code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
import mbuild as mb
from mbuild.lib.recipes.polymer import Polymer
# -
# # The basic process for creating a polymer in mBuild
#
# The `Polymer` class exists in mBuild's library of recipes `mbuild/lib/recipes/polymer.py`
#
# In order to create polymers, mbuild Compounds of the monomer(s) and any end groups must first be created, and the necesasry mbuild Ports added to the correct atoms with the desired orientations. The compounds in mBuild's library contain structures that are essentially ready to go for building polymers. However, the ultimate goal of mBuild's polymer tool is to make it easier to build polymers from nearly any structures that are given to it whether in the form of a file or SMILES string.
#
# There are multiple ways to input the needed information to build a polymer. When creating a polymer instance, monomer and end group compounds can be directly passed in. This approach can be used when the compounds already contain the needed ports, there will be examples below.
#
# The other, much more flexible, option is to use the `add_monomer` and `add_end_groups` methods on a `Polymer` instnace.
#
# # Building a polymer from SMILES
#
# In the example below, we will build up polyethylene, and add carboxylic acid end groups.
# There are two primary bits of input required:
#
# 1. A SMILES string of the monomer structure (and end group structure)
# 2. The indices of the hydrogen atoms that will be replaced to make room for the monomer-monomer bonding
#
# Step 2 will require a bit of trial and error
# +
# Quick example of the API and workflow
comp = mb.load('CC', smiles=True) # mBuild compound of the monomer unit
chain = Polymer()
chain.add_monomer(compound=comp,
indices=[2, -2],
separation=.15,
replace=True)
chain.add_end_groups(mb.load('C(=O)O', # Capping off this polymer with Carboxylic acid groups
smiles=True,
name='acid'),
index=3,
separation=0.15)
chain.build(n=7, sequence='A') # After monomers and end groups are added, call the build function
chain.visualize(show_ports=True).show()
# +
# Doing the same thing, but this time without adding an end group
# Now, the chain is capped with hydrogens (default behavior)
comp = mb.load('CC', smiles=True) # mBuild compound of the monomer unit
chain = Polymer()
chain.add_monomer(compound=comp,
indices=[2, -2],
separation=.15,
replace=True)
chain.build(n=10, sequence='A')
chain.visualize(show_ports=True).show()
# -
# # Selecting the correct values for `indices`
#
# As mentioned earlier, you have to provide information about which atoms on the monomer and end group will form the bonds between repeating monomers, and between the end groups. In this case, you are specificy the index of the hydrogen atoms that will be removed.
#
# Above, we chose `indices = [2, -2]` for the monomer units. What would happen if we chose the "wrong" indices?
#
# The cell below shows the incorrect structures that can come about from choosing the wrong indices.
# +
# Doing the same thing, but this time without adding an end group
# Now, the chain is capped with hydrogens (default behavior)
comp = mb.load('CC', smiles=True) # mBuild compound of the monomer unit
chain = Polymer()
chain.add_monomer(compound=comp,
indices=[-1, -2],
separation=.15,
replace=True)
chain.build(n=10, sequence='A')
chain.visualize(show_ports=True).show()
# -
# # Flexibility with end groups
# - You can only add one end group, while the other is capped off with hydrogen.
# - You can add 2 different end groups, labeled by 'head' and 'tail'
# - Or you can leave the ends open with an available port
# +
# Creating and adding 2 different end groups.
# The add_end_groups() function has a duplicate parameter (defaulted to True)
# When this is true, calling add_end_group will create 2 of the same compound
# and the polymer is capped off with each.
# If you want different end groups, change Duplicate to False and call the add_end_groups() function twice
comp = mb.load('CC', smiles=True) # mBuild compound of the monomer unit
chain = Polymer()
chain.add_monomer(compound=comp,
indices=[2, -2],
separation=.15,
replace=True)
chain.add_end_groups(mb.load('C(=O)O',smiles=True), # Capping off this polymer with Carboxylic acid groups
index=3,
separation=0.15,
duplicate=False) # Change duplicate to false
chain.add_end_groups(mb.load('N', smiles=True),
index=-1, separation=0.13,
duplicate=False, label="tail") # label this one tail
chain.build(n=10, sequence='A')
chain.visualize(show_ports=True).show()
# +
# If you only add one end group, you can still have the other end capped off with hydrogen
comp = mb.load('CC', smiles=True) # mBuild compound of the monomer unit
chain = Polymer()
chain.add_monomer(compound=comp,
indices=[2, -2],
separation=.15)
chain.add_end_groups(mb.load('C(=O)O',smiles=True), # Capping off this polymer with Carboxylic acid groups
index=3,
separation=0.15,
duplicate=False) # Change duplicate to false
chain.build(n=10, sequence='A')
chain.visualize(show_ports=True).show()
# +
#Or to leave the ends of the polymer open with ports, change the add_hydrogens parameter in build() to False
# This would still work when adding only one end group, then leaving the other end open.
comp = mb.load('CC', smiles=True) # mBuild compound of the monomer unit
chain = Polymer()
chain.add_monomer(compound=comp,
indices=[2, -2],
separation=.15,
replace=True)
chain.build(n=10, sequence='A', add_hydrogens=False)
chain.visualize(show_ports=True).show()
# -
# # Here is an example building up a Nylon polymer
# +
nylon = mb.load("NCCCCCCNOC(=O)CCCCC(=O)O", smiles=True)
chain = Polymer()
chain.add_monomer(compound=nylon, indices=[-23, -1], separation=.15)
chain.build(n=4, sequence='A')
chain.visualize().show()
# -
# ## Here's an example with a more complicated monomer and a little more detail into what's going on
#
# SMILES strings for Poly-ether-ether-ketone (PEEK)
#
# One has para linkages, the other has meta
#
# Goal is to build up a polymer with alternating PARA-META monomers
# +
peek_para = mb.load("Oc1ccc(Oc2ccc(C(=O)c3ccccc3)cc2)cc1", smiles=True)
peek_meta = mb.load("Oc1cc(Oc2ccc(C(=O)c3ccccc3)cc2)ccc1", smiles=True)
peek_polymer = Polymer()
peek_polymer.add_monomer(compound=peek_para,
indices = [22, 29],
separation = 0.1376,
replace=True
)
peek_polymer.add_monomer(compound=peek_meta,
indices = [22, 28],
separation = 0.1376,
replace=True
)
peek_polymer.build(n=3, sequence="AB")
peek_polymer.visualize().show()
# -
# ## A look at what is actually happening with each of the class methods
#
# The `add_monomer` and `add_end_group` functions are handling the creation of ports.
#
# The key is in the `bond_indices` and `replace` parameters.
# `bond_indices` points to the hydrogen atoms that are occupying the polymer bonding site and
# `replace` says to remove those atoms, and replace them with a port
#
# When the port is created, it defaults to using the orientation that already existed between the hydrogen atom and the atom it was bonded to.
# +
peek_para = mb.load("Oc1ccc(Oc2ccc(C(=O)c3ccccc3)cc2)cc1",smiles=True)
print('Before passing the compound into add_monomer()')
peek_para.visualize(show_ports=True).show()
peek_polymer = Polymer()
peek_polymer.add_monomer(compound=peek_para,
indices = [22, 29],
separation = 0.1376,
replace=True
)
print('After passing the compound into add_monomer()')
peek_polymer.monomers[0].visualize(show_ports=True).show()
# +
# Same thing with the end group
ca = mb.load('C(=O)O', smiles=True)
print('Before passing the compound into add_end_groups()')
ca.visualize(show_ports=True).show()
peek_polymer.add_end_groups(ca,
index=3,
separation=0.13,
replace=True)
# ca[3] is the hydrogen bonded to the carbon atom
print('After passing the compound into add_end_groups()')
peek_polymer.end_groups[0].visualize(show_ports=True).show()
# -
# # Not using the add_monomer and add_end_group functions
# - So far, the examples call these functions everytime to create a polymer, but they are optional
# - These functions basically handle the creation of ports.
# - If you have a compound where the ports already exist, or you want to add them yourself you can pass them into the polymer class instance, and go straight to the build() function.
# - However, you can use both approaches for the same polymer. Passing in compounds when initializing a polymer, and then adding to self.monomers or self.end_groups with the add functions.
# +
# In this example, we're using a CH2 compound from mBuild's library. This compound already has ports created
# So, we can skip the add_monomer method and go straight to build.
from mbuild.lib.moieties.ch2 import CH2
chain = Polymer(monomers=[CH2()])
chain.build(n=10, add_hydrogens=True)
chain.visualize().show()
# +
# You can combine passing in compounds, and using the available
# add_end_group and add_monomer functions
chain = Polymer(monomers=[CH2()])
chain.add_monomer(mb.load('c1ccccc1', smiles=True), indices=[-1, -4])
chain.add_end_groups(mb.load('C(=O)O',smiles=True), # Capping off this polymer with Carboxylic acid groups
index=3,
separation=0.15,
duplicate=False)
chain.build(n=5,sequence='AB', add_hydrogens=True)
chain.visualize().show()
# -
# ## Some more use case examples
#
# - Creating regio-regular backbone
# - Creating a poly disperse system
#
p3ht = mb.load('c1cscc1', smiles=True)
p3ht_flip = mb.load('c1cscc1', smiles=True)
p3ht_flip.rotate(np.pi, [0,0,1])
chain = Polymer()
chain.add_monomer(
compound=p3ht,
indices=[6, 7],
separation=.15,
replace=True
)
chain.add_monomer(
compound=p3ht_flip,
indices=[7, 6],
separation=.15,
replace=True
)
chain.build(n=5, sequence='AB')
chain.visualize(show_ports=True).show()
import random
#Regiorandom
chain = Polymer()
chain.add_monomer(
compound=p3ht,
indices=[6, 7],
separation=.15,
replace=True
)
chain.add_monomer(
compound=p3ht_flip,
indices=[7, 6],
separation=.15,
replace=True
)
seq = "".join([random.choice("AB") for i in range(10)])
chain.build(n=1, sequence=seq)
print(seq)
chain.visualize(show_ports=True).show()
# #### Initializing a polydisperse system
# +
polymer_lengths = [5, 10, 15, 20, 25, 30]
num_polymers = [3, 5, 6, 5, 3, 2]
polymers = []
ethane = mb.load('CC', smiles=True)
for length in polymer_lengths:
chain = Polymer()
chain.add_monomer(
compound=ethane,
indices=[2, -2],
separation=.15,
replace=True
)
chain.add_end_groups(
mb.load('C(=O)O',
smiles=True,
name='acid'),
index=3,
separation=0.15
)
chain.build(n=length)
polymers.append(chain)
system = mb.fill_box(polymers, num_polymers, density=50)
system.visualize().show()
# -
width = 1 # the width of the bars
fig, ax = plt.subplots()
rects1 = ax.bar(polymer_lengths, num_polymers, width)
ax.set_xlabel("polymer length (# monomers)")
ax.set_ylabel("# polymers")
ax.set_title("Polydispersity profile")
plt.show()
# # Reproducing the alkane chain recipe
#
# There currently exists a recipe that produces a simple alkane chain. Below I'll use the new `polymer.py` functionality to re-create the same alkane chain.
# +
ch2 = CH2()
chain = Polymer(monomers=[ch2])
chain.build(n=7, sequence='A')
chain.visualize(show_ports=True).show()
# +
# It's super easy to throw different end groups on there
chain = Polymer(monomers=[ch2])
chain.add_end_groups(mb.load('c1ccccc1', smiles=True),
index=-1,
separation=0.15,
replace=True)
chain.build(n=8, sequence='A')
chain.visualize().show()
| polymers/polymers-example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Gitignore files
#
# The process of doing homeworks or working on projects will inevitably create files that you might not want to sync to the remote repo on GitHub.com. Because the remote repo is "public" facing, the goal is to have a clean remote repo without miscellaneous files.
#
# Common versions of files you might not want on the remote repo:
# - `.DS_Store` (a temporary file generated by macOS)
# - `ipynb` checkpoint files
# - temporary data files (such as mid point saves, data files you examined to check)
#
# So how do we prevent git from syncing these files? The `.gitignore` file!
#
# ## How to create and edit a gitignore file
#
# There are several ways to create one:
#
# 1. When you create a repo, select to add a gitignore file. I usually pick the python template option.
# 2. You can add it later on github or on your computer. The file is a simple text file and should be named `.gitignore`
# 3. In GitHub desktop, choose the "Repository" menu, then "Repository settings", then "ignored files" and type in what you'll ignore.
#
# Editting it: Any text editor!
#
# ## Syntax
#
# Each line lists a file(s) or folder(s) that shouldn't be tracked. Wildcards are allowed. For example,
# ```
# .ipynb_checkpoints
# */.ipynb_checkpoints/*
# ```
# should prevent the checkpoints Jupyter Lab makes from being synced.
#
# ## I have a file in gitignore, but it's still on the remote repo!
#
# That's because it was already on the remote repo when you added it to the gitignore; **The gitignore file simply tells git to "not sync" those files any more. It doesn't mean "delete these files from the remote repo".**
#
# There are a few ways to delete the extra files in the remote repo. The simplest: Go to the repo on Github.com and delete them individually.
#
# If you have lots of files you want to suddenly gitignore/delete and you are comfortable running some git commands, you can do it programmatically. Open terminal/powershell, **move to the current directory to the root folder of your repo,** and then run these commands in order:
#
# ```
# git rm -r --cached .
# git add .
# git commit -m 'Removed all files that are in the .gitignore'
# git push origin master
# ```
#
# ## More resources
#
# [The manual for git](https://git-scm.com/docs/gitignore) describes how to use these files in much more detail.
| content/01/09_gitignore.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Assignment 3
# #### Student ID: *Double click here to fill the Student ID*
#
# #### Name: *Double click here to fill the name*
# # 3
# Suppose we have a data set with five predictors, $X_1$ = GPA, $X_2$ = IQ, $X_3$ = Level (1 for College and 0 for High School), $X_4$ = Interaction between GPA and IQ, and $X_5$ = Interaction between GPA and Level. The response is starting salary after graduation (in thousands of dollars). Suppose we use least squares to fit the model, and get $\hat{\beta}_0=50,\hat{\beta}_1=20,\hat{\beta}_2=0.07,\hat{\beta}_3=35,\hat{\beta}_4=0.01,\hat{\beta}_5=-10$.
# (a) Which answer is correct, and why?
# > (i) For a fixed value of IQ and GPA, high school graduates earn more, on average, than college graduates.
#
# > (ii) For a fixed value of IQ and GPA, college graduates earn more, on average, than high school graduates.
#
# > (iii) For a fixed value of IQ and GPA, high school graduates earn more, on average, than college graduates provided that the GPA is high enough.
#
# > (iv) For a fixed value of IQ and GPA, college graduates earn more, on average, than high school graduates provided that the GPA is high enough.
# > Ans: *double click here to answer the question.*
# (b) Predict the salary of a college graduate with IQ of 110 and a GPA of 4.0.
# > Ans: *double click here to answer the question.*
# (c) True or false: Since the coefficient for the GPA/IQ interaction term is very small, there is very little evidence of an interaction effect. Justify your answer.
# > Ans: *double click here to answer the question.*
# # 4
# I collect a set of data ($n = 100$ observations) containing a single predictor and a quantitative response. I then fit a linear regression model to the data, as well as a separate cubic regression, i.e. $Y = \beta_0 + \beta_1X + \beta_2X^2 + \beta_3X^3 + \epsilon$.
# (a) Suppose that the true relationship between X and Y is linear, i.e. $Y = \beta_0 + \beta_1X + \epsilon$. Consider the training residual sum of squares (RSS) for the linear regression, and also the training RSS for the cubic regression. Would we expect one to be lower than the other, would we expect them to be the same, or is there not enough information to tell? Justify your answer.
# > Ans: *double click here to answer the question.*
# (b) Answer (a) using test rather than training RSS.
# > Ans: *double click here to answer the question.*
# (c) Suppose that the true relationship between X and Y is not linear, but we don’t know how far it is from linear. Consider the training RSS for the linear regression, and also the training RSS for the cubic regression. Would we expect one to be lower than the other, would we expect them to be the same, or is there not enough information to tell? Justify your answer.
# > Ans: *double click here to answer the question.*
# (d) Answer (c) using test rather than training RSS.
# > Ans: *double click here to answer the question.*
# ## Applied
# # 10
# This question should be answered using the <span style='color:red'>Carseats</span> data set.
# (a) Fit a multiple regression model to predict <span style='color:red'>Sales</span> using <span style='color:red'>Price</span>, <span style='color:red'>Urban</span>, and <span style='color:red'>US</span>.
# (b) Provide an interpretation of each coefficient in the model. Be
# careful—some of the variables in the model are qualitative!
# > Ans: *double click here to answer the question.*
# (c) Write out the model in equation form, being careful to handle the qualitative variables properly.
# > Ans: *double click here to answer the question.*
# (d) For which of the predictors can you reject the null hypothesis $H_0 : \beta_j = 0$?
# > Ans: *double click here to answer the question.*
# (e) On the basis of your response to the previous question, fit a smaller model that only uses the predictors for which there is evidence of association with the outcome.
# (f) How well do the models in (a) and (e) fit the data?
# > Ans: *double click here to answer the question.*
# (g) Using the model from (e), obtain $95\%$ confidence intervals for
# the coefficient(s).
# # 14
# This problem focuses on the *collinearity* problem.
# (a) Perform the following commands in <span style='color:red'>Python</span>:
#
# ```python
# > np.random.seed(1)
# > x1 = np.random.normal(size=100)
# > x2 = 0.5*x1 + np.random.normal(size=100)/10
# > y = 2 + 2*x1 + 0.3*x2 + np.random.normal(size=100)
# ```
#
# The last line corresponds to creating a linear model in which <span style='color:red'>y</span> is
# a function of <span style='color:red'>x1</span> and <span style='color:red'>x2</span>. Write out the form of the linear model.
# What are the regression coefficients?
# > Ans: *double click here to answer the question.*
# (b) What is the correlation between <span style='color:red'>x1</span> and <span style='color:red'>x2</span>? Create a scatterplot
# displaying the relationship between the variables.
# > Ans: *double click here to answer the question.*
# (c) Using this data, fit a least squares regression to predict <span style='color:red'>y</span> using <span style='color:red'>x1</span> and <span style='color:red'>x2</span>. Describe the results obtained. What are $\hat{\beta}_0$, $\hat{\beta}_1$, and $\hat{\beta}_2$? How do these relate to the true $\beta_0$, $\beta_1$, and $\beta_2$? Can you reject the null hypothesis $H_0 : \beta_1 = 0$? How about the null hypothesis $H_0 : \beta_2 = 0$?
# > Ans: *double click here to answer the question.*
# (d) Now fit a least squares regression to predict <span style='color:red'>y</span> using only <span style='color:red'>x1</span>. Comment on your results. Can you reject the null hypothesis $H_0 : \beta_1 = 0$?
# > Ans: *double click here to answer the question.*
# (e) Now fit a least squares regression to predict <span style='color:red'>y</span> using only <span style='color:red'>x2</span>. Comment on your results. Can you reject the null hypothesis $H_0 : \beta_2 = 0$?
# > Ans: *double click here to answer the question.*
# (f) Do the results obtained in (c)–(e) contradict each other? Explain your answer.
# > Ans: *double click here to answer the question.*
# (g) Now suppose we obtain one additional observation, which was
# unfortunately mismeasured.
# ```python
# > x1 = np.append(x1,0.1)
# > x2 = np.append(x2,0.8)
# > y = np.append(y,6)
# ```
# Re-fit the linear models from (c) to (e) using this new data. What
# effect does this new observation have on the each of the models?
# In each model, is this observation an outlier? A high-leverage
# point? Both? Explain your answers.
# > Ans: *double click here to answer the question.*
| Assignment3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# [](https://colab.research.google.com/github/MonashDataFluency/python-web-scraping/blob/master/notebooks/section-3-API-based-scraping.ipynb)
# <img src="../images/api.png">
# ### A brief introduction to APIs
# ---
# In this section, we will take a look at an alternative way to gather data than the previous pattern based HTML scraping. Sometimes websites offer an API (or Application Programming Interface) as a service which provides a high level interface to directly retrieve data from their repositories or databases at the backend.
#
# From Wikipedia,
#
# > "*An API is typically defined as a set of specifications, such as Hypertext Transfer Protocol (HTTP) request messages, along with a definition of the structure of response messages, usually in an Extensible Markup Language (XML) or JavaScript Object Notation (JSON) format.*"
#
# They typically tend to be URL endpoints (to be fired as requests) that need to be modified based on our requirements (what we desire in the response body) which then returns some a payload (data) within the response, formatted as either JSON, XML or HTML.
#
# A popular web architecture style called `REST` (or representational state transfer) allows users to interact with web services via `GET` and `POST` calls (two most commonly used) which we briefly saw in the previous section.
#
# For example, Twitter's REST API allows developers to access core Twitter data and the Search API provides methods for developers to interact with Twitter Search and trends data.
#
# There are primarily two ways to use APIs :
#
# - Through the command terminal using URL endpoints, or
# - Through programming language specific *wrappers*
#
# For example, `Tweepy` is a famous python wrapper for Twitter API whereas `twurl` is a command line interface (CLI) tool but both can achieve the same outcomes.
#
# Here we focus on the latter approach and will use a Python library (a wrapper) called `wptools` based around the original MediaWiki API.
#
# One advantage of using official APIs is that they are usually compliant of the terms of service (ToS) of a particular service that researchers are looking to gather data from. However, third-party libraries or packages which claim to provide more throughput than the official APIs (rate limits, number of requests/sec) generally operate in a gray area as they tend to violate ToS. Always be sure to read their documentation throughly.
# ### Wikipedia API
# ---
# Let's say we want to gather some additional data about the Fortune 500 companies and since wikipedia is a rich source for data we decide to use the MediaWiki API to scrape this data. One very good place to start would be to look at the **infoboxes** (as wikipedia defines them) of articles corresponsing to each company on the list. They essentially contain a wealth of metadata about a particular entity the article belongs to which in our case is a company.
#
# For e.g. consider the wikipedia article for **Walmart** (https://en.wikipedia.org/wiki/Walmart) which includes the following infobox :
#
# 
#
# As we can see from above, the infoboxes could provide us with a lot of valuable information such as :
#
# - Year of founding
# - Industry
# - Founder(s)
# - Products
# - Services
# - Operating income
# - Net income
# - Total assets
# - Total equity
# - Number of employees etc
#
# Although we expect this data to be fairly organized, it would require some post-processing which we will tackle in our next section. We pick a subset of our data and focus only on the top **20** of the Fortune 500 from the full list.
#
# Let's begin by installing some of libraries we will use for this excercise as follows,
# sudo apt install libcurl4-openssl-dev libssl-dev
# !pip install wptools
# !pip install wikipedia
# !pip install wordcloud
# Importing the same,
# +
import json
import wptools
import wikipedia
import pandas as pd
print('wptools version : {}'.format(wptools.__version__)) # checking the installed version
# -
# Now let's load the data which we scrapped in the previous section as follows,
# If you dont have the file, you can use the below code to fetch it:
import urllib.request
url = 'https://raw.githubusercontent.com/MonashDataFluency/python-web-scraping/master/data/fortune_500_companies.csv'
urllib.request.urlretrieve(url, 'fortune_500_companies.csv')
fname = 'fortune_500_companies.csv' # scrapped data from previous section
df = pd.read_csv(fname) # reading the csv file as a pandas df
df.head() # displaying the first 5 rows
# | | rank | company_name | company_website |
# |---:|-------:|:-------------------|:---------------------------------|
# | 0 | 1 | Walmart | http://www.stock.walmart.com |
# | 1 | 2 | Exxon Mobil | http://www.exxonmobil.com |
# | 2 | 3 | Berkshire Hathaway | http://www.berkshirehathaway.com |
# | 3 | 4 | Apple | http://www.apple.com |
# | 4 | 5 | UnitedHealth Group | http://www.unitedhealthgroup.com |
#
# Let's focus and select only the top 20 companies from the list as follows,
no_of_companies = 20 # no of companies we are interested
df_sub = df.iloc[:no_of_companies, :].copy() # only selecting the top 20 companies
companies = df_sub['company_name'].tolist() # converting the column to a list
# Taking a brief look at the same,
for i, j in enumerate(companies): # looping through the list of 20 company
print('{}. {}'.format(i+1, j)) # printing out the same
# ### Getting article names from wiki
# Right off the bat, as you might have guessed, one issue with matching the top 20 Fortune 500 companies to their wikipedia article names is that both of them would not be exactly the same i.e. they match character for character. There will be slight variation in their names.
#
# To overcome this problem and ensure that we have all the company names and its corresponding wikipedia article, we will use the `wikipedia` package to get suggestions for the company names and their equivalent in wikipedia.
wiki_search = [{company : wikipedia.search(company)} for company in companies]
# Inspecting the same,
for idx, company in enumerate(wiki_search):
for i, j in company.items():
print('{}. {} :\n{}'.format(idx+1, i ,', '.join(j)))
print('\n')
# Now let's get the most probable ones (the first suggestion) for each of the first 20 companies on the Fortune 500 list,
# +
most_probable = [(company, wiki_search[i][company][0]) for i, company in enumerate(companies)]
companies = [x[1] for x in most_probable]
print(most_probable)
# -
# We can notice that most of the wiki article titles make sense. However, **Apple** is quite ambiguous in this regard as it can indicate the fruit as well as the company. However we can see that the second suggestion returned by was **Apple Inc.**. Hence, we can manually replace it with **Apple Inc.** as follows,
companies[companies.index('Apple')] = 'Apple Inc.' # replacing "Apple"
print(companies) # final list of wikipedia article titles
# ### Retrieving the infoboxes
# Now that we have mapped the names of the companies to their corresponding wikipedia article let's retrieve the infobox data from those pages.
#
# `wptools` provides easy to use methods to directly call the MediaWiki API on our behalf and get us all the wikipedia data. Let's try retrieving data for **Walmart** as follows,
page = wptools.page('Walmart')
page.get_parse() # parses the wikipedia article
# As we can see from the output above, `wptools` successfully retrieved the wikipedia and wikidata corresponding to the query **Walmart**. Now inspecting the fetched attributes,
page.data.keys()
# The attribute **infobox** contains the data we require,
page.data['infobox']
# Let's define a list of features that we want from the infoboxes as follows,
wiki_data = []
# attributes of interest contained within the wiki infoboxes
features = ['founder', 'location_country', 'revenue', 'operating_income', 'net_income', 'assets',
'equity', 'type', 'industry', 'products', 'num_employees']
# Now fetching the data for all the companies (this may take a while),
for company in companies:
page = wptools.page(company) # create a page object
try:
page.get_parse() # call the API and parse the data
if page.data['infobox'] != None:
# if infobox is present
infobox = page.data['infobox']
# get data for the interested features/attributes
data = { feature : infobox[feature] if feature in infobox else ''
for feature in features }
else:
data = { feature : '' for feature in features }
data['company_name'] = company
wiki_data.append(data)
except KeyError:
pass
# Let's take a look at the first instance in `wiki_data` i.e. **Walmart**,
wiki_data[0]
# So, we have successfully retrieved all the infobox data for the companies. Also we can notice that some additional wrangling and cleaning is required which we will perform in the next section.
# Finally, let's export the scraped infoboxes as a single JSON file to a convenient location as follows,
with open('infoboxes.json', 'w') as file:
json.dump(wiki_data, file)
# ### References
#
# - https://phpenthusiast.com/blog/what-is-rest-api
# - https://github.com/siznax/wptools/wiki/Data-captured
# - https://en.wikipedia.org/w/api.php
# - https://wikipedia.readthedocs.io/en/latest/code.html
| notebooks/section-3-API-based-scraping.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
import matplotlib.pyplot as plt
from IPython.display import display, HTML
from pandas.plotting import scatter_matrix
from sklearn.linear_model import LogisticRegression
from sklearn.preprocessing import PolynomialFeatures
from sklearn.linear_model import Lasso
from sklearn.linear_model import Ridge
# # DATASET
# The dataset below has been explained in earlier notebooks and is being used here to carry on the data.
#
contraceptive_data = pd.read_csv("contraceptive_method_dataset.csv",
encoding = "ISO-8859-1", engine='python')
X = contraceptive_data.drop('children', axis=1).copy()
contraceptive_data['predictor_population']= pd.cut(contraceptive_data['children'],
[-1,2,16], labels=[0,1])
contraceptive_data['predictor_population_i']= contraceptive_data['predictor_population'].astype(int)
y= contraceptive_data['predictor_population_i']
# # Splitting of the Training and Testing Data
# Training mean and std calculated when the X train was fitted into the std scaler.
# For the X train scaled, the training mean and std was used only.
# While for the X test scaled, the test data was transformed.
#
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.3,
random_state = 24, stratify = y )
std_scaler = StandardScaler()
std_scaler.fit(X_train)
X_train_scaled = std_scaler.transform(X_train)
X_test_scaled = std_scaler.transform(X_test)
# # K-Mean
# ## The Calinski Harbasz Method
# +
## install yellowbrick library
from yellowbrick.cluster import KElbowVisualizer
## From sklearn.cluster we will call KMeans
from sklearn.cluster import KMeans
# Instantiate the clustering model and visualizer
model = KMeans()
# -
visualizer = KElbowVisualizer(model, k=(2,10), metric = 'calinski_harabasz', timings=False)
visualizer.fit(X) # Fit the data to the visualizer
visualizer.show() # Finalize and render the figure
# ## The Silhouette Method
visualizer = KElbowVisualizer(model, k=(2,10), metric = 'silhouette', timings=False)
visualizer.fit(X) # Fit the data to the visualizer
visualizer.show() # Finalize and render the figure
# _Note_
# - Two methods of Kmeans are used in this project which are the Calinski Harbabasz and the Silhouette. The Calinski Harbabasz's elbow is more pronounced which can be found in 3 and the score of 3428.76. On the other hand, Silhouette elbow can not be detected in this chart.
# # Ridge, Lasso and Logistic Regression Test Model
#
# +
def model_experiment(num_iter = 5,
models = ['logreg', 'ridge', 'lasso'], alpha= 10,
complexity = 'simple', degree = 3):
x_axis = np.arange(num_iter)
y_logreg_test = []
y_lasso_test = []
y_ridge_test = []
sample_models = {}
for i in range(num_iter):
if complexity == 'simple':
## split train_test
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.2)
elif complexity == 'polynomial':
## Create higher order terms
poly = PolynomialFeatures(degree=degree)
Xp = poly.fit_transform(X)
## test-train split
X_train, X_test, y_train, y_test = train_test_split(Xp, y, test_size = 0.2)
## Standard scale mean = 0, variance = 1
sd = StandardScaler()
sd.fit(X_train)
X_train = sd.transform(X_train)
X_test = sd.transform(X_test)
## Be careful about the leakage
## Vanilla model
if 'logreg' in models:
lr = LogisticRegression()
lr.fit(X_train, y_train)
sample_models['logreg'] = lr
test_score = lr.score(X_test, y_test)
train_score = lr.score(X_train, y_train)
y_logreg_test.append(test_score)
# print('test score logreg is %.2f and train score is %.2f'%(test_score, train_score))
if 'ridge' in models:
## Ridge in the simple setting
ridge = Ridge(alpha = alpha, max_iter= 10000)
ridge.fit(X_train, y_train)
sample_models['ridge'] = ridge
y_ridge_test.append(ridge.score(X_test, y_test))
# print('test score Ridge is %.2f and train score is %.2f'%(ridge.score(X_test, y_test),
# ridge.score(X_train, y_train)))
if 'lasso' in models:
## Lasso in the simple setting
lasso = Lasso(alpha = alpha, max_iter= 10000)
lasso.fit(X_train, y_train)
sample_models['lasso'] = lasso
y_lasso_test.append(lasso.score(X_test, y_test))
# print('test score Lasso is %.2f and train score is %.2f'%(lasso.score(X_test, y_test),
# lasso.score(X_train, y_train)))
i+=1
if 'logreg' in models:
plt.plot(y_logreg_test, label = 'logreg')
if 'ridge' in models:
plt.plot(y_ridge_test, label = 'ridge')
if 'lasso' in models:
plt.plot(y_lasso_test, label = 'lasso')
plt.ylabel('R2 test score')
plt.xlabel('number of iterations')
all_results = y_logreg_test + y_lasso_test + y_ridge_test
plt.ylim((np.min(all_results), np.max(all_results)))
plt.legend()
return sample_models
trained_models = model_experiment(num_iter=30, alpha = 15,
models = ['ridge', 'lasso'],
complexity= 'polynomial', degree = 2)
# -
# _Notes_
# - The R2 test score for ridge has more dips and peaks than lasso. Lasso and Ridge graph has a variance between 0 to .40. There difference lies in the question which becomes evident in the outcome. The ridge equation shrinks the slope asymphotically close to 0 while the lasso can shrink all the way to 0. Lasso can exclude useless variables from the equations while ridge performs better when most variables are useful.
#
trained_models = model_experiment(num_iter=30, alpha = 15,
models = ['logreg', 'ridge', 'lasso'],
complexity= 'polynomial', degree = 2)
# _Notes_
# - Based on the chart, the lasso is flatter than logistic regression and ridge, but lasso is closer in (variance) distance from ridge than logistic regression. However, logistic regression have a very similar dip and peaks with ridge. As mentioned earlier, lasso shrinks variables that it deems useless.
# +
# After run model_experiment with complexity == 'polynomial'
lr_logreg = trained_models['logreg']
lr_lasso = trained_models['lasso']
lr_ridge =trained_models['ridge']
# -
lr_lasso.coef_
lr_ridge.coef_
lr_logreg.coef_
# _Notes_
# - As we see from above, lasso uses 0 coefficient which allows more predictability, and there are many useless variables in the data which is why it shrinks to 0.
| Final Project/CM_K-Means, LogReg, Lasso and Ridge Charts.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
A = np.matrix('1 2 3; 4 5 6')
print(A)
A_t = A.transpose()
print(A_t)
print(A.T)
A = np.matrix('1 2 3; 4 5 6')
print(A)
R = (A.T).T
print(R)
A = np.matrix('1 2 3; 4 5 6')
B = np.matrix('7 8 9; 0 7 5')
L = (A + B).T
R = A.T + B.T
print(L)
print(R)
A = np.matrix('1 2; 3 4')
B = np.matrix('5 6; 7 8')
L = (A.dot(B)).T
R = (B.T).dot(A.T)
print(L)
print(R)
A = np.matrix('1 2 3; 4 5 6')
k = 3
L = (k * A).T
R = k * (A.T)
print(L)
print(R)
A = np.matrix('1 2; 3 4')
A_det = np.linalg.det(A)
A_T_det = np.linalg.det(A.T)
print(format(A_det, '.9g'))
A = np.matrix('1 2 3; 4 5 6')
C = 3 * A
print(C)
A = np.matrix('1 2; 3 4')
p = 2
q = 3
L = (p + q) * A
R = p * A + q * A
print(L)
print(R)
A = np.matrix('1 2; 3 4')
p = 2
q = 3
L = (p * q) * A
R = p * (q * A)
print(L)
print(R)
A = np.matrix('1 2; 3 4')
B = np.matrix('5 6; 7 8')
k = 3
L = k * (A + B)
R = k * A + k * B
print(L)
print(R)
A = np.matrix('1 6 3; 8 2 7')
B = np.matrix('8 1 5; 6 9 12')
C = A + B
print(C)
A = np.matrix('1 2; 3 4')
B = np.matrix('5 6; 7 8')
L = A + B
R = B + A
print(L)
print(R)
A = np.matrix('1 2; 3 4')
B = np.matrix('5 6; 7 8')
C = np.matrix('1 7; 9 3')
L = A + (B + C)
R = (A + B) + C
print(L)
print(R)
A = np.matrix('1 2; 3 4')
Z = np.matrix('0 0; 0 0')
L = A + (-1)*A
print(L)
print(Z)
A = np.matrix('1 2 3; 4 5 6')
B = np.matrix('7 8; 9 1; 2 3')
C = A.dot(B)
print(C)
A = np.matrix('1 2; 3 4')
B = np.matrix('5 6; 7 8')
C = np.matrix('2 4; 7 8')
L = A.dot(B.dot(C))
R = (A.dot(B)).dot(C)
print(L)
print(R)
A = np.matrix('1 2; 3 4')
B = np.matrix('5 6; 7 8')
C = np.matrix('2 4; 7 8')
L = A.dot(B + C)
R = A.dot(B) + A.dot(C)
print(L)
print(R)
A = np.matrix('1 2; 3 4')
E = np.matrix('1 0; 0 1')
L = E.dot(A)
R = A.dot(E)
print(L)
print(R)
print(A)
A = np.matrix('-4 -1 2; 10 4 -1; 8 3 1')
print(A)
A = np.matrix('-4 -1 2; 10 4 -1; 8 3 1')
print(A)
print(A.T)
det_A = round(np.linalg.det(A), 3)
det_A_t = round(np.linalg.det(A.T), 3)
print(det_A)
print(det_A_t)
A = np.matrix('-4 -1 2; -4 -1 2; 8 3 1')
print(A)
np.linalg.det(A)
round(np.linalg.det(B), 3)
A = np.matrix('-4 -1 2; -4 -1 2; 8 3 1')
print(A)
np.linalg.det(A)
A = np.matrix('-4 -1 2; 10 4 -1; 8 3 1')
print(A)
k = 2
B = A.copy()
B[2, :] = k * B[2, :]
print(B)
det_A = round(np.linalg.det(A), 3)
det_B = round(np.linalg.det(B), 3)
det_A * k
det_B
A = np.matrix('-4 -1 2; 10 4 -1; 8 3 1')
print(A)
k = 2
A[1, :] = A[0, :] + k * A[2, :]
round(np.linalg.det(A), 3)
A = np.matrix('-4 -1 2; 10 4 -1; 8 3 1')
print(A)
k = 2
A[1, :] = k * A[0, :]
print(A)
round(np.linalg.det(A), 3)
A = np.matrix('1 -3; 2 5')
A_inv = np.linalg.inv(A)
print(A_inv)
A = np.matrix('1. -3.; 2. 5.')
A_inv = np.linalg.inv(A)
A_inv_inv = np.linalg.inv(A_inv)
print(A)
print(A_inv_inv)
A = np.matrix('1. -3.; 2. 5.')
L = np.linalg.inv(A.T)
R = (np.linalg.inv(A)).T
print(L)
print(R)
m_eye = np.eye(4)
print(m_eye)
rank = np.linalg.matrix_rank(m_eye)
print(rank)
| pr_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
x=np.random.randn(32)
sns.displot(x, kind="kde",fill=True)
x.mean(), x.std()
x_shift=5*x+3
x_shift.mean(), x.std()
sns.displot(x_shift, kind="kde", fill=True)
x_original=(1/5)*x_shift -3/5
unscaled=x_shift -3/5
unscaled_1=x_shift +10
x_original.mean()
sns.distplot(x_shift, kind="kde", fill=True)
sns.distplot(x_original, kind="kde", fill=True)
plt.show()
sns.distplot(x_shift, hist=False, rug=True)
sns.distplot(x_original, hist=False, rug=True)
plt.show()
sns.distplot(x_shift, hist=False, rug=True)
sns.distplot(x_original, hist=False, rug=True)
sns.distplot(unscaled, hist=False, rug=True)
plt.show()
sns.distplot(x_shift, hist=False, rug=True)
sns.distplot(x_original, hist=False, rug=True)
sns.distplot(unscaled_1, hist=False, rug=True)
plt.show()
| Optimizer/Batch_Norm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:miniconda3-s2s]
# language: python
# name: conda-env-miniconda3-s2s-py
# ---
# # 0.0 Launch Cluster
#
# ---
#
# Use this to start your `dask` cluster of workers. Uncomment `Cheyenne` if you're using Cheyenne. You can then open the `dask` tab in JupyterLab and type `proxy/####` where `####` is the proxy number from the Dashboard link under Client. Then you can drag tabs like Dask Workers, Dask Progress, etc. to the sidebar. You'll copy and paste the Scheduler tcp number into subsequent notebooks.
# +
#from ncar_jobqueue import NCARCluster
#from dask.distributed import Client
# Import dask
import dask
# Use dask jobqueue
from dask_jobqueue import PBSCluster
# Import a client
from dask.distributed import Client
# +
# Casper
NUMNODES = 3
num_processes = 9
#num_processes = 27
num_threads = 18
MEM = "200GB"
# Cheyenne
# NUMNODES = 5
# num_processes = 18
# num_threads = 36
# MEM = "100GB"
#cluster = NCARCluster(cores=num_threads,
# processes=num_processes,
# memory=MEM, walltime="03:00:00",
# project="p66770001")
#cluster = NCARCluster(cores=num_threads,
# processes=num_processes,
# memory=MEM,
# walltime="02:00:00",
# )
#cluster.scale(NUMNODES * num_processes)
# Setup your PBSCluster
cluster = PBSCluster(
cores=num_threads, # The number of cores you want
memory=MEM, # Amount of memory
processes=num_processes, # How many processes
queue='casper', # The type of queue to utilize (/glade/u/apps/dav/opt/usr/bin/execcasper)
local_directory='$TMPDIR', # Use your local directory
#resource_spec='select=1:ncpus=1:mem=10GB', # Specify resources
project='NASP0002', # Input your project ID here
walltime='02:00:00', # Amount of wall time
#interface='ib0', # Interface to use
)
# Scale up
cluster.scale(NUMNODES * num_processes)
# -
dask.config.set({'distributed.dashboard.link':'https://jupyterhub.hpc.ucar.edu/stable/user/{USER}/proxy/{port}/status'})
client = Client(cluster)
cluster
client
| 0.0_launch_cluster.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from matplotlib import colors
from matplotlib.ticker import PercentFormatter
import seaborn as sns
from sklearn.model_selection import train_test_split
from sklearn import preprocessing
from sklearn.preprocessing import MinMaxScaler
from sklearn.metrics import classification_report, confusion_matrix
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Dropout
from tensorflow.keras.callbacks import EarlyStopping
from tensorflow.keras.utils import to_categorical
from tensorflow.keras.regularizers import l1
from tensorflow import keras
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import LSTM,Dense ,Dropout
from tensorflow.keras import layers
import datetime
import math
# -
df = pd.read_excel('MaruteruOriginalDatainCM.xlsx')
df.head()
df.head()
df = df.rename(columns={"Year": "year", 'MaxT':'MAX','MinT':'MIN', 'RH1': 'RHI', 'RH2':'RHII', 'EVPcm':'EVAP'})
df.info()
df.describe().transpose()
df.isnull().sum()
sns.set_theme(style="ticks", color_codes=True)
#Drop Emp_Title as it is not required
df = df.drop(["SSH","SMW"], axis=1)
gby_df = df.groupby(['year'],as_index=False).mean()
gby_df
gby_df.plot(x="year", y=['MAX', 'MIN', 'RHI', 'RHII', 'WS', 'RF',
'EVAP'], kind="bar", title="Compare All vs Year (Avg)", figsize=(12,8))
plt.box(False)
gby_df.plot(x="year", y=['MAX', 'MIN', 'RHI', 'RHII', 'WS', 'RF',
'EVAP'], kind="line", title="Compare All vs Year (Avg)", figsize=(12,8))
plt.box(False)
gby_df.plot(x="year", y=['MAX', 'MIN', 'RHI', 'RHII', 'WS', 'RF',
'EVAP'], kind="area", title="Compare All vs Year (Avg)", figsize=(12,8))
plt.box(False)
gby_df.plot(x="year", y=["RF", "EVAP"], kind="bar", title="Avg of Rainfall vs Evaporation per year", figsize=(10,6))
plt.box(True)
ax = gby_df.plot(x="year", y=["RF"], kind="bar")
gby_df.plot(ax=ax, x="year", y=["EVAP"], kind="bar", color="orange", title="Rainfall vs Evaporation", figsize=(10,6))
plt.box(False)
s = sns.catplot(x="year", y="RF", kind="swarm", data=gby_df, height=8.27, aspect=11.7/8.27)
s.set_xlabels("Year")
s.set_ylabels("RainFall")
s.fig.suptitle('Rainfall vs Year')
s = sns.catplot(x="year", y="RF", kind="bar", data=gby_df, height=5.27, aspect=9.7/6.27)
s.set_xlabels("Year")
s.set_ylabels("RainFall")
s.fig.suptitle('Rainfall vs Year')
#Drop year as it is not required
df = df.drop(["year"], axis=1)
df.head()
# +
# correlation heat map
plt.figure(figsize=(12, 6))
df_corr = df.corr()
# min -1 and max +1 as we are plotting correlation
sns.heatmap(df.corr(), cmap="Blues", annot=True, fmt=".2f", vmin= -1.0, vmax=1.0,
linewidth=0.3, cbar_kws={"shrink": .8})
# yticks
plt.yticks(rotation=0)
plt.show()
# -
df = df.drop("RHII", axis=1)
df.columns
from scipy import stats
df[(np.abs(stats.zscore(df)) < 3).all(axis=1)]
X = df.drop('EVAP', axis=1).values
y = df['EVAP'].values
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2)
scaler = MinMaxScaler()
X_train = scaler.fit_transform(X_train)
X_test = scaler.transform(X_test)
X_train.shape
early_stop = EarlyStopping(monitor='val_loss', mode='min', verbose=1, patience=10)
from tensorflow.keras.models import load_model
from tensorflow.keras import layers, Input
from tensorflow.keras import Model
def build_model():
# Define model layers.
input_layer = Input(shape=(5,))
first_dense = Dense(units='128', activation='relu')(input_layer)
# Y1 output will be fed from the first dense
y1_output = Dense(units='1', name='layer1_output')(first_dense)
second_dense = Dense(units='128',activation='relu')(first_dense)
# Y2 output will be fed from the second dense
y2_output = Dense(units='1',name='layer2_output')(second_dense)
# Define the model with the input layer
# and a list of output layers
model = Model(inputs=input_layer,outputs=[y1_output, y2_output])
return model
model = build_model()
model.compile(loss="mean_squared_error", optimizer='adam')
model.summary()
model.fit(X_train, y_train, validation_data = (X_test, y_test), epochs=100, batch_size=5, callbacks=[early_stop])
losses = pd.DataFrame(model.history.history)
losses.plot()
test_predictions = model.predict(X_test)
test_predictions
pred_df = pd.DataFrame(y_test,columns=['Test Y'])
pred_df
test_predictions = pd.Series(test_predictions[0].reshape(177,))
pred_df = pd.concat([pred_df,test_predictions],axis=1)
pred_df.columns = ['Test Y','Model Predictions']
pred_df
sns.scatterplot(x='Test Y',y='Model Predictions',data=pred_df)
pred_df['Error'] = pred_df['Test Y'] - pred_df['Model Predictions']
sns.distplot(pred_df['Error'],bins=50)
# # Mean Absolute Error
from sklearn.metrics import mean_absolute_error,mean_squared_error
mean_absolute_error(pred_df['Test Y'],pred_df['Model Predictions'])
# # Mean Squared Error
mean_squared_error(pred_df['Test Y'],pred_df['Model Predictions'])
# # Predicting on brand new data
new_data = [[29.2,12.2,67.6,5.7,6.0]]
# Don't forget to scale!
scaler.transform(new_data)
new_data = scaler.transform(new_data)
model.predict(new_data)
# saving the model by serializing it using json (same thing can be done using YAML)
model_json = model.to_json()
with open("model.json","w") as json_file:
json_file.write(model_json)
model.save_weights("model.h5")
# ## Saving Scaler
import joblib
scaler_filename = "scaler.save"
joblib.dump(scaler, scaler_filename)
# ## Using saved scaler and model to predict the results
# ### Loading Saved Sacler
savedscaler = joblib.load("model/scaler.save")
# ### Loading Saved Model
from tensorflow.keras.models import model_from_json
json_file = open("model/model.json","r")
loaded_model_json = json_file.read()
json_file.close()
loaded_model = model_from_json(loaded_model_json)
loaded_model.load_weights("model/model.h5")
new_data = [[29.2,12.2,67.6,5.7,6.0]]
new_data = savedscaler.transform(new_data)
loaded_model.predict(new_data)
| Evapuration Prediction v2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Definition of the Business Problem
# A predictive model will be created to predict whether a customer is satisfied or dissatisfied. Historical data provided by Santandar will be used.
#
# Dataset: https://www.kaggle.com/c/santander-customer-satisfaction/overview
#
# The dataset has anonymous data from more than 70 thousand Santander customers, separated by the bank itself into two datasets, the first for training and the second for testing.
#
# The "TARGET" column is the variable to be predicted. It is equal to one for dissatisfied customers and 0 for satisfied customers.
#
# The task is to predict the likelihood that each customer in the test suite is a dissatisfied customer.
# # Imports
# +
# Import from libraries
from sklearn.model_selection import KFold
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import train_test_split
from xgboost import XGBClassifier
import pandas as pd
from sklearn.preprocessing import Normalizer
from sklearn.linear_model import LogisticRegression
from sklearn.tree import DecisionTreeClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
from sklearn.naive_bayes import GaussianNB
from sklearn.svm import SVC
import matplotlib.pyplot as plt
from sklearn.metrics import confusion_matrix
from sklearn.metrics import classification_report
import seaborn as sns
import pickle
import warnings
warnings.filterwarnings("ignore")
# -
# # Extracting and Loading Data
# The files were uploaded in CSV format, Santander provided one file for training and another for testing. However, during the project it was decided to use the training base for training and tests, since the test base does not have a "TARGET" column for future performance checks on the model. At the end of the data processing, the base is separated from the sklearn.
# Loading the training dataset in CSV format
training_file = 'data/train.csv'
test_file = 'data/test.csv'
data_training = pd.read_csv(training_file)
test_data = pd.read_csv (test_file)
print(data_training.shape)
print(test_data.shape)
# # Exploratory Data Analysis
# Viewing the first 20 lines
data_training.head (20)
# Data type of each attribute
data_training.dtypes
# Statistical Summary
data_training.describe()
# Distribution of classes
data_training.groupby("TARGET").size()
# # DATA PROCESSING
# As seen above, there are many more satisfied customers (class 0) than dissatisfied customers (class 1), so the dataset is totally unbalanced. Thus, it is chosen to perform a simple balancing, based on the division of data by class and capturing a sample of class 0 that has more data, this sample is the same size as class 1. Thus, you will have a fully balanced dataset with aa Class 0 and 1 in the same quantity.
# +
# Dividing by class
data_class_0 = data_training[data_training['TARGET'] == 0]
data_class_1 = data_training[data_training['TARGET'] == 1]
counter_class_0 = data_class_0.shape[0]
contador_classe_1 = data_class_1.shape[0]
data_class_0_sample = data_class_0.sample(counter_class_0)
training_data = pd.concat([data_class_0_sample, data_class_1], axis = 0)
# -
# Below, Pearson's correlation is used to identify the attributes that have minimal correlation above the limit. In this way it is possible to guarantee the variables with the best performance. As it is a dataset with many columns (371), no variable has a prominent correlation, so I chose to put a significant minimum value to reduce the variables by at least half.
# Pearson correlation
data_training.corr(method = 'pearson')
# Finding the correlation between the target variable and the predictor variables
corr = training_data[training_data.columns [1:]].corr()['TARGET'][:].abs()
minimal_correlation = 0.02
corr2 = corr[corr > minimal_correlation]
corr2.shape
corr2
corr_keys = corr2.index.tolist()
data_filter = data_training[corr_keys]
data_filter.head(20)
data_filter.dtypes
# Finally, the columns are filtered according to the Pearson correlation and the normalization of the predictive data is performed.
# +
# Filtering only the columns that have a correlation above the minimum variable
array_treino = data_training[corr_keys].values
# Separating the array into input and output components for training data
X = array_treino[:, 0:array_treino.shape[1] - 1]
Y = array_treino[:, array_treino.shape[1] - 1]
# Creating the training and test dataset
test_size = 0.30
X_training, X_testing, Y_training, Y_testing = train_test_split(X, Y, test_size = test_size)
# Generating normalized data
scaler = Normalizer (). fit (X_training)
normalizedX_treino = scaler.transform(X_training)
scaler = Normalizer().fit(X_testing)
normalizedX_teste = scaler.transform(X_testing)
Y_training = Y_training.astype('int')
Y_testing = Y_testing.astype('int')
# -
# # TRAINING
# Execution of a series of classification algorithms is based on those that have the best result. For this test, the training base is used without any treatment or data selection.
# +
# Setting the number of folds for cross validation
num_folds = 10
# Preparing the list of models
models = []
models.append(('LR', LogisticRegression()))
models.append(('LDA', LinearDiscriminantAnalysis()))
models.append(('NB', GaussianNB()))
models.append(('KNN', KNeighborsClassifier()))
models.append(('CART', DecisionTreeClassifier()))
models.append(('SVM', SVC()))
# +
results = []
names = []
for name, model in models:
kfold = KFold (n_splits = num_folds)
cv_results = cross_val_score (model, X_training, Y_training, cv = kfold, scoring = 'accuracy')
results.append (cv_results)
names.append (name)
msg = "% s:% f (% f)"% (name, cv_results.mean (), cv_results.std ())
print (msg)
# Boxplot to compare the algorithms
fig = plt.figure ()
fig.suptitle ('Comparison of Classification Algorithms')
ax = fig.add_subplot (111)
plt.boxplot (results)
ax.set_xticklabels (names)
plt.show ()
# -
# After some tests, the final training is started with the chosen algorithms, based on their respective performances.
# Function to evaluate the performance of the model and save it in a pickle format for future reuse.
def model_report(model_name):
# Print result
print("Accuracy:% .3f"% score)
# Making predictions and building the Confusion Matrix
predictions = result.predict(X_testing)
matrix = confusion_matrix(Y_testing, predictions)
print(matrix)
report = classification_report(Y_testing, predictions)
print(report)
# The precision matrix is created to visualize the number of correct cases
labels = ['SATISFIED', 'UNSATISFIED']
cm = confusion_matrix(Y_testing, predictions)
cm = pd.DataFrame(cm, index = ['0', '1'], columns = ['0', '1'])
plt.figure(figsize = (10.10))
sns.heatmap(cm, cmap = "Blues", linecolor = 'black', linewidth = 1, annot = True, fmt = '', xticklabels = labels, yticklabels = labels)
# Saving the model
file = 'models/final_classifier_model' + model_name + '.sav'
pickle.dump (model, open(file, 'wb'))
print("Saved Model!")
# +
# Linear Regression
model = LogisticRegression()
result = model.fit(normalizedX_treino, Y_testing)
score = result.score(normalizedX_treino, Y_testing)
model_report("LR")
# Linear Discriminant Analysis
model = LinearDiscriminantAnalysis()
result = model.fit(X_training, Y_testing)
score = result.score(X_training, Y_testing)
model_report("LDA")
# KNN
model = KNeighborsClassifier()
result = model.fit(normalizedX_treino, Y_testing)
score = result.score(normalizedX_treino, Y_testing)
model_report("KNN")
# CART
model = DecisionTreeClassifier()
result = model.fit(X_training, Y_testing)
score = result.score(X_training, Y_testing)
model_report("CART")
# XGBOOST
model = XGBClassifier()
result = model.fit(X_training, Y_testing)
score = result.score(X_training, Y_testing)
model_report("XGBOOST")
# +
# Loading the model
file = 'models model_classifier_final_XGBOOST.sav'
model_classifier = pickle.load(open(file, 'rb'))
model_prod = model_classifier.score(X_testing, Y_testing)
print("Uploaded Model")
# Print Result
print("Accuracy:% .3f"% (model_prod.mean () * 100))
# -
# After performing several tests it was seen that the models with the best accuracy were LDA, KNN AND XGBOOST. These models showed accuracy greater than 70%. Even so, the XGBOOST is more accurate with 75%.
| Santander-jupyter-EN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (Data Science)
# language: python
# name: python3__SAGEMAKER_INTERNAL__arn:aws:sagemaker:eu-central-1:936697816551:image/datascience-1.0
# ---
import sagemaker
import boto3
from sagemaker import get_execution_role
# +
region = boto3.Session().region_name
sm_rt = boto3.Session().client('runtime.sagemaker', region_name=region)
ep_name = 'd-scs-model'
id_name = "ID"
target = 'TARGET'
file_name = "./d-scs/test_Santander Customer Satisfaction.csv"
test_file_name = "./d-scs/test_Santander Customer Satisfaction-no-label.csv"
kaggle_file_name = "./d-scs/kaggle-test-Santander Customer Satisfaction.csv"
kaggle_test_file_name = "./d-scs/kaggle-test-Santander Customer Satisfaction-no-label.csv"
kaggle_pred_file_name = "./d-scs/kaggle-test-Santander Customer Satisfaction-predictions.csv"
# +
import pandas as pd
import numpy as np
df = pd.read_csv(file_name)
df = df.drop([target],axis=1)
print(df.head())
print(df.shape)
df.to_csv(test_file_name, index=False)
# +
f = open(test_file_name)
count = 0
pred_label = []
pred_neg_prpb = []
pred_pos_prpb = []
for line in f:
if count==0:
count=1
continue
response = sm_rt.invoke_endpoint(EndpointName=ep_name, ContentType='text/csv', Accept='text/csv', Body=line.encode('utf-8'))
response = response['Body'].read().decode("utf-8")
#print(response)
pred_label.append(int(response[0]))
ind1 = response.index("[")
ind2 = response.index(",", ind1)
neg_prob = float(response[ind1+1:ind2])
#print(neg_prob)
pred_neg_prpb.append(neg_prob)
ind1 = response.index("]", ind2)
pos_prob = float(response[ind2+2:ind1])
#print(pos_prob)
pred_pos_prpb.append(pos_prob)
# +
print(pred_label[:10])
print(pred_neg_prpb[:10])
print(pred_pos_prpb[:10])
import pandas as pd
import numpy as np
pd = pd.read_csv(file_name)
y_true = np.array(pd[target])
y_pred = np.array(pred_label)
print(np.mean(y_true==y_pred))
from sklearn import metrics
#fpr, tpr, thresholds = metrics.roc_curve(y_true, pred_pos_prpb, pos_label=1)
#metrics.auc(fpr, tpr)
metrics.roc_auc_score(y_true, pred_pos_prpb)
# +
import pandas as pd
import numpy as np
df = pd.read_csv(kaggle_file_name)
kaggle_test_ids = np.array(df[id_name])
print(df.head())
print(df.shape)
df.to_csv(kaggle_test_file_name, index=False)
f = open(kaggle_test_file_name)
count = 0
kaggle_pred = []
for line in f:
if count==0:
count=1
continue
response = sm_rt.invoke_endpoint(EndpointName=ep_name, ContentType='text/csv', Accept='text/csv', Body=line.encode('utf-8'))
response = response['Body'].read().decode("utf-8")
test_id = kaggle_test_ids[count-1]
pred = int(response[0])
kaggle_pred.append([test_id, pred])
count = count + 1
kaggle_pred_df = pd.DataFrame(kaggle_pred, columns = [id_name, target])
kaggle_pred_df.to_csv(kaggle_pred_file_name, index=False)
# -
print(kaggle_pred_df.head())
print(kaggle_pred_df.shape)
import boto3
sm = boto3.client('sagemaker')
#sm.delete_endpoint_config(EndpointConfigName=endpoint_config_name)
sm.delete_endpoint(EndpointName=ep_name)
| code/aws_sagemaker/d-scs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="7_B-El2kGidl"
import pandas as pd
import datetime
import requests, io
import numpy as np
# + id="_B2VWVdmGfKi"
#importação da base
url = 'http://www.anatel.gov.br/dadosabertos/PDA/PBLE/PBLE.csv'
pble = pd.read_csv(url, sep=';', encoding='latin-1')
# + id="E7sP5ioa20_9"
#renomeando variáveis
pble.rename(columns={'Prestadora':'empresa', 'Nº INEP': 'id_escola', 'UF':'sigla_uf', 'Municipio':'municipio', 'Situação':'situacao',
'Tipo de Obrigação':'programa', 'Velocidade de Acesso Instalada':'conexao', 'Tecnologia':'tecnologia',
'Nome da Escola':'nome_escola', 'Tipo Escola':'rede', 'Data de Ativação':'data_ativacao'}, inplace=True)
# + id="LkeX12Q15Wgd"
#exclusão de variaveis unicas: todas instaladas, programa PBLE, mes/ano de atualizacao dos dados
pble.drop(['situacao', 'programa', 'Mes', 'Ano', 'nome_escola'], axis=1, inplace=True)
# + id="yC6HseGm2tXL"
#transformar pra formato data
pble['data_ativacao']=pd.to_datetime(pble['data_ativacao'].astype(str), format='%d/%m/%Y')
# + id="NC7mHilwAski"
#criacao de colunas separadas por ano/mes/dia
pble['ano'] = pble['data_ativacao'].dt.year
pble['mes'] = pble['data_ativacao'].dt.month
pble['dia'] = pble['data_ativacao'].dt.day
# + id="BxAzjmgMkDmR"
#retirar [*] de algumas observações
pble['id_escola'] = pble['id_escola'].apply(lambda x: str(x).replace('[*]',''))
# + id="03Rj3wZKDqHT"
sigla_uf_dic ={'RO':11, 'AC':12, 'AM':13, 'RR': 14, 'PA':15, 'AP':16, 'TO':17,
'MA':21, 'PI':22, 'CE':23, 'RN':24, 'PB':25, 'PE':26, 'AL': 27, 'SE':28, 'BA':29,
'MG':31, 'ES':32, 'RJ':33, 'SP':35,
'PR': 41, 'SC':42, 'RS':43,
'MS':50, 'MT':51, 'GO':52, 'DF':53}
pble['id_uf'] = pble['sigla_uf'].map(sigla_uf_dic)
# + id="L9vCgM1cMfUj"
pble.replace(regex={r'São Luís do Paraitinga': 'São Luiz do Paraitinga', "Pau D'Arco": "Pau d'Arco", 'Augusto Severo':'Augusto Severo (Campo Grande)',
'Januário Cicco':'Januário Cicco (Boa Saúde)', 'Quixaba':'Quixabá', 'São Caitano':'São Caetano', 'Graccho Cardoso':'Gracho Cardoso',
"Pau d'Arco do Piauí":"Pau D'Arco do Piauí", "Olhos-d'Água":"Olhos d'Água"}, inplace=True)
# + colab={"base_uri": "https://localhost:8080/"} id="3QG0azRlBbb1" outputId="ee319eec-b39f-4f9a-d533-d5914f3e50a8"
#inclusão do id_municipio (código do ibge) pelo nome do municipio
url = 'https://raw.githubusercontent.com/kelvins/Municipios-Brasileiros/main/csv/municipios.csv'
read_data = requests.get(url).content
read_data
# + id="HnKf0K9QB1L3"
id_municipio = pd.read_csv(io.StringIO(read_data.decode('utf-8')))
id_municipio.drop(['capital', 'latitude', 'longitude'], axis=1, inplace=True)
# + id="Q6GxHsHWCd3E"
pble = pd.merge(pble, id_municipio,
how='left', left_on=['municipio', 'id_uf'],
right_on=['nome', 'codigo_uf'])
# + colab={"base_uri": "https://localhost:8080/"} id="LjhEjONbFkgu" outputId="938536f4-531d-41a4-92c7-e2a0a0c329e8"
pble['codigo_ibge'].loc[(pble['sigla_uf'] =='TO') & (pble['municipio']=="Pau d'Arco") & (pble['codigo_ibge'].isnull())]=1716307
pble['codigo_ibge'].loc[(pble['sigla_uf'] =='PI') & (pble['municipio']=="Pau d'Arco do Piauí") & (pble['codigo_ibge'].isnull())]=2207793
pble['codigo_ibge'].loc[(pble['sigla_uf'] =='PE') & (pble['municipio']=="Quixabá") & (pble['codigo_ibge'].isnull())]=2611533
# + id="h0u_109kDcqd"
#organização das variaveis
pble.drop(['municipio', 'data_ativacao','id_uf', 'nome', 'codigo_uf'], axis=1, inplace=True)
pble.rename(columns={'codigo_ibge':'id_municipio'}, inplace=True)
pble = pble[['ano', 'mes', 'dia', 'sigla_uf', 'id_municipio', 'rede', 'id_escola', 'empresa', 'tecnologia', 'conexao']]
# + colab={"base_uri": "https://localhost:8080/"} id="dSMIaY_vk4oW" outputId="a1fa83cf-657e-47d3-b821-c4ce2a6619e9"
#salvar arquivos no Drive
from google.colab import drive
drive.mount('/content/drive')
# + id="BDbIOPFck7PW"
#exportação para csv
pble.to_csv('/content/drive/MyDrive/br_anatel/banda_larga_fixa/output/banda_larga_pble', index=False)
| bases/br_anatel_banda_larga_fixa/code/br_anatel_banda_larga_pble.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/souravgopal25/DeepLearnigNanoDegree/blob/master/GradientDescent.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="QTvvtZCTY_Zl" colab_type="text"
# #GRADIENT DESCENT
# + id="CJcgbev8YpbX" colab_type="code" colab={}
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
#Some helper functions for plotting and drawing lines
def plot_points(X, y):
admitted = X[np.argwhere(y==1)]
rejected = X[np.argwhere(y==0)]
plt.scatter([s[0][0] for s in rejected], [s[0][1] for s in rejected], s = 25, color = 'blue', edgecolor = 'k')
plt.scatter([s[0][0] for s in admitted], [s[0][1] for s in admitted], s = 25, color = 'red', edgecolor = 'k')
def display(m, b, color='g--'):
plt.xlim(-0.05,1.05)
plt.ylim(-0.05,1.05)
x = np.arange(-10, 10, 0.1)
plt.plot(x, m*x+b, color)
# + id="V92OhGojZMQk" colab_type="code" colab={}
# Activation (sigmoid) function
def sigmoid(x):
return 1 / (1 + np.exp(-x))
def output_formula(features, weights, bias):
return sigmoid(np.dot(features, weights) + bias)
def error_formula(y, output):
return - y*np.log(output) - (1 - y) * np.log(1-output)
def update_weights(x, y, weights, bias, learnrate):
output = output_formula(x, weights, bias)
d_error = y - output
weights += learnrate * d_error * x
bias += learnrate * d_error
return weights, bias
# + id="EW8Ns-vtZbeE" colab_type="code" colab={}
np.random.seed(44)
epochs = 100
learnrate = 0.01
def train(features, targets, epochs, learnrate, graph_lines=False):
errors = []
n_records, n_features = features.shape
last_loss = None
weights = np.random.normal(scale=1 / n_features**.5, size=n_features)
bias = 0
for e in range(epochs):
del_w = np.zeros(weights.shape)
for x, y in zip(features, targets):
output = output_formula(x, weights, bias)
error = error_formula(y, output)
weights, bias = update_weights(x, y, weights, bias, learnrate)
# Printing out the log-loss error on the training set
out = output_formula(features, weights, bias)
loss = np.mean(error_formula(targets, out))
errors.append(loss)
if e % (epochs / 10) == 0:
print("\n========== Epoch", e,"==========")
if last_loss and last_loss < loss:
print("Train loss: ", loss, " WARNING - Loss Increasing")
else:
print("Train loss: ", loss)
last_loss = loss
predictions = out > 0.5
accuracy = np.mean(predictions == targets)
print("Accuracy: ", accuracy)
if graph_lines and e % (epochs / 100) == 0:
display(-weights[0]/weights[1], -bias/weights[1])
# Plotting the solution boundary
plt.title("Solution boundary")
display(-weights[0]/weights[1], -bias/weights[1], 'black')
# Plotting the data
plot_points(features, targets)
plt.show()
# Plotting the error
plt.title("Error Plot")
plt.xlabel('Number of epochs')
plt.ylabel('Error')
plt.plot(errors)
plt.show()
# + id="_CBh6FKrZ2J5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 264} outputId="76f23361-a47b-4e55-d25a-8ad8a5d5e2eb"
data = pd.read_csv('data.csv', header=None)
X = np.array(data[[0,1]])
y = np.array(data[2])
plot_points(X,y)
plt.show()
# + id="aa9J9Q1jZovj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="2cceec55-8b6f-4b71-c8a2-09b52a33a47a"
train(X, y, epochs, learnrate, True)
| GradientDescent.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Regression of pH
from tpot import TPOTRegressor
import matplotlib.pyplot as plt
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn import metrics
import pandas as pd
import seaborn as sns
import numpy as np
import datetime
# data
acidoCEST_ML = pd.read_csv('acido_CEST_MRI_MegaBox_01_to_08_clean.csv')
acidoCEST_ML = acidoCEST_ML.drop(['Unnamed: 0','ApproT1(sec)','Temp','FILE','Conc(mM)'], axis = 1)
print(acidoCEST_ML.shape)
acidoCEST_ML.shape
acidoCEST_ML
# +
# sample for development
#acidoCEST_ML = acidoCEST_ML.sample(n=2000)
# -
# ## TPO training to get pipeline all features we can measure
# +
# %%time
X_train, X_test, y_train, y_test = train_test_split( acidoCEST_ML.drop('pH',axis=1)
, acidoCEST_ML.pH
, test_size=0.30, random_state=42)
tpot = TPOTRegressor( generations= 10, population_size=10, verbosity=2, n_jobs= 4 , cv = 3
, early_stop=3
, max_time_mins= 30
, template = 'StandardScaler-Transformer-Selector-Regressor'
, scoring = metrics.make_scorer(metrics.median_absolute_error,greater_is_better=False) )
tpot.fit(X_train,y_train)
date_time = datetime.datetime.now().strftime("%m_%d_%Y_%H:%M")
tpot.export('acidoCEST_ML_tpot_pH_regressor_'+date_time+'.py')
print('Score on test set: \n',tpot.score(X_test, y_test))
# +
sat_powers = X_test['SatPower(uT)'].unique().tolist(); sat_powers.sort()
sat_times = X_test['SatTime(ms)'].unique().tolist(); sat_times.sort()
E = pd.DataFrame(data = np.zeros( ( len(sat_times) , len(sat_powers) ) )
, columns=sat_powers
, index = sat_times )
for t in sat_times:
for p in sat_powers:
f = (X_test['SatPower(uT)'] == p) & (X_test['SatTime(ms)']== t)
yhat = tpot.predict(X_test[f])
error = metrics.mean_absolute_error(y_test[f], yhat)
E.loc[t,p] = error
# -
fig, ax = plt.subplots(dpi = 140)
sns.heatmap(E, cmap='inferno',ax = ax, vmin=.1, vmax=0.50,annot=True)
plt.xlabel('\n Sat Power (uT)')
plt.ylabel('Sat Time (msec) \n')
plt.title('Median Absolure Error in pH regression using all data \n')
# ## TPO training to get pipeline with `only` CEST data & `B1` and `B0`
# - note: You cannot compare the results with the model using the entire data if the models are different
# +
# %%time
drop_cols = ['ExpT1(ms)', 'ExpT2(ms)','ExpB0(Hz)', 'SatPower(uT)', 'SatTime(ms)']
tpot2 = TPOTRegressor( generations= 10, population_size=10, verbosity=2, n_jobs= 4 , cv = 3
, early_stop=3
, max_time_mins= 30
, template = 'StandardScaler-Transformer-Selector-Regressor'
, scoring = metrics.make_scorer(metrics.median_absolute_error,greater_is_better=False) )
tpot2.fit(X_train.drop(drop_cols,axis=1),y_train)
date_time = datetime.datetime.now().strftime("%m_%d_%Y_%H:%M")
tpot2.export('acidoCEST_ML_tpot_pH_regressor_CEST_only'+date_time+'.py')
print('Score on test set: \n',tpot2.score(X_test.drop(drop_cols,axis=1), y_test))
# +
sat_powers = X_test['SatPower(uT)'].unique().tolist(); sat_powers.sort()
sat_times = X_test['SatTime(ms)'].unique().tolist(); sat_times.sort()
E2 = pd.DataFrame(data = np.zeros( ( len(sat_times) , len(sat_powers) ) )
, columns=sat_powers
, index = sat_times )
for t in sat_times:
for p in sat_powers:
f = (X_test['SatPower(uT)'] == p) & (X_test['SatTime(ms)']== t)
yhat = tpot2.predict(X_test[f].drop(drop_cols,axis=1))
error = metrics.mean_absolute_error(y_test[f], yhat)
E2.loc[t,p] = error
fig, ax = plt.subplots(dpi = 140)
sns.heatmap(E2, cmap='inferno',ax = ax, vmin=.1, vmax=0.50,annot=True)
plt.xlabel('\n Sat Power (uT)')
plt.ylabel('Sat Time (msec) \n')
plt.title('Median Absolure Error in pH regression using only CEST data, B0, and B1 \n')
# -
# ## TPO training to get pipeline with `only` CEST data & `T1` and `T2`
# - note: You cannot compare the results with the model using the entire data if the models are different
# +
# %%time
drop_cols = ['ExpB1(percent)', 'ExpB0(ppm)', 'ExpB0(Hz)','SatPower(uT)', 'SatTime(ms)']
tpot3 = TPOTRegressor( generations= 10, population_size=10, verbosity=2, n_jobs= 4 , cv = 3
, early_stop=3
, max_time_mins= 30
, template = 'StandardScaler-Transformer-Selector-Regressor'
, scoring = metrics.make_scorer(metrics.median_absolute_error,greater_is_better=False) )
tpot3.fit(X_train.drop(drop_cols,axis=1),y_train)
print('Score on test set: \n',tpot3.score(X_test.drop(drop_cols,axis=1), y_test))
# +
sat_powers = X_test['SatPower(uT)'].unique().tolist(); sat_powers.sort()
sat_times = X_test['SatTime(ms)'].unique().tolist(); sat_times.sort()
E3 = pd.DataFrame(data = np.zeros( ( len(sat_times) , len(sat_powers) ) )
, columns=sat_powers
, index = sat_times )
for t in sat_times:
for p in sat_powers:
f = (X_test['SatPower(uT)'] == p) & (X_test['SatTime(ms)']== t)
yhat = tpot3.predict(X_test[f].drop(drop_cols,axis=1))
error = metrics.mean_absolute_error(y_test[f], yhat)
E3.loc[t,p] = error
fig, ax = plt.subplots(dpi = 140)
sns.heatmap(E3, cmap='inferno',ax = ax, vmin=.1, vmax=0.50,annot=True)
plt.xlabel('\n Sat Power (uT)')
plt.ylabel('Sat Time (msec) \n')
plt.title('Median Absolure Error in pH regression using only CEST , T1, and T2 data')
# -
# ## TPO training to get pipeline with `only` CEST data , `sat time`, and `sat power`
# - note: You cannot compare the results with the model using the entire data if the models are different
# +
# %%time
drop_cols = ['ExpT1(ms)', 'ExpT2(ms)', 'ExpB1(percent)', 'ExpB0(ppm)', 'ExpB0(Hz)']
tpot4 = TPOTRegressor( generations= 10, population_size=10, verbosity=2, n_jobs= 4 , cv = 3
, early_stop=3
, max_time_mins= 30
, template = 'StandardScaler-Transformer-Selector-Regressor'
, scoring = metrics.make_scorer(metrics.median_absolute_error,greater_is_better=False) )
tpot4.fit(X_train.drop(drop_cols,axis=1),y_train)
print('Score on test set: \n',tpot4.score(X_test.drop(drop_cols,axis=1), y_test))
# +
sat_powers = X_test['SatPower(uT)'].unique().tolist(); sat_powers.sort()
sat_times = X_test['SatTime(ms)'].unique().tolist(); sat_times.sort()
E4 = pd.DataFrame(data = np.zeros( ( len(sat_times) , len(sat_powers) ) )
, columns=sat_powers
, index = sat_times )
for t in sat_times:
for p in sat_powers:
f = (X_test['SatPower(uT)'] == p) & (X_test['SatTime(ms)']== t)
yhat = tpot4.predict(X_test[f].drop(drop_cols,axis=1))
error = metrics.mean_absolute_error(y_test[f], yhat)
E4.loc[t,p] = error
fig, ax = plt.subplots(dpi = 140)
sns.heatmap(E4, cmap='inferno',ax = ax, vmin=.1, vmax=0.50,annot=True)
plt.xlabel('\n Sat Power (uT)')
plt.ylabel('Sat Time (msec) \n')
plt.title('Median Absolure Error in pH regression using only CEST, sat time, and sat power')
# -
| 1-TPOT regresor_B_and_T_exploration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# In this module, we make the Markov chain approximation for the Markovian inflow energy $X_t$. The generator function obtained in TS.py is used to train the Markov chain.
import pandas
import numpy
from msppy.utils.plot import fan_plot
from msppy.discretize import Markovian
import matplotlib.pyplot as plt
gamma = numpy.array(pandas.read_csv(
"./data/gamma.csv",
names=[0,1,2,3],
index_col=0,
skiprows=1,
))
sigma = [
numpy.array(pandas.read_csv(
"./data/sigma_{}.csv".format(i),
names=[0,1,2,3],
index_col=0,
skiprows=1,
)) for i in range(12)
]
exp_mu = numpy.array(pandas.read_csv(
"./data/exp_mu.csv",
names=[0,1,2,3],
index_col=0,
skiprows=1,
))
inflow_initial = numpy.array([41248.7153,7386.860854,10124.56146,6123.808537])
T = 12
def generator(random_state,size):
inflow = numpy.empty([size,T,4])
inflow[:,0,:] = inflow_initial[numpy.newaxis:,]
for t in range(1,T):
noise = numpy.exp(random_state.multivariate_normal(mean=[0]*4, cov=sigma[t%12],size=size))
inflow[:,t,:] = noise * (
(1-gamma[t%12]) * exp_mu[t%12]
+ gamma[t%12] * exp_mu[t%12]/exp_mu[(t-1)%12] * inflow[:,t-1,:]
)
return inflow
# Use robust stochastic approximation to iteratively train a non-homogenuous four-dimensional Markov chain with one initial Markov states and one hundred Markov states from stage two on. We make 1000 iterations to train the Markov states and then 10000 iterations to train the transition matrix.
# Refer to:
#
markovian = Markovian(
f=generator,
T=T,
n_Markov_states=[1]+[12]*(T-1),
n_sample_paths=1000)
Markov_states, transition_matrix = markovian.SA()
s = generator(numpy.random,100)
pandas.DataFrame(Markov_states[1]).head()
markovian.Markov_states[1]
pandas.DataFrame(Markov_states[1]).head()
pandas.DataFrame(transition_matrix[1])
# Let us see how do the returned markov_states and transition_matrix look like
# stage 0: The initial four-dimensional inflow
Markov_states[0]
# stage 1: The trained 100 Markov states. Each column is a Markov state.
pandas.DataFrame(Markov_states[1]).head()
# Stage 0: transition matrix always begins with [[1]] since the first stage is always
# assumed to be deterministic.
transition_matrix[0]
# stage 1: transition matrix between the initial single Markov state
# and the 100 Markov states in the second stage, so it is 1 by 100.
pandas.DataFrame(transition_matrix[1])
# stage 2: transition matrix between the 100 Markov states in the second stage
# and the 100 Markov states in the third stage, so it is 100 by 100.
pandas.DataFrame(transition_matrix[2]).head()
# Use the trained Markov state space and transition matrix to simulate inflow data. It is clear that the fan plot of the simulated sample path is very similar to that of the historical data (see TS.py).
sim = markovian.simulate(100)
fig = plt.figure(figsize=(10,5))
ax = [None] * 4
for i in range(4):
ax[i] = plt.subplot(221+i)
fan_plot(pandas.DataFrame(sim[:,:,i]),ax[i])
# Let us also try the SAA, SA and RSA approaches to make MC approximation.
markovian = Markovian(
f=generator,
T=T,
n_Markov_states=[1]+[100]*(T-1),
n_sample_paths=10000,
)
Markov_states, transition_matrix = markovian.SAA()
fig = plt.figure(figsize=(10,5))
sim = markovian.simulate(100)
ax = [None]*4
for i in range(4):
ax[i] = plt.subplot(221+i)
fan_plot(pandas.DataFrame(sim[:,:,i]),ax[i])
markovian = Markovian(
f=generator,
T=T,
n_Markov_states=[1]+[100]*(T-1),
n_sample_paths=10000,
)
Markov_states, transition_matrix = markovian.SA()
fig = plt.figure(figsize=(10,5))
sim = markovian.simulate(100)
ax = [None]*4
for i in range(4):
ax[i] = plt.subplot(221+i)
fan_plot(pandas.DataFrame(sim[:,:,i]),ax[i])
markovian = Markovian(
f=generator,
T=T,
n_Markov_states=[1]+[100]*(T-1),
n_sample_paths=10000,
)
Markov_states, transition_matrix = markovian.RSA()
fig = plt.figure(figsize=(10,5))
sim = markovian.simulate(100)
ax = [None]*4
for i in range(4):
ax[i] = plt.subplot(221+i)
fan_plot(pandas.DataFrame(sim[:,:,i]),ax[i])
| tutorials/hydro_thermal/Markov_chain_approximation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# %%writefile myfile.txt
Hello there, this is a text file
this is the second line
this is the third line
myfile = open('myfile.txt')
myfile = open('whoops_wrong.txt')
myfile = open('myfile.txt')
myfile.read()
myfile.read()
myfile.seek(0)
myfile.read()
myfile.seek(0)
contents = myfile.read()
contents
myfile.seek(0)
myfile.readlines()
myfile.close()
with open('myfile.txt') as my_new_file:
contents = my_new_file.read()
contents
with open('myfile.txt', mode = 'r') as myfile:
contents = myfile.read()
contents
# %%writefile my_new_file.txt
ONE ON FIRST
TWO ON SECOND
THREE ON THIRD
with open('my_new_file.txt', mode='r') as f:
print(f.read())
with open('my_new_file.txt', mode='a') as f:
f.write('FOUR ON FOURTH')
with open('my_new_file.txt', mode='r') as f:
print(f.read())
with open('absgctaj.txt', mode='w') as f:
f.write('I CREATED THIS FILE')
with open('absgctaj.txt', mode='r') as f:
print(f.read())
| Section 3: Python Object and Data Structure Basics/Files.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="GNkzTFfynsmV" colab_type="code" colab={}
# !pip install tensorflow==2.0.0b1
# + id="56XEQOGknrAk" colab_type="code" colab={}
import tensorflow as tf
print(tf.__version__)
# + id="sLl52leVp5wU" colab_type="code" colab={}
import numpy as np
import matplotlib.pyplot as plt
def plot_series(time, series, format="-", start=0, end=None):
plt.plot(time[start:end], series[start:end], format)
plt.xlabel("Time")
plt.ylabel("Value")
plt.grid(True)
# + id="tP7oqUdkk0gY" colab_type="code" colab={}
# !wget --no-check-certificate \
# https://raw.githubusercontent.com/jbrownlee/Datasets/master/daily-min-temperatures.csv \
# -O /tmp/daily-min-temperatures.csv
# + id="NcG9r1eClbTh" colab_type="code" colab={}
import csv
time_step = []
temps = []
with open('/tmp/daily-min-temperatures.csv') as csvfile:
# YOUR CODE HERE. READ TEMPERATURES INTO TEMPS
# HAVE TIME STEPS BE A SIMPLE ARRAY OF 1, 2, 3, 4 etc
series = np.array(temps)
time = np.array(time_step)
plt.figure(figsize=(10, 6))
plot_series(time, series)
# + id="L92YRw_IpCFG" colab_type="code" colab={}
split_time = 2500
time_train = # YOUR CODE HERE
x_train = # YOUR CODE HERE
time_valid = # YOUR CODE HERE
x_valid = # YOUR CODE HERE
window_size = 30
batch_size = 32
shuffle_buffer_size = 1000
# + id="lJwUUZscnG38" colab_type="code" colab={}
def windowed_dataset(series, window_size, batch_size, shuffle_buffer):
# YOUR CODE HERE
# + id="4XwGrf-A_wF0" colab_type="code" colab={}
def model_forecast(model, series, window_size):
# YOUR CODE HERE
# + id="AclfYY3Mn6Ph" colab_type="code" outputId="dd1fef93-d819-4d56-df20-330169907e16" executionInfo={"status": "ok", "timestamp": 1563513375680, "user_tz": 420, "elapsed": 370419, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-wUzpekukCVw/AAAAAAAAAAI/AAAAAAAAAHw/pQPstOOJqqE/s64/photo.jpg", "userId": "17858265307580721507"}} colab={"base_uri": "https://localhost:8080/", "height": 1000}
tf.keras.backend.clear_session()
tf.random.set_seed(51)
np.random.seed(51)
window_size = 64
batch_size = 256
train_set = windowed_dataset(x_train, window_size, batch_size, shuffle_buffer_size)
print(train_set)
print(x_train.shape)
model = tf.keras.models.Sequential([
# YOUR CODE HERE
])
lr_schedule = tf.keras.callbacks.LearningRateScheduler(
lambda epoch: 1e-8 * 10**(epoch / 20))
optimizer = tf.keras.optimizers.SGD(lr=1e-8, momentum=0.9)
model.compile(loss=tf.keras.losses.Huber(),
optimizer=optimizer,
metrics=["mae"])
history = model.fit(train_set, epochs=100, callbacks=[lr_schedule])
# + id="vVcKmg7Q_7rD" colab_type="code" outputId="5e9b8029-e996-4a2b-e016-666c69865b11" executionInfo={"status": "ok", "timestamp": 1563513433981, "user_tz": 420, "elapsed": 871, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-wUzpekukCVw/AAAAAAAAAAI/AAAAAAAAAHw/pQPstOOJqqE/s64/photo.jpg", "userId": "17858265307580721507"}} colab={"base_uri": "https://localhost:8080/", "height": 290}
plt.semilogx(history.history["lr"], history.history["loss"])
plt.axis([1e-8, 1e-4, 0, 60])
# + id="QsksvkcXAAgq" colab_type="code" outputId="70263fd4-3c3a-4e93-a451-ee942131e0d4" executionInfo={"status": "ok", "timestamp": 1563514553059, "user_tz": 420, "elapsed": 508595, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-wUzpekukCVw/AAAAAAAAAAI/AAAAAAAAAHw/pQPstOOJqqE/s64/photo.jpg", "userId": "17858265307580721507"}} colab={"base_uri": "https://localhost:8080/", "height": 1000}
tf.keras.backend.clear_session()
tf.random.set_seed(51)
np.random.seed(51)
train_set = windowed_dataset(x_train, window_size=60, batch_size=100, shuffle_buffer=shuffle_buffer_size)
model = tf.keras.models.Sequential([
# YOUR CODE HERE
])
optimizer = tf.keras.optimizers.SGD(lr=# YOUR CODE HERE, momentum=0.9)
model.compile(loss=tf.keras.losses.Huber(),
optimizer=optimizer,
metrics=["mae"])
history = model.fit(train_set,epochs=# YOUR CODE HERE)
# EXPECTED OUTPUT SHOULD SEE AN MAE OF <2 WITHIN ABOUT 30 EPOCHS
# + id="GaC6NNMRp0lb" colab_type="code" colab={}
rnn_forecast = model_forecast(model, series[..., np.newaxis], window_size)
rnn_forecast = rnn_forecast[split_time - window_size:-1, -1, 0]
# + colab_type="code" id="izy6wl2S9d-2" colab={}
plt.figure(figsize=(10, 6))
plot_series(time_valid, x_valid)
plot_series(time_valid, rnn_forecast)
# EXPECTED OUTPUT. PLOT SHOULD SHOW PROJECTIONS FOLLOWING ORIGINAL DATA CLOSELY
# + id="13XrorC5wQoE" colab_type="code" outputId="7f5bda4a-160c-4c0f-e511-a5a1b6945d35" executionInfo={"status": "ok", "timestamp": 1563515289044, "user_tz": 420, "elapsed": 301, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-wUzpekukCVw/AAAAAAAAAAI/AAAAAAAAAHw/pQPstOOJqqE/s64/photo.jpg", "userId": "17858265307580721507"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
tf.keras.metrics.mean_absolute_error(x_valid, rnn_forecast).numpy()
# EXPECTED OUTPUT MAE < 2 -- I GOT 1.789626
# + id="AOVzQXxCwkzP" colab_type="code" outputId="3b3fd11c-f9b3-4cbd-e32d-2d7d22b6ff7f" executionInfo={"status": "ok", "timestamp": 1563515322455, "user_tz": 420, "elapsed": 284, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-wUzpekukCVw/AAAAAAAAAAI/AAAAAAAAAHw/pQPstOOJqqE/s64/photo.jpg", "userId": "17858265307580721507"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
print(rnn_forecast)
# EXPECTED OUTPUT -- ARRAY OF VALUES IN THE LOW TEENS
| deep-learning/Tensorflow-2.x/Browser-Based-Models/TensorFlow In Practice/Course 4 - S+P/S+P Week 4 Exercise Question.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="nOzdBDxUdCuw" executionInfo={"status": "ok", "timestamp": 1626194516132, "user_tz": -360, "elapsed": 34347, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}} outputId="c617ef12-17d9-48ab-e5e5-55610164d555"
from google.colab import drive
drive.mount('/content/drive', force_remount=True)
# + [markdown] id="2ZpyR3xc2Ol3"
# #Load Libraries:
# + id="ki_Fq2_5edIl" executionInfo={"status": "ok", "timestamp": 1626194518476, "user_tz": -360, "elapsed": 2358, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://<KEY>", "userId": "07063771605322056910"}}
import tensorflow as tf
from tensorflow.keras.datasets import mnist
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Dropout, Flatten
from tensorflow.keras.layers import Conv2D, MaxPooling2D
from tensorflow.keras import backend as K
from tensorflow.keras import activations
from tensorflow.keras.layers import *
from tensorflow.keras.models import Model, load_model
from tensorflow.keras import models
from tensorflow.keras import layers
import cv2
import numpy as np
from tqdm import tqdm
import math
import os
import matplotlib.pyplot as plt
# + id="Nc1Kih9dedY_" executionInfo={"status": "ok", "timestamp": 1626194518482, "user_tz": -360, "elapsed": 16, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}}
# + [markdown] id="KMMyHBOZ19x0"
# #Load Model:
# + id="-_8AF-yhdOZ3" executionInfo={"status": "ok", "timestamp": 1626194518483, "user_tz": -360, "elapsed": 15, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}}
work_dir = "drive/My Drive/Plant_Leaf_MalayaKew_MK_Dataset/Records/"
# + id="EUOK04obeQVm" executionInfo={"status": "ok", "timestamp": 1626194518484, "user_tz": -360, "elapsed": 14, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}}
checkpointer_name = "weights.MK.D2.TTV.rgb.256p.DataAug4.DataFlow.pad0.TL.3D.DenseNet201.wInit.imagenet.TrainableAfter.allDefault.Dense.1024.1024.2048.actF.elu.opt.Adam.drop.0.5.batch16.Flatten.l2.0.001.run_1.hdf5"
# + colab={"base_uri": "https://localhost:8080/"} id="ysU6atkXdOcd" executionInfo={"status": "ok", "timestamp": 1626194545637, "user_tz": -360, "elapsed": 27164, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}} outputId="2d28dfaa-5d6a-4903-aec3-3d36193ecfd3"
model_loaded = load_model(work_dir+checkpointer_name)
print("Loaded "+work_dir+checkpointer_name+".")
# + colab={"base_uri": "https://localhost:8080/"} id="tHNURUPWdOfJ" executionInfo={"status": "ok", "timestamp": 1626194546982, "user_tz": -360, "elapsed": 1349, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}} outputId="77ea4aa6-f237-4444-ff6a-e8a4216640c5"
model_loaded.summary()
# + id="ViXRmtqjdOhz" executionInfo={"status": "ok", "timestamp": 1626194547000, "user_tz": -360, "elapsed": 31, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}}
# + [markdown] id="8hbpPCCO4Jx1"
# #Model Layers:
# + colab={"base_uri": "https://localhost:8080/"} id="wcjgpg8o4JPU" executionInfo={"status": "ok", "timestamp": 1626194547882, "user_tz": -360, "elapsed": 907, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}} outputId="83d0f8e4-2a5f-4058-c7c3-17173bef9e43"
layer_names = [] # conv4_block48_2_conv, conv3_block12_2_conv
for layer in model_loaded.layers:
layer_names.append(layer.name)
print(layer_names)
# + colab={"base_uri": "https://localhost:8080/"} id="3ATZmTBE4lI_" executionInfo={"status": "ok", "timestamp": 1626194547884, "user_tz": -360, "elapsed": 38, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}} outputId="e0c9a10c-f963-49bb-e42b-161345e4337a"
layer_no = -9
print(f"layer_names[{layer_no}] = {layer_names[layer_no]}")
# + id="1FwONIc74ZLa" executionInfo={"status": "ok", "timestamp": 1626194547885, "user_tz": -360, "elapsed": 37, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}}
# + [markdown] id="95hQ7EFG1du9"
# #By Loading Entire Test at Once:
# + id="sZetvikYywKT" colab={"base_uri": "https://localhost:8080/", "height": 0} executionInfo={"status": "ok", "timestamp": 1626194547888, "user_tz": -360, "elapsed": 38, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}} outputId="47f0b55c-2ea9-4352-928f-33ab9eea7060"
'''
input_path = "drive/My Drive/Plant_Leaf_MalayaKew_MK_Dataset/"
filename = "D2_Plant_Leaf_MalayaKew_MK_impl_1_Original_RGB_test_X.pkl.npy"
#'''
# + id="lLXSPXtMzZcQ" executionInfo={"status": "ok", "timestamp": 1626194547895, "user_tz": -360, "elapsed": 40, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}}
#input_test = np.load(f"{input_path}{filename}", allow_pickle=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="_W5hkjTzzeWx" executionInfo={"status": "ok", "timestamp": 1626194547897, "user_tz": -360, "elapsed": 40, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}} outputId="5543bb6a-8cb7-4ab2-f2e7-4e94cbab0e1a"
'''
print(f"input_test.shape = {input_test.shape}")
#'''
# + id="klW0J1i7ytUg" colab={"base_uri": "https://localhost:8080/", "height": 0} executionInfo={"status": "ok", "timestamp": 1626194547898, "user_tz": -360, "elapsed": 39, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}} outputId="2a43262c-b93b-41fe-b7b8-7c698ffbf10c"
'''
layer_outputs = [layer.output for layer in model_loaded.layers]
activation_model = models.Model(inputs=model_loaded.input, outputs=layer_outputs)
activations = activation_model.predict(input_test)
#'''
# + id="PXhv6uNf4Kqy" executionInfo={"status": "ok", "timestamp": 1626194547899, "user_tz": -360, "elapsed": 38, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}}
# + [markdown] id="W648c3vC1pIL"
# #By Loading Single at a Time:
# + colab={"base_uri": "https://localhost:8080/"} id="3lXRE2oAz-FK" executionInfo={"status": "ok", "timestamp": 1626194547901, "user_tz": -360, "elapsed": 39, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://<KEY>", "userId": "07063771605322056910"}} outputId="a94396c8-0fc0-4c73-8186-3e0a4d41a1a9"
root_path = "drive/My Drive/Plant_Leaf_MalayaKew_MK_Dataset/MK/D2/test_patch/"
#num_classes = 44
#list_classes = [f"Class{i+1}" for i in range(num_classes)]
list_classes = [f"Class{i}" for i in [1,11,22,33,44]]
list_input_path = []
for class_name in list_classes:
list_input_path.append(f"{root_path}{class_name}/")
print(f"len(list_input_path) = {len(list_input_path)}")
# + colab={"base_uri": "https://localhost:8080/", "height": 36} id="Ov1bmBMav-Wg" executionInfo={"status": "ok", "timestamp": 1626194550414, "user_tz": -360, "elapsed": 2551, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}} outputId="b5ada165-183e-4cbf-e4da-031bb9c7a092"
os.listdir(list_input_path[0])[0]
# + colab={"base_uri": "https://localhost:8080/"} id="lbhQGHoKvq2p" executionInfo={"status": "ok", "timestamp": 1626194555034, "user_tz": -360, "elapsed": 4625, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}} outputId="55e4ba40-8a5a-4684-c38b-512b356e25be"
list_full_paths = []
choose_different_index = 0
for input_path in list_input_path:
filename = os.listdir(input_path)[choose_different_index]
choose_different_index += 15
list_full_paths.append(f"{input_path}{filename}")
print(f"len(list_full_paths) = {len(list_full_paths)}")
# + colab={"base_uri": "https://localhost:8080/"} id="Td7X_qteAyLq" executionInfo={"status": "ok", "timestamp": 1626194555035, "user_tz": -360, "elapsed": 29, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}} outputId="05febc10-c747-4624-e68a-4dcd5ecc11e6"
list_full_paths
# + colab={"base_uri": "https://localhost:8080/", "height": 54} id="tmeR_0Lcz-H7" executionInfo={"status": "ok", "timestamp": 1626194555036, "user_tz": -360, "elapsed": 20, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}} outputId="950b57f3-6480-497e-b64d-19bb7c2d4d6d"
'''
filename = "Class44(8)R315_00277.jpg"
test_image = cv2.imread(f"{input_path}{filename}")
print(f"test_image.shape = {test_image.shape}")
input_test = np.expand_dims(test_image, 0)
print(f"input_test.shape = {input_test.shape}")
#'''
# + colab={"base_uri": "https://localhost:8080/"} id="lsi51g-Bz-K0" executionInfo={"status": "ok", "timestamp": 1626194557027, "user_tz": -360, "elapsed": 2001, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}} outputId="c08a70f4-4194-4712-dd27-4cff403ec43e"
list_test_images = []
for file_full_path in list_full_paths:
test_image = cv2.imread(file_full_path)
print(f"file_full_path: {file_full_path}")
list_test_images.append(test_image)
np_test_images = np.array(list_test_images)
print(f"np_test_images.shape = {np_test_images.shape}")
# + [markdown] id="l6Rwo5t8xorx"
# #Get Layer Activation Outputs:
# + id="zvSYJVJW3Wk9" executionInfo={"status": "ok", "timestamp": 1626194557029, "user_tz": -360, "elapsed": 14, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}}
layer_outputs = [layer.output for layer in model_loaded.layers]
activation_model = models.Model(inputs=model_loaded.input, outputs=layer_outputs)
#activations = activation_model.predict(input_test)
# + colab={"base_uri": "https://localhost:8080/"} id="wF_6m3ltfqFo" executionInfo={"status": "ok", "timestamp": 1626194590397, "user_tz": -360, "elapsed": 33381, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}} outputId="2c47f1bd-33d4-4e7b-9fa7-abad5454d201"
list_activations = []
for test_image in tqdm(np_test_images):
activations = activation_model.predict(np.array([test_image]))
list_activations.append(activations)
print(f"\nlen(list_activations) = {len(list_activations)}")
# + id="CDp35uWo2eTS" executionInfo={"status": "ok", "timestamp": 1626194590398, "user_tz": -360, "elapsed": 34, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}}
# + [markdown] id="6bjLJKIciHZE"
# ##Visualize:
# + colab={"base_uri": "https://localhost:8080/"} id="Yymbnrsj3flS" executionInfo={"status": "ok", "timestamp": 1626194590399, "user_tz": -360, "elapsed": 33, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}} outputId="7fea9658-6d77-4de7-94be-e75dec0abf25"
'''
input_1(256,256,3), conv1/relu(128,128,64), pool2_relu(64,64,256), pool3_relu(32,32,512), pool4_relu(16,16,1792), relu(8,8,1920)
'''
#target_layer_name = "conv3_block12_concat"
list_target_layer_names = ['input_1', 'conv1/relu', 'pool2_relu', 'pool3_relu', 'pool4_relu', 'relu']
list_layer_indices = []
for target_layer_name in list_target_layer_names:
for target_layer_index in range(len(layer_names)):
if layer_names[target_layer_index]==target_layer_name:
#layer_no = target_layer_index
list_layer_indices.append(target_layer_index)
#print(f"layer_names[{layer_no}] = {layer_names[layer_no]}")
print(f"list_layer_indices = {list_layer_indices}")
# + colab={"base_uri": "https://localhost:8080/"} id="nXvvBns6aPX5" executionInfo={"status": "ok", "timestamp": 1626194590401, "user_tz": -360, "elapsed": 31, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}} outputId="6dc4a4a7-030d-4706-bf0e-9ed160167ee8"
for activations in list_activations:
print(len(activations))
# + colab={"base_uri": "https://localhost:8080/", "height": 54} id="pSpNzilQ46Mo" executionInfo={"status": "ok", "timestamp": 1626194590402, "user_tz": -360, "elapsed": 25, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}} outputId="06e93821-b8f4-4516-e6a0-99c148fe2520"
'''
current_layer = activations[layer_no]
num_neurons = current_layer.shape[1:][-1]
print(f"current_layer.shape = {current_layer.shape}")
print(f"image_dimension = {current_layer.shape[1:][:-1]}")
print(f"num_neurons = {num_neurons}")
#'''
# + colab={"base_uri": "https://localhost:8080/"} id="LzJO_Wgcke9Y" executionInfo={"status": "ok", "timestamp": 1626194590403, "user_tz": -360, "elapsed": 23, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}} outputId="528db7b1-bb7e-466f-b058-de682e1fa865"
list_all_activations_layers = []
list_all_num_neurons = []
# list_all_activations_layers -> list_activations_layers -> current_layer -> activations[layer_no]
for activations in list_activations:
list_activations_layers = []
list_neurons = []
for layer_no in list_layer_indices:
current_layer = activations[layer_no]
#print(f"current_layer.shape = {current_layer.shape}")
list_activations_layers.append(current_layer)
#list_current_layers.append(current_layer)
list_neurons.append(current_layer.shape[1:][-1])
list_all_activations_layers.append(list_activations_layers)
list_all_num_neurons.append(list_neurons)
print(f"len(list_all_activations_layers) = {len(list_all_activations_layers)}")
print(f"len(list_all_activations_layers[0]) = {len(list_all_activations_layers[0])}")
print(f"list_all_activations_layers[0][0] = {list_all_activations_layers[0][0].shape}")
print(f"list_all_num_neurons = {list_all_num_neurons}")
print(f"list_all_num_neurons[0] = {list_all_num_neurons[0]}")
# + colab={"base_uri": "https://localhost:8080/"} id="_kTGmOSqrEOU" executionInfo={"status": "ok", "timestamp": 1626194590407, "user_tz": -360, "elapsed": 24, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}} outputId="c1595381-f8cd-4d1c-ab23-f176fcbd8f00"
print(f"list_all_activations_layers[0][0] = {list_all_activations_layers[0][0].shape}")
print(f"list_all_activations_layers[0][1] = {list_all_activations_layers[0][1].shape}")
print(f"list_all_activations_layers[0][2] = {list_all_activations_layers[0][2].shape}")
print(f"list_all_activations_layers[0][3] = {list_all_activations_layers[0][3].shape}")
print(f"list_all_activations_layers[0][4] = {list_all_activations_layers[0][4].shape}")
#print(f"list_all_activations_layers[0][5] = {list_all_activations_layers[0][5].shape}")
# + colab={"base_uri": "https://localhost:8080/", "height": 286} id="uPAhAkCgrdT-" executionInfo={"status": "ok", "timestamp": 1626194592179, "user_tz": -360, "elapsed": 1794, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}} outputId="04ab37dd-fb27-4085-bdf5-e6eb2131536f"
#'''
current_layer = list_all_activations_layers[0][1]
superimposed_activation_image = current_layer[0, :, :, 0]
for activation_image_index in range(1, current_layer.shape[-1]):
current_activation_image = current_layer[0, :, :, activation_image_index]
superimposed_activation_image = np.add(superimposed_activation_image, current_activation_image) # elementwise addition
plt.imshow(superimposed_activation_image, cmap='viridis')
#'''
# + colab={"base_uri": "https://localhost:8080/", "height": 286} id="0sSMu2wzsGVg" executionInfo={"status": "ok", "timestamp": 1626194592182, "user_tz": -360, "elapsed": 28, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}} outputId="ecf0ba81-5c08-4384-ef1d-4dcea749bed2"
#'''
current_layer = list_all_activations_layers[-1][1]
superimposed_activation_image = current_layer[0, :, :, 0]
for activation_image_index in range(1, current_layer.shape[-1]):
current_activation_image = current_layer[0, :, :, activation_image_index]
superimposed_activation_image = np.add(superimposed_activation_image, current_activation_image) # elementwise addition
plt.imshow(superimposed_activation_image, cmap='viridis')
#'''
# + id="VNU8xoUL51LV" executionInfo={"status": "ok", "timestamp": 1626194592183, "user_tz": -360, "elapsed": 20, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}}
#plt.matshow(current_layer[0, :, :, -1], cmap ='PiYG')
#plt.matshow(current_layer[0, :, :, -1], cmap ='viridis')
# + id="YxIBc2E_SzGa" colab={"base_uri": "https://localhost:8080/", "height": 91} executionInfo={"status": "ok", "timestamp": 1626194592184, "user_tz": -360, "elapsed": 20, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}} outputId="43d0845d-a952-4c9a-c6f0-94c9a2ee5123"
'''
superimposed_activation_image = current_layer[0, :, :, 0]
for activation_image_index in range(1, num_neurons):
current_activation_image = current_layer[0, :, :, activation_image_index]
#superimposed_activation_image = np.multiply(superimposed_activation_image, current_activation_image) # elementwise multiplication
superimposed_activation_image = np.add(superimposed_activation_image, current_activation_image) # elementwise addition
plt.imshow(superimposed_activation_image, cmap='viridis')
#'''
# + id="SeA7aw_HWoMN" colab={"base_uri": "https://localhost:8080/", "height": 72} executionInfo={"status": "ok", "timestamp": 1626194592185, "user_tz": -360, "elapsed": 19, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}} outputId="aa6169f2-dbad-4fc0-b51c-8672657c720c"
'''
superimposed_activation_image = current_layer[0, :, :, 0]
for activation_image_index in range(1, len(num_neurons)):
current_activation_image = current_layer[0, :, :, activation_image_index]
superimposed_activation_image = np.add(superimposed_activation_image, current_activation_image) # elementwise addition
plt.imshow(superimposed_activation_image, cmap='viridis')
#'''
# + colab={"base_uri": "https://localhost:8080/"} id="uydb14reCfwW" executionInfo={"status": "ok", "timestamp": 1626194593081, "user_tz": -360, "elapsed": 913, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}} outputId="815ca7cf-f5af-4982-cf00-db15516e8610"
#'''
# list_all_activations_layers -> list_activations_layers -> current_layer -> activations[layer_no]
#num_activation_per_test_image = len(list_target_layer_names)
list_all_superimposed_activation_image = []
for list_activations_layers_index in range(len(list_all_activations_layers)):
list_activations_layers = list_all_activations_layers[list_activations_layers_index]
list_current_num_neurons = list_all_num_neurons[list_activations_layers_index]
#print(f"list_activations_layers_index = {list_activations_layers_index}")
#print(f"list_all_num_neurons = {list_all_num_neurons}")
#print(f"list_current_num_neurons = {list_current_num_neurons}")
list_superimposed_activation_image = []
for activations_layer_index in range(len(list_activations_layers)):
activations_layers = list_activations_layers[activations_layer_index]
#print(f"activations_layers.shape = {activations_layers.shape}")
num_neurons = list_current_num_neurons[activations_layer_index]
superimposed_activation_image = activations_layers[0, :, :, 0]
for activation_image_index in range(1, num_neurons):
current_activation_image = activations_layers[0, :, :, activation_image_index]
superimposed_activation_image = np.add(superimposed_activation_image, current_activation_image) # elementwise addition
#print(f"superimposed_activation_image.shape = {superimposed_activation_image.shape}")
list_superimposed_activation_image.append(superimposed_activation_image)
#print(f"list_superimposed_activation_image[0].shape = {list_superimposed_activation_image[0].shape}")
list_all_superimposed_activation_image.append(list_superimposed_activation_image)
#print(f"list_all_superimposed_activation_image[0][0].shape = {list_all_superimposed_activation_image[0][0].shape}")
#plt.imshow(superimposed_activation_image, cmap='viridis')
print(f"len(list_all_superimposed_activation_image) = {len(list_all_superimposed_activation_image)}")
print(f"len(list_all_superimposed_activation_image[0]) = {len(list_all_superimposed_activation_image[0])}")
print(f"len(list_all_superimposed_activation_image[0][0]) = {len(list_all_superimposed_activation_image[0][0])}")
print(f"list_all_superimposed_activation_image[0][0].shape = {list_all_superimposed_activation_image[0][0].shape}")
#'''
# + colab={"base_uri": "https://localhost:8080/", "height": 146} id="sATiS-4tXC2S" executionInfo={"status": "ok", "timestamp": 1626194593085, "user_tz": -360, "elapsed": 66, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}} outputId="1dc7f8c8-e456-4fde-b663-3fae182c1fca"
'''
interpolation = cv2.INTER_LINEAR # INTER_LINEAR, INTER_CUBIC, INTER_NEAREST
# list_all_activations_layers -> list_activations_layers -> current_layer -> activations[layer_no]
#num_activation_per_test_image = len(list_target_layer_names)
list_all_superimposed_activation_image = []
for list_activations_layers_index in range(len(list_all_activations_layers)):
list_activations_layers = list_all_activations_layers[list_activations_layers_index]
list_current_num_neurons = list_all_num_neurons[list_activations_layers_index]
#print(f"list_activations_layers_index = {list_activations_layers_index}")
#print(f"list_all_num_neurons = {list_all_num_neurons}")
#print(f"list_current_num_neurons = {list_current_num_neurons}")
list_superimposed_activation_image = []
for activations_layer_index in range(len(list_activations_layers)):
activations_layers = list_activations_layers[activations_layer_index]
#print(f"activations_layers.shape = {activations_layers.shape}")
num_neurons = list_current_num_neurons[activations_layer_index]
superimposed_activation_image = activations_layers[0, :, :, 0]
superimposed_activation_image_resized = cv2.resize(superimposed_activation_image, (256,256), interpolation = interpolation)
for activation_image_index in range(1, num_neurons):
current_activation_image = activations_layers[0, :, :, activation_image_index]
#superimposed_activation_image = np.add(superimposed_activation_image, current_activation_image) # elementwise addition
current_activation_image_resized = cv2.resize(current_activation_image, (256,256), interpolation = interpolation)
superimposed_activation_image_resized = np.add(superimposed_activation_image_resized, current_activation_image_resized) # elementwise addition
#print(f"superimposed_activation_image.shape = {superimposed_activation_image.shape}")
#list_superimposed_activation_image.append(superimposed_activation_image)
list_superimposed_activation_image.append(superimposed_activation_image_resized)
#print(f"list_superimposed_activation_image[0].shape = {list_superimposed_activation_image[0].shape}")
list_all_superimposed_activation_image.append(list_superimposed_activation_image)
#print(f"list_all_superimposed_activation_image[0][0].shape = {list_all_superimposed_activation_image[0][0].shape}")
#plt.imshow(superimposed_activation_image, cmap='viridis')
print(f"len(list_all_superimposed_activation_image) = {len(list_all_superimposed_activation_image)}")
print(f"len(list_all_superimposed_activation_image[0]) = {len(list_all_superimposed_activation_image[0])}")
print(f"len(list_all_superimposed_activation_image[0][0]) = {len(list_all_superimposed_activation_image[0][0])}")
print(f"list_all_superimposed_activation_image[0][0].shape = {list_all_superimposed_activation_image[0][0].shape}")
print(f"list_all_superimposed_activation_image[0][-1].shape = {list_all_superimposed_activation_image[0][-1].shape}")
#'''
# + id="ox4o4lMddoii" executionInfo={"status": "ok", "timestamp": 1626194593092, "user_tz": -360, "elapsed": 63, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}}
# + colab={"base_uri": "https://localhost:8080/", "height": 146} id="V4yXgCD1ObB1" executionInfo={"status": "ok", "timestamp": 1626194593096, "user_tz": -360, "elapsed": 61, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}} outputId="435c0fbf-76b7-4eff-c68d-6956cd1f8e81"
'''
supported cmap values are: 'Accent', 'Accent_r', 'Blues', 'Blues_r', 'BrBG', 'BrBG_r', 'BuGn', 'BuGn_r', 'BuPu', 'BuPu_r', 'CMRmap', 'CMRmap_r',
'Dark2', 'Dark2_r', 'GnBu', 'GnBu_r', 'Greens', 'Greens_r', 'Greys', 'Greys_r', 'OrRd', 'OrRd_r', 'Oranges', 'Oranges_r', 'PRGn', 'PRGn_r',
'Paired', 'Paired_r', 'Pastel1', 'Pastel1_r', 'Pastel2', 'Pastel2_r', 'PiYG', 'PiYG_r', 'PuBu', 'PuBuGn', 'PuBuGn_r', 'PuBu_r', 'PuOr',
'PuOr_r', 'PuRd', 'PuRd_r', 'Purples', 'Purples_r', 'RdBu', 'RdBu_r', 'RdGy', 'RdGy_r', 'RdPu', 'RdPu_r', 'RdYlBu', 'RdYlBu_r', 'RdYlGn',
'RdYlGn_r', 'Reds', 'Reds_r', 'Set1', 'Set1_r', 'Set2', 'Set2_r', 'Set3', 'Set3_r', 'Spectral', 'Spectral_r', 'Wistia', 'Wistia_r', 'YlGn',
'YlGnBu', 'YlGnBu_r', 'YlGn_r', 'YlOrBr', 'YlOrBr_r', 'YlOrRd', 'YlOrRd_r', 'afmhot', 'afmhot_r', 'autumn', 'autumn_r', 'binary', 'binary_r',
'bone', 'bone_r', 'brg', 'brg_r', 'bwr', 'bwr_r', 'cividis', 'cividis_r', 'cool', 'cool_r', 'coolwarm', 'coolwarm_r', 'copper', 'copper_r',
'cubehelix', 'cubehelix_r', 'flag', 'flag_r', 'gist_earth', 'gist_earth_r', 'gist_gray', 'gist_gray_r', 'gist_heat', 'gist_heat_r', 'gist_ncar',
'gist_ncar_r', 'gist_rainbow', 'gist_rainbow_r', 'gist_stern', 'gist_stern_r', 'gist_yarg', 'gist_yarg_r', 'gnuplot', 'gnuplot2', 'gnuplot2_r',
'gnuplot_r', 'gray', 'gray_r', 'hot', 'hot_r', 'hsv', 'hsv_r', 'inferno', 'inferno_r', 'jet', 'jet_r', 'magma', 'magma_r', 'nipy_spectral',
'nipy_spectral_r', 'ocean', 'oc...
'''
# + colab={"base_uri": "https://localhost:8080/", "height": 682} id="F3zED9xEpjQM" executionInfo={"status": "ok", "timestamp": 1626194603182, "user_tz": -360, "elapsed": 10135, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}} outputId="73af4d11-2862-411a-e563-6ba3ce58a294"
sub_fig_num_rows = len(list_test_images)
sub_fig_num_cols = len(list_target_layer_names)
fig_heigth = 11
fig_width = 11
cmap = "Greens" # PuOr_r, Dark2, Dark2_r, RdBu, RdBu_r, coolwarm, viridis, PiYG, gray, binary, afmhot, PuBu, copper
fig, axes = plt.subplots(sub_fig_num_rows,sub_fig_num_cols, figsize=(fig_width,fig_heigth))
#plt.suptitle(f"Layer {str(layer_no+1)}: {layer_names[layer_no]} {str(current_layer.shape[1:])}", fontsize=20, y=1.1)
for i,ax in enumerate(axes.flat):
row = i//sub_fig_num_cols
col = i%sub_fig_num_cols
#print(f"i={i}; row={row}, col={col}")
#'''
ax.imshow(list_all_superimposed_activation_image[row][col], cmap=cmap)
#ax.imshow(list_all_superimposed_activation_image[row][col])
ax.set_xticks([])
ax.set_yticks([])
if col == 0:
ax.set_ylabel(f"{list_classes[row]}")
if row == 0:
#ax.set_xlabel(f"Layer {str(list_layer_indices[col])}") # , rotation=0, ha='right'
ax.set_xlabel(str(list_target_layer_names[col]))
#ax.set_xlabel(f"Layer {str(list_layer_indices[col])}: {str(list_target_layer_names[col])}") # , rotation=0, ha='right'
ax.xaxis.set_label_position('top')
ax.set_aspect('auto')
plt.subplots_adjust(wspace=0.02, hspace=0.05)
img_path = 'drive/My Drive/Visualizations/'+checkpointer_name[8:-5]+'.png'
plt.savefig(img_path, dpi=600)
plt.show()
print('img_path =', img_path)
#'''
# + colab={"base_uri": "https://localhost:8080/", "height": 283} id="ZKKjgDVNJ46o" executionInfo={"status": "ok", "timestamp": 1626194603184, "user_tz": -360, "elapsed": 34, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}} outputId="9ea90970-bf7d-427e-e9af-8455e3c4cca3"
# good cmap for this work: PuOr_r, Dark2_r, RdBu, RdBu_r, coolwarm, viridis, PiYG
'''
for activation_image_index in range(num_neurons):
plt.imshow(current_layer[0, :, :, activation_image_index], cmap='PiYG')
#'''
plt.imshow(superimposed_activation_image, cmap='gray')
# + id="212AvSgY39Dk" executionInfo={"status": "ok", "timestamp": 1626194603187, "user_tz": -360, "elapsed": 24, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}}
# + [markdown] id="yTVHjDQyxrYP"
# #Weight Visualization:
# + id="kCX4ucXMdOku" executionInfo={"status": "ok", "timestamp": 1626194603194, "user_tz": -360, "elapsed": 29, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}}
layer_outputs = [layer.output for layer in model_loaded.layers]
#activation_model = models.Model(inputs=model_loaded.input, outputs=layer_outputs)
#activations = activation_model.predict(input_test)
# + colab={"base_uri": "https://localhost:8080/"} id="dCOrNeGEdOsu" executionInfo={"status": "ok", "timestamp": 1626194604236, "user_tz": -360, "elapsed": 1069, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}} outputId="67532d6b-d799-444d-8e33-f3a9d7a98d43"
layer_configs = []
layer_weights = []
for layer in model_loaded.layers:
layer_configs.append(layer.get_config())
layer_weights.append(layer.get_weights())
print(f"len(layer_configs) = {len(layer_configs)}")
print(f"len(layer_weights) = {len(layer_weights)}")
# + colab={"base_uri": "https://localhost:8080/"} id="fw6huxYTdOvo" executionInfo={"status": "ok", "timestamp": 1626194604240, "user_tz": -360, "elapsed": 48, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}} outputId="d1b02cef-6cdd-4714-f58f-895e3859db67"
layer_configs[-9]
# + id="x1VI_tY6dOyc" executionInfo={"status": "ok", "timestamp": 1626194604242, "user_tz": -360, "elapsed": 47, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}}
layer_name = 'conv2_block1_1_conv' # conv5_block32_1_conv
model_weight = model_loaded.get_layer(layer_name).get_weights()[0]
#model_biases = model_loaded.get_layer(layer_name).get_weights()[1]
# + colab={"base_uri": "https://localhost:8080/"} id="F1FeqY8bdO1E" executionInfo={"status": "ok", "timestamp": 1626194604245, "user_tz": -360, "elapsed": 47, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}} outputId="2f1acd6f-b224-497c-ce0d-12e7e453a13a"
print(f"type(model_weight) = {type(model_weight)}")
print(f"model_weight.shape = {model_weight.shape}")
# + colab={"base_uri": "https://localhost:8080/"} id="iZxPM8ocdO37" executionInfo={"status": "ok", "timestamp": 1626194604247, "user_tz": -360, "elapsed": 47, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}} outputId="67a02489-7e73-459c-e4af-33d81f3cc923"
model_weight[0][0].shape
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="KICZImKDdO6T" executionInfo={"status": "ok", "timestamp": 1626194604249, "user_tz": -360, "elapsed": 46, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}} outputId="2d5509a5-106e-4c53-8490-3eb2c83b639e"
plt.matshow(model_weight[0, 0, :, :], cmap ='viridis')
# + id="m85ei0WAdO9L" executionInfo={"status": "ok", "timestamp": 1626194604249, "user_tz": -360, "elapsed": 38, "user": {"displayName": "ForOnline InternetUse", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgbMAzZA0UZsjrcX05klvyZp6cS9VKiHlpS3ggl=s64", "userId": "07063771605322056910"}}
| Layer_Activation_Visualization_from_Saved_Model_Malaya_Kew.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="_jQ1tEQCxwRx"
# ##### Copyright 2019 The TensorFlow Authors.
# + cellView="form" colab={} colab_type="code" id="V_sgB_5dx1f1"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="rF2x3qooyBTI"
# # 深度卷积生成对抗网络
# + [markdown] colab_type="text" id="0TD5ZrvEMbhZ"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://tensorflow.google.cn/tutorials/generative/dcgan">
# <img src="https://tensorflow.google.cn/images/tf_logo_32px.png" />
# 在 tensorFlow.google.cn 上查看</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/zh-cn/tutorials/generative/dcgan.ipynb">
# <img src="https://tensorflow.google.cn/images/colab_logo_32px.png" />
# 在 Google Colab 中运行</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/zh-cn/tutorials/generative/dcgan.ipynb">
# <img src="https://tensorflow.google.cn/images/GitHub-Mark-32px.png" />
# 在 GitHub 上查看源代码</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/docs-l10n/site/zh-cn/tutorials/generative/dcgan.ipynb"><img src="https://tensorflow.google.cn/images/download_logo_32px.png" />下载 notebook</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="M0gHG-LEgLZx"
# Note: 我们的 TensorFlow 社区翻译了这些文档。因为社区翻译是尽力而为, 所以无法保证它们是最准确的,并且反映了最新的
# [官方英文文档](https://tensorflow.google.cn/?hl=en)。如果您有改进此翻译的建议, 请提交 pull request 到
# [tensorflow/docs](https://github.com/tensorflow/docs) GitHub 仓库。要志愿地撰写或者审核译文,请加入
# [<EMAIL> Google Group](https://groups.google.com/a/tensorflow.org/forum/#!forum/docs-zh-cn)。
# + [markdown] colab_type="text" id="ITZuApL56Mny"
# 本教程演示了如何使用[深度卷积生成对抗网络](https://arxiv.org/pdf/1511.06434.pdf)(DCGAN)生成手写数字图片。该代码是使用 [Keras Sequential API](https://tensorflow.google.cn/guide/keras) 与 `tf.GradientTape` 训练循环编写的。
# + [markdown] colab_type="text" id="2MbKJY38Puy9"
# ## 什么是生成对抗网络?
#
# [生成对抗网络](https://arxiv.org/abs/1406.2661)(GANs)是当今计算机科学领域最有趣的想法之一。两个模型通过对抗过程同时训练。一个*生成器*(“艺术家”)学习创造看起来真实的图像,而*判别器*(“艺术评论家”)学习区分真假图像。
#
# 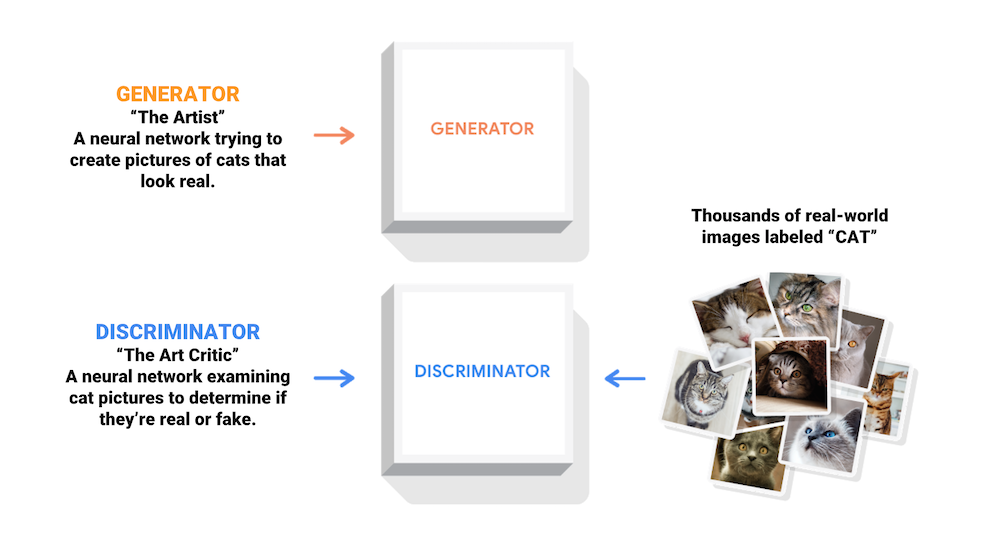
#
# 训练过程中,*生成器*在生成逼真图像方面逐渐变强,而*判别器*在辨别这些图像的能力上逐渐变强。当*判别器*不再能够区分真实图片和伪造图片时,训练过程达到平衡。
#
# 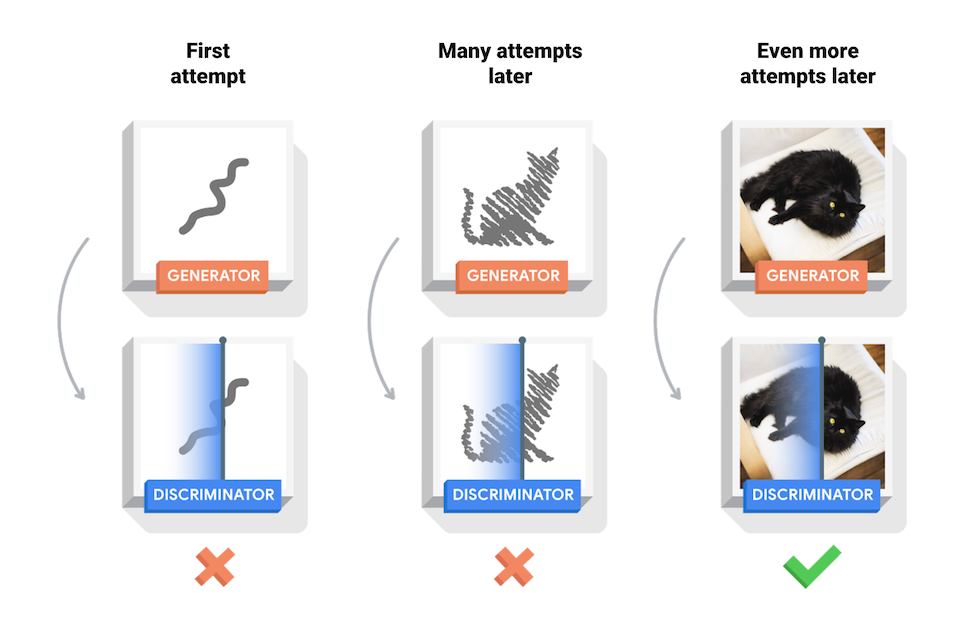
#
# 本笔记在 MNIST 数据集上演示了该过程。下方动画展示了当训练了 50 个epoch (全部数据集迭代50次) 时*生成器*所生成的一系列图片。图片从随机噪声开始,随着时间的推移越来越像手写数字。
#
# 
#
# 要了解关于 GANs 的更多信息,我们建议参阅 MIT的 [深度学习入门](http://introtodeeplearning.com/) 课程。
# + [markdown] colab_type="text" id="e1_Y75QXJS6h"
# ### Import TensorFlow and other libraries
# + colab={} colab_type="code" id="WZKbyU2-AiY-"
import tensorflow as tf
# + colab={} colab_type="code" id="57FFuKn4gLZ9"
tf.__version__
# + colab={} colab_type="code" id="YzTlj4YdCip_"
# 用于生成 GIF 图片
# !pip install imageio
# + colab={} colab_type="code" id="YfIk2es3hJEd"
import glob
import imageio
import matplotlib.pyplot as plt
import numpy as np
import os
import PIL
from tensorflow.keras import layers
import time
from IPython import display
# + [markdown] colab_type="text" id="iYn4MdZnKCey"
# ### 加载和准备数据集
#
# 您将使用 MNIST 数据集来训练生成器和判别器。生成器将生成类似于 MNIST 数据集的手写数字。
# + colab={} colab_type="code" id="a4fYMGxGhrna"
(train_images, train_labels), (_, _) = tf.keras.datasets.mnist.load_data()
# + colab={} colab_type="code" id="NFC2ghIdiZYE"
train_images = train_images.reshape(train_images.shape[0], 28, 28, 1).astype('float32')
train_images = (train_images - 127.5) / 127.5 # 将图片标准化到 [-1, 1] 区间内
# + colab={} colab_type="code" id="S4PIDhoDLbsZ"
BUFFER_SIZE = 60000
BATCH_SIZE = 256
# + colab={} colab_type="code" id="-yKCCQOoJ7cn"
# 批量化和打乱数据
train_dataset = tf.data.Dataset.from_tensor_slices(train_images).shuffle(BUFFER_SIZE).batch(BATCH_SIZE)
# + [markdown] colab_type="text" id="THY-sZMiQ4UV"
# ## 创建模型
#
# 生成器和判别器均使用 [Keras Sequential API](https://tensorflow.google.cn/guide/keras#sequential_model) 定义。
# + [markdown] colab_type="text" id="-tEyxE-GMC48"
# ### 生成器
#
# 生成器使用 `tf.keras.layers.Conv2DTranspose` (上采样)层来从种子(随机噪声)中产生图片。以一个使用该种子作为输入的 `Dense` 层开始,然后多次上采样直到达到所期望的 28x28x1 的图片尺寸。注意除了输出层使用 tanh 之外,其他每层均使用 `tf.keras.layers.LeakyReLU` 作为激活函数。
# + colab={} colab_type="code" id="6bpTcDqoLWjY"
def make_generator_model():
model = tf.keras.Sequential()
model.add(layers.Dense(7*7*256, use_bias=False, input_shape=(100,)))
model.add(layers.BatchNormalization())
model.add(layers.LeakyReLU())
model.add(layers.Reshape((7, 7, 256)))
assert model.output_shape == (None, 7, 7, 256) # 注意:batch size 没有限制
model.add(layers.Conv2DTranspose(128, (5, 5), strides=(1, 1), padding='same', use_bias=False))
assert model.output_shape == (None, 7, 7, 128)
model.add(layers.BatchNormalization())
model.add(layers.LeakyReLU())
model.add(layers.Conv2DTranspose(64, (5, 5), strides=(2, 2), padding='same', use_bias=False))
assert model.output_shape == (None, 14, 14, 64)
model.add(layers.BatchNormalization())
model.add(layers.LeakyReLU())
model.add(layers.Conv2DTranspose(1, (5, 5), strides=(2, 2), padding='same', use_bias=False, activation='tanh'))
assert model.output_shape == (None, 28, 28, 1)
return model
# + [markdown] colab_type="text" id="GyWgG09LCSJl"
# 使用(尚未训练的)生成器创建一张图片。
# + colab={} colab_type="code" id="o6VvUbMqgLaS"
generator = make_generator_model()
noise = tf.random.normal([1, 100])
generated_image = generator(noise, training=False)
plt.imshow(generated_image[0, :, :, 0], cmap='gray')
# + [markdown] colab_type="text" id="D0IKnaCtg6WE"
# ### 判别器
#
# 判别器是一个基于 CNN 的图片分类器。
# + colab={} colab_type="code" id="dw2tPLmk2pEP"
def make_discriminator_model():
model = tf.keras.Sequential()
model.add(layers.Conv2D(64, (5, 5), strides=(2, 2), padding='same',
input_shape=[28, 28, 1]))
model.add(layers.LeakyReLU())
model.add(layers.Dropout(0.3))
model.add(layers.Conv2D(128, (5, 5), strides=(2, 2), padding='same'))
model.add(layers.LeakyReLU())
model.add(layers.Dropout(0.3))
model.add(layers.Flatten())
model.add(layers.Dense(1))
return model
# + [markdown] colab_type="text" id="QhPneagzCaQv"
# 使用(尚未训练的)判别器来对图片的真伪进行判断。模型将被训练为为真实图片输出正值,为伪造图片输出负值。
# + colab={} colab_type="code" id="-nnSVbzhgLaX"
discriminator = make_discriminator_model()
decision = discriminator(generated_image)
print (decision)
# + [markdown] colab_type="text" id="0FMYgY_mPfTi"
# ## 定义损失函数和优化器
#
# 为两个模型定义损失函数和优化器。
# + colab={} colab_type="code" id="psQfmXxYKU3X"
# 该方法返回计算交叉熵损失的辅助函数
cross_entropy = tf.keras.losses.BinaryCrossentropy(from_logits=True)
# + [markdown] colab_type="text" id="PKY_iPSPNWoj"
# ### 判别器损失
#
# 该方法量化判别器从判断真伪图片的能力。它将判别器对真实图片的预测值与值全为 1 的数组进行对比,将判别器对伪造(生成的)图片的预测值与值全为 0 的数组进行对比。
# + colab={} colab_type="code" id="wkMNfBWlT-PV"
def discriminator_loss(real_output, fake_output):
real_loss = cross_entropy(tf.ones_like(real_output), real_output)
fake_loss = cross_entropy(tf.zeros_like(fake_output), fake_output)
total_loss = real_loss + fake_loss
return total_loss
# + [markdown] colab_type="text" id="Jd-3GCUEiKtv"
# ### 生成器损失
#
# 生成器损失量化其欺骗判别器的能力。直观来讲,如果生成器表现良好,判别器将会把伪造图片判断为真实图片(或 1)。这里我们将把判别器在生成图片上的判断结果与一个值全为 1 的数组进行对比。
# + colab={} colab_type="code" id="90BIcCKcDMxz"
def generator_loss(fake_output):
return cross_entropy(tf.ones_like(fake_output), fake_output)
# + [markdown] colab_type="text" id="MgIc7i0th_Iu"
# 由于我们需要分别训练两个网络,判别器和生成器的优化器是不同的。
# + colab={} colab_type="code" id="iWCn_PVdEJZ7"
generator_optimizer = tf.keras.optimizers.Adam(1e-4)
discriminator_optimizer = tf.keras.optimizers.Adam(1e-4)
# + [markdown] colab_type="text" id="mWtinsGDPJlV"
# ### 保存检查点
#
# 本笔记还演示了如何保存和恢复模型,这在长时间训练任务被中断的情况下比较有帮助。
# + colab={} colab_type="code" id="CA1w-7s2POEy"
checkpoint_dir = './training_checkpoints'
checkpoint_prefix = os.path.join(checkpoint_dir, "ckpt")
checkpoint = tf.train.Checkpoint(generator_optimizer=generator_optimizer,
discriminator_optimizer=discriminator_optimizer,
generator=generator,
discriminator=discriminator)
# + [markdown] colab_type="text" id="Rw1fkAczTQYh"
# ## 定义训练循环
#
# + colab={} colab_type="code" id="NS2GWywBbAWo"
EPOCHS = 50
noise_dim = 100
num_examples_to_generate = 16
# 我们将重复使用该种子(因此在动画 GIF 中更容易可视化进度)
seed = tf.random.normal([num_examples_to_generate, noise_dim])
# + [markdown] colab_type="text" id="jylSonrqSWfi"
# 训练循环在生成器接收到一个随机种子作为输入时开始。该种子用于生产一张图片。判别器随后被用于区分真实图片(选自训练集)和伪造图片(由生成器生成)。针对这里的每一个模型都计算损失函数,并且计算梯度用于更新生成器与判别器。
# + colab={} colab_type="code" id="3t5ibNo05jCB"
# 注意 `tf.function` 的使用
# 该注解使函数被“编译”
@tf.function
def train_step(images):
noise = tf.random.normal([BATCH_SIZE, noise_dim])
with tf.GradientTape() as gen_tape, tf.GradientTape() as disc_tape:
generated_images = generator(noise, training=True)
real_output = discriminator(images, training=True)
fake_output = discriminator(generated_images, training=True)
gen_loss = generator_loss(fake_output)
disc_loss = discriminator_loss(real_output, fake_output)
gradients_of_generator = gen_tape.gradient(gen_loss, generator.trainable_variables)
gradients_of_discriminator = disc_tape.gradient(disc_loss, discriminator.trainable_variables)
generator_optimizer.apply_gradients(zip(gradients_of_generator, generator.trainable_variables))
discriminator_optimizer.apply_gradients(zip(gradients_of_discriminator, discriminator.trainable_variables))
# + colab={} colab_type="code" id="2M7LmLtGEMQJ"
def train(dataset, epochs):
for epoch in range(epochs):
start = time.time()
for image_batch in dataset:
train_step(image_batch)
# 继续进行时为 GIF 生成图像
display.clear_output(wait=True)
generate_and_save_images(generator,
epoch + 1,
seed)
# 每 15 个 epoch 保存一次模型
if (epoch + 1) % 15 == 0:
checkpoint.save(file_prefix = checkpoint_prefix)
print ('Time for epoch {} is {} sec'.format(epoch + 1, time.time()-start))
# 最后一个 epoch 结束后生成图片
display.clear_output(wait=True)
generate_and_save_images(generator,
epochs,
seed)
# + [markdown] colab_type="text" id="2aFF7Hk3XdeW"
# **生成与保存图片**
#
# + colab={} colab_type="code" id="RmdVsmvhPxyy"
def generate_and_save_images(model, epoch, test_input):
# 注意 training` 设定为 False
# 因此,所有层都在推理模式下运行(batchnorm)。
predictions = model(test_input, training=False)
fig = plt.figure(figsize=(4,4))
for i in range(predictions.shape[0]):
plt.subplot(4, 4, i+1)
plt.imshow(predictions[i, :, :, 0] * 127.5 + 127.5, cmap='gray')
plt.axis('off')
plt.savefig('image_at_epoch_{:04d}.png'.format(epoch))
plt.show()
# + [markdown] colab_type="text" id="dZrd4CdjR-Fp"
# ## 训练模型
# 调用上面定义的 `train()` 方法来同时训练生成器和判别器。注意,训练 GANs 可能是棘手的。重要的是,生成器和判别器不能够互相压制对方(例如,他们以相似的学习率训练)。
#
# 在训练之初,生成的图片看起来像是随机噪声。随着训练过程的进行,生成的数字将越来越真实。在大概 50 个 epoch 之后,这些图片看起来像是 MNIST 数字。使用 Colab 中的默认设置可能需要大约 1 分钟每 epoch。
# + colab={} colab_type="code" id="Ly3UN0SLLY2l"
# %%time
train(train_dataset, EPOCHS)
# + [markdown] colab_type="text" id="rfM4YcPVPkNO"
# 恢复最新的检查点。
# + colab={} colab_type="code" id="SKeqh9GlgLa7"
checkpoint.restore(tf.train.latest_checkpoint(checkpoint_dir))
# + [markdown] colab_type="text" id="P4M_vIbUi7c0"
# ## 创建 GIF
#
# + colab={} colab_type="code" id="WfO5wCdclHGL"
# 使用 epoch 数生成单张图片
def display_image(epoch_no):
return PIL.Image.open('image_at_epoch_{:04d}.png'.format(epoch_no))
# + colab={} colab_type="code" id="TsUudzLCgLbA"
display_image(EPOCHS)
# + [markdown] colab_type="text" id="NywiH3nL8guF"
# 使用训练过程中生成的图片通过 `imageio` 生成动态 gif
# + colab={} colab_type="code" id="IGKQgENQ8lEI"
anim_file = 'dcgan.gif'
with imageio.get_writer(anim_file, mode='I') as writer:
filenames = glob.glob('image*.png')
filenames = sorted(filenames)
last = -1
for i,filename in enumerate(filenames):
frame = 2*(i**0.5)
if round(frame) > round(last):
last = frame
else:
continue
image = imageio.imread(filename)
writer.append_data(image)
image = imageio.imread(filename)
writer.append_data(image)
import IPython
if IPython.version_info > (6,2,0,''):
display.Image(filename=anim_file)
# + [markdown] colab_type="text" id="cGhC3-fMWSwl"
# 如果您正在使用 Colab,您可以通过如下代码下载动画:
# + colab={} colab_type="code" id="uV0yiKpzNP1b"
try:
from google.colab import files
except ImportError:
pass
else:
files.download(anim_file)
# + [markdown] colab_type="text" id="k6qC-SbjK0yW"
# ## 下一步
#
# + [markdown] colab_type="text" id="xjjkT9KAK6H7"
# 本教程展示了实现和训练 GAN 模型所需的全部必要代码。接下来,您可能想尝试其他数据集,例如大规模名人面部属性(CelebA)数据集 [在 Kaggle 上获取](https://www.kaggle.com/jessicali9530/celeba-dataset)。要了解更多关于 GANs 的信息,我们推荐参阅 [NIPS 2016 教程: 生成对抗网络](https://arxiv.org/abs/1701.00160)。
| site/zh-cn/tutorials/generative/dcgan.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: OSMNx Python3
# language: python
# name: osmnx_py3
# ---
# # Notebook 5: Identify Route Parish Intersections
# ## Introduction to Noteboook 5
# Why are route-parish intersections important? https://cw.fel.cvut.cz/b181/_media/courses/cg/lectures/09-intersect.pdf
#
#
# +
import osmnx as ox
ox.settings.log_console=False
ox.settings.use_cache=True
ox.settings.log_file=True
ox.settings.overpass_endpoint='https://overpass-api.de/api/interpreter'
# +
# %%time
import geopandas as gpd
if 'G_proj_u' not in globals():
# Load projected graph, G_proj
g_projected_nodes = gpd.read_feather('data/g_projected_nodes.feather')
g_projected_edges = gpd.read_feather('data/g_projected_edges.feather')
G_proj = ox.utils_graph.graph_from_gdfs(g_projected_nodes, g_projected_edges)
G_proj_u = ox.utils_graph.get_undirected(G_proj)
if 'G_u' not in globals():
# Load projected graph, G_proj
g_unprojected_nodes = gpd.read_feather('data/g_unprojected_nodes.feather')
g_unprojected_edges = gpd.read_feather('data/g_unprojected_edges.feather')
G = ox.utils_graph.graph_from_gdfs(g_unprojected_nodes, g_unprojected_edges)
G_u = ox.utils_graph.get_undirected(G)
# -
# ## Data sources
# +
import pandas as pd, geopandas as gpd, folium
pd.options.display.float_format = '{:.10f}'.format
# -
# ### Paired cache GDF, parishes GDF
parishes_gdf = gpd.read_file('data/parishes_gdf.gpkg')
paired_cache_df = pd.read_pickle('data/paired_cache_df.pickle')
parishes_gdf
# ## Create GDF from paired cache DF
# +
import yaml
with open("proj_crs.yml", "r") as stream:
try:
epsg_dict = yaml.safe_load(stream)
except yaml.YAMLError as exc:
print(exc)
proj_epsg_str = str(epsg_dict).replace("{","").replace("}", "").replace("'","").replace(" ","")
print('Projected graph EPSG code:',proj_epsg_str)
# -
paired_cache_df.info()
# + tags=[]
# %%time
from oxtools.compute_d3 import nodes_to_linestring
paired_cache_df['geometry'] = paired_cache_df['path_node_list'].map(\
lambda x: nodes_to_linestring(x, G_proj_u))
# +
paired_cache_gdf = gpd.GeoDataFrame(paired_cache_df, geometry='geometry', crs=proj_epsg_str)
paired_cache_gdf
# -
# ### Change GDF projection
paired_cache_proj_gdf = paired_cache_gdf.to_crs(proj_epsg_str)
parishes_proj_gdf = parishes_gdf.to_crs(proj_epsg_str)
# ### Add two new columns
paired_cache_proj_gdf['parish_route'] = ''
paired_cache_proj_gdf['parish_num'] = 0
paired_cache_proj_gdf
# References:
# 1. Convex Hull: https://geopandas.org/docs/user_guide/geometric_manipulations.html#GeoSeries.convex_hull
# 2. Distance between geoseries: https://geopandas.org/docs/reference/api/geopandas.GeoSeries.distance.html#geopandas.GeoSeries.distance
# 3. Geoseries contains: https://geopandas.org/docs/reference/api/geopandas.GeoSeries.contains.html#
# ## Set Parish GDF index to `NAME_4` (parish name)
parishes_proj_gdf = parishes_proj_gdf.set_index(['NAME_4'])
parishes_proj_gdf.loc[['Kisugu']]
# ## Identify route-parish intersections and parish routes
parishes = [ row.Index for row in parishes_proj_gdf.itertuples() ]
paired_cache_proj_gdf
# +
# %%time
from shapely.geometry import Point
from tqdm import tqdm, notebook
from oxtools.compute_d3 import get_parish_route
# tqdm parameters
total_rows=paired_cache_proj_gdf.shape[0]
# route loop
for r_row in tqdm(paired_cache_proj_gdf.itertuples(), total=total_rows):
# parishes intersected by route
parish_list = [ p_row.Index for p_row in parishes_proj_gdf.itertuples() \
if r_row.geometry.distance(p_row.geometry.convex_hull) == 0 ]
# go through path nodes to get right order of parishes (parish_route)
parish_route = get_parish_route(r_row.path_node_list, parishes_proj_gdf, parish_list, G_proj)
paired_cache_proj_gdf.at[r_row.Index, 'parish_route'] = parish_route
paired_cache_proj_gdf.at[r_row.Index, 'parish_num'] = len(parish_route)
# +
import matplotlib.pyplot as plt
fig, ax1 = plt.subplots(figsize=(24, 12))
plt.title('Sample Residence-to-Test-Facility Routes through Parishes in Kampala')
parishes_gdf.plot(ax=ax1, color='orange', edgecolor='black', linewidth=0.25)
paired_cache_proj_gdf.to_crs('EPSG:4326').plot(ax=ax1, color='darkblue', linewidth=1)
# -
paired_cache_proj_gdf.info()
paired_cache_proj_gdf[['parish_name', 'parish_num', 'd3_edge_attrs']].groupby(['parish_name']).mean()
paired_cache_proj_gdf[['parish_name', 'd3_edge_attrs']]\
.groupby(['parish_name']).mean()\
.plot.barh(\
figsize=(8, 6),
title='Mean D3 by edge attributes of Selected Kampala Parishes')
paired_cache_proj_gdf[['parish_name', 'parish_num']]\
.groupby(['parish_name']).mean()\
.plot.barh(\
figsize=(8, 6),
title='Number of Parishes Traversed from '+\
'Residence to Testing Site\n in Selected Kampala Parishes')
| NB05-Identify-Route-Parish-Intersections.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="vuVTg8VLViNd" colab_type="code" outputId="eab0d8ed-4187-4daa-abe7-7ca4232456d6" colab={"base_uri": "https://localhost:8080/", "height": 661} language="bash"
# pip install torch pytorch-transformers tensorboardX
# + id="gpgcK6edVyt7" colab_type="code" outputId="72087bc9-96ef-4d02-d094-f1aee0336b94" colab={"base_uri": "https://localhost:8080/", "height": 34}
from google.colab import drive
drive.mount('/content/gdrive')
# + id="rKblDQmBV5ky" colab_type="code" outputId="7760c0b0-711c-4e9a-e9d4-9ca1cf8c5d3c" colab={"base_uri": "https://localhost:8080/", "height": 34}
# %env TRAC_PATH /content/gdrive/My Drive/TRAC2020/
# + id="xM-Kk2LHWD5J" colab_type="code" outputId="fd7c47af-c7a7-45d9-f624-d28efe001e37" colab={"base_uri": "https://localhost:8080/", "height": 467} language="bash"
# echo "${TRAC_PATH}"
# ls -ltrh "${TRAC_PATH}/data"
# realpath "${TRAC_PATH}"
# ls -ltrh "${TRAC_PATH}"/data/**/*
# + id="7tLA7i6-WI6W" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 398} outputId="55a36aeb-0fc3-40d3-e1ca-70a2386fc40e" language="bash"
# python "${TRAC_PATH}"/src/generate_data.py
# + id="CLHYsWUCTzQV" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="3e9abfd0-5edf-442b-f925-4d26138ee34d" language="bash"
# python "${TRAC_PATH}"/src/run_experiment.py --help
# + [markdown] id="IuiCojl3h-dq" colab_type="text"
# # English
#
# ## bert-base-cased
# + id="gjwd1ny1UpCm" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="b9151ec6-c344-46a6-96d3-2dd9f559a362" language="bash"
# TASK="Sub-task A"
# LANG="ENG"
# MODEL="bert-base-cased"
# python "${TRAC_PATH}"/src/run_experiment.py \
# --data_dir "./${LANG}/${TASK}" \
# --bert_model_type "${MODEL}" \
# --task "${TASK}" \
# --lang ${LANG} \
# --output_dir "./${LANG}/${TASK}/output" \
# --cache_dir "./${LANG}/${TASK}" \
# --num_train_epochs 5 \
# --train_batch_size 32 \
# --eval_batch_size 32 \
# --save_steps 50 \
# --logging_steps 5 \
# --gradient_accumulation_steps 2 \
# + id="v2iXQ1dRVOUA" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="4994457f-85b6-4278-af4e-d3e11959b490" language="bash"
# TASK="Sub-task B"
# LANG="ENG"
# MODEL="bert-base-cased"
# python "${TRAC_PATH}"/src/run_experiment.py \
# --data_dir "./${LANG}/${TASK}" \
# --bert_model_type "${MODEL}" \
# --task "${TASK}" \
# --lang ${LANG} \
# --output_dir "./${LANG}/${TASK}/output" \
# --cache_dir "./${LANG}/${TASK}" \
# --num_train_epochs 5 \
# --train_batch_size 32 \
# --eval_batch_size 32 \
# --save_steps 50 \
# --logging_steps 5 \
# --gradient_accumulation_steps 2 \
# + id="m_wd33YPipfi" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="2ef9f25b-1904-4056-8694-5cb386cbc621" language="bash"
# TASK="Sub-task A"
# LANG="ENG"
# MODEL="bert-base-cased"
# mkdir -p "./${LANG}/${TASK}/${MODEL}"
# mv "./${LANG}/${TASK}/output/"/* "./${LANG}/${TASK}/${MODEL}/"
# mv "./${LANG}/${TASK}/${MODEL}" "./${LANG}/${TASK}/output/"
# ls "./${LANG}/${TASK}/output/"
# + [markdown] id="rW4x7kmriG3y" colab_type="text"
# ## bert-base-uncased
# + id="JEz1mhMgVpgc" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="13f6430c-711d-4da8-8dfe-1911f02439d0" language="bash"
# TASK="Sub-task A"
# LANG="ENG"
# MODEL="bert-base-uncased"
# python "${TRAC_PATH}"/src/run_experiment.py \
# --data_dir "./${LANG}/${TASK}" \
# --bert_model_type "${MODEL}" \
# --task "${TASK}" \
# --lang ${LANG} \
# --output_dir "./${LANG}/${TASK}/output/${MODEL}" \
# --cache_dir "./${LANG}/${TASK}" \
# --num_train_epochs 5 \
# --train_batch_size 32 \
# --eval_batch_size 32 \
# --save_steps 50 \
# --logging_steps 5 \
# --gradient_accumulation_steps 2 \
# + id="NJF_a8vDkrRt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="ba41b582-f69a-4be1-9899-46b3fc2567df" language="bash"
# TASK="Sub-task B"
# LANG="ENG"
# MODEL="bert-base-uncased"
# python "${TRAC_PATH}"/src/run_experiment.py \
# --data_dir "./${LANG}/${TASK}" \
# --bert_model_type "${MODEL}" \
# --task "${TASK}" \
# --lang ${LANG} \
# --output_dir "./${LANG}/${TASK}/output/${MODEL}" \
# --cache_dir "./${LANG}/${TASK}" \
# --num_train_epochs 5 \
# --train_batch_size 32 \
# --eval_batch_size 32 \
# --save_steps 50 \
# --logging_steps 5 \
# --gradient_accumulation_steps 2 \
# + [markdown] id="j4m5lPEiBBJ_" colab_type="text"
# # Hindi
#
# ## bert-base-cased
# + id="4rzn-iaYQ6kD" colab_type="code" colab={} language="bash"
# TASK="Sub-task A"
# LANG="HIN"
# MODEL="bert-base-multilingual-uncased"
# python "${TRAC_PATH}"/src/run_experiment.py \
# --data_dir "./${LANG}/${TASK}" \
# --bert_model_type "${MODEL}" \
# --task "${TASK}" \
# --lang ${LANG} \
# --output_dir "./${LANG}/${TASK}/output/${MODEL}" \
# --cache_dir "./${LANG}/${TASK}" \
# --num_train_epochs 5 \
# --train_batch_size 32 \
# --eval_batch_size 32 \
# --save_steps 50 \
# --logging_steps 5 \
# --gradient_accumulation_steps 2 \
# + id="A1nJOivHBo8j" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="de213e0a-52d5-4232-ad0e-29e5e5de0915" language="bash"
# TASK="Sub-task B"
# LANG="HIN"
# MODEL="bert-base-multilingual-uncased"
# python "${TRAC_PATH}"/src/run_experiment.py \
# --data_dir "./${LANG}/${TASK}" \
# --bert_model_type "${MODEL}" \
# --task "${TASK}" \
# --lang ${LANG} \
# --output_dir "./${LANG}/${TASK}/output/${MODEL}" \
# --cache_dir "./${LANG}/${TASK}" \
# --num_train_epochs 5 \
# --train_batch_size 32 \
# --eval_batch_size 32 \
# --save_steps 50 \
# --logging_steps 5 \
# --gradient_accumulation_steps 2 \
# + [markdown] id="0blaIwr_RLV5" colab_type="text"
# # Bengali
#
# ## bert-base-multilingual-uncased
# + id="YK12SX8_RRg-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="a1f8f19a-23e0-41f3-927d-9bc432630c35" language="bash"
# TASK="Sub-task A"
# LANG="IBEN"
# MODEL="bert-base-multilingual-uncased"
# python "${TRAC_PATH}"/src/run_experiment.py \
# --data_dir "./${LANG}/${TASK}" \
# --bert_model_type "${MODEL}" \
# --task "${TASK}" \
# --lang ${LANG} \
# --output_dir "./${LANG}/${TASK}/output/${MODEL}" \
# --cache_dir "./${LANG}/${TASK}" \
# --num_train_epochs 5 \
# --train_batch_size 32 \
# --eval_batch_size 32 \
# --save_steps 50 \
# --logging_steps 5 \
# --gradient_accumulation_steps 2 \
# + id="vCX2BLPKRZVZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="59c6e4bb-acf5-486c-b63f-63a2d68d4239" language="bash"
# TASK="Sub-task B"
# LANG="IBEN"
# MODEL="bert-base-multilingual-uncased"
# python "${TRAC_PATH}"/src/run_experiment.py \
# --data_dir "./${LANG}/${TASK}" \
# --bert_model_type "${MODEL}" \
# --task "${TASK}" \
# --lang ${LANG} \
# --output_dir "./${LANG}/${TASK}/output/${MODEL}" \
# --cache_dir "./${LANG}/${TASK}" \
# --num_train_epochs 5 \
# --train_batch_size 32 \
# --eval_batch_size 32 \
# --save_steps 50 \
# --logging_steps 5 \
# --gradient_accumulation_steps 2 \
# + [markdown] id="5LwEBfsbA3Dy" colab_type="text"
# ## Upload outputs
# + id="jjDJ0fGXtTFh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 294} outputId="bddf3605-a53b-4d49-ceba-46424bfff11f" language="bash"
# TASK="Sub-task B"
# LANG="IBEN"
# MODEL="bert-base-multilingual-uncased"
# tar -czvf "./${LANG}/${TASK}/output/${MODEL}.tar.gz" "./${LANG}/${TASK}/output/${MODEL}"/{*.tsv,*.json,events.*,model/config.json}
# curl "https://bashupload.com/${LANG}_${TASK}_${MODEL}.tar.gz" --data-binary @"./${LANG}/${TASK}/output/${MODEL}.tar.gz"
# + id="3xb_3bqt-woS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 173} outputId="4fae94f1-9edc-4ef6-a3c8-6c13d2018af3" language="bash"
# TASK="Sub-task A"
# LANG="ENG"
# MODEL="bert-base-cased"
# curl "https://bashupload.com/${LANG}_${TASK}_${MODEL}.tar.gz" --data-binary @"./${LANG}/${TASK}/output/${MODEL}.tar.gz"
# + id="3z3SYpxE3OtE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 138} outputId="37eb7fa3-36de-4542-eef1-55434cb44764" language="bash"
# TASK="Sub-task A"
# LANG="ENG"
# MODEL="bert-base-cased"
# tar -tzf "./${LANG}/${TASK}/output/${MODEL}.tar.gz"
# + id="I_0cHYvo3ZUm" colab_type="code" colab={}
| notebooks/TRAC_Experiments.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Moodle Database: Educational Data Log Analysis
#
# ## Analysing the 2019 10 Academy learners activity in the Moodle Learning Management System.
# The Moodle LMS is a free and open-source learning management system written in PHP and distributed under the GNU General Public License. It is used for blended learning, distance education, flipped classroom and other e-learning projects in schools, universities, workplaces and other sectors. With customizable management features, it is used to create private websites with online courses for educators and trainers to achieve learning goals. Moodle allows for extending and tailoring learning environments using community-sourced plugins.
#
# In 2019, 10 Academy used the Moodle LMS to manage about 1000 students in their 6 months data science training. Learners, course instructors, and all admins interacted with the Moodle system for almost all the training activities. All events from these activities are logged in the moodle postgres database.
# Moodle database is complex - with more than 400 connected tables! In this project we are interested only in the subset of the tables. The most important tables we will consider in this challenge are (tables in bold are VIP)
# * mdl_logstore_standard_log
# * mdl_context
# * mdl_user
# * mdl_course
# * mdl_modules
# * mdl_course_modules
# * mdl_course_modules_completion
# * mdl_grade_items
# * mdl_grade_grades
# * mdl_grade_categories
# * mdl_grade_items_history
# * mdl_grade_grades_history
# * mdl_grade_categories_history
# * mdl_forum
# * mdl_forum_discussions
# * mdl_forum_posts
#
# ## Task 2 - Data Extraction Transformation and Loading (ETL)
#
#Installing psycopg2 which is a DB API 2.0 compliant PostgreSQL driver
# !pip install psycopg2
# +
# %load_ext sql
# -
import pandas as pd
import os
import psycopg2
from sqlalchemy import create_engine
engine = create_engine('postgresql://postgres:Komboelvis01@localhost/moodle')
# %sql postgresql://postgres:Komboelvis01@localhost/moodle
# After connecting our postgres database, specifically to the database where the moodle log data is stored, we can start querying from the notebook
# +
# This Python script connects to a PostgreSQL database and utilizes Pandas to obtain data and create a data frame
# Establish a connection to the database by creating a cursor object
# Connect to the PostgreSQL database
host = "localhost" # either "localhost", a domain name, or an IP address.
port = "5432" # default postgres port
dbname = "moodle"
user = "postgres"
pw = "<PASSWORD>"
conn = psycopg2.connect(host=host, port=port, dbname=dbname, user=user, password=pw)
# Create a new cursor
cur = conn.cursor()
# A function that takes in a PostgreSQL query and outputs a pandas database
def pull(sql_query, database = conn):
table = pd.read_sql_query(sql_query, database)
return table
# Utilize the create_pandas_table function to create a Pandas data frame
# Store the data as a variable
overall_grade = pull("select AVG(finalgrade) as overall_grade_of_learners from mdl_grade_grades")
print('Overall grade of learners ')
print(overall_grade)
print('\n')
forum_posts = pull("select count(*) as number_of_forum_posts from mdl_forum_posts")
print('Number of forum posts')
print(forum_posts)
# Close the cursor and connection to so the server can allocate
# bandwidth to other requests
cur.close()
conn.close()
# -
# Python function to communicate with the database, they are not part of the assignment but i was practising
# +
from requests import get, post
# Module variables to connect to moodle api
KEY = "SECRET API KEY"
URL = "https://moodle.site.com"
ENDPOINT="/webservice/rest/server.php"
def rest_api_parameters(in_args, prefix='', out_dict=None):
"""Transform dictionary/array structure to a flat dictionary, with key names
defining the structure.
Example usage:
>>> rest_api_parameters({'courses':[{'id':1,'name': 'course1'}]})
{'courses[0][id]':1,
'courses[0][name]':'course1'}
"""
if out_dict==None:
out_dict = {}
if not type(in_args) in (list,dict):
out_dict[prefix] = in_args
return out_dict
if prefix == '':
prefix = prefix + '{0}'
else:
prefix = prefix + '[{0}]'
if type(in_args)==list:
for idx, item in enumerate(in_args):
rest_api_parameters(item, prefix.format(idx), out_dict)
elif type(in_args)==dict:
for key, item in in_args.items():
rest_api_parameters(item, prefix.format(key), out_dict)
return out_dict
def call(fname, **kwargs):
"""Calls moodle API function with function name fname and keyword arguments.
Example:
>>> call_mdl_function('core_course_update_courses',
courses = [{'id': 1, 'fullname': 'My favorite course'}])
"""
parameters = rest_api_parameters(kwargs)
parameters.update({"wstoken": KEY, 'moodlewsrestformat': 'json', "wsfunction": fname})
response = post(URL+ENDPOINT, parameters)
response = response.json()
if type(response) == dict and response.get('exception'):
raise SystemError("Error calling Moodle API\n", response)
return response
class CourseList():
"""Class for list of all courses in Moodle and order them by id and idnumber."""
def __init__(self):
# TODO fullname atribute is filtered
# (no <span class="multilang" lang="sl">)
courses_data = call('core_course_get_courses')
self.courses = []
for data in courses_data:
self.courses.append(Course(**data))
self.id_dict = {}
self.idnumber_dict = {}
for course in self.courses:
self.id_dict[course.id] = course
if course.idnumber:
self.idnumber_dict[course.idnumber] = course
def __getitem__(self, key):
if 0<= key < len(self.courses):
return self.courses[key]
else:
raise IndexError
def by_id(self, id):
"Return course with given id."
return self.id_dict.get(id)
def by_idnumber(self, idnumber):
"Course with given idnumber"
return self.idnumber_dict.get(idnumber)
def update_courses(courses_to_update, fields):
"Update a list of courses in one go."
if not ('id' in fields):
fields.append('id')
courses = [{k: c.__dict__[k] for k in fields} for c in courses_to_update]
return call("core_course_update_courses",
courses = courses)
class Course():
"""Class for a single course.
Example:
>>> Course(name="Example course", shortname="example", categoryid=1, idnumber=123)
"""
def __init__(self, **data):
self.__dict__.update(data)
def create(self):
"Create this course on moodle"
res = call('core_course_create_courses', courses = [self.__dict__])
if type(res) == list:
self.id = res[0].get('id')
def update(self):
"Update course"
r = call('core_course_update_courses', courses = [self.__dict__])
class User():
"""Class for a single user.
Example:
>>> User(name="Janez", surname="Novak", email="<EMAIL>", username="jnovak", password="<PASSWORD>")"""
def __init__(self, **data):
self.__dict__.update(data)
def create(self):
"Create new user on moodle site"
valid_keys = ['username',
'firstname',
'lastname',
'email',
'auth',
'idnumber',
'password']
values = {key: self.__dict__[key] for key in valid_keys}
res = call('core_user_create_users', users = [values])
if type(res) == list:
self.id = res[0].get('id')
def update(self, field=None):
"Upadte user data on moodle site"
if field:
values = {"id": self.id, field: self.__dict__[field]}
else:
values = self.__dict__
r = call('core_user_update_users', users = [values])
def get_by_field(self, field='username'):
"Create new user if it does not exist, otherwise update data"
res = call('core_user_get_users_by_field', field = field, values = [self.__dict__[field]])
if (type(res) == list) and len(res) > 0:
self.__dict__.update(res[0])
return self
else:
return None
def create_or_get_id(self):
"Get Moodle id of the user or create one if it does not exists."
if not self.get_by_field():
self.create()
def enroll(self, roleid=5):
"Enroll users in courses with specific role"
if len(self.courses)<=0:
return None
enrolments = []
for course in self.courses:
enrolments.append({'roleid': roleid, 'userid': self.id, 'courseid': course.id})
r = call('enrol_manual_enrol_users', enrolments = enrolments)
return r
def enrolments(self, m_courses):
"Get moodle courses, the user has to enroll"
self.courses = []
for idnumber in self.course_idnumbers:
course = m_courses.by_idnumber(idnumber)
if course:
self.courses.append(course)
return self.courses
class Cathegory():
pass
class Enrolments():
pass
# -
# Python class to fetch the overall grade of learners and the number of forum posts plus the login counts
class ForumGrades():
"""
This class would can be used:
1. get a connection with the moodle postgres db
2. calcualte the sum/average/count of grades
3. calculate the number of forum posts
"""
import pandas as pd
import os
import psycopg2
def __init__(self, user:str, db:str, host:str,password:str, port:str=None):
self.user = user
self.db = db
self.host = host
self.port = port
self.password = password
self._make_connection_()
def __repr__(self):
return "Object to get Overall grade of learners for Number of forum posts"
def _make_connection_(self): # this is private method not accessible by the object
try:
if self.password is None:
self.password = os.getenv('PASSWORD')
self.connection = psycopg2.connect(user = self.user,
password = self.password,
host = self.host,
port = self.port,
database = self.db)
# perform autocommit on queries
self.connection.set_session(autocommit=True)
self.cursor = self.connection.cursor()
# Print PostgreSQL version
self.cursor.execute("SELECT version();")
self.record = self.cursor.fetchone()
print("You are connected to - ", self.record,"\n")
except (Exception, psycopg2.Error) as error :
print ("Error while connecting to PostgreSQL", error)
sys.exit(1)
return self
def get_grades(self):
result = pd.read_sql_query( sql = "SELECT AVG(finalgrade) AS grade_avg, \
COUNT(finalgrade) AS grade_counts, \
SUM(finalgrade) AS grades_sum FROM mdl_grade_grades",
con = self.connection, )
return result
def user_dedication_time(self):
result = pd.read_sql_query( sql = "select id,\
EXTRACT(HOURS FROM to_timestamp(firstaccess)) as firstaccess,\
EXTRACT(HOUR FROM to_timestamp(lastlogin)) as lastlogin\
from mdl_user\
order by id",
con = self.connection)
return result
def activity_count(self):
result=pd.read_sql_query( sql = "select userid, COUNT(action) as Activity_count\
from mdl_logstore_standard_log\
group by userid\
order by userid",
con = self.connection)
return result
def login_count(self):
result=pd.read_sql_query( sql ="")
def get_forum(self):
result = pd.read_sql_query( sql = "SELECT COUNT(id) AS forum_counts \
FROM mdl_forum_posts;",
con = self.connection, )
return result
def close(self):
return self.connection.close()
forum_grades = ForumGrades(user='postgres', host='127.0.0.1',
db='moodle',password='<PASSWORD>')
forum_grades.activity_count()
# Login_Count
# Using the sqlalchemy connection we made at the beginning, we now fetch two tables from our moodle database; mdl_user and mdl_logstore_standard_log tables
log_df = pd.read_sql("select * from mdl_logstore_standard_log", engine)
log_df1 = pd.read_sql("select * from mdl_user", engine)
log_df1
new_df
def top_x(df, percent):
tot_len = df.shape[0]
top = int((tot_len * percent)/100)
return df.iloc[:top,]
log_df_logged_in = log_df[log_df.action == 'loggedin'][['userid', 'action']]
login_by_user = log_df_logged_in.groupby('userid').count().sort_values('action', ascending=False)
login_by_user.columns = ['login_count']
top_x(login_by_user, 5)
login_by_user
activity_log = log_df[['userid', 'action']]
activity_log_by_user = activity_log.groupby('userid').count().sort_values('action', ascending=False)
activity_log_by_user.columns = ['activity_count']
top_x(activity_log_by_user, 1)
log_df.info()
log_in_out = log_df[(log_df.action == "loggedin") | (log_df.action == "loggedout")]
# +
user_id = log_df.userid.unique()
d_times = {}
l = 0
for user in user_id:
l += 1
log_user = log_in_out[log_in_out.userid == user].sort_values('timecreated')
d_time = 0
isLoggedIn = 0
loggedIn_timecreated = 0
for i, row in log_user.iterrows():
if(row.action == "loggedin"):
isLoggedIn = 1
loggedIn_timecreated = row.timecreated
if((row.action == "loggedout") & (isLoggedIn == 1)):
d_time += row.timecreated - loggedIn_timecreated
isLoggedIn = 0
d_times[user] = d_time
# +
id = log_df.userid.unique()
d_times = {}
for user in id:
log_user = log_df[log_df.userid == user].sort_values('timecreated')
d_time = 0
isLoggedIn = 0
loggedIn_timecreated = 0
for i in range(len(log_user)):
row = log_user.iloc[i,]
row_next = log_user.iloc[i+1,] if i+1 < len(log_user) else row
if(row.action == "loggedin"):
isLoggedIn = 1
loggedIn_timecreated = row.timecreated
if( (i+1 == len(log_user)) | ( (row_next.action == "loggedin") & (isLoggedIn == 1) ) ):
d_time += row.timecreated - loggedIn_timecreated
isLoggedIn = 0
d_times[user] = d_time
# -
dedication_time_df = pd.DataFrame( {'id':list(d_times.keys()),
'dedication_time':list(d_times.values())})
dedication_time_df
new_df=log_df1.merge(dedication_time_df, on='id')
new_df.to_csv('user_data.csv',index=False)
dedication_time_df.to_csv('dedi_df.csv',index=False)
data=top_x(dedication_time_df.sort_values('dedication_time', ascending=False), 1)
import seaborn as sns
ax = sns.barplot(x="id", y="dedication_time", data=data)
| Moodle_databaseTask2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="RiQPyEYUwGRY" colab={"base_uri": "https://localhost:8080/"} outputId="91cd9b2c-42d4-4404-8878-72636fa05d04"
from google.colab import drive
drive.mount('/content/drive')
# + id="KjZyKQNUwTig"
# !cp -r '/content/drive/My Drive/mycode_dataset' '/subject101.dat'
# + id="mt5d6la87ko2"
import numpy as np
import pandas as pd
import seaborn as sns
from matplotlib import pyplot as plt
from IPython.display import display
from sklearn.model_selection import train_test_split
from sklearn import svm
from sklearn.model_selection import GridSearchCV
from sklearn import preprocessing
from sklearn import metrics
from sklearn.metrics import precision_score, recall_score
from sklearn.metrics import confusion_matrix
from sklearn.metrics import confusion_matrix,accuracy_score,log_loss
# + id="rSdiekvj4P-W" colab={"base_uri": "https://localhost:8080/"} outputId="fdc5ff5d-d3a1-4ebe-df00-1b2bef991988"
df1=pd.read_table(r'/content/drive/My Drive/mycode_dataset' '/subject101.dat')
print(df1)
# + id="DTgTinTU7pC9"
def load_activity_map():
map = {}
map[0] = 'transient'
map[1] = 'lying'
map[2] = 'sitting'
map[3] = 'standing'
map[4] = 'walking'
map[5] = 'running'
map[6] = 'cycling'
map[7] = 'Nordic_walking'
map[9] = 'watching_TV'
map[10] = 'computer_work'
map[11] = 'car driving'
map[12] = 'ascending_stairs'
map[13] = 'descending_stairs'
map[16] = 'vacuum_cleaning'
map[17] = 'ironing'
map[18] = 'folding_laundry'
map[19] = 'house_cleaning'
map[20] = 'playing_soccer'
map[24] = 'rope_jumping'
return map
# + id="eaP5IhH874qV"
def generate_three_IMU(name):
x = name +'_x'
y = name +'_y'
z = name +'_z'
return [x,y,z]
def generate_four_IMU(name):
x = name +'_x'
y = name +'_y'
z = name +'_z'
w = name +'_w'
return [x,y,z,w]
def generate_cols_IMU(name):
# temp
temp = name+'_temperature'
output = [temp]
# acceleration 16
acceleration16 = name+'_3D_acceleration_16'
acceleration16 = generate_three_IMU(acceleration16)
output.extend(acceleration16)
# acceleration 6
acceleration6 = name+'_3D_acceleration_6'
acceleration6 = generate_three_IMU(acceleration6)
output.extend(acceleration6)
# gyroscope
gyroscope = name+'_3D_gyroscope'
gyroscope = generate_three_IMU(gyroscope)
output.extend(gyroscope)
# magnometer
magnometer = name+'_3D_magnetometer'
magnometer = generate_three_IMU(magnometer)
output.extend(magnometer)
# oreintation
oreintation = name+'_4D_orientation'
oreintation = generate_four_IMU(oreintation)
output.extend(oreintation)
return output
def load_IMU():
output = ['time_stamp','activity_id', 'heart_rate']
hand = 'hand'
hand = generate_cols_IMU(hand)
output.extend(hand)
chest = 'chest'
chest = generate_cols_IMU(chest)
output.extend(chest)
ankle = 'ankle'
ankle = generate_cols_IMU(ankle)
output.extend(ankle)
return output
def load_subjects(root='/content/drive/My Drive/mycode_dataset' '/subject'):
output = pd.DataFrame()
cols = load_IMU()
for i in range(101,102):
path = root + str(i) +'.dat'
subject = pd.read_table(path, header=None, sep='\s+')
subject.columns = cols
subject['id'] = i
output = output.append(subject, ignore_index=True)
output.reset_index(drop=True, inplace=True)
return output
data = load_subjects()
# + id="HD2ux0hA8xYv" colab={"base_uri": "https://localhost:8080/", "height": 437} outputId="e5de1808-4547-4e04-b0a8-fc970f23aa18"
data
# + id="5zwqg3D3DqMs"
def fix_data(data):
select = [4,5,7,12,13,24]
#data = data.drop(data[data['activity_id']==0].index)
data=data[data['activity_id'].isin(select)]
data=data[['time_stamp','activity_id','ankle_3D_acceleration_6_x','ankle_3D_acceleration_6_y','ankle_3D_acceleration_6_z','ankle_3D_gyroscope_x','ankle_3D_gyroscope_y','ankle_3D_gyroscope_z','id']]
data = data.interpolate()
# fill all the NaN values in a coulmn with the mean values of the column
for colName in data.columns:
data[colName] = data[colName].fillna(data[colName].mean())
activity_mean = data.groupby(['activity_id']).mean().reset_index()
return data
data = fix_data(data)
# + id="ZvD-r0MnA1ml" colab={"base_uri": "https://localhost:8080/", "height": 417} outputId="a93afbbb-60f1-4498-d574-52579889d3a3"
data
# + id="Te0UTxxZH5eK" colab={"base_uri": "https://localhost:8080/"} outputId="b7d662a0-922d-4200-abce-528cb94c06a4"
data['activity_id'].value_counts()
# + id="5qEaCoyhEB96" colab={"base_uri": "https://localhost:8080/", "height": 315} outputId="e8e22e51-13c4-4afd-8997-700285cb0271"
data.describe()
# + id="IZmf6UumEsgL" colab={"base_uri": "https://localhost:8080/"} outputId="f8d75aef-6da2-4b33-9553-76956358dd0d"
print('Size of the data: ', data.size)
print('Shape of the data: ', data.shape)
print('Number of columns in the data: ', len(data.columns))
#result_id = data.groupby(['id']).mean().reset_index()
#print('Number of unique ids in the data: ', len(result_id))
result_act = data.groupby(['activity_id']).mean().reset_index()
print('Numbe of unique activities in the data: ',len(result_act))
# + id="hc-iy0kcFO7T"
def pd_fast_plot(pd,column_a,column_b,title, figsize=(10,6)):
plt.rcParams.update({'font.size': 16})
size = range(len(pd))
f, ax = plt.subplots(figsize=figsize)
plt.bar(size, pd[column_a], color=plt.cm.Paired(size))
a = ax.set_xticklabels(pd[column_b])
b = ax.legend(fontsize = 20)
c = ax.set_xticks(np.arange(len(pd)))
d = ax.set_title(title)
plt.show()
# + id="gY3UlhsFFXbT" colab={"base_uri": "https://localhost:8080/", "height": 417} outputId="c726b9d8-2046-4a6d-b33c-7ba4b88b2593"
samples = data.groupby(['id']).count().reset_index()
samples_to_subject = pd.DataFrame()
samples_to_subject['id'] = samples['id']
samples_to_subject['samples'] = samples['time_stamp']
samples_to_subject = samples_to_subject.sort_values(by=['samples'])
pd_fast_plot(samples_to_subject,'samples','id','Number Of Samepls By Users')
# + id="5FzRQmJMFncz" colab={"base_uri": "https://localhost:8080/", "height": 303} outputId="3f53a54c-a680-4132-d478-9b1d0de8dd54"
map_ac = load_activity_map()
samples = data.groupby(['activity_id']).count().reset_index()
samples_to_subject = pd.DataFrame()
samples_to_subject['activity'] = [map_ac[x] for x in samples['activity_id']]
samples_to_subject['samples'] = samples['time_stamp']
samples_to_subject = samples_to_subject.sort_values(by=['samples'])
pd_fast_plot(samples_to_subject,'samples','activity','Number Of Samepls By Activity',figsize=(40,7))
# + id="OqubkeYjTufO"
features = data[['ankle_3D_acceleration_6_x','ankle_3D_acceleration_6_y','ankle_3D_acceleration_6_z','ankle_3D_gyroscope_x','ankle_3D_gyroscope_y','ankle_3D_gyroscope_z']]
label = data['activity_id']
# + colab={"base_uri": "https://localhost:8080/", "height": 417} id="Tl9ljq4-VZgO" outputId="5e16faa2-ce48-4b77-ed3a-09d89ed17f32"
features
# + colab={"base_uri": "https://localhost:8080/"} id="kEcwa9-JVc8F" outputId="8f7ddc1e-b9aa-442d-f980-452c9b84476a"
label
# + id="iAoR0MS0VsGW"
x_train, x_test, y_train, y_test = train_test_split(features, label ,test_size = 0.3, shuffle=True)
# + colab={"base_uri": "https://localhost:8080/"} id="SNq4rKCWWKWO" outputId="ccb28c1c-5aa6-447f-ba7b-55b90d7eb63b"
#Dimension of train and test set
print('Train set dimension : ',x_train.shape)
print('Test set dimension : ' ,x_test.shape)
# + id="zjnkXQN2Wy7u"
x_train = preprocessing.scale(x_train)
x_test = preprocessing.scale(x_test)
parameters = {'kernel':(['rbf']), 'gamma':[0.00001, 0.0001,0.001,0.1], 'C':[ 1, 10, 100, 1000]} #,{'kernel':(['poly']), 'C':[100], 'gamma':[0.0001]}]
svr = svm.SVC()
clf = GridSearchCV(svr, parameters,cv=5)
#Create a svm Classifier
#clf = svm.SVC(kernel='rbf',gamma=0.001,C=1000) # Linear Kernel
#Train the model using the training sets
clf.fit(x_train, y_train)
#Predict the response for test dataset
y_pred = clf.predict(x_test)
# + colab={"base_uri": "https://localhost:8080/"} id="vO2wyv6iwg4H" outputId="88ef6f06-c74e-43fc-a29b-63e6476c8ec0"
parameters
# + colab={"base_uri": "https://localhost:8080/"} id="YEQhn1wgrmwm" outputId="0e10ae27-7df5-42f9-b21b-a848299de98a"
clf
# + colab={"base_uri": "https://localhost:8080/"} id="XfRbaDINnV9-" outputId="47eaf07e-6f2e-4c6a-a244-39ccce934660"
# Model Accuracy: how often is the classifier correct?
print("Accuracy:",metrics.accuracy_score(y_test, y_pred))
# + colab={"base_uri": "https://localhost:8080/"} id="kuy3myXWaYoQ" outputId="2979a508-68a1-4a22-fe9a-989764c3a7e7"
# Model Precision: what percentage of positive tuples are labeled as such?
print("Precision:",metrics.precision_score(y_test, y_pred, average='macro'))
# Model Recall: what percentage of positive tuples are labelled as such?
print("Recall:",metrics.recall_score(y_test, y_pred, average='macro'))
# + colab={"base_uri": "https://localhost:8080/"} id="6Skk2AHHcku4" outputId="380d7142-8f4f-48b7-fb28-6f6c0454cba5"
confusion_matrix(y_test, y_pred)
# + colab={"base_uri": "https://localhost:8080/", "height": 639} id="H4b041lDfV-g" outputId="2f636948-a4c0-476b-f636-9197483a6d45"
def quick_plot_con_matrix(y, results,labels):
# now print confusion metrix
con = confusion_matrix(y,results)
a = plt.figure(figsize=(25,15), dpi=50)
a = sns.heatmap(con, cmap='YlGnBu', annot=True, fmt = 'd', xticklabels=labels, yticklabels=labels)
a = plt.rcParams.update({'font.size': 20})
a = plt.title('Confusion Matrix')
a = plt.xlabel('Predictions')
a = plt.ylabel('Accpected')
#activity_map = load_activity_map()
labels = [map_ac[x] for x in samples['activity_id']]
quick_plot_con_matrix(y_test,y_pred, labels)
| SVM and CNN/SVM_HAR.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#libraries
import pandas as pd
import numpy as np
import plotly.express as px
import plotly.figure_factory as ff
data_HP = "https://data.baltimorecity.gov/resource/fesm-tgxf.json"
df_HP = pd.read_json(data_HP)
df_HP.info()
df_HP.head()
#Removing Rows that have Null Values
df_HP = df_HP[df_HP["cost_est"].notnull()]
df_HP = df_HP[df_HP["neighborhood"].notnull()]
df_HP = df_HP[df_HP["policedistrict"].notnull()]
df_HP = df_HP[df_HP["councildistrict"].notnull()]
# rename corrupted column names
df_HP = df_HP.rename(columns = {":@computed_region_5kre_ccpb": "2010 Census Neighborhoods",
":@computed_region_gwq4_fjxs": "2010 Census Wards Precincts",
":@computed_region_s6p5_2pgr": "Zip Codes"})
df_HP.info()
| .ipynb_checkpoints/BAProject-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7
# language: python
# name: python3
# ---
# +
# The code was removed by Watson Studio for sharing.
# +
#For accessing files
f = open('accessfiles.py', 'wb')
f.write(streaming_body_1.read())
import accessfiles
# -
# !pip install pyspark==2.4.5
# +
from zipfile import ZipFile
import time
import numpy as np
from operator import add
from pyspark import SparkContext, SparkConf
from pyspark.sql import SQLContext, SparkSession
from pyspark.sql.types import StructType, StructField, DoubleType, IntegerType, StringType
import os
from pyspark.sql.functions import unix_timestamp, to_date
import pyspark.sql.functions as F
sc = SparkContext.getOrCreate(SparkConf().setMaster("local[*]"))
spark = SparkSession \
.builder \
.getOrCreate()
spark.conf.set("spark.sql.repl.eagerEval.enabled", True)
from accessfiles import load_files
load_files()
# +
ratingsRDD=sc.textFile("ratings.dat")
movies=ratingsRDD.map(lambda line:int(line.split("::")[1]))
movies_pair=movies.map(lambda mv:(mv,1))
movies_count=movies_pair.reduceByKey(add)
movies_sorted=movies_count.sortBy(lambda x: x[1], False, 1)
mv_top10List=movies_sorted.take(10)
mv_top10RDD=sc.parallelize(mv_top10List)
mv_names=sc.textFile("movies.dat").map(lambda line:(int(line.split("::")[0]),line.split("::")[1]))
print(mv_top10RDD.collect())
join_out=mv_names.join(mv_top10RDD)
top10_rdd = join_out.sortBy(lambda x: x[1][1],False).map(lambda x: (x[0],x[1][0],x[1][1])).repartition(1)
top10_df = top10_rdd.toDF().toPandas()
top10_df.columns=['Movie_ID','Name','Views']
# -
from accessfiles import create_download_link
create_download_link(top10_df, filename='Most_viewed_top10movies.csv')
| SparkRDD/Most_viewed_movies_top10.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="58EU409UZrOI"
# **Project Contributors: 18111006 (<NAME>), 18111015 (<NAME>), 18111016 (<NAME>), 18111025 (<NAME>)**
#
# **ResNet50**
# + colab={"base_uri": "https://localhost:8080/"} id="O5X3ASNLkv3Y" outputId="79e7e2dd-4c4c-4702-fc63-1576af3d7f93"
from google.colab import drive
drive.mount('/content/drive')
# + colab={"base_uri": "https://localhost:8080/"} id="qTi0RAUGEfoM" outputId="e0b05ca6-0ebd-4ce0-926b-512e8122501f"
from zipfile import ZipFile
filename="/content/drive/My Drive/HAM10000.zip"
with ZipFile(filename,'r') as zip:
zip.extractall()
print("done")
# + id="Aa36bMKLze3z"
import pandas as pd
import numpy as np
import os
import tensorflow as tf
import cv2
from keras import backend as K
from keras.layers import Layer,InputSpec
import keras.layers as kl
from glob import glob
from sklearn.metrics import roc_curve, auc
from keras.preprocessing import image
from tensorflow.keras.models import Sequential
from sklearn.metrics import roc_auc_score
from tensorflow.keras import callbacks
from tensorflow.keras.callbacks import ModelCheckpoint,EarlyStopping
from matplotlib import pyplot as plt
from tensorflow.keras import Model
from tensorflow.keras.layers import concatenate,Dense, Conv2D, MaxPooling2D, Flatten,Input,Activation,add,AveragePooling2D,GlobalAveragePooling2D,BatchNormalization,Dropout
# %matplotlib inline
import shutil
from sklearn.metrics import precision_score, recall_score, accuracy_score,classification_report ,confusion_matrix
from tensorflow.python.platform import build_info as tf_build_info
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from sklearn.model_selection import train_test_split
from PIL import ImageFile
ImageFile.LOAD_TRUNCATED_IMAGES = True
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="lnzRzk7e44HL" outputId="6d1e2d2f-1669-42a6-f6a7-45a52b73ce41"
data_pd = pd.read_csv('/content/drive/MyDrive/HAM10000_metadata.csv')
data_pd.head()
# + id="qlR6SjeEzXsm"
train_dir = os.path.join('HAM10000', 'train_dir')
test_dir = os.path.join('HAM10000', 'test_dir')
# + colab={"base_uri": "https://localhost:8080/", "height": 237} id="_IFqPgUu5jPj" outputId="979e5d51-8319-435c-f4db-1981b452c849"
df_count = data_pd.groupby('lesion_id').count()
df_count.head()
# + id="QjMQNZRI2xl7"
df_count = df_count[df_count['dx'] == 1]
df_count.reset_index(inplace=True)
# + id="NeVfs-Ly95gs"
def duplicates(x):
unique = set(df_count['lesion_id'])
if x in unique:
return 'no'
else:
return 'duplicates'
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="2WZZRSzO5v8t" outputId="5042fd49-03f1-48a6-ed7b-ac849f1fe0c9"
data_pd['is_duplicate'] = data_pd['lesion_id'].apply(duplicates)
data_pd.head()
# + id="3BhGlAv0yAHu"
df_count = data_pd[data_pd['is_duplicate'] == 'no']
# + id="Y3ndAO_Ex5fb"
train, test_df = train_test_split(df_count, test_size=0.15, stratify=df_count['dx'])
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="T7w2kYUdNkjX" outputId="5b6b9660-79f4-4fdb-ec79-4d5366f99a3e"
def identify_trainOrtest(x):
test_data = set(test_df['image_id'])
if str(x) in test_data:
return 'test'
else:
return 'train'
#creating train_df
data_pd['train_test_split'] = data_pd['image_id'].apply(identify_trainOrtest)
train_df = data_pd[data_pd['train_test_split'] == 'train']
train_df.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="FPySEG1m58pu" outputId="18fd6e44-8d62-4a88-efa7-d29aa64637ae"
test_df.head()
# + id="Ja7jQJQb39wi"
# Image id of train and test images
train_list = list(train_df['image_id'])
test_list = list(test_df['image_id'])
# + colab={"base_uri": "https://localhost:8080/"} id="lBJgBAjP13q5" outputId="463d1b3f-d77c-49b0-d89e-9ce5d21a47e0"
len(test_list)
# + colab={"base_uri": "https://localhost:8080/"} id="eEChk1DK-H8Z" outputId="612e1f17-9db3-4dea-c99e-0b7aa5ee566a"
len(train_list)
# + id="PIoMqylGAYYZ"
# Set the image_id as the index in data_pd
data_pd.set_index('image_id', inplace=True)
# + id="Ja_PtDYyDPMM"
os.mkdir(train_dir)
os.mkdir(test_dir)
# + id="PsoqCvNsgmHP"
targetnames = ['akiec', 'bcc', 'bkl', 'df', 'mel', 'nv', 'vasc']
# + id="9KYMTQugCmRR"
for i in targetnames:
directory1=train_dir+'/'+i
directory2=test_dir+'/'+i
os.mkdir(directory1)
os.mkdir(directory2)
# + id="GL9vFa3X-ty1"
for image in train_list:
file_name = image+'.jpg'
label = data_pd.loc[image, 'dx']
# path of source image
source = os.path.join('HAM10000', file_name)
# copying the image from the source to target file
target = os.path.join(train_dir, label, file_name)
shutil.copyfile(source, target)
# + id="hwbKrEzJ_if2"
for image in test_list:
file_name = image+'.jpg'
label = data_pd.loc[image, 'dx']
# path of source image
source = os.path.join('HAM10000', file_name)
# copying the image from the source to target file
target = os.path.join(test_dir, label, file_name)
shutil.copyfile(source, target)
# + colab={"base_uri": "https://localhost:8080/"} id="4W8hmE2OHjQa" outputId="e57e21fb-559b-40f8-8755-d8e45bd3e8a4"
targetnames = ['akiec', 'bcc', 'bkl', 'df', 'mel', 'nv', 'vasc']
# Augmenting images and storing them in temporary directories
for img_class in targetnames:
#creating temporary directories
# creating a base directory
aug_dir = 'aug_dir'
os.mkdir(aug_dir)
# creating a subdirectory inside the base directory for images of the same class
img_dir = os.path.join(aug_dir, 'img_dir')
os.mkdir(img_dir)
img_list = os.listdir('HAM10000/train_dir/' + img_class)
# Copy images from the class train dir to the img_dir
for file_name in img_list:
# path of source image in training directory
source = os.path.join('HAM10000/train_dir/' + img_class, file_name)
# creating a target directory to send images
target = os.path.join(img_dir, file_name)
# copying the image from the source to target file
shutil.copyfile(source, target)
# Temporary augumented dataset directory.
source_path = aug_dir
# Augmented images will be saved to training directory
save_path = 'HAM10000/train_dir/' + img_class
# Creating Image Data Generator to augment images
datagen = tf.keras.preprocessing.image.ImageDataGenerator(
rotation_range=180,
width_shift_range=0.1,
height_shift_range=0.1,
zoom_range=0.1,
horizontal_flip=True,
vertical_flip=True,
fill_mode='nearest'
)
batch_size = 50
aug_datagen = datagen.flow_from_directory(source_path,save_to_dir=save_path,save_format='jpg',target_size=(224, 224),batch_size=batch_size)
# Generate the augmented images
aug_images = 8000
num_files = len(os.listdir(img_dir))
num_batches = int(np.ceil((aug_images - num_files) / batch_size))
# creating 8000 augmented images per class
for i in range(0, num_batches):
images, labels = next(aug_datagen)
# delete temporary directory
shutil.rmtree('aug_dir')
# + id="wNisha_gM3_Z"
train_path = 'HAM10000/train_dir'
test_path = 'HAM10000/test_dir'
batch_size=16
# + id="zhQWqdRN79B3"
datagen=ImageDataGenerator(preprocessing_function=tf.keras.applications.inception_resnet_v2.preprocess_input)
# + colab={"base_uri": "https://localhost:8080/"} id="w9_8FvOO7Rtu" outputId="7ca88c24-fda8-45d6-cb32-f3a43ae6ba5b"
image_size = 224
print("\nTrain Batches: ")
train_batches = datagen.flow_from_directory(directory=train_path,
target_size=(image_size,image_size),
batch_size=batch_size,
shuffle=True)
print("\nTest Batches: ")
test_batches =datagen.flow_from_directory(test_path,
target_size=(image_size,image_size),
batch_size=batch_size,
shuffle=False)
# + colab={"base_uri": "https://localhost:8080/"} id="VrlJwba5By1A" outputId="1735d18d-bc75-472c-b83d-1a2bb050d853"
resnet = tf.keras.applications.ResNet50(
include_top=True,
weights="imagenet",
input_tensor=None,
input_shape=None,
pooling=None,
classes=1000,
)
# Exclude the last 3 layers of the model.
conv = resnet.layers[-3].output
# + id="R13YR5JxVpOg"
output = GlobalAveragePooling2D()(conv)
output = Dense(7, activation='softmax')(output)
model = Model(inputs=resnet.input, outputs=output)
# + colab={"base_uri": "https://localhost:8080/"} id="BD2GGKGNV23W" outputId="cd97c665-9eef-4165-ec1a-8c6a7f0392ab"
model.summary()
# + id="WR0fUpy18vAZ"
opt1=tf.keras.optimizers.Adam(learning_rate=0.01,epsilon=0.1)
model.compile(optimizer=opt1,
loss='categorical_crossentropy',
metrics=['accuracy'])
# + id="LAf5ha295reS"
class_weights = {
0: 1.0, # akiec
1: 1.0, # bcc
2: 1.0, # bkl
3: 1.0, # df
4: 5.0, # mel
5: 1.0, # nv
6: 1.0, # vasc
}
checkpoint= ModelCheckpoint(filepath = 'ResNet50.hdf5',monitor='val_accuracy',save_best_only=True,save_weights_only=True)
# + colab={"base_uri": "https://localhost:8080/"} id="NUzTmiZ-8hL3" outputId="46eeb3e9-4356-45d4-910e-1532766e318a"
Earlystop = EarlyStopping(monitor='val_loss', mode='min',patience=40, min_delta=0.001)
history = model.fit(train_batches,
steps_per_epoch=(len(train_df)/10),
epochs=300,
verbose=2,
validation_data=test_batches,validation_steps=len(test_df)/batch_size,callbacks=[checkpoint,Earlystop],class_weight=class_weights)
# + id="zm_AewFBXTj8"
from tensorflow.keras import models
model.load_weights("ResNet50.hdf5")
# + id="LeebvJFEZlAS"
predictions = model.predict(test_batches, steps=len(test_df)/batch_size, verbose=0)
# + colab={"base_uri": "https://localhost:8080/"} id="KYxCDDjusR-S" outputId="64c8e9d3-d9d3-4880-d26a-91414471fb64"
#geting predictions on test dataset
y_pred = np.argmax(predictions, axis=1)
targetnames = ['akiec', 'bcc', 'bkl', 'df', 'mel', 'nv', 'vasc']
#getting the true labels per image
y_true = test_batches.classes
#getting the predicted labels per image
y_prob=predictions
from tensorflow.keras.utils import to_categorical
y_test = to_categorical(y_true)
# Creating classification report
report = classification_report(y_true, y_pred, target_names=targetnames)
print("\nClassification Report:")
print(report)
# + colab={"base_uri": "https://localhost:8080/"} id="yy59Zs1jqylz" outputId="48925225-2fa5-4ed6-9439-6ae2adef1b6a"
print("Precision: "+ str(precision_score(y_true, y_pred, average='weighted')))
print("Recall: "+ str(recall_score(y_true, y_pred, average='weighted')))
print("Accuracy: " + str(accuracy_score(y_true, y_pred)))
print("weighted Roc score: " + str(roc_auc_score(y_test,y_prob,multi_class='ovr',average='weighted')))
# + colab={"base_uri": "https://localhost:8080/"} id="vFRWOB82sDKi" outputId="2813e781-55a8-4203-8ef0-7b94ca628e30"
print("Precision: "+ str(precision_score(y_true, y_pred, average='macro')))
print("Recall: "+ str(recall_score(y_true, y_pred, average='macro')))
print("Accuracy: " + str(accuracy_score(y_true, y_pred)))
print("Macro Roc score: " + str(roc_auc_score(y_test,y_prob,multi_class='ovr',average='macro')))
# + colab={"base_uri": "https://localhost:8080/"} id="nDNAPv9OsRVg" outputId="1f1d443e-020a-4af0-b8ad-01f2fcd7f20a"
print("Precision: "+ str(precision_score(y_true, y_pred, average='micro')))
print("Recall: "+ str(recall_score(y_true, y_pred, average='micro')))
print("Accuracy: " + str(accuracy_score(y_true, y_pred)))
tpr={}
fpr={}
roc_auc={}
fpr["micro"], tpr["micro"], _ = roc_curve(y_test.ravel(), y_prob.ravel())
roc_auc["micro"] = auc(fpr["micro"], tpr["micro"])
print("Micro Roc score: " + str(roc_auc["micro"]))
# + colab={"base_uri": "https://localhost:8080/"} id="U03sRDM2sudx" outputId="235ea641-5f7c-421e-cfa5-e27b99ad0eb8"
fpr = {}
tpr = {}
roc_auc = {}
for i in range(7):
r = roc_auc_score(y_test[:, i], y_prob[:, i])
print("The ROC AUC score of "+targetnames[i]+" is: "+str(r))
# + id="A5nG-b11wkep"
# Compute ROC curve and ROC area for each class
fpr = {}
tpr = {}
roc_auc = dict()
for i in range(7):
fpr[i], tpr[i], _ = roc_curve(y_test[:, i], y_prob[:, i], drop_intermediate=False)
roc_auc[i] = auc(fpr[i], tpr[i])
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="9wz2--WDwHQ4" outputId="4f5ef06b-8812-4a4f-ac8f-53041987d159"
plt.plot(fpr[0], tpr[0],'v-',label='akiec: ROC curve of (area = %0.2f)' % roc_auc[0])
plt.plot(fpr[1], tpr[1],'c',label='bcc: ROC curve of (area = %0.2f)' % roc_auc[1])
plt.plot(fpr[2], tpr[2],'b',label='bkl: ROC curve of (area = %0.2f)' % roc_auc[2])
plt.plot(fpr[3], tpr[3],'g',label='df: ROC curve of (area = %0.2f)' % roc_auc[3])
plt.plot(fpr[4], tpr[4],'y',label='mel: ROC curve of (area = %0.2f)' % roc_auc[4])
plt.plot(fpr[5], tpr[5],'o-',label='nv: ROC curve of (area = %0.2f)' % roc_auc[5])
plt.plot(fpr[6], tpr[6],'r',label='vasc: ROC curve of (area = %0.2f)' % roc_auc[6])
plt.plot([0, 1], [0, 1], 'k--')
plt.xlim([-0.1, 1.1])
plt.ylim([-0.1, 1.1])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Receiver operating characteristic of %s'%targetnames[i])
plt.legend(loc="lower right")
plt.show()
| Skin_Lesion_ResNet50.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Protein Folding Problem
# We can solve the protein folding problem based on some steps:
#
# i. ProteinFoldingProblem function (Convert a protein into qubits)<br>
# ii. VQE algorithm (finding the lowest energy configuration)
from qiskit_nature.problems.sampling.protein_folding.peptide.peptide import Peptide
from qiskit_nature.problems.sampling.protein_folding.interactions.miyazawa_jernigan_interaction import MiyazawaJerniganInteraction
from qiskit_nature.problems.sampling.protein_folding.penalty_parameters import PenaltyParameters
from qiskit_nature.problems.sampling.protein_folding.protein_folding_problem import ProteinFoldingProblem
from qiskit.circuit.library import RealAmplitudes
from qiskit.algorithms.optimizers import COBYLA
from qiskit.utils import QuantumInstance
from qiskit import execute, Aer
from qiskit.opflow import PauliExpectation, CVaRExpectation
from qiskit.algorithms import VQE
# ## i. ProteinFoldingProblem function
# We can define a protein folding function to solve this problem based on:
# 1. Definition of a protein
# 2. The interaction method
# 3. Penalty terms
#
# +
### 1. Defining a protein
main_chain = "APRLRFY" #Alanine,Proline,Arginine,Leucine,Arginine,Phenylalanine,Tyrosine / 7 amino acids
side_chains = [""] * 7 # seven empty side chains = no side chains
protein1 = Peptide(main_chain, side_chains)
### 2. The interaction method
miyazawa_jernigan = MiyazawaJerniganInteraction()
### 3. Penalty terms
penalty_terms = PenaltyParameters(10,10,10) #(penalty_chiral, penalty_back, penalty_1)
### using ProteinFoldingProblem Function
protein_folding_1 = ProteinFoldingProblem(protein1, miyazawa_jernigan, penalty_terms)
qubit_operations_1 = protein_folding_1.qubit_op()
print(qubit_operations_1)
# -
# ## ii. VQE with CVaR expectation
#
# We can use VQE algorithm to solve the problem. First, we should define these parts:
#
# 1. ansatz
# 2. optimizer
# 3. backend
# +
### 1. ansatz
ansatz = RealAmplitudes(reps=1)
### 2. optimizer
optimizer = COBYLA(maxiter=50)
### 3. backend
backend = "aer_simulator"
backend = QuantumInstance(
Aer.get_backend(backend),
shots=1000,
)
### callback function
counts = []
values = []
def callback_function(eval_count, parameters, mean, std):
counts.append(eval_count)
values.append(mean)
### expectation
cvar_exp = CVaRExpectation(0.1, PauliExpectation())
### VQE using CVaR
vqe = VQE(
expectation=cvar_exp,
optimizer=optimizer,
ansatz=ansatz,
quantum_instance=backend,
callback=callback_function,
)
result = vqe.compute_minimum_eigenvalue(qubit_operations_1)
print(result)
# -
# ### Plot the results
# +
import matplotlib.pyplot as plt
figure = plt.figure()
plt.plot(counts, values)
plt.ylabel("Conformation Energy")
plt.xlabel("VQE Iterations")
plt.show()
# -
| ProteinFolding_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="GzkG4LcQt4-1"
# # Introduction to Python 5
#
# ## Modules and Numpy
#
# ## Metodos Computacionales I
# ## Uniandes
#
# Based on [SoloLearn Python](https://www.sololearn.com/Play/Python)
# + [markdown] id="raGL9qCjjX5H"
# ## Pip Install Modules
# + colab={"base_uri": "https://localhost:8080/", "height": 105} executionInfo={"elapsed": 4642, "status": "ok", "timestamp": 1602636112060, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}, "user_tz": 300} id="Hs-MV0QhjW2k" outputId="e5b03c9d-06c4-49f2-973d-628fd02fa2ac"
# To install a library we use a Linux command
# !pip install colorama
# + id="Sct5guZwjjyk"
import colorama
from colorama import Fore, Style
# + id="K9CPjbTwjn0s"
# Fore??
# + colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"elapsed": 569, "status": "ok", "timestamp": 1602636468795, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}, "user_tz": 300} id="U0QzDXxsjrOc" outputId="127b3bf7-52cd-4174-9d40-81e8a5d808ac"
print(Fore.BLUE + "Hello World")
# + executionInfo={"elapsed": 2396, "status": "ok", "timestamp": 1602700583951, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}, "user_tz": 300} id="dClOv_Tsk277"
import tensorflow as tf
from tensorflow.keras.layers import Dense
# + executionInfo={"elapsed": 475, "status": "ok", "timestamp": 1602700600686, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}, "user_tz": 300} id="M3U9D24alZXq"
# Dense??
# + [markdown] id="vXRBKOJll4P8"
# ## Numpy Arrays
# + executionInfo={"elapsed": 535, "status": "ok", "timestamp": 1602700711382, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}, "user_tz": 300} id="2hkxhwAGmAS5"
import numpy as np
# + colab={"base_uri": "https://localhost:8080/", "height": 51} executionInfo={"elapsed": 475, "status": "ok", "timestamp": 1602700741249, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}, "user_tz": 300} id="rFhg_BoRllqh" outputId="e80f50b5-c002-4d32-ce84-0f798977ab4d"
# Creating a numpy array
a = np.array([[1,2,3,4],[5,6,7,8]])
a
# + colab={"base_uri": "https://localhost:8080/", "height": 51} executionInfo={"elapsed": 375, "status": "ok", "timestamp": 1602700821873, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}, "user_tz": 300} id="Rm0bWTALl--2" outputId="6a721ce1-bfc7-4ac2-be28-9858b8effdc5"
# Selecting an element in two diferent ways
print(a[0][1])
print(a[0, 1])
# + colab={"base_uri": "https://localhost:8080/", "height": 85} executionInfo={"elapsed": 498, "status": "ok", "timestamp": 1602700928786, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}, "user_tz": 300} id="dUgXwr_JmM_G" outputId="dd141f83-6440-4e9d-990f-2ee02a0dc37c"
# Attributes of a numpy array
print(a.ndim)
print(a.shape)
print(a.size)
print(type(a))
# + [markdown] id="lddi4VpLmlJY"
# ## Initializing a numpy array
# + colab={"base_uri": "https://localhost:8080/", "height": 51} executionInfo={"elapsed": 510, "status": "ok", "timestamp": 1602700966823, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}, "user_tz": 300} id="LJyYXf1omTg9" outputId="bbdb93e7-8bcf-4c59-f91a-6356d5590a68"
# Array of ones
b = np.ones((2,3))
b
# + colab={"base_uri": "https://localhost:8080/", "height": 51} executionInfo={"elapsed": 475, "status": "ok", "timestamp": 1602700986416, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}, "user_tz": 300} id="MY9YAQhEm7Ai" outputId="85643ee8-de4c-41c4-ff06-9b1a5d2b09ed"
# Array of zeros
c = np.zeros((2,4))
c
# + colab={"base_uri": "https://localhost:8080/", "height": 51} executionInfo={"elapsed": 451, "status": "ok", "timestamp": 1602701042930, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}, "user_tz": 300} id="RXDXbh6zbK4W" outputId="9f58f451-dc0e-4cd2-c43c-54ce2a991b6b"
h = np.full((2,4), 3)
h
# + colab={"base_uri": "https://localhost:8080/", "height": 51} executionInfo={"elapsed": 579, "status": "ok", "timestamp": 1602701073025, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}, "user_tz": 300} id="yI3mbhZpnHo-" outputId="80f36184-37b6-4676-881c-2117a72342f1"
# Array of random numbers
d = np.random.random((2,4))
d
# + colab={"base_uri": "https://localhost:8080/", "height": 85} executionInfo={"elapsed": 457, "status": "ok", "timestamp": 1602701153971, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}, "user_tz": 300} id="A3u-9oBInSzW" outputId="86abc891-fc27-446e-91da-b3716b39d1ad"
# Identity matrix
#g = np.eye(4)
g = np.identity(4)
g
# + [markdown] id="Ah6L5oq_nlas"
# ## Creating ranges in numpy arrays
# + colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"elapsed": 526, "status": "ok", "timestamp": 1602701255427, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}, "user_tz": 300} id="21Uc8RMEbxUA" outputId="0f9c01d9-4764-4514-962c-e0a9a3ae1dc0"
l = range(10)
print([i for i in range(10)])
# + colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"elapsed": 707, "status": "ok", "timestamp": 1602701275082, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}, "user_tz": 300} id="t-xoWO6DnaCP" outputId="526f2db7-3d33-4302-c1c5-7133aae27e44"
# numpy range
j = np.arange(10)
print(j)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"elapsed": 522, "status": "ok", "timestamp": 1602701299021, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}, "user_tz": 300} id="3_yBsXg_n3T7" outputId="ecd25188-a7e9-49ba-9a95-38c13928346d"
# numpy range
i = np.arange(0,10,2)
print(i)
# + colab={"base_uri": "https://localhost:8080/", "height": 119} executionInfo={"elapsed": 674, "status": "ok", "timestamp": 1602701357405, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}, "user_tz": 300} id="W3EIYq22n_cp" outputId="6bf42054-2286-43e6-aa63-9b4841da756c"
# Creating a list with numbers in between
k= np.linspace(0,10,36)
print(k)
# + [markdown] id="d8uE2mJ4oRcS"
# ## Utils Functions
# + colab={"base_uri": "https://localhost:8080/", "height": 102} executionInfo={"elapsed": 744, "status": "ok", "timestamp": 1602703838725, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}, "user_tz": 300} id="KhJOaJK3oHA8" outputId="5a4f58f2-aed5-49b0-a91f-777e5db1b34c"
# Reshaping a matrix
l = np.array([[1,2,3],[4,5,6]])
print(l)
#print(l.shape)
print(l.reshape(3,2))
# + [markdown] id="ZtRNOBeeoj3X"
# ## Slicing numpy arrays
# + colab={"base_uri": "https://localhost:8080/", "height": 85} executionInfo={"elapsed": 587, "status": "ok", "timestamp": 1602701846637, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}, "user_tz": 300} id="9r1sBn51oW5J" outputId="49a0f389-e462-44f3-bd9f-25eb56ee1953"
# Create an array
m = np.arange(8)
print(m)
# Slicing
n = m[::-1]
print(n)
o = m[:5]
print(o)
p = m[6:]
print(p)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"elapsed": 626, "status": "ok", "timestamp": 1602701868412, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}, "user_tz": 300} id="IuHLcrEcpbOe" outputId="ebd5f19c-6b15-4790-889c-0cc2ab8248d1"
# Changing a value of the array
m[4] = 100
print(m)
# + colab={"base_uri": "https://localhost:8080/", "height": 68} executionInfo={"elapsed": 1064, "status": "ok", "timestamp": 1602702329031, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}, "user_tz": 300} id="ZXsrlc3ZpnzN" outputId="fff44b21-e159-4dcb-f9c3-712d6974eb5b"
# Multidimensional slicing
q = np.array([[10,11,12,13],[20,22,23,25]])
print(q)
# # Select some elements
#print(q[0:1,1])
# Select a column
# print(q[:,0])
# # Select a row
print(q[1,:])
# + [markdown] id="FLjC0RJRqacW"
# ## Arithmetic Operations
# + executionInfo={"elapsed": 561, "status": "ok", "timestamp": 1602702354926, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}, "user_tz": 300} id="4PZOpD6nqQXl"
x = np.array([[1,2,3,4],[5,6,7,8]])
y = np.array([[9,10,11,12],[13,14,15,16]])
# + colab={"base_uri": "https://localhost:8080/", "height": 51} executionInfo={"elapsed": 450, "status": "ok", "timestamp": 1602702360981, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}, "user_tz": 300} id="7jOH7SUkqhNI" outputId="13d67060-5f39-4156-8cd8-8b2ffb1c0054"
# Add two matrices
z = x + y
print(z)
# + colab={"base_uri": "https://localhost:8080/", "height": 85} executionInfo={"elapsed": 630, "status": "ok", "timestamp": 1602702465025, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}, "user_tz": 300} id="RUpxRGoHqjKn" outputId="6dae3e8e-6b90-4ffb-f40c-cf6f5433b3e0"
# Numpy functions
a = np.exp(x)
print(a)
b = np.sqrt(x)
print(b)
# + colab={"base_uri": "https://localhost:8080/", "height": 231} executionInfo={"elapsed": 608, "status": "error", "timestamp": 1602702541872, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}, "user_tz": 300} id="h3FGzIdnqw2C" outputId="93560351-fda5-4f0b-a0a8-ddd3b76a421f"
# Matrix multiplication, the shapes do not align for matrix multiplication
c = np.dot(x, y)
print(c)
# + colab={"base_uri": "https://localhost:8080/", "height": 51} executionInfo={"elapsed": 446, "status": "ok", "timestamp": 1602702613218, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}, "user_tz": 300} id="qhxInczArNbQ" outputId="a7d5ed20-dd3b-4327-84fb-117de0358c6d"
# Matrix multiplication
c = np.dot(x, y.T)
print(c)
# + colab={"base_uri": "https://localhost:8080/", "height": 51} executionInfo={"elapsed": 532, "status": "ok", "timestamp": 1602702691342, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}, "user_tz": 300} id="FXgSkjwprecm" outputId="23f006f4-5ce8-402c-df07-e4496cf84384"
# Matrix multiplication
print(x @ y.T)
# + id="rQNS6btKhkj0"
| Notebooks/04 - modules and numpy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **This notebook is an exercise in the [Python](https://www.kaggle.com/learn/python) course. You can reference the tutorial at [this link](https://www.kaggle.com/colinmorris/working-with-external-libraries).**
#
# ---
#
# # Try It Yourself
#
# There are only three problems in this last set of exercises, but they're all pretty tricky, so be on guard!
#
# Run the setup code below before working on the questions.
from learntools.core import binder; binder.bind(globals())
from learntools.python.ex7 import *
print('Setup complete.')
# # 1.
#
# After completing the exercises on lists and tuples, Jimmy noticed that, according to his `estimate_average_slot_payout` function, the slot machines at the Learn Python Casino are actually rigged *against* the house, and are profitable to play in the long run.
#
# Starting with $200 in his pocket, Jimmy has played the slots 500 times, recording his new balance in a list after each spin. He used Python's `matplotlib` library to make a graph of his balance over time:
# Import the jimmy_slots submodule
from learntools.python import jimmy_slots
# Call the get_graph() function to get Jimmy's graph
graph = jimmy_slots.get_graph()
graph
# As you can see, he's hit a bit of bad luck recently. He wants to tweet this along with some choice emojis, but, as it looks right now, his followers will probably find it confusing. He's asked if you can help him make the following changes:
#
# 1. Add the title "Results of 500 slot machine pulls"
# 2. Make the y-axis start at 0.
# 3. Add the label "Balance" to the y-axis
#
# After calling `type(graph)` you see that Jimmy's graph is of type `matplotlib.axes._subplots.AxesSubplot`. Hm, that's a new one. By calling `dir(graph)`, you find three methods that seem like they'll be useful: `.set_title()`, `.set_ylim()`, and `.set_ylabel()`.
#
# Use these methods to complete the function `prettify_graph` according to Jimmy's requests. We've already checked off the first request for you (setting a title).
#
# (Remember: if you don't know what these methods do, use the `help()` function!)
type(graph)
dir(graph)
# +
def prettify_graph(graph):
"""Modify the given graph according to Jimmy's requests: add a title, make the y-axis
start at 0, label the y-axis. (And, if you're feeling ambitious, format the tick marks
as dollar amounts using the "$" symbol.)
"""
graph.set_title("Results of 500 slot machine pulls")
graph.set_ylim(0)
graph.set_ylabel("Balance")
# Complete steps 2 and 3 here
graph = jimmy_slots.get_graph()
prettify_graph(graph)
graph
# -
# **Bonus:** Can you format the numbers on the y-axis so they look like dollar amounts? e.g. $200 instead of just 200.
#
# (We're not going to tell you what method(s) to use here. You'll need to go digging yourself with `dir(graph)` and/or `help(graph)`.)
def prettify_graph(graph):
"""Modify the given graph according to Jimmy's requests: add a title, make the y-axis
start at 0, label the y-axis. (And, if you're feeling ambitious, format the tick marks
as dollar amounts using the "$" symbol.)
"""
graph.set_title("Results of 500 slot machine pulls")
graph.set_ylim(0)
graph.set_ylabel("Balance")
ticks = graph.get_yticks()
new_label = ['${}'.format(int(label)) for label in ticks]
graph.set_yticklabels(new_label)
graph = jimmy_slots.get_graph()
prettify_graph(graph)
graph
# Check your answer (Run this code cell to receive credit!)
q1.solution()
# # 2. <span title="Spicy" style="color: coral">🌶️🌶️</span>
#
# This is a very challenging problem. Don't forget that you can receive a hint!
#
# Luigi is trying to perform an analysis to determine the best items for winning races on the Mario Kart circuit. He has some data in the form of lists of dictionaries that look like...
#
# [
# {'name': 'Peach', 'items': ['green shell', 'banana', 'green shell',], 'finish': 3},
# {'name': 'Bowser', 'items': ['green shell',], 'finish': 1},
# # Sometimes the racer's name wasn't recorded
# {'name': None, 'items': ['mushroom',], 'finish': 2},
# {'name': 'Toad', 'items': ['green shell', 'mushroom'], 'finish': 1},
# ]
#
# `'items'` is a list of all the power-up items the racer picked up in that race, and `'finish'` was their placement in the race (1 for first place, 3 for third, etc.).
#
# He wrote the function below to take a list like this and return a dictionary mapping each item to how many times it was picked up by first-place finishers.
def best_items(racers):
"""Given a list of racer dictionaries, return a dictionary mapping items to the number
of times those items were picked up by racers who finished in first place.
"""
winner_item_counts = {}
for i in range(len(racers)):
# The i'th racer dictionary
racer = racers[i]
# We're only interested in racers who finished in first
if racer['finish'] == 1:
for i in racer['items']:
# Add one to the count for this item (adding it to the dict if necessary)
if i not in winner_item_counts:
winner_item_counts[i] = 0
winner_item_counts[i] += 1
# Data quality issues :/ Print a warning about racers with no name set. We'll take care of it later.
if racer['name'] is None:
print("WARNING: Encountered racer with unknown name on iteration {}/{} (racer = {})".format(
i+1, len(racers), racer['name'])
)
return winner_item_counts
# He tried it on a small example list above and it seemed to work correctly:
sample = [
{'name': 'Peach', 'items': ['green shell', 'banana', 'green shell',], 'finish': 3},
{'name': 'Bowser', 'items': ['green shell',], 'finish': 1},
{'name': None, 'items': ['mushroom',], 'finish': 2},
{'name': 'Toad', 'items': ['green shell', 'mushroom'], 'finish': 1},
]
best_items(sample)
# However, when he tried running it on his full dataset, the program crashed with a `TypeError`.
#
# Can you guess why? Try running the code cell below to see the error message Luigi is getting. Once you've identified the bug, fix it in the cell below (so that it runs without any errors).
#
# Hint: Luigi's bug is similar to one we encountered in the [tutorial](https://www.kaggle.com/colinmorris/working-with-external-libraries) when we talked about star imports.
# + tags=["raises-exception"]
# Import luigi's full dataset of race data
from learntools.python.luigi_analysis import full_dataset
# Fix me!
def best_items(racers):
winner_item_counts = {}
for i in range(len(racers)):
# The i'th racer dictionary
racer = racers[i]
# We're only interested in racers who finished in first
if racer['finish'] == 1:
for item in racer['items']:
# Add one to the count for this item (adding it to the dict if necessary)
if item not in winner_item_counts:
winner_item_counts[i] = 0
winner_item_counts[i] += 1
# Data quality issues :/ Print a warning about racers with no name set. We'll take care of it later.
if racer['name'] is None:
print("WARNING: Encountered racer with unknown name on iteration {}/{} (racer = {})".format(
i+1, len(racers), racer['name'])
)
return winner_item_counts
# Try analyzing the imported full dataset
best_items(full_dataset)
# -
q2.hint()
# Check your answer (Run this code cell to receive credit!)
q2.solution()
# # 3. <span title="A bit spicy" style="color: darkgreen ">🌶️</span>
#
# Suppose we wanted to create a new type to represent hands in blackjack. One thing we might want to do with this type is overload the comparison operators like `>` and `<=` so that we could use them to check whether one hand beats another. e.g. it'd be cool if we could do this:
#
# ```python
# >>> hand1 = BlackjackHand(['K', 'A'])
# >>> hand2 = BlackjackHand(['7', '10', 'A'])
# >>> hand1 > hand2
# True
# ```
#
# Well, we're not going to do all that in this question (defining custom classes is a bit beyond the scope of these lessons), but the code we're asking you to write in the function below is very similar to what we'd have to write if we were defining our own `BlackjackHand` class. (We'd put it in the `__gt__` magic method to define our custom behaviour for `>`.)
#
# Fill in the body of the `blackjack_hand_greater_than` function according to the docstring.
# +
def hand_total(hand):
total = 0
# Count the number of aces and deal with how to apply them at the end.
aces = 0
for card in hand:
if card in ['J', 'Q', 'K']:
total += 10
elif card == 'A':
aces += 1
else:
# Convert number cards (e.g. '7') to ints
total += int(card)
# At this point, total is the sum of this hand's cards *not counting aces*.
# Add aces, counting them as 1 for now. This is the smallest total we can make from this hand
total += aces
# "Upgrade" aces from 1 to 11 as long as it helps us get closer to 21
# without busting
while total + 10 <= 21 and aces > 0:
# Upgrade an ace from 1 to 11
total += 10
aces -= 1
return total
def blackjack_hand_greater_than(hand_1, hand_2):
"""
Return True if hand_1 beats hand_2, and False otherwise.
In order for hand_1 to beat hand_2 the following must be true:
- The total of hand_1 must not exceed 21
- The total of hand_1 must exceed the total of hand_2 OR hand_2's total must exceed 21
Hands are represented as a list of cards. Each card is represented by a string.
When adding up a hand's total, cards with numbers count for that many points. Face
cards ('J', 'Q', and 'K') are worth 10 points. 'A' can count for 1 or 11.
When determining a hand's total, you should try to count aces in the way that
maximizes the hand's total without going over 21. e.g. the total of ['A', 'A', '9'] is 21,
the total of ['A', 'A', '9', '3'] is 14.
Examples:
>>> blackjack_hand_greater_than(['K'], ['3', '4'])
True
>>> blackjack_hand_greater_than(['K'], ['10'])
False
>>> blackjack_hand_greater_than(['K', 'K', '2'], ['3'])
False
"""
total_1 = hand_total(hand_1)
total_2 = hand_total(hand_2)
return total_1 <= 21 and (total_1 > total_2 or total_2 > 21)
# Check your answer
q3.check()
# +
#q3.hint()
#q3.solution()
# -
# # The End
#
# You've finished the Python course. Congrats!
#
# You probably didn't put in all these hours of learning Python just to play silly games of chance, right? If you're interested in applying your newfound Python skills to some data science tasks, we strongly recommend **[this tutorial](https://www.kaggle.com/alexisbcook/titanic-tutorial)**, which will teach you how to make your very first submission to a Kaggle competition.
#
# You can also check out some of our other **[Kaggle Courses](https://www.kaggle.com/learn/overview)**. Some good next steps are:
#
# 1. [Intro to Machine Learning](https://www.kaggle.com/learn/intro-to-machine-learning)
# 2. [Pandas for data manipulation](https://www.kaggle.com/learn/pandas)
# 3. [Data Visualization](https://www.kaggle.com/learn/data-visualization)
#
# Happy Pythoning!
# ---
#
#
#
#
# *Have questions or comments? Visit the [course discussion forum](https://www.kaggle.com/learn/python/discussion) to chat with other learners.*
| Kaggle Course/Course Codes/exercise-working-with-external-libraries.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="AhHzWM5vOn8I"
#
#
# ```
# import torch
# import torch.nn as nn
# import numpy as np
# import pandas as pd
# import matplotlib.pyplot as plt
# ```
#
#
# + id="81KAOOnEOpsZ" executionInfo={"status": "ok", "timestamp": 1604711648281, "user_tz": -540, "elapsed": 3505, "user": {"displayName": "olto col", "photoUrl": "", "userId": "03275196441143151671"}}
import torch
import torch.nn as nn
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# + [markdown] id="zqfXVLz_LtOf"
#
#
# ```
# # Start: 1970-10-01
# # End: 2020-09-31
#
# stock_data = pd.read_csv(
# "/content/drive/My Drive/data/stock_data/^GSCP.csv",
# index_col=0,
# parse_dates=True
# )
#
# stock_data
# ```
#
#
# + id="k2IJ9b9pLrqb" executionInfo={"status": "ok", "timestamp": 1604711648686, "user_tz": -540, "elapsed": 3893, "user": {"displayName": "olto col", "photoUrl": "", "userId": "03275196441143151671"}} outputId="3aff6e7b-b2fa-469e-c47c-e5264b594612" colab={"base_uri": "https://localhost:8080/", "height": 437}
stock_data = pd.read_csv(
"/content/drive/My Drive/data/stock_data/^GSPC.csv",
index_col = 0,
parse_dates=True
)
stock_data
# + [markdown] id="pnqYH2cSPOp2"
#
#
# ```
# stock_data.drop(
# ["Open", "High", "Low", "Close", "Volume"],
# axis="columns",
# inplace=True
# )
#
# stock_data
# ```
#
#
# + id="VgY7eMyxPQI9" executionInfo={"status": "ok", "timestamp": 1604711648687, "user_tz": -540, "elapsed": 3889, "user": {"displayName": "olto col", "photoUrl": "", "userId": "03275196441143151671"}} outputId="4ba77769-3774-4592-c738-c90b26bcf5f1" colab={"base_uri": "https://localhost:8080/", "height": 437}
stock_data.drop(
["Open", "High", "Low", "Close", "Volume"],
axis="columns",
inplace=True
)
stock_data
# + [markdown] id="AfTvt8SxPXb-"
#
#
# ```
# stock_data.plot(figsize=(12, 4))
# ```
#
#
# + id="xxmWcL7GPYt8" executionInfo={"status": "ok", "timestamp": 1604711649270, "user_tz": -540, "elapsed": 4467, "user": {"displayName": "olto col", "photoUrl": "", "userId": "03275196441143151671"}} outputId="2b481a9e-696f-4c72-a30d-f5b99d7311fb" colab={"base_uri": "https://localhost:8080/", "height": 296}
stock_data.plot(figsize=(12, 4))
# + [markdown] id="89ku6_qAP5_l"
#
#
# ```
# # Convert a feature into a one-dimensional Numpy Array
# y = stock_data["Adj Close"].values
# y
# ```
#
#
# + id="k0f48y34q-z8" executionInfo={"status": "ok", "timestamp": 1604711649270, "user_tz": -540, "elapsed": 4462, "user": {"displayName": "olto col", "photoUrl": "", "userId": "03275196441143151671"}} outputId="02d5dcb8-bd11-434b-fc39-cb4e839174e9" colab={"base_uri": "https://localhost:8080/"}
y = stock_data["Adj Close"].values
y
# + [markdown] id="-QT2HXZlQUMU"
#
#
# ```
# # Normalization:
# from sklearn.preprocessing import MinMaxScaler
# ```
#
#
# + id="NA5Fr0-mrAgu" executionInfo={"status": "ok", "timestamp": 1604711649668, "user_tz": -540, "elapsed": 4859, "user": {"displayName": "olto col", "photoUrl": "", "userId": "03275196441143151671"}}
from sklearn.preprocessing import MinMaxScaler
# + [markdown] id="-R6teByhQf4D"
#
#
# ```
# # Converts a one-dimensional Numpy Array to a two-dimensional Numpy Array
# scaler = MinMaxScaler(feature_range=(-1, 1))
# scaler.fit(y.reshape(-1, 1))
# y = scaler.transform(y.reshape(-1, 1))
# y
# ```
#
#
# + id="TBjWj9dltpgg" executionInfo={"status": "ok", "timestamp": 1604711649668, "user_tz": -540, "elapsed": 4853, "user": {"displayName": "olto col", "photoUrl": "", "userId": "03275196441143151671"}} outputId="598b7df9-72b9-4feb-d18a-28e2f85c0b91" colab={"base_uri": "https://localhost:8080/"}
scaler = MinMaxScaler(feature_range=(-1, 1))
scaler.fit(y.reshape(-1, 1))
y = scaler.transform(y.reshape(-1, 1))
y
# + [markdown] id="VONKxqqrQ0FP"
#
#
# ```
# # Convert a two-dimensional Numpy Array to a one-dimensional Pytorch Tensor
# y = torch.FloatTensor(y).view(-1)
# y
# ```
#
#
# + id="zFccyligmN6y" executionInfo={"status": "ok", "timestamp": 1604711650179, "user_tz": -540, "elapsed": 5360, "user": {"displayName": "olto col", "photoUrl": "", "userId": "03275196441143151671"}} outputId="e84485ff-0222-4268-e043-822de88afa39" colab={"base_uri": "https://localhost:8080/"}
y = torch.FloatTensor(y).view(-1)
y
# + [markdown] id="vLjRCVhRhaoI"
#
#
# ```
# # Separate normalized data for training and testing
# test_size = 24
#
# train_seq = y[:-test_size]
# test_seq = y[-test_size:]
# ```
#
#
# + id="WHGPglJRmfGL" executionInfo={"status": "ok", "timestamp": 1604711650179, "user_tz": -540, "elapsed": 5359, "user": {"displayName": "olto col", "photoUrl": "", "userId": "03275196441143151671"}}
test_size = 24
train_seq = y[:-test_size]
test_seq = y[-test_size:]
# + [markdown] id="a8WqFINwm1-o"
#
#
# ```
# # Plot y, train_seq and test_seq
# plt.figure(figsize=(12, 4))
# plt.xlim(-20, len(y)+20)
# plt.grid(True)
# plt.plot(y)
# ```
#
#
# + id="aRdNZxtrmeWL" executionInfo={"status": "ok", "timestamp": 1604711650180, "user_tz": -540, "elapsed": 5356, "user": {"displayName": "olto col", "photoUrl": "", "userId": "03275196441143151671"}} outputId="c1e7f50f-3d5b-4f8c-9ae1-adcbaf8e3dfe" colab={"base_uri": "https://localhost:8080/", "height": 282}
plt.figure(figsize=(12, 4))
plt.xlim(-20, len(test_seq)+20)
plt.grid(True)
plt.plot(test_seq)
# + [markdown] id="yx9ooXxgZFLs"
#
#
# ```
# train_window_size = 12
# ```
#
#
# + id="D_y-rhTAmiia" executionInfo={"status": "ok", "timestamp": 1604711650180, "user_tz": -540, "elapsed": 5354, "user": {"displayName": "olto col", "photoUrl": "", "userId": "03275196441143151671"}}
train_window_size = 12
# + [markdown] id="bXNO9NZaRREX"
#
#
# ```
# def input_data(seq, ws):
# out = []
# L = len(seq)
#
# for i in range(L-ws):
# window = seq[i:i+ws]
# label = seq[i+ws:i+ws+1]
# out.append((window, label))
#
# return out
# ```
#
#
# + id="W7UbcxzTvtzM" executionInfo={"status": "ok", "timestamp": 1604711650181, "user_tz": -540, "elapsed": 5354, "user": {"displayName": "olto col", "photoUrl": "", "userId": "03275196441143151671"}}
def input_data(seq, ws):
out = []
L = len(seq)
for i in range(L-ws):
window = seq[i:i+ws]
label = seq[i+ws:i+ws+1]
out.append((window, label))
return out
# + [markdown] id="EHkkBGz0RVC5"
#
#
# ```
# train_data = input_data(train_seq, train_window_size)
# ```
#
#
# + id="KNDizU0zvvn5" executionInfo={"status": "ok", "timestamp": 1604711650181, "user_tz": -540, "elapsed": 5352, "user": {"displayName": "olto col", "photoUrl": "", "userId": "03275196441143151671"}}
train_data = input_data(train_seq, train_window_size)
# + [markdown] id="upqk87KJRnH2"
#
#
# ```
# print("The Number of Training Data: ", len(train_data))
# ```
#
#
# + id="yRItxzlOvyO3" executionInfo={"status": "ok", "timestamp": 1604711650181, "user_tz": -540, "elapsed": 5348, "user": {"displayName": "olto col", "photoUrl": "", "userId": "03275196441143151671"}} outputId="e6e3ad66-7a50-450a-9e88-7bbbda9087b7" colab={"base_uri": "https://localhost:8080/"}
# 600-24-12=564
print("The Nunber of Training Data: ", len(train_data))
# + [markdown] id="u9TaCjHqR4Po"
#
#
# ```
# class Model(nn.Module):
#
#
# def __init__(self, input=1, h=50, output=1):
#
# super().__init__()
# self.hidden_size = h
#
# self.lstm = nn.LSTM(input, h)
# self.fc = nn.Linear(h, output)
#
# self.hidden = (
# torch.zeros(1, 1, h),
# torch.zeros(1, 1, h)
# )
#
#
# def forward(self, seq):
#
# out, _ = self.lstm(
# seq.view(len(seq), 1, -1),
# self.hidden
# )
#
# out = self.fc(
# out.view(len(seq), -1)
# )
#
# return out[-1]
# ```
#
#
# + id="y8VVV2nnwEn-" executionInfo={"status": "ok", "timestamp": 1604711650182, "user_tz": -540, "elapsed": 5347, "user": {"displayName": "olto col", "photoUrl": "", "userId": "03275196441143151671"}}
class Model(nn.Module):
def __init__(self, input=1, h=50, output=1):
super().__init__()
self.hidden_size = h
self.lstm = nn.LSTM(input, h)
self.fc = nn.Linear(h, output)
self.hidden = (
torch.zeros(1, 1, h),
torch.zeros(1, 1, h)
)
def forward(self, seq):
out, _ = self.lstm(
seq.view(len(seq), 1, -1),
self.hidden
)
out = self.fc(
out.view(len(seq), -1)
)
return out[-1]
# + [markdown] id="7U9yzbavR-v3"
#
#
# ```
# torch.manual_seed(123)
# model = Model()
# # mean squared error loss 平均二乗誤差損失
# criterion = nn.MSELoss()
# # stochastic gradient descent 確率的勾配降下
# optimizer = torch.optim.SGD(model.parameters(), lr=0.01)
# ```
#
#
# + id="qrNo0p6OwGOD" executionInfo={"status": "ok", "timestamp": 1604711650182, "user_tz": -540, "elapsed": 5345, "user": {"displayName": "olto col", "photoUrl": "", "userId": "03275196441143151671"}}
torch.manual_seed(123)
model = Model()
criterion = nn.MSELoss()
optimizer = torch.optim.SGD(model.parameters(), lr=0.01)
# + [markdown] id="pNhKgh-jSC9K"
#
#
# ```
# epochs = 10
# train_losses = []
# test_losses = []
# ```
#
#
# + id="8KQyl1ahwITd" executionInfo={"status": "ok", "timestamp": 1604711650182, "user_tz": -540, "elapsed": 5343, "user": {"displayName": "olto col", "photoUrl": "", "userId": "03275196441143151671"}}
epochs = 10
train_losses = []
test_losses = []
# + [markdown] id="M3SwAdHFYWYC"
#
#
# ```
# def run_train():
# model.train()
#
# for train_window, correct_label in train_data:
#
# optimizer.zero_grad()
#
# model.hidden = (
# torch.zeros(1, 1, model.hidden_size),
# torch.zeros(1, 1, model.hidden_size)
# )
#
# train_predicted_label = model.forward(train_window)
# train_loss = criterion(train_predicted_label, correct_label)
#
# train_loss.backward()
# optimizer.step()
#
# train_losses.append(train_loss)
# ```
#
#
# + id="FLfKjJ1ywJ6q" executionInfo={"status": "ok", "timestamp": 1604711650183, "user_tz": -540, "elapsed": 5342, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "03275196441143151671"}}
def run_train():
model.train()
for train_window, correct_label in train_data:
optimizer.zero_grad()
model.hidden = (
torch.zeros(1, 1, model.hidden_size),
torch.zeros(1, 1, model.hidden_size)
)
train_predicted_label = model.forward(train_window)
train_loss = criterion(train_predicted_label, correct_label)
train_loss.backward()
optimizer.step()
train_losses.append(train_loss)
# + [markdown] id="m1629zz39ZKB"
#
#
# ```
# # Extract the value of an element from a one-dimensional Tensor with a single element
# a = torch.tensor([3])
# a
# # a.item()
# ```
#
#
# + id="4h582NPewM30" executionInfo={"status": "ok", "timestamp": 1604711650183, "user_tz": -540, "elapsed": 5336, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "03275196441143151671"}} outputId="bd4112c5-2781-45ef-c49b-cff5c9937ba1" colab={"base_uri": "https://localhost:8080/"}
a = torch.tensor([3])
a.item()
# + [markdown] id="3Y37jq2XYht0"
#
#
# ```
# def run_test():
# model.eval()
#
# for i in range(test_size):
#
# test_window = torch.FloatTensor(extending_seq[-test_size:])
#
#
# # print()
# # print("The Length of Extending Sequence: ", len(extending_seq))
# # print("The Length of window", len(test_window))
# # print()
#
#
# # Stop storing parameters by not computing the slope, so as not to consume memory
# with torch.no_grad():
#
# model.hidden = (
# torch.zeros(1, 1, model.hidden_size),
# torch.zeros(1, 1, model.hidden_size)
# )
#
# test_predicted_label = model.forward(test_window)
# extending_seq.append(test_predicted_label.item())
#
# test_loss = criterion(
# torch.FloatTensor(extending_seq[-test_size:]),
# y[len(y)-test_size:]
# )
#
# test_losses.append(test_loss)
# ```
#
#
# + id="wz9BjkmZwOkF" executionInfo={"status": "ok", "timestamp": 1604711650183, "user_tz": -540, "elapsed": 5335, "user": {"displayName": "olto col", "photoUrl": "", "userId": "03275196441143151671"}}
def run_test():
model.eval()
for i in range(test_size):
test_window = torch.FloatTensor(extending_seq[-test_size:])
with torch.no_grad():
model.hidden = (
torch.zeros(1, 1, model.hidden_size),
torch.zeros(1, 1, model.hidden_size)
)
test_predicted_label = model.forward(test_window)
extending_seq.append(test_predicted_label.item())
test_loss = criterion(
torch.FloatTensor(extending_seq[-test_size:]),
y[len(y)-test_size:]
)
test_losses.append(test_loss)
# + [markdown] id="fJBA9IUL0Qh7"
#
#
# ```
# train_seq[-test_size:]
# ```
#
#
# + id="7Iauk-ALwQU7" executionInfo={"status": "ok", "timestamp": 1604711650635, "user_tz": -540, "elapsed": 5780, "user": {"displayName": "olto col", "photoUrl": "", "userId": "03275196441143151671"}} outputId="aed573b3-bb68-46df-afdf-41fe11e37ab5" colab={"base_uri": "https://localhost:8080/"}
train_seq[-test_size:]
# + [markdown] id="LmEFvWCjxfF-"
#
#
# ```
# train_seq[-test_size:].tolist()
# ```
#
#
# + id="0XO6oolswSZm" executionInfo={"status": "ok", "timestamp": 1604711650635, "user_tz": -540, "elapsed": 5775, "user": {"displayName": "olto col", "photoUrl": "", "userId": "03275196441143151671"}} outputId="f7ebcf27-1115-4371-d9f4-04dddaa4fbd0" colab={"base_uri": "https://localhost:8080/"}
train_seq[-test_size:].tolist()
# + [markdown] id="VrBD4Y6yYqBV"
#
#
# ```
# for epoch in range(epochs):
#
# print()
# print(f'Epoch: {epoch+1}')
#
# run_train()
#
# extending_seq = train_seq[-test_size:].tolist()
#
# run_test()
#
# plt.figure(figsize=(12, 4))
# plt.xlim(-20, len(y)+20)
# plt.grid(True)
#
# plt.plot(y.numpy())
#
# plt.plot(
# range(len(y)-test_size, len(y)),
# extending_seq[-test_size:]
# )
#
# plt.show()
# ```
#
#
# + id="_krXgg9rwV9w" executionInfo={"status": "ok", "timestamp": 1604711672357, "user_tz": -540, "elapsed": 27492, "user": {"displayName": "olto col", "photoUrl": "", "userId": "03275196441143151671"}} outputId="c0058c58-6343-4f1e-b0af-55c3186bdaa6" colab={"base_uri": "https://localhost:8080/", "height": 1000}
for epoch in range(epochs):
print()
print(f'Epoch: {epoch+1}')
run_train()
extending_seq = train_seq[-test_size:].tolist()
run_test()
plt.figure(figsize=(12, 4))
plt.xlim(-20, len(y)+20)
plt.grid(True)
plt.plot(y.numpy())
plt.plot(
range(len(y)-test_size, len(y)),
extending_seq[-test_size:]
)
plt.show()
# + [markdown] id="EvEKb4rcTJtp"
#
#
# ```
# plt.plot(train_losses)
# ```
#
#
# + id="W3OU6nTTwb8S" executionInfo={"status": "ok", "timestamp": 1604711672358, "user_tz": -540, "elapsed": 27488, "user": {"displayName": "olto col", "photoUrl": "", "userId": "03275196441143151671"}} outputId="d569edf9-d31d-4cf5-8d86-d6109083363d" colab={"base_uri": "https://localhost:8080/", "height": 282}
plt.plot(train_losses)
# + [markdown] id="NFw7mUShTMB0"
#
#
# ```
# plt.plot(test_losses)
# ```
#
#
# + id="W4fpWYftwdSu" executionInfo={"status": "ok", "timestamp": 1604711672358, "user_tz": -540, "elapsed": 27483, "user": {"displayName": "olto col", "photoUrl": "", "userId": "03275196441143151671"}} outputId="a136a384-69b4-40cb-cb2d-25b89fca01ca" colab={"base_uri": "https://localhost:8080/", "height": 282}
plt.plot(test_losses)
# + [markdown] id="mF7ZLg06SRtx"
#
#
# ```
# # List
# predicted_normalized_labels_list = extending_seq[-test_size:]
# predicted_normalized_labels_list
# ```
#
#
# + id="3skli4Z6whoi" executionInfo={"status": "ok", "timestamp": 1604711672358, "user_tz": -540, "elapsed": 27481, "user": {"displayName": "olto col", "photoUrl": "", "userId": "03275196441143151671"}}
predicted_normalized_labels_list = extending_seq[-test_size:]
# + [markdown] id="mCaHuToHSXIw"
#
#
# ```
# # Convert a list to a one-dimensional Numpy Array
# predicted_normalized_labels_array_1d = np.array(predicted_normalized_labels_list)
# predicted_normalized_labels_array_1d
# ```
#
#
# + id="CIWbQ3HnwjVb" executionInfo={"status": "ok", "timestamp": 1604711672359, "user_tz": -540, "elapsed": 27478, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "03275196441143151671"}} outputId="5bd32e91-a5bf-4a21-fc40-290ee53cab9d" colab={"base_uri": "https://localhost:8080/"}
predicted_normalized_labels_array_1d = np.array(predicted_normalized_labels_list)
predicted_normalized_labels_array_1d
# + [markdown] id="lZbJI1ceSdKZ"
#
#
# ```
# # Converts a one-dimensional Numpy Array to a two-dimensional Numpy Array
# predicted_normalized_labels_array_2d = predicted_normalized_labels_array_1d.reshape(-1, 1)
# predicted_normalized_labels_array_2d
# ```
#
#
# + id="SOtxS0ujwlDH" executionInfo={"status": "ok", "timestamp": 1604711672359, "user_tz": -540, "elapsed": 27473, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "03275196441143151671"}} outputId="3c459d87-6a89-4245-be66-7d47d2a6d32d" colab={"base_uri": "https://localhost:8080/"}
predicted_normalized_labels_array_2d = predicted_normalized_labels_array_1d.reshape(-1, 1)
predicted_normalized_labels_array_2d
# + [markdown] id="0N2-bR0bzmiT"
#
#
# ```
# # From a normalized number to a true number.
# predicted_labels_array_2d = scaler.inverse_transform(predicted_normalized_labels_array_2d)
# predicted_labels_array_2d
# ```
#
#
# + id="yjQC3Etwwm92" executionInfo={"status": "ok", "timestamp": 1604711672360, "user_tz": -540, "elapsed": 27470, "user": {"displayName": "olto col", "photoUrl": "", "userId": "03275196441143151671"}} outputId="dc4a903b-a11f-4caf-9168-062c15de4b26" colab={"base_uri": "https://localhost:8080/"}
predicted_labels_array_2d = scaler.inverse_transform(predicted_normalized_labels_array_2d)
predicted_labels_array_2d
# + [markdown] id="lyyywhh0Db-i"
#
#
# ```
# len(predicted_labels_array_2d)
# ```
#
#
# + id="gQsa4TKgwooK" executionInfo={"status": "ok", "timestamp": 1604711672360, "user_tz": -540, "elapsed": 27465, "user": {"displayName": "olto col", "photoUrl": "", "userId": "03275196441143151671"}} outputId="5f6b0663-b4b1-484e-80b0-f064026eac21" colab={"base_uri": "https://localhost:8080/"}
len(predicted_labels_array_2d)
# + [markdown] id="75jenTK1LeqJ"
#
#
# ```
# stock_data["Adj Close"][-test_size:]
# ```
#
#
# + id="OnjOHIN9wqCs" executionInfo={"status": "ok", "timestamp": 1604711672360, "user_tz": -540, "elapsed": 27461, "user": {"displayName": "olto col", "photoUrl": "", "userId": "03275196441143151671"}} outputId="0b67fcc5-f93c-40b0-80c9-35607caea152" colab={"base_uri": "https://localhost:8080/"}
stock_data["Adj Close"][-test_size:]
# + [markdown] id="-ON2ghHXLqjw"
#
#
# ```
# len(stock_data["Adj Close"][-test_size:])
# ```
#
#
# + id="5c7ZbPFZwrcJ" executionInfo={"status": "ok", "timestamp": 1604711672361, "user_tz": -540, "elapsed": 27458, "user": {"displayName": "olto col", "photoUrl": "", "userId": "03275196441143151671"}} outputId="f0aa88c9-8047-40eb-bb89-db0e044da187" colab={"base_uri": "https://localhost:8080/"}
len(stock_data["Adj Close"][-test_size:])
# + [markdown] id="unjjD1CPPcOU"
#
#
# ```
# stock_data.index
# ```
#
#
# + id="umyhpAOUwtI7" executionInfo={"status": "ok", "timestamp": 1604711672361, "user_tz": -540, "elapsed": 27453, "user": {"displayName": "olto col", "photoUrl": "", "userId": "03275196441143151671"}} outputId="16e2dd5b-f840-483d-e40a-f52a4c6a883a" colab={"base_uri": "https://localhost:8080/"}
stock_data.index
# + [markdown] id="7XPU1LAdDjDL"
#
#
# ```
# # Either way of writing works.
# x_2018_10_to_2020_09 = np.arange('2018-10', '2020-10', dtype='datetime64[M]')
# # x_2018_10_to_2020_09 = np.arange('2018-10-01', '2020-10-31', dtype='datetime64[M]')
#
# x_2018_10_to_2020_09
# ```
#
#
# + id="xFffUWW4wu6q" executionInfo={"status": "ok", "timestamp": 1604711672361, "user_tz": -540, "elapsed": 27448, "user": {"displayName": "olto col", "photoUrl": "", "userId": "03275196441143151671"}} outputId="69a372e8-abda-4381-9d56-5a2ae323ecb8" colab={"base_uri": "https://localhost:8080/"}
x_2018_10_to_2020_09 = np.arange('2018-10', '2020-10', dtype='datetime64[M]')
x_2018_10_to_2020_09
# + [markdown] id="LBBClsvJxRy3"
#
#
# ```
# len(x_2018_10_to_2020_09)
# ```
#
#
# + id="sArfpJfpwwwm" executionInfo={"status": "ok", "timestamp": 1604711672362, "user_tz": -540, "elapsed": 27444, "user": {"displayName": "olto col", "photoUrl": "", "userId": "03275196441143151671"}} outputId="22ef5f7d-3152-4bc7-8bba-7fdb0728a048" colab={"base_uri": "https://localhost:8080/"}
len(x_2018_10_to_2020_09)
# + [markdown] id="E51Z4mD_TX_i"
#
#
# ```
# fig = plt.figure(figsize=(12, 4))
# plt.title('Stock Price Prediction')
# plt.ylabel('Price')
# plt.grid(True)
# plt.autoscale(axis='x', tight=True)
# fig.autofmt_xdate()
#
# plt.plot(stock_data["Adj Close"]['2016-01':])
# plt.plot(x_2018_10_to_2020_09, predicted_labels_array_2d)
# plt.show()
# ```
#
#
# + id="fBt99jM-wyPX" executionInfo={"status": "ok", "timestamp": 1604711673042, "user_tz": -540, "elapsed": 28120, "user": {"displayName": "olto col", "photoUrl": "", "userId": "03275196441143151671"}} outputId="d37d4c59-4bbf-4789-c909-61bfdd4dcf75" colab={"base_uri": "https://localhost:8080/", "height": 279}
fig = plt.figure(figsize=(12, 4))
plt.title('Stock Price Prediction')
plt.ylabel('Price')
plt.grid(True)
plt.autoscale(axis='x', tight=True)
fig.autofmt_xdate()
plt.plot(stock_data["Adj Close"]['2016-01':])
plt.plot(x_2018_10_to_2020_09, predicted_labels_array_2d)
plt.show()
# + [markdown] id="G6ObN0pHTieq"
#
#
# ```
# # Either way of writing works.
# stock_data["Adj Close"]['2018-10':]
# # stock_data["Adj Close"]['2018-10-01':]
# ```
#
#
# + id="KNb-nBIjw1XX" executionInfo={"status": "ok", "timestamp": 1604711673043, "user_tz": -540, "elapsed": 28116, "user": {"displayName": "olto col", "photoUrl": "", "userId": "03275196441143151671"}} outputId="042bcbc5-c762-4aed-f6ac-444f1fe3ed19" colab={"base_uri": "https://localhost:8080/"}
stock_data["Adj Close"]['2018-10':]
# + [markdown] id="SgVymGPoRYrD"
#
#
# ```
# len(stock_data["Adj Close"]['2018-10':])
# ```
#
#
# + id="ehkSX0THw2-H" executionInfo={"status": "ok", "timestamp": 1604711673043, "user_tz": -540, "elapsed": 28111, "user": {"displayName": "olto col", "photoUrl": "", "userId": "03275196441143151671"}} outputId="a1f80d63-5a87-44a1-bd84-11f14533d258" colab={"base_uri": "https://localhost:8080/"}
len(stock_data["Adj Close"]['2018-10':])
# + [markdown] id="YoOYZpf5TrD9"
#
#
# ```
# # Convert to a one-dimensional Numpy Array
# real_labels_array_1d = stock_data["Adj Close"]['2018-10':].values
# real_labels_array_1d
# ```
#
#
# + id="nzArbu-Uw4jH" executionInfo={"status": "ok", "timestamp": 1604711673044, "user_tz": -540, "elapsed": 28107, "user": {"displayName": "olto col", "photoUrl": "", "userId": "03275196441143151671"}} outputId="155eb0fa-39f7-4e11-8e72-14a4a18628ba" colab={"base_uri": "https://localhost:8080/"}
real_labels_array_1d = stock_data["Adj Close"]['2018-10':].values
real_labels_array_1d
# + [markdown] id="KqZC7R_eT6I_"
#
#
# ```
# # Two-Dimensional Numpy Array
# predicted_labels_array_2d
# ```
#
#
# + id="d6eD4z2Pw6Gb" executionInfo={"status": "ok", "timestamp": 1604711673045, "user_tz": -540, "elapsed": 28103, "user": {"displayName": "olto col", "photoUrl": "", "userId": "03275196441143151671"}} outputId="3e5b3603-ea98-47e4-d13c-e3546e91df24" colab={"base_uri": "https://localhost:8080/"}
predicted_labels_array_2d
# + [markdown] id="kQw2k9suT_8a"
#
#
# ```
# # Converts a two-dimensional Numpy Array to a one-dimensional Numpy Array
# predicted_labels_array_1d = predicted_labels_array_2d.flatten()
# predicted_labels_array_1d
# ```
#
#
# + id="lOhWb7cWw8bB" executionInfo={"status": "ok", "timestamp": 1604711673045, "user_tz": -540, "elapsed": 28099, "user": {"displayName": "olto col", "photoUrl": "", "userId": "03275196441143151671"}} outputId="462a8c08-f010-45ce-865c-9b1858874c80" colab={"base_uri": "https://localhost:8080/"}
predicted_labels_array_1d = predicted_labels_array_2d.flatten()
predicted_labels_array_1d
# + [markdown] id="ijiWL9sDUIR3"
#
#
# ```
# len(predicted_labels_array_1d)
# ```
#
#
# + id="in7QeL6hw-Tw" executionInfo={"status": "ok", "timestamp": 1604711673045, "user_tz": -540, "elapsed": 28095, "user": {"displayName": "olto col", "photoUrl": "", "userId": "03275196441143151671"}} outputId="938e1937-8a00-4ce0-8524-7ab216191410" colab={"base_uri": "https://localhost:8080/"}
len(predicted_labels_array_1d)
# + [markdown] id="uF1dlRx9UMpt"
#
#
# ```
# up_and_down_list = []
#
# for i in range(len(real_labels_array_1d)):
# difference = real_labels_array_1d[i] - predicted_labels_array_1d[i]
# up_and_down.append(difference)
#
# up_and_down_list
# ```
#
#
# + id="DZYiHCHJxAJ3" executionInfo={"status": "ok", "timestamp": 1604711673046, "user_tz": -540, "elapsed": 28091, "user": {"displayName": "olto col", "photoUrl": "", "userId": "03275196441143151671"}} outputId="3dd78c60-63b4-41a2-a6c6-ca449fb07897" colab={"base_uri": "https://localhost:8080/"}
up_and_down_list = []
for i in range(len(real_labels_array_1d)):
difference = real_labels_array_1d[i] - predicted_labels_array_1d[i]
up_and_down_list.append(difference)
up_and_down_list
# + [markdown] id="BblefX9RUVDE"
#
#
# ```
# up_and_down_array_1d = np.array(up_and_down_list)
# up_and_down_array_1d
# ```
#
#
# + id="4m7JMMZhxBti" executionInfo={"status": "ok", "timestamp": 1604711673046, "user_tz": -540, "elapsed": 28087, "user": {"displayName": "olto col", "photoUrl": "", "userId": "03275196441143151671"}} outputId="36be1985-8f3e-4e09-e809-536c8f6852bd" colab={"base_uri": "https://localhost:8080/"}
up_and_down_array_1d = np.array(up_and_down_list)
up_and_down_array_1d
# + [markdown] id="pAW0avLYUgiN"
#
#
# ```
# plt.plot(up_and_down_array_1d)
# ```
#
#
# + id="_LJ2TsZXxDMN" executionInfo={"status": "ok", "timestamp": 1604711673047, "user_tz": -540, "elapsed": 28084, "user": {"displayName": "olto col", "photoUrl": "", "userId": "03275196441143151671"}} outputId="db5b7cf0-4245-4f1d-b080-c52dbc6dd797" colab={"base_uri": "https://localhost:8080/", "height": 282}
plt.plot(up_and_down_array_1d)
# + id="gvdXGsfzvA9H" executionInfo={"status": "ok", "timestamp": 1604711673047, "user_tz": -540, "elapsed": 28082, "user": {"displayName": "olto col", "photoUrl": "", "userId": "03275196441143151671"}}
| Udemy/StockPrediction/google_colab/1-1_sp500_monthly_with_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + tags=["active-ipynb", "remove-input", "remove-output"]
# try:
# import openmdao.api as om
# import dymos as dm
# except ImportError:
# !python -m pip install openmdao[notebooks]
# !python -m pip install dymos[docs]
# import openmdao.api as om
# import dymos as dm
# -
# # The Brachistochrone
#
# ```{admonition} Things you'll learn through this example
# - How to define a basic Dymos ODE system.
# - How to test the partials of your ODE system.
# - Adding a Trajectory object with a single Phase to an OpenMDAO Problem.
# - Imposing boundary conditions on states with simple bounds via `fix_initial` and `fix_final`.
# - Using the Phase.interpolate` method to set a linear guess for state and control values across the Phase.
# - Checking the validity of the result through explicit simulation via the `Trajectory.simulate` method.
# ```
#
# The brachistochrone is one of the most well-known optimal control problems.
# It was originally posed as a challenge by <NAME>.
#
# ```{admonition} The brachistochrone problem
# _Given two points A and B in a vertical plane, find the path AMB
# down which a movable point M must by virtue of its weight fall from
# A to B in the shortest possible time._
#
# - <NAME>, <NAME>, June 1696
# ```
#
# We seek to find the optimal shape of a wire between two points (A and B) such that a bead sliding
# without friction along the wire moves from point A to point B in minimum time.
# + tags=["remove-input"]
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.patches import FancyArrowPatch, Arc
LW = 2
fig, ax = plt.subplots(1, 1, figsize=(5, 5))
ax.axis('off')
ax.set_xlim(-1, 11)
ax.set_ylim(-1, 11)
circle = plt.Circle((0, 10), radius=0.1, fc='k')
ax.add_patch(circle)
plt.text(0.2, 10.2, 'A')
circle = plt.Circle((10, 5), radius=0.1, fc='k')
ax.add_patch(circle)
plt.text(10.2, 5.2, 'B')
# Choose a to suite, compute b
a = 0.1
b = -0.5 - 10*a
c = 10
def y_wire(x):
return a*x**2 + b*x + c, 2*a*x + b
x = np.linspace(0, 10, 100)
y, _ = y_wire(x)
plt.plot(x, y, 'b-')
# Add the bead to the wire
x = 3
y, dy_dx = y_wire(x)
plt.plot(x, y, 'ro', ms=10)
# Draw and label the gravity vector
gvec = FancyArrowPatch((x, y), (x, y-2), arrowstyle='->', mutation_scale=10, linewidth=LW, color='k')
lv_line = plt.Line2D((x, x), (y, y-2), visible=False) # Local vertical
ax.add_patch(gvec)
plt.text(x - 0.5, y-1, 'g')
# Draw and label the velocity vector
dx = 2
dy = dy_dx * dx
vvec = FancyArrowPatch((x, y), (x+dx, y+dy), arrowstyle='->', mutation_scale=10, linewidth=LW, color='k')
ax.add_patch(vvec)
plt.text(x+dx-0.25, y+dy-0.25, 'v')
# Draw angle theta
vvec_line = plt.Line2D((x, x+dx), (y, y+dy), visible=False)
# angle_plot = get_angle_plot(lv_line, vvec_line, color='k', origin=(x, y), radius=3)
# ax.add_patch(angle_plot)
ax.text(x+0.25, y-1.25, r'$\theta$')
# Draw the axes
x = 0
y = 2
dx = 5
dy = 0
xhat = FancyArrowPatch((x, y), (x+dx, y+dy), arrowstyle='->', mutation_scale=10, linewidth=LW, color='k')
ax.add_patch(xhat)
plt.text(x+dx/2.0-0.5, y+dy/2.0-0.5, 'x')
dx = 0
dy = 5
yhat = FancyArrowPatch((x, y), (x+dx, y+dy), arrowstyle='->', mutation_scale=10, linewidth=LW, color='k')
ax.add_patch(yhat)
plt.text(x+dx/2.0-0.5, y+dy/2.0-0.5, 'y')
plt.ylim(1, 11)
plt.xlim(-0.5, 10.5)
plt.savefig('brachistochrone_fbd.png')
plt.show()
# -
# ## State variables
#
# In this implementation, three _state_ variables are used to define the configuration of the system at any given instant in time.
#
# - **x**: The horizontal position of the particle at an instant in time.
# - **y**: The vertical position of the particle at an instant in time.
# - **v**: The speed of the particle at an instant in time.
#
# ## System dynamics
#
# From the free-body diagram above, the evolution of the state variables is given by the following ordinary differential equations (ODE).
#
# \begin{align}
# \frac{d x}{d t} &= v \sin(\theta) \\
# \frac{d y}{d t} &= -v \cos(\theta) \\
# \frac{d v}{d t} &= g \cos(\theta)
# \end{align}
#
# ## Control variables
#
# This system has a single control variable.
#
# - **$\theta$**: The angle between the gravity vector and the tangent to the curve at the current instant in time.
#
# ## The initial and final conditions
#
# In this case, starting point **A** is given as _(0, 10)_.
# The point moving along the curve will begin there with zero initial velocity.
#
# The initial conditions are:
#
# \begin{align}
# x_0 &= 0 \\
# y_0 &= 10 \\
# v_0 &= 0
# \end{align}
#
# The end point **B** is given as _(10, 5)_.
# The point will end there, but the velocity at that point is not constrained.
#
# The final conditions are:
#
# \begin{align}
# x_f &= 10 \\
# y_f &= 5 \\
# v_f &= \mathrm{free}
# \end{align}
#
# ## Defining the ODE as an OpenMDAO System
#
# In Dymos, the ODE is an OpenMDAO System (a Component, or a Group of components).
# The following ExplicitComponent computes the state rates for the brachistochrone problem.
#
# More detail on the workings of an ExplicitComponent can be found in the OpenMDAO documentation. In summary:
#
# - **initialize**: Called at setup, and used to define options for the component. **ALL** Dymos ODE components should have the property `num_nodes`, which defines the number of points at which the outputs are simultaneously computed.
# - **setup**: Used to add inputs and outputs to the component, and declare which outputs (and indices of outputs) are dependent on each of the inputs.
# - **compute**: Used to compute the outputs, given the inputs.
# - **compute_partials**: Used to compute the derivatives of the outputs w.r.t. each of the inputs analytically. This method may be omitted if finite difference or complex-step approximations are used, though analytic is recommended.
# +
import numpy as np
import openmdao.api as om
class BrachistochroneODE(om.ExplicitComponent):
def initialize(self):
self.options.declare('num_nodes', types=int)
self.options.declare('static_gravity', types=(bool,), default=False,
desc='If True, treat gravity as a static (scalar) input, rather than '
'having different values at each node.')
def setup(self):
nn = self.options['num_nodes']
# Inputs
self.add_input('v', val=np.zeros(nn), desc='velocity', units='m/s')
if self.options['static_gravity']:
self.add_input('g', val=9.80665, desc='grav. acceleration', units='m/s/s',
tags=['dymos.static_target'])
else:
self.add_input('g', val=9.80665 * np.ones(nn), desc='grav. acceleration', units='m/s/s')
self.add_input('theta', val=np.ones(nn), desc='angle of wire', units='rad')
self.add_output('xdot', val=np.zeros(nn), desc='velocity component in x', units='m/s',
tags=['dymos.state_rate_source:x', 'dymos.state_units:m'])
self.add_output('ydot', val=np.zeros(nn), desc='velocity component in y', units='m/s',
tags=['dymos.state_rate_source:y', 'dymos.state_units:m'])
self.add_output('vdot', val=np.zeros(nn), desc='acceleration magnitude', units='m/s**2',
tags=['dymos.state_rate_source:v', 'dymos.state_units:m/s'])
self.add_output('check', val=np.zeros(nn), desc='check solution: v/sin(theta) = constant',
units='m/s')
# Setup partials
arange = np.arange(self.options['num_nodes'])
self.declare_partials(of='vdot', wrt='theta', rows=arange, cols=arange)
self.declare_partials(of='xdot', wrt='v', rows=arange, cols=arange)
self.declare_partials(of='xdot', wrt='theta', rows=arange, cols=arange)
self.declare_partials(of='ydot', wrt='v', rows=arange, cols=arange)
self.declare_partials(of='ydot', wrt='theta', rows=arange, cols=arange)
self.declare_partials(of='check', wrt='v', rows=arange, cols=arange)
self.declare_partials(of='check', wrt='theta', rows=arange, cols=arange)
if self.options['static_gravity']:
c = np.zeros(self.options['num_nodes'])
self.declare_partials(of='vdot', wrt='g', rows=arange, cols=c)
else:
self.declare_partials(of='vdot', wrt='g', rows=arange, cols=arange)
def compute(self, inputs, outputs):
theta = inputs['theta']
cos_theta = np.cos(theta)
sin_theta = np.sin(theta)
g = inputs['g']
v = inputs['v']
outputs['vdot'] = g * cos_theta
outputs['xdot'] = v * sin_theta
outputs['ydot'] = -v * cos_theta
outputs['check'] = v / sin_theta
def compute_partials(self, inputs, partials):
theta = inputs['theta']
cos_theta = np.cos(theta)
sin_theta = np.sin(theta)
g = inputs['g']
v = inputs['v']
partials['vdot', 'g'] = cos_theta
partials['vdot', 'theta'] = -g * sin_theta
partials['xdot', 'v'] = sin_theta
partials['xdot', 'theta'] = v * cos_theta
partials['ydot', 'v'] = -cos_theta
partials['ydot', 'theta'] = v * sin_theta
partials['check', 'v'] = 1 / sin_theta
partials['check', 'theta'] = -v * cos_theta / sin_theta ** 2
# -
# ```{admonition} "Things to note about the ODE system"
# - There is no input for the position states ($x$ and $y$). The dynamics aren't functions of these states, so they aren't needed as inputs.
# - While $g$ is an input to the system, since it will never change throughout the trajectory, it can be an option on the system. This way we don't have to define any partials w.r.t. $g$.
# - The output `check` is an _auxiliary_ output, not a rate of the state variables. In this case, optimal control theory tells us that `check` should be constant throughout the trajectory, so it's a useful output from the ODE.
# ```
#
# ## Testing the ODE
#
# Now that the ODE system is defined, it is strongly recommended to test the analytic partials before using it in optimization.
# If the partials are incorrect, then the optimization will almost certainly fail.
# Fortunately, OpenMDAO makes testing derivatives easy with the `check_partials` method.
# The `assert_check_partials` method in `openmdao.utils.assert_utils` can be used in test frameworks to verify the correctness of the partial derivatives in a model.
#
# The following is a test method which creates a new OpenMDAO problem whose model contains the ODE class.
# The problem is setup with the `force_alloc_complex=True` argument to enable complex-step approximation of the derivatives.
# Complex step typically produces derivative approximations with an error on the order of 1.0E-16, as opposed to ~1.0E-6 for forward finite difference approximations.
# +
import numpy as np
import openmdao.api as om
num_nodes = 5
p = om.Problem(model=om.Group())
ivc = p.model.add_subsystem('vars', om.IndepVarComp())
ivc.add_output('v', shape=(num_nodes,), units='m/s')
ivc.add_output('theta', shape=(num_nodes,), units='deg')
p.model.add_subsystem('ode', BrachistochroneODE(num_nodes=num_nodes))
p.model.connect('vars.v', 'ode.v')
p.model.connect('vars.theta', 'ode.theta')
p.setup(force_alloc_complex=True)
p.set_val('vars.v', 10*np.random.random(num_nodes))
p.set_val('vars.theta', 10*np.random.uniform(1, 179, num_nodes))
p.run_model()
cpd = p.check_partials(method='cs', compact_print=True)
# + tags=["remove-input", "remove-output"]
from dymos.utils.testing_utils import assert_check_partials
assert_check_partials(cpd)
# -
# ## Solving the Problem
#
# The following script fully defines the brachistochrone problem with Dymos and solves it. In this section we'll walk through each step.
om.display_source("dymos.examples.brachistochrone")
# +
import openmdao.api as om
import dymos as dm
from dymos.examples.plotting import plot_results
from dymos.examples.brachistochrone import BrachistochroneODE
import matplotlib.pyplot as plt
#
# Initialize the Problem and the optimization driver
#
p = om.Problem(model=om.Group())
p.driver = om.ScipyOptimizeDriver()
p.driver.declare_coloring()
#
# Create a trajectory and add a phase to it
#
traj = p.model.add_subsystem('traj', dm.Trajectory())
phase = traj.add_phase('phase0',
dm.Phase(ode_class=BrachistochroneODE,
transcription=dm.GaussLobatto(num_segments=10)))
#
# Set the variables
#
phase.set_time_options(fix_initial=True, duration_bounds=(.5, 10))
phase.add_state('x', fix_initial=True, fix_final=True)
phase.add_state('y', fix_initial=True, fix_final=True)
phase.add_state('v', fix_initial=True, fix_final=False)
phase.add_control('theta', continuity=True, rate_continuity=True,
units='deg', lower=0.01, upper=179.9)
phase.add_parameter('g', units='m/s**2', val=9.80665)
#
# Minimize time at the end of the phase
#
phase.add_objective('time', loc='final', scaler=10)
p.model.linear_solver = om.DirectSolver()
#
# Setup the Problem
#
p.setup()
#
# Set the initial values
#
p['traj.phase0.t_initial'] = 0.0
p['traj.phase0.t_duration'] = 2.0
p.set_val('traj.phase0.states:x', phase.interp('x', ys=[0, 10]))
p.set_val('traj.phase0.states:y', phase.interp('y', ys=[10, 5]))
p.set_val('traj.phase0.states:v', phase.interp('v', ys=[0, 9.9]))
p.set_val('traj.phase0.controls:theta', phase.interp('theta', ys=[5, 100.5]))
#
# Solve for the optimal trajectory
#
dm.run_problem(p)
# Test the results
print(p.get_val('traj.phase0.timeseries.time')[-1])
# + tags=["remove-input", "remove-output"]
from openmdao.utils.assert_utils import assert_near_equal
assert_near_equal(p.get_val('traj.phase0.timeseries.time')[-1], 1.8016, tolerance=1.0E-3)
# +
# Generate the explicitly simulated trajectory
exp_out = traj.simulate()
plot_results([('traj.phase0.timeseries.states:x', 'traj.phase0.timeseries.states:y',
'x (m)', 'y (m)'),
('traj.phase0.timeseries.time', 'traj.phase0.timeseries.controls:theta',
'time (s)', 'theta (deg)')],
title='Brachistochrone Solution\nHigh-Order Gauss-Lobatto Method',
p_sol=p, p_sim=exp_out)
plt.show()
| docs/examples/brachistochrone/brachistochrone.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: geo_dev
# language: python
# name: geo_dev
# ---
# # Data preprocessing
#
# Sometimes we need to preprocess data. `momepy` has a set of tools to edit given geometry to make it fit for morphological analysis.
#
# This guide introduces a selection of tools to preprocess the street network and eliminate unwanted gaps in the network and fix its topology.
# +
import momepy
import geopandas as gpd
from shapely.geometry import LineString
# -
# ## Close gaps
#
# The first issue which may happen is the network which is disconnected. The endpoints do not match. Such a network would result in incorrect results of any graph-based analysis. `momepy.close_gaps` can fix the issue by snapping nearby endpoints to a midpoint between the two.
l1 = LineString([(1, 0), (2, 1)])
l2 = LineString([(2.1, 1), (3, 2)])
l3 = LineString([(3.1, 2), (4, 0)])
l4 = LineString([(4.1, 0), (5, 0)])
l5 = LineString([(5.1, 0), (6, 0)])
df = gpd.GeoDataFrame(['a', 'b', 'c', 'd', 'e'], geometry=[l1, l2, l3, l4, l5])
df.plot(figsize=(10, 10)).set_axis_off()
# All LineStrings above need to be fixed.
df.geometry = momepy.close_gaps(df, .25)
df.plot(figsize=(10, 10)).set_axis_off()
# Now we can compare how the fixed network looks compared to the original one.
ax = df.plot(alpha=.5, figsize=(10, 10))
gpd.GeoDataFrame(geometry=[l1, l2, l3, l4, l5]).plot(ax=ax, color='r', alpha=.5)
ax.set_axis_off()
# ## Remove false nodes
#
# A very common issue is incorrect topology. LineString should end either at road intersections or in dead-ends. However, we often see geometry split randomly along the way. `momepy.remove_false_nodes` can fix that.
#
# We will use `mapclassify.greedy` to highlight each segment.
from mapclassify import greedy
df = gpd.read_file(momepy.datasets.get_path('tests'), layer='broken_network')
df.plot(greedy(df), categorical=True, figsize=(10, 10), cmap="Set3").set_axis_off()
# You can see that the topology of the network above is not as it should be.
#
# For a reference, let's check how many geometries we have now:
len(df)
# Okay, 83 is a starting value. Now let's remove false nodes.
fixed = momepy.remove_false_nodes(df)
fixed.plot(greedy(fixed), categorical=True, figsize=(10, 10), cmap="Set3").set_axis_off()
# From the figure above, it is clear that the network is now topologically correct. How many features are there now?
len(fixed)
# We have been able to represent the same network using 27 features less.
#
# ## Extend lines
#
# In some cases, like in generation of [enclosures](enclosed.ipynb), we may want to close some gaps by extending existing LineStrings until they meet other geometry.
l1 = LineString([(0, 0), (2, 0)])
l2 = LineString([(2.1, -1), (2.1, 1)])
l3 = LineString([(3.1, 2), (4, 0.1)])
l4 = LineString([(3.5, 0), (5, 0)])
l5 = LineString([(2.2, 0), (3.5, 1)])
df = gpd.GeoDataFrame(['a', 'b', 'c', 'd', 'e'], geometry=[l1, l2, l3, l4, l5])
df.plot(figsize=(10, 10)).set_axis_off()
# The situation above is typical. The network is almost connected, but there are gaps. Let's extend geometries and close them. Note that we cannot use `momepy.close_gaps` in this situation as we are nor snapping endpoints to endpoints.
extended = momepy.extend_lines(df, tolerance=.2)
extended.plot(figsize=(10, 10)).set_axis_off()
ax = extended.plot(figsize=(10, 10), color='r')
df.plot(ax=ax)
ax.set_axis_off()
# The figures above are self-explanatory. However, remember that the extended network is not topologically correct and is not suitable for network analysis directly. For `enclosures` it is perfect though.
#
# For more details and further options, see the [API documentation](../../api.rst).
| docs/user_guide/elements/preprocessing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import requests
import json
import string
import hashlib
# +
url_request = 'https://api.codenation.dev/v1/challenge/dev-ps/generate-data?token=<PASSWORD>'
response = requests.get(url_request)
if (200 == response.status_code):
print("Sucesso")
else:
print("ERRO: " + response.text)
# -
jsonString = response.text
json_obj = json.loads(jsonString)
json_obj
def translate(x, n):
x = x - n
while(x > 122):
x = x - 26
while(x < 97):
x = x + 26
return x
def encr(char, num):
x = ord(char)
if(x >= 65 and x <= 90):
return encr(char.toLower(), num)
if(x >= 97 and x <= 122):
return chr(translate(x, num))
return char
# +
quantidade = int(json_obj["numero_casas"])
lista_de_chars = [encr(x, quantidade) for x in json_obj["cifrado"]]
json_obj["decifrado"] = "".join(lista_de_chars)
json_obj
# -
hashe = hashlib.sha1()
hashe.update(json_obj["decifrado"].encode('utf-8'))
json_obj["resumo_criptografico"] = str(hashe.hexdigest())
json_obj
jstr = json.dumps(json_obj)
jstr
File_object = open("answer.json","w")
File_object.write(jstr)
File_object.close()
# +
url = 'https://api.codenation.dev/v1/challenge/dev-ps/submit-solution?token=2447422f85d6d8dceea4439c347f3071eec5f5a8'
files = {'answer': ('answer.json', open('answer.json', 'rb'))}
r = requests.post(url, files=files)
r.text
# +
print("Importando libs e criando metodos... ", end="")
#imports
import requests
import json
import string
import hashlib
#minhas funcoes
def translate(x, n):
x = x - n
while(x > 122):
x = x - 26
while(x < 97):
x = x + 26
return x
def encr(char, num):
x = ord(char)
if(x >= 65 and x <= 90):
return encr(char.toLower(), num)
if(x >= 97 and x <= 122):
return chr(translate(x, num))
return char
print("Ok")
print("Realizando requisição... ", end="")
url_request = 'https://api.codenation.dev/v1/challenge/dev-ps/generate-data?token=2447422f85d6d8dceea4439c347f3071eec5f5a8'
response = requests.get(url_request)
if (200 == response.status_code):
print("Sucesso")
else:
print("ERRO: " + response.text)
print("Decifrando texto... ", end="")
jsonString = response.text
quantidade = int(json_obj["numero_casas"])
lista_de_chars = [encr(x, quantidade) for x in json_obj["cifrado"]]
json_obj["decifrado"] = "".join(lista_de_chars)
print("pronto!")
print("Encriptando com SHA-1... ", end="")
hashe = hashlib.sha1()
hashe.update(json_obj["decifrado"].encode('utf-8'))
json_obj["resumo_criptografico"] = str(hashe.hexdigest())
print("pronto!!")
print("Salvando a J... ", end="")
jstr = json.dumps(json_obj)
File_object = open("answer.json","w")
File_object.write(jstr)
File_object.close()
print("pronto!!!")
print("Enviando essa Joça... ", end="")
url = 'https://api.codenation.dev/v1/challenge/dev-ps/submit-solution?token=2447422f85d6d8dceea4439c347f3071eec5f5a8'
files = {'answer': ('answer.json', open('answer.json', 'rb'))}
resposta = requests.post(url, files=files)
print("pronto!!!\nSua nota de espião: " + resposta.text)
# -
| Challenges/Codenation/Caesar_cipher.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: thesis-venv
# language: python
# name: thesis-venv
# ---
from ipfml import processing
from ipfml.processing import segmentation, transform
from ipfml import utils
from ipfml import metrics
from PIL import Image
from scipy import signal
from skimage import color
import scipy.stats as stats
import seaborn as sns
import cv2
import numpy as np
import matplotlib.pyplot as plt
from numpy.linalg import svd
import os
data_folder = "../dataset"
# # SVD reconstruction analysis on Synthesis Images
# ## Utils functions definition
def compute_images_path(dict_data):
scene = dict_data['name']
prefix = dict_data['prefix']
indices = dict_data['indices']
images_path = []
for index in indices:
path = os.path.join(data_folder, os.path.join(scene, prefix + index + ".png"))
print(path)
images_path.append(path)
return images_path
def get_images_zones(dict_data, images_path):
id_zone = dict_data['zone']
zones_img = []
for path in images_path:
img = Image.open(path)
zones = segmentation.divide_in_blocks(img, (200, 200))
zones_img.append(zones[id_zone])
return zones_img
def display_svd_reconstruction(interval, zones):
output_images = []
begin, end = interval
for zone in zones:
lab_img = transform.get_LAB_L(zone)
lab_img = np.array(lab_img, 'uint8')
U, s, V = svd(lab_img, full_matrices=True)
smat = np.zeros((end-begin, end-begin), dtype=complex)
smat[:, :] = np.diag(s[begin:end])
output_img = np.dot(U[:, begin:end], np.dot(smat, V[begin:end, :]))
print(output_img)
print(np.allclose(lab_img, output_img))
output_img = np.array(output_img, 'uint8')
output_images.append(Image.fromarray(output_img))
return output_images
def display_images(dict_data, rec_images):
indices = dict_data['indices']
scene = dict_data['name']
fig=plt.figure(figsize=(15, 8))
columns = len(zones)
rows = 1
for i in range(1, columns*rows +1):
index = i - 1
fig.add_subplot(rows, columns, i)
plt.imshow(rec_images[index], label=scene + '_' + str(indices[index]))
img_path = 'tmp_images/' + dict_data['prefix'] + 'zone'+ str(current_dict['zone']) + '_reconstruct_' + str(indices[index]) + '.png'
Image.fromarray(np.asarray(rec_images[index], 'uint8')).save(img_path)
plt.show()
def diff_between_images(noisy, ref):
noisy = np.asarray(noisy)
ref = np.asarray(ref)
return ref - noisy
def display_sv_data(dict_data, zones_data, interval, reduced=True):
scene_name = dict_data['name']
image_indices = dict_data['indices']
zone_indice = dict_data['zone']
plt.figure(figsize=(25, 20))
sv_data = []
begin, end = interval
for id_img, zone in enumerate(zones_data):
zone = np.asarray(zone)
print(zone.shape)
U, s, V = metrics.get_SVD(zone)
data = s[begin:end]
label_plt = 'Zone ' + str(zone_indice)
if reduced:
label_plt += ' reduced info '
label_plt += 'of ' + scene_name + '_' + str(image_indices[id_img])
plt.plot(data, label=label_plt)
plt.legend(fontsize=18)
plt.show()
# ## Scenes information data
# +
# start 00020 - ref 00900 - step 10
dict_appart = {'name': 'Appart1opt02',
'prefix': 'appartAopt_',
'indices': ["00050","00250","00400","00550"],
'zone': 9}
# start 00050 - ref 01200 - step 10
dict_cuisine = {'name': 'Cuisine01',
'prefix': 'cuisine01_',
'indices': ["00050", "00400", "01200"],
'zone': 3}
# start 00020 - ref 00950 - step 10
dict_sdb_c = {'name': 'SdbCentre',
'prefix': 'SdB2_',
'indices': ["00020", "00400", "00950"],
'zone': 3}
# start 00020 - ref 00950 - step 10
dict_sdb_d = {'name': 'SdbDroite',
'prefix': 'SdB2_D_',
'indices': ["00020", "00400", "00950"],
'zone': 3}
# -
# ### Definition of parameters
# Here we define parameters for the rest of this study :
# - the scene used
# - the reconstructed interval (give reduced information from SVD decomposition)
# - the displayed interval of SVD values
current_dict = dict_appart
displayed_interval = (50, 200)
reconstructed_interval = (90, 200)
images_path = compute_images_path(current_dict)
# +
zones = get_images_zones(current_dict, images_path)
# save each zone
for id, zone in enumerate(zones):
img_name = current_dict['prefix'] + 'zone'+ str(current_dict['zone']) + '_' + current_dict['indices'][id] + '.png'
zone.save('tmp_images/' + img_name)
Image.fromarray(np.array(transform.get_LAB_L(zones[0]), 'uint8'))
# -
zone = zones[0]
Image.fromarray(np.array(transform.get_LAB_L(zones[0]), 'uint8')).save('tmp_images/initial_image.png')
tuples = [(0,50), (50,200)]
for t in tuples:
begin, end = t
lab_img = transform.get_LAB_L(zone)
lab_img = np.array(lab_img, 'uint8')
U, s, V = svd(lab_img, full_matrices=True)
smat = np.zeros((end-begin, end-begin), dtype=complex)
smat[:, :] = np.diag(s[begin:end])
output_img = np.dot(U[:, begin:end], np.dot(smat, V[begin:end, :]))
print(output_img)
print(np.allclose(lab_img, output_img))
output_img = np.array(output_img, 'uint8')
Image.fromarray(output_img).save('tmp_images/' + str(begin) + '_' + str(end) + '_reconstructed.png')
#output_images.append(Image.fromarray(output_img))
reconstructed_images = display_svd_reconstruction(reconstructed_interval, zones)
# Overview information for each images (with samples : \["00200","00250","00300","00350"\]) reconstructed with reduced information. Images are displayed following this samples list from left to right.
display_images(current_dict, reconstructed_images)
# ## Display of SV values obtained to compare
# ### With reduction of reconstructed images
display_sv_data(current_dict, reconstructed_images, displayed_interval)
# ### Without reduction of information
zones_lab = [Image.fromarray(np.array(metrics.get_LAB_L(z), 'uint8')) for z in zones]
display_sv_data(current_dict, zones_lab, displayed_interval, reduced=False)
# ## Noise mask analysis
# Next part to explore (extraction of noise filter)..
# +
# start 00020 - ref 00900 - step 10
dict_appart = {'name': 'Appart1opt02',
'prefix': 'appartAopt_',
'indices': ["00020", "00900"],
'zone': 3}
# start 00050 - ref 01200 - step 10
dict_cuisine = {'name': 'Cuisine01',
'prefix': 'cuisine01_',
'indices': ["00050", "00400", "01200"],
'zone': 3}
# start 00020 - ref 00950 - step 10
dict_sdb_c = {'name': 'SdbCentre',
'prefix': 'SdB2_',
'indices': ["00020", "00400", "00950"],
'zone': 3}
# start 00020 - ref 00950 - step 10
dict_sdb_d = {'name': 'SdbDroite',
'prefix': 'SdB2_D_',
'indices': ["00020", "00400", "00950"],
'zone': 3}
# -
mask_current_dict = dict_appart
mask_images_path = compute_images_path(mask_current_dict)
mask_zones = get_images_zones(mask_current_dict, mask_images_path)
Image.fromarray(diff_between_images(mask_zones[0], mask_zones[1]))
| analysis/svd_reconstruction_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a id='start'></a>
# # Deep Learning
#
# In questo notebook vengono presentati degli esercizi su come creare una rete neurale deep learning con Keras.
#
# Provate a svolgere il seguente esercizio:<br>
# 1) [Introduzione alle Reti Neurali](#section0)<br>
# 2) [Deep Learning](#section1)<br>
# 3) [Come impara un algoritmo di Deep Learning](#section2)<br>
# 4) [Funzioni di attivazione](#section3)<br>
# 5) [Cataloghiamo Iris](#section4)<br>
# 6) [Riconosciamo la scrittura](#section5)<br>
# <a id='section0'></a>
# ## Introduzione alle Reti Neurali
# In questo corso abbiamo già fatto uso di Reti Neurali senza averne visto nel dettaglio il funzionamento e le caratteristiche.
#
# Un algoritmo di Machine learning può essere visto come una black box che riceve degli input e produce degli output. Per esempio potremmo creare un modello che predice il tempo metereologico di domani, partendo da alcuni dati metereologici sugli ultimi giorni passati.
#
# <img src="img/ml.png" width="50%">
#
# La "black box" è infatti un modello matematico. L'algoritmo di machine learning seguirà una sorta di metodo a prove ed errori per individuare il modello che meglio stimi gli output, dati gli input.
#
# Una volta creato il modello, questo modello deve essere **addestrato**. L'addestramento è il processo attraverso il quale il modello **apprende** come dare un senso ai dati di input.
# <a id='section1-1'></a>
# ### I tipi di reti neurali
# Abbiamo visto nella prima lezione introduttiva che esistono tre tipi di Machine Learning:
#
# 1) **Supervised**: questo metodo viene chiamato supervisionato poiché forniamo all'algoritmo non solo gli input, ma anche gli obiettivi, ovvero gli output desiderati. Questo apprendimento automatico supervisionato è stato al centro di quasi tutti gli esempi che abbiamo visto finora.<br>
# Sulla base di tali informazioni, l'algoritmo impara a produrre output il più vicino possibile agli obiettivi.<br>
# La funzione obiettivo nell'apprendimento supervisionato è chiamata *loss function* (anche costo o errore). Stiamo cercando di ridurre al minimo la perdita in quanto minore è la funzione di perdita, maggiore è la precisione del modello.<br>
# Metodi comuni:
# - Regressione
# - Classificazione
#
#
# 2) **Unsupervised**: nell'apprendimento automatico senza supervisione, il modello viene alimentato con gli input, ma non con obiettivi. Al modello viene invece cheisto di trovare una sorta di dipendenza o logica sottostante nei dati forniti.<br>
# Ad esempio, potremmo avere i dati finanziari per 100 paesi. Il modello riesce a dividerli (cluster) in 5 gruppi. Esaminate quindi i 5 cluster e giungete alla conclusione che i gruppi sono: “Developed”, “Developing but overachieving”, “Developing but underachieving”, “Stagnating”, e “Worsening”.<br>
# L'algoritmo ha diviso i paesi in 5 gruppi in base alle somiglianze dei loro dati osservati, ma non è in grado di sapere quali somiglianze ha ravvisato e quindi quali gruppi sono emersi dalla clusterizzazione.. Potrebbe averli divisi per posizione invece.<br>
# Metodi comuni:
# - Clustering
#
#
# 3) **Reinforcement**: nel ML reinforcement, l'obiettivo dell'algoritmo è massimizzare la sua ricompensa. Questo metodo è ispirato dal comportamento umano e dal modo in cui le persone cambiano le loro azioni in base a degli incentivi: ottenere una ricompensa o evitare la punizione.<br>
# La funzione obiettivo è chiamata funzione di ricompensa, la fase di apprendimento del modello mira a massimizzare la funzione di ricompensa.<br>
# Un esempio è un computer che gioca a <NAME>. Più alto è il punteggio raggiunto, migliore è il rendimento. Il punteggio in questo caso è la funzione obiettivo.<br>
# Metodi comuni:
# - Processo decisionale
# - Sistema di ricompensa
# <a id='section1'></a>
# ## Deep Learning
# Il Deep Learning, la cui traduzione letterale significa apprendimento profondo, è una sottocategoria del Machine Learning (che letteralmente viene tradotto come apprendimento automatico) e indica quella branca dell’Intelligenza Artificiale che fa riferimento agli algoritmi ispirati alla struttura e alla funzione del cervello chiamate reti neurali artificiali.
#
# Il Deep Learning (noto anche come apprendimento strutturato profondo o apprendimento gerarchico) fa parte di una più ampia famiglia di metodi di Machine Learning basati sull’assimilazione di rappresentazioni di dati, al contrario di algoritmi per l’esecuzione di task specifici.
#
# Le architetture di Deep Learning (con le quali oggi si riporta all’attenzione anche del grande pubblico il concetto di rete neurale artificiale) sono per esempio state applicate nella computer vision, nel riconoscimento automatico della lingua parlata, nell’elaborazione del linguaggio naturale, nel riconoscimento audio e nella bioinformatica (l’utilizzo di strumenti informatici per descrivere dal punto di vista numerico e statistico determinati fenomeni biologici come le sequenze di geni, la composizione e la struttura delle proteine, i processi biochimici nelle cellule, ecc.).
#
# Raccogliendo le diverse interpretazioni di alcuni tra i più noti ricercatori e scienziati nel campo dell’apprendimento profondo (quali: <NAME>-<NAME>, fondatore di Google Brain, Chief Scientist di Baidu e professore e Director dell’AI Lab alla Stanford University; <NAME>, ricercatore riconosciuto come uno dei migliori innovatori del mondo – under 35 – dal MIT di Boston; <NAME>, uno degli scienziati più riconosciuti proprio nel campo del Deep Learning; <NAME>, Reasearch Director di OpenAI; <NAME>, una delle figure chiave del Deep Learning e dell’Intelligenza artificiale, primo ricercatore ad aver dimostrato l’uso di un algoritmo di backpropagation generalizzato per l’addestramento di reti neurali multistrato…), potremmo definire il Deep Learning come un sistema che sfrutta una classe di algoritmi di apprendimento automatico che:
#
# 1. usano vari livelli di unità non lineari a cascata per svolgere compiti di estrazione di caratteristiche e di trasformazione. Ciascun livello successivo utilizza l’uscita del livello precedente come input. Gli algoritmi possono essere sia di tipo supervisionato sia non supervisionato e le applicazioni includono l’analisi di pattern (apprendimento non supervisionato) e classificazione (apprendimento supervisionato);
#
# 2. sono basati sull’apprendimento non supervisionato di livelli gerarchici multipli di caratteristiche (e di rappresentazioni) dei dati. Le caratteristiche di più alto livello vengono derivate da quelle di livello più basso per creare una rappresentazione gerarchica;
#
# 3. fanno parte della più ampia classe di algoritmi di apprendimento della rappresentazione dei dati all’interno dell’apprendimento automatico (Machine Learning);
#
# 4. apprendono multipli livelli di rappresentazione che corrispondono a differenti livelli di astrazione; questi livelli formano una gerarchia di concetti.
# <img src="img/Deep-Learning.jpg">
# In generale, il Deep Learning è un metodo di apprendimento automatico che accetta un input X e lo utilizza per prevedere un output di Y. Ad esempio, dati i prezzi delle scorte della settimana passata come input, il mio algoritmo di deep learning proverà a prevedere il prezzo delle azioni del giorno successivo.
#
# Dato un grande set di dati di coppie di input e output, un algoritmo di deep learning proverà a minimizzare la differenza tra la sua previsione e l'output atteso. In questo modo, cerca di apprendere l'associazione / modello tra input e output dati - questo a sua volta consente a un modello di deep learning di generalizzare a input che non ha mai visto prima.
# <img src="img/deep.gif">
# <a id='section2'></a>
# ## Come impara un algoritmo di Deep Learning
# Gli algoritmi di Deep Learning usano qualcosa chiamata rete neurale per trovare associazioni tra un insieme di input e output. La struttura di base è la seguente:
# <img src="img/dl.jpg">
# Una rete neurale è composta da input, hidden e livelli di output, tutti composti da "nodi". I livelli di input comprendono una rappresentazione numerica dei dati (ad esempio immagini con specifiche dei pixel), le previsioni di output dei livelli di output, mentre i livelli nascosti sono correlati con la maggior parte del calcolo.
# Alla base di una rete neurale vi è un processo di regressione e quindi di algebra lineare.
# Le reti neurali si basano principalmente sulla simulazione di neuroni artificiali opportunamente collegati. Il modello rappresentato in figura è quello proposto da McCulloch e Pitts.
#
# <img src="img/neurone.png">
# <img src="img/neuron.png">
# <a id='section3'></a>
# ## Funzioni di attivazione
# Lo scopo di ciascun nodo nella rete neurale feed forward consiste nell’accettare valori di input e immettere un valore di output nello strato immediatamente successivo. I nodi di input inseriscono i valori delle variabili nello strato nascosto senza modificarli.
# Ogni nodo di uno strato nascosto riceve l’input dai nodi connessi dello strato precedente (output del nodo moltiplicato per il peso della connessione), combina tali valori in un valore singolo che utilizza come input della funzione di attivazione.
# La funzione di attivazione deve soddisfare alcuni criteri:
#
# - deve avere valori di output compresi nell’intervallo {0, 1};
# - deve fornire un valore di output vicino ad 1 quando viene sufficientemente stimolata (effetto soglia), per propagare l’attività all’interno della rete (come avviene per i neuroni naturali).
#
# La cosiddetta funzione sigmoide soddisfa entrambi questi criteri ed è calcolata come segue:
#
# $$F(x) = {1 \over {1 + e ^ {-x}}}$$
# Grafico della funzione sigmoide:
# <img src="img/sigmoide.png">
# Altri esempi di funzioni di attivazione sono visibli qui [su WikiPedia](https://en.wikipedia.org/wiki/Activation_function).
# <a id='section4'></a>
# ## Cataloghiamo Iris
# Ricorderete il dataset della vostra scorsa che contiene i fiori Iris:
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.cm as cm
from sklearn.cluster import KMeans
# %matplotlib inline
iris = pd.read_csv('data/iris-with-answers.csv')
# Create a scatter plot based on two corresponding features (sepal_length and sepal_width; OR petal_length and petal_width)
plt.scatter(iris['sepal_length'], iris['sepal_width'])
# Name your axes
plt.xlabel('Lenght of sepal')
plt.ylabel('Width of sepal')
plt.show()
# -
iris.head()
# Possiamo uasre la libreria seaborn per vedere rapidamente quali sono i dati contenuti nel dataset e per produrre dei pairplot:
import seaborn as sns
sns.pairplot(data=iris, hue='species')
plt.show()
# Normalizziamo nuovamente i dati, come fatto a lezione la volta scorsa:
# +
# import some preprocessing module
from sklearn import preprocessing
# scale the data for better results
x_scaled = preprocessing.scale(iris.iloc[:,0:4])
# -
# Ora proviamo, con Keras, a creare un modello in grado di classificare i fiori nelle 3 specie in base alle caratteristiche del sepalo e del petalo:
# +
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
import tensorflow.keras as keras
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, ReLU
from sklearn.preprocessing import StandardScaler, LabelBinarizer
# -
# Selezioniamo le variaibli indipendenti e dipendenti, scaliamole e mappiamo la feature 'species' in modo che possa essere utilizzata come variabile nell'algoritmo di ML:
# +
X = iris[['sepal_length', 'sepal_width', 'petal_length', 'petal_width']]
y = iris['species']
X = StandardScaler().fit_transform(X)
y = LabelBinarizer().fit_transform(y)
# -
# Dividiamo il dataset tra training e learning:
from sklearn.model_selection import train_test_split
x_train, x_test, y_train, y_test = train_test_split(X, y, test_size=0.2)
# Creiamo il modello con Keras aggiungendo i vari layer della nostra rete di deep learning:
# +
model = Sequential()
model.add(Dense(12, input_dim=4))
model.add(ReLU())
model.add(Dense(units=15))
model.add(ReLU())
model.add(Dense(units=8))
model.add(ReLU())
model.add(Dense(units=10))
model.add(ReLU())
model.add(Dense(units=3, activation='softmax'))
model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
history = model.fit(x_train, y_train, epochs=120, validation_data=(x_test, y_test))
# -
# Plottiamo prima le curve della funzione di loss sugli insiemi di training e di test:
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('Loss')
plt.legend(['Train','Test'])
plt.show()
# Quindi plottiamo l'accuratezza del modello, sempre per gli insiemi di trianing e di test:
plt.plot(history.history['accuracy'])
plt.plot(history.history['val_accuracy'])
plt.title("Accuracy")
plt.legend(['Train', 'Test'])
plt.show()
# +
from sklearn.metrics import accuracy_score
preds = model.predict(x_test)
acc = accuracy_score(np.argmax(y_test, axis=1), np.argmax(preds, axis=1))
print("Accuracy: %.2f" % acc)
# -
# Proviamo ora a creare una rete del tutto simile ma usando pyTorch invece che TensorFlow.
#
# Da principio creiamo il modello usando le classi di pyTorch, la rete create sarà del tutto identica a quella creata con TensorFlow nei passaggi precedenti.
# +
import torch
import torch.nn as nn
import torch.nn.functional as F
#build model
class Net(nn.Module):
def __init__(self):
super().__init__()
self.fc1 = nn.Linear(in_features=4, out_features=12)
self.fc2 = nn.Linear(in_features=12, out_features=15)
self.fc3 = nn.Linear(in_features=15, out_features=8)
self.fc4 = nn.Linear(in_features=8, out_features=10)
self.output = nn.Linear(in_features=10, out_features=3)
def forward(self, x):
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = F.relu(self.fc3(x))
x = F.relu(self.fc4(x))
x = self.output(x)
return x
net = Net()
print(net)
criterion = nn.CrossEntropyLoss()
optimizer = torch.optim.Adam(net.parameters(), lr=0.01)
# -
# A questo prunto prepariamo gli input e gli output per essere utilizzati da pyTorch.
#
# Riparticamo dai dati letti dal file csv, mappiamo la label che vogliamo il nostro modello apprenda a prevedere e quindi trasformiamo tutti i dati in tensori.
# +
from sklearn.model_selection import train_test_split
X = iris[['sepal_length', 'sepal_width', 'petal_length', 'petal_width']]
y = iris['species'].map({
'setosa': 0,
'versicolor': 1,
'virginica': 2 })
x_train, x_test, y_train, y_test = train_test_split(X.values, y.values, test_size=0.2)
x_train = torch.FloatTensor(x_train)
x_test = torch.FloatTensor(x_test)
y_train = torch.LongTensor(y_train)
y_test = torch.LongTensor(y_test)
# -
# Siamo ora pronti ad addestrare la nostra rete.
#
# L'addestramento, a differenza di quanto avveniva con Keras, necessita di un po' di passaggi manuali ma, proprio per questo motivo, può essere adattato e customizzato alle nostre esigenze specifiche.
# +
# train the network
num_epoch = 120
history = { 'accuracy': [], 'loss': [], 'val_accuracy': [], 'val_loss': [] }
for epoch in range(num_epoch):
# Facciamo apprendimento sul training e valutiamo i risultati sul test senza applicare
# backpropagation e quindi senza fare ulteriore apprendimento.
y_hat = net.forward(x_train)
with torch.no_grad():
y_test_hat = net.forward(x_test)
# Calcoliamo loss e accuracy sul training
loss = criterion(y_hat, y_train)
y_out = torch.FloatTensor([z.tolist().index(max(z)) for z in y_hat])
acc = torch.sum(y_out == y_train).double() / len(y_train)
# Calcoliamo loss e accuracy sul test
val_loss = criterion(y_test_hat, y_test)
y_test_out = torch.FloatTensor([z.tolist().index(max(z)) for z in y_test_hat])
val_acc = torch.sum(y_test_out == y_test).double() / len(y_test)
# Storicizziamo i valori nella variabile history per plottare i grafici successivamente
history['loss'].append(loss)
history['accuracy'].append(acc)
history['val_loss'].append(val_loss)
history['val_accuracy'].append(val_acc)
# Diamo un output sull'avazamento a console
if epoch % 10 == 0:
print('Epoch: %d - loss: %1.4f - accuracy: %1.4f - val_loss: %1.4f - val_accuracy: %1.4f' % (epoch, loss, acc, val_loss, val_acc))
# Ottimizziamo l'apprendiamento
optimizer.zero_grad()
loss.backward()
optimizer.step()
# -
plt.plot(history['loss'])
plt.plot(history['val_loss'])
plt.title('Loss')
plt.legend(['Train','Test'])
plt.show()
plt.plot(history['accuracy'])
plt.plot(history['val_accuracy'])
plt.title("Accuracy")
plt.legend(['Train', 'Test'])
plt.show()
# +
from sklearn.metrics import accuracy_score
preds = []
with torch.no_grad():
for val in x_test:
y_hat = net.forward(val)
preds.append(y_hat.argmax().item())
acc = accuracy_score(y_test, preds)
print("Accuracy: %.2f" % acc)
# -
# <a id='section5'></a>
# ## Riconosciamo la scrittura
# Creiamo una rete neurale deeplearning che permetta di riconoscere dei numeri scritti a mano:
# +
# We must always import the relevant libraries for our problem at hand. NumPy and TensorFlow are required for this example.
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
import tensorflow.keras as keras
# -
# Una volta importato tensorflow, possiamo quindi iniziare a preparare i nostri dati, modellarli e quindi addestrarli. Per semplicità, utilizzeremo il più comune esempio di "ciao mondo" per l'apprendimento approfondito, che è il set di dati mnist. È un insieme di cifre scritte a mano, da 0 a 9. Sono le immagini 28x28 di queste cifre scritte a mano. Mostreremo un esempio di utilizzo di dati esterni, ma, per ora, carichiamo questi dati:
mnist = tf.keras.datasets.mnist
(x_train, y_train),(x_test, y_test) = mnist.load_data()
# Quando lavori con i tuoi dati raccolti, è probabile che non siano impacchettato così bene. In questo caso è necessario dedicare un poì di tempo e di impegno in questo passaggio.
#
# Quindi i dati x_train sono le "caratteristiche". In questo caso, le funzionalità sono valori in pixel delle immagini 28x28 di queste cifre 0-9. L'y_train è l'etichetta (è un 0,1,2,3,4,5,6,7,8 o un 9?)
#
# Le varianti di test di queste variabili sono gli esempi "fuori campione" che useremo. Questi sono esempi dai nostri dati che metteremo da parte, riservandoli per testare il modello.
#
# Le reti neurali sono eccezionalmente buone per adattarsi ai dati, al punto che generalmente si adattano troppo ai dati. La nostra vera speranza è che la rete neurale non memorizzi solo i nostri dati e che invece "generalizzi" e impari il vero problema e i modelli ad esso associati.
#
# Diamo un'occhiata a questi dati reali:
x_train[0]
# Bene, possiamo povare a visualizzare questa osservazione:
# +
import matplotlib.pyplot as plt
plt.imshow(x_train[0],cmap=plt.cm.binary)
plt.show()
# -
# Generalmente è una buona idea "normalizzare" i tuoi dati. Questo in genere comporta il ridimensionamento dei dati tra 0 e 1, o forse -1 e positivo 1. Nel nostro caso, ogni "pixel" è una caratteristica, e ogni caratteristica attualmente varia da 0 a 255. Non abbastanza da 0 a 1. Facciamo cambiala con una comoda funzione di utilità:
x_train = tf.keras.utils.normalize(x_train, axis=1)
x_test = tf.keras.utils.normalize(x_test, axis=1)
# Ora costruiamo il nostro modello!
model = tf.keras.models.Sequential()
# Un modello sequenziale è quello che userai la maggior parte del tempo. Usiamolo in questo caso.
#
# Ora dobbiamo inserire tutti gli strati. Ricordi la nostra immagine di rete neurale? Lo strato di input era piatto. Quindi, dobbiamo prendere questa immagine 28x28 e renderla una piatta 1x784. Ci sono molti modi per farlo, ma keras ha uno strato di Flatten creato appositamente per noi, quindi lo useremo.
model.add(tf.keras.layers.Flatten())
# Questo servirà come nostro livello di input. Prenderà i dati che gli forniamo e li appiattiremo per noi. Successivamente, vogliamo i nostri livelli nascosti. Andremo con il livello di rete neurale più semplice, che è solo un livello denso. Questo si riferisce al fatto che si tratta di uno strato densamente connesso, il che significa che è "completamente connesso", in cui ogni nodo si collega a ciascun nodo precedente e successivo. Proprio come la nostra immagine.
model.add(tf.keras.layers.Dense(128, activation=tf.nn.relu))
# Questo strato ha 128 unità. La funzione di attivazione è relu, abbreviazione di linear rettificata. Attualmente, relu è la funzione di attivazione che dovrebbe essere predefinita. Ce ne sono molti altri da verificare, ma, se non sai cosa usare, usa relu per iniziare.
#
# Aggiungiamo un altro livello identico per una buona misura.
model.add(tf.keras.layers.Dense(128, activation=tf.nn.relu))
# Ora siamo pronti per un livello di output:
model.add(tf.keras.layers.Dense(10, activation=tf.nn.softmax))
# Questo è il nostro ultimo livello. Ha 10 nodi. 1 nodo per la previsione del numero possibile. In questo caso, la nostra funzione di attivazione è una funzione softmax, poiché in realtà stiamo davvero cercando qualcosa di più simile a una distribuzione di probabilità di quale delle possibili opzioni di previsione questa cosa che stiamo passando attraverso le caratteristiche di è. Grande, il nostro modello è fatto.
#
# Ora dobbiamo "compilare" il modello. Qui è dove passiamo le impostazioni per ottimizzare / allenare effettivamente il modello che abbiamo definito.
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
# Ricorda perché abbiamo scelto relu come funzione di attivazione? La stessa cosa è vera per l'ottimizzatore Adam. È solo un ottimo default per iniziare.
#
# Successivamente, abbiamo la nostra metrica di perdita. La perdita è un calcolo dell'errore. Una rete neurale in realtà non tenta di massimizzare la precisione. Cerca di minimizzare la perdita. Di nuovo, ci sono molte scelte, ma una qualche forma di crossentropia categoriale è un buon inizio per un compito di classificazione come questo.
#
# Ora, facciamo il fit!
history = model.fit(x_train, y_train, epochs=4)
# Mentre ci alleniamo, possiamo vedere la perdita va giù (yay), e la precisione migliora abbastanza rapidamente al 98-99% (doppio yay!)
#
# Questa è la perdita e l'accuratezza dei dati nel campione. Ottenere un'accuratezza elevata e una perdita bassa potrebbe significare che il tuo modello ha imparato come classificare le cifre in generale (è generalizzato) ... o semplicemente memorizzato ogni singolo esempio che hai mostrato (overfit). Questo è il motivo per cui dobbiamo testare dati fuori campione (dati che non abbiamo usato per addestrare il modello).
val_loss, val_acc = model.evaluate(x_test, y_test)
print(val_loss)
print(val_acc)
# Infine, fai previsioni!
predictions = model.predict(x_test)
predictions
# Quel certo non inizia come utile, ma ricorda che queste sono distribuzioni di probabilità. Possiamo ottenere il numero reale semplicemente:
# +
import numpy as np
case_id = 4242
plt.imshow(x_test[case_id], cmap=plt.cm.binary)
plt.show()
print("Il nostro algoritmo legge il numero %d" % np.argmax(predictions[case_id]))
# -
# Anche in questo caso proviamo a ricosturire la stessa rete utilizzando pyTorch invece che TensorFlow:
# +
import torch
import torch.nn as nn
import torch.nn.functional as F
#build model
class Net(nn.Module):
def __init__(self):
super().__init__()
self.fc1 = nn.Linear(in_features=784, out_features=128)
self.fc2 = nn.Linear(in_features=128, out_features=128)
self.output = nn.Linear(in_features=128, out_features=10)
def forward(self, x):
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = self.output(x)
return x
net = Net()
print(net)
criterion = nn.CrossEntropyLoss()
optimizer = torch.optim.Adam(net.parameters(), lr=0.01)
# +
x_train = torch.FloatTensor(x_train)
x_test = torch.FloatTensor(x_test)
y_train = torch.LongTensor(y_train)
y_test = torch.LongTensor(y_test)
x_train = torch.FloatTensor([torch.flatten(e).tolist() for e in x_train])
x_test = torch.FloatTensor([torch.flatten(e).tolist() for e in x_test])
# +
# train the network
num_epoch = 6
history = { 'accuracy': [], 'loss': [], 'val_accuracy': [], 'val_loss': [] }
for epoch in range(num_epoch):
y_hat = net.forward(x_train)
with torch.no_grad():
y_test_hat = net.forward(x_test)
loss = criterion(y_hat, y_train)
y_out = torch.FloatTensor([z.tolist().index(max(z)) for z in y_hat])
acc = torch.sum(y_out == y_train).double() / len(y_train)
val_loss = criterion(y_test_hat, y_test)
y_test_out = torch.FloatTensor([z.tolist().index(max(z)) for z in y_test_hat])
val_acc = torch.sum(y_test_out == y_test).double() / len(y_test)
history['loss'].append(loss)
history['accuracy'].append(acc)
history['val_loss'].append(val_loss)
history['val_accuracy'].append(val_acc)
print('Epoch: %d - loss: %1.4f - accuracy: %1.4f - val_loss: %1.4f - val_accuracy: %1.4f' % (epoch, loss, acc, val_loss, val_acc))
optimizer.zero_grad()
loss.backward()
optimizer.step()
# -
plt.plot(history['loss'])
plt.plot(history['val_loss'])
plt.title('Loss')
plt.legend(['Train','Test'])
plt.show()
plt.plot(history['accuracy'])
plt.plot(history['val_accuracy'])
plt.title("Accuracy")
plt.legend(['Train', 'Test'])
plt.show()
# +
from sklearn.metrics import accuracy_score
preds = []
with torch.no_grad():
for val in x_test:
y_hat = net.forward(val)
preds.append(y_hat.argmax().item())
accuracy_score(y_test, preds)
# -
with torch.no_grad():
torch_predictions = net(x_test)
torch_predictions
# +
import numpy as np
case_id = 23
plt.imshow(torch.reshape(x_test[case_id], (28, 28)), cmap=plt.cm.binary)
plt.show()
print("Il nostro algoritmo legge il numero %d" % np.argmax(torch_predictions[case_id]))
# -
# Eccezionale!
# Con questo esempio abbiamo appena scalfito la superficie di ciò che si può fare con queste librerie.
# Puoi trovare maggiore documentazione qui:
# - [Tensorflow](https://www.tensorflow.org/api_docs/python/)
# - [Keras](https://keras.io/layers/about-keras-layers/)
# - [pyTorch](https://pytorch.org/docs/stable/index.html)
# [Clicca qui per tornare all'inizio della pagina](#start)<a id='start'></a>
| Lezioni/Lezione 7.A - Deep Learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a id='top'></a>
# <h1 style="text-align:center;font-size:200%;;">Real or Not? NLP with Disaster Tweets</h1>
# <img src="https://dataxboost.files.wordpress.com/2018/03/nlp.jpg">
# ## Competition Description
# * Twitter has become an important communication channel in times of emergency.
# The ubiquitousness of smartphones enables people to announce an emergency they’re observing in real-time. Because of this, more agencies are interested in programatically monitoring Twitter (i.e. disaster relief organizations and news agencies).
# <div class="list-group" id="list-tab" role="tablist">
# <h3 class="list-group-item list-group-item-action active" data-toggle="list" role="tab" aria-controls="home">Notebook Content!</h3>
# <a class="list-group-item list-group-item-action" data-toggle="list" href="#libraries" role="tab" aria-controls="profile">Import Libraries<span class="badge badge-primary badge-pill">1</span></a>
# <a class="list-group-item list-group-item-action" data-toggle="list" href="#load" role="tab" aria-controls="messages">Load Data<span class="badge badge-primary badge-pill">2</span></a>
# <a class="list-group-item list-group-item-action" data-toggle="list" href="#visual" role="tab" aria-controls="settings">Visualization of data<span class="badge badge-primary badge-pill">3</span></a>
# <a class="list-group-item list-group-item-action" data-toggle="list" href="#word" role="tab" aria-controls="settings">WordCloud<span class="badge badge-primary badge-pill">4</span></a>
# <a class="list-group-item list-group-item-action" data-toggle="list" href="#clean" role="tab" aria-controls="settings">Cleaning the text<span class="badge badge-primary badge-pill">5</span></a>
# <a class="list-group-item list-group-item-action" data-toggle="list" href="#split" role="tab" aria-controls="settings">Train and test Split<span class="badge badge-primary badge-pill">6</span></a>
# <a class="list-group-item list-group-item-action" data-toggle="list" href="#model" role="tab" aria-controls="settings"> Creating the Model<span class="badge badge-primary badge-pill">7</span></a>
# <a class="list-group-item list-group-item-action" data-toggle="list" href="#eval" role="tab" aria-controls="settings">Model Evaluation<span class="badge badge-primary badge-pill">8</span></a>
# <a id='libraries'></a>
# ## 1. Import Libraries
# + _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19"
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
# Input data files are available in the "../input/" directory.
# For example, running this (by clicking run or pressing Shift+Enter) will list all files under the input directory
import seaborn as sns
import matplotlib.pyplot as plt
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.preprocessing.sequence import pad_sequences
import tensorflow as tf
import time
from wordcloud import WordCloud, STOPWORDS
import matplotlib.pyplot as plt
from IPython.core.display import display, HTML
import plotly.graph_objects as go
import re
# Natural Language Tool Kit
import nltk
nltk.download('stopwords')
from nltk.corpus import stopwords
from nltk.stem.porter import PorterStemmer
from collections import Counter
import cufflinks as cf
cf.go_offline()
# -
# <a id='load'></a>
# # 2. Load Data
# + _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a" _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0"
train = pd.read_csv("/kaggle/input/nlp-getting-started/train.csv")
test = pd.read_csv("/kaggle/input/nlp-getting-started/test.csv")
submission = pd.read_csv("/kaggle/input/nlp-getting-started/sample_submission.csv")
# -
train.head()
# <ul style="list-style-type:square;">
# <li><span class="label label-default">id</span> a unique identifier for each tweet</li>
# <li><span class="label label-default">text </span> the text of the tweet</li>
# <li><span class="label label-default">location</span> the location the tweet was sent from (may be blank)</li>
# <li><span class="label label-default">keyword</span> a particular keyword from the tweet (may be blank)</li>
# <li><span class="label label-default">target</span> in train.csv only, this denotes whether a tweet is about a real disaster (1) or not (0)</li>
# </ul>
#
test.head()
# + _kg_hide-input=true
display(HTML(f"""
<ul class="list-group">
<li class="list-group-item disabled" aria-disabled="true"><h4>Shape of Train and Test Dataset</h4></li>
<li class="list-group-item"><h4>Number of rows in Train dataset is: <span class="label label-primary">{ train.shape[0]:,}</span></h4></li>
<li class="list-group-item"> <h4>Number of columns Train dataset is <span class="label label-primary">{train.shape[1]}</span></h4></li>
<li class="list-group-item"><h4>Number of rows in Test dataset is: <span class="label label-success">{ test.shape[0]:,}</span></h4></li>
<li class="list-group-item"><h4>Number of columns Test dataset is <span class="label label-success">{test.shape[1]}</span></h4></li>
</ul>
"""))
# -
train.info()
# <a id='visual'></a>
# # 3. Visualization of data
missing = train.isnull().sum()
missing[missing>0].sort_values(ascending=False).iplot(kind='bar',title='Null values present in train Dataset', color=['red'])
train.target.value_counts().iplot(kind='bar',text=['Fake', 'Real'], title='Comparing Tweet is a real disaster (1) or not (0)',color=['blue'])
# + _kg_hide-input=true
counts_train = train.target.value_counts(sort=False)
labels = counts_train.index
values_train = counts_train.values
data = go.Pie(labels=labels, values=values_train ,pull=[0.03, 0])
layout = go.Layout(title='Comparing Tweet is a real disaster (1) or not (0) in %')
fig = go.Figure(data=[data], layout=layout)
fig.update_traces(hole=.3, hoverinfo="label+percent+value")
fig.update_layout(
# Add annotations in the center of the donut pies.
annotations=[dict(text='Train', x=0.5, y=0.5, font_size=20, showarrow=False)])
fig.show()
# -
train['length'] = train['text'].apply(len)
# + _kg_hide-input=true
data = [
go.Box(
y=train[train['target']==0]['length'],
name='Fake'
),
go.Box(
y=train[train['target']==1]['length'],
name='Real'
)
]
layout = go.Layout(
title = 'Comparison of text length in Tweets '
)
fig = go.Figure(data=data, layout=layout)
fig.show()
# -
train.keyword.nunique() # Total of 221 unique keywords
# +
train.keyword.value_counts()[:20].iplot(kind='bar', title='Top 20 keywords in text', color='red')
# -
train.location.value_counts()[:20].iplot(kind='bar', title='Top 20 location in tweet', color='blue') # Check the top 15 locations
# <a id='word'></a>
# # 4. WordCloud
# +
STOPWORDS.add('https') # remove htps to the world Cloud
def Plot_world(text):
comment_words = ' '
stopwords = set(STOPWORDS)
for val in text:
# typecaste each val to string
val = str(val)
# split the value
tokens = val.split()
# Converts each token into lowercase
for i in range(len(tokens)):
tokens[i] = tokens[i].lower()
for words in tokens:
comment_words = comment_words + words + ' '
wordcloud = WordCloud(width = 5000, height = 4000,
background_color ='black',
stopwords = stopwords,
min_font_size = 10).generate(comment_words)
# plot the WordCloud image
plt.figure(figsize = (12, 12), facecolor = 'k', edgecolor = 'k' )
plt.imshow(wordcloud)
plt.axis("off")
plt.tight_layout(pad = 0)
plt.show()
# -
# +
text = train.text.values
Plot_world(text)
# -
# <a id='clean'></a>
# # 5. Cleaning the text
#How many http words has this text?
train.loc[train['text'].str.contains('http')].target.value_counts()
# +
pattern = re.compile('http[s]?://(?:[a-zA-Z]|[0-9]|[$-_@.&+]|[!*\(\),]|(?:%[0-9a-fA-F][0-9a-fA-F]))+')
def remove_html(text):
no_html= pattern.sub('',text)
return no_html
# -
# Remove all text that start with html
train['text']=train['text'].apply(lambda x : remove_html(x))
# lets check if this clean works
train.loc[train['text'].str.contains('http')].target.value_counts()
# Remove all text that start with html in test
test['text']=test['text'].apply(lambda x : remove_html(x))
# ### Now remove stopwords, pass to lower add delimiter and more
def clean_text(text):
text = re.sub('[^a-zA-Z]', ' ', text)
text = text.lower()
# split to array(default delimiter is " ")
text = text.split()
text = [w for w in text if not w in set(stopwords.words('english'))]
text = ' '.join(text)
return text
text = train.text[3]
print(text)
clean_text(text)
# Apply clean text
train['text'] = train['text'].apply(lambda x : clean_text(x))
# Apply clean text
test['text']=test['text'].apply(lambda x : clean_text(x))
# How many unique words have this text
def counter_word (text):
count = Counter()
for i in text.values:
for word in i.split():
count[word] += 1
return count
# +
text_values = train["text"]
counter = counter_word(text_values)
# -
print(f"The len of unique words is: {len(counter)}")
list(counter.items())[:10]
# <a id='split'></a>
# # 6. Train Test Split
# +
# The maximum number of words to be used. (most frequent)
vocab_size = len(counter)
embedding_dim = 32
# Max number of words in each complaint.
max_length = 20
trunc_type='post'
padding_type='post'
# oov_took its set for words out our word index
oov_tok = "<XXX>"
training_size = 6090
seq_len = 12
# +
# this is base in 80% of the data, an only text and targert at this moment
training_sentences = train.text[0:training_size]
training_labels = train.target[0:training_size]
testing_sentences = train.text[training_size:]
testing_labels = train.target[training_size:]
# +
print('The Shape of training ',training_sentences.shape)
print('The Shape of testing',testing_sentences.shape)
# -
tokenizer = Tokenizer(num_words=vocab_size, oov_token=oov_tok)
tokenizer.fit_on_texts(training_sentences)
word_index = tokenizer.word_index
# +
# Lets see the first 10 elements
print("THe first word Index are: ")
for x in list(word_index)[0:15]:
print (" {}, {} ".format(x, word_index[x]))
# If you want to see completed -> word_index
# -
training_sequences = tokenizer.texts_to_sequences(training_sentences)
training_padded = pad_sequences(training_sequences, maxlen=max_length, padding=padding_type, truncating=trunc_type)
print(train.text[1])
print(training_sequences[1])
# ## check Inverse for see how it works
# + _kg_hide-output=true
reverse_word_index = dict([(value, key) for (key, value) in word_index.items()])
# +
# Lets see the first 10 elements
print("THe first reverse word Index are: ")
for x in list(reverse_word_index)[0:15]:
print (" {}, {} ".format(x, reverse_word_index[x]))
# If you want to see completed -> reverse_word_index
# -
def decode(text):
return ' '.join([reverse_word_index.get(i, '?') for i in text])
decode(training_sequences[1]) # this can be usefull for check predictions
training_padded[1628]
# +
testing_sequences = tokenizer.texts_to_sequences(testing_sentences)
testing_padded = pad_sequences(testing_sequences, maxlen=max_length, padding=padding_type, truncating=trunc_type)
# -
# <a id='model'></a>
# # 7. Creating the Model
#
# # For a binary classification problem
# model.compile(optimizer='adam',
# loss='binary_crossentropy',
# metrics=['accuracy'])
#
# +
# Model Definition with LSTM
model = tf.keras.Sequential([
tf.keras.layers.Embedding(vocab_size, embedding_dim, input_length=max_length),
tf.keras.layers.Bidirectional(tf.keras.layers.LSTM(64)),
tf.keras.layers.Dense(14, activation='relu'),
tf.keras.layers.Dense(1, activation='sigmoid') # remember this is a binary clasification
])
model.compile(loss='binary_crossentropy',optimizer='adam',metrics=['accuracy'])
# -
model.summary()
from tensorflow.keras.utils import plot_model
plot_model(model, to_file='model_plot4a.png', show_shapes=True, show_layer_names=True)
# +
start_time = time.time()
num_epochs = 10
history = model.fit(training_padded, training_labels, epochs=num_epochs, validation_data=(testing_padded, testing_labels))
final_time = (time.time()- start_time)/60
print(f'The time in minutos: {final_time}')
# -
model_loss = pd.DataFrame(model.history.history)
model_loss.head()
model_loss[['accuracy','val_accuracy']].plot(ylim=[0,1]);
# <a id='eval'></a>
# # 8. Model Evaluation
predictions = model.predict_classes(testing_padded) # predict_ clases because is classification problem with the split test
predictions
from sklearn.metrics import classification_report,confusion_matrix
# Showing Confusion Matrix
def plot_cm(y_true, y_pred, title, figsize=(5,4)):
cm = confusion_matrix(y_true, y_pred, labels=np.unique(y_true))
cm_sum = np.sum(cm, axis=1, keepdims=True)
cm_perc = cm / cm_sum.astype(float) * 100
annot = np.empty_like(cm).astype(str)
nrows, ncols = cm.shape
for i in range(nrows):
for j in range(ncols):
c = cm[i, j]
p = cm_perc[i, j]
if i == j:
s = cm_sum[i]
annot[i, j] = '%.1f%%\n%d/%d' % (p, c, s)
elif c == 0:
annot[i, j] = ''
else:
annot[i, j] = '%.1f%%\n%d' % (p, c)
cm = pd.DataFrame(cm, index=np.unique(y_true), columns=np.unique(y_true))
cm.index.name = 'Actual'
cm.columns.name = 'Predicted'
fig, ax = plt.subplots(figsize=figsize)
plt.title(title)
sns.heatmap(cm, cmap= "YlGnBu", annot=annot, fmt='', ax=ax)
# Showing Confusion Matrix
plot_cm(testing_labels,predictions, 'Confution matrix of Tweets', figsize=(7,7))
# +
testing_sequences2 = tokenizer.texts_to_sequences(test.text)
testing_padded2 = pad_sequences(testing_sequences2, maxlen=max_length, padding=padding_type, truncating=trunc_type)
# -
predictions = model.predict(testing_padded2)
# sample of submission
submission.head()
submission['target'] = (predictions > 0.5).astype(int)
submission
submission.to_csv("submission.csv", index=False, header=True)
# <a href="#top" class="btn btn-primary btn-lg active" role="button" aria-pressed="true">Go to TOP</a>
#
| static/img/BasicNLPbyusingTensorFlow.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9 (tensorflow)
# language: python
# name: tensorflow
# ---
# + [markdown] id="XKzF6dMaiLyP"
# <a href="https://colab.research.google.com/github/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_06_5_yolo.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="YDTXd8-Lmp8Q"
# # T81-558: Applications of Deep Neural Networks
# **Module 6: Convolutional Neural Networks (CNN) for Computer Vision**
# * Instructor: [<NAME>](https://sites.wustl.edu/jeffheaton/), McKelvey School of Engineering, [Washington University in St. Louis](https://engineering.wustl.edu/Programs/Pages/default.aspx)
# * For more information visit the [class website](https://sites.wustl.edu/jeffheaton/t81-558/).
# + [markdown] id="ncNrAEpzmp8S"
# # Module 6 Material
#
# * Part 6.1: Image Processing in Python [[Video]](https://www.youtube.com/watch?v=V-IUrfTJMm4&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_06_1_python_images.ipynb)
# * Part 6.2: Using Convolutional Neural Networks [[Video]](https://www.youtube.com/watch?v=nU_T2PPigUQ&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_06_2_cnn.ipynb)
# * Part 6.3: Using Pretrained Neural Networks with Keras [[Video]](https://www.youtube.com/watch?v=TXqI9fp0imI&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_06_3_resnet.ipynb)
# * Part 6.4: Looking at Keras Generators and Image Augmentation [[Video]](https://www.youtube.com/watch?v=epfpxiXRL3U&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_06_4_keras_images.ipynb)
# * **Part 6.5: Recognizing Multiple Images with YOLOv5** [[Video]](https://www.youtube.com/watch?v=zwEmzElquHw&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_06_5_yolo.ipynb)
# + [markdown] id="z0stsqSVoKD0"
# # Google CoLab Instructions
#
# The following code ensures that Google CoLab is running the correct version of TensorFlow.
# Running the following code will map your GDrive to ```/content/drive```.
# + colab={"base_uri": "https://localhost:8080/"} id="fU9UhAxTmp8S" outputId="0d05ed99-7bdf-4855-ef71-478f68afabb1"
try:
from google.colab import drive
COLAB = True
print("Note: using Google CoLab")
# %tensorflow_version 2.x
except:
print("Note: not using Google CoLab")
COLAB = False
# + [markdown] id="QSKZqD1Mmp-C"
# # Part 6.5: Recognizing Multiple Images with YOLO5
#
# Programmers typically design convolutional neural networks to classify a single item centered in an image. However, as humans, we can recognize many items in our field of view in real-time. It is advantageous to recognize multiple items in a single image. One of the most advanced means of doing this is YOLOv5. You Only Look Once (YOLO) was introduced by <NAME>, who supported YOLO up through V3. [[Cite:redmon2016you]](https://arxiv.org/abs/1506.02640) The fact that YOLO must only look once speaks to the efficiency of the algorithm. In this context, to "look" means to perform one scan over the image. It is also possible to run YOLO on live video streams.
#
#
# <NAME> left computer vision to pursue other interests. The current version, YOLOv5 is supported by the startup company [Ultralytics](https://ultralytics.com/), who released the open-source library that we use in this class.[[Cite:zhu2021tph]](https://arxiv.org/abs/2108.11539)
#
# Researchers have trained YOLO on a variety of different computer image datasets. The version of YOLO weights used in this course is from the dataset Common Objects in Context (COCO). [[Cite: lin2014microsoft]](https://arxiv.org/abs/1405.0312) This dataset contains images labeled into 80 different classes. COCO is the source of the file coco.txt used in this module.
#
# ## Using YOLO in Python
#
# To use YOLO in Python, we will use the open-source library provided by Ultralytics.
#
# * [YOLOv5 GitHub](https://github.com/ultralytics/yolov5)
#
# The code provided in this notebook works equally well when run either locally or from Google CoLab. It is easier to run YOLOv5 from CoLab, which is recommended for this course.
#
# We begin by obtaining an image to classify.
#
# + colab={"base_uri": "https://localhost:8080/", "height": 337} id="WXEobzvGlFim" outputId="e3b8422a-c417-450e-dbc8-704e768f67ac"
import urllib.request
import shutil
from IPython.display import Image
# !mkdir /content/images/
URL = "https://github.com/jeffheaton/t81_558_deep_learning"
URL += "/raw/master/photos/jeff_cook.jpg"
LOCAL_IMG_FILE = "/content/images/jeff_cook.jpg"
with urllib.request.urlopen(URL) as response, \
open(LOCAL_IMG_FILE, 'wb') as out_file:
shutil.copyfileobj(response, out_file)
Image(filename=LOCAL_IMG_FILE)
# + [markdown] id="Ym5_juokofQl"
# ## Installing YOLOv5
#
# YOLO is not available directly through either PIP or CONDA. Additionally, YOLO is not installed in Google CoLab by default. Therefore, whether you wish to use YOLO through CoLab or run it locally, you need to go through several steps to install it. This section describes the process of installing YOLO. The same steps apply to either CoLab or a local install. For CoLab, you must repeat these steps each time the system restarts your virtual environment. You must perform these steps only once for your virtual Python environment for a local install. If you are installing locally, install to the same virtual environment you created for this course. The following commands install YOLO directly from its GitHub repository.
# + colab={"base_uri": "https://localhost:8080/"} id="VuTjby5MzEre" outputId="7232ded0-18af-47ba-e760-2bedf4cb3dc5"
# !git clone https://github.com/ultralytics/yolov5 --tag 6.1
# !mv /content/6.1 /content/yolov5
# %cd /content/yolov5
# %pip install -qr requirements.txt
from yolov5 import utils
display = utils.notebook_init()
# + [markdown] id="9PSttcoraUlb"
# Next, we will run YOLO from the command line and classify the previously downloaded kitchen picture. You can run this classification on any image you choose.
# + colab={"base_uri": "https://localhost:8080/", "height": 651} id="R8zq_6akz64w" outputId="c72c4105-7e04-42c6-eec5-8f9fcf05b5f4"
# !python detect.py --weights yolov5s.pt --img 640 \
# --conf 0.25 --source /content/images/
URL = '/content/yolov5/runs/detect/exp/jeff_cook.jpg'
display.Image(filename=URL, width=300)
# + [markdown] id="qYOvD3M7ofQl"
# ## Running YOLOv5
#
# In addition to the command line execution, we just saw. The following code adds the downloaded YOLOv5 to Python's environment, allowing **yolov5** to be imported like a regular Python library.
# + colab={"base_uri": "https://localhost:8080/"} id="MY3gVyidmp-K" outputId="8da0b9a2-1752-43ee-c990-e30b93ca8d18"
import sys
sys.path.append(str("/content/yolov5"))
from yolov5 import utils
display = utils.notebook_init()
# + [markdown] id="CYhbOw1roKEN"
# Next, we obtain an image to classify. For this example, the program loads the image from a URL. YOLOv5 expects that the image is in the format of a Numpy array. We use PIL to obtain this image. We will convert it to the proper format for PyTorch and YOLOv5 later.
# + id="J5ilnhhNiLyq"
from PIL import Image
import requests
from io import BytesIO
import torchvision.transforms.functional as TF
url = "https://raw.githubusercontent.com/jeffheaton/"\
"t81_558_deep_learning/master/images/cook.jpg"
response = requests.get(url,headers={'User-Agent': 'Mozilla/5.0'})
img = Image.open(BytesIO(response.content))
# + [markdown] id="JYr64eMMoKER"
# The following libraries are needed to classify this image.
# + id="W3jTpQzj7RXk"
import argparse
import os
import sys
from pathlib import Path
import cv2
import torch
import torch.backends.cudnn as cudnn
from models.common import DetectMultiBackend
from utils.datasets import IMG_FORMATS, VID_FORMATS, LoadImages, LoadStreams
from utils.general import (LOGGER, check_file, check_img_size, check_imshow,
check_requirements, colorstr,
increment_path, non_max_suppression,
print_args, scale_coords, strip_optimizer,
xyxy2xywh)
from utils.plots import Annotator, colors, save_one_box
from utils.torch_utils import select_device, time_sync
# + [markdown] id="URXt2GjY8RBD"
# We are now ready to load YOLO with pretrained weights provided by the creators of YOLO. It is also possible to train YOLO to recognize images of your own.
# + colab={"base_uri": "https://localhost:8080/"} id="LVCuZN2p7Yyy" outputId="3a88397d-06fa-478e-a690-438d839b04aa"
device = select_device('')
weights = '/content/yolov5/yolov5s.pt'
imgsz = [img.height, img.width]
original_size = imgsz
model = DetectMultiBackend(weights, device=device, dnn=False)
stride, names, pt, jit, onnx, engine = model.stride, model.names, \
model.pt, model.jit, model.onnx, model.engine
imgsz = check_img_size(imgsz, s=stride) # check image size
print(f"Original size: {original_size}")
print(f"YOLO input size: {imgsz}")
# + [markdown] id="sGbur-vdZWyz"
# The creators of YOLOv5 built upon PyTorch, which has a particular format for images. PyTorch images are generally a 4D matrix of the following dimensions:
#
# * batch_size, channels, height, width
#
# This code converts the previously loaded PIL image into this format.
# + colab={"base_uri": "https://localhost:8080/"} id="o_oqpylt9I_n" outputId="a592010a-978e-4119-9e80-7aa0b5e8e175"
import numpy as np
source = '/content/images/'
conf_thres=0.25 # confidence threshold
iou_thres=0.45 # NMS IOU threshold
classes = None
agnostic_nms=False, # class-agnostic NMS
max_det=1000
model.warmup(imgsz=(1, 3, *imgsz)) # warmup
dt, seen = [0.0, 0.0, 0.0], 0
# https://stackoverflow.com/questions/50657449/
# convert-image-to-proper-dimension-pytorch
img2 = img.resize([imgsz[1],imgsz[0]], Image.ANTIALIAS)
img_raw = torch.from_numpy(np.asarray(img2)).to(device)
img_raw = img_raw.float() # uint8 to fp16/32
img_raw /= 255 # 0 - 255 to 0.0 - 1.0
img_raw = img_raw.unsqueeze_(0)
img_raw = img_raw.permute(0, 3, 1, 2)
print(img_raw.shape)
# + [markdown] id="xFglXpJc8cxY"
# With the image converted, we are now ready to present the image to YOLO and obtain predictions.
# + id="zGtkQOH1nv10"
pred = model(img_raw, augment=False, visualize=False)
pred = non_max_suppression(pred, conf_thres, iou_thres, classes,
agnostic_nms, max_det=max_det)
# + [markdown] id="bftSuKLo8ipJ"
# We now convert these raw predictions into the bounding boxes, labels, and confidences for each of the images that YOLO recognized.
# + id="fAk8F8bFAPFA"
results = []
for i, det in enumerate(pred): # per image
gn = torch.tensor(img_raw.shape)[[1, 0, 1, 0]]
if len(det):
# Rescale boxes from img_size to im0 size
det[:, :4] = scale_coords(original_size, det[:, :4], imgsz).round()
# Write results
for *xyxy, conf, cls in reversed(det):
xywh = (xyxy2xywh(torch.tensor(xyxy).view(1, 4)) / \
gn).view(-1).tolist()
# Choose between xyxy and xywh as your desired format.
results.append([names[int(cls)], float(conf), [*xyxy]])
# + [markdown] id="L37owfBG8tmQ"
# We can now see the results from the classification. We will display the first 3.
# + colab={"base_uri": "https://localhost:8080/"} id="4mcP90gYnjaY" outputId="fd55b7e3-1961-4c62-9666-2368f37f3c32"
for itm in results[0:3]:
print(itm)
# + [markdown] id="DL173C9_oKEU"
# It is important to note that the **yolo** class instantiated here is a callable object, which can fill the role of both an object and a function. Acting as a function, *yolo* returns three arrays named **boxes**, **scores**, and **classes** that are of the same length. The function returns all sub-images found with a score above the minimum threshold. Additionally, the **yolo** function returns an array named called **nums**. The first element of the **nums** array specifies how many sub-images YOLO found to be above the score threshold.
#
# * **boxes** - The bounding boxes for each sub-image detected in the image sent to YOLO.
# * **scores** - The confidence for each of the sub-images detected.
# * **classes** - The string class names for each item. These are COCO names such as "person" or "dog."
# * **nums** - The number of images above the threshold.
#
# Your program should use these values to perform whatever actions you wish due to the input image. The following code displays the images detected above the threshold.
#
# To demonstrate the correctness of the results obtained, we draw bounding boxes over the original image.
# + colab={"base_uri": "https://localhost:8080/", "height": 497} id="5d_vJoYn3tIX" outputId="1413d548-8b9e-4e0a-9a76-d59f3da595c3"
from PIL import Image, ImageDraw
img3 = img.copy()
draw = ImageDraw.Draw(img3)
for itm in results:
b = itm[2]
print(b)
draw.rectangle(b)
img3
# + [markdown] id="0eBtaFbimp-M"
# # Module 6 Assignment
#
# You can find the first assignment here: [assignment 6](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/assignments/assignment_yourname_class6.ipynb)
| t81_558_class_06_5_yolo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Reference text
# **bold**
# *italics*
#
# bullets
# * point 1
# * point 2
# # Title
#
# ## Subtitle
#
# ### SubSubtitle
#
# LaTeX equations
# $ 1 + 1 = 2 $
# $$ x + y = z $$
#
# Markdown cheat sheet
# https://guides.github.com/pdfs/markdown-cheatsheet-online.pdf
# +
# %matplotlib inline
import matplotlib.pyplot as plt
# -
# # Steps for run RCA tools
#
# The
#
# 1) Create a daily clutter map
#
# 2) Repeat (1) as needed
#
# 3) Create a composite clutter map
#
# 4) Calculate a baseline value
#
# 5) Calculate a daily median RCA
#
# ## 1) Create a daily clutter map
#
# Daily clutter map(s) must be created FIRST before any RCA calculations may be made.
#
# A daily clutter map is generated from a day's worth of radar files. You need only specify the directory of the radar files, the desired date (YYYYMMDD), the scan type (PPI or RHI), and if you're using horizontally or dual-polarized data. This is all done in the a handful of radar- and scan-specific JSON files, which should be modifed by the user for their specifications.
#
# The current defaults in, for example, kaband_rhi.json are:
#
# "config": "kaband_rhi",
# "scan_type": "rhi",
# "polarization": "horizontal",
# "data_directory": "/home/user/data/cor/corkasacrcfrhsrhiM1.a1/",
# "file_extension": ".nc",
# "cluttermap_directory": "/home/user/clutter_maps/",
# "cluttermap_date": "20181105",
# "baseline_directory": "/home/user/baselines/",
# "baseline_date": "20181108",
# "daily_csv_dir": "/home/user/dailies/",
# "site_abbrev": "cor",
# "instrument_abbrev": "kasacr",
# "range_limit": 10000,
# "z_threshold": 30
# Call the clutter_map function from clutter_map.py. You will need to adjust the paramters in the clutter_maps.json configuration file before using the JSON file as input for the funtion.
#
# You will need to import the following packages to use the clutter_map function:
# +
import numpy as np
import pyart
import os
import glob
import json
from netCDF4 import Dataset
from modules.create_clutter_flag import create_clutter_flag_ppi, create_clutter_flag_hsrhi
from modules.file_to_radar_object import file_to_radar_object
from clutter_map import clutter_map
# Replace the input with the JSON file named for the radar band and scan type
# band_scan.json
clutter_map('kaband_rhi.json','20181105')
# -
# Repeat this for all desired days you want to generate a daily clutter map.
#
# Save the output .nc file to a directory specifically for clutter maps (recommended).
#
# This directory will be used as the input for the next step, composite clutter map generation.
# # Create a composite clutter map
#
# A composite clutter map is composed of all specified daily clutter maps that must be previously generated. The composite clutter map identifies points where at least 80% of the daily clutter maps have identified as clutter points. This ensures the clutter signature is stable, especially in regions where the ground clutter varies or fluctuates day-to-day.
#
# Use the composite_clutter_map function and a new netCDF is written with the composite clutter map in an array. The function takes the clutter_maps JSON file as input with current deafults the same as listed above.
#
# The following new packages are required to run composite_clutter_map:
# +
from composite_clutter_map import composite_clutter_map
from get_pct_on_clutter_map import get_pct_on_clutter_map_ppi, get_pct_on_clutter_map_hsrhi
composite_clutter_map('clutter_maps.json')
# -
# ## Plotting a clutter map
# A simple plot may be generated using the plot_clutter_map function. This helps visualize where the clutter points calculated previously are located spatially and how many points did or did not make the threshold.
#
# Specify either "day" or "composite" in the function input to represent the appropriate data.
#
# ### Daily clutter map
# +
import matplotlib.pyplot as plt
from netCDF4 import Dataset
from plot_clutter_map import plot_clutter_map
clutter_map_netcdf = '/Users/hunz743/projects/rca_auxillary/datafiles/cluttermaps/cluttermap_ppi_corcsapr2_composite.nc'
output_directory = '/Users/hunz743/projects/rca_auxillary/figures/'
scan_type = 'ppi'
map_type = 'composite'
polarization = 'horizontal'
max_range = 40
site = 'cor'
inst = 'csapr2'
location = 'CORCSAPR2'
plot_clutter_map(clutter_map_netcdf,output_directory,scan_type,map_type,polarization,max_range,site,inst,location)
# -
# ### Composite clutter map
#
# Change the inputs and use the same function to generate a composite clutter map. The color bar differs from the daily map, showing determined clutter points in black. These are the elements that are used to calculate clutter area reflectivity in the baseline and daily calculations.
map_type = 'composite'
plot_clutter_map(clutter_map_netcdf,output_directory,scan_type,map_type,polarization,max_range,site,inst,location)
# # Calculate the baseline value
#
# The baseline is calculated using a day of radar files and (preferably) the composite clutter map.
#
# Simply use the baseline function with relevant input (may use and adjust any of the radar_scan.json files).
# +
import numpy as np
import os
import glob
import json
from netCDF4 import Dataset
from file_to_radar_object import file_to_radar_object
from calculate_dbz95 import calculate_dbz95_ppi, calculate_dbz95_hsrhi
from baseline import baseline
radar_config_file = 'kaband_rhi.json'
dbz95_h = baseline(radar_config_file)
print('Baseline value: ', dbz95_h, 'dBZ')
# -
# # Calculate the RCA value
#
# The daily RCA value is calculated using a day of radar files, the composite clutter map, and the baseline value.
#
# All of these are read in using the daily_rca function. Again, specify which radar_scan.json to use and adjust accordingly. This function may be looped over for all days of interest. Each day has a mean RCA calculated and written to a CSV file.
#
# (It is recommended - and even necessary - to create the CSV file with appropriate headers before running the function. Daily CSV headers are: DATE, RCA_H, RCA_V, BASELINE)
# +
import numpy as np
import os
import glob
import json
from netCDF4 import Dataset
import pandas as pd
from file_to_radar_object import file_to_radar_object
from calculate_dbz95 import calculate_dbz95_ppi, calculate_dbz95_hsrhi
from daily_rca import daily_rca
radar_config_file = 'kaband_rhi.json'
daily_rca(radar_config_file,'20181109')
# -
# ## Plot a timeseries of daily RCA values
#
| src/rca/.ipynb_checkpoints/RCA_Demo-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="RxhYJEQn6gdj" colab_type="text"
# 1. Mana dari statement berikut ini yang benar mengenai Pandas Series dan DataFrame?
# Semuanya Benar
#
# 2. Apakah objek dataframe teridiri dari objek-objek series?
# True
#
# 3. Bagaimana process grouping dengan method groupby di pandas
# Splitting - Applying - Combining
#
# 4. Mana di bawah ini statement yang benar mengenai Pandas index?
# Semuanya Benar
#
# 5. Mana method berikut yang akan menghasilkan summary statistic dari dataframe?
# .describe()
#
# 6. Diantara opsi berikut ini, operasi mana di pandas yang syntax nya sama seperti di dictionary?
# Semuanya Benar
#
# 7. Series adalah array satu dimensi yang memiliki label dan bisa berisi berbagai tipe data.
# True
#
# 8. Apa jenis penggabungan yang diterapkan pada parameter 'how' jika df_gabung adalah hasil dari df1.merge(df2, on='....', how='....')?
# right
#
# 9. Apa Jenis penggabungan data berikut
# Left Join
#
# 10. Mana berikut di bawah ini yang bukan merupkan fungsi reshaping data?
# Tidak ada, Semuanya merupakan fungsi reshaping data
#
# 11. Mana code berikut adalah cara yang benar untuk mengambil baris Umur dan column Nama dari sebuah dataframe bernama variabel df?
# df['Umur']['Nama']
#
# 12. Jika data berbentuk objek array, maka index dan value harus memiliki jumlah elemen yang sama?
# True
#
# 13. Kita dapat memfilter saat akses dataframe menggunakan teknik berikut, KECUALI?
# Grouping
#
# 14. Mana objek berikut ini yang dapat di jadikan DataFrame?
# Semuanya
#
# 15. Mana attribute dibawah ini yang bisa di gunakan untuk mendapatkan numpy array dari DataFrame?
# .values
# + id="9zYxV7bZ_7Jv" colab_type="code" outputId="9555c26d-7b9d-4952-8562-23b31d9020f0" colab={"base_uri": "https://localhost:8080/", "height": 86}
import pandas as pd
x = pd.DataFrame([6,'aa',True][])
x
# + id="4X8V0inBB_45" colab_type="code" outputId="20467e06-6251-4d26-d756-79763e3730e1" colab={"base_uri": "https://localhost:8080/", "height": 34}
df = [
[0,2],
[1,2]
]
df[0][1]
| Learn/Week 2 Pandas/Week_2_Quiz_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# plotting libraries
import matplotlib
import matplotlib.pyplot as plt
# numpy (math) libary
import numpy as np
import sys
# +
### import PARAMETERS and CONSTANTS
from modules.ConstantsAndParameters import *
### import UTILITY functions
from modules.utils import *
#print_const()
#print_const(normalized=True)
### import model's EQUATIONS
from modules.model import *
# +
# greek letters and other symbols (Ctrl+Shift+u):
# Γ = u0393
# Δ = u0394
# Ω = u03a9
# α = u03b1
# β = u03b2
# γ = u03b3, 𝛾 = u1D6FE
# δ = u03b4
# ε = u03b5
# λ = u03bb
# σ = u03c3
# τ = u03c4
# ψ = u03c8
# ω = u03c9
# √ = u221a
# +
history = 10
error = 1e-16
ω_size = 150
#ω_range = np.linspace(wlen_to_freq(1.552e-6), wlen_to_freq(1.548e-6), ω_size)
ω_range = np.linspace(w2f(1.548e-6), w2f(1.552e-6), ω_size) # [Hz]
E_size = 6
E_range = np.power(np.logspace(-3,-2, num=E_size, endpoint=True), 0.5) # [10^x Watt]
# +
verbose = True
loops = True
d = {}
d['xUp'] = np.zeros((ω_size,E_size,history), dtype=complex)
d['xUs'] = np.zeros((ω_size,E_size,history), dtype=complex)
d['xUtot']= np.zeros((ω_size,E_size,history), dtype=float)
d['xΔN'] = np.zeros((ω_size,E_size,history), dtype=float)
d['xΔT'] = np.zeros((ω_size,E_size,history), dtype=float)
d['xΔω'] = np.zeros((ω_size,E_size,history), dtype=complex)
for E_idx in range(E_size):
for ω_idx in range(ω_size):
it = 0
#clear_output(wait=True)
#print("\rP %.2f"%E_range[E_idx], ", λ %.4f"%freq_to_wlen(ω_range[ω_idx]*1e-6), ", iteration # %d"%it, end='')
#params = (ωs, ωp, ω0, Ep, Es, τa, τb, τ0, 𝛾TH, 𝛾FC, MCp, n0, n2, dndT, dndN, dαdN, βtpa, Γ, V, Veff)
#params = (ω_range[ω_idx], ωp, ω0, E_range[E_idx], Es, τa, τb, τ0, 𝛾TH, 𝛾FC, MCp, n0, n2, dndT, dndN, dαdN, βtpa, Γ, V, Veff)
params = (ω_range[ω_idx], ωp, ω0, E_range[E_idx], Es, τa, τb, τ0, 𝛾TH, 1, MCp, n0, 0.0, dndT, 0.0, 0.0, 0.0, Γ, V, Veff)
### generate first set of data, with all initial condition set to zero ("cold resonance")
new_var = equations((0.0, 0.0, 0.0, 0.0, 0.0, 0.0), params)
d['xUp'][ω_idx,E_idx,0] = new_var[0]
d['xUs'][ω_idx,E_idx,0] = new_var[1]
d['xUtot'][ω_idx,E_idx,0] = new_var[2]
d['xΔN'][ω_idx,E_idx,0] = new_var[3]
d['xΔT'][ω_idx,E_idx,0] = new_var[4]
d['xΔω'][ω_idx,E_idx,0] = new_var[5]
del new_var
while (np.abs(d['xUtot'][ω_idx,E_idx,0]-d['xUtot'][ω_idx,E_idx,1]) >= error) & (it<=2e3):
it += 1
#clear_output()#wait=True)
#print("\rP %.2f"%E_range[E_idx], ", λ %.4f"%freq_to_wlen(ω_range[ω_idx]*1e-6), end='')
#print(", iteration # %d"%it , "error %.4e"%np.abs(d['xUtot'][ω_idx,E_idx,0]-d['xUtot'][ω_idx,E_idx,1]), end='')
### shift the history data
d['xUp'][ω_idx,E_idx,1:history] = d['xUp'][ω_idx,E_idx,0:history-1]
d['xUs'][ω_idx,E_idx,1:history] = d['xUs'][ω_idx,E_idx,0:history-1]
d['xUtot'][ω_idx,E_idx,1:history] = d['xUtot'][ω_idx,E_idx,0:history-1]
d['xΔN'][ω_idx,E_idx,1:history] = d['xΔN'][ω_idx,E_idx,0:history-1]
d['xΔT'][ω_idx,E_idx,1:history] = d['xΔT'][ω_idx,E_idx,0:history-1]
d['xΔω'][ω_idx,E_idx,1:history] = d['xΔω'][ω_idx,E_idx,0:history-1]
### put old data in vector, to become input of equations()
old_var = (d['xUp'][ω_idx,E_idx,1], d['xUs'][ω_idx,E_idx,1], d['xUtot'][ω_idx,E_idx,1], d['xΔN'][ω_idx,E_idx,1], d['xΔT'][ω_idx,E_idx,1], d['xΔω'][ω_idx,E_idx,1])
### generate new values
new_var = equations(old_var, params)
### store new values
d['xUp'][ω_idx,E_idx,0] = new_var[0]
d['xUs'][ω_idx,E_idx,0] = new_var[1]
d['xUtot'][ω_idx,E_idx,0] = new_var[2]
d['xΔN'][ω_idx,E_idx,0] = new_var[3]
d['xΔT'][ω_idx,E_idx,0] = new_var[4]
d['xΔω'][ω_idx,E_idx,0] = new_var[5]
if verbose:
print("P %.2e,"%(E_range[E_idx]**2*P0), " λ %.5f"%f2w(ω_range[ω_idx]*1e-6*ω0), end='')
print(", iteration # %d,"%it , " error %.4e"%np.abs(d['xUtot'][ω_idx,E_idx,0]-d['xUtot'][ω_idx,E_idx,1]))
if loops:
print("P %.2f"%(E_range[E_idx]**2),"mW ended")
else:
print()
print("\nall loops have ended")
# +
linear = True
fig = plt.figure(figsize=(3*6.4, 2*4.8)) # default = 6.4, 4.8
ax1 = fig.add_subplot(111)
if not linear:
ax1.set_yscale('log')
ax1.set_ylim([1e-20, 1e-2])
for E_idx in range(E_size):
#if E_idx <= 3:
#ax1.plot(1e9*f2w(ω_range[:],), d['xUtot'][:,E_idx,0], label='Uint')
#ax1.plot(1e9*f2w(ω_range[:],), d['xUtot'][:,E_idx,0]-np.mean(d['xUtot'][:,E_idx,0]), label='Uint, P %.3f'%E_range[E_idx]**2 )
ax1.plot(1e9*f2w(ω_range[:],), np.abs(d['xUs'][:,E_idx,0])**2, label='Us, P %.3f'%E_range[E_idx]**2)
#else:
#ax1.plot(1e9*freq_to_wlen(ω_range[:],), , label='Uint P %.3f'%E_range[E_idx] )
ax1.set_title( r'Internal Power', fontsize=16)
ax1.set_xlabel(r'Wavelength $\lambda$ $[nm]$', fontsize=16)
ax1.set_ylabel(r'Internal Power $[$not exactly $a.u.]$', fontsize=16)
legend = ax1.legend(loc='upper right', fontsize=16)#, bbox_to_anchor=(1, 0.5))
plt.show()
plt.close()
# +
#for E_idx in range(E_size):
# print()
# indx = np.where( d['xUtot'][:,E_idx,0]==max(d['xUtot'][:,E_idx,0]) )
# tmp = f2w(ω_range[indx])
# print(tmp[0])
#
# indx = np.where( d['xUtot'][:,E_idx,0]>=max(d['xUtot'][:,E_idx,0])/2 )
# tmp = f2w(ω_range[indx])
# print(tmp[0]-tmp[-1])
#
# del indx, tmp
#print()
for E_idx in range(E_size):
print()
indx = np.where( d['xUs'][:,E_idx,0]**2==max(d['xUs'][:,E_idx,0]**2) )
tmp = f2w(ω_range[indx])
print(tmp[0])
indx = np.where( d['xUs'][:,E_idx,0]**2>=max(d['xUs'][:,E_idx,0]**2)/2 )
tmp = f2w(ω_range[indx])
print(tmp[0]-tmp[-1])
del indx, tmp
# +
linear = True
fig = plt.figure(figsize=(3*6.4, 2*4.8)) # default = 6.4, 4.8
ax1 = fig.add_subplot(111)
if not linear:
ax1.set_yscale('log')
ax1.set_ylim([1e-20, 1e-2])
for E_idx in range(E_size):
#if E_idx <4:
#ax1.plot(1e9*f2w(ω_range,), d['xΔN'][:,E_idx,0], label='ΔN')
ax1.plot(1e9*f2w(ω_range,), d['xΔT'][:,E_idx,0], label='ΔT')
#ax1.plot(1e9*f2w(ω_range,), np.real(d['xΔω'][:,E_idx,0])/ω0, label='RE[Δω]')
#ax1.plot(1e9*f2w(ω_range,), np.imag(d['xΔω'][:,E_idx,0])/ω0, label='IM[Δω]', ls='--')
ax1.set_title( r'Internal Power', fontsize=16)
ax1.set_xlabel(r'Wavelength $\lambda$ $[nm]$', fontsize=16)
ax1.set_ylabel(r'Internal Power $[$not exactly $a.u.]$', fontsize=16)
legend = ax1.legend(loc='upper right', fontsize=16)#, bbox_to_anchor=(1, 0.5))
plt.show()
plt.close()
# -
| code/Jupyter/QuasiStatic-analysis/NOT-normalized-Time-Domain.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
from hans.input import Input
from hans.plottools import DatasetSelector
myProblem = Input("channel1D_DH.yaml", None).getProblem()
myProblem.run(out_dir="data")
files = DatasetSelector("data", mode="single")
# +
# %matplotlib notebook
gr = 1.618
pt_per_inch = 72
width_pt = 500
size_x = width_pt / pt_per_inch
size_y = size_x / gr
fig, ax = plt.subplots(figsize=(size_x, size_y))
data = files.get_centerlines("p")
cmap = plt.cm.coolwarm
for fn, cl in data.items():
maxT = max(cl["p"])
sm = plt.cm.ScalarMappable(cmap=cmap, norm=plt.Normalize(vmin=0, vmax=maxT*1e6))
for t, (xdata, ydata) in cl["p"].items():
ax.plot(xdata*1e3, ydata/101325, color=cmap(t/maxT))
fig.colorbar(sm, ax=ax, label='time $t$ (µs)', extend='max')
ax.set_xlabel(r"Distance $x\,\mathrm{(mm)}$")
ax.set_ylabel(r"Pressure $p\,\mathrm{(atm)}$");
| examples/channel1D_DH.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] run_control={"frozen": false, "read_only": false}
# # Welcome to NeuNorm
#
# Package to normalize data using Open Beam (OB) and, optionally Dark Field (DF).
#
# The program allows you to select a background region to allow data to be normalized by OB that do not have the same acquisition time.
# Cropping the image is also possible using the *crop* method
#
# + [markdown] run_control={"frozen": false, "read_only": false}
# In this notebook, we are going to load a data array manually into NeuNorm and export it without doing any normalizaton on it.
#
# + [markdown] run_control={"frozen": false, "read_only": false}
# # Set up system
# + run_control={"frozen": false, "read_only": false}
import os
import sys
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.patches as patches
from matplotlib import gridspec
import glob
from PIL import Image
# %matplotlib notebook
# + [markdown] run_control={"frozen": false, "read_only": false}
# Add NeuNorm to python path
# + run_control={"frozen": false, "read_only": false}
root_folder = os.path.dirname(os.getcwd())
sys.path.insert(0, root_folder)
import NeuNorm as neunorm
from NeuNorm.normalization import Normalization
from NeuNorm.roi import ROI
# + [markdown] run_control={"frozen": false, "read_only": false}
# # Loading Data Manually
# + [markdown] run_control={"frozen": false, "read_only": false}
# Sample data path
# + run_control={"frozen": false, "read_only": false}
path_im = '../data/sample'
sample_data = os.path.join(path_im, '0001.tif')
_sample_data = np.array(Image.open(sample_data))
# + [markdown] run_control={"frozen": false, "read_only": false}
# # Adding the data to the Object
# + run_control={"frozen": false, "read_only": false}
o_norm = Normalization()
o_norm.load(data=_sample_data)
# + [markdown] run_control={"frozen": false, "read_only": false}
# # Exporting the data
# + run_control={"frozen": false, "read_only": false}
output_folder = '/Users/j35/Desktop/'
output_name = os.path.basename(sample_data)
# -
o_norm.export(folder=output_folder, data_type='sample')
| notebooks/tutorial_using_array_input_and_export_array.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Auto generated report for France
#
# This notebook generates for each available geographic zone a summary of the most recent simulation.
#
# ## Preparation
# ### Loading libraries
# %load_ext autoreload
# %autoreload 2
from IPython.display import display, Markdown, Latex
import os
from functools import partial
from matplotlib import pyplot as plt
import matplotlib
import pandas as pd
from numpy import unique
from model_analysis import *
# ### Loading data
run_manifest = read_manifest('run-manifest.csv')
analysis_set = get_analysis_set(run_manifest)
# +
def add_to_zone_dict(zone_dict, row, i, zone):
zone_dict[zone] = {
"index": i,
"date": row["last available data"],
"version": row["version"],
}
zone_dict = {}
for i, row in analysis_set.iterrows():
for zone in row["modelling zones"]:
if zone not in zone_dict:
add_to_zone_dict(zone_dict, row, i, zone)
elif (zone_dict[zone]["date"] < row["last available data"])\
or (zone_dict[zone]["date"] == row["last available data"]
and zone_dict[zone]["version"] < row["version"]):
add_to_zone_dict(zone_dict, row, i, zone)
# -
indices = unique([zone_dict[d]['index'] for d in zone_dict])
country_2_region = {}
for idx in indices:
for country in analysis_set.loc[idx,].model.data['ifr']['country'].unique():
country_2_region[country] = {}
for _, row in analysis_set.loc[idx,].model.data['ifr'].iterrows():
country_2_region[row['country']][row['region']] = zone_dict[row['region']]['index']
# ## Plotting functions
# First we define a custom process that will print for each zone.
def process_zone(zone, zone_dict, model_data, title_ext=""):
display(Markdown(
f"### {zone}{title_ext} \n\n Simulation sur la base de donnees du {zone_dict['date']}"
+ f" avec la version {zone_dict['version']} du code d'Imperial College a Londres"
))
axs = plot_zones_summary(zone, model_data)
axis_date_limits(axs, max_date='2020-05-20')
translate_axes_to_french(axs)
plt.show()
plt.close()
def print_zone(file_in, img_dir, file_dir, zone, zone_dict, model_data, title_ext="", img_ext=".png"):
img_file = os.path.join(img_dir, (zone + '_' + zone_dict['date'].strip() + img_ext).replace(" ", "_"))
unix_rel_img_path = os.path.relpath(img_file, file_dir).replace('\\','/')
file_in.write(
f"### {zone}{title_ext} \n\n Simulation sur la base de données du {zone_dict['date']}"
+ f" avec la version {zone_dict['version']} du code d'Imperial College à Londres"
+ f" \n\n \n\n"
)
axs = plot_zones_summary(zone, model_data)
axis_date_limits(axs, max_date='2020-05-20')
translate_axes_to_french(axs)
axs[0].figure.savefig(img_file, bbox_inches='tight')
plt.close(axs[0].figure)
# And then we apply this function to every zone that has been identified.
def apply_process_to_France(
process_func, country_2_region, analysis_set, zone_dict,
display_func=lambda x:display(Markdown(x))):
country = "France"
display_func(f"# Rapport de simulation sur la progression et l'impact du COViD19 en {country}\n\n")
display_func(
"Ce rapport présente les resultats du travail de réplication d'Imperial College sur la progression de l'épidémie COVID-19 en Europe. Ce travail, reprend les mêmes méthodes et les appliquent à la France et ses régions.\n\n" +
"Toutes les régions françaises sont passées au crible. À chaque fois trois graphiques illustrent la progression du nombre de décès, un quatrième montre l'effondrement du taux de reproduction initial et le dernier extrapole la progression du nombre de cas confirmés, à partir d'un taux de mortalité estimé à 1,12% (scénario plutôt pessimiste).\n\n" +
"D'une manière générale le projet vise à estimer au plus près l'impact de cinq interventions politiques (distanciation physique, confinement, annulation des grands événements, fermeture des écoles et université, mise en quarantaine des personnes infectée) sur la courbe des décès liés à la pandémie. En ce sens, les données de 'Google Community Reports' sont en cours d'intégration afin d'améliorer la capacité prédictive du modèle.\n\n" +
"Les prédictions presentées dans les graphes ci-dessous (lignes pointillées) ne prennent pas en compte le déconfinement. Elles ne sont présentées ici que pour accentuer les tendances observées durant le confinement.\n\n"
)
display_func(f"## Résultats au niveau du Pays\n\n")
display_func(
"Dans cette section vous retrouverez des délimitations non géographiques"
+ "de la population, nottament une modelisation sur la base des données:\n\n"
+ "- Santé publique France (SPF) via le ECDC sous la dénomination 'France'\n"
+ "- SPF via la collecte de opencovid19-fr sous la dénomination 'France-OC19'\n"
+ "- Des décès en EHPAD sous la dénomination 'France-EHPAD'\n"
+ "- Des décès en hopitaux sous la dénomination 'France-hopitaux'\n\n"
)
country_level_list = []
region_level_list = []
for region in country_2_region[country]:
if region[:len(country)] == country:
country_level_list.append(region)
else:
region_level_list.append(region)
for zone in sorted(country_level_list):
idx = zone_dict[zone]['index']
process_func(
zone, zone_dict[zone],
analysis_set.loc[idx, 'model'].data)
display_func(f"## Résultats régionaux \n")
for zone in sorted(region_level_list):
idx = zone_dict[zone]['index']
process_func(
zone, zone_dict[zone],
analysis_set.loc[idx, 'model'].data)
# +
#apply_process_to_France(process_zone, country_2_region, analysis_set, zone_dict)
# +
report_dir = os.path.join('public_reports', 'france_report')
img_dir = os.path.join(report_dir, 'img')
os.makedirs(report_dir, exist_ok=True)
os.makedirs(img_dir, exist_ok=True)
report_file = os.path.join(report_dir, "france_report.md")
with open(report_file, "w", encoding="utf-8") as file_in:
print_partial = partial(print_zone, file_in, img_dir, report_dir)
apply_process_to_France(
print_partial, country_2_region, analysis_set, zone_dict,
file_in.write)
| france_report.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tutorial: Analytic continuation of SrVO$_3$ self-energy
# This tutorial covers, in a pedagogical way, a real-world use case of the `ana_cont` library.
# We will show how to read Matsubara data from a w2dynamics output hdf5 file and prepare the data for analytic continuation.
# Then, analytic continuation is performed by two different methods: MaxEnt and Pade.
# Finally, we can use the Kramers-Kronig relations to get the full complex self-energy, including real and imaginary part on the real-frequency axis.
# First, we need some standard imports: sys, os, numpy, h5py, matplotlib.pyplot.
#
# Then, we insert the path of the ana_cont repository into the python-path. This allows us to import ana_cont without having to install it. But note that, for using Pade later in the notebook, it is necessary that you have already compiled it (`python setup.py build_ext --inplace`)
import sys, os
import numpy as np
import h5py
import matplotlib.pyplot as plt
sys.path.insert(0, os.environ['HOME'] + '/Programs/ana_cont')
import ana_cont.continuation as cont
# First, download the DMFT file [from here](https://github.com/josefkaufmann/ana_cont/wiki/datafiles/dmft_svo.hdf5).
#
# Load DMFT data from w2dynamics output file. It contains a single-shot symmetric improved estimators calculation on top of a converged DMFT calculation of SrVO$_3$ with three degenerate orbitals. The system is paramagnetic.
#
# If you have not saved the DMFT file in the current working directory, you have to add the path to it in the file name in the first line of the following cell.
f = h5py.File('dmft_svo.hdf5', 'r')
beta = f['.config'].attrs['general.beta']
siw_full = f['stat-001/ineq-001/siw-full/value'][()]
siw_err_full = f['stat-001/ineq-001/siw-full/error'][()]
smom = f['stat-001/ineq-001/smom/value'][()]
siw = np.diagonal(np.diagonal(siw_full, axis1=0, axis2=2), axis1=0, axis2=1).transpose((1, 2, 0))
err = np.diagonal(np.diagonal(siw_err_full, axis1=0, axis2=2), axis1=0, axis2=1).transpose((1, 2, 0))
smom = np.mean(smom, axis=(0, 1)) # degenerate spins and orbitals -> average over 6 components
siw = np.mean(siw, axis=(0, 1))
err = np.mean(err, axis=(0, 1)) / np.sqrt(6.)
niw_full = siw.shape[-1] // 2
f.close()
# Now prepare the data for analytic contination, i.e. subtract the Hartree term and select a Matsubara frequency range. We then plot the data to make sure we reach the asymptotic region. We also (logarithmically) plot the QMC error, which does not increase at high frequencies, because we have used symmetric improved estimators.
# +
niw = 250
siw_cont = siw[niw_full:niw_full+niw] - smom[0]
err_cont = err[niw_full:niw_full+niw]
iw = np.pi / beta * (2. * np.arange(niw) + 1.)
w = 15. * np.tan(np.linspace(-np.pi / 2.1, np.pi / 2.1, num=1001, endpoint=True)) / np.tan(np.pi / 2.1)
model = np.ones_like(w)
model /= np.trapz(model, w)
model *= -smom[1]
plt.plot(iw, siw_cont.real)
plt.plot(iw, siw_cont.imag)
plt.show()
plt.semilogy(iw, err_cont)
plt.show()
# -
# Define the `AnalyticContinuationProblem` for MaxEnt, and solve it. As a reasonable value for the blur width we take 0.16, but feel free to change it and see what happens.
probl_maxent = cont.AnalyticContinuationProblem(im_axis=iw, re_axis=w, im_data=siw_cont,
kernel_mode='freq_fermionic', beta=beta)
sol_maxent = probl_maxent.solve(method='maxent_svd', optimizer='newton', alpha_determination='chi2kink',
model=model, stdev=err_cont,
preblur=True, blur_width=0.16,
alpha_start=1e14, alpha_end=1e0, alpha_div=10., fit_position=2.,
interactive=True)
# Now we can plot the resulting spectral function. Here it is extremely important to look also at the backtransform (middle panel) and the difference of data and backtransform (right panel). The range between the error bars is shaded.
fig, ax = plt.subplots(ncols=3, nrows=1, figsize=(14, 4))
ax[0].plot(w, sol_maxent[0].A_opt)
ax[0].set_xlim(-5, 10)
ax[0].set_xlabel(r'$\omega$')
ax[0].set_ylabel(r'$-\frac{1}{\pi} \mathrm{Im} \Sigma_R(\omega)$')
ax[1].plot(iw, siw_cont.real, ls='None', marker='+', color='black')
ax[1].plot(iw, siw_cont.imag, ls='None', marker='x', color='black')
ax[1].plot(iw, sol_maxent[0].backtransform.real, color='red')
ax[1].plot(iw, sol_maxent[0].backtransform.imag, color='blue')
ax[1].set_xlabel(r'$i\omega_n$')
ax[1].set_ylabel(r'$\Sigma(i\omega_n)$')
ax[2].plot(iw, (siw_cont - sol_maxent[0].backtransform).real)
ax[2].plot(iw, (siw_cont - sol_maxent[0].backtransform).imag)
ax[2].fill_between(iw, -err_cont, err_cont, alpha=0.4)
ax[2].set_xlabel(r'$i\omega_n$')
plt.tight_layout()
plt.show()
# Perform the analytic continuation by Pade. The selected `indices_pade` represent a reasonable choice, but are by no means unique. Feel free to experiment with their number and spacing. Keep in mind, however, that the calculation time steeply increases when using more than approximately 25 data points on the imaginary axis.
#
# We plot the imaginary-axis data that were used for the calculation of the Pade coefficients. Also, we plot the Pade-interpolation on a fine grid on the imaginary axis. If there are pole-zero pairs along the imaginary axis as a consequence of noise, these are usually visible in such a plot. It may then be indicated to choose a different set of Matsubara frequencies, or get better data.
indices_pade = np.concatenate((np.arange(12), np.arange(12, 100, 8)))
print(indices_pade, indices_pade.shape)
iw_pade = iw[indices_pade]
siw_pade = siw_cont[indices_pade]
w_pade = np.linspace(-5., 10., num=401)
probl_pade = cont.AnalyticContinuationProblem(im_axis=iw_pade,
re_axis=w_pade,
im_data=siw_pade,
kernel_mode='freq_fermionic')
sol_pade = probl_pade.solve(method='pade')
check_axis = np.linspace(0., 1.25 * iw_pade[-1], num=500)
check = probl_pade.solver.check(im_axis_fine=check_axis)
plt.plot(iw_pade, siw_pade.imag, ls='None', marker='x')
plt.plot(iw, siw_cont.imag, ls='None', marker='.')
plt.plot(check_axis, check.imag)
plt.xlim(0., check_axis[-1])
plt.show()
# Plot both spectral function on the full frequency range and in the low-frequency region.
# +
plt.plot(w, sol_maxent[0].A_opt, label='maxent')
plt.plot(w_pade, sol_pade.A_opt, label='pade')
plt.xlim(-5., 10.)
plt.ylim(0., 2.)
plt.legend()
plt.show()
plt.plot(w, sol_maxent[0].A_opt, label='maxent')
plt.plot(w_pade, sol_pade.A_opt, label='pade')
plt.xlim(-1., 1.)
plt.ylim(0., 0.2)
plt.legend()
plt.show()
# -
# Just out of curiosity, also look at the QMC error (logplot!) at the points that were used for the Pade interpolation.
plt.semilogy(iw, err_cont)
plt.semilogy(iw_pade, err_cont[indices_pade], linestyle='None', marker='.')
plt.show()
| doc/tutorial_svo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
# machine learning models
from sklearn.linear_model import LogisticRegression, Perceptron, SGDClassifier
from sklearn.svm import SVC, LinearSVC
from sklearn.ensemble import RandomForestClassifier, ExtraTreesClassifier, GradientBoostingClassifier, VotingClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.naive_bayes import GaussianNB, MultinomialNB
from sklearn.tree import DecisionTreeClassifier
from sklearn.neural_network import MLPClassifier
#feature scaling
from sklearn.preprocessing import StandardScaler, RobustScaler
#Pipeline
from sklearn.pipeline import Pipeline, FeatureUnion
#Cross validation
from sklearn.model_selection import cross_val_score, train_test_split
#Model persistence
from sklearn.externals import joblib
# -
train = pd.read_csv('train.gz', index_col=0)
test = pd.read_csv('test.gz', index_col=0)
train.describe()
print(train.columns.values)
train.info()
#Let's drop all calculated features
col = [c for c in train.columns if not c.startswith('ps_calc_')]
train=train[col]
col = [c for c in test.columns if not c.startswith('ps_calc_')]
test=test[col]
corr_matrix = train.corr()
corr_matrix["target"].sort_values(ascending=False)
# +
def change_datatype(df):
float_cols = list(df.select_dtypes(include=['int']).columns)
for col in float_cols:
if ((np.max(df[col]) <= 127) and(np.min(df[col] >= -128))):
df[col] = df[col].astype(np.int8)
elif ((np.max(df[col]) <= 32767) and(np.min(df[col] >= -32768))):
df[col] = df[col].astype(np.int16)
elif ((np.max(df[col]) <= 2147483647) and(np.min(df[col] >= -2147483648))):
df[col] = df[col].astype(np.int32)
else:
df[col] = df[col].astype(np.int64)
change_datatype(train)
change_datatype(test)
# +
def change_datatype_float(df):
float_cols = list(df.select_dtypes(include=['float']).columns)
for col in float_cols:
df[col] = df[col].astype(np.float32)
change_datatype_float(train)
change_datatype_float(test)
# -
train, validation, target, target_val = train_test_split(train, train['target'], test_size=0.10, random_state=42)
# drop id y target from train and validation sets
train_tr=train.drop("target", axis=1).drop("id", axis=1)
validation_tr=validation.drop("target", axis=1).drop("id", axis=1)
test_tr=test.drop("id", axis=1)
# +
# Normalize the data
scaler=StandardScaler()
train_tr = scaler.fit_transform(train_tr)
validation_tr = scaler.fit_transform(validation_tr)
test_tr = scaler.fit_transform(test_tr)
# -
train_tr.shape, validation_tr.shape, test_tr.shape
# +
# Gaussian Naive Bayes
gaussian = GaussianNB()
# %time gaussian.fit(train_tr, target)
# %time prediction = gaussian.predict_proba(test_tr)
prediction_nb=prediction[:,1]
joblib.dump(gaussian, 'gaussian.pkl', compress=True)
# %time acc_gaussian = (cross_val_score(gaussian, train_tr, target, cv=5, scoring="accuracy").mean()) * 100
acc_gaussian
# -
np.savez('prediction_nb.npz', prediction_nb)
# +
# Submission
# He didn't confess yet, but he will...
# -
submission = pd.DataFrame({
"id": test["id"],
"target": prediction_nb
})
submission.to_csv('submission_nb.csv', index=False)
| Porto Seguro/Naive Bayes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (Data Languages Lecture)
# language: python
# name: datalanguages
# ---
# # `Astropy` models and fitting
# If you need to do least square fitting for data to a model a good place to start is `astropy`'s modeling and fitting code.
#
# ## Packages being used
# + `astropy`: for modeling and fitting
# + `matplotlib`: for plotting
#
# ## Relevant documentation
# + `astropy`: http://docs.astropy.org/en/stable/modeling/index.html
import numpy as np
import matplotlib.pyplot as plt
from astropy.modeling import models, fitting
import mpl_style
# %matplotlib inline
plt.style.use(mpl_style.style1)
# ## 1-D model fitting
# For an example lets look at the problem of fitting a 1-D model to a spectral line. First we need to create some fake data:
x = np.linspace(-5., 5., 200)
y = 3 * np.exp(-0.5 * (x - 1.3)**2 / 0.8**2)
y += np.random.normal(0., 0.2, x.shape)
# ### A trapezoid model
t_init = models.Trapezoid1D(amplitude=1.0, x_0=0.1, slope=0.5)
fit_t = fitting.LevMarLSQFitter()
t = fit_t(t_init, x, y)
print(t)
# ### A Gaussian model
g_init = models.Gaussian1D(amplitude=1., mean=0, stddev=1.)
fit_g = fitting.LevMarLSQFitter()
g = fit_g(g_init, x, y)
print(g)
# ### Plotting the results
plt.figure(1, figsize=(8,5))
plt.plot(x, y, 'o', mfc='none')
plt.plot(x, t(x), label='Trapezoid')
plt.plot(x, g(x), label='Gaussian')
plt.xlabel('Position')
plt.ylabel('Flux')
plt.legend(loc=2)
plt.tight_layout()
# ## Compound models
# Models can also be 'added' together before fitting. To demonstrate lets make a new dataset made up to two Gaussians.
np.random.seed(42)
g1 = models.Gaussian1D(1, 0, 0.2)
g2 = models.Gaussian1D(2.5, 0.5, 0.1)
x = np.linspace(-1, 1, 200)
y = g1(x) + g2(x) + np.random.normal(0., 0.2, x.shape)
# ### Make the model
# The model can be 'added' just like arrays:
gg_init = models.Gaussian1D(1, 0, 0.1) + models.Gaussian1D(2, 0.5, 0.1)
fit_gg = fitting.SLSQPLSQFitter()
gg = fit_gg(gg_init, x, y)
print(gg)
# ### Plot the result
plt.figure(2, figsize=(8, 5))
plt.plot(x, y, 'o', mfc='none')
plt.plot(x, gg(x), label='2 x Gaussian')
plt.xlabel('Position')
plt.ylabel('Flux')
plt.legend(loc=2)
plt.tight_layout()
# ## `Astropy`'s models
# `Astropy` has a large number of 1- and 2-D models built in. Check out https://docs.astropy.org/en/stable/modeling/index.html#module-astropy.modeling.functional_models for a full list. If the model you are looking for is not built in, you can always define your own: https://docs.astropy.org/en/stable/modeling/new-model.html.
#
# ## Limitations
# + Uses OLS (or similar) to maximize an objective function (and all the assumptions about the data that go into this, e.g. Gaussian errors)
# + Cov of fit only returned for some fitters (fond on the `fitter.fit_info()` method)
| Astropy_fitting.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Regressione lineare su polinomio con determinazione della distribuzione predittiva
# %matplotlib inline
import numpy as np
import scipy.stats as st
# +
import matplotlib.pyplot as plt
import matplotlib.colors as mcolors
from matplotlib import cm
plt.style.use('fivethirtyeight')
plt.rcParams['font.family'] = 'sans-serif'
plt.rcParams['font.serif'] = 'Ubuntu'
plt.rcParams['font.monospace'] = 'Ubuntu Mono'
plt.rcParams['font.size'] = 10
plt.rcParams['axes.labelsize'] = 10
plt.rcParams['axes.labelweight'] = 'bold'
plt.rcParams['axes.titlesize'] = 10
plt.rcParams['xtick.labelsize'] = 8
plt.rcParams['ytick.labelsize'] = 8
plt.rcParams['legend.fontsize'] = 10
plt.rcParams['figure.titlesize'] = 12
plt.rcParams['image.cmap'] = 'jet'
plt.rcParams['image.interpolation'] = 'none'
plt.rcParams['figure.figsize'] = (12, 6)
plt.rcParams['lines.linewidth'] = 2
colors = ['xkcd:pale orange', 'xkcd:sea blue', 'xkcd:pale red', 'xkcd:sage green', 'xkcd:terra cotta', 'xkcd:dull purple', 'xkcd:teal', 'xkcd:goldenrod', 'xkcd:cadet blue',
'xkcd:scarlet']
cmap_big = cm.get_cmap('Spectral', 512)
cmap = mcolors.ListedColormap(cmap_big(np.linspace(0.7, 0.95, 256)))
bbox_props = dict(boxstyle="round,pad=0.3", fc=colors[0], alpha=.5)
# -
# $d$ funzioni base gaussiane, con medie intervallate in modo costante nel dominio considerato e varianza unitaria
def vphi(x, d, dom):
l = np.linspace(domain[0], domain[1], d+1)
mus = [(l[i]+l[i+1])/2.0 for i in range(len(l)-1)]
return np.array([gaussian_basis(x, mus[i], 1) for i in range(d)]).T
# Funzione base gaussiana
def gaussian_basis(x, m, s):
return np.exp(-((x-m)**2)/(2*s**2))
# Genera la matrice delle features e il vettore target
# +
# dominio della feature
domain=(0,2*np.pi)
# numero di elementi da generare
n=30
# array delle feature generato uniformemente nel dominio
X=np.random.uniform(domain[0], domain[1], n)
# genera il vettore target mediante la funzione f e l'aggiunta di rumore gaussiano
# funzione
def f(x):
return np.sin(x)
# sd del rumore
noise = .05
#genera target
t=np.array([(f(v)+np.random.normal(0,noise,1))[0] for v in X]).reshape(-1,1)
# numero di funzioni base
d=8
# genera immagine di X per la regressione
Phi = vphi(X,d, domain)
# -
# Iperparametri
# +
# iperparametro per il prior
alfa=.2
# parametri del prior
mu=np.zeros(d+1)
sigma=np.eye(d+1)*alfa
# parametro per la verosimiglianza
beta=9
# -
# Distribuzione predittiva dato un valore $v$
# media della distribuzione predittiva
def m_pred(v):
return m.T.dot(vphi(v,d,domain))
# varianza della distribuzione predittiva
def var_pred(v):
v1=vphi(v,d,domain)
return 1.0/beta+v1.dot(s.dot(v1.T))
# +
# numero elementi considerati per il training
l=2
# estrazione del training set
X_t, t_t = Phi[:l,:], t[:l]
# derivazione di media e matrice di covarianza a posteriori
s = np.linalg.inv(np.eye(d)+beta*np.dot(X_t.T,X_t))
m=beta*s.dot(X_t.T.dot(t_t))
# -
# Plot della distribuzione predittiva
# +
# insieme dei valori considerati per il plot
xx=np.linspace(min(X),max(X),500)
# loro immagini per il calcolo della regressione
phix=vphi(xx,d,domain)
# calcolo di media e varianza della distribuzione predittiva per tutti i valori
mx = np.vectorize(m_pred)(xx)
sx= np.vectorize(var_pred)(xx)
# visualizzazione
fig = plt.figure(figsize=(16,8))
ax = fig.gca()
# plot della media
ax.plot(xx,mx,'-', c=colors[9], alpha=1)
# riempimento della regione a distanza minore di una sd dalla media
ax.fill_between(xx, mx-np.sqrt(sx), mx+np.sqrt(sx), facecolor=colors[9], alpha=.05)
# elementi dell'insieme
ax.scatter(X[l:], t[l:], c=colors[0], marker='o', alpha=1)
ax.scatter(X[:l], t[:l], c=colors[1], marker='o', alpha=1)
# plot funzione originale
ax.plot(xx,f(xx),'--',c=colors[1],alpha=1)
plt.xlabel(u'$x$', fontsize=10)
plt.ylabel(u'$y$', fontsize=10)
plt.xticks(fontsize=10)
plt.yticks(fontsize=10)
plt.show()
# -
| codici/linregr_predict.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="4VivGJAWcRNn" colab_type="text"
# ### This notebook is optionally accelerated with a GPU runtime.
# ### If you would like to use this acceleration, please select the menu option "Runtime" -> "Change runtime type", select "Hardware Accelerator" -> "GPU" and click "SAVE"
#
# ----------------------------------------------------------------------
#
# # BERT
#
# *Author: HuggingFace Team*
#
# **Bidirectional Encoder Representations from Transformers.**
#
# _ | _
# - | -
# 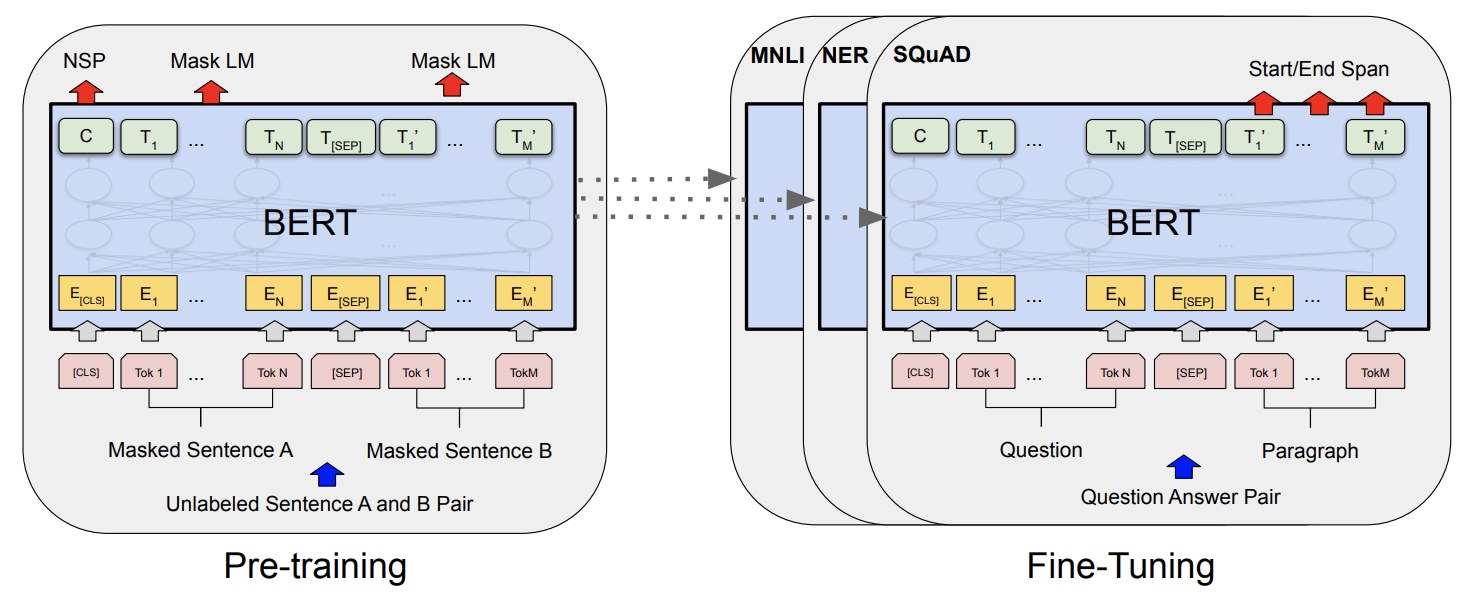 | 
#
#
# ### Model Description
#
# BERT was released together with the paper [BERT: Pre-training of Deep Bidirectional Transformers for Language Understanding](https://arxiv.org/abs/1810.04805) by <NAME> et al. The model is based on the Transformer architecture introduced in [Attention Is All You Need](https://arxiv.org/abs/1706.03762) by <NAME> et al and has led to significant improvements on a wide range of downstream tasks.
#
# Here are 8 models based on BERT with [Google's pre-trained models](https://github.com/google-research/bert) along with the associated Tokenizer.
# It includes:
# - `bertTokenizer`: perform end-to-end tokenization, i.e. basic tokenization followed by WordPiece tokenization
# - `bertModel`: raw BERT Transformer model (fully pre-trained)
# - `bertForMaskedLM`: BERT Transformer with the pre-trained masked language modeling head on top (fully pre-trained)
# - `bertForNextSentencePrediction`: BERT Transformer with the pre-trained next sentence prediction classifier on top (fully pre-trained)
# - `bertForPreTraining`: BERT Transformer with masked language modeling head and next sentence prediction classifier on top (fully pre-trained)
# - `bertForSequenceClassification`: BERT Transformer with a sequence classification head on top (BERT Transformer is pre-trained, the sequence classification head is only initialized and has to be trained)
# - `bertForMultipleChoice`: BERT Transformer with a multiple choice head on top (used for task like Swag) (BERT Transformer is pre-trained, the multiple choice classification head is only initialized and has to be trained)
# - `bertForTokenClassification`: BERT Transformer with a token classification head on top (BERT Transformer is pre-trained, the token classification head is only initialized and has to be trained)
# - `bertForQuestionAnswering`: BERT Transformer with a token classification head on top (BERT Transformer is pre-trained, the token classification head is only initialized and has to be trained)
#
# ### Requirements
#
# Unlike most other PyTorch Hub models, BERT requires a few additional Python packages to be installed.
# + id="mY19uZF8cRNw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 391} outputId="a2fc46a7-bdf9-46fd-ec9c-465d5c81009c" language="bash"
#
# pip install tqdm boto3 requests regex
# + id="HdLb-2MffrOG" colab_type="code" colab={}
from sklearn.model_selection import train_test_split
import pandas as pd
import tensorflow as tf
import tensorflow_hub as hub
from datetime import datetime
from tensorflow import keras
import os
import re
# Load all files from a directory in a DataFrame.
def load_directory_data(directory):
data = {}
data["sentence"] = []
data["sentiment"] = []
for file_path in os.listdir(directory):
with tf.gfile.GFile(os.path.join(directory, file_path), "r") as f:
data["sentence"].append(f.read())
data["sentiment"].append(re.match("\d+_(\d+)\.txt", file_path).group(1))
return pd.DataFrame.from_dict(data)
# Merge positive and negative examples, add a polarity column and shuffle.
def load_dataset(directory):
pos_df = load_directory_data(os.path.join(directory, "pos"))
neg_df = load_directory_data(os.path.join(directory, "neg"))
pos_df["polarity"] = 1
neg_df["polarity"] = 0
return pd.concat([pos_df, neg_df]).sample(frac=1).reset_index(drop=True)
# Download and process the dataset files.
def download_and_load_datasets(force_download=False):
dataset = tf.keras.utils.get_file(
fname="aclImdb.tar.gz",
origin="http://ai.stanford.edu/~amaas/data/sentiment/aclImdb_v1.tar.gz",
extract=True)
train_df = load_dataset(os.path.join(os.path.dirname(dataset),
"aclImdb", "train"))
test_df = load_dataset(os.path.join(os.path.dirname(dataset),
"aclImdb", "test"))
return train_df, test_df
# + id="OtCFIhO6ftDq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="f15c1747-9734-4301-af86-7c72a59d5c57"
train, test = download_and_load_datasets()
# + id="_v0a9QtZgELi" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="68dbb643-6819-48e2-877b-60615957161b"
train.head()
# + id="ogGE5S4gx0aK" colab_type="code" colab={}
# + id="WPMcBSbG2pZ9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 207} outputId="44f90409-d901-4cfd-aedd-0ad925cdba83"
[i for i in train['sentence'][0:10]]
# + id="1_ZdxmqoxFkC" colab_type="code" colab={}
trainloader = torch.utils.data.DataLoader(train, batch_size=4,
shuffle=True, num_workers=2)
# + id="GCVJ_Ntmxo5z" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 163} outputId="abbefa56-c4a0-4493-c3fe-47582590de8f"
bert.run_classifer.InputExample
# + [markdown] id="6TomBlhNcRN6" colab_type="text"
# ### Example
#
# Here is an example on how to tokenize the input text with `bertTokenizer`, and then get the hidden states computed by `bertModel` or predict masked tokens using `bertForMaskedLM`. The example also includes snippets showcasing how to use `bertForNextSentencePrediction`, `bertForQuestionAnswering`, `bertForSequenceClassification`, `bertForMultipleChoice`, `bertForTokenClassification`, and `bertForPreTraining`.
# + id="uBHCeul6cRN8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 105} outputId="3ccb8274-8714-45c0-fb17-e0a83f81975b"
### First, tokenize the input
import torch
tokenizer = torch.hub.load('huggingface/pytorch-pretrained-BERT', 'bertTokenizer', 'bert-base-cased', do_basic_tokenize=False)
# Tokenized input
text = "[CLS] Who was <NAME> ? [SEP] <NAME> was a puppeteer [SEP]"
tokenized_text = tokenizer.tokenize(text)
indexed_tokens = tokenizer.convert_tokens_to_ids(tokenized_text)
# + id="oNMdHnSKglRK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="b78c9958-22a3-4472-9123-8fec6382fdcb"
print(text)
print(tokenized_text)
print(indexed_tokens)
# + id="u3IRRm8tcROD" colab_type="code" colab={}
### Get the hidden states computed by `bertModel`
# Define sentence A and B indices associated to 1st and 2nd sentences (see paper)
segments_ids = [0, 0, 0, 0, 0, 0, 0, 0, 1, 1, 1, 1, 1, 1, 1, 1]
# Convert inputs to PyTorch tensors
segments_tensors = torch.tensor([segments_ids])
tokens_tensor = torch.tensor([indexed_tokens])
# + id="gcgpfbsHs3OY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="a75fd9d2-a13f-40fd-eddb-b6519b773187"
print(tokenizer.encode("Hello, my dog is cute"))
print(tokenizer.convert_tokens_to_ids(tokenizer.tokenize("Hello, my dog is cute")))
# + id="OV0H7IN7tQ9g" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="981fc613-fc3f-4715-e814-54e0233b5d1c"
model = torch.hub.load('huggingface/pytorch-pretrained-BERT', 'bertForSequenceClassification', 'bert-base-cased', num_labels=2)
input_id = torch.tensor(tokenizer.encode("Hello, my dog is cute")).unsqueeze(0) #batch_size 1
label = torch.tensor(1).unsqueeze(0) # batch_size 1
outputs = model(input_id, labels = label)
loss, logit = outputs
# + id="6pq3QM7zvRhg" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="ee91a02b-4b25-46c7-d90a-453b59fe73c1"
torch.nn.functional.softmax(logit, dim=1)
# + id="3GmdZblfv0cJ" colab_type="code" colab={}
lr = 1e-3
num_total_steps = 1000
num_warmup_steps = 100
warmup_proportion = float(num_warmup_steps) / float(num_total_steps) # 0.1
### Previously BertAdam optimizer was instantiated like this:
optimizer = BertAdam(model.parameters(), lr=lr, schedule='warmup_linear', warmup=warmup_proportion, t_total=num_total_steps)
### and used like this:
for batch in train_data:
loss = model(batch)
loss.backward()
optimizer.step()
# + id="H_eQyZpLcROl" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="115f47f1-3e94-491a-94a3-f0238582e968"
model = torch.hub.load('huggingface/pytorch-pretrained-BERT', 'bertForSequenceClassification', 'bert-base-cased', num_labels=2)
model.eval()
# Predict the sequence classification logits
with torch.no_grad():
seq_classif_logits = model(tokens_tensor, segments_tensors)
# Or get the sequence classification loss (set model to train mode before if used for training)
labels = torch.tensor([1])
seq_classif_loss = model(tokens_tensor, segments_tensors, labels=labels)
| huggingface_pytorch_pretrained_bert_bert.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + raw_mimetype="text/restructuredtext" active=""
# .. _nb_misc:
# -
# ## Miscellaneous
# Topics which did not directly fit into on of our main categories:
# + raw_mimetype="text/restructuredtext" active=""
#
# .. toctree::
# :maxdepth: 1
#
# termination_criterion.ipynb
# reference_directions.ipynb
# constraint_handling.ipynb
# decomposition.ipynb
# callback.ipynb
# display.ipynb
# results.ipynb
#
#
| doc/source/misc/index.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="ICqO4nV1fKrA" colab_type="code" colab={}
import pandas as pd
import numpy as np
from sklearn.tree import DecisionTreeRegressor
from sklearn.metrics import mean_absolute_error
from sklearn.model_selection import cross_val_score
# + id="u3KgcHVDga2v" colab_type="code" outputId="b9c54c8f-6826-4e8e-fda6-19bd182a8596" executionInfo={"status": "ok", "timestamp": 1581630275609, "user_tz": -60, "elapsed": 5909, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBbVS0exRFMvJLxGv-ZtTyGfxdhMPjX7mq78-H-4MQ=s64", "userId": "08144210629576263422"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# cd "/content/drive/My Drive/Colab Notebooks/dataworkshop_matrix"
# + id="Bw-DwQ91gsll" colab_type="code" outputId="373a5ceb-d499-414d-e9d7-efc092e84568" executionInfo={"status": "ok", "timestamp": 1581630363848, "user_tz": -60, "elapsed": 1248, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBbVS0exRFMvJLxGv-ZtTyGfxdhMPjX7mq78-H-4MQ=s64", "userId": "08144210629576263422"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
df = pd.read_csv('data/men_shoes.csv', low_memory = False)
df.shape
# + id="grDzB6BehFJR" colab_type="code" outputId="175009ca-ccf2-4fc1-9046-d6e5651cea74" executionInfo={"status": "ok", "timestamp": 1581630384668, "user_tz": -60, "elapsed": 622, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBbVS0exRFMvJLxGv-ZtTyGfxdhMPjX7mq78-H-4MQ=s64", "userId": "08144210629576263422"}} colab={"base_uri": "https://localhost:8080/", "height": 221}
df.columns
# + id="gYlaIbvIhO8N" colab_type="code" outputId="c3146552-1a3f-42d3-8ee4-c5333bd0dba8" executionInfo={"status": "ok", "timestamp": 1581630483638, "user_tz": -60, "elapsed": 626, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBbVS0exRFMvJLxGv-ZtTyGfxdhMPjX7mq78-H-4MQ=s64", "userId": "08144210629576263422"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
mean_price = np.mean(df['prices_amountmin'])
mean_price
# + id="KUPuBDDqhfgd" colab_type="code" outputId="b3683ac9-cb38-4ac8-99de-3d9363d7256e" executionInfo={"status": "ok", "timestamp": 1581630664351, "user_tz": -60, "elapsed": 659, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBbVS0exRFMvJLxGv-ZtTyGfxdhMPjX7mq78-H-4MQ=s64", "userId": "08144210629576263422"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
y_true = df['prices_amountmin']
y_pred = [mean_price] * y_true.shape[0]
mean_absolute_error(y_true, y_pred)
# + id="rVvw2x3MiZAb" colab_type="code" outputId="426aa7d0-a689-4b32-80a8-71f66ab4b101" executionInfo={"status": "ok", "timestamp": 1581630748456, "user_tz": -60, "elapsed": 1043, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBbVS0exRFMvJLxGv-ZtTyGfxdhMPjX7mq78-H-4MQ=s64", "userId": "08144210629576263422"}} colab={"base_uri": "https://localhost:8080/", "height": 282}
df['prices_amountmin'].hist(bins=100)
# + id="SQCkYlMbij48" colab_type="code" outputId="0d288255-4ea6-48f7-894f-5eefb5645ad0" executionInfo={"status": "ok", "timestamp": 1581630871937, "user_tz": -60, "elapsed": 990, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBbVS0exRFMvJLxGv-ZtTyGfxdhMPjX7mq78-H-4MQ=s64", "userId": "08144210629576263422"}} colab={"base_uri": "https://localhost:8080/", "height": 282}
np.log1p( df['prices_amountmin'] ).hist(bins=100)
# + id="_hfWpz40jMYq" colab_type="code" outputId="a4ca6947-3e13-4e25-c387-b7b6241f5c6b" executionInfo={"status": "ok", "timestamp": 1581630987555, "user_tz": -60, "elapsed": 651, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBbVS0exRFMvJLxGv-ZtTyGfxdhMPjX7mq78-H-4MQ=s64", "userId": "08144210629576263422"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
y_true = df['prices_amountmin']
y_pred = [np.median(y_true)] * y_true.shape[0]
mean_absolute_error(y_true, y_pred)
# + id="zJQh9P2sj8RQ" colab_type="code" outputId="4f21999e-7022-433f-c2ce-03658d840e50" executionInfo={"status": "ok", "timestamp": 1581631433934, "user_tz": -60, "elapsed": 839, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBbVS0exRFMvJLxGv-ZtTyGfxdhMPjX7mq78-H-4MQ=s64", "userId": "08144210629576263422"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
y_true = df['prices_amountmin']
price_log_mean = np.expm1( np.mean ( np.log1p(y_true) ) )
y_pred = [price_log_mean] * y_true.shape[0]
mean_absolute_error(y_true, y_pred)
# + id="Q66jCNIYlPYr" colab_type="code" outputId="fc89e9ec-10ec-4702-a927-9e83c3ad219a" executionInfo={"status": "ok", "timestamp": 1581631475063, "user_tz": -60, "elapsed": 633, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBbVS0exRFMvJLxGv-ZtTyGfxdhMPjX7mq78-H-4MQ=s64", "userId": "08144210629576263422"}} colab={"base_uri": "https://localhost:8080/", "height": 221}
df.columns
# + id="-dzb9t2vlRu4" colab_type="code" outputId="ae44a911-48f0-47a3-de8b-260b654775ed" executionInfo={"status": "ok", "timestamp": 1581631525010, "user_tz": -60, "elapsed": 4055, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBbVS0exRFMvJLxGv-ZtTyGfxdhMPjX7mq78-H-4MQ=s64", "userId": "08144210629576263422"}} colab={"base_uri": "https://localhost:8080/", "height": 221}
df.brand.value_counts()
# + id="8INWBYoLljb5" colab_type="code" outputId="1e3785b9-800d-4f3b-896b-3526b5b3a4fc" executionInfo={"status": "ok", "timestamp": 1581631637970, "user_tz": -60, "elapsed": 652, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBbVS0exRFMvJLxGv-ZtTyGfxdhMPjX7mq78-H-4MQ=s64", "userId": "08144210629576263422"}} colab={"base_uri": "https://localhost:8080/", "height": 136}
df['brand'].factorize()
# + id="KKPNZqoCl-lt" colab_type="code" colab={}
df['brand_cat'] = df['brand'].factorize()[0]
# + id="WB8VO0OBmdV7" colab_type="code" outputId="f6549a0f-6f8a-468d-c5a3-e6fe3adb2b24" executionInfo={"status": "ok", "timestamp": 1581632959734, "user_tz": -60, "elapsed": 621, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBbVS0exRFMvJLxGv-ZtTyGfxdhMPjX7mq78-H-4MQ=s64", "userId": "08144210629576263422"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
feats = ['brand_cat']
x = df[ feats ].values
y = df['prices_amountmin'].values
model = DecisionTreeRegressor(max_depth=5)
scores = cross_val_score(model, x, y, scoring = 'neg_mean_absolute_error')
np.mean(scores), np.std(scores)
# + id="Gc19FuwqpiG6" colab_type="code" outputId="3fb29662-51e7-48b7-be3e-1d460a41f0a1" executionInfo={"status": "ok", "timestamp": 1581632817341, "user_tz": -60, "elapsed": 692, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBbVS0exRFMvJLxGv-ZtTyGfxdhMPjX7mq78-H-4MQ=s64", "userId": "08144210629576263422"}} colab={"base_uri": "https://localhost:8080/", "height": 54}
import sklearn
sklearn.metrics.SCORERS.keys()
# + id="GKjagqgarN6d" colab_type="code" colab={}
def run_model(feats):
x = df[ feats ].values
y = df['prices_amountmin'].values
model = DecisionTreeRegressor(max_depth=5)
scores = cross_val_score(model, x, y, scoring = 'neg_mean_absolute_error')
return np.mean(scores), np.std(scores)
# + id="urC7SAS7rjoT" colab_type="code" outputId="14a5a7a2-352c-46a9-9329-c3e20827a776" executionInfo={"status": "ok", "timestamp": 1581633172537, "user_tz": -60, "elapsed": 627, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBbVS0exRFMvJLxGv-ZtTyGfxdhMPjX7mq78-H-4MQ=s64", "userId": "08144210629576263422"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
run_model(['brand_cat'])
# + id="WfZLUYNnsYjL" colab_type="code" outputId="acb1f3d6-7d37-40ba-ed1c-3d87bef12c1b" executionInfo={"status": "ok", "timestamp": 1581633356438, "user_tz": -60, "elapsed": 639, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBbVS0exRFMvJLxGv-ZtTyGfxdhMPjX7mq78-H-4MQ=s64", "userId": "08144210629576263422"}} colab={"base_uri": "https://localhost:8080/", "height": 221}
df.manufacturer.value_counts()
# + id="7jWbY270sz75" colab_type="code" outputId="a10ff407-30bd-4831-b7e6-0b1ab3e26dea" executionInfo={"status": "ok", "timestamp": 1581633463567, "user_tz": -60, "elapsed": 647, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBbVS0exRFMvJLxGv-ZtTyGfxdhMPjX7mq78-H-4MQ=s64", "userId": "08144210629576263422"}} colab={"base_uri": "https://localhost:8080/", "height": 153}
df['manufacturer'].factorize()
# + id="xqmQHmP5s-8J" colab_type="code" colab={}
df['manufacturer_type'] = df['manufacturer'].factorize()[0]
# + id="P3aQErR-tJc3" colab_type="code" colab={}
def run_model(feats):
x = df[ feats ].values
y = df['prices_amountmin'].values
model = DecisionTreeRegressor(max_depth=5)
scores = cross_val_score(model, x, y, scoring = 'neg_mean_absolute_error')
return np.mean(scores), np.std(scores)
# + id="-wzxo7XNtVcQ" colab_type="code" outputId="63923ac8-1e5e-487d-eae6-fe1291d7e7c1" executionInfo={"status": "ok", "timestamp": 1581633606998, "user_tz": -60, "elapsed": 644, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBbVS0exRFMvJLxGv-ZtTyGfxdhMPjX7mq78-H-4MQ=s64", "userId": "08144210629576263422"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
run_model(['manufacturer_type'])
# + id="MOrf548RtcWv" colab_type="code" outputId="88895164-0545-4d34-c13c-271e718a091f" executionInfo={"status": "ok", "timestamp": 1581633669134, "user_tz": -60, "elapsed": 619, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBbVS0exRFMvJLxGv-ZtTyGfxdhMPjX7mq78-H-4MQ=s64", "userId": "08144210629576263422"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
run_model(['brand_cat', 'manufacturer_type'])
# + id="TwK_81veuA5H" colab_type="code" outputId="8f45ac5c-4632-43cb-8b29-e784a8a028ef" executionInfo={"status": "ok", "timestamp": 1581633781585, "user_tz": -60, "elapsed": 644, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBbVS0exRFMvJLxGv-ZtTyGfxdhMPjX7mq78-H-4MQ=s64", "userId": "08144210629576263422"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
run_model(['brand_cat'])
# + id="ksEXJCyGzZSZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="c3a877f2-175d-4df5-e811-f9f0f0e95c10" executionInfo={"status": "ok", "timestamp": 1581635204616, "user_tz": -60, "elapsed": 12378, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBbVS0exRFMvJLxGv-ZtTyGfxdhMPjX7mq78-H-4MQ=s64", "userId": "08144210629576263422"}}
# ls
# + id="CWw2lzx6zlfs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="d2a024ea-89d9-4733-ca44-d3c7501122ad" executionInfo={"status": "ok", "timestamp": 1581635244696, "user_tz": -60, "elapsed": 4194, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBbVS0exRFMvJLxGv-ZtTyGfxdhMPjX7mq78-H-4MQ=s64", "userId": "08144210629576263422"}}
# ls day4_matrix_one_two
# + id="D-O8PFZjzrQs" colab_type="code" colab={}
# !git add day3_matrix_one/day3.ipynb
| day4_matrix_one_two/day4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# +
# Input data (baseline trial)
orig_data = pd.DataFrame(
data = np.array([ # Absorbance
[0.003, 0.076, 0.162],
[0.005, 0.126, 0.189],
[0.010, 0.127, 0.224],
[0.027, 0.173, 0.292],
[0.053, 0.183, 0.317]
]),
index = [ # Row labels = mL 25 mM NPP
0.05,
0.1,
0.2,
0.5,
1.0
],
columns = [ # Column labels: time (min)
0,
10,
20
]
)
orig_data.rename_axis(index='Substrate amt. (mL 25 mM NPP)',
columns='Time (min)')
# +
# Input data (inhibition trial)
inhb_data = pd.DataFrame(
data = np.array([ # Absorbance
[0.004, 0.011, 0.022],
[-0.002, 0.019, 0.053],
[0.009, 0.051, 0.103],
[0.018, 0.079, 0.131],
[0.043, 0.092, 0.193]
]),
index = [ # Row labels = mL 25 mM NPP
0.05,
0.1,
0.2,
0.5,
1.0
],
columns = [ # Column labels: time (min)
0,
10,
20
]
)
inhb_data.rename_axis(index='Substrate amt. (mL 25 mM NPP)',
columns='Time (min)')
# -
# Experimental constants
sample_vol = 0.0045 # (L) = 0.001 L/mL * 4.5 mL
absorptivity = 11.8e3 # cm^-1 M^-1 (measured in class by Prof.) SEE Exp 3-BC
# +
# Michaelis-Menten equation
def michaelis_menten_rxn_vel(Vmax, Km, S):
V0 = (Vmax * S) / (Km + S)
return V0
# residuals for fitting the michaelis-menten equation
def michaelis_menten_resid(params,
conc,
obs_vel):
Vmax = params[0]
Km = params[1]
return obs_vel - michaelis_menten_rxn_vel(Vmax, Km, conc)
# residuals for scaled values (can use w/ absorbance)
def absrb_fit_resid(params,
conc,
obs_absrb):
Vmax = params[0]
Km = params[1]
scl = params[2] # arbitrary scaling factor
return obs_absrb - (scl * michaelis_menten_rxn_vel(Vmax, Km, conc))
# +
# Scatter plot of Absorbance vs. Concentration
from scipy.optimize import least_squares
# Compute molar concentratio of substrate
# conc. of substrate stock sol'n * mL subst. stock * 0.001 L/mL / sample_vol
S = 0.025 * (0.001 * orig_data.index) / sample_vol # (mol/L)
plt.close()
plt.figure(figsize=(8.0,5.0))
plt.plot(S, orig_data[20] - orig_data[0], 'kx')
fit_orig = least_squares(absrb_fit_resid,
[1.0, 1.0, 1.0],
args=(S, orig_data[20] - orig_data[0]),
method='lm',
verbose=2)
ss = np.linspace(0,S.max()*1.1,50)
fit_orig_data = fit_orig.x[2] * michaelis_menten_rxn_vel(fit_orig.x[0], fit_orig.x[1], ss)
plt.plot(ss, fit_orig_data, 'k:')
plt.title('Absorbance vs. Concentration (at 20 min)')
plt.xlabel('Substrate Concentration [S] (mol/L)')
plt.ylabel('Absorbance (a.u.)')
plt.xlim([ss[0],ss[-1]])
ybnds = plt.gca().get_ylim()
plt.ylim([0,ybnds[1]])
plt.legend([
'Measured data',
'Scaled Michaelis-Menten\nderived least squares fit_orig'
], loc='lower right')
plt.gcf().set_facecolor('white')
plt.gcf().savefig('Abs_v_Conc_orig_20.png')
plt.show()
# +
# Scatter plot of Absorbance vs. Concentration
from scipy.optimize import least_squares
# Compute molar concentratio of substrate
# conc. of substrate stock sol'n * mL subst. stock * 0.001 L/mL / sample_vol
S = 0.025 * (0.001 * orig_data.index) / sample_vol # (mol/L)
plt.close()
plt.figure(figsize=(8.0,5.0))
plt.plot(S, orig_data[10] - orig_data[0], 'kx')
fit_orig = least_squares(absrb_fit_resid,
[1.0, 1.0, 1.0],
args=(S, orig_data[10] - orig_data[0]),
method='lm',
verbose=2)
ss = np.linspace(0,S.max()*1.1,50)
fit_orig_data = fit_orig.x[2] * michaelis_menten_rxn_vel(fit_orig.x[0], fit_orig.x[1], ss)
plt.plot(ss, fit_orig_data, 'k:')
plt.title('Absorbance vs. Concentration (at 10 min)')
plt.xlabel('Substrate Concentration [S] (mol/L)')
plt.ylabel('Absorbance (a.u.)')
plt.xlim([ss[0],ss[-1]])
ybnds = plt.gca().get_ylim()
plt.ylim([0,ybnds[1]])
plt.legend([
'Measured data',
'Scaled Michaelis-Menten\nderived least squares fit_orig'
], loc='lower right')
plt.gcf().set_facecolor('white')
plt.gcf().savefig('Abs_v_Conc_orig_10.png')
plt.show()
# +
# Scatter plot of Absorbance vs. Concentration
from scipy.optimize import least_squares
# Compute molar concentratio of substrate
# conc. of substrate stock sol'n * mL subst. stock * 0.001 L/mL / sample_vol
S = 0.025 * (0.001 * inhb_data.index) / sample_vol # (mol/L)
plt.close()
plt.figure(figsize=(8.0,5.0))
plt.plot(S, inhb_data[20] - inhb_data[0], 'kx')
fit_inhb = least_squares(absrb_fit_resid,
[1.0, 1.0, 1.0],
args=(S, inhb_data[20] - inhb_data[0]),
method='lm',
verbose=2)
ss = np.linspace(0,S.max()*1.1,50)
fit_inhb_data = fit_inhb.x[2] * michaelis_menten_rxn_vel(fit_inhb.x[0], fit_inhb.x[1], ss)
plt.plot(ss, fit_inhb_data, 'k:')
plt.title('Absorbance vs. Concentration (at 20 min)')
plt.xlabel('Substrate Concentration [S] (mol/L)')
plt.ylabel('Absorbance (a.u.)')
plt.xlim([ss[0],ss[-1]])
ybnds = plt.gca().get_ylim()
plt.ylim([0,ybnds[1]])
plt.legend([
'Measured data',
'Scaled Michaelis-Menten\nderived least squares fit_inhb'
], loc='lower right')
plt.gcf().set_facecolor('white')
plt.gcf().savefig('Abs_v_Conc_inhb_20.png')
plt.show()
# +
# Scatter plot of Absorbance vs. Concentration
from scipy.optimize import least_squares
# Compute molar concentratio of substrate
# conc. of substrate stock sol'n * mL subst. stock * 0.001 L/mL / sample_vol
S = 0.025 * (0.001 * inhb_data.index) / sample_vol # (mol/L)
plt.close()
plt.figure(figsize=(8.0,5.0))
plt.plot(S, inhb_data[10] - inhb_data[0], 'kx')
fit_inhb = least_squares(absrb_fit_resid,
[1.0, 1.0, 1.0],
args=(S, inhb_data[10] - inhb_data[0]),
method='lm',
verbose=2)
ss = np.linspace(0,S.max()*1.1,50)
fit_inhb_data = fit_inhb.x[2] * michaelis_menten_rxn_vel(fit_inhb.x[0], fit_inhb.x[1], ss)
plt.plot(ss, fit_inhb_data, 'k:')
plt.title('Absorbance vs. Concentration (at 10 min)')
plt.xlabel('Substrate Concentration [S] (mol/L)')
plt.ylabel('Absorbance (a.u.)')
plt.xlim([ss[0],ss[-1]])
ybnds = plt.gca().get_ylim()
plt.ylim([0,ybnds[1]])
plt.legend([
'Measured data',
'Scaled Michaelis-Menten\nderived least squares fit_inhb'
], loc='lower right')
plt.gcf().set_facecolor('white')
plt.gcf().savefig('Abs_v_Conc_inhb_10.png')
plt.show()
# +
# Scatter plot of Absorbance vs. Concentration
from scipy.optimize import least_squares
# Compute molar concentratio of substrate
# conc. of substrate stock sol'n * mL subst. stock * 0.001 L/mL / sample_vol
S = 0.025 * (0.001 * orig_data.index) / sample_vol # (mol/L)
plt.close()
plt.figure(figsize=(10.0,6.0))
plt.subplot(122)
plt.plot(S, orig_data[20] - orig_data[0], 'kx')
fit_orig = least_squares(absrb_fit_resid,
[1.0, 1.0, 1.0],
args=(S, orig_data[20] - orig_data[0]),
method='lm',
verbose=2)
ss = np.linspace(0,S.max()*1.1,50)
fit_orig_data = fit_orig.x[2] * michaelis_menten_rxn_vel(fit_orig.x[0], fit_orig.x[1], ss)
plt.plot(ss, fit_orig_data, 'k:')
plt.title('Absorbance vs. Concentration\n(at 20 min)')
plt.xlabel('Substrate Concentration\n[S] (mol/L)')
plt.ylabel('Absorbance (a.u.)')
plt.xlim([ss[0],ss[-1]])
ybnds = plt.gca().get_ylim()
plt.ylim([0,ybnds[1]])
plt.legend([
'Measured data',
'Scaled Michaelis-Menten\nderived least squares fit'
], loc='lower right')
plt.subplot(121)
plt.plot(S, orig_data[10] - orig_data[0], 'kx')
fit_orig = least_squares(absrb_fit_resid,
[1.0, 1.0, 1.0],
args=(S, orig_data[10] - orig_data[0]),
method='lm',
verbose=2)
ss = np.linspace(0,S.max()*1.1,50)
fit_orig_data = fit_orig.x[2] * michaelis_menten_rxn_vel(fit_orig.x[0], fit_orig.x[1], ss)
plt.plot(ss, fit_orig_data, 'k:')
plt.title('Absorbance vs. Concentration\n(at 10 min)')
plt.xlabel('Substrate Concentration\n[S] (mol/L)')
plt.ylabel('Absorbance (a.u.)')
plt.xlim([ss[0],ss[-1]])
ybnds = plt.gca().get_ylim()
plt.ylim([0,ybnds[1]])
plt.legend([
'Measured data',
'Scaled Michaelis-Menten\nderived least squares fit'
], loc='lower right')
plt.gcf().set_facecolor('white')
plt.gcf().savefig('Abs_v_Conc_orig.png')
plt.show()
# +
# Scatter plot of Absorbance vs. Concentration
from scipy.optimize import least_squares
# Compute molar concentratio of substrate
# conc. of substrate stock sol'n * mL subst. stock * 0.001 L/mL / sample_vol
S = 0.025 * (0.001 * inhb_data.index) / sample_vol # (mol/L)
plt.close()
plt.figure(figsize=(10.0,6.0))
plt.subplot(122)
plt.plot(S, inhb_data[20] - inhb_data[0], 'kx')
fit_inhb = least_squares(absrb_fit_resid,
[1.0, 1.0, 1.0],
args=(S, inhb_data[20] - inhb_data[0]),
method='lm',
verbose=2)
ss = np.linspace(0,S.max()*1.1,50)
fit_inhb_data = fit_inhb.x[2] * michaelis_menten_rxn_vel(fit_inhb.x[0], fit_inhb.x[1], ss)
plt.plot(ss, fit_inhb_data, 'k:')
plt.title('Absorbance vs. Concentration\n(at 20 min)')
plt.xlabel('Substrate Concentration\n[S] (mol/L)')
plt.ylabel('Absorbance (a.u.)')
plt.xlim([ss[0],ss[-1]])
ybnds = plt.gca().get_ylim()
plt.ylim([0,ybnds[1]])
plt.legend([
'Measured data',
'Scaled Michaelis-Menten\nderived least squares fit'
], loc='lower right')
plt.subplot(121)
plt.plot(S, inhb_data[10] - inhb_data[0], 'kx')
fit_inhb = least_squares(absrb_fit_resid,
[1.0, 1.0, 1.0],
args=(S, inhb_data[10] - inhb_data[0]),
method='lm',
verbose=2)
ss = np.linspace(0,S.max()*1.1,50)
fit_inhb_data = fit_inhb.x[2] * michaelis_menten_rxn_vel(fit_inhb.x[0], fit_inhb.x[1], ss)
plt.plot(ss, fit_inhb_data, 'k:')
plt.title('Absorbance vs. Concentration\n(at 10 min)')
plt.xlabel('Substrate Concentration\n[S] (mol/L)')
plt.ylabel('Absorbance (a.u.)')
plt.xlim([ss[0],ss[-1]])
ybnds = plt.gca().get_ylim()
plt.ylim([0,ybnds[1]])
plt.legend([
'Measured data',
'Scaled Michaelis-Menten\nderived least squares fit'
], loc='lower right')
plt.gcf().set_facecolor('white')
plt.gcf().savefig('Abs_v_Conc_inhb.png')
plt.show()
# +
# Comparison figure
plt.close()
plt.figure(figsize=(10.0, 6.0))
plt.subplot(122)
plt.plot(S, orig_data[20] - orig_data[0], 'ks')
fit_orig = least_squares(absrb_fit_resid,
[1.0, 1.0, 1.0],
args=(S, orig_data[20] - orig_data[0]),
method='lm',
verbose=2)
ss = np.linspace(0,S.max()*1.1,50)
fit_orig_data = fit_orig.x[2] * michaelis_menten_rxn_vel(fit_orig.x[0], fit_orig.x[1], ss)
plt.plot(ss, fit_orig_data, 'k--')
plt.plot(S, inhb_data[20] - inhb_data[0], 'kd')
fit_inhb = least_squares(absrb_fit_resid,
[1.0, 1.0, 1.0],
args=(S, inhb_data[20] - inhb_data[0]),
method='lm',
verbose=2)
ss = np.linspace(0,S.max()*1.1,50)
fit_inhb_data = fit_inhb.x[2] * michaelis_menten_rxn_vel(fit_inhb.x[0], fit_inhb.x[1], ss)
plt.plot(ss, fit_inhb_data, 'k-.')
plt.title('Absorbance vs. Concentration\n(at 20 min)')
plt.xlabel('Substrate Concentration\n[S] (mol/L)')
plt.ylabel('Absorbance (a.u.)')
plt.xlim([ss[0],ss[-1]])
ybnds = plt.gca().get_ylim()
plt.ylim([0,ybnds[1]])
plt.legend([
'Measured data\n(baseline condition)',
'Michaelis-Menten fit\n(baseline condition)',
'Measured data\n(inhibition condition)',
'Michaelis-Menten fit\n(inhibition condition)'
], loc='lower right')
plt.subplot(121)
plt.plot(S, orig_data[10] - orig_data[0], 'ks')
fit_orig = least_squares(absrb_fit_resid,
[1.0, 1.0, 1.0],
args=(S, orig_data[10] - orig_data[0]),
method='lm',
verbose=2)
ss = np.linspace(0,S.max()*1.1,50)
fit_orig_data = fit_orig.x[2] * michaelis_menten_rxn_vel(fit_orig.x[0], fit_orig.x[1], ss)
plt.plot(ss, fit_orig_data, 'k--')
plt.plot(S, inhb_data[10] - inhb_data[0], 'kd')
fit_inhb = least_squares(absrb_fit_resid,
[1.0, 1.0, 1.0],
args=(S, inhb_data[10] - inhb_data[0]),
method='lm',
verbose=2)
ss = np.linspace(0,S.max()*1.1,50)
fit_inhb_data = fit_inhb.x[2] * michaelis_menten_rxn_vel(fit_inhb.x[0], fit_inhb.x[1], ss)
plt.plot(ss, fit_inhb_data, 'k-.')
plt.title('Absorbance vs. Concentration\n(at 10 min)')
plt.xlabel('Substrate Concentration\n[S] (mol/L)')
plt.ylabel('Absorbance (a.u.)')
plt.xlim([ss[0],ss[-1]])
ybnds = plt.gca().get_ylim()
plt.ylim([0,ybnds[1]])
plt.legend([
'Measured data\n(baseline condition)',
'Michaelis-Menten fit\n(baseline condition)',
'Measured data\n(inhibition condition)',
'Michaelis-Menten fit\n(inhibition condition)'
], loc='lower right')
plt.gcf().set_facecolor('white')
plt.gcf().savefig('Abs_v_Conc_comp.png')
plt.show()
# +
# Lineweaver-Burke plot (scaled)
# NOTE: Absorbance stands in for velocity
plt.close()
plt.figure(figsize=(10.0, 6.0))
plt.subplot(122)
plt.plot(1/S, 1/(orig_data[20] - orig_data[0]), 'ks')
#Linear fit
fit = np.polyfit(1/S, 1/(orig_data[20] - orig_data[0]), 1)
fit_orig_data = np.poly1d(fit)
ss = np.linspace(0,(1/S).max()*1.1,50)
plt.plot(ss, fit_orig_data(ss), 'k--')
plt.plot(1/S, 1/(inhb_data[20] - inhb_data[0]), 'kd')
#Linear fit
fit = np.polyfit(1/S, 1/(inhb_data[20] - inhb_data[0]), 1)
fit_inhb_data = np.poly1d(fit)
ss = np.linspace(0,(1/S).max()*1.1,50)
plt.plot(ss, fit_inhb_data(ss), 'k-.')
plt.title('Approximated Lineweaver-Burke plot\n(at 20 min)')
plt.xlabel('1/[S] (L/mol)')
plt.ylabel('Absorbance$^{-1}$ (a.u.) ($\propto 1/V_0$)')
plt.xlim([ss[0],ss[-1]])
ybnds = plt.gca().get_ylim()
plt.ylim([0,ybnds[1]])
plt.legend([
'Measured data\n(baseline condition)',
'Lineweaver-Burke fit\n(baseline condition)',
'Measured data\n(inhibition condition)',
'Lineweaver-Burke fit\n(inhibition condition)'
], loc='upper left')
plt.subplot(121)
plt.plot(1/S, 1/(orig_data[10] - orig_data[0]), 'ks')
#Linear fit
fit = np.polyfit(1/S, 1/(orig_data[10] - orig_data[0]), 1)
fit_orig_data = np.poly1d(fit)
ss = np.linspace(0,(1/S).max()*1.1,50)
plt.plot(ss, fit_orig_data(ss), 'k--')
plt.plot(1/S, 1/(inhb_data[10] - inhb_data[0]), 'kd')
#Linear fit
fit = np.polyfit(1/S, 1/(inhb_data[10] - inhb_data[0]), 1)
fit_inhb_data = np.poly1d(fit)
ss = np.linspace(0,(1/S).max()*1.1,50)
plt.plot(ss, fit_inhb_data(ss), 'k-.')
plt.title('Approximated Lineweaver-Burke plot\n(at 10 min)')
plt.xlabel('1/[S] (L/mol)')
plt.ylabel('Absorbance$^{-1}$ (a.u.) ($\propto 1/V_0$)')
plt.xlim([ss[0],ss[-1]])
ybnds = plt.gca().get_ylim()
plt.ylim([0,ybnds[1]])
plt.legend([
'Measured data\n(baseline condition)',
'Lineweaver-Burke fit\n(baseline condition)',
'Measured data\n(inhibition condition)',
'Lineweaver-Burke fit\n(inhibition condition)'
], loc='upper left')
plt.gcf().set_facecolor('white')
plt.gcf().savefig('Approx_LWB_comp.png')
plt.show()
# +
# Lineweaver-Burke plot
# Calculate velocity from absorbance using last week's
# measured absorptivity
plt.close()
plt.figure(figsize=(10.0, 6.0))
plt.subplot(122)
# Calculate moles
prod_orig_molar_conc_20 = (orig_data[20] - orig_data[0]) / absorptivity
prod_orig_moles_20 = prod_orig_molar_conc_20 * sample_vol
V0_orig_20 = prod_orig_moles_20 / 20 # mol/min
plt.plot(1/S, 1/V0_orig_20, 'ks')
#Linear fit
fit_o20 = np.polyfit(1/S, 1/V0_orig_20, 1)
fit_orig_data = np.poly1d(fit_o20)
ss = np.linspace(0,(1/S).max()*1.1,50)
plt.plot(ss, fit_orig_data(ss), 'k--')
# Calculate moles
prod_inhb_molar_conc_20 = (inhb_data[20] - inhb_data[0]) / absorptivity
prod_inhb_moles_20 = prod_inhb_molar_conc_20 * sample_vol
V0_inhb_20 = prod_inhb_moles_20 / 20 # mol/min
plt.plot(1/S, 1/V0_inhb_20, 'kd')
#Linear fit
fit_i20 = np.polyfit(1/S, 1/V0_inhb_20, 1)
fit_inhb_data = np.poly1d(fit_i20)
ss = np.linspace(0,(1/S).max()*1.1,50)
plt.plot(ss, fit_inhb_data(ss), 'k-.')
plt.title('Lineweaver-Burke plot\n(at 20 min)')
plt.xlabel('$1/[S]$ (L/mol)')
plt.ylabel('$1/V_0$ (min/mol)')
plt.xlim([ss[0],ss[-1]])
ybnds = plt.gca().get_ylim()
plt.ylim([0,ybnds[1]])
plt.legend([
'Measured data\n(baseline condition)',
'Lineweaver-Burke fit\n(baseline condition)',
'Measured data\n(inhibition condition)',
'Lineweaver-Burke fit\n(inhibition condition)'
], loc='upper left')
plt.subplot(121)
# Calculate moles
prod_orig_molar_conc_10 = (orig_data[10] - orig_data[0]) / absorptivity
prod_orig_moles_10 = prod_orig_molar_conc_10 * sample_vol
V0_orig_10 = prod_orig_moles_10 / 20 # mol/min
plt.plot(1/S, 1/V0_orig_10, 'ks')
#Linear fit
fit_o10 = np.polyfit(1/S, 1/V0_orig_10, 1)
fit_orig_data = np.poly1d(fit_o10)
ss = np.linspace(0,(1/S).max()*1.1,50)
plt.plot(ss, fit_orig_data(ss), 'k--')
# Calculate moles
prod_inhb_molar_conc_10 = (inhb_data[10] - inhb_data[0]) / absorptivity
prod_inhb_moles_10 = prod_inhb_molar_conc_10 * sample_vol
V0_inhb_10 = prod_inhb_moles_10 / 20 # mol/min
plt.plot(1/S, 1/V0_inhb_10, 'kd')
#Linear fit
fit_i10 = np.polyfit(1/S, 1/V0_inhb_10, 1)
fit_inhb_data = np.poly1d(fit_i10)
ss = np.linspace(0,(1/S).max()*1.1,50)
plt.plot(ss, fit_inhb_data(ss), 'k-.')
plt.title('Lineweaver-Burke plot\n(at 10 min)')
plt.xlabel('$1/[S]$ (L/mol)')
plt.ylabel('$1/V_0$ (min/mol)')
plt.xlim([ss[0],ss[-1]])
ybnds = plt.gca().get_ylim()
plt.ylim([0,ybnds[1]])
plt.legend([
'Measured data\n(baseline condition)',
'Lineweaver-Burke fit\n(baseline condition)',
'Measured data\n(inhibition condition)',
'Lineweaver-Burke fit\n(inhibition condition)'
], loc='upper left')
plt.gcf().set_facecolor('white')
plt.gcf().savefig('LWB_comp.png')
plt.show()
# -
print('Baseline @ 10 min: Km = {:f} mol/L, Vmax = {:0.12f} mol/min'.format(
fit_o10[0] / fit_o10[1],
1 / fit_o10[1]
))
print('Baseline @ 20 min: Km = {:f} mol/L, Vmax = {:0.12f} mol/min'.format(
fit_o20[0] / fit_o20[1],
1 / fit_o20[1]
))
print('Inhibition @ 10 min: Km = {:f} mol/L, Vmax = {:0.12f} mol/min'.format(
fit_i10[0] / fit_i10[1],
1 / fit_i10[1]
))
print('Inhibition @ 20 min: Km = {:f} mol/L, Vmax = {:0.12f} mol/min'.format(
fit_i20[0] / fit_i20[1],
1 / fit_i20[1]
))
# molar mass of coloured product p-nitrophenol = 139.110 g/mol
print('Baseline @ 10 min: Km = {:f} mM, Vmax = {:0.12f} mg/(L*hr)'.format(
1e3 * fit_o10[0] / fit_o10[1],
(1 / fit_o10[1]) * 60 * 139.110 * 1e3 / sample_vol
))
print('Baseline @ 20 min: Km = {:f} mM, Vmax = {:0.12f} mg/(L*hr)'.format(
1e3 * fit_o20[0] / fit_o20[1],
(1 / fit_o20[1]) * 60 * 139.110 * 1e3 / sample_vol
))
print('Inhibition @ 10 min: Km = {:f} mM, Vmax = {:0.12f} mg/(L*hr)'.format(
1e3 * fit_i10[0] / fit_i10[1],
(1 / fit_i10[1]) * 60 * 139.110 * 1e3 / sample_vol
))
print('Inhibition @ 20 min: Km = {:f} mM, Vmax = {:0.12f} mg/(L*hr)'.format(
1e3 * fit_i20[0] / fit_i20[1],
(1 / fit_i20[1]) * 60 * 139.110 * 1e3 / sample_vol
))
| exp_4/Enzyme Kinetics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="xJzN8XWEaDv1" colab_type="code" colab={}
DATA_PATH = 'Project_2_Data.xlsx'
# + id="F2rYl-nCaDv4" colab_type="code" colab={}
import pandas as pd
# + id="FNwGJJGYaDv6" colab_type="code" colab={}
df = pd.read_excel(DATA_PATH)
# + id="jCfTBsjdaDv_" colab_type="code" colab={}
df.head(20)
# + id="dsyE8mSuaDwE" colab_type="code" colab={}
# choose your target. which column in will you predict?
# target: 2019 points
# + id="EIeAemRLaDwI" colab_type="code" colab={}
# regression or classification?
# regression
# + id="z10ABpxcaDwK" colab_type="code" colab={}
# evaluation metrics: MAE
# + id="3zxEAMidaDwM" colab_type="code" colab={}
train = df[df['Season'] < 2018]
val = df[df['Season'] == 2018]
test = df[df['Season'] == 2019]
# + id="2eXmSdZ2aDwO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="ea1a1e68-d227-486a-ba6a-45dd8da25275"
train.shape, val.shape, test.shape
# + id="-eg8tvW9Iq5w" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="ac7409a9-c03a-475a-a43d-c638b0183a8a"
df['Points'].value_counts(1)
# + id="rFdnXjzAmdv1" colab_type="code" colab={}
# assign variables
target = 'Points'
X_train = train.drop(columns=target)
y_train = train[target]
X_val = val.drop(columns=target)
y_val = val[target]
X_test = test
# + id="qESwRMeaQEED" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="9801fee4-8e85-47e5-8551-cd2c26b55966"
y_train.value_counts(normalize=True)
# + id="zaNzKABxQWXj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="01986b33-b521-463e-a8e3-d6f8c1fee068"
y_train.nunique()
# + id="dIX6Wye03DvO" colab_type="code" colab={}
# Split train into train & val
from sklearn.model_selection import train_test_split
train, val = train_test_split(train, train_size=0.80, test_size=0.20,
stratify= y_train, random_state=42)
# + id="C_k7_nChDe6q" colab_type="code" colab={}
X_train, X_val , y_train, y_val = train_test_split(X_train, y_train, test_size=0.2, stratify=y_train, random_state=42)
# + id="rN7864i9aDwR" colab_type="code" colab={}
# # null?
df.isna().sum()
# + id="UeDjr902aDwZ" colab_type="code" colab={}
df.nunique()
# + id="raHMvhJwaDw4" colab_type="code" colab={}
import pandas_profiling
# + id="z6Ibiw7GaDw8" colab_type="code" colab={}
#profile_report()
# + id="0cQHMAOCaY-p" colab_type="code" colab={}
import numpy as np
import pandas as pd
from sklearn.ensemble import RandomForestClassifier
import category_encoders as ce
from sklearn.impute import SimpleImputer
from sklearn.pipeline import make_pipeline
# + id="18pPFElYohDn" colab_type="code" colab={}
pipeline = make_pipeline(
ce.OrdinalEncoder(),
SimpleImputer(strategy='median'),
RandomForestClassifier(n_estimators=100, random_state=42, n_jobs=-1)
)
# Fit on train, score on val
pipeline.fit(X_train, y_train)
print('Validation Accuracy', pipeline.score(X_val, y_val))
# + id="cm45tSv0qL-k" colab_type="code" colab={}
# Get feature importances
rf = pipeline.named_steps['randomforestclassifier']
importances = pd.Series(rf.feature_importances_, X_train.columns)
# Plot feature importances
# %matplotlib inline
import matplotlib.pyplot as plt
n = 8
plt.figure(figsize=(10,n/2))
plt.title(f'Top {n} features')
importances.sort_values()[-n:].plot.barh(color='grey');
# + id="mLzNPrlttib5" colab_type="code" colab={}
column = 'Expenditure per Point'
# Fit without column
pipeline = make_pipeline(
ce.OrdinalEncoder(),
SimpleImputer(strategy='median'),
RandomForestClassifier(n_estimators=100, random_state=42, n_jobs=-1)
)
pipeline.fit(X_train.drop(columns=column), y_train)
score_without = pipeline.score(X_val.drop(columns=column), y_val)
print(f'Validation Accuracy without {column}: {score_without}')
# Fit with column
pipeline = make_pipeline(
ce.OrdinalEncoder(),
SimpleImputer(strategy='median'),
RandomForestClassifier(n_estimators=100, random_state=42, n_jobs=-1)
)
pipeline.fit(X_train, y_train)
score_with = pipeline.score(X_val, y_val)
print(f'Validation Accuracy with {column}: {score_with}')
# Compare the error with & without column
print(f'Drop-Column Importance for {column}: {score_with - score_without}')
# + id="cjezmanxtCMB" colab_type="code" colab={}
feature = 'Expenditure per Point'
X_val[feature].value_counts()
# + id="aXM2nMUgsoXI" colab_type="code" colab={}
X_val_permuted = X_val.copy()
X_val_permuted[feature] = np.random.permutation(X_val[feature])
# + id="v6hf0P3LtRWg" colab_type="code" colab={}
X_val_permuted[feature].head()
# + id="8dCBEVmNuHIQ" colab_type="code" colab={}
# Get the permutation importance
score_permuted = pipeline.score(X_val_permuted, y_val)
print(f'Validation accuracy with {feature}: {score_with}')
print(f'Validation accuracy with {feature} permuted: {score_permuted}')
print(f'Permutation importance: {score_with - score_permuted}')
# + id="wmQgnxVIuRAa" colab_type="code" colab={}
# Rerun the permutation importance process, but for a different feature
feature = 'Club'
X_val_permuted = X_val.copy()
X_val_permuted[feature] = np.random.permutation(X_val[feature])
score_permuted = pipeline.score(X_val_permuted, y_val)
print(f'Validation accuracy with {feature}: {score_with}')
print(f'Validation accuracy with {feature} permuted: {score_permuted}')
print(f'Permutation importance: {score_with - score_permuted}')
# + id="5PIRyC8GbszT" colab_type="code" colab={}
# eli5
# + id="jA2ZxGz2zpFv" colab_type="code" colab={}
# assign variables
target = 'Points'
X_train = train.drop(columns=target)
y_train = train[target]
X_val = val.drop(columns=target)
y_val = val[target]
X_test = test
# + id="Ufq9rXieaDw_" colab_type="code" colab={}
import eli5
from eli5.sklearn import PermutationImportance
# + id="LfTxzHzUuaLZ" colab_type="code" colab={}
transformers = make_pipeline(
ce.OrdinalEncoder(),
SimpleImputer(strategy='mean')
)
# + id="d2ZJYLeKu2Qz" colab_type="code" colab={}
X_train_transformed = transformers.fit_transform(X_train)
X_val_transformed = transformers.transform(X_val)
model = RandomForestClassifier(n_estimators=100, random_state=42, n_jobs=-1)
model.fit(X_train_transformed, y_train)
# + id="tmyxsKvku2Bv" colab_type="code" colab={}
# 1. Calculate permutation importances
permuter = PermutationImportance(
model,
scoring='neg_mean_absolute_error',
n_iter=5,
random_state=42
)
permuter.fit(X_val_transformed, y_val)
# + id="gCgp2y7Eu1zT" colab_type="code" colab={}
feature_names = X_val.columns.tolist()
pd.Series(permuter.feature_importances_, feature_names).sort_values()
# + id="5eAjisgmvHM4" colab_type="code" colab={}
# 2. Display permutation importances
eli5.show_weights(
permuter,
top=None, # show permutation importances for all features
feature_names=feature_names # must be a list
)
# + id="fiObqiGovMI9" colab_type="code" colab={}
# Use XGBoost for Gradient Boosting
# + id="nhtcUH72v3mf" colab_type="code" colab={}
from xgboost import XGBClassifier
pipeline = make_pipeline(
ce.OrdinalEncoder(),
XGBClassifier(n_estimators=100, random_state=42, n_jobs=-1)
)
pipeline.fit(X_train, y_train)
# + id="N2IQxY_7v3cV" colab_type="code" colab={}
from sklearn.metrics import accuracy_score
y_pred = pipeline.predict(X_val)
print('Validation Accuracy', accuracy_score(y_val, y_pred))
# + id="UW1Za2Fuv3Tv" colab_type="code" colab={}
# fit_transfom on train, transform on val
encoder = ce.OrdinalEncoder()
X_train_encoded = encoder.fit_transform(X_train)
X_val_encoded = encoder.transform(X_val)
model = XGBClassifier(
n_estimators=1000,
max_depth=7,
learning_rate=0.5,
n_jobs=-1
)
eval_set = [(X_train_encoded, y_train),
(X_val_encoded, y_val)]
model.fit(X_train_encoded, y_train,
eval_set=eval_set,
eval_metric='merror',
early_stopping_rounds=50)
| module3/Project_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Ответ на вопрос [об инвертировании матрицы методом Гаусса](https://ru.stackoverflow.com/questions/1279499)
import numpy as np
def invert(A):
assert np.ndim(A) == 2
assert A.shape[0] == A.shape[1]
n = A.shape[0]
matrix = np.hstack((A, np.eye(n)))
# Приводим матрицу А к диагональному виду
for nrow, row in enumerate(matrix):
# nrow равен номеру строки
# row содержит саму строку матрицы
divider = row[nrow] # диагональный элемент
# делим на диагональный элемент.
row /= divider
# теперь надо вычесть приведённую строку из всех нижележащих строчек
for lower_row in matrix[nrow+1:]:
factor = lower_row[nrow] # элемент строки в колонке nrow
lower_row -= factor*row # вычитаем, чтобы получить ноль в колонке nrow
# обратный ход
for nrow in range(len(matrix)-1,0,-1):
row = matrix[nrow]
for upper_row in matrix[:nrow]:
factor = upper_row[nrow]
# Вычитание целой строки на 15% быстрее, чем вычитание только правой части
upper_row -= factor*row
return matrix[:, n:].copy()
A = np.array([[3.8, 6.7, -1.2],
[6.4, 1.3, -2.7],
[2.4, -4.5, 3.5]])
I = invert(A)
I
# Проверка - произведение обратной матрицы и исходной матрицы
np.matmul(A, I)
# Максимальное отклонение от нуля в произведении обратной и исходной матриц
np.max(np.abs(np.matmul(A, I) - np.eye(3)))
# Исправленный вариант топик-стартера
# +
def inverse_matrix(matrix_origin):
"""
Функция получает на вход матрицу, затем добавляет к ней единичную матрицу,
проводит элементарные преобразования по строкам с первоначальной, добиваясь получения слева единичной матрицы.
В этом случае справа окажется матрица, которая является обратной к заданнй первоначально
"""
# Склеиваем 2 матрицы: слева - первоначальная, справа - единичная
n = matrix_origin.shape[0]
m = np.hstack((matrix_origin, np.eye(n)))
for nrow, row in enumerate(m):
# nrow равен номеру строки
# row содержит саму строку матрицы
divider = row[nrow] # диагональный элемент
# делим на диагональный элемент:
row /= divider
# теперь вычитаем приведённую строку из всех нижележащих строк:
for lower_row in m[nrow+1:]:
factor = lower_row[nrow] # элемент строки в колонке nrow
lower_row -= factor*row # вычитаем, чтобы получить ноль в колонке nrow
# обратный ход:
for k in range(n - 1, 0, -1):
for row_ in range(k - 1, -1, -1):
if m[row_, k]:
# 1) Все элементы выше главной диагонали делаем равными нулю
m[row_, :] -= m[k, :] * m[row_, k]
return m[:,n:].copy()
# +
matrix = np.array([[3.8, 6.7, -1.2],
[6.4, 1.3, -2.7],
[2.4, -4.5, 3.5]])
inverse_matrix(np.copy(matrix))
# -
B = np.random.rand(100,100)
# %timeit -n 10 -r 3 invert(B)
B = np.random.rand(100,100)
# %timeit -n 10 -r 3 inverse_matrix(B)
| python/1271321/invert_gauss.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="JmvzTcYice-_"
# ##### Copyright 2020 The TensorFlow Authors.
# + id="zlvAS8a9cD_t"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="b2VYQpTttmVN"
# # TensorFlow Constrained Optimization Example Using CelebA Dataset
# + [markdown] id="3iFsS2WSeRwe"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/responsible_ai/fairness_indicators/tutorials/Fairness_Indicators_TFCO_CelebA_Case_Study"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/fairness-indicators/blob/master/g3doc/tutorials/Fairness_Indicators_TFCO_CelebA_Case_Study.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/fairness-indicators/tree/master/g3doc/tutorials/Fairness_Indicators_TFCO_CelebA_Case_Study.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/fairness-indicators/g3doc/tutorials/Fairness_Indicators_TFCO_CelebA_Case_Study.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# </table>
# + [markdown] id="-DQoReGDeN16"
# This notebook demonstrates an easy way to create and optimize constrained problems using the TFCO library. This method can be useful in improving models when we find that they’re not performing equally well across different slices of our data, which we can identify using [Fairness Indicators](https://www.tensorflow.org/responsible_ai/fairness_indicators/guide). The second of Google’s AI principles states that our technology should avoid creating or reinforcing unfair bias, and we believe this technique can help improve model fairness in some situations. In particular, this notebook will:
#
#
# * Train a simple, *unconstrained* neural network model to detect a person's smile in images using [`tf.keras`](https://www.tensorflow.org/guide/keras) and the large-scale CelebFaces Attributes ([CelebA](http://mmlab.ie.cuhk.edu.hk/projects/CelebA.html)) dataset.
# * Evaluate model performance against a commonly used fairness metric across age groups, using Fairness Indicators.
# * Set up a simple constrained optimization problem to achieve fairer performance across age groups.
# * Retrain the now *constrained* model and evaluate performance again, ensuring that our chosen fairness metric has improved.
#
# Last updated: 3/11 Feb 2020
# + [markdown] id="JyCbEWt5Zxe2"
# # Installation
# This notebook was created in [Colaboratory](https://research.google.com/colaboratory/faq.html), connected to the Python 3 Google Compute Engine backend. If you wish to host this notebook in a different environment, then you should not experience any major issues provided you include all the required packages in the cells below.
#
# Note that the very first time you run the pip installs, you may be asked to restart the runtime because of preinstalled out of date packages. Once you do so, the correct packages will be used.
# + id="T-Zm-KDdt0bn"
#@title Pip installs
# !pip install -q -U pip==20.2
# !pip install git+https://github.com/google-research/tensorflow_constrained_optimization
# !pip install -q tensorflow-datasets tensorflow
# !pip install fairness-indicators \
# "absl-py==0.12.0" \
# "apache-beam<3,>=2.38" \
# "avro-python3==1.9.1" \
# "pyzmq==17.0.0"
# + [markdown] id="UXWXhBLvISOY"
# Note that depending on when you run the cell below, you may receive a warning about the default version of TensorFlow in Colab switching to TensorFlow 2.X soon. You can safely ignore that warning as this notebook was designed to be compatible with TensorFlow 1.X and 2.X.
# + id="UTBBdSGaZ8aW"
#@title Import Modules
import os
import sys
import tempfile
import urllib
import tensorflow as tf
from tensorflow import keras
import tensorflow_datasets as tfds
tfds.disable_progress_bar()
import numpy as np
import tensorflow_constrained_optimization as tfco
from tensorflow_metadata.proto.v0 import schema_pb2
from tfx_bsl.tfxio import tensor_adapter
from tfx_bsl.tfxio import tf_example_record
# + [markdown] id="70tLum8uIZUm"
# Additionally, we add a few imports that are specific to Fairness Indicators which we will use to evaluate and visualize the model's performance.
# + cellView="form" id="7Se0Z0Bo9K-5"
#@title Fairness Indicators related imports
import tensorflow_model_analysis as tfma
import fairness_indicators as fi
from google.protobuf import text_format
import apache_beam as beam
# + [markdown] id="xSG2HP7goGrj"
# Although TFCO is compatible with eager and graph execution, this notebook assumes that eager execution is enabled by default as it is in TensorFlow 2.x. To ensure that nothing breaks, eager execution will be enabled in the cell below.
# + id="W0ZusW1-lBao"
#@title Enable Eager Execution and Print Versions
if tf.__version__ < "2.0.0":
tf.compat.v1.enable_eager_execution()
print("Eager execution enabled.")
else:
print("Eager execution enabled by default.")
print("TensorFlow " + tf.__version__)
print("TFMA " + tfma.VERSION_STRING)
print("TFDS " + tfds.version.__version__)
print("FI " + fi.version.__version__)
# + [markdown] id="idY3Uuk3yvty"
# # CelebA Dataset
# [CelebA](http://mmlab.ie.cuhk.edu.hk/projects/CelebA.html) is a large-scale face attributes dataset with more than 200,000 celebrity images, each with 40 attribute annotations (such as hair type, fashion accessories, facial features, etc.) and 5 landmark locations (eyes, mouth and nose positions). For more details take a look at [the paper](https://liuziwei7.github.io/projects/FaceAttributes.html).
# With the permission of the owners, we have stored this dataset on Google Cloud Storage and mostly access it via [TensorFlow Datasets(`tfds`)](https://www.tensorflow.org/datasets).
#
# In this notebook:
# * Our model will attempt to classify whether the subject of the image is smiling, as represented by the "Smiling" attribute<sup>*</sup>.
# * Images will be resized from 218x178 to 28x28 to reduce the execution time and memory when training.
# * Our model's performance will be evaluated across age groups, using the binary "Young" attribute. We will call this "age group" in this notebook.
#
# ___
#
# <sup>*</sup> While there is little information available about the labeling methodology for this dataset, we will assume that the "Smiling" attribute was determined by a pleased, kind, or amused expression on the subject's face. For the purpose of this case study, we will take these labels as ground truth.
#
#
#
# + id="zCSemFST0b89"
gcs_base_dir = "gs://celeb_a_dataset/"
celeb_a_builder = tfds.builder("celeb_a", data_dir=gcs_base_dir, version='2.0.0')
celeb_a_builder.download_and_prepare()
num_test_shards_dict = {'0.3.0': 4, '2.0.0': 2} # Used because we download the test dataset separately
version = str(celeb_a_builder.info.version)
print('Celeb_A dataset version: %s' % version)
# + cellView="form" id="Ocqv3R06APfW"
#@title Test dataset helper functions
local_root = tempfile.mkdtemp(prefix='test-data')
def local_test_filename_base():
return local_root
def local_test_file_full_prefix():
return os.path.join(local_test_filename_base(), "celeb_a-test.tfrecord")
def copy_test_files_to_local():
filename_base = local_test_file_full_prefix()
num_test_shards = num_test_shards_dict[version]
for shard in range(num_test_shards):
url = "https://storage.googleapis.com/celeb_a_dataset/celeb_a/%s/celeb_a-test.tfrecord-0000%s-of-0000%s" % (version, shard, num_test_shards)
filename = "%s-0000%s-of-0000%s" % (filename_base, shard, num_test_shards)
res = urllib.request.urlretrieve(url, filename)
# + [markdown] id="u5PDLXZb_uIj"
# ## Caveats
# Before moving forward, there are several considerations to keep in mind in using CelebA:
# * Although in principle this notebook could use any dataset of face images, CelebA was chosen because it contains public domain images of public figures.
# * All of the attribute annotations in CelebA are operationalized as binary categories. For example, the "Young" attribute (as determined by the dataset labelers) is denoted as either present or absent in the image.
# * CelebA's categorizations do not reflect real human diversity of attributes.
# * For the purposes of this notebook, the feature containing the "Young" attribute is referred to as "age group", where the presence of the "Young" attribute in an image is labeled as a member of the "Young" age group and the absence of the "Young" attribute is labeled as a member of the "Not Young" age group. These are assumptions made as this information is not mentioned in the [original paper](http://openaccess.thecvf.com/content_iccv_2015/html/Liu_Deep_Learning_Face_ICCV_2015_paper.html).
# * As such, performance in the models trained in this notebook is tied to the ways the attributes have been operationalized and annotated by the authors of CelebA.
# * This model should not be used for commercial purposes as that would violate [CelebA's non-commercial research agreement](http://mmlab.ie.cuhk.edu.hk/projects/CelebA.html).
# + [markdown] id="Elkiu92cY2bY"
# # Setting Up Input Functions
# The subsequent cells will help streamline the input pipeline as well as visualize performance.
#
# First we define some data-related variables and define a requisite preprocessing function.
# + id="gDdarTZxk6y4"
#@title Define Variables
ATTR_KEY = "attributes"
IMAGE_KEY = "image"
LABEL_KEY = "Smiling"
GROUP_KEY = "Young"
IMAGE_SIZE = 28
# + cellView="form" id="SD-H70Je0cTp"
#@title Define Preprocessing Functions
def preprocess_input_dict(feat_dict):
# Separate out the image and target variable from the feature dictionary.
image = feat_dict[IMAGE_KEY]
label = feat_dict[ATTR_KEY][LABEL_KEY]
group = feat_dict[ATTR_KEY][GROUP_KEY]
# Resize and normalize image.
image = tf.cast(image, tf.float32)
image = tf.image.resize(image, [IMAGE_SIZE, IMAGE_SIZE])
image /= 255.0
# Cast label and group to float32.
label = tf.cast(label, tf.float32)
group = tf.cast(group, tf.float32)
feat_dict[IMAGE_KEY] = image
feat_dict[ATTR_KEY][LABEL_KEY] = label
feat_dict[ATTR_KEY][GROUP_KEY] = group
return feat_dict
get_image_and_label = lambda feat_dict: (feat_dict[IMAGE_KEY], feat_dict[ATTR_KEY][LABEL_KEY])
get_image_label_and_group = lambda feat_dict: (feat_dict[IMAGE_KEY], feat_dict[ATTR_KEY][LABEL_KEY], feat_dict[ATTR_KEY][GROUP_KEY])
# + [markdown] id="iwg3sPmExciD"
# Then, we build out the data functions we need in the rest of the colab.
# + id="KbR64r0VVG5h"
# Train data returning either 2 or 3 elements (the third element being the group)
def celeb_a_train_data_wo_group(batch_size):
celeb_a_train_data = celeb_a_builder.as_dataset(split='train').shuffle(1024).repeat().batch(batch_size).map(preprocess_input_dict)
return celeb_a_train_data.map(get_image_and_label)
def celeb_a_train_data_w_group(batch_size):
celeb_a_train_data = celeb_a_builder.as_dataset(split='train').shuffle(1024).repeat().batch(batch_size).map(preprocess_input_dict)
return celeb_a_train_data.map(get_image_label_and_group)
# Test data for the overall evaluation
celeb_a_test_data = celeb_a_builder.as_dataset(split='test').batch(1).map(preprocess_input_dict).map(get_image_label_and_group)
# Copy test data locally to be able to read it into tfma
copy_test_files_to_local()
# + [markdown] id="NXO3woTxiCk0"
# # Build a simple DNN Model
# Because this notebook focuses on TFCO, we will assemble a simple, unconstrained `tf.keras.Sequential` model.
#
# We may be able to greatly improve model performance by adding some complexity (e.g., more densely-connected layers, exploring different activation functions, increasing image size), but that may distract from the goal of demonstrating how easy it is to apply the TFCO library when working with Keras. For that reason, the model will be kept simple — but feel encouraged to explore this space.
# + id="RNZhN_zU8DRD"
def create_model():
# For this notebook, accuracy will be used to evaluate performance.
METRICS = [
tf.keras.metrics.BinaryAccuracy(name='accuracy')
]
# The model consists of:
# 1. An input layer that represents the 28x28x3 image flatten.
# 2. A fully connected layer with 64 units activated by a ReLU function.
# 3. A single-unit readout layer to output real-scores instead of probabilities.
model = keras.Sequential([
keras.layers.Flatten(input_shape=(IMAGE_SIZE, IMAGE_SIZE, 3), name='image'),
keras.layers.Dense(64, activation='relu'),
keras.layers.Dense(1, activation=None)
])
# TFCO by default uses hinge loss — and that will also be used in the model.
model.compile(
optimizer=tf.keras.optimizers.Adam(0.001),
loss='hinge',
metrics=METRICS)
return model
# + [markdown] id="7A4uKPNVzPVO"
# We also define a function to set seeds to ensure reproducible results. Note that this colab is meant as an educational tool and does not have the stability of a finely tuned production pipeline. Running without setting a seed may lead to varied results.
# + id="-IVw4EgKzqSF"
def set_seeds():
np.random.seed(121212)
tf.compat.v1.set_random_seed(212121)
# + [markdown] id="Xrbjmmeom8pA"
# # Fairness Indicators Helper Functions
# Before training our model, we define a number of helper functions that will allow us to evaluate the model's performance via Fairness Indicators.
#
# + [markdown] id="1EPF_k620CRN"
# First, we create a helper function to save our model once we train it.
# + id="ejHbhLW5epar"
def save_model(model, subdir):
base_dir = tempfile.mkdtemp(prefix='saved_models')
model_location = os.path.join(base_dir, subdir)
model.save(model_location, save_format='tf')
return model_location
# + [markdown] id="erhKEvqByCNj"
# Next, we define functions used to preprocess the data in order to correctly pass it through to TFMA.
# + id="D2qa8Okwj_U3"
#@title Data Preprocessing functions for
def tfds_filepattern_for_split(dataset_name, split):
return f"{local_test_file_full_prefix()}*"
class PreprocessCelebA(object):
"""Class that deserializes, decodes and applies additional preprocessing for CelebA input."""
def __init__(self, dataset_name):
builder = tfds.builder(dataset_name)
self.features = builder.info.features
example_specs = self.features.get_serialized_info()
self.parser = tfds.core.example_parser.ExampleParser(example_specs)
def __call__(self, serialized_example):
# Deserialize
deserialized_example = self.parser.parse_example(serialized_example)
# Decode
decoded_example = self.features.decode_example(deserialized_example)
# Additional preprocessing
image = decoded_example[IMAGE_KEY]
label = decoded_example[ATTR_KEY][LABEL_KEY]
# Resize and scale image.
image = tf.cast(image, tf.float32)
image = tf.image.resize(image, [IMAGE_SIZE, IMAGE_SIZE])
image /= 255.0
image = tf.reshape(image, [-1])
# Cast label and group to float32.
label = tf.cast(label, tf.float32)
group = decoded_example[ATTR_KEY][GROUP_KEY]
output = tf.train.Example()
output.features.feature[IMAGE_KEY].float_list.value.extend(image.numpy().tolist())
output.features.feature[LABEL_KEY].float_list.value.append(label.numpy())
output.features.feature[GROUP_KEY].bytes_list.value.append(b"Young" if group.numpy() else b'Not Young')
return output.SerializeToString()
def tfds_as_pcollection(beam_pipeline, dataset_name, split):
return (
beam_pipeline
| 'Read records' >> beam.io.ReadFromTFRecord(tfds_filepattern_for_split(dataset_name, split))
| 'Preprocess' >> beam.Map(PreprocessCelebA(dataset_name))
)
# + [markdown] id="fBKvxd2Tz3hK"
# Finally, we define a function that evaluates the results in TFMA.
# + id="30YduitftaNB"
def get_eval_results(model_location, eval_subdir):
base_dir = tempfile.mkdtemp(prefix='saved_eval_results')
tfma_eval_result_path = os.path.join(base_dir, eval_subdir)
eval_config_pbtxt = """
model_specs {
label_key: "%s"
}
metrics_specs {
metrics {
class_name: "FairnessIndicators"
config: '{ "thresholds": [0.22, 0.5, 0.75] }'
}
metrics {
class_name: "ExampleCount"
}
}
slicing_specs {}
slicing_specs { feature_keys: "%s" }
options {
compute_confidence_intervals { value: False }
disabled_outputs{values: "analysis"}
}
""" % (LABEL_KEY, GROUP_KEY)
eval_config = text_format.Parse(eval_config_pbtxt, tfma.EvalConfig())
eval_shared_model = tfma.default_eval_shared_model(
eval_saved_model_path=model_location, tags=[tf.saved_model.SERVING])
schema_pbtxt = """
tensor_representation_group {
key: ""
value {
tensor_representation {
key: "%s"
value {
dense_tensor {
column_name: "%s"
shape {
dim { size: 28 }
dim { size: 28 }
dim { size: 3 }
}
}
}
}
}
}
feature {
name: "%s"
type: FLOAT
}
feature {
name: "%s"
type: FLOAT
}
feature {
name: "%s"
type: BYTES
}
""" % (IMAGE_KEY, IMAGE_KEY, IMAGE_KEY, LABEL_KEY, GROUP_KEY)
schema = text_format.Parse(schema_pbtxt, schema_pb2.Schema())
coder = tf_example_record.TFExampleBeamRecord(
physical_format='inmem', schema=schema,
raw_record_column_name=tfma.ARROW_INPUT_COLUMN)
tensor_adapter_config = tensor_adapter.TensorAdapterConfig(
arrow_schema=coder.ArrowSchema(),
tensor_representations=coder.TensorRepresentations())
# Run the fairness evaluation.
with beam.Pipeline() as pipeline:
_ = (
tfds_as_pcollection(pipeline, 'celeb_a', 'test')
| 'ExamplesToRecordBatch' >> coder.BeamSource()
| 'ExtractEvaluateAndWriteResults' >>
tfma.ExtractEvaluateAndWriteResults(
eval_config=eval_config,
eval_shared_model=eval_shared_model,
output_path=tfma_eval_result_path,
tensor_adapter_config=tensor_adapter_config)
)
return tfma.load_eval_result(output_path=tfma_eval_result_path)
# + [markdown] id="76tZ3vk-tyo9"
# # Train & Evaluate Unconstrained Model
#
# With the model now defined and the input pipeline in place, we’re now ready to train our model. To cut back on the amount of execution time and memory, we will train the model by slicing the data into small batches with only a few repeated iterations.
#
# Note that running this notebook in TensorFlow < 2.0.0 may result in a deprecation warning for `np.where`. Safely ignore this warning as TensorFlow addresses this in 2.X by using `tf.where` in place of `np.where`.
# + id="3m9OOdU_8GWo"
BATCH_SIZE = 32
# Set seeds to get reproducible results
set_seeds()
model_unconstrained = create_model()
model_unconstrained.fit(celeb_a_train_data_wo_group(BATCH_SIZE), epochs=5, steps_per_epoch=1000)
# + [markdown] id="nCtBH9DkvtUy"
# Evaluating the model on the test data should result in a final accuracy score of just over 85%. Not bad for a simple model with no fine tuning.
# + id="mgsjbxpTIdZf"
print('Overall Results, Unconstrained')
celeb_a_test_data = celeb_a_builder.as_dataset(split='test').batch(1).map(preprocess_input_dict).map(get_image_label_and_group)
results = model_unconstrained.evaluate(celeb_a_test_data)
# + [markdown] id="L5jslIrzwIKo"
# However, performance evaluated across age groups may reveal some shortcomings.
#
# To explore this further, we evaluate the model with Fairness Indicators (via TFMA). In particular, we are interested in seeing whether there is a significant gap in performance between "Young" and "Not Young" categories when evaluated on false positive rate.
#
# A false positive error occurs when the model incorrectly predicts the positive class. In this context, a false positive outcome occurs when the ground truth is an image of a celebrity 'Not Smiling' and the model predicts 'Smiling'. By extension, the false positive rate, which is used in the visualization above, is a measure of accuracy for a test. While this is a relatively mundane error to make in this context, false positive errors can sometimes cause more problematic behaviors. For instance, a false positive error in a spam classifier could cause a user to miss an important email.
# + id="nFL91nZF1V8D"
model_location = save_model(model_unconstrained, 'model_export_unconstrained')
eval_results_unconstrained = get_eval_results(model_location, 'eval_results_unconstrained')
# + [markdown] id="34zHIMW0NHld"
# As mentioned above, we are concentrating on the false positive rate. The current version of Fairness Indicators (0.1.2) selects false negative rate by default. After running the line below, deselect false_negative_rate and select false_positive_rate to look at the metric we are interested in.
# + id="KXMVmUMi0ydk"
tfma.addons.fairness.view.widget_view.render_fairness_indicator(eval_results_unconstrained)
# + [markdown] id="zYVpZ-DpBsfD"
# As the results show above, we do see a **disproportionate gap between "Young" and "Not Young" categories**.
#
# This is where TFCO can help by constraining the false positive rate to be within a more acceptable criterion.
#
# + [markdown] id="ZNnI_Eu70gVp"
# # Constrained Model Set Up
# As documented in [TFCO's library](https://github.com/google-research/tensorflow_constrained_optimization/blob/master/README.md), there are several helpers that will make it easier to constrain the problem:
#
# 1. `tfco.rate_context()` – This is what will be used in constructing a constraint for each age group category.
# 2. `tfco.RateMinimizationProblem()`– The rate expression to be minimized here will be the false positive rate subject to age group. In other words, performance now will be evaluated based on the difference between the false positive rates of the age group and that of the overall dataset. For this demonstration, a false positive rate of less than or equal to 5% will be set as the constraint.
# 3. `tfco.ProxyLagrangianOptimizerV2()` – This is the helper that will actually solve the rate constraint problem.
#
# The cell below will call on these helpers to set up model training with the fairness constraint.
#
#
#
# + id="BTukzvfD6iWr"
# The batch size is needed to create the input, labels and group tensors.
# These tensors are initialized with all 0's. They will eventually be assigned
# the batch content to them. A large batch size is chosen so that there are
# enough number of "Young" and "Not Young" examples in each batch.
set_seeds()
model_constrained = create_model()
BATCH_SIZE = 32
# Create input tensor.
input_tensor = tf.Variable(
np.zeros((BATCH_SIZE, IMAGE_SIZE, IMAGE_SIZE, 3), dtype="float32"),
name="input")
# Create labels and group tensors (assuming both labels and groups are binary).
labels_tensor = tf.Variable(
np.zeros(BATCH_SIZE, dtype="float32"), name="labels")
groups_tensor = tf.Variable(
np.zeros(BATCH_SIZE, dtype="float32"), name="groups")
# Create a function that returns the applied 'model' to the input tensor
# and generates constrained predictions.
def predictions():
return model_constrained(input_tensor)
# Create overall context and subsetted context.
# The subsetted context contains subset of examples where group attribute < 1
# (i.e. the subset of "Not Young" celebrity images).
# "groups_tensor < 1" is used instead of "groups_tensor == 0" as the former
# would be a comparison on the tensor value, while the latter would be a
# comparison on the Tensor object.
context = tfco.rate_context(predictions, labels=lambda:labels_tensor)
context_subset = context.subset(lambda:groups_tensor < 1)
# Setup list of constraints.
# In this notebook, the constraint will just be: FPR to less or equal to 5%.
constraints = [tfco.false_positive_rate(context_subset) <= 0.05]
# Setup rate minimization problem: minimize overall error rate s.t. constraints.
problem = tfco.RateMinimizationProblem(tfco.error_rate(context), constraints)
# Create constrained optimizer and obtain train_op.
# Separate optimizers are specified for the objective and constraints
optimizer = tfco.ProxyLagrangianOptimizerV2(
optimizer=tf.keras.optimizers.Adam(learning_rate=0.001),
constraint_optimizer=tf.keras.optimizers.Adam(learning_rate=0.001),
num_constraints=problem.num_constraints)
# A list of all trainable variables is also needed to use TFCO.
var_list = (model_constrained.trainable_weights + list(problem.trainable_variables) +
optimizer.trainable_variables())
# + [markdown] id="thEe8A8UYbrO"
# The model is now set up and ready to be trained with the false positive rate constraint across age group.
#
# Now, because the last iteration of the constrained model may not necessarily be the best performing model in terms of the defined constraint, the TFCO library comes equipped with `tfco.find_best_candidate_index()` that can help choose the best iterate out of the ones found after each epoch. Think of `tfco.find_best_candidate_index()` as an added heuristic that ranks each of the outcomes based on accuracy and fairness constraint (in this case, false positive rate across age group) separately with respect to the training data. That way, it can search for a better trade-off between overall accuracy and the fairness constraint.
#
# The following cells will start the training with constraints while also finding the best performing model per iteration.
# + id="73doG4HL6nPS"
# Obtain train set batches.
NUM_ITERATIONS = 100 # Number of training iterations.
SKIP_ITERATIONS = 10 # Print training stats once in this many iterations.
# Create temp directory for saving snapshots of models.
temp_directory = tempfile.mktemp()
os.mkdir(temp_directory)
# List of objective and constraints across iterations.
objective_list = []
violations_list = []
# Training iterations.
iteration_count = 0
for (image, label, group) in celeb_a_train_data_w_group(BATCH_SIZE):
# Assign current batch to input, labels and groups tensors.
input_tensor.assign(image)
labels_tensor.assign(label)
groups_tensor.assign(group)
# Run gradient update.
optimizer.minimize(problem, var_list=var_list)
# Record objective and violations.
objective = problem.objective()
violations = problem.constraints()
sys.stdout.write(
"\r Iteration %d: Hinge Loss = %.3f, Max. Constraint Violation = %.3f"
% (iteration_count + 1, objective, max(violations)))
# Snapshot model once in SKIP_ITERATIONS iterations.
if iteration_count % SKIP_ITERATIONS == 0:
objective_list.append(objective)
violations_list.append(violations)
# Save snapshot of model weights.
model_constrained.save_weights(
temp_directory + "/celeb_a_constrained_" +
str(iteration_count / SKIP_ITERATIONS) + ".h5")
iteration_count += 1
if iteration_count >= NUM_ITERATIONS:
break
# Choose best model from recorded iterates and load that model.
best_index = tfco.find_best_candidate_index(
np.array(objective_list), np.array(violations_list))
model_constrained.load_weights(
temp_directory + "/celeb_a_constrained_" + str(best_index) + ".0.h5")
# Remove temp directory.
os.system("rm -r " + temp_directory)
# + [markdown] id="6r-6_R_gSrsT"
# After having applied the constraint, we evaluate the results once again using Fairness Indicators.
# + id="5G6B3OR9CUmo"
model_location = save_model(model_constrained, 'model_export_constrained')
eval_result_constrained = get_eval_results(model_location, 'eval_results_constrained')
# + [markdown] id="sVteOnE80ATS"
# As with the previous time we used Fairness Indicators, deselect false_negative_rate and select false_positive_rate to look at the metric we are interested in.
#
# Note that to fairly compare the two versions of our model, it is important to use thresholds that set the overall false positive rate to be roughly equal. This ensures that we are looking at actual change as opposed to just a shift in the model equivalent to simply moving the threshold boundary. In our case, comparing the unconstrained model at 0.5 and the constrained model at 0.22 provides a fair comparison for the models.
# + id="GRIjYftvuc7b"
eval_results_dict = {
'constrained': eval_result_constrained,
'unconstrained': eval_results_unconstrained,
}
tfma.addons.fairness.view.widget_view.render_fairness_indicator(multi_eval_results=eval_results_dict)
# + [markdown] id="lrT-7EBrcBvV"
# With TFCO's ability to express a more complex requirement as a rate constraint, we helped this model achieve a more desirable outcome with little impact to the overall performance. There is, of course, still room for improvement, but at least TFCO was able to find a model that gets close to satisfying the constraint and reduces the disparity between the groups as much as possible.
| g3doc/tutorials/Fairness_Indicators_TFCO_CelebA_Case_Study.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="me5o4Lx0TxLE"
# ### Install Pygeohash library
#
# This library provides functions for computing geohash
#
# + [markdown] id="r9KEiCHOKNmL"
# ## Step 5 - Develop Model - Task 6 - Connect the dots & Task 7 - Graph Analytics - CLASS ASSIGNMENTS
# + colab={"base_uri": "https://localhost:8080/"} id="b9R7tS923Q4O" outputId="6380f385-25d7-4568-8b27-3758449c4520"
# !pip install pygeohash
# + [markdown] id="yjjmQTJvUBRh"
# ### Import pygeohash, networkx and Pandas libraries
#
# Pygeohash - functions for converting latitude, longitude to geohash and related distance measurement utilities
#
# Networkx - functions for creating, manipulating and querying open source network graphs
#
# Pandas - Python functions for table manipuation
# + id="Og2j0Urk3K2D"
import pygeohash as pgh
import networkx as nx
import pandas as pd
# + [markdown] id="uSm82HpWUnuB"
# ### Connect to datasets using Google drive or local files
# + id="Y8GrIKFJ3YuE"
using_Google_colab = True
using_Anaconda_on_Mac_or_Linux = False
using_Anaconda_on_windows = False
# + colab={"base_uri": "https://localhost:8080/"} id="1S4wXHrh4AJd" outputId="fec0d4c9-93ba-4403-9655-ef727018390c"
if using_Google_colab:
from google.colab import drive
drive.mount('/content/drive')
# + [markdown] id="z0q-b28UKv6e"
# ##DM6.1 Open Notebook, read Lat, Long and compute Geohash - Activity 1
# + colab={"base_uri": "https://localhost:8080/", "height": 235} id="Hbdkra_c4Jvf" outputId="25644740-042e-4fa3-8f83-f2c39c599266"
if using_Google_colab:
state_location = pd.read_csv('/content/drive/MyDrive/COVID_Project/input/state_lat_long.csv')
if using_Anaconda_on_Mac_or_Linux:
state_location = pd.read_csv('../input/state_lat_long.csv')
if using_Anaconda_on_windows:
state_location = pd.read_csv(r'..\input\state_lat_long.csv')
state_location.loc[0:5,]
# + [markdown] id="rqQ623gxVWA6"
# ### Apply a function call to convert Lat, Long to Geohash
# + id="rxHCyTgl6hZw"
def lat_long_to_geohash(lat_long):
return pgh.encode(lat_long[0], lat_long[1])
# + colab={"base_uri": "https://localhost:8080/", "height": 359} id="UGO4CqdJ5UV9" outputId="7a679901-2433-4a8f-e25b-67acf5def33b"
state_location['geohash'] = state_location[['latitude',
'longitude']].apply(lat_long_to_geohash,
axis=1)
state_location.iloc[0:10,]
# + [markdown] id="RGwAoz-UVlWI"
# ### Truncate geohash to first two characters
# + colab={"base_uri": "https://localhost:8080/", "height": 359} id="HdBh95xJ86qh" outputId="d55917a2-de55-4b5d-e430-569ec057b79c"
state_location['geohash'] = state_location.geohash.str.slice(stop=2)
state_location.iloc[0:10,]
# + [markdown] id="UZD4qG6SLYcM"
# ## DM6.2 - Design Graph representaing States and Geohash
# + [markdown] id="gSlx7_kEV2XV"
# ### Find neighbors by sorting the states by 2 character geohash codes attached to each state
# + [markdown] id="8xqPTiNKWCdR"
# ### Initialize Graph and create state and geohash concepts as nodes
# + id="fOyOGSfNDBFM"
GRAPH_ID = nx.DiGraph()
GRAPH_ID.add_node('state')
GRAPH_ID.add_node('geohash')
# + [markdown] id="JhdS8SEzWOnV"
# ### Create a node for each state
# + id="GL8rq-ylKwAz"
state_list = state_location.state.values
for state in state_list:
GRAPH_ID.add_node(state)
GRAPH_ID.add_edge('state', state, label='instance')
# + [markdown] id="4gCHHKClWfDE"
# ### Create a list of unique geohash codes and create a node for each geohash
# + id="KzUVFIDQLk0M"
geohash_list = state_location.geohash.values
for geohash in geohash_list:
GRAPH_ID.add_node(geohash)
GRAPH_ID.add_edge('geohash', geohash, label='instance')
# + id="3HXi1jtNLMIR"
df_state_geohash = state_location[['state', 'geohash']]
for state_geohash in df_state_geohash.itertuples():
GRAPH_ID.add_edge(state_geohash.state, state_geohash.geohash,
label='located_at')
GRAPH_ID.add_edge(state_geohash.geohash, state_geohash.state,
label='locates',
distance=0.0)
# + [markdown] id="ctpFnpaPLxmO"
# ## DM6.3 - Which states are in Geohash 9q
# + [markdown] id="7vKnRysJOv-Y"
# ### Find geohash associated with California and Naveda
#
# + colab={"base_uri": "https://localhost:8080/"} id="th1YHdHsVIDa" outputId="8dc7ac5e-cb8c-404f-dc64-d5983331cf58"
list(GRAPH_ID.neighbors('CA'))
# + colab={"base_uri": "https://localhost:8080/"} id="RrTXjBV9VbgU" outputId="5e2d9c4b-a09e-4c47-99e4-b3cc386f142c"
list(GRAPH_ID.neighbors('NV'))
# + [markdown] id="4SiWFFecPHCR"
# ### Find States locsted with geohash '9q'
# + colab={"base_uri": "https://localhost:8080/"} id="v3zjYR4RVgzx" outputId="c22ab496-0c55-4574-d0b3-60bbd4aba6fb"
list(GRAPH_ID.neighbors('9q'))
# + [markdown] id="OoNCNf3xMJxS"
# ## DM6.4 Sort the data and find neighbors sharing geohash
# + [markdown] id="hIUHCvSBPSuw"
# ### Find states located with geohah for all geohashes
# + colab={"base_uri": "https://localhost:8080/"} id="unzACFJoWAaX" outputId="eff5d67d-11a7-48f8-cec5-68650adc494c"
for geohash in GRAPH_ID['geohash']:
print("Geohash: ", geohash, "States: ", list(GRAPH_ID.neighbors(geohash)))
# + [markdown] id="ecPQw61cMc55"
# ## DM6.5 Use Graph to find Geohash associated with NY - CLASS ASSIGNMENT
# + colab={"base_uri": "https://localhost:8080/"} id="tE-A4H4jMuT-" outputId="183070d5-0423-4773-9b24-c00d1e9033d8"
list(GRAPH_ID.neighbors('NY'))
# + id="WxUhz4ynMuh9"
# + [markdown] id="NFNlWHfQMvNR"
# ## DM6.6 Use Graph to find which states are in Geohash 'dr' - CLASS ASSIGNMENT
# + colab={"base_uri": "https://localhost:8080/"} id="FhlOwhntM8Qt" outputId="60b4cc2b-7f56-4900-f0c6-d9f98a38597b"
list(GRAPH_ID.neighbors('dr'))
# + id="d-4QW8DsM8dh"
# + [markdown] id="srwyqkTORLVV"
# ## Step 4 - Develop Model - Task 7 - Graph Analytics - DM7.1 Activity 1 - Find number of state and geohash nodes in a graph
# + colab={"base_uri": "https://localhost:8080/"} id="goJ2RMcxFAuF" outputId="59268590-ad42-44cf-ae36-38c79db436ef"
len(list (GRAPH_ID.neighbors('geohash')))
# + colab={"base_uri": "https://localhost:8080/"} id="8SOFj2yjRplz" outputId="1c9f2ad3-2e4e-4602-a51c-b867173aaa8e"
len(list (GRAPH_ID.neighbors('state')))
# + [markdown] id="ULxy711bOpyA"
# ## DM7.2 - Find all neighboring states for NY
# + [markdown] id="sQUUYKEgKmDc"
# ### Connect neighboring geohash codes if the distance is less than 1,000 km
# + id="WIyPeG5iJhUo"
for geohash_1 in geohash_list:
for geohash_2 in geohash_list:
if geohash_1 != geohash_2:
distance = pgh.geohash_haversine_distance(geohash_1, geohash_2)
if distance < 1000000:
GRAPH_ID.add_edge(geohash_1, geohash_2, label='near')
# + [markdown] id="rbEYIVBIdd7D"
# ### Find path length from NY to all nodes (states and geohashes)
# + colab={"base_uri": "https://localhost:8080/"} id="FZS8GFS-L-Tl" outputId="0a868104-4470-4436-c8ef-afcc85b2d983"
neighbor_path_length = nx.single_source_dijkstra_path_length(GRAPH_ID, 'NY', weight='distance')
neighbor_path_length
# + [markdown] id="LZ0UzPx2dyVc"
# ### Make a list of all nodes covered in the path length and then find those nodes which are states and less than or equal to 3 hops
# + id="cglrEuNVMacW"
neighbor_states = neighbor_path_length.keys()
# + colab={"base_uri": "https://localhost:8080/"} id="c24d1JfBHtBm" outputId="9a96479b-23a3-4dc0-cdb2-997ee0def8ea"
state_list = (list (GRAPH_ID.neighbors('state')))
for state in state_list:
if state in neighbor_states:
if neighbor_path_length[state] <= 3:
print(state)
# + [markdown] id="RTAk14L9PAlK"
# ## DM7.3 - Find all neighboring states for each state
# + colab={"base_uri": "https://localhost:8080/"} id="mBflHi8qRKSj" outputId="790f60a5-8c33-4667-dd99-394e36f040c1"
for state_1 in state_list:
neighbor_path_length = nx.single_source_dijkstra_path_length(GRAPH_ID, state_1)
neighbor_state_list = neighbor_path_length.keys()
next_door_list = []
for state_2 in neighbor_state_list:
if state_1 != state_2:
if state_2 in state_list:
if neighbor_path_length[state_2] <=3:
next_door_list.append(state_2)
if next_door_list:
print(state_1, next_door_list)
# + [markdown] id="hblgSpHlPsf0"
# ## DM7.4 - Find path between two states
# + colab={"base_uri": "https://localhost:8080/"} id="WXblSdaKMo6d" outputId="23e02536-04e2-4906-f883-7feeefd7fe49"
nx.dijkstra_path(GRAPH_ID, 'NY', 'CA', weight='distance')
# + colab={"base_uri": "https://localhost:8080/"} id="8WYbKR6qVwKp" outputId="50aa27b8-2471-419f-8d12-b0cc29f5785f"
nx.dijkstra_path(GRAPH_ID, 'OR', 'CA', weight='distance')
# + colab={"base_uri": "https://localhost:8080/"} id="zgM81gMmOQUA" outputId="9aa11245-c15d-483f-917a-e2806711a937"
GRAPH_ID.nodes()
# + colab={"base_uri": "https://localhost:8080/"} id="IbFV9hyqRzhZ" outputId="0e7d645e-0076-4a53-d9da-1fd0271732c5"
nx.single_source_dijkstra_path_length(GRAPH_ID, 'NY')
| Notebook-Class-Assignment-Answers/Step-4-Develop-Model-Task-6-Connect-the-Dots-Class-Assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
#hide
from nbdev_testing.core import *
# # Nbdev Test Run!
#
# > Running through the nbdev process, end to end
# This file becomes your README and also the index of your documentation.
# ## Install
# `pip install nbdev_testing`
# ## How to use
# Using the functions in nbdev testing is easy!
say_sup('Klara')
| index.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Isolated skyrmion in confined helimagnetic nanostructure
# **Authors**: <NAME>, <NAME>, <NAME>
#
# **Date**: 26 June 2016
#
# This notebook can be downloaded from the github repository, found [here](https://github.com/computationalmodelling/fidimag/blob/master/doc/ipynb/isolated_skyrmion.ipynb).
# ### Problem specification
# A thin film disk sample with thickness $t=10 \,\text{nm}$ and diameter $d=150 \,\text{nm}$ is simulated. The material is FeGe with material parameters [1]:
#
# - exchange energy constant $A = 8.78 \times 10^{-12} \,\text{J/m}$,
# - magnetisation saturation $M_\text{s} = 3.84 \times 10^{5} \,\text{A/m}$, and
# - Dzyaloshinskii-Moriya energy constant $D = 1.58 \times 10^{-3} \,\text{J/m}^{2}$.
#
# It is expected that when the system is initialised in the uniform out-of-plane direction $\mathbf{m}_\text{init} = (0, 0, 1)$, it relaxes to the isolated Skyrmion (Sk) state (See Supplementary Information in Ref. 1). (Note that LLG dynamics is important, which means that artificially disable the precession term in LLG may lead to other states).
# ### Simulation
from fidimag.micro import Sim
from fidimag.common import CuboidMesh
from fidimag.micro import UniformExchange, Demag, DMI
# The cuboidal thin film mesh which contains the disk is created:
# +
# Mesh dimensions.
d = 150 # diameter (nm)
t = 10 # thickness (nm)
# Mesh discretisation.
dx = dy = 2.5 # nm
dz = 5
mesh = CuboidMesh(nx=int(d/dx), ny=int(d/dy), nz=int(t/dz), dx=dx, dy=dy, dz=dz, unit_length=1e-9)
# -
# Since the disk geometry is simulated, it is required to set the saturation magnetisation to zero in the regions of the mesh outside the disk. In order to do that, the following function is created:
def Ms_function(Ms):
def wrapped_function(pos):
x, y, z = pos[0], pos[1], pos[2]
r = ((x-d/2.)**2 + (y-d/2.)**2)**0.5 # distance from the centre
if r <= d/2:
# Mesh point is inside the disk.
return Ms
else:
# Mesh point is outside the disk.
return 0
return wrapped_function
# To reduce the relaxation time, we define a state using a python function.
def init_m(pos):
x,y,z = pos
x0, y0 = d/2., d/2.
r = ((x-x0)**2 + (y-y0)**2)**0.5
if r<10:
return (0,0, 1)
elif r<30:
return (0,0, -1)
elif r<60:
return (0, 0, 1)
else:
return (0, 0, -1)
# Having the magnetisation saturation function, the simulation object can be created:
# +
# FeGe material paremeters.
Ms = 3.84e5 # saturation magnetisation (A/m)
A = 8.78e-12 # exchange energy constant (J/m)
D = 1.58e-3 # Dzyaloshinkii-Moriya energy constant (J/m**2)
alpha = 1 # Gilbert damping
gamma = 2.211e5 # gyromagnetic ration (m/As)
# Create simulation object.
sim = Sim(mesh)
sim.Ms = Ms_function(Ms)
sim.driver.alpha = alpha
sim.driver.gamma = gamma
# Add energies.
sim.add(UniformExchange(A=A))
sim.add(DMI(D=D))
sim.add(Demag())
# Since the magnetisation dynamics is not important in this stage,
# the precession term in LLG equation can be set to artificially zero.
sim.driver.do_precession = False
# Initialise the system.
sim.set_m(init_m)
# -
# Now the system is relaxed:
# PYTEST_VALIDATE_IGNORE_OUTPUT
# Relax the system to its equilibrium.
sim.driver.relax(dt=1e-13, stopping_dmdt=0.01, max_steps=5000, save_m_steps=None, save_vtk_steps=None)
# The magnetisation components of obtained equilibrium configuration can be plotted in the following way:
# +
import matplotlib.pyplot as plt
# %matplotlib inline
def plot_magnetisation(m, layer=0):
n_layer = int(d/dx) * int(d/dy)
m.shape = (-1, 3)
mx = m[:, 0][layer*n_layer:(layer+1)*n_layer]
my = m[:, 1][layer*n_layer:(layer+1)*n_layer]
mz = m[:, 2][layer*n_layer:(layer+1)*n_layer]
mx.shape = (int(d/dx), int(d/dy))
my.shape = (int(d/dx), int(d/dy))
mz.shape = (int(d/dx), int(d/dy))
extent = [0, d, 0, d]
plt.figure(figsize=(20, 10))
plt.subplot(1, 3, 1)
plt.imshow(mx, extent=extent)
plt.title('mx')
plt.xlabel('x (nm)')
plt.ylabel('y (nm)')
plt.subplot(1, 3, 2)
plt.imshow(my, extent=extent)
plt.xlabel('x (nm)')
plt.ylabel('y (nm)')
plt.title('my')
plt.subplot(1, 3, 3)
plt.imshow(mz, extent=extent)
plt.xlabel('x (nm)')
plt.ylabel('y (nm)')
plt.title('mz')
plot_magnetisation(sim.spin, layer=0)
# -
# ### References
#
# [1] <NAME>. Ground state search, hysteretic behaviour, and reversal mechanism of skyrmionic textures in confined helimagnetic nanostructures. *Sci. Rep.* **5**, 17137 (2015).
| sandbox/more-dmi/isolated_skyrmion.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Finding the Optimal Learning Rate on K-MNIST using Baysian Optimization
#
# Within this blog, I am giving a short introduction into Bayesian optimization to find a near optimal learning rate. There exists a lot of great tutorials regarding the theory of Bayesian optimization. The main objective of this blog is to give a hands-on tutorial for hyperparameter optimization. As I will cover the theory only very briefly, it is recommend to read about the latter first before going through this tutorial. I am training a small ResNet implemented in PyTorch on the Kuzushiji-MNIST (or K-MNIST) dataset. This tutorial covers the following steps:
#
# 1. **Download and import the K-MNIST dataset into our project**
# 2. **Define a small ResNet in Pytorch**
# 3. **Define everything needed for Bayesian Optimization**
# 4. **Using Bayesian Optimization to find the optimal learning rate**
# 4. **Some practical approaches for learning rate optimization (logarithmic feature transformation)**
#
# What this tutorial will **not** cover:
# * Introduction to PyTorch
# * Gaussian Processes
#
# A basic understanding of Python and PyTorch are required.
# ***
# But first things first, let's make some general imports and set the seed so we do not need to worry about them later:
#
# # Finding the Optimal Learning Rate on K-MNIST using Baysian Optimization
#
# Within this blog, I am giving a short introduction into Bayesian
# optimization to find a near optimal learning rate. There exists a
# lot of great tutorials regarding the theory of Bayesian optimization.
# The main objective of this blog is to give a hands-on tutorial for
# hyperparameter optimization. As I will cover the theory only very briefly,
# it is recommend the read about the latter first before going through this
# tutorial. I am training a small ResNet implemented in PyTorch on the
# Kuzushiji-MNIST (or K-MNIST) dataset. This tutorial covers the following
# steps:
#
# 1. **Download and import the K-MNIST dataset into our project**
# 2. **Define a small ResNet in Pytorch**
# 3. **Define everything needed for Bayesian Optimization**
# 4. **Using Bayesian Optimization to find optimal learning rate**
# 4. **Some practical approaches for learning rate optimization**
#
# What this tutorial will **not** cover:
# * Introduction to PyTorch
# * Gaussian Processes
#
# ***
# A basic understanding of Python and PyTorch are required. But first things first, let's make some general imports and set some seeds so we do not need to worry about them later:
#
# +
import os
import torch
import torchvision
import numpy as np
from tqdm.notebook import tqdm
import matplotlib.pyplot as plt
# set seeds
seed=94
np.random.seed(seed)
torch.manual_seed(seed)
# -
# Done! So let's dive into the first step...
# ***
# ## Download and import the K-MNIST dataset
#
# As mentioned before, we are training on the K-MNIST dataset. Luckily, this dataset is part of the `torchvision` package, which we have just imported! This makes it very straightforward to create a training and validation dataset as the dataset is downloaded autonomously (if it is not already downloaded) and imported in the desired format. We can pass each dataset to PyTorch's `DataLoader`, which represents an iterable over the dataset.
# +
# define a batch size
batch_size = 32
# define transformations we want to apply to our images
transform = torchvision.transforms.Compose(
[torchvision.transforms.ToTensor()])
# create a dataset and dataloader for training
train_ds = torchvision.datasets.KMNIST(root='./data', train=True,
download=True, transform=transform)
train_loader = torch.utils.data.DataLoader(train_ds, batch_size=batch_size,
shuffle=True, num_workers=2)
# create a dataset and dataloader for validation
val_ds = torchvision.datasets.KMNIST(root='./data', train=False,
download=True, transform=transform)
val_loader = torch.utils.data.DataLoader(val_ds, batch_size=batch_size,
shuffle=True, num_workers=2)
# -
# That's it! We have just prepared our data. Let's see how our images look like.
# +
def show_batch(images):
"""
Prints one batch of images in a single image.
Parameters
----------
images:
Batch of images in training data from the DataLoader.
"""
images = torchvision.utils.make_grid(images).numpy()
plt.imshow(np.transpose(images, (1, 2, 0)))
plt.show()
# get one batch
images, labels = next(iter(train_loader))
# plot
show_batch(images)
# -
# Looks like some japanese characters! **Make sure to run the DataLoader cell again, as we took one batch already.**
#
# It's time to create the model.
# ___
# ## Defining a small ResNet
#
# We use a small ResNet9 (8 convolutional layers and 1 fully-connected layer) as it is small yet provides reasonable performance. This is the structure of the network:
#
# <br/>
# <img src="img/ResNet9.png" alt="drawing" width="400"/>
# <br/>
#
# #### Convolutional Blocks
# In my eyes, it looks less cluttered if we aggregate multiple layers to blocks. So let's start by defining the convolutional blocks shown above. These consist of a convolutional layer, batch normalization, ReLU activation and (eventually) MaxPooling:
# +
import torch.nn as nn
import torch.nn.functional as F
class ConvBlock(nn.Module):
"""
Block consisting of a convolutional layer, batch-norm, relu activation and max-pooling (if needed).
"""
def __init__(self, in_channels, out_channels, kernel_size=3, padding=1, pool=False, pool_kernel_size=2):
super(ConvBlock, self).__init__()
self.conv = nn.Conv2d(in_channels, out_channels, kernel_size=kernel_size, padding=padding)
self.conv_bn = nn.BatchNorm2d(out_channels)
if pool:
self.pooling = nn.MaxPool2d(pool_kernel_size)
else:
self.pooling = None
def forward(self, x):
out = F.relu(self.conv_bn(self.conv(x)))
if self.pooling is not None:
out = self.pooling(out)
return out
# -
# #### Residual Blocks
#
# Next, let's define the residual blocks shown above. These blocks consists of two convolutional blocks without MaxPooling:
class ResidualBlock(nn.Module):
"""
Residual block consisting of 2 convolutional blocks.
"""
def __init__(self, in_channels, out_channels, kernel_size=3, padding=1):
super(ResidualBlock, self).__init__()
self.conv_block1 = ConvBlock(in_channels, out_channels, kernel_size, padding)
self.conv_block2 = ConvBlock(in_channels, out_channels, kernel_size, padding)
def forward(self, x):
residual = x
out = self.conv_block1(x)
out = self.conv_block2(out)
out += residual
return out
# ### ResNet9
#
# Now it is straightforward to define our ResNet. Simply concatenate the blocks as shown above and add an additional MaxPolling and a fully-connected layer at the end. Note that we do not add a Softmax layer at the end, as the Cross-Entropy loss, which we are going to use later, includes this already:
class ResNet9(nn.Module):
"""
ResNet consisting of 8 convolutional layers, 1 fully-connected layer and some forward paths for residuals.
"""
def __init__(self, in_channels, num_classes):
super(ResNet9, self).__init__()
# 1st and 2nd convolutional layer
self.conv_block1 = ConvBlock(in_channels, 64)
self.conv_block2 = ConvBlock(64, 128, pool=True)
# residual block consisting of the 3rd and 4th convolutional layer
self.res_block1 = ResidualBlock(128, 128)
# 5th and 6th convolutional layers
self.conv_block3 = ConvBlock(128, 256, pool=True)
self.conv_block4 = ConvBlock(256, 512, pool=True)
# residual block consisting of the 7th and 8th convolutional layer
self.res_block2 = ResidualBlock(512, 512)
# final fully-connected layer
self.classifier = nn.Sequential(nn.MaxPool2d(3),
nn.Flatten(),
nn.Linear(512, num_classes))
def forward(self, x):
out = self.conv_block1(x)
out = self.conv_block2(out)
out = self.res_block1(out)
out = self.conv_block3(out)
out = self.conv_block4(out)
out = self.res_block2(out)
out = self.classifier(out)
return out
# That's it! We have our model now. So let's dive into the next step.
#
# ___
#
# ## Bayesian Optimization
#
# A naive solution to find promising learning rates is to sample learning rates equidistantly or randomly in the search space. This is the concept behind grid and random search. While this is easy to use when function evaluation is cheap, it becomes infeasible when the function evaluation is costly. The latter is typically the case in deep learning. However, we can do better using Bayesian optimization. Bayesian optimization uses probability theory to predict the *most promising* learning rate candidate based on previously evaluated learning rates.
#
# ### Objective Function and Surrogate Model
#
# But how do we predict the most promising next learning rate? Well, if you think about it, what we actually would like to know is some kind of function, which maps a learning rate to a performance metric, for instance the loss. If we would have such a function, we can simply take the minimum of it to find the best learning rate possible. Let's call the latter the *objective function*. Obviously, we don't have access to the objective function (otherwise, you wouldn't be here). And evaluating the objective function for a huge number of learning rates is also infeasible, as we already said. However, what we can do is to evaluate a few learning rates and try to fit a *model* to the objective function. The key idea is to fit the model until it decently represents the objective function. Then, we can instead search for the model's minimum to find a surrogate optimal learning rate. This is why we call such a model a *surrogate model* in Bayesian optimization. We are going to use a *Gaussian Process (GP)* as our surrogate model. Without observation noise, a GP can be interpreted as an interpolator, which - in contrast to other interpolators - additionally gives us information about its uncertainty between two data samples. The uncertainty measure between two data samples is crucial and one of the most distinct features of Bayesian optimization. The latter is used to exert exploration in the search space(or rather the learning rate's space). The higher the uncertainty within a certain search space region the more exploration we need to do. Note that we are not going to implement GPs ourselves here. There are tons of libraries out there. Instead, we are using `sklearn`'s implementation, as shown later.
#
# ### Acquisition Function
#
# Okay, assume for now that we have a surrogate model, which does not yet fits the objective function very well. How do we actually choose the next most promising learning rate to evaluate? This is where the *aquisition funtion* comes into play. We are using it to determine what learning rate is the most promising for the *current* GP fitting. Hence, the acquisition function can be interpreted as a one-step utility measure. A popular choice for the acquisition function is *Expected Improvement* (EI). For our task, the improvement is defined as the improvement over the current best learning rate. Hence, the improvement $I$ at time step $t$ is defined as
#
# \begin{equation}
# I^{(t)}(\lambda) = \max (0, L_{inc}^{(t)} - L(\lambda)) \,, \label{eq:improvement}
# \end{equation}
#
# where $\lambda$ is the learning rate and $L_{inc}^{(t)}$ is the best loss experienced so far, which we call current incumbent. The corresponding learning rate is $\lambda_{inc}^{(t)} = \mathrm{argmin}_{\lambda' \in \mathcal{D}^{(t)}} L(\lambda')$, where $\mathcal{D}^{(t)}$ is the dataset containing all learning rates $\lambda'$ evaluated until time step $t$. This equation has an intuitive appeal; an improvement is achieved if our model predicts a loss smaller than the loss of the current incumbent. The best improvement possible can be achieved at the smallest loss, $\min\, L(\lambda)$.
#
# The Expected Improvement additionally considers uncertainty and is defined - as the name suggests - as the expectation over the improvement $I^{(t)}$
#
# \begin{equation}
# u_{EI}^{(t)}(\lambda) = \mathop{\mathbb{E}}[I^{(t)}(\lambda)] = \int_{-\infty}^{\infty} p^{(t)}(L|\lambda) \times I^{(t)}(\lambda) \, dL \,.
# \end{equation}
#
# The latter can be computed in the closed form yielding:
#
# \begin{equation}
# u_{EI}^{(t)}(\lambda)=
# \begin{cases}
# \sigma^{(t)}(\lambda) [ Z \Phi(Z) + \phi(Z) ],& \text{if } \sigma^{(t)}(\lambda) > 0\\
# 0, & \text{if } \sigma^{(t)}(\lambda) = 0
# \end{cases} \,,
# \end{equation}
#
# where $Z = \frac{L_{inc}^{(t)} - \mu^{(t)} (\lambda) - \xi }{\sigma^{(t)}(\lambda)}$ and $\xi$ is an optional exploration parameter. Note that $\phi$ is the PDF and $\Phi$ is the CDF of the standard normal distribution.
#
# Now we can predict the next promising learning rate using our utility function
#
# \begin{equation}
# \lambda^{(t+1)} = \mathrm{argmax}_{\lambda \in \Lambda} u_{EI}^{(t)}(\lambda) \,,
# \end{equation}
#
# where $\Lambda$ is the search space.
#
# That's it! We now know everything to write our own Bayesian optimizer. Let's start coding! We are going to define a class, which contains everything needed for Bayesian Optimization. Below, you can see the respective class. Let me first show you the code before explaining.
# +
from scipy.stats import norm
from matplotlib import gridspec
class BayesianOptimizer:
"""
This is a Bayesian Optimizer, which takes in a function to optimize, and finds the
maximum value of a parameter within a bounded search space. It uses Expected Improvement as the
acquisition function.
Attributes
----------
f: function
Function to optimize.
gp: GaussianProcessRegressor
Gaussian Process used for regression.
mode: str
Either "linear" or "logarithmic".
bound: list
List containing the lower and upper bound of the search space. IMPORTANT: If mode is "logarithmic",
the bound specifies the minimum and maximum exponents!
size_search_space: int
Number of evaluation points used for finding the maximum of the acquisition function. Can be interpreted
as the size of our discrete search space.
search_space: ndarray
Vector covering the search space.
gp_search_space: ndarray
The search space of GP might be transformed logarithmically depending on the mode, which is why it
might differ from our defined search space.
dataset: list
List containing all data samples used for fitting (empty at the beginning).
states: list
List containing the state of each iteration in the optimization process (used for later plotting).
"""
def __init__(self, f, gp, mode, bound, size_search_space=250):
if mode not in ["linear", "logarithmic"]:
raise ValueError("%s mode not supported! Chose either linear or logarithmic." % mode)
else:
self.mode = mode
self.f = f
self.gp = gp
self.min = bound[0]
self.max = bound[1]
self.size_search_space = size_search_space
if mode == "linear":
self.search_space = np.linspace(self.min, self.max, num=size_search_space).reshape(-1, 1)
self.gp_search_space = self.search_space
else:
self.search_space = np.logspace(self.min, self.max, num=size_search_space).reshape(-1, 1)
self.gp_search_space = np.log10(self.search_space)
self.dataset = []
self.states = []
def _ei(self, c_inc, xi=0.05):
"""
Expected Improvement (EI) acquisition function used for maximization.
Parameters
----------
c_inc: float
Utility of current incumbent.
xi: float
Optional exploration parameter.
Returns
-------
util: ndarray
Utilization given the current Gaussian Process and incumbent
"""
# calculate the current mean and std for the search space
mean, std = self.gp.predict(self.gp_search_space, return_std=True)
std = np.array(std).reshape(-1, 1)
# calculate the utilization
a = (mean - c_inc - xi)
z = a / std
util = a * norm.cdf(z) + std * norm.pdf(z)
return util
def _max_acq(self):
"""
Calculates the next best incumbent for the current dataset D.
Returns
-------
x_max: float
Location (x-coordinate) of the next best incumbent
util_max: float
Utility of the next best incumbent.
util: ndarray
Utility function for the search space.
"""
# get the value of the current best incumbent
c_inc = np.max(np.array(self.dataset)[:, 1])
# calculate the utility function
util = self._ei(c_inc)
# check if the utilization is all zero
if np.all((util == 0.)):
print("Warning! Utilization function is all zero. Returning a random point for evaluation.")
x_max = self.search_space.reshape(-1)[np.random.randint(len(self.search_space))]
util_max = 0.0
else:
# get the maximum's location and utility
x_max = self.search_space.reshape(-1)[util.argmax()]
util_max = util.max()
return x_max, util_max, util
def eval(self, n_iter=10, init_x_max=None):
"""
Runs n_iter evaluations of function f and optimizes its parameter using Bayesian Optimization.
Parameters
----------
n_iter: int
Number of iterations used for optimization
init_x_max: float
Initial guess of the parameter. If none, a random initial guess is sampled in the search space.
Returns
-------
best_return_x: float
Best sample found during optimization
best_return_param:
Parameters defining the best function (e.g., torch model).
"""
# get a random initial value for the incumbent from our search space if not specified
if not init_x_max:
x_max = self.search_space[np.random.randint(len(self.search_space))]
x_max = x_max.item()
else:
x_max = init_x_max
# for storing the best return and some parameters specifying it
best_return = None
best_return_x = None
best_return_param = None
for i in range(n_iter):
# print some information
print("\nBO Iteration %d --> Chosen parameter: %f %s" % (i, x_max,
"" if (init_x_max or i != 0) else "(randomly)"))
# evaluate the function
y, param = self.f(x_max)
# store if it is the best
if not best_return or y > best_return:
best_return = y
best_return_x = x_max
best_return_param = param
# add the new sample to the dataset
self.dataset.append([x_max, y])
# get all the data samples in the dataset
xs = np.array(self.dataset)[:, 0].reshape(-1, 1)
ys = np.array(self.dataset)[:, 1].reshape(-1, 1)
# fit the GP with the updated dataset
if self.mode == "linear":
self.gp.fit(xs, ys)
else:
self.gp.fit(np.log10(xs), ys)
# calculate the maximum utilization and its position
x_max, util_max, util = self._max_acq()
# save the state for later plotting
self.states.append({"dataset": self.dataset.copy(),
"util": util,
"GP": self.gp.predict(self.gp_search_space, return_std=True)})
return best_return_x, best_return_param
def save_all_plots(self):
"""
Saves all plots.
"""
self.plot_all(show=False, save=True)
def plot_all(self, show=True, save=True):
"""
Plots all states/iterations made during optimization until now.
Parameters
----------
show: bool
If true, plot is shown directly.
save: bool
If true, plot is saved.
"""
for id, state in enumerate(self.states):
self.plot_state(state, id, show=False, save=save)
if show:
plt.show()
def plot_iteration(self, it, show=True, save=True):
"""
Plots a certain iteration of the optimization process.
Parameters
----------
it: int
Iteration of the optimization process
show: bool
If true, plot is shown directly.
save: bool
If true, plot is saved.
"""
# get the corresponding state
state = self.states[it]
self.plot_state(state, it, show=show, save=save)
def plot_state(self, state, fig_number, show=True, save=True, additional_func=None):
"""
Plots a state of the optimization process.
Parameters
----------
state: dict
Dictionary storing the dataset, utilization and GP describing one state during optimization.
fig_number: int
Id of the figure to plot.
show: bool
If true, plot is shown directly.
save: bool
If true, plot is saved.
additional_func: (function, name)
Additional function to plot.
"""
# reshape search space as this is more convenient for plotting
search_space = self.search_space.reshape(-1)
# get all information of the corresponding state
dataset = state["dataset"]
util = state["util"].reshape(-1)
gp = state["GP"]
# create figure with two plots (ax1: GP fitting, ax2: utility function)
figure = plt.figure(fig_number)
gs = gridspec.GridSpec(nrows=2, ncols=1, height_ratios=[3, 1], figure=figure)
ax1 = figure.add_subplot(gs[0])
ax1.set_xticklabels([]) # turn off x labeling of upper plot
ax1.set_title("Iteration %d" % fig_number)
ax2 = figure.add_subplot(gs[1])
# check if we need to set a logarithmic scale
if self.mode == "logarithmic":
ax1.set_xscale("log")
ax2.set_xscale("log")
# adjust borders to make it look better
figure.subplots_adjust(bottom=.14, top=.95)
# plot an additional function if given
if additional_func:
func, name = additional_func
add_ys, _ = func(search_space)
ax1.plot(search_space, add_ys, color="red", label=name)
# plot the GP mean and std
mu, std = gp
mu = mu.reshape(-1)
ax1.plot(search_space, mu,
color="blue", label="GP mean")
ax1.fill_between(search_space,
mu - (std * 1), mu + (std * 1),
color="blue", alpha=0.3, label="GP std")
# plot the dataset
xs = np.array(dataset)[:, 0]
ys = np.array(dataset)[:, 1]
ax1.scatter(xs, ys, color="blue", label="Dataset")
# plot the utility function
ax2.plot(search_space, util, color="green", label="Utility function")
ax2.fill_between(search_space,
np.zeros_like(util),
util.reshape(-1), alpha=0.3, color="green")
figure.legend(loc="lower center", ncol=5 if additional_func else 4)
if save:
if not os.path.exists('./plots'):
os.makedirs('./plots')
fig_name = "./plots/BO_iteration_%d" % fig_number
plt.savefig(fig_name)
if show:
plt.show()
# -
# The latter might look overwhelming at first, but it's actually straightforward. Let's go through it function by function:
#
# - `__init__()`: Here we initialize everything needed for our optimizer. The most important parts are the objective function (`self.f`), the Gaussian proccess (`self.gp`, defined later), the search space (`self.search_space`) and the search space for the Gaussian process (`self.gp_search_space`). But why do we have two search spaces? Well, you'll see later that it might be very beneficial to transform the GP's search space to a logarithmic space. More on that later!
#
#
# - `_ei()`: This function defines the Expected Improvement (EI) acquisition function as described above.
#
#
# - `_max_acq()`: This function calculates the best next incumbent based on the acquisition function. It simply calculates the utility function for our bounded and discrete search space $\Lambda$ (`self.search_space`) and determines where the maximum is.
#
#
# - `eval()`: This function evaluates the given function (`self.f`), fits the GP and determines the next incumbent using `_max_acq`. This is done for `n_iter` iterations.
#
#
# Note that we have defined the Bayesian optimizer in a way that it is *maximizing* the objective function. That is, we need to take the negative of the objective function in case of a minimization problem (as it is the case for the loss).
#
# ***
# ### Testing our Bayesian Optimizer
#
# Okay, now we have our Bayesian optimizer. Let's try it on a simple example. Therefore, we need to define an objective function first:
def objective(x):
return x**2 * np.sin(5 * np.pi * x)**6.0, None
# I have taken this objective function from another [blog](https://machinelearningmastery.com/what-is-bayesian-optimization/), which provides a great tutorial for basic Bayesian optimization. I can recommend to check it out as well!
#
# Note that the objective function returns a tuple consisting of the actual return of the function and an additional parameter, which is `None` in this case. The latter is used later when we want to know, which ResNet model yielded what loss in order to save its parameters and probably continue training from there on. Let's take a quick look at our objective function:
#
# +
def plot_function(func, xs, fig_id=0):
fig = plt.figure(0)
ax = fig.add_subplot()
ys, _ = objective(xs)
ax.plot(xs, ys, color="red")
ax.set_title("Objective Function")
xs = np.linspace(0,1, 250)
plot_function(objective, xs)
plt.show()
# -
# Okay, seems like our objective has several maxima from which the one at $x=0.9$ is the best in our interval. Let's see if our optimizer can find it.
#
# However, we need to define the kernel, the Gaussian Process and some bounds first. We use a product kernel here consisting of a constant kernel and a Radial Basis Function (RBF) kernel. This is the default setting for sklearn as well.
# +
from sklearn.gaussian_process import GaussianProcessRegressor
from sklearn.gaussian_process.kernels import RBF, ConstantKernel as C
kernel = C(1.0, (1e-5, 1e5)) * RBF(10, (1e-2, 1e2))
gp = GaussianProcessRegressor(kernel, n_restarts_optimizer=10)
bo = BayesianOptimizer(objective, gp, mode="linear", bound=[0, 1])
# -
# As mentioned above, we are not going to talk much about the theory of GPs. If you want to learn more, you might want to checkout this [blog](https://peterroelants.github.io/posts/gaussian-process-tutorial/). However, we are going to talk a bit about the importance of kernels later.
#
# For now, let's run 10 iterations of Bayesian optimization using our class and plot the results. Note that our optimizer stores everything we need to know about an iteration during the optimization process in `self.states`, so that plotting is easier.
# +
# sklearn might throw some annoying warnings, let's surpress them
import warnings
warnings.filterwarnings("ignore")
# give an initial guess of 0.5 and optimize
maximum, _ = bo.eval(10, 0.5)
print("\n--> Best Value found: ", maximum)
# plot the results
for i, state in enumerate(bo.states, start=1):
bo.plot_state(state, i, show=False, additional_func=(objective, "Objective"))
plt.show()
# -
# The red line shows the objective function (as shown above), the blue line shows the mean of the GP, the light blue area shows the GP's standard deviation and the green line shows the EI utility function. Go through the images and track the way our optimizer works. On the last image you can see that our optimizer found $0.892$ as the best result after 10 iterations, which is quite near the global maximum at $0.9$. However, it is not guaranteed that the optimizer finds the global maximum on a small number of iterations. It might only find a local maximum, as the one at $0.5$. Okay, now we can go on to the main part of this blog.
#
# ___
# # Using Bayesian Optimization to find the Optimal Learning Rate
#
# We want to find an optimal (or near optimal) learning rate for our classification task on K-MNIST. Therefore, we need to think more thoroughly about what our objective functions is. As mentioned before, we are using the loss $L$. But which loss exactly? The one calculated on a batch? Or the one after ten batches? Are we using the training or the validation loss?
#
# Well, our main goal in a classification task is to decrease the loss on validation data. And even though, function evaluation is expensive, K-MNIST is a rather small dataset. This is why we are going to evaluate on the **validation loss after training one epoch**. In doing so, we are optimizing the learning rate with respect to the loss we care most about and on all data provided.
#
# That is, our function to evaluate, which is given to the Bayesian optimizer, takes the learning rate and the dataset (training + validation) as the input and returns the average validation loss (as well as the torch model). As we are evaluating on one epoch, the function is called `run_one_epoch()`, as shown below.
#
# Note that, our function is returning the negative loss since our optimizer tries to maximize the objective function (and we are interested in a small loss). Moreover, we are also calculating the accuracy, as it is more human-readable.
# +
def accuracy(pred, true):
class_index_pred = pred.detach().numpy().argmax(axis=1)
return np.sum(true.detach().numpy() == class_index_pred) / len(pred)
def run_one_epoch(lr, train_l, val_l, seed):
"""
Runs one epoch of training using the specified learning rate lr and returns the negative average
validation loss.
Parameters
----------
lr: float
Learning rate of the model.
train_l: DataLoader
Torch's DataLoaders constituting an iterator over the training dataset.
val_l: DataLoader
Torch's DataLoaders constituting an iterator over the validation dataset.
seed: int
Seed for Numpy and Torch.
Returns
-------
Tuple containing the negative validation loss and the model trained on the specified learning rate.
"""
# set the seed to initialize same model and randomness on all epochs to allow fair comparison
np.random.seed(seed)
torch.manual_seed(seed)
# get our model and define the optimizer as well as the loss criterion
model = ResNet9(in_channels=1, num_classes=10)
optimizer = torch.optim.SGD(model.parameters(), lr=lr)
criterion = torch.nn.CrossEntropyLoss()
train_loop = tqdm(train_l) # tqdm wrapper used to print progress bar
for data in train_loop:
# unpack images and labels
images, labels = data
# zero the parameter gradients
optimizer.zero_grad()
# calculate loss
outputs = model(images)
loss = criterion(outputs, labels)
# calculate and apply the gradients
loss.backward()
optimizer.step()
# print some training information
train_loop.set_postfix({"Loss": loss.item(), "Accuracy": accuracy(outputs, labels)})
# let's validate our model
print("Validating ...")
with torch.no_grad():
cum_val_loss = 0.0
cum_acc = 0.0
for data in val_l:
# unpack images and labels
images, labels = data
# calculate loss
outputs = model(images)
cum_val_loss += criterion(outputs, labels)
cum_acc += accuracy(outputs, labels)
# print some validation information
avg_val_loss = cum_val_loss / len(val_loader)
avg_val_acc = cum_acc / len(val_loader)
print("---> Validation-Loss: %.4f & Validation-Accuracy: %.4f" % (avg_val_loss, avg_val_acc))
print("\n", "-"*60,"\n")
return -avg_val_loss, model
# -
# Basically, the `run_one_epoch()` method consists of two loops; the training and the validation loop. While the model is optimized during the training loop, it is kept fixed during validation. We have everything needed now to find the optimal learning rate. However, as for our example, we need to define a kernel, a GP and some bounds. As can be seen from our bounds defined below, our search space covers learning rates from $0.00001$ to $1.0$ since learning rates smaller than that are very uncommon on the first epoch. Let's run it for 10 iterations and see what happens!
# +
n_iter = 10 # number of iterations
np.random.seed(seed) # set seed to allow fair comparison to logarithmic
# define the GP
kernel = C(1.0, (1e-5, 1e5)) * RBF(10, (1e-2, 1e2))
gp = GaussianProcessRegressor(kernel, n_restarts_optimizer=10)
# optmize the learning rate
bo = BayesianOptimizer(lambda x: run_one_epoch(x, train_loader, val_loader, seed),
gp,
mode="linear",
bound=[1e-5, 1.0])
found_lr, best_model = bo.eval(n_iter=10)
print("\n--> Found learning-rate after %d iterations: %f" % (n_iter, found_lr))
# plot all iterations
bo.plot_all()
# save the best model (this is the one returned from BO)
torch.save(best_model.state_dict(), "./ResNet9_linear")
# -
# Okay, let's take a look at the results. The optimizer is giving us a learning rate of $0.205$ that was achieved in the 8th iteration with a validation loss of $0.2946$ and an accuracy of $91.47\%$, which is quite good for the first epoch. However, I don't know about you, but I was wondering a bit about the way the optimizer was searching for the learning rate. If you take a closer look at the graph, you'll see that nearly all evaluations were done between $0.1$ and $1.0$. If you have ever tried to manually tune a learning rate on a classification task or at least read about commonly used learning rates, you'll find these learning rates quite huge. Moreover, you'll find the way the optimizer is searching for the optimal learning rate quite counterintuitive. Usually, we tune learning rates with an exponential decay, e.g., starting from $1\times 10^{-1}$ and going smaller to $1\times 10^{-2}$, $1\times 10^{-3}$, and so on.
#
# Let's take a moment and think about why the optimizer is evaluating learning rates in a way, which might look counterintuitive to us. At first, let's take a closer look at the search space defined in the `BayesianOptimizer` class (for the linear case, ignore the logarithmic case for the moment). It uses **linear spacing**. Let's take a look at such a search space with the same bounds yet a smaller number of samples:
np.linspace(1e-5, 1.0, 100)
# You will quickly realize that most samples lie between $1\times 10^{-1}$ and $1\times 10^{0}$. This is due to the fact that linear spacing causes equidistant spreading, which is different to our logarithmic way of tuning. As you might know, this is an easy fix. We can just use logarithmic spacing (note that `np.logspace` is expecting you to give *exponents* as the bound):
# uses base 10 by default
np.logspace(-5,0, 100)
# Great! We can now evaluate each exponent equally in our search space, similar to what we would do when manually tuning the learning rate.
#
# However, we can't just use logarithmic spacing. Something you might not know (because we skipped it here) is that the main component for calculating the covariance in GPs is the *kernel*. As mentioned before, we took a RBF kernel (Constant kernel is not important here). Let's take a look at the kernel function for our case
#
# \begin{equation}
# k(\lambda_i, \lambda_j) = \exp\left( - \frac{d(\lambda_i, \lambda_j)^2}{2l^2} \right)\, .
# \end{equation}
#
# $\lambda_i$ and $\lambda_j$ are two learning rates, $d(.,.)$ is the Euclidean distance between those points and $l$ is the length parameter for scaling the covariance.
#
# The part I want to point your attention to is the distance $d(.,.)$. As for a lot of kernels, this is the main metric for calculating the covariance between two points. Our intention when using logarithmic spacing is that we would like to explore each exponent equally. However, because our kernel is using the distance for calculating the covariance, it yields higher covariance for greater distance and vice versa. And since Expected Improvement yields higher utility with higher variance (c.f., equations above), our optimizer would still favour greater exponents.
#
# However, the fix is easy here as well. We can simply **transform the search space logarithmically** when working with GPs. That is, the kernel is computed on a transformed learning rate $\psi(\lambda) = (\log_{10}(\lambda))$. The kernel is then
#
# \begin{equation}
# k(\psi(\lambda_i), \psi(\lambda_j)) = \exp\left( - \frac{d(\psi(\lambda_i), \psi(\lambda_j))^2}{2l^2} \right)\, .
# \end{equation}
#
# Note that we do not transform the targets during GP fitting. This means that the loss is equal for both cases $L(\lambda) = L(\psi(\lambda))$.
#
# Let's recap briefly before we try that.
#
# 1. We are using logarithmic *spacing* to have a search space, which contains the same amount of data points for each exponent. In doing so, we pay equal attention to each exponent.
#
# 2. We are using logarithmic *feature transformation* when working with GPs. This turns our non-linearly (here logarithmically) spaced search space into a linearly spaced search space. In doing so, we are encouraging the optimizer to search on small exponents as well.
#
# Okay, enough theory! Let's see the results. Everything needed is already implemented in the `BayesianOptimizer` class. We only need to switch the mode from linear to logarithmic and change the bounds to exponents.
# +
n_iter = 10 # number of iterations
np.random.seed(seed) # set seed to allow fair comparison to linear
# define the GP
kernel = C(1.0, (1e-5, 1e5)) * RBF(10, (1e-2, 1e2))
gp = GaussianProcessRegressor(kernel, n_restarts_optimizer=10)
# optmize the learning rate
bo = BayesianOptimizer(lambda x: run_one_epoch(x, train_loader, val_loader, seed),
gp,
mode="logarithmic",
bound=[-5, 0])
found_lr, best_model = bo.eval(n_iter=10)
print("\n--> Found learning-rate after %d iterations: %f" % (n_iter, found_lr))
# plot all iterations
bo.plot_all()
# save the best model (this is the one returned from BO)
torch.save(best_model.state_dict(), "./ResNet9_log")
# -
# Yeay! We were able to decrease the loss from $0.2946$ to $0.1843$ and increase our accuracy from $91.47\%$ to $94.45\%$. The learning rate found is significantly smaller then the one found before: linear-mode $\rightarrow 0.204$ vs. logarithmic-mode $\rightarrow 0.0037$! Especially in the first iterations, you can see the that the variance is high in *both* directions, which is exactly what we wanted. In the last iteration, you can see that our optmizer paid equal attention to all exponents in our search space.
#
# ## Conclusion
#
# One might argument that the difference between linear and logarithmic mode in performance isn't that high. Actually, that's true. However, paying more attention to smaller learning rates becomes more important in later epochs, where greater learning rates often cause stagnation in learning.
#
# Finally, I want to point out that, even though it might be useful to exploit our domain knowledge to transform the learning rate's search space logarithmically, it might be different for other hyperparameters, which we want to optimize as well. So, be careful with that!
| Optimal_Learning_Rate_KMNIST.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exercícios Python e Numpy
#
# O objetivo desse notebook é ajudar na fixação dos conteúdos que passamos na aula de Python e Numpy. Sabemos que acabamos passando meio rápido durante a aula, então o objetivo aqui é conseguir treinar os conceitos para você conseguir usa-los na prática mais tarde.
#
# Qualquer dúvida pode ficar a vontade para perguntar no Slack ou diretamente para gente, vamos ficar felizes em ajudar :)
# ## Python
#
# Aqui na parte de Python vamos passar por algumas das principais coisas que você precisa saber, é claro que não colocamos tudo de importante da linguagem, só o mais necessário.
# ### Variáveis
# +
# Declare uma variavel chamada a e atribua o valor 10 a ela
# +
# Imprime essa variavel que você acabou de criar
# +
# Crie uma outra variavel b que recebe o valor de a so que como string
# -
# Combine sua variavel b a variavel abaixo para obter a string "Hello 10" em uma variavel d
c = "Hello "
# +
# Imprima a variável d
# -
# ### Strings
my_str = 'Insira uma frase aqui!'
# +
# Substitua a exclamação da frase por uma interrogação
# (Dica: A funcão altera inplace ou retorna uma copia modificada?)
# +
# Crie uma lista "my_words" com cada palavra a frase
# -
# ### Listas
lista = [1, 23, 31, 40, 56, 16]
# +
# Faça um for que imprima cada elemento da lista "lista" (Lembre-se que o for em python é um for each)
# +
# Faça um for que imprima o dobro de cada elemento da lista "lista"
# +
# Gere uma lista chamada "dobro" com o dobro de cada elemento de "lista" usando list comprehension
# +
# Crie uma nova lista chamada "pares"
# Faça um for que itetere sobre a lista "lista" e para cada elemento impar coloque ele no fim da lista "pares"
# Imprima a lista "pares"
# -
lista2 = ['oi', 2, 2.5, 'top', 'python', 45]
# +
# Faça um for pela "lista2" e imprima todos os elementos que são strings (Dica: pesquise pela função type())
# -
# #### Indexando
my_list = [0, 10, 20, 30, 40, 50, 60, 70]
# +
# Selecione o ultimo elemento da lista
# +
# Selecione do primeiro até o 4 elemento da lista
# +
# Selecione do segundo elemento da lista até o quinto
# +
# Selecione do primeiro elemento da lista até o penultimo
# -
# ### Dicionários
lista = ['a', 'a', 'b', 'a', 'c', 'd', 'e', 'b', 'b', 'c']
# Crie um dicionario que contenha a contagem de cada elemento do vetor
my_dict = {}
#...
print(my_dict)
# ### Funções
# Crie uma função soma_elementos() que recebe uma lista e retorna a soma de todos os seus elementos
def soma_elementos(lista):
return 0
soma_elementos([1, 2, 3, 4, 5])
soma_elementos([-1, 5, 7, -2])
# +
# Crie uma função produto_escalar() que recebe duas listas de tamanho igual e calcula o produto escalar entre elas
# Dica: Utilize a função zip
# -
produto_escalar([1, 2, 3], [0, 4, 7])
produto_escalar([10, 20, 40, 1], [23, 4, 2, 1])
# +
# Crie uma função par_ou_impar() que recebe um numero n é para cada numero de 1 a n imprime o numero
# seguido de par ou impar, dependendo do que ele seja. Caso o usuário não coloque nada n deve valer 20
# Exemplo: par_ou_impar(4)
# 1 Impar
# 2 Par
# 3 Impar
# 4 Par
# -
par_ou_impar(15)
par_ou_impar(15)
# +
# Crie uma função diga_indice() que recebe uma lista e imprime o indice de cada elemento e em seguida
# o proprio elemento
# Exemplo: diga_indice(['oi', 'tudo', 'bem'])
# 0 oi
# 1 tudo
# 2 bem
# (DICA: Pesquise pela função enumerate)
# -
diga_indice(['1', '2', '3'])
diga_indice(['a', 'b', 'c', 'd', 'e'])
# ## Numpy
#
# O elemento central do numpy são os arrays, então aqui vamos treinar muitas coisas sobre eles
# Importando a biblioteca
import numpy as np
# ### Arrays
# +
a = np.array([1, 2, 3, 4, 5, 6, 7])
b = np.array([[1, 2, 3, 4],
[5, 6, 7, 8]])
c = np.zeros((3,4))
# +
# Pense em qual é a shape de cada um dos arrays acima
# Depois de pensar imprima cada um deles para conferir sua resposta
# +
# Crie um array de uma dimenção com 20 elementos inteiros aletórios entre 0 e 23bb
# +
# Crie um array de uns com shape (4, 5)
# +
# Crie um array shape (4, 2) onde cada entrada vale 77
# (Dica: Talvez vc tenha que usar uma multiplicação)
# +
# Gere um array chamado my_sequence com os numeros 0, 10, 20, 30, ..., 90, 100]
# -
# ### Indexando
my_array = np.random.randint(50, size=(15,))
print(my_array)
# +
# Selecione todos os elementos entre o quinto e o decimo primeiro (intervalo fechado)
# +
# Selecione todos os elementos maiores que 20
# -
my_matrix = np.array([[1, 2, 3, 4],
[5, 6, 7, 8],
[9, 10, 11, 12],
[13, 14, 15, 16]])
# +
# Selecione o elemento na primeira linha da terceira coluna
# +
# Selecione o elemento na primeira linha da ultima coluna
# +
# Selecione os elementos da matriz para obter o seguinte
# [[6, 7],
# [10, 11]]
# +
# Selecione os elementos da matriz para obter o seguinte
# [[2, 3, 4],
# [6, 7, 8]]
# +
# Selecione os elementos da ultima coluna inteira
# +
# Selecione os elementos da 2a linha inteira
# -
# ### Operações
my_array = np.random.randint(10, size=(5,))
print(my_array)
# +
# Some 10 a todos os elementos de my_array
# +
# Multiplique todos os elementos de my_array por 4
# +
# Obtenha a soma de todos os elementos de my_array
# +
# Obtenha a média de todos os elementos de my_array
# +
# Obtenha o indice do maior elemento de my_array
# -
my_array = np.random.randint(10, size=(5,))
my_other_array = np.random.randint(10, size=(5,))
print(my_array, '\n')
print(my_other_array)
my_array = np.random.randint(10, size=(5,))
my_other_array = np.random.randint(10, size=(10,5))
print(my_array, '\n')
print(my_other_array)
# +
# Some my_array elemento por elemento em cada linha de my_other_array
# -
my_array = np.random.randint(10, size=(5,4))
my_other_array = np.random.randint(10, size=(10,5))
print(my_array, '\n')
print(my_other_array)
# +
# Faça a multiplicação entre my_other_array e my_array
# +
# Descubra a soma dos valores de cada linha de my_other_array
# (Dica: Pesquise sobre o atributo axis da função de soma)
# -
my_array = np.random.randint(10, size=(5,4))
print(my_array)
# +
# Usando reshape transforme a matriz acima em um vetor (Concatendo a linha de baixo na de cima)
# -
np.array([[ 0, 1, 2, 3, 4, 5, 6, 7],
[ 8, 9, 10, 11, 12, 13, 14, 15]])
# +
# Gere a array anterior usando np.arange e a função reshape
| Aula0/exercicios_python_numpy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19"
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load in
import numpy as np
np.random.seed(42)
import pandas as pd
import string
import re
import gensim
from collections import Counter
import pickle
import tensorflow as tf
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split
from sklearn.metrics import roc_auc_score, f1_score
from sklearn import metrics
from keras.models import Model
from keras.layers import Input, Dense, Dropout, Conv1D, Embedding, SpatialDropout1D, concatenate
from keras.layers import GRU, LSTM,Bidirectional, GlobalAveragePooling1D, GlobalMaxPooling1D
from keras.layers import CuDNNLSTM, CuDNNGRU
from keras.preprocessing import text, sequence
from keras.callbacks import Callback
from keras import optimizers
from keras.layers import Lambda
from keras.callbacks import *
import warnings
warnings.filterwarnings('ignore')
from nltk.corpus import stopwords
import os
os.environ['OMP_NUM_THREADS'] = '4'
import gc
from keras import backend as K
from sklearn.model_selection import KFold
from unidecode import unidecode
import time
eng_stopwords = set(stopwords.words("english"))
# + _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a"
# 1. preprocessing
train = pd.read_csv("../input/train.csv")
test = pd.read_csv("../input/test.csv")
print("Train shape : ",train.shape)
print("Test shape : ",test.shape)
# + _uuid="5b42925bff7c0275450d2cb4e2fcbfeef00f5252"
# 1-a. Count non ascii characters
special_character = re.compile(r'[A-Za-z0-9\.\-\?\!\,\#\@\% \'\/\"]',re.IGNORECASE)
train['spl_chars'] = train['question_text'].apply(lambda x: len(special_character.sub('', str(x))))
test['spl_chars'] = test['question_text'].apply(lambda x: len(special_character.sub('', str(x))))
# + _uuid="1b2aa6f947fc1850b8644e461090a83e59337c6c"
#pd.set_option('display.max_colwidth', -1)
#train.head()
# + _uuid="e173c94e68a61528291cc9473cba37f30847bef8"
#train.loc[train.target==0]['spl_chars'].mean()
# + _uuid="ab6d45bc22305cfe00ec7bdceb8a9764f67bd346"
# 2. remove numbers
def clean_numbers(x):
x = re.sub('[0-9]{5,}', '#####', x)
x = re.sub('[0-9]{4}', '####', x)
x = re.sub('[0-9]{3}', '###', x)
x = re.sub('[0-9]{2}', '##', x)
return x
train['clean_text'] = train['question_text'].apply(lambda x: clean_numbers(str(x)))
test['clean_text'] = test['question_text'].apply(lambda x: clean_numbers(str(x)))
# + _uuid="91b0a8237489bbc87efb394ce8a83dc9988831fd"
#train['clean_text']
# + _uuid="848325747b9d511eb48b382a2c565324908e5a83"
#3. remove non-ascii
special_character_removal = re.compile(r'[^A-Za-z\.\-\?\!\,\#\@\% ]',re.IGNORECASE)
def clean_text(x):
x_ascii = unidecode(x)
x_clean = special_character_removal.sub('',x_ascii)
return x_clean
train['clean_text'] = train['clean_text'].apply(lambda x: clean_text(str(x)))
test['clean_text'] = test['clean_text'].apply(lambda x: clean_text(str(x)))
# + _uuid="5c21bc47b5bd5476b7632c43f6dd4546798508b4"
X_train = train['clean_text'].fillna("something").values
y_train = train.target.values
X_test = test['clean_text'].fillna("something").values
# + _uuid="478cbe489de98a6366b82e65ebbbf83264014365"
#X_train
# + _uuid="0a1ddc438df214446749d198ae34fc910ce9e7d9"
def add_features(df):
df['comment_text'] = df['clean_text'].fillna('something').apply(lambda x:str(x))
df['total_length'] = df['comment_text'].apply(len)
df['capitals'] = df['comment_text'].apply(lambda comment: sum(1 for c in comment if c.isupper()))
df['caps_vs_length'] = df['capitals']/df['total_length']
df['num_words'] = df.comment_text.str.count('\S+')
df['num_unique_words'] = df['comment_text'].apply(lambda comment: len(set(w for w in comment.split())))
df['words_vs_unique'] = df['num_unique_words'] / df['num_words']
df['spl_chars_vs_len'] = df['spl_chars']/df['total_length']
return df
train = add_features(train)
test = add_features(test)
# + _uuid="30b2b9829e2c92852aa32f4f00b6f72b351e6440"
train.loc[np.isinf(train.caps_vs_length),'caps_vs_length'] =0
train.loc[np.isinf(train.words_vs_unique),'words_vs_unique'] =0
train.loc[np.isinf(train.spl_chars_vs_len),'spl_chars_vs_len'] =0
# + _uuid="d8e0ed4c1f9e2e3ed785155b1c4fd7609e66eed8"
features = train[['caps_vs_length', 'words_vs_unique', 'spl_chars_vs_len']].fillna(0)
test_features = test[['caps_vs_length', 'words_vs_unique', 'spl_chars_vs_len']].fillna(0)
# + _uuid="de760831762e47616e9f29d89096d1149506b0f9"
#test[test.num_words>=50].count()
# + _uuid="df06403bb00b1af4fe06bca7dc490e8890988d80"
ss = StandardScaler()
ss.fit(np.vstack((features, test_features)))
features = ss.transform(features)
test_features = ss.transform(test_features)
# + _uuid="da25bfaddf3ace9b4319fa5c226813bd429aa4f2"
max_features = 180000
maxlen = 50
tokenizer = text.Tokenizer(num_words=max_features)
tokenizer.fit_on_texts(list(X_train) + list(X_test))
X_train_sequence = tokenizer.texts_to_sequences(X_train)
X_test_sequence = tokenizer.texts_to_sequences(X_test)
x_train = sequence.pad_sequences(X_train_sequence, maxlen=maxlen)
x_test = sequence.pad_sequences(X_test_sequence, maxlen=maxlen)
print(len(tokenizer.word_index))
# + _uuid="fcb923a725451e2e03a6ae5b04d5be496f3c78a0"
# Load the FastText Web Crawl vectors
EMBEDDING_FILE_FASTTEXT='../input/embeddings/wiki-news-300d-1M/wiki-news-300d-1M.vec'
EMBEDDING_FILE_TWITTER='../input/embeddings/glove.840B.300d/glove.840B.300d.txt'
def get_coefs(word, *arr): return word, np.asarray(arr, dtype='float32')
# switching as glove has better support fot this text
embeddings_index_tw = dict(get_coefs(*o.rstrip().rsplit(' ')) for o in open(EMBEDDING_FILE_FASTTEXT,encoding='utf-8'))
embeddings_index_ft = dict(get_coefs(*o.strip().split(' ')) for o in open(EMBEDDING_FILE_TWITTER,encoding='utf-8'))
spell_model = gensim.models.KeyedVectors.load_word2vec_format(EMBEDDING_FILE_FASTTEXT)
# + _uuid="5e8f9f2bb8a57148e11ed3661d0a834d70a83556"
# This code is based on: Spellchecker using Word2vec by CPMP
# https://www.kaggle.com/cpmpml/spell-checker-using-word2vec
words = spell_model.index2word
w_rank = {}
for i,word in enumerate(words):
w_rank[word] = i
WORDS = w_rank
# Use fast text as vocabulary
def words(text): return re.findall(r'\w+', text.lower())
def P(word):
"Probability of `word`."
# use inverse of rank as proxy
# returns 0 if the word isn't in the dictionary
return - WORDS.get(word, 0)
def correction(word):
"Most probable spelling correction for word."
return max(candidates(word), key=P)
def candidates(word):
"Generate possible spelling corrections for word."
return (known([word]) or [word])# or known(edits1(word)) or known(edits2(word))
def known(words):
"The subset of `words` that appear in the dictionary of WORDS."
return set(w for w in words if w in WORDS)
def edits1(word):
"All edits that are one edit away from `word`."
letters = 'abcdefghijklmnopqrstuvwxyz'
splits = [(word[:i], word[i:]) for i in range(len(word) + 1)]
deletes = [L + R[1:] for L, R in splits if R]
transposes = [L + R[1] + R[0] + R[2:] for L, R in splits if len(R)>1]
replaces = [L + c + R[1:] for L, R in splits if R for c in letters]
inserts = [L + c + R for L, R in splits for c in letters]
return set(deletes + transposes + replaces + inserts)
def edits2(word):
"All edits that are two edits away from `word`."
return (e2 for e1 in edits1(word) for e2 in edits1(e1))
def singlify(word):
return "".join([letter for i,letter in enumerate(word) if i == 0 or letter != word[i-1]])
# + _uuid="1a5245ab137d836d57f56eb0f71392b28f049da7"
WORDS
# + _uuid="d9063c74e4b0dcae6bc499b4de88c3dc7ce99e84"
word_index = tokenizer.word_index
nb_words = min(max_features, len(word_index))
embedding_matrix = np.zeros((nb_words,601))
something_tw = embeddings_index_tw.get("something")
something_ft = embeddings_index_ft.get("something")
something = np.zeros((601,))
something[:300,] = something_ft
something[300:600,] = something_tw
something[600,] = 0
# + _uuid="14bf0e68627109d96ded3a11c511d9c829a5c5c2"
def all_caps(word):
return len(word) > 1 and word.isupper()
def embed_word(embedding_matrix,i,word):
embedding_vector_ft = embeddings_index_ft.get(word)
if embedding_vector_ft is not None:
if all_caps(word):
last_value = np.array([1])
else:
last_value = np.array([0])
embedding_matrix[i,:300] = embedding_vector_ft
embedding_matrix[i,600] = last_value
embedding_vector_tw = embeddings_index_tw.get(word)
if embedding_vector_tw is not None:
embedding_matrix[i,300:600] = embedding_vector_tw
# Glove vector is used by itself if there is no glove vector but not the other way around.
for word, i in word_index.items():
if i >= max_features: continue
if embeddings_index_ft.get(word) is not None:
embed_word(embedding_matrix,i,word)
else:
# change to > 20 for better score.
if len(word) > 26:
embedding_matrix[i] = something
#print(word)
else:
word2 = correction(word)
#print(word2)
if embeddings_index_ft.get(word2) is not None:
embed_word(embedding_matrix,i,word2)
else:
word2 = correction(singlify(word))
if embeddings_index_ft.get(word2) is not None:
embed_word(embedding_matrix,i,word2)
else:
embedding_matrix[i] = something
# + _uuid="746f34d01074e5536b5b85b74084b1a378fffce0"
embedding_matrix.shape
# + _uuid="0d20c805d3c19e56a2418f2e1c90ebcd97999cb5"
del(embeddings_index_tw, embeddings_index_ft); gc.collect()
# + _uuid="caa275f237ec4a5eceb3f6329e0c40ec6bce78a6"
class RocAucEvaluation(Callback):
def __init__(self, validation_data=(), interval=1):
super(Callback, self).__init__()
self.interval = interval
self.X_val, self.y_val = validation_data
self.max_score = 0
self.not_better_count = 0
def on_epoch_end(self, epoch, logs={}):
if epoch % self.interval == 0:
y_pred = self.model.predict(self.X_val, verbose=1)
score = roc_auc_score(self.y_val, y_pred)
print("\n ROC-AUC - epoch: %d - score: %.6f \n" % (epoch+1, score))
if (score > self.max_score):
print("*** New High Score (previous: %.6f) \n" % self.max_score)
model.save_weights("best_weights.h5")
self.max_score=score
self.not_better_count = 0
else:
self.not_better_count += 1
if self.not_better_count > 3:
print("Epoch %05d: early stopping, high score = %.6f" % (epoch,self.max_score))
self.model.stop_training = True
# + _uuid="9ff70836e3f36bcbd8a1fe096ad275c8af1e5ede"
def get_model(features,clipvalue=1.,num_filters=40,dropout=0.5,embed_size=601):
features_input = Input(shape=(features.shape[1],))
inp = Input(shape=(maxlen, ))
# Layer 1: concatenated fasttext and glove twitter embeddings.
x = Embedding(max_features, embed_size, weights=[embedding_matrix], trainable=False)(inp)
# Uncomment for best result
# Layer 2: SpatialDropout1D(0.5)
x = SpatialDropout1D(dropout)(x)
# Uncomment for best result
# Layer 3: Bidirectional CuDNNLSTM
x = Bidirectional(CuDNNLSTM(num_filters, return_sequences=True))(x)
# Layer 4: Bidirectional CuDNNGRU
x, x_h, x_c = Bidirectional(CuDNNGRU(num_filters, return_sequences=True, return_state = True))(x)
# Layer 5: A concatenation of the last state, maximum pool, average pool and
# two features: "Unique words rate" and "Rate of all-caps words"
avg_pool = GlobalAveragePooling1D()(x)
max_pool = GlobalMaxPooling1D()(x)
x = concatenate([avg_pool, x_h, max_pool,features_input])
# Layer 6: output dense layer.
outp = Dense(1, activation="sigmoid")(x)
model = Model(inputs=[inp,features_input], outputs=outp)
adam = optimizers.adam(clipvalue=clipvalue)
model.compile(loss='binary_crossentropy',
optimizer=adam,
metrics=['accuracy'])
return model
# + _uuid="48c3d67be5fe44871b5b6c926740f1d93761e9db"
model = get_model(features)
batch_size = 512
# Used epochs=100 with early exiting for best score.
epochs = 7
gc.collect()
K.clear_session()
# Change to 5
num_folds = 5 #number of folds
y_test = np.zeros((test.shape[0],1))
# Uncomment for out-of-fold predictions
scores = []
oof_predict = np.zeros((train.shape[0],1))
kf = KFold(n_splits=num_folds, shuffle=True, random_state=239)
# + _uuid="d0fe5c5f79a3857df30bfbfda75ed44fbbddf3e2"
def f1_smart(y_true, y_pred):
args = np.argsort(y_pred)
tp = y_true.sum()
fs = (tp - np.cumsum(y_true[args[:-1]])) / np.arange(y_true.shape[0] + tp - 1, tp, -1)
res_idx = np.argmax(fs)
return 2 * fs[res_idx], (y_pred[args[res_idx]] + y_pred[args[res_idx + 1]]) / 2
# + _uuid="f77fd6268d31e91d85af3b9714398393737de8df"
bestscore = []
for train_index, test_index in kf.split(x_train):
filepath="weights_best.h5"
kfold_y_train,kfold_y_test = y_train[train_index], y_train[test_index]
kfold_X_train = x_train[train_index]
kfold_X_features = features[train_index]
kfold_X_valid = x_train[test_index]
kfold_X_valid_features = features[test_index]
gc.collect()
K.clear_session()
model = get_model(features)
#ra_val = RocAucEvaluation(validation_data=([kfold_X_valid,kfold_X_valid_features], kfold_y_test), interval = 1)
checkpoint = ModelCheckpoint(filepath, monitor='val_loss', verbose=2, save_best_only=True, mode='min')
reduce_lr = ReduceLROnPlateau(monitor='val_loss', factor=0.5, patience=1, min_lr=0.0001, verbose=2)
earlystopping = EarlyStopping(monitor='val_loss', min_delta=0.0001, patience=2, verbose=2, mode='auto')
if i == 0:print(model.summary())
model.fit([kfold_X_train,kfold_X_features], kfold_y_train, batch_size=batch_size, epochs=epochs, verbose=1,
validation_data=([kfold_X_valid,kfold_X_valid_features], kfold_y_test),
callbacks = [checkpoint, reduce_lr, earlystopping])#ra_val,
gc.collect()
#model.load_weights(bst_model_path)
model.load_weights(filepath)
y_test += model.predict([x_test,test_features], batch_size=1024,verbose=1) / num_folds
gc.collect()
# uncomment for out of fold predictions
oof_predict[test_index] = model.predict([kfold_X_valid, kfold_X_valid_features],batch_size=batch_size, verbose=1)
cv_score = roc_auc_score(kfold_y_test, oof_predict[test_index])
f1, threshold = f1_smart(np.squeeze(kfold_y_test), np.squeeze(oof_predict[test_index]))
print('Optimal F1: {:.4f} at threshold: {:.4f}'.format(f1, threshold))
bestscore.append(threshold)
scores.append(cv_score)
print('score: ',cv_score)
print("Done")
print('Total CV score is {}'.format(np.mean(scores)))
# + _uuid="2361998e3847bc4a874a46cc0b27b510826a7ded"
# + _uuid="0f3013249d7f16ca1fa6dea824a4c399046715a6"
from sklearn.metrics import f1_score
def threshold_search(y_true, y_proba):
best_threshold =0
best_score = 0
for threshold in [i * 0.01 for i in range(100)]:
score = f1_score(y_true=y_true, y_pred=y_proba > threshold)
if score > best_score:
best_threshold = threshold
best_score = score
search_result = {'threshold': best_threshold, 'f1': best_score}
return search_result
search_result = threshold_search(y_train, oof_predict)
print(search_result)
print("Mean of Best Score ::: {}".format(np.mean(bestscore)))
# + _uuid="d9850e3a5ff5045d549fe66cc873b037f5c05d47"
#sum((y_test>.38).reshape(-1)==1)
#sum(y_train)
# + _uuid="72a6df50370a51a36fbcb3203f8cd33eb8d2e0e7"
sub = test[['qid']]
y_test = y_test.reshape((-1, 1))
pred_test_y = (y_test>search_result['threshold']).astype(int)#np.mean(bestscore)
sub['prediction'] = pred_test_y
sub.to_csv("submission.csv", index=False)
# + _uuid="a86ceaa0021086bc57f4deff49cbfeeb2828ebdf"
| kaggle-quora-insincere-question/clean-slate-v3 (1).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # [L&S 88] Open Science -- Project 1, part 1
#
# ---
#
# ### Instructors <NAME> and <NAME>
#
# In this notebook we will be covering different approaches to Exploratory Data Analysis (EDA), exploring how different techniques and approachs can lead to different results and conclusions about data.
#
# We will be exploring a controversial dataset which has led many data scientists down different analytical paths. This notebook contains autograder tests from [Gofer Grader](https://github.com/data-8/Gofer-Grader). Some of the tests check that you answered correctly, but some are not comprehensive. Autograder cells look like this:
#
#
# ```python
# check('tests/q1-1.py')
# ```
#
# If you pass the tests, the cell will output `All tests passed!`.
#
# *Estimated Time: 120 minutes*
#
# ---
#
# ### Topics Covered
# - Exploratory Data Analysis
# - Understanding past studies with data
#
# ### Table of Contents
#
# 1 - [Introduction to Study](#section1)<br>
#
#
# 2 - [Introduction to EDA](#section2)<br>
#
#
# 3 - [More EDA and Visualizations](#section3)<br>
#
# 4 - [More Practice](#section4)<br>
#
# 5 - [Free Response](#section5)<br>
#
# **Dependencies:**
# Please consult the `datascience` library [documentation](http://data8.org/datascience/tables.html) for useful help on functions and visualizations throughout the assignment, as needed.
# +
#Just run me
# if this cell errors, uncomment the line below and rerun
# !pip install gofer-grader
from gofer.ok import check
from datascience import *
import numpy as np
import pandas as pd
import matplotlib
# %matplotlib inline
import matplotlib.pyplot as plt
plt.style.use('fivethirtyeight')
import warnings
warnings.simplefilter('ignore', FutureWarning)
# -
# ---
# ## 1. Introduction to the Study <a id='section 1'></a>
# 
# Creator: <NAME>
#
# Credit: AFP/Getty Images
# Nothing frustrates both soccer fans and players as much as being [red-carded](https://en.wikipedia.org/wiki/Penalty_card#Red_card). In soccer, receiving a red card from the referee means that the player awarded the red card is expelled from the game, and consequently his team must play with one fewer player for the remainder of the game.
#
# Due to the inherently subjective nature of referees' judgments, questions involving the fairness of red card decisions crop up frequently, especially when soccer players with darker complexions are red-carded.
#
# For the remainder of this project, we will explore a dataset on red-cards and skin color and attempt to understand how different approachs to analysis can lead to different conclusions to the general question: "Are referees more likely to give red cards to darker-skinned players?"
#
# ---
#
# ## The Data <a id='data'></a>
#
# In this notebook, you'll be working with a dataset containing entries for many European soccer players, containing variables such as club, position, games, and skin complexion.
#
# Important to note about this dataset is that it was generated as the result of an [observational study](https://en.wikipedia.org/wiki/Observational_study), rather than a [randomized controlled experiment](https://en.wikipedia.org/wiki/Randomized_controlled_trial). In an observational study, entities' independent variables (such as race, height, zip code) are observed, rather than controlled as in the randomized controlled experiment. Though data scientists often prefer the control and accuracy of controlled experiments, often performing one is either too costly or poses ethical questions (e.g., testing trial drugs and placebo treatments on cancer patients at random). Though our dataset was generated organically--in the real world rather than in a laboratory--it is statistically more challenging to prove causation among variables for these kinds of observational studies (more on this in Question 2).
#
#
# Please read this summary of the [dataset's description](https://osf.io/9yh4x/) to familiarize yourself with the context of the data:
#
# *"...we obtained data and profile photos from all soccer players (N = 2053) playing in the first male divisions of England, Germany, France and Spain in the 2012-2013 season and all referees (N = 3147) that these players played under in their professional career. We created a dataset of player dyads including the number of matches players and referees encountered each other and our dependent variable, the number of red cards given to a player by a particular referee throughout all matches the two encountered each other."*
#
# *...implicit bias scores for each referee country were calculated using a race implicit association test (IAT), with higher values corresponding to faster white | good, black | bad associations. Explicit bias scores for each referee country were calculated using a racial thermometer task, with higher values corresponding to greater feelings of warmth toward whites versus blacks."*
#
# Run the cell below to load in the data into a `Table` object from the `datascience` library used in Data 8.
# +
# Just run me
data = pd.read_csv("CrowdstormingDataJuly1st.csv").dropna()
data = Table.from_df(data)
# -
# Here are some of the important fields in our data set that we will focus on:
#
# |Variable Name | Description |
# |--------------|------------|
# |`player` | player's name |
# |`club` | player's soccer club (team) |
# |`leagueCountry`| country of player club (England, Germany, France, and Spain) |
# |`height` | player height (in cm) |
# |`games`| number of games in the player-referee dyad |
# |`position` | detailed player position |
# |`goals`| goals scored by a player in the player-referee dyad |
# |`yellowCards`| number of yellow cards player received from referee |
# |`yellowReds`| number of yellow-red cards player received from referee |
# |`redCards`| number of red cards player received from referee |
# |`rater1`| skin rating of photo by rater 1 (5-point scale ranging from very light skin to very dark skin |
# |`rater2`| skin rating of photo by rater 2 (5-point scale ranging from very light skin to very dark skin |
# |`meanIAT`| mean implicit bias score (using the race IAT) for referee country, higher values correspond to faster white good, black bad associations |
# |`meanExp`| mean explicit bias score (using a racial thermometer task) for referee country, higher values correspond to greater feelings of warmth toward whites versus blacks |
#
#
# As you can see on the table above, two of the variables we will be exploring is the ratings on skin tone (1-5) measured by two raters, Lisa and Shareef. For context, we have added a series of images that were given to them so that you can better understand their perspective on skin tones. Keep in mind that this might affect our hypothesis and drive our conclusions.
#
# Note: On the following images, the only two were the rating for the two raters coincide is image #3 on the top and image #6 on the bottom.
# <img src="L1S1.jpg" style="float: left; width: 30%; margin-right: 1%; margin-bottom: 0.5em;">
# <img src="L1S2.jpg" style="float: left; width: 30%; margin-right: 1%; margin-bottom: 0.5em;">
# <img src="L2S2.jpg" style="float: left; width: 30%; margin-right: 1%; margin-bottom: 0.5em;">
# <img src="L3S4.jpg" style="float: left; width: 30%; margin-right: 1%; margin-bottom: 0.5em;">
# <img src="L4S5.jpg" style="float: left; width: 30%; margin-right: 1%; margin-bottom: 0.5em;">
# <img src="L5S5.jpg" style="float: left; width: 30%; margin-right: 1%; margin-bottom: 0.5em;">
# <p style="clear: both;">
# Run this cell to peek at the data
data
# <b> Question 1.1: </b>
# What is the shape of data? Save the number of variables (columns) as `num_columns` and the number of players (rows) as `num_rows`.
# +
num_columns = ...
num_rows = ...
print("Our dataset has {0} variables and {1} players.".format(num_columns, num_rows))
# -
check('tests/q1-1.py')
# <b>Question 1.2:</b> Which columns should we focus our analysis on? Drop the columns which contain variables which we're not going to analyze. You might consider using the `Table.drop` method to create a transformed copy of our `data`.
# +
cols_to_drop = ["birthday", "victories", "ties", "defeats", "goals"\
, "photoID", "Alpha_3", "nIAT", "nExp"]
data = ...
# Make sure data no longer contains those columns
data
# -
check('tests/q1-2.py')
# <b> Question 1.3: </b>
# Let's break down our remaining variables by type. Create a table with each of the variables' names and their classifications as either "categorical" or "quantitative" variables. In order to do this, use their Python types. *Hint*: Python's `type()` function might be helpful.
# +
python_types = []
# Get the Python type of each variable by looking at the first value in each column of the data
for ... in data:
column_type = type(data[...].item(0))
python_types.append(...)
label_classifications = []
numeric_categorical_vars = ["refNum", "refCountry"] # Numerical variables that aren't quantitative
# Loop through the array of variable Python types and classify them as quantitative or categorical
for ... in np.arange(len(python_types)):
if python_types[...] == str: # If label is a string...
label_classifications.append(...)
elif data.labels[...] in ...: # If label is a categorical numerical variable...
label_classifications.append(...)
else: # If label isn't categorical...
label_classifications.append(...)
# Create a table with the data's labels and label_classifications array
variables = Table().with_columns("variable name", data.labels\
, "classification", ...)
variables.show()
# -
check('tests/q1-3.py')
# <b> Question 1.4:</b> If we're trying to examine the relationship between red cards given and skin color, which variables ought we to consider? Classify the ones you choose as either independent or dependent variables and explain your choices.
# Independent Variables (variables that may correlate or cause red cards):
# ***YOUR ANSWER HERE***
# Dependent Variables (variables which indicate red cards):
# ***YOUR ANSWER HERE***
# ---
# ## 2. Introduction to EDA <a id='section 2'></a>
# 
# An overview of the Data Science Process with EDA highlighted. By Farcaster at English Wikipedia, CC BY-SA 3.0, https://commons.wikimedia.org/w/index.php?curid=40129394
# Exploratory data analysis (EDA) is the stage of the data science process that takes the processed, cleaned dataset and attempts to learn a bit about its structure. In so doing, the data scientist ties to understand basic properties of the data. They seek to:
# - Craft hypotheses about the data
# - Make and assess assumptions for future statistical inference
# - Select useful statistical tools and techniques
# - Create explanatory of suggestive data visualizations
# <b> Question 2.1: </b>
# First, let's compute the minimum, maximum, mean and standard deviation of our data to understand how our data are distributed. *Hint*: look up the `Numpy` documentation for the relevant functions.
stats = data.stats(ops=(min, max, ..., ...))
stats
check('tests/q2-1.py')
# <b> Question 2.2: </b> Now let's take our `statistics` table and enhance it a bit. First, drop the columns with categorical variables. *Hint*: Use the `variables` table we created in Question 1.3.
# +
categorical_vars = variables.where(..., ...)
cols_to_drop = categorical_vars.column(...)
stats = stats.drop(...)
stats
# -
check('tests/q2-2.py')
# <b> Question 2.3: </b> Now that we have some information about the distribution of our data, let's try to get rid of some statistical outliers. Assume that data points with variables valued plus or minus 2 standard deviations (SDs) below those variables' means are statistical outliers. In other words, only data points whose variables' values are within 2 standard deviations on either side of the corresponding means are valid. Get rid of the outliers accordingly.
#
# Formally, we can describe the set of outliers for the $i$th variable, $O_i$, as:
#
# $$O_i = \{\text{values} \mid \text{values} < \mu_i - 2 \sigma_i\}\cup\{\text{values} \mid \text{values} > \mu_i + 2 \sigma_i\}$$
#
# In words, we want the "union of all values of the $i$th variable $\pm$ 2 standard deviations from the mean"
#
# *Hint*: You'll need to look up values in your `stats` table to find the means and SDs for each variable.
# Just run me to drop remaining categorical variables
data = data.drop("player", "position", "leagueCountry", "club", "playerShort", "refNum", "refCountry")
# +
for ... in data.labels:
data = data.where(..., are.above_or_equal_to(
stats.column(...)[...] - 2 * stats.column(...)[...] # mean - 2 * SD
)).where(..., are.below_or_equal_to(
stats.column(...)[...] + 2 * stats.column(...)[...] # mean + 2 * SD
))
data
# -
check('tests/q2-3.py')
# ## 3. More EDA and Visualizations <a id='section 3'></a>
# ##### Hypotheses:
# Two types of general hypotheses can be made about the data. Either:
# ##### $H_A:$ Referees give red cards to darker skinned players with higher (or lower) frequency.
# or
# ##### $H_0:$ Referees give red cards to all players at similar frequencies.
#
# Where $H_A$ and $H_0$ denote a "alternative" hypothesis and a ["null" hypothesis](https://en.wikipedia.org/wiki/Null_hypothesis), respectively.
#
#
# As mentioned before, we typically cannot prove causation in an observational study such as our dataset. Then we can only "reject" our null hypothesis if the our independent variable(s) and dependent variable have a statistically significant correlation, or "fail to reject" (basically, to accept the null) if there is no statistically significant correlation between the variables.
# <b> Question 3.1: </b>
# ##### Scatter plots:
# To analyze the correlation between independent and dependent variables, we may use a scatter plot as a simple form of data visualization between one numerical "x" (independent) variable and one numerical "y" (dependent) variable. Below are a few scatterplot examples a data scientist might generate when asking the questions,"How are implicit and explicit bias correlated?", and "Is a player's height correlated with the number of yellow cards he receives?", respectively.
# +
# Just run this. You don't need to understand this cell
data_df = pd.read_csv("CrowdstormingDataJuly1st.csv")
meanExp = []
meanIAT = []
for index, row in data_df.iterrows():
if row["meanExp"] not in meanExp:
meanExp.append(row["meanExp"])
meanIAT.append(row["meanIAT"])
exps = np.nan_to_num(meanExp)
iats = np.nan_to_num(meanIAT)
# +
# Run to create a table of means
means = Table().with_columns("meanExps", exps, "meanIATs", iats)
means
# +
# Run to display scatter plot meaEXPS vs meanIATS
means.select("meanIATs", "meanExps").scatter( "meanExps", fit_line=True)
# -
# What do you observe from the scatter plot? Why might these two variables be related in this way? Why might this be a coincidence?
# ***YOUR ANSWER HERE***
# Run to display scatter plot
height_yellowCards = data.select("yellowCards", "height")
height_yellowCards.scatter("height", fit_line=True)
# What do you observe from this scatter plot? Why might these two variables be related in this way? Why might this be a coincidence?
# ***YOUR ANSWER HERE***
# <b> Question 3.2: </b>
# ##### Histograms:
# Histograms are a data visualization tool that helps one understand the shape of a single variable's distribution of values. Each bin in a histogram has a width such that the sum of all bin_widths * bin_heights = 1. Histograms are used to plot the empirical (observed) distribution of values. Below is an example histogram a data scientist might generate when asking the questions, "what is the empirical distribution of the `goals` variable?"
# +
# Run to display histogram of skin colors, as measure by rater 1
goals = data.select("rater1")
goals.hist("rater1", bins=np.arange(0, 1, 0.2))
# +
# Run to display histogram of skin colors, as measure by rater 2
goals = data.select("rater2")
goals.hist("rater2", bins=np.arange(0, 1, 0.2))
# -
# What do you observe from the histograms? Why could this be?
# ***YOUR ANSWER HERE***
# <b> Question 3.3: </b>
# Now create a histogram with the empirical distribution of red cards and a histogram with the empirical distribution of yellow cards. Then create a histogram that displays both simultaneously and describe your findings and offer an explanation.
# +
yellows = ...
reds = ...
yellows.hist(...)
reds.hist()
data.hist([...])
# -
# *Describe and explain your findings*
# ***YOUR ANSWER HERE***
# <b> Question 3.4: </b>
# ##### Box plots:
# Box plots are a data visualization that also allows a data scientist to understand the distribution of a particular variable or variables. In particular, it presents data in percentiles (25, 50, 75%) to give a more standardized picture of the spread of the data. Below is an example of box plots in a side-by-side layout describing the distribution of mean and explicit biases, respectively. Please refer to [this article](https://pro.arcgis.com/en/pro-app/help/analysis/geoprocessing/charts/box-plot.htm) for more information on what the components of a box plot mean.
# +
# Run to create boxplot. We will be using the table of "means" we created in 3.1
means.select("meanIATs", "meanExps").boxplot()
# -
# What do you observe from the box plots? Why might each distribution may be shaped like this?
# ***YOUR ANSWER HERE***
# <b> Question 3.5: </b>
# Now create a pair of side-by-side box plots analyzing the distribution of two comparable variables (i.e., red and yellow cards). Then describe your findings and offer an explanation.
# +
### Create an illustrative data visualization ###
# -
# *Describe your findings and explain why the data may be distributed this way.*
# ***YOUR ANSWER HERE***
# ---
# ## 4. More Practice<a id='section 4'></a>
# +
# Just run me to reload our dropped variables into our data table
data = pd.read_csv("CrowdstormingDataJuly1st.csv").dropna()
data = Table.from_df(data)
# -
# Observe below how we're able to use a pivot table to make an insightful series of bar plots on the number of red cards awarded by referees officiating in different leagues across Europe. The number to the left of the plots' y axes represents the number of red cards awarded in those kinds of games. The labels of the plots' y axes is the number of games in that particular referee/league combination for the given number of red cards.
agg = data.pivot("redCards", "leagueCountry")
agg
agg.bar("leagueCountry", overlay=False)
# ### Question 4.1:
# Interpret what you see.
# *** YOUR ANSWER HERE ***
# Observe below how we are again able to use a pivot table to make a similar bar plot--this time aggregating the number of games with certain amounts of red cards given by referees of different countries. Note: the referees' countries are anonimized as random, numerical IDs.
agg = data.pivot("redCards", "refCountry")
agg
agg.bar("refCountry", overlay=False)
# ### Question 4.2:
# Interpret each plot. Explain what the peaks in these bar plots represent.
# ***YOUR ANSWER HERE ***
# Observe below the further use of pivot tables to break down the distribution of red cards by player position.
agg = data.pivot("redCards", "position")
agg
agg.bar("position", overlay=False, width=20)
# ### Question 4.3:
# Interpret each plot. What [positions](https://en.wikipedia.org/wiki/Association_football_positions) stand out and why might this be?
# ***YOUR ANSWER HERE***
# Observe a scatter plot between victories and games. Intuitively, the two variables are positively correlate and the best fit line has a slope of about 1. This slope is consistent with the fact--with the exception of ties--a win by one team must be accompanied by a loss for the opposing team.
data.scatter("victories", "games", fit_line=True)
# Observe a histogram of the number of games each player has appeared in the dataset.
data.hist("games", bins=np.arange(1, 20))
# ## 5. Free Response <a id='section 5'></a>
# Suppose you wish to analyze the relationship between two variables which you believe are correlated, within the context of the debate on whether skin color correlates with red cards. First, propose a null hypothesis $H_0$ about this relationship, and an alternative hypothesis $H_A$. Then create a data visualization that examines this relationship graphically, explaining why you used that visualization. Finally, describe your observations and decide whether $H_0$ or $H_A$ is more plausible based on your results.
# $H_0$: ***YOUR ANSWER HERE***
#
# $H_A$: ***YOUR ANSWER HERE***
# +
### Create an illustrative data visualization ###
# -
# *Why did you decide to use this type of visualization?*
# ***YOUR ANSWER HERE***
# *Describe your observations and decide which hypothesis is more plausible in that context.*
# ***YOUR ANSWER HERE***
# ---
# Notebook developed by: <NAME>
#
# Data Science Modules: http://data.berkeley.edu/education/modules
#
| Project_2/Soccer/ls88/student-final.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from qutip import *
import numpy as np
basis(2) * 2
# Seems to work just like matrix multiplication
basis(2) + 2
# Oh! It looks like it only applies
# to the "excited" value
np.array([[1], [0]]) + 2
# Just had to double check I
# was remembering matrix arithmetic correctly
identity(4)
# Creates a (4,4) matrix, where the "excited" value
# is moved down a column in each subsequent row
# +
Qobj(identity(4)[:, 0])
# We can retrieve the first column
# by using normal Numpy slicing
# and then putting the result back
# into a Qobj.
# -
basis(4).dag()
# Seems like it just transposes the matrix?
create(4)
# I'm not quite sure what is happening here.
# I can see that the "excited" value is rising,
# but how/why I'm not sure
destroy(4)
# This seems like it's doing the same thing
# as create(N), except the values seem
# to start on the first row, second column.
| Day 55/moremath.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import json
import pandas as pd
import pprint
pp = pprint.PrettyPrinter(indent=2)
from collections import Counter
import re
import numpy as np
dir_data_raw = os.path.join("..", "data", "raw")
data_dir_interim = os.path.join("..", "data", "interim")
datasets = ['biorxiv_medrxiv', 'comm_use_subset', 'noncomm_use_subset', 'pmc_custom_license']
# ## Formatting
# These formatting helper functions are courtesy of [xhlulu](https://www.kaggle.com/xhlulu/cord-19-eda-parse-json-and-generate-clean-csv)
# +
def format_name(author):
middle_name = " ".join(author['middle'])
if author['middle']:
return " ".join([author['first'], middle_name, author['last']])
else:
return " ".join([author['first'], author['last']])
def format_affiliation(affiliation):
text = []
location = affiliation.get('location')
if location:
text.extend(list(affiliation['location'].values()))
institution = affiliation.get('institution')
if institution:
text = [institution] + text
return ", ".join(text)
def format_authors(authors, with_affiliation=False):
name_ls = []
for author in authors:
name = format_name(author)
if with_affiliation:
affiliation = format_affiliation(author['affiliation'])
if affiliation:
name_ls.append(f"{name} ({affiliation})")
else:
name_ls.append(name)
else:
name_ls.append(name)
return ", ".join(name_ls)
def format_body(body_text):
texts = [(di['section'], di['text']) for di in body_text]
texts_di = {di['section']: "" for di in body_text}
for section, text in texts:
texts_di[section] += text
body = ""
for section, text in texts_di.items():
body += section
body += "\n\n"
body += text
body += "\n\n"
return body
def format_bib(bibs):
if type(bibs) == dict:
bibs = list(bibs.values())
bibs = deepcopy(bibs)
formatted = []
for bib in bibs:
bib['authors'] = format_authors(
bib['authors'],
with_affiliation=False
)
formatted_ls = [str(bib[k]) for k in ['title', 'authors', 'venue', 'year']]
formatted.append(", ".join(formatted_ls))
return "; ".join(formatted)
# -
metadata = pd.read_csv(os.path.join(dir_data_raw, "all_sources_metadata_2020-03-13.csv"))
metadata["full_text"] = ""
metadata["file_path"] = None
metadata["results"] = ""
metadata["conclusion"] = ""
def parse_article(full_path, file_path):
with open(full_path) as file:
json_article = json.load(file)["body_text"]
article_sections = []
metadata.loc[index, 'full_text'] = format_body(json_article)
for body_text in json_article:
section_heading = re.sub(r'[^a-zA-Z0-9 ]', '', body_text["section"]).lower().strip()
for section, headings in section_headings.items():
metadata.loc[index, 'full_text'] = article["full_text"] + body_text["text"]
if section_heading in headings:
metadata.loc[index, section] = article[section] + body_text["text"]
# +
section_headings = {
"results": ["results and discussion", "results"],
"conclusion": ["conclusion", "conclusions", "discussion and conclusions"]
}
for index, article in metadata.iterrows():
# We only need to update if there's a full text
if article["has_full_text"]:
for dataset in datasets:
file_path = os.path.join(dataset, dataset, str(article["sha"]) + ".json")
metadata.loc[index, "file_path"] = file_path
full_path = os.path.join(dir_data_raw, file_path)
if os.path.exists(full_path):
parse_article(full_path, file_path)
# -
metadata.to_csv(os.path.join(data_dir_interim, "1_full_data.csv"))
| notebooks/1_extract_dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# # Robust covariance estimation and Mahalanobis distances relevance
#
#
# An example to show covariance estimation with the Mahalanobis
# distances on Gaussian distributed data.
#
# For Gaussian distributed data, the distance of an observation
# $x_i$ to the mode of the distribution can be computed using its
# Mahalanobis distance: $d_{(\mu,\Sigma)}(x_i)^2 = (x_i -
# \mu)'\Sigma^{-1}(x_i - \mu)$ where $\mu$ and $\Sigma$ are
# the location and the covariance of the underlying Gaussian
# distribution.
#
# In practice, $\mu$ and $\Sigma$ are replaced by some
# estimates. The usual covariance maximum likelihood estimate is very
# sensitive to the presence of outliers in the data set and therefor,
# the corresponding Mahalanobis distances are. One would better have to
# use a robust estimator of covariance to guarantee that the estimation is
# resistant to "erroneous" observations in the data set and that the
# associated Mahalanobis distances accurately reflect the true
# organisation of the observations.
#
# The Minimum Covariance Determinant estimator is a robust,
# high-breakdown point (i.e. it can be used to estimate the covariance
# matrix of highly contaminated datasets, up to
# $\frac{n_\text{samples}-n_\text{features}-1}{2}$ outliers)
# estimator of covariance. The idea is to find
# $\frac{n_\text{samples}+n_\text{features}+1}{2}$
# observations whose empirical covariance has the smallest determinant,
# yielding a "pure" subset of observations from which to compute
# standards estimates of location and covariance.
#
# The Minimum Covariance Determinant estimator (MCD) has been introduced
# by P.J.Rousseuw in [1].
#
# This example illustrates how the Mahalanobis distances are affected by
# outlying data: observations drawn from a contaminating distribution
# are not distinguishable from the observations coming from the real,
# Gaussian distribution that one may want to work with. Using MCD-based
# Mahalanobis distances, the two populations become
# distinguishable. Associated applications are outliers detection,
# observations ranking, clustering, ...
# For visualization purpose, the cubic root of the Mahalanobis distances
# are represented in the boxplot, as Wilson and Hilferty suggest [2]
#
# [1] <NAME>. Least median of squares regression. J. Am
# Stat Ass, 79:871, 1984.
# [2] <NAME>., & <NAME>. (1931). The distribution of chi-square.
# Proceedings of the National Academy of Sciences of the United States
# of America, 17, 684-688.
#
#
#
# +
print(__doc__)
import numpy as np
import matplotlib.pyplot as plt
from sklearn.covariance import EmpiricalCovariance, MinCovDet
n_samples = 125
n_outliers = 25
n_features = 2
# generate data
gen_cov = np.eye(n_features)
gen_cov[0, 0] = 2.
X = np.dot(np.random.randn(n_samples, n_features), gen_cov)
# add some outliers
outliers_cov = np.eye(n_features)
outliers_cov[np.arange(1, n_features), np.arange(1, n_features)] = 7.
X[-n_outliers:] = np.dot(np.random.randn(n_outliers, n_features), outliers_cov)
# fit a Minimum Covariance Determinant (MCD) robust estimator to data
robust_cov = MinCovDet().fit(X)
# compare estimators learnt from the full data set with true parameters
emp_cov = EmpiricalCovariance().fit(X)
# #############################################################################
# Display results
fig = plt.figure()
plt.subplots_adjust(hspace=-.1, wspace=.4, top=.95, bottom=.05)
# Show data set
subfig1 = plt.subplot(3, 1, 1)
inlier_plot = subfig1.scatter(X[:, 0], X[:, 1],
color='black', label='inliers')
outlier_plot = subfig1.scatter(X[:, 0][-n_outliers:], X[:, 1][-n_outliers:],
color='red', label='outliers')
subfig1.set_xlim(subfig1.get_xlim()[0], 11.)
subfig1.set_title("Mahalanobis distances of a contaminated data set:")
# Show contours of the distance functions
xx, yy = np.meshgrid(np.linspace(plt.xlim()[0], plt.xlim()[1], 100),
np.linspace(plt.ylim()[0], plt.ylim()[1], 100))
zz = np.c_[xx.ravel(), yy.ravel()]
mahal_emp_cov = emp_cov.mahalanobis(zz)
mahal_emp_cov = mahal_emp_cov.reshape(xx.shape)
emp_cov_contour = subfig1.contour(xx, yy, np.sqrt(mahal_emp_cov),
cmap=plt.cm.PuBu_r,
linestyles='dashed')
mahal_robust_cov = robust_cov.mahalanobis(zz)
mahal_robust_cov = mahal_robust_cov.reshape(xx.shape)
robust_contour = subfig1.contour(xx, yy, np.sqrt(mahal_robust_cov),
cmap=plt.cm.YlOrBr_r, linestyles='dotted')
subfig1.legend([emp_cov_contour.collections[1], robust_contour.collections[1],
inlier_plot, outlier_plot],
['MLE dist', 'robust dist', 'inliers', 'outliers'],
loc="upper right", borderaxespad=0)
plt.xticks(())
plt.yticks(())
# Plot the scores for each point
emp_mahal = emp_cov.mahalanobis(X - np.mean(X, 0)) ** (0.33)
subfig2 = plt.subplot(2, 2, 3)
subfig2.boxplot([emp_mahal[:-n_outliers], emp_mahal[-n_outliers:]], widths=.25)
subfig2.plot(1.26 * np.ones(n_samples - n_outliers),
emp_mahal[:-n_outliers], '+k', markeredgewidth=1)
subfig2.plot(2.26 * np.ones(n_outliers),
emp_mahal[-n_outliers:], '+k', markeredgewidth=1)
subfig2.axes.set_xticklabels(('inliers', 'outliers'), size=15)
subfig2.set_ylabel(r"$\sqrt[3]{\rm{(Mahal. dist.)}}$", size=16)
subfig2.set_title("1. from non-robust estimates\n(Maximum Likelihood)")
plt.yticks(())
robust_mahal = robust_cov.mahalanobis(X - robust_cov.location_) ** (0.33)
subfig3 = plt.subplot(2, 2, 4)
subfig3.boxplot([robust_mahal[:-n_outliers], robust_mahal[-n_outliers:]],
widths=.25)
subfig3.plot(1.26 * np.ones(n_samples - n_outliers),
robust_mahal[:-n_outliers], '+k', markeredgewidth=1)
subfig3.plot(2.26 * np.ones(n_outliers),
robust_mahal[-n_outliers:], '+k', markeredgewidth=1)
subfig3.axes.set_xticklabels(('inliers', 'outliers'), size=15)
subfig3.set_ylabel(r"$\sqrt[3]{\rm{(Mahal. dist.)}}$", size=16)
subfig3.set_title("2. from robust estimates\n(Minimum Covariance Determinant)")
plt.yticks(())
plt.show()
| scikit-learn-official-examples/covariance/plot_mahalanobis_distances.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.6.9 64-bit (''torch_rl'': venv)'
# name: python3
# ---
# +
# {'pkts_dropped': 0.0,
# 'pkts_transmitted': 75.0,
# 'timestamp': '1626810037874447104',
# 'obj': 'simulation_pedestrian1',
# 'pos_x': '11.417197',
# 'pos_y': '37.027515',
# 'pos_z': '7.4369965',
# 'orien_x': '-0.0',
# 'orien_y': '0.0',
# 'orien_z': '0.9999752',
# 'orien_w': '0.0070461035',
# 'linear_acc_x': '',
# 'linear_acc_y': '',
# 'linear_acc_z': '',
# 'linear_vel_x': '',
# 'linear_vel_y': '',
# 'linear_vel_z': '',
# 'angular_acc_x': '',
# 'angular_acc_y': '',
# 'angular_acc_z': '',
# 'angular_vel_x': '',
# 'angular_vel_y': '',
# 'angular_vel_z': '',
# 'pkts_buffered': 0.0,
# 'bit_rate': 4949598.859792932,
# 'chosen_ue': 'simulation_pedestrian1',
# 'packets': 14627.0,
# 'channel_mag': array(0.00890296)}
# +
import torch
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
from torch.utils.tensorboard import SummaryWriter
import tqdm
import math
import random
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
from collections import namedtuple, deque
from itertools import count
import copy
import gym
# set up matplotlib
is_ipython = 'inline' in matplotlib.get_backend()
if is_ipython:
from IPython import display
plt.ion()
# -
device = torch.device("cpu")
# # Gym Environment
import caviar_tools
from beamselect_env import BeamSelectionEnv
# ### Hyper Params
# +
reward_type = 'train' # 'test' or 'train'
epi = [0,9] #[start,end]
epi_val = [500,549]
gym_env = BeamSelectionEnv(epi, reward_type)
gym_env_val = BeamSelectionEnv(epi_val)
n_steps = caviar_tools.linecount(epi)
n_steps_val = caviar_tools.linecount(epi_val)
# -
n_steps
# +
# chan_mag = []
# pckts = []
# for i in tqdm.tqdm_notebook(range(n_steps)):
# next_state, _, _, _ = gym_env.step([np.random.randint(0,3), np.random.randint(0, 64)])
# pckts.append(next_state[7])
# chan_mag.append(next_state[8])
# +
# print(max(np.array(pckts).astype('float')), min(np.array(pckts).astype('float')))
# print(max(np.array(chan_mag).astype('float')), min(np.array(chan_mag).astype('float')))
# -
# Observation Space : X,Y,Z,pkts_dropped,pkts_transmitted,pkts_buffered,bit_rate
#
# Action Space : [3,64] -> [UE,Possible beams]
# ## Replay Memory
# +
Transition = namedtuple('Transition',
('state', 'action', 'next_state', 'reward'))
class ReplayMemory(object):
def __init__(self, capacity):
self.memory = deque([],maxlen=capacity)
def push(self, *args):
"""Save a transition"""
self.memory.append(Transition(*args))
def sample(self, batch_size):
return random.sample(self.memory, batch_size)
def __len__(self):
return len(self.memory)
# -
# ## DQN
# +
class DQN(nn.Module):
def __init__(self, inputs:int=7, outputs:int=64*3):
super().__init__()
self.linear = nn.Sequential(
self.create_linear(inputs,16),
self.create_linear(16, 32),
self.create_linear(32,64),
self.create_linear(64,256)
)
self.value_linear = self.create_linear(256,1)
self.action_linear = self.create_linear(256,outputs)
def create_linear(self,inp:int,out:int)-> nn.Module:
return nn.Sequential(
nn.Linear(inp,out),
nn.ELU()
# nn.BatchNorm1d(out)
)
def forward(self, x):
x = x.to(device)
x = self.linear(x)
value = self.value_linear(x)
adv = self.action_linear(x)
advAverage = torch.mean(adv, dim=1, keepdim=True)
out = value + adv - advAverage
return out
# -
# ## Hyperparams
# +
BATCH_SIZE = 128
GAMMA = 0.99
EPS_START = 0.9
EPS_END = 0.5
# It depends on overall number of steps, basic intitution is that
# once steps_done == EPS_DECAY then the probablity of choosing
# random action is 33%; considering EPS_END is zero
# As for ep = [0,10]; approx ep is 80k therefore exploration can be reduced to 33% around 50k
# Also because of this factor smoothed accuracy matters more for training then seeing the average
EPS_DECAY = n_steps*0.3
TARGET_UPDATE = 1000
VAL_STEP = 100000
Replay = 500000
EPOCH = 10
# -
# ## Action
# +
policy_net = DQN(13, 192).to(device)
target_net = DQN(13, 192).to(device)
target_net.load_state_dict(policy_net.state_dict())
target_net.eval()
optimizer = optim.Adam(policy_net.parameters())
memory = ReplayMemory(500000)
n_actions = 64*3
steps_done = 0
def select_action(state,steps_done:int=0,val:bool=False):
if val:
with torch.no_grad():
# t.max(1) will return largest column value of each row.
# second column on max result is index of where max element was
# found, so we pick action with the larger expected reward.
flattened_action = policy_net(state).max(dim = 1).indices
return torch.tensor([[flattened_action]], device=device, dtype=torch.long)
else:
sample = random.random()
eps_threshold = EPS_END + (EPS_START - EPS_END) * \
math.exp(-1. * steps_done / EPS_DECAY)
if sample > eps_threshold:
with torch.no_grad():
# t.max(1) will return largest column value of each row.
# second column on max result is index of where max element was
# found, so we pick action with the larger expected reward.
flattened_action = policy_net(state).max(dim = 1).indices
return torch.tensor([[flattened_action]], device=device, dtype=torch.long)
else:
flattened_action = random.randrange(n_actions)
return torch.tensor([[flattened_action]], device=device, dtype=torch.long)
# -
# ## Optimize Model
def optimize_model(memory:ReplayMemory):
if len(memory) < BATCH_SIZE:
return
transitions = memory.sample(BATCH_SIZE)
# Transpose the batch (see https://stackoverflow.com/a/19343/3343043 for
# detailed explanation). This converts batch-array of Transitions
# to Transition of batch-arrays.
batch = Transition(*zip(*transitions))
# Compute a mask of non-final states and concatenate the batch elements
# (a final state would've been the one after which simulation ended)
non_final_mask = torch.tensor(tuple(map(lambda s: s is not None,
batch.next_state)), device=device, dtype=torch.bool)
non_final_next_states = torch.cat([s for s in batch.next_state
if s is not None])
state_batch = torch.cat(batch.state)
action_batch = torch.cat(batch.action)
reward_batch = torch.cat(batch.reward)
# Compute Q(s_t, a) - the model computes Q(s_t), then we select the
# columns of actions taken. These are the actions which would've been taken
# for each batch state according to policy_net
state_action_values = policy_net(state_batch).gather(1, action_batch)
# Compute V(s_{t+1}) for all next states.
# Expected values of actions for non_final_next_states are computed based
# on the "older" target_net; selecting their best reward with max(1)[0].
# This is merged based on the mask, such that we'll have either the expected
# state value or 0 in case the state was final.
next_state_values = torch.zeros(BATCH_SIZE, device=device)
next_state_values[non_final_mask] = target_net(non_final_next_states).max(1)[0].detach()
# Compute the expected Q values
expected_state_action_values = (next_state_values * GAMMA) + reward_batch
# Compute Huber loss
criterion = nn.SmoothL1Loss()
loss = criterion(state_action_values, expected_state_action_values.unsqueeze(1))
# Optimize the model
optimizer.zero_grad()
loss.backward()
for param in policy_net.parameters():
param.grad.data.clamp_(-1, 1)
optimizer.step()
# ## Training loop
# +
# Tensorboard
log_dir = './mini_logs_with_test/train_10_epoch_10_reward_train_replay_500k'
writer = SummaryWriter(log_dir=log_dir)
# -
def val(train_step:int,gym_env_val:gym.Env,n_steps_val:int=n_steps_val,
writer:SummaryWriter=writer):
state = torch.zeros((1,13), dtype=torch.float32)
ovr_reward = 0
for i_episode in tqdm.tqdm_notebook(range(n_steps_val),desc='Validation'):
# Select and perform an action
action = select_action(state,val=True)
# Observe new state
next_state, reward, done, info = gym_env_val.step([action.item()//64, action.item()%64])
ovr_reward+=reward.item()
next_state = next_state.astype(np.float32).reshape(1, next_state.shape[0])
next_state = torch.tensor(next_state)
reward = torch.tensor([reward], device=device).float()
# Move to the next state
state = next_state
writer.add_scalar('Val Overall Reward',ovr_reward,train_step)
writer.add_scalar('Val Average Rewad',(ovr_reward/n_steps_val),train_step)
print(f'Validation Overall reward = {ovr_reward:.2f}. ' \
f' Validation Average Reward = {ovr_reward/n_steps_val:.4f}')
gym_env_val.close()
# +
# Initialize the environment and state
cnt = 0
gym_env_cp = copy.deepcopy(gym_env)
for i in range(0,EPOCH):
steps_done = 0
memory = ReplayMemory(Replay)
gym_env = copy.deepcopy(gym_env_cp)
state = torch.zeros((1,13), dtype=torch.float32)
ovr_reward = 0
for i_episode in tqdm.tqdm_notebook(range(n_steps),desc='Train'):
# Select and perform an action
action = select_action(state,steps_done)
steps_done+=1
# Observe new state
next_state, reward, done, info = gym_env.step([action.item()//64, action.item()%64])
ovr_reward+=reward.item()
next_state = next_state.astype(np.float32).reshape(1, next_state.shape[0])
next_state = torch.tensor(next_state)
reward = torch.tensor([reward], device=device).float()
writer.add_scalar('episode_reward',reward,cnt)
# print(next_state)
# print(state)
# Store the transition in memory
memory.push(state, action, next_state, reward)
# Move to the next state
state = next_state
# Perform one step of the optimization (on the policy network)
optimize_model(memory)
# Update the target network, copying all weights and biases in DQN
if i_episode % TARGET_UPDATE == 0:
target_net.load_state_dict(policy_net.state_dict())
if (i_episode) == 0:
val(cnt, copy.deepcopy(gym_env_val))
# cnt+=1
print(f'Overall Train reward after epoch {i} = {ovr_reward:.2f}. ' \
f'Average Reward = {ovr_reward/n_steps:.4f}')
gym_env.close()
writer.add_hparams(
{'BATCH_SIZE' : BATCH_SIZE,
'GAMMA' : GAMMA,
'EPS_START' : EPS_START,
'EPS_END' : EPS_END,
'EPS_DECAY' : EPS_DECAY,
'TARGET_UPDATE' :TARGET_UPDATE},
{
'Overall Reward':ovr_reward,
'Average Reward': ovr_reward/n_steps
}
)
gym_env_cp.close()
val(cnt, copy.deepcopy(gym_env_val))
| RL.ipynb |
# import dependencies
from pyspark.sql import SparkSession # needed to use spark DataFarmes
spark = SparkSession.builder.appName("recommender").getOrCreate()
from pyspark.ml.recommendation import ALS
from pyspark.ml.evaluation import RegressionEvaluator
# import data
df = spark.read.csv('/FileStore/tables/ratings.csv', inferSchema=True, header=True)
# explore data columns
df.printSchema()
# un-select timestamp column
df = df.select(['userId', 'movieId', 'rating'])
df.head(5)
df.show()
df.describe().show()
# create evaluation data
training, test = df.randomSplit([0.8,0.2])
# create model
als = ALS(maxIter=5, regParam=0.01, userCol='userId', itemCol='movieId', ratingCol='rating')
# train model
model = als.fit(training)
# get predictions
predictions = model.transform(test)
# look into predictions
predictions.describe().show()
# drop rows with non predictions
predictions = predictions.na.drop()
# again look into predictions
predictions.describe().show()
# evaluate
evaluator = RegressionEvaluator(metricName='rmse', labelCol='rating')
rmse = evaluator.evaluate(predictions)
rmse
| notebooks/0.1.als.recommender.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Get crystal structures Metadata from PDB
# - This notebook allows to download a pdb id associated metadata.
# - The metadata from the pdb ids belonging to the target protein are gathered into a dataframe named: `TABLA_MTDATA_{prot_name}_{number_of_structures}_crys.json`
from pathlib import Path
from glob import glob
import pandas as pd
import pickle
import json
import sys
import pypdb
from Bio import pairwise2, SeqIO
sys.path.insert(0, '../..')
from helper_modules.get_pdb_ids_from_uniport import *
from helper_modules.find_gaps import *
# ## Protein data
# ### Inputs
# This notebook requires to specify the following values
# - `prot_name`: the name of the protein.
# - `uniprot_id`: Uniprot Accession number.
# - `ref_struc_id`: PDB id of the reference structure.
# - `ref_ligand_name`: Three letters name of the cocrystalized ligand in the reference structure.
#
prot_name = 'egfr'
uniprot_id = 'P00533'
ref_struc_id = '7a2a'
ref_ligand_name = '7G9'
# ### Get the protein sequence
seq_prot = get_seq_from_uniprot(uniprot_id)
print(seq_prot)
print(f'\nThere are {len(seq_prot)} residues.')
# ### Get the pdbids of the downloaded structures
# +
# Get the pdbids and some metadata from uniprot
df_pdb_ids = pdb_ids_from_uniprot(uniprot_id)
# Get the reference structure and its sequence and residue positions
seq_cry, positions_cry = get_structure_sequence(ref_struc_id)
print(positions_cry[0], positions_cry[-1])
# Select only those entries that include the binding site
df_sel_pdbids = get_useful_pdbids(df_pdb_ids,
positions_cry,
thr_tol = (3,3))
# Get the list of input files
INPUT_DIR = f'./pdb_structures/pdb_chains'
pdbids_list = [i.split('/')[-1].split('_')[0] for i in sorted(glob(f'{INPUT_DIR}/*pdb'))]
n_pdb_id = len(pdbids_list)
# Update selection only those ids which files were downloaded
df_sel_pdbids = df_sel_pdbids[df_sel_pdbids.pdb_id.isin(pdbids_list)]
# -
df_sel_pdbids
# ### Get the available metadata
# +
mtdat_pkl_file = f"./MTDATA_{prot_name.upper()}_{n_pdb_id}_crys.pkl"
# If the dictionary already exists, load it!
if Path(mtdat_pkl_file).exists():
with open(mtdat_pkl_file, 'rb') as f:
prot_crys_mtd_dict = pickle.load(f)
else:
# Create an empty dict to fill it with the pdbs metadata
prot_crys_mtd_dict = {}
for i, (pdb_id, chain) in df_sel_pdbids[['pdb_id', 'chain']].iterrows():
print(i, '->', pdb_id)
# Get the metadata using pypdb and in-house functions
descrip_pdb = pypdb.describe_pdb(pdb_id)
cristal = pypdb.get_entity_info(pdb_id)
ligs_names = get_bounded_ligands(pdb_id)
num_ligs = len(ligs_names)
secuencia = get_pdb_sequence(pdb_id)
# Find number of gaps and coverage
seq_alg, coverage, gaps = get_gaps_and_coverage(
pdb_file = f'./pdb_structures/pdb_chains/{pdb_id}_A.pdb',
full_sequence = seq_prot,
chain = 'A') # Hardcoded as the chain was renamed in the previous notebook
identity = get_identity(seq_alg, seq_prot, gap_char = '-')
prot_crys_mtd_dict.update(
{pdb_id :
{'describe_pdb': descrip_pdb,
'pdb_info': cristal,
'identity': identity,
'num_ligs': num_ligs,
'name_ligs': ligs_names,
'seq': secuencia,
'seq_alg': seq_alg,
'coverage': coverage,
'gaps': gaps,
'chain': chain
}
})
# Save the dict as a json file
with open(mtdat_pkl_file, 'wb') as file:
pickle.dump(prot_crys_mtd_dict, file)
# -
# ## Create the final Metadata table
# +
def get_data_rows(pdb_entry: dict) -> pd.Series:
'''
This function takes a pdb_entry dictionary,
nested in `prot_crys_mtd_dict` created in the previous cell,
and returns a pandas series containing the entry metadata
'''
e = pdb_entry
d = e['describe_pdb']
l = e
dic = {"PDB_ID" : d['rcsb_id'].lower(),
"Title" : d['citation'][0]['title'].lower(),
"Entities" : d['pdbx_vrpt_summary']['protein_dnarnaentities'],
"ChainID" : e['chain'],
"Identity" : round(e['identity'], 3),
"Coverage" : round(e['coverage'], 3),
"NumGaps" : e['gaps']['num_gaps'],
"GapLen" : e['gaps']['gap_lengths'],
"GapPos" : e['gaps']['gap_list'],
"Resolution" : d['pdbx_vrpt_summary']['pdbresolution'],
"Date" : d['rcsb_accession_info']['initial_release_date'].split('T')[0],
"NumLigs" : e['num_ligs'],
"NameLigs" : e['name_ligs']
}
entry_series = pd.Series(dic)
return entry_series
def get_mtd_table(dic: dict):
'''
This function is designed to take the
`prot_crys_mtd_dict` dictionary
and extract a pandas series for each pdb entry
'''
df = pd.DataFrame()
for pdb_id in dic.keys():
row = get_data_rows(dic[pdb_id])
df = df.append(row, ignore_index=True)
return df
# +
# Now we create the dataframe
df_prot = get_mtd_table(prot_crys_mtd_dict).sort_values("PDB_ID").set_index("PDB_ID")
# Reorder the columns
df_prot = df_prot[['Title','Date', 'Entities', 'ChainID','Resolution',
'Identity', 'Coverage', 'NumGaps', 'GapLen', 'GapPos',
'NumLigs', 'NameLigs']]
df_prot
# -
# ## Save the metadata Dataframe to a file
n_pdb_id_to_use = len(df_prot)
dataframe_file = f"./TABLA_MTDATA_{prot_name.upper()}_{n_pdb_id_to_use}_crys.json"
if not Path(dataframe_file).exists:
df_prot.to_json(dataframe_file)
# ## Data exploration
from wordcloud import WordCloud, STOPWORDS, ImageColorGenerator
import matplotlib.pyplot as plt
import seaborn as sns
sns.set(font_scale = 1.2, style = 'white')
# +
text = " ".join(titulo for titulo in df_prot.Title)
text = text.replace('-', ' ').upper()
stopwords = set(STOPWORDS)
stopwords.update([ "structure", "ray", "crystal", "yl"])
# Create and generate a word cloud image:
wordcloud = WordCloud(stopwords = stopwords,
max_font_size = 100,
collocations = False,
min_font_size = 4,
contour_width = 100,
colormap = "Spectral",
random_state = 15,
max_words = 100,
width=800, height=400,
background_color = "black").generate(text)
plt.figure(figsize = (16, 7))
plt.imshow(wordcloud, interpolation = "bilinear")
plt.axis("off")
plt.show()
# -
def plot_property(prop, color = 'red', title = ''):
df_prot[prop] = df_prot[prop].astype(float)
fig, ax = plt.subplots(figsize = (8, 5))
sns.histplot(df_prot[prop] ,
color = color,
kde = True,
label = prop,
ax = ax)
plt.legend()
plt.title(title)
plt.show()
df_prot.Date = pd.to_datetime(df_prot.Date)
df_prot_dates = df_prot.set_index('Date')
df_prot_dates.tail()
fig, ax = plt.subplots(figsize = (8, 5))
ax = df_prot.groupby(df_prot.Date.dt.year)['Date'].count().plot(kind='bar')
ax.set(ylabel = 'Count', title = 'Number of structures published by year')
prop = 'Resolution'
plot_property(prop, title = f'Number of structures by {prop}')
prop = 'Coverage'
plot_property(prop, color = 'teal',
title = f'Number of structures by {prop}')
prop = 'NumLigs'
plot_property(prop, color = 'orange',
title = f'Number of structures by {prop}')
| egfr/1_Download_and_prepare_protein_ensembles/2_Get_PDB_structures_metadata.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # 成為初級資料分析師 | R 程式設計與資料科學應用
#
# > 程式封裝:自訂函數
#
# ## 郭耀仁
# + [markdown] slideshow={"slide_type": "subslide"}
# > The way R works is pretty straightforward, you apply functions to objects.
# >
# > <NAME>
# + [markdown] slideshow={"slide_type": "subslide"}
# ## 大綱
#
# - 關於函數
# - 常用的 R 內建函數
# - 自訂函數
# - 全域(Global)與區域(Local)
# - 向量化函數
# - 遞迴(Recursion)
# + [markdown] slideshow={"slide_type": "slide"}
# ## 關於函數
# + [markdown] slideshow={"slide_type": "subslide"}
# 
# + [markdown] slideshow={"slide_type": "subslide"}
# ## 函數由四個元件組合而成
#
# - 輸入 INPUTS
# - 參數 ARGUMENTS
# - 主體 BODY
# - 輸出 OUTPUTS
# + [markdown] slideshow={"slide_type": "slide"}
# ## 常用的 R 內建函數
# + [markdown] slideshow={"slide_type": "subslide"}
# ## 內建數值函數
# + [markdown] slideshow={"slide_type": "subslide"}
# ## `abs()`
#
# 取絕對值
# -
abs(-5566)
abs(5566)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## `sqrt()`
#
# 平方根
# -
sqrt(2)
sqrt(3)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## `ceiling()`
#
# 無條件進位
# -
ceiling(sqrt(2))
ceiling(sqrt(3))
# + [markdown] slideshow={"slide_type": "subslide"}
# ## `floor()`
#
# 無條件捨去
# -
floor(sqrt(2))
floor(sqrt(3))
# + [markdown] slideshow={"slide_type": "subslide"}
# ## `round()`
#
# 四捨五入
# -
round(sqrt(2))
round(sqrt(3))
# + [markdown] slideshow={"slide_type": "subslide"}
# ## `exp()`
#
# 指數函數
#
# $$exp(x) = e^x \text{ , where } e = 2.71828...
# $$
# -
exp(1)
exp(2)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## `log()`
#
# 以 $e$ 為底的對數函數
# -
e = exp(1)
ee = exp(2)
log(e)
log(ee)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## `log10()`
#
# 以 10 為底的對數函數
# -
log10(10)
log10(10**2)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## 內建描述性統計函數
# + [markdown] slideshow={"slide_type": "subslide"}
# ## `mean()`
#
# 平均數
# -
mean(1:10)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## `sd()`
#
# 標準差
# -
sd(1:10)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## `median()`
#
# 中位數
# -
median(1:9)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## `range()`
#
# 最大值與最小值
# -
range(11:100)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## `sum()`
#
# 加總
# -
sum(1:100)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## `max()`
#
# 最大值
# -
max(11:100)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## `min()`
#
# 最小值
# -
min(11:100)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## 內建文字處理函數
# + [markdown] slideshow={"slide_type": "subslide"}
# ## `unique()`
#
# 取獨一值
# -
cities <- c("New York", "Boston", "Tokyo", "Kyoto", "Taipei")
countries <- c("United States", "United States", "Japan", "Japan", "Taiwan")
unique(cities)
unique(countries)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## `toupper()`
#
# 轉換為大寫
# -
toupper("Luke, use the Force!")
# + [markdown] slideshow={"slide_type": "subslide"}
# ## `tolower()`
#
# 轉換為小寫
# -
tolower("Luke, use the Force!")
# + [markdown] slideshow={"slide_type": "subslide"}
# ## `substr()`
#
# 擷取部分字串
# -
luke <- "Luke, use the Force!"
substr(luke, start = 1, stop = 4)
substr(luke, start = 11, stop = nchar(luke))
# + [markdown] slideshow={"slide_type": "subslide"}
# ## `grep()`
#
# 回傳指定特徵有出現的索引值
# -
avengers <- c("The Avengers", "Avengers: Age of Ultron", "Avengers: Infinity War", "Avengers: Endgame")
grep(pattern = "Avengers", avengers)
grep(pattern = "Endgame", avengers)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## `sub()`
#
# 取代指定特徵
# -
skywalker <- "<NAME>"
sub(pattern = "Anakin", replacement = "Luke", skywalker)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## `strsplit()`
#
# 依據指定特徵分割文字
# -
avengers <- c("The Avengers", "Avengers: Age of Ultron", "Avengers: Infinity War", "Avengers: Endgame")
strsplit(avengers, split = " ")
strsplit(avengers, split = ":")
# + [markdown] slideshow={"slide_type": "subslide"}
# ## `paste()` 與 `paste0()`
#
# 連接文字
# -
paste("Avengers:", "Endgame")
paste0("Avengers:", "Endgame")
# + [markdown] slideshow={"slide_type": "subslide"}
# ## `trimws()`
#
# 消除前(leading)或後(trailing)的空白
# -
luke <- " Luke, use the Force! "
trimws(luke)
trimws(luke, which = "left")
trimws(luke, which = "right")
# + [markdown] slideshow={"slide_type": "slide"}
# ## 自訂函數
# + [markdown] slideshow={"slide_type": "subslide"}
# ## 在自訂函數時考慮 6 個元件
#
# - 函數名稱(命名風格與物件命名風格相同,儘量使用動詞)
# - 輸入的命名與設計
# - 參數的命名、設計與預設值
# - 函數主體
# - 輸出的命名與設計
# - 使用保留字 `function` 與 `return()`
# + [markdown] slideshow={"slide_type": "subslide"}
# ## 自訂函數的 Code Block
#
# 
# + [markdown] slideshow={"slide_type": "subslide"}
# ```r
# FUNCTION_NAME <- function(INPUTS, ARGUMENTS, ...) {
# # BODY
# return(OUTPUTS)
# }
# ```
# + [markdown] slideshow={"slide_type": "subslide"}
# ## 單一輸入的函數
#
# 將攝氏溫度轉換為華氏溫度的函數 `celsius_to_fahrenheit()`
#
# $$Fahrenheit_{(°F)} = Celsius_{(°C)} \times \frac{9}{5} + 32$$
# + slideshow={"slide_type": "-"}
celsius_to_fahrenheit <- function(x) {
return(x*9/5 + 32)
}
celsius_to_fahrenheit(20)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## 隨堂練習:將公里轉換為英里的函數 `km_to_mile()`
#
# $$Miles = Kilometers \times 0.62137$$
# + slideshow={"slide_type": "skip"}
km_to_mile <- function(x) {
return(x*0.62137)
}
# + slideshow={"slide_type": "fragment"}
km_to_mile(21.095)
km_to_mile(42.195)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## 隨堂練習:判斷輸入 x 是否為質數的函數 `is_prime()`
# + slideshow={"slide_type": "skip"}
is_prime <- function(x) {
divisors_cnt <- 0
for (i in 1:x) {
if (x %% i == 0) {
divisors_cnt <- divisors_cnt + 1
}
}
return(divisors_cnt == 2)
}
# + slideshow={"slide_type": "fragment"}
is_prime(1)
is_prime(2)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## 兩個以上輸入的函數
# + slideshow={"slide_type": "-"}
get_bmi <- function(height, weight) {
return(weight / (height*0.01)**2)
}
get_bmi(191, 91)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## 隨堂練習:回傳介於 x 與 y 之間的質數個數(包含 x 與 y 如果他們也是質數)的函數 `count_primes()`
# + slideshow={"slide_type": "skip"}
count_primes <- function(x, y) {
primes_cnt <- 0
for (i in x:y) {
if (is_prime(i)) {
primes_cnt <- primes_cnt + 1
}
}
return(primes_cnt)
}
# + slideshow={"slide_type": "fragment"}
count_primes(1, 5)
count_primes(9, 19)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## 參數有預設值的函數
# + slideshow={"slide_type": "-"}
temperature_converter <- function(x, to_fahrenheit = TRUE) {
if (to_fahrenheit) {
return(x*9/5 + 32)
} else {
return((x - 32)*5/9)
}
}
temperature_converter(20)
temperature_converter(68, to_fahrenheit = FALSE)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## 隨堂練習:計算圓面積或圓周長的函數 `circle_calculator()`(預設計算圓面積)
# + slideshow={"slide_type": "skip"}
circle_calculator <- function(r, is_area = TRUE) {
if (is_area) {
return(pi*r**2)
} else {
return(2*pi*r)
}
}
# + slideshow={"slide_type": "fragment"}
circle_calculator(3)
circle_calculator(3, is_area = FALSE)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## 多個輸出的函數
# + slideshow={"slide_type": "-"}
get_bmi_and_label <- function(height, weight) {
bmi <- weight / (height/100)**2
if (bmi > 30) {
label <- "Obese"
} else if (bmi < 18.5) {
label <- "Underweight"
} else if (bmi > 25) {
label <- "Overweight"
} else {
label <- "Normal weight"
}
bmi_and_label <- list(
bmi = bmi,
bmi_label = label
)
return(bmi_and_label)
}
# -
get_bmi_and_label(216, 147) # <NAME>
get_bmi_and_label(203, 113) # LeBron James
get_bmi_and_label(191, 82) # <NAME>
get_bmi_and_label(231, 91) # <NAME>
# + [markdown] slideshow={"slide_type": "subslide"}
# ## 隨堂練習:回傳介於 x 與 y 之間的質數個數(包含 x 與 y 如果他們也是質數)以及質數明細的函數 `get_primes_and_counts()`
# + slideshow={"slide_type": "skip"}
get_primes_and_counts <- function(x, y) {
primes <- c()
for (i in x:y) {
if (is_prime(i)) {
primes <- c(primes, i)
}
}
primes_and_counts <- list(
primes = primes,
prime_counts = length(primes)
)
return(primes_and_counts)
}
# + slideshow={"slide_type": "fragment"}
get_primes_and_counts(1, 5)
get_primes_and_counts(9, 19)
# + [markdown] slideshow={"slide_type": "slide"}
# ## 全域(Global)與區域(Local)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## 什麼是全域與區域?
#
# - 在函數的 Code Block 以外所建立的物件屬於全域
# - 在函數的 Code Block 中建立的物件屬於區域
# + [markdown] slideshow={"slide_type": "subslide"}
# ## 全域物件與區域物件的差別?
#
# - 區域物件**僅可**在區域中使用
# - 全域物件可以在全域以及區域中使用
# + slideshow={"slide_type": "subslide"}
# 區域物件僅可在區域中使用
get_sqrt <- function(x) {
sqrt_x <- x**0.5
return(sqrt_x)
}
get_sqrt(2)
sqrt_x # Local object cannot be accessed in global
# + slideshow={"slide_type": "subslide"}
# 全域物件可以在全域以及區域中使用
x <- 2
sqrt_x <- x**0.5
sqrt_x # Global object can be accessed in global, of course
# + slideshow={"slide_type": "subslide"}
# 全域物件可以在全域以及區域中使用
x <- 2
sqrt_x <- x**0.5
get_sqrt <- function() {
return(sqrt_x) # Global object can be accessed in local
}
get_sqrt()
# + [markdown] slideshow={"slide_type": "slide"}
# ## 向量化函數
# + [markdown] slideshow={"slide_type": "subslide"}
# ## 什麼是向量化函數?
#
# 利用 A 函數將 B 函數**向量化**至某個資料結構的方法,其中 A 函數是特定的函數型函數(Functional Functions)
# + slideshow={"slide_type": "subslide"}
# 將一個 list 中的每個數字都平方
my_list <- list(
11,
12,
13,
14,
15
)
my_list
# + [markdown] slideshow={"slide_type": "subslide"}
# ## 老實說,我們會有個衝動這樣寫
# -
my_list**2
# + [markdown] slideshow={"slide_type": "subslide"}
# ## 注意這是一個 `list` 而不是向量
# -
for (i in my_list) {
print(i**2)
}
# + [markdown] slideshow={"slide_type": "subslide"}
# ## 使用 `lapply()` 函數將 `get_squared()` 函數向量化至 `my_list` 之上
# +
get_squared <- function(x) {
return(x**2)
}
lapply(my_list, FUN = get_squared)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## 搭配匿名函數(Anonymous Function)將平方運算向量化至 `my_list` 之上
# -
lapply(my_list, FUN = function(x) return(x**2))
# + [markdown] slideshow={"slide_type": "subslide"}
# ## R 常用的函數型函數(functional functions)
#
# - `lapply()`
# - `sapply()`
# - `apply()`
# - `mapply()`
# + [markdown] slideshow={"slide_type": "subslide"}
# ## `sapply()`
#
# - Simplified apply
# - 回傳向量而非 `list`
# + slideshow={"slide_type": "-"}
sapply(my_list, FUN = function(x) return(x**2))
# + [markdown] slideshow={"slide_type": "subslide"}
# ## `apply()`
#
# 將函數向量化至二維的資料結構(`matrix` 與 `data.frame`)
# -
my_matrix <- matrix(1:12, nrow = 2)
my_matrix
apply(my_matrix, MARGIN = 1, FUN = sum)
apply(my_matrix, MARGIN = 2, FUN = sum)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## `mapply()`
#
# 將具有多個輸入的函數向量化
# -
weights <- list(91, 82, 113, 147)
heights <- list(231, 191, 203, 216)
bmis <- mapply(FUN = function(h, w) return(w/(h*0.01)**2), heights, weights)
bmis
# + [markdown] slideshow={"slide_type": "subslide"}
# ## 隨堂練習:使用向量化函數將 5 個球員的姓氏(last name)擷取出來並轉換成大寫
# -
fav_players <- list("<NAME>", "<NAME>", "<NAME>", "<NAME>", "<NAME>")
# + slideshow={"slide_type": "skip"}
get_uppercased_lastname <- function(x) {
last_name <- strsplit(x, split = " ")[[1]][2]
return(toupper(last_name))
}
ans <- lapply(fav_players, FUN = get_uppercased_lastname)
# + slideshow={"slide_type": "fragment"}
ans
# + [markdown] slideshow={"slide_type": "slide"}
# ## 遞迴(Recursion)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## 什麼是遞迴(Recursion)?
#
# 在一個函數中呼叫函數本身的技法
# + [markdown] slideshow={"slide_type": "subslide"}
# 
#
# Source: <https://twitter.com/ProgrammersMeme/status/1147050956821008384>
# + [markdown] slideshow={"slide_type": "subslide"}
# ## 階乘(factorial)的計算
#
# $$n! = 1 \times 2 \times 3 \times ... \times n$$
# -
factorial <- function(n) {
if (n == 1) {
return(n)
} else {
return(n * factorial(n-1))
}
}
factorial(1)
factorial(2)
factorial(3)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## 隨堂練習:建立 fibonacci 數列
#
# $$
# F_0 = 0, F_1 = 1 \\
# F_n = F_{n-1} + F_{n-2} \text{ , For } n > 1
# $$
# + slideshow={"slide_type": "skip"}
fib <- function(n) {
if (n == 1) {
return(0)
} else if (n == 2) {
return(1)
} else {
return(fib(n-1) + fib(n-2))
}
}
fibonacci <- function(N) {
fib_seq <- c()
for (i in 1:N) {
fib_seq <- c(fib_seq, fib(i))
}
return(fib_seq)
}
# + slideshow={"slide_type": "fragment"}
fibonacci(1)
fibonacci(2)
fibonacci(20)
| notebooks/08-functions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
plt.style.use('seaborn')
# %matplotlib inline
import json
import requests
# -
from jupyterdataworkflow.data import get_Fremont_data
data = get_Fremont_data()
data.head()
data.plot()
data.resample('W').sum().plot()
ax = data.resample('D').sum().rolling(365).sum().plot()
ax.set_ylim(0, None)
data.groupby(data.index.time).mean().plot()
pivoted = data.pivot_table('Total', index = data.index.time, columns = data.index.date)
pivoted.iloc[:5, :5]
pivoted.plot(legend=False, alpha = 0.01)
| Data analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] heading_collapsed=true
# # 0.0 - IMPORTS
# + hidden=true
import math
import random
import datetime
import warnings
import inflection
import numpy as np
import pandas as pd
import seaborn as sns
import xgboost as xgb
from IPython.core.display import HTML
from IPython.display import Image
from boruta import BorutaPy
from scipy import stats as ss
from matplotlib import pyplot as plt
from sklearn.ensemble import RandomForestRegressor
from sklearn.linear_model import LinearRegression, Lasso
from sklearn.preprocessing import MinMaxScaler, LabelEncoder, RobustScaler
from sklearn.metrics import mean_absolute_error, mean_absolute_percentage_error, mean_squared_error
warnings.filterwarnings( 'ignore' )
# + [markdown] heading_collapsed=true hidden=true
# ## 0.1 Helper Functions
# + hidden=true
def cross_validation(x_training, kfold, model_name, model, verbose=False ):
mae_list= []
mape_list = []
rmse_list = []
for k in reversed( range(1, kfold+1 ) ):
if verbose:
print('\nKFold Number: {}'.format(k) )
# start and end date for validation
validation_start_date = x_training['date'].max() - datetime.timedelta( days=k*6*7 )
validadtion_end_date = x_training['date'].max() - datetime.timedelta( days=(k-1)*6*7 )
# filtering dataset
training = x_training[x_training['date'] < validation_start_date]
validation = x_training[(x_training['date'] >= validation_start_date) & (x_training['date'] <= validadtion_end_date)]
# training and validation datasel
# training
xtraining = training.drop( ['date','sales'], axis=1 )
ytraining = training['sales']
# validation
xvalidation = validation.drop( ['date','sales'], axis=1 )
yvalidation = validation['sales']
# model
m = model.fit( xtraining, ytraining )
#prediction
yhat = m.predict( xvalidation )
#performance
m_result = ml_error( model_name, np.expm1( yvalidation), np.expm1(yhat) )
# store performance of each kfold interation
mae_list.append( m_result['MAE'] )
mape_list.append( m_result['MAPE'] )
rmse_list.append( m_result['RMSE'] )
return pd.DataFrame( {'Model Name': model_name,
'MAE CV': np.round( np.mean( mae_list), 2 ).astype( str ) + ' +/- ' + np.round( np.std( mae_list ), 2 ).astype( str ),
'MAPE CV': np.round( np.mean( mape_list), 2 ).astype( str ) + ' +/- ' + np.round( np.std( mape_list ), 2 ).astype( str ),
'RMSE CV': np.round( np.mean( rmse_list), 2 ).astype( str ) + ' +/- ' + np.round( np.std( rmse_list ), 2 ).astype( str ) }, index=[0] )
def ml_error( model_name, y, yhat ):
mae = mean_absolute_error( y, yhat )
mape = mean_absolute_percentage_error( y, yhat )
rmse = np.sqrt( mean_squared_error( y, yhat ) )
return pd.DataFrame( {'Model Name': model_name,
'MAE': mae,
'MAPE': mape,
'RMSE': rmse}, index=[0])
def mean_percentage_error ( y, yhat ):
return np.mean( ( y- yhat ) /y )
def cramer_v( x, y):
cm = pd.crosstab( x, y).values
n = cm.sum()
r, k = cm.shape
chi2 = ss.chi2_contingency( cm )[0]
chi2corr = max( 0, chi2 - (k-1)*(r-1)/(n-1) )
kcorr = k - (k-1)**2/(n-1)
rcorr = r - (r-1)**2/(n-1)
return np.sqrt( (chi2corr/n) / ( min( kcorr-1, rcorr-1 ) ) )
def jupyter_settings():
# %matplotlib inline
# %pylab inline
plt.style.use( 'bmh' )
plt.rcParams['figure.figsize'] = [25,12]
plt.rcParams['font.size'] = 24
display( HTML( '<style>.container {width:100% !important; }</style>') )
pd.options.display.max_columns = None
pd.options.display.max_rows = None
pd.set_option( 'display.expand_frame_repr', False )
sns.set()
# + hidden=true
jupyter_settings()
# + [markdown] hidden=true
# ## 0.2 Loading Data
# + hidden=true
df_sales_raw = pd.read_csv( 'data_csv/train.csv', low_memory= False)
df_store_raw = pd.read_csv( 'data_csv/store.csv', low_memory= False)
# Merge
df_raw = pd.merge( df_sales_raw, df_store_raw, how='left', on='Store' )
# + [markdown] heading_collapsed=true
# # 1.0 PASSO 1 - DATA DESCRIPTION
# + [markdown] heading_collapsed=true hidden=true
# ## 1.1 Rename Columns.
# + hidden=true
df1 = df_raw.copy()
# + hidden=true
cols_old = ['Store', 'DayOfWeek', 'Date', 'Sales', 'Customers', 'Open', 'Promo', 'StateHoliday', 'SchoolHoliday', 'StoreType',
'Assortment', 'CompetitionDistance', 'CompetitionOpenSinceMonth', 'CompetitionOpenSinceYear', 'Promo2',
'Promo2SinceWeek','Promo2SinceYear', 'PromoInterval']
snakecase = lambda x: inflection.underscore( x )
cols_new = list( map( snakecase, cols_old ) )
# rename
df1.columns = cols_new
# + [markdown] heading_collapsed=true hidden=true
# ## 1.2 Data Dimensions
# + hidden=true
print( 'Number of Rows: {}'.format( df1.shape[0] ) )
print( 'Number of Cols: {}'.format( df1.shape[1] ) )
# + [markdown] heading_collapsed=true hidden=true
# ## 1.3 Data Types
# + hidden=true
df1['date'] = pd.to_datetime (df1['date'] )
# + [markdown] hidden=true
# ## 1.4 Check NA
# + [markdown] heading_collapsed=true hidden=true
# ## 1.5 Fillout NA
# + hidden=true
# Competition Distance
df1['competition_distance'] = df1['competition_distance'].apply( lambda x: 200000.0 if math.isnan( x ) else x )
# Competition Open Since Month
df1['competition_open_since_month'] = df1.apply( lambda x: x['date'].month if math.isnan( x['competition_open_since_month'] ) else x['competition_open_since_month'], axis=1 )
# Competition Open Since Year
df1['competition_open_since_year'] = df1.apply( lambda x: x['date'].year if math.isnan( x['competition_open_since_year'] ) else x['competition_open_since_year'], axis=1 )
# Promo2 Since Week
df1['promo2_since_week'] = df1.apply( lambda x: x['date'].week if math.isnan( x['promo2_since_week'] ) else x['promo2_since_week'], axis=1 )
# Promo2 Since Year
df1['promo2_since_year'] = df1.apply( lambda x: x['date'].year if math.isnan( x['promo2_since_year'] ) else x['promo2_since_year'], axis=1 )
# Promo Interval
month_map = {1: 'Jan', 2: 'Fev', 3: 'Mar', 4: 'Apr', 5: 'May', 6: 'Jun', 7: 'Jul', 8: 'Aug', 9: 'Sept', 10: 'Oct', 11: 'Nov', 12: 'Dec'}
df1['promo_interval'].fillna(0, inplace=True)
df1['month_map'] = df1['date'].dt.month.map( month_map )
df1['is_promo'] = df1[['promo_interval', 'month_map']].apply( lambda x: 0 if x['promo_interval'] == 0 else 1 if x['month_map'] in x['promo_interval'].split( ',' ) else 0, axis=1 )
# + [markdown] heading_collapsed=true hidden=true
# ## 1.6 Change Types
# + hidden=true
df1['competition_open_since_month'] = df1['competition_open_since_month'].astype( int )
df1['competition_open_since_year'] = df1['competition_open_since_year'].astype( int )
df1['promo2_since_week'] = df1['promo2_since_week'].astype( int )
df1['promo2_since_year'] = df1['promo2_since_year'].astype( int )
# + [markdown] heading_collapsed=true hidden=true
# ## 1.7 Descriptive Statistical
# + [markdown] hidden=true
# ### 1.7.1 Numerical Categorics
# + hidden=true
num_attributes = df1.select_dtypes( include=['int32', 'int64', 'float64'] )
cat_attributes = df1.select_dtypes( exclude=['int32', 'int64', 'float64', 'datetime64[ns]'] )
# + hidden=true
# Central Tendency - mean, median
ct1 = pd.DataFrame( num_attributes.apply( np.mean ) ).T
ct2 = pd.DataFrame( num_attributes.apply( np.median ) ).T
# Dispersion - std, min, max, range, skew, kurtosis
d1 = pd.DataFrame( num_attributes.apply( np.std ) ).T
d2 = pd.DataFrame( num_attributes.apply( min ) ).T
d3 = pd.DataFrame( num_attributes.apply( max ) ).T
d4 = pd.DataFrame( num_attributes.apply( lambda x: x.max() - x.min() ) ).T
d5 = pd.DataFrame( num_attributes.apply( lambda x: x.skew() ) ).T
d6 = pd.DataFrame( num_attributes.apply( lambda x: x.kurtosis() ) ).T
# Concatenar
m = pd.concat( [ d2, d3, d4, ct1, ct2, d1, d5, d6 ] ).T.reset_index()
m.columns = ['attributes', 'min', 'max', 'range', 'mean', 'median', 'std', 'skew', 'kurtosis']
m
# + hidden=true
sns.displot( df1['competition_distance'] )
# + [markdown] hidden=true
# ### 1.7.2 Categorical Attributes
# + hidden=true
cat_attributes.apply( lambda x: x.unique().shape[0] )
# + hidden=true
aux1 = df1[(df1['state_holiday'] != '0') & (df1['sales'] > 0)]
plt.subplot(1,3,1)
sns.boxplot( x='state_holiday', y='sales', data=aux1 )
plt.subplot(1,3,2)
sns.boxplot( x='store_type', y='sales', data=aux1 )
plt.subplot(1,3,3)
sns.boxplot( x='assortment', y='sales', data=aux1 )
# + [markdown] heading_collapsed=true
# # 2.0 PASSO 2 - FEATURE ENGINEERING
# + hidden=true
df2 = df1.copy()
# + [markdown] hidden=true
# ## 2.1 Mapa mental de Hipoteses
# + hidden=true
Image('img/mindmaphypothesis.png')
# + [markdown] heading_collapsed=true hidden=true
# ## 2.2 Criação das Hipoteses
# + [markdown] hidden=true
# ### 2.2.1 Hipoteses Loja
# + [markdown] hidden=true
# **1.** Lojas com número maior de funcionários deveriam vender mais.
#
# **2.** Lojas com maior capacidade de estoque deveriam vender mais.
#
# **3.** Lojas com maior porte deveriam vender mais.
#
# **4.** Lojas com maior sortimentos deveriam vender mais.
#
# **5.** Lojas com competidores mais próximos deveriam vender menos.
#
# **6.** Lojas com competidores à mais tempo deveriam vendem mais.
# + [markdown] hidden=true
# ### 2.2.2 Hipoteses Produto
# + [markdown] hidden=true
# **1.** Lojas que investem mais em Marketing deveriam vender mais.
#
# **2.** Lojas com maior exposição de produto deveriam vender mais.
#
# **3.** Lojas com produtos com preço menor deveriam vender mais.
#
# **5.** Lojas com promoções mais agressivas ( descontos maiores ), deveriam vender mais.
#
# **6.** Lojas com promoções ativas por mais tempo deveriam vender mais.
#
# **7.** Lojas com mais dias de promoção deveriam vender mais.
#
# **8.** Lojas com mais promoções consecutivas deveriam vender mais.
# + [markdown] hidden=true
# ### 2.2.3 Hipoteses Tempo
# + [markdown] hidden=true
# **1.** Lojas abertas durante o feriado de Natal deveriam vender mais.
#
# **2.** Lojas deveriam vender mais ao longo dos anos.
#
# **3.** Lojas deveriam vender mais no segundo semestre do ano.
#
# **4.** Lojas deveriam vender mais depois do dia 10 de cada mês.
#
# **5.** Lojas deveriam vender menos aos finais de semana.
#
# **6.** Lojas deveriam vender menos durante os feriados escolares.
#
# + [markdown] hidden=true
# ### 2.2.4 Lista Final de Hipoteses
# + [markdown] hidden=true
# **1.** Lojas com maior sortimentos deveriam vender mais.
#
# **2.** Lojas com competidores mais próximos deveriam vender menos.
#
# **3.** Lojas com competidores à mais tempo deveriam vendem mais.
#
# **4.** Lojas com promoções ativas por mais tempo deveriam vender mais.
#
# **5.** Lojas com mais dias de promoção deveriam vender mais.
#
# **7.** Lojas com mais promoções consecutivas deveriam vender mais.
#
# **8.** Lojas abertas durante o feriado de Natal deveriam vender mais.
#
# **9.** Lojas deveriam vender mais ao longo dos anos.
#
# **10.** Lojas deveriam vender mais no segundo semestre do ano.
#
# **11.** Lojas deveriam vender mais depois do dia 10 de cada mês.
#
# **12.** Lojas deveriam vender menos aos finais de semana.
#
# **13.** Lojas deveriam vender menos durante os feriados escolares.
# + [markdown] hidden=true
# ## 2.3 Feature Engineering
# + hidden=true
# year
df2['year'] = df2['date'].dt.year
# month
df2['month'] = df2['date'].dt.month
# day
df2['day'] = df2['date'].dt.day
# week of year
df2['week_of_year'] = df2['date'].dt.weekofyear
# year week
df2['year_week'] = df2['date'].dt.strftime( '%Y-%W' )
# competition since
df2['competition_since'] = df2.apply( lambda x: datetime.datetime( year=x['competition_open_since_year'], month=x['competition_open_since_month'], day=1 ), axis=1 )
df2['competition_time_month'] = ( ( df2['date'] - df2['competition_since'] ) / 30 ).apply( lambda x: x.days ).astype( int )
# promo since
df2['promo_since'] = df2['promo2_since_year'].astype( str ) + '-' + df2['promo2_since_week'].astype( str )
df2['promo_since'] = df2['promo_since'].apply( lambda x: datetime.datetime.strptime( x + '-1', '%Y-%W-%w' ) - datetime.timedelta( days=7 ) )
df2['promo_time_week'] = ( ( df2['date'] - df2['promo_since'] ) /7 ).apply( lambda x: x.days ).astype( int )
# assortment
df2['assortment'] = df2['assortment'].apply(lambda x: 'basic' if x == 'a' else 'extra' if x == 'b' else 'extended' )
# state holiday
df2['state_holiday'] = df2['state_holiday'].apply( lambda x:'public_holiday' if x == 'a' else 'easter_holiday' if x == 'b' else 'christmas' if x == 'c' else 'regular_day' )
# + hidden=true
df2.head().T
# + [markdown] heading_collapsed=true
# # 3.0 PASSO 3 - FEATURE FILTERING
# + hidden=true
df3 = df2.copy()
# + hidden=true
df3.head()
# + [markdown] heading_collapsed=true hidden=true
# ## 3.1 Filtragem das Linhas
# + hidden=true
df3 = df3[ (df3['open'] != 0) & (df3['sales'] > 0) ]
# + [markdown] hidden=true
# ## 3.2 Seleção das Colunas
# + hidden=true
cols_drop = ['customers', 'open', 'promo_interval', 'month_map']
df3 = df3.drop( cols_drop, axis = 1)
# + hidden=true
df3.columns
# + [markdown] heading_collapsed=true
# # 4.0 PASSO 4 - EXPLORATORY DATA ANALYSIS
# + hidden=true
df4 = df3.copy()
# + [markdown] hidden=true
# ## 4.1 Análise Univariadaabs
# + [markdown] hidden=true
# ### 4.1.1 Response Variable
# + hidden=true hide_input=false
sns.distplot(df4['sales'])
# + [markdown] hidden=true
# ### 4.1.2 Numerical Variable
# + hidden=true hide_input=false
num_attributes.hist( bins = 25 );
# + [markdown] hidden=true
# ### 4.1.3 Categorical Variable
# + hidden=true
df4['state_holiday'].drop_duplicates()
# + hidden=true hide_input=false
# state holiday
plt.subplot( 3, 2, 1 )
a = df4[df4['state_holiday'] != 'regular_day']
sns.countplot( a['state_holiday'] )
plt.subplot( 3, 2, 2 )
sns.kdeplot( df4[df4['state_holiday'] == 'public_holiday']['sales'], label='public_holiday', shade=True )
sns.kdeplot( df4[df4['state_holiday'] == 'easter_holiday']['sales'], label='easter_holiday', shade=True )
sns.kdeplot( df4[df4['state_holiday'] == 'christmas']['sales'], label='christmas', shade=True )
# store type
plt.subplot( 3, 2, 3 )
sns.countplot( a['store_type'] )
plt.subplot( 3, 2, 4 )
sns.kdeplot( df4[df4['store_type'] == 'a']['sales'], label='a', shade=True )
sns.kdeplot( df4[df4['store_type'] == 'b']['sales'], label='b', shade=True )
sns.kdeplot( df4[df4['store_type'] == 'c']['sales'], label='c', shade=True )
sns.kdeplot( df4[df4['store_type'] == 'd']['sales'], label='d', shade=True )
# assortment
plt.subplot( 3, 2, 5 )
sns.countplot( a['assortment'] )
plt.subplot( 3, 2, 6 )
sns.kdeplot( df4[df4['assortment'] == 'extended']['sales'], label='extended', shade=True )
sns.kdeplot( df4[df4['assortment'] == 'basic']['sales'], label='basic', shade=True )
sns.kdeplot( df4[df4['assortment'] == 'extra']['sales'], label='extra', shade=True )
# + [markdown] heading_collapsed=true hidden=true
# ## 4.2 Análise Bivariada
# + [markdown] hidden=true
# ### H1. Lojas com maior sortimentos deveriam vender mais.
# Hipótese **FALSA**: lojas com maior sortimento vendem menos.
# + hidden=true
aux1 = df4[['assortment', 'sales']].groupby( 'assortment' ).sum().reset_index()
sns.barplot( x='assortment', y='sales', data=aux1 );
aux2 = df4[['year_week', 'assortment', 'sales']].groupby(['year_week','assortment'] ).sum().reset_index()
aux2.pivot( index='year_week', columns='assortment', values='sales' ).plot()
aux3 = aux2[aux2['assortment'] == 'extra']
aux3.pivot( index='year_week', columns='assortment', values='sales' ).plot()
# + hidden=true hide_input=false
aux1 = df4[['assortment','sales']].groupby('assortment').sum().reset_index()
sns.barplot( x='assortment', y='sales', data=aux1 );
aux2 = df4[['year_week','assortment','sales']].groupby(['year_week','assortment']).sum().reset_index()
aux2.pivot( index='year_week', columns='assortment', values='sales').plot()
aux3 = aux2[aux2['assortment'] == 'extra']
aux3.pivot( index='year_week', columns='assortment', values='sales').plot()
# + [markdown] hidden=true
# ### H2. Lojas com competidores mais próximos deveriam vender menos.
# Hipótese **FALSA**: lojas com **COMPETIDORES** mais próximos vendem **MAIS**.
# + hidden=true
aux1 = df4[['competition_distance','sales']].groupby('competition_distance').sum().reset_index()
plt.subplot( 1, 3, 1 )
sns.scatterplot( x='competition_distance', y='sales', data=aux1 )
plt.subplot( 1, 3, 2)
bins = list( np.arange(0, 20000, 1000) )
aux1['competition_distance_binned'] = pd.cut( aux1['competition_distance'], bins=bins )
aux2 = aux1[['competition_distance_binned','sales']].groupby('competition_distance_binned').sum().reset_index()
sns.barplot( x='competition_distance_binned', y='sales', data=aux2);
plt.xticks( rotation=90 )
plt.subplot( 1, 3, 3 )
sns.heatmap( aux1.corr( method='pearson' ), annot=True );
# + [markdown] hidden=true
# ### H3. Lojas com competidores à mais tempo deveriam vendem mais.
# Hipótese **FALSA**: lojas com **COMPETIDORES** à mais tempos vendem **MENOS**
# + hidden=true
plt.subplot( 1, 3, 1)
aux1 = df4[['competition_time_month','sales']].groupby( 'competition_time_month' ).sum().reset_index()
aux2 = aux1[(aux1['competition_time_month'] < 120) & (aux1['competition_time_month'] != 0)]
sns.barplot( x='competition_time_month', y='sales', data=aux2);
plt.xticks( rotation=90 );
plt.subplot( 1, 3, 2)
sns.regplot( x='competition_time_month', y='sales', data=aux2);
plt.subplot( 1, 3, 3 )
sns.heatmap( aux1.corr( method='pearson'), annot=True );
# + [markdown] hidden=true
# ### H4. Lojas com promoções ativas por mais tempo deveriam vender mais.
# Hipótese **FALSA**: Lojas com promoções ativas por mais tempo vendem menos, depois de um certo período de tempo de promoção.
# + hidden=true
aux1 = df4[['promo_time_week','sales']].groupby('promo_time_week').sum().reset_index()
plt.subplot(2,1,1)
aux2 = aux1[aux1['promo_time_week'] > 0 ] # promo extendida
sns.barplot(x='promo_time_week', y='sales', data=aux2);
plt.xticks( rotation=90 );
plt.subplot(2,1,2)
aux3 = aux1[aux1['promo_time_week'] < 0 ] # promo regular
sns.barplot(x='promo_time_week', y='sales', data=aux3);
plt.xticks( rotation=90 );
# + [markdown] hidden=true
# ###<s>H5. Lojas com mais dias de promoção deveriam vender mais.</s>
# + [markdown] hidden=true
# ### H7. Lojas com mais promoções consecutivas deveriam vender mais.
# Hipótese **FALSA**: Lojas com promoções consecutivas vendem menos.
# + hidden=true
df4[['promo','promo2','sales']].groupby(['promo','promo2']).sum().reset_index()
# + hidden=true
aux1 = df4[( df4['promo'] == 1 ) & (df4['promo2'] == 1)][['year_week','sales']].groupby('year_week').sum().reset_index()
ax = aux1.plot()
aux2 = df4[( df4['promo'] == 1 ) & (df4['promo2'] == 0)][['year_week','sales']].groupby('year_week').sum().reset_index()
aux2.plot( ax=ax )
ax.legend(labels=['Tradicional e Extendida','Tradicional'])
# + [markdown] heading_collapsed=true hidden=true
# ### **H8.** Lojas abertas durante o feriado de Natal deveriam vender mais.
# Hipótese **FALSA**: Lojas abertas durante o feriado de Natal vendem menos.
# + hidden=true
plt.subplot(1,2,1)
aux = df4[df4['state_holiday'] != 'regular_day']
aux1 = aux[['state_holiday', 'sales']].groupby('state_holiday').sum().reset_index()
sns.barplot( x='state_holiday', y='sales', data=aux1 );
plt.subplot(1,2,2)
aux2 = aux[['year','state_holiday', 'sales']].groupby( ['year', 'state_holiday'] ).sum().reset_index()
sns.barplot( x='year', y='sales', hue='state_holiday', data=aux2 );
# + [markdown] hidden=true
# ### **H9.** Lojas deveriam vender mais ao longo dos anos.
# Hipótese **FALSA**: Lojas vendem menos ao longo dos anos.
# + hidden=true
aux1 = df4[['year','sales']].groupby( 'year' ).sum().reset_index()
plt.subplot(1,3,1)
sns.barplot(x='year', y='sales', data=aux1);
plt.subplot(1,3,2)
sns.regplot(x='year', y='sales', data=aux1);
plt.subplot(1,3,3)
sns.heatmap(aux1.corr( method='pearson'), annot=True);
# + [markdown] hidden=true
# ### **H10.** Lojas deveriam vender mais no segundo semestre do ano.
# Hipótese **FALSA**: Lojas vendem menos durante o segundo semestre do ano.
# + hidden=true
aux1 = df4[['month','sales']].groupby( 'month' ).sum().reset_index()
plt.subplot(1,3,1)
sns.barplot(x='month', y='sales', data=aux1);
plt.subplot(1,3,2)
sns.regplot(x='month', y='sales', data=aux1);
plt.subplot(1,3,3)
sns.heatmap(aux1.corr( method='pearson'), annot=True);
# + [markdown] hidden=true
# ### **H11.** Lojas deveriam vender mais depois do dia 10 de cada mês.
# Hipótese **'VERDADEIRA**: Lojas vendem mais depois do dia 10 de cada mês.
# + hidden=true
aux1 = df4[['day','sales']].groupby( 'day' ).sum().reset_index()
plt.subplot(2,2,1)
sns.barplot(x='day', y='sales', data=aux1);
plt.subplot(2,2,2)
sns.regplot(x='day', y='sales', data=aux1);
plt.subplot(2,2,3)
sns.heatmap(aux1.corr( method='pearson'), annot=True);
aux1['before_after'] = aux1['day'].apply( lambda x: 'before_10_days' if x <= 10 else 'after_10_days' )
aux2 = aux1[['before_after','sales']].groupby( 'before_after' ).sum().reset_index()
plt.subplot(2,2,4)
sns.barplot( x ='before_after', y='sales', data=aux2 )
# + [markdown] hidden=true
# ### **H12.** Lojas deveriam vender menos aos finais de semana.
# Hipótese **VERDADEIRA**: Lojas vendem menos aos finais de semana.
# + hidden=true
aux1 = df4[['day_of_week','sales']].groupby( 'day_of_week' ).sum().reset_index()
plt.subplot(1,3,1)
sns.barplot(x='day_of_week', y='sales', data=aux1);
plt.subplot(1,3,2)
sns.regplot(x='day_of_week', y='sales', data=aux1);
plt.subplot(1,3,3)
sns.heatmap(aux1.corr( method='pearson'), annot=True);
# + [markdown] hidden=true
# ### **H13.**Lojas deveriam vender menos durante os feriados escolares.
# Hipótese **VERDADEIRA**: Lojas vendem menos durante os feriados escolares, exceto nos meses <NAME> e Agosto.
# + hidden=true
aux1 = df4[['school_holiday', 'sales']].groupby('school_holiday').sum().reset_index()
plt.subplot(2,1,1)
sns.barplot( x='school_holiday', y='sales', data=aux1);
aux2 = df4[['month', 'school_holiday', 'sales']].groupby(['month', 'school_holiday']).sum().reset_index()
plt.subplot(2,1,2)
sns.barplot(x='month', y='sales', hue='school_holiday', data=aux2)
# + [markdown] hidden=true
# ## 4.3 Analise Multivariada
# + [markdown] hidden=true hide_input=false
# ### 4.3.1 Numerical Attributes
# + hidden=true hide_input=false
# Only categorical data
a = df4.select_dtypes( include='object')
# Calculate cramer v
a1 = cramer_v( a['state_holiday'], a['state_holiday'] )
a2 = cramer_v( a['state_holiday'], a['store_type'] )
a3 = cramer_v( a['state_holiday'], a['assortment'] )
a4 = cramer_v( a['store_type'], a['state_holiday'] )
a5 = cramer_v( a['store_type'], a['store_type'] )
a6 = cramer_v( a['store_type'], a['assortment'] )
a7 = cramer_v( a['assortment'], a['state_holiday'] )
a8 = cramer_v( a['assortment'], a['store_type'] )
a9 = cramer_v( a['assortment'], a['assortment'] )
# Final DataSet
d = pd.DataFrame( {'state_holiday': [a1, a2, a3],
'store_type': [a4, a5, a6],
'assortment': [a7, a8, a9] })
d = d.set_index( d.columns )
# Heatmap
sns.heatmap( d, annot=True )
# + [markdown] heading_collapsed=true
# # 5.0 PASSO 5 - DATA PREPARATION
# + hidden=true
df5 = df4.copy()
# + [markdown] hidden=true
# ## 5.1 - Normalização
# + hidden=true
df5.head().T
# + [markdown] hidden=true
# ## 5.2 - Reescala
# + hidden=true
a = df5.select_dtypes( include=['int64', 'float64'] )
rs = RobustScaler()
mms = MinMaxScaler()
# Year
df5['year'] = mms.fit_transform( df5[['year']].values )
# competition distance
df5['competition_distance'] = rs.fit_transform( df5[['competition_distance']].values )
# competition time month
df5['competition_time_month'] = rs.fit_transform( df5[['competition_time_month']].values )
# promo time week
df5['promo_time_week'] = mms.fit_transform( df5[['promo_time_week']].values )
# + [markdown] hidden=true
# ## 5.3 - Transformação
# + [markdown] hidden=true
# ### 5.3.1 - Encoding
# + hidden=true
# state_holiday - Hot Encoding
df5 = pd.get_dummies( df5, prefix=['state_holiday'], columns=['state_holiday'] )
# store_type - Label Encoding
le = LabelEncoder()
df5['store_type'] = le.fit_transform( df5['store_type'] )
# assortment - Ordinal Encoding
assortment_dict={'basic':1, 'extra':2, 'extended':3 }
df5['assortment'] = df5['assortment'].map( assortment_dict )
# + [markdown] hidden=true
# ### 5.3.2 - Transformação da Variavel Resposta
# + hidden=true
df5['sales'] = np.log1p( df5['sales'] )
# + hidden=true
# day of week
df5['day_of_week_sin'] = df5['day_of_week'].apply( lambda x: np.sin( x * ( 2. * np.pi/7 ) ) )
df5['day_of_week_cos'] = df5['day_of_week'].apply( lambda x: np.sin( x * ( 2. * np.pi/7 ) ) )
# month
df5['month_sin'] = df5['month'].apply( lambda x: np.sin( x * ( 2. * np.pi/12 ) ) )
df5['month_cos'] = df5['month'].apply( lambda x: np.sin( x * ( 2. * np.pi/12 ) ) )
# day
df5['day_sin'] = df5['day'].apply( lambda x: np.sin( x * ( 2. * np.pi/30 ) ) )
df5['day_cos'] = df5['day'].apply( lambda x: np.sin( x * ( 2. * np.pi/30 ) ) )
# week of year
df5['week_of_year_sin'] = df5['week_of_year'].apply( lambda x: np.sin( x * ( 2. * np.pi/52 ) ) )
df5['week_of_year_cos'] = df5['week_of_year'].apply( lambda x: np.sin( x * ( 2. * np.pi/52 ) ) )
# + hidden=true
df5.head().T
# + [markdown] heading_collapsed=true
# # 6.0 PASSO 6 - FEATURE SELECTION
# + hidden=true
df6 = df5.copy()
# + [markdown] hidden=true
# ## 6.1 - Split dataframe into training and test dataset
# + hidden=true
cols_drop = ['week_of_year','day','month','day_of_week','promo_since','competition_since','year_week']
df6 = df6.drop( cols_drop, axis=1 )
# + hidden=true
df6[['store','date']].groupby('store').max().reset_index()['date'][0] - datetime.timedelta( days=6*7 )
# + hidden=true
# Training dataset
x_train = df6[df6['date'] < '2015-06-19']
y_train = x_train['sales']
# Test dataset
x_test = df6[df6['date'] >= '2015-06-19']
y_test = x_test['sales']
print('Training min date: {}'.format( x_train['date'].min() ) )
print('Training max date: {}'.format( x_train['date'].max() ) )
print('\nTest min date: {}'.format( x_test['date'].min() ) )
print('Test min date: {}'.format( x_test['date'].max() ) )
# + [markdown] hidden=true
# ## 6.2 - Boruta as Feature Selector
# + hidden=true
# training and test dataset for Boruta
# x_train_n = x_train.drop(['date','sales'], axis=1 ).values
# y_train_n = y_train.values.ravel()
# Define RandomForest Regressor
# rf = RandomForestRegressor( n_jobs=-1 )
# Define Boruta
# boruta = BorutaPy( rf, n_estimators='auto', verbose=2, random_state=42 ).fit( x_train_n, y_train_n )
# + [markdown] hidden=true
# ### 6.2.1 - Best features from Boruta
# + hidden=true
#cols_selected = boruta.support_.tolist()
# Best Features
#x_train_fs = x_train.drop(['date','sales'], axis=1 )
#cols_selected_boruta = x_train_fs.iloc[:, cols_selected].columns.to_list()
#cols_not_selected_boruta = list(np.setdiff1d(x_train_fs.columns, cols_selected_boruta ) )
# + [markdown] hidden=true
# ## 6.3 - Manual Feature Selection
# + hidden=true
cols_selected_boruta = [
'store',
'promo',
'month_cos',
'month_sin',
'store_type',
'assortment',
'competition_distance',
'competition_open_since_month',
'competition_open_since_year',
'promo2',
'promo2_since_week',
'promo2_since_year',
'competition_time_month',
'promo_time_week',
'day_of_week_sin',
'day_of_week_cos',
'week_of_year_cos',
'week_of_year_sin',
'day_sin',
'day_cos']
# Columns to add
feat_to_add = ['date','sales']
# final features
cols_selected_boruta_full = cols_selected_boruta.copy()
cols_selected_boruta_full.extend( feat_to_add )
# + [markdown] heading_collapsed=true
# # 7.0 PASSO 7 - ML MODEL'S
# + hidden=true
X_train = x_train[ cols_selected_boruta ]
X_test = x_test[ cols_selected_boruta ]
# time series data preparation
x_training = x_train[ cols_selected_boruta_full ]
# + [markdown] hidden=true
# ## 7.1 - Average Model
# + hidden=true
aux1 = X_test.copy()
aux1['sales'] = y_test.copy()
# Predictions
aux2 = aux1[['store', 'sales']].groupby('store').mean().reset_index().rename( columns={'sales': 'predictions'} )
aux1 = pd.merge( aux1, aux2, how='left', on='store' )
yhat_baseline= aux1['predictions']
# Performance
baseline_result = ml_error( 'Average Model', np.expm1( y_test ), np.expm1( yhat_baseline ) )
baseline_result
# + [markdown] heading_collapsed=true hidden=true
# ## 7.2 - Linear Regression Model
# + hidden=true
# model
lr = LinearRegression().fit( X_train, y_train )
# prediction
yhat_lr = lr.predict( X_test )
# performance
lr_result = ml_error( 'Linear Regression', np.expm1( y_test ), np.expm1( yhat_lr ) )
lr_result
# + [markdown] hidden=true
# ### 7.2.1 - Linear Regression Model: Cross Validation
# + hidden=true
lr_result_cv = cross_validation( x_training, 5, 'Linear Regression', lr, verbose=False )
lr_result_cv
# + [markdown] heading_collapsed=true hidden=true
# ## 7.3 - Linear Regression Regularized Model
# + hidden=true
# model
lrr = Lasso( alpha=0.01 ).fit( X_train, y_train )
# prediction
yhat_lrr = lrr.predict( X_test )
# performance
lrr_result = ml_error( 'Linear Regression - Lasso', np.expm1( y_test ), np.expm1( yhat_lrr ) )
lrr_result
# + [markdown] hidden=true
# ### 7.3.1 - Linear Regression Regularized Model - Cross Validation
# + hidden=true
lrr_result_cv = cross_validation( x_training, 5, 'Lasso', lrr, verbose=False )
lrr_result_cv
# + [markdown] heading_collapsed=true hidden=true
# ## 7.4 - Random Forest Regressor
# + hidden=true
# model
rf = RandomForestRegressor( n_estimators=100, n_jobs=-1, random_state=42 ).fit( X_train, y_train )
# prediction
yhat_rf = rf.predict( X_test )
# performance
rf_result = ml_error( 'Random Forest Regressor', np.expm1( y_test ), np.expm1( yhat_rf ) )
rf_result
# + [markdown] heading_collapsed=true hidden=true
# ### 7.4.1 - Randon Forest Regresson - Cross Validation
# + hidden=true
rf_result_cv = cross_validation( x_training, 5, 'Randon Forest Regressor', rf, verbose=True )
rf_result_cv
# + [markdown] heading_collapsed=true hidden=true
# ## 7.5 - XGBoost Regressor
# + hidden=true
# model
model_xgb = xgb.XGBRegressor( objective='reg:squarederror',
n_estimators=100,
eta=0.01,
max_depth=10,
subsample=0.7,
colsample_bytree=0.9 ).fit( X_train, y_train )
# prediction
yhat_xgb = model_xgb.predict( X_test )
# performance
xgb_result = ml_error( 'XGBoost Regressor', np.expm1( y_test ), np.expm1( yhat_xgb ) )
xgb_result
# + [markdown] hidden=true
# ### 7.5.1 - XGBoost Regressor - Cross Validation
# + hidden=true
xgb_result_cv = cross_validation( x_training, 5, 'XGboost Regressor', model_xgb, verbose=True )
xgb_result_cv
# + [markdown] heading_collapsed=true hidden=true
# ## 7.6 - Compare Model's Performance
# + [markdown] heading_collapsed=true hidden=true
# ### 7.6.1 - Single Performance
# + hidden=true
modelling_result = pd.concat( [baseline_result, lr_result, lrr_result, rf_result, xgb_result] )
modelling_result.sort_values( 'RMSE' )
# + [markdown] heading_collapsed=true hidden=true
# ### 7.6.2 - Real Performance: Cross Validation
# + hidden=true
modelling_result_cv = pd.concat( [lr_result_cv, lrr_result_cv, rf_result_cv, xgb_result_cv] )
modelling_result_cv.sort_values('RMSE CV')
# + [markdown] heading_collapsed=true
# # 8.0 PASSO 8 - HYPERPARAMETER FINE TUNING
# + [markdown] heading_collapsed=true hidden=true
# ## 8.1 - Random Search
# + hidden=true
# param = {
# 'n_estimators': [1500, 1700, 2500, 3000, 3500],
# 'eta': [0.01, 0.03],
# 'max_depth': [3, 5, 9],
# 'subsample': [0.1, 0.5, 0.7],
# 'colsample_bytree': [0.3, 0.7, 0.9],
# 'min_child_weight':[3, 8, 15] }
# MAX_EVAL = 5
# + hidden=true
# final_result = pd.DataFrame()
# for i in range( MAX_EVAL ):
# # choose values for parameters randomly
# hp = { k: random.sample(v, 1)[0] for k, v in param.items() }
# print( hp )
# # model
# model_xgb = xgb.XGBRegressor( objective='reg:squarederror',
# n_estimators=hp['n_estimators'],
# eta=hp['eta'],
# max_depth=hp['max_depth'],
# subsample=hp['subsample'],
# colsample_bytree=hp['colsample_bytree'],
# min_child_weight=hp['min_child_weight'] )
# # performance
# result = cross_validation( x_training, 5, 'XGBoost Regressor', model_xgb, verbose=True )
# final_result = pd.concat( [final_result, result] )
# final_result
# + [markdown] heading_collapsed=true hidden=true
# ## 8.2 - Final Model
# + hidden=true
param_tuned = {
'n_estimators':1500,
'eta':0.03,
'max_depth':9,
'subsample':0.1,
'colsample_bytree':0.3,
'min_child_weight':3}
# + hidden=true
model_xgb_tuned = xgb.XGBRegressor( objective='reg:squarederror',
n_estimators=param_tuned['n_estimators'],
eta=param_tuned['eta'],
max_depth=param_tuned['max_depth'],
subsample=param_tuned['subsample'],
colsample_bytree=param_tuned['colsample_bytree'],
min_child_weight=param_tuned['min_child_weight'] ).fit( X_train, y_train )
# prediction
yhat_xgb_tuned = model_xgb_tuned.predict( X_test )
# performance
xgb_result_tuned = ml_error( 'XGBoost Regressor', np.expm1( y_test), np.expm1( yhat_xgb_tuned ) )
xgb_result_tuned
# + hidden=true
mpe = mean_percentage_error( np.expm1(y_test), np.expm1(yhat_xgb_tuned) )
mpe
# + hidden=true
# + [markdown] heading_collapsed=true
# # 9.0 PASSO 9 - EVALUATION
# + hidden=true
df9 = x_test[ cols_selected_boruta_full ]
# rescale
df9['sales'] = np.expm1( df9['sales'] )
df9['predictions'] = np.expm1( yhat_xgb_tuned )
# + [markdown] heading_collapsed=true hidden=true
# ## 9.1 - Business Performance
# + hidden=true
# sum of predictions
df91 = df9[['store', 'predictions']].groupby( 'store' ).sum().reset_index()
# MAE and MAPE
df9_aux1 = df9[['store','sales','predictions']].groupby( 'store' ).apply( lambda x: mean_absolute_error( x['sales'], x['predictions'] ) ).reset_index().rename( columns={0:'MAE'} )
df9_aux2 = df9[['store','sales','predictions']].groupby( 'store' ).apply( lambda x: mean_absolute_percentage_error( x['sales'], x['predictions'] ) ).reset_index().rename( columns={0:'MAPE'} )
# Merge
df9_aux3 = pd.merge( df9_aux1, df9_aux2, how='inner', on='store')
df92 = pd.merge( df91, df9_aux3, how='inner', on='store')
# Scenarios
df92['worst_scenario'] = df92['predictions'] - df92['MAE']
df92['best_scenario'] = df92['predictions'] + df92['MAE']
# order columns
df92 = df92[['store','predictions','worst_scenario','best_scenario','MAE','MAPE']]
# + hidden=true
df92.head()
# + hidden=true
df92.sort_values( 'MAPE', ascending=False).head()
# + hidden=true
sns.scatterplot( x='store', y='MAPE', data=df92)
# + [markdown] heading_collapsed=true hidden=true
# ## 9.2 - Total Performance
# + hidden=true
df93 = df92[['predictions','worst_scenario','best_scenario']].apply( lambda x: np.sum( x ), axis=0 ).reset_index().rename( columns={'index': 'Scenario', 0: 'Values'})
df93['Values'] = df93['Values'].map( 'R${:,.2f}'.format )
df93
# + [markdown] heading_collapsed=true hidden=true
# ### 9.3 - ML Performance
# + hidden=true
df9['error'] = df9['sales'] - df9['predictions']
df9['error_rate'] = df9['predictions'] / df9['sales']
# + hidden=true
plt.subplot( 2, 2, 1 )
sns.lineplot( x='date', y='sales', data=df9, label='SALES')
sns.lineplot( x='date', y='predictions', data=df9, label='PREDICTIONS')
plt.subplot( 2, 2, 2 )
sns.lineplot( x='date', y='error_rate', data=df9)
plt.axhline(1, linestyle='--')
plt.subplot( 2, 2, 3 )
sns.distplot( df9['error'] )
plt.subplot( 2, 2, 4 )
sns.scatterplot( df9['predictions'], df9['error'] )
# + hidden=true
import pickle
# ML trained saving
pickle.dump( model_xgb_tuned, open('/Users/ricadesk/Documents/ds_repos/ds_em_producao/model/model_rossmann.pkl', 'wb' ) )
infile = open('/Users/ricadesk/Documents/ds_repos/ds_em_producao/model/model_rossmann.pkl', 'rb')
model_xgb_tuned = pickle.load(infile)
xgb_result_tuned
| notebooks/m09_v01_store_sales_prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Gather
# +
import pandas as pd
import requests
import tweepy
import json
import re
import matplotlib.pyplot as plt
import numpy as np
import statsmodels.api as sm
import random
# %matplotlib inline
random.seed(42)
# -
# read in the twitter data from file on hand
twitter_arch = pd.read_csv('twitter-archive-enhanced.csv')
# download the image predictions data
url = 'https://d17h27t6h515a5.cloudfront.net/topher/2017/August/599fd2ad_image-predictions/image-predictions.tsv'
r = requests.get(url)
with open(url.split('/')[-1],'wb') as file:
file.write(r.content)
# read in the image predictions data
image_preds = pd.read_csv('image-predictions.tsv',sep='\t')
# +
# access the twitter api
with open('twitter_dev_keys.txt','r') as file:
keys = file.readlines()
keys = [key.rstrip('\n') for key in keys]
consumer_key = keys[1]
consumer_secret = keys[3]
access_token = keys[7]
access_token_secret = keys[9]
auth = tweepy.OAuthHandler(consumer_key, consumer_secret)
auth.set_access_token(access_token, access_token_secret)
api = tweepy.API(auth, wait_on_rate_limit=True, wait_on_rate_limit_notify=True)
# -
# download the tweets & write them
with open('tweet_json.txt','a') as file:
for i in twitter_arch['tweet_id'].items():
# in case tweet has been deleted
try:
tweet = api.get_status(i[1], tweet_mode='extended')
json.dump(tweet._json, file)
file.write('\n')
except tweepy.TweepError:
print('id {} not available'.format(i[1]))
# read json data into dataframe
# note here this is a solution from Stack Overflow; json.load(file) fails, apparently if have multiple objects
with open('tweet_json.txt') as jfile:
jtweet = [json.loads(line) for line in jfile]
ext_tweet = pd.DataFrame.from_dict(jtweet)
# just keep the relevant columns
ext_tweet = ext_tweet[['id','id_str','retweet_count','favorite_count']]
# # Assess
display(twitter_arch.head())
display(twitter_arch.tail())
display(twitter_arch.sample(20))
display(twitter_arch.info())
display(twitter_arch.describe())
display(twitter_arch['rating_numerator'].value_counts())
display(twitter_arch['rating_denominator'].value_counts())
display(twitter_arch[twitter_arch['rating_denominator'] != 10])
display(twitter_arch['text'][313])
display(twitter_arch['text'][342])
display(twitter_arch['text'][433])
display(twitter_arch['text'][516])
display(twitter_arch['text'][902])
display(twitter_arch['text'][1068])
display(twitter_arch['text'][1120])
display(twitter_arch['text'][1165])
display(twitter_arch['text'][1202])
display(twitter_arch['text'][1228])
display(twitter_arch['text'][1254])
display(twitter_arch['text'][1274])
display(twitter_arch['text'][1351])
display(twitter_arch['text'][1433])
display(twitter_arch['text'][1598])
display(twitter_arch['text'][1634])
display(twitter_arch['text'][1635])
display(twitter_arch['text'][1662])
display(twitter_arch['text'][1663])
display(twitter_arch['text'][1779])
display(twitter_arch['text'][1843])
display(twitter_arch['text'][2335])
display(twitter_arch[(twitter_arch['rating_numerator'] > 20) & (twitter_arch['rating_denominator'] == 10)])
display(twitter_arch['text'][188])
display(twitter_arch['text'][189])
display(twitter_arch['text'][290])
display(twitter_arch['text'][340])
display(twitter_arch['text'][695])
display(twitter_arch['text'][763])
display(twitter_arch['text'][979])
display(twitter_arch['text'][1712])
display(twitter_arch['text'][2074])
display(twitter_arch['name'].value_counts()[0:59])
display(twitter_arch['name'].value_counts()[60:119])
# non-names appear to be lowercase.
not_names = twitter_arch['name'][twitter_arch['name'].str.match(r'^[a-z]')==True].value_counts()
display(not_names)
# see if some of the names are discernible from text
display(twitter_arch['text'][twitter_arch['name'].str.match(r'^[a-z]')==True])
display(twitter_arch['text'][twitter_arch['name'].str.match(r'^[a-z]')==True][22])
display(twitter_arch['text'][twitter_arch['name'].str.match(r'^[a-z]')==True][56])
display(twitter_arch['text'][twitter_arch['name'].str.match(r'^[a-z]')==True][118])
display(twitter_arch['text'][twitter_arch['name'].str.match(r'^[a-z]')==True][2354])
display(twitter_arch['text'][twitter_arch['name'].str.match(r'^[a-z]')==True][2353])
display(twitter_arch['text'][twitter_arch['name'].str.match(r'^[a-z]')==True][2352])
# see if consistent tagging of non-dog entries
# see if some of the names are discernible from text
display(twitter_arch[twitter_arch['text'].str.match(r'.+[Ww]e only rate dogs.+')==True])
# most of those look like non-dogs
display(twitter_arch['text'][25])
display(twitter_arch['doggo'].value_counts())
display(twitter_arch['floofer'].value_counts())
display(twitter_arch['pupper'].value_counts())
display(twitter_arch['puppo'].value_counts())
# these fields should be exclusive, but in tidying found some have multiple stage entries
twitter_arch[(twitter_arch['doggo'] != "None") & (twitter_arch['floofer']!="None")]
# this appears to be a non-dog
display(twitter_arch['text'][200])
twitter_arch[(twitter_arch['doggo'] != "None") & (twitter_arch['puppo']!="None")]
# this appears to be parsed incorrectly due to "doggo" also being part of the tweet
display(twitter_arch['text'][191])
# based on the texts of these, moost appear to be multiple or non-dog
twitter_arch[(twitter_arch['doggo'] != "None") & (twitter_arch['pupper']!="None")]
# look at individual entries for those not obvious from snippets above
display(twitter_arch['text'][460])
display(twitter_arch['text'][575])
display(twitter_arch['text'][705])
display(twitter_arch['text'][889])
display(twitter_arch['text'][1063])
twitter_arch[(twitter_arch['puppo'] != "None") & (twitter_arch['pupper']!="None")]
twitter_arch[(twitter_arch['floofer'] != "None") & (twitter_arch['pupper']!="None")]
twitter_arch[(twitter_arch['puppo'] != "None") & (twitter_arch['floofer']!="None")]
display(image_preds.head())
display(image_preds.tail())
display(image_preds.sample(20))
display(image_preds.info())
display(image_preds.describe())
display(image_preds['tweet_id'].nunique())
display(image_preds['p1'].value_counts())
display(ext_tweet.head())
display(ext_tweet.tail())
display(ext_tweet.info())
display(ext_tweet.describe())
# # Clean
#
# ### Quality
#
# `twitter_arch` contains retweets, which we want to exclude
#
# ##### Define
# Delete retweet entries
#
# ##### Code
# back up data first
twitter_archbu = twitter_arch.copy()
twitter_archbu = twitter_archbu[twitter_archbu['retweeted_status_id'].isnull()]
# ##### Test
twitter_archbu['retweeted_status_id'].value_counts()
twitter_archbu.sample(10)
# ##### Code
# the retweeted columns are now empty, so delete them
twitter_archbu.drop(columns=['retweeted_status_id','retweeted_status_user_id','retweeted_status_timestamp'], inplace=True)
# ##### Test
twitter_archbu.head()
# looks good, copy back
twitter_arch = twitter_archbu.copy()
# `twitter_arch` contains tweets about non-dog things, which we want to exclude
#
# ##### Define
# Delete the non-dog entries. Since [@dograte's](https://twitter.com/dog_rates) replies to these consistently include "we only rate dogs", it is assumed any entry with this phrase in `text` is a non-dog entry.
#
# ##### Code
twitter_archbu = twitter_archbu[twitter_archbu['text'].str.match(r'.+[Ww]e only rate dogs.+')==False]
# ##### Test
display(twitter_archbu[twitter_archbu['text'].str.match(r'.+[Ww]e only rate dogs.+')==True])
# looks good, so copy back
twitter_arch = twitter_archbu.copy()
# `twitter_arch` contains ratings data that aren't ratings
#
# ##### Define
# Replace the erroneous ratings with actual ratings from `text` or delete if no rating was found in `text`.
#
# ##### Code
# these are the entries that appear to be parsing errors and had correct-appearing ratings in the tweet
rating_fix = [{'index': 313, 'numerator':13},{'index':1068, 'numerator':14},{'index':1165, 'numerator':13},\
{'index':1202, 'numerator':11},{'index':1662, 'numerator':10}, {'index':2335, 'numerator':9},\
{'index':695, 'numerator':10},{'index':763, 'numerator':11}]
rate_fix = pd.DataFrame(rating_fix)
rate_fix.head()
twitter_archbu = twitter_arch.copy()
for i in rate_fix['index']:
twitter_archbu.loc[i,'rating_numerator'] = rate_fix.loc[(rate_fix['index']==i),'numerator'].values
twitter_archbu.loc[i,'rating_denominator'] = 10
# ##### Test
display(twitter_archbu.loc[rate_fix['index']])
# looks good, copy back
twitter_arch = twitter_archbu.copy()
# ##### Code
# these did not have obvious ratings in the text, so delete the entries
rate_del = [342, 516, 1598, 1663, 1712]
twitter_archbu.drop(rate_del, inplace=True)
# ##### Test
try:
display(twitter_archbu.loc[rate_del])
except KeyError:
print('values not found')
# looks good, copy back
twitter_arch = twitter_archbu.copy()
# `twitter_arch` contains ratings for multiple-dog groups
#
# ##### Define
# Remove the multi-dog entries, which have denominators <> 10
#
# ##### Code
twitter_archbu = twitter_archbu[twitter_archbu['rating_denominator'] == 10]
# ##### Test
display(twitter_archbu[twitter_archbu['rating_denominator'] != 10])
display(twitter_archbu.head())
# looks good, copy back
twitter_arch = twitter_archbu.copy()
# `twitter_arch` contains non-name entries in `name`
#
# ##### Define
# Remove the multi-dog entries, which have denominators <> 10
#
# ##### Code
no_names = twitter_archbu['name'][twitter_archbu['name'].str.match(r'^[a-z]')==True].index
twitter_archbu.loc[no_names,'name'] = 'None'
# ##### Test
display(twitter_archbu['name'][twitter_archbu['name'].str.match(r'^[a-z]')==True].value_counts())
# check one of the entries to make sure it's set correctly
twitter_archbu.loc[56,'name']
# looks good, copy back
twitter_arch = twitter_archbu.copy()
# `twitter_arch` fields `timestamp` and `retweeted_status_timestamp` are strings, not datetime
#
# ##### Define
# Convert `timestamp` to datetime. `retweeted_status_timestamp` has been deleted.
#
# ##### Code
twitter_archbu['timestamp'] = pd.to_datetime(twitter_archbu['timestamp'])
# ##### Test
display(twitter_archbu.info())
display(twitter_archbu.head())
# looks good, copy back
twitter_arch = twitter_archbu.copy()
# `twitter_arch` fields `in_reply_to_status_id`,`in_reply_to_user_id`, `retweeted_status_id` and `retweeted_user_id` are floats, not ints
#
# ##### Define
# Convert `in_reply_to_status_id` and `in_reply_to_user_id` to int. `retweeted_status_id` and `retweeted_user_id` have been deleted.
#
# ##### Code
# the pd.Int64DType() is to get around a error due to float NaNs
twitter_archbu['in_reply_to_status_id'] = twitter_archbu['in_reply_to_status_id'].astype(pd.Int64Dtype())
twitter_archbu['in_reply_to_user_id'] = twitter_archbu['in_reply_to_user_id'].astype(pd.Int64Dtype())
# ##### Test
display(twitter_archbu.info())
display(twitter_archbu.head())
# looks good, copy back
twitter_arch = twitter_archbu.copy()
# `twitter_arch` dog-stage fields `floofer`,`pupper`, `puppo` and `doggo` should be mutually exclusive, but some entries have more than one stage
#
# ##### Define
# Delete the entries that appear to be non-dog or multi-dog. Correct the ones that appear to be parsed incorrectly.
#
# ##### Code
# These two are parsed incorrectly. Set doggo field to none
twitter_archbu.loc[191,'doggo'] = "None"
twitter_archbu.loc[460,'doggo'] = "None"
# remove the other entries as are multi-dog or non-dog
twitter_archbu = twitter_archbu[~((twitter_archbu['doggo'] != "None") & (twitter_archbu['floofer']!="None"))]
twitter_archbu = twitter_archbu[~((twitter_archbu['doggo'] != "None") & (twitter_archbu['pupper']!="None"))]
# ##### Test
twitter_archbu[(twitter_archbu['doggo'] != "None") & (twitter_archbu['floofer']!="None")]
twitter_archbu[(twitter_archbu['doggo'] != "None") & (twitter_archbu['pupper']!="None")]
# looks good, copy back
twitter_arch = twitter_archbu.copy()
# `image_preds` fields `p1`, `p2`, and `p3` are inconsistently capitalized
#
# ##### Define
# Make all entries in `p1`, `p2`, and `p3` lowercase for consistency.
#
# ##### Code
image_predsbu = image_preds.copy()
image_predsbu['p1'] = image_predsbu['p1'].str.lower()
image_predsbu['p2'] = image_predsbu['p2'].str.lower()
image_predsbu['p3'] = image_predsbu['p3'].str.lower()
# ##### Test
image_predsbu.sample(10)
# looks good, copy back
image_preds = image_predsbu.copy()
# `image_preds` contains fewer entries than the twitter archive.
#
# ##### Define
# This means some tweets have no images. This will be addressed in "Tidiness" when the tables are joined.
# `ext_tweet` contains fewer entries than the twitter archive.
#
# ##### Define
# This means some tweets were deleted or are otherwise not accessible. This will be addressed in "Tidiness" when the tables are joined.
# ### Tidiness
#
# `twitter_arch` fields `doggo`, `floofer`, `pupper` and `puppo` are mutually exclusive descriptions of dog stage
#
# ##### Define
# Make a single categorical field `stage` containing each stage name or "none".
#
# ##### Code
twitter_archmelt = pd.melt(twitter_archbu, id_vars=['tweet_id','in_reply_to_status_id','in_reply_to_user_id',\
'timestamp','source','expanded_urls','rating_numerator',\
'rating_denominator','name'],value_vars=['doggo','floofer',\
'pupper','puppo'])
# first get rid of all the duplicate "None" entries
twitter_archmelt.drop_duplicates(subset=['tweet_id','value'],inplace=True)
twitter_archmelt.info()
twitter_archmelt['value'].value_counts()
twitter_archmelt['drop'] = twitter_archmelt.duplicated(subset=['tweet_id'], keep=False)
twitter_archmelt['drop'].value_counts()
# find the "None" entries that are duplicates of those with a stage entry
drop_rows = twitter_archmelt[(twitter_archmelt['drop']==True) & (twitter_archmelt['value']=="None")].index
len(drop_rows)
# drop those rows
twitter_archmelt.drop(drop_rows,inplace=True)
# clean up the columns & names
twitter_archmelt.drop(columns=['variable','drop'], inplace=True)
twitter_archmelt.rename(columns={'value':'stage'}, inplace=True)
twitter_archmelt['stage'] = twitter_archmelt['stage'].astype('category')
# ##### Test
twitter_archmelt['stage'].value_counts()
twitter_archmelt.duplicated(subset=['tweet_id'],keep=False).sum()
display(twitter_archmelt.head())
display(twitter_archmelt.info())
#looks good, copy back
twitter_archbu = twitter_archmelt.copy()
twitter_arch = twitter_archmelt.copy()
# `image_preds`should be combined with the main archive table since there is one entry per tweet
#
# ##### Define
# Merge `image_preds` with `twitter_arch`. Inner join so non-image-containing tweets are eliminated.
#
# ##### Code
twitter_archbu = twitter_archbu.merge(image_predsbu,on='tweet_id')
# ##### Test
display(twitter_archbu.head())
display(twitter_archbu.info())
#looks good, copy back
twitter_arch = twitter_archbu.copy()
# `ext_tweets`should be combined with the main archive table since it has additional data about each tweet
#
# ##### Define
# Merge `ext_tweet` with `twitter_arch`. Inner join so no-longer-accessible tweets are eliminated.
#
# ##### Code
twitter_archbu = twitter_archbu.merge(ext_tweet,left_on='tweet_id',right_on='id')
# drop extra columns
twitter_archbu.drop(columns=['id','id_str'], inplace=True)
# ##### Test
display(twitter_archbu.head())
display(twitter_archbu.info())
#looks good, copy back
twitter_arch = twitter_archbu.copy()
# store the cleaned data
twitter_arch.to_csv('twitter_archive_master.csv')
# # Analyze
#
# Are dogs with higher ratings more frequently favorited and retweeted?
twitter_arch['rating_numerator'].describe()
twitter_arch['retweet_count'].describe()
twitter_arch['favorite_count'].describe()
twitter_arch.plot(x='rating_numerator', y='retweet_count', kind='scatter')
plt.xlabel('Rating')
plt.ylabel('Retweets')
plt.title('Retweet Count as Function of Rating');
# remove the 2 outliers
rate_nooutl = twitter_arch[twitter_arch['rating_numerator']< 250]
rate_nooutl.plot(x='rating_numerator', y='retweet_count', kind='scatter')
plt.xlabel('Rating')
plt.ylabel('Retweets')
plt.title('Retweet Count as Function of Rating');
rate_nooutl.plot(x='rating_numerator', y='favorite_count', kind='scatter')
plt.xlabel('Rating')
plt.ylabel('Favorites')
plt.title('Favorite Count as Function of Rating');
# split ratings into < 10 vs >= 10
rate_nooutl['rate_hilo'] = rate_nooutl.apply(lambda row: row['rating_numerator']>=10, axis=1)
rate_nooutl.head()
rate_avg = rate_nooutl.groupby(['rate_hilo'], as_index=False)[['favorite_count','retweet_count']].mean()
rate_avg.head()
ind = pd.array([1.0, 2.0])
plt.bar(ind, rate_avg['favorite_count'])
plt.xticks(ind, ['< 10', '>= 10'])
plt.xlabel('Rating')
plt.ylabel('Favorites')
plt.title('Figure 1. Mean Favorites by Low vs. High Rating');
# Higher-rated dogs appear to be favorited more frequently.
#
# Check if this is statistically significant:
obs_diff_favs = rate_avg.loc[1,'favorite_count'] - rate_avg.loc[0,'favorite_count']
obs_diff_favs
# create sampling distribution of difference in favorites
# with boostrapping
diffs = np.empty(10000, dtype=float)
size = rate_nooutl.shape[0]
for x in range(10000):
smplx = rate_nooutl.sample(size,replace=True)
lo_mn = smplx.query('rate_hilo == False').favorite_count.mean()
hi_mn = smplx.query('rate_hilo == True').favorite_count.mean()
diffs[x] = hi_mn - lo_mn
plt.hist(diffs);
np.std(diffs)
# simulate distribution under the null hypothesis
null_vals = np.random.normal(0, np.std(diffs), 10000)
# +
# plot null distribution
plt.hist(null_vals)
# plot line for observed statistic
plt.axvline(obs_diff_favs,color='red',lw=2);
# -
# compute p value
pval = (null_vals > obs_diff_favs).mean()
pval
# The difference is highly significant.
ind = pd.array([1.0, 2.0])
plt.bar(ind, rate_avg['retweet_count'])
plt.xticks(ind, ['< 10', '>= 10'])
plt.xlabel('Rating')
plt.ylabel('Retweets')
plt.title('Figure 2. Mean Retweets by Low vs. High Rating');
# Higher-rated dogs appear to be retweeted more frequently.
#
# Check if this is statistically significant:
obs_diff_rts = rate_avg.loc[1,'retweet_count'] - rate_avg.loc[0,'retweet_count']
obs_diff_rts
# create sampling distribution of difference in favorites
# with boostrapping
diffs = np.empty(10000, dtype=float)
size = rate_nooutl.shape[0]
for x in range(10000):
smplx = rate_nooutl.sample(size,replace=True)
lo_mn = smplx.query('rate_hilo == False').retweet_count.mean()
hi_mn = smplx.query('rate_hilo == True').retweet_count.mean()
diffs[x] = hi_mn - lo_mn
plt.hist(diffs);
np.std(diffs)
# simulate distribution under the null hypothesis
null_vals = np.random.normal(0, np.std(diffs), 10000)
# +
# plot null distribution
plt.hist(null_vals)
# plot line for observed statistic
plt.axvline(obs_diff_rts,color='red',lw=2);
# -
# compute p value
pval = (null_vals > obs_diff_favs).mean()
pval
# Again, the difference is highly significant
# Do dog ratings differ by stage?
rate_stg = rate_nooutl.groupby(['stage'], as_index=False)[['rating_numerator']].mean()
rate_stg
ind2 = pd.array([1.0, 2.0, 3.0, 4.0, 5.0])
plt.bar(ind2, rate_stg['rating_numerator'])
plt.xticks(ind2, ['none','doggo','floofer','pupper','puppo'])
plt.xlabel('Stage')
plt.ylabel('Rating')
plt.title('Figure 3. Mean Rating by Dog Stage');
# Dog ratings do seem to vary slightly by stage. Dogs staged as `doggo`, `floofer`, and `puppo` may have slighltly higher ratings than dogs with no stage or dogs staged as `pupper`.
#
# To see if there are significant differences, perform a linear regression on ratings as function of stage:
# mucked up dummy cols, restart
rate_nooutl.drop(columns=['doggo','floofer','none','pupper','puppo','None'],inplace=True)
display(rate_nooutl.head())
# make dummy variables
rate_nooutl[['None','doggo','floofer','pupper','puppo']]=pd.get_dummies(rate_nooutl['stage'])
rate_nooutl.head()
# perform linear regression on ratings as fxn of stage
rate_nooutl['intercept'] = 1
mdl_stg = sm.OLS(rate_nooutl['rating_numerator'],rate_nooutl[['intercept','doggo','floofer','pupper','puppo']])
stg_res = mdl_stg.fit()
stg_res.summary()
# The linear regression does show some significant differences, although the R-squared value is low (0.02), indicating the model does not explain much of the variation in ratings.
#
# Compared to dogs with no stage classification, those classified as `doggo` and `puppo` have significantly higher ratings.
#
# The confidence interval for `pupper` does not overlap with those of `doggo` or `puppo`, indicating that pupper have significantly lower ratings (based on the coefficients) than doggos and puppos.
#
# The confidence intervals for `doggo`, `floofer`, and `puppo` all overlap, indicating no significant difference between these groups. This is also true for `floofer` and `pupper`.
# What are dogs most frequently mis-classified as by the neural network?
top10_miscl = rate_nooutl[rate_nooutl['p1_dog']==False]['p1'].value_counts()[0:10]
top10_miscl
ind3 = range(1,11)
plt.bar(ind3,top10_miscl)
plt.xticks(ind3, top10_miscl.index, rotation='vertical')
plt.xlabel('Thing')
plt.ylabel('Frequency')
plt.title('Figure 4. Top 10 Misclassifications of Dogs');
# Dogs are most frequently mis-classified as seat belts (?). Possibly due to presence of collar or leash.
#
# Other furry (teddy, hamster) and very dog-like (dingo) animals are also common.
#
# Does the neural network's confidence in its classification of the dogs as dogs correlate with dog rating?
corr_class = rate_nooutl[rate_nooutl['p1_dog']==True]
corr_class.plot(x='rating_numerator', y='p1_conf', kind='scatter')
plt.xlabel('Rating')
plt.ylabel('Pred. 1 Conf.')
plt.title('Figure 5. Prediction 1 Confidence for Dog Predictions as Function of Rating');
# There doesn't seem to be an obvious relationship between rating and the neural network's confidence in its first prediction when it predicts that the picture is a dog, although there may be more predictions near a confidence of 1.0 at higher ratings.
#
# Check this with linear regression:
mdl_ccl = sm.OLS(corr_class['p1_conf'],corr_class[['intercept','rating_numerator']])
ccl_res = mdl_ccl.fit()
ccl_res.summary()
# The linear regression does a significant increase in prediction confidence with rating, although the R-squared value is low (0.01), indicating the model does not explain much of the variation in ratings.
| wrangle_act.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
type(123)
type(123.0)
type("Solomon")
type(False)
a = "Evelyn"
print(a)
type(a)
b = 1 + 12
print(b)
type(b)
type("b")
c = False
print(c)
type(c)
d = 123.0 + 123
type(d)
| 4. Data Types.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="yizCV24kz9qQ" colab_type="text"
# #Baseline rhythm generation model
# - modified based on Hungyi-Lee's HW8 structure: https://colab.research.google.com/drive/11iwJbQv9iScRo6kGP7YfyHaaorlHhzMT
# + id="8ZtzTW0ipIxR" colab_type="code" colab={}
import torch
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
import torch.utils.data as data
import torch.utils.data.sampler as sampler
import torchvision
from torchvision import datasets, transforms
import numpy as np
import sys
import os
import random
import json
import re
import pickle
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
# + id="tqcT8J2Jwcgk" colab_type="code" colab={}
class LabelTransform(object):
def __init__(self, size, pad):
self.size = size
self.pad = pad
def __call__(self, label):
label = np.pad(label, (0, (self.size - label.shape[0])), mode='constant', constant_values=self.pad)
return label
# + id="mg4hW-oBwckD" colab_type="code" colab={}
class RhythmDataset(data.Dataset):
def __init__(self, root, max_output_len, set_name):
self.root = root
self.max_output_len = max_output_len
self.word2int, self.int2word = self.get_dictionary()
# 載入資料
self.data = []
with open(os.path.join(self.root, f'{set_name}'), "rb") as f:
self.data=pickle.load(f)
print (f'{set_name} dataset size: {len(self.data)}')
self.vocab_size = len(self.word2int)
self.transform = LabelTransform(max_output_len, self.word2int['<PAD>'])
def get_dictionary(self):
# 載入字典
with open(os.path.join(self.root+'/rhythm_dict', f'vocab_word2int.json'), "r") as f:
word2int = json.load(f)
with open(os.path.join(self.root+'/rhythm_dict', f'vocab_int2word.json'), "r") as f:
int2word = json.load(f)
return word2int, int2word
def __len__(self):
return len(self.data)
def __getitem__(self, Index, pad=False):
global device
sentence = self.data[Index]
sentence_idx = []
for word in sentence:
if (word in self.word2int.keys()):
sentence_idx.append(self.word2int[word])
else:#如果遇到不会的单词就赋给不知道
sentence_idx.append(self.word2int["<UNK>"])
if pad:
sentence_idx = np.asarray(sentence_idx)
sentence_idx = self.transform(sentence_idx)
sentence_idx = torch.LongTensor(sentence_idx).to(device)
target = sentence_idx.clone()
return sentence_idx, target
# + id="XqmiDklbwvQR" colab_type="code" colab={}
class Encoder(nn.Module):
def __init__(self, vocab_size, emb_dim, hid_dim, n_layers, dropout):
super().__init__()
self.embedding = nn.Embedding(vocab_size, emb_dim)
self.hid_dim = hid_dim
self.n_layers = n_layers
self.rnn = nn.GRU(emb_dim, hid_dim, n_layers, dropout=dropout, batch_first=True, bidirectional=True)
self.dropout = nn.Dropout(dropout)
def forward(self, input):
# input: [batch_size, sequence_len]
embedding = self.embedding(input)
outputs, hidden = self.rnn(self.dropout(embedding))
# outputs: [batch_size, sequence_len, hid_dim * directions]
# hidden: [num_layers * directions, batch_size , hid_dim]
return outputs, hidden
def get_embedding(self, input):
# input: [batch_size, sequence_len]
embedding = self.embedding(input)
return embedding
# + id="1tRBlSUgw7-t" colab_type="code" colab={}
class Decoder(nn.Module):
def __init__(self, vocab_size, emb_dim, hid_dim, n_layers, dropout):
super().__init__()
self.vocab_size = vocab_size
self.hid_dim = hid_dim * 2
self.n_layers = n_layers
self.embedding = nn.Embedding(vocab_size, config.emb_dim)
self.input_dim = emb_dim
self.rnn = nn.GRU(self.input_dim, self.hid_dim, self.n_layers, dropout = dropout, batch_first=True)
self.embedding2vocab1 = nn.Linear(self.hid_dim, self.hid_dim * 2)
self.embedding2vocab2 = nn.Linear(self.hid_dim * 2, self.hid_dim * 4)
self.embedding2vocab3 = nn.Linear(self.hid_dim * 4, self.vocab_size)
self.dropout = nn.Dropout(dropout)
def forward(self, input, hidden, encoder_outputs):
# input: [batch_size, 1]
# hidden: [batch_size, n_layers * directions, hid_dim]
# Decoder's directions=1
input = input.unsqueeze(1)
embedded = self.dropout(self.embedding(input))
# embedded: [batch_size, 1, emb_dim]
output, hidden = self.rnn(embedded, hidden)
# output: [batch_size, 1, hid_dim]
# hidden: [num_layers, batch_size, hid_dim]
output = self.embedding2vocab1(output.squeeze(1))
output = self.embedding2vocab2(output)
prediction = self.embedding2vocab3(output)
# prediction: [batch_size, vocab_size]
return prediction, hidden
def get_embedding(self, input):
# input: [batch_size, sequence_len]
embedding = self.embedding(input)
return embedding
# + id="RC4ZcOq2xuBz" colab_type="code" colab={}
class Seq2Seq(nn.Module):
def __init__(self, encoder, decoder, device):
super().__init__()
self.encoder = encoder
self.decoder = decoder
self.device = device
assert encoder.n_layers == decoder.n_layers, \
"Encoder and decoder must have equal number of layers!"
def forward(self, input, target, teacher_forcing_ratio):
# input: [batch_size, input_len]
# target: [batch_size, target_len]
batch_size = target.shape[0]
target_len = target.shape[1]
vocab_size = self.decoder.vocab_size
outputs = torch.zeros(batch_size, target_len, vocab_size).to(self.device)
encoder_outputs, hidden = self.encoder(input)
# hidden = [num_layers * directions, batch_size, hid_dim] --> [num_layers, directions, batch_size, hid_dim]
hidden = hidden.view(self.encoder.n_layers, 2, batch_size, -1)
hidden = torch.cat((hidden[:, -2, :, :], hidden[:, -1, :, :]), dim=2)
# <BOS> token
input = target[:, 0]
preds = []
for t in range(1, target_len):
output, hidden = self.decoder(input, hidden, encoder_outputs)
outputs[:, t] = output
teacher_force = random.random() <= teacher_forcing_ratio
top1 = output.argmax(1)
input = target[:, t] if teacher_force and t < target_len else top1
preds.append(top1.unsqueeze(1))
preds = torch.cat(preds, 1)
#print(preds)
return outputs, preds
def inference(self, input, target):
# TODO: Beam Search
batch_size = input.shape[0]
input_len = input.shape[1]
vocab_size = self.decoder.vocab_size
#print('input_len=',input_len)
outputs = torch.zeros(batch_size, input_len, vocab_size).to(self.device)
encoder_outputs, hidden = self.encoder(input)
hidden = hidden.view(self.encoder.n_layers, 2, batch_size, -1)
hidden = torch.cat((hidden[:, -2, :, :], hidden[:, -1, :, :]), dim=2)
input = target[:, 0]
preds = []
for t in range(1, input_len):
output, hidden = self.decoder(input, hidden, encoder_outputs)
outputs[:, t] = output
top1 = output.argmax(1)
input = top1
preds.append(top1.unsqueeze(1))
preds = torch.cat(preds, 1)
#print(preds)
return outputs, preds
# + id="Uo_SfO9iwvtG" colab_type="code" colab={}
def build_model(config, vocab_size):
encoder = Encoder(vocab_size, config.emb_dim, config.hid_dim, config.n_layers, config.dropout)
decoder = Decoder(vocab_size, config.emb_dim, config.hid_dim, config.n_layers, config.dropout)
model = Seq2Seq(encoder, decoder, device)
print(model)
optimizer = torch.optim.Adam(model.parameters(), lr=config.learning_rate)
print(optimizer)
if config.load_model:
model = load_model(model, config.load_model_path)
model = model.to(device)
return model, optimizer
def save_model(model, optimizer, store_model_path, step):
torch.save(model.state_dict(), f'{store_model_path}/model_{step}.ckpt')
return
def load_model(model, load_model_path):
print(f'Load model from {load_model_path}')
model.load_state_dict(torch.load(f'{load_model_path}.ckpt'))
return model
# + id="axXTd1Jdwvvm" colab_type="code" colab={}
def tokens2sentence(outputs, int2word):
sentences = []
for tokens in outputs:
sentence = []
for token in tokens:
word = int2word[str(int(token))]
if word == '<EOS>':
break
sentence.append(word)
sentences.append(sentence)
return sentences
# + id="WgINQTpLyvJR" colab_type="code" colab={}
import nltk
from nltk.translate.bleu_score import sentence_bleu
from nltk.translate.bleu_score import SmoothingFunction
def computebleu(sentences, targets):
score = 0
if len(sentences) < len(targets):
#print(sentences)
#print(targets)
to_add = len(targets)-len(sentences)
for i in range(to_add):
sentences.append('<PAD>')
else:
assert (len(sentences) == len(targets))
def cut_token(sentence):
tmp = []
for token in sentence:
if token == '<UNK>' or token.isdigit() or len(bytes(token[0], encoding='utf-8')) == 1:
tmp.append(token)
else:
tmp += [word for word in token]
return tmp
for sentence, target in zip(sentences, targets):
sentence = cut_token(sentence)
target = cut_token(target)
score += sentence_bleu([target], sentence, weights=(1, 0, 0, 0))
return score
# + id="2tBSGZOsy2Rw" colab_type="code" colab={}
def infinite_iter(data_loader):
it = iter(data_loader)
while True:
try:
ret = next(it)
yield ret
except StopIteration:
it = iter(data_loader)
# + id="ltp5-iA1y_6u" colab_type="code" colab={}
def schedule_sampling(step,summary_steps):
return 1-0.8*step/summary_steps
# + id="h4rar2KtzCZj" colab_type="code" colab={}
def train(model, optimizer, train_iter, loss_function, total_steps, summary_steps, train_dataset):
model.train()
model.zero_grad()
losses = []
loss_sum = 0.0
for step in range(summary_steps):
sources, targets = next(train_iter)
sources, targets = sources.to(device), targets.to(device)
outputs, preds = model(sources, targets, schedule_sampling(step,summary_steps))
outputs = outputs.reshape(-1, outputs.size(2))
targets = targets.reshape(-1)
loss = loss_function(outputs, targets)
optimizer.zero_grad()
loss.backward()
grad_norm = torch.nn.utils.clip_grad_norm_(model.parameters(), 1)
optimizer.step()
loss_sum += loss.item()
if (step + 1) % 5 == 0:
loss_sum = loss_sum / 5
print ("\r", "train [{}] loss: {:.3f}, Perplexity: {:.3f} ".format(total_steps + step + 1, loss_sum, np.exp(loss_sum)), end=" ")
losses.append(loss_sum)
loss_sum = 0.0
return model, optimizer, losses
# + id="t7Z-Ic4XzJ4t" colab_type="code" colab={}
def test(model, dataloader, loss_function):
model.eval()
loss_sum, bleu_score= 0.0, 0.0
n = 0
result = []
for sources, targets in dataloader:
sources, targets = sources.to(device), targets.to(device)
batch_size = sources.size(0)
outputs, preds = model.inference(sources, targets)
outputs = outputs.reshape(-1, outputs.size(2))
targets = targets.reshape(-1)
loss = loss_function(outputs, targets)
loss_sum += loss.item()
# result2text
targets = targets.view(batch_size, -1)
# print(preds.shape)
# print(targets.shape)
# assert 0
preds = tokens2sentence(preds, dataloader.dataset.int2word)
targets = tokens2sentence(targets, dataloader.dataset.int2word)
for pred, target in zip(preds, targets):
result.append((pred, target))
bleu_score += computebleu(preds, targets)
n += batch_size
return loss_sum / len(dataloader), bleu_score / n, result
# + id="eyeF1jjzzOuy" colab_type="code" colab={}
def train_process(config):
train_dataset = RhythmDataset(config.data_path, config.max_output_len, 'rhythm_pattern_list_all.data')
train_loader = data.DataLoader(train_dataset, batch_size=config.batch_size, shuffle=True)
train_iter = infinite_iter(train_loader)
val_dataset = RhythmDataset(config.data_path, config.max_output_len, 'rhythm_pattern_list_all.data')
val_loader = data.DataLoader(val_dataset, batch_size=1)
model, optimizer = build_model(config, train_dataset.vocab_size)
loss_function = nn.CrossEntropyLoss(ignore_index=0)
train_losses, val_losses, bleu_scores = [], [], []
total_steps = 0
while (total_steps < config.num_steps):
#train
model, optimizer, loss = train(model, optimizer, train_iter, loss_function, total_steps, config.summary_steps, train_dataset)
train_losses += loss
#test
val_loss, bleu_score, result = test(model, val_loader, loss_function)
val_losses.append(val_loss)
bleu_scores.append(bleu_score)
total_steps += config.summary_steps
print ("\r", "val [{}] loss: {:.3f}, Perplexity: {:.3f}, blue score: {:.3f} ".format(total_steps, val_loss, np.exp(val_loss), bleu_score))
#save
if total_steps % config.store_steps == 0 or total_steps >= config.num_steps:
save_model(model, optimizer, config.store_model_path, total_steps)
with open(f'{config.store_model_path}/output_{total_steps}.txt', 'w') as f:
for line in result:
print (line, file=f)
return train_losses, val_losses, bleu_scores
# + id="mDjTVG7dzeQY" colab_type="code" colab={}
def test_process(config):
test_dataset = RhythmDataset(config.data_path, config.max_output_len, 'rhythm_pattern_list_all.data')
test_loader = data.DataLoader(test_dataset, batch_size=1)
model, optimizer = build_model(config, test_dataset.vocab_size)
print ("Finish build model")
loss_function = nn.CrossEntropyLoss(ignore_index=0)
model.eval()
#test
test_loss, bleu_score, result = test(model, test_loader, loss_function)
#save
with open(f'{config.store_model_path}/test_output.txt', 'w') as f:
for line in result:
print (line, file=f)
return test_loss, bleu_score, result
# + id="mH5vUWb3zm42" colab_type="code" colab={}
class configurations(object):
def __init__(self):
self.batch_size = 1
self.emb_dim = 256
self.hid_dim = 512
self.n_layers = 3
self.dropout = 0.5
self.learning_rate = 0.00005
self.max_output_len = 100
self.num_steps = 9000
self.store_steps = 2000
self.summary_steps = 300
self.load_model = False
self.store_model_path = "/content/drive/My Drive/Colab Notebooks/music_GAN_rhythm_seed/model_seq2seq_baseline"
self.load_model_path = "/content/drive/My Drive/Colab Notebooks/music_GAN_rhythm_seed/model_seq2seq_baseline/model_"+str(12000)
self.data_path = "/content/drive/My Drive/Colab Notebooks/music_GAN_rhythm_seed/data_folder/"
# + id="GLwQFOzHzeab" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 818} executionInfo={"status": "ok", "timestamp": 1592572665976, "user_tz": -480, "elapsed": 850836, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13618470284889852279"}} outputId="9f229a5f-9a30-4c79-82ce-f21e01f4c598"
config = configurations()
print ('config:\n', vars(config))
train_losses, val_losses, bleu_scores = train_process(config)
# + id="3pFFO7IjIZIM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1592569441699, "user_tz": -480, "elapsed": 1745, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13618470284889852279"}} outputId="4c849306-0751-48af-8ede-a8c1a9ae769d"
train_dataset = RhythmDataset(config.data_path, config.max_output_len, 'rhythm_pattern_list_all.data')
train_loader = data.DataLoader(train_dataset, batch_size=config.batch_size, shuffle=True)
train_iter = infinite_iter(train_loader)
# + id="uialyivgIZ_L" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 70} executionInfo={"status": "ok", "timestamp": 1592569474630, "user_tz": -480, "elapsed": 1879, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13618470284889852279"}} outputId="261b62cd-6a35-435c-e70c-a7d23b4baeb2"
sources, targets = next(train_iter)
print(sources)
# + id="5gNPaTPfj3O_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 621} executionInfo={"status": "ok", "timestamp": 1592569987154, "user_tz": -480, "elapsed": 1141, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13618470284889852279"}} outputId="2c5f4d2b-9739-4c58-bd9c-8533993ae18c"
tokens2sentence(sources, train_loader.dataset.int2word)
| colab_pruned/model_seq2seq_baseline/rhythm_word_seq2seq_baseline_master.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Quantum Ensemble for Classification
#
# ## <NAME>
#
#
#
# Department of Computer Science and Engineering, University of Bologna, Italy
#
#
# [preprint arXiv:2007.01028](https://arxiv.org/abs/2007.01028)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Contribution
#
# * A novel quantum algorithm to perform ensemble classification
#
# * Superposition and Entanglement to generate a large number of different models
#
# * Large ensembles feasible with relatively small circuits
#
# * Experiments using qiskit
# + [markdown] slideshow={"slide_type": "slide"}
# # Classical Ensemble Learning
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Supervised Machine Learning as Aggregator of Functions
#
# Standard formulation for supervised ML:
#
# \begin{align}
# y = f(x; \Theta) + \epsilon.
# \end{align}
# where $x \in \mathbb{R}^p$ and $y$ is the target variable
#
# Objectives:
# * Useful approximation $\hat{f}(x; \cdot)$
# * Optimal set of parameters $\Theta$
#
# Alternative formulation:         $f(x;\beta, \theta) = \sum_{i=1}^{B} \beta_i g(x;\theta_i)$
#
# where $g(x; \cdot)$ is the base model and $\theta = \{ \theta_i\}_{i=1, \dots, B}$ and $\beta = \{ \beta_i\}_{i=1, \dots, B}$
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Ensemble Learning (Bagging)
#
# # + $g(x, \cdot)$ is a ML model (under/over-fitting)
# # + Fit the same model under different and independent training conditions
# # + Necessary and sufficient conditions: the base models **accurate** and **diverse**
#
#
# Ensemble prediction:
# \begin{align}
# f_{\text{ens}}(x) = \frac{1}{B}\sum_{i=1}^B \beta_i \hspace{0.2em} g(x; \theta_i)
# \end{align}
#
# where $g(x;\cdot)$ is the base model
#
# Approaches: *Bagging, Boosting, Randomisation, Stacking*
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Example of Bagging: Random Forest
#
# Bagging averages the predictions over a collection of bootstrap samples
#
# Random Forest uses a decision tree as based model
#
# \begin{align}
# \hat{f}^B_{\text{rf}}(x) = \frac{1}{B} \sum_{i=1}^B g(x; \theta_i)
# =
# \frac{1}{B} \sum_{i=1}^B \left[ \sum_{m=1}^{M} c^{(i)}_{m} \boldsymbol{I}_{\left(x\in R^{(i)}_m \right)}\right]
# \end{align}
#
# $\beta_i = \frac{1}{B}$ for $i=1, \dots, B$
#
# $\theta_i$ characterises the $b$th tree in terms of split variables, cutpoints, terminal-node values
#
# <br></br>
#
# *<NAME>, <NAME>, and <NAME>. The elements of statistical learning. Vol. 1. No. 10. New York: Springer series in statistics, 2001*
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Time Complexity of Classical Ensemble (Bagging)
# Classical Ensemble scaling:
#
# \begin{align*}
# \underbrace{\mathcal{O}\left(B \times N^{a}p^{b}\right)}_\text{Training} + \underbrace{\mathcal{O}(Bp)}_\text{Testing} \qquad a, b \geq 1,
# \end{align*}
#
# where $N$ is the number of training points and $p$ is the number of features
#
#
# - Linearity in $B$ (ensemble size)
#
# - Multiplicative cost of the classifier
# + [markdown] slideshow={"slide_type": "skip"}
# $$\newcommand{\ket}[1]{\left|{#1}\right\rangle}$$
# $$\newcommand{\bra}[1]{\left\langle{#1}\right|}$$
# $$\newcommand{\braket}[1]{\left\langle{#1}\right\rangle}$$
# + [markdown] slideshow={"slide_type": "slide"}
# # Quantum Algorithm for Ensemble Learning
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Quantum Computing Properties
# Superposition
# # + $n$ qubits $\xrightarrow{}$ $2^n$ states $\implies$ $n+1$ qubits $\xrightarrow{}$ $2^{n+1}$ states
# # + single model/prediction $\rightarrow$ single quantum state
#
# Entanglement
# # + Each base model $g(x; \cdot)$ is strictly associated to a single weight
#
# Interference
# # + Train and Test "interact" each other to compute test prediction
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Quantum Ensemble: requirements
#
# - Quantum gate for sampling:                     $\ket{x,y} \xrightarrow{V_b}\ket{x_b,y_b}$
#
# - Quantum classifier $F$ (working via interference):   $\ket{x_b,y_b}\ket{\tilde{x}}\ket{0}\xrightarrow{F} \ket{x_b,y_b}\ket{\tilde{x}}\ket{f_b}$
#
# Input:
# - $d$-qubits $control$ register
# - $data$ register encodes the training set
# - $test$ register encodes the test set
#
# Output: the average of $B=2^d$ different predictions of the same classifier
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Quantum Ensemble using Bagging strategy
#
# # + **(Step 1) State Preparation**
#
# # + **(Step 2) Sampling in Superposition**
#
# # + **(Step 3) Learning via Interference**
#
# # + **(Step 4) Measurement**
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Step 1: State Preparation
# * The training set is encoded into the $data$ quantum register
#
# * The $control$ register is initialised into uniform superposition
#
# \begin{align}
# \ket{\Phi_0} & = \big(W \otimes S_{(x,y)}\big)\overset{d}{\underset{j=1}{\otimes}} \ket{0} \otimes \ket{0} \nonumber \\
# & = \overset{d}{\underset{j=1}{\otimes}} \left(\frac{\ket{0}+\ket{1}}{\sqrt{2}}\right) \otimes \ket{x,y} = \frac{1}{\sqrt{2^d}}\sum_{b=1}^{2^d} \ket{b} \otimes \ket{x,y},
# \end{align}
#
# $S_{(x,y)}$ is the unitary that encodes data into a quantum state
#
# $W=H^{\otimes d}$ is the Walsh-Hadamard gate
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Step 2: Sampling in Superposition
#
# $2^d$ different transformations (*quantum trajectories*) of the input in superposition
#
# Entanglement between $data$ and $control$ registers
# \begin{align}
# \ket{\Phi_{d}}
# = & \frac{1}{\sqrt{2^d}} \sum_{b = 1}^{2^{d}} \ket{b} V_b\ket{x,y} = \frac{1}{\sqrt{2^d}} \sum_{b = 1}^{2^{d}} \ket{b}\ket{x_b,y_b}
# \end{align}
#
# Each $V_b$ represents a single "quantum trajectory"
#
#
# \begin{equation}\label{eq:single_subsample}
# \ket{x,y} \xrightarrow{V_{b}} \ket{x_b, y_b} %\text{where} \quad U_{t_1}U_{t_2}\dots U_{t_q} = V_t
# \end{equation}
#
#
# $V_b$ transforms the original training set to obtain a random sub-sample of it
#
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Step 3: Learning via Interference
#
# The *test* register is initialised to encode the test set, considering also a set of qubits to store the final predictions ($target$ sub-register):
#
# \begin{align}
# (S_{\tilde{x}} \otimes \mathbb{1}) \ket{0}\ket{0} =\ket{\tilde{x}}\ket{0}
# \end{align}
#
# The $data$ and $test$ registers interact via interference to compute the target variable:
#
# \begin{align}\label{eq:classification via interference}
# \ket{\Phi_{f}} & = \Big(\mathbb{1}^{\otimes d} \otimes F \Big) \ket{\Phi_d} = (\mathbb{1}^{\otimes d} \otimes F )\Bigg[\frac{1}{\sqrt{2^d}}\sum_{b=1}^{2^d} \ket{b} \ket{x_b, y_b}\Bigg] \otimes
# \ket{\tilde{x}}
# \ket{0} \nonumber \\
# & = \frac{1}{\sqrt{2^d}}\sum_{b=1}^{2^d} \ket{b} \ket{x_b, y_b} \ket{\tilde{x}}\ket{f_b}
# # %CV_{t}\ket{\psi} = CV_{t}\ket{x, y}\ket{0} \ket{x^{(\text{test})}, 0} = C \ket{x, y}\ket{x_t, x_y} \ket{x^{(\text{test})}, 0} = \ket{x, y}\ket{x_t, x_t} \ket{x^{(\text{test})}, \tilde{y}} %= \ket{x, y, D_t, x^{(\text{test})}, f(x^{(\text{test})}|x_t,y_t)}
# \end{align}
#
# where $\tilde{y} = f_{b}$ is computed through the quantum gate $F$
#
# A single execution of $F$ allows propagating the use of the classifier to all $2^d$ sub-samples
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Step 4: Measurement
# $\newcommand{\braket}[1]{\left\langle{#1}\right\rangle}$
# The expectation $measurement$ on the target qubit provides a sum of expectation values:
#
# \begin{align}
# \left\langle M \right\rangle & = \braket{\Phi_f|\mathbb{1}^{\otimes d} \otimes \mathbb{1} \otimes \mathbb{1} \otimes M|\Phi_f} \nonumber \\
# & =
# \frac{1}{2^d}\sum_{b=1}^{2^d}\braket{b|b} \otimes \braket{(x_b,y_b)|(x_b,y_b)} \otimes\braket{\tilde{x}|\tilde{x}} \otimes\braket{f_b|M|f_b} \nonumber \\
# & = \frac{1}{2^d}\sum_{b=1}^{2^d}\left\langle M_b \right\rangle = \frac{1}{B} \sum_{b=1}^B \hat{f}_b = \hat{f}_{bag}(\tilde{x}|x,y)
# \end{align}
# E.g.
#
# $$f_{bag} = \sqrt{a_0}\ket{0}+\sqrt{a_1}\ket{1} $$
# + [markdown] slideshow={"slide_type": "slide"}
# # Quantum Circuit
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Quantum Circuit for Ensemble
#
# <br></br>
#
# <center><img src="IMG/circuit.JPG" width="100%" height="100%"/></center>
# + [markdown] slideshow={"slide_type": "subslide"}
# The generic $i$-th step involves the following transformations:
#
# \begin{align}
# \ket{\Phi_{i,1}} = \frac{1}{\sqrt{2}}\Big(\ket{0}\ket{x,y} + \ket{1} U_{(i,1)}\ket{x,y} \Big)
# \end{align}
#
#
# \begin{align}
# \ket{\Phi_{i,2}} = & ( X \otimes \mathbb{1}) \ket{\Phi_{i,1}}
# = \frac{1}{\sqrt{2}}\Big(\ket{1}\ket{x,y} + \ket{0} U_{(i,1)}\ket{x,y} \Big)
# \end{align}
#
#
# \begin{align}
# \ket{\Phi_{i}}= \frac{1}{\sqrt{2}}\Big(\ket{1}U_{(i,2)}\ket{x,y} + \ket{0} U_{(i,1)}\ket{x,y} \Big)
# \end{align}
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Sampling in superposition steps
# The output of the State preparation is the following:
#     $\ket{\Phi_0} = \overset{d}{\underset{i=1}{\otimes}} \left(\frac{\ket{0}+\ket{1}}{\sqrt{2}}\right) \otimes \ket{x,y}$
# + [markdown] slideshow={"slide_type": "fragment"}
# ### Sampling in superposition steps ($1st$ step)
#
# \begin{align}
# \ket{\Phi_{1}} = \overset{d-1}{\underset{i=1}{\otimes}} \left(\frac{\ket{0}+\ket{1}}{\sqrt{2}}\right) \otimes \frac{1}{\sqrt{2}}\Big(\ket{1}U_{(1,2)}\ket{x,y} + \ket{0} U_{(1,1)}\ket{x,y} \Big) \end{align}
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Sampling in superposition steps ($2nd$ step)
#
# \begin{align}
# \ket{\Phi_{2}}
# = \overset{d-2}{\underset{i=1}{\otimes}} \left(\frac{\ket{0}+\ket{1}}{\sqrt{2}}\right) \otimes \frac{1}{2}\Big[
# \hspace{.2em} &\ket{00} U_{(2,1)}U_{(1,1)}\ket{x,y} +
# %\nonumber \\ + &
# \ket{01} U_{(2,1)}U_{(1,2)}\ket{x,y} +
# \nonumber \\ + &
# \ket{10} U_{(2,2)}U_{(1,1)}\ket{x,y} +
# %\nonumber \\ + &
# \ket{11} U_{(2,2)}U_{(1,2)}\ket{x,y}
# \Big] \nonumber \\
# \hspace{-10.75em} = \frac{1}{\sqrt{4}} \sum_{b=1}^{4} \ket{b} V_b\ket{x,y}
# \end{align}
#
# where each $V_b$ is the product of $2$ unitaries $U_{(i,j)}$ for $j=1,2$ and $i=1,2$
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Sampling in Superposition ($3rd$ step)
#
# \begin{align}
# \ket{\Phi_{3}}
# = \overset{d-3}{\underset{i=1}{\otimes}} \left(\frac{\ket{0}+\ket{1}}{\sqrt{2}}\right) \otimes \frac{1}{\sqrt{8}}\Big[
# & \ket{000} U_{(3,1)}U_{(2,1)}U_{(1,1)}\ket{x,y} + \ket{001}U_{(3,1)}U_{(2,1)}U_{(1,2)}\ket{x,y}
# \nonumber \\ + &
# \ket{010} U_{(3,1)}U_{(2,2)}U_{(1,1)}\ket{x,y} + \ket{011}U_{(3,1)}U_{(2,2)}U_{(1,2)}\ket{x,y}
# \nonumber \\ + &
# \ket{100} U_{(3,2)}U_{(2,1)}U_{(1,1)}\ket{x,y} +
# \ket{101} U_{(3,2)}U_{(2,1)}U_{(1,2)}\ket{x,y}
# \nonumber \\ + &
# \ket{110} U_{(3,2)}U_{(2,2)}U_{(1,1)}\ket{x,y} + \ket{111}U_{(3,2)}U_{(2,2)}U_{(1,2)}\ket{x,y}
# \Big] \nonumber \\
# & \hspace{-11.5em} = \frac{1}{\sqrt{2^3}} \sum_{b=1}^{8} \ket{b} V_b\ket{x,y}
# \end{align}
#
# where each $V_b$ is the product of $2$ unitaries $U_{(i,j)}$ for $j=1,2$ and $i=1,2,3$
#
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Sampling in superposition after d steps:
#
#
# \begin{align}
# \ket{\Phi_{d}}
# & = \frac{1}{\sqrt{2^d}} \sum_{b = 1}^{2^{d}} \ket{b} V_b\ket{x,y} = \frac{1}{\sqrt{2^d}} \sum_{b = 1}^{2^{d}} \ket{b}\ket{x_b,y_b}
# \end{align}
#
# where each $V_b$ is the product of $d$ unitaries $U_{(i,j)}$ for $i=1, \cdots, d$ and $j=1,2$.
#
#
#
# **$2^d$ different transformations** (*quantum trajectories*) of the input in superposition in only **$d$ steps**
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Time Complexity
#
# Classical Ensemble scaling:
#
# \begin{align*}
# \underbrace{\mathcal{O}\left(B \times N^{a}p^{b}\right)}_\text{Training} + \underbrace{\mathcal{O}(Bp)}_\text{Testing} \qquad a, b \geq 1,
# \end{align*}
#
# Quantum Ensemble scaling:
#
# \begin{align*}
# \mathcal{O}(\underbrace{d \times 2 cost({CU_{(i,j)}})}_\text{Sampling in Sup.} + \underbrace{cost(F)}_\text{Learning} )
# %\Big)
# \end{align*}
#
# $B =2^d$ is the ensemble size, $N$ is the number of obs., and $p$ is the number of features
#
# Additive cost of the classifier (instead of multiplicative)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Aggregation Strategy
#
# Majority voting vs Simple averaging
#
# \begin{align}
# f_{\text{avg}}^{(i)}(x) = \frac{1}{B}\sum_{b=1}^B f_b^{(i)}(x),
# \end{align}
#
# $B$ is the ensemble size and $f_b^{(i)}(x)$ is the probability for $x$ to be classified in the $i$-th class provided by the $b$-th classifier
#
# Performance:
#
# \begin{align}\label{perf_ens}
# E_{\text{ens}} = \frac{1+\rho(B-1)}{B}E_{\text{model}}
# \end{align}
# where $E_{\text{model}}$ is the expected error of the single models and $\rho$ is the average correlation among their errors
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Theoretical Performance
# + slideshow={"slide_type": "fragment"}
import sys
sys.path.insert(1, '../')
from Utils import *
# parameters
d = np.arange(6) # Number of control qubits
B = 2**d # Number of models: ensemble size
errs = np.array([0.1, 0.2, 0.3, 0.4]) # base model error
ro = [0.00, 0.25, 0.5] # correlation amontg errors
colors = sns.diverging_palette(220, 20, n=6)
# create figure
fig, ax = plt.subplots(1, figsize=(8,5))
#for each possible ensemble size
for i in np.arange(len(errs)):
err = errs[i]
color = colors[i]
mean = ((1+ro[1]*(B-1))/B)*err
lower = mean - ((1+ro[0]*(B-1))/B)*err
upper = ((1 + ro[2] * (B - 1)) / B) * err - mean
ax.plot(d, mean, lw=2, label = str(int(err*100))+'%', color=color)
ax.fill_between(d, mean + upper, mean - lower, facecolor=color, alpha=0.5)
# title
ax.set_title('Theoretical performance of Bagging Ensemble', size=14).set_position([.5, 1.05])
# legend
ax.legend(loc='upper right', title = '$E_{model}$', prop=dict(size=12))
# label of x-axis
ax.set_xlabel(r'Number of control qubits $(d=log_2B)$', size=14)
## label of x-axis
ax.set_ylabel('Prediction Error ($E_{ens}$)', size=14)
# set grid
ax.grid(alpha=.3)
# + [markdown] slideshow={"slide_type": "skip"}
# ## Boosting and Randomisation
#
# Quantum Boosting:
# \begin{align} \label{eq:quantum_boosting}
# \ket{\Phi_{f}}
# = \frac{1}{\sqrt{2^d}}\sum_{b=1}^{2^d} \alpha_b \ket{b} \ket{f_b}
# \end{align}
#
# where the contribution of $f_b$ to the ensemble is weighted by $\alpha_b$
# + [markdown] slideshow={"slide_type": "skip"}
# Quantum Randomisation:
#
# \begin{align}
# \ket{\Phi_{f}}
# =\frac{1}{\sqrt{2^d}}\sum_{b=1}^{2^d}\ket{b} G_b \ket{x,y}\ket{\tilde{x}}\ket{0} = \frac{1}{\sqrt{2^d}}\sum_{b=1}^{2^d} \ket{b} \ket{x,y}\ket{\tilde{x}}\ket{f_b}
# \end{align}
#
# where $G_b=V_b F$ is the quantum classifier
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
# ## (Theoretical) Advantages and Limitations
#
#
# # + <p>$2^d$ different transformations of the input in $d$ steps: exponential speed-up <span style="color:green">(+)</span></p>
#
# # + Quantum parallelism allows propagation of $F$ to all quantum trajectories <span style="color:green">(+)</span></p>
#
# # + Independence from the encoding strategy chosen for data <span style="color:green">(+)</span></p>
#
# # + Measurement is limited to a subset of qubits <span style="color:green">(+)</span></p>
#
# <br></br>
#
# - Evaluation of the single functions implies exponential cost <span style="color:red">(-)</span></p>
#
# - Additional cost of state preparation <span style="color:red">(-)</span></p>
| presentation/1. Theory.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 学習に関するテクニック
# ## パラメータの更新
# ### SGD(確率的勾配降下法 stochastic gradient descent)
# パラメータの勾配を使って、勾配方向にパラメータを更新するというステップを繰り返す
class SGD:
def __init__(self, lr=0.01):
self.lr = lr
def update(self, params, grads):
for key in params.keys():
params[key] -= self.lr * grads[key]
# ### Momentum
# 物体が勾配方向に力を受け、その力によって物体の速度が加算される物理法則、イメージは坂を転がるボール
class Momentum:
def __init__(self, lr=0.01, momentum=0.9):
self.lr = lr
self.momentum = momentum
self.v = None
def update(self, params, grads):
if self.v is None:
self.v = {}
for key, val in params.items():
self.v[key] = np.zeros_like(val)
for key in params.keys():
self.v[key] = self.momentum*self.v[key] - self.lr*grads[key]
params[key] += self.v[key]
# ### AdaGrad
# 学習率の減衰(learning rate decay)を用いる、パラメータの要素ごとに適応的に学習係数を調整しながら学習を行う。
# 具体的にはパラメータの要素の中で大きく更新された要素は、学習係数が小さくなっていく
#
# AdaはAdaptiveの意
class AdaGrad:
def __init__(self, lr=0.01):
self.lr = lr
self.h = None
def update(self, params, grads):
if self.h is None:
self.h = {}
for key, val in params.items():
self.h[key] = np.zeros_like(val)
for key in params.keys():
self.h[key] += grads[key] * grads[key]
params[key] -= self.lr * grads[key] / (np.sqrt(self.h[key]) + 1e-7) # 0除算を防ぐため分母にεを加算
# ## アクティベーション分布
# アクティベーションに偏りがあると、表現力に問題があるということになる。
# +
import numpy as np
import matplotlib.pyplot as plt
def sigmoid(x):
return 1/(1+np.exp(-x))
x = np.random.randn(1000, 100)
node_num = 100
hidden_layer_size = 5
activations = {}
for i in range(hidden_layer_size):
if i != 0:
x = activations[i-1]
w = np.random.randn(node_num, node_num) * 1
z = np.dot(x, w)
a = sigmoid(z)
activations[i] = a
# -
plt.figure(figsize=(20,5)) #横幅,縦幅
for i, a in activations.items():
plt.subplot(1, len(activations), i+1)
plt.title(str(i+1) + "-layer")
plt.hist(a.flatten(), 30, range=(0, 1))
plt.show()
# ### Xavierの初期値
x = np.random.randn(1000, 100)
for i in range(hidden_layer_size):
if i != 0:
x = activations[i-1]
w = np.random.randn(node_num, node_num) /np.sqrt(node_num)
z = np.dot(x, w)
a = sigmoid(z)
activations[i] = a
plt.figure(figsize=(20,5)) #横幅,縦幅
for i, a in activations.items():
plt.subplot(1, len(activations), i+1)
plt.title(str(i+1) + "-layer")
plt.hist(a.flatten(), 30, range=(0, 1))
plt.show()
# ### ReLU + Heの初期値
# +
import numpy as np
import matplotlib.pyplot as plt
def Relu(x):
return np.maximum(0, x)
x = np.random.randn(1000, 100)
node_num = 100
hidden_layer_size = 5
activations = {}
for i in range(hidden_layer_size):
if i != 0:
x = activations[i-1]
w = np.random.randn(node_num, node_num) * np.sqrt(2.0/node_num)
z = np.dot(x, w)
a = Relu(z)
activations[i] = a
# -
plt.figure(figsize=(20,5)) #横幅,縦幅
for i, a in activations.items():
plt.subplot(1, len(activations), i+1)
plt.title(str(i+1) + "-layer")
plt.hist(a.flatten(), 30, range=(0, 1))
plt.show()
# ## Dropout
class Dropout:
def __init__(self, dropout_ratio = 0.5):
self.dropout_ratio = dropout_ratio
self.mask = None
def forward(self, x, train_flg = True):
if train_flg:
self.mask = np.random.rand(*x.shape) > self.dropout_ratio
return x * self.mask
else:
return x * (1.0 - self.dropout_ratio)
def backward(self, dout):
return dout * self.mask
| other/techniques.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
import numpy as np
import pandas as pd
import allel
import matplotlib.pyplot as plt
from sklearn.decomposition import PCA
# # Import variants VCF file & Data Cleaning
vcf = allel.read_vcf("data/raw/1349 sample and all 253k unfiltered SNPs.vcf", )
variants = np.char.array(vcf["variants/CHROM"].astype(str)) + ":" + np.char.array(vcf["variants/POS"].astype(str))
vcf_arr = vcf["calldata/GT"].astype("float")
vcf_arr[vcf_arr == -1] = np.nan
mutations = vcf_arr
# mutations = np.abs(mutations)
mutations = mutations.sum(axis=2)
mutations = mutations.T
mutations.shape
mutations_df = pd.DataFrame(data=mutations, index=vcf["samples"], columns=variants)
mutations_df.shape
mutations_df.dropna(axis=1, how="any", thresh=800, inplace=True)
mutations_df.shape
mutations_df.dropna(axis=0, how="any", thresh=200000, inplace=True)
mutations_df.fillna(value=0, inplace=True)
mutations_df.isna().sum().sum()
# # Subset patients
samples_phenotypes = pd.read_table("data/raw/Sample metadata.csv", sep=",")
samples_phenotypes.set_index("ID", inplace=True)
good_samples = pd.read_table("data/interim/samples_metadata.csv", sep=",")
good_samples.set_index("ID", inplace=True)
good_samples = good_samples[good_samples["SRC"] != "LGS"]
good_samples = good_samples[good_samples["SRC"] != "D2"]
good_samples = good_samples[good_samples["SRC"] != "U2"]
SLE_samples = good_samples[good_samples["SLE"] == 1]
SLE_samples.shape
hla_protein_samples = pd.Index(['55062', '56104', '34903', '16820', '41060', '54687', '44119', '48523',
'33287', '14947', '21560', '87483', '42335', '30146', '28289', '40007'])
highdsdna_samples = pd.Index(["32588", "55062"]) # High dsDNA
lowdsdna_samples = pd.Index(["54687", "16820"]) # low dsDNA
validation_samples = highdsdna_samples.append(lowdsdna_samples)
validation_samples.shape
training_samples = SLE_samples.index[~SLE_samples.index.isin(validation_samples)]
training_samples.shape
# # Filtered data
# +
# filtered_df = mutations_df
training_df = mutations_df.filter(items=training_samples, axis=0)
training_df.shape
validation_df = mutations_df.filter(items=validation_samples, axis=0)
validation_df.shape
# -
# # Use PCA feature selction model
from src.features.PCA_feature_selection import PCA_Variants2Gene_FeatureSelection
pca_v2g = PCA_Variants2Gene_FeatureSelection(variants_genes_path="data/interim/variants_top56_genes.csv",
variance_threshold=0.80)
training_X1 = pca_v2g.fit_transform(training_df)
training_X1.index.name = "patient_id"
validation_X1 = pca_v2g.transform(validation_df)
validation_X1.index.name = "patient_id"
validation_X1.shape
training_gene_scores = pca_v2g.get_gene_variability_score(pd.concat([training_df, validation_df], axis=0))
training_gene_scores
training_gene_scores.to_csv("data/processed/allsamples_gene_variability_scores.csv")
# +
# training_X1.to_csv("data/processed/training_pca_projs.csv")
# validation_X1.to_csv("data/processed/validation_pca_projs.csv")
# -
# # Gene - mutations
variant_genes = pd.read_table("../data/raw/216k Variants with gene name.csv", sep=",")
variant_genes = variant_genes[variant_genes["Gene(s)"] != "?"]
# variant_genes = variant_genes.filter(items=["Gene(s)", "Variant ID"])
variant_genes["Position"] = variant_genes["Position"].astype("object")
genes_56 = variant_genes["Gene(s)"].value_counts()[:56]
# # Compute PCA projections of all SNP data
mutation_gene_dict = pca_v2g.get_mutations_by_gene_dict(validation_df)
genes = list(mutation_gene_dict.keys())
num_top_components = [mutation_gene_dict[gene].shape[1] for gene in genes]
plt.figure(figsize=(12, 4), dpi=150)
plt.title("Number of Mutation Sites by Gene")
plt.xlabel("Genes")
plt.xticks(rotation=90)
plt.ylabel("# of Mutations")
plt.bar(x=genes, height=num_top_components)
training_X1.shape
genes = list(pca_v2g.top_k_PC_by_gene.keys())
num_top_components = [pca_v2g.top_k_PC_by_gene[gene] for gene in genes]
print(np.sum(num_top_components))
plt.figure(figsize=(12, 3), dpi=150)
plt.title("Top-k PCA components selected with 90% variance threshold, split by genes, only SLE")
plt.xlabel("Genes")
plt.xticks(rotation=90)
plt.ylabel("# of PCA components selected")
plt.bar(x=genes, height=num_top_components)
patient_gene_proj_scores.to_csv("../data/interim/patient_gene_proj_scores.csv")
# # Select variants by percentile
top_variants_by_gene = {}
for gene in mutations_by_gene.keys():
top_variants_by_gene[gene] = select_top_variants(mutations_by_gene[gene], var_threshold=0.80, coef_percentile=70)
print(gene, mutations_by_gene[gene].shape[1], top_variants_by_gene[gene].shape[0])
all_top_variants = []
for gene in top_variants_by_gene.keys():
all_top_variants.extend(top_variants_by_gene[gene])
len(all_top_variants)
# # Concatenate all selected variant into one DataFrame
mutations_df = mutations_df.loc[:,~mutations_df.columns.duplicated()]
mutations_top_variants_df = mutations_df.filter(items=all_top_variants)
mutations_top_variants_df
mutations_top_variants_df.to_csv("../data/interim/mutations_top_variants_80_variance.csv")
# # Graph the eigenvalues
import matplotlib.pyplot as plt
# +
plt.figure(figsize=(10, 7))
plt.bar(x=range(pca.singular_values_.shape[0]),
height=pca.singular_values_, )
plt.ylabel('log of singular values')
plt.xlabel('Principal components')
plt.title('Elbow plot from PCA of mutations data')
plt.xlim(-1, 45)
# plt.ylim(0, 10)
plt.show()
# -
| variants_PCA_feature_selection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Welcome to the Intro to Astronomy Research Gaia Data Release 2 (DR2) Tutorial.
#
# #### Written by <NAME>, 2018
#
# ### In this tutorial, you will:
# - learn about the Astronomical Data Query Language (ADQL)
# - use the Gaia DR2 Database and the NASA Exoplanet Archive to get Gaia parameters for the 10,000 closest stars
# - plot a color-magnitude diagram
#
# ### Notes:
# - This tutorial is challenging! If you spend more than 15 minutes stuck on a task, post about it on Piazza (you can post anonymously if you want). An instructor will help you out! Don't struggle needlessly.
# - Make sure you complete the pandas and matplotlib tutorials before attempting this tutorial.
# # Learn About ADQL
#
# - Navigate to the Gaia ADQL interface. First, go [here](https://gea.esac.esa.int/archive/). Click "Search," then click "Advanced (ADQL)" (in the top left corner of the page).
#
# - Read [this webpage](https://gea.esac.esa.int/archive-help/adql/index.html).
#
# - Read slides 6-17 in [this powerpoint](https://www.cosmos.esa.int/documents/915837/915858/ADQL_handson_slides.pdf/652b9120-a3fe-4857-b5eb-933b476687ad).
#
# Try out some of the commands for yourself in the Gaia ADQL search bar you opened in step 1. Using the buttons that pop up to the right of your query results, you can download the results or view them in your browser. Hover over the buttons to see what they do.
#
# Don't worry if you don't understand everything in the powerpoint! Pick out the 7 most important slides and summarize them for yourself.
#
# ** Note: wherever the powerpoint uses "gaiadr1.tgas_source," replace with "gaiadr2.gaia_source."
# # Use the Gaia DR2 Database and the NASA Exoplanet Archive to Get Gaia Parameters for the 10,000 Closest Stars
# ### Write an ADQL query to get parameters of the 10,000 closest stars. Your query should return the following parameters:
#
# - BP - RP color (bp_rp in the Gaia database)
# - absolute g-band photometric magnitude
# - distance
#
# <span style="color:red">ADQL QUERY COMMAND:</span>
#
# ```
# SELECT TOP 10000
# phot_g_mean_mag + 5 * log10(parallax/1000) + 5 AS g_abs, bp_rp, 1/parallax AS dist
# FROM gaiadr2.gaia_source
# WHERE parallax > 0
# ORDER BY parallax DESC
# ```
#
#
# ### Download your Query Results as a csv file.
#
# ### Hints:
# - Distance is the inverse of parallax.
# - You can calculate absolute photometric magnitude in the gband using this formula: phot_g_mean_mag + 5 + 5 * log10(parallax/1000)
# - You'll need to use "ORDER BY" in your ADQL command.
# - Some Gaia sources have negative parallaxes due to instrumental imperfections. You'll need to add a line to your query specifying that parallax must be greater than 0.
# - Using the buttons that pop up to the right of your query results, you can download the results or view them in your browser. Hover over the buttons to see what they do.
# # Plot a Color-Magnitude Diagram of the 10,000 Closest Stars
# ### 1. Using [pandas.read_csv](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.read_csv.html), read your downloaded csv file into a pandas DataFrame
# +
import pandas as pd
import matplotlib.pyplot as plt
# %pylab inline
# Type your pd.read_csv command here:
data = pd.read_csv('gaia-query-results.csv')
# HINTS:
# - make sure the jupyter notebook and your csv file are in the same directory
# - your read_csv command should be 1 line of code
# -
# ### 2. Using matplotlib.pyplot, make a scatterplot of BP-RP color vs absolute g-band magnitude. This is a [color-magnitude diagram](https://en.wikipedia.org/wiki/Hertzsprung%E2%80%93Russell_diagram)!
# +
plt.figure()
# Type your plotting code here:
plt.scatter(data.bp_rp, data.g_abs, s=.1, color='red')
# More Fun Things to Try if You're Interested:
# - use plt.ylim to reverse the direction of the y axis.
plt.ylim(30,-7)
# - give your plot x and y labels.
plt.xlabel('G$_{BP}$ - G$_{RP}$')
plt.ylabel('M$_G$')
# - make the points red
# (changed in initial plotting command)
# - make the 10 closest stars red
data = data.sort_values(by = 'dist')
data_no_nans = data.dropna() # remove NaN values
plt.scatter(
data_no_nans.bp_rp.iloc[0:10],
data_no_nans.g_abs.iloc[0:10],
color='blue', # blue instead of red for clarity in answer key
s=10. # make these points bigger for clarity in answer key
)
# - compare your results against Figure 1 in this paper: https://arxiv.org/pdf/1804.09378.pdf.
# What similarities and differences do you notice?
"""
In the paper, a heat map is presented, so relative density is easier to see. They also have more datapoints
than we do. I see some of the same features, though, notably thick diagonal line going from top left
to bottom right.
"""
# Challenge: read section 2.1 of this paper and try to reproduce their plot exactly.
# To make the plot below, I used the ADQL query shown in the below jupyter notebook cell.
# Note that I only selected the top 100,000 results for simplicity, not all 13,000,000 as the paper does.
challenge_data = pd.read_csv('gaia-query-challenge-results.csv')
# make a density heatmap
plt.figure()
heatmap, xedges, yedges = np.histogram2d(
challenge_data.bp_rp.values,
challenge_data.g_abs.values,
bins=100
)
extent = [xedges[0], xedges[-1], yedges[0], yedges[-1]]
heatmap = np.ma.masked_where(heatmap == 0.0, heatmap)
color_map = plt.cm.hot
color_map.set_bad(color='white')
plt.imshow(
np.sqrt(heatmap.T),
extent=extent,
cmap=color_map,
aspect=(extent[1]-extent[0])/(extent[3]-extent[2]),
origin='lower'
)
plt.colorbar()
plt.xlabel('G$_{BP}$ - G$_{RP}$')
plt.ylabel('M$_G$')
plt.gca().invert_yaxis()
# To make this look exactly like the figure from the paper, we'd just need to plot all stars in the Gaia catalogue
# -
# <span style="color:red">ADQL Query Command for Data from Babusiaux et al. (2018) (Appendix B):</span>
#
# ```
# SELECT TOP 100000
# phot_g_mean_mag+5*log10(parallax)-10 AS g_abs, bp_rp FROM gaiadr2.gaia_source
# WHERE parallax_over_error > 10
# AND phot_g_mean_flux_over_error > 50
# AND phot_rp_mean_flux_over_error > 20
# AND phot_bp_mean_flux_over_error > 20
# AND phot_bp_rp_excess_factor < 1.3+0.06*power(phot_bp_mean_mag-phot_rp_mean_mag,2)
# AND phot_bp_rp_excess_factor > 1.0+0.015*power(phot_bp_mean_mag-phot_rp_mean_mag,2)
# AND visibility_periods_used > 8
# AND astrometric_chi2_al/(astrometric_n_good_obs_al-5) < 1.44*greatest(1,exp(-0.4*(phot_g_mean_mag-19.5)))
# ```
| Week3_gaia_nasa_exoplanet_archive/GaiaTutorial_KEY.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.0.3
# language: julia
# name: julia-1.0
# ---
# # Examine IHT reconstruction results
# This notebook examines IHT's reconstruction result with and without debiasing. Overall, debiasing does not affect model selection nor parameter estimation.
using DelimitedFiles
using Random
using DataFrames
using StatsBase
using Statistics
using Plots
using Plotly
# # Below are 100 simulations of y where X is 5k by 100k matrix
# +
#debiasing simulation results
normal_5k_by_100k_100 = readdlm("repeats/Normal_100")
logistic_5k_by_100k_100 = readdlm("repeats/Bernoulli_100")
poisson_5k_by_100k_100 = readdlm("repeats/Poisson_100")
negativebinomial_5k_by_100k_100 = readdlm("repeats/NegativeBinomial_100")
#non-debiasing simulation results
# normal_5k_by_100k_100_nodebias = readdlm("repeats_nodebias/Normal_100")
# logistic_5k_by_100k_100_nodebias = readdlm("repeats_nodebias/Bernoulli_100")
# poisson_5k_by_100k_100_nodebias = [readdlm("repeats_nodebias/Poisson_50_1") readdlm("repeats_nodebias/Poisson_50_2")]
# negativebinomial_5k_by_100k_100_nodebias = readdlm("repeats_nodebias/NegativeBinomial_100")
#true model
true_b = [0.25; 0.5; 0.1; 0.8]
# -
# normal_5k_by_100k_100
# logistic_5k_by_100k_100
# poisson_5k_by_100k_100
negativebinomial_5k_by_100k_100
# # Plots reconstruction results (debiasing)
# +
normal = copy(normal_5k_by_100k_100)
logistic = copy(logistic_5k_by_100k_100)
poisson = copy(poisson_5k_by_100k_100)
negatibebinomial = copy(negativebinomial_5k_by_100k_100)
beta_10 = zeros(4, 100)
beta_25 = zeros(4, 100)
beta_50 = zeros(4, 100)
beta_80 = zeros(4, 100)
beta_10[1, :] .= negatibebinomial[3, :]
beta_10[2, :] .= poisson[3, :]
beta_10[3, :] .= logistic[3, :]
beta_10[4, :] .= normal[3, :]
beta_25[1, :] .= negatibebinomial[1, :]
beta_25[2, :] .= poisson[1, :]
beta_25[3, :] .= logistic[1, :]
beta_25[4, :] .= normal[1, :]
beta_50[1, :] .= negatibebinomial[2, :]
beta_50[2, :] .= poisson[2, :]
beta_50[3, :] .= logistic[2, :]
beta_50[4, :] .= normal[2, :]
beta_80[1, :] .= negatibebinomial[4, :]
beta_80[2, :] .= poisson[4, :]
beta_80[3, :] .= logistic[4, :]
beta_80[4, :] .= normal[4, :];
# +
plt = Plots.scatter(beta_10[:, 1], collect(1:1:4), color=:red, xlabel = "Estimated beta Values",
markersize=6, legendfontsize=8, guidefontsize=15, grid=false, framestyle=:box,
xlim=(-0.15, 1.35), label=L"\mathbf{\beta}_{true} = 0.1", legend=:right)
scatter!(beta_25[:, 1], collect(1:1:4), markershape=:hex, color=:lightblue, markersize=6, label=L"\mathbf{\beta}_{true} = 0.25")
scatter!(beta_50[:, 1], collect(1:1:4), markershape=:star5, color=:lightgreen, markersize=6, label=L"\mathbf{\beta}_{true} = 0.5")
scatter!(beta_80[:, 1], collect(1:1:4), markershape=:diamond, color=:lightyellow, markersize=6, label=L"\mathbf{\beta}_{true} = 0.8")
yticks!([1:1:4;], ["NegBin", "Poisson", "Logistic", "Normal"], tickfontsize=15)
xticks!([0.1, 0.25, 0.5, 0.8], ["0.1", "0.25", "0.5", "0.8"], tickfontsize=11)
Random.seed!(111)
for i in 2:100
scatter!(beta_10[:, i], 0.05*randn() .+ collect(1:1:4), markershape=:circle, color=:red, markersize=6, label = "")
scatter!(beta_25[:, i], 0.05*randn() .+ collect(1:1:4), markershape=:hex, color=:lightblue, markersize=6, label = "")
scatter!(beta_50[:, i], 0.05*randn() .+ collect(1:1:4), markershape=:star5, color=:lightgreen, markersize=6, label = "")
scatter!(beta_80[:, i], 0.05*randn() .+ collect(1:1:4), markershape=:diamond, color=:lightyellow, markersize=6, label = "")
end
vline!([0.1,0.25,0.5,0.8], linestyle=:dash, linecolor=:black, linewidth=0.5, label = "")
annotate!([(-0.05, 4.0, text("0/100\n not found", 8, :topleft)),
(-0.05, 3.35, text("84/100\n not found", 8, :topleft)),
(-0.05, 2.23, text("3/100\n not found", 8, :topleft)),
(-0.06, 1.25, text("2/100\n not found", 8, :topleft))])
vspan!(plt,[-Inf,0.04], color = :red, alpha = 0.1, labels = "Not found zone");
plt
# -
Plots.savefig(plt, "repeats.png")
# # Construct Table
#
# ### First compute the probability that reach predictor is found
# +
k = size(true_b, 1)
normal_found = zeros(k)
logistic_found = zeros(k)
poisson_found = zeros(k)
negativebinomial_found = zeros(k)
# normal_found_nodebias = zeros(k)
# logistic_found_nodebias = zeros(k)
# poisson_found_nodebias = zeros(k)
# negativebinomial_found_nodebias = zeros(k)
for i in 1:k
normal_found[i] = sum(normal_5k_by_100k_100[i, :] .!= 0)
logistic_found[i] = sum(logistic_5k_by_100k_100[i, :] .!= 0)
poisson_found[i] = sum(poisson_5k_by_100k_100[i, :] .!= 0)
negativebinomial_found[i] = sum(negativebinomial_5k_by_100k_100[i, :] .!= 0)
# normal_found_nodebias[i] = sum(normal_5k_by_100k_100_nodebias[i, :] .!= 0)
# logistic_found_nodebias[i] = sum(logistic_5k_by_100k_100_nodebias[i, :] .!= 0)
# poisson_found_nodebias[i] = sum(poisson_5k_by_100k_100_nodebias[i, :] .!= 0)
# negativebinomial_found_nodebias[i] = sum(negativebinomial_5k_by_100k_100_nodebias[i, :] .!= 0)
end
# -
negativebinomial_found
# # Found proportion (debiasing)
find_probability = DataFrame(
true_b = true_b[:],
normal_prob_find = normal_found,
logistic_prob_find = logistic_found,
poisson_prob_find = poisson_found,
negativebinomial_prob_find = negativebinomial_found)
#sort!(find_probability, rev=true) #sort later
# # Found proportion (no debiasing)
find_probability_nodebias = DataFrame(
true_b = true_b[:],
normal_prob_find_nodebias = normal_found_nodebias,
logistic_prob_find_nodebias = logistic_found_nodebias,
poisson_prob_find_nodebias = poisson_found_nodebias,
negativebinomial_prob_find_nodebias = negativebinomial_found_nodebias)
#sort!(find_probability, rev=true) #sort later
# # Mean and standard deviation (debiasing)
# +
k = size(true_b, 1)
normal_mean = zeros(k)
normal_std = zeros(k)
logistic_mean = zeros(k)
logistic_std = zeros(k)
poisson_mean = zeros(k)
poisson_std = zeros(k)
negativebinomial_mean = zeros(k)
negativebinomial_std = zeros(k)
for i in 1:k
#compute mean and std if at least 1 found
if normal_found[i] != 0
normal_cur_row = normal_5k_by_100k_100[i, :] .!= 0
normal_mean[i] = mean(normal_5k_by_100k_100[i, :][normal_cur_row])
normal_std[i] = std(normal_5k_by_100k_100[i, :][normal_cur_row])
end
if logistic_found[i] != 0
logistic_cur_row = logistic_5k_by_100k_100[i, :] .!= 0
logistic_mean[i] = mean(logistic_5k_by_100k_100[i, :][logistic_cur_row])
logistic_std[i] = std(logistic_5k_by_100k_100[i, :][logistic_cur_row])
end
if poisson_found[i] != 0
poisson_cur_row = poisson_5k_by_100k_100[i, :] .!= 0
poisson_mean[i] = mean(poisson_5k_by_100k_100[i, :][poisson_cur_row])
poisson_std[i] = std(poisson_5k_by_100k_100[i, :][poisson_cur_row])
end
if negativebinomial_found[i] != 0
negativebinomial_cur_row = negativebinomial_5k_by_100k_100[i, :] .!= 0
negativebinomial_mean[i] = mean(negativebinomial_5k_by_100k_100[i, :][negativebinomial_cur_row])
negativebinomial_std[i] = std(negativebinomial_5k_by_100k_100[i, :][negativebinomial_cur_row])
end
end
# -
found_mean_and_std = DataFrame(
true_b = true_b,
normal_mean = normal_mean,
normal_std = normal_std,
logistic_mean = logistic_mean,
logistic_std = logistic_std,
poisson_mean = poisson_mean,
poisson_std = poisson_std,
negativebinomial_mean = negativebinomial_mean,
negativebinomial_std = negativebinomial_std)
# sort!(found_mean_and_std, rev=true) #sort later
# # Mean and standard deviation (non-debiasing)
# +
k = size(true_b, 1)
normal_mean_nodebias = zeros(k)
normal_std_nodebias = zeros(k)
logistic_mean_nodebias = zeros(k)
logistic_std_nodebias = zeros(k)
poisson_mean_nodebias = zeros(k)
poisson_std_nodebias = zeros(k)
negativebinomial_mean_nodebias = zeros(k)
negativebinomial_std_nodebias = zeros(k)
for i in 1:k
#compute mean and std if at least 1 found
if normal_found_nodebias[i] != 0
normal_cur_row = normal_5k_by_100k_100_nodebias[i, :] .!= 0
normal_mean_nodebias[i] = mean(normal_5k_by_100k_100_nodebias[i, :][normal_cur_row])
normal_std_nodebias[i] = std(normal_5k_by_100k_100_nodebias[i, :][normal_cur_row])
end
if logistic_found_nodebias[i] != 0
logistic_cur_row = logistic_5k_by_100k_100_nodebias[i, :] .!= 0
logistic_mean_nodebias[i] = mean(logistic_5k_by_100k_100_nodebias[i, :][logistic_cur_row])
logistic_std_nodebias[i] = std(logistic_5k_by_100k_100_nodebias[i, :][logistic_cur_row])
end
if poisson_found_nodebias[i] != 0
poisson_cur_row = poisson_5k_by_100k_100_nodebias[i, :] .!= 0
poisson_mean_nodebias[i] = mean(poisson_5k_by_100k_100_nodebias[i, :][poisson_cur_row])
poisson_std_nodebias[i] = std(poisson_5k_by_100k_100_nodebias[i, :][poisson_cur_row])
end
if negativebinomial_found_nodebias[i] != 0
negativebinomial_cur_row = negativebinomial_5k_by_100k_100_nodebias[i, :] .!= 0
negativebinomial_mean_nodebias[i] = mean(negativebinomial_5k_by_100k_100_nodebias[i, :][negativebinomial_cur_row])
negativebinomial_std_nodebias[i] = std(negativebinomial_5k_by_100k_100_nodebias[i, :][negativebinomial_cur_row])
end
end
# -
found_mean_and_std_nodebias = DataFrame(
true_b = true_b[:],
normal_mean_nodebias = normal_mean_nodebias,
normal_std_nodebias = normal_std_nodebias,
logistic_mean_nodebias = logistic_mean_nodebias,
logistic_std_nodebias = logistic_std_nodebias,
poisson_mean_nodebias = poisson_mean_nodebias,
poisson_std_nodebias = poisson_std_nodebias,
negativebinomial_mean_nodebias = negativebinomial_mean_nodebias,
negativebinomial_std_nodebias = negativebinomial_std_nodebias)
# sort!(found_mean_and_std_nodebias, rev=true) #sort later
# # Sort and round results (debiasing)
sort!(found_mean_and_std, rev=true)
for i in 1:size(found_mean_and_std, 2)
found_mean_and_std[:, i] = round.(found_mean_and_std[:, i], digits=3)
end
found_mean_and_std
# # Sort and round results (non-debiasing)
sort!(found_mean_and_std_nodebias, rev=true)
for i in 1:size(found_mean_and_std_nodebias, 2)
found_mean_and_std_nodebias[:, i] = round.(found_mean_and_std_nodebias[:, i], digits=3)
end
found_mean_and_std_nodebias
| figures/repeats/old4/IHT_reconstruction_results.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="FMd9ebo7QYDh"
# 이번엔 와인을 분류해 보자
# + id="9Yla7ABmQcHS"
#필요한 모듈 import 하기
from sklearn.datasets import load_wine
from sklearn.model_selection import train_test_split
from sklearn.metrics import classification_report
wine = load_wine()
# + colab={"base_uri": "https://localhost:8080/"} id="73VpqWWeiCOs" outputId="5a6ec325-3d3f-46d2-9b22-834c983e38f6"
print(type(dir(wine)))
# + colab={"base_uri": "https://localhost:8080/"} id="2qPlrCPZen_Q" outputId="7dce1272-ff03-49e8-c5ed-6cc4c58ad178"
wine.keys()
# + colab={"base_uri": "https://localhost:8080/"} id="a-4WstV-e0T2" outputId="26f1d9f1-90c8-4103-881a-654ebbb7822a"
print(wine.DESCR)
# + id="9-KWFiy5fFD9"
wine_data = wine.data
# + colab={"base_uri": "https://localhost:8080/"} id="QTaGIHqegFc_" outputId="5701c25a-4e37-4225-de0d-d34f5b621a02"
print(wine_data.shape)
# + colab={"base_uri": "https://localhost:8080/"} id="TcXbQxire2ZO" outputId="3e4fdf3e-3177-458e-8b98-3e251a823268"
wine.feature_names
# + colab={"base_uri": "https://localhost:8080/"} id="lisz1dqTgKCD" outputId="7b813eee-ad17-43d5-85cd-625a867f236e"
wine_data[0]
# + colab={"base_uri": "https://localhost:8080/"} id="md8L0sb1eunI" outputId="5bf58218-4a83-4309-c5a2-bebc96095e08"
wine.target_names
# + id="d5_r2rHGfN0N"
wine_label = wine.target
# + colab={"base_uri": "https://localhost:8080/", "height": 215} id="1wSUU460jRzU" outputId="6128029a-e238-46dc-90b7-e9077fb4ba73"
import pandas as pd
wine_df = pd.DataFrame(data=wine_data, columns=wine.feature_names)
wine_df.head()
# + colab={"base_uri": "https://localhost:8080/"} id="jT7h1Eh_jh3r" outputId="4645f35d-c533-43a9-da55-77e3e6279268"
wine_df.info()
# + colab={"base_uri": "https://localhost:8080/", "height": 304} id="-nxcTqG8joaC" outputId="acee3128-e801-483f-c28a-dd3e92b8e1f6"
wine_df.describe()
# + id="YltoU0DykAqq"
# + colab={"base_uri": "https://localhost:8080/"} id="SwK-CLNZfRba" outputId="0b9ca877-9b5e-4d14-cae4-53e6e384001d"
X_train, X_test, y_train, y_test = train_test_split(wine_data,
wine_label,
test_size=0.2,
random_state = 3
)
from sklearn.tree import DecisionTreeClassifier
from sklearn.metrics import accuracy_score
#DecisionTree를 사용
decision_tree = DecisionTreeClassifier(random_state=3)
decision_tree.fit(X_train, y_train)
y_pred = decision_tree.predict(X_test)
from sklearn.metrics import classification_report
print(classification_report(y_test, y_pred))
accuracy = accuracy_score(y_test,y_pred)
accuracy
# + colab={"base_uri": "https://localhost:8080/"} id="q2jl05kyfXcR" outputId="f6edc06b-ce0e-4b63-b3d8-ac0089fdb41c"
from sklearn.ensemble import RandomForestClassifier
X_train, X_test, y_train, y_test = train_test_split(wine_data,
wine_label,
test_size=0.2,
random_state=3)
random_forest = RandomForestClassifier(random_state=3)
random_forest.fit(X_train, y_train)
y_pred = random_forest.predict(X_test)
print(classification_report(y_test, y_pred))
accuracy = accuracy_score(y_test,y_pred)
accuracy
# + colab={"base_uri": "https://localhost:8080/"} id="rP0e3mshuzyU" outputId="b22d3204-46fc-43e3-b880-b2735280a007"
from sklearn.metrics import confusion_matrix
confusion_matrix(y_test, y_pred)
# + colab={"base_uri": "https://localhost:8080/"} id="b1BSjmWmvMX5" outputId="1ec88df9-1f67-4d1f-b524-9061d67f38df"
#SVM을 사옹해 보자
from sklearn import svm
svm_model = svm.SVC()
print(svm_model._estimator_type)
# + colab={"base_uri": "https://localhost:8080/"} id="eKOSF8TkvwZM" outputId="ba053740-ede4-477a-e052-2120fdaaee84"
svm_model.fit(X_train, y_train)
y_pred = svm_model.predict(X_test)
print(classification_report(y_test, y_pred))
accuracy = accuracy_score(y_test, y_pred)
accuracy
# + colab={"base_uri": "https://localhost:8080/"} id="fCzuljw_wElt" outputId="5bbe22c4-ad29-4ea8-9533-88f832498684"
confusion_matrix(y_test, y_pred)
# + colab={"base_uri": "https://localhost:8080/"} id="D9MOw_EywLh6" outputId="a837124b-f8f7-4afb-ea87-4a4435b84b70"
#SGDClassifier
from sklearn.linear_model import SGDClassifier
sgd_model = SGDClassifier()
sgd_model.fit(X_train, y_train)
y_pred = sgd_model.predict(X_test)
print(classification_report(y_test, y_pred))
accuracy = accuracy_score(y_test, y_pred)
accuracy
#적합한 모델이 아닌듯하다.
# + colab={"base_uri": "https://localhost:8080/"} id="hsgMZeuYww8j" outputId="c2568b98-3c14-4d37-983c-b9eb2956fefa"
#Logistic Regression
from sklearn.linear_model import LogisticRegression
logistic_model = LogisticRegression()
print(logistic_model._estimator_type)
# + colab={"base_uri": "https://localhost:8080/"} id="eCW_6f-UxN4H" outputId="9b64cf58-06d1-4e30-bb55-f8a49563d81b"
logistic_model.fit(X_train, y_train)
y_pred = logistic_model.predict(X_test)
print(classification_report(y_test, y_pred))
accuracy = accuracy_score(y_test, y_pred)
accuracy
# + colab={"base_uri": "https://localhost:8080/"} id="l16u6ZmExyUF" outputId="f82e0b06-85aa-431a-d92e-59de8bbf56aa"
confusion_matrix(y_test, y_pred)
# + id="LKBZI0nux2en"
# + [markdown] id="YAf9XkQnyI1n"
# 와인의 경우 랜덤포레스트가 정확도가 가장 높게 나왔다.
# + id="lbIwQf60yLq3"
| Exloratory/exp2/exp2_project_wine.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.4 64-bit (''.venv'': venv)'
# language: python
# name: python394jvsc74a57bd0e7a5033076b597f81e5ae7ba1abbcc3e4e4f13c94c7fa961e9c6d3be8b84fac4
# ---
# # Usage: scenario analysis
#
# [](https://colab.research.google.com/github/lisphilar/covid19-sir/blob/master/example/usage_quick.ipynb)
#
# This is a quick tour of CovsirPhy. Details scenario analysis will be explained.
# "Scenario analysis" means that we calculate the number of cases in the future phases with some sets of ODE parameter values. With this analysis, we can estimate the impact of our activities against the outbreak on the number of cases.
# ### Preparation
# Prepare the packages.
# + tags=[]
# # !pip install covsirphy --upgrade
from pprint import pprint
import covsirphy as cs
cs.__version__
# -
# ### Dataset preparation
# Download the datasets to "../input" directory and load them.
# Please refer to [Usage: datasets](https://lisphilar.github.io/covid19-sir/usage_dataset.html) for the details.
# + tags=[]
data_loader = cs.DataLoader("../input")
# The number of cases (JHU style)
jhu_data = data_loader.jhu()
# Population in each country
population_data = data_loader.population()
# Government Response Tracker (OxCGRT)
oxcgrt_data = data_loader.oxcgrt()
# The number of tests
pcr_data = data_loader.pcr()
# The number of vaccinations
vaccine_data = data_loader.vaccine()
# -
# ### Start scenario analysis
# As an example, we will analysis the number of cases in Japan. `covsirphy.Scenario` is the interface for analysis. Please specify the area (country: required, province: optional) when creating the instance and register the datasets with `Scenario.register()`. As the extra datasets, we can select `OxCGRTData`, `PCRData`, `VaccineData` and `CountryData`.
# Specify country and province (optinal) names
snl = cs.Scenario(country="Japan", province=None)
# Register datasets
snl.register(jhu_data, population_data, extras=[oxcgrt_data, pcr_data, vaccine_data])
# We call `JHUData` and `PopulationData` as "the main datasets" because they are required to calculate the number of susceptible/infected/recovered/fatal cases. These variables are used in SIR-F model.
# The other datasets are called as "the extra datasets" and they will be used to predict the futute parameter values of SIR-F model for forecasting the number of cases with some scenarios.
#
# Additional information:
#
# - Details of the datasets: [Usage: datasets](https://lisphilar.github.io/covid19-sir/usage_dataset.html)
# - Details of SIR-F model: [Usage: SIR-derived models](https://lisphilar.github.io/covid19-sir/usage_theoretical.html)
# #### Display/save figures
# We have intactive mode and script mode to display/save figures.
# When use use interactive shells, including Jupyter Notebook, we can choose either "interactive shell mode" or "script mode" as follows.
# Interactive mode:
# Figures will be displayed as the output of code cells.
# Choose interactive mode (default is True when we use interactive shells)
snl.interactive = True
# When you want to turn-off interactive mode temporarlly, set `False` as `Scenario.interactive` or apply `show_figure=False` as an argument of methods, including `Scenario.records()`. Methods with figures will be shown later in this tutorial.
# +
# apply "show_figures=False" to turn-off intaractive mode temporally
# snl.records(show_figure=False)
# -
# Script mode:
# In script mode, figures will not be displayed. When filenames were applied to the methods as `filename` argument, figures will be saved in your local environment.
# +
# Stop displaying figures
# snl.interactive = False
# With this mode we can save figures, specifying "filename" argument
# snl.records(filename="records.jpg")
# -
# When we run codes as a script (eg. `python scenario_analysis.py`), only "script mode" is selectable and `Scenario.interactive` is always `False`. Figures will be saved when filenames are specified with `filename` argument.
# Because some methods, including `Scenario.summary()`, return dataframes (`pandas.DataFrame`), we can save them as CSV files etc. using `.to_csv(filename, index=True)`.
# We can produce filenames more easily with `Filer` class. Please refer to the scripts in [example directory of the repository](https://github.com/lisphilar/covid19-sir/tree/master/example).
filer = cs.Filer(directory="output", prefix="jpn", suffix=None, numbering="01")
# filer.png("records")
# -> {"filename": "<absolute path>/output/jpn_01_records.png"}
# filer.jpg("records")
# -> {"filename": "<absolute path>/output/jpn_01_records.jpg"}
# filer.csv("records", index=True)
# -> {"path_or_buf": "<absolute path>/output/jpn_01_records.csv", index: True}
# We can save files more easily with `Filer` as follows.
# +
# record_df = snl.records(**filer.png("records"))
# record_df.to_csv(**filer.csv("records", index=False))
# -
# ### Check records
# Let's see the records at first. `Scenario.records()` method return the records as a pandas dataframe and show a line plot. Some kind of complement will be done for analysis, if necessary.
# `Scenario.records()` shows the number of infected/recovered/fatal cases as default. Using `variables` argument, we can set the variables to show. Here, we chack the number of confirmed/fatal/recovered cases. They are cumurative values.
snl.records(variables="CFR").tail()
# This is the same as
# snl.records(variables=["Confirmed", "Fatal", "Recovered"])
# The number of infected cases on date:
_ = snl.records(variables="I")
# This is the same as
# snl.records(variables=["Infected"])
# All available variables can be retrieved with `variables="all"`.
df = snl.records(variables="all", show_figure=False)
pprint(df.set_index("Date").columns.tolist(), compact=True)
# We can specify the variables to show.
snl.records(variables=["Vaccinated_once"]).tail()
# We can calculate the number of daily new cases with `Scenario.record_diff()` method.
# Acceptable variables are the same as Scenario.records()
_ = snl.records_diff(variables="C", window=7)
# `Scenario.show_complement()` method is useful to show the kinds of complement. The details of complement are explained in [Usage: datasets](https://lisphilar.github.io/covid19-sir/usage_dataset.html#The-number-of-cases-(JHU-style)) section.
# Show the details of complement
snl.show_complement()
# ### S-R trend analysis
# S-R trend analysis finds the change points of SIR-derived ODE parameters. Details will be explained in [Usage (details: phases)](https://lisphilar.github.io/covid19-sir/usage_phases.html). Phases will be separated with dotted lines. i.e. Dot lines indicate the start dates of phases.
snl.trend().summary()
# ### Parameter estimation of ODE models
# Here, we will estimate the tau value [min] (using grid search) and parameter values of SIR-derived models using [Optuna](https://github.com/optuna/optuna) package (automated hyperparameter optimization framework). As an example, we use SIR-F model. Details of models will be explained in [Usage (details: theoritical datasets)](https://lisphilar.github.io/covid19-sir/usage_theoretical.html).
#
# **We can select the model from SIR, SIRD and SIR-F model for parameter estimation. SIR-FV model (completely deprecated) and SEWIR-F model cannot be used.**
# + tags=[]
# Estimate the tau value and parameter values of SIR-F model
# Default value of timeout in each phase is 180 sec
snl.estimate(cs.SIRF, timeout=180)
# -
# Show the summary of parameter estimation
snl.summary()
# ### Evaluation of estimation accuracy
# Accuracy of parameter estimation can be evaluated with RMSLE (Root Mean Squared Log Error) score.
#
# \begin{align*}
# \mathrm{RMSLE} = \sqrt{\cfrac{1}{n}\sum_{i=1}^{n}(log_{10}(A_{i} + 1) - log_{10}(P_{i} + 1))^2}
# \end{align*}
#
# Where $A$ is the observed (actual) values, $P$ is estimated (predicted) values. Variables are $S (i=1), I (i=2), R (i=3)\ \mathrm{and}\ F (i=n=4)$ for SIR-F model. When RMSLE score is low, hyperparameter estimation is highly accurate.
# Please refer to external sites, including [Medium: What’s the Difference Between RMSE and RMSLE?](https://medium.com/analytics-vidhya/root-mean-square-log-error-rmse-vs-rmlse-935c6cc1802a)
# Show RMSLE scores with the number of optimization trials and runtime for phases
snl.summary(columns=["Start", "End", "RMSLE", "Trials", "Runtime"])
# Additionally, we can visualize the accuracy with `Scenario.estimate_accuracy()`, specifing phase name.
# Visualize the accuracy for the 2nd phase
snl.estimate_accuracy(phase="2nd")
# phase="last" means the last phases
# snl.estimate_accuracy(phase="last")
# We can calculate total score for all phases using `Scenario.score()` method. Metrics can be selected from MAE, MSE, MSLE, RMSE and RMSLE.
# Get total score: metrics="MAE", "MSE", "MSLE", "RMSE" or "RMSLE"
# snl.score(metrics="RMSLE")
metrics_list = ["MAE", "MSE", "MSLE", "RMSE", "RMSLE"]
for metrics in metrics_list:
metrics_name = metrics.rjust(len(max(metrics_list, key=len)))
print(f"{metrics_name}: {snl.score(metrics=metrics):.3f}")
# ### Get parameter value
# We can get the parameter values of a phase using `Scenario.get()` method.
# Get parameter values
snl.get("Rt", phase="4th")
# phase="last" means the last phases
snl.get("Rt", phase="last")
# ### Show parameter history
# We can get the history of parameter values with a dataframe and a figure.
# Get the parameter values as a dataframe
snl.summary(columns=[*cs.SIRF.PARAMETERS, "Rt"])
# `Scenario.history()` method shows the trajectories of parameters (and the number of cases).
_ = snl.history(target="theta", show_legend=False)
_ = snl.history(target="kappa", show_legend=False)
_ = snl.history(target="rho", show_legend=False)
_ = snl.history(target="sigma", show_legend=False)
# Notes on the history of $\sigma$ value in japan (last updated: 28Dec2020):
# In Japan, we experienced two waves and we are in third wave. In the first wave (Apr - May), recovery period was too long because collapse of the medical care system occurred and no medicines were found.
#
# Sigma values: the first wave < the second wave > the third wave
#
# However, in the second wave (Jul - Oct), recovery period appears short because we have some effective medicines (not approved, in clinical study), yonger people (people un-associated to sever diseases) were infected.
#
# In the third wave (Nov - ), older people tend to be infected and we are facing with medical collapse at this time...
# ### Show the history of reproduction number
# $R_0$ ("R naught") means "the average number of secondary infections caused by an infected host" ([Infection Modeling — Part 1](https://towardsdatascience.com/infection-modeling-part-1-87e74645568a)). When this value is larger than 1, the infection disease is outbreaking.
_ = snl.history(target="Rt", show_legend=False)
# ### Simulate the number of cases
# We can compare the actual and simulated (with estimated parameter values) number of confirmed/infected/recovered/fatal cases using `Scenario.history()` method.
# Compare the actual values and the main scenario
_ = snl.history("Infected")
# When we want to show only one scenario with all variables, we use `Scenario.simulate()` method.
_ =snl.simulate(name="Main")
# We can select variables and phases with arguments.
_ = snl.simulate(name="Main", variables=["Confirmed", "Infected"], phases=["9th", "10th"])
# ### Main scenario
# To investigate the effect of parameter changes, we will perform scenario analysis. In the main scenario, we will assume that the parameter values do not change after the last past phase.
#
# i.e. If the parameter velues will not be changed until 31Jul2021, how many cases will be? We call this scenario as "Main" scenario.
# Clear future phases in Main scenario
snl.clear(name="Main")
# Add one future phase 30 days with the parameter set of the last past phase
snl.add(days=30, name="Main")
# Add one future phase until 31Jul2021 with the same parameter set
snl.add(end_date="31Jul2021", name="Main")
# Simulate the number of cases
snl.simulate(name="Main").tail()
# ### Medicine scenario
# To investigate the effect of new medicines, we will assume that $\sigma$ will be changed in the future phases.
#
# If $\sigma$ will be 1.2 times in 30 days, how many cases will be? We will call this scenario as "Medicine" scenario.
# Calcuate the current sigma value of the last phase
sigma_current = snl.get("sigma", name="Main", phase="last")
sigma_current
# Sigma value will be double
sigma_new = sigma_current * 1.2
sigma_new
# Initialize "Medicine" scenario (with the same past phases as that of Main scenario)
snl.clear(name="Medicine")
# Add 30 days as a new future phases with the same parameter set
snl.add(name="Medicine", days=30, sigma=sigma_current)
# Add a phase with doubled sigma value and the same end date with main date
snl.add(name="Medicine", end_date="31Jul2021", sigma=sigma_new)
snl.summary(name="Medicine")
# Simulate the number of cases.
_ = snl.simulate(name="Medicine").tail()
# ### Short-term prediction of parameter values
# With extra datasets, we can predict the parameter values of the future phases because [OxCGRT indicators](https://github.com/OxCGRT/covid-policy-tracker) (policy measures), vaccinations and so on impact on parameter values with the delay period. Delay period will be calculated with `cenario.estimate_delay()` automatically.
#
# OxCGRT indicators are
#
# - school_closing,
# - workplace_closing,
# - cancel_events,
# - gatherings_restrictions,
# - transport_closing,
# - stay_home_restrictions,
# - internal_movement_restrictions,
# - international_movement_restrictions,
# - information_campaigns,
# - testing_policy, and
# - contact_tracing.
# `Scenario.fit()` method learns the relationship of indicators (X) and the parameter values (y) with regresion model (Elastic Net regression). X was registered with `Scenario.register()` and y was calculated with `Scenario.estimate()` in advance respectively.
# The indicator used to estimate delay period ("Stringency_index" as default) will be automatically from X dataset. If you have some indicators to be removed additionally, please use `removed_cols` argument (list of indicator names) of `Scenario.fit()`.
# Here, we will remove "Tests" and "Tests_diff" as an example. (This is because we have "Testing_policy" indicator and they may be overlaped.)
# Create Forecast scenario (copy Main scenario and delete future phases)
snl.clear(name="Forecast", template="Main")
# Fitting with linear regression model (Elastic Net regression)
fit_dict = snl.fit(name="Forecast", removed_cols=["Tests", "Tests_diff"])
# Show determination coeefients of training/test dataset.
print(f"Determination coefficient: {fit_dict['score_train']:.3f} (train)")
print(f"Determination coefficient: {fit_dict['score_test']:.3f} (test)")
# Show the intercept and coefficient values of the regression model.
print("Intercept and coefficient:")
fit_dict["intercept"].style.background_gradient(axis=None)
# `Scenario.predict()` predicts the parameter values of future phases.
# Short-term prediction
snl.predict(name="Forecast").summary(name="Forecast")
# We can select list of days to predict optionally
# snl.predict(days=[1, 4], name="Forecast").summary(name="Forecast")
# Or, short-cut `.fit()` and `.predict()` with `Scenario.fit_predict()`.
# +
# Or, when you do not need 'fit_dict',
# snl.fit_predict(name="Forecast").summary(name="Forecast")
# -
# To compare this scenario with the other scenarios, we should adjust the last end date with `Scenario.adjust_end()` because the last end date is differenct from the other scenarios at this time.
# Adjust the last end dates
snl.adjust_end()
# Show the last phases of all scenarios
all_df = snl.summary().reset_index()
for name in all_df["Scenario"].unique():
df = snl.summary(name=name)
last_end_date = df.loc[df.index[-1], "End"]
print(f"{name} scenario: to {last_end_date}")
# Simulate the number of cases of forecast scenario.
_ = snl.simulate(variables="CFR", name="Forecast").tail()
# Compare the number of cases of the all scenario with `Scenario.history()` and variable name.
_ = snl.history("Infected")
# From version 2.19.1-eta, we can focus on the values in specified date range with the following arguments.
#
# - `dates`: tuple of start date and end date
# - `past_days` (integer): how many past days to use in calculation from today (`Scenario.today` property)
# - `phases` (list of str): phase names to use in calculation
#
# These arguments are effective with `Scenario.history()`, `Scenario.simulate()`, `Scenario.track()` and `Scenario.score()`.
# Get the minimum value (from today to future) to set lower limit of y-axis
lower_limit = snl.history("Infected", dates=(snl.today, None), show_figure=False).min().min()
# From today to future (no limitation regarding end date)
_ = snl.history("Infected", dates=(snl.today, None), ylim=(lower_limit, None))
# In the past 20 days. Reference date is today (`Scenario.today` property).
_ = snl.history("Infected", past_days=20)
# In the selected phases. Here, we will show the 3rd, 4th and 5th phase.
_ = snl.history("Infected", phases=["3rd", "4th", "5th"])
# ### Compare the scenarios
# We will compare the scenarios with representative values, reproduction number and parameter values. Currently, we can compare the scenarios with the following indexes.
#
# - max(Infected): max value of Infected
# - argmax(Infected): the date when Infected shows max value
# - Infected on …: Infected on the end date of the last phase
# - Fatal on …: Fatal on the end date of the last phase
snl.describe()
_ = snl.history(target="Infected")
_ = snl.history(target="Rt")
_ = snl.history(target="rho")
_ = snl.history(target="sigma")
_ = snl.history(target="theta")
_ = snl.history(target="kappa")
# ### Change rate of parameters in main scenario
# History of each parameter will be shown. Values will be divided by the values in 0th phase.
_ = snl.history_rate(name="Main")
# ### Retrospective analysis
# We can evaluate the impact of measures using past records. How many people were infected if the parameter values have not changed sinse 01Sep2020?
# Perform retrospective analysis
snl_retro = cs.Scenario(jhu_data, population_data, "Japan")
snl_retro.retrospective(
"01Jan2021", model=cs.SIRF, control="Main", target="Retrospective", timeout=10)
# Show the summary of estimation
cols = ["Start", "End", "ODE", "Rt", *cs.SIRF.PARAMETERS] + ["RMSLE", "Trials", "Runtime"]
snl_retro.summary(columns=cols)
# History of reproduction number
_ = snl_retro.history("Rt")
# History of Infected
_ = snl_retro.history("Infected")
# Show the representative values
snl_retro.describe()
| example/usage_quick.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import keras
os.environ["CUDA_VISIBLE_DEVICES"] = "" # 使用 CPU
import cv2 # 載入 cv2 套件
import matplotlib.pyplot as plt
train, test = keras.datasets.cifar10.load_data()
# +
image = train[0][0] # 讀取圖片
plt.imshow(image)
plt.show()
# -
# 把彩色的圖片轉為灰度圖
gray = cv2.cvtColor(image, cv2.COLOR_RGB2GRAY)
plt.imshow(gray)
plt.show()
# 通過調用 __cv2.calcHist(images, channels, mask, histSize, ranges)__ 函數來得到統計後的直方圖值
#
# * imaages (list of array):要分析的圖片
# * channels:產生的直方圖類型。例:[0]→灰度圖,[0, 1, 2]→RGB三色。
# * mask:optional,若有提供則僅計算 mask 部份的直方圖。
# * histSize:要切分的像素強度值範圍,預設為256。每個channel皆可指定一個範圍。例如,[32,32,32] 表示RGB三個channels皆切分為32區段。
# * ranges:像素的範圍,預設為[0,256],表示<256。
# 調用 cv2.calcHist 函數,回傳值就是 histogram
hist = cv2.calcHist([gray], [0], None, [256], [0, 256])
plt.figure()
plt.title("Grayscale Histogram")
plt.xlabel("Bins")
plt.ylabel("# of Pixels")
plt.plot(hist)
plt.xlim([0, 256])
plt.show()
print("hist shape:", hist.shape, "\n直方圖中前兩個值:", hist[:2]) # 1 表示該灰度圖中,只有 1 個 pixel 的值是 0,0 個 pixel 的值是 1
# +
chans = cv2.split(image) # 把圖像的 3 個 channel 切分出來
colors = ("r", "g", "b")
plt.figure()
plt.title("'Flattened' Color Histogram")
plt.xlabel("Bins")
plt.ylabel("# of Pixels")
# 對於所有 channel
for (chan, color) in zip(chans, colors):
# 計算該 channel 的直方圖
hist = cv2.calcHist([chan], [0], None, [256], [0, 256])
# 畫出該 channel 的直方圖
plt.plot(hist, color = color)
plt.xlim([0, 256])
plt.show()
# -
# ## 作業
# 畫出 16 個 bin 的顏色直方圖,並嘗試回答每個 channel 在 [16, 32] 這個 bin 中有多少個 pixel?
| homeworks/D090/Day090_color_histogram.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Austrian energy system Tutorial Part 1: Building an Energy Model
#
# For information on how to install *MESSAGEix*, please refer to [Installation page](https://docs.messageix.org/en/stable/#getting-started) and for getting *MESSAGEix* tutorials, please follow the steps mentioned in [Tutorials](https://docs.messageix.org/en/stable/tutorials.html).
#
# Please refer to the [user guidelines](https://github.com/iiasa/message_ix/blob/master/NOTICE.rst)
# for additional information on using *MESSAGEix*, including the recommended citation and how to name new models.
#
# **Pre-requisites**
# - You have the *MESSAGEix* framework installed and working
#
# **Structure of these tutorials.** After having run this baseline tutorial, you are able to start with any of the other tutorials, but we recommend to follow the order below for going through the information step-wise:
#
# 1. Prepare the base model version (Python: ``austria.ipynb``, also available in R: ``austria_reticulate.ipynb``)
# 2. Plot the results of the baseline runs (Python: ``austria_load_scenario.ipynb``, also available in R: ``austria_load_scenario_R.ipynb``).
# 3. Run a single policy scenario (``austria_single_policy.ipynb``).
# 4. Run multiple policy scenarios. This tutorial has two notebooks: an introduction with some exercises and completed code for the exercises (exercises: ``austria_multiple_policies.ipynb``, answers: ``austria_multiple_policies-answers.ipynb``).
#
# **Introduction**
#
# In this notebook, we will build a model of the Austrian energy system from scratch. The process will involve defining our model's time horizon and spatial extent, and then populating the model with data associated with model parameters. Once we have a baseline model, we will then move on to investigating policy scenarios.
#
# We will be populating different kinds of parameters including:
#
# ### Economic Parameters
#
# - `interestrate`
# - `demand`
#
# ### Technology Parameters
#
# #### Engineering Parameters
#
# - `input`
# - `output`
# - `technical_lifetime`
# - `capacity_factor`
#
#
# #### Technoeconomic Parameters
#
# - `inv_cost`
# - `fix_cost`
# - `var_cost`
#
# ### Dynamic Behavior Parameters
#
# - `bound_activity_up`
# - `bound_activity_lo`
# - `bound_new_capacity_up`
# - `initial_activity_up`
# - `growth_activity_up`
#
# ### Emissions
#
# - `emission_factor`
#
# A full list of parameters can be found in the [MESSAGEix documentation](http://messageix.iiasa.ac.at/model/MESSAGE/parameter_def.html). (If you have cloned the MESSAGEix [Github repository](https://github.com/iiasa/message_ix), the documentation can also be built offline; see `doc/README.md`.)
#
# ## The Final Product
#
# At the completion of this exercise, we will have developed an energy model is comprised of the below Reference Energy System (RES):
#
# 
#
#
# ## Setup
# +
# load required packages
import itertools
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
plt.style.use('ggplot')
import ixmp as ix
import message_ix
from message_ix.utils import make_df
# -
# launch the IX modeling platform using the local default database
mp = ix.Platform()
# +
model = "Austrian energy model"
scen = "baseline"
annot = "developing a stylized energy system model for illustration and testing"
scenario = message_ix.Scenario(mp, model, scen, version='new', annotation=annot)
# -
# ## Time and Spatial Detail
#
# The model includes the time periods 2010, 2020, 2030 and 2040.
horizon = range(2010, 2041, 10)
scenario.add_horizon(year=horizon)
country = 'Austria'
scenario.add_spatial_sets({'country': country})
# ## Model Structure
scenario.add_set("commodity", ["electricity", "light", "other_electricity"])
scenario.add_set("level", ["secondary", "final", "useful"])
scenario.add_set("mode", "standard")
# ## Economic Parameters
# Definition of the socio-economic discount rate:
scenario.add_par("interestrate", horizon, value=0.05, unit='-')
# The fundamental premise of the model is to satisfy demand for energy (services). To first order, demands for services (e.g. electricity) track with economic productivity (GDP). Therefore, as a simple example, we define both a GDP profile and a correlation factor between GDP growth and demand, called beta. Beta will then be used to obtain a simplistic demand profile.
gdp = pd.Series([1., 1.21631, 1.4108, 1.63746], index=horizon)
beta = 0.7
demand = gdp ** beta
# ## Technologies
# +
plants = [
"coal_ppl",
"gas_ppl",
"oil_ppl",
"bio_ppl",
"hydro_ppl",
"wind_ppl",
"solar_pv_ppl", # actually primary -> final
]
secondary_energy_techs = plants + ['import']
final_energy_techs = ['electricity_grid']
lights = [
"bulb",
"cfl",
]
useful_energy_techs = lights + ['appliances']
# -
technologies = secondary_energy_techs + final_energy_techs + useful_energy_techs
scenario.add_set("technology", technologies)
# +
demand_per_year = 55209. / 8760 # from IEA statistics
elec_demand = pd.DataFrame({
'node': country,
'commodity': 'other_electricity',
'level': 'useful',
'year': horizon,
'time': 'year',
'value': demand_per_year * demand,
'unit': 'GWa',
})
scenario.add_par("demand", elec_demand)
demand_per_year = 6134. / 8760 # from IEA statistics
light_demand = pd.DataFrame({
'node': country,
'commodity': 'light',
'level': 'useful',
'year': horizon,
'time': 'year',
'value': demand_per_year * demand,
'unit': 'GWa',
})
scenario.add_par("demand", light_demand)
# -
# ### Engineering Parameters
year_df = scenario.vintage_and_active_years()
vintage_years, act_years = year_df['year_vtg'], year_df['year_act']
# +
base_input = {
'node_loc': country,
'year_vtg': vintage_years,
'year_act': act_years,
'mode': 'standard',
'node_origin': country,
'commodity': 'electricity',
'time': 'year',
'time_origin': 'year',
}
grid = pd.DataFrame(dict(
technology = 'electricity_grid',
level = 'secondary',
value = 1.0,
unit = '-',
**base_input
))
scenario.add_par("input", grid)
bulb = pd.DataFrame(dict(
technology = 'bulb',
level = 'final',
value = 1.0,
unit = '-',
**base_input
))
scenario.add_par("input", bulb)
cfl = pd.DataFrame(dict(
technology = 'cfl',
level = 'final',
value = 0.3, # LED and CFL lighting equipment are more efficient than conventional light bulbs,
#so they need less input electricity to produce the same quantity of 'light'
#compared to conventional light bulbs (0.3 units vs 1.0, respectively)
unit = '-',
**base_input
))
scenario.add_par("input", cfl)
app = pd.DataFrame(dict(
technology = 'appliances',
level = 'final',
value = 1.0,
unit = '-',
**base_input
))
scenario.add_par("input", app)
# +
# make_df?
# to see what the funcion "df" does.
# +
base_output = {
'node_loc': country,
'year_vtg': vintage_years,
'year_act': act_years,
'mode': 'standard',
'node_dest': country,
'time': 'year',
'time_dest': 'year',
'unit': '-',
}
imports = make_df(base_output, technology='import', commodity='electricity',
level='secondary', value=1.)
scenario.add_par('output', imports)
grid = make_df(base_output, technology='electricity_grid', commodity='electricity',
level='final', value=0.873)
scenario.add_par('output', grid)
bulb = make_df(base_output, technology='bulb', commodity='light',
level='useful', value=1.)
scenario.add_par('output', bulb)
cfl = make_df(base_output, technology='cfl', commodity='light',
level='useful', value=1.)
scenario.add_par('output', cfl)
app = make_df(base_output, technology='appliances', commodity='other_electricity',
level='useful', value=1.)
scenario.add_par('output', app)
coal = make_df(base_output, technology='coal_ppl', commodity='electricity',
level='secondary', value=1.)
scenario.add_par('output', coal)
gas = make_df(base_output, technology='gas_ppl', commodity='electricity',
level='secondary', value=1.)
scenario.add_par('output', gas)
oil = make_df(base_output, technology='oil_ppl', commodity='electricity',
level='secondary', value=1.)
scenario.add_par('output', oil)
bio = make_df(base_output, technology='bio_ppl', commodity='electricity',
level='secondary', value=1.)
scenario.add_par('output', bio)
hydro = make_df(base_output, technology='hydro_ppl', commodity='electricity',
level='secondary', value=1.)
scenario.add_par('output', hydro)
wind = make_df(base_output, technology='wind_ppl', commodity='electricity',
level='secondary', value=1.)
scenario.add_par('output', wind)
solar_pv = make_df(base_output, technology='solar_pv_ppl', commodity='electricity',
level='final', value=1.)
scenario.add_par('output', solar_pv)
# +
base_technical_lifetime = {
'node_loc': country,
'year_vtg': horizon,
'unit': 'y',
}
lifetimes = {
'coal_ppl': 40,
'gas_ppl': 30,
'oil_ppl': 30,
'bio_ppl': 30,
'hydro_ppl': 60,
'wind_ppl': 20,
'solar_pv_ppl': 20,
'bulb': 1,
'cfl': 10,
}
for tec, val in lifetimes.items():
df = make_df(base_technical_lifetime, technology=tec, value=val)
scenario.add_par('technical_lifetime', df)
# +
base_capacity_factor = {
'node_loc': country,
'year_vtg': vintage_years,
'year_act': act_years,
'time': 'year',
'unit': '-',
}
capacity_factor = {
'coal_ppl': 0.85,
'gas_ppl': 0.75,
'oil_ppl': 0.75,
'bio_ppl': 0.75,
'hydro_ppl': 0.5,
'wind_ppl': 0.2,
'solar_pv_ppl': 0.15,
'bulb': 0.1,
'cfl': 0.1,
}
for tec, val in capacity_factor.items():
df = make_df(base_capacity_factor, technology=tec, value=val)
scenario.add_par('capacity_factor', df)
# -
# ### Technoeconomic Parameters
# +
base_inv_cost = {
'node_loc': country,
'year_vtg': horizon,
'unit': 'USD/kW',
}
# Adding a new unit to the library
mp.add_unit('USD/kW')
# in $ / kW (specific investment cost)
costs = {
'coal_ppl': 1500,
'gas_ppl': 870,
'oil_ppl': 950,
'hydro_ppl': 3000,
'bio_ppl': 1600,
'wind_ppl': 1100,
'solar_pv_ppl': 4000,
'bulb': 5,
'cfl': 900,
}
for tec, val in costs.items():
df = make_df(base_inv_cost, technology=tec, value=val)
scenario.add_par('inv_cost', df)
# +
base_fix_cost = {
'node_loc': country,
'year_vtg': vintage_years,
'year_act': act_years,
'unit': 'USD/kWa',
}
# Adding a new unit to the library
mp.add_unit('USD/kWa')
# in $ / kW / year (every year a fixed quantity is destinated to cover part of the O&M costs
# based on the size of the plant, e.g. lightning, labor, scheduled maintenance, etc.)
costs = {
'coal_ppl': 40,
'gas_ppl': 25,
'oil_ppl': 25,
'hydro_ppl': 60,
'bio_ppl': 30,
'wind_ppl': 40,
'solar_pv_ppl': 25,
}
for tec, val in costs.items():
df = make_df(base_fix_cost, technology=tec, value=val)
scenario.add_par('fix_cost', df)
# +
base_var_cost = {
'node_loc': country,
'year_vtg': vintage_years,
'year_act': act_years,
'mode': 'standard',
'time': 'year',
'unit': 'USD/kWa',
}
# Variable O&M (costs associatied to the degradation of equipment when the plant is functioning
# per unit of energy produced)
# kWa = kW·year = 8760 kWh. Therefore this costs represents USD per 8760 kWh of energy.
# Do not confuse with fixed O&M units.
#var O&M in $ / MWh
costs = {
'coal_ppl': 24.4,
'gas_ppl': 42.4,
'oil_ppl': 77.8,
'bio_ppl': 48.2,
'electricity_grid': 47.8,
}
for tec, val in costs.items():
df = make_df(base_var_cost, technology=tec, value=val * 8760. / 1e3) # to convert it into USD/kWa
scenario.add_par('var_cost', df)
# -
# ## Dynamic Behavior Parameters
# In this section the following parameters will be added to the different technologies:
# - `bound_activity_up`
# - `bound_activity_lo`
# - `bound_new_capacity_up`
# - `initial_activity_up`
# - `growth_activity_up`
#
# As stated in the **Introduction**, a full list of parameters can be found in the *MESSAGEix* documentation. Specifically for this list, please refer to the section [Bounds on capacity and activity](https://docs.messageix.org/en/stable/model/MESSAGE/parameter_def.html#bounds-on-capacity-and-activity)
# +
base_growth = {
'node_loc': country,
'year_act': horizon[1:],
'value': 0.05,
'time': 'year',
'unit': '%',
}
growth_technologies = [
"coal_ppl",
"gas_ppl",
"oil_ppl",
"bio_ppl",
"hydro_ppl",
"wind_ppl",
"solar_pv_ppl",
"cfl",
"bulb",
]
for tec in growth_technologies:
df = make_df(base_growth, technology=tec)
scenario.add_par('growth_activity_up', df)
# +
base_initial = {
'node_loc': country,
'year_act': horizon[1:],
'time': 'year',
'unit': '%',
}
for tec in lights:
df = make_df(base_initial, technology=tec, value=0.01 * light_demand['value'].loc[horizon[1:]])
scenario.add_par('initial_activity_up', df)
# +
base_activity = {
'node_loc': country,
'year_act': [2010],
'mode': 'standard',
'time': 'year',
'unit': 'GWa',
}
# in GWh - from IEA Electricity Output
activity = {
'coal_ppl': 7184,
'gas_ppl': 14346,
'oil_ppl': 1275,
'hydro_ppl': 38406,
'bio_ppl': 4554,
'wind_ppl': 2064,
'solar_pv_ppl': 89,
'import': 2340,
'cfl': 0,
}
#MODEL CALIBRATION: by inserting an upper and lower bound to the same quantity we are ensuring
#that the model is calibrated at that value that year, so we are at the right starting point.
for tec, val in activity.items():
df = make_df(base_activity, technology=tec, value=val / 8760.)
scenario.add_par('bound_activity_up', df)
scenario.add_par('bound_activity_lo', df)
# +
base_capacity = {
'node_loc': country,
'year_vtg': [2010],
'unit': 'GW',
}
cf = pd.Series(capacity_factor)
act = pd.Series(activity)
capacity = (act / 8760 / cf).dropna().to_dict()
for tec, val in capacity.items():
df = make_df(base_capacity, technology=tec, value=val)
scenario.add_par('bound_new_capacity_up', df)
# +
base_activity = {
'node_loc': country,
'year_act': horizon[1:],
'mode': 'standard',
'time': 'year',
'unit': 'GWa',
}
# in GWh - base value from IEA Electricity Output
keep_activity = {
'hydro_ppl': 38406,
'bio_ppl': 4554,
'import': 2340,
}
for tec, val in keep_activity.items():
df = make_df(base_activity, technology=tec, value=val / 8760.)
scenario.add_par('bound_activity_up', df)
# -
# ## Emissions
scenario.add_set('emission', 'CO2')
scenario.add_cat('emission', 'GHGs', 'CO2')
# +
base_emissions = {
'node_loc': country,
'year_vtg': vintage_years,
'year_act': act_years,
'mode': 'standard',
'unit': 'tCO2/kWa',
}
# adding new units to the model library (needed only once)
mp.add_unit('tCO2/kWa')
mp.add_unit('MtCO2')
emissions = {
'coal_ppl': ('CO2', 0.854), # units: tCO2/MWh
'gas_ppl': ('CO2', 0.339), # units: tCO2/MWh
'oil_ppl': ('CO2', 0.57), # units: tCO2/MWh
}
for tec, (species, val) in emissions.items():
df = make_df(base_emissions, technology=tec, emission=species, value=val * 8760. / 1000) #to convert tCO2/MWh into tCO2/kWa
scenario.add_par('emission_factor', df)
# -
# ## Commit the datastructure and solve the model
comment = 'initial commit for Austria model'
scenario.commit(comment)
scenario.set_as_default()
scenario.solve()
# + jupyter={"name": "solve-objective-value"}
scenario.var('OBJ')['lvl']
# -
# # Plotting Results
from tools import Plots
p = Plots(scenario, country)
p.plot_new_capacity(baseyear=True, subset=plants)
p.plot_new_capacity(baseyear=True, subset=lights)
p.plot_capacity(baseyear=True, subset=plants)
p.plot_capacity(baseyear=True, subset=lights)
p.plot_demand(light_demand, elec_demand)
p.plot_activity(baseyear=True, subset=plants)
p.plot_activity(baseyear=True, subset=lights)
p.plot_prices(baseyear=False, subset=['light', 'other_electricity'])
mp.close_db()
| tutorial/Austrian_energy_system/austria.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **Experiment for obtaining 24 Hr prediction from Dense Model in rainymotion library**
#
# Author: <NAME>
#
# File use: For predicting 24 Hr precipitation images.
#
# Date Created: 19-03-21
#
# Last Updated: 19-03-21
#
# Python version: 3.8.2
# +
import h5py
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
import scipy.misc
from rainymotion.models import Dense
import os
import cv2
import pandas as pd
import wradlib.ipol as ipol # for interpolation
from rainymotion import metrics
from rainymotion import utils
from scipy.ndimage import map_coordinates
import timeit
from matplotlib.cm import get_cmap
os.environ['PROJ_LIB'] = '/anaconda3/pkgs/proj4-5.2.0-h0a44026_1/share/proj/'
from mpl_toolkits.basemap import Basemap
import imageio
#from tvl1sindysupport import tvl1utilities -in future our own library
# +
#For plotting map - currently using function as in source code Need to change to Cartopy
def plotMap(title,img, lat1, lat2, long1, long2, outputPath,last=0):
(height, width) = img.shape
# print(img.min(), img.max())
intensity = np.asarray(img, dtype=np.float32)
# print(intensity.min(), intensity.max())
#intensity_level = convert_rep_to_level(intensity).reshape(height, width)
# print(intensity.min(), intensity.max())
intensity_level = np.flipud(intensity)
dLon = (long2 - long1) / width
dLat = (lat2 - lat1) / height
lon = np.arange(long1, long2, dLon)
lat = np.arange(lat1, lat2, dLat)
lons, lats = np.meshgrid(lon, lat)
# print(lons.shape, lats.shape)
fig = plt.figure(figsize=(12, 8))
# Set up Basemap instance
m = Basemap(projection="cyl",
llcrnrlon=long1, urcrnrlon=long2,
llcrnrlat=lat1, urcrnrlat=lat2,
resolution='h')
# Add geographic outlines
m.drawcoastlines(color='black')
m.drawstates()
m.drawcountries()
m.drawmeridians(np.arange(long1, long2, 1), labels=[True, False, False, True])
m.drawparallels(np.arange(lat1, lat2, 1), labels=[True, False, True, False])
#m.drawmeridians(np.arange(new_lon_min, new_lon_max, 1), labels=[False, False, False, False])
#m.drawparallels(np.arange(new_lat_min, new_lat_max, 1), labels=[False, False, False, False])
# Plot Data
#cs = m.contourf(lons, lats, intensity_level, shading='flat', levels=list(range(1, 65)), cmap=get_cmap("jet"))
#cs = m.contourf(lons, lats, intensity_level,shading='flat', levels=list(range(1,65)), cmap=get_cmap("gist_earth"))
cs = m.contourf(lons, lats, intensity_level,shading='flat', levels=list(range(1,65)), cmap=discrete_cmap(8,"jet"))
# Add Colorbar
if last==1:
cb = plt.colorbar(cs ,shrink=1.0) #, extend='both')
# Add Title
plt.title(title)
plt.savefig(outputPath, bbox_inches='tight', pad_inches=0.0)
plt.close()
# -
# For reading data from .h5 files see http://docs.h5py.org/en/stable/quick.html
def readInputData(inputFile):
initialDataSetNo =60 # The dataset no. to start with
frames = []
file = h5py.File(inputFile, 'r')
datasets = list(file.keys())
print(len(datasets)) # There are 178 datasets in this file
for i in range(3):
print('The item is',datasets[i+initialDataSetNo])
dset = file[datasets[i+initialDataSetNo]]
data=np.asarray(dset.value)
frames.append(data)
outFrameName=datasets[i+initialDataSetNo]+'_'+str(i)+'.png'
matplotlib.image.imsave(outFrameName, frames[i])
frames = np.stack(frames, axis=0)
file.close()
print(frames.shape)
return frames
# Radar images - For example,to read radar images of Typhoon Faxai
def readRadarImages(inputFolder, startHr,startMin, timeStep,height,width, noOfImages, fileType):
files = (os.listdir(inputFolder))
files.sort()
inputRadarImages = []
firstImgTime = startHr*100+startMin
listTime = [startHr*100+startMin]
startTime = startHr
initialTime = startHr
startTime = startTime*100+startMin
for i in range(noOfImages-1):
if "60" in str(startTime+10):
startTime = initialTime + 1
initialTime = startTime
startTime = startTime*100
listTime.append((startTime))
else:
listTime.append((startTime)+10)
startTime = startTime+10
print(listTime)
for itemNo in range(noOfImages):
for fileName in files:
if str(listTime[itemNo]) in fileName:
#print(fileName)
if fileName.endswith(fileType):
inputFileName =inputFolder+'/'+fileName
fd = open(inputFileName,'rb')
#print(inputFileName)
# straight to numpy data (no buffering)
recentFrame = np.fromfile(fd, dtype = np.dtype('float32'), count = 2*height*width)
recentFrame = np.reshape(recentFrame,(height,width))
recentFrame = recentFrame.astype('float16')
inputRadarImages.append(recentFrame)
inputRadarImages = np.stack(inputRadarImages, axis=0)
return inputRadarImages
# +
# Common Initialization
eventName = "TyphoneFaxai"
eventDate ="20190908"
eventNameDate = eventName + "_" + eventDate
# For radar images
inputFolder = "./ForExperiments/Exp1/RadarImages/TyphoonFaxai/For21/"
outputFolder= "./ForExperiments/Exp1/Results/"
height = 781
width = 561
fileType='.bin'
timeStep = 10 # for Japan Radar Data
modelName = "Sparse SD"
startHr = 20
startMin = 30
noOfImages = 3
leadSteps = 12
stepRainyMotion = 5 # 5 minutes
outputFilePath = outputFolder+modelName+'_'
outputFilePath = outputFilePath + eventNameDate
print(outputFilePath)
#Latitude and Longitude of Typhoon Faxai
lat1 = 32.5
lat2 = 39
long1 = 136
long2 = 143
# -
# **1.3 Dense**
# +
modelName = "Dense"
startHr = 20
startMin= 40
noOfImages = 2# Sparse Model needs 24 frames
predStartHr = 2100
step = 5
leadSteps = 12
outputFilePath = outputFolder+'/'+modelName+'/'
outputFilePath = outputFilePath + eventNameDate
print(outputFilePath)
# of_method = "DIS"
# direction = "backward"
# advection = "constant-vector"
# interpolation = "idw"
model = Dense()
model.input_data = readRadarImages(inputFolder, startHr,startMin,timeStep,height,width, noOfImages, fileType)
start = timeit.timeit()
nowcastDense = model.run()
end = timeit.timeit()
sparseTime = end - start
print("Dense took ",end - start)
nowcastDense.shape
print("Saving the nowcast images. Please wait...")
for i in range(leadSteps):
outFrameName = outputFilePath + '_'+str(predStartHr+(i*5))+'.png'
#matplotlib.image.imsave(outFrameName, nowcastDense[i])
if i == leadSteps-1:
last = 1
else:
last = 0
plotMap(modelName+' '+str(predStartHr+(i*5)),nowcastDense[i], lat1, lat2, long1, long2, outFrameName,last)
print("Finished Dense model nowcasting!")
# -
import numpy as np
def getGroundTruthImages(recentFramePath,groundTruthTime,height,width,fileType):
files = (os.listdir(recentFramePath))
files.sort()
groundTruthImages = []
for fileName in files:
if fileName.endswith(fileType):
#if groundTruthTime in fileName:
#print("The ground truth at %s is available",groundTruthTime)
inputFileName =recentFramePath+'/'+fileName
fd = open(inputFileName,'rb')
#print(inputFileName)
# straight to numpy data (no buffering)
recentFrame = np.fromfile(fd, dtype = np.dtype('float32'), count = 2*height*width)
recentFrame = np.reshape(recentFrame,(height,width))
recentFrame = recentFrame.astype('float16')
#print(recentFrame.shape)
groundTruthImages.append(recentFrame)
#else:
# print("Sorry, unable to find file.")
groundTruthImages = np.moveaxis(np.dstack(groundTruthImages), -1, 0)
#print(groundTruthImages.shape)
return groundTruthImages
# **2.1 Mean Absolute Error**
# +
from rainymotion import metrics
def MAE(obs, sim):
"""
Mean absolute error
Reference: https://en.wikipedia.org/wiki/Mean_absolute_error
Args:
obs (numpy.ndarray): observations
sim (numpy.ndarray): simulations
Returns:
float: mean absolute error between observed and simulated values
"""
obs = obs.flatten()
sim = sim.flatten()
return np.mean(np.abs(sim - obs))
def prep_clf(obs, sim, threshold=0.1):
obs = np.where(obs >= threshold, 1, 0)
sim = np.where(sim >= threshold, 1, 0)
# True positive (TP)
hits = np.sum((obs == 1) & (sim == 1))
# False negative (FN)
misses = np.sum((obs == 1) & (sim == 0))
# False positive (FP)
falsealarms = np.sum((obs == 0) & (sim == 1))
# True negative (TN)
correctnegatives = np.sum((obs == 0) & (sim == 0))
return hits, misses, falsealarms, correctnegatives
def CSI(obs, sim, threshold=0.1):
"""
CSI - critical success index
details in the paper:
<NAME>., & <NAME>. (2017).
Operational Application of Optical Flow Techniques to Radar-Based
Rainfall Nowcasting.
Atmosphere, 8(3), 48. https://doi.org/10.3390/atmos8030048
Args:
obs (numpy.ndarray): observations
sim (numpy.ndarray): simulations
threshold (float) : threshold for rainfall values binaryzation
(rain/no rain)
Returns:
float: CSI value
"""
hits, misses, falsealarms, correctnegatives = prep_clf(obs=obs, sim=sim,
threshold=threshold)
return hits / (hits + misses + falsealarms)
# -
event_name = "<NAME> 09 August, 2019"
start = "21:00"
end = "21:50"
t = ['21:00','21:10','21:20','21:30','21:40', '21:50']
# +
groundTruthPath = "./ForExperiments/Exp1/GroundTruth/TyphoonFaxai"
groundTruthTime = "2100"
groundTruthImgs = getGroundTruthImages(groundTruthPath,groundTruthTime,height,width,fileType)
#print("Ground truth images shape", groundTruthImgs.shape)
MAEDense = []
noOfPrecipitationImages = 6
j = 0 # using another index to skip 5min interval data from rainymotion
for i in range(noOfPrecipitationImages):
#print(groundTruthImgs[i].shape)
#print(nowcast[j].shape)
mae = MAE(groundTruthImgs[i],nowcastDense[j])
MAEDense.append(mae)
j = j + 2
# -
# **2.2 Critical Success Index**
# +
CSIDense = []
noOfPrecipitationImages = 6
thres=1.0 #0.1 default
j = 0 # using another index to skip 5min interval data from rainymotion
for i in range(noOfPrecipitationImages):
#print(groundTruthImgs[i].shape)
#print(nowcast[j].shape)
csi = CSI(groundTruthImgs[i],nowcastDense[j],thres)
CSIDense.append(csi)
j = j + 2
# +
print(MAEDense)
# +
print(CSIDense)
| examples/Dense24HrPrediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Approximate q-learning
#
# In this notebook you will teach a __tensorflow__ neural network to do Q-learning.
# __Frameworks__ - we'll accept this homework in any deep learning framework. This particular notebook was designed for tensorflow, but you will find it easy to adapt it to almost any python-based deep learning framework.
#XVFB will be launched if you run on a server
import os
if os.environ.get("DISPLAY") is not str or len(os.environ.get("DISPLAY"))==0:
# !bash ../xvfb start
# %env DISPLAY=:1
import gym
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
# +
env = gym.make("CartPole-v0").env
env.reset()
n_actions = env.action_space.n
state_dim = env.observation_space.shape
plt.imshow(env.render("rgb_array"))
# -
# # Approximate (deep) Q-learning: building the network
#
# To train a neural network policy one must have a neural network policy. Let's build it.
#
#
# Since we're working with a pre-extracted features (cart positions, angles and velocities), we don't need a complicated network yet. In fact, let's build something like this for starters:
#
# 
#
# For your first run, please only use linear layers (L.Dense) and activations. Stuff like batch normalization or dropout may ruin everything if used haphazardly.
#
# Also please avoid using nonlinearities like sigmoid & tanh: agent's observations are not normalized so sigmoids may become saturated from init.
#
# Ideally you should start small with maybe 1-2 hidden layers with < 200 neurons and then increase network size if agent doesn't beat the target score.
import tensorflow as tf
import keras
import keras.layers as L
tf.reset_default_graph()
sess = tf.InteractiveSession()
keras.backend.set_session(sess)
# +
network = keras.models.Sequential()
network.add(L.InputLayer(state_dim))
# let's create a network for approximate q-learning following guidelines above
network.add(L.Dense(100, activation='relu'))
network.add(L.Dense(100, activation='relu'))
network.add(L.Dense(n_actions, activation='linear'))
# -
import random
def get_action(state, epsilon=0):
"""
sample actions with epsilon-greedy policy
recap: with p = epsilon pick random action, else pick action with highest Q(s,a)
"""
q_values = network.predict(state[None])[0]
###YOUR CODE
action = np.random.choice(n_actions)
chosen_action = np.argmax(q_values) if random.random() > epsilon else action
return chosen_action
# +
assert network.output_shape == (None, n_actions), "please make sure your model maps state s -> [Q(s,a0), ..., Q(s, a_last)]"
assert network.layers[-1].activation == keras.activations.linear, "please make sure you predict q-values without nonlinearity"
# test epsilon-greedy exploration
s = env.reset()
assert np.shape(get_action(s)) == (), "please return just one action (integer)"
for eps in [0., 0.1, 0.5, 1.0]:
state_frequencies = np.bincount([get_action(s, epsilon=eps) for i in range(10000)], minlength=n_actions)
best_action = state_frequencies.argmax()
assert abs(state_frequencies[best_action] - 10000 * (1 - eps + eps / n_actions)) < 200
for other_action in range(n_actions):
if other_action != best_action:
assert abs(state_frequencies[other_action] - 10000 * (eps / n_actions)) < 200
print('e=%.1f tests passed'%eps)
# -
# ### Q-learning via gradient descent
#
# We shall now train our agent's Q-function by minimizing the TD loss:
# $$ L = { 1 \over N} \sum_i (Q_{\theta}(s,a) - [r(s,a) + \gamma \cdot max_{a'} Q_{-}(s', a')]) ^2 $$
#
#
# Where
# * $s, a, r, s'$ are current state, action, reward and next state respectively
# * $\gamma$ is a discount factor defined two cells above.
#
# The tricky part is with $Q_{-}(s',a')$. From an engineering standpoint, it's the same as $Q_{\theta}$ - the output of your neural network policy. However, when doing gradient descent, __we won't propagate gradients through it__ to make training more stable (see lectures).
#
# To do so, we shall use `tf.stop_gradient` function which basically says "consider this thing constant when doingbackprop".
# Create placeholders for the <s, a, r, s'> tuple and a special indicator for game end (is_done = True)
states_ph = keras.backend.placeholder(dtype='float32', shape=(None,) + state_dim)
actions_ph = keras.backend.placeholder(dtype='int32', shape=[None])
rewards_ph = keras.backend.placeholder(dtype='float32', shape=[None])
next_states_ph = keras.backend.placeholder(dtype='float32', shape=(None,) + state_dim)
is_done_ph = keras.backend.placeholder(dtype='bool', shape=[None])
# +
#get q-values for all actions in current states
predicted_qvalues = network(states_ph)
#select q-values for chosen actions
predicted_qvalues_for_actions = tf.reduce_sum(predicted_qvalues * tf.one_hot(actions_ph, n_actions), axis=1)
# +
gamma = 0.99
# compute q-values for all actions in next states
predicted_next_qvalues = network(next_states_ph)
# compute V*(next_states) using predicted next q-values
next_state_values = tf.reduce_max(predicted_next_qvalues, axis=1)
# compute "target q-values" for loss - it's what's inside square parentheses in the above formula.
target_qvalues_for_actions = rewards_ph + gamma*next_state_values
# at the last state we shall use simplified formula: Q(s,a) = r(s,a) since s' doesn't exist
target_qvalues_for_actions = tf.where(is_done_ph, rewards_ph, target_qvalues_for_actions)
# +
#mean squared error loss to minimize
loss = (predicted_qvalues_for_actions - tf.stop_gradient(target_qvalues_for_actions)) ** 2
loss = tf.reduce_mean(loss)
# training function that resembles agent.update(state, action, reward, next_state) from tabular agent
train_step = tf.train.AdamOptimizer(1e-4).minimize(loss)
# -
assert tf.gradients(loss, [predicted_qvalues_for_actions])[0] is not None, "make sure you update q-values for chosen actions and not just all actions"
assert tf.gradients(loss, [predicted_next_qvalues])[0] is None, "make sure you don't propagate gradient w.r.t. Q_(s',a')"
assert predicted_next_qvalues.shape.ndims == 2, "make sure you predicted q-values for all actions in next state"
assert next_state_values.shape.ndims == 1, "make sure you computed V(s') as maximum over just the actions axis and not all axes"
assert target_qvalues_for_actions.shape.ndims == 1, "there's something wrong with target q-values, they must be a vector"
# ### Playing the game
def generate_session(t_max=1000, epsilon=0, train=False):
"""play env with approximate q-learning agent and train it at the same time"""
total_reward = 0
s = env.reset()
for t in range(t_max):
a = get_action(s, epsilon=epsilon)
next_s, r, done, _ = env.step(a)
if train:
sess.run(train_step,{
states_ph: [s], actions_ph: [a], rewards_ph: [r],
next_states_ph: [next_s], is_done_ph: [done]
})
total_reward += r
s = next_s
if done: break
return total_reward
epsilon = 0.5
for i in range(1000):
session_rewards = [generate_session(epsilon=epsilon, train=True) for _ in range(100)]
print("epoch #{}\tmean reward = {:.3f}\tepsilon = {:.3f}".format(i, np.mean(session_rewards), epsilon))
epsilon *= 0.99
assert epsilon >= 1e-4, "Make sure epsilon is always nonzero during training"
if np.mean(session_rewards) > 300:
print ("You Win!")
break
# ### How to interpret results
#
#
# Welcome to the f.. world of deep f...n reinforcement learning. Don't expect agent's reward to smoothly go up. Hope for it to go increase eventually. If it deems you worthy.
#
# Seriously though,
# * __ mean reward__ is the average reward per game. For a correct implementation it may stay low for some 10 epochs, then start growing while oscilating insanely and converges by ~50-100 steps depending on the network architecture.
# * If it never reaches target score by the end of for loop, try increasing the number of hidden neurons or look at the epsilon.
# * __ epsilon__ - agent's willingness to explore. If you see that agent's already at < 0.01 epsilon before it's is at least 200, just reset it back to 0.1 - 0.5.
# ### Record videos
#
# As usual, we now use `gym.wrappers.Monitor` to record a video of our agent playing the game. Unlike our previous attempts with state binarization, this time we expect our agent to act ~~(or fail)~~ more smoothly since there's no more binarization error at play.
#
# As you already did with tabular q-learning, we set epsilon=0 for final evaluation to prevent agent from exploring himself to death.
#record sessions
import gym.wrappers
env = gym.wrappers.Monitor(gym.make("CartPole-v0"),directory="videos",force=True)
sessions = [generate_session(epsilon=0, train=False) for _ in range(100)]
env.close()
# +
#show video
from IPython.display import HTML
import os
video_names = list(filter(lambda s:s.endswith(".mp4"),os.listdir("./videos/")))
HTML("""
<video width="640" height="480" controls>
<source src="{}" type="video/mp4">
</video>
""".format("./videos/"+video_names[-1])) #this may or may not be _last_ video. Try other indices
# -
# ---
# ### Submit to coursera
# %load_ext autoreload
# %autoreload 2
from submit import submit_cartpole
submit_cartpole(generate_session, '', '')
| Practical Reinforcement Learning/Week4_approx/practice_approx_qlearning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_tensorflow_p27
# language: python
# name: conda_tensorflow_p27
# ---
from keras.models import load_model
import numpy as np
import os
model = None
if os.path.isfile('./additionrnn_model.h5') and os.path.isfile('./additionrnn_weightsandbias.h5'):
model = load_model('additionrnn_model.h5')
model.load_weights('additionrnn_weightsandbias.h5')
class CharacterTable(object):
def __init__(self, chars):
self.chars = sorted(set(chars))
self.char_indices = dict((c, i) for i, c in enumerate(self.chars))
self.indices_char = dict((i, c) for i, c in enumerate(self.chars))
def encode(self, C, num_rows):
x = np.zeros((num_rows, len(self.chars)))
for i, c in enumerate(C):
x[i, self.char_indices[c]] = 1
return x
def decode(self, x, calc_argmax=True):
if calc_argmax:
x = x.argmax(axis=-1)
return ''.join(self.indices_char[x] for x in x)
class colors:
ok = '\033[92m'
fail = '\033[91m'
close = '\033[0m'
chars = '0123456789+ '
ctable = CharacterTable(chars)
# +
DIGITS = 9
MAXLEN = DIGITS + 1 + DIGITS
a=923676789
b=167456780
c=a+b
q = '{}+{}'.format(a, b)
query = q + ' ' * (MAXLEN - len(q))
query = query[::-1]
ans=str(c)
ans += ' ' * (DIGITS + 1 - len(ans))
X = np.array([ctable.encode(query, MAXLEN)])
print(X.shape)
y = model.predict_classes(X, verbose=0)
y = ctable.decode(y[0], calc_argmax=False)
if ans == y:
print q,'=',ans,colors.ok + '☑' + colors.close, y
else:
print q,'=',ans,colors.fail + '☒' + colors.close, y
# -
| seq2seq/AdditionRNNTest.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Rascal Kernel
# language: java
# name: javakernel
# ---
# +
module execution
function levelMask(img<int32_t> param[x,y,c]):
r[x] -> randomInt();
level[x,y,c] -> do:
if c==2 || c==0
if param[x,y,c] == 67
if r[x]==1
10
else
30
end
else
param[x,y,c]
end
else
if param[x,y,c] == 67 && (r[x]==10)
10
else
param[x,y,c]
end
end
od;
return level;
end
exec tst1{
img<int32_t> img1= loadImage("images/rgb.png");
img<int32_t> output = run(levelMask(img1), img1.width(), img1.height(), img1.channels());
save(output, "images/levelOutput.png");
##renderImage(output);
##assert output != input;
}
exec test2{
img<int32_t> img2= loadImage("images/gray.png");
img<int32_t> output2 = run(levelMask(img2), img2.width(), img2.height(), img2.channels());
save(output2, "images/output/levelOutput2.png");
}
| rascal-kernel/main/resources/nb-examples/amalga.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + tags=["hide-cell"]
import numpy as np
import matplotlib.pyplot as plt
plt.style.use('fivethirtyeight')
from IPython.core.display import SVG
# -
# # Mechanical advantage
#
# In the diagram below, there is a 50-N force, $F$, applied downwards on a
# constrained linkage system. There is a restoring force, $R$, that
# maintains a slow change in the angle of the mechanism, $\theta$, as it
# changes from $89^o$ to $-89^o$. Your
# goal is to determine the necessary restoring force $R$ as a
# function of angle $\theta$.
#
# 
#
# ## Kinematics - geometry of motion
#
# This system has two rigid bodies connected at point $A$ by a pin and the
# whole system is pinned to the ground at $O$. Point $B$ slides
# along a horizontal surface. The total degrees of freedom are
#
# 3 (link 1) + 3 (link 2) - 2 constraints ($O$) - 2 constraints ($A$) - 1
# constraint ($B$) - 1 constraint ($\theta$-fixed) = __0 DOF__
#
# For a given angle $\theta$, there can only be one configuration in the
# system. Create the constraint equations using the relative position of
# point $A$ as such
#
# $\mathbf{r}_A = \mathbf{r}_{A/B} + \mathbf{r}_B$
#
# 
#
# where
#
# $\mathbf{r}_A = L(\cos\theta \hat{i}+ \sin\theta \hat{j})$
#
# $\mathbf{r}_B = d\hat{i}$
#
# $\mathbf{r}_{A/B} = L(-\cos\theta \hat{i} + \sin\theta \hat{j})$
#
# solving for the $\hat{i}-$ and $\hat{j}-$components creates two
# equations that describe the state of the system
#
# * $\theta = \theta$ which states because we have an isosceles triangle,
# two angles have to be equal
# * $d = 2 L \cos\theta$ so $\mathbf{r}_{B} = 2L \cos\theta \hat{i}$
# + tags=["hide-input"]
theta = np.linspace(89,-89)*np.pi/180
d = 2*1*np.cos(theta)
plt.plot(theta*180/np.pi, d)
plt.xticks(np.arange(-90,91,30))
plt.xlabel(r'$\theta$ (deg)')
plt.xlim(90,-90)
plt.ylabel('d (m)');
# -
# ## Kinetics - applied forces and constraint forces
#
# The applied force, $F=50~N$ is constant, but $R$ is dependent upon the
# geometry of the system. You solved for the kinematics in the first part,
# here you can use the Newton-Euler equations to solve for $R$ given $\theta$ and
# $F$. Separate the system into the left and right links,
#
# 
# The Newton-Euler equations:
#
# * $\mathbf{F} = m\mathbf{a} = \mathbf{0}$ links moving slowly
# * $M_G = I\alpha = 0$ links rotating slowly
# Newton-Euler equations for the left bar:
#
# 1. $\mathbf{F}\cdot \hat{i} = N_{x1}+N_{x2} = 0$
# 2. $\mathbf{F}\cdot \hat{i} = N_{y1}+N_{y2} - F = 0$
# 3. $M_O = l\hat{b}_1 \times (-F\hat{j} + N_{x2}\hat{i} + N_{y2}\hat{j})
# = 0$
#
# Newton-Euler equations for the right bar:
#
# 1. $\mathbf{F}\cdot \hat{i} = -N_{x2}-R = 0$
# 2. $\mathbf{F}\cdot \hat{i} = -N_{y2}+N_{y3} = 0$
# 3. $M_A = l\hat{c}_1 \times (-R\hat{i} + N_{y3}\hat{j})= 0$
#
# > __Note:__ Don't count $F$ twice! You can use the applied force on the
# > left or right, but not both. Try solving the equations placing it on
# > the right bar.
#
# The four equations for $\mathbf{F}$ relate the reaction forces
# $\mathbf{N}_{1},~\mathbf{N}_{2},~and~\mathbf{N}_{3}$. The two moment
# equations relate $R$ to $F$ and $\theta$ as such
#
# $N_{y2}L\cos\theta - RL\sin\theta = 0$
#
# and
#
# $-FL\cos\theta +N_{y2}\cos\theta+RL\sin\theta = 0$
#
# combining there results
#
# $F\cos\theta = 2R\sin\theta\rightarrow R = \frac{F}{2}\cot\theta$
# + tags=["hide-input"]
R = 50/2*np.tan(theta)**-1
plt.plot(theta*180/np.pi, R)
plt.xticks(np.arange(-90,91,30))
plt.xlabel(r'$\theta$ (deg)')
plt.ylabel('restoring force, R (N)')
plt.xlim(90,-90)
plt.ylim(-200,200);
# -
# ## Wrapping up
#
# Take a look at the mechanical advantage and disadvantage this system can create. For angles close to $\theta\approx 90^o$, the restoring force is close to zero. In this case, the applied force is mostly directed at constraints on the system. When the angles are close to $\theta \approx 0^o$, the required restoring force can be $>100\times$ the input force.
#
# Have you seen this type of linkage system in engineering devices?
| _build/jupyter_execute/module_02/mechanical-advantage.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from alpha_vantage.timeseries import TimeSeries
from alpha_vantage.techindicators import TechIndicators
from alpha_vantage.sectorperformance import SectorPerformances
from alpha_vantage.cryptocurrencies import CryptoCurrencies
from alpha_vantage.foreignexchange import ForeignExchange
from alpha_vantage.fundamentaldata import FundamentalData
import matplotlib
import matplotlib.pyplot as plt
import os
#making plots bigger
#matplotlib.rcParams['figure.figsize'] = (50, 20)
# magic function (idk what it does)
# #%matplotlib inline
# -
# ## Some Disclaimer
# AlphaVantage isn't perfect. It is a free open source finance API that comes with lots of imcomplete information and functions. Particularly:
#
# - TimeSeries do not have an argument to edit the number of observations. 'outputsize' is limited to 'compact or full'.
# - TechIndicators do not have argument to edit number of observations. 'timeperiod' is refered to the number of observations used to obtain averages, not "to be included as data points".
# - Intraday data are very outdated. You cannot use it for live trading. This is particularly important if you plan on using volume data to make informed trades.
# - at interval = '60min', there are 16 observations per day.
# - at interval = '30min', there are 26 observations per day.
# - Daily observations starts 0830 hrs and ends at 2000 hrs.
#
# ## Time Series
# +
from alpha_vantage.timeseries import TimeSeries
ts = TimeSeries(key = '<KEY>', output_format = 'pandas')
#important to have 'pandas' to print
# +
pton_p, pton_m = ts.get_daily_adjusted(symbol = "PTON", outputsize = 'full')
print(pton_p.head(2))
pton_p['5. adjusted close'].plot()
# choose either '1. open', '2. high', etc. It is found from the columns printed from head()
plt.title('Epic PTON stonks')
plt.show()
# -
# ## Volume Analysis
ti = TechIndicators(key = '<KEY>', output_format = 'pandas')
# ### Chaikin A/D Line
# Accumulation/distribution is a cumulative indicator that uses volume and price to assess whether a stock is being accumulated or distributed. The accumulation/distribution measure seeks to identify divergences between the stock price and volume flow. This provides insight into how strong a trend is.
#
# - The A/D indicator is cumulative, meaning one period's value is added or subtracted from the last.
# - If the price is rising but the indicator is falling this indicates that buying or accumulation volume may not be enough to support the price rise and a price decline could be forthcoming.
# - If the price of an asset is falling but A/D is rising, it signals underlying strength and the price may start to rise.
#
# +
pton_ad, q = ti.get_ad(symbol = 'pton', interval = '30min')
pton_p, pton_m = ts.get_daily(symbol = "pton")
print(pton_p.head(2))
pton_ad.plot()
pton_p['4. close'].plot()
plt.title('AD Indicator for PTON')
plt.show()
# how do you do a side-by-side chart on python?
# note the yellow straight line. It is the price chart that it's so small compared to the volume chart.
# -
# ### Chaikin A/D Oscillator
# As many traders say, price follows volume, and this indicator is used to analyze the current momentum of the price to predict future price movements. Chaikin A/D Oscillator fluctuates around the value zero. Every time that the indicator crosses goes changes from positive to negative, it suggests that the momentum of the price is changing.
#
# A Chaikin Oscillator reading above zero indicates net buying pressure, while below zero registers net selling pressure. Divergence between the indicator and pure price moves are the most common signals from the indicator, and often flag market turning points.
#
# +
pton_ado, q = ti.get_adosc(symbol = 'pton', interval = '30min')
print(pton_ado)
# at interval = '60min', there are 16 observations per day.
# at interval = '30min', there are 26 observations per day.
# Daily observations starts 0830 hrs and ends at 2000 hrs.
pton_p, pton_m = ts.get_daily(symbol = "pton")
print(pton_p.head(2))
pton_ado.plot()
pton_p['4. close'].plot()
plt.title('ADOSC for PTON')
plt.show()
# -
# ### On Balance Volume Value
# On Balance Volume (OBV) measures buying and selling pressure as a cumulative indicator that adds volume on up days and subtracts volume on down days. When the security closes higher than the previous close, all of the day’s volume is considered up-volume. When the security closes lower than the previous close, all of the day’s volume is considered down-volume.
#
# How this indicator works:
# The actual value of the OBV is unimportant; concentrate on its direction.
#
# - When both price and OBV are making higher peaks and higher troughs, the upward trend is likely to continue.
# - When both price and OBV are making lower peaks and lower troughs, the downward trend is likely to continue.
# - During a trading range, if the OBV is rising, accumulation may be taking place—a warning of an upward breakout.
# - During a trading range, if the OBV is falling, distribution may be taking place—a warning of a downward breakout.
# - When price continues to make higher peaks and OBV fails to make higher peaks, the upward trend is likely to stall or fail. This is called a negative divergence.
# - When price continues to make lower troughs and OBV fails to make lower troughs, the downward trend is likely to stall or fail. This is called a positive divergence.
#
#
# +
pton_obv, q = ti.get_obv(symbol = 'pton', interval = '30min')
print(pton_ado)
pton_p, pton_m = ts.get_daily(symbol = "pton")
print(pton_p.head(2))
pton_obv.plot()
pton_p['4. close'].plot()
plt.title('OBV for PTON')
plt.show()
# looks very similar to pton_ad chart.
# -
# ### Money Flow Index
# Instead of looking at the volume (number of shares traded) or price (RSI) as an indicator of identifying overbought or oversold conditions, the Money Flow Index incorporates both price and volume data. For this reason, some analysts call MFI the volume-weighted RSI.
#
# The Formulas for the Money Flow Index Are:
#
# Money Flow Index = 100 − 100/(1 + Money Flow Ratio)
#
# where:
#
# Money Flow Ratio = 14 Period Positive Money Flow / 14 Period Negative Money Flow
# Raw Money Flow = Typical Price * Volume
#
# +
pton_mfi, q = ti.get_mfi(symbol = 'pton', interval = 'daily', time_period = 14)
print(pton_mfi.tail(2))
pton_p, pton_m = ts.get_daily(symbol = "pton")
print(pton_p.tail(2))
pton_mfi.plot()
plt.title('MFI for PTON')
plt.show()
pton_p['4. close'].plot()
plt.title('Price chart for PTON')
plt.show()
# -
# ### Hilbert Transformed Instantaneous Trendline
# The [Hilbert Transform](http://www2.wealth-lab.com/WL5Wiki/HTTrendLine.ashx) is a technique used to generate inphase and quadrature components of a de-trended real-valued "analytic-like" signal (such as a Price Series) in order to analyze variations of the instantaneous phase and amplitude. HTTrendline (or MESA Instantaneous Trendline) returns the Price Series value after the Dominant Cycle of the analytic signal as generated by the Hilbert Transform has been removed. The Dominant Cycle can be thought of as being the "most likely" period (in the range of 10 to 40) of a sine function of the Price Series.
#
# Basically, lots of fancy math going behind that is still out of my expertise to describe.
# +
pton_htt, q = ti.get_ht_trendline(symbol = 'pton', interval = '60min')
print(pton_htt.tail(2))
pton_p, pton_m = ts.get_intraday(symbol = "pton", interval = '60min')
print(pton_p.tail(2))
pton_htt.plot()
pton_p['4. close'].plot()
plt.title('HTT for PTON')
plt.show()
# -
# ### Hilbert Transformed Sine Waves Value
# Again, I have no idea what it does.
# +
pton_hts, q = ti.get_ht_sine(symbol = 'pton', interval = '60min')
print(pton_htt.tail(2))
pton_p, pton_m = ts.get_intraday(symbol = "pton", interval = '60min')
print(pton_p.tail(2))
pton_hts.plot()
#pton_p['4. close'].plot()
plt.title('HTS for PTON')
plt.show()
# -
# ## Moving Averages
# Moving averages are commonly used in finances and are probably too overused to get any useful alpha. Nonetheless, it remains an important indicator at least to understand what the majority of traders are thinking. Most of the time, traders refer to these MA lines (aka lookback period) to confirm which MAs are more useful at predicting the direction of a security.
#
# Mathematically, they are unreliable at determining the value or even the demand of a security (via volume). But well, it's still useful at understanding how technical traders think.
#
# The **Simple Moving Average (SMA)** is calculated by adding the price of an instrument over a number of time periods and then dividing the sum by the number of time periods. The SMA is basically the average price of the given time period, with equal weighting given to the price of each period. **The 'timeperiod' argument within each moving average functions will reflect the statement before it.**
#
# An **exponential moving average (EMA)** places a greater weight and significance on the most recent data points, meaning it reacts more significantly to recent price changes than a simple moving average (SMA), which applies an equal weight to all observations in the period. **Weighted moving averages (WMA)** also assign a heavier weighting to more current data points similar to EMA, just that the formula is different.
#
# Prepare for trouble. Make it double. The **double exponential moving average (DEMA)** is an expansion of EMA, but responds quicker to price changes (less lag) than a normal exponential moving average (EMA). Note that less lag isn't always a good thing because lag helps filter out noise. An indicator with less lag is more prone to reacting to noise or small inconsequential price moves. A 100 periods DEMA will react slower to shorter time frame DEMAs, like 10 periods.
#
# If double isn't enough, there's a **triple exponential moving average (TEMA)** to consider. Obviously, they are more sensitive to price changes than DEMA or EMA and they do not tell you anything about the company or the demand for their shares. But it is still a useful indicator for lookbacks.
#
# Other MA methods that are available in this package include: **Triangular Moving Average (TRIMA), Kaufman Adaptive Moving Average (KAMA), MESA Adaptive Moving Average (MAMA).** There is also **Volume Weighted Moving Average (VWAP)** for intraday analysis.
#
#
# +
#pton_wma, q = ti.get_wma(symbol = 'pton', interval = 'daily', time_period = 20)
#pton_dema, q = ti.get_dema(symbol = 'pton', interval = 'daily', time_period = 20)
#pton_tema, q = ti.get_tema(symbol = 'pton', interval = 'daily', time_period = 20)
#pton_trima, q = ti.get_trima(symbol = 'pton', interval = 'daily', time_period = 20)
pton_sma, q = ti.get_sma(symbol = 'pton', interval = 'daily', time_period = 20)
pton_ema, q = ti.get_ema(symbol = 'pton', interval = 'daily', time_period = 20)
pton_kama, q = ti.get_kama(symbol = 'pton', interval = 'daily', time_period = 20)
pton_mama, q = ti.get_mama(symbol = 'pton', interval = 'daily')
print(pton_sma.tail(2))
pton_p, pton_m = ts.get_daily(symbol = "pton")
print(pton_p.tail(2))
#pton_wma.plot()
#pton_dema.plot()
#pton_tema.plot()
#pton_trima.plot()
pton_sma.plot()
pton_ema.plot()
pton_kama.plot()
pton_mama.plot()
pton_p['4. close'].plot()
plt.title('Moving Averages for PTON')
plt.show()
# You will likely get an error for exceeding AlphaVantage's maximum API call. Simply comment out uninteresting MAs and plot again.
# -
# ## Other Technical Analysis Methods
# ### Moving Average Convergence Divergence
#
# Moving Average Convergence Divergence (MACD) is a trend-following momentum indicator that shows the relationship between two moving averages of a security’s price. [Most commonly](https://school.stockcharts.com/doku.php?id=technical_indicators:moving_average_convergence_divergence_macd), the MACD is calculated by subtracting the "slow" 26-period Exponential Moving Average (EMA) from the "fast" 12-period EMA. Time period can be edited accordingly.
#
# Essential Keyword Arguments:
# - symbol: the symbol for the equity we want to get its data
# - interval: time interval between two conscutive values, supported values are '1min', '5min', '15min', '30min', '60min', 'daily', 'weekly', 'monthly' (default 'daily')
# - fastperiod: Positive integers are accepted (default=None)
# - slowperiod: Positive integers are accepted (default=None)
# - signalperiod: Positive integers are accepted (default=None)
# - fastmatype: Moving average type for the faster moving average. By default, fastmatype=0. Integers 0 - 8 are accepted
# - slowmatype: Moving average type for the slower moving average. By default, slowmatype=0. Integers 0 - 8 are accepted
# - signalmatype: Moving average type for the signal moving average. By default, signalmatype=0. Integers 0 - 8 are accepted
#
# * 0 = Simple Moving Average (SMA),
# * 1 = Exponential Moving Average (EMA),
# * 2 = Weighted Moving Average (WMA),
# * 3 = Double Exponential Moving Average (DEMA),
# * 4 = Triple Exponential Moving Average (TEMA),
# * 5 = Triangular Moving Average (TRIMA),
# * 6 = T3 Moving Average,
# * 7 = Kaufman Adaptive Moving Average (KAMA),
# * 8 = MESA Adaptive Moving Average (MAMA)
#
# +
pton_macd, q = ti.get_macd(symbol = 'pton', interval = 'daily', fastperiod = 12, slowperiod = 26, signalperiod = 9)
print(pton_macd.tail(2))
# On customizing MA for MACD, use 'get_macdext'
# pton_macdext, q = ti.get_macdext(symbol = 'pton', interval = 'daily', fastperiod = 12, slowperiod = 26, signalperiod = 9,
# fastmatype = 1, slowmatype = 1, signalmatype = 1)
# print(pton_macdext.tail(2))
pton_p, pton_m = ts.get_daily(symbol = "pton")
print(pton_p.tail(2))
pton_macd.plot()
#
# pton_macdext.plot()
#
# You will also likely get an error for exceeding AV's call limit. Apparently, macdext() counts as 3 calls.
plt.title('MACD for PTON')
plt.show()
pton_p['4. close'].plot()
plt.title('Price chart for PTON')
plt.show()
# -
# ### Bollinger Bands
# Bollinger Bands are price range plotted at a standard deviation level above and below a simple moving average of the price. Because the distance of the bands is based on standard deviation, they adjust to volatility swings in the underlying price. Bollinger bands help determine whether prices are high or low on a relative basis.
#
# [Commonly used settings](https://www.fidelity.com/learning-center/trading-investing/technical-analysis/technical-indicator-guide/bollinger-bands):
# - Short term: 10 day moving average, bands at 1.5 standard deviations.
# - Medium term: 20 day moving average, bands at 2 standard deviations.
# - Long term: 50 day moving average, bands at 2.5 standard deviations.
# +
pton_bbands, q = ti.get_bbands(symbol = 'pton', interval = 'daily', time_period = 20,
nbdevup = 2, nbdevdn = 2, matype = 0)
print(pton_bbands.tail(2))
pton_p, pton_m = ts.get_daily(symbol = "pton")
print(pton_p.tail(2))
pton_bbands.plot()
pton_p['4. close'].plot()
plt.title('Bollinger Bands for PTON')
plt.show()
# -
# ### Stochastic Oscillator
#
# A [stochastic oscillator](https://www.investopedia.com/terms/s/stochasticoscillator.asp) is a momentum indicator comparing a particular closing price of a security to a range of its prices over a certain period of time. The sensitivity of the oscillator to market movements is reducible by adjusting that time period or by taking a moving average of the result. It is used to generate overbought and oversold trading signals, utilizing a 0-100 bounded range of values.
#
# Readings over 80 are considered in the overbought range, and readings under 20 are considered oversold. However, these are not always a rule of thumb to follow; very strong trends can maintain overbought or oversold conditions for an extended period. It still takes human judgment to make an accurate deduction.
#
# The Stochastic Oscillator is displayed as two lines. The main line is called **"K"**. The second line, called **"D"**, is a moving average of **K**. There are several ways to interpret a Stochastic Oscillator. Three popular methods include:
#
# - Buy when the Oscillator (either K or D) falls below a specific level (e.g., 20) and then rises above that level.
# - Sell when the Oscillator rises above a specific level (e.g., 80) and then falls below that level.
# - Buy when the K line rises above the D line and sell when the K line falls below the D line.
#
# +
pton_stoch, q = ti.get_stoch(symbol = 'pton', interval = 'daily', fastkperiod = 12, slowkperiod = 26, slowdperiod = 12,
slowkmatype = 1, slowdmatype = 1)
print(pton_stoch.tail(2))
# Only fast
pton_stochf, q = ti.get_stochf(symbol = 'pton', interval = 'daily', fastkperiod = 12, fastdperiod = 12,
fastdmatype = 1)
print(pton_stochf.tail(2))
# StochRSI
pton_stochrsi, q = ti.get_stochrsi(symbol = 'pton', interval ='daily', time_period = 14,
fastkperiod = 12, fastdperiod = 12, fastdmatype = 0)
print(pton_stochrsi.tail(2))
pton_p, pton_m = ts.get_daily(symbol = "pton")
print(pton_p.tail(2))
pton_stoch.plot()
plt.title('Stoch for PTON')
plt.show()
pton_stochf.plot()
plt.title('Stoch (fast) for PTON')
plt.show()
pton_stochrsi.plot()
plt.title('Stoch RSI for PTON')
plt.show()
pton_p['4. close'].plot()
plt.title('Price chart for PTON')
plt.show()
# -
# ### Relative Strength Index (RSI)
# The relative strength index (RSI) is a momentum indicator used in technical analysis that measures the magnitude of recent price changes to evaluate overbought or oversold conditions in the price of a stock or other asset. The RSI is displayed as an oscillator (a line graph that moves between two extremes) and can have a reading from 0 to 100. The RSI will rise as the number and size of positive closes increase, and it will fall as the number and size of losses increase.
#
# Rule of thumb for the RSI are values of 70 or above indicate overbought or overvalued and may be primed for a trend reversal. An RSI reading of 30 or below indicates an oversold or undervalued condition. The standard is to use 14 periods to calculate the initial RSI value.
#
# ### RSI vs Stochastic Oscillator
# Relative strength index was designed to measure the speed of price movements, the stochastic oscillator formula works best when the market is trading in consistent ranges. Generally speaking, RSI is more useful in trending markets, and stochastics are more useful in sideways or choppy markets.
# +
pton_rsi, q = ti.get_rsi(symbol = 'pton', interval = 'daily', time_period = 14)
print(pton_rsi.tail(2))
pton_p, pton_m = ts.get_daily(symbol = "pton")
print(pton_p.tail(2))
pton_rsi.plot()
plt.title('RSI for PTON')
plt.show()
pton_p['4. close'].plot()
plt.title('Price chart for PTON')
plt.show()
# -
# ### Average Direction Movement Index (ADX)
# ADX stands for Average Directional Movement Index and can be used to help measure the overall strength of a trend.
#
# [How this indicator works](https://www.fidelity.com/learning-center/trading-investing/technical-analysis/technical-indicator-guide/adx)
# - Wilder suggests that a strong trend is present when ADX is above 25 and no trend is present when below 20.
# - When the ADX turns down from high values, then the trend may be ending. You may want to do additional research to determine if closing open positions is appropriate for you.
# - If the ADX is declining, it could be an indication that the market is becoming less directional, and the current trend is weakening. You may want to avoid trading trend systems as the trend changes.
# - If after staying low for a lengthy time, the ADX rises by 4 or 5 units, (for example, from 15 to 20), it may be giving a signal to trade the current trend.
# - If the ADX is rising then the market is showing a strengthening trend. The value of the ADX is proportional to the slope of the trend. The slope of the ADX line is proportional to the acceleration of the price movement (changing trend slope). If the trend is a constant slope then the ADX value tends to flatten out.
#
# ### Average Direction Movement Index Rating (ADXR)
# ADXR quantifies the change in momentum of the Average Directional Index (ADX).
#
# ADXR = (ADX + ADX n-periods ago) / 2
#
# The ADXR is used in the same way as the ADX – the higher the ADXR reading, the stronger the trend. As a rule of thumb, trend-following strategies are used when the ADXR shows a reading higher than 25. ADXR values above 40 are indicative of very strong trending environment, while values below 20 suggest that the market is in a trading range.
#
# ADXR is a lagging indicator (slow) and will usually provide signals after the ADX (fast) does. Technical analysts can use it like the MA methods and refer ADXR in the form of a signal line applied to the ADX. When ADX goes above ADXR, buy. When it goes lower, sell.
#
#
# +
pton_adx, q = ti.get_adx(symbol = 'pton', interval = 'daily', time_period = 20)
print(pton_adx.tail(2))
pton_adxr, q = ti.get_adxr(symbol = 'pton', interval = 'daily', time_period = 20)
print(pton_adxr.tail(2))
pton_p, pton_m = ts.get_daily(symbol = "pton")
print(pton_p.tail(2))
pton_adx.plot()
pton_adxr.plot()
plt.title('ADX, ADXR for PTON')
plt.show()
# why can't I stack the plots?
pton_p['4. close'].plot()
plt.title('Price chart for PTON')
plt.show()
# -
# ### Momentum Values
# Momentum measures the rate of the rise or fall in stock prices. For trending analysis, momentum is a useful indicator of strength or weakness in the issue's price. The formula for momentum is:
#
# Momentum = V−Vx
# where: V = Latest price
# Vx = Closing price X number of days ago
#
# Technicians typically use a 10-day time frame when measuring momentum. If the most recent closing price of the index is more than the closing price 10 trading days ago, the positive number (from the equation) is plotted above the zero line. Conversely, if the latest closing price is lower than the closing price 10 days ago, the negative measurement is plotted below the zero line.
#
# When the momentum indicator slides below the zero line and then reverses in an upward direction, it doesn't necessarily mean that the downtrend is over. It merely means that the downtrend is slowing down.
#
# +
pton_mom, q = ti.get_mom(symbol = 'pton', interval = 'daily', time_period = 10)
print(pton_mom.tail(2))
pton_p, pton_m = ts.get_daily(symbol = "pton")
print(pton_p.tail(2))
pton_mom.plot()
plt.title('Momentum value for PTON')
plt.show()
pton_p['4. close'].plot()
plt.title('Price chart for PTON')
plt.show()
# -
# ### Balance of Power Indicator
# The Balance of Power indicator measures the market strength of buyers against sellers by assessing the ability of each side to drive prices to an extreme level. IE: who are the players in control of the prices (sellers or buyers). The calculation is:
#
# Balance of Power = (Close price – Open price) / (High price – Low price)
#
# Traders may use this indicator to help:
# - Identify the direction of a trend.
# - Find divergences between the price and the BoP in order to identify a potential trend reversal or trend continuation setup.
# - Take advantage of overbought and oversold conditions.
# +
pton_bop, q = ti.get_bop(symbol = 'pton', interval = 'daily', time_period = 10)
print(pton_bop.tail(2))
pton_p, pton_m = ts.get_daily(symbol = "pton")
print(pton_p.tail(2))
pton_bop.plot()
plt.title('Balance of Power for PTON')
plt.show()
pton_p['4. close'].plot()
plt.title('Price chart for PTON')
plt.show()
# -
# ### Commodity Channel Index
# The CCI measures the difference between the current price and the historical average price. When the CCI is above zero it indicates the price is above the historic average. When CCI is below zero, the price is below the historic average.
#
# Unlike RSI or most momentum indicators ranging between 0 to 100, CCI is an unbounded indicator meaning it can go higher or lower indefinitely. For this reason, overbought and oversold levels are typically determined for each individual asset by looking at historical extreme CCI levels where the price reversed from. Example: Going from negative or near-zero readings to +100 can be used as a signal to watch for an emerging uptrend.
#
# The time period of 20 is most commonly used. Fewer periods results in a more volatile indicator, while more periods will make it smoother.
# +
pton_cci, q = ti.get_cci(symbol = 'pton', interval = 'daily', time_period = 20)
print(pton_cci.tail(2))
pton_p, pton_m = ts.get_daily(symbol = "pton")
print(pton_p.tail(2))
pton_cci.plot()
plt.title('CCI for PTON')
plt.show()
pton_p['4. close'].plot()
plt.title('Price chart for PTON')
plt.show()
# -
# ### Rate of Change
# The rate of change, commonly referred to as delta, is the speed at which a variable changes over a specific period of time. The formula for ROC is:
#
# Price ROC = (B − A)/A *100
#
# where
#
# B = price at current time
# A = price at previous time periods ago (20, if time_period = 20)
#
# Rate of change is also a [good indicator](https://www.investopedia.com/terms/r/rateofchange.asp) of market bubbles. Even though momentum is good and traders look for securities with a positive ROC, if a broad-market ETF, index, or mutual fund has a sharp increase in its ROC in the short term, it may be a sign that the market is unsustainable. If the ROC of an index or other broad-market security is over 50%, investors should be wary of a bubble.
# +
pton_roc, q = ti.get_roc(symbol = 'pton', interval = 'daily', time_period = 20)
print(pton_roc.tail(2))
pton_rocr, q = ti.get_rocr(symbol = 'pton', interval = 'daily', time_period = 20)
print(pton_rocr.tail(2))
pton_p, pton_m = ts.get_daily(symbol = "pton")
print(pton_p.tail(2))
pton_roc.plot()
plt.title('ROC for PTON')
plt.show()
pton_rocr.plot()
plt.title('ROC ratio for PTON')
plt.show()
pton_p['4. close'].plot()
plt.title('Price chart for PTON')
plt.show()
# -
# ## Closing
# There are a lot of technical indicators present within the financial industry. AlphaVantage has most of the essential indicators within their package, and this notebook has covered most of the important ones that are available within the package.
#
# The charts here are pretty bad and certainly require lots of help to improve their presentability. Do not be complacent with my work and try to come up with interesting combinations.
| Exploring Technical Analysis with AV.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Interruptible optimization runs with checkpoints
#
# <NAME>, Mai 2018
import numpy as np
np.random.seed(777)
# ## Problem statement
#
# Optimization runs can take a very long time and even run for multiple days. If for some reason the process has to be interrupted results are irreversibly lost, and the routine has to start over from the beginning.
#
# With the help of the `CheckpointSaver` callback the optimizer's current state can be saved after each iteration, allowing to restart from that point at any time.
#
# This is useful, for example,
#
# * if you don't know how long the process will take and cannot hog computational resources forever
# * if there might be system failures due to shaky infrastructure (or colleagues...)
# * if you want to adjust some parameters and continue with the already obtained results
# ## Simple example
#
# We will use pretty much the same optimization problem as in the [`bayesian-optimization.ipynb`](https://github.com/scikit-optimize/scikit-optimize/blob/master/examples/bayesian-optimization.ipynb) notebook. Additionaly we will instantiate the `CheckpointSaver` and pass it to the minimizer:
# +
from skopt import gp_minimize
from skopt import callbacks
from skopt.callbacks import CheckpointSaver
noise_level = 0.1
def obj_fun(x, noise_level=noise_level):
return np.sin(5 * x[0]) * (1 - np.tanh(x[0] ** 2)) + np.random.randn() * noise_level
checkpoint_saver = CheckpointSaver("./checkpoint.pkl", compress=9) # keyword arguments will be passed to `skopt.dump`
gp_minimize(obj_fun, # the function to minimize
[(-20.0, 20.0)], # the bounds on each dimension of x
x0=[-20.], # the starting point
acq_func="LCB", # the acquisition function (optional)
n_calls=10, # the number of evaluations of f including at x0
n_random_starts=0, # the number of random initialization points
callback=[checkpoint_saver], # a list of callbacks including the checkpoint saver
random_state=777);
# -
# Now let's assume this did not finish at once but took some long time: you started this on Friday night, went out for the weekend and now, Monday morning, you're eager to see the results. However, instead of the notebook server you only see a blank page and your colleague Garry tells you that he had had an update scheduled for Sunday noon – who doesn't like updates?
#
# TL;DR: `gp_minimize` did not finish, and there is no `res` variable with the actual results!
# ## Restoring the last checkpoint
#
# Luckily we employed the `CheckpointSaver` and can now restore the latest result with `skopt.load` (see [store and load results](./store-and-load-results.ipynb) for more information on that)
# +
from skopt import load
res = load('./checkpoint.pkl')
res.fun
# -
# ## Continue the search
#
# The previous results can then be used to continue the optimization process:
# +
x0 = res.x_iters
y0 = res.func_vals
gp_minimize(obj_fun, # the function to minimize
[(-20.0, 20.0)], # the bounds on each dimension of x
x0=x0, # already examined values for x
y0=y0, # observed values for x0
acq_func="LCB", # the acquisition function (optional)
n_calls=10, # the number of evaluations of f including at x0
n_random_starts=0, # the number of random initialization points
callback=[checkpoint_saver],
random_state=777);
# -
# ## Possible problems
#
# * __changes in search space:__ You can use this technique to interrupt the search, tune the search space and continue the optimization. Note that the optimizers will complain if `x0` contains parameter values not covered by the dimension definitions, so in many cases shrinking the search space will not work without deleting the offending runs from `x0` and `y0`.
# * see [store and load results](./store-and-load-results.ipynb) for more information on how the results get saved and possible caveats
| examples/interruptible-optimization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # An introduction to the Trove API
# + [markdown] slideshow={"slide_type": "slide"}
# ## What's an API?
#
# An API is an Application Programming Interface. It's a set of predefined requests and responses that enables computer programs talk to each other.
#
# Web APIs are generally used to deliver data. While humans can easily interpret information on a web page, computers need more help. APIs provide data in a form that computers can understand and use (we call this *machine-readable* data).
#
# The [Trove API](http://help.nla.gov.au/trove/building-with-trove/api) works much like the Trove website. You make queries and you get back results. But instead of going through a nicely-designed web interface, requests to the API are just URLs, and the results are just structured data.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Cracking the code
#
# While you can just type an API request into the location box of your web browser, most of the time requests and responses will be handled by a computer script or program. APIs don't care what programming language you use as long as you structure requests in the way they expect.
#
# In these notebooks we'll be using the programming language Python. No prior knowledge of Python is expected or required -- just follow along! The examples and approaches used could be easily translated into any another programming language.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Using this notebook
#
# This tutorial is created using Jupyter notebooks. Jupyter lets you combine text, images, and live code within a single web page. So not only can you read about collections data, you can download it, analyse it, and visualise it – all within your browser!
# + [markdown] slideshow={"slide_type": "slide"}
# ## Running code cells
#
# This notebook includes snippets of real code contained within cells. To run these snippets, just click on the cell and hit Shift+Enter. Try it with the cell below!
# +
# CLICK ON ME AND THEN HIT SHIFT+ENTER!
# This makes the datetime module available to use
import datetime
# This creates a variable called 'date_now' and uses the datetime.date.today() function to set it to today's date.
date_now = datetime.date.today()
# This displays a nicely-formatted string containing the date
print("Congratulations! On {} you ran the code in this cell.".format(date_now))
# + [markdown] slideshow={"slide_type": "skip"}
# You can also run the code in a cell by clicking on the 'Run' icon in the toolbar, or by hitting **Control+Enter**. You'll notice that **Shift+Enter** runs the code and moves you on to the next cell, while **Control+Enter** leaves you where you are.
# -
| Trove/Trove-API-Slideshow.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Task 1: Importing Libraries
import pandas as pd
import numpy as np
import seaborn as sns
from scipy.stats import skew
# %matplotlib inline
import matplotlib.pyplot as plt
plt.style.use("ggplot")
plt.rcParams['figure.figsize'] = (12, 8)
#
# ### Task 2: Load the Data
# The adverstiting dataset captures sales revenue generated with respect to advertisement spends across multiple channles like radio, tv and newspaper.
advert = pd.read_csv('Advertising.csv')
advert.head()
advert.info()
# ### Task 3: Relationship between Features and Response
sns.pairplot(advert, x_vars=['TV','radio','newspaper'], y_vars='sales', height=7, aspect=0.7);
#
# ### Task 4: Multiple Linear Regression - Estimating Coefficients
# +
from sklearn.linear_model import LinearRegression
# create X and y
feature_cols = ['TV', 'radio', 'newspaper']
X = advert[feature_cols]
y = advert.sales
# instantiate and fit
lm1 = LinearRegression()
lm1.fit(X, y)
# print the coefficients
print(lm1.intercept_)
print(lm1.coef_)
# -
# pair the feature names with the coefficients
list(zip(feature_cols, lm1.coef_))
sns.heatmap(advert.corr(), annot=True)
#
#
#
# ### Task 5: Feature Selection
#
# +
from sklearn.metrics import r2_score
lm2 = LinearRegression().fit(X[['TV', 'radio']], y)
lm2_preds = lm2.predict(X[['TV', 'radio']])
print("R^2: ", r2_score(y, lm2_preds))
# +
lm3 = LinearRegression().fit(X[['TV', 'radio', 'newspaper']], y)
lm3_preds = lm3.predict(X[['TV', 'radio', 'newspaper']])
print("R^2: ", r2_score(y, lm3_preds))
# -
#
# ### Task 6: Model Evaluation Using Train/Test Split and Metrics
#
# **Mean Absolute Error** (MAE) is the mean of the absolute value of the errors: <h5 align=center>$$\frac{1}{n}\sum_{i=1}^{n} \left |y_i - \hat{y_i} \right |$$</h5>
# **Mean Squared Error** (MSE) is the mean of the squared errors: <h5 align=center>$$\frac{1}{n}\sum_{i=1}^{n} (y_i - \hat{y_i})^2$$</h5>
# **Root Mean Squared Error** (RMSE) is the mean of the squared errors: <h5 align=center>$$\sqrt{\frac{1}{n}\sum_{i=1}^{n} (y_i - \hat{y_i})^2}$$</h5>
#
#
#
# Let's use train/test split with RMSE to see whether newspaper should be kept in the model:
# +
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_squared_error
X = advert[['TV', 'radio', 'newspaper']]
y = advert.sales
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state = 1)
lm4 = LinearRegression()
lm4.fit(X_train, y_train)
lm4_preds = lm4.predict(X_test)
print("RMSE :", np.sqrt(mean_squared_error(y_test, lm4_preds)))
print("R^2: ", r2_score(y_test, lm4_preds))
# +
X = advert[['TV', 'radio']]
y = advert.sales
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state = 1)
lm5 = LinearRegression()
lm5.fit(X_train, y_train)
lm5_preds = lm5.predict(X_test)
print("RMSE :", np.sqrt(mean_squared_error(y_test, lm5_preds)))
print("R^2: ", r2_score(y_test, lm5_preds))
# +
from yellowbrick.regressor import PredictionError, ResidualsPlot
visualizer = PredictionError(lm5)
visualizer.fit(X_train, y_train) # Fit the training data to the visualizer
visualizer.score(X_test, y_test) # Evaluate the model on the test data
visualizer.poof()
# -
visualizer = ResidualsPlot(lm5)
visualizer.fit(X_train, y_train)
visualizer.score(X_test, y_test)
visualizer.poof()
#
# ### Task 7: Interaction Effect (Synergy)
advert['interaction'] = advert['TV'] * advert['radio']
# +
X = advert[['TV', 'radio', 'interaction']]
y = advert.sales
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state = 1)
lm6 = LinearRegression()
lm6.fit(X_train, y_train)
lm6_preds = lm6.predict(X_test)
print("RMSE :", np.sqrt(mean_squared_error(y_test, lm6_preds)))
print("R^2: ", r2_score(y_test, lm6_preds))
# +
visualizer = PredictionError(lm6)
visualizer.fit(X_train, y_train) # Fit the training data to the visualizer
visualizer.score(X_test, y_test) # Evaluate the model on the test data
visualizer.poof()
| Linear Regression/Multiple Linear Regression/Predicting-sales-with-multiple-linear-regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="USuqdV4w8hDT"
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
from keras.datasets import mnist
from keras.layers import Dense, Flatten, Reshape
from keras.layers.advanced_activations import LeakyReLU
from keras.models import Sequential
from tensorflow.keras.optimizers import Adam
# + id="77Soc-9D8pzs"
img_rows = 28
img_cols = 28
channels = 1
img_shape = (img_rows, img_cols, channels) # input image dimensions
z_dim = 100 # size of the input noise vector to the generator
# + id="XONCjw_79DnE"
# Generator
def build_generator(img_shape, z_dim):
model = Sequential()
model.add(Dense(128, input_dim=z_dim))
model.add(LeakyReLU(alpha=0.1))
model.add(Dense(28 * 28 * 1, activation='tanh'))
model.add(Reshape(img_shape))
return model
# + id="OHOUBjxt-Q_W"
# Discriminator
def build_discriminator(img_shape):
model = Sequential()
model.add(Flatten(input_shape=img_shape))
model.add(Dense(128))
model.add(LeakyReLU(alpha=0.1))
model.add(Dense(1, activation='sigmoid'))
return model
# + id="n2Y-w9aYABhp"
# build the GAN model
def build_gan(generator, discriminator):
model = Sequential()
model.add(generator)
model.add(discriminator)
return model
# + id="1kbOxH3uBZyb"
discriminator = build_discriminator(img_shape)
discriminator.compile(loss='binary_crossentropy', optimizer=Adam(), metrics=['accuracy'])
generator = build_generator(img_shape, z_dim)
discriminator.trainable = False
gan = build_gan(generator, discriminator)
gan.compile(loss='binary_crossentropy', optimizer=Adam())
# + id="L_uuwqC0B3xN"
# GAN training loop
losses = []
accuracies = []
iteration_checkpoints = []
def train(iterations, batch_size, sample_interval):
(X_train, _), (_,_) = mnist.load_data()
X_train = X_train / 127.5 - 1 # rescale pixel values to [-1, 1] as tanh gives in range [-1,1]
X_train = np.expand_dims(X_train, axis=3) # add a channel dimension
real = np.ones((batch_size, 1)) # labels for real images
fake = np.zeros((batch_size, 1)) # labels for fake images
for iteration in range(iterations):
idx = np.random.randint(0, X_train.shape[0], batch_size) # gets a batch of real images
imgs = X_train[idx]
z = np.random.normal(0, 1, (batch_size, 100)) # generate a batch of fake images
gen_imgs = generator.predict(z)
d_loss_real = discriminator.train_on_batch(imgs, real) # train the discriminator
d_loss_fake = discriminator.train_on_batch(gen_imgs, fake)
d_loss, accuracy = 0.5 * np.add(d_loss_real, d_loss_fake)
z = np.random.normal(0, 1, (batch_size, 100)) # generate a batch of fake images
gen_imgs = generator.predict(z)
g_loss = gan.train_on_batch(z, real) # trian the generator
if (iteration + 1) % sample_interval == 0:
losses.append((d_loss, g_loss))
accuracies.append(100.0 * accuracy)
iteration_checkpoints.append(iteration + 1)
print("%d [D loss: %f, acc.: %.2f%%] [G loss: %f]" %
(iteration + 1, d_loss, 100.0 * accuracy, g_loss))
sample_images(generator)
# + id="Lk0VxJ_WKngo"
def sample_images(generator, image_grid_rows = 4, image_grid_columns=4):
z = np.random.normal(0, 1, (image_grid_rows*image_grid_columns, z_dim)) # sample random noise
gen_imgs = generator.predict(z)
gen_imgs = 0.5 * gen_imgs + 0.5 # rescale pixels to [0,1]
fig, axs = plt.subplots(image_grid_rows,
image_grid_columns,
figsize=(4,4),
sharey=True,
sharex=True)
cnt = 0
for i in range(image_grid_rows):
for j in range(image_grid_columns):
axs[i,j]. imshow(gen_imgs[cnt, :, :, 0], cmap='gray')
axs[i,j].axis('off')
cnt+=1
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="5BYdLnT-FV7i" outputId="5396b847-8d6f-4858-b301-d7c05ff9e1e4"
iterations = 10000
batch_size = 128
sample_interval = 1000
train(iterations, batch_size, sample_interval)
# + id="SToGLuY-Ffhw" colab={"base_uri": "https://localhost:8080/", "height": 389} outputId="73602ba4-358b-4359-e83a-70d4fd8e7929"
losses = np.array(losses)
# Plot training losses for Discriminator and Generator
plt.figure(figsize=(15, 5))
plt.plot(iteration_checkpoints, losses.T[0], label="Discriminator loss")
plt.plot(iteration_checkpoints, losses.T[1], label="Generator loss")
plt.xticks(iteration_checkpoints, rotation=90)
plt.title("Training Loss")
plt.xlabel("Iteration")
plt.ylabel("Loss")
plt.legend()
| GAN1_MNIST.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.7.1
# language: julia
# name: julia-1.7
# ---
# Before running this, please make sure to activate and instantiate the
# environment with [this `Project.toml`](https://raw.githubusercontent.com/juliaai/DataScienceTutorials.jl/gh-pages/__generated/A-ensembles-3/Project.toml) and
# [this `Manifest.toml`](https://raw.githubusercontent.com/juliaai/DataScienceTutorials.jl/gh-pages/__generated/A-ensembles-3/Manifest.toml).
# For instance, copy these files to a folder 'A-ensembles-3', `cd` to it and
#
# ```julia
# using Pkg; Pkg.activate("."); Pkg.instantiate()
# ```
# # Simple example of a homogeneous ensemble using learning networks
# In this simple example, no bagging is used, so every atomic model
# gets the same learned parameters, unless the atomic model training
# algorithm has randomness, eg, DecisionTree with random subsampling
# of features at nodes.
# Note that MLJ has a built in model wrapper called `EnsembleModel`
# for creating bagged ensembles with a few lines of code.
# ## Definition of composite model type
using MLJ
using PyPlot
import Statistics
# Defining the learning network (composite model spec):
# +
Xs = source()
ys = source()
DecisionTreeRegressor = @load DecisionTreeRegressor pkg=DecisionTree
atom = DecisionTreeRegressor()
machines = (machine(atom, Xs, ys) for i in 1:100)
# -
# Overloading `mean` for nodes:
# +
Statistics.mean(v...) = mean(v)
Statistics.mean(v::AbstractVector{<:AbstractNode}) = node(mean, v...)
yhat = mean([predict(m, Xs) for m in machines]);
# -
# Defining the new composite model type and instance:
# +
surrogate = Deterministic()
mach = machine(surrogate, Xs, ys; predict=yhat)
@from_network mach begin
mutable struct OneHundredModels
atom=atom
end
end
one_hundred_models = OneHundredModels()
# -
# ## Application to data
X, y = @load_boston;
# tune regularization parameter for a *single* tree:
# +
r = range(atom,
:min_samples_split,
lower=2,
upper=100, scale=:log)
mach = machine(atom, X, y)
figure()
curve = learning_curve!(mach,
range=r,
measure=mav,
resampling=CV(nfolds=9),
verbosity=0)
plot(curve.parameter_values, curve.measurements)
xlabel(curve.parameter_name)
# -
# \fig{e1.svg}
# tune regularization parameter for all trees in ensemble simultaneously:
# +
r = range(one_hundred_models,
:(atom.min_samples_split),
lower=2,
upper=100, scale=:log)
mach = machine(one_hundred_models, X, y)
figure()
curve = learning_curve!(mach,
range=r,
measure=mav,
resampling=CV(nfolds=9),
verbosity=0)
plot(curve.parameter_values, curve.measurements)
xlabel(curve.parameter_name)
# -
# \fig{e2}
# ---
#
# *This notebook was generated using [Literate.jl](https://github.com/fredrikekre/Literate.jl).*
| __site/__generated/A-ensembles-3/tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="lmTQJGoCdYDW" colab={"base_uri": "https://localhost:8080/"} outputId="d208cc97-ab09-4817-fe6a-b108af00f0df"
# !pip install transformers
# !pip install sentencepiece
# + id="QAhlLwQ4jZja"
# This class uses a long text as input and summarizes it
from transformers import T5ForConditionalGeneration, T5Tokenizer
class ComplexString(str):
__model_name = 't5-base'
def summary(self) -> str:
text = 'summarize: %s' % self
model = T5ForConditionalGeneration.from_pretrained(self.__model_name)
tokenizer = T5Tokenizer.from_pretrained(self.__model_name)
features = tokenizer(text, return_tensors='pt')
input_ids = features['input_ids']
attention_mask = features['attention_mask']
output = model.generate(input_ids = input_ids, attention_mask = attention_mask, max_length=512)
if len(output) > 0:
return tokenizer.decode(output[0]).replace('<pad>', '').replace('</s>', '').strip()
else:
return None
# + colab={"base_uri": "https://localhost:8080/", "height": 54} id="pNLSAl2vlZzO" outputId="f0d12a78-05a1-42ab-b887-7f1d4acf570c"
text = """
The National Union of Freedom Fighters (NUFF) was an armed Marxist revolutionary group in Trinidad and Tobago. The group fought a guerrilla campaign to overthrow the government of Prime Minister <NAME> following the failed 1970 Black Power uprising and a mutiny in the Trinidad and Tobago Regiment. NUFF formed from the Western United Liberation Front, a loose grouping of largely unemployed men from the western suburbs of Port of Spain. NUFF drew disaffected members of the National Joint Action Committee, a Black Power organisation, and established a training camp in south Trinidad. In 1972 and 1973 NUFF attacked police posts to acquire weapons, robbed banks, and carried out an insurgent campaign against the government. With improved intelligence capabilities, the government eventually killed or captured most of its leadership. Eighteen NUFF members and three policemen were killed over the course of the insurgency. NUFF was anti-imperialist and anti-capitalist and was notable for the extent to which women played an active role in the organisation, including among its guerrilla fighters.
"""
c_string = ComplexString(text)
c_string.summary()
# + id="ZCc7eOAGlnmW"
# Prediction of answer given a context and question
context = 'the national union of freedom fighters was an armed revolutionary group. it fought a guerrilla campaign to overthrow the government of prime minister <NAME>. NUFF attacked police posts to acquire weapons, robbed banks, carried out insurgent campaign.'
question = "Who is the prime minister"
input = 'question: %s context: %s' % (question, context)
# + colab={"base_uri": "https://localhost:8080/", "height": 37} id="-KTd84lyqwUb" outputId="75a24541-1c17-4b70-c7e7-29e0bfd82c90"
features = tokenizer(input, return_tensors='pt')
input_ids = features['input_ids']
attention_mask = features['attention_mask']
outcome = model.generate(input_ids = input_ids, attention_mask = attention_mask, max_length=255)
tokenizer.decode(outcome[0])
# + id="6NzHG1DEqxBq"
| generators/text_summarization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="mwbb5IvNfqi7"
# ### Linguistic Features
# + id="sutS0dAj79S6"
# !python -m spacy download en_core_web_md
# + [markdown] id="3xY1K-Kv9J8B"
# ### POS Tagging
# + colab={"base_uri": "https://localhost:8080/"} id="NQfxZmSi7smo" outputId="364c0db4-b517-4327-a3a3-e704d8a557aa"
import spacy
nlp = spacy.load('en_core_web_md')
doc = nlp("Alicia and me went to the school by bus")
for token in doc:
print(token.text, token.pos_, token.tag_, spacy.explain(token.pos_), spacy.explain(token.tag_))
# + colab={"base_uri": "https://localhost:8080/"} id="wcgzhVGs8gnP" outputId="a68b43ef-e336-40e4-adb1-300dd2988356"
doc = nlp("My friend will fly to New York fast and she is staying there for 3 days.")
for token in doc:
print(token.text, token.pos_, token.tag_, spacy.explain(token.pos_), spacy.explain(token.tag_))
# + colab={"base_uri": "https://localhost:8080/"} id="YDv-ino7BIyY" outputId="55d3d014-90c4-42fc-8cd4-dad4acea7f4d"
doc = nlp("My cat will fish for a fish tomorrow in a fishy way.")
for token in doc:
print(token.text, token.pos_, token.tag_, spacy.explain(token.pos_), spacy.explain(token.tag_))
# + colab={"base_uri": "https://localhost:8080/"} id="fTRX6GxVBpJ4" outputId="ac7a3314-3737-4b00-b6e1-95c5dbea8387"
doc = nlp("He earned $5.5 million in 2020 and paid %35 tax.")
for token in doc:
print(token.text, token.pos_, token.tag_, spacy.explain(token.pos_), spacy.explain(token.tag_))
# + [markdown] id="UZe8X3joIlqH"
# ### Dependency
# + colab={"base_uri": "https://localhost:8080/"} id="2UNaLH9QBwpQ" outputId="f5aaed13-8f16-4ad2-eb87-194cc23312da"
doc = nlp("I counted white sheep.")
for token in doc:
print(token.text, token.pos_, token.tag_, token.dep_, token.head)
# + colab={"base_uri": "https://localhost:8080/", "height": 333} id="Fao21VZuD6N_" outputId="633503dd-fb93-41bc-d643-7bbff0977336"
from spacy import displacy
displacy.render(doc, jupyter=True, style='dep')
# + [markdown] id="D4ouMGaYMFNI"
# ### NER
# + colab={"base_uri": "https://localhost:8080/"} id="BvlUQk3DHE_I" outputId="2db11533-4f02-4d8a-90ef-2512fea37e0b"
doc = nlp("The president <NAME> visited France.")
print(doc.ents)
print(type(doc.ents[1]))
# + colab={"base_uri": "https://localhost:8080/"} id="fHLqgu5oHqY4" outputId="3ab4161b-376c-4f9e-ca47-ca525a240cf3"
print(spacy.explain("ORG"))
# + colab={"base_uri": "https://localhost:8080/"} id="NdkF5cHPHweQ" outputId="7649724c-c318-4135-f7d4-8ff6f9dee51a"
doc2 = nlp("He worked for NASA")
token = doc2[3]
print(token.text, token.ent_type_, spacy.explain(token.ent_type_))
# + colab={"base_uri": "https://localhost:8080/"} id="hTTeY5jyH1Fn" outputId="b3a07fba-e175-49fe-f750-dcbe80e8c3ae"
doc3 = nlp("“<NAME> was born in Ulm on 1987. He studied electronical engineering at ETH Zurich.")
print(doc3.ents)
# + colab={"base_uri": "https://localhost:8080/"} id="BvVWJbU4ICf_" outputId="2af1bc9b-adf9-42f1-f3ce-02f64359c259"
for token in doc3:
print(token.text, token.ent_type_, spacy.explain(token.ent_type_))
# + [markdown] id="iLT3J4BxPeUP"
# ### Merging-Splitting-Merge
# + colab={"base_uri": "https://localhost:8080/"} id="glXWsaHPIPGn" outputId="090c7e9b-64f8-45ef-a14c-352d56f607c4"
doc = nlp("She lived in New Hampshire.")
print(doc.ents)
print([(token.text, token.i) for token in doc])
print(len(doc))
# + id="Bod5QVkwOmz2"
with doc.retokenize() as retokenizer:
retokenizer.merge(doc[3:5], attrs={"LEMMA":"new hampshire"})
# + colab={"base_uri": "https://localhost:8080/"} id="S974ZYSkPGjv" outputId="ab66646b-4116-44c1-e992-fd1c67542b97"
print(doc.ents)
print([(token.text, token.i) for token in doc])
# + colab={"base_uri": "https://localhost:8080/"} id="g0L2tZaNPK8X" outputId="dd6feed4-e966-4a8f-a7d9-68b92c0f139f"
print(len(doc))
print([(token.lemma_) for token in doc])
# + colab={"base_uri": "https://localhost:8080/"} id="OLptmTy-PXnH" outputId="d6eea4f6-2382-4eeb-cd42-4982799d830e"
doc = nlp("She lived in NewHampshire.")
print(len(doc))
print([(token.text, token.lemma_, token.i) for token in doc])
for token in doc:
print(token.text, token.pos_, token.tag_, token.dep_)
# + id="YxfxUUdpQkDO"
with doc.retokenize() as retokenizer:
heads = [(doc[3], 1), doc[2]]
attrs = {"TAG":["NNP", "NNP"], "DEP":["compound", "pobj"]}
retokenizer.split(doc[3], ["New", "Hampshire"], heads=heads, attrs=attrs)
# + colab={"base_uri": "https://localhost:8080/"} id="X3y6-Fx-Qt7X" outputId="69592ab8-1e42-4b9a-cb5d-be585914405a"
print(len(doc))
print([(token.text, token.lemma_, token.i) for token in doc])
for token in doc:
print(token.text, token.pos_, token.tag_, token.dep_)
# + [markdown] id="avZcuWNifUVL"
# ### Rule-Based Matching - Matcher Class
#
# Matching a pattern
# + id="QHhKl-x8QxOe"
from spacy.matcher import Matcher
# + colab={"base_uri": "https://localhost:8080/"} id="y8ppvYMEjBXa" outputId="a64f83a1-2526-4ef3-945a-b31251695cfd"
doc = nlp("Good morning, I want to reserve a ticket.")
matcher = Matcher(nlp.vocab) # Matcher needs to be intialized with vocabulary object
pattern = [{"LOWER": "good"}, {"LOWER": "morning"}, {"IS_PUNCT": True}]
matcher.add("morningGreeting", [pattern])
matches = matcher(doc)
for match_id, start, end in matches:
m_span = doc[start:end]
print(start, end, m_span.text)
# + [markdown] id="DL6kZu_M5Rie"
# Matching two patterns
# + colab={"base_uri": "https://localhost:8080/"} id="PjE4HC4slcIz" outputId="a6662844-274f-4d82-e410-77a2c47b92cb"
doc = nlp("Good morning, I want to reserve a ticket. I will then say good evening!")
# Initialize the Matcher
matcher = Matcher(nlp.vocab)
# Define the patterns
pattern1 = [{"LOWER": "good"}, {"LOWER": "morning"}, {"IS_PUNCT": True}]
pattern2 = [{"LOWER": "good"}, {"LOWER": "evening"}, {"IS_PUNCT": True}]
# name and add the patterns
matcher.add("morningGreeting", [pattern1])
matcher.add("eveningGreeting", [pattern2])
# get the matches
matches = matcher(doc)
for match_id, start, end in matches:
pattern_name = nlp.vocab.strings[match_id]
m_span = doc[start:end]
print(start, end, m_span.text)
# + [markdown] id="_LAoqbw-9pX9"
# While matching pattern ORTH and TEXT are similar to LOWER: they mean an exact match of the token text, including the case.
# + colab={"base_uri": "https://localhost:8080/"} id="EFXfEoJn2bH2" outputId="69f12d91-5d2b-4522-93f5-786f79b5e7ec"
doc = nlp("I bought a pineapple.")
matcher = Matcher(nlp.vocab)
pattern = [{"LENGTH": 1}]
matcher.add("onlyShort", [pattern])
matches = matcher(doc)
print("no.of matches:", len(matches))
for mid, start, end in matches:
print(start, end, doc[start:end])
# + [markdown] id="PaHpPu52-PXc"
# The next block of token attributes is IS_ALPHA, IS_ASCII, and IS_DIGIT. These features are handy for finding number tokens and ordinary words (which do not include any interesting characters). The following pattern matches a sequence of two tokens, a number followed by an ordinary word:
# + colab={"base_uri": "https://localhost:8080/"} id="2380v65K9_vr" outputId="5ad6cfd3-e489-4b31-9447-145d638df981"
doc1 = nlp("I met him at 2 o'clock.")
matcher = Matcher(nlp.vocab)
pattern = [{"IS_DIGIT": True},{"IS_ALPHA": True}]
matcher.add("numberAndPlainWord", [pattern])
matches = matcher(doc1)
print(len(matches))
for mid, start, end in matches:
print(start, end, doc1[start:end])
# + colab={"base_uri": "https://localhost:8080/"} id="xmLioJoC-yc1" outputId="80f64c1d-ab5c-431d-9a20-813d6c5598d2"
doc2 = nlp("He brought me 2 apples.")
matcher = Matcher(nlp.vocab)
pattern = [{"IS_DIGIT": True},{"IS_ALPHA": True}]
matcher.add("numberAndPlainWord", [pattern])
matches = matcher(doc2)
print(len(matches))
for mid, start, end in matches:
print(start, end, doc2[start:end])
# + [markdown] id="vs257RcVOH7M"
# In the preceding code segment, 2 o'clock didn't match the pattern because o'clock contains an apostrophe, which is not an alphabetic character (alphabetic characters are digits, letters, and the underscore character). 2 apples matched because the token apples consists of letters.
# + colab={"base_uri": "https://localhost:8080/"} id="qaM7q81n_O3s" outputId="eab1c1f3-17f8-46ae-840b-3e950556b821"
doc = nlp("Take me out of your SPAM list. We never asked you to contact me. If you write again we'll SUE!!!!")
matcher = Matcher(nlp.vocab)
pattern = [{"IS_UPPER": True}]
matcher.add("capitals", [pattern])
matches = matcher(doc)
for mid, start, end in matches:
print(start, end, doc[start:end])
# + id="yrxVBGibQ4Jy"
doc1 = nlp("Can you swim?")
matcher = Matcher(nlp.vocab)
# In here we put two attributes in one brace
pattern = [{"IS_SENT_START": True, "LOWER": "can"}, {"IS_TITLE": True}]
matcher.add("canThenCapitalized", [pattern])
matches = matcher(doc1)
for mid, start, end in matches:
print(start, end, doc1[start:end])
# + colab={"base_uri": "https://localhost:8080/"} id="AWMq_HKoRBZ7" outputId="4070a8f8-a7f0-4c29-d28c-2bcbc51d9639"
doc2 = nlp("Can Sally swim?")
matcher = Matcher(nlp.vocab)
pattern = [{"IS_SENT_START": True, "LOWER": "can"}, {"IS_TITLE": True}]
matcher.add("canThenCapitalized", [pattern])
matches = matcher(doc2)
for mid, start, end in matches:
print(start, end, doc2[start:end])
# + [markdown] id="DUj7P6RHSkok"
# LIKE_NUM, LIKE_URL, and LIKE_EMAIL are attributes that are related to token shape.
#
# After seeing the shape attributes, let's see the POS, TAG, DEP, LEMMA, and SHAPE linguistic attributes
# + colab={"base_uri": "https://localhost:8080/"} id="b2Rz5BiPRZf7" outputId="7fdbd4b0-c186-4184-d711-91dc4e383ea8"
doc = nlp("Will you go there?")
matcher = Matcher(nlp.vocab)
pattern = [{"IS_SENT_START": True, "TAG": "MD"}]
matcher.add("sentStart",[pattern])
matches = matcher(doc)
for mid, start, end in matches:
print(start, end, doc[start:end])
# + [markdown] id="nI-2phepVcld"
# Extended syntax support - IN, NOT_IN, IS_SUBSET, IS_SUPERSET, INTESECTS and comparison operators.
# + colab={"base_uri": "https://localhost:8080/"} id="lUbhjgX0TNvL" outputId="01ad9aff-1cdf-461d-aa92-350c4200b4e2"
doc = nlp("Good morning, I'm here. I'll say good evening!!")
matcher = Matcher(nlp.vocab)
pattern = [{"LOWER": "good"},{"LOWER": {"IN": ["morning", "evening"]}},{"IS_PUNCT": True}]
matcher.add("greetings", [pattern])
matches = matcher(doc)
for mid, start, end in matches:
print(start, end, doc[start:end])
# + colab={"base_uri": "https://localhost:8080/"} id="y7EtCuPGVhs7" outputId="0876cd62-4b9d-46c4-9bcc-c6859c150bf6"
doc = nlp("I suffered from Trichotillomania when I was in college. The doctor prescribed me Psychosomatic medicine.")
matcher = Matcher(nlp.vocab)
pattern = [{"LENGTH": {">=" : 10}}]
matcher.add("longWords", [pattern])
matches = matcher(doc)
for mid, start, end in matches:
print(start, end, doc[start:end])
# + [markdown] id="iDbbEWskWLLy"
# Regex-like operators - OP
# ```
# # ! Negate the pattern, by requiring it to match exactly 0 times.
# # ? Make the pattern optional, by allowing it to match 0 or 1 times.
# + Require the pattern to match 1 or more times.
# * Allow the pattern to match 0 or more times.
# ```
# + colab={"base_uri": "https://localhost:8080/"} id="HR6d1Lu6WOKC" outputId="237148f5-37e1-4a6f-e90d-8d61a0fc8640"
doc1 = nlp("Barack Obama visited France.")
doc2 = nlp("<NAME> Obama visited France.")
matcher = Matcher(nlp.vocab)
pattern = [{"LOWER": "barack"}, {"LOWER": "hussein", "OP": "?"},{"LOWER": "obama"}]
matcher.add("obamaNames", [pattern])
print(matcher(doc1))
print(matcher(doc2))
# + colab={"base_uri": "https://localhost:8080/"} id="W5TZVOkwXt2R" outputId="8e01dc62-f31a-46ad-be93-633dea3f4f42"
doc1 = nlp("Hello hello hello, how are you?")
doc2 = nlp("Hello, how are you?")
doc3 = nlp("How are you?")
matcher = Matcher(nlp.vocab)
pattern = [{"LOWER": {"IN": ["hello", "hi", "hallo"]}, "OP": "*"}, {"IS_PUNCT": True}]
matcher.add("greetings", [pattern])
print("**************")
for mid, start, end in matcher(doc1):
print(start, end, doc1[start:end])
print("**************")
for mid, start, end in matcher(doc2):
print(start, end, doc1[start:end])
print("**************")
for mid, start, end in matcher(doc3):
print(start, end, doc1[start:end])
print("**************")
# + colab={"base_uri": "https://localhost:8080/"} id="11u5Q6QabgQ0" outputId="a56640e0-825a-4141-f730-639e83cb863e"
doc1 = nlp("Hello hello hello, how are you?")
doc2 = nlp("Hello, how are you?")
doc3 = nlp("How are you?")
matcher = Matcher(nlp.vocab)
pattern = [{"LOWER": {"IN": ["hello", "hi", "hallo"]}, "OP": "+"}, {"IS_PUNCT": True}]
matcher.add("greetings", [pattern])
print("**************")
for mid, start, end in matcher(doc1):
print(start, end, doc1[start:end])
print("**************")
for mid, start, end in matcher(doc2):
print(start, end, doc1[start:end])
print("**************")
for mid, start, end in matcher(doc3):
print(start, end, doc1[start:end])
print("**************")
# + [markdown] id="XI-1fH06YjEa"
# Regex support - spaCy Matcher offers full support for token-level regex matching
# + colab={"base_uri": "https://localhost:8080/"} id="qhzDrjbEYfhz" outputId="d8436bf4-f730-4a80-a597-9dfd44e3d3c0"
doc1 = nlp("I travelled by bus.")
doc2 = nlp("She traveled by bike.")
matcher = Matcher(nlp.vocab)
pattern = [{"POS": "PRON"}, {"TEXT": {"REGEX": "[Tt]ravell?ed"}}]
matcher.add("travelRegex", [pattern])
for mid, start, end in matcher(doc1):
print(start, end, doc1[start:end])
for mid, start, end in matcher(doc2):
print(start, end, doc2[start:end])
# + colab={"base_uri": "https://localhost:8080/"} id="3uICLAFmffhJ" outputId="a4df2f03-d200-419d-aae5-54f298082538"
doc = nlp("I went to Italy; he has been there too. His mother also has told me she wants to visit Rome.")
matcher = Matcher(nlp.vocab)
# Using regex with POS tags
pattern = [{"TAG": {"REGEX": "^V"}}]
matcher.add("verbs", [pattern])
for mid, start, end in matcher(doc):
print(start, end, doc1[start:end])
# + [markdown] id="rNE2o8zQcVzs"
# We have extracted all the finite verbs (you can think of a finite verb as a non-modal verb). How did we do it? Our token pattern includes the regex ^V, which means all fine-grained POS tags that start with V: VB, VGD, VBG, VBN, VBP, and VBZ. Then we extracted tokens with verbal POS tags.
# + [markdown] id="B_mmJ2ALgFWi"
# Wild Card Matching
# + colab={"base_uri": "https://localhost:8080/"} id="lZFTHiXCYgg6" outputId="32a1d0f9-3a01-4480-9978-57326a946d9e"
doc = nlp("My name is Alice and his name was Elliot.")
matcher = Matcher(nlp.vocab)
pattern = [{"LOWER": "name"},{"LEMMA": "be"},{}]
matcher.add("pickName", [pattern])
for mid, start, end in matcher(doc):
print(start, end, doc[start:end])
# + colab={"base_uri": "https://localhost:8080/"} id="NoUbQfM9ciYq" outputId="c99f9cfb-2cda-4e8b-ec86-ae80d32027a9"
doc1 = nlp("I forwarded his email to you.")
doc2 = nlp("I forwarded an email to you.")
doc3 = nlp("I forwarded the email to you.")
matcher = Matcher(nlp.vocab)
pattern = [{"LEMMA": "forward"}, {}, {"LOWER": "email"}]
matcher.add("forwardMail", [pattern])
print("****************************")
for mid, start, end in matcher(doc1):
print(start, end, doc1[start:end])
print("****************************")
for mid, start, end in matcher(doc2):
print(start, end, doc2[start:end])
print("****************************")
for mid, start, end in matcher(doc3):
print(start, end, doc3[start:end])
print("****************************")
# + [markdown] id="xDsNdDlt4fiT"
# To check regex and Matcher these sites are useful:
#
# https://regex101.com/
#
# https://explosion.ai/demos/matcher
# + [markdown] id="h9qqYwPv4y5A"
# Phrase Matcher
# + id="UB6EyoJOc8qp"
from spacy.matcher import PhraseMatcher
# + colab={"base_uri": "https://localhost:8080/"} id="LWOlfDpd43l6" outputId="edab1481-6820-471a-8ad4-99621abb4433"
doc = nlp("3 EU leaders met in Berlin. German chancellor <NAME> first welcomed the US president <NAME>. The following day <NAME> joined them in Brandenburg.")
matcher = PhraseMatcher(nlp.vocab)
terms = ["<NAME>", "<NAME>", "<NAME>"]
patterns = [nlp.make_doc(term) for term in terms]
# make_doc() creates a Doc from every term, and it's quite efficient in terms
# of processing because instead of the whole pipeline, it only calls the Tokenizer
matcher.add("politiciansList", None, *patterns)
matches = matcher(doc)
for mid, start, end in matches:
print(start, end, doc[start:end])
# + [markdown] id="u43ksZ7395lI"
# Example of matching by the LOWER attribute
# + colab={"base_uri": "https://localhost:8080/"} id="73GcQzIY5N3f" outputId="c9d7921e-e350-46f0-b4cf-0ff9a337f9d1"
doc = nlp("During the last decade, derivatives market became an asset class of their own and influenced the financial landscape strongly.")
matcher = PhraseMatcher(nlp.vocab, attr="LOWER")
terms = ["Asset", "Investment", "Derivatives", "Demand", "Market"]
patterns = [nlp.make_doc(term) for term in terms]
matcher.add("financeTerms", None, *patterns)
matches = matcher(doc)
for mid, start, end in matches:
print(start, end, doc[start:end])
# + [markdown] id="RPpsRlrZ-Ds3"
# Example of matching by the SHAPE attribute
# + colab={"base_uri": "https://localhost:8080/"} id="J4gCpprl98bI" outputId="5e7844bf-7d64-4730-863a-935f7931dc67"
doc = nlp("This log contains the following IP addresses: 172.16.58.3 and 172.16.58.3 and 172.16.58.3 .")
matcher = PhraseMatcher(nlp.vocab, attr="SHAPE")
ip_nums = ["127.0.0.0", "127.256.0.0"]
patterns = [nlp.make_doc(ip) for ip in ip_nums]
matcher.add("IPNums", None, *patterns)
for mid, start, end in matcher(doc):
print(start, end, doc[start:end])
# + [markdown] id="Vb-vr2_o-lcy"
# ### Entity Ruler
# + colab={"base_uri": "https://localhost:8080/"} id="5XwDkZur-Z8n" outputId="c57be757-ca54-436d-b463-f2169bbd2f1f"
doc = nlp("<NAME> visited Berlin.")
matcher = Matcher(nlp.vocab)
pattern = [{"ENT_TYPE": "PERSON"}]
matcher.add("personEnt", [pattern])
matches = matcher(doc)
for mid, start, end in matches:
print(start, end, doc[start:end])
# + colab={"base_uri": "https://localhost:8080/"} id="viT_v9TC--J2" outputId="bdef605e-cc3d-4430-9261-3c60b6200a0f"
doc = nlp("Today German chancellor <NAME> met with the US president.")
matcher = Matcher(nlp.vocab)
pattern = [{"ENT_TYPE": "PERSON", "OP": "+"}, {"POS" : "VERB"}]
matcher.add("personEntAction", [pattern])
matches = matcher(doc)
for mid, start, end in matches:
print(start, end, doc[start:end])
# + [markdown] id="SI8Nkylb_cyw"
# spaCy's EntityRuler is the component that allows us to add rules on top of the statistical model and creates an even more powerful NER model.
#
# EntityRuler is not a matcher, it's a pipeline component that we can add to our pipeline via nlp.add_pipe. When it finds a match, the match is appended to doc.ents and ent_type will be the label we pass in the pattern
# + colab={"base_uri": "https://localhost:8080/"} id="HZF_i8pvpdqv" outputId="247c3edc-9b4f-4935-a424-dbd8cf685326"
nlp.pipe_names
# + colab={"base_uri": "https://localhost:8080/"} id="_39AmppI_IIA" outputId="bfa99b33-907a-4e3b-a6c2-c13974583cb5"
doc = nlp("I have an acccount with chime since 2017")
for ent in doc.ents:
print(ent.text, ent.label_)
# + id="c5dZAEmdPOQ2"
ruler = nlp.add_pipe('entity_ruler')
# + colab={"base_uri": "https://localhost:8080/"} id="FnZRBePhRGth" outputId="07d80f47-0e0e-4e18-92e9-08f04774b117"
nlp.pipe_names
# + id="QGb4Id1i1KCY"
patterns = [{"label": "ORG", "pattern": [{"LOWER": "chime"}]}]
# + id="fNIIC8372Tmh"
ruler.add_patterns(patterns)
# + colab={"base_uri": "https://localhost:8080/"} id="DnUG-5AJ_zNd" outputId="e5f2679b-f810-41a4-d0e7-1949c7741932"
doc2 = nlp("I have an acccount with chime since 2017")
print(doc2.ents)
print(doc2[5].ent_type_)
# + [markdown] id="_KOHJcDf9nby"
# ### Combining spaCy models and matchers
# Extracting IBAN and account numbers
# + colab={"base_uri": "https://localhost:8080/"} id="aoFDtpzmoCb2" outputId="cb3eeda2-d4f0-4afe-c52c-8b3f515e38f8"
doc = nlp("My IBAN number is BE71 0961 2345 6769, please send the money there.")
doc1 = nlp("My IBAN number is FR76 3000 6000 0112 3456 7890 189, please send the money there.")
matcher = Matcher(nlp.vocab)
pattern = [{"SHAPE": "XXdd"}, {"TEXT": {"REGEX": "\d{1,4}"}, "OP":"+"}]
matcher.add("ibanNum", [pattern])
for mid, start, end in matcher(doc):
print(start, end, doc[start:end])
print("**************************************")
for mid, start, end in matcher(doc1):
print(start, end, doc1[start:end])
# + id="jv6yFxtj99WW" outputId="828b9da0-0539-4128-f5e5-08b5a51f3ebc" colab={"base_uri": "https://localhost:8080/"}
doc = nlp("My account number is 8921273.")
matcher = Matcher(nlp.vocab)
pattern = [{"LOWER": "account"},{"LOWER": {"IN": ["num", "number"]}},{},{"IS_DIGIT": True}]
matcher.add("accountNum", [pattern])
for mid, start, end in matcher(doc):
print(start, end, doc[start:end])
# + [markdown] id="Jv9OP98m_lBf"
# Extracting phone number
# + id="GA2Hs07l-geH" outputId="e2d47752-6558-4fad-d5a5-dc6f4cb46533" colab={"base_uri": "https://localhost:8080/"}
doc1 = nlp("You can call my office on +1 (221) 102-2423 or email me directly.")
doc2 = nlp("You can call me on (221) 102 2423 or text me.")
matcher = Matcher(nlp.vocab)
pattern = [{"TEXT": "+1", "OP": "?"}, {"TEXT": "("}, {"SHAPE": "ddd"}, {"TEXT": ")"}, {"SHAPE": "ddd"}, {"TEXT": "-", "OP": "?"}, {"SHAPE": "dddd"}]
matcher.add("usPhonNum", [pattern])
for mid, start, end in matcher(doc1):
print(start, end, doc1[start:end])
print("**************************************")
for mid, start, end in matcher(doc2):
print(start, end, doc2[start:end])
# + [markdown] id="T-HcgMtDAcWF"
#
#
# Extracting mentions
# ```
# pattern = [{"ENT_TYPE": "ORG"}, {"LEMMA": "be"}, {"POS": "ADV", "OP":"*" {"POS": "ADJ"}]
# ```
# Hashtag and emoji extraction
#
# Expanding named entities
# + id="kJtOvUhUAKyI" outputId="be6f2b2f-b184-4140-ab70-45f60ee5e107" colab={"base_uri": "https://localhost:8080/"}
doc = nlp("<NAME> left her house 2 hours ago.")
doc.ents
# + id="vY1l6qHEFsz9" outputId="dc9c8b49-c2f2-4f88-db3a-c6f7697a638b" colab={"base_uri": "https://localhost:8080/"}
doc = nlp("<NAME> left her house")
patterns = [{"label": "TITLE", "pattern": [{"LOWER": {"IN": ["ms.", "mr.", "mrs.", "prof.", "dr."]}}]}]
ruler.add_patterns(patterns)
print([(ent.text, ent.label_) for ent in doc.ents])
# + [markdown] id="CozTRn52KM7V"
# Combining linguistic features and named entities
# + id="PqQlduTBF428" outputId="47ac6133-d8cd-443e-c9e4-2b0f0ecc5f31" colab={"base_uri": "https://localhost:8080/"}
doc = nlp("Einstein lived in Zurich.")
print([(ent.text, ent.label_) for ent in doc.ents])
# + id="GlzPtgCfKaD7" outputId="22015b46-d8ce-46f3-ace4-653be4747ec0" colab={"base_uri": "https://localhost:8080/"}
person_ents = [ent for ent in doc.ents if ent.label_ == "PERSON"]
for person_ent in person_ents:
# We use head of the entity's last token
head = person_ent[-1].head
if head.lemma_ == "live":
#Check if the children of live contains prepositional attachment
preps = [token for token in head.children if token.dep_ == "prep"]
for prep in preps:
places = [token for token in prep.children if token.ent_type_ == "GPE"]
# Verb is in past or present tense
print({'person': person_ent, 'city': places, 'past': head.tag_ == "VBD"})
# + id="S36JCKNmMTI-"
| Exploring Spacy3/Linguistic_Features_and_Rule_Based_Matching.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # TF-Slim Walkthrough
#
# This notebook will walk you through the basics of using TF-Slim to define, train and evaluate neural networks on various tasks. It assumes a basic knowledge of neural networks.
# ## Table of contents
#
# <a href="#Install">Installation and setup</a><br>
# <a href='#MLP'>Creating your first neural network with TF-Slim</a><br>
# <a href='#ReadingTFSlimDatasets'>Reading Data with TF-Slim</a><br>
# <a href='#CNN'>Training a convolutional neural network (CNN)</a><br>
# <a href='#Pretained'>Using pre-trained models</a><br>
#
# ## Installation and setup
# <a id='Install'></a>
#
# As of 8/28/16, the latest stable release of TF is r0.10, which does not contain the latest version of slim.
# To obtain the latest version of TF-Slim, please install the most recent nightly build of TF
# as explained [here](https://github.com/tensorflow/models/tree/master/slim#installing-latest-version-of-tf-slim).
#
# To use TF-Slim for image classification (as we do in this notebook), you also have to install the TF-Slim image models library from [here](https://github.com/tensorflow/models/tree/master/slim). Let's suppose you install this into a directory called TF_MODELS. Then you should change directory to TF_MODELS/slim **before** running this notebook, so that these files are in your python path.
#
# To check you've got these two steps to work, just execute the cell below. If it complains about unknown modules, restart the notebook after moving to the TF-Slim models directory.
#
# +
from __future__ import print_function
import matplotlib
# %matplotlib inline
import matplotlib.pyplot as plt
import math
import numpy as np
import os
import tensorflow as tf
import time
from datasets import dataset_utils
# Main slim library
slim = tf.contrib.slim
# -
from six.moves import xrange
from six import iteritems
# ## Creating your first neural network with TF-Slim
# <a id='MLP'></a>
#
# Below we give some code to create a simple multilayer perceptron (MLP) which can be used
# for regression problems. The model has 2 hidden layers.
# The output is a single node.
# When this function is called, it will create various nodes, and silently add them to whichever global TF graph is currently in scope. When a node which corresponds to a layer with adjustable parameters (eg., a fully connected layer) is created, additional parameter variable nodes are silently created, and added to the graph. (We will discuss how to train the parameters later.)
#
# We use variable scope to put all the nodes under a common name,
# so that the graph has some hierarchical structure.
# This is useful when we want to visualize the TF graph in tensorboard, or if we want to query related
# variables.
# The fully connected layers all use the same L2 weight decay and ReLu activations, as specified by **arg_scope**. (However, the final layer overrides these defaults, and uses an identity activation function.)
#
# We also illustrate how to add a dropout layer after the first fully connected layer (FC1). Note that at test time,
# we do not drop out nodes, but instead use the average activations; hence we need to know whether the model is being
# constructed for training or testing, since the computational graph will be different in the two cases
# (although the variables, storing the model parameters, will be shared, since they have the same name/scope).
def regression_model(inputs, is_training=True, scope="deep_regression"):
"""Creates the regression model.
Args:
inputs: A node that yields a `Tensor` of size [batch_size, dimensions].
is_training: Whether or not we're currently training the model.
scope: An optional variable_op scope for the model.
Returns:
predictions: 1-D `Tensor` of shape [batch_size] of responses.
end_points: A dict of end points representing the hidden layers.
"""
with tf.variable_scope(scope, 'deep_regression', [inputs]):
end_points = {}
# Set the default weight _regularizer and acvitation for each fully_connected layer.
with slim.arg_scope([slim.fully_connected],
activation_fn=tf.nn.relu,
weights_regularizer=slim.l2_regularizer(0.01)):
# Creates a fully connected layer from the inputs with 32 hidden units.
net = slim.fully_connected(inputs, 32, scope='fc1')
end_points['fc1'] = net
# Adds a dropout layer to prevent over-fitting.
net = slim.dropout(net, 0.8, is_training=is_training)
# Adds another fully connected layer with 16 hidden units.
net = slim.fully_connected(net, 16, scope='fc2')
end_points['fc2'] = net
# Creates a fully-connected layer with a single hidden unit. Note that the
# layer is made linear by setting activation_fn=None.
predictions = slim.fully_connected(net, 1, activation_fn=None, scope='prediction')
end_points['out'] = predictions
return predictions, end_points
# ### Let's create the model and examine its structure.
#
# We create a TF graph and call regression_model(), which adds nodes (tensors) to the graph. We then examine their shape, and print the names of all the model variables which have been implicitly created inside of each layer. We see that the names of the variables follow the scopes that we specified.
with tf.Graph().as_default():
# Dummy placeholders for arbitrary number of 1d inputs and outputs
inputs = tf.placeholder(tf.float32, shape=(None, 1))
outputs = tf.placeholder(tf.float32, shape=(None, 1))
# Build model
predictions, end_points = regression_model(inputs)
# Print name and shape of each tensor.
print("Layers")
for k, v in iteritems(end_points):
print('name = {}, shape = {}'.format(v.name, v.get_shape()))
# Print name and shape of parameter nodes (values not yet initialized)
print("\n")
print("Parameters")
for v in slim.get_model_variables():
print('name = {}, shape = {}'.format(v.name, v.get_shape()))
# ### Let's create some 1d regression data .
#
# We will train and test the model on some noisy observations of a nonlinear function.
#
# +
def produce_batch(batch_size, noise=0.3):
xs = np.random.random(size=[batch_size, 1]) * 10
ys = np.sin(xs) + 5 + np.random.normal(size=[batch_size, 1], scale=noise)
return [xs.astype(np.float32), ys.astype(np.float32)]
x_train, y_train = produce_batch(200)
x_test, y_test = produce_batch(200)
plt.scatter(x_train, y_train)
# -
# ### Let's fit the model to the data
#
# The user has to specify the loss function and the optimizer, and slim does the rest.
# In particular, the slim.learning.train function does the following:
#
# - For each iteration, evaluate the train_op, which updates the parameters using the optimizer applied to the current minibatch. Also, update the global_step.
# - Occasionally store the model checkpoint in the specified directory. This is useful in case your machine crashes - then you can simply restart from the specified checkpoint.
def convert_data_to_tensors(x, y):
inputs = tf.constant(x)
inputs.set_shape([None, 1])
outputs = tf.constant(y)
outputs.set_shape([None, 1])
return inputs, outputs
# +
# The following snippet trains the regression model using a sum_of_squares loss.
ckpt_dir = '/tmp/regression_model/'
with tf.Graph().as_default():
tf.logging.set_verbosity(tf.logging.INFO)
inputs, targets = convert_data_to_tensors(x_train, y_train)
# Make the model.
predictions, nodes = regression_model(inputs, is_training=True)
# Add the loss function to the graph.
loss = tf.losses.mean_squared_error(targets, predictions)
# The total loss is the uers's loss plus any regularization losses.
total_loss = slim.losses.get_total_loss()
# Specify the optimizer and create the train op:
optimizer = tf.train.AdamOptimizer(learning_rate=0.005)
train_op = slim.learning.create_train_op(total_loss, optimizer)
# Run the training inside a session.
final_loss = slim.learning.train(
train_op,
logdir=ckpt_dir,
number_of_steps=5000,
save_summaries_secs=5,
log_every_n_steps=500)
print("Finished training. Last batch loss:", final_loss)
print("Checkpoint saved in %s" % ckpt_dir)
# -
# ### Training with multiple loss functions.
#
# Sometimes we have multiple objectives we want to simultaneously optimize.
# In slim, it is easy to add more losses, as we show below. (We do not optimize the total loss in this example,
# but we show how to compute it.)
with tf.Graph().as_default():
inputs, targets = convert_data_to_tensors(x_train, y_train)
predictions, end_points = regression_model(inputs, is_training=True)
# Add multiple loss nodes.
sum_of_squares_loss = tf.losses.mean_squared_error(targets, predictions)
absolute_difference_loss = slim.losses.absolute_difference(predictions, targets)
# The following two ways to compute the total loss are equivalent
regularization_loss = tf.add_n(slim.losses.get_regularization_losses())
total_loss1 = sum_of_squares_loss + absolute_difference_loss + regularization_loss
# Regularization Loss is included in the total loss by default.
# This is good for training, but not for testing.
total_loss2 = slim.losses.get_total_loss(add_regularization_losses=True)
init_op = tf.global_variables_initializer()
with tf.Session() as sess:
sess.run(init_op) # Will initialize the parameters with random weights.
total_loss1, total_loss2 = sess.run([total_loss1, total_loss2])
print('Total Loss1: %f' % total_loss1)
print('Total Loss2: %f' % total_loss2)
print('Regularization Losses:')
for loss in slim.losses.get_regularization_losses():
print(loss)
print('Loss Functions:')
for loss in slim.losses.get_losses():
print(loss)
# ### Let's load the saved model and use it for prediction.
# +
with tf.Graph().as_default():
inputs, targets = convert_data_to_tensors(x_test, y_test)
# Create the model structure. (Parameters will be loaded below.)
predictions, end_points = regression_model(inputs, is_training=False)
# Make a session which restores the old parameters from a checkpoint.
sv = tf.train.Supervisor(logdir=ckpt_dir)
with sv.managed_session() as sess:
inputs, predictions, targets = sess.run([inputs, predictions, targets])
plt.scatter(inputs, targets, c='r');
plt.scatter(inputs, predictions, c='b');
plt.title('red=true, blue=predicted')
# -
# ### Let's compute various evaluation metrics on the test set.
#
# In TF-Slim termiology, losses are optimized, but metrics (which may not be differentiable, e.g., precision and recall) are just measured. As an illustration, the code below computes mean squared error and mean absolute error metrics on the test set.
#
# Each metric declaration creates several local variables (which must be initialized via tf.initialize_local_variables()) and returns both a value_op and an update_op. When evaluated, the value_op returns the current value of the metric. The update_op loads a new batch of data, runs the model, obtains the predictions and accumulates the metric statistics appropriately before returning the current value of the metric. We store these value nodes and update nodes in 2 dictionaries.
#
# After creating the metric nodes, we can pass them to slim.evaluation.evaluation, which repeatedly evaluates these nodes the specified number of times. (This allows us to compute the evaluation in a streaming fashion across minibatches, which is usefulf for large datasets.) Finally, we print the final value of each metric.
#
# +
# Updated with code from section "Evaluating a Checkpointed Model with Metrics"
# of https://github.com/tensorflow/tensorflow/blob/master/tensorflow/contrib/training/python/training/evaluation.py
with tf.Graph().as_default():
inputs, targets = convert_data_to_tensors(x_test, y_test)
predictions, end_points = regression_model(inputs, is_training=False)
# Specify metrics to evaluate:
names_to_value_nodes, names_to_update_nodes = slim.metrics.aggregate_metric_map({
'Mean Squared Error': slim.metrics.streaming_mean_squared_error(predictions, targets),
'Mean Absolute Error': slim.metrics.streaming_mean_absolute_error(predictions, targets)
})
metric_values = tf.contrib.training.evaluate_once(
os.path.join(ckpt_dir, 'model.ckpt'),
eval_ops=list(names_to_update_nodes.values()),
final_ops=list(names_to_value_nodes.values()),
hooks=[
tf.contrib.training.StopAfterNEvalsHook(1)
],)
names_to_values = dict(zip(names_to_value_nodes.keys(), metric_values))
for key, value in iteritems(names_to_values):
print('%s: %f' % (key, value))
# -
# # Reading Data with TF-Slim
# <a id='ReadingTFSlimDatasets'></a>
#
# Reading data with TF-Slim has two main components: A
# [Dataset](https://github.com/tensorflow/tensorflow/blob/master/tensorflow/contrib/slim/python/slim/data/dataset.py) and a
# [DatasetDataProvider](https://github.com/tensorflow/tensorflow/blob/master/tensorflow/contrib/slim/python/slim/data/dataset_data_provider.py). The former is a descriptor of a dataset, while the latter performs the actions necessary for actually reading the data. Lets look at each one in detail:
#
#
# ## Dataset
# A TF-Slim
# [Dataset](https://github.com/tensorflow/tensorflow/blob/master/tensorflow/contrib/slim/python/slim/data/dataset.py)
# contains descriptive information about a dataset necessary for reading it, such as the list of data files and how to decode them. It also contains metadata including class labels, the size of the train/test splits and descriptions of the tensors that the dataset provides. For example, some datasets contain images with labels. Others augment this data with bounding box annotations, etc. The Dataset object allows us to write generic code using the same API, regardless of the data content and encoding type.
#
# TF-Slim's Dataset works especially well when the data is stored as a (possibly sharded)
# [TFRecords file](https://www.tensorflow.org/versions/r0.10/how_tos/reading_data/index.html#file-formats), where each record contains a [tf.train.Example protocol buffer](https://github.com/tensorflow/tensorflow/blob/r0.10/tensorflow/core/example/example.proto).
# TF-Slim uses a consistent convention for naming the keys and values inside each Example record.
#
# ## DatasetDataProvider
#
# A
# [DatasetDataProvider](https://github.com/tensorflow/tensorflow/blob/master/tensorflow/contrib/slim/python/slim/data/dataset_data_provider.py) is a class which actually reads the data from a dataset. It is highly configurable to read the data in various ways that may make a big impact on the efficiency of your training process. For example, it can be single or multi-threaded. If your data is sharded across many files, it can read each files serially, or from every file simultaneously.
#
# ## Demo: The Flowers Dataset
#
# For convenience, we've include scripts to convert several common image datasets into TFRecord format and have provided
# the Dataset descriptor files necessary for reading them. We demonstrate how easy it is to use these dataset via the Flowers dataset below.
# ### Download the Flowers Dataset
# <a id='DownloadFlowers'></a>
#
# We've made available a tarball of the Flowers dataset which has already been converted to TFRecord format.
# +
import tensorflow as tf
from datasets import dataset_utils
url = "http://download.tensorflow.org/data/flowers.tar.gz"
flowers_data_dir = '/tmp/flowers'
if not tf.gfile.Exists(flowers_data_dir):
tf.gfile.MakeDirs(flowers_data_dir)
dataset_utils.download_and_uncompress_tarball(url, flowers_data_dir)
# -
# ### Display some of the data.
# +
from datasets import flowers
import tensorflow as tf
slim = tf.contrib.slim
with tf.Graph().as_default():
dataset = flowers.get_split('train', flowers_data_dir)
data_provider = slim.dataset_data_provider.DatasetDataProvider(
dataset, common_queue_capacity=32, common_queue_min=1)
image, label = data_provider.get(['image', 'label'])
with tf.Session() as sess:
with slim.queues.QueueRunners(sess):
for i in xrange(4):
np_image, np_label = sess.run([image, label])
height, width, _ = np_image.shape
class_name = name = dataset.labels_to_names[np_label]
plt.figure()
plt.imshow(np_image)
plt.title('%s, %d x %d' % (name, height, width))
plt.axis('off')
plt.show()
# -
# # Convolutional neural nets (CNNs).
# <a id='CNN'></a>
#
# In this section, we show how to train an image classifier using a simple CNN.
#
# ### Define the model.
#
# Below we define a simple CNN. Note that the output layer is linear function - we will apply softmax transformation externally to the model, either in the loss function (for training), or in the prediction function (during testing).
def my_cnn(images, num_classes, is_training): # is_training is not used...
with slim.arg_scope([slim.max_pool2d], kernel_size=[3, 3], stride=2):
net = slim.conv2d(images, 64, [5, 5])
net = slim.max_pool2d(net)
net = slim.conv2d(net, 64, [5, 5])
net = slim.max_pool2d(net)
net = slim.flatten(net)
net = slim.fully_connected(net, 192)
net = slim.fully_connected(net, num_classes, activation_fn=None)
return net
# ### Apply the model to some randomly generated images.
# +
import tensorflow as tf
with tf.Graph().as_default():
# The model can handle any input size because the first layer is convolutional.
# The size of the model is determined when image_node is first passed into the my_cnn function.
# Once the variables are initialized, the size of all the weight matrices is fixed.
# Because of the fully connected layers, this means that all subsequent images must have the same
# input size as the first image.
batch_size, height, width, channels = 3, 28, 28, 3
images = tf.random_uniform([batch_size, height, width, channels], maxval=1)
# Create the model.
num_classes = 10
logits = my_cnn(images, num_classes, is_training=True)
probabilities = tf.nn.softmax(logits)
# Initialize all the variables (including parameters) randomly.
init_op = tf.global_variables_initializer()
with tf.Session() as sess:
# Run the init_op, evaluate the model outputs and print the results:
sess.run(init_op)
probabilities = sess.run(probabilities)
print('Probabilities Shape:')
print(probabilities.shape) # batch_size x num_classes
print('\nProbabilities:')
print(probabilities)
print('\nSumming across all classes (Should equal 1):')
print(np.sum(probabilities, 1)) # Each row sums to 1
# -
# ### Train the model on the Flowers dataset.
#
# Before starting, make sure you've run the code to <a href="#DownloadFlowers">Download the Flowers</a> dataset. Now, we'll get a sense of what it looks like to use TF-Slim's training functions found in
# [learning.py](https://github.com/tensorflow/tensorflow/blob/master/tensorflow/contrib/slim/python/slim/learning.py). First, we'll create a function, `load_batch`, that loads batches of dataset from a dataset. Next, we'll train a model for a single step (just to demonstrate the API), and evaluate the results.
# +
from preprocessing import inception_preprocessing
import tensorflow as tf
slim = tf.contrib.slim
def load_batch(dataset, batch_size=32, height=299, width=299, is_training=False):
"""Loads a single batch of data.
Args:
dataset: The dataset to load.
batch_size: The number of images in the batch.
height: The size of each image after preprocessing.
width: The size of each image after preprocessing.
is_training: Whether or not we're currently training or evaluating.
Returns:
images: A Tensor of size [batch_size, height, width, 3], image samples that have been preprocessed.
images_raw: A Tensor of size [batch_size, height, width, 3], image samples that can be used for visualization.
labels: A Tensor of size [batch_size], whose values range between 0 and dataset.num_classes.
"""
data_provider = slim.dataset_data_provider.DatasetDataProvider(
dataset, common_queue_capacity=32,
common_queue_min=8)
image_raw, label = data_provider.get(['image', 'label'])
# Preprocess image for usage by Inception.
image = inception_preprocessing.preprocess_image(image_raw, height, width, is_training=is_training)
# Preprocess the image for display purposes.
image_raw = tf.expand_dims(image_raw, 0)
image_raw = tf.image.resize_images(image_raw, [height, width])
image_raw = tf.squeeze(image_raw)
# Batch it up.
images, images_raw, labels = tf.train.batch(
[image, image_raw, label],
batch_size=batch_size,
num_threads=1,
capacity=2 * batch_size)
return images, images_raw, labels
# +
from datasets import flowers
# This might take a few minutes.
train_dir = '/tmp/tfslim_model/'
print('Will save model to %s' % train_dir)
with tf.Graph().as_default():
tf.logging.set_verbosity(tf.logging.INFO)
dataset = flowers.get_split('train', flowers_data_dir)
images, _, labels = load_batch(dataset)
# Create the model:
logits = my_cnn(images, num_classes=dataset.num_classes, is_training=True)
# Specify the loss function:
one_hot_labels = slim.one_hot_encoding(labels, dataset.num_classes)
slim.losses.softmax_cross_entropy(logits, one_hot_labels)
total_loss = slim.losses.get_total_loss()
# Create some summaries to visualize the training process:
tf.summary.scalar('losses/Total Loss', total_loss)
# Specify the optimizer and create the train op:
optimizer = tf.train.AdamOptimizer(learning_rate=0.01)
train_op = slim.learning.create_train_op(total_loss, optimizer)
# Run the training:
final_loss = slim.learning.train(
train_op,
logdir=train_dir,
number_of_steps=10, # For speed, we just do 1 epoch
save_summaries_secs=1)
print('Finished training. Final batch loss %d' % final_loss)
# -
# ### Evaluate some metrics.
#
# As we discussed above, we can compute various metrics besides the loss.
# Below we show how to compute prediction accuracy of the trained model, as well as top-5 classification accuracy. (The difference between evaluation and evaluation_loop is that the latter writes the results to a log directory, so they can be viewed in tensorboard.)
# +
from datasets import flowers
# This might take a few minutes.
with tf.Graph().as_default():
tf.logging.set_verbosity(tf.logging.DEBUG)
dataset = flowers.get_split('train', flowers_data_dir)
images, _, labels = load_batch(dataset)
logits = my_cnn(images, num_classes=dataset.num_classes, is_training=False)
predictions = tf.argmax(logits, 1)
# Define the metrics:
names_to_values, names_to_updates = slim.metrics.aggregate_metric_map({
'eval/Accuracy': slim.metrics.streaming_accuracy(predictions, labels),
'eval/Recall@5': slim.metrics.streaming_recall_at_k(logits, labels, 5),
})
print('Running evaluation Loop...')
checkpoint_path = tf.train.latest_checkpoint(train_dir)
metric_values = slim.evaluation.evaluate_once(
master='',
checkpoint_path=checkpoint_path,
logdir=train_dir,
eval_op=list(names_to_updates.values()),
final_op=list(names_to_values.values()))
names_to_values = dict(zip(names_to_values.keys(), metric_values))
for name in names_to_values:
print('%s: %f' % (name, names_to_values[name]))
# -
# # Using pre-trained models
# <a id='Pretrained'></a>
#
# Neural nets work best when they have many parameters, making them very flexible function approximators.
# However, this means they must be trained on big datasets. Since this process is slow, we provide various pre-trained models - see the list [here](https://github.com/tensorflow/models/tree/master/slim#pre-trained-models).
#
#
# You can either use these models as-is, or you can perform "surgery" on them, to modify them for some other task. For example, it is common to "chop off" the final pre-softmax layer, and replace it with a new set of weights corresponding to some new set of labels. You can then quickly fine tune the new model on a small new dataset. We illustrate this below, using inception-v1 as the base model. While models like Inception V3 are more powerful, Inception V1 is used for speed purposes.
#
# ### Download the Inception V1 checkpoint
#
#
#
# +
from datasets import dataset_utils
url = "http://download.tensorflow.org/models/inception_v1_2016_08_28.tar.gz"
checkpoints_dir = '/tmp/checkpoints'
if not tf.gfile.Exists(checkpoints_dir):
tf.gfile.MakeDirs(checkpoints_dir)
dataset_utils.download_and_uncompress_tarball(url, checkpoints_dir)
# -
#
# ### Apply Pre-trained model to Images.
#
# We have to convert each image to the size expected by the model checkpoint.
# There is no easy way to determine this size from the checkpoint itself.
# So we use a preprocessor to enforce this.
# +
import numpy as np
import tensorflow as tf
#import urllib2
from datasets import imagenet
from nets import inception
from preprocessing import inception_preprocessing
slim = tf.contrib.slim
batch_size = 3
image_size = inception.inception_v1.default_image_size
with tf.Graph().as_default():
#url = 'https://upload.wikimedia.org/wikipedia/commons/7/70/EnglishCockerSpaniel_simon.jpg'
#image_string = urllib2.urlopen(url).read()
image_string = open("/tmp/EnglishCockerSpaniel_simon.jpg", mode='rb').read()
image = tf.image.decode_jpeg(image_string, channels=3)
processed_image = inception_preprocessing.preprocess_image(image, image_size, image_size, is_training=False)
processed_images = tf.expand_dims(processed_image, 0)
# Create the model, use the default arg scope to configure the batch norm parameters.
with slim.arg_scope(inception.inception_v1_arg_scope()):
logits, _ = inception.inception_v1(processed_images, num_classes=1001, is_training=False)
probabilities = tf.nn.softmax(logits)
init_fn = slim.assign_from_checkpoint_fn(
os.path.join(checkpoints_dir, 'inception_v1.ckpt'),
slim.get_model_variables('InceptionV1'))
with tf.Session() as sess:
init_fn(sess)
np_image, probabilities = sess.run([image, probabilities])
probabilities = probabilities[0, 0:]
sorted_inds = [i[0] for i in sorted(enumerate(-probabilities), key=lambda x:x[1])]
plt.figure()
plt.imshow(np_image.astype(np.uint8))
plt.axis('off')
plt.show()
names = imagenet.create_readable_names_for_imagenet_labels()
for i in xrange(5):
index = sorted_inds[i]
print('Probability %0.2f%% => [%s]' % (probabilities[index]*100, names[index]))
# -
# ### Fine-tune the model on a different set of labels.
#
# We will fine tune the inception model on the Flowers dataset.
# +
# Note that this may take several minutes.
import os
from datasets import flowers
from nets import inception
from preprocessing import inception_preprocessing
slim = tf.contrib.slim
image_size = inception.inception_v1.default_image_size
def get_init_fn():
"""Returns a function run by the chief worker to warm-start the training."""
checkpoint_exclude_scopes=["InceptionV1/Logits", "InceptionV1/AuxLogits"]
exclusions = [scope.strip() for scope in checkpoint_exclude_scopes]
variables_to_restore = []
for var in slim.get_model_variables():
excluded = False
for exclusion in exclusions:
if var.op.name.startswith(exclusion):
excluded = True
break
if not excluded:
variables_to_restore.append(var)
return slim.assign_from_checkpoint_fn(
os.path.join(checkpoints_dir, 'inception_v1.ckpt'),
variables_to_restore)
train_dir = '/tmp/inception_finetuned/'
with tf.Graph().as_default():
tf.logging.set_verbosity(tf.logging.INFO)
dataset = flowers.get_split('train', flowers_data_dir)
images, _, labels = load_batch(dataset, height=image_size, width=image_size)
# Create the model, use the default arg scope to configure the batch norm parameters.
with slim.arg_scope(inception.inception_v1_arg_scope()):
logits, _ = inception.inception_v1(images, num_classes=dataset.num_classes, is_training=True)
# Specify the loss function:
one_hot_labels = slim.one_hot_encoding(labels, dataset.num_classes)
slim.losses.softmax_cross_entropy(logits, one_hot_labels)
total_loss = slim.losses.get_total_loss()
# Create some summaries to visualize the training process:
tf.summary.scalar('losses/Total Loss', total_loss)
# Specify the optimizer and create the train op:
optimizer = tf.train.AdamOptimizer(learning_rate=0.01)
train_op = slim.learning.create_train_op(total_loss, optimizer)
# Run the training:
final_loss = slim.learning.train(
train_op,
logdir=train_dir,
init_fn=get_init_fn(),
number_of_steps=100)
print('Finished training. Last batch loss %f' % final_loss)
# -
# ### Apply fine tuned model to some images.
# +
import numpy as np
import tensorflow as tf
from datasets import flowers
from nets import inception
slim = tf.contrib.slim
image_size = inception.inception_v1.default_image_size
batch_size = 3
with tf.Graph().as_default():
tf.logging.set_verbosity(tf.logging.INFO)
dataset = flowers.get_split('train', flowers_data_dir)
images, images_raw, labels = load_batch(dataset, height=image_size, width=image_size)
# Create the model, use the default arg scope to configure the batch norm parameters.
with slim.arg_scope(inception.inception_v1_arg_scope()):
logits, _ = inception.inception_v1(images, num_classes=dataset.num_classes, is_training=True)
probabilities = tf.nn.softmax(logits)
checkpoint_path = tf.train.latest_checkpoint(train_dir)
init_fn = slim.assign_from_checkpoint_fn(
checkpoint_path,
slim.get_variables_to_restore())
with tf.Session() as sess:
with slim.queues.QueueRunners(sess):
sess.run(tf.initialize_local_variables())
init_fn(sess)
np_probabilities, np_images_raw, np_labels = sess.run([probabilities, images_raw, labels])
for i in xrange(batch_size):
image = np_images_raw[i, :, :, :]
true_label = np_labels[i]
predicted_label = np.argmax(np_probabilities[i, :])
predicted_name = dataset.labels_to_names[predicted_label]
true_name = dataset.labels_to_names[true_label]
plt.figure()
plt.imshow(image.astype(np.uint8))
plt.title('Ground Truth: [%s], Prediction [%s]' % (true_name, predicted_name))
plt.axis('off')
plt.show()
# -
| slim/slim_walkthough.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.4 64-bit (''var-gp'': conda)'
# name: python394jvsc74a57bd0f6f4ef2e4dfc72dba41423bb399455b0e2be86e7ec916ab82ad022c9707621b9
# ---
# +
import os
import sys
import numpy as np
import torch
from torch.utils.data import DataLoader
import matplotlib.pyplot as plt
import seaborn as sns
import pandas as pd
sns.set(font_scale=2, style='whitegrid')
device = 'cuda' if torch.cuda.is_available() else 'cpu'
save = False
if os.path.abspath('..') not in sys.path:
sys.path.append(os.path.abspath('..'))
# -
from var_gp.datasets import ToyDataset
from var_gp.vargp import VARGP
from var_gp.train_utils import set_seeds
# +
set_seeds(1)
toy_ds = ToyDataset()
df = pd.DataFrame({ 'x': toy_ds.data[:, 0].numpy(), 'y': toy_ds.data[:, 1].numpy(), 'Class': toy_ds.targets.numpy() })
fig, ax = plt.subplots(figsize=(9,9))
sns.scatterplot(ax=ax, data=df, x='x', y='y', hue='Class', palette='Set2', s=200, edgecolor='black', linewidth=2)
ax.set_xlabel('')
ax.set_ylabel('')
handles, labels = ax.get_legend_handles_labels()
for h, l in zip(handles, labels):
h.set_edgecolor('black')
h.set_linewidth(2)
h.set_sizes([200])
ax.legend(handles=handles, labels=labels, title='Class');
# fig.savefig('toy_data.pdf', bbox_inches='tight')
# +
grid_data = torch.cat([v.unsqueeze(-1) for v in torch.meshgrid([torch.arange(-3,3,0.1), torch.arange(-3,3,0.1)])], dim=-1).permute(1, 0, 2)
def plot_task(preds):
out = preds.reshape(preds.size(0), *grid_data.shape[:-1], -1)
fig, axes = plt.subplots(2, 4, sharey=True, sharex=True, figsize=(40, 20))
for r in range(2):
for i in range(preds.size(-1)):
toy_ds.filter_by_class([i])
axes[r, i].contourf(out[r, ..., i], cmap=sns.color_palette("Blues_r", as_cmap=True),
extent=(-3,3,-3,3), origin='lower')
axes[r, i].set(aspect='equal')
axes[r, i].set_xlim(-3, 3)
axes[r, i].set_ylim(-3, 3)
axes[r, i].grid(False)
axes[r, i].set_xticks([])
axes[r, i].set_yticks([])
if r == 0:
axes[r, i].set_title(f'Class {i}', fontsize=75)
axes[r, i].scatter(toy_ds.data[toy_ds.task_ids][:, 0], toy_ds.data[toy_ds.task_ids][:, 1],
marker='o', facecolor='red', s=400, edgecolor='black', linewidth=2)
axes[r, 0].set_ylabel(f'After Task {r}', fontsize=75)
# Reset filter.
toy_ds.filter_by_class()
fig.tight_layout()
return fig, axes
# +
run_dir = 'results/vargp-toy-seed1'
prev_params = []
preds = []
for t in range(2):
with torch.no_grad():
cur_params = torch.load(f'{run_dir}/ckpt{t}.pt')
gp = VARGP.create_clf(toy_ds, M=20, n_f=100, n_var_samples=20, prev_params=prev_params).to(device)
gp.load_state_dict(cur_params)
preds.append(gp.predict(grid_data.reshape(-1, 2)))
prev_params.append(cur_params)
preds = torch.cat([p.unsqueeze(0) for p in preds], axis=0)
fig, _ = plot_task(preds)
# fig.savefig(f'toy_vargp_density.pdf', bbox_inches='tight')
# +
from var_gp.vargp_retrain import VARGPRetrain
run_dir = 'results/re-vargp-toy'
prev_params = []
preds = []
for t in range(2):
with torch.no_grad():
cur_params = torch.load(f'{run_dir}/ckpt{t}.pt')
gp = VARGPRetrain.create_clf(toy_ds, M=20, n_f=100, n_var_samples=20, prev_params=prev_params).to(device)
gp.load_state_dict(cur_params)
preds.append(gp.predict(grid_data.reshape(-1, 2)))
prev_params.append(cur_params)
preds = torch.cat([p.unsqueeze(0) for p in preds], axis=0)
fig, _ = plot_task(preds)
# fig.savefig(f'toy_vargp_retrain_density.pdf', bbox_inches='tight')
# +
pred_dump = 'results/vcl-toy-seed1'
preds = []
for t in range(2):
data = np.load(f'{pred_dump}/grid_pred_probs_{t}.npz')
preds.append(torch.from_numpy(np.squeeze(data['probs'], axis=-1).T).float())
preds = torch.cat([p.unsqueeze(0) for p in preds], axis=0)
fig, _ = plot_task(preds)
# fig.savefig(f'toy_vcl_density.pdf', bbox_inches='tight')
# -
| notebooks/toy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Extract MP components from synthetic data and write to MAT file
# +
import h5py
import numpy as np
import matplotlib.pyplot as plt
from skimage.util import img_as_float
from scipy.io import savemat
from imagerep import mp_gaussian, reconstruct_image
# +
# Input and output paths
IN_FPATH = '/home/mn2822/Desktop/WormTraces/animal_056_head/run401.mat'
OUT_FPATH = '/home/mn2822/Desktop/WormTracking/data/synthetic/syn_data_mp.mat'
# Start and stop times for extraction
T_START = 0
T_STOP = 50
# Covariance values for each dimension
#COV_DIAG = [4.0, 4.0, 1.0]
COV_DIAG = [5.0, 5.0, 5.0]
# Number of MP iterations to run
#N_ITER = 500
N_ITER = 300
# +
cov = np.diag(COV_DIAG)
means = []
weights = []
with h5py.File(IN_FPATH, 'r') as f:
dset = f.get('data')
#for t in range(T_START, T_STOP):
for t in range(1):
print(f'Frame: {t}')
# Load frame
img_raw = dset[t, :, :, :]
img_raw = np.moveaxis(img_raw, [0, 1, 2], [2, 0, 1])
img = img_as_float(img_raw)
# Extract MP components from frame
mus, wts, _ = mp_gaussian(img, cov, N_ITER)
means.append(mus)
weights.append(wts)
# +
img_recon = reconstruct_image(mus, [cov] * N_ITER, wts, img.shape)
plt.figure(figsize=(15, 15))
plt.subplot(121)
plt.imshow(np.max(img, 2).T)
plt.subplot(122)
plt.imshow(np.max(img_recon, 2).T)
# -
| python/archive/mp_vivek_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Energy Efficiency
#
# Following image represents the energy consumption in US buildings:
#
# <img src='Pictures/Statistics.gif'/>
#
# (Couresty: http://homepages.cae.wisc.edu/~ece539/fall13/project/Wysocki.pptx)
#
# Heating, ventilation, and air conditioning(HVAC) contributes to higher % of energy consumption. In this study, we are focusing on accurately predicting the energy consumption (heating and colling load) of equipments to choose building shapes efficiently.
#
# > i.e., this notebook demonstrates the how to effectively design the energy requirements for residential buildings.
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.feature_selection import RFE
from sklearn.ensemble import ExtraTreesRegressor
from sklearn.model_selection import train_test_split
import warnings
warnings.filterwarnings('ignore')
import xgboost
from sklearn.metrics import accuracy_score
from sklearn import model_selection
# ### Data Set Information
#
# (Courtesy: https://archive.ics.uci.edu/ml/datasets/Energy+efficiency)
#
# We perform energy analysis using 12 different building shapes simulated in Ecotect. The buildings differ with respect to the glazing area, the glazing area distribution, and the orientation, amongst other parameters. We simulate various settings as functions of the afore-mentioned characteristics to obtain 768 building shapes. The dataset comprises 768 samples and 8 features, aiming to predict two real valued responses.
# ### Load Dataset
energy = pd.read_excel('Dataset/data14.xlsx')
energy
# ### Attribute Information:
#
# The dataset contains eight attributes (or features, denoted by X1...X8) and two responses (or outcomes, denoted by y1 and y2). The aim is to use the eight features to predict each of the two responses.
#
# Specifically:<br/>
# X1 Relative Compactness <br/>
# X2 Surface Area<br/>
# X3 Wall Area<br/>
# X4 Roof Area<br/>
# X5 Overall Height<br/>
# X6 Orientation<br/>
# X7 Glazing Area<br/>
# X8 Glazing Area Distribution<br/>
# Y1 Heating Load<br/>
# Y2 Cooling Load<br/>
#
# (Courtesy: https://archive.ics.uci.edu/ml/datasets/Energy+efficiency)
# ### Data cleaning and Data Analysis
# Dimensions
energy.shape
# Check for missing Values
energy.isna().sum()
# <b>There are No missing values.</b>
#
# Renaming the column names of the Dataset
energy.columns= [
'rel_compactness', 'surface_area', 'wall_area', 'roof_area', 'overall_height','orientation',
'glazing_area', 'glazing_area_dist', 'heating_load', 'cooling_load'
]
energy
# ### Finding correlation between features
#
# We systematically investigate the association strength of each input variable with each of the output variables in order to identify the most strongly related input variables.
correlation = energy.corr()
correlation.round(decimals=4)
correlation.style.background_gradient(cmap='coolwarm').set_precision(4)
# ### Observations
# As we can see from correlation matrix:
# > 1. rel_compactness, wall_area and overall_height are highly positively correlated with heating and cooling load
# > 2. surface area and roof_area is highly negatively correlated with heating_load and cooling load.
# Getting Features for Input (X) and Response (Y)
X = energy.drop(['heating_load', 'cooling_load'], axis=1)
X
Y = energy['heating_load']
Y
# ### Model Building (for Y='Heating Load') using RFE
#
# #### Build a model to predict the Heating Load of Heating equipment.
# Feature Selection and Model Building
#
# > ExtraTreesRegressor() and RandomForestRegression() perform with almost same efficiency.
# >(Courtesy: https://quantdare.com/what-is-the-difference-between-extra-trees-and-random-forest/)
model = ExtraTreesRegressor()
# #### ABOUT RFE
#
# Feature ranking with recursive feature elimination.
#
# >This technique begins by building a model on the entire set of predictors and computing an importance score for each predictor. The least important predictor(s) are then removed, the model is re-built, and importance scores are computed again. In practice, the analyst specifies the number of predictor subsets to evaluate as well as each subset’s size. Therefore, the subset size is a tuning parameter for RFE. The subset size that optimizes the performance criteria is used to select the predictors based on the importance rankings. The optimal subset is then used to train the final model.
#
# (Courtesy: https://towardsdatascience.com/feature-selection-in-python-recursive-feature-elimination-19f1c39b8d15)
# For syntax of RFE: https://scikit-learn.org/stable/modules/generated/sklearn.feature_selection.RFE.html
rfe = RFE(estimator=model, n_features_to_select=3, step=1)
rfe
fit = rfe.fit(X, Y)
print('Number of Features: ', fit.n_features_)
# Selected Features
fit.support_
selected_features = [each_feature for each_feature, status in zip(X.columns, fit.support_) if status == True]
print('Selected Features:\n', *selected_features, sep="\n")
for feature, value in zip(X.columns, fit.ranking_):
print(feature, "--- (", value, ")")
# ### Plotting Importance of Features
imp_values = list(fit.estimator_.feature_importances_)
imp_values
imp_values.reverse()
imp_values
# +
importance = []
for value in fit.support_:
if value == True:
importance.append(imp_values.pop())
else:
importance.append(0)
importance
# -
plotting_data = pd.DataFrame({'Features': X.columns, 'Importance':importance}, index=X.columns)
plotting_data
plotting_data.plot.barh()
# ### Fit the Model
seed = 7 # Can be any value. Keep it same to repoduce the same results
n_estimators = len(X)
model.fit(X, Y)
# ### Evaluate the Model
# >Repeated k-fold cross-validation provides a way to improve the estimated performance of a machine learning model. This involves simply repeating the cross-validation procedure multiple times and reporting the mean result across all folds from all runs. This mean result is expected to be a more accurate estimate of the true unknown underlying mean performance of the model on the dataset, as calculated using the standard error.
#
# (Courtesy: https://machinelearningmastery.com/repeated-k-fold-cross-validation-with-python/)
kfold = model_selection.RepeatedKFold(n_splits=n_estimators, n_repeats=10, random_state=seed)
cv_results = model_selection.cross_val_score(model, X, Y, cv=10)
cv_results
msg = "%s: %f (%f)" % ("ExtraTreesRegressor", cv_results.mean(), cv_results.std())
msg
# ### Model Building (for Y='Cooling Load')
# >The main issue of RFE is that it can be expensive to run.
#
# Removing correlated features are beneficial as the highly correlated features in the dataset provide the same information
#
# (Courtesy: https://towardsdatascience.com/feature-selection-in-python-recursive-feature-elimination-19f1c39b8d15)
data = energy.drop(['heating_load', 'cooling_load'], axis=1)
data
correlation_matrix = data.corr()
correlation_matrix.round(decimals=4)
correlation_matrix.style.background_gradient(cmap='coolwarm').set_precision(4)
# Omitting highly correlated features
# +
correlated_features = set()
for i in range(len(correlation_matrix.columns)):
for j in range(i):
if abs(correlation_matrix.iloc[i, j]) > 0.8:
colname = correlation_matrix.columns[i]
correlated_features.add(colname)
correlated_features.add('heating_load')
print("Highly Correlated Features: ", correlated_features)
# -
# Get new Input and Target data
req_data = energy.drop(list(correlated_features), axis=1)
req_data
X = req_data.drop(['cooling_load'], axis=1)
target = req_data['cooling_load']
X
target
#
# #### Build a model to predict the Cooling Load of Heating equipment.
# Feature Selection and Model Building
model = ExtraTreesRegressor()
from sklearn.feature_selection import RFECV
# #### ABOUT RFECV
#
# Feature ranking with recursive feature elimination and cross-validated selection of the best number of features.
#
# <img src='Pictures/14.png'/>
#
# (Courtesy: https://scikit-learn.org/stable/modules/generated/sklearn.feature_selection.RFECV.html)
rfecv = RFECV(model, step=1)
fit = rfecv.fit(X, target)
print('Optimal number of features: {}'.format(rfecv.n_features_))
# ### Plot the accuracy obtained with every number of features used
#
# >From the plot we can identify how many features has to be selected (instead of explicity choosing a value without prior knowledge of dataset).
#
# (Courtesy: https://towardsdatascience.com/feature-selection-in-python-recursive-feature-elimination-19f1c39b8d15)
# +
plt.figure(figsize=(16, 9))
plt.title('Recursive Feature Elimination with Cross-Validation', fontsize=18, fontweight='bold', pad=20)
plt.xlabel('Number of features Choosen', fontsize=14, labelpad=20)
plt.ylabel('% Correct Classification', fontsize=14, labelpad=20)
plt.plot(range(1, len(rfecv.grid_scores_) + 1), rfecv.grid_scores_, color='#303F9F', linewidth=3)
plt.show()
# -
rfecv.estimator_.feature_importances_
# Selected Features
rfecv.support_
selected_features = [each_feature for each_feature, status in zip(X.columns, rfecv.support_) if status == True]
print('Selected Features:\n', *selected_features, sep="\n")
for feature, value in zip(X.columns, rfecv.ranking_):
print(feature, "--- (", value, ")")
imp_values = list(rfecv.estimator_.feature_importances_)
imp_values
imp_values.reverse()
imp_values
# +
importance = []
for value in fit.support_:
if value == True:
importance.append(imp_values.pop())
else:
importance.append(0)
importance
# -
plotting_data = pd.DataFrame({'Features': X.columns, 'Importance':importance}, index=X.columns)
plotting_data
plotting_data.plot.barh()
# ### Fit the Model
seed = 7
n_estimators = len(X)
model.fit(X, target)
# ### Evaluate the Model
kfold = model_selection.RepeatedKFold(n_splits=n_estimators, n_repeats=10, random_state=seed)
cv_results = model_selection.cross_val_score(model, X, target, cv=10)
cv_results
msg = "%s: %f (%f)" % ("ExtraTreesRegressor", cv_results.mean(), cv_results.std())
msg
# ## Summary
#
# For Y1 (Heating Load)
# > 1. 'surface_area', 'roof_area' and 'overall_height' were top 3 selected for predicting 'heating_load'.
# > 2. The accuracy of prediction by ExtraTreeRegressor is ~97%.
#
# For Y2 (Cooling Load)
# > 1. 'rel_compactness', 'wall_area' and 'glazing_area' were top 3 selected for predicting 'cooling_load'.
# > 2. The accuracy of prediction by ExtraTreeRegressor is ~95%.
#
# The results of this study support the feasibility of using machine learning tools to estimate building parameters as a convenient and accurate approach, as long as the requested query bears resemblance to the data actually used to train the mathematical model in the first place.
#
# (Courtesy: https://www.semanticscholar.org/paper/Accurate-quantitative-estimation-of-energy-of-using-Tsanas-Xifara/719e65379c5959141180a45f540f707d583b8ce2)
# Prepared by J.Haripriya
| Energy Efficiency Dataset Analysis/Energy Efficiency Dataset Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from surprise import KNNBasic
from surprise import Reader
from surprise import Dataset
import pandas as pd
# Cargar los registros de usuarios
ratings_df = pd.read_csv('ratings.dat', sep='::', header=None)
del (ratings_df[3])# borrar la tercera fila correspondiente a la fecha
reader = Reader(rating_scale=(1,5))# Clase usada 'parsear' el dataframe de ratings
data = Dataset.load_from_df(ratings_df,reader)# cargar el dataset desde dataframe y el reader
train_set = data.build_full_trainset() # tomar un conjunto datos de entrenamiento
algo = KNNBasic()# uso de algoritmo
algo.fit(train_set)# ajuste del algoritmo con conjunto de datos de entrenamiento
# seleccion de variables para la prediccion: usuario, pelicula
uid = 196
iid = 302
pre = algo.predict(uid, iid, r_ui=4, verbose=True)
| 3. KNNBasic.ipynb |