code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + active=""
# Imports:
# -
import pickle
import cv2
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
from glob import glob
from os.path import join, exists, splitext, split
from statistics import mean
# Import everything needed to edit/save/watch video clips
from moviepy.editor import VideoFileClip
from IPython.display import HTML
# + active=""
# Constants:
# -
nx = 9 #number of inside corners in x
ny = 6 #number of inside corners in y
# Define conversions in x and y from pixels space to meters
ym_per_pix = 30/720 # meters per pixel in y dimension
xm_per_pix = 3.7/700 # meters per pixel in x dimension
# + active=""
# Help functions:
# +
def configure_calibration(calibration_images_dir):
'''
Get main path for images to caliprate camera with as input then save a pickle file
with the configuration to be used in undistortion later
'''
#if not exists('calibration_conf.p'):
# For every calibration image, get object points and image points by finding chessboard corners.
objpoints = [] # 3D points in real world space.
imgpoints = [] # 2D points in image space.
# Prepare constant object points, like (0,0,0), (1,0,0), (2,0,0) ....,(9,6,0).
objpoints_const = np.zeros((nx * ny, 3), np.float32)
objpoints_const[:, :2] = np.mgrid[0:nx, 0:ny].T.reshape(-1, 2)
images = glob(join(calibration_images_dir, '*.jpg'))
for img_path in images:
img = cv2.imread(img_path)
# Convert to grayscale
gray = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
# Find the chessboard corners
ret, corners = cv2.findChessboardCorners(gray, (nx, ny), None)
if ret == True:
# append found corners to imgpoints & prepared constants object points to objpoints for mapping
objpoints.append(objpoints_const)
imgpoints.append(corners)
#use all point got from images for calibration
ret, mtx, dist, rvecs, tvecs = cv2.calibrateCamera(objpoints, imgpoints, gray.shape[::-1], None, None)
# Save to pickle file
pickle.dump({'mtx': mtx, 'dist': dist, 'corners': corners}, open('calibration_conf.p', 'wb'))
def undistort_image(img, calibration_images_dir = './camera_cal/'):
'''
Get calibration configuration from config file to undistort images
Takes image as input then
Returns undistorted image
'''
if not exists('calibration_conf.p'):
configure_calibration(calibration_images_dir)
# Return pickled calibration data.
pickle_dict = pickle.load(open('calibration_conf.p', 'rb'))
mtx = pickle_dict['mtx']
dist = pickle_dict['dist']
corners = pickle_dict['corners']
# return undistorted image
undist = cv2.undistort(img, mtx, dist, None, mtx)
return undist, corners
def abs_sobel_thresh(img, sobel_thresh=(0, 255)):
# Apply the following steps to img
# Convert to grayscale
gray = cv2.cvtColor(img, cv2.COLOR_RGB2GRAY)
# Take the derivative in x & take the absolute value of the result
abs_sobel = np.absolute(cv2.Sobel(gray, cv2.CV_64F, 1, 0))
# Scale to 8-bit (0 - 255) then convert to type = np.uint8
scaled_sobel = np.uint8(255*abs_sobel/np.max(abs_sobel))
# Create a mask of 1's where the scaled gradient magnitude
# is > thresh_min and < thresh_max
binary_output = np.zeros_like(scaled_sobel)
# Return this mask as your binary_output image
binary_output[(scaled_sobel > sobel_thresh[0]) & (scaled_sobel < sobel_thresh[1])] = 1
return binary_output
def dir_threshold(img, sobel_kernel=3, dir_thresh=(0, np.pi/2)):
# Grayscale
gray = cv2.cvtColor(img, cv2.COLOR_RGB2GRAY)
# Calculate the x and y gradients
sobelx = cv2.Sobel(gray, cv2.CV_64F, 1, 0, ksize=sobel_kernel)
sobely = cv2.Sobel(gray, cv2.CV_64F, 0, 1, ksize=sobel_kernel)
# Take the absolute value of the gradient direction,
# apply a threshold, and create a binary image result
absgraddir = np.arctan2(np.absolute(sobely), np.absolute(sobelx))
binary_output = np.zeros_like(absgraddir)
binary_output[(absgraddir >= dir_thresh[0]) & (absgraddir <= dir_thresh[1])] = 1
# Return the binary image
return binary_output
def hls_select(img, s_thresh=(0, 255), l_thresh=(0, 255)):
# Get hls of the image
hls = cv2.cvtColor(img, cv2.COLOR_RGB2HLS)
# Get the s & l channels to apply thresholds
s_channel = hls[:,:,2]
l_channel = hls[:,:,1]
# Get binary output of s_channel applying thresholds
binary_output_s = np.zeros_like(s_channel)
binary_output_s[(s_channel > s_thresh[0]) & (s_channel <= s_thresh[1])] = 1
# Get binary output of l_channel applying thresholds
binary_output_l = np.zeros_like(l_channel)
binary_output_l[(l_channel > l_thresh[0]) & (l_channel <= l_thresh[1])] = 1
# Combine the thresholds of both S & L
combined_binary = np.zeros_like(l_channel)
combined_binary[(binary_output_l == 1) & (binary_output_s == 1)] = 1
return combined_binary
def birdeye(undist, corners, inverse = False):
'''
Get undistort image, corners got from findChessboardCorners as input
Returns birdeye image for this image
'''
# Grab the image shape
img_size = (undist.shape[1], undist.shape[0])
# The lower points should be as close to the lower edge of the image as possible.
# The length of the road in the selected area should be around 30m.
src = np.float32([(250, 680), (1050, 680), (600, 470), (730, 470)])
# For destination points, I'm arbitrarily choosing some points to be
# a nice fit for displaying our warped result
# again, not exact, but close enough to make lines appear parraled
dst = np.float32([(280, 720), (1000, 720), (280, 0), (1000, 0)])
# Given src and dst points, calculate the perspective transform matrix
if inverse:
M = cv2.getPerspectiveTransform(dst, src)
else:
M = cv2.getPerspectiveTransform(src, dst)
# Warp the image using OpenCV warpPerspective()
warped = cv2.warpPerspective(undist, M, img_size)
return warped
def combined_thresholds(img, sobel_kernel = 3, sobel_thresh=(0, 255), dir_thresh=(0, np.pi/2), s_thresh=(0, 255), l_thresh=(0, 255)):
'''
Takes warped image with thresholds as input,
Calculates the drivative in x direction then returns the result
'''
# Convert to HLS color space and separate the S channel
# Note: img is the undistorted image
hls = cv2.cvtColor(img, cv2.COLOR_RGB2HLS)
s_channel = hls[:,:,2]
# Sobel filtering in x direction with thresholds to get better view of the edges
sobel_binary = abs_sobel_thresh(img, sobel_thresh)
# Get the direction thresholds between 40 to 90 degrees as lane lines are nearly vertical
dir_binary = dir_threshold(img, sobel_kernel, dir_thresh)
# Get the S & L saturation of the image thersholded for better color saturation & to remove shadows
sl_binary = hls_select(img, s_thresh, l_thresh)
# Combine the two binary thresholds
combined_binary = np.zeros_like(sobel_binary)
combined_binary[(sobel_binary == 1) & (dir_binary == 1) | (sl_binary == 1) ] = 1
return combined_binary
def find_lane_pixels(binary_warped):
# Take a histogram of the bottom half of the image
histogram = np.sum(binary_warped[binary_warped.shape[0]//2:,:], axis=0)
# Create an output image to draw on and visualize the result
out_img = np.dstack((binary_warped, binary_warped, binary_warped))
# Find the peak of the left and right halves of the histogram
# These will be the starting point for the left and right lines
midpoint = np.int(histogram.shape[0]//2)
leftx_base = np.argmax(histogram[:midpoint])
rightx_base = np.argmax(histogram[midpoint:]) + midpoint
# HYPERPARAMETERS
# Choose the number of sliding windows
nwindows = 9
# Set the width of the windows +/- margin
margin = 100
# Set minimum number of pixels found to recenter window
minpix = 50
# Set height of windows - based on nwindows above and image shape
window_height = np.int(binary_warped.shape[0]//nwindows)
# Identify the x and y positions of all nonzero pixels in the image
nonzero = binary_warped.nonzero()
nonzeroy = np.array(nonzero[0])
nonzerox = np.array(nonzero[1])
# Current positions to be updated later for each window in nwindows
leftx_current = leftx_base
rightx_current = rightx_base
# Create empty lists to receive left and right lane pixel indices
left_lane_inds = []
right_lane_inds = []
# Step through the windows one by one
for window in range(nwindows):
# Identify window boundaries in x and y (and right and left)
win_y_low = binary_warped.shape[0] - (window+1)*window_height
win_y_high = binary_warped.shape[0] - window*window_height
win_xleft_low = leftx_current - margin
win_xleft_high = leftx_current + margin
win_xright_low = rightx_current - margin
win_xright_high = rightx_current + margin
# Draw the windows on the visualization image
cv2.rectangle(out_img,(win_xleft_low,win_y_low),
(win_xleft_high,win_y_high),(0,255,0), 2)
cv2.rectangle(out_img,(win_xright_low,win_y_low),
(win_xright_high,win_y_high),(0,255,0), 2)
# Identify the nonzero pixels in x and y within the window #
good_left_inds = ((nonzeroy >= win_y_low) & (nonzeroy < win_y_high) &
(nonzerox >= win_xleft_low) & (nonzerox < win_xleft_high)).nonzero()[0]
good_right_inds = ((nonzeroy >= win_y_low) & (nonzeroy < win_y_high) &
(nonzerox >= win_xright_low) & (nonzerox < win_xright_high)).nonzero()[0]
# Append these indices to the lists
left_lane_inds.append(good_left_inds)
right_lane_inds.append(good_right_inds)
# If you found > minpix pixels, recenter next window on their mean position
if len(good_left_inds) > minpix:
leftx_current = np.int(np.mean(nonzerox[good_left_inds]))
if len(good_right_inds) > minpix:
rightx_current = np.int(np.mean(nonzerox[good_right_inds]))
# Concatenate the arrays of indices (previously was a list of lists of pixels)
try:
left_lane_inds = np.concatenate(left_lane_inds)
right_lane_inds = np.concatenate(right_lane_inds)
except ValueError:
# Avoids an error if the above is not implemented fully
pass
# Extract left and right line pixel positions
leftx = nonzerox[left_lane_inds]
lefty = nonzeroy[left_lane_inds]
rightx = nonzerox[right_lane_inds]
righty = nonzeroy[right_lane_inds]
return leftx, lefty, rightx, righty, out_img
def fit_polynomial(binary_warped):
# Find our lane pixels first
leftx, lefty, rightx, righty, out_img = find_lane_pixels(binary_warped)
# Fit a second order polynomial to each using `np.polyfit`
left_fit = np.polyfit(lefty, leftx, 2)
right_fit = np.polyfit(righty, rightx, 2)
# Generate x and y values for plotting
ploty = np.linspace(0, binary_warped.shape[0]-1, binary_warped.shape[0] )
# Fit a second order polynomial to each using `np.polyfit` real world data
left_fit_cr = np.polyfit(lefty * ym_per_pix, leftx * xm_per_pix, 2)
right_fit_cr = np.polyfit(righty * ym_per_pix, rightx * xm_per_pix, 2)
try:
left_fitx = left_fit[0]*ploty**2 + left_fit[1]*ploty + left_fit[2]
right_fitx = right_fit[0]*ploty**2 + right_fit[1]*ploty + right_fit[2]
except TypeError:
# Avoids an error if `left` and `right_fit` are still none or incorrect
print('The function failed to fit a line!')
left_fitx = 1*ploty**2 + 1*ploty
right_fitx = 1*ploty**2 + 1*ploty
## Visualization ##
# Colors in the left and right lane regions
out_img[lefty, leftx] = [255, 0, 0]
out_img[righty, rightx] = [0, 0, 255]
return out_img, left_fitx, right_fitx, ploty, left_fit_cr, right_fit_cr
def measure_curvature_pixels(left_fit_cr, right_fit_cr, ploty):
'''
Calculates the curvature of polynomial functions in pixels.
'''
# Define y-value where we want radius of curvature
# We'll choose the maximum y-value, corresponding to the bottom of the image
y_eval = np.max(ploty)
left_curverad = ((1 + (2*left_fit_cr[0]*y_eval*ym_per_pix + left_fit_cr[1])**2)**1.5) / np.absolute(2*left_fit_cr[0])
right_curverad = ((1 + (2*right_fit_cr[0]*y_eval*ym_per_pix + right_fit_cr[1])**2)**1.5) / np.absolute(2*right_fit_cr[0])
return left_curverad, right_curverad
# + active=""
# Process Image:
# +
def process_image(img):
undistort, corners = undistort_image(img, calibration_images_dir = './camera_cal/')
thresholded = combined_thresholds(img, sobel_kernel = 7, sobel_thresh=(10, 100), dir_thresh=(0.69, 1.3), s_thresh=(90, 255), l_thresh=(140, 255))
#thresholded = hls_select(undistort, s_thresh=(90, 255), l_thresh=(140, 255))
warped = birdeye(thresholded, corners)
out_img, left_fitx, right_fitx, ploty, left_fit_cr, right_fit_cr = fit_polynomial(warped)
left_curverad, right_curverad = measure_curvature_pixels(left_fit_cr, right_fit_cr, ploty)
#f, (ax1, ax2) = plt.subplots(1, 2, figsize=(24, 9))
#f.tight_layout()
#ax1.imshow(undistort)
#ax1.set_title('Original Image', fontsize=50)
#ax2.imshow(warped)
#ax2.set_title('Thresholded Grad. Dir.', fontsize=50)
#plt.subplots_adjust(left=0., right=1, top=0.9, bottom=0.)
#plt.imshow(warped)
#print(left_curverad, right_curverad)
# Create an image to draw the lines on
warp_zero = np.zeros_like(warped).astype(np.uint8)
color_warp = np.dstack((warp_zero, warp_zero, warp_zero))
# Recast the x and y points into usable format for cv2.fillPoly()
pts_left = np.array([np.transpose(np.vstack([left_fitx, ploty]))])
pts_right = np.array([np.flipud(np.transpose(np.vstack([right_fitx, ploty])))])
pts = np.hstack((pts_left, pts_right))
# Draw the lane onto the warped blank image
cv2.fillPoly(color_warp, np.int_([pts]), (0,255, 0))
# Warp the blank back to original image space
newwarp = birdeye(color_warp, corners, inverse = True)
# Combine the result with the original image
result = cv2.addWeighted(undistort, 1, newwarp, 0.3, 0)
font = cv2.FONT_HERSHEY_SIMPLEX
deviation = (mean(left_fitx) + mean(right_fitx))//2 - 650
cv2.putText(result,"Left Carvature: "+ str(left_curverad)+" Right Carvature: "+ str(right_curverad),(100,100), font, 1,(255,255,255),2,cv2.LINE_AA)
cv2.putText(result,"Deviation from the center: "+ str(deviation * xm_per_pix),(100,150), font, 1,(255,255,255),2,cv2.LINE_AA)
return result
# +
image_paths = glob(join('./test_images/', '*.jpg'))
for img_path in image_paths:
img = cv2.imread(img_path)
result = process_image(img)
head, tail = split(img_path)
cv2.imwrite('./output_images/output_' + tail , result)
# + active=""
# Video:
# +
output = 'test_videos_output/project_video.mp4'
clip2 = VideoFileClip('project_video.mp4')
clip = clip2.fl_image(process_image)
# %time clip.write_videofile(output, audio=False)
# -
HTML("""
<video width="960" height="540" controls>
<source src="{0}">
</video>
""".format(output))
# Test Image:
# +
img_path = './test_images/straight_lines1.jpg'
head, tail = split(img_path)
img = cv2.imread(img_path)
undistort, corners = undistort_image(img, calibration_images_dir = './camera_cal/')
cv2.imwrite('./output_images/undistorted_' + tail , undistort)
thresholded = combined_thresholds(img, sobel_kernel = 7, sobel_thresh=(10, 100), dir_thresh=(0.69, 1.3), s_thresh=(90, 255), l_thresh=(140, 255))
cv2.imwrite('./output_images/thresholded_' + tail , thresholded)
#thresholded = hls_select(undistort, s_thresh=(90, 255), l_thresh=(140, 255))
warped = birdeye(thresholded, corners)
cv2.imwrite('./output_images/warped_' + tail , warped)
out_img, left_fitx, right_fitx, ploty, left_fit_cr, right_fit_cr = fit_polynomial(warped)
cv2.imwrite('./output_images/color_fit_lines_' + tail , out_img)
left_curverad, right_curverad = measure_curvature_pixels(left_fit_cr, right_fit_cr, ploty)
#f, (ax1, ax2) = plt.subplots(1, 2, figsize=(24, 9))
#f.tight_layout()
#ax1.imshow(undistort)
#ax1.set_title('Original Image', fontsize=50)
#ax2.imshow(warped)
#ax2.set_title('Thresholded Grad. Dir.', fontsize=50)
#plt.subplots_adjust(left=0., right=1, top=0.9, bottom=0.)
#plt.imshow(warped)
#print(left_curverad, right_curverad)
# Create an image to draw the lines on
warp_zero = np.zeros_like(warped).astype(np.uint8)
color_warp = np.dstack((warp_zero, warp_zero, warp_zero))
# Recast the x and y points into usable format for cv2.fillPoly()
pts_left = np.array([np.transpose(np.vstack([left_fitx, ploty]))])
pts_right = np.array([np.flipud(np.transpose(np.vstack([right_fitx, ploty])))])
pts = np.hstack((pts_left, pts_right))
# Draw the lane onto the warped blank image
cv2.fillPoly(color_warp, np.int_([pts]), (0,255, 0))
# Warp the blank back to original image space
newwarp = birdeye(color_warp, corners, inverse = True)
# Combine the result with the original image
result = cv2.addWeighted(undistort, 1, newwarp, 0.3, 0)
font = cv2.FONT_HERSHEY_SIMPLEX
deviation = (mean(left_fitx) + mean(right_fitx))//2 - 650
cv2.putText(result,"Left Carvature: "+ str(left_curverad)+" Right Carvature: "+ str(right_curverad),(100,100), font, 1,(255,255,255),2,cv2.LINE_AA)
cv2.putText(result,"Deviation from the center: "+ str(deviation * xm_per_pix),(100,150), font, 1,(255,255,255),2,cv2.LINE_AA)
cv2.imwrite('./output_images/example_output_' + tail , result)
| Advanced Lane Finding.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:py2.7]
# language: python
# name: conda-env-py2.7-py
# ---
pwd
# cd Desktop/BIA-660/Final Project/data
# ### Increasing rate of Cumulative trip since 2015 becomes small, we consider it plateaued
#
# so, I used 2015,2016 data for training, and 2017 for testing.
# More specific, I trained data by season, so for each season there are 3 (month/year) * 2 (year) = 6 months
# For later testing, you only need to identify which season it belongs to.
# +
# glob.glob("2016*.csv")
# +
# all_data = pd.DataFrame()
# for f in glob.glob("2016*.csv"):
# df = pd.read_csv(f)
# all_data = all_data.append(df,ignore_index=True)
# +
#del all_data['Birth Year']
#del all_data['User Type']
#del all_data['Gender']
# -
import pandas as pd
import numpy as np
import re
import datetime
from sklearn.linear_model import LinearRegression
from sklearn import metrics
from sklearn.cross_validation import train_test_split
from six.moves import urllib
import json
import collections
import itertools
import requests
import time
import pandas as pd
from __builtin__ import any as b_any
import math
from sklearn.cross_validation import cross_val_score
from sklearn.ensemble import RandomForestRegressor
from sklearn import linear_model
from sklearn.neural_network import MLPRegressor
from collections import Counter
import matplotlib.pyplot as plt
from sklearn.linear_model import Lasso
# %matplotlib inline
# ## Integrate Variables & Data Preprocess
# +
# =========== Pre-process ==========
df = pd.read_csv('201612-citibike-tripdata.csv')
#df = pd.read_csv('201608-citibike-tripdata.csv')
#df = all_data
# Missing values in columns
#print(df.isnull().sum())
# Snake_case the columns
def camel_to_snake(name):
s1 = re.sub('(.)([A-Z][a-z]+)', r'\1_\2', name)
return re.sub('([a-z0-9])([A-Z])', r'\1_\2', s1).replace(' ','').lower()
df.columns = [camel_to_snake(col) for col in df.columns]
# Parse start_time and stop_time
try:
df['start_datetime'] = [datetime.datetime.strptime(x, '%m/%d/%Y %H:%M:%S') for x in df.start_time]
#except ValueError:
#df['start_datetime'] = [datetime.datetime.strptime(x, '%m/%d/%Y %H:%M') for x in df.start_time]
except ValueError:
df['start_datetime'] = [datetime.datetime.strptime(x, '%Y-%m-%d %H:%M:%S') for x in df.start_time]
df['start_day'] = [x.strftime('%Y-%m-%d') for x in df.start_datetime]
# =========== Group by date and station ===========
df['bikein'] = 1
df['bikeout'] = 1
df_bikeout = df.groupby(by=['start_stationid', 'start_day'])['bikeout'].count()
df_bikeout = pd.DataFrame(df_bikeout)
df_out = df_bikeout.reset_index(drop = False)
df_out.rename_axis({'start_stationid':'station_id'}, axis='columns', inplace=True)
print(df_out.head())
df_bikein = df.groupby(by=['end_stationid', 'start_day'])['bikein'].count()
df_bikein = pd.DataFrame(df_bikein)
df_in = df_bikein.reset_index(drop = False)
df_in.rename_axis({'end_stationid':'station_id'}, axis='columns', inplace=True)
#print(df_in.head())
# =========== Bike loss per day for each station ===========
# Merge two DFs by station_id
df_dayloss = df_out.merge(right=df_in, left_on=['station_id', 'start_day'], right_on=['station_id', 'start_day'],
how='outer')
df_dayloss = df_dayloss.sort_values(['station_id', 'start_day'])
df_dayloss = df_dayloss.fillna(0)
df_dayloss['bike_loss'] = df_dayloss['bikeout'] - df_dayloss['bikein']
print(df_dayloss[1:5])
# -
df_dayloss[df_dayloss.station_id==72][1:5]
# append weekday_index column
#wekday_name=pd.to_datetime(df_dayloss.start_day).dt.weekday_name #name
df_wekday_index=pd.to_datetime(df_dayloss.start_day).dt.weekday # index Monday=0, Sunday=6
# Append weekday column to df_dayloss
df_dayloss['wek_index']=df_wekday_index
# +
# =========== count docks Then append to dayloss============
jsonurl = urllib.request.urlopen('https://gbfs.citibikenyc.com/gbfs/en/station_status.json')
text = json.loads(jsonurl.read()) # <-- read from it
data=text['data']
#print(len(data['stations'])) 664 stations
#print(data['stations'][0]['station_id']) # station id
# station id Extract:
total_stations_id = []
for i in range(len(data['stations'])):
id=data['stations'][i]['station_id']
total_stations_id.append(id)
total_stations_id=[int(x) for x in total_stations_id]
#print('station_id list:'+str(total_stations_id))
# num_bikes_available Extract:
total_num_bikes_available = []
for i in range(len(data['stations'])):
num_bikes_available = data['stations'][i]['num_bikes_available']
total_num_bikes_available.append(num_bikes_available)
#print('bikes availble:'+str(total_num_bikes_available))
# num_docks_available Extract:
total_docks_available = []
for i in range(len(data['stations'])):
docks_availble=data['stations'][i]['num_docks_available']
total_docks_available.append(docks_availble)
#print('docks availble:'+str(total_docks_available))
total_docks=[x + y for x, y in zip(total_num_bikes_available, total_docks_available)]
#print('total docks for each station:'+str(total_docks))
#to_dict
new_dict = dict(zip(total_stations_id,total_docks))
#print('zip station_id and total docks:'+str(new_dict))
# append docks to df_dayloss
df_dayloss['docks'] = df_dayloss['station_id'].map(new_dict)
# -
# Append holiday column to df_dayloss
holiday = ['2016-01-01', '2016-01-18', '2016-02-12', '2016-02-15',
'2016-05-08', '2016-05-30', '2016-06-19', '2016-07-04',
'2016-09-05', '2016-10-10', '2016-11-11', '2016-11-24',
'2016-12-26']
df_dayloss['holiday_idx'] = df_dayloss['start_day'].apply(lambda x: 1 if (x in holiday) else 0)
# ## Get Weather Data
#
# 1. Get highest temperature of that day
# 2. Get Weather Condition (Sunny, Rainy...)
# Note: Since the Weather Condition we extract for the day is the condition that appears most frequently in 8 equal
# time piece of the day(0-3, 3-6...), so we didn't use the amount of rain, instead we just simply used whether there is rain or not.
#
def weather_Data(df_climate):
dates = []
maxtempF = []
cltype = []
each_day_weather_set = []
dk = df_climate.drop_duplicates(['start_day'])
for x in dk['start_day']:
dates.append(x)
api_str = 'http://api.worldweatheronline.com/premium/v1/past-weather.ashx?key=c3e90d1cd020454594c203537172604&q=NY&date=' + x + '&format=json' # print api_str
time.sleep(0.02)
r = requests.get(api_str)
#print "request over"
k = r.json()
maxtempF.append(k['data']['weather'][0]['maxtempF'])
size = len(k['data']['weather'][0]['hourly'])
climate = []
for l in range(size):
climate.append(str(k['data']['weather'][0]['hourly'][l]['weatherDesc'][0].values()).split(" "))
to_str = str(climate)
r1=to_str.replace('[','').replace(']','').replace('"','').replace("u'",'').replace("'",'').replace(' ','')
r2=r1.split(",")
weather_condition_count=Counter(r2)
most_common_weather=weather_condition_count.most_common(1)[0][0]
each_day_weather_set.append(most_common_weather)
new_dict = dict(zip(dates, maxtempF))
new_dict2 = dict(zip(dates, each_day_weather_set))
#print each_day_weather_set
#print dates
df_climate['High_temp'] = df_climate['start_day'].map(new_dict)
df_climate['Climate_type'] = df_climate['start_day'].map(new_dict2)
return df_climate
time.sleep(0.2)
weather_Data(df_dayloss)
set(df_dayloss.Climate_type)
# ## change categorical variables to dummy variables:
#
# 1. wek_index: Weekday:0; Weekend:1
# 2. holiday_index: Non-holiday:0; Holiday:1
# 3. High_temp: continuous
# 4. Climate_type(Sunny, Rainy....): Dummy Variables.
# +
# Assign weekday = 0 and weekend = 1
df_dayloss['wek_index']=df_dayloss['wek_index'].replace([1,2,3,4],[0,0,0,0])
df_dayloss['wek_index']=df_dayloss['wek_index'].replace([5,6],[1,1]) # weekend
# Get Dummy Variables for 'Climate Type'
dummy_weather=pd.get_dummies(df_dayloss['Climate_type'])
df_dayloss = pd.concat([df_dayloss, dummy_weather], axis=1)
# Remove docks col
del df_dayloss['docks']
# Remove Climate_type col
del df_dayloss['Climate_type']
# -
df_dayloss[31:60]
#df_dayloss=df_dayloss.drop(df_dayloss.columns[8:11], axis=1)
# Make variables for regression in continuous columns
# cols = df_dayloss.columns.tolist()
# cols.insert(len(cols)-1, cols.pop(cols.index('Climate_type')))
# df_dayloss = df_dayloss.reindex(columns=cols)
# ## Some Visilizations
# 1. Average bike_loss for each station
avg_loss=df_dayloss.groupby('station_id', as_index=False)['bike_loss'].mean()
plt.scatter(avg_loss.station_id, avg_loss.bike_loss,s=50)
axes = plt.gca()
axes.set_xlim([0,3500])
axes.set_ylim([-5,5])
small_sta_id=avg_loss[avg_loss.station_id<=2000]
print small_sta_id.describe()
large_sta_id=avg_loss[avg_loss.station_id>2000]
print large_sta_id.describe()
# ## Tune parameters --- Random Search & Grid Search
#
# 1. Random Search: use random search to find good starting points,
# 2. Grid search: to zoom in and find the local optima (or close to it) for those good starting points
# ## Tune Parameters For Each Model-- Grid Search
#
# Description:
# Grid search is an approach to parameter tuning that will methodically build and evaluate a model for each combination of algorithm parameters specified in a grid.
#
# 1. Random Forest
# 2. Lasso
# 3. Ridge
# 4. MLP
#
# NOTE: GRID SEARCH WILL FILTER STATIONS THAT DESN'T MEET THE REQUIREMENT!(SAMPLE SIZE AT THAT STATION IS TOO SMALL TO CV)
len(set(df['start_stationid']))
# Delete station that has less than 20 samples (Not good for CV)
station_id_set = list(set(df_dayloss['station_id']))
useless_ids_index = []
for i in station_id_set:
sample_size=len(df_dayloss[df_dayloss['station_id']==i])
if sample_size < 20:
useless_ids_index.append(station_id_set.index(i))
station_id_set=np.delete(station_id_set, useless_ids_index).tolist()
len(station_id_set)
station_id_set
# +
from sklearn.model_selection import GridSearchCV
parameter_box=[]
# RF:
reg_type = RandomForestRegressor()
param_grid = [
{'n_estimators': [3, 10, 30], 'max_features': [2, 4, 6, 8, 9]}
]
# Lasso:
# reg_type = linear_model.Lasso()
# param_grid = [{'alpha':[1,0.1,0.01,0.001,0.0001,0,10.2,12.4,15,18]}]
#MLPRegressor -- Neural Network
# reg_type = MLPRegressor()
# param_grid = [{ 'hidden_layer_sizes':[(5),(50),(50,100),(100,50,100)],
# 'activation':['relu','identity', 'logistic', 'tanh'] ,
# 'max_iter':[50,200,500],
# }]
def grid_search(reg_type,param_grid):
grid_search = GridSearchCV(reg_type, param_grid, cv=5,
scoring='neg_mean_squared_error',n_jobs=-1)
for i in station_id_set:
a=df_dayloss[df_dayloss['station_id']==i]
X=a.ix[:,5:16]
y=a.ix[:,4]
grid_search.fit(X, y)
print grid_search.best_params_
parameter_box.append(grid_search.best_params_)
grid_search(reg_type,param_grid)
# -
Lasso_para_value = [x['alpha'] for x in parameter_box]
MLP_para_value1=[x['activation'] for x in parameter_box]
MLP_para_value2=[x['hidden_layer_sizes'] for x in parameter_box]
MLP_para_value3=[x['max_iter'] for x in parameter_box]
RF_para_value1=[x['max_features'] for x in parameter_box]
RF_para_value2=[x['n_estimators'] for x in parameter_box]
para_all=pd.concat([pd.DataFrame(Lasso_para_value),pd.DataFrame(MLP_para_value1),pd.DataFrame(MLP_para_value2),
pd.DataFrame(MLP_para_value3),pd.DataFrame(RF_para_value1),pd.DataFrame(RF_para_value2)],axis=1)
para_all.columns=['Lasso-Alpha','MLP-Activation','MLP-HiddenLayerSize','MLP-MaxIter','RF-MaxFeatures','RF-NEstimates']
print len(para_all)
print len(station_id_set)
para_all.index=station_id_set
# Tune Para Result: For each station, each model.
para_all
# ## Linear Regression
#
# 1. Linear regression for each station(December)
# 2. Coff Analysis (Larger coff represents for larger effect of that variable)
# 3. Significant test for each independent variable
# 4. R^2 & accuracy of Linear Regression Model (When test data coming in)
#
# # Cross Validation
#
# 1. CV for different models.
# note: useless_id_index generated in linear regression CV
# ## Cross Validation -- Linear Regression
# +
LR_CV_score_set = []
LR_culmilative_set = []
# Count RMSE (Root Mean Squared Error) for every station
for i in station_id_set:
a=df_dayloss[df_dayloss['station_id']==i]
X=a.ix[:,5:16]
y=a.ix[:,4]
linearReg = LinearRegression()
mean_CV_score = math.sqrt(-(cross_val_score(linearReg, X, y, cv=10, scoring='neg_mean_squared_error').mean()))
#print 'CV score for station %s is: '%i +str(mean_CV_score)
LR_CV_score_set.append(mean_CV_score)
LR_culmilative_set.append(np.mean(LR_CV_score_set))
print 'Mean Score of Cross Validation using Linear Regression is:' + str(np.mean(LR_CV_score_set))
print 'Standard Deviation Score of Cross Validation using Linear Regression is:' + str(np.std(LR_CV_score_set))
# -
# ## Cross Validation - Lasso
lasso_CV_score_set = []
lasso_culmilative_set = []
for i, j in zip(list(station_id_set), para_value): # loop simultaneously
a=df_dayloss[df_dayloss['station_id']==i]
X=a.ix[:,5:16]
y=a.ix[:,4]
lasso = linear_model.Lasso(alpha=j)
mean_CV_score = math.sqrt(-(cross_val_score(lasso, X, y,cv=5, scoring='neg_mean_squared_error').mean()))
print 'CV score for station %s is: '%i +str(mean_CV_score)
lasso_CV_score_set.append(mean_CV_score)
lasso_culmilative_set.append(np.mean(lasso_CV_score_set))
print 'Mean Score of Cross Validation using Lasso Regression is: ' + str(np.mean(lasso_CV_score_set))
print 'Standard Deviation Score of Cross Validation using Lasso Regression is: ' + str(np.std(lasso_CV_score_set))
print len(station_id_set)
print len(dict_value)
# ## Random Forest
#
# 1. Importance of Each Feature
# 2. Accuracy of Random Forest Model
# ## Cross Validation --- Random Forest
RF_CV_score_set = []
RF_culmilative_set = []
for i,j,k in zip(list(station_id_set),RF_para_value1,RF_para_value2):
a=df_dayloss[df_dayloss['station_id']==i]
X=a.ix[:,5:16]
y=a.ix[:,4]
rf = RandomForestRegressor(max_features=j,n_estimators=k)
mean_CV_score = math.sqrt(-(cross_val_score(rf, X, y, cv=10, scoring='neg_mean_squared_error').mean()))
print 'CV score for station %s is: '%i +str(mean_CV_score)
RF_CV_score_set.append(mean_CV_score)
RF_culmilative_set.append(np.mean(RF_CV_score_set))
print 'Mean Score of Cross Validation using Random Forest is: ' + str(np.mean(RF_CV_score_set))
print 'Standard Deviation Score of Cross Validation using Random Forest is: ' + str(np.std(RF_CV_score_set))
# ## Cross Validation -- Neural Network
#
# 1. MLP: Multi-layer Perceptron
from sklearn.neural_network import MLPRegressor
MLP_CV_score_set = []
MLP_culmilative_set = []
for i,j,k,p in zip(list(station_id_set),MLP_para_value1,MLP_para_value2,MLP_para_value3):
a=df_dayloss[df_dayloss['station_id']==i]
X=a.ix[:,5:16]
y=a.ix[:,4]
MLP = MLPRegressor(activation=j,hidden_layer_sizes=k,max_iter=p)
mean_CV_score = math.sqrt(-(cross_val_score(MLP, X, y, cv=10, scoring='neg_mean_squared_error').mean()))
print 'CV score for station %s is: '%i +str(mean_CV_score)
MLP_CV_score_set.append(mean_CV_score)
MLP_culmilative_set.append(np.mean(MLP_CV_score_set))
print 'Mean Score of Cross Validation using MLP is: ' + str(np.mean(MLP_CV_score_set))
print 'Standard Deviation Score of Cross Validation using MLP is: ' + str(np.std(MLP_CV_score_set))
# ## RF better than LR !
# +
# The total RMASE is counted by average rmse of all stations
# For each model(LR & RF), the culmulative RMSE changes when new station coming in.
from pylab import *
t1 = LR_culmilative_set
t2 = lasso_culmilative_set
t3 = RF_culmilative_set
t4 = MLP_culmilative_set
s = station_id_set
plt.plot(s, t1, label='LR')
plt.plot(s, t2, label='Lasso')
plot(s, t3, label='RF')
plot(s, t4, label='NN')
xlabel('Station ID')
ylabel('Culmilative RMSE')
title('Python Line Chart: Culmulative RMSE plot for LR, Lasso, RF & NN')
grid(True)
plt.legend(loc='lower right',ncol=2)
plt.show()
# -
# ## Testing
# =========== Preparing test data==========
df = pd.read_csv('201611-citibike-tripdata.csv')
def camel_to_snake(name):
s1 = re.sub('(.)([A-Z][a-z]+)', r'\1_\2', name)
return re.sub('([a-z0-9])([A-Z])', r'\1_\2', s1).replace(' ','').lower()
df.columns = [camel_to_snake(col) for col in df.columns]
try:
df['start_datetime'] = [datetime.datetime.strptime(x, '%m/%d/%Y %H:%M:%S') for x in df.start_time]
#except ValueError:
#df['start_datetime'] = [datetime.datetime.strptime(x, '%m/%d/%Y %H:%M') for x in df.start_time]
except ValueError:
df['start_datetime'] = [datetime.datetime.strptime(x, '%Y-%m-%d %H:%M:%S') for x in df.start_time]
df['start_day'] = [x.strftime('%Y-%m-%d') for x in df.start_datetime]
# =========== Group by date and station ===========
df['bikein'] = 1
df['bikeout'] = 1
df_bikeout = df.groupby(by=['start_stationid', 'start_day'])['bikeout'].count()
df_bikeout = pd.DataFrame(df_bikeout)
df_out = df_bikeout.reset_index(drop = False)
df_out.rename_axis({'start_stationid':'station_id'}, axis='columns', inplace=True)
df_bikein = df.groupby(by=['end_stationid', 'start_day'])['bikein'].count()
df_bikein = pd.DataFrame(df_bikein)
df_in = df_bikein.reset_index(drop = False)
df_in.rename_axis({'end_stationid':'station_id'}, axis='columns', inplace=True)
# =========== Bike loss per day for each station ===========
df_dayloss_test = df_out.merge(right=df_in, left_on=['station_id', 'start_day'], right_on=['station_id', 'start_day'],
how='outer')
df_dayloss_test = df_dayloss_test.sort_values(['station_id', 'start_day'])
df_dayloss_test = df_dayloss_test.fillna(0)
df_dayloss_test['bike_loss'] = df_dayloss_test['bikeout'] - df_dayloss_test['bikein']
print(df_dayloss_test[1:5])
df_wekday_index=pd.to_datetime(df_dayloss_test.start_day).dt.weekday # index Monday=0, Sunday=6
df_dayloss_test['wek_index']=df_wekday_index
holiday = ['2016-01-01', '2016-01-18', '2016-02-12', '2016-02-15',
'2016-05-08', '2016-05-30', '2016-06-19', '2016-07-04',
'2016-09-05', '2016-10-10', '2016-11-11', '2016-11-24',
'2016-12-26']
df_dayloss_test['holiday_idx'] = df_dayloss_test['start_day'].apply(lambda x: 1 if (x in holiday) else 0)
def weather_Data(df_climate):
dates = []
maxtempF = []
cltype = []
each_day_weather_set = []
dk = df_climate.drop_duplicates(['start_day'])
for x in dk['start_day']:
dates.append(x)
api_str = 'http://api.worldweatheronline.com/premium/v1/past-weather.ashx?key=c3e90d1cd020454594c203537172604&q=NY&date=' + x + '&format=json' # print api_str
time.sleep(0.02)
r = requests.get(api_str)
#print "request over"
k = r.json()
maxtempF.append(k['data']['weather'][0]['maxtempF'])
size = len(k['data']['weather'][0]['hourly'])
climate = []
for l in range(size):
climate.append(str(k['data']['weather'][0]['hourly'][l]['weatherDesc'][0].values()).split(" "))
to_str = str(climate)
r1=to_str.replace('[','').replace(']','').replace('"','').replace("u'",'').replace("'",'').replace(' ','')
r2=r1.split(",")
weather_condition_count=Counter(r2)
most_common_weather=weather_condition_count.most_common(1)[0][0]
each_day_weather_set.append(most_common_weather)
new_dict = dict(zip(dates, maxtempF))
new_dict2 = dict(zip(dates, each_day_weather_set))
#print each_day_weather_set
#print dates
df_climate['High_temp'] = df_climate['start_day'].map(new_dict)
df_climate['Climate_type'] = df_climate['start_day'].map(new_dict2)
return df_climate
time.sleep(0.2)
weather_Data(df_dayloss_test)
# +
# Assign weekday = 0 and weekend = 1
df_dayloss_test['wek_index']=df_dayloss_test['wek_index'].replace([1,2,3,4],[0,0,0,0])
df_dayloss_test['wek_index']=df_dayloss_test['wek_index'].replace([5,6],[1,1]) # weekend
# Get Dummy Variables for 'Climate Type'
dummy_weather=pd.get_dummies(df_dayloss_test['Climate_type'])
df_dayloss_test = pd.concat([df_dayloss_test, dummy_weather], axis=1)
# Remove docks col
#del df_dayloss['docks']
# Remove Climate_type col
del df_dayloss_test['Climate_type']
# -
len(set(df_dayloss_test.station_id))
len(df_dayloss_test)
training = df_dayloss
del training['Fog']
del training['Freezing']
training
# ## Predict Using Linear Regression
final_predict_set = []
for i,j in zip(station_id_set,station_id_set):
a=training[training['station_id']==i]
X=a.ix[:,5:14]
Y=a.ix[:,4]# Assign features to be X and target to be Y.
try:
mdl = LinearRegression()
mdl.fit(X,Y)
b=df_dayloss_test[df_dayloss_test['station_id']==j]
test_data=b.ix[:,5:14]
final_predictions = mdl.predict(test_data) # I just used trained X to test.
#print final_predictions
final_predict_set.append(final_predictions)
continue
except:
continue
LR_preditct = pd.DataFrame(final_predict_set)
LR_preditct=pd.DataFrame(LR_preditct)
one_col=pd.concat([LR_preditct[0],LR_preditct[1],LR_preditct[2],LR_preditct[3],LR_preditct[4],LR_preditct[5],LR_preditct[6],LR_preditct[7],LR_preditct[8],
LR_preditct[9],LR_preditct[10],LR_preditct[11],LR_preditct[12],LR_preditct[13],LR_preditct[14],LR_preditct[15],
LR_preditct[16],LR_preditct[17],LR_preditct[18],LR_preditct[19],LR_preditct[20],LR_preditct[21],LR_preditct[22],
LR_preditct[23],LR_preditct[24],LR_preditct[25],LR_preditct[26],LR_preditct[27],LR_preditct[28],
LR_preditct[29]],axis=0)
# +
index_sorted=pd.DataFrame(one_col.sort_index(axis=0))
index_sorted.isnull().sum()
#len(index_sorted)-447
print index_sorted
len(index_sorted)
# -
sum(LR_preditct.isnull().sum())
# ## Predict Using Random Forest
#
# +
final_predict_set = []
for i in station_id_set:
a=training[training['station_id']==i]
X=a.ix[:,5:14]
y=a.ix[:,4]
final_model = RandomForestRegressor(n_estimators=30,max_features=4)
try:
final_model.fit(X,y) # Here fit the train model
except:
continue
# -
station_id_set = set(df_dayloss_test['station_id'])
final_predict_set = []
for j in station_id_set:
b=df_dayloss_test[df_dayloss_test['station_id']==j]
test_data=b.ix[:,5:14]
final_predictions = final_model.predict(test_data) # I just used trained X to test.
# print final_predictions
final_predict_set.append(final_predictions)
pd.DataFrame(final_predict_set)
# # Other Stuff
# Linear regession equations
station_id_set = list(set(df_dayloss['station_id']))
for i in station_id_set:
a=df_dayloss[df_dayloss['station_id']==i]
X=a.ix[:,5:16]
Y=a.ix[:,4]# Assign features to be X and target to be Y.
try:
mdl = LinearRegression()
mdl.fit(X,Y)
mdl.predict
m = mdl.coef_
b = mdl.intercept_
#print 'y={}x+{}'.format(m,b)
except:
continue
# +
#Feature Importance using RF(If needed)
from sklearn.linear_model import RandomizedLasso
from sklearn.ensemble import RandomForestRegressor
from sklearn.feature_selection import RFE
for i in station_id_set:
a=df_dayloss[df_dayloss['station_id']==i]
X=a.ix[:,5:16]
y=a.ix[:,4]# Assign features to be X and target to be Y.
# rf = RandomForestRegressor(n_estimators=30)
# rf.fit(X, y)
# print rf.feature_importances_
# lr = LinearRegression()
# rfe = RFE(lr, n_features_to_select=1)
# rfe.fit(X,y)
rlasso = RandomizedLasso(alpha=0.1)
try:
rlasso.fit(X, y)
except:
continue
# rfe = RFE(rf, n_features_to_select=1)
# rfe.fit(X,Y)
# -
varible_names=list(df_dayloss)[5:]
# from sklearn.linear_model import RandomizedLasso
# #Feacture Importance continue
names = varible_names
names = np.array(names)
#sorted(zip(names, rf.feature_importances_),key=lambda x: x[1],reverse=True)
print 'feature imoirtance rank using lasso_score is: '
sorted(zip(names, rlasso.scores_),key=lambda x: x[1],reverse=True)
#print sorted(zip(map(lambda x: round(x, 4), rfe.ranking_), names))
# ### training data split
dayloss_72 =df_dayloss[df_dayloss['station_id'] == 79]
from sklearn.cross_validation import train_test_split
import matplotlib.pyplot as plt
# %matplotlib inline
# training data split
X = dayloss_72[['wek_index','holiday_idx','docks']]
y = dayloss_72['bike_loss']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.25, random_state = 123)
#y_train, y_test = cross_validation(y, test_size = 0.25, random_state = 123)
X_train = X_train.sort_index()
X_test = X_test.sort_index()
y_train = y_train.sort_index()
y_test = y_test.sort_index()
X_train[1:5]
plt.plot(dayloss_72.index, dayloss_72.bike_loss)
# ### Lasso
from sklearn.linear_model import Lasso
from sklearn.model_selection import KFold
import numpy as np
import math
def RMSPE(y, y_pred):
return math.sqrt( ((1- y_pred / y)**2).mean() )
def test_lasso(alphas):
scores_lasso = []
kf = KFold(n_splits=10, random_state = 123)
for alpha in alphas:
lasso = Lasso(alpha)
for train, val in kf.split(X_train):
score_cv = []
y_pred = lasso.fit(X_train.values[train],y_train.values[train]).predict(X_train.values[val])
score_cv.append(RMSPE(y_train.values[val], y_pred))
score = np.mean(score_cv)
scores_lasso.append(score)
plt.figure(figsize=(14,8))
cv_lasso = pd.Series(scores_lasso, index = alphas)
cv_lasso.plot(title = "Validation - Lasso")
plt.xlabel("alpha")
plt.ylabel("RMSPE")
print(cv_lasso.argmin())
alphas = [1e-4, 1e-3,1e-2, 0.1, 1, 5, 10, 20]
test_lasso(alphas)
alphas = [10, 12, 14, 18, 20]
test_lasso(alphas)
alphas = [10, 10.5, 11, 11.25, 11.5, 11.75, 12, 12.25, 12.5, 13]
test_lasso(alphas)
alphas = [10, 10.10,10.15, 10.20, 10.25, 10.30,10.40,10.55,10.7]
test_lasso(alphas)
# +
lasso = Lasso(alpha=10.2)
scores_lasso = []
kf = KFold(n_splits=10, random_state = 123)
for alpha in alphas:
lasso = Lasso(alpha)
for train, val in kf.split(X_train):
score_cv = []
y_pred = lasso.fit(X_train.values[train],y_train.values[train]).predict(X_train.values[val])
score_cv.append(RMSPE(y_train.values[val], y_pred))
score = np.mean(score_cv)
scores_lasso.append(score)
# -
# prediction of bike loss of station 72
y_pred
| 660-final_4-29.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ## Cleaning up Atlas data - ETHZ ClimWIP
# **Function** : Preprocess netCDF files and restructure the dataset<br>
# **Description** : In this notebook serves to clean up Atlas data which is given in netcdf format and aggregate the data into a single file.<br>
# **Return Values : .nc files**<br>
# **Note** : All the data is saved to netCDF4 format. Note that data from different models may vary concerning the resolution and coordinates.<br>
# +
import os
from pathlib import Path
import numpy as np
import pandas as pd
import xarray as xr
# -
# ### Path
# Specify the path to the dataset and the place to save the outputs. <br>
# +
# please specify data path
datapath = Path("./AtlasData/raw")
# please specify output path
output_path = Path("./AtlasData/preprocess")
os.makedirs(output_path, exist_ok=True)
# -
# Components used to create the output file names. Here, only `institution_id` and `cmor_var` is based on on CMIP DRS conventions.
# + tags=[]
output_file_name = {
"prefix": "atlas",
"activity": "EUCP", # project name e.g. EUCP
"institution_id": "ETHZ", # ETHZ
"source": "CMIP6", # e.g. CMIP6 or CMIP5
"method": "ClimWIP", # e.g. ClimWIP
"sub_method": "cons", # e.g. cons or uncons
"cmor_var": "tas", # e.g. tas or pr
}
# -
# ### Load and process raw data
# Make some functions to combining multiple dimensions with a preprocessor and load data
# +
INSTITUTION_ID = output_file_name["institution_id"]
METHOD = output_file_name["method"]
def add_percentile_tas(ds):
filename = ds.encoding["source"]
percentile = int(filename.split("_")[-1][:2])
_, variable, future, season, dataset, _ = filename.split("_")[1:]
return ds.assign_coords(percentile=percentile).expand_dims("percentile")
def add_percentile_pr(ds):
filename = ds.encoding["source"]
percentile = int(filename.split("_")[-2][:2])
_, variable, future, season, dataset, _ = filename.split("_")[1:-1]
return ds.assign_coords(percentile=percentile).expand_dims("percentile")
# data loader and batch processing
def load_data(project, season, variable):
# open multiple files with xarray
if variable == "tas":
ds = xr.open_mfdataset(
str(
Path(
datapath,
f"{INSTITUTION_ID}_{METHOD}",
f"eur_{variable}_41-60_{season}_{project}_*perc.nc",
)
),
preprocess=add_percentile_tas,
)
elif variable == "pr":
ds = xr.open_mfdataset(
str(
Path(
datapath,
f"{INSTITUTION_ID}_{METHOD}",
f"eur_{variable}_41-60_{season}_{project}_*perc_rel.nc",
)
),
preprocess=add_percentile_pr,
)
else:
raise ValueError("Given variable is not valid.")
weighted = (
ds[f"{variable}_mean_weighted"]
.rename(variable)
.assign_coords(constrained=1)
.expand_dims("constrained")
)
unweighted = (
ds[f"{variable}_mean"]
.rename(variable)
.assign_coords(constrained=0)
.expand_dims("constrained")
)
return xr.concat([weighted, unweighted], dim="constrained")
# -
# Call functions
project = output_file_name["source"].lower()
seasons = []
for season in ["djf", "jja"]:
tas = load_data(project, season, "tas")
pr = load_data(project, season, "pr")
ds = xr.merge([tas, pr]).assign_coords(season=season.upper())
seasons.append(ds)
ethz_climwip_ds = xr.concat(seasons, dim="season")
ethz_climwip_ds
# Make some metadata. Here, we follow CF-conventions as much as possible.
attrs = {
"tas": {
"description": "Change in Air Temperature",
"standard_name": "Change in Air Temperature",
"long_name": "Change in Near-Surface Air Temperature",
"units": "K", # in line with raw data
"cell_methods": "time: mean changes over 20 years 2041-2060 vs 1995-2014",
},
"pr": {
"description": "Relative precipitation",
"standard_name": "Relative precipitation",
"long_name": "Relative precipitation",
"units": "%", # in line with raw data
"cell_methods": "time: mean changes over 20 years 2041-2060 vs 1995-2014",
},
"latitude": {"units": "degrees_north", "long_name": "latitude", "axis": "Y"},
"longitude": {"units": "degrees_east", "long_name": "longitude", "axis": "X"},
"time": {
"climatology": "climatology_bounds",
"long_name": "time",
"axis": "T",
"climatology_bounds": ["2050-6-1", "2050-9-1", "2050-12-1", "2051-3-1"],
"description": "mean changes over 20 years 2041-2060 vs 1995-2014. The mid point 2050 is chosen as the representative time.",
},
"percentile": {"units": "%", "long_name": "percentile", "axis": "Z"},
}
# ### Assemble data and save to netcdf
# Make a function to assemble and save data
# +
TIMES = {
"JJA": "2050-7-16",
"DJF": "2051-1-16",
} # "0000-4-16", "0000-7-16", "0000-10-16", "0000-1-16" MAM JJA SON DJF
PERCENTILES = [10, 25, 50, 75, 90]
LAT = ethz_climwip_ds.coords["lat"]
LON = ethz_climwip_ds.coords["lon"]
def assembly(ds_original, var, cons):
"""
Select data from original nc files and save the target fields.
"""
ds_target = xr.Dataset(
{
var: (
("time", "latitude", "longitude", "percentile"),
np.full([len(TIMES), len(LAT), len(LON), len(PERCENTILES)], np.nan),
),
"climatology_bounds": (
pd.to_datetime(["2050-6-1", "2050-9-1", "2050-12-1", "2051-3-1"])
),
},
coords={
"time": pd.to_datetime(list(TIMES.values())),
"latitude": LAT.values,
"longitude": LON.values,
"percentile": PERCENTILES,
},
attrs={
"description": f"Contains modified {INSTITUTION_ID} {METHOD} data used for Atlas in EUCP project.",
"history": f"original {INSTITUTION_ID} {METHOD} data files eur_pr_41-60_djf_cmip6_*perc_rel.nc, eur_tas_41-60_djf_cmip6_*perc.nc",
},
)
for season in ["JJA", "DJF"]:
for j, p in enumerate(PERCENTILES):
ds_target[var].values[list(TIMES).index(season), :, :, j] = (
ds_original[var]
.sel(percentile=p, season=season, constrained=cons)
.values
)
return ds_target
# -
# Call the function
for VAR_NAME in ["tas", "pr"]:
output_file_name["cmor_var"] = VAR_NAME
for i, sub_method in enumerate(["uncons", "cons"]):
output_file_name["sub_method"] = sub_method
new_ds = assembly(ethz_climwip_ds, VAR_NAME, i)
# Fix attributes
for key in new_ds.keys():
new_ds[key].attrs = attrs[key]
file_name = f"{'_'.join(output_file_name.values())}.nc"
print(f"one dataset is saved to {file_name}")
new_ds.to_netcdf(output_path / file_name)
# ### Check output
# Load one of the saved data.
ds = xr.open_dataset(output_path / "atlas_EUCP_ETHZ_CMIP6_ClimWIP_cons_tas.nc")
ds
| python/cleanup_ETHZ_ClimWIP_atlas_netcdf.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# Find the bounding box of an object
# ===================================
#
# This example shows how to extract the bounding box of the largest object
#
#
#
# +
import numpy as np
from scipy import ndimage
import matplotlib.pyplot as plt
np.random.seed(1)
n = 10
l = 256
im = np.zeros((l, l))
points = l*np.random.random((2, n**2))
im[(points[0]).astype(np.int), (points[1]).astype(np.int)] = 1
im = ndimage.gaussian_filter(im, sigma=l/(4.*n))
mask = im > im.mean()
label_im, nb_labels = ndimage.label(mask)
# Find the largest connected component
sizes = ndimage.sum(mask, label_im, range(nb_labels + 1))
mask_size = sizes < 1000
remove_pixel = mask_size[label_im]
label_im[remove_pixel] = 0
labels = np.unique(label_im)
label_im = np.searchsorted(labels, label_im)
# Now that we have only one connected component, extract it's bounding box
slice_x, slice_y = ndimage.find_objects(label_im==4)[0]
roi = im[slice_x, slice_y]
plt.figure(figsize=(4, 2))
plt.axes([0, 0, 1, 1])
plt.imshow(roi)
plt.axis('off')
plt.show()
| _downloads/plot_find_object.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
fruitdata = pd.read_table("C:/Users/home/Downloads/fruit_data_with_colors.txt")
fruitdata.head()
fruitdata.fruit_name.value_counts().plot.pie(autopct= "%1.1f%%")
sns.countplot(fruitdata["fruit_name"])
plt.show()
fruitdata.fruit_subtype.value_counts().plot.pie(autopct= "%1.1f%%")
sns.countplot(fruitdata["fruit_subtype"])
locs, labels = plt.xticks()
plt.setp(labels, rotation=75)
fruitdata.mass.hist()
fruitdata.width.hist()
fruitdata.height.hist()
fruitdata.color_score.hist()
plt.scatter(fruitdata["mass"], fruitdata["width"], edgecolors='y')
plt.xlabel('Mass')
plt.ylabel('Width')
plt.title('Fruit Mass to Width')
plt.show()
plt.scatter(fruitdata["mass"], fruitdata["height"], edgecolors='y')
plt.xlabel('Mass')
plt.ylabel('Height')
plt.title('Fruit Mass to Height')
plt.show()
sns.boxplot(fruitdata["mass"])
sns.boxplot(fruitdata["height"])
sns.boxplot(fruitdata["width"])
sns.boxplot(fruitdata["color_score"])
corr=fruitdata.corr()
corr.nlargest(5,'fruit_label')['fruit_label']
fruitdata.head()
fruitdata =fruitdata.drop(["fruit_name"],axis=1)
fruitdata.head()
x = np.array(fruitdata.drop(["fruit_subtype","fruit_label"],axis=1))
x[0:5]
y = np.array(fruitdata["fruit_label"])
y[0:5]
from sklearn.model_selection import train_test_split
x_train,x_test,y_train,y_test=train_test_split(x,y,test_size=0.3,random_state=6)
from sklearn.naive_bayes import GaussianNB
gnb = GaussianNB()
gnb.fit(x_train,y_train)
pred=gnb.predict(x_test)
pred
gnb.score(x_train,y_train)
gnb.score(x_test,y_test)
x_test[0:10]
y_test[0:10]
gnb.predict(x_test)
fruitdata = fruitdata.drop(["fruit_subtype"],axis=1)
fruitdata.head()
fruitdata.shape
a= fruitdata.drop(["fruit_label"],axis=1)
a[20:30]
gnb.predict(a)
b= fruitdata.iloc[:,1:5]
b
| FruitGaussianNBClassifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Examples
#
# Below we show three examples of the mx_viz functions. Please see the `slide_figures.ipynb` and the OHBM 2020 multilayer network educational presentation for more details.
# +
## Import packages
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import networkx as nx
import multinetx as mx
import scipy.io as sio
import pandas as pd
import os
import sys
import mx_viz
print("Done importing packages :)")
# -
# ## Create a multilayer network
#
# Using multinetx, we will create a small multilayer (mx) network for plotting later. Note that the visualization functions will work best with _small_ mx networks. For larger mx networks, consider alternative plotting strategies found in this [2019 paper](https://onlinelibrary.wiley.com/doi/full/10.1111/cgf.13610).
#
# Note below we will assume that all nodes exist in every layer and that nodes only connect to their counterparts in other layers.
# +
# Define number of nodes (number of nodes in largest layer).
nNodes = 10
# Define number of levels.
nLayers = 3
# Use multinetx to generate three graphs each on nNodes nodes.
g1 = mx.generators.erdos_renyi_graph(nNodes,0.5,seed=216)
g2 = mx.generators.erdos_renyi_graph(nNodes,0.5,seed=130)
g3 = mx.generators.erdos_renyi_graph(nNodes,0.5,seed=81)
# Define adjacency between layers. Here we only assign nodes to themselves in each layer.
adj_block = mx.lil_matrix(np.zeros((nNodes*nLayers,nNodes*nLayers)))
for i in np.arange(nLayers-1):
for l in np.arange(i+1,nLayers):
adj_block[i*nNodes:(i+1)*nNodes, (l)*nNodes:(l+1)*nNodes] = np.identity(nNodes)
adj_block += adj_block.T
# Create multilayer graph with mx.
mg = mx.MultilayerGraph(list_of_layers=[g1,g2,g3],
inter_adjacency_matrix=adj_block)
# Here we can set the edge weights to different values just so we can see which are inter- and intra-layer edges.
mg.set_edges_weights(intra_layer_edges_weight=2,
inter_layer_edges_weight=1)
## Plot the supra-adjacency matrix to check that we actually made a multilayer network.
fig = plt.figure(figsize=(6,5))
sns.heatmap(mx.adjacency_matrix(mg,weight='weight').todense())
plt.title('supra adjacency matrix');
# -
# ### Write to json and create visualization.
#
# The first mx_viz function writes the graph to a json file, while the second reads the file and creates an html file with the visualization.
#
# Below we will create two html files - one using the `theme="light"` flag and the other with `theme="dark"`.
# +
# We'll use networkx positioning to get nice layouts. However pos is changed (for example by
# using a different nx.layout function), it should remain a dictionary mapping nodes to coordinate
# arrays with at least an x and y position.
pos = nx.layout.fruchterman_reingold_layout(g1, dim=3, k=2)
filename_json = "data/example1.json"
G2 = mx_viz.write_mx_to_json(filename_json,mg, nNodes, pos, nLayers)
filename_html_light = "visualization_output_example_light.html"
mx_viz.visualize(G2,theme="light",path_html=filename_html_light)
filename_html_dark = "visualization_output_example_dark.html"
mx_viz.visualize(G2,theme="dark",path_html=filename_html_dark);
# -
# ## Create a temporal network
#
# Temporal networks are special types of multilayer networks in which the layers correspond to timepoints. Time has a natural ordering, so we create a slightly different visualization that respects the ordered layers.
#
# Again, we assume that all nodes exist in every layer and that nodes connect only to their counterparts in every layer.
#
# +
# Define number of nodes (number of nodes in largest layer)
nNodes = 10
# Define number of timepoints (levels)
nLayers = 14
# Use multinetx to generate fourteen graphs each on nNodes nodes
graph_layers = [mx.generators.erdos_renyi_graph(nNodes,((i+1)/(nLayers*2+2)),seed=np.random.randint(1,300)) for i in np.arange(nLayers)]
# Define adjacency between layers. Here, again, we only assign nodes to themselves in each layer.
adj_block = mx.lil_matrix(np.zeros((nNodes*nLayers,nNodes*nLayers)))
for i in np.arange(nLayers-1):
for l in np.arange(i+1,nLayers):
adj_block[i*nNodes:(i+1)*nNodes, (l)*nNodes:(l+1)*nNodes] = np.identity(nNodes)
adj_block += adj_block.T
# Create multilayer graph
mg = mx.MultilayerGraph(list_of_layers=graph_layers,
inter_adjacency_matrix=adj_block)
# Set edge weights
mg.set_edges_weights(intra_layer_edges_weight=2,
inter_layer_edges_weight=1)
## Plot the supra-adjacency matrix
fig = plt.figure(figsize=(6,5))
sns.heatmap(mx.adjacency_matrix(mg,weight='weight').todense())
plt.title('supra adjacency matrix');
# +
# As before, generate positions for nodes in the first layer.
pos = nx.layout.fruchterman_reingold_layout(graph_layers[0], dim=3)
filename_json = "data/example2.json"
G2 = mx_viz.write_mx_to_json(filename_json,mg, nNodes, pos, nLayers)
filename_html = "visualization_output_example_timeseries.html"
mx_viz.visualize_timeseries(G2, path_html=filename_html);
# -
# Open the html files in your browser and enjoy!
| mx_viz_examples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="u8QcXQ_4POSt" colab_type="text"
#
#
#
# ### **<font> Emotion Recognizer </font>**
# + [markdown] id="ttDOvPzPPXky" colab_type="text"
# #◢ Mevon-AI - Recognize Emotions in Speech
#
# This program is for recognizing emotions from audio files generated in a customer care call center. A customer care call center of any company receives many calls from customers every day. Every call is recorded for analysis purposes. The program aims to analyse the emotions of the customer and employees from these recordings.
#
# The emotions are classified into 6 categories: 'Neutral', 'Happy', 'Sad', 'Angry', 'Fearful', 'Disgusted', 'Surprised'
#
# Analysing the emotions of the customer after they have spoken with the company's employee in the call center can allow the company to understand the customer's behaviour and rate the performance of its employees accordingly.
#
#
# ####**Credits:**
#
# * [Speech Emotion Recognition from Saaket Agashe's Github](https://github.com/saa1605/speech-emotion-recognition)
# * [Speech Emotion Recognition with CNN](https://towardsdatascience.com/speech-emotion-recognition-with-convolution-neural-network-1e6bb7130ce3)
# * [MFCCs Tutorial](http://practicalcryptography.com/miscellaneous/machine-learning/guide-mel-frequency-cepstral-coefficients-mfccs/)
# * [UIS-RNN Fully Supervised Speaker Diarization](https://github.com/google/uis-rnn)
# * [uis-rnn and speaker embedding by vgg-speaker-recognition by taylorlu](https://github.com/taylorlu/Speaker-Diarization)
#
# + [markdown] id="0nJiZwrIPqPK" colab_type="text"
#
#
# ---
#
#
# #◢ Verify Correct Runtime Settings
#
# **<font color='#FF000'> IMPORTANT </font>**
#
# In the "Runtime" menu for the notebook window, select "Change runtime type." Ensure that the following are selected:
# * Runtime Type = Python 3
# * Hardware Accelerator = GPU
#
# + [markdown] id="bg1lRc5VQG8f" colab_type="text"
# #◢ Git clone and install Mevon-AI Speech Emotion Recognition
# + id="PTWK6HAPPL43" colab_type="code" colab={}
# !git clone https://github.com/SuyashMore/MevonAI-Speech-Emotion-Recognition.git
# + id="NNOzO-tyQzdL" colab_type="code" colab={}
# cd MevonAI-Speech-Emotion-Recognition/src
# + [markdown] id="5vFfeI0IQYhD" colab_type="text"
# #◢ Setup
# + id="S9zJdxSLY_rr" colab_type="code" colab={}
# !chmod +x setup.sh
# !./setup.sh
# + [markdown] id="Eam-QOmHajCk" colab_type="text"
# #◢ Instructions
#
# ## Add Audio Files
# You can add audio files in any language inside input/ folder.
#
# For eg. currently, there are 3 folders for 3 different Employees inside the input/ directory.
# Each folder contains 1 audio file of conversation between that employee with a customer. You can add many more files in each of the employee's folder.
#
# If you have 5 employees, then create 5 folders inside the **input/** directory. Then add audio files of conversation with customer of each employee in the respective folders.
#
# ## Run Mevon_AI
# Demo for running the main program is given in the next section.
#
# ## Diarization Output
# Since each audio file has 2 speakers: customer and employee of the customer care call center, we split the audio file into 2 such that one audio file contains the audio of customer and other contains the audio of employee.
#
# These splitted audio files are stored in **output**/ folder
#
# ##Predicted Emotions
# The audio file of each customer is analysed by the CNN model and a **.csv** file is generated which contains the predicted emotion
#
# + [markdown] id="iXB2kKqRZN-Q" colab_type="text"
# #◢ Recognize Emotions!!
# + id="KXD8AbtrZgqY" colab_type="code" colab={}
import os
os.environ['TF_CPP_MIN_LOG_LEVEL'] = '3'
# + id="0rBWrBWlW9Jd" colab_type="code" colab={}
# !python3 speechEmotionRecognition.py
| src/notebooks/Mevon_AI_Speech_Emotion_Recognition_Demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] pycharm={"name": "#%% md\n"}
# # Data Visualization semester project
# ## Meteorite landings data pre-processing
#
# #### Libraries:
# + pycharm={"name": "#%%\n"}
import pandas as pd
# + [markdown] pycharm={"name": "#%% md\n"}
# Import the meteorite landings dataset from the NASA website.
# + pycharm={"name": "#%%\n"}
URL: str ="https://data.nasa.gov/api/views/gh4g-9sfh/rows.csv?accessType=DOWNLOAD"
df: pd.DataFrame = pd.read_csv(URL)
# + [markdown] pycharm={"name": "#%% md\n"}
# ### Pre processing:
#
# #### Brief exploration
# + pycharm={"name": "#%%\n"}
print(df.shape)
df.dtypes
# + pycharm={"name": "#%%\n"}
df.head()
# + [markdown] pycharm={"name": "#%% md\n"}
# We have:
# - 6 categorical variables of interest: id, name, nametype, recclass fall and year.
# - if there are no duplicate names id does not provide added information.
# - year is stored in the wrong format
# - 3 numerical variables of interest: mass, longitude and latitude.
# - mass does not follow the naming convention of the rest of the dataset.
# - GeoLocation does not contain additional information.
# + pycharm={"name": "#%%\n"}
df['name'].duplicated().any()
# + [markdown] pycharm={"name": "#%% md\n"}
# Since the name uniquely identifies the meteorite we can drop the id together with GeoLocation
# + pycharm={"name": "#%%\n"}
df.drop(['id', 'GeoLocation'], axis='columns', inplace=True)
# + [markdown] pycharm={"name": "#%% md\n"}
# renaming mass (g) to mass and giving more meaningful names to nametype,
# which represents whether the meteorite landed as a meteorite or is now a
# relic after landing, renamed to status and fall, which represents how the meteorite
# was detected, while falling or found, to detection method.
#
# Renaming reclat and reclong and recclass to latitude and longitude and recommended
# classification, a standard on how to categorize meteorites.
# + pycharm={"name": "#%%\n"}
df.rename(columns={'mass (g)': 'mass',
'nametype': 'status',
'fall': 'detection method',
'year': 'year fell or found',
'reclat': 'latitude',
'reclong': 'longitude',
'recclass': 'recommended classification'}, inplace=True)
df.head()
# + [markdown] pycharm={"name": "#%% md\n"}
# Fixing the years
# + pycharm={"name": "#%%\n"}
from numpy.array_api import astype
df["year fell or found"] = df["year fell or found"].str[6:10]
df["year fell or found"].head()
# + [markdown] pycharm={"name": "#%% md\n"}
# Exploring the missing values:
# + pycharm={"name": "#%%\n"}
df.isnull().sum()
# -
# Our visualization requires geolocation, hence we drop the meteorites with
# missing latitude and longitude coordinates.
# + pycharm={"name": "#%%\n"}
df = df[df['latitude'].notna()]
df.isnull().sum()
# + pycharm={"name": "#%%\n"}
df.shape
# + [markdown] pycharm={"name": "#%% md\n"}
# Year and mass are not as important for us, hence we decided to keep those rows.
# Saving the dataset:
# + pycharm={"name": "#%%\n"}
df.to_csv("meteorite_landings_preprocessed.csv", index=False)
# + [markdown] pycharm={"name": "#%% md\n"}
# ## Below we show our aborted attempt to geo-locate the landing sites by country.
# #### Assigning an address to each meteorite landing found by the geo-locator with its coordinates.
#
# ##### Connection done using GeoPy's git-hub repository code:
# https://github.com/geopy/geopy/blob/master/geopy/geocoders/mapbox.py
# + pycharm={"name": "#%%\n"}
from functools import partial
from urllib.parse import quote, urlencode
from geopy.geocoders.base import DEFAULT_SENTINEL, Geocoder
from geopy.location import Location
from geopy.util import logger
__all__ = ("MapBox", )
class MapBox(Geocoder):
"""Geocoder using the Mapbox API.
Documentation at:
https://www.mapbox.com/api-documentation/
"""
api_path = '/geocoding/v5/mapbox.places/%(query)s.json/'
def __init__(
self,
api_key,
*,
scheme=None,
timeout=DEFAULT_SENTINEL,
proxies=DEFAULT_SENTINEL,
user_agent=None,
ssl_context=DEFAULT_SENTINEL,
adapter_factory=None,
domain='api.mapbox.com'
):
"""
:param str api_key: The API key required by Mapbox to perform
geocoding requests. API keys are managed through Mapox's account
page (https://www.mapbox.com/account/access-tokens).
:param str scheme:
See :attr:`geopy.geocoders.options.default_scheme`.
:param int timeout:
See :attr:`geopy.geocoders.options.default_timeout`.
:param dict proxies:
See :attr:`geopy.geocoders.options.default_proxies`.
:param str user_agent:
See :attr:`geopy.geocoders.options.default_user_agent`.
:type ssl_context: :class:`ssl.SSLContext`
:param ssl_context:
See :attr:`geopy.geocoders.options.default_ssl_context`.
:param callable adapter_factory:
See :attr:`geopy.geocoders.options.default_adapter_factory`.
.. versionadded:: 2.0
:param str domain: base api domain for mapbox
"""
super().__init__(
scheme=scheme,
timeout=timeout,
proxies=proxies,
user_agent=user_agent,
ssl_context=ssl_context,
adapter_factory=adapter_factory,
)
self.api_key = api_key
self.domain = domain.strip('/')
self.api = "%s://%s%s" % (self.scheme, self.domain, self.api_path)
def _parse_json(self, json, exactly_one=True):
"""Returns location, (latitude, longitude) from json feed."""
features = json['features']
if not features:
return None
def parse_feature(feature):
location = feature['place_name']
longitude = feature['geometry']['coordinates'][0]
latitude = feature['geometry']['coordinates'][1]
return Location(location, (latitude, longitude), feature)
if exactly_one:
return parse_feature(features[0])
else:
return [parse_feature(feature) for feature in features]
def reverse(
self,
query,
*,
exactly_one=True,
timeout=DEFAULT_SENTINEL
):
"""
Return an address by location point.
:param query: The coordinates for which you wish to obtain the
closest human-readable addresses.
:type query: :class:`geopy.point.Point`, list or tuple of ``(latitude,
longitude)``, or string as ``"%(latitude)s, %(longitude)s"``.
:param bool exactly_one: Return one result or a list of results, if
available.
:param int timeout: Time, in seconds, to wait for the geocoding service
to respond before raising a :class:`geopy.exc.GeocoderTimedOut`
exception. Set this only if you wish to override, on this call
only, the value set during the geocoder's initialization.
:rtype: ``None``, :class:`geopy.location.Location` or a list of them, if
``exactly_one=False``.
"""
params = {'access_token': self.api_key}
point = self._coerce_point_to_string(query, "%(lon)s,%(lat)s")
quoted_query = quote(point.encode('utf-8'))
url = "?".join((self.api % dict(query=quoted_query),
urlencode(params)))
logger.debug("%s.reverse: %s", self.__class__.__name__, url)
callback = partial(self._parse_json, exactly_one=exactly_one)
return self._call_geocoder(url, callback, timeout=timeout)
# + pycharm={"name": "#%%\n"}
from typing import Tuple
from typing import List
import time
mapbox: MapBox = MapBox('<KEY>')
points: pd.DataFrame = df[['latitude','longitude']]
point_list: List[Tuple] = [tuple(x) for x in points.to_numpy()]
# + pycharm={"name": "#%%\n"}
country = list()
def reverse_geo_locate():
for i in range(0, len(point_list)): # len(point_list)):
# do not exceed API calls per second
time.sleep(0.005)
with open('countries.csv', 'a', encoding='utf-8') as f:
location = mapbox.reverse(point_list[i])
country.append(location if location else ("no address","no country"))
# + pycharm={"name": "#%%\n"}
new_df = pd.DataFrame(country)
# + pycharm={"name": "#%%\n"}
new_df.to_csv('countries_last_attempt')
# + [markdown] pycharm={"name": "#%% md\n"}
# After checking the quality of the geo-located data, in particular the number of missing nations, we decided
# to use another strategy to represent the population density.
| preprocessing/meteorite_landings_preprocessing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# from now on, I will arrange the vector of 2D
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
# if you want to make vec in here, change the following variables
# set np.linspace function
lineSizePositive = 5
lineSizeNegative = -5
numberOfSampleOfLine = lineSizePositive * 2 + 1
# choose one of {'b', 'g', 'r', 'c', 'm', 'y', 'k', 'w'} as vector color
# 'b' : blue, 'g' : green, 'r' : red, 'c' : cyan,
# 'm' : meganta, 'y':yellow, 'k' : black, 'w' : whilte from https://matplotlib.org/api/colors_api.html
vectors = {"StartingPointOfX" : 0,"StartingPointOfY" : 0,
"SizeOfXvector" : 1,"StartingPointOfYvector" : 2,
"VectorColor" : "r"}
vectors2 = {"StartingPointOfX" : 1,"StartingPointOfY" : 0,
"SizeOfXvector" : 2,"StartingPointOfYvector" : 3,
"VectorColor" : "k"}
vectorList = [vectors, vectors2]
# if you want to make it better, just use array
# you might need the function meshgrid
# But In my case, just this function for linear algebra. I think I couldn't need it.
x = np.arange(25).reshape(5,5)
a = np.array([0,1,2,3, 'r'])
print (type(34))
for i in x :
print (i)
for i in a :
print (i)
#for vec in vectorList :
# print (vec["StartingPointOfX"]);
def PlotVector(legend=True) :
# Create a figure of size 8x6 inches, 80 dots per inch
fig = plt.figure(figsize=(8, 6), dpi=80)
# for the origin
#ax = fig.gca()
#ax.scatter([0],[0],color="k",s=100)
# line size of each axes
line = np.linspace(lineSizeNegative, lineSizePositive, numberOfSampleOfLine, endpoint=True, dtype="int")
# print varialble line
print ("line : \n {0}".format(line))
# Set x limits, * 1.1 just express more space for drawing
xlim = plt.xlim(line.min() * 1.1, line.max() * 1.1)
# print varialble line
#print ("xlim : \n {0}".format(xlim))
# Set x ticks
xticks = plt.xticks(line)
# print varialble line
#print ("xticks : \n {0}".format(xticks))
# Set x limits, * 1.1 just express more space for drawing
ylim = plt.ylim(line.min() * 1.1, line.max() * 1.1)
# print varialble line
#rint ("ylim : \n {0}".format(ylim))
# Set x ticks
yticks = plt.yticks(line)
# print varialble line
#rint ("yticks : \n {0}".format(yticks))
# set label name of x axis
plt.xlabel("x axis")
# set label name of y axis
plt.ylabel("y axis")
# set grid of coordinate to True
plt.grid(linestyle = "-")
# choose one of {'b', 'g', 'r', 'c', 'm', 'y', 'k', 'w'} as vector color
# 'b' : blue, 'g' : green, 'r' : red, 'c' : cyan,
# 'm' : meganta, 'y':yellow, 'k' : black, 'w' : whilte from https://matplotlib.org/api/colors_api.html
for vec in vectorList :
vecs = plt.quiver(vec["StartingPointOfX"], vec["StartingPointOfY"], vec["SizeOfXvector"], vec["StartingPointOfYvector"]
, angles="xy", scale_units="xy",scale=1, color=vec["VectorColor"])
#if (legend) :
#plt.legend(["The origin",r'$\vec a$',"test 2"])
plt.show()
PlotVector()
if __name__ == "__main__" :
print ("test")
# -
| img/Image/Languages/Python/2017-05-16-How_To_Plot_Vector_And_Plane_With_Python/2D_vector.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Implementation of Ada-Boost Algorithm
#Importing neccesary packages
# Load libraries
from sklearn.ensemble import AdaBoostClassifier
from sklearn import datasets
from sklearn.model_selection import train_test_split
from sklearn import metrics
# Load data
iris = datasets.load_iris()
X = iris.data
y = iris.target
# Split dataset into training set and test set
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2) # 80% training and 20% test
# +
# Create adaboost classifer object
AdaModel = AdaBoostClassifier(n_estimators=100,
learning_rate=1)
# Train Adaboost Classifer
model = AdaModel.fit(X_train, y_train)
#Predict the response for test dataset
y_pred = model.predict(X_test)
# -
# #Important Parameters
# base_estimator: It is a weak learner used to train the model. It uses DecisionTreeClassifier as default weak learner for training purpose. You can also specify different machine learning algorithms.
#
# n_estimators: Number of weak learners to train iteratively.
#
# learning_rate: It contributes to the weights of weak learners. It uses 1 as a default value.
# Model Accuracy, how often is the classifier correct?
print("Accuracy:",metrics.accuracy_score(y_test, y_pred))
# +
# Import Support Vector Classifier
from sklearn.svm import SVC
#Import scikit-learn metrics module for accuracy calculation
from sklearn import metrics
svc=SVC(probability=True, kernel='linear')
# Create adaboost classifer object
abc =AdaBoostClassifier(n_estimators=50, base_estimator=svc,learning_rate=1)
# +
# Train Adaboost Classifer
model = abc.fit(X_train, y_train)
#Predict the response for test dataset
y_pred = model.predict(X_test)
# -
# Model Accuracy, how often is the classifier correct?
print("Accuracy:",metrics.accuracy_score(y_test, y_pred))
# Pros
# AdaBoost is easy to implement. It iteratively corrects the mistakes of the weak classifier and improves accuracy by combining weak learners. You can use many base classifiers with AdaBoost. AdaBoost is not prone to overfitting. This can be found out via experiment results, but there is no concrete reason available.
#
# Cons
# AdaBoost is sensitive to noise data. It is highly affected by outliers because it tries to fit each point perfectly. AdaBoost is slower compared to XGBoost.
| Examples/Ada Boost/Ada-Boost Implementation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + pycharm={"name": "#%%\n"}
# Imports
import os
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import random
import re
import json
from sklearn.utils import shuffle
from sklearn.model_selection import train_test_split
from sklearn.utils import class_weight
from sklearn.metrics import f1_score
from sklearn.metrics import confusion_matrix, classification_report, accuracy_score
from sklearn.preprocessing import LabelEncoder
from tensorflow.keras.utils import to_categorical
from tensorflow.python.keras.callbacks import EarlyStopping
import tensorflow as tf
import tensorflow_addons as tfa
import seaborn as sn
from nltk.corpus import stopwords
import pickle
import tensorflow_hub as hub
import tensorflow_datasets as tfds
from official.modeling import tf_utils
from official import nlp
from official.nlp import bert
import official.nlp.optimization
import official.nlp.bert.bert_models
import official.nlp.bert.configs
import official.nlp.bert.run_classifier
import official.nlp.bert.tokenization
import official.nlp.data.classifier_data_lib
import official.nlp.modeling.losses
import official.nlp.modeling.models
import official.nlp.modeling.networks
# BERT model, tokenizer
from tensorflow.python.keras.layers import Flatten
from transformers import TFBertForSequenceClassification, TFAutoModel , AutoTokenizer
# + pycharm={"name": "#%%\n"}
# Constants
ITER = "1"
# Specialis karakterekkel a maximalis kodolt szoveg max hossza
SHUFFLE_RANDOM_STATE = 42
TRAIN_RANDOM_STATE = 42
TEST_RANDOM_STATE = 42
USE_STOPWORDS = True
LOAD_CHECKPOINT = False
CHECKPOINT_SUBDIR = 'sport/'
CHECKPOINT_PREFIX = 'restore_best_cased_stopwords_9/'
TEXT = 'Sentence'
START_TOKEN = 'START'
TOKEN_LEN = 'LEN'
Y_HEADER = 'LABEL'
LABELS = {
"SPORT": 0,
"VIDEÓJÁTÉK": 1
}
MAX_SEQUENCE_LENGTH = 64
BATCH_SIZE = 16
EPOCHS = 10
# Maximum hány dokumentumot nézzünk. None esetén mindet
TRAIN_PROCESSED_MAX_DOCUMENTS = None
# Maximum hány dokumentumot nézzünk. None esetén mindet
TEST_PROCESSED_MAX_DOCUMENTS = None
# + pycharm={"name": "#%%\n"}
physical_devices = tf.config.experimental.list_physical_devices('GPU')
print(physical_devices)
if physical_devices:
tf.config.experimental.set_memory_growth(physical_devices[0], True)
path = 'checkpoints/' + CHECKPOINT_SUBDIR + CHECKPOINT_PREFIX
# + pycharm={"name": "#%%\n"}
URL_RE = 'https?:\/\/(www\.)?[-a-zA-Z0-9@:%._\+~#=]{1,256}\.[a-zA-Z0-9()]{1,6}([-a-zA-Z0-9()@:%_+.~#?&/=]*)'
WHITELIST_RE = '[^a-zA-Z0-9íÍöÖüÜóÓőŐúÚáÁéÉűŰ]'
ures = 0
def cleanse(i):
global ures
text = dataset[TEXT].iloc[i]
text = re.sub(URL_RE, ' ', text)
text = re.sub(WHITELIST_RE, ' ', text)
text = ' '.join([text])
return re.sub(' +', ' ', text)
def delete_empty_rows(dataset):
ids_to_delete = dataset.index[dataset[TEXT] == ' '].tolist()
return dataset.drop(ids_to_delete)
# + pycharm={"name": "#%%\n"}
# Load and set up data
dataset = pd.read_csv('db/sport_or_e_sport.csv', sep=';', error_bad_lines=False)
dataset[Y_HEADER] = dataset[Y_HEADER].map(LABELS)
dataset = shuffle(dataset, random_state=SHUFFLE_RANDOM_STATE)
dataset.info()
dataset = delete_empty_rows(dataset)
for i in range(len(dataset.index)):
dataset[TEXT].iloc[i] = cleanse(i)
dataset = delete_empty_rows(dataset)
dataset.head(5)
X = dataset[TEXT].values
y = dataset[Y_HEADER].values
X_train, X_rem, y_train, y_rem = train_test_split(X, y, train_size=0.8, random_state=TRAIN_RANDOM_STATE)
X_dev, X_test, y_dev, y_test = train_test_split(X_rem, y_rem, train_size=0.5, random_state=TEST_RANDOM_STATE)
y_train_labels = y_train
y_dev_labels = y_dev
y_test_labels = y_test
y_train = to_categorical(y_train, 2)
y_dev = to_categorical(y_dev, 2)
y_test = to_categorical(y_test, 2)
# +
# eloszlas
def plot_label_counts(y, title='y labels'):
unique, counts = np.unique(y, return_counts=True)
b = dict(zip(unique, counts))
plt.barh(range(len(b)), list(b.values()), align='center', color=['lightblue', 'lightgreen'])
y_values = ["Sport", "Videójáték"]
y_axis = np.arange(0, 2, 1)
plt.yticks(y_axis, y_values)
plt.title(title)
plt.xlabel('Number of Samples in training Set')
plt.ylabel('Label')
ax = plt.gca()
for i, v in enumerate(b.values()):
plt.text(ax.get_xlim()[1]/100, i, str(v), color='blue', fontweight='bold')
plt.show()
plot_label_counts(y_train_labels, 'Train eloszlas')
plot_label_counts(y_dev_labels, 'Dev eloszlas')
plot_label_counts(y_test_labels, 'Test eloszlas')
# + pycharm={"name": "#%%\n"}
bert_tokenizer = AutoTokenizer.from_pretrained("SZTAKI-HLT/hubert-base-cc")
bert_model = TFBertForSequenceClassification.from_pretrained("SZTAKI-HLT/hubert-base-cc", num_labels=2)
# +
all_stopwords = []
def apply_stopwords(sentences):
global all_stopwords
corpus = []
for sen in sentences:
sentence = sen.split()
all_stopwords = stopwords.words('hungarian')
whitelist = ["ne", "nem", "se", "sem"]
sentence = [word for word in sentence if (word.lower() not in all_stopwords or word.lower() in whitelist)
and len(word) > 1]
sentence = ' '.join(sentence)
corpus.append(sentence)
return corpus
# + pycharm={"name": "#%%\n"}
def batch_encode(X):
return bert_tokenizer.batch_encode_plus(
X,
truncation=True,
max_length=MAX_SEQUENCE_LENGTH,
add_special_tokens=True, # add [CLS] and [SEP] tokens
return_attention_mask=True,
return_token_type_ids=False, # not needed for this type of ML task
padding='max_length', # add 0 pad tokens to the sequences less than max_length
return_tensors='tf'
)
X_train = X_train.tolist() if not USE_STOPWORDS else apply_stopwords(X_train.tolist())
X_dev = X_dev.tolist() if not USE_STOPWORDS else apply_stopwords(X_dev.tolist())
X_test = X_test.tolist() if not USE_STOPWORDS else apply_stopwords(X_test.tolist())
X_train = batch_encode(X_train[:TRAIN_PROCESSED_MAX_DOCUMENTS])
X_dev = batch_encode(X_dev[:TEST_PROCESSED_MAX_DOCUMENTS])
X_test = batch_encode(X_test[:TEST_PROCESSED_MAX_DOCUMENTS])
y_train = y_train[:TRAIN_PROCESSED_MAX_DOCUMENTS]
y_dev = y_dev[:TEST_PROCESSED_MAX_DOCUMENTS]
y_test = y_test[:TEST_PROCESSED_MAX_DOCUMENTS]
# + pycharm={"name": "#%%\n"}
def create_model():
input_ids = tf.keras.layers.Input(shape=(64,), dtype=tf.int32, name='input_ids')
attention_mask = tf.keras.layers.Input((64,), dtype=tf.int32, name='attention_mask')
output = bert_model([input_ids, attention_mask])[0]
output = tf.keras.layers.Dropout(rate=0.15)(output)
output = tf.keras.layers.Dense(2, activation='softmax')(output)
result = tf.keras.models.Model(inputs=[input_ids, attention_mask], outputs=output)
return result
model = create_model()
opt = tf.keras.optimizers.Adam(learning_rate=3e-5)
model.compile(optimizer=opt, loss='categorical_crossentropy', metrics=['accuracy'])
# + pycharm={"name": "#%%\n"}
print(bert_model.config)
model.summary()
# + pycharm={"name": "#%%\n"}
checkpoint_path = path + 'cp.ckpt'
cp_callback = tf.keras.callbacks.ModelCheckpoint(filepath=checkpoint_path,
save_weights_only=True,
verbose=1)
# + pycharm={"is_executing": true, "name": "#%%\n"}
def get_history_as_text(history, epoch):
return f'Epoch {epoch+1: <3}: loss: {format(history["loss"][epoch], ".4f")} - accuracy: {format(history["accuracy"][epoch], ".4f")} - val_loss: {format(history["val_loss"][epoch], ".4f")} - val_accuracy: {format(history["val_accuracy"][epoch], ".4f")}'
def fit_model():
if not LOAD_CHECKPOINT:
early_stopping_callback = EarlyStopping(monitor='val_loss', mode='min', verbose=1, patience=2, restore_best_weights=True)
if ITER == "11":
history = model.fit(
x=X_train.values(),
y=y_train,
validation_data=(X_dev.values(), y_dev),
epochs=EPOCHS,
batch_size=BATCH_SIZE,
callbacks=[early_stopping_callback, cp_callback]
)
else:
history = model.fit(
x=X_train.values(),
y=y_train,
validation_data=(X_dev.values(), y_dev),
epochs=EPOCHS,
batch_size=BATCH_SIZE,
callbacks=[early_stopping_callback]
)
with open(path + '.history', 'wb') as file_pi:
pickle.dump(history.history, file_pi)
text_file = open(path + "history" + ITER + ".txt", "w")
print(len(history.history['loss']))
for i in range(len(history.history['loss'])):
history_text = get_history_as_text(history.history, i)
print(history_text)
text_file.writelines(history_text + '\n')
text_file.close()
else:
model.load_weights(checkpoint_path)
history = pickle.load(open(path + '.history', "rb"))
for i in range(len(history['loss'])):
history_text = get_history_as_text(history, i)
print(history_text)
result = model.evaluate(X_test.values(), y_test)
predict = model.predict(X_test.values())
np_predict = np.argmax(predict,axis=1)
return history, result, np_predict
# + pycharm={"name": "#%%\n"}
#history, result, predict = fit_model()
# + pycharm={"name": "#%%\n"}
le = LabelEncoder()
def evaluate(predict, result, y):
y_le = le.fit_transform(y[:TEST_PROCESSED_MAX_DOCUMENTS])
print('Classification report:')
print(classification_report(y_le, predict))
print(f'Accuracy: {accuracy_score(y_le, predict): >43}')
print(f'Accuracy from evaluation: {result[1]: >27}')
print('Confusion matrix:')
df_cm = pd.DataFrame(confusion_matrix(y_le, predict),
index=[i for i in ['sport', 'videójáték']],
columns=[i for i in ['sport', 'videójáték']])
if not LOAD_CHECKPOINT:
with open(path + "results" + ITER + ".txt", "w") as text_file:
df_cm_string = df_cm.to_string(header=False, index=False)
text_file.write('Classification report:\n')
text_file.write(classification_report(y_le, predict))
text_file.write(f'\nAccuracy: {accuracy_score(y_le, predict): >43}\n')
text_file.write(f'Accuracy from evaluation: {result[1]: >27}\n')
text_file.write('\nConfusion matrix:\n')
text_file.write(df_cm_string)
plt.figure(figsize=(10,7))
plt.title(CHECKPOINT_PREFIX[:-1])
hm = sn.heatmap(df_cm, annot=True, fmt='g', cmap="Blues")
hm.set(ylabel='True label', xlabel='Predicted label')
if not LOAD_CHECKPOINT:
plt.savefig(path + 'accuracy-' + format(result[1], ".4f") + ITER + '.jpg')
plt.show()
# + pycharm={"name": "#%%\n"}
le = LabelEncoder()
def evaluate(predict, result, y):
y_le = le.fit_transform(y[:TEST_PROCESSED_MAX_DOCUMENTS])
print('Classification report:')
print(classification_report(y_le, predict))
print(f'Accuracy: {accuracy_score(y_le, predict): >43}')
print(f'Accuracy from evaluation: {result[1]: >27}')
print('Confusion matrix:')
df_cm = pd.DataFrame(confusion_matrix(y_le, predict),
index=[i for i in ['sport', 'videójáték']],
columns=[i for i in ['sport', 'videójáték']])
if not LOAD_CHECKPOINT:
with open(path + "results" + ITER + ".txt", "w") as text_file:
df_cm_string = df_cm.to_string(header=False, index=False)
text_file.write('Classification report:\n')
text_file.write(classification_report(y_le, predict))
text_file.write(f'\nAccuracy: {accuracy_score(y_le, predict): >43}\n')
text_file.write(f'Accuracy from evaluation: {result[1]: >27}\n')
text_file.write('\nConfusion matrix:\n')
text_file.write(df_cm_string)
plt.figure(figsize=(10,7))
plt.title(CHECKPOINT_PREFIX[:-1])
hm = sn.heatmap(df_cm, annot=True, fmt='g', cmap="Blues")
hm.set(ylabel='True label', xlabel='Predicted label')
if not LOAD_CHECKPOINT:
plt.savefig(path + 'accuracy-' + format(result[1], ".4f") + ITER + '.jpg')
plt.show()
# -
for i in range(9, 12):
ITER = str(i)
print("Iteration ", ITER)
history, result, predict = fit_model()
evaluate(predict, result, y_test_labels)
import winsound
#for i in range(2, 12):
# winsound.Beep(100 + i * 100, 400)
| second_semester/docby/sport_bert_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 3. Missing Data and Imputation
# Compiled by [<NAME>](mailto:<EMAIL>) for C3DIS 2018
# The 'achilles heel' of compositional data analysis is the incompatibility with 'null' or zero components. In practice, most compositional data contains such values - as either true zeros (e.g. count data as found in surveys), below-detection values (e.g. geochemistry) or components which are simply missing (e.g. components are not measured, but may be of significant quantity).
#
# The simplest but ultimately least practical solution to this problem is to use subcompositions - a set of components free of missing/zero data. Beyond this approach, missing data may be able to be imputed using relationships between variables. In the absence of additional information, typical data imputation methods utilise some measure of central tendency (e.g. median, mode, mean) as a fill value for missing data; this preserves the expected values, and largely preserves the covariance stucture of the dataset. However, we can use information we already have to provide more accurate imputation methods - leading to more robust analysis workflows.
#
# For compositional data, missing values commonly represent a 'detection limit' - a specific value below which the variable cannot be quantified, nor certified to be above background/zero (e.g. commonly 2$\sigma$ above zero). With regards to this form of missing data, a measure of central tendency is not likely an accurate representation - instead imputed values should lie below some threshold corresponding to the 'detction limit'. Here, the simplest form of imputation uses an arbitrarily small fill value. Imputation using nominal values may be marginally valid in some cases (i.e. using small values to represent values below decection, without altering the closure operation greatly), but overall this approach typically serves as a confounding factor (e.g. creating bimodal distributions and spurious clusters).
#
# Regardless of the method of imputation, when values are imputed from low density dataset, the output is strongly dependent on the data quality and 'representiveness' of the present values. For this reason, using imputed values for geological inference may be misleading - especially for rarely recorded parameters.
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
# +
import sys
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
sys.path.insert(0, './src')
from geochem import *
from compositions import *
from EMimputation import EMCOMP
# -
def simple_impute(arr, method='mean', threshold=0.01, limits=[]):
"""
"""
if method == 'mean':
# Note this is not a compositional mean, which should be: inv_alr(np.nanmean(alr(arr), axis=0)); this would require 1 nan-free column
replace_vals = np.nanmean(arr, axis=0)
elif method == 'eps': # constant arbitrarily small value
replace_vals = 2*np.finfo(float).eps * np.ones(arr.shape)
elif method == 'threshold': # detection limits or some ratio of the minimum values
if not len(limits):
replace_vals = np.tile(np.nanmin(arr, axis=0), (arr.shape[0])).reshape(arr.shape)
else:
assert len(limits) == arr.shape[1]
replace_vals = np.tile(np.array(limits), (arr.shape[0])).reshape(arr.shape)
else:
raise NotImplementedError
return np.where(~np.isfinite(arr), replace_vals, arr)
r1 = pd.DataFrame(dict(SiO2=[45.0], Al2O3=[10.0], MgO=[20.0], CaO=[25.0]),)
r2 = pd.DataFrame(dict(SiO2=[40.0], Al2O3=[12.0], MgO=[23.0], CaO=[25.0]),)
r3 = pd.DataFrame(dict(SiO2=[10.0], Ti=[10.0], Ge=[20.0], Tm=[25.0]),)
r4 = pd.DataFrame(dict(SiO2=[10.0], Al2O3=[3.0], Ti=[10.0], Ge=[20.0], Tm=[25.0]),)
aggdf = r1
for r in [r2, r3, r4]:
aggdf = aggdf.append(r, sort=False)
aggdf = aggdf.reset_index(drop=True)
aggdf
mean_imputed = aggdf.copy()
mean_imputed.loc[:, :] = simple_impute(aggdf.values, method='mean')
mean_imputed
eps_imputed = aggdf.copy()
eps_imputed.loc[:, :] = simple_impute(aggdf.values, method='eps')
eps_imputed
limit_imputed = aggdf.copy()
limit_imputed.loc[:, :] = simple_impute(aggdf.values, method='threshold', limits=0.01*np.nanmin(aggdf.values, axis=0))
limit_imputed
# #### Parametric Imputation using Multivariate Regression
# Parametric imputation attempts to preserve - and ideally *restore* (detection-limits effectively truncate distributions) - the distribution of multivariate compositional data. Missing values are imputed using regression against other variables. However, there remain difficulties with regards to imputation:
# * Omitted/not measured vs. 'below detection' values have different overall expectations: below detection values have upper bounds (and also upper error bounds), omitted values may be significant quantities, but are not well constrained
# * An iterative algorithm is needed (e.g. expectation-maximisation) as imputed values alter the closure operator and hence adjust other compostional components, if only slightly. Iteration continues until the imputed dataset resembles the original dataset within some specified tolerance (i.e. minimal perturbation to attain workable data).
# Here we use the EMCOMP algorithm of Palarea-Albaladejo and Martin-Fernandez (2008) to impute below-detection limit values. This requires at least one component free of zeros as a divisor. The method attempts to preserve the overall non-nan mean and covariance structure. This algorithm works well for relatively low-dimension data; future work will attempt to replace this algorithm with a bayesian imputation method along similar lines.
# +
def replace_below_detection(arr, limits=[0.1]):
if len(limits) == 1:
arr[arr < lim] = np.nan
else:
for ix, lim in enumerate(limits):
arr[arr[:, ix] < lim, ix] = np.nan
return arr
for pctl in [0.1, 1, 10., 20.]:
print(f'\nLimits at {pctl} percentile level.')
arrshape = (1000, 5)
arr = np.random.rand(*arrshape) * [1.0, 0.2, 0.01, 1.0, 0.5]
arr = np.divide(arr, np.nansum(arr, axis=1)[:, np.newaxis])
limits = np.percentile(arr, pctl, axis=0)
arr2 = arr.copy()
arr2[:, 2:] = replace_below_detection(arr2[:, 2:], limits=limits[2:])
imputed_arr, s, n = EMCOMP(arr2, limits)
print(f'Proportion BDL: \t\t{arr2[~np.isfinite(arr2)].size / arr2.size:2.3f}')
print(f'Maximum Abs. Difference: \t{(imputed_arr-arr).max()*100:2.2f}%')
# -
| 03_Missing_Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ランダムなエルミート行列の量子位相推定
#
# Authors: <NAME> (Blueqat inc.), <NAME> (Riken), <NAME>, <NAME> (Quemix inc.), <NAME> (Blueqat inc.)
#
# [前回](113_pea_ja.ipynb)はZゲートやXゲートの行列の固有値を求めました。今回は、ランダムな2x2エルミート行列の量子位相推定をします。
#
# エルミート行列の固有値を求めることは量子力学において物理量を求めることに相当し、量子化学や量子シミュレーションなどの分野で広く応用が期待されます。
# ## 固有値の和が0になるエルミート行列の量子位相推定
#
# エルミート行列の固有値を量子位相推定アルゴリズムで求める方法も、大まかな原理は前回と全く同じです。ですが、前回はZ行列やX行列の固有値を求めるために、Controlled-Uゲートとして、Controlled-ZゲートやControlled-Xゲートを用意しましたが、今回、エルミート行列はユニタリ行列になるとは限らないため、Controlled-Uゲートを直接用意することができません。
#
# 結論だけを言うと、エルミート行列$\hat H$の固有ベクトルを作るような量子回路を用意して、また、Controlled-UゲートとしてControlled-$e^{2\pi i\hat H}$ゲートを作ります。そうして量子位相推定を行うことで、固有値を求めることができます。
#
# 以下の説明は少し難しいので、飛ばしてコードを読んでも構いません。また、ここに書かれていることを理解するには[行列指数関数](https://ja.wikipedia.org/wiki/%E8%A1%8C%E5%88%97%E6%8C%87%E6%95%B0%E9%96%A2%E6%95%B0)についての知識が必要です。
#
# 固有値の和が0になるようなエルミート行列$\hat H$を考えます。
#
# $\hat H$が固有値と固有ベクトルの組$\{(\lambda_j, \left|\psi_j\right\rangle)\}$を持つとき、任意の$i$について
# $$\hat H\left|\psi_j\right\rangle = \lambda_j \left|\psi_j\right\rangle$$
# の関係を満たします。$\hat H$がエルミート行列の場合、$\lambda_j$が必ず実数となることが知られています。
#
# 前回は、$\hat H = Z$または$\hat H = X$のようにして、位相キックバックにはControlled-ZゲートやControlled-Xゲートを使いました。ですが、量子ゲートを並べて作れるのはユニタリ行列だけなので、一般のエルミート行列については、全く同じ方法は取ることができません。
#
# そこで、天下り式ですが、ユニタリ行列 $U = e^{2\pi i\hat H}$を使って量子位相推定することを考えてみます。
#
# ### $U = e^{2\pi i\hat H}$はユニタリ行列になるか?
#
# $U U^\dagger = I$を示したいので、$U^\dagger$を考えます。
# $$\begin{eqnarray}
# U^\dagger &=& (e^{2\pi i\hat H})^\dagger\nonumber\\
# &=& (\sum_n^\infty \frac{(2\pi i\hat H)^n}{n!})^\dagger\nonumber\\
# &=& \sum_n^\infty \frac{((2\pi i\hat H)^n)^\dagger}{n!}\nonumber\\
# &=& \sum_n^\infty \frac{(-2\pi i\hat H^\dagger)^n)}{n!}\nonumber\\
# &=& \sum_n^\infty \frac{(-2\pi i\hat H)^n)}{n!}\nonumber\\
# &=& e^{-2\pi i\hat H}\nonumber
# \end{eqnarray}$$
# なので、$U U^\dagger = e^{2\pi i\hat H} e^{-2\pi i\hat H}$となります。ここで、$[2\pi i\hat H, -2\pi i\hat H] = 0$なので、$e^{2\pi i\hat H} e^{-2\pi i\hat H} = e^{2\pi i\hat H -2\pi i\hat H} = e^{0\hat H} = I$とできます。すなわち$U U^\dagger = I$なので$U$はユニタリ行列です。
#
# ### $U$の固有値と固有ベクトルは何になるか。$\hat H$のそれとの関係は?
#
# $U$の固有ベクトルが$\hat H$の固有ベクトルと同じであることを示し、その固有値を求めます。
#
# ある$\hat H$の固有値と固有ベクトルの組$\lambda_j, \left|\psi_j\right\rangle$を考えます。このとき、
# $$\begin{eqnarray}
# U \left|\psi_j\right\rangle &=& (\sum_n^\infty \frac{(2\pi i\hat H)^n}{n!})\left|\psi_j\right\rangle\nonumber\\
# &=& \sum_n^\infty \frac{(2\pi i)^n \hat H^n \left|\psi_j\right\rangle}{n!}\nonumber\\
# &=& \sum_n^\infty \frac{(2\pi i)^n \lambda_j^n \left|\psi_j\right\rangle}{n!}\nonumber\\
# &=& \sum_n^\infty \frac{(2\pi i)^n \lambda_j^n}{n!}\left|\psi_j\right\rangle\nonumber\\
# &=& \sum_n^\infty \frac{(2\pi i \lambda_j)^n}{n!}\left|\psi_j\right\rangle\nonumber\\
# &=& e^{2\pi i \lambda_j}\left|\psi_j\right\rangle\nonumber
# \end{eqnarray}$$
# なので、$\left|\psi_j\right\rangle$は$U$のひとつの固有ベクトルで、対応する固有値は$e^{2\pi i \lambda_j}$です。
#
# これにより、
# $$U\left|\psi_j\right\rangle = e^{2\pi i\lambda_j} \left|\psi_j\right\rangle$$
# を量子位相推定することで、$\hat H$の固有値$\lambda_j$を計算できることが分かりました。
# ## 実装
# まず、必要なライブラリをインポートします。また、今回はnumbaバックエンドを使用します。
# +
import math
import cmath
import random
import numpy as np
from blueqat import *
from blueqat.pauli import X, Y, Z, I
BlueqatGlobalSetting.set_default_backend('numba')
pi = math.pi
# Blueqat バージョンチェック
try:
Circuit().r(0.1)[0].run()
except AttributeError:
raise ImportError('Blueqat version is old.')
# -
# 次に、ランダムなエルミート行列を作ります。
#
# 位相推定により固有値を求めるには、ターゲットとなるエルミート行列$\hat H$の他に
# - 固有ベクトル(またはその近似値)を与える量子回路
# - Controlled-$e^{i2\pi \hat H 2^n}$を与える量子回路
#
# が必要です。これらも得られるような形で、エルミート行列を作ります。
#
#
# エルミート行列は$\hat H = P D P^\dagger$
# (ここで、$P$はユニタリ行列、$D$は実対角行列)の形で書き表せます。
#
# このとき、$D$の対角成分が固有値になり、$P$の各列が固有ベクトルとなります。今は、固有値の合計が0になる2x2のエルミート行列を考えるので、$D$の成分は$\pm E$となります(ただし、$\hat H$の固有値のひとつを$E$とおいた)。
#
# そのため、$D$の成分となる$E$をランダムに生成し、また、$P$は、任意のユニタリ行列を与えるゲートであるU3ゲートのパラメータ$\theta, \phi, \lambda$の3つ組をランダムに生成し、
#
# - エルミート行列$\hat H$
# - 答え合わせのための固有値$E$
# - 固有ベクトルを与える量子回路を作るために必要なU3ゲートのパラメータ$\theta, \phi, \lambda$
#
# を返すような関数を定義します。
# +
def is_hermitian(mat):
"""matがエルミート行列かどうかを判定する"""
# matの転置共役がmatと等しい場合、matはエルミート行列
return np.allclose(mat, mat.T.conjugate())
def get_u3_matrix(theta, phi, lam):
"""U3をユニタリ行列の形で得る"""
# Blueqatのto_unitary()で得られる行列を、numpy形式に変換して作ります。
u = Circuit().u3(theta, phi, lam)[0].to_unitary()
return np.array(u.tolist()).astype(np.complex64)
def random_hermitian():
"""ランダムにエルミート行列を作り、
エルミート行列, 固有値, パラメータ(3つ組)
を返す。
"""
# 固有値のひとつをランダムに決める
eigval = random.random()
# U3ゲートのパラメータをランダムに決める
theta = random.random()
phi = random.random()
lam = random.random()
# これらから、エルミート行列を作る
u3 = get_u3_matrix(theta, phi, lam)
hermitian = u3 @ np.diag([eigval, -eigval]) @ u3.T.conjugate()
# エルミート行列であることを確認
assert is_hermitian(hermitian)
# エルミート行列, 固有値, パラメータを返す
return hermitian, eigval, (theta, phi, lam)
# -
# エルミート行列を作ってみます。
H, E, (theta, phi, lam) = random_hermitian()
print(H)
# 固有ベクトルは、`theta, phi, lam`とU3ゲートから作ることができます。
vec = Circuit().u3(theta, phi, lam)[0].run()
print(vec)
# これが固有ベクトルであることを確かめるために、`H vec = E vec`が成り立つことを確かめます。
np.allclose(np.dot(H, vec), E * vec)
# 準備ができたので、これから量子位相推定を実装していきます。具体的には、`theta, phi, lam`から量子回路を作って、量子位相推定により`E`を求めます。
# +
def iqft(c, q0, n_qubits):
"""回路のq0〜q0 + n_qubits - 1番目ビットに量子逆フーリエ変換の操作を付け加える"""
for i in reversed(range(n_qubits)):
angle = -0.5
for j in range(i + 1, n_qubits):
c.cr(angle * pi)[q0 + j, q0 + i]
angle *= 0.5
c.h[q0 + i]
return c
def initial_circuit(theta, phi, lam):
"""初期回路(つまり、固有ベクトルを表す回路)を用意します"""
return Circuit().u3(theta, phi, lam)[0]
def apply_cu(c, ctrl, theta, phi, lam, eigval, n):
"""Controlled-U^(2^n)を量子回路cに付け加えて返します。
制御ビットをctrl, 標的ビットを0としています。
ここで、この関数は固有値eigvalを引数にとっていることに気をつけて下さい。
今回は、固有値を使って(ズルをして)Controlled-U^(2^n)を作ります。
固有値を使わなくても、鈴木-トロッター展開などにより、近似的に作ることはできますが、
その場合、精度には十分注意が必要です。
このようなズルをせず、効率よく高精度に、こういった回路を作ることは、一般には困難です。
"""
return c.u3(-theta, -lam, -phi)[0].crz(-2 * pi * eigval * (2**n))[ctrl, 0].u3(theta, phi, lam)[0]
def qpe_circuit(eigval, theta, phi, lam, precision):
"""固有値、U3のパラメータ、位相推定の精度(何桁求めるか)から、量子位相推定の回路を作成して返します。"""
c = initial_circuit(theta, phi, lam)
c.h[1:1 + precision]
for i in range(precision):
apply_cu(c, i + 1, theta, phi, lam, eigval, i)
iqft(c, 1, precision)
return c
# -
# 試しに、量子回路を見てみましょう。
qpe_circuit(E, theta, phi, lam, 4).run_with_ibmq(returns='draw', output='mpl')
# 続いて、測定結果から固有値を計算する関数を作ります。
def run_qpe(c, shots=1000, max_candidates=5):
"""量子位相推定の回路を実行し、実行結果から固有値の候補を求めます。
shots: 量子回路をrunする際のショット数, max_candidates: 固有値の候補をいくつ返すか?
"""
cnt = c.m[1:].run(shots=shots)
# 測定結果を値に変換する
def to_value(k):
k = k[1:] # 測定結果のうち、最初のビットは無関係なので捨てる
val = 0 # 値
a = 1.0
for ch in k:
if ch == '1':
val += a
a *= 0.5
if val > 1:
# 位相がπを越えたら、2π引いて、マイナスで考える
val = val - 2
return val
return [(to_value(k), v) for k, v in cnt.most_common(max_candidates)]
# いよいよ、量子位相推定を行い、結果を見ていきます。
print('Eigenvalue (expected):', E) # これが答え。これ(に近い値が出てほしい)
# 精度が小さいときと大きいときの違いを見るため、精度を振って動かしてみる。
for precision in range(3, 16):
print(precision, 'bit precision:')
c = qpe_circuit(E, theta, phi, lam, precision)
result = run_qpe(c, 1000, 3)
for value, count in result:
# 1000 shotsのうち、その数が出た回数 得られた固有値 実際の固有値との差分 を表示します。
print(f'{count:<5}{value:<18}(deviation: {value - E: .3e})')
print('')
# 比較的精度よく求まっていることが分かります。
# ## 固有値の和が0にならないエルミート行列の量子位相推定
# 一般には、エルミート行列の固有値は和が0になりません。その場合も基本的には、固有値の和が0の場合と変わらないのですが、Controlled-Uゲートの作り方が少し変わってきます。
#
# $U = e^{2\pi i\hat H}$ の部分に注目します。固有値の和は、行列のトレース(対角和)に相当しましたので、$U$のトレースを考えると、
# $$\begin{eqnarray}
# \mathrm{tr}(U) &=& \mathrm{tr}\left(\sum_{n=0}^{\infty}\frac{(2\pi i \hat H)^n}{n!}\right)\nonumber\\
# &=&\sum_{n=0}^{\infty}\mathrm{tr}\left(\frac{(2\pi i \hat H)^n}{n!}\right)\nonumber\\
# &=&\sum_{n=0}^{\infty}\frac{(2\pi i \mathrm{tr}(\hat H))^n}{n!}\nonumber\\
# &=&e^{2\pi i \mathrm{tr}(\hat H)}\nonumber\\
# \end{eqnarray}$$
# となります。このトレース部分は$U$のグローバル位相として現れます。$U$のグローバル位相を考慮に入れたControlled-Uゲートを作ることで、先ほどと同じように量子位相推定ができます。このようなControlled-Uゲートは、制御ビットにRZゲートを余分に入れることで作ることができます。
def random_hermitian2():
"""ランダムにエルミート行列を作り、
エルミート行列, 固有値(2つ組), パラメータ(3つ組)
を返す。
"""
# 固有値をランダムに決める。先程は固有値の範囲を0〜1にしたが、今回は-1〜1にしてみる。
eigvals = [random.random() * 2 - 1, random.random() * 2 - 1]
# 固有値を小さい順に並べる(この処理は、望まないならコメントアウトしてもよい)
eigvals.sort()
# U3ゲートのパラメータをランダムに決める
theta = random.random()
phi = random.random()
lam = random.random()
# これらから、エルミート行列を作る
u3 = get_u3_matrix(theta, phi, lam)
hermitian = u3 @ np.diag(eigvals) @ u3.T.conjugate()
# エルミート行列であることを確認
assert is_hermitian(hermitian)
# エルミート行列, 固有値, パラメータを返す
return hermitian, eigvals, (theta, phi, lam)
H, eigvals, (theta, phi, lam) = random_hermitian2()
print(H)
# +
def apply_cu2(c, ctrl, theta, phi, lam, eigvals, n):
"""apply_cuを改造し、グローバル位相も考慮に入れられるようにした。
eigvals引数には、固有値の組を入れる。
"""
bias = sum(eigvals) / 2
angle = (eigvals[0] - eigvals[1]) / 2
return c.u3(-theta, -lam, -phi)[0].crz(-2 * pi * angle * (2**n))[ctrl, 0].u3(theta, phi, lam)[0].rz(pi * bias * (2**n))[ctrl]
def qpe_circuit2(eigvals, theta, phi, lam, precision):
"""apply_cu2を使うよう、改造した。"""
c = initial_circuit(theta, phi, lam)
c.h[1:1 + precision]
for i in range(precision):
apply_cu2(c, i + 1, theta, phi, lam, eigvals, i)
iqft(c, 1, precision)
return c
# -
# まず、1番目の固有値を求めます。
print('Eigenvalue (expected):', eigvals[0]) # これが答え。これ(に近い値が出てほしい)
# 精度が小さいときと大きいときの違いを見るため、精度を振って動かしてみる。
for precision in range(3, 16):
print(precision, 'bit precision:')
c = qpe_circuit2(eigvals, theta, phi, lam, precision)
result = run_qpe(c, 1000, 3)
for value, count in result:
# 1000 shotsのうち、その数が出た回数 得られた固有値 実際の固有値との差分 を表示します。
print(f'{count:<5}{value:<18}(deviation: {value - eigvals[0]: .3e})')
print('')
# Controlled-Uゲートを工夫することで、同じように求まりました。
#
# 続いて、もうひとつの固有値を求められないか考えてみましょう。もうひとつの固有値は、もうひとつの固有ベクトルを用意して同じように量子位相推定を行うことで求まります。
#
# 今回、エルミート行列を$\hat H = P D P^\dagger$で書けるようにして、$P$は、U3ゲートを使って$\mathrm{U3}(\theta, \phi, \lambda)$でできるユニタリ行列としました。つまり、$\hat H$の固有ベクトルは、$\mathrm{U3}(\theta, \phi, \lambda)$のユニタリ行列の1列目、2列目ということになります。これまでは、空の量子回路に$\mathrm{U3}(\theta, \phi, \lambda)$ゲートを適用するとユニタリ行列の1列目の固有ベクトルになることを利用して、1つ目の固有ベクトルを用意して、量子位相推定を行いました。2列目の固有ベクトルは$\mathrm{U3}(\theta, \phi, \lambda)$の前に$X$ゲートを付け加えることで得られます。
#
# それをやってみましょう。
# +
def initial_circuit2(theta, phi, lam):
"""2つ目の固有ベクトルを用意する初期回路を作ります。"""
return Circuit().x[0].u3(theta, phi, lam)[0]
def qpe_circuit3(eigvals, theta, phi, lam, precision):
"""apply_cu2とinitial_circuit2を使うよう、改造した。"""
c = initial_circuit2(theta, phi, lam)
c.h[1:1 + precision]
for i in range(precision):
apply_cu2(c, i + 1, theta, phi, lam, eigvals, i)
iqft(c, 1, precision)
return c
# -
print('Eigenvalue (expected):', eigvals[1]) # これが答え。これ(に近い値が出てほしい)
# 精度が小さいときと大きいときの違いを見るため、精度を振って動かしてみる。
for precision in range(3, 16):
print(precision, 'bit precision:')
c = qpe_circuit3(eigvals, theta, phi, lam, precision)
result = run_qpe(c, 1000, 3)
for value, count in result:
# 1000 shotsのうち、その数が出た回数 得られた固有値 実際の固有値との差分 を表示します。
print(f'{count:<5}{value:<18}(deviation: {value - eigvals[1]: .3e})')
print('')
# 2つ目の固有値も求まりました。
| tutorial-ja/114_pea2_ja.ipynb |
# + [markdown] gradient={"editing": false, "id": "e9728fae-bfa6-42b6-80a3-038bb55fc7af", "kernelId": "2f0a71fd-3b47-4853-b105-e7740b79e3ff"}
# # Weights & Biases with Gradient
# + [markdown] gradient={"editing": false, "id": "883566ec-a8d4-4275-8ade-8e6d5400475f", "kernelId": "2f0a71fd-3b47-4853-b105-e7740b79e3ff"}
# # Preface
#
# Weights and Biases is a ML Ops platform that has useful features around model tracking, hyperparameter tuning, and artifact saving during model training steps. Integrating with Weights and Biases provides Gradient users access to world-class model experimenting features while taking advantage of Gradient easy-to-use development platform and access to accelerated hardware.
#
# The goal of this tutorial is to highlight Weights and Biases features and how to use those within Gradient to scale up model training. During this tutorial you will learn to initiate W&B model runs, log metrics, save artifacts, tune hyperparameters, and determine the best performing model. The models trained during this tutorial can be saved in a Gradient Dataset and then be leveraged within Gradient Workflows and Deployments.
# + [markdown] gradient={"editing": false, "id": "eb3ee5bc-0b15-49bd-afaf-c7332018886c", "kernelId": "2f0a71fd-3b47-4853-b105-e7740b79e3ff"}
#
# # Installation and Setup
# + gradient={"editing": false, "execution_count": 2, "id": "2ec09825-b8c8-4e3b-b094-686b93c964e2", "kernelId": "2f0a71fd-3b47-4853-b105-e7740b79e3ff", "source_hidden": false}
# !pip install wandb -q
# + gradient={"editing": false, "execution_count": 3, "id": "9a48fc36-dbab-4d2b-a579-61fc8122dd20", "kernelId": "2f0a71fd-3b47-4853-b105-e7740b79e3ff", "source_hidden": false}
import wandb
import os
os.environ["WANDB_NOTEBOOK_NAME"] = "./train_model_wandb.ipynb"
# + [markdown] gradient={"editing": false, "id": "c585a73f-8953-47d2-94f7-731f1a28204d", "kernelId": "2f0a71fd-3b47-4853-b105-e7740b79e3ff"}
# # Login
# + gradient={"editing": false, "execution_count": 4, "id": "1f154991-d6c5-4544-b949-9ce02a36137d", "kernelId": "2f0a71fd-3b47-4853-b105-e7740b79e3ff", "source_hidden": false}
wandb.login(key='your-api-key')
# + [markdown] gradient={"editing": false, "id": "8e32c911-123d-40a2-a45c-473200b97a13", "kernelId": "2f0a71fd-3b47-4853-b105-e7740b79e3ff"}
# # Initalizing a Model Run and Logging
# + gradient={"editing": false, "execution_count": 5, "id": "aea40ccc-0eb1-4638-8983-1e7fce90a891", "kernelId": "2f0a71fd-3b47-4853-b105-e7740b79e3ff", "source_hidden": false}
config={
"epochs": 5,
"batch_size": 128,
"lr": 1e-3,
"model": 'ResNet18'
}
# + gradient={"editing": false, "execution_count": 6, "id": "9329f15f-6b9f-439f-858f-0f978620df47", "kernelId": "2f0a71fd-3b47-4853-b105-e7740b79e3ff", "source_hidden": false}
import time
import torch.nn as nn
import torch.optim as optim
import torch
from resnet import resnet18, resnet34
from load_data import load_data
# + gradient={"editing": false, "execution_count": 7, "id": "54f772b1-3f33-4daa-ab3d-ef52b054b0ff", "kernelId": "2f0a71fd-3b47-4853-b105-e7740b79e3ff", "source_hidden": false}
def validate_model(model, valid_dl, loss_func, device):
# Compute performance of the model on the validation dataset
model.eval()
val_loss = 0.
with torch.inference_mode():
correct = 0
for i, (images, labels) in enumerate(valid_dl, 0):
# Move data to GPU if available
images, labels = images.to(device), labels.to(device)
# Forward pass
outputs = model(images)
val_loss += loss_func(outputs, labels)*labels.size(0)
# Compute accuracy and accumulate
_, predicted = torch.max(outputs.data, 1)
correct += (predicted == labels).sum().item()
return val_loss / len(valid_dl.dataset), correct / len(valid_dl.dataset)
# + [markdown] gradient={"editing": false, "id": "1caacdd4-c8a7-4c19-968c-117ca42f8656", "kernelId": "2f0a71fd-3b47-4853-b105-e7740b79e3ff"}
# ### Saving a model as a Gradient artifact
# + gradient={"editing": false, "execution_count": 8, "id": "46634cf2-fcd1-4646-924a-dca081339ea0", "kernelId": "2f0a71fd-3b47-4853-b105-e7740b79e3ff", "source_hidden": false}
# !pip install gradient -q
# + gradient={"editing": false, "execution_count": 9, "id": "71136ae9-9df2-4f14-86e2-a95ed2d6123f", "kernelId": "2f0a71fd-3b47-4853-b105-e7740b79e3ff", "source_hidden": false}
from gradient import ModelsClient
model_client = ModelsClient(api_key='your-gradient-api-key')
# + gradient={"editing": true, "id": "092f035c-42a9-404a-b193-f6e01b6a9380", "kernelId": "2f0a71fd-3b47-4853-b105-e7740b79e3ff", "source_hidden": false}
def upload_model(config, model_client, model_dir='models'):
# Create model directory
if not os.path.exists(model_dir):
os.makedirs(model_dir)
# Save model file
params = [config['model'], 'epchs', str(config['epochs']), 'bs', str(config['batch_size']), 'lr', str(round(config['lr'], 6))]
full_model_name = '-'.join(params)
model_path = os.path.join(model_dir, full_model_name + '.pth')
torch.save(model.state_dict(), model_path)
# Upload model as a Gradient artifact
model_client.upload(path=model_path, name=full_model_name, model_type='Custom', project_id='your-project-id')
return full_model_name
# + [markdown] gradient={"editing": false, "id": "49898835-d09b-48fc-8cfb-221af48644b5", "kernelId": "2f0a71fd-3b47-4853-b105-e7740b79e3ff"}
# ### Train ResNet 18
# + gradient={"editing": false, "execution_count": 15, "id": "6df04398-e565-4b08-812b-d85c79046e90", "kernelId": "2f0a71fd-3b47-4853-b105-e7740b79e3ff", "source_hidden": false}
model_name = 'ResNet18'
# Initialize W&B run
with wandb.init(project="test-project", config=config, name=model_name):
# Create Data Loader objects
trainloader, valloader, testloader = load_data(config)
# Create ResNet18 Model with 3 channel inputs (colored image) and 10 output classes
model = resnet18(3, 10)
# Define loss and optimization functions
criterion = nn.CrossEntropyLoss()
optimizer = optim.SGD(model.parameters(), lr=config['lr'], momentum=0.9)
# Move the model to GPU if accessible
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
model.to(device)
step = 0
epoch_durations = []
for epoch in range(config['epochs']):
epoch_start_time = time.time()
batch_checkpoint=50
running_loss = 0.0
model.train()
for i, data in enumerate(trainloader, 0):
# Move data to GPU if available
inputs, labels = data[0].to(device), data[1].to(device)
# Zero the parameter gradients
optimizer.zero_grad()
# Forward + Backward + Optimize
outputs = model(inputs)
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
running_loss += loss.item()
# Log every 50 mini-batches
if i % batch_checkpoint == batch_checkpoint-1: # log every 50 mini-batches
step +=1
print(f'epoch: {epoch + ((i+1)/len(trainloader)):.2f}')
wandb.log({"train_loss": running_loss/batch_checkpoint, "epoch": epoch + ((i+1)/len(trainloader))}, step=step)
print('[%d, %5d] loss: %.3f' %
(epoch + 1, i + 1, running_loss / batch_checkpoint))
running_loss = 0.0
# Log validation metrics
val_loss, accuracy = validate_model(model, valloader, criterion, device)
wandb.log({"val_loss": val_loss, "val_accuracy": accuracy}, step=step)
print(f"Valid Loss: {val_loss:3f}, accuracy: {accuracy:.2f}")
# Log epoch duration
epoch_duration = time.time() - epoch_start_time
wandb.log({"epoch_runtime (seconds)": epoch_duration}, step=step)
epoch_durations.append(epoch_duration)
# Log average epoch duration
avg_epoch_runtime = sum(epoch_durations) / len(epoch_durations)
wandb.log({"avg epoch runtime (seconds)": avg_epoch_runtime})
#Upload model artifact to Gradient and log model name to W&B
full_model_name = upload_model(config, model_client)
wandb.log({"Notes": full_model_name})
print('Training Finished')
# + [markdown] gradient={"editing": false, "id": "ba5ab816-312d-4aa6-b4b5-36c0b6482c58", "kernelId": "2f0a71fd-3b47-4853-b105-e7740b79e3ff"}
# ### Train ResNet34
# + gradient={"editing": false, "execution_count": 1, "id": "120b9fc6-e027-4443-9197-42fe8fe109bf", "kernelId": "2f0a71fd-3b47-4853-b105-e7740b79e3ff", "source_hidden": false}
model_name = 'ResNet34'
config['model'] = model_name
# Initialize W&B run
with wandb.init(project="test-project", config=config, name=model_name):
# Create Data Loader objects
trainloader, valloader, testloader = load_data(config)
# Create ResNet34 Model with 3 channel inputs (colored image) and 10 output classes
model = resnet34(3, 10)
# Define loss and optimization functions
criterion = nn.CrossEntropyLoss()
optimizer = optim.SGD(model.parameters(), lr=config['lr'], momentum=0.9)
# Move the model to GPU if accessible
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
model.to(device)
step = 0
epoch_durations = []
for epoch in range(config['epochs']):
epoch_start_time = time.time()
batch_checkpoint=50
running_loss = 0.0
model.train()
for i, data in enumerate(trainloader, 0):
# Move data to GPU if available
inputs, labels = data[0].to(device), data[1].to(device)
# Zero the parameter gradients
optimizer.zero_grad()
# Forward + Backward + Optimize
outputs = model(inputs)
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
running_loss += loss.item()
# log every 50 mini-batches
if i % batch_checkpoint == batch_checkpoint-1: # log every 50 mini-batches
step +=1
print(f'epoch: {epoch + ((i+1)/len(trainloader)):.2f}')
wandb.log({"train_loss": running_loss/batch_checkpoint, "epoch": epoch + ((i+1)/len(trainloader))}, step=step)
print('[%d, %5d] loss: %.3f' %
(epoch + 1, i + 1, running_loss / batch_checkpoint))
running_loss = 0.0
# Log validation metrics
val_loss, accuracy = validate_model(model, valloader, criterion, device)
wandb.log({"val_loss": val_loss, "val_accuracy": accuracy}, step=step)
print(f"Valid Loss: {val_loss:3f}, accuracy: {accuracy:.2f}")
# Log epoch duration
epoch_duration = time.time() - epoch_start_time
wandb.log({"epoch_runtime (seconds)": epoch_duration}, step=step)
epoch_durations.append(epoch_duration)
# Log average epoch duration
avg_epoch_runtime = sum(epoch_durations) / len(epoch_durations)
wandb.log({"avg epoch runtime (seconds)": avg_epoch_runtime})
#Upload model artifact to Gradient and log model name to W&B
full_model_name = upload_model(config, model_client)
wandb.log({"Notes": full_model_name})
print('Training Finished')
# + [markdown] gradient={"editing": false, "id": "b635c65c-702a-42e6-bdd6-24f2e742d897", "kernelId": "2f0a71fd-3b47-4853-b105-e7740b79e3ff"}
# # Artifacts
# + gradient={"editing": false, "id": "42c1d34a-d8dc-4eac-8b95-6c11417c6c9d", "kernelId": "2f0a71fd-3b47-4853-b105-e7740b79e3ff", "source_hidden": false}
# Classes of images in CIFAR-10 dataset
classes = ('plane', 'car', 'bird', 'cat',
'deer', 'dog', 'frog', 'horse', 'ship', 'truck')
# Initialize W&B run
with wandb.init(project='test-project'):
# Create W&B artifact
artifact = wandb.Artifact('cifar10_image_predictions', type='predictions')
# Create Data Loader objects
trainloader, valloader, testloader = load_data(config)
# Create columns for W&B table
columns=['image', 'label', 'prediction']
for digit in range(10):
columns.append("score_" + classes[digit])
# Create W&B table
pred_table = wandb.Table(columns=columns)
with torch.no_grad():
for i, data in enumerate(testloader, 0):
# Move data to GPU if available
inputs, labels = data[0].to(device), data[1].to(device)
# Calculate model outputs and predictions
outputs = model(inputs)
_, predicted = torch.max(outputs.data, 1)
# Loop through first batch of images and add data to the table
for j, image in enumerate(inputs, 0):
pred_table.add_data(wandb.Image(image), classes[labels[j].item()], classes[predicted[j]], *outputs[j])
break
# Log W&B model artifact
artifact.add(pred_table, "cifar10_predictions")
wandb.log_artifact(artifact)
# + [markdown] gradient={"editing": false, "id": "3b0ff48f-8fe4-4eb1-a683-c74d9e8dc4d6", "kernelId": "2f0a71fd-3b47-4853-b105-e7740b79e3ff"}
# # Sweeps
# + gradient={"editing": false, "id": "91010533-80af-4bb2-abc9-1467e70f6eb5", "kernelId": "2f0a71fd-3b47-4853-b105-e7740b79e3ff", "source_hidden": false}
sweep_config = {
'method': 'bayes',
'metric': {'goal': 'minimize', 'name': 'val_loss'},
'parameters': {
'batch_size': {'values': [32, 128]},
'epochs': {'value': 5},
'lr': {'distribution': 'uniform',
'max': 1e-2,
'min': 1e-4},
'model': {'values': ['ResNet18', 'ResNet34']}
}
}
# + gradient={"editing": false, "id": "3b27ef13-210b-407e-a6f1-2020429d1522", "kernelId": "2f0a71fd-3b47-4853-b105-e7740b79e3ff", "source_hidden": false}
def train(config = None):
with wandb.init(project='test-project', config=config):
config = wandb.config
trainloader, valloader, testloader = load_data(config)
if config['model']=='ResNet18':
model = resnet18(3,10)
else:
model = resnet34(3,10)
criterion = nn.CrossEntropyLoss()
optimizer = optim.SGD(model.parameters(), lr=config['lr'], momentum=0.9)
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
model.to(device)
step = 0
batch_checkpoint=50
epoch_durations = []
for epoch in range(config['epochs']):
epoch_start_time = time.time()
running_loss = 0.0
model.train()
for i, data in enumerate(trainloader, 0):
# Move data to GPU if available
inputs, labels = data[0].to(device), data[1].to(device)
# Zero the parameter gradients
optimizer.zero_grad()
# Forward + Backward + Optimize
outputs = model(inputs)
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
running_loss += loss.item()
# Log every 50 batches
if i % batch_checkpoint == batch_checkpoint-1:
step +=1
print(f'epoch: {epoch + ((i+1)/len(trainloader)):.2f}')
wandb.log({"train_loss": running_loss/batch_checkpoint, "epoch": epoch + ((i+1)/len(trainloader))}, step=step)
print('[%d, %5d] loss: %.3f' %
(epoch + 1, i + 1, running_loss / batch_checkpoint))
running_loss = 0.0
# Log at the end of each epoch
step +=1
print(f'epoch: {epoch + ((i+1)/len(trainloader)):.2f}')
wandb.log({"train_loss": running_loss/batch_checkpoint, "epoch": epoch + ((i+1)/len(trainloader))}, step=step)
print('[%d, %5d] loss: %.3f' %
(epoch + 1, i + 1, running_loss / batch_checkpoint))
# Log validation metrics
val_loss, accuracy = validate_model(model, valloader, criterion, device)
wandb.log({"val_loss": val_loss, "val_accuracy": accuracy}, step=step)
print(f"Valid Loss: {val_loss:3f}, accuracy: {accuracy:.2f}")
epoch_duration = time.time() - epoch_start_time
wandb.log({"epoch_runtime (seconds)": epoch_duration}, step=step)
epoch_durations.append(epoch_duration)
avg_epoch_runtime = sum(epoch_durations) / len(epoch_durations)
wandb.log({"avg epoch runtime (seconds)": avg_epoch_runtime})
#Upload model artifact to Gradient and log model name to W&B
full_model_name = upload_model(config, model_client)
wandb.log({"Notes": full_model_name})
print('Training Finished')
# + gradient={"editing": false, "id": "fc38bc28-6519-4bb5-9ef4-82cc7b1ec7dc", "kernelId": "2f0a71fd-3b47-4853-b105-e7740b79e3ff", "source_hidden": false}
sweep_id = wandb.sweep(sweep_config, project="test-project")
wandb.agent(sweep_id, function=train, count=10)
# + gradient={"editing": false, "id": "1b0b2aad-62f7-4412-9782-ca2f96ca0454", "kernelId": "2f0a71fd-3b47-4853-b105-e7740b79e3ff", "source_hidden": false}
| train_model_wandb.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="Dd-3r8P9TQ_G"
# ## 0. Вещественные числа и погрешности
#
# Вещественные числа в программировании не так просты. Вот, например, посчитаем синус числа $\pi$:
# + id="OBACHtO-TQ_G" colab={"base_uri": "https://localhost:8080/"} outputId="463e9278-9d89-4e38-83ce-95068d2cc0b1"
from math import pi, sin
sin(pi) #думаете, получится 0? Ха-ха!
# + [markdown] id="k3U4SPDETQ_G"
# Непонятный ответ? Во-первых, это так называемая [компьютерная форма экспоненциальной записи чисел.](https://ru.wikipedia.org/wiki/Экспоненциальная_запись#.D0.9A.D0.BE.D0.BC.D0.BF.D1.8C.D1.8E.D1.82.D0.B5.D1.80.D0.BD.D1.8B.D0.B9_.D1.81.D0.BF.D0.BE.D1.81.D0.BE.D0.B1_.D1.8D.D0.BA.D1.81.D0.BF.D0.BE.D0.BD.D0.B5.D0.BD.D1.86.D0.B8.D0.B0.D0.BB.D1.8C.D0.BD.D0.BE.D0.B9_.D0.B7.D0.B0.D0.BF.D0.B8.D1.81.D0.B8) Она удобна, если нужно уметь записывать очень большие или очень маленькие числа:`1.2E2` означает `1.2⋅102`, то есть `1200`, а `2.4e-3` — то же самое, что `2.4⋅10−3=00024`.
#
# Результат, посчитанный Python для $\sin \pi$, имеет порядок `10^(−16)` — это очень маленькое число, близкое к нулю. Почему не «настоящий» ноль? Все вычисления в вещественных числах делаются компьютером с некоторой ограниченной точностью, поэтому зачастую вместо «честных» ответов получаются такие приближенные. К этому надо быть готовым.
# + id="K31XqHzUTQ_G" colab={"base_uri": "https://localhost:8080/"} outputId="172c1453-0720-47d5-c9e9-16ef91d577cb"
0.4 - 0.3 == 0.1
# + id="sMyy3Nq8TQ_H"
a = 0.4 - 0.3
# + id="ij3srYLbTQ_H" colab={"base_uri": "https://localhost:8080/"} outputId="330a3483-a3df-4105-bafa-3978d3c1fa1f"
a
# + id="Cg8mAv3fTQ_H" colab={"base_uri": "https://localhost:8080/"} outputId="71fbf04b-78fc-4d1b-b720-21d49c975761"
a == 0.1
# + id="SKfZl7r7TQ_I" colab={"base_uri": "https://localhost:8080/"} outputId="60f5ea78-bf4e-428b-d4c6-b0c0937b7932"
import sys
a - 0.1 < sys.float_info.epsilon
# + colab={"base_uri": "https://localhost:8080/"} id="GpxaD05Ik0SJ" outputId="40d33f67-25a2-4e42-c624-5224e7f18366"
import sys
sys.float_info.epsilon
# + id="oszKlePpTQ_I" colab={"base_uri": "https://localhost:8080/"} outputId="3d9e4b00-6683-4014-8787-a72e870c1c22"
0.4 - 0.3
# + [markdown] id="3lNhbqzUTQ_I"
# Когда сравниваете вещественные числа -- будьте осторожнее.
#
# Представление вещественного числа:
# + [markdown] id="i5at7BWoLxGk"
# 
# + [markdown] id="HH0DOmryL1tr"
# S -- Знаковый бит
#
# E -- Порядок
#
# M -- Мантисса
#
# B -- Основание (10)
#
# X = (-1)^S * B^E * M
# + [markdown] id="rR7DsI9MMpar"
# 64-битный float (double precision):
# + [markdown] id="X9umx1mGMmjv"
# 
# + [markdown] id="pHnHc9QcTQ_M"
# ## 1. Коллекции: списки, кортежи, range, (строки), множества, словари и др. Понятие последовательности
# + [markdown] id="RF2vHIXgTQ_N"
# ### Строки
#
# Строковые константы можно задавать и с помощью одинарных кавычек, и с помощью двойных, и с помощью тройных:
# + id="cFV-T3PkTQ_N" colab={"base_uri": "https://localhost:8080/"} outputId="dac8cd69-74a3-4a58-b82b-bde2bd559b2c"
s1 = 'ab"c'
s2 = "ab'c"
s3 = """abc
aaaaaa
aaaaa
a
aa
"""
s1 == s2 == s3
# + id="N6yhUHhFTQ_N" colab={"base_uri": "https://localhost:8080/", "height": 153} outputId="57a8175d-8d81-4bf6-fa6d-18721f7589b2"
print(s3)
print()
s3
# + id="e0rrMxLeTQ_O" colab={"base_uri": "https://localhost:8080/", "height": 69} outputId="274488b2-d280-444e-f542-7f457decfcd1"
a = 'a\naaaa'
print(a)
a
# + id="UsJC0wKRTQ_O" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="ec1c2a37-c1ca-4991-f549-04180a57fdb8"
a[3]
# + [markdown] id="yNQ3sZUqVlrG"
# **Внимание:** строки иммутабельны (неизменяемы)!
# + colab={"base_uri": "https://localhost:8080/", "height": 162} id="ZXOY4q7IVeFC" outputId="2b469ceb-794d-4fe1-dc8b-faad7287049b"
a[1] = 'b'
# + [markdown] id="JxnDeKcGTQ_O"
# Функция `len` — получить длину строки (определена для всех коллекций):
# + id="PItdK3IkTQ_P" colab={"base_uri": "https://localhost:8080/"} outputId="dcce39b4-d803-46b8-9cb3-812c4b9120bd"
len("abc")
# + colab={"base_uri": "https://localhost:8080/"} id="B2ejXNTSpufi" outputId="f91fdeaf-0051-4233-ecdb-279e96f9fdd7"
# + [markdown] id="SUXyABVVTQ_P"
# У типа str большое количество встроенных строковых методов:
# * .find — найти подстроку
# * .upper — перевести строку в верхний регистр
# * .lower — перевести строку в нижний регистр
# * .strip — обрезать по краям поданные символы (по умолчанию пробельные)
# * ...
#
# https://docs.python.org/3/library/stdtypes.html#text-sequence-type-str
#
# Все они возвращают новый экземпляр строки!
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="In4EVqKGpJa8" outputId="48e415d0-433e-4642-8748-cc18f45ed037"
s3.replace('a', '10')
# + id="fqYN8DWxTQ_P" colab={"base_uri": "https://localhost:8080/"} outputId="fb9841a9-01b9-4ff7-d072-b9043c32e3d8"
s1.find('b')
# + id="Cgh8Pxa2TQ_P" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="6752be33-6e39-4ea0-c078-bb80053edf88"
s1[-1]
# + id="JaIENCdJTQ_Q" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="ac7d6406-dcdd-4673-e1ba-0d89cdb30a14"
sl = s1.upper()
sl
# + id="1pfwI3PyTQ_R" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="d900e1cb-efa5-4d0d-e4e6-7217fd4a5f54"
s1.upper().lower()
# + id="8U4pWhAhTQ_S" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="42c21d3f-095a-463c-c1e7-06857a22e9f0"
"a ".strip()
# + id="5pZKZ0ciTQ_S"
z = ' z. '
# + id="yU6QnUaXTQ_S" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="ad2cecfe-537a-490e-d814-f03992a4cbaf"
z.strip()
# + id="8ZQTSIdJTQ_T" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="363416ab-cf2a-4fea-aa9d-6b7708326b2e"
z.rstrip()
# + id="MOdP1OB_TQ_T" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="d7480213-9c60-4491-f2d5-effb4c1cd4d6"
z.lstrip()
# + [markdown] id="9fSBFpcCTQ_T"
# Продвинутая индексация: Слайсы
# + id="qd8uP8faWaPP"
s = 'abcde'
# + id="OOSkwN5kTQ_U" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="199a3621-d345-45ff-eba3-3cf04da9728e"
'b' + s[1:]
# + id="DbPRzvskTQ_V" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="8991ca18-84af-425b-a54a-0edbcaaeb137"
s[1:]
# + id="BM5wuNtGTQ_V" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="0cbb49a6-9deb-45f9-94ba-677f92a0c10d"
s[1:-1]
# + id="YzBOr4aOTQ_V" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="1e8b439b-9ed5-4235-c7b6-8213e60a18b3"
s_new = s[2:4]
print(s)
s_new
# + [markdown] id="sxPM6ymATQ_W"
# s[start:finish:step]
#
# * start — начало среза
# * finish — конец среза (не входит в срез!!!)
# * step — шаг
#
# Индексы в питоне могут быть отрицательными! В таком случае считаем с конца (или двигаемся в обратном направлении, если это step)
# + id="kpPkoaRoTQ_W"
s = '0123456789'
# + id="eTyH9cocTQ_W" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="1045c066-36d2-45c2-de19-dcd774553c6d"
s[::2]
# + id="_yDlc6jUTQ_X" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="2d0a3a57-d582-497b-a851-fbfaa144d222"
s[-1:1:-1]
# + id="AV1vo-sJTQ_X" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="ca768bfc-6c7f-407c-e33d-a1546cde9d15"
s[::-2]
# + id="povXyxhBTQ_X" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="5a4f954c-87a1-4d36-ac2e-8446e505733e"
s[::3]
# + [markdown] id="Jbq5g3zQTQ_X"
# Срез равный всей строке:
# + id="Q8DaL9S5TQ_X" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="2539698b-f6c5-4362-86bf-9ce692587cca"
s[0:len(s):1]
# + [markdown] id="ozt9hMs0XhEo"
# Cрез с шагом 2:
# + id="uSELbfiXTQ_Y" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="d28bb9ec-b452-41b1-d48d-026c3b898e47"
s[0:len(s):2]
# + [markdown] id="EF0SeaUvTQ_Y"
# Срез равный одному элементу:
# + id="zJAZKSh1TQ_Y" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="3ed7e371-8c19-4c1c-efc9-59add7dbbe96"
s[5:6:1]
# + [markdown] id="LNW3ujCkYdCS"
# Строки, как мы помним, можно складывать и умножать:
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="d2Bf8UNBYgh8" outputId="c2b37047-ebb9-4e48-e34b-2a6306b73e50"
s_new + s3
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="Mk_cmOfyYmV9" outputId="14005139-9a79-4550-f73b-9698c5e627f1"
s_new * 5
# + [markdown] id="mlwchb0uTQ_Z"
# ### Списки
# + id="2j1Jh-dBTQ_Z"
a = [1, 2, 3, 2, 0]
# + [markdown] id="UIPNButLTQ_Z"
# В список могут входить любые объекты:
# + id="SfW2ugcOTQ_Z" colab={"base_uri": "https://localhost:8080/"} outputId="ba2f2b9f-4044-48d2-d757-5f20cb7567f6"
a = [1, '2', '3', [4, 5]]
a.append(1.2)
a
# + [markdown] id="GS9IYQXWTQ_a"
# Списки тоже поддерживают многие операции!
#
# * cложение списков (+) — конкатенация списков
# * умножение списка на целое число n (*) — повтороение списка n раз
# * слайсы!
# + [markdown] id="xbAw9kukYAxt"
# Всё это вместе называется концепцией последовательности. К "последовательным" типам относятся списки, кортежи (иммутабельные списки), range(), строки (в качестве символьных последовательностей)
#
# Помимо упомянутых операций сложения, умножения, слайсов, измерения длины, поддерживаются операции in (not in), min, max, index (поиск), count
# + id="deDtUNHuTQ_a" colab={"base_uri": "https://localhost:8080/"} outputId="a69d5c0a-2aba-4c8e-b666-99c0406cdbd0"
b = [1, '2']
a + b
# + id="7As9bcWdTQ_a" colab={"base_uri": "https://localhost:8080/"} outputId="00cc2d6f-18f2-4e33-eacc-023ca4b83a2a"
a * 2
# + id="5sk_pB9NTQ_a" colab={"base_uri": "https://localhost:8080/"} outputId="50a98e61-a972-404c-e155-14ff73523a12"
a[::-1]
# + colab={"base_uri": "https://localhost:8080/"} id="MPYnM1evZsl0" outputId="ed23c49c-9c93-4d3c-88fa-7578f810fa4b"
'2' in a + b
# + colab={"base_uri": "https://localhost:8080/"} id="hDlSSXEKZwQr" outputId="133c72be-4024-4b3f-ca50-1dad7edb5eb5"
2 not in a + b
# + colab={"base_uri": "https://localhost:8080/", "height": 162} id="AJqGhrVYZ2OH" outputId="9915ca06-eda9-48cf-b8a0-05293000ddff"
min(a) # работает не всегда!
# + colab={"base_uri": "https://localhost:8080/"} id="Al7ILE7IaGsZ" outputId="840c86b1-cf0d-4eba-c10f-b62223147aea"
max([10, 1000, 1, 2, 5000, 3])
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="y-gx-hysaL9Y" outputId="96a19f14-c404-4014-9c38-7a5daa99d3a8"
min(['1', '2', '3', 'abc', "абв"])
# + colab={"base_uri": "https://localhost:8080/"} id="MzJK7IPEaOi1" outputId="b79effb9-4f29-4e16-e1fe-c9ea892b2892"
a.index([4, 5])
[4, 5] in a
# + colab={"base_uri": "https://localhost:8080/"} id="-u2UCZCNbGQM" outputId="5ed2aa8b-eb6f-41bd-ace6-c8ef0fffde3e"
a.count(2)
# + [markdown] id="GnNs_UuAbJIR"
# Другие коллекции тоже поддерживают многое из этого!
# + [markdown] id="taZ930Q1TQ_b"
# **Внимание:** список — изменяемый объект!
#
# Можно менять элементы списка, можно добавлять новые элементы и удалять старые (`.append` и `.pop`).
# + id="rWorHSr4TQ_b" colab={"base_uri": "https://localhost:8080/"} outputId="7353bb5a-a858-49d8-89a0-bd3a616eeea6"
a[0] = '123'
a
# + id="2emIWM10TQ_b" colab={"base_uri": "https://localhost:8080/"} outputId="a4a525a0-9443-48e5-d227-a461b1d4dcbb"
a.append('data science')
a
# + [markdown] id="rbcPkpo5TQ_b"
# ### Преобразования списков и строк
#
# Два основных строковых метода для взаимодействия списков и строк:
#
# * .split — разделить строку на список строк по определённому сепаратору
# * .join — соеденить список строк в одну большую строку по определённому сепаратору
# + id="_LeWfmEDTQ_b" colab={"base_uri": "https://localhost:8080/"} outputId="a9e6f4b7-0f7c-47b5-a6eb-4d41c06c0989"
s = 'one,two,three'
s.split(',')
# + id="rxH5fsewTQ_b" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="f401eb55-5d15-4da6-9066-ad62fdfe9ca1"
", ".join(['arb', 'borb', 'kork'])
# + [markdown] id="lCtGLsl0b2og"
# ### Другие коллекции
# + [markdown] id="UjhNi0AZb9Vi"
# **Кортежи**: иммутабельные списки
# + colab={"base_uri": "https://localhost:8080/"} id="5ibLcu73b7lc" outputId="677357d7-5ad1-48c0-fac3-2e3fce0637e8"
a = (1, )
a
# + colab={"base_uri": "https://localhost:8080/"} id="8znJhYcvcLH2" outputId="3705e1a1-885b-47f8-b363-445f518792a5"
a = (1, 2, 3)
a
# + colab={"base_uri": "https://localhost:8080/", "height": 162} id="1tVLlpUrxKhB" outputId="6590c540-a46f-4950-990a-e3734429f7e7"
a[0] = 7
# + colab={"base_uri": "https://localhost:8080/"} id="omvDoFu-xKZf" outputId="f94f1a54-ff73-4aae-b208-821717aa34db"
c = ('1', 2, 3)
c
# + [markdown] id="pbimSrJ-cytT"
# **range**: итерируемый объект
# + colab={"base_uri": "https://localhost:8080/"} id="Vy53syFwc4Fo" outputId="87a86a15-dd4e-49fd-8e3a-a9f850ef316f"
a = range(3, 17, 3)
a
# + colab={"base_uri": "https://localhost:8080/"} id="dAyd1DtSdPm9" outputId="724e0b77-1a97-40b6-bd25-4fbb92633d8d"
for x in a:
print(type(x))
# + [markdown] id="-r4xZBuNcSdb"
# **Множество**: коллекция без повторов
# + colab={"base_uri": "https://localhost:8080/"} id="2V2L7HjDcWZ1" outputId="b71f20dc-c02d-4aec-b376-76598b275da8"
a = {1, 2, 3, 3, 2, 4}
a
a.add('1')
a
a.add((1, 2, 3))
a
# + colab={"base_uri": "https://localhost:8080/"} id="1yDfkU8VcjQK" outputId="db254a06-4840-4e29-ca4a-4f473e7e7721"
b = frozenset('aabc')
b
b[2] =
# + [markdown] id="7oX4W6GCctCK"
# **Словари**: набор пар ключ-значение, индексируемый ключами (хеши)
# + colab={"base_uri": "https://localhost:8080/"} id="WFZ2lWSvdpbZ" outputId="912332e0-476f-4fc5-b0f4-846ccdbf42ed"
d = {1: 3, '1': 4, (1, 2, 3): 5, b: 10}
d
# + colab={"base_uri": "https://localhost:8080/"} id="6hef2oHBdz1y" outputId="1e3929c9-facc-41c8-e578-0851819e2f4e"
d[b]
# + colab={"base_uri": "https://localhost:8080/", "height": 162} id="in7ZY65Od290" outputId="e81eb487-8cde-46e6-96c9-5260ba4a54b1"
c = {a: 3}
# + [markdown] id="51XZNUnnTQ_c"
# ## 3. Итерируемся!
#
# В анализе данных нужно много итерироваться по коллекциям!
# + id="_z_AwuTjTQ_c" colab={"base_uri": "https://localhost:8080/"} outputId="a1e8d405-c6cd-40dc-d8a9-eff1b7326a20"
for i in a:
print(i)
# + colab={"base_uri": "https://localhost:8080/"} id="O2HaCRnaeS4y" outputId="26f5b925-2628-46c1-9a3f-0338129c2992"
for i, j in enumerate(a):
print(i, j)
k, b, c = (1, 2, 3)
print(k, b, c)
# + colab={"base_uri": "https://localhost:8080/"} id="t05plRbteWiv" outputId="e9b933f4-ebc7-42b6-a1bb-bc1a38261ab6"
for k, v in d.items():
print(k, v)
# + colab={"base_uri": "https://localhost:8080/", "height": 178} id="Al-dmD4heb4m" outputId="f2e30335-eb0a-4e3d-ae4b-344c43a86eb8"
for k, v in sorted(d.items()):
print(k, v)
# + colab={"base_uri": "https://localhost:8080/"} id="jkA3Hffjei5J" outputId="47740616-f0c0-4e26-e648-0200b93ccec6"
a = [1, 2, 3, 4]
b = [1, 2, 3, 4, 5, 6]
for i in zip(a * 2, b):
print(i)
# + colab={"base_uri": "https://localhost:8080/"} id="8ysUEq0Jey5z" outputId="984bae99-972a-48c1-cf28-0b1a807d4b65"
l = [1, 2, 3, 4, 5, 5]
for i in reversed(l):
print(i)
# + colab={"base_uri": "https://localhost:8080/"} id="njiKvAzGe5d-" outputId="ea1b2a73-58f0-4981-afd8-a0b85bac634b"
for i in sorted(set(reversed(l))):
print(i)
# + id="EEY2LuSQe9pT"
# + [markdown] id="pDl99gFKTQ_d"
# ## 4. Функции
# + id="oxhvu2ZCTQ_d"
def plus(a, b):
return a + b
print(plus)
plus
# + id="e8g3hr4Ffj3x"
locals()
globals()
# + [markdown] id="rUswkPCrgS9H"
# **Замыкание:**
# + id="5IsN8YS1fvjz"
def make_multiplier_of(n):
def multiplier(x):
return x * n
return multiplier
# + id="yR9Iqzg0gVoc"
times3 = make_multiplier_of(3):
print(times3(10))
# + id="rM70O64jgcSc"
times7 = make_multiplier_of(7)
print(times7(5))
# + [markdown] id="9zTUJYrKgj4K"
# **Декораторы:**
# + id="OsDGSgXigmNh"
import math
def cache(function):
CACHE = {}
def decorator(value):
try: return CACHE[value]
except:
result = function(value)
CACHE[value] = result
return result
return decorator
@cache
def is_prime(n):
if n < 2: return False
if n % 2 == 0: return n == 2
limit = int(math.sqrt(n))
for d in range(3, limit+1, 2):
if n % d == 0: return False
return True
print([n for n in range(20+1) if is_prime(n)])
print([n for n in range(20+1) if is_prime(n)])
print([n for n in range(20+1) if is_prime(n)])
| WS02-basics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introdução Data Science - <NAME><img src="https://octocat-generator-assets.githubusercontent.com/my-octocat-1626096942740.png" width="324" height="324" align="right">
# ## Link para download: https://github.com/AnabeatrizMacedo241/DataScience-101
# ## Github: https://github.com/AnabeatrizMacedo241
# ## Linkedin: https://www.linkedin.com/in/ana-beatriz-oliveira-de-macedo-85b05b215/
# 
# ### Nessa oitava parte veremos como utilizar o `SQL` em um caderno Jupyter
import pandas as pd
import sqlite3
pip install ipython-sql
df = pd.DataFrame({'Times': ['Boston Celtics', 'Los Angeles Lakers', 'Golden State Warriors', 'Chicago Bulls',
'San Antonio Spurs', 'Philadelphia 76ers', 'Detroit Pistons', 'Miami Heat',
'New York Knicks', 'Houston Rockets'],
'Títulos': [17, 17, 6, 6, 5, 3, 3, 3, 2, 2],
'Finais': [21, 33, 11, 6, 6, 9, 7, 6, 8, 4],
})
df
connection = sqlite3.connect('data_science101.db')
# +
df.to_sql('nbaTeams', connection) #Parâmetros: (nome_tabela, variável da conexão)
# %load_ext sql
# %sql sqlite:///data_science101.db
# -
# `SELECT` é um comando para retornar informações de um banco de dados e `*` significa que irá retornar tudo.
# + language="sql"
# SELECT * FROM nbaTeams #Retorna nossa tabela em linguagem SQL
# -
# `COUNT()` é usado para contar a quantidade de 'linhas'
# + language="sql"
# SELECT COUNT(*) FROM nbaTeams
# -
# Usamos o `WHERE` para retornar informações específicas. Neste caso, retorna apenas os time com mais de 10 títulos
# + language="sql"
# SELECT * FROM nbaTeams WHERE Títulos>10
# -
# Com os comandos `BETWEEN` e `AND` retornamos informação dentro de um limite. Neste caso, todos os times que participaram entre 5 ou 10 finais
# + language="sql"
# SELECT * FROM nbaTeams WHERE Finais BETWEEN 5 AND 10
# -
# `ORDER BY` retornará alguma coluna em certa ordem. `asc` é ascendente e `desc` seria descendente.
# + language="sql"
# select * from nbaTeams order by Finais asc
# + language="sql"
# select * from nbaTeams order by Finais desc
# -
# Assim como no pandas, SQL também possui alguns métodos de estatística.
# - `min()`retornará o menor número de uma certa coluna
# - `max()` retornará o maior número de uma certa coluna
# - `avg()` retornara a média dessa coluna
# - `sum()` retornará a soma de todos os elementos dessa coluna
# + language="sql"
# select min(Títulos), max(Títulos), avg(Títulos), count(*), sum(Títulos) from nbaTeams
# -
# Se quisermos inserer novas informações podemos usar o comando `INSERT INTO` colocando o nome das colunas e seus valores(`values`) em seguida.
# + language="sql"
# insert into nbaTeams(Times, Títulos, Finais) values("Toronto Raptors", 1, 1)
# + language="sql"
# SELECT * FROM nbaTeams
# -
# ### Até a próxima aula e bons estudos!
# ## <NAME>
# 
| Aulas/Caderno-08(SQL).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
# heterogenous data type
# store the value and datatype for each value
my_list = [1, 'Colorado', 4.8]
my_list
my_list[0]
my_list[2]
my_list[-1]
# homogeneous datatypes
# datatype stored only the 1st time
# faster when using in functions
my_array = np.array([1, 2, 3])
my_array
my_array = np.array((2,3,4))
my_array
my_array[0]
my_list = [1,2,3]
my_array = np.array([1,2,3])
my_array/3
my_list/3
# appended in place
my_list.append('new thing')
my_list
# returns a new array; not in-place
#
my_array = np.append(my_array, 5)
my_array
# +
# dictionaries
# k-v pairs: k - indexes used to find the values
# keys should be unique
my_dict = {'key1': 'first value', 'key2': 'second value'}
my_dict
# -
my_dict['key1']
my_dict['new key'] = 'newest value'
my_dict['new key']
my_dict['int'] = 5
my_dict['int']
# dictionaries imp in creating dataframes and changing dataframes
df = pd.DataFrame({'col1':range(0,3), 'col2':range(3,6)})
df
df.rename(columns = {'col1':'apples', 'col2':'oranges'})
df
| python_basics_lists_arrays_dicts.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import xarray as xr
import iris
import cf_units
import glob
infile_list = glob.glob('/g/data/p66/cm2704/archive/bj594/history/ocn/ocean_month.nc-*')
infile_list.sort()
# ## Iris
cube = iris.load_cube(infile, 'implicit vert diffusion of heat')
cube
coord_names = [coord.name() for coord in cube.dim_coords]
coord_names
cube.coord('tcell zstar depth')
cube.coord('tcell latitude')
# ## xarray
dset = xr.open_mfdataset(infile_list, decode_times=False)
dset
dset['time']
dset['temp'].data.max()
dset['geolat_t']
# ## New cube
ref_cube = iris.load_cube('/g/data/fs38/publications/CMIP6/CMIP/CSIRO-ARCCSS/ACCESS-CM2/historical/r1i1p1f1/Omon/thetao/gn/v20191108/thetao_Omon_ACCESS-CM2_historical_r1i1p1f1_gn_185001-185912.nc', 'sea_water_potential_temperature')
ref_cube
ref_cube.shape
dset['temp_eta_smooth'].shape
ref_cube.dim_coords
time_coord = iris.coords.DimCoord(dset['time'].values,
standard_name='time',
long_name='time',
var_name='time',
units=cf_units.Unit(dset['time'].units, calendar=dset['time'].calendar.lower()))
new_standard_name = dset['temp_eta_smooth'].long_name.replace(' ', '_')
iris.std_names.STD_NAMES[new_standard_name] = {'canonical_units': 'W m-2'}
new_cube = iris.cube.Cube(dset['temp_eta_smooth'].to_masked_array(),
standard_name=new_standard_name,
long_name=dset['temp_eta_smooth'].long_name,
var_name=dset['temp_eta_smooth'].name,
units='W m-2',
attributes=ref_cube.attributes,
dim_coords_and_dims=[(time_coord, 0),
(ref_cube.dim_coords[-2], 1),
(ref_cube.dim_coords[-1], 2)],
aux_coords_and_dims=[(ref_cube.aux_coords[0], (1, 2)),
(ref_cube.aux_coords[1], (1, 2))])
new_cube
dset['temp_eta_smooth'].coords
dset['geolat_t'].values.shape
ref_cube.coord('latitude').points.shape
dset['geolat_t'].values[10, 10]
ref_cube.coord('latitude').points[10, 10]
import numpy as np
np.allclose(ref_cube.coord('latitude').points, dset['geolat_t'].values)
dset['temp_eta_smooth'].ndim
ref_cube.ndim
dset['geolon_t'].values + 360
ref_cube.coord('longitude').points.max()
ref_cube.coord('longitude').points.min()
dset['geolon_t'].values.max()
dset['geolon_t'].values.min()
test_lons = np.where(dset['geolon_t'].values < 0.0, dset['geolon_t'].values + 360, dset['geolon_t'].values)
test_lons
ref_cube.coord('longitude').points
# ## Temperature check
cmip_file = '/g/data/fs38/publications/CMIP6/CMIP/CSIRO-ARCCSS/ACCESS-CM2/historical/r1i1p1f1/Omon/thetao/gn/v20191108/thetao_Omon_ACCESS-CM2_historical_r1i1p1f1_gn_185001-185912.nc'
mom_file = '/g/data/r87/dbi599/ACCESS-CM2-MOM5/thetao_Omon_ACCESS-CM2-MOM5_historical_r1i1p1f1_gn_185001-185912.nc'
cmip_cube = iris.load_cube(cmip_file, 'sea_water_potential_temperature')
mom_cube = iris.load_cube(mom_file, 'sea_water_potential_temperature')
cmip_cube[10, ::].data.mean()
mom_cube[10, ::].data.mean() + 273.15
mom_cube.data[0, 0, 50:55, 10:15] + 273.15
cmip_cube.data[0, 0, 50:55, 10:15]
import iris
import iris.plot as iplt
import matplotlib.pyplot as plt
test_file = '/g/data/r87/dbi599/CMIP6/CMIP/CSIRO-ARCCSS/ACCESS-CM2/historical/r1i1p1f1/Omon/sw-heat/gn/v20191108/sw-heat_Omon_ACCESS-CM2_historical_r1i1p1f1_gn_185001-185912.nc'
cube = iris.load(test_file)
cube = cube[0]
vmin = -100
vmax = 100
#iplt.pcolormesh(plot_cube, cmap='RdBu_r', vmin=vmin, vmax=vmax)
plt.pcolormesh(cube[0, 1, ::].data, cmap='RdBu_r', vmin=vmin, vmax=vmax)
cb = plt.colorbar()
cb.set_label(str(cube[0, 0, ::].units))
plt.show()
for level in range(50):
print(cube[0, level, ::].data.sum())
cube[0, 1, ::].data.sum()
cube[0, 2, ::].data.sum()
cube[0, 0, 30:40, :].data
| development/mom_data.ipynb |
from bokeh.models import ColumnDataSource, DataRange1d, Plot, LinearAxis, Grid, GlyphRenderer, Circle, HoverTool, BoxSelectTool
from bokeh.models.widgets import VBox, DataTable, TableColumn, StringFormatter, NumberFormatter, StringEditor, IntEditor, NumberEditor, SelectEditor
from bokeh.sampledata.autompg2 import autompg2 as mpg
source = ColumnDataSource(mpg)
manufacturers = sorted(mpg["manufacturer"].unique())
models = sorted(mpg["model"].unique())
transmissions = sorted(mpg["trans"].unique())
drives = sorted(mpg["drv"].unique())
classes = sorted(mpg["class"].unique())
columns = [
TableColumn(field="manufacturer", title="Manufacturer", editor=SelectEditor(options=manufacturers), formatter=StringFormatter(font_style="bold")),
TableColumn(field="model", title="Model", editor=StringEditor(completions=models)),
TableColumn(field="displ", title="Displacement", editor=NumberEditor(step=0.1), formatter=NumberFormatter(format="0.0")),
TableColumn(field="year", title="Year", editor=IntEditor()),
TableColumn(field="cyl", title="Cylinders", editor=IntEditor()),
TableColumn(field="trans", title="Transmission", editor=SelectEditor(options=transmissions)),
TableColumn(field="drv", title="Drive", editor=SelectEditor(options=drives)),
TableColumn(field="class", title="Class", editor=SelectEditor(options=classes)),
TableColumn(field="cty", title="City MPG", editor=IntEditor()),
TableColumn(field="hwy", title="Highway MPG", editor=IntEditor()),
]
data_table = DataTable(source=source, columns=columns, editable=True)
xdr = DataRange1d(sources=[source.columns("index")])
ydr = DataRange1d(sources=[source.columns("cty"), source.columns("hwy")])
plot = Plot(title=None, x_range=xdr, y_range=ydr, plot_width=1000, plot_height=300)
xaxis = LinearAxis(plot=plot)
plot.below.append(xaxis)
yaxis = LinearAxis(plot=plot)
ygrid = Grid(plot=plot, dimension=1, ticker=yaxis.ticker)
plot.left.append(yaxis)
cty_glyph = Circle(x="index", y="cty", fill_color="#396285", size=8, fill_alpha=0.5, line_alpha=0.5)
hwy_glyph = Circle(x="index", y="hwy", fill_color="#CE603D", size=8, fill_alpha=0.5, line_alpha=0.5)
cty = GlyphRenderer(data_source=source, glyph=cty_glyph)
hwy = GlyphRenderer(data_source=source, glyph=hwy_glyph)
tooltips = [
("Manufacturer", "@manufacturer"),
("Model", "@model"),
("Displacement", "@displ"),
("Year", "@year"),
("Cylinders", "@cyl"),
("Transmission", "@trans"),
("Drive", "@drv"),
("Class", "@class"),
]
cty_hover_tool = HoverTool(plot=plot, renderers=[cty], tooltips=tooltips + [("City MPG", "@cty")])
hwy_hover_tool = HoverTool(plot=plot, renderers=[hwy], tooltips=tooltips + [("Highway MPG", "@hwy")])
select_tool = BoxSelectTool(plot=plot, renderers=[cty, hwy], dimensions=['width'])
plot.tools.extend([cty_hover_tool, hwy_hover_tool, select_tool])
plot.renderers.extend([cty, hwy, ygrid])
VBox(children=[plot, data_table]).html
| examples/glyphs/data_tables.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: nlpbook
# language: python
# name: nlpbook
# ---
# +
import os
from argparse import Namespace
import collections
import nltk.data
import numpy as np
import pandas as pd
import re
import string
from tqdm import tqdm_notebook
# -
args = Namespace(
raw_dataset_txt="data/books/frankenstein.txt",
window_size=5,
train_proportion=0.7,
val_proportion=0.15,
test_proportion=0.15,
output_munged_csv="data/books/frankenstein_with_splits.csv",
seed=1337
)
# Split the raw text book into sentences
tokenizer = nltk.data.load('tokenizers/punkt/english.pickle')
with open(args.raw_dataset_txt) as fp:
book = fp.read()
sentences = tokenizer.tokenize(book)
print (len(sentences), "sentences")
print ("Sample:", sentences[100])
# Clean sentences
def preprocess_text(text):
text = ' '.join(word.lower() for word in text.split(" "))
text = re.sub(r"([.,!?])", r" \1 ", text)
text = re.sub(r"[^a-zA-Z.,!?]+", r" ", text)
return text
cleaned_sentences = [preprocess_text(sentence) for sentence in sentences]
# Global vars
MASK_TOKEN = "<MASK>"
# +
# Create windows
flatten = lambda outer_list: [item for inner_list in outer_list for item in inner_list]
windows = flatten([list(nltk.ngrams([MASK_TOKEN] * args.window_size + sentence.split(' ') + \
[MASK_TOKEN] * args.window_size, args.window_size * 2 + 1)) \
for sentence in tqdm_notebook(cleaned_sentences)])
# Create cbow data
data = []
for window in tqdm_notebook(windows):
target_token = window[args.window_size]
context = []
for i, token in enumerate(window):
if token == MASK_TOKEN or i == args.window_size:
continue
else:
context.append(token)
data.append([' '.join(token for token in context), target_token])
# Convert to dataframe
cbow_data = pd.DataFrame(data, columns=["context", "target"])
# -
# Create split data
n = len(cbow_data)
def get_split(row_num):
if row_num <= n*args.train_proportion:
return 'train'
elif (row_num > n*args.train_proportion) and (row_num <= n*args.train_proportion + n*args.val_proportion):
return 'val'
else:
return 'test'
cbow_data['split']= cbow_data.apply(lambda row: get_split(row.name), axis=1)
cbow_data.head()
# Write split data to file
cbow_data.to_csv(args.output_munged_csv, index=False)
| chapters/chapter_5/5_2_CBOW/5_2_munging_frankenstein.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.7 64-bit (''flaml'': conda)'
# metadata:
# interpreter:
# hash: bfcd9a6a9254a5e160761a1fd7a9e444f011592c6770d9f4180dde058a9df5dd
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# Copyright (c) 2020-2021 Microsoft Corporation. All rights reserved.
#
# Licensed under the MIT License.
#
# # Tune XGBoost with FLAML Library
#
#
# ## 1. Introduction
#
# FLAML is a Python library (https://github.com/microsoft/FLAML) designed to automatically produce accurate machine learning models
# with low computational cost. It is fast and cheap. The simple and lightweight design makes it easy
# to use and extend, such as adding new learners. FLAML can
# - serve as an economical AutoML engine,
# - be used as a fast hyperparameter tuning tool, or
# - be embedded in self-tuning software that requires low latency & resource in repetitive
# tuning tasks.
#
# In this notebook, we demonstrate how to use FLAML library to tune hyperparameters of XGBoost with a regression example.
#
# FLAML requires `Python>=3.6`. To run this notebook example, please install flaml with the `notebook` option:
# ```bash
# pip install flaml[notebook]
# ```
# -
# !pip install flaml[notebook];
# + [markdown] slideshow={"slide_type": "slide"}
# ## 2. Regression Example
# ### Load data and preprocess
#
# Download [houses dataset](https://www.openml.org/d/537) from OpenML. The task is to predict median price of the house in the region based on demographic composition and a state of housing market in the region.
# + slideshow={"slide_type": "subslide"} tags=[]
from flaml.data import load_openml_dataset
X_train, X_test, y_train, y_test = load_openml_dataset(dataset_id = 537, data_dir = './')
# + [markdown] slideshow={"slide_type": "slide"}
# ### Run FLAML
# In the FLAML automl run configuration, users can specify the task type, time budget, error metric, learner list, whether to subsample, resampling strategy type, and so on. All these arguments have default values which will be used if users do not provide them.
# + slideshow={"slide_type": "slide"}
''' import AutoML class from flaml package '''
from flaml import AutoML
automl = AutoML()
# + slideshow={"slide_type": "slide"}
settings = {
"time_budget": 60, # total running time in seconds
"metric": 'r2', # primary metrics for regression can be chosen from: ['mae','mse','r2']
"estimator_list": ['xgboost'], # list of ML learners; we tune xgboost in this example
"task": 'regression', # task type
"log_file_name": 'houses_experiment.log', # flaml log file
}
# + slideshow={"slide_type": "slide"} tags=[]
'''The main flaml automl API'''
automl.fit(X_train = X_train, y_train = y_train, **settings)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Best model and metric
# + slideshow={"slide_type": "slide"} tags=[]
''' retrieve best config'''
print('Best hyperparmeter config:', automl.best_config)
print('Best r2 on validation data: {0:.4g}'.format(1-automl.best_loss))
print('Training duration of best run: {0:.4g} s'.format(automl.best_config_train_time))
# + slideshow={"slide_type": "slide"}
automl.model
# + slideshow={"slide_type": "slide"}
''' pickle and save the best model '''
import pickle
with open('best_model.pkl', 'wb') as f:
pickle.dump(automl.model, f, pickle.HIGHEST_PROTOCOL)
# + slideshow={"slide_type": "slide"} tags=[]
''' compute predictions of testing dataset '''
y_pred = automl.predict(X_test)
print('Predicted labels', y_pred)
print('True labels', y_test)
# + slideshow={"slide_type": "slide"} tags=[]
''' compute different metric values on testing dataset'''
from flaml.ml import sklearn_metric_loss_score
print('r2', '=', 1 - sklearn_metric_loss_score('r2', y_pred, y_test))
print('mse', '=', sklearn_metric_loss_score('mse', y_pred, y_test))
print('mae', '=', sklearn_metric_loss_score('mae', y_pred, y_test))
# + slideshow={"slide_type": "subslide"} tags=[]
from flaml.data import get_output_from_log
time_history, best_valid_loss_history, valid_loss_history, config_history, train_loss_history = \
get_output_from_log(filename = settings['log_file_name'], time_budget = 60)
for config in config_history:
print(config)
# + slideshow={"slide_type": "slide"}
import matplotlib.pyplot as plt
import numpy as np
plt.title('Learning Curve')
plt.xlabel('Wall Clock Time (s)')
plt.ylabel('Validation r2')
plt.scatter(time_history, 1-np.array(valid_loss_history))
plt.step(time_history, 1-np.array(best_valid_loss_history), where='post')
plt.show()
# -
# ## 3. Comparison with untuned XGBoost
#
# ### FLAML's accuracy
print('flaml (60s) r2', '=', 1 - sklearn_metric_loss_score('r2', y_pred, y_test))
# ### Default XGBoost
from xgboost import XGBRegressor
xgb = XGBRegressor()
xgb.fit(X_train, y_train)
y_pred = xgb.predict(X_test)
from flaml.ml import sklearn_metric_loss_score
print('default xgboost r2', '=', 1 - sklearn_metric_loss_score('r2', y_pred, y_test))
| notebook/flaml_xgboost.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Assign place name and county information to zip-level data. Add county-level cases, deaths, caseRate, and deathRate for specific days and weeks for waves 1 - 3.
#
# **[Work in progress]**
#
# Author: <NAME> (<EMAIL>)
import os
import pandas as pd
from pathlib import Path
from py2neo import Graph
import time
import unidecode
import difflib
from functools import reduce
pd.options.display.max_rows = None # display all rows
pd.options.display.max_columns = None # display all columsns
# #### Connect to COVID-19-Net Knowledge Graph
graph = Graph("bolt://172.16.58.3:7687", user="reader", password="<PASSWORD>")
df = pd.read_csv("COVID_waves1to3_zipcodefocused.csv", dtype='str')
df.head()
# Zip codes must be 5 digits, pad 4 digit zip code with zeros
df['S1_ZipCode'] = df['S1_ZipCode'].apply(lambda x: x if len(x) != 4 else '0' + x)
df.shape
df_zip = df.query("S1_ZipCode != ' '").copy()
df_zip['S1_ZipCode'] = df_zip['S1_ZipCode'].astype(int)
zip_unique = df_zip['S1_ZipCode'].unique()
len(zip_unique)
zip_unique
df_zip.info()
df.drop_duplicates(inplace=True)
df.shape
query = """
OPTIONAL MATCH (p:PostalCode{name:$zip_code})-[i:IN]->(a2:Admin2)-[:IN]->(a1:Admin1),
(a2)-[:HAS_DEMOGRAPHICS]->(d:Demographics{aggregationLevel: 'Admin2'})
RETURN a1.name AS state, a2.name AS county, i.resRatio AS resRatio, p.placeName AS placeName, a1.code AS code,
$zip_code AS S1_ZipCode, d.totalPopulation as population
"""
loc = pd.concat((graph.run(query, zip_code=row.S1_ZipCode).to_data_frame()
for row in df.itertuples()))
loc.head(10)
df_loc = df.merge(loc, on='S1_ZipCode', how='left')
df_loc.shape
df_loc.head()
df_loc.to_csv("COVID_waves1to3_zipcodefocused_Locations.csv", index=False)
query = """
MATCH (p:PostalCode{name:$zip_code})-[i:IN]->(a2:Admin2)-[:IN]->(a1:Admin1),
(a2)<-[:REPORTED_IN]-(c:Cases{source:'JHU'}),
(a2)-[:HAS_DEMOGRAPHICS]->(d:Demographics{aggregationLevel: 'Admin2'})
WHERE c.date = date($date)
RETURN $date AS date, a1.name AS state, a2.name AS county, i.resRatio AS resRatio, p.placeName AS placeName, a1.code AS code,
$zip_code AS S1_ZipCode,
d.totalPopulation as population, c.cases AS cases, c.deaths AS deaths,
c.cases*100000.0/d.totalPopulation AS caseRate,
c.deaths*100000.0/d.totalPopulation AS deathRate
"""
zip_code = '32754'
graph.run(query, zip_code=zip_code, date='2020-04-27').to_data_frame()
# ### Data for 2020-02-28
data_20200228 = pd.concat((graph.run(query, zip_code=row.S1_ZipCode, date='2020-02-28').to_data_frame()
for row in df.itertuples()))
data_20200228.rename(columns={'cases': 'cases_2020-02-28', 'deaths': 'death_2020-02-28',
'caseRate': 'caseRate_2020-02-28', 'deathRate': 'deathRate_2020-02-28'}, inplace=True)
data_20200228.drop(['date'], axis=1, inplace=True)
data_20200228.drop_duplicates(inplace=True)
data_20200228.shape
data_20200228.head()
# ### Data for 2020-04-27
data_20200427 = pd.concat((graph.run(query, zip_code=row.S1_ZipCode, date='2020-04-27').to_data_frame()
for row in df.itertuples()))
data_20200427.rename(columns={'cases': 'cases_2020-04-27', 'deaths': 'death_2020-04-27',
'caseRate': 'caseRate_2020-04-27', 'deathRate': 'deathRate_2020-04-27'}, inplace=True)
data_20200427.drop(['date'], axis=1, inplace=True)
data_20200427.drop_duplicates(inplace=True)
data_20200427.shape
data_20200427.head()
# ### Data for 2020-08-05
data_20200805 = pd.concat((graph.run(query, zip_code=row.S1_ZipCode, date='2020-08-05').to_data_frame()
for row in df.itertuples()))
data_20200805.rename(columns={'cases': 'cases_2020-08-05', 'deaths': 'death_2020-08-05',
'caseRate': 'caseRate_2020-08-05', 'deathRate': 'deathRate_2020-08-05'}, inplace=True)
data_20200805.drop(['date'], axis=1, inplace=True)
data_20200805.drop_duplicates(inplace=True)
data_20200805.shape
data_20200805.head()
# ### Get 7-day averages
# +
# Wave 1: Feb. 21-27 (pre-launch) versus Jan 24-Jan 30 (comparison baseline)
# Wave 2: Apr 20-26 versus Mar 23-29
# Wave 3: July 29-Aug 4 versus June 30-July 6
# -
query = """
MATCH (p:PostalCode{name:$zip_code})-[i:IN]->(a2:Admin2)-[:IN]->(a1:Admin1),
(a2)<-[:REPORTED_IN]-(c:Cases{source:'JHU'}),
(a2)-[:HAS_DEMOGRAPHICS]->(d:Demographics{aggregationLevel: 'Admin2'})
WHERE c.date >= date($startDate) AND c.date <= date($endDate)
RETURN a1.name AS state, a2.name AS county, i.resRatio AS resRatio, p.placeName AS placeName, a1.code AS code,
$zip_code AS S1_ZipCode,
d.totalPopulation as population, avg(c.cases) AS cases, avg(c.deaths) AS deaths,
avg(c.cases)*100000.0/d.totalPopulation AS caseRate,
avg(c.deaths)*100000.0/d.totalPopulation AS deathRate
"""
data_20200124_20200130 = pd.concat((graph.run(query, zip_code=row.S1_ZipCode,
startDate='2020-01-24', endDate='2020-01-30').to_data_frame()
for row in df.itertuples()))
data_20200124_20200130.rename(columns={'cases': 'avgcases_2020-01-24_2020-01-30', 'deaths': 'avgdeath_2020-01-24_2020-01-30',
'caseRate': 'avgcaseRate_2020-01-24_2020-01-30', 'deathRate': 'avgdeathRate_2020-01-24_2020-01-30'}, inplace=True)
data_20200124_20200130.drop_duplicates(inplace=True)
data_20200124_20200130.shape
data_20200221_20200227 = pd.concat((graph.run(query, zip_code=row.S1_ZipCode,
startDate='2020-02-21', endDate='2020-02-27').to_data_frame()
for row in df.itertuples()))
data_20200221_20200227.rename(columns={'cases': 'avgcases_2020-02-21_2020-02-27', 'deaths': 'avgdeath_2020-02-21_2020-02-27',
'caseRate': 'avgcaseRate_2020-02-21_2020-02-27', 'deathRate': 'avgdeathRate_2020-02-21_2020-02-27'}, inplace=True)
data_20200221_20200227.drop_duplicates(inplace=True)
data_20200221_20200227.shape
data_20200323_20200329 = pd.concat((graph.run(query, zip_code=row.S1_ZipCode,
startDate='2020-03-23', endDate='2020-03-29').to_data_frame()
for row in df.itertuples()))
data_20200323_20200329.rename(columns={'cases': 'avgcases_2020-03-23_2020-03-29', 'deaths': 'avgdeath_2020-03-23_2020-03-29',
'caseRate': 'avgcaseRate_2020-03-23_2020-03-29', 'deathRate': 'avgdeathRate_2020-03-23_2020-03-29'}, inplace=True)
data_20200323_20200329.drop_duplicates(inplace=True)
data_20200323_20200329.shape
data_20200420_20200426 = pd.concat((graph.run(query, zip_code=row.S1_ZipCode,
startDate='2020-04-20', endDate='2020-04-26').to_data_frame()
for row in df.itertuples()))
data_20200420_20200426.rename(columns={'cases': 'avgcases_2020-04-20_2020-04-26', 'deaths': 'avgdeath_2020-04-20_2020-04-26',
'caseRate': 'avgcaseRate_2020-04-20_2020-04-26', 'deathRate': 'avgdeathRate_2020-04-20_2020-04-26'}, inplace=True)
data_20200420_20200426.drop_duplicates(inplace=True)
data_20200420_20200426.shape
data_20200630_20200706 = pd.concat((graph.run(query, zip_code=row.S1_ZipCode,
startDate='2020-06-30', endDate='2020-07-06').to_data_frame()
for row in df.itertuples()))
data_20200630_20200706.rename(columns={'cases': 'avgcases_2020-06-30_2020-07-06', 'deaths': 'avgdeath_2020-04-20_2020-07-06',
'caseRate': 'avgcaseRate_2020-06-30_2020-07-06', 'deathRate': 'avgdeathRate_2020-06-30_2020-07-06'}, inplace=True)
data_20200630_20200706.drop_duplicates(inplace=True)
data_20200630_20200706.shape
data_20200729_20200804 = pd.concat((graph.run(query, zip_code=row.S1_ZipCode,
startDate='2020-07-29', endDate='2020-08-04').to_data_frame()
for row in df.itertuples()))
data_20200729_20200804.rename(columns={'cases': 'avgcases_2020-07-29_2020-08-04', 'deaths': 'avgdeath_2020-07-29_2020-08-04',
'caseRate': 'avgcaseRate_2020-07-29_2020-08-04', 'deathRate': 'avgdeathRate_2020-07-29_2020-08-04'}, inplace=True)
data_20200729_20200804.drop_duplicates(inplace=True)
data_20200729_20200804.shape
data_20200729_20200804.query("county == 'Los Angeles County'").head(20)
# ### Merge dataframes
data_frames = [data_20200228, data_20200427, data_20200805, data_20200124_20200130, data_20200221_20200227,
data_20200323_20200329, data_20200420_20200426, data_20200630_20200706, data_20200729_20200804]
data = reduce(lambda left,right: pd.merge(left,right,on=['S1_ZipCode','state','county', 'resRatio', 'placeName','code', 'population'], how='outer'), data_frames).fillna(0.0)
data.shape
data.head()
# ### Merge
df_loc_cases = df_loc.merge(data, on=['S1_ZipCode', 'state', 'county', 'resRatio', 'placeName', 'resRatio', 'code', 'population'], how='left')
df_loc_cases.shape
df_loc_cases.head()
df_loc_cases.shape
df_loc_cases.drop_duplicates(inplace=True)
df_loc_cases.shape
df_loc_cases.to_csv("COVID_waves1to3_zipcodefocused_JHU_20201003.csv", index=False)
df_loc_cases.query("S1_ZipCode == '92301'")
| notebooks/dev/Waves1-3-Zip.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ipycytoscape from scratch. Part 2.
# ### The Train rail net grows up.
# ### Objective
# This jupyter is the continuation of part one:
# Ipycytoscape from Scratch.ipynb
#
# The goal of this article is to learn graphs from scratch using ipycytoscape. In order to get into graphs I had made the point that it is better starting by visualizing because an image is worth a million words and also a million equations.
# A small graph was built in the first part, and we made little by little some modifications to the graph showing how modify the graph using ipycytoscape.
#
# ### For whom is this article/notebook? What do you need to know?
# You code in python but somehow you had never had anything to do with graphs.
# You have read the first part notebook.
# You are familiar with dictionaries and JSON structures.
# You have some knowledge of pandas. (If you don't have, you might skip the last part).
#
# ### Our project
# You have to imagine that you are a coder that is ultimately in charge of creating a GUI that shows the German rail net in an interactive way showing different kinds of information for the operators to be able to take decisions.
#
# ### Some notes:
# As a coder after having read the first article you are at a point where you might be facing the following situations:
# - A friend told you that visualizing graphs in notebooks is not the way to go: I assure you that you will be able to tell him "voilà". ("here you are" in French)
# - Another friend might have told you that in order to really work in python with graphs you better start digging into Networkx. Well you might tell him that a pic is worth a thousand words, and for what it takes, a pic is worth a thousand equations. You will be able to jump from visualizing with ipycytoscape to Networkx, better than the other way around, that is my point here.
#
# ### That said lets see where we stand and move forward.
# We have built a 'mini' German rail net with rail connections between 6 cities. BER (Berlin), MUN(Munich), HAM (Hamburg), FRA (Frankfurt) and LEP(LEIPZIG). That resulted in a graph.
# We added also some information to the graphs.
# Note: I will tend to paste here all the code AND DATA necessary in order to avoid you to have to pick it up in GitHub or read it from disc.
#
# What comes next is the situation we had in part one.
# Lets go
# +
import ipycytoscape
import json
import ipywidgets
# I paste here allthe data in order to allow you to copy and paste it without loading data from files.
railnet= '''{
"nodes": [
{"data": { "id": "BER", "label":"HBf BER", "classes":"east"}},
{"data": { "id": "MUN", "label":"HBf MUN", "classes":"west"}},
{"data": { "id": "FRA", "label":"HBf FRA", "classes":"west"}},
{"data": { "id": "HAM", "label":"HBf HAM", "classes":"west"}},
{"data": { "id": "LEP", "label":"HBf LEP", "classes":"east"}}
],
"edges": [
{"data": { "id": "line1", "source": "BER", "target": "MUN","label":"200km/h"}},
{"data": { "id": "line2", "source": "MUN", "target": "FRA","label":"200km/h"}},
{"data": { "id": "line3", "source": "FRA", "target": "BER","label":"250km/h" }},
{"data": { "id": "line4", "source": "BER", "target": "HAM","label":"300km/h" }},
{"data": { "id": "line5", "source": "BER", "target": "LEP","label":"300km/h" }}
]
}'''
train_style = [
{'selector': 'node','style': {
'font-family': 'arial',
'font-size': '10px',
'label': 'data(label)',}},
{'selector': 'node[classes="east"]','style': {
'background-color': 'yellow'}},
{'selector': 'node[classes="west"]','style': {
'background-color': 'blue'}},
{'selector': 'node[id = "BER"]','style': {
'font-family': 'arial',
'font-size': '10px',
'label': 'data(label)',
'background-color': 'green'}},
{'selector': 'edge[id = "line1"]','style': {
'font-family': 'arial',
'font-size': '10px',
'label': 'data(label)'}},
{'selector': 'edge[id = "line2"]','style': {
'font-family': 'arial',
'font-size': '10px',
'label': 'data(label)'}},
{'selector': 'edge[id = "line3"]','style': {
'font-family': 'arial',
'font-size': '10px',
'label': 'data(label)'}},
{'selector': 'edge[id = "line4"]','style': {
'font-family': 'arial',
'font-size': '10px',
'label': 'data(label)'}},
{'selector': 'edge[id = "line5"]','style': {
'font-family': 'arial',
'font-size': '10px',
'label': 'data(label)'}}
]
railnetJSON = json.loads(railnet)
ipycytoscape_obj = ipycytoscape.CytoscapeWidget()
ipycytoscape_obj.graph.add_graph_from_json(railnetJSON)
ipycytoscape_obj.set_style(train_style)
ipycytoscape_obj
# -
# ## Adding nodes and edges.
#
# Nuremberg is a city in former West Germany. A new station was built in Nuremberg (NUR) and it has to be added to the train rail net. Nuremberg's station is joined to the net in Leipzig and Frankfurt. And the connections between NUR and the net is done only with regional rail connections. How do we add that to the net?
# A new node and a new edge have to be added. In this case I don't built a new graph from scratch in order to show that adding the nodes and edges will happened in the already visualized graph when you run the code.
# Be aware that the rail station in Nuremberg is still being built, so we want to reflect that on the graph. The desired label is "HBf NUR in construction"
# +
station_NUR = ipycytoscape.Node()
station_NUR.data['id'] = "NUR"
station_NUR.data['label'] = "HBf NUR in construction"
station_NUR.data['classes'] = "west"
ipycytoscape_obj.graph.add_node(station_NUR)
ipycytoscape_obj
# -
# Ups! The station is built but the rail connections are missing.
# Let's add the rail connections (edges).
# The info that we got is that there will be two connections built as follows:
# - connection NUR-LEP will be a regional train running only at 150km/h.
# - connection NUR-FRA will be a regional train running only at 150km/h.
# +
# rail connection NUR-LEP
new_edge1 = ipycytoscape.Edge()
new_edge1.data['id'] = "line6"
new_edge1.data['source'] = "NUR"
new_edge1.data['target'] = "LEP"
new_edge1.data['label'] = "150km/h"
# rail connection NUR-FRA
new_edge2 = ipycytoscape.Edge()
new_edge2.data['id'] = "line7"
new_edge2.data['source'] = "NUR"
new_edge2.data['target'] = "FRA"
new_edge2.data['label'] = "150km/h"
ipycytoscape_obj.graph.add_edges([new_edge1,new_edge2])
ipycytoscape_obj
# -
# You see that already a little bit of magic happens. Since NUR is in the west (class="west") it got automatically the blue colour.
# Mmmmm seems that there is still something missing. We don't see the speed of the new regional trains added to the net.
# ## Style of new edges
#
# my_style was defined as a list with all the requirements for visualizing the graph.
# Let's add the new necessary styles.
train_style.append({'selector': 'edge[id = "line6"]',
'style': {
'font-family': 'arial',
'font-size': '10px',
'label': 'data(label)'}})
train_style.append({'selector': 'edge[id = "line7"]',
'style': {
'font-family': 'arial',
'font-size': '10px',
'label': 'data(label)'}})
ipycytoscape_obj2 = ipycytoscape.CytoscapeWidget()
ipycytoscape_obj2.graph.add_graph_from_json(railnetJSON)
ipycytoscape_obj2.graph.add_node(station_NUR)
ipycytoscape_obj2.graph.add_edges([new_edge1,new_edge2])
ipycytoscape_obj2.set_style(train_style)
ipycytoscape_obj2
# NUR station is not anymore in construction.
# So I want to change the label of the NUR station from Hbf NUR in construction by simply "Hbf NUR".
#
ipycytoscape_obj2.graph.nodes
for node in ipycytoscape_obj2.graph.nodes:
if node.data['id'] == 'NUR':
node.data['label'] = 'Hbf NUR'
ipycytoscape_obj2
# ### Adding more nodes, classes and edges
#
# The EU decides to promote a transnational European Union rail connection between mayor cities. New train stations in France, Italy and Spain have to be added to the net.
#
# So far it is planned a 400km/h connection between BER, FRA in Germany, and from FRA to Paris (PAR) to Lyon (LYO) and from Lyon to Barcelona (BAR) and to Milan (MIL).
#
# Besides, We want to visually separate the German train stations from the non-German ones.
#
# So we will add the class "Germany" to all the German stations, and the new stations added will have the class EU.
# +
EU_stations = ['PAR','MIL','BAR','LYO']
new_EU_stations = []
for station in EU_stations:
new_station = ipycytoscape.Node()
new_station.data['id'] = station
new_station.data['label'] = f"CS {station}" # CS = Central Station
new_station.data['classes'] = "EU"
new_EU_stations.append(new_station)
connections = [('BER','PAR'),('PAR','LYO'),('LYO','BAR'),('LYO','MIL')]
lines = [8,9,10,11]
new_EU_rails = []
for i,connection in enumerate(connections):
new_edge = ipycytoscape.Edge()
new_edge.data['id'] = f"line{lines[i]}"
new_edge.data['source'] = connection[0]
new_edge.data['target'] = connection[1]
new_edge.data['classes'] = "EU"
new_edge.data['label'] = "400km/h"
new_EU_rails.append(new_edge)
# -
# We wanted to differentiate the German from the rest of the EU stations. Let's give an orange colour to all the non-German stations.
# Since we added the class "EU" to the new added stations we can add a style to our list of styles as follows.
# Let's check out the new stations added as nodes. And finally build a new graph rail net.
train_style.append({'selector': 'node[classes="EU"]',
'style': {'background-color': 'orange'}})
train_style.append({'selector': 'edge[classes="EU"]',
'style': {'font-family': 'arial',
'font-size': '10px',
'label': 'data(label)'}})
ipycytoscape_obj3 = ipycytoscape.CytoscapeWidget()
ipycytoscape_obj3.graph.add_graph_from_json(railnetJSON)
ipycytoscape_obj3.graph.add_node(station_NUR)
ipycytoscape_obj3.graph.add_edges([new_edge1,new_edge2])
ipycytoscape_obj3.graph.add_nodes(new_EU_stations)
ipycytoscape_obj3.graph.add_edges(new_EU_rails)
ipycytoscape_obj3.set_style(train_style)
ipycytoscape_obj3
# Btw, you might be thinking that the stations (nodes) are not really plotted according to the real geographical location. You can also modify location of nodes. But that is at the moment not part of the scope of this notebook.
# You see that we did not add many nodes, and we are dealing with a tiny graph and our code got already quite verbose for changing here and there properties and attributes.
# Imagine you have to work with the real German rail system. Thousands of stations and thousands of rail connections.
# Only with these few nodes you might have had this feeling of being lost, imagine with some thousand of those!!
# Hence of course the data would be separated from the code in a real project.
#
# ### Conclusion
# We have learn to make more modifications to the ipycytoscape graph. In coming notebooks/articles the European rail net will go on growing.
| examples/Ipycytoscape_from_Scratch_part_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # EXERCÍCIOS FACTORY
#
# [](https://colab.research.google.com/github/catolicasc-joinville/lp1-notebooks/blob/master/3-padroes-de-projeto/3.1-exercicios.ipynb) [launch](https://colab.research.google.com/github/catolicasc-joinville/lp1-notebooks/blob/master/3-padroes-de-projeto/3.1-exercicios.ipynb)
# 1) Crie uma Factory `ShapeFactory` que cria formas geométricas. Faça esta Factory ser capaz de criar objetos dos tipos `Circle` e `Rectangle`. A Factory escolhe qual tipo quer criar baseado no número de parâmetros passados. Esses objetos devem estender da classe `Shape` e devem implementar o método `calculate_area()`.
#
# ```python
#
# class Shape:
# def area(self):
# raise NotImplementedError
#
# class ShapeFactory:
# def factory(self, *size):
# pass
#
# ShapeFactory().factory('circle', 10)
# ShapeFactory().factory('rectangle', 10, 20)
# ```
#
# 2) [Pesquise](https://sourcemaking.com/design_patterns/abstract_factory) sobre o padrão `AbstractFactory` e implemente uma Factory que cria e retorna um Carro. Seja criativo, um carro precisa de motos, rodas, portas, etc.
| python/padroes-de-projeto/factory-exercicios.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 比赛和数据概述:
#
# 在这场比赛中,我们面临的挑战是,如何预测俄罗斯软件公司[1c公司](http://1c.ru/eng/title.htm)下个月每种产品和商店的总销售额。
#
# > 1C公司是干什么的?
# >
# > 1C公司的 Enterprise 8 程序系统旨在实现日常企业活动的自动化:经济和管理活动的各种业务,例如管理会计、业务会计、HR管理、CRM、SRM
# MRP、MRP等。
#
# **数据**:我们现在有每种商品组合的日销售数据,我们的任务是预测每月的销售量。
# +
# Basic packages
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import random as rd # generating random numbers
import datetime # manipulating date formats
# 可视化
import matplotlib.pyplot as plt # basic plotting
import seaborn as sns # for prettier plots
# TIME SERIES
from statsmodels.tsa.arima_model import ARIMA
from statsmodels.tsa.statespace.sarimax import SARIMAX
from pandas.plotting import autocorrelation_plot
from statsmodels.tsa.stattools import adfuller, acf, pacf,arma_order_select_ic
import statsmodels.formula.api as smf
import statsmodels.tsa.api as smt
import statsmodels.api as sm
import scipy.stats as scs
# settings
import warnings
warnings.filterwarnings("ignore")
# -
# 导入数据:
# +
# settings
import warnings
warnings.filterwarnings("ignore")
# 导入销售数据
sales=pd.read_csv("../datasets/Predict Future Sales/sales_train.csv")
item_cat=pd.read_csv("../datasets/Predict Future Sales/item_categories.csv")
item=pd.read_csv("../datasets/Predict Future Sales/items.csv")
sub=pd.read_csv("../datasets/Predict Future Sales/sample_submission.csv")
shops=pd.read_csv("../datasets/Predict Future Sales/shops.csv")
test=pd.read_csv("../datasets/Predict Future Sales/test.csv")
# +
# 正确处理时间列
sales['date'] = sales['date'].apply(lambda x:datetime.datetime.strptime(x, '%d.%m.%Y'))
# 检查
print(sales.info())
# -
sales.head(10)
# +
# Aggregate to monthly level the required metrics
monthly_sales=sales.groupby(["date_block_num","shop_id","item_id"])[
"date","item_price","item_cnt_day"].agg({"date":["min",'max'],"item_price":"mean","item_cnt_day":"sum"})
## Lets break down the line of code here:
# aggregate by date-block(month),shop_id and item_id
# select the columns date,item_price and item_cnt(sales)
# Provide a dictionary which says what aggregation to perform on which column
# min and max on the date
# average of the item_price
# sum of the sales
# -
# take a peak
monthly_sales.head(20)
# +
# 每个类型的商品数量
x = item.groupby(['item_category_id']).count()
x = x.sort_values(by='item_id',ascending=False)
x = x.iloc[0:10].reset_index()
print(x)
# plot
plt.figure(figsize=(8,4))
ax = sns.barplot(x.item_category_id, x.item_id, alpha=0.8)
plt.title("Items per Category")
plt.ylabel('# of items', fontsize=12)
plt.xlabel('Category', fontsize=12)
plt.show()
# -
# 当然,我们可以在此数据集中挖掘更多内容。但是,首先我们还是来深入了解“时间序列”部分。
#
# ### Single series:
#
# 该目标要求我们以 store-item 组合来预测下个月的销售额。
#
# 每个 store-item 随着时间的销售情况本身就是一个时间序列。在深入研究所有组合之前,首先让我们了解如何预测单个序列。
#
# 首先,我们选择预测整个公司每月的总销售额。让我们计算每月的总销售额并绘制该数据。
ts=sales.groupby(["date_block_num"])["item_cnt_day"].sum()
ts.astype('float')
plt.figure(figsize=(16,8))
plt.title('Total Sales of the company')
plt.xlabel('Time')
plt.ylabel('Sales')
plt.plot(ts);
plt.figure(figsize=(16,6))
plt.plot(ts.rolling(window=12,center=False).mean(),label='Rolling Mean');
plt.plot(ts.rolling(window=12,center=False).std(),label='Rolling sd');
plt.legend();
# #### 快速观察
#
# 存在明显的“季节性”(例如:一年中某个时间的销售高峰)和“趋势”下降。
#
# 让我们通过快速分解为趋势,季节性和残差进行检查。
import statsmodels.api as sm
# multiplicative
res = sm.tsa.seasonal_decompose(ts.values,freq=12,model="multiplicative")
fig = res.plot()
# Additive model
res = sm.tsa.seasonal_decompose(ts.values,freq=12,model="additive")
#plt.figure(figsize=(16,12))
fig = res.plot()
#fig.show()
# ### Stationarity - 稳态
#
# 
#
# 稳态指的是序列的时间不变性。即时间序列中的两个点之间的关系仅仅取决于彼此之间的距离,而不是方向(向前、向后。)
#
# 当时间序列固定时,更容易建模,这里我们假定时间序列是稳态的。
#
# 有多种方法可以检测时间序列是不是稳态的:
#
# - ADF( Augmented Dicky Fuller Test)
# - KPSS
# - PP (Phillips-Perron test)
#
# 这里我们使用 ADF 这种常用方法。
# +
# Stationarity tests
def test_stationarity(timeseries):
#Perform Dickey-Fuller test:
print('Results of Dickey-Fuller Test:')
dftest = adfuller(timeseries, autolag='AIC')
dfoutput = pd.Series(dftest[0:4], index=['Test Statistic','p-value','#Lags Used','Number of Observations Used'])
for key,value in dftest[4].items():
dfoutput['Critical Value (%s)'%key] = value
print (dfoutput)
test_stationarity(ts)
# -
| 04其他模型教程/4.04 时间序列问题.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
import matplotlib.pyplot as plt
from sklearn import datasets
from matplotlib import rc
rc("font", family = "serif",size=20)
rc("figure",figsize=(9,6))
rc("figure",facecolor="white")
# %config InlineBackend.figure_format = 'retina'
import cvxpy as cp
import numpy as np
from loss import *
from reg import *
from util import *
from glrm import *
# +
iris = datasets.load_iris()
A = iris.data
colors = np.array(['tab:blue','tab:orange','tab:green'])[iris.target]
loss_list = [(np.arange(A.shape[1]),QuadraticLoss())]
glrm = GLRM(A,loss_list,2,regX = QuadraticReg(),regY=QuadraticReg(.1) )
# -
X,Y = glrm.fit()
plt.scatter(X[:,0],X[:,1],color=colors)
| glrm/regularized PCA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # `PyLammpsMPI`
#
# ## Running on queueing systems
# `pylammpsmpi` integrates with [dask distributed](https://distributed.dask.org/en/latest/) and [dask jobqueue](https://jobqueue.dask.org/en/latest/) to enable running lammps interactively on queueing systems.
# For this example, [slurm cluster](https://jobqueue.dask.org/en/latest/generated/dask_jobqueue.SLURMCluster.html) provided by dask-jobqueue will be used to set up a cluster on which pylammpsmpi will be run.
from dask.distributed import Client
from dask_jobqueue import SLURMCluster
# Create a slurm cluster worker and ten cores. The optional would need to be tweaked according to the SLURM specifications.
cluster = SLURMCluster(queue='shorttime', cores=10, processes=1, job_cpu=10, memory="3GB", walltime="05:59:00")
# Create a client and connect to the cluster
client = Client(cluster)
client
# By default the cluster has no workers. We add a worker,
cluster.scale(1)
# A visual dashboard of jobs running on the cluster is also available. Once a cluster is set up, it can be provided to pylammpsmpi
from pylammpsmpi import LammpsLibrary
# Specify two cores for the job(since we created a cluster with 2), choose the `mode` as `dask`, and pass the `client` to pylammpsmpi object
lmp = LammpsLibrary(cores=10, mode='dask', client=client)
# The rest is similar as you would run on a local machine - except the calculations are run on the cluster.
# Read an input file
lmp.file("../tests/in.simple")
# Check version of lammps
lmp.version
# Check number of atoms
lmp.natoms
# ### Run commands
lmp.command("run 1")
lmp.command(["run 1", "run 1"])
# Commands can also be direct
lmp.run(10)
lmp.mass(1, 20)
# ### Extract a global property
lmp.extract_global("boxxhi")
# ### Access thermo quantities
lmp.get_thermo("temp")
# Thermo quantities can also be accessed directly,
lmp.temp
lmp.press
# ### Accessing simulation box
lmp.extract_box()
# ### Accessing and changing atom properties
# Get individual atom properties, for example force on each atoms
ff = lmp.gather_atoms("f")
print(type(ff))
print(len(ff))
# Get atom properties by their ids
ids = lmp.gather_atoms("id")
ff = lmp.gather_atoms("f", ids=ids[:10])
len(ff)
# Change atom properties
ff = ff*0.5
lmp.scatter_atoms("f", ff, ids=ids[:10])
# ### Access value of variables
temp = lmp.extract_variable("tt", "all", 0)
temp
# ### Access value of computes
ke = lmp.extract_compute("ke", 1, 1)
len(ke)
v = lmp.extract_compute("v", 1, 2, width=3)
v.shape
lmp.extract_compute("1", 0, 0)
msd = lmp.extract_compute("msd", 0, 1, length=4)
msd[0]
# ### Access values from fix
x = lmp.extract_fix("2", 0, 1, 1)
x
# ### Change the simulation box
lmp.delete_atoms("group", "all")
lmp.reset_box([0.0,0.0,0.0], [8.0,8.0,8.0], 0.0,0.0,0.0)
# Finally, the cluster is closed.
client.close()
cluster.close()
| notebooks/lammps_slurm_cluster.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"} tags=[]
# # Feature Engineering
#
# 
# + [markdown] slideshow={"slide_type": "slide"} tags=[]
# ## Objective
#
# Data preprocessing and engineering techniques generally refer to the addition, deletion, or transformation of data.
#
# The time spent on identifying data engineering needs can be significant and requires you to spend substantial time understanding your data...
#
# > _"Live with your data before you plunge into modeling"_ - <NAME>
# + [markdown] slideshow={"slide_type": "fragment"} tags=[]
# In this module we introduce:
#
# - an example of preprocessing numerical features,
# - two common ways to preprocess categorical features,
# - using a scikit-learn pipeline to chain preprocessing and model training.
# + [markdown] slideshow={"slide_type": "slide"} tags=[]
# ## Basic prerequisites
#
# Let's go ahead and import a couple required libraries and import our data.
#
# <div class="admonition note alert alert-info">
# <p class="first admonition-title" style="font-weight: bold;"><b>Note</b></p>
# <p class="last">We will import additional libraries and functions as we proceed but we do so at the time of using the libraries and functions as that provides better learning context.</p>
# </div>
# + tags=[]
import pandas as pd
# to display nice model diagram
from sklearn import set_config
set_config(display='diagram')
# import data
adult_census = pd.read_csv('../data/adult-census.csv')
# separate feature & target data
target = adult_census['class']
features = adult_census.drop(columns='class')
# + [markdown] slideshow={"slide_type": "slide"} tags=[]
# ## Selection based on data types
#
# Typically, data types fall into two categories:
#
# * __Numeric__: a quantity represented by a real or integer number.
# * __Categorical__: a discrete value, typically represented by string labels (but not only) taken from a finite list of possible choices.
# + slideshow={"slide_type": "fragment"} tags=[]
features.dtypes
# + [markdown] slideshow={"slide_type": "skip"} tags=[]
# <div class="admonition warning alert alert-danger">
# <p class="first admonition-title" style="font-weight: bold;"><b>Warning</b></p>
# <p class="last">Do not take dtype output at face value! It is possible to have categorical data represented by numbers (i.e. <tt class="docutils literal">education_num</tt>. And <tt class="docutils literal">object</tt> dtypes can represent data that would be better represented as continuous numbers (i.e. dates).
#
# Bottom line, always understand how your data is representing your features!
# </p>
# </div>
# + [markdown] slideshow={"slide_type": "slide"} tags=[]
# We can separate categorical and numerical variables using their data types to identify them.
#
# There are a few ways we can do this. Here, we make use of [`make_column_selector`](https://scikit-learn.org/stable/modules/generated/sklearn.compose.make_column_selector.html) helper to select the corresponding columns.
# +
from sklearn.compose import make_column_selector as selector
# create selector object based on data type
numerical_columns_selector = selector(dtype_exclude=object)
categorical_columns_selector = selector(dtype_include=object)
# get columns of interest
numerical_columns = numerical_columns_selector(features)
categorical_columns = categorical_columns_selector(features)
# results in a list containing relevant column names
numerical_columns
# + [markdown] slideshow={"slide_type": "slide"} tags=[]
# ## Preprocessing numerical data
# -
# Scikit-learn works "out of the box" with numeric features. However, some algorithms make some assumptions regarding the distribution of our features.
#
# We see that our numeric features span across different ranges:
numerical_features = features[numerical_columns]
numerical_features.describe()
# + [markdown] slideshow={"slide_type": "slide"} tags=[]
# Normalizing our features so that they have mean = 0 and standard deviation = 1, helps to ensure our features align to algorithm assumptions.
#
# <div class="admonition tip alert alert-warning">
# <p class="first admonition-title" style="font-weight: bold;"><b>Tip</b></p>
# <p>Here are some reasons for scaling features:</p>
# <ul class="last simple">
# <li>Models that rely on the distance between a pair of samples, for instance
# k-nearest neighbors, should be trained on normalized features to make each
# feature contribute approximately equally to the distance computations.</li>
# <li>Many models such as logistic regression use a numerical solver (based on
# gradient descent) to find their optimal parameters. This solver converges
# faster when the features are scaled.</li>
# </ul>
# </div>
# + [markdown] slideshow={"slide_type": "fragment"} tags=[]
# Whether or not a machine learning model requires normalization of the features depends on the model family. Linear models such as logistic regression generally benefit from scaling the features while other models such as tree-based models (i.e. decision trees, random forests) do not need such preprocessing (but will not suffer from it).
# + [markdown] slideshow={"slide_type": "fragment"} tags=[]
# We can apply such normalization using a scikit-learn transformer called [`StandardScaler`](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.StandardScaler.html).
# +
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
scaler.fit(numerical_features)
# + [markdown] slideshow={"slide_type": "slide"} tags=[]
# The `fit` method for transformers is similar to the `fit` method for
# predictors. The main difference is that the former has a single argument (the
# feature matrix), whereas the latter has two arguments (the feature matrix and the
# target).
#
# 
# + [markdown] slideshow={"slide_type": "slide"} tags=[]
# In this case, the algorithm needs to compute the mean and standard deviation
# for each feature and store them into some NumPy arrays. Here, these
# statistics are the model states.
#
# <div class="admonition note alert alert-info">
# <p class="first admonition-title" style="font-weight: bold;"><b>Note</b></p>
# <p class="last">The fact that the model states of this scaler are arrays of means and
# standard deviations is specific to the <tt class="docutils literal">StandardScaler</tt>. Other
# scikit-learn transformers will compute different statistics and store them
# as model states, in the same fashion.</p>
# </div>
#
# -
# We can inspect the computed means and standard deviations.
scaler.mean_
scaler.scale_
# + [markdown] slideshow={"slide_type": "skip"} tags=[]
# <div class="admonition tip alert alert-warning">
# <p class="first admonition-title" style="font-weight: bold;"><b>Tip</b></p>
# <p class="last">Scikit-learn convention: if an attribute is learned from the data, its name
# ends with an underscore (i.e. <tt class="docutils literal">_</tt>), as in <tt class="docutils literal">mean_</tt> and <tt class="docutils literal">scale_</tt> for the
# <tt class="docutils literal">StandardScaler</tt>.</p>
# </ul>
# </div>
# + [markdown] slideshow={"slide_type": "slide"} tags=[]
# Once we have called the `fit` method, we can perform data transformation by
# calling the method `transform`.
# + slideshow={"slide_type": "fragment"} tags=[]
numerical_features_scaled = scaler.transform(numerical_features)
numerical_features_scaled
# + [markdown] slideshow={"slide_type": "slide"} tags=[]
# Let's illustrate the internal mechanism of the `transform` method and put it
# to perspective with what we already saw with predictors.
#
# 
#
# The `transform` method for transformers is similar to the `predict` method
# for predictors. It uses a predefined function, called a **transformation
# function**, and uses the model states and the input data. However, instead of
# outputting predictions, the job of the `transform` method is to output a
# transformed version of the input data.
# + [markdown] slideshow={"slide_type": "slide"} tags=[]
# Finally, the method `fit_transform` is a shorthand method to call
# successively `fit` and then `transform`.
#
# 
# + slideshow={"slide_type": "slide"} tags=[]
# fitting and transforming in one step
scaler.fit_transform(numerical_features)
# + [markdown] slideshow={"slide_type": "slide"} tags=[]
# Notice that the mean of all the columns is close to 0 and the standard deviation in all cases is close to 1:
# +
numerical_features = pd.DataFrame(
numerical_features_scaled,
columns=numerical_columns
)
numerical_features.describe()
# + [markdown] slideshow={"slide_type": "slide"} tags=[]
# ## Model pipelines
# -
# We can easily combine sequential operations with a scikit-learn
# [`Pipeline`](https://scikit-learn.org/stable/modules/generated/sklearn.pipeline.Pipeline.html), which chains together operations and is used as any other
# classifier or regressor. The helper function [`make_pipeline`](https://scikit-learn.org/stable/modules/generated/sklearn.pipeline.make_pipeline.html#sklearn.pipeline.make_pipeline) will create a
# `Pipeline`: it takes as arguments the successive transformations to perform,
# followed by the classifier or regressor model.
# + slideshow={"slide_type": "fragment"} tags=[]
from sklearn.linear_model import LogisticRegression
from sklearn.pipeline import make_pipeline
model = make_pipeline(StandardScaler(), LogisticRegression())
model
# + [markdown] slideshow={"slide_type": "slide"} tags=[]
# Let's divide our data into train and test sets and then apply and score our logistic regression model:
# +
from sklearn.model_selection import train_test_split
# split our data into train & test
X_train, X_test, y_train, y_test = train_test_split(numerical_features, target, random_state=123)
# fit our pipeline model
model.fit(X_train, y_train)
# score our model on the test data
model.score(X_test, y_test)
# + [markdown] slideshow={"slide_type": "slide"} tags=[]
# ## Preprocessing categorical data
# -
# Unfortunately, Scikit-learn does not accept categorical features in their raw form. Consequently, we need to transform them into numerical representations.
#
# The following presents typical ways of dealing with categorical variables by encoding them, namely **ordinal encoding** and **one-hot encoding**.
# + [markdown] slideshow={"slide_type": "slide"} tags=[]
# ### Encoding ordinal categories
#
# The most intuitive strategy is to encode each category with a different
# number. The [`OrdinalEncoder`](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.OrdinalEncoder.html) will transform the data in such manner.
# We will start by encoding a single column to understand how the encoding
# works.
# +
from sklearn.preprocessing import OrdinalEncoder
# let's illustrate with the 'education' feature
education_column = features[["education"]]
encoder = OrdinalEncoder()
education_encoded = encoder.fit_transform(education_column)
education_encoded
# + [markdown] slideshow={"slide_type": "slide"} tags=[]
# We see that each category in `"education"` has been replaced by a numeric
# value. We could check the mapping between the categories and the numerical
# values by checking the fitted attribute `categories_`.
# -
encoder.categories_
# + [markdown] slideshow={"slide_type": "skip"} tags=[]
# <div class="admonition note alert alert-info">
# <p class="first admonition-title" style="font-weight: bold;"><b>Note</b></p>
# <p class="last"><tt class="docutils literal">OrindalEncoder</tt> transforms the category value into the corresponding index value of <tt class="docutils literal">encoder.categories_</tt>.</p>
# </div>
# + [markdown] slideshow={"slide_type": "slide"} tags=[]
# However, be careful when applying this encoding strategy:
# using this integer representation leads downstream predictive models
# to assume that the values are ordered (0 < 1 < 2 < 3... for instance).
#
# By default, `OrdinalEncoder` uses a lexicographical strategy to map string
# category labels to integers. This strategy is arbitrary and often
# meaningless. For instance, suppose the dataset has a categorical variable
# named `"size"` with categories such as "S", "M", "L", "XL". We would like the
# integer representation to respect the meaning of the sizes by mapping them to
# increasing integers such as `0, 1, 2, 3`.
# However, the lexicographical strategy used by default would map the labels
# "S", "M", "L", "XL" to 2, 1, 0, 3, by following the alphabetical order.
# + [markdown] slideshow={"slide_type": "slide"} tags=[]
# The `OrdinalEncoder` class accepts a `categories` argument to
# pass categories in the expected ordering explicitly (`categories[i]` holds the categories expected in the ith column).
# + slideshow={"slide_type": "fragment"} tags=[]
ed_levels = [' Preschool', ' 1st-4th', ' 5th-6th', ' 7th-8th', ' 9th', ' 10th', ' 11th',
' 12th', ' HS-grad', ' Prof-school', ' Some-college', ' Assoc-acdm',
' Assoc-voc', ' Bachelors', ' Masters', ' Doctorate']
encoder = OrdinalEncoder(categories=[ed_levels])
education_encoded = encoder.fit_transform(education_column)
education_encoded
# -
encoder.categories_
# + [markdown] slideshow={"slide_type": "skip"} tags=[]
# If a categorical variable does not carry any meaningful order information
# then this encoding might be misleading to downstream statistical models and
# you might consider using one-hot encoding instead (discussed next).
# + [markdown] slideshow={"slide_type": "slide"} tags=[]
# ### Ecoding nominal categories
#
# [`OneHotEncoder`](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.OneHotEncoder.html) is an alternative encoder that converts the categorical levels into new columns.
#
# We will start by encoding a single feature (e.g. `"education"`) to illustrate
# how the encoding works.
# + slideshow={"slide_type": "fragment"} tags=[]
from sklearn.preprocessing import OneHotEncoder
encoder = OneHotEncoder(sparse=False)
education_encoded = encoder.fit_transform(education_column)
education_encoded
# + [markdown] slideshow={"slide_type": "skip"} tags=[]
# <div class="admonition note alert alert-info">
# <p class="first admonition-title" style="font-weight: bold;"><b>Note</b></p>
# <p><tt class="docutils literal">sparse=False</tt> is used in the <tt class="docutils literal">OneHotEncoder</tt> for didactic purposes, namely
# easier visualization of the data.</p>
# <p class="last">Sparse matrices are efficient data structures when most of your matrix
# elements are zero. They won't be covered in detail in this workshop. If you
# want more details about them, you can look at
# <a class="reference external" href="https://scipy-lectures.org/advanced/scipy_sparse/introduction.html#why-sparse-matrices">this</a>.</p>
# </div>
# + [markdown] slideshow={"slide_type": "slide"} tags=[]
# Viewing this as a data frame provides a more intuitive illustration:
# -
feature_names = encoder.get_feature_names(input_features=["education"])
pd.DataFrame(education_encoded, columns=feature_names)
# + [markdown] slideshow={"slide_type": "skip"} tags=[]
# As we can see, each category (unique value) became a column; the encoding
# returned, for each sample, a 1 to specify which category it belongs to.
# + [markdown] slideshow={"slide_type": "slide"} tags=[]
# Let's apply this encoding to all the categorical features:
# +
# get all categorical features
categorical_features = features[categorical_columns]
# one-hot encode all features
categorical_features_encoded = encoder.fit_transform(categorical_features)
# view as a data frame
columns_encoded = encoder.get_feature_names(categorical_features.columns)
pd.DataFrame(categorical_features_encoded, columns=columns_encoded).head()
# + [markdown] slideshow={"slide_type": "fragment"} tags=[]
# <div class="admonition warning alert alert-danger">
# <p class="first admonition-title" style="font-weight: bold;"><b>Warning</b></p>
# <p class="last">One-hot encoding can significantly increase the number of features in our data. In this case we went from 8 features to 102! If you have a data set with many categorical variables and those categorical variables in turn have many unique levels, the number of features can explode. In these cases you may want to explore ordinal encoding or some other alternative.</p>
# </ul>
# </div>
# + [markdown] slideshow={"slide_type": "slide"} tags=[]
# ### Choosing an encoding strategy
#
# Choosing an encoding strategy will depend on the underlying models and the
# type of categories (i.e. ordinal vs. nominal).
#
# <div class="admonition note alert alert-info">
# <p class="first admonition-title" style="font-weight: bold;"><b>Tip</b></p>
# <p class="last">In general <tt class="docutils literal">OneHotEncoder</tt> is the encoding strategy used when the
# downstream models are <strong>linear models</strong> while <tt class="docutils literal">OrdinalEncoder</tt> is often a
# good strategy with <strong>tree-based models</strong>.</p>
# </div>
# + [markdown] slideshow={"slide_type": "fragment"} tags=[]
# Using an `OrdinalEncoder` will output ordinal categories. This means
# that there is an order in the resulting categories (e.g. `0 < 1 < 2`). The
# impact of violating this ordering assumption is really dependent on the
# downstream models. Linear models will be impacted by misordered categories
# while tree-based models will not.
#
# You can still use an `OrdinalEncoder` with linear models but you need to be
# sure that:
# - the original categories (before encoding) have an ordering;
# - the encoded categories follow the same ordering than the original
# categories.
#
# One-hot encoding categorical variables with high cardinality can cause
# computational inefficiency in tree-based models. Because of this, it is not recommended
# to use `OneHotEncoder` in such cases even if the original categories do not
# have a given order.
# + [markdown] slideshow={"slide_type": "slide"} tags=[]
# ## Using numerical and categorical variables together
#
# Now let's look at how to combine some of these tasks so we can preprocess both numeric and categorical data.
#
# First, let's get our train & test data established:
# + slideshow={"slide_type": "fragment"} tags=[]
# drop the duplicated column `"education-num"` as stated in the data exploration notebook
features = features.drop(columns='education-num')
# create selector object based on data type
numerical_columns_selector = selector(dtype_exclude=object)
categorical_columns_selector = selector(dtype_include=object)
# get columns of interest
numerical_columns = numerical_columns_selector(features)
categorical_columns = categorical_columns_selector(features)
# split into train & test sets
X_train, X_test, y_train, y_test = train_test_split(features, target, random_state=123)
# + [markdown] slideshow={"slide_type": "slide"} tags=[]
# Scikit-learn provides a [`ColumnTransformer`](https://scikit-learn.org/stable/modules/generated/sklearn.compose.ColumnTransformer.html) class which will send specific
# columns to a specific transformer, making it easy to fit a single predictive
# model on a dataset that combines both kinds of variables together.
#
# We first define the columns depending on their data type:
#
# * **one-hot encoding** will be applied to categorical columns.
# * **numerical scaling** numerical features which will be standardized.
#
# We then create our `ColumnTransfomer` by specifying three values:
#
# 1. the preprocessor name,
# 2. the transformer, and
# 3. the columns.
#
# First, let's create the preprocessors for the numerical and categorical
# parts.
# + slideshow={"slide_type": "fragment"} tags=[]
categorical_preprocessor = OneHotEncoder(handle_unknown="ignore")
numerical_preprocessor = StandardScaler()
# + [markdown] slideshow={"slide_type": "skip"} tags=[]
#
# <div class="admonition tip alert alert-warning">
# <p class="first admonition-title" style="font-weight: bold;"><b>Tip</b></p>
# <p class="last">We can use the <tt class="docutils literal">handle_unknown</tt> parameter to ignore rare categories that may show up in test data but were not present in the training data.</p>
# </ul>
# </div>
# + [markdown] slideshow={"slide_type": "slide"} tags=[]
# Now, we create the transformer and associate each of these preprocessors
# with their respective columns.
# +
from sklearn.compose import ColumnTransformer
preprocessor = ColumnTransformer([
('one-hot-encoder', categorical_preprocessor, categorical_columns),
('standard_scaler', numerical_preprocessor, numerical_columns)
])
# + [markdown] slideshow={"slide_type": "slide"} tags=[]
# We can take a minute to represent graphically the structure of a
# `ColumnTransformer`:
#
# 
# + [markdown] slideshow={"slide_type": "skip"} tags=[]
# A `ColumnTransformer` does the following:
#
# * It **splits the columns** of the original dataset based on the column names
# or indices provided. We will obtain as many subsets as the number of
# transformers passed into the `ColumnTransformer`.
# * It **transforms each subset**. A specific transformer is applied to
# each subset: it will internally call `fit_transform` or `transform`. The
# output of this step is a set of transformed datasets.
# * It then **concatenates the transformed datasets** into a single dataset.
#
# The important thing is that `ColumnTransformer` is like any other
# scikit-learn transformer. In particular it can be combined with a classifier
# in a `Pipeline`:
# + slideshow={"slide_type": "slide"} tags=[]
model = make_pipeline(preprocessor, LogisticRegression(max_iter=500))
model
# + [markdown] slideshow={"slide_type": "skip"} tags=[]
# <div class="admonition warning alert alert-danger">
# <p class="first admonition-title" style="font-weight: bold;"><b>Warning</b></p>
# <p class="last">Including non-scaled data can cause some algorithms to iterate
# longer in order to converge. Since our categorical features are not scaled it's often recommended to increase the number of allowed iterations for linear models.</p>
# </div>
# + slideshow={"slide_type": "fragment"} tags=[]
# fit our model
_ = model.fit(X_train, y_train)
# score on test set
model.score(X_test, y_test)
# + [markdown] slideshow={"slide_type": "slide"} tags=[]
# ## Wrapping up
# -
# Unfortunately, we only have time to scratch the surface of feature engineering in this workshop. However, this module should provide you with a strong foundation of how to apply the more common feature preprocessing tasks.
#
# <div class="admonition tip alert alert-warning">
# <p class="first admonition-title" style="font-weight: bold;"><b>Tip</b></p>
# <p class="last">Scikit-learn provides many feature engineering options. Learn more here: <a href="https://scikit-learn.org/stable/modules/preprocessing.html">https://scikit-learn.org/stable/modules/preprocessing.html</a></p>
# </ul>
# </div>
# + [markdown] slideshow={"slide_type": "fragment"} tags=[]
# In this module we learned how to:
#
# - normalize numerical features with `StandardScaler`,
# - ordinal and one-hot encode categorical features with `OrdinalEncoder` and `OneHotEncoder`, and
# - chain feature preprocessing and model training steps together with `ColumnTransformer` and `make_pipeline`.
# -
| notebooks/05-feat_eng.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#single image RPCA
import sys, os
import numpy as np
import glob
from matplotlib import pylab as plt
from numpy.linalg import svd
from PIL import Image
from RPCA_ADMM import TRPCA
import time
img =Image.open('E:/cjh/CNN/admm/oriface/4/2.jpg')
R,G,B= img.split()
R = np.asarray(R.convert("L"))
G = np.asarray(G.convert("L"))
B = np.asarray(B.convert("L"))
TRPCA = TRPCA()
RL, RS = TRPCA.ADMM(R)
GL, GS = TRPCA.ADMM(G)
BL, BS = TRPCA.ADMM(B)
imB=Image.fromarray(BL).convert("L")
imG=Image.fromarray(GL).convert("L")
imR=Image.fromarray(RL).convert("L")
S1=Image.fromarray(BS).convert("L")
S2=Image.fromarray(GS).convert("L")
S3=Image.fromarray(RS).convert("L")
S=Image.merge('RGB',(S1,S2,S3))
L=Image.merge('RGB',(imR,imG,imB))
L.save('singleL.jpg')
S.save('singleS.jpg')
# +
#multi
#make sure that the images all have the same size
import sys, os
import numpy as np
import glob
from matplotlib import pylab as plt
from numpy.linalg import svd
from PIL import Image
from RPCA_ADMM import TRPCA
import time
import glob2
faces = glob2.glob('E:/cjh/CNN/admm/oriface/4/*.jpg')
matrixB = []
matrixG = []
matrixR = []
for face in faces:
img = Image.open(face)
R,G,B=img.split()
#R=img[ :, :,0]
#G=img[ :, :,1]
#B=img[ :, :,2]
h,w=np.array(B).shape
pixelsB = list(B.getdata())
matrixB.append(pixelsB)
pixelsG = list(G.getdata())
matrixG.append(pixelsG)
pixelsR = list(R.getdata())
matrixR.append(pixelsR)
B=(np.array(matrixB).astype(np.float64))
G=(np.array(matrixG).astype(np.float64))
R=(np.array(matrixR).astype(np.float64))
TRPCA = TRPCA()
RL, RS = TRPCA.ADMM(R)
GL, GS = TRPCA.ADMM(G)
BL, BS = TRPCA.ADMM(B)
BL = BL[1,:].reshape((h,w))
BS = BS[1,:].reshape((h,w))
GL = GL[1,:].reshape((h,w))
GS = GS[1,:].reshape((h,w))
RL = RL[1,:].reshape((h,w))
RS = RS[1,:].reshape((h,w))
imB=Image.fromarray(BL).convert("L")
imG=Image.fromarray(GL).convert("L")
imR=Image.fromarray(RL).convert("L")
S1=Image.fromarray(BS).convert("L")
S2=Image.fromarray(GS).convert("L")
S3=Image.fromarray(RS).convert("L")
"""plt.subplot(131)
plt.imshow(L)
plt.gray()
plt.subplot(132)
plt.imshow(S)
plt.gray()
plt.subplot(133)
plt.imshow(X)
plt.gray()
plt.show()
"""
S=Image.merge('RGB',(S1,S2,S3))
L=Image.merge('RGB',(imR,imG,imB))
L.save('multiL.jpg')
S.save('multiS.jpg')
# -
| RPCA/RPCA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/piotr14prywatny/dw_matrix/blob/master/day1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="ud9UU5Xn-UoX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="0a8f0019-beab-4fde-fa4e-9896af60ae07"
# !pwd
# + id="BgXuUg8SAMkD" colab_type="code" colab={}
# mkdir -p '/content/drive/My Drive/Colab Notebooks/matrix/matrix_three'
# + id="J0v7ZnmaBDPp" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="2f85c575-1936-4f78-f59f-534ebf0a16e0"
# cd '/content/drive/My Drive/Colab Notebooks/matrix/matrix_three'
# + id="Fh307RAdBMJa" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="148161f3-fbff-40a3-f523-98f3976b1020"
pwd
# + id="jCyGevjFBtqU" colab_type="code" colab={}
GIT_TOKEN = 'ffad4953eb7c51200ba5cc086a39b0ff97bd7d96'
GIT_URL = 'https://{0}@github.com/piotr14prywatny/dw_matrix_road_sign.git'.format(GIT_TOKEN)
# + id="4EnnzztVDYZe" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 118} outputId="5d82f397-979a-44da-909e-03147a908c58"
# !git clone $GIT_URL
# + id="CkJDXUsWDuzu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="0de44364-a516-475c-8a90-a43ff986dea5"
# ls
# + id="NdSRZrTDD0e9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="1b81792f-ac49-4a72-d4dc-7ba84a3dfe6e"
# cd dw_matrix_road_sign
# + id="cmlq4JSOD_xC" colab_type="code" colab={}
# !mkdir data
# + id="CjGdxjeDEEm9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="9a668cae-6530-471c-9886-22dc4cd4f65c"
# cd data
# + id="ExqGmRQREIb5" colab_type="code" colab={}
- bit.ly/train_road_sign
- bit.ly/test_road_sign
- bit.ly/dw_signnames
# + id="GtfppSGDE0qR" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 118} outputId="9056f727-11d7-4a02-ef82-2994432f0c41"
# !curl -L bit.ly/train_road_sign -o train.p
# + id="-_pMfU3kF3j4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 118} outputId="0e2f7512-48e8-4ebc-dd50-444844a182e8"
# !curl -L bit.ly/test_road_sign -o test.p
# + id="xiTHPWJfGFos" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 118} outputId="a23b3a84-8d34-48e0-bd39-e5f2673fe46c"
# !curl -L bit.ly/dw_signnames -o signnames.csv
# + id="mQUlL3bDGsxn" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="4fb99712-c895-45a0-82da-e48c4d3f7955"
# ls
# + id="y-cttXErG7qP" colab_type="code" colab={}
import pandas as pd
# + id="tExWMWoRHEiN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="125186bb-42b4-41e4-f22d-6157b150377f"
train = pd.read_pickle('train.p')
train.keys()
# + id="dHgGP3pBHRp-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="0d7ad047-cb79-4b21-f3df-f0c9b463fec2"
X_train, y_train = train['features'], train['labels']
X_train.shape, y_train.shape
# + id="7E23k-3pH-34" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 343} outputId="4f6a3301-c66e-47ce-f683-6ce3ce6d4909"
pd.read_csv('signnames.csv').sample(10)
# + id="km9Pt_MrIcFe" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="d868a29f-8927-42e8-b358-a7b01aa2127f"
# ls
# + id="OscoQJUBJLzc" colab_type="code" colab={}
| day1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/JimKing100/DS-Unit-3-Sprint-2-SQL-and-Databases/blob/master/module3-nosql-and-document-oriented-databases/Module3-Assignment1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="PD-9ATok-T_W" colab_type="code" colab={}
import sqlite3
import pymongo
# + id="G6seilCv-yC5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="4005fdec-a15d-403e-b78c-29cd5345b7a3"
# Queries
query1 = 'SELECT COUNT(character_id) \
FROM charactercreator_character;'
# File is uploaded to Colab
conn = sqlite3.connect('rpg_db.sqlite3')
# Execute queries
curs1 = conn.cursor()
print('Total characters =', curs1.execute(query1).fetchall())
# + id="xeQaFp30_PID" colab_type="code" colab={}
# Close cursors and commit
curs1.close()
conn.commit()
# + id="e5y4u4GS_bwq" colab_type="code" colab={}
# Connect to sqlite RPG DB
sl_conn =sqlite3.connect('rpg_db.sqlite3')
sl_cur = sl_conn.cursor()
# + id="WHR2DjeU_iwX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="bc6603c9-2e14-4c9b-bb8b-5184194a2938"
# Test the Connection
sl_cur.execute('SELECT * FROM charactercreator_character')
sl_cc_table = sl_cur.fetchall()
sl_cc_table[0]
# + id="JXkU_jjQAZW6" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="34a4af15-c670-4690-a6e8-8eada7cb6b28"
len(sl_cc_table)
# + id="G60d7vQeBVk6" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="395ef1ae-9af8-452f-9379-0fdac15bf331"
sl_cc_table[0][1]
# + id="r4wSndDC-Zqe" colab_type="code" colab={}
client = pymongo.MongoClient("mongodb://admin:uvuUMd5mEVbtYEG4@cluster0-shard-00-00-tlxq2.mongodb.net:27017,cluster0-shard-00-01-tlxq2.mongodb.net:27017,cluster0-shard-00-02-tlxq2.mongodb.net:27017/test?ssl=true&replicaSet=Cluster0-shard-0&authSource=admin&retryWrites=true&w=majority")
rpg_data = db.rpg_data
# + id="tkdelLEOARcI" colab_type="code" colab={}
for i in range(0, len(sl_cc_table)-1):
rpg_data.insert_one({'id': i,
'char_id': sl_cc_table[i][0],
'name': sl_cc_table[i][1],
'level': sl_cc_table[i][2],
'exp': sl_cc_table[i][3],
'hp': sl_cc_table[i][4],
'strength': sl_cc_table[i][5],
'intelligence': sl_cc_table[i][6],
'dexterity': sl_cc_table[i][7],
'wisdom': sl_cc_table[i][8]
})
# + id="s5zpa7cFDyTu" colab_type="code" colab={}
cur = rpg_data.find()
# + id="wBJCv0HPGE7Y" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="fd0d49cd-388b-40ee-f7ef-0e1c4ed4b6f0"
for i in cur:
print(i)
| module3-nosql-and-document-oriented-databases/Module3-Assignment1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:c3dev] *
# language: python
# name: conda-env-c3dev-py
# ---
# # Applying a discrete-time, non-stationary nucleotide model
#
# We fit a discrete-time Markov nucleotide model. This corresponds to a Barry and Hartigan 1987 model.
# +
from cogent3.app import io, evo
loader = io.load_aligned(format="fasta", moltype="dna")
aln = loader("../data/primate_brca1.fasta")
model = evo.model("BH", tree="../data/primate_brca1.tree")
result = model(aln)
result
# -
# **NOTE:** DLC stands for diagonal largest in column and the value is a check on the identifiability of the model. `unique_Q` is not applicable to a discrete-time model and so remains as `None`.
#
# Looking at the likelihood function, you will
result.lf
# ## Get a tree with branch lengths as paralinear
#
# This is the only possible length metric for a discrete-time process.
tree = result.tree
fig = tree.get_figure()
fig.scale_bar = "top right"
fig.show(width=500, height=500)
# ## Getting parameter estimates
#
# For a discrete-time model, aside from the root motif probabilities, everything is edge specific. But note that the `tabular_result` has different keys from the continuous-time case, as demonstrated below.
tabulator = evo.tabulate_stats()
stats = tabulator(result)
stats
stats['edge motif motif2 params']
| doc/app/evo-dt-nuc-model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import datetime as dt
sns.set()
df = pd.read_csv('../TICKER.csv')
df.head()
count = int(np.ceil(len(df) * 0.1))
signals = pd.DataFrame(index=df.index)
signals['signal'] = 0.0
signals['trend'] = df['Close']
signals['RollingMax'] = (signals.trend.shift(1).rolling(count).max())
signals['RollingMin'] = (signals.trend.shift(1).rolling(count).min())
signals.loc[signals['RollingMax'] < signals.trend, 'signal'] = -1
signals.loc[signals['RollingMin'] > signals.trend, 'signal'] = 1
signals
def buy_stock(
real_movement,
signal,
initial_money = 10000,
max_buy = 1,
max_sell = 1,
):
"""
real_movement = actual movement in the real world
delay = how much interval you want to delay to change our decision from buy to sell, vice versa
initial_state = 1 is buy, 0 is sell
initial_money = 1000, ignore what kind of currency
max_buy = max quantity for share to buy
max_sell = max quantity for share to sell
"""
starting_money = initial_money
states_sell = []
states_buy = []
current_inventory = 0
def buy(i, initial_money, current_inventory):
shares = initial_money // real_movement[i]
if shares < 1:
print(
'at %d: total balances %f, not enough money to buy a unit price %f'
% (, initial_money, real_movement[i])
)
else:
if shares > max_buy:
buy_units = max_buy
else:
buy_units = shares
initial_money -= buy_units * real_movement[i]
current_inventory += buy_units
print(
'day %s: buy %d units at price %f, total balance %f'
% (dt.datetime., buy_units, buy_units * real_movement[i], initial_money)
)
states_buy.append(0)
return initial_money, current_inventory
for i in range(real_movement.shape[0] - int(0.025 * len(df))):
state = signal[i]
if state == 1:
initial_money, current_inventory = buy(
i, initial_money, current_inventory
)
states_buy.append(i)
elif state == -1:
if current_inventory == 0:
print('day %d: cannot sell anything, inventory 0' % (i))
else:
if current_inventory > max_sell:
sell_units = max_sell
else:
sell_units = current_inventory
current_inventory -= sell_units
total_sell = sell_units * real_movement[i]
initial_money += total_sell
try:
invest = (
(real_movement[i] - real_movement[states_buy[-1]])
/ real_movement[states_buy[-1]]
) * 100
except:
invest = 0
print(
'day %d, sell %d units at price %f, investment %f %%, total balance %f,'
% (i, sell_units, total_sell, invest, initial_money)
)
states_sell.append(i)
invest = ((initial_money - starting_money) / starting_money) * 100
total_gains = initial_money - starting_money
return states_buy, states_sell, total_gains, invest
states_buy, states_sell, total_gains, invest = buy_stock(df.Close, signals['signal'])
close = df['Close']
fig = plt.figure(figsize = (15,5))
plt.plot(close, color='r', lw=2.)
plt.plot(close, '^', markersize=10, color='m', label = 'buying signal', markevery = states_buy)
plt.plot(close, 'v', markersize=10, color='k', label = 'selling signal', markevery = states_sell)
plt.title('total gains %f, total investment %f%%'%(total_gains, invest))
plt.legend()
plt.show()
| agent/1.turtle-agent.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from pyspark import SparkContext, SparkConf
from pyspark.sql import SQLContext
from pyspark.sql.types import *
# Assure Hadoop conf dir is correct for DC/OS Data Science Engine
import os,subprocess
myenv = dict(os.environ, HADOOP_CONF_DIR="/mnt/mesos/sandbox")
# +
# # Move data to HDFS for shared access if directory not already provided
# subprocess.check_output('hdfs dfs -mkdir -p churndata/', shell=True, env=myenv)
# subprocess.check_output('hdfs dfs -put -f ../data/churn.all churndata/', shell=True, env=myenv);
# +
conf = SparkConf().setAppName("test-me-some-s3")
sc = SparkContext(conf=conf)
sql = SQLContext(sc)
# -
# sql.read.csv('s3a://gregorygrubbs/datasets/churn.all')
gsdf = sql.read.csv('s3a://gregoryg/datasets/github_stats/github_stats.csv.gz', inferSchema=True, header=True)
os.getenv('SPARK_HADOOP_FS_S3A_AWS_CREDENTIALS_PROVIDER')
gsdf.createTempView('githubstats')
sql.sql('select * FROM githubstats').toPandas()
# +
# sc.stop()
| notebooks/broken/s3-pyspark.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import time
import torch
from torch import nn
from ml_models import SuperResolutionNet
from datasets import SRDataset
from utils import *
import numpy as np
from scipy.interpolate import interp2d, RectBivariateSpline
from importlib import reload
import pylab as plt
# %matplotlib inline
import matplotlib
font = {'family' : 'normal',
'weight' : 'bold',
'size' : 10}
matplotlib.rc('font', **font)
# Data parameters
data_folder = './exp2' # folder with JSON data files
scaling_factor = 4 # the scaling factor for the generator; the input LR images will be downsampled from the target HR images by this factor
# +
import diagnostics
def cal_enstrophy_spec(field2d):
spec = diagnostics.grid2spec(field2d)
return diagnostics.prod_spectrum(spec, spec)
def BivariateSplineInterp(low_res, scaling_factor):
num_rows, num_cols = low_res.shape
interp_f = RectBivariateSpline(np.linspace(0, 1, num_rows), np.linspace(0, 1, num_cols), low_res)
interp_array = np.zeros((num_rows*scaling_factor, num_cols*scaling_factor))
high_res_x = np.linspace(0, 1, num_rows * scaling_factor)
high_res_y = np.linspace(0, 1, num_cols * scaling_factor)
for i in range(num_rows * scaling_factor):
for j in range(num_cols * scaling_factor):
interp_array[i, j] = interp_f(high_res_x[i], high_res_y[j])
return interp_array
# +
train_dataset = SRDataset(data_folder,
scaling_factor=scaling_factor)
low_res, high_res = train_dataset[25]
interped_res = BivariateSplineInterp(low_res[0, :, :], scaling_factor)
# -
checkpoint = torch.load("/home/juchai/azblob/an_model/epoch195_checkpoint_srgan.pth.tar")
model = checkpoint['generator'].cpu()
# +
with torch.no_grad():
super_res = model(low_res).reshape(high_res.shape).detach().cpu().numpy()
super_res = super_res.reshape(super_res.shape[1:])
if len(low_res.shape) == 2:
low_res = low_res.reshape(low_res.shape[1:])
# +
# %matplotlib inline
plt.figure(dpi= 300)
high_res_xlim = [200, 300]
high_res_ylim = [100, 250]
plt.subplot(1, 5, 1)
plt.imshow(low_res.T, interpolation='none')
if high_res_xlim is not None:
plt.xlim([high_res_xlim[0]//scaling_factor, high_res_xlim[1]//scaling_factor])
plt.ylim([high_res_ylim[0]//scaling_factor, high_res_ylim[1]//scaling_factor])
plt.title("low res")
plt.subplot(1, 5, 2)
plt.imshow(high_res.T, interpolation='none')
if high_res_xlim is not None:
plt.xlim([high_res_xlim[0], high_res_xlim[1]])
plt.ylim([high_res_ylim[0], high_res_ylim[1]])
plt.title("high res")
plt.subplot(1, 5, 3)
plt.imshow(super_res.T, interpolation='none')
if high_res_xlim is not None:
plt.xlim([high_res_xlim[0], high_res_xlim[1]])
plt.ylim([high_res_ylim[0], high_res_ylim[1]])
plt.title("super res")
plt.subplot(1, 5, 4)
plt.imshow(interped_res.T, interpolation='none')
if high_res_xlim is not None:
plt.xlim([high_res_xlim[0], high_res_xlim[1]])
plt.ylim([high_res_ylim[0], high_res_ylim[1]])
plt.title("SplineInterp")
plt.subplot(1, 5, 5)
plt.imshow(super_res.T - interped_res.T, interpolation='none')
if high_res_xlim is not None:
plt.xlim([high_res_xlim[0], high_res_xlim[1]])
plt.ylim([high_res_ylim[0], high_res_ylim[1]])
plt.title("SRes-Interp")
# +
for field_name, field_value in [("low_res", low_res), ("high_res", high_res),
("sup_res", super_res), ("interped_res", interped_res)]:
if isinstance(field_value, torch.Tensor):
field_value = field_value.numpy()
spec = cal_enstrophy_spec(field_value)
plt.loglog(*spec, label=field_name)
orignal_size = low_res.shape[-1] / 2
plt.plot([orignal_size, orignal_size], [1e-4, 1e-2], "--")
plt.ylim([1e-6, 1])
plt.plot([orignal_size*2, orignal_size*2], [1e-4, 1e-2], "--")
plt.ylim([1e-6, 1])
plt.legend()
# +
#plt.figure(dpi= 300)
plt.imshow(super_res.T - interped_res.T, interpolation='nearest')
# if high_res_xlim is not None:
# plt.xlim([high_res_xlim[0], high_res_xlim[1]])
# plt.ylim([high_res_ylim[0], high_res_ylim[1]])
plt.title("super res - Interp")
plt.xlim([200, 400])
plt.ylim([100, 300])
plt.colorbar()
# -
| visualize_super_resolution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + deletable=true editable=true
import matplotlib.pyplot as plt
# %matplotlib inline
import cv2
import numpy as np
import pickle
from scipy.misc import imread
from birdseye import BirdsEye
from helpers import show_images, save_image, roi
from lanefilter import LaneFilter
# + deletable=true editable=true
calibration_data = pickle.load(open("calibration_data.p", "rb" ))
matrix = calibration_data['camera_matrix']
dist_coef = calibration_data['distortion_coefficient']
source_points = [(580, 460), (205, 720), (1110, 720), (703, 460)]
dest_points = [(320, 0), (320, 720), (960, 720), (960, 0)]
p = { 'sat_thresh': 120, 'light_thresh': 40, 'light_thresh_agr': 205,
'grad_thresh': (0.7, 1.4), 'mag_thresh': 40, 'x_thresh': 20 }
birdsEye = BirdsEye(source_points, dest_points, matrix, dist_coef)
laneFilter = LaneFilter(p)
# + deletable=true editable=true
def lane_filter_test(path):
img = imread(path)
img = birdsEye.undistort(img)
binary = laneFilter.apply(img)
masked_lane = np.logical_and(birdsEye.sky_view(binary), roi(binary))
sobel_img = birdsEye.sky_view(laneFilter.sobel_breakdown(img))
color_img = birdsEye.sky_view(laneFilter.color_breakdown(img))
show_images([color_img, sobel_img, masked_lane], per_row = 3, per_col = 1, W = 15, H = 5)
# + deletable=true editable=true
for i in range(1, 7):
lane_filter_test("test_images/test" + str(i) + ".jpg")
# + deletable=true editable=true
lane_filter_test("test_images/straight_lines1.jpg")
# + deletable=true editable=true
lane_filter_test("test_images/straight_lines2.jpg")
# + deletable=true editable=true
| notebooks/gradient_and_color_thresholding.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # The Capacitor
# + slideshow={"slide_type": "-"}
# Additional styling ; should be moved into helpers
from IPython.core.display import display, HTML
HTML('<style>{}</style>'.format(open('styler.css').read()))
# + [markdown] slideshow={"slide_type": "slide"}
# ## Welcome!
#
# To run a cell, click the cell and hit "Shift+Enter" on your keyboard.
# + slideshow={"slide_type": "fragment"}
from helpers import *
# + [markdown] cell_style="split" slideshow={"slide_type": "subslide"}
# ### Reminder & Review!
#
# - Electric fields $\vec{E}$ are formed between positive and negative charges
# - $\vec{E}$ has magnitude (units of $N/C$ or $V/m$) and a direction
# - The work done on a charge $q$ by the electric field is:
#
# $$ W = \vec{F} d $$
#
# - The voltage difference between two plates can be expressed as:
#
# $$ V = \frac{\vec{F} d}{q} = \vec{E}d$$
#
# + [markdown] cell_style="split" slideshow={"slide_type": "fragment"}
# <center>
# <img src="../../fig7_2.jpeg" width=50%>
# </center>
#
# <p style="text-align:right; font-size:40%;"><a href="https://openstax.org/books/university-physics-volume-2/pages/8-1-capacitors-and-capacitance#91635">Source: OpenStax Textbook</a></p>
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Learning Objectives
#
# <br><br>
#
# - Identify and summarize key functions of the capacitor
# - Explain the relationship between the capacitance and the plate separation and area
# - Describe applications of the capacitor based on its properties
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Stud finder!
#
# <br>
# Today you will learn how a stud-finder works - it uses electric fields and a capacitor!
# + slideshow={"slide_type": "fragment"}
IFrame(stud_finder,width='100%',height=600)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## What is a capacitor?
# + [markdown] cell_style="split" slideshow={"slide_type": "fragment"}
# - Capacitors are very important components of many electronic devices including:
# - Pacemakers
# - Flash lamps on cameras
# - Defribrillators
# - Cell phones and computers
# + [markdown] cell_style="split" slideshow={"slide_type": "fragment"}
# <center>
# <img src="../../fig_8_16.jpeg" width=50%>
# </center>
#
# <p style="text-align:right; font-size:40%;"><a href="https://openstax.org/books/university-physics-volume-2/pages/8-3-energy-stored-in-a-capacitor">Source: OpenStax Textbook</a></p>
# + [markdown] cell_style="split" slideshow={"slide_type": "fragment"}
# - A capacitor is a device that stores electrical charge.
#
# - The energy is stored within an electric field (between two conducting plates).
#
# - Capacitance is defined as the ratio of the charge $Q$ on a conductor to the potential difference $V$ between them:
#
# $$C \equiv \frac{Q}{\Delta V}$$
#
# - On the right, is the **parallel plate capacitor**.
# + [markdown] cell_style="split" slideshow={"slide_type": "fragment"}
# <center>
# <img src="../../fig8_2.jpeg" width=30%>
# </center>
#
# <p style="text-align:right; font-size:30%;"><a href="https://openstax.org/books/university-physics-volume-2/pages/8-1-capacitors-and-capacitance#91635">Source: OpenStax Textbook</a></p>
# + [markdown] cell_style="center" slideshow={"slide_type": "fragment"}
# - The unit of capacitance is a "Farad", or F (in honour of <NAME>)
#
# $$ 1 F = \frac{1C}{1V} $$
# + [markdown] cell_style="split" slideshow={"slide_type": "subslide"}
# ### Exploring Capacitors
#
# We will explore the dependence of three parameters on the capacitance:
#
# 1. Voltage $V$
# 2. Plate separation $d$
# 3. Plate area $A$
#
# We will use a simulation app to build and develop some intuition!
# + [markdown] cell_style="split" slideshow={"slide_type": "fragment"}
# <center>
# <img src="../../fig8_5_orig.jpeg" width=50%>
# </center>
#
# <p style="text-align:right; font-size:30%;"><a href="https://openstax.org/books/university-physics-volume-2/pages/8-1-capacitors-and-capacitance#91635">Source: OpenStax Textbook</a></p>
# + slideshow={"slide_type": "slide"}
display(Markdown("## Capacitor Simulation Demo"))
IFrame(capacitor_phet, width='100%',height=900)
# + [markdown] cell_style="split" slideshow={"slide_type": "subslide"}
# ### Activity 1: Does the battery affect the Capacitance?
#
# <br><br>
# What do you think happens to the Capacitance if:
#
# 1. the battery voltage is maximum ?
#
# 2. the battery voltage is 0 ?
#
# 3. the battery voltage is negative?
# + cell_style="split" slideshow={"slide_type": "fragment"}
IFrame('https://app.sli.do/event/gkmj65xm/embed/polls/43a660b4-f549-4cde-b9b3-91165a66a42a', width=700,height=500)
# + cell_style="split" slideshow={"slide_type": "subslide"}
IFrame(capacitor_phet, width='100%',height=600)
# + [markdown] cell_style="split" slideshow={"slide_type": "fragment"}
# ### Instructions
#
# 1. Keep all parameters except the **Voltage** constant.
# 2. Step through the voltages from 0.3 to 1.0 in units of 0.1 V and record the top-plate charge.
# 3. Type in the data for V and Q into the lists below.
# 4. Observe the relationship between Voltage ($V$) and Charge ($Q$) in the plot.
# + cell_style="split" slideshow={"slide_type": "fragment"}
voltage = [0.30, 0.40, 0.50, 0.60, 0.70, 0.80, 0.90, 1.0] # units: V
charge = [0.09, 0.12, 0.15, 0.18, 0.21, 0.24, 0.27, 0.30] # units: pC
# Plot here
plot_VQ(voltage,charge)
# + [markdown] cell_style="split" slideshow={"slide_type": "subslide"}
# ### Activity 2: Effect of plate separation on capacitance
#
# <br><br>
# What do you think happens to the $\vec{E}$ field when:
#
# 1. the plates touch ($d = 0$) ?
#
# 1. the plates are extremely far apart ($d \to \infty$) ?
# + cell_style="split" slideshow={"slide_type": "fragment"}
IFrame('https://app.sli.do/event/gkmj65xm/embed/polls/07567c0d-6ab5-4a01-a7e6-ec99cf8c8eaf', width=700,height=500)
# + cell_style="split" slideshow={"slide_type": "subslide"}
IFrame(capacitor_phet, width='100%',height=600)
# + [markdown] cell_style="split" slideshow={"slide_type": "fragment"}
# ### Instructions
#
# 1. Keep all parameters except the **plate separation** constant.
# 2. Charge the capacitor (set $V$ to 1.5V).
# 2. Step through at least 6 separation distances between 2.0 mm to 10mm and record the Capacitance.
# 3. Type in the data for capacitance and separation into the lists below.
# 4. Identify the relationship between $C$ and $d$.
# + cell_style="split" slideshow={"slide_type": "fragment"}
capacitance = [0.18, 0.30, 0.35, 0.44, 0.59, 0.89] # units: pF
separation = [1/10, 1/6.0, 1/5.0, 1/4.0, 1/3.0, 1/2.0] # units: mm
# Plot here
plot_dC(capacitance,separation)
# + slideshow={"slide_type": "skip"}
# + [markdown] cell_style="split" slideshow={"slide_type": "subslide"}
# ### Activity 3: Effect of plate area on capacitance
#
# <br><br>
# What do you think happens to the charges when:
#
# 1. the plate area A increases, $A\to \infty$ ?
#
# 1. the plate area A decreases, $A \to 0$ ?
#
# + cell_style="split" slideshow={"slide_type": "fragment"}
IFrame('https://app.sli.do/event/gkmj65xm/embed/polls/6881fa0b-2c6f-4f38-aaae-967d10c8129e', width=700,height=500)
# + cell_style="split" slideshow={"slide_type": "subslide"}
IFrame(capacitor_phet, width='100%',height=600)
# + [markdown] cell_style="split" slideshow={"slide_type": "fragment"}
# ### Instructions
#
# 1. Keep all parameters except the **plate area** constant.
# 2. Step through at least 4 areas between 100 and 400 mm$^2$ and record the Capacitance.
# 3. Type in the data for capacitance and areas into the lists below.
# 4. Identify the relationship between $C$ and $A$.
# + cell_style="split" slideshow={"slide_type": "fragment"}
capacitance = [0.15, 0.30, 0.44, 0.59 ] # units: pF
area = [100,200,300,400 ]# units: mm^2
plot_AC(capacitance,area)
# + [markdown] cell_style="split" slideshow={"slide_type": "subslide"}
# ### Recap: Parallel Plate Capacitor Activity
#
# 1. C and Voltage $V$:
# <br><br><br>
# 2. C and Plate separation $d$:
# <br><br><br>
# 3. C and Plate area $A$:
# <br><br><br>
# + [markdown] cell_style="split" slideshow={"slide_type": "fragment"}
# #### Capacitance of a Parallel Plate Capacitor:
# <br><br>
# $$ C = \frac{\epsilon_0 A}{d} $$
# <br><br><br><br><br>
# Where does $\epsilon_0$ come from? Go back to [this pencast](https://youtu.be/gTfEFwVsgKA) for the derivation of the Electric Field between parallel plates. More in the Tutorial this week!
# + [markdown] cell_style="center" slideshow={"slide_type": "slide"}
# ### Capacitors in practice
# + [markdown] cell_style="split" slideshow={"slide_type": "fragment"}
# - Capacitors come in many shapes and geometries:
# <br>
# <br>
# <center>
# <img src="../../fig8_4.jpeg" width=50%>
# </center>
#
# <p style="text-align:right; font-size:30%;"><a href="https://openstax.org/books/university-physics-volume-2/pages/8-1-capacitors-and-capacitance#91635">Source: <NAME></a></p>
#
# + [markdown] cell_style="split" slideshow={"slide_type": "fragment"}
# - Typically an insulating material is placed between conducting sheets (dielectric) to make the capacitor more effective
#
# <center>
# <img src="../../fig8_2b.jpeg" width=70%>
# </center>
#
# <p style="text-align:right; font-size:30%;"><a href="https://openstax.org/books/university-physics-volume-2/pages/8-1-capacitors-and-capacitance#91635">Source: OpenStax Textbook</a></p>
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Between the plates
# + [markdown] cell_style="split" slideshow={"slide_type": "fragment"}
# This is a molecular model of an insulating material that contains polar (one side positive, one side negative) molecules:
#
# <img src="../../fig8_20_a.jpeg" width=35%>
#
# Notice that the molecules are currently randomly assorted and distributed.
#
# <p style="text-align:right; font-size:30%;"><a href="https://openstax.org/books/university-physics-volume-2/pages/8-1-capacitors-and-capacitance#91635">Source: OpenStax Textbook</a></p>
#
# + [markdown] cell_style="split" slideshow={"slide_type": "fragment"}
# If an electric field is applied across an insulator (for e.g., when placing this material between the plates of a capacitor), individual molecules become aligned with the electric field.
#
# <img src="../../fig8_20_b.jpeg" width=40%>
#
# <p style="text-align:right; font-size:30%;"><a href="https://openstax.org/books/university-physics-volume-2/pages/8-1-capacitors-and-capacitance#91635">Source: OpenStax Textbook</a></p>
# + [markdown] slideshow={"slide_type": "subslide"}
# Thus, the main electric field $E_0$ is "reduced" by the electric field in the opposite direction $E_i$ induced by the dielectric material.
#
# <img src="../../fig8_20_c.jpeg" width=20%>
#
# Reducing of the electric field results in a DECREASE in voltage ($V = E\cdot d$), and consequently, an INCREASE in capacitance ($C=Q/V$) since the charge Q on the plates is constant.
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Back to the Studfinder
# + [markdown] cell_style="split" slideshow={"slide_type": "fragment"}
# Now you have all the physics and intuition you need to work out how a stud finder works (hint: it uses capacitance and dielectrics)!
# <br><br>
# There is just one more key piece of insight, but it's "geometry"...
# <br><br><br>
# + [markdown] cell_style="split" slideshow={"slide_type": "fragment"}
# <center>
# <img src="assets/stud1.png" width=80%>
# </center>
# + [markdown] slideshow={"slide_type": "fragment"}
# ### Instructions: In break-out rooms, discuss and see if you can figure out how the stud finder works!
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Watch the magic (geometry)!
# + [markdown] slideshow={"slide_type": "fragment"}
# <center>
# <img src="../../stud.gif" width=50%>
# </center>
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Stud Finder
# <center>
# <img src="../../stud2.png" width=90%>
# </center>
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Stud Finder
#
# <center>
# <img src="../../stud3.png" width=90%>
# </center>
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Challenge Question
# + slideshow={"slide_type": "fragment"}
IFrame(capacitor_phet, width='100%',height=550)
# + [markdown] slideshow={"slide_type": "fragment"}
# Let's disconnect the battery, and then observe the voltage $V$ across the parallel plates after we decrease the plate separation $d$ and increase the plate area $A$.
# + [markdown] slideshow={"slide_type": "fragment"}
# Question: Can you explain how the voltage across the capacitor can be larger than the initially supplied voltage? Answer in 3-4 sentences.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Recap Learning Objectives
# + [markdown] slideshow={"slide_type": "fragment"}
# - Identify and summarize key functions of the capacitor
# - <span style="color:red">Capacitors store and release energy in a controlled manner</style>
# - <span style="color:red">Energy is stored within the electric field
# + [markdown] slideshow={"slide_type": "fragment"}
# - Explain the relationship between the capacitance and the plate separation and area
# - <span style="color:red">$C$ is inversely proportional to plate separation $d$</style>
# - <span style="color:red">$C$ is proportional to plate area $A$</style>
# + [markdown] slideshow={"slide_type": "fragment"}
# - Describe applications of the capacitor based on its properties
# - <span style="color:red">Defribrillator: Controlled storage and release of energy</style>
# - <span style="color:red">Stud finder: plates are planar and the stud (different dielectric) changes the capacitance </style>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Post-lecture Conceptual Questions
# + slideshow={"slide_type": "fragment"}
qs = display(Q1), display(Q2), display(Q3), display(Q4), display(Q5)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Next class
#
# - Quantifying the energy stored in a capacitor
# - Dielectric constant $\kappa$ of different materials
# - Adding capacitors in series and parallel
#
# + [markdown] slideshow={"slide_type": "fragment"}
# Reminder: In Tutorial this week, you will work through the derivation of the Capacitance in a parallel plate capacitor!
# <br><br><br>
# ### Have a great week!
# + [markdown] slideshow={"slide_type": "skip"}
# ## Appendix
# + [markdown] slideshow={"slide_type": "skip"}
# ### Deriving the E field within parallel plates using Gauss' Law
# + cell_style="center" slideshow={"slide_type": "skip"}
### Gauss' Law Derivation
IFrame('https://www.youtube-nocookie.com/embed/gTfEFwVsgKA', width=700,height=400)
| jupyterdays/capacitor_talk/Capacitor.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/WillMartin7/edgetpu-ggcnn/blob/main/convert_ggcnn.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="5AVHyl9OpIuz"
#
# # **Install libraries**
# Tensorflow nightly is needed. Sometimes the most recent version breaks the TFLiteConverter and an older version should be used.
# + colab={"base_uri": "https://localhost:8080/"} id="_E7LVaNDRq3n" outputId="f7d59156-86d1-40f8-fc8e-754c565dd09e"
# !pip install tf-nightly
# + [markdown] id="AuEOjIudNU3O"
# # **Add model and dataset to accessible files**
#
# Model file (e.g. epoch_50_model.hdf5) and dataset file (e.g. dataset_210516_1505.hdf5) need to be placed in google drive folder (e.g. ggcnn_files/) so they can be accessed.
# + colab={"base_uri": "https://localhost:8080/"} id="U4njHUTESRZ0" outputId="c11da4c7-a50f-4251-cd75-401706cf44c8"
# mount google drive to google colab
from google.colab import drive
drive.mount('/content/drive')
# + [markdown] id="MRFp-7FWMv2Y"
# # **Convert to TFLite model**
# + colab={"base_uri": "https://localhost:8080/"} id="8ze-x9b4TX3E" outputId="2ed20465-3f8e-41f9-aadc-f5d4849112c7"
import tensorflow as tf
from tensorflow import keras
import h5py
import numpy as np
# load dataset
dataset_fn = '/content/drive/MyDrive/ggcnn_files/dataset_210516_1505.hdf5'
f = h5py.File(dataset_fn, 'r')
#representative dataset
rep_imgs = np.array(f['test/rgb'], dtype=np.float32)
def representative_dataset():
for data in tf.data.Dataset.from_tensor_slices((rep_imgs)).batch(1).take(150):
yield [data]
# load model
model_checkpoint_fn = '/content/drive/MyDrive/ggcnn_files/epoch_50_model.hdf5'
model = keras.models.load_model(model_checkpoint_fn)
model.summary()
# convert to tflite model with integer only quantization
# https://www.tensorflow.org/lite/performance/post_training_quantization#integer_only
converter = tf.lite.TFLiteConverter.from_keras_model(model)
converter.optimizations = [tf.lite.Optimize.DEFAULT]
converter.representative_dataset = representative_dataset
converter.target_spec.supported_ops = [tf.lite.OpsSet.TFLITE_BUILTINS_INT8]
converter.inference_input_type = tf.int8 # or tf.uint8
converter.inference_output_type = tf.int8 # or tf.uint8
converter.experimental_new_converter = False
tflite_quant_model = converter.convert()
# save tflite model as ggcnn_model.tflite
with open('ggcnn_model.tflite', 'wb') as g:
g.write(tflite_quant_model)
# + [markdown] id="eM9MCEL9M3XH"
# # **Convert to Edge TPU model**
# + colab={"base_uri": "https://localhost:8080/"} id="rw33C3gFl-ay" outputId="4bb83008-d8af-4ad3-a576-80b5cfaa18fb"
# install edge tpu compiler: https://coral.ai/docs/edgetpu/compiler/
# !curl https://packages.cloud.google.com/apt/doc/apt-key.gpg | apt-key add -
# !echo "deb https://packages.cloud.google.com/apt coral-edgetpu-stable main" | sudo tee /etc/apt/sources.list.d/coral-edgetpu.list
# !sudo apt-get update
# !sudo apt-get install edgetpu-compiler
# + colab={"base_uri": "https://localhost:8080/"} id="XLKxBNPimNhX" outputId="a7e7b66e-37d0-45a4-f1f3-b29efa39a33a"
# convert and save model as ggcnn_model_edgetpu.tflite
# verify that all operators are mapped to TPU
# !edgetpu_compiler -s ggcnn_model.tflite
| convert_ggcnn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
pip install psycopg2-binary
# +
from sqlalchemy import create_engine
import pandas as pd
import psycopg2
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib inline
# -
from password import admin_key
engine = create_engine(f'postgresql+psycopg2://postgres:{admin_key}@localhost:5432/sql_challenge')
connection = engine.connect()
# +
sql = '''
SELECT *
FROM salaries
'''
salaries_df = pd.read_sql(sql, connection)
salaries_df.head()
# +
fig = plt.figure(figsize = (10,7))
plt.title('Salary Frequencies')
plt.xlabel("Salary")
plt.ylabel('Frequency')
plt.hist(salaries_df['salary'])
plt.grid()
plt.tight_layout()
plt.show()
plt.savefig("output/salaries_hist.png")
# -
#ranges
bins = [0,40000,50000,60000,70000,80000,90000,100000,130000]
labels = ['≤40', '40-50','50-60','60-70','70-80','80-90','90-100','≥100']
salaries_df['Salary range'] = pd.cut(salaries_df['salary'],bins=bins, labels=labels)
grouped_df = salaries_df.groupby('Salary range')
salary_range_df = grouped_df.count().iloc[:,1].reset_index().rename({'salary': 'count'},axis=1)
salary_range_df
# +
# Create a bar chart of average salary by title.
sql = '''
SELECT t.title, ROUND(AVG(s.salary), 2) AS "average_salary"
FROM titles t
INNER JOIN employees e ON t.title_id = e.emp_title_id
INNER JOIN salaries s ON e.emp_no = s.emp_no
GROUP BY t.title;
'''
avg_salaries = pd.read_sql(sql, connection)
avg_salaries.head(10)
# +
#plot average salaries
plot = avg_salaries.plot(kind = 'bar', figsize = (6,6))
# Set a title for the chart
plt.title("Average Salaries Per Title")
plot.set_xticklabels(avg_salaries['title'], rotation=55,horizontalalignment="right")
plt.show()
plt.tight_layout()
# -
| sqlchallenge_bonus_connect.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import requests
import json
import pandas as pd
import numpy as np
url_matomo = "https://stats.data.gouv.fr/index.php?module=API&\
format=json&idSite=145&filter_column=label&filter_pattern=selectedSiren%3D&showColumns=label,\
nb_events&method=Events.getName&secondaryDimension=eventAction&expanded=1&filter_limit=9999\
&period=year&date=2022-01-01"
response_matomo = requests.get(url_matomo)
print(response_matomo.url)
content = json.loads(response_matomo.content)
content
len(content)
df_data = pd.DataFrame(columns=["terms", "siren", "url"])
df_data
for index, item in enumerate(content):
siren = item["label"].split("selectedSiren=")[1][:9]
list_terms = []
for term in item["subtable"]:
list_terms.append(term["label"])
url = f"https://annuaire-entreprises.data.gouv.fr/entreprise/{siren}"
df_data = df_data.append(
{"terms": list_terms, "siren": str(siren), "url": url}, ignore_index=True
)
df_data.head(30)
df_data.to_csv("./elastic_test_set.csv", header=True, index=False)
df_test = pd.read_csv("./elastic_test_set.csv", dtype=str)
df_test.head(30)
| testing/get_matomo_logs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# import dependants
import csv
import os
# add file path
BankData = os.path.join("budget_data.csv")
# Set list variables
Profits = []
MonthlyChange = []
Date = []
# Set variables to zero
MonthCount = 0
TotalProfits = 0
Change = 0
StartingProfit = 0
with open(BankData, newline = "") as csvfile:
csvreader = csv.reader(csvfile, delimiter = ",")
csvheader = next(csvreader)
# Count the months in the sheet
for row in csvreader:
MonthCount = MonthCount + 1
# Set the date column
Date.append(row[0])
# Set the profits column
Profits.append(row[1])
# Calculate total profits
TotalProfits = TotalProfits + int(row[1])
# Calculate average change in profits
FinalProfit = int(row[1])
Monthly_Change= FinalProfit - StartingProfit
MonthlyChange.append(Monthly_Change)
Change = Change + Monthly_Change
StartingProfit = FinalProfit
ProfitChanges = round((Change/MonthCount))
GreatestIncrease = max(MonthlyChange)
IncreaseDate = Date[MonthlyChange.index(GreatestIncrease)]
GreatestLoss = min(MonthlyChange)
LossDate = Date[MonthlyChange.index(GreatestLoss)]
print("-----------------------------------------------")
print(" Financial Analysis")
print("-----------------------------------------------")
print("Total Months: " + str(MonthCount))
print("Total Profits: $" + str(TotalProfits))
print("Average Profit Change: $" + str(ProfitChanges))
print('Greatest Increase in Profits:' + str(IncreaseDate) +" ($" + str(GreatestIncrease) + ")")
print('Greatest Loss in Profits:' + str(LossDate) + '($' + str(GreatestLoss) + ')')
with open('Financial_Analysis.txt', 'w') as text:
text.write("-----------------------------------------------\n")
text.write(" Financial Analysis" + "\n")
text.write("-----------------------------------------------\n")
text.write("Total Months: " + str(MonthCount) + "\n")
text.write("Total Profits: $" + str(TotalProfits) + "\n")
text.write("Average Profit Change: $" +str(ProfitChanges) + "\n")
text.write('Greatest Increase in Profits:' + str(IncreaseDate) +" ($" + str(GreatestIncrease) + ")" + "\n")
text.write('Greatest Loss in Profits:' + str(LossDate) + '($' + str(GreatestLoss) + ')' + "\n")
# -
| pybank/Untitled1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
from utilities_namespace import *
# %%capture
# %load_ext rpy2.ipython
# %R require(ggplot2)
# To better understand the CMap 2.0 enrichment score I have answered this question: https://www.biostars.org/p/140821/#364663
#
# Below is the code that I used to reproduce the enrichment score calculation:
# +
from config import DATA_DIR
from pathlib import Path
data_path = Path(DATA_DIR)
# -
# Loading the CMap 2.0:
metadata = pd.read_excel(data_path / 'cmap/cmap_instances_02.xls')
substances = metadata[metadata.isna().sum(axis=1) <= 1].dropna(how='all').set_index('instance_id')
substances.head()
substances.columns
instances_ranks = {
5941: 174,
5968: 305,
5963: 339,
5936: 368
}
# +
n = 6100
t = len(instances_ranks)
dd = DataFrame([
{
'x': j,
'y': (j +1 ) / t,
'group': 'j/t',
'label': f'{i}: {substances.loc[i].cmap_name}\n{substances.loc[i].batch_id} batch, {substances.loc[i].cell2}',
'case': 'j/t - V(j)/n'
}
for j, (i, vj) in enumerate(instances_ranks.items())
] + [
{
'x': j,
'y': vj / n,
'group': 'V(j) / n',
'label': f'{i}: {substances.loc[i].cmap_name}\n{substances.loc[i].batch_id} batch, {substances.loc[i].cell2}',
'case': 'j/t - V(j)/n'
}
for j, (i, vj) in enumerate(instances_ranks.items())
] +
[
{
'x': j,
'y': (j) / t,
'group': '(j-1)/t',
'label': f'{i}: {substances.loc[i].cmap_name}\n{substances.loc[i].batch_id} batch, {substances.loc[i].cell2}',
'case': 'V(j)/n - (j-1)/t'
}
for j, (i, vj) in enumerate(instances_ranks.items())
] + [
{
'x': j,
'y': vj / n,
'group': 'V(j) / n',
'label': f'{i}: {substances.loc[i].cmap_name}\n{substances.loc[i].batch_id} batch, {substances.loc[i].cell2}',
'case': 'V(j)/n - (j-1)/t'
}
for j, (i, vj) in enumerate(instances_ranks.items())
]
)
# -
from numpy import argmax
show_table(dd)
1.000000 - 0.060328
0.750000 - 0.060328
# +
vj_j = [
vj/n - j/t
for j, (i, vj) in enumerate(instances_ranks.items())
]
j_vj = [
(j+1)/t - vj/n
for j, (i, vj) in enumerate(instances_ranks.items())
]
annotations = DataFrame([{
'x': argmax(vj_j) + 1 - 0.25,
'text': f'{max(vj_j) :.4f}',#[vj/n for j, (i, vj) in enumerate(instances_ranks.items())][argmax(vj_j)]
'y': max(vj_j) / 2,
'case': 'V(j)/n - (j-1)/t',
'group': 'max(V(j)/n - (j-1)/t)'
}, {
'x': argmax(j_vj) + 1,
'text': f'{max(j_vj):.4f}',#[vj/n for j, (i, vj) in enumerate(instances_ranks.items())][argmax(vj_j)]
'y': max(j_vj)/2,
'case': 'j/t - V(j)/n',
'group': 'max(j/t - V(j)/n)'
}
])
annotations
# -
segments = DataFrame([
dict(x=4.2, xend=4.2, y=0.060328, yend=1, group='max(j/t - V(j)/n)', label='', case='j/t - V(j)/n', color='max(j/t - V(j)/n)'),
dict(x=.92, xend=.92, y=0, yend=0.028525, group='max(V(j)/n - (j-1)/t)', label='', case='V(j)/n - (j-1)/t', color='max(V(j)/n - (j-1)/t)')
])
# + magic_args="-i dd -i annotations -w 700 -i segments -h 300" language="R"
# dd$label <- reorder(dd$label, dd$x)
# (
# ggplot(dd, aes(x=label, y=y, group=group, color=group))
# + geom_step()
# + facet_wrap(. ~ case)
# + geom_segment(aes(x=x, y=y,xend=xend,yend=yend,color=color, group=group), data=segments)
# + geom_text(aes(x=x, y=y, label=text, group=-Inf), data=annotations, angle=90)
# + xlab('')
# +ylab('')
# )
# -
vj = (Series([174, 305, 339, 368]) / 6100)
j = Series(range(1,5)) / 4
j
', '.join((j - vj).apply(lambda x: f'{x:.4f}').tolist())
| notebooks/Connectivity_Map_2.0/How_does_enrichment_in_CMap_2.0_work.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="-iA_qC5BAv6w"
# ### Load preprocessed data
# -
# Run the script that downloads and processes the MovieLens data.
# Uncomment it to run the download & processing script.
# +
# #!python ../src/download.py
# + colab={"base_uri": "https://localhost:8080/", "height": 272} colab_type="code" id="POjwTTneAv6y" outputId="b3acebb0-47b2-405c-eb40-5474b7aab5c2"
import numpy as np
from sklearn.model_selection import train_test_split
from torch import from_numpy
from torch.utils.data import DataLoader
from torch.utils.data import TensorDataset
from torch.utils.data import BatchSampler
from torch.utils.data import SequentialSampler
fh = np.load('data/dataset.npz')
# We have a bunch of feature columns and last column is the y-target
# Note pytorch is finicky about need int64 types
train_x = fh['train_x'].astype(np.int64)
train_y = fh['train_y']
# We've already split into train & test
X_test = fh['test_x'].astype(np.int64)
Y_test = fh['test_y']
X_train, X_val, Y_train, Y_val = train_test_split(train_x, train_y)
n_user = int(fh['n_user'])
n_item = int(fh['n_item'])
n_occu = int(fh['n_occu'])
# columns are user_id, item_id and other features
# we won't use the 3rd and 4th columns
print(X_train)
print(' ')
print(Y_train)
def dataloader(*arrs, batch_size=1024):
dataset = TensorDataset(*arrs)
arr_size = len(arrs[0])
bs = BatchSampler(SequentialSampler(range(arr_size)),
batch_size=batch_size, drop_last=False)
return DataLoader(dataset, batch_sampler=bs, shuffle=False)
train = dataloader(from_numpy(X_train), from_numpy(Y_train))
test = dataloader(from_numpy(X_test), from_numpy(Y_test))
val = dataloader(from_numpy(X_val), from_numpy(Y_val))
# -
from abstract_model import AbstractModel
# +
import torch
from torch import nn
import torch.nn.functional as F
import pytorch_lightning as pl
def l2_regularize(array):
return torch.sum(array ** 2.0)
class MF(AbstractModel):
def __init__(self, n_user, n_item, n_occu, k=18, c_ovector=1.0,
c_vector=1.0, c_bias=1.0, batch_size=128):
super().__init__()
self.k = k
self.n_user = n_user
self.n_item = n_item
self.c_bias = c_bias
self.c_vector = c_vector
self.c_ovector = c_ovector
self.batch_size = batch_size
self.save_hyperparameters()
# These are learned and fit by PyTorch
self.user = nn.Embedding(n_user, k)
self.item = nn.Embedding(n_item, k)
self.bias_user = nn.Embedding(n_user, 1)
self.bias_item = nn.Embedding(n_item, 1)
self.bias = nn.Parameter(torch.ones(1))
# **NEW: occupation vectors
self.occu = nn.Embedding(n_occu, k)
def forward(self, inputs):
# This is the most import function in this script
# These are the user indices, and correspond to "u" variable
user_id = inputs[:, 0]
# Item indices, correspond to the "i" variable
item_id = inputs[:, 1]
# vector user = p_u
vector_user = self.user(user_id)
# vector item = q_i
vector_item = self.item(item_id)
# Pull out biases
bias_user = self.bias_user(user_id).squeeze()
bias_item = self.bias_item(item_id).squeeze()
biases = (self.bias + bias_user + bias_item)
# NEW: occupation-item interaction
occu_id = inputs[:, 3]
vector_occu = self.occu(occu_id)
vector_user_occu = vector_user + vector_occu
# this is a dot product & a user-item interaction: p_u * q_i
ui_interaction = torch.sum(vector_user_occu * vector_item, dim=1)
prediction = ui_interaction + biases
return prediction
def loss(self, prediction, target):
# MSE error between target = R_ui and prediction = p_u * q_i
loss_mse = F.mse_loss(prediction, target.squeeze())
log = {"mse": loss_mse}
return loss_mse, log
def reg(self):
# Add new regularization to the biases
reg_bias_user = l2_regularize(self.bias_user.weight) * self.c_bias
reg_bias_item = l2_regularize(self.bias_item.weight) * self.c_bias
reg_user = l2_regularize(self.user.weight) * self.c_vector
reg_item = l2_regularize(self.item.weight) * self.c_vector
# Compute new occupation regularization
reg_occu = l2_regularize(self.occu.weight) * self.c_ovector
log = {"reg_user": reg_user, "reg_item": reg_item,
"reg_bias_user": reg_bias_user, "reg_bias_item": reg_bias_item,
"reg_occu": reg_occu
}
total = reg_user + reg_item + reg_bias_user + reg_bias_item + reg_occu
return total, log
# +
from pytorch_lightning.loggers.wandb import WandbLogger
k = 6
c_bias = 1e-3
c_vector = 1e-5
c_ovector = 1e-8
model = MF(n_user, n_item, n_occu,
k=k, c_bias=c_bias, c_vector=c_vector,
c_ovector=c_ovector)
# add a logger
logger = WandbLogger(name="03_mf", project="simple_mf")
trainer = pl.Trainer(max_epochs=100, logger=logger,
early_stop_callback=True,
progress_bar_refresh_rate=1)
# -
trainer.fit(model, train, val)
| notebooks/03 MF model with side-features.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.6 64-bit ('venv')
# metadata:
# interpreter:
# hash: 15a2b6a943499ec644758d74268c29165fcd196c0e2707856e04b7f4372516d9
# name: python3
# ---
# + [markdown] id="T2FKiBhp05aD"
# # RNNによるfxデータ予測
# !対数差分を取らずに予測
# ## 流れ
# - データ読み込み
# - データをtensorflow用(feature, target)に変換.depend: `delay`
# - モデル構築,コンパイル.depend: `delay, n_hidden`
# - 学習
# - 学習評価, テスト評価
# + id="H2tzYvc6z02w" executionInfo={"status": "ok", "timestamp": 1609405121546, "user_tz": -540, "elapsed": 855, "user": {"displayName": "\u7af9\u7530\u822a\u592a", "photoUrl": "", "userId": "07685491189999509781"}}
import tensorflow
from tensorflow.keras import models, layers, callbacks, optimizers
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
import sys
sys.path.append('../module')
from utils import load_fx_data, nrmse
np.random.seed(0)
# + [markdown] id="bcmaqNYG_7XR"
# # データ読み込み
# +
instrument_list = ['USD_JPY', 'GBP_JPY', 'EUR_JPY']
df_dict_train = load_fx_data(instrument_list, data_kind='train')
df_dict_test = load_fx_data(instrument_list, data_kind='test')
df_dict_train['EUR_JPY']
# +
train = df_dict_train['USD_JPY']['Close_ask'].values[1:]
test = df_dict_test['USD_JPY']['Close_ask'].values[1:]
print(f'shape: {train.shape, test.shape}')
# 正規化
train = (train - train.mean())/train.std()
test = (test - test.mean())/test.std()
# -
# 参考までに現在の値を明日の予測値とする場合の誤差
rough_nrmse = nrmse(train[1:],train[:-1], 1)
print(f'rough forecast nrmse: {rough_nrmse}')
# + [markdown] id="909EO1VvAR8h"
# ## RNN用feature, targetに変換
# + id="JzJ_jmWd0cEz" executionInfo={"status": "ok", "timestamp": 1609405122265, "user_tz": -540, "elapsed": 1550, "user": {"displayName": "\u7af9\u7530\u822a\u592a", "photoUrl": "", "userId": "07685491189999509781"}}
# RNN用feature, targetを生成 in: raw_data, delay out: feature, targets
def convert_data_for_RNN(raw_data, delay=25):
"""
targetに対してdelay個の過去データをfeatureとする
"""
features = np.array([raw_data[n:n+delay] for n in range(len(raw_data)-delay)]).reshape(-1, delay,1)
targets = raw_data[delay:].reshape(-1,1,1)
return features, targets
# + colab={"base_uri": "https://localhost:8080/"} id="eq1YHZ_p_5Ka" executionInfo={"status": "ok", "timestamp": 1609405122265, "user_tz": -540, "elapsed": 1545, "user": {"displayName": "\u7af9\u7530\u822a\u592a", "photoUrl": "", "userId": "07685491189999509781"}} outputId="6802ac89-1459-45cc-c131-d3cab71efa9e"
delay = 25
train_features, train_targets = convert_data_for_RNN(train, delay=delay)
test_features, test_targets = convert_data_for_RNN(test, delay=delay)
print(f'train: {train_features.shape, train_targets.shape}')
print(f'test: {test_features.shape, test_targets.shape}')
# + [markdown] id="EI2BtifmAY8-"
# # モデル作成
# -
train_std = train.std()
print(train_std)
# + colab={"base_uri": "https://localhost:8080/"} id="auI34MRQ0iab" executionInfo={"status": "ok", "timestamp": 1609405337779, "user_tz": -540, "elapsed": 677, "user": {"displayName": "\u7af9\u7530\u822a\u592a", "photoUrl": "", "userId": "07685491189999509781"}} outputId="93362488-2037-4daf-df1b-9653180f865d"
# モデル構築
"""
n_hidden:
dropout, recurrent_dropout
regulalizer
deep化
"""
print(f'delay: {delay}') # 時間遅れstep
out_size = 1 # targetのsize
n_hidden = 300 # 中間層のノード数
model = models.Sequential(name='RNN')
model.add(layers.SimpleRNN(n_hidden, input_shape=(delay, out_size), return_sequences=False, dropout=0., recurrent_dropout=0.,))
model.add(layers.Dense(out_size, activation='linear'))
model.compile(loss='mean_squared_error', optimizer=optimizers.Adam(learning_rate=1e-3))
model.summary()
# + colab={"base_uri": "https://localhost:8080/"} id="mlYCRDvJ0lFN" executionInfo={"status": "ok", "timestamp": 1609405346051, "user_tz": -540, "elapsed": 7440, "user": {"displayName": "\u7af9\u7530\u822a\u592a", "photoUrl": "", "userId": "07685491189999509781"}} outputId="e9f1fe3d-0a3f-4095-a879-ff9811dd069d"
# 学習
callbacks.EarlyStopping()
model.fit(train_features,
train_targets,
batch_size=20,
epochs=10,
validation_split=0.2,
callbacks=[callbacks.EarlyStopping(monitor='val_loss', patience=5, mode='min')]
)
# + colab={"base_uri": "https://localhost:8080/", "height": 305} id="dSCG1wAN0tlv" executionInfo={"status": "ok", "timestamp": 1609405348348, "user_tz": -540, "elapsed": 1437, "user": {"displayName": "\u7af9\u7530\u822a\u592a", "photoUrl": "", "userId": "07685491189999509781"}} outputId="ba504d0c-2aa6-4cb8-d827-b71ae009f84d"
# fittingの確認
train_predict = model.predict(train_features)
start = 1200
fig = plt.figure()
plt.plot(train_targets[start:, 0,0], label='train targets')
plt.plot(train_predict[start:], label='train predict')
plt.legend()
plt.title('USD JPY Close Ask RNN')
model.evaluate(train_features, train_targets)
train_nrmse = nrmse(train_targets, train_predict, train_std)
print(f'nrmse: {train_nrmse}')
# fig.savefig('../data/img/usd_jpy_close_ask_rnn.png')
fig.savefig('../data/img/usd_jpy_close_ask_rnn_detail.png')
# + colab={"base_uri": "https://localhost:8080/", "height": 305} id="FejQzBPQ034j" executionInfo={"status": "ok", "timestamp": 1609405350313, "user_tz": -540, "elapsed": 999, "user": {"displayName": "\u7af9\u7530\u822a\u592a", "photoUrl": "", "userId": "07685491189999509781"}} outputId="afafec25-cb43-44ec-8f14-924d0d53195d"
# 未来の予測
start = 0
test_predict = model.predict(test_features)
fig = plt.figure()
plt.plot(test_targets[start:,0,0], label='test target')
plt.plot(test_predict[start:], label='test pred')
plt.legend()
plt.title('USD JPY Close Ask RNN detail')
model.evaluate(test_features, test_targets)
test_nrmse = nrmse(test_targets, test_predict, 1)
print(f'nrmse: {test_nrmse}')
# + [markdown] id="_vddfE2gCGir"
# ## 2021/01/10 観察
# - 隠れ層のノード数は100以上にすると結果の振動が小さくなる?
# - 評価方法はtrainのstdで正規化したrmseがいいかも
# -
# ## 2021/01/17 観察
# - 1step前の状態をそのまま予測値にしているのとほとんど変わらない.(25,30)
# ## パラメータを変えてみる
# パラメータ
# - `delay`: 時間遅れ
# - `n_hidden`: 中間層ノード数
# - dropout(オプション)
# - `dropout`
# - `recurrent_dropout`
#
# 条件
# - パラメータ:
# - `delay`: `[10,25,50,100]`
# - `n_hidden`: `[5,10,25,50,100,200]`
# - 学習:
# - `epoch=30`
# - `batch_size=20`
# - `learning_rate=1e-3`
delays = [10, 25, 50, 100, 200]
n_hiddens = [5, 10, 25, 50, 100, 200,300]
nrmse_results = np.zeros((len(delays), len(n_hiddens)))
for i, delay in enumerate(delays):
print(f'delay: {delay}') # 時間遅れstep
# 時間遅れ
train_features, train_targets = convert_data_for_RNN(train, delay=delay)
test_features, test_targets = convert_data_for_RNN(test, delay=delay)
print(f'train: {train_features.shape, train_targets.shape}')
print(f'test: {test_features.shape, test_targets.shape}')
for j, n_hidden in enumerate(n_hiddens):
print(f'n_hidden: {n_hidden}')
# モデル構築
model = models.Sequential(name='RNN')
model.add(layers.SimpleRNN(n_hidden, input_shape=(delay, out_size), return_sequences=False, dropout=0., recurrent_dropout=0.,))
model.add(layers.Dense(out_size, activation='linear'))
model.compile(loss='mean_squared_error', optimizer=optimizers.Adam(learning_rate=1e-3))
# model.summary()
# 学習
callbacks.EarlyStopping()
model.fit(train_features,
train_targets,
batch_size=20,
epochs=30,
validation_split=0.2,
callbacks=[callbacks.EarlyStopping(monitor='val_loss', patience=5, mode='min')],
verbose=0
)
train_predict = model.predict(train_features)
train_nrmse = nrmse(train_targets, train_predict, train_std)
print(f'nrmse: {train_nrmse}')
nrmse_results[i,j] = train_nrmse
print('---------------------')
print('=========================')
# df = pd.DataFrame(nrmse_results, index=delays, columns=n_hiddens)
# df.to_csv('../data/csv/delay_nhidden_nrmse.csv')
df = pd.read_csv('../data/csv/delay_nhidden_nrmse.csv', header=0, index_col=0)
fig = plt.figure()
sns.heatmap(df, annot=True, vmax=rough_nrmse)
plt.title('Grid Search for Delay and N_hidden')
plt.xlabel('delay')
plt.ylabel('n_hidden')
fig.savefig('../data/img/delay_nhidden_nrmse.png')
| forecast/rnn_to_fx.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import geopandas as gpd
import matplotlib.pyplot as plt
import sys
sys.path.append('../')
# -
from src.d01_data.block_data_api import BlockDataApi
from src.d02_intermediate import block_data_cleaning as b_clean
from src.d02_intermediate.classifier_data_api import ClassifierDataApi
# # Exploratory Data Analysis of Block Data
# Loading the data:
block_data_api = BlockDataApi()
classifier_data_api = ClassifierDataApi()
SFHA_block_df = block_data_api.get_data(sfha=True)
block_df = block_data_api.get_data(sfha=False)
field_descriptions_df = block_data_api.get_fields()
frl_df_raw = block_data_api.get_data(frl=True)
frl_fields = block_data_api.get_fields(frl=True)
full_data = classifier_data_api.get_block_data(pct_frl=True)
# ## SFHA Block Data
# We want to know whether the SFHA dataset is redundant or it provides relevant information. We can see that the information it contains is entirely contained in the main Block dataset, so we can ignore it:
SFHA_fields = list(SFHA_block_df.columns)
SFHA_fields
# +
block_df_new = block_df.filter(['ID','Block','SFHA_Hsng', "SA_Hsng"], axis=1)
SFHA_block_df_new = SFHA_block_df.filter(['ID','Block','SFHA_Hsng', "SA_Hsng"], axis=1)
merge_df = pd.merge(SFHA_block_df_new, block_df_new, on=['ID'], how='inner')
merge_df.head()
# -
# Any difference would appear here (can be tried with any other pair of columns):
count = 0
L = list(merge_df["SA_Hsng_x"] == merge_df["SA_Hsng_y"])
for i in range(len(L)):
if L[i] == False:
print(i)
count += 1
# We are good! We do not need to use the SFHA dataset!
# ## Block Data
pd.set_option("max_rows", None)
field_descriptions_df[["Field Name", "Field Description", "Data Source"]]
# Grouping the columns in useful "thematic" groups:
#
# 1. IDENTIFICATION: Reflects geographic characteristics and id numbers (Census Tracts, area, FIPS, block type)
# 2. CURRENT: Columns referring to current CTIP assignment and simulations of variations in the exisiting model by the district
# 3. POPULATION: Demographic information such as population by age and enrollment in schools, also parents educational level and language status
# 4. ETHNICITY: Information on ethnicity of residents and students
# 5. ETHNICITY_DETAILED: Breakdown of ethnicity by grade, detailed ethnic group, and year (district)
# 6. ETHNICITY_DETAILED_GROUP: Above data but grouped for subsequent grades
# 6. INCOME: Data referrent to income and wealth of block and families
# 7. TEST SCORES: Academic data on CST and SBAC
# 8. HOUSING: San Francisco and Federal Hosuing Authority information
#
# To retrieve the dictionary we can use the appropriate method in the block api class
group_dict = block_data_api.get_classification()
group_dict
# BIG REMARK: Some columns are empty (or non-informative)!
# Search for a specific description:
name = "NH White students 2006-2010 K-8"
field_descriptions_df.loc[field_descriptions_df["Field Name"] == name, "Field Description"].iloc[0]
# Verifying if the field names and block data all match:
block_fields = list(block_df.columns)
field_fields = list(field_descriptions_df["Field Name"].values)
# +
def Diff(li1, li2):
return list(set(li1) - set(li2))
Diff(block_fields, field_fields)
# -
Diff(field_fields, block_fields)
# The FRL column is a mystery, and so is the DATA in the block dataset. All other mismatches have been fixed in the initialization on the class.
# ## Focal Students Data
# +
focal_columns = ["n", "nFocal", "nAALPI", "nFRL", "nBoth", "pctFocal", "pctAALPI", "pctFRL", "pctBoth"]
focal_data = full_data[focal_columns]
focal_data_map = classifier_data_api.get_map_df_data(cols=focal_columns)
# -
# This focal student dataset is extremely big. The total number of focal students amounts to over 60% of the student body. The intersection students are a better dataset to consider, but it is still too large:
# +
s_df = focal_data.sum()
print("Percentage of focal students in SF: {:.2%}".format(s_df["nFocal"]/s_df["n"]))
print("Percentage of intersection focal students in SF: {:.2%}".format(s_df["nBoth"]/s_df["n"]))
# -
# Most blocks are very heterogeneous. The median focal student percentage in a block is 50%, whereas the median intersectional focal students per block is 3%. This means the classification will have many false positives.
focal_data.median()
# We can view the distribution of percentages per bloc as histograms to understand the trade-offs:
ax = focal_data.hist(column=["pctBoth", "pctFocal"], grid=False, bins=20,
layout=(2,1), figsize=(20,30),
ylabelsize=20, xlabelsize=20,
sharey=True)
# We can try to remove the zero and one blocks:
heterogeneous_focal_data = focal_data[focal_data["nBoth"] > 0]
heterogeneous_focal_data.median()["pctBoth"]
ax = heterogeneous_focal_data.hist(column=["pctBoth"], grid=False, bins=20,
figsize=(20,15),
ylabelsize=20, xlabelsize=20,
sharey=True)
| notebooks/20210624-gsa-block_eda.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import os, sys, io
import json
from pprint import pprint
import codecs
# -
data_path = '../../data'
# +
# Given the juristidction, file type and root path to data
# Returns a list of case ids in that jurisdiction
def get_cases_in_jurisdiction( juris_abv = 'nced', file_type = 'opinions', data_path = '../../data'):
# path leading to the jurisdiction files
path = data_path + '/'+ file_type + '/' + juris_abv + '/'
# TODO: throw an exception
# Check that the directory exists
if not os.path.isdir(path):
print 'not a legal path'
return []
else:
return [int(f.split('.json')[0]) for f in os.listdir(path)]
# -
nced_case_ids = get_cases_in_jurisdiction('nced')
# +
def get_case_attributes(cl_file, op_file):
print 'blah'
# -
cl_file = data_path + '/clusters/nced/1361899.json'
op_file = data_path + '/opinions/nced/1361899.json'
# +
# Open the cluster and opinion json files
with open(cl_file) as data_file:
cl_data_temp = json.load(data_file)
with open(op_file) as data_file:
op_data_temp = json.load(data_file)
# TODO: do this more succinctly
# Convert to utf8 from unicode
cl_data = {}
for k in cl_data_temp.keys():
value = cl_data_temp[k]
if type(value) == unicode:
cl_data[k.encode('utf8')] = value.encode('utf8')
else:
cl_data[k.encode('utf8')] = value
op_data = {}
for k in op_data_temp.keys():
value = op_data_temp[k]
if type(value) == unicode:
op_data[k.encode('utf8')] = value.encode('utf8')
else:
op_data[k.encode('utf8')] = value
# -
pprint(cl_data)
| python_code/ipynb/.ipynb_checkpoints/case_class-checkpoint.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .groovy
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Groovy
// language: groovy
// name: groovy
// ---
// + deletable=true editable=true
import com.twosigma.beaker.table.*
import com.twosigma.beaker.fileloader.CsvPlotReader
new TableDisplay( new CsvPlotReader().readAsList("tableRows.csv"))
// + deletable=true editable=true
display
// + deletable=true editable=true
| demoFiles/tableApi.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lab 5: NLI with Attention
# In this assignment we train an attention model for NLI based on [Rocktäschel's](https://arxiv.org/pdf/1509.06664v4.pdf) initial (non word-by-word) model.
# ## Setup
# You'll need to download and unzip SNLI, which you can find [here](http://nlp.stanford.edu/projects/snli/). Set `snli_home` below to point to it. The following block of code loads it.
#
# This will take a couple of minutes to run if you load whole data, so you may want to start editing the code below at the same time. As it is now, the code is ready for debugging, it only loads 1000 examples.
#
# You'll use Glove embeddings as well. If you dowloaded for the assignment 3 then edit `glove_home` to point the correct embedding file. Otherwise, if need to download them, You can do it at the following [url](http://nlp.stanford.edu/data/glove.6B.zip) (1GB). The zip file includes embeddings of different dimensionality (50d, 100d, 200d, 300d) for a vocabulary of 400000 words. Decompress them and place somewhere, for example in `./embeddings/` folder.
# +
snli_home = './data/snli_1.0/'
import re
import random
import json
LABEL_MAP = {
"entailment": 0,
"neutral": 1,
"contradiction": 2
}
def load_snli_data(path):
data = []
with open(path) as f:
for i, line in enumerate(f):
if i >= 1000000: # Edit to use less data for debugging. set to 1000000 for testing.
break
loaded_example = json.loads(line)
if loaded_example["gold_label"] not in LABEL_MAP:
continue
loaded_example["label"] = LABEL_MAP[loaded_example["gold_label"]]
data.append(loaded_example)
random.seed(1)
random.shuffle(data)
return data
training_set = load_snli_data(snli_home + '/snli_1.0_train.jsonl')
dev_set = load_snli_data(snli_home + '/snli_1.0_dev.jsonl')
test_set = load_snli_data(snli_home + '/snli_1.0_test.jsonl')
# -
# Next, we'll convert the data to index vectors in the same way that we've done for in-class exercises with RNN-based sentiment models. A few notes:
#
# - We use a sequence length of only 10, which is short enough that we're truncating a large fraction of sentences.
# - Tokenization is easy here because we're relying on the output of a parser (which does tokenization as part of parsing), just as with the SST corpus that we've been using until now. Note that we use the 'sentence1_binary_parse' field of each example rather than the human-readable 'sentence1'.
# - We're using a moderately large vocabulary (for a class exercise) of about 36k words.
# +
SEQ_LEN = 10
import collections
import numpy as np
def sentences_to_padded_index_sequences(datasets):
'''Annotates datasets with feature vectors.'''
PADDING = "<PAD>"
UNKNOWN = "<UNK>"
# Extract vocabulary
def tokenize(string):
string = re.sub(r'\(|\)', '', string)
return string.lower().split()
word_counter = collections.Counter()
for example in datasets[0]:
word_counter.update(tokenize(example['sentence1_binary_parse']))
word_counter.update(tokenize(example['sentence2_binary_parse']))
vocabulary = set([word for word in word_counter])
vocabulary = list(vocabulary)
vocabulary = [PADDING, UNKNOWN] + vocabulary
word_indices = dict(zip(vocabulary, range(len(vocabulary))))
indices_to_words = {v: k for k, v in word_indices.items()}
for i, dataset in enumerate(datasets):
for example in dataset:
for sentence in ['sentence1_binary_parse', 'sentence2_binary_parse']:
example[sentence + '_index_sequence'] = np.zeros((SEQ_LEN), dtype=np.int32)
token_sequence = tokenize(example[sentence])
padding = SEQ_LEN - len(token_sequence)
for i in range(SEQ_LEN):
if i >= padding:
if token_sequence[i - padding] in word_indices:
index = word_indices[token_sequence[i - padding]]
else:
index = word_indices[UNKNOWN]
else:
index = word_indices[PADDING]
example[sentence + '_index_sequence'][i] = index
return indices_to_words, word_indices
indices_to_words, word_indices = sentences_to_padded_index_sequences([training_set, dev_set, test_set])
# -
print(training_set[6])
print(len(word_indices))
# Now we load GloVe. You'll need to decompress the file in 'data' folder. You'll need the same file that you used for the in-class exercise on word embeddings.
# +
glove_home = './data/'
words_to_load = 45000
with open(glove_home + 'glove.6B.50d.txt') as f:
loaded_embeddings = np.zeros((len(word_indices), 50), dtype='float32')
for i, line in enumerate(f):
if i >= words_to_load:
break
s = line.split()
if s[0] in word_indices:
loaded_embeddings[word_indices[s[0]], :] = np.asarray(s[1:])
word = 'dog'
word_ind = word_indices[word]
print('Loaded embedding for {}:\n{}'.format(word,loaded_embeddings[word_ind]))
# -
# Now we set up an evaluation function as before.
def evaluate_classifier(classifier, eval_set):
correct = 0
hypotheses = classifier(eval_set)
for i, example in enumerate(eval_set):
hypothesis = hypotheses[i]
if hypothesis == example['label']:
correct += 1
return correct / float(len(eval_set))
# ## Assignments: Build GRU pair with attention
# Run the first block of code right away to make sure you have the proper dependencies. If you are working on your machine, you may need to install [matplotlib](http://matplotlib.org/users/installing.html), which should be fairly straightforward.
#
# ### TODO:
# - Fill in the missing componant below to complete Rocktäschel-style attention. Details of the attention mechanism are described in Section 2.3 of the [paper](https://arxiv.org/pdf/1509.06664v4.pdf).
#
# 
# - More specifically you need to implement the following two equations:
# 
# %matplotlib inline
import matplotlib.pyplot as plt
import tensorflow as tf
class RNNEntailmentClassifier:
def __init__(self, vocab_size, sequence_length):
# Define the hyperparameters
self.training_epochs = 20 # How long to train for - chosen to fit within class time
self.display_epoch_freq = 1 # How often to print out cost statistics (in epochs)
self.display_step_freq = 250 # How often to test (in steps)
self.dim = 24 # The dimension of the hidden state of the RNN
self.embedding_dim = 50 # The dimension of the learned word embeddings
self.batch_size = 64 # Somewhat arbitrary - can be tuned, but often tune for speed, not accuracy
self.vocab_size = vocab_size # Defined by the file reader above
self.sequence_length = sequence_length # Defined by the file reader above
# Define the parameters
self.E = tf.Variable(loaded_embeddings, trainable=False)
self.W_rnn = {}
self.W_r = {}
self.W_z = {}
self.b_rnn = {}
self.b_r = {}
self.b_z = {}
for name in ['p', 'h']:
in_dim = self.embedding_dim
self.W_rnn[name] = tf.Variable(tf.random_normal([in_dim + self.dim, self.dim], stddev=0.1))
self.b_rnn[name] = tf.Variable(tf.random_normal([self.dim], stddev=0.1))
self.W_r[name] = tf.Variable(tf.random_normal([in_dim + self.dim, self.dim], stddev=0.1))
self.b_r[name] = tf.Variable(tf.random_normal([self.dim], stddev=0.1))
self.W_z[name] = tf.Variable(tf.random_normal([in_dim + self.dim, self.dim], stddev=0.1))
self.b_z[name] = tf.Variable(tf.random_normal([self.dim], stddev=0.1))
# TODO: You'll need to use these three parameters.
self.W_h_attn = tf.Variable(tf.random_normal([self.dim, self.dim], stddev=0.1))
self.W_y_attn = tf.Variable(tf.random_normal([self.dim, self.dim], stddev=0.1))
self.w_attn = tf.Variable(tf.random_normal([self.dim, 1], stddev=0.1))
self.W_combination = tf.Variable(tf.random_normal([2 * self.dim, self.dim], stddev=0.1))
self.b_combination = tf.Variable(tf.random_normal([self.dim], stddev=0.1))
self.W_cl = tf.Variable(tf.random_normal([self.dim, 3], stddev=0.1))
self.b_cl = tf.Variable(tf.random_normal([3], stddev=0.1))
# Define the placeholders
self.premise_x = tf.placeholder(tf.int32, [None, self.sequence_length])
self.hypothesis_x = tf.placeholder(tf.int32, [None, self.sequence_length])
self.y = tf.placeholder(tf.int32, [None])
# Define the GRU function
def gru(emb, h_prev, name):
emb_h_prev = tf.concat([emb, h_prev], 1, name=name + '_emb_h_prev')
z = tf.nn.sigmoid(tf.matmul(emb_h_prev, self.W_z[name]) + self.b_z[name], name=name + '_z')
r = tf.nn.sigmoid(tf.matmul(emb_h_prev, self.W_r[name]) + self.b_r[name], name=name + '_r')
emb_r_h_prev = tf.concat([emb, r * h_prev], 1, name=name + '_emb_r_h_prev')
h_tilde = tf.nn.tanh(tf.matmul(emb_r_h_prev, self.W_rnn[name]) + self.b_rnn[name], name=name + '_h_tilde')
h = (1. - z) * h_prev + z * h_tilde
return h
# Define one step of the premise encoder RNN
def premise_step(x, h_prev):
emb = tf.nn.embedding_lookup(self.E, x)
return gru(emb, h_prev, 'p')
# Define one step of the hypothesis encoder RNN
def hypothesis_step(x, h_prev):
emb = tf.nn.embedding_lookup(self.E, x)
return gru(emb, h_prev, 'h')
# Split up the inputs into individual tensors
self.x_premise_slices = tf.split(self.premise_x, self.sequence_length, 1)
self.x_hypothesis_slices = tf.split(self.hypothesis_x, self.sequence_length, 1)
self.h_zero = tf.zeros(tf.stack([tf.shape(self.premise_x)[0], self.dim]))
# Unroll the first RNN
premise_h_prev = self.h_zero
premise_steps_list = []
for t in range(self.sequence_length):
x_t = tf.reshape(self.x_premise_slices[t], [-1])
premise_h_prev = premise_step(x_t, premise_h_prev)
premise_steps_list.append(premise_h_prev)
premise_steps = tf.stack(premise_steps_list, axis=1, name='premise_steps')
# Unroll the second RNN
h_prev_hypothesis = premise_h_prev # Continue running the same RNN
for t in range(self.sequence_length):
x_t = tf.reshape(self.x_hypothesis_slices[t], [-1])
h_prev_hypothesis = hypothesis_step(x_t, h_prev_hypothesis)
# Do attention
wm_list = []
wh_hn=tf.matmul(h_prev_hypothesis,self.W_h_attn)
# TODO: Fill wm_list with one scalar (a.k.a., vector with one scalar for each batch entry)
# for each word in the premise.
# This'll likely be easiest if you use a loop to iterate over timesteps.
# You should use the three `attn` parameters defined above, as well as `premise_steps_list`.
for t in range(self.sequence_length):
y = premise_steps_list[t]
M = tf.tanh(tf.matmul(y,self.W_y_attn) + wh_hn)
attention_t = tf.matmul(M,self.w_attn)
wm_list.append(attention_t)
# End above TODO
wm = tf.stack(wm_list, axis=1)
self.attn_weights = tf.nn.softmax(wm, dim=1)
attn_result = tf.reduce_sum(tf.multiply(self.attn_weights, premise_steps, name='attn_result_unsummed'),
1, name='attn_result')
# Combine the results of attention with the final GRU state
concat_features = tf.concat([attn_result, h_prev_hypothesis], 1, name=name + '_emb_h_prev')
pair_features = tf.nn.tanh(tf.matmul(concat_features, self.W_combination) + self.b_combination)
# Compute the logits
self.logits = tf.matmul(pair_features, self.W_cl) + self.b_cl
# Define the cost function (here, the softmax exp and sum are built in)
self.total_cost = tf.reduce_mean(tf.nn.sparse_softmax_cross_entropy_with_logits(logits=self.logits, labels=self.y))
# This performs the main SGD update equation with gradient clipping
optimizer_obj = tf.train.AdamOptimizer()
gvs = optimizer_obj.compute_gradients(self.total_cost)
capped_gvs = [(tf.clip_by_norm(grad, 5.0), var) for grad, var in gvs if grad is not None]
self.optimizer = optimizer_obj.apply_gradients(capped_gvs)
# Create an operation to fill zero values in for W and b
self.init = tf.global_variables_initializer()
# Create a placeholder for the session that will be shared between training and evaluation
self.sess = tf.Session()
# Initialize variables
self.sess.run(self.init)
self.step = 1
self.epoch = 1
def train(self, training_data, dev_data):
def get_minibatch(dataset, start_index, end_index):
indices = range(start_index, end_index)
premise_vectors = np.vstack([dataset[i]['sentence1_binary_parse_index_sequence'] for i in indices])
hypothesis_vectors = np.vstack([dataset[i]['sentence2_binary_parse_index_sequence'] for i in indices])
labels = [dataset[i]['label'] for i in indices]
return premise_vectors, hypothesis_vectors, labels
print('Training.')
# Training cycle
for _ in range(self.training_epochs):
random.shuffle(training_data)
avg_cost = 0.
total_batch = int(len(training_data) / self.batch_size)
# Loop over all batches in epoch
for i in range(total_batch):
# Assemble a minibatch of the next B examples
minibatch_premise_vectors, minibatch_hypothesis_vectors, minibatch_labels = get_minibatch(
training_data, self.batch_size * i, self.batch_size * (i + 1))
# Run the optimizer to take a gradient step, and also fetch the value of the
# cost function for logging
_, c = self.sess.run(
[self.optimizer, self.total_cost],
feed_dict={self.premise_x: minibatch_premise_vectors,
self.hypothesis_x: minibatch_hypothesis_vectors,
self.y: minibatch_labels})
if self.step % self.display_step_freq == 0:
print("Step:", self.step, "Dev acc:", evaluate_classifier(self.classify, dev_data[0:1000]), \
"Train acc:", evaluate_classifier(self.classify, training_data[0:1000]))
self.step += 1
avg_cost += c / (total_batch * self.batch_size)
# Display some statistics about the step
# Evaluating only one batch worth of data -- simplifies implementation slightly
if self.epoch % self.display_epoch_freq == 0:
print ("Epoch:", self.epoch, "Cost:", avg_cost)
self.epoch += 1
def classify(self, examples):
# This classifies a list of examples
premise_vectors = np.vstack([example['sentence1_binary_parse_index_sequence'] for example in examples])
hypothesis_vectors = np.vstack([example['sentence2_binary_parse_index_sequence'] for example in examples])
logits = self.sess.run(self.logits,
feed_dict={self.premise_x: premise_vectors,
self.hypothesis_x: hypothesis_vectors})
return np.argmax(logits, axis=1)
def get_attn(self, examples):
premise_vectors = np.vstack([example['sentence1_binary_parse_index_sequence'] for example in examples])
hypothesis_vectors = np.vstack([example['sentence2_binary_parse_index_sequence'] for example in examples])
attn_weights = self.sess.run(self.attn_weights,
feed_dict={self.premise_x: premise_vectors,
self.hypothesis_x: hypothesis_vectors})
return np.reshape(attn_weights, [len(examples), 10, 1])
def plot_attn(self, examples):
attn_weights = self.get_attn(examples)
for i in range(len(examples)):
fig = plt.figure()
ax = fig.add_subplot(111)
ax.matshow(np.transpose(attn_weights[i,:,:]), vmin=0., vmax=1., cmap=plt.cm.inferno)
premise_tokens = [indices_to_words[index] for index in examples[i]['sentence1_binary_parse_index_sequence']]
hypothesis_tokens = [indices_to_words[index] for index in examples[i]['sentence2_binary_parse_index_sequence']]
plt.text(0, 1, 'H: ' + ' '.join(hypothesis_tokens))
ax.set_xticklabels(premise_tokens, rotation=45)
plt.xticks(np.arange(0, 10, 1.0))
# Next, create an instance of the model. Unlike in previous exercises, initialization happens here, rather than at the start of training. You can now initialize a model once and start and stop training as needed.
classifier = RNNEntailmentClassifier(len(word_indices), SEQ_LEN)
# In implementing attention, it's easy to accidentally mix information between different examples in a batch. This assertion will fail if you've done so. Run it whenever you edit core model code.
assert (classifier.get_attn(training_set[0:2])[0, :, :] == \
classifier.get_attn(training_set[0:3])[0, :, :]).all(), \
'Warning: There is cross-example information flow.'
# Running the training long enough __on the whole training set__, you may get around 62% dev accuracy. This should take around five or ten minutes (or maybe more). This isn't great performance, but it's good enough that we should start to see attention play a role. If you have extra time, run longer.
classifier.train(training_set, dev_set)
# ## Visualization
# This will print some (NYU-colored) visualizations for the first ten dev examples. Explore these examples and more, and see if you can identify any patterns in what the model has learned.
classifier.plot_attn(dev_set[0:50])
# # Atribution:
# Adapted by <NAME>, based on a notebook by <NAME> at NYU
| labs/lab5/Attention Model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
from __future__ import division
import tensorflow as tf
import numpy as np
import tarfile
import os
import matplotlib.pyplot as plt
import time
# +
def csv_to_numpy_array(filePath, delimiter):
return np.genfromtxt(filePath, delimiter=delimiter, dtype='float32')
def import_data():
if "data" not in os.listdir(os.getcwd()):
# Untar directory of data if we haven't already
tarObject = tarfile.open("data.tar.gz")
tarObject.extractall()
tarObject.close()
print("Extracted tar to current directory")
else:
# we've already extracted the files
pass
print("loading training data")
trainX = csv_to_numpy_array("data/trainX.csv", delimiter=",")
trainY = csv_to_numpy_array("data/trainY.csv", delimiter=",")
print("loading test data")
testX = csv_to_numpy_array("data/testX.csv", delimiter=",")
testY = csv_to_numpy_array("data/testY.csv", delimiter=",")
return trainX,trainY,testX,testY
trainX,trainY,testX,testY = import_data()
# -
def accuracy(predictions, labels):
return (100.0 * np.sum(np.argmax(predictions, 1) == np.argmax(labels, 1))
/ predictions.shape[0])
trainX.shape
# +
num_hidden_nodes1 = 2000
num_hidden_nodes2 = 1000
num_hidden_nodes3 = 256
keep_prob = 0.5
# numFeatures = the number of words extracted from each email
numFeatures = trainX.shape[1]
# numLabels = number of classes we are predicting (here just 2: Spam or Ham)
numLabels = trainY.shape[1]
graph = tf.Graph()
with graph.as_default():
# Input data.
tf_train_dataset = tf.constant(trainX)
tf_train_labels = tf.constant(trainY)
tf_test_dataset = tf.constant(testX)
# Single mail input.
tf_mail = tf.placeholder(tf.float32, shape=(1, numFeatures))
# Variables.
weights1 = tf.Variable(tf.truncated_normal([numFeatures, num_hidden_nodes1],
stddev=np.sqrt(2.0 / (numFeatures))),name="v1")
biases1 = tf.Variable(tf.zeros([num_hidden_nodes1]),name="v2")
weights2 = tf.Variable(
tf.truncated_normal([num_hidden_nodes1, num_hidden_nodes2], stddev=np.sqrt(2.0 / num_hidden_nodes1)),name="v3")
biases2 = tf.Variable(tf.zeros([num_hidden_nodes2]),name="v4")
weights3 = tf.Variable(
tf.truncated_normal([num_hidden_nodes2, num_hidden_nodes3], stddev=np.sqrt(2.0 / num_hidden_nodes2)),name="v5")
biases3 = tf.Variable(tf.zeros([num_hidden_nodes3]),name="v6")
weights4 = tf.Variable(
tf.truncated_normal([num_hidden_nodes3, numLabels], stddev=np.sqrt(2.0 / num_hidden_nodes3)),name="v7")
biases4 = tf.Variable(tf.zeros([numLabels]),name="v8")
# Add ops to save and restore all the variables.
saver = tf.train.Saver()
# Training computation.
layer1_train = tf.nn.relu(tf.matmul(tf_train_dataset, weights1) + biases1)
drop1 = tf.nn.dropout(layer1_train, keep_prob)
layer2_train = tf.nn.relu(tf.matmul(drop1, weights2) + biases2)
drop2 = tf.nn.dropout(layer2_train, keep_prob)
layer3_train = tf.nn.relu(tf.matmul(drop2, weights3) + biases3)
drop3 = tf.nn.dropout(layer3_train, keep_prob)
logits = tf.matmul(drop3, weights4) + biases4
loss = tf.reduce_mean(
tf.nn.sigmoid_cross_entropy_with_logits(logits, tf_train_labels))
# Optimizer.
optimizer = tf.train.AdamOptimizer(learning_rate=0.1,
beta1=0.9, beta2=0.999,
epsilon=1e-08).minimize(loss)
# Predictions for the training, test data, and single mail.
train_prediction = tf.nn.sigmoid(logits)
layer1_test = tf.nn.relu(tf.matmul(tf_test_dataset, weights1) + biases1)
layer2_test = tf.nn.relu(tf.matmul(layer1_test, weights2) + biases2)
layer3_test = tf.nn.relu(tf.matmul(layer2_test, weights3) + biases3)
test_prediction = tf.nn.sigmoid(tf.matmul(layer3_test, weights4) + biases4)
layer1_mail = tf.nn.relu(tf.matmul(tf_mail, weights1) + biases1)
layer2_mail = tf.nn.relu(tf.matmul(layer1_mail, weights2) + biases2)
layer3_mail = tf.nn.relu(tf.matmul(layer2_mail, weights3) + biases3)
prediction_mail = tf.nn.sigmoid(tf.matmul(layer3_mail, weights4) + biases4)
# +
num_steps = 151
start = time.time()
with tf.Session(graph=graph) as session:
tf.initialize_all_variables().run()
print("Initialized")
for step in range(num_steps):
_, l, predictions = session.run(
[optimizer, loss, train_prediction])
acc = accuracy(predictions,trainY)
if (step % 10 == 0):
print("Loss at step %d: %f" % (step, l))
print("Accuracy: %.1f%%" % acc)
print("Test accuracy: %.1f%%" % accuracy(test_prediction.eval(), testY))
# Save the variables to disk.
save_path = saver.save(session, "./model.ckpt")
print("Model saved in file: %s" % save_path)
end = time.time()
duration = end - start
print("time consumed in training: %f seconds" % duration)
# -
import cPickle
from collections import Counter
# Load Bag of words that were in training data
BagOfWords = cPickle.load(open('BagOfWords.p', 'rb'))
features = set(BagOfWords)
featureDict = {feature:i for i,feature in enumerate(features)}
def get_feature_vector(email_text, featureDict):
'''
create feature/x vector from email text
row = email, cols = features
'''
featurevector = np.zeros(shape=(1,len(featureDict)),dtype=float)
tokens = email_text.split()
UniDist = Counter(tokens)
for key,value in UniDist.items():
if key in featureDict:
featurevector[0,featureDict[key]] = value
return featurevector
def regularize_vectors(featurevector):
'''
Input:
featurevector: vector, where single email is a row and features are columns
Returns:
featurevector: vector, updated by dividing each feature value by the total
number of features
'''
totalWords = np.sum(featurevector[0,:],axis=0)
featurevector[0,:] = np.multiply(featurevector[0,:],(1/(totalWords + 1e-5)))
return featurevector
# ## Enter your username and password
# If ur email is <EMAIL> , only enter a.b
#
# And before running the cell below, u need to enable IMAP from [HERE](https://support.google.com/mail/answer/7126229?hl=en&visit_id=1-636175756291815919-2042916920&rd=1)
#
# First time Gmail rejected the loggin , and sent mail to my inbox reviewing the block and asked me to allow loggin or not .
#
# And u just need to allow Less secure apps from [HERE](https://www.google.com/settings/security/lesssecureapps?rfn=27&rfnc=1&et=0&asae=2&anexp=ire-f3)
#
# 
import email, getpass, imaplib
user = raw_input("Enter your GMail username --> ")
pwd = <PASSWORD>("Enter your password --> ")
m = imaplib.IMAP4_SSL("imap.gmail.com")
m.login(user, pwd)
print('OK, <EMAIL> authenticated (Success)' % user)
def get_text(mail,i):
result, data = mail.uid('search', None, "ALL")
# search and return uids instead
latest_email_uid = data[0].split()[i] # unique ids wrt label selected
result, email_data = mail.uid('fetch', latest_email_uid, '(RFC822)')
# fetch the email body (RFC822) for the given ID
raw_email = email_data[0][1]
raw_email_string = raw_email.decode('utf-8')
# converts byte literal to string removing b''
email_message = email.message_from_string(raw_email_string)
# this will loop through all the available multiparts in mail
for part in email_message.walk():
if part.get_content_type() == "text/plain": # ignore attachments/html
body = part.get_payload(decode=True)
return str(body)
# +
with tf.Session(graph=graph) as session:
m.list() # Lists all labels in GMail
m.select('Inbox') # Connected to 'Inbox' or '[Gmail]/Spam'
for i in range(0,7,1):
email_text = get_text(m,i)
print(email_text[:9])
email_test = get_feature_vector(email_text,featureDict)
email_test = regularize_vectors(email_test)
# Restore variables from disk.
saver.restore(session, "./model.ckpt")
#print("Model restored.")
# Do some work with the model
feed_dict={tf_mail:email_test}
emailpred = session.run(prediction_mail,feed_dict=feed_dict)
#check on the first column of the single row
print(emailpred[0])
if emailpred[0][0] > emailpred[0][1] :
print("Spam")
else:
print("Not Spam")
# -
m.list() # Lists all labels in GMail
m.select('[Gmail]/Spam') # Connected to 'Inbox' or '[Gmail]/Spam'
for i in range(-1,-3,-1):
email_text = get_text(m,i)
#print(email_text)
email_test = get_feature_vector(email_text,featureDict)
email_test = regularize_vectors(email_test)
with tf.Session(graph=graph) as session:
# Restore variables from disk.
saver.restore(session, "./model.ckpt")
#print("Model restored.")
# Do some work with the model
feed_dict={tf_mail:email_test}
emailpred = session.run(prediction_mail,feed_dict=feed_dict)
#check on the first column of the single row
print(emailpred[0])
if emailpred[0][0] > emailpred[0][1] :
print("Spam")
else:
print("Not Spam")
# ## Logic used
# I used One-hot encoding which means that one column is One and other is Zero.
#
# prediction consists of two columns [Spam, Not Spam] , the prediction is normalized probability if the first column is greater than the second then it is 'Spam' and else is 'Not Spam'
testY[0]
| Spam detection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 3.2. Querying an encrypted model
# **protecting privacy and IP simultaneously**
#
# ### Set up
# +
# Import dependencies
import grid as gr
from grid import syft as sy
import torch as th
import skin_cancer_model_utils as scmu
# Ignore warnings
import warnings
warnings.filterwarnings('ignore')
# Hook torch
hook = sy.TorchHook(th)
me = hook.local_worker
me.is_client_worker = False
# Connect to nodes
grid_server = gr.WebsocketGridClient(hook, "http://localhost:3001", id="grid_server")
patient_server = gr.WebsocketGridClient(hook, "http://localhost:3000", id="patient_server")
hospital_server = gr.WebsocketGridClient(hook, "http://localhost:3002", id="hospital_server")
crypto_provider = gr.WebsocketGridClient(hook, "http://localhost:3003", id="crypto_provider")
grid_server.connect()
patient_server.connect()
hospital_server.connect()
crypto_provider.connect()
# Connect nodes to each other
gr.connect_all_nodes([grid_server, patient_server, hospital_server, crypto_provider])
# -
# ### Query model
# #### Get a copy of the encrypted model
encrypted_model = grid_server.download_model("skin-cancer-model-encrypted")
# **Encrypted data for running through the model**
data, target = scmu.get_data_sample()
x_sh = data.encrypt(patient_server, hospital_server, crypto_provider=crypto_provider)
# #### Run encrypted inference
# %%time
print(encrypted_model(x_sh).request_decryption(), target)
| examples/Serving and Querying models on Grid/3.2. Querying an encrypted model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: newpy36
# language: python
# name: newpy36
# ---
# # Spark EDA
# +
from pyspark.sql import SparkSession
spark = SparkSession.builder \
.appName('Spark EDA') \
.getOrCreate()
# +
data_path = '/home/lorenzo/spark-repo/1_spark_dataframes/data/california.csv'
df1 = spark.read.option('header', 'True') \
.option('inferSchema', 'True') \
.csv(data_path)
df1 = df1.withColumnRenamed('latitude', 'lat') \
.withColumnRenamed('longitude', 'lng') \
.withColumnRenamed('total_rooms', 'rooms') \
.withColumnRenamed('total_bedrooms', 'bedrooms') \
.withColumnRenamed('median_income', 'income') \
.withColumnRenamed('median_house_value', 'value') \
.withColumnRenamed('housing_median_age', 'age')
df1.createOrReplaceTempView('california')
# -
# ### Descriptive statistics
#
# **Describe() method:**
df1.select('rooms', 'income', 'age', 'value').describe().show()
# **Correlations:**
df1.stat.corr('rooms', 'value')
df1 = df1.withColumn('rooms_per_person', df1.rooms/df1.population)
df1.stat.corr('rooms_per_person', 'value')
# **Frequent values:**
df1.stat.freqItems(['age']).show()
df1.stat.freqItems(['age', 'value']).show()
# **Crosstable:**
df1.filter(df1.age <= 10).stat.crosstab('ocean_proximity', 'age').show()
# **Bucketing:**
spark.sql("SELECT count(*), FLOOR(age*10) as age_bucket \
FROM california GROUP BY age_bucket ORDER BY age_bucket").show()
spark.sql("SELECT count(*), round(mean(value), 2), FLOOR(age*10) as age_bucket \
FROM california GROUP BY age_bucket ORDER BY age_bucket").show()
# ### Time-series data
# +
data_path = '/home/lorenzo/Desktop/utilization.csv'
df2 = spark.read.option('header', 'False') \
.option('inferSchema', 'True') \
.csv(data_path)
# +
df2 = df2.withColumnRenamed("_c0", "event_datetime") \
.withColumnRenamed ("_c1", "server_id") \
.withColumnRenamed("_c2", "cpu_utilization") \
.withColumnRenamed("_c3", "free_memory") \
.withColumnRenamed("_c4", "session_count")
df2.createOrReplaceTempView('utilization')
# -
df2.show(5)
spark.sql("SELECT server_id, min(cpu_utilization), max(cpu_utilization) \
FROM utilization \
GROUP BY server_id").show(5)
# **Windowing:**
spark.sql("SELECT event_datetime, server_id, cpu_utilization, \
avg(cpu_utilization) OVER (PARTITION BY server_id) as avg_cpu_util \
FROM utilization").show(10)
spark.sql("SELECT event_datetime, server_id, cpu_utilization, \
avg(cpu_utilization) OVER (PARTITION BY server_id) as avg_cpu_util, \
cpu_utilization - avg(cpu_utilization) OVER (PARTITION BY server_id) as delta_cpu_util\
FROM utilization").show(10)
# **Moving window:**
spark.sql("SELECT event_datetime, server_id, cpu_utilization, \
avg(cpu_utilization) OVER (PARTITION BY server_id ORDER BY event_datetime \
ROWS BETWEEN 1 PRECEDING AND 1 FOLLOWING) as avg_cpu_util \
FROM utilization").show(10)
| 3_spark_eda/spark_eda.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/wesleybeckner/technology_fundamentals/blob/main/C4%20Machine%20Learning%20II/LABS_PROJECT/Tech_Fun_C4_L1_NN_Linearity.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="3rZ3ZWSkhxyS"
# # Technology Fundamentals Course 4, Lab 1: Practice with FFNNs
#
# **Instructor**: <NAME>
#
# **Contact**: <EMAIL>
#
# **Teaching Assitants**: <NAME>, <NAME>
#
# **Contact**: <EMAIL>, <EMAIL>
#
# <br>
#
# ---
#
# <br>
#
# In this lab we will compare the FFNN to the classification algorithms we created in Course 2.
#
# <br>
#
# ---
#
#
#
# + [markdown] id="t6pFVfpXtx9Q"
# # Data and Helper Functions
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="zTnsH0OQhoFn" outputId="186e2476-3496-4a27-feeb-782d43824a22"
import plotly.express as px
from sklearn.datasets import make_blobs, make_moons
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
X, y = make_blobs(random_state=42, centers=2, cluster_std=3)
X, y = make_moons(random_state=42, noise=.05, n_samples=1000)
px.scatter(x=X[:,0],y=X[:,1],color=y.astype(str))
# + id="9AXDQU8Ztrci"
def plot_boundaries(X, clf, ax=False):
plot_step = 0.02
x_min, x_max = X[:, 0].min() - 1, X[:, 0].max() + 1
y_min, y_max = X[:, 1].min() - 1, X[:, 1].max() + 1
xx, yy = np.meshgrid(np.arange(x_min, x_max, plot_step),
np.arange(y_min, y_max, plot_step))
Z = clf.predict(np.c_[xx.ravel(), yy.ravel()])
Z = Z.reshape(xx.shape)
if ax:
cs = ax.contourf(xx, yy, Z, cmap='viridis', alpha=0.2)
ax.scatter(X[:,0], X[:,1], c=y, cmap='viridis', edgecolor='grey', alpha=0.9)
return ax
else:
cs = plt.contourf(xx, yy, Z, cmap='viridis', alpha=0.2)
plt.scatter(X[:,0], X[:,1], c=y, cmap='viridis', edgecolor='grey', alpha=0.9)
# + [markdown] id="GA0l_pwYn9g1"
# # L1 Q1:
#
# Build and train a linear classification model using keras tf. Verify that the model is linear by either showing the weights or plotting the decision boundary (hint: you can use `plot_boundaries` above).
# + id="BR11MzzIj_ga" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="4533abaa-f035-4989-cf2a-ef8023daafa9"
# Code Cell for L1 Q1
from tensorflow import keras
from tensorflow.keras import layers
model = keras.Sequential([
#### YOUR CODE HERE ###
])
model.compile(
optimizer='adam',
loss='binary_crossentropy',
metrics=['binary_accuracy'],
)
history = model.fit(X,y,
batch_size=100,
epochs=500,
verbose=0)
model.summary()
results = pd.DataFrame(history.history)
display(results.tail())
y_pred = model.predict(X) > 0.5
px.scatter(x=X[:,0],y=X[:,1], color=y_pred.astype(str))
# + colab={"base_uri": "https://localhost:8080/", "height": 266} id="e2cZfcqKtiEz" outputId="7dfe98ea-b8aa-4c3a-f950-b583a888ca6e"
plot_boundaries(X, model)
# + [markdown] id="Sp832EkKi67C"
# # L1 Q2:
#
# Now add an activation function to your previous model. Does the model become non-linear?
#
# + id="Vb1NyIqZmqYg" colab={"base_uri": "https://localhost:8080/", "height": 729} outputId="a6a4fc3c-ec24-4e53-947e-053a95aac259"
# Code Cell for L1 Q2
model = keras.Sequential([
#### YOUR CODE HERE ###
])
model.compile(
optimizer='adam',
loss='binary_crossentropy',
metrics=['binary_accuracy'],
)
history = model.fit(X,y,
batch_size=100,
epochs=500,
verbose=0)
results = pd.DataFrame(history.history)
display(results.tail())
y_pred = model.predict(X) > 0.5
px.scatter(x=X[:,0],y=X[:,1],color=y_pred.astype(str))
# + colab={"base_uri": "https://localhost:8080/", "height": 266} id="Bc0rnsX9uOFA" outputId="187c91ad-2d0b-4e08-cfcb-1e376950de59"
plot_boundaries(X, model)
# + [markdown] id="e9MwkxIGlc0k"
# # L1 Q3:
#
# Continue to add complexity to your Q3 model until you get an accuracy above 99%
# + id="w-mykt4KhnA2" colab={"base_uri": "https://localhost:8080/", "height": 729} outputId="ff712966-9fe3-4b7f-886b-0ed822aef72b"
# Code Cell for L1 Q3
model = keras.Sequential([
#### YOUR CODE HERE ###
])
model.compile(
optimizer='adam',
loss='binary_crossentropy',
metrics=['binary_accuracy'],
)
history = model.fit(X,y,
batch_size=100,
epochs=100,
verbose=0)
results = pd.DataFrame(history.history)
display(results.tail())
y_pred = model.predict(X) > 0.5
px.scatter(x=X[:,0],y=X[:,1],color=y_pred.astype(str))
# + colab={"base_uri": "https://localhost:8080/", "height": 266} id="uHcqTSU6s0i5" outputId="658eec48-647e-49f2-f671-e5ea17c84797"
plot_boundaries(X, model)
# + id="S6lpJU18L2S0"
| C4 Machine Learning II/LABS_PROJECT/Tech_Fun_C4_L1_NN_Linearity.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Homework 2.2: The Quest For A Better Network
#
# In this assignment you will build a monster network to solve Tiny ImageNet image classification.
#
# This notebook is intended as a sequel to seminar 3, please give it a try if you haven't done so yet.
# (please read it at least diagonally)
#
# * The ultimate quest is to create a network that has as high __accuracy__ as you can push it.
# * There is a __mini-report__ at the end that you will have to fill in. We recommend reading it first and filling it while you iterate.
#
# ## Grading
# * starting at zero points
# * +20% for describing your iteration path in a report below.
# * +20% for building a network that gets above 20% accuracy
# * +10% for beating each of these milestones on __TEST__ dataset:
# * 25% (50% points)
# * 30% (60% points)
# * 32.5% (70% points)
# * 35% (80% points)
# * 37.5% (90% points)
# * 40% (full points)
#
# ## Restrictions
# * Please do NOT use pre-trained networks for this assignment until you reach 40%.
# * In other words, base milestones must be beaten without pre-trained nets (and such net must be present in the anytask atttachments). After that, you can use whatever you want.
# * you __can't__ do anything with validation data apart from running the evaluation procedure. Please, split train images on train and validation parts
#
# ## Tips on what can be done:
#
#
# * __Network size__
# * MOAR neurons,
# * MOAR layers, ([torch.nn docs](http://pytorch.org/docs/master/nn.html))
#
# * Nonlinearities in the hidden layers
# * tanh, relu, leaky relu, etc
# * Larger networks may take more epochs to train, so don't discard your net just because it could didn't beat the baseline in 5 epochs.
#
# * Ph'nglui mglw'nafh Cthulhu R'lyeh wgah'nagl fhtagn!
#
#
# ### The main rule of prototyping: one change at a time
# * By now you probably have several ideas on what to change. By all means, try them out! But there's a catch: __never test several new things at once__.
#
#
# ### Optimization
# * Training for 100 epochs regardless of anything is probably a bad idea.
# * Some networks converge over 5 epochs, others - over 500.
# * Way to go: stop when validation score is 10 iterations past maximum
# * You should certainly use adaptive optimizers
# * rmsprop, nesterov_momentum, adam, adagrad and so on.
# * Converge faster and sometimes reach better optima
# * It might make sense to tweak learning rate/momentum, other learning parameters, batch size and number of epochs
# * __BatchNormalization__ (nn.BatchNorm2d) for the win!
# * Sometimes more batch normalization is better.
# * __Regularize__ to prevent overfitting
# * Add some L2 weight norm to the loss function, PyTorch will do the rest
# * Can be done manually or like [this](https://discuss.pytorch.org/t/simple-l2-regularization/139/2).
# * Dropout (`nn.Dropout`) - to prevent overfitting
# * Don't overdo it. Check if it actually makes your network better
#
# ### Convolution architectures
# * This task __can__ be solved by a sequence of convolutions and poolings with batch_norm and ReLU seasoning, but you shouldn't necessarily stop there.
# * [Inception family](https://hacktilldawn.com/2016/09/25/inception-modules-explained-and-implemented/), [ResNet family](https://towardsdatascience.com/an-overview-of-resnet-and-its-variants-5281e2f56035?gi=9018057983ca), [Densely-connected convolutions (exotic)](https://arxiv.org/abs/1608.06993), [Capsule networks (exotic)](https://arxiv.org/abs/1710.09829)
# * Please do try a few simple architectures before you go for resnet-152.
# * Warning! Training convolutional networks can take long without GPU. That's okay.
# * If you are CPU-only, we still recomment that you try a simple convolutional architecture
# * a perfect option is if you can set it up to run at nighttime and check it up at the morning.
# * Make reasonable layer size estimates. A 128-neuron first convolution is likely an overkill.
# * __To reduce computation__ time by a factor in exchange for some accuracy drop, try using __stride__ parameter. A stride=2 convolution should take roughly 1/4 of the default (stride=1) one.
#
#
# ### Data augmemntation
# * getting 5x as large dataset for free is a great
# * Zoom-in+slice = move
# * Rotate+zoom(to remove black stripes)
# * Add Noize (gaussian or bernoulli)
# * Simple way to do that (if you have PIL/Image):
# * ```from scipy.misc import imrotate,imresize```
# * and a few slicing
# * Other cool libraries: cv2, skimake, PIL/Pillow
# * A more advanced way is to use torchvision transforms:
# ```
# transform_train = transforms.Compose([
# transforms.RandomCrop(32, padding=4),
# transforms.RandomHorizontalFlip(),
# transforms.ToTensor(),
# transforms.Normalize((0.4914, 0.4822, 0.4465), (0.2023, 0.1994, 0.2010)),
# ])
# trainset = torchvision.datasets.ImageFolder(root=path_to_tiny_imagenet, train=True, download=True, transform=transform_train)
# trainloader = torch.utils.data.DataLoader(trainset, batch_size=128, shuffle=True, num_workers=2)
#
# ```
# * Or use this tool from Keras (requires theano/tensorflow): [tutorial](https://blog.keras.io/building-powerful-image-classification-models-using-very-little-data.html), [docs](https://keras.io/preprocessing/image/)
# * Stay realistic. There's usually no point in flipping dogs upside down as that is not the way you usually see them.
#
#
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
from tiny_img import download_tinyImg200
data_path = '.'
download_tinyImg200(data_path)
dataset = torchvision.datasets.ImageFolder('tiny-imagenet-200/train', transform=transforms.ToTensor())
train_dataset, val_dataset = torch.utils.data.random_split(dataset, [80000, 20000])
# +
# feel free to copypaste code from seminar03 as a basic template for training
# -
# When everything is done, please calculate accuracy on `tiny-imagenet-200/val`
test_accuracy = .... # YOUR CODE
# +
print("Final results:")
print(" test accuracy:\t\t{:.2f} %".format(
test_accuracy * 100))
if test_accuracy * 100 > 40:
print("Achievement unlocked: 110lvl Warlock!")
elif test_accuracy * 100 > 35:
print("Achievement unlocked: 80lvl Warlock!")
elif test_accuracy * 100 > 30:
print("Achievement unlocked: 70lvl Warlock!")
elif test_accuracy * 100 > 25:
print("Achievement unlocked: 60lvl Warlock!")
else:
print("We need more magic! Follow instructons below")
# -
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
#
# # Report
#
# All creative approaches are highly welcome, but at the very least it would be great to mention
# * the idea;
# * brief history of tweaks and improvements;
# * what is the final architecture and why?
# * what is the training method and, again, why?
# * Any regularizations and other techniques applied and their effects;
#
#
# There is no need to write strict mathematical proofs (unless you want to).
# * "I tried this, this and this, and the second one turned out to be better. And i just didn't like the name of that one" - OK, but can be better
# * "I have analized these and these articles|sources|blog posts, tried that and that to adapt them to my problem and the conclusions are such and such" - the ideal one
# * "I took that code that demo without understanding it, but i'll never confess that and instead i'll make up some pseudoscientific explaination" - __not_ok__
# ### Hi, my name is `___ ___`, and here's my story
#
# A long time ago in a galaxy far far away, when it was still more than an hour before the deadline, i got an idea:
#
# ##### I gonna build a neural network, that
# * brief text on what was
# * the original idea
# * and why it was so
#
# How could i be so naive?!
#
# ##### One day, with no signs of warning,
# This thing has finally converged and
# * Some explaination about what were the results,
# * what worked and what didn't
# * most importantly - what next steps were taken, if any
# * and what were their respective outcomes
#
# ##### Finally, after __ iterations, __ mugs of [tea/coffee]
# * what was the final architecture
# * as well as training method and tricks
#
# That, having wasted ____ [minutes, hours or days] of my life training, got
#
# * accuracy on training: __
# * accuracy on validation: __
# * accuracy on test: __
#
#
# [an optional afterword and mortal curses on assignment authors]
| homework02/homework_part2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Environment (conda_tensorflow_p36)
# language: python
# name: conda_tensorflow_p36
# ---
# On this hands-on lab we will perform few activities related to Sequence Models - in special Recurrent Neural Networks applied to sentiment analysis and time series forecasting.
#
# To perform those activities it is important to address some requirements beforehand:
#
# 1) deploy one AWS EC2 instance (P2.8x type) to be used as sandbox (it could be destroyed after the lab execution)
#
# 2) After logging in the instance, run 'source activate tensorflow_p36'
#
# 3) Create a directory as 'mkdir -p /models/ai-conference' and enter on it 'cd /models/ai-conference'
#
# 4) Clone the github repository containing the labs 'git clone github link'
#
# This notebook includes the following activities:
#
# - building a first sample RNN (LSTM) on NLP
# - train the neural network using the IMDB reviews dataset and evaluate its performance
# - report the performance metrics for that model, including precision, recall, f1score and support
# - performing transfer learning to speed up model creation process
# - building a second sample RNN (LSTM) network on Time Series forecasting
#
# ## Part I - Sequence Models basics
# +
# validate that the required python modules are installed before starting
# !conda install -y seaborn Pillow scikit-learn
# +
# importing required modules
import os
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from IPython.display import Image
from operator import itemgetter
from keras import models, regularizers, layers, optimizers, losses, metrics
from keras.models import Sequential
from keras.layers import Dense
from keras.utils import np_utils, to_categorical
from keras.datasets import imdb
from keras.utils.training_utils import multi_gpu_model
from sklearn.metrics import confusion_matrix, classification_report
# -
# ### "Recurrent means the output at the current time step becomes the input to the next time step. At each element of the sequence, the model considers not just the current input, but what it remembers about the preceding elements."
#
# 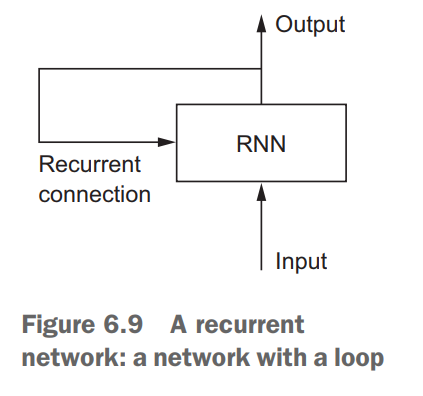
# +
# For reproducibility
np.random.seed(1000)
# model configuration -- number of GPUs and training option (Yes or No)
n_gpus = 8 # knob to make the model parallel or not
train_model = False # knob to decide if the model will be trained or imported
# +
# loading IMDB dataset
(train_data, train_labels), (test_data, test_labels) = imdb.load_data(num_words=10000)
# +
# taking a look on the dataset info
print("_"*50)
print("\ntrain_data ", train_data.shape)
print("train_labels ", train_labels.shape)
print("_"*50)
print("\ntest_data ", test_data.shape)
print("test_labels ", test_labels.shape)
print("_"*50)
print("\nMaximum value of a word index ")
print(max([max(sequence) for sequence in train_data]))
print("\nMaximum length num words of review in train ")
print(max([len(sequence) for sequence in train_data]))
print("_"*50)
# checking a sample from the dataset
word_index = imdb.get_word_index()
reverse_word_index = dict([(value, key) for (key, value) in word_index.items()])
decoded_review = ' '.join([reverse_word_index.get(i - 3, '?') for i in train_data[123]])
print('decoded text:\n\n', decoded_review)
# -
# 
# +
# function to vectorize the dataset information
def vectorize_sequences(sequences, dimension=10000):
results = np.zeros((len(sequences), dimension))
for i, sequence in enumerate(sequences):
results[i, sequence] = 1.
return results
# vectorizing the datasets
X_train = vectorize_sequences(train_data)
X_test = vectorize_sequences(test_data)
print("x_train ", X_train.shape)
print("x_test ", X_test.shape)
# vectorizing the labels
y_train = np.asarray(train_labels).astype('float32')
y_test = np.asarray(test_labels).astype('float32')
print("y_train ", y_train.shape)
print("y_test ", y_test.shape)
# +
# creating a validation set
X_val = X_train[:10000]
X_train = X_train[10000:]
y_val = y_train[:10000]
y_train = y_train[10000:]
print("x_val ", X_val.shape)
print("X_train ", X_train.shape)
print("y_val ", y_val.shape)
print("y_train ", y_train.shape)
# -
# 
# +
# creating the RNN model
if train_model is True:
model = models.Sequential()
model.add(layers.Dense(16, kernel_regularizer=regularizers.l1(0.001), activation='relu', input_shape=(10000,)))
model.add(layers.Dropout(0.5))
model.add(layers.Dense(16, kernel_regularizer=regularizers.l1(0.001),activation='relu'))
model.add(layers.Dropout(0.5))
model.add(layers.Dense(1, activation='sigmoid'))
# making the model aware of multiple GPUs
if n_gpus > 1:
final_model = multi_gpu_model(model, gpus=8)
else:
final_model = model
# compile the model
final_model.compile(optimizer='rmsprop',
loss='binary_crossentropy',
metrics=['accuracy'])
# summarize the model
final_model.summary()
# +
# training the model
if train_model is True:
n_epochs = 20
batch_size = 512
history = final_model.fit(X_train,
y_train,
epochs=n_epochs,
batch_size=batch_size,
validation_data=(X_val, y_val))
# +
# save the model details
if train_model is True:
# save the model
model.save('rnn_model.h5')
# +
# summarize history for accuracy
if train_model is True:
plt.plot(history.history['acc'])
plt.plot(history.history['val_acc'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'validation'], loc='upper left')
plt.show()
# +
# summarize history for loss
if train_model is True:
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'validation'], loc='upper left')
plt.show()
# +
# evaluating the model
if train_model is True:
results = final_model.evaluate(X_test, y_test)
print("_"*50)
print("Test Loss and Accuracy")
print("results ", results)
history_dict = history.history
# +
# evaluating the model accuracy
if train_model is not True:
from keras.models import load_model
final_model = load_model('rnn_model.h5')
final_model.compile(optimizer='adam',
loss='categorical_crossentropy',
metrics=['accuracy'])
predictions = final_model.predict(X_test)
predictions = (predictions > 0.5)
cm = confusion_matrix(y_test, predictions)
plt.imshow(cm, cmap=plt.cm.Blues)
classNames = ['Negative','Positive']
plt.title('IMDB reviews sentiment analysis')
plt.xlabel("Predicted labels")
plt.ylabel("True labels")
tick_marks = np.arange(len(classNames))
plt.xticks(tick_marks, classNames, rotation=45)
plt.yticks(tick_marks, classNames)
s = [['TN','FP'], ['FN', 'TP']]
for i in range(2):
for j in range(2):
plt.text(j,i, str(s[i][j])+" = "+str(cm[i][j]))
plt.colorbar()
plt.show()
# -
print(classification_report(predictions, y_test))
# ## Part II - Time Series Forecasting
#
# To avoid memory and/or cpu usage issues, it is important to reset the Jupyter Notebook kernel.
#
# This task can be performed as:
#
# - go to the Jupyter notebook menu (up there)
# - click on 'Kernel'
# - click on 'Restart'
# - wait for the kernel to restart
#
# Once the restart procedure is finished, go ahead on the next steps.
# +
# importing modules
from datetime import datetime
from math import sqrt
import numpy as np
from numpy import concatenate
from matplotlib import pyplot as plt
import pandas as pd
from pandas import read_csv, DataFrame, concat
from sklearn.preprocessing import MinMaxScaler, LabelEncoder
from sklearn.metrics import mean_squared_error
from keras.models import Sequential
from keras.layers import Dense, LSTM, Dropout
from keras.utils.training_utils import multi_gpu_model
# +
# model configuration -- number of GPUs and training option (Yes or No)
n_gpus = 1 # knob to make the model parallel or not
# +
# load and process data
def parse(x):
return datetime.strptime(x, '%Y %m %d %H')
dataset = read_csv('pollution.csv',
parse_dates = [['year', 'month', 'day', 'hour']],
index_col=0,
date_parser=parse)
dataset.drop('No', axis=1, inplace=True)
dataset.columns = ['pollution', 'dew', 'temp', 'press',
'wnd_dir', 'wnd_spd', 'snow', 'rain']
dataset.index.name = 'date'
dataset['pollution'].fillna(0, inplace=True)
dataset = dataset[24:]
print('-'*100)
print(dataset.head(5))
print('-'*100)
dataset.to_csv('pollution_parsed.csv')
# +
# visualizing the dataset
dataset = read_csv('pollution_parsed.csv', header=0, index_col=0)
values = dataset.values
groups = [0, 1, 2, 3, 5, 6, 7]
i = 1
plt.figure()
for group in groups:
plt.subplot(len(groups), 1, i)
plt.plot(values[:, group],'k')
plt.title(dataset.columns[group], y=0.5, loc='right')
i += 1
plt.show()
# +
# loading the dataset
def organize_series(data, n_in=1, n_out=1, dropnan=True):
n_vars = 1 if type(data) is list else data.shape[1]
df = pd.DataFrame(data)
cols, names = list(), list()
for i in range(n_in, 0, -1):
cols.append(df.shift(i))
names += [('var%d(t-%d)' % (j+1, i)) for j in range(n_vars)]
for i in range(0, n_out):
cols.append(df.shift(-i))
if i == 0:
names += [('var%d(t)' % (j+1)) for j in range(n_vars)]
else:
names += [('var%d(t+%d)' % (j+1, i)) for j in range(n_vars)]
agg = pd.concat(cols, axis=1)
agg.columns = names
if dropnan:
agg.dropna(inplace=True)
return agg
# processing the dataset
dataset = read_csv('pollution_parsed.csv', header=0, index_col=0)
values = dataset.values
encoder = LabelEncoder()
values[:,4] = encoder.fit_transform(values[:,4])
values = values.astype('float32')
scaler = MinMaxScaler(feature_range=(0, 1))
scaled = scaler.fit_transform(values)
reframed = organize_series(scaled, 1, 1)
# +
# split data into training and testing
values = reframed.values
n_train_hours = 365 * 24
test = values[:n_train_hours, :]
train = values[n_train_hours:, :]
# split into input and outputs
X_train, y_train = train[:, :-1], train[:, -1]
X_test, y_test = test[:, :-1], test[:, -1]
X_train = X_train.reshape((X_train.shape[0], 1, X_train.shape[1]))
X_test = X_test.reshape((X_test.shape[0], 1, X_test.shape[1]))
print(" Training data shape X, y => ",X_train.shape, y_train.shape," Testing data shape X, y => ", X_test.shape, y_test.shape)
# +
# defining the RNN/LSTM model
model = Sequential()
model.add(LSTM(50, input_shape=(X_train.shape[1], X_train.shape[2])))
model.add(Dropout(0.3))
model.add(Dense(1,kernel_initializer='normal', activation='sigmoid'))
# make the model aware of multi GPU
if n_gpus > 1:
final_model = multi_gpu_model(model, gpus=8)
else:
final_model = model
# compile the model
final_model.compile(loss='mae', optimizer='adam')
# summarize the model
final_model.summary()
# +
# training the RNN/LSTM model
epochs = 5
batch_size = 72
history = final_model.fit(X_train,
y_train,
epochs=epochs,
batch_size=batch_size,
validation_data=(X_test, y_test),
shuffle=False)
# +
# visualizing the loss during training
plt.plot(history.history['loss'], 'b', label='Training')
plt.plot(history.history['val_loss'], 'r',label='Validation')
plt.title("Train and Test Loss for the LSTM")
plt.legend()
plt.show()
# -
# ## Cleaning things up
#
# Not much actions must be taken to clean the environment used on this lab.
#
# As a new EC2 instance was created for this purpose, simply terminate the instance.
| Sequence/AI ML LatAm Conference 2019 - Sequence Models HandsOn.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.3.1
# language: julia
# name: julia-1.3
# ---
# ## Reading data from ROOT using UpROOT
#
# _For flat data structures, it's very simple to read in data using uproot,
# especially when that data can fit in memory. In this example we read a ROOT file
# in and store the data in a DataFrame. The data is then split 80/20, with the first
# 80 used to form the predicted hypothesis/spectrum, and the last 20 used for validation._
push!(LOAD_PATH, "../src/")
using Batman
using DataFrames
# import Random: rand
using Distributions
# using StatsPlots; pyplot()
using PyPlot
plt.style.use("bat.mplstyle")
# ### Readiness Checklist
# _Once this checklist is completed, it can be removed and this module is complete_
# - [x] Read in an arbitrary root file (produced externally)
# - [x] Produce multiple 1D PDFs
# - [ ] Produce a 2D PDF
# - [ ] Produce a ... 4D PDF?
# - [x] Mock dataset to fit to a 1D PDF
# - [ ] Fit to the 2D distribution (extended likelihood)
# - [ ] Produce uncertainties
# - [x] Bias
# - [x] Pull
# MODEL MC
signalMC = DataStructures.rootreader("assets/signal.root", "bat")
bkgMC = DataStructures.rootreader("assets/background.root", "bat")
bins = collect(-20:0.1:20);
# +
import Interpolations
# Build Model: First take PDF from histogram to get generic function
sig_itp = HistogramPDF(signalMC, :energy; bins=bins, extrapolate=Interpolations.Flat())
sig_pos = HistogramPDF(signalMC, :position; bins=bins)
bkg_itp = HistogramPDF(bkgMC, :energy, bins=bins)
bkg_pos = HistogramPDF(bkgMC, :position; bins=bins)
model = SpectralMonofit()
s = add_parameter!(model, "signal", 20.0)
b = add_parameter!(model, "background", 500.0; σ=1.0 )
add_observable!(model, :energy, -20.0, 20.0)
add_observable!(model, :position, -20.0, 20.0)
add_dataset(:tb1, DataFrame(energy=Float64[], position=Float64[]))
# Each dataset should have a LogPDF of its own
#add_dataset(:tb1, data)
#f1 = constructPDF!(model, s, [sig_itp], [:energy], :tb1)
#f2 = constructPDF!(model, b, [bkg_itp], [:energy], :tb1)
#f1 = constructPDF!(model, s, [sig_pos], [:position], :tb1)
#f2 = constructPDF!(model, b, [bkg_pos], [:position], :tb1)
f1 = constructPDF!(model, s, [sig_itp,sig_pos], [:energy, :position], :tb1)
f2 = constructPDF!(model, b, [bkg_itp,bkg_pos], [:energy, :position], :tb1)
#f1 = constructPDF!(model, s, [sig_itp,sig_itp], [:energy, :energy], :tb1)
#f2 = constructPDF!(model, b, [bkg_itp,bkg_itp], [:energy, :energy], :tb1)
@show f1
#combinePDFs!(model, [f1, f2], :tb1)
combinePDFs!(model, [f1, f2], :tb1)
generate_mock_dataset(model)
options = Dict(
"ftol_abs"=>0,
"ftol_rel"=>1e-6,
"initial_step"=>[10.0, 10.0]
)
results = minimize!(model; options=options);
compute_profiled_uncertainties!(results; σ=1, init_step=[1.0, 1.0], step=0.1)
pretty_results(results)
# -
# ### 1D component fit: Signal v Background
# +
## Plot spectra
sx = collect(-10:0.01:10)
p = getparam(model, "signal")
signal_y = getparam(model, "signal").fit * sig_itp(sx) * (bins[2] - bins[1])
bkg_y = getparam(model, "background").fit * bkg_itp(sx) * (bins[2] - bins[1])
fs_y = 1000 * sig_itp(sx) * (bins[2]-bins[1])
fb_y = 350 * bkg_itp(sx) * (bins[2]-bins[1])
#plt.plot(sx, fs_y, label="fake signal", color="blue" )
#plt.plot(sx, fb_y, label="fake signal", color="red")
#plt.plot(sx, fs_y+fb_y, label="fake signal", color="black")
plt.plot(sx, signal_y+bkg_y, label="Total", color="black")
plt.plot(sx, signal_y, label="signal")
plt.plot(sx, bkg_y, label="bkg")
plt.hist(Batman.tb1.energy, bins=bins, label="Data")
plt.legend()
# -
## Plot spectra
sx = collect(-10:0.01:10)
p = getparam(model, "signal")
signal_y = getparam(model, "signal").fit * sig_pos(sx) * (bins[2] - bins[1])
bkg_y = getparam(model, "background").fit * bkg_pos(sx) * (bins[2] - bins[1])
plt.plot(sx, signal_y+bkg_y, label="Total", color="black")
plt.plot(sx, signal_y, label="signal")
plt.plot(sx, bkg_y, label="bkg")
plt.hist(Batman.tb1.position, bins=bins, label="Data")
plt.legend()
# +
hs = x -> x >= 0 ? 1 : 0
#profile!("Signal", results; prior=nothing)
#uncertainty!("Signal", results )
interval_plot(results, "signal")
plt.savefig("profile.svg")
plt.show()
# -
correlation_plots(results)
plt.show()
correlation_plots2(results)
# +
## Bias/Pull Testing
bias_vector = []
pull_vector = []
trials = collect(1:1000)
errors = 0
for t in trials
try
#mock_dataset()
generate_mock_dataset(model)
if size(model.datasets[1], 1) < 1
continue
end
results = minimize!(model; options=options)
profile!("signal", results; init_step=[1.0, 1.0], step=0.1)
uncertainty!("signal", results;σ=1)
sig_stats = getparam(model, "signal")
bias = sig_stats.fit - sig_stats.init
if bias >= 0
pull = bias / abs(sig_stats.fit - sig_stats.low)
else
pull = bias / abs(sig_stats.fit - sig_stats.high)
end
push!(bias_vector, bias)
push!(pull_vector, pull)
catch e
errors += 1
end
print(t,"\r")
end
println("Failure rate: ", errors/maximum(trials))
# -
@show mean = sum(bias_vector) / length(bias_vector)
plt.hist(bias_vector, bins=100);
# +
@show mean = sum(pull_vector)/length(pull_vector)
@show dev = sqrt(sum((pull_vector.-mean).^2)/length(pull_vector))
plt.hist(pull_vector, bins=collect(-3:0.1:3))
println("Pull distribution: ", mean, " +- ", dev);
# -
| examples/4_SpectralMonofit.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Resolución del problema de Laplace 1D con condiciones Dirichlet
# Calcular $u:[a,b] \to R$ solución de
# $$
# \begin{align}
# \label{pb1d}
# -\frac{d^2 u(x) }{dx^2} = f(x) \mbox{ en } [a,b], \\
# u(a)=u_a, \ u(b)=u_b,
# \end{align}
# $$
#
# ## Sistema de ecuaciones MDF 1D
from numpy import *
# Datos:
a, b = 0, 1 #Extremos del intervalo
n = 10 # Números de puntos interiores en la partición, x_1,..., x_n
h = 1./(n+1) # Tamaño de la partición
ua, ub = 0, 1 # Datos de contorno
f = lambda x: pi**2/4 * sin(x*pi/2)
# ### Matriz
# Caso n=3
A=array(
[
[2,-1,0], # Fila1
[-1,2,-1], # Fila 2
[0,-1,2] # Fila 3
]
)
print("n=3:\n", A)
# Caso general:
A_h = (1./h**2) * (
2*diag( ones(n) )
- diag( ones(n-1), +1 )
- diag( ones(n-1), -1)
)
print("Caso general:\n", A_h)
# ### Segundo miembro
f_h = []
x = linspace(0, 1, num=n) # x_0, ..., x_{n-1}
#for i in range(n): # i 0,...n-1
# f_h.append( f(x[i]) )
f_h = f(x) # f_h es el array resultante de aplicar f a cada elmento del array x
f_h[0] += ua/h**2
f_h[-1] += ub/h**2
f_h
# ### Resolver sistema
# +
from numpy.linalg import solve
u_h = solve(A_h, f_h)
from matplotlib.pylab import plot, show, grid, legend
plot(x, u_h, label="Solución aproximada", linewidth=3, color="green")
grid()
legend()
show()
# -
# ## Papel sucio....
# +
# Trabajando...
size = 24
print("El tamaño es %i" %size)
# Mucho más abajo:
from numpy import *
x=array([1,2,3])
size(x) # No me he dado cuenta de que el import ha pisado la variable size
# De nuevo más abajo
print("El tamaño es %i", size)
# -
import numpy as np
x = np.array([1,2,3])
y = np.sin(x)
y
2*identity(3)
ones(3)
diag([1,3,88],-2)
x = linspace(0,1, n) # x_0, ..., x_{n-1}
x[9]
lista = [1,2,3,4,5,6]
lista[:5:2]
# %pylab
plo
| diferencias-finitas/Laplace MDF 1D.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:py27]
# language: python
# name: conda-env-py27-py
# ---
# #Functions
def double(x):
"""optional doc string
where you can specifiy what this does
ex: this function multiplies input by 2"""
return x*2
| .ipynb_checkpoints/chp02 crash course-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Analysis with 2-by-2 model Hamiltonian
# Based on <NAME> *et al.*, *J. Phys. Chem. Lett.* **12**, 1202-1206 (2021)
#
# * Take 70% of a crossing as a data set
# * Select all sets that
# * include the mid point
# * consist of 8 or more connected points
# * Print the average and do some analysis
# +
import sys
import numpy as np
import pandas as pd
import matplotlib
import matplotlib.pyplot as plt
sys.path.append('../../Python_libs')
import stabtools as st
matplotlib.use('Qt5Agg')
# %matplotlib qt5
# -
# ### Read-in stabilization data
Angs2Bohr=1.8897259886
au2eV=27.211386027
au2cm=219474.63068
#
# files in the current directory do not need the path name
#
dfin=pd.read_csv("DVR_stab_plot.csv")
#dfin=pd.read_csv("GTO_DZ_stab_plot.csv")
dfin.head()
# +
z_col=dfin.columns[1]
all_zs=dfin[z_col].values
zmin, zmax = all_zs[0], all_zs[-1]
if zmax < zmin:
zmin, zmax = zmax, zmin
all_zs=(all_zs-zmin)/(zmax-zmin)
E_labels=dfin.columns[2:]
plt.cla()
plt.xlabel('Z')
plt.ylabel('E')
for E in E_labels:
plt.plot(all_zs, dfin[E].values, '-')
plt.ylim(0,7)
plt.show()
# -
# Set the following parameters after inspecting the stabilization plot:
# * lroot: the lower root; the 2nd root is lroot+1
# * curvature_cutoff: the crossing is defined by the minimum and maximum curvature positions; the selection range is determined by the drop off of the abs(curvature)
# +
lroot = 3
curvature_cutoff=0.3
E_lw=dfin[E_labels[lroot]].values
E_up=dfin[E_labels[lroot+1]].values
success, cross_center, zs, E1s, E2s = st.crossing(all_zs, E_lw, E_up, curvature_cutoff)
if success:
print(f'Center of the crossing at z={cross_center:.8f}')
npts = len(zs)
print(f'N = {npts} points on each curve')
plt.cla()
plt.plot(all_zs, E_lw, all_zs, E_up)
plt.plot(zs, E1s, 'o', zs, E2s, 'o')
plt.ylabel('roots '+str(lroot)+' and '+str(lroot+1))
plt.show()
else:
print('No crossing found.')
zcs = (all_zs[zs[0]], all_zs[zs[1]])
print('min/max curvature at z=%f and z=%f' % zcs)
plt.cla()
plt.plot(all_zs, E1s, all_zs, E2s)
plt.ylabel('curvature '+str(lroot)+' and '+str(lroot+1))
plt.show()
# +
N=len(zs)
j_mid=np.argmin(abs(zs-cross_center))
min_data=8 #
ils=[]
irs=[]
Ers=[]
Eis=[]
chi2s=[]
for ilft in range(j_mid-1):
for irht in range(N,j_mid+1,-1):
if irht - ilft > min_data:
zsel, E1sel, E2sel = zs[ilft:irht], E1s[ilft:irht], E2s[ilft:irht]
zreal, zimag, Er, Ei, chi2 = st.tbt_ana_lsq(zsel, E1sel, E2sel, cross_center)
ils.append(ilft)
irs.append(N-irht)
Ers.append(Er)
Eis.append(Ei)
chi2s.append(chi2)
print(f'{ilft:4d} {N-irht:4d} {Er:8f} {Ei:8f} {chi2:.3e}')
# -
dic={'left':ils, 'right':irs, 'Er':Ers, 'Ei':Eis, 'chi2':chi2s}
df=pd.DataFrame(dic)
df.plot.scatter(x='Er', y='Ei')
df.describe()
xs=df.Er.values
ys=df.Ei.values
#
# long vs short datasets
#
j_left=df.left.values*1.0
j_right=df.right.values*1.0
sum_pts=j_left + j_right
#sum_pts=left + right
plt.cla()
plt.scatter(xs, ys, marker='.', c=sum_pts, cmap='viridis')
plt.colorbar()
plt.show()
#
# symmetric vs unsymmetric datasets
#
diff_pts=abs(j_left - j_right)
plt.cla()
plt.scatter(xs, ys, marker='.', c=diff_pts, cmap='viridis')
plt.colorbar()
plt.show()
# +
# both
fig, axs = plt.subplots(1, 2, sharex=True, sharey=True)
fig.set_figwidth(6.4)
fig.set_figheight(3.2)
#axs[0].set_xticks([3.15, 3.18])
axs[0].scatter(xs, ys, marker='.', c=sum_pts, cmap='viridis')
axs[1].scatter(xs, ys, marker='.', c=diff_pts, cmap='viridis')
axs[0].plot(3.17296,-0.160848,'k+',markersize=10)
axs[1].plot(3.17296,-0.160848,'k+',markersize=10)
axs[0].set_xlabel("$E_r$ [eV]", fontsize=12)
axs[1].set_xlabel("$E_r$ [eV]", fontsize=12)
axs[0].set_ylabel("$E_i$ [eV]", fontsize=12)
plt.tight_layout()
plt.show()
# -
| notebooks/Stab_3D/2x2_scatter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="9sQDpSAlqjgp"
# %matplotlib inline
# + [markdown] id="vWvPmOS-xGHQ"
# code for mounting the google drive if dataset from google drive is used.
# + colab={"base_uri": "https://localhost:8080/"} id="49et9uctojCy" outputId="345a9f5a-2e3c-469e-fb01-e5125b663775"
from google.colab import drive
drive.mount('/content/drive')
# + [markdown] id="vkNBu01exO4W"
# Code for archiving the dataset.
# + colab={"base_uri": "https://localhost:8080/"} id="FJ1RY2AGokv0" outputId="a96064b1-1cc2-4095-db1b-a12dc01bc5a9"
# !pip install pyunpack
# !pip install patool
from pyunpack import Archive
Archive('/content/drive/MyDrive/arc.zip').extractall('/content/sample_data')
# + colab={"base_uri": "https://localhost:8080/"} id="LlrGHhq_rCCb" outputId="ea99f82b-048b-4075-c4d5-75f6bb6c269c"
# !pip3 install --user Flask
# + id="yskEGxBlttRg"
# #!pip3 install --user tensorflow
# #!pip3 install --user keras
# #!pip3 install --user pandas
# + [markdown] id="Y7OSe7PAxwaZ"
# Importations
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" id="KTkYPxellFWx"
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import os
# + _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a" id="b8q_mFYplFWz"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
import seaborn as sns
import cv2
import os
from tqdm import tqdm
from sklearn.metrics import confusion_matrix
from sklearn.model_selection import train_test_split
from keras.utils.np_utils import to_categorical
from keras.models import Model,Sequential, Input, load_model
from keras.layers import Dense, Dropout, Flatten, Conv2D, MaxPool2D, BatchNormalization, AveragePooling2D, GlobalAveragePooling2D
from keras.optimizers import Adam
from keras.preprocessing.image import ImageDataGenerator
from keras.callbacks import ModelCheckpoint, ReduceLROnPlateau
from keras.applications import DenseNet121
# + [markdown] id="ESjhVF5LlWYC"
# Loading the dataset
# + id="ikMgCltmlFW0"
disease_types = ['Pepper__bell___Bacterial_spot','Pepper__bell___healthy','Potato___Early_blight','Potato___Late_blight','Potato___healthy','Tomato_Bacterial_spot','Tomato_Early_blight','Tomato_Late_blight','Tomato_Leaf_Mold','Tomato_Septoria_leaf_spot','Tomato_Spider_mites_Two_spotted_spider_mite','Tomato__Target_Spot','Tomato__Tomato_YellowLeaf__Curl_Virus','Tomato__Tomato_mosaic_virus','Tomato_healthy']
data_dir = '/content/sample_data/PlantVillage'
train_dir = os.path.join(data_dir)
# + [markdown] id="xDA0WqMDyDae"
# Preprocessing of data
# + colab={"base_uri": "https://localhost:8080/", "height": 195} id="S19_ezJalFW0" outputId="0b090052-3fc4-4db4-de33-4f3bc1595c50"
train_data = []
for defects_id, sp in enumerate(disease_types):
for file in os.listdir(os.path.join(train_dir, sp)):
train_data.append(['{}/{}'.format(sp, file), defects_id, sp])
train = pd.DataFrame(train_data, columns=['File', 'DiseaseID','Disease Type'])
train.tail()
# + [markdown] id="f6ytpnXFyIGp"
# Taking seed in order to get consistent results.
# + colab={"base_uri": "https://localhost:8080/", "height": 195} id="9pU-hQNLlFW1" outputId="ab4a02a6-73b7-40e7-ea74-66902d246818"
# Randomize the order of training set
SEED = 42
train = train.sample(frac=1, random_state=SEED)
train.index = np.arange(len(train)) # Reset indices
train.head()
# + [markdown] id="3Xuty6hZyPwB"
# Plotting a histogram between disease ID and its frequency.
# + colab={"base_uri": "https://localhost:8080/", "height": 298} id="TtLApBc9lFW1" outputId="4b4bf3ae-3a14-4d00-884e-9ac83f1d5897"
# Plot a histogram
plt.hist(train['DiseaseID'])
plt.title('Frequency Histogram of Species')
plt.figure(figsize=(12, 12))
plt.show()
# + [markdown] id="9GzpBm7NybD2"
# Displaying images for different species
# + colab={"base_uri": "https://localhost:8080/", "height": 687} id="CUmriTdAlFW1" outputId="e291f7aa-19f5-43c8-915a-97b2f2fd1cf9"
# Display images for different species
def plot_defects(defect_types, rows, cols):
fig, ax = plt.subplots(rows, cols, figsize=(12, 12))
defect_files = train['File'][train['Disease Type'] == defect_types].values
n = 0
for i in range(rows):
for j in range(cols):
image_path = os.path.join(data_dir, defect_files[n])
ax[i, j].set_xticks([])
ax[i, j].set_yticks([])
ax[i, j].imshow(cv2.imread(image_path))
n += 1
# Displays first n images of class from training set
plot_defects('Tomato_Bacterial_spot', 5, 5)
# + [markdown] id="fFT0MeAdyl6O"
# Color and size preprocessing of dataset
# + id="mjaLZ8XTlFW2"
IMAGE_SIZE = 64
def read_image(filepath):
return cv2.imread(os.path.join(data_dir, filepath)) # Loading a color image is the default flag
# Resize image to target size
def resize_image(image, image_size):
return cv2.resize(image.copy(), image_size, interpolation=cv2.INTER_AREA)
# + [markdown] id="Z_XPeSl4ytwg"
# Data Normalization
# + colab={"base_uri": "https://localhost:8080/"} id="MD_Y1Fr6lFW2" outputId="705f8d40-8d69-4290-f068-e50425448c20"
X_train = np.zeros((train.shape[0], IMAGE_SIZE, IMAGE_SIZE, 3))
for i, file in tqdm(enumerate(train['File'].values)):
image = read_image(file)
if image is not None:
X_train[i] = resize_image(image, (IMAGE_SIZE, IMAGE_SIZE))
# Normalize the data
X_Train = X_train / 255.
print('Train Shape: {}'.format(X_Train.shape))
# + [markdown] id="JgNSDr5Sy2QG"
# Data Encoding
# + id="InG5RyTflFW2"
Y_train = train['DiseaseID'].values
Y_train = to_categorical(Y_train, num_classes=15)
# + [markdown] id="j5FcbJcDy1X0"
# Splitting the dataset in to testing and training sub-datasets.
# + id="cbAHp6BwlFW3"
BATCH_SIZE = 64
# Split the train and validation sets
X_train, X_val, Y_train, Y_val = train_test_split(X_Train, Y_train, test_size=0.2, random_state=SEED)
# + [markdown] id="dSUQ-s37zE-3"
# Displaying the images with their labels in training dataset.
# + colab={"base_uri": "https://localhost:8080/", "height": 293} id="mgytCMtulFW3" outputId="84f4a090-0df4-43b9-e517-f61095536dff"
fig, ax = plt.subplots(1, 3, figsize=(15, 15))
for i in range(3):
ax[i].set_axis_off()
ax[i].imshow(X_train[i])
ax[i].set_title(disease_types[np.argmax(Y_train[i])])
# + [markdown] id="Hrzx_dTNzPH6"
# Parametrization
# + id="_xfwcJQ6lFW3"
EPOCHS = 50
SIZE=64
N_ch=3
# + [markdown] id="-q5hv2AvzRhU"
# Building DenseNet 121 Model
# + id="Ta6F8JVflFW3"
def build_densenet():
densenet = DenseNet121(weights='imagenet', include_top=False)
input = Input(shape=(SIZE, SIZE, N_ch))
x = Conv2D(3, (3, 3), padding='same')(input)
x = densenet(x)
x = GlobalAveragePooling2D()(x)
x = BatchNormalization()(x)
x = Dropout(0.5)(x)
x = Dense(256, activation='relu')(x)
x = BatchNormalization()(x)
x = Dropout(0.5)(x)
# multi output
output = Dense(15,activation = 'softmax', name='root')(x)
# model
model = Model(input,output)
optimizer = Adam(lr=0.002, beta_1=0.9, beta_2=0.999, epsilon=0.1, decay=0.0)
model.compile(loss='categorical_crossentropy', optimizer=optimizer, metrics=['accuracy'])
model.summary()
return model
# + [markdown] id="_pSRv727zYQf"
# Fitting the model onto the dataset. ( To increase the accurary increase the number of epocs upto 50)
# + colab={"base_uri": "https://localhost:8080/"} id="JWp53s6HlFW4" outputId="0255311a-bb41-4123-9b87-01f1d2ef11d1"
model = build_densenet()
annealer = ReduceLROnPlateau(monitor='val_accuracy', factor=0.5, patience=5, verbose=1, min_lr=1e-3)
checkpoint = ModelCheckpoint('model.h5', verbose=1, save_best_only=True)
# Generates batches of image data with data augmentation
datagen = ImageDataGenerator(rotation_range=360, # Degree range for random rotations
width_shift_range=0.2, # Range for random horizontal shifts
height_shift_range=0.2, # Range for random vertical shifts
zoom_range=0.2, # Range for random zoom
horizontal_flip=True, # Randomly flip inputs horizontally
vertical_flip=True) # Randomly flip inputs vertically
datagen.fit(X_train)
# Fits the model on batches with real-time data augmentation
hist = model.fit(datagen.flow(X_train, Y_train, batch_size=BATCH_SIZE),
steps_per_epoch=X_train.shape[0] // BATCH_SIZE,
epochs=EPOCHS,
verbose=2,
callbacks=[annealer, checkpoint],
validation_data=(X_val, Y_val))
model.save("dhaan.h5")
# + [markdown] id="wt-WhKB_hOdo"
# Creating the tar file of the model
# + colab={"base_uri": "https://localhost:8080/"} id="PFRAoLJ5Z-V1" outputId="39252d8d-d0b3-46e2-d293-d7f06f0bac8d"
# !tar -zcvf model.tgz model.h5
# + [markdown] id="vUD1z0iOhT1K"
# Checking whether all the required files are there.
# + colab={"base_uri": "https://localhost:8080/"} id="Ia8K3ncOaJby" outputId="b6ec75a2-8438-4113-b134-c9ec7c9bc64c"
# !ls -l
# + [markdown] id="aZoF9PSihZuv"
# Loading the model.
# + id="7YhqKwiYXqWp"
from keras.models import load_model
# returns a compiled model
# identical to the previous one
model = load_model('//content/sample_data/arc/dhaani.h5')
# + id="mYhf8DrwGbRB"
# + colab={"base_uri": "https://localhost:8080/", "height": 337} id="Z2btESQalFW5" outputId="fb8e6515-60ef-4cf5-d243-1a187f4398ff"
import numpy as np
import matplotlib.pyplot as plt
from skimage import io
from keras.preprocessing import image
#path='imbalanced/Scratch/Scratch_400.jpg'
img = image.load_img('/content/sample_data/arc/0a8a68ee-f587-4dea-beec-79d02e7d3fa4___RS_Early.B 8461.JPG', grayscale=False, target_size=(64, 64))
show_img=image.load_img('/content/sample_data/arc/0a8a68ee-f587-4dea-beec-79d02e7d3fa4___RS_Early.B 8461.JPG', grayscale=False, target_size=(200, 200))
disease_class = ['Pepper__bell___Bacterial_spot','Pepper__bell___healthy','Potato___Early_blight','Potato___Late_blight','Potato___healthy','Tomato_Bacterial_spot','Tomato_Early_blight','Tomato_Late_blight','Tomato_Leaf_Mold','Tomato_Septoria_leaf_spot','Tomato_Spider_mites_Two_spotted_spider_mite','Tomato__Target_Spot','Tomato__Tomato_YellowLeaf__Curl_Virus','Tomato__Tomato_mosaic_virus','Tomato_healthy']
x = image.img_to_array(img)
x = np.expand_dims(x, axis = 0)
#x = np.array(x, 'float32')
x /= 255
custom = model.predict(x)
print(custom[0])
#x = x.reshape([64, 64]);
#plt.gray()
plt.imshow(show_img)
plt.show()
a=custom[0]
ind=np.argmax(a)
print('Prediction:',disease_class[ind])
Disease_name = disease_class[ind]
# + colab={"base_uri": "https://localhost:8080/"} id="SW1j-d-vmOXk" outputId="df35c96d-fb85-4fb1-eb65-e78ca8b169cc"
if(Disease_name == "Tomato__Tomato_mosaic_virus"):
print("""Cultural control :\n\nUse certified disease-free seed or treat your own seed.\n
Soak seeds in a 10% solution of trisodium phosphate (Na3PO4) for at least 15 minutes.\n
Or heat dry seeds to 158 °F and hold them at that temperature for two to four days.\n
Purchase transplants only from reputable sources. Ask about the sanitation procedures they use to prevent disease.\n
Inspect transplants prior to purchase. Choose only transplants showing no clear symptoms.\n
Avoid planting in fields where tomato root debris is present, as the virus can survive long-term in roots.\n
Wash hands with soap and water before and during the handling of plants to reduce potential spread between plants.\n
Disinfect tools regularly — ideally between each plant, as plants can be infected before showing obvious symptoms.\n
Soaking tools for 1 minute in a 1:9 dilution of germicidal bleach is highly effective.\n
Or a 1-minute soak in a 20% weight/volume solution of nonfat dry milk and water is also very effective.\n
When pruning plants, have two pruners and alternate between them to allow proper soaking time between plants.\n
Avoid using tobacco products around tomato plants, and wash hands after using tobacco products and before working with the plants.\n
Tobacco in cigarettes and other tobacco products may be infected with either ToMV or TMV, both of which could spread to the tomato plants.\n
Scout plants regularly. If plants displaying symptoms of ToMV or TMV are found, remove the entire plant (including roots), bag the plant, and send it to the University of Minnesota Plant Diagnostic Clinic for diagnosis.\n
If ToMV or TMV is confirmed, employ stringent sanitation procedures to reduce spread to other plants, fields, tunnels and greenhouses.\n
Completely pull up and burn infected plants. Do not compost infected plant material.\n
After working with diseased plants, thoroughly disinfect all tools and hands as outlined above.\n
For added security against spread, keep separate tools for working in the diseased area and avoid working with healthy plants after working in an area with diseased plants.\n
At the end of the season, burn all plants from diseased areas, even healthy-appearing ones, or bury them away from vegetable production areas.\n
Disinfect stakes, ties, wires or any other equipment between growing seasons using the methods noted above.\n
Chemical control\n
There are currently no chemical options that are effective against either virus.\n""")
print("Control strategy would be provided for all the diseases in future")
| Dhaan_Crop_Doctor.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # scmap for label transfer between our scRNAseq ref and snRNA from multiomics dualRNA-ATAC
#
# +
import numpy as np
import pandas as pd
import scanpy as sc
import os
import sys
import warnings
import anndata
warnings.filterwarnings('ignore')
def MovePlots(plotpattern, subplotdir):
os.system('mkdir -p '+str(sc.settings.figdir)+'/'+subplotdir)
os.system('mv '+str(sc.settings.figdir)+'/*'+plotpattern+'** '+str(sc.settings.figdir)+'/'+subplotdir)
sc.settings.verbosity = 3 # verbosity: errors (0), warnings (1), info (2), hints (3)
sc.settings.figdir = './figures_sn/scmap/'
sc.logging.print_header()
sc.settings.set_figure_params(dpi=80) # low dpi (dots per inch) yields small inline figures
sys.executable
# -
# ## Load data
# ### SC ref
sc_adataR = sc.read('/nfs/team292/lg18/with_valentina/FCA-M5-annotatedCluster4Seurat.h5ad')
# +
# Add cluster name and doublet information
clu_annot = pd.read_csv('/nfs/team292/lg18/with_valentina/FCA-M5-annotatedCluster4Seurat.csv', header=0, index_col=0)
import collections
if collections.Counter(sc_adataR.obs.index) == collections.Counter(clu_annot.index):
print ("The lists are identical")
else :
print ("The lists are not identical")
vars2import = ['clusters_manual','louvain', 'is_doublet', 'scrublet_cluster_score', 'scrublet_score']
for var in vars2import:
sc_adataR.obs[var] = clu_annot[var]
# -
sc_adataR.obs['clusters'] = sc_adataR.obs['clusters_manual']
sc_adataR.obs['clusters'].value_counts(dropna = False)
sc_adataR = sc_adataR[[ 'female' in i for i in sc_adataR.obs['sex'] ]]
sc_adataR = sc_adataR[[ i==14 for i in sc_adataR.obs['PCW'] ]]
sc_adataR = sc_adataR[[ 'Doublet' not in i for i in sc_adataR.obs['clusters'] ]]
sc_adataR = sc_adataR[[ 'Metanephros' not in i for i in sc_adataR.obs['clusters'] ]]
sc_adataR = sc_adataR[[ 'lowQC' not in i for i in sc_adataR.obs['clusters'] ]]
supporting = pd.read_csv('/nfs/team292/lg18/with_valentina/supporting_nocycling_annotation.csv', index_col = 0)
print(supporting['annotated_clusters'].value_counts())
supporting = supporting[supporting['annotated_clusters'].isin(['sLGR5', 'sKITLG', 'coelEpi', 'sPAX8b', 'sPAX8m', 'preGC_I_OSR1',
'ovarianSurf', 'preGC_II', 'preGC_II_hypoxia', 'preGC_III', 'preGC_III_Notch'])]
print(supporting['annotated_clusters'].value_counts())
stromal = pd.read_csv('/nfs/team292/lg18/with_valentina/mesenchymal_annotated.csv', index_col = 0)
print(stromal['annotated_clusters'].value_counts())
# stromal = stromal.replace({'annotated_clusters': ['M_HOXC6', 'M_SFRP2', 'M_cycling', 'MullDuctFib_LGR5', 'M_ALDH1A2', 'M_CRABP1_BMP4', 'M_ISL1_BMP4', 'M_CD24']}, 'M_mesonephros')
stromal = stromal[stromal['annotated_clusters'].isin(['M_MGP', 'M_ALX1', 'M_prog_ISL1', 'M_MullDuct_LGR5', 'Gi', 'Oi'])]
print(stromal['annotated_clusters'].value_counts())
germcells = pd.read_csv('/nfs/team292/lg18/with_valentina/germcells_annotation.csv', index_col = 0)
print(germcells['annotated_clusters'].value_counts())
germcells = germcells[germcells['annotated_clusters'].isin(['PGC', 'oogonia_meiosis', 'oogonia_STRA8', 'oocyte'])]
print(stromal['annotated_clusters'].value_counts())
supporting_germ = supporting.append(germcells)
supporting_germ = supporting_germ.append(stromal)
mapping_dict = supporting_germ['annotated_clusters'].to_dict()
sc_adataR.obs['annotated_clusters'] = sc_adataR.obs_names.map(mapping_dict)
sc_adataR.obs['annotated_clusters'].value_counts(dropna = False)
# +
# Replace NaN with big clusters
sc_adataR.obs['annotated_clusters'] = sc_adataR.obs['annotated_clusters'].fillna(sc_adataR.obs['clusters'])
# Remove
sc_adataR = sc_adataR[[i not in ['Supporting_female', 'Mesenchymal_GATA2_NR2F1', 'Sertoli', 'Mesenchymal_LHX9_ARX', 'Coel Epi', 'Germ cells', 'Stromal', 'lowQC'] for i in sc_adataR.obs['annotated_clusters']]]
sc_adataR.shape
sc_adataR.obs['annotated_clusters'].value_counts(dropna = False)
# -
# ### SN
sn_adataR = sc.read('/nfs/team292/lg18/with_valentina/gonadsV1_freezed/dual_snRNAsnATAC_female_Hrv39.h5ad')
sn_adataR = sn_adataR[[i not in ['1'] for i in sn_adataR.obs['leiden']]]
sn_adata = anndata.AnnData(X=sn_adataR.raw.X, var=sn_adataR.raw.var, obs=sn_adataR.obs, asview=False)
# ## Intersect genes
# +
sc_genes = sc_adataR.var_names.to_list()
sn_genes = sn_adata.var_names.to_list()
common_genes = list(set(sc_adataR.var.index.values).intersection(sn_adata.var.index.values))
print('common genes: ', len(common_genes))
sn_adata = sn_adata[:,common_genes]
sc_adataR = sc_adataR[:,common_genes]
print(sn_adata.X.shape)
print(sc_adataR.X.shape)
# -
# ## R2PY setup
#
import rpy2.rinterface_lib.callbacks
import logging
# Ignore R warning messages
#Note: this can be commented out to get more verbose R output
rpy2.rinterface_lib.callbacks.logger.setLevel(logging.ERROR)
import anndata2ri
anndata2ri.activate()
# %load_ext rpy2.ipython
# ## scmap
# + language="R"
# library(Matrix)
# library(scmap)
# + magic_args="-i sc_adataR" language="R"
# sc_adataR
# + magic_args="-i sn_adata" language="R"
# rowData(sn_adata)$feature_symbol <- rownames(sn_adata)
# sn_adata
# + language="R"
#
# counts <- assay(sc_adataR, "X")
# libsizes <- colSums(counts)
# size.factors <- libsizes/mean(libsizes)
# logcounts(sc_adataR) <- log2(t(t(counts)/size.factors) + 1)
# assayNames(sc_adataR)
# + language="R"
#
# rowData(sc_adataR)$feature_symbol <- rownames(sc_adataR)
# logcounts(sc_adataR) <- as.matrix(logcounts(sc_adataR))
# sc_adataR <- selectFeatures(sc_adataR, n_features = 400, suppress_plot = FALSE)
# print(table(rowData(sc_adataR)$scmap_features))
# + language="R"
#
# sc_adataR <- indexCluster(sc_adataR, cluster_col = "annotated_clusters")
# print(head(metadata(sc_adataR)$scmap_cluster_index))
# heatmap(as.matrix(metadata(sc_adataR)$scmap_cluster_index))
# + language="R"
#
# counts <- assay(sn_adata, "X")
# libsizes <- colSums(counts)
# size.factors <- libsizes/mean(libsizes)
# logcounts(sn_adata) <- log2(t(t(counts)/size.factors) + 1)
# logcounts(sn_adata) <- as.matrix(logcounts(sn_adata))
# assayNames(sn_adata)
# + magic_args="-o scmapCluster_results" language="R"
#
# scmapCluster_results <- scmapCluster(
# projection = sn_adata, threshold = 0.5,
# index_list = list(
# sc_adataR = metadata(sc_adataR)$scmap_cluster_index
# )
# )
# + language="R"
#
# print(head(scmapCluster_results$scmap_cluster_labs))
# print(head(scmapCluster_results$scmap_cluster_siml))
# print(head(scmapCluster_results$combined_labs))
# -
scmapCluster_results
labels = list(scmapCluster_results[0])
sn_adataR.obs['scmap_labels'] = labels
sn_adataR.obs['scmap_labels'].value_counts(dropna = False)
similarities = scmapCluster_results[1]
similarities = [val for sublist in similarities for val in sublist]
#similarities
sn_adataR.obs['scmap_similarities'] = similarities
sn_adataR.obs[['scmap_labels', 'scmap_similarities']].to_csv('/nfs/team292/lg18/with_valentina/dual_snRNAsnATAC_female_Hrv39_scmap_predictions_highRes.csv')
sc.pl.umap(sn_adataR, color='scmap_labels')
sc.pl.umap(sn_adataR, color='scmap_similarities')
sc.pl.umap(sn_adataR, color='clusters')
| sn5b_scmap_scRNAref_female.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Likelihood - Demo
# Welcome to the Likelihood Demo! This will present a **truncated** version of Likelihood, one which utilizes working features to give the user a good idea of what Likelihood actually does, and how it can be implemented on a dataset!
# ### A Quick Rundown:
#
# Likelihood is a data quality monitoring engine that measures the surprise, or entropy, of members of a given dataset. To learn more about the basic theory behind it, one may click on the 2 links below:
#
# https://en.wikipedia.org/wiki/Entropy_(information_theory)
#
# http://people.math.harvard.edu/~ctm/home/text/others/shannon/entropy/entropy.pdf
#
# The basic essence is: uncertainty is maximized (it is most regular) in cases where the probability structure of a dataset is chaotic, meaning we don't have much information about it. However, when we can identify some given patterns about the probability structure of a dataset, we know that data members following these rules are not particularly chaotic. They are structured, and thus unsurprising. It is when these patterns are defied that the entropy shoots up to irregular heights. It is this percise rule defying approach that Likelihood uses to find outliers within a structured dataset.
#
# Likelihood began as a numerical-estimation focused tool, but currently it works quite well with numerical, categorical, and timestamp data. Its functional approaches are mapped out below:
#
# 1. **Bootstrapping** - Building a distribution using the properties of a bootstrap, this approach uses the bootstrap to capture standard counts for values by TimeStamp and finds anomaly if test-set counts are a certain level off expected training set ratios.
#
#
# 2. **Time Series** - Using Facebook Prophet, Time Series evaluation puts surprising event in the context of the time in which they happened. Building a pattern off these approximations and understanding, the Time Series tool predicts the future for the test-set and raises an issue if expected future counts fall off.
#
#
# 3. **Kernel Density** - Smoothing a distribution so that certain properties can be utilized, Kernel Density fits the data under a curve depending on the data's variation and approximates which values in a distribution are unlikely by virtue of magnitude, thus finding the most surprising Data.
#
#
# 4. **PCA** - Using Dimensionality Reduction, PCA attributes the variation in the data to several significant columns which are used to compute bias row-wise. This approach is combined with the column based kernel density approach to truly triangulate the percise location of numeric data-error, and PCA's surprise metric is thus grouped with Kernel Density's.
#
#
# 5. **Relative Entropy Model for Categorical Data** - Much in the spirit of grammar, this relative entropy its own rules (expected formatting and behavior) for data, and obtains surprise based off the strictness of the rule that the data defies (defying a stricter rule would inherently be more chaotic)
#
#
# 6. **TimeStamp Intervals** - This Kernel Density approach computes similarly to the numerical Kernel Density, but this time orders the time intervals in the dataset and procceeds to test if there is a weird interval in which no data/ too much data was recorded.
#
#
# 7. **In Progress**: Mutual Entropy for Mixed Numeric and Categorical Data
#
# Ultimately, Likelihood should become a functional tool that can build functional distributions without the need for any context. Currently it functions more as a copilot
# Imports for project purposes
# Full Project imports
import pandas as pd
import math as mt
import dateutil
from datetime import datetime, timedelta
import requests as rd
import numpy as np
from sklearn import neighbors, decomposition
from sklearn.decomposition import PCA
from sklearn.preprocessing import StandardScaler
import smtplib
import scipy.stats as st
import os
from datetime import datetime, timedelta
from pandas.api.types import is_numeric_dtype
import copy
# parameters (will put into JSON config file later)
params = {"fName": "pd_calls_for_service_2020_datasd.csv", # local CSV file only right now
"ts": "date_time", # Timestamp for when the event happened in the world
"bootstrapResamples":1000, # should probably be 10k for better accuracy, but much worse speed
"maxTrainingSizeMultiple":10, # if there is more than X times more training data, trim to most recent
"maxCategories":100, # maximum number of categories in a column - if higher we skip
"minCategoryCount":10, # don't report boostrap surprise if a category has lower count
}
# +
# Loading data into project
def load_data(dataset_link, category):
if(category == "html"):
return pd.read_html(dataset_link)
else:
if(category == "excel"):
return pd.read_excel(dataset_link)
else:
return pd.read_csv(dataset_link)
df = load_data("pd_calls_for_service_2020_datasd.csv", "csv")
# -
# The Data used throughout this part of the demo comes from the San Diego County Police Calls for Service Dataset, it will be used to show the effect of Likelihood's Time-Series Methods
df
# +
# Converts Timetamp column of DataFrame to a legitimate timestamp
def convertToDateTime(df, timestamp):
df[timestamp] = pd.to_datetime(df[timestamp], format='%Y%m%d %H:%M:%S')
return df
# Assignments for computational purposes
df['ts'] = df['date_time']
batchHours = 24*7
df = convertToDateTime(df, 'ts')
df
# +
# Splits data into train and test set based on date/time
def split_train_test(df, batchHours):
maxTs = max(df['ts'])
batchTs = maxTs - timedelta(hours = batchHours)
testDf = df[df['ts'] > batchTs]
trainDf = df[df['ts'] < batchTs]
return trainDf, testDf
trainDf, testDf = split_train_test(df, batchHours)
# +
# Helpers and Math
def pValue(data, threshold, result):
p_larger = sum(np.array(data) >= threshold) / len(data)
p_smaller = sum(np.array(data) <= threshold) / len(data)
p = min(p_larger, p_smaller)
# only use gaussian p-value when there is variation, but bootsrap p = 0
stdev = np.std(data)
if stdev == 0 or p != 0:
p_gauss = p
else:
p_gauss = st.norm(np.mean(result['bootstrap_counts']), stdev).cdf(result['count'])
p_gauss = min(p_gauss,1-p_gauss)
return p_gauss
def trimTraining(trainDf, params):
# trim to most recent
trainDf = trainDf.sort_values(params['ts'], ascending =False)
trainDfTrimmed = trainDf[:params['maxTrainingSizeMultiple']*len(testDf)]
return trainDfTrimmed
# -
# Returns names of categorical columns
def getCategoricalColumnNames(df):
columnNames = []
for columnName in df.keys():
if (type (df[columnName].iloc[0])) == str:
columnNames.append(columnName)
return columnNames
# +
def bootstrap(trainDf, testDf, params):
# get all of the string columns
columnNames = getCategoricalColumnNames(testDf)
bootstrapDf = trimTraining(trainDf, params)
# set up dict, add counts
results = {}
for columnName in columnNames:
# if it isn't a string column, reject it
if type(testDf[columnName].iloc[0]) != str:
continue
categories = (bootstrapDf[columnName].append(testDf[columnName])).unique()
if len(categories) > params['maxCategories']:
continue
results[columnName] = {}
testCounts = testDf[columnName].value_counts(dropna = False)
for i in np.arange(1,len(categories) -1):
if(pd.isna(categories[i])):
categories = np.delete(categories, i)
for category in categories:
results[columnName][category] = {'bootstrap_counts':[],
'count':testCounts.get(category,0)}
# resample, add boostrap counts
for ii in range(params['bootstrapResamples']):
# Draw random sample from training
sampleDf = bootstrapDf.sample(len(testDf), replace=True)
for columnName in results.keys():
# count by category
trainCounts = sampleDf[columnName].value_counts(dropna = False)
# put results in dict
for category in results[columnName].keys():
boostrapCount = trainCounts.get(category,0)
results[columnName][category]['bootstrap_counts'].append(boostrapCount)
# convert to records, add p-values
bootstrap_results = []
for columnName in results.keys():
for category in results[columnName].keys():
result = results[columnName][category]
estimatedCount = int(np.round(np.mean(result['bootstrap_counts'])))
# don't report entries with very low predicted and actual counts
if estimatedCount < params['minCategoryCount'] and result['count'] < params['minCategoryCount']:
continue
p = pValue(result['bootstrap_counts'],result['count'], result)
categoryName = category
# Backup
if not category:
categoryName = "NULL"
bootstrap_results.append({"column":columnName,
"category":categoryName,
"count":result['count'],
"p": p,
"estimated_count":estimatedCount,
})
# Sorting by P-values and obtaining Surprise of each
if(np.count_nonzero(p)>0):
resultsDf = pd.DataFrame.from_records(bootstrap_results).sort_values('p')
resultsDf['surprise'] = -np.log2(resultsDf['p'])
return resultsDf
bootstrap(trainDf, testDf, params)
# -
# ## TimeSeries Approximation
from fbprophet import Prophet
def truncateTs(ts):
return ts.replace(minute=0, second=0, microsecond=0)
# +
# Groups data by value counts and returns a table with corresponding y values according to
def group_and_build_time_table(truncatedData):
groupedCounts = truncatedData.value_counts()
prophetDf = pd.DataFrame({'ds':groupedCounts.index,'y':np.log10(groupedCounts.values)})
return prophetDf
truncatedData = trainDf['ts'].apply(truncateTs)
prophetDf = group_and_build_time_table(truncatedData)
prophetDf
# +
# Takes in the the dataset and the prophet dataset returned by the ast option
def train_model_on_country(testDf, prophetDf, country = "US"):
# Train model
m = Prophet(#daily_seasonality = True,
#yearly_seasonality = False,
#weekly_seasonality = True,
#growth='linear',
interval_width=0.68 # one sigma
)
m.add_country_holidays(country_name=country)
m.fit(prophetDf)
return m
# Applies Prophet analytics to create a forecast based on hours
def predict_future(testDf,m, timestamp = "date_time"):
# Takes in trained model and predicts the future
# find number of hours to preduct: ceil of hours in testDf
testDf = testDf.assign(ts = testDf.get(timestamp))
#If a column is string, convert to date/time
if(testDf.applymap(type).eq(str).any()['ts']):
testDf['ts'] = pd.to_datetime(testDf['ts'])
timeDelta = max(testDf['ts']) -min(testDf['ts'])
hours = int(timeDelta.days*24 + timeDelta.seconds/(60*60))+1
future = m.make_future_dataframe(periods = hours, freq = 'H')
forecast = m.predict(future)
return forecast, testDf
m = train_model_on_country(testDf, prophetDf)
forecast, testDf = predict_future(testDf, m)
forecast
# +
# Takes in truncated test data (column), spits out out the prophet results
def find_surprise(truncatedData, forecast):
groupedCounts = truncatedData.value_counts()
prophetTestDf = pd.DataFrame({'ds':groupedCounts.index,
'y':np.log10(groupedCounts.values),
'y_linear':groupedCounts.values})
# find p-value
prophet_results = []
# Comparing test and training set data for identical intervals
for ii in range(len(prophetTestDf)):
ts = prophetTestDf['ds'][ii]
fcstExample = forecast[forecast['ds'] == ts]
mean = fcstExample['yhat'].iloc[0]
stdev = (fcstExample['yhat_upper'].iloc[0] - fcstExample['yhat_lower'].iloc[0])/2
# Calculating the P-value
p = st.norm(mean, stdev).cdf(prophetTestDf['y'][ii])
p = min(p,1-p)
prophet_results.append({"column":"Forecast",
"category":str(ts),
"count":prophetTestDf['y_linear'][ii],
"p": p,
"estimated_count":int(np.round(np.power(10,mean))),
})
# Obtaining Entropy of Time-Series values
prophetResultsDf = pd.DataFrame.from_records(prophet_results).sort_values('p')
prophetResultsDf['surprise'] = -np.log2(prophetResultsDf['p'])
return prophetResultsDf
#Group the test data
truncatedData = testDf['ts'].apply(truncateTs)
find_surprise(truncatedData, forecast)
# +
# Takes in a model that has been trained on country, plots graphs for visualization
def visualize(m, forecast):
# Model visualization
fig = m.plot(forecast)
fig = m.plot_components(forecast)
visualize(m, forecast)
# -
# # Kernel Density
# +
#https://www.nbastuffer.com/2019-2020-nba-team-stats/
def inp(default = 1, default2 = "https://www.nbastuffer.com/2019-2020-nba-team-stats/"):
# If our default dataset is changed, obtain some input
if(default2 != "https://www.nbastuffer.com/2019-2020-nba-team-stats/"):
nam = input()
else:
nam = default2
frame = pd.read_html(nam)
first_table = frame[default]
return first_table
first_table = inp(0)
first_table
# -
# ## Different Kernels Attached Below
# Using cosine kernel function to get estimate for log density
def cosKernel(stat):
stat = stat.to_numpy().reshape(-1,1)
l = neighbors.KernelDensity(kernel = 'cosine').fit(stat)
cos_density = l.score_samples(stat)
return cos_density
# Using gaussian kernel function to get estimate for log density
def gaussKernel(stat):
stat = stat.to_numpy().reshape(-1,1)
l = neighbors.KernelDensity(kernel = 'gaussian').fit(stat)
density = l.score_samples(stat)
return density
# Using linear kernel function to get estimate for log density
def expKernel(stat):
stat = stat.to_numpy().reshape(-1,1)
l = neighbors.KernelDensity(kernel = 'exponential').fit(stat)
triDensity = l.score_samples(stat)
return triDensity
# Using epanechnikov kernel function to get estimate for log density
def parabolicKernel(stat):
stat = stat.to_numpy().reshape(-1,1)
l = neighbors.KernelDensity(kernel = 'epanechnikov').fit(stat)
epDensity = l.score_samples(stat)
return epDensity
# Drops non-numerical and nan values from a table
def pcaPrep(first_table):
# Finding all numerical components of the table so that pca can function
tabl = first_table.select_dtypes(include = [np.number])
tabl = tabl.dropna(1)
return tabl
# Specialized column based P-value function: double ended
def retPVal(col):
#Since we have a normal distribution, starting by obtaining the z-score
mean = col.mean()
std = np.std(col)
array = np.array([])
for i in np.arange(len(col)):
array = np.append(array, col.iloc[i] - mean)
#Now obtaining legitimate p-values
z_scores = array/std
for l in np.arange(len(z_scores)):
cdf = st.norm.cdf(z_scores[l])
z_scores[l] = min(cdf, 1-cdf)
return pd.Series(z_scores, index = col.index)
# Assigning initial kernal estimations
def kernelEstimator(indx, stat):
kernelEstimate = pd.DataFrame()
kernelEstimate = kernelEstimate.assign(Data_Index = indx, Data_Point = stat,Gaussian = gaussKernel(stat),
Epanechnikov = parabolicKernel(stat), Exponential = expKernel(stat),
Cosine = cosKernel(stat))
# temporary sort for some visualization of surprise
kernelEstimate = kernelEstimate.sort_values(by = "Gaussian", ascending = False)
return kernelEstimate
# Calculating their average
def surprise_estimator(kernelEstimation):
# Calculating maximum number of deviations from the mean
numDevMax = (kernelEstimation.get("Data_Point").max() - kernelEstimation.get("Data_Point").mean())/kernelEstimation.get("Data_Point").std()
numDevMin = (kernelEstimation.get("Data_Point").min() - kernelEstimation.get("Data_Point").mean())/kernelEstimation.get("Data_Point").std()
numDev = max(numDevMax, numDevMin)
# Assigning appropriate Kernel Estimator
if(numDev > 3.2):
metric = retPVal(kernelEstimation.get("Exponential"))
elif((numDev <=3.2) & (numDev >= 2)):
metric = retPVal(kernelEstimation.get("Gaussian"))
else:
metric = retPVal(kernelEstimation.get("Exponential")+kernelEstimation.get("Epanechnikov"))
# Surprise Metric
kernelEstimation = kernelEstimation.assign(Surprise = -np.log2(metric))
kernelEstimation = kernelEstimation.sort_values(by = "Surprise", ascending = False)
return kernelEstimation
# A grouping of the entire kernel estimation process
def surprise_Table(Table, index = "TEAM"):
temp = pcaPrep(Table)
# Checking if index given
if(isinstance(index, str)):
index = Table.get(index)
#Obtaining surprise of every individual column
sum_surprise = pd.Series(np.zeros(Table.shape[0]))
for col in temp.columns:
stat = temp.get(col)
KernelTable = kernelEstimator(index, stat)
KernelTable = surprise_estimator(KernelTable)
Table[col] = KernelTable.get("Surprise")
sum_surprise+=Table[col]
# Averaging our surprise so we can sort by it
sum_surprise = sum_surprise.array
Table = Table.set_index(index)
Table = Table.assign(mean_surprise = np.round(sum_surprise/Table.shape[1],2))
# Sorting table for easier visualization
Table = Table.sort_values(by = "mean_surprise", ascending = False)
return Table
surpriseTable = surprise_Table(first_table)
surpriseTable
# +
def obtain_variance_table(first_table):
# Scaling and preparing values for PCA
tabl = pcaPrep(first_table)
scaled_data = StandardScaler().fit_transform(tabl)
# Creating a PCA object
pca = PCA(n_components = (tabl.shape[1]))
pcaData = pca.fit_transform(scaled_data)
infoFrame = pd.DataFrame().assign(Column = ["PC" + str(i) for i in range(tabl.shape[1])], Variance_ratio = pca.explained_variance_ratio_ )
return infoFrame
obtain_variance_table(first_table)
# -
# Fit PCA model onto the data
def obtainPCAVals(componentNum, scaled_data):
pca = PCA(n_components = componentNum)
pcaData = pca.fit_transform(scaled_data)
return pcaData
# +
# Deciding how many columns need to be used: utilizing threashold of 95% of the explained variance
def elementDecider(infoFrame):
numSum = 0
counter = 0
# Continuing until we have accounted for 95% of the variance
for i in infoFrame.get("Variance_ratio"):
if(numSum < .95):
numSum += i
counter+=1
return counter
# Reducing dimensionality of data into pc's, only storing what is neccessary
def reducedData(infoFrame, scaled_data, indx):
numCols = elementDecider(infoFrame)
pcaData = obtainPCAVals(numCols, scaled_data)
pcaFrame = pd.DataFrame(pcaData)
# Dealing with potential index issues
pcaFrame = pcaFrame.set_index(indx)
return pcaFrame
# Visualization tool for seeing grouping of elements by pc
def displayReducedData(pcaVals, xNum = 0, yNum = 1):
# Ensuring that the elements given do not overacess table
if(xNum < pcaVals.shape[1]) & (yNum < pcaVals.shape[1]):
pcaVals.plot(kind = "scatter", x = 2, y = 3)
else:
print("You have overaccessed the number of elements, keep in mind there are only " + str(pcaVals.shape[1]) + " elements")
# +
#Summing p-values because PCA serves to check for systematic bias, whereas kernel density checks for accuracy
def sumRows(pcaVals):
sumArray = np.zeros(pcaVals.shape[0])
for i in np.arange(pcaVals.shape[1]):
values = pcaVals.get(str(i)).array
sumArray += abs(values)
sumArray /= pcaVals.shape[1]
#After obtaining sum, the average deviation from the expected value is averaged out, not taking in absolute value
# to check for systematic error
return sumArray
# Tests for systematic bias by row
def pcaRowOutliers(pcaVals):
P_val_table = pd.DataFrame()
#Creating a table of all the PCA p-values
for col in np.arange(0,pcaVals.shape[1]):
P_vals = retPVal(pcaVals.get(col))
i = str(col)
P_val_table[i] = P_vals
totalVar = sumRows(P_val_table)
#Calculating surprise by taking negative log
newVals = pcaVals.assign(Surprise = -np.log2(totalVar))
newVals = newVals.sort_values(by = "Surprise", ascending = False)
return newVals
# +
# Master method to run PCA as a whole
def runPCA(table, index):
processing_table = pcaPrep(table)
variance_table = obtain_variance_table(table)
pcaVals = reducedData(variance_table, StandardScaler().fit_transform(processing_table), table.get(index))
new_pca = pcaRowOutliers(pcaVals)
return new_pca
new_pca = runPCA(first_table, 'TEAM')
# -
new_pca
# +
# Combining PCA and Kernel density into one approach to obtain join probabilities
def pca_kernel_combo(pcaTable,kernelTable):
pcaSurpriseCol = new_pca.get("Surprise")
temp = pcaPrep(kernelTable)
for column in temp.columns:
# Finding geometric mean of two factors individually (updating our beliefs in a Bayesian manner)
kernelTable[column] = (kernelTable[column].multiply(pcaSurpriseCol)).apply(np.sqrt)
kernelTable = kernelTable.sort_values(by = "mean_surprise", ascending = False)
return kernelTable
surpriseTable = pca_kernel_combo(new_pca, surpriseTable)
surpriseTable
# -
# # Categorical Data
# Will examine whether or not a column is categorical, giving the user the opportunity to add additional numeric columns
def identifyCategorical(surpriseFrame):
categorical_list = []
for col in surpriseFrame.columns:
if(not(is_numeric_dtype(surpriseFrame[col]))):
categorical_list.append(col)
# Allows fixing of default assumption that numeric columns aren't categorical
print("Are there any numeric Columns you would consider categorical?(yes/no)")
while(input().upper() == "YES"):
print("Enter one such column:")
categorical_list.append(input())
print("Any more?")
return categorical_list
# ### **Running tests for: value type, its length, and whether or not it is missing (NaN)**
# +
# Returns suprise of type classification
def types(column):
value_types = column.apply(classifier)
counts = value_types.value_counts(normalize = True)
index = counts.index
values = counts.values
probs = value_types.apply(giveProb, args = (np.array(index), np.array(values)))
surpriseVal = probs.apply(surprise)
return surpriseVal
# Obtains the type of value, even if it is currently contained within a string
def classifier(value):
value = str(value)
# Boolean check done manually: this is an easy check
if(('True' in value) or ('False' in value )):
return 'boolean'
else:
if(value.isnumeric()):
return 'number'
else:
return 'string'
# Takes in a column and returns the surprise of each nan value being present (True) or not being present (False)
def nans(column):
nan_values = column.apply(isNan)
counts = nan_values.value_counts(normalize = True)
index = counts.index
values = counts.values
probs = nan_values.apply(giveProb, args = (np.array(index), np.array(values)))
surpriseVal = probs.apply(surprise)
return surpriseVal
# Takes in a column and returns the surprise of the length of each value in the column: the first and simplest of probabilistic tests
def lenCount(column):
column = column.apply(str)
counts = column.apply(len).value_counts(normalize = True)
index = counts.index
values = counts.values
column = column.apply(len)
probs = column.apply(giveProb, args = (np.array(index), np.array(values)))
surpriseVal = probs.apply(surprise)
return surpriseVal
# Calculates the surprise of a given value
def surprise(value):
return -np.log2(value)
# Given a numerical value, finds it equivalent within the set of indices and assigns it the proper probability
def giveProb(value, index, values):
for num in np.arange(len(index)):
if(value == index[num]):
return values[num]
return values[0]
# NaN's aren't equal to themselves
def isNan(x):
return x!=x
# -
# ### **Running tests for: Special Character sequence and Number of Unique Values**
# +
# Checks for special characters within a string, calculating surprise so as to identify which character combinations are chaotic
def special_char(column):
characters = column.apply(str).apply(char_identifier)
counts = characters.value_counts(normalize = True)
index = counts.index
values = counts.values
probs = characters.apply(giveProb, args = (np.array(index), np.array(values)))
surpriseVal = probs.apply(surprise)
return surpriseVal
# Checks if a single entry of any data type contains special symbols and returns all that it contains
def char_identifier(entry):
charList = np.array(['<', '>', '!', '#','_','@','$','&','*','^', ' ', '/', '-','"','(', ',', ')', '?', '.'])
ret_string = ""
for i in charList:
if(i in entry):
ret_string += i
return ret_string
# Simpler approach here: if the value counts of certain elements are greater when they should be unique, they are more suprising
# If they are non-unique when they are supposed to be unique, also more surprising. Done with binary classification
def uniques(column):
# Counting number of each value and returning whether or not it is a singular unique value,
#then counting truly unique values
column = column.replace({np.nan: "np.nan"})
vals = column.value_counts().apply(isunique)
vals = column.apply(unique_assignment, args = [vals])
counts = vals.value_counts(normalize = True)
index = counts.index
values = counts.values
probs = vals.apply(giveProb, args = (np.array(index), np.array(values)))
surpriseVal = probs.apply(surprise)
# Note: if all values unique/non unique this will provide definite outcome because no room for uncertainty
return surpriseVal
# Returns whether the count of a value is 1
def isunique(val):
return (val == 1)
# Maintains individual values without grouping while assigning unique / nonunique probabilities. To be used on original column
def unique_assignment(val, column):
value = column[column.index == val]
return value.iloc[0]
# +
# Obtains a date time object and treats this as numerical rather than categorical value
def obtainCatSurprise(table):
cols = identifyCategorical(table)
for col in cols:
# Obtaining individual relative entropies, averaging them out, finding their p-values and calculating final surprise
values = table.get(col)
temp = (uniques(values)+special_char(values)+ nans(values) + types(values)+lenCount(values))/5
table[col] = -np.log2(retPVal(temp))
table = table.replace({np.nan:0})
return table
dataset = pd.read_excel("sampleDataSet.xlsx")
dataset.head(40)
# -
categoricalSurprise = obtainCatSurprise(dataset).head(80)
# +
# Assigning colors to problematic values (still grouped with indices so easy to tell)
# Yellow: mild concern, Orange: serious concern, red - major concern
def designer(frame):
threshold1 = 5
threshold2 = 10
threshold3 = 15
print("Would you like to reset default issue alert thresholds?")
if(input().upper() == 'YES'):
print("Mild concern threshold (in probability (percentage) of issue being present):")
threshold1 = float(input())
print("Moderate concern threshold (in probability (percentage) of issue being present)")
threshold2 = float(input())
print("Serious concern threshold (in probability (percentage) of issue being present)")
threshold3 = float(input())
temp = pcaPrep(frame)
styler = frame.style
for col in temp.columns:
frame = styler.applymap(lambda x: 'background-color: %s' % 'yellow' if x > threshold1 else 'background-color: %s' % 'light-gray', subset = [col])
frame = styler.applymap(lambda x: 'background-color: %s' % 'orange' if x > threshold2 else 'background-color: %s' % 'light-gray', subset = [col])
frame = styler.applymap(lambda x: 'background-color: %s' % 'red' if x > threshold3 else 'background-color: %s' % 'light-gray', subset = [col])
return frame
designer(categoricalSurprise)
# -
# ## Date/Time Interval Approximation
# +
# Calculation of date time entropies
def dateTimeClassifier(column):
# Conversion to proper format
if (type(column.iloc[0]) == str):
column = convert_to_dateTime(column)
# Unix timestamps for ease of calculation
unixCol = column.apply(convertToUnix).to_numpy()
# Finding time intervals
difference_array = np.append(np.array([]), np.diff(unixCol))
timeFrame = (pd.DataFrame().assign(index = np.arange(1,len(unixCol)), Times_diff = difference_array))
dateSurprise = surprise_Table(timeFrame, 'index')
return (dateSurprise.sort_values(by = ['Times_diff']))
# If date-value is given as a string, convert to date- time format first
def convert_to_dateTime(column):
return pd.to_datetime(column, format='%Y%m%d %H:%M:%S')
# Converting the date to unix format for ease of calculations
def convertToUnix(value):
return (value - datetime(1970, 1, 1)).total_seconds()
dateTimeClassifier(trainDf[:3000].get("date_time"))
# -
# # The Next Step
# The next step in the proccess as of now is releasing the Time Series as a Python package and layering the rest of the functionality on top of it. In terms of the actual functionality of the project, the next step is mutual entropy, or the correlation of columns as a means of obtaining more information (context) for the column itself!
# ## Thank you!
| Demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="Jray-y0klNsT"
import numpy as np
import time
import matplotlib.pyplot as plt
import sys
sys.path.insert(0, 'dummydirectory') #'/content/drive/My Drive/Colab Notebooks/gamenight'
from arena import letsplay
from scoring import score_stich, score_game, did_cheat
from group0 import play as player0
from group1 import play as player1
from group2 import play as player2
from group3 import play as player3
players = [player0, player1, player2, player3]
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="5ClkKgO2wTcQ" outputId="603931e6-9695-49a3-e6a4-09ee2476c9e9"
ncards = 10
toprint = False
score, history = letsplay(players,ncards,toprint,score_stich,score_game,did_cheat);
# -
| Competition.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # DSCI 525 - Web and Cloud Computing
# Milestone 2: Your team is planning to migrate to the cloud. AWS gave 400$ (100$ each) to your team to support this. As part of this initiative, your team needs to set up a server in the cloud, a collaborative environment for your team, and later move your data to the cloud. After that, your team can wrangle the data in preparation for machine learning.
#
# ## Milestone 2 checklist
# You will have mainly 2 tasks. Here is the checklist...
# - To set up a collaborative environment
# - Setup your EC2 instance with JupyterHub.
# - Install all necessary things needed in your UNIX server (amazon ec2 instance).
# - Set up your S3 bucket.
# - Move the data that you wrangled in your last milestone to s3.
# - To move data from s3.
# - Wrangle the data in preparation for machine learning
# - Get the data from S3 in your notebook and make data ready for machine learning.
# **Keep in mind:**
#
# - _All services you use are in region us-west-2._
#
# - _Don't store anything in these servers or storage that represents your identity as a student (like your student ID number) ._
#
# - _Use only default VPC and subnet._
#
# - _No IP addresses are visible when you provide the screenshot._
#
# - _You do proper budgeting so that you don't run out of credits._
#
# - _We want one single notebook for grading, and it's up to your discretion on how you do it. ***So only one person in your group needs to spin up a big instance and a ```t2.xlarge``` is of decent size.***_
#
# - _Please stop the instance when not in use. This can save you some bucks, but it's again up to you and how you budget your money. Maybe stop it if you or your team won't use it for the next 5 hours?
#
# - _Your AWS lab will shut down after 3 hours 30 min. When you start it again, your AWS credentials (***access key***,***secret***, and ***session token***) will change, and you want to update your credentials file with the new one. _
#
# - _Say something went wrong and you want to spin up another EC2 instance, then make sure you terminate the previous one._
#
# - _We will be choosing the storage to be ```Delete on Termination```, which means that stored data in your instance will be lost upon termination. Make sure you save any data to S3 and download the notebooks to your laptop so that next time you have your jupyterHub in a different instance, you can upload your notebook there._
#
# _***Outside of Milestone:*** If you are working as an individual just to practice setting up EC2 instances, make sure you select ```t2.large``` instance (not anything bigger than that as it can cost you money). I strongly recommend you spin up your own instance and experiment with the s3 bucket in doing something (there are many things that we learned and practical work from additional instructions and video series) to get comfortable with AWS. But we won't be looking at it for a grading purpose._
#
# ***NOTE:*** Everything you want for this notebook is discussed in lecture 3, lecture 4, and setup instructions.
# ### 1. Setup your EC2 instance
# rubric={correctness:20}
# #### Please attach this screen shots from your group for grading.
# https://github.ubc.ca/mds-2021-22/DSCI_525_web-cloud-comp_students/blob/master/release/milestone2/image/1_result.png
from PIL import Image
Image.open("img/525_m2_1.png")
# ### 2. Setup your JupyterHub
# rubric={correctness:20}
# #### Please attach this screen shots from your group for grading
# I want to see all the group members here in this screenshot https://github.ubc.ca/mds-2021-22/DSCI_525_web-cloud-comp_students/blob/master/release/milestone2/image/2_result.png
Image.open("img/525_m2_2.png")
# ### 3. Setup the server
# rubric={correctness:20}
# 3.1) Add your team members to EC2 instance.
#
# 3.2) Setup a common data folder to download data, and this folder should be accessible by all users in the JupyterHub.
#
# 3.3)(***OPTIONAL***) Setup a sharing notebook environment.
#
# 3.4) Install and configure AWS CLI.
# + [markdown] tags=[]
# #### Please attach this screen shots from your group for grading
#
# Make sure you mask the IP address refer [here](https://www.anysoftwaretools.com/blur-part-picture-mac/).
#
# https://github.ubc.ca/mds-2021-22/DSCI_525_web-cloud-comp_students/blob/master/release/milestone2/image/3_result.png
# -
Image.open("img/525_m2_3.png")
# ### 4. Get the data what we wrangled in our first milestone.
# You have to install the packages that are needed. Refer this TLJH [document]( https://tljh.jupyter.org/en/latest/howto/env/user-environment.html).Refer ```pip``` section.
#
# Don't forget to add option -E. This way, all packages that you install will be available to other users in your JupyterHub.
# These packages you must install and install other packages needed for your wrangling.
#
# sudo -E pip install pandas
# sudo -E pip install pyarrow
# sudo -E pip install s3fs
# As in the last milestone, we looked at getting the data transferred from Python to R, and we have different solutions. Henceforth, I uploaded the parquet file format, which we can use moving forward.
import re
import os
import glob
import zipfile
import requests
from urllib.request import urlretrieve
import json
import pandas as pd
# Rememeber here we gave the folder that we created in Step 3.2 as we made it available for all the users in a group.
# Necessary metadata
article_id = 14226968 # this is the unique identifier of the article on figshare
url = f"https://api.figshare.com/v2/articles/{article_id}"
headers = {"Content-Type": "application/json"}
output_directory = os.path.abspath(os.path.join(os.getcwd(), os.pardir)) + "/data/shared/"
print(output_directory)
response = requests.request("GET", url, headers=headers)
data = json.loads(response.text) # this contains all the articles data, feel free to check it out
files = data["files"] # this is just the data about the files, which is what we want
files
files_to_dl = ["combined_model_data_parti.parquet.zip"] ## Please download the partitioned
for file in files:
if file["name"] in files_to_dl:
os.makedirs(output_directory, exist_ok=True)
urlretrieve(file["download_url"], output_directory + file["name"])
with zipfile.ZipFile(os.path.join(output_directory, "combined_model_data_parti.parquet.zip"), 'r') as f:
f.extractall(output_directory)
# ### 5. Setup your S3 bucket and move data
# rubric={correctness:20}
# 5.1) Create a bucket name should be mds-s3-xxx. Replace xxx with your "groupnumber".
#
# 5.2) Create your first folder called "output".
#
# 5.3) Move the "observed_daily_rainfall_SYD.csv" file from the Milestone1 data folder to your s3 bucket from your local computer.
#
# 5.4) Moving the parquet file we downloaded(combined_model_data_parti.parquet) in step 4 to S3 using the cli what we installed in step 3.4.
# !aws configure set aws_access_key_id "<KEY>"
# !aws configure set aws_secret_access_key "<KEY>"
# !aws configure set aws_session_token "<KEY>
# !aws s3 cp ../data/shared/ "s3://mds-s3-17" --recursive
# !aws s3 cp ../data/raw/observed_daily_rainfall_SYD.csv "s3://mds-s3-17" --recursive
# #### Please attach this screen shots from your group for grading
#
# Make sure it has 3 objects.
#
# https://github.ubc.ca/mds-2021-22/DSCI_525_web-cloud-comp_students/blob/master/release/milestone2/image/4_result.png
# ### 6. Wrangle the data in preparation for machine learning
# rubric={correctness:20}
# Our data currently covers all of NSW, but say that our client wants us to create a machine learning model to predict rainfall over Sydney only. There's a bit of wrangling that needs to be done for that:
# 1. We need to query our data for only the rows that contain information covering Sydney
# 2. We need to wrangle our data into a format suitable for training a machine learning model. That will require pivoting, resampling, grouping, etc.
#
# To train an ML algorithm we need it to look like this:
#
# ||model-1_rainfall|model-2_rainfall|model-3_rainfall|...|observed_rainfall|
# |---|---|---|---|---|---|
# |0|0.12|0.43|0.35|...|0.31|
# |1|1.22|0.91|1.68|...|1.34|
# |2|0.68|0.29|0.41|...|0.57|
# 6.1) Get the data from s3 (```combined_model_data_parti.parquet``` and ```observed_daily_rainfall_SYD.csv```)
#
# 6.2) First query for Sydney data and then drop the lat and lon columns (we don't need them).
# ```
# syd_lat = -33.86
# syd_lon = 151.21
# ```
# Expected shape ```(1150049, 2)```.
#
# 6.3) Save this processed file to s3 for later use:
#
# Save as a csv file ```ml_data_SYD.csv``` to ```s3://mds-s3-xxx/output/```
# expected shape ```(46020,26)``` - This includes all the models as columns and also adding additional column ```Observed``` loaded from ```observed_daily_rainfall_SYD.csv``` from s3.
# +
### Do all your coding here
# -
import pandas as pd
import pyarrow.parquet as pq
aws_credentials ={"key": "<KEY>","secret": "<KEY>", "token": "<KEY>}
df = pd.read_parquet("s3://mds-s3-17/combined_model_data_parti.parquet/", storage_options=aws_credentials)
# Examining the file:
df.head()
# +
# Using the provided coordinates to filter Sydney / observed model:
syd_lat = -33.86
syd_lon = 151.21
df = df[((df["lat_min"] < syd_lat) & (df["lat_max"] > syd_lat))]
df = df[((df["lon_min"] < syd_lon) & (df["lon_max"] > syd_lon))]
df_sydney = df
# Drop the lat and lon columns as instructed:
df_sydney = df_sydney.drop(columns = ["lat_min", "lat_max", "lon_min", "lon_max"])
# df_sydney = df_sydney.reset_index(drop=True).set_index('time')
df_sydney.head()
# -
# Check the shape of Sydney data
df_sydney_ti = df_sydney.reset_index(drop=True).set_index('time')
df_sydney_ti.shape
# Load observed data:
df_observed = pd.read_csv("s3://mds-s3-17/observed_daily_rainfall_SYD.csv", storage_options=aws_credentials)
df_observed["model"] = "observed_rainfall"
df_observed.head()
# +
# combine the two df:
df_sydney = pd.concat([df_sydney, df_observed])
# Set the index properly:
df_sydney["time"] = pd.to_datetime(df_sydney["time"]).dt.date
df_sydney.set_index("time", inplace=True)
df_sydney.head()
# -
# Pivot the new df
df_final = df_sydney.reset_index().pivot(index = 'time', columns = "model", values = "rain (mm/day)")
df_final = df_final.reset_index(drop=True)
df_final.head()
df_final.shape
# save the file
df_final.to_csv("s3://mds-s3-17/output/ml_data_SYD.csv", storage_options=aws_credentials, index=False)
# How the final file format looks like
# https://github.ubc.ca/mds-2021-22/DSCI_525_web-cloud-comp_students/blob/master/release/milestone2/image/finaloutput.png
#
# Shape ```(46020,26 )```
# (***OPTIONAL***) If you are interested in doing some benchmarking!! How much time it took to read..
# - Parquet file from your local disk ?
# - Parquet file from s3 ?
# - CSV file from s3 ?
# For that, upload the CSV file (```combined_model_data.csv```
# )to S3 and try to read it instead of parquet.
| notebook/.ipynb_checkpoints/Milestone2-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import csv
import pickle
import datetime
from sklearn.model_selection import train_test_split
# %matplotlib inline
# %config InlineBackend.figure_formats = {'png','retina'}
def import_data():
df = pd.read_csv('data/final_df.csv')
df = df.iloc[:,1:]
return df
# +
# data = pd.read_csv('data/data.csv')
# data=data.iloc[:,1:]
# +
# new_order = ['date', 'store_nbr', 'item_nbr', 'station_nbr', 'tmax', 'tmin',
# 'tavg', 'depart', 'dewpoint', 'wetbulb', 'heat', 'cool', 'sunrise',
# 'sunset', 'snowfall', 'preciptotal', 'stnpressure', 'sealevel',
# 'avgspeed', 'resultspeed', 'resultdir', 'TS', 'GR', 'RA', 'DZ', 'SN',
# 'SG', 'GS', 'PL', 'FG+', 'FG', 'BR', 'UP', 'HZ', 'FU', 'DU', 'SQ', 'FZ',
# 'MI', 'PR', 'BC', 'BL', 'VC', 'units']
# +
# data = data[new_order]
# +
# data.columns = ['date', 'store_nbr', 'item_nbr', 'station_nbr', 'tmax', 'tmin', 'tavg',
# 'depart', 'dewpoint', 'wetbulb', 'heat', 'cool', 'sunrise', 'sunset',
# 'snowfall', 'preciptotal', 'stnpressure', 'sealevel', 'avgspeed',
# 'resultspeed', 'resultdir', 'TS', 'GR', 'RA', 'DZ', 'SN', 'SG', 'GS',
# 'PL', 'FG2', 'FG', 'BR', 'UP', 'HZ', 'FU', 'DU', 'SQ', 'FZ', 'MI', 'PR',
# 'BC', 'BL', 'VC', 'units']
# +
def fulldf(df):
df.snowfall = df.snowfall.replace(["T"," T"],0.05)
df.preciptotal = df.preciptotal.replace(["T"," T"],0.005)
df.depart = pd.to_numeric(df.depart, errors='coerce')
df = type_change_numeric(df,[ 'store_nbr', 'item_nbr', 'units', 'station_nbr', 'tmax', 'tmin',
'tavg', 'depart', 'dewpoint', 'wetbulb', 'heat', 'cool', 'snowfall', 'preciptotal', 'stnpressure', 'sealevel',
'avgspeed', 'resultspeed', 'resultdir' ])
df['date'] = pd.to_datetime(df['date'])
df["day_of_year"] = df['date'].dt.dayofyear
df["year"] = df['date'].dt.year
df["month"] = df["date"].dt.month
for idx in range(5, 8):
df.iloc[:,idx].fillna(df.groupby(["day_of_year","store_nbr"])[df.columns[idx]].\
transform('mean'), inplace=True)
for idx in range(16, 23):
df.iloc[:,idx].fillna(df.groupby(["day_of_year","store_nbr"])[df.columns[idx]].\
transform('mean'), inplace=True)
add_depart1(df)
return df
# for column in item37.columns:
# item37[column].interpolate()
# -
def type_change_numeric(df, ls = []):
#ls에 있는 column name은 numeric형으로 바꾸지 않는다.
cols = df.columns
for i in cols:
if i in ls:
#df = df.replace(["M",None], '')
df.snowfall = df.snowfall.replace(["T"," T"],0.05)
df.preciptotal = df.preciptotal.replace(["T"," T"],0.005)
df[i] = pd.to_numeric(df[i], errors='coerce')
return df
def add_depart1(x):
x.depart.fillna(x.tavg - x.groupby(["day_of_year","store_nbr"])["tavg"].transform('mean'),inplace = True)
x.depart = x.depart.round(2)
return x.sort_values(["store_nbr","date","item_nbr"])
def reorder_df(df):
#Column 정렬 (y값을 마지막으로 ) 후 FG+ -> FG2 변환 (formula에 인식시키기위해 )
new_order = ['date', 'store_nbr', 'item_nbr', 'station_nbr', 'tmax', 'tmin',
'tavg', 'depart', 'dewpoint', 'wetbulb', 'heat', 'cool', 'snowfall', 'preciptotal', 'stnpressure', 'sealevel',
'avgspeed', 'resultspeed', 'TS', 'GR', 'RA', 'DZ', 'SN',
'SG', 'GS', 'PL', 'FG+', 'FG', 'BR', 'UP', 'HZ', 'FU', 'DU', 'SQ', 'FZ',
'MI', 'PR', 'BC', 'BL', 'VC', 'day_of_year', 'year', 'month', 'units' ]
df = df[new_order]
df.columns = ['date', 'store_nbr', 'item_nbr', 'station_nbr', 'tmax', 'tmin',
'tavg', 'depart', 'dewpoint', 'wetbulb', 'heat', 'cool', 'snowfall', 'preciptotal', 'stnpressure', 'sealevel',
'avgspeed', 'resultspeed', 'TS', 'GR', 'RA', 'DZ', 'SN',
'SG', 'GS', 'PL', 'FG2', 'FG', 'BR', 'UP', 'HZ', 'FU', 'DU', 'SQ', 'FZ',
'MI', 'PR', 'BC', 'BL', 'VC', 'day_of_year', 'year', 'month', 'units' ]
return df
def add_cat_columns(df):
df['date'] = pd.to_datetime(df['date'])
# df['sunrise'] = pd.to_datetime(df['sunrise'], format='%H%M')
# df['sunset'] = pd.to_datetime(df['sunset'], format='%H%M')
blackfriday = ["2012-11-21","2012-11-22","2012-11-23", "2012-11-24","2012-11-25",
"2013-11-27","2013-11-28", "2013-11-29","2013-11-30","2013-11-31",
"2014-11-26", "2014-11-27", "2014-11-28","2014-11-29","2014-11-30"]
df["week_day_name"] = df['date'].dt.weekday_name
df['weekend'] = False
df.weekend[df['week_day_name'] == 'Sunday'] = True
df.weekend[df['week_day_name'] == 'Saturday'] = True
df.weekend[df['week_day_name'] == 'Friday'] = True
df["is_blackfriday"] = df.date.apply(lambda x : str(x)[:10] in blackfriday).astype(int)
df['hardrain'] = [(((4 if i > 8 else 3) if i > 6 else 2) if i > 1 else 1) if i > 0 else 0 for i in df['preciptotal']]
# rain 1 snow 2
# 득정조건열 추가 ()
df['hardsnow'] = [(( 3 if i > 3.5 else 2) if i > 1 else 1) if i > 0 else 0 for i in df['snowfall']]
df['log_units'] = df.units.apply(lambda x: np.log(x + 1)).astype(float)
return df
def df_sampling(df):
new_order = ['date', 'store_nbr', 'item_nbr', 'station_nbr', 'tmax', 'tmin', 'tavg',
'depart', 'dewpoint', 'wetbulb', 'heat', 'cool', 'preciptotal', 'stnpressure', 'sealevel', 'avgspeed',
'resultspeed', 'TS', 'GR', 'RA', 'DZ', 'SN', 'SG', 'GS',
'PL', 'FG2', 'FG', 'BR', 'UP', 'HZ', 'FU', 'DU', 'SQ', 'FZ', 'MI', 'PR',
'BC', 'BL', 'VC', 'day_of_year', 'year', 'month',
'week_day_name', 'weekend', 'is_blackfriday', 'hardrain', 'hardsnow',
'log_units', 'units' ]
df = df[new_order]
X, y = df.iloc[:,:-1], df.units
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.3, random_state = 2018)
train = pd.concat([X_train, y_train] ,axis = 1)
train = train.sort_values(by=['date', 'store_nbr', 'item_nbr']).reset_index(drop= True)
X_test = X_test.sort_index()
y_train = y_train.sort_index()
y_test = y_test.sort_index()
return train, y_train, X_test ,y_test
data = import_data()
data_t = fulldf(data)
data_t = reorder_df(data_t)
data_t = add_cat_columns(data_t)
del data_t['snowfall']
data_t = data_t.dropna()
train0, y_train, X_test, y_test = df_sampling(data_t)
train = sm.add_constant(train0)
# +
# X, y = data_t.iloc[:,:-1], data_t.units
# X_train, X_testm, y_train, y_test = train_test_split(X, y, test_size = 0.3, random_state = 2018)
# train = pd.concat([X_train, y_train] ,axis = 1)
# train = train.sort_values(by=['date', 'store_nbr', 'item_nbr']).reset_index(drop= True)
# +
# X_train['date'] = pd.to_datetime(X_train['date'])
# X_train['sunrise'] = pd.to_datetime(X_train['sunrise'], format='%H%M')
# X_train['sunset'] = pd.to_datetime(X_train['sunset'], format='%H%M')
# X_testm['date'] = pd.to_datetime(X_testm['date'])
# X_testm['sunrise'] = pd.to_datetime(X_testm['sunrise'], format='%H%M')
# X_testm['sunset'] = pd.to_datetime(X_testm['sunset'], format='%H%M')
# +
# train['log_units'] = train.units.apply(lambda x: np.log(x + 1))
# +
# train['month'] =train["date"].dt.month
# train['weekend'] = False
# train.weekend[train['week_day_name'] == 'Sunday'] = True
# train.weekend[train['week_day_name'] == 'Saturday'] = True
# train.weekend[train['week_day_name'] == 'Friday'] = True
# train["is_blackfriday"] = train.date.apply(lambda x : str(x)[:10] in blackfriday).astype(int)
# +
# blackfriday = ["2012-11-21","2012-11-22","2012-11-23", "2012-11-24","2012-11-25",
# "2013-11-27","2013-11-28", "2013-11-29","2013-11-30","2013-11-31",
# "2014-11-26", "2014-11-27", "2014-11-28","2014-11-29","2014-11-30"]
# +
# train[train > 3.5].preciptotal.hist(bins = 20)
# +
# train['hardrain'] = [(((4 if i > 8 else 3) if i > 6 else 2) if i > 1 else 1) if i > 0 else 0 for i in train['preciptotal']]
# # rain 1 snow 2
# # 득정조건열 추가 ()
# train['hardsnow'] = [(( 3 if i > 3.5 else 2) if i > 1 else 1) if i > 0 else 0 for i in train['snowfall']]
# +
# del train['sunrise']
# del train['sunset']
# del train['snowfall']
# train = train.dropna()
# -
model_OLS = sm.OLS.from_formula("I(np.log(units+1)**3) ~ C(store_nbr) + C(item_nbr) + scale(preciptotal) + C(weekend) + C(month) + C(year) + C(is_blackfriday) + 0" , data = train)
results_OLS = model_OLS.fit()
print(results_OLS.summary())
sns.jointplot(results_OLS.predict(X_test), np.log(y_test + 1))
plt.show()
model_OLS2 = sm.OLS.from_formula("log_units ~ C(store_nbr) + C(item_nbr) + scale(preciptotal) + C(weekend) + C(month) + C(year) + C(is_blackfriday) + 0" , data = train)
results_OLS2 = model_OLS2.fit()
print(results_OLS2.summary())
| team_project/OLS.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Training a better model
from theano.sandbox import cuda
# %matplotlib inline
from imp import reload
import utils; reload(utils)
from utils import *
from __future__ import division, print_function
# +
#path = "data/dogscats/sample/"
path = "/usr/local/share/dsLab/datasets/dogscats/sample/"
model_path = path + 'models/'
if not os.path.exists(model_path): os.mkdir(model_path)
batch_size=64
# + [markdown] heading_collapsed=true
# ## Are we underfitting?
# + [markdown] hidden=true
# Our validation accuracy so far has generally been higher than our training accuracy. That leads to two obvious questions:
#
# 1. How is this possible?
# 2. Is this desirable?
#
# The answer to (1) is that this is happening because of *dropout*. Dropout refers to a layer that randomly deletes (i.e. sets to zero) each activation in the previous layer with probability *p* (generally 0.5). This only happens during training, not when calculating the accuracy on the validation set, which is why the validation set can show higher accuracy than the training set.
#
# The purpose of dropout is to avoid overfitting. By deleting parts of the neural network at random during training, it ensures that no one part of the network can overfit to one part of the training set. The creation of dropout was one of the key developments in deep learning, and has allowed us to create rich models without overfitting. However, it can also result in underfitting if overused, and this is something we should be careful of with our model.
#
# So the answer to (2) is: this is probably not desirable. It is likely that we can get better validation set results with less (or no) dropout, if we're seeing that validation accuracy is higher than training accuracy - a strong sign of underfitting. So let's try removing dropout entirely, and see what happens!
#
# (We had dropout in this model already because the VGG authors found it necessary for the imagenet competition. But that doesn't mean it's necessary for dogs v cats, so we will do our own analysis of regularization approaches from scratch.)
# -
# ## Removing dropout
# Our high level approach here will be to start with our fine-tuned cats vs dogs model (with dropout), then fine-tune all the dense layers, after removing dropout from them. The steps we will take are:
# - Re-create and load our modified VGG model with binary dependent (i.e. dogs v cats)
# - Split the model between the convolutional (*conv*) layers and the dense layers
# - Pre-calculate the output of the conv layers, so that we don't have to redundently re-calculate them on every epoch
# - Create a new model with just the dense layers, and dropout p set to zero
# - Train this new model using the output of the conv layers as training data.
# As before we need to start with a working model, so let's bring in our working VGG 16 model and change it to predict our binary dependent...
model = vgg_ft(2)
# ...and load our fine-tuned weights.
model.load_weights(model_path+'finetune3.h5')
# We're going to be training a number of iterations without dropout, so it would be best for us to pre-calculate the input to the fully connected layers - i.e. the *Flatten()* layer. We'll start by finding this layer in our model, and creating a new model that contains just the layers up to and including this layer:
layers = model.layers
last_conv_idx = [index for index,layer in enumerate(layers)
if type(layer) is Convolution2D][-1]
last_conv_idx
layers[last_conv_idx]
conv_layers = layers[:last_conv_idx+1]
conv_model = Sequential(conv_layers)
# Dense layers - also known as fully connected or 'FC' layers
fc_layers = layers[last_conv_idx+1:]
# Now we can use the exact same approach to creating features as we used when we created the linear model from the imagenet predictions in the last lesson - it's only the model that has changed. As you're seeing, there's a fairly small number of "recipes" that can get us a long way!
# +
batches = get_batches(path+'train', shuffle=False, batch_size=batch_size)
val_batches = get_batches(path+'valid', shuffle=False, batch_size=batch_size)
val_classes = val_batches.classes
trn_classes = batches.classes
val_labels = onehot(val_classes)
trn_labels = onehot(trn_classes)
# -
batches.class_indices
val_features = conv_model.predict_generator(val_batches, val_batches.nb_sample)
trn_features = conv_model.predict_generator(batches, batches.nb_sample)
save_array(model_path + 'train_convlayer_features.bc', trn_features)
save_array(model_path + 'valid_convlayer_features.bc', val_features)
trn_features = load_array(model_path+'train_convlayer_features.bc')
val_features = load_array(model_path+'valid_convlayer_features.bc')
trn_features.shape
# For our new fully connected model, we'll create it using the exact same architecture as the last layers of VGG 16, so that we can conveniently copy pre-trained weights over from that model. However, we'll set the dropout layer's p values to zero, so as to effectively remove dropout.
# Copy the weights from the pre-trained model.
# NB: Since we're removing dropout, we want to half the weights
def proc_wgts(layer): return [o/2 for o in layer.get_weights()]
# Such a finely tuned model needs to be updated very slowly!
opt = RMSprop(lr=0.00001, rho=0.7)
def get_fc_model():
model = Sequential([
MaxPooling2D(input_shape=conv_layers[-1].output_shape[1:]),
Flatten(),
Dense(4096, activation='relu'),
Dropout(0.),
Dense(4096, activation='relu'),
Dropout(0.),
Dense(2, activation='softmax')
])
for l1,l2 in zip(model.layers, fc_layers): l1.set_weights(proc_wgts(l2))
model.compile(optimizer=opt, loss='categorical_crossentropy', metrics=['accuracy'])
return model
fc_model = get_fc_model()
# And fit the model in the usual way:
fc_model.fit(trn_features, trn_labels, nb_epoch=8,
batch_size=batch_size, validation_data=(val_features, val_labels))
fc_model.save_weights(model_path+'no_dropout.h5')
fc_model.load_weights(model_path+'no_dropout.h5')
# # Reducing overfitting
# Now that we've gotten the model to overfit, we can take a number of steps to reduce this.
# + [markdown] heading_collapsed=true
# ## Approaches to reducing overfitting
# + [markdown] hidden=true
# We do not necessarily need to rely on dropout or other regularization approaches to reduce overfitting. There are other techniques we should try first, since regularlization, by definition, biases our model towards simplicity - which we only want to do if we know that's necessary. This is the order that we recommend using for reducing overfitting (more details about each in a moment):
#
# 1. Add more data
# 2. Use data augmentation
# 3. Use architectures that generalize well
# 4. Add regularization
# 5. Reduce architecture complexity.
#
# We'll assume that you've already collected as much data as you can, so step (1) isn't relevant (this is true for most Kaggle competitions, for instance). So the next step (2) is data augmentation. This refers to creating additional synthetic data, based on reasonable modifications of your input data. For images, this is likely to involve one or more of: flipping, rotation, zooming, cropping, panning, minor color changes.
#
# Which types of augmentation are appropriate depends on your data. For regular photos, for instance, you'll want to use horizontal flipping, but not vertical flipping (since an upside down car is much less common than a car the right way up, for instance!)
#
# We recommend *always* using at least some light data augmentation, unless you have so much data that your model will never see the same input twice.
# -
# ## About data augmentation
# Keras comes with very convenient features for automating data augmentation. You simply define what types and maximum amounts of augmentation you want, and keras ensures that every item of every batch randomly is changed according to these settings. Here's how to define a generator that includes data augmentation:
# dim_ordering='tf' uses tensorflow dimension ordering,
# which is the same order as matplotlib uses for display.
# Therefore when just using for display purposes, this is more convenient
gen = image.ImageDataGenerator(rotation_range=10, width_shift_range=0.1,
height_shift_range=0.1, shear_range=0.15, zoom_range=0.1,
channel_shift_range=10., horizontal_flip=True, dim_ordering='tf')
# Let's take a look at how this generator changes a single image (the details of this code don't matter much, but feel free to read the comments and keras docs to understand the details if you're interested).
# Create a 'batch' of a single image
img = np.expand_dims(ndimage.imread('data/dogscats/test/7.jpg'),0)
# Request the generator to create batches from this image
aug_iter = gen.flow(img)
# Get eight examples of these augmented images
aug_imgs = [next(aug_iter)[0].astype(np.uint8) for i in range(8)]
# The original
plt.imshow(img[0])
# As you can see below, there's no magic to data augmentation - it's a very intuitive approach to generating richer input data. Generally speaking, your intuition should be a good guide to appropriate data augmentation, although it's a good idea to test your intuition by checking the results of different augmentation approaches.
# Augmented data
plots(aug_imgs, (20,7), 2)
# Ensure that we return to theano dimension ordering
K.set_image_dim_ordering('th')
# + [markdown] heading_collapsed=true
# ## Adding data augmentation
# + [markdown] hidden=true
# Let's try adding a small amount of data augmentation, and see if we reduce overfitting as a result. The approach will be identical to the method we used to finetune the dense layers in lesson 2, except that we will use a generator with augmentation configured. Here's how we set up the generator, and create batches from it:
# + hidden=true
gen = image.ImageDataGenerator(rotation_range=15, width_shift_range=0.1,
height_shift_range=0.1, zoom_range=0.1, horizontal_flip=True)
# + hidden=true
batches = get_batches(path+'train', gen, batch_size=batch_size)
# NB: We don't want to augment or shuffle the validation set
val_batches = get_batches(path+'valid', shuffle=False, batch_size=batch_size)
# + [markdown] hidden=true
# When using data augmentation, we can't pre-compute our convolutional layer features, since randomized changes are being made to every input image. That is, even if the training process sees the same image multiple times, each time it will have undergone different data augmentation, so the results of the convolutional layers will be different.
#
# Therefore, in order to allow data to flow through all the conv layers and our new dense layers, we attach our fully connected model to the convolutional model--after ensuring that the convolutional layers are not trainable:
# + hidden=true
fc_model = get_fc_model()
# + hidden=true
for layer in conv_model.layers: layer.trainable = False
# Look how easy it is to connect two models together!
conv_model.add(fc_model)
# + [markdown] hidden=true
# Now we can compile, train, and save our model as usual - note that we use *fit_generator()* since we want to pull random images from the directories on every batch.
# + hidden=true
conv_model.compile(optimizer=opt, loss='categorical_crossentropy', metrics=['accuracy'])
# + hidden=true
conv_model.fit_generator(batches, samples_per_epoch=batches.nb_sample, nb_epoch=8,
validation_data=val_batches, nb_val_samples=val_batches.nb_sample)
# + hidden=true
conv_model.fit_generator(batches, samples_per_epoch=batches.nb_sample, nb_epoch=3,
validation_data=val_batches, nb_val_samples=val_batches.nb_sample)
# + hidden=true
conv_model.save_weights(model_path + 'aug1.h5')
# + hidden=true
conv_model.load_weights(model_path + 'aug1.h5')
# -
# ## Batch normalization
# + [markdown] heading_collapsed=true
# ### About batch normalization
# + [markdown] hidden=true
# Batch normalization (*batchnorm*) is a way to ensure that activations don't become too high or too low at any point in the model. Adjusting activations so they are of similar scales is called *normalization*. Normalization is very helpful for fast training - if some activations are very high, they will saturate the model and create very large gradients, causing training to fail; if very low, they will cause training to proceed very slowly. Furthermore, large or small activations in one layer will tend to result in even larger or smaller activations in later layers, since the activations get multiplied repeatedly across the layers.
#
# Prior to the development of batchnorm in 2015, only the inputs to a model could be effectively normalized - by simply subtracting their mean and dividing by their standard deviation. However, weights in intermediate layers could easily become poorly scaled, due to problems in weight initialization, or a high learning rate combined with random fluctuations in weights.
# + [markdown] hidden=true
# Batchnorm resolves this problem by normalizing each intermediate layer as well. The details of how it works are not terribly important (although I will outline them in a moment) - the important takeaway is that **all modern networks should use batchnorm, or something equivalent**. There are two reasons for this:
# 1. Adding batchnorm to a model can result in **10x or more improvements in training speed**
# 2. Because normalization greatly reduces the ability of a small number of outlying inputs to over-influence the training, it also tends to **reduce overfitting**.
# + [markdown] hidden=true
# As promised, here's a brief outline of how batchnorm works. As a first step, it normalizes intermediate layers in the same way as input layers can be normalized. But this on its own would not be enough, since the model would then just push the weights up or down indefinitely to try to undo this normalization. Therefore, batchnorm takes two additional steps:
# 1. Add two more trainable parameters to each layer - one to multiply all activations to set an arbitrary standard deviation, and one to add to all activations to set an arbitary mean
# 2. Incorporate both the normalization, and the learnt multiply/add parameters, into the gradient calculations during backprop.
#
# This ensures that the weights don't tend to push very high or very low (since the normalization is included in the gradient calculations, so the updates are aware of the normalization). But it also ensures that if a layer does need to change the overall mean or standard deviation in order to match the output scale, it can do so.
# + [markdown] heading_collapsed=true
# ### Adding batchnorm to the model
# + [markdown] hidden=true
# We can use nearly the same approach as before - but this time we'll add batchnorm layers (and dropout layers):
# + hidden=true
conv_layers[-1].output_shape[1:]
# + hidden=true
def get_bn_layers(p):
return [
MaxPooling2D(input_shape=conv_layers[-1].output_shape[1:]),
Flatten(),
Dense(4096, activation='relu'),
BatchNormalization(),
Dropout(p),
Dense(4096, activation='relu'),
BatchNormalization(),
Dropout(p),
Dense(1000, activation='softmax')
]
# -
def load_fc_weights_from_vgg16bn(model):
"Load weights for model from the dense layers of the Vgg16BN model."
# See imagenet_batchnorm.ipynb for info on how the weights for
# Vgg16BN can be generated from the standard Vgg16 weights.
from vgg16bn import Vgg16BN
vgg16_bn = Vgg16BN()
_, fc_layers = split_at(vgg16_bn.model, Convolution2D)
copy_weights(fc_layers, model.layers)
# + hidden=true
p=0.6
# + hidden=true
bn_model = Sequential(get_bn_layers(0.6))
# + hidden=true
load_fc_weights_from_vgg16bn(bn_model)
# + hidden=true
def proc_wgts(layer, prev_p, new_p):
scal = (1-prev_p)/(1-new_p)
return [o*scal for o in layer.get_weights()]
# + hidden=true
for l in bn_model.layers:
if type(l)==Dense: l.set_weights(proc_wgts(l, 0.5, 0.6))
# + hidden=true
bn_model.pop()
for layer in bn_model.layers: layer.trainable=False
# + hidden=true
bn_model.add(Dense(2,activation='softmax'))
# + hidden=true
bn_model.compile(Adam(), 'categorical_crossentropy', metrics=['accuracy'])
# + hidden=true
bn_model.fit(trn_features, trn_labels, nb_epoch=8, validation_data=(val_features, val_labels))
# + hidden=true
bn_model.save_weights(model_path+'bn.h5')
# + hidden=true
bn_model.load_weights(model_path+'bn.h5')
# + hidden=true
bn_layers = get_bn_layers(0.6)
bn_layers.pop()
bn_layers.append(Dense(2,activation='softmax'))
# + hidden=true
final_model = Sequential(conv_layers)
for layer in final_model.layers: layer.trainable = False
for layer in bn_layers: final_model.add(layer)
# + hidden=true
for l1,l2 in zip(bn_model.layers, bn_layers):
l2.set_weights(l1.get_weights())
# + hidden=true
final_model.compile(optimizer=Adam(),
loss='categorical_crossentropy', metrics=['accuracy'])
# + hidden=true
final_model.fit_generator(batches, samples_per_epoch=batches.nb_sample, nb_epoch=1,
validation_data=val_batches, nb_val_samples=val_batches.nb_sample)
# + hidden=true
final_model.save_weights(model_path + 'final1.h5')
# + hidden=true
final_model.fit_generator(batches, samples_per_epoch=batches.nb_sample, nb_epoch=4,
validation_data=val_batches, nb_val_samples=val_batches.nb_sample)
# + hidden=true
final_model.save_weights(model_path + 'final2.h5')
# + hidden=true
final_model.optimizer.lr=0.001
# + hidden=true
final_model.fit_generator(batches, samples_per_epoch=batches.nb_sample, nb_epoch=4,
validation_data=val_batches, nb_val_samples=val_batches.nb_sample)
# + hidden=true
bn_model.save_weights(model_path + 'final3.h5')
# + hidden=true
| JupyterNotebooks/Training/FastAI/DeepLearning1/lesson3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (matenv)
# language: python
# name: matenv
# ---
# # Predicting the refractive index for the Materials Project
#
# This notebook will predict the refractive index for ~68,000 materials from the Materials Project (MP).
# This is done by using the pretrained model "refractive_index" available in the following directory: modnet/pretrained.
#
# ## Load model
# First the MODNetModel is loaded:
import sys
#sys.path.append('..')
from modnet.models import MODNetModel
from modnet.preprocessing import MODData
model = MODNetModel.load('../pretrained/refractive_index')
# ## Load MODData
# In order to predict it on new structures a MODData should be created. Here we want to predict on structures from the MP, and therefore load the precomputed MP MODData.
# This takes some time and memory.
MP_data = MODData.load("../moddata/MP_2018.6")
# ## Predict on new structures
# By using the predict() method on the MP_data, a dataframe containing the predictions is obtained.
df = model.predict(MP_data)
df
# As a test, 3 materials with id mp-19033, mp-559175, and mp-6930, not included in the training set, were computed by DFT with corresponding 'true' refractive index: 2.59, 2.53 and 1.56.
#
# The following approximation was found by our model, in a fraction of time:
df.loc[['mp-19033', 'mp-559175', 'mp-6930']]
| example_notebooks/predicting_ref_index.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:py27]
# language: python
# name: conda-env-py27-py
# ---
# # KNN & DTW
# +
# -*- coding: utf-8 -*-
class Dtw(object):
def __init__(self, seq1, seq2,
patterns = [(-1,-1), (-1,0), (0,-1)],
weights = [{(0,0):2}, {(0,0):1}, {(0,0):1}],
band_r=0.05):
self._seq1 = seq1
self._seq2 = seq2
self.len_seq1 = len(seq1)
self.len_seq2 = len(seq2)
self.len_pattern = len(patterns)
self.sum_w = [sum(ws.values()) for ws in weights]
self._r = int(len(seq1)*band_r)
assert len(patterns) == len(weights)
self._patterns = patterns
self._weights = weights
def get_distance(self, i1, i2):
return abs(self._seq1[i1] - self._seq2[i2])
def calculate(self):
g = list([float('inf')]*self.len_seq2 for i in range(self.len_seq1))
cost = list([0]*self.len_seq2 for i in range(self.len_seq1))
g[0][0] = 2*self.get_distance(0, 0)
for i in range(self.len_seq1):
for j in range(max(0,i-self._r), min(i+self._r+1, self.len_seq2)):
for pat_i in range(self.len_pattern):
coor = (i+self._patterns[pat_i][0], j+self._patterns[pat_i][1])
if coor[0]<0 or coor[1]<0:
continue
dist = 0
for w_coor_offset, d_w in self._weights[pat_i].items():
w_coor = (i+w_coor_offset[0], j+w_coor_offset[1])
dist += d_w*self.get_distance(w_coor[0], w_coor[1])
this_val = g[coor[0]][coor[1]] + dist
this_cost = cost[coor[0]][coor[1]] + self.sum_w[pat_i]
if this_val < g[i][j]:
g[i][j] = this_val
cost[i][j] = this_cost
return g[self.len_seq1-1][self.len_seq2-1]/cost[self.len_seq1-1][self.len_seq2-1], g, cost
def print_table(self, tb):
print(' '+' '.join(["{:^7d}".format(i) for i in range(self.len_seq2)]))
for i in range(self.len_seq1):
str = "{:^4d}: ".format(i)
for j in range(self.len_seq2):
str += "{:^7.3f} ".format(tb[i][j])
print (str)
def print_g_matrix(self):
_, tb, _ = self.calculate()
self.print_table(tb)
def print_cost_matrix(self):
_, _, tb = self.calculate()
self.print_table(tb)
def get_dtw(self):
ans, _, _ = self.calculate()
return ans
# +
import csv
import random
import math
import operator
import numpy as np
def loadDataset(filename, data=[]):
with open(filename, 'rb') as csvfile:
lines = csv.reader(csvfile,delimiter=' ')
dataset = list(lines)
for x in range(len(dataset)):
dataset[x] = filter(None, dataset[x])
dataset[x] = list(map(float, dataset[x]))
data.append(dataset[x])
def euclideanDistance(instance1, instance2, length):
distance = 0
for x in range(length):
if x == 0:
continue
distance += pow((instance1[x] - instance2[x]), 2)
return math.sqrt(distance)
def getNeighbors(trainingSet, testInstance, k, pattern, weight):
distances = []
length = len(testInstance)
for x in range(len(trainingSet)):
# z-normalization
new_testInstance = (np.array(testInstance)-np.mean(testInstance))/np.std(testInstance)
new_trainingSet = (np.array(trainingSet[x])-np.mean(trainingSet[x]))/np.std(trainingSet[x])
d = Dtw(testInstance[1:], trainingSet[1:], pattern, weight)
dist = d.get_dtw()
# dist = euclideanDistance(testInstance, trainingSet[x], length)
distances.append((trainingSet[x], dist))
distances.sort(key=operator.itemgetter(1))
# print "dist >>>> ",distances
neighbors = []
for x in range(k):
neighbors.append(distances[x][0])
return neighbors
def getResponse(neighbors):
classVotes = {}
for x in range(len(neighbors)):
response = neighbors[x][0]
if response in classVotes:
classVotes[response] += 1
else:
classVotes[response] = 1
sortedVotes = sorted(classVotes.iteritems(), key=operator.itemgetter(1), reverse=True)
return sortedVotes[0][0]
def getAccuracy(testSet, predictions):
correct = 0
for x in range(len(testSet)):
if testSet[x][0] == predictions[x]:
correct += 1
return (correct/float(len(testSet))) * 100.0
def knn(train_data, test_data, k, pattern, weight):
# prepare data
trainingSet=[]
testSet=[]
loadDataset(train_data, trainingSet)
loadDataset(test_data, testSet)
# print 'Train set: ' + repr(len(trainingSet))
# print trainingSet
# print 'Test set: ' + repr(len(testSet))
# print testSet
# generate predictions
predictions=[]
for x in range(len(testSet)):
# print ">>",testSet[x]
neighbors = getNeighbors(trainingSet, testSet[x], k, pattern, weight)
# print "neighbors >>", neighbors
result = getResponse(neighbors)
# print "result >>", result
predictions.append(result)
# print('> predicted=' + repr(result) + ', actual=' + repr(testSet[x][0]))
accuracy = getAccuracy(testSet, predictions)
return accuracy
# -
# # Main
PATTERNS_1 = [(0,-1), (-1,-1), (-1,0)]
WEIGHTS_SYM_1 = [{(0,0):1}, {(0,0):2}, {(0,0):1}]
COUNT = 10
weights = []
for i in range(COUNT+1):
for j in range(COUNT-i+1):
k = COUNT - j - i
weights.append([{(0,0):i}, {(0,0):j}, {(0,0):k}])
TRAIN_DATA = 'dataset/Coffee_TRAIN'
TEST_DATA = 'dataset/Coffee_TEST'
OUTPUT_FILE = 'COUNT10_acc_coffee_0.01band.csv'
knn(TRAIN_DATA, TEST_DATA, 1, PATTERNS_1, WEIGHTS_SYM_1)
with open(OUTPUT_FILE, "w") as myfile:
myfile.write("i,j,k,accuracy\n")
for weight in weights:
i = weight[0][(0,0)]
j = weight[1][(0,0)]
k = weight[2][(0,0)]
print "i:", i, "j:", j,"k:", k
acc = knn(TRAIN_DATA, TEST_DATA, 1, PATTERNS_1, weight)
print acc
with open(OUTPUT_FILE, "a") as myfile:
myfile.write(str(i)+","+str(j)+","+str(k)+","+str(acc)+"\n")
| knn_Coffee_0.1band_COUNT10.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# In this notebook, we explore the GOLEM tokamak database using the _pygolem_ Python module
# Python 3
import configparser as cp
import numpy as np
import matplotlib.pyplot as plt
from urllib.request import urlopen, HTTPError
# %matplotlib inline
def get_shot_config(shot):
"""
Get the GOLEM shot configuration.
Parameters
----------
shot : int
GOLEM shot number
Returns
-------
dict
GOLEM shot configuration dictionnary
"""
url = 'http://golem.fjfi.cvut.cz/shots/{}/data_configuration.cfg'.format(shot)
try:
with urlopen(url) as response:
config_str = response.read().decode('utf-8')
config = cp.RawConfigParser()
data_types = dict()
config.read_string(config_str)
for data_type in config.sections():
data_types[data_type] = dict(config.items(data_type))
return data_types
except HTTPError:
print('Problem with the network? Can''t open the config file')
return None
def get_shot_data_dict(shot, signame):
"""
Returns the data dictionnary of a signal for given shot.
"""
baseURL = "http://golem.fjfi.cvut.cz/utils/data/"
url = baseURL + str(shot) +'/' + signame + '.npz'
print('Openning {} ...'.format(url))
# The source file gets downloaded and saved into a temporary directory
ds = np.DataSource()
return np.load(ds.open(url, mode='br'))# as npz: # Python 3 needs to open the file in binary mode
def dict_to_y(data_dict):
y = data_dict['data']
t = np.linspace(data_dict['t_start'], data_dict['t_end'], len(y))
return t, y
cfg = get_shot_config(22668)
irog = get_shot_data_dict(22668, 'irog')
t, _irog = dict_to_y(irog)
plt.plot(t, _irog)
# # Basic Parameters
shot = 22667
# Gas Pressure
pressure = get_shot_data_dict(shot, 'pressure')['data'] # mPa
print(pressure)
# Gas Specie
gas = get_shot_data_dict(shot, 'working_gas')['data']
print(gas)
# Plasma lifetime
is_plasma = get_shot_data_dict(shot, 'plasma')['data'] # 1 or 0
t_plasma = get_shot_data_dict(shot, 'plasma_life')['data']
ub, ubd, ucd, ust = get_shot_data_dict(shot, 'ub')['data'], get_shot_data_dict(shot, 'ubd')['data'], get_shot_data_dict(shot, 'ucd')['data'], get_shot_data_dict(shot, 'ust')['data']
tb, tbd, tcd, tst = get_shot_data_dict(shot, 'tb')['data'], get_shot_data_dict(shot, 'tbd')['data'], get_shot_data_dict(shot, 'tcd')['data'], get_shot_data_dict(shot, 'tst')['data'],
print(pressure, gas, t_plasma)
print(ub, ubd, ucd, ust)
print(tb, tbd, tcd, tst)
| GOLEM database analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h4> Importing libraries </h4>
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
sns.set(style="darkgrid")
import plotly.offline as py
import plotly.figure_factory as ff
import plotly.graph_objs as go
from plotly import tools
from plotly.offline import download_plotlyjs, init_notebook_mode, iplot, plot
# -
# ## 1 . EXPLORATORY DATA ANALYSIS
# ### Exploratory Data Analysis refers to the critical process of performing initial investigations on data so as to discover patterns,to spot anomalies,to test hypothesis and to check assumptions with the help of summary statistics and graphical representations.
#
# ## <b>Client:</b> <NAME> (Risk averse)
# #### Available data : Extracted 800,000 data records from kaggle dataset with complete loan data issued through the 2007-2015
# <h3>1.1 Reading lending club data</h3>
#
dataset = pd.read_csv('../Data/loan.csv',low_memory=False)
data_dictionary =pd.read_excel('../Data/LCDataDictionary.xlsx')
dataset.describe()
dataset.info()
# <h3>1.2 Understanding the data (categorical and numerical) </h3>
datatypes=dataset.dtypes.value_counts()
datatypes_list = datatypes.index.tolist()
datatypesdf=pd.DataFrame()
datatypesdf["datatype"]=datatypes_list
datatypesdf["total"]=[datatypes[0],datatypes[1],datatypes[2]]
datatypesdf
# <h4>Given dataset have total 53 numerical features and 24 non-numeric
datatypesdf.plot(x="datatype",y="total", kind='bar',title="Available data types")
# <h3> 1.3 Missing Value Analysis</h3>
#dataset['desc']
dataset.isna().sum()
columns_with_most_nulls = []
columns_with_data=[]
for key,null_value in (dataset.isnull().sum()/len(dataset)*100).sort_values().iteritems():
if null_value > 50:
columns_with_most_nulls.append([key,null_value])
else:
columns_with_data.append([key,null_value])
print("No of columns with more than 50% missing data: ",len(columns_with_most_nulls))
print("No of columns with considerable amount of data: ",len(columns_with_data))
missingdataanalysis_df=pd.DataFrame({"Column Analysis":["columns_with_most_nulls","columns_with_data"],"count":[21,53]})
missingdataanalysis_df.plot.barh(x='Column Analysis',y="count")
#
# <h3> 1.4 Statistical Analysis</h3>
set(dataset['loan_status'])
loan_status = dataset['loan_status'].value_counts()
dataset['addr_state'].value_counts().plot.bar(title = 'Loan Counts by USA states')
# +
data = []
names=[]
for name in set(dataset.loc[:,"addr_state"]):
data.append(dataset[dataset["addr_state"]==name]["int_rate"])
names.append(name)
# layout = go.Layout(title = 'Interested Rate based on Loan Status',
# xaxis = dict(title = 'State'),
# yaxis = dict(title = 'Interest Rate'))
# fig = dict(data = data, layout = layout)
# py.iplot(fig)
# +
# Create a figure instance
fig = plt.figure(1, figsize=(40, 10))
# Create an axes instance
ax = fig.add_subplot(111)
# Create the boxplot
bp = ax.boxplot(data)
bp = ax.boxplot(data, patch_artist=True)
## change outline color, fill color and linewidth of the boxes
for box in bp['boxes']:
# change outline color
box.set( color='Black', linewidth=2)
# change fill color
box.set( facecolor = 'Indigo' )
## change color and linewidth of the whiskers
for whisker in bp['whiskers']:
whisker.set(color='Black', linewidth=2)
## change color and linewidth of the caps
for cap in bp['caps']:
cap.set(color='Red', linewidth=2)
## change color and linewidth of the medians
for median in bp['medians']:
median.set(color='Yellow', linewidth=2)
## change the style of fliers and their fill
for flier in bp['fliers']:
flier.set(marker='o', color='Green', alpha=0.5)
ax.set_xticklabels(names,fontsize=20)
ax.set_yticklabels(np.arange(0,31,5),fontsize=20)
ax.set_title("Interest rates vs States",fontsize=40)
ax.get_yaxis().tick_left()
# -
print(names)
status_data = []
stat_names=[]
for name in set(dataset.loc[:,"loan_status"]):
status_data.append(dataset[dataset["loan_status"]==name]["int_rate"])
stat_names.append(name)
# +
# Create a figure instance
fig = plt.figure(1, figsize=(40, 10))
# Create an axes instance
ax = fig.add_subplot(111)
# Create the boxplot
bp = ax.boxplot(status_data)
bp = ax.boxplot(status_data, patch_artist=True)
## change outline color, fill color and linewidth of the boxes
for box in bp['boxes']:
# change outline color
box.set( color='Black', linewidth=2)
# change fill color
box.set( facecolor = 'Indigo' )
## change color and linewidth of the whiskers
for whisker in bp['whiskers']:
whisker.set(color='Black', linewidth=2)
## change color and linewidth of the caps
for cap in bp['caps']:
cap.set(color='Red', linewidth=2)
## change color and linewidth of the medians
for median in bp['medians']:
median.set(color='Yellow', linewidth=2)
## change the style of fliers and their fill
for flier in bp['fliers']:
flier.set(marker='o', color='Green', alpha=0.5)
ax.set_xticklabels(stat_names,rotation=90,fontsize=20)
ax.set_yticklabels(np.arange(0,31,5),fontsize=20)
ax.set_title("Interest Rates vs Loan_Status",fontsize=40)
ax.set_ylabel("Interest Rates",rotation=90,fontsize=30)
ax.get_yaxis().tick_left()
# -
completed_loans = dataset.loc[dataset["loan_status"].isin(["Fully Paid","Charged Off"]),:]
completed_loans
index = loan_status.index.tolist()
value = loan_status.tolist()
loan_statusdf = pd.DataFrame()
loan_statusdf['name'] = index
loan_statusdf['value'] = value
loan_statusdf
# +
# Plot the figure.
plt.figure(figsize=(7, 5))
ax = loan_status.plot(kind='bar')
ax.set_title('Loan status')
#ax.set_xlabel('value ($)')
#ax.set_ylabel('index ($)')
#ax.set_yticklabels(value)
rects = ax.patches
# Make some labels.
labels = [i for i in (loan_statusdf['value'].tolist())]
for rect, label in zip(rects, labels):
height = rect.get_height()
ax.text(rect.get_x() + rect.get_width() / 2, height + 5, label,
ha='center', va='bottom')
# -
issue_dates=dataset['issue_d']
year = []
for i in issue_dates:
year.append(i[4:])
dataset['year'] = year
dataset['sum'] =1
dataset['count'] =1
year = dataset.groupby(by=["year"])['int_rate'].mean().index.values.tolist()
int_rate_avg_year = dataset.groupby(by=["year"])['int_rate'].mean().tolist()
int_rate_avg_year
plt.scatter(year, int_rate_avg_year, color = 'Orange')
plt.grid
plt.title('Year vs Average Intrest rate')
dataset.groupby(by=["year"])['int_rate'].mean().sort_values().plot(kind='line',title='Interest rate by year',figsize=(20,10),xticks=['2007', '2010', '2008', '2011', '2009', '2015', '2012', '2014','2013'])
dataset.groupby(by=["year"]).loan_status.value_counts().to_csv('byyear.csv')
dataset.groupby(by=["purpose"]).mean()["int_rate"].sort_values().plot.bar(title = 'Average intrest rate vs purpose')
dataset.groupby(by=["grade"]).mean()["int_rate"].sort_values().plot.bar(title = 'Average intrest rate vs Grade')
dataset.groupby(by=["sub_grade"]).mean()["int_rate"].sort_values().plot.bar(title = 'Average intrest rate vs SUb Grade')
| Problem_Framing/.ipynb_checkpoints/Understanding_Data-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="farifxiKU1aB" papermill={"duration": 6.366772, "end_time": "2021-08-08T17:54:06.029281", "exception": false, "start_time": "2021-08-08T17:53:59.662509", "status": "completed"} tags=[]
import warnings
import gc
import tensorflow as tf
from tensorflow import keras
from random import choice
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, GRU, Concatenate, Embedding, Flatten, Activation, Dropout
from sklearn.model_selection import KFold
from tensorflow.python.client import device_lib
warnings.filterwarnings('ignore')
import random
# + id="9kZqV9siDyNb" papermill={"duration": 0.36364, "end_time": "2021-08-08T18:00:28.005034", "exception": false, "start_time": "2021-08-08T18:00:27.641394", "status": "completed"} tags=[]
MAXLENGTH = 400
EMBEDDING_DIM = 128
# + id="1MksD1JizpPn" papermill={"duration": 0.36214, "end_time": "2021-08-08T18:00:28.725215", "exception": false, "start_time": "2021-08-08T18:00:28.363075", "status": "completed"} tags=[]
FEATURES_SIZE = 2
CHAPTER_SIZE = 38
SUB_CHAPTER_SIZE = 223
QUESTION_SIZE = 1069
# + id="mY3Thp6d0NaT" papermill={"duration": 1.487414, "end_time": "2021-08-08T18:00:31.293949", "exception": false, "start_time": "2021-08-08T18:00:29.806535", "status": "completed"} tags=[]
#create dataset class
#to prepare it for train, valid, and test sets
from torch.utils.data import Dataset, DataLoader
class SPACE_DATASET(Dataset):
def __init__(self, data, maxlength = 400):
super(SPACE_DATASET, self).__init__()
self.maxlength = maxlength
self.data = data
self.users = list()
for user in data.index:
self.users.append(user)
def __len__(self):
return len(self.users)
def __getitem__(self, ix):
user = self.users[ix]
user = user
target, term, ch_label, sub_ch_label, ques_name, features = self.data[user]
#0s should be used as padding values
ori_target = target.values
term = term.values
ch_label = ch_label.values + 1
sub_ch_label = sub_ch_label.values +1
ques_name = ques_name.values + 1
n = len(ch_label)
# one hot for term
term_encode = [0]*7
term_encode[term[0]] = 1
shifted_target= []
# get user interaction informations in the previous MAXLEN interactions
if n > self.maxlength:
ch_label = ch_label[-self.maxlength:]
sub_ch_label = sub_ch_label[-self.maxlength:]
ques_name = ques_name[-self.maxlength:]
features = features[-self.maxlength:]
target = ori_target[-self.maxlength:]
shifted_target = ori_target[ (-self.maxlength - 1) :-1]
else:
ch_label = [0]*(self.maxlength - n)+list(ch_label[:])
sub_ch_label = [0]*(self.maxlength - n)+list(sub_ch_label[:])
ques_name = [0]*(self.maxlength - n)+list(ques_name[:])
features = [[0]*len(features[0])]*(self.maxlength - n)+list(features[:])
target = [-1]*(self.maxlength - n) + list(ori_target[:])
shifted_target = [2]*(self.maxlength + 1 - n) + list(ori_target[:-1])
new_features = []
count = 0
for f in features:
temp = list(f)
# temp.extend(term_encode)
#temp.append(shifted_target[count]) #uncomment this line for include previous response feature
new_features.append(temp)
count += 1
features = new_features
return ch_label,sub_ch_label,ques_name,features, shifted_target, target
# + papermill={"duration": 0.366532, "end_time": "2021-08-08T18:00:32.015989", "exception": false, "start_time": "2021-08-08T18:00:31.649457", "status": "completed"} tags=[]
SUB_CHAPTER_SIZE
# + [markdown] id="4xc90-aLzxat" papermill={"duration": 0.356272, "end_time": "2021-08-08T18:00:32.730579", "exception": false, "start_time": "2021-08-08T18:00:32.374307", "status": "completed"} tags=[]
# ## KFOLD - GRU
#
# + id="gzJrljnjzypP" outputId="87abe488-b493-4f8f-9d71-45cb1d2ddf51" papermill={"duration": 190.404719, "end_time": "2021-08-08T18:03:43.528203", "exception": false, "start_time": "2021-08-08T18:00:33.123484", "status": "completed"} tags=[]
# 5 fold cross validation with LSTM-based model
X = np.array(grouped_data.keys())
kfold = KFold(n_splits=5, shuffle=True)
train_losses = list()
train_aucs = list()
val_losses = list()
val_aucs = list()
train_eval = list()
test_eval = list()
for train, test in kfold.split(X):
users_train, users_test = X[train], X[test]
n = len(users_test)//2
users_test, users_val = users_test[:n], users_test[n: ]
train_data_space = SPACE_DATASET(grouped_data[users_train], MAXLENGTH)
val_data_space = SPACE_DATASET(grouped_data[users_val], MAXLENGTH)
test_data_space = SPACE_DATASET(grouped_data[users_test], MAXLENGTH)
#construct training input
train_chapter=[]
train_sub_chapter=[]
train_question = []
train_features=[]
train_labels=[]
for i in range(len(users_train)):
user = train_data_space.__getitem__(i)
train_chapter.append(user[0])
train_sub_chapter.append(user[1])
train_question.append(user[2])
train_features.append(user[3])
train_labels.append(user[4])
train_chapter = np.array(train_chapter)
train_sub_chapter = np.array(train_sub_chapter)
train_question = np.array(train_question)
train_features = np.array(train_features)
train_labels= np.array(train_labels)[..., np.newaxis]
#construct validation input
val_chapter=[]
val_sub_chapter=[]
val_question = []
val_features=[]
val_labels=[]
for i in range(len(users_val)):
user = val_data_space.__getitem__(i)
val_chapter.append(user[0])
val_sub_chapter.append(user[1])
val_question.append(user[2])
val_features.append(user[3])
val_labels.append(user[4])
val_chapter = np.array(val_chapter)
val_sub_chapter = np.array(val_sub_chapter)
val_features = np.array(val_features)
val_question = np.array(val_question)
val_labels= np.array(val_labels)[..., np.newaxis]
# construct test input
test_chapter=[]
test_sub_chapter=[]
test_features=[]
test_question=[]
test_labels=[]
for i in range(len(users_test)):
user = test_data_space.__getitem__(i)
test_chapter.append(user[0])
test_sub_chapter.append(user[1])
test_question.append(user[2])
test_features.append(user[3])
test_labels.append(user[4])
test_chapter = np.array(test_chapter)
test_sub_chapter = np.array(test_sub_chapter)
test_features = np.array(test_features)
test_question = np.array(test_question)
test_labels= np.array(test_labels)[..., np.newaxis]
# define loss function and evaluation metrics
bce = tf.keras.losses.BinaryCrossentropy(from_logits=True)
acc = tf.keras.metrics.Accuracy()
auc = tf.keras.metrics.AUC()
def masked_bce(y_true, y_pred):
flat_pred = y_pred
flat_ground_truth = y_true
label_mask = tf.math.not_equal(flat_ground_truth, -1)
return bce(flat_ground_truth, flat_pred, sample_weight=label_mask)
def masked_acc(y_true, y_pred):
flat_pred = y_pred
flat_ground_truth = y_true
flat_pred = (flat_pred >= 0.5)
label_mask = tf.math.not_equal(flat_ground_truth, -1)
return acc(flat_ground_truth, flat_pred, sample_weight=label_mask)
def masked_auc(y_true, y_pred):
flat_pred = y_pred
flat_ground_truth = y_true
label_mask = tf.math.not_equal(flat_ground_truth, -1)
return auc(flat_ground_truth, flat_pred, sample_weight=label_mask)
# input layer
input_chap = tf.keras.Input(shape=(MAXLENGTH))
input_sub_chap = tf.keras.Input(shape=(MAXLENGTH))
input_ques = tf.keras.Input(shape=(MAXLENGTH))
input_features = tf.keras.Input(shape=(MAXLENGTH, FEATURES_SIZE))
# embedding layer for categorical features
embedding_chap = Embedding(input_dim = CHAPTER_SIZE, output_dim = EMBEDDING_DIM)(input_chap)
embedding_sub_chap = Embedding(input_dim = SUB_CHAPTER_SIZE, output_dim = EMBEDDING_DIM)(input_sub_chap)
embedding_ques = Embedding(input_dim = QUESTION_SIZE, output_dim = EMBEDDING_DIM)(input_ques)
# dense layer for numeric features
dense_features = Dense(EMBEDDING_DIM,input_shape = (None, MAXLENGTH))(input_features)
output = tf.concat([embedding_chap, embedding_sub_chap, embedding_ques, dense_features], axis = 2)
pred = Dense(1, input_shape = (None, 4*EMBEDDING_DIM), activation='sigmoid')(output)
model = tf.keras.Model(
inputs=[input_chap, input_sub_chap,input_ques, input_features],
outputs=pred,
name='logistic_regression'
)
callback = tf.keras.callbacks.EarlyStopping(monitor='val_loss', patience=10)
opt_adam = Adam(learning_rate = 0.005)
model.compile(
optimizer=opt_adam,
loss= masked_bce,
metrics = [masked_acc, masked_auc]
)
history = model.fit(
[train_chapter, train_sub_chapter, train_question, train_features],
train_labels,
batch_size = 64,
epochs = 100,
validation_data=([val_chapter, val_sub_chapter, val_question, val_features], val_labels),
callbacks=[callback]
)
val_losses.append(list(history.history['val_loss']))
train_losses.append(list(history.history['loss']))
val_aucs.append(list(history.history['val_masked_auc']))
train_aucs.append(list(history.history['masked_auc']))
train_score = model.evaluate([train_chapter, train_sub_chapter, train_question, train_features], train_labels)
train_eval.append(train_score)
test_score = model.evaluate([test_chapter, test_sub_chapter, test_question, test_features], test_labels)
test_eval.append(test_score)
print("Test: ", test_score)
def reset_weights(model):
for layer in model.layers:
if isinstance(layer, tf.keras.Model):
reset_weights(layer)
continue
for k, initializer in layer.__dict__.items():
if "initializer" not in k:
continue
# find the corresponding variable
var = getattr(layer, k.replace("_initializer", ""))
var.assign(initializer(var.shape, var.dtype))
reset_weights(model)
# + id="QsVmumHMz3lx" outputId="4ff1e2fa-6abb-458e-c729-495b456f53e5" papermill={"duration": 0.910577, "end_time": "2021-08-08T18:03:45.409722", "exception": false, "start_time": "2021-08-08T18:03:44.499145", "status": "completed"} tags=[]
t_eval = np.array(test_eval)
print("test avg loss: ", np.mean(t_eval[:, 0]), "+/-" ,np.std(t_eval[:, 0]))
print("test avg acc: ", np.mean(t_eval[:, 1]), "+/-" ,np.std(t_eval[:, 1]))
print("test avg auc: ", np.mean(t_eval[:, 2]), "+/-" ,np.std(t_eval[:, 2]))
# + id="b9MM_CXWz5K6" outputId="4cf88e1d-3a74-4e7d-f92c-d01522e91757" papermill={"duration": 0.906324, "end_time": "2021-08-08T18:03:47.217207", "exception": false, "start_time": "2021-08-08T18:03:46.310883", "status": "completed"} tags=[]
t_eval = np.array(train_eval)
print("train avg loss: ", np.mean(t_eval[:, 0]), "+/-" ,np.std(t_eval[:, 0]))
print("train avg acc: ", np.mean(t_eval[:, 1]), "+/-" ,np.std(t_eval[:, 1]))
print("train avg auc: ", np.mean(t_eval[:, 2]), "+/-" ,np.std(t_eval[:, 2]))
| additional_features/baseline/logistic_regression/s400/logistic_regression_TF.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import re
import requests
from bs4 import BeautifulSoup
import json
import warnings
warnings.filterwarnings(action='once')
'''
Example format for extraction:
Count | Card Name | (Set# Card#)
2 Trail Stories (Set1 #188)
4 Horus Traver (Set1002 #23)
4 Longbarrel (Set4 #5)
4 On the Hunt (Set2 #5)
4 Oni Dragonsmith (Set1003 #2)
'''
# -
# # First step: Login and extract collection fully from eternal warcry.
# ## Note: Remove user/pass lane later!
# +
"""
For those paying attention mechanize is python 2.x only, someone made mechanicalsoup
(Love you), which if you read the tutorial we give you a brief intro along with show you how
to login. This is compatible /w python 3.6. The code below takes in two command
line parameterse delimited by a space; username password.
ie: aggregate_card_json.py <username> <password>
For now I have my u+p information manually punched in and works!
"""
# import argparse
# parser = argparse.ArgumentParser(description="Login to EternalWarcry.")
# parser.add_argument("username")
# parser.add_argument("password")
# args = parser.parse_args()
user = '<user>'
passwd = '<<PASSWORD>>'
import mechanicalsoup
browser = mechanicalsoup.Browser(soup_config={'features': 'lxml'})
# request eternalWarcry login page. the result is a requests.Response object
login_page = browser.get("https://eternalwarcry.com/login")
# login_page.soup is a BeautifulSoup object
# we grab the login form
login_form = mechanicalsoup.Form(login_page.soup.select_one('form[action="/login"]'))
# specify username and password
login_form.input({"LoginEmail": user, "LoginPassword": <PASSWORD>})# REPLACE WITH args.username & args.password
# submit form
page2 = browser.submit(login_form, login_page.url)
# Verify we logged in at Eternal Warcry...
print(page2.soup.title.text)
"""
STEP TWO: ITERATE THROUGH EACH PAGE AND GRAB ALL OF THE INFORMATION FOR EACH CARD,
PUT IT INTO A STRING IN THE PRETTY FORMAT THAT ETERNAL FOLLOWS, AND APPEND INTO A BIG LIST
OUTPUT WHICH IS YOUR COLLECTION FROM ETERNALWARCRY
(All this really does is save you time by not having to go into ->"Deck Builder" and
add ALL your cards to a deck and exporting it.... :))
"""
page_num = 1
check_last_pg = re.compile(r'\b(No cards found in your collection)\b')
final_collection = []
while True:
# Now that were logged in pull each link for our collection and grab the info
current_page = 'https://eternalwarcry.com/collection?view=oo&p=' + str(page_num)
page_num = page_num + 1 # Iterate don't wanna forget ;)
collection_page = browser.get(current_page) # Response object with .soup object
# Get out of loop if we reach 'no cards exists' page.
if bool(check_last_pg.search(collection_page.text)):
break
# Current Page. #collection_page.text because its response object
html_page = collection_page.soup
# Get all cards on this page.
divs = html_page.findAll(class_= 'card-search-item col-lg-3 col-sm-4 col-xs-6 add-card-main element-relative')
for div in divs:
# This is where it gets fun.
# Each div is a card from the search_view. a href has the link which contains info.
card_name = str(div.find_all('a'))
#print(card_name) # This is to be searched for the name of card; can also acquire set and card#
card_details = str(div.find(class_ = 'display-count'))
# STEP TWO: FORMULATE CARD
# First part of string is count
count = str(re.search(r'data-count="(\d)"', card_details, re.IGNORECASE).group(1))
# Now get set - card#
card_set = str(re.search(r'data-card="(\d+-\d+)"', card_details, re.IGNORECASE).group(1))
card_set = card_set.split('-')
details = '(Set' + str(card_set[0]) + ' #' + str(card_set[1]) + ')'
# Now get card name with grammar
name = str(re.search(r'<img alt="(.+)" class=', card_name, re.IGNORECASE).group(1))
# Put it altogether....
item = count + ' ' + name + ' ' + details
#print(item)
# Drop each page into a list for later...
final_collection.append(item)
# final_collection # Uncomment to see list.
# -
len(final_collection) # Wow that numbers off! Eternal Statistics say I should have 68% 3,691/5,424 cards total.
# But let's think: IT's because it is only looking at the unique cards! So lets do math.
# +
# Grab first character of each string (which we know is the count of each card in our collection up to 4) and add it.
total_cards = 0
for x in final_collection:
total_cards = total_cards + int(x[0])
total_cards
output_json = {'total_cards': total_cards, 'my_collection': final_collection}
# -
# Cool! WE got an accurate record of our cards, minus the Commons and UnCommons. Success for tonight.
# Output to a JSON file because sexy
with open('my_collection.json', 'w') as fp:
json.dump(output_json, fp)
| Old/AutoEternal/.ipynb_checkpoints/get_my_collection-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Sameem25/2015-PyDataParis/blob/master/XGBoost_basic.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="FDK8pEtI6kje" colab_type="code" colab={}
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# + id="1zKpURa26yAD" colab_type="code" colab={}
import xgboost as xgb
from sklearn.model_selection import train_test_split
# + id="yKd-fgTC7pbQ" colab_type="code" colab={}
#reading the data (csv file)
data = pd.read_csv('data.csv')
# + id="h0bnDjvHLtvS" colab_type="code" outputId="beea6807-20dd-414b-d8e5-807dbf744373" colab={"base_uri": "https://localhost:8080/", "height": 419}
data
# + id="eyL7lWqGLv48" colab_type="code" colab={}
X, y = data.iloc[:,:-1], data.iloc[:,-1]
# + id="cTdaXW9NSK4K" colab_type="code" colab={}
X_train, X_test, y_train, y_test = train_test_split(X,y, test_size = 0.2)
# + id="4m-eePU4SrnY" colab_type="code" colab={}
xg_cl = xgb.XGBClassifier(objective = 'binary:logistic', n_estimators= 1000, learning_rate= 0.1)
# + id="jd5HYkScTeB6" colab_type="code" outputId="90a18e79-4bbc-4fd9-c4b4-76fa4bc4c985" colab={"base_uri": "https://localhost:8080/", "height": 136}
xg_cl.fit(X_train, y_train)
# + id="jWB2UIAL8LZX" colab_type="code" colab={}
preds = xg_cl.predict(X_test)
# + id="_iDmDgGHT9yi" colab_type="code" outputId="8c3d7d76-f142-46c8-f657-b9ba85f51d73" colab={"base_uri": "https://localhost:8080/", "height": 34}
accuracy = float(np.sum(preds==y_test))/y_test.shape[0]
accuracy
# + id="CEYbi2ZDNcYC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 102} outputId="874aee59-a445-4d3c-dce2-59d1041c0774"
from sklearn.model_selection import GridSearchCV
xgb_model = xgb.XGBClassifier()
optimization_dict = {'max_depth': [2,3,4,5,6],
'n_estimators': [50,100,200,500,1000,2000],
'learning_rate':[0.01,0.1,0.2,0.05]}
model = GridSearchCV(xgb_model, optimization_dict,
scoring='accuracy', verbose=1)
model.fit(X_train,y_train)
print(model.best_score_)
print(model.best_params_)
| XGBoost_basic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import dask
import numpy as np
import xarray as xr
import glob
import matplotlib.pyplot as plt
import time
import datetime
from dask_jobqueue import SLURMCluster
from dask.distributed import Client, progress
import warnings
warnings.filterwarnings('ignore')
now = datetime.datetime.now()
now_string = str(now.strftime("%Y-%m-%d_%A_%H%M%S"))
now_string
cluster = SLURMCluster(cores=16, memory='20GB', project='pi_jianwu', queue='high_mem', job_extra=['--qos=medium+','--exclusive'])
cluster.scheduler
cluster.scale(4)
cluster.start_workers()
cluster.scheduler_address
# !squeue -u savio1
client = Client('tcp://10.2.1.32:46292')
cluster.dashboard_link
cluster.running_jobs
cluster.pending_jobs
print(client)
t0 = time.time()
total_pix = np.zeros((180, 360))
cloud_pix = np.zeros((180, 360))
def ingest_data(M03_dir, M06_dir):
M03_files = sorted(glob.glob(M03_dir + "MYD03.A2008*.hdf"))
M06_files = sorted(glob.glob(M06_dir + "MYD06_L2.A2008*.hdf"))
for M03, M06 in zip (M03_files, M06_files):
d06 = xr.open_mfdataset(M06[:], parallel=True)['Cloud_Mask_1km'][:,:,:].values
d06CM = d06[::3,::3,0]
ds06_decoded = (np.array(d06CM, dtype = "byte") & 0b00000110) >> 1
d03_lat = xr.open_mfdataset(M03[:], drop_variables = "Scan Type", parallel=True)['Latitude'][:,:].values
d03_lon = xr.open_mfdataset(M03[:], drop_variables = "Scan Type", parallel=True)['Longitude'][:,:].values
lat = d03_lat[::3,::3]
lon = d03_lon[::3,::3]
l_index = (lat + 89.5).astype(int).reshape(lat.shape[0]*lat.shape[1])
lat_index = np.where(l_index > -1, l_index, 0)
ll_index = (lon + 179.5).astype(int).reshape(lon.shape[0]*lon.shape[1])
lon_index = np.where(ll_index > -1, ll_index, 0)
for i, j in zip(lat_index, lon_index):
total_pix[i,j] += 1
indicies = np.nonzero(ds06_decoded <= 0)
row_i = indicies[0]
column_i = indicies[1]
cloud_lon = [lon_index.reshape(ds06_decoded.shape[0],ds06_decoded.shape[1])[i,j] for i, j in zip(row_i, column_i)]
cloud_lat = [lat_index.reshape(ds06_decoded.shape[0],ds06_decoded.shape[1])[i,j] for i, j in zip(row_i, column_i)]
for x, y in zip(cloud_lat, cloud_lon):
cloud_pix[int(x),int(y)] += 1
return cloud_pix, total_pix
t0 = time.time()
import dask.multiprocessing
dask.config.set(num_workers=5)
M03_dir = '/umbc/xfs1/jianwu/common/MODIS_Aggregation/MODIS_one_day_data/'
M06_dir = '/umbc/xfs1/jianwu/common/MODIS_Aggregation/MODIS_one_day_data/'
cluster.running_jobs
t0 = time.time()
import dask.multiprocessing
dask.config.set(num_workers=5)
M03_dir = "/umbc/xfs1/jianwu/common/MODIS_Aggregation/MODIS_one_day_data/"
M06_dir = "/umbc/xfs1/jianwu/common/MODIS_Aggregation/MODIS_one_day_data/"
future1 = client.submit(ingest_data,M03_dir,M06_dir)
y = future1.result()
progress(y)
cf1 = future1.result()[0]/future1.result()[1]
progress(cf1)
plt.figure(figsize=(14,7))
plt.contourf(range(-180,180), range(-90,90), cf1, 100, cmap = "jet")
plt.xlabel("Longitude", fontsize = 14)
plt.ylabel("Latitude", fontsize = 14)
plt.title("Level 3 Cloud Fraction Aggregation For One Month %s" %now_string, fontsize = 16)
plt.colorbar()
plt.savefig("/umbc/xfs1/jianwu/common/MODIS_Aggregation/savioexe/test/8/%s.png" %now_string)
cf2 = xr.DataArray(cf1)
cf2.to_netcdf("/umbc/xfs1/jianwu/common/MODIS_Aggregation/savioexe/test/8/%s.hdf" %now_string)
t1 = time.time()
total = t1-t0
print(total,"seconds")
print(total/60,"minutes")
cluster.pending_jobs
cluster.close()
client.close()
# !squeue -u savio1
| source/8/DaskAttemptOneDay.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# Author: <NAME> (@thalesians) - Managing Director & Co-founder of [the Thalesians](http://www.thalesians.com)
# ## Introduction
# With the UK general election in early May 2015, we thought it would be a fun exercise to demonstrate how you can investigate market price action over historial elections. We shall be using Python, together with Plotly for plotting. Plotly is a free web-based platform for making graphs. You can keep graphs private, make them public, and run Plotly on your [Chart Studio Enterprise on your own servers](https://plotly.com/product/enterprise/). You can find more details [here](https://plotly.com/python/getting-started/).
# ## Getting market data with Bloomberg
# To get market data, we shall be using Bloomberg. As a starting point, we have used bbg_py from [Brian Smith's TIA project](https://github.com/bpsmith/tia/tree/master/tia/bbg), which allows you to access Bloomberg via COM (older method), modifying it to make it compatible for Python 3.4. Whilst, we shall note use it to access historical daily data, there are functions which enable us to download intraday data. This method is only compatible with 32 bit versions of Python and assumes you are running the code on a Bloomberg terminal (it won't work without a valid Bloomberg licence).
#
# In my opinion a better way to access Bloomberg via Python, is via the official Bloomberg open source Python Open Source Graphing Library, however, at time of writing the official version is not yet compatible with Python 3.4. Fil Mackay has created a Python 3.4 compatible version of this [here](https://github.com/filmackay/blpapi-py), which I have used successfully. Whilst it takes slightly more time to configure (and compile using Windows SDK 7.1), it has the benefit of being compatible with 64 bit Python, which I have found invaluable in my analysis (have a read of [this](http://ta.speot.is/2012/04/09/visual-studio-2010-sp1-windows-sdk-7-1-install-order/) in case of failed installations of Windows SDK 7.1).
#
# Quandl can be used as an alternative data source, if you don't have access to a Bloomberg terminal, which I have also included in the code.
# ## Breaking down the steps in Python
# Our project will consist of several parts:
# - bbg_com - low level interaction with BBG COM object (adapted for Python 3.4) (which we are simply calling)
# - datadownloader - wrapper for BBG COM, Quandl and CSV access to data
# - eventplot - reusuable functions for interacting with Plotly and creating event studies
# - ukelection - kicks off the whole script process
# ### Downloading the market data
# As with any sort of financial market analysis, the first step is obtaining market data. We create the DataDownloader class, which acts a wrapper for Bloomberg, Quandl and CSV market data. We write a single function "download_time_series" for this. We could of course extend this for other data sources such as Yahoo Finance. Our output will be Pandas based dataframes. We want to make this code generic, so the tickers are not hard coded.
# +
# for time series manipulation
import pandas
class DataDownloader:
def download_time_series(self, vendor_ticker, pretty_ticker, start_date, source, csv_file = None):
if source == 'Quandl':
import Quandl
# Quandl requires API key for large number of daily downloads
# https://www.quandl.com/help/api
spot = Quandl.get(vendor_ticker) # Bank of England's database on Quandl
spot = pandas.DataFrame(data=spot['Value'], index=spot.index)
spot.columns = [pretty_ticker]
elif source == 'Bloomberg':
from bbg_com import HistoricalDataRequest
req = HistoricalDataRequest([vendor_ticker], ['PX_LAST'], start = start_date)
req.execute()
spot = req.response_as_single()
spot.columns = [pretty_ticker]
elif source == 'CSV':
dateparse = lambda x: pandas.datetime.strptime(x, '%Y-%m-%d')
# in case you want to use a source other than Bloomberg/Quandl
spot = pandas.read_csv(csv_file, index_col=0, parse_dates=0, date_parser=dateparse)
return spot
# -
# ### Generic functions for event study and Plotly plotting
# We now focus our efforts on the EventPlot class. Here we shall do our basic analysis. We shall aslo create functions for creating plotly traces and layouts that we shall reuse a number of times. The analysis we shall conduct is fairly simple. Given a time series of spot, and a number of dates, we shall create an event study around these times for that asset. We also include the "Mean" move over all the various dates.
# +
# for dates
import datetime
# time series manipulation
import pandas
# for plotting data
import plotly
from plotly.graph_objs import *
class EventPlot:
def event_study(self, spot, dates, pre, post, mean_label = 'Mean'):
# event_study - calculates the asset price moves over windows around event days
#
# spot = price of asset to study
# dates = event days to anchor our event study
# pre = days before the event day to start our study
# post = days after the event day to start our study
#
data_frame = pandas.DataFrame()
# for each date grab spot data the days before and after
for i in range(0, len(dates)):
mid_index = spot.index.searchsorted(dates[i])
start_index = mid_index + pre
finish_index = mid_index + post + 1
x = (spot.ix[start_index:finish_index])[spot.columns.values[0]]
data_frame[dates[i]] = x.values
data_frame.index = range(pre, post + 1)
data_frame = data_frame / data_frame.shift(1) - 1 # returns
# add the mean on to the end
data_frame[mean_label] = data_frame.mean(axis=1)
data_frame = 100.0 * (1.0 + data_frame).cumprod() # index
data_frame.ix[pre,:] = 100
return data_frame
# -
# We write a function to convert dates represented in a string format to Python format.
def parse_dates(self, str_dates):
# parse_dates - parses string dates into Python format
#
# str_dates = dates to be parsed in the format of day/month/year
#
dates = []
for d in str_dates:
dates.append(datetime.datetime.strptime(d, '%d/%m/%Y'))
return dates
EventPlot.parse_dates = parse_dates
# Our next focus is on the Plotly functions which create a layout. This enables us to specify axes labels, the width and height of the final plot and so on. We could of course add further properties into it.
def create_layout(self, title, xaxis, yaxis, width = -1, height = -1):
# create_layout - populates a layout object
# title = title of the plot
# xaxis = xaxis label
# yaxis = yaxis label
# width (optional) = width of plot
# height (optional) = height of plot
#
layout = Layout(
title = title,
xaxis = plotly.graph_objs.XAxis(
title = xaxis,
showgrid = False
),
yaxis = plotly.graph_objs.YAxis(
title= yaxis,
showline = False
)
)
if width > 0 and height > 0:
layout['width'] = width
layout['height'] = height
return layout
EventPlot.create_layout = create_layout
# Earlier, in the DataDownloader class, our output was Pandas based dataframes. Our convert_df_plotly function will convert these each series from Pandas dataframe into plotly traces. Along the way, we shall add various properties such as markers with varying levels of opacity, graduated coloring of lines (which uses colorlover) and so on.
def convert_df_plotly(self, dataframe, axis_no = 1, color_def = ['default'],
special_line = 'Mean', showlegend = True, addmarker = False, gradcolor = None):
# convert_df_plotly - converts a Pandas data frame to Plotly format for line plots
# dataframe = data frame due to be converted
# axis_no = axis for plot to be drawn (default = 1)
# special_line = make lines named this extra thick
# color_def = color scheme to be used (default = ['default']), colour will alternate in the list
# showlegend = True or False to show legend of this line on plot
# addmarker = True or False to add markers
# gradcolor = Create a graduated color scheme for the lines
#
# Also see http://nbviewer.ipython.org/gist/nipunreddevil/7734529 for converting dataframe to traces
# Also see http://moderndata.plot.ly/color-scales-in-ipython-notebook/
x = dataframe.index.values
traces = []
# will be used for market opacity for the markers
increments = 0.95 / float(len(dataframe.columns))
if gradcolor is not None:
try:
import colorlover as cl
color_def = cl.scales[str(len(dataframe.columns))]['seq'][gradcolor]
except:
print('Check colorlover installation...')
i = 0
for key in dataframe:
scatter = plotly.graph_objs.Scatter(
x = x,
y = dataframe[key].values,
name = key,
xaxis = 'x' + str(axis_no),
yaxis = 'y' + str(axis_no),
showlegend = showlegend)
# only apply color/marker properties if not "default"
if color_def[i % len(color_def)] != "default":
if special_line in str(key):
# special case for lines labelled "mean"
# make line thicker
scatter['mode'] = 'lines'
scatter['line'] = plotly.graph_objs.Line(
color = color_def[i % len(color_def)],
width = 2
)
else:
line_width = 1
# set properties for the markers which change opacity
# for markers make lines thinner
if addmarker:
opacity = 0.05 + (increments * i)
scatter['mode'] = 'markers+lines'
scatter['marker'] = plotly.graph_objs.Marker(
color=color_def[i % len(color_def)], # marker color
opacity = opacity,
size = 5)
line_width = 0.2
else:
scatter['mode'] = 'lines'
scatter['line'] = plotly.graph_objs.Line(
color = color_def[i % len(color_def)],
width = line_width)
i = i + 1
traces.append(scatter)
return traces
EventPlot.convert_df_plotly = convert_df_plotly
# ### UK election analysis
# We've now created several generic functions for downloading data, doing an event study and also for helping us out with plotting via Plotly. We now start work on the ukelection.py script, for pulling it all together. As a very first step we need to provide credentials for Plotly (you can get your own Plotly key and username [here](https://plotly.com/python/getting-started/)).
# +
# for time series/maths
import pandas
# for plotting data
import plotly
import plotly.plotly as py
from plotly.graph_objs import *
def ukelection():
# Learn about API authentication here: https://plotly.com/python/getting-started
# Find your api_key here: https://plotly.com/settings/api
plotly_username = "thalesians"
plotly_api_key = "<KEY>"
plotly.tools.set_credentials_file(username=plotly_username, api_key=plotly_api_key)
# -
# Let's download our market data that we need (GBP/USD spot data) using the DataDownloader class. As a default, I've opted to use Bloomberg data. You can try other currency pairs or markets (for example FTSE), to compare results for the event study. Note that obviously each data vendor will have a different ticker in their system for what could well be the same asset. With FX, care must be taken to know which close the vendor is snapping. As a default we have opted for BGN, which for GBP/USD is the NY close value.
ticker = 'GBPUSD' # will use in plot titles later (and for creating Plotly URL)
##### download market GBP/USD data from Quandl, Bloomberg or CSV file
source = "Bloomberg"
# source = "Quandl"
# source = "CSV"
csv_file = None
event_plot = EventPlot()
data_downloader = DataDownloader()
start_date = event_plot.parse_dates(['01/01/1975'])
if source == 'Quandl':
vendor_ticker = "BOE/XUDLUSS"
elif source == 'Bloomberg':
vendor_ticker = 'GBPUSD BGN Curncy'
elif source == 'CSV':
vendor_ticker = 'GBPUSD'
csv_file = 'D:/GBPUSD.csv'
spot = data_downloader.download_time_series(vendor_ticker, ticker, start_date[0], source, csv_file = csv_file)
# The most important part of the study is getting the historical UK election dates! We can obtain these from Wikipedia. We then convert into Python format. We need to make sure we filter the UK election dates, for where we have spot data available.
labour_wins = ['28/02/1974', '10/10/1974', '01/05/1997', '07/06/2001', '05/05/2005']
conservative_wins = ['03/05/1979', '09/06/1983', '11/06/1987', '09/04/1992', '06/05/2010']
# convert to more easily readable format
labour_wins_d = event_plot.parse_dates(labour_wins)
conservative_wins_d = event_plot.parse_dates(conservative_wins)
# only takes those elections where we have data
labour_wins_d = [d for d in labour_wins_d if d > spot.index[0].to_pydatetime()]
conservative_wins_d = [d for d in conservative_wins_d if d > spot.index[0].to_pydatetime()]
spot.index.name = 'Date'
# We then call our event study function in EventPlot on our spot data, which compromises of the 20 days before up till the 20 days after the UK general election. We shall plot these lines later.
# number of days before and after for our event study
pre = -20
post = 20
# calculate spot path during Labour wins
labour_wins_spot = event_plot.event_study(spot, labour_wins_d, pre, post, mean_label = 'Labour Mean')
# calculate spot path during Conservative wins
conservative_wins_spot = event_plot.event_study(spot, conservative_wins_d, pre, post, mean_label = 'Conservative Mean')
# Define our xaxis and yaxis labels, as well as our source, which we shall later include in the title.
##### Create separate plots of price action during Labour and Conservative wins
xaxis = 'Days'
yaxis = 'Index'
source_label = "Source: @thalesians/BBG/Wikipedia"
# We're finally ready for our first plot! We shall plot GBP/USD moves over Labour election wins, using the default palette and then we shall embed it into the sheet, using the URL given to us from the Plotly website.
###### Plot market reaction during Labour UK election wins
###### Using default color scheme
title = ticker + ' during UK gen elect - Lab wins' + '<BR>' + source_label
fig = Figure(data=event_plot.convert_df_plotly(labour_wins_spot),
layout=event_plot.create_layout(title, xaxis, yaxis)
)
py.iplot(fig, filename='labour-wins-' + ticker)
# The "iplot" function will send it to Plotly's server (provided we have all the dependencies installed).
# Alternatively, we could embed the HTML as an image, which we have taken from the Plotly website. Note this approach will yield a static image which is fetched from Plotly's servers. It also possible to write the image to disk. Later we shall show the embed function.
# <div>
# <a href="https://plotly.com/~thalesians/244/" target="_blank" title="GBPUSD during UK gen elect - Lab wins<br>Source: @thalesians/BBG/Wikipedia" style="display: block; text-align: center;"><img src="https://plotly.com/~thalesians/244.png" alt="GBPUSD during UK gen elect - Lab wins<br>Source: @thalesians/BBG/Wikipedia" style="max-width: 100%;" onerror="this.onerror=null;this.src='https://plotly.com/404.png';" /></a>
# <script data-plotly="thalesians:244" src="https://plotly.com/embed.js" async></script>
# </div>
#
# We next plot GBP/USD over Conservative wins. In this instance, however, we have a graduated 'Blues' color scheme, given obviously that blue is the color of the Conserative party in the UK!
###### Plot market reaction during Conservative UK election wins
###### Using varying shades of blue for each line (helped by colorlover library)
title = ticker + ' during UK gen elect - Con wins ' + '<BR>' + source_label
# also apply graduated color scheme of blues (from light to dark)
# see http://moderndata.plot.ly/color-scales-in-ipython-notebook/ for details on colorlover package
# which allows you to set scales
fig = Figure(data=event_plot.convert_df_plotly(conservative_wins_spot, gradcolor='Blues', addmarker=False),
layout=event_plot.create_layout(title, xaxis, yaxis),
)
plot_url = py.iplot(fig, filename='conservative-wins-' + ticker)
# Embed the chart into the document using "embed". This essentially embeds the JavaScript code, necessary to make it interactive.
# +
import plotly.tools as tls
tls.embed("https://plotly.com/~thalesians/245")
# -
# Our final plot, will consist of three subplots, Labour wins, Conservative wins, and average moves for both. We also add a grid and a grey background for each plot.
##### Plot market reaction during Conservative UK election wins
##### create a plot consisting of 3 subplots (from left to right)
##### 1. Labour wins, 2. Conservative wins, 3. Conservative/Labour mean move
# create a dataframe which grabs the mean from the respective Lab & Con election wins
mean_wins_spot = pandas.DataFrame()
mean_wins_spot['Labour Mean'] = labour_wins_spot['Labour Mean']
mean_wins_spot['Conservative Mean'] = conservative_wins_spot['Conservative Mean']
fig = plotly.tools.make_subplots(rows=1, cols=3)
# apply different color scheme (red = Lab, blue = Con)
# also add markets, which will have varying levels of opacity
fig['data'] += Data(
event_plot.convert_df_plotly(conservative_wins_spot, axis_no=1,
color_def=['blue'], addmarker=True) +
event_plot.convert_df_plotly(labour_wins_spot, axis_no=2,
color_def=['red'], addmarker=True) +
event_plot.convert_df_plotly(mean_wins_spot, axis_no=3,
color_def=['red', 'blue'], addmarker=True, showlegend = False)
)
fig['layout'].update(title=ticker + ' during UK gen elects by winning party ' + '<BR>' + source_label)
# use the scheme from https://plotly.com/python/bubble-charts-tutorial/
# can use dict approach, rather than specifying each separately
axis_style = dict(
gridcolor='#FFFFFF', # white grid lines
ticks='outside', # draw ticks outside axes
ticklen=8, # tick length
tickwidth=1.5 # and width
)
# create the various axes for the three separate charts
fig['layout'].update(xaxis1=plotly.graph_objs.XAxis(axis_style, title=xaxis))
fig['layout'].update(yaxis1=plotly.graph_objs.YAxis(axis_style, title=yaxis))
fig['layout'].update(xaxis2=plotly.graph_objs.XAxis(axis_style, title=xaxis))
fig['layout'].update(yaxis2=plotly.graph_objs.YAxis(axis_style))
fig['layout'].update(xaxis3=plotly.graph_objs.XAxis(axis_style, title=xaxis))
fig['layout'].update(yaxis3=plotly.graph_objs.YAxis(axis_style))
fig['layout'].update(plot_bgcolor='#EFECEA') # set plot background to grey
plot_url = py.iplot(fig, filename='labour-conservative-wins-'+ ticker + '-subplot')
# This time we use "embed", which grab the plot from Plotly's server, we did earlier (given we have already uploaded it).
# +
import plotly.tools as tls
tls.embed("https://plotly.com/~thalesians/246")
# -
# <B>That's about it!</B> I hope the code I've written proves fruitful for creating some very cool Plotly plots and also for doing some very timely analysis ahead of the UK general election! Hoping this will be first of many blogs on using Plotly data.
# The analysis in this blog is based on a report I wrote for Thalesians, a quant finance thinktank. If you are interested in getting access to the full copy of the report (Thalesians: My kingdom for a vote - The definitive quant guide to UK general elections), feel free to e-mail me at <b><EMAIL></b> or tweet me <b>@thalesians</b>
# ## Want to hear more about global macro and UK election developments?
# If you're interested in FX and the UK general election, come to our Thalesians panel in London on April 29th 2015 at 7.30pm in Canary Wharf, which will feature, <NAME> (Reuters - FX Buzz Editor), <NAME> (Bloomberg - First Word EM Strategist), <NAME> (Nomura - FX strategist), <NAME> (Independent FX trader) and myself as the moderator. Tickets are available [here](http://www.meetup.com/thalesians/events/221147156/)
# ## Biography
# <b><NAME></b> is the managing director and co-founder of the Thalesians. He has a decade of experience creating and successfully running systematic trading models at Lehman Brothers, Nomura and now at the Thalesians. Independently, he runs a systematic trading model with proprietary capital. He is the author of Trading Thalesians – What the ancient world can teach us about trading today (Palgrave Macmillan). He graduated with a first class honours master’s degree from Imperial College in Mathematics & Computer Science. He is also a fan of Python and has written an extensive library for financial market backtesting called PyThalesians.
# <BR>
#
# Follow the Thalesians on Twitter @thalesians and get my book on Amazon [here](http://www.amazon.co.uk/Trading-Thalesians-Saeed-Amen/dp/113739952X)
# All the code here is available to download from the [Thalesians GitHub page](https://github.com/thalesians/pythalesians)
# +
from IPython.display import display, HTML
display(HTML('<link href="//fonts.googleapis.com/css?family=Open+Sans:600,400,300,200|Inconsolata|Ubuntu+Mono:400,700" rel="stylesheet" type="text/css" />'))
display(HTML('<link rel="stylesheet" type="text/css" href="http://help.plot.ly/documentation/all_static/css/ipython-notebook-custom.css">'))
# ! pip install publisher --upgrade
import publisher
publisher.publish(
'ukelectionbbg.ipynb', 'ipython-notebooks/ukelectionbbg/', 'Plotting GBP/USD price action around UK general elections',
'Create interactive graphs with market data, IPython Notebook and Plotly', name='Plot MP Action in GBP/USD around UK General Elections')
# -
| _posts/python-v3/ipython-notebooks/ukelectionbbg.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import torch
from torch import nn, optim
from torch.utils.data import DataLoader, Dataset
from torchvision import datasets, transforms
from torchvision.utils import make_grid
import matplotlib
from matplotlib import pyplot as plt
import seaborn as sns
from IPython import display
import torchsummary as ts
import numpy as np
sns.set()
display.set_matplotlib_formats("svg")
plt.rcParams['font.sans-serif'] = "Liberation Sans"
device = torch.device("cuda")
trans = transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))
])
dataset = datasets.ImageFolder("dataset/faces/", transform=trans)
data_loader = DataLoader(dataset, batch_size=16, shuffle=True, num_workers=4,
drop_last=True)
images = make_grid(next(iter(data_loader))[0], normalize=True, padding=5, pad_value=1)
plt.imshow(images.permute(1, 2, 0))
plt.axis("off")
plt.grid(False)
def imshow(data):
images = make_grid(data.detach().cpu() , normalize=True, padding=5, pad_value=1)
plt.imshow(images.permute(1, 2, 0))
plt.axis("off")
plt.grid(False)
plt.pause(0.0001)
def weights_init(m):
classname = m.__class__.__name__
if classname.find('Conv') != -1:
nn.init.normal_(m.weight.data, 0.0, 0.02)
elif classname.find('BatchNorm') != -1:
nn.init.normal_(m.weight.data, 1.0, 0.02)
nn.init.constant_(m.bias.data, 0)
class Discriminator(nn.Module):
def __init__(self,nc=3,ndf=32):
super().__init__()
self.layer1 = nn.Sequential(nn.Conv2d(nc,ndf,kernel_size=4,stride=2,padding=1),
nn.BatchNorm2d(ndf),
nn.LeakyReLU(0.2,inplace=True))
self.layer2 = nn.Sequential(nn.Conv2d(ndf,ndf*2,kernel_size=4,stride=2,padding=1),
nn.BatchNorm2d(ndf*2),
nn.LeakyReLU(0.2,inplace=True))
self.layer3 = nn.Sequential(nn.Conv2d(ndf*2,ndf*4,kernel_size=4,stride=2,padding=1),
nn.BatchNorm2d(ndf*4),
nn.LeakyReLU(0.2,inplace=True))
self.layer4 = nn.Sequential(nn.Conv2d(ndf*4,ndf*8,kernel_size=4,stride=2,padding=1),
nn.BatchNorm2d(ndf*8),
nn.LeakyReLU(0.2,inplace=True))
self.fc = nn.Sequential(nn.Linear(256*6*6,1),nn.Sigmoid())
def forward(self,x):
out = self.layer1(x)
out = self.layer2(out)
out = self.layer3(out)
out = self.layer4(out)
out = out.view(-1,256*6*6)
out = self.fc(out)
out = out.reshape(-1)
return out
class Generator(nn.Module):
def __init__(self,nc=3, ngf=128,nz=1024,feature_size=100):
super().__init__()
self.prj = nn.Linear(feature_size,nz*6*6)
self.layer1 = nn.Sequential(nn.ConvTranspose2d(nz,ngf*4,kernel_size=4,stride=2,padding=1),
nn.BatchNorm2d(ngf*4),
nn.ReLU())
self.layer2 = nn.Sequential(nn.ConvTranspose2d(ngf*4,ngf*2,kernel_size=4,stride=2,padding=1),
nn.BatchNorm2d(ngf*2),
nn.ReLU())
self.layer3 = nn.Sequential(nn.ConvTranspose2d(ngf*2,ngf,kernel_size=4,stride=2,padding=1),
nn.BatchNorm2d(ngf),
nn.ReLU())
self.layer4 = nn.Sequential(nn.ConvTranspose2d(ngf,nc,kernel_size=4,stride=2,padding=1),
nn.Tanh())
def forward(self,x):
out = self.prj(x)
out = out.view(-1,1024,6,6)
out = self.layer1(out)
out = self.layer2(out)
out = self.layer3(out)
out = self.layer4(out)
return out
netD = Discriminator()
netD(torch.randn(16, 3, 96, 96)).shape
netG = Generator()
netG(torch.randn(16, 100)).shape
BATCH_SIZE = 256
ININT_SIZE = 100
data_loader = DataLoader(dataset, batch_size=BATCH_SIZE,
shuffle=True, num_workers=4, drop_last=True)
Epoch = 500
current_epoch = 1
D_losses = []
G_losses = []
generator = Generator().to(device)
discirminator = Discriminator().to(device)
generator.apply(weights_init)
discirminator.apply(weights_init)
criterion = nn.BCELoss()
OPTIMIZER_G = optim.Adam(generator.parameters(), lr=4e-4, betas=(0.5, 0.999))
OPTIMIZER_D = optim.Adam(discirminator.parameters(), lr=1e-4, betas=(0.5, 0.999))
prebs = []
def display_prebs(data):
fig, ax = plt.subplots()
ax.plot(np.arange(len(data)) + 1,
[item[0] for item in data], label="pdr", ls="-.")
ax.plot(np.arange(len(data)) + 1,
[item[1] for item in data], label="pdf", ls="--")
ax.plot(np.arange(len(data)) + 1,
[item[2] for item in data], label="pg", ls=":")
ax.set_xlabel("Epoch")
ax.set_ylabel("Acc")
ax.set_title("Probs in GAN Training Process")
plt.savefig("GNA-Probs.svg", format="svg")
ax.legend()
plt.pause(0.00001)
discirminator = nn.DataParallel(discirminator, device_ids=[0, 1])
generator = nn.DataParallel(generator, device_ids=[0, 1])
discirminator = discirminator.to("cuda")
generator = generator.to("cuda")
Epoch = 1000
pdr, pdf, pg = None, None, None
for epoch in range(current_epoch, 1 + Epoch):
dis_temp_loss = []
gen_temp_loss = []
temp_pdr = []
temp_pdf = []
temp_pg =[]
for idx, (d, l) in enumerate(data_loader):
d = d.to(device)
l = l.float().to(device)
out = discirminator(d)
pdr = out.mean().item()
real_loss = criterion(out, torch.ones_like(l))
noise = torch.randn(BATCH_SIZE, ININT_SIZE).to(device)
images = generator(noise)
out = discirminator(images.detach().to(device))
pdf = out.mean().item()
fake_loss = criterion(out, torch.zeros_like(l))
OPTIMIZER_D.zero_grad()
real_loss.backward()
fake_loss.backward()
OPTIMIZER_D.step()
noise = torch.randn(BATCH_SIZE, ININT_SIZE).to(device)
images = generator(noise)
out = discirminator(images)
pg = out.mean().item()
loss = criterion(out, torch.ones_like(l))
OPTIMIZER_G.zero_grad()
loss.backward()
OPTIMIZER_G.step()
d_loss = fake_loss + real_loss
temp_pdr.append(pdr)
temp_pdf.append(pdf)
temp_pg.append(pg)
print("Epoch = {:<2} Step[{:3}/{:3}] Dis-Loss = {:.5f} Gen-Loss = {:.5f} Acc = {:.2f} {:.2f} {:.2f}"\
.format(current_epoch, idx + 1, len(data_loader), d_loss.item(),
loss.item(), pdr, pdf, pg))
dis_temp_loss.append(d_loss.item())
gen_temp_loss.append(loss.item())
D_losses.append(np.mean(dis_temp_loss))
G_losses.append(np.mean(gen_temp_loss))
current_epoch +=1
prebs.append((np.mean(temp_pdr), np.mean(temp_pdf), np.mean(temp_pg)))
if epoch > 1:
fig, ax = plt.subplots()
ax.plot(np.arange(len(D_losses)) + 1,
D_losses, label="Discriminator", ls="-.")
ax.plot(np.arange(len(G_losses)) + 1,
G_losses, label="Generator", ls="--")
ax.set_xlabel("Epoch")
ax.set_ylabel("Loss")
ax.set_title("GAN Training process")
ax.legend(bbox_to_anchor=[1, 1.02])
plt.pause(0.0001)
imshow(images[:16])
imshow(d[:16])
display_prebs(prebs)
if epoch % 10 == 0:
display.clear_output()
torch.save(generator.state_dict(), "gen_final.pkl")
torch.save(discirminator.state_dict(), "dis_final.pkl")
fig, ax = plt.subplots()
ax.plot(np.arange(len(D_losses)) + 1,
D_losses, label="Discriminator", ls="-.")
ax.plot(np.arange(len(G_losses)) + 1,
G_losses, label="Generator", ls="--")
ax.set_xlabel("Epoch")
ax.set_ylabel("Loss")
ax.set_title("GAN Training process")
ax.legend()
plt.savefig("GNA-Loss.svg", format="svg")
generator.load_state_dict(torch.load("gen_final.pkl"))
generator = generator.eval()
with torch.no_grad():
noise = torch.randn(16, ININT_SIZE).to(device)
images = generator(noise)
data = make_grid(d[:16].detach().cpu() , nrow=4,
normalize=True, padding=5, pad_value=1)
plt.imshow(data.permute(1, 2, 0))
plt.axis("off")
plt.grid(False)
plt.savefig("RealAnimation.png", format="png", dpi=200)
display_prebs(prebs)
| ModelCollaps.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <div align='center'><font size="5" color='#353B47'>Predicting Stock Prices</font></div>
#
# <br>
# <hr>
# <img src="https://toocooltrafficschool.com/wp-content/uploads/2019/02/what-is-tlsae740.png" width="500">
# ## <div id="summary">Summary</div>
#
# **<font size="2"><a href="#chap1">1. Import libraries</a></font>**
# **<br><font size="2"><a href="#chap2">2. Preprocessing</a></font>**
# **<br><font size="2"><a href="#chap3">3. Build LSTM model</a></font>**
# **<br><font size="2"><a href="#chap4">4. Training</a></font>**
# **<br><font size="2"><a href="#chap5">5. Predictions</a></font>**
# # <div id="chap1">1. Import libraries
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5"
import numpy as np
import pandas as pd
import os
import matplotlib.pyplot as plt
import pandas_datareader as web
import datetime as dt
from sklearn.preprocessing import MinMaxScaler
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Dropout, LSTM
from tensorflow.keras.callbacks import ModelCheckpoint, EarlyStopping
# -
# **<font size="2"><a href="#summary">Back to summary</a></font>**
#
# ----
# # <div id="chap2">2. Preprocessing
def load_data(company, start, end):
data = web.DataReader(company, 'yahoo', start, end)
return data
# Company to be focused on: facebook
company = 'WIPRO.NS'
# +
# data = load_data(company = company,
# start = dt.datetime(2012,1,1),
# end = dt.datetime(2019,1,1))
# import pandas_datareader as pdr
# import datetime as dt
# ticker = "WIPRO.NS"
# start = dt.datetime(2019, 1, 1)
# end = dt.datetime(2020, 12, 31)
# data = pdr.get_data_yahoo(ticker, start, end)
# print(data)
# -
data= pd.read_csv("./WIPRO.NS.csv")
# Normalize data
scaler = MinMaxScaler(feature_range=(0,1))
scaled_data = scaler.fit_transform(data['Close'].values.reshape(-1,1))
# +
# how many days do i want to base my predictions on ?
prediction_days = 60
x_train = []
y_train = []
for x in range(prediction_days, len(scaled_data)):
x_train.append(scaled_data[x - prediction_days:x, 0])
y_train.append(scaled_data[x, 0])
x_train, y_train = np.array(x_train), np.array(y_train)
x_train = np.reshape(x_train, (x_train.shape[0], x_train.shape[1], 1))
# -
# **<font size="2"><a href="#summary">Back to summary</a></font>**
#
# ----
# # <div id="chap3">3. Build LSTM model
# **<font color="blue" size="4">What is a LSTM ?</font>**
#
# > Long Short Term Memory networks – usually just called “LSTMs” – are a special kind of RNN, capable of learning long-term dependencies. Introduced by Hochreiter & Schmidhuber (1997), and were refined and popularized by many people in following work. They work tremendously well on a large variety of problems, and are now widely used.
# >
# > LSTMs are explicitly designed to avoid the long-term dependency problem. Remembering information for long periods of time is practically their default behavior, not something they struggle to learn!
# >
# > All recurrent neural networks have the form of a chain of repeating modules of neural network.
# You can find more details here: http://colah.github.io/posts/2015-08-Understanding-LSTMs/
def LSTM_model():
model = Sequential()
model.add(LSTM(units = 50, return_sequences = True, input_shape = (x_train.shape[1],1)))
model.add(Dropout(0.2))
model.add(LSTM(units = 50, return_sequences = True))
model.add(Dropout(0.2))
model.add(LSTM(units = 50))
model.add(Dropout(0.2))
model.add(Dense(units=1))
return model
# **<font size="2"><a href="#summary">Back to summary</a></font>**
#
# ----
# # <div id="chap4">4. Training
model = LSTM_model()
model.summary()
model.compile(optimizer='adam',
loss='mean_squared_error')
# +
# Define callbacks
# Save weights only for best model
checkpointer = ModelCheckpoint(filepath = 'weights_best.hdf5',
verbose = 2,
save_best_only = True)
model.fit(x_train,
y_train,
epochs=25,
batch_size = 32,
callbacks = [checkpointer])
# -
# **<font size="2"><a href="#summary">Back to summary</a></font>**
#
# ----
model.save("model.h5")
# # <div id="chap5">5. Predictions
# +
# test model accuracy on existing data
test_data = data
actual_prices = test_data['Close'].values
total_dataset = pd.concat((data['Close'], test_data['Close']), axis=0)
model_inputs = total_dataset[len(total_dataset) - len(test_data) - prediction_days:].values
model_inputs = model_inputs.reshape(-1,1)
model_inputs = scaler.transform(model_inputs)
# +
x_test = []
for x in range(prediction_days, len(model_inputs)):
x_test.append(model_inputs[x-prediction_days:x, 0])
x_test = np.array(x_test)
x_test = np.reshape(x_test, (x_test.shape[0], x_test.shape[1] ,1))
predicted_prices = model.predict(x_test)
predicted_prices = scaler.inverse_transform(predicted_prices)
# -
plt.plot(actual_prices, color='black', label=f"Actual {company} price")
plt.plot(predicted_prices, color= 'green', label=f"predicted {company} price")
plt.title(f"{company} share price")
plt.xlabel("time")
plt.ylabel(f"{company} share price")
plt.legend()
plt.show()
# +
# predicting next day
real_data = [model_inputs[len(model_inputs)+1 - prediction_days:len(model_inputs+1),0]]
real_data = np.array(real_data)
print(real_data.shape)
real_data = np.reshape(real_data, (real_data.shape[0], real_data.shape[1], 1))
# -
prediction = model.predict(real_data)
prediction = scaler.inverse_transform(prediction)
print(f"prediction: {prediction}")
# **<font size="2"><a href="#summary">Back to summary</a></font>**
#
# ----
# # References
#
# * https://www.youtube.com/c/Neural
#
# * https://www.kaggle.com/bryanb/introduction-to-time-series-analysis/edit/run/53321733
#
# * http://colah.github.io/posts/2015-08-Understanding-LSTMs/
# <hr>
# <br>
# <div align='justify'><font color="#353B47" size="4">Thank you for taking the time to read this notebook. I hope that I was able to answer your questions or your curiosity and that it was quite understandable. <u>any constructive comments are welcome</u>. They help me progress and motivate me to share better quality content. I am above all a passionate person who tries to advance my knowledge but also that of others. If you liked it, feel free to <u>upvote and share my work.</u> </font></div>
# <br>
# <div align='center'><font color="#353B47" size="3">Thank you and may passion guide you.</font></div>
| stock-prices-forecasting-with-lstm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import os
import glob
import librosa
from librosa import load
from librosa import cqt
from librosa.display import specshow
from tqdm import tqdm
# -
data_dir = '/Users/sripathisridhar/Desktop/SOL'
hop_size= 512
q= 24
# +
file_paths= sorted(glob.glob(os.path.join(data_dir, '**', '*.wav')))
file_names= []
for file_path in file_paths:
file_names.append(os.path.basename(file_path))
# +
features_dict= {}
feature_key= ''
for file_path in tqdm(file_paths, disable=False):
# Read audio files
waveform, sample_rate= load(file_path, sr=None)
# Compute CQTs
cqt_complex= cqt(y=waveform,
sr=sample_rate,
hop_length=hop_size,
bins_per_octave=q,
n_bins=q*7,
sparsity=1e-6,
)
scalogram= np.abs(cqt_complex)**2
# Find frame with maximum RMS value
rms= librosa.feature.rms(y=waveform, hop_length=hop_size)
rms_argmax= np.argmax(rms)
feature= scalogram[:,rms_argmax]
# Stack in dict
file_name= os.path.basename(file_path)
feature_key= f'{file_name}'
features_dict[feature_key]= feature
# +
import h5py
with h5py.File("SOL.h5", "w") as f:
for key in features_dict.keys():
f[key] = features_dict[key]
# -
with h5py.File("SOL.h5", "r") as f:
my_dict = {key:f[key][()] for key in f.keys()}
[key for i,key in enumerate(my_dict.keys()) if i in range(500,510)]
| FeatureExtraction/preCompute_SOL.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# %matplotlib inline
import matplotlib.pyplot as plt
import netCDF4 as nc
import numpy as np
from salishsea_tools import bathy_tools
from salishsea_tools import nc_tools
factor_of_grid = 1.0 # amount of a grid point we deepen by
min_depth_deepen = 4.7 # minimum depth that we deepen
bathy = nc.Dataset('../../nemo-forcing/grid/bathy_meter_SalishSea6.nc', 'r')
print bathy.file_format
print bathy.variables.keys
depths = bathy.variables['Bathymetry'][:]
fig = bathy_tools.plot_colourmesh(
bathy, 'NEMO - Salish Sea Bathymetry',
axis_limits=(-124, -122.9, 48.5, 49.2), colour_map='spectral', bins=30)
grid = nc.Dataset('/ocean/sallen/allen/research/MEOPAR/NEMO-forcing/grid/mesh_mask_SalishSea2.nc')
#nc_tools.show_variable_attrs(grid)
depth_chunks = grid.variables['e3t_0'][0,:]
print depth_chunks, depth_chunks.shape
depth_places = grid.variables['gdept_0'][0,:]
print depth_places, depth_places.shape
new_depths = np.copy(depths)
mask = depths > 0
notmask = depths == 0
# stackoverflow by unutbu
def find_nearest(array, value):
idx = (np.abs(array - value)).argmin()
return idx
imin = 0; imax = 898
jmin = 0; jmax = 398
for i in range(imin, imax):
for j in range(jmin, jmax):
if depths[i,j] > min_depth_deepen:
idx = find_nearest(depth_places, depths[i,j])
new_depths[i,j] = depths[i,j] + factor_of_grid * depth_chunks[idx]
new_depths_masked = np.ma.array(new_depths, mask=notmask)
fig, axs = plt.subplots(1, 3, figsize=(15, 10))
axs[0].pcolormesh(depths[imin:imax, jmin:jmax], vmax=450, vmin=0)
axs[1].pcolormesh(new_depths_masked[imin:imax, jmin:jmax], vmax=450, vmin=0)
mesh = axs[2].pcolormesh(new_depths_masked - depths)
fig.colorbar(mesh, ax=axs[2])
plt.plot(-depths[250:400,75])
plt.plot(-new_depths_masked[250:400,75])
# lets smooth
max_dh_over_hbar = 0.8
depthsmooth = bathy_tools.smooth(new_depths_masked[:], max_norm_depth_diff=max_dh_over_hbar, smooth_factor=0.2)
fig, axs = plt.subplots(1, 3, figsize=(15, 10))
axs[0].pcolormesh(new_depths_masked[imin:imax, jmin:jmax], vmax=450, vmin=0)
axs[1].pcolormesh(depthsmooth[imin:imax, jmin:jmax], vmax=450, vmin=0)
mesh = axs[2].pcolormesh(depthsmooth - new_depths_masked)
fig.colorbar(mesh, ax=axs[2])
plt.pcolormesh(depthsmooth[imin-10:imax+10, jmin-10:jmax+10]-new_depths_masked[imin-10:imax+10, jmin-10:jmax+10], cmap = 'bwr')
plt.colorbar()
# +
ysize = depthsmooth.shape[0]; xsize = depthsmooth.shape[1]
print ysize, xsize
new_bathy = nc.Dataset('bathy_downonegrid.nc', 'w')
new_bathy.createDimension('y', ysize)
new_bathy.createDimension('x', xsize)
nc_tools.show_dimensions(new_bathy)
# +
newlons = new_bathy.createVariable('nav_lon', float, ('y', 'x'), zlib=True)
newlons.setncattr('units', 'degrees_east')
newlats = new_bathy.createVariable('nav_lat', float, ('y', 'x'), zlib=True)
newlats.setncattr('units', 'degrees_north')
newdepths = new_bathy.createVariable(
'Bathymetry', float, ('y', 'x'),
zlib=True, least_significant_digit=0.1, fill_value=0)
newdepths.setncattr('units', 'metres')
newlons[:] = bathy.variables['nav_lon']
newlats[:] = bathy.variables['nav_lat']
newdepths[:] = depthsmooth[:]
# -
nc_tools.show_dataset_attrs(new_bathy)
print bathy.history
new_bathy.history = """
[2013-10-30 13:18] Created netCDF4 zlib=True dataset.
[2013-10-30 15:22] Set depths between 0 and 4m to 4m and those >428m to 428m.
[2013-10-31 17:10] Algorithmic smoothing.
[2013-11-21 19:53] Reverted to pre-smothing dataset (repo rev 3b301b5b9b6d).
[2013-11-21 20:14] Updated dataset and variable attributes to CF-1.6 conventions & project standards.
[2013-11-21 20:47] Removed east end of Jervis Inlet and Toba Inlet region due to deficient source bathymetry data in Cascadia dataset.
[2013-11-21 21:52] Algorithmic smoothing.
[2014-01-01 14:44] Smoothed mouth of Juan de Fuca
[2015-] Jie made Fraser River
[2015-12-24 11:46] dropped by one grid thickness, smoothed to 0.8
"""
# +
new_bathy.conventions = """
CF-1.6"""
new_bathy.title= """
Salish Sea NEMO Bathymetry"""
new_bathy.institution= """
Dept of Earth, Ocean & Atmospheric Sciences, University of British Columbia"""
new_bathy.references= """
https://bitbucket.org/salishsea/nemo-forcing/src/tip/grid/bathy_downonegrid.nc"""
new_bathy.comment= """
Based on 1_bathymetry_seagrid_WestCoast.nc file from 2-Oct-2013 WCSD_PREP tarball provided by J-<NAME>.
"""
new_bathy.source= """
https://github.com/SalishSeaCast/tools/blob/master/bathymetry/SalishSeaBathy.ipynb
https://github.com/SalishSeaCast/tools/blob/master/bathymetry/SmoothMouthJdF.ipynb
https://github.com/SalishSeaCast/tools/blob/master/bathymetry/Deepen by Grid Thickness.ipynb
"""
new_bathy.close()
# -
| bathymetry/Deepen by Grid Thickness.ipynb |
;; -*- coding: utf-8 -*-
;; ---
;; jupyter:
;; jupytext:
;; text_representation:
;; extension: .scm
;; format_name: light
;; format_version: '1.5'
;; jupytext_version: 1.14.4
;; kernelspec:
;; display_name: Calysto Scheme (Python)
;; language: scheme
;; name: calysto_scheme
;; ---
;; <img src="images/logo-64x64.png"/>
;; <h1>Reference Guide for Calysto Scheme</h1>
;;
;; [Calysto Scheme](https://github.com/Calysto/calysto_scheme) is a real Scheme programming language, with full support for continuations, including call/cc. It can also use all Python libraries. Also has some extensions that make it more useful (stepper-debugger, choose/fail, stack traces), or make it better integrated with Python.
;;
;; In Jupyter notebooks, because Calysto Scheme uses [MetaKernel](https://github.com/Calysto/metakernel/blob/master/README.rst), it has a fully-supported set of "magics"---meta-commands for additional functionality. This includes running Scheme in parallel. See all of the [MetaKernel Magics](https://github.com/Calysto/metakernel/blob/master/metakernel/magics/README.md).
;;
;; Calysto Scheme is written in Scheme, and then translated into Python (and other backends). The entire functionality lies in a single Python file: https://github.com/Calysto/calysto_scheme/blob/master/calysto_scheme/scheme.py However, you can easily install it (see below).
;;
;; Please see [Calysto Scheme Language](Calysto%20Scheme%20Language.ipynb) for more details on the Calysto Scheme language.
;;
;; ## Installation
;;
;; You can install Calysto Scheme with Python3:
;;
;; ```
;; pip3 install --upgrade calysto-scheme --user -U
;; python3 -m calysto_kernel install --user
;; ```
;;
;; or in the system kernel folder with:
;;
;; ```
;; sudo pip3 install --upgrade calysto-scheme -U
;; sudo python3 -m calysto_kernel install
;; ```
;;
;; Change pip3/python3 to use a different pip or Python. The version of Python used will determine how Calysto Scheme is run.
;;
;; Use it in the console, qtconsole, or notebook with IPython 3:
;;
;; ```
;; ipython console --kernel calysto_scheme
;; ipython qtconsole --kernel calysto_scheme
;; ipython notebook --kernel calysto_scheme
;; ```
;;
;; In addition to all of the following items, Calysto Scheme also has access to all of Python's builtin functions, and all of Python's libraries. For example, you can use `(complex 3 2)` to create a complex number by calling Python's complex function.
;;
;; ## Jupyter Enhancements
;;
;; When you run Calysto Scheme in Jupyter (console, notebook, qtconsole, etc.) you get:
;;
;; * TAB completions of Scheme functions and variable names
;; * display of rich media
;; * stepper/debugger
;; * magics (% macros)
;; * shell commands (! command)
;; * LaTeX equations
;; * LaTeX-style variables
;; * Python integration
;; ### LaTeX-style variables
;;
;; Calysto Scheme allows you to use LaTeX-style variables in code. For example, if you type:
;;
;; ```
;; \beta
;; ```
;;
;; with the cursor right after the 'a' in beta, and then press TAB, it will turn into the unicode character:
;;
;; ```
;; β
;; ```
;;
;; There are nearly 1300 different symbols defined (thanks to the Julia language) and documented here:
;;
;; http://docs.julialang.org/en/release-0.4/manual/unicode-input/#man-unicode-input
;;
;; Calysto Scheme may not implement all of those. Some useful and suggestive ones:
;;
;; * \pi - π
;; * \Pi - Π
;; * \Sigma - Σ
;; * \_i - subscript i, such as vectorᵢ
(define α 67)
α
(define i 2)
(define vectorᵢ (vector-ref (vector 0 6 3 2) i))
vectorᵢ
;; ### Rich media
(import "calysto.display")
(calysto.display.HTML "This is <b>bold</b>, <i>italics</i>, <u>underlined</u>.")
(import "calysto.graphics")
(define canvas (calysto.graphics.Canvas))
(define ball (calysto.graphics.Circle '(150 150) 100))
(ball.draw canvas)
;; ### Shell commands
! ls /tmp
;; ### Stepper/Debugger
;;
;; Here is what the debugger looks like:
;;
;; <img src="images/stepper_debugger.png">
;;
;; It has breakpoints (click in left margin). You must press Stop to exit the debugger.
;; ```scheme
;; ;; %%debug
;;
;; (begin
;; (define x 1)
;; (set! x 2)
;; )
;; ```
;; ### Python Integration
;;
;; You can import and use any Python library in Calysto Scheme.
;;
;; In addition, if you wish, you can execute expressions and statements in a Python environment:
(python-eval "1 + 2")
(python-exec
"
def mypyfunc(a, b):
return a * b
")
;; This is a shared environment with Scheme:
(mypyfunc 4 5)
;; You can use `func` to turn a Scheme procedure into a Python function, and `define!` to put it into the shared environment with Python:
(define! mypyfunc2 (func (lambda (n) n)))
(python-eval "mypyfunc2(34)")
;; # Differences Between Languages
;;
;; ## Major differences between Scheme and Python
;;
;; 1. In Scheme, double quotes are used for strings and may contain newlines
;; 1. In Scheme, a single quote is short for (quote ...) and means "literal"
;; 1. In Scheme, everything is an expression and has a return value
;; 1. Python does not support macros (e.g., extending syntax)
;; 1. In Python, "if X" is false if X is None, False, [], (,) or 0. In Scheme, "if X" is only false if X is #f or 0
;; 1. Calysto Scheme uses continuations, not the call stack. However, for debugging there is a pseudo-stack when an error is raised. You can turn that off with (use-stack-trace #f)
;; 1. Scheme procedures are not Python functions, but there are means to use one as the other.
;;
;; ## Major Differences Between Calysto Scheme and other Schemes
;;
;; 1. define-syntax works slightly differently
;; 1. In Calysto Scheme, #(...) is short for '#(...)
;; 1. Calysto Scheme is missing many standard functions (see list at bottom)
;; 1. Calysto Scheme has a built-in amb operator called `choose`
;; 1. For debugging there is a pseudo-stack when errors are raised in Calysto Scheme. You can turn that off with (use-stack-trace #f)
;;
;; ### Stack Trace
;;
;; Calysto Scheme acts as if it has a call stack, for easier debugging. For example:
(define fact
(lambda (n)
(if (= n 1)
q
(* n (fact (- n 1))))))
(fact 5)
;; To turn off the stack trace on error:
;;
;; ```scheme
;; (use-stack-trace #f)
;; ```
;; That will allow infinite recursive loops without keeping track of the "stack".
;; # Calysto Scheme Variables
;;
;; ## SCHEMEPATH
;; SCHEMEPATH is a list of search directories used with (load NAME). This is a reference, so you should append to it rather than attempting to redefine it.
SCHEMEPATH
(set-cdr! (cdr SCHEMEPATH) (list "/var/modules"))
SCHEMEPATH
;; ## Getting Started
;;
;; Note that you can use the word `lambda` or \lambda and then press [TAB]
(define factorial
(λ (n)
(cond
((zero? n) 1)
(else (* n (factorial (- n 1)))))))
(factorial 5)
;; ## define-syntax
;; (define-syntax NAME RULES): a method for creating macros
(define-syntax time
[(time ?exp) (let ((start (current-time)))
?exp
(- (current-time) start))])
(time (car '(1 2 3 4)))
;; +
;;---------------------------------------------------------------------
;; collect is like list comprehension in Python
(define-syntax collect
[(collect ?exp for ?var in ?list)
(filter-map (lambda (?var) ?exp) (lambda (?var) #t) ?list)]
[(collect ?exp for ?var in ?list if ?condition)
(filter-map (lambda (?var) ?exp) (lambda (?var) ?condition) ?list)])
(define filter-map
(lambda (f pred? values)
(if (null? values)
'()
(if (pred? (car values))
(cons (f (car values)) (filter-map f pred? (cdr values)))
(filter-map f pred? (cdr values))))))
;; -
(collect (* n n) for n in (range 10))
(collect (* n n) for n in (range 5 20 3))
(collect (* n n) for n in (range 10) if (> n 5))
;; +
;;---------------------------------------------------------------------
;; for loops
(define-syntax for
[(for ?exp times do . ?bodies)
(for-repeat ?exp (lambda () . ?bodies))]
[(for ?var in ?exp do . ?bodies)
(for-iterate1 ?exp (lambda (?var) . ?bodies))]
[(for ?var at (?i) in ?exp do . ?bodies)
(for-iterate2 0 ?exp (lambda (?var ?i) . ?bodies))]
[(for ?var at (?i ?j . ?rest) in ?exp do . ?bodies)
(for ?var at (?i) in ?exp do
(for ?var at (?j . ?rest) in ?var do . ?bodies))])
(define for-repeat
(lambda (n f)
(if (< n 1)
'done
(begin
(f)
(for-repeat (- n 1) f)))))
(define for-iterate1
(lambda (values f)
(if (null? values)
'done
(begin
(f (car values))
(for-iterate1 (cdr values) f)))))
(define for-iterate2
(lambda (i values f)
(if (null? values)
'done
(begin
(f (car values) i)
(for-iterate2 (+ i 1) (cdr values) f)))))
;; +
(define matrix2d
'((10 20)
(30 40)
(50 60)
(70 80)))
(define matrix3d
'(((10 20 30) (40 50 60))
((70 80 90) (100 110 120))
((130 140 150) (160 170 180))
((190 200 210) (220 230 240))))
;; -
(begin
(define hello 0)
(for 5 times do (set! hello (+ hello 1)))
hello
)
(for sym in '(a b c d) do (define x 1) (set! x sym) (print x))
(for n in (range 10 20 2) do (print n))
(for n at (i j) in matrix2d do (print (list n 'coords: i j)))
(for n at (i j k) in matrix3d do (print (list n 'coords: i j k)))
;; +
(define-syntax scons
[(scons ?x ?y) (cons ?x (lambda () ?y))])
(define scar car)
(define scdr
(lambda (s)
(let ((result ((cdr s))))
(set-cdr! s (lambda () result))
result)))
(define first
(lambda (n s)
(if (= n 0)
'()
(cons (scar s) (first (- n 1) (scdr s))))))
(define nth
(lambda (n s)
(if (= n 0)
(scar s)
(nth (- n 1) (scdr s)))))
(define smap
(lambda (f s)
(scons (f (scar s)) (smap f (scdr s)))))
(define ones (scons 1 ones))
(define nats (scons 0 (combine nats + ones)))
(define combine
(lambda (s1 op s2)
(scons (op (scar s1) (scar s2)) (combine (scdr s1) op (scdr s2)))))
(define fibs (scons 1 (scons 1 (combine fibs + (scdr fibs)))))
(define facts (scons 1 (combine facts * (scdr nats))))
(define ! (lambda (n) (nth n facts)))
;; -
(! 5)
(nth 10 facts)
(nth 20 fibs)
(first 30 fibs)
;; ## for-each
;; (for-each PROCEDURE LIST): apply PROCEDURE to each item in LIST; like `map` but don't return results
(for-each (lambda (n) (print n)) '(3 4 5))
;; ## format
;; (format STRING ITEM ...): format the string with ITEMS as arguments
(format "This uses formatting ~a ~s ~%" 'apple 'apple)
;; ## func
;;
;; Turns a lambda into a Python function.
;;
;; (func (lambda ...))
(func (lambda (n) n))
;; ## There's more!
;;
;; Please see [Calysto Scheme Language](Calysto%20Scheme%20Language.ipynb) for more details on the Calysto Scheme language.
| tests/notebooks/ipynb_scheme/Reference Guide for Calysto Scheme.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
from dotenv import load_dotenv, find_dotenv
from os.path import join, dirname, basename, exists, isdir
### Load environmental variables from the project root directory ###
# find .env automagically by walking up directories until it's found
dotenv_path = find_dotenv()
# load up the entries as environment variables
load_dotenv(dotenv_path)
# now you can get the variables using their names
# Check whether a network drive has been specified
DATABASE = os.environ.get("NETWORK_URL")
if DATABASE == 'None':
pass
else:
pass
#mount network drive here
# set up directory paths
CURRENT_DIR = os.getcwd()
PROJ = dirname(dotenv_path) # project root directory
DATA = join(PROJ, 'data') #data directory
RAW_EXTERNAL = join(DATA, 'raw_external') # external data raw directory
RAW_INTERNAL = join(DATA, 'raw_internal') # internal data raw directory
INTERMEDIATE = join(DATA, 'intermediate') # intermediate data directory
FINAL = join(DATA, 'final') # final data directory
RESULTS = join(PROJ, 'results') # output directory
FIGURES = join(RESULTS, 'figures') # figure output directory
PICTURES = join(RESULTS, 'pictures') # picture output directory
# make folders specific for certain data
folder_name = ''
if folder_name != '':
#make folders if they don't exist
if not exists(join(RAW_EXTERNAL, folder_name)):
os.makedirs(join(RAW_EXTERNAL, folder_name))
if not exists(join(INTERMEDIATE, folder_name)):
os.makedirs(join(INTERMEDIATE, folder_name))
if not exists(join(FINAL, folder_name)):
os.makedirs(join(FINAL, folder_name))
print('Standard variables loaded, you are good to go!')
# -
# 1. Abundance [mmol/cell] = Abundance [mmol/gDW] * ( cell volume [fL/cell] * cell density [g/fL] * dry content [gDW/g] )
# 2. Abundance [molecules/cell] = Abundance [mmol/cell] * Na [molecules/mol] * 1000 [mmol/mol]
#
# +
import pandas as pd
import re
# import data
data = pd.read_csv(f"{INTERMEDIATE}/proteomics_concentrations.csv", index_col=0)
# get cell volumes
cell_volumes = pd.read_csv(f"{RAW_INTERNAL}/proteomics/growth_conditions.csv", index_col=0)
cell_volumes = cell_volumes["Single cell volume [fl]1"]
# remove the first two rows of LB
cell_volumes = cell_volumes.loc[~cell_volumes.index.duplicated(keep='first')]
# rename the number 3 in there
cell_volumes = cell_volumes.rename({'Osmotic-stress glucose3':'Osmotic-stress glucose_uncertainty'}, axis='index')
rename_dict = {i:re.sub(r'\W+', '', i).lower() for i in cell_volumes.index}
cell_volumes = cell_volumes.rename(rename_dict, axis='index')
# Finally, convert to mmol/gDW:
water_content = 0.3
cell_density = 1.105e-12
# Iterate through the dataset and multiply by the corresponding cell volume, to get mmol/fL:
for (col_name, d) in data.iteritems():
chemo_name = col_name.replace("_uncertainty", "").replace("_mean", "")
try:
data[col_name] = data[col_name] * cell_volumes.loc[chemo_name]#["cell_volume"]
except:
print(chemo_name)
data = data * cell_density * water_content
# convert into counts
data = data * 6.022e+23 / 1000
data
# -
original_data = pd.read_csv(f"{RAW_INTERNAL}/proteomics/protein_values.csv", index_col=0)
original_data
| data_science/code/.ipynb_checkpoints/conversion_proteomics_conc2count-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## What is Supervised Learning
#
# There are few types of machine learning. Supervised Learning is one of them.
#
# The fundamental concept is letting a system learn from lots of **labeled** data.
#
# After the learning, the system will be able to predict the result when new data come.
#
# This is supervised learning.
#
# **Labeled** data means, we know the meaning of our data. Example can be,
#
# - Given fact, like house size, location, year of build, we know the price. Here the price is the label. House size, location and year of build are called features.
# - Given a photo, we know whether is a cat. Here whether is a cat is the label. And photo is the features.
#
# ## How Learn a Cat is a Cat
#
# When we were young, someone, likely our parents, told us that is a cat when we see a cat.
#
# So we just looked at the cat visually and label it as a cat.
#
# And late on, when we see a cat, we predict it.
#
# - Sometime we are right, our parents will say good job.
# - Sometime we are left, our parents will say, no, that is not a cat.
#
# Over time, we will get better predication, super close to 100%.
#
# Our parents are our supervisor in that case. We are learning in supervised way.
# ## Before Learning
#
# Without previous knowledge, let a system tell whether a photo is a cat.
#
# The system is a plain cubic box, it can only guess randomly. Either yes, it's a cat. Or no, it's not a cat. Just like toss a coin. So the accuracy will be just 50%.
#
# 
# readmore
# ## After Learning
#
# While if we give the cubic box enough **labeled** data, and let it learn **long enough**.
#
# The plain cubic box will become a magic box. It will have high accuracy to tell whether a photo is a cat.
#
# 
# ## How Good a Magic Box is
#
# Of course we want our magic box able to:
#
# - classify a photo is a cat if a coming photo is a cat
# - classify a photo is not a cat if a coming photo is not a cat
#
# We can measure it with following steps:
#
# 1. give the magic box few **new** labeled photos
# 2. the magic box do the classification
# 3. compare the predicted result and expected result to get the accuracy.
#
# > **New** photos, means that the magic box has never see this photo before.
# ## How to Make a Good Magic Box
#
# There are few general ways to get a better magic box.
#
# 1. Give it more photos
# 2. Use better algorithms
# 3. Buy more powerful machines
# 4. Let it learn long enough
#
# We can always letting the box learn long enough. Though it would not be an option most of the time.
#
# We could also buy more powerful machines, if we got enough money. Again, might not an option.
#
# So most of time, we are spending time either getting more data or looking for better algorithms.
# ## Hello World Algorithm - Logistic Regression
#
# Just like any other programming language, there is Hello World.
#
# In supervised learning world, it's Logistic Regression. **Logistics Regression** is a supervised learning algorithm to solve classification problems.
#
# Yes. It's strange, **Logistics Regression** is used to solve classification problems.
#
# If we take cat classification problem, denote
#
# - every pixel as an input feature \\(x\\), denote as \\(x_1, x_2, x_3 ..., x_n\\)
# - every pixel has a weight \\(w\\), denote as \\(w_1, w_2, w_3 ..., w_n\\)
# - a bias value regardless of any pixel \\(b\\)
#
# Logistics Regression use equations
#
# $$z=w_1x_1+w_2x_2+w_3x_3+...+w_nx_n+b$$
#
# $$y=\sigma(z)=\frac 1{1+e^{-z}}$$
#
# By training it long enough, and we get a set of value \\(w_1, w_2, w_3 ..., w_n\\) and \\(b\\).
#
# We will able to calculate the result by substitute the value \\(x_1, x_2, x_3 ..., x_n\\), \\(w_1, w_2, w_3 ..., w_n\\), \\(b\\) with the previous equation.
# ## How About Other Algorithms
#
# So Logistic Regression sounds simple. How about other algorithms?
#
# Theroatically it's just more equoations.
#
# And of course, it will take lots of effort to make it work, and make it work better
#
# Here is an example of a bit complex algorithm. It still able to draw every connections.
#
# 
#
# For modern deep learning, it will not able to draw every connections since there are more than million connections.
# ## Why Machine Learning is Difficult
#
# ### Hard to Understand
#
# One dimension is easy. We can easily figure it out. How about 2, 3, 4 dimensions?
#
# Our brain will not able to plot it when its over 3 dimension. It will be difficult, if we can not make the algorithm a picture in our head
#
# ### Hard to Try
#
# We could just try every combination if
#
# - Combination are in finite space
# - Time cost is small
# - Financial cost is small
#
# While that is not the case for machine learning. It can not be done by brutal force.
#
# ### Hard to Get Enough Cleaning Data
#
# The world is complex. Data could be here, there, in this format, in that format, correct or incorrect.
#
# It takes lots of time and effort to get **clean** data.
#
# If we could not get enough clean data, no matter how good algorithm we have. It will goes to idiom "garbage in, garbage out".
# ## A Little Bit on Linear Algbra
#
# It will not take more than 3 minutes.
#
# 
# 
#
# $$
# \begin{align*}
# 1 * 7 + 2 * 9 + 3 * 11 &= 58\\
# 1 * 8 + 2 * 10 + 3 * 12 &= 64
# \end{align*}
# $$
#
# The first one [2, 3] matrix, and the second one is [3, 2] matrix, the result will be [2, 2] matrix.
#
# In general, if we have a [m, n] matrix dot product [n, o] matrix, the result will be [m, o] matrix.
# ## Vectorize Logistic Regression
#
# Recall the equations used Ligistic Regression
# $$
# \begin{align*}
# z&=w_1x_1+w_2x_2+w_3x_3+...+w_nx_n+b \\
# \hat y=a&=\sigma(z)=\frac 1{1+e^{-z}}
# \end{align*}
# $$
#
# If we set w as [1, dim] matrix, and x as [dim, 1] matrix. We can rewrite the previous equation as.
#
# $$\begin{align*}
# z&=w\cdot x \\
# [1, 1] &\Leftarrow [1, dim] \cdot [dim, 1]
# \end{align*}$$
#
# If we stack all samples of x as [dim, m] matrix. Each column is one example. and stack all labels y together as [1, m]. Z has shape, [dim, m]. We can write the equation of the whole dataset as
#
# $$\begin{align*}
# Z &= w\cdot X\\
# [1, m] &\Leftarrow [1, dim] \cdot [dim, m]
# \end{align*}$$
#
# So after vectorize, we have following parameter and with the shape
#
# |parameter|shape|
# |:--------|:----|
# | X | [dim, m]|
# | Y,A,Z | [1, m]|
# | w | [1, dim]|
#
# ## Implement Logistic Regression Forward Propagation
#
# With the equations we have. We can simply implement **Logistic Regression** with **numpy**, which is a linear algebra library in Python.
#
# We can create a test data and implement logistic regression forward propagation like this
# +
import numpy as np
## Generate test data
dim = 3 # just tested with 3 dimentions
m = 10 # just tested with 10 samples
np.random.seed(1) # set seed, so that we will get predictable value for random
X = np.random.randn(dim, m) # generate random [dim, m] matrix
Y = np.random.randint(2, size=(1, m)) # generate random int 0 or 1, matrix [1, m]
## Initialize parameter
w = np.random.randn(1, dim) # generate inital weight with random value
b = 0.0 # inital bias
## The following two lines are logistic regression forward propagation
Z = np.dot(w, X) + b # dot product w and X, then plus b. numpy will broadcast b
A = 1.0 / (1.0 + np.exp(-Z)) # sigmod function
print(A)
# -
# ## Cost Function
#
# **Lost function** is used to define how close a predict result to expected result.
#
# In logistic regression, lost function for each example defined as
#
# $$ \mathcal{L}(a, y) = - y \log(a) - (1-y) \log(1-a)\tag{8}$$
#
# There is an explanation by <NAME> about [why use this definition](https://www.coursera.org/learn/neural-networks-deep-learning/lecture/SmIbQ/explanation-of-logistic-regression-cost-function-optional).
#
# **Cost function** is used to define how close for all predict result to expected result (**label**). The **cost function** defines as the average lost over all the training examples.
#
# $$ J = \frac{1}{m} \sum_{i=1}^m \mathcal{L}(a^{(i)}, y^{(i)})\tag{9}$$
#
# Recall that A and Y are both is [1, m]. In order to get sum. We can write one line to get cost from A and Y. We will get an [1, 1] matrix just by dot product [1, m] and [m, 1] .
#
# $$J = \frac{1}{m} \bigg(-Y \cdot log(A.T) - (1 - Y) \cdot log(1-A.T) \bigg)$$
#
# And implemented with numpy in one line
cost = (-np.dot(Y, np.log(A.T)) - np.dot(Y, np.log(1 - A.T))) / m
print(np.squeeze(cost))
# ## Gradient Decent
#
# If I'm dropped randomly in a ski resort, it is less likely that I'm dropped off at the lowest bottom.
#
# The gravity will pull you down on the slope when you ski.
#
# | | |
# |:-|:-|
# | | |
#
# We also need get the slope when training our model. By moving on the slope direction towards smaller cost, our model will getting better and better. This slope called gradient decent. It is the derivative of the cost function.
#
# As we define the cost function as
#
# $$ \mathcal{L}(\hat{y}, y^{(i)}) = - y^{(i)} \log(\hat y^{(i)}) - (1-y^{(i)} ) \log(1-\hat y^{(i)})\tag{8}$$
#
# Based on the basic calculus equations.
#
# $$
# \begin{align*}
# \big(f(x)g(x)\big)' &= f(x)g'(x)+f'(x)g(x) \\
# \big(log(x)\big)'&=\frac1x\\
# \frac{dz}{dx}=\frac{dz}{dy}.\frac{dy}{dx}=g'(y)f'(x)&=g'\big(f(x)\big)f'(x) \\
# J &= \frac{1}{m} \sum_{i=1}^m \mathcal{L}(a^{(i)}, y^{(i)})\tag{9}
# \end{align*}
# $$
#
# We can get
#
# $$\begin{align*}
# \frac{\partial \mathcal{L}}{\partial z} &=
# \frac{\partial \mathcal{L}}{\partial \mathcal{\hat y}}\cdot&
# \frac{\partial \hat{y}}{\partial {z}}\\
# &=-(\frac y{\hat y}+\frac{1-y}{1-\hat y}) \cdot&\hat y(1-\hat y)\\
# &=\hat y - y\\
# \\
# \frac{\partial J}{\partial z} &= \frac{1}{m} \sum_{i=1}^m\frac{\partial \mathcal{L}}{\partial z}
# \end{align*} $$
#
#
# $$\begin{align*}
# dw&=\frac{1}{m}(A - Y)\cdot X^T\\
# [1, dim] &\Leftarrow [1, m] \cdot [dim, m]^T \\
# db&=\frac{1}{m}\sum_{i=1}^m(A-Y)
# \end{align*} $$
# With the equation for \\(dw\\) and \\(db\\), we can easily implemented as.
dw = np.dot(A - Y, X.T) / m
db = np.sum(A - Y) / m
print(dw, db)
# ## Backward Propogation
#
# Just like when we are in a mountain, we can easy follow the slope to get to the valley.
# Since we as human is much smaller than the mountain.
#
# How about a giant in the mountain. He might never get to the valley by fixed large step.
#
# 
#
#
# Same apply to machine learning. We need to control the step, which is called learning rate \\(\alpha\\)to avoid to over shooting.
#
# After knowing the gradient decent \\(dw\\), \\(db\\) and controlling the learning rate \\(\alpha\\) we can update the weights \\(w\\) and bias \\(b\\) with following code.
learning_rate = 0.005
w = w - learning_rate * dw
b = b - learning_rate * b
# ## Whole Algorithm
#
# Just like we are in mountain. It's less likely that we will arrive to the valley in one step
#
# The same applies to here, we need iterate many times, with all the previous preparations.
#
# we can write Pseudo code like this.
#
# ```python
# init weight
# init bias
#
# for i in range(number_iterations):
# forward_progation
# calculate cost
# stop iterate if cost already small enough
#
# calculate gradient decent
# update weights and bias
# ```
#
# We can implement the previous pseudo code in two functions.
def propagate(w, b, X, Y):
"""
w: weights, [1, m]
b: bias, scalar value
X: features, [dim, m]
Y: labels, [1, m]
"""
m = X.shape[1]
Z = np.dot(w, X) + b
A = 1.0 / (1.0 + np.exp(-Z))
dw = np.dot(A - Y, X.T) / m
db = np.sum(A - Y) / m
return dw, db, A
def logistic_regression(X, Y, num_iterations=10, learning_rate=0.01):
dim, m = X.shape
w = np.zeros((1, dim)) # Initialize weights to zero
b = 0.0 # Initialize bias to zero
costs = [] # save cost for each iteration
for i in range(num_iterations):
dw, db, A = propagate(w, b, X, Y)
cost = -(np.dot(Y, np.log(A.T)) + np.dot(1-Y, np.log(1-A.T))) / m
# update weights and bias
w = w - learning_rate * dw
b = b - learning_rate * db
if i % 100 == 0:
print(i, cost)
costs.append(cost)
return w, b, costs, A
# ## Get a Data Set
#
# There is a cat data set from [coursera deep learning course](https://www.coursera.org/learn/neural-networks-deep-learning/notebook/zAgPl/logistic-regression-with-a-neural-network-mindset)
#
# The data set is encoded in HDF5 format which is typically used to store numeric data.
#
# The following piece of code is copied from deep learning course to load the cat data
# +
import numpy as np
import h5py
train_dataset = h5py.File('../datasets/train_catvnoncat.h5', "r")
train_set_x_orig = np.array(train_dataset["train_set_x"][:]) # your train set features
train_set_y_orig = np.array(train_dataset["train_set_y"][:]) # your train set labels
test_dataset = h5py.File('../datasets/test_catvnoncat.h5', "r")
test_set_x_orig = np.array(test_dataset["test_set_x"][:]) # your test set features
test_set_y_orig = np.array(test_dataset["test_set_y"][:]) # your test set labels
classes = np.array(test_dataset["list_classes"][:]) # the list of classes
train_set_y = train_set_y_orig.reshape((1, train_set_y_orig.shape[0]))
test_set_y = test_set_y_orig.reshape((1, test_set_y_orig.shape[0]))
# -
# The shape of each sample is [64, 64, 3]. Both height and width are 64.
#
# Each pixel has 3 values for each channel, Red Green and Blue.
# The value range for each channel is from 0 which is darkest, to 255 which is the lightest.
#
# we can use matplotlib plot a sample.
# +
import matplotlib.pyplot as plt
# %matplotlib inline
plt.imshow(train_set_x_orig[10])
print(train_set_x_orig[10].shape)
print(train_set_x_orig[10][0, 0])
# -
# ## Preprocess the Data
#
# A photo has two dimension, x and y. Each point is a pixel. Each pixel in RGB photo has 3 value, Red, Green and Blue. In **Logistic Regression**, we need to convert to one dimension.
#
# **Normalize** data will be an important step. Since machine Learning will typically has better result by normalize the value to range [-1, 1], or [0, 1].
#
# We can pre-process with the following code.
#
# And get dimension 12288 which is 64*64*3. 209 sample for training set and 50 for test set.
# +
m_train = train_set_x_orig.shape[0] # number of train samples
m_test = test_set_x_orig.shape[0] # number of test samples
num_px = train_set_x_orig.shape[1] # number pixel on x and y dimension
train_set_x = train_set_x_orig.reshape(m_train, -1).T / 255 # normalize pixel value to [0, 1]
test_set_x = test_set_x_orig.reshape(m_test, -1).T / 255
print(train_set_x.shape, test_set_x.shape)
# -
# ## Train the Model with Logistic Regression
#
# We can train model with the data set get weights, bias.
w, b, costs, A=logistic_regression(X=train_set_x, Y=train_set_y, num_iterations = 2001, learning_rate = 0.005)
plt.plot(np.squeeze(costs))
plt.ylabel('cost')
plt.xlabel('iterations (per hundreds)')
plt.title("Learning rate =" + str(learning_rate))
plt.show()
costs[:10]
# ## How Good is the Model on the Sample it has Seen
# +
train_predict = np.where(A >= 0.5, 1, 0)
train_accuracy = np.sum(train_predict == train_set_y) / train_set_y.shape[1]
print("train accuracy", train_accuracy)
wrong_index = np.argmax(train_predict != train_set_y)
print("wrong predict on sample ", wrong_index, " to ", train_predict[:, wrong_index], "which should be", train_set_y[:, wrong_index])
plt.imshow(train_set_x_orig[wrong_index])
# -
# Will you say that is a cat? :-(
# ## How Good is the Model on the Sample it has not seen
# +
def predict(w, b, X):
Z = np.dot(w, X)
A = 1.0 / (1.0 + np.exp(-Z))
return np.where(A >= 0.5, 1, 0)
test_predict = predict(w, b, test_set_x)
test_accuracy = np.sum(test_predict == test_set_y) / test_set_y.shape[1]
print("test accuracy", test_accuracy)
wrong_index = np.argmax(test_predict != test_set_y)
print("wrong predict on sample ", wrong_index, " to ", test_predict[:, wrong_index], "which should be", test_set_y[:, wrong_index])
plt.imshow(test_set_x_orig[wrong_index])
# -
# This seems is not a cat. :-(
# ## Why does the Model have Low Accuracy?
#
# With previous logistic regression. We got training set accuracy 99%, and test set accuracy 70%.
#
# 70%, not very impressive, right. But why?
#
# A simple answer is that the model is too simple to catch the characteristics.
#
# We will try to improve it in the next blog.
# ### Reference
#
# 1. [deep learning](https://www.coursera.org/learn/neural-networks-deep-learning)
| notebooks/2018-03-05-Supervised-Learning-Explained-1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#default_exp core
# -
#hide
from nbdev.showdoc import *
# %load_ext autoreload
# %autoreload 2
# + active=""
# #export
#
# import sys,os,re,typing,itertools,operator,functools,math,warnings,functools,io,enum, array
#
# from operator import itemgetter,attrgetter
# from warnings import warn
# from typing import Iterable,Generator,Sequence,Iterator
# from functools import partial,reduce
# from pathlib import Path
#
# try:
# from types import WrapperDescriptorType,MethodWrapperType,MethodDescriptorType
# except ImportError:
# WrapperDescriptorType = type(object.__init__)
# MethodWrapperType = type(object().__str__)
# MethodDescriptorType = type(str.join)
# from types import BuiltinFunctionType,BuiltinMethodType,MethodType,FunctionType,SimpleNamespace
#
# NoneType = type(None)
# string_classes = (str,bytes)
#
# def is_iter(o):
# "Test whether `o` can be used in a `for` loop"
# #Rank 0 tensors in PyTorch are not really iterable
# return isinstance(o, (Iterable,Generator)) and getattr(o,'ndim',1)
#
# def is_coll(o):
# "Test whether `o` is a collection (i.e. has a usable `len`)"
# #Rank 0 tensors in PyTorch do not have working `len`
# return hasattr(o, '__len__') and getattr(o,'ndim',1)
#
# def all_equal(a,b):
# "Compares whether `a` and `b` are the same length and have the same contents"
# if not is_iter(b): return False
# return all(equals(a_,b_) for a_,b_ in itertools.zip_longest(a,b))
#
# def noop (x=None, *args, **kwargs):
# "Do nothing"
# return x
#
# def noops(self, x=None, *args, **kwargs):
# "Do nothing (method)"
# return x
#
# def any_is_instance(t, *args): return any(isinstance(a,t) for a in args)
#
# def isinstance_str(x, cls_name):
# "Like `isinstance`, except takes a type name instead of a type"
# return cls_name in [t.__name__ for t in type(x).__mro__]
#
# def array_equal(a,b):
# if hasattr(a, '__array__'): a = a.__array__()
# if hasattr(b, '__array__'): b = b.__array__()
# return (a==b).all()
#
# def df_equal(a,b): return a.equals(b) if isinstance_str(a, 'NDFrame') else b.equals(a)
#
# def equals(a,b):
# "Compares `a` and `b` for equality; supports sublists, tensors and arrays too"
# if (a is None) ^ (b is None): return False
# if any_is_instance(type,a,b): return a==b
# if hasattr(a, '__array_eq__'): return a.__array_eq__(b)
# if hasattr(b, '__array_eq__'): return b.__array_eq__(a)
# cmp = (array_equal if isinstance_str(a, 'ndarray') or isinstance_str(b, 'ndarray') else
# array_equal if isinstance_str(a, 'Tensor') or isinstance_str(b, 'Tensor') else
# df_equal if isinstance_str(a, 'NDFrame') or isinstance_str(b, 'NDFrame') else
# operator.eq if any_is_instance((str,dict,set), a, b) else
# all_equal if is_iter(a) or is_iter(b) else
# operator.eq)
# return cmp(a,b)
#
# def ipython_shell():
# "Same as `get_ipython` but returns `False` if not in IPython"
# try: return get_ipython()
# except NameError: return False
#
# def in_ipython():
# "Check if code is running in some kind of IPython environment"
# return bool(ipython_shell())
#
# def in_colab():
# "Check if the code is running in Google Colaboratory"
# return 'google.colab' in sys.modules
#
# def in_jupyter():
# "Check if the code is running in a jupyter notebook"
# if not in_ipython(): return False
# return ipython_shell().__class__.__name__ == 'ZMQInteractiveShell'
#
# def in_notebook():
# "Check if the code is running in a jupyter notebook"
# return in_colab() or in_jupyter()
#
# IN_IPYTHON,IN_JUPYTER,IN_COLAB,IN_NOTEBOOK = in_ipython(),in_jupyter(),in_colab(),in_notebook()
#
# def remove_prefix(text, prefix):
# "Temporary until py39 is a prereq"
# return text[text.startswith(prefix) and len(prefix):]
#
# def remove_suffix(text, suffix):
# "Temporary until py39 is a prereq"
# return text[:-len(suffix)] if text.endswith(suffix) else text
#
# -
# ## Export -
#hide
from nbdev.export import notebook2script
notebook2script()
| ignore/00_core.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Overview
#
# I tried to use a CFG feature grammar with lambda calculus to parse a natural language queries into a first order logic
# representation. My plan was to use the FOL as intermediate representation from which I could translate into other query
# languages such as Xpath or SQL. While I was successful in building gramars that achieved this for simple queries
# the translation from FOL to Xpath or SQL proved to difficult in the given time frame. I therefore returned to my first approach
# of parsing NL queries directly into Xpath.
# # Attempting Context Free Feature Grammar with Lambda Calculus
# +
import nltk
from nltk import grammar, parse
sents = ['which planets have a radius of number']
g = """
% start S
S[SEM = <?subj(?vp)>] -> NP[NUM=?n,SEM=?subj] VP[NUM=?n,SEM=?vp]
VP[NUM=?n,SEM=?obj] -> TV[NUM=?n] NP[SEM=?obj]
NP[+INT,NUM=?n,SEM=<?det(?nom)>] -> Det[+INT, NUM=?n,SEM=?det] Nom[NUM=?n,SEM=?nom]
NP[-INT,NUM=?n,SEM=?nom] -> Det[-INT, NUM=?n] Nom[NUM=?n,SEM=?nom]
Nom[NUM=?n,SEM=?nom] -> N[NUM=?n,SEM=?nom]
Nom[NUM=?n,SEM=<?pp(?nom)>] -> N[NUM=?n,SEM=?nom] PP[SEM=?pp]
PP[SEM=?u] -> P[-LOC] UNIT[SEM=?u]
N[NUM=sg,SEM=<\\x.planet(x)>] -> 'planet'
N[NUM=pl,SEM=<\\x.planet(x)>] -> 'planets'
N[NUM=sg,SEM=<\\x y.radius(x,y)>] -> 'radius'
UNIT[SEM=<\\P x.P(x,number)>] -> 'number'
Det[+INT,NUM=pl,SEM=<\\P \\Q.exists x.(P(x) & Q(x))>] -> 'which'
Det[+INT,NUM=sg,SEM=<\\P \\Q.exists x.(P(x) & Q(x))>] -> 'which'
Det[-INT,NUM=sg,SEM=<\\P \\Q.exists x.(P(x) & Q(x))>] -> 'a'
TV[NUM=sg,SEM="",TNS=pres] -> 'has'
TV[NUM=pl,SEM="",TNS=pres] -> 'have'
P[-LOC,SEM=""] -> 'of'
"""
gram = grammar.FeatureGrammar.fromstring(g)
parser = parse.FeatureEarleyChartParser(gram,trace=2)
#trees = list(parser.parse(sents[0].split()))
for results in nltk.interpret_sents(sents, gram):
for (synrep, semrep) in results:
print(semrep)
# +
import nltk
from nltk import grammar, parse
sents = ['which planets have a radius of greater #NUM#']
g = """
% start S
S[SEM = <?subj(?vp)>] -> NP[NUM=?n,SEM=?subj] VP[NUM=?n,SEM=?vp]
VP[NUM=?n,SEM=?obj] -> TV[NUM=?n] NP[SEM=?obj]
NP[+INT,NUM=?n,SEM=<?det(?nom)>] -> Det[+INT, NUM=?n,SEM=?det] Nom[NUM=?n,SEM=?nom]
NP[-INT,NUM=?n,SEM=?nom] -> Det[-INT, NUM=?n] Nom[NUM=?n,SEM=?nom]
Nom[NUM=?n,SEM=?nom] -> N[NUM=?n,SEM=?nom]
Nom[NUM=?n,SEM=<?nom(?pp)>] -> N[NUM=?n,SEM=?nom] PP[SEM=?pp]
Nom[SEM=<?u(?a)>] -> A[SEM=?a] UNIT[SEM=?u]
PP[SEM=?nom] -> P[-LOC] Nom[SEM=?nom]
N[NUM=sg,SEM=<\\x.planet(x)>] -> 'planet'
N[NUM=pl,SEM=<\\x.planet(x)>] -> 'planets'
N[NUM=sg,SEM=<\\P x .exists y.(radius(x,y) & P(y))>] -> 'radius'
UNIT[SEM=<\\P.P(number)>] -> '#NUM#'
A[SEM=<\\x y.greater(x,y)>] -> 'greater'
Det[+INT,NUM=pl,SEM=<\\P \\Q.exists x.(P(x) & Q(x))>] -> 'which'
Det[+INT,NUM=sg,SEM=<\\P \\Q.exists x.(P(x) & Q(x))>] -> 'which'
Det[-INT,NUM=sg,SEM=<\\P \\Q.exists x.(P(x) & Q(x))>] -> 'a'
TV[NUM=sg,SEM="",TNS=pres] -> 'has'
TV[NUM=pl,SEM="",TNS=pres] -> 'have'
P[-LOC,SEM=""] -> 'of'
"""
gram = grammar.FeatureGrammar.fromstring(g)
parser = parse.FeatureEarleyChartParser(gram,trace=0)
trees = list(parser.parse(sents[0].split()))
for results in nltk.interpret_sents(sents, gram):
for (synrep, semrep) in results:
print(semrep)
# +
import nltk
from nltk import grammar, parse
sents = ['which planets have a radius of greater #NUM#']
g = """
% start S
S[SEM = <?subj(?vp)>] -> NP[NUM=?n,SEM=?subj] VP[NUM=?n,SEM=?vp]
VP[NUM=?n,SEM=<?v(?obj)>] -> TV[NUM=?n,SEM=?v] NP[SEM=?obj]
NP[+INT,NUM=?n,SEM=<?det(?nom)>] -> Det[+INT, NUM=?n,SEM=?det] Nom[NUM=?n,SEM=?nom]
NP[-INT,NUM=?n,SEM=<?det(?nom)>] -> Det[-INT, NUM=?n,SEM=?det] Nom[NUM=?n,SEM=?nom]
Nom[NUM=?n,SEM=?nom] -> N[NUM=?n,SEM=?nom]
Nom[NUM=?n,SEM=<?pp(?nom)>] -> N[NUM=?n,SEM=?nom] PP[SEM=?pp]
Nom[SEM=<?u(?a)>] -> A[SEM=?a] UNIT[SEM=?u]
Nom[SEM=?a] -> A[SEM=?a]
PP[SEM=<?p(?nom)>] -> P[-LOC,SEM=?p] Nom[SEM=?nom]
N[NUM=sg,SEM=<\\x.planet(x)>] -> 'planet'
N[NUM=pl,SEM=<\\x.planet(x)>] -> 'planets'
N[NUM=sg,SEM=<\\x.radius(x)>] -> 'radius'
UNIT[SEM=<\\P.P(number)>] -> '#NUM#'
A[SEM=<\\x.great(x)>] -> 'great'
A[SEM=<\\x y.greater(x,y)>] -> 'greater'
Det[+INT,NUM=pl,SEM=<\\P \\Q.exists x.(P(x) & Q(x))>] -> 'which'
Det[+INT,NUM=sg,SEM=<\\P \\Q.exists x.(P(x) & Q(x))>] -> 'which'
Det[-INT,NUM=sg,SEM=<\\P \\Q.exists x.(P(x) & Q(x))>] -> 'a'
TV[NUM=sg,TNS=pres,SEM=<\\X y.(X(\\x.have(y, x)))>] -> 'has'
TV[NUM=pl,TNS=pres,SEM=<\\X y.(X(\\x.have(y, x)))>] -> 'have'
P[-LOC,SEM=<\\P \\Q x.(P(x) & Q(x))>] -> 'of'
"""
gram = grammar.FeatureGrammar.fromstring(g)
parser = parse.FeatureEarleyChartParser(gram,trace=2)
#trees = list(parser.parse(sents[0].split()))
for results in nltk.interpret_sents(sents, gram):
for (synrep, semrep) in results:
print(semrep)
# +
import nltk
from nltk import grammar, parse
sents = ['which planets have a radius of greater #NUM#']
g = """
% start S
S[SEM = <?subj(?vp)>] -> NP[NUM=?n,SEM=?subj] VP[NUM=?n,SEM=?vp]
VP[NUM=?n,SEM=<?v(?obj)>] -> TV[NUM=?n,SEM=?v] NP[SEM=?obj]
NP[+INT,NUM=?n,SEM=<?det(?nom)>] -> Det[+INT, NUM=?n,SEM=?det] Nom[NUM=?n,SEM=?nom]
NP[-INT,NUM=?n,SEM=<?det(?nom)>] -> Det[-INT, NUM=?n,SEM=?det] Nom[NUM=?n,SEM=?nom]
Nom[NUM=?n,SEM=?nom] -> N[NUM=?n,SEM=?nom]
Nom[NUM=?n,SEM=<?pp(?nom)>] -> N[NUM=?n,SEM=?nom] PP[SEM=?pp]
Nom[SEM=<?u(?a)>] -> A[SEM=?a] UNIT[SEM=?u]
Nom[SEM=?a] -> A[SEM=?a]
PP[SEM=<?p(?nom)>] -> P[-LOC,SEM=?p] Nom[SEM=?nom]
N[NUM=sg,SEM=<\\x.planet(x)>] -> 'planet'
N[NUM=pl,SEM=<\\x.planet(x)>] -> 'planets'
N[NUM=sg,SEM=<\\x.radius(x)>] -> 'radius'
UNIT[SEM=<\\P.P(number)>] -> '#NUM#'
A[SEM=<\\x.great(x)>] -> 'great'
A[SEM=<\\x y.greater(x,y)>] -> 'greater'
Det[+INT,NUM=pl,SEM=<\\P \\Q.exists x.(P(x) & Q(x))>] -> 'which'
Det[+INT,NUM=sg,SEM=<\\P \\Q.exists x.(P(x) & Q(x))>] -> 'which'
Det[-INT,NUM=sg,SEM=<\\P \\Q.exists x.(P(x) & Q(x))>] -> 'a'
TV[NUM=sg,TNS=pres,SEM=<\\X y.(X(\\x.have(y, x)))>] -> 'has'
TV[NUM=pl,TNS=pres,SEM=<\\X y.(X(\\x.have(y, x)))>] -> 'have'
P[-LOC,SEM=<\\P \\Q x.(P(x) & Q(x))>] -> 'of'
"""
gram = grammar.FeatureGrammar.fromstring(g)
parser = parse.FeatureEarleyChartParser(gram,trace=2)
#trees = list(parser.parse(sents[0].split()))
for results in nltk.interpret_sents(sents, gram):
for (synrep, semrep) in results:
print(semrep)
# +
import nltk
from nltk import grammar, parse
sents = ['which planets have a radius of greater #NUM#']
g = """
% start S
S[SEM = <?subj(?vp)>] -> NP[NUM=?n,SEM=?subj] VP[NUM=?n,SEM=?vp]
VP[NUM=?n,SEM=<?v(?obj)>] -> TV[NUM=?n,SEM=?v] NP[SEM=?obj]
NP[+INT,NUM=?n,SEM=<?det(?nom)>] -> Det[+INT, NUM=?n,SEM=?det] Nom[NUM=?n,SEM=?nom]
NP[-INT,NUM=?n,SEM=<?det(?nom)>] -> Det[-INT, NUM=?n,SEM=?det] Nom[NUM=?n,SEM=?nom]
Nom[NUM=?n,SEM=?nom] -> N[NUM=?n,SEM=?nom]
Nom[NUM=?n,SEM=<?pp(?nom)>] -> N[NUM=?n,SEM=?nom] PP[SEM=?pp]
Nom[SEM=<?u(?a)>] -> A[SEM=?a] UNIT[SEM=?u]
Nom[SEM=?a] -> A[SEM=?a]
PP[SEM=<?p(?nom)>] -> P[-LOC,SEM=?p] Nom[SEM=?nom]
N[NUM=sg,SEM=<\\x.planet(x)>] -> 'planet'
N[NUM=pl,SEM=<\\x.planet(x)>] -> 'planets'
N[NUM=sg,SEM=<\\x.radius(x)>] -> 'radius'
UNIT[SEM=<\\P.P(number)>] -> '#NUM#'
A[SEM=<\\x.great(x)>] -> 'great'
A[SEM=<\\x y.greater(x,y)>] -> 'greater'
Det[+INT,NUM=pl,SEM=<\\P \\Q.exists x.(P(x) & Q(x))>] -> 'which'
Det[+INT,NUM=sg,SEM=<\\P \\Q.exists x.(P(x) & Q(x))>] -> 'which'
Det[-INT,NUM=sg,SEM=<\\P \\Q.exists x.(P(x) & Q(x))>] -> 'a'
TV[NUM=sg,TNS=pres,SEM=<\\X y.(X(\\x.have(y, x)))>] -> 'has'
TV[NUM=pl,TNS=pres,SEM=<\\X y.(X(\\x.have(y, x)))>] -> 'have'
P[-LOC,SEM=<\\P \\Q x.(P(x) & Q(x))>] -> 'of'
"""
gram = grammar.FeatureGrammar.fromstring(g)
parser = parse.FeatureEarleyChartParser(gram,trace=2)
#trees = list(parser.parse(sents[0].split()))
for results in nltk.interpret_sents(sents, gram):
for (synrep, semrep) in results:
print(semrep)
# -
# # Returning to the Simpler Approach
#
# I decided to build the Xpath queries directly from the parse trees. However, I refined the CFG to be based on X-Bar syntax and strictly in Chomsky Normal Form. Here a first attempt with example syntactic tree:
from nltk import grammar, parse
import nltk,urllib.request, gzip, io
from nltk import grammar, parse
from nltk.tokenize import word_tokenize
import re
from lxml import etree
# +
sents = ['what planets have a mass of 19.4','which planets have a mass of 19.4',
'what planets have a radius of 0.188','which planets have a mass of at least 19.4',
'which planets have a mass of at most 0.001','which planets have a mass smaller than 0.001',
'which planets have a mass greater than 19.4','what planets were discovered before 2001',
'what planets were discovered after 2021','which planets have a mass smaller than 0.001',
'which planets have a mass smaller than 0.001 and a radius of less than 0.188',
'what planets have a mass larger than 0.001 and a radius of at least 0.188 and a mass of at least 0.188',
'what planets have a mass larger than 0.001 and have a radius of at least 0.188']
g = """
% start S
S[SEM=(?n + '[' + ?v+']')] -> N[BAR=2,SEM=?n] V[BAR=2,SEM=?v]
V[BAR=2,SEM=(?v + ?n)] -> V[BAR=1,SEM=?v] N[BAR=2,SEM=?n]
V[BAR=2,SEM=(?v + ?p)] -> V[BAR=1,SEM=?v] P[BAR=2,SEM=?p]
V[BAR=2,SEM=(?v + ?c)] -> V[BAR=1,SEM=?v] CONJ[BAR=2,SEM=?c]
V[BAR=1,SEM=?p] -> AUX PART[SEM=?p]
V[BAR=1,SEM=?v] -> V[BAR=0,SEM=?v]
CONJ[BAR=2,SEM=(?n + ?c)] -> N[BAR=2,SEM=?n] CONJ[BAR=1,SEM=?c]
CONJ[BAR=2,SEM=(?v + ?c)] -> V[BAR=2,SEM=?v] CONJ[BAR=1,SEM=?c]
CONJ[BAR=1,SEM=(?c+ ?cp)] -> CONJ[BAR=0,SEM=?c] CONJ[BAR=2,SEM=?cp]
CONJ[BAR=1,SEM=(?c + ?n)] -> CONJ[BAR=0,SEM=?c] N[BAR=2,SEM=?n]
CONJ[BAR=1,SEM=(?c + ?v)] -> CONJ[BAR=0,SEM=?c] V[BAR=2,SEM=?v]
N[BAR=2,SEM=(?det + ?n)] -> Art[SEM=?det] N[BAR=1,SEM=?n] | Int[SEM=?det] N[BAR=1,SEM=?n]
N[BAR=2,SEM=(?a + ?n)] -> A[BAR=2,SEM=?a] N[BAR=1,SEM=?n]
N[BAR=1,SEM=(?n + ?p)] -> N[BAR=0,SEM=?n] P[-TIME, BAR=2,SEM=?p]
N[BAR=1,SEM=(?n + ?a)] -> N[BAR=0,SEM=?n] A[BAR=2,SEM=?a]
N[BAR=2,SEM=?n] -> N[BAR=1,SEM=?n]
N[BAR=1,SEM=?n] -> N[BAR=0,SEM=?n]
P[BAR=2,SEM=(?p + ?n)] -> P[BAR=1,SEM=?p] A[BAR=2,SEM=?n]
P[BAR=2,SEM=(?p + ?n)] -> P[BAR=1,SEM=?p] N[BAR=2,SEM=?n]
P[BAR=1,SEM=?p] -> P[BAR=0,SEM=?p]
A[BAR=2,SEM=(?a+?n)] -> A[BAR=1,SEM=?a] N[BAR=1,SEM=?n]
A[BAR=1,SEM=?a] -> A[BAR=0,SEM=?a]
A[BAR=1,SEM=?a] -> A[BAR=0,SEM=?a] CONJ[BAR=0]
A[BAR=1,SEM=?a] -> P[BAR=1] A[BAR=1,SEM=?a]
PART[SEM='discoveryyear'] -> 'discovered'
Int[SEM='.//'] -> 'which' | 'what'
AUX -> 'were'
V[BAR=0,SEM=''] -> 'have' | 'possess'
Art[SEM=''] -> 'a'
N[BAR=0,SEM='mass'] -> 'mass'
N[BAR=0,SEM='radius'] -> 'radius'
N[BAR=0,SEM='planet'] -> 'planets'
N[BAR=0,SEM="=#NUM#"] -> '#NUM#'
P[-TIME, BAR=0,SEM=''] -> 'of' |'at'
P[-TIME, BAR=0,SEM='<'] -> 'before'
P[+TIME, BAR=0,SEM='>'] -> 'after'
CONJ[BAR=0,SEM=''] -> 'than'
CONJ[BAR=0,SEM=' and '] -> 'and'
A[BAR=0,SEM='>'] -> 'bigger' | 'larger' | 'greater' | 'more' | 'least'
A[BAR=0,SEM='<'] -> 'smaller' | 'less' | 'most'
"""
sents = [re.sub(r'\d+\.*\d*','#NUM#',s).split() for s in sents]
gram = grammar.FeatureGrammar.fromstring(g)
parser = parse.FeatureEarleyChartParser(gram)
trees = [list(parser.parse(s)) for _,s in enumerate(sents)]
#trees = list(parser.parse(sents[0].split()))
from nltk.draw.tree import draw_trees
draw_trees(trees[-1][0])
# -
# The (almost) finished version has more rules and tow different sentence types. It becomes apparent that the number of trees is exploding fast.
# I am printing the results of the parser at the end of this code chunk:
# +
g = """
% start S
S[SEM=?s] -> S1[SEM=?s] | S2[SEM=?s]
S1[SEM=('(.//' + ?n + ')' + '[position()<' + ?count + ']')] -> N[BAR=2,NUM=?num,SEM=?n,COUNT=?count]
S1[NUM=?num,SEM=(?s+ ';' + ?c)] -> S1[NUM=?num,SEM=?s] CONJ[BAR=1,SEM=?c]
S2[SEM=(?n + '[' + ?v+']'),NUM=?num] -> N[BAR=2,SEM=?n,NUM=?num] V[BAR=2,SEM=?v,NUM=?num]
V[BAR=2,SEM=(?v + ?n),NUM=?num] -> V[BAR=1,SEM=?v,NUM=?num] N[BAR=2,SEM=?n]
V[BAR=2,SEM=(?v + ?p),NUM=?num] -> V[BAR=1,SEM=?v,NUM=?num] P[BAR=2,SEM=?p,SUBC=?subc]
V[BAR=2,SEM=(?v + ?c),NUM=?num] -> V[BAR=1,SEM=?v,NUM=?num] CONJ[BAR=2,SEM=?c]
V[BAR=1,SEM=?p] -> AUX PART[SEM=?p]
V[BAR=1,SEM=?v,NUM=?num] -> V[BAR=0,SEM=?v,NUM=?num]
C[BAR=2,SEM=?v,NUM=?num] -> C[BAR=1] V[BAR=2,SEM=?v,NUM=?num]
C[BAR=2,SEM=?c] -> C[BAR=1] CONJ[BAR=2,SEM=?c]
C[BAR=1] -> C[BAR=0]
CONJ[BAR=2,SEM=(?n + ?c)] -> N[BAR=2,SEM=?n] CONJ[BAR=1,SEM=?c]
CONJ[BAR=2,SEM=(?v + ?c)] -> V[BAR=2,SEM=?v] CONJ[BAR=1,SEM=?c]
CONJ[BAR=2,SEM=(?p + ?c)] -> P[BAR=2,SEM=?p,SUBC=?subc] CONJ[BAR=1,SEM=?c]
CONJ[BAR=2,SEM=(?c + ?con)] -> C[BAR=2,SEM=?c] CONJ[BAR=1,SEM=?con]
CONJ[BAR=1,SEM=(?c+ ?cp)] -> CONJ[BAR=0,SEM=?c] CONJ[BAR=2,SEM=?cp]
CONJ[BAR=1,SEM=(?c + ?n)] -> CONJ[BAR=0,SEM=?c] N[BAR=2,SEM=?n]
CONJ[BAR=1,SEM=(?c + ?n)] -> CONJ[BAR=0,SEM=?c] N[BAR=2,SEM=?n]
CONJ[BAR=1,SEM=(?c + ?v)] -> CONJ[BAR=0,SEM=?c] V[BAR=2,SEM=?v]
CONJ[BAR=1,SEM=(?c + ?v)] -> CONJ[BAR=0,SEM=?c] P[BAR=2,SEM=?v,SUBC=?subc]
CONJ[BAR=1,SEM=(?con + ?c)] -> CONJ[BAR=0,SEM=?con] C[BAR=2,SEM=?c]
CONJ[BAR=1,SEM=?s] -> CONJ[BAR=0,SEM=?c] S[SEM=?s]
N[BAR=2,NUM=?num,SEM=(?det + ?n),COUNT='=count(//*)'] -> Art[NUM=?num,SEM=?det] N[BAR=1,NUM=?num,SEM=?n] | Int[NUM=?num,SEM=?det] N[BAR=1,NUM=?num,SEM=?n]
N[BAR=2,NUM=?num,SEM=?n,COUNT=?count] -> Num[SEM=?count] N[BAR=1,NUM=?num,SEM=?n]
N[BAR=2,SEM=(?n + ' and ' + ?c),NUM=?num,COUNT='=count(//*)'] -> N[BAR=2,SEM=?n,NUM=?num] C[BAR=2,SEM=?c,NUM=?num]
N[BAR=2,SEM=(?a + ?n)] -> A[BAR=2,SEM=?a] N[BAR=1,SEM=?n]
N[BAR=1,SEM=(?n + ?p)] -> N[BAR=0,SEM=?n] P[BAR=2,SEM=?p,SUBC='-Adj']
N[BAR=1,SEM=(?n + '[' + ?p +']')] -> N[BAR=0,SEM=?n] P[BAR=2,SEM=?p,SUBC='+Adj']
N[BAR=1,SEM=(?n + '[' + ?c +']')] -> N[BAR=0,SEM=?n] C[BAR=2,SEM=?c]
N[BAR=1,SEM=(?n + ?a)] -> N[BAR=0,SEM=?n] A[BAR=2,SEM=?a]
N[BAR=1,NUM=?num,SEM=?n] -> N[BAR=0,NUM=?num,SEM=?n]
P[BAR=2,SEM=(?p + ?n),SUBC=?subc] -> P[BAR=1,SEM=?p,SUBC=?subc] A[BAR=2,SEM=?n]
P[BAR=2,SEM=(?p + ?n),SUBC=?subc] -> P[BAR=1,SEM=?p,SUBC=?subc] N[BAR=2,SEM=?n]
P[BAR=2,SEM=(?p + ?n),SUBC=?subc] -> P[BAR=1,SEM=?p,SUBC=?subc] Num[SEM=?n]
P[BAR=2,SEM=(?p + ?c),SUBC=?subc] -> P[BAR=1,SEM=?p,SUBC=?subc] CONJ[BAR=2,SEM=?c]
P[BAR=1,SEM=?p,SUBC=?subc] -> P[BAR=0,SEM=?p,SUBC=?subc]
A[BAR=2,SEM=(?a+?n)] -> A[BAR=1,SEM=?a] Num[SEM=?n]
A[BAR=1,SEM=?a] -> A[BAR=0,SEM=?a]
A[BAR=1,SEM=?a] -> A[BAR=0,SEM=?a] CONJ[BAR=0]
A[BAR=1,SEM=?a] -> P[BAR=1] A[BAR=1,SEM=?a]
PART[SEM='discoveryyear'] -> 'discovered'
Int[NUM='sg',SEM='.//'] -> 'which' | 'what'
Int[NUM='pl',SEM='.//'] -> 'which' | 'what'
AUX[NUM='pl'] -> 'were'
AUX[NUM='sg'] -> 'was'
V[NUM='pl',BAR=0,SEM=''] -> 'have' | 'possess'
V[NUM='sg',BAR=0,SEM=''] -> 'has' | 'possesses'
Art[NUM='sg',SEM=''] -> 'a' | 'any'
Art[NUM='pl',SEM=''] -> | 'any'
N[BAR=0,NUM='sg',SEM='mass'] -> 'mass'
N[BAR=0,NUM='sg',SEM='radius'] -> 'radius'
N[NUM='pl',BAR=0,SEM='planet'] -> 'planets'
N[NUM='sg',BAR=0,SEM='planet'] -> 'planet'
Num[NUM='na',SEM="=#NUM#"] -> '#NUM#'
Num[NUM='na',SEM="=#NUM0"] -> '#NUM0'
Num[NUM='na',SEM="=#NUM1"] -> '#NUM1'
Num[NUM='na',SEM="=#NUM2"] -> '#NUM2'
Num[NUM='na',SEM="=#NUM3"] -> '#NUM3'
P[BAR=0,SEM='',SUBC='-Adj'] -> 'of' |'at'
P[BAR=0,SEM='<',SUBC='-Adj'] -> 'before'
P[BAR=0,SEM='>',SUBC='-Adj'] -> 'after'
P[BAR=0,SEM='',SUBC='+Adj'] -> 'with'
C[BAR=0] -> 'that' | 'which'
CONJ[BAR=0,SEM=''] -> 'than'
CONJ[BAR=0,SEM=' and '] -> 'and'
A[BAR=0,SEM='>'] -> 'bigger' | 'larger' | 'greater' | 'more' | 'least'
A[BAR=0,SEM='<'] -> 'smaller' | 'less' | 'most' | 'maximally'
"""
queries = ['planets with a mass of 19.4','planets with a radius of 1',
'planets with a radius of 1 and with a mass of 19.4',
'planets with a mass of 19.4 and planets with a radius of 1',
'planets with a mass of 19.4 and with a radius of 1 and planets with a radius of 1',
'planets that were discovered before 2002','planets with a radius of 1 that were discovered before 2015',
'what planets have a mass of 19.4','which planets have a mass of 19.4',
'what planets have a radius of 0.188','which planets have a mass of at least 19.4',
'which planets have a mass of at most 0.001','which planets have a mass smaller than 0.001',
'which planets have a mass greater than 19.4','what planets were discovered before 2001',
'what planets were discovered after 2021','which planets have a mass smaller than 0.001',
'which planets have a mass smaller than 0.001 and a radius of less than 0.188',
'what planets have a mass larger than 0.001 and a radius of at least 0.188 and a mass of at least 0.188',
'what planets have a mass larger than 0.001 and have a radius of at least 0.188','5 planets with a mass larger than 1 and 1 planet with a mass larger than 2',
'planets with a radius of 1 and with a mass of 19.4','planets that were discovered before 2001 and have a mass greater than 3']
gram = grammar.FeatureGrammar.fromstring(g)
parser = parse.FeatureEarleyChartParser(gram)
for query in queries:
query = re.sub(r'\d+\.*\d*','#NUM#',query)
print('\n',query)
trees = list(parser.parse(query.split()))
for i,t in enumerate(trees):
answer = trees[i].label()['SEM']
print(i,answer)
# -
# # Putting it together with German
#
# I now added in rules for German. For the most part, the rules for English could be reused because the syntactic patterns are similar for the queries that I use. The terminal nodes have to be duplicated and a 'de' tag has to be added to those. The language feature is added to all nodes such that it can later be used to select language-specific responses by the dialogue agent. A complicating factor is that German uses verb-final word order in relative clauses (e.g., 'a planet that a radius of 1 has') and I had to create a German-specific feature +VFINAL and introduce additional rules making use of this feature. The final Grammar looked like this:
# +
import re
g = """
% start S
S[SEM=?s,NUM=?num,L=?l] -> S1[SEM=?s,NUM=?num,L=?l] | S2[SEM=?s,NUM=?num,L=?l]
S1[SEM=('(.//' + ?n + ')' + '[position()<' + ?count + ']'),L=?l] -> N[BAR=2,NUM=?num,SEM=?n,COUNT=?count,L=?l]
S1[NUM=?num,SEM=(?s+ ';' + ?c),L=?l] -> S1[NUM=?num,SEM=?s,L=?l] CONJ[BAR=1,SEM=?c,L=?l]
S2[SEM=(?n + '[' + ?v+']'),NUM=?num,L=?l] -> N[BAR=2,SEM=?n,NUM=?num,L=?l] V[BAR=2,SEM=?v,NUM=?num,L=?l]
S2[NUM=?num,SEM=(?s+ ';' + ?c),L=?l] -> S2[NUM=?num,SEM=?s,L=?l] CONJ[BAR=1,SEM=?c,L=?l]
V[BAR=2,SEM=(?v + ?n),NUM=?num,L=?l] -> V[BAR=1,SEM=?v,NUM=?num,L=?l] N[BAR=2,SEM=?n,L=?l]
V[BAR=2,SEM=(?v + ?n),NUM=?num,L='de',+VFINAL] -> N[BAR=2,SEM=?n] V[BAR=1,SEM=?v,NUM=?num,L='de']
V[BAR=2,SEM=(?v + ?p),NUM=?num,L=?l] -> V[BAR=1,SEM=?v,NUM=?num,L=?l] P[BAR=2,SEM=?p,SUBC=?subc,L=?l]
V[BAR=2,SEM=(?v + ?p),NUM=?num,L='de',+VFINAL] -> P[BAR=2,SEM=?p,SUBC=?subc,L='de'] V[BAR=1,SEM=?v,NUM=?num,L='de',+VFINAL]
V[BAR=2,SEM=(?v + ?c),NUM=?num,L=?l] -> V[BAR=1,SEM=?v,NUM=?num,L=?l] CONJ[BAR=2,SEM=?c,L=?l]
V[BAR=1,SEM=?p] -> AUX PART[SEM=?p,L=?l]
V[BAR=1,SEM=?p,L='de',+VFINAL] -> PART[SEM=?p,L='de'] AUX[L='de']
V[BAR=1,SEM='',L='de'] -> AUX[L='de']
V[BAR=1,SEM=?v,NUM=?num,L=?l] -> V[BAR=0,SEM=?v,NUM=?num,L=?l]
C[BAR=2,SEM=?v,NUM=?num,L=?l] -> C[BAR=1,L=?l] V[BAR=2,SEM=?v,NUM=?num,L=?l]
C[BAR=2,SEM=?v,NUM=?num,L='de'] -> C[BAR=1,L='de'] V[BAR=2,SEM=?v,NUM=?num,L='de',+VFINAL]
C[BAR=2,SEM=?c,L=?l] -> C[BAR=1,L=?l] CONJ[BAR=2,SEM=?c,L=?l]
C[BAR=1,L=?l] -> C[BAR=0,L=?l]
CONJ[BAR=2,SEM=(?n + ?c),L=?l] -> N[BAR=2,SEM=?n,L=?l] CONJ[BAR=1,SEM=?c,L=?l]
CONJ[BAR=2,SEM=(?v + ?c),L=?l] -> V[BAR=2,SEM=?v,L=?l] CONJ[BAR=1,SEM=?c,L=?l]
CONJ[BAR=2,SEM=(?p + ?c),L=?l] -> P[BAR=2,SEM=?p,SUBC=?subc,L=?l] CONJ[BAR=1,SEM=?c,L=?l]
CONJ[BAR=2,SEM=(?c + ?con),L=?l] -> C[BAR=2,SEM=?c,L=?l] CONJ[BAR=1,SEM=?con,L=?l]
CONJ[BAR=1,SEM=(?c+ ?cp),L=?l] -> CONJ[BAR=0,SEM=?c,L=?l] CONJ[BAR=2,SEM=?cp,L=?l]
CONJ[BAR=1,SEM=(?c + ?n),L=?l] -> CONJ[BAR=0,SEM=?c,L=?l] N[BAR=2,SEM=?n,L=?l]
CONJ[BAR=1,SEM=(?c + ?n),L=?l] -> CONJ[BAR=0,SEM=?c,L=?l] N[BAR=2,SEM=?n,L=?l]
CONJ[BAR=1,SEM=(?c + ?v),L=?l] -> CONJ[BAR=0,SEM=?c,L=?l] V[BAR=2,SEM=?v,L=?l]
CONJ[BAR=1,SEM=(?c + ?v),L=?l] -> CONJ[BAR=0,SEM=?c,L=?l] P[BAR=2,SEM=?v,SUBC=?subc,L=?l]
CONJ[BAR=1,SEM=(?con + ?c),L=?l] -> CONJ[BAR=0,SEM=?con,L=?l] C[BAR=2,SEM=?c,L=?l]
CONJ[BAR=1,SEM=?s,L=?l] -> CONJ[BAR=0,SEM=?c,L=?l] S[SEM=?s,L=?l]
N[BAR=2,NUM=?num,SEM=(?det + ?n),COUNT='=count(//*)',L=?l] -> Art[NUM=?num,SEM=?det,L=?l] N[BAR=1,NUM=?num,SEM=?n,L=?l] | Int[NUM=?num,SEM=?det,L=?l] N[BAR=1,NUM=?num,SEM=?n,L=?l]
N[BAR=2,NUM=?num,SEM=?n,COUNT=?count,L=?l] -> Num[SEM=?count,L=?l] N[BAR=1,NUM=?num,SEM=?n,L=?l]
N[BAR=2,SEM=(?n + ' and ' + ?c),NUM=?num,COUNT='=count(//*)',L=?l] -> N[BAR=2,SEM=?n,NUM=?num,L=?l] C[BAR=2,SEM=?c,NUM=?num,L=?l]
N[BAR=2,SEM=(?a + ?n),L=?l] -> A[BAR=2,SEM=?a,L=?l] N[BAR=1,SEM=?n,L=?l]
N[BAR=1,SEM=(?n + ?p),L=?l] -> N[BAR=0,SEM=?n,L=?l] P[BAR=2,SEM=?p,SUBC='-Adj',L=?l]
N[BAR=1,SEM=(?n + '[' + ?p +']'),L=?l] -> N[BAR=1,SEM=?n,L=?l] P[BAR=2,SEM=?p,SUBC='+Adj',L=?l]
N[BAR=1,SEM=(?n + '[' + ?c +']'),L=?l] -> N[BAR=0,SEM=?n,L=?l] C[BAR=2,SEM=?c,L=?l]
N[BAR=1,SEM=(?n + ?a),L=?l] -> N[BAR=0,SEM=?n,L=?l] A[BAR=2,SEM=?a,L=?l]
N[BAR=1,NUM=?num,SEM=?n,L=?l] -> N[BAR=0,NUM=?num,SEM=?n,L=?l]
P[BAR=2,SEM=(?p + ?n),SUBC=?subc,L=?l] -> P[BAR=1,SEM=?p,SUBC=?subc,L=?l] A[BAR=2,SEM=?n,L=?l]
P[BAR=2,SEM=(?p + ?n),SUBC=?subc,L=?l] -> P[BAR=1,SEM=?p,SUBC=?subc,L=?l] N[BAR=2,SEM=?n,L=?l]
P[BAR=2,SEM=(?p + ?n),SUBC=?subc,L=?l] -> P[BAR=1,SEM=?p,SUBC=?subc,L=?l] Num[SEM=?n,L=?l]
P[BAR=2,SEM=(?s + ?p + ?n),SUBC=?subc,L='de'] -> P[BAR=1,SEM=?p,SUBC=?subc,L='de'] Num[SEM=?n] PART[SEM=?s,L='de']
P[BAR=2,SEM=(?p + ?c),SUBC=?subc,L=?l] -> P[BAR=1,SEM=?p,SUBC=?subc,L=?l] CONJ[BAR=2,SEM=?c,L=?l]
P[BAR=1,SEM=?p,SUBC=?subc,L=?l] -> P[BAR=0,SEM=?p,SUBC=?subc,L=?l]
A[BAR=2,SEM=(?a+?n),L=?l] -> A[BAR=1,SEM=?a,L=?l] Num[SEM=?n,L=?l]
A[BAR=1,SEM=?a,L=?l] -> A[BAR=0,SEM=?a,L=?l]
A[BAR=1,SEM=?a,L=?l] -> A[BAR=0,SEM=?a,L=?l] CONJ[BAR=0,L=?l]
A[BAR=1,SEM=?a,L=?l] -> P[BAR=1,L=?l] A[BAR=1,SEM=?a,L=?l]
PART[SEM='discoveryyear',L='en'] -> 'discovered'
Int[NUM='sg',SEM='.//',L='en'] -> 'which' | 'what'
Int[NUM='pl',SEM='.//',L='en'] -> 'which' | 'what'
AUX[NUM='pl',L='en'] -> 'were'
AUX[NUM='sg',L='en'] -> 'was'
V[NUM='pl',BAR=0,SEM='',L='en'] -> 'have' | 'possess'
V[NUM='sg',BAR=0,SEM='',L='en'] -> 'has' | 'possesses'
Art[NUM='sg',SEM='',L='en'] -> 'a' | 'any' | 'an'
Art[NUM='pl',SEM='',L='en'] -> | 'any'
N[BAR=0,NUM='sg',SEM='mass',L='en'] -> 'mass'
N[BAR=0,NUM='sg',SEM='radius',L='en'] -> 'radius'
N[BAR=0,NUM='sg',SEM='age',L='en'] -> 'age'
N[BAR=0,NUM='sg',SEM='temperature',L='en'] -> 'temperature'
N[NUM='pl',BAR=0,SEM='planet',L='en'] -> 'planets'
N[NUM='sg',BAR=0,SEM='planet',L='en'] -> 'planet'
Num[NUM='na',SEM="=#NUM#"] -> '#num#'
Num[NUM='na',SEM="=#NUM0"] -> '#num0'
Num[NUM='na',SEM="=#NUM1"] -> '#num1'
Num[NUM='na',SEM="=#NUM2"] -> '#num2'
Num[NUM='na',SEM="=#NUM3"] -> '#num3'
Num[NUM='na',SEM="=#NUM4"] -> '#num4'
Num[NUM='na',SEM="=#NUM5"] -> '#num5'
Num[NUM='na',SEM="=#NUM6"] -> '#num6'
Num[NUM='na',SEM="=#NUM7"] -> '#num7'
Num[NUM='na',SEM="=#NUM8"] -> '#num8'
Num[NUM='na',SEM="=#NUM9"] -> '#num9'
Num[NUM='na',SEM="=#NUM10"] -> '#num10'
P[BAR=0,SEM='',SUBC='-Adj',L='en'] -> 'of' |'at'
P[BAR=0,SEM='<',SUBC='-Adj',L='en'] -> 'before'
P[BAR=0,SEM='>',SUBC='-Adj',L='en'] -> 'after'
P[BAR=0,SEM='',SUBC='-Adj',L='en'] -> 'in'
P[BAR=0,SEM='',SUBC='+Adj',L='en'] -> 'with'
C[BAR=0,L='en'] -> 'that' | 'which'
CONJ[BAR=0,SEM='',L='en'] -> 'than'
CONJ[BAR=0,SEM=' and ',L='en'] -> 'and'
A[BAR=0,SEM='>',L='en'] -> 'bigger' | 'larger' | 'greater' | 'more' | 'least' | 'above' | 'over'
A[BAR=0,SEM='<',L='en'] -> 'smaller' | 'less' | 'most' | 'maximally' | 'below' | 'under'
PART[SEM='discoveryyear',L='de'] -> 'entdeckt'
Int[NUM='sg',SEM='.//',L='de'] -> 'welcher'
Int[NUM='pl',SEM='.//',L='de'] -> 'welche'
AUX[NUM='pl',L='de'] -> 'wurden'
AUX[NUM='sg',L='de'] -> 'wurde'
V[NUM='pl',BAR=0,SEM='',L='de'] -> 'haben' | 'besitzen'
V[NUM='sg',BAR=0,SEM='',L='de'] -> 'hat' | 'besitzt'
Art[NUM='sg',SEM='',L='de'] -> 'ein' | 'einen' | 'eine' | 'einem' | 'einer'
Art[NUM='pl',SEM='',L='de'] ->
N[BAR=0,NUM='sg',SEM='mass',L='de'] -> 'masse'
N[BAR=0,NUM='sg',SEM='radius',L='de'] -> 'radius'
N[BAR=0,NUM='sg',SEM='age',L='de'] -> 'alter'
N[BAR=0,NUM='sg',SEM='temperature',L='en'] -> 'temperatur'
N[NUM='pl',BAR=0,SEM='planet',L='de'] -> 'planeten'
N[NUM='sg',BAR=0,SEM='planet',L='de'] -> 'planet' | 'planeten'
P[BAR=0,SEM='',SUBC='-Adj',L='de'] -> 'von'
P[BAR=0,SEM='<',SUBC='-Adj',L='de'] -> 'vor'
P[BAR=0,SEM='>',SUBC='-Adj',L='de'] -> 'nach'
P[BAR=0,SEM='',SUBC='-Adj',L='de'] -> 'in'
P[BAR=0,SEM='',SUBC='+Adj',L='de'] -> 'mit'
C[BAR=0,L='de'] -> 'der' | 'die'
CONJ[BAR=0,SEM='',L='de'] -> 'als'
CONJ[BAR=0,SEM=' and ',L='de'] -> 'und'
A[BAR=0,SEM='>',L='de'] -> 'größer' | 'mehr' | 'mindestens'
A[BAR=0,SEM='<',L='de'] -> 'kleiner' | 'weniger' | 'maximal' | 'unter'
"""
queries = ['planets with a mass of 19.4','planets with a radius of 1',
'planets with a radius of 1 and with a mass of 19.4',
'planets with a mass of 19.4 and planets with a radius of 1',
'planets with a mass of 19.4 and with a radius of 1 and planets with a radius of 1',
'planets that were discovered before 2002','planets with a radius of 1 that were discovered before 2015',
'what planets have a mass of 19.4','which planets have a mass of 19.4',
'what planets have a radius of 0.188','which planets have a mass of at least 19.4',
'which planets have a mass of at most 0.001','which planets have a mass smaller than 0.001',
'which planets have a mass greater than 19.4','what planets were discovered before 2001',
'what planets were discovered after 2021','which planets have a mass smaller than 0.001',
'which planets have a mass smaller than 0.001 and a radius of less than 0.188',
'what planets have a mass larger than 0.001 and a radius of at least 0.188 and a mass of at least 0.188',
'what planets have a mass larger than 0.001 and have a radius of at least 0.188','5 planets with a mass larger than 1 and 1 planet with a mass larger than 2',
'planets with a radius of 1 and with a mass of 19.4','planets that were discovered before 2001 and have a mass greater than 3',
'planeten die vor 1 entdeckt wurden und einen radius von 1 haben und eine masse von 1',
'Gibt es Planeten, die vor 2001 entdeckt wurden?',
'Show me 1 planet with an age of maximally 0.1 and a mass of at least 1 that was discovered in 2020 and 3 planets that have a radius of at least 1']
def normalize(query):
query = query.lower()
query = re.sub(r'[^\w\s#]','',query)
query = re.sub(
r'zeig mir|show me|are there any|are there|gibt es|can you show me|look for|search|suche?|finde?',
'',query)
return query
gram = grammar.FeatureGrammar.fromstring(g)
parser = parse.FeatureEarleyChartParser(gram,trace=0)
for query in queries:
query = re.sub(r'\d+\.*\d*','#NUM#',query)
query = normalize(query)
print('\n',query)
trees = list(parser.parse(query.split()))
for i,t in enumerate(trees):
answer = trees[i].label()['SEM']
print(i,answer)
# -
| dff_space_skill/Week 4 - Code Report.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Exploring S1-NRB data cubes
# + [markdown] tags=[]
# ## Introduction
# -
# **Sentinel-1 Normalised Radar Backscatter**
# Sentinel-1 Normalised Radar Backscatter (S1-NRB) is a newly developed Analysis Ready Data (ARD) product for the European Space Agency that offers high-quality, radiometrically terrain corrected (RTC) Synthetic Aperture Radar (SAR) backscatter and is designed to be compliant with the CEOS ARD for Land (CARD4L) [NRB specification](https://ceos.org/ard/files/PFS/NRB/v5.5/CARD4L-PFS_NRB_v5.5.pdf).
# You can find more detailed information about the S1-NRB product [here](https://sentinel.esa.int/web/sentinel/sentinel-1-ard-normalised-radar-backscatter-nrb-product).
#
# **SpatioTemporal Asset Catalog (STAC)**
# All S1-NRB products include metadata in JSON format compliant with the [SpatioTemporal Asset Catalog (STAC)](https://stacspec.org/) specification.
# STAC uses several sub-specifications ([Item](https://github.com/radiantearth/stac-spec/blob/master/item-spec/item-spec.md), [Collection](https://github.com/radiantearth/stac-spec/blob/master/collection-spec/collection-spec.md) & [Catalog](https://github.com/radiantearth/stac-spec/blob/master/catalog-spec/catalog-spec.md)) to create a hierarchical structure that enables efficient querying and access of large volumes of geospatial data.
# **This example notebook will give a short demonstration of how S1-NRB products can be explored as on-the-fly data cubes with little effort by utilizing the STAC metadata provided with each product. It is not intended to demonstrate how to process the S1-NRB products in the first place. For this information please refer to the [usage instructions](https://s1-nrb.readthedocs.io/en/docs/general/usage.html).**
# + [markdown] tags=[]
# ## Getting started
# -
# After following the [installation instructions](https://s1-nrb.readthedocs.io/en/latest/general/installation.html) you need to install a few additional packages into the activated conda environment to reproduce all steps presented in the following example notebook.
#
# ```bash
# conda activate nrb_env
# conda install jupyterlab stackstac rioxarray xarray_leaflet
# ```
# Instead of importing all packages now, they will successively be imported throughout the notebook:
import numpy as np
import stackstac
from S1_NRB.metadata.stac import make_catalog
# Let's assume you have a collection of S1-NRB scenes located on your local disk, a fileserver or somewhere in the cloud. As mentioned in the [Introduction](#Introduction), each S1-NRB scene includes metadata as a STAC Item, describing the scene's temporal, spatial and product specific properties.
#
# The **only step necessary to get started** with analysing your collection of scenes, is the creation of STAC Collection and Catalog files, which connect individual STAC Items and thereby create a hierarchy of STAC objects. `S1_NRB` includes the utility function [make_catalog](https://s1-nrb.readthedocs.io/en/latest/api.html#S1_NRB.metadata.stac.make_catalog), which will create these files for you. Please note that `make_catalog` expects a directory structure based on MGRS tile IDs, which allows for efficient data querying and access. After user confirmation it will take care of reorganizing your S1-NRB scenes if this directory structure doesn't exist yet.
nrb_catalog = make_catalog(directory='./NRB_thuringia', silent=True)
#
# The STAC Catalog can then be used with libraries such as [stackstac](https://github.com/gjoseph92/stackstac), which _"turns a STAC Collection into a lazy xarray.DataArray, backed by dask"._
#
# The term _lazy_ describes a [method of execution](https://tutorial.dask.org/01x_lazy.html) that only computes results when actually needed and thereby enables computations on larger-than-memory datasets. _[xarray](https://xarray.pydata.org/en/stable/index.html)_ is a Python library for working with labeled multi-dimensional arrays of data, while the Python library _[dask](https://docs.dask.org/en/latest/)_ facilitates parallel computing in a flexible way.
#
# Compatibility with [odc-stac](https://github.com/opendatacube/odc-stac), a very [similar library](https://github.com/opendatacube/odc-stac/issues/54) to stackstac, will be tested in the near future.
aoi = (10.638066, 50.708415, 11.686751, 50.975775)
ds = stackstac.stack(items=nrb_catalog, bounds_latlon=aoi,
dtype=np.dtype('float32'), chunksize=(-1, 1, 1024, 1024))
ds
# As you can see in the output above, the collection of S1-NRB scenes was successfully loaded as an `xarray.DataArray`. The metadata attributes included in all STAC Items are now available as coordinate arrays (see [here](https://docs.xarray.dev/en/stable/user-guide/terminology.html#term-Coordinate) for clarification of Xarray's terminology) and can be utilized during analysis. This will be explored in the next section.
# + [markdown] tags=[]
# ## Data Exploration
#
# *coming soon*
# -
# ### Spatial
# + pycharm={"name": "#%%\n"}
# + pycharm={"name": "#%%\n"}
# -
# ### Temporal
# ### Bands / Attributes
# + [markdown] tags=[]
# ## Example Analysis
#
# *coming soon*
# -
| docs/examples/nrb_cube.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Emulators: Measuring performance
#
# This example illustrates how different neural networks perform in emulating the log-likelihood surface of a time series and in Bayesian inference, using a two-step MCMC procedure with emulator neural networks [Emulated Metropolis MCMC](../sampling/first-example.ipynb).
#
# It follows on from [Emulators: First example](../mcmc/first-example-emulator.ipynb)
#
# Like in the first example, I start by importing pints:
import pints
# Next, I create a model class using the "Logistic" toy model included in pints:
# +
import pints.toy as toy
class RescaledModel(pints.ForwardModel):
def __init__(self):
self.base_model = toy.LogisticModel()
def simulate(self, parameters, times):
# Run a simulation with the given parameters for the
# given times and return the simulated values
r, k = parameters
r = r / 50
k = k * 500
return self.base_model.simulate([r, k], times)
def simulateS1(self, parameters, times):
# Run a simulation with the given parameters for the
# given times and return the simulated values
r, k = parameters
r = r / 50
k = k * 500
return self.base_model.simulateS1([r, k], times)
def n_parameters(self):
# Return the dimension of the parameter vector
return 2
model = toy.LogisticModel()
# -
# In order to generate some test data, I choose an arbitrary set of "true" parameters:
true_parameters = [0.015, 500]
start_parameters = [0.75, 1.0] # rescaled true parameters
# And a number of time points at which to sample the time series:
import numpy as np
times = np.linspace(0, 1000, 400)
# Using these parameters and time points, I generate an example dataset:
org_values = model.simulate(true_parameters, times)
range_values = max(org_values) - min(org_values)
# And make it more realistic by adding gaussian noise:
noise = 0.05 * range_values
print("Gaussian noise:", noise)
values = org_values + np.random.normal(0, noise, org_values.shape)
values = org_values + np.random.normal(0, noise, org_values.shape)
# Using matplotlib and seaborn (optional - for styling), I look at the noisy time series I just simulated:
import matplotlib.pyplot as plt
import seaborn as sns
sns.set(context='notebook', style='whitegrid', palette='deep', font='Times New Roman',
font_scale=1.5, color_codes=True, rc={"grid.linewidth": 1})
plt.figure(figsize=(12,4.5))
plt.xlabel('Time')
plt.ylabel('Values')
plt.plot(times, values, label='Noisy data')
plt.plot(times, org_values, lw=2, label='Original data')
plt.legend()
plt.show()
# Now, I have enough data (a model, a list of times, and a list of values) to formulate a PINTS problem:
model = RescaledModel()
problem = pints.SingleOutputProblem(model, times, values)
# I now have some toy data, and a model that can be used for forward simulations. To make it into a probabilistic problem, a _noise model_ needs to be added. This can be done using the `GaussianLogLikelihood` function, which assumes independently distributed Gaussian noise over the data, and can calculate log-likelihoods:
log_likelihood = pints.GaussianKnownSigmaLogLikelihood(problem, noise)
# This `log_likelihood` represents the _conditional probability_ $p(y|\theta)$, given a set of parameters $\theta$ and a series of $y=$ `values`, it can calculate the probability of finding those values if the real parameters are $\theta$.
#
# This can be used in a Bayesian inference scheme to find the quantity of interest:
#
# $p(\theta|y) = \frac{p(\theta)p(y|\theta)}{p(y)} \propto p(\theta)p(y|\theta)$
#
# To solve this, a _prior_ is defined, indicating an initial guess about what the parameters should be.
# Similarly as using a _log-likelihood_ (the natural logarithm of a likelihood), this is defined by using a _log-prior_. Hence, the above equation simplifies to:
#
# $\log p(\theta|y) \propto \log p(\theta) + \log p(y|\theta)$
#
# In this example, it is assumed that we don't know too much about the prior except lower and upper bounds for each variable: We assume the first model parameter is somewhere on the interval $[0.01, 0.02]$, the second model parameter on $[400, 600]$, and the standard deviation of the noise is somewhere on $[1, 100]$.
# Create (rescaled) bounds for our parameters and get prior
bounds = pints.RectangularBoundaries([0.5, 0.8], [1.0, 1.2])
log_prior = pints.UniformLogPrior(bounds)
# With this prior, the numerator of Bayes' rule can be defined -- the unnormalised log posterior, $\log \left[ p(y|\theta) p(\theta) \right]$, which is the natural logarithm of the likelihood times the prior:
# Create a posterior log-likelihood (log(likelihood * prior))
log_posterior = pints.LogPosterior(log_likelihood, log_prior)
# Finally we create a list of guesses to use as initial positions. We'll run three MCMC chains so we create three initial positions, using the rescaled true parameters:
x0 = [
np.array(start_parameters) * 0.9,
np.array(start_parameters) * 1.05,
np.array(start_parameters) * 1.15,
]
# ## Creating training data
# +
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
from sklearn.model_selection import train_test_split
input_parameters = log_prior.sample(2000)
x = [p[0] for p in input_parameters]
y = [p[1] for p in input_parameters]
likelihoods = np.apply_along_axis(log_likelihood, 1, input_parameters)
likelihoods[:5]
X_train, X_valid, y_train, y_valid = train_test_split(input_parameters, likelihoods, test_size=0.3, random_state=0)
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
ax.scatter(x, y, list(likelihoods))
plt.show()
# -
# ## Training various NNs with different setups
# +
layers = list(range(1, 11, 1))
neurons = [ 2**j for j in range(4,6+1) ]
epochs = 1000
print("NN layers:", layers)
print("NN hidden units:", neurons)
# Compute all possible permutations of NN parameters
hyperparams = [[i, j] for i in layers
for j in neurons]
print ("All possible layer and hidden units permutations are: " + str(hyperparams))
print(len(hyperparams), "sets of hyperparameters")
# +
from sklearn.preprocessing import StandardScaler, MinMaxScaler
from scipy import stats
sigma0 = [[ 1.01547594e-05, -2.58358260e-06], [-2.58358260e-06, 1.22093040e-05]]
scaling_factors = [1/50, 500]
runs = 5
mapes = []
rhats_nn = []
rhats_emu = []
w_distances_nn1 = []
w_distances_nn2 = []
w_distances_nn3 = []
w_distances_emu1 = []
w_distances_emu2 = []
w_distances_emu3 = []
acceptance_rates1 = []
acceptance_rates2 = []
acceptance_rates3 = []
# Run standard Metropolis Hastings MCMC
# Create mcmc routine
mcmc = pints.MCMCController(log_posterior, 3, x0, sigma0=sigma0, method=pints.MetropolisRandomWalkMCMC)
mcmc.set_max_iterations(30000) # Add stopping criterion
mcmc.set_log_to_screen(False) # Disable logging mode
metropolis_chains = mcmc.run()
# Revert scaling
metropolis_chains_rescaled = np.copy(metropolis_chains)
metropolis_chain_rescaled = metropolis_chains_rescaled[0]
metropolis_chain_rescaled = metropolis_chain_rescaled[10000:]
metropolis_chains = np.array([[[s*f for s,f in zip(samples, scaling_factors)] for samples in chain]
for chain in metropolis_chains])
metropolis_chain = metropolis_chains[0][10000:]
metropolis_chain2 = metropolis_chains[1][10000:]
metropolis_chain3 = metropolis_chains[2][10000:]
# Split chains by parameter for computing Wasserstein distance
metropolis_chain_r = np.array([sample[0] for sample in metropolis_chain])
metropolis_chain_k = np.array([sample[1] for sample in metropolis_chain])
metropolis_chain2_r = np.array([sample[0] for sample in metropolis_chain2])
metropolis_chain2_k = np.array([sample[1] for sample in metropolis_chain2])
metropolis_chain3_r = np.array([sample[0] for sample in metropolis_chain3])
metropolis_chain3_k = np.array([sample[1] for sample in metropolis_chain3])
# Check convergence using rhat criterion
rhat = pints.rhat_all_params(metropolis_chains_rescaled[:, 10000:, :])
# Create a number of splits along each axis
test_splits = 50
r_grid, k_grid, test_data = pints.generate_grid(bounds.lower(), bounds.upper(), test_splits)
model_prediction = pints.predict_grid(log_likelihood, test_data)
for i, p in enumerate(hyperparams):
print("Testing NN with parameters (layers, neurons):", p)
mape = 0
w_distance_r_nn, w_distance_k_nn = 0, 0
w_distance_r_emu, w_distance_k_emu = 0, 0
w_distance_r_nn2, w_distance_k_nn2 = 0, 0
w_distance_r_emu2, w_distance_k_emu2 = 0, 0
w_distance_r_nn3, w_distance_k_nn3 = 0, 0
w_distance_r_emu3, w_distance_k_emu3 = 0, 0
rates1 = [0, 0, 0]
rates2 = [0, 0, 0]
rates3 = [0, 0, 0]
for n in range(runs):
print(" Run", n+1, "/", runs)
# Train NN with given parameters
emu = pints.MultiLayerNN(problem, X_train, y_train, input_scaler=MinMaxScaler(), output_scaler=StandardScaler())
emu.set_parameters(layers=p[0], neurons=p[1], hidden_activation='relu', activation='linear', learning_rate=0.0001)
hist = emu.fit(epochs=epochs, batch_size=32, X_val=X_valid, y_val=y_valid, verbose=0)
log_posterior_emu = pints.LogPosterior(emu, log_prior)
print(" Done in", len(hist.history['loss']), "epochs")
# Compute mean abs. percentage error between likelihood surfaces
emu_prediction = pints.predict_grid(emu, test_data)
mape += np.mean(np.abs((model_prediction - emu_prediction) / model_prediction))
# Run Metropolis Hastings MCMC using NN posterior
# Create mcmc routine
mcmc = pints.MCMCController(log_posterior_emu, 3, x0, sigma0=sigma0, method=pints.MetropolisRandomWalkMCMC)
mcmc.set_max_iterations(30000) # Add stopping criterion
mcmc.set_log_to_screen(False) # Disable logging mode
chains_emu = mcmc.run()
# Revert scaling
chains_emu_rescaled = np.copy(chains_emu)
chain_emu_rescaled = chains_emu_rescaled[0]
chain_emu_rescaled = chain_emu_rescaled[10000:]
chains_emu = np.array([[[s*f for s,f in zip(samples, scaling_factors)] for samples in chain] for chain in chains_emu])
chain_emu = chains_emu[0][10000:]
chain_emu2 = chains_emu[1][10000:]
chain_emu3 = chains_emu[2][10000:]
# Check convergence using rhat criterion
rhat = pints.rhat_all_params(chains_emu_rescaled[:, 10000:, :])
rhats_nn.append(rhat)
# Run 2-step Metropolis Hastings MCMC
# Create mcmc routine
mcmc = pints.MCMCController(log_posterior_emu, 3, x0, sigma0=sigma0, method=pints.EmulatedMetropolisMCMC, f=log_posterior)
mcmc.set_max_iterations(30000) # Add stopping criterion
mcmc.set_log_to_screen(False) # Disable logging mode
emulated_chains = mcmc.run()
# Get acceptance rates per stage of the 2-step procedure
rates = mcmc.acceptance_rates()
rates1 = [sum(r) for r in zip(rates1, rates[0])] # Chain 1
rates2 = [sum(r) for r in zip(rates2, rates[1])] # Chain 2
rates3 = [sum(r) for r in zip(rates3, rates[2])] # Chain 3
# Revert scaling
emulated_chains_rescaled = np.copy(emulated_chains)
emulated_chain_rescaled = emulated_chains_rescaled[0]
emulated_chain_rescaled = emulated_chain_rescaled[10000:]
emulated_chains = np.array([[[s*f for s,f in zip(samples, scaling_factors)] for samples in chain]
for chain in emulated_chains])
emulated_chain = emulated_chains[0][10000:]
emulated_chain2 = emulated_chains[1][10000:]
emulated_chain3 = emulated_chains[2][10000:]
# Check convergence using rhat criterion
rhat = pints.rhat_all_params(emulated_chains_rescaled[:, 10000:, :])
rhats_emu.append(rhat)
# Split chains by parameter for computing Wasserstein distance
chain_emu_r = np.array([sample[0] for sample in chain_emu])
chain_emu_k = np.array([sample[1] for sample in chain_emu])
chain2_emu_r = np.array([sample[0] for sample in chain_emu2])
chain2_emu_k = np.array([sample[1] for sample in chain_emu2])
chain3_emu_r = np.array([sample[0] for sample in chain_emu3])
chain3_emu_k = np.array([sample[1] for sample in chain_emu3])
emulated_chain_r = np.array([sample[0] for sample in emulated_chain])
emulated_chain_k = np.array([sample[1] for sample in emulated_chain])
emulated_chain2_r = np.array([sample[0] for sample in emulated_chain2])
emulated_chain2_k = np.array([sample[1] for sample in emulated_chain2])
emulated_chain3_r = np.array([sample[0] for sample in emulated_chain3])
emulated_chain3_k = np.array([sample[1] for sample in emulated_chain3])
# Compute Wasserstein distances
w_distance_r_nn = stats.wasserstein_distance(metropolis_chain_r, chain_emu_r)
w_distance_k_nn = stats.wasserstein_distance(metropolis_chain_k, chain_emu_k)
w_distance_r_emu = stats.wasserstein_distance(metropolis_chain_r, emulated_chain_r)
w_distance_k_emu = stats.wasserstein_distance(metropolis_chain_k, emulated_chain_k)
w_distance_r_nn2 = stats.wasserstein_distance(metropolis_chain2_r, chain2_emu_r)
w_distance_k_nn2 = stats.wasserstein_distance(metropolis_chain2_k, chain2_emu_k)
w_distance_r_emu2 = stats.wasserstein_distance(metropolis_chain2_r, emulated_chain2_r)
w_distance_k_emu2 = stats.wasserstein_distance(metropolis_chain2_k, emulated_chain2_k)
w_distance_r_nn3 = stats.wasserstein_distance(metropolis_chain3_r, chain3_emu_r)
w_distance_k_nn3 = stats.wasserstein_distance(metropolis_chain3_k, chain3_emu_k)
w_distance_r_emu3 = stats.wasserstein_distance(metropolis_chain3_r, emulated_chain3_r)
w_distance_k_emu3 = stats.wasserstein_distance(metropolis_chain3_k, emulated_chain3_k)
w_distances_nn1.append((w_distance_r_nn/runs, w_distance_k_nn/runs))
w_distances_nn2.append((w_distance_r_nn2/runs, w_distance_k_nn2/runs))
w_distances_nn3.append((w_distance_r_nn3/runs, w_distance_k_nn3/runs))
w_distances_emu1.append((w_distance_r_emu/runs, w_distance_k_emu/runs))
w_distances_emu2.append((w_distance_r_emu2/runs, w_distance_k_emu2/runs))
w_distances_emu3.append((w_distance_r_emu3/runs, w_distance_k_emu3/runs))
mapes.append(mape/runs)
acceptance_rates1.append([r/runs for r in rates1]) # Averages for chain 1
acceptance_rates2.append([r/runs for r in rates2]) # Averages for chain 2
acceptance_rates3.append([r/runs for r in rates3]) # Averages for chain 3
print(p, mape/runs, (w_distance_r_nn/runs, w_distance_k_nn/runs), [r/runs for r in rates1])
# -
mapes[:5]
w_distances_nn1[:5]
w_distances_nn2[:5]
acceptance_rates1[:5]
#self._acceptance, self._acceptance1, self._acceptance2
chain1_acceptance_rates2 = [r[2] for r in acceptance_rates1]
chain2_acceptance_rates2 = [r[2] for r in acceptance_rates2]
chain3_acceptance_rates2 = [r[2] for r in acceptance_rates3]
chain1_acceptance_rates2[:5]
sns.set(context='notebook', style='whitegrid', palette='deep', font='Times New Roman',
font_scale=1.5, color_codes=True, rc={"grid.linewidth": 1})
fig, ax = plt.subplots(figsize=(15,6))
plt.xlabel('Mean Absolute Percentage Error')
plt.ylabel('Acceptance rate in step 2')
ax.axhline(y=0.565, ls='--',lw=2, c='k', alpha=0.7)
ax.scatter(mapes, chain1_acceptance_rates2, lw=2, label='Chain 1')
ax.scatter(mapes, chain2_acceptance_rates2, lw=2, label='Chain 2')
ax.scatter(mapes, chain3_acceptance_rates2, lw=2, label='Chain 3')
for i, txt in enumerate(hyperparams):
ax.annotate(txt, (mapes[i], chain1_acceptance_rates2[i]))
plt.legend()
plt.show()
# Extract Wasserstein distances by parameter
w_distances_r1 = [d[0] for d in w_distances_nn1]
w_distances_k1 = [d[1] for d in w_distances_nn1]
w_distances_sum1 = [r+k for r, k in zip(w_distances_r1, w_distances_k1)]
# +
w_distances_r = []
w_distances_k = []
w_distances_sum = []
for dist in [w_distances_nn1, w_distances_nn2, w_distances_nn3]:
# Extract Wasserstein distances by parameter
dist_r = [d[0] for d in dist]
dist_k = [d[1] for d in dist]
# Compute sum of rescaled distances
scaler_r = MinMaxScaler()
scaler_k = MinMaxScaler()
dist_r = scaler_r.fit_transform(np.array(dist_r).reshape(-1, 1))
dist_k = scaler_k.fit_transform(np.array(dist_k).reshape(-1, 1))
w_distances_r.append(list(dist_r))
w_distances_k.append(list(dist_k))
w_distances_sum.append([r+k for r, k in zip(list(dist_r), list(dist_k))])
w_distances_sum[0]
# +
sns.set(context='notebook', style='whitegrid', palette='deep', font='Times New Roman',
font_scale=1.5, color_codes=True, rc={"grid.linewidth": 1})
fig, ax = plt.subplots(figsize=(15,6))
plt.xlabel('Mean Absolute Percentage Error')
plt.ylabel('Acceptance rate in step 2')
ax.axhline(y=0.565, ls='--',lw=2, c='k', alpha=0.7)
ax.scatter(mapes, chain1_acceptance_rates2, lw=3, label='Chain 1')
ax.scatter(mapes, chain2_acceptance_rates2, lw=3, label='Chain 2')
ax.scatter(mapes, chain3_acceptance_rates2, lw=3, label='Chain 3')
for i, txt in enumerate(hyperparams):
ax.annotate(txt, (mapes[i], chain1_acceptance_rates2[i]))
plt.legend()
plt.show()
fig.savefig("figures/nn-comparisons/mae.png", bbox_inches='tight', dpi=600)
# +
sns.set(context='notebook', style='whitegrid', palette='deep', font='Times New Roman',
font_scale=1.5, color_codes=True, rc={"grid.linewidth": 1})
fig, ax = plt.subplots(figsize=(15,6))
plt.xlabel('Sum of rescaled Wasserstein distances')
plt.ylabel('Acceptance rate in step 2')
ax.axhline(y=0.565, ls='--',lw=2, c='k', alpha=0.7)
ax.scatter(w_distances_sum[0], chain1_acceptance_rates2, lw=3, label='Chain 1')
ax.scatter(w_distances_sum[1], chain2_acceptance_rates2, lw=3, label='Chain 2')
ax.scatter(w_distances_sum[2], chain3_acceptance_rates2, lw=3, label='Chain 3')
for i, txt in enumerate(hyperparams):
ax.annotate(txt, (w_distances_sum[0][i], chain1_acceptance_rates2[i]))
plt.legend()
plt.show()
fig.savefig("figures/nn-comparisons/rescaled-wasserstein-sum.png", bbox_inches='tight', dpi=600)
# +
sns.set(context='notebook', style='whitegrid', palette='deep', font='Times New Roman',
font_scale=1.5, color_codes=True, rc={"grid.linewidth": 1})
fig, ax = plt.subplots(figsize=(15,6))
plt.xlabel('Sum of Wasserstein distances')
plt.ylabel('Acceptance rate in step 2')
ax.axhline(y=0.565, ls='--',lw=2, c='k', alpha=0.7)
ax.scatter(w_distances_sum1, chain1_acceptance_rates2, lw=3, label='Chain 1')
for i, txt in enumerate(hyperparams):
ax.annotate(txt, (w_distances_sum1[i], chain1_acceptance_rates2[i]))
plt.legend()
plt.show()
fig.savefig("figures/nn-comparisons/wasserstein-sum.png", bbox_inches='tight', dpi=600)
# -
| examples/emulators/mcmc/nn-quality-vs-mcmc-acceptance-rates-different-nns-3-wasserstein.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ETS models
#
# The ETS models are a family of time series models with an underlying state space model consisting of a level component, a trend component (T), a seasonal component (S), and an error term (E).
#
# This notebook gives a very brief introduction to these models and shows how they can be used with statsmodels. For a more thorough treatment we refer to [1], chapter 8 (free online resource), on which the implementation in statsmodels and the examples used in this notebook are based.
#
# [1] Hyndman, <NAME>., and <NAME>. *Forecasting: principles and practice*, 3rd edition, OTexts, 2019. https://www.otexts.org/fpp3/7
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# %matplotlib inline
from statsmodels.tsa.exponential_smoothing.ets import ETSModel
plt.rcParams['figure.figsize'] = (12, 8)
# ## Simple exponential smoothing
#
# The simplest of the ETS models is also known as *simple exponential smoothing*. In ETS terms, it corresponds to the (A, N, N) model, that is, a model with additive errors, no trend, and no seasonality. The state space formulation of Holt's method is:
#
# \begin{align}
# y_{t} &= y_{t-1} + e_t\\
# l_{t} &= l_{t-1} + \alpha e_t\\
# \end{align}
#
# This state space formulation can be turned into a different formulation, a forecast and a smoothing equation (as can be done with all ETS models):
#
# \begin{align}
# \hat{y}_{t|t-1} &= l_{t-1}\\
# l_{t} &= \alpha y_{t-1} + (1 - \alpha) l_{t-1}
# \end{align}
#
# Here, $\hat{y}_{t|t-1}$ is the forecast/expectation of $y_t$ given the information of the previous step. In the simple exponential smoothing model, the forecast corresponds to the previous level. The second equation (smoothing equation) calculates the next level as weighted average of the previous level and the previous observation.
oildata = [
111.0091, 130.8284, 141.2871, 154.2278,
162.7409, 192.1665, 240.7997, 304.2174,
384.0046, 429.6622, 359.3169, 437.2519,
468.4008, 424.4353, 487.9794, 509.8284,
506.3473, 340.1842, 240.2589, 219.0328,
172.0747, 252.5901, 221.0711, 276.5188,
271.1480, 342.6186, 428.3558, 442.3946,
432.7851, 437.2497, 437.2092, 445.3641,
453.1950, 454.4096, 422.3789, 456.0371,
440.3866, 425.1944, 486.2052, 500.4291,
521.2759, 508.9476, 488.8889, 509.8706,
456.7229, 473.8166, 525.9509, 549.8338,
542.3405
]
oil = pd.Series(oildata, index=pd.date_range('1965', '2013', freq='AS'))
oil.plot()
plt.ylabel("Annual oil production in Saudi Arabia (Mt)");
# The plot above shows annual oil production in Saudia Arabia in million tonnes. The data are taken from the R package `fpp2` (companion package to prior version [1]).
# Below you can see how to fit a simple exponential smoothing model using statsmodel's ETS implementation to this data. Additionally, the fit using `forecast` in R is shown as comparison.
# +
model = ETSModel(oil, error='add', trend='add', damped_trend=True)
fit = model.fit(maxiter=10000)
oil.plot(label='data')
fit.fittedvalues.plot(label='statsmodels fit')
plt.ylabel("Annual oil production in Saudi Arabia (Mt)");
# obtained from R
params_R = [0.99989969, 0.11888177503085334, 0.80000197, 36.46466837, 34.72584983]
yhat = model.smooth(params_R).fittedvalues
yhat.plot(label='R fit', linestyle='--')
plt.legend();
# -
# By default the initial states are considered to be fitting parameters and are estimated by maximizing log-likelihood. Additionally it is possible to only use a heuristic for the initial values. In this case this leads to better agreement with the R implementation.
# +
model_heuristic = ETSModel(oil, error='add', trend='add', damped_trend=True,
initialization_method='heuristic')
fit_heuristic = model_heuristic.fit()
oil.plot(label='data')
fit.fittedvalues.plot(label='estimated')
fit_heuristic.fittedvalues.plot(label='heuristic', linestyle='--')
plt.ylabel("Annual oil production in Saudi Arabia (Mt)");
# obtained from R
params = [0.99989969, 0.11888177503085334, 0.80000197, 36.46466837, 34.72584983]
yhat = model.smooth(params).fittedvalues
yhat.plot(label='with R params', linestyle=':')
plt.legend();
# -
fit.summary()
fit_heuristic.summary()
# ## Holt-Winters' seasonal method
#
# The exponential smoothing method can be modified to incorporate a trend and a seasonal component. In the additive Holt-Winters' method, the seasonal component is added to the rest. This model corresponds to the ETS(A, A, A) model, and has the following state space formulation:
#
# \begin{align}
# y_t &= l_{t-1} + b_{t-1} + s_{t-m} + e_t\\
# l_{t} &= l_{t-1} + b_{t-1} + \alpha e_t\\
# b_{t} &= b_{t-1} + \beta e_t\\
# s_{t} &= s_{t-m} + \gamma e_t
# \end{align}
#
#
austourists_data = [
30.05251300, 19.14849600, 25.31769200, 27.59143700,
32.07645600, 23.48796100, 28.47594000, 35.12375300,
36.83848500, 25.00701700, 30.72223000, 28.69375900,
36.64098600, 23.82460900, 29.31168300, 31.77030900,
35.17787700, 19.77524400, 29.60175000, 34.53884200,
41.27359900, 26.65586200, 28.27985900, 35.19115300,
42.20566386, 24.64917133, 32.66733514, 37.25735401,
45.24246027, 29.35048127, 36.34420728, 41.78208136,
49.27659843, 31.27540139, 37.85062549, 38.83704413,
51.23690034, 31.83855162, 41.32342126, 42.79900337,
55.70835836, 33.40714492, 42.31663797, 45.15712257,
59.57607996, 34.83733016, 44.84168072, 46.97124960,
60.01903094, 38.37117851, 46.97586413, 50.73379646,
61.64687319, 39.29956937, 52.67120908, 54.33231689,
66.83435838, 40.87118847, 51.82853579, 57.49190993,
65.25146985, 43.06120822, 54.76075713, 59.83447494,
73.25702747, 47.69662373, 61.09776802, 66.05576122,
]
index = pd.date_range("1999-03-01", "2015-12-01", freq="3MS")
austourists = pd.Series(austourists_data, index=index)
austourists.plot()
plt.ylabel('Australian Tourists');
# +
# fit in statsmodels
model = ETSModel(austourists, error="add", trend="add", seasonal="add",
damped_trend=True, seasonal_periods=4)
fit = model.fit()
# fit with R params
params_R = [
0.35445427, 0.03200749, 0.39993387, 0.97999997, 24.01278357,
0.97770147, 1.76951063, -0.50735902, -6.61171798, 5.34956637
]
fit_R = model.smooth(params_R)
austourists.plot(label='data')
plt.ylabel('Australian Tourists')
fit.fittedvalues.plot(label='statsmodels fit')
fit_R.fittedvalues.plot(label='R fit', linestyle='--')
plt.legend();
# -
fit.summary()
fit._rank
| examples/notebooks/ets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import MPO_class as MPO
import MPS_class as MPS
import SpinSystems_1d as SS1d
import dmrg1 as dmrg
import thermal_tdvp2 as ttdvp
from MixedMPS_class import mix_compute_corr
import SpinSystems_2d as SS2d
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from ncon import ncon
from compute_connected_correlation_function import compute_corr
# +
h_space = [0.1,.15,.2]#np.arange(0.01,0.4,0.005)
L = 5
N = 2**L
J = 0.1
dbeta = 1e-3*5
sigma_z = np.array([[1,0],[0,-1]])
G = []; op1 = []; op2 = []
T = [];
for h in h_space:
print('h',h,end=' ')
beta = 0.
H = SS2d.IsingMPO_2D_diagonal_MPS(L,h=h,J=J)
engine = ttdvp.thermalTDVP2(H)
engine.initialize()
E = []
E.append(engine.MPS.contractMPOmixMPS(H))
engine.chi_MAX = 128
while(E[-1]+2*L*J*(L-1)>0):
#print('diff:',E[-1]+2*L*J*(L-1))
if beta<3*dbeta:
engine.truncate_info = False
else:
engine.truncate_info = True
engine.beta_step(dbeta,1e-11)
E.append(engine.MPS.contractMPOmixMPS(H))
beta += dbeta
corr_stuff = mix_compute_corr(engine.MPS,sigma_z)
G.append(corr_stuff[0])
op1.append(corr_stuff[1][0])
op2.append(corr_stuff[1][1])
T.append(1/beta)
print('T',1/beta)
# -
for x in engine.MPS.M:
print(x.shape)
h_space.size
# +
plt.plot(h_space,[(op2[x]).sum()/(16**2) for x in range(h_space.size)])
np.savetxt('4x4_J0.25.dat',np.array([(op2[x]).sum()/(16**2) for x in range(h_space.size)]))
np.savetxt('Temperature_4x4_J0.25.dat',T)
# -
plt.plot(T)
plt.imshow(op2[-1,:,:],aspect='auto',origin='lower')
plt.imshow(op2[:,:].reshape(-1,256),aspect='auto')
plt.colorbar()
corr_stuff[0]
plt.plot(dbeta*np.arange(len(mx)),np.abs(mx-ncon([engine_GS.MPS.M[0],sigma_x,engine_GS.MPS.M[0]],[[1,2,3],[2,5],[1,5,3]])))
plt.yscale('log')
engine_GS = dmrg.DMRG1(H)
engine_GS.initialize(64)
for _ in range(10):
engine_GS.dmrg_step()
ncon([engine_GS.MPS.M[0],sigma_x,engine_GS.MPS.M[0]],[[1,2,3],[2,5],[1,5,3]])
sigma_z.shape
2*L*(L-1)
engine.MPS.M[10]
# +
plt.plot(1/beta_space[:G.size],[(op2[x] ).sum()/(64**2) for x in range(G.size)])
plt.plot(1/beta_space[:G.size],np.ones(G.size)*(corr_gs ).sum()/(64**2))
plt.xlim([-5,50])
# -
op2[0].sum()/(64**2)
import compute_connected_correlation_function
stuffs = compute_connected_correlation_function.compute_corr(engine_GS.MPS,sigma_z)
corr_gs = stuffs[1][1]
plt.imshow(corr_gs-(op2[-1] - np.eye(64)))
plt.colorbar()
| .ipynb_checkpoints/Test-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: lang101
# language: python
# name: lang101
# ---
# ## Import libraries
# +
# standard library
import sys,os
sys.path.append(os.path.join(".."))
# data and nlp
import pandas as pd
import spacy
nlp = spacy.load("en_core_web_sm", disable=["ner"])
# visualisation
import pyLDAvis.gensim
pyLDAvis.enable_notebook()
import seaborn as sns
from matplotlib import rcParams
# figure size in inches
rcParams['figure.figsize'] = 20,10
# LDA tools
import gensim
import gensim.corpora as corpora
from gensim.models import CoherenceModel
from utils import lda_utils
# warnings
import logging, warnings
warnings.filterwarnings('ignore')
logging.basicConfig(format='%(asctime)s : %(levelname)s : %(message)s', level=logging.ERROR)
# -
# ## Read in text
# +
#with open("Austen_Pride.txt") as f:
# text = f.read()
with open(os.path.join("..", "data", "100_english_novels", "corpus", "Anon_Clara_1864.txt")) as f:
text = f.read()
text = text[:20000]
# -
# __Split into individual sentences__
doc = nlp(text)
doc.sents[0]
sentences = [sent.string.strip() for sent in doc.sents]
# __Create chunks of 10 sentences at a time__
chunks = []
for i in range(0, len(sentences), 10):
chunks.append(' '.join(sentences[i:i+10]))
# ## Process using ```gensim```
#
# Here we're using ```gensim``` to produce efficiently procude a model of bigrams and trigrams in the data.
#
# We first create bigrams based on words appearing one after another frequently. These bigrams are then fed into a trigram generator, which takes the bigram as the second part of a bigram.
#
# - bigram model = (a,b)
# - trigram model = (x, (a,b))
# Build the bigram and trigram models
bigram = gensim.models.Phrases(chunks, min_count=3, threshold=100) # higher threshold fewer phrases.
trigram = gensim.models.Phrases(bigram[chunks], threshold=100)
bigram_mod = gensim.models.phrases.Phraser(bigram)
trigram_mod = gensim.models.phrases.Phraser(trigram)
# We can use the ```lda_utils.process_words()``` function to further process the data using ```spaCy```.
#
# Make sure to check the util script and see what arguments this function takes!
data_processed = lda_utils.process_words(chunks,nlp, bigram_mod, trigram_mod, allowed_postags=["NOUN"])
# We then need to create a ```gensim``` dictionary and a ```gensim``` corpus.
#
# The dictionary converts each word into an integer value; the corpus creates a 'bag of words' model for all of the data - in this case, chunks of 10 sentences at a time.
# +
# Create Dictionary
id2word = corpora.Dictionary(data_processed)
# Create Corpus: Term Document Frequency
corpus = [id2word.doc2bow(text) for text in data_processed]
# -
# We then feed all of this information into our LDA algorithm using ```gensim```.
# Build LDA model
lda_model = gensim.models.LdaMulticore(corpus=corpus,
id2word=id2word,
num_topics=10,
random_state=100,
chunksize=10,
passes=10,
iterations=100,
per_word_topics=True,
minimum_probability=0.0)
# We can then calculate model perplexity and coherence in order to interpret how well the model performs.
# +
# Compute Perplexity
print('\nPerplexity: ', lda_model.log_perplexity(corpus)) # a measure of how good the model is. lower the better.
# Compute Coherence Score
coherence_model_lda = CoherenceModel(model=lda_model,
texts=data_processed,
dictionary=id2word,
coherence='c_v')
#coherence_lda = coherence_model_lda.get_coherence()
#print('\nCoherence Score: ', coherence_lda)
# -
# Lastly, we can inspect the 'contents' of the model
pprint(lda_model.print_topics())
# ## Run model multiple times to find best fit
# Can take a long time to run.
model_list, coherence_values = lda_utils.compute_coherence_values(texts=data_processed,
corpus=corpus,
dictionary=id2word,
start=5,
limit=40,
step=5)
# ## Create dataframe showing most dominant topic per chunk
# +
df_topic_keywords = lda_utils.format_topics_sentences(ldamodel=lda_model,
corpus=corpus,
texts=data_processed)
# Format
df_dominant_topic = df_topic_keywords.reset_index()
df_dominant_topic.columns = ['Document_No', 'Dominant_Topic', 'Topic_Perc_Contrib', 'Keywords', 'Text']
df_dominant_topic.sample(10)
# +
# Display setting to show more characters in column
pd.options.display.max_colwidth = 100
sent_topics_sorteddf = pd.DataFrame()
sent_topics_outdf_grpd = df_topic_keywords.groupby('Dominant_Topic')
for i, grp in sent_topics_outdf_grpd:
sent_topics_sorteddf = pd.concat([sent_topics_sorteddf,
grp.sort_values(['Perc_Contribution'], ascending=False).head(1)],
axis=0)
# Reset Index
sent_topics_sorteddf.reset_index(drop=True, inplace=True)
# Format
sent_topics_sorteddf.columns = ['Topic_Num', "Topic_Perc_Contrib", "Keywords", "Representative Text"]
# Show
sent_topics_sorteddf.head(10)
# -
# ## Visualise topics
vis = pyLDAvis.gensim.prepare(lda_model, corpus, dictionary=lda_model.id2word)
vis
# ## To dataframe
values = list(lda_model.get_document_topics(corpus))
# __Split tuples and keep only values per topic__
split = []
for entry in values:
topic_prevelance = []
for topic in entry:
topic_prevelance.append(topic[1])
split.append(topic_prevelance)
# __Create document-topic matrix__
df = pd.DataFrame(map(list,zip(*split)))
df
# ## Plot with seaborn
sns.lineplot(data=df.T.rolling(50).mean())
| notebooks/session8.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.3 64-bit (''base'': conda)'
# language: python
# name: python373jvsc74a57bd0210f9608a45c0278a93c9e0b10db32a427986ab48cfc0d20c139811eb78c4bbc
# ---
import nltk
from nltk.stem.porter import *
from torch.nn import *
from torch.optim import *
import numpy as np
import pandas as pd
import torch,torchvision
import random
from tqdm import *
from torch.utils.data import Dataset,DataLoader
stemmer = PorterStemmer()
def tokenize(sentence):
return nltk.word_tokenize(sentence)
tokenize('$100')
def stem(word):
return stemmer.stem(word.lower())
stem('organic')
def bag_of_words(tokenized_words,all_words):
tokenized_words = [stem(w) for w in tokenized_words]
bag = np.zeros(len(all_words))
for idx,w in enumerate(all_words):
if w in tokenized_words:
bag[idx] = 1.0
return bag
bag_of_words(['hi'],['how','hi'])
data = pd.read_csv('./data.csv')
data = data[:1000]
X = data['Text']
y = data['Summary']
X_words = []
data = []
y_words = []
for X_batch,y_batch in tqdm(zip(X,y)):
X_batch = tokenize(X_batch)
y_batch = tokenize(y_batch)
new_X = []
new_y = []
for Xb in X_batch:
new_X.append(stem(Xb))
for yb in y_batch:
new_y.append(stem(yb))
X_words.extend(new_X)
y_words.extend(new_y)
data.append([new_X,new_y])
X_words = sorted(set(X_words))
y_words = sorted(set(y_words))
np.random.shuffle(data)
X = []
y = []
for X_batch,y_batch in tqdm(data):
X.append(bag_of_words(X_batch,X_words))
y.append(bag_of_words(y_batch,y_words))
from sklearn.model_selection import *
X_train,X_test,y_train,y_test = train_test_split(X,y,test_size=0.25,shuffle=False)
device = 'cuda'
X_train = torch.from_numpy(np.array(X_train)).to(device).float()
y_train = torch.from_numpy(np.array(y_train)).to(device).float()
X_test = torch.from_numpy(np.array(X_test)).to(device).float()
y_test = torch.from_numpy(np.array(y_test)).to(device).float()
# +
# torch.save(X_train,'X_train.pt')
# torch.save(X_test,'X_test.pth')
# torch.save(y_train,'y_train.pt')
# torch.save(y_test,'y_test.pth')
# torch.save(X,'X.pt')
# torch.save(X,'X.pth')
# torch.save(y,'y.pt')
# torch.save(y,'y.pth')
# +
# torch.save(X_words,'X_words.pt')
# torch.save(X_words,'X_words.pth')
# torch.save(data,'data.pt')
# torch.save(data,'data.pth')
# torch.save(y_words,'y_words.pt')
# torch.save(y_words,'y_words.pth')
# -
def get_accuracy(model,X,y):
accs = []
preds = model(X)
correct = 0
total = 0
for pred,yb in zip(preds,y):
for pred_in_pred,yb_in_yb in zip(pred,yb):
pred_in_pred = int(torch.argmax(pred_in_pred))
yb_in_yb = int(torch.argmax(yb_in_yb))
if pred_in_pred == yb_in_yb:
correct += 1
total += 1
acc = round(correct/total,3)*100
accs.append(acc)
print(accs)
print(yb_in_yb)
print(pred_in_pred)
acc = np.mean(accs)
return acc
def get_loss(model,X,y,criterion):
preds = model(X)
loss = criterion(preds,y)
return loss.item()
class Model(Module):
def __init__(self):
super().__init__()
self.activation = ReLU()
self.iters = 10
self.linear1 = Linear(len(X_words),256)
self.linear2 = Linear(256,256)
self.linear2bn = BatchNorm1d(256)
self.output = Linear(256,len(y_words))
def forward(self,X):
preds = self.linear1(X)
for _ in range(self.iters):
preds = self.activation(self.linear2bn(self.linear2(preds)))
preds = self.output(preds)
return preds
model = Model().to(device)
criterion = MSELoss()
optimizer = Adam(model.parameters(),lr=0.001)
batch_size = 32
epochs = 100
import wandb
PROJECT_NAME = 'Summarize-Text-Review'
wandb.init(project=PROJECT_NAME,name='baseline')
for _ in tqdm(range(epochs)):
for i in range(0,len(X_train),batch_size):
X_batch = X_train[i:i+batch_size].to(device)
y_batch = y_train[i:i+batch_size].to(device)
preds = model(X_batch)
loss = criterion(preds,y_batch)
optimizer.zero_grad()
loss.backward()
optimizer.step()
model.eval()
torch.cuda.empty_cache()
wandb.log({'Loss':get_loss(model,X_train,y_train,criterion)})
torch.cuda.empty_cache()
wandb.log({'Val Loss':get_loss(model,X_test,y_test,criterion)})
torch.cuda.empty_cache()
wandb.log({'Acc':get_accuracy(model,X_train,y_train)})
torch.cuda.empty_cache()
wandb.log({'Val Acc':get_accuracy(model,X_test,y_test)})
torch.cuda.empty_cache()
model.train()
wandb.finish()
torch.save(model,'model.pt')
torch.save(model,'model.pth')
torch.save(model.state_dict(),'model-sd.pt')
torch.save(model.state_dict(),'model-sd.pth')
| wandb/run-20210917_201500-362pocpc/tmp/code/00.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 実験B
import numpy as np
import scipy as sp
import scipy.constants as const
import matplotlib.pyplot as plt
import japanize_matplotlib
import pandas as pd
import sympy as sym
x=[1,2,3,4,5,6,7]
y=[0.085, 0.3, 0.77, 0.41, 0.33, 0.15, 0.11]
label = ['10:0', '9:1', '8:2', '7:3', '6:4', '5:5', '4:6']
plt.bar(x, y)
plt.ylabel('吸光度')
plt.xlabel('A液、B液の体積比')
plt.xticks(x, label)
plt.savefig('1.pdf')
plt.show()
from scipy import optimize
from sklearn.linear_model import LinearRegression as LR
bunsi = [10,15,20, 25, 37, 50, 75, 100, 150, 250][::-1]
kyori = np.array([22, 41, 67, 95, 155, 215, 318, 372, 457, 526])
x = np.array([85, 289, 437]) / kyori[0]
kyori = kyori / kyori[0]
reg = LR().fit(kyori[3:10].reshape((-1, 1)), np.log(bunsi)[3:10])
plt.yscale('log')
plt.xlim(0, 25)
plt.ylim(9, 300)
plt.scatter(kyori, bunsi)
X = np.linspace(3.5, 30)
plt.plot(X, np.exp(reg.predict(X.reshape((-1, 1)))))
plt.xlabel('相対移動度')
plt.ylabel('分子量 [kDa]')
plt.savefig('kenryo.pdf')
plt.show()
np.exp(reg.predict(x.reshape((-1, 1))))
a = {}
for i in range(len(bunsi) + 1):
for j in range(i + 2, len(bunsi) + 1):
a[(i, j)] = LR().fit(kyori[2:].reshape((-1, 1)), np.log(bunsi)[2:]).score(kyori[i:j].reshape((-1, 1)), np.log(bunsi)[i:j])
a[max(a, key=a.get)]
max(a, key=a.get)
a
# # 実験A
# 試験管番号、吸光度洗浄あり、洗浄なし
idx, a1, a2 = list(zip(*[
[16, 0.044, 0.045],
[15, 0.067, 0.061],
[14, 0.069, 0.072],
[13, 0.098, 0.099],
[12, 0.153, 0.172],
[11, 0.233, 0.288],
[10, 0.372, 0.445],
[9, 0.59, 0.716],
[8, 0.849, 0.918],
[7, 1.014, 0.877],
[6, 1.176, 0.595],
[5, 1.218, 0.411],
[4, 1.209, 0.299],
[3, 1.226, 0.22],
[2, 1.219, 0.157],
[1, 1.241, 0.118],
]))
# 濃度 ug / mL
c = 32 / (2 ** np.array(idx))
c[0] = 1e-10
c
# +
plt.scatter(c, a1, label='洗浄あり')
# plt.plot(X, np.poly1d(np.polyfit(np.log(c[1:-7]), a1[1:-7], 2))(np.log(X)))
X = np.logspace(-4, 2)
sig = lambda x, a, b, c, d: d / (1 + np.exp(-a * np.log(x) - b)) + c
param = sp.optimize.curve_fit(sig, c, a1)[0]
plt.plot(X, sig(X, *param))
plt.scatter(c, a2, label='洗浄なし')
plt.xlabel('ヒトIgG濃度 [ug / mL]')
plt.ylabel('吸光度')
plt.xscale('log')
plt.xlim((1e-4, 1e2))
plt.ylim((0, 1.3))
plt.legend()
plt.savefig('A-kenryo.pdf')
plt.show()
# -
sig = lambda x, a, b, c, d: d / (1 + np.exp(-a * np.log(x) - b)) + c
param = sp.optimize.curve_fit(sig, c, a1)
param
| example/jikken-report/results.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/bessx/aniclass/blob/master/AniClass.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="gt5_ryTQZJut" colab_type="text"
# # Performance Comparison of Convolutional Neural Networks for Animal Classification on Disproportionate Datasets
#
# by <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>
#
# Dataset downloader: https://github.com/bessx/aniclass
#
# [(•̀ᴗ•́)و ̑̑](http://git.bess.ai/aniclass)
# + [markdown] id="CL5IzoySxn61" colab_type="text"
# # Environment Setup
# + id="EPDtz0OEwCFq" colab_type="code" cellView="form" outputId="a3ed6ddc-2425-4fd2-a91d-814aa879d372" colab={"base_uri": "https://localhost:8080/", "height": 171}
#@title Download Image Datasets
# !wget -nc http://csc.lsu.edu/~bess/aniclass/test.tgz
# !wget -nc http://csc.lsu.edu/~bess/aniclass/teddy.tgz
# !wget -nc http://csc.lsu.edu/~bess/aniclass/train100.tgz
# !wget -nc http://csc.lsu.edu/~bess/aniclass/train1000.tgz
# !wget -nc http://csc.lsu.edu/~bess/aniclass/train10000.tgz
# + id="y4WnD64ZO8v5" colab_type="code" cellView="form" outputId="1bef3c50-1967-4872-9fb2-c4d65cfd5e72" colab={"base_uri": "https://localhost:8080/", "height": 225}
#@title Unpack Dataset Tars
# !tar xzkf test.tgz
# !tar xzkf teddy.tgz
# !tar xzkf train100.tgz
# !tar xzkf train1000.tgz
# !tar xzkf train10000.tgz
# + id="ZxePr-Sppn4f" colab_type="code" outputId="91c273e9-8795-4674-afe1-53a86887cd2b" cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 34}
#@title Import Commands
from keras.applications import xception, vgg16, inception_v3
# from keras.applications.resnet import ResNet101,ResNet152,ResNet50
# from keras.applications.densenet import DenseNet121,DenseNet169,DenseNet201
# from keras.applications.nasnet import NASNetMobile,NASNetLarge
#from keras.applications.vgg16 import VGG16
from keras.preprocessing import image
from keras.preprocessing.image import ImageDataGenerator, load_img, img_to_array
from keras.applications.vgg16 import preprocess_input, decode_predictions
from keras.applications.inception_v3 import InceptionV3
from keras.models import Model, Sequential, clone_model, load_model
from keras import layers, utils
from keras.layers import Dense, Activation, Flatten, add
from keras.layers.pooling import GlobalAveragePooling2D
from keras.utils.np_utils import to_categorical
from keras.optimizers import SGD
from keras.callbacks import History, ModelCheckpoint
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from PIL import ImageFile
ImageFile.LOAD_TRUNCATED_IMAGES = True
# %matplotlib inline
# + [markdown] id="MCus3K8By8Xk" colab_type="text"
# # Models Setup
# + id="bMX-8HHBqE54" colab_type="code" outputId="2c15d473-c773-4ba1-f147-04543b982744" colab={"base_uri": "https://localhost:8080/", "height": 51}
#@title Load Xception and freeze final layers
# Import Xception without bottom two layers
xcept_base = xception.Xception(include_top=False, weights='imagenet')
xcept_base.name = 'xcept_base'
# Create two layers with 10 classes in final prediction layer
x = xcept_base.output
x = GlobalAveragePooling2D()(x)
predictions = Dense(10, activation='softmax')(x)
# Add the top layer block to the base model
xcept_train = Model(xcept_base.input, predictions)
xcept_train.name = 'xcept'
# Freeze all but final two layers of xcept model
for ind, layer in enumerate(xcept_train.layers):
if ind >= (len(xcept_train.layers)-2):
layer.trainable = True
# print(ind, layer, "if")
else:
layer.trainable = False
# print(ind, layer, "else")
# + id="z5YzIQSHxxsx" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="014ef185-07c2-4b8a-907c-7edd267e1a21"
#@title Load Inception and freeze final layers
# Import Inception without bottom two layers
incept_base = inception_v3.InceptionV3(include_top=False, weights='imagenet')
incept_base.name = 'incept_base'
# Create two layers with 10 classes in final prediction layer
y = incept_base.output
y = GlobalAveragePooling2D()(y)
predictions = Dense(10, activation='softmax')(y)
# Add the top layer block to the base model
incept_train = Model(incept_base.input, predictions)
incept_train.name = 'incept'
#@title Freeze all but final two layers of incept model
for ind, layer in enumerate(incept_train.layers):
if ind >= (len(incept_train.layers)-2):
layer.trainable = True
# print(ind, layer, "if")
else:
layer.trainable = False
# print(ind, layer, "else")
# + id="miIxpooG0BZu" colab_type="code" colab={}
#@title Load VGG16 and freeze final layers
vgg16_base = vgg16.VGG16(include_top=True, weights='imagenet')
vgg16_base.name = 'vgg16_pre'
vgg16_input = vgg16_base.get_layer(index=0).input
vgg16_output = vgg16_base.get_layer(index=-2).output
vgg16_model = Model(inputs=vgg16_input, outputs=vgg16_output)
vgg16_model.name = 'vgg16_base'
vgg16_train = Sequential()
vgg16_train.add(vgg16_model)
vgg16_train.add(Dense(10, activation='softmax', input_dim=4096, name='predictions'))
vgg16_train.name = 'vgg16'
# Freeze all but final two layers of VGG16 model
for ind, layer in enumerate(vgg16_train.layers):
if ind >= (len(vgg16_train.layers)-1):
layer.trainable = True
# print(ind, layer, "if")
else:
layer.trainable = False
# print(ind, layer, "else")
# + id="3QeGrSuwtJWy" colab_type="code" colab={}
# Save Models to List
model_list = [xcept_train, vgg16_train, incept_train]
# Make Models Folder
# !mkdir models
# Make Results Folder
# !mkdir results
# Save Models to Folder
for mdl in model_list:
mdl.save('models/' + mdl.name + '_train.h5')
print('Saved ' + mdl.name + '_train to disk')
# + [markdown] id="NMe12PFaxhHP" colab_type="text"
# # Retrain Last Layers
# + id="oskPzF6HI-fI" colab_type="code" cellView="form" colab={}
#@title Set Variables for Training: { run: "auto" }
trainmodel = incept_train #@param ["xcept_train", "incept_train", "vgg16_train"] {type:"raw"}
traindir = 'train100' #@param ["train100", "train1000", "train10000"]
img_height = 299 #@param {type:"integer"}
img_width = 299 #@param {type:"integer"}
batch_size = 50 #@param {type:"integer"}
nb_epochs = 20 #@param {type:"integer"}
# Set Save Name
sname = trainmodel.name + '_' + traindir
# + id="qDzo-0f3kdxB" colab_type="code" cellView="both" outputId="f8ec09a7-3247-4953-d6ce-544be32329a9" colab={"base_uri": "https://localhost:8080/", "height": 1000}
#@title Retrain final 2 layers of model { vertical-output: true }
print('Loading model ' + trainmodel.name + '...')
model = load_model('models/' + trainmodel.name + '_train.h5')
train_datagen = ImageDataGenerator(rescale=1./255,
# shear_range=0.2,
# zoom_range=0.2,
# horizontal_flip=True,
validation_split=0.15) # set validation split
train_generator = train_datagen.flow_from_directory(
traindir,
target_size=(img_height, img_width),
batch_size=batch_size,
class_mode='binary',
subset='training') # set as training data
validation_generator = train_datagen.flow_from_directory(
traindir, # same directory as training data
target_size=(img_height, img_width),
batch_size=batch_size,
class_mode='binary',
subset='validation') # set as validation data
test_datagen = ImageDataGenerator(rescale=1./255)
test_generator = test_datagen.flow_from_directory(
'test',
target_size=(img_height, img_width),
batch_size=batch_size,
class_mode='binary')
# Fix for Corrupt Images
def my_gen(gen):
while True:
try:
data, labels = next(gen)
yield data, labels
except:
pass
# Learning rate is changed to 0.001
sgd = SGD(lr=1e-3, decay=1e-6, momentum=0.9, nesterov=True)
model.compile(optimizer=sgd, loss='sparse_categorical_crossentropy', metrics=['acc'])
# Save accuracy to history
filepath = 'models/' + sname + ".h5"
cblist = [
History(),
ModelCheckpoint(filepath, monitor='val_acc', verbose=1, save_best_only=True)
]
train = model.fit_generator(
my_gen(train_generator),
steps_per_epoch = train_generator.samples // batch_size,
validation_data = my_gen(validation_generator),
validation_steps = validation_generator.samples // batch_size,
epochs = nb_epochs,
callbacks=cblist)
# Print model evaluation and save to results
meval = model.evaluate_generator(test_generator, verbose=1)
mev = 'test loss: %.5f - test acc: %.2f%%' % (meval[0], meval[1]*100)
print(mev)
# %store meval > models/"$sname"_eval.txt
# %store mev > results/"$sname"_ev.txt
# + id="O1ELb6v7UKIe" colab_type="code" outputId="3f61201a-4473-48d7-ee1d-fb341e776606" colab={"base_uri": "https://localhost:8080/", "height": 34}
# plt.clf()
# plt.close('all')
# + id="w0rNBeOH47dx" colab_type="code" outputId="ca3df425-7391-4cee-8b56-665002e08537" cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 608}
#@title Plot accuracy and loss and save results
# summarize history for accuracy
plt.plot(train.history['acc'])
plt.plot(train.history['val_acc'])
plt.title('Model Accuracy')
plt.ylabel('Accuracy')
plt.xlabel('Epoch')
plt.xticks(range(0, nb_epochs))
plt.legend(['Training', 'Validation'], loc='lower right')
plt.savefig('results/' + sname + '_acc.png', bbox_inches='tight')
plt.show()
# summarize history for loss
plt.plot(train.history['loss'])
plt.plot(train.history['val_loss'])
plt.title('Model Loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.xticks(range(0, nb_epochs))
plt.legend(['Training', 'Validation'], loc='upper right')
plt.savefig('results/' + sname + '_loss.png', bbox_inches='tight')
plt.show()
# build history dataframe and save
hist_df = pd.DataFrame(train.history)
# %store train.history > results/"$sname"_history.txt
# %store hist_df > results/"$sname"_histdf.txt
# + id="eph_fa6-YHda" colab_type="code" outputId="53ab4480-ca67-4e23-c695-1b861b872720" colab={"base_uri": "https://localhost:8080/", "height": 669}
hist_df
# + id="dIi2Z899I7g8" colab_type="code" cellView="form" outputId="2a8ebb38-1d32-4bda-e379-c76d985e21a5" colab={"base_uri": "https://localhost:8080/", "height": 69}
#@title Predictions with current model of directory: { vertical-output: true }
pred_model = 'incept_train100' #@param ['xcept_train100','xcept_train1000','xcept_train10000', 'vgg16_train100', 'vgg16_train1000', 'vgg16_train10000','incept_train100','incept_train1000','incept_train10000']
pred_dir = 'test' #@param ['test', 'train100','train1000', 'train10000', 'teddy']
print('Loading model models/' + pred_model + '.h5...')
model = load_model('models/' + pred_model + '.h5')
pred_datagen = ImageDataGenerator(rescale=1./255)
pred_generator = test_datagen.flow_from_directory(
pred_dir,
target_size=(img_height, img_width),
batch_size=batch_size,
class_mode='binary')
pred = model.predict_generator(pred_generator, pred_generator.samples // batch_size, verbose=1)
# + id="7obtE3uLQSEj" colab_type="code" cellView="form" outputId="f27c17ec-382d-446a-b38d-c90cde925111" colab={"base_uri": "https://localhost:8080/", "height": 514}
#@title Create Confusion Matrix from Predictions { vertical-output: true }
import seaborn as sn
from sklearn.metrics import confusion_matrix
y_true = pred_generator.classes
y_pred = np.argmax(pred, axis=-1)
cm = confusion_matrix(y_true, y_pred)
labels = ['bear', 'bird','cat', 'cow', 'dog', 'elephant', 'giraffe', 'other',
'sheep', 'zebra']
df_cm = pd.DataFrame(cm, index = [i for i in labels],
columns = [i for i in labels])
df_cm.index.name = 'Actual'
df_cm.columns.name = 'Predicted'
plt.figure(figsize = (10,7))
sn.heatmap(df_cm, annot=True, annot_kws={"size": 16})
# sn.set(font_scale=1.4)
plt.savefig('results/'+ pred_model + '_pred_' + pred_dir + '_cm.png', bbox_inches='tight')
print('results/'+ pred_model + '_pred_' + pred_dir + '_cm.png')
plt.show()
# + id="Muli3dG_LsJC" colab_type="code" outputId="d9fd8c53-1f9a-426f-8fb6-6022ad6cd870" colab={"base_uri": "https://localhost:8080/", "height": 111}
#@title Predictions as Pandas Dataframe
pred_files = pred_generator.filenames
pred_df = pd.DataFrame(pred)
pred_df.columns = ['bear', 'bird','cat', 'cow', 'dog', 'elephant', 'giraffe', 'other', 'sheep', 'zebra']
# [i.split('\t', 1)[0] for i in l]
pred_df['file'] = pred_files
pred_df.insert(0, 'class', [item.split('/')[0] for item in pred_files])
pred_df
# Average of predicted class by actual class
class_preds = pred_df.groupby('class').mean().round(5).astype(object)*100
class_preds
# + id="JDjeQHxkIPkA" colab_type="code" outputId="d1631274-2d5d-436d-8129-58a33388fb5c" colab={"base_uri": "https://localhost:8080/", "height": 51}
# %store pred_df > results/"$pred_model"_teddy_all.txt
# %store class_preds > results/"$pred_model"_teddy.txt
# + [markdown] id="2WThzzssF5vm" colab_type="text"
# # Results File Download
# + id="r2IuPwLf2MC8" colab_type="code" cellView="form" outputId="fb4845c4-8649-4323-f8c2-881fb34b9f75" colab={"base_uri": "https://localhost:8080/", "height": 595}
#@title Save results to csc server:
#@markdown http://csc.lsu.edu/~bess/aniclass/results
# !rm results.tgz
# !tar czvf results.tgz results/
from google.colab import files
files.download('results.tgz')
# + [markdown] id="2LFDFyQqUW3I" colab_type="text"
# # The following code is for individual image and model tests
# + id="_F7pj2zCEC_N" colab_type="code" colab={}
# Get Test Images
# !wget -nc -q -P example_images/ https://upload.wikimedia.org/wikipedia/commons/f/f9/Zoorashia_elephant.jpg
# !wget -nc -q http://farm1.static.flickr.com/145/430300483_21e993670c.jpg -O example_images/eagle.jpg
# + id="CFafrbq9Dnjs" colab_type="code" colab={}
# Test Images Individually
# load an image from file
image = load_img('example_images/eagle.jpg', target_size=(224, 224))
image = img_to_array(image)
# reshape data for the model
image = image.reshape((1, image.shape[0], image.shape[1], image.shape[2]))
# prepare the image for the VGG model
image = preprocess_input(image)
features = vgg16_pre.predict(image)
# Print Predictions
print('Predictions:')
preds = decode_predictions(features, top=5)[0]
for p in preds:
print('%s: %s (%.2f%%)' % (p[0], p[1], p[2]*100))
# + id="Tvx4sHdZGxt7" colab_type="code" colab={}
def show_batch(image_batch, label_batch):
plt.figure(figsize=(10,10))
for n in range(25):
ax = plt.subplot(5,5,n+1)
plt.imshow(image_batch[n])
plt.title(CLASS_NAMES[label_batch[n]==1][0].title())
plt.axis('off')
# + id="KjZQLcsAETop" colab_type="code" outputId="55d242ea-5c1f-4d95-c6b5-b2a9b9734d70" colab={"base_uri": "https://localhost:8080/", "height": 51}
import pathlib
# data_dir = utils.get_file(origin='https://storage.googleapis.com/download.tensorflow.org/example_images/flower_photos.tgz',fname='flower_photos', untar=True)
data_dir = pathlib.Path('ImageNetOther/')
image_count = len(list(data_dir.glob('*/*.jpg')))
image_count
CLASS_NAMES = np.array([item.name for item in data_dir.glob('*') if item.name != "LICENSE.txt"])
CLASS_NAMES
image_generator = ImageDataGenerator(rescale=1./255)
BATCH_SIZE = 32
IMG_HEIGHT = 224
IMG_WIDTH = 224
STEPS_PER_EPOCH = np.ceil(image_count/BATCH_SIZE)
train_data_gen = image_generator.flow_from_directory(directory=str(data_dir),
batch_size=BATCH_SIZE,
shuffle=True,
target_size=(IMG_HEIGHT, IMG_WIDTH),
classes = list(CLASS_NAMES))
test_generator = image_generator.flow_from_directory(directory=str(data_dir),
target_size=(IMG_HEIGHT, IMG_WIDTH),
batch_size=1,
class_mode=None,
shuffle=False,
seed=42
)
# test_gen = datagen.flow_from_directory('/content/drive/My Drive/CSC7333/test/', target_size=(224,224), class_mode='binary')
# test_samples = len(test_gen.filenames)
# + id="Ej9je0T6E0SP" colab_type="code" colab={}
image_batch, label_batch = next(train_data_gen)
show_batch(image_batch, label_batch)
# + id="eVWD1QbuIrkq" colab_type="code" colab={}
print(test_generator.filenames[82])
print(test_generator.filenames[83])
# + id="nZiYefW-HQqL" colab_type="code" colab={}
STEP_SIZE_TEST = test_generator.n//test_generator.batch_size
test_generator.reset()
pred = ovgg16.predict_generator(test_generator, steps=STEP_SIZE_TEST, verbose=1)
# predicted_class_indices=np.argmax(pred,axis=1)
# + id="CjzGdppoJb8P" colab_type="code" colab={}
predicted_class_indices=np.argmax(pred,axis=1)
print(predicted_class_indices)
# + id="cV5KN-tpJyt6" colab_type="code" colab={}
# Print Predictions
print('Predictions:')
preds = decode_predictions(pred, top=5)[0]
for p in preds:
print('%s: %s (%.2f%%)' % (p[0], p[1], p[2]*100))
| AniClass.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Raster data
#
# ```{admonition} Learning Objectives
# *A 30 minute guide to raster data for SnowEX*
# - find, visualize, interpret raster data formats
# - reproject rasters to different coordinate reference systems
# - use Python raster libraries [rasterio](https://github.com/mapbox/rasterio) and [rioxarray](https://corteva.github.io/rioxarray)
# ```
# ## Raster Basics
#
# *Raster data is stored as a grid of values which are rendered on a map as pixels. Each pixel value represents an area on the Earth's surface.* Pixel values can be continuous (elevation) or categorical (land use). This data structure is very common - jpg images on the web, photos from your digital camera. A geospatial raster is only unique in that it is accompanied by metadata that connects the pixel grid to a location on Earth's surface.
#
# ### Coordinate Reference System or "CRS"
# This specifies the datum, projection, and additional parameters needed to place the raster in geographic space. For a dedicated lesson on CRSs, see: https://datacarpentry.org/organization-geospatial/03-crs/index.html
#
# The natural representation of an image in programming is as an array, or matrix, with accompanying metadata to keep track of the geospatial information such as CRS.
# +
# It's good practice to keep track of the versions of various packages you are using
import rioxarray
import xarray as xr
import rasterio
import numpy as np
import os
print('rioxarray version:', rioxarray.__version__)
print('xarray version:', xr.__version__)
print('rasterio version:', rasterio.__version__)
# Work in a temporary directory
os.chdir('/tmp')
# Plotting setup
import matplotlib.pyplot as plt
# %config InlineBackend.figure_format='retina'
#plt.rcParams.update({'font.size': 16}) # make matplotlib font sizes bigger
# + language="bash"
#
# # Retrieve a copy of data files used in this tutorial from Zenodo.org:
# # Re-running this cell will not re-download things if they already exist
#
# mkdir -p /tmp/tutorial-data
# cd /tmp/tutorial-data
# wget -q -nc -O data.zip https://zenodo.org/record/5504396/files/geospatial.zip
# unzip -q -n data.zip
# rm data.zip
# -
# ## Elevation rasters
#
#
# Let's compare a few elevation rasters over Grand Mesa, CO.
#
# ### NASADEM
# First, let's look at NASADEM, which uses data from NASA's Shuttle Radar Topography Mission from February 11, 2000, but also fills in data gaps with the ASTER GDEM product.
# Read more about this data set at https://lpdaac.usgs.gov/products/nasadem_hgtv001/. You can rind URLs to data via NASA's Earthdata search https://search.earthdata.nasa.gov:
#
# * https://e4ftl01.cr.usgs.gov//DP132/MEASURES/NASADEM_HGT.001/2000.02.11/NASADEM_HGT_n39w109.zip
#
# +
# #%%bash
# Get data directly from NASA LPDAAC and unzip
#DATADIR='/tmp/tutorial-data/geospatial/raster'
# #mkdir -p ${DATADIR}
#wget -q -nc https://e4ftl01.cr.usgs.gov/DP132/MEASURES/NASADEM_HGT.001/2000.02.11/NASADEM_HGT_n39w109.zip
#unzip -n NASADEM_HGT_n39w109.zip -d ${DATADIR}
# -
# ### Rasterio
#
# [rasterio](https://rasterio.readthedocs.io/en/latest/) is a foundational geospatial Python library to work with raster data. It is a Python library that builds on top of the [Geospatial Data Abstraction Library (GDAL)](https://gdal.org), a well-established and critical geospatial library that underpins most GIS software.
# Open a raster image in a zipped archive
# https://rasterio.readthedocs.io/en/latest/topics/datasets.html
path = 'zip:///tmp/tutorial-data/geospatial/NASADEM_HGT_n39w109.zip!n39w109.hgt'
with rasterio.open(path) as src:
print(src.profile)
# We can read this raster data as a numpy array to perform calculations
with rasterio.open(path) as src:
data = src.read(1) #read first band
print(type(data))
plt.imshow(data, cmap='gray')
plt.colorbar()
plt.title(f'{path} (m)')
# Rasterio has a convenience function for plotting with geospatial coordinates
import rasterio.plot
with rasterio.open(path) as src:
rasterio.plot.show(src)
# +
# rasterio knows about the 2D geospatial coordinates, so you can easily take a profile at a certain latitude
import rasterio.plot
latitude = 39.3
longitude = -108.8
with rasterio.open(path) as src:
row, col = src.index(longitude, latitude)
profile = data[row,:] # we already read in data earlier
plt.plot(profile)
plt.xlabel('column')
plt.ylabel('elevation (m)')
plt.title(path)
# -
# ### Rioxarray
#
# As the volume of digital data grows, it is increasingly common to work with n-dimensional data or "datacubes". The most basic example of a data cube is a stack of co-located images over time. Another example is multiband imagery where each band is acquired simultaneously. You can find a nice walk-through of concepts in this documentation from the [OpenEO project](https://openeo.org/documentation/1.0/datacubes.html). The Python library [Xarray](http://xarray.pydata.org/en/stable/) is designed to work efficiently with multidimensional datasets, and the extension [RioXarray](https://corteva.github.io/rioxarray/stable/) adds geospatial functionality such as reading and writing GDAL-supported data formats, CRS management, reprojection, etc.
#
# Rioxarray depends on rasterio for functionality (which in turn depends on GDAL), so you can see that it's software libraries all the way down!
da = rioxarray.open_rasterio(path, masked=True)
da
# ```{note}
# We have an `xarray DataArray`, which is convenient for datasets of one variable (in this case, elevation). The `xarray Dataset` is a related data intended for multiple data variables (elevation, precipitation, snow depth etc.), Read more about xarray datastructures in the [documentation](http://xarray.pydata.org/en/stable/user-guide/data-structures.html).
# ```
# Drop the 'band' dimension since we don't have multiband data
da = da.squeeze('band', drop=True)
da.name = 'nasadem'
da
# the rioxarray 'rio' accessor gives us access to geospatial information and other methods
print(da.rio.crs)
print(da.rio.encoded_nodata)
# xarray, like rasterio has built-in convenience plotting with matplotib
# http://xarray.pydata.org/en/stable/user-guide/plotting.html
da.plot();
# xarray is also integrated into holoviz plotting tools
# which are great for interactive data exploration in a browser
import hvplot.xarray
da.hvplot.image(x='x', y='y', rasterize=True, cmap='gray',
aspect=1/np.cos(np.deg2rad(39)))
# Xarray uses pandas-style indexing to select data. This makes it very easy to plot a profile
profile = da.sel(y=39.3, method='nearest')
profile.hvplot.line(x='x')
# Finally, reproject this data to a different CRS and save it
# https://epsg.io/26912
da_reproject = da.rio.reproject("EPSG:26912")
da_reproject
da_reproject.hvplot.image(x='x', y='y', rasterize=True, cmap='gray',
aspect=1, # NOTE: we change this since we're in UTM now
)
# We can easily save the entire raster or a subset
da_reproject.rio.to_raster('n39w109_epsg26912.tif', driver='COG') #https://www.cogeo.org
# Load the saved dataset back in
#ds = xr.open_dataset('n39w109_epsg26912.tif', engine='rasterio')
# ## Comparing rasters
#
# ### Copernicus DEM
# Second, let's get rasters from the European Space Agency Copernicus DEM, which is available as a public dataset on AWS: https://registry.opendata.aws/copernicus-dem/. This is a global digital elevation model derived from the TanDEM-X Synthetic Aperture Radar Mission, see [ESA's website for product details](https://spacedata.copernicus.eu/explore-more/news-archive/-/asset_publisher/Ye8egYeRPLEs/blog/id/434960)
#
# * s3://copernicus-dem-30m/Copernicus_DSM_COG_10_N39_00_W109_00_DEM/Copernicus_DSM_COG_10_N39_00_W109_00_DEM.tif
# Can use AWS CLI to interact with this open data
# !aws --no-sign-request s3 ls s3://copernicus-dem-30m/Copernicus_DSM_COG_10_N39_00_W109_00_DEM/Copernicus_DSM_COG_10_N39_00_W109_00_DEM.tif
# +
# Rasterio has some capabilities to read URLs in addition to local file paths
url = 's3://copernicus-dem-30m/Copernicus_DSM_COG_10_N39_00_W109_00_DEM/Copernicus_DSM_COG_10_N39_00_W109_00_DEM.tif'
# need to use environment variables to change default GDAL settings when reading URLs
Env = rasterio.Env(GDAL_DISABLE_READDIR_ON_OPEN='EMPTY_DIR',
AWS_NO_SIGN_REQUEST='YES')
# NOTE: this reads metadata only
with Env:
with rasterio.open(url) as src:
print(src.profile)
# +
# Because rioxarray uses rasterio & gdal it can also read urls
with Env:
daC = rioxarray.open_rasterio(url).squeeze('band', drop=True)
daC.name = 'copernicus_dem'
# NOTE: this data doesn't have NODATA set in metadata, so let's use the same value as SRTM
daC.rio.write_nodata(-32768.0, encoded=True, inplace=True)
daC
# -
daC.hvplot.image(x='x', y='y', rasterize=True, cmap='gray',
aspect=1/np.cos(np.deg2rad(latitude)))
# +
# Ensure the grid of one raster exactly matches another (same projection, resolution, and extents)
# NOTE: these two rasters happen to already be on an aligned grid
# There are many options for how to resample a warped raster grid (nearest, bilinear, etc)
#print(list(rasterio.enums.Resampling))
daR = daC.rio.reproject_match(da, resampling=rasterio.enums.Resampling.nearest)
daR
# -
difference = daR - da
difference.name = 'cop30 - nasadem'
plot = difference.hvplot.image(x='x', y='y', rasterize=True, cmap='bwr', clim=(-50,50),
aspect=1/np.cos(np.deg2rad(latitude)))
plot
import holoviews as hv
mean = difference.mean().values
print(mean)
mean_hline = hv.VLine(mean)
# Composite plot with holoviz (histogram with line overlay)
difference.hvplot.hist(bins=100, xlim=(-50,50), color='gray') * mean_hline.opts(color='red')
# ```{warning}
#
# NOTE that both the NASA DEM and Copernicus DEM report a CRS of `CRS.from_epsg(4326)`. This is the 2D horizontal coordinate reference. Elevation values also are with respect to a reference surface known as a datum, which is commonly an ellipsoid representation of the Earth, or a spatially varying "geoid" which is an equipotential surface. For high-precision geodetic applications it is common to use even more accurate time-varying [3D coordinate reference systems](https://proj.org/usage/transformation.html#).
#
# For NASADEM, according to the documentation page the datum is WGS84/EGM96, where 'EGM96' is the 'Earth Geoid Model from 1996'. But the Copernicus DEM uses EGM2008, a slightly updated model. Often, GNSS elevation datasets and SAR products are relative to elliposid heights (EPSG:4979), and if you do not convert between these systems you might end up with elevation discrepancies on the order of 100 meters!
#
# Datum shift grids can be used to vertically shift rasters and can be read about here https://proj.org/usage/network.html
# ```
with Env:
daS = rioxarray.open_rasterio('https://cdn.proj.org/us_nga_egm96_15.tif').squeeze(dim='band', drop=True)
daS.name = 'us_nga_egm96_15'
# +
# World - WGS 84 (EPSG:4979) to EGM96 height (EPSG:5773). Size: 2.6 MB. Last modified: 2020-01-24
from cartopy import crs
daS.hvplot.image(x='x', y='y', rasterize=True,
geo=True, global_extent=True, coastline=True,
cmap='bwr', projection=crs.Mollweide(),
title='WGS 84 (EPSG:4979) to EGM96 height (EPSG:5773)')
# -
# What is the approximate grid shift at a specific location?
daS.sel(x=longitude, y=latitude, method='nearest').values
# ```{admonition} execercises
# - find longitude, latitude point of maximum EPSG:4979 -> EPSG:5773 datum shift magnitude
# - convert both elevation datasets to ellipsoid height (EPSG:4979) using data shift grids
# - save a small subset of a raster dataset
# ```
| book/tutorials/geospatial/raster.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Lecture 3
#
# ## Differentiation I:
#
# ### Introduction and Interpretation
# + slideshow={"slide_type": "skip"}
import numpy as np
##################################################
##### Matplotlib boilerplate for consistency #####
##################################################
from ipywidgets import interact
from ipywidgets import FloatSlider
from matplotlib import pyplot as plt
# %matplotlib inline
from IPython.display import set_matplotlib_formats
set_matplotlib_formats('svg')
global_fig_width = 10
global_fig_height = global_fig_width / 1.61803399
font_size = 12
plt.rcParams['axes.axisbelow'] = True
plt.rcParams['axes.edgecolor'] = '0.8'
plt.rcParams['axes.grid'] = True
plt.rcParams['axes.labelpad'] = 8
plt.rcParams['axes.linewidth'] = 2
plt.rcParams['axes.titlepad'] = 16.0
plt.rcParams['axes.titlesize'] = font_size * 1.4
plt.rcParams['figure.figsize'] = (global_fig_width, global_fig_height)
plt.rcParams['font.sans-serif'] = ['Computer Modern Sans Serif', 'DejaVu Sans', 'sans-serif']
plt.rcParams['font.size'] = font_size
plt.rcParams['grid.color'] = '0.8'
plt.rcParams['grid.linestyle'] = 'dashed'
plt.rcParams['grid.linewidth'] = 2
plt.rcParams['lines.dash_capstyle'] = 'round'
plt.rcParams['lines.dashed_pattern'] = [1, 4]
plt.rcParams['xtick.labelsize'] = font_size
plt.rcParams['xtick.major.pad'] = 4
plt.rcParams['xtick.major.size'] = 0
plt.rcParams['ytick.labelsize'] = font_size
plt.rcParams['ytick.major.pad'] = 4
plt.rcParams['ytick.major.size'] = 0
##################################################
# + [markdown] slideshow={"slide_type": "slide"}
# ## Gradients
#
# We often want to know about the *rate* at which one quantity changes over time.
# Examples:
# 1. The rate of disappearance of substrate with time in an enzyme reaction.
# 1. The rate of decay of a radioactive substance (how long will it have activity above a certain level?)
# 1. The rate of bacterial cell growth over time.
# 1. How quickly an epidemic is growing.
#
# + [markdown] slideshow={"slide_type": "slide"}
# ### Defining the gradient
#
# * The **gradient of a curve** at a point $P$ is the slope of the tangent of the curve at that point.
# * The **tangent** is the line that "just touches" (but doesn't cross) the curve.
# * The gradient is also known as the **rate of change** or **derivative**, and the process of finding the gradient is called **differentiation**.
# * The gradient of the curve $\;y = f(x)\;$ is denoted in a few different ways, the three most common are:
#
# $$ y', \quad f'(x), \quad \frac{dy}{dx}. $$
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Example, $y = x^2$
# + slideshow={"slide_type": "skip"}
x1_widget = FloatSlider(value=1.0, min=-3., max=3., step=0.2, continuous_update=False)
_x = np.linspace(-5,5,50)
def add_line(x):
plt.title('$y=x^2$')
plt.xlabel('$x$')
plt.ylabel('$y=x^2$')
plt.xlim((-4.,4.))
plt.ylim((-5.,17.))
plt.plot(_x, _x**2);
plt.plot([x-10., x, x+10.], [x*x-20.*x, x*x, x*x+20.*x]);
plt.plot(x, x*x, 'ko')
# -
interact(add_line, x=x1_widget, continuous_update=False);
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Example, $y = \log(x)$
# + slideshow={"slide_type": "skip"}
x_widget = FloatSlider(value=1.0, min=.4, max=1.8, step=0.05, continuous_update=False)
_x2 = np.linspace(0.2,2.,50)
def add_line(x):
plt.title('$y=log(x)$')
plt.xlabel('$x$')
plt.ylabel('$y=\log(x)$')
plt.xlim((0.2,2.))
plt.ylim((-2.,1.))
plt.plot(_x2, np.log(_x2));
plt.plot([x-10., x, x+10.], [np.log(x)-10./x, np.log(x), np.log(x)+10./x]);
plt.plot(x, np.log(x), 'ko')
# + slideshow={"slide_type": "subslide"}
interact(add_line, x=x_widget, continuous_update=False);
# + [markdown] slideshow={"slide_type": "slide"}
# ### Algebraic example
#
#
# If we want to find $y'(x)$ for $y = x^3 + 2$:
#
#
# $$ \text{Gradient} = \frac{y_2 - y_1}{x_2-x_1} = \frac{\Delta y}{\Delta x}$$
#
#
# Try with
#
# $x_1 = 1.5,\;1.9,\;1.99,\;\ldots$
#
# $x_2 = 2.5,\;2.1,\;2.01,\;\ldots$
# + slideshow={"slide_type": "subslide"}
x_1 = 1.5; x_2 = 2.5
y_1 = x_1**3 + 2; y_2 = x_2**3 + 2
print((y_2-y_1)/(x_2-x_1))
x_1 = 1.9; x_2 = 2.1
y_1 = x_1**3 + 2; y_2 = x_2**3 + 2
print((y_2-y_1)/(x_2-x_1))
x_1 = 1.99; x_2 = 2.01
y_1 = x_1**3 + 2; y_2 = x_2**3 + 2
print((y_2-y_1)/(x_2-x_1))
# + [markdown] slideshow={"slide_type": "subslide"}
# As the difference between $x_1$ and $x_2$ gets smaller, the gradient stabilises. The value it converges to is the gradient at the midway point of $x_1$ and $x_2$.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Calculating gradients exactly
#
# $\text{Gradient} \approx \frac{\Delta y}{\Delta x} = \frac{f(x+h) - f(x)}{h}$
#
# This is called a finite difference approximation to the gradient. The approximation becomes more accurate the smaller h is.
#
# When using the approximation, we denote the changes as $\frac{\Delta y}{\Delta x}$, in the limit as h goes to 0, this becomes $\frac{dy}{dx}$.
#
# In this way, $\frac{d}{dx}$ is an operator, acting on $y$.
#
# Note, the $d$s cannot be cancelled out, as they aren't variables, they denote an infinitely small change.
# + slideshow={"slide_type": "skip"}
h_widget = FloatSlider(value=5.0, min=0.05, max=9., step=0.05, continuous_update=False)
_x3 = np.linspace(-2,11,50)
def add_line(h):
plt.title('$y=x^2$')
plt.xlabel('$x$')
plt.ylabel('$y=x^2$')
plt.xlim((-2.,11.))
plt.ylim((-15.,121.))
plt.plot(_x3, _x3**2);
plt.plot([-8., 12.], [-36., 44.]);
plt.plot([12, -8], [4. + 10.*((2+h)**2-4)/h, 4. - 10.*((2+h)**2-4)/h]);
plt.plot([2., 2.+h], [4., (2.+h)**2], 'ko')
# + slideshow={"slide_type": "subslide"}
interact(add_line, h=h_widget, continuous_update=False);
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Example
#
# Find the gradient of $y = f(x) = x^3 + 2$.
#
# $\frac{dy}{dx} = \frac{f(x+h) - f(x)}{h}$
#
# $\frac{dy}{dx} = \frac{(x+h)^3 + 2 - (x^3 + 2)}{h}$
#
# $\frac{dy}{dx} = \frac{x^3 + 3x^2 h + 3xh^2 + h^3 + 2 - x^3 - 2}{h}$
#
# $\frac{dy}{dx} = \frac{3x^2h + 3xh^2 + h^3}{h}$
#
# $\frac{dy}{dx} = 3x^2 + 3xh + h^3$
#
# Now this is only exactly right when $h \rightarrow 0$. So letting that happen, we have
# $\frac{dy}{dx} = 3x^2$
# + [markdown] slideshow={"slide_type": "slide"}
# ## Derivative of polynomial functions
# Using techniques like the one above (which is called differentiation from first principles), one can generalise the connection between powers of $x$ and their derivatives:
#
# If $y = a x^n$, then its **derivative** is
# $\frac{dy}{dx} = y'(x) = a n x^{n-1}$
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Examples to try
# 1. $y = x^4$
# 2. $y = 7x^5$
# 3. $y = x^{-2} = \frac{1}{x^2}$
# 4. $y = \sqrt{1/x} = (1/x)^{1/2} = x^{-1/2}$
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Summing and multiplying derivatives
# ### Summing
#
# $(f(x) \pm g(x))' = f'(x) \pm g'(x)$
#
# e.g.
#
# $y = x^2 + x^3, \quad y' = 2x + 3x^2$
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Multiplying (by a scalar)
# $ (a f(x))' = a f'(x)$
#
# e.g.
#
# $y = 6x^3, \quad y' = 6 \cdot 3x^2 = 18 x^2$
#
# **This only works for scalars**.
#
# In most circumstances $(f(x) g(x))' \neq f(x)' g(x)'$
#
# e.g.
#
# $y = x\cdot x = x^2, \quad y' \neq 1$
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Higher-order derivatives
# You can take a derivative of a function multiple times in a row. This is usually denoted either $y''(x),\;\;f''(x)\;$ or $\;\frac{d^2 y}{dx^2}\;$ for second-order derivatives (differentiating twice), and similar for higher orders.
#
# e.g.
#
# $y = x^3$
#
# $y' = 3x^2$
#
# $y'' = \frac{d^2 y}{dx^2} = 6 x$
# + [markdown] slideshow={"slide_type": "slide"}
# ## Interpreting derivatives:
#
# The sign of the first derivative $\;f'(x)\;$ tells us how $\;f(x)\;$ is growing
#
# - Positive gradient: If $\;y' > 0\;$ then $\;y\;$ is **increasing** at $\;x\;$
# - Negative gradient: If $\;y' < 0\;$ then $\;y\;$ is **decreasing** at $\;x\;$
# - Zero gradient: If $\;y' = 0\;$ then $\;y\;$ is not changing (flat) at $\;x\;$
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Extreme values (turning points and points of inflection)
# (a) Local maximum: $\;\frac{dy}{dx} = 0,\;$ and $\;\frac{d^2y}{dx^2} < 0\;$
#
# (b) Local minimum: $\;\frac{dy}{dx} = 0,\;$ and $\;\frac{d^2y}{dx^2} > 0\;$
#
# (c) Inflection: $\;\frac{d^2y}{dx^2} = 0\;$
#
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Example: Find the stationary points of $\;y = 2x^3 - 5x^2 - 4x\;$
# To do this, we need to know both $\;y'(x)\;$ and $\;y''(x)\;$.
#
# $y'(x) = 6x^2 - 10x - 4$
#
# $y''(x) = 12x - 10$
#
# Stationary points occur when $\;y'(x) = 0\;$
#
# $6x^2 - 10x - 4 = 0$
#
# $(3x + 1)(2x - 4) = 0$
#
# $x = -1/3,\;2$
# + [markdown] slideshow={"slide_type": "subslide"}
# At $x = -1/3$
#
# $y''(-1/3) = 12 \times -1/3 - 10 = -14 < 0$
#
# So this point is a **maximum**.
#
# At $x = 2$
#
# $y''(2) = 12 \times 2 - 10 = 14 > 0$
#
# So this point is a **mimimum**.
#
# Inflection points occur whenever $y''(x) = 0$
#
# $y''(x) = 12x - 10 = 0$
#
# $x = \frac{10}{12} = \frac{5}{6}$
#
# This is an **inflection point**
# + slideshow={"slide_type": "subslide"}
x = np.linspace(-2, 3.5, 100)
y = 2*x**3 - 5*x**2 - 4*x
plt.plot(x,y, label='y = 2x^3 - 5x^2 - 4x')
plt.plot([2., -1./3., 5./6.], [-12., 19./27., -305./54.], 'ko')
plt.xlabel('x')
plt.ylabel('y')
# + [markdown] slideshow={"slide_type": "subslide"}
# **Note**: Points of inflection do not require that $\;y'(x) = 0\;$, only that $\;y''(x) = 0\;$.
#
# Points of inflection are important in biology as they define conditions where a response (e.g. reaction rate) is most or least sensitive to a change in conditions (e.g. the concentration of a metabolite).
# + [markdown] slideshow={"slide_type": "slide"}
# ## Reminder on curve sketching
#
#
# - Aim to evaluate and identify key values of the function (i.e. turning points, points of inflection)
#
#
# - Look at the limit behaviour as $\;x \to \pm \infty\;$ and as $\;x\;$ approaches any points where the function is undefined (e.g. $\;x \to 0\;$ for $\;y = 1/x\;$).
#
#
# - Determine the first and second order derivatives to find turning points and points of inflection.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Real life example
# The number $n$ (in thousands) of bacteria on an agar plate at time $t$ (in days) is given by the expression:
#
# $n = 15.42 + 6t - t^2$
#
# 1. Find the time at which the greatest number of bacteria are present on the plate.
# 1. Find the number of bacteria on the plate at this time.
#
# To do this we must find the turning points of the function.
# + [markdown] slideshow={"slide_type": "subslide"}
# ##### 1. Find the time at which the greatest number of bacteria are present on the plate
#
# - $n(t) = 15.42 + 6t - t^2$
# - $n'(t) = 6 - 2t$
# - $n'(t) = 0 \quad\implies\quad6-2t=0\quad\implies t=3$
#
# To show this is a maximum, we need to check $n''(t)$
#
# $n''(t) = -2$
#
# Therefore, $n''(t)<0$, for $t = 3$. This means that a maximum occurs at $t = 3$ days.
# + [markdown] slideshow={"slide_type": "fragment"}
# ##### 2. Find the number of bacteria on the plate at this time
#
# $n(3) = 15.42 + 6 \times 3 - 3^2 = 24.42$
#
# The greatest number of bacteria on the plate is **24,420**.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Real life example 2
# The growth rate $R$ of a cell colony with $N$ cells at time $t$ can be represented by the equation
#
# $R = \frac{d N}{d t} = kN - bN^2$
#
# For this example take the constants $k$ and $b$ as $k = 3.8$/hr, and $b = 0.01$/hr. This is called a **logistic** model.
#
# (a) What is the equilibrium size of the population?
#
# (b) What population size leads to the largest growth rate?
# + [markdown] slideshow={"slide_type": "subslide"}
# (a) The equilibrium will occur when the population stops changing, i.e. when $R = 0$. Meaning:
#
# $R = 3.8 N - 0.01 N^2 = 0$
#
# $N (3.8 - 0.01 N) = 0$
#
# We can disregard the $N = 0$ solution, as it represents population extinction. This means that
#
# $N = \frac{3.8}{0.01} = 380$.
# + [markdown] slideshow={"slide_type": "subslide"}
# (b) To find the largest growth rate, we want the maximal value of $R(N)$. This means we need to find $R'(N) = 0$.
#
# $R(N) = 3.8 N - 0.01 N^2$
#
# $R'(N) = 3.8 - 0.02 N$
#
# If $R'(N) = 0$
#
# $3.8 - 0.02N = 0$
#
# $N = 190$
#
# Since $R''(N) = -0.02 < 0$, we can be sure that this is a maximum.
| lectures/lecture-03-differentiation-01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Planning observations with `astroplan`
# + jupyter={"outputs_hidden": false}
import matplotlib.pyplot as plt
import numpy as np
import astropy.units as u
from astropy.time import Time
import pytz
from astroplan.plots import plot_airmass
from astroplan import Observer, FixedTarget, time_grid_from_range
# -
# ---
#
# <img style="float: right;" src="./images/UTC2.jpg" width="300"/>
#
#
# # Time and Dates
# - ### All dates and times in are UTC: *Coordinated Universal Time*
# - All `Time` calculation assume that the time is UTC.
# - UTC is related to Greenwich Mean Time (GMT) but does not change with a change of seasons.
# - Time will default to 00:00:00 UTC.
# + jupyter={"outputs_hidden": false}
my_date = Time("2021-02-22 22:15")
my_date
# + jupyter={"outputs_hidden": false}
my_date.iso
# + jupyter={"outputs_hidden": false}
f"The value of my_date is {my_date.iso}"
# -
# ### Different Date Formats
# + jupyter={"outputs_hidden": false}
print(f"The Julian Date is {my_date.jd:.2f}")
print(f"The Modified Julian Date is {my_date.mjd:.2f}")
print(f"The unix Epoch is {my_date.unix:.2f}") # Seconds since (Jan 01, 1970 00:00:00 UTC)
print(f"The fraction of a year is {my_date.decimalyear:.2f}")
# -
# ---
#
# ### [Accurate Time](https://www.ucolick.org/~sla/leapsecs/amsci.html) - `UT1`
#
# * `AstroPy` calculates the times of events to a very high accuracy.
# * Earth's rotation period is constantly changing due to tidal forces and changes in the Earth's moment of inertia.
# * `AstroPy` uses a time convention called `UT1`.
# * `UT1` is constanly changing with repect to `UTC`.
# * This system is tied to the rotation of the Earth with repect to the positions of distant quasars (International Celestial Reference Frame).
#
# [`UT1` is not really a time, it’s a way to express the Earth’s rotation angle and it should not be thought of as a real time scale](http://mperdikeas.github.io/utc-vs-ut1-time.html)
#
# The orientation of the Earth, which must be measured continuously to keep `UT1` accurate. This measurement is logged by the International Earth Rotation and Reference Systems Service (IERS). They publish a "bulletin" with the most recent measurements of the Earth's orientation. This bulletin is constantly being updated.
#
# You will run into occasions when you will get a warning that your dates are out of range of the IERS bulletin. To update the bulletin, run the follow block of code (this can take a while to run - so be patient):
#
# ---
# + jupyter={"outputs_hidden": false}
from astroplan import download_IERS_A
download_IERS_A()
# -
# ---
#
# <img style="float: right;" src="./images/LatLon.jpg" width="500"/>
#
# # Places
#
# ## Setting your location - `Observer`
#
# * `longitude` and `latitude` - any angular unit
# * `timezone` - see below
# * `name` - any string
# + jupyter={"outputs_hidden": false}
computer_room = Observer(longitude = -122.311473 * u.deg,
latitude = 47 * u.deg + 39 * u.arcmin + 15 * u.arcsec,
timezone = 'US/Pacific',
name = "UW Astro Computer Lab"
)
# + jupyter={"outputs_hidden": false}
computer_room
# + jupyter={"outputs_hidden": false}
computer_room.name
# -
# ## Note about negative values for latitude and longitude
#
# * If you use decimal degrees, this format is good: `longitude = -122.311473 * u.deg`
# * If you are using deg, min, sec, make sure to use `()`: `longitude = -(122 * u.deg + 18 * u.arcmin + 38.5 * u.arcsec)`
# * Note the `-` outside the `()`
# ---
# ## Working with timezones (local time)
#
# * [Timezone List](https://en.wikipedia.org/wiki/List_of_tz_database_time_zones)
# * Use the name in the **TZ database name** column.
# * Only use timezone conversions for printouts, NEVER calculations!
# * If you cannot find the exact timezone but know the offset from GMT use: `Etc/GMT(+/-)OFFSET` - (Example: `Etc/GMT-8`)
# * Working with tomezones is a [quick path to maddness!](https://xkcd.com/1883/)
# + jupyter={"outputs_hidden": false}
computer_room.timezone
# + jupyter={"outputs_hidden": false}
local_now = my_date.to_datetime(computer_room.timezone)
print(local_now)
# -
# ---
#
# <img style="float: right;" src="./images/SunNoon.jpg" width="300"/>
#
# ## Information at your location
#
# ### `noon` - output: UTC datetime of noon at your location
#
# * `LOCATION.noon(DATE, WHICH)`
# * `DATE` : Time of observation
# * `WHICH`: Choose which noon relative to the present `DATE` would you like to calculate:
# * next
# * previous
# * nearest
# + jupyter={"outputs_hidden": false}
my_noon = computer_room.noon(my_date, which = "previous")
# + jupyter={"outputs_hidden": false}
my_noon
# + jupyter={"outputs_hidden": false}
print(my_noon.to_datetime(computer_room.timezone))
# -
# ### `tonight` - output `[UTC datetime sunset, UTC datetime sunrise]`
#
# * `LOCATION.tonight(TIME, HORIZON)`
# * The `TIME` has to be before local sunset!
# * `HORIZON`: Sun position above/below horizon to use for calculating set/rise time
# + jupyter={"outputs_hidden": false}
tonight_here = computer_room.tonight(my_noon, horizon = 0 * u.deg)
# + jupyter={"outputs_hidden": false}
tonight_here
# + jupyter={"outputs_hidden": false}
print(f"Sunset will be at {tonight_here[0].iso} UTC")
print(f"Sunrise will be at {tonight_here[1].iso} UTC")
# + jupyter={"outputs_hidden": false}
local_sunset = tonight_here[0].to_datetime(computer_room.timezone)
local_sunrise = tonight_here[1].to_datetime(computer_room.timezone)
print(f"Sunset will be at {local_sunset} local time")
print(f"Sunrise will be at {local_sunrise} local time")
# -
# ### Astronomical twilight is when the Sun is 18 degrees below the horizon
#
# * This is often the time to begin/end observation
# + jupyter={"outputs_hidden": false}
observe_night = computer_room.tonight(my_noon, horizon = -18 * u.deg)
# + jupyter={"outputs_hidden": false}
print(f"Start observations tonight at {observe_night[0].iso} UTC")
print(f"End observations tonight at {observe_night[1].iso} UTC")
# + jupyter={"outputs_hidden": false}
observing_length = (observe_night[1] - observe_night[0]).to(u.h)
print(f"You can observe for {observing_length:.1f} tonight")
# + jupyter={"outputs_hidden": false}
# Local Times
print(f"Tonight's observing starts at {observe_night[0].to_datetime(computer_room.timezone)} local time")
print(f"Tonight's observing ends at {observe_night[1].to_datetime(computer_room.timezone)} local time")
# -
# ---
#
# <img style="float: right;" src="./images/NGC2403.jpg" width="250"/>
#
# # Things
#
# ## Objects in the sky - `FixedTarget`
#
# ### Most targets can be defined by name
# * ### Uses [SIMBAD](http://simbad.u-strasbg.fr/simbad/sim-fbasicwill)
# + jupyter={"outputs_hidden": false}
my_target_one = FixedTarget.from_name("ngc2403")
# + jupyter={"outputs_hidden": false}
my_target_one
# + jupyter={"outputs_hidden": false}
my_target_one.coord
# -
my_target_one.dec.degree
my_target_one.dec.dms
# + jupyter={"outputs_hidden": false}
my_target_one.ra.hour
# -
my_target_one.ra.hms
# ### Can you see the object at midnight tonight?
# + jupyter={"outputs_hidden": false}
my_midnight = computer_room.midnight(my_noon, which='next')
# + jupyter={"outputs_hidden": false}
computer_room.target_is_up(my_midnight, my_target_one)
# -
# ---
#
# <img style="float: right;" src="./images/AltAz.gif" width="400"/>
#
# ## Where in the sky?
#
# * Altitude (Alt) - Degrees above horizon
# * Azimuth (Az) - Compass degrees from North
# * `LOCATION.altaz(TIME, TARGET)`
#
# + jupyter={"outputs_hidden": false}
where_to_look = computer_room.altaz(my_midnight, my_target_one)
# + jupyter={"outputs_hidden": false}
where_to_look.alt
# + jupyter={"outputs_hidden": false}
where_to_look.az
# -
# ---
#
# <img style="float: right;" src="./images/Airmass.png" width="200"/>
#
# ### [Air Mass](https://en.wikipedia.org/wiki/Air_mass_%28astronomy%29) is the optical path length through Earth’s atmosphere.
#
# * At sea-level, the air mass at the zenith is 1.
# * Air mass increases as you move toward the horizon.
# * Air mass at the horizon is approximately 38.
# * The best time to observe a target is at minimum airmass.
# * When the airmass of your target is getting close to 2, you should be observing another target.
# + jupyter={"outputs_hidden": false}
# You can find the airmass by using the .secz method
where_to_look.secz
# -
# ##### Airmass < 2, you are good to go.
# ### Not all targets can (or should) be observed at all locations
# + jupyter={"outputs_hidden": false}
my_target_two = FixedTarget.from_name("Sirius")
# + jupyter={"outputs_hidden": false}
computer_room.target_is_up(my_midnight, my_target_two)
# + jupyter={"outputs_hidden": false}
where_to_look_two = computer_room.altaz(my_midnight, my_target_two)
# + jupyter={"outputs_hidden": false}
where_to_look_two.alt, where_to_look_two.az
# + jupyter={"outputs_hidden": false}
where_to_look_two.secz
# -
# ##### Airmass > 2, a big NOPE!
# ---
#
# ## Planning observation is better with plots
# #### Setup our observing window
# + jupyter={"outputs_hidden": false}
my_time_grid = time_grid_from_range(observe_night, time_resolution = 0.5 * u.h)
# + jupyter={"outputs_hidden": false}
my_time_grid
# -
# ### Simple Plot
# + jupyter={"outputs_hidden": false}
plot_airmass(my_target_one, computer_room, my_time_grid);
# -
# ### Better Plot
# ##### Make a target list
# + jupyter={"outputs_hidden": false}
target_list = [my_target_one, my_target_two]
# -
target_list
# + jupyter={"outputs_hidden": false}
fig,ax = plt.subplots(1,1)
fig.set_size_inches(12,6)
fig.tight_layout()
for my_object in target_list:
ax = plot_airmass(my_object, computer_room, my_time_grid)
ax.legend(loc=0,shadow=True);
# -
# ---
# ### [There is lots of information available for your location](https://astroplan.readthedocs.io/en/latest/api/astroplan.Observer.html)
#
# + jupyter={"outputs_hidden": false}
computer_room.is_night(my_noon)
# + jupyter={"outputs_hidden": false}
computer_room.sun_altaz(my_noon)
# + jupyter={"outputs_hidden": false}
computer_room.moon_illumination(my_midnight)
# + jupyter={"outputs_hidden": false}
computer_room.local_sidereal_time(my_midnight)
# + [markdown] jupyter={"outputs_hidden": false}
# <p align="center">
# <img src="./images/A11StarChart.jpg" width = "1000">
# </p>
# -
| Python_Observing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pyEX
c = pyEX.Client()
# c = pyEX.Client(version='sandbox')
# # Symbols data
c.symbolsDF().head()
# ## Symbols
c.symbols()[:5]
c.symbolsDF().head()
c.symbolsList()[:5]
# ## IEX Symbols
c.iexSymbols()[:5]
c.iexSymbolsDF().head()
c.iexSymbolsList()[:10]
# ## International Symbols
c.internationalSymbols()[:5]
c.internationalSymbolsDF().head()
c.internationalSymbolsList()[:5]
# ## FX Symbols
c.fxSymbols()
dfs = c.fxSymbolsDF()
dfs[0].head()
dfs[1].head()
l = c.fxSymbolsList()
l[0][:5]
l[1][:5]
# ## Options Symbols
d = c.optionsSymbols()
list(d.keys())[:5]
c.optionsSymbolsDF().head()
c.optionsSymbolsList()[:5]
# ## International Exchanges
c.internationalExchanges()[:5]
c.internationalExchangesDF().head()
# ## US Exchanges
c.exchanges()[:5]
c.exchangesDF().head()
# ## US Holidays
c.holidays()[:5]
c.holidaysDF().head()
# ## Sectors
c.sectors()[:5]
c.sectorsDF().head()
# ## Tags
c.tags()[:5]
c.tagsDF().head()
# # Data Points API
c.points()[:5]
c.pointsDF().head()
c.points(key='DGS10')
c.pointsDF('aapl').head()
c.points('aapl', 'QUOTE-LATESTPRICE')
# # Stock API
symbol = 'AAPL'
# ## Advanced Stats
c.advancedStats(symbol)
c.advancedStatsDF(symbol)
# ## Balance Sheet
c.balanceSheet(symbol)
c.balanceSheetDF(symbol)
# ## Book
c.book(symbol)
c.bookDF(symbol).head()
# ## Cashflow
c.cashFlow(symbol)
c.cashFlowDF(symbol)
# ## Collections
# +
# c.collections?
# +
# c.collectionsDF?
# -
# ## Company
c.company(symbol)
c.companyDF(symbol)
# ## Delayed Quote
c.delayedQuote(symbol)
c.delayedQuoteDF(symbol)
# ## Dividends
c.dividends(symbol)
c.dividendsDF(symbol)
# ## Earnings
c.earnings(symbol)
c.earningsDF(symbol)
# ## Earnings Today
c.earningsToday()
c.earningsTodayDF().head()
# ## Effective Spread
c.spread(symbol)
c.spreadDF(symbol)
# ## Estimates
c.estimates(symbol)
c.estimatesDF(symbol)
# ## Financials
c.financials(symbol)
c.financialsDF(symbol)
# ## Fund Ownership
c.fundOwnership(symbol)
c.fundOwnershipDF(symbol)
# ## Historical Prices
c.chart(symbol)
c.chartDF(symbol)
# ## Income Statement
c.incomeStatement(symbol)
c.incomeStatementDF(symbol)
# ## Insider Roster
c.insiderRoster(symbol)
c.insiderRosterDF(symbol)
# ## Insider Summary
c.insiderSummary(symbol)
c.insiderSummaryDF(symbol)
# ## Insider Transactions
c.insiderTransactions(symbol)[:5]
c.insiderTransactionsDF(symbol)
# ## Institutional Ownership
c.institutionalOwnership(symbol)
c.institutionalOwnershipDF(symbol)
# ## Intraday Prices
c.intraday(symbol)[:5]
c.intradayDF(symbol).head()
# ## IPO Calendar
c.ipoToday()
c.ipoTodayDF()
c.ipoUpcoming()
c.ipoUpcomingDF()
# ## Key Stats
c.keyStats(symbol)
c.keyStatsDF(symbol)
# ## Largest Trades
c.largestTrades(symbol)
c.largestTradesDF(symbol)
# ## List
c.list()[:5]
c.listDF().head()
# ## Logo
c.logo(symbol)
c.logoPNG(symbol) # Not available for sandbox
c.logoNotebook(symbol) # Not available for sandbox
# ## Market Volume
c.marketVolume()[:5]
c.marketVolumeDF()
# ## News
c.news(symbol)[:5]
c.newsDF(symbol).head()
c.marketNews()[:5]
c.marketNewsDF().head()
# ## OHLC
c.ohlc(symbol)
c.ohlcDF(symbol)
# ## Open/Close Price
c.ohlc(symbol)
c.ohlcDF(symbol)
# ## Options
exps = c.optionExpirations(symbol)
exps
c.options(symbol, exps[0])[:5]
c.optionsDF(symbol, exps[0]).head()
# ## Peers
c.peers(symbol)
c.peersDF(symbol)
# ## Previous Day Price
c.yesterday(symbol)
c.yesterdayDF(symbol)
c.marketYesterday()['A']
c.marketYesterdayDF().head()
# ## Price
c.price(symbol)
c.priceDF(symbol)
# ## Price Target
c.priceTarget(symbol)
c.priceTargetDF(symbol)
# ## Quote
c.quote(symbol)
c.quoteDF(symbol)
# # Recommendation Trends
# # Sector Performance
c.sectorPerformance()
c.sectorPerformanceDF()
# ## Splits
c.splits(symbol)
c.splitsDF(symbol)
# ## Upcoming Events
# ## Volume By Venue
c.volumeByVenue(symbol)
c.volumeByVenueDF(symbol)
# # Alternative
# ## crypto
# +
# c.crypto()
# +
# c.cryptoDF()
# -
# ## sentiment
c.sentiment(symbol)
c.sentimentDF(symbol)
# ## CEO Compensation
c.ceoCompensation(symbol)
c.ceoCompensationDF(symbol)
# # Forex
# ## Exchange Rates
# # IEX Data
# ## TOPS
# ## Last
# ## DEEP
# ## DEEP Auction
# ## DEEP Book
# ## DEEP Operational Halt Status
# ## DEEP Official Price
# ## DEEP Security Event
# ## DEEP Short Sale Price Test Status
# ## DEEP System Event
# ## DEEP Trades
# ## DEEP Trade Break
# ## DEEP Trading Status
# ## Listed Regulation SHO Threshold Securities List In Dev
# ## Listed Short Interest List In Dev
# ## Stats Historical Daily In Dev
# ## Stats Historical Summary
# ## Stats Intraday
# ## Stats Recent
# ## Stats Records
# # API Metadata
| examples/all.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import sys
import importlib
import copy
from collections import defaultdict
sys.path.insert(0, '/n/groups/htem/Segmentation/shared-nondev/cb2_segmentation/analysis_mf_grc')
from tools_pattern import get_eucledean_dist
# script_n = os.path.basename(__file__).split('.')[0]
script_n = 'distribution_123share_vs_random_210407'
import my_plot
importlib.reload(my_plot)
from my_plot import MyPlotData, my_box_plot
def to_ng_coord(coord):
return (
int(coord[0]/4),
int(coord[1]/4),
int(coord[2]/40),
)
import compress_pickle
# input_graph = compress_pickle.load('/n/groups/htem/Segmentation/shared-nondev/cb2_segmentation/analysis_mf_grc/'\
# 'mf_grc_model/input_graph_201114_restricted_z.gz')
input_graph = compress_pickle.load('/n/groups/htem/Segmentation/shared-nondev/cb2_segmentation/analysis_mf_grc/'\
# 'mf_grc_model/input_graph_210407_xlim_90_140_zlim_17_27.gz')
'mf_grc_model/input_graph_210407_xlim_90_140_zlim_2.5_41.5.gz')
# input_graph = compress_pickle.load('/n/groups/htem/Segmentation/shared-nondev/cb2_segmentation/analysis_mf_grc/mf_grc_model/input_graph_201114.gz')
# grcs = [k for k in input_graph.grcs.keys()]
z_min = 10
z_max = 40
# z_min = 20000
# z_max = 30000
x_min = 360
x_max = 520
# radius = 200
n_randoms = 5
def get_prob(in_graph, unique_count=False):
n_common_pairs = 0
processed = set()
total_n_pairs = 0
hist = defaultdict(int)
n = 0
for grc_i_id in in_graph.grcs:
n += 1
if n % 1000 == 0:
print(n, end=', ')
grc_i = in_graph.grcs[grc_i_id]
rosettes_i = set([mf[0] for mf in grc_i.edges])
for grc_j_id in in_graph.grcs:
if grc_i_id == grc_j_id:
continue
if unique_count and (grc_i_id, grc_j_id) in processed:
continue
processed.add((grc_i_id, grc_j_id))
processed.add((grc_j_id, grc_i_id))
grc_j = in_graph.grcs[grc_j_id]
common_rosettes = set([mf[0] for mf in grc_j.edges])
common_rosettes = common_rosettes & rosettes_i
hist[len(common_rosettes)] += 1
for k in hist:
# fix 0 datapoint plots
if hist[k] == 0:
hist[k] = 1
return hist
input_observed = copy.deepcopy(input_graph)
hist_data = get_prob(input_observed)
n_grcs = len(input_graph.grcs)
# +
def check_share(fname, depth):
graph = compress_pickle.load(fname)
graph.remove_empty_mfs()
hist_data = get_prob(graph)
n_grcs = len(graph.grcs)
n_mfs = len(graph.mfs)
# print(hist_data)
print(fname)
print(f'n_grcs: {n_grcs} ({n_grcs/depth}), n_mfs: {n_mfs}({n_mfs/depth})')
print(f'1: {hist_data[1]/n_grcs}, 2: {hist_data[2]/n_grcs}')
print()
fname = ('/n/groups/htem/Segmentation/shared-nondev/cb2_segmentation/analysis_mf_grc/'\
'mf_grc_model/input_graph_210407_xlim_90_140_zlim_17_27.gz')
check_share(fname, depth=10)
fname = ('/n/groups/htem/Segmentation/shared-nondev/cb2_segmentation/analysis_mf_grc/'\
'mf_grc_model/input_graph_210407_xlim_90_140_zlim_14.5_29.5.gz')
check_share(fname, depth=15)
fname = ('/n/groups/htem/Segmentation/shared-nondev/cb2_segmentation/analysis_mf_grc/'\
'mf_grc_model/input_graph_210407_xlim_90_140_zlim_12.0_32.0.gz')
check_share(fname, depth=20)
fname = ('/n/groups/htem/Segmentation/shared-nondev/cb2_segmentation/analysis_mf_grc/'\
'mf_grc_model/input_graph_210407_xlim_90_140_zlim_9.5_34.5.gz')
check_share(fname, depth=25)
fname = ('/n/groups/htem/Segmentation/shared-nondev/cb2_segmentation/analysis_mf_grc/'\
'mf_grc_model/input_graph_210407_xlim_90_140_zlim_7.0_37.0.gz')
check_share(fname, depth=30)
fname = ('/n/groups/htem/Segmentation/shared-nondev/cb2_segmentation/analysis_mf_grc/'\
'mf_grc_model/input_graph_210407_xlim_90_140_zlim_4.5_39.5.gz')
check_share(fname, depth=35)
fname = ('/n/groups/htem/Segmentation/shared-nondev/cb2_segmentation/analysis_mf_grc/'\
'mf_grc_model/input_graph_210407_xlim_90_140_zlim_2.5_41.5.gz')
check_share(fname, depth=39)
# +
# import tools_mf_graph
# importlib.reload(tools_mf_graph)
# input_graph = compress_pickle.load('/n/groups/htem/Segmentation/shared-nondev/cb2_segmentation/analysis_mf_grc/'\
# # 'mf_grc_model/input_graph_210407_xlim_90_140_zlim_17_27.gz')
# 'mf_grc_model/input_graph_210407_xlim_90_140_zlim_2.5_41.5.gz')
n_randoms = 5
hist_random_21 = []
for n in range(n_randoms):
# input_graph.randomize_graph(random_model=True)
input_graph.randomize_graph_by_grc(
single_connection_per_pair=True,
constant_grc_degree=True,
constant_dendrite_length=21000,
# always_pick_closest_rosette=True,
)
hist_random = get_prob(input_graph)
hist_random_21.append(hist_random)
# +
n_randoms = 5
hist_random_17 = []
for n in range(n_randoms):
# input_graph.randomize_graph(random_model=True)
input_graph.randomize_graph_by_grc(
single_connection_per_pair=True,
constant_grc_degree=True,
constant_dendrite_length=17000,
# always_pick_closest_rosette=True,
)
hist_random = get_prob(input_graph)
hist_random_17.append(hist_random)
# +
global_random_n_grcs, hist_global_random = compress_pickle.load('/n/groups/htem/Segmentation/shared-nondev/cb2_segmentation/analysis_mf_grc/mf_grc_analysis/share_distribution/gen_global_random_7k_204k_data_2000.gz')
import compress_pickle
compress_pickle.dump((
hist_data,
hist_random,
global_random_n_grcs,
hist_global_random,
), f"{script_n}_data.gz")
# normalize
# total_n_pairs = hist_data[0] + hist_data[1] + hist_data[2] + hist_data[3]
# global_random_n_grcs = 204000
# +
mpd_data = MyPlotData()
for n_share in [1, 2, 3]:
if n_share in hist_data:
mpd_data.add_data_point(
n_share=n_share,
count=hist_data[n_share]/n_grcs,
type='Observation',
)
# for hist_random in hist_randoms:
# if n_share in hist_random:
# mpd_data.add_data_point(
# n_share=n_share,
# count=hist_random[n_share]/n_grcs,
# type='Local Random',
# )
# for hist_random in hist_random2s:
# if n_share in hist_random:
# mpd_data.add_data_point(
# n_share=n_share,
# count=hist_random[n_share]/n_grcs,
# type='Local Random',
# )
# for hist_ex_30 in hist_ex_30s:
# if n_share in hist_ex_30:
# mpd_data.add_data_point(
# n_share=n_share,
# count=hist_ex_30[n_share]/n_grcs,
# type='LocalEx Random',
# )
if n_share in hist_global_random:
mpd_data.add_data_point(
n_share=n_share,
# need to divide by 2 because we're sampling only 2/200 grcs
# (or multiply others by 2)
count=hist_global_random[n_share]/global_random_n_grcs/2,
type='Global Random',
)
def custom_legend_fn(plt):
# plt.legend(bbox_to_anchor=(1.025, .8), loc='upper left', borderaxespad=0.)
plt.legend(loc='upper right', frameon=False, markerscale=2, prop={'size': 8})
# importlib.reload(my_plot); my_plot.my_relplot(
# mpd_data,
# x='n_share',
# y='count',
# hue='type',
# # hue_order=['Data', 'Random Model'],
# kind='scatter',
# context='paper',
# # ylim=[.005, 50],
# ylim=[.005, 2000],
# xlim=[.7, 3.3],
# log_scale_y=True,
# s=150,
# # xticklabels=['', 1, '', 2, '', 3, ''],
# xticks=[1, 2, 3],
# height=4,
# aspect=1.1,
# custom_legend_fn=custom_legend_fn,
# y_axis_label='GrC pairs / # GrCs',
# x_axis_label='Shared Inputs',
# # save_filename=f'{script_n}.pdf',
# save_filename=f'{script_n}.svg',
# show=True,
# )
importlib.reload(my_plot); my_plot.my_relplot(
mpd_data,
x='n_share',
y='count',
hue='type',
# hue_order=['Data', 'Random Model'],
kind='line',
err_style="bars",
ci=68,
markers=True,
dashes=False,
# s=150,
context='paper',
# ylim=[.005, 50],
ylim=[.005, 2000],
xlim=[.9, 3.1],
log_scale_y=True,
# s=150,
# xticklabels=['', 1, '', 2, '', 3, ''],
xticks=[1, 2, 3],
# height=4,
width=3,
aspect=1.4,
font_scale=1,
custom_legend_fn=custom_legend_fn,
y_axis_label='GrC pairs / # GrCs',
x_axis_label='Shared MF inputs',
# save_filename=f'{script_n}.pdf',
save_filename=f'{script_n}_line.svg',
show=True,
)
# +
n_grcs = len(input_graph.grcs)
mpd_data = MyPlotData()
for n_share in [1, 2, 3]:
if n_share in hist_data:
mpd_data.add_data_point(
n_share=n_share,
count=hist_data[n_share]/n_grcs,
type='Observation',
)
for hist_random in hist_random_17:
if n_share in hist_random:
mpd_data.add_data_point(
n_share=n_share,
count=hist_random[n_share]/n_grcs,
type='Local Random',
)
def custom_legend_fn(plt):
# plt.legend(bbox_to_anchor=(1.025, .8), loc='upper left', borderaxespad=0.)
plt.legend(loc='upper right', frameon=False, markerscale=2, prop={'size': 8})
importlib.reload(my_plot); my_plot.my_relplot(
mpd_data,
x='n_share',
y='count',
hue='type',
# hue_order=['Data', 'Random Model'],
kind='line',
err_style="bars",
# ci=68,
markers=True,
dashes=False,
# s=150,
context='paper',
# ylim=[.005, 50],
ylim=[None, 70],
xlim=[.7, 3.3],
linewidth=2.5,
# log_scale_y=True,
# s=150,
# xticklabels=['', 1, '', 2, '', 3, ''],
xticks=[1, 2, 3],
width=3.5,
height=3,
# aspect=1.1,
custom_legend_fn=custom_legend_fn,
# y_axis_label='GrC pairs / # GrCs',
y_axis_label='Average pairs per GrC',
x_axis_label='Shared MF inputs',
# save_filename=f'{script_n}.pdf',
save_filename=f'{script_n}_line_linear.svg',
show=True,
)
# -
print(hist_data)
print(hist_random_17[0])
print(hist_data[2]/n_grcs)
print(hist_random_17[2][2]/n_grcs)
print(hist_random_21[2][2]/n_grcs)
print(hist_data[1]/n_grcs)
print(hist_random_17[2][1]/n_grcs)
print(hist_random_21[2][1]/n_grcs)
| analysis/mf_grc_analysis/share_distribution/distribution_123share_vs_random_210409_compare.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _execution_state="idle" _uuid="5280abe23d5ec188e9e671a1e292942f0487756e" _cell_guid="b6d17237-6780-4797-8ea5-6e71083c869d"
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load in
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import matplotlib.pyplot as plt
# Input data files are available in the "../input/" directory.
# For example, running this (by clicking run or pressing Shift+Enter) will list the files in the input directory
df = pd.read_csv("../input/timesData.csv")
from subprocess import check_output
print(check_output(["ls", "../input"]).decode("utf8"))
# Any results you write to the current directory are saved as output.
# + _uuid="c96488f9e744f09a04ec7a937782175b015c6eab" _execution_state="idle" _cell_guid="8132fdc4-a7a7-47c7-bb57-0b67352f9a83"
df.info()
# + _execution_state="idle" _uuid="aa2397c9bf284fd6ed6202be5d64b5ffa6eaa465" _cell_guid="b86dcc66-1622-469f-87fb-689f4d74aca4"
df.head()
# + _execution_state="idle" _uuid="d95281ea331f8dfbceac2da8ccd468865ae72bfa" _cell_guid="647db394-87d2-4234-8211-d0b5c211b1cf"
# Create variable with TRUE if year is equal to 2016
yr_2016 = df['year'] == 2016
# + _execution_state="idle" _uuid="553a8140949b5569db07ab130949425aeb65c403" _cell_guid="ac320010-173c-43b8-8037-4155c300c481"
#for Nigeria
# Create variable with TRUE if country is Nigeria
Nigeria = df['country'] == "Nigeria"
# Select all cases where the country is Nigeria and year is equal to 2016
Nigeria = df[Nigeria & yr_2016]
Nigeria
# + _execution_state="idle" _uuid="3a970c3a2b0a6dd6e2cdfd3460f3058480c2629c" _cell_guid="d5d098ef-9bec-44e0-8972-148dbf7bbf49"
#for Ghana
# Create variable with TRUE if country is Ghana
Ghana = df['country'] == "Ghana"
#Select all cases where the country is Ghana and year is equal to 2016
Ghana = df[Ghana & yr_2016]
Ghana
# + _execution_state="idle" _uuid="8f7a2057e905220ba856471b0009649792db7169" _cell_guid="14bd8859-43df-4f55-a25a-40b113201744"
#for Kenya
# Create variable with TRUE if country is Kenya
Kenya = df['country'] == "Kenya"
# Select all cases where the country is and year is equal to 2016
Kenya = df[Kenya & yr_2016]
Kenya
# + _execution_state="idle" _uuid="f7b3a8afe336418dbd3c56a3bad6af5cafdf9452" _cell_guid="641d72c4-1d00-41e8-aea6-a60616290ea3"
#for South Africa
# Create variable with TRUE if country is South Africa
SA = df['country'] == "South Africa"
# Select all cases where the country is South Africa and year is equal to 2016
SA = df[SA & yr_2016]
SA
# + _execution_state="idle" _uuid="c315360f6009c47b254978ba5fb2fb5ae64fe21b" _cell_guid="994640cb-df06-4010-b827-6b0d01f60d8b"
#for Egypt
# Create variable with TRUE if country is Egypt
Egypt = df['country'] == "Egypt"
# Select all cases where the country is Egypt and year is equal to 2016
Egypt = df[Egypt & yr_2016]
Egypt
# + _execution_state="idle" _uuid="d8cee398a8e7614cf918f1e388da48a592f32d3f" _cell_guid="8012410b-5d3b-4723-a2a4-a5d5aa4f02f7"
#for Morocco
# Create variable with TRUE if country is Morocco
Morocco = df['country'] == "Morocco"
# Select all casess country is Morocco and year is equal to 2016
Morocco = df[Morocco & yr_2016]
Morocco
# + _execution_state="idle" _uuid="af272075ece8e20f1012acb9e3e313912b218c38" _cell_guid="1a7d1370-cc21-4eef-8cda-50c289e73080"
#for Uganda
# Create variable with TRUE if country is Uganda
Uganda = df['country'] == "Uganda"
# Select all cases country is Uganda and year is equal to 2016
Uganda = df[Uganda & yr_2016]
Uganda
# + _uuid="9060bda5a8dd6aed93ae061b64a5e8e7ecad8d10" _cell_guid="1ea11dd7-6cbd-4f99-b3ac-de8f472b44ac"
frames = [Nigeria, Ghana, Kenya, Uganda, SA, Morocco, Egypt]
# + _uuid="cd35f342deffc0afbedb43dd33cdb6bbce706a77" _cell_guid="70d35130-6cc1-4c33-8ce6-2fffcc2fc980"
result = pd.concat(frames)
# + _uuid="75c216509b0c21cf2352cffae742fd1f0648115a" _cell_guid="2b9c8a23-0972-49a7-9d86-6728fe91ef45"
result
# + _uuid="953d9e06bdb8d2a57976b61da465b403ec423bbb" _cell_guid="1970402d-f449-4df2-a65e-dc97e6a28360"
#columns for visualizations
uni_name = result['university_name']
teaching = result['teaching']
research = result['research']
international_coll = result['international']
citations = result['citations']
stud_pop = result['num_students']
# + _uuid="93e6b9b16f7f8d1927a848ce72d7d99f97f711a8" _cell_guid="72eabd22-2e23-4ac1-acc2-366367ee8255"
stud_num = [x.replace(",", "") for x in stud_pop] #taking out the commas in student population data
# + _uuid="8c4e2763aafc7d79f8dbdf28a028bae8d5da0f9e" _cell_guid="976df661-f342-46a9-b38e-e882307abcc6"
#plotting universities vs student population
universities = uni_name.values.tolist() #converting the data frame to a list
objects = universities
y_pos = np.arange(len(objects))
# + _uuid="f5d928342cd6ea162048d1c167e631e6413d580d" _cell_guid="ddf02d8f-115d-49d8-b243-aa080e18f649"
#first plot-Student Population of Selected African Universities
plt.bar(y_pos, stud_num, align='center', alpha=0.5)
plt.xticks(y_pos, objects, rotation=90)
plt.ylabel('Student Population')
plt.title('Plot 1 - 2016 Student Population of selected African Universities')
plt.show()
# + _uuid="38557b6ae2ca71f24790f6fd4e49953f239124e9" _cell_guid="1cf871a3-217d-4037-8240-6b4cf5619f7b"
universities
# + _uuid="02340874baa6741247a2de57c59140d5ceb9cf35" _cell_guid="9650066c-c99a-4d4e-b98c-629290e0a123"
#Plot 2-Group bar charts for Teaching, Research, International Collaborations and citations
# Setting the positions and width for the bars
pos = np.arange(len(universities))
width = 0.2
# + _uuid="0f3b066e93a258cc51a5bccff5918a83e28a9e80" _cell_guid="ec0412e8-929b-4f9f-891b-3f364d441a30"
# Plotting the bars
fig, ax = plt.subplots(figsize=(10,5))
y1 = teaching
y2 = research
y3 = international_coll
y4 = citations
# Create a bar with Teaching data,
# in position pos,
plt.bar(pos,y1, width, alpha=0.5, color='b', label='Teaching')
# Create a bar with research data,
# in position pos + some width buffer,
plt.bar([p + width for p in pos], y2, width, alpha=0.5, color='r', label='Research')
# Create a bar with international collaborations data,
# in position pos + some width buffer,
plt.bar([p + width*2 for p in pos], y3, width, alpha=0.5, color='g', label='International Collaborations')
# Create a bar with citations data,
# in position pos + some width buffer,
plt.bar([p + width*3 for p in pos], y4, width, alpha=0.5, color='y', label='Citations')
# Set the y axis label
ax.set_ylabel('Ranking Parameters')
# Set the chart's title
ax.set_title('Bar chart for different ranking parameters')
# Set the position of the x ticks
ax.set_xticks([p + 1.5 * width for p in pos])
# Set the labels for the x ticks
ax.set_xticklabels(universities, rotation=90)
# Setting the x-axis and y-axis limits
plt.xlim(min(pos)-width, max(pos)+width*4)
plt.ylim([0, 100] )
# Adding the legend and showing the plot
plt.legend(['Teaching', 'Research', 'International Collaborations', 'Citations'], loc='upper left')
plt.grid()
plt.show()
# + _uuid="ec9728b7e21d6e15677d4d7765a794056288cd1e" _cell_guid="16aeeb3f-6089-474a-abfd-04a69d00090d"
# pairwise correlation between columns, dropping certain columns
result.drop(['world_rank', 'income', 'total_score', 'year'], axis=1).corr(method='spearman').style.format("{:.2}").background_gradient(cmap=plt.get_cmap('coolwarm'), axis=1)
# + _uuid="a89f15c277ce7740a5f700d78b10cfdc8aec77f7" _cell_guid="5c3f4e44-2514-40c4-b528-6d9ce061ceb1"
| downloaded_kernels/university_rankings/kernel_19.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] graffitiCellId="id_4zl18if"
# # Trie
# You've learned about Trees and Binary Search Trees. In this notebook, you'll learn about a new type of Tree called Trie. Before we dive into the details, let's talk about the kind of problem Trie can help with.
#
# Let's say you want to build software that provides spell check. This software will only say if the word is valid or not. It doesn't give suggested words. From the knowledge you've already learned, how would you build this?
#
# The simplest solution is to have a hashmap of all known words. It would take O(1) to see if a word exists, but the memory size would be O(n\*m), where n is the number of words and m is the length of the word. Let's see how a Trie can help decrease the memory usage while sacrificing a little on performance.
#
# ## Basic Trie
# Let's look at a basic Trie with the following words: "a", "add", and "hi"
# + graffitiCellId="id_gum3me0"
basic_trie = {
# a and add word
'a': {
'd': {
'd': {'word_end': True},
'word_end': False},
'word_end': True},
# hi word
'h': {
'i': {'word_end': True},
'word_end': False}}
print('Is "a" a word: {}'.format(basic_trie['a']['word_end']))
print('Is "ad" a word: {}'.format(basic_trie['a']['d']['word_end']))
print('Is "add" a word: {}'.format(basic_trie['a']['d']['d']['word_end']))
# + [markdown] graffitiCellId="id_8a5nwsy"
# You can lookup a word by checking if `word_end` is `True` after traversing all the characters in the word. Let's look at the word "hi". The first letter is "h", so you would call `basic_trie['h']`. The second letter is "i", so you would call `basic_trie['h']['i']`. Since there's no more letters left, you would see if this is a valid word by getting the value of `word_end`. Now you have `basic_trie['h']['i']['word_end']` with `True` or `False` if the word exists.
#
# In `basic_trie`, words "a" and "add" overlapp. This is where a Trie saves memory. Instead of having "a" and "add" in different cells, their characters treated like nodes in a tree. Let's see how we would check if a word exists in `basic_trie`.
# + graffitiCellId="id_r22pqse"
def is_word(word):
"""
Look for the word in `basic_trie`
"""
current_node = basic_trie
for char in word:
if char not in current_node:
return False
current_node = current_node[char]
return current_node['word_end']
# Test words
test_words = ['ap', 'add']
for word in test_words:
if is_word(word):
print('"{}" is a word.'.format(word))
else:
print('"{}" is not a word.'.format(word))
# + [markdown] graffitiCellId="id_rvrc2k0"
# The `is_word` starts with the root node, `basic_trie`. It traverses each character (`char`) in the word (`word`). If a character doesn't exist while traversing, this means the word doesn't exist in the trie. Once all the characters are traversed, the function returns the value of `current_node['word_end']`.
#
# You might notice the function `is_word` is similar to a binary search tree traversal. Since Trie is a tree, it makes sense that we would use a type of tree traversal. Now that you've seen a basic example of a Trie, let's build something more familiar.
# ## Trie Using a Class
# Just like most tree data structures, let's use classes to build the Trie. Implement two functions for the `Trie` class below. Implement `add` to add a word to the Trie. Implement `exists` to return `True` if the word exist in the trie and `False` if the word doesn't exist in the trie.
#
# + graffitiCellId="id_ripuwyf"
class TrieNode(object):
def __init__(self):
self.is_word = False
self.children = {}
class Trie(object):
def __init__(self):
self.root = TrieNode()
def add(self, word):
"""
Add `word` to trie
"""
pass
def exists(self, word):
"""
Check if word exists in trie
"""
pass
# + [markdown] graffitiCellId="id_h7y0qpa"
# <span class="graffiti-highlight graffiti-id_h7y0qpa-id_pncadbt"><i></i><button>Show Solution</button></span>
# + graffitiCellId="id_9l7z1sf"
word_list = ['apple', 'bear', 'goo', 'good', 'goodbye', 'goods', 'goodwill', 'gooses' ,'zebra']
word_trie = Trie()
# Add words
for word in word_list:
word_trie.add(word)
# Test words
test_words = ['bear', 'goo', 'good', 'goos']
for word in test_words:
if word_trie.exists(word):
print('"{}" is a word.'.format(word))
else:
print('"{}" is not a word.'.format(word))
# + graffitiCellId="id_pncadbt"
class TrieNode(object):
def __init__(self):
self.is_word = False
self.children = {}
class Trie(object):
def __init__(self):
self.root = TrieNode()
def add(self, word):
"""
Add `word` to trie
"""
current_node = self.root
for char in word:
if char not in current_node.children:
current_node.children[char] = TrieNode()
current_node = current_node.children[char]
current_node.is_word = True
def exists(self, word):
"""
Check if word exists in trie
"""
current_node = self.root
for char in word:
if char not in current_node.children:
return False
current_node = current_node.children[char]
return current_node.is_word
# + [markdown] graffitiCellId="id_8irwsjx"
# ## Trie using Defaultdict (Optional)
# This is an optional section. Feel free to skip this and go to the next section of the classroom.
#
# A cleaner way to build a trie is with a Python default dictionary. The following `TrieNod` class is using `collections.defaultdict` instead of a normal dictionary.
# + graffitiCellId="id_9cezzui"
import collections
class TrieNode:
def __init__(self):
self.children = collections.defaultdict(TrieNode)
self.is_word = False
# + [markdown] graffitiCellId="id_m7tuw99"
# Implement the `add` and `exists` function below using the new `TrieNode` class.
# + graffitiCellId="id_8y03wp7"
class Trie(object):
def __init__(self):
self.root = TrieNode()
def add(self, word):
"""
Add `word` to trie
"""
pass
def exists(self, word):
"""
Check if word exists in trie
"""
pass
# + [markdown] graffitiCellId="id_158726u"
# <span class="graffiti-highlight graffiti-id_158726u-id_461jk1b"><i></i><button>Hide Solution</button></span>
# + graffitiCellId="id_461jk1b"
class Trie(object):
def __init__(self):
self.root = TrieNode()
def add(self, word):
"""
Add `word` to trie
"""
current_node = self.root
for char in word:
current_node = current_node.children[char]
current_node.is_word = True
def exists(self, word):
"""
Check if word exists in trie
"""
current_node = self.root
for char in word:
if char not in current_node.children:
return False
current_node = current_node.children[char]
return current_node.is_word
# + graffitiCellId="id_6uuui8u"
# Add words
valid_words = ['the', 'a', 'there', 'answer', 'any', 'by', 'bye', 'their']
word_trie = Trie()
for valid_word in valid_words:
word_trie.add(valid_word)
# Tests
assert word_trie.exists('the')
assert word_trie.exists('any')
assert not word_trie.exists('these')
assert not word_trie.exists('zzz')
print('All tests passed!')
# + [markdown] graffitiCellId="id_bcwz5dp"
# The Trie data structure is part of the family of Tree data structures. It shines when dealing with sequence data, whether it's characters, words, or network nodes. When working on a problem with sequence data, ask yourself if a Trie is right for the job.
| Course/Data structures and algorithms/3.Basic algorithm/1.Basic algorithms/6.trie_introduction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: MoST_venv
# language: python
# name: most_venv
# ---
# # MoST: high-level scheduling tools
import MoST.MoST_base as most
import MoST.qast_utils.loopReader as lr
import MoST.transforms.TilingSchedule as ts
import MoST.transforms.ReorderingSchedule as rs
# We start by generating a new kernel, here for SGEMM. This is actually the code the user writes:
sg = lr.__debug_new_sgemm()
print(sg)
# The first thing we do is to lock this to a specific problem size:
sg_const = sg.partial_eval(512, 512, 512)
print(sg_const)
# We can generate specific transformations, for instance, tiling:
tile_8_16_8 = ts.TilingSchedule({'i':8, 'j':16, 'k':8})
# can run for sg or sg_const
print(tile_8_16_8.apply(sg_const))
# ... or reordering...
reorder_kij = rs.ReorderingSchedule(['k', 'i', 'j'])
print(reorder_kij.apply(sg_const))
# These transforms can be combined to generate more complicated, higher-level transforms, e.g.:
cs = most.CompoundSchedule([reorder_kij, tile_8_16_8])
print(cs.apply(sg_const))
# One could, also do multilevel tiling (CoSA style) as follows:
t1 = ts.TilingSchedule({'i':128, 'j':128, 'k':128})
t2 = ts.TilingSchedule({'i_in':8, 'j_in':16, 'k_in':8})
multilevel = most.CompoundSchedule([t1,t2])
print(multilevel.apply(sg_const))
# These scheduling elements can be defined manually as above, or through static algorithms, such as this HBL thing here...
memsize = 768
bounds = lr.getFixedLoopBounds(sg_const)
accesses = lr.getProjectiveDataAccesses(sg_const)
opt_tile = ts.TilingSchedule.generateHBLProjectiveTile(bounds, accesses, memsize, False)
print(opt_tile.apply(sg_const))
# ... or autotuning (in progress!)
| docs/matmap_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: openvino_env
# language: python
# name: openvino_env
# ---
# # Hello Segmentation
#
# A very basic introduction to OpenVINO segmentation model.
#
# We use the [road-segmentation-adas-0001](https://docs.openvinotoolkit.org/latest/omz_models_model_road_segmentation_adas_0001.html) model from [Open Model Zoo](https://github.com/openvinotoolkit/open_model_zoo/). ADAS stands for Advanced Driver Assistance Services. The model recognizes four classes: background, road, curb and mark.
#
# ## Imports
# +
import cv2
import matplotlib.pyplot as plt
import numpy as np
import sys
from openvino.inference_engine import IECore
sys.path.append("../utils")
from notebook_utils import segmentation_map_to_image
# -
# ## Load the network
# +
ie = IECore()
net = ie.read_network(
model="model/road-segmentation-adas-0001.xml")
exec_net = ie.load_network(net, "CPU")
output_layer_ir = next(iter(exec_net.outputs))
input_layer_ir = next(iter(exec_net.input_info))
# -
# ## Load an Image
# +
# The segmentation network expects images in BGR format
image = cv2.imread("data/empty_road_mapillary.jpg")
rgb_image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
image_h, image_w, _ = image.shape
# N,C,H,W = batch size, number of channels, height, width
N, C, H, W = net.input_info[input_layer_ir].tensor_desc.dims
# OpenCV resize expects the destination size as (width, height)
resized_image = cv2.resize(image, (W, H))
# reshape to network input shape
input_image = np.expand_dims(
resized_image.transpose(2, 0, 1), 0
)
plt.imshow(rgb_image)
# -
# ## Do Inference
# +
# Run the infernece
result = exec_net.infer(inputs={input_layer_ir: input_image})
result_ir = result[output_layer_ir]
# Prepare data for visualization
segmentation_mask = np.argmax(result_ir, axis=1)
plt.imshow(segmentation_mask[0])
# -
# ## Prepare data for visualization
# +
# Define colormap, each color represents a class
colormap = np.array([[68, 1, 84], [48, 103, 141], [53, 183, 120], [199, 216, 52]])
# Define the transparency of the segmentation mask on the photo
alpha = 0.3
# Use function from notebook_utils.py to transform mask to an RGB image
mask = segmentation_map_to_image(segmentation_mask, colormap)
resized_mask = cv2.resize(mask, (image_w, image_h))
# Create image with mask put on
image_with_mask = cv2.addWeighted(resized_mask, alpha, rgb_image, 1 - alpha, 0)
# -
# ## Visualize data
# +
# Define titles with images
data = {"Base Photo": rgb_image, "Segmentation": mask, "Masked Photo": image_with_mask}
# Create subplot to visualize images
f, axs = plt.subplots(1, len(data.items()), figsize=(15, 10))
# Fill subplot
for ax, (name, image) in zip(axs, data.items()):
ax.axis('off')
ax.set_title(name)
ax.imshow(image)
# Display image
plt.show(f)
| notebooks/003-hello-segmentation/003-hello-segmentation.ipynb |