w])
sentence_embeddings.append(sentence_emb)
# + [markdown] colab_type="text" id="Nr4iEyNTjmlu"
# Then we split the dataset into test and train.
# + colab_type="code" id="GiNZ-iI_jnOF" colab={}
from sklearn.model_selection import train_test_split
import numpy as np
X_train, X_test, y_train, y_test = train_test_split(sentence_embeddings, np.array(tags_encoding))
# + [markdown] colab_type="text" id="_RqMeWe_jron"
# And pad the sentences with zero to make all sentences of equal length.
#
# + colab_type="code" id="yqD7DvzRGRY7" colab={}
MAX_LENGTH = 137
# + colab_type="code" id="Ai9cwv82jufe" colab={}
from keras.preprocessing.sequence import pad_sequences
train_sentences_X = pad_sequences(X_train, maxlen=MAX_LENGTH, padding='post')
test_sentences_X = pad_sequences(X_test, maxlen=MAX_LENGTH, padding='post')
# + [markdown] id="BKi2cvNKi-1Q" colab_type="text"
# Split Train into Train and Validation - about 10% into validation - In order to validate the model as it is training
# + colab_type="code" id="517zYSQLXkbn" colab={}
train_input = train_sentences_X[:140000]
val_input = train_sentences_X[140000:]
train_labels = y_train[:140000]
val_labels = y_train[140000:]
# + [markdown] colab_type="text" id="kHJbZDtk7N-3"
# # Model 1 -
#
# The first approach we'll try is to treat DA tagging as a standard multi-class text classification task, in the way you've done before with sentiment analysis and other tasks. Each utterance will be treated independently as a text to be classified with its DA tag label. This model has an architecture of:
#
# - Embedding
# - BLSTM
# - Fully Connected Layer
# - Softmax Activation
# + [markdown] colab_type="text" id="FItlHC1Fjz6y"
# The model architecture is as follows: Embedding Layer (to generate word embeddings) Next layer Bidirectional LSTM. Feed forward layer with number of neurons = number of tags. Softmax activation to get the probabilities.
#
# + colab_type="code" id="M97Sw5iv-lEU" colab={}
VOCAB_SIZE = len(wordvectors) # 43,731
MAX_LENGTH = len(max(sentences, key=len))
EMBED_SIZE = 100 # arbitary
HIDDEN_SIZE = len(unique_tags)
# + [markdown] id="LU9zEH6dba0G" colab_type="text"
# **The model below is created with one embedding layer, two biLSTM Layers and one dense layer and softmax activation layer. Adam optimiser is used with categorical_crossentropy loss for multiclass classification. This model treats the problem as a multiclass classification problem.**
#
#
# + colab_type="code" id="LCaX-ptaj8G2" outputId="6931fb78-f5e0-4ed8-b7e5-ed4bb7644b5b" colab={"base_uri": "https://localhost:8080/", "height": 317}
from keras.models import Sequential
from keras.layers import LSTM
from keras.layers import Dense
from keras.layers import Dropout, InputLayer, Bidirectional, TimeDistributed, Activation, Embedding
from keras.optimizers import Adam
#Building the network
# Include 2 BLSTM layers, in order to capture both the forward and backward hidden states
model = Sequential()
# Embedding layer with output size = EMBED_SIZE
model.add(Embedding(VOCAB_SIZE, EMBED_SIZE, input_length=MAX_LENGTH))
# Bidirectional 1 returns the full sequence
model.add(Bidirectional(LSTM(HIDDEN_SIZE, return_sequences=True)))
# Bidirectional 2
model.add(Bidirectional(LSTM(HIDDEN_SIZE)))
# Dense layer
model.add(Dense(HIDDEN_SIZE))
# Activation - softmax for multiclass classification
model.add(Activation('softmax'))
optimizer = Adam()
model.compile(loss='categorical_crossentropy',optimizer='adam',metrics=['accuracy'])
model.summary()
# + [markdown] id="Vu7wT539bebF" colab_type="text"
# **The layers of the model are given below**
# + id="LGTTgAa2IKut" colab_type="code" outputId="80fb7d6c-d713-4dee-cac2-d8da20ab890f" colab={"base_uri": "https://localhost:8080/", "height": 631}
from IPython.display import SVG
from keras.utils import vis_utils
SVG(vis_utils.model_to_dot(model, show_shapes=True, show_layer_names=True, dpi='70').create(prog='dot', format='svg'))
# + [markdown] id="YVMyGM5VcSs8" colab_type="text"
# **The model is trained until the val_accuracy starts to decrease. The callback of the fit function, called "EarlyStopping" is used. This stops the training as soon as the val_accuracy is lower for the next three epochs. The weights from the best epoch is restored.**
# + colab_type="code" id="OeiLkgD3Arpl" outputId="f981839f-0b8b-48ea-c3cd-7509ede3e691" colab={"base_uri": "https://localhost:8080/", "height": 487}
# Train the model - using validation
from keras.callbacks import EarlyStopping
history = model.fit(x=train_input, y=train_labels, epochs=50, batch_size=500, validation_data=(val_input, val_labels),
callbacks=[EarlyStopping(monitor='val_accuracy', patience=3, verbose=1, restore_best_weights=True)])
# + colab_type="code" id="2LkONUKQkSrL" outputId="deb41746-6eaa-47d0-d8d3-e7c0189e32ed" colab={"base_uri": "https://localhost:8080/", "height": 33}
score = model.evaluate(test_sentences_X, y_test, batch_size=100)
# + colab_type="code" id="Ab0ZL1dqkTY4" outputId="79ff535e-6f94-409c-f5d4-9ff09015b684" colab={"base_uri": "https://localhost:8080/", "height": 33}
print("Overall Accuracy:", score[1]*100)
# + [markdown] colab_type="text" id="LhMViQVSPY1J"
# ## Evaluation
#
#
# The overall accuracy is 67%, an effective accuracy for this task.
# + [markdown] colab_type="text" id="XHwoVCEwjEz7"
# In addition to overall accuracy, you need to look at the accuracy of some minority classes. Signal-non-understanding ('br') is a good indicator of "other-repair" or cases in which the other conversational participant attempts to repair the speaker's error. Summarize/reformulate ('bf') has been used in dialogue summarization. Report the accuracy for these classes and some frequent errors you notice the system makes in predicting them. What do you think the reasons are?
# + [markdown] colab_type="text" id="H7owA1f27se8"
# ## Minority Classes
# + colab_type="code" id="UZ8BwgDxNcIr" colab={}
# Generate predictions for the test data
predictions = model.predict(test_sentences_X, batch_size=100)
# + colab_type="code" id="5I26g20qQdzF" colab={}
# Build the confusion matrix off these predictions
matrix = sklearn.metrics.confusion_matrix(y_test.argmax(axis=1), predictions.argmax(axis=1))
# + [markdown] id="7iSCP9NFE22Z" colab_type="text"
# **Get the acuracy of the minority classes as given below.**
# + id="qwzKei_bgQoo" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 50} outputId="178ca6f3-7b7b-468f-c42f-f66d09573db4"
acc_class = matrix.diagonal()/matrix.sum(axis=1)
index_br = list(one_hot_encoding_dic["br"][one_hot_encoding_dic["br"]==1].index)[0]
br_accuracy = acc_class[index_br]*100
print("br accuracy: {}".format(br_accuracy))
index_bf = list(one_hot_encoding_dic["bf"][one_hot_encoding_dic["bf"]==1].index)[0]
bf_accuracy = acc_class[index_bf]*100
print("bf accuracy: {}".format(bf_accuracy))
# + [markdown] colab_type="text" id="HdnpWLggZ-6z"
#
# Due to the reduced lack of training data for the minority classes, these minority classifiers will not be very confident in classification, as they have not been fully optimised. The frequent classifiers will be more optimised and will generate more confident scores for all examples, effectively crowding out the less confident minority classifiers.
#
#
#
# + [markdown] colab_type="text" id="BZ16sE5F7x9e"
# # Model 2 - Balanced Network
# + [markdown] colab_type="text" id="hKHbOs4WkFaP"
#
# One thing we can do to try to improve performance is therefore to balance the data more sensibly. As the dataset is highly imbalanced, we can simply weight up the minority classes proportionally to their underrepresentation while training.
# + colab_type="code" id="6L4kNdf6kGEa" colab={}
import numpy as np
from sklearn.utils.class_weight import compute_class_weight
y_integers = np.argmax(tags_encoding, axis=1)
class_weights = compute_class_weight('balanced', np.unique(y_integers), y_integers)
d_class_weights = dict(enumerate(class_weights))
# + [markdown] colab_type="text" id="zF1UM-ZMZoa1"
# ## Define & Train the model
# + colab_type="code" id="xIRgRAzOPSAZ" outputId="e43860fe-a436-4220-faa4-feabe4e367a4" colab={"base_uri": "https://localhost:8080/", "height": 317}
# Re-built the model for the balanced training
# Include 2 BLSTM layers, in order to capture both the forward and backward hidden states
model_balanced = Sequential()
# Embedding layer
model_balanced.add(Embedding(VOCAB_SIZE, EMBED_SIZE, input_length=MAX_LENGTH))
# Bidirectional 1
model_balanced.add(Bidirectional(LSTM(HIDDEN_SIZE, return_sequences=True)))
# Bidirectional 2
model_balanced.add(Bidirectional(LSTM(HIDDEN_SIZE)))
# Dense layer
model_balanced.add(Dense(HIDDEN_SIZE))
# Activation
model_balanced.add(Activation('softmax'))
optimizer = Adam()
model_balanced.compile(loss='categorical_crossentropy',optimizer='adam',metrics=['accuracy'])
model_balanced.summary()
# + [markdown] id="oRmFk9y9FCRk" colab_type="text"
# **Adding class_weight=d_class_weights for a balanced model. This adds a higher penalty for the missclassification of minority classes. This weights the loss function during training. It is used to pay more attention to the minority classes.**
# + colab_type="code" id="xB2McUREkL4B" outputId="fb84c8ec-6e81-464b-df7b-b7023e173d67" colab={"base_uri": "https://localhost:8080/", "height": 420}
# Train the balanced network - takes time to achieve good accuracy
from keras.callbacks import EarlyStopping
history_balanced = model_balanced.fit(train_input, train_labels, epochs=50, batch_size=500, validation_data=(val_input, val_labels), class_weight=d_class_weights,
callbacks=[EarlyStopping(monitor='val_accuracy', patience=3, verbose=1, restore_best_weights=True)])
# + [markdown] colab_type="text" id="DJPjlMclZtw2"
# ## Test the model
# + colab_type="code" id="8UMAMGpJRINC" outputId="d84cff7c-e939-4549-beec-57bd1ebcd699" colab={"base_uri": "https://localhost:8080/", "height": 33}
# Overall Accuracy
score = model_balanced.evaluate(test_sentences_X, y_test, batch_size=100)
# + colab_type="code" id="0xzLIkTarjei" outputId="910dbf5c-6c7d-4444-91d7-a869b072e6f3" colab={"base_uri": "https://localhost:8080/", "height": 33}
print("Overall Accuracy:", score[1]*100)
# + colab_type="code" id="qkULcz2igEW3" colab={}
# Generate predictions for the test data
label_pred = model_balanced.predict(test_sentences_X, batch_size=100)
# + [markdown] colab_type="text" id="hq7i7giWZ4_l"
# ## Balanced network evaluation
# + [markdown] colab_type="text" id="fM7VWweco0Et"
# Report the overall accuracy and the accuracy of 'br' and 'bf' classes. Suggest other ways to handle imbalanced classes.
# + colab_type="code" id="4jNfWmSNgRvT" colab={"base_uri": "https://localhost:8080/", "height": 50} outputId="5952909a-d54f-42e1-ad65-ba6e0df006b1"
# Build the confusion matrix off these predictions
matrix_balanced = sklearn.metrics.confusion_matrix(y_test.argmax(axis=1), label_pred.argmax(axis=1))
acc_class_balanced = matrix_balanced.diagonal()/matrix_balanced.sum(axis=1)
index_br = list(one_hot_encoding_dic["br"][one_hot_encoding_dic["br"]==1].index)[0]
br_accuracy = acc_class_balanced[index_br]*100
print("br accuracy: {}".format(br_accuracy))
index_bf = list(one_hot_encoding_dic["bf"][one_hot_encoding_dic["bf"]==1].index)[0]
bf_accuracy = acc_class_balanced[index_bf]*100
print("bf accuracy: {}".format(bf_accuracy))
# Calculate Accuracies for "br" and "bf"
# + [markdown] id="HYC8XaZdq8EV" colab_type="text"
# **The overall accuracy of the balanced model is lower than the accuracy of the unbalanced model. But the individual classes have a much better accuracy in balanced model than in unbalanced model. In balanced model, there is a higher penalty when the classification of minority classes goes wrong. Hence the overall accuracy is lower.**
# + [markdown] colab_type="text" id="Zi9GyVUvPcrF"
#
#
# ### Accuracies
#
#
#
# ### Explanation
#
#
# ### Other ways to handle imbalanced classes
#
#
# - Over-sampling: this means to increase instances of minority classes on the training set by duplication. The advantage is that no information is lost. The disadvantage is that the model becomes prone to overfitting.
#
# - Under-sampling: This means to decrease the instances of majority classes until it is comparable with the minority class. As this method removes the data from dataset, useful information may be lost.
# + [markdown] colab_type="text" id="fW4g5mQkkaFv"
# Can we improve things by using context information? Next we try to build a model which predicts DA tag from the sequence of
# previous DA tags, plus the utterance representation.
# + [markdown] colab_type="text" id="WfrGWuZ6nk4y"
# # Using Context for Dialog Act Classification
#
# The second approach we will try is a hierarchical approach to DA tagging. We expect there is valuable sequential information among the DA tags. So in this section we apply a BiLSTM on top of the sentence CNN representation. The CNN model learns textual information in each utterance for DA classification, acting like the text classifier from Model 1 above. Then we use a bidirectional-LSTM (BLSTM) above that to learn how to use the context before and after the current utterance to improve the output.
# + [markdown] colab_type="text" id="7qyPpNaK-2mb"
# ## Define the model
#
# This model has an architecture of:
#
# - Word Embedding
# - CNN
# - Bidirectional LSTM
# - Fully-Connected output
#
#
# + [markdown] colab_type="text" id="DuJLqgjWqcIf"
# ## CNN
#
#
# This is a classical CNN layer used to convolve over embedings tensor and gether useful information from it. The data is represented by hierarchy of features, which can be modelled using a CNN. We transform/reshape conv output to 2d matrix. Then we pass it to the max pooling layer that applies the max pool operation on windows of different sizes.
# + colab_type="code" id="XA5INtFl-fM0" colab={}
from keras.layers import Input, Reshape, Conv2D, BatchNormalization, MaxPool2D, concatenate, Flatten
from keras.models import Model
filter_sizes = [3,4,5]
num_filters = 64
drop = 0.2
VOCAB_SIZE = len(wordvectors) # 43,731
MAX_LENGTH = len(max(sentences, key=len))
EMBED_SIZE = 100 # arbitary
HIDDEN_SIZE = len(unique_tags)
# CNN model
inputs = Input(shape=(MAX_LENGTH, ), dtype='int32')
embedding = Embedding(input_dim=VOCAB_SIZE, output_dim=EMBED_SIZE, input_length=MAX_LENGTH)(inputs)
reshape = Reshape((MAX_LENGTH, EMBED_SIZE, 1))(embedding)
# 3 convolutions
conv_0 = Conv2D(num_filters, kernel_size=(filter_sizes[0], EMBED_SIZE), strides=1, padding='valid', kernel_initializer='normal', activation='relu')(reshape)
bn_0 = BatchNormalization()(conv_0)
conv_1 = Conv2D(num_filters, kernel_size=(filter_sizes[1], EMBED_SIZE), strides=1, padding='valid', kernel_initializer='normal', activation='relu')(reshape)
bn_1 = BatchNormalization()(conv_1)
conv_2 = Conv2D(num_filters, kernel_size=(filter_sizes[2], EMBED_SIZE), strides=1, padding='valid', kernel_initializer='normal', activation='relu')(reshape)
bn_2 = BatchNormalization()(conv_2)
# maxpool for 3 layers
maxpool_0 = MaxPool2D(pool_size=(MAX_LENGTH - filter_sizes[0] + 1, 1), padding='valid')(bn_0)
maxpool_1 = MaxPool2D(pool_size=(MAX_LENGTH - filter_sizes[1] + 1, 1), padding='valid')(bn_1)
maxpool_2 = MaxPool2D(pool_size=(MAX_LENGTH - filter_sizes[2] + 1, 1), padding='valid')(bn_2)
# concatenate tensors
merged_1 = concatenate([maxpool_0, maxpool_1, maxpool_2])
# flatten concatenated tensors
# applying time distributed layer so that cnn output is compatible with BiLSTM input
flat = TimeDistributed(Flatten())(merged_1)
# dense layer (dense_1)
dense_1 = Dense(HIDDEN_SIZE, activation='relu')(flat)
# dropout_1
dropout_1 = Dropout(drop)(dense_1)
# + [markdown] colab_type="text" id="wDuERMw7-rAV"
# ## BLSTM
#
# This is used to create LSTM layers. The data we’re working with has temporal properties which we want to model as well — hence the use of a LSTM. You should create a BiLSTM.
# + colab_type="code" id="pFGp2EWI-fM7" colab={}
# BLSTM model
# using CNN + dense + dropout's output as input to the BiLSTM layer.
biLSTM1 = Bidirectional(LSTM(HIDDEN_SIZE, return_sequences='true'))(dropout_1)
# Bidirectional 2
biLSTM2 = Bidirectional(LSTM(HIDDEN_SIZE))(biLSTM1)
# Dense layer (dense_2)
dense_2 = Dense(HIDDEN_SIZE, activation='relu')(biLSTM2)
# dropout_2
dropout_2 = Dropout(drop)(dense_2)
# + [markdown] colab_type="text" id="7wluAkx6AQUb"
# Concatenate 2 last layers and create the output layer
# + colab_type="code" id="kzrhgkX2-fNE" outputId="092afa28-4079-4e83-c442-f9295f889525" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# concatenate 2 final layers
# flatten the output of the CNN + dense + dropout so that it can be concatenated with the output of BiLSTM
dropout_flat = Flatten()(dropout_1)
# concatenating the output of CNN + dense + dropout with the output of BiLSTM + dense + dropout
merged_2 = concatenate([dropout_flat, dropout_2])
# merged_2 has the dimension of (None, 86)
# adding a dense layer to get the output of the dim (None, 43) for the multiclass classification problem
dense_3 = Dense(units=HIDDEN_SIZE, input_shape=(1,))(merged_2)
# adding softmax for multiclass classification
output = Activation('softmax')(dense_3)
optimizer = Adam()
model_cnn = Model(inputs=[inputs], outputs=[output])
model_cnn.compile(loss='categorical_crossentropy',optimizer='adam',metrics=['accuracy'])
model_cnn.summary()
# + id="JfNrZtnCHxSU" colab_type="code" outputId="883824c0-6ab3-4a05-c9cd-1f4bfb8b69c7" colab={"base_uri": "https://localhost:8080/", "height": 1000}
from IPython.display import SVG
from keras.utils import vis_utils
SVG(vis_utils.model_to_dot(model_cnn, show_shapes=True, show_layer_names=True, dpi='50').create(prog='dot', format='svg'))
# + colab_type="code" id="3Jneg-GD-fNJ" outputId="3a101608-4b60-48b9-ae08-7435101d0b01" colab={"base_uri": "https://localhost:8080/", "height": 320}
# Train the model - using validation
from keras.callbacks import EarlyStopping
history = model_cnn.fit(train_input, train_labels, epochs=50, batch_size=500, validation_data=(val_input, val_labels),
callbacks=[EarlyStopping(monitor='val_accuracy', patience=3, verbose=1, restore_best_weights=True)])
# + colab_type="code" id="MSMRSX1u-fNO" outputId="5f2c64ca-6d07-4656-e8bc-9e8f6ba19471" colab={"base_uri": "https://localhost:8080/", "height": 33}
score = model_cnn.evaluate(test_sentences_X, y_test, batch_size=100)
# + colab_type="code" id="3qFMsXNS-fNS" outputId="66e27b1d-218c-4653-8a70-5c82411732e8" colab={"base_uri": "https://localhost:8080/", "height": 33}
print("Overall Accuracy:", score[1]*100)
# + [markdown] colab_type="text" id="QmO6hVsWTaNr"
# ### Minority Classes
#
#
# + id="wTjOQhWptIGb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 50} outputId="4a586f73-39aa-405a-8d3e-b1635c9934c5"
predictions_cnn = model_cnn.predict(test_sentences_X, batch_size=100)
matrix_cnn = sklearn.metrics.confusion_matrix(y_test.argmax(axis=1), predictions_cnn.argmax(axis=1))
acc_class_cnn = matrix_cnn.diagonal()/matrix_cnn.sum(axis=1)
index_br = list(one_hot_encoding_dic["br"][one_hot_encoding_dic["br"]==1].index)[0]
br_accuracy = acc_class_cnn[index_br]*100
print("br accuracy: {}".format(br_accuracy))
index_bf = list(one_hot_encoding_dic["bf"][one_hot_encoding_dic["bf"]==1].index)[0]
bf_accuracy = acc_class_cnn[index_bf]*100
print("bf accuracy: {}".format(bf_accuracy))
# + [markdown] id="Nj5TKy_Etu6H" colab_type="text"
# # CNN balanced
# + [markdown] id="tI0_jM926hFM" colab_type="text"
# Balanced CNN+BiLSTM model - architecture remains the same
# + id="5BOoRUPQtw5X" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="e4dbc741-64c7-48f2-b54b-9e08f008e8cb"
from keras.layers import Input, Reshape, Conv2D, BatchNormalization, MaxPool2D, concatenate, Flatten
from keras.models import Model
filter_sizes = [3,4,5]
num_filters = 64
drop = 0.2
VOCAB_SIZE = len(wordvectors) # 43,731
MAX_LENGTH = len(max(sentences, key=len))
EMBED_SIZE = 100 # arbitary
HIDDEN_SIZE = len(unique_tags)
# CNN model
inputs = Input(shape=(MAX_LENGTH, ), dtype='int32')
embedding = Embedding(input_dim=VOCAB_SIZE, output_dim=EMBED_SIZE, input_length=MAX_LENGTH)(inputs)
reshape = Reshape((MAX_LENGTH, EMBED_SIZE, 1))(embedding)
# 3 convolutions
conv_0 = Conv2D(num_filters, kernel_size=(filter_sizes[0], EMBED_SIZE), strides=1, padding='valid', kernel_initializer='normal', activation='relu')(reshape)
bn_0 = BatchNormalization()(conv_0)
conv_1 = Conv2D(num_filters, kernel_size=(filter_sizes[1], EMBED_SIZE), strides=1, padding='valid', kernel_initializer='normal', activation='relu')(reshape)
bn_1 = BatchNormalization()(conv_1)
conv_2 = Conv2D(num_filters, kernel_size=(filter_sizes[2], EMBED_SIZE), strides=1, padding='valid', kernel_initializer='normal', activation='relu')(reshape)
bn_2 = BatchNormalization()(conv_2)
# maxpool for 3 layers
maxpool_0 = MaxPool2D(pool_size=(MAX_LENGTH - filter_sizes[0] + 1, 1), padding='valid')(bn_0)
maxpool_1 = MaxPool2D(pool_size=(MAX_LENGTH - filter_sizes[1] + 1, 1), padding='valid')(bn_1)
maxpool_2 = MaxPool2D(pool_size=(MAX_LENGTH - filter_sizes[2] + 1, 1), padding='valid')(bn_2)
# concatenate tensors
merged_1 = concatenate([maxpool_0, maxpool_1, maxpool_2])
# flatten concatenated tensors
# applying time distributed layer so that cnn output is compatible with BiLSTM input
flat = TimeDistributed(Flatten())(merged_1)
# dense layer (dense_1)
dense_1 = Dense(HIDDEN_SIZE, activation='relu')(flat)
# dropout_1
dropout_1 = Dropout(drop)(dense_1)
# BLSTM model
# using CNN + dense + dropout's output as input to the BiLSTM layer.
biLSTM1 = Bidirectional(LSTM(HIDDEN_SIZE, return_sequences='true'))(dropout_1)
# Bidirectional 2
biLSTM2 = Bidirectional(LSTM(HIDDEN_SIZE))(biLSTM1)
# Dense layer (dense_2)
dense_2 = Dense(HIDDEN_SIZE, activation='relu')(biLSTM2)
# dropout_2
dropout_2 = Dropout(drop)(dense_2)
# concatenate 2 final layers
# flatten the output of the CNN + dense + dropout so that it can be concatenated with the output of BiLSTM
dropout_flat = Flatten()(dropout_1)
# concatenating the output of CNN + dense + dropout with the output of BiLSTM + dense + dropout
merged_2 = concatenate([dropout_flat, dropout_2])
# merged_2 has the dimension of (None, 86)
# adding a dense layer to get the output of the dim (None, 43) for the multiclass classification problem
dense_3 = Dense(units=HIDDEN_SIZE, input_shape=(1,))(merged_2)
# adding softmax for multiclass classification
output = Activation('softmax')(dense_3)
optimizer = Adam()
model_cnn_balanced = Model(inputs=[inputs], outputs=[output])
model_cnn_balanced.compile(loss='categorical_crossentropy',optimizer='adam',metrics=['accuracy'])
model_cnn_balanced.summary()
# + id="vCSALauPuJ0O" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 553} outputId="2dae30e8-9ff7-4295-b466-042edeb9a7fd"
from keras.callbacks import EarlyStopping
history = model_cnn_balanced.fit(train_input, train_labels, epochs=50, batch_size=500, validation_data=(val_input, val_labels), class_weight=d_class_weights,
callbacks=[EarlyStopping(monitor='val_accuracy', patience=3, verbose=1, restore_best_weights=True)])
# + id="0QvPZmUpu8h9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 33} outputId="0c5cbefe-45b3-42bb-9de6-ec53864bd52a"
score = model_cnn_balanced.evaluate(test_sentences_X, y_test, batch_size=100)
# + id="DM9ZUKfwvB0O" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 33} outputId="67008e02-c202-46e8-ea78-38d04f3b4bc2"
print("Overall Accuracy:", score[1]*100)
# + [markdown] id="YLDWjELU6sgh" colab_type="text"
# **Getting the accuracy for "br" and "bf" class**
# + id="0NKfAe2xvGo-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 50} outputId="77164526-57c0-4089-d844-949ef9cc9b2f"
predictions_cnn_bal = model_cnn_balanced.predict(test_sentences_X, batch_size=100)
matrix_cnn_bal = sklearn.metrics.confusion_matrix(y_test.argmax(axis=1), predictions_cnn_bal.argmax(axis=1))
acc_class_cnn_bal = matrix_cnn_bal.diagonal()/matrix_cnn_bal.sum(axis=1)
index_br = list(one_hot_encoding_dic["br"][one_hot_encoding_dic["br"]==1].index)[0]
br_accuracy = acc_class_cnn_bal[index_br]*100
print("br accuracy: {}".format(br_accuracy))
index_bf = list(one_hot_encoding_dic["bf"][one_hot_encoding_dic["bf"]==1].index)[0]
bf_accuracy = acc_class_cnn_bal[index_bf]*100
print("bf accuracy: {}".format(bf_accuracy))
# + [markdown] id="ZJ5wxnqv7u7s" colab_type="text"
# Report your overall accuracy. Did context help disambiguate and better predict the minority classes ('br' and 'bf')? What are frequent errors? Show one positive example where adding context changed the prediction.
# + [markdown] id="0IHOqL0V61vT" colab_type="text"
# **The accuracy for br has increased from 46% initially (BiLSTM - Balanced) to now 50%. The accuracy pf bf has reduced from 14% initially (BiLSTM - Balanced) to 9% now.**
# + [markdown] id="ggmKyE_8v2Fp" colab_type="text"
# # Prediction Examples
# + [markdown] id="a_EVm9n34J2y" colab_type="text"
# **To get positive change from the BiLSTM model to CNN+BiLSTM model. If the model initially predicted the wrong class (BiLSTM) but it later got rectified by CNN+BiLSTM then it is noted as positive change. The code below gets the indexof all positive changes as well as negative changes.**
# + id="GHcwvINNzak9" colab_type="code" colab={}
index_pos_change = []
index_neg_change = []
for i in range(len(y_test)):
true_label = y_test[i].argmax(axis=0)
bLISTM_pred = label_pred[i].argmax(axis=0)
cnn_pred = predictions_cnn_bal[i].argmax(axis=0)
if true_label == bLISTM_pred and true_label != cnn_pred:
index_neg_change.append(i)
elif true_label != bLISTM_pred and true_label == cnn_pred:
index_pos_change.append(i)
# + [markdown] id="iTXK7P-C4kvq" colab_type="text"
# **Printing the values of the sentences which changed from BiLSTM model to CNN+BiLSTM. It can be seen that the sequences are quite short. CNN+LSTM model helps in classifying the short sentences correctly.**
# + id="a19XiffhBEZ0" colab_type="code" colab={}
# creating a reverse mapping from word_ids to words
reverse_word_index = dict([(value, key) for (key, value) in wordvectors.items()])
# method to decode the sentence from a list of IDs to a string
def decode_sentence(text):
return ' '.join([reverse_word_index.get(i, '?') for i in text])
# + id="JqFCmXkf1Fmh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="5fc5121a-ffaf-4dbc-c89c-9e38aa20b467"
# printing all the sentences which were missclassified by BiLSTM but correctly classified by CNN + BiLSTM
for i, val in enumerate(index_pos_change):
print(decode_sentence(X_test[val]))
# + [markdown] id="07JxrFgpBgKx" colab_type="text"
# **One of the frequent errors the BiLSTM model made was of the missclassification of smaller sentences which doesn't have any context. For example "How funny. /", "yeah. /", "# Right. # /", "How true. /" etc.**
# + [markdown] colab_type="text" id="gUZt48JgrE34"
# # Advanced: Bert-Based Model for Dialogue Act Tagging
# + [markdown] colab_type="text" id="zE63Q5guuPdA"
# In the last section we want to use BERT and leverage contextual word embeddings, following on from the last lab you've
# just done. This is an advanced part of the assignment and worth 10 marks (20%) in total. You could use your BERT-based text classifier here (instead of the CNN utterance-level classifier) and see if a pre-trained BERT language model helps. The domain difference from conversational data is one possible downside to using BERT. Explore some techniques to efficiently transfer the knowledge from conversational data and to improve model performance on DA tagging.
# + [markdown] id="AkU3ofnHR8qR" colab_type="text"
# **Refer to the notebook “PART_B_Task_3.ipynb”**
| coursework-2/PART_B_Task _1_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import numpy as np
import cv2
import matplotlib.pyplot as plt
from PIL import Image
from scipy import ndimage
from skimage.morphology import erosion, dilation, opening, closing, white_tophat
from skimage.morphology import disk
# %matplotlib inline
# +
def adaptive_thresh(input_img):
h, w = input_img.shape
S = int(w/8)
s2 = int(S/2)
T = 10.0
#integral img
int_img = np.zeros_like(input_img, dtype=np.uint32)
for col in range(w):
for row in range(h):
int_img[row,col] = input_img[0:row,0:col].sum()
#output img
out_img = np.zeros_like(input_img)
for col in range(w):
for row in range(h):
#SxS region
y0 = max(row-s2, 0)
y1 = min(row+s2, h-1)
x0 = max(col-s2, 0)
x1 = min(col+s2, w-1)
count = (y1-y0)*(x1-x0)
#print(x0,y0,x1,y1)
sum_ = int_img[y1, x1]-int_img[y0, x1]-int_img[y1, x0]+int_img[y0, x0]
if input_img[row, col]*count < sum_*(100.-T)/100.:
out_img[row,col] = 0
else:
out_img[row,col] = 255
return out_img
def faster_bradley_threshold(image, threshold=90, window_r=45):
percentage = threshold / 100.
window_diam = 2*window_r + 1
# convert image to numpy array of grayscale values
img = np.array(image).astype(np.float) # float for mean precision
# matrix of local means with scipy
means = ndimage.uniform_filter(img, window_diam)
# result: 0 for entry less than percentage*mean, 255 otherwise
height, width = img.shape[:2]
result = np.zeros((height,width), np.uint8) # initially all 0
result[img >= percentage * means] = 255
return np.array(result)
# +
def get_shading(rim,mask):
im=rim
im=cv2.copyMakeBorder(rim, top=100, bottom=100, left=100, right=100, borderType= cv2.BORDER_CONSTANT, value=255)
mask=cv2.copyMakeBorder(mask, top=100, bottom=100, left=100, right=100, borderType= cv2.BORDER_CONSTANT, value=255)
mask=mask/255
shading=np.zeros(im.shape)
for i in range(len(im)):
#print(i)
for j in range(len(im[i])):
if mask[i][j]>0:
shading[i][j]=im[i][j]
else:
dx=3
dy=3
while np.count_nonzero(mask[i-dx:i+dx,j-dy:j+dy])<25:
dx+=2
dy+=1
curr=0.0
tot=0.0
for it1 in range(max(i-dx,0),min(i+dx,len(mask))):
for it2 in range(max(j-dy,0),min(j+dy,len(mask[0]))):
if mask[it1][it2]>0:
curr+=im[it1][it2]
tot+=1
shading[i][j]=curr/tot
return shading[100:-100,100:-100]
def get_reflectance(im,shading):
reflectance=np.zeros(im.shape)
for i in range(len(im)):
for j in range(len(im[i])):
reflectance[i][j]=((im[i][j]*1.0)/shading[i][j])
#reflectance=reflectance[30:-30,30:-30]
return reflectance
# +
rim=cv2.imread('reflectance.png',0)
mask=cv2.imread('0mask.png',0)
c=0
for i in range(len(mask)):
for j in range(len(mask[i])):
if mask[i][j]>0:
mask[i][j]=255
else:
mask[i][j]=0
c+=1
print(c)
# -
shading=get_shading(rim,mask)
nshading = (255*(shading-np.min(shading)))/(np.max(shading)-np.min(shading))
ref=get_reflectance(rim,shading)
plt.imshow(ref,cmap='gray')
cv2.imwrite('1.png',ref)
rim=cv2.imread('./ACCV-Dataset/RealWorld/032.png',0)
rim=np.array(rim,dtype='float32')
circle = disk(5)
#rim=cv2.resize(rim,None,fx=0.2,fy=0.2)
for i in range(10):
print("Iteration",i)
#mask=cv2.adaptiveThreshold(rim, 255,cv2.ADAPTIVE_THRESH_GAUSSIAN_C, cv2.THRESH_BINARY,41,20)
mask=adaptive_thresh(rim)
#mask=cv2.dilate(mask,circle,iterations = 1)
print("Thresholding done")
n3=str(i)+'mask.png'
cv2.imwrite(n3,mask)
shading=get_shading(rim,mask)
print("shading done")
#plt.imshow(shading,cmap='gray')
rim=get_reflectance(rim,shading)
ms=np.max(rim)
mi=np.min(rim)
rim=255*(rim-np.min(rim))
rim=rim/(ms-mi)
n1=str(i)+'shading.png'
n2=str(i)+'reflectance.png'
cv2.imwrite(n1,shading)
cv2.imwrite(n2,rim)
# +
#rim=cv2.imread('reflectance.png',0)
plt.imshow(mask,cmap='gray')
nmask=cv2.dilate(mask,circle,iterations = 5)
| src/Document/ShadowRemoval.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import base64
import os
import requests
import numpy as np
import io
# This notebook shows some examples of how to call a model trained and deployed in the Peltarion platform from Python using JSON requests.
#
# The advantage of using JSON is that you can score several examples in a batch rather than one at a time as with curl or POST requests with forms.
# ## Image classification
#
# ### Classify a single image
#
# The following example assumes that you have a model trained on [MNIST](http://docs.peltarion.com/new/en/datasets-view/datasets-used-in-tutorials/mnist-dataset.html) data (28x28 pixels, 3 color channels) and want to classify new images that the model has not seen. Pretty much the same thing what you do in the tutorial [Deploy an operational AI model](http://docs.peltarion.com/new/en/tutorials/tutorial---deploy-an-operational-ai-model.html).
#
# We will hide the deployment URL and the authentication token for security reasons. You can substitute the values found on the Deployment view for the model we want to use.
url = ' -- insert the URL you find on the deployment view -- '
token = ' -- insert the token you find on the deployment view --'
# The file called `three.png` can be found in the images folder of this repo.
#
# In order to feed the image to the deployment API, we need to encode it in base64 format and prepend a short string explaining what type of data it is.
img_file = "images/three.png"
img_type = os.path.splitext(img_file)[-1][1:]
with open(img_file, "rb") as image_file:
encoded_img = 'data:image/{};base64,'.format(img_type) + base64.b64encode(image_file.read()).decode('ascii')
# The structure of the JSON string that we will send is shown below. The "rows" key must always be present and its value is a list of strings that represent different examples that we want to classify. Each entry of the list contains a comma-separated set of key-value pairs where the key is the feature name and the value is the feature value (which will be a base64 string for images.)
#
# ```
# {"rows":
# [{"feature1": "value1", "feature2": "value2"},
# {"feature1": "value1", "feature2": "value2"}
# ]
# }
# ```
# For the current example, we only have one example and one feature called "Image". (You can find the feature names on the Deployment view.) The structure of the JSON will be fairly simple:
#
# ```
# {"rows":
# [{"Image": "<base64 encoded image>"}]
# }
# ```
# +
payload = "{\"rows\": [{\"Image\":\"" + encoded_img + "\"}]}"
headers = {
'Content-Type': "application/json",
'Authorization': "Bearer {}".format(token),
}
response = requests.request("POST", url, data=payload, headers=headers)
print(response.json())
# -
# **Result**: ‘3’ gets the highest value, 0.99999976. This means that the model predicts the image to be a ‘3’.
# ### Classify several images
# To simplify our life, we might want to write a small function that encodes an image to base64 given a file path.
def encode_img(path):
img_type = os.path.splitext(path)[-1][1:]
with open(path, "rb") as image_file:
encoded_img = 'data:image/{};base64,'.format(img_type) + base64.b64encode(image_file.read()).decode('ascii')
return encoded_img
# Now we can classify a batch of images, in this case just two, but it would work with a much larger batch too. The files can be found in the images folder of this repo.
img_files = ['images/three.png', 'images/Six.png']
encoded_imgs = [encode_img(f) for f in img_files]
input_batch = ','.join(["{\"Image\":\"" + encoded_img + "\"}" for encoded_img in encoded_imgs])
payload = "{\"rows\": [" + input_batch + "]}"
response = requests.request("POST", url, data=payload, headers=headers)
response.json()
# **Result**: The first image is predicted to be a '3' and the second to a '6'.
# ## Tabular data
#
# This example assumes that we have trained a model on the California housing dataset, where we try to predict latitude from some numeric features.
# In this example we try to predict on which latitude a house is situated. We assume that the deployed model has been trained on the [Calihouse dataset](http://docs.peltarion.com/new/en/datasets-view/datasets-used-in-tutorials/calihouse-dataset.html) as in the tutorial [Predict California house prices](http://docs.peltarion.com/new/en/tutorials/tutorial---predict-california-house-prices.html).
url = ' -- insert the URL you find on the deployment view -- '
token = ' -- insert the token you find on the deployment view --'
# We can define a short utility function to construct a row for a training example in the right format.
def input_row(input_params):
return '{' + ','.join(["\"" + name + "\":" + value for (name, value) in input_params.items()]) + '}'
# +
ex1 = {
"population": "1551.0",
"totalBedrooms": "434.0",
"totalRooms": "2202.0",
"housingMedianAge": "52.0",
"medianHouseValue": "261100.0",
"medianIncome": "3.12",
"households": "514.0"
}
ex2 = {
"population": "3551.0",
"totalBedrooms": "834.0",
"totalRooms": "2902.0",
"housingMedianAge": "76.0",
"medianHouseValue": "111100.0",
"medianIncome": "2.12",
"households": "1000.0"
}
examples = [ex1, ex2]
input_batch = ','.join([input_row(ex) for ex in examples])
payload = "{\"rows\": [" + input_batch + "]}"
# -
payload
# +
headers = {
'Content-Type': "application/json",
'Authorization': "Bearer {}".format(token),
}
response = requests.request("POST", url, data=payload, headers=headers)
print(response.json())
# -
# **Result**: The model predicts that the second house is situated slightly north of the first house.
# ## Images and tabular data
#
# In this example, we will predict the mean house value in a specific area, just as in the tutorial [Predict California house prices](http://docs.peltarion.com/new/en/tutorials/tutorial---predict-california-house-prices.html). We use a model trained on the [Calihouse dataset](http://docs.peltarion.com/new/en/datasets-view/datasets-used-in-tutorials/calihouse-dataset.html) that consists of map images from [Open street map](https://www.openstreetmap.org/about) and tabular demographic data collected from the California 1990 Census.
url = ' -- insert the URL you find on the Deployment view -- '
token = ' -- insert the token you find on the Deployment view --'
# We will re-use the `encode_imgs()` function defined above here.
img_files = ['images/15_5256_12656.png', 'images/15_5258_12653.png']
encoded_imgs = [encode_img(f) for f in img_files]
# We can now populate the examples with numerical values and encoded images.
# +
ex1 = {
"population": "1551.0",
"totalBedrooms": "434.0",
"totalRooms": "2202.0",
"housingMedianAge": "52.0",
"medianIncome": "3.12",
"households": "514.0",
"image_path": "\"" + encoded_imgs[0] + "\"",
"latitude": "37.88",
"longitude": "-122.25"
}
ex2 = {
"population": "3551.0",
"totalBedrooms": "834.0",
"totalRooms": "2902.0",
"housingMedianAge": "76.0",
"medianIncome": "2.12",
"households": "1000.0",
"image_path": "\"" + encoded_imgs[1] + "\"",
"latitude": "37.88",
"longitude": "-122.25"
}
examples = [ex1,ex2]
input_batch = ','.join([input_row(ex) for ex in examples])
payload = "{\"rows\": [" + input_batch + "]}"
headers = {
'Content-Type': "application/json",
'Authorization': "Bearer {}".format(token),
}
response = requests.request("POST", url, data=payload, headers=headers)
print(response.json())
# -
# **Result**: The model predicts that the area where the second house is situated is more expensive than the first houses' area.
# ## Image to image
#
# Here we will send two images to a deployment and get two images back. This is useful for image-to-image mapping problems, such as image segmentation or image denoising/reconstruction.
# The images in this example come from the [NoisyOffice dataset](https://archive.ics.uci.edu/ml/datasets/NoisyOffice) where the task is to clean images from stains and other imperfections.
url = ' -- insert the URL you find on the Deployment view -- '
token = ' -- insert the token you find on the Deployment view --'
# +
img_files = ['images/FontLrm_Noisec_TE.png', 'images/FontLrm_Noisew_TE.png']
encoded_imgs = [encode_img(f) for f in img_files]
input_batch = ','.join(["{\"path_noisy\":\"" + encoded_img + "\"}" for encoded_img in encoded_imgs])
payload = "{\"rows\": [" + input_batch + "]}"
headers = {
'Content-Type': "application/json",
'Authorization': "Bearer {}".format(token),
}
response = requests.request("POST", url, data=payload, headers=headers)
# -
results = response.json()['rows']
# Now you can, for example, save the generated images to file.
for i, res in enumerate(results):
decoded = base64.b64decode(res['path_clean'].split(',')[-1])
with open('images/image{}.png'.format(i), 'bw') as outf:
outf.write(decoded)
# ## Numpy to numpy
#
# Here we will send input data represented as numpy arrays to the deployment API, and get a numpy array of predictions back.
#
# The numpy data type can be used to build several models, e.g. auto-encoders, segmentation models, or multi-label classification of vectors or images.
# +
# Get predictions from deployment api
# Return the response as json
def get_predictions(data, token, url):
headers = {
'Content-Type': "application/json",
'Authorization': "Bearer {}".format(token),
}
response = requests.request("POST", url, data=data, headers=headers)
return response.json()
# Prepare a json data structure from numpy array
# Assume first axis in the numpy array arr represents samples
def prepare_api_data(arr, input_param_name="input"):
encoded_arrs = [encode_numpy(a) for a in arr]
input_batch = ','.join(["{\"" + input_param_name + "\":\"" + encoded_arr + "\"}" for encoded_arr in encoded_arrs])
payload = "{\"rows\": [" + input_batch + "]}"
return payload
# Encode a numpy array in base64 format and add data application type
def encode_numpy(arr):
# Need to temp save the arr to a buffer to get the npy headers not just the raw data
buffer = io.BytesIO()
np.save(buffer, arr)
encoded_arr = base64.b64encode(buffer.getvalue()).decode('ascii')
return 'data:application/x.peltarion.npy;base64,' + encoded_arr
# Decode a base64 string into a numpy array
def decode_base64(base64_string):
decoded = base64.decodebytes(base64_string.encode('ascii'))
buffer = io.BytesIO(decoded)
return np.load(buffer)
# Decode a json response from Peltarion deployment API into a numpy array
# The resulting array represents one or several samples
def decode_api_response(response_json, output_param_name='output'):
res = []
for sample in response_json['rows']:
data_base64 = sample[output_param_name].split(',')[1]
data_numpy = decode_base64(data_base64)
res.append(data_numpy)
return np.array(res)
# -
url = ' -- insert the URL you find on the Deployment view -- '
token = ' -- insert the token you find on the Deployment view --'
features= np.load('/home/asa/projects/potkaista_dev/features.npy')
print("Shape of the input numpy array:", features.shape)
api_data = prepare_api_data(features, input_param_name="features.npy_0")
preds = get_predictions(api_data, token, url)
decoded = decode_api_response(preds, output_param_name="labels.npy_0")
print("Shape of the returned numpy array: ", decoded.shape)
| Peltarion_deployment_JSON_examples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/uetsuji-kaito/ML-learn/blob/master/basic/2nd.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="AcjoSI-gemVt" colab_type="code" colab={}
# !pip install tensorflow==2.1.0
# + id="Zg2SZT6xk6y6" colab_type="code" colab={}
from tensorflow import keras
import numpy as np
import matplotlib.pyplot as plt
# + id="c7Va4HMXfFCJ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="0939ea98-6680-4d80-e5bc-4336fca7b552"
(x_train, y_train), (x_test, y_test) = keras.datasets.mnist.load_data()
# + id="quxUTT3ejAUQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 542} outputId="ceaeae1a-57c3-487a-9bf4-93dd0bfd0404"
plt.figure(figsize=(10,10))
for cl in range(10):
for idx in range(5):
plt.subplot(5, 10, 10*idx + cl + 1)
plt.xticks([])
plt.yticks([])
plt.grid(False)
plt.imshow(x_train[y_train==cl][idx], cmap=plt.cm.binary)
plt.xlabel(cl)
plt.show()
# + id="F0q3PSYRgRGa" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="7028139f-44ec-4f58-d215-64ec26980b67"
print("x_train shape:", x_train.shape)
print("y_train shape:", y_train.shape)
print("x_test shape:", x_test.shape)
print("y_test shape:", y_test.shape)
# + id="McDn2N8mm0ko" colab_type="code" colab={}
print(x_train[0])
# + id="XGFfJmeBGQIn" colab_type="code" colab={}
x_train = x_train / 255.
x_test = x_test / 255.
# + id="QCzs1Xh2GdQj" colab_type="code" colab={}
y_train = keras.utils.to_categorical(y_train)
y_test = keras.utils.to_categorical(y_test)
# + id="zt22vEFzEids" colab_type="code" colab={}
model = keras.Sequential()
model.add(keras.layers.Flatten(input_shape=(28, 28)))
model.add(keras.layers.Dense(units=128, activation=keras.activations.relu))
model.add(keras.layers.Dense(units=10, activation=keras.activations.softmax))
model.compile(
optimizer=keras.optimizers.SGD(),
loss=keras.losses.categorical_crossentropy,
metrics=["accuracy"]
)
# + id="tTFbpc87GN0I" colab_type="code" colab={}
model.fit(
x_train,
y_train,
batch_size=64,
epochs=30
)
# + id="2arMm1WcHYxr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="6f90a394-b86a-401b-ddea-3c2179e3e79b"
score = model.evaluate(x_test, y_test)
for key, value in zip(model.metrics_names, score):
print(key, value)
| basic/2nd.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Create Analytical Data
# This notebook covers the loading of the various data collected in order to build a forecasting model for contraceptive medical products order in Cote d'Ivoire. The data is loaded into a postgres database. The data is then explored to determine how the data can be appropropriately merged to facilitate future analytical work including exploratory data analysis, model development and data visualization.
# +
import os
import pandas as pd
import numpy as np
# -
# %matplotlib inline
lc_data = pd.read_csv("final_data/contraceptive_logistics_data.csv")
lcd_date_dict = dict(year = lc_data.year,
month = lc_data.month,
day = [28]*len(lc_data))
lc_data['year_month'] = pd.to_datetime(lcd_date_dict)
lc_data.to_csv("lc_data.csv",index=False)
lc_data.head()
ccm_data = pd.read_csv("final_data/contraceptive_case_data_monthly.csv")
ccm_date_dict = dict(year = ccm_data.year,
month = ccm_data.month,
day = [28]*len(ccm_data))
ccm_data['year_month'] = pd.to_datetime(ccm_date_dict)
ccm_data.to_csv("ccm_data.csv",index=False)
ccm_data.head()
cca_data = pd.read_csv("final_data/contraceptive_case_data_annual.csv")
cca_data.head()
site_data = pd.read_csv("final_data/service_delivery_site_data.csv")
site_data.head()
product_data = pd.read_csv("final_data/product.csv")
product_data.head()
from sqlalchemy import create_engine
from dotenv import load_dotenv
load_dotenv()
def get_posgres_connection():
db_name = os.getenv("PSQL_DB_NAME")
db_user = os.getenv("PSQL_DB_USER")
db_password = <PASSWORD>("<PASSWORD>")
db_host = os.getenv("PSQL_DB_HOST")
sql_engine = create_engine(f'postgresql://{db_user}:{db_password}@{db_host}:5432/{db_name}')
return sql_engine
# ?lc_data.to_sql
# ### Load data to the database
# +
# Create db engine
psql_engine = get_posgres_connection()
# ci_logistics_contraceptive_cases
lc_data.to_sql('ci_logistics_contraceptive_cases', psql_engine, index=False,if_exists='replace')
# ci_monthly_cases
ccm_data.to_sql('ci_monthly_cases', psql_engine, index=False,if_exists='replace')
# ci_annual_cases
cca_data.to_sql('ci_annual_cases', psql_engine, index=False,if_exists='replace')
# ci_med_products
product_data.to_sql('ci_med_products', psql_engine, index=False,if_exists='replace')
# ci_med_sites
site_data.to_sql('ci_med_sites', psql_engine, index=False,if_exists='replace')
# -
# #### Load spatial data to the database
# ##### Get SRID
# +
from osgeo import ogr
driver = ogr.GetDriverByName('ESRI Shapefile')
shape = driver.Open('civ_admbnda_adm3_cntig_ocha_itos_20180706/civ_admbnda_adm3_cntig_ocha_itos_20180706.shp')
layer= shape.GetLayer()
# the crs
crs = layer.GetSpatialRef()
# from Geometry
feature = layer.GetNextFeature()
geom = feature.GetGeometryRef()
print(crs)
# -
# Imports
from geoalchemy2 import Geometry, WKTElement
from sqlalchemy import *
import pandas as pd
import geopandas as gpd
def load_shape_file_to_db(shape_file,table,engine,geom_type ='MULTIPOLYGON',if_exists ='replace',srid = 4326):
"""
This function will load a shape file to a PostGIS database system.
shape_file str: path to shapefile
table str:name of database file to load to POSTGIS
engine sqlalchemy.Engine: A sqlAlchemy POSTGIS Engine Object
if_exists str: method of upload to database, default is "replace"
srid int: format ID of shape files, this can be discovered by using the
"""
# Load shape file with geopandas
gdf = gpd.read_file(shape_file)
#Convert geometry column to WKTElement
gdf['geom'] = gdf['geometry'].apply(lambda x: WKTElement(x.wkt, srid= 4326))
#drop the geometry column as it is now duplicative
gdf.drop('geometry', 1, inplace=True)
# Use 'dtype' to specify column's type
# For the geom column, we will use GeoAlchemy's type 'Geometry'
gdf.to_sql(table, engine, if_exists= if_exists, index=False,
dtype={'geom': Geometry(geom_type, srid= 4326)})
return None
# +
# district
#Establish a postgres connection
psql_engine = get_posgres_connection()
load_shape_file_to_db('civ_admbnda_adm3_cntig_ocha_itos_20180706/civ_admbnda_adm3_cntig_ocha_itos_20180706.shp',
"cdi_shp_adm3",
psql_engine,
geom_type="POLYGON")
# +
gdf = gpd.read_file('civ_admbnda_adm3_cntig_ocha_itos_20180706/civ_admbnda_adm3_cntig_ocha_itos_20180706.shp')
gdf.head()
# -
# ### Lets start exploring the data to make an analytical dataset
districts = site_data['site_district'].unique()
print(len(districts))
districts.sort()
districts
site_data.head()
# 'ABOBO-EST' is a neighborhood in Abidjan
#
# Match city to district then aggregate the district level.
#
# Need to do some manual analysis to figure this whole matching up.
# Each city should be rolled-up into a district
region = gdf['ADM1_FR'].unique()
print(len(region))
region.sort()
region
districts = gdf['ADM2_FR'].unique()
print(len(districts))
districts.sort()
districts
site_data['site_code'].head()
gdf['ADM2_PCODE'].head()
# ### The codes are not matching up between the two dataframes
# - On inspection we can see there is an I missing, lets try tp add that and see if that fixes thing
# Insert String
ins_char = lambda x: x[0:1]+"I"+x[1:]
site_data['ADM2_PCODE'] = site_data['site_code'].apply(ins_char)
len(site_data)
len(gdf)
# #### Lets merge the two dataframes and see the overlap
mgdf = gdf.merge(site_data, on = "ADM2_PCODE")
print( len(mgdf) )
mgdf.head()
# #### Analysis
# The dataframes are not able to merged. We would like to be able to do additional analysis on the sites based on their regional geographic characteristics but currently the data does not support their merging.
#
# ## Next Steps - Questions?
# - How can we merge the data we have available into a single dataset?
# - What transformations and processing stems should we make on the data prior to loading into our database?
# - What analytical transformations on the data should we make prior to modeling of the data?
# +
import numpy as np
import matplotlib.pyplot as plt
from sklearn import datasets
from sklearn.model_selection import train_test_split
from sklearn.decomposition import PCA
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
from sklearn.neighbors import (KNeighborsClassifier,
NeighborhoodComponentsAnalysis)
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import StandardScaler
print(__doc__)
n_neighbors = 3
random_state = 42
# Load Digits dataset
X, y = datasets.load_digits(return_X_y=True)
# Split into train/test
X_train, X_test, y_train, y_test = \
train_test_split(X, y, test_size=0.5, stratify=y,
random_state=random_state)
dim = len(X[0])
n_classes = len(np.unique(y))
# Reduce dimension to 2 with PCA
pca = make_pipeline(StandardScaler(),
PCA(n_components=2, random_state=random_state))
# Reduce dimension to 2 with LinearDiscriminantAnalysis
lda = make_pipeline(StandardScaler(),
LinearDiscriminantAnalysis(n_components=2))
# Reduce dimension to 2 with NeighborhoodComponentAnalysis
nca = make_pipeline(StandardScaler(),
NeighborhoodComponentsAnalysis(n_components=2,
random_state=random_state))
# Use a nearest neighbor classifier to evaluate the methods
knn = KNeighborsClassifier(n_neighbors=n_neighbors)
# Make a list of the methods to be compared
dim_reduction_methods = [('PCA', pca), ('LDA', lda), ('NCA', nca)]
# plt.figure()
for i, (name, model) in enumerate(dim_reduction_methods):
plt.figure()
# plt.subplot(1, 3, i + 1, aspect=1)
# Fit the method's model
model.fit(X_train, y_train)
# Fit a nearest neighbor classifier on the embedded training set
knn.fit(model.transform(X_train), y_train)
# Compute the nearest neighbor accuracy on the embedded test set
acc_knn = knn.score(model.transform(X_test), y_test)
# Embed the data set in 2 dimensions using the fitted model
X_embedded = model.transform(X)
# Plot the projected points and show the evaluation score
plt.scatter(X_embedded[:, 0], X_embedded[:, 1], c=y, s=30, cmap='Set1')
plt.title("{}, KNN (k={})\nTest accuracy = {:.2f}".format(name,
n_neighbors,
acc_knn))
plt.show()
# -
# ### Data treatment
len(ccm_data["district"].unique())
#.head()
ccm_data.columns
#lc_data.head()
lc_data.columns
# +
# Either will drop 2019 or have to adjust it with the data we have for that year
by_district_lc_data = lc_data.copy()
lc_data_numeric = ["stock_received","stock_distributed","stock_adjustment",
"average_monthly_consumption","stock_stockout_days"]#,"stock_change"]
by_district_lc_data['year'] = by_district_lc_data['year'].astype(str)
by_district_lc_data = by_district_lc_data[by_district_lc_data['year'] != "2019"]
agg_by_district_lc_data = by_district_lc_data.groupby(["region","district","year"])[lc_data_numeric].agg(['count','sum','mean','std'])
agg_by_district_lc_data = agg_by_district_lc_data.unstack()
agg_by_district_lc_data.columns = ['_'.join(col) for col in agg_by_district_lc_data.columns.values]
agg_by_district_lc_data.reset_index(inplace=True)
agg_by_district_lc_data.head()
# -
#agg_by_district_lc_data.sort_values("district")
len(agg_by_district_lc_data["region"].unique())
# +
#product_df = pd.merge(product_data,agg_product_lc_data,on='product_code')
#product_df.head()
#agg_by_district_lc_data.iloc[:,2:]
# -
# ## Dimentionality reduction
# +
from sklearn.preprocessing import StandardScaler
# Select which features as numeric
agg_by_district_lc_data_numerics = agg_by_district_lc_data.iloc[:,2:]
# Standardizing the features
standardized_district_df = StandardScaler().fit_transform(agg_by_district_lc_data_numerics)
standardized_district_df = pd.DataFrame(standardized_district_df)
standardized_district_df.columns = agg_by_district_lc_data_numerics.columns.values
# -
import seaborn as sns
import matplotlib.pyplot as plt
standardized_district_df
# +
# Compute the correlation matrix
# log transform metrics for analysis
#corr = np.log(standardized_product_df+0.01).corr()
corr = standardized_district_df.corr()
# Generate a mask for the upper triangle
mask = np.triu(np.ones_like(corr, dtype=np.bool))
# Set up the matplotlib figure
f, ax = plt.subplots(figsize=(11, 9))
# Generate a custom diverging colormap
cmap = sns.diverging_palette(220, 10, as_cmap=True)
# Draw the heatmap with the mask and correct aspect ratio
sns.heatmap(corr, mask=mask, cmap=cmap,center=0,
square=True, linewidths=.5, cbar_kws={"shrink": .75})
# -
standardized_district_df = standardized_district_df.fillna(0)
from sklearn.decomposition import PCA
n_components = 10 #len(standardized_district_df) -1
pca = PCA(n_components=n_components)
principalComponents = pca.fit_transform(standardized_district_df)
principalDf = pd.DataFrame(data = principalComponents , columns = ["comp_"+str(x) for x in range(n_components)])
principalDf.plot.scatter(x="comp_0",y="comp_1")
pca_district_df = pd.concat([agg_by_district_lc_data,principalDf],axis=1)
# How much of the variance is explained
plt.plot(np.cumsum(pca.explained_variance_ratio_))
plt.xlabel('number of components')
plt.ylabel('cumulative explained variance');
plt.figure(figsize=(16,10))
sns.scatterplot(
x="comp_0", y="comp_1",
hue="region",
palette=sns.color_palette("hls", 20),
data=pca_district_df,
legend="full",
alpha=0.9
)
from sklearn.manifold import TSNE
tsne = TSNE(n_components=2, verbose=1, perplexity=40, n_iter=300)
tsne_results = tsne.fit_transform(standardized_district_df)
pca_district_df['tsne_1']= tsne_results[:,0]
pca_district_df['tsne_2'] = tsne_results[:,1]
plt.figure(figsize=(16,10))
sns.scatterplot(
x="tsne_1", y="tsne_2",
hue="region",
palette=sns.color_palette("hls", 20),
data=pca_district_df,
legend="full",
alpha=0.9
)
# +
from sklearn.cluster import KMeans
from yellowbrick.cluster.elbow import kelbow_visualizer
# Use the quick method and immediately show the figure
kelbow_visualizer(KMeans(random_state=4), principalComponents, k=(2,10))
# +
from sklearn.cluster import KMeans
from yellowbrick.cluster.elbow import kelbow_visualizer
# Use the quick method and immediately show the figure
kelbow_visualizer(KMeans(random_state=4), standardized_district_df, k=(2,10))
# -
| Create Analytical Dataset with district.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Py-ART 2.0: Field manipulation and Saving the Radar Object to a file.
#
# In this section we will go over the basics of how to change the data inside a radar file and save a new radar file. We will do this by creating a new field that stores a reflectivity mask.
# +
import pyart
import numpy as np
import os
from matplotlib import pyplot as plt
# %matplotlib inline
# -
# As before, we will load our trusty radar file!
radar = pyart.io.read(os.path.expanduser('~/data/arm/csapr_test_case.nc'))
# Let's store the radar field into a variable.
ref_field = radar.fields['reflectivity']['data']
# To create an array that is zero when the condition is false and one when it is true, we can make use of the np.ma.where command.
ref_gt_0 = np.ma.where(ref_field > 0, 1, 0)
print(ref_gt_0)
# To create a new field, we need to create a dictionary with keys containing the data, the long name, the units, the fill value, and the standard name.
mask_dict = {'data': ref_gt_0, 'units': '0 = Z < 0, 1 = Z >= 0', 'long_name': 'reflectivity_mask',
'_FillValue': ref_gt_0.fill_value, 'standard_name': 'reflectivity_mask'}
# Adding this field into the radar object is as simple as doing radar.add_field()!
radar.add_field('reflectivity_mask', mask_dict, replace_existing=True)
# We can now plot the field using RadarMapDisplayCartopy!
plt.figure(figsize=[12, 8])
display = pyart.graph.RadarMapDisplay(radar)
display.plot_ppi_map('reflectivity_mask', cmap='coolwarm', vmin=0, vmax=1)
# As you can see, we have now located all of the regions where the reflectivity is greater than 0 dBZ.
plt.figure(figsize=[12, 8])
display.plot_ppi_map('reflectivity', vmin=-10, vmax=60)
# Writing this radar object to a new file is as simple as uing pyart.io.write_cfradial()!
pyart.io.write_cfradial(os.path.expanduser('~/data/arm/new_radar.nc'), radar)
# Now, if you look in the data folder within the pyart_short_course, you should see new_radar.nc
# ## Exercise
#
# Create a field that is zero when the normalized coherent power is < 0.5 and is one when it is > 0.5
# %load section_2_answer.py
| pyart/2_pyart_changing_fields_and_saving.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import pandas as pd
from pathlib import Path
import os
import datetime
# +
sheet_id = "1vQJ3psvi0Kl6_tecq3SV_mCMuQPvnXxnQ2MLN2uc128"
sheet_name = "Sheet1"
url = f"https://docs.google.com/spreadsheets/d/{sheet_id}/gviz/tq?tqx=out:csv&sheet={sheet_name}"
df = pd.read_csv(url)
df['announce_date'] = pd.to_datetime(df['announce_date'])
df['first_release_date'] = pd.to_datetime(df['first_release_date'])
df.head()
# -
def get_suggested_links(host, row):
suggested_links = """
You might also like:
"""
if row.name in ['Apple Watch', 'iPhone']:
suggested_links += f"- [{row.category} Release Dates]({host}/categories/{row.category.lower().replace(' ','-')}/)\n"
elif row.name in ['iPad']:
suggested_links += f"- [{row.name} Release Dates]({host}/tags/{row.name.lower().replace(' ','-')}/)\n"
else:
suggested_links += f"- [All {row.category} Release Dates]({host}/categories/{row.category.lower().replace(' ','-')}/)\n"
suggested_links += f"- [{row.name} Release Dates]({host}/tags/{row.name.lower().replace(' ','-')}/)\n"
suggested_links += f"""- [Releases in {str(row.first_release_date.year)}]({host}/tags/{str(row.first_release_date.year).lower().replace(' ','-')}/)
- [All Apple Products]({host}/categories/)\n"""
return suggested_links
def get_page_content(row):
# print(type(row.first_release_date))
host = 'https://AppleReleaseDate.com'
categories = [row.category]
tags = [row.name, str(row.first_release_date.year)]
if row.release_series_tag not in tags:
tags.append(row.release_series_tag)
post_content = f"""+++
ShowToc = false
categories = {categories}
date = {str(row.first_release_date)}
title = "{get_page_name(row)}"
tags = {tags}
+++
### The {get_content_name(row)} was released on:
#### {row.first_release_date.strftime('%B %d, %Y')}
<!--more-->
{get_suggested_links(host=host,row=row)}
<kbd> {row.page_content}</kbd>
"""
return post_content
def get_content_name(row):
if row.page_name == row.full_name:
return row.full_name
return f'{row.page_name} / {row.full_name}'
def get_page_name(row):
year = str(row.first_release_date.year)
if year in row.page_name:
return f'{row.page_name}'
else:
return f'{row.page_name} ({year})'
def update_posts(git_push=True):
export_root = Path('/Users/arjun921/working_directory/apple-release-dates/content/posts')
commit_message = f"Autogenerated commit message @ {str(datetime.datetime.now())}\n\n"
for row in df.itertuples():
print('.',end=' ')
folder_path = export_root / Path(row.category)
post_path = folder_path / Path(get_page_name(row)+'.md')
os.makedirs(folder_path, exist_ok=True)
log = f'Generated/Updated post for: {get_page_name(row)}'
commit_message += f'{log}\n'
fopen = open(post_path,'w')
page_content = get_page_content(row)
fopen.write(page_content)
fopen.close()
print('')
if git_push:
os.system(f'cd {folder_path.parent.parent.parent} && git add . && git commit -m "{commit_message}" && git push')
update_posts(True)
| generator/generate_blogposts.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import tridy
from tridy import GeoConcept, SubGeoConcept, MetaData, Table, View, DBStorage, DataSource, Feature, FeatureWithID, AdmUnitFeature, OLUFeature, Grid, Imagee, ds_from_metadata, xml_lpis_cz_reader, lpis_cz__posledni_aktualizace, get_listvalues_from_generator, apply_function, select_nodes_from_graph, unzip_file, find_neighbors_till, connection_parameters_to_pg, transform_name_to_postgresql_format, world_to_pixel
from importlib import reload
import requests
import datetime
import re
from io import BytesIO
from requests.adapters import HTTPAdapter
from requests.packages.urllib3.util.retry import Retry
from osgeo import ogr, osr, gdal
import networkx as nx
import numpy as np
import json
import binascii
import copy
import time
from lxml import etree
import random
from ipyleaflet import Map, GeoJSON
# -
del(GeoConcept, SubGeoConcept, MetaData, Table, View, DBStorage, DataSource, Feature, FeatureWithID, AdmUnitFeature, OLUFeature, Grid, Imagee, ds_from_metadata,xml_lpis_cz_reader,get_listvalues_from_generator,apply_function,select_nodes_from_graph,world_to_pixel)
reload(tridy)
from tridy import GeoConcept, SubGeoConcept, MetaData, Table, View, DBStorage, DataSource, Feature, FeatureWithID, AdmUnitFeature, OLUFeature, Grid, Imagee, ds_from_metadata, xml_lpis_cz_reader, get_listvalues_from_generator, apply_function, select_nodes_from_graph,world_to_pixel
def random_color(feature):
return {
'color': 'black',
'fillColor': random.choice(['red', 'yellow', 'green', 'orange']),
}
#administrative territorial units
dbs_admin_connection={'dbname':'olu_administrative_units','user':'euxdat_admin','host':'euxdat-db-svc','port':'5432','password':'<PASSWORD>'}
dbs_admin=DBStorage(dbs_admin_connection)
dbs_admin.connect()
dbs_admin.disconnect()
dbs_admin.connect()
replacement_dictionary = {"[posledni_den_mesice]":(datetime.datetime.today().replace(day=1)-datetime.timedelta(days=1)).strftime('%Y%m%d'),"[lpis_cz__posledni_aktualizace]":lpis_cz__posledni_aktualizace().strftime('%Y%m%d'), "[vcera]":(datetime.datetime.today().replace(day=1)-datetime.timedelta(days=1)).strftime('%Y%m%d')}
json_feature_structure=[{"name":"id","type":"serial primary key"},{"name":"geom","type":"geometry"},{"name":"data","type":"json"}]
json_feature_with_bigid_structure=[{"name":"id","type":"bigint primary key"},{"name":"geom","type":"geometry"},{"name":"data","type":"json"}]
json_admin_unit_structure=[{"name":"id","type":"integer primary key"},{"name":"geom","type":"geometry"},{"name":"data","type":"json"},{"name":"level","type":"integer"},{"name":"parent_id","type":"text"}]
json_admin_unit_structure_at=[{"name":"id","type":"text primary key"},{"name":"geom","type":"geometry"},{"name":"data","type":"json"},{"name":"level","type":"integer"},{"name":"parent_id","type":"text"}]
json_feature_with_raster_structure=[{"name":"id","type":"serial primary key"},{"name":"geom","type":"geometry"},{"name":"data","type":"json"},{"name":"raster_maps","type":"raster"}]
# +
admunit_at__metadata=MetaData('Administrative units in Austria',
{"url":"http://gis.lesprojekt.cz/admunit_at.zip", "format":"ESRI Shapefile","compression":"zip"},
'data')
admunit_at__ds=ds_from_metadata(admunit_at__metadata)
admunit_at=GeoConcept('Administrative units in Austria','Administrative units in Austria. All levels.',
'AdmUnitFeature',json_admin_unit_structure_at, data_source=admunit_at__ds, subgeoconcepts=[] )
# -
#admunit_at.get_data_source().download_data('admunit_at.zip', s, 'all', folder='admunit_at/')
#admunit_at.get_data_source().set_data_file([file for file in admunit_at.get_data_source().get_data_file() if file.endswith('shp')][0])
admunit_at.get_data_source().set_data_file('admunit_at/AU_AdministrativeUnit_extended.shp')
concept_list=['1stOrder','2ndOrder','3rdOrder','4thOrder','5thOrder']
concept_additional_attributes={'1stOrder':{'level_value':0,'parent_value':'null','id_attribute':'inspireId'},
'2ndOrder':{'level_value':1,'parent_attribute':'AU_Attri_6','id_attribute':'inspireId'},
'3rdOrder':{'level_value':2,'parent_attribute':'AU_Attri_6','id_attribute':'inspireId'},
'4thOrder':{'level_value':3,'parent_attribute':'AU_Attri_6','id_attribute':'inspireId'},
'5thOrder':{'level_value':4,'parent_attribute':'AU_Attri_6','id_attribute':'inspireId'}}
for l in concept_list:
admunit_at.append_subgeoconcept(SubGeoConcept(l,l,'AdmUnitFeature',admunit_at.get_attributes(),data_source=DataSource(admunit_at.get_data_source().get_type(),admunit_at.get_data_source().get_name(),({**admunit_at.get_data_source().get_attributes(),**{'attribute_filter':"AU_Attri_2='%s'"%l}}),None,admunit_at.get_data_source().get_data_file()),supergeoconcept=admunit_at,table_inheritance=False,subgeoconcepts=[]))
admunit_at.create_table(dbs_admin, name='default',scheme='at',conflict='append')
# +
#for sub in admunit_at.get_subgeoconcepts():
# features=sub.get_data_source().read_features('admunitfeature',concept_additional_attributes[sub.get_name()],number=10)
# dbs_admin.insert_many('insert into %s.%s (geom,data,id,level,parent_id) ' % (admunit_at.get_table().get_scheme(),admunit_at.get_table().get_name()) ,features,20)
# -
for sub in admunit_at.get_subgeoconcepts():
sub.set_table(View(transform_name_to_postgresql_format(sub.get_name()),sub.get_attributes(), sub.get_supergeoconcept().get_table(),"level=%s" % (concept_additional_attributes[sub.get_name()]['level_value']), dbs=dbs_admin, scheme='public', type='usual'))
dbs_admin.execute(sub.get_table().create_script())
sub=admunit_at.get_subgeoconcept_by_name('2ndOrder')
with open('2ndOrder.geojson', 'w', encoding='utf-8') as file:
geojson={"type": "FeatureCollection", "features": [] }
features=sub.read_features_from_table(100)
for f in features:
if len(f)>0:
for feature in f:
geojson["features"].append(feature.export_to_geojson())
else:
break
json.dump(geojson, file, ensure_ascii=False, indent=4)
# +
with open('2ndOrder.geojson', 'r') as f:
data = json.load(f)
m = Map(center=(47.8,13), zoom=7)
geo_json = GeoJSON(
data=data,
style={
'opacity': 1, 'dashArray': '9', 'fillOpacity': 0.1, 'weight': 1
},
hover_style={
'color': 'white', 'dashArray': '0', 'fillOpacity': 0.5
},
style_callback=random_color
)
m.add_layer(geo_json)
m
# -
| jupyter_examples/administrative_units_in_austria.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Given an array of integers, return indices of the two numbers such that they add up to a specific target.
#
# You may assume that each input would have exactly one solution, and you may not use the same element twice.
#
# Example:
# ```
# Given nums = [2, 7, 11, 15], target = 9,
#
# Because nums[0] + nums[1] = 2 + 7 = 9,
# return [0, 1].
# ```
class Solution(object):
def twoSum(self, nums, target):
"""
:type nums: List[int]
:type target: int
:rtype: List[int]
"""
lookup = dict((v, i) for i, v in enumerate(nums)) # dict, later replace previous idxs
print(lookup)
return next(
(
(i, lookup.get(target-v)) # get index of num with its pair which sums up to target
for i,v in enumerate(nums) # enumerate nums
if lookup.get(target-v, i) != i # if the two indexes are not equal, i.e. same number doesn't count
),
None)
if __name__ == "__main__":
nums=(6, 7, 11, 15, 3, 6, 5, 3)
target=6
print(Solution().twoSum(nums,target))
# +
nums=(6, 7, 11, 15, 3, 6, 5, 3)
lookup = dict((v, i) for i, v in enumerate(nums))
lookup.get(3)
# -
next((i, lookup.get(target-v)) # get index of num with its pair which sums up to target
for i,v in enumerate(nums) # enumerate nums
if lookup.get(target-v, i) != i # v = 3, target = 6, i = 4,
)
lookup.get(3,4)
| 1.TwoSum.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # PIV analysis and Plotting
# Code set using data from PIV experiments on 09-21-17<br>
# $U_\infty = 4.5 (400rpm)$<br>
# Test 0: <br>
# Data taken at 500Hz continuously<br>
# Test 1: <br>
# Data taken at 500Hz for 100 images on a 1Hz loop<br>
# Test 2: <br>
# Data taken at 500Hz for 500 images on a .5Hz loop<br>
# Test 3: <br>
# Data taken at 500Hz continuously<br><br>
# Laser Power = 14amps <br>
# Last updated: 09-26-17 <br>
# Code Strucutre: <br>
# - import libraries <br>
# - run analysis codes <br>
# - read in data <br>
# - plot outer <br>
# - plot inner <br>
import pandas as pd
import numpy as np
import PIV as piv
import time_series as ts
import time
import sys
import h5py
from scipy.signal import medfilt
import matplotlib.pyplot as plt
import hotwire as hw
import imp
from datetime import datetime
# %matplotlib inline
now = datetime.now()
#for setting movie
import time
import pylab as pl
from IPython import display
# import functions to be run
imp.reload(ts)
imp.reload(piv)
imp.reload(hw)
# %run 'air_prop.py'
# %run 'piv_outer.py'
# %run 'piv_readin.py'
# %run 'piv_inner.py'
# # Read in and Filter Datasets
## DATA SET READ IN ##
#data sets taken continuously (test_0, test_3)
#Parameter set
date = '092117_0'
data_delimiter = '\t'
num_images = 10917
sizex = 128
sizey = 129
walloffset = 2 #mm
side_error = 5
#determine file name
file_name = dict()
for j in range(1, num_images+1):
file_name[j] = '/B' + str('{0:05}'.format(j)) + '.txt'
#list name of data set folders
base_name = dict()
#List the base name for each test to be read in and analyzed, names taken directly from folder
base_name[0] = '/media/drummond/My Passport/DATA/FPF/test_092117/Cam_Date=170921_Time=120913_TR_SeqPIV_MP(1x16x16_50ov_ImgCorr)=unknown'
base_name[1] = '/media/drummond/My Passport/DATA/FPF/test_092117/Cam_Date=170921_Time=140859_TR_SeqPIV_MP(1x16x16_50ov_ImgCorr)=unknown'
[u, v, x, y, bad_im] = piv_readin(date, file_name, base_name, num_images, data_delimiter, sizex, sizey, walloffset, side_error)
# +
## DATA SET READ IN ##
#data set taken on cycle, 100 images every 1hz (test_1)
#Parameter set
date = '092117_1'
data_delimiter = '\t'
num_images = 10907
sizex = 128
sizey = 129
walloffset = 2 #mm
side_error = 5
#determine file name
file_name = dict()
for j in range(1, num_images+1):
file_name[j] = '/B' + str('{0:05}'.format(j)) + '.txt'
#list name of data set folders
base_name = dict()
#List the base name for each test to be read in and analyzed, names taken directly from folder
base_name[0] = '/media/drummond/My Passport/DATA/FPF/test_092117/Cam_Date=170921_Time=124152_TR_SeqPIV_MP(1x16x16_50ov_ImgCorr)=unknown'
piv_readin(date, file_name, base_name, num_images, data_delimiter, sizex, sizey, walloffset, side_error)
# -
## DATA SET READ IN ##
#data set taken on cycle, 500 images every .5hz (test_1)
#Parameter set
date = '092117_2'
data_delimiter = '\t'
num_images = 10520
sizex = 128
sizey = 129
walloffset = 2 #mm
side_error = 5
#determine file name
file_name = dict()
for j in range(1, num_images+1):
file_name[j] = '/B' + str('{0:05}'.format(j)) + '.txt'
#list name of data set folders
base_name = dict()
#List the base name for each test to be read in and analyzed, names taken directly from folder
base_name[0] = '/media/drummond/My Passport/DATA/FPF/test_092117/Cam_Date=170921_Time=130741_TR_SeqPIV_MP(1x16x16_50ov_ImgCorr)=unknown'
piv_readin(date, file_name, base_name, num_images, data_delimiter, sizex, sizey, walloffset, side_error)
# # Mean Velocity Plots
# Plot Outer Normalized Data
date = '092117'
legend = [r'$Re_{\theta}=$30288, Cont.', r'$Re_{\theta}=$30288, 100im', r'$Re_{\theta}=$30288, 500im']
num_tests = 3
piv_outer(date, num_tests, legend)
# # Inner Normalized Plots
##Plot Inner Normalized Data##
date = '092117'
num_tests = 3
utau = .15
legend = [r'$Re_{\theta}=$30288, Cont.', r'$Re_{\theta}=$30288, 100im', r'$Re_{\theta}=$30288, 500im']
piv_inner(date, num_tests, utau, legend)
# # Control Volume Analysis
# Procedure: <br>
# 1). Create mean velocity field from mean u and v velocity fields ($(u^2 + v^2)^{1/2}$)<br>
# 2). Integrate left side of image (control volume in) <br>
# 3). Integrate right side of image (control volumne out) <br>
# 4). Calculate difference (control volume delta) and divide by streamwise length of FOV
# +
## Control Volume Analysis ##
umean = np.nanmean(u[0], axis=0)
vmean = np.nanmean(v[0], axis=0)
mean_vel = np.sqrt(umean**2 + vmean**2)
#print(np.shape(mean_vel))
cv_in = np.trapz(mean_vel[:, 0], x = y)*-1
cv_out = np.trapz(mean_vel[:, -1], x = y)*-1
cv_delta = cv_out - cv_in
vel_out_y = cv_delta / (x[-1] - x[0])
plt.figure(num=None, figsize=(8, 6), dpi=100, facecolor='w', edgecolor='k')
plt.semilogx(y, mean_vel[:, 0], '-xb')
plt.semilogx(y, mean_vel[:, -1], '-xr')
plt.legend(['Control Volume In', 'Control Volume Out'])
plt.ylabel('Velocity Magnitude (m/sec)')
plt.xlabel('Wall Normal Position (m)')
# + [markdown] variables={" x[-1] - x[0] ": "0.11123059999999999", "print('%.4g'%(cv_delta))": "0.001073", "print('%.4g'%(cv_in))": "0.6526", "print('%.4g'%(cv_out))": "0.6537", "print('%.4g'%(vel_out_y))": "0.009646"}
# The control volume input is {{print('%.4g'%(cv_in))}} $m^2/sec$ <br>
# The control volume ouput is {{print('%.4g'%(cv_out))}} $m^2/sec$ <br>
# Giving a difference of {{print('%.4g'%(cv_delta))}} <br>
# For which the top length of the control volume is {{ x[-1] - x[0] }}m <br>
# Giving the average v velocity to be {{print('%.4g'%(vel_out_y))}} $m/sec$
#
# -
# # Pixel Locking
# Procedure: Take masked and filtered datasets <br>
# 1). Convert into 1-D vector of all velocities (10520 images x 127 rows x 97 columns) <br>
# 2). Convert into displacement using known image frequency ($500hz$) <br>
# 3). Convert into # of pixel displacement by using calibration size ($.2mm/pixel$)
freq = 500 #hz
pixel_size = 0.0002 #mm/pixel
#calculate displacement in x dir
x_disp = u[0]*(1/freq)
#organize into 1-d vector
x_disp = np.array(np.reshape(x_disp, [1, 127*68*10917]))[0]
#calculate in pixel disp
x_disp = x_disp / pixel_size
#plot
plt.figure(num=None, figsize=(10, 8), dpi=100, facecolor='w', edgecolor='k')
plt.hist(x_disp[0:1000000], bins=5000, range=[20, 40], normed=True)
plt.title('Streamwise Velocity pixel displacement PDF')
plt.xlabel('Pixel Displacement')
plt.ylabel('Normalize Counts')
plt.show()
np.shape(u)
#calculate displacement in y dir
y_disp = v[0]*(1/freq)
#organize into 1-d vector
y_disp = np.array(np.reshape(y_disp, [1, 127*68*10917]))[0]
#calculate in pixel disp
y_disp = y_disp / pixel_size
#plot
plt.figure(num=None, figsize=(10, 8), dpi=100, facecolor='w', edgecolor='k')
plt.hist(y_disp[:1000000], bins=5000, normed=True)
plt.title('Wall-normal Velocity pixel displacement PDF')
plt.xlabel('Pixel Displacement')
plt.ylabel('Normalize Counts')
plt.show()
# # Autocorrelation Plot
# Procedure: <br>
# -working to examine when each point in the velocity field becomes time indep. <br>
# - in development
np.shape(u)
# # Conculsions
# Velocity plots: <br>
# - Mean data matches well with other experimental and DNS datasets <br>
# - U component collapes for all statistics examined <br>
# - V component is correct order but is not resolved due to low dynamic range <br>
# - CONTROL VOLUME ANALYSIS: V is order .001 $m/sec$ <br>
# - PIXEL DISP PDFs: No evidence of pixel locking <br>
#
# # To Do
# - Increase dynamic range through increasing resolution of v velocity displacement by increasing $Re$ and decreasing image frame rate ($\Delta T$)
| PIV_092117.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/NeuromatchAcademy/course-content-dl/blob/main/projects/ComputerVision/transfer_learning.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a> <a href="https://kaggle.com/kernels/welcome?src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content-dl/main/projects/ComputerVision/transfer_learning.ipynb" target="_parent"><img src="https://kaggle.com/static/images/open-in-kaggle.svg" alt="Open in Kaggle"/></a>
# -
# # Transfer Learning
#
# **By Neuromatch Academy**
#
# __Content creators:__ [<NAME>](https://engmubarak48.github.io/jmohamud/index.html) & [<NAME>](https://alexhernandezgarcia.github.io/)
#
# __Production editors:__ <NAME>, <NAME>
#
#
# **Our 2021 Sponsors, including Presenting Sponsor Facebook Reality Labs**
#
# <p align='center'><img src='https://github.com/NeuromatchAcademy/widgets/blob/master/sponsors.png?raw=True'/></p>
# ---
# # Objective
#
# One desired capability for machines is the ability to transfer the knowledge (features) learned on one domain to another This can potentially save compute time, enable training when data is scarce, and even improve performance. Unfortunately, there is no single recipe for transfer learning and instead multiple options are possible and much remains to be well understood. In this project, you will explore how transfer learning works in different scenarios.
# ---
# # Setup
# +
# imports
import os
import gc
import csv
import glob
import torch
import multiprocessing
import numpy as np
import pandas as pd
import torch.nn as nn
import matplotlib.pyplot as plt
import torch.optim as optim
import torch.nn.functional as F
import torch.backends.cudnn as cudnn
from torch.autograd import Variable
import torchvision
import torchvision.transforms as transforms
# + cellView="form"
# @title Set random seed
# @markdown Executing `set_seed(seed=seed)` you are setting the seed
# for DL its critical to set the random seed so that students can have a
# baseline to compare their results to expected results.
# Read more here: https://pytorch.org/docs/stable/notes/randomness.html
# Call `set_seed` function in the exercises to ensure reproducibility.
import random
import torch
def set_seed(seed=None, seed_torch=True):
if seed is None:
seed = np.random.choice(2 ** 32)
random.seed(seed)
np.random.seed(seed)
if seed_torch:
torch.manual_seed(seed)
torch.cuda.manual_seed_all(seed)
torch.cuda.manual_seed(seed)
torch.backends.cudnn.benchmark = False
torch.backends.cudnn.deterministic = True
print(f'Random seed {seed} has been set.')
# In case that `DataLoader` is used
def seed_worker(worker_id):
worker_seed = torch.initial_seed() % 2**32
np.random.seed(worker_seed)
random.seed(worker_seed)
# + cellView="form"
# @title Set device (GPU or CPU)
# inform the user if the notebook uses GPU or CPU.
def set_device():
device = "cuda" if torch.cuda.is_available() else "cpu"
if device != "cuda":
print("WARNING: For this notebook to perform best, "
"if possible, in the menu under `Runtime` -> "
"`Change runtime type.` select `GPU` ")
else:
print("GPU is enabled in this notebook.")
return device
# -
# ### Random seeds
#
# If you want to obtain reproducible results, it is a good practice to set seeds for the random number generators of the various libraries
set_seed(seed=2021)
device = set_device()
# ### Training hyperparameters
#
# Here we set some general training hyperparameters such as the learning rate, batch size, etc. as well as other training options such as including data augmentation (`torchvision_transforms`).
# hyper-parameters
use_cuda = torch.cuda.is_available()
best_acc = 0 # best test accuracy
start_epoch = 0 # start from epoch 0 or last checkpoint epoch
batch_size = 128
max_epochs = 15 # Please change this to 200
max_epochs_target = 10
base_learning_rate = 0.1
torchvision_transforms = True # True/False if you want use torchvision augmentations
# ---
# # Data
# ## Source dataset
#
# We will train the source model using CIFAR-100 data set from PyTorch, but with small tweaks we can get any other data we are interested in.
#
# Note that the data set is normalised by substracted the mean and dividing by the standard deviation (pre-computed) of the training set. Also, if `torchvision_transforms` is `True`, data augmentation will be applied during training.
# + cellView="form"
# @markdown Download and prepare Data
print('==> Preparing data..')
def percentageSplit(full_dataset, percent = 0.0):
set1_size = int(percent * len(full_dataset))
set2_size = len(full_dataset) - set1_size
final_dataset, _ = torch.utils.data.random_split(full_dataset, [set1_size, set2_size])
return final_dataset
# CIFAR100 normalizing
mean = [0.5071, 0.4866, 0.4409]
std = [0.2673, 0.2564, 0.2762]
# CIFAR10 normalizing
# mean = (0.4914, 0.4822, 0.4465)
# std = (0.2023, 0.1994, 0.2010)
# torchvision transforms
transform_train = transforms.Compose([])
if torchvision_transforms:
transform_train.transforms.append(transforms.RandomCrop(32, padding=4))
transform_train.transforms.append(transforms.RandomHorizontalFlip())
transform_train.transforms.append(transforms.ToTensor())
transform_train.transforms.append(transforms.Normalize(mean, std))
transform_test = transforms.Compose([
transforms.ToTensor(),
transforms.Normalize(mean, std),
])
trainset = torchvision.datasets.CIFAR100(
root='./CIFAR100', train=True, download=True, transform=transform_train)
testset = torchvision.datasets.CIFAR100(
root='./CIFAR100', train=False, download=True, transform=transform_test)
# -
# ### CIFAR-100
#
# CIFAR-100 is a data set of 50,000 colour (RGB) training images and 10,000 test images, of size 32 x 32 pixels. Each image is labelled as 1 of 100 possible classes.
#
# The data set is stored as a custom `torchvision.datasets.cifar.CIFAR` object. You can check some of its properties with the following code:
print(f"Object type: {type(trainset)}")
print(f"Training data shape: {trainset.data.shape}")
print(f"Test data shape: {testset.data.shape}")
print(f"Number of classes: {np.unique(trainset.targets).shape[0]}")
# ## Data loaders
#
# A dataloader is an optimized data iterator that provides functionality for efficient shuffling, transformation and batching of the data.
# + cellView="form"
##@title Dataloader
num_workers = multiprocessing.cpu_count()
print(f'----> number of workers: {num_workers}')
trainloader = torch.utils.data.DataLoader(
trainset, batch_size=batch_size, shuffle=True, num_workers=num_workers)
testloader = torch.utils.data.DataLoader(
testset, batch_size=batch_size, shuffle=False, num_workers=num_workers)
# -
# ## Architecture: ResNet
#
# ResNet is a family of network architectures whose main property is that the network is organised as a stack of _residual blocks_. Residual blocks consist of a stack of layers whose output is added the input, making a _shortcut connection_.
#
# See the [original paper](https://arxiv.org/abs/1512.03385) for more details.
#
# ResNet is just a popular choice out of many others, but data augmentation works well in general. We just picked ResNet for illustration purposes.
# + cellView="form"
# @title ResNet model in PyTorch
class BasicBlock(nn.Module):
"""ResNet in PyTorch.
Reference:
[1] <NAME>, <NAME>, <NAME>, <NAME>
Deep Residual Learning for Image Recognition. arXiv:1512.03385
"""
expansion = 1
def __init__(self, in_planes, planes, stride=1):
super(BasicBlock, self).__init__()
self.conv1 = nn.Conv2d(in_planes, planes, kernel_size=3, stride=stride, padding=1, bias=False)
self.bn1 = nn.BatchNorm2d(planes)
self.conv2 = nn.Conv2d(planes, planes, kernel_size=3, stride=1, padding=1, bias=False)
self.bn2 = nn.BatchNorm2d(planes)
self.shortcut = nn.Sequential()
if stride != 1 or in_planes != self.expansion*planes:
self.shortcut = nn.Sequential(
nn.Conv2d(in_planes, self.expansion*planes, kernel_size=1, stride=stride, bias=False),
nn.BatchNorm2d(self.expansion*planes)
)
def forward(self, x):
out = F.relu(self.bn1(self.conv1(x)))
out = self.bn2(self.conv2(out))
out += self.shortcut(x)
out = F.relu(out)
return out
class Bottleneck(nn.Module):
expansion = 4
def __init__(self, in_planes, planes, stride=1):
super(Bottleneck, self).__init__()
self.conv1 = nn.Conv2d(in_planes, planes, kernel_size=1, bias=False)
self.bn1 = nn.BatchNorm2d(planes)
self.conv2 = nn.Conv2d(planes, planes, kernel_size=3, stride=stride, padding=1, bias=False)
self.bn2 = nn.BatchNorm2d(planes)
self.conv3 = nn.Conv2d(planes, self.expansion*planes, kernel_size=1, bias=False)
self.bn3 = nn.BatchNorm2d(self.expansion*planes)
self.shortcut = nn.Sequential()
if stride != 1 or in_planes != self.expansion*planes:
self.shortcut = nn.Sequential(
nn.Conv2d(in_planes, self.expansion*planes, kernel_size=1, stride=stride, bias=False),
nn.BatchNorm2d(self.expansion*planes)
)
def forward(self, x):
out = F.relu(self.bn1(self.conv1(x)))
out = F.relu(self.bn2(self.conv2(out)))
out = self.bn3(self.conv3(out))
out += self.shortcut(x)
out = F.relu(out)
return out
class ResNet(nn.Module):
def __init__(self, block, num_blocks, num_classes=100):
super(ResNet, self).__init__()
self.in_planes = 64
self.conv1 = nn.Conv2d(3, 64, kernel_size=3, stride=1, padding=1, bias=False)
self.bn1 = nn.BatchNorm2d(64)
self.layer1 = self._make_layer(block, 64, num_blocks[0], stride=1)
self.layer2 = self._make_layer(block, 128, num_blocks[1], stride=2)
self.layer3 = self._make_layer(block, 256, num_blocks[2], stride=2)
self.layer4 = self._make_layer(block, 512, num_blocks[3], stride=2)
self.linear = nn.Linear(512*block.expansion, num_classes)
def _make_layer(self, block, planes, num_blocks, stride):
strides = [stride] + [1]*(num_blocks-1)
layers = []
for stride in strides:
layers.append(block(self.in_planes, planes, stride))
self.in_planes = planes * block.expansion
return nn.Sequential(*layers)
def forward(self, x):
out = F.relu(self.bn1(self.conv1(x)))
out = self.layer1(out)
out = self.layer2(out)
out = self.layer3(out)
out = self.layer4(out)
out = F.avg_pool2d(out, 4)
out = out.view(out.size(0), -1)
out = self.linear(out)
return out
def ResNet18():
return ResNet(BasicBlock, [2, 2, 2, 2])
def ResNet34():
return ResNet(BasicBlock, [3, 4, 6, 3])
def ResNet50():
return ResNet(Bottleneck, [3, 4, 6, 3])
# -
# #### Test on random data
# +
# Load the Model
net = ResNet18()
print('-----> verify if model is run on random data')
y = net(Variable(torch.randn(1,3,32,32)))
print('model loaded')
result_folder = './results/'
if not os.path.exists(result_folder):
os.makedirs(result_folder)
logname = result_folder + net.__class__.__name__ + '_pretrain' + '.csv'
if use_cuda:
net.cuda()
net = torch.nn.DataParallel(net)
print('Using', torch.cuda.device_count(), 'GPUs.')
cudnn.benchmark = True
print('Using CUDA..')
# -
# ## Set up training
# ### Set loss function and optimizer
#
# We use the cross entropy loss, commonly used for classification, and stochastic gradient descent (SGD) as optimizer, with momentum and weight decay.
# +
# Optimizer and criterion
criterion = nn.CrossEntropyLoss()
optimizer = optim.SGD(net.parameters(), lr=base_learning_rate, momentum=0.9, weight_decay=1e-4)
# -
# ### Train and test loops
# +
# Training & Test functions
def train(net, epoch, use_cuda=True):
print('\nEpoch: %d' % epoch)
net.train()
train_loss = 0
correct = 0
total = 0
for batch_idx, (inputs, targets) in enumerate(trainloader):
if use_cuda:
inputs, targets = inputs.cuda(), targets.cuda()
optimizer.zero_grad()
inputs, targets = Variable(inputs), Variable(targets)
outputs = net(inputs)
loss = criterion(outputs, targets)
loss.backward()
optimizer.step()
train_loss += loss.item()
_, predicted = torch.max(outputs.data, 1)
total += targets.size(0)
correct += predicted.eq(targets.data).cpu().sum()
if batch_idx % 500 == 0:
print(batch_idx, len(trainloader), 'Loss: %.3f | Acc: %.3f%% (%d/%d)'
% (train_loss/(batch_idx+1), 100.*correct/total, correct, total))
return (train_loss/batch_idx, 100.*correct/total)
def test(net, epoch, outModelName, use_cuda=True):
global best_acc
net.eval()
test_loss, correct, total = 0, 0, 0
with torch.no_grad():
for batch_idx, (inputs, targets) in enumerate(testloader):
if use_cuda:
inputs, targets = inputs.cuda(), targets.cuda()
outputs = net(inputs)
loss = criterion(outputs, targets)
test_loss += loss.item()
_, predicted = torch.max(outputs.data, 1)
total += targets.size(0)
correct += predicted.eq(targets.data).cpu().sum()
if batch_idx % 200 == 0:
print(batch_idx, len(testloader), 'Loss: %.3f | Acc: %.3f%% (%d/%d)'
% (test_loss/(batch_idx+1), 100.*correct/total, correct, total))
# Save checkpoint.
acc = 100.*correct/total
if acc > best_acc:
best_acc = acc
checkpoint(net, acc, epoch, outModelName)
return (test_loss/batch_idx, 100.*correct/total)
# -
# ### Auxiliary functions
#
# * `checkpoint()`: Store checkpoints of the model
# * `adjust_learning_rate()`: Decreases the learning rate (learning rate decay) at certain epochs of training.
# +
# checkpoint & adjust_learning_rate
def checkpoint(model, acc, epoch, outModelName):
# Save checkpoint.
print('Saving..')
state = {
'state_dict': model.state_dict(),
'acc': acc,
'epoch': epoch,
'rng_state': torch.get_rng_state()
}
if not os.path.isdir('checkpoint'):
os.mkdir('checkpoint')
torch.save(state, f'./checkpoint/{outModelName}.t7')
def adjust_learning_rate(optimizer, epoch):
"""decrease the learning rate at 100 and 150 epoch"""
lr = base_learning_rate
if epoch <= 9 and lr > 0.1:
# warm-up training for large minibatch
lr = 0.1 + (base_learning_rate - 0.1) * epoch / 10.
if epoch >= 100:
lr /= 10
if epoch >= 150:
lr /= 10
for param_group in optimizer.param_groups:
param_group['lr'] = lr
# -
# ### Train the model
#
# This is the loop where the model is trained for `max_epochs` epochs.
# +
# Start training
outModelName = 'pretrain'
if not os.path.exists(logname):
with open(logname, 'w') as logfile:
logwriter = csv.writer(logfile, delimiter=',')
logwriter.writerow(['epoch', 'train loss', 'train acc', 'test loss', 'test acc'])
for epoch in range(start_epoch, max_epochs):
adjust_learning_rate(optimizer, epoch)
train_loss, train_acc = train(net, epoch, use_cuda=use_cuda)
test_loss, test_acc = test(net, epoch, outModelName, use_cuda=use_cuda)
with open(logname, 'a') as logfile:
logwriter = csv.writer(logfile, delimiter=',')
logwriter.writerow([epoch, train_loss, train_acc.item(), test_loss, test_acc.item()])
print(f'Epoch: {epoch} | train acc: {train_acc} | test acc: {test_acc}')
# -
# ## Transfer learning
# ### Re-use the trained model to improve training on a different data set
# ### Delete variables from the previous model
# delete the backbone network
delete = True
if delete:
del net
del trainset
del testset
del trainloader
del testloader
gc.collect()
# #### Target dataset
#
# We will now use CIFAR-10 as _target_ data set. Again, with small tweaks we can get any other data we are interested in.
#
# CIFAR-10 is very similar to CIFAR-100, but it contains only 10 classes instead of 100.
# +
# Target domain Data
print('==> Preparing target domain data..')
# CIFAR10 normalizing
mean = (0.4914, 0.4822, 0.4465)
std = (0.2023, 0.1994, 0.2010)
num_classes = 10
lr = 0.0001
# torchvision transforms
transform_train = transforms.Compose([])
if torchvision_transforms:
transform_train.transforms.append(transforms.RandomCrop(32, padding=4))
transform_train.transforms.append(transforms.RandomHorizontalFlip())
transform_train.transforms.append(transforms.ToTensor())
transform_train.transforms.append(transforms.Normalize(mean, std))
transform_test = transforms.Compose([
transforms.ToTensor(),
transforms.Normalize(mean, std),
])
trainset = torchvision.datasets.CIFAR10(
root='./CIFAR10', train=True, download=True, transform=transform_train)
testset = torchvision.datasets.CIFAR10(
root='./CIFAR10', train=False, download=True, transform=transform_test)
# -
# #### Select a subset of the data
#
# To simulate a lower data regime, where transfer learning can be useful.
#
# Choose percentage from the trainset. Set `percent = 1.0` to use the whole train data
# +
percent = 0.6
trainset = percentageSplit(trainset, percent = percent)
print('size of the new trainset: ', len(trainset))
# -
# #### Dataloaders
#
# As before
# +
# Dataloader
num_workers = multiprocessing.cpu_count()
print(f'----> number of workers: {num_workers}')
trainloader = torch.utils.data.DataLoader(
trainset, batch_size=batch_size, shuffle=True, num_workers=num_workers)
testloader = torch.utils.data.DataLoader(
testset, batch_size=batch_size, shuffle=False, num_workers=num_workers)
# -
# ### Load pre-trained model
#
# Load the checkpoint of the model previously trained on CIFAR-100
# +
model = ResNet18()
checkpointPath = '/content/checkpoint/pretrain.t7'
print(' ===> loading pretrained model from: ', checkpointPath)
if os.path.isfile(checkpointPath):
state_dict = torch.load(checkpointPath)
best_acc = state_dict['acc']
print('Best Accuracy:', best_acc)
if "state_dict" in state_dict:
state_dict = state_dict["state_dict"]
# remove prefixe "module."
state_dict = {k.replace("module.", ""): v for k, v in state_dict.items()}
for k, v in model.state_dict().items():
if k not in list(state_dict):
print('key "{}" could not be found in provided state dict'.format(k))
elif state_dict[k].shape != v.shape:
print('key "{}" is of different shape in model and provided state dict'.format(k))
state_dict[k] = v
msg = model.load_state_dict(state_dict, strict=False)
print("Load pretrained model with msg: {}".format(msg))
else:
raise Exception('No pretrained weights found')
# -
# ### Freeze model parameters
#
# In transfer learning, we usually do not re-train all the weights of the model, but only a subset of them, for instance the last layer. Here we first _freeze_ all the parameters of the model, and we will _unfreeze_ one layer below.
# +
# Freeze the model parameters, you can also freeze some layers only
for param in model.parameters():
param.requires_grad = False
# -
# ### Loss function, optimizer and _unfreeze_ last layer
# +
num_ftrs = model.linear.in_features
model.linear = nn.Linear(num_ftrs, num_classes)
model.to(device)
criterion = nn.CrossEntropyLoss()
optimizer = torch.optim.SGD(
model.linear.parameters(),
lr=lr,
momentum=0.9,
weight_decay=1e-4,
)
# -
# #### Check number of parameters
#
# We can calculate the number of total parameters and the number of trainable parameters, that is those that will be updated during training. Since we have freezed most of the parameters, the number of training parameters should be much smaller.
# +
total_params = sum(p.numel() for p in model.parameters())
trainable_total_params = sum(p.numel() for p in model.parameters() if p.requires_grad)
print('Total Parameters:', total_params, 'Trainable parameters: ', trainable_total_params)
# -
# ### Train the target model
# +
outModelName = 'finetuned'
logname = result_folder + model.__class__.__name__ + f'_{outModelName}.csv'
if not os.path.exists(logname):
with open(logname, 'w') as logfile:
logwriter = csv.writer(logfile, delimiter=',')
logwriter.writerow(['epoch', 'train loss', 'train acc', 'test loss', 'test acc'])
for epoch in range(start_epoch, max_epochs_target):
adjust_learning_rate(optimizer, epoch)
train_loss, train_acc = train(model, epoch, use_cuda=use_cuda)
test_loss, test_acc = test(model, epoch, outModelName, use_cuda=use_cuda)
with open(logname, 'a') as logfile:
logwriter = csv.writer(logfile, delimiter=',')
logwriter.writerow([epoch, train_loss, train_acc.item(), test_loss, test_acc.item()])
print(f'Epoch: {epoch} | train acc: {train_acc} | test acc: {test_acc}')
# -
# ## Plot results
# title plot results
results = pd.read_csv(f'/content/results/ResNet_{outModelName}.csv', sep =',')
results.head()
# +
train_accuracy = results['train acc'].values
test_accuracy = results['test acc'].values
print(f'Average Accuracy over {max_epochs_target} epochs:', sum(test_accuracy)//len(test_accuracy))
print(f'best accuraccy over {max_epochs_target} epochs:', max(test_accuracy))
# +
figureName = 'figure' # change figure name
plt.plot(results['epoch'].values, train_accuracy, label='train')
plt.plot(results['epoch'].values, test_accuracy, label='test')
plt.xlabel('Number of epochs')
plt.ylabel('Accuracy')
plt.title(f'Train/Test Accuracy curve for {max_epochs} epochs')
plt.savefig(f'/content/results/{figureName}.png')
plt.legend()
plt.show()
| projects/ComputerVision/transfer_learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/VindhyaHV/AppliedAI_Assignments/blob/main/5_Performance_metrics_Instructions.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="s0Ej_bXyQvnV"
# # Compute performance metrics for the given Y and Y_score without sklearn
# + id="4CHb6NE7Qvnc"
import numpy as np
import pandas as pd
# other than these two you should not import any other packages
import warnings
warnings.filterwarnings('ignore')
# + [markdown] id="KbsWXuDaQvnq"
# <pre>
# <font color='red'><b>A.</b></font> Compute performance metrics for the given data <strong>5_a.csv</strong>
# <b>Note 1:</b> in this data you can see number of positive points >> number of negatives points
# <b>Note 2:</b> use pandas or numpy to read the data from <b>5_a.csv</b>
# <b>Note 3:</b> you need to derive the class labels from given score</pre> $y^{pred}= \text{[0 if y_score < 0.5 else 1]}$
#
# <pre>
# <ol>
# <li> Compute Confusion Matrix </li>
# <li> Compute F1 Score </li>
# <li> Compute AUC Score, you need to compute different thresholds and for each threshold compute tpr,fpr and then use numpy.trapz(tpr_array, fpr_array) <a href='https://stackoverflow.com/q/53603376/4084039'>https://stackoverflow.com/q/53603376/4084039</a>, <a href='https://stackoverflow.com/a/39678975/4084039'>https://stackoverflow.com/a/39678975/4084039</a> Note: it should be numpy.trapz(tpr_array, fpr_array) not numpy.trapz(fpr_array, tpr_array)</li>
# <li> Compute Accuracy Score </li>
# </ol>
# </pre>
# + colab={"base_uri": "https://localhost:8080/"} id="H1K07ajpawOX" outputId="cd06e3bb-548c-4593-b0df-31b41ca3e6f9"
from google.colab import drive
drive.mount('/content/drive')
# + colab={"base_uri": "https://localhost:8080/"} id="6-52YX6mbgZZ" outputId="60b01659-6f50-4555-9075-6aff743f919a"
# !cd drive
# !ls
# + colab={"base_uri": "https://localhost:8080/"} id="R58WgCftYI_V" outputId="13027c9f-ce74-48a5-aaab-8f0b14d41c93"
a=pd.read_csv('drive/My Drive/5_a.csv')
a.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 363} id="KaCETD0yYGWa" outputId="70be0cb1-7809-4509-df3d-b501e648dacb"
a.head()
a.shape
decision = []
def decision_fun(input,threshold):
for i in input:
if i >= threshold:
decision.append(1.)
else:
decision.append(0.)
return decision
pred = decision_fun(a['proba'],0.8)
a['y_pred'] = pred
a.head(10)
# + colab={"base_uri": "https://localhost:8080/"} id="CUFEMK0CYFMB" outputId="ee58773a-929d-400c-e651-8934364dc02b"
tp = a.loc[a['y']==1.0][a['y_pred']==1.0]
tn = a.loc[a['y']==0.0][a['y_pred']==0.0]
fp = a.loc[a['y']==0.0][a['y_pred']==1.0]
fn = a.loc[a['y']==1.0][a['y_pred']==0.0]
tp = len(tp)
tn = len(tn)
fp = len(fp)
fn = len(fn)
print('tp',tp,'tn',tn,'fp',fp,'fn',fn)
confusion_matrix = [[tp , fp],[fn , tn]]
confusion_matrix
# + colab={"base_uri": "https://localhost:8080/"} id="AnJQBbU-zcxb" outputId="e86fdcfb-e899-48e8-d2bf-d797067b88c9"
precision = tp/(tp+fp)
print('precision',precision)
recall = tp/(tp+fn)
print('recall',recall)
f1_score = 2*(recall*precision)/(recall+precision)
print(f1_score)
print(a.head())
# + colab={"base_uri": "https://localhost:8080/", "height": 131} id="bso7Vwhc7ZIo" outputId="4e0cdcad-776e-4fe7-ca73-f9c6336f2<PASSWORD>"
#https://www.drjamesfroggatt.com/python-and-neural-networks/iopub-data-rate-exceeded-the-notebook-server-will-temporarily-stop-sending-output-to-the-client-in-order-to-avoid-crashing-it/
# + id="WaFLW7oBQvnt" colab={"base_uri": "https://localhost:8080/"} outputId="027d541b-646a-4ca9-eafd-42c7ddfc53fd"
#Compute AUC Score, you need to compute different thresholds and for each threshold compute tpr,fpr and then use numpy.trapz(tpr_array, fpr_array) https://stackoverflow.com/q/53603376/4084039,
#https://stackoverflow.com/a/39678975/4084039 Note: it should be numpy.trapz(tpr_array, fpr_array) not numpy.trapz(fpr_array, tpr_array)
thresholds = len(a)
print(thresholds)
interval = 1/10
prob_thresholds = np.linspace(0.5,1,2)
print(prob_thresholds)
for i in prob_thresholds:
x = decision_fun(a['proba'],i)
print(x)
# + [markdown] id="V5KZem1BQvn2"
# <pre>
# <font color='red'><b>B.</b></font> Compute performance metrics for the given data <strong>5_b.csv</strong>
# <b>Note 1:</b> in this data you can see number of positive points << number of negatives points
# <b>Note 2:</b> use pandas or numpy to read the data from <b>5_b.csv</b>
# <b>Note 3:</b> you need to derive the class labels from given score</pre> $y^{pred}= \text{[0 if y_score < 0.5 else 1]}$
#
# <pre>
# <ol>
# <li> Compute Confusion Matrix </li>
# <li> Compute F1 Score </li>
# <li> Compute AUC Score, you need to compute different thresholds and for each threshold compute tpr,fpr and then use numpy.trapz(tpr_array, fpr_array) <a href='https://stackoverflow.com/q/53603376/4084039'>https://stackoverflow.com/q/53603376/4084039</a>, <a href='https://stackoverflow.com/a/39678975/4084039'>https://stackoverflow.com/a/39678975/4084039</a></li>
# <li> Compute Accuracy Score </li>
# </ol>
# </pre>
# + id="U2sKlq0YQvn5"
# write your code
# + [markdown] id="GiPGonTzQvoB"
# <font color='red'><b>C.</b></font> Compute the best threshold (similarly to ROC curve computation) of probability which gives lowest values of metric <b>A</b> for the given data <strong>5_c.csv</strong>
# <br>
#
# you will be predicting label of a data points like this: $y^{pred}= \text{[0 if y_score < threshold else 1]}$
#
# $ A = 500 \times \text{number of false negative} + 100 \times \text{numebr of false positive}$
#
# <pre>
# <b>Note 1:</b> in this data you can see number of negative points > number of positive points
# <b>Note 2:</b> use pandas or numpy to read the data from <b>5_c.csv</b>
# </pre>
# + id="x5HIJzq1QvoE"
# write your code
# + [markdown] id="sD4CcgjXQvoL"
# <pre>
# <font color='red'><b>D.</b></font> Compute performance metrics(for regression) for the given data <strong>5_d.csv</strong>
# <b>Note 2:</b> use pandas or numpy to read the data from <b>5_d.csv</b>
# <b>Note 1:</b> <b>5_d.csv</b> will having two columns Y and predicted_Y both are real valued features
# <ol>
# <li> Compute Mean Square Error </li>
# <li> Compute MAPE: https://www.youtube.com/watch?v=ly6ztgIkUxk</li>
# <li> Compute R^2 error: https://en.wikipedia.org/wiki/Coefficient_of_determination#Definitions </li>
# </ol>
# </pre>
| 5_Performance_metrics_Instructions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
# get ANOVA table as R like output
import statsmodels.api as sm
from statsmodels.formula.api import ols
# post-hoc tests:
from bioinfokit.analys import stat
from scipy.stats import ttest_rel
import math
# helper function
# function to calculate Cohen's d for independent samples
def cohend(d1, d2):
# calculate the size of samples
n1, n2 = len(d1), len(d2)
# calculate the variance of the samples
s1, s2 = np.var(d1, ddof=1), np.var(d2, ddof=1)
# calculate the pooled standard deviation
s = math.sqrt(((n1 - 1) * s1 + (n2 - 1) * s2) / (n1 + n2 - 2))
# calculate the means of the samples
u1, u2 = np.mean(d1), np.mean(d2)
# calculate the effect size
return (u1 - u2) / s
# -
# ## 1. Loading the data
#
# Data is available from the EDMOND repository [Modulation of behavior in zebrafish by the neuropeptide PTH2](https://dx.doi.org/10.17617/3.6v). Individual data files should be placed within the same folder as the scripts, otherwise, file locations need to be updated in the cell below.
#
# The _social preference_ dataframe contains 5 columns:
# - **experiment** either _tripartite_ or _uchamber_, indicating whether the data was obtained in the open field or the forced-choice paradigm.
# - **genotype**: either _wt_ (for $pth2^{+/+}$) or _ko_ (fo $pth2^{-/-}$).
# - **rearing** either _social_ (rearing in groups of defined size as of 3 days post fertilization) or _isolated_ (rearing alone as of 3 dpf).
# - **age**: either 21 or 56 dpf.
# - **value**: number between -1 and 1, indicating the social preference index.
df = pd.read_excel(r'.\SocialPreference.xlsx' , sheet_name = 'import')
df.head()
# ## Figure 3 b - d and f - h
# +
g = sns.FacetGrid(data = df[df['rearing']=='social'], col = 'age' , row = 'experiment')
g.map_dataframe(sns.boxplot , x = 'condition', y = 'value' , hue = 'genotype' , showfliers = 0, palette = ['w', 'g'])
g.map_dataframe(sns.stripplot , x = 'condition' , y = 'value' , hue = 'genotype' , size = 7, dodge = True , palette = ['k', 'g'])
g.set_axis_labels("condition", "social preference index")
g.set_titles(col_template="{col_name} dpf", row_template="{row_name}")
g.set(ylim=(-1.1, 1.1))
f = sns.FacetGrid(data = df[df['rearing']=='isolated'], row = 'experiment')
f.map_dataframe(sns.boxplot , x = 'condition', y = 'value' , hue = 'genotype' , showfliers = 0, palette = ['w'])
f.map_dataframe(sns.stripplot , x = 'condition' , y = 'value' , hue = 'genotype' , size = 7, dodge = True)
f.set(ylim=(-1.1, 1.1))
# +
# Statistical Evaluation
df_uchamber = df[df['experiment'] == 'uchamber']
df_uchamber_21 = df_uchamber[df_uchamber['age'] == 21]
df_uchamber_56 = df_uchamber[df_uchamber['age'] == 56]
df_uchamber_56_social = df_uchamber_56[df_uchamber_56['rearing'] == "social"]
df_uchamber_56_iso = df_uchamber_56[df_uchamber_56['rearing'] == "isolated"]
df_tripartite = df[df['experiment'] == 'tripartite']
df_tripartite_21 = df_tripartite[df_tripartite['age'] == 21]
df_tripartite_56 = df_tripartite[df_tripartite['age'] == 56]
df_tripartite_56_social = df_tripartite_56[df_tripartite_56['rearing'] == "social"]
df_tripartite_56_iso = df_tripartite_56[df_tripartite_56['rearing'] == "isolated"]
# for 21 dpf, u-chamber paradigm:
# Ordinary Least Squares (OLS) model
model = ols('value ~ genotype * condition', data=df_uchamber_21).fit()
anova_table = sm.stats.anova_lm(model, typ=2)
anova_table
#np.sum(df_uchamber_21["genotype"] == "ko")/2
# -
# perform multiple pairwise comparison (Tukey's HSD)
# for unequal sample size data, tukey_hsd uses Tukey-Kramer test
res = stat()
res.tukey_hsd(df=df_uchamber_21, res_var='value', xfac_var=['genotype', 'condition'], anova_model='value ~ genotype * condition')
res.tukey_summary
# for 21 dpf, open-field paradigm:
# Ordinary Least Squares (OLS) model
model = ols('value ~ genotype * condition', data=df_tripartite_21).fit()
anova_table = sm.stats.anova_lm(model, typ=2)
anova_table
# perform multiple pairwise comparison (Tukey's HSD)
# for unequal sample size data, tukey_hsd uses Tukey-Kramer test
res = stat()
res.tukey_hsd(df=df_tripartite_21, res_var='value', xfac_var=['genotype', 'condition'], anova_model='value ~ genotype * condition')
res.tukey_summary
# for 56 dpf, socially reared, u-chamber paradigm:
# Ordinary Least Squares (OLS) model
model = ols('value ~ genotype * condition', data=df_uchamber_56_social).fit()
anova_table = sm.stats.anova_lm(model, typ=2)
anova_table
# +
# perform multiple pairwise comparison (Tukey's HSD)
# for unequal sample size data, tukey_hsd uses Tukey-Kramer test
res = stat()
res.tukey_hsd(df=df_uchamber_56_social, res_var='value', xfac_var=['genotype', 'condition'], anova_model='value ~ genotype * condition')
res.tukey_summary
# +
# for 56 dpf, reared in isolation, forced-choice paradigm:
ttest_rel(df_uchamber_56_iso[df_uchamber_56_iso['condition'] == 'habituation']['value'] ,
df_uchamber_56_iso[df_uchamber_56_iso['condition'] == 'test']['value'])
cohend(df_uchamber_56_iso[df_uchamber_56_iso['condition'] == 'habituation']['value'] ,
df_uchamber_56_iso[df_uchamber_56_iso['condition'] == 'test']['value'])
np.mean(df_uchamber_56_iso[df_uchamber_56_iso['condition'] == 'test']['value'])
# +
# for 56 dpf, open-field paradigm:
# Ordinary Least Squares (OLS) model
model = ols('value ~ genotype * condition', data=df_tripartite_56_social).fit()
anova_table = sm.stats.anova_lm(model, typ=2)
anova_table
wt_56 = df_tripartite_56_social[df_tripartite_56_social["genotype"] == "wt"]
ko_56 = df_tripartite_56_social[df_tripartite_56_social["genotype"] == "ko"]
cohend(ko_56[ko_56["condition"]=="habituation"]["value"],
ko_56[ko_56["condition"]=="test"]["value"])
# -
# perform multiple pairwise comparison (Tukey's HSD)
# for unequal sample size data, tukey_hsd uses Tukey-Kramer test
res = stat()
res.tukey_hsd(df=df_tripartite_56_social, res_var='value', xfac_var=['genotype', 'condition'], anova_model='value ~ genotype * condition')
res.tukey_summary
# for 56 dpf, reared in isolation, open-field paradigm:
ttest_rel(df_tripartite_56_iso[df_tripartite_56_iso['condition'] == 'habituation']['value'] ,
df_tripartite_56_iso[df_tripartite_56_iso['condition'] == 'test']['value'])
# +
# Effect sizes and central tendencies
# Tripartite chamber, 21 dpf: 0.12
print('Tripartite, 21 dpf, effect size (Cohen) for wildtype: ')
print(cohend(df_tripartite_21[df_tripartite_21['condition'] == 'habituation'][df_tripartite_21[df_tripartite_21['condition'] == 'habituation']['genotype']=='wt']['value'],
df_tripartite_21[df_tripartite_21['condition'] == 'test'][df_tripartite_21[df_tripartite_21['condition'] == 'test']['genotype']=='wt']['value']))
print('Tripartite, 21 dpf, effect size (Cohen) for mutants: ')
print(cohend(df_tripartite_21[df_tripartite_21['condition'] == 'habituation'][df_tripartite_21[df_tripartite_21['condition'] == 'habituation']['genotype']=='ko']['value'],
df_tripartite_21[df_tripartite_21['condition'] == 'test'][df_tripartite_21[df_tripartite_21['condition'] == 'test']['genotype']=='ko']['value']))
print('\n')
print('median of wildtype in this experiment, habituation: ')
print(np.median(df_tripartite_21[df_tripartite_21['condition'] == 'habituation'][df_tripartite_21[df_tripartite_21['condition'] == 'habituation']['genotype']=='wt']['value']))
print('median of wildtype in this experiment, test: ')
print(np.median(df_tripartite_21[df_tripartite_21['condition'] == 'test'][df_tripartite_21[df_tripartite_21['condition'] == 'test']['genotype']=='wt']['value']))
print('\n')
print('median of mutants in this experiment, habituation: ')
print(np.median(df_tripartite_21[df_tripartite_21['condition'] == 'habituation'][df_tripartite_21[df_tripartite_21['condition'] == 'habituation']['genotype']=='ko']['value']))
print('median of mutants in this experiment, test: ')
print(np.median(df_tripartite_21[df_tripartite_21['condition'] == 'test'][df_tripartite_21[df_tripartite_21['condition'] == 'test']['genotype']=='ko']['value']))
print('\n')
print('----------------------')
# U-chamber, 21 dpf: 0.49
print('U-chamber, 21 dpf, effect size (Cohen) for wildtype: ')
print(cohend(df_uchamber_21[df_uchamber_21['condition'] == 'habituation'][df_uchamber_21[df_uchamber_21['condition'] == 'habituation']['genotype']=='wt']['value'],
df_uchamber_21[df_uchamber_21['condition'] == 'test'][df_uchamber_21[df_uchamber_21['condition'] == 'test']['genotype']=='wt']['value']))
print('U-chamber, 21 dpf, effect size (Cohen) for mutants: ')
print(cohend(df_uchamber_21[df_uchamber_21['condition'] == 'habituation'][df_uchamber_21[df_uchamber_21['condition'] == 'habituation']['genotype']=='ko']['value'],
df_uchamber_21[df_uchamber_21['condition'] == 'test'][df_uchamber_21[df_uchamber_21['condition'] == 'test']['genotype']=='ko']['value']))
print('\n')
print('median of wildtype in this experiment, habituation: ')
print(np.median(df_uchamber_21[df_uchamber_21['condition'] == 'habituation'][df_uchamber_21[df_uchamber_21['condition'] == 'habituation']['genotype']=='wt']['value']))
print('median of wildtype in this experiment, test: ')
print(np.median(df_uchamber_21[df_uchamber_21['condition'] == 'test'][df_uchamber_21[df_uchamber_21['condition'] == 'test']['genotype']=='wt']['value']))
print('\n')
print('median of mutant in this experiment, habituation: ')
print(np.median(df_uchamber_21[df_uchamber_21['condition'] == 'habituation'][df_uchamber_21[df_uchamber_21['condition'] == 'habituation']['genotype']=='ko']['value']))
print('median of mutant in this experiment, test: ')
print(np.median(df_uchamber_21[df_uchamber_21['condition'] == 'test'][df_uchamber_21[df_uchamber_21['condition'] == 'test']['genotype']=='ko']['value']))
print('\n')
print('----------------------')
# Tripartite chamber, 56 dpf: 0.23
print('Tripartite, 56 dpf, effect size (Cohen) for wildtype: ')
print(cohend(df_tripartite_56[df_tripartite_56['condition'] == 'habituation'][df_tripartite_56[df_tripartite_56['condition'] == 'habituation']['genotype']=='wt']['value'],
df_tripartite_56[df_tripartite_56['condition'] == 'test'][df_tripartite_56[df_tripartite_56['condition'] == 'test']['genotype']=='wt']['value']))
print('Tripartite, 56 dpf, effect size (Cohen) for mutants: ')
print(cohend(df_tripartite_56[df_tripartite_56['condition'] == 'habituation'][df_tripartite_56[df_tripartite_56['condition'] == 'habituation']['genotype']=='ko']['value'],
df_tripartite_56[df_tripartite_56['condition'] == 'test'][df_tripartite_56[df_tripartite_56['condition'] == 'test']['genotype']=='ko']['value']))
print('\n')
print('median of wildtype in this experiment, habituation: ')
print(np.median(df_tripartite_56[df_tripartite_56['condition'] == 'habituation'][df_tripartite_56[df_tripartite_56['condition'] == 'habituation']['genotype']=='wt']['value']))
print('median of wildtype in this experiment, test: ')
print(np.median(df_tripartite_56[df_tripartite_56['condition'] == 'test'][df_tripartite_56[df_tripartite_56['condition'] == 'test']['genotype']=='wt']['value']))
print('\n')
print('median of mutant in this experiment, habituation: ')
print(np.median(df_tripartite_56[df_tripartite_56['condition'] == 'habituation'][df_tripartite_56[df_tripartite_56['condition'] == 'habituation']['genotype']=='ko']['value']))
print('median of mutant in this experiment, test: ')
print(np.median(df_tripartite_56[df_tripartite_56['condition'] == 'test'][df_tripartite_56[df_tripartite_56['condition'] == 'test']['genotype']=='ko']['value']))
print('\n')
print('----------------------')
# U-chamber, 56 dpf: 0.57
print('U-chamber, 56 dpf, effect size (Cohen), wildtype: ')
print(cohend(df_uchamber_56[df_uchamber_56['condition'] == 'habituation'][df_uchamber_56[df_uchamber_56['condition'] == 'habituation']['genotype']=='wt']['value'],
df_uchamber_56[df_uchamber_56['condition'] == 'test'][df_uchamber_56[df_uchamber_56['condition'] == 'test']['genotype']=='wt']['value']))
print('U-chamber, 56 dpf, effect size (Cohen), mutant: ')
print(cohend(df_uchamber_56[df_uchamber_56['condition'] == 'habituation'][df_uchamber_56[df_uchamber_56['condition'] == 'habituation']['genotype']=='ko']['value'],
df_uchamber_56[df_uchamber_56['condition'] == 'test'][df_uchamber_56[df_uchamber_56['condition'] == 'test']['genotype']=='ko']['value']))
print('\n')
print('median of wildtype in this experiment, habituation: ')
print(np.median(df_uchamber_56[df_uchamber_56['condition'] == 'habituation'][df_uchamber_56[df_uchamber_56['condition'] == 'habituation']['genotype']=='wt']['value']))
print('median of wildtype in this experiment, test: ')
print(np.median(df_uchamber_56[df_uchamber_56['condition'] == 'test'][df_uchamber_56[df_uchamber_56['condition'] == 'test']['genotype']=='wt']['value']))
print('\n')
print('median of mutant in this experiment, habituation: ')
print(np.median(df_uchamber_56[df_uchamber_56['condition'] == 'habituation'][df_uchamber_56[df_uchamber_56['condition'] == 'habituation']['genotype']=='ko']['value']))
print('median of mutant in this experiment, test: ')
print(np.median(df_uchamber_56[df_uchamber_56['condition'] == 'test'][df_uchamber_56[df_uchamber_56['condition'] == 'test']['genotype']=='ko']['value']))
print('\n')
print('----------------------')
# Tripartite chamber, 56 dpf, isolated: 0.75
print('Tripartite, 56 dpf, isolated wildtype, effect size (Cohen): ')
print(cohend(df_tripartite_56_iso[df_tripartite_56_iso['condition'] == 'habituation']['value'],
df_tripartite_56_iso[df_tripartite_56_iso['condition'] == 'test']['value']))
# U-chamber, 56 dpf, isolated: 0.18
print('U-chamber, 56 dpf, isolated wildtype, effect size (Cohen): ')
print(cohend(df_uchamber_56_iso[df_uchamber_56_iso['condition'] == 'habituation']['value'],
df_uchamber_56_iso[df_uchamber_56_iso['condition'] == 'test']['value']))
print('\n')
print('median of habituation phase in Tripartite chamber: ')
print(np.median(df_tripartite_56_iso[df_tripartite_56_iso['condition'] == 'habituation']['value']))
print('median of test phase in Tripartite chamber: ')
print(np.median(df_tripartite_56_iso[df_tripartite_56_iso['condition'] == 'test']['value']))
print('\n')
print('median of habituation phase in U-chamber chamber: ')
print(np.median(df_uchamber_56_iso[df_uchamber_56_iso['condition'] == 'habituation']['value']))
print('median of test phase in U-chamber chamber: ')
print(np.median(df_uchamber_56_iso[df_uchamber_56_iso['condition'] == 'test']['value']))
# -
df_tripartite_56_iso
| Figure3_SocialPreference.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Question 1:
# + active=""
# Create a function to perform basic arithmetic operations that includes
# addition, subtraction, multiplication and division on a string number
# (e.g. "12 + 24" or "23 - 21" or "12 // 12" or "12 * 21").
# Here, we have 1 followed by a space, operator followed by
# another space
# and 2. For the challenge, we are going to have only two numbers between 1 valid operator.
# The return value should be a number. eval() is not allowed.
# In case of division, whenever the second number equals "0" return -1.
#
# For example:
#
# "15 // 0" -1
#
# Examples
#
# arithmetic_operation("12 + 12") 24 // 12 + 12 = 24
# arithmetic_operation("12 - 12") 24 // 12 - 12 = 0
# arithmetic_operation("12 * 12") 144 // 12 * 12 = 144
# arithmetic_operation("12 // 0") -1 // 12 / 0 = -1.
# -
# Answer :
# +
def arithmetic_operation(expr):
lst = expr.split(" ")
if lst[1]=="+":
return int(lst[0])+int(lst[2])
elif lst[1]=="-":
return int(lst[0])-int(lst[2])
elif lst[1]=="*":
return int(lst[0])*int(lst[2])
else:
if int(lst[2]) == 0:
return -1
else:
return int(lst[0]//lst[2])
print(arithmetic_operation("12 + 12"))
print(arithmetic_operation("12 - 12"))
print(arithmetic_operation("12 * 12"))
print(arithmetic_operation("12 // 0"))
# -
# Question 2:
# + active=""
# Write a function that takes the coordinates of three points in the form of a 2d array and returns
# the perimeter of the triangle. The given points are the vertices of a triangle on a two-dimensional plane.
#
# Examples
#
# perimeter( [ [15, 7], [5, 22], [11, 1] ] ) 47.08
# perimeter( [ [0, 0], [0, 1], [1, 0] ] ) 3.42
# perimeter( [ [-10, -10], [10, 10 ], [-10, 10] ] ) 68.28
# +
def distance_2d(a,b):
return ((a[0]-b[0])**2 + (a[1]-b[1])**2)**(1/2)
def perimeter(coord):
side1 = distance_2d(coord[0],coord[1])
side2 = distance_2d(coord[1],coord[2])
side3 = distance_2d(coord[2],coord[0])
return round(side1+side2+side3,2)
print(perimeter([ [15, 7], [5, 22], [11, 1] ]))
print(perimeter([ [0, 0], [0, 1], [1, 0] ]))
print(perimeter([ [-10, -10], [10, 10 ], [-10, 10] ]))
# -
# Question 3:
# + active=""
# A city skyline can be represented as a 2-D list with 1s representing
# buildings. In the example below, the height of the tallest building is 4 (secondmost right column).
#
# [[0, 0, 0, 0, 0, 0],
# [0, 0, 0, 0, 1, 0],
# [0, 0, 1, 0, 1, 0],
# [0, 1, 1, 1, 1, 0],
# [1, 1, 1, 1, 1, 1]]
#
# Create a function that takes a skyline (2-D list of 0's and 1's) and returns
# the height of the tallest skyscraper.
#
# Examples
#
# tallest_skyscraper([
# [0, 0, 0, 0],
# [0, 1, 0, 0],
# [0, 1, 1, 0],
# [1, 1, 1, 1]
# ]) 3
# tallest_skyscraper([
# [0, 1, 0, 0],
# [0, 1, 0, 0],
# [0, 1, 1, 0],
# [1, 1, 1, 1]
# ]) 4
# tallest_skyscraper([
# [0, 0, 0, 0],
# [0, 0, 0, 0],
# [1, 1, 1, 0],
# [1, 1, 1, 1]
# ]) 2
# -
# Answer :
# +
def tallest_skyscraper(lst):
columns = list(zip(*lst))
sums = list(map(sum , columns))
return max(sums)
print(tallest_skyscraper([[0, 0, 0, 0, 0, 0],
[0, 0, 0, 0, 1, 0],
[0, 0, 1, 0, 1, 0],
[0, 1, 1, 1, 1, 0],
[1, 1, 1, 1, 1, 1]]))
print(tallest_skyscraper([
[0, 0, 0, 0],
[0, 1, 0, 0],
[0, 1, 1, 0],
[1, 1, 1, 1]
]))
print(tallest_skyscraper([
[0, 1, 0, 0],
[0, 1, 0, 0],
[0, 1, 1, 0],
[1, 1, 1, 1]
]))
print(tallest_skyscraper([
[0, 0, 0, 0],
[0, 0, 0, 0],
[1, 1, 1, 0],
[1, 1, 1, 1]
]))
# -
# Question 4:
# + active=""
# A financial institution provides professional services to banks and claims charges from
# the customers based on the number of man-days provided.
# Internally, it has set a scheme to motivate and reward staff to meet and
# exceed targeted billable utilization and revenues by paying a bonus for
# each day claimed from customers in excess of a threshold target.
# This quarterly scheme is calculated with a threshold target of 32 days per
# quarter, and the incentive payment for each billable day in excess of such
# threshold target is shown as follows:
#
# Days Bonus
# 0 to 32 days Zero
# 33 to 40 days SGD$325 per billable day
# 41 to 48 days SGD$550 per billable day
#
# Greater than 48 days SGD$600 per billable day
# Please note that incentive payment is calculated progressively. As an
# example, if an employee reached total billable days of 45 in a quarter,
# his/her incentive payment is computed as follows:
#
# 32*0 + 8*325 + 5*550 = 5350
#
# Write a function to read the billable days of an employee and return the bonus
# he/she has obtained in that quarter.
#
# Examples
#
# bonus(15) 0
# bonus(37) 1625
# bonus(50) 8200
# -
# Answer :
# +
def bonus(days):
tot_bonus = 0
if 0<=days<=32:
tot_bonus += 0
elif 33<=days<=40:
tot_bonus += (days-32)*325
elif 41<=days<=48:
tot_bonus += 0 + 8*325 + (days-40)*550
else:
tot_bonus += 0 + 8*325 + 8*550 + (days-48)*600
return tot_bonus
print(bonus(15))
print(bonus(37))
print(bonus(50))
# -
# Question 5:
# + active=""
# A number is said to be Disarium if the sum of its digits raised to their
# respective positions is the number itself.
# Create a function that determines whether a number is a Disarium or not.
#
# Examples
#
# is_disarium(75) False
# # 7^1 + 5^2 = 7 + 25 = 32
# is_disarium(135) True
# # 1^1 + 3^2 + 5^3 = 1 + 9 + 125 = 135
# is_disarium(544) False
# is_disarium(518) True
# is_disarium(466) False
# is_disarium(8) True
# -
# Answer :
# +
def is_disarium(num):
st = str(num)
sum_ = sum([int(st[i-1])**(i) for i in range(1,len(st)+1)])
if sum_ == num:
return True
else:
return False
print(is_disarium(75))
print(is_disarium(135))
print(is_disarium(544))
print(is_disarium(518))
print(is_disarium(466))
print(is_disarium(8))
# -
| Python Advance Programming Assignment/Assignment_03.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### This notebook is to create the preparation information and sample information files for Qiita, as well as the mapping file for Qiime.
import numpy as np
import pandas as pd
# ## create preparation_info document
info = pd.read_csv('../data/Demultiplex_Sheet.txt', sep='\t')
info = info.sort_values(['#SampleID'])
info.head()
info = info.drop_duplicates(subset=['#SampleID', 'ReversePrimer'])
info.head()
info.shape[0]
n = info.shape[0]
data = {'#SampleID': info.loc[:,'#SampleID'].str[7:13].values, # .values takes only value not index
'Barcode': info.loc[:,'BarcodeSequence'].values,
'LinkerPrimerSequence': info.loc[:, 'LinkerPrimerSequence'].values,
'Description': info.loc[:,'Description'].values,
'Experiment_Design_Description': np.repeat(
'16S stool samples sequenced for MrOS Vitamin D study', n),
'Library_Construction_Protocol': np.repeat('16S rRNA v4', n),
'Linker': np.repeat('GT', n),
'Platform': np.repeat('Illumina', n),
'Center_Name': info.loc[:,'#SampleID'].str[14:16].values,
'Center_Project': np.repeat('MrOS', n),
'Instrument_Model': np.repeat('Illumina MiSeq', n)}
prep_info = pd.DataFrame(data, columns=['#SampleID', 'Barcode', 'LinkerPrimerSequence',
'Description',
'Experiment_Design_Description',
'Library_Construction_Protocol', 'Linker', 'Platform',
'Center_Name', 'Center_Project', 'Instrument_Model'])
prep_info.shape
prep_info.head(10)
# fix mismatch sample name 'BI0778' should be 'BIO778' (Nora's email on08/07/2017)
prep_info = prep_info.replace(to_replace='BIO778', value='BI0778')
# ## Create sample_info document
# +
# note AMPHIT_15SD was missing in Nov6th metadata, thus copied from file Variable_sleepV4updated.csv
# -
samples = pd.read_csv('../data/SleepMetadataV4_Nov6th2017.csv', sep=',')
samples.shape
# +
# error in categories of SLEEPHRS (additional space in second '2')
samples.SLEEPHRS.value_counts()
# correct
samples.loc[samples.SLEEPHRS == '2: 7 HRS ', 'SLEEPHRS'] = '2: 7 HRS'
# check
samples.SLEEPHRS.value_counts()
# -
samples.TUDRAMT.value_counts() # alcohol drinking variable is wrong
# replace alcohol drinking variable
tmp = pd.read_csv('../data/mapping_vitd_v4.txt', sep='\t')
tmp = tmp[['ID', 'TUDRAMT']]
tmp.rename(columns={'TUDRAMT':'TUDRAMT_REVISED'}, inplace=True)
tmp.shape
tmp.TUDRAMT_REVISED.value_counts()
tmp.head(3)
samples.head(3)
samples = pd.merge(samples, tmp, left_on='ID', right_on='ID')
samples.columns
samples.shape
len(pd.unique(samples.ID))
# reference: http://www.latlong.net/
Latitude = []
Longitude = []
sites = samples.loc[:, 'SITE']
for i in range(samples.shape[0]):
if sites[i] == 'Birmingham':
latitude = '33.520661'
longitude = '-86.80249'
elif sites[i] == 'San Diego':
latitude = '32.715738'
longitude = '-117.1611'
elif sites[i] == 'Pittsburgh':
latitude = '40.440625'
longitude = '-79.99589'
elif sites[i] == 'Palo Alto':
latitude = '37.441883'
longitude = '-122.143'
elif sites[i] == 'Portland':
latitude = '45.523062'
longitude = '-122.6765'
elif sites[i] == 'Minneapolis':
latitude = '44.977753'
longitude = '-93.26501'
Latitude.append(latitude)
Longitude.append(longitude)
# simple check
print(samples['SITE'][[1,90,200,300,400, 500]])
print(np.array(Latitude)[[1,90,200,300,400, 500]])
print(np.array(Longitude)[[1,90,200,300,400, 500]])
m = samples.shape[0]
required = {'#SampleID': samples.loc[:,'ID'].values,
'Title': np.repeat('MrOS_VitaminD', m),
'Anonymized_Name': samples.loc[:,'ID'].values,
'Scientific_Name': np.repeat('human gut metagenome', m),
'Taxon_ID': np.repeat('Not applicable', m),
#'Description': np.repeat('Not applicable', m),
'Sample_Type': np.repeat('stool', m),
'Geo_Loc_Name': samples.loc[:, 'SITE'].values,
'Elevation': np.repeat('Not applicable', m),
'Env_Biome': np.repeat('urban biome', m),
'Env_Feature': np.repeat('human-associated habitat', m),
'Env_Material': np.repeat('feces', m),
'Env_Package': np.repeat('human-gut', m),
'Latitude': Latitude,
'Longitude': Longitude,
'Collection_Timestamp': np.repeat('Not applicable', m),
'DNA_Extracted': np.repeat('Not applicable', m),
'Physical_Specimen_Location': np.repeat('Not applicable', m),
'Physical_Specimen_Remaining': np.repeat('Not applicable', m),
'Age': samples.loc[:,'V4AGE1'].values,
'Age_Units': np.repeat('years', m),
'Host_Subject_ID': np.repeat('Not applicable', m),
'Host_Taxid': np.repeat('Not applicable', m),
'Host_Scientific_Name': np.repeat('Homo sapiens', m),
'Host_Common_Name': np.repeat('human', m),
'Life_Stage': np.repeat('adult', m),
'Sex': np.repeat('male', m),
'Height': samples.loc[:, 'HWHGT'].values,
'Height_Units': np.repeat('cm', m),
'Weight': samples.loc[:, 'HWWGT'].values,
'Weight_Units': np.repeat('kg', m),
'BMI': samples.loc[:, 'HWBMI'].values,
'Body_Habitat': np.repeat('UBERON:feces', m),
'Body_Site': np.repeat('UBERON:feces', m),
'Body_Product': np.repeat('UBERON:feces', m)}
len(required)
sample_info = pd.concat([pd.DataFrame(required), samples], axis=1)
sample_info = pd.DataFrame(sample_info, columns = ['#SampleID', 'Title', 'Anonymized_Name',
'Scientific_Name', 'Taxon_ID',
#'Description',
'Sample_Type', 'Geo_Loc_Name', 'Elevation',
'Env_Biome', 'Env_Feature', 'Env_Material', 'Env_Package', 'Latitude', 'Longitude',
'Collection_Timestamp', 'DNA_Extracted', 'Physical_Specimen_Location', 'Physical_Specimen_Remaining',
'Age', 'Age_Units', 'Host_Subject_ID', 'Host_Taxid', 'Host_Scientific_Name', 'Host_Common_Name',
'Life_Stage', 'Sex', 'Height', 'Height_Units', 'Weight', 'Weight_Units', 'BMI', 'Body_Habitat',
'Body_Site', 'Body_Product', 'GIERACE', 'SITE', 'AMAMPT', 'AMPHIT', 'AMFVT', 'MIDATA', 'V4DATE',
'GIMSTAT', 'MHDIAB', 'MHRHEU1', 'MHOA', 'MHCHF', 'MHMI', 'MHDEPR', 'MHDEPRT', 'TUDRAMT_REVISED', 'PQPSLMED',
'PQPSQUAL', 'PASCORE', 'QLCOMP', 'SLSLPHRS', 'SLEEPHRS', 'PQPEFFCY', 'PQPEFFIC', 'PQPSQI', 'PQBADSLP', 'TURSMOKE',
'M1ADEPR', 'M1BENZO', 'AMAMPT_C1', 'AMFVT_C1', 'AMPHIT_15SD'])
sample_info.shape
sample_info.head(10)
# ## Match sample_name in two documents
id_prep=list(prep_info.loc[:,'#SampleID'].values)
id_sample=list(sample_info.loc[:,'#SampleID'].values)
# samples with metadata yet no sequencing data
for i in id_sample:
if i not in id_prep:
print(i)
# samples with sequencing data yet no metadata
no_sample_info = []
for j in id_prep:
if j not in id_sample:
no_sample_info.append(j)
print(len(no_sample_info))
print(no_sample_info)
# ## Drop un-needed samples
# exclude PO7100, as it has no microbiome data (see <NAME>'s email on 08/07/2017)
prep_info = prep_info.loc[~prep_info['#SampleID'].isin(['PO7100'])]
sample_info = sample_info.loc[~sample_info['#SampleID'].isin(['PO7100'])]
print(prep_info.shape)
print(sample_info.shape)
# ## create Qiita prep and Qiime mapping files
prep_info.head(5)
mapping = pd.merge(prep_info, sample_info, on='#SampleID')
print(prep_info.shape)
print(sample_info.shape)
print(mapping.shape)
#mapping.rename(columns={'Sample_Name': '#SampleID'}, inplace=True)
mapping.rename(columns={'Barcode': 'BarcodeSequence'}, inplace=True)
mapping.shape
mapping.head(5)
# move 'Description' to the end
cols = list(mapping)
cols.insert(mapping.shape[1], cols.pop(cols.index('Description')))
mapping = mapping.loc[:, cols]
mapping.head(5)
mapping.columns
# convert to proper variable types (categorical or continuous)
vars_cat = np.array(['BarcodeSequence', 'LinkerPrimerSequence', 'Experiment_Design_Description',
'Library_Construction_Protocol', 'Linker', 'Platform', 'Center_Name', 'Center_Project', 'Instrument_Model',
'Title', 'Anonymized_Name', 'Scientific_Name', 'Taxon_ID', 'Sample_Type', 'Geo_Loc_Name', 'Elevation', 'Env_Biome',
'Env_Feature', 'Env_Material', 'Env_Package', 'Collection_Timestamp', 'DNA_Extracted', 'Physical_Specimen_Location',
'Physical_Specimen_Remaining', 'Age_Units', 'Host_Subject_ID', 'Host_Taxid','Host_Scientific_Name', 'Host_Common_Name',
'Life_Stage', 'Sex', 'Height_Units', 'Weight_Units', 'Body_Habitat', 'Body_Site', 'Body_Product', 'GIERACE', 'SITE',
'SLEEPHRS', 'MIDATA', 'AMPHIT_15SD', 'MHDIAB', 'MHRHEU1', 'MHOA', 'MHCHF', 'MHMI', 'MHDEPR', 'GIMSTAT',
'MHDEPRT', 'TUDRAMT_REVISED', 'PQPSQUAL', 'QLCOMP', 'PQBADSLP', 'TURSMOKE', 'M1ADEPR', 'M1BENZO', 'Description',
'AMAMPT_C1', 'AMFVT_C1', 'V4DATE', 'PQPSLMED', 'PQPEFFIC'])
vars_cts = np.array(['Latitude', 'Longitude', 'Age', 'Height', 'Weight', 'BMI', 'PASCORE', 'AMAMPT',
'AMPHIT', 'AMFVT', 'PQPSQI', 'PQPEFFCY', 'SLSLPHRS'])
df = mapping.copy()
df[vars_cts] = df[vars_cts].apply(pd.to_numeric, errors='coerce')
df[vars_cat] = df[vars_cat].apply(lambda x: x.astype('category'))
prep_info.to_csv('../data/prep_sleep.txt', sep= '\t', index=False)
df.to_csv('../data/mapping_sleep.txt', sep= '\t', na_rep='Missing:not collected', index=False)
| 1.1 prep_sample_info (Python).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: u4-s3-dnn
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/JAaron93/DS-Unit-4-Sprint-3-Deep-Learning/blob/main/LS_DS_Unit_4_Sprint_Challenge_3_AG.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="EPIia3JW7K_B"
#
# ## Autograded Notebook (Canvas & CodeGrade)
#
# This notebook will be automatically graded. It is designed to test your answers and award points for the correct answers. Following the instructions for each Task carefully.
# Instructions
#
# - **Download** this notebook as you would any other ipynb file
# - **Upload** to Google Colab or work locally (if you have that set-up)
# - **Delete** `raise NotImplementedError()`
#
# - **Write** your code in the `# YOUR CODE HERE` space
#
#
# - **Execute** the Test cells that contain assert statements - these help you check your work (others contain hidden tests that will be checked when you submit through Canvas)
#
# - **Save** your notebook when you are finished
# - **Download** as a ipynb file (if working in Colab)
# - **Upload** your complete notebook to Canvas (there will be additional instructions in Slack and/or Canvas)
#
#
# + [markdown] id="4b8-4bzQ7K_D"
# # Major Neural Network Architectures Challenge
# ## *Data Science Unit 4 Sprint 3 Challenge*
#
# In this sprint challenge, you'll explore some of the cutting edge of Deep Learning. This week we studied several famous neural network architectures:
# recurrent neural networks (RNNs), long short-term memory (LSTMs), convolutional neural networks (CNNs), and Autoencoders. In this sprint challenge, you will revisit these models. Remember, we are testing your knowledge of these architectures not your ability to fit a model with high accuracy.
#
# __*Caution:*__ these approaches can be pretty heavy computationally. All problems were designed so that you should be able to achieve results within at most 5-10 minutes of runtime locally, on AWS SageMaker, on Colab or on a comparable environment. If something is running longer, double check your approach!
#
# __*GridSearch:*__ CodeGrade will likely break if it is asked to run a gridsearch for a deep learning model (CodeGrade instances run on a single processor). So while you may choose to run a gridsearch locally to find the optimum hyper-parameter values for your model, please delete (or comment out) the gridsearch code and simply instantiate a model with the optimum parameter values to get the performance that you want out of your model prior to submission.
#
#
# ## Challenge Objectives
# *You should be able to:*
# * <a href="#p1">Part 1</a>: Train a LSTM classification model
# * <a href="#p2">Part 2</a>: Utilize a pre-trained CNN for object detection
# * <a href="#p3">Part 3</a>: Describe a use case for an autoencoder
# * <a href="#p4">Part 4</a>: Describe yourself as a Data Science and elucidate your vision of AI
#
# ____
#
# # (CodeGrade) Before you submit your notebook you must first
#
# 1) Restart your notebook's Kernel
#
# 2) Run all cells sequentially, from top to bottom, so that cell numbers are sequential numbers (i.e. 1,2,3,4,5...)
# - Easiest way to do this is to click on the **Cell** tab at the top of your notebook and select **Run All** from the drop down menu.
#
# 3) If you have gridsearch code, now is when you either delete it or comment out that code so CodeGrade doesn't run it and crash.
#
# 4) Read the directions in **Part 2** of this notebook for specific instructions on how to prep that section for CodeGrade.
#
# ____
# + [markdown] id="-5UwGRnJOmD4"
# <a id="p1"></a>
# ## Part 1 - LSTMs
#
# Use a LSTM to fit a multi-class classification model on Reuters news articles to distinguish topics of articles. The data is already encoded properly for use in a LSTM model.
#
# Your Tasks:
# - Use Keras to fit a predictive model, classifying news articles into topics.
# - Name your model as `model`
# - Use a `single hidden layer`
# - Use `sparse_categorical_crossentropy` as your loss function
# - Use `accuracy` as your metric
# - Report your overall score and accuracy
# - Due to resource concerns on CodeGrade, `set your model's epochs=1`
#
# For reference, the LSTM code we used in class will be useful.
#
# __*Note:*__ Focus on getting a running model, not on maxing accuracy with extreme data size or epoch numbers. Only revisit and push accuracy if you get everything else done!
# + id="DS-9ksWjoJit"
# Import data (don't alter the code in this cell)
from tensorflow.keras.datasets import reuters
# Suppress some warnings from deprecated reuters.load_data
import warnings
warnings.filterwarnings('ignore')
# Load data
(X_train, y_train), (X_test, y_test) = reuters.load_data(num_words=None,
skip_top=0,
maxlen=None,
test_split=0.2,
seed=723812,
start_char=1,
oov_char=2,
index_from=3)
# Due to limited computational resources on CodeGrade, take the following subsample
train_size = 1000
X_train = X_train[:train_size]
y_train = y_train[:train_size]
# + colab={"base_uri": "https://localhost:8080/"} id="fLKqFh8DovaN" outputId="8d7ce88e-6664-4ead-b350-b8662675376c"
# Demo of encoding
word_index = reuters.get_word_index(path="reuters_word_index.json")
print(f"Iran is encoded as {word_index['iran']} in the data")
print(f"London is encoded as {word_index['london']} in the data")
print("Words are encoded as numbers in our dataset.")
# + id="HhA7FgKW7K_I"
# Imports (don't alter this code)
from tensorflow.keras.preprocessing import sequence
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Embedding, LSTM
# DO NOT CHANGE THESE VALUES
# Keras docs say that the + 1 is needed: https://keras.io/api/layers/core_layers/embedding/
MAX_FEATURES = len(word_index.values()) + 1
# maxlen is the length of each sequence (i.e. document length)
MAXLEN = 200
# + colab={"base_uri": "https://localhost:8080/"} id="CSwKaoPt5zmn" outputId="b11a5556-1e9d-4380-b856-a2f45885898b"
MAX_FEATURES
# + deletable=false nbgrader={"cell_type": "code", "checksum": "36031ab4b52ab8412c65a71e101baaf5", "grade": false, "grade_id": "cell-471d7f5819bebff6", "locked": false, "schema_version": 3, "solution": true, "task": false} id="HdQM5rfb7K_J"
# Pre-process your data by creating sequences
# Save your transformed data to the same variable name:
# example: X_train = some_transformation(X_train)
# YOUR CODE HERE
X_train = sequence.pad_sequences(X_train, maxlen=MAXLEN)
X_test = sequence.pad_sequences(X_test, maxlen=MAXLEN)
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "cef4a5ae6ec56bee0d3121c7d8d37f3e", "grade": true, "grade_id": "cell-b46c98c26266363a", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false} id="ShEoVtC37K_K"
# Visible tests
assert X_train.shape[1] == MAXLEN, "Your train input sequences are the wrong length. Did you use the sequence import?"
assert X_test.shape[1] == MAXLEN, "Your test input sequences are the wrong length. Did you use the sequence import?"
# + [markdown] id="e_Fu3wjk7K_K"
# ### Create your model
#
# Make sure to follow these instructions (also listed above):
# - Name your model as `model`
# - Use a `single hidden layer`
# - Use `sparse_categorical_crossentropy` as your loss function
# - Use `accuracy` as your metric
#
# **Additional considerations**
#
# The number of nodes in your output layer should be equal to the number of **unique** values in the sequences you are training and testing on. For this text, that value is equal to 46.
#
# - Set the number of nodes in your output layer equal to 46
# + deletable=false nbgrader={"cell_type": "code", "checksum": "cc18fbac87183e4e19e0342541d5684b", "grade": false, "grade_id": "cell-5e7ea9089f827793", "locked": false, "schema_version": 3, "solution": true, "task": false} colab={"base_uri": "https://localhost:8080/"} id="yzvKQsmy7K_L" outputId="804d494e-3388-448e-eca1-f3817983f5d6"
# Build and complie your model here
# YOUR CODE HERE
# Values for output layer
num_output_values = 46
# Instantiating the sequential class
model = Sequential()
# Creating the input layer, explicitly!
model.add(Embedding(MAX_FEATURES, 128))
# creating a hidden layer
model.add(LSTM(128, recurrent_dropout=0.2))
# creating output layer
model.add(Dense(num_output_values, activation="sigmoid"))
# compiling the entire model
model.compile(loss="sparse_categorical_crossentropy", optimizer="adam", metrics=["accuracy"])
# summarizing our model
model.summary()
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "ff2e025e7744c0524ecebcb854ae632d", "grade": true, "grade_id": "cell-54f4676c642d2c94", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false} id="2jqSN88g7K_M"
# Visible Test
assert model.get_config()["layers"][1]["class_name"] == "Embedding", "Layer 1 should be an Embedding layer."
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "cc9bb4e19f4d22eb6a243d76024fb637", "grade": true, "grade_id": "cell-974465c65fe51083", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false} id="Z1Yt_eYP7K_M"
# Hidden Test
# + [markdown] id="HOzGMZQQ7K_M"
# ### Fit your model
#
# Now, fit the model that you built and compiled in the previous cells. Remember to set your `epochs=1`!
# + colab={"base_uri": "https://localhost:8080/"} deletable=false id="_QVSlFEAqWJM" nbgrader={"cell_type": "code", "checksum": "214ed02cdf6fe3f25483d81c2f4dd09c", "grade": false, "grade_id": "cell-10c20c87933d059c", "locked": false, "schema_version": 3, "solution": true, "task": false} outputId="a264fb05-000d-43a6-8d6f-843c05f4f2f0"
# Fit your model here
# REMEMBER to set epochs=1
# YOUR CODE HERE
model.fit(X_train,
y_train,
batch_size=42, # Tried using a batch_size that worked well in a prior experiment
epochs=1,
validation_data=(X_test, y_test))
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "ca6a93fe03cb86e1d3ba3d38fc848102", "grade": true, "grade_id": "cell-277a7dc0b08b9a29", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false} id="wmUmG1FM7K_N"
# Visible Test
n_epochs = len(model.history.history["loss"])
assert n_epochs == 1, "Verify that you set epochs to 1."
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "5035264cce6916dab4735e61ab5a92a0", "grade": false, "grade_id": "cell-e46402041c52cd24", "locked": true, "points": 0, "schema_version": 3, "solution": false, "task": true} nteract={"transient": {"deleting": false}} id="UagRTqpH7K_O"
# ## Sequence Data Question
# #### *Describe the `pad_sequences` method used on the training dataset. What does it do? Why do you need it?*
# + [markdown] deletable=false nbgrader={"cell_type": "markdown", "checksum": "466921cbc36892fb12aa9a3f0c2424a3", "grade": true, "grade_id": "cell-92a7ebc76ad66f05", "locked": false, "points": 0, "schema_version": 3, "solution": true, "task": false} id="uxEmJ3hn7K_O"
# This method is used to transform lists ( such as the length of samples we're using from our dataset ) of sequences, which are just lists of integers, into the shape of a Rank 2 Tensor (2D Numpy array).
#
# We use padded sequences in Recurrent Neural Networks because regular NNs, and even Convolutional Neural Networks, are too restrictive with their inputs and outputs. They only accept fixed-sized vectors while using a limited number of hidden layers. RNNs, such as the LSTM model, allow us to operate over sequences of vectors, allowing us to overcome the limitations of other NNs.
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "1162a5c137e05e13d9c1275bf89709c2", "grade": false, "grade_id": "cell-a7a697b125edb2b7", "locked": true, "points": 0, "schema_version": 3, "solution": false, "task": true} nteract={"transient": {"deleting": false}} id="w2HYgj4Q7K_O"
# ## RNNs versus LSTMs
# #### *What are the primary motivations behind using Long-ShortTerm Memory Cell unit over traditional Recurrent Neural Networks?*
# + [markdown] deletable=false nbgrader={"cell_type": "markdown", "checksum": "e88cf9a290290a182ca8381c5055b218", "grade": true, "grade_id": "cell-bfe45496d78d39bb", "locked": false, "points": 0, "schema_version": 3, "solution": true, "task": false} id="odU8mfAd7K_P"
# LSTM networks has a more robust update equation and backpropagation dynamics. Base RNNs lack memory cells that allow it to remove or add information to the cell.
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "72cdfd02468a598b328fef9d0fb2c449", "grade": false, "grade_id": "cell-ca70eabc807f8f52", "locked": true, "points": 0, "schema_version": 3, "solution": false, "task": true} nteract={"transient": {"deleting": false}} id="a61SkHJI7K_P"
# ## RNN / LSTM Use Cases
# #### *Name and Describe 3 Use Cases of LSTMs or RNNs and why they are suited to that use case*
# + [markdown] deletable=false nbgrader={"cell_type": "markdown", "checksum": "91b3a655469df8a2d64e7c7c0c0aed02", "grade": true, "grade_id": "cell-eeaef2336d124b88", "locked": false, "points": 0, "schema_version": 3, "solution": true, "task": false} id="4gk-bmuE7K_P"
# RNNs/LSTM models are best suited for speech recognition, text classification and generation, stocks and cryptocurrency, and weather forecasting.
#
# The memory cells of these models allow them to tackle almost any problem a ML engineer could throw at them!
# + [markdown] id="yz0LCZd_O4IG"
# <a id="p2"></a>
# ## Part 2- CNNs
#
# ### Find the Frog
#
# Time to play "find the frog!" Use Keras and [ResNet50v2](https://www.tensorflow.org/api_docs/python/tf/keras/applications/resnet_v2) (pre-trained) to detect which of the images with the `frog_images` subdirectory has a frog in it.
#
# <img align="left" src="https://d3i6fh83elv35t.cloudfront.net/newshour/app/uploads/2017/03/GettyImages-654745934-1024x687.jpg" width=400>
# + [markdown] id="GnG4Qo-37K_Q"
# The skimage function below will help you read in all the frog images into memory at once. You should use the preprocessing functions that come with ResnetV2, and you should also resize the images using scikit-image.
#
# ### Reading in the images
#
# The code in the following cell will download the images to your notebook (either in your local Jupyter notebook or in Google colab).
# + id="fxFfw6Ve7K_Q"
# Prep to import images (don't alter the code in this cell)
import urllib.request
# Text file of image URLs
text_file = "https://raw.githubusercontent.com/LambdaSchool/data-science-canvas-images/main/unit_4/sprint_challenge_files/frog_image_url.txt"
data = urllib.request.urlopen(text_file)
# Create list of image URLs
url_list = []
for line in data:
url_list.append(line.decode('utf-8'))
# + colab={"base_uri": "https://localhost:8080/"} id="whIqEWR236Af" outputId="fc346b81-9c39-40b5-ebf4-70006205d178"
# Import images (don't alter the code in this cell)
from skimage.io import imread
from skimage.transform import resize
# instantiate list to hold images
image_list = []
### UNCOMMENT THE FOLLOWING CODE TO LOAD YOUR IMAGES
#loop through URLs and load each image
for url in url_list:
image_list.append(imread(url))
## UNCOMMENT THE FOLLOWING CODE TO VIEW AN EXAMPLE IMAGE SIZE
#What is an "image"?
print(type(image_list[0]), end="\n\n")
print("Each of the Images is a Different Size")
print(image_list[0].shape)
print(image_list[1].shape)
# + [markdown] id="si5YfNqS50QU"
# ### Run ResNet50v2
#
# Your goal is to validly run ResNet50v2 on the input images - don't worry about tuning or improving the model. You can print out or view the predictions in any way you see fit. In order to receive credit, you need to have made predictions at some point in the following cells.
#
# *Hint* - ResNet 50v2 doesn't just return "frog". The three labels it has for frogs are: `bullfrog, tree frog, tailed frog`
#
# **Autograded tasks**
#
# * Instantiate your ResNet 50v2 and save to a variable named `resnet_model`
#
# **Other tasks**
# * Re-size your images
# * Use `resnet_model` to predict if each image contains a frog
# * Decode your predictions
# * Hint: the lesson on CNNs will have some helpful code
#
# **Stretch goals***
# * Check for other things such as fish
# * Print out the image with its predicted label
# * Wrap everything nicely in well documented functions
#
# ## Important note!
#
# To increase the chances that your notebook will run in CodeGrade, when you **submit** your notebook:
#
# * comment out the code where you load the images
# * comment out the code where you make the predictions
# * comment out any plots or image displays you create
#
# **MAKE SURE YOUR NOTEBOOK RUNS COMPLETELY BEFORE YOU SUBMIT!**
# + id="FaT07ddW3nHz"
# Imports
import numpy as np
import matplotlib.pyplot as plt
from tensorflow.keras.applications.resnet_v2 import ResNet50V2 # <-- pre-trained model
from tensorflow.keras.preprocessing import image
from tensorflow.keras.applications.resnet_v2 import preprocess_input, decode_predictions
# + colab={"base_uri": "https://localhost:8080/"} id="Xr4FWcdyMpsk" outputId="038483d3-8ac9-436a-9156-67b6f6cce0d1"
# Checking what images we're attempting to decode and display
url_list
# + colab={"base_uri": "https://localhost:8080/", "height": 53} id="GbUoA87LIf_k" outputId="e7459c9c-5ae2-41db-bede-de6b30fb0c32"
# This is the one I need! It's the only one without a \n attached to the end of its URL.
url_list[5]
# + id="Kx3iQr-hZNQy"
# Have to use image.load_image in the code cell below, but it only accepts filepaths! So I need to trick it into thinking
# the URL from url_list[5] is actually a filepath with this here code
import pathlib
import tensorflow as tf
# Because I couldn't get to this image programmatically from url_list, I'm assigning it here manually, thus I could have chosen any of the images form that list
image_dir = tf.keras.utils.get_file(origin="https://raw.githubusercontent.com/LambdaSchool/data-science-canvas-images/main/unit_4/sprint_challenge_files/frog_images/jared-evans-VgRnolD7OIw-unsplash.jpg", fname='frog_photo')
image_dir = pathlib.Path(image_dir)
# + colab={"base_uri": "https://localhost:8080/", "height": 287} id="SCL6noHHWsiE" outputId="c5d1b7f2-2f50-477c-c2e4-977c3b58f2ef"
# Now I can use our 'image directory' with image.load_image and it actually works! This method is far more reproducible than just uploading
# the image from that URL into Colab or Jupyter Notebook and assigning it to a filepath. This way, anyone can run this!
img = image.load_img(image_dir,
target_size = (224,224))
# Displaying our frog
plt.imshow(img)
# + id="naQ9KsWdXgt0"
# Preproccessing our frog image
img = image.img_to_array(img)
img = np.expand_dims(img, axis=0)
img = preprocess_input(img)
# + id="gMrgFHA_Xpnm"
# Instantiating our resnet model
resnet_model = ResNet50V2(weights='imagenet')
# + colab={"base_uri": "https://localhost:8080/"} id="mYBnQ2zzXwvC" outputId="c6bb9919-ffe8-4b98-e6e8-c7ab9c4626c7"
# Creating our predictions
preds = resnet_model.predict(img)
# Printing said preditions
print('Predicted:', decode_predictions(preds, top=3)[0])
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "dc052ce43e9c6139f9049a72613c53cb", "grade": true, "grade_id": "cell-6e0982cb9f7775ef", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false} id="N6SGfzF07K_T"
# Visible test
assert resnet_model.get_config()["name"] == "resnet50v2", "Did you instantiate the resnet model?"
# + [markdown] deletable=false editable=false id="XEuhvSu7O5Rf" nbgrader={"cell_type": "markdown", "checksum": "07da5c698aa6b4bfc985abf74be530d0", "grade": false, "grade_id": "cell-98f795ea1478ba74", "locked": true, "points": 0, "schema_version": 3, "solution": false, "task": true}
# <a id="p3"></a>
# ## Part 3 - Autoencoders
#
# **Describe a use case for an autoencoder given that an autoencoder tries to predict its own input.**
# + [markdown] deletable=false nbgrader={"cell_type": "markdown", "checksum": "b9def07acfd8d9a0bb2fb9b3a9d20170", "grade": true, "grade_id": "cell-1ec34a8c8251db51", "locked": false, "points": 0, "schema_version": 3, "solution": true, "task": false} id="yhHz8Lus7K_T"
# Autoencoders are great for text generation. We may desire a completely automated poem, song, or even an entire novel!
#
# What is this 'black-box' capable of when left to it's own backpropagated devices? What will it output? These questions inspire curiosity from people. This intrigue leads text generation into full-blown chatbots, where people can actually test whether these autoencoded DL models can actually mimic a proper interlocuter or not.
#
# Will they pass the Turing Test? It's all up to the effifacy of the autoencoder model.
# + [markdown] id="626zYgjkO7Vq"
# <a id="p4"></a>
# ## Part 4 - More...
# + [markdown] deletable=false editable=false id="__lDWfcUO8oo" nbgrader={"cell_type": "markdown", "checksum": "061b43713725ca6955ee708b4495d0df", "grade": false, "grade_id": "cell-75a20bfb51f81e3b", "locked": true, "points": 0, "schema_version": 3, "solution": false, "task": true}
# **Answer the following questions, with a target audience of a fellow Data Scientist:**
#
# - What do you consider your strongest area as a Data Scientist?
# - What area of Data Science would you most like to learn more about, and why?
# - Where do you think Data Science will be in 5 years?
#
# A few sentences per answer is fine - only elaborate if time allows.
# + [markdown] deletable=false nbgrader={"cell_type": "markdown", "checksum": "7e4b53bf4d823ee49c9fcb0e91c0dae3", "grade": true, "grade_id": "cell-7133ec302afe51d8", "locked": false, "points": 0, "schema_version": 3, "solution": true, "task": false} id="xUEyFs4h7K_U"
# My strongest area as a data scientist is with regard to machine learning engineering. Considering I actually scored in the Top Three in one of two Kaggle competitions I participated in. I just love experimenting with different types of feature engineering, data preprocessing, and **especially** hyper-parameter tuning!
#
# Or maybe my strongest suit is as a science communicator? Both of my Unit 1 & 2 Portfolio Projects were picked up by separate publications, so perhaps that's what I should pursue in the future.
#
# I would love to learn more about data infrastructure engineering,considering I have a startup idea that I'll be pursuing with Draper University a month after graduating from Lambda School.
#
# Considering the advent and adoption of AutoML, I believe data science will actually dry up within the next 5 years. Why would anyone need computational statisticians or ML engineers when Google Vertex exists? There will be a shift toward data engineering, where the creation of data scrapers, backend development and deployment will keep data scientists employed as the handling of APIs isn't as easily automated(for the near future anyway).
# + [markdown] id="_Hoqe3mM_Mtc"
# ## Congratulations!
#
# Thank you for your hard work, and [congratulations](https://giphy.com/embed/26xivLqkv86uJzqWk)!!! You've learned a lot, and you should proudly call yourself a Data Scientist.
#
| LS_DS_Unit_4_Sprint_Challenge_3_AG.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Classification on the Titanic Dataset
# The following example gives an idea about how you could run basic classification using a Gaussian mixture model on the Titanic dataset, using a latent node, continuous variables as well as discrete variables. The example uses cross validation to get a more robust accuracy score across the training and testing data sets.
#
# The initial step is our imports, and a bit of code for extracting floor and room number.
# +
# %matplotlib inline
import pandas as pd
import numpy as np
import re
import sys
sys.path.append("../../../bayesianpy")
import bayesianpy
import bayesianpy.visual
import logging
import os
from sklearn.cross_validation import KFold
from sklearn.metrics import accuracy_score
pattern = re.compile("([A-Z]{1})([0-9]{1,3})")
def get_cabin_floor_and_number(cabin):
if not isinstance(cabin, str):
return "", np.nan
cabins = cabin.split(" ")
for cabin in cabins:
match = re.match(pattern, cabin)
if match is not None:
floor = match.group(1)
number = match.group(2)
return floor, number
return "", np.nan
logger = logging.getLogger()
logger.addHandler(logging.StreamHandler())
logger.setLevel(logging.INFO)
# -
# The first step is a bit of preprocessing to get the data in the required format.
# +
db_folder = bayesianpy.utils.get_path_to_parent_dir("")
titanic = pd.read_csv(os.path.join(db_folder, "data/titanic.csv"))
titanic['Floor'], titanic['CabinNumber'] = zip(*titanic.Cabin.map(get_cabin_floor_and_number))
titanic.CabinNumber = titanic.CabinNumber.astype(float)
titanic.Floor.replace("", np.nan, inplace=True)
# drop variables that vary too much, e.g. with almost every row
titanic.drop(['Cabin', 'Ticket', 'Name', 'PassengerId'], inplace=True, axis=1)
# -
# It's then necessary to attach the thread to the JVM through a pipe created by Jpype (otherwise you get a recursion error message).
bayesianpy.jni.attach(logger)
# There are a few basic utility functions for deciding on the type of the data provided - obviously if you're already aware of the type then it's more accurate to manually specify datatypes.
# +
auto = bayesianpy.data.AutoType(titanic)
network_factory = bayesianpy.network.NetworkFactory(logger)
discrete = titanic[list(auto.get_discrete_variables())]
continuous = titanic[list(auto.get_continuous_variables())]
print("Discrete variables: {}".format(discrete.columns.tolist()))
print("Continuous variables: {}".format(continuous.columns.tolist()))
# -
# The structure will look something like the following (as visualised in networkx). Bayes Server does have a UI, so you could save the model that you generate through the API.
# +
# write data to the temporary sqllite db
with bayesianpy.data.DataSet(titanic, db_folder, logger) as dataset:
# Use a standard template, which generally gives good performance
mixture_naive_bayes_tpl = bayesianpy.template.MixtureNaiveBayes(logger, discrete=discrete, continuous=continuous)
model = bayesianpy.model.NetworkModel(
mixture_naive_bayes_tpl.create(network_factory),
logger)
# result contains a bunch of metrics regarding the training step
results = model.train(dataset)
layout = bayesianpy.visual.NetworkLayout(results.get_network())
graph = layout.build_graph()
pos = layout.fruchterman_reingold_layout(graph)
layout.visualise(graph, pos)
# -
# Finally, run the code through 3 folds to get an average score from three different models.
# +
# write data to the temporary sqllite db
with bayesianpy.data.DataSet(titanic, db_folder, logger) as dataset:
# Use a standard template, which generally gives good performance
mixture_naive_bayes_tpl = bayesianpy.template.MixtureNaiveBayes(logger, discrete=discrete, continuous=continuous)
k_folds = 3
kf = KFold(titanic.shape[0], n_folds=k_folds, shuffle=True)
score = 0
# use cross validation to try and predict whether the individual survived or not
for k, (train_indexes, test_indexes) in enumerate(kf):
model = bayesianpy.model.NetworkModel(
mixture_naive_bayes_tpl.create(network_factory),
logger)
# result contains a bunch of metrics regarding the training step
model.train(dataset.subset(train_indexes))
# note that we've not 'dropped' the target data anywhere, this will be retracted when it's queried,
# by specifying query_options.setQueryEvidenceMode(bayesServerInference().QueryEvidenceMode.RETRACT_QUERY_EVIDENCE)
results = model.batch_query(dataset.subset(test_indexes), bayesianpy.model.QueryMostLikelyState("Survived",
output_dtype=titanic['Survived'].dtype))
# Each query just appends a column/ columns on to the original dataframe, so results is the same as titanic.iloc[test_indexes],
# with (in this case) one additional column called 'Survived_maxlikelihood', joined to the original.
score += accuracy_score(y_pred=results['Survived_maxlikelihood'].tolist(),
y_true=results['Survived'].tolist())
print("Average score was {}. Baseline accuracy is about 0.61.".format(score / k_folds))
# -
| examples/notebook/titanic_classification.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .fs
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: .NET (F#)
// language: F#
// name: .net-fsharp
// ---
// [this doc on github](https://github.com/dotnet/interactive/tree/master/samples/notebooks/fsharp/Docs)
//
// # Charts with XPlot
// Charts can be rendered using [Xplot.Plotly](https://fslab.org/XPlot/).
// We will cover some example on how to use XPlot in a notebook with the .NET Kernel.
//
// First, import the `XPlot.Plotly` namespace:
open XPlot.Plotly
// The next cell sets up some helpers for data generation.
let generator = new Random()
// # Rendering Scatter plots
// One of the most commonly used type of chart to explore data set. Use the type `Scatter`.
// +
let openSeries =
Scatter(
name = "Open",
x = [1; 2; 3; 4],
y = [10; 15; 13; 17])
let closeSeries =
Scatter(
name = "Close",
x = [2; 3; 4; 5],
y = [16; 5; 11; 9])
[openSeries; closeSeries]
|> Chart.Plot
|> Chart.WithTitle "Open vs Close"
// -
// Let's change it to be markers style, so more like a scatter plot.
// +
openSeries.mode <- "markers"
closeSeries.mode <- "markers"
[openSeries; closeSeries]
|> Chart.Plot
// -
// `Scatter` can also produce polar charts by setting the radial property `r` and angular proeprty `t`
// +
let openSeries =
Scatter(
name = "Open",
r = [1.; 2.; 3.; 4.],
t = [45.; 100.; 150.; 290.])
let closeSeries =
Scatter(
name = "Close",
r = [2.; 3.; 4.; 5. ],
t = [16.; 45.; 118.; 90.])
[openSeries; closeSeries]
|> Chart.Plot
|> Chart.WithLayout(Layout(orientation = -90.))
// -
// ## Large scatter plots and performance
// It is not uncommong to have scatter plots with a large dataset, it is a common scenario at the beginning of a data exploration process. Using the default `svg` based rendering will create performace issues as the dom will become very large.
// We can then use `web-gl` support to address the problem.
// +
#!time
let series =
[|
for a in 1 .. 10 ->
Scattergl(
name = sprintf "Series %i" a,
mode = "markers",
x = [ for ax in 1 .. 100000 -> generator.Next(-200, 200) * 1000 * generator.Next(-2000, 2000)],
y = [ for ay in 1 .. 100000 -> generator.Next(-200, 200) * 1000 * generator.Next(-2000, 2000)])
|]
series
|> Chart.Plot
|> Chart.WithTitle "Large Dataset"
// -
// Can provide custom marker `colour`, `size` and `colorscale` to display even more information to the user.
// +
let generatePoint () = generator.Next(-200, 200) * 1000 * generator.Next(-2000, 2000)
let sizes =
[ for s in 1..100 ->
if generator.NextDouble() < 0.75 then
generator.Next(1, 5)
else
generator.Next(10, 15) ]
let temperatures = sizes |> Seq.map (fun x -> x * 10 - 100)
let series =
[|
for a in 1 .. 10 ->
Scattergl(
name = sprintf "Series %i" a,
mode = "markers",
x = [ for _ in 1 .. 100 -> generatePoint () ],
y = [ for ay in 1 .. 100 -> generatePoint () ],
marker =
Marker(
colorscale = "hot",
color = temperatures,
size = sizes))
|]
series
|> Chart.Plot
|> Chart.WithTitle "Size and Colour"
// -
// Plotly provides some additional `color scales` to use. Note that we use `display` explicitly to display each graph with separate titles, rather than a single chart.
// +
for s in series do s.marker.colorscale <- "Viridis"
series
|> Chart.Plot
|> Chart.WithTitle "Viridis scale"
|> display
for s in series do s.marker.colorscale <- "Hot"
series
|> Chart.Plot
|> Chart.WithTitle "Hot scale"
|> display
for s in series do s.marker.colorscale <- "Jet"
series
|> Chart.Plot
|> Chart.WithTitle "Jet scale"
|> display
// -
// # Rendering Histograms
// Let's have a look at using histograms, the next cell sets up some generators.
let count = 20
let dates = [for d in 1 .. count -> DateTime.Now.AddMinutes(float(generator.Next(d, d + 30)))]
// Now let's define histogram traces:
// +
let openByTime =
Histogram(
x = dates,
y = [for y in 1 .. count -> generator.Next(0, 200)],
name = "Open")
let closeByTime =
Histogram(
x = dates,
y = [for y in 1 .. count -> generator.Next(0, 200)],
name = "Close")
[openByTime; closeByTime]
|> Chart.Plot
// -
// The Histogram generator will automatically count the number of items per bin.
//
// Setting `histfunc` to `"sum"` we can now add up all the values contained in each bin.
// Note that we are creatng bin using the `x` data point and we are using bydefault autobinx
// +
let openByTime =
Histogram(
x = dates,
y = [for y in 1 .. count -> generator.Next(0, 200)],
name = "Open",
histfunc = "sum")
let closeByTime =
Histogram(
x = dates,
y = [for y in 1 .. count -> generator.Next(0, 200)],
name = "Close",
histfunc = "sum")
[openByTime; closeByTime]
|> Chart.Plot
// -
// # Area chart and Polar Area chart
// By populating hte property `fill` of a `Scatter` trace the chart will render as area chart.
//
// Here is set to `"tozeroy"` which will create a fill zone underneath the line reachin to the 0 of the y axis.
// +
let openSeries =
Scatter(
name = "Open",
x = [1; 2; 3; 4],
y = [10; 15; 13; 17],
fill = "tozeroy",
mode= "lines")
let closeSeries =
Scatter(
name = "Close",
x = [1; 2; 3; 4],
y = [3; 5; 11; 9],
fill = "tozeroy",
mode= "lines")
[openSeries; closeSeries]
|> Chart.Plot
|> Chart.WithTitle "Open vs Close"
// -
// With one `fill` set to `"tonexty"` the cahrt will fill the aread between traces.
// +
openSeries.fill <- None
closeSeries.fill <- "tonexty"
[openSeries; closeSeries]
|> Chart.Plot
|> Chart.WithTitle "Open vs Close"
// -
// Using `Area` traces we can generate radial area chart. In this example we are using cardinal points to xpress angular values.
// The list `["North"; "N-E"; "East"; "S-E"; "South"; "S-W"; "West"; "N-W"]` will be autoimatically translated to angular values.
// +
let winDirections = ["North"; "N-E"; "East"; "S-E"; "South"; "S-W"; "West"; "N-W"]
let areaTrace1 =
Area(
r = [77.5; 72.5; 70.0; 45.0; 22.5; 42.5; 40.0; 62.5],
t = winDirections,
name = "11-14 m/s",
marker = Marker(color = "rgb(106,81,163)"))
let areaTrace2 =
Area(
r = [57.49999999999999; 50.0; 45.0; 35.0; 20.0; 22.5; 37.5; 55.00000000000001],
t = winDirections,
name = "8-11 m/s",
marker = Marker(color = "rgb(158,154,200)"))
let areaTrace3 =
Area(
r = [40.0; 30.0; 30.0; 35.0; 7.5; 7.5; 32.5; 40.0],
t = winDirections,
name = "5-8 m/s",
marker = Marker(color = "rgb(203,201,226)"))
let areaTrace4 =
Area(
r = [20.0; 7.5; 15.0; 22.5; 2.5; 2.5; 12.5; 22.5],
t = winDirections,
name = "< 5 m/s",
marker = Marker(color = "rgb(242,240,247)"))
let areaLayout =
Layout(
title = "Wind Speed Distribution in Laurel, NE",
font = Font(size = 16.),
legend = Legend(font = Font(size = 16.)),
radialaxis = Radialaxis(ticksuffix = "%"),
orientation = 270.)
[areaTrace1; areaTrace2; areaTrace3; areaTrace4]
|> Chart.Plot
|> Chart.WithLayout areaLayout
// -
| samples/notebooks/fsharp/Docs/Plotting with Xplot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Deepu-Pk/AI-lab/blob/main/Experiment.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/", "height": 294} id="fEDyovG6a6nx" outputId="6e8bf857-da3d-4a34-e45b-80d5edd7a2f7"
# plotting a Normal distribution PDF with mean=0 and standard deviation = 1. 50 points taken.
# for univariate normal distribution : [-5,5] with step 0.2 taken for x axis.
import math
import random
import numpy as np
import matplotlib.pyplot as plt
from scipy.stats import norm
# %matplotlib inline
# Plot between -5 and 5 with 0.2 steps amounting to 50 data points
x_axis = np.arange(-5, 5, 0.2)
# Mean = 0, SD = 2.
# norm.pdf(array, mean, std)
plt.plot(x_axis, norm.pdf(x_axis,0,1))
plt.suptitle('A Univariate Normal Distribution')
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 428} id="XqbNJB4KbUG-" outputId="1255837a-0ed8-41a8-f26d-4ad03f6a7adb"
# Normal Distribution
# drawing i.i.d samples randomly from the normal distribution.
# defining a sample with 500 poplulation..
# samples taken random from this is to be used for all other purposes..
mu = 0
sigma = 1
parentDatapoints = np.random.normal(mu, sigma, 500).tolist()
# datapoints = np.random.normal(mu, sigma, 50)
datapoints = np.asarray(random.sample(parentDatapoints, 50))
#defining the size of the figure.
figure, ax = plt.subplots(1, figsize=(10,7))
# for plotting the histogram and making use of the bins for plotting normal
#distribution.
count, bins, ignored = plt.hist(datapoints,30, density=True)
# scipy's norm function can also be used inplace of this equation.
ax.plot(np.sort(datapoints),
1/(sigma * np.sqrt(2 * np.pi)) * np.exp( - (np.sort(datapoints) - mu)**2 / (2 * sigma**2) ),
linewidth=2, color='r')
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 431} id="qXXfhz80b9Qi" outputId="db2dcb82-8140-45d4-830c-eac09f42c234"
# plotting Log likelihood values for different mu and sigma..
# for the same datapoints obatined above
# This can also be used inplace of above function.
# This is the better alternative.
def LogLikelihoodFunction( mu, sigma, data):
value = np.sum(np.log(norm.pdf(data, mu, sigma)))
return value
mu1 = np.array([-2,-.8,-1,0,1.1,1.5, 1, 2])
sigma1 = np.array([2,2,1,1,1.1,1.6, 1.2, 1])
LogLikelihood=[] # defining a empty list for storing log likelihood
# calulating the log likelihood values of above mu1 and sigma1 for analysis
for i in range(len(mu1)):
LogLikelihood.append(LogLikelihoodFunction(mu1[i], sigma1[i], datapoints))
# Likelihood for a big array of samples as shown below
mu2 = np.arange(-6, 6, 0.2)
# Taking random values between o and 5 which has the same length as mu2 array
sigma2 = []
for i in range(len(mu2)):
sigma2.append(1)
# creating a list containing all the likelihood valus from above mu2 and sigma2
LogLikelihood2 = []
for i in range(len(mu2)):
LogLikelihood2.append(LogLikelihoodFunction(mu2[i], sigma2[i], datapoints))
fig,ax = plt.subplots(1,figsize=(10,6))
# plotting the log-likelihood values obtained above
ax.plot(np.arange(1, len(mu2)+1, 1), LogLikelihood2,c="r",
label="mu vs likelihood")
# doing the annotation for arg max only
max = np.argmax(LogLikelihood2)
ax.annotate(" N ($\mu$={}, $\sigma$={}) ".format(np.round(mu2[max],3),sigma2[max]),(max, LogLikelihood2[max]), c='black', )
plt.xlabel("Different Values of MU and SIGMA --- N(Mu,SIgma)")
plt.ylabel("Log Likelihood")
# to find which observation has the maximum log-likelihood value.
plt.axvline(max + 1, ls='--', c='black')
# np.argmax() returns the index of the value which is the highest in the list
fig.suptitle("Plotting the log-likelihood values for {} different mu and sigma".format(len(mu2)))
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 554} id="4FFvEa_CdGSQ" outputId="b88e01f0-6441-4289-9d32-e40ae2f65989"
figure, ax = plt.subplots(1, figsize=(10,8))
count, bins, ignored = ax.hist(datapoints,30, density=True, color='yellow')
# we already proved that Maximum-Likelihood Estimate of mu is the mean of
# the measurements and for sigma = standard deviation.
estimated_mu = np.mean(datapoints)
estimated_sigma= np.std(datapoints)
# plotting the PDF of Estimated Distribution.
ax.plot(bins, norm.pdf(bins,estimated_mu,estimated_sigma),'--', lw=3.5,c='b',label='mean={}, std={}'.format(round(np.mean(datapoints), 3),round(np.std(datapoints),3)))
ax.annotate(" Estimated Distribution", (0,0.4), c='b')
# plotting the PDF of True Distribution
ax.plot(bins, norm.pdf(bins,0,1),'-.', lw=3,color='green',label='mean={}, std={}'.format(0,1))
ax.annotate(" True Distribution", (0.7,0.3), c='g')
# plotting the PDF of all the tested mu and std values.
for i in range(len(mu1)):
if mu1[i]!= 0 or sigma1[i]!= 1:
# print(mu1[i])
ax.plot(bins, norm.pdf(bins,mu1[i],sigma1[i]),label='mean={}, std={}'.format(mu1[i], sigma1[i]))
ax.legend() # to show the legend
plt.axvline(0, ls='--', c='b') # for the vertical line passing through origin
plt.xlabel("Mean")
plt.ylabel("Normal Distribution")
figure.suptitle("Plotting for 50 sample datasets with different values of $\mu$ and $\sigma$")
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 863} id="pt4KklDZddQS" outputId="b3d5feea-ba31-47c4-f05e-021ac7f9ed45"
#Plotting the true and estimated distributions with increasing number of samples.
fig = plt.figure(figsize=(20,15))
mu = 0
sigma = 1
# 4 arrays containing datapoints from a normal distribution
datapoints1 =np.asarray(random.sample(parentDatapoints, 50))
datapoints2 = np.asarray(random.sample(parentDatapoints, 100))
datapoints3 = np.asarray(random.sample(parentDatapoints, 250))
datapoints4 = np.asarray(random.sample(parentDatapoints, 500))
# bins and hists for 4 arrays
count1, bins1, ignored1 = plt.hist(datapoints1,30, density=True, color='yellow')
count2, bins2, ignored2 = plt.hist(datapoints2,30, density=True, color='yellow')
count3, bins3, ignored3 = plt.hist(datapoints3,30, density=True, color='yellow')
count4, bins4, ignored4 = plt.hist(datapoints4,30, density=True, color='yellow')
# estimating mu and sigma for each array of samples
estimated_mu1 = np.mean(datapoints1)
estimated_sigma1= np.std(datapoints1)
estimated_mu2 = np.mean(datapoints2)
estimated_sigma2= np.std(datapoints2)
estimated_mu3 = np.mean(datapoints3)
estimated_sigma3= np.std(datapoints3)
estimated_mu4 = np.mean(datapoints4)
estimated_sigma4= np.std(datapoints4)
# PLOTTING
# plotting the PDF of Estimated Distribution for subplot 1
plt.plot(bins1, norm.pdf(bins1,estimated_mu1,estimated_sigma1),'--', lw=2.5,label='50 samples mean={}, std={}'.format(round(np.mean(datapoints1), 3),round(np.std(datapoints1),3)))
# plotting the PDF of True Distribution for subplot 1
# plt.plot(bins1, norm.pdf(bins1,0,1),'-.', lw=2,
# label='50 samples true mean=0, std=1')
# plt.title.set_text('50 Samples')
plt.legend()
# plotting the PDF of Estimated Distribution for subplot 2
plt.plot(bins2, norm.pdf(bins2,estimated_mu2,estimated_sigma2),'--', lw=2.5,
label='100 samples mean={}, std={}'.format(round(np.mean(datapoints2), 3),
round(np.std(datapoints2),3)))
plt.legend()
# plotting the PDF of Estimated Distribution for subplot 3
plt.plot(bins3, norm.pdf(bins3,estimated_mu3,estimated_sigma3),'--', lw=2.5,
label='250 samples mean={}, std={}'.format(round(np.mean(datapoints3), 3),
round(np.std(datapoints3),3)))
plt.legend()
# plotting the PDF of Estimated Distribution for subplot 4
plt.plot(bins4, norm.pdf(bins4,estimated_mu4,estimated_sigma4),'--', lw=2.5,
label='500 samples mean={}, std={}'.format(round(np.mean(datapoints4), 3),
round(np.std(datapoints4),3)))
# plotting the PDF of True Distribution for subplot 4.
plt.plot(bins4, norm.pdf(bins4,0,1), lw=5,
label='500 sample True mean=0, std=1')
# plt.title.set_text('500 Samples')
plt.legend()
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 428} id="EzdP3S2KeXwX" outputId="913ed06e-0c2f-44b6-a6ef-093f3cf9e1f8"
# taking the same samples which MLE was trained On.
# Implementing a function which returns the
# MAP estimate of the parameters given the data and the prior probability
# parameters
sigma_squared = 1 # setting variance of sample to 1
# DEFINING FUNCTION FOR MAP ESTIMATION
def MAP_Estimate(data, prior_nu, prior_beta_squared):
n = len(data) # no of samples
X_bar = np.mean(data) # measured mean of samples
# numerator of equation (1) in above derivation
numerator = sigma_squared * prior_nu + n * prior_beta_squared * X_bar
# denomenator
denomenator = sigma_squared + n * prior_beta_squared
# returns the MAP of mu
return numerator / denomenator
# logLikelihoodfunction already defined
# MAP_estimated_values for 50, 100, 250, 500 samples
map_estimate_50_samples = MAP_Estimate(datapoints1, .5, .5)
map_estimate_100_samples = MAP_Estimate(datapoints2, .5, .5)
map_estimate_250_samples = MAP_Estimate(datapoints3, .5, .5)
map_estimate_500_samples = MAP_Estimate(datapoints4, .5, .5)
# plotting likelihood, prior and MAP estimate next to each other
figure, ax = plt.subplots(1, figsize=(10,7))
count, bins1, ignored = plt.hist(datapoints1,30, density=True, color = 'yellow')
# scipy's norm function can also be used inplace of this equation.
ax.plot(bins1,norm.pdf(bins1, map_estimate_50_samples , sigma_squared) ,
linewidth=3, color='r', label = '$\hat{\mu}_{MAP}$ ')
ax.plot(bins1, norm.pdf(bins1,estimated_mu1,estimated_sigma1),'--', lw=2.5,c='b',
label='$\hat{\mu}_{MLE}$')
ax.plot(bins1, norm.pdf(bins1,0,1),'-.', lw=2.5,c='g',
label='True Value')
plt.legend()
ax.plot()
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 971} id="tFzrwizDezo0" outputId="36b66c8f-a820-4cc2-c238-62e574b561c9"
# Examining the behavior of the MAP estimate compared with the MLE estimate and
# the prior
# 1. repeatedly sampling the dataset and observing the variations in the estimate
# for this we use datapoints1, datapoints2, etc.,
fig = plt.figure(figsize=(15,15))
fig.suptitle('Plotting MLE, MAP and True Value based normal curves')
# PLOTTING
# plotting the PDF of Estimated Distribution for subplot 1
plt.plot(bins1, norm.pdf(bins1,estimated_mu1,estimated_sigma1),'--', lw=2.5,
label='50 samples $\hat{\mu}_{MLE}$')
# plotting the PDF of True Distribution for subplot 1
# ax1.plot(bins1, norm.pdf(bins1,0,1),'-.', lw=2,c='r',
# label='True')
# plotting the PDF of MAP for subplot 1.
plt.plot(bins1,norm.pdf(bins1, map_estimate_50_samples , sigma_squared) ,
linewidth=3, label = '50 samples $\hat{\mu}_{MAP}$ ')
# ax1.title.set_text('50 Samples')
plt.legend()
# plotting the PDF of Estimated Distribution for subplot 2
plt.plot(bins2, norm.pdf(bins2,estimated_mu2,estimated_sigma2),'--', lw=2.5,
label='100 samples $\hat{\mu}_{MLE}$ ')
# # plotting the PDF of True Distribution for subplot 2
# plt.plot(bins2, norm.pdf(bins2,0,1),'-.', lw=2,c='r',
# label='True')
# plotting the PDF of MAP for subplot 2.
plt.plot(bins2,norm.pdf(bins2, map_estimate_100_samples , sigma_squared) ,
linewidth=3, label = '100 samples $\hat{\mu}_{MAP}$ ')
# plt.title.set_text('100 Samples')
plt.legend()
# plotting the PDF of Estimated Distribution for subplot 3
plt.plot(bins3, norm.pdf(bins3,estimated_mu3,estimated_sigma3),'--', lw=2.5,
label='$250 samples \hat{\mu}_{MLE}$ ')
# # plotting the PDF of True Distribution for subplot 3
# plt.plot(bins3, norm.pdf(bins3,0,1),'-.', lw=2,c='r',
# label='True')
# plotting the PDF of MAP for subplot 2.
plt.plot(bins3,norm.pdf(bins3, map_estimate_250_samples , sigma_squared) ,
linewidth=3, label = '250 samples $\hat{\mu}_{MAP}$ ')
# plt.title.set_text('250 Samples')
plt.legend()
# plotting the PDF of Estimated Distribution for subplot 4
plt.plot(bins4, norm.pdf(bins4,estimated_mu4,estimated_sigma4),'--', lw=2.5,
label='$ 500 samples \hat{\mu}_{MLE}$')
# plotting the PDF of True Distribution for subplot 4.
plt.plot(bins4, norm.pdf(bins4,0,1), lw=6,c = 'g',
label='True')
# plotting the PDF of MAP for subplot 4.
plt.plot(bins4,norm.pdf(bins4, map_estimate_500_samples , sigma_squared) ,
linewidth=5,c = 'r' ,label = '500 samples $\hat{\mu}_{MAP}$ ')
# plt.title.set_text('500 Samples')
plt.legend()
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 602} id="yIyPo5V0fL18" outputId="f1629efc-b88c-4a96-fd48-ff5442d44ffd"
# Changing the prior parameters to see their effect on the estimate
# MAP_estimated_values on 50 samples and different prior parameters
# using MAP_Estimate(data, prior_nu, prior_beta_squared)
map_estimate_50_samples1 = MAP_Estimate(datapoints1, -5,1.5 )
map_estimate_50_samples2 = MAP_Estimate(datapoints1, -4, .09)
map_estimate_50_samples3 = MAP_Estimate(datapoints1, +10, .5)
map_estimate_50_samples4 = MAP_Estimate(datapoints1, 0, 1)
fig = plt.figure(figsize=(10,9))
ax1 = fig.add_subplot(221)
ax2 = fig.add_subplot(222)
ax3 = fig.add_subplot(223)
ax4 = fig.add_subplot(224)
fig.suptitle('MLE, MAP, True for 50 samples for differnet prior parameters')
# PLOTTING
# plotting the PDF of Estimated Distribution for subplot 1
ax1.plot(bins1, norm.pdf(bins1,estimated_mu1,estimated_sigma1),'--', lw=2.5,c='b',
label='$\hat{\mu}_{MLE}$')
# plotting the PDF of True Distribution for subplot 1
ax1.plot(bins1, norm.pdf(bins1,0,1),'-.', lw=2,c='r',
label='True')
# plotting the PDF of MAP for subplot 1.
ax1.plot(bins1,norm.pdf(bins1, map_estimate_50_samples1 , sigma_squared) ,
linewidth=3, color='g', label = '$\hat{\mu}_{MAP}$ ')
ax1.title.set_text('prior mean = -5 , std = 1.5')
ax1.legend()
# plotting the PDF of Estimated Distribution for subplot 2
ax2.plot(bins1, norm.pdf(bins1,estimated_mu1,estimated_sigma1),'--', lw=2.5,c='b',
label='$\hat{\mu}_{MLE}$')
# plotting the PDF of True Distribution for subplot 2
ax2.plot(bins1, norm.pdf(bins1,0,1),'-.', lw=2,c='r',
label='True')
# plotting the PDF of MAP for subplot 2
ax2.plot(bins1,norm.pdf(bins1, map_estimate_50_samples2 , sigma_squared) ,
linewidth=3, color='g', label = '$\hat{\mu}_{MAP}$ ')
ax2.title.set_text('prior mean = -4, std = 0.09')
ax2.legend()
# plotting the PDF of Estimated Distribution for subplot 3
ax3.plot(bins1, norm.pdf(bins1,estimated_mu1,estimated_sigma1),'--', lw=2.5,c='b',
label='$\hat{\mu}_{MLE}$')
# plotting the PDF of True Distribution for subplot 3
ax3.plot(bins1, norm.pdf(bins1,0,1),'-.', lw=2,c='r',
label='True')
# plotting the PDF of MAP for subplot 3
ax3.plot(bins1,norm.pdf(bins1, map_estimate_50_samples3 , sigma_squared) ,
linewidth=3, color='g', label = '$\hat{\mu}_{MAP}$ ')
ax3.title.set_text('prior mean = 10, std = 0.5 ')
ax3.legend()
# plotting the PDF of Estimated Distribution for subplot 4
ax4.plot(bins1, norm.pdf(bins1,estimated_mu1,estimated_sigma1),'--', lw=2.5,c='b',
label='$\hat{\mu}_{MLE}$')
# plotting the PDF of True Distribution for subplot 4
ax4.plot(bins1, norm.pdf(bins1,0,1),'-.', lw=2,c='r',
label='True')
# plotting the PDF of MAP for subplot 4
ax4.plot(bins1,norm.pdf(bins1, map_estimate_50_samples4 , sigma_squared) ,
linewidth=3, color='g', ls='-.', label = '$\hat{\mu}_{MAP}$ ')
ax4.title.set_text('prior mean = 0, std =1')
ax4.legend()
plt.show()
| Experiment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="toWe1IoH7X35"
# # IllusTrip: Text to Video 3D
#
# Part of [Aphantasia](https://github.com/eps696/aphantasia) suite, made by <NAME> [[eps696](https://github.com/eps696)]
# Based on [CLIP](https://github.com/openai/CLIP) + FFT/pixel ops from [Lucent](https://github.com/greentfrapp/lucent).
# 3D part by [deKxi](https://twitter.com/deKxi), based on [AdaBins](https://github.com/shariqfarooq123/AdaBins) depth.
# thanks to [<NAME>](https://twitter.com/advadnoun), [<NAME>](https://twitter.com/jonathanfly), [@eduwatch2](https://twitter.com/eduwatch2) for ideas.
#
# ## Features
# * continuously processes **multiple sentences** (e.g. illustrating lyrics or poems)
# * makes **videos**, evolving with pan/zoom/rotate motion
# * works with [inverse FFT](https://github.com/greentfrapp/lucent/blob/master/lucent/optvis/param/spatial.py) representation of the image or **directly with RGB** pixels (no GANs involved)
# * generates massive detailed textures (a la deepdream), **unlimited resolution**
# * optional **depth** processing for 3D look
# * various CLIP models
# * can start/resume from an image
#
# + [markdown] id="QytcEMSKBtN-"
# **Run the cell below after each session restart**
#
# Ensure that you're given Tesla T4/P4/P100 GPU, not K80!
# + id="etzxXVZ_r-Nf" cellView="form"
#@title General setup
# !pip install ftfy==5.8 transformers
# !pip install gputil ffpb
try:
# !pip3 install googletrans==3.1.0a0
from googletrans import Translator, constants
translator = Translator()
except: pass
# # !apt-get -qq install ffmpeg
work_dir = '/content/illustrip'
import os
os.makedirs(work_dir, exist_ok=True)
# %cd $work_dir
import os
import io
import time
import math
import random
import imageio
import numpy as np
import PIL
from base64 import b64encode
import shutil
from easydict import EasyDict as edict
a = edict()
import torch
import torch.nn as nn
import torch.nn.functional as F
import torchvision
from torchvision import transforms as T
from torch.autograd import Variable
from IPython.display import HTML, Image, display, clear_output
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
import ipywidgets as ipy
from google.colab import output, files
import warnings
warnings.filterwarnings("ignore")
# !pip install git+https://github.com/openai/CLIP.git --no-deps
import clip
# !pip install sentence_transformers
from sentence_transformers import SentenceTransformer
# !pip install kornia
import kornia
# !pip install lpips
import lpips
# !pip install PyWavelets==1.1.1
# !pip install git+https://github.com/fbcotter/pytorch_wavelets
# %cd /content
# !rm -rf aphantasia
# !git clone https://github.com/eps696/aphantasia
# %cd aphantasia/
from clip_fft import to_valid_rgb, fft_image, rfft2d_freqs, img2fft, pixel_image, un_rgb
from utils import basename, file_list, img_list, img_read, txt_clean, plot_text, old_torch
from utils import slice_imgs, derivat, pad_up_to, slerp, checkout, sim_func, latent_anima
import transforms
from progress_bar import ProgressIPy as ProgressBar
shutil.copy('mask.jpg', work_dir)
depth_mask_file = os.path.join(work_dir, 'mask.jpg')
clear_output()
def save_img(img, fname=None):
img = np.array(img)[:,:,:]
img = np.transpose(img, (1,2,0))
img = np.clip(img*255, 0, 255).astype(np.uint8)
if fname is not None:
imageio.imsave(fname, np.array(img))
imageio.imsave('result.jpg', np.array(img))
def makevid(seq_dir, size=None):
char_len = len(basename(img_list(seq_dir)[0]))
out_sequence = seq_dir + '/%0{}d.jpg'.format(char_len)
out_video = seq_dir + '.mp4'
# !ffpb -y -i $out_sequence -codec nvenc -crf 18 $out_video
data_url = "data:video/mp4;base64," + b64encode(open(out_video,'rb').read()).decode()
wh = '' if size is None else 'width=%d height=%d' % (size, size)
return """<video %s controls><source src="%s" type="video/mp4"></video>""" % (wh, data_url)
# Hardware check
# !ln -sf /opt/bin/nvidia-smi /usr/bin/nvidia-smi
import GPUtil as GPU
gpu = GPU.getGPUs()[0] # XXX: only one GPU on Colab and isn’t guaranteed
# !nvidia-smi -L
print("GPU RAM {0:.0f}MB | Free {1:.0f}MB)".format(gpu.memoryTotal, gpu.memoryFree))
# + id="JUvpdy8BWGuM" cellView="form"
#@title Load inputs
#@markdown **Content** (either type a text string, or upload a text file):
content = "" #@param {type:"string"}
upload_texts = False #@param {type:"boolean"}
#@markdown **Style** (either type a text string, or upload a text file):
style = "" #@param {type:"string"}
upload_styles = False #@param {type:"boolean"}
#@markdown For non-English languages use Google translation:
translate = False #@param {type:"boolean"}
#@markdown Resume from the saved `.pt` snapshot, or from an image
#@markdown (resolution settings below will be ignored in this case):
if upload_texts:
print('Upload main text file')
uploaded = files.upload()
text_file = list(uploaded)[0]
texts = list(uploaded.values())[0].decode().split('\n')
texts = [tt.strip() for tt in texts if len(tt.strip())>0 and tt[0] != '#']
print(' main text:', text_file, len(texts), 'lines')
workname = txt_clean(basename(text_file))
else:
texts = [content]
workname = txt_clean(content)[:44]
if upload_styles:
print('Upload styles text file')
uploaded = files.upload()
text_file = list(uploaded)[0]
styles = list(uploaded.values())[0].decode().split('\n')
styles = [tt.strip() for tt in styles if len(tt.strip())>0 and tt[0] != '#']
print(' styles:', text_file, len(styles), 'lines')
else:
styles = [style]
resume = False #@param {type:"boolean"}
if resume:
print('Upload file to resume from')
resumed = files.upload()
resumed_filename = list(resumed)[0]
resumed_bytes = list(resumed.values())[0]
assert len(texts) > 0 and len(texts[0]) > 0, 'No input text[s] found!'
tempdir = os.path.join(work_dir, workname)
os.makedirs(tempdir, exist_ok=True)
print('main dir', tempdir)
# + [markdown] id="PQFGziYKtHSa"
# **`content`** (what to draw) is your primary input; **`style`** (how to draw) is optional, if you want to separate such descriptions.
# If you load text file[s], the imagery will interpolate from line to line (ensure equal line counts for content and style lists, for their accordance).
# + id="Uti6XrqiQumf" cellView="form"
#@title Google Drive [optional]
#@markdown Run this cell, if you want to store results on your Google Drive.
using_GDrive = True#@param{type:"boolean"}
if using_GDrive:
import os
from google.colab import drive
if not os.path.isdir('/G/MyDrive'):
drive.mount('/G', force_remount=True)
gdir = '/G/MyDrive'
tempdir = os.path.join(gdir, 'illustrip', workname)
os.makedirs(tempdir, exist_ok=True)
print('main dir', tempdir)
# + id="64mlBCAYeOrB" cellView="form"
#@title Main settings
sideX = 1280 #@param {type:"integer"}
sideY = 720 #@param {type:"integer"}
steps = 200 #@param {type:"integer"}
frame_step = 100 #@param {type:"integer"}
#@markdown > Config
method = 'RGB' #@param ['FFT', 'RGB']
model = 'ViT-B/32' #@param ['ViT-B/16', 'ViT-B/32', 'RN101', 'RN50x16', 'RN50x4', 'RN50']
# Default settings
if method == 'RGB':
align = 'overscan'
colors = 2
contrast = 1.2
sharpness = -1.
aug_noise = 0.
smooth = False
else:
align = 'uniform'
colors = 1.8
contrast = 1.1
sharpness = 1.
aug_noise = 2.
smooth = True
interpolate_topics = True
style_power = 1.
samples = 200
save_step = 1
learning_rate = 1.
aug_transform = 'custom'
similarity_function = 'cossim'
macro = 0.4
enforce = 0.
expand = 0.
zoom = 0.012
shift = 10
rotate = 0.8
distort = 0.3
animate_them = True
sample_decrease = 1.
DepthStrength = 0.
print(' loading CLIP model..')
model_clip, _ = clip.load(model, jit=old_torch())
modsize = model_clip.visual.input_resolution
xmem = {'ViT-B/16':0.25, 'RN50':0.5, 'RN50x4':0.16, 'RN50x16':0.06, 'RN101':0.33}
if model in xmem.keys():
sample_decrease *= xmem[model]
clear_output()
print(' using CLIP model', model)
# + [markdown] id="JIWNmmd5uuSn"
# **`FFT`** method uses inverse FFT representation of the image. It allows flexible motion, but is either blurry (if smoothed) or noisy (if not).
# **`RGB`** method directly optimizes image pixels (without FFT parameterization). It's more clean and stable, when zooming in.
# There are few choices for CLIP `model` (results do vary!). I prefer ViT-B/32 for consistency, next best bet is ViT-B/16.
#
# **`steps`** defines the length of animation per text line (multiply it to the inputs line count to get total video duration in frames).
# `frame_step` sets frequency of the changes in animation (how many frames between motion keypoints).
#
#
# + [markdown] id="f3Sj0fxmtw6K"
# ## Other settings [optional]
# + id="P88_xcpAIXlq" cellView="form"
#@title Run this cell to override settings, if needed
#@markdown [to roll back defaults, run "Main settings" cell again]
style_power = 1. #@param {type:"number"}
overscan = True #@param {type:"boolean"}
align = 'overscan' if overscan else 'uniform'
interpolate_topics = True #@param {type:"boolean"}
#@markdown > Look
colors = 2 #@param {type:"number"}
contrast = 1.2 #@param {type:"number"}
sharpness = 0. #@param {type:"number"}
#@markdown > Training
samples = 200 #@param {type:"integer"}
save_step = 1 #@param {type:"integer"}
learning_rate = 1. #@param {type:"number"}
#@markdown > Tricks
aug_transform = 'custom' #@param ['elastic', 'custom', 'none']
aug_noise = 0. #@param {type:"number"}
macro = 0.4 #@param {type:"number"}
enforce = 0. #@param {type:"number"}
expand = 0. #@param {type:"number"}
similarity_function = 'cossim' #@param ['cossim', 'spherical', 'mixed', 'angular', 'dot']
#@markdown > Motion
zoom = 0.012 #@param {type:"number"}
shift = 10 #@param {type:"number"}
rotate = 0.8 #@param {type:"number"}
distort = 0.3 #@param {type:"number"}
animate_them = True #@param {type:"boolean"}
smooth = True #@param {type:"boolean"}
if method == 'RGB': smooth = False
# + [markdown] id="QYrJTb8xDm9C"
# `style_power` controls the strength of the style descriptions, comparing to the main input.
# `overscan` provides better frame coverage (needed for RGB method).
# `interpolate_topics` changes the subjects smoothly, otherwise they're switched by cut, making sharper transitions.
#
# Decrease **`samples`** if you face OOM (it's the main RAM eater), or just to speed up the process (with the cost of quality).
# `save_step` defines, how many optimization steps are taken between saved frames. Set it >1 for stronger image processing.
#
# Experimental tricks:
# `aug_transform` applies some augmentations, which quite radically change the output of this method (and slow down the process). Try yourself to see which is good for your case. `aug_noise` augmentation [FFT only!] seems to enhance optimization with transforms.
# `macro` boosts bigger forms.
# `enforce` adds more details by enforcing similarity between two parallel samples.
# `expand` boosts diversity (up to irrelevant) by enforcing difference between prev/next samples.
#
# Motion section:
# `shift` is in pixels, `rotate` in degrees. The values will be used as limits, if you mark `animate_them`.
#
# `smooth` reduces blinking, but induces motion blur with subtle screen-fixed patterns (valid only for FFT method, disabled for RGB).
# + [markdown] id="YdVubN0vb3TU"
# ## Add 3D depth [optional]
# + id="vl-rm1Nm03lK" cellView="form"
### deKxi:: This whole cell contains most of whats needed,
# with just a few changes to hook it up via frame_transform
# (also glob_step now as global var)
# I highly recommend performing the frame transformations and depth *after* saving,
# (or just the depth warp if you prefer to keep the other affines as they are)
# from my testing it reduces any noticeable stretching and allows the new areas
# revealed from the changed perspective to be filled/detailed
# pretrained models: Nyu is much better but Kitti is an option too
depth_model = 'nyu'#@param ["nyu","kitti"]
workdir_depth = "/content"
DepthStrength = 0.01 #@param{type:"number"}
MaskBlurAmt = 33 #@param{type:"integer"}
if DepthStrength > 0:
depthdir = os.path.join(tempdir, 'depth')
os.makedirs(depthdir, exist_ok=True)
print('depth dir', depthdir)
#@markdown NB: depth computing may take up to ~3x more time. Read the comments inside for more info.
#@markdown Courtesy of [deKxi](https://twitter.com/deKxi)
# Some useful misc funcs used
ToImage = T.ToPILImage()
def numpy2tensor(imgArray):
im = torch.unsqueeze(torchvision.transforms.ToTensor()(imgArray), 0)
return im
def triangle_blur(x, kernel_size=3, pow=1.0):
padding = (kernel_size-1) // 2
b,c,h,w = x.shape
kernel = torch.linspace(-1,1,kernel_size+2)[1:-1].abs().neg().add(1).reshape(1,1,1,kernel_size).pow(pow).cuda()
kernel = kernel / kernel.sum()
x = x.reshape(b*c,1,h,w)
x = F.pad(x, (padding,padding,padding,padding), mode='reflect')
x = F.conv2d(x, kernel)
x = F.conv2d(x, kernel.permute(0,1,3,2))
x = x.reshape(b,c,h,w)
return x
# Imports, some necessary and some just handy
###############
import cv2
import matplotlib.pyplot as plt
import os
if not os.path.exists(os.path.join(workdir_depth, 'AdaBins')):
# %cd $workdir_depth
# !git clone https://github.com/shariqfarooq123/AdaBins.git
workdir_depth = os.path.join(workdir_depth, "AdaBins")
# %cd $workdir_depth
if not os.path.exists('pretrained'):
# !mkdir pretrained
if depth_model=='nyu' and not os.path.exists(os.path.join(workdir_depth, "pretrained/AdaBins_nyu.pt")):
# !gdown https://drive.google.com/uc?id=1lvyZZbC9NLcS8a__YPcUP7rDiIpbRpoF
# !mv AdaBins_nyu.pt pretrained/AdaBins_nyu.pt
if depth_model=='kitti' and not os.path.exists(os.path.join(workdir_depth, "pretrained/AdaBins_kitti.pt")):
# !gdown https://drive.google.com/uc?id=1HMgff-FV6qw1L0ywQZJ7ECa9VPq1bIoj
# !mv AdaBins_kitti.pt pretrained/AdaBins_kitti.pt
############ Mask is for blending multi-crop depth
global mask_blurred
masksize = (830, 500) # I've hardcoded this but it doesn't have to be this exact number, this is just the max for what works at 16:9 for each crop
mask = cv2.imread(depth_mask_file, cv2.IMREAD_GRAYSCALE)
mask = cv2.resize(mask, masksize)
ch = sideY//2
cw = sideX//2
mask_blur = cv2.GaussianBlur(mask,(MaskBlurAmt,MaskBlurAmt),0)
mask_blurred = cv2.resize(mask_blur,(cw,ch)) / 255.
############
from infer import InferenceHelper
infer_helper = InferenceHelper(dataset='nyu')
# You can adjust AdaBins' internal depth max and min here, but unsure if it makes a huge difference - haven't tested it too much yet
#infer_helper.max_depth = infer_helper.max_depth * 50
#infer_helper.min_depth = infer_helper.min_depth * 1
# %cd /content/aphantasia/
def depthwarp(img, strength=0, rescale=0, midpoint=0.5, depth_origin=(0,0), clip_range=0, save_depth=True, multicrop=True):
if strength==0: return img
img2 = img.clone().detach() # Most of the pre-inference operations will take place on a dummy cloned tensor for simplicity sake
# Blurring first can somewhat mitigate the inherent noise from pixelgen method. Feel free to change these values if the depthmap is unsatisfactory
img2 = torch.lerp(img2, triangle_blur((img2), 5, 2), 0.5)
_, _, H, W = img2.shape
# This will define the centre/origin point for the depth extrusion
centre = torch.as_tensor([depth_origin[0],depth_origin[1]]).cpu()
# Converting the tensor to image in order to perform inference, probably a cleaner way to do this though as this was quick and dirty
par, imag, _ = pixel_image([1,3,H,W], resume=img2)
img2 = to_valid_rgb(imag, colors=colors)()
img2 = img2.detach().cpu().numpy()[0]
img2 = (np.transpose(img2, (1,2,0))) # convert image back to Height,Width,Channels
img2 = np.clip(img2*255, 0, 255).astype(np.uint8)
image = ToImage(img2)
del img2
# Resize down for inference
if H < W: # 500p on either dimension was the limit I found for AdaBins
r = 500 / float(H)
dim = (int(W * r), 500)
else:
r = 500 / float(W)
dim = (500, int(H * r))
image = image.resize(dim,3)
bin_centres, predicted_depth = infer_helper.predict_pil(image)
# Resize back to original before (optionally) adding the cropped versions
predicted_depth = cv2.resize(predicted_depth[0][0],(W,H))
if multicrop:
# This code is very jank as I threw it together as a quick proof-of-concept, and it miraculously worked
# There's very likely to be some improvements that can be made
clone = predicted_depth.copy()
# Splitting the image into separate crops, probably inefficiently
TL = torchvision.transforms.functional.crop(image.resize((H,W),3), top=0, left=0, height=cw, width=ch).resize(dim,3)
TR = torchvision.transforms.functional.crop(image.resize((H,W),3), top=0, left=ch, height=cw, width=ch).resize(dim,3)
BL = torchvision.transforms.functional.crop(image.resize((H,W),3), top=cw, left=0, height=cw, width=ch).resize(dim,3)
BR = torchvision.transforms.functional.crop(image.resize((H,W),3), top=cw, left=ch, height=cw, width=ch).resize(dim,3)
# Inference on crops
_, predicted_TL = infer_helper.predict_pil(TL)
_, predicted_TR = infer_helper.predict_pil(TR)
_, predicted_BL = infer_helper.predict_pil(BL)
_, predicted_BR = infer_helper.predict_pil(BR)
# Rescale will increase per object depth difference, but may cause more depth fluctuations if set too high
# This likely results in the depth map being less "accurate" to any real world units.. not that it was particularly in the first place lol
if rescale != 0:
# Histogram equalize requires a range of 0-255, but I'm recombining later in 0-1 hence this mess
TL = cv2.addWeighted(cv2.equalizeHist(predicted_TL.astype(np.uint8) * 255) / 255., 1-rescale,
predicted_TL.astype(np.uint8),rescale,0)
TR = cv2.addWeighted(cv2.equalizeHist(predicted_TR.astype(np.uint8) * 255) / 255., 1-rescale,
predicted_TR.astype(np.uint8),rescale,0)
BL = cv2.addWeighted(cv2.equalizeHist(predicted_BL.astype(np.uint8) * 255) / 255., 1-rescale,
predicted_BL.astype(np.uint8),rescale,0)
BR = cv2.addWeighted(cv2.equalizeHist(predicted_BR.astype(np.uint8) * 255) / 255., 1-rescale,
predicted_BR.astype(np.uint8),rescale,0)
# Combining / blending the crops with the original, quite a janky solution admittedly
TL = clone[0: ch, 0: cw] * (1 - mask_blurred) + cv2.resize(predicted_TL[0][0],(cw,ch)) * mask_blurred
TR = clone[0: ch, cw: cw+cw] * (1 - mask_blurred) + cv2.resize(predicted_TR[0][0],(cw,ch)) * mask_blurred
BL = clone[ch: ch+ch, 0: cw] * (1 - mask_blurred) + cv2.resize(predicted_BL[0][0],(cw,ch)) * mask_blurred
BR = clone[ch: ch+ch, cw: cw+cw] * (1 - mask_blurred) + cv2.resize(predicted_BR[0][0],(cw,ch)) * mask_blurred
# If you wish to display the depth map for each crop and for the merged version, uncomment these
#with outpic:
#plt.imshow(TL, cmap='plasma')
#plt.show()
#plt.imshow(TR, cmap='plasma')
#plt.show()
#plt.imshow(BL, cmap='plasma')
#plt.show()
#plt.imshow(BR, cmap='plasma')
#plt.show()
clone[0: ch, 0: cw] = TL
clone[0: ch, cw: cw+cw] = TR
clone[ch: ch+ch, 0: cw] = BL
clone[ch: ch+ch, cw: cw+cw] = BR
# I'm just multiplying the depths currently, but performing a pixel average is possibly a better idea
predicted_depth = predicted_depth * clone
predicted_depth /= np.max(predicted_depth) # Renormalize so we don't blow the image out of range
# Display combined end result
with outpic:
plt.imshow(predicted_depth, cmap='plasma')
plt.show()
#### Generating a new depth map on each frame can sometimes cause temporal depth fluctuations and "popping".
#### This part is just some of my experiments trying to mitigate that
# Dividing by average
#ave = np.mean(predicted_depth)
#predicted_depth = np.true_divide(predicted_depth, ave)
# Clipping the very end values that often throw the histogram equalize balance off which can mitigate rescales negative effects
gmin = np.percentile(predicted_depth,0+clip_range)#5
gmax = np.percentile(predicted_depth,100-clip_range)#8
clipped = np.clip(predicted_depth, gmin, gmax)
# Depth is reversed, hence the "1 - x"
predicted_depth = (1 - ((clipped - gmin) / (gmax - gmin))) * 255
# Rescaling helps emphasise the depth difference but is less "accurate". The amount gets mixed in via lerp
if rescale != 0:
rescaled = numpy2tensor(cv2.equalizeHist(predicted_depth.astype(np.uint8)))
rescaled = torchvision.transforms.Resize((H,W))(rescaled.cuda())
# Renormalizing again before converting back to tensor
predicted_depth = predicted_depth.astype(np.uint8) / np.max(predicted_depth.astype(np.uint8))
dtensor = numpy2tensor(PIL.Image.fromarray(predicted_depth)).cuda()
#dtensor = torchvision.transforms.Resize((H,W))(dtensor.cuda())
if rescale != 0: # Mixin amount for rescale, from 0-1
dtensor = torch.lerp(dtensor, rescaled, rescale)
if save_depth: # Save depth map out, currently its as its own image but it could just be added as an alpha channel to main image
global glob_step
saveddepth = dtensor.detach().clone().cpu().squeeze(0)
save_img(saveddepth, os.path.join(depthdir, '%05d.jpg' % glob_step))
dtensor = dtensor.squeeze(0)
# Building the coordinates, most of this is on CPU since it uses numpy
xx = torch.linspace(-1, 1, W)
yy = torch.linspace(-1, 1, H)
gy, gx = torch.meshgrid(yy, xx)
grid = torch.stack([gx, gy], dim=-1).cpu()
d = (centre-grid).cpu()
# Simple lens distortion that can help mitigate the "stretching" that appears in the periphery
lens_distortion = torch.sqrt((d**2).sum(axis=-1)).cpu()
#grid2 = torch.stack([gx, gy], dim=-1)
d_sum = dtensor[0]
# Adjust midpoint / move direction
d_sum = (d_sum - (torch.max(d_sum) * midpoint)).cpu()
# Apply the depth map (and lens distortion) to the grid coordinates
grid += d * d_sum.unsqueeze(-1) * strength
del image, bin_centres, predicted_depth
# Perform the depth warp
img = torch.nn.functional.grid_sample(img, grid.unsqueeze(0).cuda(), align_corners=True, padding_mode='reflection')
# Reset and perform the lens distortion warp (with reduced strength)
grid = torch.stack([gx, gy], dim=-1).cpu()
grid += d * lens_distortion.unsqueeze(-1) * (strength*0.31)
img = torch.nn.functional.grid_sample(img, grid.unsqueeze(0).cuda(), align_corners=True, padding_mode='reflection')
return img
# + [markdown] id="YZFuwNux8oEg"
# ## Generate
# + id="Nq0wA-wc-P-s" cellView="form"
#@title Generate
if aug_transform == 'elastic':
trform_f = transforms.transforms_elastic
sample_decrease *= 0.95
elif aug_transform == 'custom':
trform_f = transforms.transforms_custom
sample_decrease *= 0.95
else:
trform_f = transforms.normalize()
if enforce != 0:
sample_decrease *= 0.5
samples = int(samples * sample_decrease)
print(' using %s method, %d samples' % (method, samples))
if translate:
translator = Translator()
def enc_text(txt):
if translate:
txt = translator.translate(txt, dest='en').text
emb = model_clip.encode_text(clip.tokenize(txt).cuda()[:77])
return emb.detach().clone()
# Encode inputs
count = 0 # max count of texts and styles
key_txt_encs = [enc_text(txt) for txt in texts]
count = max(count, len(key_txt_encs))
key_styl_encs = [enc_text(style) for style in styles]
count = max(count, len(key_styl_encs))
assert count > 0, "No inputs found!"
# # !rm -rf $tempdir
# os.makedirs(tempdir, exist_ok=True)
opt_steps = steps * save_step # for optimization
glob_steps = count * steps # saving
if glob_steps == frame_step: frame_step = glob_steps // 2 # otherwise no motion
outpic = ipy.Output()
outpic
if method == 'RGB':
if resume:
img_in = imageio.imread(resumed_bytes) / 255.
params_tmp = torch.Tensor(img_in).permute(2,0,1).unsqueeze(0).float().cuda()
params_tmp = un_rgb(params_tmp, colors=1.)
sideY, sideX = img_in.shape[0], img_in.shape[1]
else:
params_tmp = torch.randn(1, 3, sideY, sideX).cuda() # * 0.01
else: # FFT
if resume:
if os.path.splitext(resumed_filename)[1].lower()[1:] in ['jpg','png','tif','bmp']:
img_in = imageio.imread(resumed_bytes)
params_tmp = img2fft(img_in, 1.5, 1.) * 2.
else:
params_tmp = torch.load(io.BytesIO(resumed_bytes))
if isinstance(params_tmp, list): params_tmp = params_tmp[0]
params_tmp = params_tmp.cuda()
sideY, sideX = params_tmp.shape[2], (params_tmp.shape[3]-1)*2
else:
params_shape = [1, 3, sideY, sideX//2+1, 2]
params_tmp = torch.randn(*params_shape).cuda() * 0.01
params_tmp = params_tmp.detach()
# function() = torch.transformation(linear)
# animation
if animate_them:
if method == 'RGB':
m_scale = latent_anima([1], glob_steps, frame_step, uniform=True, cubic=True, start_lat=[-0.3])
m_scale = 1 + (m_scale + 0.3) * zoom # only zoom in
else:
m_scale = latent_anima([1], glob_steps, frame_step, uniform=True, cubic=True, start_lat=[0.6])
m_scale = 1 - (m_scale-0.6) * zoom # ping pong
m_shift = latent_anima([2], glob_steps, frame_step, uniform=True, cubic=True, start_lat=[0.5,0.5])
m_angle = latent_anima([1], glob_steps, frame_step, uniform=True, cubic=True, start_lat=[0.5])
m_shear = latent_anima([1], glob_steps, frame_step, uniform=True, cubic=True, start_lat=[0.5])
m_shift = (m_shift-0.5) * shift * abs(m_scale-1.) / zoom
m_angle = (m_angle-0.5) * rotate * abs(m_scale-1.) / zoom
m_shear = (m_shear-0.5) * distort * abs(m_scale-1.) / zoom
def get_encs(encs, num):
cnt = len(encs)
if cnt == 0: return []
enc_1 = encs[min(num, cnt-1)]
enc_2 = encs[min(num+1, cnt-1)]
return slerp(enc_1, enc_2, opt_steps)
def frame_transform(img, size, angle, shift, scale, shear):
### deKxi:: Performing depth warp first so the standard affine zoom can remove any funkiness at the edges
if DepthStrength > 0:
# Some quick sine animating, didnt bother hooking them up to latent_anima since I replaced it with a different animation method in my own version
# d X/Y define the origin point of the depth warp, effectively a "3D pan zoom". Range is '-1 -> 1', with the ends being quite extreme
dX = 0.45 * float(math.sin(((glob_step % 114)/114) * math.pi * 2))
dY = -0.45 * float(math.sin(((glob_step % 166)/166) * math.pi * 2))
# # Midpointoffset == Movement Direction:
# 1 == everything recedes away, 0 == everything moves towards.
# (and oscillating/animating this value is quite visually pleasing IMO)
midpointOffset = 0.5 + (0.5 * float(math.sin(((glob_step % 70)/70) * math.pi * 2)))
depthOrigin = (dX, dY)
# Perform the warp
depthAmt = DepthStrength*scale # I like to multiply by zoom amount
# It might be worth combining shift with dX/Y change as well
# Rescale combined with clipping end values can improve temporal consistency of depth map
# but generally speaking the technique itself inherently has that fluctuation due to
# independently inferred frames. Best combined with a low depth strength to effectively
# average out the fluctuates
# ^ Performing a batch of depth inference with some augments and averaging could help
# alleviate this, but could end up being performance heavy. Was on my test todo list
# prior to posting publically, but haven't had the time to try it yet
img = depthwarp(img, strength=depthAmt, rescale=0.5, midpoint=midpointOffset, depth_origin=depthOrigin, clip_range=2, save_depth=True, multicrop=True)
if old_torch(): # 1.7.1
img = T.functional.affine(img, angle, shift, scale, shear, fillcolor=0, resample=PIL.Image.BILINEAR)
img = T.functional.center_crop(img, size)
img = pad_up_to(img, size)
else: # 1.8+
img = T.functional.affine(img, angle, shift, scale, shear, fill=0, interpolation=T.InterpolationMode.BILINEAR)
img = T.functional.center_crop(img, size) # on 1.8+ also pads
return img
prev_enc = 0
def process(num):
global params_tmp, opt_state, params, image_f, optimizer, pbar
if interpolate_topics:
txt_encs = get_encs(key_txt_encs, num)
styl_encs = get_encs(key_styl_encs, num)
else:
txt_encs = [key_txt_encs[min(num, len(key_txt_encs)-1)][0]] * opt_steps if len(key_txt_encs) > 0 else []
styl_encs = [key_styl_encs[min(num, len(key_styl_encs)-1)][0]] * opt_steps if len(key_styl_encs) > 0 else []
if len(texts) > 0: print(' ref text: ', texts[min(num, len(texts)-1)][:80])
if len(styles) > 0: print(' ref style: ', styles[min(num, len(styles)-1)][:80])
for ii in range(opt_steps):
global glob_step ### deKxi:: Making this global since I use it everywhere, but especially for saving the depth images out
glob_step = num * steps + ii // save_step # saving/transforming
loss = 0
txt_enc = txt_encs[ii].unsqueeze(0)
# motion: transform frame, reload params
if ii % save_step == 0:
# get encoded inputs
txt_enc = txt_encs[ii % len(txt_encs)].unsqueeze(0) if len(txt_encs) > 0 else None
styl_enc = styl_encs[ii % len(styl_encs)].unsqueeze(0) if len(styl_encs) > 0 else None
# render test frame
h, w = sideY, sideX
# transform frame for motion
scale = m_scale[glob_step] if animate_them else 1-zoom
trans = tuple(m_shift[glob_step]) if animate_them else [0, shift]
angle = m_angle[glob_step][0] if animate_them else rotate
shear = m_shear[glob_step][0] if animate_them else distort
if method == 'RGB':
img_tmp = frame_transform(params_tmp, (h,w), angle, trans, scale, shear)
params, image_f, _ = pixel_image([1,3,h,w], resume=img_tmp)
else: # FFT
if old_torch(): # 1.7.1
img_tmp = torch.irfft(params_tmp, 2, normalized=True, signal_sizes=(h,w))
img_tmp = frame_transform(img_tmp, (h,w), angle, trans, scale, shear)
params_tmp = torch.rfft(img_tmp, 2, normalized=True)
else: # 1.8+
if type(params_tmp) is not torch.complex64:
params_tmp = torch.view_as_complex(params_tmp)
img_tmp = torch.fft.irfftn(params_tmp, s=(h,w), norm='ortho')
img_tmp = frame_transform(img_tmp, (h,w), angle, trans, scale, shear)
params_tmp = torch.fft.rfftn(img_tmp, s=[h,w], dim=[2,3], norm='ortho')
params_tmp = torch.view_as_real(params_tmp)
params, image_f, _ = fft_image([1,3,h,w], resume=params_tmp, sd=1.)
image_f = to_valid_rgb(image_f, colors=colors)
del img_tmp
optimizer = torch.optim.Adam(params, learning_rate)
# optimizer = torch.optim.AdamW(params, learning_rate, weight_decay=0.01, amsgrad=True)
if smooth is True and num + ii > 0:
optimizer.load_state_dict(opt_state)
noise = aug_noise * (torch.rand(1, 1, *params[0].shape[2:4], 1)-0.5).cuda() if aug_noise > 0 else 0.
img_out = image_f(noise)
img_sliced = slice_imgs([img_out], samples, modsize, trform_f, align, macro)[0]
out_enc = model_clip.encode_image(img_sliced)
if method == 'RGB': # empirical hack
loss += 1.5 * abs(img_out.mean((2,3)) - 0.45).mean() # fix brightness
loss += 1.5 * abs(img_out.std((2,3)) - 0.17).sum() # fix contrast
if txt_enc is not None:
loss -= sim_func(txt_enc, out_enc, similarity_function)
if styl_enc is not None:
loss -= style_power * sim_func(styl_enc, out_enc, similarity_function)
if sharpness != 0: # mode = scharr|sobel|naive
loss -= sharpness * derivat(img_out, mode='naive')
# loss -= sharpness * derivat(img_sliced, mode='scharr')
if enforce != 0:
img_sliced = slice_imgs([image_f(noise)], samples, modsize, trform_f, align, macro)[0]
out_enc2 = model_clip.encode_image(img_sliced)
loss -= enforce * sim_func(out_enc, out_enc2, similarity_function)
del out_enc2; torch.cuda.empty_cache()
if expand > 0:
global prev_enc
if ii > 0:
loss += expand * sim_func(prev_enc, out_enc, similarity_function)
prev_enc = out_enc.detach().clone()
del img_out, img_sliced, out_enc; torch.cuda.empty_cache()
optimizer.zero_grad()
loss.backward()
optimizer.step()
if ii % save_step == save_step-1:
params_tmp = params[0].detach().clone()
if smooth is True:
opt_state = optimizer.state_dict()
if ii % save_step == 0:
with torch.no_grad():
img = image_f(contrast=contrast).cpu().numpy()[0]
save_img(img, os.path.join(tempdir, '%05d.jpg' % glob_step))
outpic.clear_output()
with outpic:
display(Image('result.jpg'))
del img
pbar.upd()
params_tmp = params[0].detach().clone()
outpic = ipy.Output()
outpic
pbar = ProgressBar(glob_steps)
for i in range(count):
process(i)
HTML(makevid(tempdir))
files.download(tempdir + '.mp4')
## deKxi: downloading depth video
if DepthStrength > 0:
HTML(makevid(depthdir))
files.download(depthdir + '.mp4')
# + [markdown] id="BsehQRircaw7"
# If video is not auto-downloaded after generation (for whatever reason), run this cell to do that:
# + id="m0BZVNi8cZUP"
files.download(tempdir + '.mp4')
if DepthStrength > 0:
files.download(depthdir + '.mp4')
| IllusTrip3D.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Car Price Predictor
# ### Importing the libraries
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
import warnings
warnings.filterwarnings('ignore')
# %matplotlib inline
sns.set_style('whitegrid')
# ### Importing the dataset
car_df = pd.read_csv('car_data.csv')
car_df.head()
car_df.tail()
car_df.shape
car_df.info()
car_df.isnull().sum()
car_df.company.unique()
print(car_df.year.unique())
car_df.Price.unique()
car_df.kms_driven.unique()
car_df.fuel_type.unique()
# ## From the above car data:
# - Year has many non-year values and is of object type
# - Price is also object type and has 'Ask for Price' data
# - kms_drive has kms with integers, object type and has nan values
# - fuel_type has nan values
# - keep first words of name
# ### Data cleaning and EDA
backup = car_df.copy()
# #### Cleaning the non-year values and shnaged to integer values
car_df = car_df[car_df['year'].str.isnumeric()]
car_df['year'] = car_df['year'].astype(int)
# #### Cleaned the non numeric price value and changed to integer type
car_df = car_df[car_df['Price'] != 'Ask For Price']
car_df['Price'] = car_df['Price'].str.replace(',', '').astype(int)
# #### Changed the kms driven value to integer value
car_df['kms_driven'] = car_df['kms_driven'].str.split(' ').str.get(0).str.replace(',', '')
car_df = car_df[car_df['kms_driven'].str.isnumeric()]
car_df['kms_driven'] = car_df['kms_driven'].astype(int)
# #### Dropped the nan fuel_type row
car_df = car_df[~car_df['fuel_type'].isna()]
car_df.info()
# #### Now dealing with the name of the cars
car_df['name'] = car_df['name'].str.split(' ').str.slice(0, 3).str.join(' ')
car_df.reset_index(drop=True)
car_df.describe()
# ##### As we see here our car's price to 75% is 4.912500e+05 but max price is too much i.e. 8.500003e+06.
car_df[car_df['Price'] > 6e6]
# ##### Soo we can say it as a outlier in our dataset soo we remove it.
car_df = car_df[car_df['Price'] < 6e6].reset_index(drop=True)
car_df
car_df.describe()
car_df.to_csv('car_data_cleaned.csv')
# ### Splitting the data into dependent and independent variables
X = car_df.drop('Price', axis=1)
y = car_df.Price
# ### Now dealing with categorial values which is turned into numbers by One Hot Encoder
# +
# Turn the categories into numbers
from sklearn.preprocessing import OneHotEncoder
from sklearn.compose import ColumnTransformer
categorial_features = ["name", "company", "fuel_type"]
one_hot = OneHotEncoder()
transformer = ColumnTransformer([("one_hot",
one_hot,
categorial_features)],
remainder="passthrough")
X = transformer.fit_transform(X)
X
# -
# ### Now splitting the dataset into training and test set
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=0)
# ### Fitting the model into the training dataset
from sklearn.linear_model import LinearRegression
from sklearn.tree import DecisionTreeRegressor
from sklearn.ensemble import RandomForestRegressor
from sklearn.svm import SVR
from xgboost import XGBRegressor
# ### Making a function fit_model for different ML algorithms
def fit_model(model, X_train, X_test, y_train, y_test):
model.fit(X_train, y_train)
print('The predicted values:\n ', model.predict(X_test))
print('\nThe accuracy score: ', model.score(X_test, y_test))
# ### 1. Linear Regression
fit_model(LinearRegression(), X_train, X_test, y_train, y_test)
# ### 2. Decision Tree Regressor
fit_model(DecisionTreeRegressor(), X_train, X_test, y_train, y_test)
# ### 3. Random Forest Regressor
fit_model(RandomForestRegressor(), X_train, X_test, y_train, y_test)
# ### 4. Support Vector Regressor
fit_model(SVR(), X_train, X_test, y_train, y_test)
# ### 5. XGB Regressor
fit_model(XGBRegressor(), X_train, X_test, y_train, y_test)
| 3. Car Price Prediction/Car Price Predictor.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:margin-gan] *
# language: python
# name: conda-env-margin-gan-py
# ---
# # MarginGAN: Adversarial Training in Semi-Supervised Learning
#
# **Authors:** <NAME>, <NAME>
#
# PyTorch implementation of the NIPS 2019 paper "MarginGAN".
#
# **Link:** https://papers.nips.cc/paper/9231-margingan-adversarial-training-in-semi-supervised-learning7
#
# **Code Authors:** <br/>
# <NAME> - <EMAIL> <br/>
# <NAME> - <EMAIL>
# We start with a brief overview of MarginGAN: Adversarial Training in Semi-Supervised Learning. To this end, we first introduce the concepts of semi-supervised learning and margin.
# ### Semi-Supervised Learning
# Semi-supervised learning (SSL) is a learning method that aims to train high accuracy classifiers utilizing only a limited amount of labeled data and large amounts of unlabeled data in the training process. This method is practical since labeled data is expensive, and unlabeled data is not hard to come by. In SSL, pseudo-label techniques are commonly used to assign labels to unlabeled data in order to incorporate unlabeled data to the training process to improve model robustness.
# ### Margin
# Next, we give the definition of margin, directly from the paper: “The margin of a single data point is defined to be the distance from that data point to a decision boundary, which can be used to bound the generalization error of the classifier.” In the SSL setting, the goal is to train high accuracy classifiers. But, we also want to have large-margin classifiers so that our predictions will be robust.
#
# To understand the idea of large-margin classifiers, a four-class problem is illustrated in Fig. 1. If the classifier believes the wrong pseudo-label, the decision boundary will be updated to cover the gap between the two classes of examples, the margin between the decision boundaries decreases, which hurts the generalization accuracy. Therefore, large-margin classifiers should ignore those wrong pseudo-labels to achieve higher accuracy. To construct MarginGAN, it is crucial to understand the margin notion in terms of SSL and multi-class classification.
#
# <p><img src="https://raw.githubusercontent.com/sonatbaltaci/marginGAN/master/figures/marginGAN_fig2.png" height=300px></p>
#
# Fig. 1: Four class toy example to illustrate the idea of large-margin classifiers [1]
#
#
# In SSL, the margin of an unlabeled example is denoted as $|f(x)|$, which is also represented as $\tilde{y}f(x)$ where $\tilde{y} = \text{sign}(f(x))$ [2].With this definition, the current prediction is regarded as correct and makes the classifier confident of what is predicted. If we define the margin in multi-class classification, it is the difference between the probability for the true class, and the maximal probability for the false classes,
#
# $$
# \text{Margin}(x,y) = C_{y}(x) - \max_{i \neq y}C_{i}(x)
# $$
#
# which is the evidence that if the margin is large, the probability of the correct class has peaked in the classification distribution.
#
# ### MarginGAN
# MarginGAN is a semi-supervised learning model that utilizes a standard GAN architecture to improve the performance of a classifier by increasing the margin of real examples and decreasing the margin of generated examples in parallel.
#
# In addition to a generator G and a discriminator D in a GAN architecture, there is also a classifier C in MarginGAN. With the addition of C, the standard two-player minimax game between G and D, now transforms into a three-player minimax game. The overall architecture of MarginGAN is provided in Fig. 2.
#
# <p><img src="https://raw.githubusercontent.com/sonatbaltaci/marginGAN/master/figures/marginGAN_fig1.png" height=300px></p>
#
# Fig. 2: Overall architecture of MarginGAN [1]
#
# #### Discriminator
#
# D’s objective is unchanged. D still tries to classify real and fake images. It considers both labeled and unlabeled images as real and generated images as fake. Therefore, loss of the discriminator is calculated as follows:
#
# $$
# Loss(D) = -\{ \mathbb{E}_{x \sim p^{[l]}(x)} [log(D(x))] + \mathbb{E}_{\tilde{x} \sim p^{[u]}(\tilde{x})} [log(D(\tilde{x}))] + \mathbb{E}_{z \sim p(z)} [log(1-D(G(z)))]\},
# $$
# where $p^{[l]}(x)$ is the distribution of the labeled images, $p^{[u]}(\tilde{x})$ is the distribution of the unlabeled images and $p(z)$ is the noise distribution from which the fake images are generated.
#
# #### Classifier
#
# In this three-player minimax game, the main idea is to increase the accuracy of C. The loss of C is calculated as the sum of the losses for three different types of samples, namely, labeled, unlabeled, and generated, and is thus threefold.
#
# For labeled samples, C has the same objective as ordinary multi-class classifiers, that is to classify each image correctly. For this purpose, standard cross-entropy loss is used.
#
# $$
# Loss(C^{[l]}) = \mathbb{E}_{(x,y) \sim p^{[l]}(x,y)} \left[ -\sum_{i=1}^{k} y_ilog(C(x)_i) \right],
# $$
#
# where $(x,y)$ is an image-label pair and $p^{[l]}(x,y)$ is the distribution from which the image-label pair is sampled and $k$ is the number of classes. Minimizing cross-entropy encourages the increase of the true class while suppressing other classes. This way the margin for these data points is increased.
#
#
# For unlabeled samples, the goal of C is to again increase the margin. However, since unlabeled samples have no known “true” label, a one-hot encoded pseudo-label concept is leveraged. The one-hot encoded vectors have 1 for the class with the maximum predicted probability of the current C, while other entries are 0. To increase the margin of the unlabeled samples, cross-entropy loss is minimized but this time, the image-label pair consists of unlabeled images and pseudo-labels.
#
# $$
# Loss(C^{[u]}) = \mathbb{E}_{\tilde{x} \sim p^{[u]}(\tilde{x})} \left[ -\sum_{i=1}^{k} \tilde{y}^{[u]}_ilog(C(\tilde{x})_i) \right],
# $$
#
# where $p^{[u]}(\tilde{x})$ is the distribution of the unlabeled images, $\tilde{y}^{[u]}$ is the one-hot encoded pseudo-label vector for the unlabeled examples. The idea is to boost the confidence of the current predictions and thus this process increases the margins.
#
# For the generated examples, C tries to decrease the margin in order to make the distribution flat. For this reason, “Inverted Cross-Entropy (ICE) loss” is introduced. Since generated examples are also unlabeled, the pseudo-label concept is employed again. ICE loss is defined as follows:
#
# $$
# Loss(C^{[g]}) = \mathbb{E}_{z\sim p(z)} \left[ -\sum_{i=1}^{k} \tilde{y}^{[g]}_ilog(1-C(G(z))_i) \right],
# $$
#
# where $p(z)$ is the noise distribution and $\tilde{y}^{[g]}$ is the one-hot encoded pseudo-label vector for the generated examples. Minimizing ICE loss will increase the cross-entropy between the pseudo-label and $C(G(z))$, so that the prediction distribution will be flat.
#
# The loss of the classifier is thus:
#
# $$
# Loss(C) = Loss(C^{[l]}) + Loss(C^{[u]}) + Loss(C^{[g]})
# $$
#
# #### Generator
#
# As in the standard GAN, G tries to fool D by generating realistic examples. On the other hand, because C increases the margin of real examples and decreases the margin of fake examples, G generates data points having large margin to fool C. Therefore, in order to fool both D and C, G tries to generate realistic and large-margin examples simultaneously such that the generated fake data points cannot easily be separated from real examples. So, the loss for the generator is calculated as:
#
# $$
# Loss(G) = - \mathbb{E}_{z \sim p(z)} [log(D(G(z)))] + \mathbb{E}_{z \sim p(z)} \left[ -\sum_{i=1}^{k} \tilde{y}^{[g]}_ilog(C(G(z))_i) \right],
# $$
#
# where $p(z)$ is the noise distribution and $\tilde{y}^{[g]}$ is the one-hot encoded pseudo label vector for the generated examples.
#
# #### Pre-training
#
# The paper uses 4 different numbers of labeled examples, $100$, $600$, $1000$, and $3000$ to conduct the experiments. For G and D, MarginGAN utilizes the well-known architecture of InfoGAN [3]. To predict the pseudo-labels, the classifier architecture of MarginGAN is pre-trained for all label sizes to be used within the training of the corresponding architecture. The pre-training is performed until C reached the accuracy levels of $80\%$, $93\%$, $95\%$, and $97\%$, respectively for each label size. Later, it is fine-tuned along with G and D. The generated examples of the final model are near the decision boundary, that is their class labels cannot be determined with ease, which is in line with the aim of MarginGAN.
#
#
# ### Experimental setup
#
# We perform our experiments using Python 3.7.6 and PyTorch 1.5.0. For the full details of the setup, we include an environment.yml file. We provide both CPU and GPU support to execute the code. Here is a complete list of all hyperparameters:
# * Learning Rate of Classifier: $0.01$
# * Learning Rate of Generator: $0.0002$
# * Learning Rate of Discriminator: $0.0002$
# * Momentum for SGD Optimizer: $0.8$
# * $\beta_1$ for Adam Optimizer: $0.5$
# * $\beta_2$ for Adam Optimizer: $0.999$
# * Batch Size: $50$
# * Torch Seed: $0$
#
# ### Workflow of the code
#
# * We first pre-process the data before training and testing processes. For this reason, we normalize both the training set and the test set.
# * Although it is not stated in the paper, we reserve 10k samples from the 60k training samples to be the validation set.
# * We experiment with 4 different number of labels: $100$, $600$, $1000$, and $3000$ while keeping the number of samples from each class balanced.
# * The partitioning of the dataset is kept the same for all the pre-training and fine-tuning procedures.
# * For each number of labels, classifier is pre-trained to ensure certain percentages of accuracies: $80\%$, $93\%$, $95\%$, and $97\%$, respectively.
# * To train the MarginGAN for each number of labels separately, the model is initialized with a random generator, a random discriminator and a pre-trained classifier.
# * The hyperparameters are determined via experimentation and commonly used values.
# * To feed the labeled and unlabeled data to the network in parallel, the labeled data is cycled with respect to its ratio to unlabeled data.
# * For each epoch, the model is trained with labeled and unlabeled data and it is evaluated on the validation set afterward while keeping track of the best accuracy achieved.
# * The training and validation processes are the same with many deep learning models. The main difference for this study is the use of different loss functions during training.
# * For each number of labels, the best models and their results are saved.
# * Each hyperparameter setting has been trained 5 times as in [1] to observe the variation of the error rates.
# * The best models of the 5 runs are then tested on the test set. Mean and standard errors of the 5 runs are reported as in [1]. Also, Table-1 from [1] is recreated comparing our results with [1].
#
#
# Our code is presented below.
# ### Import the libraries
import torch.optim as optim
import torchvision.transforms as transforms
import argparse
import warnings
from itertools import cycle
from marginGAN import *
from utils import *
from dataset import *
from model_paths import *
# ### Set the hyperparameters
batch_size = 50 # batch size
seed = 0 # manual seed
num_epochs = 50 # number of epochs
lrC = 0.01 # learning rate of classifier
lrG = 0.0002 # learning rate of generator
lrD = 0.0002 # learning rate of discriminator
beta_1 = 0.5 # beta 1 for Adam optimizer
beta_2 = 0.999 # beta 2 for Adam optimizer
momentum = 0.8 # momentum for SGD optimizer
log_every = 1000 # log frequency
job_id = 1 # log id
device = "cuda:0" # device
# ### Select the device
if torch.cuda.is_available():
device = torch.device(device)
else:
device = torch.device("cpu")
# +
# Ignore warnings to avoid clutter when using different PyTorch versions
warnings.filterwarnings("ignore")
# Provide manual seed for reproducibility
torch.manual_seed(seed)
# Create save directories
create_dirs(job_id)
# -
# ### Log parameters
# +
params = {'job_id': job_id,
'batch_size': batch_size,
'seed': seed,
'num_epochs': num_epochs,
'lrC': lrC,
'lrG': lrG,
'lrD': lrD,
'beta_1': beta_1,
'beta_2': beta_2,
'momentum': momentum,
}
# Save the hyperparameters
param_log(params)
# -
# ### Load the datasets
# +
# Reserve 10,000 samples for validation
dataset = torchvision.datasets.MNIST("./data",train=True,transform=transforms.Compose([transforms.ToTensor(),transforms.Normalize((0.5,),(0.5,))]),target_transform=None,download=True)
trainset, valset = torch.utils.data.random_split(dataset, [50000, 10000])
testset = torchvision.datasets.MNIST("./data",train=False,transform=transforms.Compose([transforms.ToTensor(),transforms.Normalize((0.5,), (0.5,))]),target_transform=None,download=True)
valloader = torch.utils.data.DataLoader(
dataset=valset,
batch_size=batch_size,
shuffle=False)
testloader = torch.utils.data.DataLoader(
dataset=testset,
batch_size=batch_size,
shuffle=False)
# -
# ### Sample a fixed noise for the reproducibility of generated samples
latent_dim = 62
fixed_noise = torch.rand(batch_size,latent_dim).to(device)
# ### Set the number of labeled examples
label_sizes = [100,600,1000,3000]
# ### Print the model
marginGAN = MarginGAN().print_model()
# ### Train and save the model
# Perform training on 4 different number of labeled examples
for label_size in label_sizes:
# Divide the dataset into labeled and unlabeled samples
labeled_set, unlabeled_set = divide_dataset(trainset, label_size)
labeled_loader = torch.utils.data.DataLoader(dataset=labeled_set,batch_size=batch_size,shuffle=True)
unlabeled_loader = torch.utils.data.DataLoader(dataset=unlabeled_set,batch_size=batch_size,shuffle=True)
# Load the model
marginGAN = MarginGAN(label_size=label_size,
batch_size=batch_size,
device=device,
lrs=[lrD, lrG, lrC],
beta_1=beta_1,
beta_2=beta_2,
momentum=momentum,
pretrained="pretrained_classifiers/pre_cls_label_model_"+str(label_size)+".pt")
# Cycle over the same labeled examples during training
zipped_loader = zip(cycle(labeled_loader), unlabeled_loader)
best_correct = 0
for epoch in range(num_epochs):
# Train for one epoch
marginGAN.train(zipped_loader,epoch,log_every)
# Reinitialize the iterator
zipped_loader = zip(cycle(labeled_loader), unlabeled_loader)
if (epoch+1) % 1 == 0:
# Evaluate the model on the validation set
correct_guesses = marginGAN.eval(valloader)
if correct_guesses > best_correct:
best_correct = correct_guesses
# Save the model
marginGAN.save(epoch,fixed_noise,job_id)
# Log the accuracy
print('[',epoch+1,']','Accuracy: %d %%' % (100 * correct_guesses / 10000), correct_guesses,"/", 10000)
# ### Load and test the model
# +
# Create a model instance
marginGAN = MarginGAN()
err_hist = []
var_hist = []
for label_size in label_sizes:
all_model_paths = model_paths()
models = all_model_paths[str(label_size)]
worst_err = 100
best_err = 0
acc_hist = []
# Compute the average, best and worst error rates over 5 runs
for i in range(5):
marginGAN.load(models[i])
correct = marginGAN.eval(testloader)
acc_hist.append(correct)
# 10,000 test examples in MNIST
total_samples = 10000
avg_err = (total_samples - np.mean(acc_hist))/100
worst_err = (total_samples - min(acc_hist))/100
best_err = (total_samples - max(acc_hist))/100
# Fill the table
if abs(avg_err - worst_err) > abs(avg_err - best_err):
err_hist.append(str(round(avg_err,2)))
var_hist.append(str(round(abs(avg_err-worst_err),2)))
else:
err_hist.append(str(round(avg_err,2)))
var_hist.append(str(round(abs(avg_err-best_err),2)))
from IPython.display import HTML, display
import tabulate
table = [["Method","Label Size: 100","Label Size: 600","Label Size: 1000","Label Size: 3000"],
["Ours",err_hist[0]+" \u00B1 "+var_hist[0],err_hist[1]+" \u00B1 "+var_hist[1],err_hist[2]+" \u00B1 "+var_hist[2],err_hist[3]+" \u00B1 "+var_hist[3]],
["Paper [1]","3.53 \u00B1 0.57","3.03 \u00B1 0.60","2.87 \u00B1 0.71","2.06 \u00B1 0.20"]]
display(HTML(tabulate.tabulate(table, tablefmt='html',floatfmt=".2f")))
# -
# Table 1: Mean and standard error rates of the classifier over 5 runs (Reproduction of Table 1 from [1])
# <br/>
#
# ### Reproduce the qualitative results
# +
# Generate two random integers to select a random model that is to be used in generating a fake image batch with MarginGAN's generator
lab = np.random.randint(0,4)
ind = np.random.randint(0,5)
all_model_paths = model_paths()
model = all_model_paths[str(label_sizes[lab])][ind]
# Create a model instance
marginGAN = MarginGAN()
# Load the randomly selected model
marginGAN.load(model)
# Show the randomly sampled image
marginGAN.imshow(fixed_noise)
# -
# Fig. 3: Reproduction of Figure 3-a from [1]
#
# <br/>
#
# <p><img src="https://raw.githubusercontent.com/sonatbaltaci/marginGAN/master/figures/marginGAN_fig3.png" height=300px></p>
#
# Fig. 4: Figure 3-a from [1]
#
# <br/>
# ## Challenges Faced
#
# Although the overall idea to utilize the GAN architecture to enhance the semi-supervised learning process was clearly explained, crucial architectural details were missing:
#
# ### Dataset
# Firstly, since MNIST has no specific validation set, we separate 10000 samples to be used as the validation set for hyperparameter optimization. This was not mentioned in the paper. Additionally, no information provided about the normalization of the data. It is stated in various studies that the normalization of data helps the training process. Thus we normalized training, validation, and test data.
#
# We know that the paper uses 4 different number of labeled examples for their experiments. These are stated to be $100$, $600$, $1000$, and $3000$. However, randomly sampling 100 examples from the training set may cause a class imbalance. Therefore, to create a balanced labeled training set, we choose an equal number of examples from each class. After we divided the dataset into labeled and unlabeled subsets, we observed that there were different techniques in literature to feed the data into the model in semi-supervised learning. In various research, all labeled and unlabeled images are passed into the architecture only once per epoch. On the other hand, since the number of unlabeled images is much higher than the number of labeled images within one iteration, some researchers prefer to iterate the labeled images in a cycle until the unlabeled images are all passed. This choice wasn’t stated in the paper. We experimented with both options and iterating the labels gave better training performance and validation results.
#
# ### Model Architecture
# Architecture of C was proposed to be a “simple 6-layer” C, but no further detail was given. The number of convolutional and fully-connected layers, the channel and kernel sizes of the layers, and the type of the activation functions were unclear. We first experimented with several architectures having a simple 6-layer structure. However, we observed that increasing the number of layers by adding batch normalization following the convolutional layers improved our pre-training speed and performance (of the main architecture) remarkably. Therefore, we set the architecture of C a little bigger than it is in the paper.
#
# ### Optimizers and Hyperparameters
# Most of the hyperparameters and which optimizers were used by the authors were not specified in the paper. Thus, when choosing the hyperparameters and optimizers, we follow common practices used in research, check recent important works on generative models and experiment with different settings and test our results on the validation set.
#
# For the optimizer selection, since InfoGAN [3] and DCGAN [4] uses Adam optimizers for their G and D with $\beta_1$ value of $0.5$ and $\beta_2$ value of $0.999$, we also use Adam optimizer for G and D with these $\beta$ values. For classifiers, it is common practice to use an SGD optimizer. Therefore, we use an SGD optimizer for C.
#
# Since MarginGAN uses the G and D of the InfoGAN [3], we experimented with the hyperparameter settings in the utilized in InfoGAN [3]. Furthermore, we checked popular generative model papers like DCGAN [4] to find a good range of learning rates for our discriminator and generator. We also investigated general practices used in research and experimented with different settings. For the learning rate value of D, we tried different values and for most cases D was stable. Therefore, we choose $0.0002$ to be the learning rate of D, which was the setting in InfoGAN [3] and DCGAN [4], both.
#
# Choosing the learning rate for G was trickier. The learning rate used in InfoGAN [3] was $0.001$ but we found that this value was too high and made the training process for G unstable. In addition, the images generated with this learning rate were meaningless. Therefore, after experimenting with lower values, we choose $0.0002$ to be the learning rate of our G.
#
# In the paper, they mention in Chapter 4.2 that they reduce the learning rate of C from $0.1$ to $0.01$ in these experiments. We deduce that in Chapter 4.1 (where we reproduce the results) the paper uses $0.1$ learning rate for C. However, after experimenting with different values of the learning rate and the momentum parameter of SGD optimizer, we found that using a $0.01$ learning rate for C was much more suitable along with a momentum value of $0.8$. Again, there was no mention of the momentum value that the authors used, in the paper.
#
#
# ### Pre-training
#
# In [1], it was stated that the pre-training procedure was performed until C reached the error rates of $8.0\%, 9.3\%, 9.5\%$, and $9.7\%$, respectively for each label size. This seemed illogical because we expect to see better accuracies as the number of labeled examples increase. Also in [5], before including the pseudo-labels, the Drop-NN model is pre-trained with the same numbers of labeled data ($100, 600, 1000, 3000$), and obtained $78\%$, $91.5\%$, $93.5\%$, and $96.5\%$ accuracy respectively. This is a contradiction and we believe this is a mistake on [1]’s side. Therefore, we interpret the error rates in [1] similar to [5] and use $80\%$, $93\%$, $95\%$, and $97\%$ accuracy for each label as to stop the pre-training process.
#
#
# ### References
#
# [1] Dong, Jinhao, and <NAME>. "MarginGAN: Adversarial Training in Semi-Supervised Learning." Advances in Neural Information Processing Systems. 2019.
#
# [2] Bennett, <NAME>., <NAME>, and <NAME>. "Exploiting unlabeled data in ensemble methods." Proceedings of the eighth ACM SIGKDD international conference on Knowledge discovery and data mining. 2002.
#
# [3] <NAME>, et al. "Infogan: Interpretable representation learning by information maximizing generative adversarial nets." Advances in neural information processing systems. 2016.
#
# [4] <NAME>, <NAME>, and <NAME>. "Unsupervised representation learning with deep convolutional generative adversarial networks." arXiv preprint arXiv:1511.06434 (2015).
#
# [5] <NAME>. "Pseudo-label: The simple and efficient semi-supervised learning method for deep neural networks." Workshop on challenges in representation learning, ICML. Vol. 3. 2013.
#
| MarginGAN/main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:.conda-alphapept] *
# language: python
# name: conda-env-.conda-alphapept-py
# ---
# +
# default_exp score
# -
# # Score
#
# > Functions related to the score
# This notebook contains all functions related to the scoring of peptide-spectrum-matches (PSMS).
#
# In brief, this notebook includes the following:
#
# - Functions for target-decoy based FDR estimation
# - X!tandem based scoring of PSMs
# - Machine learning based scoring of PSMs
# - Protein grouping by the razor approach
# ## Filtering
#
# The filtering functions are essential base functions for scoring in AlphaPept. They make sure that only the 'best precursor per spectum' and the 'best spectrum per precursor' is used.
#
# Recall from the search that when having feautres, `raw_idx` refers to the actual index from the raw data. Otherwise it is`query_data`.
#
# For filtering, we have several functions. When applying for a score, we first use `filter_score` and then `filter_precursor`.
# `filter_score` is keeping the best score per experimental spectrum. First we rank by score for each `query_idx`. As we have multiple hits for each experimental spectrum from the search we only want to keep the best one.
#
# When performing feature finding, we assign multiple possible features to each experimental spectrum. The idea here is that a spectrum could originate from various precursors. To disentangle these psms we can use the following modes:
#
# * `single`: This mode will only keep one feature per experimental spectrum (the one with the highest score and the closest distance). Each feature can only occur once.
# * `multiple`: Allow multiple features per experimental spectrum. Each feature can only occur once.
#
# `filter_precusor` is intended for the case that a precursor (charge + sequence) occurs more than once. Only the one with the highest score will be kept.
# +
#export
import numpy as np
import pandas as pd
import logging
import alphapept.io
def filter_score(df: pd.DataFrame, mode: str='multiple') -> pd.DataFrame:
"""
Filter psms feature table by keeping only the best scoring psm per experimental spectrum.
TODO: psms could still have the same score when having modifications at multiple positions that are not distinguishable.
Only keep one.
Args:
df (pd.DataFrame): psms table of search results from alphapept.
mode (str, optional): string specifying which mode to use for psms filtering. The two options are 'single' and 'multiple'. 'single' will only keep one feature per experimental spectrum. 'multiple' will allow multiple features per experimental spectrum. In either option, each feature can only occur once. Defaults to 'multiple'.
Returns:
pd.DataFrame: table containing the filtered psms results.
"""
df["rank"] = df.groupby("query_idx")["score"].rank("dense", ascending=False).astype("int")
df = df[df["rank"] == 1]
# in case two hits have the same score and therfore the same rank only accept the first one
df = df.drop_duplicates("query_idx")
if 'dist' in df.columns:
df["feature_rank"] = df.groupby("feature_idx")["dist"].rank("dense", ascending=True).astype("int")
df["raw_rank"] = df.groupby("raw_idx")["score"].rank("dense", ascending=False).astype("int")
if mode == 'single':
df_filtered = df[(df["feature_rank"] == 1) & (df["raw_rank"] == 1) ]
df_filtered = df_filtered.drop_duplicates("raw_idx")
elif mode == 'multiple':
df_filtered = df[(df["feature_rank"] == 1)]
else:
raise NotImplementedError('Mode {} not implemented yet'.format(mode))
else:
df_filtered = df
# TOD: this needs to be sorted out, for modifications -> What if we have MoxM -> oxMM, this will screw up with the filter sequence part
return df_filtered
# +
#hide
def test_filter_score():
## DataFrame with unique assignments
df = pd.DataFrame({'query_idx':[1,1,2,2,3,3], 'score':[1,2,3,4,5,6],'feature_idx':[1,1,1,3,4,5],'raw_idx':[1,1,2,2,3,3]})
assert len(filter_score(df)) == 3
## Spectra competing for a feature, only keep one
df = pd.DataFrame({'query_idx':[1,2], 'score':[1,2],'feature_idx':[1,1],'raw_idx':[1,2], 'dist':[1,2]})
assert len(filter_score(df) == 1)
## Same spectra multiple features
df = pd.DataFrame({'query_idx':[1,2], 'score':[1,2],'feature_idx':[1,2],'raw_idx':[1,1], 'dist':[1,2]})
assert len(filter_score(df, mode='single')) == 1
assert len(filter_score(df, mode='multiple')) == 2
test_filter_score()
# +
#export
def filter_precursor(df: pd.DataFrame) -> pd.DataFrame:
"""
Filter psms feature table by precursor.
Allow each precursor only once.
Args:
df (pd.DataFrame): psms table of search results from alphapept.
Returns:
pd.DataFrame: table containing the filtered psms results.
"""
df["rank_precursor"] = (
df.groupby("precursor")["score"].rank("dense", ascending=False).astype("int")
)
df_filtered = df[df["rank_precursor"] == 1]
return df_filtered
# +
#hide
def test_filter_precursor():
df = pd.DataFrame({'precursor':['A','A','A'],'score':[1,2,3]})
assert len(filter_precursor(df)) == 1
df = pd.DataFrame({'precursor':['A','A','B'],'score':[1,2,3]})
assert len(filter_precursor(df)) == 2
df = pd.DataFrame({'precursor':['A','B','C'],'score':[1,2,3]})
assert len(filter_precursor(df)) == 3
test_filter_precursor()
# -
# ## Q-Values
#
# `get_q_values` is used to calculate q-values from FDR values. The direct relationship is illustrated further down in the notebook.
#export
from numba import njit
@njit
def get_q_values(fdr_values: np.ndarray) -> np.ndarray:
"""
Calculate q-values from fdr_values.
Args:
fdr_values (np.ndarray): np.ndarray of fdr values.
Returns:
np.ndarray: np.ndarray of q-values.
"""
q_values = np.zeros_like(fdr_values)
min_q_value = np.max(fdr_values)
for i in range(len(fdr_values) - 1, -1, -1):
fdr = fdr_values[i]
if fdr < min_q_value:
min_q_value = fdr
q_values[i] = min_q_value
return q_values
# +
#hide
def test_get_q_values():
assert np.allclose(get_q_values(np.array([1,2,3,4])), np.array([1,2,3,4]))
assert np.allclose(get_q_values(np.array([3,3,3,3])), np.array([3,3,3,3]))
assert np.allclose(get_q_values(np.array([4,3,2,1])), np.array([1,1,1,1]))
test_get_q_values()
# -
# ## FDR
#
# The employed FDR strategy is based on a classical target-decoy competition approach. The procedure works as follows:
# 1. Consider only the best scoring target or decoy PSM per spectrum.
# 2. Sort all PSMs by decreasing scores.
# 3. Estimate the FDR as #decoys / #targets, where #targets (#decoys) is the number of positive target (decoy) PSMs at a given score threshold t (i.e. PSMs with scores higher than t).
# 4. Convert the estimated FDR to q-values by selecting the minimum FDR at which the identification could be made, i.e. the lowest score threshold t that could be set to include an identification without increasing the number of false positives.
# 5. Report the set of target PSMs with q-values smaller or equal to the selected `fdr_level`.
#
# Informative literature describing and discussing different FDR estimation approaches for shotgun proteomics can be found here (the implemented strategy in alphapept is referred to as T-TDC in this article):
# > Keich, <NAME>. "Improved False Discovery Rate Estimation Procedure for Shotgun Proteomics." Journal of proteome research vol. 14,8 (2015): 3148-61. <https://pubs.acs.org/doi/10.1021/acs.jproteome.5b00081>
#
# +
#export
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
#Note that the test function for cut_fdr is further down in the notebook to also test protein-level FDR.
def cut_fdr(df: pd.DataFrame, fdr_level:float=0.01, plot:bool=True) -> (float, pd.DataFrame):
"""
Cuts a dataframe with a given fdr level
Args:
df (pd.DataFrame): psms table of search results from alphapept.
fdr_level (float, optional): fdr level that should be used for filtering. The value should lie between 0 and 1. Defaults to 0.01.
plot (bool, optional): flag to enable plot. Defaults to 'True'.
Returns:
float: numerical value of the applied score cutoff
pd.DataFrame: df with psms within fdr
"""
df["target"] = ~df["decoy"]
df = df.sort_values(by=["score","decoy"], ascending=False)
df = df.reset_index()
df["target_cum"] = np.cumsum(df["target"])
df["decoys_cum"] = np.cumsum(df["decoy"])
df["fdr"] = df["decoys_cum"] / df["target_cum"]
df["q_value"] = get_q_values(df["fdr"].values)
last_q_value = df["q_value"].iloc[-1]
first_q_value = df["q_value"].iloc[0]
if last_q_value <= fdr_level:
logging.info('Last q_value {:.3f} of dataset is smaller than fdr_level {:.3f}'.format(last_q_value, fdr_level))
cutoff_index = len(df)-1
elif first_q_value >= fdr_level:
logging.info('First q_value {:.3f} of dataset is larger than fdr_level {:.3f}'.format(last_q_value, fdr_level))
cutoff_index = 0
else:
cutoff_index = df[df["q_value"].gt(fdr_level)].index[0] - 1
cutoff_value = df.loc[cutoff_index]["score"]
cutoff = df[df["score"] >= cutoff_value]
targets = df.loc[cutoff_index, "target_cum"]
decoy = df.loc[cutoff_index, "decoys_cum"]
fdr = df.loc[cutoff_index, "fdr"]
logging.info(f"{targets:,} target ({decoy:,} decoy) of {len(df)} PSMs. fdr {fdr:.6f} for a cutoff of {cutoff_value:.2f} (set fdr was {fdr_level})")
if plot:
import matplotlib.pyplot as plt
plt.figure(figsize=(10, 5))
plt.plot(df["score"], df["fdr"])
plt.axhline(0.01, color="k", linestyle="--")
plt.axvline(cutoff_value, color="r", linestyle="--")
plt.title("fdr vs Cutoff value")
plt.xlabel("Score")
plt.ylabel("fdr")
# plt.savefig('fdr.png')
plt.show()
bins = np.linspace(np.min(df["score"]), np.max(df["score"]), 100)
plt.figure(figsize=(10, 5))
plt.hist(df[df["decoy"]]["score"].values, label="decoy", bins=bins, alpha=0.5)
plt.hist(df[~df["decoy"]]["score"].values, label="target", bins=bins, alpha=0.5)
plt.xlabel("Score")
plt.ylabel("Frequency")
plt.title("Score vs Class")
plt.legend()
plt.show()
cutoff = cutoff.reset_index(drop=True)
return cutoff_value, cutoff
# -
# Note that the test function for cut_fdr is further down in the notebook to also test protein-level FDR.
# +
#hide
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
#Helper functions to create in-silico data.
def simulateTargetDecoyScores(n, target_mu=4.0, stdev=1.0, pi0=0.5):
decoys = np.random.normal(loc=0.0, scale=stdev, size=n)
false_targets = np.random.normal(loc=0.0, scale=stdev, size=int(np.round(n*pi0)))
true_targets = np.random.normal(loc=target_mu, scale=stdev, size=int(np.round(n*(1-pi0))))
df = pd.DataFrame({
'TD':np.append(np.append(np.repeat('TT',len(true_targets)),np.repeat('FT',len(false_targets))),np.repeat('D',len(decoys))),
'decoy':np.append(np.repeat(False,len(true_targets)+len(false_targets)), np.repeat(True,len(decoys))),
'score':np.append(np.append(true_targets,false_targets),decoys),
'sequence':np.append(np.arange(0,n),np.arange(0,n)),
'protein':np.append(np.arange(0,n),np.arange(0,n))})
return df
def simulateProteinLevelTargetDecoyScores(n, target_mu=4.0, stdev=1.0, pi0=0.5, plot=True):
idx = np.arange(0,n)
protein_size = np.random.poisson(lam=3.0, size=n) + 1
if plot:
plt.hist(protein_size)
plt.title("Number of peptides per protein")
plt.xlabel("Number of peptides per protein")
plt.ylabel("Count")
plt.show()
TT_protein_size = protein_size[idx[0:int(np.round(1-(n*pi0)))]]
FT_protein_size = protein_size[idx[int(np.round(1-(n*pi0))):n]]
D_protein_size = protein_size
true_targets = np.random.normal(loc=target_mu, scale=stdev, size=sum(TT_protein_size))
false_targets = np.random.normal(loc=0.0, scale=stdev, size=sum(FT_protein_size))
decoys = np.random.normal(loc=0.0, scale=stdev, size=sum(D_protein_size))
D_proteins = np.repeat(idx,D_protein_size)
TT_proteins = np.repeat(idx[0:int(np.round(1-(n*pi0)))],TT_protein_size)
FT_proteins = np.repeat(idx[int(np.round(1-(n*pi0))):n],FT_protein_size)
df = pd.DataFrame({
'TD':np.append(np.append(np.repeat('TT',len(TT_proteins)),np.repeat('FT',len(FT_proteins))),np.repeat('D',len(D_proteins))),
'decoy':np.append(np.repeat(False,len(TT_proteins)+len(FT_proteins)), np.repeat(True,len(D_proteins))),
'score':np.append(np.append(true_targets,false_targets),decoys),
'sequence':np.append(np.arange(0,sum(protein_size)),np.arange(0,sum(protein_size))),
'protein':np.append(np.append(TT_proteins,FT_proteins),D_proteins)})
return df
def plot_score_hist(df, analyte_level='sequence'):
if analyte_level=='protein':
df = df.sort_values(by=['protein','score'], ascending=False)
df = df.drop_duplicates(subset='protein', keep="first")
decoys=df[df.decoy].score.values
false_targets= df[df.TD == 'FT'].score.values
true_targets= df[df.TD == 'TT'].score.values
minS = int(np.round(np.min(np.append(decoys, np.append(false_targets, true_targets)))))
maxS = int(np.round(np.max(np.append(decoys, np.append(false_targets, true_targets)))))
plt.hist(false_targets, rwidth=.4, bins=range(minS,maxS), range=[minS,maxS], align='mid', label='false targets')
plt.hist(true_targets, rwidth=.4, bins=range(minS,maxS), range=[minS,maxS], align='mid', label='true targets')
plt.hist(decoys, rwidth=.4, bins=range(minS,maxS), range=[minS,maxS], align='left', label='decoys')
plt.legend(loc='best')
plt.title("score histogram")
plt.xlabel("score")
plt.ylabel("count")
plt.xlim(-5,10)
plt.show()
def score_TDcompetition(df):
td_dataframe = pd.DataFrame({'T':df[~df.decoy].score.values, 'D':df[df.decoy].score.values, 'label':df[~df.decoy].TD.values})
td_dataframe['win'] = td_dataframe.apply(lambda x: 'T' if x['T'] > x['D'] else 'D', axis = 1)
target_in = np.where(td_dataframe.win=='T')
decoy_in = np.where(td_dataframe.win=='D')
T_df = df[(~df.decoy) & (np.isin(df.sequence, target_in))]
D_df = df[(df.decoy) & (np.isin(df.sequence, decoy_in))]
df = T_df.append(D_df)
return df
def get_simulated_stat_rates(df, TDcompetition = False, analyte_level='sequence', df_ini = None):
alpha = np.arange(0.002,1,0.002)
stat_rates = pd.DataFrame(columns=['alpha','TP','FP','TN','FN','TPR','FPR','FDR','FNR','ACC'])
if analyte_level=='protein':
df = df.drop_duplicates(subset='protein', keep="first")
for idx in range(len(alpha)):
sig = df[df.q_value <= alpha[idx]]
not_sig = df[df.q_value > alpha[idx]]
TP = len(sig[sig.TD == 'TT'][analyte_level].unique())
FP = len(sig[sig.TD == 'FT'][analyte_level].unique())
TN = len(not_sig[not_sig.TD == 'FT'][analyte_level].unique())
FN = len(not_sig[not_sig.TD == 'TT'][analyte_level].unique())
if TDcompetition:
TN = TN + (len(df_ini[df_ini.TD == 'FT'][analyte_level].unique()) - len(df[df.TD == 'FT'][analyte_level].unique()))
FN = FN + (len(df_ini[df_ini.TD == 'TT'][analyte_level].unique()) - len(df[df.TD == 'TT'][analyte_level].unique()))
TPR = TP/(TP+FN)
FPR = FP/(FP+TN)
if (FP+TP)==0:
FDR = 0
else:
FDR = FP/(FP+TP)
FNR = FN/(FN+TP)
ACC = (TP+TN)/(TP+TN+FP+FN)
stat_rates.loc[idx] = [alpha[idx], TP, FP, TN, FN, TPR, FPR, FDR, FNR, ACC]
border = 0.1
plt.plot([-1,2], [-1,2], linestyle="--", color='red')
plt.scatter(stat_rates.alpha, stat_rates.FDR)
plt.ylim(0-border,1+border)
plt.xlim(0-border,1+border)
plt.title("decoy vs. true FDR")
plt.xlabel("decoy FDR")
plt.ylabel("true FDR")
plt.show()
plt.plot([-1,1], [-1,1], linestyle="--", color='red')
plt.scatter(stat_rates.alpha, stat_rates.FDR)
plt.ylim(0-(0.01),0.1+(0.01))
plt.xlim(0-(0.01),0.1+(0.01))
plt.title("decoy vs. true FDR (zoom)")
plt.xlabel("decoy FDR")
plt.ylabel("true FDR")
plt.show()
plt.plot([-1,2], [1,1], linestyle="--", color='red')
plt.scatter(stat_rates.FPR, stat_rates.TPR)
plt.ylim(0-border,1+border)
plt.xlim(0-border,1+border)
plt.title("ROC curve")
plt.xlabel("FPR")
plt.ylabel("TPR")
plt.show()
plt.plot([-1,2], [1,1], linestyle="--", color='red')
plt.scatter(stat_rates.FPR, stat_rates.TPR)
plt.ylim(0-border,1+border)
plt.xlim(0-0.01,0.1+0.01)
plt.title("ROC curve (zoom)")
plt.xlabel("FPR")
plt.ylabel("TPR")
plt.show()
return stat_rates
def plot_qvalue_vs_fdr(df):
plt.plot(df.fdr, df.target_cum, label='FDR')
plt.plot(df.q_value, df.target_cum, label='q-value')
plt.xlim(0-0.0001,0.005)
plt.ylim(0-100,7000)
plt.legend(loc='best')
plt.title("Difference between q-value and FDR")
plt.xlabel("q-value / FDR")
plt.ylabel("Cummulative number of accepted targets")
plt.show()
# -
#hide
TD = simulateTargetDecoyScores(n=50000, pi0=0.8, target_mu=3.5)
TDC = score_TDcompetition(TD)
# Simulation of random scores for 50'000 measurements (corresponding to spectra). Simulated are decoys, true targets and false targets. We assume a false traget raio (pi0) of 0.8 and a mean score difference of 3.5.
# Simulated score distribution for a separate target and decoy database search:
#hide
plot_score_hist(TD, analyte_level='sequence')
# Simulated score distribution for a corresponding concatinated target-decoy database search with target-decoy-competition:
#hide
plot_score_hist(TDC, analyte_level='sequence')
# Application of the `cut_fdr` function to the simulated target-decoy competition dataset saved in `TDC`:
cval, cut_TDC = cut_fdr(TDC, fdr_level=0.01)
# Evaluation of the FDR estimated by the target-decoy approach versus the true FDR confirms accurate FDR estimation by our approach. The true FDR is capped by the selected fraction of false targets (pi0 = 0.8) and by the effect of target decoy competition. Similarly, the true positive rate (TPR) is limited by the effect of target decoy competition and can only reach 1 in cases where not a single decoy scores higher than a true target.
# hide
cval_, cut_TDC_ = cut_fdr(TDC, fdr_level=100, plot=False)
stat = get_simulated_stat_rates(cut_TDC_, TDcompetition = True, analyte_level='sequence', df_ini = TD)
# The following figure illustrates the difference between `fdr` and `q_value`.
# hide
plot_qvalue_vs_fdr(cut_TDC_)
# Please be aware that the shown simulations are not an accurate model for PSMS scoring and they were designed only for illustrative purposes and to test the implemeted functions.
# ## Global FDR
#
# The `cut_global_fdr` function has two specific applications:
# 1. **Estimate q-values on the peptide and protein level** <br/>
# The concept here is based on selecting the best scoring precursor per peptide (or protein) to then estimate the FDR by target-decoy competition using the `cut_fdr` function.
# 2. **Estimate q-values across an entire dataset on either precursor, peptide or protein level** <br/>
# The concept here is based on selecting the best scoring precursor, peptide or protein signal across an entire dataset to then estimate the FDR by target-decoy competition using the `cut_fdr` function.
#
# This strategy was extensively tested and discussed in the following publications:
#
# * Nesvizhskii, <NAME>. "A survey of computational methods and error rate estimation procedures for peptide and protein identification in shotgun proteomics." Journal of proteomics vol. 73,11 (2010): 2092-123. <https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2956504/>
#
# * Savitski, <NAME> et al. "A Scalable Approach for Protein False Discovery Rate Estimation in Large Proteomic Data Sets." Molecular & cellular proteomics : MCP vol. 14,9 (2015): 2394-404. <https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4563723/>
#
# * The, Matthew et al. "Fast and Accurate Protein False Discovery Rates on Large-Scale Proteomics Data Sets with Percolator 3.0." Journal of the American Society for Mass Spectrometry vol. 27,11 (2016): 1719-1727. <https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5059416/>
#
# * <NAME>, and <NAME>. "False discovery rates of protein identifications: a strike against the two-peptide rule." Journal of proteome research vol. 8,9 (2009): 4173-81.
# <https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3398614/>
# +
#export
def cut_global_fdr(data: pd.DataFrame, analyte_level: str='sequence', fdr_level: float=0.01, plot: bool=True, **kwargs) -> pd.DataFrame:
"""
Function to estimate and filter by global peptide or protein fdr
Args:
data (pd.DataFrame): psms table of search results from alphapept.
analyte_level (str, optional): string specifying the analyte level to apply the fdr threshold. Options include: 'precursor', 'sequence', 'protein_group' and 'protein'. Defaults to 'sequence'.
fdr_level (float, optional): fdr level that should be used for filtering. The value should lie between 0 and 1. Defaults to 0.01.
plot (bool, optional): flag to enable plot. Defaults to 'True'.
Returns:
pd.DataFrame: df with filtered results
"""
logging.info('Global FDR on {}'.format(analyte_level))
data_sub = data[[analyte_level,'score','decoy']]
data_sub_unique = data_sub.groupby([analyte_level,'decoy'], as_index=False).agg({"score": "max"})
analyte_levels = ['precursor', 'sequence', 'protein_group','protein']
if analyte_level in analyte_levels:
agg_score = data_sub_unique.groupby([analyte_level,'decoy'])['score'].max().reset_index()
else:
raise Exception('analyte_level should be either sequence or protein. The selected analyte_level was: {}'.format(analyte_level))
agg_cval, agg_cutoff = cut_fdr(agg_score, fdr_level=fdr_level, plot=plot)
agg_report = pd.merge(data,
agg_cutoff,
how = 'inner',
on = [analyte_level,'decoy'],
suffixes=('', '_'+analyte_level),
validate="many_to_one")
return agg_report
# -
# Similar to the sequence level simulations we can simulatae score distributions for peptides beloning to proteins. In our simulation we assumed a poisson distribution for the number of peptides for each protein centered at 4 peptides.
# +
#hide
TD_prot = simulateProteinLevelTargetDecoyScores(n=8000, pi0=0.8, target_mu=3.5)
TDC_prot = score_TDcompetition(TD_prot)
# -
# Application of the `cut_global_fdr` function to the simulated protein-level target-decoy competition dataset saved in `TDC_prot`:
cut_TDC_prot = cut_global_fdr(TDC_prot, fdr_level=0.01, analyte_level='protein')
# Evaluation of the protein-level FDR estimated by the target-decoy approach versus the true FDR confirms accurate FDR estimation by our approach:
#hide
cut_TDC_prot_ = cut_global_fdr(TDC_prot, fdr_level=100, analyte_level='protein', plot=False)
stat_prot = get_simulated_stat_rates(cut_TDC_prot_, TDcompetition = True, analyte_level='protein', df_ini = TD_prot)
# Investigating the peptide-level statistics after protein-level FDR filtering shows a conservative pattern.
#hide
stat_prot = get_simulated_stat_rates(cut_TDC_prot_, TDcompetition = True, analyte_level='sequence', df_ini = TD_prot)
# Please be aware that the shown simulations are not an accurate model for PSMS scoring and they were designed only for illustrative purposes and to test the implemeted functions.
# ## Scoring
# ### X!tandem scoring
#
# * `get_x_tandem_score` performs scoring of PSMs according to the X!tandem strategy:
#
# * `score_x_tandem` first calls `get_x_tandem_score` and and subsequently applies the `cut_fdr` function to filter PSMs at the specified `fdr_level`.
#
# > X!Tandem, Craig,R. and Beavis,R.C. (2003) Rapid Commun. Mass Spectrom., 17, 2310-2316
# +
#export
import networkx as nx
def get_x_tandem_score(df: pd.DataFrame) -> np.ndarray:
"""
Function to calculate the x tandem score
Args:
df (pd.DataFrame): psms table of search results from alphapept.
Returns:
np.ndarray: np.ndarray with x_tandem scores
"""
b = df['b_hits'].astype('int').apply(lambda x: np.math.factorial(x)).values
y = df['y_hits'].astype('int').apply(lambda x: np.math.factorial(x)).values
x_tandem = np.log(b.astype('float')*y.astype('float')*df['matched_int'].values)
x_tandem[x_tandem==-np.inf] = 0
return x_tandem
def score_x_tandem(df: pd.DataFrame, fdr_level: float = 0.01, plot: bool = True, **kwargs) -> pd.DataFrame:
"""
Filters the psms table by using the x_tandem score and filtering the results for fdr_level.
Args:
df (pd.DataFrame): psms table of search results from alphapept.
fdr_level (float, optional): fdr level that should be used for filtering. The value should lie between 0 and 1. Defaults to 0.01.
Returns:
pd.DataFrame: psms table with an extra 'score' column for x_tandem, filtered for no feature or precursor to be assigned multiple times.
"""
logging.info('Scoring using X-Tandem')
df['score'] = get_x_tandem_score(df)
df['decoy'] = df['sequence'].str[-1].str.islower()
df = filter_score(df)
df = filter_precursor(df)
cval, cutoff = cut_fdr(df, fdr_level, plot)
return cutoff
def filter_with_x_tandem(df: pd.DataFrame) -> pd.DataFrame:
"""
Filters the psms table by using the x_tandem score, no fdr filter.
TODO: Remove redundancy with score functions, see issue: #275
Args:
df (pd.DataFrame): psms table of search results from alphapept.
Returns:
pd.DataFrame: psms table with an extra 'score' column for x_tandem, filtered for no feature or precursor to be assigned multiple times.
"""
logging.info('Filter df with x_tandem score')
df['score'] = get_x_tandem_score(df)
df['decoy'] = df['sequence'].str[-1].str.islower()
df = filter_score(df)
df = filter_precursor(df)
return df
def filter_with_score(df: pd.DataFrame):
"""
Filters the psms table by using the score column, no fdr filter.
TODO: Remove redundancy with score functions, see issue: #275
Args:
df (pd.DataFrame): psms table of search results from alphapept.
Returns:
pd.DataFrame: psms table filtered for no feature or precursor to be assigned multiple times.
"""
logging.info('Filter df with custom score')
df['decoy'] = df['sequence'].str[-1].str.islower()
df = filter_score(df)
df = filter_precursor(df)
return df
# +
#hide
def test_get_x_tandem_score():
y_hits = np.array([1,2,3,0])
b_hits = np.array([0,1,2,1])
matched_int = np.array([1000,1000,1000,1000])
df = pd.DataFrame({'y_hits':y_hits,'b_hits':b_hits,'matched_int':matched_int})
np.testing.assert_almost_equal(get_x_tandem_score(df), np.array([6.90775528, 7.60090246, 9.39266193, 6.90775528]))
test_get_x_tandem_score()
def test_score_x_tandem():
y_hits = np.array([1,2,3,0])
b_hits = np.array([0,1,2,1])
matched_int = np.array([1000,1000,1000,1000])
sequence = np.array(['A','A','B','C_decoy'])
precursor = np.array(['A1','A1','B','C_decoy'])
query_idx = np.array([1,2,3,4])
df = pd.DataFrame({'y_hits':y_hits,'b_hits':b_hits,'matched_int':matched_int,
'sequence':sequence,'precursor':precursor,'query_idx':query_idx})
res = score_x_tandem(df, fdr_level=1, plot=False)
assert all(res.precursor == ['B','A1','C_decoy'])
assert all(res.q_value == [0,0,0.5])
test_score_x_tandem()
# -
# ### Score and filter PSMs by any specified score
#
# `score_psms` uses the specified `score` and applies the `cut_fdr` function to filter PSMs at the specified `fdr_level`. `filter_score` and `filter_precursor` are applied to only report the best PSM per acquired spectrum and the best signal per precursor (i.e. sequence + charge combination).
# +
#export
def score_psms(df: pd.DataFrame, score: str='y_hits', fdr_level: float=0.01, plot: bool=True, **kwargs) -> pd.DataFrame:
"""
Uses the specified score in df to filter psms and to apply the fdr_level threshold.
Args:
df (pd.DataFrame): psms table of search results from alphapept.
score (str, optional): string specifying the column in df to use as score. Defaults to 'y_hits'.
fdr_level (float, optional): fdr level that should be used for filtering. The value should lie between 0 and 1. Defaults to 0.01.
plot (bool, optional): flag to enable plot. Defaults to 'True'.
Returns:
pd.DataFrame: filtered df with psms within fdr
"""
if score in df.columns:
df['score'] = df[score]
else:
raise ValueError("The specified 'score' {} is not available in 'df'.".format(score))
df['decoy'] = df['sequence'].str[-1].str.islower()
df = filter_score(df)
df = filter_precursor(df)
cval, cutoff = cut_fdr(df, fdr_level, plot)
return cutoff
# +
#hide
def test_score_psms():
y_hits = np.array([1,2,3,0])
b_hits = np.array([0,1,2,1])
matched_int = np.array([1000,1000,1000,1000])
sequence = np.array(['A','A','B','C_decoy'])
precursor = np.array(['A1','A1','B','C_decoy'])
query_idx = np.array([1,2,3,4])
df = pd.DataFrame({'y_hits':y_hits,'b_hits':b_hits,'matched_int':matched_int,
'sequence':sequence,'precursor':precursor,'query_idx':query_idx})
res = score_psms(df, fdr_level=1, plot=False)
assert all(res.precursor == ['B','A1','C_decoy'])
assert all(res.q_value == [0,0,0.5])
res = score_psms(df, score='b_hits', fdr_level=1, plot=False)
assert all(res.precursor == ['B','C_decoy','A1'])
assert all(res.q_value == [0,0.5,0.5])
test_score_x_tandem()
# -
# ## Machine learning based scoring of PSMs
#
# * `get_ML_features` extracts additional scoring metrics for the machine learning, including the number of amino acids per precursor, the number of missed cleavages and the logarithmic number of times the same peptide occurs in the set of PSMs
#
# * `train_RF` trains a random forest classifier for scoring all PSMs. For this, we use the scikit-learn library.
# * First, a machine learning pipeline is created including the sklearn `StandardScaler` and `RandomForestClassifier`. The `StandardScaler` is used to standardize all features by removing the mean and scaling to unit variance. For details on the `RandomForestClassifier` see: <https://scikit-learn.org/stable/modules/generated/sklearn.ensemble.RandomForestClassifier.html>.
# * Next, a grid search is initialized for testing the hyperparameter space (`max_depth` and `max_leaf_nodes`) of the random forest classifier by a 5-fold cross-validation using `GridSearchCV`.
# * To train the classifier, we first select a suitable set of PSMSs. This is achieved by an initial scoring and FDR estimation of the input PSMs based on the `ini_score`. Only targets below the `train_fdr_level` cutoff are considered for training the classifier. To ensure a balanced dataset for training (i.e. same number of targets and decoys), the number of PSMs per category is selected to be the minimum of either the number of high scoring targets below the `train_fdr_level` cutoff or the overall number of decoys among the PSMs. `min_train` specifies the minimum number of targets and decoys that should be available.
# * Once a balanced set of PSMs is established, the PSMs are split into a training and test set accoring to the `test_size` argument using `train_test_split` from sklearn while ensuring the PSMs are split in a stratified fashion (i.e. equal number of targets and decoys in both the training and test sets).
# * The grid search and training of the random forest classifier is performed on the training set of PSMs. The `GridSearchCV` returns the classifier which performed best across all cross-validation hold-out sets according to the `scoring` function (classification *'accuracy'* is set as default score). The grid search is parallelize dinto `n_jobs`.
# * Next, the trained classifier is applied to the testing set of PSMs and the test score is reported.
# * If `plot` is enabled, a figure illustrating the weights of each feature is produced.
# * Finally the function returns the trained random forest classifier for subsequent application to the entire set of PSMs or for transfering to a different dataset.
#
# * `score_ML` applies a classifier trained by `train_RF` to a complete set of PSMs. It calls the `cut_fdr` function and filters for the specified `fdr_level`. `filter_score` and `filter_precursor` are applied to only report the best PSM per acquired spectrum and the best signal per precursor (i.e. sequence + charge combination).
# +
#export
import numpy as np
import pandas as pd
import sys
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
from sklearn.ensemble import RandomForestClassifier
from sklearn.pipeline import Pipeline
from sklearn.model_selection import GridSearchCV
import matplotlib.pyplot as plt
from alphapept.fasta import count_missed_cleavages, count_internal_cleavages
def get_ML_features(df: pd.DataFrame, protease: str='trypsin', **kwargs) -> pd.DataFrame:
"""
Uses the specified score in df to filter psms and to apply the fdr_level threshold.
Args:
df (pd.DataFrame): psms table of search results from alphapept.
protease (str, optional): string specifying the protease that was used for proteolytic digestion. Defaults to 'trypsin'.
Returns:
pd.DataFrame: df including additional scores for subsequent ML.
"""
df['decoy'] = df['sequence'].str[-1].str.islower()
df['abs_delta_m_ppm'] = np.abs(df['delta_m_ppm'])
df['naked_sequence'] = df['sequence'].apply(lambda x: ''.join([_ for _ in x if _.isupper()]))
df['n_AA']= df['naked_sequence'].str.len()
df['matched_ion_fraction'] = df['hits']/(2*df['n_AA'])
df['n_missed'] = df['naked_sequence'].apply(lambda x: count_missed_cleavages(x, protease))
df['n_internal'] = df['naked_sequence'].apply(lambda x: count_internal_cleavages(x, protease))
df['x_tandem'] = get_x_tandem_score(df)
return df
def train_RF(df: pd.DataFrame,
exclude_features: list = ['precursor_idx','ion_idx','fasta_index','feature_rank','raw_rank','rank','db_idx', 'feature_idx', 'precursor', 'query_idx', 'raw_idx','sequence','decoy','naked_sequence','target'],
train_fdr_level: float = 0.1,
ini_score: str = 'x_tandem',
min_train: int = 1000,
test_size: float = 0.8,
max_depth: list = [5,25,50],
max_leaf_nodes: list = [150,200,250],
n_jobs: int = -1,
scoring: str = 'accuracy',
plot:bool = False,
random_state: int = 42,
**kwargs) -> (GridSearchCV, list):
"""
Function to train a random forest classifier to separate targets from decoys via semi-supervised learning.
Args:
df (pd.DataFrame): psms table of search results from alphapept.
exclude_features (list, optional): list with features to exclude for ML. Defaults to ['precursor_idx','ion_idx','fasta_index','feature_rank','raw_rank','rank','db_idx', 'feature_idx', 'precursor', 'query_idx', 'raw_idx','sequence','decoy','naked_sequence','target'].
train_fdr_level (float, optional): Only targets below the train_fdr_level cutoff are considered for training the classifier. Defaults to 0.1.
ini_score (str, optional): Initial score to select psms set for semi-supervised learning. Defaults to 'x_tandem'.
min_train (int, optional): Minimum number of psms in the training set. Defaults to 1000.
test_size (float, optional): Fraction of psms used for testing. Defaults to 0.8.
max_depth (list, optional): List of clf__max_depth parameters to test in the grid search. Defaults to [5,25,50].
max_leaf_nodes (list, optional): List of clf__max_leaf_nodes parameters to test in the grid search. Defaults to [150,200,250].
n_jobs (int, optional): Number of jobs to use for parallelizing the gridsearch. Defaults to -1.
scoring (str, optional): Scoring method for the gridsearch. Defaults to'accuracy'.
plot (bool, optional): flag to enable plot. Defaults to 'False'.
random_state (int, optional): Random state for initializing the RandomForestClassifier. Defaults to 42.
Returns:
[GridSearchCV, list]: GridSearchCV: GridSearchCV object with trained RandomForestClassifier. list: list of features used for training the classifier.
"""
if getattr(sys, 'frozen', False):
logging.info('Using frozen pyinstaller version. Setting n_jobs to 1')
n_jobs = 1
features = [_ for _ in df.columns if _ not in exclude_features]
# Setup ML pipeline
scaler = StandardScaler()
rfc = RandomForestClassifier(random_state=random_state) # class_weight={False:1,True:5},
## Initiate scaling + classification pipeline
pipeline = Pipeline([('scaler', scaler), ('clf', rfc)])
parameters = {'clf__max_depth':(max_depth), 'clf__max_leaf_nodes': (max_leaf_nodes)}
## Setup grid search framework for parameter selection and internal cross validation
cv = GridSearchCV(pipeline, param_grid=parameters, cv=5, scoring=scoring,
verbose=0,return_train_score=True,n_jobs=n_jobs)
# Prepare target and decoy df
df['decoy'] = df['sequence'].str[-1].str.islower()
df['target'] = ~df['decoy']
df['score'] = df[ini_score]
dfT = df[~df.decoy]
dfD = df[df.decoy]
# Select high scoring targets (<= train_fdr_level)
df_prescore = filter_score(df)
df_prescore = filter_precursor(df_prescore)
scored = cut_fdr(df_prescore, fdr_level = train_fdr_level, plot=False)[1]
highT = scored[scored.decoy==False]
dfT_high = dfT[dfT['query_idx'].isin(highT.query_idx)]
dfT_high = dfT_high[dfT_high['db_idx'].isin(highT.db_idx)]
# Determine the number of psms for semi-supervised learning
n_train = int(dfT_high.shape[0])
if dfD.shape[0] < n_train:
n_train = int(dfD.shape[0])
logging.info("The total number of available decoys is lower than the initial set of high scoring targets.")
if n_train < min_train:
raise ValueError("There are fewer high scoring targets or decoys than required by 'min_train'.")
# Subset the targets and decoys datasets to result in a balanced dataset
df_training = dfT_high.sample(n=n_train, random_state=random_state).append(dfD.sample(n=n_train, random_state=random_state))
# Select training and test sets
X = df_training[features]
y = df_training['target'].astype(int)
X_train, X_test, y_train, y_test = train_test_split(X.values, y.values, test_size=test_size, random_state=random_state, stratify=y.values)
# Train the classifier on the training set via 5-fold cross-validation and subsequently test on the test set
logging.info('Training & cross-validation on {} targets and {} decoys'.format(np.sum(y_train),X_train.shape[0]-np.sum(y_train)))
cv.fit(X_train,y_train)
logging.info('The best parameters selected by 5-fold cross-validation were {}'.format(cv.best_params_))
logging.info('The train {} was {}'.format(scoring, cv.score(X_train, y_train)))
logging.info('Testing on {} targets and {} decoys'.format(np.sum(y_test),X_test.shape[0]-np.sum(y_test)))
logging.info('The test {} was {}'.format(scoring, cv.score(X_test, y_test)))
feature_importances=cv.best_estimator_.named_steps['clf'].feature_importances_
indices = np.argsort(feature_importances)[::-1][:40]
top_features = X.columns[indices][:40]
top_score = feature_importances[indices][:40]
feature_dict = dict(zip(top_features, top_score))
logging.info(f"Top features {feature_dict}")
# Inspect feature importances
if plot:
import seaborn as sns
g = sns.barplot(y=X.columns[indices][:40],
x = feature_importances[indices][:40],
orient='h', palette='RdBu')
g.set_xlabel("Relative importance",fontsize=12)
g.set_ylabel("Features",fontsize=12)
g.tick_params(labelsize=9)
g.set_title("Feature importance")
plt.show()
return cv, features
def score_ML(df: pd.DataFrame,
trained_classifier: GridSearchCV,
features: list = None,
fdr_level: float = 0.01,
plot: bool = True,
**kwargs) -> pd.DataFrame:
"""
Applies a trained ML classifier to df and uses the ML score to filter psms and to apply the fdr_level threshold.
Args:
df (pd.DataFrame): psms table of search results from alphapept.
trained_classifier (GridSearchCV): GridSearchCV object returned by train_RF.
features (list): list with features returned by train_RF. Defaults to 'None'.
fdr_level (float, optional): fdr level that should be used for filtering. The value should lie between 0 and 1. Defaults to 0.01.
plot (bool, optional): flag to enable plot. Defaults to 'True'.
Returns:
pd.DataFrame: filtered df with psms within fdr
"""
logging.info('Scoring using Machine Learning')
# Apply the classifier to the entire dataset
df_new = df.copy()
df_new['score'] = trained_classifier.predict_proba(df_new[features])[:,1]
df_new = filter_score(df_new)
df_new = filter_precursor(df_new)
cval, cutoff = cut_fdr(df_new, fdr_level, plot)
return cutoff
def filter_with_ML(df: pd.DataFrame,
trained_classifier: GridSearchCV,
features: list = None,
**kwargs) -> pd.DataFrame:
"""
Filters the psms table by using the x_tandem score, no fdr filter.
TODO: Remove redundancy with score functions, see issue: #275
Args:
df (pd.DataFrame): psms table of search results from alphapept.
trained_classifier (GridSearchCV): GridSearchCV object returned by train_RF.
features (list): list with features returned by train_RF. Defaults to 'None'.
Returns:
pd.DataFrame: psms table with an extra 'score' column from the trained_classifier by ML, filtered for no feature or precursor to be assigned multiple times.
"""
logging.info('Filter df with x_tandem score')
# Apply the classifier to the entire dataset
df_new = df.copy()
df_new['score'] = trained_classifier.predict_proba(df_new[features])[:,1]
df_new = filter_score(df_new)
df_new = filter_precursor(df_new)
return df_new
# -
# ## Protein grouping
#
# What is a protein group? A introduction and explanation can be found here [1]:
#
# ```
# The proteome is characterized by a relatively high sequence redundancy. This results from different evolutionary processes and the presence of isoforms. In bottom-up proteomics, this situation leads to the problem that often a peptide cannot be uniquely associated with one protein of origin, which makes it impossible to unambiguously claim the presence of one protein over another. MaxQuant resolves this issue by collapsing all proteins that cannot be distinguished based on the identified peptides into protein groups.
# The rule is that if all peptides of a given protein are a subset of the peptides used for identification of another protein, these proteins will be merged in a protein group. A more complex situation arises when two protein groups are identified with distinct peptides, except for one that is shared between the two. In this case, the two groups cannot be combined, as they contain group-unique peptides and will thus be reported separately in the MaxQuant output table. Depending on the user-defined setting, the shared peptide will not be used for quantification (unique peptides only), or it will be used for the quantification of the protein group with a larger number of associated peptides (unique + razor peptides).
#
# ```
#
# In AlphaPept we employ the following strategy:
# First, we check whether a peptide is proteotypic, meaning that the peptide can only belong to one protein. For peptides that are shared between multiple proteins, we employ a razor approach.
#
# We create a network and add all connections between the peptides and proteins. Then, we extract all connected components, referring to all peptides and proteins that are connected. For a cluster of connected components, we then iterate over all proteins and count the number of peptides that are connected to the particular protein. The protein with the most peptides will then be the razor protein.
#
# We remove this protein and the respective peptides and continue with the extraction from the cluster until no more peptides are present.
#
# For efficient implementation, the proteins and peptides are encoded as indexes. To distinguish proteins from peptides, proteins have a leading 'p'.
#
# * [1] <NAME>., <NAME>. & <NAME>. The MaxQuant computational platform for mass spectrometry-based shotgun proteomics. Nat Protoc 11, 2301–2319 (2016). https://doi.org/10.1038/nprot.2016.136
# +
#export
import networkx as nx
def assign_proteins(data: pd.DataFrame, pept_dict: dict) -> (pd.DataFrame, dict):
"""
Assign psms to proteins.
This function appends the dataframe with a column 'n_possible_proteins' which indicates how many proteins a psm could be matched to.
It returns the appended dataframe and a dictionary `found_proteins` where each protein is mapped to the psms indices.
Args:
data (pd.DataFrame): psms table of scored and filtered search results from alphapept.
pept_dict (dict): dictionary that matches peptide sequences to proteins
Returns:
pd.DataFrame: psms table of search results from alphapept appended with the number of matched proteins.
dict: dictionary mapping psms indices to proteins.
"""
data = data.reset_index(drop=True)
data['n_possible_proteins'] = data['sequence'].apply(lambda x: len(pept_dict[x]))
unique_peptides = (data['n_possible_proteins'] == 1).sum()
shared_peptides = (data['n_possible_proteins'] > 1).sum()
logging.info(f'A total of {unique_peptides:,} unique and {shared_peptides:,} shared peptides.')
sub = data[data['n_possible_proteins']==1]
psms_to_protein = sub['sequence'].apply(lambda x: pept_dict[x])
found_proteins = {}
for idx, _ in enumerate(psms_to_protein):
idx_ = psms_to_protein.index[idx]
p_str = 'p' + str(_[0])
if p_str in found_proteins:
found_proteins[p_str] = found_proteins[p_str] + [str(idx_)]
else:
found_proteins[p_str] = [str(idx_)]
return data, found_proteins
def get_shared_proteins(data: pd.DataFrame, found_proteins: dict, pept_dict: dict) -> dict:
"""
Assign peptides to razor proteins.
Args:
data (pd.DataFrame): psms table of scored and filtered search results from alphapept, appended with `n_possible_proteins`.
found_proteins (dict): dictionary mapping psms indices to proteins
pept_dict (dict): dictionary mapping peptide indices to the originating proteins as a list
Returns:
dict: dictionary mapping peptides to razor proteins
"""
G = nx.Graph()
sub = data[data['n_possible_proteins']>1]
for i in range(len(sub)):
seq, score = sub.iloc[i][['sequence','score']]
idx = sub.index[i]
possible_proteins = pept_dict[seq]
for p in possible_proteins:
G.add_edge(str(idx), 'p'+str(p), score=score)
connected_groups = np.array([list(c) for c in sorted(nx.connected_components(G), key=len, reverse=True)], dtype=object)
n_groups = len(connected_groups)
logging.info('A total of {} ambigious proteins'.format(len(connected_groups)))
#Solving with razor:
found_proteins_razor = {}
for a in connected_groups[::-1]:
H = G.subgraph(a).copy()
shared_proteins = list(np.array(a)[np.array(list(i[0] == 'p' for i in a))])
while len(shared_proteins) > 0:
neighbors_list = []
for node in shared_proteins:
shared_peptides = list(H.neighbors(node))
if node in G:
if node in found_proteins.keys():
shared_peptides += found_proteins[node]
n_neigbhors = len(shared_peptides)
neighbors_list.append((n_neigbhors, node, shared_peptides))
#Check if we have a protein_group (e.g. they share the same everythin)
neighbors_list.sort()
# Check for protein group
node_ = [neighbors_list[-1][1]]
idx = 1
while idx < len(neighbors_list): #Check for protein groups
if neighbors_list[-idx][0] == neighbors_list[-idx-1][0]: #lenght check
if set(neighbors_list[-idx][2]) == set(neighbors_list[-idx-1][2]): #identical peptides
node_.append(neighbors_list[-idx-1][1])
idx += 1
else:
break
else:
break
#Remove the last entry:
shared_peptides = neighbors_list[-1][2]
for node in node_:
shared_proteins.remove(node)
for _ in shared_peptides:
if _ in H:
H.remove_node(_)
if len(shared_peptides) > 0:
if len(node_) > 1:
node_ = tuple(node_)
else:
node_ = node_[0]
found_proteins_razor[node_] = shared_peptides
return found_proteins_razor
def get_protein_groups(data: pd.DataFrame, pept_dict: dict, fasta_dict: dict, decoy = False, callback = None, **kwargs) -> pd.DataFrame:
"""
Function to perform protein grouping by razor approach.
This function calls `assign_proteins` and `get_shared_proteins`.
ToDo: implement callback for solving
Each protein is indicated with a p -> protein index
Args:
data (pd.DataFrame): psms table of scored and filtered search results from alphapept.
pept_dict (dict): A dictionary mapping peptide indices to the originating proteins as a list.
fasta_dict (dict): A dictionary with fasta sequences.
decoy (bool, optional): Defaults to False.
callback (bool, optional): Defaults to None.
Returns:
pd.DataFrame: alphapept results table now including protein level information.
"""
data, found_proteins = assign_proteins(data, pept_dict)
found_proteins_razor = get_shared_proteins(data, found_proteins, pept_dict)
report = data.copy()
assignment = np.zeros(len(report), dtype=object)
assignment[:] = ''
assignment_pg = assignment.copy()
assignment_idx = assignment.copy()
assignment_idx[:] = ''
razor = assignment.copy()
razor[:] = False
if decoy:
add = 'REV__'
else:
add = ''
for protein_str in found_proteins.keys():
protein = int(protein_str[1:])
protein_name = add+fasta_dict[protein]['name']
indexes = [int(_) for _ in found_proteins[protein_str]]
assignment[indexes] = protein_name
assignment_pg[indexes] = protein_name
assignment_idx[indexes] = str(protein)
for protein_str in found_proteins_razor.keys():
indexes = [int(_) for _ in found_proteins_razor[protein_str]]
if isinstance(protein_str, tuple):
proteins = [int(_[1:]) for _ in protein_str]
protein_name = ','.join([add+fasta_dict[_]['name'] for _ in proteins])
protein = ','.join([str(_) for _ in proteins])
else:
protein = int(protein_str[1:])
protein_name = add+fasta_dict[protein]['name']
assignment[indexes] = protein_name
assignment_pg[indexes] = protein_name
assignment_idx[indexes] = str(protein)
razor[indexes] = True
report['protein'] = assignment
report['protein_group'] = assignment_pg
report['razor'] = razor
report['protein_idx'] = assignment_idx
return report
def perform_protein_grouping(data: pd.DataFrame, pept_dict: dict, fasta_dict: dict, **kwargs) -> pd.DataFrame:
"""
Wrapper function to perform protein grouping by razor approach
Args:
data (pd.DataFrame): psms table of scored and filtered search results from alphapept.
pept_dict (dict): A dictionary mapping peptide indices to the originating proteins as a list.
fasta_dict (dict): A dictionary with fasta sequences.
Returns:
pd.DataFrame: alphapept results table now including protein level information.
"""
data_sub = data[['sequence','score','decoy']]
data_sub_unique = data_sub.groupby(['sequence','decoy'], as_index=False).agg({"score": "max"})
targets = data_sub_unique[data_sub_unique.decoy == False]
targets = targets.reset_index(drop=True)
protein_targets = get_protein_groups(targets, pept_dict, fasta_dict, **kwargs)
protein_targets['decoy_protein'] = False
decoys = data_sub_unique[data_sub_unique.decoy == True]
decoys = decoys.reset_index(drop=True)
protein_decoys = get_protein_groups(decoys, pept_dict, fasta_dict, decoy=True, **kwargs)
protein_decoys['decoy_protein'] = True
protein_groups = protein_targets.append(protein_decoys)
protein_groups_app = protein_groups[['sequence','decoy','protein','protein_group','razor','protein_idx','decoy_protein','n_possible_proteins']]
protein_report = pd.merge(data,
protein_groups_app,
how = 'inner',
on = ['sequence','decoy'],
validate="many_to_one")
return protein_report
# +
#hide
def test_get_protein_groups():
pept_dict = {}
pept_dict['seq0'] = [0] #unique
pept_dict['seq1'] = [1] #unique
pept_dict['seq2'] = [2] #unique
pept_dict['seq3'] = [3] #unique
pept_dict['seq4'] = [4] #unique
pept_dict['seq5'] = [5] #unique
pept_dict['seq345'] = [3,4,5] #multiple
pept_dict['seq34'] = [3,4] #multiple
pept_dict['seq45'] = [4,5] #multiple
pept_dict['seq35'] = [3,5] #multiple
fasta_dict = {}
fasta_dict[0] = {'name':'P0'}
fasta_dict[1] = {'name':'P1'}
fasta_dict[2] = {'name':'P2'}
fasta_dict[3] = {'name':'P3'}
fasta_dict[4] = {'name':'P4'}
fasta_dict[5] = {'name':'P5'}
test_case = ['seq0','seq1','seq2','seq3','seq4','seq5']
data = pd.DataFrame({'sequence':test_case, 'score':[1 for _ in test_case]})
res = get_protein_groups(data, pept_dict, fasta_dict)
assert res['razor'].sum() == 0
#sequence 3,4 & 3,5 are present -> P3 will be razor
test_case = ['seq0','seq1','seq2','seq3','seq4','seq5','seq34','seq35']
data = pd.DataFrame({'sequence':test_case, 'score':[1 for _ in test_case]})
res = get_protein_groups(data, pept_dict, fasta_dict)
assert res[res['sequence'] == 'seq34'][['protein', 'razor']].values.tolist()[0] == ['P3', True]
assert res[res['sequence'] == 'seq35'][['protein', 'razor']].values.tolist()[0] == ['P3', True]
#sequence 3,4,5 & 3,4, & 4,5 are present -> P4 will be razor
test_case = ['seq0','seq1','seq2','seq3','seq4','seq5','seq345','seq34','seq45']
data = pd.DataFrame({'sequence':test_case, 'score':[1 for _ in test_case]})
res = get_protein_groups(data, pept_dict, fasta_dict)
assert res[res['sequence'] == 'seq345'][['protein', 'razor']].values.tolist()[0] == ['P4', True]
assert res[res['sequence'] == 'seq34'][['protein', 'razor']].values.tolist()[0] == ['P4', True]
assert res[res['sequence'] == 'seq45'][['protein', 'razor']].values.tolist()[0] == ['P4', True]
# protein group case:
test_case = ['seq35']
data = pd.DataFrame({'sequence':test_case, 'score':[1 for _ in test_case]})
res = get_protein_groups(data, pept_dict, fasta_dict)
assert res[res['sequence'] == 'seq35'][['protein', 'razor']].values.tolist()[0] == ['P5,P3', True]
test_get_protein_groups()
# +
#hide
def test_cut_fdr():
import random
import string
from collections import Counter
# Generate dummy data
n_samples = 10000
test_data = np.random.rand(n_samples)
df = pd.DataFrame(test_data, columns=['score'])
df['decoy'] = (np.random.rand(n_samples) + df['score']) < 0.5
df['filename'] = np.repeat(['file1','file2','file3','file4'], 2500)
sequences = []
i = 0
while i < 5000:
i += 1
sequences.append(''.join(random.choices(string.ascii_uppercase, k=50)))
df['sequence'] = np.random.choice(sequences, 10000, replace=True)
proteins = []
i = 0
while i < 500:
i += 1
proteins.append(''.join(random.choices(string.ascii_uppercase, k=50)))
df['protein'] = np.random.choice(proteins, 10000, replace=True)
for fdr_level in [0.01, 0.02, 0.05, 0.1, 0.2, 0.4]:
cutoff_value, cutoff = cut_fdr(df,fdr_level = fdr_level, plot=False)
assert cutoff.iloc[-1]['fdr'] <= fdr_level
count_fdr = len(cutoff[cutoff.decoy])/len(cutoff[cutoff.target])
assert count_fdr <= fdr_level
sequence_res = cut_global_fdr(df, plot=False)
sequence_count_fdr = len(np.unique(sequence_res[sequence_res.decoy].sequence))/len(np.unique(sequence_res[~ sequence_res.decoy].sequence))
assert len(np.unique(sequence_res.filename)) == 4
assert Counter(sequence_res.sequence).most_common(1)[0][1] > 1
assert sequence_count_fdr <= fdr_level
protein_res = cut_global_fdr(df, analyte_level="protein", plot=False)
protein_count_fdr = len(np.unique(protein_res[protein_res.decoy].protein))/len(np.unique(protein_res[~ protein_res.decoy].protein))
assert len(np.unique(protein_res.filename)) == 4
assert protein_count_fdr <= fdr_level
assert Counter(protein_res.sequence).most_common(1)[0][1] > 1
test_cut_fdr()
# +
#export
ion_dict = {}
ion_dict[0] = ''
ion_dict[1] = '-H20'
ion_dict[2] = '-NH3'
def get_ion(i: int, df: pd.DataFrame, ions: pd.DataFrame)-> (list, np.ndarray):
"""
Helper function to extract the ion-hits for a given DataFrame index.
This function extracts the hit type and the intensities.
E.g.: ['b1','y1'], np.array([10,20]).
Args:
i (int): Row index for the DataFrame
df (pd.DataFrame): DataFrame with PSMs
ions (pd.DataFrame): DataFrame with ion hits
Returns:
list: List with strings that describe the ion type.
np.ndarray: Array with intensity information
"""
start = df['ion_idx'].iloc[i]
end = df['n_ions'].iloc[i]+start
ion = [('b'+str(int(_))).replace('b-','y') for _ in ions.iloc[start:end]['ion_index']]
losses = [ion_dict[int(_)] for _ in ions.iloc[start:end]['ion_type']]
ion = [a+b for a,b in zip(ion, losses)]
ints = ions.iloc[start:end]['ion_int'].astype('int').values
return ion, ints
# -
#hide
def test_get_ion():
df = pd.DataFrame({'ion_idx':[1], 'n_ions':[3]})
ions = pd.DataFrame({'ion_index':[-1,1,-1,1],'ion_type':[0,0,1,2],'ion_int':[1,2,3,4]})
i = 0
ion, ints = get_ion(i, df, ions)
assert ion == ['b1', 'y1-H20', 'b1-NH3']
assert np.allclose(ints, np.array([2,3,4]))
#export
def ecdf(data:np.ndarray)-> (np.ndarray, np.ndarray):
"""Compute ECDF.
Helper function to calculate the ECDF of a score distribution.
This is later used to normalize the score from an arbitrary range to [0,1].
Args:
data (np.ndarray): Array containting the score.
Returns:
np.ndarray: Array containg the score, sorted.
np.ndarray: Noramalized counts.
"""
x = np.sort(data)
n = x.size
y = np.arange(1, n+1) / n
return (x,y)
# +
#hide
def test_ecdf():
x,y = ecdf(np.array([4, 2, 3, 1]))
assert np.allclose(x, np.array([1, 2, 3, 4]))
assert np.allclose(y, np.array([0.25, 0.5 , 0.75, 1. ]))
test_ecdf()
# -
# ## Helper functions
#
# To call the functions from the interface with a process pool, we define the helper functions `score_hdf` and `protein_grouping_all`.
# +
#export
import os
from multiprocessing import Pool
from scipy.interpolate import interp1d
from typing import Callable, Union
#This function has no unit test and is covered by the quick_test
def score_hdf(to_process: tuple, callback: Callable = None, parallel: bool=False) -> Union[bool, str]:
"""Apply scoring on an hdf file to be called from a parallel pool.
This function does not raise errors but returns the exception as a string.
Args:
to_process: (int, dict): Tuple containg a file index and the settings.
callback: (Callable): Optional callback
parallel: (bool): Parallel flag (unused).
Returns:
Union[bool, str]: True if no eo exception occured, the exception if things failed.
"""
try:
index, settings = to_process
file_name = settings['experiment']['file_paths'][index]
base_file_name, ext = os.path.splitext(file_name)
ms_file = base_file_name+".ms_data.hdf"
skip = False
ms_file_ = alphapept.io.MS_Data_File(ms_file, is_overwritable=True)
try:
df = ms_file_.read(dataset_name='second_search')
logging.info('Found second search psms for scoring.')
except KeyError:
try:
df = ms_file_.read(dataset_name='first_search')
logging.info('No second search psms for scoring found. Using first search.')
except KeyError:
df = pd.DataFrame()
if len(df) == 0:
skip = True
logging.info('Dataframe does not contain data. Skipping scoring step.')
if not skip:
df_ = get_ML_features(df, **settings['fasta'])
if settings["score"]["method"] == 'random_forest':
try:
cv, features = train_RF(df)
df = filter_with_ML(df_, cv, features = features)
except ValueError as e:
logging.info('ML failed. Defaulting to x_tandem score')
logging.info(f"{e}")
logging.info('Converting x_tandem score to probabilities')
x_, y_ = ecdf(df_[~df_['decoy']]['score'].values)
f = interp1d(x_, y_, bounds_error = False, fill_value=(y_.min(), y_.max()))
df_['score'] = df_['score'].apply(lambda x: f(x))
df = filter_with_score(df_)
elif settings["score"]["method"] == 'x_tandem':
df = filter_with_x_tandem(df)
else:
try:
import importlib
alphapept_plugin = importlib.import_module(settings["score"]["method"]+".alphapept_plugin")
df = alphapept_plugin.score_alphapept(df, index, settings)
except Exception as e:
raise NotImplementedError('Scoring method {} not implemented. Other exception info: {}'.format(settings["score"]["method"], e))
df = cut_global_fdr(df, analyte_level='precursor', plot=False, fdr_level = settings["search"]["peptide_fdr"], **settings['search'])
logging.info('FDR on peptides complete. For {} FDR found {:,} targets and {:,} decoys.'.format(settings["search"]["peptide_fdr"], df['target'].sum(), df['decoy'].sum()) )
# Insert here
try:
logging.info('Extracting ions')
ions = ms_file_.read(dataset_name='ions')
ion_list = []
ion_ints = []
for i in range(len(df)):
ion, ints = get_ion(i, df, ions)
ion_list.append(ion)
ion_ints.append(ints)
df['ion_int'] = ion_ints
df['ion_types'] = ion_list
logging.info('Extracting ions complete.')
except KeyError:
logging.info('No ions present.')
ms_file_.write(df, dataset_name="peptide_fdr")
logging.info(f'Scoring of file {ms_file} complete.')
return True
except Exception as e:
logging.error(f'Scoring of file {ms_file} failed. Exception {e}')
return f"{e}" #Can't return exception object, cast as string
import alphapept.utils
#This function has no unit test and is covered by the quick_test
def protein_grouping_all(settings:dict, pept_dict:dict, fasta_dict:dict, callback=None):
"""Apply protein grouping on all files in an experiment.
This function will load all dataframes (peptide_fdr level) and perform protein grouping.
Args:
settings: (dict): Settings file for the experiment
pept_dict: (dict): A peptide dictionary.
fast_dict: (dict): A FASTA dictionary.
callback: (Callable): Optional callback.
"""
df = alphapept.utils.assemble_df(settings, field = 'peptide_fdr', callback=None)
if len(df) > 0:
df_pg = perform_protein_grouping(df, pept_dict, fasta_dict, callback = None)
df_pg = cut_global_fdr(df_pg, analyte_level='protein_group', plot=False, fdr_level = settings["search"]["protein_fdr"], **settings['search'])
logging.info('FDR on proteins complete. For {} FDR found {:,} targets and {:,} decoys. A total of {:,} proteins found.'.format(settings["search"]["protein_fdr"], df_pg['target'].sum(), df_pg['decoy'].sum(), len(set(df_pg['protein']))))
path = settings['experiment']['results_path']
base, ext = os.path.splitext(path)
df_pg.to_csv(base+'_protein_fdr.csv')
df_pg.to_hdf(
path,
'protein_fdr'
)
logging.info('Saving complete.')
else:
logging.info('No peptides for grouping present. Skipping.')
# -
#hide
from nbdev.showdoc import *
#hide
from nbdev.export import *
notebook2script()
| nbs/06_score.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="VKFOucyKzOdX"
# # Problem 1:
# Set random seed = 42. Generate a sample of 500 observations of the following random variables: <br>
# $X = 2 + u$, where $u$ is the random float in the range (0,2). <br>
# $Z = 3 - e$, where $e$ is the random number drawn from the standard normal distribution. <br>
# $Y = 5 + 2X + Z + o$, where $o$ is the random noise in the range (0,1). <br>
# Estimate the model: $Y = 5 + 2X + Z + o$
#
# + colab={} colab_type="code" id="DTdOjdWczOdZ"
np.random.seed(42)
# + colab={} colab_type="code" id="NGky7JiFBexs"
# + colab={} colab_type="code" id="wAW2uqvwzOdf"
# + [markdown] colab_type="text" id="_KzKUZmEzOdj"
# # Problem 2:
# Estimate the model from problem 1 Batch Gradient Descent (BGD) with eta = 0.3, 100 iterations, and 500 observations.
# Set starting theta at random number with mean 3 and sd = 1. <br>
# Print a graph of MSE and iterations.
# What do you think is the problem? Is MSE is getting smaller or bigger with each iteration?
# + colab={} colab_type="code" id="q5lK7nOpzOdk"
# + [markdown] colab_type="text" id="XQGPn6D-zOdn"
# The fit is getting worse as we iterate.
# + [markdown] colab_type="text" id="0ovBklhVzOdo"
# # Problem 3
# Let's try two ways to fix the problem. <br>
# 3.A Find smaller eta to reduce MSE. Search from 0.001 to 0.1, using a step of 0.001. What is the optimal eta? What is the MSE with the optimal eta? Plot the graph of eta and MSE around the optimal eta (+5,-5 observations).
#
# + colab={} colab_type="code" id="X_4eRlag9KYB"
np.random.seed(42)
# + colab={} colab_type="code" id="eW4arJKdzOdt"
# + colab={} colab_type="code" id="xpc9_nniXhrv"
# + [markdown] colab_type="text" id="S32XJoT0zOdv"
# # Problem 4:
# Let's try to fix the model fit in Problem 2 by using optimal early stopping. At what iteration should we stop to minimize MSE? What is the value of MSE at this step? Compare our MSE with the one from Problem 3.
# + colab={} colab_type="code" id="3ESXhLhMzOdw"
np.random.seed(42)
# + colab={} colab_type="code" id="O-SYONC8zOdy"
# + [markdown] colab_type="text" id="VZP0bwrPzOd1"
# # Problem 5:
#
# To solve this problem we need to learn how to time the how long it takes to run our code:
# + colab={} colab_type="code" id="JZnjulfCzOd2"
# + [markdown] colab_type="text" id="QhSnTaNuzOd4"
# Create create Y, X, and Z samples from problem 1 with 100,000,000 observations (100 million). Measure how long does it take to estimate linear regression of Y on X and Z using: Stochastic Gradient Descent, Batch Gradient Descent, Linear Regression from sklearn, and Linear regression estimated using matrix multiplication $$\hat{\theta} = (\pmb{X}^T \cdot \pmb{X})^{-1} \cdot \pmb{X}^T \cdot \pmb{y}$$, and Linear regression estimated preudo-inverse. For Batch Gradient Descent use 1000 iterations and eta = 0.01. For Stochastic Gradient Descent use 5 iterations and eta = 0.01. Use SGDRegressior imported from sklearn.model.
#
# For each estimation import **MSE** and **time**
# + colab={} colab_type="code" id="umHo4KqtzOd4"
import numpy as np
np.random.seed(42)
N = 100000000
# + colab={} colab_type="code" id="8qd4avSDzOd6"
from sklearn.linear_model import LinearRegression
# + colab={} colab_type="code" id="-UadqbuZzOd7"
np.random.seed(42)
# + colab={} colab_type="code" id="MOu5Zx59zOd9"
# + colab={} colab_type="code" id="Cpd0rakAzOd_"
# + colab={} colab_type="code" id="EfUx-157zOeA"
from sklearn.linear_model import SGDRegressor
# + [markdown] colab_type="text" id="bJzlwYlCzOeD"
# # Problem 6
# Recreate a X,Y data from problem 1 using 500 observations. Create a 25-degree polynomial for X and scale the data using standard scaler. Estimate the regression model predicting Y via Ridge regression. Calculate MSE (average 'neg_mean_squared_error') using cross-validation with cv = 3. Find optimal alpha by looping from 0.0001 to 1 using step size 0.0001.
# + colab={} colab_type="code" id="1KDDgdA1zOeD"
import numpy as np
np.random.seed(42)
# + colab={} colab_type="code" id="S9eUvNluzOeF"
# + colab={} colab_type="code" id="NXdGbzlpzOeG"
# + colab={} colab_type="code" id="ojWqgmTwwxKm"
# + [markdown] colab_type="text" id="GCFfHfwPzOeI"
# # Problem 7:
# Repeat the problem 6 using Lasso regression. In addition to MSE and alpha, report how many coefficients are equal to zero in the optimal model.
# + colab={} colab_type="code" id="mB5U8ReCzOeI"
# + colab={} colab_type="code" id="AUM-0TyWzOeJ"
# + colab={} colab_type="code" id="5CxkVLCIzOeK"
# + colab={} colab_type="code" id="AQp67KD3zOeM"
# + colab={} colab_type="code" id="rp-lsnenzOeO"
| lesson_4_Training/HW_4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import myutil as mu
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from torch.utils.data import TensorDataset # 텐서데이터셋
from torch.utils.data import DataLoader # 데이터로더
from torch.utils.data import Dataset
# -
# ---
# - 로지스틱 회귀(Logistic Regression)
# - 일상 속 풀고자하는 많은 문제 중에서는 두 개의 선택지 중에서 정답을 고르는 문제가 많습니다.
# - 예를 들어 시험을 봤는데 이 시험 점수가 합격인지 불합격인지가 궁금할 수도 있고, 어떤 메일을 받았을 때 이게 정상 메일인지 스팸 메일인지를 분류하는 문제도 그렇습니다. 이렇게 둘 중 하나를 결정하는 문제를 이진 분류(Binary Classification)라고 합니다. 그리고 이진 분류를 풀기 위한 대표적인 알고리즘으로 로지스틱 회귀(Logistic Regression)가 있습니다.
# - 로지스틱 회귀는 알고리즘의 이름은 회귀이지만 실제로는 분류(Classification) 작업에 사용할 수 있습니다.
#
# +
# -
# ---
# - 시그모이드 함수(Sigmoid function)
# - 위와 같이 S자 형태로 그래프를 그려주는 시그모이드 함수의 방정식은 아래와 같습니다.
# 
# - 선형 회귀에서는 최적의 W와 b를 찾는 것이 목표였습니다.
# - 여기서도 마찬가지입니다.
# - 선형 회귀에서는 W가 직선의 기울기, b가 y절편을 의미했습니다.
# - 그렇다면 여기에서는 W와 b가 함수의 그래프에 어떤 영향을 주는지 직접 그래프를 그려서 알아보겠습니다.
#
# +
# # %matplotlib inline
import numpy as np # 넘파이 사용
import matplotlib.pyplot as plt # 맷플롯립사용
def sigmoid(x):
res = 1 / (1 + np.exp(-x))
return res
x = np.arange(-5.0, 5.0, 0.1)
y = sigmoid(x)
plt.plot(x, y, "g")
plt.plot([0, 0], [1, 0], ":")
plt.title("sigmoid function")
plt.show()
# -
# ---
# - 비용 함수(Cost function)
# - y 의 실제값이 1일 때 −logH(x) 그래프를 사용하고
# - y의 실제값이 0일 때 −log(1−H(X)) 그래프를 사용해야 합니다.
# - 이는 다음과 같이 하나의 식으로 통합할 수 있습니다.
#  = -\frac{1}{n} \sum_{i=1}^{n} [y^{(i)}logH(x^{(i)}) %2B (1-y^{(i)})log(1-H(x^{(i)}))])
#
# +
torch.manual_seed(1)
x_data = [[1, 2], [2, 3], [3, 1], [4, 3], [5, 3], [6, 2]]
y_data = [[0], [0], [0], [1], [1], [1]]
x_train = torch.FloatTensor(x_data)
y_train = torch.FloatTensor(y_data)
W = torch.zeros((2, 1), requires_grad=True)
b = torch.zeros(1, requires_grad=True)
hypothesis = 1 / (1 + torch.exp(-(x_train.matmul(W) + b)))
mu.log("hypothesis", hypothesis)
mu.log("y_train", y_train)
hypothesis = torch.sigmoid(x_train.matmul(W) + b)
mu.log("hypothesis", hypothesis)
mu.log("y_train", y_train)
losses = -(y_train * torch.log(hypothesis)) + (1 - y_train) * torch.log(1 - hypothesis)
cost = losses.mean()
mu.log("losses", losses)
mu.log("cost", cost)
loss = F.binary_cross_entropy(hypothesis, y_train)
mu.log("loss.item()", loss.item())
# -
# ---
# 모델의 훈련 과정까지 추가한 전체 코드는 아래와 같습니다.
#
# +
x_data = [[1, 2], [2, 3], [3, 1], [4, 3], [5, 3], [6, 2]]
y_data = [[0], [0], [0], [1], [1], [1]]
x_train = torch.FloatTensor(x_data)
y_train = torch.FloatTensor(y_data)
W = torch.zeros((2, 1), requires_grad=True)
b = torch.zeros(1, requires_grad=True)
optimizer = optim.SGD([W, b], lr=1)
nb_epoches = 1000
mu.plt_init()
for epoch in range(nb_epoches + 1):
hypothesis = torch.sigmoid(x_train.matmul(W) + b)
cost = -(y_train * torch.log(hypothesis) + (1 - y_train) * torch.log(1 - hypothesis)).mean()
accuracy = mu.get_regression_accuracy(hypothesis, y_train)
optimizer.zero_grad()
cost.backward()
optimizer.step()
if epoch % 100 == 0:
mu.log_epoch(epoch, nb_epoches, cost, accuracy)
mu.plt_show()
mu.log("W", W)
mu.log("b", b)
prediction = hypothesis >= torch.FloatTensor([0.5])
mu.log("prediction", prediction)
mu.log("y_data", y_data)
| 0401_logistic_regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: car-behavioral-cloning-keras2
# language: python
# name: car-behavioral-cloning-keras2
# ---
# ## Fix for Reproducibility
# +
# check
#import os
#if os.environ.get("PYTHONHASHSEED") != "0":
# raise Exception("You must set PYTHONHASHSEED=0 when starting the Jupyter server to get reproducible results.")
# +
import numpy as np
import tensorflow as tf
import random as rn
# The below is necessary in Python 3.2.3 onwards to
# have reproducible behavior for certain hash-based operations.
# See these references for further details:
# https://docs.python.org/3.4/using/cmdline.html#envvar-PYTHONHASHSEED
# https://github.com/fchollet/keras/issues/2280#issuecomment-306959926
import os
os.environ['PYTHONHASHSEED'] = '0'
# The below is necessary for starting Numpy generated random numbers
# in a well-defined initial state.
SEED=42
np.random.seed(SEED)
# The below is necessary for starting core Python generated random numbers
# in a well-defined state.
rn.seed(SEED)
# Force TensorFlow to use single thread.
# Multiple threads are a potential source of
# non-reproducible results.
# For further details, see: https://stackoverflow.com/questions/42022950/which-seeds-have-to-be-set-where-to-realize-100-reproducibility-of-training-res
session_conf = tf.ConfigProto(intra_op_parallelism_threads=1, inter_op_parallelism_threads=1)
from keras import backend as K
# The below tf.set_random_seed() will make random number generation
# in the TensorFlow backend have a well-defined initial state.
# For further details, see: https://www.tensorflow.org/api_docs/python/tf/set_random_seed
tf.set_random_seed(SEED)
sess = tf.Session(graph=tf.get_default_graph(), config=session_conf)
K.set_session(sess)
# -
from keras import __version__ as keras_version
print("keras version: {}".format(keras_version))
from tensorflow import __version__ as tf_version
print("tensorflow version:{}".format(tf_version ))
# ## Get Data and Generate Dataset
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="S-_m1d42Z_Ix" outputId="59301455-6419-453c-e0d5-c08eda2c310a"
# #!wget https://d17h27t6h515a5.cloudfront.net/topher/2016/December/584f6edd_data/data.zip
# #!unzip data.zip
# -
DATASET_PATH="/home/milhouse/datasets/steering_angle/data2"
# !ls $DATASET_PATH
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="ApygA6cKetP8" outputId="7ca7de34-d5fb-4873-990b-f2dcf68e5b39"
#import numpy as np
import matplotlib.pyplot as plt
import matplotlib.image as npimg
import os
## Keras
import keras
from keras.models import Sequential
from keras.optimizers import Adam
from keras.layers import Convolution2D, MaxPooling2D, Dropout, Flatten, Dense
import cv2
import pandas as pd
#import random
import ntpath
## Sklearn
from sklearn.utils import shuffle
from sklearn.model_selection import train_test_split
# + colab={"base_uri": "https://localhost:8080/", "height": 400} colab_type="code" id="-vW2OAepfCY7" outputId="a481b688-42e5-46be-f9c4-c7a66af787c7"
## Store data
#datadir = 'self-driving-car-data-track-1'
datadir = DATASET_PATH
#columns = ['center', 'left', 'right', 'steering', 'throttle', 'reverse', 'speed']
data = pd.read_csv(os.path.join(datadir, 'driving_log.csv'))
pd.set_option('display.max_colwidth', -1)
data.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 226} colab_type="code" id="DPesS6YZhmiR" outputId="0b57fe4a-2f17-4916-adc2-3008cf331dd1"
def path_leaf(path):
"""Get tail of path"""
head, tail = ntpath.split(path)
return tail
## Remove path of images
data['center'] = data['center'].apply(path_leaf)
data['left'] = data['left'].apply(path_leaf)
data['right'] = data['right'].apply(path_leaf)
print(data.shape)
data.head()
# + colab={} colab_type="code" id="mujQNa09pSdD"
def load_img_steering(datadir, df):
"""Get img and steering data into arrays"""
image_path = []
steering = []
for i in range(len(data)):
indexed_data = data.iloc[i]
center, left, right = indexed_data[0], indexed_data[1], indexed_data[2]
image_path.append(os.path.join(datadir, center.strip()))
steering.append(float(indexed_data[3]))
image_paths = np.asarray(image_path)
steerings = np.asarray(steering)
return image_paths, steerings
image_paths, steerings = load_img_steering(datadir + '/IMG', data)
# + colab={"base_uri": "https://localhost:8080/", "height": 333} colab_type="code" id="3vuJ1oMTvnHE" outputId="72584d95-ba75-4632-c6a9-35a54071ed58"
## Split data into training and validation
X_train, X_valid, Y_train, Y_valid = train_test_split(image_paths, steerings, test_size=0.2, random_state=0)
# -
X_train = np.array(list(map(lambda x:npimg.imread(x), X_train)))
X_valid = np.array(list(map(lambda x:npimg.imread(x), X_valid)))
plt.imshow(X_train[rn.randint(0, len(X_train)-1)])
plt.axis('off')
print(X_train.shape)
IMG_SIZE = X_train[0].shape
# ## Nvidia convolutional network. First flavor
# https://github.com/tylerlum/self_driving_car
# + colab={} colab_type="code" id="EMEwn5h50ZxX"
def nvidia_model_tylerlum():
model = Sequential()
model.add(Convolution2D(24, (5, 5), strides=(2, 2), input_shape=IMG_SIZE, activation='elu'))
#model.add(Convolution2D(24, 5, 5, subsample=(2, 2), input_shape=(66, 200, 3), activation='elu'))
model.add(Convolution2D(36, (5, 5), strides=(2, 2), activation='elu'))
model.add(Convolution2D(48, (5, 5), strides=(2, 2), activation='elu'))
model.add(Convolution2D(64, (3, 3), activation='elu'))
model.add(Convolution2D(64, (3, 3), activation='elu'))
model.add(Dropout(0.5))
model.add(Flatten())
model.add(Dense(100, activation='elu'))
model.add(Dropout(0.5))
model.add(Dense(50, activation='elu'))
model.add(Dropout(0.5))
model.add(Dense(10, activation='elu'))
model.add(Dropout(0.5))
model.add(Dense(1))
optimizer = Adam(lr=1e-3)
model.compile(loss='mse', optimizer=optimizer)
return model
# + colab={"base_uri": "https://localhost:8080/", "height": 642} colab_type="code" id="7lEbwjbA2Tt8" outputId="d24cc318-a7d8-401d-86b2-35d1a91e940c"
model = nvidia_model_tylerlum()
print(model.summary())
# + colab={"base_uri": "https://localhost:8080/", "height": 1148} colab_type="code" id="tA0727Hn2ldx" outputId="a9e7758e-e619-49cc-d583-cb3056f9124e"
np.random.seed(SEED)
history = model.fit(X_train, Y_train, epochs=30, validation_data=(X_valid, Y_valid), batch_size=100, verbose=1, shuffle=1)
# -
plt.plot(history.history['loss'][1:])
plt.plot(history.history['val_loss'][1:])
plt.legend(['training', 'validation'])
plt.title('Loss')
plt.xlabel('Epoch')
# 
PROJECT_PATH="/home/milhouse/projects/car-behavioral-cloning/"
# ## Let's try on the simulator
model.save('{}/models/model_meetup.h5'.format(PROJECT_PATH))
# !/home/milhouse/miniconda3/envs/car-behavioral-cloning-keras2/bin/python $PROJECT_PATH/drive_original.py $PROJECT_PATH/models/model_meetup.h5 $PROJECT_PATH/models/images_model_meetup & $PROJECT_PATH/beta_simulator_linux/beta_simulator.x86_64
# !convert $PROJECT_PATH/models/images_model_meetup/*5.jpg $PROJECT_PATH/models/images_model_tylerlum_zero.gif
# <img src="../models/images_model_tylerlum_zero.gif" width="800">
# ## So...
# 
| notebooks/Behavioral_Cloning_with_Keras_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from glob import glob
import pickle
import numpy as np
import sklearn
import matplotlib.pyplot as plt
import sys
import pandas as pd
# +
# Import from adjacent scripts.
import sys
import os
sys.path.append('..')
sys.path.append('../src')
sys.path.append('../audio')
import src.data_util as du
from src.defaults import DATA_PATH, MAX_INTEGER
# +
# Get STRFs from one study type.
study_type = 1
df = du.load_and_clean_data()
df = du.min_max_norm(df)
df = du.isolate_study(df, study_type)
df = df.reset_index(drop=True)
with open('modulation_features.pickle', 'rb') as handle:
tmp = pickle.load(handle)
print(tmp[0]['strf'].shape)
tmp = pd.DataFrame(tmp)
df = pd.merge(tmp, df, on='stimulus')
df['strf'] = df['strf'].transform(lambda x: x.flatten())
# +
from sklearn.linear_model import LogisticRegression
# Flatten STRF representation.
X = df['strf']
X = np.stack(X.to_numpy())
# +
means_ = np.mean(X, axis=0)
stds_ = np.std(X, axis=0)
X = (X - means_)/stds_
# +
# Convert response to binary above/below quantile.
quantile = 0.5
y = df.groupby('subjectNo')['response'].transform(
lambda x: (x < np.quantile(x, q=quantile)).astype(int)
).to_numpy()
# +
reg = LogisticRegression(penalty='l2', max_iter=400).fit(X, y)
print(f"Score: {reg.score(X, y)}")
learned_weights = reg.coef_.reshape(128, 11, 22)
strf_scale_rate = np.mean(learned_weights, axis=0)
strf_freq_rate = np.mean(learned_weights, axis=1)
strf_freq_scale = np.mean(learned_weights, axis=2)
plt.imshow(strf_scale_rate, origin='lower', aspect='auto')
plt.show()
plt.imshow(strf_freq_rate, origin='lower', aspect='auto')
plt.show()
plt.imshow(strf_freq_scale, origin='lower', aspect='auto')
plt.show()
# -
def plotStrfavg(strf_scale_rate, strf_freq_rate, strf_freq_scale,aspect_='auto', interpolation_='none',figname='defaut',show='true'):
rates = [4.0, 5.7, 8.0, 11.3, 16.0, 22.6, 32.0, 45.3, 64.0, 90.5, 128.0]
rates = np.append(-1 * rates[::-1], rates)
scales = [0.25, 0.35, 0.50, 0.71, 1.00, 1.41, 2.00, 2.83, 4.00, 5.66, 8.00]
plt.suptitle(figname, fontsize=10)
plt.subplot(1,3,1)
plt.imshow(strf_scale_rate, aspect=aspect_, interpolation=interpolation_,origin='lower')
plt.xlabel('Rates (Hz)', fontsize=10)
plt.ylabel('Scales (c/o)', fontsize=10)
plt.xticks(np.arange(len(rates)), rates)
plt.yticks([])
plt.subplot(1,3,2)
plt.imshow(strf_freq_rate, aspect=aspect_, interpolation=interpolation_,origin='lower')
plt.xticks(np.arange(len(rates)), rates)
plt.yticks([])
plt.xlabel('Rates (Hz)', fontsize=10)
plt.ylabel('Freq Channel', fontsize=10)
plt.subplot(1,3,3)
plt.imshow(np.transpose(strf_freq_scale), aspect=aspect_, interpolation=interpolation_,origin='lower')
plt.xticks([])
plt.yticks([])
plt.ylabel('Scales (c/o)', fontsize=10)
plt.xlabel('Freq Channel', fontsize=10)
plt.savefig(figname+'.png')
if show=='true':
plt.show()
plotStrfavg(strf_scale_rate, strf_freq_rate, strf_freq_scale, aspect_='auto', interpolation_='sinc',figname='defaut',show='true')
| data/strf_learn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: irl_python3
# language: python
# name: irl_python3
# ---
# # Navigation MDP [1]
# +
import numpy as np
from simple_rl.tasks import NavigationMDP
from simple_rl.agents import QLearningAgent
from simple_rl.planning import ValueIteration
from simple_rl.tasks.grid_world.GridWorldStateClass import GridWorldState
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
# -
np.random.seed(0)
nvmdp = NavigationMDP(width=30,
height=30,
goal_locs=[(21,21)],
init_loc=(1,1),
rand_init=True,
cell_types=['white', 'yellow', 'red', 'lime', 'magenta'],
cell_type_rewards=[0, 0, -10, -10, -10],
goal_reward=1.,
slip_prob=0.00,
step_cost=0.0,
vacancy_prob=0.85,
gamma=.9999)
N_tau = 8
# Use "init_states" to request specific init states while sampling trajectories
# if "n_trajectory" is greater than # of init_states, remaining init states will be sampled randomly
D_traj_states, D_traj_actions = nvmdp.sample_data(n_trajectory=N_tau, init_states=[GridWorldState(1,1)], init_repetition=False)
nvmdp.visualize_grid(trajectories=D_traj_states, show_rewards_cbar=True)
# ## Features used for short horizon
[nvmdp.feature_long_at_state(s, normalize=False) for s in D_traj_states[0]]
# ## Features used for long horizon
[nvmdp.feature_short_at_state(s) for s in D_traj_states[0]]
# ## Reference
#
# [1] MacGlashan, James, and <NAME>. "Between Imitation and Intention Learning." IJCAI. 2015.
| simple_rl/tasks/navigation/Usage_NavigationMDP.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: forecasting_env
# language: python
# name: forecasting_env
# ---
# <i>Copyright (c) Microsoft Corporation.</i>
#
# <i>Licensed under the MIT License.</i>
# # Orange Juice Data Exploration in Python
#
# In this notebook, we use Python to explore the Orange Juice (OJ) dataset. The OJ dataset is from R package [bayesm](https://cran.r-project.org/web/packages/bayesm/index.html) and is part of the [Dominick's dataset](https://www.chicagobooth.edu/research/kilts/datasets/dominicks).
#
# ## Dataset description
#
# This dataset contains the following two tables:
#
# - **yx.csv** - Weekly sales of refrigerated orange juice at 83 stores. This table has 106139 rows and 19 columns. It includes weekly sales and prices of 11 orange juice
# brands as well as information about profit, deal, and advertisement for each brand. Note that the weekly sales is captured by a column named `logmove` which
# corresponds to the natural logarithm of the number of units sold. To get the number of units sold, you need to apply an exponential transform to this column.
#
# - **storedemo.csv** - Demographic information on those stores. This table has 83 rows and 13 columns. For every store, the table describes demographic information of its consumers,
# distance to the nearest warehouse store, average distance to the nearest 5 supermarkets, ratio of its sales to the nearest warehouse store, and ratio of its sales
# to the average of the nearest 5 stores.
#
# Note that the week number starts from 40 in this dataset, while the full Dominick's dataset has week number from 1 to 400. According to [Dominick's Data Manual](https://www.chicagobooth.edu/-/media/enterprise/centers/kilts/datasets/dominicks-dataset/dominicks-manual-and-codebook_kiltscenter.aspx), week 1 starts on 09/14/1989.
# Please see pages 40 and 41 of the [bayesm reference manual](https://cran.r-project.org/web/packages/bayesm/bayesm.pdf) and the [Dominick's Data Manual](https://www.chicagobooth.edu/-/media/enterprise/centers/kilts/datasets/dominicks-dataset/dominicks-manual-and-codebook_kiltscenter.aspx) for more details about the data.
#
#
# ## Global setting and imports
# To run this notebook, please ensure to run through the `SETUP.md` guide in the top-level directory, which will create and activate `forecasting_env` conda environment, as well as, register that environment as a Jupyter kernel.
#
# You can then launch the Jupyter notebook by running `jupyter notebook` and select the kernel named `forecasting_env` in the list of kernels under Kernel tab.
# +
# import packages
import os
import math
import pandas as pd
import numpy as np
import itertools
import matplotlib.pyplot as plt
import statsmodels.api as sm
from fclib.common.utils import git_repo_path
from fclib.dataset.ojdata import download_ojdata
# -
# ## Dataset
#
# Let's read in the OJ dataset and look at what it contains.
# +
# Use False if you've already downloaded the data
DOWNLOAD_DATA = True
# Data directory
DATA_DIR = os.path.join(git_repo_path(), "ojdata")
if DOWNLOAD_DATA:
download_ojdata(DATA_DIR)
print("Data download completed. Data saved to " + DATA_DIR)
# +
sales_file = os.path.join(DATA_DIR, "yx.csv")
store_file = os.path.join(DATA_DIR, "storedemo.csv")
sales = pd.read_csv(sales_file, index_col=False)
storedemo = pd.read_csv(store_file, index_col=False)
# -
# First few rows of sales data
sales.head(5)
# First few rows of store demographic data
storedemo.head(5)
# Check number of time series and lengths
print("Number of stores is {}.".format(len(sales.groupby(["store"]).groups.keys())))
print("Number of brands is {}.".format(len(sales.groupby(["brand"]).groups.keys())))
print("Number of time series is {}.".format(len(sales.groupby(["store", "brand"]).groups.keys())))
print("\nLenth distribution of the time series:")
print(sales.groupby(["store", "brand"]).size().describe())
# +
# Fill missing gaps
store_list = sales["store"].unique()
brand_list = sales["brand"].unique()
week_list = range(sales["week"].min(), sales["week"].max() + 1)
item_list = list(itertools.product(store_list, brand_list, week_list))
item_df = pd.DataFrame.from_records(item_list, columns=["store", "brand", "week"])
print(f"Total number of rows before filling gaps is {len(sales)}.")
sales = item_df.merge(sales, how="left", on=["store", "brand", "week"])
print(f"Total number of rows after filling gaps is {len(sales)}.")
# Fill the missing `logmove` with zeros
sales["logmove"] = sales["logmove"].fillna(value=0)
# Merge sales and store demographics
sales = sales.merge(storedemo, how="left", left_on="store", right_on="STORE")
# Compute unit sales for later analysis
sales["move"] = sales["logmove"].apply(lambda x: round(math.exp(x)) if x > 0 else 0)
# -
# ## Visualize sample time series
#
# We look at some examples of weekly sales time series for sample store and brand.
# Plot sample time series of sales
sample_store = 2
sample_brand = 1
sales_sub = sales.loc[(sales["store"] == sample_store) & (sales["brand"] == sample_brand)]
plt.plot(sales_sub["week"], sales_sub["move"])
plt.plot(
sales_sub["week"].loc[sales_sub["move"] > 0],
sales_sub["move"].loc[sales_sub["move"] > 0],
linestyle="",
marker="o",
color="red",
)
plt.gcf().autofmt_xdate()
plt.xlabel("week")
plt.ylabel("move")
plt.title(f"Weekly sales of store {sample_store} brand {sample_brand} \n missing values are filled with zero")
plt.grid(True)
plt.show()
# +
# Plot sales of all brands in a sample store
sample_store = 2
brand_list = sales.loc[(sales["store"] == sample_store), "brand"].unique()
fig, axes = plt.subplots(nrows=3, ncols=4, figsize=(15, 15))
print("Weekly sales of all brands in store {}.".format(sample_store))
brand_num = 0
for row in axes:
for col in row:
if brand_num < len(brand_list):
brand = brand_list[brand_num]
sales_sub = sales.loc[(sales["store"] == sample_store) & (sales["brand"] == brand)]
col.plot(sales_sub["week"], sales_sub["move"])
col.set_ylim(0, 150000)
col.set_title("brand {}".format(brand))
col.set_xlabel("week")
col.set_ylabel("move")
brand_num += 1
else:
col.axis("off")
plt.tight_layout()
# -
# ## Impact of demographics, brand, and store
#
# In this section, we plot the boxplot of the sales across different stores, brands and different values of the demographics variables. There are variations of the sales observed across these variables. We also observed through our modeling experience that once we included the store and brand variables, the contribution of the demographic variables seems to be limited.
# +
print("Correlation between unit sales and each demographic feature:")
print(sales[storedemo.columns[1:]].corrwith(sales["move"]))
print("\nCorrelation between log-scale sales and each demographic feature:")
print(sales[storedemo.columns[1:]].corrwith(sales["logmove"]))
# -
# Plot box plot of logmove vs. store demographic columns
for cl in storedemo.columns[1:]:
p = sales.loc[sales["logmove"] != 0].boxplot(column="logmove", by=cl)
p.set_title("logmove by {}".format(cl), linespacing=3)
p.set_xlabel(cl)
p.set_ylabel("logmove")
p.get_xaxis().set_ticks([])
plt.suptitle("")
# Plot box plot of logmove across different brands and stores
for by_cl in ["brand", "store"]:
p = sales.loc[sales["logmove"] != 0].boxplot(column="logmove", by=by_cl)
p.set_title("logmove by {}".format(by_cl), linespacing=3)
p.set_xlabel(by_cl)
p.set_ylabel("logmove")
p.get_xaxis().set_ticks([])
plt.suptitle("")
# ## Check seasonality and autocorrelation
#
# Overall, we don't find a strong seasonality in the data. It seems that there is a weak yearly-seasonality according to the seasonal-trend-level decomposition and the autocorrelation values around the lag of 52 weeks. As a rough estimate, autocorrelation beyond 20 weeks is usually very small.
# Check Seasonality
# Not much seasonality is found
d = sales.loc[(sales["store"] == 2) & (sales["brand"] == 1)].copy()
decom = sm.tsa.seasonal_decompose(d["move"].values, freq=52)
print(decom.plot())
# autocorrealtion: weekly, monthly, quarterly, yearly
def single_autocorr(series, lag):
"""
Autocorrelation for single data series
:param series: traffic series
:param lag: lag, days
:return:
"""
s1 = series[lag:]
s2 = series[:-lag]
ms1 = np.mean(s1)
ms2 = np.mean(s2)
ds1 = s1 - ms1
ds2 = s2 - ms2
divider = np.sqrt(np.sum(ds1 * ds1)) * np.sqrt(np.sum(ds2 * ds2))
return np.sum(ds1 * ds2) / divider if divider != 0 else 0
# +
sample_store = 2
brand_list = sales["brand"].unique()
l_range = list(range(1, 53))
for j in range(len(brand_list)):
brand = brand_list[j]
d = sales.loc[(sales["store"] == sample_store) & (sales["brand"] == brand)].copy()
cor = []
for l in l_range:
cor.append(single_autocorr(d["logmove"].values, l))
l_range.insert(0, 0)
cor.insert(0, 1)
plt.scatter(list(l_range), cor)
plt.plot(list(l_range), cor)
plt.title("time series for autocorrelation for store {} brand {}".format(sample_store, brand))
plt.xlabel("lag")
plt.ylabel("autocorrelation")
plt.show()
# -
# ## Impact of promotional information: deal and feat
#
# We find that deal column has a very significant impact on sales. The impact of feat column also looks strong although the pattern shown in the scatter plot is a bit noisy.
# Check the impact of deal, feat by plotting logmove vs feat and deal
# These two features significantly impact the sales
plt.scatter(sales["feat"], sales["logmove"])
plt.title("logmove vs feat")
plt.xlabel("feat")
plt.ylabel("logmove")
p = sales.boxplot(column="logmove", by="deal")
plt.suptitle("")
p.set_title("logmove by deal", linespacing=3)
p.set_xlabel("deal")
p.set_ylabel("logmove")
# ## Impact of price
#
# We find that the sales does typically decrease when the absolute price or the relative price of the product increases.
# correlation between the sales and price, sales and relative price
sales["price"] = sales.apply(lambda x: x.loc["price" + str(int(x.loc["brand"]))], axis=1)
price_cols = [
"price1",
"price2",
"price3",
"price4",
"price5",
"price6",
"price7",
"price8",
"price9",
"price10",
"price11",
]
sales["avg_price"] = sales[price_cols].sum(axis=1).apply(lambda x: x / len(price_cols))
sales["price_ratio"] = sales.apply(lambda x: x["price"] / x["avg_price"], axis=1)
plt.scatter(sales["price"], sales["logmove"])
plt.title("logmove vs price")
plt.xlabel("price")
plt.ylabel("logmove")
plt.show()
plt.scatter(sales["price_ratio"], sales["logmove"])
plt.title("logmove vs price_ratio")
plt.xlabel("price ratio")
plt.ylabel("logmove")
plt.show()
| examples/grocery_sales/python/01_prepare_data/ojdata_exploration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/", "height": 71} colab_type="code" executionInfo={"elapsed": 40178, "status": "ok", "timestamp": 1588213047201, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10844282398210252241"}, "user_tz": -420} id="rPwL9bdoBNzQ" outputId="553f83f0-cbf1-48d5-a184-4f4c8ff055ac"
import os
import re
import PIL
import sys
import json
import time
import timm
import math
import copy
import torch
import pickle
import logging
import fnmatch
import argparse
import torchvision
import numpy as np
# %matplotlib inline
import pandas as pd
import seaborn as sns
import torch.nn as nn
from PIL import Image
from pathlib import Path
from copy import deepcopy
from sklearn import metrics
import torch.optim as optim
from datetime import datetime
from torchvision import models
import matplotlib.pyplot as plt
import torch.nn.functional as F
import torch.utils.data as data
from torch.autograd import Variable
from tqdm import tqdm, tqdm_notebook
from torch.optim import lr_scheduler
#from pytorch_metric_learning import loss
import torch.utils.model_zoo as model_zoo
from timm.models.layers.activations import *
# %config InlineBackend.figure_format = 'retina'
from collections import OrderedDict, defaultdict
from torch.utils.tensorboard import SummaryWriter
from torchvision import transforms, models, datasets
from torch.utils.data.sampler import SubsetRandomSampler
from randaugment import RandAugment, ImageNetPolicy, Cutout
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
from timm.loss import LabelSmoothingCrossEntropy, SoftTargetCrossEntropy
from sklearn.metrics import confusion_matrix,accuracy_score, classification_report
# + colab={"base_uri": "https://localhost:8080/", "height": 102} colab_type="code" executionInfo={"elapsed": 179460, "status": "ok", "timestamp": 1588213186502, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10844282398210252241"}, "user_tz": -420} id="yyGpxuktB96O" outputId="584ea32f-dbe1-4465-8e60-e0f4e5c96a6f"
data_dir = '/home/linh/Downloads/Brain/'
# Define your transforms for the training and testing sets
data_transforms = {
'train': transforms.Compose([
transforms.RandomRotation(30),
transforms.RandomResizedCrop(224),
transforms.RandomHorizontalFlip(),
RandAugment(),
ImageNetPolicy(),
Cutout(size=16),
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406],
[0.229, 0.224, 0.225])
]),
'val': transforms.Compose([
transforms.Resize(256),
transforms.CenterCrop(224),
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406],
[0.229, 0.224, 0.225])
])
}
# Load the datasets with ImageFolder
image_datasets = {x: datasets.ImageFolder(os.path.join(data_dir, x),
data_transforms[x])
for x in ['train', 'val']}
batch_size = 70
data_loader = {x: torch.utils.data.DataLoader(image_datasets[x], batch_size=batch_size,
shuffle=True, num_workers=4, pin_memory = True)
for x in ['train', 'val']}
dataset_sizes = {x: len(image_datasets[x]) for x in ['train', 'val']}
class_names = image_datasets['train'].classes
print(class_names)
print(dataset_sizes)
print(device)
### we get the class_to_index in the data_Set but what we really need is the cat_to_names so we will create
_ = image_datasets['train'].class_to_idx
cat_to_name = {_[i]: i for i in list(_.keys())}
print(cat_to_name)
# Run this to test the data loader
images, labels = next(iter(data_loader['val']))
images.size()
# + colab={"base_uri": "https://localhost:8080/", "height": 603} colab_type="code" executionInfo={"elapsed": 226470, "status": "ok", "timestamp": 1588213233519, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10844282398210252241"}, "user_tz": -420} id="N350JAHpu8c3" outputId="96a2d095-f78f-4ca5-eb0c-c5390e367831"
"""def showimage(data_loader, number_images, cat_to_name):
dataiter = iter(data_loader)
images, labels = dataiter.next()
images = images.numpy() # convert images to numpy for display
# plot the images in the batch, along with the corresponding labels
fig = plt.figure(figsize=(number_images, 4))
for idx in np.arange(number_images):
ax = fig.add_subplot(2, number_images/2, idx+1, xticks=[], yticks=[])
img = np.transpose(images[idx])
plt.imshow(img)
ax.set_title(cat_to_name[labels.tolist()[idx]])
#### to show some images
showimage(data_loader['test'], 20, cat_to_name)"""
# + colab={"base_uri": "https://localhost:8080/", "height": 71} colab_type="code" executionInfo={"elapsed": 226461, "status": "ok", "timestamp": 1588213233520, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10844282398210252241"}, "user_tz": -420} id="L9jdFtBjSAE6" outputId="f0f393c5-4369-422c-9aef-fc290ccc941d"
#model = models.resnet50(pretrained=True)
#model = timm.create_model('resnet50', pretrained=True)
model = timm.create_model('tresnet_xl', pretrained=True)
#model.fc #show fully connected layer for ResNet family
model.head #show the classifier layer (fully connected layer) for EfficientNets
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" executionInfo={"elapsed": 226454, "status": "ok", "timestamp": 1588213233520, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10844282398210252241"}, "user_tz": -420} id="w6QP4CFPBNzg" outputId="6beb0600-5fdf-4ae6-a216-40c32a13bb9f"
# Create classifier
for param in model.parameters():
param.requires_grad = True
# define `classifier` for ResNet
# Otherwise, define `fc` for EfficientNet family
#because the definition of the full connection/classifier of 2 CNN families is differnt
classifier = nn.Sequential(OrderedDict([('fc1', nn.Linear(2048, 1000, bias=True)),
('BN1', nn.BatchNorm2d(1000, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)),
('dropout1', nn.Dropout(0.7)),
('fc2', nn.Linear(1000, 512)),
('BN2', nn.BatchNorm2d(512, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)),
('swish1', Swish()),
('dropout2', nn.Dropout(0.5)),
('fc3', nn.Linear(512, 128)),
('BN3', nn.BatchNorm2d(128, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)),
('swish2', Swish()),
('fc4', nn.Linear(128, 4)),
('output', nn.Softmax(dim=1))
]))
# connect base model (EfficientNet_B0) with modified classifier layer
model.fc = classifier
criterion = LabelSmoothingCrossEntropy()
#criterion = nn.CrossEntropyLoss()
#optimizer = Nadam(model.parameters(), lr=0.001)
#optimizer = optim.Adam(model.parameters(), lr=0.001, weight_decay=0.0001)
optimizer = optim.SGD(model.parameters(),
lr=0.01,momentum=0.9,
nesterov=True,
weight_decay=0.0001)
scheduler = optim.lr_scheduler.StepLR(optimizer, step_size=80, gamma=0.1)
#show our model architechture and send to GPU
def count_parameters(model):
return sum(p.numel() for p in model.parameters() if p.requires_grad)
count = count_parameters(model)
print(count)
# + colab={} colab_type="code" id="iPNx-TodPpVA"
model.to(device)
def train_model(model, criterion, optimizer, scheduler, num_epochs=200, checkpoint = None):
since = time.time()
if checkpoint is None:
best_model_wts = copy.deepcopy(model.state_dict())
best_loss = math.inf
best_acc = 0.
else:
print(f'Val loss: {checkpoint["best_val_loss"]}, Val accuracy: {checkpoint["best_val_accuracy"]}')
model.load_state_dict(checkpoint['model_state_dict'])
best_model_wts = copy.deepcopy(model.state_dict())
optimizer.load_state_dict(checkpoint['optimizer_state_dict'])
scheduler.load_state_dict(checkpoint['scheduler_state_dict'])
best_loss = checkpoint['best_val_loss']
best_acc = checkpoint['best_val_accuracy']
# Tensorboard summary
writer = SummaryWriter()
for epoch in range(num_epochs):
print('Epoch {}/{}'.format(epoch + 1, num_epochs)) #(epoch, num_epochs -1)
print('-' * 20)
# Each epoch has a training and validation phase
for phase in ['train', 'val']:
if phase == 'train':
model.train() # Set model to training mode
else:
model.eval() # Set model to evaluate mode
running_loss = 0.0
running_corrects = 0
# Iterate over data.
for i, (inputs, labels) in enumerate(data_loader[phase]):
inputs = inputs.to(device)
labels = labels.to(device)
# zero the parameter gradients
optimizer.zero_grad()
if i % 1000 == 999:
print('[%d, %d] loss: %.8f' %
(epoch + 1, i, running_loss / (i * inputs.size(0))))
# forward
# track history if only in train
with torch.set_grad_enabled(phase == 'train'):
outputs = model(inputs)
_, preds = torch.max(outputs, 1)
loss = criterion(outputs, labels)
# backward + optimize only if in training phase
if phase == 'train':
loss.backward()
optimizer.step()
# statistics
running_loss += loss.item() * inputs.size(0)
running_corrects += torch.sum(preds == labels.data)
if phase == 'train':
scheduler.step()
epoch_loss = running_loss / dataset_sizes[phase]
epoch_acc = running_corrects.double() / dataset_sizes[phase]
print('{} Loss: {:.8f} Acc: {:.8f}'.format(
phase, epoch_loss, epoch_acc))
# Record training loss and accuracy for each phase
if phase == 'train':
writer.add_scalar('Train/Loss', epoch_loss, epoch)
writer.add_scalar('Train/Accuracy', epoch_acc, epoch)
writer.flush()
else:
writer.add_scalar('Valid/Loss', epoch_loss, epoch)
writer.add_scalar('Valid/Accuracy', epoch_acc, epoch)
writer.flush()
# deep copy the model
if phase == 'val' and epoch_acc > best_acc:
print(f'New best model found!')
print(f'New record ACC: {epoch_acc}, previous record acc: {best_acc}')
best_loss = epoch_loss
best_acc = epoch_acc
best_model_wts = copy.deepcopy(model.state_dict())
torch.save({'model_state_dict': model.state_dict(),
'optimizer_state_dict': optimizer.state_dict(),
'best_val_loss': best_loss,
'best_val_accuracy': best_acc,
'scheduler_state_dict' : scheduler.state_dict(),
},
CHECK_POINT_PATH
)
print(f'New record acc is SAVED: {epoch_acc}')
print()
time_elapsed = time.time() - since
print('Training complete in {:.0f}m {:.0f}s'.format(
time_elapsed // 60, time_elapsed % 60))
print('Best val Acc: {:.8f} Best val loss: {:.8f}'.format(best_acc, best_loss))
# load best model weights
model.load_state_dict(best_model_wts)
return model, best_loss, best_acc
# + colab={"base_uri": "https://localhost:8080/", "height": 85} colab_type="code" id="vcXkJFOlP4NJ" outputId="e47fadb8-c292-4051-8a56-bbdc5868abe8"
CHECK_POINT_PATH = '/home/linh/Downloads/Brain/weights/TResNet_Extra_Large.pth'
try:
checkpoint = torch.load(CHECK_POINT_PATH)
print("checkpoint loaded")
except:
checkpoint = None
print("checkpoint not found")
if checkpoint == None:
CHECK_POINT_PATH = CHECK_POINT_PATH
model, best_val_loss, best_val_acc = train_model(model,
criterion,
optimizer,
scheduler,
num_epochs = 300,
checkpoint = None #torch.load(CHECK_POINT_PATH)
)
torch.save({'model_state_dict': model.state_dict(),
'optimizer_state_dict': optimizer.state_dict(),
'best_val_loss': best_val_loss,
'best_val_accuracy': best_val_acc,
'scheduler_state_dict': scheduler.state_dict(),
}, CHECK_POINT_PATH)
# -
| notebooks/TResNet_XL.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### New to Plotly?
# Plotly's Python library is free and open source! [Get started](https://plot.ly/python/getting-started/) by downloading the client and [reading the primer](https://plot.ly/python/getting-started/).
# <br>You can set up Plotly to work in [online](https://plot.ly/python/getting-started/#initialization-for-online-plotting) or [offline](https://plot.ly/python/getting-started/#initialization-for-offline-plotting) mode, or in [jupyter notebooks](https://plot.ly/python/getting-started/#start-plotting-online).
# <br>We also have a quick-reference [cheatsheet](https://images.plot.ly/plotly-documentation/images/python_cheat_sheet.pdf) (new!) to help you get started!
# #### Version Check
# Plotly's python package is updated frequently. Run `pip install plotly --upgrade` to use the latest version.
import plotly
plotly.__version__
# #### Overwriting existing graphs and updating a graph at its unique URL
# By default, Plotly will overwrite files made with the same filename. For example, if a graph named 'my plot' already exists in your account, then it will be overwritten with this new version and the URL of the graph will persist.
# +
import plotly.plotly as py
import plotly.graph_objs as go
data = [
go.Scatter(
x=[1, 2],
y=[3, 4]
)
]
plot_url = py.plot(data, filename='my plot')
# -
# #### Saving to a folder
# Filenames that contain `"/"` be treated as a Plotly directory and will be saved to your Plotly account in a folder tree. For example, to save your graphs to the folder `my-graphs` use the `filename = "my-graphs/my plot"` (if it doesn't already exist it will be created)
# +
import plotly.plotly as py
import plotly.graph_objs as go
data = [
go.Scatter(
x=[1, 2],
y=[3, 4]
)
]
plot_url = py.plot(data, filename='my-graphs/my plot')
# -
# #### Creating new files
# With `fileopt='new'`, Plotly will always create a new file. If a file with the same name already exists, then Plotly will append a '(1)' to the end of the filename, e.g. `new plot (1)` and create a unique URL.
# +
import plotly.plotly as py
import plotly.graph_objs as go
data = [
go.Scatter(
x=[1, 2],
y=[3, 4]
)
]
plot_url = py.plot(data, filename='new plot', fileopt='new')
# -
# #### Extending traces in an existing graph
# To extend existing traces with your new data, use `fileopt='extend'`.
# +
import plotly.plotly as py
import plotly.graph_objs as go
trace0 = go.Scatter(
x=[1, 2],
y=[1, 2]
)
trace1 = go.Scatter(
x=[1, 2],
y=[2, 3]
)
trace2 = go.Scatter(
x=[1, 2],
y=[3, 4]
)
data = [trace0, trace1, trace2]
# Take 1: if there is no data in the plot, 'extend' will create new traces.
plot_url = py.plot(data, filename='extend plot', fileopt='extend')
# -
# Then, extend the traces with more data.
# +
import plotly.plotly as py
import plotly.graph_objs as go
trace0 = go.Scatter(
x=[3, 4],
y=[2, 1]
)
trace1 = go.Scatter(
x=[3, 4],
y=[3, 2]
)
trace2 = go.Scatter(
x=[3, 4],
y=[4, 3]
)
data = [trace0, trace1, trace2]
# Take 2: extend the traces on the plot with the data in the order supplied.
py.iplot(data, filename='extend plot', fileopt='extend')
# +
from IPython.display import display, HTML
display(HTML('<link href="//fonts.googleapis.com/css?family=Open+Sans:600,400,300,200|Inconsolata|Ubuntu+Mono:400,700" rel="stylesheet" type="text/css" />'))
display(HTML('<link rel="stylesheet" type="text/css" href="http://help.plot.ly/documentation/all_static/css/ipython-notebook-custom.css">'))
# ! pip install git+https://github.com/plotly/publisher.git --upgrade
import publisher
publisher.publish(
'fileopts.ipynb', 'python/file-options/', 'Updating Plotly Graphs',
'How to update your graphs in Python with the fileopt parameter.',
title = 'Python Filenames Options | Plotly',
has_thumbnail='true',
thumbnail='thumbnail/horizontal-bar.jpg',
language='python',
page_type='example_index',
display_as='file_settings',
order=3,
#ipynb='~notebook_demo/1'
)
# -
| _posts/python/fundamentals/fileopt/fileopts.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import warnings; warnings.filterwarnings('ignore')
# %matplotlib inline
from sklearn.datasets import load_boston
from sklearn.datasets import load_diabetes
import sys
sys.path.append('..')
from korr import pearson, corr_vs_pval
# -
# ## Example 1 (Boston Dataset)
# Load the demo dataset
data = load_boston()
cols = [0,1,2,4,5,6,7,8,9,10,11,12]
x = data.data[:,cols]
# Compute Pearson correlations and its p-values.
r, pval = pearson(x)
# Display the results in one histogram.
# The default `rlim=0.4` is usually ok.
# Play with `plim` argument until three colored categories emerge.
corr_vs_pval(r, pval, plim=0.0001, dpi=120);
# The correlation coefficients among the variables of the Boston dataset exhibit low p-values.
# I set the threshold at `0.0001` (or `0.01%`) and still very few correlation coefficients had higher p-values (orange bar).
# ## Interpretations
#
# **Unrelated variables as x predictors (orange)**.
# Two unrelated variables will yield higher p-values, i.e. its correlation coefficient has any value just by chance, or resp. its correlation coefficient is meaningless or even misleading.
# Such a result can be actually helpful. If the p-Value of two variables' correlation coefficient is very high, then both variables are favorable candidates as explanatory variables (or predictors, features) in a multivariate linear regression model or resp. any other supervised learning algorithm.
#
# **Candiates for y and x (green)**.
# As a rule of thumb a correlation coefficient above $|r|>0.4$ can be considered useful.
# If the accompanying p-value is below the accepted threshold (e.g. `plim=0.0001` or `0.01%`) then these high correlations can be considered significant. The use case would be to take such a pair of significant highly correlated variables as `y` and `x` variables in a linear regression model (or resp . any other supervised learning algorithm).
#
# **Dangerous Results (gray)**.
# There is not point of using variable pairs with significant low correlation coefficient (e.g. $|r|\leq 0.4$ and p-value less than `0.0001`) if you have other significantly highly correlated pairs (green) or unrelated pairs (orange).
# ## Example 2 (Diabetes Dataset)
# Load the demo dataset, compute the correlations and its coefficients, display everything in a histogram.
x = load_diabetes().data
r, pval = pearson(x)
corr_vs_pval(r, pval, plim=0.001, rlim=0.4, dpi=120);
# The diabetes dataset is provided for regression tasks by the sklearn package.
# As we only considered predictor variables `x`, the interpretation is as follows
#
# * low p-value (orange): Start with these predictor pairs for a multivariate regression model.
# * high rho, high p-value (green): These predictors might cause multicollinarity issue when used in the same multivariate regression model. Consider dimensionality reduction (e.g. PCA) to combine these predictors
# * gray: Try to add these predictors lateron (e.g. forward variable selection)
| examples/corr_vs_pval.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # `ipywidgets` Interactive Demo
#
# Simple demonstration of rendering Interactive widgets in a `jupyterlite` notebook.
#
# `ipywidgets` can be installed in this deployment (it provides the @jupyter-widgets/jupyterlab-manager federated extension), but you will need to make your own deployment to have access to other interactive widgets libraries.
# +
import micropip
await micropip.install('ipywidgets')
# -
from ipywidgets import IntSlider
slider = IntSlider()
slider
slider
slider.value
slider.value = 5
from ipywidgets import IntText, link
text = IntText()
text
link((slider, 'value'), (text, 'value'));
# +
from ipywidgets import interact
from IPython.display import Markdown, display
@interact(cookies=slider)
def cookies(cookies=slider.value, calories=(0,150)):
total_calories = calories * cookies
if cookies:
display(Markdown(f"If each cookie contains _{calories} calories_, _{cookies} cookies_ contain **{total_calories} calories**!"))
else:
display(Markdown(f"No cookies!"))
if total_calories > 2000:
display(Markdown(f"> Maybe that's too many cookies..."))
| examples/pyolite - interactive-widgets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a href="https://colab.research.google.com/github/Tessellate-Imaging/monk_v1/blob/master/study_roadmaps/4_image_classification_zoo/Classifier%20-%20Auto%20tag%20gala%20Hackerearth%20Dataset%20.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# # Monk Library - https://github.com/Tessellate-Imaging/monk_v1¶
#
# ### Monk is an opensource low-code tool for computer vision and deep learning
#
#
# ## Monk features
#
# low-code
# unified wrapper over major deep learning framework - keras, pytorch, gluoncv
# syntax invariant wrapper
#
#
# ## Enables
#
# to create, manage and version control deep learning experiments
# to compare experiments across training metrics
# to quickly find best hyper-parameters
#
#
# ## To contribute to Monk AI or Pytorch RoadMap repository raise an issue in the git-repo or dm us on linkedin
#
# Abhishek - https://www.linkedin.com/in/abhishek-kumar-annamraju/
# Akash - https://www.linkedin.com/in/akashdeepsingh01/
# # Competition
# - https://www.hackerearth.com/challenges/competitive/hackerearth-deep-learning-challenge-auto-tag-images-gala/machine-learning/auto-tag-images-of-the-gala-9e47fb31/
#
#
# ### Score achieved with this tutorial: 83.54047
#
#
# ### Monk team's final score: 85.785
# # Table of contents
#
#
# ## Install Monk
#
#
# ## Using pretrained model for adding tags to gala imagery
#
#
# ## Training a classifier from scratch
# # Install Monk
#
# - git clone https://github.com/Tessellate-Imaging/monk_v1.git
#
# - cd monk_v1/installation/Linux && pip install -r requirements_cu9.txt
# (Select the requirements file as per OS and CUDA version)
# ! git clone https://github.com/Tessellate-Imaging/monk_v1.git
# +
# If using Colab install using the commands below
# ! cd monk_v1/installation/Misc && pip install -r requirements_colab.txt
# If using Kaggle uncomment the following command
# #! cd monk_v1/installation/Misc && pip install -r requirements_kaggle.txt
# Select the requirements file as per OS and CUDA version when using a local system or cloud
# #! cd monk_v1/installation/Linux && pip install -r requirements_cu9.txt
# -
# # Used trained classifier for demo
# +
# Import monk
import os
import sys
sys.path.append("monk_v1/monk/");
# -
# +
# Download trained weights
# -
# ! wget --load-cookies /tmp/cookies.txt "https://docs.google.com/uc?export=download&confirm=$(wget --save-cookies /tmp/cookies.txt --keep-session-cookies --no-check-certificate 'https://docs.google.com/uc?export=download&id=1cHoSP0UJK3sI5oPWtWMoDpdbpwy2wo5z' -O- | sed -rn 's/.*confirm=([0-9A-Za-z_]+).*/\1\n/p')&id=1cHoSP0UJK3sI5oPWtWMoDpdbpwy2wo5z" -O cls_autogala_trained.zip && rm -rf /tmp/cookies.txt
# ! unzip -qq cls_autogala_trained.zip
# ls workspace/Project-Autogala
# Gluon project
from gluon_prototype import prototype
# +
# Load project in inference mode
gtf = prototype(verbose=1);
gtf.Prototype("Project-Autogala", "Gluon-resnet50_v2", eval_infer=True);
#Other trained models - uncomment
#gtf.Prototype("Project-Autogala", "Gluon-resnet101_v2", eval_infer=True);
#gtf.Prototype("Project-Autogala", "Gluon-resnet152_v2", eval_infer=True);
# -
# +
#Infer
# -
img_name = "workspace/test/1.jpg"
predictions = gtf.Infer(img_name=img_name);
from IPython.display import Image
Image(filename=img_name)
img_name = "workspace/test/2.jpg"
predictions = gtf.Infer(img_name=img_name);
from IPython.display import Image
Image(filename=img_name)
img_name = "workspace/test/3.jpg"
predictions = gtf.Infer(img_name=img_name);
from IPython.display import Image
Image(filename=img_name)
# # Training custom classifier from scratch
# # Goals
# - Train a classifier on an expanded, manually augmented dataset
# - Use the final model from the first classifier as a pretrained model for the seond classifier
# - Train this second classifier on original training set
# - Compare the experiments
# - Run inference to generate test script
#
# # Table of contents
#
#
# ## [0. Install Monk](#0)
#
#
# ## [1. Train on Augmented Set](#1)
#
#
# ## [2. Copy the experiment and train on Original Set](#2)
#
#
# ## [3. Compare the three experiments](#3)
#
#
# ## [4. Run inference using final model from 2nd experiment](#4)
# <a id='0'></a>
# # Install Monk
#
# - git clone https://github.com/Tessellate-Imaging/monk_v1.git
#
# - cd monk_v1/installation && pip install -r requirements_cu9.txt
# - (Select the requirements file as per OS and CUDA version)
# !git clone https://github.com/Tessellate-Imaging/monk_v1.git
# +
# If using Colab install using the commands below
# !cd monk_v1/installation/Misc && pip install -r requirements_colab.txt
# If using Kaggle uncomment the following command
# #!cd monk_v1/installation/Misc && pip install -r requirements_kaggle.txt
# Select the requirements file as per OS and CUDA version when using a local system or cloud
# #!cd monk_v1/installation/Linux && pip install -r requirements_cu9.txt
# -
# # Dataset
# - Credits: https://www.hackerearth.com/challenges/competitive/hackerearth-deep-learning-challenge-auto-tag-images-gala/machine-learning/auto-tag-images-of-the-gala-9e47fb31/
# ! wget --load-cookies /tmp/cookies.txt "https://docs.google.com/uc?export=download&confirm=$(wget --save-cookies /tmp/cookies.txt --keep-session-cookies --no-check-certificate 'https://docs.google.com/uc?export=download&id=1wUEvnf_x4hpCBXrdbSEFeG-Gyv_fOP3a' -O- | sed -rn 's/.*confirm=([0-9A-Za-z_]+).*/\1\n/p')&id=1wUEvnf_x4hpCBXrdbSEFeG-Gyv_fOP3a" -O gala_dataset.zip && rm -rf /tmp/cookies.txt
# ! unzip -qq gala_dataset.zip
# <a id='1'></a>
# # Experiment - 1 - On manually augmented data
import os
import sys
sys.path.append("monk_v1/monk/")
# +
# Using Mxnet Backend
from gluon_prototype import prototype
# To use pytorch backend
#from pytorch_prototype import prototype
# To use keras backend
#from keras_prototype import prototype
# -
# ## Creating and managing experiments
# - Provide project name
# - Provide experiment name
# - For a specific data create a single project
# - Inside each project multiple experiments can be created
# - Every experiment can be have diferent hyper-parameters attached to it
gtf = prototype(verbose=1);
gtf.Prototype("Project", "Experiment-On-Augmented-Data");
# ## This creates files and directories as per the following structure
#
#
# workspace
# |
# |--------Project (Project name can be different)
# |
# |
# |-----Experiment-On-Augmented-Data
# |
# |-----experiment-state.json
# |
# |-----output
# |
# |------logs (All training logs and graphs saved here)
# |
# |------models (all trained models saved here)
# ## Load the data and the model
#
# - Docs on quick mode loading of data and model: https://github.com/Tessellate-Imaging/monk_v1#4
#
# - Tutorials on Monk: https://github.com/Tessellate-Imaging/monk_v1/tree/master/study_roadmaps/1_getting_started_roadmap
gtf.Default(dataset_path="dataset/Train_Images_Foldered",
model_name="resnet152_v2",
freeze_base_network=False,
num_epochs=10);
# ### Docs on how to update hyper-parameters
#
# - Update hyperparams using update mode - https://clever-noyce-f9d43f.netlify.com/#/update_mode/update_dataset
#
# - Tutorials on how to update hyper-params - https://github.com/Tessellate-Imaging/monk_v1/tree/master/study_roadmaps/1_getting_started_roadmap/5_update_hyperparams
# +
gtf.update_batch_size(12)
gtf.optimizer_sgd(0.01);
gtf.lr_multistep_decrease([7, 9]);
gtf.update_save_intermediate_models(False)
# Very important to reload post updates
gtf.Reload();
# -
# ## Train
gtf.Train();
# <a id='2'></a>
# # Experiment - 2 - Further training on original data
from gluon_prototype import prototype
# +
## Copy the previous experiment to use the final model of experiment 1 as pretrained model in experiment 2
# -
gtf = prototype(verbose=1);
gtf.Prototype("Project", "Experiment-On-Original-Data",
copy_from=["Project", "Experiment-On-Augmented-Data"]);
# ## This creates files and directories as per the following structure
#
#
# workspace
# |
# |--------Project (Project name can be different)
# |
# |
# |-----Experiment-On-Augmented-Data (Already created)
# |
# |-----experiment-state.json
# |
# |-----output
# |
# |------logs (All training logs and graphs saved here)
# |
# |------models (all trained models saved here)
# |
# |
# |-----Experiment-On-Original-Data (Created Now)
# |
# |-----experiment-state.json
# |
# |-----output
# |
# |------logs (All training logs and graphs saved here)
# |
# |------models (all trained models saved here)
# +
gtf.update_dataset(dataset_path="dataset/Train_Images",
path_to_csv="dataset/train.csv");
# Very important to reload post Updates
gtf.Reload();
# -
# +
gtf.update_batch_size(16)
gtf.optimizer_sgd(0.001);
gtf.lr_multistep_decrease([8, 9]);
gtf.update_save_intermediate_models(False)
# Very important to reload post updates
gtf.Reload();
# -
# ## Train
gtf.Train();
# <a id='3'></a>
# # Compare experiments
# Invoke the comparison class
from compare_prototype import compare
# ### Creating and managing comparison experiments
# - Provide project name
# Create a project
gtf = compare(verbose=1);
gtf.Comparison("Analysis");
# ### This creates files and directories as per the following structure
#
# workspace
# |
# |--------comparison
# |
# |
# |-----Analysis
# |
# |------stats_best_val_acc.png
# |------stats_max_gpu_usage.png
# |------stats_training_time.png
# |------train_accuracy.png
# |------train_loss.png
# |------val_accuracy.png
# |------val_loss.png
#
# |
# |-----comparison.csv (Contains necessary details of all experiments)
# ## Add experiments
gtf.Add_Experiment("Project", "Experiment-On-Original-Data");
gtf.Add_Experiment("Project", "Experiment-On-Augmented-Data");
# ## Run Analysis
gtf.Generate_Statistics();
# ## Visualize comparisons
# ### Training Accuracy Curves
from IPython.display import Image
Image(filename="workspace/comparison/Analysis/train_accuracy.png")
# ### Training Loss Curves
from IPython.display import Image
Image(filename="workspace/comparison/Analysis/train_loss.png")
# ### Validation Accuracy Curves
from IPython.display import Image
Image(filename="workspace/comparison/Analysis/val_accuracy.png")
# ### Validation loss curves
from IPython.display import Image
Image(filename="workspace/comparison/Analysis/val_loss.png")
# ### Training time curves
from IPython.display import Image
Image(filename="workspace/comparison/Analysis/stats_training_time.png")
# ### Best Validation accuracies
from IPython.display import Image
Image(filename="workspace/comparison/Analysis/stats_best_val_acc.png")
# <a id='4'></a>
# # Inferencing on Experiment - 2
# +
from gluon_prototype import prototype
gtf = prototype(verbose=0);
# -
# To load experiment in evaluation mode, set eval_infer can be set as True
gtf.Prototype("Project", "Experiment-On-Original-Data", eval_infer=True);
import pandas as pd
df = pd.read_csv("dataset/test.csv");
# +
columns = list(df.columns)
combined = [];
from tqdm import tqdm_notebook as tqdm
for i in tqdm(range(len(df))):
img_name = df[columns[0]][i];
#Monk inference Engine
predictions = gtf.Infer(img_name="dataset/Test_Images/" + img_name, return_raw=False);
combined.append([img_name, predictions['predicted_class']])
# -
df = pd.DataFrame(combined, columns = ['Image', 'Class'])
df.to_csv("submission.csv", index=False)
| study_roadmaps/4_image_classification_zoo/Classifier - Auto tag gala Hackerearth Dataset .ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1 Short Answer
# #### 1 False. MV optimization aims at minimizing the overall portfolio volatility given the expected return.
#
# #### 2 True. We believe the overall market goes up in the long run.
#
# #### 3 With an intercept. The regressor doesn't include T-bill. We'll get a better estimate with an intercept.
#
# #### 4 Yes. HDG has high correlaion with HFRI both in-sample and out of sample.
#
# #### 5 The residual may have high information ratio although having a negative alpha.
# # 2 Allocation
import pandas as pd
import numpy as np
import statsmodels.api as sm
from statsmodels.regression.rolling import RollingOLS
#from sklearn.linear_model import LinearRegression
import matplotlib.pyplot as plt
import seaborn as sns
import warnings
warnings.filterwarnings("ignore")
# #### (a)
factor_data = pd.read_excel('../data/proshares_analysis_data.xlsx', sheet_name = 'merrill_factors')
factor_data = factor_data.set_index('date')
column = ['SPY US Equity', 'EEM US Equity', 'EFA US Equity',
'EUO US Equity', 'IWM US Equity']
for i in column:
factor_data[i] = factor_data[i] - factor_data['USGG3M Index']
y = factor_data[['USGG3M Index']]
x = factor_data[['SPY US Equity', 'EEM US Equity', 'EFA US Equity', 'EUO US Equity', 'IWM US Equity']]
x.head()
y.head()
def compute_tangency(df_tilde, diagonalize_Sigma=False):
"""Compute tangency portfolio given a set of excess returns.
Also, for convenience, this returns the associated vector of average
returns and the variance-covariance matrix.
Parameters
----------
diagonalize_Sigma: bool
When `True`, set the off diagonal elements of the variance-covariance
matrix to zero.
"""
Sigma = df_tilde.cov()
# N is the number of assets
N = Sigma.shape[0]
Sigma_adj = Sigma.copy()
if diagonalize_Sigma:
Sigma_adj.loc[:,:] = np.diag(np.diag(Sigma_adj))
mu_tilde = df_tilde.mean()
Sigma_inv = np.linalg.inv(Sigma_adj)
weights = Sigma_inv @ mu_tilde / (np.ones(N) @ Sigma_inv @ mu_tilde)
# For convenience, I'll wrap the solution back into a pandas.Series object.
omega_tangency = pd.Series(weights, index=mu_tilde.index)
return omega_tangency, mu_tilde, Sigma_adj
# +
omega_tangency, mu_tilde, Sigma = compute_tangency(x)
omega_tangency.to_frame('Tangency Weights')
# -
# #### (b)
# +
def target_mv_portfolio(df_tilde, target_return=0.02, diagonalize_Sigma=False):
"""Compute MV optimal portfolio, given target return and set of excess returns.
Parameters
----------
diagonalize_Sigma: bool
When `True`, set the off diagonal elements of the variance-covariance
matrix to zero.
"""
omega_tangency, mu_tilde, Sigma = compute_tangency(df_tilde, diagonalize_Sigma=diagonalize_Sigma)
Sigma_adj = Sigma.copy()
if diagonalize_Sigma:
Sigma_adj.loc[:,:] = np.diag(np.diag(Sigma_adj))
Sigma_inv = np.linalg.inv(Sigma_adj)
N = Sigma_adj.shape[0]
delta_tilde = ((np.ones(N) @ Sigma_inv @ mu_tilde)/(mu_tilde @ Sigma_inv @ mu_tilde)) * target_return
omega_star = delta_tilde * omega_tangency
return omega_star, mu_tilde, Sigma_adj
omega_star, mu_tilde, Sigma = target_mv_portfolio(x)
omega_star_df = omega_star.to_frame('MV Portfolio Weights')
omega_star_df
# -
# #### (c)
# +
def portfolio_stats(omega, mu_tilde, Sigma, annualize_fac):
# Mean
mean = (mu_tilde @ omega) * annualize_fac
# Volatility
vol = np.sqrt(omega @ Sigma @ omega) * np.sqrt(annualize_fac)
# Sharpe ratio
sharpe_ratio = mean / vol
return round(pd.DataFrame(data = [mean, vol, sharpe_ratio],
index = ['Mean', 'Volatility', 'Sharpe'],
columns = ['Portfolio Stats']), 4)
portfolio_stats(omega_star, mu_tilde, Sigma, 12)
# -
# #### (d)
x_2018 = x.loc[:'2019-01-01']
x_2021 = x.loc['2019-01-01':]
# +
omega_star, mu_tilde, Sigma = target_mv_portfolio(x_2018)
omega_star_df = omega_star.to_frame('MV Portfolio Weights')
omega_star_df
# -
r_2021 = x_2021@omega_star_df
r_2021.head()
r_2021.mean()*12 #mean
r_2021.std()*np.sqrt(12) #volatility
r_2021.mean()*12/(r_2021.std()*np.sqrt(12)) #sharpe ratio
# #### (d) commodities would be better as they have more stable price.
# # 3
# #### (a)
from sklearn.linear_model import LinearRegression
y = factor_data['EEM US Equity']
X = factor_data['SPY US Equity']
static_model_noint = sm.OLS(y,X).fit()
static_model_noint.params #buy -0.92566 SPY
# #### (b)
r = factor_data['EEM US Equity'] - 0.92566 * factor_data['SPY US Equity']
r.head()
r.mean()*12
r.std()*np.sqrt(12)
r.mean()*12 / (r.std()*np.sqrt(12))
# #### (c) No. Because it hedged out the SPY return.
# #### (d) Because SPY and IWM have high similarity.
# # 4
factor_data = pd.read_excel('../data/proshares_analysis_data.xlsx', sheet_name = 'merrill_factors')
factor_data = factor_data.set_index('date')
EFA = factor_data['EFA US Equity']
EFA.head()
val_rolling = EFA.shift(1).dropna().rolling(60).apply(lambda x: x.std()*np.sqrt(12))
val_rolling
val_rolling.iloc[-1]
VaR = val_rolling.iloc[-1]*phi(1)
| solutions/mid1/submissions/zhengxing_173045_6241862_Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <center><h1>My Home Nutrition Analysis</h1><center>
# <img src='https://www.fda.gov/files/nfl-howtounderstand-labeled.png' height=500 width=500 >
# ## Objective:
#
# Over the past few years, I have started paying more attention to my nutrition since nutrition is a large part in maintaining a healthy lifestyle. With this analysis I want to observe how the nutrition of foods within my home varies. In addition to our household foods, I collected some nutritional data for fast food items we commonly enjoy.
#
# ### About The Data:
#
# I collected the data by reading the nutrition fact labels on food items within my house. For nutrition on whole fruits and fast food items I took the nutrition from the company's web site.
#
# Each item contains its macronutrients and micronutrients with the measures given from the nutrition label. Some items contain a value of `0.99` which is a placeholder I used if the nutrition label said **'< 1g'**. Some items were given a value of zero in certain micronutrients where the micronutrient was labeled with a percentage as opposed to an actual measurement.
#
# Not all of the foods in my house were used for this data, I mainly collected the nutrition facts for foods and drinks we regularly consume since there are some items in our pantry which never get touched.
#
# **NOTE:**
# I use the term **"Caution Zone"** to describe foods which I feel should should be consumed cautiously. This is not a definitive categorization since most foods can be consumed in moderation and just because an item did not meet the caution criteria does not mean it is exempt from moderation. The values for the cautious zones I used correspond to being 16-20% of my personal daily intake which I determined from a macronutrient calculator which is linked at the end of this notebook.
#
# ## Analysis
#
# I will begin by importing the modules, importing the excel file with the data, and making sure there's no missing data or incorrect data types
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import plotly as py
import plotly.express as px
from plotly.subplots import make_subplots
import plotly.graph_objects as go
df = pd.read_excel('https://github.com/a-camarillo/nutrition-analysis/blob/master/data/nutrition_facts.xlsx?raw=true')
df.head()
df.isna().sum()
df.dtypes
# There are no missing values and all of the data types are as expected so now I'm going to do some quick cleaning of the column names to make things a little easier
"hide_cell"
df.columns = map(str.lower, df.columns)
df.columns = [column.replace(' ','_') for column in df.columns]
df.columns = [column.replace('(grams)','_g') for column in df.columns]
df.columns = [column.replace('(milligrams)','_mg') for column in df.columns]
df.columns = [column.replace('(micrograms)','_mcg') for column in df.columns]
"hide_cell"
df.columns
"hide_cell"
#One more rename to deal with sneaky folate
df.rename(columns={'folate(micrograms':'folate_mcg'},inplace=True)
df.columns
# Before I begin visualizing the data, I am going to create a function for normalization allowing for another comparison of each food item.
def per_100g(Series):
''' Pass in a macronutrient series and find it's value per 100 grams
for each item '''
value = (Series/df['serving_measurement_g']) * 100
return value
# ## Fat
#
# First macronutrient I want analyze is total fat, I will begin by adding a column for fat per 100 grams, and looking at some of the top results
"hide_cell"
fat = df[['food_item','serving_size','serving_measurement_g','total_fat_g']]
"hide_cell"
fat.head(2)
"hide_cell"
fat.insert(loc=len(fat.columns),column='fat_per_100g',value=per_100g(fat['total_fat_g']))
"hide_cell"
#create a 'caution' column for foods which contain high fat content
fat.insert(loc=len(fat.columns),column='caution',value='No')
for row in fat.index:
if fat.at[row,'total_fat_g'] >= 10:
fat.at[row,'caution'] = 'Yes'
"hide_cell"
fat_20 = fat.nlargest(20,'fat_per_100g',keep='all')
"hide_input"
plt.figure(figsize=(8,5))
sns.barplot(y='food_item',x='fat_per_100g',data=fat_20)
plt.title('Top 20 Items by Fat Per 100g')
# Much to my surprise, Best Foods Mayonnaise and Skippy Peanut Butter are considerably high in fat per 100 grams. As someone who frequently consumes both of these products I will definitely have to monitor my intake to avoid having too much fat in my diet.
#
# Another surprise high contender for me is the Ritz Crackers, it's hard to not eat an entire pack of these in one sitting but I might have to reconsider next time a craving hits.
#
# Among the top 20 items for fat per 100 grams, the expected fast food items are there but much lower than some of the household items.
#
# I want to also look at how the fat per 100 grams compares to a single serving since I am curious to see if the same items are as fatty relative to serving size.
"hide_input"
fat_fig = px.scatter(fat,x='serving_measurement_g',y='fat_per_100g',hover_data=['food_item'],
labels = {'serving_measurement_g':'Single Serving In Grams','fat_per_100g':'Total Fat Per 100 Grams'},
title='Serving Size vs. Fat Per 100 Grams(Interactive Plot)',
color='caution',
color_discrete_map={'No':'Blue','Yes':'Orange'})
fat_fig.show()
#
# The above plot compares each item's single serving to its respective total fat per 100 grams. Some takeaways from this plot are:
#
# **Assuming you adhere to proper serving sizes**, Ritz Crackers and the Sabra Hummus are not as fattening as the previous plot might have indicated. Due to each having a small serving size relative to fat per 100 grams, the actual fat per serving becomes relatively small(about 5g each).
#
# Lucerne Cheese Blend is also not as bad as the fat per 100 grams alone might have indicated, however it should still be consumed cautiously since the fat for a single serving is still about.
#
# **THE CAUTION ZONE:**
# I am considering the caution zone(for total fat) to be foods that are shown to have high fat content per serving(greater than or equal to 10g). These can easily be identified as the items around the 10g mark for Total Fat Per 100 Grams and 100 gram Serving Size or greater.
#
# Looking all the way to the right is my go to Rubio's choice, the Ancho Citrus Shrimp Burrito. At about 450 grams for the burrito and 10 grams of fat per 100 grams of serving, this burrito packs a whopping <ins>45 grams of fat</ins>. This is definitely something to take note of as I have never shied away from eating the whole thing in one sitting.
#
# On the opposite side of the graph, but should be noted as well is one of my favorites, Skippy Creamy Peanut Butter. Although its serving size is on the lower end, the high fat per 100 grams reveals a single serving of peanut butter to have about 16 grams of fat. Again, the amount of Peanut Butter I use is something I will have to keep in mind the next I go to make a sandwich.
#
# Other culprits of high fat vary from fast food items like fries to some favorite household foods like tortillas.
#
# I would also like to reiterate, as I likely will in each section, the caution zone is not definitive and does not mean these items have to be exempt from one's diet rather I feel they should be consumed moderately.
# ## Carbohydrates
#
# For Carbohydrates and Protein I will perform analysis similar to Fats.
"hide_cell"
carbs=df[['food_item','serving_size','serving_measurement_g','total_carbohydrates_g']]
"hide_cell"
carbs.insert(loc=len(carbs.columns),column='carbohydrates_per_100g',value=per_100g(carbs['total_carbohydrates_g']))
"hide_cell"
#create a 'caution' column for foods which contain high fat content
carbs.insert(loc=len(carbs.columns),column='caution',value='No')
for row in carbs.index:
if carbs.at[row,'total_carbohydrates_g'] >= 44:
carbs.at[row,'caution'] = 'Yes'
"hide_cell"
carbs_20 = carbs.nlargest(20,'carbohydrates_per_100g',keep='all')
"hide_input"
plt.figure(figsize=(8,5))
sns.barplot(y='food_item',x='carbohydrates_per_100g',data=carbs_20)
plt.title('Top 20 Items by Carbohydrates Per 100g')
# Looking at the carbohydrates per 100 gram the main culprits are, for the most part, as expected. A lot of items in this list are grain based products which are known to have a higher carbohydrate content.
#
# The surprise items for this list are the fruit snacks, Fruit By The Foot and Welch's mixed fruit. Being fruit based foods I did not expect these to rank high in carbohydrates.
"hide_input"
carbs_fig = px.scatter(carbs,x='serving_measurement_g',y='carbohydrates_per_100g',hover_data=['food_item'],
labels = {'serving_measurement_g':'Single Serving In Grams','carbohydrates_per_100g':'Total Carbs Per 100 Grams'},
title='Serving Size vs. Carbohydrates Per 100 Grams(Interactive Plot)',
color='caution',
color_discrete_map={'No':'Blue','Yes':'Orange'})
carbs_fig.show()
# The first thing I noted from this visualization is that the Annie's Organic Macaroni and Cheese actually contains more carbs than the Kraft Single Serving. However, the Kraft Macaroni and Cheese does contain more fat so there is the trade-off.
#
# The second, more obvious, thing I noted is how few items there are in the cautious zone for carbohydrates. The criteria for the carb cautious zone was for a single serving to contain <ins>44 grams of carbs or more</ins>.
#
# So despite cereal topping the charts for carbs per 100 grams, **if you adhere to the single serving size** they are actually an adequate source of carbohydrates.
#
#
# ### Sugars
#
# Sugars are actually a form of carbohydrates and contribute to overall carbohydrate intake so for the sake of consistency I will analyze sugar content next.
"hide_cell"
sugars = df[['food_item','serving_size','serving_measurement_g','total_sugars_g','added_sugars_g']]
"hide_cell"
sugars.insert(loc=len(sugars.columns),column='total_sugars_per_100g',value=per_100g(sugars['total_sugars_g']))
sugars.insert(loc=len(sugars.columns),column='added_sugars_per_100g',value=per_100g(sugars['added_sugars_g']))
"hide_cell"
sugars.insert(loc=len(sugars.columns),column='caution',value='No')
for row in sugars.index:
if sugars.at[row,'total_sugars_g'] >= 9:
sugars.at[row,'caution'] = 'Yes'
"hide_cell"
sug_20 = sugars.nlargest(20,'total_sugars_per_100g')
add_sug_20 = sugars.nlargest(20,'added_sugars_per_100g')
"hide_input"
plt.figure(figsize=(11,7.5))
plt.subplot(211)
sns.barplot(data=sug_20,y='food_item',x='total_sugars_per_100g')
plt.title('Top 20 Items by Total Sugars Per 100 Grams')
plt.subplot(212)
sns.barplot(data=add_sug_20,y='food_item',x='added_sugars_per_100g')
plt.title('Top 20 Items by Added Sugars Per 100 Grams')
plt.tight_layout()
# Interestingly enough, there appears to be quite some overlap not only with the total sugars and added sugars items, but with the high carb items as well. This makes sense since sugar content makes up part of the total carbohydrate content, so any item with both high carbohydrates and high sugar could be a potential red flag.
#
# One other thing I find interesting from these charts is the Snickers Candy Bar is second highest in terms of total sugars per 100 grams but does not appear in the top 20 of added sugar per 100 grams. This indicates that in terms of sugar content, a Snickers Bar might actually be better than some other food choices here.
"hide_cell"
caution=sugars[sugars['caution']=='Yes']
no_caution=sugars[sugars['caution']=='No']
added_sugars=sugars[sugars['added_sugars_g']>0]
# +
"hide_input"
sugars_fig = py.subplots.make_subplots(rows=2,cols=1,
subplot_titles=('Total Sugars',
'Added Sugars'),
vertical_spacing=0.07,
shared_xaxes=True)
sugars_fig.add_trace(go.Scatter(x=no_caution['serving_measurement_g'],y=no_caution['total_sugars_per_100g'],mode='markers',
name='Total Sugars',
hovertemplate=
'Single Serving in Grams: %{x}'+
'<br>Total Sugars Per 100 Grams: %{y}<br>'+
'Food Item: %{text}',
text=no_caution['food_item']),
row=1, col=1)
sugars_fig.add_trace(go.Scatter(x=caution['serving_measurement_g'],y=caution['total_sugars_per_100g'],mode='markers',
name='Caution',
hovertemplate=
'Single Serving in Grams: %{x}'+
'<br>Total Sugars Per 100 Grams: %{y}<br>'+
'Food Item: %{text}',
text=caution['food_item']),
row=1, col=1)
sugars_fig.add_trace(go.Scatter(x=added_sugars['serving_measurement_g'],
y=added_sugars['added_sugars_per_100g'],mode='markers',
name='Added Sugars',
hovertemplate=
'Single Serving in Grams: %{x}'+
'<br>Added Sugars Per 100 Grams: %{y}<br>'+
'Food Item: %{text}',
text=added_sugars['food_item'],
marker={'color':'Orange'}),
row=2, col=1)
sugars_fig.update_layout(title_text='Serving Size vs. Sugars Per 100 Grams')
sugars_fig.update_xaxes(title_text='Single Serving In Grams',row=2,col=1)
sugars_fig.update_yaxes(title_text='Sugars Per 100 Grams(Interactive Plot)')
sugars_fig.show()
# -
# The first graph here displays the <ins>total sugars</ins> per 100 grams versus single serving size in grams with **caution** on any items that contain 9 or more grams of sugar per serving. The second graph contains all of the items which contain <ins>added sugars</ins> and since added sugars generally want to be avoided, this can be considered its own caution zone.
#
# Surprisingly all of the fruit/fruit-based items, except the single clementine, met the criteria for being high in sugar. Although fruits, particularly whole fruits, are generally considered to be essential in a well-balanced diet, those generally consumed in my household are still high in **natural sugar**.
#
# The big culprits from this sugar analysis are cereals. In the previous section I noted how cereal can still be considered an adequate source of carbohydrates but after some further investigation an overwhelming amount of total carbohydrates comes from **added sugar**.
#
# To get a better understanding, below is a plot showing the relationship between food items' carbohydrates content and their added sugar content.
#
"hide_input"
carbs_sug = px.scatter(df,x='total_carbohydrates_g',y='added_sugars_g' ,hover_data=['food_item'],
labels = {'total_carbohydrates_g':'Single Serving Carbs(g)','added_sugars_g':'Single Serving Added Sugars(g)'},
title='Carbohydrate content vs. Added Sugar Content(Interactive Plot)')
carbs_sug.show()
# The cereals from this data reside in the middle of the plot and it can be seen that added sugar makes up between 1/3 to 1/2 of total carbohydrates for a single serving. Some huge red flags are the Coca-Cola and Aunt Jemima Syrup which both contain 100% of their carbohydrates from added sugars.
#
# The American Heart Association reccomended added sugar intake is no more than 36 grams a day for men and 25 grams for women so it is quite alarming that some of these foods contain half or exceed that amount in just a single serving.<a href="https://www.heart.org/en/healthy-living/healthy-eating/eat-smart/sugar/added-sugars#:~:text=The%20American%20Heart%20Association%20(AHA,day%2C%20or%20about%209%20teaspoons.">[5]</a>
# ## Protein
#
# Now for the analysis of protein content. Generally high protein is recommended in a nutritious diet so I will choose to omit a caution zone for the items.
#
# However, it is important to note excess protein can actually be detrimental since excess protein is stored as fats.[[6]](https://www.healthline.com/health/too-much-protein)
"hide_cell"
protein = df[['food_item','serving_size','serving_measurement_g','protein_g']]
"hide_cell"
protein.insert(loc=len(protein.columns),column='protein_per_100g',value=per_100g(protein['protein_g']))
"hide_cell"
protein_20 = protein.nlargest(20,'protein_per_100g',keep='all')
"hide_input"
plt.figure(figsize=(8,5))
sns.barplot(y='food_item',x='protein_per_100g',data=protein_20)
plt.title('Top 20 Items by Protein Per 100g')
"hide_input"
protein_fig = px.scatter(protein,x='serving_measurement_g',y='protein_per_100g',hover_data=['food_item'],
labels = {'serving_measurement_g':'Single Serving In Grams','protein_per_100g':'Total Protein Per 100 Grams'},
title='Serving Size vs. Protein Per 100 Grams(Interactive Plot)')
protein_fig.show()
# There isn't a whole lot of surprise from the protein results but I was surprised to see the El Pollo Loco Chicken Breast to have the highest protein content. I expected either the Foster Farms Chicken Breast or Foster Farms Ground Turkey to contain the most protein.
#
# One other surprise to me is how little protein the Ancho Citrus Burrito has, especially in comparison to the amount of fat and carbohydrates it contains.
#
# The rest of the high protein items are what I expected being lean meat, fish, or legumes.
# ## Conclusion
#
# The first thing I conclude from this analysis is that I should definitely reconsider ordering the Ancho Citrus Shrimp Burrito since it does not seem nutritionally worth it to me. Additionally I want to avoid cereal as much as possible due to the high content of added sugar in a single serving alone.
#
# Overall, anything in excess can be detrimental and moderation is important in sustaining nutrition. No one of the cautious items will destroy nutrition the same as no single non-cautious item will fix nutrition, the critical part is to maintain balance.
# ### Resources
#
# [1] A link to an overview provided by USDA for more resources related to macronutrients:
#
# https://www.nal.usda.gov/fnic/macronutrients
#
# [2] A link to to the Dietary Reference Intake for macronutrients(A consensus study publication):
#
# https://www.nap.edu/catalog/10490/dietary-reference-intakes-for-energy-carbohydrate-fiber-fat-fatty-acids-cholesterol-protein-and-amino-acids
#
# [3] A link to current USDA dietary guidelines which focus more on substance of foods(vegetables, fruits, etc.) consumed as opposed to simply the macronutrients:
#
# https://health.gov/our-work/food-nutrition/2015-2020-dietary-guidelines
#
# [4] A link for a macronutrient calculator:
#
# https://www.calculator.net/macro-calculator.html
#
# [5] A link to an article on sugar intake:
#
# https://www.heart.org/en/healthy-living/healthy-eating/eat-smart/sugar/added-sugars#:~:text=The%20American%20Heart%20Association%20
#
# [6] A link to an article on protein intake:
#
# https://www.healthline.com/health/too-much-protein
| nutrition_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
def sol(X,Y):
# set union
print("Union of X and Y is",X | Y)
# set intersection
print("Intersection of X and Y is",X & Y)
# set difference
print("Difference of X and Y is",X - Y)
# set symmetric difference
print("Symmetric difference of X and Y is",X ^ Y)
X = {0, 2, 4, 6, 8}
Y = {1, 2, 3, 4, 5}
sol(X,Y)
| Intermediate programs/ 7. Perform different Set Operations..ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.10 64-bit (''RL'': conda)'
# name: python3
# ---
import torch
import numpy as np
from Traj2Dataset import TrajDataset, DatasetTransform
from torch.utils.data import DataLoader
from pytorch_lightning import seed_everything
from model import LitMLP
root_dir = 'dataset'
system = 'great-piquant-bumblebee'
SEED = 42
num_workers = 0
seed_everything(SEED)
# +
state_norms, action_norms = TrajDataset.norms(system)
state_dim = len(state_norms)
action_dim = len(action_norms)
mean = state_norms[:, 0] # mean
std = state_norms[:, 1] # std_dev
transform = DatasetTransform(mean, std)
target_mean = action_norms[:, 0] # mean
target_std = action_norms[:, 1] # std_dev
target_transform = DatasetTransform(target_mean, target_std)
test_dataset = TrajDataset(system, root_dir, train=False,
transform=transform,
target_transform=target_transform)
# -
path = "./submission/models/great-piquant-bumblebee-v1.ckpt"
model = LitMLP.load_from_checkpoint(
path, in_dims=state_dim, out_dims=action_dim)
# +
def upscale(x):
return x * torch.Tensor(target_std) + torch.Tensor(target_mean)
test_dataloader = DataLoader(test_dataset,
batch_size=1, shuffle=False,
num_workers=0)
for i, (x, y) in enumerate(test_dataloader):
print(f'Time-step: {i}')
print(f'Target: {y}')
print(f'Prediction: {model(x)}')
# -
| Imitation-Learning/Test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # HK Housing Analysis
# ## Introduction
#
# Hong Kong is home to one of the most expensive housing markets in the world. This Jupyter Notebook will be charting and interpreting the data provided by the Hong Kong Government.
#
# Since this analysis is being done in the year 2020, economic data is also available to analyze for the months where the Hong Kong economy was influenced by COVID-19 (which covers most of 2020).
# ## Data Sources
# - [Hong Kong Monetary Authority (HKMA)](https://www.hkma.gov.hk/)
# - [Ratings and Valuation Department (RVD)](https://www.rvd.gov.hk/)
# - [Census and Statistics Department (Censtatd)](https://www.censtatd.gov.hk/home.html)
# +
# Prerequisites
import pandas as panda
import pymysql
import matplotlib.pyplot as plot
# Create database connection
host = "localhost";
database = "hkhousinganalysis";
username = "root";
password = "";
connection = pymysql.connect(host=host, user=username, password=password, db=database)
cursor = connection.cursor()
# -
# ## Consumer Price Index Performance
#
# Below charts the CPI performance in Hong Kong. Also known as "Consumer Price Index", is essentially to measure purchase power. Despite not being exactly part of housing, CPI is still important to determine the purchasing power and economics of renters and buyers in the housing market.
# +
# CPI Data
cpiDataQuery = cursor.execute("SELECT `year`, `cpi` FROM `censtats_cpi_data`;")
cpiDataQuery = cursor.fetchall()
cpiDF = panda.DataFrame(cpiDataQuery)
yearToNum = []
cpiToNum = []
for row in cpiDF.iloc(0):
if str(row[1]) != "":
yearToNum.append(int(row[0]))
cpiToNum.append(float(row[1]))
plot.plot(yearToNum, cpiToNum)
plot.title("Yearly CPI Performance")
plot.xlabel("Year")
plot.ylabel("CPI (Pts)")
plot.show()
# -
# As seen in the chart above, the CPI for Hong Kong residents has increased over the years from the start of the data set, with the uptrend expected to continue in the long-term.
# ## House Prices
#
# Below charts the price averages per square meter for bought housing across Grades A to E housing properties on the market.
# +
# Housing Price Data
# Hong Kong Island Data
priceByClassHKIDataQuery = cursor.execute("SELECT * FROM `rv_price_by_class_data_hki`;")
pricebyClassHKIDataQuery = cursor.fetchall()
priceByClassHKIDF = panda.DataFrame(pricebyClassHKIDataQuery)
yearToNum = []
avgPriceHKI = []
for row in priceByClassHKIDF.iloc(0):
yearToNum.append(int(row[0]))
avgPriceHKI.append((float(row[1]) + float(row[2]) + float(row[3]) + float(row[4]) + float(row[5]))/5)
try:
plot.plot(yearToNum, avgPriceHKI, label="Hong Kong Island")
plot.title("Housing Price Per Square Meter")
plot.xlabel("Year")
plot.xticks(rotation=90)
plot.ylabel("Price Per Square Meter (HKD)")
except TypeError:
pass
# Kowloon Data
priceByClassKOWDataQuery = cursor.execute("SELECT * FROM `rv_price_by_class_data_kow`;")
pricebyClassKOWDataQuery = cursor.fetchall()
priceByClassKOWDF = panda.DataFrame(pricebyClassKOWDataQuery)
yearToNum = []
avgPriceKOW = []
for row in priceByClassKOWDF.iloc(0):
yearToNum.append(int(row[0]))
avgPriceKOW.append((float(row[1]) + float(row[2]) + float(row[3]) + float(row[4]) + float(row[5]))/5)
try:
plot.plot(yearToNum, avgPriceKOW, label="Kowloon")
except TypeError:
pass
# New Territories Data
priceByClassNTDataQuery = cursor.execute("SELECT * FROM `rv_price_by_class_data_nt`;")
pricebyClassNTDataQuery = cursor.fetchall()
priceByClassNTDF = panda.DataFrame(pricebyClassNTDataQuery)
yearToNum = []
avgPriceNT = []
for row in priceByClassNTDF.iloc(0):
yearToNum.append(int(row[0]))
avgPriceNT.append((float(row[1]) + float(row[2]) + float(row[3]) + float(row[4]) + float(row[5]))/5)
try:
plot.plot(yearToNum, avgPriceNT, label="New Territories")
except TypeError:
pass
# All Territories
avgPriceAllTerritories = []
for index in range(len(avgPriceNT)):
avgPriceAllTerritories.append((float(avgPriceHKI[index]) + float(avgPriceKOW[index]) + float(avgPriceNT[index]))/3)
try:
plot.plot(yearToNum, avgPriceAllTerritories, label="All Territories")
plot.legend()
plot.show()
except TypeError:
pass
# -
# The chart itself has charted lines for all 3 territories of Hong Kong, as well the average price line for all territories together.
#
# As seen in the chart, New Territories rent per square meter is the lowest out of the 3 territories. This could be due to New Territories being much farther away from locations stretching across Queen's Road, making it less appealing compared to Kowloon and Hong Kong Island.
#
# Properties in Hong Kong Island are as expected, much more expensive as there is no need to cross Victoria Harbour to reach offices in Central, Causeway Bay, and so on.
#
# Kowloon, despite being densely populated and retaining Hong Kong's old and classic architecture, it seems to fall in the middle between Hong Kong Island and New Territories properties. It is also geographically in the middle of the two territories, making it easy to get to Central.
# ## Rent Prices
#
# Below charts the price averages per square meter for rentals across Grades A to E housing properties on the market:
# +
# Housing Rent Data
# Hong Kong Island Data
rentByClassHKIDataQuery = cursor.execute("SELECT * FROM `rv_rent_by_class_data_hki`;")
rentByClassHKIDataQuery = cursor.fetchall()
rentByClassHKIDF = panda.DataFrame(rentByClassHKIDataQuery)
yearToNum = []
avgRentHKI = []
for row in rentByClassHKIDF.iloc(0):
yearToNum.append(int(row[0]))
avgRentHKI.append((float(row[1]) + float(row[2]) + float(row[3]) + float(row[4]) + float(row[5]))/5)
try:
plot.plot(yearToNum, avgRentHKI, label="Hong Kong Island")
plot.title("Housing Rent Per Square Meter")
plot.xlabel("Year")
plot.xticks(rotation=90)
plot.ylabel("Price Per Square Meter (HKD)")
except TypeError:
pass
# Kowloon Data
rentByClassKOWDataQuery = cursor.execute("SELECT * FROM `rv_rent_by_class_data_kow`;")
rentbyClassKOWDataQuery = cursor.fetchall()
rentByClassKOWDF = panda.DataFrame(rentbyClassKOWDataQuery)
yearToNum = []
avgRentKOW = []
for row in rentByClassKOWDF.iloc(0):
yearToNum.append(int(row[0]))
avgRentKOW.append((float(row[1]) + float(row[2]) + float(row[3]) + float(row[4]) + float(row[5]))/5)
try:
plot.plot(yearToNum, avgRentKOW, label="Kowloon")
except TypeError:
pass
# New Territories Data
rentByClassNTDataQuery = cursor.execute("SELECT * FROM `rv_rent_by_class_data_nt`;")
rentbyClassNTDataQuery = cursor.fetchall()
rentByClassNTDF = panda.DataFrame(rentbyClassNTDataQuery)
yearToNum = []
avgRentNT = []
for row in rentByClassNTDF.iloc(0):
yearToNum.append(int(row[0]))
avgRentNT.append((float(row[1]) + float(row[2]) + float(row[3]) + float(row[4]) + float(row[5]))/5)
try:
plot.plot(yearToNum, avgRentNT, label="New Territories")
except TypeError:
pass
# All Territories
avgRentAllTerritories = []
for index in range(len(avgPriceNT)):
avgRentAllTerritories.append((float(avgRentHKI[index]) + float(avgRentKOW[index]) + float(avgRentNT[index]))/3)
try:
plot.plot(yearToNum, avgRentAllTerritories, label="All Territories")
plot.legend()
plot.show()
except TypeError:
pass
# -
# Similar to the chart for buying, the rent chart seems to follow the same trend with Hong Kong Island being the most expensive and New Territories being the least expensive out of all territories.
# ## Monthly Salaries
#
# The values in the table below represent the monthly salaries of Hong Kong residents based on occupation.
# +
# Monthly Salaries Data
monthlySalariesDataQuery = cursor.execute("SELECT * FROM `censtats_monthlypay_amount_data`;")
monthlySalariesDataQuery = cursor.fetchall()
tableRetrieveColumnNames = []
for tableColumn in cursor.description:
tableRetrieveColumnNames.append(tableColumn[0])
unemploymentDF = panda.DataFrame(monthlySalariesDataQuery, columns=tableRetrieveColumnNames)
tableColumns = unemploymentDF.columns
filteredTableColumns = [tableColumns[0]]
tableColumns = tableColumns[12:42]
for column in tableColumns:
filteredTableColumns.append(column)
unemploymentDF[filteredTableColumns]
# -
# In general, the overall salary of a Hong Kong resident has increased over the years from 2011, with education and public admin workers making the most average monthly salary across all occupations for the year 2019. It seems that monthly salary is also following the uptrend that both CPI and housing is experiencing, which shows signs of a growing economy, or inflation of the US Dollar, which the Hong Kong Dollar is pegged to.
# +
# Unemployment Rate Data
unemploymentDataQuery = cursor.execute("SELECT `end_of_month`, `unemploy_rate` FROM `hkma_econ_data`;")
unemploymentDataQuery = cursor.fetchall()
unemploymentDF = panda.DataFrame(unemploymentDataQuery)
endOfMonthToNum = []
unemploymentRateToNum = []
for row in unemploymentDF.iloc(0):
if str(row[0])[5:7] != "00":
endOfMonthToNum.append(str(row[0]))
unemploymentRateToNum.append(panda.to_numeric(row[1]))
try:
plot.plot(endOfMonthToNum, unemploymentRateToNum)
plot.title("Unemployment Rate Performance")
plot.xlabel("Date Recorded")
plot.xticks(rotation=90)
plot.ylabel("Rate (%)")
plot.show()
except TypeError:
pass
# -
# Despite monthly salaries growing year over year, unemployment has increased sharply due to the COVID-19 pandemic, affecting Hong Kong's tourism business and financial hub in Asia. Despite increasing from 3.5% to over 6.0% over the course of the pandemic, Hong Kong seems to be on the road of a slow recovery as unemployment has dropped slightly between June and July of 2020.
# ## Hang Seng Index vs Housing Indices
#
# The Hang Seng Index is an index comprised of some of the largest companies in Hong Kong, which makes it very good to guage how well the Hong Kong economy is doing.
#
# Other than HSI, the housing indices for price and rent are also charted below.
# +
# HSI Data
hsiDataQuery = cursor.execute("SELECT `end_of_month`, `eq_mkt_hs_index` FROM `hkma_capitalmarkets_data`;")
hsiDataQuery = cursor.fetchall()
hsiDataDF = panda.DataFrame(hsiDataQuery)
endOfMonthToNum = []
hsiPriceToNum = []
for row in hsiDataDF.iloc(0):
endOfMonthToNum.append(str(row[0]))
hsiPriceToNum.append(panda.to_numeric(row[1]))
try:
plot.plot(endOfMonthToNum, hsiPriceToNum, label="HSI Price")
plot.plot([endOfMonthToNum[0], endOfMonthToNum[len(endOfMonthToNum)-1]], [hsiPriceToNum[0], hsiPriceToNum[len(hsiPriceToNum)-1]], label="Return from start of data set to latest price")
plot.title("Hang Seng Index")
plot.xlabel("Date Recorded")
plot.xticks(rotation=90)
plot.ylabel("Price (HKD)")
plot.legend()
plot.show()
except TypeError:
pass
# Housing Price Index Data
housingPriceIndexDataQuery = cursor.execute("SELECT `year`, `All_Classes` FROM `rv_price_index_by_class_data`;")
housingPriceIndexDataQuery = cursor.fetchall()
housingPriceIndexDF = panda.DataFrame(housingPriceIndexDataQuery)
yearToNum = []
housingPriceToNum = []
for row in housingPriceIndexDF.iloc(0):
if str(row[1]) != "":
yearToNum.append(int(row[0]))
housingPriceToNum.append(float(row[1]))
try:
plot.plot(yearToNum, housingPriceToNum, label="Price Index")
plot.title("Housing Price Index vs Housing Rent Index")
plot.xlabel("Date Recorded")
plot.xticks(rotation=90)
plot.ylabel("Index (Pts)")
except TypeError:
pass
# Housing Rent Index Data
housingRentIndexDataQuery = cursor.execute("SELECT `year`, `All_Classes` FROM `rv_rent_index_by_class_data`;")
housingRentIndexDataQuery = cursor.fetchall()
housingRentIndexDF = panda.DataFrame(housingRentIndexDataQuery)
yearToNum = []
housingRentToNum = []
for row in housingRentIndexDF.iloc(0):
if str(row[1]) != "":
yearToNum.append(int(row[0]))
housingRentToNum.append(float(row[1]))
try:
plot.plot(yearToNum, housingRentToNum, label="Rent Index")
plot.legend()
plot.show()
except TypeError:
pass
# -
# Both historical data sets for Hang Seng and housing indices show that the overall trend is upwards. Do note that the time series is spanned from 2015 to 2020 comapred to the housing indices that span from 1980 to 2020, however the HSI has been trending upward for the data set, and as shown in the chart above for housing indices, between 2015 and 2020, both price and rent indices trend up as well.
# ## Conclusion
#
# With a global COVID-19 pandemic, many Hong Kong businesses have shut down permanently, especially restaurants and luxury clothing stores. This proved costly to the economy as the Hang Seng Index has dropped from well over 28000 HKD at the end of 2019 to 23000 HKD in May 2020.
#
# However, Hong Kong house prices continue to increase over the years, matching the year-over-year increase in CPI, prices per square meter, and the overall HK economy. All charts show that the latest data points have shown a slow down in the uptrend, which should recover within the next few years once Hong Kong's tourism levels return to normal to stimulate the local economy.
#
| HK Housing Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: tf1.11_gpu
# language: python
# name: tf111_gpu
# ---
# <img src="../Pics/MLSb-T.png" width="160">
# <br><br>
# <center><u><H1>Encoder-Decoder with Attention</H1></u></center>
import tensorflow as tf
from keras.backend.tensorflow_backend import set_session
config = tf.ConfigProto()
config.gpu_options.allow_growth = True
config.log_device_placement = True
sess = tf.Session(config=config)
set_session(sess)
from keras.models import Sequential
from keras.layers import CuDNNLSTM, LSTM
from keras.layers import Dense
from keras.layers import TimeDistributed
from keras.layers import RepeatVector
from random import randint
from numpy import array
from numpy import argmax
from numpy import array_equal
# function to generate random integer values between 0 and n. The number of timestpes is: length
def gen_seq(length, n):
return [randint(0, n-1) for _ in range(length)]
# ## Generate sequence:
sequence = gen_seq(6, 30)
sequence
def onehot_encoder(seq, n):
encod = []
for s in seq:
v = [0 for _ in range(n)]
v[s] = 1
encod.append(v)
return array(encod)
def onehot_decoder(encod_seq):
return [argmax(idx) for idx in encod_seq]
onehot = onehot_encoder(sequence, 30)
print(onehot)
decoded = onehot_decoder(onehot)
print(decoded)
def generate_pair(n_in, n_out, n_total):
# generating random sequences
seq_in = gen_seq(n_in, n_total)
seq_out = seq_in[:n_out] + [0 for _ in range(n_in-n_out)]
X = onehot_encoder(seq_in, n_total)
y = onehot_encoder(seq_out, n_total)
# reshaping as 3D tensor
X = X.reshape((1, X.shape[0], X.shape[1]))
y = y.reshape((1, y.shape[0], y.shape[1]))
return X,y
X, y = generate_pair(6, 3, 30)
print('X=%s, y=%s' % (onehot_decoder(X[0]), onehot_decoder(y[0])))
print(X.shape, y.shape)
# ## Encoder-Decoder without Attention
n_features = 50
n_timesteps_in = 5
n_timesteps_out = 3
# ## Creating the model:
model = Sequential()
model.add(CuDNNLSTM(150, input_shape=(n_timesteps_in, n_features)))
model.add(RepeatVector(n_timesteps_in))
model.add(CuDNNLSTM(150, return_sequences=True))
model.add(TimeDistributed(Dense(n_features, activation='softmax')))
model.summary()
model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['acc'])
# ## Training the model:
for epoch in range(5000):
X,y = generate_pair(n_timesteps_in, n_timesteps_out, n_features)
model.fit(X, y, epochs=1, verbose=1)
# ## Testing the model:
epochs = 100
correct = 0
# Testing the model with new 100 new randomly generated integer sequences
for _ in range(epochs):
X,y = generate_pair(n_timesteps_in, n_timesteps_out, n_features)
pred = model.predict(X)
if array_equal(onehot_decoder(y[0]), onehot_decoder(pred[0])):
correct += 1
print('Accuracy: %.2f%%' % (float(correct)/float(epochs)*100.0))
# ## Checking samples:
#checking 20 examples of expected output sequences and predictions
for _ in range(20):
X,y = generate_pair(n_timesteps_in, n_timesteps_out, n_features)
pred = model.predict(X)
print('Expected:', onehot_decoder(y[0]), 'Predicted', onehot_decoder(pred[0]))
# ## Encoder-Decoder with Attention Layer
from attention_decoder import AttentionDecoder
# ## Creating the model with Attention:
model_att = Sequential()
model_att.add(CuDNNLSTM(150, input_shape=(n_timesteps_in, n_features), return_sequences=True))
model_att.add(CuDNNLSTM(150, return_sequences=True))
model_att.add(CuDNNLSTM(150, return_sequences=True))
model_att.add(AttentionDecoder(150, n_features))
model_att.summary()
model_att.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['acc'])
for epoch in range(5000):
X, y = generate_pair(n_timesteps_in, n_timesteps_out, n_features)
model_att.fit(X, y, epochs=1, verbose=1)
# ## Testing the attention model:
for _ in range(epochs):
X, y = generate_pair(n_timesteps_in, n_timesteps_out, n_features)
pred_att = model_att.predict(X)
if array_equal(onehot_decoder(y[0]), onehot_decoder(pred_att[0])):
correct += 1
print('Accuracy: %.2f%%' % (float(correct)/float(epochs)*100.0))
# ## Checking samples:
for _ in range(20):
X,y = generate_pair(n_timesteps_in, n_timesteps_out, n_features)
pred = model_att.predict(X)
print('Expected:', onehot_decoder(y[0]), 'Predicted', onehot_decoder(pred[0]))
| DEEP_NLP_resources/3-Deep-Learning for NLP/Encoder-Decoder with Attention.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
# matplotlibをjupyter内で表示させるおまじない
# %matplotlib inline
from IPython.display import Audio # jupyter上で音声を再生するためのおまじない
# +
# 白色雑音のサンプル数を設定
n_samples = 16000
# サンプリング周波数
sample_freq = 16000
# -
# 白色雑音を生成
data = np.random.normal(size=n_samples)
# +
# 時間軸の設定
time = np.arange(n_samples) / sample_freq
# 音声データのプロット
plt.plot(time, data)
# x軸のラベル
plt.xlabel("Time (sec)")
# y軸のラベル
plt.ylabel("Amplitude")
# 画像のタイトル
plt.title("Waveform")
# -
# 白色雑音を再生(音量注意!)
Audio(data, rate=sample_freq)
| WarmUp/plt_whitenoise.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import datetime
# +
date_time_format = "%Y/%m/%d"
users = pd.read_csv("./Dataset/JData_User.csv", encoding="gbk")
comments = pd.read_csv("./Dataset/JData_Comment.csv", encoding="gbk")
ac02 = pd.read_csv("./Dataset/JData_Action_201602.csv", encoding="gbk")
ac03 = pd.read_csv("./Dataset/JData_Action_201603.csv", encoding="gbk")
ac03ex = pd.read_csv("./Dataset/JData_Action_201603_extra.csv", encoding="gbk")
ac04 = pd.read_csv("./Dataset/JData_Action_201604.csv", encoding="gbk")
actions = pd.concat([ac02, ac03, ac03ex, ac04])
product = pd.read_csv("./Dataset/JData_Product.csv", encoding="gbk")
users['user_reg_dt'] = pd.to_datetime(users['user_reg_dt'], format=date_time_format)
comments['dt'] = pd.to_datetime(comments['dt'], format=date_time_format)
# -
ac02.groupby('model_id').count()
# ## 用户分布
# +
x_pos = np.array([1, 5, 9])
plt.bar(x_pos, users.groupby('sex').count()['user_id'])
plt.xticks(x_pos + .35, (u'男性', u'女性', u'未知'))
plt.ylabel(u'数量')
plt.title(u'性别分布')
plt.figure()
x_pos = np.array([1, 3, 5, 7, 9])
plt.bar(x_pos, users.groupby('user_lv_cd').count()['user_id'])
plt.xticks(x_pos + .35, users.groupby('user_lv_cd').count()['user_id'].index)
plt.ylabel(u'数量')
plt.title(u'活跃等级分布')
plt.figure()
age_count = users.groupby('age').count()['user_id']
x_pos = np.arange(len(age_count)) + 1
plt.bar(x_pos, age_count)
plt.xticks(x_pos + .25, age_count.index)
plt.ylabel(u'数量')
plt.title(u'年龄')
plt.figure()
date_2016_2_1 = datetime.date(2016, 2, 1)
register_after = users[users['user_reg_dt'] > date_2016_2_1]
register_before = users[users['user_reg_dt'] < date_2016_2_1]
plt.plot(register_before.groupby('user_reg_dt').count()['user_id'], color='b', label=u'之前注册')
plt.plot(register_after.groupby('user_reg_dt').count()['user_id'], color='r', label=u'之后注册')
plt.xlabel(u'日期')
plt.ylabel(u'数量')
plt.title(u'注册日期')
plt.legend()
plt.show()
# +
def cut_comment(comment_mean, low, high):
left = comment_mean[comment_mean > low]
right = left[left <= high]
return right.count()
def draw_comment(comment_mean, name):
com_count = [
comment_mean[comment_mean==0].count(),
cut_comment(comment_mean, 0, .1),
cut_comment(comment_mean, .1, .2),
cut_comment(comment_mean, .2, .4),
cut_comment(comment_mean, .4, .6),
cut_comment(comment_mean, .6, .8),
cut_comment(comment_mean, .8, 1),
]
x_pos = np.arange(len(com_count)) + 1
plt.bar(x_pos, com_count)
# plt.xticks(x_pos + .25, ("0%", "0% - 10%","10% - 20%" ,"20% - 40%", "40% - 60%", "60% - 80%", "80% - 100%"))
plt.ylabel(u'数量')
plt.title(u'{0}'.format(name))
plt.show()
comment_mean = comments.groupby('sku_id')['bad_comment_rate'].mean()
draw_comment(comment_mean, u"平均差评率")
# i = 1
# for name, comm in comments.groupby('dt'):
# plt.figure()
# i += 1
# com = comm['bad_comment_rate']
# draw_comment(com, name)
| Competition/JData/view.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # This notebook create plots for data-model-comparison
# This notebook create zonal lat. averaged temperatures for Darrell.
import xarray as xr
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import xesmf as xe
# %matplotlib inline
import cartopy
import cartopy.crs as ccrs
import matplotlib
from netCDF4 import Dataset
from mpl_toolkits.axes_grid1 import make_axes_locatable
import numpy.ma as ma
import math
import xlrd
import os
import matplotlib.colors as colors
import seaborn as sns
import scipy
# +
#No change needs here
#PMIP model list used in this notebook represents CMIP models included in PMIP
'''Define CMIP list'''
def whole_model_list():
filename_xls='PMIP & CMIP model data status.xlsx'
wb=xlrd.open_workbook(filename_xls)
sh_pmip4 = wb.sheet_by_name('PMIP4-CMIP6')
model_id4=sh_pmip4.col_values(14, 6, 28)
cmip6=[]
cmip6_generation=sh_pmip4.col_values(20, 6, 28)
pmip4_generation=sh_pmip4.col_values(21, 6, 28)
for i in range(len(model_id4)):
if (cmip6_generation[i]=='CMIP6' and pmip4_generation[i]=='PMIP4'):
cmip6.append(model_id4[i])
cmip5=[]
sh_pmip3 = wb.sheet_by_name('PMIP3-CMIP5')
model_id3=sh_pmip3.col_values(12, 2, 20)
cmip5_generation=sh_pmip3.col_values(18, 2, 20)
pmip3_generation=sh_pmip3.col_values(19, 2, 20)
for i in range(len(model_id3)):
if (cmip5_generation[i]=='CMIP5' and pmip3_generation[i]=='PMIP3'):
cmip5.append(model_id3[i])
return model_id4,cmip6,model_id3,cmip5
#PMIP4-CMIP6
#Cell range: B6 - X28
#Below: No values are corresponed No in python
# model row No.: 6-27
# col_No. col. experiment/column content
# 4 E 0kpiControl
# 5 F LMpast1000(1000 years)
# 6 G 6kmidHolocene
# 7 H 21klgm
# 8 I Last Interglaciallig127k
# 9 J Mid Pliocene warm periodmidPliocene-eoi400
# 14 O Model id
# 20 U CMIP6
# 21 V PMIP4
#PMIP3-CMIP5
#Cell range: B2 - U20
#Below: No values are corresponed No in python
# model row No.: 2-19
# col_No. col. experiment/column content
# 3 D 0kpiControl
# 4 E LMpast1000(1000 years)
# 5 F 6kmidHolocene
# 6 G 21klgm
# 7 H Last Interglaciallig127k
# 8 I Mid Pliocene warm periodmidPliocene-eoi400
# 12 M Model id
# 18 S CMIP6
# 19 T PMIP4
pmip4,cmip6,pmip3,cmip5=whole_model_list()
pmip={}
pmip['PMIP3']=cmip5
pmip['PMIP4']=cmip6
pmip_v4='PMIP4'
pmip_v3='PMIP3'
# +
pmip4=['IPSL-CM6A-LR',
'HadGEM3-GC31',
'AWI-ESM']
pmip3=['GISS-E2-1-G',
'COSMOS-ASO',
'MRI-CGCM3',
'CNRM-CM5',
'FGOALS-g2',
'GISS-E2-R',
'EC-EARTH-2-2',
'IPSL-CM5A-LR',
'bcc-csm1-1',
'HadGEM2-CC',
'HadCM3',
'MPI-ESM-P',
'CCSM4',
'CSIRO-Mk3L-1-2',
'CSIRO-Mk3-6-0',
'FGOALS-s2',
'MIROC-ESM']
pmip_all=['GISS-E2-1-G',
'COSMOS-ASO',
'AWI-ESM',
'MRI-CGCM3',
'CNRM-CM5',
'FGOALS-g2',
'GISS-E2-R',
'EC-EARTH-2-2',
'IPSL-CM5A-LR',
'IPSL-CM6A-LR',
'bcc-csm1-1',
'HadGEM3-GC31',
'HadGEM2-CC',
'HadCM3',
'MPI-ESM-P',
'CCSM4',
'CSIRO-Mk3L-1-2',
'CSIRO-Mk3-6-0',
'FGOALS-s2',
'MIROC-ESM']
pmip={}
pmip['PMIP3']=pmip3
pmip['PMIP4']=pmip4
pmip['PMIP']=pmip_all
pmip_v3='PMIP3'
pmip_v4='PMIP4'
# +
#No change needs here
'''Define calculating functions'''
#This function will get all available experiment names
def experimentlist():
exps=[]
file_path = "data"
for dirpaths, dirnames, filenames in os.walk(file_path):
for d in dirnames:
exps.append(d)
return exps
#This function will get all available model names in the experiment
def modellist(experiment_name):
models=[]
file_path = "data/%s" %(experiment_name)
for dirpaths, dirnames, filenames in os.walk(file_path):
for f in filenames:
mname=f.split("_")[0]
models.append(mname)
return models
#This function will get all available filenames in the experiment
def filenamelist(experiment_name):
filenames=[]
file_path = "data/%s" %(experiment_name)
for dirpaths, dirnames, files in os.walk(file_path):
for f in files:
ff='data/%s/%s'%(experiment_name,f)
filenames.append(ff)
return filenames
#This function will identify models in the ensemble
def identify_ensemble_members(variable_name,experiment_name):
datadir="data/%s" %(experiment_name)
# ensemble_members=!scripts/find_experiment_ensemble_members.bash {experiment_name} {variable_name} {datadir}
return ensemble_members
#This function will list excat model name
def extract_model_name(filename):
file_no_path=filename.rpartition("/")
file_strings=file_no_path[2].partition("_")
model_name=file_strings[0]
return model_name
def ensemble_members_dict(variable_name,experiment_name):
ens_mems=identify_ensemble_members(variable_name,experiment_name)
ens_mems_dict={extract_model_name(ens_mems[0]):ens_mems[0]}
for mem in ens_mems[1:]:
ens_mems_dict[extract_model_name(mem)]=mem
return ens_mems_dict
#This function will find an regird avaiable models, calculate changes, and return model data and model names
def ensemble_diffence(pmip_v):
dataset={}
model=[]
A_dict=ensemble_members_dict(variable_name,experiment_name)
B_dict=ensemble_members_dict(variable_name,'piControl')
grid_1x1= xr.Dataset({'lat': (['lat'], np.arange(-89.5, 90., 1.0)),
'lon': (['lon'], np.arange(-0, 360., 1.0))})
for gcm in A_dict:
if gcm in B_dict:
if gcm in pmip[pmip_v]:
expt_a_file=xr.open_dataset(A_dict.get(gcm),decode_times=False)
expt_a=expt_a_file[variable_name]
expt_b_file=xr.open_dataset(B_dict.get(gcm),decode_times=False)
expt_b=expt_b_file[variable_name]
diff=expt_a-expt_b
this_regridder=xe.Regridder(expt_a_file,grid_1x1,'bilinear', reuse_weights=True,periodic=True)
diff_1x1=this_regridder(diff)
dataset[gcm]=diff_1x1
model.append(gcm)
return dataset,model
#x=np.arange(0,181,20)
#xla=['-90 ~ -70','-70 ~ -50','-50 ~ -30','-30 ~ -10','-10 ~ 10','10 ~ 30','30 ~ 50','50 ~ 70','70 ~ 90']
x=np.arange(0,181,30)
xla=['-90 ~ -60','-60 ~ -30','-30 ~ 0','0 ~ 30','30 ~ 60','60 ~ 90']
#This function will calculate zonal means of every 20 deg in latitude
def zonal_mean(pmip_v):
ensemble_diff_v,model_diff_v=ensemble_diffence(pmip_v)
data={}
for m in model_diff_v:
data[m]=[]
for i in range(len(x)-1):
men=ensemble_diff_v[m][x[i]:x[i+1]]
xx=np.average(men)
data[m].append(xx)
return data
#This function will plot zonal data
def zonal_scatter(data):
fig=plt.figure(figsize=(10,7))
for m in model_diff_v:
plt.scatter(data[m],xla,marker='o',s=50,label=m)
plt.legend()
plt.axvline(x=0,color='k',linestyle="--")
plt.ylabel('latitude (deg_N)')
plt.xlabel('changes in TAS (midHolocene - piControl)')
return fig
#This function will combine all functions above
def zonal_plot(pmip_v):
ensemble_diff_v,model_diff_v=ensemble_diffence(pmip_v)
data={}
for m in model_diff_v:
data[m]=[]
for i in range(len(x)-1):
men=ensemble_diff_v[m][x[i]:x[i+1]]
xx=np.average(men)
data[m].append(xx)
plt.figure(figsize=(10,10))
plt.axvline(x=0,color='k',linestyle="--")
for m in model_diff_v:
plt.scatter(data[m],xla,marker='o',s=50,label=m)
plt.legend()
plt.ylabel('latitude (deg_N)')
x_label='changes in %s (%s - piControl)' %(variable_name,experiment_name)
plt.xlabel(x_label)
plt.plot(0,)
plt.title(pmip_v)
# +
'''Set experiment and variable names'''
experiment_name='midHolocene-cal-adj'
variable_name='tas_spatialmean_ann'
zonal_plot(pmip_v3)
# +
experiment_name='midHolocene-cal-adj'
variable_name='tas_spatialmean_ann'
zonal_plot(pmip_v3)
| notebooks/PMIP4_MH_DMC.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
import matplotlib as mpl
import seaborn as sns
# Style
sns.set() #Set Seaborn style
sns.set_style('darkgrid') # Tick style 'darkgrid', 'whitegrid', 'dark', 'white'
sns.set_color_codes("deep") # How Color is interpreted 'deep', 'muted', 'pastel', 'bright', 'colorblind'
# _ = plt.plot([0, 2], color="c")
# _ = plt.plot([0, 1], color="g")
# RC Setting
# mpl.rc('font', family='nanumgothic') # for font
# mpl.rc('axes', unicode_minus=False) # for unicode
mpl.rc('figure', figsize=(8, 5)) # figure size (unit: inch)
mpl.rc('figure', dpi=300) # figure resolution
# Color Setting
# sns.palplot(sns.color_palette("hls", 8)) #hue, light, saturation
sns.palplot(sns.hls_palette(8, l=.3, s=.8))
# sns.palplot(sns.color_palette("Paired"))
# sns.palplot(sns.color_palette("Set2"))
# flatui = ["#9b59b6", "#3498db", "#95a5a6", "#e74c3c", "#34495e", "#2ecc71"]
# sns.palplot(sns.color_palette(flatui)) # User Defined Color
# colors = ["windows blue", "amber", "greyish", "faded green", "dusty purple"]
# sns.palplot(sns.xkcd_palette(colors)) #xkcd Color Set
# sns.palplot(sns.color_palette("Blues"))
# sns.palplot(sns.color_palette("BuGn_r"))
# sns.palplot(sns.color_palette("GnBu_d"))
# sns.palplot(sns.color_palette("cubehelix", 8))
# +
# Matplotlib Plot
# figure 객체 / Axes 객체(2개 이상의 Axis객체 갖는다) / Axis 객체
# plt.figure(figsize=(10,2)) # figure 객체 생성 그러나 plot으로도 생성되서 figsize나 여러개의 plot을 그리지 않으
plt.title("Plot")
면 필요 없다.
# ax = plt.subplot(2, 1, 1) # 그리드 형태의 Axes 객체 생성
# plt.plot([1,4,9,16]) # F(0) = 4 , F(1) = 4 ...
plt.plot([10,20,30,40], [1,4,9,16], label="just line") # x,y function
# plt.plot([10, 20, 30, 40], [1, 4, 9, 16], 'rs--')
# c,m,lw 순으로 스타일 지정
# Color : b(blue), g(green), r(red), c(cyan), m(magenta), y(yellow), k(black), w(white)
# Marker : .(point), ,(pixel), o(circle), v(triangle_down), ^(tri_up), <(tri_left), >(tri_right)
# *(star), p(pentagon), +(plus), D(diamond), d(thin_diamond)
plt.plot([10, 20, 30, 40], [1, 4, 9, 16], c="b",
lw=5, ls="--", marker="d", ms=15, mec="g", mew=5, mfc="r")
# plt.plot(t, t, 'r--', t, 0.5 * t**2, 'bs:', t, 0.2 * t**3, 'g^-') #동시에 여러개 그리기
# c(color), lw(linewidth), ls(linestyle), marker, ms(markersize),
# mec(markeredgecolor), mew(markeredgewidth), mfc(markerfacecolor)
plt.xlim(12, 50) # x range
plt.ylim(-10,30) # y range
plt.xticks([11, 19, 29]) # mark on axis
plt.yticks([1, 3, 8, 15])
plt.grid(False) # grid 설정
plt.legend(loc=2) # 범례 표시
plt.show()
# -
fig, ax0 = plt.subplots()
ax1 = ax0.twinx() # x축을 공유하는 Axes 객체
ax0.set_title("2개의 y축 한 figure에서 사용하기")
ax0.plot([10, 5, 2, 9, 7], 'r-', label="y0")
ax0.set_ylabel("y0")
ax0.grid(False)
ax1.plot([100, 200, 220, 180, 120], 'g:', label="y1")
ax1.set_ylabel("y1")
ax1.grid(False)
ax0.set_xlabel("공유되는 x축")
plt.show()
# +
# 1차원 실수 분포
# sns.rugplot(x) # 꽃잎의 길이 분포
# sns.kdeplot(x) # 꽃잎 길이에 대한 density 그래프
# sns.distplot(x, kde=True, rug=True) # 막대분포 rug + kde
# sns.countplot(x="column_name", data=dataframe) # 각 카테고리별 데이터 count
# 2차원 실수 데이터
# sns.pairplot(iris, hue="species", markers=["o", "s", "D"])
# 2차원 카테고리 데이터
# sns.heatmap(titanic_size, cmap=sns.light_palette("gray", as_cmap=True), annot=True, fmt="d")
# 2차원 복합 데이터
# sns.barplot(x="day", y="total_bill", data=tips) # 요일별 팁의 평균(막대바), 편차(에러바)
# sns.boxplot(x="day", y="total_bill", data=tips) #
# sns.violinplot(x="day", y="total_bill", data=tips) #
# sns.stripplot(x="day", y="total_bill", data=tips, jitter=True) # 스캐터 플롯처럼 점으로 그려준다. Jitter True 설정시 겹치지 않게 보여줌
# 다차원 복합 데이터
# sns.barplot(x="day", y="total_bill", hue="smoker",data=tips) # 두가지 카테고리 값에 의한 변화를 볼 수 있게 hue값을 제공
# sns.boxplot(x="day", y="total_bill", hue="sex", data=tips)
# sns.violinplot(x="day", y="total_bill", hue="sex", data=tips)
# sns.stripplot(x="day", y="total_bill", hue="sex", data=tips, jitter=True)
# sns.swarmplot(x="day", y="total_bill", hue="sex", data=tips)
# sns.violinplot(x="day", y="total_bill", hue="sex", data=tips, split=True)
# sns.heatmap(flights_passengers, annot=True, fmt="d", linewidths=1) # 두개의 카테고리 값에 의한 실수 값 변화들
# 3개 이상의 카테고리
# sns.catplot(x="age", y="sex", hue="survived", row="class", data=data,kind="violin", split=True, height=2, aspect=4)
# -
| Visualization/Drawing Tool.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/ayulockin/DLshots/blob/master/PyTorch_Classification_Example.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="CufETwmZN-vF" colab_type="text"
# ## Setups, Imports and Installations
# + id="HESv1l--3AAa" colab_type="code" colab={}
# %%capture
# !pip install wandb -q
# + id="vg0iv_RPT3YG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 108} outputId="10566000-0a50-4c41-dcfb-b9065768835b"
import wandb
wandb.login()
# + id="_RFZ81yY3Rfl" colab_type="code" colab={}
import torch
from torch import nn
from torch import optim
from torch.nn import functional as F
import torchvision
from torchvision import datasets, transforms
from torch.utils.data import DataLoader
import matplotlib.pyplot as plt
import numpy as np
# + id="NiEHlMTh4ecC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="ee7eaa37-2d6a-46e1-870f-2f81be8b8116"
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
print(device)
# + [markdown] id="XogH13qlOF-Q" colab_type="text"
# # Download Dataset and Prepare Dataloader
# + id="FtM4FMPP3imH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 104, "referenced_widgets": ["cd117ce687f5496bb4cb366fb42eacd1", "2382275b1ef94088a861ca194ee75da7", "<KEY>", "<KEY>", "<KEY>", "cdbe7c33455f49689e89a1b16109eb88", "<KEY>", "0399c819d531404898a23bf1836212e7"]} outputId="043f640e-baa6-4a24-acf5-3133d3a9409e"
BATCH_SIZE = 32
transform = transforms.Compose(
[transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
trainset = torchvision.datasets.CIFAR10(root='./data', train=True,
download=True, transform=transform)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=BATCH_SIZE,
shuffle=True, num_workers=2)
testset = torchvision.datasets.CIFAR10(root='./data', train=False,
download=True, transform=transform)
testloader = torch.utils.data.DataLoader(testset, batch_size=BATCH_SIZE,
shuffle=False, num_workers=2)
CLASS_NAMES = ('plane', 'car', 'bird', 'cat',
'deer', 'dog', 'frog', 'horse', 'ship', 'truck')
# + [markdown] id="XyG3USzkON-d" colab_type="text"
# # Visualize Data
# + id="NWEveFVYN9SE" colab_type="code" colab={}
def show_batch(image_batch, label_batch):
plt.figure(figsize=(10,10))
for n in range(25):
ax = plt.subplot(5,5,n+1)
img = image_batch[n] / 2 + 0.5 # unnormalize
img = img.numpy()
plt.imshow(np.transpose(img, (1, 2, 0)))
plt.title(CLASS_NAMES[label_batch[n]])
plt.axis('off')
# + id="0XVa6VnyOlej" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 591} outputId="f5aeeaf9-8041-413b-b880-59f21339acc5"
sample_images, sample_labels = next(iter(trainloader))
show_batch(sample_images, sample_labels)
# + [markdown] colab_type="text" id="oibAId_GOfJP"
# # Model
# + colab_type="code" id="IzqTsmA_OfJU" colab={}
class Net(nn.Module):
def __init__(self, input_shape=(3,32,32)):
super(Net, self).__init__()
self.conv1 = nn.Conv2d(3, 32, 3)
self.conv2 = nn.Conv2d(32, 64, 3)
self.conv3 = nn.Conv2d(64, 128, 3)
self.pool = nn.MaxPool2d(2,2)
n_size = self._get_conv_output(input_shape)
self.fc1 = nn.Linear(n_size, 512)
self.fc2 = nn.Linear(512, 10)
self.dropout = nn.Dropout(0.25)
def _get_conv_output(self, shape):
batch_size = 1
input = torch.autograd.Variable(torch.rand(batch_size, *shape))
output_feat = self._forward_features(input)
n_size = output_feat.data.view(batch_size, -1).size(1)
return n_size
def _forward_features(self, x):
x = self.pool(F.relu(self.conv1(x)))
x = self.pool(F.relu(self.conv2(x)))
x = self.pool(F.relu(self.conv3(x)))
return x
def forward(self, x):
x = self._forward_features(x)
x = x.view(x.size(0), -1)
x = self.dropout(x)
x = F.relu(self.fc1(x))
x = self.dropout(x)
x = self.fc2(x)
return x
# + [markdown] id="wuhjN4PONzr2" colab_type="text"
# # Train Step
# + id="Z3Cv0XLX54Xo" colab_type="code" colab={}
def train(model, device, train_loader, optimizer, criterion, epoch, steps_per_epoch=20):
# Switch model to training mode. This is necessary for layers like dropout, batchnorm etc which behave differently in training and evaluation mode
model.train()
train_loss = 0
train_total = 0
train_correct = 0
# We loop over the data iterator, and feed the inputs to the network and adjust the weights.
for batch_idx, (data, target) in enumerate(train_loader, start=0):
# Load the input features and labels from the training dataset
data, target = data.to(device), target.to(device)
# Reset the gradients to 0 for all learnable weight parameters
optimizer.zero_grad()
# Forward pass: Pass image data from training dataset, make predictions about class image belongs to (0-9 in this case)
output = model(data)
# Define our loss function, and compute the loss
loss = criterion(output, target)
train_loss += loss.item()
scores, predictions = torch.max(output.data, 1)
train_total += target.size(0)
train_correct += int(sum(predictions == target))
# Reset the gradients to 0 for all learnable weight parameters
optimizer.zero_grad()
# Backward pass: compute the gradients of the loss w.r.t. the model's parameters
loss.backward()
# Update the neural network weights
optimizer.step()
acc = round((train_correct / train_total) * 100, 2)
print('Epoch [{}], Loss: {}, Accuracy: {}'.format(epoch, train_loss/train_total, acc), end='')
wandb.log({'Train Loss': train_loss/train_total, 'Train Accuracy': acc, 'Epoch': epoch})
# + [markdown] id="KeWfs6B9N2uX" colab_type="text"
# # Test Step
# + id="vHcCrrPJ65p-" colab_type="code" colab={}
def test(model, device, test_loader, criterion, classes):
# Switch model to evaluation mode. This is necessary for layers like dropout, batchnorm etc which behave differently in training and evaluation mode
model.eval()
test_loss = 0
test_total = 0
test_correct = 0
example_images = []
with torch.no_grad():
for data, target in test_loader:
# Load the input features and labels from the test dataset
data, target = data.to(device), target.to(device)
# Make predictions: Pass image data from test dataset, make predictions about class image belongs to (0-9 in this case)
output = model(data)
# Compute the loss sum up batch loss
test_loss += criterion(output, target).item()
scores, predictions = torch.max(output.data, 1)
test_total += target.size(0)
test_correct += int(sum(predictions == target))
acc = round((test_correct / test_total) * 100, 2)
print(' Test_loss: {}, Test_accuracy: {}'.format(test_loss/test_total, acc))
wandb.log({'Test Loss': test_loss/test_total, 'Test Accuracy': acc})
# + [markdown] colab_type="text" id="Y5AMPT-jOfJm"
# # Initialize Model, Loss and Optimizer
# + colab_type="code" id="OnkABSPdOfJo" colab={"base_uri": "https://localhost:8080/", "height": 182} outputId="82db0b5a-532d-4200-b609-51a65e3351ec"
net = Net().to(device)
print(net)
criterion = nn.CrossEntropyLoss()
optimizer = optim.Adam(net.parameters())
# + [markdown] colab_type="text" id="a3uSKprMOfJ0"
# #Train
# + colab_type="code" id="nuzN_vcHOfJ3" colab={"base_uri": "https://localhost:8080/", "height": 455} outputId="dd33f7cf-f18d-4084-effc-f669c1e62a59"
wandb.init(entity='authors', project='seo')
wandb.watch(net, log='all')
for epoch in range(20):
train(net, device, trainloader, optimizer, criterion, epoch)
test(net, device, testloader, criterion, CLASS_NAMES)
print('Finished Training')
# + id="N-yq9hDJy1u8" colab_type="code" colab={}
| PyTorch_Classification_Example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] heading_collapsed=true
# # Processing your Data with topsApp.py
#
# **Author**: <NAME> - Jet Propulsion Laboratory
#
# In this notebook, we will walk through the various steps of processing with topsApp.py.
#
# topsApp.py is a pair-by-pair interferometric processor that takes as input two Sentinel-1 SAR acquisitions acquired in TOPS mode. topsApp.py will not work for other Sentinel-1 acquisition formats such as Stripmap and ScanSAR. ISCE's stripmapApp.py supports interferometric stripmap processing of Sentinel-1 and other sensors. At this time, topsApp only supports SLC data from Sentinel-1 A and B. Processing is supported across the Sentinel-1 constellation, i.e. data acquired from A and B can be combined.
#
# To illustrate the usage of topsApp.py, we will use a Sentinel-1 dataset capturing the surface deformation as result of the 12 November 2017 Mw7.3 earthquake that occurred in Iran. The exercise runs the workflow step by step to generate a geocoded interferogram and offsets map.
# + [markdown] heading_collapsed=true hidden=true
# ## 0. Initial setup of the notebook
# + [markdown] hidden=true
# The cell below performs initial setup of the notebook and must be run every time the notebook is used. It is possible to partially complete the exercise, close the notebook, and come back and continue later from that point, but this initialization must be re-run before restarting (as well as Step 1.2.1 if the slc download step 1.2 is skipped). Initialization defines the processing locations as well as a few plotting routines that will be used throughout the tutorial, and this information is lost when the notebook is closed.
# + hidden=true
from shutil import copyfile
from osgeo import gdal ## GDAL support for reading virtual files
import os ## To create and remove directories
import matplotlib.pyplot as plt ## For plotting
import numpy as np ## Matrix calculations
import glob ## Retrieving list of files
## NOTE: When retrieiving the TOPS notebook from a different location,
## it is assumed that you have put it in your home dir under /work/notebooks/TOPS
## Defining the home and data directories at the processing location
home_dir = os.getenv("HOME")
tutorial_home_dir = os.path.abspath(os.path.join(home_dir, "work/notebooks/TOPS"))
slc_dir = os.path.join(tutorial_home_dir,'slc')
processing_DEM_dir = os.path.join(tutorial_home_dir,'DEM/DEM1')
geocoding_DEM_dir = os.path.join(tutorial_home_dir,'DEM/DEM3')
print("home directory: ", tutorial_home_dir)
# generate all the folders in case they do not exist yet
if not os.path.exists(tutorial_home_dir):
os.makedirs(tutorial_home_dir)
if not os.path.exists(slc_dir):
os.makedirs(slc_dir)
if not os.path.exists(processing_DEM_dir):
os.makedirs(processing_DEM_dir)
if not os.path.exists(geocoding_DEM_dir):
os.makedirs(geocoding_DEM_dir)
os.chdir(tutorial_home_dir)
# defining backup dirs in case of download issues on the local server
data_backup_dir = os.path.abspath(os.path.join(home_dir,"course-material/datasets/TOPS"))
slc_backup_dir = os.path.join(data_backup_dir,'slc')
processing_DEM_backup_dir = os.path.join(data_backup_dir,'DEM/DEM1')
geocoding_DEM_backup_dir = os.path.join(data_backup_dir,'DEM/DEM3')
def plotdata(GDALfilename,band=1,title=None,colormap='gray',aspect=1, datamin=None, datamax=None,draw_colorbar=True,colorbar_orientation="horizontal",background=None):
ds = gdal.Open(GDALfilename, gdal.GA_ReadOnly)
data = ds.GetRasterBand(band).ReadAsArray()
transform = ds.GetGeoTransform()
ds = None
# getting the min max of the axes
firstx = transform[0]
firsty = transform[3]
deltay = transform[5]
deltax = transform[1]
lastx = firstx+data.shape[1]*deltax
lasty = firsty+data.shape[0]*deltay
ymin = np.min([lasty,firsty])
ymax = np.max([lasty,firsty])
xmin = np.min([lastx,firstx])
xmax = np.max([lastx,firstx])
# put all zero values to nan and do not plot nan
if background is None:
try:
data[data==0]=np.nan
except:
pass
fig = plt.figure(figsize=(18, 16))
ax = fig.add_subplot(111)
cax = ax.imshow(data, vmin = datamin, vmax=datamax, cmap=colormap,extent=[xmin,xmax,ymin,ymax])
ax.set_title(title)
if draw_colorbar is not None:
cbar = fig.colorbar(cax,orientation=colorbar_orientation)
ax.set_aspect(aspect)
plt.show()
# clearing the data
data = None
def plotcomplexdata(GDALfilename,title=None,aspect=1,datamin=None, datamax=None,draw_colorbar=None,colorbar_orientation="horizontal"):
ds = gdal.Open(GDALfilename, gdal.GA_ReadOnly)
slc = ds.GetRasterBand(1).ReadAsArray()
transform = ds.GetGeoTransform()
ds = None
# getting the min max of the axes
firstx = transform[0]
firsty = transform[3]
deltay = transform[5]
deltax = transform[1]
lastx = firstx+slc.shape[1]*deltax
lasty = firsty+slc.shape[0]*deltay
ymin = np.min([lasty,firsty])
ymax = np.max([lasty,firsty])
xmin = np.min([lastx,firstx])
xmax = np.max([lastx,firstx])
# put all zero values to nan and do not plot nan
try:
slc[slc==0]=np.nan
except:
pass
fig = plt.figure(figsize=(18, 16))
ax = fig.add_subplot(1,2,1)
cax1=ax.imshow(np.abs(slc),vmin = datamin, vmax=datamax, cmap='gray',extent=[xmin,xmax,ymin,ymax])
ax.set_title(title + " (amplitude)")
if draw_colorbar is not None:
cbar1 = fig.colorbar(cax1,orientation=colorbar_orientation)
ax.set_aspect(aspect)
ax = fig.add_subplot(1,2,2)
cax2 =ax.imshow(np.angle(slc),cmap='rainbow',extent=[xmin,xmax,ymin,ymax])
ax.set_title(title + " (phase [rad])")
if draw_colorbar is not None:
cbar2 = fig.colorbar(cax2,orientation=colorbar_orientation)
ax.set_aspect(aspect)
plt.show()
# clearing the data
slc = None
def plotstackdata(GDALfilename_wildcard,band=1,title=None,colormap='gray',aspect=1, datamin=None, datamax=None,draw_colorbar=True,colorbar_orientation="horizontal"):
# get a list of all files matching the filename wildcard criteria
GDALfilenames = glob.glob(os.path.abspath(GDALfilename_wildcard))
# initialize empty numpy array
for GDALfilename in GDALfilenames:
ds = gdal.Open(GDALfilename, gdal.GA_ReadOnly)
data_temp = ds.GetRasterBand(band).ReadAsArray()
ds = None
try:
data
except NameError:
data = data_temp
else:
data = np.vstack((data,data_temp))
# put all zero values to nan and do not plot nan
try:
data[data==0]=np.nan
except:
pass
fig = plt.figure(figsize=(18, 16))
ax = fig.add_subplot(111)
cax = ax.imshow(data, vmin = datamin, vmax=datamax, cmap=colormap)
ax.set_title(title)
if draw_colorbar is not None:
cbar = fig.colorbar(cax,orientation=colorbar_orientation)
ax.set_aspect(aspect)
plt.show()
# clearing the data
data = None
def plotstackcomplexdata(GDALfilename_wildcard,title=None,aspect=1, datamin=None, datamax=None,draw_colorbar=True,colorbar_orientation="horizontal"):
# get a list of all files matching the filename wildcard criteria
GDALfilenames = glob.glob(os.path.abspath(GDALfilename_wildcard))
# initialize empty numpy array
for GDALfilename in GDALfilenames:
ds = gdal.Open(GDALfilename, gdal.GA_ReadOnly)
data_temp = ds.GetRasterBand(1).ReadAsArray()
ds = None
try:
data
except NameError:
data = data_temp
else:
data = np.vstack((data,data_temp))
# put all zero values to nan and do not plot nan
try:
data[data==0]=np.nan
except:
pass
fig = plt.figure(figsize=(18, 16))
ax = fig.add_subplot(1,2,1)
cax1=ax.imshow(np.abs(data),vmin = datamin, vmax=datamax, cmap='gray')
ax.set_title(title + " (amplitude)")
if draw_colorbar is not None:
cbar1 = fig.colorbar(cax1,orientation=colorbar_orientation)
ax.set_aspect(aspect)
ax = fig.add_subplot(1,2,2)
cax2 =ax.imshow(np.angle(data),cmap='rainbow')
ax.set_title(title + " (phase [rad])")
if draw_colorbar is not None:
cbar2 = fig.colorbar(cax2,orientation=colorbar_orientation)
ax.set_aspect(aspect)
plt.show()
# clearing the data
data = None
# + [markdown] heading_collapsed=true hidden=true
# ## 1. Overview of the tutorial input dataset
# + [markdown] hidden=true
# Let us first take a look at the dataset. For our dataset we are focusing on Ascending track 72 and acquisitions dates of 20171112 and 20171118. As the earthquake occurs along the frame boundaries we will use multiple Sentinel-1 SLC products for each acquisition.
#
# 
# + [markdown] heading_collapsed=true hidden=true
# ### 1.1 Background on TOPS mode
# + [markdown] hidden=true
# The TOPS acquisition strategy is different than conventional stripmap mode. TOPS stands for Terrain Observation with Progressive Scans. As the name indicates, the radar sensor performs a scan of the surface by electronically steering the antenna beam from a backward-pointing along-track direction to a forward-pointing along-track direction for a fixed range swath (also called a subswath).
# 
# After a successful scan at that range extent, the antenna beam is electronically rolled back to its initial position, the range swath is electronically directed outward to a new area to increase coverage, and the next scan is made from backward to forward along track at this new range swath. After a third scan at a third range swath, the entire process is repeated over and over to create a continuous image as the satellite flies along. The timing of the scans is such that there is a small geographic overlap between successive scans at a given range to ensure continuous coverage. Each scan at a given range swath is known as a "burst". When inspecting one of the downloaded SLC products you will note that data are provided in 3 individual subswaths (IW1, IW2, IW3) each with a set of bursts. All together they form a Sentinel-1 frame, typically ~250 x 250 km in size. The bursts are approximately 20 km in length and overlap by 2 km. Due to the TOPS acquisition mode, the overlap region in successive bursts is seen from two different directions (forward looking and backward looking).
# 
# The ESA website provides detailed background information on Sentinel-1 products and technical information of the TOPS sensors. Whenever during the practical you are waiting for the processing to complete you could explore their webpage at https://sentinel.esa.int/web/sentinel/user-guides/sentinel-1-sar
# + [markdown] heading_collapsed=true hidden=true
# ### 1.2 SLC download
# + [markdown] hidden=true
# The ASF vertex page (https://www.asf.alaska.edu/sentinel/) and the SSARA GUI (http://web-services.unavco.org/brokered/ssara/gui) both offer a GUI to visually search for available Sentinel-1 data over your area of interest. Once you have found your data, you can download it from the GUI. ASF provides a bulk-download python script.
#
# Alternatively, you can use SSARA from command line.
# + hidden=true
# !git clone https://github.com/bakerunavco/SSARA.git
# + [markdown] hidden=true
# Running the client without any options will print the help message and describe the options. Some usage examples are given in the help as well.
# + hidden=true
# !SSARA/ssara_federated_query.py
# + [markdown] hidden=true
# Take some time to perform some example searches in a terminal window to get a feel from how the client works.
# <br>
# <div class="alert alert-warning">
# <b>TIP:</b>
# In case you are not sure about some of the SSARA API fields (e.g. the platform name, the polygon etc.), you can experiment with the SSARA GUI first. One potential solution is to draw a polygon without specifying much additional information, see what the search table returns for keywords, and use those on the command line API.
# </div>
#
# The following command provides the necessary filters for the search and returns the 4 scenes we need for processing:
# + hidden=true
# !SSARA/ssara_federated_query.py --platform=sentinel-1B,sentinel-1A --relativeOrbit=72 -s 2017-11-11 -e 2017-11-18 --intersectsWith='POLYGON((45 35.25,45 34,46.5 34,46.5 35.25,45 35.25))' --print
# + [markdown] hidden=true
# By changing the **--print** to **--download** you can download the files once you set up your password and username for SSARA in the **password_config.py** file.
#
# Below we include a small python script to download the data which allows you to directly parse your pasword and username at the top of the function (this will take a few minutes for each file).
# + hidden=true
ASF_USER = ""
ASF_PASS = ""
files = ['https://datapool.asf.alaska.edu/SLC/SA/S1A_IW_SLC__1SDV_20171111T150004_20171111T150032_019219_0208AF_EE89.zip',
'https://datapool.asf.alaska.edu/SLC/SA/S1A_IW_SLC__1SDV_20171111T150029_20171111T150056_019219_0208AF_BF55.zip',
'https://datapool.asf.alaska.edu/SLC/SB/S1B_IW_SLC__1SDV_20171117T145900_20171117T145928_008323_00EBAB_B716.zip',
'https://datapool.asf.alaska.edu/SLC/SB/S1B_IW_SLC__1SDV_20171117T145926_20171117T145953_008323_00EBAB_AFB8.zip']
if len(ASF_USER)==0 or len(ASF_PASS)==0:
raise Exception("Specifiy your ASF password and user (earthdata log-in)")
for file in files:
filename = os.path.basename(file)
if not os.path.exists(os.path.join(slc_dir,filename)):
cmd = "wget {0} --user={1} --password={2}".format(file,ASF_USER, ASF_PASS)
print(cmd)
os.chdir(slc_dir)
os.system(cmd)
else:
print(filename + " already exists")
os.chdir(tutorial_home_dir)
# + [markdown] hidden=true
# Files size are around 4.3GB
# + hidden=true
# ls -lh slc
# + [markdown] heading_collapsed=true hidden=true
# #### Download issues
# + [markdown] hidden=true
# In case the download does not work, we have pre-downloaded the required files. Run the routine below to copy these files into your work directory.
# + hidden=true
files = ['https://datapool.asf.alaska.edu/SLC/SA/S1A_IW_SLC__1SDV_20171111T150004_20171111T150032_019219_0208AF_EE89.zip',
'https://datapool.asf.alaska.edu/SLC/SA/S1A_IW_SLC__1SDV_20171111T150029_20171111T150056_019219_0208AF_BF55.zip',
'https://datapool.asf.alaska.edu/SLC/SB/S1B_IW_SLC__1SDV_20171117T145900_20171117T145928_008323_00EBAB_B716.zip',
'https://datapool.asf.alaska.edu/SLC/SB/S1B_IW_SLC__1SDV_20171117T145926_20171117T145953_008323_00EBAB_AFB8.zip']
for file in files:
filename = os.path.basename(file)
if not os.path.exists(os.path.join(slc_dir,filename)):
cmd = 'cp -r ' + os.path.join(slc_backup_dir,filename) + " " + slc_dir
os.system(cmd)
print(filename + " done")
else:
print(filename + " already exists")
os.chdir(tutorial_home_dir)
# + [markdown] hidden=true
# Files size are around 4.3GB
# -
# ### 1.2.1 Set/Reset to home directory
#
# This step is required each time the notebook is restarted and the slc download is skipped. It can also be run in the initial run, though it is not needed since the directory is reset during the slc download process.
os.chdir(tutorial_home_dir)
# + hidden=true
# !ls -lh slc
# + [markdown] heading_collapsed=true hidden=true
# ### 1.3 SLC filenaming convention
# + [markdown] hidden=true
# TOPS SLC product files delivered from ESA are zip archives. When unpacked the zip extension will be replaced by SAFE. The products are therefore also frequently called SAFE files. topsApp.py can read the data from either a zip file or a SAFE file. To limit disk usage, it is recommended to not unzip the individual files.
#
# The zip or SAFE filenames provide information on the product type, the polarization, and the start and stop acquisition time. For example: S1A_IW_SLC__1SDV_20171111T150004_20171111T150032_019219_0208AF_EE89.zip
# - Type = slc
# - Polarization = Dual pole
# - Date = 20171111
# - UTC time of acquisition = ~15:00
# - Sensing start for the acquisition was 20171111 at 15:00:04
#
# + [markdown] heading_collapsed=true hidden=true
# ### 1.3 Orbits and Instrument file download
# + [markdown] hidden=true
# In addition to the **SAFE files**, **orbit files** and the **auxiliary instrument files** are required for ISCE processing. Both the orbit and instrument files are provided by ESA and can be downloaded at: https://qc.sentinel1.eo.esa.int/.
#
# We have pre-downloaded the precise orbit files as well as the instrument files for our test dataset. Run the routine below to copy these files into the aux folder.
# + hidden=true
aux_dir = os.path.join(tutorial_home_dir,'aux')
files = ['aux_cal/S1A_AUX_CAL_V20140908T000000_G20140909T130257.SAFE',
'aux_cal/S1A_AUX_CAL_V20140915T100000_G20151125T103928.SAFE',
'aux_poeorb/S1A_OPER_AUX_POEORB_OPOD_20171201T121227_V20171110T225942_20171112T005942.EOF',
'aux_poeorb/S1B_OPER_AUX_POEORB_OPOD_20171207T111223_V20171116T225942_20171118T005942.EOF',]
for file in files:
if not os.path.exists(os.path.join(aux_dir,file)):
cmd = "cp -r " + os.path.join(data_backup_dir,'aux',file) + " " + os.path.join(aux_dir,file)
os.system(cmd)
print(file + " done")
else:
print(file + " already exists")
os.chdir(tutorial_home_dir)
# + hidden=true
# ls aux/*
# + [markdown] hidden=true
# Although Sentinel-1 restituted orbits (RESORB) are of good quality, it is recommended to use the precise orbits (POEORB) when available. Typically, precise orbits are available with a 15 to 20-day lag from the day of the acquisition.
#
# <div class="alert alert-danger">
# <b>AUX_CAL:</b>
# AUX_CAL information is used for **antenna pattern correction** of SAFE products with **IPF verison 002.36**. If all your SAFE products are from another IPF version, then no AUX files are needed.
# </div>
#
# + [markdown] heading_collapsed=true hidden=true
# ## 2. topsApp.py input variables
# + [markdown] hidden=true
# Like the other apps in ISCE, the input variables to topsApp.py are controlled through an app xml file. All apps in ISCE have example xml files included in the ISCE distribution. You can find these under **/examples/input_files**. For convenience, we have included the *topsApp.xml* and *master_TOPS_SENTINEL1.xml* example in the support_docs folder.
# + [markdown] heading_collapsed=true hidden=true
# ### 2.1 Required versus optional topsApp.py inputs
# + [markdown] hidden=true
# The example *topsApp.xml* contains all input variables with a description. Let us first read this file. You can open the file by launching a **terminal** and using your preferred editor to open the file. For vim type:
# ```
# vim support_docs/example/topsApp.xml
# vim support_docs/example/master_TOPS_SENTINEL1.xml
# ```
# When it comes to the actual processing with topsApp.py, you do not need to specify all the input variables as shown in the example topsApp.xml. Defaults will be assumed when properties are not set by the user. You can get a simple table overview of the required variables by calling the help of topsApp.py.
# + hidden=true
# !topsApp.py --help
# + [markdown] hidden=true
# From the table, you can see that the master and slave components are to be specified. This can be done directly with its specific properties in the topsApp.xml, or alternatively, one can point to a dedicated xml file for the master and slave, each of which contain their individual properties. The required properties for the master and slave should at least include orbit information as "orbit directory", the auxiliary instrument information as "auxiliary data directory", and a list of products under the "safe" property tag.
#
# <br>
# <div class="alert alert-danger">
# <b>POTENTIAL ISSUE:</b>
# If you specify a **region of interest**, make sure it covers at last two bursts. If this is not the case, ESD will fail in the processing as it is being estimated from the burst overlap region. You could always decide to only geocode a smaller region with the ** ** property.
# </div>
# + [markdown] heading_collapsed=true hidden=true
# ### 2.2 topsApp inputs for this tutorial
# + [markdown] hidden=true
# Let us now examine the files *topsApp.xml*, *master.xml* and *slave.xml* that will be used for this tutorial.
# + [markdown] heading_collapsed=true hidden=true
# #### topsApp.xml
# + [markdown] hidden=true
# ```xml
# <?xml version="1.0" encoding="UTF-8"?>
# <topsApp>
# <component name="topsinsar">
# <property name="Sensor name">SENTINEL1</property>
# <component name="master">
# <catalog>master.xml</catalog>
# </component>
# <component name="slave">
# <catalog>slave.xml</catalog>
# </component>
# <property name="swaths">[3]</property>
# <property name="range looks">19</property>
# <property name="azimuth looks">7</property>
# <property name="region of interest">[34, 35.25 , 44.75, 46.5]</property>
# <!--<property name="geocode demfilename">../../DEM/DEM3/Coarse_demLat_N33_N36_Lon_E045_E047.dem.wgs84</property>-->
# <!--<property name="demFilename">../../DEM/DEM1/demLat_N33_N36_Lon_E045_E047.dem.wgs84</property>-->
# <property name="do unwrap">True</property>
# <property name="unwrapper name">snaphu_mcf</property>
# <!--<property name="do unwrap2stage">True</property>-->
# <property name="do denseoffsets">True</property>
# <!--property name="geocode list">['merged/phsig.cor', 'merged/filt_topophase.unw', 'merged/los.rdr', 'merged/topophase.flat', 'merged/filt_topophase.flat','merged/topophase.cor','merged/filt_topophase.unw.conncomp']</property>-->
# </component>
# </topsApp>
# ```
#
#
# - The master and slave components refer to their own *.xml* files
# - The **swaths** property controls the number of swaths to be processed. As the earthquake occurred in the subswath three, we can directly limit the list to the single entry list [3] only.
# - We specify the **range looks** and **azimuth looks** to be 19 and 7. The range resolution for sentinel varies from near to far range, but is roughly 5m, while the azimuth resolution is approximately 15m, leading to a multi-looked product that will be approximately 95m by 105m.
# - By specifying the **region of interest** as [S, N, W, E] to only capture the extent of the earthquake, topsApp.py will only extract those bursts from subswath 3 needed to cover the earthquake.
# - By default, topsApp can download a DEM on the fly. By including **demFilename** a local DEM can be specified as input for the processing. For this notebook exercise, a sample 1-arc second DEM in WGS84 is provided in the DEM/DEM1 folder.
# - By default, the geocoding in topsApp.py is performed at the same sampling as processing DEM. However, a different DEM *to be used specifically for geocoding* can be specified using the **geocode demfilename** property. In this notebook, we provide a DEM in WGS84 sampled at 3-arc seconds to be used specifically for geocoding. It is stored in the DEM/DEM3 folder.
# - By default, no unwrapping is done. In order to turn it on, set the property **do unwrap** to *True*.
# - In case unwrapping is requested, the default unwrapping strategy to be applied is the *icu* unwrapping method. For this tutorial, we will use *snaphu_mcf*.
# - Lastly, we request topsApp.py to run the dense-offsets using the **do denseoffsets** property. By enabling this, topsApp.py will estimate the range and azimuth offsets on the amplitude of the master and slave SLC files.
#
#
# You will see that a few of the above properties are commented out in the xml files provided. We will come back to these later in the tutorial. The commented properties have the form:
# ```xml
# <!--<property> ... </property>-->
# ```
# + [markdown] heading_collapsed=true hidden=true
# #### master.xml and slave.xml
# + [markdown] hidden=true
# **master.xml**
# ``` xml
# <component name="master">
# <property name="orbit directory">../../aux/aux_poeorb</property>
# <property name="auxiliary data directory">../../aux/aux_cal</property>
# <property name="output directory">master</property>
# <property name="safe">['../../slc/S1B_IW_SLC__1SDV_20171117T145900_20171117T145928_008323_00EBAB_B716.zip','../../slc/S1B_IW_SLC__1SDV_20171117T145926_20171117T145953_008323_00EBAB_AFB8.zip']</property>
# </component>
# ```
#
# **slave.xml**
# ``` xml
# <component name="slave">
# <property name="orbit directory">../../aux/aux_poeorb</property>
# <property name="auxiliary data directory">../../aux/aux_cal</property>
# <property name="output directory">slave</property>
# <property name="safe">['../../slc/S1A_IW_SLC__1SDV_20171111T150004_20171111T150032_019219_0208AF_EE89.zip']</property>
# </component>
# ```
#
# - The value associated with the master **safe** property corresponds to a list of SAFE files acquired on 20171117. For the slave only, one SAFE files is included, acquired on 20171111. The paths to each SAFE file points to the slc directory.
# - The **orbit directory** and **auxiliary data directory** point respectively to the directory where we have stored the POEORB (precise) orbits for the example, and the Auxiliary instrument files for Sentinel-1.
#
#
# <br>
# <div class="alert alert-warning">
# <b>SIGN CONVENTION:</b>
# By selecting the master to be acquired after the slave, and keeping in mind that the interferogram formation is master* conj(slave), then a positive phase value for the interferogram indicates the surface has moved away from the satellite between 20171111 and 20171117.
# </div>
#
# + [markdown] heading_collapsed=true hidden=true
# ## 3. topsApp.py processing steps
# + [markdown] hidden=true
# The topsApp.py workflow can be called with a single command-line call to topsApp.py; by default it will run all the required processing steps with inputs pulled from the topsApp.xml file. Although this is an attractive feature, it is recommended to run topsApp.py with “steps” enabled. This will allow you to re-start the processing from a given processing step. If “steps” are not used, users must restart processing from the beginning of the workflow after fixing any downstream issues with the processing.
#
# The "--help" switch lists all the steps involved in the processing:
# + hidden=true
# !topsApp.py --help --steps
# + [markdown] hidden=true
# <br>
# <div class="alert alert-danger">
# <b>POTENTIAL ISSUE:</b>
# **Steps are to be run in the following prescribed order**:
# *<center>
# ['startup', 'preprocess', 'computeBaselines', 'verifyDEM', 'topo']
# ['subsetoverlaps', 'coarseoffsets', 'coarseresamp', 'overlapifg', 'prepesd']
# ['esd', 'rangecoreg', 'fineoffsets', 'fineresamp', 'burstifg']
# ['mergebursts', 'filter', 'unwrap', 'unwrap2stage', 'geocode']
# ['denseoffsets', 'filteroffsets', 'geocodeoffsets']
# </center>*
# </div>
#
# + [markdown] heading_collapsed=true hidden=true
# ### 3.0 Directory setup
# + [markdown] hidden=true
# Before we begin step by step processing with topsApp.py, we first setup the processing directory in the insar folder as **masteryyyymmdd_slaveyyyymmdd**, and copy the template topsApp.xml file (*ls support_docs/insar/topsApp.xml*) to the processing location.
# + hidden=true
import os
from shutil import copyfile
## Directory where we will perform the topsApp processing
processing_dir = os.path.join(tutorial_home_dir,'insar/20171117_20171111')
## Template xml for this tutorial
topsAppXml_original = os.path.join(tutorial_home_dir,'support_docs/insar/topsApp.xml')
masterXml_original = os.path.join(tutorial_home_dir,'support_docs/insar/master.xml')
slaveXml_original = os.path.join(tutorial_home_dir,'support_docs/insar/slave.xml')
## Check if output directory already exists. If not create it
if not os.path.isdir(processing_dir):
os.makedirs(processing_dir)
## Check if the topsApp.xml file already exists, if not copy the example for the excerisize
if not os.path.isfile(os.path.join(processing_dir,'topsApp.xml')):
copyfile(topsAppXml_original,os.path.join(processing_dir,'topsApp.xml'))
else:
print(os.path.join(processing_dir,'topsApp.xml') + " already exist, will not overwrite")
if not os.path.isfile(os.path.join(processing_dir,'master.xml')):
copyfile(masterXml_original,os.path.join(processing_dir,'master.xml'))
else:
print(os.path.join(processing_dir,'master.xml') + " already exist, will not overwrite")
if not os.path.isfile(os.path.join(processing_dir,'slave.xml')):
copyfile(slaveXml_original,os.path.join(processing_dir,'slave.xml'))
else:
print(os.path.join(processing_dir,'slave.xml') + " already exist, will not overwrite")
# + [markdown] hidden=true
# We will now "cd" into this directory where the processing will be done.
# + hidden=true
os.chdir(processing_dir)
# + [markdown] heading_collapsed=true hidden=true
# ### 3.1 Step Startup
# + [markdown] hidden=true
# We will start with running the first step **startup**.
# + hidden=true
# !topsApp.py --dostep=startup
# + [markdown] hidden=true
# When topsApp.py is run in steps, PICKLE files are used to store state information between the steps. A PICKLE folder is created during the startup step. The PICKLE folder is used to store the processing parameters for each processing step. By exploring the pickle folder, you will find a binary pickle file and an *xml* file associated with the **startup** step.
# + hidden=true
# ls PICKLE
# + [markdown] hidden=true
# The information contained within the **startup** pickle and *xml* files allows topsApp to start or re-start processing from where **startup** was completed.
# + [markdown] heading_collapsed=true hidden=true
# ### 3.2 Step Preprocess
# + [markdown] hidden=true
# Keeping in mind that the order of steps matter, we move to the second step of topsApp.py processing, which is to **preprocess** the data.
# + hidden=true
# !topsApp.py --dostep=preprocess
# + [markdown] hidden=true
# During preprocessing, the orbits, the IPF (Instrument Processing Facility, which is the processing version used by ESA to focus the data), the bursts, and, if needed, the antenna pattern are extracted.
#
# Note if you had processed the complete region without limitation, there would be three subswaths (IW1,IW2,IW3). For our example,
# we limited the processing to IW3 alone. Therefore, for our tutorial, within the slave and master folder you will find only an **IW3** folder and an **IW3.xml** file.
# + hidden=true
# ls master/*
# + [markdown] hidden=true
# The **IW3.xml** file contains metadata information specific for each subswath (e.g. orbit state vectors, doppler, sensing times, IPF version). You can open the file using your preferred editor from the command line:
# ```
# vim master/IW3.xml
# ```
# The **IW3** folder contains the unpacked bursts and their meta-data.
# Typically you will find *.xml* and *.vrt* files, and in certain cases *.slc* files, as described below.
# - If a bounding box is specified, only the master bursts covering the box are unpacked, otherwise the complete master is used. The bursts of the master in master/IW* are relabeled from 1 to n.
# - The slave/IW* folders typically contain a larger set of bursts to cover the master extent completely, where the bursts are labelled from 1 to m. The burst numbering is not coordinated between master and slave folders: burst 1 in the master does not necessarily correspond to burst 1 in the slave.
# - All data are unpacked virtually (i.e., only *.xml* and *.vrt* files in *IW* folders) unless: (1) user requested to physically unpack the data, which can be requested in topsApp.xml by property **usevirtualfiles**, or (2) if an antenna pattern correction needs to be applied (not controlled by user) which happens mainly for the initially acquired Sentinel-1 data.
# - The IPF version of unpacked bursts is tracked and a combined IFP is assigned for the master and one for the slave. Note: bursts with varying IPF cannot be stitched and attempting to do so will cause an error message.
#
# <br>
# <div class="alert alert-danger">
# <b>POTENTIAL ISSUE:</b>
# There is a gap in spatial coverage, e.g. you are missing the middle SAFE file.
# </div>
# <div class="alert alert-danger">
# <b>POTENTIAL ISSUE:</b>
# The SAFE frames that need to be stitched do not have a consistent IPF version
# </div>
# + [markdown] heading_collapsed=true hidden=true
# ### 3.3 Step computeBaselines
# + hidden=true
# !topsApp.py --dostep=computeBaselines
# + [markdown] hidden=true
# In this step, the perpendicular and parallel baselines are computed using the orbit (state vector information) for the first and last burst of the master image (center range). Each subswath is processed individually, with output sent to the screen. As the processing for the tutorial is limited to IW3, the baselines are only computed for this subswath.
# + [markdown] heading_collapsed=true hidden=true
# ### 3.4 Step verifyDEM
# + hidden=true
# !topsApp.py --dostep=verifyDEM
# + [markdown] hidden=true
# This step will check the DEM file specified in the topsApp.xml. If no DEM file has been specified, topsApp.py will download the DEM on the fly and track the filename for subsequent processing.
#
# <div class="alert alert-danger">
# <b>POTENTIAL ISSUE:</b>
# You did not set your earthdata credentials as detailed in the ISCE installation.
# </div>
# <div class="alert alert-danger">
# <b>POTENTIAL ISSUE:</b>
# The DEM ftp site is down and returns no data tiles.
# </div>
#
# + [markdown] heading_collapsed=true hidden=true
# #### Directly move to 3.5 if verifyDEM does not fail:
# + [markdown] hidden=true
# If your DEM download failed and you are unable to resolve the issue, you can use the DEM provided with this tutorial. This requires three steps to complete, before you can rerun the **verifyDEM** step:
#
# 1) Copy over the DEM from the backup folder to your processing location
# + hidden=true
cmd = "cp -r " + os.path.join(processing_DEM_backup_dir,'*') + " " + processing_DEM_dir
print(cmd)
os.system(cmd)
print('done')
# + [markdown] hidden=true
# 2) To use this DEM, you will need to edit the **topsApp.xml** file and uncomment the property for the **demFilename**.
# ``` vim
# vim insar/20171117_20171111/topsApp.xml
# ```
# Remember the *xml* guidelines:
# ```xml
# <!--<property> ..COMMENTED.. </property>-->
# <property> ..UNCOMMENTED.. </property>
# ```
#
# 3) Update the DEM metadata information. In the processing the DEM metadata such as the DEM path is read from the *xml* information. As the DEM was copied from another machine, we need to update the path. **fixImageXml.py** allows you to do this from the command line.
# + hidden=true
# !fixImageXml.py -h
# + [markdown] hidden=true
# From the help, you can see that the **-f** option will update the path in the xml file with an absolute path. Now let us run the command:
# + hidden=true
os.chdir(processing_DEM_dir)
# !fixImageXml.py -i demLat_N33_N36_Lon_E045_E047.dem.wgs84 -f
os.chdir(processing_dir)
# + [markdown] hidden=true
# Now rerun the **verifyDEM** step to ensure the new DEM information is correctly loaded into the processing:
# + hidden=true
# !topsApp.py --dostep=verifyDEM
# + [markdown] heading_collapsed=true hidden=true
# ### 3.5 Step topo
# + [markdown] hidden=true
# During topo processing, the DEM is mapped into the radar coordinates of the master. As output this generates the **master_geom** folder containing the longitude (*lon_XX.rdr*), latitude (*lat_XX.rdr*), height (*hgt_XX.rdr*) and LOS angles (*los_XX.rdr*) on a pixel by pixel grid for each burst (*XX*). This step is the most time-consuming step. It is parallelized for performance and can also be ran with GPU support enabled.
#
# For our tutorial, anticipate this step will take 20+ min in CPU mode with 4 treads.
# ***Depending on the size of the class the instructor might recommend to decrease the number of treads, pair up in teams, or stagger this processing step***.
#
# This is a good opportunity to familiarize yourself a bit more with the TOPS mode and input parameters.
# - Can you find which property in the topsApp.xml controls the GPU processing? Tip: see “topsApp.py --help”.
# - Can you find the typical incidence angle range for IW3? Tip: You could try to load the los.rdr file for a burst or search the ESA website for TOPS documentation.
#
# Once the step is complete you can have a look at the files that have been generated for each burst:
# + hidden=true
# !export OMP_NUM_TREADS=4; topsApp.py --dostep=topo
# + hidden=true
# ls geom_master/*
# + [markdown] hidden=true
# Were you able to figure out the incidence angle for subswath 3? It is about 41$^\circ$-46$^\circ$. This infomation is contained in the *los.XX.rdr* files. Its first band contains the incidence angle and the second band the azimuth angle of the satellite. There is an easy way to retrieve its average value with GDAL:
# + hidden=true
# !gdalinfo -stats geom_master/IW3/los_07.rdr.vrt
# + hidden=true
plotdata('geom_master/IW3/hgt_01.rdr',1,'IW3: Height of Burst 1 [meter]','terrain')
plotdata('geom_master/IW3/los_01.rdr',1,'IW3: Incidence angle of Burst 1 [degrees]','jet')
plotdata('geom_master/IW3/los_01.rdr',2,'IW3: Azimuth angle of Burst 1 [degrees]','jet')
# + [markdown] heading_collapsed=true hidden=true
# ### 3.6 Step subsetoverlaps
# + [markdown] hidden=true
# Due to the large doppler frequency variation inherent to the TOPS acquisition mode, images supporting interferometry must be coregistered in the azimuth direction to better than 0.001 pixels, compared to regular stripmap data (0.1 of a pixel). While conventional cross-correlation of the amplitude works for the range direction, it does not provide sufficient accuracy for azimuth. The solution is to apply an Enhanced Spectral Diversity (ESD) approach to estimate the azimuth coregistration. In ESD processing, a double difference interferogram is made between the master and slave in the burst overlap region. As the range displacement is cancelled out, this interferogram will only show azimuthal motion. In absence of large ground deformation this azimuthal motion can be interpreted as an azimuthal coregistration offset.
#
# The following processing steps up to ESD are specific to burst overlap regions alone.
#
# By running the **subsetoverlaps** step, the top and bottom overlap between bursts is computed for the master geometry. The information is then stored within the *overlaps* folder of the master directory.
#
# Note, the nomenclature "top" and "bottom" can be confusing. In this case, it is **not** referring to the top and bottom of the burst. It is referring to burst *n* as "top" and burst *n+1* as "bottom", visualizing it as the earlier image being laid on top of the later image, which is on the bottom.
#
# 
# (Figure from Fattahi et. al., SCEC)
# + hidden=true
# !topsApp.py --dostep=subsetoverlaps
# + hidden=true
# ls master/overlaps/*
# + [markdown] hidden=true
# For each subswath being processed, you will find a *bottom_IW[].xml* and a *top_IW[].xml* file, associated with the overlap region in the the bottom and top overlapping bursts, respectively. Like before, only a IW3 folder is present for this tutorial. If you explore the file, you will find additional information such as the FM rate and the doppler information.
# ```
# vim master/overlaps/bottom_IW3.xml
# ```
# Within the *overlaps* directory you will also find for each processed subswath a folder containing the cropped SLC's for each burst overlap region. The convention for top and bottom is as follows:
# - Burst_bot_01_02.slc and Burst_top_01_02.slc both refer to the same burst overlap region, where:
# - Burst_bot_01_02.slc is the part of burst 2 (bot burst)
# - Burst_top_01_02.slc is the part of burst 1 (top burst)
# - Burst_bot_02_03.slc and Burst_top_02_03.slc both refer to the same burst overlap region, where:
# - Burst_bot_02_03.slc is the part of burst 3
# - Burst_top_02_03.slc is the part of burst 2
# - etc...
#
# Though the overlap is not large, we will try to visualize the overlap by comparing:
# - Burst_bot_01_02.slc and Burst_top_01_02.slc
#
# Can you spot the overlap with respect to the full burst?
# + hidden=true
plotcomplexdata('master/overlaps/IW3/burst_bot_01_02.slc.vrt','bot_01_02 (from Burst 2)',aspect=10,datamin=0,datamax=150)
plotcomplexdata('master/overlaps/IW3/burst_top_01_02.slc.vrt','top_01_02 (from Burst 1)',aspect=10,datamin=0,datamax=150)
plotcomplexdata('master/IW3/burst_01.slc.vrt','Burst 1',aspect=10,datamin=0,datamax=150)
plotcomplexdata('master/IW3/burst_02.slc.vrt','Burst 2',aspect=10,datamin=0,datamax=150,draw_colorbar=True,colorbar_orientation="horizontal")
# + [markdown] heading_collapsed=true hidden=true
# ### 3.7 Step coarseoffsets
# + [markdown] hidden=true
# A coarse coregistration can be done using the orbits alone. This is also referred to as geometric coregistration. As ESD is applied in burst overlap region, there is no need to apply this **coarseoffsets** estimation step to the complete SLC. Instead, we estimate the coarse offset for each burst overlap as identified in the previous step.
# + hidden=true
# !topsApp.py --dostep=coarseoffsets
# + [markdown] hidden=true
# As output this generates a "*coarse_offsets/overlaps*" folder, within which is a subswath breakdown (*IW*) containing pixel-by-pixel azimuth offsets (*azimuth_[top,bot]_XX_YY.off*) and range offsets (*range_[top,bot]_XX_YY.off*) for each burst overlap.
# + hidden=true
# ls coarse_offsets/overlaps/IW3/*
# + [markdown] hidden=true
# Let us plot the average azimuth offset is for burst overlap: bot_01_02
# + hidden=true
plotdata('coarse_offsets/overlaps/IW3/azimuth_bot_01_02.off.vrt',1,'Azimuth bot_01_02','gray',aspect=10)
# !gdalinfo -stats coarse_offsets/overlaps/IW3/azimuth_bot_01_02.off.vrt
# + [markdown] hidden=true
# Now do the same for the range offset.
# + hidden=true
plotdata('coarse_offsets/overlaps/IW3/range_bot_01_02.off.vrt',1,'Range bot_01_02','gray',aspect=10)
# !gdalinfo -stats coarse_offsets/overlaps/IW3/range_bot_01_02.off.vrt
# + [markdown] heading_collapsed=true hidden=true
# ### 3.8 Step coarseresamp
# + [markdown] hidden=true
# Both master and slave need to be in a common geometry prior to interferogram formation. In this step we will be taking the outputs from the coarse offset estimation and resample the slave burst overlaps into the master geometry.
# + hidden=true
# !topsApp.py --dostep=coarseresamp
# + [markdown] hidden=true
# The output of this step is stored in the "*coarse_coreg/overlaps/*" folder for each subswath.
# + hidden=true
# ls coarse_coreg/overlaps/IW3/*
# + [markdown] heading_collapsed=true hidden=true
# ### 3.9 Step overlapifg
# + [markdown] hidden=true
# Now that the master and slave burst overlap SLC's are in the same geometry we can compute the interferogram between the master and the slave. As there is a bottom and top set of SLC's, there will be two interferograms for each burst overlap:
# $$\Delta\Phi_{\text{bot}} = \Phi_{\text{bot}}^{\text{master}}*conj\left( \Phi_{\text{bot}}^{\text{slave}}\right) $$
# $$\Delta\Phi_{\text{top}} = \Phi_{\text{top}}^{\text{master}}*conj\left( \Phi_{\text{top}}^{\text{slave}}\right) $$
# + hidden=true
# !topsApp.py --dostep=overlapifg
# + [markdown] hidden=true
# As output this generates a "*coarse_interferogram/overlaps*" folder, with in it a subswath breakdown (*IW*) containing the inteferograms (*burst_[top,bot]_XX_YY.int*) and multi-looked inteferograms (*burst_[top,bot]_XX_YY.7alks_19rlks.int*) for each burst overlap.
# + hidden=true
# ls coarse_interferogram/overlaps/IW3*
# + [markdown] hidden=true
# Let us plot all the multi-looked interferograms for the "top burst" overlaps together.
# + hidden=true
plotstackcomplexdata('coarse_interferogram/overlaps/IW3/burst_top_*7alks_19rlks.int',title="Top overlaps ",aspect=10, datamin=0, datamax=6000,draw_colorbar=True,colorbar_orientation="horizontal")
# + [markdown] hidden=true
# As you can see there is a large fringe-rate in the interferograms, especially towards the center of our study area. Do you have any thoughts why this might be?
# + [markdown] heading_collapsed=true hidden=true
# ### 3.10 Step prepesd
# + [markdown] hidden=true
# Now that the bottom and top interferograms have been calculated the double difference interferogram can be calculated, this procedure is also referred to as spectral diversity:
# $$\Delta\Phi_{\text{ESD}} = \Delta\Phi_{\text{bot}}*conj\left(\Delta\Phi_{\text{top}}\right)$$
# + hidden=true
# !topsApp.py --dostep=prepesd
# + [markdown] hidden=true
# As output this generates a "*ESD*" folder, with in it the inteferograms (*overlap_[IW?]_XX.int*) and multi-looked interferograms (*overlap_[IW?]_XX.5alks_15rlks.int*) for each burst overlap *XX*.
# + [markdown] heading_collapsed=true hidden=true
# ### 3.11 Step esd
# + [markdown] hidden=true
# The double difference interferogram only captures displacements in the azimuth direction. During the **esd** step, the phase of the double difference interferogram for each burst overlap is converted to pixel offsets using the azimuth frequency and difference of the Doppler Centroid frequency between the forward and backward-looking geometry.
#
# $$ \text{offset}_{\text{azimuth}}= \frac{f_{\text{azimuth}}}{2\pi} \frac{\text{arg}\left[e^{ j\Delta\Phi_{\text{ESD}}} \right]}{\Delta f_{\text{DC}}^{\text{overlap}}}$$
#
# Once completed, all the offsets are combined together, and the mean of the distribution is taken as the azimuth coregistration offset. Note only those pixels with a coherence > than 0.85 are considered.
#
# <div class="alert alert-danger">
# <b>POTENTIAL ISSUE:</b>
# Coherence is too bad to find reliable point to estimate the ESD threshold. Either take one the following two options listed below.
# </div>
# <br>
# <div class="alert alert-warning">
# <b>ESD threshold:</b>
# The ESD coherence threshold can be changed by the user in the **topsApp.xml** using the property **ESD coherence threshold"**. It is recommended not to go below 0.7.
# </div>
# <br>
# <div class="alert alert-warning">
# <b>SKIP ESD:</b>
# In case you do not want to perform ESD, e.g. the coherence is to bad in the overlap regions, you could opt to skip ESD and only use the orbit information. This is controlled via **topsApp.xml** with the property **do ESD**.
# </div>
# + hidden=true
# !topsApp.py --dostep=esd
# + [markdown] hidden=true
# At the completion of the **esd** step you will see that *combined_*\**[.cor,.int,freq.bin]* files as well as an *ESDmisregistration.jpg* figure have been added to the "*ESD*" folder.
# + hidden=true
# ls ESD/combined* ESD/ESDmisregistration*
# + [markdown] hidden=true
# The combined coherence and interferogram files are used as input to the ESD azimuth offset estimation:
# + hidden=true
plotdata('ESD/combined_IW3.cor',1,title="coherence [-]",aspect=10, datamin=0, datamax=1,background='yes')
plotcomplexdata('ESD/combined_IW3.int',title="Diff IFG [-]",aspect=10,datamin=100000, datamax=100000000,draw_colorbar=True)
# + [markdown] hidden=true
# In general, high coherence can be observed (> 0.85), and thus most pixels will be used in the ESD histogram generation. The interferogram shows strong azimuthal motion at the location of the earthquake. Away from the earthquake the impact is less, showing on average ~0 rad of azimuthal phase. Let us investigate if this potentially has biased the ESD estimation by inspecting the historgram figure that was created as part of the **esd** step:
# + hidden=true
import matplotlib.image as mpimg
img= mpimg.imread("ESD/ESDmisregistration.png")
fig = plt.figure(figsize=(10, 11))
plt.imshow(img);
# + [markdown] hidden=true
# <div class="alert alert-warning">
# <b>TIP:</b>
# It is always good to inspect the inputs of the ESD calculation, as there are sources such as ionosphere and tectonic deformation that can cause an apparent azimuthal shift, potentially biasing the mean of the ESD estimation.
# </div>
# + [markdown] heading_collapsed=true hidden=true
# ### 3.12 Step rangecoreg
# + [markdown] hidden=true
# In the next step we will estimate the offset in the range direction by doing an amplitude cross-correlation between the master and slave burst overlaps. As for the azimuth offsets, a histogram of the range offset is used to estimate the mean range offset for the complete SLC. Note only pixels with an SNR of 8 or above are considered.
# + hidden=true
# !topsApp.py --dostep=rangecoreg
# + [markdown] hidden=true
# Let us vizualize the results:
# + hidden=true
# ls ESD
# + hidden=true
import matplotlib.image as mpimg
img= mpimg.imread("ESD/rangeMisregistration.jpg")
fig = plt.figure(figsize=(10, 11))
plt.imshow(img);
# + [markdown] heading_collapsed=true hidden=true
# ### 3.13 Step fineoffsets
# + [markdown] hidden=true
# Now that we have estimated the range and azimuth offsets using the burst overlap regions, we will apply them to the full bursts in the **fineoffsets** step.
# + hidden=true
# !topsApp.py --dostep=fineoffsets
# + [markdown] hidden=true
# The output is stored in the *fine_offsets* folder for each subswath, with an azimuth offset file (*azimuth_XX.off*) and a range offset file (*range_XX.off*) for each burst *XX*.
# + hidden=true
# ls fine_offsets/*
# + [markdown] heading_collapsed=true hidden=true
# ### 3.14 Step fineresamp
# + [markdown] hidden=true
# The burst fine offset files just created are used to resample the slave burst SLC's into the master geometry using the **fineresamp** step.
# + hidden=true
# !topsApp.py --dostep=fineresamp
# + [markdown] hidden=true
# The output is stored in the *coreg_fine* folder for each subswath, containing the resampled slave burst SLC's (*burst_XX.slc*).
# + [markdown] heading_collapsed=true hidden=true
# ### 3.15 Step burstifg
# + [markdown] hidden=true
# The slave and master burst SLC's are now in a common geometry, and interferograms can be computed for each burst. We will do this as part of the **burstifg** step.
# + hidden=true
# !topsApp.py --dostep=burstifg
# + [markdown] hidden=true
# This step generates a "*fine_interferogram*" folder, within which is a subswath breakdown (*IW*), each containing their respective interferograms (*burst_XX.int*), the coherence (*burst_XX.cor*), and the multilooked interferograms (*burst_XX.7alks_19rlks.int*) for each burst *XX*.
# + hidden=true
# ls fine_interferogram/*
# + [markdown] hidden=true
# Here is a plot of one of the burst interferograms:
# + hidden=true
plotcomplexdata('fine_interferogram/IW3/burst_04.7alks_19rlks.int.vrt',title="Burst 4 IFG ",aspect=3,datamin=0, datamax=10000,draw_colorbar=True)
# + [markdown] heading_collapsed=true hidden=true
# ### 3.16 Step mergebursts
# + [markdown] hidden=true
# As you can tell from the figure, it is hard to capture the full extent of the earthquake deformation from a single burst. In the **mergebursts** step, we will combine the different bursts together to generate a combined product.
# + hidden=true
# !topsApp.py --dostep=mergebursts
# + [markdown] hidden=true
# The output is written into the "*merged*" folder. This folder contains the merged interferogram (**topophase.flat**), the coherence (**topophase.cor**), the line-of-sight (LOS) angles (**los.rdr**), the heights (**z.rdr**), the longitude (**lon.rdr**), and the latitude information (**lat.rdr**). These files are multilooked, while files with a *.full* suffix refer to the full resolution data. You can also find a merged SLC for the master (**master.slc.full.vrt**) and the slave (**master.slc.full.vrt**). These files are generated if **do denseoffsets** property is set to *True* in **topsApp.xml**. You can use *gdalinfo* to explore the format of these files and the number of bands.
#
#
# Here is a plot of the merged intergerogram:
# + hidden=true
plotcomplexdata('merged/topophase.flat.vrt',title="MERGED IFG ",aspect=3,datamin=0, datamax=10000,draw_colorbar=True)
# + [markdown] hidden=true
# At first sight the stitching appears to be successful, with no obvious discontinuities in the earthquake region across bursts. A closer inspection shows that the two top burst edges have a slight misalignment. This is likely introduced by either the ionosphere or leakage of the earthquake azimuth deformation into the ESD estimation procedure. Two potential avenues can be further explored:
# #1) In addition to the ESD threshold, mask out pixels within the earthquake region. i.e. let the ESD estimation be described by the far-field.
# #2) Specify not to use ESD in the topsApp.xml. Instead only use the orbit information for coregistration purposes.
# + [markdown] heading_collapsed=true hidden=true
# ### 3.17 Step filter
# + [markdown] hidden=true
# Multilooking reduces noise, but it is often insufficient for interpreting the interferogram or unwrapping. We often apply additional filtering to improve the SNR.
# + hidden=true
# !topsApp.py --dostep=filter
# + [markdown] hidden=true
# The outputs of the filtering step are added to the merged folder, where the filtered interferogram is called **filt_topophase.flat**, and the filtered coherence **phsig.cor**. Note that the latter is different from conventional coherence: it is based on an estimate of the standard deviation of the phase translated back to coherence through the Cramer-Rao bound relationship:
#
# $\gamma = \frac{1}{\sqrt{1+2N\sigma_\phi^2}}$
#
# Let us compare the filtered and unfiltered interferograms:
# + hidden=true
plotcomplexdata('merged/topophase.flat.vrt',title="MERGED IFG ",aspect=3,datamin=0, datamax=10000,draw_colorbar=True)
plotcomplexdata('merged/filt_topophase.flat.vrt',title="MERGED FILT IFG ",aspect=1,datamin=0, datamax=10000,draw_colorbar=True)
# + [markdown] hidden=true
# <div class="alert alert-warning">
# <b>FILTER STRENGTH:</b>
# The amount of filtering can be controlled through the **"filter strength"** property in the **topsApp.xml file**.</div>
# <div class="alert alert-danger">
# <b>POTENTIAL ISSUE:</b>
# As a user you can change the filtering strength. Care should be taken, as a smoother and cleaner interferogram does not necessarily imply a better interferogram. Always inspect the interferogram before and after filtering and assess if false signals are introduced or if dense fringes are oversmoothed due to overfiltering. Look carefully at noisy regions, where fringes from coherent areas "bleed" into noisy areas.
# </div>
#
# + [markdown] heading_collapsed=true hidden=true
# ### 3.18 Step unwrap
# + [markdown] hidden=true
# In the next step we will use **snaphu** to unwrap the interferogram:
# + hidden=true
# !topsApp.py --dostep=unwrap
# + [markdown] hidden=true
# The unwrapped output is added to the "*merged*" folder. The unwrapped interferogram is contained in **filt_topophase.unw**. You will also find the connected components stored in **filt_topophase.conncomp**. We plot them below and inspect the results for potential errors.
#
# <div class="alert alert-warning">
# <b>INSPECT CONNECTED COMPONENT OUPUT:</b>
# The connected component file is a metric of the unwrapping performance. It provides information on which parts of the unwrapped image are connected. Regions with a different connected component can have a phase jump equivalent to a multiple of 2 $\pi$. It is therefore important to always inspect both the phase and the connected component output together. </div>
#
# <div class="alert alert-warning">
# <b>CONNECTED COMPONENT NUMBERS:</b>
# Connected component 0 refers to those pixels which cannot be reliably unwrapped. These should be masked out before doing any sort of modeling. The other connected components increment by 1 per isolated region.
# </div>
# <div class="alert alert-danger">
# <b>UNWRAPPING ERROR EXAMPLE:</b>
# To see unwrapped result with non-physical phase jumps, run the "icu" unwrapper by changing the **unwrapper name** property in **topsApp.xml** to *icu*. Note that the filename for the icu connected component is **filt_topophase.conncomp**.
# </div>
#
# + hidden=true
plotdata('merged/filt_topophase.unw',2,title="UNW FILT IFG [rad] ",colormap='jet',colorbar_orientation="vertical")
plotdata('merged/filt_topophase.unw.conncomp',1,title="UNW CONN COMP [-] ",colormap='jet',colorbar_orientation="vertical")
# + [markdown] heading_collapsed=true hidden=true
# ### 3.19 Step unwrap2stage
# + [markdown] hidden=true
# There is also a two-stage unwrapper where the outputs of the connected component file are used to adjust the unwrapped interferogram. During this optimization procedure, the phase offset between connected component boundaries are minimized. This step can only be run if the unwrapping was done with the *snaphu_mcf* unwrapper, and also requires the **"do unwrap2stage"** property to be set to *True*. Both are controlled in the topsApp.xml.
#
# We did not enable the two stage unwrapper. The impact should be small given only small scale connected components. Executing the **unwrap2stage** will therefore complete instantly.
# + hidden=true
# !topsApp.py --dostep=unwrap2stage
# + [markdown] heading_collapsed=true hidden=true
# ### 3.20 Step geocode
# + [markdown] hidden=true
# In the next step we will **geocode** many of the output products created to this point. By default topsApp.py geocodes the following files: **filt_topophase.flat**, **filt_topophase.unw**, **los.rdr**, **phsig.cor**, **topophase.cor** and **topophase.flat**. The user can control which files are geocoded using the **"geocode list"** property in **topsApp.xml**.
# + hidden=true
# !topsApp.py --dostep=geocode
# + hidden=true
plotdata('merged/filt_topophase.unw.geo',2,title="UNW GEO FILT IFG [rad] ",colormap='jet',colorbar_orientation="vertical",datamin=-50, datamax=190)
#plotdata('merged/filt_topophase_2stage.unw.geo',2,title="UNW 2-stage GEO FILT IFG [rad] ",colormap='jet',colorbar_orientation="vertical",datamin=-40, datamax=200)
# + [markdown] heading_collapsed=true hidden=true
# #### Changing your geocode DEM:
# + [markdown] hidden=true
# To experiment with a coarser resolution DEM, proceed with this section. By default, the geocoding in topsApp.py is done at the same sampling as processing DEM. However, a different DEM can be specified using the **geocode demfilename** property.
# + [markdown] heading_collapsed=true hidden=true
# ##### Option 1: Downsample your processing DEM
# + [markdown] hidden=true
# The ISCE contrib folder has a script called **downsampleDEM.py** that allows you to downsample a DEM to a user-defined sampling. To see how to call the function type:
# + hidden=true
# !downsampleDEM.py -h
# + [markdown] hidden=true
# For example let us downsample the processing DEM that you have been using to 120m resolution. First find out what the name and the path of the DEM is. Either you are using the DEM as specified in the topsApp.xml file or a DEM has been downloaded to your local directory as part of the processing.
# + hidden=true
if not os.path.isfile('demLat_N33_N36_Lon_E045_E047.dem.wgs84'):
SRC_processing_DEM_dir = processing_DEM_dir
processing_DEM ="demLat_N33_N36_Lon_E045_E047.dem.wgs84"
else:
SRC_processing_DEM_dir = '.'
processing_DEM = "demLat_N33_N36_Lon_E045_E047.dem.wgs84"
# making the paths absolute for dir changes
SRC_processing_DEM_dir = os.path.abspath(SRC_processing_DEM_dir)
# providing results to user
print("Your processing DEM is called " + processing_DEM )
print("Stored at: " + SRC_processing_DEM_dir)
# + [markdown] hidden=true
# Let us now locally downsample this dem to 120m using **downsampleDEM.py**
# + hidden=true
cmd = "downsampleDEM.py -i " + processing_DEM + ".vrt -r 120"
print(cmd)
os.chdir(SRC_processing_DEM_dir)
os.system(cmd)
expected_DEM = os.path.join(SRC_processing_DEM_dir,"Coarse_" + processing_DEM)
if os.path.isfile(expected_DEM):
print("Downsampled DEM:")
print(expected_DEM)
os.chdir(processing_dir)
# + [markdown] hidden=true
# Now update your **geocode demfilename** in the topsApp.xml file with the new coarse DEM. Once done, you can run the geocode step again:
# + hidden=true
# !topsApp.py --dostep=geocode
plotdata('merged/filt_topophase.unw.geo',2,title="UNW GEO FILT IFG [rad] ",colormap='jet',colorbar_orientation="vertical")
# + [markdown] heading_collapsed=true hidden=true
# ##### Option 2: Use of a different DEM
# + [markdown] hidden=true
# A 3-arc second DEM has been provided with this tutorial. You can use this to geocode at ~90 m resolution by rerunning **geocode** after completing the following two steps:
#
# 1) The processing information has already been loaded in the **startup** step through the **topsProc.xml** file, so the geocode DEM is already stored. We will therefore need to manually update the geocode DEM in it. It is a good practice to set it directly in the topsApp.xml file with the property for the *geocode demFilename*.
#
# ``` vim
# vim topsProc.xml
# ```
#
# Remember the xml guidelines:
# ``` xml
# <Dem_Used>/Users/dbekaert/UNAVCO/TopsApp/DEM/DEM1/demLat_N33_N36_Lon_E045_E047.dem.wgs84</Dem_Used>
# to
# <Dem_Used>../../DEM/DEM3/Coarse_demLat_N33_N36_Lon_E045_E047.dem.wgs84</Dem_Used>
# ```
#
# 2) Update the DEM metadata information. The DEM metadata such as the DEM path is read from the xml information. As the DEM was copied from another machine we need to update the path. fixImageXml.py allows you to do this from the command line.
#
#
# + hidden=true
# !fixImageXml.py -h
# + [markdown] hidden=true
# From the help, you can see that the **-f** option will update the path in the xml file with an absolute path. Now run the command:
# + hidden=true
os.chdir(geocoding_DEM_dir)
# !fixImageXml.py -i Coarse_demLat_N33_N36_Lon_E045_E047.dem.wgs84 -f
os.chdir(processing_dir)
# + [markdown] hidden=true
# Now rerun the **geocode** step again. Do you see the decrease in file size when plotting the result?
# + hidden=true
# !topsApp.py --dostep=geocode
plotdata('merged/filt_topophase.unw.geo',2,title="UNW GEO FILT IFG [rad] ",colormap='jet',colorbar_orientation="vertical")
# + [markdown] heading_collapsed=true hidden=true
# ### 3.21 Step denseoffsets
# + [markdown] hidden=true
# In addition to the interferometric processing, we also requested to run the **denseoffsets** step in the **topsApp.xml**. The azimuth and range offsets are estimated by doing a cross-correlation between the coregistered master and slave SLC's on a dense grid, providing a low resolution image of displacement in the range and azimuth domain.
# + hidden=true
# !topsApp.py --dostep=denseoffsets
# + [markdown] hidden=true
# The output file, **dense_offsets.bil**, is a two-band file containing the azimuth and range offsets, placed in the "*merged*" folder . The correlation coefficient is contained in the **dense_offsets_snr.bil** file.
#
# Here is a plot of the azimuth and range offsets:
# + hidden=true
plotdata('merged/dense_offsets.bil',2,title="DENSE RANGE OFFSETS [pixels] ",colormap='jet',datamin=-0.5, datamax=0.5,colorbar_orientation="vertical")
plotdata('merged/dense_offsets.bil',1,title="DENSE AZIMUTH OFFSETS [pixels] ",colormap='jet',datamin=-0.25, datamax=0.25,colorbar_orientation="vertical")
# + [markdown] heading_collapsed=true hidden=true
# ### 3.22 Step filteroffsets
# + [markdown] hidden=true
# The denseoffsets can be a bit noisy. In the following step, we will clean this up by applying a filter. The output will be added to the "*merged*" folder under the filename **filt_dense_offsets.bil**.
# + hidden=true
# !topsApp.py --dostep=filteroffsets
# + [markdown] heading_collapsed=true hidden=true
# ### 3.23 Step geocodeoffsets
# + [markdown] hidden=true
# Geocoding the filtered dense offsets generates a **filt_dense_offsets.bil.geo** file in the "*merged*" folder.
# + hidden=true
# !topsApp.py --dostep=geocodeoffsets
# + [markdown] hidden=true
# Here is a plot of the final geocoded filtered dense offsets:
# + hidden=true
plotdata('merged/filt_dense_offsets.bil.geo',2,title="DENSE AZIMUTH OFFSETS [pixels] ",colormap='jet',datamin=-0.5, datamax=0.5,colorbar_orientation="vertical")
plotdata('merged/filt_dense_offsets.bil.geo',1,title="DENSE RANGE OFFSETS [pixels] ",colormap='jet',datamin=-0.25, datamax=0.25,colorbar_orientation="vertical")
# -
# # Relevant references:
# - <NAME>, <NAME>, and <NAME>, *Precise coregistration of Sentinel-1A TOPS data*, https://files.scec.org/s3fs-public/0129_1400_1530_Fattahi.pdf
# - ESA, *Sentinel-1 and TOPS overview*, https://sentinel.esa.int/web/sentinel/user-guides/sentinel-1-sar
# - <NAME>, <NAME>, <NAME>,<NAME>, <NAME>, <NAME>, <NAME>, and <NAME>, *Interferometric Processing of Sentinel-1 TOPS Data*, IEEE, doi:10.1109/TGRS.2015.2497902, https://ieeexplore.ieee.org/document/7390052/
#
| docs/Notebooks/TOPS/Tops.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import cv2
import numpy as np
fold_path = './data/msnn4_fold4'
model = 'mcnn4'
out_img_ori = "ucf-cc-50-gt.png"
out_img_model = "ucf-cc-50-et-msnn4.png"
image = "./data/utils/ucf-cc-50/26.jpg"
den_map = "./data/utils/ucf-cc-50/26.npy"
alpha = 0.35
den_scale_factor = 1e3
# +
from architecture.crowd_count import CrowdCounter
import architecture.network as network
import os.path as osp
# gt
img = cv2.imread(image, cv2.IMREAD_COLOR)
den = np.load(den_map)
den = (den/np.max(den) * 255).astype("uint8")
heatmap = cv2.applyColorMap(den, cv2.COLORMAP_JET)
super_img = (1-alpha) * img + alpha * heatmap
cv2.imwrite(out_img_ori, super_img)
#model
imgt = cv2.imread(image, 0)
net = CrowdCounter(model = model).cuda().eval()
pretrained_model = osp.join(fold_path, 'best_model.h5')
network.load_net(pretrained_model, net)
imgt = imgt.astype(np.float32, copy=False)
imgt = imgt.reshape((1,imgt.shape[0],imgt.shape[1]))
imgt = imgt / 127.5 - 1
imgt = np.array([imgt])
#imgt = network.np_to_variable(imgt, is_cuda=True, is_training=False)
den = net(imgt)
den = den.data.cpu().numpy()
den /= den_scale_factor
den = den.reshape((den.shape[2], den.shape[3]))
ht = den.shape[0]
wd = den.shape[1]
wd_1 = img.shape[1]
ht_1 = img.shape[0]
den = cv2.resize(den,(wd_1,ht_1))
den = den * ((wd*ht)/(wd_1*ht_1)) #fix people count
print(img.shape, den.shape, imgt.shape)
den = (den/np.max(den) * 255).astype("uint8")
heatmap = cv2.applyColorMap(den, cv2.COLORMAP_JET)
super_img = (1-alpha) * img + alpha * heatmap
cv2.imwrite(out_img_model, super_img)
# -
| make_den_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Diagnostics for Spliner (3d, Least squares)
# ### Configure environment
# +
import os
os.chdir("/home/hbabcock/Data/storm_analysis/sa_diagnostics/spliner_ls")
print(os.getcwd())
import numpy
numpy.random.seed(1)
# -
import storm_analysis.diagnostics.spliner.settings as settings
import storm_analysis.diagnostics.spliner.configure as configure
import storm_analysis.diagnostics.spliner.make_data as makeData
import storm_analysis.diagnostics.spliner.analyze_data as analyzeData
import storm_analysis.diagnostics.spliner.collate as collate
# +
settings.photons = [[20, 500], [20, 1000]]
print(settings.photons)
settings.fit_error_model = "LS"
settings.iterations = 20
settings.n_frames = 10
settings.peak_locations = None
settings.smooth_psf = False
# -
# ### Configure
configure.configure(False)
# ### Make Data
makeData.makeData()
# ### Analyze data
# %time analyzeData.analyzeData()
# ### Collate data
collate.collate()
# ### Reference results
# + active=""
# 2019-04-01
# commit <PASSWORD>
#
# Processing test_01
# Using max_distance 200.0 nm for error calcuations.
# Processing test_02
# Using max_distance 200.0 nm for error calcuations.
#
# Analysis Summary:
# Processed 2425 localizations in 3.68 seconds, 659.46/sec
# Recall 0.53492
# Noise 0.46508
# XYZ Error Standard Deviation (nm):
# test_01 36.57 35.99 61.95
# test_02 21.25 21.27 33.85
#
# XYZ RMSE Accuracy (nm):
# test_01 36.61 36.00 61.97
# test_02 21.25 21.27 33.85
| storm_analysis/diagnostics/jpy_notebooks/spliner_ls.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:PythonData] *
# language: python
# name: conda-env-PythonData-py
# ---
# # Import dependencies
# +
# Dependencies
import pandas as pd
import numpy as np
import requests
from census import Census
# Census API Key
from config import api_key
#change year as needed
c = Census(api_key, year=2015)
# SQLAlchemy
from sqlalchemy import create_engine
from secret import username, password
import psycopg2
# -
# # Store CSV into DataFrame
accident_csv_file = "Resources/US_Accidents_June20.csv"
accident_data_df = pd.read_csv(accident_csv_file)
accident_data_df.head()
state_csv_file = "Resources/state_csvData.csv"
state_df = pd.read_csv(state_csv_file)
state_df.head()
# Rename columns to match accident df for future merging
state_df = state_df.drop(columns={"Abbrev"})
state_df = state_df.rename(columns={"State": "state_name",
"Code": "State"})
state_df.head()
# # accident_df
# +
# view data before transformation
# accident_data_df.nunique()
# +
# accident_data_df.info()
# -
# # Create new accident df
new_accident_data_df = accident_data_df[['ID', 'Start_Time', 'City', 'County', 'State', 'Zipcode', 'Street', 'Timezone', 'Temperature(F)', 'Visibility(mi)', 'Weather_Condition']].copy()
new_accident_data_df.head()
# # merge state and accident df
# Merge accident_df with state_df to obtain state_name for future merging with population df
clean_accident_data_df = pd.merge(new_accident_data_df, state_df, on= "State")
clean_accident_data_df.head()
# convert start_time from object to datetime
clean_accident_data_df['Start_Time'] = pd.to_datetime(clean_accident_data_df['Start_Time'])
clean_accident_data_df.info()
# +
# clean_accident_data_df.nunique()
# -
# Limit data from 2016-2020 to 2016 only
clean_accident_data_df = clean_accident_data_df[(clean_accident_data_df['Start_Time']>= "2016-01-01") &
(clean_accident_data_df['Start_Time']<= "2018-12-31")]
# clean_accident_data_df.nunique()
clean_accident_data_df.head()
# # Check to see if all 2016 data is present
# Sort by datetime
clean_accident_data_df = clean_accident_data_df.sort_values('Start_Time')
clean_accident_data_df.head()
# +
# clean_accident_data_df.count()
# +
# clean_accident_data_df.nunique()
# -
# # Test to see if correct data is included
# +
# clean_accident_data_df['normalised_date'] = clean_accident_data_df['Start_Time'].dt.normalize()
# clean_accident_data_df.head()
# -
# # Reorganize df to look pretty
clean_accident_data_df = clean_accident_data_df[['ID', 'Start_Time', 'City', 'County', 'State', 'state_name', 'Zipcode', 'Street', 'Timezone', 'Temperature(F)', 'Visibility(mi)', 'Weather_Condition']]
clean_accident_data_df.head()
clean_accident_data_df = clean_accident_data_df.rename(columns={"State": "state_abbrev",
"state_name":"State",
"Start_Time": "Date_Time"})
# # Population from 2015-2018
#2015
c = Census(api_key, year=2015)
county_pop_data= c.acs5.get(("NAME","B01001_001E"), {"for": "county:*"})
county_pop15_df = pd.DataFrame(county_pop_data)
county_pop15_df.rename(columns = {"B01001_001E":"2015"}, inplace=True)
county_pop15_df.head()
#2016
c = Census(api_key, year=2016)
county_pop_data= c.acs5.get(("NAME","B01001_001E"), {"for": "county:*"})
county_pop16_df = pd.DataFrame(county_pop_data)
county_pop16_df.rename(columns = {"B01001_001E":"2016"}, inplace=True)
county_pop16_df.head()
#2017
c = Census(api_key, year=2017)
county_pop_data= c.acs5.get(("NAME","B01001_001E"), {"for": "county:*"})
county_pop17_df = pd.DataFrame(county_pop_data)
county_pop17_df.rename(columns = {"B01001_001E":"2017"}, inplace=True)
county_pop17_df.head()
#2018
c = Census(api_key, year=2018)
county_pop_data= c.acs5.get(("NAME","B01001_001E"), {"for": "county:*"})
county_pop18_df = pd.DataFrame(county_pop_data)
county_pop18_df.rename(columns = {"B01001_001E":"2018"}, inplace=True)
county_pop18_df.head()
df_1516 = pd.merge(county_pop15_df, county_pop16_df, on="NAME")
df_1718 = pd.merge(county_pop17_df, county_pop18_df, on="NAME")
county_pop_df = pd.merge(df_1516, df_1718, on="NAME")
county_pop_df.head()
county_pop_df.drop(columns=["state_x_x","county_x_x","state_y_x","county_y_x","state_x_y","county_x_y"], inplace=True)
county_pop_df.rename(columns = {"NAME":"county", "state_y_y":"state_id", "county_y_y":"county_id"}, inplace=True)
county_pop_df = county_pop_df[["county","county_id","state_id","2015","2016","2017","2018"]]
county_pop_df.head()
#make another call for state population just to get a list of states since county infor only have state census id
state_pop_data= c.acs5.get(("NAME","B01001_001E"), {"for": "state:*"})
#pass on the state pop data to a df
state_pop_df = pd.DataFrame(state_pop_data)
state_pop_df.head()
#drop population column from the state pop data df
state_pop1_df = state_pop_df.drop(columns=["B01001_001E"])
state_pop1_df.head()
#rename "Name" column to prevent duplication when merging with county pop df
state_pop1_df.rename(columns = {"NAME":"state", "state":"state_id"}, inplace=True)
state_pop1_df.head()
#merge state and county df on the "state" the numeric identifier for states
county_pop1_df = pd.merge(county_pop_df, state_pop1_df, on="state_id")
county_pop1_df.head()
#reorder columns
county_pop1_df = county_pop1_df[["county","county_id","state","state_id","2015","2016","2017","2018"]]
#convert population columns to numeric
county_pop1_df[["2015","2016","2017","2018"]] = county_pop1_df[["2015","2016","2017","2018"]].apply(pd.to_numeric)
county_pop1_df.info()
#import US state 2 letter abbreviation dictionary to create a state table
us_state_abbrev = {
'Alabama': 'AL',
'Alaska': 'AK',
'American Samoa': 'AS',
'Arizona': 'AZ',
'Arkansas': 'AR',
'California': 'CA',
'Colorado': 'CO',
'Connecticut': 'CT',
'Delaware': 'DE',
'District of Columbia': 'DC',
'Florida': 'FL',
'Georgia': 'GA',
'Guam': 'GU',
'Hawaii': 'HI',
'Idaho': 'ID',
'Illinois': 'IL',
'Indiana': 'IN',
'Iowa': 'IA',
'Kansas': 'KS',
'Kentucky': 'KY',
'Louisiana': 'LA',
'Maine': 'ME',
'Maryland': 'MD',
'Massachusetts': 'MA',
'Michigan': 'MI',
'Minnesota': 'MN',
'Mississippi': 'MS',
'Missouri': 'MO',
'Montana': 'MT',
'Nebraska': 'NE',
'Nevada': 'NV',
'New Hampshire': 'NH',
'New Jersey': 'NJ',
'New Mexico': 'NM',
'New York': 'NY',
'North Carolina': 'NC',
'North Dakota': 'ND',
'Northern Mariana Islands':'MP',
'Ohio': 'OH',
'Oklahoma': 'OK',
'Oregon': 'OR',
'Pennsylvania': 'PA',
'Puerto Rico': 'PR',
'Rhode Island': 'RI',
'South Carolina': 'SC',
'South Dakota': 'SD',
'Tennessee': 'TN',
'Texas': 'TX',
'Utah': 'UT',
'Vermont': 'VT',
'Virgin Islands': 'VI',
'Virginia': 'VA',
'Washington': 'WA',
'West Virginia': 'WV',
'Wisconsin': 'WI',
'Wyoming': 'WY'
}
state_series = pd.Series(us_state_abbrev)
state_df = state_series.to_frame()
state_df.reset_index(inplace=True)
state_df.rename(columns={"index":"state",0:"statecode"}, inplace=True)
state1_df = pd.merge(state_df,state_pop1_df, on="state" )
#County Estimated Population 2015 to 2018
county_pop1_df.head()
#STATE TABLE
state1_df.head()
# # edit population
county_pop1_df.rename(columns = {"2015":"2015 Population",
"2016":"2016 Population",
"2017":"2017 Population",
"2018":"2018 Population"}, inplace=True)
county_pop1_df.head(300)
clean_pop_df = county_pop1_df.copy()
clean_pop_df.head()
# Cleaned county column
clean_pop_df['county'] = clean_pop_df['county'].str.split(",", expand=True)[0]
clean_pop_df['county'] = clean_pop_df['county'].str.replace('County', '')
clean_pop_df.head()
# clean_pop_df = clean_pop_df.drop(columns={"state"})
clean_pop_df = clean_pop_df.rename(columns={"county": "County",
"state": "State"})
clean_pop_df.head()
# ## Extract,Clean,Transform DL data
#Extract Licensed Drivers by state (DL) csv into pandas DF for year 2014,2015,2016
DL_DF = pd.read_csv("Resources/DL.csv", usecols=[0,66,67,68])
DL_DF
# clean Licensed Drivers by state (DL)
DL_DF_Clean = DL_DF.rename(columns={'2014':'num_licenced_driver_2014',
'2015':'num_licenced_driver_2015',
'2016':'num_licenced_driver_2016',
'STATE':'state_name'})
DL_DF_Clean
DL_DF_Clean.info()
# +
# DL_DF_Clean['num_licenced_driver_2014'] = pd.to_numeric(DL_DF_Clean['num_licenced_driver_2014'],errors='coerce')
# DL_DF_Clean['num_licenced_driver_2015'] = pd.to_numeric(DL_DF_Clean['num_licenced_driver_2015'],errors='coerce')
# DL_DF_Clean['num_licenced_driver_2016'] = pd.to_numeric(DL_DF_Clean['num_licenced_driver_2016'],errors='coerce')
# +
# DL_DF_Clean.info()
# +
# DL_DF_Clean.info()
# -
# ## Extract,Clean,Transform RMV data
#Extract Registered Motor Vehicles(RMV) csv into pandas DF
RMV_DF = pd.read_csv("Resources/RMV.csv", usecols=[0,15], skiprows=4)
RMV_DF
# clean egistered Motor Veicheles(RMV)
RMV_DF_Clean = RMV_DF.rename(columns={'Unnamed: 0':'state_name','Unnamed: 15':'num_reg_vehicle'})
RMV_DF_Clean
# +
# RMV_DF_Clean['num_reg_vehicle'] = pd.to_numeric(RMV_DF_Clean['num_reg_vehicle'],errors='coerce')
# RMV_DF_Clean.info()
# -
# # Export csv
RMV_DF_Clean.head()
RMV_DF_Clean.to_csv("Resources/Output/reg_vehicle.csv", index=False)
DL_DF_Clean.head()
DL_DF_Clean.to_csv("Resources/Output/license.csv", index=False)
clean_pop_df.head()
clean_pop_df_rename = clean_pop_df.copy()
clean_pop_df_rename = clean_pop_df_rename.rename(columns= {"2015 Population": "population_2015",
"2016 Population": "population_2016",
"2017 Population": "population_2017",
"2018 Population": "population_2018",
"County": "county"})
clean_pop_df_rename = clean_pop_df_rename.sort_values(by='State')
clean_pop_df_rename.drop(columns= ["State", "state_id", "county_id"], inplace=True)
clean_pop_df_rename= clean_pop_df_rename[["county", "population_2015", "population_2016","population_2017", "population_2018"]]
clean_pop_df_rename.head()
# +
# clean_pop_df_rename.nunique()
# -
clean_pop_df_rename.to_csv("Resources/Output/population.csv", index=False)
clean_accident_data_df.head()
clean_accident_data_df_rename = clean_accident_data_df.copy()
clean_accident_data_df_rename = clean_accident_data_df_rename.rename(columns= {"ID": "id",
"Date_Time": "datetime",
"City": "city",
"County": "county",
"State": "state_name",
"Zipcode": "zipcode",
"Timezone": "timezone",
"Temperature(F)": "temperature",
"Visibility(mi)": "visibility",
"Weather_Condition": "weather_condition"})
# drop street and state abbrev
clean_accident_data_df_rename.drop(columns= ["state_abbrev", "Street"], inplace=True)
clean_accident_data_df_rename.head()
clean_accident_data_df_rename = clean_accident_data_df_rename.sort_values(by='datetime')
clean_accident_data_df_rename.head()
clean_accident_data_df_rename = clean_accident_data_df_rename[['id', 'datetime', 'state_name', 'city', 'county', 'zipcode', 'timezone', 'temperature', 'visibility', 'weather_condition']]
clean_accident_data_df_rename.head()
# +
# clean_accident_data_df_rename.nunique()
# -
# ## Run Accident_Schema_Final.sql in Postgres now
clean_accident_data_df_rename.to_csv("Resources/Output/accident.csv", index=False)
rds_connection_string =f'{username}:{password}@localhost:5432/us_accidents'
engine = create_engine(f'postgresql+psycopg2://{rds_connection_string}')
engine.table_names()
RMV_DF_Clean.to_sql(name='reg_vehicle', con=engine, if_exists='append', index=False)
RMV_DF_Clean.info()
DL_DF_Clean.to_sql(name='license', con=engine, if_exists='append', index=False)
RMV_DF_Clean
DL_DF_Clean
clean_accident_data_df_rename.info()
clean_pop_df_rename.to_sql(name='population', con=engine, if_exists='append', index=False)
clean_accident_data_df_rename.to_sql(name='accident', con=engine, if_exists='append', index=False)
| ETL_Proj_Final.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="BtMkTMxui6Ut"
# # Урок 4
#
# ## Непрерывные случайные величины. Функция распределения и плотность распределения вероятностей. Равномерное и нормальное распределение. Центральная предельная теорема
#
# ### Непрерывные случайные величины
#
# Ранее мы познакомились с дискретными случайными величинами. Такие величины принимают дискретные, т.е. разделимые значения. Например, это может быть конечное или счётное множество значений.
#
# __Непрерывные случайные величины__ принимают все значения, содержащиеся в заданном промежутке. Промежуток может быть конечным или бесконечным.
#
# Например, рост или вес человека — непрерывные случайные величины: они могут принимать любое значение в некоторых пределах.
#
# Закон распределения вероятностей дискретной случайной величины мы задавали как соответствие между значениями $a_i$ случайной величины и соответствующими вероятностями $P(X = a_i)$.
#
# Для непрерывных случайных величин аналогичный подход невозможен, поскольку вероятность $P(X = a)$ для непрерывной случайной величины $X$ равна $0$ для любого $a$. Поэтому распределение вероятностей непрерывных случайных величин характеризуют с помощью __функции распределения__:
# $$F(x) = P(X < x).$$
#
# Функция распределения показывает, какова для каждого $x$ вероятность того, что случайная величина $X$ принимает значение меньше $x$. (Для дискретных распределений эта функция ступенчатая.)
#
# Эта функция монотонно возрастает на отрезке, на котором определена случайная величина. Кроме того, $F(-\infty) = 0$ и $F(\infty) = 1$.
#
# __Пример 1__
#
# Рассмотрим опять данные по хоккеистам и построим по ним _эмпирическую_ (т.е. полученную опытным путём) функцию распределения возраста хоккеистов.
# +
import pandas as pd
import numpy as np
from matplotlib import pyplot as plt
from scipy import stats
# %config InlineBackend.figure_formats = ['svg']
# -
plt.style.use('seaborn-whitegrid')
plt.rcParams['figure.facecolor'] = 'white'
# + pycharm={"is_executing": false}
df = pd.read_csv('hockey_players.csv', encoding='cp1251')
df = df.drop_duplicates(['firstName', 'lastName', 'birthYear', 'birthMon', 'birthDay'])
df.head()
# -
def edf(samples, x: float) -> float:
"""Эмпирическая функция распределения (empirical distribution
function).
"""
return samples[samples < x].shape[0] / samples.shape[0]
# С помощью `matplotlib` построим график данной функции в пределах от $10$ до $50$.
# +
ox = np.linspace(10, 50, 1000)
oy = np.array([edf(df['age'], x) for x in ox])
plt.plot(ox, oy)
# -
# Всё же функция распределения не даёт представления о распределении, аналогичного тому, что даёт закон распределения дискретных случайных величин. Хотелось бы понять, какие значения случайной величины более «вероятно» наблюдать, чем другие.
#
# Для таких целей удобно использовать __функцию плотности__:
# $$f(x) = F'(x)$$
#
# Геометрический смысл функции плотности таков: вероятность того, что случайная величина $X$ будет лежать в отрезке $(a, b)$, равна площади под графиком функции плотности $f(x)$ в пределах от $a$ до $b$.
#
# Общая площадь под графиком функции $f(x)$ равна $1$, аналогично тому, что сумма вероятностей значений дискретной случайной величины равна $1$.
#
# Однако, стоит помнить, что _значение $f(x)$ не является вероятностью $P(X = x)$_. Оно лишь отражает _плотность_ случайной величины в окрестности точки $x$.
#
# Плотность распределения можно визуализировать с помощью гистограммы, передав в неё значение параметра `density=True`.
#
# __Пример 2__
#
# Построим гистограмму значений возраста хоккеистов, чтобы примерно представить, как выглядит соответствующая функция плотности распределения.
plt.hist(df['age'], density=True, bins=15);
# Математическое ожидание и дисперсия для непрерывной случайной величины также считаются иначе, чем для дискретной.
#
# Формула для математического ожидания:
#
# $$M(X) = \displaystyle\int_{-\infty}^\infty x \cdot f(x) \: dx$$
#
# Формула для дисперсии:
#
# $$D(X) = \displaystyle\int_{-\infty}^\infty (x - M(X))^2 \cdot f(x) \: dx$$
# ### Примеры непрерывных распределений
#
# Непрерывная случайная величина $X$ имеет __равномерное распределение__ на отрезке $[a, b]$, если её плотность внутри этого отрезка постоянна, а вне этого отрезка равна 0. Другими словами:
# $$
# f(x) = \begin{cases}
# \dfrac{1}{b - a}, & x \in [a, b], \\
# 0, & x \not\in [a, b].
# \end{cases}
# $$
#
# Не путать с _дискретным равномерным_ распределением.
#
# Математическое ожидание и дисперсия равномерного распределения:
# $$M(X) = \dfrac{a + b}{2}, \:\: D(X) = \dfrac{(b - a)^2}{12}.$$
#
# __Пример 3__
#
# Выборку из равномерного распределения можно получить с помощью функции `uniform` из модуля `numpy.random`. Построим выборку из равномерного распределения с параметрами $a = -1.5$, $b = 2.5$.
# +
a = -1.5
b = 2.5
samples = np.random.uniform(a, b, size=10000)
print(samples)
# -
# Убедимся, что каждый элемент в этой выборке встречается лишь однажды:
len(samples) - len(set(samples))
# Математическое ожидание:
# $$M(X) = \dfrac{a + b}{2} = \dfrac{-1.5 + 2.5}{2} = \dfrac{1}{2}.$$
#
# Выборочное среднее:
samples.mean()
# Дисперсия:
# $$D(X) = \dfrac{(b - a)^2}{12} = \dfrac{4^2}{12} = \dfrac{4}{3} \approx 1.333.$$
#
# Выборочная дисперсия (несмещённая) от выборки:
samples.var(ddof=1)
# Формула функции равномерного распределения:
#
# $
# F(x) = \begin{cases}
# 0, & x < a, \\
# \dfrac{x - a}{b - a}, & x \in [a, b], \\
# 1, & x > b.
# \end{cases}
# $
#
# __Пример 4__
#
# Построим эмпирическую и теоретическую функции распределения.
def cdf_uniform(a: float, b: float, x: float) -> float:
"""Функция распределения (cumulative distribution function)
для равномерного распределения.
"""
if x < a:
return 0
if x > b:
return 1
else:
return (x - a) / (b - a)
# +
ox = np.linspace(-4, 4, 1000)
oy1 = np.array([edf(samples=samples, x=x) for x in ox])
oy2 = np.array([cdf_uniform(a=a, b=b, x=x) for x in ox])
plt.plot(ox, oy1, label='edf')
plt.plot(ox, oy2, linestyle='dashed', label='cdf')
plt.legend()
# -
# __Пример 5__
#
# Построим гистограмму, чтобы визуализировать функцию плотности. Вместе с ней построим теоретическую функцию плотности.
def pdf_uniform(a: float, b: float, x: float) -> float:
"""Функция плотности (probability density function)
для равномерного распределения.
"""
return 1 / (b - a) if a <= x <= b else 0
# +
oy = np.array([pdf_uniform(a, b, x) for x in ox])
plt.hist(samples, density=True)
plt.plot(ox, oy, linestyle='dashed')
# -
# Непрерывная случайная величина $X$ имеет __нормальное распределение__ с параметрами $a$ и $\sigma > 0$, если её плотность распределения задаётся формулой
# $$f(x) = \dfrac{1}{\sqrt{2 \pi \sigma^2}} \cdot \exp \left(- \frac{(x - a)^2}{2 \sigma^2} \right)$$
#
# Параметры $a$ и $\sigma$ задают, соответственно, математическое ожидание и среднее квадратическое отклонение случайной величины:
# $$M(X) = a, \:\: D(X) = \sigma^2$$
#
# Нормальное распределение с параметрами $a = 0$ и $\sigma = 1$ называется __стандартным нормальным распределением__.
#
# Нормальное распределение является одним из наиболее распространённых на практике. Например, нормально распределены:
# * рост, вес людей
# * показатели IQ
# * время прихода на работу
# * скорость движения молекул в жидкостях и газах
#
# Как правило, нормально распределёнными являются случайные величины, описывающие события, которые зависят от большого числа слабо связанных случайных факторов.
#
# __Пример 6__
#
# Выборку из нормального распределения можно получить с помощью функции `normal` из модуля `numpy.random`. Построим такую выборку с параметрами $a = 8$, $\sigma = 2$.
loc = 8
scale = 2
# +
samples = np.random.normal(loc, scale, size=1000)
# print(samples)
# -
# Выборочные мат. ожидание и дисперсия:
samples.mean(), samples.var(ddof=1)
# Функция нормального распределения:
# $$F(x) = \dfrac{1}{2} \left[ 1 + \operatorname{erf} \left( \dfrac{x - a}{\sigma \sqrt{2}} \right) \right],$$
# где $\operatorname{erf}$ — __функция ошибок__.
#
# Функция ошибок представляет собой интеграл
# $$\operatorname{erf}(x) = \dfrac{2}{\pi}\displaystyle\int_0^x e^{- t^2} dt,$$
# который аналитически не считается. Численная реализация этой функции доступна как функция `erf` из модуля `scipy.special`.
#
# __Пример 7__
#
# Эмпирическая и теоретическая функции распределения:
from scipy.special import erf
def cdf_normal(loc: float, scale: float, x: float) -> float:
"""Функция распределения для нормального распределения.
"""
return (1 + erf((x - loc) / (scale * np.sqrt(2)))) / 2
# +
ox = np.linspace(0, 16, 1000)
oy1 = np.array([edf(samples, x) for x in ox])
oy2 = np.array([cdf_normal(loc, scale, x) for x in ox])
plt.plot(ox, oy1, label='edf')
plt.plot(ox, oy2, label='cdf', linestyle='dashed')
plt.legend()
# -
# __Пример 8__
#
# Гистограмма выборки и теоретическая функция плотности:
def pdf_normal(loc: float, scale: float, x: float) -> float:
"""Функция плотности для нормального распределения.
"""
return np.exp(- ((x - loc) ** 2) / (2 * scale ** 2)) / (scale * np.sqrt(2 * np.pi))
# +
oy = np.array([pdf_normal(loc, scale, x) for x in ox])
plt.hist(samples, density=True, bins=15)
plt.plot(ox, oy, linestyle='dashed')
# -
# Кстати, в модуле `scipy.stats` есть готовые функции распределения и плотности для многих известных распределений.
#
# ### `scipy.stats`
#
# В модуле `scipy.stats` содержатся реализации основных функций для различных распределений:
# * `binom` — биномиальное,
# * `poisson` — Пуассоновское,
# * `norm` — нормальное,
# * `uniform` — непрерывное равномерное,
# * `t` — распределение Стьюдента (о нём позже).
#
# и др.
#
# Доступные функции:
# * `pmf` — закон распределения для дискретных величин,
# * `pdf` — функция плотности для непрерывных величин,
# * `cdf` — функция распределения,
# * `ppf` — квантильная функция (обратная к функции распределения).
#
# и др. Например, с помощью этого модуля можно получить значения функции нормального распределения, которая не вычисляется аналитически и доступна лишь численно.
# +
ox = np.linspace(0, 16, 1000)
oy = np.array([stats.norm.cdf(x, loc=8, scale=2) for x in ox])
plt.plot(ox, oy)
# -
# При многократном использовании функций одного и того же распределения удобнее сразу зафиксировать объект:
# +
norm = stats.norm(loc=8, scale=2)
norm.pdf(6), norm.cdf(6), norm.ppf(0.16)
# -
# Для вычисления разброса значений нормально распределённой случайной величины можно использовать следующие правила:
#
# * Интервал от $a - \sigma$ до $a + \sigma$ (__стандартное отклонение__) содержит около $68\%$ вероятностной массы (т.е. с вероятностью $68\%$ данная величина попадает в этот интервал).
# * От $a - 2\sigma$ до $a + 2\sigma$ — около $95\%$ массы (__правило двух сигм__).
# * От $a - 3\sigma$ до $a + 3\sigma$ — около $99.7\%$ массы (__правило трёх сигм__).
#
# Посчитаем точные значения для этих интервалов. Поскольку эти правила не зависят от конкретных параметров, посчитаем интервалы для стандартного нормального распределения. (Оно используется по умолчанию в `scipy.stats.norm`.)
#
# Стандартное отклонение:
stats.norm.cdf(1) - stats.norm.cdf(-1)
# Правило двух сигм:
stats.norm.cdf(2) - stats.norm.cdf(-2)
# Правило трёх сигм:
stats.norm.cdf(3) - stats.norm.cdf(-3)
# +
loc = 800
scale = 500
stats.norm.cdf(loc + scale ** 2, loc=loc, scale=scale) - stats.norm.cdf(loc - scale ** 2, loc=loc, scale=scale)
# -
# __Пример 9__
#
# Вернёмся к данным о хоккеистах и рассмотрим столбец с ростом. Построим его гистограмму.
plt.hist(df['height'], bins=20);
# Мы знаем, что рост — одна из величин, которая может распределяться нормально. Восстановим по выборке значения $a$ и $\sigma$ и построим поверх гистограммы теоретическую функцию плотности соответствующего нормального распределения.
# +
loc = df['height'].mean()
scale = df['height'].std(ddof=1)
loc, scale
# +
ox = np.linspace(165, 205, 1000)
oy = np.array([pdf_normal(loc, scale, x) for x in ox])
plt.hist(df['height'], density=True, bins=20)
plt.plot(ox, oy, linestyle='dashed')
# -
# Проверим на этих данных правило двух сигм. В отрезке `(loc - 2 * scale, loc + 2 * scale)` должно содержаться $95\%$ значений.
# +
two_sigmas_condition = (df['height'] > loc - 2 * scale) & (df['height'] < loc + 2 * scale)
df[two_sigmas_condition].shape[0] / df.shape[0]
# -
# __Другие непрерывные распределения__
#
# * __Экспоненциальное__ (или __показательное__): время между последовательными свершениями одного и того же события. Является непрерывным аналогом геометрического распределения. Функция плотности:
#
# $$
# F(x) = \begin{cases}
# 1 - e^{- \lambda x}, & x \geq 0, \\
# 0, & x < 0.
# \end{cases}
# $$
x = np.random.normal(1, 2, size=10000)
y = np.random.normal(1, 3, size=10000)
plt.hist(x, bins=20);
plt.hist(y, bins=20);
plt.hist(x + y, bins=20);
# ### Центральная предельная теорема
#
# Одно из практически уникальных свойств нормального распределения — __устойчивость__ — означает, что если $X$ и $Y$ — _независимые нормально распределённые_ случайные величины, то их комбинация $Z = a \cdot X + b \cdot Y$ ($a$, $b$ — числа) также имеет нормальное распределение. Более того, для распределения $Z$ верны следующие равенства. Математическое ожидание:
# $$M(Z) = a \cdot M(X) + b \cdot M(Y)$$
# Дисперсия:
# $$D(Z) = |a| \cdot D(X) + |b| \cdot D(Y)$$
#
# Большинство других распределений не являются устойчивыми. Например, сумма двух равномерно распределённых случайных величин не является равномерно распределённой. Вместо этого неустойчивые распределения «стремятся» к нормальному. Это хорошо иллюстрирует центральная предельная теорема.
#
# Рассмотрим выборку из $n$ значений случайной величины $X$, имеющей произвольное распределение, и пусть $Y$ — случайная величина, равная сумме этих значений.
#
# __Центральная предельная теорема__ утверждает: чем больше $n$, тем _ближе_ распределение величины $Y$ к нормальному распределению с параметрами
# $$a = n \cdot M(X), \:\: \sigma^2 = n \cdot D(X)$$
#
#
# _Другая версия_ этой теоремы: пусть $Z$ — случайная величина, равная среднему арифметическому значений из выборки. Тогда с увеличением $n$ распределение этой величины становится всё ближе к нормальному распределению с параметрами
# $$a = M(X), \:\: \sigma^2 = \dfrac{D(X)}{n}$$
#
# Центральная предельная теорема согласуется со сделанным ранее наблюдением, что, как правило, случайные величины, описывающие события, которые зависят от большого числа слабо связанных случайных факторов, являются нормально распределёнными.
#
# Отметим, что центральная предельная теорема работает не только для непрерывных случайных величин, но и для дискретных.
#
# __Пример 8__
#
# Проверим утверждение центральной предельной теоремы на равномерном распределении $X$ с параметрами $a = -1.5$,
# $b = 2.5$. Для этого при различных $n$ сформируем $n$ выборок размера, например, $10^4$, а затем посчитаем от них поэлементно среднее. Таким образом, мы получим одну выборку размера $10^4$. Теорема утверждает, что чем больше $n$, тем больше эта выборка похожа на выборку из некоторого нормального распределения $Y$.
#
# Математическое ожидание величины $X$ равно $\dfrac{a + b}{2} = \dfrac{1}{2}$. Дисперсия:
# $\dfrac{(b - a)^2}{12} = \dfrac{4^2}{12} = \dfrac{4}{3}$. Это означает, что нормальное распределение $Y$ имеет параметры $a = \dfrac{1}{2}$, $\sigma^2 = \dfrac{4}{3n}$.
# +
a = -1.5
b = 2.5
m = (a + b) / 2
d = ((b - a) ** 2) / 12
n_samples = 10 ** 4
# -
def get_mean_samples_uniform(n: int) -> np.ndarray:
"""Возвращает выборку из значений среднего от `n` равномерно распределённых
случайных величин.
"""
samples = np.random.uniform(a, b, size=(n, n_samples))
return samples.mean(axis=0)
# +
n = 500
loc = m
scale = np.sqrt(d / n)
samples = get_mean_samples_uniform(n)
ox = np.linspace(loc - 4 * scale, loc + 4 * scale, 1000)
oy = np.array([pdf_normal(loc, scale, x) for x in ox])
plt.hist(samples, density=True, bins=20)
plt.plot(ox, oy, linestyle='dashed')
# -
# __Пример 9__
#
# Аналогично продемонстрируем центральную предельную теорему на примере дискретного распределения, а именно биномиального с параметрами $n = 30$, $p = 0.2$. Параметр $n$ обозначим переменной `n_`, чтобы не конфликтовать с переменной `n`, отвечающей за число случайных величин, от которых берётся среднее.
#
# Математическое ожидание в данном случае равно $np = 6$, дисперсия: $np(1 - p) = 4.8$.
30 * 0.2 * 0.8
# +
n_ = 30
p = 0.2
m = n_ * p
d = n_ * p * (1 - p)
n_samples = 10 ** 4
# -
def get_mean_samples_binomial(n: int) -> np.ndarray:
"""Возвращает выборку из значений среднего от `n` биномиально распределённых
случайных величин.
"""
samples = np.random.binomial(n_, p, size=(n, n_samples))
return samples.mean(axis=0)
# +
n = 1000
loc = m
scale = np.sqrt(d / n)
samples = get_mean_samples_binomial(n)
ox = np.linspace(loc - 4 * scale, loc + 4 * scale, 1000)
oy = np.array([pdf_normal(loc, scale, x) for x in ox])
plt.hist(samples, density=True, bins=20)
plt.plot(ox, oy, linestyle='dashed')
# -
# ### Упражнения
# __Задача 1__
#
# О непрерывной равномерно распределённой случайной величине `B` известно, что её дисперсия равна 0.2. Можно ли найти правую границу величины `B` и её среднее значение зная, что левая граница равна 0.5? Если да, найдите их.
#
# __Задача 2__
#
# Коробки с шоколадом упаковываются автоматически. Их средняя масса равна 1.06 кг. Известно, что 5% коробок имеют массу, меньшую 1 кг. Найдите: а) среднее квадратическое отклонение массы коробки, б) процент коробок, имеющих массу больше 1.1 кг.
#
# _Подсказка_. Найдите такое значение `scale`, для которого значение `cdf(x=1, loc=1.06, scale=scale)` близко к 0.05. Точности 0.0001 будет достаточно.
#
# __Задача 3__
#
# Коробка содержит 30 конфет. Известно, что масса каждой конфеты распределена равномерно в промежутке от 12 до 14 граммов. Используя центральную предельную теорему, найти вероятность, что масса всей коробки будет: а) меньше 390 граммов, б) больше 395 граммов, в) от 380 до 400 граммов.
#
# Массой самой коробки можно пренебречь.
#
# __Задача 4__ (на подумать)
#
# Каким образом связаны между собой понятие квантиля и правила двух и трёх сигм?
#
# Данное задание - на подумать, оформлять его не обязательно. Этот вопрос мы обсудим в начале следующего занятия.
| practice4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Template Matching
#
# Template Matching is a method for searching and finding the location of a template image in a larger image. OpenCV comes with a function cv2.matchTemplate() for this purpose. It simply slides the template image over the input image (as in 2D convolution) and compares the template and patch of input image under the template image. Several comparison methods are implemented in OpenCV. (You can check docs for more details). It returns a grayscale image, where each pixel denotes how much does the neighbourhood of that pixel match with template.
#
# If input image is of size (WxH) and template image is of size (wxh), output image will have a size of (W-w+1, H-h+1). Once you got the result, you can use cv2.minMaxLoc() function to find where is the maximum/minimum value. Take it as the top-left corner of rectangle and take (w,h) as width and height of the rectangle. That rectangle is your region of template.
#
# For Template Matching with Multiple Objects: Suppose you are searching for an object which has multiple occurances, cv2.minMaxLoc() won’t give you all the locations. In that case, we will use thresholding.
# # Hough Line Transform - cv2.HoughLines()
# The Hough transform is a technique which can be used to isolate features of a particular shape within an image. Because it requires that the desired features be specified in some parametric form, the classical Hough transform is most commonly used for the detection of regular curves such as lines, circles, ellipses, etc. A generalized Hough transform can be employed in applications where a simple analytic description of a feature(s) is not possible. Due to the computational complexity of the generalized Hough algorithm, we restrict the main focus of this discussion to the classical Hough transform. Despite its domain restrictions, the classical Hough transform (hereafter referred to without the classical prefix) retains many applications, as most manufactured parts (and many anatomical parts investigated in medical imagery) contain feature boundaries which can be described by regular curves. The main advantage of the Hough transform technique is that it is tolerant of gaps in feature boundary descriptions and is relatively unaffected by image noise.
#
# # Hough Circle Transform - cv2.HoughCircles()
#
# +
import cv2
import numpy as np
img = cv2.imread('opencv_logo.png',0)
img = cv2.medianBlur(img,5)
cimg = cv2.cvtColor(img,cv2.COLOR_GRAY2BGR)
circles = cv2.HoughCircles(img,cv2.HOUGH_GRADIENT,1,20,
param1=50,param2=30,minRadius=0,maxRadius=0)
circles = np.uint16(np.around(circles))
for i in circles[0,:]:
# draw the outer circle
cv2.circle(cimg,(i[0],i[1]),i[2],(0,255,0),2)
# draw the center of the circle
cv2.circle(cimg,(i[0],i[1]),2,(0,0,255),3)
cv2.imshow('detected circles',cimg)
cv2.waitKey(3000)
cv2.destroyAllWindows()
# -
# # Image Segmentation with Watershed Algorithm - cv2.watershed()
#
# Any grayscale image can be viewed as a topographic surface where high intensity denotes peaks and hills while low intensity denotes valleys. You start filling every isolated valleys (local minima) with different colored water (labels). As the water rises, depending on the peaks (gradients) nearby, water from different valleys, obviously with different colors will start to merge. To avoid that, you build barriers in the locations where water merges. You continue the work of filling water and building barriers until all the peaks are under water. Then the barriers you created gives you the segmentation result. This is the “philosophy” behind the watershed. You can visit the CMM webpage on watershed to understand it with the help of some animations.
#
# But this approach gives you oversegmented result due to noise or any other irregularities in the image. So OpenCV implemented a marker-based watershed algorithm where you specify which are all valley points are to be merged and which are not. It is an interactive image segmentation. What we do is to give different labels for our object we know. Label the region which we are sure of being the foreground or object with one color (or intensity), label the region which we are sure of being background or non-object with another color and finally the region which we are not sure of anything, label it with 0. That is our marker. Then apply watershed algorithm. Then our marker will be updated with the labels we gave, and the boundaries of objects will have a value of -1.
#
# # Interactive Foreground Extraction using GrabCut Algorithm
#
# GrubCut was created for foreground extraction with minimal user interaction.
#
# How it works from user point of view ? Initially user draws a rectangle around the foreground region (foreground region shoule be completely inside the rectangle). Then algorithm segments it iteratively to get the best result. Done. But in some cases, the segmentation won’t be fine, like, it may have marked some foreground region as background and vice versa. In that case, user need to do fine touch-ups. Just give some strokes on the images where some faulty results are there. Strokes basically says “Hey, this region should be foreground, you marked it background, correct it in next iteration” or its opposite for background. Then in the next iteration, you get better results.
| Books and Courses/OpenCV/6 - Template Matching, Hough Transformation, Watershed Algorithm, GrabCut Algorithm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import glob
import os
import matplotlib.pyplot as plt
os_sep = os.path.abspath(os.sep)
wd = os.getcwd()
dfs = os.path.join(os_sep, wd, 'opensmile', 'egemaps_summary_turns_zero_filtered') #the feature dfs of the interviews
dem_dir = os.path.join(os_sep, 'dem_dir') #where to find the txt files with the group information of each participant
# +
pauses = 'MeanUnvoicedSegmentLength'
syll_rate = 'VoicedSegmentsPerSec'
pitch = 'F0semitoneFrom27.5Hz_sma3nz_amean'
loudness = 'loudness_sma3_amean'
pitch_var = 'F0semitoneFrom27.5Hz_sma3nz_stddevNorm'
features = [pauses, syll_rate, pitch, loudness, pitch_var]
# -
# #### Split each interview in conversation halves
#
# For each of the loaded dataframes, split the conversation in first and second half, separately for each speaker since we correlate the first and second part individually.
# +
ch1_first_half = []
ch1_second_half = []
ch2_first_half = []
ch2_second_half = []
for file in sorted(glob.glob(dfs + '/*.csv')):
df = pd.read_csv(file, sep = ';', index_col= [0])
first_half, second_half = np.array_split(df.index, 2)
if 'ch1' in file:
ch1_first_half.append(df.loc[first_half])
ch1_second_half.append(df.loc[second_half])
else:
ch2_first_half.append(df.loc[first_half])
ch2_second_half.append(df.loc[second_half])
# -
def calculateSynchronyFromDF(ch1_dfs, ch2_dfs, features):
import pandas as pd
import scipy.stats as stats
#ToDo: fix later with loop
feature_rows = {'MeanUnvoicedSegmentLength' : [],
'VoicedSegmentsPerSec' : [],
'F0semitoneFrom27.5Hz_sma3nz_amean' : [],
'loudness_sma3_amean' : [],
'F0semitoneFrom27.5Hz_sma3nz_stddevNorm' : []}
for ch1, ch2 in zip(ch1_dfs, ch2_dfs):
sub_id = ch1['sub_id'].unique()[0]
for feature in features:
speaker_1 = ch1[feature].to_numpy()
speaker_2 = ch2[feature].to_numpy()
#sometimes turns will be unequal, in that case drop the last one from the array
if len(speaker_1) > len(speaker_2):
speaker_1 = speaker_1[:-1]
elif len(speaker_1) < len(speaker_2):
speaker_2 = speaker_2[:-1]
speaker_1 = speaker_1[~np.isnan(speaker_2)] #drop nan turns from ch2 also from ch1
speaker_2 = speaker_2[~np.isnan(speaker_2)]
x = speaker_1[~np.isnan(speaker_1)] #drop nan turns from ch1 also from ch2
y = speaker_2[~np.isnan(speaker_1)]
#calculate synchrony using spearman r
r, p = stats.spearmanr(x, y)
#transform to z scores
r_z = np.arctanh(r)
#create dictionary with all the information
row = {'soundname': sub_id,
'r': r,
'p': p,
'r_z': r_z}
feature_rows[feature] += [row]
return feature_rows
# #### Calculate speech accommodation for the first and the second halves of the interview
feature_rows_first_half = calculateSynchronyFromDF(ch1_first_half, ch2_first_half, features)
feature_rows_second_half = calculateSynchronyFromDF(ch1_second_half, ch2_second_half, features)
# +
summary_dfs_first_half = {}
summary_dfs_second_half = {}
for feature, rows in feature_rows_first_half.items():
summary_dfs_first_half[feature] = pd.DataFrame(rows)
for feature, rows in feature_rows_second_half.items():
summary_dfs_second_half[feature] = pd.DataFrame(rows)
# -
# #### Load the group splits and compare halves of healthy controls and SZ patients separately
controls = np.loadtxt(os.path.join(dem_dir, 'control_subs.txt'), dtype= str)
patients = np.loadtxt(os.path.join(dem_dir, 'patient_subs.txt'), dtype= str)
def getGroupIndices(df, group):
group_indices = [k for k in df['soundname'] if k[:4] in group]
return group_indices
def pairedTestPerFeature(features, dfs_condition1, dfs_condition2, group):
import scipy.stats as stats
rows = {}
for feature in features:
row = {}
cond1 = dfs_condition1[feature]
cond2 = dfs_condition2[feature]
idxs_group = getGroupIndices(cond1, group) #the matching group subjects in the dataframe
x = cond1[cond1['soundname'].isin(idxs_group)]['r_z'] #select converted r value
y = cond2[cond2['soundname'].isin(idxs_group)]['r_z']
#paired ttest!
t, p = stats.ttest_rel(x, y)
row['T'] = t
row['p'] = p
rows[feature] = row
df = pd.DataFrame(rows)
return df.T
t_df_controls = pairedTestPerFeature(features, summary_dfs_first_half, summary_dfs_second_half, controls)
t_df_patients = pairedTestPerFeature(features, summary_dfs_first_half, summary_dfs_second_half, patients)
# #### Print results
t_df_controls
t_df_patients
# #### Repeat same process with conversation thirds instead of halves
# +
ch1_first = []
ch1_second = []
ch1_third = []
ch2_first = []
ch2_second = []
ch2_third = []
for file in sorted(glob.glob(dfs + '/*.csv')):
df = pd.read_csv(file, sep = ';', index_col= [0])
first, second, third = np.array_split(df.index, 3)
if 'ch1' in file:
ch1_first.append(df.loc[first])
ch1_second.append(df.loc[second])
ch1_third.append(df.loc[third])
else:
ch2_first.append(df.loc[first])
ch2_second.append(df.loc[second])
ch2_third.append(df.loc[third])
# -
# #### Calculate speech accommodation for each third of the interview
feature_rows_first = calculateSynchronyFromDF(ch1_first, ch2_first, features)
feature_rows_second = calculateSynchronyFromDF(ch1_second, ch2_second, features)
feature_rows_third = calculateSynchronyFromDF(ch1_third, ch2_third, features)
def makeDFsFromDict(feature_dict):
import pandas as pd
summary_dfs = {}
for feature, rows in feature_dict.items():
summary_dfs[feature] = pd.DataFrame(rows)
return summary_dfs
summary_dfs_first = makeDFsFromDict(feature_rows_first)
summary_dfs_second = makeDFsFromDict(feature_rows_second)
summary_dfs_third = makeDFsFromDict(feature_rows_third)
# #### Perform t-tests between the first and second and the second and third conversation part
# +
first_vs_second_cntrl = pairedTestPerFeature(features, summary_dfs_first, summary_dfs_second, controls)
second_vs_third_cntrl = pairedTestPerFeature(features, summary_dfs_second, summary_dfs_third, controls)
first_vs_second_sz = pairedTestPerFeature(features, summary_dfs_first, summary_dfs_second, patients)
second_vs_third_sz = pairedTestPerFeature(features, summary_dfs_second, summary_dfs_third, patients)
# -
# #### print results
first_vs_second_cntrl
second_vs_third_cntrl
first_vs_second_sz
second_vs_third_sz
# #### Plot the results
#
# To make use of seaborn's high level integration of dataframes we reshape the data a bit and take the mean for each conversation part, so that all values from all channels, features and time points are in one dataframe. For that we merge the individual dfs for each interview and add a few columns with extra information for conversation halves and thirds respectively.
import matplotlib.pyplot as plt
import seaborn as sns
# +
ch1_first_mean = []
ch2_first_mean = []
ch1_second_mean = []
ch2_second_mean = []
#for all interviews, take the mean
for ch1, ch2 in zip(ch1_first_half, ch2_first_half):
sub_id = pd.Series(data = ch1['sub_id'].unique()[0])
ch1_first_mean.append(ch1.mean().append(sub_id))
ch2_first_mean.append(ch2.mean().append(sub_id))
for ch1, ch2 in zip(ch1_second_half, ch2_second_half):
sub_id = pd.Series(data = ch1['sub_id'].unique()[0])
ch1_second_mean.append(ch1.mean().append(sub_id))
ch2_second_mean.append(ch2.mean().append(sub_id))
# +
#merge all interviews in one dataframe
ch1_first_df = pd.DataFrame(ch1_first_mean)
ch2_first_df = pd.DataFrame(ch2_first_mean)
ch1_second_df = pd.DataFrame(ch1_second_mean)
ch2_second_df = pd.DataFrame(ch2_second_mean)
# +
#merge the first and second half, add information which part each conversation belongs to
ch1_first_df['time'] = '1/2'
ch2_first_df['time'] = '1/2'
ch1_second_df['time'] = '2/2'
ch2_second_df['time'] = '2/2'
ch1 = pd.concat([ch1_first_df, ch1_second_df])
ch2 = pd.concat([ch2_first_df, ch2_second_df])
# +
ch1['channel'] = 'Interviewer'
ch2['channel'] = 'Participant'
conversation_halves = pd.concat([ch1, ch2])
# -
conversation_halves
# +
ch1_first_mean = []
ch1_second_mean = []
ch1_third_mean = []
ch2_first_mean = []
ch2_second_mean = []
ch2_third_mean = []
#for each interview, take the mean of the first, second and third part
for ch1, ch2 in zip(ch1_first, ch2_first):
sub_id = pd.Series(data = ch1['sub_id'].unique()[0])
ch1_first_mean.append(ch1.mean().append(sub_id))
ch2_first_mean.append(ch2.mean().append(sub_id))
for ch1, ch2 in zip(ch1_second, ch2_second):
sub_id = pd.Series(data = ch1['sub_id'].unique()[0])
ch1_second_mean.append(ch1.mean().append(sub_id))
ch2_second_mean.append(ch2.mean().append(sub_id))
for ch1, ch2 in zip(ch1_third, ch2_third):
sub_id = pd.Series(data = ch1['sub_id'].unique()[0])
ch1_third_mean.append(ch1.mean().append(sub_id))
ch2_third_mean.append(ch2.mean().append(sub_id))
# +
#merge all interviews into a dataframe
ch1_first_df = pd.DataFrame(ch1_first_mean)
ch2_first_df = pd.DataFrame(ch2_first_mean)
ch1_second_df = pd.DataFrame(ch1_second_mean)
ch2_second_df = pd.DataFrame(ch2_second_mean)
ch1_third_df = pd.DataFrame(ch1_third_mean)
ch2_third_df = pd.DataFrame(ch2_third_mean)
# +
#concatenate all parts, add information which third the row belongs to
ch1_first_df['time'] = '1/3'
ch2_first_df['time'] = '1/3'
ch1_second_df['time'] = '2/3'
ch2_second_df['time'] = '2/3'
ch1_third_df['time'] = '3/3'
ch2_third_df['time'] = '3/3'
ch1 = pd.concat([ch1_first_df, ch1_second_df, ch1_third_df])
ch2 = pd.concat([ch2_first_df, ch2_second_df, ch2_third_df])
# +
ch1['channel'] = 'Interviewer'
ch2['channel'] = 'Participant'
conversation_thirds = pd.concat([ch1, ch2])
# -
conversation_halves
conversation_thirds
# +
#filter the dataframes for controls and patients
conversation_halves_controls = conversation_halves.loc[conversation_halves[0].isin(controls)]
conversation_halves_patients = conversation_halves.loc[conversation_halves[0].isin(patients)]
conversation_thirds_controls = conversation_thirds.loc[conversation_thirds[0].isin(controls)]
conversation_thirds_patients = conversation_thirds.loc[conversation_thirds[0].isin(patients)]
# -
# #### Make a plot that shows the mean values for all speech features across the different time splits
# +
fig, axs = plt.subplots(nrows=len(features), ncols=2, figsize = (10, 20))
features_to_plot = sorted(features * 2)
#one column contains halves, the other thirds all other labels stay the same
y_labels = ['Pitch', 'Pitch',
'Pitch Variability', 'Pitch Variability',
'Average Pause Duration', 'Average Pause Duration',
'Syllable Rate', 'Syllable Rate',
'Loudness', 'Loudness']
#defining these manually so the y axes for halves and thirds are the same
y_lims = [(20, 35), (20, 35),
(0.1, 0.2), (0.1, 0.2),
(0, 0.6), (0, 0.6),
(1.5, 6), (1.5, 6),
(0.2, 0.8), (0.2, 0.8)]
#custom legend showing speaker and group attribution
legend_elements = [
Line2D([0], [0], marker='o', label='Interviewer', markerfacecolor='lightgrey', markersize=10, color = 'lightgrey'),
Line2D([0], [0], marker='x', label='Participant', markerfacecolor='grey', markersize=10, color = 'dimgrey'),
Line2D([0], [0], label='Control Group', linestyle = '--'),
Line2D([0], [0], label='Patient Group', color = 'red'),
]
plt.suptitle('Average Speech Features Across the Interviews', fontsize = 15, y=1.0, x =0.45)
for i, ax in enumerate(axs.flatten()):
#plot conversation halves on the left column
if (i % 2) == 0:
#plot interviewer + control participant
sns.pointplot(x="time", y=features_to_plot[i], hue="channel",
capsize=.2, height=6, aspect=.75,
kind="point", data=conversation_halves_controls, palette = "Blues", ax = ax,
markers=["o", "x"], linestyles=["--", "--"])
#plot interviewer + patient
sns.pointplot(x="time", y=features_to_plot[i], hue="channel",
capsize=.2, height=6, aspect=.75,
kind="point", data=conversation_halves_patients, ax = ax, palette = 'Reds',
markers = ['o', 'x'])
ax.get_legend().remove() #one legend per row
ax.set_ylabel(y_labels[i], fontsize = 14)
ax.set_xlabel('Conversation Halves', fontsize = 14)
ax.set_ylim(y_lims[i])
else:
#plot interviewer + control participant
sns.pointplot(x="time", y=features_to_plot[i], hue="channel",
capsize=.2, height=6, aspect=.75,
kind="point", data=conversation_thirds_controls, palette = "Blues", ax = ax,
markers=["o", "x"], linestyles=["--", "--"])
#plot interviewer + patient
sns.pointplot(x="time", y=features_to_plot[i], hue="channel",
capsize=.2, height=6, aspect=.75,
kind="point", data=conversation_thirds_patients, ax = ax, palette = 'Reds',
markers = ['o', 'x'])
#add custom legend
ax.legend(handles=legend_elements, loc='center left', bbox_to_anchor=(1, 0.5), fontsize = 13)
ax.set_ylabel('')
ax.set_xlabel('Conversation Thirds', fontsize = 14)
ax.set_ylim(y_lims[i])
plt.tight_layout()
| analysis/comparisons_over_time.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Join surrogate classes that are under the same node
#
#
# +
import numpy as np
import os
from shutil import copyfile, copytree
from tqdm import tqdm
import sys
sys.path.append('../../python_scripts')
from utils import read_images_stl10 as read_images
from torchvision import transforms
from PIL import Image
from matplotlib import pyplot as plt
import shutil
from os.path import join
sys.path.insert(0, './../scikit_learn')
sys.path.insert(0, './../')
from utils_clust import normalizing_samples_L2, loading_images, searching_similar_images
# +
# To load pairs of children and...
iteration_nb = '001_retrieval'
childrens_nb = '001'
path_pairs_out = './clusters/best_model/image_pairs' + childrens_nb
path_pairs_out_larger_clusters = './clusters/best_model/image_pairs_mixed_nodes' + childrens_nb
# other paths
path_target_dset_s_cl = './clusters/best_model/dset' + iteration_nb + '_short_cl/'
path_target_dset_l_cl = './clusters/best_model/dset' + iteration_nb + '_large_cl/'
path_new_classes = './clusters/best_model/new_classes_' + iteration_nb + '/'
path_target_dset_single = './clusters/best_model/dset' + iteration_nb + '_clusters_from_single_images'
path_single_old_classes = './clusters/best_model/ld_classes_' + iteration_nb + '/'
# source images
# 1st iteration path
#images_path = '../../surrogate_dataset/classes_folder_16000_set/'
images_path = '../../surrogate_dataset/training_set_100000/'
# 2nd iteration path
#images_path = './images_2nd_iteration/'
# 3rd iteration path
#images_path = './images_3rd_iteration/'
# +
short_clusters = []
larger_clusters = []
if os.path.exists(path_pairs_out + '.npy'):
short_clusters = np.load(path_pairs_out +'.npy')
if os.path.exists(path_pairs_out_larger_clusters + '.npy'):
larger_clusters = np.load(path_pairs_out_larger_clusters + '.npy')
# +
print 'Arrays loaded stats: '
print "children_array:",
print len(short_clusters), type(short_clusters), short_clusters.shape
#print "sub_child_int_array:",
#print len(sub_child_int_array)
print "sub_child_mixed_array:",
print len(larger_clusters), type(larger_clusters), larger_clusters.shape
# -
# ## Build first the large clusters
# +
# We need a set of all the images selected for clustering.
# From there we will remove each image when it is added to a cluster.
samples_set = set()
for cluster_i in larger_clusters:
for sample_i in cluster_i:
samples_set.add(sample_i)
print "Larger cplusters introduced. Length: ",
print len(samples_set)
# I introduce the samples from the simple clusters as well
for cluster_i in short_clusters:
for sample_i in cluster_i:
samples_set.add(sample_i)
print "Short clusters (pairs) introduced. Length: ",
print len(samples_set)
# +
# we need a set with all the 100k images from the dataset
samples_full = os.listdir(images_path)
samples_full_set = set([int(sample_i[:-4]) for sample_i in samples_full])
print len(samples_full), len(samples_full_set), type(samples_full_set)
# +
# first we invert the larger_cluster array. This is to start clustering the larger groups.
# This is needed because the tree structure made clusters that are inside other clusters...
larger_clusters = larger_clusters[::-1]
# Note that we do not need to do that with the simple clusters
# -
cluster_number = 0 # variable to count clusters
for cluster_i in larger_clusters:
nb_img = len(cluster_i)
num = 1
temp = 0 # variable to count clusters
for sample_i in cluster_i:
if sample_i in samples_set:
temp +=1 # variable to count clusters
## defining paths
if not os.path.exists(join(path_target_dset_l_cl, 'cl_l_' + str(cluster_number).zfill(6))):
os.makedirs(join(path_target_dset_l_cl, 'cl_l_' + str(cluster_number).zfill(6)))
# 'cl_l' stands for clusters large
src_path = join(images_path, str(sample_i).zfill(6) + '.png')
dst_path = join(path_target_dset_l_cl, 'cl_l_' + str(cluster_number).zfill(6), str(sample_i).zfill(6) + '.png')
## moving files to the new clusterred dataset
shutil.copyfile(src_path, dst_path)
### ploting...
#image = Image.open(src_path)
#plt.subplot(1,nb_img, num)
#plt.imshow(np.asarray(image))
#num += 1
### removing processed samples
samples_set.remove(sample_i)
samples_full_set.remove(sample_i)
if temp > 0:
cluster_number += 1 # variable to count clusters
plt.show()
print "Number of larger clusters:",
print cluster_number
# +
print "Set length after processing the large clusters: ",
print len(samples_set)
print "Set length of the full set after processing the large clusters: ",
print len(samples_full_set)
# -
cluster_number = 0 # variable to count clusters
for cluster_i in short_clusters:
nb_img = len(cluster_i)
num = 1
temp = 0 # variable to count clusters
for sample_i in cluster_i:
if sample_i in samples_set:
temp +=1 # variable to count clusters
## defining paths
if not os.path.exists(join(path_target_dset_s_cl, 'cl_s_' + str(cluster_number).zfill(6))):
os.makedirs(join(path_target_dset_s_cl, 'cl_s_' + str(cluster_number).zfill(6)))
# 'cl_l' stands for clusters large
src_path = join(images_path, str(sample_i).zfill(6) + '.png')
dst_path = join(path_target_dset_s_cl, 'cl_s_' + str(cluster_number).zfill(6), str(sample_i).zfill(6) + '.png')
## moving files to the new clusterred dataset
shutil.copyfile(src_path, dst_path)
### ploting...
#image = Image.open(images_path + str(sample_i).zfill(6) + '.png')
#plt.subplot(1,nb_img, num)
#plt.imshow(np.asarray(image))
#num += 1.
### removing processed samples
samples_set.remove(sample_i)
samples_full_set.remove(sample_i)
if temp > 0:
cluster_number += 1 # variable to count clusters
plt.show()
print "Number of smaller clusters:",
print cluster_number
# +
print "Set length after processing the short clusters: ",
print len(samples_set)
print "Set length of the full set after processing the large clusters: ",
print len(samples_full_set)
# -
# # Introduce "single cluster" (the images not clustered) to the dataset
# +
# first we extract the images that we need to reach again the 16000 classes
max_classes = 16000
nb_new_classes = max_classes - (len(os.listdir(path_target_dset_l_cl)) + len(os.listdir(path_target_dset_s_cl)))
print "Checking numbers..."
print nb_new_classes
# +
# I will get the images from the unsup set
single_samples = [image_i for image_i in list(samples_full_set)[:nb_new_classes]]
# Save the images in a folder
if not os.path.exists(path_new_classes):
os.makedirs(path_new_classes)
for idx in tqdm(single_samples):
path = os.path.join(path_new_classes, str(idx).zfill(6))
image = Image.open(images_path + str(idx).zfill(6) + '.png')
samples_full_set.remove(idx)
image.save(path + '.png')
# +
print "Set length after processing the short clusters: ",
print len(samples_set)
print "Set length of the full set after processing the large clusters: ",
print len(samples_full_set)
# -
# ## Searching the nearest images from unlabeled images - Single images
single_images = os.listdir(path_new_classes)
single_images.sort()
# +
# We need the features to do the query search
# paths
features_path = './features_maxpool_allConv.hdf5'
# load images
samples = loading_images(features_path)
# normalize images
samples_L2 = normalizing_samples_L2(samples)
# +
query_mtx = np.zeros([len(single_images), len(samples_L2[0])])
query_names = []
for num, sample_i in enumerate(single_images):
query = samples_L2[int(sample_i[:-4])]
query_names.append(sample_i)
query_mtx[num] = query
# compute cosine similarity
sim = np.dot(query_mtx, samples_L2.T)
# sort ranking
ranks = np.argsort(sim, axis=1)[:,::-1]
# +
cluster_size = 4
for num, sample_i in enumerate(query_names):
cluster_i = []
cluster_i.append(sample_i)
instance_num = 1
while len(cluster_i)<cluster_size:
image_nb = ranks[num][instance_num]
image_name = str(image_nb).zfill(6) + '.png'
if image_nb in samples_full_set:
cluster_i.append(image_name)
samples_full_set.remove(image_nb)
instance_num += 1
# move the images fom the cluster_i to the new folder
src_path = join(path_new_classes, cluster_i[0])
dst_folder = join(path_target_dset_single, cluster_i[0][:-4])
if not os.path.exists(dst_folder):
os.makedirs(dst_folder)
dst_path = join(dst_folder, cluster_i[0])
shutil.copyfile(src_path, dst_path)
for num_cl, img_i in enumerate(cluster_i[1:]):
src_path = join(images_path, cluster_i[num_cl+1])
dst_path = join(dst_folder, img_i)
shutil.copyfile(src_path, dst_path)
# +
print "Set length after processing the short clusters: ",
print len(samples_set)
print "Set length of the full set after processing the large clusters: ",
print len(samples_full_set)
# +
# after that i should do the same with the other clusters with 2 images each...
# -
print sim.shape
print sim[:5]
print sim[-5:]
print ''
print max(sim)
print min(sim)
sim_ord = np.argsort(sim)
print ''
print sim_ord[:5]
print sim_ord[-5:]
...
# finally we move all the single classes to a folder inside the "dataset00?" folder
list_files = os.listdir(path_new_classes)
for sample_i in tqdm(list_files):
src_path = join(path_new_classes, sample_i)
dst_folder = join(path_target_dset, sample_i[:-4])
if not os.path.exists(dst_folder):
os.makedirs(dst_folder)
dst_path = join(dst_folder, sample_i)
shutil.copyfile(src_path, dst_path)
# ## Searching the nearest images from unlabeled images - Simple clusters
# #### This one is the same but changing the last cell for some code to search for near images in the 50000 samples left. The aim is get larger clusters for the single images
#
# paths
features_path = './features_maxpool_allConv.hdf5'
# load images
samples = loading_images(features_path)
# normalize images
samples_L2 = normalizing_samples_L2(samples)
current_images = len(os.listdir(images_path))
current_images
# +
unlab_set = read_images('../../data/stl10_binary/unlabeled_X.bin')
np.random.seed(42)
# indexes drawn in a set to avoid duplicates
indexes = set()
while len(indexes) < (current_images + nb_new_classes): # we do that to get only the last ones
indexes.add(np.random.randint(unlab_set.shape[0]))
# Save the images in a folder
toPill = transforms.Compose([transforms.ToPILImage()])
if not os.path.exists(path_new_classes):
os.makedirs(path_new_classes)
for num, idx in tqdm(enumerate(list(indexes)[-nb_new_classes:])):
path = os.path.join(path_new_classes, str(num + current_images).zfill(len(str(max_classes))))
image = toPill(unlab_set[idx])
image.save(path + '.png')
# +
# now we copy the images that we did not use during the clustering in another folder
if not os.path.exists(path_single_old_classes):
os.makedirs(path_single_old_classes)
for sample_i in tqdm(samples_full_set):
src_path = join(images_path, str(sample_i).zfill(5) + '.png')
dst_path = join(path_single_old_classes, str(sample_i).zfill(5) + '.png')
shutil.copyfile(src_path, dst_path)
# +
# finally we move all the single classes to a folder inside the "dataset00?" folder
list_files = os.listdir(path_new_classes)
for sample_i in tqdm(list_files):
src_path = join(path_new_classes, sample_i)
dst_folder = join(path_target_dset, sample_i[:-4])
if not os.path.exists(dst_folder):
os.makedirs(dst_folder)
dst_path = join(dst_folder, sample_i)
shutil.copyfile(src_path, dst_path)
list_files = os.listdir(path_single_old_classes)
for sample_i in tqdm(list_files):
src_path = join(path_single_old_classes, sample_i)
dst_folder = join(path_target_dset, sample_i[:-4])
if not os.path.exists(dst_folder):
os.makedirs(dst_folder)
dst_path = join(dst_folder, sample_i)
shutil.copyfile(src_path, dst_path)
# -
# ### Now is time to compute the transformations from the new clustered dataset...
| clustering/code_to_do_the_clustering/clustering_children_005_not_iterative_retrieval.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.7 64-bit
# language: python
# name: python3
# ---
# # Jupyter Notebooks
# **Jupyter** notebooks are commonly used to combine Markdown and Python into a single file. For example, this file is a notebook file! Imagine a blog post where you can talk about your thought process, show your code, and also let readers run the code.
#
# The great things about cells is that you can run specific sections of code at a time. You can also quickly see the return values of each cell without running the entire file.
# Try running this cell!
print("Hello, World")
# # Pandas
# 1. What happens when you run the code below? Make sure you change the path to point to where `penguins.csv` is stored on your own computer.
# +
# We're going to work with a new library called pandas
import pandas as pd
penguins = pd.read_csv("../../Datasets/penguins.csv")
penguins
# -
# 2. What happens when you run the code below? Add a cell block to access the column for species.
penguins["bill_length_mm"]
# 3. What happens when you run the code below? Add a cell block to list all Adelie penguins.
penguins[(penguins["species"] == "Gentoo") | (penguins["species"] == "Adelie")][["species","flipper_length_mm"]]
# You have been working with the **Dataframe** class that is provided by the `pandas` module! As you can see, Dataframes allow you to quickly store and access tabular information.
#
# 4. Add a cell block that _describes_ the flipper length of all penguins.
penguins["bill_length_mm"].describe()
penguins.bill_length_mm
| art of data/topics/descriptive stats/intro_to_pandas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="9oAXesa2Rcg-" colab_type="code" colab={}
#Program on Loops
#program1:Write a Python program to find those numbers which are divisible by 7 and multiple of 5, between 1500 and 2700 (both included).
for i in range(1500,2700):
list=[]
if (i%7==0) and (i%5==0):
print(i)
# + id="p8OY-8PUSjui" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 87} outputId="04369a12-1f91-4e10-af03-40d810e5810c"
#program2:Write a Python program to convert temperatures from celsius, fahrenheit.
a=int(input(''' *enter 1 to convert temperature from celsius to farenheit
*enter 2 to convert temperature from farenheit to celsius'''))
b=int(input('Enter the temperature'))
if (a==1):
c=(b-(32/9))*5
print(c)
else:
f=((b/5)+(32/9))
print(f)
# + id="oUwVzISHUQZE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="e5cdf354-c026-42c1-cb50-896915349cc0"
#Program3:Write a Python program to guess a number between 1 to 9.
from random import randint #Importing randint from random module
a=int(input('Enter the number between 0 to 9:'))
b=random.randint(0,9) #This generates the numbers between 0 to 9
if (a==b):
print("It is a right match")
else:
print("The entered number is not a right match")
# + id="eLMHZITmSj0K" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="cc33b9ef-2f8b-4817-f667-7f2ed6384573"
#program5:Write a Python program that accepts a word from the user and reverse it.
a='Google'
b=''
for i in a:
b=i+b
print(b)
# + id="BIS-C9y8Sj2o" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="9a660875-3cc2-4c78-a86c-06dcc50779d7"
#program6:Write a Python program to count the number of even and odd numbers from a series of numbers.
a= (1, 2, 3, 4, 5, 6, 7, 8, 9)
j=0
for i in range(0,len(a)):
if (i%2==0):
j=j+1
print("The total number of even numbers",j)
print("The total number of odd numbers",len(a)-j)
# + id="Ep30Dyr9WRnj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 158} outputId="883de4e6-f7aa-41cd-af2a-c4f7a2be965c"
#Program7:Write a Python program that prints each item and its corresponding type from the following list.
a= [1452, 11.23, 1+2j, True, 'w3resource', (0, -1), [5, 12], {"class":'V', "section":'A'}]
for i in a:
print(type(i))
# + id="ac6l-C9OSj5H" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 105} outputId="ea9e9fbf-00c7-42b9-ac22-bb0d962b86b7"
#program8:Write a Python program that prints all the numbers from 0 to 6 except 3 and 6.
for i in range(0,7):
if (i==3) or (i==6):
continue
else:
print(i)
# + id="uffJ5rPrSj7s" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 176} outputId="d04b0cab-427c-4b8d-fa20-e17003f7f694"
#program9:Write a Python program to get the Fibonacci series between 0 to 50.
a,b=0,1
while(b<50):
print(b)
a,b=b,a+b
# + id="j52O9YlmSj-D" colab_type="code" colab={}
#program 10:Write a Python program which iterates the integers from 1 to 50. For multiples of three print "Fizz" instead of the number and for the multiples of five print "Buzz". For numbers which are multiples of both three and five print "FizzBuzz".
for i in range(0,50):
if (i%5==0) and (i%3==0):
print("FizzBuzz")
elif (i%3==0):
print("Fizz")
elif (i%5==0):
print("Buzz")
else:
print(i)
# + id="g5uJ5s5BSkAb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="116eda77-35a4-4828-9ed8-15b704e4f4f8"
#program11:Write a Python program which takes two digits m (row) and n (column) as input and generates a two-dimensional array. The element value in the i-th row and j-th column of the array should be i*j.
b=[]
for i in range(0,3):
b.append([])
for j in range(0,4):
b[i].append(j)
print(b)
# + id="GMCekVNhSkC8" colab_type="code" colab={}
#program 12:Write a Python program that accepts a sequence of lines (blank line to terminate) as input and prints the lines as output (all characters in lower case).
a="This is Python Class"
b=[]
while True:
if a:
b.append(a.lower())
for a in b:
print(a)
# + id="8qSFbEuwSkF2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="f3fbfd3f-4693-4895-cdd9-121f74128ead"
#program 13:Write a Python program which accepts a sequence of comma separated 4 digit binary numbers as its input and print the numbers that are divisible by 5 in a comma separated sequence
list=[]
num=[x for x in input().split(',')]
for i in num:
x=int(i,2)
if not x%5:
list.append(i)
print(','.join(list))
# + id="1OvxhHnRSkIS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="be0a363e-895a-4a94-8ed2-b96da6a2e393"
#program 14:Write a Python program that accepts a string and calculate the number of digits and letters.
a='Python 3.2'
j=0
k=0
for i in a:
if i.isalpha():
j+=1
elif i.isdigit():
k+=1
print("The number of letters",j)
print("The number of digits",k)
# + id="h6iE_tFqSkLE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="4a31d175-cb2e-4706-c7a6-4ab0ce846239"
#program 15: Write a Python program to check the validity of password input by users. Go to the editor
#Validation
import re
a=input('input the password')
b=True
while b:
if (len(a)<6) or (len(a)>12):
break
elif not re.search("[a-z]",a):
break
elif not re.search("[A-Z]",a):
break
elif not re.search("[0-9]",a):
break
else:
print("valid password")
x=False
break
# + id="7kEFFGI2SkN6" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="ccc4b573-2bb6-4bad-8219-d70d17a947d6"
#program 16:Write a Python program to find numbers between 100 and 400 (both included) where each digit of a number is an even number. The numbers obtained should be printed in a comma-separated sequence.
list=[]
for i in range(100,400):
s=str(i)
if (int(s[0])%2==0) and (int(s[1])%2==0) and (int(s[2])%2==0):
list.append(s)
print(",".join(list))
# + id="jzAgK4Vdr7M4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="879eff9b-a117-4f01-c80a-9a74b2a93563"
#program 17:Write a Python program to calculate a dog's age in dog's years.
a=int(input("Enter the dogs age"))
if (a>2):
x=a-2
y=2
sum1=2*10.5
sum2=x*4
total=sum1+sum2
print("The dogs age in dogs years are:",total)
# + id="da4mvKm2sWb_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="2edfaf8b-1ad6-464c-e024-63b4b8a467bf"
#program 18:Write a Python program to check whether an alphabet is a vowel or consonant.
a=input('enter the character:')
for m in a:
if (m=='a'or m== 'e' or m=='i' or m=='o' or m=='u'):
print("The entered character is vowel")
else:
print("The entered charcter is consonant")
# + id="QoB3gAkPsWfj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="2fdabb5c-7615-47a5-9d7b-864e8f27201f"
#program 19:Write a Python program to convert month name to a number of days.
a=input("Enter the month")
if (a=='January') or (a=='July') or (a=='August') or (a=='December') or (a=='March'):
print("The number of days are 31 DAYS")
elif (a=='April' or a=='June' or a=='September' or a=='October' or a=='November'):
print("The number of days are 30 DAYS")
else:
print("The number of days are 28 Days")
# + id="azx7MrZTsWjT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 70} outputId="1ad6612f-6212-4cad-a23d-f616ecf8870d"
#program 20: Write a Python program to sum of two given integers. However, if the sum is between 15 to 20 it will return 20.
a=int(input("enter the number"))
b=int(input("enter the number"))
sum=a+b
if sum in range(15,20):
print(20)
else:
print(sum)
# + id="SYso9yqHIzxf" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 87} outputId="74d62dd3-5d22-402e-ff1d-71422491c8d7"
#program 21:Write a Python program to check a triangle is equilateral, isosceles or scalene.
a=int(input("enter the length of side 1"))
b=int(input("enter the length of side 2"))
c=int(input("enter the length of side 3"))
if (a==b==c):
print("The given triangle is equilateral traiangle")
elif (a==b or a==c):
print("The given traiangle is Isoceles triangle")
else:
print("The given traingle is Scalene traingle")
# + id="1sSKd1JAgRDh" colab_type="code" colab={}
#Program 22:Write a Python program that reads two integers representing a month and day and prints the season for that month and day.
month=input('ENTER THE MONTH')
day=int(input("Input the day"))
if month in ('January','February','March'):
season='winter'
elif month in ('April','May','June'):
season='Spring'
elif month in ('July','August','September'):
season='summer'
else:
season='autumn'
if (month=='March') and (day>19):
season='spring'
elif (month=='June') and (day>20):
season='summer'
elif (month=='September') and (day>21):
season='autumn'
elif (month=='December') and (day>20):
season='winter'
print(season)
# + id="H9Xl-6VhHKNT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 87} outputId="76b9a7c5-3926-40d4-9407-dd39d066f656"
#program 23:Write a Python program to find the median of three values.
a= int(input("enter the number"))
b= int(input("enter the number"))
c= int(input("enter the number"))
if a>b and a>c:
if b>c:
print("the median is ",b)
else:
print("the mediam is",c)
if b>a and b>c:
if a>c:
print("the median is ",a)
else:
print("the mediam is",c)
if c>a and c>b:
if a>b:
print("the median is ",a)
else:
print("the mediam is",b)
# + id="PDKDFbL9HKP-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 87} outputId="48fcffa8-8165-4b42-e6f8-9f497890f8fb"
#program 24:Write a Python program to calculate the sum and average of n integer numbers (input from the user).
a=int(input('enter the starting number'))
b=int(input('enter the last number'))
c=b+1
sum=0
for i in range(a,c):
n=(c-a)
sum=sum+i
average=sum/n
print(sum)
print(average)
# + id="6QzjGMllHKSl" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 212} outputId="34653dbb-3e2d-42cd-fee7-87d34d2c6a3c"
#program 25:Write a Python program to create the multiplication table (from 1 to 10) of a number.
a=int(input('enter the number'))
for i in range(1,11):
print(a*i)
# + id="Kw9RmI1UkiLe" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 194} outputId="e0a3f0de-30ca-4a90-a441-a807678e49fb"
#program 26:Write a Python program to construct the following pattern, using a nested loop number.
for i in range(10):
print(str(i)*i)
# + id="T90wgKdWHKVL" colab_type="code" colab={}
| Harshitha/Practice problems/loops.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # OP2: Numpy Demo #1 (Displacement, Solid Stress)
#
# The Jupyter notebook for this demo can be found in:
# - docs/quick_start/demo/op2_demo_numpy2.ipynb
# - https://github.com/SteveDoyle2/pyNastran/tree/master/docs/quick_start/demo/op2_demo_numpy2.ipynb
#
# It's recommended that you first go through:
# - https://github.com/SteveDoyle2/pyNastran/tree/master/docs/quick_start/demo/op2_intro.ipynb
# The previous demo was intentionally clunky to demonstrate how one might think of a single element.
#
# If you code like that, your code will be slow, so let's show you how to really use the numpy-style with the OP2.
#
# 
#
# ### Import the packages
# +
import os
import copy
import numpy as np
np.set_printoptions(precision=2, threshold=20, suppress=True, linewidth=100)
import pyNastran
pkg_path = pyNastran.__path__[0]
model_path = os.path.join(pkg_path, '..', 'models')
from pyNastran.utils import print_bad_path
from pyNastran.op2.op2 import read_op2
from pyNastran.utils import object_methods, object_attributes
np.set_printoptions(precision=3, threshold=20, edgeitems=10)
# -
# ### Load the model
op2_filename = os.path.join(model_path, 'solid_bending', 'solid_bending.op2')
model = read_op2(op2_filename, build_dataframe=False, debug=False)
# ### Find the min/max Displacement magnitude
#
# In this example, we access the 3D "data" numpy array object. Then we take the L2-norm of the translations to determine the magnitude. We broadcast the L2-norm across the column (x, y, z) to end up with **nnodes** results. It's good practice to verify the shapes of your arrays just to make sure you get the **axis=1** parameter correct.
# +
subcase_id = 1
disp = model.displacements[subcase_id]
disp_headers = disp.get_headers()
print('disp_headers = %s' % disp_headers)
nnodes = disp.node_gridtype.shape[0]
txyz = disp.data[0, :, :3]
txyz_mag = np.linalg.norm(txyz, axis=1)
assert len(txyz_mag) == nnodes
txyz_mag_max = txyz_mag.max()
txyz_mag_min = txyz_mag.min()
inid_max = np.where(txyz_mag == txyz_mag_max)[0]
inid_min = np.where(txyz_mag == txyz_mag_min)[0]
all_nodes = disp.node_gridtype[:, 0]
max_nodes = all_nodes[inid_max]
min_nodes = all_nodes[inid_min]
print('max displacement=%s max_nodes=%s' % (txyz_mag_max, max_nodes))
print('min displacement=%s min_nodes=%s' % (txyz_mag_min, min_nodes))
# -
# ### Find the max centroidal stress on the CTETRA elements
# +
subcase_id = 1
stress = model.ctetra_stress[subcase_id]
stress_headers = stress.get_headers()
print('stress_headers = %s' % stress_headers)
element_node = stress.element_node
elements = element_node[:, 0]
nodes = element_node[:, 1]
#print(element_node)
# -
# ### The 0 location is the centroid
#
# You can either query the 0 location or calculate it with a numpy arange. CTETRA elements have 4 nodes (even 10 noded CTETRA elements) in the OP2.
# +
izero = np.where(nodes == 0)[0]
izero2 = np.arange(0, len(nodes), step=5, dtype='int32')
#print(izero)
#print(izero2)
eids_centroid = elements[izero2]
print('eids_centroid = %s' % eids_centroid)
ivm = stress_headers.index('von_mises')
vm_stress = stress.data[0, izero2, ivm]
print(vm_stress)
vm_stress_max = vm_stress.max()
vm_stress_min = vm_stress.min()
icentroid_max = np.where(vm_stress == vm_stress_max)[0]
icentroid_min = np.where(vm_stress == vm_stress_min)[0]
eids_max = eids_centroid[icentroid_max]
eids_min = eids_centroid[icentroid_min]
print('max_stress=%s eids=%s' % (vm_stress_max, eids_max))
print('min_stress=%s eids=%s' % (vm_stress_min, eids_min))
# -
# ## Finding the VM stress associated with a single node ID
#
# One node in a tet mesh may be shared by many elements. In this case, 26 elements share 1 node!
# +
subcase_id = 1
stress = model.ctetra_stress[subcase_id]
stress_headers = stress.get_headers()
print('stress_headers = %s' % stress_headers)
element_node = stress.element_node
elements = element_node[:, 0]
nelements = len(elements) // 5
nodes = element_node[:, 1]#.reshape(nelements, 5)
#------------------------------
ivm = -1
print('nodes =', nodes)
ifour = np.where(nodes == 4)[0]
eids_four = elements[ifour].tolist()
print('eids4 =', eids_four)
print('ifour =', ifour)
vm_stress = stress.data[0, ifour, ivm]
print('vm_stress =', vm_stress, len(vm_stress))
# -
# ## Finding the centroidal VM stress for a set of elements
#
# Some fancy numpy code will be used for this case. Your code will be much faster if you are familiar with numpy.
# +
subcase_id = 1
stress = model.ctetra_stress[subcase_id]
stress_headers = stress.get_headers()
print('stress_headers = %s' % stress_headers)
element_node = stress.element_node
elements = element_node[:, 0]
nodes = element_node[:, 1]
# the slow way to get the unique elements
izero = np.where(nodes == 0)[0]
ueids_slow = elements[izero]
# the fast way
ueids = np.unique(elements)
assert np.array_equal(ueids, ueids_slow)
eids_to_lookup = [5, 7, 10]
ilookup = np.searchsorted(ueids, eids_to_lookup)
ivm = stress_headers.index('von_mises')
vm_stress = stress.data[0, ilookup, ivm]
print('eids_to_lookup =', eids_to_lookup)
print('vm_stress =', vm_stress)
# -
# ## Finding the centroidal VM stress for a set of elements when you have multiple element types
#
# In this case, we'll assume the set of element_ids to lookup contain CHEXAs as well as CTETRAs.
# Thus, we need to filter the data.
# +
subcase_id = 1
stress = model.ctetra_stress[subcase_id]
stress_headers = stress.get_headers()
print('stress_headers = %s' % stress_headers)
element_node = stress.element_node
elements = element_node[:, 0]
nodes = element_node[:, 1]
ueids = np.unique(elements)
print('ueids', ueids)
eids_to_lookup = [5, 7, 10, 186, 1000000]
ilookup = np.searchsorted(ueids, eids_to_lookup)
ivm = stress_headers.index('von_mises')
vm_stress = stress.data[0, ilookup, ivm]
print('eids_to_lookup =', eids_to_lookup)
print('vm_stress =', vm_stress)
# -
# ### We have a problem where our element_id (1000000) is out of range
#
# Searchsorted is fast, but you need to make sure your data actually exists. Otherwise, you'll end
# up finding the data for the next element in the sorted list.
#
# Let's filter the data using sets and then use searchsorted.
# +
eids_to_lookup = [5, 7, 10, 186, 1000000]
filtered_eids = np.intersect1d(elements, eids_to_lookup)
ilookup = np.searchsorted(ueids, filtered_eids)
vm_stress = stress.data[0, ilookup, ivm]
print('filtered_eids =', filtered_eids)
print('vm_stress =', vm_stress)
# -
# ### Other Elements that are Similar
#
# - Rod Stress/strain
# - Beam Stress/strain
# - Bar Stress/strain
# - Isotropic CQUAD4 stress/strain
| docs/quick_start/demo/op2_demo_numpy1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# <img src="http://csdms.colorado.edu/mediawiki/images/CSDMS_high_res_weblogo.jpg">
# # HydroTrend Study with the Stochastic Collocation Method
# [HydroTrend](https://csdms.colorado.edu/wiki/Model:HydroTrend) is a numerical model that creates synthetic river discharge and sediment load time series as a function of climate trends and basin morphology.
#
# In this example, we'll perform an experiment on HydroTrend using the [stoch_collocation](https://dakota.sandia.gov//sites/default/files/docs/6.4/html-ref/method-stoch_collocation.html) method
# to evaluate how changing two input parameters:
#
# * `starting_mean_annual_temperature` (_T_) and
# * `total_annual_precipitation` (_P_)
#
# affects the median values of one output parameter,
# long-term suspended sediment load at the river mouth (_Qs_),
# over a 10-year run.
# Before we start, make sure that you've installed Dakota, HydroTrend, and this package on your computer, using the instructions in the [README](https://github.com/csdms/dakota/blob/master/README.md) file.
# Start by importing the Dakota class.
from dakotathon import Dakota
# Create a Dakota instance to perform a study of HydroTrend using the stochastic collocation method and input parameters characterized by normal distributions.
d = Dakota(method='stoch_collocation', variables='normal_uncertain', plugin='hydrotrend')
# Define the HydroTrend input variables (_T_ and _P_) to be used in the study.
# We assume they're random variables with normal distributions,
# and
# we can obtain their default mean and standard deviation values
# from the HydroTrend parameters file (discussed below).
d.variables.descriptors = ['starting_mean_annual_temperature', 'total_annual_precipitation'] # T and P
d.variables.means = [14.26, 1.59]
d.variables.std_deviations = [0.55, 0.30]
# Define the HydroTrend outputs to be used in the study, as well as the statistics to be calculated from them.
d.responses.response_descriptors = 'Qs_median'
d.responses.response_files = 'HYDROASCII.QS'
d.responses.response_statistics = 'median'
# In the stochastic collocation method,
# a polynomial expansion is computed for the response function
# from four Gauss points in both _T_ and _P_,
# for a total of 16 integration points.
# From this expansion, 1000 samples are selected using Latin hypercube sampling to perform the analysis.
# The method calculates the PDF and the CDF of _Qs_ using the bins specified by the `response_levels` keyword.
# Variance-based decomposition is turned on to provide additional sensitivity statistics,
# such as Sobol' indices.
d.method.quadrature_order = 4
d.method.sample_type = 'lhs'
d.method.samples = 1000
d.method.seed = 17
d.method.response_levels = [2.5, 3.0, 3.5, 4.0, 4.5, 5.0, 5.5]
d.method.variance_based_decomp = True
# HydroTrend requires a set of files to run.
# They're included in the **data** directory of the repository containing this example.
# They can also be obtained directly from the HydroTrend [GitHub repository](https://github.com/mcflugen/hydrotrend/tree/add-bmi-metadata).
# Set paths to these files with the following statements.
# +
import os
data_dir = os.path.join(os.getcwd(), 'data')
template_file = os.path.join(data_dir, 'hydrotrend.in.tmpl')
parameters_file = os.path.join(data_dir, 'parameters.yaml')
hypsometry_file = os.path.join(data_dir, 'HYDRO0.HYPS')
# -
# The *template file* provides the configuration file for HydroTrend, but with all parameter values replaced by variables in the form `{parameter_name}`. The *parameters file* provides descriptions, ranges, and default values for all of the parameters represented in the template file. The *hypsometry file* describes the change in elevation along the river's course from source to sea.
# From the template and parameters files,
# we can create an input file that HydroTrend can run.
# Included in the CSDMS Dakota package is a routine that replaces the variables in the template file with default values from the parameters file.
# Import this routine and use it to create a HydroTrend input file.
# +
from dakotathon.plugins.base import write_dflt_file
default_input_file = write_dflt_file(template_file, parameters_file, run_duration=10*365)
print default_input_file
# -
# Next, we must replace the default values for the variables for `starting_mean_annual_temperature` and `total_annual_precipitation` with variable names for Dakota to substitute into. The CSDMS Dakota package also includes a routine to do this. Import this routine and use it to create a Dakota template file.
# +
from dakotathon.plugins.base import write_dtmpl_file
dakota_template_file = write_dtmpl_file(template_file, default_input_file, d.variables.descriptors)
print dakota_template_file
# -
# Associate the Dakota template file and the hypsometry file with the Dakota instance.
d.template_file = dakota_template_file
d.auxiliary_files = hypsometry_file
# Call the setup method to create files needed by Dakota, then run the experiment.
d.setup()
d.run()
# Check the output. First, the **dakota.dat** file. It shows the quadrature points, and the median values of _Qs_ calculated at those points.
# %cat dakota.dat
# Next, the **dakota.out** file.
# At the end of this file,
# statistics calculated by Dakota are reported.
# %cat dakota.out
# The coefficient of variation, $c_v = \sigma / \mu$,
# can be used as a measure of the uncertainty in the inputs, _T_ and _P_.
# For _T_, $c_v = 1.59/14.26 = 3.9\%$,
# while for _P_, $c_v = 0.30/1.59 = 18.9\%$.
# These uncertainties are propagated through the model by Dakota,
# which calculates the first four moments of the response, _Qs_.
# From these moments, we can also calculate a coefficient of variation for _Qs_,
# $c_v = 0.346/4.34 = 8.0\%$.
#
# Sobol' indices are calculated, which provide a measure of sensitivity of _Qs_ to the model inputs.
# For this study,
# 75 percent of the variance in _Qs_ is caused by variance in _T_,
# while only about 25 percent is caused by _P_,
# and with marginal interactions between the two.
| examples/hydrotrend-stochastic-collocation-study.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# > Created on Sun Jan 17 09:46:23 2021 @author: <NAME>-Chicago.IIT
# # [Guide to the knowledge structure of th digital LA|UP|Archi design](https://github.com/richieBao/guide_to_digitalDesign_of_LAUPArhi_knowledgeStruc)
#
# The rapid rise of parametric design in landscape architecture in 2007 was attributed to the visualization programming tool Grasshopper(GH) conceived by <NAME> in Robert McNeel and Associates ( Rhino 3D design model software creators). It is based on a Graphical User Interface(GUI) scripting engine that uses drag-and-drop data blocks and functions (called components) wired together. And it allows users who do not understand text programming to manipulate the Rhino tool using computational logic and generate highly complex 3-dimensional models. In the landscape architecture field, GH is used not only for design projects but also for academic projects. For example, Fletcher Studio used GH simulation environments extensively and created concept programming packages and construction documentation for a 2.8 million dollars renovation project in San Francisco's South Park. Influenced by GH, to simplify the AEC(Architecture, Engineering, and Construction) workflow in Revit software used for Building Information Modeling(BIM) construction, visual programming tool Dynamo was developed in 2013 to realize the integration of parameterization in BIM software.
#
# Parametric tools for visual programming are often not closed design environment, but rather textual scripting languages such as Python, commonly used, and C#. It dramatically expands the freedom and extensibility of designers to write design logic or algorithms. For example, based on the extension of the GH library of about 434 groups, in the interface, data management, spatial geometrical construction and analysis, algorithm, intelligent design, structure, and sustainable design, BIM and manufacturing and robot construction, 3d printing, GIS(Geographic Information System), and the interaction of hardware and so on.。
#
# Simultaneously, another voice is also rising, big data, machine learning, deep learning, and intelligent design.
#
# > About this lecture's format, I thought about some forms of presentation, PowerPoint presentation? MindManager mind map? Markdown text files? I remember that I used PPT long ago and used it to form planning and design text directly. Later, I was adapted to the form of a mind map, and I could freely think about the relationship between the issues. I did not have to worry about how the PPT should be typeset or even think about the base map, which restricted the thinking content's speed. While making fair use of the mind map, when I am a coder, I find that the coder's main thinking habit is 'laziness', which is focused on what should concentrate and leave the rest to the code to implement. Markdown then became the unwritten popular note of the code world. Then there was the Jupyter (a mashup of Markdown, code interactive interpreters, and plain text), which wrote code and took notes as it ran. So I thought about explaining digital design in the form of JupyterLab, which seems to fit the theme.
#
# > At the same time, I also thought that the maximum effect of teaching should be to share as much as possible and practice, which is especially important in the world of code. So you can get the content of the handout through [guide_to_digitalDesign_of_LAUPArhi_knowledgeStruc](https://github.com/richieBao/guide_to_digitalDesign_of_LAUPArhi_knowledgeStruc),GitHub code hosting.
#
# ## 1.Before and after 2010
#
# > *It does not matter what tools you use, as long as it is a good design or planning.*
#
# 2010 may be a better demarcation point, the necessary components of Grasshopper almost finalize, related extension components also probably includes all directions. At the same time, big data and machine learning began to emerge. The appearance of the Sklearn library of Python marked the popularization of machine learning. The emergence of TensorFlow marked the beginning of the popularity of deep learning, and the emergence of PyTorch accelerated this trend.
#
# ### 1.1 In the era of handcraft
# This stage will not go away because it is a necessary stage to learn design. Regardless of logical thinking, algorithms, machine/deep learning, it is essential to think about design itself. In the days of SketchUp, I did a lot of design work, and I knew the pain of SketchUp. We have to modify the model to make it clean, and we even can learn a lot about the model editing secrets. When the design is changed repeatedly, the design concept step by step to achieve and to perfect, really can feel the joy of creation. (The lumbar disc protrusion is so come!)
#
# This design took more than half of a year to elaborate. Because it also took a month to study the Song Dynasty architecture-related monographs, such as '营造法式 /building mode,' '梁思成全集6-7卷 /Liang Sicheng Complete Works 6-7 volumes', and a Japnese garden treatise from the parallel period of the Song Dynasty, '作庭记 /garden design.' We studied it all over and investigated the relevant legacy Tang and Song buildings and imitation. However, I left the planning institute to finish a Ph.D. and engage in digital research. Much of what happened later was lost. I only know that this project was rated as an excellent project in our institute. Of course, we received the design fee(Things designers have to relate to).
# +
def imgs_slideShow(imgs_fp,suffix=r'jpg',scale=0.7):
from PIL import Image
import glob
import numpy as np
from tqdm.auto import tqdm
import matplotlib.pyplot as plt
import matplotlib.animation as animation
'''
function - To dynamically show the images in a specified folder.
'''
imgs_f=glob.glob(imgs_fp+'/*.'+suffix)
#print(imgs_f)
imgs=[Image.open(i) for i in imgs_f]
img_resize=lambda img:img.resize([int(scale * s) for s in img.size] )
imgs_resize=[img_resize(i) for i in imgs]
fig=plt.figure(figsize=(20,10))
ims=[[plt.imshow(f,animated=True,)] for f in imgs_resize]
anima=animation.ArtistAnimation(fig,ims,interval=1000, blit=True,repeat_delay=1000)
return anima,ims
ancient_buildings_fp=r'.\imgs\ancientArhi_perspective'
anima,ims=imgs_slideShow(ancient_buildings_fp)
from IPython.display import HTML
HTML(anima.to_html5_video())
#anima.save('./video/g_antiqueArchi.mp4',fps=1)
# -
import utils
imgs_fn=utils.filePath_extraction(r'.\imgs\ancientArchi_section',["jpg"])
imgs_root=list(imgs_fn.keys())[0]
imgsFn_lst=imgs_fn[imgs_root]
columns=1
scale=1
utils.imgs_layoutShow(imgs_root,imgsFn_lst,columns,scale,figsize=(20,20))
# ### 1.2 In the era of parametric design
#
# > *Designers have gone from being tool users to tool makers.*
#
# The manual way is time-consuming and laborious, especially in constant adjustment and modification. Built half or the whole design of the model, at any time to modify, and the original model does not have the flexibility to adjust, so usually need to deliberate while rebuilding. Parametric design improves the above environment to a large extent. Although it takes a lot of effort to build formal logic initially, it can flexibly implement multiple variation structures of different sizes under the same design logic once it is completed.
#
# The various DouGong parts of ancient buildings in the Song Dynasty and the DouGong constructed by parts are compiled here. We can observe the construction relationship between elements and some antique buildings structure, which is also the parametric model construction logic. Later, some scholars specialized in studying ancient buildings' parametric systems, forming more rich and detailed results.
import glob
import utils
imgs_dougong_fps=glob.glob(r'.\code\12_DouGong(Qing)\*.jpg')
columns=3;scale=1
utils.imgs_layoutShow_FPList(imgs_dougong_fps,columns,scale,figsize=(20,20))
import glob
import utils
imgs_dougong_fps=glob.glob(r'.\code\11_AncientArchi\*.jpg')
columns=3;scale=1
utils.imgs_layoutShow_FPList(imgs_dougong_fps,columns,scale,figsize=(20,20))
# Due to parameterization technology, which expands the possibility of design forms, there are occasional special-form buildings. Cooperate with the structural engineer, adjust the parameter relationship, carry out the preliminary structural calculation, and meet the structure's requirements. The construction/building structure simulated by the designer in the design stage is different from the structural engineer's professional simulation. Still, it makes the design structure reasonable and avoids the unreasonable design structure in the structural analysis stage, resulting in extensive modifications.
# +
def get_multipleFiletypes_fps_list(files_root,suffix=['jpg','png']):
import os
import glob
'''
function - Returns a list of file paths from a folder that specified multiple file types.
'''
exts=['*.{}'.format(i) for i in suffix]
file_paths=[f for ext in exts for f in glob.glob(os.path.join(files_root, ext))]
return file_paths
imgs_fish_root=r'.\code\44_architecture_fish'
imgs_fish_fps=get_multipleFiletypes_fps_list(files_root=imgs_fish_root,suffix=['jpg','png'])
columns=5;scale=1
utils.imgs_layoutShow_FPList(imgs_fish_fps,columns,scale,figsize=(20,20))
# -
# The parametric design contains rich content, which can be embedded in many directions. The following are several cases of solving different problems, which were studied around 2012, and can be quickly looked over. And there is a lot of information available on the internet.
import os
video_root=r'./video'
video_fps=get_multipleFiletypes_fps_list(files_root=video_root,suffix=['mp4'])
video_dic={os.path.splitext(os.path.basename(v))[-2]:v for v in video_fps}
print("files info:\n")
import pandas as pd
print(pd.DataFrame.from_dict(video_dic,orient='index'))
from IPython.display import Video
Video(video_dic['02_Box flattened'])
# Estimated Grasshopper extension Add-Ons in 2014 or so, there are more than 30 groups. Still, up to 434 groups, recent statistics are already a large number, which has covered each direction because more and more designers and scholars form all kinds of components(being libraries of GH) while doing design and research. Of course, many unannounced extensions are designed for use within the company to increase competitiveness; Or researchers whose academic work is not published.
#
# Here is a piece of a code snippet in the form of a Sankey diagram of the more than 400 extension libraries. Let us look at the main research directions in the parametric design field at present, which also references the research in this field. It is also easy to find relevant functional components, quickly locate the plug-in name, and download the application.
# +
def Sankey(material_yaml_path,title_text,html_savePath,font_size=10,height=7000,): #font_size=10,height=7000
import yaml,random,copy
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from collections import OrderedDict
cmaps=OrderedDict()
'''
function - 读取自定义的.yaml文件,建立桑基图(Sankey diagram)
Paras:
material_yaml_path - 数据文件路径
title_text - 桑基图标题
html_savePath - 桑基图保存为html文件路径
font_size=10 - 图表字体大小
height=7000 - 图表高度
'''
#initialize parameters
material_yaml_path=material_yaml_path
title_text=title_text
font_size=font_size
height=height
html_savePath=html_savePath
with open(material_yaml_path,encoding='utf-8') as file:
materials=yaml.safe_load(file)
cmaps['Sequential']=['Greys', 'Purples', 'Blues', 'Greens', 'Oranges', 'Reds','YlOrBr', 'YlOrRd', 'OrRd', 'PuRd', 'RdPu', 'BuPu','GnBu', 'PuBu', 'YlGnBu', 'PuBuGn', 'BuGn', 'YlGn']
classification=materials['mapping']['classification']
classification_level_1=classification.keys()
#classification_level_2=classification.values()
cmaps_sequential=list(cmaps.values())[0]
random.shuffle(cmaps_sequential)
classification_level_1_colorSequential={k:v for k,v in zip(classification_level_1,cmaps_sequential[:len(classification_level_1)])}
classification_level_1_color={key:list(plt.cm.get_cmap(v)(0))[:3]+[0.8] for key,v in classification_level_1_colorSequential.items()} #configure level_1 color
classification_level_2_color={}
for k,v in classification.items():
classification_level_2_color.update({i:list(plt.cm.get_cmap(classification_level_1_colorSequential[k])(c_v)[:3])+[0.8] for i,c_v in zip(v,np.linspace(0.1,0.9,len(v)))}) #configure leve_2 color
materials_data=pd.DataFrame([dic['data'] for dic in materials['materials'] if dic['data'][0]!=''],columns=['name','date','case','international','source'])
materials_data['color']=materials_data.source.apply(lambda row:{0:[31, 119, 180, 0.8],1:[255, 127, 14, 0.8],2:[44, 160, 44, 0.8],3:[214, 39, 40, 0.8],4:[148, 103, 189, 0.8],5:[140, 86, 75, 0.8]}[row])
flatten_lst=lambda lst: [m for n_lst in lst for m in flatten_lst(n_lst)] if type(lst) is list else [lst]
label_idx={v:idx for idx,v in enumerate(flatten_lst(list(classification.values()))+list(classification.keys())+materials_data.name.to_list())}
node_label=label_idx.keys()
classification_level_1_color_255={k:[round(i*255) for i in v[:3]]+[v[-1]] for k,v in classification_level_1_color.items()}
classification_level_2_color_255={k:[round(i*255) for i in v[:3]]+[v[-1]] for k,v in classification_level_2_color.items()}
classification_color=copy.deepcopy(classification_level_1_color_255)
classification_color.update(classification_level_2_color_255)
classification_color.update({row['name']:row['color'] for idx,row in materials_data[['name','color']].iterrows()})
node_color=[classification_color[label] for label in node_label]
node_color_rgba=["rgba(%d,%d,%d,%.1f)"%(r,g,b,a) for r,g,b,a in node_color]
link_data_=[]
for dic in materials['materials']:
if dic['data'][0]!='':
link_data_.append([(dic['data'][0],link) for link in dic['link']])
link_data=flatten_lst(link_data_)
link_classification=flatten_lst([[(i,k) for i in v] for k,v in classification.items()])
classification_sub=materials['mapping']['classification_sub']
link_classification_sub=flatten_lst([[(i,k) for i in v] for k,v in classification_sub.items()])
link_S_T=link_data+link_classification_sub #+link_classification
link_source=[label_idx[s] for s,t in link_S_T]
link_target=[label_idx[t] for s,t in link_S_T]
link_num=len(link_S_T)
link_value=[1]*link_num #further adjust
opacity = 0.9
link_color_rgba=[node_color_rgba[src].replace("0.8", str(opacity)) for src in link_target]
link_label=[""]*link_num
import plotly.graph_objects as go
fig = go.Figure(data=[go.Sankey(
valueformat = ".0f",
valuesuffix = "TWh",
# Define nodes
node = dict(
pad = 20,
thickness = 15,
line = dict(color = "black", width = 0.5),
label=list(label_idx.keys()),
color=node_color_rgba
),
#domain=dict(column= ),
# Add links
link = dict(
source=link_source,
target=link_target,
value=link_value,
label=link_label,
color=link_color_rgba
))])
fig.update_layout(title_text=title_text,font_size=font_size,height=height) #fig.update_layout(title_text="参数化研究内容关联 <a href=''>link</a>",font_size=10)
fig.show()
fig.write_html("%s"%html_savePath)
material_yaml_path=r'./data/materials.yaml'
title_text=r"Grasshopper App(add-on),) content correlation _Created on Sat Oct 3 21:18:36 2020; updated on Sun Jan 24 21:52:32 2021 @author:<NAME>"
font_size=10
height=7000
html_savePath=r"./html/Parameterized overview chart.html"
Sankey(material_yaml_path,title_text=title_text,html_savePath=html_savePath,font_size=font_size,height=height) #Sankey(material_yaml_path,html_savePath,font_size,height)
# -
# In the driverless city project, the last stage mainly focused on the analysis of space mode. This phase is more in planning/landscape, exploring data analysis and information exchange approaches, such as database, OSM data exchange, 3D building data exchange, model data writing, and later related to various kinds of intelligent analysis. Work is still going on. The project is uploaded to the GitHub repository, [driverlessCity_LIM] (https://github.com/richieBao/driverlessCity_LIM), you can download the developed components, as well as database files. You can experiment with it yourself or use it directly in your design project.
# +
import matplotlib.pyplot as plt
from PIL import Image
img_LIM_fp=r'./imgs/misc/LIM.png'
img_LIM=Image.open(img_LIM_fp)
img_LIM
# -
# ### 1.3 It is back to manual work.--VR
#
# Oculus Quest 2+Gravity sketch
from IPython.display import Video
Video('./video/vr-01.mp4')
# ## 2.Coding in the present and future
# Focusing solely on parameterization does not maximize the benefits of the Code Age. Although GH(Dynamo) programming has dramatically improved the environment in which design tools are used, some design ideas are still difficult to implement with existing components, which is why it is necessary to drill down to the code level. There are 137,000+198,826(python libraries+python packages)in Python. What does it mean? It represents that we have countless tools to use. If you can not write code, then this precious wealth of knowledge may not be with you; what a pity!
#
# The studies in this field are all in (https://richiebao.github.io/Urban-Spatial-Data-Analysis_python/#/) CH,和[Urban spatial data analysis method——python language implementation](https://richiebao.github.io/Urban-Spatial-Data-Analysis_python_EN/#/)EN, with an estimated 50 chapters. Since there is a lot of content, you can browse it by yourself. Here is just a method to obtain image object through object detection of deep learning and apply it to urban spatial feature analysis.
from IPython.display import Video
Video('./video/17_13.mp4')
Video('./video/segmentation_FCN_RESNET101_animation.mp4')
# What knowledge do we need to achieve intelligent analysis and design? The following code categorizes the existing chapters' knowledge points and analyzes the relationships between them in a network structure. With this network structure, we can realize that you can know what knowledge you need by looking up the connection edges to determine how to perform an analysis. However, the roughly written code implementation of the network structure, its structure diagram expression is not very clear, can be further improved
import pandas as pd
USDA_kp_fp=r'./data/urban spatial data analysis method knowledge points structure_CH_EN.xlsx'
USDA_kp=pd.read_excel(USDA_kp_fp,sheet_name='EN',header=[0,1],engine='openpyxl')
USDA_kp.columns = USDA_kp.columns.get_level_values(1)
USDA_kp
# +
def G_draw(G,layout='spring_layout',node_color=None,node_size=None,figsize=(30, 30),font_size=12):
import matplotlib
import matplotlib.pyplot as plt
import networkx as nx
'''
function - To show a networkx graph
'''
#解决中文显示问题
plt.rcParams['font.sans-serif'] = ['DengXian'] # 指定默认字体 'KaiTi','SimHei'
plt.rcParams['axes.unicode_minus'] = False # 解决保存图像是负号'-'显示为方块的问题
fig, ax = plt.subplots(figsize=figsize)
#nx.draw_shell(G, with_labels=True)
layout_dic={
'spring_layout':nx.spring_layout,
'random_layout':nx.random_layout,
'circular_layout':nx.circular_layout,
'kamada_kawai_layout':nx.kamada_kawai_layout,
'shell_layout':nx.shell_layout,
'spiral_layout':nx.spiral_layout,
}
pos=layout_dic[layout](G)
nx.draw(G_1,pos,with_labels=True,node_color=node_color,node_size=node_size,font_size=font_size) #nx.draw(G, pos, font_size=16, with_labels=False)
import plotly.graph_objects as go
import networkx as nx
G_1=nx.from_pandas_edgelist(USDA_kp, source='chapter', target='knowledge point',create_using=nx.DiGraph()) #,create_using=nx.DiGraph();G=G.to_directed()
#G_1=nx.from_pandas_edgelist(USDA_kp, '章节','知识点')
classi=['algebra_regression','statistics', 'linear algebra', 'calculus', 'data processing','data analysis', 'data visulization',
'computer vision', 'machine learning', 'deep learning', 'framwork_platform','data available', 'darabase', 'GIS', 'RS', 'examples']
USDA_kp['edges']=USDA_kp.apply(lambda row:[(row['knowledge point'],i) for i in classi if row[i]==1],axis=1)
import utils
edges=utils.flatten_lst(USDA_kp['edges'].tolist())
G_1.add_edges_from(edges)
G_1.add_nodes_from(pd.unique(USDA_kp['chapter']).tolist(),layer=0)
G_1.add_nodes_from(pd.unique(USDA_kp['knowledge point']).tolist(),layer=2)
G_1.add_nodes_from(classi,layer=3)
colors=[
"gold",
"violet",
"limegreen",
"darkorange",
]
layer_colors=[colors[data["layer"]] for v, data in G_1.nodes(data=True)]
connections_num=[(len(list(nx.edges(G_1,[n])))+2)*50 for n in list(G_1.nodes())]
print("_"*50)
# -
G_draw(G_1,font_size=11,node_color=layer_colors,node_size=connections_num,layout='spring_layout')
nx.write_gpickle(G_1,'./model/G_1.pkl') #nx.read_gpickle('./model/G_1.pkl')
import hvplot.networkx as hvnx
hvnx.draw_spring(G_1, node_color=layer_colors, font_size='10pt',with_labels=True,arrowstyle='->',arrowsize=0.1,width=2000, height=2000) #labels='club', font_size='10pt', node_color='club', cmap='Category10',
import pandas as pd
USDA_kp_fp=r'./data/urban spatial data analysis method knowledge points structure_CH_EN.xlsx'
USDA_kp=pd.read_excel(USDA_kp_fp,sheet_name='EN',header=[0,1],engine='openpyxl')
USDA_kp.columns = USDA_kp.columns.get_level_values(1)
classi=['algebra_regression','statistics', 'linear algebra', 'calculus', 'data processing','data analysis', 'data visulization',
'computer vision', 'machine learning', 'deep learning', 'framwork_platform','data available', 'darabase', 'GIS', 'RS', 'examples']
USDA_kp['edges']=USDA_kp.apply(lambda row:[(row['knowledge point'],i) for i in classi if row[i]==1],axis=1)
# +
import plotly.graph_objects as go
import matplotlib.pyplot as plt
from matplotlib import colors
import pandas as pd
import numpy as np
import itertools
import random
#colors = [colors.to_rgba(c) for c in plt.rcParams['axes.prop_cycle'].by_key()['color']]
chapter=pd.unique(USDA_kp['chapter']).tolist()
Kp=pd.unique(USDA_kp['knowledge point']).tolist()
classi=list(USDA_kp.columns)[2:-1]
nodes=chapter+Kp+classi
nodes_idx_mapping={v:i for i,v in enumerate(nodes)}
opacity=0.4
cmap=plt.get_cmap('gist_ncar') #'gnuplot'
list_itemReplace=lambda lst,idx,v:[lst[i] if i!=idx else v for i in range(len(lst)) ]
nodes_colors=['rgba'+str(tuple(list_itemReplace([int(j*255) for j in cmap(i)],3,opacity))) for i in np.linspace(0, 1, len(nodes))]
random.shuffle(nodes_colors)
chapter2Kp=list(USDA_kp[['chapter','knowledge point']].itertuples(index=False,name=None))
Kp_classi=list(itertools.chain(*USDA_kp['edges'].to_list()))
edges_mapping=chapter2Kp+Kp_classi
edge_source=[nodes_idx_mapping[i[0]] for i in edges_mapping]
edge_target=[nodes_idx_mapping[i[1]] for i in edges_mapping]
link_num=len(edge_source)
link_value=[1]*link_num #further adjust
opacity_edge=0.2
link_color_rgba=[nodes_colors[src].replace(str(opacity),str(opacity_edge)) for src in edge_target]
link_label=[""]*link_num
# +
fig = go.Figure(data=[go.Sankey(
valueformat = ".0f",
valuesuffix = "TWh",
# Define nodes
node = dict(
pad = 15,
thickness = 15,
line = dict(color = "black", width = 0.5),
label =nodes,
color =nodes_colors
),
# Add links
link = dict(
source =edge_source,
target =edge_target,
value=link_value,
label=link_label,
color =link_color_rgba
))])
title_text="Urban spatial data analysis method--knowledge points correlation"
font_size=13
height=2700
fig.update_layout(title_text=title_text,font_size=font_size,height=height)
fig.show()
html_savePath='./html/Urban spatial data analysis method--knowledge points correlation.html'
fig.write_html("%s"%html_savePath)
# -
# # It is a great opportunity to share with you some research in the field of digital design that may plant a seed in your mind that you may not notice until later.
#
# # **THX!**
| .ipynb_checkpoints/guide_to_digitalDesignOFknowledgeStruc_EN_torun-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import pandas as pd
import matplotlib.pyplot as plt
from sklearn import datasets
import numpy as np
from sklearn.model_selection import train_test_split
from sklearn.tree import DecisionTreeClassifier
from sklearn import tree
from sklearn.metrics import classification_report
# !pip install category_encoders
a=pd.read_csv('Company_Data.csv')
a
a.info()
a.describe()
import category_encoders as ce
encoder=ce.OrdinalEncoder(cols=['ShelveLoc','Urban', 'US'])
sales1=encoder.fit_transform(a)
# +
sales_val = []
for value in a["Sales"]:
if value<=7.49:
sales_val.append("low")
else:
sales_val.append("high")
sales1["sales_val"]= sales_val
# -
sales1
x=sales1.drop(['sales_val','Sales'],axis=1)
y=sales1['sales_val']
x
y
x_train,x_test,y_train,y_test=train_test_split(x,y,test_size=0.3,random_state=40)
# # building decision tree classifier using entropy criteria
model=DecisionTreeClassifier(criterion='entropy',max_depth=5)
model.fit(x_train,y_train)
preds = model.predict(x_test) # predicting on test data set
pd.Series(preds).value_counts()
preds
pd.crosstab(y_test,preds)
#accuracy
np.mean(preds==y_test)
(34+47)/(34+18+21+47)
print(classification_report(preds,y_test))
fig = plt.figure(figsize=(25,20))
fig = tree.plot_tree(model,
feature_names= ['CompPrice','Income','Advertising','Population','Price', 'ShelveLoc', 'Age', 'Education',
'Urban', 'US'], class_names= ['low', 'high'], filled=True)
plt.title('Decision tree using Entropy',fontsize=22)
# # Building Decision Tree Classifier (CART) using Gini Criteria
from sklearn.tree import DecisionTreeClassifier
model_gini=DecisionTreeClassifier(criterion='gini',max_depth=5)
model_gini.fit(x_train,y_train)
pred=model.predict(x_test)
np.mean(preds==y_test)
fig = plt.figure(figsize=(25,20))
fig = tree.plot_tree(model_gini,
feature_names= ['CompPrice','Income','Advertising','Population','Price', 'ShelveLoc', 'Age', 'Education',
'Urban', 'US'], class_names= ['low', 'high'], filled=True)
plt.title('Decision tree using Entropy',fontsize=22)
# +
# hence from the both of the tree's we can understand that the most immportant constraints of the given data are price ,advertising and comp price
| decision tree company_data.csv.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# source: https://www.kaggle.com/ejunichi/m5-simple-fe from https://www.kaggle.com/ejunichi/m5-three-shades-of-dark-darker-magic
# +
import sys
import os
import pathlib
import gc
import pandas as pd
pd.set_option('display.max_columns', 500)
# pd.set_option('display.max_rows', 500)
import numpy as np
import math
import random
import pickle
import time
import psutil
import warnings
# custom import
from sklearn.preprocessing import LabelEncoder
from multiprocessing import Pool # Multiprocess Runs
from sklearn.preprocessing import PowerTransformer
# warnings.filterwarnings('ignore')
# -
# # constant variables for helper functions
N_CORES = psutil.cpu_count() # Available CPU cores
print(f"N_CORES: {N_CORES}")
# # function nicely diplaying a head of Pandas DataFrame
# +
import IPython
def display(*dfs, head=True):
for df in dfs:
IPython.display.display(df.head() if head else df)
# -
# # function fixing random seeds
# +
def seed_everything(seed=0):
"""Sets seed to make all processes deterministic # type: int
"""
random.seed(seed)
np.random.seed(seed)
SEED = 42
seed_everything(SEED)
# -
# # function processing df in multiprocess
def run_df_in_multiprocess(func, t_split):
"""Process ds in Multiprocess
"""
num_cores = np.min([N_CORES,len(t_split)])
pool = Pool(num_cores)
df = pd.concat(pool.map(func, t_split), axis=1)
pool.close()
pool.join()
return df
# # other helper functions
# +
def get_memory_usage():
"""メモリ使用量を確認するためのシンプルな「メモリプロファイラ」
"""
return np.round(psutil.Process(os.getpid()).memory_info()[0]/2.**30, 2)
def sizeof_fmt(num, suffix='B'):
for unit in ['','Ki','Mi','Gi','Ti','Pi','Ei','Zi']:
if abs(num) < 1024.0:
return "%3.1f%s%s" % (num, unit, suffix)
num /= 1024.0
return "%.1f%s%s" % (num, 'Yi', suffix)
def merge_by_concat(df1, df2, merge_on):
"""
dtypesを失わないための連結による結合
"""
merged_gf = df1[merge_on]
merged_gf = merged_gf.merge(df2, on=merge_on, how='left')
new_columns = [col for col in list(merged_gf) if col not in merge_on]
df1 = pd.concat([df1, merged_gf[new_columns]], axis=1)
return df1
# -
# # constant variables for data import
# +
# _DATA_DIR = os.path.sep.join(["data", "M5_Three_shades_of_Dark_Darker_magic", "sample"])
# _DATA_DIR = os.path.sep.join(["data", "M5_Three_shades_of_Dark_Darker_magic"])
_DATA_DIR = os.path.sep.join(["data", "M5_Three_shades_of_Dark_Darker_magic"])
_CALENDAR_CSV_FILE = "calendar.csv"
_SAMPLE_SUBMISSION_CSV_FILE = "sample_submission.csv"
# _SALES_TRAIN_VALIDATION_CSV_FILE = "sales_train_validation.csv"
_SALES_TRAIN_EVALUATION_CSV_FILE = "sales_train_evaluation.csv"
_SELL_PRICES_CSV_FILE = "sell_prices.csv"
# S3より取得済み。
_IS_RUN_ON_SAGEMAKER = False
# -
# # downlaod data (only on sagemaker)
# +
if _IS_RUN_ON_SAGEMAKER:
import sagemaker
# print(sagemaker.s3.parse_s3_url('s3://sagemaker-m5-forecasting-okada/accuracy/original/calendar.csv'))
parent_dir = pathlib.Path(os.path.abspath(os.curdir)).parent.parent
local_path = os.path.sep.join([str(parent_dir), _DATA_DIR])
print(local_path)
def import_one_object_from_s3(s3_uri, local_path):
sagemaker.s3.S3Downloader.download(
s3_uri=s3_uri,
local_path=local_path
)
# !ls $local_path
calendar_data_s3_uri = 's3://sagemaker-m5-forecasting-okada/accuracy/original/calendar.csv'
sales_train_evaluation_data_s3_uri = 's3://sagemaker-m5-forecasting-okada/accuracy/original/sales_train_evaluation.csv'
sales_train_validation_data_s3_uri = 's3://sagemaker-m5-forecasting-okada/accuracy/original/sales_train_validation.csv'
sample_submission_data_s3_uri = 's3://sagemaker-m5-forecasting-okada/accuracy/original/sample_submission.csv'
sell_prices_data_s3_uri = 's3://sagemaker-m5-forecasting-okada/accuracy/original/sell_prices.csv'
s3_uris = [
calendar_data_s3_uri,
sales_train_evaluation_data_s3_uri,
sales_train_validation_data_s3_uri,
sample_submission_data_s3_uri,
sell_prices_data_s3_uri
]
for s3_uri in s3_uris:
import_one_object_from_s3(s3_uri, local_path)
# -
# # import data
# +
def reduce_mem_usage(df, verbose=True):
"""
reduce the memory usage of the given dataframe.
https://qiita.com/hiroyuki_kageyama/items/02865616811022f79754
Args:
df: Dataframe
verbose:
Returns:
df, whose memory usage is reduced.
Raises:
None
"""
numerics = ['int16', 'int32', 'int64', 'float16', 'float32', 'float64']
start_mem = df.memory_usage().sum() / 1024**2
for col in df.columns: #columns毎に処理
col_type = df[col].dtypes
if col_type in numerics: #numericsのデータ型の範囲内のときに処理を実行. データの最大最小値を元にデータ型を効率的なものに変更
c_min = df[col].min()
c_max = df[col].max()
if str(col_type)[:3] == 'int':
if c_min > np.iinfo(np.int8).min and c_max < np.iinfo(np.int8).max:
df[col] = df[col].astype(np.int8)
elif c_min > np.iinfo(np.int16).min and c_max < np.iinfo(np.int16).max:
df[col] = df[col].astype(np.int16)
elif c_min > np.iinfo(np.int32).min and c_max < np.iinfo(np.int32).max:
df[col] = df[col].astype(np.int32)
elif c_min > np.iinfo(np.int64).min and c_max < np.iinfo(np.int64).max:
df[col] = df[col].astype(np.int64)
else:
if c_min > np.finfo(np.float16).min and c_max < np.finfo(np.float16).max:
df[col] = df[col].astype(np.float16)
elif c_min > np.finfo(np.float32).min and c_max < np.finfo(np.float32).max:
df[col] = df[col].astype(np.float32)
else:
df[col] = df[col].astype(np.float64)
end_mem = df.memory_usage().sum() / 1024**2
if verbose: print('Mem. usage decreased to {:5.2f} Mb ({:.1f}% reduction)'.format(end_mem, 100 * (start_mem - end_mem) / start_mem))
return df
def read_csv_data(directory, file_name):
print('Reading files...')
df = pd.read_csv(os.path.sep.join([str(directory), _DATA_DIR, file_name]))
df = reduce_mem_usage(df)
print('{} has {} rows and {} columns'.format(file_name, df.shape[0], df.shape[1]))
return df
# -
# # read csv data
# +
parent_dir = pathlib.Path(os.path.abspath(os.curdir)).parent.parent
print(f"parent_dir: {parent_dir}")
df_sales_train_evaluation = read_csv_data(parent_dir, _SALES_TRAIN_EVALUATION_CSV_FILE)
# -
df_sell_prices = read_csv_data(parent_dir, _SELL_PRICES_CSV_FILE)
df_calendar = read_csv_data(parent_dir, _CALENDAR_CSV_FILE)
df_sample_submission = read_csv_data(parent_dir, _SAMPLE_SUBMISSION_CSV_FILE)
print(f"df_sales_train_evaluation: {df_sales_train_evaluation.head()}")
print(f"df_sample_submission: {df_sample_submission.head()}")
# # constant variables for preprocessing/prediction
# +
# 予測期間とitem数の定義 / number of items, and number of prediction period
_NUM_UNIQUE_ITEM_ID = df_sales_train_evaluation.shape[0] # 30490
print(f"_NUM_UNIQUE_ITEM_ID: {_NUM_UNIQUE_ITEM_ID}")
_DAYS_FOR_PREDICTION = df_sample_submission.shape[1] - 1 # 28
print(f"_DAYS_FOR_PREDICTION: {_DAYS_FOR_PREDICTION}")
# DAYS_PER_YEAR = 365
# _NUM_YEARS_FOR_MELT = 2
# _NUM_IMPORT_ROWS_FOR_MELT = DAYS_PER_YEAR * _NUM_YEARS_FOR_MELT * _NUM_UNIQUE_ITEM_ID
# print(f"_NUM_IMPORT_ROWS_FOR_MELT: {_NUM_IMPORT_ROWS_FOR_MELT}")
_SALES_HISTORY_DAYS = 1913
_SALES_HISTORY_START_DAYS_FOR_VALIDATION = _SALES_HISTORY_DAYS + 1
_SALES_HISTORY_START_DAYS_FOR_EVALUATION = _SALES_HISTORY_START_DAYS_FOR_VALIDATION + _DAYS_FOR_PREDICTION
print(f"_SALES_HISTORY_START_DAYS_FOR_EVALUATION: {_SALES_HISTORY_START_DAYS_FOR_EVALUATION}")
TARGET = 'sales'
MAIN_INDEX = ['id','d'] # We can identify items by these columns
# -
# # Create Grid from df_sales_train_evaluation by melting
# +
# We can tranform horizontal representation to vertical "view"
# Our "index" will be 'id','item_id','dept_id','cat_id','store_id','state_id' and labels are 'd_' coulmns
index_columns = ['id','item_id','dept_id','cat_id','store_id','state_id']
grid_df = pd.melt(df_sales_train_evaluation,
id_vars = index_columns,
var_name = 'd',
value_name = TARGET)
print(f"Train shapes: {df_sales_train_evaluation.shape}, {grid_df.shape}")
print(f"grid_df: {grid_df}")
# -
# # テスト結果を格納するためのRowを追加
# +
# add the test result receiver column to grid_df
df_test_result_receiver = pd.DataFrame()
for i in range(0,_DAYS_FOR_PREDICTION):
temp_df = df_sales_train_evaluation[index_columns]
temp_df = temp_df.drop_duplicates() # snince "id" is unique, this is not necessary
temp_df['d'] = 'd_'+ str(_SALES_HISTORY_START_DAYS_FOR_EVALUATION + i)
temp_df[TARGET] = np.nan
df_test_result_receiver = pd.concat([df_test_result_receiver,temp_df])
print(f"df_test_result_receiver: {df_test_result_receiver}")
grid_df = pd.concat([grid_df, df_test_result_receiver])
grid_df = grid_df.reset_index(drop=True)
print(f"grid_df concatenated with df_test_result_receiver: {grid_df}")
# 一時的なDFを削除する
# We will not need original train_df anymore and can remove it
del temp_df, df_test_result_receiver
# del temp_df, df_test_result_receiver, df_sales_train_evaluation
gc.collect()
# check memory usage
print("{:>20}: {:>8}".format('Original grid_df',sizeof_fmt(grid_df.memory_usage(index=True).sum())))
# change the data type from string into category for memory reduction
for col in index_columns:
print(f"grid_df[col]: {grid_df[col]}")
grid_df[col] = grid_df[col].astype('category')
# check memory usage
print("{:>20}: {:>8}".format('Reduced grid_df',sizeof_fmt(grid_df.memory_usage(index=True).sum())))
# -
# # remove some 0 sale price rows which actually means that products do not exist yet
# +
# 各train_dfアイテム行の先行ゼロ値は実際の0売上ではなく、店にアイテムがないことを意味する。
# そのようなゼロを削除することで、一部のメモリを安全にする。
# 価格は州ごとに設定されるので、 リリース週があまり正確ではありません
# find the oldest release week (= smallest wm_yr_wk) of each item at each store
item_release_week_df = df_sell_prices.groupby(['store_id','item_id'])['wm_yr_wk'].agg(['min']).reset_index()
# display(df_sell_prices.groupby(['store_id','item_id'])['wm_yr_wk'].agg(['min']))
# display(df_sell_prices.groupby(['store_id','item_id']).get_group(("CA_1", "FOODS_1_001")))
# display(df_sell_prices.groupby(['store_id','item_id']).get_group(("CA_1", "FOODS_1_001")).agg(['min']))
# display(df_sell_prices.groupby(['store_id','item_id'])['wm_yr_wk'].get_group(("CA_1", "FOODS_1_001")))
# just change the column name from "min" to "release"
item_release_week_df.columns = ['store_id','item_id','release']
print(f"item_release_week_df: {item_release_week_df}")
# concat with grid_d
grid_df = merge_by_concat(grid_df, item_release_week_df, ['store_id','item_id'])
del item_release_week_df
gc.collect()
print(f"grid_df after concatenating with item_release_week_df: {grid_df}")
# grid_dfから「ゼロ」行をいくつか削除したい
# それを行うには、wm_yr_wk列が必要
# 部分的にcalendar_dfを結合
grid_df = merge_by_concat(grid_df, df_calendar[['wm_yr_wk','d']], ['d'])
print(f"grid_df after concatenating with df_calendar: {grid_df}")
# +
# これで、いくつかの行をカットして安全なメモリにできます
# remove the rows whose release week is earlier than 'wm_yr_wk' when products should not be started to be sold.
grid_df = grid_df[grid_df['wm_yr_wk'] >= grid_df['release']]
grid_df = grid_df.reset_index(drop=True)
# メモリ使用量を確認しましょう
print("{:>20}: {:>8}".format('Original grid_df',sizeof_fmt(grid_df.memory_usage(index=True).sum())))
print(f"grid_df after removing uneccesary rows: {grid_df}")
# +
# 特徴量の1つとしてリリース週を維持する必要がありますか?
# 良いCVだけが答えを出すことができます。
# リリース値を縮小してみましょう。
# 最小変換はここでは役に立たない
# int16→integer(-32768から32767) grid_df ['release'].max()→int16のような変換は。
# しかし、必要な場合に備えて、変換するある方法があります。
grid_df['release'] = grid_df['release'] - grid_df['release'].min()
grid_df['release'] = grid_df['release'].astype(np.int16)
# メモリ使用量をもう一度確認してみましょう
print("{:>20}: {:>8}".format('Reduced grid_df',sizeof_fmt(grid_df.memory_usage(index=True).sum())))
print(f"grid_df:{grid_df}")
# -
# # export the base grid (grid_part_1)
# +
# save the base grid file as pickle for later usage.
_EXPORT_FILE_NAME = "base_grid_for_darker_magic_evaluation.pkl"
print("data export start")
grid_df.to_pickle(os.path.sep.join([str(parent_dir), _DATA_DIR, _EXPORT_FILE_NAME]))
print('data export finished. Size:', grid_df.shape)
# -
# # ----- you can split the notebook here -----
# # preprocessing df_sell_prices: add features
display(df_sell_prices.head())
# +
# 基本的な集計を行う
# the max, min, std, mean price among the same item at the same store
df_sell_prices['price_max'] = df_sell_prices.groupby(['store_id','item_id'])['sell_price'].transform('max')
df_sell_prices['price_min'] = df_sell_prices.groupby(['store_id','item_id'])['sell_price'].transform('min')
df_sell_prices['price_std'] = df_sell_prices.groupby(['store_id','item_id'])['sell_price'].transform('std')
df_sell_prices['price_mean'] = df_sell_prices.groupby(['store_id','item_id'])['sell_price'].transform('mean')
print(f"df_sell_prices['price_max']: {df_sell_prices['price_max']}")
# 価格正規化を行う(min-max scaling)。priceのレンジは大きくないので、Logを行う意味なし。(むしろ、一ドル以下の商品が多いので悪い)
df_sell_prices['price_norm'] = df_sell_prices['sell_price']/df_sell_prices['price_max']
print(f"df_sell_prices['price_norm']: {df_sell_prices['price_norm']}")
print(f"df_sell_prices['price_norm'].shape: {df_sell_prices['price_norm'].shape}")
# # since the price distribution is not very skewed or long-tailed, I decided not to take log scale. see: https://www.kaggle.com/headsortails/back-to-predict-the-future-interactive-m5-eda
# df_sell_prices['price_norm'] = np.log1p(df_sell_prices['sell_price'])
# print(f"log1p df_sell_prices['price_norm']: {df_sell_prices['price_norm']}")
# print(f"log1p df_sell_prices['price_norm'].shape: {df_sell_prices['price_norm'].shape}")
# since "pt.fit(sell_price_2d_array)" caused "ValueError: Input contains infinity or a value too large for dtype('float16').", box-cox transformation was discarded: from scipy import stats also did not make propcer transform.
# # https://note.com/mikiokubo/n/n42417e5d0f6c
# # https://gakushukun1.hatenablog.com/entry/2019/04/29/112424
# pt = PowerTransformer(method="box-cox")
# sell_price_2d_array = df_sell_prices['sell_price'].values.reshape(-1,1)
# # data = df["B"].values
# print(f"sell_price_2d_array: {sell_price_2d_array}")
# pt.fit(sell_price_2d_array)
# df_sell_prices['price_norm'] = pt.transform(sell_price_2d_array)
# df_sell_prices['price_norm'].hist()
# print(f"box-coxed df_sell_prices['price_norm']: {df_sell_prices['price_norm']}")
# print(f"box-coxed df_sell_prices['price_norm'].shape: {df_sell_prices['price_norm'].shape}")
# 一部のアイテムはインフレに依存する可能性があります。いくつかのアイテムは非常に「安定」しています
# count the kinds of sell_price per ['store_id','item_id']. the smaller, the more stable the price is.
df_sell_prices['price_nunique'] = df_sell_prices.groupby(['store_id','item_id'])['sell_price'].transform('nunique')
# count the kinds of item_id per ['store_id','sell_price']. the smaller, the less frequent the price is, whether the price is large or small. todo@kensakuokada: this feature may not be necessary.
df_sell_prices['item_nunique'] = df_sell_prices.groupby(['store_id','sell_price'])['item_id'].transform('nunique')
# I would like some "rolling" aggregations but would like months and years as "window"
calendar_prices = df_calendar[['wm_yr_wk','month','year']]
# the rows are duplicated in each week. reduce rows per week
calendar_prices = calendar_prices.drop_duplicates(subset=['wm_yr_wk'])
# add month and year
df_sell_prices = df_sell_prices.merge(calendar_prices[['wm_yr_wk','month','year']], on=['wm_yr_wk'], how='left')
del calendar_prices
gc.collect()
display(df_sell_prices.head())
# +
# 週ごとにシフト in each (['store_id','item_id']) group. todo@kensakuokada: add more momentum by shifting more, which may be better.
df_sell_prices['price_momentum'] = df_sell_prices['sell_price']/df_sell_prices.groupby(['store_id','item_id'])['sell_price'].transform(lambda x: x.shift(1))
# each sell price/月平均 in ['store_id','item_id','month'] group
df_sell_prices['price_momentum_m'] = df_sell_prices['sell_price']/df_sell_prices.groupby(['store_id','item_id','month'])['sell_price'].transform('mean')
# each sell price/年平均 in ['store_id','item_id','year'] group
df_sell_prices['price_momentum_y'] = df_sell_prices['sell_price']/df_sell_prices.groupby(['store_id','item_id','year'])['sell_price'].transform('mean')
del df_sell_prices['month'], df_sell_prices['year'] # todo@kensakuokada: try not to remove month and year, which may be better.
gc.collect()
display(df_sell_prices.head())
# -
# # preprocessing df_sell_prices: concat df_sell_prices with MAIN_INDEX (connecting with other dataframes)
# +
# base columns
original_columns = list(grid_df)
print(f"original_columns: {original_columns}")
grid_df = grid_df.merge(df_sell_prices, on=['store_id','item_id','wm_yr_wk'], how='left')
print(f"grid_df: {grid_df}")
keep_columns = [col for col in list(grid_df) if col not in original_columns]
print(f"keep_columns: {keep_columns}")
grid_df = grid_df[MAIN_INDEX+keep_columns]
grid_df = reduce_mem_usage(grid_df)
print(f"grid_df: {grid_df}")
# We don't need prices_df anymore
del df_sell_prices
gc.collect()
# -
# # export grid_df having sales_price features and MAIN_INDEX (grid_part_2)
# 今後のモデルトレーニングのためpickleファイルとして保存
_EXPORT_FILE_NAME = "base_grid_with_sales_price_features_for_darker_magic_evaluation.pkl"
print("data export start")
grid_df.to_pickle(os.path.sep.join([str(parent_dir), _DATA_DIR, _EXPORT_FILE_NAME]))
print('data export finished. Size:', grid_df.shape)
# # ----- you can split the notebook here -----
# # preprocessing df_calendar
# +
grid_df = grid_df[MAIN_INDEX]
# カレンダーを部分的に結合
icols = ['date',
'd',
'event_name_1',
'event_type_1',
'event_name_2',
'event_type_2',
'snap_CA',
'snap_TX',
'snap_WI']
grid_df = grid_df.merge(df_calendar[icols], on=['d'], how='left')
# データを縮小する
# 'snap_'列はboolまたはint8に変換できる
icols = ['event_name_1',
'event_type_1',
'event_name_2',
'event_type_2',
'snap_CA',
'snap_TX',
'snap_WI']
for col in icols:
grid_df[col] = grid_df[col].astype('category')
# 日時に変換
grid_df['date'] = pd.to_datetime(grid_df['date'])
# 日付からいくつかの特徴量を作る
grid_df['tm_d'] = grid_df['date'].dt.day.astype(np.int8)
grid_df['tm_w'] = grid_df['date'].dt.week.astype(np.int8)
grid_df['tm_m'] = grid_df['date'].dt.month.astype(np.int8)
grid_df['tm_y'] = grid_df['date'].dt.year
grid_df['tm_y'] = (grid_df['tm_y'] - grid_df['tm_y'].min()).astype(np.int8)
grid_df['tm_wm'] = grid_df['tm_d'].apply(lambda x: math.ceil(x/7)).astype(np.int8)
grid_df['tm_dw'] = grid_df['date'].dt.dayofweek.astype(np.int8)
grid_df['tm_w_end'] = (grid_df['tm_dw']>=5).astype(np.int8)
print(f"grid_df: {grid_df}")
# -
# 日付の削除
del grid_df['date']
grid_df.info()
# # export grid_df having sales_price features and MAIN_INDEX (grid_part_3)
_EXPORT_FILE_NAME = "base_grid_with_calendar_features_for_darker_magic_evaluation.pkl"
print("data export start")
grid_df.to_pickle(os.path.sep.join([str(parent_dir), _DATA_DIR, _EXPORT_FILE_NAME]))
print('data export finished. Size:', grid_df.shape)
| accuracy/M5_Three_shades_of_Dark_Darker_magic/preprocessing_for_base_grid_having_item_properties_and_sales_prices_evaluation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Arithmetic operators
# Combine 15 and 23.
15 + 23
# Subtract 50 from 26.
# 50 - 26
# Divide 20 by 4.
20/4
# Divide 22 by 4.
22/4
# Obtain the remainder of the division of 22 by 4.
22 % 4
# Divide the float 22 by 4.
22.0 / 4
# Multiply 6 by 8.
6 * 8
# Raise 15 to the power of 2.
15 ** 2
# source:
# https://www.udemy.com/course/the-data-science-course-complete-data-science-bootcamp/
| Arithmetic Operators - Exercise_Py3.ipynb |