code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Multiple linear regression
# ## Import the relevant libraries
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
sns.set()
from sklearn.linear_model import LinearRegression
# -
# ## Load the data
data = pd.read_csv('1.02. Multiple linear regression.csv')
data.head()
data.describe()
# ## Create the multiple linear regression
# ### Declare the dependent and independent variables
x = data[['SAT','Rand 1,2,3']]
y = data['GPA']
# ### Regression itself
reg = LinearRegression()
reg.fit(x,y)
reg.coef_
reg.intercept_
| 15 - Advanced Statistical Methods in Python/3_Linear Regression with sklearn/6_Multiple Linear Regression with sklearn (3:10)/sklearn - Multiple Linear Regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from config import get_config
import argparse
from ShapeLearner import ShapeLearner
from ShapeLoader import ShapeDataSet
from torchvision import transforms as trans
from torch.utils.data import Dataset, ConcatDataset, DataLoader, RandomSampler
from torchvision import transforms as trans
from torchvision.datasets import ImageFolder
from PIL import Image, ImageFile
import numpy as np
import cv2
import pickle
import torch
import sys
from torch.autograd import Variable
import torch.nn.functional as F
import matplotlib.pyplot as plt
import PIL
# %matplotlib inline
from pathlib import Path
from torch import nn
from torchvision import transforms as trans
from os import path
from pathlib import Path
import os
from itertools import product
import re
from tqdm.notebook import tqdm as tqdm_notebook
import math
conf = get_config(2)
conf.device = 'cpu'
conf.net_mode = 'resnet18'
conf.n_shapes = 1
conf.n_colors = 3
conf.shape_only = False
conf.color_only = False
learner = ShapeLearner(conf, inference=True)
from itertools import product
triangle_ds = ShapeDataSet(no_bkg=True)
triangle_ds.shapes = ['triangle']
triangle_ds.colors = [[(255, 255), (0, 0), (0, 0)],
[(0, 0), (255, 255), (0, 0)],
[(0, 0), (0, 0), (255, 255)]]
triangle_ds.n_shapes = 1
triangle_ds.n_colors = 3
ziped_classes = enumerate(product(range(1), range(3)))
triangle_ds.label_map = {v:-1 for k, v in ziped_classes}
triangle_ds.label_names = [-1]
learner.ds = triangle_ds
# ## generate OOD shape/color
# +
def set_distractors(learner):
# set OOD data
triangle_ds = ShapeDataSet(no_bkg=True)
triangle_ds.shapes = ['triangle']
triangle_ds.colors = [[(255, 255), (0, 0), (0, 0)],
[(0, 0), (255, 255), (0, 0)],
[(0, 0), (0, 0), (255, 255)]]
triangle_ds.n_shapes = 1
triangle_ds.n_colors = 3
ziped_classes = enumerate(product(range(1), range(3)))
triangle_ds.label_map = {v:-1 for k, v in ziped_classes}
triangle_ds.label_names = [-1]
learner.ds = triangle_ds
dloader_args = {
'batch_size': conf.batch_size,
'pin_memory': True,
'num_workers': conf.num_workers,
'drop_last': False,
}
learner.loader = DataLoader(learner.ds, **dloader_args)
eval_sampler = RandomSampler(learner.ds, replacement=True, num_samples=len(learner.ds) // 10)
learner.eval_loader = DataLoader(learner.ds, sampler=eval_sampler, **dloader_args)
def set_probes(learner):
# set OOD data
triangle_ds = ShapeDataSet(no_bkg=True)
triangle_ds.shapes = ['rectangle', 'circle']
triangle_ds.colors = [[(255, 255), (0, 0), (0, 0)],
[(0, 0), (255, 255), (0, 0)],
]#[(0, 0), (0, 0), (255, 255)]]
triangle_ds.n_shapes = 2
triangle_ds.n_colors = 2 # TODO fix this! we need the right 2 in the right order!
ziped_classes = enumerate(product(range(triangle_ds.n_shapes), range(triangle_ds.n_colors)))
triangle_ds.label_map = {v:k for k, v in ziped_classes}
triangle_ds.label_names = [str(x) for x in product(triangle_ds.shapes, range(triangle_ds.n_colors))]
learner.ds = triangle_ds
dloader_args = {
'batch_size': conf.batch_size,
'pin_memory': True,
'num_workers': conf.num_workers,
'drop_last': False,
}
learner.loader = DataLoader(learner.ds, **dloader_args)
eval_sampler = RandomSampler(learner.ds, replacement=True, num_samples=len(learner.ds) // 10)
learner.eval_loader = DataLoader(learner.ds, sampler=eval_sampler, **dloader_args)
def get_evaluation(learner):
# evaluate OOD data
for i in range(len(learner.models)):
learner.models[i].eval()
do_mean = -1 if len(learner.models) > 1 else 0
ind_iter = range(do_mean, len(learner.models))
predictions = dict(zip(ind_iter, [[] for i in ind_iter]))
prob = dict(zip(ind_iter, [[] for i in ind_iter]))
labels = []
learner.eval_loader.dataset.set_mode('test') # todo check this works :)
for imgs, label in tqdm_notebook(learner.eval_loader, total=len(learner.eval_loader)):
imgs = imgs.to(conf.device)
thetas = [model(imgs).detach() for model in learner.models]
if len(learner.models) > 1: thetas = [torch.mean(torch.stack(thetas), 0)] + thetas
for ind, theta in zip(range(do_mean, len(learner.models)), thetas):
val, arg = torch.max(theta, dim=1)
predictions[ind].append(arg.cpu().numpy())
prob[ind].append(theta.cpu().numpy())
labels.append(label.detach().cpu().numpy())
labels = np.hstack(labels)
for ind in range(do_mean, len(learner.models)):
predictions[ind] = np.hstack(predictions[ind])
prob[ind] = np.vstack(prob[ind])
return prob, predictions, labels
# -
# ## Load Models
dloader_args = {
'batch_size': conf.batch_size,
'pin_memory': True,
'num_workers': conf.num_workers,
'drop_last': False,
}
learner.loader = DataLoader(learner.ds, **dloader_args)
eval_sampler = RandomSampler(learner.ds, replacement=True, num_samples=len(learner.ds) // 10)
learner.eval_loader = DataLoader(learner.ds, sampler=eval_sampler, **dloader_args)
MODEL_DIR = 'work_space/save/shapes_pre_v2'
rel_dirs = os.listdir(MODEL_DIR)
alpha = [re.findall('a=([0-9, \.]*)_', d)[0] for d in rel_dirs]
fix_str = [x for x in os.listdir(path.join(MODEL_DIR, rel_dirs[0])) if 'model' in x][0][8:]
model_path = rel_dirs[0]
conf.save_path = Path(path.join(MODEL_DIR, model_path))
learner.load_state(conf, fix_str, model_only=True, from_save_folder=True)
# ## evaluate OOD samples
def get_TTR_FTR_curve():
open_set_1st_labels_0 = np.argmax(distractors_prob[0], 1)
open_set_1st_labels_1 = np.argmax(distractors_prob[1], 1)
open_set_1st_scores_0 = np.max(distractors_prob[0], 1)
open_set_1st_scores_1 = np.max(distractors_prob[1], 1)
mean_pred = np.argmax(prob_prob[-1], 1)
mean_score = np.max(prob_prob[-1], 1)
a = np.sum((open_set_1st_labels_0 == open_set_1st_labels_1))
b = len(prob_labels)
corr = (100.0*a / b)
prev_FTR = -1
prev_TTR = -1
THs = []
TTRs = []
FTRs = []
for i, TH in enumerate(np.arange(0, 1, 0.00001)):
FTR = np.sum((open_set_1st_labels_0 == open_set_1st_labels_1) & (open_set_1st_scores_0 > TH) & (open_set_1st_scores_1 > TH)) / b
TTR = np.sum((mean_score > TH) & (mean_pred == prob_labels)) / b
if (prev_FTR != FTR and prev_TTR != TTR) or (i%100 == 0):
prev_FTR = FTR
prev_TTR = TTR
THs.append(TH)
TTRs.append(TTR)
FTRs.append(FTR)
return THs, TTRs, FTRs, corr
res_dir = dict.fromkeys(alpha)
for model_path, curr_alpha in zip(rel_dirs, alpha):
conf.save_path = Path(path.join(MODEL_DIR, model_path))
fix_str = [x for x in os.listdir(path.join(MODEL_DIR, model_path)) if 'model' in x][0][8:]
learner.load_state(conf, fix_str, model_only=True, from_save_folder=True)
# distractors
set_distractors(learner)
distractors_prob, distractors_predictions, distractors_labels = get_evaluation(learner)
# probs
set_probes(learner)
prob_prob, prob_predictions, prob_labels = get_evaluation(learner)
THs, TTRs, FTRs, corr = get_TTR_FTR_curve()
print(curr_alpha, corr)
res_dir[curr_alpha] = [THs, TTRs, FTRs, corr]
fig, ax = plt.subplots(5, figsize=(7,7))
ax = ax[:]
for i, (curr_alpha, curr_color) in enumerate(zip(sorted(alpha), 'rgbyc')):
THs, TTRs, FTRs, corr = res_dir[curr_alpha]
ax[i].plot(THs, TTRs, label=str(curr_alpha)+': TTR', color=curr_color, linestyle='dotted')
ax[i].plot(THs, FTRs, label=str(curr_alpha)+': FTR', color=curr_color, linestyle='--')
ax[i].legend(loc='center left', bbox_to_anchor=(-.4, 0.5))
ax[i].set_ylim([-.1,1.1])
ax[i].set_ylabel('corr: '+ str(corr))
plt.xlabel('Thresholds')
plt.tight_layout()
import torchvision.models as models
import eagerpy as ep
from foolbox import PyTorchModel, accuracy, samples
from foolbox.attacks import LinfPGD
# instantiate a model
model = learner.models[0]
#preprocessing = dict(mean=[0.485, 0.456, 0.406], std=[0.229, 0.224, 0.225], axis=-3)
fmodel = PyTorchModel(model, bounds=(0, 1))#, preprocessing=preprocessing)
images, labels = ep.astensors(*samples(fmodel, dataset="cifar10", batchsize=16))
for imgs, labels in learner.eval_loader:
imgs = ep.astensors(imgs)
labels = ep.astensors(labels)
break
print(accuracy(fmodel, images, labels))
# +
# get data and test the model
# wrapping the tensors with ep.astensors is optional, but it allows
# us to work with EagerPy tensors in the following
print(accuracy(fmodel, images, labels))
# apply the attack
attack = LinfPGD()
epsilons = [0.0, 0.001, 0.01, 0.03, 0.1, 0.3, 0.5, 1.0]
advs, _, success = attack(fmodel, images, labels, epsilons=epsilons)
# calculate and report the robust accuracy
robust_accuracy = 1 - success.float32().mean(axis=-1)
for eps, acc in zip(epsilons, robust_accuracy):
print(eps, acc.item())
# we can also manually check this
for eps, advs_ in zip(epsilons, advs):
print(eps, accuracy(fmodel, advs_, labels))
# but then we also need to look at the perturbation sizes
# and check if they are smaller than eps
print((advs_ - images).norms.linf(axis=(1, 2, 3)).numpy())
| .ipynb_checkpoints/evaluate_open_set_shape-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="My1_5AFmSUf5"
# <table align="left">
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/<<<https://github.com 제외한 깃허브 파일 주소 입력>>>"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />구글 코랩에서 실행하기</a>
# </td>
# </table>
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 20812, "status": "ok", "timestamp": 1643350260283, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": -540} id="oa1Z1w-IFEyi" outputId="89161d45-e32a-4a20-8ce7-01b050968bbb"
from google.colab import drive
drive.mount('/content/drive')
# %cd /content/drive
# + id="76IwwD9GJG8k" executionInfo={"status": "ok", "timestamp": 1643350261202, "user_tz": -540, "elapsed": 957, "user": {"displayName": "", "photoUrl": "", "userId": ""}} outputId="62d6c23e-6c16-4b51-fe8a-b7d51ddc31e9" colab={"base_uri": "https://localhost:8080/"}
# %cd /content/drive/MyDrive/
# !git clone https://github.com/just-benedict-it/u2net.git
# + [markdown] id="kRXT145yTwYJ"
# download u2net.pth from https://drive.google.com/file/d/1ao1ovG1Qtx4b7EoskHXmi2E9rp5CHLcZ/view?usp=sharing
#
# + [markdown] id="dPA_aO_QT-JR"
# put u2net.pth to u2net/U-2-Net/saved_models/u2net
# + id="nezfRkLG8TDO" executionInfo={"status": "ok", "timestamp": 1643350567487, "user_tz": -540, "elapsed": 344, "user": {"displayName": "", "photoUrl": "", "userId": ""}}
# 백그라운드를 제거하고 싶은 이미지가 있는 폴더경로.
# input_path = "/content/drive/MyDrive/<write your image folder dir>"
input_path = "/content/drive/MyDrive/images"
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 334, "status": "ok", "timestamp": 1643350603459, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": -540} id="kch3rMF16vWx" outputId="7ffa453a-fa1d-4217-e30d-0b88bf47cebb"
# 이미지들이 있는 폴더경로를 주면 이미지의 이름을 바꾸줌 ex) 0.png
# change images' name
# %cd /content/drive/MyDrive/u2net
# !python rename.py --image_path {input_path}
# + colab={"base_uri": "https://localhost:8080/"} id="-vw_CRJaek4j" outputId="b65c4a37-4b75-4c31-efcb-41121776e8c8" executionInfo={"status": "ok", "timestamp": 1643350831149, "user_tz": -540, "elapsed": 15711, "user": {"displayName": "", "photoUrl": "", "userId": ""}}
# 이미지 경로를 주면 이미지 경로_mask 폴더를 만든 후, 그 폴더에 mask된 이미지를 저장.
# make masked images
# %cd /content/drive/MyDrive/u2net/U-2-Net
# !python u2net_test.py --image_path {input_path}
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 538, "status": "ok", "timestamp": 1643350851431, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": -540} id="kM8LApKCc_da" outputId="3490b8ae-fcb8-43a4-a29c-d320fe775c56"
# 이미지 경로를 주면 이미지 경로_rm_bg 폴더를 만든 후, 그 폴더에 백그라운드 제거된 이미지를 저장.
# make background_removed images
# %cd /content/drive/MyDrive/u2net
# !python mask.py --image_path {input_path}
| U2net.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="Z9L6Iq0ilFi5"
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import math
from sklearn.linear_model import LinearRegression
# + id="2CGw4KMplLrw"
data = pd.read_csv('topdown_data.csv')
# + id="fWyLaTsNlPe3"
Y = data['time'][0:57]
X = data['temp'][0:57]
Z = data['var'][0:57]
#print(Y)
Y = Y.values.reshape(-1, 1)
# -
#n*2^(len(bit(W)))
X_ = Z
X_ = X_.values.reshape(-1, 1)
ones = np.ones((X_.shape[0], 1))
X_ = np.concatenate((ones, X_), axis = 1)
model = LinearRegression(fit_intercept=False)
model.fit(X_, Y)
print(model.coef_)
X_ = Z
f = open("nbit.csv", "w")
X_ =5.33949020e-04*(X_) + -3.32804428e+01
for ii in X_:
f.write(str(ii))
f.write('\n')
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="kG7l9Im9wBqG" outputId="bca743e8-bcaa-4d76-d592-49c7d52ecd00"
#log2(x)
X_ = np.log2(X)
X_ = X_.values.reshape(-1, 1)
ones = np.ones((X_.shape[0], 1))
X_ = np.concatenate((ones, X_), axis = 1)
model = LinearRegression(fit_intercept=False)
model.fit(X_, Y)
print(model.coef_)
# + id="j4HKGECspa9E"
X_ = np.log2(X)
f = open("log.csv", "w")
X_ =62.15103496*(X_) + -841.04006535
for ii in X_:
f.write(str(ii))
f.write('\n')
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="JBeAECvKv9XT" outputId="3b300529-d61b-4bc9-ad0d-6e57fe2e97c2"
#sqrt(x)
X_ = np.sqrt(X)
X_ = X_.values.reshape(-1, 1)
ones = np.ones((X_.shape[0], 1))
X_ = np.concatenate((ones, X_), axis = 1)
model = LinearRegression(fit_intercept=False)
model.fit(X_, Y)
print(model.coef_)
# + id="yRB_Xl8rqx_5"
X_ = np.sqrt(X)
f = open("sqrt.csv", "w")
X_ = 0.73302702*(X_) + -119.31451633
for ii in X_:
f.write(str(ii))
f.write('\n')
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="F4uYAzj4wmQH" outputId="eee747dc-3cdf-4671-b2bc-b15b13344540"
#x
X_ = X
X_ = X_.values.reshape(-1, 1)
ones = np.ones((X_.shape[0], 1))
X_ = np.concatenate((ones, X_), axis = 1)
model = LinearRegression(fit_intercept=False)
model.fit(X_, Y)
print(model.coef_)
# + id="Z-G-MCxLr_R0"
X_ = X
f = open("x.csv", "w")
X_ = 8.26970152e-04*(X_) + -8.39905974e+00
for ii in X_:
f.write(str(ii))
f.write('\n')
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="WfavD0QAwrud" outputId="e7bbcf81-e6c1-4ad8-f79c-0afefc450a1a"
#xlogx
X_ = X * np.log2(X)
X_ = X_.values.reshape(-1, 1)
ones = np.ones((X_.shape[0], 1))
X_ = np.concatenate((ones, X_), axis = 1)
model = LinearRegression(fit_intercept=False)
model.fit(X_, Y)
print(model.coef_)
# + id="lz_t_832sXeW"
X_ = X*np.log2(X)
f = open("xlogx.csv", "w")
X_ = 4.24393127e-05*(X_) + 1.46416058e+00
for ii in X_:
f.write(str(ii))
f.write('\n')
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="eM5Z4rsfvxL-" outputId="ebd8f88f-ec77-4359-a2f8-37aab3207c0d"
#x2
X_ = X * X
X_ = X_.values.reshape(-1, 1)
ones = np.ones((X_.shape[0], 1))
X_ = np.concatenate((ones, X_), axis = 1)
model = LinearRegression(fit_intercept=False)
model.fit(X_, Y)
print(model.coef_)
# + id="46ur7BBssqsN"
X_ = X*X
f = open("x2.csv", "w")
X_ = 1.14152606e-09*(X_) + 6.76555542e+01
for ii in X_:
f.write(str(ii))
f.write('\n')
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="jaTBUwyAmsI9" outputId="0f4a86b2-f2e1-40bc-cbff-5bf21a02cb82"
#x3
X_ = X * X * X
X_ = X_.values.reshape(-1, 1)
ones = np.ones((X_.shape[0], 1))
X_ = np.concatenate((ones, X_), axis = 1)
model = LinearRegression(fit_intercept=False)
model.fit(X_, Y)
print(model.coef_)
# + id="QW6Z9Nbisv93"
X_ = X*X*X
f = open("x3.csv", "w")
X_ = 1.95031195e-15*(X_) + 7.78951264e-33
for ii in X_:
f.write(str(ii))
f.write('\n')
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="djwWU35kncya" outputId="92537f98-6bfd-4afa-d8fc-99f3460bcea1"
#2^n
X_ = 2**X
print(X_)
X_ = X_.values.reshape(-1, 1)
ones = np.ones((X_.shape[0], 1))
X_ = np.concatenate((ones, X_), axis = 1)
model = LinearRegression(fit_intercept=False)
model.fit(X_, Y)
print(model.coef_)
# + id="fc2aIse6s8Jy"
X_ = 2**X
f = open("2x.csv", "w")
X_ = 0. *(X_) + 206.62219641
for ii in X_:
f.write(str(ii))
f.write('\n')
# + id="_HuKWfgpz54r"
from decimal import *
# + colab={"base_uri": "https://localhost:8080/", "height": 54} id="sSrzOPfL_m9f" outputId="da89c052-8bbd-4e6e-9961-88063802dc7b"
#x!
sum = 0
Y = data['time']
f = open("finallll.csv", "w")
for i in range(61):
#tính bình phương độ lệch giữa mỗi giá trị n! và t(n) tương ứng với n
x = math.factorial(X[i]) - Y[i]
x = x*x
sum = sum+x
x = Decimal(x)
if len(str(x)) > 6:
x = '{:.5E}'.format(x)
f.write(x)
f.write('\n')
sum = Decimal(sum//61)
#MSE
if len(str(sum)) > 6:
sum = '{:.5E}'.format(sum)
print(sum)
f.write(str(sum))
# + id="9UReZ-G-fC33"
'''
Cấu hình thử nghiệm gồm:
CPU: I5-9400F
RAM: 16gb
GPU: NVIDIA GeForce GTX 1650
'''
| cs112.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # FFT and Frequency Resolution
#
# *NOTE: This entire notebook is a complete ripoff from the Bitweenie Blog reference below. I claim no credit for the insightful explanations and examples*
#
# Refs
# + [Bitweenie blog article](https://www.bitweenie.com/listings/fft-zero-padding/) by "<NAME>" on FFT and zero-padding
# + (heh heh--think that's a cleverly chosen handle, but if it's a real name, they are definitely in the right business...)
# + [Matplotlib tutorial](https://matplotlib.org/3.3.0/gallery/lines_bars_and_markers/psd_demo.html#sphx-glr-gallery-lines-bars-and-markers-psd-demo-py) on PSD plotting
import numpy as np
import matplotlib.pyplot as plt
from scipy import signal
# ## Zero padding
# +
# Parameters:
n = 1000 # number of samples
f = 1e6 # Hz (first sinusoid)
f2 = 1.05e6 # Hz (second sinusoid)
Fs = 100*f # samples/cycle
# For exponential decay:
half_lives = 10.0 # number of half-lives for exponential decay damping function display
t_half = (n/Fs)*(1/half_lives) # half-life
el = np.log(2)/t_half # decay constant
t = np.linspace(0, (n-1)/Fs, n)
# +
fig, (ax1, ax2) = plt.subplots(ncols=2, figsize=(15,4))
# Damping/Envelope Signal
damping = np.repeat(1, len(t)) # no damping
# damping = np.exp(-el*t) # exponential decay
# damping = (-Fs/n)*t + 1.0 # linear damping
# damping = np.real(np.exp(-1j*(2*np.pi*f/40*t + 0))) # envelope, not really damping
ax1.plot(damping)
ax1.set_title('Damping Signal')
# Sinusoid Signal
#s_t = np.exp(-1j*(2*np.pi*f*t + 0)) # single sinusoid
s_t = np.exp(-1j*2*np.pi*f*t) + np.exp(-1j*2*np.pi*f2*t)
ax2.plot(np.real(s_t))
ax2.set_title('Sinusoid Signal')
plt.show()
# +
#s_t = np.exp(-(el+1j*2*np.pi*f)*t)
damped = damping*s_t
padded = np.concatenate((damped, np.zeros(n, dtype=np.complex128)))
fig, (ax1, ax2) = plt.subplots(ncols=2, figsize=(15,4))
ax1.plot(np.real(damped))
ax1.set_title('Damped Sinusoid')
ax2.plot(np.real(padded))
ax2.set_title('Zero-padded Signal')
plt.show()
# -
# From [Bitweenie Blog](https://www.bitweenie.com/listings/fft-zero-padding/):
#
# There are a few reasons why you might want to zero pad time-domain data. The most common reason is to make a waveform have a power-of-two number of samples. When the time-domain length of a waveform is a power of two, radix-2 FFT algorithms, which are extremely efficient, can be used to speed up processing time. FFT algorithms made for FPGAs also typically only work on lengths of power two.
#
# While it’s often necessary to stick to powers of two in your time-domain waveform length, it’s important to keep in mind how doing that affects the resolution of your frequency-domain output.
# ## Frequency Resolution
#
# There are two aspects of FFT resolution. I’ll call the first one **“waveform frequency resolution”** and the second one **“FFT resolution”**. These are not technical names, but I find them helpful for the sake of this discussion. The two can often be confused because when the signal is not zero padded, the two resolutions are equivalent.
#
# + The **“waveform frequency resolution”** is the minimum spacing between two frequencies that can be resolved.
#
# + The **“FFT resolution”** is the number of points in the spectrum, which is directly proportional to the number points used in the FFT.
#
# It is possible to have extremely fine FFT resolution, yet not be able to resolve two coarsely separated frequencies.
#
# It is also possible to have fine waveform frequency resolution, but have the peak energy of the sinusoid spread throughout the entire spectrum (this is called FFT **spectral leakage**).
#
# ### Waveform Frequency Resolution
#
# The **waveform frequency resolution** is defined by the following equation:
#
# $$
# {\Delta}R_{waveform} = \frac{1}{{\Delta}T}
# $$
#
# where ${\Delta}T$ is the time length of the signal with data. It’s important to note here that you should not include any zero padding in this time! Only consider the actual data samples.
# +
delta_T = n*(1/Fs)
print('Signal Length =', delta_T, 'seconds') # 10 us
delta_R_wf = 1/delta_T
print('Waveform Freq Resolution (Delta R) =', delta_R_wf, 'Hz') # 100 kHz
# -
# It’s important to make the connection here that the discrete time Fourier transform (DTFT) or FFT operates on the data as if it were an infinite sequence with zeros on either side of the waveform. This is why the FFT has the distinctive `sinc` function shape at each frequency bin.
#
# You should recognize the waveform resolution equation $1/T$ is the same as the space between nulls of a `sinc` function.
#
# ### FFT Resolution
#
# The **FFT resolution** is defined by the following equation:
#
# $$
# {\Delta}R_{FFT} = \frac{f_s}{N_{FFT}}
# $$
print('For a sampling frequency of', Fs, 'sps:')
for NFFT in [256, 512, 1024, 2048, 4096]:
print(' - FFT Resolution a', NFFT, 'point FFT is', Fs/NFFT, 'Hz')
# Plot the power spectral density (PSD)
# + *NOTE: I used [this stack overflow post](https://stackoverflow.com/questions/48129222/matplotlib-make-plots-in-functions-and-then-add-each-to-a-single-subplot-figure) to figure out how to use custom plotting functions within subplots*
def psd(data, Fs, ax=None, truncate=None, log=False, **kwargs):
if ax is None:
ax = plt.gca()
else:
ax = ax
if (type(truncate) == tuple) and (len(truncate) == 2):
start = truncate[0]
stop = truncate[1]
elif type(truncate) == int:
start = 0
stop = truncate
# elif (len(truncate) == len(data)) and (truncate.dtype == 'bool'):
else:
start = 0
stop = len(data)
f, Pxx_den = signal.periodogram(data, Fs)
if log:
ax.semilogy(f[start:stop], Pxx_den[start:stop], **kwargs)
else:
ax.plot(f[start:stop], Pxx_den[start:stop], **kwargs)
ax.set_xlabel('frequency [Hz]')
ax.set_ylabel('PSD [V**2/Hz]')
return f, Pxx_den, ax
# +
fig, (ax1, ax2) = plt.subplots(ncols=2, figsize=(14,4))
*_, psd_ax1 = psd(np.real(damped), Fs, ax=ax1)
psd_ax1.set_title("PSD Plot (Linear Scale)")
freqs, psds, psd_log_ax2 = psd(np.real(damped), Fs, ax=ax2, truncate=(5,16), log=True)
psd_log_ax2.set_title("Zoomed in on Peak (Log Scale)")
plt.show()
# -
print(len(psds))
print(max(psds))
# Two distinct peaks are not shown, and the single wide peak has an amplitude of about $6.75{\times}10^{-6} \frac{V^2}{Hz}$ (11.4 dBm ?). Clearly these results don’t give an accurate picture of the spectrum. There is not enough resolution in the frequency domain to see both peaks.
#
# Let’s try to resolve the two peaks in the frequency domain by using a larger FFT, thus adding more points to the spectrum along the frequency axis. Let’s use a 7000-point FFT. This is done by zero padding the time-domain signal with 6000 zeros (60 us). The zero-padded time-domain signal is shown here:
# +
damped = damping*s_t
padded = np.concatenate((damped, np.zeros(6000, dtype=np.complex128)))
fig, (ax1, ax2) = plt.subplots(ncols=2, figsize=(15,4))
ax1.plot(np.real(damped))
ax1.set_title('Damped Sinusoid')
ax2.plot(np.real(padded))
ax2.set_title('Zero-padded Signal')
plt.show()
# +
fig, (ax1, ax2) = plt.subplots(ncols=2, figsize=(15,4))
# plot PSD
freqs, psds, psd_ax1 = psd(np.real(padded), Fs, ax=ax1)
psd_ax1.set_title("PSD Plot (Linear Scale)")
# zoom in on peak to see what is going on
mask = (0.5e6 <= freqs) & (freqs <= 1.5e6)
# mask = (-1.5e6 <= freqs) & (freqs <= -0.5e6) # use if complex signal
start = min(np.nonzero(mask)[0])
stop = max(np.nonzero(mask)[0])
*_, psd_log_ax2 = psd(np.real(padded), Fs, ax=ax2, truncate=(start,stop), log=True, marker='D', markersize=3)
psd_log_ax2.set_title("Zoomed in on Peak (Log Scale)")
plt.show()
# -
# Although we’ve added many more frequency points, we still cannot resolve the two sinuoids; we are also still not getting the expected power.
#
# Taking a closer look at what this plot is telling us, we see that all we have done by adding more FFT points is to more clearly define the underlying `sinc` function arising from the waveform frequency resolution equation. You can see that the `sinc` nulls are spaced at about 0.1 MHz.
#
# Because our two sinusoids are spaced only 0.05 MHz apart, no matter how many FFT points (zero padding) we use, we will never be able to resolve the two sinusoids.
#
# Let’s look at what the resolution equations are telling us. Although the **FFT resolution** is about 14 kHz (more than enough resoution), the **waveform frequency resolution** is only 100 kHz. The spacing between signals is 50 kHz, so we are being limited by the waveform frequency resolution.
#
# To resolve the spectrum properly, we need to increase the amount of time-domain data we are using. Instead of zero padding the signal out to 70 us (7000 points), let’s capture 7000 points of the waveform. The time-domain and frequency domain results are shown here, respectively.
# +
# Parameters:
n = 7000 # number of samples
f = 1e6 # Hz (first sinusoid)
f2 = 1.05e6 # Hz (second sinusoid)
Fs = 100*f # samples/cycle
t = np.linspace(0, (n-1)/Fs, n)
# +
fig, (ax1, ax2) = plt.subplots(ncols=2, figsize=(15,4))
# Damping/Envelope Signal
damping = np.repeat(1, len(t)) # no damping
# damping = np.exp(-el*t) # exponential decay
# damping = (-Fs/n)*t + 1.0 # linear damping
# damping = np.real(np.exp(-1j*(2*np.pi*f/40*t + 0))) # envelope, not really damping
ax1.plot(damping)
ax1.set_title('Damping Signal')
# Sinusoid Signal
s_t = np.real(np.exp(-1j*(2*np.pi*f*t + 0)) + np.exp(-1j*(2*np.pi*f2*t + 0))) # two sinusoids
# Damped Signal
damped = damping*s_t
ax2.plot(np.real(damped))
ax2.set_title('Damped Sinusoid')
plt.show()
# +
delta_T = n*(1/Fs)
#delta_T = n*np.power(Fs, -1)
print('Signal Length =', delta_T, 'seconds')
delta_R_wf = 1/delta_T
#delta_R_wf = np.power(delta_T, -1)
print('Waveform Freq Resolution (Delta R) =', delta_R_wf, 'Hz')
# +
fig, (ax1, ax2) = plt.subplots(ncols=2, figsize=(14,4))
# plot PSD
freqs, psds, psd_ax1 = psd(np.real(damped), Fs, ax=ax1)
psd_ax1.set_title("PSD Plot (Linear Scale)")
# zoom in on peak to see what is going on
mask = (0.5e6 <= freqs) & (freqs <= 1.5e6)
start = min(np.nonzero(mask)[0])
stop = max(np.nonzero(mask)[0])
*_, psd_log_ax2 = psd(np.real(damped), Fs, ax=ax2, truncate=(start,stop), log=True, marker='D', markersize=3)
psd_log_ax2.set_title("Zoomed in on Peak (Log Scale)")
plt.show()
# -
psds[np.argmax(freqs==1e6)]
# TODO: convert V^2/Hz into dBm
# With the expanded time-domain data, the waveform frequency resolution is now about 14 kHz as well. As seen in the power spectrum plot, the two sinusoids are not seen. The 1 MHz signal is clearly represented and [**is at the correct power level of 10 dBm (?)**], but the 1.05 MHz signal is wider and not showing the expected power level of 10 dBm. What gives?
#
# What is happening with the 1.05 MHz signal is that we don’t have an FFT point at 1.05 MHz, so the energy is split between multiple FFT bins.
#
# The spacing between FFT points follows the equation:
#
# $$
# {\Delta}R_{FFT} = \frac{f_s}{N_{FFT}}
# $$
#
# where $N_{FFT}$ is the number of FFT points and $f_s$ is the sampling frequency.
#
# In our example, we’re using a sampling frequency of 100 MHz and a 7000-point FFT. This gives us a spacing between points of 14.28 kHz. The frequency of 1 MHz is a multiple of the spacing, but 1.05 MHz is not. The closest frequencies to 1.05 MHz are 1.043 MHz 1.057 MHz, so the energy is split between the two FFT bins.
#
# To solve this issue, we can choose the FFT size so that both frequencies are single points along the frequency axis. Since we don’t need finer waveform frequency resolution, it’s okay to just zero pad the time-domain data to adjust the FFT point spacing.
padded = np.concatenate((damped, np.repeat(0, 1000)))
len(padded)
# +
fig, (ax1, ax2) = plt.subplots(ncols=2, figsize=(14,4))
# plot PSD
freqs, psds, psd_ax1 = psd(np.real(padded), Fs, ax=ax1)
psd_ax1.set_title("PSD Plot (Linear Scale)")
# zoom in on peak to see what is going on
mask = (0.5e6 <= freqs) & (freqs <= 1.5e6)
start = min(np.nonzero(mask)[0])
stop = max(np.nonzero(mask)[0])
*_, psd_log_ax2 = psd(np.real(padded), Fs, ax=ax2, truncate=(start,stop), log=True, marker='D', markersize=3)
psd_log_ax2.set_title("Zoomed in on Peak (Log Scale)")
plt.show()
# -
# Now both frequencies are resolved [**and at the expected power of 10 dBm (?)**].
#
# For the sake of overkill, you can always add more points to your FFT through zero padding (ensuring that you have the correct waveform resolution) to see the shape of the FFT bins as well. This is shown in the following figure:
padded = np.concatenate((damped, np.repeat(0, 100000)))
len(padded)
# +
fig, (ax1, ax2) = plt.subplots(ncols=2, figsize=(14,4))
# plot PSD
freqs, psds, psd_ax1 = psd(np.real(padded), Fs, ax=ax1)
psd_ax1.set_title("PSD Plot (Linear Scale)")
# zoom in on peak to see what is going on
mask = (0.5e6 <= freqs) & (freqs <= 1.5e6)
start = min(np.nonzero(mask)[0])
stop = max(np.nonzero(mask)[0])
*_, psd_log_ax2 = psd(np.real(padded), Fs, ax=ax2, truncate=(start,stop), log=True, marker='D', markersize=2)
psd_log_ax2.set_title("Zoomed in on Peak (Log Scale)")
plt.show()
# -
# ## Choosing the Right FFT Size
#
# Three considerations should factor into your choice of FFT size, zero padding, and time-domain data length.
#
# ### What **waveform frequency resolution** do you need?
# + How close together are your frequencies of interest?
# + The **waveform frequency resolution** should be smaller than the minimum spacing between frequencies of interest in order for them to be distinguishable
# + ${\Delta}R_{waveform} = \frac{1}{{\Delta}T}$
#
# ### What **FFT resolution** do you need?
# + The **FFT resolution** should at least support the same resolution as your waveform frequency resolution. Additionally, some highly-efficient implementations of the FFT require that the number of FFT points be a power of two.
# + Choose $f_s$ and $N_{FFT}$ so that ${\Delta}R_{FFT} = \frac{f_s}{N_{FFT}}$ is at least as fine as ${\Delta}R_{waveform}$
# + Consider choosing $N_{FFT}$ that is a power of 2 for more efficient computations
#
# ### Does your choice of FFT size allow you to inspect particular **frequencies of interest**?
# + You should ensure that there are enough points in the FFT, or the **FFT has the correct spacing** set, so that your frequencies of interest are not split between multiple FFT points.
#
# One final thought on zero padding the FFT:
#
# If you apply a **windowing function** to your waveform, the windowing function needs to be applied *before zero padding the data*. This ensures that your real waveform data starts and ends at zero, which is the point of most windowing functions.
# ## The "knobs" you can turn
#
# ### You can raise the sampling rate
# + This raises the **maximum component frequency** you can observe (Nyquist frequency), $\frac{1}{2}{\cdot}F_s$
# + But this will *lower* your **FFT resolution** (i.e., each FFT bin will represent a wider frequency bandwidth)
#
# ### You can choose a higher $N_{FFT}$
# + This will *raise* your **FFT resolution**, ${\Delta}R_{FFT} = \frac{f_s}{N_{FFT}}$
# + If you don't raise the sampling rate correspondingly, you will need to **zero-pad** the signal, which has the effect of further defining the underlying shape of the `sinc` function which characterizes the FFT bins, but does not actually give you additional information on the frequency content of the signal.
#
# ### You can examine a **longer length of time** for the signal
# + This will *raise* your **waveform resolution**, ${\Delta}R_{waveform} = \frac{1}{{\Delta}T}$
#
| Frequency Resolution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.6.0
# language: julia
# name: julia-1.6
# ---
using PolyharmonicSplines
using PyPlot
# # Polyharmonic Splines
# Polyharmonic splines are used for function approximation and data interpolation. They are very useful for interpolation of scattered data in many dimensions. A special case ($k=2$) are thin plate splines.
#
# The basis functions of polyharmonic splines are radial basis functions of the form:
#
# $$
# \begin{matrix}
# \phi(r) = \begin{cases}
# r^k \qquad \quad \,\mbox{with } k=1,3,5,\dots, \\
# r^k \ln(r) \quad \mbox{with } k=2,4,6,\dots
# \end{cases} \\[5mm]
# r = ||\mathbf{x} - \mathbf{c}_i||_2
# = \sqrt{ (\mathbf{x} - \mathbf{c}_i)^T \, (\mathbf{x} - \mathbf{c}_i) }
# \end{matrix}
# $$
# ## 1D Splines
# +
x = 2pi*rand(10)
y = sin.(x)
S = [PolyharmonicSpline(k,x,y) for k=1:5]
fig, ax = plt.subplots()
xx = range(0,2pi,length=100)
ax.plot(x, y,"o",label="Data")
ax.plot(xx, sin.(xx),label="True")
for k=1:5
ax.plot(xx, S[k].(xx), label="K = $k")
end
ax.set_xlabel("x")
ax.set_ylabel("sin(x)")
ax.set_xlim(0,2pi)
ax.legend()
# -
# ## 1D Smoothed Splines
# +
yn = y .+ 0.1*randn(10)
S2n = PolyharmonicSpline(2,x,yn,s=1)
fig, ax = plt.subplots()
ax.plot(xx, sin.(xx),label="True")
ax.plot(x, yn,"o",label="Noisy Data")
ax.plot(xx, S2n.(xx),label="K=2, s=1")
ax.set_xlabel("x")
ax.set_ylabel("sin(x)")
ax.set_xlim(0,2pi)
ax.legend()
# -
# ## 2D Splines
# +
x,y = randn(50),randn(50)
z = exp.(-(x.^2 .+ y.^2))
S2 = PolyharmonicSpline(2,[x y],z)
n=100
xgrid = ones(n)*LinRange(-3,3,n)'
ygrid = LinRange(-3,3,n)*ones(n)'
xx = reshape(xgrid,n*n)
yy = reshape(ygrid,n*n)
zz = S2.(xx,yy)
zgrid = reshape(zz,n,n);
plot_surface(xgrid,ygrid,zgrid,alpha=0.5)
scatter3D(x,y,z,color="r")
# -
| examples/PolyharmonicSplines.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# +
# # !pip install oolearning --upgrade
# +
import math
import os
import time
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.ensemble import AdaBoostClassifier, ExtraTreesClassifier, RandomForestClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import make_scorer, roc_auc_score
from sklearn.model_selection import GridSearchCV
from sklearn.model_selection import RepeatedKFold
from sklearn.pipeline import Pipeline, FeatureUnion
from sklearn.preprocessing import StandardScaler, Imputer, MinMaxScaler
from sklearn.decomposition import PCA
from xgboost import XGBClassifier
import oolearning as oo
from helpers import DataFrameSelector, CustomLogTransform, ChooserTransform, CombineAgeHoursTransform, CombineCapitalGainLossTransform
pd.set_option('display.width', 500)
pd.set_option('display.max_colwidth', -1)
pd.set_option('display.max_columns', None)
pd.set_option('display.max_rows', None)
width = 10
plt.rcParams['figure.figsize'] = [width, width/1.333]
# +
working_directory = os.path.join(os.getcwd(), '../')
csv_file = os.path.join(working_directory, 'census.csv')
target_variable = 'income'
positive_class = '>50K'
negative_class = '<=50K'
#target_mapping = {0: 'died', 1: 'lived'} # so we can convert from numeric to categoric
explore = oo.ExploreClassificationDataset.from_csv(csv_file_path=csv_file,
target_variable=target_variable)
# map_numeric_target=target_mapping)
# look at data
explore.dataset.head()
# -
explore.numeric_summary()
explore.categoric_summary()
explore.plot_correlation_heatmap()
# NOTE: since I will be cross-validating transformations e.g. GridSearchCV, it typically won't work to
# one-hot-encode during cross-validation because the holdout fold will tend to have categoric values that
# weren't found in the training folds, and therefore will break during transformation because it will encode
# a value (i.e. add a column) that didn't exist in the training folds.
# So, for this, we need to fit ALL data. Then, below if we have new data e.g. Kaggle, we have to apply
# the same pipeline (i.e. cat_encoding_pipeline.transform()
# TODO: this breaks though if there are any categorical features with missing values in the final test/Kaggle set
one_hot_transformer = oo.DummyEncodeTransformer(encoding=oo.CategoricalEncoding.ONE_HOT)
transformed_data = one_hot_transformer.fit_transform(explore.dataset.drop(columns=target_variable))
transformed_data[target_variable] = explore.dataset[target_variable]
# # Transformations & Transformation Tuning Parameters
#
# define the transformations we want to do, some transformations will have parameters (e.g. base of log tranform (or no transform), type of scaling, whether or not to add column combinations (e.g. age * hours-per-week)
# Below is the pipeline for captail-gain/lost.
#
# We want to tune whether or not we should log transform. We need to do this after imputing but before scaling, so it needs to be it's own pipeline.
cap_gain_loss_pipeline = Pipeline([
('selector', DataFrameSelector(attribute_names=['capital-gain', 'capital-loss'])),
('imputer', Imputer()),
# tune Log trasformation base (or no transformation); update: tuned - chose base e
('custom_transform', CustomLogTransform(base=math.e)),
# tune "net gain" (have to do it after log transform; log of <=0 doesn't exist)
('custom_cap_gain_minus_loss', CombineCapitalGainLossTransform(combine=True)),
# tune MinMax vs StandardScaler; we chose MinMax; update: tuned - chose MinMax
('custom_scaler', ChooserTransform(base_transformer=MinMaxScaler())),
])
# Below is the pipeline for the rest of numeric features:
num_pipeline = Pipeline([
('selector', DataFrameSelector(attribute_names=['age', 'education-num', 'hours-per-week'])),
('imputer', Imputer()),
# tune age * hours-per-week; update: tuned -chose not to include
#('combine_agehours', CombineAgeHoursTransform()),
# tune MinMax vs StandardScaler; update: tuned - chose MinMax
('custom_scaler', ChooserTransform(base_transformer=MinMaxScaler())),
])
# Pipeline that simply gets the categorical/encoded columns from the previous transformation (which used `oo-learning`)
append_categoricals = Pipeline([
('append_cats', DataFrameSelector(attribute_names=one_hot_transformer.encoded_columns)) # already encoded
])
# Below is the pipeline for combining all of the other pipelines
# combine pipelines
transformations_pipeline = FeatureUnion(transformer_list=[
("cap_gain_loss_pipeline", cap_gain_loss_pipeline),
("num_pipeline", num_pipeline),
("cat_pipeline", append_categoricals),
])
# Choose the transformations to tune, below:
model = RandomForestClassifier(
random_state=42,
#oob_score=True,
#criterion="gini",
# max_features="auto",
# n_estimators=10,
# max_depth=None,
# min_samples_split=2,
# min_samples_leaf=1,
# min_weight_fraction_leaf=0.,
# max_leaf_nodes=None,
# min_impurity_decrease=0.,
# min_impurity_split=None,
)
full_pipeline = Pipeline([
('preparation', transformations_pipeline),
#('pca_chooser', ChooserTransform()), # PCA option lost; didn't include
#('feature_selection', TopFeatureSelector(feature_importances, k)),
('model', model)
])
# Tuning strategy according to https://www.analyticsvidhya.com/blog/2016/03/complete-guide-parameter-tuning-xgboost-with-codes-python/
from scipy.stats import randint, uniform, expon
model_param_dict = {
'model__max_features': uniform(.2, .8),
'model__n_estimators': randint(50, 2000),
'model__max_depth': randint(3, 100),
'model__min_samples_split': uniform(0.001, 0.4),
'model__min_samples_leaf': uniform(0.001, 0.4),
}
# actual hyper-parameters/options to tune for transformations.
transformation_parameters = {
#'preparation__cap_gain_loss_pipeline__custom_transform__base': [None, math.e], # Log transform (base e) or not
#'preparation__cap_gain_loss_pipeline__custom_cap_gain_minus_loss__combine': [True, False],
#'preparation__cap_gain_loss_pipeline__custom_scaler__base_transformer': [MinMaxScaler(), StandardScaler()],
#'preparation__num_pipeline__imputer__strategy': ['mean', 'median', 'most_frequent'], # tune strategy
#'preparation__num_pipeline__custom_scaler__base_transformer': [MinMaxScaler(), StandardScaler()],
#'preparation__num_pipeline__combine_agehours__combine': [True, False],
#'pca_chooser__base_transformer': [PCA(n_components=0.95, random_state=42), None], # PCA vs not
}
param_grid = {**transformation_parameters, **model_param_dict}
param_grid
# +
# def binary_roc_auc(y_true, y_score):
# return roc_auc_score(y_true=y_true,
# # binary makes it so it converts the "scores" to predictions
# y_score=[1 if x > 0.5 else 0 for x in y_score])
scorer = make_scorer(roc_auc_score, greater_is_better=True)
# -
y = transformed_data[target_variable].apply(lambda x: 1 if x == positive_class else 0)
transformed_data[target_variable].values[0:10]
y[0:10]
print('Starting....')
time_start = time.time()
from sklearn.model_selection import RandomizedSearchCV
grid_search = RandomizedSearchCV(estimator=full_pipeline,
param_distributions=param_grid,
n_iter=40,
cv=RepeatedKFold(n_splits=5, n_repeats=1),
scoring=scorer,
return_train_score=True,
n_jobs=-1,
verbose=2)
grid_search.fit(transformed_data.drop(columns=target_variable), y)
time_end = time.time()
print('Time: {}m'.format(round((time_end-time_start)/60, 1)))
results_df = pd.concat([pd.DataFrame({'mean_score': grid_search.cv_results_["mean_test_score"],
'st_dev_score': grid_search.cv_results_["std_test_score"]}),
pd.DataFrame(grid_search.cv_results_["params"])],
axis=1)
results_df.sort_values(by=['mean_score'], ascending=False).head(10)
grid_search.best_score_, grid_search.best_params_
rescaled_means = MinMaxScaler(feature_range=(100, 1000)).fit_transform(results_df['mean_score'].values.reshape(-1, 1))
rescaled_means = rescaled_means.flatten() # reshape back to array
#rescaled_means
def compare_two_parameters(x_label, y_label):
x = results_df[x_label]
y = results_df[y_label]
plt.scatter(x,y,c=rescaled_means, s=rescaled_means, alpha=0.5)
plt.xlabel(x_label)
plt.ylabel(y_label)
index_of_best = np.argmax(rescaled_means)
plt.scatter(x[index_of_best], y[index_of_best], marker= 'x', s=200, color='red')
x_label = 'model__max_depth'
y_label = 'model__max_features'
compare_two_parameters(x_label, y_label)
grid_search.best_score_, grid_search.best_params_
x_label = 'model__max_depth'
y_label = 'model__n_estimators'
compare_two_parameters(x_label, y_label)
grid_search.best_score_, grid_search.best_params_
x_label = 'model__max_features'
y_label = 'model__n_estimators'
compare_two_parameters(x_label, y_label)
grid_search.best_score_, grid_search.best_params_
x_label = 'model__max_features'
y_label = 'model__n_estimators'
compare_two_parameters(x_label, y_label)
x_label = 'model__min_samples_leaf'
y_label = 'model__n_estimators'
compare_two_parameters(x_label, y_label)
x_label = 'model__min_samples_leaf'
y_label = 'model__max_features'
compare_two_parameters(x_label, y_label)
grid_search.best_score_, grid_search.best_params_
x_label = 'model__min_samples_split'
y_label = 'model__n_estimators'
compare_two_parameters(x_label, y_label)
x_label = 'model__min_samples_split'
y_label = 'model__max_features'
compare_two_parameters(x_label, y_label)
grid_search.best_score_, grid_search.best_params_
# reference:
#
# ```
# N = 1000
# r = range(0, N)
# x = r
# y = r
# colors = r
# area = r
#
# plt.scatter(x, y, s=area, c=colors, alpha=0.5)
# plt.show()
# ```
# column order is based off of the pipeline and FeatureUnion
# cap_gain_loss_pipeline
# num_pipeline
# cat_pipeline
features = ['capital-gain', 'capital-loss'] + ['Net Capital'] + ['age', 'education-num', 'hours-per-week'] + one_hot_transformer.encoded_columns
importances = grid_search.best_estimator_.steps[1][1].feature_importances_
import pandas as pd
pd.DataFrame({'feature': features, 'importance': importances}).sort_values(by=['importance'], ascending=False).head(20)
| data_scientist_nanodegree/projects/p1_charityml/custom/RandomForest_RandomizedSearch_sklearn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: py36
# language: python
# name: py36
# ---
# # pdf_hunter
#
# Search for and download PDF file links from a webpage.
# ## Installation
#
# This has been tested using Python 3 and Python 2.7.
#
# ```
# pip install pdf_hunter
# ```
# ## Usage
# +
import pdf_hunter
url = "https://github.com/EbookFoundation/free-programming-books/blob/master/free-programming-books.md"
# -
pdf_urls = pdf_hunter.get_pdf_urls(url)
pdf_urls[:10]
# ## We can download a single PDF file from a given url
pdf_url = pdf_urls[0]
pdf_url
file_name = pdf_hunter.get_pdf_name(pdf_url)
file_name
# +
import os
os.path.isfile(file_name)
# +
pdf_hunter.download_file(pdf_url, folder_path=os.getcwd())
os.path.isfile(file_name)
# -
# ## Or download all PDF files from the page
pdf_hunter.download_pdf_files(url, folder_path=os.getcwd())
# ***
| notes/Overview.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="D8EJFDz91oTY" colab_type="code" outputId="3c7e0d35-5ddc-4fcb-e2b2-a1e29439bc01" colab={"base_uri": "https://localhost:8080/", "height": 204}
# !rm -rf /content/*
# !wget http://restbfiles-eu.s3.eu-west-1.amazonaws.com/users/albert/articles_2.csv
# + [markdown] id="cbtbPD5Z2tiH" colab_type="text"
#
# """
# import pandas as pd
#
# filePath = "./articles.csv"
# df = pd.read_csv(filePath,encoding='utf-8',header=None,error_bad_lines=False)
# """
# + id="ICYu3w9hIJkC" colab_type="code" outputId="0f5a0ba8-cad8-4a9b-b2d2-6a97106efcdb" colab={"base_uri": "https://localhost:8080/", "height": 187}
# %cd /content
# !rm -rf gpt-2
# !git clone https://github.com/xlacasa/gpt-2.git
# !mv articles_2.csv gpt-2/
# %cd gpt-2
# + [markdown] id="Qtn1qZPgZLb0" colab_type="text"
# install requirements
# + id="434oOx0bZH6J" colab_type="code" outputId="f9d236c1-52ae-407e-c46d-76a1c6fb652d" colab={"base_uri": "https://localhost:8080/", "height": 785}
# !pip3 install -r requirements.txt
# + colab_type="code" id="A498TySgHYyF" outputId="e51323f5-58b0-4926-b8f3-812751710e98" colab={"base_uri": "https://localhost:8080/", "height": 136}
# !python3 download_model.py 117M
# + id="7oJPQtdLbbeK" colab_type="code" colab={}
# !export PYTHONIOENCODING=UTF-8
# + [markdown] id="0KzSbAvePgsI" colab_type="text"
# fetch checkpoints if you have them saved in google drive
# + id="cA2Wk7yIPmS6" colab_type="code" colab={}
# # !cp -r /content/drive/My\ Drive/checkpoint/ /content/gpt-2/
# + [markdown] id="0p--9zwqQRTc" colab_type="text"
#
# lets get our text to train on, in this case from project gutenberg, A Tale of Two Cities, by <NAME>
# + id="QOCvrs-DHvxa" colab_type="code" colab={}
# # !wget https://www.gutenberg.org/files/98/98-0.txt
# + [markdown] id="yPfJ5b3CQXqr" colab_type="text"
#
# start training
# + id="pEn_ihcGI00T" colab_type="code" colab={}
# # !PYTHONPATH=src ./train.py --dataset /content/gpt-2/dog.txt
# + id="QQxwl2exsnHd" colab_type="code" colab={}
# # !PYTHONPATH=src ./train.py -h
# + id="gl4fEhpK0xqU" colab_type="code" colab={}
# # !ls driveCheckpoints/190405-1
# + id="AYcRIKPXUEFi" colab_type="code" outputId="2ab8f9ef-e8bb-4094-fc57-1ad43f15189f" colab={"base_uri": "https://localhost:8080/", "height": 289}
# !ls
# !rm -rf train
# !mkdir train
# !split -b 24576000 -d --additional-suffix=.txt articles_2.csv train/wikiSmall
# !ls train -lh
# + id="AflW4efWuKMd" colab_type="code" outputId="c01b9797-3905-404b-f144-8b6230911c6d" colab={"base_uri": "https://localhost:8080/", "height": 60659}
# !PYTHONPATH=src ./train.py --dataset /content/gpt-2/train --save_every 1000 --sample_every 500 --batch_size 1 --run_name "HELLYEAH"
# + [markdown] id="vS1RJJDFOPnb" colab_type="text"
# save our checkpoints to start training again later
# + id="0XyhsjhTOaTw" colab_type="code" outputId="551f1349-8880-4299-9ce2-337ad9a9037d" colab={"base_uri": "https://localhost:8080/", "height": 102}
# # !rm /content/gpt-2/checkpoint/190405-1/
# !ls
# + id="V3lqshXQM9pp" colab_type="code" colab={}
# # !cp -r /content/drive/My\ Drive/checkpoint/190405-1 /content/gpt-2/models/117M/
# + id="JretqG1zOXdi" colab_type="code" outputId="4f229022-85f3-4d55-e392-ed1f20266ae3" colab={"base_uri": "https://localhost:8080/", "height": 122}
from google.colab import files
from google.colab import drive
drive.mount('/content/drive')
# !cp -r checkpoint/ /content/drive/My\ Drive/
# + [markdown] id="6D-i7vERWbNS" colab_type="text"
# use your trained model
# + id="Zt3fJihsgwkE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 102} outputId="35111ec0-c245-4ba5-8b67-45e43673b0d1"
# #!rm -rf /content/gpt-2/models/117M/
# !cd checkpoint/
# !ls
# + id="VeETvWvrbKga" colab_type="code" colab={}
# !cp -r /content/gpt-2/checkpoint/190405-2/* /content/gpt-2/models/117M/
# + [markdown] id="GmnSrXqtfRbq" colab_type="text"
# Conditional samples
# + id="utJj-iY4gHwE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 6004} outputId="cf314da9-bfac-4474-d882-52d9882cf540"
# !python3 src/interactive_conditional_samples.py --top_k 100 --temperature=0.85
# + [markdown] id="K8rSqkGxg5OK" colab_type="text"
# Unconditional samples
# + id="LaQUEnRxWc3c" colab_type="code" colab={}
# !python3 src/generate_unconditional_samples.py | tee /tmp/samples
| src/gpt_2/Training/NotebookForTraining.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Jupyter[Lab] Language Server Protocol
#
# This is the documentation for:
#
# - [jupyterlab-lsp](https://pypi.org/project/jupyterlab-lsp/)
# - [jupyter-lsp](https://pypi.org/project/jupyter-lsp/)
#
# Both are managed on [GitHub](https://github.com/jupyter-lsp/jupyterlab-lsp),
# where you can find the
# [issue tracker](https://github.com/jupyter-lsp/jupyterlab-lsp/issues).
# + [markdown] nbsphinx-toctree={"maxdepth": 2}
# ```{toctree}
# ---
# caption: Getting Started
# maxdepth: 2
# ---
# Installation
# Language Servers
# Servers configuration <Configuring>
# ```
# + [markdown] nbsphinx-toctree={"maxdepth": 2}
# ```{toctree}
# ---
# caption: Advanced Usage
# maxdepth: 2
# ---
# Contributing
# Extending
# Releasing
# ```
# + [markdown] nbsphinx-toctree={"maxdepth": 2}
# ```{toctree}
# ---
# caption: Project Information
# maxdepth: 1
# ---
# Changelog <CHANGELOG>
# Roadmap
# Architecture
# GitHub repo <https://github.com/jupyter-lsp/jupyterlab-lsp>
# ```
| docs/index.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python
# language: python
# name: conda-env-python-py
# ---
# <a href="https://cognitiveclass.ai"><img src = "https://ibm.box.com/shared/static/ugcqz6ohbvff804xp84y4kqnvvk3bq1g.png" width = 300, align = "center"></a>
#
# <h1 align=center><font size = 5>Lab: Working with a real world data-set using SQL and Python</font></h1>
# # Introduction
#
# This notebook shows how to work with a real world dataset using SQL and Python. In this lab you will:
# 1. Understand the dataset for Chicago Public School level performance
# 1. Store the dataset in an Db2 database on IBM Cloud instance
# 1. Retrieve metadata about tables and columns and query data from mixed case columns
# 1. Solve example problems to practice your SQL skills including using built-in database functions
# ## Chicago Public Schools - Progress Report Cards (2011-2012)
#
# The city of Chicago released a dataset showing all school level performance data used to create School Report Cards for the 2011-2012 school year. The dataset is available from the Chicago Data Portal: https://data.cityofchicago.org/Education/Chicago-Public-Schools-Progress-Report-Cards-2011-/9xs2-f89t
#
# This dataset includes a large number of metrics. Start by familiarizing yourself with the types of metrics in the database: https://data.cityofchicago.org/api/assets/AAD41A13-BE8A-4E67-B1F5-86E711E09D5F?download=true
#
# __NOTE__: Do not download the dataset directly from City of Chicago portal. Instead download a more database friendly version from the link below.
# Now download a static copy of this database and review some of its contents:
# https://ibm.box.com/shared/static/f9gjvj1gjmxxzycdhplzt01qtz0s7ew7.csv
#
#
# ### Store the dataset in a Table
# In many cases the dataset to be analyzed is available as a .CSV (comma separated values) file, perhaps on the internet. To analyze the data using SQL, it first needs to be stored in the database.
#
# While it is easier to read the dataset into a Pandas dataframe and then PERSIST it into the database as we saw in the previous lab, it results in mapping to default datatypes which may not be optimal for SQL querying. For example a long textual field may map to a CLOB instead of a VARCHAR.
#
# Therefore, __it is highly recommended to manually load the table using the database console LOAD tool, as indicated in Week 2 Lab 1 Part II__. The only difference with that lab is that in Step 5 of the instructions you will need to click on create "(+) New Table" and specify the name of the table you want to create and then click "Next".
#
# ##### Now open the Db2 console, open the LOAD tool, Select / Drag the .CSV file for the CHICAGO PUBLIC SCHOOLS dataset and load the dataset into a new table called __SCHOOLS__.
#
# <a href="https://cognitiveclass.ai"><img src = "https://ibm.box.com/shared/static/uc4xjh1uxcc78ks1i18v668simioz4es.jpg"></a>
# ### Connect to the database
# Let us now load the ipython-sql extension and establish a connection with the database
# %load_ext sql
# # %reload_ext sql
# Enter the connection string for your Db2 on Cloud database instance below
# # %sql ibm_db_sa://my-username:my-password@my-hostname:my-port/my-db-name
# %sql ibm_db_sa://hzv87509:DA7KM007rDA7KM007r!@dashdb-txn-sbox-yp-lon02-01.services.eu-gb.bluemix.net:50000/BLUDB
# ### Query the database system catalog to retrieve table metadata
#
# ##### You can verify that the table creation was successful by retrieving the list of all tables in your schema and checking whether the SCHOOLS table was created
# +
# type in your query to retrieve list of all tables in the database for your db2 schema (username)
# # %sql select TABSCHEMA, TABNAME, CREATE_TIME from SYSCAT.TABLES where TABSCHEMA='HZV87509'
# %sql select * from SYSCAT.TABLES where TABNAME = 'SCHOOLS'
# -
# Double-click __here__ for a hint
#
# <!--
# In Db2 the system catalog table called SYSCAT.TABLES contains the table metadata
# -->
# Double-click __here__ for the solution.
#
# <!-- Solution:
#
# # %sql select TABSCHEMA, TABNAME, CREATE_TIME from SYSCAT.TABLES where TABSCHEMA='YOUR-DB2-USERNAME'
#
# or, you can retrieve list of all tables where the schema name is not one of the system created ones:
#
# # %sql select TABSCHEMA, TABNAME, CREATE_TIME from SYSCAT.TABLES \
# # where TABSCHEMA not in ('SYSIBM', 'SYSCAT', 'SYSSTAT', 'SYSIBMADM', 'SYSTOOLS', 'SYSPUBLIC')
#
# or, just query for a specifc table that you want to verify exists in the database
# # %sql select * from SYSCAT.TABLES where TABNAME = 'SCHOOLS'
#
# -->
# ### Query the database system catalog to retrieve column metadata
#
# ##### The SCHOOLS table contains a large number of columns. How many columns does this table have?
# +
# type in your query to retrieve the number of columns in the SCHOOLS table
# %sql select count(*) from SYSCAT.COLUMNS where TABNAME = 'SCHOOLS'
# -
# Double-click __here__ for a hint
#
# <!--
# In Db2 the system catalog table called SYSCAT.COLUMNS contains the column metadata
# -->
# Double-click __here__ for the solution.
#
# <!-- Solution:
#
# # %sql select count(*) from SYSCAT.COLUMNS where TABNAME = 'SCHOOLS'
#
# -->
# Now retrieve the the list of columns in SCHOOLS table and their column type (datatype) and length.
# +
# type in your query to retrieve all column names in the SCHOOLS table along with their datatypes and length
# # %sql select distinct(NAME), COLTYPE, LENGTH from SYSIBM.SYSCOLUMNS where TBNAME = 'SCHOOLS'
# %sql select COLNAME, TYPENAME, LENGTH from SYSCAT.COLUMNS where TABNAME = 'SCHOOLS'
# -
# Double-click __here__ for the solution.
#
# <!-- Solution:
#
# # %sql select COLNAME, TYPENAME, LENGTH from SYSCAT.COLUMNS where TABNAME = 'SCHOOLS'
#
# or
#
# # %sql select distinct(NAME), COLTYPE, LENGTH from SYSIBM.SYSCOLUMNS where TBNAME = 'SCHOOLS'
#
# -->
# ### Questions
# 1. Is the column name for the "SCHOOL ID" attribute in upper or mixed case?
# 1. What is the name of "Community Area Name" column in your table? Does it have spaces?
# 1. Are there any columns in whose names the spaces and paranthesis (round brackets) have been replaced by the underscore character "_"?
# ## Problems
#
# ### Problem 1
#
# ##### How many Elementary Schools are in the dataset?
# %sql select count(*) from SCHOOLS where "Elementary, Middle, or High School" = 'ES'
# Double-click __here__ for a hint
#
# <!--
# Which column specifies the school type e.g. 'ES', 'MS', 'HS'?
# -->
# Double-click __here__ for another hint
#
# <!--
# Does the column name have mixed case, spaces or other special characters?
# If so, ensure you use double quotes around the "Name of the Column"
# -->
# Double-click __here__ for the solution.
#
# <!-- Solution:
#
# # %sql select count(*) from SCHOOLS where "Elementary, Middle, or High School" = 'ES'
#
# Correct answer: 462
#
# -->
# ### Problem 2
#
# ##### What is the highest Safety Score?
# %sql select max(SAFETY_SCORE) from SCHOOLS
# Double-click __here__ for a hint
#
# <!--
# Use the MAX() function
# -->
# Double-click __here__ for the solution.
#
# <!-- Hint:
#
# # %sql select MAX(Safety_Score) AS MAX_SAFETY_SCORE from SCHOOLS
#
# Correct answer: 99
# -->
#
# ### Problem 3
#
# ##### Which schools have highest Safety Score?
# %sql select NAME_OF_SCHOOL, SAFETY_SCORE from SCHOOLS \
# where SAFETY_SCORE = (select max(SAFETY_SCORE) from SCHOOLS)
# Double-click __here__ for the solution.
#
# <!-- Solution:
# In the previous problem we found out that the highest Safety Score is 99, so we can use that as an input in the where clause:
#
# # %sql select Name_of_School, Safety_Score from SCHOOLS where Safety_Score = 99
#
# or, a better way:
#
# # %sql select Name_of_School, Safety_Score from SCHOOLS where \
# # Safety_Score= (select MAX(Safety_Score) from SCHOOLS)
#
#
# Correct answer: several schools with with Safety Score of 99.
# -->
#
# ### Problem 4
#
# ##### What are the top 10 schools with the highest "Average Student Attendance"?
#
# %sql select Name_of_School, Average_Student_Attendance from SCHOOLS \
# order by Average_Student_Attendance desc nulls last limit 10
# Double-click __here__ for the solution.
#
# <!-- Solution:
#
# # %sql select Name_of_School, Average_Student_Attendance from SCHOOLS \
# # order by Average_Student_Attendance desc nulls last limit 10
#
# -->
# ### Problem 5
#
# ##### Retrieve the list of 5 Schools with the lowest Average Student Attendance sorted in ascending order based on attendance
# %sql select Name_of_School, Average_Student_Attendance from SCHOOLS \
# order by Average_Student_Attendance limit 5;
# Double-click __here__ for the solution.
#
# <!-- Solution:
#
# # %sql SELECT Name_of_School, Average_Student_Attendance \
# # from SCHOOLS \
# # order by Average_Student_Attendance \
# # fetch first 5 rows only
#
# -->
#
# ### Problem 6
#
# ##### Now remove the '%' sign from the above result set for Average Student Attendance column
# %sql SELECT Name_of_School, REPLACE(Average_Student_Attendance, '%', '') from SCHOOLS \
# order by Average_Student_Attendance limit 5;
# Double-click __here__ for a hint
#
# <!--
# Use the REPLACE() function to replace '%' with ''
# See documentation for this function at:
# https://www.ibm.com/support/knowledgecenter/en/SSEPGG_10.5.0/com.ibm.db2.luw.sql.ref.doc/doc/r0000843.html
# -->
# Double-click __here__ for the solution.
#
# <!-- Hint:
#
# # %sql SELECT Name_of_School, REPLACE(Average_Student_Attendance, '%', '') \
# # from SCHOOLS \
# # order by Average_Student_Attendance \
# # fetch first 5 rows only
#
# -->
#
# ### Problem 7
#
# ##### Which Schools have Average Student Attendance lower than 70%?
# %sql SELECT Name_of_School, Average_Student_Attendance \
# from SCHOOLS \
# where CAST ( REPLACE(Average_Student_Attendance, '%', '') AS DOUBLE ) < 70 \
# order by Average_Student_Attendance
# Double-click __here__ for a hint
#
# <!--
# The datatype of the "Average_Student_Attendance" column is varchar.
# So you cannot use it as is in the where clause for a numeric comparison.
# First use the CAST() function to cast it as a DECIMAL or DOUBLE
# e.g. CAST("Column_Name" as DOUBLE)
# or simply: DECIMAL("Column_Name")
# -->
# Double-click __here__ for another hint
#
# <!--
# Don't forget the '%' age sign needs to be removed before casting
# -->
# Double-click __here__ for the solution.
#
# <!-- Solution:
#
# # %sql SELECT Name_of_School, Average_Student_Attendance \
# # from SCHOOLS \
# # where CAST ( REPLACE(Average_Student_Attendance, '%', '') AS DOUBLE ) < 70 \
# # order by Average_Student_Attendance
#
# or,
#
# # %sql SELECT Name_of_School, Average_Student_Attendance \
# # from SCHOOLS \
# # where DECIMAL ( REPLACE(Average_Student_Attendance, '%', '') ) < 70 \
# # order by Average_Student_Attendance
#
# -->
#
# ### Problem 8
#
# ##### Get the total College Enrollment for each Community Area
# %sql select Community_Area_Name, sum(College_Enrollment) AS TOTAL_ENROLLMENT \
# from SCHOOLS \
# group by Community_Area_Name;
# Double-click __here__ for a hint
#
# <!--
# Verify the exact name of the Enrollment column in the database
# Use the SUM() function to add up the Enrollments for each Community Area
# -->
# Double-click __here__ for another hint
#
# <!--
# Don't forget to group by the Community Area
# -->
# Double-click __here__ for the solution.
#
# <!-- Solution:
#
# # %sql select Community_Area_Name, sum(College_Enrollment) AS TOTAL_ENROLLMENT \
# # from SCHOOLS \
# # group by Community_Area_Name
#
# -->
#
# ### Problem 9
#
# ##### Get the 5 Community Areas with the least total College Enrollment sorted in ascending order
# %sql select Community_Area_Name, sum(College_Enrollment) AS TOTAL_ENROLLMENT \
# from SCHOOLS \
# group by Community_Area_Name \
# order by TOTAL_ENROLLMENT limit 5;
# Double-click __here__ for a hint
#
# <!--
# Order the previous query and limit the number of rows you fetch
# -->
# Double-click __here__ for the solution.
#
# <!-- Solution:
#
# # %sql select Community_Area_Name, sum(College_Enrollment) AS TOTAL_ENROLLMENT \
# # from SCHOOLS \
# # group by Community_Area_Name \
# # order by TOTAL_ENROLLMENT asc \
# # fetch first 5 rows only
#
# -->
# ### Problem 10
#
# ##### Get the hardship index for the community area which has College Enrollment of 4638
# %sql select * from chicago_socioeconomic_data
# %sql select hardship_index from chicago_socioeconomic_data CD, schools CPS \
# where CD.ca = CPS.community_area_number and college_enrollment = 4368;
# Double-click __here__ for the solution.
#
# <!-- Solution:
# NOTE: For this solution to work the CHICAGO_SOCIOECONOMIC_DATA table
# as created in the last lab of Week 3 should already exist
#
# # %%sql
# select hardship_index
# from chicago_socioeconomic_data CD, schools CPS
# where CD.ca = CPS.community_area_number
# and college_enrollment = 4368
#
# -->
# ### Problem 11
#
# ##### Get the hardship index for the community area which has the highest value for College Enrollment
# %sql select ca, community_area_name, hardship_index from chicago_socioeconomic_data \
# where ca in \
# ( select community_area_number from schools order by college_enrollment desc limit 1 )
# Double-click __here__ for the solution.
#
# <!-- Solution:
# NOTE: For this solution to work the CHICAGO_SOCIOECONOMIC_DATA table
# as created in the last lab of Week 3 should already exist
#
# # %sql select ca, community_area_name, hardship_index from chicago_socioeconomic_data \
# # where ca in \
# # ( select community_area_number from schools order by college_enrollment desc limit 1 )
#
# -->
# ## Summary
#
# ##### In this lab you learned how to work with a real word dataset using SQL and Python. You learned how to query columns with spaces or special characters in their names and with mixed case names. You also used built in database functions and practiced how to sort, limit, and order result sets, as well as used sub-queries and worked with multiple tables.
# Copyright © 2018 [cognitiveclass.ai](cognitiveclass.ai?utm_source=bducopyrightlink&utm_medium=dswb&utm_campaign=bdu). This notebook and its source code are released under the terms of the [MIT License](https://bigdatauniversity.com/mit-license/).
#
| 5-9 Databases and SQL for Data Science/Week 4/DB0201EN-Week4-1-1-RealDataPractice-v4-py.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 0.0. Imports
# +
import numpy as np
import pandas as pd
import boruta as bt
import scikitplot as skplt
import pickle
import seaborn as sns
from matplotlib import pyplot as plt
from sklearn import preprocessing as pp
from sklearn import linear_model as lm
from sklearn import model_selection as ms
from sklearn import ensemble as en
from sklearn import neighbors as nh
# -
pd.options.mode.chained_assignment = None # default='warn'
# # 0.1. Helper Function
def jupyter_settings():
# %matplotlib inline
# %pylab inline
plt.style.use( 'bmh' )
plt.rcParams['figure.figsize'] = [25, 12]
plt.rcParams['font.size'] = 24
display( HTML( '<style>.container { width:100% !important; }</style>') )
pd.options.display.max_columns = None
pd.options.display.max_rows = None
pd.set_option( 'display.expand_frame_repr', False )
sns.set()
# # 0.2. Load dataset
# Connect database
import pandas as pd
import psycopg2 as pg
# +
#HOST = 'comunidade-ds-postgres.c50pcakiuwi3.us-east-1.rds.amazonaws.com'
#PORT = '5432'
#DATABASE = 'comunidadedsdb'
#USER = 'member'
#PASSWORD ='<PASSWORD>'
#
#conn = pg.connect( host=HOST,
# database=DATABASE,
# port=PORT,
# user=USER,
# password=PASSWORD )
#
#query = """
# SELECT *
# FROM pa004.users u INNER JOIN pa004.vehicle v ON ( u.id = v.id )
# INNER JOIN pa004.insurance i ON ( u.id = i.id )
#"""
#
#df1 = pd.read_sql( query, conn )
# -
df_raw = pd.read_csv( '../data/raw/train.csv' )
df_raw.head()
# # 1.0. Data Description
df1 = df_raw.copy()
# ## 1.1. Rename Columns
# +
cols_new = ['id', 'gender', 'age', 'driving_license', 'region_code', 'previously_insured', 'vehicle_age',
'vehicle_damage', 'annual_premium', 'policy_sales_channel', 'vintage', 'response']
# rename
df1.columns = cols_new
# -
# ## 1.2. Data Dimensions
print( 'Number of Rows: {}'.format( df1.shape[0] ) )
print( 'Number of Cols: {}'.format( df1.shape[1] ) )
# ## 1.3. Data Types
df1.dtypes
# ## 1.4. Check NA
df1.isna().sum()
# ## 1.5. Data Descriptive
num_attributes = df1.select_dtypes( include=['int64', 'float64'] )
cat_attributes = df1.select_dtypes( exclude=['int64', 'float64', 'datetime64[ns]'])
# ### 1.5.1. Numerical Attributes
# +
# Central Tendency - Mean, Median
ct1 = pd.DataFrame( num_attributes.apply( np.mean ) ).T
ct2 = pd.DataFrame( num_attributes.apply( np.median ) ).T
# dispersion - std, min, max, range, skew, kurtosis
d1 = pd.DataFrame( num_attributes.apply( np.std ) ).T
d2 = pd.DataFrame( num_attributes.apply( min ) ).T
d3 = pd.DataFrame( num_attributes.apply( max ) ).T
d4 = pd.DataFrame( num_attributes.apply( lambda x: x.max() - x.min() ) ).T
d5 = pd.DataFrame( num_attributes.apply( lambda x: x.skew() ) ).T
d6 = pd.DataFrame( num_attributes.apply( lambda x: x.kurtosis() ) ).T
# concatenar
m = pd.concat( [d2, d3, d4, ct1, ct2, d1, d5, d6] ).T.reset_index()
m.columns = ['attributes', 'min', 'max', 'range', 'mean', 'median', 'std', 'skew', 'kurtosis']
m
# -
# # 2.0. Feature Engineering
# +
df2 = df1.copy()
# Vehicle Damage Number
df2['vehicle_damage'] = df2['vehicle_damage'].apply( lambda x: 1 if x == 'Yes' else 0 )
# Vehicle Age
df2['vehicle_age'] = df2['vehicle_age'].apply( lambda x: 'over_2_years' if x == '> 2 Years' else 'between_1_2_year' if x == '1-2 Year' else 'below_1_year' )
# -
# # 3.0. Data Filtering
df3 = df2.copy()
# # 4.0. Exploratoria Data Analysis
df4 = df3.copy()
# ## 4.1. Univariate Analysis
# ### 4.1.1. Age
sns.boxplot( x='response', y='age', data=df4 )
# + cell_style="split"
aux00 = df4.loc[df4['response'] == 0, 'age']
sns.histplot( aux00 )
# + cell_style="split"
aux00 = df4.loc[df4['response'] == 1, 'age']
sns.histplot( aux00 )
# -
# ### 4.1.2. Anual Income
sns.boxplot( x='response', y='annual_premium', data=df4 )
df4 = df4[(df4['annual_premium'] > 10000) &(df4['annual_premium'] < 100000)]
# + cell_style="split"
aux00 = df4.loc[df4['response'] == 0, 'annual_premium']
sns.histplot( aux00 );
# + cell_style="split"
aux00 = df4.loc[df4['response'] == 1, 'annual_premium']
sns.histplot( aux00 );
# -
# ### 4.1.3. Driving License
aux = df4[['driving_license', 'response']].groupby( 'response' ).sum().reset_index()
sns.barplot( x='response', y='driving_license', data=aux );
# ### 4.1.4. Region Code
# +
aux0 = df4[['id', 'region_code', 'response']].groupby( ['region_code', 'response'] ).count().reset_index()
#aux0 = aux0[(aux0['id'] > 1000) & (aux0['id'] < 20000)]
sns.scatterplot( x='region_code', y='id', hue='response', data=aux0 )
# -
# ### 4.1.5. Previously Insured
# + cell_style="split"
aux0 = df4[['id', 'previously_insured', 'response']].groupby( ['previously_insured', 'response'] ).count().reset_index()
#aux0 = aux0[(aux0['id'] > 1000) & (aux0['id'] < 20000)]
sns.scatterplot( x='previously_insured', y='id', hue='response', data=aux0 )
# + cell_style="split"
pd.crosstab(df4['previously_insured'], df4['response'] ).apply( lambda x: x / x.sum(), axis=1 )
# -
# ### 4.1.6. Vehicle Age
df4[['id','vehicle_age', 'response']].groupby( ['vehicle_age', 'response'] ).count().reset_index()
# ### 4.1.7. Vehicle Damage
aux = df4[['vehicle_damage', 'response']].groupby( 'response' ).sum().reset_index()
sns.barplot( x='response', y='vehicle_damage', data=aux );
# ### 4.1.8. Policy Sales Channel - Stacked Percentage Bar
aux = df4[['policy_sales_channel', 'response']].groupby( 'policy_sales_channel' ).sum().reset_index()
sns.barplot( x='response', y='policy_sales_channel', data=aux );
# +
aux01 = df4[['policy_sales_channel', 'response']].groupby( 'policy_sales_channel' ).sum().reset_index()
aux02 = df4[['id', 'policy_sales_channel']].groupby( 'policy_sales_channel' ).size().reset_index().rename( columns={0:'total_responses'})
aux = pd.merge( aux01, aux02, how='inner', on='policy_sales_channel' )
aux.head()
# -
# ### 4.1.9. Vintage
# + cell_style="split"
aux = df4.loc[df4['response'] == 0, 'vintage']
sns.histplot( aux )
# + cell_style="split"
aux = df4.loc[df4['response'] == 1, 'vintage']
sns.histplot( aux )
# -
df = pd.pivot_table( index='vintage', columns='response', values='id', data=df4).reset_index()
df.columns = ['vintage', 'no_response', 'yes_response']
df.plot( x='vintage', kind='bar', stacked=True)
# # 5.0. Data Preparation
# +
X = df4.drop( 'response', axis=1 )
y = df4['response'].copy()
x_train, x_validation, y_train, y_validation = ms.train_test_split( X, y, test_size=0.20 )
df5 = pd.concat( [x_train, y_train], axis=1 )
# -
# ## 5.1. Standardization
# +
ss = pp.StandardScaler()
# anual premium - StandarScaler
df5['annual_premium'] = ss.fit_transform( df5[['annual_premium']].values )
pickle.dump( ss, open( '../src/features/annual_premium_scaler.pkl', 'wb' ) )
# -
# ## 5.2. Rescaling
# +
mms_age = pp.MinMaxScaler()
mms_vintage = pp.MinMaxScaler()
# Age - MinMaxScaler
df5['age'] = mms_age.fit_transform( df5[['age']].values )
pickle.dump( mms_age, open( '../src/features/age_scaler.pkl', 'wb' ) )
# Vintage - MinMaxScaler
df5['vintage'] = mms_vintage.fit_transform( df5[['vintage']].values )
pickle.dump( mms_vintage, open( '../src/features/vintage_scaler.pkl', 'wb' ) )
# -
# ## 5.3. Transformation
# ### 5.3.1. Encoding
# +
# gender - One Hot Encoding / Target Encoding
target_encode_gender = df5.groupby( 'gender' )['response'].mean()
df5.loc[:, 'gender'] = df5['gender'].map( target_encode_gender )
pickle.dump( target_encode_gender, open( '../src/features/target_encode_gender_scaler.pkl', 'wb' ) )
# region_code - Target Encoding / Frequency Encoding
target_encode_region_code = df5.groupby( 'region_code' )['response'].mean()
df5.loc[:, 'region_code'] = df5['region_code'].map( target_encode_region_code )
pickle.dump( target_encode_region_code, open( '../src/features/target_encode_region_code_scaler.pkl', 'wb' ) )
# vehicle_age - One Hot Encoding / Frequency Encoding
df5 = pd.get_dummies( df5, prefix='vehicle_age', columns=['vehicle_age'] )
# policy_sales_channel - Target Encoding / Frequency Encoding
fe_policy_sales_channel = df5.groupby( 'policy_sales_channel' ).size() / len( df5 )
df5.loc[:, 'policy_sales_channel'] = df5['policy_sales_channel'].map( fe_policy_sales_channel )
pickle.dump( fe_policy_sales_channel, open( '../src/features/fe_policy_sales_channel_scaler.pkl', 'wb' ) )
# -
# ## 5.4. Validation Preparation
# +
# gender
x_validation.loc[:, 'gender'] = x_validation.loc[:, 'gender'].map( target_encode_gender )
# age
x_validation.loc[:, 'age'] = mms_age.transform( x_validation[['age']].values )
# region_code
x_validation.loc[:, 'region_code'] = x_validation.loc[:, 'region_code'].map( target_encode_region_code )
# vehicle_age
x_validation = pd.get_dummies( x_validation, prefix='vehicle_age', columns=['vehicle_age'] )
# annual_premium
x_validation.loc[:, 'annual_premium'] = ss.transform( x_validation[['annual_premium']].values )
# policy_sales_channel
x_validation.loc[:, 'policy_sales_channel'] = x_validation['policy_sales_channel'].map( fe_policy_sales_channel )
# vintage
x_validation.loc[:, 'vintage'] = mms_vintage.transform( x_validation[['vintage']].values )
# fillna
x_validation = x_validation.fillna( 0 )
# -
# # 6.0. Feature Selection
# ## 6.1. Boruta Algorithm
# +
x_train_n = df5.drop( ['id', 'response'], axis=1 ).values
y_train_n = y_train.values.ravel()
# Define model
et = en.ExtraTreesClassifier( n_jobs=-1 )
# Define boruta
boruta = bt.BorutaPy( et, n_estimators='auto', verbose=2, random_state=42 ).fit( x_train_n, y_train_n )
# +
cols_selected = boruta.support_.tolist()
# best features
x_train_fs = df5.drop( ['id', 'response'], axis=1 )
cols_selected_boruta = x_train_fs.iloc[:, cols_selected].columns.to_list()
# not selected boruta
cols_not_selected_boruta = list( np.setdiff1d( x_train_fs.columns, cols_selected_boruta ) )
# -
# ## 6.2. Feature Importance
# +
# model definition
forest = en.ExtraTreesClassifier( n_estimators=250, random_state=0, n_jobs=-1 )
# data preparation
x_train_n = df5.drop( ['id', 'response'], axis=1 )
y_train_n = y_train.values
forest.fit( x_train_n, y_train_n )
# +
importances = forest.feature_importances_
std = np.std([tree.feature_importances_ for tree in forest.estimators_], axis=0)
indices = np.argsort(importances)[::-1]
# Print the feature ranking
print("Feature ranking:")
df = pd.DataFrame()
for i, j in zip( x_train_n, forest.feature_importances_ ):
aux = pd.DataFrame( {'feature': i, 'importance': j}, index=[0] )
df = pd.concat( [df, aux], axis=0 )
print( df.sort_values( 'importance', ascending=False ) )
# Plot the impurity-based feature importances of the forest
plt.figure()
plt.title("Feature importances")
plt.bar(range(x_train_n.shape[1]), importances[indices], color="r", yerr=std[indices], align="center")
plt.xticks(range(x_train_n.shape[1]), indices)
plt.xlim([-1, x_train_n.shape[1]])
plt.show()
# -
# # 7.0. Machine Learning Modelling
cols_selected = ['annual_premium', 'vintage', 'age', 'region_code', 'vehicle_damage', 'previously_insured',
'policy_sales_channel']
# +
x_train = df5[ cols_selected ]
x_val = x_validation[ cols_selected ]
y_val = y_validation
# -
# ## 7.1. KNN Classifier
# +
# model definition
knn_model = nh.KNeighborsClassifier( n_neighbors=7 )
# model training
knn_model.fit( x_train, y_train )
# model prediction
yhat_knn = knn_model.predict_proba( x_val )
# + cell_style="split"
# Accumulative Gain
skplt.metrics.plot_cumulative_gain( y_val, yhat_knn );
# + cell_style="split"
#Scikitplot library is there to help
skplt.metrics.plot_lift_curve( y_val, yhat_knn );
# -
# ## 7.2. Logistic Regression
# +
# model definition
lr_model = lm.LogisticRegression( random_state=42 )
# model training
lr_model.fit( x_train, y_train )
# model prediction
yhat_lr = lr_model.predict_proba( x_val )
# + cell_style="split"
# Accumulative Gain
skplt.metrics.plot_cumulative_gain( y_val, yhat_lr );
# + cell_style="split"
#Scikitplot library is there to help
skplt.metrics.plot_lift_curve( y_val, yhat_lr );
# -
# ## 7.3. Extra Trees Classifier
#
#
# +
# model definition
et = en.ExtraTreesClassifier( n_estimators=1000, n_jobs=-1, random_state=42 )
# model training
et.fit( x_train, y_train )
# model prediction
yhat_et = et.predict_proba( x_val )
# + cell_style="split"
# Accumulative Gain
skplt.metrics.plot_cumulative_gain( y_val, yhat_et );
# + cell_style="split"
#Scikitplot library is there to help
skplt.metrics.plot_lift_curve( y_val, yhat_et );
# -
# ## 7.4. Random Forest
# +
# model definition
rf = en.RandomForestClassifier( n_estimators=1000, n_jobs=-1, random_state=42 )
# model training
rf.fit( x_train, y_train )
# model prediction
yhat_rf = et.predict_proba( x_val )
# + cell_style="split"
# Accumulative Gain
skplt.metrics.plot_cumulative_gain( y_val, yhat_rf );
# + cell_style="split"
#Scikitplot library is there to help
skplt.metrics.plot_lift_curve( y_val, yhat_rf );
# -
# ## 8.0. Performance Metrics
def precision_at_k( data, k=10 ):
data = data.reset_index( drop=True )
data['ranking'] = data.index + 1
data['precision_at_k'] = data['response'].cumsum() / data['ranking']
return ( data.loc[ k, 'precision_at_k'], data )
def recall_at_k( data, k=15 ):
data = data.reset_index( drop=True )
data['ranking'] = data.index + 1
data['recall_at_k'] = data['response'].cumsum() / data['response'].sum()
return ( data.loc[ k, 'recall_at_k'], data )
# +
# # copy dataframe
df8 = x_validation.copy()
df8['response'] = y_validation.copy()
# propensity score
df8['score'] = yhat_et[:, 1].tolist()
# sort clients by propensity score
df8 = df8.sort_values( 'score', ascending=False )
# compute precision at k
precision_at_20, data = precision_at_k( df8, k=20 )
# compute recall at k
recall_at_15, data = recall_at_k( df8, k=15 )
# +
import numpy as np
from sklearn.metrics import top_k_accuracy_score
y_true = np.array([0, 1, 2, 2])
y_score = np.array([[0.5, 0.2, 0.2], # 0 is in top 2
[0.3, 0.4, 0.2], # 1 is in top 2
[0.2, 0.4, 0.3], # 2 is in top 2
[0.7, 0.2, 0.1]]) # 2 isn't in top 2
top_k_accuracy_score(y_true, y_score, k=2)
# -
y_true = np.array( [1, 0, 1, 1, 0, 1, 0, 0] )
y_score = np.array( [])
# # Cummulative Curve Manually
# +
results = pd.DataFrame()
results['prediction'] = yhat_et[:,1].tolist()
results['real'] = y_val.tolist()
# ordering by prediction
results = results.sort_values( 'prediction', ascending=False )
# Percentage of intereset ( Propensity Score )
results['real_cum'] = results['real'].cumsum()
results['real_cum_perc'] = 100*results['real_cum']/results['real'].sum()
# Percentage of Base ( Clients )
results['base'] = range( 1, len( results ) + 1 )
results['base_cum_perc'] = 100*results['base']/len( results )
# Basline model
results['baseline'] = results['base_cum_perc']
plt.figure( figsize=(12,8))
sns.lineplot( x='base_cum_perc', y='real_cum_perc', data=results )
sns.lineplot( x='base_cum_perc', y='baseline', data=results )
# -
# # Lift Curve Manually
# +
results = pd.DataFrame()
results['prediction'] = yhat_et[:,1].tolist()
results['real'] = y_val.tolist()
# ordering by prediction
results = results.sort_values( 'prediction', ascending=False )
# Percentage of intereset ( Propensity Score )
results['real_cum'] = results['real'].cumsum()
results['real_cum_perc'] = 100*results['real_cum']/results['real'].sum()
# Percentage of Base ( Clients )
results['base'] = range( 1, len( results ) + 1 )
results['base_cum_perc'] = 100*results['base']/len( results )
# Basline model
results['baseline'] = results['base_cum_perc']
# Lift Calculation
results['lift'] = results['real_cum_perc'] / results['base_cum_perc']
plt.figure( figsize=(12,8))
sns.lineplot( x='base_cum_perc', y='lift', data=results )
# -
# # ROI Curve Manually
# +
results = pd.DataFrame()
results['prediction'] = yhat_et[:,1].tolist()
results['real'] = y_val.tolist()
# ordering by prediction
results = results.sort_values( 'prediction', ascending=False )
# Percentage of intereset ( Propensity Score )
results['real_cum'] = results['real'].cumsum()
results['real_cum_perc'] = 100*results['real_cum']/results['real'].sum()
# Percentage of Base ( Clients )
results['base'] = range( 1, len( results ) + 1 )
results['base_cum_perc'] = 100*results['base']/len( results )
# Basline model
results['baseline'] = results['base_cum_perc']
# Lift Calculation
results['lift'] = results['real_cum_perc'] / results['base_cum_perc']
# ROI Curve
#plt.figure( figsize=(12,8))
#sns.lineplot( x='base_cum_perc', y='lift', data=results )
# +
# Compute bucket
results['bucket'] = results['prediction'].apply( lambda x: 0.9 if x >= 0.90 else
0.8 if ( x >= 0.80) & ( x <= 0.90 ) else
0.7 if ( x >= 0.70) & ( x <= 0.80 ) else
0.6 if ( x >= 0.60) & ( x <= 0.70 ) else
0.5 if ( x >= 0.50) & ( x <= 0.60 ) else
0.4 if ( x >= 0.40) & ( x <= 0.50 ) else
0.3 if ( x >= 0.30) & ( x <= 0.40 ) else
0.2 if ( x >= 0.20) & ( x <= 0.30 ) else
0.1 if ( x >= 0.10) & ( x <= 0.20 ) else 0.01 )
# Aggregate clients among buckets
df = results[['prediction','bucket']].groupby( 'bucket' ).agg( {'min', 'count'} ).reset_index()
df.columns = df.columns.droplevel()
df.columns = ['index', 'clients', 'propensity_score']
# Compute revenue and cost
df['gross_revenue'] = 40 * df['clients'] * df['propensity_score']
df['cost'] = 4 * df['clients']
df['base'] = df['clients'].sort_values( ascending=True ).cumsum() / df['clients'].sum()
# revenue
df['revenue'] = df['gross_revenue'] - df['cost']
df = df.sort_values( 'index', ascending=False )
df
# -
plt.figure( figsize=(12,8))
aux = df[df['propensity_score'] >= 0.1]
sns.lineplot( x='base', y='revenue', data=aux )
# # 9.0. Deploy to Production
# Save trained model
#pickle.dump( et, open( '/Users/meigarom.lopes/repos/pa004_health_insurance_cross_sell/health_insurance_cross-sell/src/models/model_health_insurance.pkl', 'wb' ) )
pickle.dump( lr_model, open( '/Users/meigarom.lopes/repos/pa004_health_insurance_cross_sell/health_insurance_cross-sell/src/models/model_linear_regression.pkl', 'wb' ) )
# ## 9.1. Health Insurance Class
# +
import pickle
import numpy as np
import pandas as pd
class HealthInsurance:
def __init__( self ):
self.home_path = '/Users/meigarom.lopes/repos/pa004_health_insurance_cross_sell/health_insurance_cross-sell/'
self.annual_premium_scaler = pickle.load( open( self.home_path + 'src/features/annual_premium_scaler.pkl' ) )
self.age_scaler = pickle.load( open( self.home_path + 'src/features/age_scaler.pkl' ) )
self.vintage_scaler = pickle.load( open( self.home_path + 'src/features/vintage_scaler.pkl' ) )
self.target_encode_gender_scaler = pickle.load( open( self.home_path + 'src/features/target_encode_gender_scaler.pkl' ) )
self.target_encode_region_code_scaler = pickle.load( open( self.home_path + 'src/features/target_encode_region_code_scaler.pkl' ) )
self.fe_policy_sales_channel_scaler = pickle.load( open( self.home_path + 'src/features/fe_policy_sales_channel_scaler.pkl' ) )
def data_cleaning( self, df1 ):
# 1.1. Rename Columns
cols_new = ['id', 'gender', 'age', 'driving_license', 'region_code', 'previously_insured', 'vehicle_age',
'vehicle_damage', 'annual_premium', 'policy_sales_channel', 'vintage', 'response']
# rename
df1.columns = cols_new
return df1
def feature_engineering( self, df2 ):
# 2.0. Feature Engineering
# Vehicle Damage Number
df2['vehicle_damage'] = df2['vehicle_damage'].apply( lambda x: 1 if x == 'Yes' else 0 )
# Vehicle Age
df2['vehicle_age'] = df2['vehicle_age'].apply( lambda x: 'over_2_years' if x == '> 2 Years' else 'between_1_2_year' if x == '1-2 Year' else 'below_1_year' )
return df2
def data_preparation( self, df5 ):
# anual premium - StandarScaler
df5['annual_premium'] = self.annual_premium_scaler.transform( df5[['annual_premium']].values )
# Age - MinMaxScaler
df5['age'] = self.age_scaler.transform( df5[['age']].values )
# Vintage - MinMaxScaler
df5['vintage'] = self.vintage_scaler.transform( df5[['vintage']].values )
# gender - One Hot Encoding / Target Encoding
df5.loc[:, 'gender'] = df5['gender'].map( self.target_encode_gender )
# region_code - Target Encoding / Frequency Encoding
df5.loc[:, 'region_code'] = df5['region_code'].map( self.target_encode_region_code )
# vehicle_age - One Hot Encoding / Frequency Encoding
df5 = pd.get_dummies( df5, prefix='vehicle_age', columns=['vehicle_age'] )
# policy_sales_channel - Target Encoding / Frequency Encoding
df5.loc[:, 'policy_sales_channel'] = df5['policy_sales_channel'].map( self.fe_policy_sales_channel )
# Feature Selection
cols_selected = ['annual_premium', 'vintage', 'age', 'region_code', 'vehicle_damage', 'previously_insured',
'policy_sales_channel']
return df5[ cols_selected ]
def get_prediction( self, model, original_data, test_data ):
# model prediction
pred = model.predict_proba( test_data )
# join prediction into original data
original_data['prediction'] = pred
return original_data.to_json( orient='records', date_format='iso' )
# -
# ## 9.2. API Handler
# +
import pickle
import pandas as pd
from flask import Flask, request, Response
from healthinsurance import HealthInsurance
# loading model
path = '/Users/meigarom.lopes/repos/pa004_health_insurance_cross_sell/health_insurance_cross-sell/'
model = pickle.load( open( path + 'src/models/model_health_insurance.pkl', 'rb' ) )
# initialize API
app = Flask( __name__ )
@app.route( '/heathinsurance/predict', methods=['POST'] )
def health_insurance_predict():
test_json = request.get_json()
if test_json: # there is data
if isinstance( test_json, dict ): # unique example
test_raw = pd.DataFrame( test_json, index=[0] )
else: # multiple example
test_raw = pd.DataFrame( test_json, columns=test_json[0].keys() )
# Instantiate Rossmann class
pipeline = HealthInsurance()
# data cleaning
df1 = pipeline.data_cleaning( test_raw )
# feature engineering
df2 = pipeline.feature_engineering( df1 )
# data preparation
df3 = pipeline.data_preparation( df2 )
# prediction
df_response = pipeline.get_prediction( model, test_raw, df3 )
return df_response
else:
return Response( '{}', status=200, mimetype='application/json' )
if __name__ == '__main__':
app.run( '0.0.0.0', debug=True )
# -
# ## 9.3. API Tester
import requests
# loading test dataset
df_test = x_validation
df_test['response'] = y_validation
df_test = df_test.sample(10)
# convert dataframe to json
data = json.dumps( df_test.to_dict( orient='records' ) )
data
# +
# API Call
#url = 'http://0.0.0.0:5000/predict'
url = 'https://health-insurance-model.herokuapp.com/predict'
header = {'Content-type': 'application/json' }
r = requests.post( url, data=data, headers=header )
print( 'Status Code {}'.format( r.status_code ) )
# -
d1 = pd.DataFrame( r.json(), columns=r.json()[0].keys() )
d1.sort_values( 'score', ascending=False ).head()
| health_insurance_cross-sell/notebooks/look_1.0-mdfl-health-insurance-cross-sell.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Profitable App Profiles for the Apple App Store and Google Play Store
# 2019-08-29
#
# last updated 2019-09-04
#
# New entrants into the mobile application space have a daunting task ahead of them with regards to standing out and being profitable in an extremely crowded marketplace [1][1]. Here I analyze data on apps in the Apple and Google app stores to identify types of apps likely to attract larger user bases, with the specific goal of identifying types of *free* apps that are likely to attract the most users on the Apple App Store and Google Play Store, with number of users as a proxy for potential ad revenue.
#
# Data sources:
#
# https://www.kaggle.com/ramamet4/app-store-apple-data-set-10k-apps/downloads/app-store-apple-data-set-10k-apps.zip/7
#
# https://www.kaggle.com/lava18/google-play-store-apps/downloads/google-play-store-apps.zip/6
#
# [1]: https://www.statista.com/statistics/276623/number-of-apps-available-in-leading-app-stores/ "app totals"
# +
# import datasets - downloaded from Kaggle
from csv import reader
androidfile = open('google-play-store-apps/googleplaystore.csv', encoding='utf8')
androidread = list(reader(androidfile))
android = androidread[1:]
androidheader = androidread[0]
applefile = open('app-store-apple-data-set-10k-apps/AppleStore.csv', encoding='utf8')
appleread = list(reader(applefile))
apple = appleread[1:]
appleheader = appleread[0]
# explore_data() function to easily read in and explore datasets
def explore_data(dataset, start, end, rows_and_columns=False):
dataset_slice = dataset[start:end]
for row in dataset_slice:
print(row)
print('\n') # adds a new (empty) line after each row for easier viewing
if rows_and_columns:
print('Number of rows:', len(dataset))
print('Number of columns:', len(dataset[0]))
explore_data(android, start=0, end=5, rows_and_columns=True)
explore_data(apple, 0, 5, True)
# -
# ## Relevant columns
#
# Based on a quick glance of the datasets, we can choose some more relevant columns for our given task.
#
# For Android apps: Category, Rating (maybe), Reviews, Installs, Type or Price, Genres. We only want to consider free apps (Type=='Free' or Price==0). Category and Genres are related, with Category looking like the first Genre (an app can have multiple genres, but only gets one Category). We'll keep it simple for now and use Category.
#
# For Apple apps: currency (keep only USD apps), price, rating_count_tot, rating_count_ver (total ratings for current version only), prime_genre
# ## Data cleaning
#
# We need to clean the data. Let Dataquest walk us through for now. DQ notes the Google Play data has a [discussion](https://www.kaggle.com/lava18/google-play-store-apps/discussion/66015) identifying an error. We can inspect and remove the offending row. Browsing the Kaggle discussion for the Apple data reveals no identified errors (doesn't mean there aren't any!). There are also duplicate entries that should be removed.
# remove entry 10472 (not including header)
print(android[10472])
#del android[10472] # only run once! Ideally would delete using a unique feature, e.g. app name
# +
### Remove duplicate entries
duplicate_apps = []
unique_apps = []
for app in android:
name = app[0]
if name in unique_apps:
duplicate_apps.append(name)
else:
unique_apps.append(name)
print('Number of duplicate apps:', len(duplicate_apps))
print('\n')
print('Examples of duplicate apps:', duplicate_apps[:15])
apple_duplicate_apps = []
apple_unique_apps = []
for app in apple:
name = app[2]
if name in apple_unique_apps:
apple_duplicate_apps.append(name)
else:
apple_unique_apps.append(name)
print('Number of duplicate apps:', len(apple_duplicate_apps))
print('\n')
print('Examples of duplicate apps:', apple_duplicate_apps[:15])
# -
# The Android data has 1181 duplicate entries. We could keep one at random, or we could keep the most recent entry, using the entry with the highest number of reviews as the most recent. (We can reasonably expect number of reviews to increase with time, so more reviews = more recent entry)
# +
# to remove duplicates, create a dictionary, where key == unique app name and value == highest number of reviews of that app
# use 'not in' operator to check for membership in a dictionary
# e.g. print('z' not in ['a', 'b', 'c'])
reviews_max = {} # empty dictionary
for app in android:
name = app[0]
n_reviews = float(app[3]) # number of reviews is the 3rd index (4th column) in the table
if name in reviews_max and reviews_max[name] < n_reviews:
reviews_max[name] = n_reviews
elif name not in reviews_max:
reviews_max[name] = n_reviews
print(len(reviews_max)) # should have 9659 entries
# -
# Now actually remove duplicate rows, using reviews_max dictionary.
# We have the highest number of reviews for a given app in the `reviews_max` dictionary. When this matches the
# number of reviews in an entry in the full `android` app table, we will keep that row.
# When the loop encounters another entry for the same app, it skips it because its name is already in the `already_added` list.
# We need this `already_added` condition in case duplicate entries have the same number of reviews.
# +
android_clean = [] # store the new cleaned dataset with all columns
already_added = [] # just store app names
for app in android:
name = app[0]
n_reviews = float(app[3])
if n_reviews == reviews_max[name] and name not in already_added:
android_clean.append(app) # appends the whole row
already_added.append(name) # appends just the name to keep track of which apps have been added
print(len(android_clean))
explore_data(android_clean, 0, 5, True)
# +
# Repeat for Apple
apple_reviews_max = {} # empty dictionary
for app in apple:
name = app[2] # app name is 2nd index in apple data
n_reviews = float(app[6]) # number of reviews is the 6th index in the apple data
if name in apple_reviews_max and apple_reviews_max[name] < n_reviews:
apple_reviews_max[name] = n_reviews
elif name not in apple_reviews_max:
apple_reviews_max[name] = n_reviews
print(len(apple_reviews_max))
apple_clean = [] # store the new cleaned dataset with all columns
apple_already_added = [] # just store app names
for app in apple:
name = app[2]
n_reviews = float(app[6])
if n_reviews == apple_reviews_max[name] and name not in apple_already_added:
apple_clean.append(app) # appends the whole row
apple_already_added.append(name) # appends just the name to keep track of which apps have been added
print(len(apple_clean))
explore_data(apple_clean, 0, 5, True)
# -
# We have more filtering we'd like to do. We are only interested in apps oriented toward English speakers, so we can check that characters in app names are Western characters.
#
#
# +
# function that returns False if any characters in a given string are not common English characters
def isenglish(string):
for char in string:
if ord(char) > 127:
return False
return True
# test it out
print(isenglish('Instagram'))
print(isenglish('爱奇艺PPS -《欢乐颂2》电视剧热播'))
print(isenglish('Docs To Go™ Free Office Suite'))
print(isenglish('Instachat 😜'))
# -
# Our `isenglish` function is a little rough - unfortunately it identifies names with emojis and other symbols as non-English. We can make it a bit more robust by only identifying app names with more than a few non-standard-English characters as non-English.
# +
# keep count of the number of non-english characters and make conditional statements based on the count
def isenglish(string):
nonengcount = 0
for char in string:
if ord(char) > 127:
nonengcount += 1
if nonengcount >= 3:
return False
return True
# test it out
print(isenglish('Instagram'))
print(isenglish('爱奇艺PPS -《欢乐颂2》电视剧热播'))
print(isenglish('Docs To Go™ Free Office Suite'))
print(isenglish('Instachat 😜'))
# +
# filter out apps from both android and apple datasets:
android_clean2 = []
for app in android_clean:
name = app[0]
if isenglish(name) == True:
android_clean2.append(app)
print(len(android), 'Android apps total')
print(len(android_clean), 'Android apps after filtering for duplicates')
print(len(android_clean2), 'Android apps after filtering for English')
explore_data(android_clean2, 0, 5, False)
# +
apple_clean2 = []
for app in apple_clean:
name = app[2]
if isenglish(name) == True:
apple_clean2.append(app)
print(len(apple), 'Apple apps total')
print(len(apple_clean), 'Apple apps after filtering for duplicates')
print(len(apple_clean2), 'Apple apps after filtering for English')
explore_data(apple_clean2, 0, 5, True)
# -
apple[0:5]
print(appleheader)
print(apple[:5], '\n')
# ## Isolate free apps
# +
# android
android_free = []
for app in android_clean2:
if app[6] == 'Free':
android_free.append(app)
print(len(android_clean2), 'filtered Android apps')
print(len(android_free), 'free filtered Android apps')
# apple
apple_free = []
for app in apple_clean2:
if float(app[5]) == 0:
apple_free.append(app)
print(len(apple_clean2), 'filtered Apple apps')
print(len(apple_free), 'free filtered Apple apps')
# -
# ## Analyzing our filtered data
# Now that we have filtered our datasets and isolated the free apps, we can come back to our main goal: identifying genres of apps that are the most popular to guide us in our choice of genre for developing our own app, in order to maximize ad revenue. With our filtered datasets, we can identify the columns we care about for addressing this and producing frequency tables from them.
# +
# print the column names again
print(appleheader) # let's use prime_genre
print(androidheader) # let's use Category
# make function to generate a frequency table of apps by column (in this case, genre or category)
def freq_table(dataset, index): # dataset is a list of lists, index is an integer
table = {} # dictionary
total = 0
for row in dataset:
total += 1 # counts all rows
value = row[index]
if value in table:
table[value] += 1 # a category, e.g. Games, would become the key, and the count of apps in the Games category is the value for that key in the dictionary
else:
table[value] = 1
table_percentages = {}
for key in table:
percentage = (table[key] / total) * 100
table_percentages[key] = percentage
return table_percentages
# we can sort our tables using built-in sorted() function, and some duct tape. Much easier ways to do this later...
def display_table(dataset, index):
table = freq_table(dataset, index)
table_display = []
for key in table:
key_val_as_tuple = (table[key], key)
table_display.append(key_val_as_tuple)
table_sorted = sorted(table_display, reverse = True)
for entry in table_sorted:
print(entry[1], ':', entry[0])
# -
display_table(apple_free, -5) # index for prime_genre is 12 or -5
# In English apps in the Apple App Store, 58% are games. Gaming/entertainment apps are the most common, while more practical apps are less common.
#
# This doesn't necessarily mean gaming and entertainment apps are the best business opportunity though - we want to examine how much they're used. It's possible that games are just easier to make and publish, but may not be the best proxy for user engagement for ad revenue.
#
# The pattern differs a bit for Android, with a more even distribution across categories, but still weighted toward entertainment (family/games).
display_table(android_free, 1)
# What else can we analyze as proxies of popularity or user engagement? We can look at the total number of downloads or number of ratings.
#
# We can use a nested loop to generate frequency tables of downloads by category.
# +
import statistics # enables median()
prime_freqtable = freq_table(apple_free, -5)
#print(prime_freqtable)
lol = [] # create empty list to store the genre and average number of ratings
for genre in prime_freqtable:
appratings = []
total = 0 # store total number of user ratings
len_genre = 0 # store number of apps in each genre
for app in apple_free:
genre_app = app[-5]
if genre_app == genre: # if the genre is the same, save the total number of reviews to "total"
total += float(app[6])
appratings.append(float(app[6])) # create a list of all the rating totals for a given genre of apps
len_genre += 1
medianrating = statistics.median(appratings) # generates the median from the list of rating totals
avgratings = total / len_genre
lol.append([genre, avgratings, medianrating]) # by storing this as a list of lists we can sort the output
#print([genre, avgratings])
print('Sorted by mean:', sorted(lol, key=lambda x: float(x[1]), reverse=True), '\n') # can use a lambda function for the key to sort by the second element in each row (here, by the number of ratings)
print('Sorted by median:', sorted(lol, key=lambda x: float(x[2]), reverse=True), '\n') # can use a lambda function for the key to sort by the second element in each row (here, by the number of ratings)
# -
# For free English Apple Store apps, Navigation apps have the highest average number of reviews. Average is a horrible metric here for determining average downloads/engagement/ratings - this is almost certainly skewed by Google Maps and Waze, with very few other competitors (meaning navigation apps are dominated by a few big players). Meanwhile, games have tons of newer entrants that will bring the average way down.
#
# If we sort by mean, we find Navigation, Reference, and Social Networking make up the top 3 most reviewed app categories.
#
# If we sort by median, we find Productivity, Reference, and Navigation make up the top 3 most reviewed app categories. Medians differ greatly from the means, implying highly skewed distributions (likely bimodal or multimodal) - median should give us a slightly better idea of how a "typical" app in a given genre performs. For planning a new app, even better would be to look at the histogram of review counts and remove the highest reviewed apps from the bimodal distribution and take the median in the absence of those outliers.
#
#
# Now for Android:
# +
import statistics # enables statistics.median()
android_freqtable = freq_table(android_free, 1)
#print(prime_freqtable)
lol = [] # create empty list to store the genre and average number of ratings
for genre in android_freqtable:
appratings = []
total = 0 # store total number of user ratings
len_genre = 0 # store number of apps in each genre
for app in android_free:
genre_app = app[1]
if genre_app == genre: # if the genre is the same, save the total number of reviews to "total"
total += float(app[3])
appratings.append(float(app[3])) # create a list of all the rating totals for a given genre of apps
len_genre += 1
medianrating = statistics.median(appratings) # generates the median from the list of rating totals
avgratings = total / len_genre
lol.append([genre, avgratings, medianrating]) # by storing this as a list of lists we can sort the output
#print([genre, avgratings])
print('Sorted by mean:', sorted(lol, key=lambda x: float(x[1]), reverse=True), '\n') # can use a lambda function for the key to sort by the second element in each row (here, by the number of ratings)
print('Sorted by median:', sorted(lol, key=lambda x: float(x[2]), reverse=True), '\n') # can use a lambda function for the key to sort by the second element in each row (here, by the number of ratings)
# -
# For free English Android apps, the top 3 app categories by mean number of reviews are Communication, Social, and Game.
#
# The top 3 app categories by median are: Game, Entertainment, and Photography.
#
# Again, we see very high differences between the mean and median numbers, so the means are being skewed by extreme outliers.
# +
# show some histograms
import numpy as np
import matplotlib.pyplot as plt
import statistics # enables statistics.median()
android_freqtable = freq_table(android_free, 1)
#print(prime_freqtable)
lol = [] # create empty list to store the genre and average number of ratings
for genre in android_freqtable:
appratings = []
total = 0 # store total number of user ratings
len_genre = 0 # store number of apps in each genre
for app in android_free:
genre_app = app[1]
if genre_app == genre: # if the genre is the same, save the total number of reviews to "total"
total += float(app[3])
appratings.append(float(app[3])) # create a list of all the rating totals for a given genre of apps
len_genre += 1
medianrating = statistics.median(appratings) # generates the median from the list of rating totals
plt.hist(appratings)
plt.xlabel('Total number of ratings')
plt.title(genre)
plt.yscale('log', nonposy='clip')
plt.show() # shows a graph for each iteration of the loop
avgratings = total / len_genre
lol.append([genre, avgratings, medianrating]) # by storing this as a list of lists we can sort the output
#print([genre, avgratings])
print('Sorted by mean:', sorted(lol, key=lambda x: float(x[1]), reverse=True), '\n') # can use a lambda function for the key to sort by the second element in each row (here, by the number of ratings)
print('Sorted by median:', sorted(lol, key=lambda x: float(x[2]), reverse=True), '\n') # can use a lambda function for the key to sort by the second element in each row (here, by the number of ratings)
# +
# scrap
print(sorted(apple_free, key=lambda x: float(x[6]), reverse=True)[:20], '\n') # print the first 10 apps, sorted by number of reviews
#
print(sorted(android_free, key=lambda x: float(x[3]), reverse=True)[:20], '\n') # print the first 10 apps, sorted by number of reviews
# -
# ## Apps by number of installs (Android)
#
# The Android data also provides number of installs, but only presents them as broad bin sizes (e.g. 1000+, 100,000+, 1M+). This might not be as useful as number of ratings, but we can try it out. Because the strings in these columns have the plus sign, we need to remove those before we can convert the numbers to floats. This will be inherently inaccurate because the values provided are only a floor for number of installs, but the exercise is good practice.
#
#
# +
categories_android = freq_table(android_free, 1)
for category in categories_android:
total = 0
len_category = 0
for app in android_free:
category_app = app[1]
if category_app == category:
n_installs = app[5]
n_installs = n_installs.replace(',', '')
n_installs = n_installs.replace('+', '')
total += float(n_installs)
len_category += 1
avg_n_installs = total / len_category
print(category, ':', avg_n_installs)
# again highly skewed, median would be better/we should remove outliers, but we already did that for the more precise number of ratings data
# +
# get a sense of the skew
for app in android_free:
if app[1] == 'COMMUNICATION' and (app[5] == '1,000,000,000+'
or app[5] == '500,000,000+'
or app[5] == '100,000,000+'):
print(app[0], ':', app[5])
# -
# # Conclusions
#
# I would be reluctant to draw conclusions from this initial analysis, based on the incredible skew caused by a few outliers in the dataset. At face value, based on median number of reviews by category, if we were looking to maximize ad revenue on the Apple App Store we may want to make an app in the Productivity, Reference, or Navigation genres. For Android, we may want to make an app in the Game, Entertainment, or Photography categories.
# +
# scrap
#print(android[3][6])
#print(appleheader)
#print(androidheader)
#print(apple_clean2[3][5])
#prime_freqtable.values() # can use .values() and .keys() to print those
| Exploring Apple and Google app stores/gp_profitableapps.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# +
import codecademylib
from matplotlib import pyplot as plt
unit_topics = ['Limits', 'Derivatives', 'Integrals', 'Diff Eq', 'Applications']
middle_school_a = [80, 85, 84, 83, 86]
middle_school_b = [73, 78, 77, 82, 86]
def create_x(t, w, n, d):
return [t*x + w*n for x in range(d)]
# Make your chart here
school_a_x = create_x(2, .8, 1, 5)
school_b_x = create_x(2, .8, 2, 5)
middle_x = [ (a + b) / 2.0 for a, b in zip(school_a_x, school_b_x)]
plt.figure(figsize=(10, 8))
ax = plt.subplot()
ax.set_xticks(middle_x)
ax.set_xticklabels(unit_topics)
plt.bar(school_a_x, middle_school_a)
plt.bar(school_b_x, middle_school_b)
plt.title('Test Averages on Different Units')
plt.xlabel('Unit')
plt.ylabel('Test Average')
plt.legend(['Middle School A', 'Middle School B'])
plt.show()
plt.savefig('my_side_by_side.png')
| Side By Side Bars.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] cell_id="8630cbcd-2dfa-43d9-9651-49063a11728b" deepnote_cell_type="markdown" deepnote_to_be_reexecuted=false execution_millis=1 execution_start=1645456539826 source_hash="3c8ecc38" tags=[]
# # Week 10 Video notebooks
# + [markdown] cell_id="00001-e42b6386-c003-4997-a184-d4c9d800db73" deepnote_cell_type="markdown" tags=[]
# ## Adding a title in Altair
#
# A few references in the Altair documentation:
# * [Customizing visualizations](https://altair-viz.github.io/user_guide/customization.html)
# * [Top-level chart configuration](https://altair-viz.github.io/user_guide/configuration.html)
# + cell_id="00002-53093b1a-2a2c-4c1f-ac4e-8e7962cb2b99" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=30 execution_start=1646146477368 source_hash="7aca45b0" tags=[]
import pandas as pd
import altair as alt
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split
from sklearn.neighbors import KNeighborsClassifier
df = pd.read_csv("../data/spotify_dataset.csv", na_values=" ")
df.dropna(inplace=True)
df = df[df["Artist"].isin(["<NAME>", "<NAME>"])]
scaler = StandardScaler()
df[["Tempo","Energy"]] = scaler.fit_transform(df[["Tempo","Energy"]])
X_train, X_test, y_train, y_test = train_test_split(df[["Tempo","Energy"]], df["Artist"], test_size=0.2)
# + cell_id="00003-632b3d52-71cd-4648-a8c4-7a6a6e861d8e" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=8 execution_start=1646146733596 source_hash="63670829" tags=[]
clf = KNeighborsClassifier(n_neighbors=1)
clf.fit(X_train, y_train)
df["pred"] = clf.predict(df[["Tempo","Energy"]])
c = alt.Chart(df).mark_circle().encode(
x="Tempo",
y="Energy",
color=alt.Color("pred", title="Legend title"),
).properties(
title="Here is a title",
width=700,
height=100,
)
c.configure_legend(
strokeColor='gray',
fillColor='#EEEEEE',
padding=10,
cornerRadius=10,
orient='top-right'
)
# + [markdown] cell_id="44f3962f-61f2-4278-a61c-bce60cee0bd6" deepnote_cell_type="markdown" tags=[]
# ## f-strings
# + cell_id="3ab14d8e-8237-4e04-b107-b623d2e796b5" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=64 execution_start=1646147372582 source_hash="a00b6146" tags=[]
k = 20
j = "hello"
clf = KNeighborsClassifier(n_neighbors=k)
clf.fit(X_train, y_train)
df["pred"] = clf.predict(df[["Tempo","Energy"]])
c = alt.Chart(df).mark_circle().encode(
x="Tempo",
y="Energy",
color=alt.Color("pred", title="Predicted Artist"),
).properties(
title=f"n_neighbors = {k}",
width=700,
height=100,
)
c.configure_legend(
strokeColor='gray',
fillColor='#EEEEEE',
padding=10,
cornerRadius=10,
orient='top-right'
)
# + [markdown] cell_id="da6669b9-f25b-422e-9553-eb3e75ef25b7" deepnote_cell_type="markdown" deepnote_to_be_reexecuted=false execution_millis=2 execution_start=1646147415058 source_hash="edf3c2f8" tags=[]
# ## DRY (Don't Repeat Yourself)
# + cell_id="8f49bda7-fcb8-411c-a079-56438d3f4ae0" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=4 execution_start=1646148257165 source_hash="b34020b0" tags=[]
def make_chart(k):
clf = KNeighborsClassifier(n_neighbors=k)
clf.fit(X_train, y_train)
df[f"pred{k}"] = clf.predict(df[["Tempo","Energy"]])
c = alt.Chart(df).mark_circle().encode(
x="Tempo",
y="Energy",
color=alt.Color(f"pred{k}", title="Predicted Artist"),
).properties(
title=f"n_neighbors = {k}",
width=700,
height=100,
)
return c
# + cell_id="08ec3c77-5876-4a6e-a619-6a66aa928556" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=514 execution_start=1646148381025 source_hash="31ce364a" tags=[]
alt.vconcat(*[make_chart(k) for k in range(1,31)])
# + [markdown] cell_id="2009562f-596a-4d53-9ed4-149c2b98e690" deepnote_cell_type="markdown" tags=[]
# ## Alternatives to Deepnote
#
# * [Jupyter notebook and Jupyter lab](https://jupyter.org/)
# * [Google colab](https://colab.research.google.com/)
| _build/html/_sources/Week10/Week10-Video-notebooks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
pip install pandas
pip install requests
pip install PyMySQL
import pandas as pd
import io
import requests
url="https://raw.githubusercontent.com/cs109/2014_data/master/countries.csv"
s=requests.get(url).content
c=pd.read_csv(io.StringIO(s.decode('utf-8')))
print(c)
# +
import pymysql.cursors
# Connect to the database
connection = pymysql.connect(host='db',
user='admin',
port=3306,
password='<PASSWORD>',
database='covid',
cursorclass=pymysql.cursors.DictCursor)
with connection:
# with connection.cursor() as cursor:
# # Create a new record
# sql = "INSERT INTO `users` (`email`, `password`) VALUES (%s, %s)"
# cursor.execute(sql, ('<EMAIL>', 'very-secret'))
# # connection is not autocommit by default. So you must commit to save
# # your changes.
# connection.commit()
with connection.cursor() as cursor:
# Read a single record
sql = "SELECT * FROM `covid_cases_by_region`"
cursor.execute(sql)
result = cursor.fetchone()
print(result)
# -
import pandas as pd
import io
import requests
url="https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/csse_covid_19_time_series/time_series_covid19_confirmed_US.csv"
content_url=requests.get(url).content
confirmed_dataframe=pd.read_csv(io.StringIO(content_url.decode('utf-8')))
print(confirmed_dataframe)
url="https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/csse_covid_19_time_series/time_series_covid19_deaths_US.csv"
content_url=requests.get(url).content
deaths_dataframe=pd.read_csv(io.StringIO(content_url.decode('utf-8')))
column_array = confirmed_dataframe.columns
print(confirmed_dataframe['1/22/20'])
print(len(column_array))
print(column_array[0])
# +
# 0,3,4,5,6 params
# 11-n dates
# -
print(confirmed_dataframe[column_array[0]])
df_column_info = confirmed_dataframe[['UID', 'code3', 'FIPS', 'Admin2', 'Province_State']]
df_column_date = confirmed_dataframe.iloc[: , 11:479]
print(df_column_info)
print(df_column_date)
final_dataframe = pd.DataFrame(columns=['uid', 'code3', 'fips', 'county', 'state', 'date', 'confirmed'])
# for i in range(5):
# df = df.append({'A': i, 'B': i}, ignore_index=True)
df_column_info.shape[0]
df_column_date.shape[1]
print(df_column_date.shape[1]*df_column_info.shape[0])
for i in range(df_column_info.shape[0]):
# print(df_column_info.iloc[i,:][0])
for j in range(df_column_date.shape[1]):
final_dataframe = final_dataframe.append({
'uid':df_column_info.iloc[i,:][0],
'code3':df_column_info.iloc[i,:][1],
'fips':df_column_info.iloc[i,:][2],
'county':df_column_info.iloc[i,:][3],
'state':df_column_info.iloc[i,:][4],
'date':df_column_date.columns[j],
'confirmed':df_column_date.iloc[i,j]
},ignore_index=True)
print(final_dataframe.shape)
for i in range(3000):
for j in range(400):
print(i,j)
new_confirmed_dataframe = pd.melt(confirmed_dataframe,
id_vars =['UID', 'code3', 'FIPS', 'Admin2','Province_State'],
value_vars =['1/31/20', '2/29/20', '3/31/20', '4/30/20', '5/31/20', '6/30/20',
'7/31/20', '8/31/20', '9/30/20', '10/31/20', '11/30/20', '12/31/20',
'1/31/21', '2/28/21', '3/31/21', '4/30/21', '5/7/21'])
print(new_confirmed_dataframe)
new_deaths_dataframe = pd.melt(deaths_dataframe,
id_vars =['UID', 'Population'],
value_vars =['1/31/20', '2/29/20', '3/31/20', '4/30/20', '5/31/20', '6/30/20',
'7/31/20', '8/31/20', '9/30/20', '10/31/20', '11/30/20', '12/31/20',
'1/31/21', '2/28/21', '3/31/21', '4/30/21', '5/7/21'])
print(new_deaths_dataframe)
new_confirmed_dataframe.columns = ['uid', 'code3', 'fips', 'county', 'state', 'date', 'confirmed']
new_deaths_dataframe.columns = ['uid', 'population','date', 'deaths']
print(new_deaths_dataframe)
new_deaths_only_val = new_deaths_dataframe[['population', 'deaths']]
print(new_deaths_only_val)
final_dataframe = new_confirmed_dataframe.join(new_deaths_only_val)
print(final_dataframe)
# +
# import the module
from sqlalchemy import create_engine
# create sqlalchemy engine
engine = create_engine("mysql+pymysql://{user}:{pw}@db/{db}"
.format(user="admin",
pw="<PASSWORD>",
db="covid"))
# -
# connection = pymysql.connect(host='db',
# user='admin',
# port=3306,
# password='<PASSWORD>',
# database='covid',
# cursorclass=pymysql.cursors.DictCursor)
print(engine)
# Insert whole DataFrame into MySQL
final_dataframe.to_sql('covid_dataset', con = engine, if_exists = 'append', chunksize = 1000)
| etl/ETL Covid.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
from matplotlib import style
style.use('fivethirtyeight')
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import datetime as dt
# # Reflect Tables into SQLAlchemy ORM
# Python SQL toolkit and Object Relational Mapper
import sqlalchemy
from sqlalchemy.ext.automap import automap_base
from sqlalchemy.orm import Session
from sqlalchemy import create_engine, func
# create engine to hawaii.sqlite
engine = create_engine("sqlite:///Resources/hawaii.sqlite")
# reflect an existing database into a new model
Base = automap_base()
# reflect the tables
Base.prepare(engine, reflect=True)
# View all of the classes that automap found
Base.classes.keys()
# Save references to each table
measure = Base.classes.measurement
station = Base.classes.station
# Create our session (link) from Python to the DB
session = Session(engine)
# # Exploratory Precipitation Analysis
# Find the most recent date in the data set.
lastdate = session.query(measure.date).order_by(measure.date.desc()).first()
lastdate
# Design a query to retrieve the last 12 months of precipitation data and plot the results.
# Starting from the most recent data point in the database.
last_date = dt.datetime.strptime(lastdate[0], '%Y-%M-%d')
last_date
# Calculate the date one year from the last date in data set.
date2016 = dt.date(last_date.year -1, last_date.month, last_date.day)
date2016
# Perform a query to retrieve the data and precipitation scores
prcpdata = session.query(measure.date, measure.prcp).filter(measure.date >= date2016).all()
prcpdata
# Save the query results as a Pandas DataFrame
prcp = pd.DataFrame(prcpdata, columns=['Date','Precipitation'])
prcp.head()
# Sort the dataframe by date
prcpsorted = prcp.sort_values(["Date"], ascending = True)
prcpsorted.head()
#set the index to the date column
prcpfinal = prcpsorted.set_index("Date")
prcpfinal.head()
# +
# Use Pandas Plotting with Matplotlib to plot the data
x = prcpsorted['Date']
y = prcpsorted['Precipitation']
plt.figure(figsize = (10,6))
plt.bar(x,y)
plt.title(f"Precipitation")
plt.xlabel("Date")
plt.ylabel("Inches")
plt.show()
# -
# Use Pandas to calcualte the summary statistics for the precipitation data
prcpfinal.describe()
# # Exploratory Station Analysis
# Design a query to calculate the total number stations in the dataset
session.query(station).count()
# Design a query to find the most active stations (i.e. what stations have the most rows?)
most_active = (session.query(measure.station, func.count(measure.id)).filter(measure.station == station.station).group_by(measure.station).order_by(func.count(measure.id).desc()).first())
most_active
# List the stations and the counts in descending order.
station_activity = (session.query(measure.station, func.count(measure.id)).filter(measure.station == station.station).group_by(measure.station).order_by(func.count(measure.id).desc()).all())
station_activity
# Using the most active station id from the previous query, calculate the lowest, highest, and average temperature.
m_a = 'USC00519281'
m_a_temps = session.query(func.min(measure.tobs), func.max(measure.tobs), func.avg(measure.tobs)).filter(measure.station == m_a).all()
m_a_temps
# Using the most active station id
# Query the last 12 months of temperature observation data for this station
ma_12month = session.query(measure.date, measure.tobs).filter(measure.station == m_a).filter(func.strftime("%Y-%m-%d", measure.date) >= date2016).all()
ma_12month
#Create Dataframe
ma12_df = pd.DataFrame(ma_12month, columns = ['Date', 'Temperature'])
ma12 = ma12_df.set_index("Date")
ma12.head()
#plot the results as a histogram
plt.hist(ma12, bins=12)
plt.ylabel("Frequency")
plt.xlabel('Temperature')
plt.show()
# # Close session
# Close Session
session.close()
| Surfs Up/climate_api.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (Data Science)
# language: python
# name: python3__SAGEMAKER_INTERNAL__arn:aws:sagemaker:eu-west-1:470317259841:image/datascience-1.0
# ---
# # Targeting Direct Marketing with Amazon SageMaker XGBoost
# _**Supervised Learning with Gradient Boosted Trees: A Binary Prediction Problem With Unbalanced Classes**_
#
# ---
#
# ## Background
# Direct marketing, either through mail, email, phone, etc., is a common tactic to acquire customers. Because resources and a customer's attention is limited, the goal is to only target the subset of prospects who are likely to engage with a specific offer. Predicting those potential customers based on readily available information like demographics, past interactions, and environmental factors is a common machine learning problem.
#
# This notebook presents an example problem to predict if a customer will enroll for a term deposit at a bank, after one or more phone calls. The steps include:
#
# * Preparing your Amazon SageMaker notebook
# * Downloading data from the internet into Amazon SageMaker
# * Investigating and transforming the data so that it can be fed to Amazon SageMaker algorithms
# * Estimating a model using the Gradient Boosting algorithm
# * Evaluating the effectiveness of the model
# * Setting the model up to make on-going predictions
#
# ---
#
# ## Preparation
#
# _This notebook was created and tested on an ml.m4.xlarge notebook instance._
#
# Let's start by specifying:
#
# - The S3 bucket and prefix that you want to use for training and model data. This should be within the same region as the Notebook Instance, training, and hosting.
# - The IAM role arn used to give training and hosting access to your data. See the documentation for how to create these. Note, if more than one role is required for notebook instances, training, and/or hosting, please replace the boto regexp with a the appropriate full IAM role arn string(s).
# + isConfigCell=true
import sagemaker
bucket=sagemaker.Session().default_bucket()
prefix = 'sagemaker/DEMO-xgboost-dm'
# Define IAM role
import boto3
import re
from sagemaker import get_execution_role
role = get_execution_role()
# -
# Now let's bring in the Python libraries that we'll use throughout the analysis
import numpy as np # For matrix operations and numerical processing
import pandas as pd # For munging tabular data
import matplotlib.pyplot as plt # For charts and visualizations
from IPython.display import Image # For displaying images in the notebook
from IPython.display import display # For displaying outputs in the notebook
from time import gmtime, strftime # For labeling SageMaker models, endpoints, etc.
import sys # For writing outputs to notebook
import math # For ceiling function
import json # For parsing hosting outputs
import os # For manipulating filepath names
import sagemaker
import zipfile # Amazon SageMaker's Python SDK provides many helper functions
# ---
#
# ## Data
# Let's start by downloading the [direct marketing dataset](https://sagemaker-sample-data-us-west-2.s3-us-west-2.amazonaws.com/autopilot/direct_marketing/bank-additional.zip) from the sample data s3 bucket.
#
# \[Moro et al., 2014\] <NAME>, <NAME> and <NAME>. A Data-Driven Approach to Predict the Success of Bank Telemarketing. Decision Support Systems, Elsevier, 62:22-31, June 2014
#
# +
# !wget https://sagemaker-sample-data-us-west-2.s3-us-west-2.amazonaws.com/autopilot/direct_marketing/bank-additional.zip
with zipfile.ZipFile('bank-additional.zip', 'r') as zip_ref:
zip_ref.extractall('.')
# -
# Now lets read this into a Pandas data frame and take a look.
data = pd.read_csv('./bank-additional/bank-additional-full.csv')
pd.set_option('display.max_columns', 500) # Make sure we can see all of the columns
pd.set_option('display.max_rows', 20) # Keep the output on one page
data
# We will store this natively in S3 to then process it with SageMaker Processing.
# +
from sagemaker import Session
sess = Session()
input_source = sess.upload_data('./bank-additional/bank-additional-full.csv', bucket=bucket, key_prefix=f'{prefix}/input_data')
input_source
# -
# # Feature Engineering with Amazon SageMaker Processing
#
# Amazon SageMaker Processing allows you to run steps for data pre- or post-processing, feature engineering, data validation, or model evaluation workloads on Amazon SageMaker. Processing jobs accept data from Amazon S3 as input and store data into Amazon S3 as output.
#
# 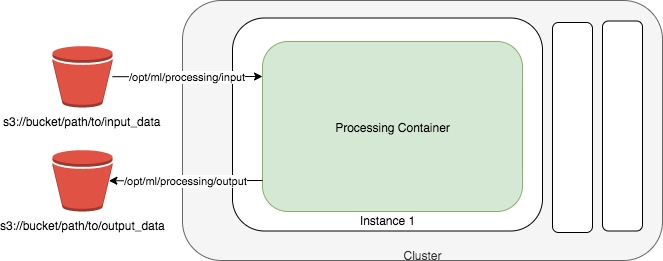
#
# Here, we'll import the dataset and transform it with SageMaker Processing, which can be used to process terabytes of data in a SageMaker-managed cluster separate from the instance running your notebook server. In a typical SageMaker workflow, notebooks are only used for prototyping and can be run on relatively inexpensive and less powerful instances, while processing, training and model hosting tasks are run on separate, more powerful SageMaker-managed instances. SageMaker Processing includes off-the-shelf support for Scikit-learn, as well as a Bring Your Own Container option, so it can be used with many different data transformation technologies and tasks.
#
# To use SageMaker Processing, simply supply a Python data preprocessing script as shown below. For this example, we're using a SageMaker prebuilt Scikit-learn container, which includes many common functions for processing data. There are few limitations on what kinds of code and operations you can run, and only a minimal contract: input and output data must be placed in specified directories. If this is done, SageMaker Processing automatically loads the input data from S3 and uploads transformed data back to S3 when the job is complete.
# +
# %%writefile preprocessing.py
import pandas as pd
import numpy as np
import argparse
import os
from sklearn.preprocessing import OrdinalEncoder
def _parse_args():
parser = argparse.ArgumentParser()
# Data, model, and output directories
# model_dir is always passed in from SageMaker. By default this is a S3 path under the default bucket.
parser.add_argument('--filepath', type=str, default='/opt/ml/processing/input/')
parser.add_argument('--filename', type=str, default='bank-additional-full.csv')
parser.add_argument('--outputpath', type=str, default='/opt/ml/processing/output/')
parser.add_argument('--categorical_features', type=str, default='y, job, marital, education, default, housing, loan, contact, month, day_of_week, poutcome')
return parser.parse_known_args()
if __name__=="__main__":
# Process arguments
args, _ = _parse_args()
# Load data
df = pd.read_csv(os.path.join(args.filepath, args.filename))
# Change the value . into _
df = df.replace(regex=r'\.', value='_')
df = df.replace(regex=r'\_$', value='')
# Add two new indicators
df["no_previous_contact"] = (df["pdays"] == 999).astype(int)
df["not_working"] = df["job"].isin(["student", "retired", "unemployed"]).astype(int)
df = df.drop(['duration', 'emp.var.rate', 'cons.price.idx', 'cons.conf.idx', 'euribor3m', 'nr.employed'], axis=1)
# Encode the categorical features
df = pd.get_dummies(df)
# Train, test, validation split
train_data, validation_data, test_data = np.split(df.sample(frac=1, random_state=42), [int(0.7 * len(df)), int(0.9 * len(df))]) # Randomly sort the data then split out first 70%, second 20%, and last 10%
# Local store
pd.concat([train_data['y_yes'], train_data.drop(['y_yes','y_no'], axis=1)], axis=1).to_csv(os.path.join(args.outputpath, 'train/train.csv'), index=False, header=False)
pd.concat([validation_data['y_yes'], validation_data.drop(['y_yes','y_no'], axis=1)], axis=1).to_csv(os.path.join(args.outputpath, 'validation/validation.csv'), index=False, header=False)
test_data['y_yes'].to_csv(os.path.join(args.outputpath, 'test/test_y.csv'), index=False, header=False)
test_data.drop(['y_yes','y_no'], axis=1).to_csv(os.path.join(args.outputpath, 'test/test_x.csv'), index=False, header=False)
print("## Processing complete. Exiting.")
# -
# Before starting the SageMaker Processing job, we instantiate a `SKLearnProcessor` object. This object allows you to specify the instance type to use in the job, as well as how many instances.
train_path = f"s3://{bucket}/{prefix}/train"
validation_path = f"s3://{bucket}/{prefix}/validation"
test_path = f"s3://{bucket}/{prefix}/test"
# +
from sagemaker.sklearn.processing import SKLearnProcessor
from sagemaker.processing import ProcessingInput, ProcessingOutput
from sagemaker import get_execution_role
sklearn_processor = SKLearnProcessor(
framework_version="0.23-1",
role=get_execution_role(),
instance_type="ml.m5.large",
instance_count=1,
base_job_name='sm-immday-skprocessing'
)
sklearn_processor.run(
code='preprocessing.py',
# arguments = ['arg1', 'arg2'],
inputs=[
ProcessingInput(
source=input_source,
destination="/opt/ml/processing/input",
s3_input_mode="File",
s3_data_distribution_type="ShardedByS3Key"
)
],
outputs=[
ProcessingOutput(
output_name="train_data",
source="/opt/ml/processing/output/train",
destination=train_path,
),
ProcessingOutput(output_name="validation_data", source="/opt/ml/processing/output/validation", destination=validation_path),
ProcessingOutput(output_name="test_data", source="/opt/ml/processing/output/test", destination=test_path),
]
)
# -
# !aws s3 ls $train_path/
# ---
#
# ## End of Lab 1
#
# ---
#
# ## Training
# Now we know that most of our features have skewed distributions, some are highly correlated with one another, and some appear to have non-linear relationships with our target variable. Also, for targeting future prospects, good predictive accuracy is preferred to being able to explain why that prospect was targeted. Taken together, these aspects make gradient boosted trees a good candidate algorithm.
#
# There are several intricacies to understanding the algorithm, but at a high level, gradient boosted trees works by combining predictions from many simple models, each of which tries to address the weaknesses of the previous models. By doing this the collection of simple models can actually outperform large, complex models. Other Amazon SageMaker notebooks elaborate on gradient boosting trees further and how they differ from similar algorithms.
#
# `xgboost` is an extremely popular, open-source package for gradient boosted trees. It is computationally powerful, fully featured, and has been successfully used in many machine learning competitions. Let's start with a simple `xgboost` model, trained using Amazon SageMaker's managed, distributed training framework.
#
# First we'll need to specify the ECR container location for Amazon SageMaker's implementation of XGBoost.
container = sagemaker.image_uris.retrieve(region=boto3.Session().region_name, framework='xgboost', version='latest')
# Then, because we're training with the CSV file format, we'll create `s3_input`s that our training function can use as a pointer to the files in S3, which also specify that the content type is CSV.
s3_input_train = sagemaker.inputs.TrainingInput(s3_data=train_path.format(bucket, prefix), content_type='csv')
s3_input_validation = sagemaker.inputs.TrainingInput(s3_data=validation_path.format(bucket, prefix), content_type='csv')
# First we'll need to specify training parameters to the estimator. This includes:
# 1. The `xgboost` algorithm container
# 1. The IAM role to use
# 1. Training instance type and count
# 1. S3 location for output data
# 1. Algorithm hyperparameters
#
# And then a `.fit()` function which specifies:
# 1. S3 location for output data. In this case we have both a training and validation set which are passed in.
# +
sess = sagemaker.Session()
xgb = sagemaker.estimator.Estimator(container,
role,
instance_count=1,
instance_type='ml.m4.xlarge',
output_path='s3://{}/{}/output'.format(bucket, prefix),
sagemaker_session=sess)
xgb.set_hyperparameters(max_depth=5,
eta=0.2,
gamma=4,
min_child_weight=6,
subsample=0.8,
silent=0,
objective='binary:logistic',
num_round=100)
xgb.fit({'train': s3_input_train, 'validation': s3_input_validation})
# -
# ---
#
# ## Hosting
# Now that we've trained the `xgboost` algorithm on our data, let's deploy a model that's hosted behind a real-time endpoint.
xgb_predictor = xgb.deploy(initial_instance_count=1,
instance_type='ml.m4.xlarge')
# ---
#
# ## Evaluation
# There are many ways to compare the performance of a machine learning model, but let's start by simply comparing actual to predicted values. In this case, we're simply predicting whether the customer subscribed to a term deposit (`1`) or not (`0`), which produces a simple confusion matrix.
#
# First we'll need to determine how we pass data into and receive data from our endpoint. Our data is currently stored as NumPy arrays in memory of our notebook instance. To send it in an HTTP POST request, we'll serialize it as a CSV string and then decode the resulting CSV.
#
# *Note: For inference with CSV format, SageMaker XGBoost requires that the data does NOT include the target variable.*
xgb_predictor.serializer = sagemaker.serializers.CSVSerializer()
# Now, we'll use a simple function to:
# 1. Loop over our test dataset
# 1. Split it into mini-batches of rows
# 1. Convert those mini-batches to CSV string payloads (notice, we drop the target variable from our dataset first)
# 1. Retrieve mini-batch predictions by invoking the XGBoost endpoint
# 1. Collect predictions and convert from the CSV output our model provides into a NumPy array
# !aws s3 cp $test_path/test_x.csv /tmp/test_x.csv
# !aws s3 cp $test_path/test_y.csv /tmp/test_y.csv
len(test_x.columns)
# +
def predict(data, predictor, rows=500 ):
split_array = np.array_split(data, int(data.shape[0] / float(rows) + 1))
predictions = ''
for array in split_array:
predictions = ','.join([predictions, predictor.predict(array).decode('utf-8')])
return np.fromstring(predictions[1:], sep=',')
test_x = pd.read_csv('/tmp/test_x.csv', names=[f'{i}' for i in range(59)])
test_y = pd.read_csv('/tmp/test_y.csv', names=['y'])
predictions = predict(test_x.drop(test_x.columns[0], axis=1).to_numpy(), xgb_predictor)
# -
# Now we'll check our confusion matrix to see how well we predicted versus actuals.
pd.crosstab(index=test_y['y'].values, columns=np.round(predictions), rownames=['actuals'], colnames=['predictions'])
# So, of the ~4000 potential customers, we predicted 136 would subscribe and 94 of them actually did. We also had 389 subscribers who subscribed that we did not predict would. This is less than desirable, but the model can (and should) be tuned to improve this. Most importantly, note that with minimal effort, our model produced accuracies similar to those published [here](http://media.salford-systems.com/video/tutorial/2015/targeted_marketing.pdf).
#
# _Note that because there is some element of randomness in the algorithm's subsample, your results may differ slightly from the text written above._
# ## Automatic model Tuning (optional)
# Amazon SageMaker automatic model tuning, also known as hyperparameter tuning, finds the best version of a model by running many training jobs on your dataset using the algorithm and ranges of hyperparameters that you specify. It then chooses the hyperparameter values that result in a model that performs the best, as measured by a metric that you choose.
# For example, suppose that you want to solve a binary classification problem on this marketing dataset. Your goal is to maximize the area under the curve (auc) metric of the algorithm by training an XGBoost Algorithm model. You don't know which values of the eta, alpha, min_child_weight, and max_depth hyperparameters to use to train the best model. To find the best values for these hyperparameters, you can specify ranges of values that Amazon SageMaker hyperparameter tuning searches to find the combination of values that results in the training job that performs the best as measured by the objective metric that you chose. Hyperparameter tuning launches training jobs that use hyperparameter values in the ranges that you specified, and returns the training job with highest auc.
#
from sagemaker.tuner import IntegerParameter, CategoricalParameter, ContinuousParameter, HyperparameterTuner
hyperparameter_ranges = {'eta': ContinuousParameter(0, 1),
'min_child_weight': ContinuousParameter(1, 10),
'alpha': ContinuousParameter(0, 2),
'max_depth': IntegerParameter(1, 10)}
objective_metric_name = 'validation:auc'
tuner = HyperparameterTuner(xgb,
objective_metric_name,
hyperparameter_ranges,
max_jobs=20,
max_parallel_jobs=3)
tuner.fit({'train': s3_input_train, 'validation': s3_input_validation})
boto3.client('sagemaker').describe_hyper_parameter_tuning_job(
HyperParameterTuningJobName=tuner.latest_tuning_job.job_name)['HyperParameterTuningJobStatus']
# return the best training job name
tuner.best_training_job()
# Deploy the best trained or user specified model to an Amazon SageMaker endpoint
tuner_predictor = tuner.deploy(initial_instance_count=1,
instance_type='ml.m4.xlarge')
# Create a serializer
tuner_predictor.serializer = sagemaker.serializers.CSVSerializer()
# Predict
predictions = predict(test_data.drop(['y_no', 'y_yes'], axis=1).to_numpy(),tuner_predictor)
# Collect predictions and convert from the CSV output our model provides into a NumPy array
pd.crosstab(index=test_data['y_yes'], columns=np.round(predictions), rownames=['actuals'], colnames=['predictions'])
# ---
#
# ## Extensions
#
# This example analyzed a relatively small dataset, but utilized Amazon SageMaker features such as distributed, managed training and real-time model hosting, which could easily be applied to much larger problems. In order to improve predictive accuracy further, we could tweak value we threshold our predictions at to alter the mix of false-positives and false-negatives, or we could explore techniques like hyperparameter tuning. In a real-world scenario, we would also spend more time engineering features by hand and would likely look for additional datasets to include which contain customer information not available in our initial dataset.
# ### (Optional) Clean-up
#
# If you are done with this notebook, please run the cell below. This will remove the hosted endpoint you created and avoid any charges from a stray instance being left on.
xgb_predictor.delete_endpoint(delete_endpoint_config=True)
tuner_predictor.delete_endpoint(delete_endpoint_config=True)
| processing_xgboost.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.0.2
# language: julia
# name: julia-1.0
# ---
using Plots, ComplexPhasePortrait, ApproxFun, SingularIntegralEquations, DifferentialEquations
gr();
# # M3M6: Methods of Mathematical Physics
#
# $$
# \def\dashint{{\int\!\!\!\!\!\!-\,}}
# \def\infdashint{\dashint_{\!\!\!-\infty}^{\,\infty}}
# \def\D{\,{\rm d}}
# \def\E{{\rm e}}
# \def\dx{\D x}
# \def\dt{\D t}
# \def\dz{\D z}
# \def\C{{\mathbb C}}
# \def\R{{\mathbb R}}
# \def\CC{{\cal C}}
# \def\HH{{\cal H}}
# \def\I{{\rm i}}
# \def\qqqquad{\qquad\qquad}
# \def\qqfor{\qquad\hbox{for}\qquad}
# \def\qqwhere{\qquad\hbox{where}\qquad}
# \def\Res_#1{\underset{#1}{\rm Res}}\,
# \def\sech{{\rm sech}\,}
# \def\acos{\,{\rm acos}\,}
# \def\vc#1{{\mathbf #1}}
# \def\ip<#1,#2>{\left\langle#1,#2\right\rangle}
# \def\norm#1{\left\|#1\right\|}
# \def\half{{1 \over 2}}
# $$
#
# Dr. <NAME>
# <br>
# <EMAIL>
#
# <br>
# Website: https://github.com/dlfivefifty/M3M6LectureNotes
#
#
# # Lecture 16: Solving differential equations with orthogonal polynomials
#
#
# This lecture we do the following:
#
# 1. Recurrence relationships for Chebyshev and ultrashperical polynomials
# - Conversion
# - Three-term recurrence and Jacobi operators
# 2. Application: solving differential equations
# - First order constant coefficients differential equations
# - Second order constant coefficient differential equations with boundary conditions
# - Non-constant coefficients
#
#
#
# That is, we introduce recurrences related to ultraspherical polynomials. This allows us to represent general linear differential equations as almost-banded systems.
# ## Recurrence relationships for Chebyshev and ultraspherical polynomials
#
#
# We have discussed general properties, but now we want to discuss some classical orthogonal polynomials, beginning with Chebyshev (first kind) $T_n(x)$, which is orthogonal w.r.t.
# $$1\over \sqrt{1-x^2}$$
# and ultraspherical $C_n^{(\lambda)}(x)$, which is orthogonal w.r.t.
# $$(1-x^2)^{\lambda - \half}$$
# for $\lambda > 0$. Note that Chebyshev (second kind) satisfies $U_n(x) = C_n^{(1)}(x)$.
#
# For Chebyshev, recall we have the normalization constant (here we use a superscript $T_n(x) = k_n^{\rm T} x^n + O(x^{n-1})$)
# $$
# k_0^{\rm T} = 1, k_n^{\rm T} = 2^{n-1}
# $$
# For Ultraspherical $C_n^{(\lambda)}$, this is
# $$
# k_n^{(\lambda)} = {2^n (\lambda)_n \over n!} = {2^n \lambda (\lambda+1) (\lambda+2) \cdots (\lambda+n-1) \over n!}
# $$
# where $(\lambda)_n$ is the Pochhammer symbol. Note for $U_n(x) = C_n^{(1)}(x)$ this simplifies to $k_n^{\rm U} = k_n^{(1)} = 2^n$.
#
# We have already found the recurrence for Chebyshev:
# $$
# x T_n(x) = {T_{n-1}(x) \over 2} + {T_{n+1}(x) \over 2}
# $$
# We will show that we can use this to find the recurrence for _all_ ultraspherical polynomials. But first we need some special recurrences.
#
# **Remark** Jacobi, Laguerre, and Hermite all have similar relationships, which will be discussed further in the problem sheet.
#
# ### Derivatives
#
# It turns out that the derivative of $T_n(x)$ is precisely a multiple of $U_{n-1}(x)$, and similarly the derivative of $C_n^{(\lambda)}$ is a multiple of $C_{n-1}^{(\lambda+1)}$.
#
# **Proposition (Chebyshev derivative)** $$T_n'(x) = n U_{n-1}(x)$$
#
# **Proof**
# We first show that $T_n'(x)$ is othogonal w.r.t. $\sqrt{1-x^2}$ to all polynomials of degree $m < n-1$, denoted $f_m$, using integration by parts:
# $$
# \ip<T_n',f_m>_{\rm U} = \int_{-1}^1 T_n'(x) f_m(x) \sqrt{1-x^2} \dx = -\int_{-1}^1 T_n(x) (f_m'(x)(1-x^2) + xf_m) {1 \over \sqrt{1-x^2}} \dx = - \ip<T_n, f_m'(1-x^2) + x f_m >_{\rm T} = 0
# $$
# since $f_m'(1-x^2) + f_m $ is degree $m-1 +2 = m+1 < n$.
#
# The constant works out since
# $$
# T_n'(x) = {\D \over \dx} (2^{n-1} x^n) + O(x^{n-2}) = n 2^{n-1} x^{n-1} + O(x^{n-2})
# $$
# ⬛️
#
# The exact same proof shows the following:
#
# **Proposition (Ultraspherical derivative)**
# $${\D \over \dx} C_n^{(\lambda)}(x) = 2 \lambda C_{n-1}^{(\lambda+1)}(x)$$
#
# Like the three-term recurrence and Jacobi operators, it is useful to express this in matrix form. That is, for the derivatives of $T_n(x)$ we get
# $$
# {\D \over \dx} \begin{pmatrix} T_0(x) \\ T_1(x) \\ T_2(x) \\ \vdots \end{pmatrix}= \begin{pmatrix}
# 0 \cr
# 1 \cr
# & 2 \cr
# && 3 \cr
# &&&\ddots
# \end{pmatrix} \begin{pmatrix} U_0(x) \\ U_1(x) \\ U_2(x) \\ \vdots \end{pmatrix}
# $$
# which let's us know that, for
# $$
# f(x) = (T_0(x),T_1(x),\ldots) \begin{pmatrix} f_0\\f_1\\\vdots \end{pmatrix}
# $$
# we have a derivative operator in coefficient space as
# $$
# f'(x) = (U_0(x),U_1(x),\ldots)\begin{pmatrix}
# 0 & 1 \cr
# && 2 \cr
# &&& 3 \cr
# &&&&\ddots
# \end{pmatrix} \begin{pmatrix} f_0\\f_1\\\vdots \end{pmatrix}
# $$
#
# _Demonstration_ Here we see that applying a matrix to a vector of coefficients successfully calculates the derivative:
f = Fun(x -> cos(x^2), Chebyshev()) # f is expanded in Chebyshev coefficients
n = ncoefficients(f) # This is the number of coefficients
D = zeros(n-1,n)
for k=1:n-1
D[k,k+1] = k
end
D
# Here `D*f.coefficients` gives the vector of coefficients corresponding to the derivative, but now the coefficients are in the $U_n(x)$ basis, that is, `Ultraspherical(1)`:
# +
fp = Fun(Ultraspherical(1), D*f.coefficients)
fp(0.1)
# -
# Indeed, it matches the "true" derivative:
f'(0.1)
-2*0.1*sin(0.1^2)
# Note that in ApproxFun.jl we can construct these operators rather nicely:
D = Derivative()
(D*f)(0.1)
# Here we see that we can write produce the ∞-dimensional version as follows:
D : Chebyshev() → Ultraspherical(1)
# ### Conversion
#
#
# We can convert between any two polynomial bases using a lower triangular operator, because their span's are equivalent. In the case of Chebyshev and ultraspherical polynomials, they have the added property that they are banded.
#
# **Proposition (Chebyshev T-to-U conversion)**
# \begin{align*}
# T_0(x) &= U_0(x) \\
# T_1(x) &= {U_1(x) \over 2} \\
# T_n(x) &= {U_n(x) \over 2} - {U_{n-2}(x) \over 2}
# \end{align*}
#
# **Proof**
#
# Before we do the proof, note that the fact that there are limited non-zero entries follows immediately: if $m < n-2$ we have
# $$
# \ip<T_n, U_m>_{\rm U} = \ip<T_n, (1-x^2)U_m>_{\rm T} = 0
# $$
#
# To actually determine the entries, we use the trigonometric formulae. Recall for $x = (z + z^{-1})/2$, $z = \E^{\I \theta}$, we have
# \begin{align*}
# T_n(x) &= \cos n \theta = {z^{-n} + z^n \over 2}\\
# U_n(x) &= {\sin (n+1) \theta \over \sin \theta} = {z^{n+1} - z^{-n-1} \over z - z^{-1}} = z^{-n} + z^{2-n} + \cdots + \cdots + z^{n-2} + z^n
# \end{align*}
# The result follows immediately.
#
# ⬛️
#
# **Corollary (Ultrapherical λ-to-(λ+1) conversion)**
# $$
# C_n^{(\lambda)}(x) = {\lambda \over n+ \lambda} (C_n^{(\lambda+1)}(x) - C_{n-2}^{(\lambda+1)}(x))
# $$
#
# **Proof** This follows from differentiating the previous result. For example:
# \begin{align*}
# {\D\over \dx} T_0(x) &= {\D\over \dx} U_0(x) \\
# {\D\over \dx} T_1(x) &= {\D\over \dx} {U_1(x) \over 2} \\
# {\D\over \dx} T_n(x) &= {\D\over \dx} \left({U_n(x) \over 2} - {U_{n-2} \over 2} \right)
# \end{align*}
# becomes
# \begin{align*}
# 0 &= 0\\
# U_0(x) &= C_1^{(2)}(x) \\
# n U_{n-1}(x) &= C_{n-1}^{(2)}(x) - C_{n-3}^{(2)}(x)
# \end{align*}
#
# Differentiating this repeatedly completes the proof.
#
# ⬛️
#
#
# Note we can write this in matrix form, for example, we have
# $$
# \underbrace{\begin{pmatrix}1 \cr
# 0 & \half\cr
# -\half & 0 & \half \cr
# &\ddots &\ddots & \ddots\end{pmatrix} }_{S_0^\top} \begin{pmatrix}
# U_0(x) \\ U_1(x) \\ U_2(x) \\ \vdots \end{pmatrix} = \begin{pmatrix} T_0(x) \\ T_1(x) \\ T_2(x) \\ \vdots \end{pmatrix}
# $$
#
# therefore,
# $$
# f(x) = (T_0(x),T_1(x),\ldots) \begin{pmatrix} f_0\\f_1\\\vdots \end{pmatrix} = (U_0(x),U_1(x),\ldots) S_0 \begin{pmatrix} f_0\\f_1\\\vdots \end{pmatrix}
# $$
#
# Again, we can construct this nicely in ApproxFun:
S₀ = I : Chebyshev() → Ultraspherical(1)
f = Fun(exp, Chebyshev())
g = S₀*f
g(0.1) - exp(0.1)
# ### Ultraspherical Three-term recurrence
#
# **Theorem (three-term recurrence for Chebyshev U)**
# \begin{align*}
# x U_0(x) &= {U_1(x) \over 2} \\
# x U_n(x) &= {U_{n-1}(x) \over 2} + {U_{n+1}(x) \over 2}
# \end{align*}
#
# **Proof**
# Differentiating
# \begin{align*}
# x T_0(x) &= T_1(x) \\
# x T_n(x) &= {T_{n-1}(x) \over 2} + {T_{n+1}(x) \over 2}
# \end{align*}
# we get
# \begin{align*}
# T_0(x) &= U_0(x) \\
# T_n(x) + n x U_{n-1}(x) &= {(n-1) U_{n-2}(x) \over 2} + {(n+1) U_n(x) \over 2}
# \end{align*}
# substituting in the conversion $T_n(x) = (U_n(x) - U_{n-2}(x))/2$ we get
# \begin{align*}
# T_0(x) &= U_0(x) \\
# n x U_{n-1}(x) &= {(n-1) U_{n-2}(x) \over 2} + {(n+1) U_n(x) \over 2} - (U_n(x) - U_{n-2}(x))/2 = {n U_{n-2}(x) \over 2} + {n U_n(x) \over 2}
# \end{align*}
#
# ⬛️
#
# Differentiating this theorem again and applying the conversion we get the following
#
# **Corollary (three-term recurrence for ultrashperical)**
# \begin{align*}
# x C_0^{(\lambda)}(x) &= {1 \over 2\lambda } C_1^{(\lambda)}(x) \\
# x C_n^{(\lambda)}(x) &= {n+2\lambda-1 \over 2(n+\lambda)} C_{n-1}^{(\lambda)}(x) + {n+1 \over 2(n+\lambda)} C_{n+1}^{(\lambda)}(x)
# \end{align*}
#
#
# Here's an example of the Jacobi operator (which is the transpose of the multiplciation by $x$ operator):
Multiplication(Fun(), Ultraspherical(2))'
# ## Application: solving differential equations
#
# The preceding results allowed us to represent
#
# 1. Differentiation
# 2. Conversion
# 3. Multiplication
#
# as banded operators. We will see that we can combine these, along with
#
# 4\. Evaluation
#
# to solve ordinary differential equations.
#
# ### First order, constant coefficient differential equations
#
# Consider the simplest ODE:
# \begin{align*}
# u(0) &= 0 \\
# u'(x) - u(x) &= 0
# \end{align*}
# and suppose represent $u(x)$ in its Chebyshev expansion, with to be determined coefficents. In other words, we want to calculate coefficients $u_k$ such that
# $$
# u(x) = \sum_{k=0}^\infty u_k T_k(x) = (T_0(x), T_1(x), \ldots) \begin{pmatrix} u_0 \\ u_1 \\ \vdots \end{pmatrix}
# $$
# In this case we know that $u(x) = \E^x$, but we would still need other means to calculate $u_k$ (They are definitely not as simple as Taylor series coefficients).
#
# We can express the constraints as acting on the coefficients. For example, we have
# $$
# u(0) = (T_0(0), T_1(0), \ldots) \begin{pmatrix} u_0\\u_1\\\vdots \end{pmatrix} = (1,0,-1,0,1,\ldots) \begin{pmatrix} u_0\\u_1\\\vdots \end{pmatrix}
# $$
# We also have
# $$u'(x) = (U_0(x),U_1(x),\ldots) \begin{pmatrix}
# 0 & 1 \cr
# && 2 \cr
# &&& 3 \cr
# &&&&\ddots
# \end{pmatrix}\begin{pmatrix} u_0\\u_1\\\vdots \end{pmatrix}
# $$
# To represent $u'(x) - u(x)$, we need to make sure the bases are compatible. In other words, we want to express $u(x)$ in it's $U_k(x)$ basis using the conversion operator $S_0$:
# $$u(x) = (U_0(x),U_1(x),\ldots) \begin{pmatrix}
# 1 &0 & -\half \cr
# & \half & 0 & -\half \cr
# &&\ddots & \ddots & \ddots
# \end{pmatrix}\begin{pmatrix} u_0\\u_1\\\vdots \end{pmatrix}
# $$
#
# Which gives us,
# $$
# u'(x) - u(x) = (U_0(x),U_1(x),\ldots) \begin{pmatrix}
# -1 &1 & \half \cr
# & -\half & 2 & \half \cr
# && -\half & 3 & \half \cr
# &&&\ddots & \ddots & \ddots
# \end{pmatrix} \begin{pmatrix} u_0\\u_1\\\vdots \end{pmatrix}
# $$
#
#
# Combing the differential part and the evaluation part, we arrive at an (infinite) system of equations for the coefficients $u_0,u_1,\dots$:
# $$
# \begin{pmatrix}
# 1 & 0 & -1 & 0 & 1 & \cdots \\
# -1 &1 & \half \cr
# & -\half & 2 & \half \cr
# && -\half & 3 & \half \cr
# &&&\ddots & \ddots & \ddots
# \end{pmatrix} \begin{pmatrix} u_0\\u_1\\\vdots \end{pmatrix} = \begin{pmatrix} 1 \\ 0 \\ 0 \\ \vdots \end{pmatrix}
# $$
#
# How to solve this system is outside the scope of this course (though a simple approach is to truncate the infinite system to finite systems). We can however do this in ApproxFun:
B = Evaluation(0.0) : Chebyshev()
D = Derivative() : Chebyshev() → Ultraspherical(1)
S₀ = I : Chebyshev() → Ultraspherical(1)
L = [B;
D - S₀]
# We can solve this system as follows:
u = L \ [1; 0]
plot(u)
# It matches the "true" result:
u(0.1) - exp(0.1)
# Note we can incorporate right-hand sides as well, for example, to solve $u'(x) - u(x) = f(x)$, by expanding $f$ in its Chebyshev U series.
#
# ### Second-order constanst coefficient equations
#
# This approach extends to second-order constant-coefficient equations by using ultraspherical polynomials. Consider
# \begin{align*}
# u(-1) &= 1\\
# u(1) &= 0\\
# u''(x) + u'(x) + u(x) &= 0
# \end{align*}
# Evaluation works as in the first-order case. To handle second-derivatives, we need $C^{(2)}$ polynomials:
D₀ = Derivative() : Chebyshev() → Ultraspherical(1)
D₁ = Derivative() : Ultraspherical(1) → Ultraspherical(2)
D₁*D₀ # 2 zeros not printed in (1,1) and (1,2) entry
# For the identity operator, we use two conversion operators:
S₀ = I : Chebyshev() → Ultraspherical(1)
S₁ = I : Ultraspherical(1) → Ultraspherical(2)
S₁*S₀
# And for the first derivative, we use a derivative and then a conversion:
S₁*D₀ # or could have been D₁*S₀
# Putting everything together we get:
# +
B₋₁ = Evaluation(-1) : Chebyshev()
B₁ = Evaluation(1) : Chebyshev()
# u(-1)
# u(1)
# u'' + u' +u
L = [B₋₁;
B₁;
D₁*D₀ + S₁*D₀ + S₁*S₀]
# -
u = L \ [1.0,0.0,0.0]
plot(u)
# ### Variable coefficients
#
# Consider the Airy ODE
# \begin{align*}
# u(-1) &= 1\\
# u(1) &= 0\\
# u''(x) - xu(x) &= 0
# \end{align*}
#
# to handle, this, we need only use the Jacobi operator to represent multiplication by $x$:
x = Fun()
Jᵗ = Multiplication(x) : Chebyshev() → Chebyshev() # transpose of the Jacobi operator
# We set op ther system as follows:
L = [B₋₁; # u(-1)
B₁ ; # u(1)
D₁*D₀ - S₁*S₀*Jᵗ] # u'' - x*u
u = L \ [1.0;0.0;0.0]
plot(u; legend=false)
# If we introduce a small parameter, that is, solve
# \begin{align*}
# u(-1) &= 1\\
# u(1) &= 0\\
# \epsilon u''(x) - xu(x) &= 0
# \end{align*}
# we can see pretty hard to compute solutions:
# +
ε = 1E-6
L = [B₋₁;
B₁ ;
ε*D₁*D₀ - S₁*S₀*Jᵗ]
u = L \ [1.0;0.0;0.0]
plot(u; legend=false)
# -
# Because of the banded structure, this can be solved fast:
# +
ε = 1E-10
L = [B₋₁;
B₁ ;
ε*D₁*D₀ - S₁*S₀*Jᵗ]
@time u = L \ [1.0;0.0;0.0]
@show ncoefficients(u);
# -
# To handle other variable coefficients, first consider a polynomial $p(x)$. If Multiplication by $x$ is represented by multiplying the coefficients by $J^\top$, then multiplication by $p$ is represented by $p(J^\top)$:
M = -I + Jᵗ + (Jᵗ)^2 # represents -1+x+x^2
# +
ε = 1E-6
L = [B₋₁;
B₁ ;
ε*D₁*D₀ - S₁*S₀*M]
@time u = L \ [1.0;0.0;0.0]
@show ε*u''(0.1) - (-1+0.1+0.1^2)*u(0.1)
plot(u)
# -
# For other smooth functions, we first approximate in a polynomial basis, and without loss of generality we use Chebyshev T basis. For example, consider
# \begin{align*}
# u(-1) &= 1\\
# u(1) &= 0\\
# \epsilon u''(x) - \E^x u(x) &= 0
# \end{align*}
# where
# $$
# \E^x \approx p(x) = \sum_{k=0}^{m-1} p_k T_k(x)
# $$
# Evaluating at a point $x$, recall Clenshaw's algorithm:
# \begin{align*}
# \gamma_{n-1} &= 2p_{n-1} \\
# \gamma_{n-2} &= 2p_{n-2} + 2x \gamma_{n-1} \\
# \gamma_{n-3} &= 2 p_{n-3} + 2x \gamma_{n-2} - \gamma_{n-1} \\
# & \vdots \\
# \gamma_1 &= p_1 + x \gamma_2 - \half \gamma_3 \\
# p(x) = \gamma_0 &= p_0 + x \gamma_1 - \half \gamma_2
# \end{align*}
# If multiplication by $x$ becomes $J^\top$, then multiplication by $p(x)$ becomes $p(J^\top)$, and hence we calculate:\
# \begin{align*}
# \Gamma_{n-1} &= 2p_{n-1}I \\
# \Gamma_{n-2} &= 2p_{n-2}I + 2J^\top \Gamma_{n-1} \\
# \Gamma_{n-3} &= 2 p_{n-3}I + 2J^\top \Gamma_{n-2} - \Gamma_{n-1} \\
# & \vdots \\
# \Gamma_1 &= p_1I + J^\top \Gamma_2 - \half \Gamma_3 \\
# p(J^\top) = \Gamma_0 &= p_0 + x \Gamma_1 - \half \Gamma_2
# \end{align*}
#
# Here is an example:
p = Fun(exp, Chebyshev()) # polynomial approximation to exp(x)
M = Multiplication(p) : Chebyshev() # constructed using Clenshaw:
ApproxFun.bandwidths(M) # still banded
# +
ε = 1E-6
L = [B₋₁;
B₁ ;
ε*D₁*D₀ + S₁*S₀*M]
@time u = L \ [1.0;0.0;0.0]
@show ε*u''(0.1) + exp(0.1)*u(0.1)
plot(u)
| Lecture 16.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Deploy a Serverless XGBoost Model Server
# --------------------------------------------------------------------
#
# The following notebook demonstrates how to deploy an XGBoost model server (a.k.a <b>Nuclio-serving</b>)
#
# #### **notebook how-to's**
# * Write and test model serving class in a notebook.
# * Deploy the model server function.
# * Invoke and test the serving function.
# <a id="top"></a>
# #### **steps**
# **[define a new function and its dependencies](#define-function)**<br>
# **[test the model serving class locally](#test-locally)**<br>
# **[deploy our serving class using as a serverless function](#deploy)**<br>
# **[test our model server using HTTP request](#test-model-server)**<br>
# nuclio: ignore
import nuclio
# <a id="define-function"></a>
# ### **define a new function and its dependencies**
# +
# %nuclio config kind="nuclio:serving"
# %nuclio env MODEL_CLASS=XGBoostModel
# %nuclio config spec.build.baseImage = "mlrun/ml-models"
# -
# ## Function Code
# +
# import kfserving
import os
import json
import numpy as np
import xgboost as xgb
from cloudpickle import load
### Model Serving Class
import mlrun
class XGBoostModel(mlrun.runtimes.MLModelServer):
def load(self):
model_file, extra_data = self.get_model(".pkl")
self.model = load(open(str(model_file), "rb"))
def predict(self, body):
try:
feats = np.asarray(body["instances"], dtype=np.float32).reshape(-1, 4)
result = self.model.predict(feats, validate_features=False)
return result.tolist()
except Exception as e:
raise Exception("Failed to predict %s" % e)
# -
# The following end-code annotation tells ```nuclio``` to stop parsing the notebook from this cell. _**Please do not remove this cell**_:
# +
# nuclio: end-code
# -
# ### mlconfig
from mlrun import mlconf
import os
mlconf.dbpath = mlconf.dbpath or "http://mlrun-api:8080"
mlconf.artifact_path = mlconf.artifact_path or f"{os.environ['HOME']}/artifacts"
# <a id="test-locally"></a>
# ## Test the function locally
#
# The class above can be tested locally. Just instantiate the class, `.load()` will load the model to a local dir.
#
# > **Verify there is a model file in the model_dir path (generated by the training notebook)**
# +
model_dir = os.path.join(mlconf.artifact_path, "models")
print(model_dir)
my_server = XGBoostModel("my-model", model_dir=model_dir)
my_server.load()
# -
REPO_URL = "https://raw.githubusercontent.com/yjb-ds/testdata/master"
DATA_PATH = "data/classifier-data.csv"
MODEL_PATH = "models/xgb_test"
import pandas as pd
xtest = pd.read_csv(f"{REPO_URL}/{DATA_PATH}")
# We can use the `.predict(body)` method to test the model.
import json, numpy as np
preds = my_server.predict({"instances":xtest.values[:10,:-1].tolist()})
print("predicted class:", preds)
# <a id="deploy"></a>
# ### **deploy our serving class using as a serverless function**
# in the following section we create a new model serving function which wraps our class , and specify model and other resources.
#
# the `models` dict store model names and the assosiated model **dir** URL (the URL can start with `S3://` and other blob store options), the faster way is to use a shared file volume, we use `.apply(mount_v3io())` to attach a v3io (iguazio data fabric) volume to our function. By default v3io will mount the current user home into the `\User` function path.
#
# **verify the model dir does contain a valid `model.bst` file**
from mlrun import new_model_server, mount_v3io
import requests
# +
fn = new_model_server("xgb-test",
model_class="XGBoostModel",
models={"xgb_serving_v2": f"{model_dir}"})
fn.spec.description = "xgboost test data classification server"
fn.metadata.categories = ["serving", "ml"]
fn.metadata.labels = {"author": "yaronh", "framework": "xgboost"}
fn.export("function.yaml")
# -
# ## tests
if "V3IO_HOME" in list(os.environ):
from mlrun import mount_v3io
fn.apply(mount_v3io())
else:
# is you set up mlrun using the instructions at
# https://github.com/mlrun/mlrun/blob/master/hack/local/README.md
from mlrun.platforms import mount_pvc
fn.apply(mount_pvc("nfsvol", "nfsvol", "/home/jovyan/data"))
addr = fn.deploy(dashboard="http://172.17.0.66:8070", project="churn-project")
addr
# <a id="test-model-server"></a>
# ### **test our model server using HTTP request**
#
#
# We invoke our model serving function using test data, the data vector is specified in the `instances` attribute.
# KFServing protocol event
event_data = {"instances": xtest.values[:10,:-1].tolist()}
# +
import json
resp = requests.put("http://192.168.99.135:30791" + "/xgb_serving_v2/predict", json=json.dumps(event_data))
# mlutils function for this?
tl = resp.text.replace("[","").replace("]","").split(",")
#assert preds == [int(i) for i in np.asarray(tl)]
# -
tl
preds
# **[back to top](#top)**
| xgb_serving/xgb_serving.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# A crash course in
# <b><font size=44px><center> Surviving Titanic</center></font></b>
# <img src='http://4.media.bustedtees.cvcdn.com/f/-/bustedtees.d6ab8f8f-a63a-45fd-acac-142e2c22.gif' width=400>
# <center> (with numpy and matplotlib)</center>
#
# ---
#
# This notebook's gonna teach you to use the basic data science stack for python: jupyter, numpy, matplotlib and sklearn.
# ### Part I: Jupyter notebooks in a nutshell
# * You are reading this line in a jupyter notebook.
# * A notebook consists of cells. A cell can contain either code or hypertext.
# * This cell contains hypertext. The next cell contains code.
# * You can __run a cell__ with code by selecting it (click) and pressing `Ctrl + Enter` to execute the code and display output(if any).
# * If you're running this on a device with no keyboard, ~~you are doing it wrong~~ use topbar (esp. play/stop/restart buttons) to run code.
# * Behind the curtains, there's a python interpreter that runs that code and remembers anything you defined.
#
# Run these cells to get started
a = 5
print(a * 2)
# * `Ctrl + S` to save changes (or use the button that looks like a floppy disk)
# * Top menu -> Kernel -> Interrupt (or Stop button) if you want it to stop running cell midway.
# * Top menu -> Kernel -> Restart (or cyclic arrow button) if interrupt doesn't fix the problem (you will lose all variables).
# * For shortcut junkies like us: Top menu -> Help -> Keyboard Shortcuts
#
#
# * More: [Hacker's guide](http://arogozhnikov.github.io/2016/09/10/jupyter-features.html), [Beginner's guide'](https://jupyter-notebook-beginner-guide.readthedocs.io/en/latest/), [Datacamp tutorial](https://www.datacamp.com/community/tutorials/tutorial-jupyter-notebook)
#
# Now __the most important feature__ of jupyter notebooks for this course:
# * if you're typing something, press `Tab` to see automatic suggestions, use arrow keys + enter to pick one.
# * if you move your cursor inside some function and press `__Shift + Tab__`, you'll get a help window. `Shift + (Tab , Tab)` will expand it.
# run this first
import math
# +
# place your cursor at the end of the unfinished line below to find a function
# that computes arctangent from two parameters (should have 2 in it's name)
# once you chose it, press shift + tab + tab(again) to see the docs
math.atan2 # <---
# -
# ### Part II: Loading data with Pandas
# Pandas is a library that helps you load the data, prepare it and perform some lightweight analysis. The god object here is the `pandas.DataFrame` - a 2d table with batteries included.
#
# In the cell below we use it to read the data on the infamous titanic shipwreck.
#
# __please keep running all the code cells as you read__
import pandas as pd
# this yields a pandas.DataFrame
data = pd.read_csv("train.csv", index_col='PassengerId')
# +
# Selecting rows
head = data[:10]
head # if you leave an expression at the end of a cell, jupyter will "display" it automatically
# -
# #### About the data
# Here's some of the columns
# * Name - a string with person's full name
# * Survived - 1 if a person survived the shipwreck, 0 otherwise.
# * Pclass - passenger class. Pclass == 3 is cheap'n'cheerful, Pclass == 1 is for moneybags.
# * Sex - a person's gender (in those good ol' times when there were just 2 of them)
# * Age - age in years, if available
# * Sibsp - number of siblings on a ship
# * Parch - number of parents on a ship
# * Fare - ticket cost
# * Embarked - port where the passenger embarked
# * C = Cherbourg; Q = Queenstown; S = Southampton
# table dimensions
print("len(data) = ", len(data))
print("data.shape = ", data.shape)
print(type(data))
# select a single row
print(data.loc[4])
# select a single column.
ages = data["Age"]
print(ages[:10]) # alternatively: data.Age
# select several columns and rows at once
# alternatively: data[["Fare","Pclass"]].loc[5:10]
data.loc[5:10, ("Fare", "Pclass")]
# ## Your turn:
#
# +
# select passengers number 13 and 666 - did they survive?
print(data.loc[13]['Survived'])
print(data.loc[666]['Survived'])
data.loc[666]
# -
# compute the overall survival rate (what fraction of passengers survived the shipwreck)
res = sum(data.Survived) / len(data.Survived)
print(res)
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
#
# Pandas also has some basic data analysis tools. For one, you can quickly display statistical aggregates for each column using `.describe()`
data.describe()
# Some columns contain __NaN__ values - this means that there is no data there. For example, passenger `#5` has unknown age. To simplify the future data analysis, we'll replace NaN values by using pandas `fillna` function.
#
# _Note: we do this so easily because it's a tutorial. In general, you think twice before you modify data like this._
data.iloc[5]
data['Age'] = data['Age'].fillna(value=data['Age'].mean())
data['Fare'] = data['Fare'].fillna(value=data['Fare'].mean())
data.iloc[5]
# More pandas:
# * A neat [tutorial](http://pandas.pydata.org/) from pydata
# * Official [tutorials](https://pandas.pydata.org/pandas-docs/stable/tutorials.html), including this [10 minutes to pandas](https://pandas.pydata.org/pandas-docs/stable/10min.html#min)
# * Bunch of cheat sheets awaits just one google query away from you (e.g. [basics](http://blog.yhat.com/static/img/datacamp-cheat.png), [combining datasets](https://pbs.twimg.com/media/C65MaMpVwAA3v0A.jpg) and so on).
# ### Part III: Numpy and vectorized computing
#
# Almost any machine learning model requires some computational heavy lifting usually involving linear algebra problems. Unfortunately, raw python is terrible at this because each operation is interpreted at runtime.
#
# So instead, we'll use `numpy` - a library that lets you run blazing fast computation with vectors, matrices and other tensors. Again, the god object here is `numpy.ndarray`:
# +
import numpy as np
a = np.array([1, 2, 3, 4, 5])
b = np.array([5, 4, 3, 2, 1])
print("a = ", a)
print("b = ", b)
# math and boolean operations can applied to each element of an array
print("a + 1 =", a + 1)
print("a * 2 =", a * 2)
print("a == 2", a == 2)
# ... or corresponding elements of two (or more) arrays
print("a + b =", a + b)
print("a * b =", a * b)
# -
# Your turn: compute half-products of a and b elements (halves of products)
print(a * b / 2)
# compute elementwise quotient between squared a and (b plus 1)
print(a ** 2 - (b + 1))
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
#
# ### How fast is it, harry?
# 
#
# Let's compare computation time for python and numpy
# * Two arrays of 10^6 elements
# * first - from 0 to 1 000 000
# * second - from 99 to 1 000 099
#
# * Computing:
# * elemwise sum
# * elemwise product
# * square root of first array
# * sum of all elements in the first array
#
# +
# %time
# ^-- this "magic" measures and prints cell computation time
# Option I: pure python
arr_1 = range(1000000)
arr_2 = range(99, 1000099)
a_sum = []
a_prod = []
sqrt_a1 = []
for i in range(len(arr_1)):
a_sum.append(arr_1[i]+arr_2[i])
a_prod.append(arr_1[i]*arr_2[i])
a_sum.append(arr_1[i]**0.5)
arr_1_sum = sum(arr_1)
# +
# %time
# Option II: start from python, convert to numpy
arr_1 = range(1000000)
arr_2 = range(99, 1000099)
arr_1, arr_2 = np.array(arr_1), np.array(arr_2)
a_sum = arr_1 + arr_2
a_prod = arr_1 * arr_2
sqrt_a1 = arr_1 ** .5
arr_1_sum = arr_1.sum()
# +
# %time
# Option III: pure numpy
arr_1 = np.arange(1000000)
arr_2 = np.arange(99, 1000099)
a_sum = arr_1 + arr_2
a_prod = arr_1 * arr_2
sqrt_a1 = arr_1 ** .5
arr_1_sum = arr_1.sum()
# -
# If you want more serious benchmarks, take a look at [this](http://brilliantlywrong.blogspot.ru/2015/01/benchmarks-of-speed-numpy-vs-all.html).
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
#
# There's also a bunch of pre-implemented operations including logarithms, trigonometry, vector/matrix products and aggregations.
# +
a = np.array([1, 2, 3, 4, 5])
b = np.array([5, 4, 3, 2, 1])
print("numpy.sum(a) = ", np.sum(a))
print("numpy.mean(a) = ", np.mean(a))
print("numpy.min(a) = ", np.min(a))
print("numpy.argmin(b) = ", np.argmin(b)) # index of minimal element
# dot product. Also used for matrix/tensor multiplication
print("numpy.dot(a,b) = ", np.dot(a, b))
print("numpy.unique(['male','male','female','female','male']) = ", np.unique(
['male', 'male', 'female', 'female', 'male']))
# and tons of other stuff. see http://bit.ly/2u5q430 .
# -
# The important part: all this functionality works with dataframes:
print("Max ticket price: ", np.max(data["Fare"]))
print("\nThe guy who paid the most:\n", data.loc[np.argmax(data["Fare"])])
# your code: compute mean passenger age and the oldest guy on the ship
np.mean(data.Age)
np.max(data.Age)
# +
print("Boolean operations")
print('a = ', a)
print('b = ', b)
print("a > 2", a > 2)
print("numpy.logical_not(a>2) = ", np.logical_not(a > 2))
print("numpy.logical_and(a>2,b>2) = ", np.logical_and(a > 2, b > 2))
print("numpy.logical_or(a>4,b<3) = ", np.logical_or(a > 2, b < 3))
print("\n shortcuts")
print("~(a > 2) = ", ~(a > 2)) # logical_not(a > 2)
print("(a > 2) & (b > 2) = ", (a > 2) & (b > 2)) # logical_and
print("(a > 2) | (b < 3) = ", (a > 2) | (b < 3)) # logical_or
# -
# The final numpy feature we'll need is indexing: selecting elements from an array.
# Aside from python indexes and slices (e.g. a[1:4]), numpy also allows you to select several elements at once.
# +
a = np.array([0, 1, 4, 9, 16, 25])
ix = np.array([1, 2, 5])
print("a = ", a)
print("Select by element index")
print("a[[1,2,5]] = ", a[ix])
print("\nSelect by boolean mask")
# select all elementts in a that are greater than 5
print("a[a > 5] = ", a[a > 5])
print("(a % 2 == 0) =", a % 2 == 0) # True for even, False for odd
print("a[a > 3] =", a[a % 2 == 0]) # select all elements in a that are even
# select male children
print("data[(data['Age'] < 18) & (data['Sex'] == 'male')] = (below)")
data[(data['Age'] < 18) & (data['Sex'] == 'male')]
# -
# ### Your turn
#
# Use numpy and pandas to answer a few questions about data
# +
# who on average paid more for their ticket, men or women?
mean_fare_men = <YOUR CODE >
mean_fare_women = <YOUR CODE >
print(mean_fare_men, mean_fare_women)
# +
# who is more likely to survive: a child (<18 yo) or an adult?
child_survival_rate = <YOUR CODE >
adult_survival_rate = <YOUR CODE >
print(child_survival_rate, adult_survival_rate)
# -
# # Part IV: plots and matplotlib
#
# Using python to visualize the data is covered by yet another library: `matplotlib`.
#
# Just like python itself, matplotlib has an awesome tendency of keeping simple things simple while still allowing you to write complicated stuff with convenience (e.g. super-detailed plots or custom animations).
# +
import matplotlib.pyplot as plt
# %matplotlib inline
# ^-- this "magic" tells all future matplotlib plots to be drawn inside notebook and not in a separate window.
# line plot
plt.plot([0, 1, 2, 3, 4, 5], [0, 1, 4, 9, 16, 25])
# +
# scatter-plot
plt.scatter([0, 1, 2, 3, 4, 5], [0, 1, 4, 9, 16, 25])
plt.show() # show the first plot and begin drawing next one
# +
# draw a scatter plot with custom markers and colors
plt.scatter([1, 1, 2, 3, 4, 4.5], [3, 2, 2, 5, 15, 24],
c=["red", "blue", "orange", "green", "cyan", "gray"], marker="x")
# without .show(), several plots will be drawn on top of one another
plt.plot([0, 1, 2, 3, 4, 5], [0, 1, 4, 9, 16, 25], c="black")
# adding more sugar
plt.title("Conspiracy theory proven!!!")
plt.xlabel("Per capita alcohol consumption")
plt.ylabel("# Layers in state of the art image classifier")
# fun with correlations: http://bit.ly/1FcNnWF
# +
# histogram - showing data density
plt.hist([0, 1, 1, 1, 2, 2, 3, 3, 3, 3, 3, 4, 4, 5, 5, 5, 6, 7, 7, 8, 9, 10])
plt.show()
plt.hist([0, 1, 1, 1, 2, 2, 3, 3, 3, 3, 3, 4,
4, 5, 5, 5, 6, 7, 7, 8, 9, 10], bins=5)
# +
# plot a histogram of age and a histogram of ticket fares on separate plots
<YOUR CODE >
# bonus: use tab shift-tab to see if there is a way to draw a 2D histogram of age vs fare.
# +
# make a scatter plot of passenger age vs ticket fare
<YOUR CODE >
# kudos if you add separate colors for men and women
# -
# * Extended [tutorial](https://matplotlib.org/2.0.2/users/pyplot_tutorial.html)
# * A [cheat sheet](http://bit.ly/2koHxNF)
# * Other libraries for more sophisticated stuff: [Plotly](https://plot.ly/python/) and [Bokeh](https://bokeh.pydata.org/en/latest/)
# ### Part V (final): machine learning with scikit-learn
#
# <img src='https://imgs.xkcd.com/comics/machine_learning.png' width=320px>
#
# Scikit-learn is _the_ tool for simple machine learning pipelines.
#
# It's a single library that unites a whole bunch of models under the common interface:
# * Create:__ `model = sklearn.whatever.ModelNameHere(parameters_if_any)`__
# * Train:__ `model.fit(X,y)`__
# * Predict:__ `model.predict(X_test)`__
#
# It also contains utilities for feature extraction, quality estimation or cross-validation.
# +
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import accuracy_score
features = data[["Fare", "SibSp"]].copy()
answers = data["Survived"]
model = RandomForestClassifier(n_estimators=100)
model.fit(features[:-100], answers[:-100])
test_predictions = model.predict(features[-100:])
print("Test accuracy:", accuracy_score(answers[-100:], test_predictions))
# -
# Final quest: add more features to achieve accuracy of at least 0.80
#
# __Hint:__ for string features like "Sex" or "Embarked" you will have to compute some kind of numeric representation.
# For example, 1 if male and 0 if female or vice versa
#
# __Hint II:__ you can use `model.feature_importances_` to get a hint on how much did it rely each of your features.
# * Sklearn [tutorials](http://scikit-learn.org/stable/tutorial/index.html)
# * Sklearn [examples](http://scikit-learn.org/stable/auto_examples/index.html)
# * SKlearn [cheat sheet](http://scikit-learn.org/stable/_static/ml_map.png)
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
#
# Okay, what we learned: to survive a shipwreck you need to become an underaged girl with parents on the ship. Try this next time you'll find yourself in a shipwreck
| week01_intro/primer_python_for_ml/recap_ml.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Programming Languages A
# * Week 1
# * Week 2
# * Week 3
# * Week 4
# * Week 5
#
# All the weeks don't give you the actual content. It's hard to get a sneak peak unless I actually get into the course. So I'll just dive right in!
# # Week 1
# First week of my journey to learning programming languages. I just hope to view programming languages in a great way.
#
# ## Start here!
# Just a few stuff talking about this and that.
#
# ## Welcome! (And Some Course Mechanics)
# I just got the welcoming and stuff. After this course, I'll most likely be viewing programming languages in a different way. Which will make me write software in many other ways. Also, actually understand the language constructs. What a programming language needs, what a programming language is different from, why that programming language has such feature, how they differentiate from the phylosophy, etc.
# All the other things were about how the course will be and what materials we have. I didn't really get the homework part, but I think we're going to be doing a huge chunk of it for each week.
#
# ## Who I Am / Acknowledgements
# Just telling who he is and acknowledging the people that made this possible. He certainly travels a lot, has two kids, likes doing activities like biking, hockey, and etc. Maybe I should find a sport quick. I'm dying lol.
#
# ## What the Course is About / Initial Motivation
# This course mostly focuses on programming language constructs. Also it goes heavily on functional programming. It's pretty easy to say that this course is somewhat of a "believe me and you'll get it" course. The lecturer doesn't really defines what exactly we'll be learning. But he says that we'll learn the language features in a whole new way.
#
# ## Recommended Background
# Implementation vs Interface, Recursion. I think I should get a grip on these two first. Since recursion is the most essential parts of functional programming. It actually is the way to use a repeat statement. Pretty much, because everything is a expression in FP.
# This course is actually for intermediate programmers.
#
# ## Why Part A, Part B, Part C
# He didn't want to do it but it's for everyone to divide it into nice separate pieces. Part A = Part B + Part C. You need everything to proceed. Don't leave things out! It builds on top of everything!
#
# ## Grading Policy
# Submit once per day! It's not going to be easy because of the peer-grading system. Not easy to actually get 100%. But well, get an A+ for 95% over! That's what you should aim for. An A+.
#
# ## Very High-Level Outline
# * Part A
# * Basics, functions, recursions, scope, variables, tuples, lists, ...
# * Data types, pattern-matching, tail recursion
# * First-class functions, closures
# * Type inference, modules
# * Part B
# * Quick "re-do" in a dynamically typed language; Delaying evaluation
# * Implementing languages with interpreters; Static vs Dynamic typing
# * Part C
# * Dynamically-typed OOP
# * OOP vs FP decomposition
# * Advanced OOP topics (e.g. mixins, double dispatch)
# * Generics vs Subtyping
#
# # Week 2
# The real thing are in here. The start of the course! Let's get it over with. I need to learn this to actually understand functional programming and get a nice grep of Closure, Scala, or any other functional language.
#
# ## Welcome Message
# * variables
# * numbers
# * conditionals
# * scope
# * shadowing
# * functions
# * pairs
# * lists
# * let expressions
# * local bindings
# * nested functions
# * options
# * booleans
#
# Those are the sneak peaks of what I'll be learning.
#
# ## Reading Notes
# There's a cool note that the professor gave us to read! It's all in a nice PDF file!
# https://d3c33hcgiwev3.cloudfront.net/_9a49a3d3d0d3db7c72aa819869f7df33_section1sum.pdf?Expires=1562371200&Signature=QDd7X0QR2B29Y1R6IAs~JPuzgQ069ynylCMUHlhEuQiP79djhI9CFRmpGPPDnTwsl-I3L-tIdjjGGxG0kLAO-0HRQnonGzMGiZ0AQtmljBBCN<KEY>&Key-Pair-Id=<KEY>
#
# ## Code Files
# Just check the course page and you'll get it lol. All the code that's involved with this course is there.
#
# ## ML Variable Bindings and Expressions
# The peices of a programming language. A really important thing! You have to understand the static environment and dynamic environment. Finding out what the types are is the static environment, and doing the evaluation is the dynamic environment.
# Syntax and Semantics. How to write the various parts of a language is snytax. Semantics tell what those features are. So we need to learn the syntax to know how to write, and then learn the semantics to know what they do.
#
# ## Rules for Expressions
# Expressions have three rules to check. The syntax, type-checking, and the evaluation. So you need to know how to write that with the syntax, and how the type checks. And then how it evaluates into a value. After all, evaluation is actually making something into a single value.
#
# ## The REPL and Errors
# I learned that in ML you have to use tilda(~) for representing negative numbers and you have to divide with the `div` keyword. Never seen anything like that. Anyways it's cool how we get to see the syntax errors first, and then type-checking, and then evaluation errors! And we use test code to actually check whether the evaluation was right or wrong. So all this is actually a compiling thing!
#
# ## Shadowing
# There's no such thing as reassignment in SML. So it just shadows that value and the later one has the real value. I don't really understand how the other values that referenced that value becomes. Is it just caching? Well it feels like that. But still, that's it. It just doesn't do compile time errors because of convenience is what I think.
#
# ## Functions Informally
# So I got to learn how to define functions in SML. What I saw is, there's no return sign. The last thing is the thing that it returns. Also, when it needs more than one parameter, you have to put parantheses! It's so similar to Ruby. But I'm getting a hang on it. Oh and the multiply sign(*) works as a delimiter of function parameters.
#
# ## Functions Formally
# Functions are shown formally with the syntax, type-checking, and evaluation rules. Functions have two states. The definition or binding state, and the call state. Both are different syntaxes since they're different! All I got was that from this video, I should've listened to all the other steps ...
#
# ## Pairs and Other Tuples
# Pairs are just Tuples that have two! Tuples, Triples, etc. The important thing to have here is, Tuples are immutable and they can have multiple types for each item. They can also be nested. The use for tuples generally are for returning more than one value. Not like a list, but a set of values. More than one return values which is a really cool thing to note.
#
# ## Introducing Lists
# Opposite to tuples, which needs to have the same types for all elements, and it's mutable. In SML there's a lot to know. Just assign it with `[]`, add elements with `::`, get the head or tail with `hd`, `tl`, and check whether the list is empty or not with `null`. All the mentioned syntax are functions, use them well. Also you need to understand how `hd` and `tl` work. Didn't understand the recursive style of getting the element I want lol.
#
# ## List Functions
# Functions that manipulate lists. There's wasn't much to see over here except using null and recursion. I think I'll get plenty of questions in the problem set about these. Better get used to recursion lol.
#
# ## Let Expressions
# The local variable in SML. `let blah in blah end` is the syntax for it. The type checking, well it's the same as returning something. Anything. So it's pretty much a local variable, that makes a new scope of all the calculation and just returns it. It's an expression, but it could look like a function. If it does all this and that of computing instead of a function, it'll be a nice expression. The `val` can do the same as this. But the important thing to remember over here is the introduction of scope.
#
# ## Nested Functions
# You can make nested functions with let expressions. The reason nested functions are there are because of its code maintainability and cleanness. You can restrict the scope of that function and use it widely. That's the main purpose of these nested functions. You may have functions that you must use inside a function or method but there will be times that those are only used in there. It's best to use nested functions for these kinds of operations instead of making a utility function that spans across all over the program.
#
# ## Let and Efficiency
# If you do recursions wrong, you're going to do it wrong by exponentionally. So you have to be really careful when you write those kinds of functions. Cache your calculations, obviously it's dynamic programming, and analyze your function with Big(O) notation. It's getting even more important than you think. Let's get on with it.
#
# ## Options
# Optionals! Null pointers will die! Is what they're really trying to do. So the user can know that this function could give an empty value is the real important thing. Something concrete is for the user to decide.
#
# ## Booleans and Comparison Operations
# The language actually only needs an ifelse statement. The or, and, ||, && are all just helpers. In SML you use andalso, and orelse for those operations above. The `not` is actually a function over here. For more things to know is, when you do comparisons, both sides must be the same type. If not, you'll get a type-checking error. `=, <>, >, <, <=, >=` for comparison operators. The thing is, you can't use equality(=, <>) operations on real numbers. Real numbers can't be the same in computing(?) so you have to put in some error rates between. A range that you'll accept it to be true! Remember, `<>` is not equal!
#
# ## Benefits of No Mutation
# We don't need to worry whether the language is actually referring to an alias or a new object! This is something that blew my mind. If you want to append things to a list, you'll have to mutate that list, but in SML, it's not like that. It just aliases it to that value! Which means you don't use much space, and you'll only have that value in the memory. That's how they made it better. But in Java, you have to create a lot of objects and you don't know when they'll change and you have to know whether you're referring to the right object. This is a really important thing to know. Aliasing in SML.
#
# ## Java Mutation
# Now this lesson was mind blowing too! In languages that are imperative, that allow mutation, have a big problem when they give aliases(references). It's becuase, references will be always 'mutable'. If you just return a reference of an object, it'll be really easy to just mutate that object! Which is really cool to know. If you don't copy lists, or references that can be mutated, you're going to get hacked. Mutation is this dangerous! But if you're using an immutable language, you're not going to have this problem since mutation doesn't exist. Everything is rather an alias or a new value!
#
# ## Pieces of a Language
# There are five parts of learning a new language.
# 1. Syntax
# 2. Semantics
# 3. Idioms
# 4. Standard Library
# 5. Tools
#
# This syntax is just fact based. How to write this and how to write that.
# From semantics, it's different. You get to know what the syntax you wrote did. How it works in the internals, what it actually does. Idioms is how you actually typical patterns of expressing your computation. How to write this and that in a paradigm. It's either going to be structured programming, object-oriented programming, and functional programming. So we'll mostly get to learn how the language constructs can express that paradigm, idion and what it does when that particular syntax is evaluated. Evaluation rules are semantics.
#
| specifications/coursera_programming_languages_a.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from IPython.core.display import HTML
display(HTML("<style>.container { width:95% !important; }</style>"))
import pandas as pd
import numpy as np
from elasticsearch import Elasticsearch, helpers
from elasticsearch_dsl import Search, Q, SF
from bs4 import BeautifulSoup
# es = Elasticsearch(http_compress=True, maxsize=1000) # Use this to index
es = Elasticsearch()
es.ping()
wiki_dataset = pd.read_pickle('../data/dbpedia_with_articles.pkl')
plato_dataset = pd.read_pickle('../data/plato.pkl')
wiki_index = wiki_dataset[['philosopher_url', 'abstract', 'text']].copy()
plato_index = plato_dataset[['title', 'abstract', 'full_article_with_tags']].copy()
wiki_index['title'] = wiki_index['philosopher_url'].apply(lambda x: x.replace('_', ' '))
wiki_index['id'] = 'W' + wiki_index.index.astype(str)
wiki_index = wiki_index[['id','title', 'abstract', 'text']]
wiki_index.head(5)
# %%time
plato_index['text'] = plato_index['full_article_with_tags'].apply(lambda x: BeautifulSoup(x).text)
plato_index['id'] = 'P' + plato_index.index.astype(str)
plato_index = plato_index[['id','title', 'abstract', 'text']]
plato_index.head(3)
to_index = pd.concat([plato_index, wiki_index])
to_index
to_index['text'] = to_index['text'].apply(lambda x: np.nan if x==-1 else x)
to_index.to_csv('../data/to_index.csv')
to_index.to_pickle('../data/to_index.pkl')
to_index.to_pickle('../data/to_index_p4.pkl', protocol=4)
body = {
'mappings': {
'properties' : {
'title': {'type': 'text'},
'abstract': {'type': 'text'},
'article': {'type': 'text'},
}
}
}
es.indices.create(index='philosophy', body=body)
def document_generator_from_dataframe(df, index):
for _, row in df.iterrows():
row_as_dict = row.replace('', 'empty').to_dict()
yield {
"_index": index,
"_id": row['id'],
"_source": {k: row_as_dict[k] for k in ['title', 'abstract', 'text']}
}
gen = document_generator_from_dataframe(to_index[~to_index[['text', 'abstract']].isna().all(1)].replace(np.nan, 'empty'), 'philosophy')
# %%time
for success, info in helpers.parallel_bulk(es, gen, thread_count=2000,chunk_size=2500, queue_size=1000):
if not success:
print('A document failed:', info)
# +
# %%time
lookup = {"P": "the Stanford Plato Encyclopedia", "W":"the Wikipedia Philosophers Collection"}
query = "language games"
s = Search(using=es, index="philosophy")
# s.query = Q("match", title=query)
s.query = Q("multi_match", query=query, fields=['title', 'abstract', 'article'])
s = s[:20]
response = s.execute()
for hit in response:
print(f"{hit.title} - ID: {hit.meta.id} from {lookup[hit.meta.id[0]]} SCORE: {hit.meta.score}")
print("*************************************************************************************************************************")
# -
| notebooks/indexing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Measuring the Jaynes-Cummings Hamiltonian with Qiskit Pulse
#
# ### Physics Background
#
# The Jaynes-Cummings model describes a two-level system (qubit) interacting with a single-mode of an electromagnetic cavity (resonator). When a two-level system is placed in a cavity, it couples to the cavity with strength $g$, spontatnously emits its excitation with rate $\gamma$, while the cavity decays with rate $\kappa$.
# <img src="images/CQED.png" width="250"/>
# This qubit-cavity interaction can be described using the Jaynes-Cummings (JC) Hamiltonian:
#
# $H_{JC}=\omega_r(a^\dagger a) + \frac{1}{2} \omega_q \sigma_z + g (\sigma_+ a + \sigma_- a^\dagger)$
#
# Let's break down this Hamiltonian in different parts: The first part of the Hamiltonian $H_r=\omega_r(a^\dagger a)$ describes the resonator. The resonator can be treated as a quantum harmonic oscillator, where $\omega_r$ is the resonator frequency, and $a$ and $a^\dagger$ are the raising a lowering operators of the resonator photons. The next term in the JC Hamiltoninan $H_q=\frac{1}{2} \omega_q \sigma_z$ describes the qubit. Here, $\omega_q$ is the qubit frequency, and $\sigma_z$ is the Pauli-Z operator. The final term of the Hamiltonian $H_{rq}=g (\sigma_+ a + \sigma_- a^\dagger)$ describes the interaction between the resonator and the qubit: $g$ is the coupling strength between the qubit and the resonator, and the operators $\sigma_+$ and $\sigma_-$ represent exciting and de-exciting the qubit. Based on this interaction term we can see that the process of exciting a qubit leads to a photon loss in the resonator and vice-versa.
#
# In the limit that detuning between the qubit and the resonator $\Delta=\omega_q-\omega_r$ is less than the coupling strength between the two, $|\Delta|\ll g$, the resonator-qubit system becomes hybridized, leading to coherent excitation swaps which can be useful for certain two-qubit operations. However, for optimal readout, we want to operate the system in the dispersive limit, where the qubit-resonator detuning is much larger than the coupling rate and the resonator decay rate: $|\Delta| \gg g,\kappa$. In this limit the interaction between the qubit and resonator influences each of their frequencies, a feature that can be used for measuring the state of the qubit. We can apply the dispersive approximation in the limit of few photons in the resonator, and approximate the JC Hamiltonian using second-order perturbation theory as:
#
# $H_{JC(disp)}=(\omega_r+ \chi \sigma_z) a^\dagger a + \frac{1}{2} \tilde{\omega}_q \sigma_z$
#
# where $\chi=-g^2/\Delta$ is the dispersive shift (the negative sign is ue to the fact that the transmon has a negative anharmonicity), and $\tilde{\omega}_q= \omega_q+g^2/\Delta$ is the modified qubit frequency, experiencing a Lamb shift.
#
# In this tutorial we measure the parameters of the JC Hamiltonian for a system consting of a superconducting qubit coupled to a superconducting resonator using Qiskit Pulse.
# ### 0. Getting started
# We'll first get our basic dependencies and helper functions set up and ready to go.
# +
# %matplotlib inline
# Importing standard Qiskit libraries and configuring account
from qiskit import QuantumCircuit, execute, Aer, IBMQ
from qiskit.compiler import transpile, assemble
from qiskit.tools.jupyter import *
from qiskit.visualization import *
# Loading your IBM Q account(s)
provider = IBMQ.load_account()
provider = IBMQ.get_provider(hub='ibm-q-internal', group='deployed', project='default')
backend = provider.get_backend('ibmq_armonk')
backend_config = backend.configuration()
backend_defaults = backend.defaults()
# +
from scipy.optimize import curve_fit
from scipy.signal import savgol_filter
# samples need to be multiples of 16
def get_closest_multiple_of_16(num):
return int(num + 8 ) - (int(num + 8 ) % 16)
# process the reflective measurement results
def process_reflective_measurement(freqs,values):
phase_grad = np.gradient(savgol_filter(np.unwrap(np.angle(values)),3,2),freqs)
return (phase_grad-min(phase_grad))/(max(phase_grad)-min(phase_grad)) - 1
# lorentzian function
def lorentzian(f, f0, k, a, offs):
return -a*k/(2*np.pi)/((k/2)**2+(f-f0)**2)+offs
# fit a lorentizan function
def fit_lorentzian(freqs,values):
p0=[freqs[np.argmin(values)],(freqs[-1]-freqs[0])/2,min(values),0]
bounds=([freqs[0],0,-np.inf,-np.inf],[freqs[-1],freqs[-1]-freqs[0],np.inf,np.inf])
popt,pcov=curve_fit(lorentzian, freqs, values, p0=p0, bounds=bounds)
return popt,pcov
# exponential function
def exponential(t,tau,a,offset):
return a*np.exp(-t/tau)+offset
# fit an exponential function
def fit_exponential(ts,values):
p0=[np.average(ts),1,0]
return curve_fit(exponential, ts, values,p0=p0)
# -
# ### 1. Measuring $\kappa$
# Photons decay out of imperfect electromagnetic cavities. The decay rate $\kappa$ for the resonator cavity can be measured by calculating the linewidth of the resonance peak in a resonator frequency scan. Larger values of $\kappa$ imply that the resonator cavity is more lossy. The resonator loss can be quantified using the quality factor $Q=\omega_r/\kappa$; higher $Q$ indicates a lower rate of energy loss from the cavity.
# +
from qiskit import pulse # This is where we access all of our Pulse features!
from qiskit.pulse import Play, Acquire
from qiskit.pulse import pulse_lib
import numpy as np
backend_config = backend.configuration()
dt=backend_config.dt
qubit=0
readout_time = 4e-6
readout_sigma = 10e-9
# low power drive for the resonator for dispersive readout
readout_drive_low_power=pulse_lib.GaussianSquare(duration = get_closest_multiple_of_16(readout_time//dt),
amp = .3,
sigma = get_closest_multiple_of_16(readout_sigma//dt),
width = get_closest_multiple_of_16((readout_time-8*readout_sigma)//dt),
name = 'low power readout tone')
meas_chan = pulse.MeasureChannel(qubit)
acq_chan = pulse.AcquireChannel(qubit)
# readout output signal acquisition setup
acquisition_time = readout_time
acquisition=Acquire(duration = get_closest_multiple_of_16(acquisition_time//dt),
channel = acq_chan,
mem_slot = pulse.MemorySlot(0))
# +
schedule_low_power = pulse.Schedule(name='Low power resonator sweep')
schedule_low_power += Play(readout_drive_low_power, meas_chan) # apply drive pulse to the resonator channel
schedule_low_power += acquisition # aquire the output signal from the resonator readout
schedule_low_power.draw(label=True, scaling=1)
# +
center_freq = backend_defaults.meas_freq_est[qubit]
freq_span = .3e6
frequencies_range = np.linspace(center_freq-freq_span/2,center_freq+freq_span/2,41)
# list of resonator frequencies for the experiment
schedule_frequencies = [{meas_chan: freq} for freq in frequencies_range]
# +
from qiskit import assemble
from qiskit.tools.monitor import job_monitor
num_shots_per_frequency = 2*1024
frequency_sweep_low_power = assemble(schedule_low_power,
backend=backend,
meas_level=1,
meas_return='avg',
shots=num_shots_per_frequency,
schedule_los=schedule_frequencies)
job_low_power = backend.run(frequency_sweep_low_power)
job_monitor(job_low_power)
low_power_sweep_results = job_low_power.result(timeout=120)
# +
import matplotlib.pyplot as plt
low_power_sweep_values = []
for i in range(len(low_power_sweep_results.results)):
res_low_power = low_power_sweep_results.get_memory(i)
low_power_sweep_values.append(res_low_power[qubit])
low_power_sweep_values = process_reflective_measurement(frequencies_range,low_power_sweep_values)
plt.plot(frequencies_range/1e3, low_power_sweep_values,'-o', color='red', lw=2)
popt_low_power,_=fit_lorentzian(frequencies_range,low_power_sweep_values)
popt_low_power,_=fit_lorentzian(frequencies_range,low_power_sweep_values)
f0, kappa, a, offset = popt_low_power
fs=np.linspace(frequencies_range[0],frequencies_range[-1],1000)
plt.plot(fs/1e3, lorentzian(fs,*popt_low_power), color='red', ls='--')
plt.annotate("", xy=((f0-kappa/2)/1e3, offset-1/2), xytext=((f0+kappa/2)/1e3, offset-1/2), arrowprops=dict(arrowstyle="<->", color='black'))
plt.annotate("$\kappa$={:d} kHz".format(int(kappa/1e3)), xy=((f0-kappa/2)/1e3, offset-.45), color='black')
plt.grid()
plt.xlabel("Frequency [kHz]")
plt.ylabel("Measured signal [a.u.]")
plt.show()
# -
# ### 2. Measuring $\chi$ and $g$
# Next, we measure the qubit-resonator coupling. One method for measuring the dispersive shift ($\chi$) and subsequently the qubit-resonator coupling ($g=\sqrt{\chi.\Delta}$) is to compare the resonator frequency in the dispersive limit with the frequency in the non-dispersive regime, where the resonator does not get shifted by $\chi$. In the non-dispersive limit the resonator photon number $n=a^\dagger a$ is larger than $n_c=\frac{\Delta^2}{4g^2}$. In experiment we can populate the resonator with more photons by driving it with more power.
readout_drive_high_power=pulse_lib.GaussianSquare(duration = get_closest_multiple_of_16(readout_time//dt),
amp = 1, # High drive amplitude
sigma = get_closest_multiple_of_16(readout_sigma//dt),
width = get_closest_multiple_of_16((readout_time-8*readout_sigma)//dt),
name = 'high power readout tone')
# +
schedule_high_power = pulse.Schedule(name='High power resonator sweep')
schedule_high_power += Play(readout_drive_high_power, meas_chan)
schedule_high_power += acquisition
schedule_high_power.draw(label=True, scaling=1)
# +
frequency_sweep_high_power = assemble(schedule_high_power,
backend=backend,
meas_level=1,
meas_return='avg',
shots=num_shots_per_frequency,
schedule_los=schedule_frequencies)
job_high_power = backend.run(frequency_sweep_high_power)
job_monitor(job_high_power)
high_power_sweep_results = job_high_power.result(timeout=120)
# +
high_power_sweep_values = []
for i in range(len(high_power_sweep_results.results)):
res_high_power = high_power_sweep_results.get_memory(i)
high_power_sweep_values.append(res_high_power[qubit])
high_power_sweep_values = process_reflective_measurement(frequencies_range,high_power_sweep_values)
popt_high_power,_=fit_lorentzian(frequencies_range,high_power_sweep_values)
# +
plt.plot(frequencies_range/1e3, high_power_sweep_values, '-o', color='black', lw=2, label='non-dispersive')
plt.plot(frequencies_range/1e3, low_power_sweep_values,'-o', color='red', lw=2, label='dispersive')
fs=np.linspace(frequencies_range[0],frequencies_range[-1],1000)
plt.plot(fs/1e3, lorentzian(fs,*popt_high_power), color='black', ls='--')
plt.plot(fs/1e3, lorentzian(fs,*popt_low_power), color='red', ls='--')
plt.axvline(x=popt_low_power[0]/1e3, color='red')
plt.axvline(x=popt_high_power[0]/1e3, color='black')
chi=popt_low_power[0]-popt_high_power[0]
plt.annotate("", xy=(popt_low_power[0]/1e3, -.1), xytext=(popt_high_power[0]/1e3, -.1), arrowprops=dict(arrowstyle="<->", color='black'))
plt.annotate("$\chi$={:d} kHz".format(int(chi/1e3)), xy=(popt_high_power[0]/1e3, -.05), color='black')
plt.grid()
plt.xlabel("Frequency [kHz]")
plt.ylabel("Measured signal [a.u.]")
plt.legend()
plt.show()
print(r'$\chi$={:.1f} kHz'.format((popt_low_power[0]-popt_high_power[0])/1e3))
Delta=abs(backend_defaults.meas_freq_est[qubit] - backend_defaults.qubit_freq_est[qubit])
print(r'$g$=$(\chi \Delta)^.5$={:.1f} MHz'.format(np.sqrt(chi*Delta)/1e6))
# -
# ### 3. Measuring $\gamma$
# A qubit coupled to a resonator will spontaneous emit photons into the cavity, and therefore relaxing from an excited state to the ground state. The spontaneous emission of photons gets enhanced by the qubit environment, a phenomenon known as the Purcell effect. We can measure the qubit decay rate $\gamma$ by exciting the qubit with a microwave drive, and measuring the decay rate $T_1=1/\gamma$ of the qubit excitation. For this experiment our microwave drive doesn't have to be $\pi$-pulse
# +
drive_sigma = 100e-9
drive_duration = 8*drive_sigma
# qubit micrwave drive
qubit_drive = pulse_lib.gaussian(duration = get_closest_multiple_of_16(drive_duration//dt),
amp = .5,
sigma = get_closest_multiple_of_16(drive_sigma//dt),
name = 'qubit tone')
drive_chan = pulse.DriveChannel(qubit)
delay_times=np.linspace(0,600e-6,61) #measurement time delays
qubit_decay_schedules = []
for delay in delay_times:
this_schedule = pulse.Schedule(name=f"decay delay = {delay * 1e6} us")
this_schedule += Play(qubit_drive, drive_chan)
this_schedule |= Play(readout_drive_low_power, meas_chan) + acquisition << int(delay//dt)
qubit_decay_schedules.append(this_schedule)
qubit_decay_schedules[1].draw(label=True, scaling=1)
# +
# setting the readout frequency to the resontor frequecy in the dispersive limit measured earlier
# setting the qubit freqency to the default value
los = [{meas_chan: f0, drive_chan: backend_defaults.qubit_freq_est[qubit]}]
num_shots = 4*1024
qubit_decay_experiment = assemble(qubit_decay_schedules,
backend=backend,
meas_level=1,
meas_return='avg',
shots=num_shots,
schedule_los= los * len(qubit_decay_schedules))
job_qubit_decay = backend.run(qubit_decay_experiment)
job_monitor(job_qubit_decay)
# +
qubit_decay_results = job_qubit_decay.result(timeout=120)
qubit_decay_values = []
for i in range(len(delay_times)):
qubit_decay_values.append(qubit_decay_results.get_memory(i)[qubit])
qubit_decay_values = np.abs(qubit_decay_values)
qubit_decay_values = (qubit_decay_values-min(qubit_decay_values))
qubit_decay_values/=max(qubit_decay_values)
decay_popt,_=fit_exponential(delay_times, qubit_decay_values)
tau=decay_popt[0]
g=1/tau
plt.scatter(delay_times*1e6, qubit_decay_values, color='black')
plt.plot(delay_times*1e6,exponential(delay_times,*decay_popt),'--',lw=2,color='red',label=r'$\tau$={:.1f} $\mu$s'.format(tau*1e6))
plt.title("$T_1$ Experiment", fontsize=15)
plt.xlabel('Delay before measurement [$\mu$s]', fontsize=15)
plt.ylabel('Signal [a.u.]', fontsize=15)
plt.legend()
plt.show()
print(r'$\gamma$= 1/$\tau$= {:.2f} kHz'.format(g/1e3))
# -
import qiskit
qiskit.__qiskit_version__
# [1] <NAME> al. A quantum engineer’s guide to superconducting qubits. Appl. Phys. Rev. 6, 021318 (2019).
| content/ch-quantum-hardware/Jaynes-Cummings-model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/itz-kin/OOP-1-2/blob/main/OOP_Concepts.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="EAfQqsE1SEtC"
# ## Python Classes and Objects
# + [markdown] id="LtkRvjs7SWk-"
# Create a Class
# + id="-GbHi2uhSa-u"
class MyClass:
pass
# + colab={"base_uri": "https://localhost:8080/"} id="0MLWp3rOSx1w" outputId="117b03b0-5144-49ae-8b26-ee99d77fb6b0"
class OOP1_2:
X = 5
print(X)
# + [markdown] id="deCe3GQnUF_R"
# Create Objects
# + colab={"base_uri": "https://localhost:8080/"} id="O3iT5f2TS-lZ" outputId="1bae6318-7616-463f-a12f-c838e6159563"
class OOP1_2:
def __init__(self,name,age): #__init__(parameter)
self.name = name #attributes
self.age = age
def identity(self):
print(self.name, self.age)
person = OOP1_2("Kinlie", 18) #create objects
print(person.name)
print(person.age)
print(person.identity)
# + colab={"base_uri": "https://localhost:8080/"} id="Kncm-9xrWRbL" outputId="74559dec-a3c3-4141-bfe0-0af7b765dfff"
#Modify the Object Name
person.name = "Wonyoung"
person.age = 17
print(person.name)
print(person.age)
# + colab={"base_uri": "https://localhost:8080/", "height": 195} id="VjcjA-7tW5Wf" outputId="8a8cd770-e73b-4c8b-86ec-385c66656058"
#Delete the Object
del person.name
# + colab={"base_uri": "https://localhost:8080/", "height": 162} id="lRPZxCrxXE3-" outputId="34886f92-1513-4bcc-d372-65c802a38bd7"
print(person.name)
# + colab={"base_uri": "https://localhost:8080/"} id="fTGY8M7CXUcN" outputId="efe9da7d-593a-4ebb-b70e-1d0484f6d82a"
print(person.age)
# + [markdown] id="sg2GtAchXkM8"
# Application 1 - Write a Python program that computes the area of a square, and name its class as Square, side as attribute
# + colab={"base_uri": "https://localhost:8080/"} id="P2zROYLLYEkV" outputId="bafc46de-70fd-4a89-daa2-bfe6bd00aee9"
class Square:
def __init__(self,sides):
self.sides = sides
def area(self):
return self.sides*self.sides #formula to compute the area of the square
def display(self):
print("the area of the square is:",self.area())
square = Square(4)
print(square.sides)
square.display()
# + [markdown] id="W3azBF59bqr3"
# Application 2 - Write a Python program that displays your full name, age, course, school. Create a class named MyClass, and name, age, school and course as attributes
# + colab={"base_uri": "https://localhost:8080/"} id="8dUUmxqwb6q2" outputId="b44e3535-0b08-4fe9-fe05-aefb4b2f17f9"
class MyClass:
def __init__(self,name,age,school,course): #__init__(parameter)
self.name = name #attributes
self.age = age
self.school = school
self.course = course
def identity(self):
print(self.name, self.age, self.school, self.course)
person = MyClass("<NAME>", 18, "Cavite State Universty", "BS Computer Engineering") #create objects
print(person.name)
print(person.age)
print(person.school)
print(person.course)
| OOP_Concepts.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Suggest Onshore Wind Turbine
#
# * RESKit can suggest a turbine design based off site conditions (average 100m wind speed)
# * Suggestions are tailored to a far future context (~2050)
# * Since the suggestion model computes a specific capacity, a desired rotor diameter must be specified
from reskit import windpower
# +
# Get suggestion for one location
design = windpower.suggestOnshoreTurbine(averageWindspeed=6.70, # Assume average 100m wind speed is 6.70 m/s
rotordiam=136 )
print("Suggested capacity is {:.0f} kW".format( design['capacity'] ) )
print("Suggested hub height is {:.0f} meters".format( design['hubHeight'] ) )
print("Suggested rotor diamter is {:.0f} meters".format( design['rotordiam'] ) )
print("Suggested specific capacity is {:.0f} W/m2".format( design['specificPower'] ) )
# +
# Get suggestion for many locations
designs = windpower.suggestOnshoreTurbine(averageWindspeed=[6.70,4.34,5.66,4.65,5.04,4.62,4.64,5.11,6.23,5.25,],
rotordiam=136 )
designs.round()
# -
| examples/3.03-WindPower-Design_Onshore_Turbine.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Essentials
import os, sys, glob
import pandas as pd
import numpy as np
import nibabel as nib
import scipy.io as sio
# Stats
import scipy as sp
from scipy import stats
import statsmodels.api as sm
import pingouin as pg
# Plotting
import seaborn as sns
import matplotlib.pyplot as plt
plt.rcParams['svg.fonttype'] = 'none'
# -
import numpy.matlib
from sklearn.utils import resample
sys.path.append('/Users/lindenmp/Google-Drive-Penn/work/research_projects/normative_neurodev_cs_t1/1_code/')
from func import set_proj_env, get_fdr_p, run_pheno_correlations, prop_bar_plot, get_fdr_p_df, get_sys_prop, update_progress, dependent_corr, my_get_cmap
train_test_str = 'train_test'
exclude_str = 't1Exclude' # 't1Exclude' 'fsFinalExclude'
parc_str = 'schaefer' # 'schaefer' 'lausanne'
parc_scale = 400 # 200 400 | 60 125 250
parcel_names, parcel_loc, drop_parcels, num_parcels, yeo_idx, yeo_labels = set_proj_env(exclude_str = exclude_str, parc_str = parc_str, parc_scale = parc_scale)
# output file prefix
outfile_prefix = exclude_str+'_'+parc_str+'_'+str(parc_scale)+'_'
outfile_prefix
# ### Setup directory variables
# +
figdir = os.path.join(os.environ['OUTPUTDIR'], 'figs')
print(figdir)
if not os.path.exists(figdir): os.makedirs(figdir)
outputdir = os.path.join(os.environ['PIPELINEDIR'], '6_results_correlations', 'out')
print(outputdir)
if not os.path.exists(outputdir): os.makedirs(outputdir)
# -
# ## Setup plots
# +
if not os.path.exists(figdir): os.makedirs(figdir)
os.chdir(figdir)
sns.set(style='white', context = 'paper', font_scale = 1)
cmap = my_get_cmap('psych_phenos')
phenos = ['Overall_Psychopathology','Psychosis_Positive','Psychosis_NegativeDisorg','AnxiousMisery','Externalizing','Fear']
phenos_label_short = ['Ov. Psych.', 'Psy. (pos.)', 'Psy. (neg.)', 'Anx.-mis.', 'Ext.', 'Fear']
phenos_label = ['Overall psychopathology','Psychosis (positive)','Psychosis (negative)','Anxious-misery','Externalizing','Fear']
metrics = ['ct', 'vol']
metrics_label_short = ['Thickness', 'Volume']
metrics_label = ['Thickness', 'Volume']
# -
# ## Load data
load = 'test'
if load == 'all':
# Train
df_train = pd.read_csv(os.path.join(os.environ['PIPELINEDIR'], '2_prepare_normative', 'out', outfile_prefix+'train.csv'))
df_train.set_index(['bblid', 'scanid'], inplace = True)
df_node_train = pd.read_csv(os.path.join(os.environ['PIPELINEDIR'], '2_prepare_normative', 'out', outfile_prefix+'resp_train.csv'))
df_node_train.set_index(['bblid', 'scanid'], inplace = True)
# Test
df_test = pd.read_csv(os.path.join(os.environ['PIPELINEDIR'], '2_prepare_normative', 'out', outfile_prefix+'test.csv'))
df_test.set_index(['bblid', 'scanid'], inplace = True)
df_node_test = pd.read_csv(os.path.join(os.environ['PIPELINEDIR'], '2_prepare_normative', 'out', outfile_prefix+'resp_test.csv'))
df_node_test.set_index(['bblid', 'scanid'], inplace = True)
# concat
df = pd.concat((df_train, df_test), axis = 0); print(df.shape)
df_node = pd.concat((df_node_train, df_node_test), axis = 0); print(df_node.shape)
elif load == 'test':
# Test
df = pd.read_csv(os.path.join(os.environ['PIPELINEDIR'], '2_prepare_normative', 'out', outfile_prefix+'test.csv'))
df.set_index(['bblid', 'scanid'], inplace = True); print(df.shape)
df_node = pd.read_csv(os.path.join(os.environ['PIPELINEDIR'], '2_prepare_normative', 'out', outfile_prefix+'resp_test.csv'))
df_node.set_index(['bblid', 'scanid'], inplace = True); print(df_node.shape)
# ## Load nispat outputs
if load == 'all':
z_cv = np.loadtxt(os.path.join(os.environ['PIPELINEDIR'], '4_run_normative', outfile_prefix+'out_cv', 'Z.txt'), delimiter = ' ').transpose()
df_z_cv = pd.DataFrame(data = z_cv, index = df_node_train.index, columns = df_node_train.columns)
z = np.loadtxt(os.path.join(os.environ['PIPELINEDIR'], '4_run_normative', outfile_prefix+'out', 'Z.txt'), delimiter = ' ').transpose()
df_z_test = pd.DataFrame(data = z, index = df_node_test.index, columns = df_node_test.columns)
# concat
df_z = pd.concat((df_z_cv,df_z_test), axis = 0); print(df_z.shape)
elif load == 'test':
z = np.loadtxt(os.path.join(os.environ['PIPELINEDIR'], '4_run_normative', outfile_prefix+'out', 'Z.txt'), delimiter = ' ').transpose()
df_z = pd.DataFrame(data = z, index = df_node.index, columns = df_node.columns); print(df_z.shape)
# ### Regress age/sex out of psychopathology phenotypes
# +
# df_nuis = df.loc[:,['ageAtScan1_Years','sex_adj']]
# # df_nuis = df.loc[:,['ageAtScan1_Years','sex_adj','medu1']]
# # df_nuis = df.loc[:,'medu1']
# df_nuis = sm.add_constant(df_nuis)
# mdl = sm.OLS(df.loc[:,phenos], df_nuis).fit()
# y_pred = mdl.predict(df_nuis)
# y_pred.columns = phenos
# df.loc[:,phenos] = df.loc[:,phenos] - y_pred
# Note, regressing out age/sex from the deviations as well as the phenotypes makes no difference to the results
# # df_z
# cols = df_z.columns
# mdl = sm.OLS(df_z.loc[:,cols], df_nuis).fit()
# y_pred = mdl.predict(df_nuis)
# y_pred.columns = cols
# df_z.loc[:,cols] = df_z.loc[:,cols] - y_pred
# -
# ## Setup region filter
# ### regions with SMSE <1 in normative model
smse = np.loadtxt(os.path.join(os.environ['PIPELINEDIR'], '4_run_normative', outfile_prefix+'out', 'smse.txt'), delimiter = ' ').transpose()
df_smse = pd.DataFrame(data = smse, index = df_node.columns)
smse_thresh = 1
region_filter = df_smse.iloc[:,0] < smse_thresh
region_filter.sum()
# ## Get pheno-nispat relationships
# +
# region_filter = region_filter.filter(regex = 'vol')
# region_filter.sum()
# +
# # drop thickness
# df_node = df_node.filter(regex = 'vol'); print(df_node.shape)
# df_z = df_z.filter(regex = 'vol'); print(df_z.shape)
# -
# ### Regional
method = 'pearson'
assign_p = 'parametric' # 'permutation' 'parametric'
if parc_str == 'lausanne':
assign_p = 'parametric'
if assign_p == 'permutation':
nulldir = os.path.join(os.environ['NORMATIVEDIR'], 'nulls_z')
# nulldir = os.path.join(os.environ['NORMATIVEDIR'], 'nulls_z_agesex')
if not os.path.exists(nulldir): os.makedirs(nulldir)
df_pheno_z = run_pheno_correlations(df.loc[:,phenos], df_z, method = method, assign_p = assign_p, nulldir = nulldir)
elif assign_p == 'parametric':
df_pheno_z = run_pheno_correlations(df.loc[:,phenos], df_z, method = method, assign_p = assign_p)
# +
# correct multiple comparisons. We do this across brain regions and phenotypes (e.g., 400*6 = 2400 tests)
df_p_corr = pd.DataFrame(index = df_pheno_z.index, columns = ['p-corr']) # output dataframe
for metric in metrics:
p_corr = get_fdr_p(df_pheno_z.loc[:,'p'].filter(regex = metric)) # correct p-values for metric
p_corr_tmp = pd.DataFrame(index = df_pheno_z.loc[:,'p'].filter(regex = metric).index, columns = ['p-corr'], data = p_corr) # set to dataframe with correct indices
df_pheno_z.loc[p_corr_tmp.index, 'p-corr'] = p_corr_tmp # store using index matching
# -
for pheno in phenos:
for metric in metrics:
print(pheno, metric, np.sum(df_pheno_z.filter(regex = metric, axis = 0).filter(regex = pheno, axis = 0).loc[:,'p-corr'] < .05) / num_parcels * 100, '% significant effects (fdr)')
print('')
alpha = 0.05
print(alpha)
# +
x = df_pheno_z['p-corr'].values < alpha
df_pheno_z['sig'] = x
x = x.reshape(1,-1)
y = np.matlib.repmat(region_filter, 1, len(phenos))
my_bool = np.concatenate((x, y), axis = 0); region_filt = np.all(my_bool, axis = 0); df_pheno_z['sig_smse'] = region_filt
print(str(np.sum(df_pheno_z['sig'] == True)) + ' significant effects (fdr)')
print(str(np.sum(df_pheno_z['sig_smse'] == True)) + ' significant effects (fdr)')
# -
for pheno in phenos:
for metric in metrics:
# print(pheno, metric, np.sum(df_pheno_z.loc[pheno,'sig_smse'].filter(regex = metric) == True) / num_parcels * 100, '% significant effects (fdr)')
print(pheno, metric, np.sum(df_pheno_z.loc[pheno,'sig'].filter(regex = metric) == True) / num_parcels * 100, '% significant effects (fdr)')
print('')
metrics = ['vol',]
metrics_label_short = ['Volume',]
metrics_label = ['Volume',]
metric = metrics[0]; print(metric)
# +
vals = np.zeros(len(phenos))
for p, pheno in enumerate(phenos):
# vals[p] = np.sum(df_pheno_z.loc[pheno,'sig_smse'].filter(regex = metric) == True) / num_parcels * 100
vals[p] = np.sum(df_pheno_z.loc[pheno,'sig'].filter(regex = metric) == True) / num_parcels * 100
idx_perc_sig = np.argsort(vals)[::-1]
phenos_ordered = [phenos[i] for i in idx_perc_sig]
phenos_label_ordered = [phenos_label[i] for i in idx_perc_sig]
phenos_ordered
# +
sns.set(style='white', context = 'talk', font_scale = 0.8)
f, ax = plt.subplots()
f.set_figwidth(3)
f.set_figheight(3.5)
ax.barh(y = np.arange(len(phenos)), width = vals[idx_perc_sig], color = 'white', edgecolor = 'black', linewidth = 3)
ax.set_yticks(np.arange(len(phenos)))
ax.set_yticklabels(phenos_label_ordered)
ax.set_xlabel('Percentage of significant correlations')
f.savefig(outfile_prefix+'percent_sig_corrs', dpi = 100, bbox_inches = 'tight', pad_inches = 0)
# -
# ## Save out
df.to_csv(os.path.join(outputdir,outfile_prefix+'df.csv'))
df_z.to_csv(os.path.join(outputdir,outfile_prefix+'df_z.csv'))
df_pheno_z.to_csv(os.path.join(outputdir,outfile_prefix+'df_pheno_z.csv'))
region_filter.to_csv(os.path.join(outputdir,outfile_prefix+'region_filter.csv'))
# # Plots
import matplotlib.image as mpimg
from brain_plot_func import roi_to_vtx, brain_plot
if parc_str == 'schaefer':
subject_id = 'fsaverage'
elif parc_str == 'lausanne':
subject_id = 'lausanne125'
# ## a-priori regions of interest
# Schaefer 200
if parc_scale == 200:
dacc_strs = ['17Networks_LH_SalVentAttnB_PFCmp_1', '17Networks_LH_ContA_Cinga_1',
'17Networks_RH_SalVentAttnB_PFCmp_1', '17Networks_RH_ContA_Cinga_1', '17Networks_RH_DefaultA_PFCm_2'] # daCC
mofc_strs = ['17Networks_LH_Limbic_OFC_1', '17Networks_LH_Limbic_OFC_2', '17Networks_LH_DefaultA_PFCm_1',
'17Networks_RH_Limbic_OFC_1', '17Networks_RH_Limbic_OFC_2', '17Networks_RH_Limbic_OFC_3'] # vmPFC/mOFC
insula_strs = ['17Networks_LH_SalVentAttnA_Ins_1', '17Networks_LH_SalVentAttnA_Ins_2', '17Networks_LH_SalVentAttnA_Ins_3',
'17Networks_RH_SalVentAttnA_Ins_1', '17Networks_RH_SalVentAttnA_Ins_2', '17Networks_RH_SalVentAttnA_Ins_3']
inftemp_strs = ['17Networks_LH_Limbic_TempPole_1', '17Networks_LH_Limbic_TempPole_2', '17Networks_LH_Limbic_TempPole_3', '17Networks_LH_Limbic_TempPole_4',
'17Networks_LH_ContB_Temp_1',
'17Networks_LH_DefaultB_Temp_1', '17Networks_LH_DefaultB_Temp_2', '17Networks_LH_DefaultB_Temp_3', '17Networks_LH_DefaultB_Temp_4',
'17Networks_RH_Limbic_TempPole_1', '17Networks_RH_Limbic_TempPole_2', '17Networks_RH_Limbic_TempPole_3','17Networks_RH_Limbic_TempPole_4',
'17Networks_RH_ContB_Temp_1', '17Networks_RH_ContB_Temp_2',
'17Networks_RH_DefaultB_Temp_1', '17Networks_RH_DefaultB_AntTemp_1']
mask_strs = {'mOFC': mofc_strs,
'Inf. temporal': inftemp_strs,
'daCC': dacc_strs,
'Insula': insula_strs}
elif parc_scale == 400:
dacc_strs = ['17Networks_LH_SalVentAttnB_PFCmp_1', '17Networks_LH_DefaultA_PFCm_6', '17Networks_LH_ContA_Cinga_1',
'17Networks_RH_SalVentAttnB_PFCmp_1', '17Networks_RH_SalVentAttnB_PFCmp_2', '17Networks_RH_DefaultA_PFCm_6', '17Networks_RH_ContA_Cinga_1'] # daCC
mofc_strs = ['17Networks_LH_Limbic_OFC_1', '17Networks_LH_Limbic_OFC_2', '17Networks_LH_Limbic_OFC_3', '17Networks_LH_Limbic_OFC_4', '17Networks_LH_Limbic_OFC_5', '17Networks_LH_SalVentAttnB_OFC_1',
'17Networks_RH_Limbic_OFC_1', '17Networks_RH_Limbic_OFC_2', '17Networks_RH_Limbic_OFC_3', '17Networks_RH_Limbic_OFC_4', '17Networks_RH_Limbic_OFC_5', '17Networks_RH_Limbic_OFC_6'] # vmPFC/mOFC
insula_strs = ['17Networks_LH_SalVentAttnA_Ins_1', '17Networks_LH_SalVentAttnA_Ins_2', '17Networks_LH_SalVentAttnA_Ins_3',
'17Networks_LH_SalVentAttnA_Ins_4', '17Networks_LH_SalVentAttnA_Ins_5', '17Networks_LH_SalVentAttnA_Ins_6'
'17Networks_RH_SalVentAttnA_Ins_1', '17Networks_RH_SalVentAttnA_Ins_2', '17Networks_RH_SalVentAttnA_Ins_3', '17Networks_RH_SalVentAttnA_Ins_4',
'17Networks_RH_SalVentAttnA_Ins_5', '17Networks_RH_SalVentAttnA_Ins_6', '17Networks_RH_SalVentAttnA_Ins_7']
inftemp_strs = ['17Networks_LH_Limbic_TempPole_1', '17Networks_LH_Limbic_TempPole_2', '17Networks_LH_Limbic_TempPole_3', '17Networks_LH_Limbic_TempPole_4',
'17Networks_LH_Limbic_TempPole_5', '17Networks_LH_Limbic_TempPole_6', '17Networks_LH_Limbic_TempPole_7',
'17Networks_LH_ContB_Temp_1',
'17Networks_LH_DefaultB_Temp_1', '17Networks_LH_DefaultB_Temp_2', '17Networks_LH_DefaultB_Temp_3', '17Networks_LH_DefaultB_Temp_4', '17Networks_LH_DefaultB_Temp_5', '17Networks_LH_DefaultB_Temp_6',
'17Networks_RH_Limbic_TempPole_1', '17Networks_RH_Limbic_TempPole_2', '17Networks_RH_Limbic_TempPole_3',
'17Networks_RH_Limbic_TempPole_4', '17Networks_RH_Limbic_TempPole_5', '17Networks_RH_Limbic_TempPole_6',
'17Networks_RH_ContB_Temp_1', '17Networks_RH_ContB_Temp_2',
'17Networks_RH_DefaultA_Temp_1', '17Networks_RH_DefaultB_Temp_1', '17Networks_RH_DefaultB_Temp_2', '17Networks_RH_DefaultB_AntTemp_1']
mask_strs = {'mOFC': mofc_strs,
'Inf. temporal': inftemp_strs,
'daCC': dacc_strs,
'Insula': insula_strs}
# +
mask = np.zeros(parcel_names.shape).astype(bool)
for roi in mask_strs.keys():
# create mask
mask = np.zeros(parcel_names.shape).astype(bool)
for i, mask_str in enumerate(mask_strs[roi]):
mask[np.where(parcel_names == mask_str)[0]] = True
mask = mask.astype(float)
mask[mask == False] = -1000
for hemi in ['lh', 'rh']:
if subject_id == 'lausanne125':
parc_file = os.path.join('/Applications/freesurfer/subjects/', subject_id, 'label', hemi + '.myaparc_' + str(parc_scale) + '.annot')
elif subject_id == 'fsaverage':
parc_file = os.path.join('/Users/lindenmp/Google-Drive-Penn/work/research_projects/normative_neurodev_cs_t1/figs_support/Parcellations/FreeSurfer5.3/fsaverage/label/',
hemi + '.Schaefer2018_' + str(parc_scale) + 'Parcels_17Networks_order.annot')
# project subject's data to vertices
brain_plot(mask, parcel_names, parc_file, hemi+'_'+roi, subject_id = subject_id, hemi = hemi, surf = 'inflated', color = 'coolwarm', center_anchor = 3)
# -
mask_strs.keys()
# +
r_vals = pd.DataFrame(index = phenos, columns = mask_strs.keys())
p_vals = pd.DataFrame(index = phenos, columns = mask_strs.keys())
for pheno in phenos:
for roi in mask_strs.keys():
# create mask
mask = np.zeros(parcel_names.shape).astype(bool)
for mask_str in mask_strs[roi]:
mask[np.where(parcel_names == mask_str)[0]] = True
x = df_z.filter(regex = metric).loc[:,mask].mean(axis = 1) # get region average deviations
y = df.loc[:,pheno]
r_vals.loc[pheno,roi] = sp.stats.pearsonr(x,y)[0]
p_vals.loc[pheno,roi] = sp.stats.pearsonr(x,y)[1]
p_vals = get_fdr_p_df(p_vals)
r_vals.index = phenos_label
# -
np.round(r_vals.astype(float),2)
p_vals<.05
# +
sns.set(style='white', context = 'paper', font_scale = 1)
f, ax = plt.subplots()
f.set_figwidth(3)
f.set_figheight(3)
mask = np.zeros_like(r_vals)
mask[p_vals>=.05] = True
sns.heatmap(r_vals.astype(float), mask=mask, center=0, cmap='coolwarm', annot=True, linewidth=.1, square=True, ax=ax)
ax.set_xticklabels(ax.get_xticklabels(), rotation=45)
ax.tick_params(pad = -6)
f.savefig(outfile_prefix+'roi_correlations.png', dpi = 300, bbox_inches = 'tight')
# -
if os.path.exists(os.path.join(outputdir,outfile_prefix+'r_bs.npy')):
r_bs = np.load(os.path.join(outputdir,outfile_prefix+'r_bs.npy'))
else: # otherwise, compute and save it out
n_boot = 10000
r_bs = np.zeros((n_boot, len(mask_strs), len(phenos)-1))
for i in np.arange(n_boot):
update_progress(i/n_boot)
for j, roi in enumerate(mask_strs.keys()):
# create mask
mask = np.zeros(parcel_names.shape).astype(bool)
for mask_str in mask_strs[roi]:
mask[np.where(parcel_names == mask_str)[0]] = True
x = df_z.filter(regex = metric).loc[:,mask].mean(axis = 1) # get region average deviations
x_bs, df_bs = resample(x, df, n_samples = int(x.shape[0]), random_state = i, replace=True) # get boot sample
xy = np.abs(sp.stats.pearsonr(x_bs, df_bs.loc[:,phenos[0]])[0]) # correlation between deviations (x) and phenotype (y)
for k, pheno in enumerate(phenos[1:]):
xz = np.abs(sp.stats.pearsonr(x_bs, df_bs.loc[:,pheno])[0]) # correlation between deviations (x) and phenotype (z)
d = xy-xz
r_bs[i,j,k] = d
update_progress(1)
np.save(os.path.join(outputdir,outfile_prefix+'r_bs'), r_bs)
alpha = 0.05/r_bs.shape[2]
# alpha = 0.05
print(alpha)
print(alpha/2*100)
print(100-alpha/2*100)
for j, roi in enumerate(mask_strs.keys()):
for k, pheno in enumerate(phenos[1:]):
lower = np.percentile(r_bs[:,j,k], alpha/2*100)
upper = np.percentile(r_bs[:,j,k], 100-alpha/2*100)
if lower > 0:
print(roi,pheno,np.mean(r_bs[:,j,k]),lower,upper)
# +
sns.set(style='white', context = 'paper', font_scale = 1)
cmap = my_get_cmap('psych_phenos')
cmap = cmap[1:]
for j, roi in enumerate(mask_strs.keys()):
print(roi)
f, ax = plt.subplots(len(phenos)-1,1)
f.set_figwidth(2)
f.set_figheight(3)
for k, pheno in enumerate(phenos[1:]):
lower = np.percentile(r_bs[:,j,k], alpha/2*100)
upper = np.percentile(r_bs[:,j,k], 100-alpha/2*100)
if lower>0:
sns.kdeplot(x=r_bs[:,j,k], ax=ax[k], bw_adjust=.75, clip_on=False, color=cmap[k], alpha=0.75, linewidth=0, fill=True)
# sns.kdeplot(x=r_bs[:,j,k], ax=ax[k], bw_adjust=.75, clip_on=False, color="gray", alpha=1, linewidth=2)
# add point estimate
ax[k].axvline(x=r_bs[:,j,k].mean(), ymax=0.25, clip_on=False, color='w', linewidth=2)
else:
sns.kdeplot(x=r_bs[:,j,k], ax=ax[k], bw_adjust=.75, clip_on=False, color=cmap[k], alpha=0.2, linewidth=0, fill=True)
ax[k].axvline(x=r_bs[:,j,k].mean(), ymax=0.25, clip_on=False, color='w', linewidth=1)
ax[k].set_xlim([-.2, .3])
ax[k].axhline(y=0, linewidth=2, clip_on=False, color=cmap[k])
ax[k].axvline(x=0, ymax=1, clip_on=False, color='gray', linestyle='--', linewidth=1.5)
for spine in ax[k].spines.values():
spine.set_visible(False)
ax[k].set_ylabel('')
ax[k].set_yticklabels([])
ax[k].set_yticks([])
# if k != len(phenos)-2:
# ax[k].set_xticklabels([])
ax[k].set_xlabel('')
ax[k].tick_params(pad = -2)
if j == 0:
ax[k].text(0, .75, phenos_label[1:][k], fontweight="regular", color=cmap[k],
ha="left", va="center", transform=ax[k].transAxes)
ax[-1].set_xlabel('Pearson''s r (delta)')
f.subplots_adjust(hspace=1)
# f.suptitle(roi)
f.savefig(outfile_prefix+'r_bs_'+roi.replace('. ','_')+'.svg', dpi = 600, bbox_inches = 'tight')
# -
# ## Figure 3
# +
figs_to_delete = []
for pheno in phenos:
for metric in metrics:
for hemi in ('lh', 'rh'):
print(pheno, metric)
# Plots of univariate pheno correlation
fig_str = hemi + '_' + pheno + '_' + metric + '_z'
figs_to_delete.append('ventral_'+fig_str)
figs_to_delete.append('med_'+fig_str)
figs_to_delete.append('lat_'+fig_str)
roi_data = df_pheno_z.loc[pheno].filter(regex = metric, axis = 0)['coef'].values
# sig = df_pheno_z.loc[pheno].filter(regex = metric, axis = 0)['sig_smse']
sig = df_pheno_z.loc[pheno].filter(regex = metric, axis = 0)['sig']
roi_data[~sig] = -1000
if subject_id == 'lausanne125':
parc_file = os.path.join('/Applications/freesurfer/subjects/', subject_id, 'label', hemi + '.myaparc_' + str(parc_scale) + '.annot')
elif subject_id == 'fsaverage':
parc_file = os.path.join('/Users/lindenmp/Google-Drive-Penn/work/research_projects/normative_neurodev_cs_t1/figs_support/Parcellations/FreeSurfer5.3/fsaverage/label/',
hemi + '.Schaefer2018_' + str(parc_scale) + 'Parcels_17Networks_order.annot')
# project subject's data to vertices
brain_plot(roi_data, parcel_names, parc_file, fig_str, subject_id = subject_id, hemi = hemi, surf = 'inflated', center_anchor = 0.2)
# -
for pheno in phenos:
for metric in metrics:
f, axes = plt.subplots(3, 2)
f.set_figwidth(3)
f.set_figheight(5)
plt.subplots_adjust(wspace=0, hspace=-0.465)
print(pheno)
print(metric)
# column 0:
fig_str = 'lh_'+pheno+'_'+metric+'_z.png'
try:
image = mpimg.imread('ventral_' + fig_str); axes[2,0].imshow(image); axes[2,0].axis('off')
except FileNotFoundError: axes[2,0].axis('off')
try:
image = mpimg.imread('med_' + fig_str); axes[1,0].imshow(image); axes[1,0].axis('off')
except FileNotFoundError: axes[1,0].axis('off')
try:
# axes[0,0].set_title('Thickness (left)')
image = mpimg.imread('lat_' + fig_str); axes[0,0].imshow(image); axes[0,0].axis('off')
except FileNotFoundError: axes[0,0].axis('off')
# column 1:
fig_str = 'rh_'+pheno+'_'+metric+'_z.png'
try:
# axes[0,1].set_title('Thickness (right)')
image = mpimg.imread('lat_' + fig_str); axes[0,1].imshow(image); axes[0,1].axis('off')
except FileNotFoundError: axes[0,1].axis('off')
try:
image = mpimg.imread('med_' + fig_str); axes[1,1].imshow(image); axes[1,1].axis('off')
except FileNotFoundError: axes[1,1].axis('off')
try:
image = mpimg.imread('ventral_' + fig_str); axes[2,1].imshow(image); axes[2,1].axis('off')
except FileNotFoundError: axes[2,1].axis('off')
plt.show()
f.savefig(outfile_prefix+metric+'_'+pheno+'_z.svg', dpi = 600, bbox_inches = 'tight', pad_inches = 0)
for file in figs_to_delete:
try:
os.remove(os.path.join(figdir,file+'.png'))
except:
print(file, 'not found')
| 1_code/6_results_correlations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Visualization Case Study
#
# ## Information on the Diamond Dataset
# In this lesson, you'll be working with a dataset regarding the prices and attributes of approximately 54,000 round-cut diamonds. You'll go through the steps of an explanatory data visualization, systematically starting from univariate visualizations, moving through bivariate visualizations, and finally multivariate visualizations. Finally, you'll work on polishing up selected plots from the analysis so that their main points can be clearly conveyed to others.
#
# You can find a copy of the dataset in the Resources tab of the classroom; it will automatically be available to you in the workspaces of this lesson. The dataset consists of almost 54,000 rows and 10 columns:
#
# - price: Price in dollars. Data was collected in 2008.
# - carat: Diamond weight. 1 carat is equal to 0.2 grams.
# - cut: Quality of diamond cut, affects its shine. Grades go from (low) Fair, Good, Very Good, Premium, Ideal (best).
# - color: Measure of diamond coloration. Increasing grades go from (some color) J, I, H, G, F, E, D (colorless).
# - clarity: Measure of diamond inclusions. Increasing grades go from (inclusions) I1, SI2, SI1, VS2, VS1, VVS2, VVS1, IF (internally flawless).
# - x, y, z: Diamond length, width, and depth, respectively, in mm.
# - table: Ratio of width of top face of diamond to its overall width, as a percentage.
# - depth: Proportional depth of diamond, as a percentage. This is computed as 2 * z / (x + y), or the ratio of the depth to the average of length and width.
#
# For the case study, we will concentrate only the variables in the top five bullet points: price and the four 'C's of diamond grade. Our focus will be on answering the question about the degree of importance that each of these quality measures has on the pricing of a diamond. You can see an example report covering all of the variables in the project information lesson.
# +
# import all packages and set plots to be embedded inline
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sb
# %matplotlib inline
# -
# load in the dataset into a pandas dataframe
diamonds = pd.read_csv('diamonds.csv')
# ## Dataset Overview
#
# Before running straight into the exploration, let's just take a moment to get a high-level overview of the dataset. We can refer back to these points if we need to get our bearings on what we should expect to see from each variable.
# high-level overview of data shape and composition
print(diamonds.shape)
print(diamonds.dtypes)
diamonds.head(10)
# descriptive statistics for numeric variables
diamonds.describe()
# ## Univariate Exploration
#
# Let's start our exploration by looking at the main variable of interest: price. Is the distribution skewed or symmetric? Is it unimodal or multimodal?
np.log10(diamonds.price).describe()
# +
# Get a histogram of base 10 log price
bins = 10**np.arange(2.5,4.3+0.05,0.05)
ticks = [300,1000,3000,10000,30000]
labels = ['{}'.format(v) for v in ticks]
plt.hist(data = diamonds, x = 'price',bins = bins)
plt.xscale('log')
plt.xticks(ticks,labels);
# -
# Under a log scale, I see that the data is roughly bimodal, with one large peak somewhere between 500 and 1000, and a second large peak between 4 and 5 thousand. The price distribution also seems to just cut off at its maximum, rather than declining in a smooth tail
#
# Next, create a plot of our first diamond 'C' metric: carat weight. Is there an interesting pattern in these values?
# univariate plot of carat weights
plt.figure(figsize= [8,8])
bins = np.arange(0.2,5+0.01,0.01)
ticks = [i/5 for i in range(20)]
labels = ['{}'.format(v) for v in ticks]
plt.hist(data = diamonds, x = 'carat',bins =bins)
plt.xlim(0,3)
plt.xticks(ticks,labels)
plt.xticks(rotation = 90);
# I see an interesting pattern in carat weights, where there is a large spike around a single decimal value, then a gradual tailing off to the right until the next spike. In this case, it's a good idea to set the bin size small (e.g. 0.01) and zoom into a narrower range of values to clearly see the spikes in the data values. Perhaps these spikes come about as part of standard diamond size conventions?
#
# Now, I will move on to exploring the other three 'C' quality measures: cut, color, and clarity. For each of these measures, does the data we have tend to be higher on the quality scale, or lower?
# +
# univariate plots of cut, color, and clarity grades
base_color = sb.color_palette()[0]
cut_order = ['Fair', 'Good', 'Very Good', 'Premium', 'Ideal']
ordered_cut_type = pd.api.types.CategoricalDtype(ordered = True, categories = cut_order)
diamonds['cut'] = diamonds.cut.astype(ordered_cut_type)
sb.countplot(data = diamonds, x = 'cut', color = base_color)
# +
color_order = ['J', 'I', 'H', 'G', 'F', 'E', 'D']
ordered_color_type = pd.api.types.CategoricalDtype(ordered = True, categories = color_order)
diamonds['color'] = diamonds.color.astype(ordered_color_type)
sb.countplot(data = diamonds, x = 'color', color = base_color)
# +
clarity_order = ['I1', 'SI2', 'SI1', 'VS2', 'VS1', 'VVS2', 'VVS1', 'IF']
ordered_clarity_type = pd.api.types.CategoricalDtype(ordered = True, categories = clarity_order)
diamonds['clarity'] = diamonds.clarity.astype(ordered_clarity_type)
sb.countplot(data = diamonds, x = 'clarity', color = base_color)
# -
# The bar chart should show that the mode cut quality for diamonds in this dataset is the highest grade, Ideal. There is a consistent decrease in number of diamonds of lower cut grades.
#
# The distribution is roughly unimodal, with the most common color grade in this dataset belonging to the central grade represented, G. (There are other diamond grades that are worse than J, but they're not part of the collected data.)
#
# There is a right-skew in the clarity grades represented in the dataset, with most of the diamonds collected having slight or very slight inclusions (SI2, SI1, VS2, VS1). There are very few diamonds with visible inclusions.
# ```
# # convert cut, color, and clarity into ordered categorical types
# ordinal_var_dict = {'cut': ['Fair','Good','Very Good','Premium','Ideal'],
# 'color': ['J', 'I', 'H', 'G', 'F', 'E', 'D'],
# 'clarity': ['I1', 'SI2', 'SI1', 'VS2', 'VS1', 'VVS2', 'VVS1', 'IF']}
#
# for var in ordinal_var_dict:
# pd_ver = pd.__version__.split(".")
# print
# if (int(pd_ver[0]) > 0) or (int(pd_ver[1]) >= 21): # v0.21 or later
# ordered_var = pd.api.types.CategoricalDtype(ordered = True,
# categories = ordinal_var_dict[var])
# diamonds[var] = diamonds[var].astype(ordered_var)
# else: # pre-v0.21
# diamonds[var] = diamonds[var].astype('category', ordered = True,
# categories = ordinal_var_dict[var])
# ```
# ## Bivariate Exploration
#
# In the previous notebook, you looked at the univariate distribution of five features in the diamonds dataset: price, carat, cut, color, and clarity. Now, we'll investigate relationships between pairs of these variables, particularly how each of them relate to diamond price.
#
# To start, construct a plot of the price against carat weight. What kind of shape does the relationship between these variables take?
ticks = [300,1000,3000,10000,30000]
labels = ['{}'.format(v) for v in ticks]
plt.scatter(data = diamonds, x = 'carat', y= 'price',alpha = 1/100)
plt.yscale('log')
plt.yticks(ticks,labels);
# Assuming that you put price on the y-axis and carat on the x-axis, the transformation should have changed the scatterplot from looking concave upwards (like an upwards-pointing bowl) to concave downards (like a downwards-pointing bowl). It would be nice to have a linear trend, wouldn't it? It turns out that the x, y, and z dimensions track a much more linear trend against price than carat does. Since carat is more recognizable a feature value, let's add a transformation to the x-axis to see the approximately linear trend. Since weight is proportional to volume, and since all the diamonds are cut in approximately the same way (round-cut), a cube-root transformation of carat should allow us to see an approximately linear trend.
# +
ticks = [300,1000,3000,10000,30000]
labels = ['{}'.format(v) for v in ticks]
plt.scatter(diamonds['carat']**(1/3),diamonds.price,alpha = 1/100)
plt.yscale('log')
plt.yticks(ticks,labels)
plt.title('Scatter plot of carat and price')
plt.xlabel('Cubic root of Carat')
plt.ylabel('price');
# -
# One thing that you might notice in the visualization after transformation is that the trend between price and carat weight is 'cut off' by the maximum price limit of diamonds available in the dataset. For diamonds above about 1.5 carats, we'd probably expect some diamonds about $20,000 in price, given the trend below 1.5 carats. This would be a point to potentially pay attention to later on with interaction effects between other factors, and when building a model of diamond prices.
# Now let's take a look at the relationship between price and the three categorical quality features, cut, color, and clarity. Are there any surprising trends to be seen here?
# +
# box plot of price and cut
sb.boxplot(data = diamonds, x = 'cut', y = 'price', color = base_color);
# +
# box plot of price and color
sb.boxplot(data = diamonds, x = 'color', y = 'price', color = base_color);
# +
# box plot of price and clarity
sb.boxplot(data = diamonds, x = 'clarity', y = 'price', color = base_color);
# -
# There's a lot of different approaches you could have taken here. If you created a box plot, then you probably noticed something interesting and unintuitive: the median price of diamonds should tend to look like it _decrease_ with _increasing_ gem quality. Shouldn't we expect this to be the other way around? This deserves a deeper look. Is there a different plot type to choose that shows more details in the data?
# +
# violin plot of price and cut
sb.violinplot(data = diamonds, x = 'cut', y = 'price', color = base_color);
# +
# violin plot of price and color
sb.violinplot(data = diamonds, x = 'color', y = 'price', color = base_color);
# +
# violin plot of price and clarity
sb.violinplot(data = diamonds, x = 'clarity', y = 'price', color = base_color);
# -
# With a violin plot, you can get more insight into what causes the trend in median prices to appear as it does. Faceted histograms will also produce a similar result, though unless the faceting keeps the price axis common across facets, the trend will be harder to see. For each ordinal variable, there are multiple modes into which prices appear to fall. Going across increasing quality levels, you should see that the modes rise in price - this should be the expected effect of quality. However, you should also see that more of the data will be located in the lower-priced modes - this explains the unintuitive result noted in the previous comment. This is clearest in the clarity variable. Let's keep searching the data to see if there's more we can say about this pattern.
# Before we complete the bivariate exploration, we should take a look at some of the relationships between the independent variables. Namely, how does carat weight change across the other three 'C' quality measures? You can take the same approach as the above investigation of price against the three categorical quality measures. What can you see in these plots, and how does this relate to your earlier investigations?
# +
# violin plot of carat and cut
sb.violinplot(data = diamonds, x = 'cut', y = 'carat', color = base_color);
# +
# violin plot of carat and color
sb.violinplot(data = diamonds, x = 'color', y = 'carat', color = base_color);
# +
# violin plot of carat and clarity
sb.violinplot(data = diamonds, x = 'clarity', y = 'carat', color = base_color);
# -
# These plots should shed a lot more light into the patterns already seen. Larger diamonds are more likely to receive lower categorical quality grades, while high quality grades are more likely to have their numbers made up of smaller diamonds. Since carat weight appears to be a major driver of price, this helps to explain the surprising marginal effect of diamond quality against price. The challenge next will be on how to depict this in a multivariate plot.
# ## Multivariate Exploration
#
# In the previous workspace, you looked at various bivariate relationships. You saw that the log of price was approximately linearly related to the cube root of carat weight, as analogy to its length, width, and depth. You also saw that there was an unintuitive relationship between price and the categorical quality measures of cut, color, and clarity, that the median price decreased with increasing quality. Investigating the distributions more clearly and looking at the relationship between carat weight with the three categorical variables showed that this was due to carat size tending to be smaller for the diamonds with higher categorical grades.
#
# The goal of this workspace will be to depict these interaction effects through the use of multivariate plots.
#
# To start off with, create a plot of the relationship between price, carat, and clarity. In the previous workspace, you saw that clarity had the clearest interactions with price and carat. How clearly does this show up in a multivariate visualization?
diamonds['cub_root_carat'] = diamonds['carat']**(1/3)
diamonds.head()
# +
# multivariate plot of price by carat weight, and clarity
ticks = [300,1000,3000,10000,30000]
labels = ['{}'.format(v) for v in ticks]
g = sb.FacetGrid(data = diamonds, col = 'clarity',col_wrap = 3)
g.map(plt.scatter,'cub_root_carat','price',alpha = 1/30)
plt.yscale('log')
plt.ylim(300,30000)
plt.yticks(ticks,labels);
# -
# You should see across facets the general movement of the points upwards and to the left, corresponding with smaller diamond sizes, but higher value for their sizes. As a final comment, did you remember to apply transformation functions to the price and carat values?
#
# Let's try a different plot, for diamond price against cut and color quality features. To avoid the trap of higher quality grades being associated with smaller diamonds, and thus lower prices, we should focus our visualization on only a small range of diamond weights. For this plot, select diamonds in a small range around 1 carat weight. Try to make it so that your plot shows the effect of each of these categorical variables on the price of diamonds.
one_carat = diamonds.query('carat > 0.9 and carat < 1.1')
one_carat
# +
# multivariate plot of price by cut and color, for approx. 1 carat diamonds
plt.figure(figsize = [10,8])
ticks = [300,1000,3000,10000,30000]
labels = ['{}'.format(v) for v in ticks]
sb.pointplot(data = one_carat, x = 'color' , y = 'price' ,hue = 'cut',
ci = 'sd', linestyles = '', palette = 'viridis_r',dodge = 0.3)
plt.yscale('log')
plt.yticks(ticks,labels)
plt.ylim(1000,30000);
# -
# Assuming you went with a clustered plot approach, you should see a gradual increase in price across the main x-value clusters, as well as generally upwards trends within each cluster for the third variable. Aesthetically, did you remember to choose a sequential color scheme for whichever variable you chose for your third variable, to override the default qualitative scheme? If you chose a point plot, did you set a dodge parameter to spread the clusters out?
# ## Explanatory Polishing
#
# Through the last few workbooks, you've performed an exploration of a dataset relating the price of diamonds to their four 'C' attributes: carat, cut, color, and clarity. During that exploration, you found that the primary driver of price was carat weight, where the log of price was approximately linearly related to the cube root of carat weight, analogous to the diamond's length, width, or depth. The price was then modified by the diamond's cut, color, and clarity grades.
#
# In this workbook, you'll polish two plots from the earlier exploration so that they can be presented to others in an understandable form.
#
# The first plot to polish depicts the base relationship between price and carat. Make additions and revisions to the code below to refine the given plot.
# +
plt.figure(figsize = [8, 6])
ticks1 = [500,1000,2000,5000,10000,20000]
labels1 = ['{}'.format(v) for v in ticks1]
ticks2 = [0.2,0.3,0.4,0.5,0.7,1.0,1.5,2.0,2.5,3]
labels2 = ['{:0.1f}'.format(v) for v in ticks2]
new = []
for i in ticks2:
new.append(i**(1/3))
ticks2 = new
plt.scatter(diamonds.cub_root_carat,diamonds.price,alpha = 1/50)
# plt.grid(axis = 'x', which = 'major')
plt.yscale('log')
plt.yticks(ticks1,labels1)
plt.ylim(300,)
plt.xlim(0.15**(1/3),3.1**(1/3))
plt.xticks(ticks2,labels2,rotation = 45)
plt.title('Scatter plot of carat and price')
plt.xlabel('Carat')
plt.ylabel('price')
plt.show();
# -
# - Does the plot has an informative title?
# - Are the axis labels informative?
# - Are the tick values interpretable?
# - Do we need to worry about any of the plot aesthetics?
#
# - The original plot's vertical axis only has two tick marks, and they're in scientific notation. Not only is it not clear that the data is on a power scale, it's also difficult to parse. We can solve both of these problems by specifying the tick marks. Take advantage of the 1-2-5-10 sequence to set which tick marks to show.
# - As for the horizontal axis, the values are in terms of the cube-root transformed values, rather than in raw carat values. This presents an obstacle to interpretability. Once again, we should specify tick locations: the peaks observed in the univariate distribution might be good choices. Note that we also need to make use of the cuberoot_trans() function since the transformation is non-standard.
# - There's a lot of overplotting in the scatterplot as given. Try playing with the alpha parameter to try and make the main trend stand out more.
#
# The second plot that you will polish is a multivariate plot of price against levels of color and cut, for diamonds of about 1 carat weight. Make additions and revisions to the code below to refine the given plot.
# +
# select diamonds of approximately 1 carat
one_carat = diamonds.query('carat >= 0.99 and carat <= 1.03')
plt.figure(figsize = [8,6])
ticks = [2000,3000,5000,8000]
labels = ['2k','3k','5k','8k']
ax = sb.pointplot(data = one_carat, x = 'color' , y = 'price' ,hue = 'cut',
linestyles = '', palette = 'Blues',dodge = 0.5)
plt.title('Price on Color and Cut (dimands around 1 carat)')
plt.xlabel('Diamond Color')
plt.yscale('log')
plt.yticks(ticks,labels)
plt.ylabel('Diamond Price')
plt.ylim(2000,10000)
ax.set_yticklabels([], minor = True)
;
# -
# - Don't forget to provide an informative title and axis labels.
# - There's a few things with the pointplot function that we can revise. First of all, the default color palette used is qualitative, but we should probably choose a sequential palette instead. We could also stand to add a dodge parameter so that the points aren't stacked directly on top of one another and to make individual points easier to read. One other possible modification we could add is to remove the line connecting points on each level of cut: with the other changes made, removing the connecting line can help reduce the ink used, cleaning up the presentation.
# +
import math
math.degrees(math.atan(1/2))
# -
# $$ \vec{a}$$
| Visualization Case Study.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
# +
# read everything in
t0 = pd.read_csv('rawdata/180302_NB501961_0075_AH5FWKBGX5_gene_cell_table.csv', index_col=0)
t1 = pd.read_csv('rawdata/180306_NB501961_0078_AH5G7VBGX5_gene_cell_table.csv', index_col=0)
t2 = pd.read_csv('rawdata/180311_NB501961_0082_AHL5LHBGX5_gene_cell_table.csv', index_col=0)
t3 = pd.read_csv('rawdata/180319_A00111_0118_BH7GTHDMXX_gene_cell_table.csv', index_col=0)
t4 = pd.read_csv('rawdata/180504_A00111_0143_BH3VG5DSXX_gene_cell_table.csv', index_col=0)
print(t0.shape)
print(t1.shape)
print(t2.shape)
print(t3.shape)
print(t4.shape)
# -
def trim_col_names(col_list):
""" removes all this extraneous shit from the cell names"""
new_cols = []
for x in list(col_list):
try:
y = x.split('-')[0] + '_' + x.split('-')[1]
except IndexError:
y = x
new_cols.append(y)
return(new_cols)
t0_new_cols = trim_col_names(t0.columns)
t1_new_cols = trim_col_names(t1.columns)
t2_new_cols = trim_col_names(t2.columns)
t3_new_cols = trim_col_names(t3.columns)
t4_new_cols = trim_col_names(t4.columns)
# shouldnt be any overlaps
set(t4_new_cols).intersection(set(t3_new_cols))
# reset col names
t0.columns = t0_new_cols
t1.columns = t1_new_cols
t2.columns = t2_new_cols
t3.columns = t3_new_cols
t4.columns = t4_new_cols
# +
# lets rename the files entirely
#t0.to_csv('180302_expression_matrix.csv')
#t1.to_csv('180306_expression_matrix.csv')
#t2.to_csv('180311_expression_matrix.csv')
#t3.to_csv('180319_expression_matrix.csv')
#t4.to_csv('180504_expression_matrix.csv')
# -
# what does this metadata obj look like?
meta = pd.read_csv('metadata/MouseTBIPlateMetadata.csv')
meta
# +
#//////////////////////////////////////////////////////////////////////////////
#//////////////////////////////////////////////////////////////////////////////
#////////////////////// combine em dataframes /////////////////////////////
#//////////////////////////////////////////////////////////////////////////////
# -
# first need to get rid of these 'Undetermined' col names
print(list(t0.columns)[-1])
print(list(t1.columns)[-1])
print(list(t2.columns)[-1])
print(list(t3.columns)[-1])
print(list(t4.columns)[-1])
t0 = t0.drop('Undetermined_S0.mus', axis=1)
t1 = t1.drop('Undetermined_S0.mus', axis=1)
t2 = t2.drop('Undetermined_S0.mus', axis=1)
# combine everyone
em_super = pd.concat([t0, t1, t2, t3, t4], axis=1)
em_super
# +
# get a list of all the M1/M2 cells (to remove)
m1_m2_cells = []
for x in list(em_super.columns):
plate = x.split('_')[1]
meta_row = meta[meta.Sort_Plate_ID == plate]
mouse_id = list(meta_row['BH_Identifier '])[0]
if mouse_id == 'M1' or mouse_id == 'M2':
m1_m2_cells.append(x)
# -
cells_to_keep = set(em_super.columns) - set(m1_m2_cells)
em_super_trimmed = em_super[list(cells_to_keep)]
em_super_trimmed
# want to get rid of these wierd guys
list(em_super_trimmed.index)[-8:]
em_super_trimmed = em_super_trimmed[:-8]
em_super_trimmed.shape
# +
#em_super_trimmed.to_csv('M3-M8_expression_matrix.csv')
# +
#//////////////////////////////////////////////////////////////////////////////
#//////////////////////////////////////////////////////////////////////////////
#//////// finding a bunch of these excel-style rownames //////////////
#//////////////////////////////////////////////////////////////////////////////
# +
# quick check
# looks like its only in t0
months = ['Jan', 'Feb', 'Mar', 'Apr', 'May', 'Jun', 'Jul', 'Aug', 'Sep', 'Oct', 'Nov', 'Dec']
for x in list(t0.index):
for month in months:
if month in x and '-' in x:
print(x)
# +
# how many are there?
corrupted_rows = []
for x in list(em_super_trimmed.index):
for month in months:
if month in x and '-' in x:
corrupted_rows.append(x)
print(len(corrupted_rows))
# -
# fuck it lets just get rid of em
em_super_trimmed_cookin = em_super_trimmed[~em_super_trimmed.index.isin(corrupted_rows)]
em_super_trimmed_cookin
# +
#//////////////////////////////////////////////////////////////////////////////////////
#//////////////////////////////////////////////////////////////////////////////////////
#//////////////////////////////////////////////////////////////////////////////////////
# -
# fuck it, im gonna remove the ERCCs too
ercc_list = []
for x in list(em_super_trimmed_cookin.index):
if 'ERCC' in x:
ercc_list.append(x)
em_super_trimmed_cookin1 = em_super_trimmed_cookin[~em_super_trimmed_cookin.index.isin(ercc_list)]
em_super_trimmed_cookin1
em_super_trimmed_cookin1.to_csv('M3-M8_expression_matrix_cookin.csv')
| reformat_expression_matricies.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# %matplotlib notebook
import numpy as np
import numpy
from tqdm import tqdm
import gensim
import random
# -
# ## Download Word2Vec Vectors from https://drive.google.com/uc?id=0B7XkCwpI5KDYNlNUTTlSS21pQmM&export=download
#
# ## Download analogies data from http://download.tensorflow.org/data/questions-words.txt
# 
model = gensim.models.KeyedVectors.load_word2vec_format('data/GoogleNews-vectors-negative300.bin', binary=True)
king_vector = model['king']
print(len(king_vector))
print(king_vector)
analogy_vector = model['king'] - model['man'] + model['queen']
print(analogy_vector)
# example analogy task like king - man + woman = queen
answer = model.most_similar(positive=['woman', 'king'], negative=['man'])
print("king - man + woman = {}".format(answer))
analogy_words = [line.rstrip('\n').split(' ') for line in open('data/questions-words.txt')]
analogy_words = [words for words in analogy_words if len(words) == 4]
np.random.seed(0)
analogy_words = random.sample(analogy_words, 100)
X = [words[:3] for words in analogy_words]
y = [words[3] for words in analogy_words]
print(X[0], y[0])
print(X[10], y[10])
print(X[50], y[50])
is_correct_list = []
top_5_predictions_list = []
for i in tqdm(range(len(X))):
components = X[i]
answer = y[i]
predictions = model.most_similar(positive=[components[1], components[2]], negative=[components[0]])
top_5_predictions = [p[0].lower() for p in sorted(predictions, key=lambda x : -x[1])[:10]]
top_5_predictions_list.append(top_5_predictions)
is_in_top_5 = 1.0 if answer.lower() in top_5_predictions else 0.0
is_correct_list.append(is_in_top_5)
for i in range(10):
components = X[i]
answer = y[i]
top5 = top_5_predictions_list[i]
correct = is_correct_list[i]
print("Components: {}, Answer: {} Top5: {} is_correct: {}".format(components, answer, top5, correct))
print("Accuracy in Analogy Task is", np.mean(is_correct_list))
| Sec7/7.2 Notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Formatting and Analysis
# # =====================
# +
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import sktime
# %matplotlib inline
# -
data = pd.read_csv('/home/zone/Documents/advantage-investing/data/raw/Goog.csv')
data
data.dtypes
data.columns
data[:][:1]
for col in data.drop(columns=['Unnamed: 0']):
if str(col) !=['Unnamed: 0']:
print(str(col))
data[col] = data[col].str.split('.').str[0]
for col in data.drop(columns=['Unnamed: 0']):
print(str(col))
data[col] = data[col].str.strip(',')
df.Close
df = data.replace(',','', regex=True)
df.Date = pd.to_datetime(df.Date)
df.Close = pd.to_numeric(df.Close)
df = df.set_index(df.Date,drop = True)
df = pd.DataFrame(df.Close)
df
# +
plt.figure(figsize=(9,5))
plt.grid(True)
plt.xlabel('')
plt.ylabel('')
plt.yticks(np.arange(0, 3000, 100))
plt.plot(df['Close'])
plt.title('goog closing price')
plt.show()
# -
plt.figure(figsize=(10,6))
df_close = df.Close
df_close.plot(style='k.')
plt.title('Scatter plot of closing price')
plt.show()
# ### Is the data stationary? Data points are often non-stationary or have means, variances, and covariances that change over time.
# ### Non-stationary behaviors like trends are unpredictable and produce unreliable forecasts
from statsmodels.tsa.stattools import adfuller
# +
def test_stationarity(timeseries):
#Determing rolling statistics
rolmean = timeseries.rolling(12).mean()
rolstd = timeseries.rolling(12).std()
#Plot rolling statistics:
plt.plot(timeseries, color='yellow',label='Original')
plt.plot(rolmean, color='red', label='Rolling Mean')
plt.plot(rolstd, color='black', label = 'Rolling Std')
plt.legend(loc='best')
plt.title('Rolling Mean and Standard Deviation')
plt.show(block=False)
print("Results of dickey fuller test")
adft = adfuller(timeseries,autolag='AIC')
# output for dft will give us without defining what the values are.
#hence we manually write what values does it explains using a for loop
output = pd.Series(adft[0:4],index=['Test Statistics','p-value','No. of lags used','Number of observations used'])
for key,values in adft[4].items():
output['critical value (%s)'%key] = values
print(output)
test_stationarity(df.Close)
# -
# ### The mean and std changes over time so our data is non stationary and looking at the graph the data is showing a trend.
# ### First we'll create train and test sets then we'll fit an ARIMA model to this data. ( ARIMA models use differencing to extract stationary data from our originally non stationary data)
#
df_log = df_log[::-1]
train_data, test_data = df_log[3:int(len(df_log)*0.9)], df_log[int(len(df_log)*0.9):]
plt.figure(figsize=(10,6))
plt.grid(True)
plt.xlabel('Dates')
plt.ylabel('Closing Prices')
plt.plot(df_log, 'green', label='Train data')
plt.plot(test_data, 'blue', label='Test data')
plt.legend()
len(test_data)
# !pip install kats
# !pip install arrow
| notebooks/1.0-xs-analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from PIL import Image
import numpy as np
from skimage import transform
def getMaskR(pic,saveName = None):
np_image = Image.open(pic)
np_image = np.array(np_image).astype('float32')/255
np_image = transform.resize(np_image,(256,256,1))
np_image = np.expand_dims(np_image, axis=0)
maskArr = modelR.predict(np_image)
maskArr = maskArr.reshape(256,256)
#np.save(saveName,maskArr)
maskArrThresh = np.zeros((maskArr.shape[0], maskArr.shape[1], 4))
for row in range(0, maskArr.shape[0]):
for col in range(0, maskArr.shape[1]):
#if maskarr[row, col] < 0.:
maskArrThresh[row, col, 0] = 255
maskArrThresh[row, col, 1] = 50
maskArrThresh[row, col, 2] = 0
maskArrThresh[row, col, 3] = int(maskArr[row, col] * 255)
#else: #maskArrThresh[row, col, 0:3] = 255 # maskArrThresh[row, col, 3] = 0
maskArrThresh = maskArrThresh.astype("uint8")
maskImg = Image.fromarray(maskArrThresh, 'RGBA')
return maskImg
def getMaskB(pic,saveName = None):
np_image = Image.open(pic)
np_image = np.array(np_image).astype('float32')/255
np_image = transform.resize(np_image,(256,256,1))
np_image = np.expand_dims(np_image, axis=0)
maskArr = modelB.predict(np_image)
maskArr = maskArr.reshape(256,256)
#np.save(saveName,maskArr)
maskArrThresh = np.zeros((maskArr.shape[0], maskArr.shape[1], 4))
for row in range(0, maskArr.shape[0]):
for col in range(0, maskArr.shape[1]):
#if maskarr[row, col] < 0.:
maskArrThresh[row, col, 0] = 50
maskArrThresh[row, col, 1] = 255
maskArrThresh[row, col, 2] = 0
maskArrThresh[row, col, 3] = 1-int(maskArr[row, col] * 255)
#else: #maskArrThresh[row, col, 0:3] = 255 # maskArrThresh[row, col, 3] = 0
maskArrThresh = maskArrThresh.astype("uint8")
maskImg = Image.fromarray(maskArrThresh, 'RGBA')
return maskImg
def displayPrediction(maskImg,pic):
PANImg = Image.open(pic)
PANImgT = PANImg.convert('RGBA')
PANDim = (PANImgT.width, PANImgT.height)
maskImgResize = maskImg.resize(PANDim)
comp = Image.alpha_composite(PANImgT,maskImgResize)
return comp
def displayBoth(maskImgR,maskImgB,pic):
PANImg = Image.open(pic)
PANImgT = PANImg.convert('RGBA')
PANDim = (PANImgT.width, PANImgT.height)
maskImgResizeR = maskImgR.resize(PANDim)
maskImgResizeB = maskImgB.resize(PANDim)
comp = Image.alpha_composite(PANImgT,maskImgResizeR)
comp1 = Image.alpha_composite(comp,maskImgResizeB)
return comp1
| testUnet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # TimeSeries - a new object for handling time domain data
#
# ### NOTE: Internet access is required in order to use this tutorial
#
# TimeSeries is a new feature in the SunPy 0.8 release, replacing the LightCurve object which is now deprecated. Similar to LightCurve, TimeSeries handles time domain data from a variety of solar instruments, including every mission that was previously supported by LightCurve. In TimeSeries, the downloading of data has been separated from the object - now, the object operates only on data files, while downloading is done via other tools such as the Unified Downloader (FIDO).
#
# In this notebook, we demonstrate how to use TimeSeries and give a brief tour of its features.
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.dates as mdates
# %matplotlib inline
import datetime
import sunpy
from sunpy.net import Fido, attrs as a
# ### 1. Download a data file to work with
# As mentioned above, TimeSeries does not download data itself. That functionality is covered by other SunPy tools. Without going into detail, here we use Fido to download a GOES XRS data file from 2011 June 7. Detailed tutorials on using FIDO may be found elsewhere.
search_results = Fido.search(a.Time('2011-06-07 00:00','2011-06-07 08:00'),a.Instrument('XRS'))
Fido.fetch(search_results[0],path='.')
# ### 2. Create a TimeSeries object from a data file
# Now that we have a GOES data file in the local directory, let's load it into a TimeSeries object and take a look at it.
from sunpy import timeseries
goes = timeseries.TimeSeries('go1520110607.fits')
goes.peek()
# Here we can see a quicklook plot of the data in the object. It shows the two GOES XRS channels, 0.5-4A and 1-8A. We can see that a solar flare occured at ~06:20 UT.
# The TimeSeries object retains the metadata from the originating file. We can query this as follows:
print(goes.meta)
# The data itself is stored in something called a pandas DataFrame. This DataFrame is designed to hold column-based data. The first column is the index (i.e. the time dimension), while the other columns correspond to the dependent variables, in this case the flux from the two GOES channels. Below, we show what the data looks like, showing only the first 5 rows for brevity.
print(goes.data[0:5])
# Data for a particular variable can easily be extracted like this (again, extracting only the first 5 rows for convenience):
xrsa = goes.data['xrsa'][0:5]
print(xrsa)
# The TimeSeries object also knows the units of the data in each column, e.g.
print(goes.units)
# ### 3. Working with TimeSeries objects
# By default, the entire file contents are converted to a TimeSeries object. A subset of the data can easily be extracted, similar to creating a SubMap from a larger Map object. This is done using the ```truncate()``` function:
goes2 = goes.truncate('2011-06-07 06:00','2011-06-07 08:00')
goes2.peek()
# #### Finding the maxima in the data
# We can easily find the maximum value for each channel, and the index location of these maxima, like so:
print(goes2.data.max())
print(goes2.data.idxmax())
# #### Subtracting the moving average of the data
# The TimeSeries object contains a number of useful tools for common time series operations. For example, to subtract a rolling mean of the data, we can use the built-in function ```rolling()```
smoothed_goes_data = goes2.data.rolling(60, center=True).mean()
# The object `smoothed_goes_data` contains the rolling mean with a 60-point window for each channel. We can easily subtract it from the original data and plot the result.
# +
fig1 = plt.figure(1,figsize=(12,6))
plt.plot(goes2.data.index,goes2.data['xrsa'] - smoothed_goes_data['xrsa'],label='xrsa')
plt.plot(goes2.data.index,goes2.data['xrsb'] - smoothed_goes_data['xrsb'],label='xrsb')
plt.xlim(datetime.datetime(2011,6,7,6,25),datetime.datetime(2011,6,7,6,45))
plt.legend(fontsize=14)
#This line just formats the x-axis to look nicer
plt.gca().xaxis.set_major_formatter(mdates.DateFormatter('%H:%M:%S'))
# -
# #### Resample the TimeSeries
# It is easy to resample the data in the TimeSeries, for example to reduce the cadence:
goes_resampled = goes2.data.resample('30S').mean()
print(goes_resampled[0:5])
# #### Other useful functions
goes2.data.diff() # take the difference between each data point
goes2.data.cumsum() # return the cumulative sum of the data.
goes2.index.duplicated() # returns True for time indexes that appear more than once in the TimeSeries
goes2.index.drop_duplicates() # just remove all duplicate rows from the TimeSeries
goes2.concatenate() # allows two DataFrames to be joined together into one.
# ...and much more!
# ### 4. Summary
# By the end of this notebook you should have some familarity with the basic functionality of TimeSeries, including:
# - how to create a TimeSeries from a data file
# - how to access the data and metadata in the TimeSeries object
# - how to create a truncated TimeSeries from a longer one
# - how to perform simple data manipulation, such as averaging, subtracting a running mean.
| 2017_SPD_Portland/2017 SPD Workshop - Creating a timeseries.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/zavorfang/Api/blob/master/yt2mega_v1_0_1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="MLh6YfpVBAKS" outputId="7b718a7c-e366-4fe7-fd13-d3ac44521ba1"
# !pip install pytube mega.py
# + colab={"base_uri": "https://localhost:8080/"} id="IFKQ05DTIGn9" outputId="4077c06d-637f-4ba1-ef0b-ff9e3498f0b0"
from pytube import YouTube, Playlist
import os, time, datetime
from mega import Mega
mega = Mega()
username = ""
password = ""
m = mega.login(username, password)
def dl_mega(path):
file = m.upload(f'{os.listdir(path)[0]}')
return m.get_upload_link(file)
def create_dir(name=None):
# Directory
directory = name or input('Enter name of folder: ')
# Parent Directory path
parent_dir = "/content"
# Path
path = os.path.join(parent_dir, directory)
# Path existence verification
if not os.path.exists(path):
# Create the directory
os.mkdir(path)
return path
def display_qualities():
print('Enter:\n[0] 720p\n[1] 480p\n[2] 360p\n[3] 240p\n[4] 144p')
num = input('Qualiy: ')
return num
def download_fn(num):
res = ["720p", "480p", "360p", "240p", "144p"]
if num.isdigit() and int(num) <= 4:
num = int(num)
begin = time.time()
# playlist
if str(url).find('playlist') > 0:
yt = Playlist(url)
filesize = 0
path = create_dir(yt.title)
for video in yt.videos:
try:
stream = video.streams.filter(res=res[num]).first()
print(f"Downloading selected quality: {res[num]}")
except:
stream = video.streams.get_highest_resolution().first()
print(f"Downloading highest quality")
filesize += stream.filesize
stream.download(path)
# Single
else:
yt = YouTube(url)
try:
stream = yt.streams.filter(res=res[num]).first()
print(f"Downloading selected quality: {res[num]}")
except:
stream = yt.streams.get_highest_resolution()
print(f"Downloading highest quality")
filesize = stream.filesize
stream.download(path)
os.chdir(path)
print(dl_mega(path))
end = time.time()
filesize = round(filesize/(1024*1024))
print("Total size copied: ", filesize, " MB")
print("Elapsed Time: ",int((end-begin)//60),"min :", int((end-begin)%60), "sec")
print(datetime.datetime.now())
return None
print('ENTER 0, 1, 2, 3 or 4')
num = display_qualities()
download_fn(num)
path = create_dir()
url = input('Video/Playlist URL: ')
num = display_qualities()
download_fn(num)
| yt2mega_v1_0_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import numpy as np
from astropy.io import fits
from astropy.visualization import astropy_mpl_style
from astropy.visualization.mpl_normalize import ImageNormalize
from astropy.visualization import SqrtStretch
import matplotlib.pyplot as plt
plt.style.use(astropy_mpl_style)
from fonctions import *
""" Test for the average of images"""
folder = 'test-bias'
avg_img = averageFolder(folder)
plt.figure()
plt.title('Average Image')
norm = ImageNormalize(stretch=SqrtStretch())
plt.imshow(avg_img, norm=norm, origin='lower', cmap='viridis', interpolation='none')
plt.colorbar()
plt.show()
| testAverage.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Alistair's test cases that I'm including with my pull requests to Tony
# Assume that [the basic test suite](TM351%20VM%20Installation%20Test.ipynb) runs OK.
# Now load in `pandas` and the `sql` extension:
# +
import pandas as pd
import sqlalchemy
# %load_ext sql
# -
# ## connect to the database using the connection string from notebook 03.3
# +
# Execute this cell if you are using the local environment
DB_USER='tm351'
DB_PWD='<PASSWORD>'
DB_NAME='tm351'
# +
# Create the connection string for the sql magic
CONNECT_DB='postgresql://{}:{}@localhost:5432/{}'.format(DB_USER, DB_PWD, DB_NAME)
CONNECT_DB
# -
# %sql $CONNECT_DB
# Good, that seems to work. Now do a quick test with the `quickdemo` table:
# + language="sql"
#
# DROP TABLE IF EXISTS quickdemo;
#
# CREATE TABLE quickdemo(id INT PRIMARY KEY, name VARCHAR(20), value INT);
#
# INSERT INTO quickdemo VALUES(1,'This',12);
# INSERT INTO quickdemo VALUES(2,'That',345);
#
# SELECT * FROM quickdemo;
# -
# result = %sql SELECT * FROM quickdemo WHERE value > 25
result
# Excellent. Those seem OK.
# ## connect to the database using the connection string from notebook 08.1
# What we want first is to check that the reset script is working OK. I've put the script and all the support files in the folder `AGW_test_files`
# I'll use `runpy` to execute the script. That'll need the student to be logged in with the correct credentials:
# +
# Execute this cell if you are using the local environment
DB_USER='tm351_student'
DB_PWD='<PASSWORD>'
DB_NAME='tm351_clean'
# -
# The `runpy` script needs to delete any tables from the `tm351_clean` database which might be called in the rest of the notebook.
import runpy
# +
runpy.run_path('./AGW_test_files/sql_initial_state.py',
{'DB_USER_CLEANUP':DB_USER,
'DB_PWD_CLEANUP':DB_PWD,
'DB_NAME_CLEANUP':DB_NAME})
pass
# -
# Check that the tables are correctly defined in the `tm351_hospital` schema:
# +
# Create the connection string for the sql magic
CONNECT_DB='postgresql://{}:{}@localhost:5432/{}'.format(DB_USER, DB_PWD, DB_NAME)
CONNECT_DB
# -
# %sql $CONNECT_DB
# + language="sql"
#
# SELECT *
# FROM tm351_hospital.patient
# -
# And check that the foreign keys are properly implemented:
try:
# %sql DELETE FROM tm351_hospital.doctor WHERE doctor_id='d06'
except sqlalchemy.exc.IntegrityError:
print("Integrity Error raised OK")
| .tests/TM351 VM AGW Tests.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import tensorflow as tf
import cProfile
tf.executing_eagerly() # In TF2 eager execution is enabled by default
# #### Tensors
# - Shape of a tensor are the number of elements in each dimension (ex. 2 rows, 3 columns -> TensorShape([2, 3]))
# - Rank of a tensor is the number of dimensions of the tensor
s1 = tf.Variable('It is done', tf.string)
s1.shape, tf.rank(s1)
x1 = tf.Variable(32, tf.int32)
x1.shape, tf.rank(x1)
x2 = tf.Variable([[1, 2, 3], [4, 5, 6]], tf.int32)
x2.shape, tf.rank(x2)
# ##### Reshaping tensors
tf.reshape(x2, [3, 2])
tf.reshape(x2, [6, -1]) # -1 implies "infer the number of elements in this dimension"
tf.reshape(x2, [-1, 6])
# ##### Constant and Variable tensors
# - You can point the constant tensor's name to another tensor
# - You cannot change the constant tensor's values inside, thus
# - it cannot be used during training
# - A variable tensor's values can be updated, thus
# - it can be used during training (default).
# - You can also create a variable tensor to be untrainable (using trainable=False)
c = tf.constant([[1, 8, 9]], tf.int32)
c = tf.constant([[1, 2, 3]], tf.int32)
c
d = tf.Variable([1, 8, 9], tf.int32)
d
d = 2 * d
d
| tf2_basics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
# Dersler listesindeki her dersi yazdıralım.
dersler = ["Matematik", "Fizik", "Kimya","Biyoloji"]
for x in dersler:
print(x)
# -
#Pythonda For Döngüsü ile bir metnin karakterleri arasında dolaşmak mümkündür.
for x in "PYTHON":
print (x)
#Break ifadesiyle, döngüyü tüm öğeler arasında dolaşmadan önce durdurabiliriz.
dersler = ["Matematik", "Fizik", "Kimya","Biyoloji"]
for x in dersler:
print(x)
if x=="Fizik":
break
#continue ifadesi ile döngünün geçerli yinelemesini durdurabilir ve sonraki adımla devam edebiliriz.
dersler = ["Matematik", "Fizik", "Kimya","Biyoloji"]
for x in dersler:
if x=="Fizik":
continue
print (x)
""" Belirli bir sayıda kod kümesi arasında dolaşmak için, range () işlevini kullanabiliriz,
range () işlevi varsayılan olarak 0’dan başlayarak ve 1 (varsayılan olarak) artışlarla belirtilen sayıda bitiyor."""
for x in range(6):
print(x)
for x in range(2,6):
print(x)
#Python For Döngüsü else Kullanımı
#Bir for döngüsünde else anahtar sözcüğü döngü bittiğinde yürütülecek kod bloğunu belirtir:
for x in range(6):
print(x)
else:
print("Döngü bitti!")
# +
"""Python İç İçe For Döngüsü Kullanımı
İç içe bir döngü, döngü içine eklenmiş bir döngüdür. İki, üç ya da daha fazla olabilir.
‘İç döngü’, ‘dış döngü’ nün her bir yinelemesi için bir kez gerçekleştirilir:"""
dersler = ["<NAME>", "<NAME>", "Ders 3"]
konular = ["Konu 1", "Konu 2", "Konu 3"]
for x in dersler:
for y in konular:
print(x, y)
# -
for x in range(1,11):
for y in range(1,11):
print(x,"x", y,"=",x*y)
| Python Loops-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/RichardFreedman/CRIM-notebooks/blob/master/CRIM/CRIM_Intervals_Basics.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="uqgPHgqXPrJa"
# ### Load CRIM Intervals Software
# + colab={"base_uri": "https://localhost:8080/"} id="JY6rMDPCmYGf" outputId="9c207571-fe6d-441e-f3a7-19130237e530"
# !git clone https://github.com/HCDigitalScholarship/intervals.git
# !pip install httpx
# + [markdown] id="sabnU304P1NU"
# ## Start CRIM Intervals
# + id="qHbslxGqmexc" outputId="3a5a2b5f-7da2-421c-8154-a7229f90a2a6" colab={"base_uri": "https://localhost:8080/", "height": 419}
from intervals.main_objs import *
# + [markdown] id="1RkHh_LKP4kt"
# ## Load MEI Files from CRIM or Github by pasting one or more of [these links](https://docs.google.com/spreadsheets/d/1TzRqnzgcYYuQqZR78c5nizIsBWp4pnblm2wbU03uuSQ/edit?auth_email=<EMAIL> <EMAIL>#gid=0)below.
#
# *Note: each file must be in quotation marks and separated by commas
#
#
#
#
#
#
# + id="WhUxkepkmmV6"
corpus = CorpusBase(['https://crimproject.org/mei/CRIM_Model_0008.mei', 'https://crimproject.org/mei/CRIM_Mass_0005_5.mei'])
# + [markdown] id="woLnjv-uSLgq"
# ## Give the scores short names, in order according to the way they were listed above
# + id="OOEp3RaPmnH-"
model, mass = corpus.scores
# + [markdown] id="q41OFP5FSVSl"
# ## Now apply various methods to the scores:
# * getNoteRest returns all the notes and rests, each voice as a column
# * getDuration returns the durations for all notes and rests, as above
# * getMelodic returns the melodic intervals in each voice as a column
# * getHarmonic returns pairs of harmonic intervals between each pair of voices
# * getNgrams returns segments of various kinds, melodic (one voice) or modular (pairs of voices, including vertical and horizontal motion)
#
#
#
#
# ---
#
# ## Documentation available via this command:
#
# print(model.getNgrams.__doc__)
#
# ---
#
#
#
#
#
#
#
# + id="TqIwSBI_TvBE"
print(model.getMelodic.__doc__)
# + id="yh6oPV2KrZaM"
notes = mass.getNoteRest()
notes.fillna(value= "-", inplace=True)
notes.head()
# + id="pmXfVpnSWrrA"
notes.stack().value_counts()
# + id="eLeZuKLvW7qn"
notes.apply(pd.Series.value_counts).fillna(0).astype(int)
# + id="loTNxtPfmyo4"
mel = mass.getMelodic(kind='d')
mel.fillna(value= "-", inplace=True)
mel.head(10)
# + id="7CYg4NqysUwF"
durs = mass.getDuration()
durs.fillna(value= "-", inplace=True)
durs.head()
# + id="gcqxZm4Isasv"
notes_durs = pd.concat([notes, durs], axis=1)
notes_durs
# + id="7wbhoYMYm49L"
mel = model.getMelodic()
model.getNgrams(df=mel, n=7)
# + id="H9pS4PKZrvAp"
modules = model.getNgrams(how='modules', cell_type=str)
modules.iloc[:, 0:6]
| CRIM_Intervals_Basics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [python3]
# language: python
# name: Python [python3]
# ---
1 + 2
x = 2+3
print(x)
# + active=""
#
# -
y = x -2
print(y)
x = 10
y = x -2
print(y)
x = 2 + 3
y = x -2
print(y)
# +
print(2 + 1)
print(5 - 2)
print(1 + 1)
print(2 - 1)
print(1 + 5)
# -
# # copy and paste code here
| .ipynb_checkpoints/Untitled-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <NAME> at Kyoto University, April 28th - April ~th, 2018
#
# 机器学习,无论单纯从理论,或者应用的角度来看,都可能会只看到冰山一角。现在,我们是从基础开始的,即fundation Oriented Machine Learning. 在学习的过程中,穿插基础在理论和实践中的实际应用,讨论各种应用场景背后的基础。
#
# The goal of this course is make students learn 'future/untaught' techniques or study deeper theory easily.
# # Learning to Answer Yes/No
#
# 回顾下course01--the learning problem: $\mathcal{A}$ (algorithm) takes $\mathcal{D}$ (database) and $\mathcal{H}$ (hypothesis set) to get $g$ (final hypothesis)
#
# 在当前的course02中,我们则要让机器学习帮助我们判断 yes or no。
#
# ## Perceptron Hypothesis Set ($\mathcal{H}$)
#
# 假设我们是一家银行,当一位客户(用户信息见下表)对我行提出信用卡申请时,我们需要判断核发或者拒发。
#
# | age | 23 years |
# | ------------- |:-------------:|
# | annual salary | NTD 1,000,000|
# | year in job | 0.5 year |
# | current debt | 200,000 |
#
# + For $\vec{x}$ = ($x_1$, $x_2$, $x_3$, ..., $x_d$) -- 'features of customer', compute a weighted 'score' and
#
# - approve credit if ${\sum_{i=1}^d w_ix_i > {\rm threshold}}$
# - deny credit if ${\sum_{i=1}^d w_ix_i < {\rm threshold}}$
#
# + $y$: {+1(good), -1(bad)}, 0 ignored -- linear formula $h$ ${\in}$ $\mathcal{H}$ are
#
# ${h({\mathbf{x}}) = {\rm sign}((\sum_{i=1}^d w_ix_i) - {\rm{threshold}})}$
#
# **Note**:
#
# 'perceptron': 感知器,来源于类神经网络中的专业名词。
#
#
# 我们可以对perceptron hypothesis ${h({\mathbf{x}})}$ 矢量化,是指具有更简单对形式:
#
# ${h({\mathbf{x}}) = {\rm sign}((\sum_{i=1}^d w_ix_i) - {\rm{threshold}})}$
#
# = sign${((\sum_{i=1}^d w_ix_i) + (-{\rm{threshold}})\cdot(+1))}$
#
# 这里,我们可以把 -threshold 这一项取为 ${w_{\rm 0}}$, +1 取作 ${x_0}$,那么我们就可以把第二项放到第一项里面,上面的公式就变为
#
# = sign(${\sum_{i=0}^d w_ix_i}$)
#
# 我们可以把 ${\sum_{i=0}^d x_i}$ 和 ${\sum_{i=0}^d w_i}$ 都写成vector form,then we have
#
# = sign(${\mathbf{w}^T \mathbf{x}}$)
#
# - each 'tall' $\mathbf{w}$ represents a hypothesis $h$ $\&$ is multiplied with 'tall' $\mathbf{x}$ -- will use tall versions to simplify notation
#
#
#
#
#
#
# what do perceptrons $h$ 'look linke'? -- Perceptrons in ${\mathbb{R}^2}$
#
# ${h({\mathbf{x}})}$ = sign(${w_0 + w_1x_1 + w_2x_2}$)
#
# 这里 $x_1$ 和 ${x_2}$ 分别代表客户features中的第一个和第二个维度。
#
# 示意图如下,
# 
#
# - customer features ${\mathbf{x}}$: points on the plane (or points in ${\mathbb{R}^d}$, 在上面的式子中,$d$ = 2)
#
# - labels $y$: $\circ$ represents (+1), $\times$ represents (-1)
#
# - hypothesis $h$: $\textbf{lines}$ (or hyperplanes in ${\mathbb{R}^d}$)
#
# --positive on one side of a line, negative on the other side.
#
# 因此在二维平面内,这个 perceptron 其实就是一条线,因此perceptron 也被称作 linear (binary) classifiers. 在三维中,这个 perceptron 就是一个面。
# ## Perceptron Learning Algorithm (PLA)
#
#
# $\mathcal{H}$ = all possible perceptrons, but how can we select $g$ from $\mathcal{H}$?
#
# 我们的出发点如下
#
# - want: $g$ $\approx$ $f$ (hard when $f$ unknown)
# - almost necessary: $g$ $\approx$ $f$ on $\mathcal{D}$, ideally ${g(\mathbf{x}_n)}$ = ${f(\mathbf{x}_n)}$ = ${y_n}$
#
# 但是我们发现,一个一个去尝试 $\mathcal{H}$ 中的 $h$ 在实际情况中是十分困难的,因为$\mathcal{H}$ 是无限的。即使是在上一个图中的二维问题,这个集合就是无限的,因为一个二维平面中有无数条直线。显然,穷尽所有,几乎是不可能的。
#
# 我们不妨先取其中的一个,例如 ${g_0}$, 也许这条线并不是很好(指代它可能离$f$仍然相差较大),然后通过数据 $\mathcal{D}$, 不断地correct它。为了方便,我们用weight vector ${w_0}$ 来代表 ${g_0}$.
#
#
# +
Cyclic PLA
For $t$ = 0, 1, ...
# -
# Some remaining issures of PLA?
#
# 它是否会停下来?假设对于 $\mathcal{D}$,我们已经找到了一个 $g$ $\approx$ $f$,那么对于 $\mathcal{D}$ 之外的数据, $g$ 是否还能保持接近于我们希望的那个 $f$呢? 这个问题留待之后的学习中解决。
| machine-learning/coursera/MachineLearningFoundations/course02.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Bumblebees and Flowers
# Lets simulate the adventures of a bumblebee in a field of flowers with the following assumptions:
#
# - In our simulation there exist a fixed number of flowers.
# - Pollen can be harvested by a bumblebee from each flower.
# - The mean amount of pollen that can be harvested is different from flower to flower.
# - The amount of pollen for a specific visit is determined by a Gaussian distribution.
# - The Bumblebee tries to figure out what the best flowers are, measured by expected amount of pollen.
#
# For the simulation we use the two classes FlowerField and Bumblebee. Lets get a feeling for them first before we start the simulation.
from course_001_FlowerField import FlowerField
ff = FlowerField()
ff.visit_flower_nr(3)
from course_001_Bumblebee import Bumblebee
bb = Bumblebee()
bb.memory_df
bb.choose_flower()
bb.update_memory(flower_id=6, amount_of_pollen=8)
# ## First contact with a reinforcement learning Setup using just numpy/pandas
# +
import matplotlib.pyplot as plt
# %matplotlib inline
import numpy as np
np.random.seed(42)
plt.xlabel("Number of Trips")
plt.ylabel("Average Reward")
ff = FlowerField()
bb = Bumblebee()
n_trips = 500
aops = []
for trip in range(n_trips):
flower_id = bb.choose_flower()
amount_of_polen = ff.visit_flower_nr(flower_id)
bb.update_memory(flower_id, amount_of_polen)
aops = [amount_of_polen] + aops
average_to_see_through_the_noise = True
if average_to_see_through_the_noise:
plt.scatter(trip, np.mean(aops[:50]))
else:
plt.scatter(trip, amount_of_polen)
| course_001_minimal-example-bumblebee.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="oXyt3vkIw7K2" outputId="4aa6d7a0-bd87-4d7d-efa5-d40c6e961cef"
'''
<NAME>, <NAME>, <NAME>, <NAME>
MSc students in Artificial Intelligence
@ Alma Mater Studiorum, University of Bologna
January, 2022
'''
from google.colab import drive
drive.mount('/content/drive')
# + [markdown] id="4847FosCyyhj"
# # Project work: **Key-Points Matching**
# Key Point Analysis (KPA) is a new NLP task, with strong relations to computational argumentation, opinion analysis, and summarization.
#
# Given an input corpus, consisting of a collection of relatively short, opinionated texts focused on a topic of interest, the goal of KPA is to produce a succinct list of the most prominent key-points in the input corpus, along with their relative prevalence. Thus, the output of KPA is a bullet-like summary, with an important quantitative angle and an associated well-defined evaluation framework.
#
# Successful solutions to KPA can be used to gain better insights from public opinions as expressed in social media, surveys, and so forth, giving rise to a new form of a communication channel between decision makers and people that might be impacted by the decision.
# + id="OxAEuybR3KvW"
import numpy as np
import pandas as pd
import tensorflow as tf
# !pip install tensorflow-text --quiet
import tensorflow_text as text
# !pip install --upgrade tensorflow_hub --quiet
import tensorflow_hub as hub
import matplotlib.pyplot as plt
from tqdm import tqdm
# !pip install contractions --quiet
import contractions
# !pip install focal-loss --quiet
import focal_loss
# !pip install -U sentence-transformers --quiet
import sentence_transformers
# + [markdown] id="k2TCu5NMy3oG"
# ## 1.0 **Data loading**
# The dataset is divided in:
# * Arguments (`arg_id`, `argument`, `topic`, `stance`);
# * Keypoint (`key_point_id`, `keypoint`, `topic`, `stance`);
# * Labels (`arg_id`, `key_point_id`, label).
#
# Each dataset has its own split (train, dev and test).
# The dataset is directly loaded via url from the GitHub repository.
# + colab={"base_uri": "https://localhost:8080/"} id="Iu4p-8vF3y-Y" outputId="a3ecad7f-1b60-4f47-b2b1-cbfb02d59ea1"
# Arguments dataset loading.
arguments_train = pd.read_csv(filepath_or_buffer = 'https://raw.githubusercontent.com/IBM/KPA_2021_shared_task/main/kpm_data/arguments_train.csv')
arguments_dev = pd.read_csv(filepath_or_buffer = 'https://raw.githubusercontent.com/IBM/KPA_2021_shared_task/main/kpm_data/arguments_dev.csv')
arguments_test = pd.read_csv(filepath_or_buffer = 'https://raw.githubusercontent.com/IBM/KPA_2021_shared_task/main/test_data/arguments_test.csv')
print(f'There are {len(arguments_train)} arguments in the train set, {len(arguments_dev)} arguments in the dev set and {len(arguments_test)} arguments in the test set.')
# Check.
# arguments_train.head()
# arguments_dev.head()
# arguments_test.head()
# + colab={"base_uri": "https://localhost:8080/"} id="N8T0uT3Z36q7" outputId="41319c3c-028c-46f6-fcde-5f0b9c7726c9"
# Key-points dataest loading.
key_points_train = pd.read_csv(filepath_or_buffer = 'https://raw.githubusercontent.com/IBM/KPA_2021_shared_task/main/kpm_data/key_points_train.csv')
key_points_dev = pd.read_csv(filepath_or_buffer = 'https://raw.githubusercontent.com/IBM/KPA_2021_shared_task/main/kpm_data/key_points_dev.csv')
key_points_test = pd.read_csv(filepath_or_buffer = 'https://raw.githubusercontent.com/IBM/KPA_2021_shared_task/main/test_data/key_points_test.csv')
print(f'There are {len(key_points_train)} keypoints in the train set, {len(key_points_dev)} keypoints in the dev set and {len(key_points_test)} keypoints in the test set.')
# Check.
# key_points_train.head()
# key_points_dev.head()
# key_points_test.head()
# + colab={"base_uri": "https://localhost:8080/"} id="xtqWZbTe36xS" outputId="eedad6ce-7b4f-4474-fe41-6c7036088752"
# Labels dataset loading.
labels_train = pd.read_csv(filepath_or_buffer = 'https://raw.githubusercontent.com/IBM/KPA_2021_shared_task/main/kpm_data/labels_train.csv')
labels_dev = pd.read_csv(filepath_or_buffer = 'https://raw.githubusercontent.com/IBM/KPA_2021_shared_task/main/kpm_data/labels_dev.csv')
labels_test = pd.read_csv(filepath_or_buffer = 'https://raw.githubusercontent.com/IBM/KPA_2021_shared_task/main/test_data/labels_test.csv')
print(f'There are {len(labels_train)} labels in the train set, {len(labels_dev)} labels in the dev set and {len(labels_test)} labels in the test set.')
# Check.
# labels_train.head()
# labels_dev.head()
# labels_test.head()
# + [markdown] id="_Sdi7-a6Ypch"
# The dataset is unbalanced.
# + colab={"base_uri": "https://localhost:8080/"} id="eZeDsyHXWrzn" outputId="ac0d7a67-1fd6-4220-e92b-b18ca4782060"
# Balance of the train dataset.
print(f"There are {np.sum(labels_train['label'].tolist())} positive samples and {len(labels_train['label']) - np.sum(labels_train['label'].tolist())} negative samples over {len(labels_train['label'])} samples of the training set.")
# Balance of the dev dataset.
print(f"There are {np.sum(labels_dev['label'].tolist())} positive samples and {len(labels_dev['label']) - np.sum(labels_dev['label'].tolist())} negative samples over {len(labels_dev['label'])} samples of the development set.")
# Balance of the test dataset.
print(f"There are {np.sum(labels_test['label'].tolist())} positive samples and {len(labels_test['label']) - np.sum(labels_test['label'].tolist())} negative samples over {len(labels_test['label'])} samples of the test set.")
# + [markdown] id="UaFHaKW9A_jt"
# ## 2.0 **Data processing**
# + [markdown] id="2UGqYY2dBb5m"
# ### 2.1 **Text cleaning**
# The text cleaning consists in lowercase every letter, expand abbreviations and verbal contractions, remove possible special characters and uncommon symbols.
#
# Notice that the new dataframes override the previous ones.
# + colab={"base_uri": "https://localhost:8080/"} id="a6YvcijoBe7J" outputId="ea0cf208-7a3a-4e32-aabd-b62d2324f64c"
import re
import nltk
from typing import List, Callable
from functools import reduce
# Lowercase everything.
def lower(text: str) -> str:
return text.lower()
# Expand abbreviations and verbal contractions.
def expand(text: str) -> str:
return contractions.fix(text)
# Remove special characters.
def replace_special_characters(text: str) -> str:
replace_symbols = re.compile('[/(){}\[\]\|@,;]')
return replace_symbols.sub(' ', text)
def filter_out_uncommon_symbols(text: str) -> str:
good_symbols = re.compile('[^0-9a-z #+_]')
return re.sub(good_symbols, '', text)
pipeline = [lower, expand, replace_special_characters, filter_out_uncommon_symbols]
def text_prepare(text: str,
filter_methods: List[Callable[[str], str]] = None) -> str:
'''
Applies a list of pre-processing functions in sequence (reduce).
Note that the order is important here!
'''
filter_methods = filter_methods if filter_methods is not None else pipeline
return reduce(lambda txt, f: f(txt), filter_methods, text)
def cleaning(dataframes: list, labels: list):
for dataframe in dataframes:
print('Before: \n')
print(dataframe[labels].head(5))
for label in labels:
dataframe[label] = dataframe[label].apply(lambda txt: text_prepare(txt))
print('After: \n')
print(dataframe[labels].head(5))
arguments_dataset_list = [arguments_train, arguments_dev, arguments_test]
arguments_labels_list = ['argument', 'topic']
cleaning(arguments_dataset_list, arguments_labels_list)
key_points_datasets_list = [key_points_train, key_points_dev, key_points_test]
key_points_labels_list = ['key_point', 'topic']
cleaning(key_points_datasets_list, key_points_labels_list)
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="gKwkADByDWSc" outputId="eff75eea-b857-43ec-f219-a9d80d77d063"
# Check.
# arguments_train.head()
# arguments_dev.head()
# arguments_test.head()
# key_points_train.head()
key_points_dev.head()
# key_points_test.head()
# + [markdown] id="wF_FoUnEbfHK"
# ### 2.2 **Text tokenization**
# Text tokenization consists into split a whole sentencences into meaningful pieces of it. It is achieved with the punkt tokenizer by NLTK.
# + id="tfkyF1albkaX"
# Create copies of the processed dataframes.
arguments_train_tok = arguments_train.copy()
arguments_dev_tok = arguments_dev.copy()
arguments_test_tok = arguments_test.copy()
key_points_train_tok = key_points_train.copy()
key_points_dev_tok = key_points_dev.copy()
key_points_test_tok = key_points_test.copy()
# Note: it takes a while.
from nltk.tokenize import wordpunct_tokenize
def tokenize_dataset(dataframes: list, labels: list):
for df in dataframes:
for lab in labels:
df[lab] = df[lab].apply(wordpunct_tokenize)
tokenize_dataset([arguments_train_tok, arguments_dev_tok, arguments_test_tok], labels = arguments_labels_list)
tokenize_dataset([key_points_train_tok, key_points_dev_tok, key_points_test_tok], labels = key_points_labels_list)
# + [markdown] id="7szBYrm3SAn_"
# ### 2.3 **Lenghts analysis**
# Padding is not needed for processor and the pre-trained models since they automatically takes into account different lengths, but is needed for the GloVe embeddings.
#
# In any case, length distribution studies are useful to decide if it makes sense to employ attention mechanisms.
#
# Key-points lengths are not considered since their maximum length is fixed.
# + colab={"base_uri": "https://localhost:8080/", "height": 563} id="5a-pU6gLSNP3" outputId="9e1ac1f0-4fae-40ae-86b7-6df885f82c81"
def extract_info(dataframe: pd.DataFrame) -> int:
plt.figure()
plt.title('Argument length distribution vs. Topic length distribution ')
no_token_argument = [len(sentence) for sentence in dataframe['argument']]
n_c, _, _ = plt.hist(no_token_argument, bins = [i for i in range(1, 50)], cumulative = False)
no_token_topic = [len(sentence) for sentence in dataframe['topic']]
n_e, _, _ = plt.hist(no_token_topic, bins = [i for i in range(1, 50)], cumulative = False)
return max(max(no_token_argument), max(no_token_topic))
max_tokens = max(extract_info(arguments_train_tok), extract_info(arguments_dev_tok))
print(f'The maximum found length is {max_tokens}.')
# + [markdown] id="vVmQwxuSiXiP"
# ## 3.0 **The Key-Points Matching problem**
# Given a controversial topic $T$ with a list of $m$ arguments and $n$ key
# points:
# * $A_1, A_2, \dots, A_m$;
# * $K_1, K_2, \dots, K_n$,
#
# along with their corresponding $m+n$ stances:
# * $S_1, S_2, \dots, S_{m+n}$;
#
# with $S_i \in \{−1, 1\}$, which
# imply the attack or support relationships against the topic, the task is to rank key points that have the same stance with an input argument by the matching score.
#
# This priority is dependent on both the topic and the semantic of statements.
# + [markdown] id="AmyX8LL-U0Em"
# ## 4.0 **Unsupervised approaches**
# + [markdown] id="tDr2J9xaVUcm"
# ### 4.1 **Pre-trained embeddings**
# It is possible to use pre-trained embeddings to measure the "distance" of each pair of keyword and argument.
#
# GloVe is a context independent model that computes a single embedding for each word, while BERT is a contextualized embedding model that takes the entire sentence into account.
#
# Again, we use cosine similarity to compute the match score.
#
# For each method it is possible to decide wether to consider also topic and stance.
# + [markdown] id="Rajy4_AbXPVa"
# #### 4.1.1 **GloVe embeddings**
# GloVe is an unsupervised learning algorithm for obtaining vector representations for words. Training is performed on aggregated global word-word co-occurrence statistics from a corpus, and the resulting representations showcase interesting linear substructures of the word vector space.
#
#
# The training objective of GloVe is to learn word vectors such that their dot product equals the logarithm of the words' probability of co-occurrence. Owing to the fact that the logarithm of a ratio equals the difference of logarithms, this objective associates (the logarithm of) ratios of co-occurrence probabilities with vector differences in the word vector space. Because these ratios can encode some form of meaning, this information gets encoded as vector differences as well.
#
# + [markdown] id="bKK1gosDXKiw"
# ##### 4.1.1.1 **Out-of-vocaboulary terms & embedding**
# At first some functions to handle OOVs terms and token embeddings are define. In particular:
# * `build_vocaboulary`: create a double vocaboulary from a dataframe to mantain encode and decode tokens;
# * `load_glove_embedding`: given a chosen embedding dimension, it downloads the corresponding pre-trained corpus;
# * `get_OOV_terms`: given a vocaboulary and a word listings returns the set of OOV terms;
# * `build_embedding_matrix`: returns the matrix to pass to the embedding layer as kernel initializer.
# + id="dwoPB90ZXkLc" colab={"base_uri": "https://localhost:8080/"} outputId="d3f3d5e2-45c0-46cf-b78e-aa33165c2ddc"
import gensim
import gensim.downloader as gloader
from collections import OrderedDict
from typing import OrderedDict
def build_vocabulary(dataset: pd.DataFrame, labels: list) -> (OrderedDict[int, str], OrderedDict[str, int], list):
idx2word = OrderedDict()
word2idx = OrderedDict()
curr_idx = 1 # Start at 1 since 0 is reserved for padding.
for label in labels:
for row in tqdm(dataset[label].values):
for token in row:
if token not in word2idx:
word2idx[token] = curr_idx
idx2word[curr_idx] = token
curr_idx += 1
word_listing = list(idx2word.values())
return idx2word, word2idx, word_listing
def load_glove_embedding(embedding_dimension: int) -> gensim.models.keyedvectors.KeyedVectors:
download_path = "glove-wiki-gigaword-{}".format(embedding_dimension)
# Sanity check.
try:
embedded_model = gloader.load(download_path)
except ValueError as e:
print("Invalid embedding dimension.")
raise e
return embedded_model
def get_OOV_terms(embedding_model: gensim.models.keyedvectors.KeyedVectors, word_listing: List[str]) -> list:
embedding_vocabulary = set(embedding_model.vocab.keys())
oov = set(word_listing).difference(embedding_vocabulary)
return list(oov)
def build_embedding_matrix(embedding_model: gensim.models.keyedvectors.KeyedVectors,
embedding_dimension: int,
word2idx: OrderedDict[str, int],
vocab_size: int) -> np.ndarray:
# Note: we take (vocab_size) + 1 since we start counting by 1, having reserved 0 for the padding.
embedding_matrix = np.zeros((vocab_size + 1, embedding_dimension), dtype = np.float32)
for word, idx in tqdm(word2idx.items()):
try:
embedding_vector = embedding_model[word]
except (KeyError, TypeError):
embedding_vector = np.random.uniform(low = -0.05, high = 0.05, size = embedding_dimension)
embedding_matrix[idx] = embedding_vector
return embedding_matrix
emb_dim = 300
embedded_model = load_glove_embedding(embedding_dimension = emb_dim)
# + [markdown] id="GQprH0lHYrib"
# At first the vocabularies of words and tags for the training set is created, the out-of-vocabularies (OOV) words extracted, assigning random vectors for the OOV words. The same process is repeated for the validation set, taking into account the fact that the OOV words already considered for the training set, are not OOV words for the validation set. The same was done for the test set.
#
# Note: the process must take into account arguments and key-points (two distinct dataframes).
#
# + colab={"base_uri": "https://localhost:8080/"} id="x2I-TwCsZQ5E" outputId="964c23b8-5994-490a-971c-f58e00cb054f"
# Step 0: load GloVe vocaboulary.
vocaboulary = list(embedded_model.vocab.keys())
# Step 1: creation of vocaboulary for the training set and extraction of his OOVs.
_, _, word_listing_train_arg = build_vocabulary(arguments_train_tok, ['argument'])
_, _, word_listing_train_kp = build_vocabulary(key_points_train_tok, ['key_point'])
word_listing_train = word_listing_train_arg + word_listing_train_kp
oov_terms_train = get_OOV_terms(embedded_model, word_listing_train)
print(f'There are {len(oov_terms_train)} OOVs in the training set.')
print('\n')
# Step 2: creation of vocabulary for the validation set and extraction of his OOVs, taking into account the ones already from training set.
_, _, word_listing_val_arg = build_vocabulary(arguments_dev_tok, ['argument'])
_, _, word_listing_val_kp = build_vocabulary(key_points_dev_tok, ['key_point'])
word_listing_val = word_listing_val_arg + word_listing_val_kp
oov_terms_val = get_OOV_terms(embedded_model, word_listing_val)
oov_terms_val = list(set(oov_terms_val).difference(set(oov_terms_train)))
print(f'There are {len(oov_terms_val)} new OOVs in the validation set.')
print('\n')
# Step 3: creation of vocabulary for the test set and extraction of his OOVs.
_, _, word_listing_test_arg = build_vocabulary(arguments_test_tok, ['argument'])
_, _, word_listing_test_kp = build_vocabulary(key_points_test_tok, ['key_point'])
word_listing_test = word_listing_test_arg + word_listing_test_kp
oov_terms_test = get_OOV_terms(embedded_model, word_listing_test)
oov_terms_test = list(set(oov_terms_test).difference((set(oov_terms_train).union(set(oov_terms_val)))))
print(f'There are {len(oov_terms_test)} new OOVs in the test set.')
print('\n')
# Step 4: merge vocabulary and OOVs.
vocaboulary = np.concatenate((vocaboulary, oov_terms_train, oov_terms_val, oov_terms_test))
# Step 5: creation of a comprehensive dictionary.
word2idx = dict(zip(vocaboulary, range(1, len(vocaboulary) + 1))) # Start from 1 since we reserve 0 for padding.
# Step 6: creation of the whole embedding matrix.
embedding_matrix = build_embedding_matrix(embedded_model, emb_dim, word2idx, len(word2idx))
print('\n')
print("Embedding matrix shape: {}.".format(embedding_matrix.shape))
# + [markdown] id="WnLsRwA1dawp"
# ##### 4.1.1.2 **Dataset encoding**
# The datasets are encoded and aligned with the respective identifiers.
# + id="8pxuQpUteQxK"
def encode_sentence(sentence, word2idx):
return [word2idx[words] for words in sentence]
def encode_dataframe(dataframe, word2idx, columns):
encoded_dataframe = pd.DataFrame(columns = columns)
for column in columns:
encoded_dataframe[column] = dataframe[column].apply(lambda sentence: encode_sentence(sentence, word2idx))
return encoded_dataframe
# The dataset is encoded and then the id columns restored.
arguments_train_enc = encode_dataframe(arguments_train_tok, word2idx, arguments_labels_list)
arguments_train_enc['arg_id'] = arguments_train['arg_id']
arguments_dev_enc = encode_dataframe(arguments_dev_tok, word2idx, arguments_labels_list)
arguments_dev_enc['arg_id'] = arguments_dev['arg_id']
arguments_test_enc = encode_dataframe(arguments_test_tok, word2idx, arguments_labels_list)
arguments_test_enc['arg_id'] = arguments_test['arg_id']
key_points_train_enc = encode_dataframe(key_points_train_tok, word2idx, key_points_labels_list)
key_points_train_enc['key_point_id'] = key_points_train['key_point_id']
key_points_dev_enc = encode_dataframe(key_points_dev_tok, word2idx, key_points_labels_list)
key_points_dev_enc['key_point_id'] = key_points_dev['key_point_id']
key_points_test_enc = encode_dataframe(key_points_test_tok, word2idx, key_points_labels_list)
key_points_test_enc['key_point_id'] = key_points_test['key_point_id']
# + [markdown] id="sRs3KYRFX9GU"
# ##### 4.1.1.3 **Labels alignment**
# The pairs of argument and key-point pairs identifier are extracted. With these pairs the encoded arguments and key-points are extracted.
#
# Topic and stance gives the “main context”. The argument and the key-point can also be integrated with this main context and then matched.
# + id="ROmFLDWeYN5J"
from keras.preprocessing.sequence import pad_sequences
def flatten(t: list) -> list:
# Flatten a nested list.
return [item for sublist in t for item in sublist]
indexes = [i for i in range(len(labels_test))]
# Generations of pairs arg-kp of the test set.
arg_id = [labels_test['arg_id'][k] for k in indexes]
kp_id = [labels_test['key_point_id'][k] for k in indexes]
# Arguments extraction.
arguments = flatten([arguments_test_enc.loc[arguments_test_enc['arg_id'] == arg]['argument'].tolist() for arg in arg_id])
arguments = pad_sequences(arguments, maxlen = max_tokens, padding = 'post')
# Key-points extraction.
keypoints = flatten([key_points_test_enc.loc[key_points_test_enc['key_point_id'] == kp]['key_point'].tolist() for kp in kp_id])
keypoints = pad_sequences(keypoints, maxlen = max_tokens, padding = 'post')
# Topics extraction.
topics = flatten([arguments_test_enc.loc[arguments_test_enc['arg_id'] == arg]['topic'].tolist() for arg in arg_id])
topics = pad_sequences(keypoints, maxlen = max_tokens, padding = 'post')
# + [markdown] id="Qk9m4lFnbyE1"
# ##### 4.1.1.4 **GloVe application**
# GloVe embedding matrix is applied, unrolling the vectors in order to apply the cosine similarity.
#
# + id="XAHdckHoZhnP"
# Application of the embedding matrix.
arguments_emb = embedding_matrix[arguments].reshape((len(labels_test), -1))
keypoints_emb = embedding_matrix[keypoints].reshape((len(labels_test), -1))
# Concatenation of the topic and application of the embedding matrix.
arguments_topic = np.concatenate((arguments, topics), axis = -1)
keypoints_topic = np.concatenate((keypoints, topics), axis = -1)
arguments_topic_emb = embedding_matrix[arguments_topic].reshape((len(labels_test), -1))
keypoints_topic_emb = embedding_matrix[keypoints_topic].reshape((len(labels_test), -1))
# + [markdown] id="duCMkydXatVA"
# ##### 4.1.1.5 **Cosine similarity calculation and evaluation**
# Cosine similarity is evaluated wether if the topic is considered or not. The general performances are worsened if the topic is taken into account, but the average precision score is higher in the latter case.
#
# The cause can be the redundacy in of the such feature.
#
# + id="r-UkjJQQeeaI"
def cos_sim(vec1: np.ndarray, vec2: np.ndarray) -> np.ndarray:
return np.dot(vec1, vec2)/(np.linalg.norm(vec1) * np.linalg.norm(vec2))
# Cosine similarity is calculated and then rounded.
predictions_glove_no_topic = np.rint([cos_sim(arguments_emb[i], keypoints_emb[i]) for i in range(len(indexes))])
predictions_glove_with_topic = np.rint([cos_sim(arguments_topic_emb[i], keypoints_topic_emb[i]) for i in range(len(indexes))])
true_values = labels_test['label'].to_numpy()
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="nJH6U2CYgqrR" outputId="83fd7526-2e7b-4ea2-e133-6bf9db7387dc"
from sklearn.metrics import classification_report, average_precision_score, PrecisionRecallDisplay, balanced_accuracy_score, ConfusionMatrixDisplay
print('Metrics when topic is not considered:')
print(classification_report(true_values, predictions_glove_no_topic))
print(f'The average precision score is: {average_precision_score(true_values, predictions_glove_no_topic)}.')
print(f'The balanced accuracy precision score is: {balanced_accuracy_score(true_values, predictions_glove_no_topic)}.')
print(f'The confusion matrix is: ')
ConfusionMatrixDisplay.from_predictions(y_true = true_values, y_pred = predictions_glove_no_topic);
print('\n')
print('Metrics when topic is considered:')
print(classification_report(true_values, predictions_glove_with_topic))
print(f'The average precision score is: {average_precision_score(true_values, predictions_glove_with_topic)}.')
print(f'The balanced accuracy score is: {balanced_accuracy_score(true_values, predictions_glove_with_topic)}.')
print(f'The confusion matrix is: ')
ConfusionMatrixDisplay.from_predictions(true_values, predictions_glove_with_topic);
# + id="bT_zTa69n05W" colab={"base_uri": "https://localhost:8080/", "height": 578} outputId="a3643ee1-688a-4c51-e918-d2f7a20b1340"
print('Metrics when topic is not considered:')
PrecisionRecallDisplay.from_predictions(true_values, [cos_sim(arguments_emb[i], keypoints_emb[i]) for i in range(len(indexes))]);
print('Metrics when topic is considered:')
PrecisionRecallDisplay.from_predictions(true_values, [cos_sim(arguments_topic_emb[i], keypoints_topic_emb[i]) for i in range(len(indexes))]);
# + [markdown] id="_iJy7Ws8pYMw"
# #### 4.1.2 **BERT embeddings**
# BERT (Bidirectional Encoder Representations from Transformers) provides dense vector representations for natural language by using a deep, pre-trained neural network with the Transformer architecture.
#
# BERT is a contextualized embedding model that takes the entire sentence into account. The following implementation takes automatically into account padding and encoding.
#
# BERT provides three different kinds of output:
# * `pooled_output`: pooled output of the entire sequence with shape `[batch_size, hidden_size]`. Can be used as sentence representation;
# * `sequence_output`: representations of every token in the input sequence with shape `[batch_size, max_sequence_length, hidden_size]`;
# * `encoder_outputs`: a list of 12 tensors of shapes `[batch_size, sequence_length, hidden_size]` with the outputs of the $i$-th Transformer block.
#
# For sake of simplicity (and RAM occupancy) the pooled output is computed with a lighter implementation.
#
# + id="8wbJPINZrO6z"
# Faster implementation to work with pooled outputs.
pretrained_bert_model = sentence_transformers.SentenceTransformer('all-mpnet-base-v2');
# + id="JrimRJtOqQxz"
indexes = [i for i in range(len(labels_test))]
# Generations of pairs arg-kp of the test set.
arg_id = [labels_test['arg_id'][k] for k in indexes]
kp_id = [labels_test['key_point_id'][k] for k in indexes]
# Arguments extraction.
arguments = flatten([arguments_test.loc[arguments_test['arg_id'] == arg]['argument'].tolist() for arg in arg_id])
encoded_arguments = pretrained_bert_model.encode(arguments)
# Key-points extraction.
keypoints = flatten([key_points_test.loc[key_points_test['key_point_id'] == kp]['key_point'].tolist() for kp in kp_id])
encoded_keypoints = pretrained_bert_model.encode(keypoints)
# Topics extraction.
topics = flatten([arguments_test.loc[arguments_test['arg_id'] == arg]['topic'].tolist() for arg in arg_id])
encoded_topics = pretrained_bert_model.encode(topics)
# + id="Dkx8q8i905N9"
arguments_topic = np.concatenate((encoded_arguments, encoded_topics), axis = -1)
keypoints_topic = np.concatenate((encoded_keypoints, encoded_topics), axis = -1)
# + [markdown] id="zaU2Yk3t3LJF"
# ##### 4.1.2.1 **Cosine similarity calculation and evaluation**
# Cosine similarity is evaluated wether if the topic is considered or not. The general performances are worsened if the topic is taken into account. The cause can be due to the redundacy of the such feature.
#
# + id="cSQBuZEk3UJ7"
def cos_sim(vec1: np.ndarray, vec2: np.ndarray) -> np.ndarray:
return np.dot(vec1, vec2)/(np.linalg.norm(vec1) * np.linalg.norm(vec2))
# Cosine similarity is calculated and then rounded.
predictions_bert_no_topic = np.rint([cos_sim(encoded_arguments[i], encoded_keypoints[i]) for i in range(len(indexes))])
predictions_bert_with_topic = np.rint([cos_sim(arguments_topic[i], keypoints_topic[i]) for i in range(len(indexes))])
true_values = labels_test['label'].to_numpy()
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="faa354f4-bb6a-46e3-8ca5-bd923c3b5f8a" id="U_xlJIbu3UJ9"
from sklearn.metrics import classification_report, average_precision_score, PrecisionRecallDisplay, ConfusionMatrixDisplay
print('Metrics when topic is not considered:')
print(classification_report(true_values, predictions_bert_no_topic))
print(f'The average precision score is: {average_precision_score(true_values, predictions_bert_no_topic)}.')
print(f'The balanced accuracy precision score is: {balanced_accuracy_score(true_values, predictions_bert_no_topic)}.')
print(f'The confusion matrix is: ')
ConfusionMatrixDisplay.from_predictions(true_values, predictions_bert_no_topic);
print('\n')
print('Metrics when topic is considered:')
print(classification_report(true_values, predictions_bert_with_topic))
print(f'The average precision score is: {average_precision_score(true_values, predictions_bert_with_topic)}.')
print(f'The balanced accuracy score is: {balanced_accuracy_score(true_values, predictions_bert_with_topic)}.')
print(f'The confusion matrix is: ')
ConfusionMatrixDisplay.from_predictions(true_values, predictions_bert_with_topic);
# + id="-sRsu3EOoCws" colab={"base_uri": "https://localhost:8080/", "height": 578} outputId="abb0fed8-953e-4ed7-ae19-8ca893b08d80"
print('Metrics when topic is not considered:')
PrecisionRecallDisplay.from_predictions(true_values, [cos_sim(encoded_arguments[i], encoded_keypoints[i]) for i in range(len(indexes))]);
print('Metrics when topic is considered:')
PrecisionRecallDisplay.from_predictions(true_values, [cos_sim(arguments_topic[i], keypoints_topic[i]) for i in range(len(indexes))]);
# + [markdown] id="3LwdSHQdVDwm"
# ### 4.2 **Tf-idf**
# In order to assess the role of lexical overlap in the matching task, we represent each argument and key point as tf-idf weighted word vectors and use their cosine similarity as the match score.
# + id="gbRR0_EIUOSX"
from sklearn.feature_extraction.text import TfidfVectorizer
vectorizer = TfidfVectorizer()
indexes = [i for i in range(len(labels_test))]
# Generations of pairs arg-kp of the test set.
arg_id = [labels_test['arg_id'][k] for k in indexes]
kp_id = [labels_test['key_point_id'][k] for k in indexes]
# Arguments extraction.
arguments = flatten([arguments_test.loc[arguments_test['arg_id'] == arg]['argument'].tolist() for arg in arg_id])
# Key-points extraction.
keypoints = flatten([key_points_test.loc[key_points_test['key_point_id'] == kp]['key_point'].tolist() for kp in kp_id])
# Topics extraction.
topics = flatten([arguments_test.loc[arguments_test['arg_id'] == arg]['topic'].tolist() for arg in arg_id])
# Pairs of arguments and key-points extraction
args_kps = [vectorizer.fit_transform([arguments[i], keypoints[i]]).toarray() for i in indexes]
args_kps_topic = [vectorizer.fit_transform([arguments[i] + topics[i], keypoints[i] + topics[i]]).toarray() for i in indexes]
# + [markdown] id="yhK8ojqjWoS3"
# ##### 4.2.2 **Cosine similarity calculation and evaluation**
# Cosine similarity is evaluated wether if the topic is considered or not. The general performances are worsened if the topic is taken into account. The cause can be due to the redundacy of the such feature.
#
# + id="dL4BMIbQUg-x"
predictions_tfidf = np.rint([cos_sim(args_kps[i][0], args_kps[i][1]) for i in indexes])
predictions_tfidf_topic = np.rint([cos_sim(args_kps_topic[i][0], args_kps_topic[i][1]) for i in indexes])
true_values = labels_test['label'].to_numpy()
# + id="Zd_ni3nlvHFX" colab={"base_uri": "https://localhost:8080/", "height": 297} outputId="9fc4e782-4347-4ad8-f3c4-0188cec2d6d8"
PrecisionRecallDisplay.from_predictions(true_values, [cos_sim(args_kps[i][0], args_kps[i][1]) for i in indexes])
PrecisionRecallDisplay.from_predictions(true_values, [cos_sim(args_kps_topic[i][0], args_kps_topic[i][1]) for i in indexes])
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="3iya3dF4WzXJ" outputId="20438bca-3f6a-4f8d-9fb1-63fb65d6c56c"
from sklearn.metrics import classification_report, average_precision_score, PrecisionRecallDisplay, ConfusionMatrixDisplay
print('Metrics when topic is not considered:')
print(classification_report(true_values, predictions_tfidf))
print(f'The average precision score is: {average_precision_score(true_values, predictions_tfidf)}.')
print(f'The accuracy precision score is: {balanced_accuracy_score(true_values, predictions_tfidf)}.')
print(f'The confusion matrix is: ')
ConfusionMatrixDisplay.from_predictions(true_values, predictions_tfidf);
print('\n')
print('Metrics when topic is considered:')
print(classification_report(true_values, predictions_tfidf_topic))
print(f'The average precision score is: {average_precision_score(true_values, predictions_tfidf_topic)}.')
print(f'The accuracy score is: {balanced_accuracy_score(true_values, predictions_tfidf_topic)}.')
print(f'The confusion matrix is: ')
ConfusionMatrixDisplay.from_predictions(true_values, predictions_tfidf_topic);
# + [markdown] id="SieCEoPSUd8J"
# ## 5.0 **Supervised approaches**
# + [markdown] id="c_HJIvZO9986"
# ### 5.1 **Metrics**
# + id="r2m21Av_4Bjs"
from keras import backend as K
def recall_m(y_true, y_pred):
true_positives = K.sum(K.round(K.clip(y_true * y_pred, 0, 1)))
possible_positives = K.sum(K.round(K.clip(y_true, 0, 1)))
recall = true_positives / (possible_positives + K.epsilon())
return recall
def precision_m(y_true, y_pred):
true_positives = K.sum(K.round(K.clip(y_true * y_pred, 0, 1)))
predicted_positives = K.sum(K.round(K.clip(y_pred, 0, 1)))
precision = true_positives / (predicted_positives + K.epsilon())
return precision
def f1_m(y_true, y_pred):
precision = precision_m(y_true, y_pred)
recall = recall_m(y_true, y_pred)
return 2 * ((precision * recall) / (precision + recall + K.epsilon()))
# + [markdown] id="3W41M7vbfG2c"
# ### 5.2 **Neural architecture #1**
# The experimented neural architecture is composed by:
# * *Stance encoder*: encode the stance value (1 or -1) employing a fully-connected network with no activation function to map the scalar input to a $N$-dimensional vector space;
# * *BERT*: extract the contextualized representation for textual inputs;
# * A context integration layer is done by stacking stance encoding, topic encoding and statements encoding;
# * *Statement encoder*: another fully-connected network on top of the context integration layer to get the final $D$-dimensional embeddings for key points or arguments.
#
# <div>
# <center>
# <img src = "https://i.ibb.co/BPGXRyJ/architecture.jpg" width = "800"/>
# <center>
# </div>
#
# The extraction of contextualized vector representation is already embedded in the generator.
# + id="UYQRPK4M-GT2"
# Load the pre-trained BERT model.
bert_preprocess_model = hub.KerasLayer("https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/3")
bert_model = hub.KerasLayer("https://tfhub.dev/tensorflow/bert_en_uncased_L-24_H-1024_A-16/4", trainable = False)
# Hidden states of BERT.
layers = 1024
# + id="TIvTHWF4Y4E8"
from tensorflow.keras.layers import Dense, TimeDistributed, Input, concatenate, add, average, Dropout, dot, Lambda, LSTM, Bidirectional
from tensorflow.keras.models import Model
from tensorflow.keras.callbacks import EarlyStopping, ModelCheckpoint
from tensorflow.keras.initializers import Constant, glorot_normal
from tensorflow.keras.optimizers import RMSprop, Adamax, SGD, Adam
from tensorflow.keras.optimizers.schedules import ExponentialDecay, CosineDecayRestarts
from tensorflow.keras import regularizers
from tensorflow.keras.metrics import Precision, Recall
from tensorflow.keras.losses import BinaryCrossentropy, CosineSimilarity
# Note: the learning rate is not parametrized since we use a scheduler.
def classifier(no_last_state: int,
dropout: float = 0.5, recurrent_dropout: float = 0.25,
initial_learning_rate: float = 0.0001, N_stance: int = 16) -> Model:
hidden_states = 32
''' Network structure '''
# Inputs.
topic_input = Input(shape = (no_last_state * layers),
#batch_size = batch_size,
name = 'topic_input')
argument_input = Input(shape = (no_last_state * layers),
#batch_size = batch_size,
name = 'argument_input')
keypoint_input = Input(shape = (no_last_state * layers),
#batch_size = batch_size,
name = 'keypoint_input')
stance_input = Input(shape = (1, ),
#batch_size = batch_size,
name = 'stance_input')
# Stance encoder.
encoded_stance = Dense(units = N_stance, activation = None, name = 'fcn')(stance_input)
# Context integration layer.
argument_integration = concatenate([encoded_stance, topic_input, argument_input])
keypoint_integration = concatenate([encoded_stance, topic_input, keypoint_input])
# Statement encoder.
mlp0 = Dense(units = (hidden_states / 2), activation = 'selu',
kernel_regularizer = regularizers.l1_l2(l1 = 1e-5, l2 = 1e-4),
bias_regularizer = regularizers.l2(1e-4),
activity_regularizer = regularizers.l2(1e-5),
name = 'st_enc_0')
mlp1 = Dropout(rate = dropout, name = 'st_enc_drp')
mlp2 = Dense(units = hidden_states, activation = 'selu',
kernel_regularizer = regularizers.l1_l2(l1 = 1e-5, l2 = 1e-4),
bias_regularizer = regularizers.l2(1e-4),
activity_regularizer = regularizers.l2(1e-5),
name = 'st_enc_2')
mlp0_argument = mlp0(argument_integration)
mlp1_argument = mlp1(mlp0_argument)
argument_output = mlp2(mlp1_argument)
mlp0_keypoint = mlp0(keypoint_integration)
mlp1_keypoint = mlp1(mlp0_keypoint)
keypoint_output = mlp2(mlp1_keypoint)
# Cosine similarity.
cos_sim = dot(inputs = [argument_output, keypoint_output], axes = -1, normalize = True)
# Rescale in range [0,1].
output = Lambda(lambda x: (x + 1) / 2)(cos_sim)
end_to_end = Model(inputs = (topic_input, argument_input, keypoint_input, stance_input), outputs = output)
# Compile the model.
scheduler_exp = ExponentialDecay(initial_learning_rate,
decay_steps = 100000,
decay_rate = 0.96,
staircase = True)
scheduler_cos = CosineDecayRestarts(initial_learning_rate,
first_decay_steps = 1000)
end_to_end.compile(loss = focal_loss.BinaryFocalLoss(gamma = 2),
optimizer = Adamax(learning_rate = scheduler_cos),
metrics = ['binary_accuracy', precision_m, recall_m, f1_m])
# Check if the structure is correct.
end_to_end.summary()
return end_to_end
# + colab={"base_uri": "https://localhost:8080/"} id="0PI8tc-49BEF" outputId="3dd34a4a-b389-4070-ab5f-86d1de30c15e"
# Number of last states to concatenate out.
l = 4
matcher = classifier(no_last_state = l, initial_learning_rate = 0.00005)
# + colab={"base_uri": "https://localhost:8080/", "height": 340} id="nTBKAxYD9F80" outputId="e3a01696-7c36-4189-861e-e7e4b427e126"
from tensorflow.keras.utils import plot_model
plot_model(matcher, show_shapes = True)
# + [markdown] id="fXgUI_zQH8Dg"
# #### 5.2.1 **Data generator with vector representation**
# To avoid high memory usage a data generator is implemented.
#
# The generator is initialized with:
# * A *shuffle* parameter to add more robustness to the batches of data;
# * A *batch_size* parameter to decide the size of the aforementioned batches;
# * The dataset parameters;
# * The *embedder* parameter to pass the chosen transformer;
# * A *no_last_state* parameter to decide how many states of the transformer's output concatenate to obtain the representation.
#
# The data is handled in order to parse and align the pairs in the labels dataframes. Then, topics, arguments and keypoints are encoded through BERT, and averaged over the last $l$ states.
#
# Zero-Shot learning is a type of machine learning technique, where the model is used without fine-tuning on a particular task.
#
# A single datapoint is output in the form of: `([topic, argument, keypoint, stance], label)`.
# + id="4cPWSRvShDuG"
from tensorflow.keras.utils import Sequence
def flatten(t: list) -> list:
# Flatten a nested list.
return [item for sublist in t for item in sublist]
class DataGenerator(Sequence):
def __init__(self, shuffle: bool, batch_size: int,
arguments_df: pd.DataFrame, key_points_df: pd.DataFrame, labels_df: pd.DataFrame,
preprocessor, pretrained_model, no_last_state: int):
self.shuffle = shuffle
self.batch_size = batch_size
self.no_last_state = no_last_state
self.preprocessor = preprocessor
self.pretrained_model = pretrained_model
self.arguments_df = arguments_df.copy()
self.key_points_df = key_points_df.copy()
self.labels_df = labels_df.copy()
# Labels.
self.labels = labels_df['label'].values
# Stance.
self.stance = arguments_df['stance'].tolist()
self.length = len(labels_df)
self.indexes = np.arange(self.length)
self.on_epoch_end()
self.n = 0
def on_epoch_end(self):
# Shuffle the indexes to have more robust batches.
if self.shuffle:
np.random.shuffle(self.indexes)
def __len__(self):
# Length of the batch.
return int(np.ceil(self.length / self.batch_size))
def __getitem__(self, idx):
'''Generate one batch of data.'''
# Select the indexes of the batch.
indexes = self.indexes[idx * self.batch_size : (idx + 1) * self.batch_size]
# For each row in the labels dataframe we have the pair argument and keypoint. These two lines retrieve the pairs in the batch
arg_id = [self.labels_df['arg_id'][k] for k in indexes]
kp_id = [self.labels_df['key_point_id'][k] for k in indexes]
''' Warning: it produces nested list! '''
# Take the topic from the arguments dataframe that match the given argument, already paired before.
topics = flatten([self.arguments_df.loc[self.arguments_df['arg_id'] == arg]['topic'].tolist() for arg in arg_id])
# Preprocces add [CLS] as 0 and [SEP] as 2 and pads with 1s.
preprocessed_topic = self.preprocessor(topics)
# Encode the topics and picks the last four states of the [CLS].
encoded_topic = np.array(self.pretrained_model(preprocessed_topic)['encoder_outputs'][-self.no_last_state:])[:,:,0,:]
# Swap axes to have the batch size at the beginning.
encoded_topic = np.swapaxes(encoded_topic, 0, 1)
encoded_topic = np.reshape(encoded_topic, (-1, self.no_last_state * layers))
# Take the arguments from the arguments dataframe that match the given argument, already paired before.
arguments = flatten([self.arguments_df.loc[self.arguments_df['arg_id'] == arg]['argument'].tolist() for arg in arg_id])
# Preprocces add [CLS] as 0 and [SEP] as 2 and pads with 1s.
preprocessed_argument = self.preprocessor(arguments)
# Encode the arguments and picks the last four states of the [CLS].
encoded_argument = np.array(self.pretrained_model(preprocessed_argument)['encoder_outputs'][-self.no_last_state:])[:,:,0,:]
# Swap axes to have the batch size at the beginning.
encoded_argument = np.swapaxes(encoded_argument, 0, 1)
encoded_argument = np.reshape(encoded_argument, (-1, self.no_last_state * layers))
# Take the keypoint from the keypoints dataframe that match the given argument, already paired before.
keypoints = flatten([self.key_points_df.loc[self.key_points_df['key_point_id'] == kp]['key_point'].tolist() for kp in kp_id])
# Preprocces add [CLS] as 0 and [SEP] as 2 and pads with 1s.
preprocessed_keypoint = self.preprocessor(keypoints)
# Encode the keypoints and picks the last four states of the [CLS].
encoded_keypoint = np.array(self.pretrained_model(preprocessed_keypoint)['encoder_outputs'][-self.no_last_state:])[:,:,0,:]
# Swap axes to have the batch size at the beginning.
encoded_keypoint = np.swapaxes(encoded_keypoint, 0, 1)
encoded_keypoint = np.reshape(encoded_keypoint, (-1, self.no_last_state * layers))
# Take the stance from the arguments dataframe that match the given argument, already paired before.
stance = np.array([self.arguments_df.loc[self.arguments_df['arg_id'] == arg]['stance'].tolist() for arg in arg_id])
labels = np.array([self.labels[k] for k in indexes])
features = [encoded_topic,
encoded_argument,
encoded_keypoint,
stance]
return features, labels
def __next__(self):
if self.n >= self.__len__():
self.n = 0
result = self.__getitem__(self.n)
self.n += 1
return result
# + id="VHgyaB52rQLs"
# Batch size.
batch_size = 32
# Instantiate generators.
train_gen = DataGenerator(shuffle = True, batch_size = batch_size,
arguments_df = arguments_train, key_points_df = key_points_train, labels_df = labels_train,
preprocessor = bert_preprocess_model, pretrained_model = bert_model, no_last_state = l)
dev_gen = DataGenerator(shuffle = True, batch_size = batch_size,
arguments_df = arguments_dev, key_points_df = key_points_dev, labels_df = labels_dev,
preprocessor = bert_preprocess_model, pretrained_model = bert_model, no_last_state = l)
# No need to shuffle test data.
test_gen = DataGenerator(shuffle = False, batch_size = batch_size,
arguments_df = arguments_test, key_points_df = key_points_test, labels_df = labels_test,
preprocessor = bert_preprocess_model, pretrained_model = bert_model, no_last_state = l)
# + [markdown] id="CaTisB1C-IQo"
# #### 5.2.2 **Training**
# + id="rkEUK8lZ-v82"
checkpoint = [EarlyStopping(monitor = 'val_loss', patience = 10, verbose = 1, mode = 'auto'),
ModelCheckpoint(filepath = "/content/drive/MyDrive/[NLP]/project/models/adamax_cd_hu64_bs128.h5",
monitor = 'val_loss', verbose = 1, save_best_only = True)]
# + id="qIJVDcWRA8pJ"
# Train the model.
matcher.fit(x = train_gen,
validation_data = dev_gen,
epochs = 100, callbacks = checkpoint)
# + id="f67V2J67Kv8t"
# Manual saving in case Colab crashes.
"""
from tensorflow.keras.models import save_model
save_model(reloaded_model, '/content/drive/MyDrive/[NLP]/project/models/adamax_cd_hu64_bs128.h5')
"""
# + id="DtgTzVwrSnTk"
# Reload the model and restart the training.
from tensorflow.keras.models import load_model
reloaded_model = load_model(filepath = '/content/drive/MyDrive/[NLP]/project/models/adamax_cd_hu64_bs128.h5',
custom_objects = {'precision_m': precision_m, 'recall_m': recall_m, 'f1_m': f1_m})
# To change optimizers in further training.
"""
scheduler_cos = CosineDecayRestarts(initial_learning_rate = 0.000001, first_decay_steps = 1000)
reloaded_model.compile(loss = focal_loss.BinaryFocalLoss(gamma = 2),
optimizer = Adamax(learning_rate = scheduler_cos),
metrics = ['binary_accuracy', precision_m, recall_m, f1_m])
"""
reloaded_model.summary()
reloaded_model.fit(x = train_gen,
validation_data = dev_gen,
epochs = 100, callbacks = checkpoint)
# + [markdown] id="xOy9zQVuWhFh"
# #### 5.2.3 **Classifier evaluation**
# + colab={"base_uri": "https://localhost:8080/"} id="64jNFl101W1r" outputId="d630f5ed-67e3-4900-951b-c0b91f368d6e"
from tensorflow.keras.models import load_model
reloaded_model = load_model(filepath = '/content/drive/MyDrive/[NLP]/project/models/adamax_cd_hu64_bs128.h5',
custom_objects = {'precision_m': precision_m, 'recall_m': recall_m, 'f1_m': f1_m})
reloaded_model.summary()
# + id="bUehIDDFr-2Z"
# Sample weighting.
total_samples = len(labels_test['label'].to_numpy())
pos_samples = sum(labels_test['label'].to_numpy())
neg_samples = total_samples - pos_samples
pos_percentage = pos_samples/total_samples
neg_percentage = neg_samples/total_samples
weights = []
for p in labels_test['label'].to_numpy():
if p == 1:
weights.append(pos_percentage)
else:
weights.append(neg_percentage)
# + id="PmyselQdZS5x" colab={"base_uri": "https://localhost:8080/", "height": 761} outputId="374f0392-a953-4ca6-9c24-c9cb76b10192"
from sklearn.metrics import accuracy_score, classification_report, f1_score, recall_score, precision_score, PrecisionRecallDisplay, roc_curve, balanced_accuracy_score, ConfusionMatrixDisplay
def evaluate_classifier(model: tf.keras.Model, test_set_generator, true_labels):
predictions = model.predict(test_set_generator)
# Threshold tuning.
fpr, tpr, thresholds = roc_curve(true_labels, predictions)
thr = thresholds[np.argmin(np.abs(fpr + tpr - 1))]
predictions_thr = np.zeros(predictions.shape)
predictions_thr[predictions >= thr] = 1
print(classification_report(predictions_thr,
true_labels,
#sample_weight = weights
))
print(f'The average precision score is: {average_precision_score(true_labels, predictions_thr)}.')
print(f'The balanced accuracy score is: {balanced_accuracy_score(true_labels, predictions_thr)}.')
print(f'The tuned threshold is: {thr}.')
print(f'The confusion matrix is: ')
ConfusionMatrixDisplay.from_predictions(true_values, predictions_thr)
PrecisionRecallDisplay.from_predictions(true_labels, predictions)
evaluate_classifier(reloaded_model, test_gen, labels_test['label'].to_numpy())
# + [markdown] id="nK-dTPyNcarV"
# ### 5.3 **Neural architecture #2**
# The extraction of contextualized vector representation is already embedded in the generator.
# Such outputs are reshaped and encoded with a fully connected layer and then the cosine similarity of the encoded vectors is computed.
#
# <div>
# <center>
# <img src = "https://i.ibb.co/3hpvmNw/acrch-Tiny.jpg" width = "800"/>
# <center>
# </div>
#
# The extraction of contextualized vector representation is already embedded in the generator.
# + id="8Dg3F-lbgwBL"
# Load the pre-trained BERT model.
bert_preprocess_model = hub.KerasLayer("https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/3")
bert_model = hub.KerasLayer("https://tfhub.dev/tensorflow/bert_en_uncased_L-24_H-1024_A-16/4", trainable = False)
# Hidden states of BERT.
layers = 1024
# + id="zBNEuRyfcie9"
from tensorflow.keras.layers import Dense, Reshape, Input, concatenate, add, average, Dropout, dot, Lambda, LSTM, Bidirectional, TimeDistributed
from tensorflow.keras.models import Model
from tensorflow.keras.callbacks import EarlyStopping, ModelCheckpoint
from tensorflow.keras.initializers import Constant, glorot_normal
from tensorflow.keras.optimizers import RMSprop, Adamax, SGD, Adam
from tensorflow.keras.optimizers.schedules import ExponentialDecay, CosineDecayRestarts
from tensorflow.keras import regularizers
from tensorflow.keras.metrics import Precision, Recall
from tensorflow.keras.losses import BinaryCrossentropy, CosineSimilarity
# Note: the learning rate is not parametrized since we use a scheduler.
def classifier(no_transformers: int,
dropout: float = 0.5, recurrent_dropout: float = 0.25,
initial_learning_rate: float = 0.0001) -> Model:
hidden_states = 32
# Inputs.
argument_input = Input(shape = (no_transformers, layers),
name = 'argument_input')
keypoint_input = Input(shape = (no_transformers, layers),
name = 'keypoint_input')
reshaper = Reshape((-1, ))
argument = reshaper(argument_input)
keypoint = reshaper(keypoint_input)
encoder = Dense(units = hidden_states, activation = 'gelu',
kernel_regularizer = regularizers.l1_l2(l1 = 1e-5, l2 = 1e-4),
bias_regularizer = regularizers.l2(1e-4),
activity_regularizer = regularizers.l2(1e-5),
name = 'encoder')
dropout = Dropout(rate = dropout, name = 'dropout')
argument_enc = encoder(argument)
argument_enc = dropout(argument_enc)
keypoint_enc = encoder(keypoint)
keypoint_enc = dropout(keypoint_enc)
# Cosine similarity.
cos_sim = dot(inputs = [argument_enc, keypoint_enc], axes = -1, normalize = True)
# Rescale in range [0,1].
output = Lambda(lambda x: (x + 1) / 2)(cos_sim)
end_to_end = Model(inputs = (argument_input, keypoint_input), outputs = output)
# Compile the model.
scheduler_exp = ExponentialDecay(initial_learning_rate,
decay_steps = 100000,
decay_rate = 0.96,
staircase = True)
scheduler_cos = CosineDecayRestarts(initial_learning_rate,
first_decay_steps = 1000)
end_to_end.compile(loss = focal_loss.BinaryFocalLoss(gamma = 2),
optimizer = Adamax(learning_rate = scheduler_cos),
metrics = ['binary_accuracy', precision_m, recall_m, f1_m])
# Check if the structure is correct.
end_to_end.summary()
return end_to_end
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="tGYSuf3Jk9qx" outputId="1e37f659-8e46-43d3-de3c-f5e7481f7896"
matcher_two = classifier(no_transformers = 24, initial_learning_rate = 0.00005)
from tensorflow.keras.utils import plot_model
plot_model(matcher_two, show_shapes = True)
# + [markdown] id="JfWoEPiUly8-"
# #### 5.3.1 **Data generator with vector representation**
# To avoid high memory usage a data generator is implemented.
#
# The generator is initialized with:
# * A *shuffle* parameter to add more robustness to the batches of data;
# * A *batch_size* parameter to decide the size of the aforementioned batches;
# * The dataset parameters;
# * The *embedder* parameter to pass the chosen transformer.
#
# The data is handled in order to parse and align the pairs in the labels dataframes. Then, topics, arguments and keypoints are encoded through BERT.
#
# Zero-Shot learning is a type of machine learning technique, where the model is used without fine-tuning on a particular task.
#
# A single datapoint is output in the form of: `([argument, keypoint], label)`.
# + id="uFHNW8t5iuSF"
from tensorflow.keras.utils import Sequence
def flatten(t: list) -> list:
# Flatten a nested list.
return [item for sublist in t for item in sublist]
class DataGeneratorReduced(Sequence):
def __init__(self, shuffle: bool, batch_size: int,
arguments_df: pd.DataFrame, key_points_df: pd.DataFrame, labels_df: pd.DataFrame,
preprocessor, pretrained_model):
self.shuffle = shuffle
self.batch_size = batch_size
self.preprocessor = preprocessor
self.pretrained_model = pretrained_model
self.arguments_df = arguments_df.copy()
self.key_points_df = key_points_df.copy()
self.labels_df = labels_df.copy()
# Labels.
self.labels = labels_df['label'].values
# Stance.
self.stance = arguments_df['stance'].tolist()
self.length = len(labels_df)
self.indexes = np.arange(self.length)
self.on_epoch_end()
self.n = 0
def on_epoch_end(self):
# Shuffle the indexes to have more robust batches.
if self.shuffle:
np.random.shuffle(self.indexes)
def __len__(self):
# Length of the batch.
return int(np.ceil(self.length / self.batch_size))
def __getitem__(self, idx):
'''Generate one batch of data.'''
# Select the indexes of the batch.
indexes = self.indexes[idx * self.batch_size : (idx + 1) * self.batch_size]
# For each row in the labels dataframe we have the pair argument and keypoint. These two lines retrieve the pairs in the batch
arg_id = [self.labels_df['arg_id'][k] for k in indexes]
kp_id = [self.labels_df['key_point_id'][k] for k in indexes]
''' Warning: it produces nested list! '''
# Take the arguments from the arguments dataframe that match the given argument, already paired before.
arguments = flatten([self.arguments_df.loc[self.arguments_df['arg_id'] == arg]['argument'].tolist() for arg in arg_id])
# Preprocces add [CLS] as 0 and [SEP] as 2 and pads with 1s.
preprocessed_argument = self.preprocessor(arguments)
# Encode the arguments and picks the last four states of the [CLS].
encoded_argument = np.array(self.pretrained_model(preprocessed_argument)['encoder_outputs'])[:,:,0,:]
# Swap axes to have the batch size at the beginning.
encoded_argument = np.swapaxes(encoded_argument, 0, 1)
# Take the keypoint from the keypoints dataframe that match the given argument, already paired before.
keypoints = flatten([self.key_points_df.loc[self.key_points_df['key_point_id'] == kp]['key_point'].tolist() for kp in kp_id])
# Preprocces add [CLS] as 0 and [SEP] as 2 and pads with 1s.
preprocessed_keypoint = self.preprocessor(keypoints)
# Encode the keypoints and picks the last four states of the [CLS].
encoded_keypoint = np.array(self.pretrained_model(preprocessed_keypoint)['encoder_outputs'])[:,:,0,:]
# Swap axes to have the batch size at the beginning.
encoded_keypoint = np.swapaxes(encoded_keypoint, 0, 1)
labels = np.array([self.labels[k] for k in indexes])
features = [encoded_argument, encoded_keypoint]
return features, labels
def __next__(self):
if self.n >= self.__len__():
self.n = 0
result = self.__getitem__(self.n)
self.n += 1
return result
# + id="atCE_u6OhU-D"
# Batch size.
batch_size = 64
# Instantiate generators.
train_gen = DataGeneratorReduced(shuffle = True, batch_size = batch_size,
arguments_df = arguments_train, key_points_df = key_points_train, labels_df = labels_train,
preprocessor = bert_preprocess_model, pretrained_model = bert_model)
dev_gen = DataGeneratorReduced(shuffle = True, batch_size = batch_size,
arguments_df = arguments_dev, key_points_df = key_points_dev, labels_df = labels_dev,
preprocessor = bert_preprocess_model, pretrained_model = bert_model)
# No need to shuffle test data.
test_gen = DataGeneratorReduced(shuffle = False, batch_size = batch_size,
arguments_df = arguments_test, key_points_df = key_points_test, labels_df = labels_test,
preprocessor = bert_preprocess_model, pretrained_model = bert_model)
# + [markdown] id="34bz-GGzmJ19"
# #### 5.3.2 **Training**
# + id="OGdDQueFmNmv"
checkpoint = [EarlyStopping(monitor = 'val_loss', patience = 10, verbose = 1, mode = 'auto'),
ModelCheckpoint(filepath = "/content/drive/MyDrive/[NLP]/project/models/adamax_cd_hu32_bs64_small.h5",
monitor = 'val_loss', verbose = 1, save_best_only = True)]
# + id="039e1Gq7mS4P"
# Train the model.
from tensorflow.keras.models import load_model
reloaded_model_two = load_model(filepath = '/content/drive/MyDrive/[NLP]/project/models/adamax_cd_hu32_bs64_small.h5',
custom_objects = {'precision_m': precision_m, 'recall_m': recall_m, 'f1_m': f1_m})
reloaded_model_two.summary()
reloaded_model_two.fit(x = train_gen,
validation_data = dev_gen,
epochs = 100, callbacks = checkpoint)
# + [markdown] id="ldvgf1y0nkkD"
# #### 5.3.3 **Classifier evaluation**
# + colab={"base_uri": "https://localhost:8080/"} outputId="84483f13-3ba6-4380-dcb2-68e55b194eda" id="Gt0WLoYHnpjk"
from tensorflow.keras.models import load_model
reloaded_model_two = load_model(filepath = '/content/drive/MyDrive/[NLP]/project/models/adamax_cd_hu32_bs64_small.h5',
custom_objects = {'precision_m': precision_m, 'recall_m': recall_m, 'f1_m': f1_m})
reloaded_model_two.summary()
# + id="JioDZrYD0oel" colab={"base_uri": "https://localhost:8080/", "height": 761} outputId="03c2d233-c370-4bc8-9036-97387844ea57"
from sklearn.metrics import accuracy_score, classification_report, f1_score, recall_score, precision_score, PrecisionRecallDisplay, roc_curve, balanced_accuracy_score, ConfusionMatrixDisplay
def evaluate_classifier(model: tf.keras.Model, test_set_generator, true_labels):
predictions = model.predict(test_set_generator)
# Threshold tuning.
fpr, tpr, thresholds = roc_curve(true_labels, predictions)
thr = thresholds[np.argmin(np.abs(fpr + tpr - 1))]
predictions_thr = np.zeros(predictions.shape)
predictions_thr[predictions >= thr] = 1
print(classification_report(predictions_thr,
true_labels,
#sample_weight = weights
))
print(f'The average precision score is: {average_precision_score(true_labels, predictions_thr)}.')
print(f'The balanced accuracy score is: {balanced_accuracy_score(true_labels, predictions_thr)}.')
print(f'The tuned threshold is: {thr}.')
print(f'The confusion matrix is: ')
ConfusionMatrixDisplay.from_predictions(true_values, predictions_thr)
PrecisionRecallDisplay.from_predictions(true_labels, predictions)
evaluate_classifier(reloaded_model_two, test_gen, labels_test['label'].to_numpy())
# + id="A8107Ph65V0U"
| notebook_key-point_matching.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Chapter 3 Pandas
# - High level data manipulation tool
# - Built on Numpy
# - DataFrame
# ### DataFrame from Dictionary
dict = {
"country":["Brazil", "Russia", "India", "China", "South Africa"],
"capital":["Brasilia", "Moscow", "New Delhi", "Beijing", "Pretoria"],
"area":[8.516, 17.10, 3.286, 9.597, 1.221],
"population":[200.4, 143.5, 1252, 1357, 52.98]
}
import pandas as pd
brics = pd.DataFrame(dict)
brics
brics.index = ["BR", "RU", "IN", "CH", "SA"]
brics
# ### DataFrame from CSV file
brics = pd.read_csv("brics.csv")
brics
brics = pd.read_csv("brics.csv", index_col = 0)
brics
# +
# Pre-defined lists
names = ['United States', 'Australia', 'Japan', 'India', 'Russia', 'Morocco', 'Egypt']
dr = [True, False, False, False, True, True, True]
cpc = [809, 731, 588, 18, 200, 70, 45]
# Import pandas as pd
import pandas as pd
# Create dictionary my_dict with three key:value pairs: my_dict
my_dict = {"country": names,
"drives_right": dr,
"cars_per_cap":cpc
}
# Build a DataFrame cars from my_dict: cars
cars = pd.DataFrame(my_dict)
# Print cars
print(cars)
# +
# Definition of row_labels
row_labels = ['US', 'AUS', 'JPN', 'IN', 'RU', 'MOR', 'EG']
# Specify row labels of cars
cars.index = row_labels
# Print cars again
print(cars)
# -
# ### Index and Select data
# - Square brackets
# - Advanced methods
# - loc(label-based)
# - iloc(integer position-based)
# #### Column Access []
brics["country"]
type(brics["country"])
brics[["country","capital"]]
# #### Row Access []
brics[1:4]
# ### loc
brics.loc["RU"]
brics.loc[["RU"]]
brics.loc[["RU","IN","CH"]]
brics.loc[["RU","IN","CH"], ["country", "capital"]]
brics.loc[:, ["country", "capital"]]
# ### iloc
brics.iloc[[1]]
brics.iloc[[1, 2, 3]]
brics.iloc[[1, 2, 3], [0, 1]]
brics.loc["RU", "country"]
| Notes-python(DS)/Pandas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from datascience import *
import numpy as np
# %matplotlib inline
import matplotlib.pyplot as plots
plots.style.use('fivethirtyeight')
# -
# ## Lecture 6 and 7 ##
# ## Census ##
full = Table.read_table('nc-est2014-agesex-res.csv')
full
# Keep only the columns we care about
partial = full.select('SEX', 'AGE', 'POPESTIMATE2010', 'POPESTIMATE2014')
partial
# Make things easier to read
simple = partial.relabeled(2, '2010').relabeled(3, '2014')
simple
# Sort by age
simple.sort('AGE')
# Sort by age (another way)
simple.sort('AGE', descending=True)
# ## Line Plots ##
# Remove the age totals
no_999 = simple.where('AGE', are.below(999))
# Remove male and female (keep only combined)
everyone = no_999.where('SEX', 0).drop('SEX')
everyone
everyone.plot('AGE', '2010')
# +
# ^^ That plot should be labeled! Here are 3 ways to label it:
# +
# US Population <--- Just add a comment
everyone.plot('AGE', '2010')
# -
everyone.plot('AGE', '2010')
print('US Population') # <--- Print out what it is
everyone.plot('AGE', '2010')
plots.title('US Population'); # <--- OPTIONAL; not needed for Data 8
# Age distribution for two different years
everyone.plot('AGE')
# ## Males and Females in 2014 ##
# Let's compare male and female counts per age
males = no_999.where('SEX', 1).drop('SEX')
females = no_999.where('SEX', 2).drop('SEX')
pop_2014 = Table().with_columns(
'Age', males.column('AGE'),
'Males', males.column('2014'),
'Females', females.column('2014')
)
pop_2014
pop_2014.plot('Age')
# Calculate the percent female for each age
total = pop_2014.column('Males') + pop_2014.column('Females')
pct_female = pop_2014.column('Females') / total * 100
pct_female
# Round it to 3 so that it's easier to read
pct_female = np.round(pct_female, 3)
pct_female
# Add female percent to our table
pop_2014 = pop_2014.with_column('Percent female', pct_female)
pop_2014
pop_2014.plot('Age', 'Percent female')
# ^^ Look at the y-axis! Trend is not as dramatic as you might think
pop_2014.plot('Age', 'Percent female')
plots.ylim(0, 100); # Optional for Data 8
# ## Scatter Plots ##
# Actors and their highest grossing movies
actors = Table.read_table('actors.csv')
actors
actors.scatter('Number of Movies', 'Total Gross')
actors.scatter('Number of Movies', 'Average per Movie')
actors.where('Average per Movie', are.above(400))
# ## Bar Charts ##
# Highest grossing movies as of 2017
top_movies = Table.read_table('top_movies_2017.csv')
top_movies
top10_adjusted = top_movies.take(np.arange(10))
top10_adjusted
# Convert to millions of dollars for readability
millions = np.round(top10_adjusted.column('Gross (Adjusted)') / 1000000, 3)
top10_adjusted = top10_adjusted.with_column('Millions', millions)
top10_adjusted
# A line plot doesn't make sense here: don't do this!
top10_adjusted.plot('Year', 'Millions')
top10_adjusted.barh('Title', 'Millions')
| lec/lec06and07/lec06and07.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + code_folding=[0] id="d70582b9"
# Basic import
import warnings
warnings.filterwarnings('ignore')
from country_code_file import country_codes
import pandas as pd
import numpy as np
import plotly.express as px
import plotly.graph_objects as go
# %matplotlib inline
# + [markdown] id="1d29fabe"
# # Overview
# > The goal of this project is to provide a detailed EDA representation of the total number of users per attributes specified. With this project I will break this into sections to ananlyse as much of the dataset.
#
# - Workflow:
# - `Basic Discovery`
# - `Filtering`
# - `Categorical features` EDA
# - `Numerical features` EDA
#
# - toc: true
# - badges: true
# - comments: true
# - categories: [EDA, Pandas]
# - image: https://duckduckgo.com/?q=data+science+image&atb=v314-1&iax=images&ia=images&iai=https%3A%2F%2Fnews.southernct.edu%2Fwp-content%2Fuploads%2F2019%2F12%2FDataScience_home.jpg
# + colab={"base_uri": "https://localhost:8080/", "height": 357} id="c2050d97" outputId="b163181c-cfe6-4c20-9780-d4365de978b7"
df = pd.read_csv('./Users_Descriptives.csv')
df.columns = df.columns.str.replace("[^\w\s]", "").str.replace(" ", "_").str.lower()
df.head()
# + [markdown] heading_collapsed=true id="4a83c5a0"
# ## Basic Discovery
# - In this part of the notebook will just go through getting to know what data we are dealing with before doing in sort of cleaning and plotting.
# + hidden=true colab={"base_uri": "https://localhost:8080/"} id="b4af4949" outputId="8f1d363b-062c-48e9-cd4a-83a8848991ad"
df.shape
# + [markdown] hidden=true id="4905c5c8"
# **Conclusion**
# - We have `3,747,583` rows of data. Since this has a huge quantity let's check out how much `memory` this data is using.
# - We see that the machine uses `1,24GB` of memory.
# + code_folding=[0] hidden=true colab={"base_uri": "https://localhost:8080/"} id="4ea5abff" outputId="9a2de144-e62e-4ffd-b892-ac9db68f6b6c"
(df
.memory_usage(deep=True)
.sum()
)
# + hidden=true colab={"base_uri": "https://localhost:8080/", "height": 426} id="c0b68050" outputId="b5c8e828-e2e4-4679-87c6-fa8878687713"
df.describe().transpose()
# + hidden=true colab={"base_uri": "https://localhost:8080/"} id="69bb96cc" outputId="d6510957-41ef-4151-ebec-592d0ce13a7e"
df.dtypes
# + code_folding=[1, 21] hidden=true id="8f8f7b92"
# Function
def missing_data(data):
"""
This function calculates missing data for each column in a pandas dataframe.
It returns a pandas dataframe containing the column name, the number of missing values,
the percentage of missing values, the data type of the column, and the set of unique values
for that column.
:param data: A pandas dataframe
:return: A pandas dataframe
"""
total = data.isnull().sum()
percent = round(data.isnull().sum()/data.isnull().count()*100,2)
tt = pd.concat([total, percent], axis=1, keys=['Total', 'Percent'])
types = []
for col in data.columns:
dtype = str(data[col].dtype)
types.append(dtype)
tt['Types'] = types
return(np.transpose(tt))
def unique_values(data):
"""
This function takes a pandas dataframe as input and returns a transposed
pandas dataframe that lists the total number of observations and the number
of unique values for each column.
Parameters:
-----------
data: A pandas dataframe.
Returns:
--------
A transposed pandas dataframe that lists the total number of observations
and the number of unique values for each column.
"""
total = data.count()
tt = pd.DataFrame(total)
tt.columns = ['Total']
uniques = []
for col in data.columns:
unique = data[col].nunique()
uniques.append(unique)
tt['Uniques'] = uniques
return(np.transpose(tt))
# + hidden=true colab={"base_uri": "https://localhost:8080/", "height": 143} id="67a5d704" outputId="7381193e-a879-4144-d3ca-b0a58f46d780"
obj = df.select_dtypes(include="object")
missing_data(obj)
# + hidden=true colab={"base_uri": "https://localhost:8080/", "height": 426} id="b76046aa" outputId="d17cb4a5-0f12-43b2-b506-b926d03aa981"
num = df.select_dtypes(include="number")
nm = missing_data(num)
nm.transpose()
# + [markdown] hidden=true id="0dd52397"
# **Conclusion**
# - The `city` column contains has the highest number of missing values with 31%
# - But since all the categorical columns have low percentages we can just drop them
# + hidden=true colab={"base_uri": "https://localhost:8080/", "height": 557} id="11abbc47" outputId="beff679c-a309-4166-fff2-18c8cd4a6751"
nan_df = df[df.isna().any(axis=1)]
nan_df
# + code_folding=[0] hidden=true id="99eda980"
def clean(data):
# Categorical
cols = ['city', 'state', 'country', 'gender']
for objs in cols:
data = data.dropna(subset=cols)
# Numerical
numerical = data.select_dtypes(include="number")
for num in numerical.columns:
data[num].fillna(data[num].median(), inplace=True)
# data[num].fillna(0, inplace=True)
data["city"] = data['city'].astype('category')
data["state"] = data['state'].astype('category')
data["country"] = data['country'].astype('category')
data["gender"] = data['gender'].astype('category')
return data
# + hidden=true colab={"base_uri": "https://localhost:8080/", "height": 551} id="34f099f5" outputId="cf2a5b1d-ba13-4746-ad70-2cc0f2716d51"
user_d = clean(df)
md = missing_data(user_d)
md.transpose()
# + [markdown] heading_collapsed=true id="ed48c44f"
# ## Filtering
# **QUESTIONS**
# - What is the average and sum of variables by gender.
# - What is the average and sum of variables by country.
# - Which gender per country as the most views, fame, questions
# + code_folding=[0] hidden=true colab={"base_uri": "https://localhost:8080/", "height": 175} id="802ef08d" outputId="aaddef66-4370-4cac-a03c-d78a588e6b83"
# Average and sum of variables by gender.
(user_d
.groupby("gender")
.agg({"questions": ["mean", "sum"],
"answers": ["mean", "sum"],
"comments": ["mean", "sum"],
"activitymonthlyduration": ["mean", "sum"]
})
)
# + [markdown] hidden=true id="6719450c"
# **Conclusion**
# - Clearly males are the ones with the highest average and sum
# - Let's see which country reflects this
# + hidden=true colab={"base_uri": "https://localhost:8080/", "height": 363} id="52390de1" outputId="6f5297e6-32fc-431b-e483-3ad2ed25127d"
# Average and sum of variables by country.
user_d = user_d.replace({'United States':'US', 'United Kingdom':'UK', 'South Africa':'SA'})
user_country_code = user_d[user_d["country"].isin(["US", "UK", "India", "Germany", "SA",
"Canada", "Australia", "France"])]
(user_country_code
.groupby("country")
.agg({"questions": ["mean", "sum"],
"answers": ["mean", "sum"],
"views": ["mean", "sum"],
"fame": ["mean", "sum"]
})
)
# + [markdown] hidden=true id="a43d95f4"
# **Conclusion**
# - The `UK` reflects the highest average and sum
# - `Germany` follows in second
# + code_folding=[1, 3] hidden=true colab={"base_uri": "https://localhost:8080/", "height": 143} id="ac1cc02b" outputId="6d178cb1-55d5-4af1-b804-0224e8606415"
# Gender per country with views
user_group = user_d[user_d["country"].isin(["US", "UK", "India", "Germany", "SA",
"Canada", "Australia", "France"])]
(user_group
.groupby(['gender','country'])
.views
.first()
.unstack()
)
# + code_folding=[0] hidden=true colab={"base_uri": "https://localhost:8080/", "height": 143} id="d0b8c529" outputId="f6b5785f-dc30-4e59-ab29-258d738ff5ea"
# gender per country with fame
user_d = user_d.replace({'United States':'US', 'United Kingdom':'UK', 'South Africa':'SA'})
user_group = user_d[user_d["country"].isin(["US", "UK", "India", "Germany", "SA",
"Canada", "Australia", "France"])]
(user_group
.groupby(['gender','country'])
.fame
.first()
.unstack()
)
# + hidden=true colab={"base_uri": "https://localhost:8080/", "height": 143} id="93967595" outputId="0d3680b5-0eda-43ab-be96-30523ff23bf7"
# gender per country with questions
user_d = user_d.replace({'United States':'US', 'United Kingdom':'UK', 'South Africa':'SA'})
user_group = user_d[user_d["country"].isin(["US", "UK", "India", "Germany", "SA",
"Canada", "Australia", "France"])]
(user_group
.groupby(['gender','country'])
.questions
.first()
.unstack()
)
# + [markdown] heading_collapsed=true id="8dd5d20b"
# ## EDA Analysis
# + [markdown] heading_collapsed=true hidden=true id="d630f6e6"
# ### Categorical
# **QUESTIONS**
# - What is the top countries with high fame by gender?
# - What is the top cities with high questions?
# - What is the total fame by country?
# - `World Map`: Total questions, answers, edits, monthly duration by country
# + code_folding=[0, 5] hidden=true colab={"base_uri": "https://localhost:8080/", "height": 542} id="d2b2ac3d" outputId="2b117af1-e1d3-4b02-f514-08336bca5471"
pie_gender_views = (user_d
.groupby('gender', as_index=False)
.sum()
.sort_values(by='views', ascending=False)
)
def pie_chart(data, col, val):
fig = px.pie(data, values=val, names=col,
color_discrete_sequence=['#14213d', '#9d0208'],
title=f'Total {val} by {col}',
hole=.6)
fig.update_traces(textposition='inside', textinfo='percent+label')
fig.layout.template = 'plotly_dark'
return fig.show()
pie_chart(pie_gender_views, 'gender', 'views')
# + code_folding=[0, 2] hidden=true colab={"base_uri": "https://localhost:8080/", "height": 1000} id="9b3e56be" outputId="f117a1da-7c00-424b-cda5-bf63852ab010"
# Histogram Gender
gender_hist = user_d.replace('male', 'Males').replace('female', 'Females')
for gen in gender_hist['gender'].unique():
dataframe = gender_hist.loc[gender_hist['gender'] == gen].groupby('country')['fame'].agg('sum').sort_values(ascending=False)[:10]
dataframe = pd.DataFrame({'Country':dataframe.index, 'Fame':dataframe.values})
fig = px.histogram(dataframe, x="Fame", y='Country', color='Country',
title=f'Top 10 countries with fame by {gen}')
fig.layout.template = 'plotly_dark'
fig.update_traces(opacity=0.7)
fig.show()
# + [markdown] hidden=true id="e60a954e"
# **Coclusion**
# - For `males`: US as expected came first but interestingly we have `UK second`. We can get a closer look at which city in the UK carry most of fame.
# - For `females`: US again first, but `India came in second` we is the opposite of the males. We will also look at which city in India carry the most fame.
# + code_folding=[1] hidden=true colab={"base_uri": "https://localhost:8080/", "height": 269} id="8ab3c7d4" outputId="56f7731b-2819-4367-c98e-f06a8b080b4b"
india_female_city = user_d[(user_d['gender']=='female') & (user_d['country']=='India')]
(india_female_city['city']
.value_counts()
.reset_index()
.rename(columns={'index':'city', 'city':'count'})[:7]
)
# + code_folding=[1] hidden=true colab={"base_uri": "https://localhost:8080/", "height": 269} id="8a233f90" outputId="09f81b84-fa61-4f92-8777-19356592224d"
uk_male_city = user_d[(user_d['gender']=='male') & (user_d['country']=='UK')]
(uk_male_city['city']
.value_counts()
.reset_index()
.rename(columns={'index':'city', 'city':'count'})[:7]
)
# + code_folding=[0, 2] hidden=true colab={"base_uri": "https://localhost:8080/", "height": 542} id="987fa516" outputId="d7817bde-cc3a-4d72-8275-089348f5dfe0"
# Histogram Countries
dataframe = user_d.groupby('state').agg('sum').sort_values(by='views', ascending=False)[:10]
fig = px.histogram(dataframe, x="views", y=dataframe.index, color=dataframe.index,
title=f'Top 10 states with high views')
fig.layout.template = 'plotly_dark'
fig.update_traces(opacity=0.7)
fig.show()
# + code_folding=[0, 3] hidden=true colab={"base_uri": "https://localhost:8080/", "height": 542} id="a9b13a00" outputId="9e292ae1-b831-4947-837d-b1d0029a7a64"
# Histogram Cities
unq_variant = user_d[user_d["country"].isin(["US", "UK", "India", "Germany"])]
dataframe = unq_variant.groupby('city').agg('sum').sort_values(by='questions', ascending=False)[:10]
fig = px.histogram(dataframe, x="questions", y=dataframe.index, color=dataframe.index,
title=f'Top 10 cities with high questions',
labels={'sum of questions':'questions'})
fig.layout.template = 'plotly_dark'
fig.update_traces(opacity=0.7)
fig.show()
# + hidden=true id="ccf60938"
def scatter_world_map(data, category, size):
user_category = data.groupby(category, as_index=False).sum()
user_category['Code'] = user_category[category].map(country_codes)
# Plot
map_fig = px.scatter_geo(user_category,
locations = 'Code',
projection = 'orthographic',
title=f'Total {size} by country',
color = category,
opacity=.7,
size=size,
hover_name=category,
hover_data=['views', 'questions', 'activitymonthlyduration', 'fame']
)
map_fig.layout.template = 'plotly_dark'
return map_fig.show()
# + hidden=true colab={"base_uri": "https://localhost:8080/", "height": 542} id="92d7e6a4" outputId="fede292e-c8b2-4a4e-d856-28a705dbc232"
scatter_world_map(user_d, 'country', 'views')
# + hidden=true colab={"base_uri": "https://localhost:8080/", "height": 542} id="8e9ce27b" outputId="8c3ba912-e7b8-418a-bc6a-c94f95b9a485"
scatter_world_map(user_d, 'country', 'questions')
# + hidden=true colab={"base_uri": "https://localhost:8080/", "height": 542} id="0ec8c7e4" outputId="7c8e5f0c-4544-4f38-e24d-6cd259087cf5"
scatter_world_map(user_d, 'country', 'fame')
# + [markdown] heading_collapsed=true hidden=true id="0a70b632"
# ### Numerical
# **QUESTIONS**
# - Which city and gender have the highest views?
# - Which country has the lowest fame?
# + code_folding=[0, 2] hidden=true colab={"base_uri": "https://localhost:8080/", "height": 542} id="bfb8bd97" outputId="cc8f167a-b79e-44a4-c4ac-96403da14e15"
def correlation(data):
corrmat = data.corr()
fig = px.imshow(corrmat, color_continuous_scale=["#caf0f8","#03071e", "#370617", "#70e000"],
title='Heatmap User Description')
fig.layout.template = 'plotly_dark'
return fig.show()
correlation(user_d)
# + [markdown] heading_collapsed=true hidden=true id="e2bcd4ae"
# #### Views
# + code_folding=[0] hidden=true id="78377633"
def numerical_histogram(data, x, color):
fig = px.histogram(data, x=x,
title=f'Histogram of {x}',
color_discrete_sequence=[color])
fig.layout.template = 'plotly_dark'
return fig.show()
# + hidden=true colab={"base_uri": "https://localhost:8080/", "height": 0} id="320c9dd8" outputId="08828c40-ee24-4b7d-fd3f-18607c90a4e7"
view_zero = user_d[user_d['views']<150]
numerical_histogram(view_zero, 'views', 'rosybrown')
# + [markdown] hidden=true id="47b3e4c3"
# **Conclusion**
# - Views is right skewed
# - There is a lot of data and `outliers in views`, which makes our data `skew`. So we will lower the amount of views in our data because it is clear that less people have more views.
# + code_folding=[1] hidden=true colab={"base_uri": "https://localhost:8080/", "height": 0} id="17cfd7e3" outputId="9ea014a3-bdb2-40ac-8566-d5c3fbd9c549"
one_million = user_d[user_d['views']>100000]
(one_million['state']
.value_counts()
.reset_index()[:10]
.rename(columns={'index':'state', 'state':'count'})
)
# + hidden=true colab={"base_uri": "https://localhost:8080/", "height": 0} id="bb5387b2" outputId="a70bbfed-3ba3-4dd6-de9f-ab84f4744c54"
user_d[user_d['views']>500000]
# + hidden=true colab={"base_uri": "https://localhost:8080/", "height": 0} id="5e816d92" outputId="2e0ec615-b03f-4960-a0d9-7765b0be26e0"
user_d[user_d['views']<5][:10]
# + code_folding=[0] hidden=true id="340face1"
def categorical_feature(x):
if x < 35:
return 'Low Views'
elif x < 70:
return 'Some Views'
elif x < 105:
return 'Medium Views'
elif pd.isnull(x):
return np.nan
else:
return 'High Views'
# + hidden=true colab={"base_uri": "https://localhost:8080/", "height": 0} id="139398ce" outputId="64255ab1-3196-4c4e-e320-70dda8a2036a"
view_zero['view_feature'] = view_zero['views'].map(categorical_feature)
view_zero[['country','city','gender','views', 'view_feature']]
# + code_folding=[0, 5] hidden=true colab={"base_uri": "https://localhost:8080/", "height": 0} id="d0132643" outputId="97ded0c9-29f6-474a-c1bf-c90084ac2ab4"
hist_categorical_views = (view_zero['view_feature']
.value_counts()
.reset_index()
.rename(columns=({'index': 'views', 'view_feature': 'count'}))
)
fig = px.histogram(hist_categorical_views, x='count', y='views',
title='Histogram of View Feature',
color='views',
color_discrete_sequence=['#005f73', '#ee9b00', '#ae2012', '#9b2226'])
fig.layout.template = 'plotly_dark'
fig.show()
# + hidden=true colab={"base_uri": "https://localhost:8080/", "height": 0} id="e9b9d079" outputId="21b5435d-fb99-43fc-a675-8ba91d1ff944"
view_zero[view_zero['view_feature']=='High Views'].sort_values(by='views', ascending=False)[:10]
# + hidden=true colab={"base_uri": "https://localhost:8080/", "height": 0} id="0093c3de" outputId="330c1242-584f-462e-ced2-b9b0865f9f37"
view_zero['country'].value_counts().reset_index()[:10]
# + [markdown] heading_collapsed=true hidden=true id="504feb03"
# #### monthly duration
# + hidden=true colab={"base_uri": "https://localhost:8080/"} id="9d5d5c5b" outputId="5d25eaec-d33d-4f01-dc35-d9e3257a5b20"
user_d['activitymonthlyduration'].value_counts()
# + hidden=true id="d4c0fce1"
amd = user_d[(user_d['activitymonthlyduration']>0.05) & (user_d['activitymonthlyduration']<12.00)]
# + [markdown] hidden=true id="da636255"
# **Conclusion**
# - There is a lot of people with less than a month so we can remove some just to see our data better
# + code_folding=[1] hidden=true colab={"base_uri": "https://localhost:8080/", "height": 0} id="295abe92" outputId="eb62e89b-6248-4b8e-a001-0daa2bec70b9"
(amd['activitymonthlyduration']
.value_counts()
.reset_index()[:10]
.rename(columns={'index':'monthly_duration', 'activitymonthlyduration':'count'})
)
# + hidden=true colab={"base_uri": "https://localhost:8080/", "height": 0} id="a320bde6" outputId="2294961f-919e-4061-b2f6-4298763fa929"
numerical_histogram(amd, 'activitymonthlyduration', 'saddlebrown')
# + code_folding=[0] hidden=true id="5425345c"
def categorical_feature(x):
if x < 1:
return 0
elif x < 2:
return 1
elif x < 3:
return 2
elif x < 4:
return 3
elif x < 5:
return 4
elif x < 6:
return 5
elif x < 7:
return 6
elif x < 8:
return 7
elif x < 9:
return 8
elif x < 10:
return 9
elif x < 11:
return 10
elif x < 12:
return 11
else:
return 12
amd['monthly_feature'] = amd['activitymonthlyduration'].map(categorical_feature)
# + hidden=true colab={"base_uri": "https://localhost:8080/", "height": 0} id="028eb198" outputId="888ab641-6120-46ce-f72c-4d3957fb4afa"
amd[['country','city','gender','activitymonthlyduration', 'monthly_feature']]
# + code_folding=[0, 5] hidden=true colab={"base_uri": "https://localhost:8080/", "height": 0} id="2a3a34f7" outputId="2b09995c-080f-4ecf-fb94-913f01b5cb96"
hist_categorical_monthly = (amd['monthly_feature']
.value_counts()
.reset_index()
.rename(columns=({'index': 'monthly', 'monthly_feature': 'count'}))
)
fig = px.histogram(hist_categorical_monthly, x='monthly', y='count',
title='Histogram of View Feature',
color_discrete_sequence=['saddlebrown'],
nbins=35
)
fig.layout.template = 'plotly_dark'
fig.show()
# + [markdown] heading_collapsed=true hidden=true id="21d71238"
# #### fame
# + hidden=true id="24c0af09"
fame_data = user_d[user_d['fame']>1]
# + code_folding=[0] hidden=true colab={"base_uri": "https://localhost:8080/", "height": 0} id="4f3a9458" outputId="14eb5ef5-6cfb-4489-a303-a9bd794e9542"
(user_d['fame']
.value_counts()
.reset_index()[:10]
.rename(columns={'index':'fame', 'fame':'count'})
)
# + hidden=true id="aa27bc40"
user_fame = user_d[(user_d['fame']>1)& (user_d['fame']<250)]
# + hidden=true colab={"base_uri": "https://localhost:8080/", "height": 0} id="823c6933" outputId="bf98fe03-18e8-4983-bd03-8353dc971b2d"
numerical_histogram(user_fame, 'fame', 'sienna')
# + code_folding=[0] hidden=true id="aa36e272"
def categorical_feature(x):
if x <= 60:
return 'Beginner'
elif x <=120:
return 'Medium'
elif x <= 180:
return 'Intermediate'
else:
return 'Famous'
# + hidden=true colab={"base_uri": "https://localhost:8080/", "height": 0} id="63e3905f" outputId="e10d8dda-ac65-437f-fade-d6f972eeeb26"
user_fame['fame_feature'] = user_fame['fame'].map(categorical_feature)
user_fame[['country','city','gender','fame', 'fame_feature']]
# + hidden=true colab={"base_uri": "https://localhost:8080/", "height": 0} id="a0666531" outputId="28a2b6f5-6030-41f4-a1c5-a2ecc3178485"
user_fame[user_fame['fame_feature']=='Famous']
# + code_folding=[0, 5] hidden=true colab={"base_uri": "https://localhost:8080/", "height": 0} id="7404f6f0" outputId="7d528ed6-dea3-4ae0-875e-51440cd61bf2"
hist_categorical_fame = (user_fame['fame_feature']
.value_counts()
.reset_index()
.rename(columns=({'index': 'fame', 'fame_feature': 'count'}))
)
fig = px.histogram(hist_categorical_fame, x='count', y='fame',
title='Histogram of Fame Feature',
color='fame',
color_discrete_sequence=['#007f5f', '#2b9348', '#55a630', '#80b918'])
fig.layout.template = 'plotly_dark'
fig.show()
# + hidden=true id="f0c9ff6d"
| _notebooks/2022_04_12_user_description.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="ixCsLMj-CAF7"
# The module below uses only the first feature of the diabetes dataset, in order to illustrate the data points within the two-dimensional plot. The straight line can be seen in the plot, showing how linear regression attempts to draw a straight line that will best minimize the residual sum of squares between the observed responses in the dataset, and the responses predicted by the linear approximation.
#
# The coefficients, residual sum of squares and the coefficient of determination are also calculated.
# + id="ANp6MJCjB7m2"
# Code source: <NAME>
# License: BSD 3 clause
import matplotlib.pyplot as plt
import numpy as np
from sklearn import datasets, linear_model
from sklearn.metrics import mean_squared_error, r2_score
# + id="IsbR7zZNCMCN"
# Load the diabetes dataset
diabetes_X, diabetes_y = datasets.load_diabetes(return_X_y=True)
# + id="BBoFQFd0CQJZ"
# Use only one feature
diabetes_X = diabetes_X[:, np.newaxis, 2]
# + id="5704_6eaCRt4"
# Split the data into training/testing sets
diabetes_X_train = diabetes_X[:-20]
diabetes_X_test = diabetes_X[-20:]
# + id="Np2tUIL3CTpS"
# Split the targets into training/testing sets
diabetes_y_train = diabetes_y[:-20]
diabetes_y_test = diabetes_y[-20:]
# + id="BN2ue318CVAL"
# Create linear regression object
regr = linear_model.LinearRegression()
# + colab={"base_uri": "https://localhost:8080/"} id="_KW5kOHxCWQv" executionInfo={"status": "ok", "timestamp": 1636399940225, "user_tz": -480, "elapsed": 859, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gict81qvzCzYor7OD2HAqOMscBAGuRrSJShYazDZg=s64", "userId": "02100760507760735849"}} outputId="48fb55d4-db59-4d24-9bd3-2777999effc3"
# Train the model using the training sets
regr.fit(diabetes_X_train, diabetes_y_train)
# + id="jIFLkuKRCXyy"
# Make predictions using the testing set
diabetes_y_pred = regr.predict(diabetes_X_test)
# + colab={"base_uri": "https://localhost:8080/"} id="_RoW1WP4CaJI" executionInfo={"status": "ok", "timestamp": 1636399961756, "user_tz": -480, "elapsed": 3, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gict81qvzCzYor7OD2HAqOMscBAGuRrSJShYazDZg=s64", "userId": "02100760507760735849"}} outputId="7c27fb0a-6984-451d-89a9-cfdb552a5896"
# The coefficients
print("Coefficients: \n", regr.coef_)
# The mean squared error
print("Mean squared error: %.2f" % mean_squared_error(diabetes_y_test, diabetes_y_pred))
# The coefficient of determination: 1 is perfect prediction
print("Coefficient of determination: %.2f" % r2_score(diabetes_y_test, diabetes_y_pred))
# + colab={"base_uri": "https://localhost:8080/", "height": 252} id="y8lcejLWCdHG" executionInfo={"status": "ok", "timestamp": 1636399995574, "user_tz": -480, "elapsed": 371, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gict81qvzCzYor7OD2HAqOMscBAGuRrSJShYazDZg=s64", "userId": "02100760507760735849"}} outputId="17dbd9a0-e87e-4127-9ade-a0bcb74ff624"
# Plot outputs
plt.scatter(diabetes_X_test, diabetes_y_test, color="black")
plt.plot(diabetes_X_test, diabetes_y_pred, color="blue", linewidth=3)
plt.xticks(())
plt.yticks(())
plt.show()
| Week 2/Linear Regression Example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Elliptic Curve Cryptography using the BitCoin curve, SECG secp256k1
# Dr. <NAME>, <EMAIL>, 2015-02-20 (Public Domain)
#
# full Python3 elliptic curve point multiplication code here - uaf-cs
# https://www.cs.uaf.edu/2015/spring/cs463/lecture/02_23_ECC_impl/ECC_bitcoin.py
# Convert a string with hex digits, colons, and whitespace to a long integer
def hex2int(hexString):
return int("".join(hexString.replace(":","").split()),16)
# Half the extended Euclidean algorithm:
# Computes gcd(a,b) = a*x + b*y
# Returns only gcd, x (not y)
# From http://rosettacode.org/wiki/Modular_inverse#Python
def half_extended_gcd(aa, bb):
lastrem, rem = abs(aa), abs(bb)
x, lastx = 0, 1
while rem:
lastrem, (quotient, rem) = rem, divmod(lastrem, rem)
x, lastx = lastx - quotient*x, x
return lastrem, lastx
# Modular inverse: compute the multiplicative inverse i of a mod m:
# i*a = a*i = 1 mod m
def modular_inverse(a, m):
g, x = half_extended_gcd(a, m)
if g != 1:
raise ValueError
return x % m
# An elliptic curve has these fields:
# p: the prime used to mod all coordinates
# a: linear part of curve: y^2 = x^3 + ax + b
# b: constant part of curve
# G: a curve point (G.x,G.y) used as a "generator"
# n: the order of the generator
class ECcurve:
def __init__(self):
return
# Prime field multiplication: return a*b mod p
def field_mul(self,a,b):
return (a*b)%self.p
# Prime field division: return num/den mod p
def field_div(self,num,den):
inverse_den=modular_inverse(den%self.p,self.p)
return self.field_mul(num%self.p,inverse_den)
# Prime field exponentiation: raise num to power mod p
def field_exp(self,num,power):
return pow(num%self.p,power,self.p)
# Return the special identity point
# We pick x=p, y=0
def identity(self):
return ECpoint(self,self.p,0)
# Return true if point Q lies on our curve
def touches(self,Q):
y2=self.field_exp(Q.y,2)%self.p
# print("y^2 = ", y2)
x3ab=(self.field_mul((Q.x*Q.x)%self.p+self.a,Q.x)+self.b)%self.p
# print("x^3 + a x + b = ",x3ab)
return y2==x3ab
# Return the slope of the tangent of this curve at point Q
def tangent(self,Q):
return self.field_div(Q.x*Q.x*3+self.a,Q.y*2)
# Return the (x,y) point where this line intersects our curve
# Q1 and Q2 are two points on the line of slope m
def line_intersect(self,Q1,Q2,m):
v=(Q1.y + self.p - (m*Q1.x)%self.p)%self.p
x=(m*m + self.p-Q1.x + self.p-Q2.x)%self.p
y=(self.p-(m*x)%self.p + self.p-v)%self.p
return ECpoint(self,x,y)
# Return a doubled version of this elliptic curve point
def double(self,Q):
if (Q.x==self.p): # doubling the identity
return Q
if (Q.y==0): # vertical tangent
return self.identity()
return self.line_intersect(Q,Q,self.tangent(Q))
# Return the "sum" of these elliptic curve points
def add(self,Q1,Q2):
# Identity special cases
if (Q1.x==self.p): # Q1 is identity
return Q2
if (Q2.x==self.p): # Q2 is identity
return Q1
# Equality special cases
if (Q1.x==Q2.x):
if (Q1.y==Q2.y): # adding point to itself
return self.double(Q1)
else: # vertical pair--result is the identity
return self.identity()
# Ordinary case
m=self.field_div(Q1.y+self.p-Q2.y,Q1.x+self.p-Q2.x)
return self.line_intersect(Q1,Q2,m)
# "Multiply" this elliptic curve point Q by the integer m
# Often the point Q will be the generator G
def mul(self,Q,m):
R=self.identity() # return point
while m!=0: # binary multiply loop
if m&1: # bit is set
# print(" mul: adding Q to R =",R);
R=self.add(R,Q)
m=m>>1
if (m!=0):
# print(" mul: doubling Q =",Q);
Q=self.double(Q)
return R
# A point on an elliptic curve: (x,y)
class ECpoint:
"""A point on an elliptic curve (x,y)"""
def __init__(self,curve, x,y):
self.curve=curve
self.x=x
self.y=y
if not x==curve.p and not curve.touches(self):
print(" ECpoint left curve: ",x,",",y)
# "Add" this point to another point on the same curve
def add(self,Q2):
return self.curve.add(self,Q2)
# "Multiply" this point by a scalar
def mul(self,m):
return self.curve.mul(self,m)
# Print this ECpoint
def __str__(self):
if (self.x==self.curve.p):
return "identity_point"
else:
return "("+str(self.x)+", "+str(self.y)+")"
# -
myECC_01 = ECcurve()
myECC_01.p = 23
myECC_01.a = 1
myECC_01.b = 1
myECC_01.G = ECpoint(curve=myECC_01,x=0,y=1)
myECC_01.n = 29
# +
# Test program:
curve=myECC_01
Q=curve.G
print("Generator touches curve? ",curve.touches(Q));
print("Tangent of generator: ",curve.tangent(Q));
print("Initial curve point ",Q);
for i in range(2,myECC_01.n):
Q=Q.add(curve.G) # repeatedly add generator
print("Curve point ",i,Q);
J=curve.mul(curve.G,i) # direct jump
if (J.x!=Q.x or J.y!=Q.y):
print(" -> MULTIPLY MISMATCH: ",J.x,",",J.y);
# -
print("9 Q ", curve.mul(curve.G,2))
144 % 23
(12*(0-6) - 1 ) % 23
# +
myECC_02 = ECcurve()
myECC_02.p = 5
myECC_02.a = 2
myECC_02.b = 3
myECC_02.G = ECpoint(curve=myECC_02,x=1,y=1)
myECC_02.n = 7
curve=myECC_02
Q=curve.G
print("Generator touches curve? ",curve.touches(Q));
# print("Tangent of generator: ",curve.tangent(Q));
# print("Initial curve point ",Q);
print("6*(1,1) ", curve.mul(curve.G, 6))
for i in range(2,myECC_02.n):
J=curve.mul(curve.G,i) # direct jump
print("Curve point direct jump ",J.x,",",J.y);
for i in range(2,myECC_02.n):
Q=Q.add(curve.G) # repeatedly add generator
print("Curve point ",i,Q);
J=curve.mul(curve.G,i) # direct jump
if (J.x!=Q.x or J.y!=Q.y):
print(" -> MULTIPLY MISMATCH: ",J.x,",",J.y);
# +
myECC_03 = ECcurve()
myECC_03.p = 5
myECC_03.a = 1
myECC_03.b = 1
myECC_03.G = ECpoint(curve=myECC_03,x=0,y=1)
myECC_03.n = 10
curve=myECC_03
Q=curve.G
print("Generator touches curve? ",curve.touches(Q));
print("Tangent of generator: ",curve.tangent(Q));
print("Initial curve point ",Q);
for i in range(2,myECC_03.n):
Q=Q.add(curve.G) # repeatedly add generator
print("Curve point ",i,Q);
J=curve.mul(curve.G,i) # direct jump
if (J.x!=Q.x or J.y!=Q.y):
print(" -> MULTIPLY MISMATCH: ",J.x,",",J.y);
# -
Q1 = ECpoint(curve=myECC_03,x=0,y=1)
print(Q1.add(Q1))
[print(Q1.mul(i)) for i in range(10)]
Q2 = ECpoint(curve=myECC_03,x=4,y=2)
Q3 = ECpoint(curve=myECC_03,x=3,y=1)
print(Q2.add(Q3))
# +
myECC_04 = ECcurve()
myECC_04.p = 23
myECC_04.a = 1
myECC_04.b = 3
myECC_04.G = ECpoint(curve=myECC_04,x=0,y=7)
myECC_04.n = 29
curve=myECC_04
Q=curve.G
print("Generator touches curve? ",curve.touches(Q));
print("Tangent of generator: ",curve.tangent(Q));
print("Initial curve point ",Q);
for i in range(2,myECC_04.n):
Q=Q.add(curve.G) # repeatedly add generator
print("Curve point ",i,Q);
J=curve.mul(curve.G,i) # direct jump
if (J.x!=Q.x or J.y!=Q.y):
print(" -> MULTIPLY MISMATCH: ",J.x,",",J.y);
# -
| SW06/Elliptic_curves_over_finite_field_01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Modelling spreading of infectious diseases
# This is an experimental model. Inspirered by https://kagr.shinyapps.io/C19DK/ but for this example without age groups
#
# The model is purely for testing the capabilities of ModelFlow, the parameters selected are for ilustration of the dynamic and are not actual estimates.
#
# This is a Jupyter Notebook running Python.
#
# The notebook is located on github here: https://github.com/IbHansen/Modelflow2
#
# Feel free to use this notebook. **To run the notebook** find the menu line above and select **cell>run all**
#
#
# THE Notebook IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR
# IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY,
# FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE
# AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER
# LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM,
# OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE
# SOFTWARE.
#
# The model is specified as equations. The equations defines the daily transition between the states:
#
# - susciptible
# - exposed
# - infectious
# - hospitalized
# - In ICU
# - recovered
# - dead
#
# # Make eksperiments
# The model will run with a baseline scenario and you can inspect the results.
#
# Then you can use the sliders to change the scenario.
# You can make your own scenario. Use the sliders to define an scenario. Then press the **Run scenario** button. The model will solve, and the results can be inspected.
#
# For instance, try to increase *Daily rate of contact* to 6 and press **Run scenario** and watch how the values changes.
#
# You can change the scenario name. After **Run scenario** a graph for each scenario will be shown.
#
# Again, you don't have to understand the Python code.
# +
import pandas as pd
from numpy import arange
from ipywidgets import interact, Dropdown, Checkbox, IntRangeSlider,SelectMultiple, Layout
from ipywidgets import interactive
from modelclass import model
import modelmf
model.modelflow_auto() # to run the model, when loaded
# -
# # Specify the model
# First a model specification **rcorona** is ceated. It specifies the dynamic of the different states.
#
# Then a model instance **mcorona** from rcorona. This is a python class object which contains the solver of the model and different relevant metohds to manipulate the model.
#
# Some conventions are used setting up the model specification:
# - (-1) after a variable means the value the day before.
# - diff means the change in variable from the day before.
rcorona = '''\
infection_rate = rate_contact * probability_transmision * infectious(-1) / population(-1)
new_exposed = infection_rate * susceptible + exo_exposed
diff(exposed) = new_exposed - new_infectious
new_infectious = new_exposed(-2)*0.8 + new_exposed(-3)*0.2
new_infectious_to_recover = new_infectious * new_infectious_to_recover_rate
exit_infectious_to_recover = new_infectious_to_recover(-14)
new_infectious_to_hospital = new_infectious * (1-new_infectious_to_recover_rate)
diff(infectious) = new_infectious - exit_infectious_to_recover-exit_intensive_to_recover -exit_hospital_to_recover -exit_intensive_to_dead
new_hospital_to_recover = new_infectious_to_hospital * new_hospital_to_recover_rate
exit_hospital_to_recover = new_hospital_to_recover(-5)
diff(hospital_to_recover) = new_hospital_to_recover - exit_hospital_to_recover
new_hospital_to_Intensive = new_infectious_to_hospital * (1-new_hospital_to_recover_rate)
new_Intensive_to_recover = new_hospital_to_Intensive * new_Intensive_to_recover_rate
exit_intensive_to_recover = new_intensive_to_recover(-7)
diff(intensive_to_recover) = new_intensive_to_recover-exit_intensive_to_recover
new_Intensive_to_dead = new_hospital_to_Intensive * (1-new_Intensive_to_recover_rate)
exit_intensive_to_dead = new_intensive_to_dead(-20)
diff(intensive_to_dead) = new_intensive_to_dead-exit_intensive_to_dead
diff(hospital_non_icu) = new_hospital_to_recover - exit_Hospital_to_recover
diff(hospital_icu) = new_Intensive_to_recover+new_Intensive_to_dead-(exit_intensive_to_recover+exit_Intensive_to_dead)
hospital = hospital_non_icu+hospital_icu
diff(dead) = exit_intensive_to_dead
diff(susceptible) = -new_exposed
diff(recovered) = exit_infectious_to_recover + exit_intensive_to_recover + exit_hospital_to_recover
diff(population) = -diff(dead)
dead_delta = diff(dead)
infectious_delta = diff(infectious)
hospital_delta = diff(hospital)
hospital_icu_delta = diff(hospital_icu)
dead_growth = 100 * dead_delta / dead(-1)
infectious_growth = 100 * infectious_delta / infectious(-1)
hospital_growth = 100 * hospital_delta / hospital(-1)
hospital_icu_growth = 100 * hospital_icu_delta / hospital_icu(-1)
'''
mcorona = model.from_eq(rcorona) # create a model instance which can solve the model
# # Specify a baseline with no infections
# A dataframe newdf is created. It contains baseline with information regarding the population, the infection, the spreading and the number of infested.
#
# In the baseline there there are no infections as
# EXO_EXPOSED is set to 0
DAYS = 500 # number of days the model will run
empty = pd.DataFrame(index=range(DAYS)) # make an empty dataframe with DAYS rows
empty.index.name = 'Day'
newdf = empty.mfcalc('''\
POPULATION = 1000000
SUSCEPTIBLE = 1000000
EXO_EXPOSED = 0
NEW_INFECTIOUS_TO_RECOVER_RATE = 0.9
NEW_HOSPITAL_TO_RECOVER_RATE = 0.01
NEW_INTENSIVE_TO_RECOVER_RATE = 0.6
RATE_CONTACT = 4.
PROBABILITY_TRANSMISION = 0.05
''' )
# ## Display the first days of the baseline
newdf.head(4).T.style.format("{:15.2f}")
# # Run the model, and show some results
# ## Update with 10 infected from the outside on day 30
baseline = model.update_from_list(newdf, f'''
EXO_EXPOSED = 10 30 31 ''') # getting 10 infected at day 30
# ## Run the model
base_result = mcorona(baseline,keep = f'baseline 10 persons infested at day 30')
# ## Dump the model and baseline
# This allows us to load the model and baseline in other notebooks
mcorona.modeldump('coronatest.pcim')
# This "precooked" model can be loaded in other notebooks like in this one [Interactive slideshow](Corona%20experiments%20with%20policy%20-%20interactive%20slideshow.ipynb) where you can look at other experiments
# # Display the results
with mcorona.set_smpl(20,300):
mcorona.keep_plot(' hospital_icu dead infectious',diff=0,legend=1,dec='0')
# # Make eksperiments with a range of initial infections
# Run and display
# +
mcorona_infection = model.from_eq(rcorona) # we make a model instance for this esperiment
start = 30 # Day with infection
for infested in arange(0,11,1): # Loop over values for number of infested from outside
eks = f'EXO_EXPOSED = {infested} {start} {start+1}' # Specify the update
eksdf = model.update_from_list(newdf,eks) # Update the baseline dataframe
mcorona_infection(eksdf,keep = f'{infested} infested at t={start}'); # Run the model on the updated dataframe
# print(eksdf.EXO_EXPOSED[30:40])
mcorona_infection.keep_viz('dead hospital_icu',smpl=(75,250))
# -
# # Try different rate of contact
# +
mcorona_rate_contact = model.from_eq(rcorona) # we make a model instance for this experiment
start = 30 # Day with infection
for RATE_CONTACT in range(0,11,1):
eks = f'''
EXO_EXPOSED = 10 {start} {start+1}
RATE_CONTACT = {RATE_CONTACT}
'''
eksdf = model.update_from_list(newdf,eks)
mcorona_rate_contact(eksdf,keep = f'Rate of contact: {RATE_CONTACT}');
mcorona_rate_contact.keep_viz('DEAD HOSPITAL_ICU')
# -
# # Try different probability of transmission
# +
mcorona_transmission = model.from_eq(rcorona) # we make a model instance for this esperiment
start = 30 # Day with infection
for PROBABILITY_TRANSMISION in arange(0.00,0.1,0.01):
eks = f'''
EXO_EXPOSED = 10 {start} {start+1}
RATE_CONTACT = 4
PROBABILITY_TRANSMISION = {PROBABILITY_TRANSMISION}
'''
eksdf = model.update_from_list(newdf,eks)
mcorona_transmission(eksdf,keep = f'Probability of transmission: {PROBABILITY_TRANSMISION}');
mcorona_transmission.keep_viz('DEAD HOSPITAL_ICU')
# -
# # An interactive interface
# We can also use an interactive interface to make experiment.
#
# Try the to change the different input parametres (try Daily rate of contact
# and Probability of transmission)
# then press **run scenario**.
# Define user interface
mcorona.inputwidget(basedf = empty,
slidedef = {
'Population ' :{'var' : 'POPULATION SUSCEPTIBLE', 'min' : 0.0, 'max' : 100_000_000, 'value' : 10_000_000,'step':100_000,'op':'=start-','dec':0},
'Number of infected t=1' :{'var' : 'EXO_EXPOSED', 'min' : 0.0, 'max' : 1000, 'value' : 1000, 'step':10,'op':'=impulse','dec':0},
'Share of infected with mild symptom' :{'var' : 'NEW_INFECTIOUS_TO_RECOVER_RATE', 'min' : 0.0, 'max' : 1.0, 'value' : 0.1, 'op':'='},
'Share in hospital no ICU and recover':{'var' : 'NEW_HOSPITAL_TO_RECOVER_RATE', 'min' : 0.0, 'max' : 1.0, 'value' : 0.01, 'op':'=' },
'Share in ICU which recovers' :{'var' : 'NEW_INTENSIVE_TO_RECOVER_RATE', 'min' : 0.0, 'max' : 1.0, 'value' : 0.1, 'op':'='},
'Daily rate of contact' :{'var' : 'RATE_CONTACT', 'min' : 0.0, 'max' : 30, 'value' : 0,'step':0.1, 'op':'='},
'Probability of transmission' :{'var' : 'PROBABILITY_TRANSMISION','min' : 0.0, 'max' : 1.0, 'value' : 0.0,'step':0.005, 'op':'=','dec':3},
},
varpat='infectious recovered dead hospital* *_growth',showvar=True);
| Examples/corona/Corona specify model and make eksperiments.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# -*- coding: utf 8
from matplotlib import pyplot as plt
from matplotlib import ticker
from datetime import datetime
import pandas as pd
import numpy as np
plt.style.use('seaborn-colorblind')
plt.ion()
# -
# Dados obtidos do site https://brasil.io/dataset/covid19/files/
#
# Arquivo 'caso_full.csv'
#
# Data de captura: 2021-02-11
players = pd.read_csv('players.csv')
players.head()
# +
jog = players.copy()
correto = np.array(jog["joined"])
l = []
for i in range(0,len(correto)):
l.append(datetime.utcfromtimestamp(correto[i]).strftime('%Y-%m-%d'))
jog["joined"] = l
jog.head()
# -
datas = jog.groupby("joined").count()
datas['new_players'] = datas['player_id']
datas = datas.drop(columns=['player_id', 'username', 'country', 'followers', 'last_online', 'is_streamer', 'status', 'title', 'best_rapid_rating', 'best_blitz_rating', 'best_bullet_rating'])
# +
#Função que retorna os novos jogadores em um determinado mês, a partir de 2015 até 2020 ou no ano de 2020
def newPlayersMes(mes, pandemia=False):
players_mes = datas.copy()
players_mes = players_mes.reset_index()
players_mes = players_mes.rename(columns={"joined": "date"})
filtro = []
for i in range(len(players_mes)):
if pandemia:
filtro.append(players_mes['date'][i][5:7] == mes and int(players_mes['date'][i][0:4]) == 2020)
else:
filtro.append(players_mes['date'][i][5:7] == mes and int(players_mes['date'][i][0:4]) < 2020 and int(players_mes['date'][i][0:4]) > 2015)
players_mes = players_mes[filtro]
return players_mes
# +
#Novos jogadores no mês de janeiro, entre 2015 e 2020
newPlayersMes('01')
# +
#Histograma com os novos jogadoes no mês de janeiro entre 2015 e 2020 (pré pandemia) e em 2020 (pós pandemia)
janeiroPre = newPlayersMes('01')['new_players']
janeiroPos = newPlayersMes('01', True)['new_players']
plt.hist(janeiroPre, ec='black', bins=30)
plt.title('Pre')
plt.show()
plt.hist(janeiroPos, ec='black', bins=30)
plt.title('Pos')
plt.show()
# +
#Função que calcula o intervalo de confiança a partir da fórmula
def calcula_IC(df):
n = df.count()
desvio = np.std(df, ddof = 1)
media = np.mean(df)
inferior = media - 1.96*(desvio/(n**(1/2))) # 5%
superior = media + 1.96*(desvio/(n**(1/2)))
return [ inferior, superior]
# +
#Intervalo de confiança da média de novos jogadores em janeiro, pré pandemia
icJaneiroPre = calcula_IC(janeiroPre)
icJaneiroPre
# -
np.mean(janeiroPre)
# +
#Intervalo de confiança da média de novos jogadores em janeiro, pós pandemia
icJaneiroPos = calcula_IC(janeiroPos)
icJaneiroPos
# -
np.mean(janeiroPos)
# +
#Teste AB
def teste_ab(df_hyp_menor, df_hyp_maior):
return calcula_IC(df_hyp_menor)[1] < calcula_IC(df_hyp_maior)[0]
# -
#Janeiro
teste_ab(newPlayersMes('01')['new_players'], newPlayersMes('01', True)['new_players'])
print("Durante 2020 a média de novos jogadores saiu do intervalo de confiança no mês")
for i in range(1,13):
mes = str(i)
if i < 10:
mes = '0' + mes
print(mes + ': ', teste_ab(newPlayersMes(mes)['new_players'], newPlayersMes(mes, True)['new_players']))
print("Durante 2020 a média de novos jogadores saiu do intervalo de confiança no mês")
ints = [[],[]]
medias = []
for i in range(1,13):
mes = str(i)
if i < 10:
mes = '0' + mes
ic = calcula_IC(newPlayersMes(mes)['new_players'])
mean = np.mean(newPlayersMes(mes, True)['new_players'])
print(mes + ': ', ic, mean, (ic[0] <= mean and ic[1] >= mean))
ints[0].append(ic[0])
ints[1].append(ic[1])
medias.append(mean)
# +
meses = ['01', '02', '03', '04', '05', '06', '07', '08', '09', '10', '11', '12']
zz = np.array(ints)
dd = pd.DataFrame(zz, columns=meses)
plt.figure(figsize=(5,3))
dd.boxplot(grid=False)
plt.title('Intervalos de Confiança de 2016 até 2019')
plt.xlabel('Mês')
plt.ylabel("Novos jogadores")
plt.savefig('ics.pdf')
plt.show()
plt.figure(figsize=(5,3))
plt.yscale('log')
dd.boxplot(grid=False)
plt.ylim(100,10000)
plt.scatter(x=range(1,13), y=medias, alpha=1)
plt.title('Comparação dos Intervalos com as médias de 2020')
plt.xlabel('Mês')
plt.ylabel("Novos jogadores")
plt.savefig('icsmedias.pdf')
plt.show()
# -
| code/analysis/intervalo_meses.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # LINEAR REGRESSION - ASSUMPTIONS AND INTERPRETATIONS
#
# by <NAME>, December, 2020
import numpy as np
import math
import scipy.stats as ss
import seaborn as sns
import sklearn
import sklearn.datasets
import sklearn.model_selection
import statsmodels.api as sm
import pandas as pd
import pickle
from matplotlib import pyplot as plt
# %matplotlib inline
# # The model
#
# In the simplest case of linear regression, sometimes called ordinary linear regression, the scalar output $y$ is assumed to be a linear combination of the inputs $\mathbf{x}$, and the observation errors follow a Gaussian white noise distribution, thus
#
# $$ y | \mathbf{w}, \sigma, \mathbf{x} \sim \mathcal{N}(w_0 + \sum_{m=1}^{M-1} w_m x_m, \sigma^2) = \mathcal{N}(\mathbf{w}^T \mathbf{x}, \sigma^2), $$
#
# where we have augmented the input vector $\mathbf{x}$ with an additional first element, which is always 1, thus $\mathbf{x} = (1, x_1, x_2, ..., x_{M-1})$.
#
# If we now consider a training data set $\mathcal{D}=\{ \mathbf{x}[n], y[n] \}_{n=1}^N = \{ \mathbf{X}, \mathbf{y} \}$, where $\mathbf{X}$ is a $ N \times M $ design matrix and $\mathbf{y}$ is a column vector of the corresponding output observations, the joint likelihood of the training data may be written:
#
# $$ \mathbf{y} | \mathbf{w}, \sigma, \mathbf{X} \sim \prod_{n=1}^N \mathcal{N}(\mathbf{w}^T \mathbf{x}[n], \sigma^2) = \mathcal{N}(\mathbf{X}\mathbf{w},\sigma^2\mathbf{I}), $$
#
# where $\mathbf{I}$ is an $ N \times N $ identity matrix. See e.g. Wasserman (2004) for further details on linear regression.
#
# ***
# <NAME>. (2004). All of statistics: a concise course in statistical inference. Springer Science & Business Media.
# ***
#
# # Sample data
#
# We start by generating a data set, which we will study in the remainder of this tutorial. In this regard, we will use functionalities from the library ``scikit-learn`` to generate a toy example with five input variables and one output variable. The inputs generated are independent, standard normal and appear in an input matrix below. This matrix is later converted to a design matrix, by including a vector of ones, to account for a potential bias in the output. The model outputs and true coefficients appear as vectors below.
# +
# generate data (matrix of inputs, vector of outputs and true underlaying coefficients)
nX = 200
mX, mXinf = 5, 3
std_y = 5
bias_y = 50
X, y, coef = sklearn.datasets.make_regression(n_samples=nX, n_features=5, n_informative=3, n_targets=1, # note that only 3 inputs are informative!
bias=bias_y, noise=std_y, shuffle=True, coef=True, random_state=100)
y = y.reshape(-1,1)
coef = np.hstack([bias_y, coef])
# Split data into training and test
Xtr, Xte, ytr, yte = sklearn.model_selection.train_test_split(X, y, train_size=.5, shuffle=True, random_state=42)
nXtr, mXtr = Xtr.shape
nXte, mXte = Xte.shape
# plotting
varNames = ['x1', 'x2', 'x3', 'x4', 'x5','y']
dfXy = pd.DataFrame(np.hstack((Xtr, ytr)), columns=varNames)
pd.plotting.scatter_matrix(dfXy, figsize=(10, 10), marker='.', hist_kwds={'bins': 20}, s=60, alpha=.8);
# -
# From the scatter matrix above, we see no apparent dependence among the inputs, and we observe a (linear) relationship between the output and input $x_1$, $x_2$ and $x_4$, respectively. There is no apparent relationship between the output and the remaining outputs.
#
# # Ordinary least squares solution
#
# See my tutorial on [*Linear regression*](https://nbviewer.jupyter.org/github/SebastianGlavind/PhD-study/blob/master/Linear-regression/LinReg.ipynb) for the mathematical details on how to conduct inference in this setting.
#
# Note that the least squares solution has the following properties
#
# $$ E[\mathbf{w}] = E[(\mathbf{X}^T \mathbf{X})^{-1} \mathbf{X}^T \mathbf{y}] = E[ (\mathbf{X}^T \mathbf{X})^{-1} \mathbf{X}^T (\mathbf{X}\mathbf{w} + \epsilon)] ] = \mathbf{w}$$
#
# $$ Var[\mathbf{w}] = Var[(\mathbf{X}^T \mathbf{X})^{-1} \mathbf{X}^T \mathbf{y}] = (\mathbf{X}^T \mathbf{X})^{-1} \mathbf{X}^T Var[Y] \mathbf{X} (\mathbf{X}^T \mathbf{X})^{-1} = (\mathbf{X}^T \mathbf{X})^{-1} \mathbf{X}^T \sigma^2 \mathbf{I} \ \mathbf{X} (\mathbf{X}^T \mathbf{X})^{-1} = \sigma^2 (\mathbf{X}^T \mathbf{X})^{-1}.$$
#
# Thus, it is unbiased with a closed form solution for the covariance, see e.g. Wasserman (2004). Under the assumption of normality of the noise precess $\epsilon$ ($\mathbf{X}$ is fixed), it follows that $\mathbf{w}$ also follow a Gaussian distribution with mean $E[\mathbf{w}]$ and covariance $Var[\mathbf{w}]$, as written above.
# +
def myOLS(XX,y):
nX, mX = XX.shape;
Wols = np.linalg.pinv(XX).dot(y);
mu_ols = np.dot(XX, Wols)
SSE_ols = np.sum( (y - mu_ols)**2)
sigma_ols_unbias = np.sqrt(SSE_ols / (nX-mX))
Rsq = 1 - SSE_ols / np.sum( (y-np.mean(y))**2 )
return(Wols, sigma_ols_unbias, Rsq)
def myMSE(XX, y, W):
nX = XX.shape[0];
yest = np.dot(XX, W)
ydiff = y - yest
mse = np.dot(ydiff.T, ydiff)/nX
return(mse[0,0], ydiff)
XXtr = np.hstack([np.ones(nXtr).reshape(-1,1), Xtr])
XXte = np.hstack([np.ones(nXte).reshape(-1,1), Xte])
Wols, sigma_ols_unbias, Rsq = myOLS(XXtr,ytr)
covW = sigma_ols_unbias**2 * np.linalg.inv( np.dot(XXtr.T, XXtr) )
mse_te, ydiff_te = myMSE(XXte, yte, Wols)
print('Point estimate for OLS parameters: ', np.round(Wols.T,2)[0])
print('Standard error for OLS parameters: ', np.round(np.diag(covW),2) )
print('Unbiased estimate of output standard error: ', round(sigma_ols_unbias,2))
print('R-squared metric: ', np.round(Rsq,2))
print('MSE on test set: ', np.round(mse_te,2))
print('-------------------------------------------------------------------------------------')
print('True parameters: ', np.round(coef,2))
print('True standard deviation: ', round(std_y,2))
# -
# The model fit is plotted against the training and test set below;
# +
# redidual scatter plot
mu_pred_tr = np.dot(XXtr, Wols)
mu_pred_te = np.dot(XXte, Wols)
mu_pred = np.concatenate([mu_pred_tr, mu_pred_te])
x_plot_fit = y_plot_fit = np.array([np.min(mu_pred), np.max(mu_pred)])
plt.figure(figsize=(10,10))
plt.plot(mu_pred_tr, ytr, 'ob', label='Training set')
plt.plot(mu_pred_te, yte, 'or', label='Test set')
plt.plot(x_plot_fit, y_plot_fit, '-k', label='Regression line')
plt.xlabel('Predicted value')
plt.ylabel('Observed value')
plt.legend()
plt.grid()
plt.show()
# -
# Another common way of visualizing the regression is by plotting the regression on the individual inputs, where the remaining inputs are held constant at their mean value.
plt.figure(figsize=(10,2))
for i in range(5):
plt.subplot(1,5,i+1)
index_not_i = np.setdiff1d(np.arange(0,5,1), i+1)
XXtr_fit_i = XXtr.copy();
XXtr_fit_i[:,index_not_i] = np.mean(XXtr[:,index_not_i],0)
x_plot_fit_i = XXtr_fit_i[:,i+1]
mu_plot_fit_i = np.dot(XXtr_fit_i, Wols) # as mu of all x_i ~ 0, we could also simply have used f = Wols[0] + XXtr[:,i+1]*Wols[i+1]
plt.plot(x_plot_fit_i, ytr, 'ob', label='Training set')
plt.plot(x_plot_fit_i, mu_plot_fit_i, '-k', label='Regression line')
plt.xlabel('x'+str(i+1))
if i==0:
plt.ylabel('y')
else:
plt.yticks([])
# plt.legend()
plt.ylim([-60,160])
# As apparent from the subplots above, we again observe a (linear) relationship between the output and input $x_1$, $x_2$ and $x_4$, respectively, and no apparent relationship between the output and the remaining outputs. This is in agreement with the insignificant regression coefficients found above for variable $x_3$ and $x_5$.
#
# # Assessment of the model assumptions
#
# Following Gelman et al. (2020), the assumptions of linear regression are listed below in decreasing order of importance.
#
# 1. Validity
# 2. Representativeness
# 3. Additivity and linearity
# 4. Independence of errors
# 5. Equal variance of errors
# 6. Normality of errors
#
# In the following, we will address these one by one, along with associated interpretations.
#
# ***
# <NAME>., <NAME>., & <NAME>. (2020). Regression and other stories. Cambridge University Press.
# ***
#
# ## Assumption 1: Validity
#
# The fundamental assumption of linear regression, and modeling in general, is that the data being analyzed reflect the research question. This means that the outcome measure should reflect the phenomena under study, including all relevant predictors (inputs), such that the model can generalize to the cases for which it will be used for inference. As an example, a model of incomes will not necessarily tell us anything about patterns of total assets, see Gelman et al. (2020, ch.11).
#
# We will not dive further into this assumption, as the objective of this tutorial is the numerical assessment of the assumptions of linear regression, which we study on a toy example only for illustration purposes.
#
# ## Assumption 2: Representativeness
#
# This assumption addresses the fact that a regression model build from sample data is generally employed to make inferences about a larger population, i.e. we assume that the data is representative of the distribution of the output $y$ given inputs $\{x_i\}_{i=1}^M$, which are included in the modeling. As an example, in a general regression of earnings on sex and height, it would be acceptable for women and tall people to be overrepresented in the sample, compared to the general population, but it would be problematic if the sample include too many rich people $-$ selection on $x_i$ does not interfere with inference from the regression model, but selection on $y$ does! See Gelman et al. (2020, ch.11)
#
# Again, we will not dive further into this assumption, as it is outside the scope of this tutorial, but ...
#
# ### Multicollinearity
#
# Both the validity and representativeness assumption points to the inclusion of more predictors in the regression, as (i) a regression should include all predictors of relevant in predicting the phenomena of interest, and (ii) it allows the representativeness, conditional on $\mathbf{X}$, to be more reasonable. Thus, choosing the inputs to a regression model is a crucial and challenging part of a regression analysis, and there are some pitfalls related to identifiability issues, which can make the model less interpretable. The most familiar and important case of non-identifiability is multicollinearity, see Gelman et al. (2020, ch.10-11)
#
# Multicollinearity is a common problem when estimating linear and generalized linear models. It occurs when there are high correlations among input variables, leading to unreliable and unstable estimates of regression coefficients. This means that multicollinearity make it difficult to choose the correct predictors (inputs) to include in the model and interferes in determining the precise effect of each predictor, but it does not affect the overall fit of the model or the predictions! In the following, we will study these effects in turn.
#
# #### The uncorrelated case (no multicollinearity)
#
# Again, as apparent from the scatter matrix above (*Sample data* section) we observe no immediate dependence among the inputs in the training set of our tay example. This may be further assessed by studying the corresponding correlation matrix;
# +
dfCorr=dfXy.corr() # correlation matrix
plt.figure(figsize = (10,10))
heatplt=sns.heatmap(dfCorr, annot=True, fmt=".3f", vmin=-1, vmax=1, cmap='gray', annot_kws={"size": 12},square=True);
cbar = heatplt.collections[0].colorbar
plt.show()
# -
# The correlation matrix tells as similar story as the scatter matrix, i.e. only week correlations exist among the inputs. Note that it is only the informative inputs, i.e. $x_1$, $x_2$ and $x_4$, which correlate significantly with the output. Thus, the correlation matrix also provides information on the impact of each input on the output.
#
# A commonly used measure for assessing multicollinearity is the variance inflation factor (VIF), which is defined as
#
# $$ \text{VIF}(X_i) = (1- \{R^2|\mathbf{X}_{\sim i}\})^{-1}, $$
#
# where $R^2$ is calculated for each input $X_i$ by performing a linear regression of that input on all the remaining inputs $\mathbf{X}_{\sim i}$. Note that the VIF may easily be found as the diagonal terms of the inverse correlation matrix for normalized inputs, see e.g. [math.stackexchange.com](https://math.stackexchange.com/questions/2681911/prove-vif-of-jth-predictor-is-jth-diagonal-entry-of-inverse-of-correlation-matri).
#
# VIFs range from 1 and upwards, where the numerical values indicate how much the variance of each coefficient is inflated. For example, a VIF of 1.9 indicates that the variance of that particular coefficient is 90% bigger than what you would expect if there was no multicollinearity, see e.g. [statisticshowto.com](https://www.statisticshowto.com/variance-inflation-factor/). As a rule of thumb, a VIF around 0 indicates no correlation, a VIF around 5 indicates moderate correlation, and a VIF around 10 indicates high correlation.
# +
VIF = np.diag(np.linalg.inv(dfCorr.to_numpy(copy=True)[:5,:5]))
print('Variance inflation factors: ', np.round(VIF,3))
# Test
# from statsmodels.stats.outliers_influence import variance_inflation_factor
# from statsmodels.tools.tools import add_constant
# dfXX = dfXy.copy()
# dfXX = dfXX.drop('y', axis=1); # print(dfXX.head())
# dfXX = add_constant(dfXX); # print(dfXX.head())
# pd.Series([variance_inflation_factor(dfXX.values, i) for i in range(dfXX.shape[1])], index=dfXX.columns)
# -
# As expected, the VIFs take on values around 1, i.e. no variance inflation, and thus we conclude that multicollinearity is not a problem in this case.
#
# #### The correlated case (multicollinearity)
#
# We can easily convert our uncorrelated (multivariate) Gaussian sample to a correlated sample using the Cholesky decomposition as
#
# $$ \mathbf{X}^{(C)} = \boldsymbol\mu + \mathbf{X}^{(I)} \mathbf{L}^T, $$
#
# where the covariance matrix $\Sigma = \mathbf{L}\mathbf{L}^T$, $\mathbf{X}^{(I)}$ and $\mathbf{X}^{(C)}$ are a matrix containing the independent input variables and dependent input variables, respectively.
np.random.seed(1000)
cov = np.array([[1. , 0.90595519, 0.67363845, 0.41111229, 0.20592425],
[0.90595519, 1. , 0.90595519, 0.67363845, 0.41111229],
[0.67363845, 0.90595519, 1. , 0.90595519, 0.67363845],
[0.41111229, 0.67363845, 0.90595519, 1. , 0.90595519],
[0.20592425, 0.41111229, 0.67363845, 0.90595519, 1. ]])
Lcl = np.linalg.cholesky(cov)
Xtr_c = np.dot(Xtr, Lcl.T) # zero mean
Xte_c = np.dot(Xte, Lcl.T) # zero mean
XXtr_c = np.hstack([np.ones(Xtr_c.shape[0]).reshape(-1,1), Xtr_c])
XXte_c = np.hstack([np.ones(Xte_c.shape[0]).reshape(-1,1), Xte_c])
ytr_c = np.dot(XXtr_c, coef).reshape(-1,1) + np.random.normal(loc=0, scale=std_y, size=nXtr).reshape(-1,1)
yte_c = np.dot(XXte_c, coef).reshape(-1,1) + np.random.normal(loc=0, scale=std_y, size=nXte).reshape(-1,1)
# For this correlated sample, we first fit the model;
# +
Wols_c, sigma_ols_unbias_c, Rsq_c = myOLS(XXtr_c,ytr_c)
covW_c = sigma_ols_unbias_c**2 * np.linalg.inv( np.dot(XXtr_c.T, XXtr_c) )
mse_te_c, _ = myMSE(XXte_c, yte_c, Wols_c)
print('Point estimate for OLS parameters: ', np.round(Wols_c.T,2)[0])
print('Standard error for OLS parameters: ', np.round(np.diag(covW_c),2) )
print('Unbiased estimate of output standard error: ', round(sigma_ols_unbias_c,2))
print('R-squared metric: ', round(Rsq_c,2))
print('MSE on test set: ', np.round(mse_te_c,2))
print('-------------------------------------------------------------------------------------')
print('True parameters: ', np.round(coef,2))
print('True standard deviation: ', round(std_y,2))
# -
# Please note that the parameters in this case are ill-specified or non-identifiable (very high coefficient of variation for the regression coefficients).
#
# Next, we repeat the procedure of plotting the correlation matrix and computing the VIFs.
# +
dfXcy = pd.DataFrame(np.hstack((Xtr_c, ytr_c)), columns=varNames)
dfCorrXc=dfXcy.corr() # correlation matrix
plt.figure(figsize = (10,10))
heatplt=sns.heatmap(dfCorrXc, annot=True, fmt=".3f", vmin=-1, vmax=1, cmap='gray', annot_kws={"size": 12},square=True);
cbar = heatplt.collections[0].colorbar
plt.show()
# -
VIFc = np.diag(np.linalg.inv(dfCorrXc.to_numpy(copy=True)[:5,:5]))
print('Variance inflation factors: ', np.round(VIFc,3))
# We observe that the estimated regression coefficients indeed do not reflect the true underlaying relationship (non-interpretable) in this case of sever multicollinearity, but the regression receives a similar $R^2$- and MSE-score, and thus does not suffer from the multicollinearity, as expected.
#
# #### Conclusion on multicollinearity
#
# In conclusion, multicollinearity leads to unreliable and unstable estimates of the regression coefficients (non-interpretable), but it does not affect the overall fit of the model or the predictions. We may deal with multicollinearity by e.g. removing one of two highly correlated inputs or combine the two inputs into one new (joint) input variable.
#
# ## Assumption 3: Additivity and linearity
#
# This assumption relates to the linear regression model as being additive and linear in its predictors (including interactions), e.g. $y = w_0 + w_1 x_1 + w_2 x_2$. If additivity is violated, we may e.g. include interactions or transforms, and if linearity is violated, we may e.g. apply a transform(s) to the input(s). For example, $1/x$ or $log(x)$, Gelman et al. (2020, ch.11).
#
# In our numerical toy example, the scatter matrix and the regression coefficients above (*Sample data* and *Ordinary least squares solution* section) tell a similar story, i.e. an apparent linear relationship between the output and the inputs $x_1$, $x_2$ and $x_4$ respectively. The scatter matrix by means of visualization and the regression coefficients by means of quantitative measures of dependence. Moreover, $R^2 \approx 1.0$ means that approximately $100\%$ of the variance in the original model outputs are captured by the linear regression model. As a rule of thumb, models for which $R^2 > 0.7$ are usually considered sufficient for the model to be representative of the data.
#
# ## Assumption 4: Independence of errors
#
# This assumption relates to the white noise assumption of the error terms, i.e. the errors should not exhibit autocorrelation, an assumption that is violated in temporal, spatial, and multilevel settings, Gelman et al. (2020, ch.11).
#
# ### The independent case
#
# Graphically, we can assess autocorrelation in the data series using the autocorrelation function;
# see https://machinelearningmastery.com/gentle-introduction-autocorrelation-partial-autocorrelation/
mse_tr, ydiff_tr = myMSE(XXtr, ytr, Wols) # residual for training set
acf_bounds_theory = 1.96/np.sqrt(nXtr) # https://otexts.com/fpp2/wn.html
plt.figure(figsize=(10,4))
ax1=plt.subplot(1, 1, 1)
sm.graphics.tsa.plot_acf(ydiff_tr, ax=ax1, title='Autocorrelation: Training set');
plt.plot(np.array([0.5,20.5]), np.array([acf_bounds_theory, acf_bounds_theory]), '--k')
plt.plot(np.array([0.5,20.5]), np.array([-acf_bounds_theory, -acf_bounds_theory]), '--k')
plt.xlabel('Lag')
plt.ylabel('ACF')
plt.show();
# The plot includes the empirical and theoretical 95% confidence interval under a white noise assumption, i.e. the shaded region and the region enclosed by the dotted lines, respectively, see e.g. [wikipedia.org](https://en.wikipedia.org/wiki/Correlogram#Statistical_inference_with_correlograms). In this regard, correlation values outside the confidence bounds are very likely due to correlation and not due to random variation. As an exception, a realization is always perfectly correlated with itself, thus an autocorrelation of 1 is always observed at lag zero. In our plot, this is the only value outside the confidence bounds, which indicate that the independence of errors assumption is appropriate in this case.
#
# Autocorrelation can also be tested by e.g. the Durbin-Watson test, where the null hypothesis is that there is no serial correlation in the data, see e.g. [www.statsmodels.org](https://www.statsmodels.org/stable/diagnostic.html) for more autocorrelation tests. For the Durbin-Watson test, the test statistic is
#
# $$ \sum_{n=2}^N ( e[n]-e[n-1] )^2 {\big/} \sum_{n=1}^N ( e[n] )^2, $$
#
# where $e[n]$ is the realized error for item $n$ in the data series. The test statistic is approximately equal to $2(1-r)$, where $r$ is the sample autocorrelation of the residuals. The test statistic is bounded between 0 and 4, with the midpoint 2 indication no autocorrelation. Values below 2 indicates a positive sample autocorrelation and values above 2 indicates a negative sample autocorrelation. Taking the theoretical bounds from before, we would expect a data series without serial correlation to fall in the range $2( 1 \pm 1.96 / \sqrt{N})$. This is the case for our data;
stat_dw = sm.stats.stattools.durbin_watson(ydiff_tr); print('Durbin-Watson test statistic: ', stat_dw)
bounds_dw = 2 * (1 + np.array([-1,1])*1.96/np.sqrt(nXtr)); print('95% confiden bounds: ', bounds_dw)
# ### Some dependent cases
#
# For illustration purposes, some examples where the independence of errors assumption is violated are depicted below.
#
# #### Moving average model of order 2 (MA(2))
# Simulate MA(2) process
np.random.seed(42)
z_autoEr = np.random.normal(loc=0, scale=1, size=1000); nZautoEr = len(z_autoEr)
ma2 = np.zeros([nZautoEr])
for i in np.arange(2,nZautoEr):
ma2[i] = z_autoEr[i] + 0.7*z_autoEr[i-1] + 0.5*z_autoEr[i-2]
# +
# Plot simulation results
plt.figure(figsize=(10,10))
ax1=plt.subplot(2, 1, 1)
plt.plot(ma2[2:])
plt.ylabel('Residual')
plt.xlabel('Samples')
plt.title('Time series of MA(2) process')
ax2=plt.subplot(2, 1, 2)
sm.graphics.tsa.plot_acf(ma2[2:], ax=ax2, title='Autocorrelation: MA(2) process');
plt.xlabel('Lag')
plt.ylabel('ACF')
plt.show();
# -
# #### Autoregressive model of order 1 (AR(1))
# Simulate AR(1) process
ar2 = z_autoEr.copy();
for i in np.arange(1,nZautoEr):
ar2[i] = 0.9*ar2[i-1] + z_autoEr[i]
# +
# Plot simulation results
plt.figure(figsize=(10,10))
ax1=plt.subplot(2, 1, 1)
plt.plot(ar2[2:])
plt.ylabel('Residual')
plt.xlabel('Samples')
plt.title('Time series of AR(1) process')
ax2=plt.subplot(2, 1, 2)
sm.graphics.tsa.plot_acf(ar2[2:], ax=ax2, title='Autocorrelation: AR(1) process');
plt.xlabel('Lag')
plt.ylabel('ACF')
plt.show();
# -
# For the MA(2) and AR(1) above, we observe that the ACF plot exhibits different characteristics. Thus, for the MA(2) process, we see that the first two lags have a significant correlation for the values in the data series, which is consistent with the MA(2) generating process; for the AR(1) process, the correlation falls off gradually, which is a general characteristic of AR processes, when depicted using an ACF plot.
#
# ## Assumption 5: Equal variance of errors (homoscedasticity)
#
# This assumption relates to the constant variance assumption of the noise term. If this is not the case, our model does not reflect the generative process of the data, and probabilistic predictions using the model would not be reliable. However, it does not affect the parametric model estimation, see Gelman et al. (2020, ch.11).
#
# ### The homoscedasticity case
#
# A scatter plot of the residual values on the predicted values is one way of assessing the constant variance assumption. Thus, there should be no clear patterns in the residuals over the predicted values in the residual plot.
mu_pred_range_plot = np.array([np.min(np.concatenate([mu_pred_tr, mu_pred_te])), np.max(np.concatenate([mu_pred_tr, mu_pred_te]))])
plt.figure(figsize=(10, 10))
plt.plot(mu_pred_tr, ydiff_tr,'ob', label='Training set')
plt.plot(mu_pred_te, ydiff_te,'or', label='Test set')
plt.plot(mu_pred_range_plot, [1.96*sigma_ols_unbias]*2, '--k', label='95% bounds')
plt.plot(mu_pred_range_plot, [-1.96*sigma_ols_unbias]*2, '--k')
plt.ylabel('Residual')
plt.xlabel('Predicted value')
plt.legend()
plt.show()
# From the residual plot, it appears that there are no clear patterns in the residuals over the predicted values in this example.
#
# Note that if we consider a plot of the residuals on the observed values, we cannot be sure not to see a pattern, as the model only assumes that the errors are independent of the inputs $\mathbf{x}[n]$ and not the output $y[n]$, see Gelman et al.(2020, ch.11).
#
# ### Some heteroscedasticity cases
#
# For illustration purposes, some examples where the constant variance assumption is violated are depicted below;
# Simulate heteroscedastic errors
y_pred_hetEr = np.arange(-3,3,0.1); nYpred_autoEr= len(y_pred_hetEr)
y_res_hetEr_ex1 = (y_pred_hetEr**2).reshape(-1,1)*np.random.normal(loc=0,scale=1, size=(len(y_pred_hetEr),10)) # Ex1
y_res_hetEr_ex2 = np.exp(-np.linspace(0, 5, num=nYpred_autoEr)).reshape(-1,1)*np.random.normal(loc=0,scale=1, size=(nYpred_autoEr,10)) # Ex2
# +
# Plot simulations
mu_pred_range_plot = np.array([np.min(np.concatenate([mu_pred_tr, mu_pred_te])), np.max(np.concatenate([mu_pred_tr, mu_pred_te]))])
plt.figure(figsize=(10, 10))
plt.subplot(2,1,1)
plt.plot(y_pred_hetEr, y_res_hetEr_ex1,'ob')
plt.ylabel('Residual')
plt.xlabel('Predicted value')
plt.title('Example 1: Large errors for large absolute values of prediction')
plt.subplot(2,1,2)
plt.plot(y_pred_hetEr, y_res_hetEr_ex2,'ob')
plt.ylabel('Residual')
plt.xlabel('Predicted value')
plt.title('Example 2: Decreasing variance in errors with prediction')
plt.show()
# -
# ## Assumption 6: Normality of errors
#
# The distribution of errors is, as in the case of Assumption 5, mostly relevant for probabilistic predictions using the model, as it usually does not affect the parametric model estimation. Note that linear regression does not assume or require that inputs follow a Gaussian distribution, and the Gaussian assumption on the output refer to the error and not the raw data. Depending on the structure of the inputs, it is possible that data $\mathbf{y}$ does not conform with a normality assumption, but the regression errors do, see Gelman et al. (2020, ch.11).
#
# In this tutorial, we will consider a set of graphical tools and statistical test (normality tests) for assessing whether a sample can be assumed to be drawn from a Gaussian distribution.
#
# ### Graphical tools
#
# First, we consider scatter plots, histograms, and Q-Q plots for assessing normality.
# +
# residuals of both training and test set (used below for assessing normality)
ydiff = np.concatenate([ydiff_tr, ydiff_te])
plt.figure(figsize=(10, 4))
plt.subplot(1, 2, 1)
plt.plot(ydiff_tr,'ob', label='Training set')
plt.plot(ydiff_te,'or', label='Test set')
plt.ylabel('Residual')
plt.xlabel('Sample')
plt.title("Scatter plot")
plt.legend()
plt.subplot(1,2,2)
plt.hist(ydiff, density=True,orientation='horizontal')
plt.xlabel('Density')
plt.title("Histogram")
plt.show()
# +
plt.figure(figsize=(10, 4))
plt.subplot(1, 2, 1)
plt.plot(ydiff_tr,'ob', label='Training set')
plt.plot(ydiff_te,'or', label='Test set')
plt.ylabel('Residual')
plt.xlabel('Sample')
plt.title("Scatter plot")
plt.legend()
plt.subplot(1,2,2)
ss.probplot(((ydiff_te-np.mean(ydiff_te))/np.std(ydiff_te)).reshape(-1), dist="norm", plot=plt)
plt.title("Normal Q-Q plot")
plt.show()
# -
# From the scatter/histogram figure, we see that the data is centered around zero and has a Gaussian-like shape (bell-shape), where most of the probability mass is located around the mean.
#
# From the scatter/Q-Q plot figure, we see that there are a few small deviations from the theoretical normal quantiles, especially in the lower and upper tail, which is to be expected given the small data sample.
#
# ### Normality tests
#
# Next, we consider the Shapiro-Wilk test, D’Agostino’s K^2 test and Anderson-Darling test for assessing normality, see e.g. Wasserman (2004) for further details on hypothesis testing, and the ``SciPy`` documentation for details on the implementation of the test.
#
# Note that in the ``SciPy`` implementations used in this tutorial, we can interpret the $p$ value as follows:
#
# - p > $\alpha$: we cannot reject $H_0$ (normal).
# - p <= $\alpha$: we reject $H_0$ (not normal).
# see https://machinelearningmastery.com/a-gentle-introduction-to-normality-tests-in-python/
stat_sha, p_sha = ss.shapiro(ydiff)
print('Statistics=%.3f, p=%.3f' % (stat_sha, p_sha))
# interpret
alpha = 0.05
if p_sha > alpha:
print('We cannot reject H0 (normal)')
else:
print('We reject H0 (not normal)')
# see https://machinelearningmastery.com/a-gentle-introduction-to-normality-tests-in-python/
stat_k2, p_k2 = ss.normaltest(ydiff)
print('Statistics=%.3f, p=%.3f' % (stat_k2, p_k2))
# interpret
alpha = 0.05
if p_k2 > alpha:
print('We cannot reject H0 (normal)')
else:
print('We reject H0 (not normal)')
result_and = ss.anderson(ydiff.reshape(-1))
# print('Statistic: %.3f' % result_and.statistic)
p = 0
for i in range(len(result_and.critical_values)):
sl, cv = result_and.significance_level[i], result_and.critical_values[i]
if result_and.statistic < result_and.critical_values[i]:
print('Significance level %.3f (critical value %.3f): We cannot reject H0 (normal)' % (sl, cv))
else:
print('Significance level %.3f (critical value %.3f): We reject H0 (not normal)' % (sl, cv))
# In line with the graphical assessment, all the normality test indicate that we cannot reject a null-hypothesis of normality with a significance level of 5% (and lower). This is not always the case in practice, where there can be disagreement between the tests (and plots). Here engineering judgement comes into play, e.g. if the data looks normal and/or some of the normality test indicate that we cannot reject normality, maybe a Gaussian assumption is reasonable, or maybe we can transform the output variable to make it conform with a Gaussian assumption, see e.g. [machinelearningmastery.com](https://machinelearningmastery.com/how-to-transform-data-to-fit-the-normal-distribution/).
#
# # Interpretations of linear models
#
# ## Regression coefficients
#
# The correct interpretation of a regression coefficient is the average difference in the outcome, when the corresponding input is changed by one unit, under the assumption of linearity in the inputs. Please note that this relates to population level inferences, whereas the causal interpretation of regression coefficients is the effect of fixing an input, and thus relates to inferences for individuals. To illustrate the difference in interpretation, consider the regression of earnings on years of education. On a population level, this related to the average difference in earnings, when the years of educations is increased by one year in the population. On an individual level, this relates to the effect on earnings for a single person, when increasing this persons years of education by one year, see Gelman et al.(2020, ch.11).
#
# ## Sensitivity analysis
#
# A well fitting linear regression model ($R^2 \geq 0.7$) also provides information regarding the output sensitivites in the inputs/factors, i.e. how much does the variance in input $x_i$ impact the variance in $y$, see my tutorial on [*Variance-based sensitivity analysis for independent inputs*](https://nbviewer.jupyter.org/github/SebastianGlavind/PhD-study/blob/master/Sensitivity-analysis/SA_varianceBased_independentInputs.ipynb) for more information.
#
# The squared, standardized regression coefficients provides the means for what is typically refrerred to as factor prioritization in sensitivity analysis, i.e. the reduction in output variance, if an input set to its true value;
# +
# Standardized regression coefficients acc. variance contribution
SRC = Wols * np.std(XXtr,0).reshape(-1,1) / np.std(ytr)
# Squared SRC
SRCsq = SRC**2;
print( 'Squared, standardized regression coefficients: ', np.round(SRCsq[range(1,6)],2).T, ', sum: ', np.round(np.sum(SRCsq[range(1,6)]),2) ) # almost the same as analytical values
print( 'R-squared for the linear regression model: ', np.round(Rsq,2) )
# -
# From the sensitivity indices above, it appears that $x_2$ is by far the one with the greatest impact on the $y$ and should be prioritized in relation to potential model refinements. We also observe an output variance contribution from $x_1$ and $x_4$ but not from $x_3$ and $x_5$, which is consistent with all findings above.
| Linear-regression/LinReg_assumptionsEtc.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lemmatization
#
# ## <NAME> (DTU/2K16/MC/013)
#
# ## Natural Language Processing (NLP) - Dr. <NAME>
#
# ### Overview
# We introduce lemmatization in this Notebook and alsous lemmatization to process our resume to lemmatize each toke i the corpus (Resume). We also compare the results with previously seen Porter Stemmer algorithm on our Resume.
#
# <img src="../assets/lemmatization.png" width="400px" />
#
# ### 1. Introduction
#
# For grammatical reasons, documents are going to use different forms of a word, such as organize, organizes, and organizing. Additionally, there are families of derivationally related words with similar meanings, such as democracy, democratic, and democratization. In many situations, it seems as if it would be useful for a search for one of these words to return documents that contain another word in the set.
#
# The goal of both stemming and lemmatization is to reduce inflectional forms and sometimes derivationally related forms of a word to a common base form. For instance:
#
#
# _am, are, is_ $\Rightarrow$ __be__
#
# _car, cars, car's, cars'_ $\Rightarrow$ __car__
#
# The Result of such mappings can b something like this:
#
# _the boy's cars are different colors_ $\Rightarrow$ _the boy car be differ color_
#
# However, the two words differ in their flavor. Stemming usually refers to a crude heuristic process that chops off the ends of words in the hope of achieving this goal correctly most of the time, and often includes the removal of derivational affixes. Lemmatization usually refers to doing things properly with the use of a vocabulary and morphological analysis of words, normally aiming to remove inflectional endings only and to return the base or dictionary form of a word, which is known as the lemma .
#
# If confronted with the token _saw_, stemming might return just _s_, whereas lemmatization would attempt to return either _see_ or _saw_ depending on whether the use of the token was as a verb or a noun. The two may also differ in that stemming most commonly collapses derivationally related words, whereas lemmatization commonly only collapses the different inflectional forms of a lemma.
#
# Linguistic processing for stemming or lemmatization is often done by an additional plug-in component to the indexing process, and a number of such components exist, both commercial and open-source.
# ## 2. Importing basic tools required for tokenization, stemming and lemmatization of the resume corpus.
import nltk
import pickle
from collections import Counter
from nltk.stem import WordNetLemmatizer
from nltk.tokenize import word_tokenize
from nltk.corpus import wordnet
nltk.download('wordnet')
# ## 3. Tokenizer
# The Tokenizer is a basic tool used in NLP that returns tokens from a given document. We can create our own token as well based on Regex.
class Tokenizer:
def __init__(self):
self._tokenizer = nltk.RegexpTokenizer(r'\w+')
def tokenize(self, document: str) -> list:
return self._tokenizer.tokenize(document)
# We define a basic tokenizer that takes all word strings from a document as a token. Test it below
# You can also modify the sentence below to see tokens
message = "Have no fear of perfection. You'll never reach it 🔥"
tokenizer = Tokenizer()
print(tokenizer.tokenize(message))
# ## 4. Porter Stemmer
# We now define the Porter Stemmer Algorithm, one of the most famous stemming algorithm for the english language created by <NAME> in 1980. This algorithm reduces a token by removing it's inflections. e.g. _running_ $\rightarrow$ _run_.
#
# We use the porter stemmer as defined in the first assignment which can be seen [here](https://github.com/anishLearnsToCode/porter-stemmer).
class PorterStemmer:
def __init__(self):
"""The word is a buffer holding a word to be stemmed. The letters are in the range
[start, offset ... offset + 1) ... ending at end."""
self.vowels = ('a', 'e', 'i', 'o', 'u')
self.word = ''
self.end = 0
self.start = 0
self.offset = 0
self._tokenizer = Tokenizer()
def is_vowel(self, letter):
return letter in self.vowels
def is_consonant(self, index):
""":returns True if word[index] is a consonant."""
if self.is_vowel(self.word[index]):
return False
if self.word[index] == 'y':
if index == self.start:
return True
else:
return not self.is_consonant(index - 1)
return True
def m(self):
"""m() measures the number of consonant sequences between start and offset.
if c is a consonant sequence and v a vowel sequence, and <..>
indicates arbitrary presence,
<c><v> gives 0
<c>vc<v> gives 1
<c>vcvc<v> gives 2
<c>vcvcvc<v> gives 3
....
"""
n = 0
i = self.start
while True:
if i > self.offset:
return n
if not self.is_consonant(i):
break
i += 1
i += 1
while True:
while True:
if i > self.offset:
return n
if self.is_consonant(i):
break
i += 1
i += 1
n += 1
while True:
if i > self.offset:
return n
if not self.is_consonant(i):
break
i += 1
i += 1
def contains_vowel(self):
""":returns TRUE if the word contains a vowel in the range [start, offset]"""
for i in range(self.start, self.offset + 1):
if not self.is_consonant(i):
return True
return False
def contains_double_consonant(self, j):
""":returns TRUE if the word contain a double consonant in the range [offset, start]"""
if j < (self.start + 1):
return False
if self.word[j] != self.word[j - 1]:
return False
return self.is_consonant(j)
def is_of_form_cvc(self, i):
""":returns TRUE for indices set {i-2, i-1, i} has the form consonant - vowel - consonant
and also if the second c is not w,x or y. this is used when trying to
restore an e at the end of a short e.g.
cav(e), lov(e), hop(e), crim(e), but
snow, box, tray.
"""
if i < (self.start + 2) or not self.is_consonant(i) or self.is_consonant(i - 1) or not self.is_consonant(i - 2):
return 0
ch = self.word[i]
if ch == 'w' or ch == 'x' or ch == 'y':
return 0
return 1
def ends_with(self, s):
""":returns TRUE when {start...end} ends with the string s."""
length = len(s)
if s[length - 1] != self.word[self.end]: # tiny speed-up
return False
if length > (self.end - self.start + 1):
return False
if self.word[self.end - length + 1: self.end + 1] != s:
return False
self.offset = self.end - length
return True
def set_to(self, s):
"""sets [offset + 1, end] to the characters in the string s, readjusting end."""
length = len(s)
self.word = self.word[:self.offset + 1] + s + self.word[self.offset + length + 1:]
self.end = self.offset + length
def replace_morpheme(self, s):
"""is a mapping function to change morphemes"""
if self.m() > 0:
self.set_to(s)
def remove_plurals(self):
"""This is step 1 ab and gets rid of plurals and -ed or -ing. e.g.
caresses -> caress
ponies -> poni
ties -> ti
caress -> caress
cats -> cat
feed -> feed
agreed -> agree
disabled -> disable
matting -> mat
mating -> mate
meeting -> meet
milling -> mill
messing -> mess
meetings -> meet
"""
if self.word[self.end] == 's':
if self.ends_with("sses"):
self.end = self.end - 2
elif self.ends_with("ies"):
self.set_to("i")
elif self.word[self.end - 1] != 's':
self.end = self.end - 1
if self.ends_with("eed"):
if self.m() > 0:
self.end = self.end - 1
elif (self.ends_with("ed") or self.ends_with("ing")) and self.contains_vowel():
self.end = self.offset
if self.ends_with("at"):
self.set_to("ate")
elif self.ends_with("bl"):
self.set_to("ble")
elif self.ends_with("iz"):
self.set_to("ize")
elif self.contains_double_consonant(self.end):
self.end = self.end - 1
ch = self.word[self.end]
if ch == 'l' or ch == 's' or ch == 'z':
self.end = self.end + 1
elif self.m() == 1 and self.is_of_form_cvc(self.end):
self.set_to("e")
def terminal_y_to_i(self):
"""This defines step 1 c which turns terminal y to i when there is another vowel in the stem."""
if self.ends_with('y') and self.contains_vowel():
self.word = self.word[:self.end] + 'i' + self.word[self.end + 1:]
def map_double_to_single_suffix(self):
"""Defines step 2 and maps double suffices to single ones.
so -ization ( = -ize plus -ation) maps to -ize etc. note that the
string before the suffix must give m() > 0.
"""
if self.word[self.end - 1] == 'a':
if self.ends_with("ational"):
self.replace_morpheme("ate")
elif self.ends_with("tional"):
self.replace_morpheme("tion")
elif self.word[self.end - 1] == 'c':
if self.ends_with("enci"):
self.replace_morpheme("ence")
elif self.ends_with("anci"):
self.replace_morpheme("ance")
elif self.word[self.end - 1] == 'e':
if self.ends_with("izer"): self.replace_morpheme("ize")
elif self.word[self.end - 1] == 'l':
if self.ends_with("bli"):
self.replace_morpheme("ble") # --DEPARTURE--
# To match the published algorithm, replace this phrase with
# if self.ends("abli"): self.r("able")
elif self.ends_with("alli"):
self.replace_morpheme("al")
elif self.ends_with("entli"):
self.replace_morpheme("ent")
elif self.ends_with("eli"):
self.replace_morpheme("e")
elif self.ends_with("ousli"):
self.replace_morpheme("ous")
elif self.word[self.end - 1] == 'o':
if self.ends_with("ization"):
self.replace_morpheme("ize")
elif self.ends_with("ation"):
self.replace_morpheme("ate")
elif self.ends_with("ator"):
self.replace_morpheme("ate")
elif self.word[self.end - 1] == 's':
if self.ends_with("alism"):
self.replace_morpheme("al")
elif self.ends_with("iveness"):
self.replace_morpheme("ive")
elif self.ends_with("fulness"):
self.replace_morpheme("ful")
elif self.ends_with("ousness"):
self.replace_morpheme("ous")
elif self.word[self.end - 1] == 't':
if self.ends_with("aliti"):
self.replace_morpheme("al")
elif self.ends_with("iviti"):
self.replace_morpheme("ive")
elif self.ends_with("biliti"):
self.replace_morpheme("ble")
elif self.word[self.end - 1] == 'g':
if self.ends_with("logi"): self.replace_morpheme("log")
def step3(self):
"""step3() deals with -ic-, -full, -ness etc."""
if self.word[self.end] == 'e':
if self.ends_with("icate"):
self.replace_morpheme("ic")
elif self.ends_with("ative"):
self.replace_morpheme("")
elif self.ends_with("alize"):
self.replace_morpheme("al")
elif self.word[self.end] == 'i':
if self.ends_with("iciti"): self.replace_morpheme("ic")
elif self.word[self.end] == 'l':
if self.ends_with("ical"):
self.replace_morpheme("ic")
elif self.ends_with("ful"):
self.replace_morpheme("")
elif self.word[self.end] == 's':
if self.ends_with("ness"): self.replace_morpheme("")
def step4(self):
"""step4() takes off -ant, -ence etc., in context <c>vcvc<v>."""
if self.word[self.end - 1] == 'a':
if self.ends_with("al"):
pass
else:
return
elif self.word[self.end - 1] == 'c':
if self.ends_with("ance"):
pass
elif self.ends_with("ence"):
pass
else:
return
elif self.word[self.end - 1] == 'e':
if self.ends_with("er"):
pass
else:
return
elif self.word[self.end - 1] == 'i':
if self.ends_with("ic"):
pass
else:
return
elif self.word[self.end - 1] == 'l':
if self.ends_with("able"):
pass
elif self.ends_with("ible"):
pass
else:
return
elif self.word[self.end - 1] == 'n':
if self.ends_with("ant"):
pass
elif self.ends_with("ement"):
pass
elif self.ends_with("ment"):
pass
elif self.ends_with("ent"):
pass
else:
return
elif self.word[self.end - 1] == 'o':
if self.ends_with("ion") and (self.word[self.offset] == 's' or self.word[self.offset] == 't'):
pass
elif self.ends_with("ou"):
pass
# takes care of -ous
else:
return
elif self.word[self.end - 1] == 's':
if self.ends_with("ism"):
pass
else:
return
elif self.word[self.end - 1] == 't':
if self.ends_with("ate"):
pass
elif self.ends_with("iti"):
pass
else:
return
elif self.word[self.end - 1] == 'u':
if self.ends_with("ous"):
pass
else:
return
elif self.word[self.end - 1] == 'v':
if self.ends_with("ive"):
pass
else:
return
elif self.word[self.end - 1] == 'z':
if self.ends_with("ize"):
pass
else:
return
else:
return
if self.m() > 1:
self.end = self.offset
def step5(self):
"""step5() removes a final -e if m() > 1, and changes -ll to -l if m > 1."""
self.offset = self.end
if self.word[self.end] == 'e':
a = self.m()
if a > 1 or (a == 1 and not self.is_of_form_cvc(self.end - 1)):
self.end = self.end - 1
if self.word[self.end] == 'l' and self.contains_double_consonant(self.end) and self.m() > 1:
self.end = self.end - 1
def stem_document(self, document):
result = []
for line in document.split('\n'):
result.append(self.stem_sentence(line))
return '\n'.join(result)
def alphabetic(self, word):
return ''.join([letter if letter.isalpha() else '' for letter in word])
def stem_sentence(self, sentence):
result = []
for word in self._tokenizer.tokenize(sentence):
result.append(self.stem_word(word))
return ' '.join(result)
def stem_word(self, word):
if word == '':
return ''
self.word = word
self.end = len(word) - 1
self.start = 0
self.remove_plurals()
self.terminal_y_to_i()
self.map_double_to_single_suffix()
self.step3()
self.step4()
self.step5()
return self.word[self.start: self.end + 1]
# test the porter stemmer with any word of your choice
stemmer = PorterStemmer()
print(stemmer.stem_word('running'))
# test the porter stemmer with any sentence of your choice
print(stemmer.stem_sentence("the boy's cars are different colors"))
# ## 5. Lemmatization
# We now create a class for lemmatization that will use the `Tokenizer` class. An instance of the Lemmatization class will help us reduce words into their __lemmas__.
class Lemmatizer:
def __init__(self):
self._lemmatizer = WordNetLemmatizer()
self._tokenizer = Tokenizer()
def _tokenize(self, document: str) -> list:
return self._tokenizer.tokenize(document)
def lemmatize_word(self, word: str, pos=None) -> str:
return self._lemmatizer.lemmatize(word, pos) if pos is not None else self._lemmatizer.lemmatize(word)
def lemmatize_sentence(self, sentence: str, pos=None) -> str:
result = []
for word in self._tokenize(sentence):
if pos is not None:
result.append(self.lemmatize_word(word, pos))
else:
result.append(self.lemmatize_word(word))
return ' '.join(result)
def lemmatize_document(self, document: str) -> str:
result = []
for line in document.split('\n'):
result.append(self.lemmatize_sentence(line))
return '\n'.join(result)
# test the lemmatizer with any word of your choice
lemmatizer = Lemmatizer()
print(lemmatizer.lemmatize_word('wolves'))
# try the lemmaztization algorithm with a sentence of your choice
print(lemmatizer.lemmatize_sentence('the quick brown fox 🦊 jumps over the lazy dog 🐶'))
# ## 6. Loading in the Resume
# We now load in our resume and see the sample text in it.
resume_file = open('../assets/resume.txt', 'r')
resume = resume_file.read().lower()
resume_file.close()
print(resume)
# ## 7. Stemming the Resume
# We now Stem our resume by applying the PorterStemmer algorithm on it and see the output.
# +
stemmer = PorterStemmer()
resume_file = open('../assets/resume.txt', 'r')
resume = resume_file.read().lower()
resume_file.close()
resume_stemmed = stemmer.stem_document(resume)
pickle.dump(obj=resume_stemmed, file=open('../assets/resume_stemmed.p', 'wb'))
resume_stemmed_file = open('../assets/resume_stemmed.txt', 'w')
resume_stemmed_file.write(resume_stemmed)
resume_stemmed_file.close()
# -
# We now display the stemmed resume
print(resume_stemmed)
# ## 8. Creating the Lemmatized Resume
# We now use our `Lemmatizer` class to lemmatize our resume and save it so that we can run analytics on it later on.
# +
lemmatizer = Lemmatizer()
resume_file = open('../assets/resume.txt')
resume = resume_file.read().lower()
resume_file.close()
resume_lemmatized = lemmatizer.lemmatize_document(resume)
pickle.dump(resume_lemmatized, open('../assets/resume_lemmatized.p', 'wb'))
resume_lemmatized_file = open('../assets/resume_lemmatized.txt', 'w')
resume_lemmatized_file.write(resume_lemmatized)
resume_lemmatized_file.close()
# -
# displaying lemmatized resume
print(resume_lemmatized)
# ## 9. Analytics
# We now run a few basic analytics and compae the output of the stemmed and lemmaztized Resumes with each other and the original resume.
# We load in the original, stemmed and lemmatized resumes
resume_file = open('../assets/resume.txt', 'r')
resume = resume_file.read().lower()
resume_file.close()
resume_stemmed = pickle.load(open('../assets/resume_stemmed.p', 'rb'))
resume_lemmatized = pickle.load(open('../assets/resume_lemmatized.p', 'rb'))
# extracting tokens from the original, stemmed and lemmatized outputs
resume_tokens = word_tokenize(resume)
stemmed_resume_tokens = word_tokenize(resume_stemmed)
lemmatized_resume_tokens = word_tokenize(resume_lemmatized)
# Comparing the number of tokens in original, stemmed and lemmatized outputs
print('No. of tokens in Resume:', len(resume_tokens))
print('No. of tokens in Stemmed Resume:', len(stemmed_resume_tokens))
print('No. of tokens in Lemmatized Resume:', len(lemmatized_resume_tokens))
# We observe that both the stemmed and lemmatized resume's have same number of tokesn which is correct as the tokenization step for both these processes uses the same Tokenization algorithm.
# comparing no. of words and word frequencies in both stemmed and lemmatized outputs
stemmed_resume_frequencies = Counter(stemmed_resume_tokens)
lemmatized_resume_frequencies = Counter(lemmatized_resume_tokens)
print('\nNo. of unique tokens/words in the stemmed output:', len(stemmed_resume_frequencies))
print('No. of unique tokens/words in the lemmatized output:', len(lemmatized_resume_frequencies))
# In the stemmed output there are less number of tokens, but the reduction in number of tokens isn't that high and if the purpose of our task is to reduce the number of tokens in the corpus, stemming is definately a way to go, but lemmatization also achieves similar percentage reduction.
# seeing the top 30 most common words in the stemmed and lemmatized outputs
print('\nTop 30 most common words/tokens in the stemmed output:\n', stemmed_resume_frequencies.most_common(30))
print('\nTop 30 most common words/tokens in the lemmatized output:\n', lemmatized_resume_frequencies.most_common(30))
# In stemming we are getting words reduced down to their roots and words are much more clearer in their meaning and are closer to their original form in the lemmatizaed format. Although in lemmatization we are also receiving a number __4__ in our most frequently ocurring characters.
#
# We now introduce a helpee method that will help us in tagging each token in the corpus with the corresponding Part of Speech Tags (POS tags). POS Tags are mainly of the following types:
#
# - Noun (n)
# - Verb (v)
# - Adjective (a)
# - Adverb (r)
# - Symbol (s)
#
# By tagging the orignal, stemmed and lemmatized resumes we can check whether we are still maintaing the same frequency of POS tags. Which will further show that the meaning or context of our words has been retained despite the pre-processing steps of Stemming or Lemmatization.
# +
def get_pos_frequency(tokens: list) -> Counter:
synsets = [wordnet.synsets(token) for token in tokens]
pos_tags = []
for synset in synsets:
if isinstance(synset, list) and len(synset) > 0:
pos_tags.append(synset[0].pos())
return Counter(pos_tags)
# Analyzing of frequency of POS tags in original, stemmed and Lemmatized resume
resume_pos_frequency = get_pos_frequency(resume_tokens)
stemmed_resume_pos_frequency = get_pos_frequency(stemmed_resume_tokens)
lemmatized_resume_pos_frequency = get_pos_frequency(lemmatized_resume_tokens)
print('\nResume POS Tags Frequency:', resume_pos_frequency)
print('Stemmed Resume POS Tags Frequency:', stemmed_resume_pos_frequency)
print('Lemmatized Resume POS Tags Frequency:', lemmatized_resume_pos_frequency)
# -
# We observe that the number of nouns and adverbs increases in the resume after performing the lemmatization step and the number of nouns clearly decreases after performing stemming. So, if in our application the user wishes to search proper nouns and obtain specific results and only those that match exactly, like searching __java__, __python__ etc. through a resume, the lemmatization will give a better result.
#
# We now see the POS (Part of speech) Tags for those words that have been reduced differently by the stemming and Lemmatization Operations.
# +
# compiling pos tags for words that have been stemmed and lemmatized differently
import pprint
printer = pprint.PrettyPrinter(width=50)
diff = []
for stemmed, lemmatized in zip(stemmed_resume_tokens, lemmatized_resume_tokens):
if not stemmed == lemmatized:
stemmed_synsets = wordnet.synsets(stemmed)
lemmatized_synsets = wordnet.synsets(lemmatized)
stemmed_synset = stemmed_synsets[0].pos() if isinstance(stemmed_synsets, list) and len(stemmed_synsets) > 0 else ''
lemmatized_synset = lemmatized_synsets[0].pos() if isinstance(lemmatized_synsets, list) and len(lemmatized_synsets) > 0 else ''
diff.append({stemmed: stemmed_synset, lemmatized: lemmatized_synset})
print('POS Tags for Words with different stemmed and Lemmatized forms:')
printer.pprint(diff)
# -
# From the above example we can clealy see that teh POS tager present inside the NLTKpackage is unable to tag email ID's, phone numbers, proper nouns such as names etc. for both the _stemmed_ and _lemmatized_ outputs, but it is able to tag ore words in the _lemmatized_ tokens than it is for _stemmed_ words.
#
# This is because in lemmatiation we are reducing the words to valid english words and the POS tagger can still tag it as a Noun, verb etc. wheras in the stemmer in many cases the words are reduced to a form that may be benificial for certain NLP tasks like searching, information extraction etc. but the reduced isn't always a valid english word e.g. _pony_ $\rightarrow$ _poni_ , _happy_ $\rightarrow$ _happi_ , _language_ $\rightarrow$ _languag_.
#
# Hence if we need to use an application where speech tagging is required such as text 2 speech or understanding speech and analyzing speech such as speech to text or machine transaltion Lemmatization can play a vital role wheras Stemming has no percievable bennefits and on the contrary might only reduce application performance.
#
# We can also see in afew examples that stemming has actually resulted in a loss of the contextual meaning of the wrd e.g. in the word _worked_. This word is stemmed down to _work_ and lemmatized down to _worked_. These are then tagged as the follows:
#
# 1. (Stemming) _worked_ $\rightarrow$ __Noun__
# 1. (Lemmatization) _worked_ $\rightarrow$ __Verb__
#
# In the context of our resume, when we wrote worked, we meant it as a verb and not a noun and that context has been lost in stemming. We can see similar results for _learning_ , _development_ and electronic_.
# ## 10. Discussion & Conclusion
# Both stemming and Lemmatization serve a very similar purpose in the NLP preprocessing pipeline and are methods used to reduce a word to it's toot form, they also have many differences.
#
# Stemming reduces a word by removing it's inflections and the morphologically reduced form that it recieives may or may not be an english language word. Stemming uses a heuristic approach to reducing inflections wherein the steps followed are not aware of the meaning of the word or even the context of the word and simply follow basic deterministic rules based on the characters in the word to add and remove character strings in the word.
#
# This isn't how lemmatization functions. The Porter Stemmer Algorithm was introduced in the 1980's whereas Lemmatization is a much more modern algorithm that isn't a simple heuristic based algorithm, but it is aware of the word meaning along with synonyms of the word.
#
# It also tags the word with the correct part of speech (POS) tag, or multiple tags and uses a pre-built dictionary to define the meaning of each and every tag with different contexts (Contexts being Noun, Verb, Adverb etc.) Lemmatization requires this pre-built dictionary to function and in the __nltk__ package that we were using, internally __nltk__ uses the __wordnet__ corpus which contains words, their definitions for all different contexts and also synonyms like a thesaurus.
#
# Lemmatiation using the wordnet corpus can reduce words to words that do not have the same character structure e.g. it can reduce _better_ $\rightarrow$ _good_ if we tell it that _better_ here is a __verb__.
lemmatizer = WordNetLemmatizer()
print(lemmatizer.lemmatize('better', pos='a'))
# reducing a word to a form with dissimilar charaters isn't possible in Stemming
stemmer = PorterStemmer()
print(stemmer.stem_word('better'))
# Stemming reduces words having the same character roots (may not have same meanings) to the same roots and this is then helpful in __IR__ (Information Retrieval) Applications as the person/user can search for small strings like _uni_ or _univer_ and these strings will automatically match to _university_ , _universities_ etc.
#
# So, stemming makes a lot of sense in Information Retrieval Applications. In advanced information retrieval applications where the user can not only enter a stemmed form of what she is searching, but can also enter the context of what she wishes to search such as _better food than x_ (where x is a restaurant). Our IR application should be able to understand that _better_ here refers to _good_ food or better ratings than the ratings for a restaurant.
#
# Or we may have a chatbot application which communicates with the user and the chatbot application needs to understand the intent of the user so that the chatbot can answer queries that the user puts forth. For answering queries or understanding speech and translating to text, or calculating the probability of a given word we require a model that can understand context and not just the root of a word.
#
# In such applications we use lemmatization along with POS (Part of Speech) tagging. Even in machine translations wherein we need to compute the probabilities of the translated text, we need a lemmatizer along with a POS tagger to compute structure and probabilities.
#
# Hence, both the stemmer and the lemmatizer cater to very different needs. In our application we decide whether to use a stemmer or lemmatizer based on what our application must do. In a system with multiple resumes, the most common thing an employer might want to do is search the corpora with specific skills such as _management_ , _java_ , _machine learning_ etc. and then receive resumes with a match for this string.
#
# We can also sort the resumes based on frequency and count of matches, where the resume with a higher number of matches might bubble up to the top. Hence, in our application which is centered more around Information Extraction/Information Retrieval than context understanding Stemming makes more sense.
#
# This may seem counterintuitive as we have seen in the analytics above that lemmatization preserves the POS tags and context structure, but preserving POS tags and context structure will not improve an IR system. Also running a lemmatizer is more compute heavy as it has a higher time complexity.
| notebook/lemmatization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
from matplotlib import pyplot
import warnings
from summer.utils import ref_times_to_dti
from autumn.models.sm_sir.constants import IMMUNITY_STRATA
from autumn.tools.project import get_project
from autumn.settings.constants import COVID_BASE_DATETIME
from autumn.tools.utils.display import pretty_print
from autumn.tools.inputs.covid_phl.queries import get_phl_vac_coverage
# -
pyplot.style.use("ggplot")
warnings.filterwarnings("ignore")
region = "national-capital-region"
project = get_project("sm_sir", region, reload=True)
baseline_params = project.param_set.baseline
model = project.run_baseline_model(baseline_params)
baseline_df = model.get_derived_outputs_df()
model_start_time = ref_times_to_dti(COVID_BASE_DATETIME, [baseline_params["time"]["start"]])[0]
sc_models = project.run_scenario_models(model, project.param_set.scenarios)
scenario_dfs = [m.get_derived_outputs_df() for m in sc_models]
# +
def convert_ts_index_to_date(ts):
ts.index = ref_times_to_dti(COVID_BASE_DATETIME, ts.index)
return ts
ts_set_dates = {project.calibration.targets[i].data.name:
convert_ts_index_to_date(project.calibration.targets[i].data) for
i in range(len(project.calibration.targets))
}
# +
# Define what we want to plot
outputs_to_plot = ["notifications", "infection_deaths", "icu_occupancy", "hospital_occupancy"]
plot_end_time = ref_times_to_dti(COVID_BASE_DATETIME, [1000.])[0]
fig = pyplot.figure(figsize=(15, 12))
for i_out, output in enumerate(outputs_to_plot):
axis = fig.add_subplot(2, 2, i_out + 1)
# Plotting
if output in ts_set_dates:
ts_set_dates[output].plot(style='.', label="")
baseline_df[output].plot(label="baseline")
# for scenario in range(len(sc_models)):
# scenario_dfs[scenario][outputs_to_plot[i_out]].plot(label=project.param_set.scenarios[scenario]["description"])
# Display
axis.set_title(output.replace("_", " "))
if i_out == 0:
axis.legend()
axis.set_xlim(left=model_start_time)
# -
baseline_df[[f"prop_immune_{stratum}" for stratum in IMMUNITY_STRATA[::-1]]].plot.area(figsize=(10, 7))
get_phl_vac_coverage(dose="BOOSTER_DOSE").plot.area()
| notebooks/user/jtrauer/ncr_manual_calibration_2.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.5.3
# language: julia
# name: julia-1.5
# ---
# # A-orthogonalization Process
# $
# \begin{align}
# d_1 . d_2 &= u_1 . (u_2 - \beta_{21} u_1) \\
# &= u_1 . (u_2 - \frac{u_2^T A u_1}{u_1^T A u_1} u_1) \\
# &= 0
# \end{align}
# $
GramSchmidt(A)
# +
using LinearAlgebra
function GramSchmidt(A)
D = A
for i = 2:size(A,2)
for j=1:(i-1)
α = D[:,j]' * A * D[:,j]
β = A[:,i]' * A * D[:,j]
D[:,i] -= D[:,j]*β/α
display([α, β])
end
end
return D
end
# -
# # Test
# +
# SPD matrix
function spd(n)
A = rand(n,n)
vec,val = eigen(A)
A = (A*A')/2
A -= val[1]*I
end
A = spd(3)
# +
D = GramSchmidt(A)
res = D[:,1]'*A*D[:,2], D[:,1]'*A*D[:,3], D[:,2]'*A*D[:,3]
# -
# ## Adjourn
using Dates
println("mahdiar")
Dates.format(now(), "Y/U/d HH:MM")
| HW05/3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ##### Import the required packages
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
import os
import seaborn as sns
import pickle
from collections import Counter
from datetime import datetime
from sortedcontainers import SortedList
# We can override the default matplotlib styles with those of Seaborn
sns.set()## Importing the relevant libraries
# Load the data from a .csv
ratings_data = pd.read_csv(os.path.join(os.path.pardir,'data','raw','ratings.csv'))
movies_data = pd.read_csv(os.path.join(os.path.pardir,'data','raw','movies.csv'))
ratings_df = ratings_data.copy()
movies_df = movies_data.copy()
ratings_df.head()
ratings_df.describe(include='all')
ratings_df.info()
# Make the user IDs go from 0 to N-1
ratings_df['userId'] = ratings_df['userId'] - 1
# +
# Create a mapper for movie IDs
unique_movie_ids = list(ratings_df['movieId'].unique())
movie_id_mapper = {}
count = 0
for movie_id in unique_movie_ids:
movie_id_mapper[movie_id] = count
count +=1
# Add them to the databrame
ratings_df['movieId'] = ratings_df['movieId'].apply(lambda x: movie_id_mapper[x])
ratings_df = ratings_df.drop(columns='timestamp', axis=1)
# +
N = ratings_df['userId'].max() + 1 # Number of users
M = ratings_df['movieId'].max() + 1 # number of movies
# user_id_count = dict(ratings_df['userId'].value_counts())
# movie_id_count = dict(ratings_df['movieId'].value_counts())
user_id_count = Counter(ratings_df['userId'])
movie_id_count = Counter(ratings_df['movieId'])
# Number of users and movies we would like to keep
n = 10000
m = 2000
user_ids = [u for u, c in user_id_count.most_common(n)]
movie_ids = [m for m, c in user_id_count.most_common(m)]
# Make a copy, otherwise the original df won't be overwritten
ratings_df_small = ratings_df[ratings_df['userId'].isin(user_ids) & ratings_df['movieId'].isin(movie_ids)].copy()
# Need to remake user_ids and movie_ids since they are no longer sequential
new_user_id_mapper = {}
i = 0
for old in user_ids:
new_user_id_mapper[old] = i
i += 1
new_movie_id_mapper = {}
j = 0
for old in movie_ids:
new_movie_id_mapper[old] = j
j += 1
ratings_df_small['userId'] = ratings_df_small['userId'].apply(lambda x: new_user_id_mapper[x])
ratings_df_small['movieId'] = ratings_df_small['movieId'].apply(lambda x: new_movie_id_mapper[x])
ratings_df_small.to_csv(os.path.join(os.path.pardir,'data','processed','small_ratings.csv'))
# +
ratings_df_processed = ratings_df_small.copy()
N_ = ratings_df_processed['userId'].max() + 1 #Number of users
M_ = ratings_df_processed['movieId'].max() + 1 # Number of movies
# Split the df into train and test
ratings_df_train, ratings_df_test = train_test_split(ratings_df_processed, test_size=0.2)
# A dictionary to tell us, which users have rated which movie
user_movie = {}
# A dictionary to tell us, which movies have been rated by which users
movie_user = {}
# A dictionary to lookup ratings
user_movie_ratings = {}
cutoff = int(0.8 * len(ratings_df_processed))
count = 0
def update_user_movie_and_movie_user(row):
global count
count += 1
if count % 100000 == 0:
print("Processed: %.3f" % (float(count)/cutoff))
i = int(row['userId'])
j = int(row['movieId'])
if i not in user_movie:
user_movie[i] = [j]
else:
user_movie[i].append(j)
if j not in movie_user:
movie_user[j] = [i]
else:
movie_user[j].append(i)
user_movie_ratings[(i,j)] = row['rating']
ratings_df_train.apply(update_user_movie_and_movie_user, axis=1)
user_movie
# +
user_movie_ratings_test = {}
def update_user_movie_and_movie_user_test(row):
global count
count += 1
if count % 100000 == 0:
print("Processed: %.3f" % (float(count)/cutoff))
i = int(row['userId'])
j = int(row['movieId'])
user_movie_ratings_test[(i, j)] = row['rating']
ratings_df_test.apply(update_user_movie_and_movie_user_test, axis=1)
with open(os.path.join(os.path.pardir,'data','interim','user_movie.json'), 'wb') as f:
pickle.dump(user_movie, f)
with open(os.path.join(os.path.pardir,'data','interim','movie_user.json'), 'wb') as f:
pickle.dump(movie_user, f)
with open(os.path.join(os.path.pardir,'data','interim','user_movie_rating.json'), 'wb') as f:
pickle.dump(user_movie_ratings_test, f)
# +
# Load the data
with open(os.path.join(os.path.pardir,'data','interim','user_movie.json'), 'rb') as f:
user_movie = pickle.load(f)
with open(os.path.join(os.path.pardir,'data','interim','movie_user.json'), 'rb') as f:
movie_user = pickle.load(f)
with open(os.path.join(os.path.pardir,'data','interim','user_movie_rating.json'), 'rb') as f:
user_movie_ratings_test = pickle.load(f)
# +
N = max(user_movie.keys()) + 1
# the test set may contain movies that the train set doesn't have
m1 = max(movie_user.keys())
m2 = max([m for (u,m), r in user_movie_ratings_test.items()])
M = max(m1, m2) + 1
# To find the user-user similarity, you have to do O(N^2 * M) calculations
# In the real world you'd have to parallelize this
# Note: we only have to do half the calculations since w_ij is symetric
K = 25 # The number of neighbours we'd like to consider
limit = 5 # Minimum number of movies the users must have in common
neighbors = [] # store the neighbors in a list
averages = [] # each user's average rating
deviations = [] #each user's deviation
for i in range(N):
# find the 25 closes users to i
try:
movies_i = user_movie[i]
movies_i_set = set(movies_i)
# calculate average and deviation
ratings_i = {movie:user_movie_ratings[(i, movie)] for movie in movies_i}
avg_i = np.mean(list(ratings_i.values()))
dev_i = {movie: (rating-avg_i) for movie, rating in ratings_i.items()}
dev_i_values = np.array(list(dev_i.values()))
# convert all the values in the deviations dictionary into a numpy array.
# This is because the denominator in the Pearson correlation is the square root of the sum of squares
# of the deviations.
sigma_i = np.sqrt(dev_i_values.dot(dev_i_values))
# Save these for later use
averages.append(avg_i)
deviations.append(dev_i)
sl = SortedList()
for j in range(N):
# Don't calculate the correlation with yourself
if j !=i:
movies_j = user_movie[j]
movies_j_set = set(movies_j)
common_movies = (movies_i_set & movies_j_set) # intersetion
if len(common_movies) > limit:
# calculate average and deviation
ratings_j = {movie:user_movie_ratings[(j, movie)] for movie in movies_j}
avg_j = np.mean(list(ratings_j.values()))
dev_j = {movie: (rating-avg_j) for movie, rating in ratings_j.items()}
dev_j_values = np.array(list(dev_j.values()))
sigma_j = np.sqrt(dev_j_values.dot(dev_j_values))
# calculate the correlation coefficient
numerator = sum(dev_i[m] for m in common_movies)
w_ij = numerator/(sigma_i * sigma_j)
# insert into a sorted list and truncate
# negate the weight, because the list is sorted in ascending
# maximum values (1) is "closest"
sl.add((w_ij, j))
if len(sl) > K:
del sl[-1]
# store the neighbors
neighbors.append(sl)
# print out useful things
# if i%i == 0:
# print(i)
except:
pass
# -
# > #### User User Collaborative filtering
# +
# using neighbors to calculate train and test MSE
def predict(i, m): # i: user, m: movie
# calculate the weighted sum of deviation
numerator = 0
denominator = 0
for neg_w, j in neighbors[i]:
# remember, the weight is sorted as its negative
# so, the negative of the negative weight is positive
try:
numerator += -neg_w * deviations[j][m]
denominator += abs(neg_w)
except KeyError:
# neighbors may not have rated the same movie
# don't want to do dictionary lookup twice
# so, just throw an exception
pass
if denominator == 0:
prediction = averages[i]
else:
prediction = numerator / denominator * averages[i]
prediction = min(5, prediction)
prediction = max(0.5, prediction) # max rating is 0.5
return prediction
train_predictions = []
train_targets = []
for (i, m), target in user_movie_ratings.items():
# calculate the predictions of the movie
prediction =predict(i, m)
# save the prediction and target
train_predictions.append(prediction)
train_targets.append(target)
test_predictions = []
test_targets = []
for (i, m), target in user_movie_ratings_test.items():
# calculate the predictions of the movie
prediction =predict(i, m)
# save the prediction and target
test_predictions.append(prediction)
test_targets.append(target)
# calculate accuracy
def mse(p, t):
p = np.array(p)
t = np.array(t)
return np.mean((p - t) ** 2)
print("Train mse", mse(train_predictions, train_targets))
print("Test mse", mse(test_predictions, test_targets))
# -
# > #### Item Item collaboprative filtering
# +
# Load the data
with open(os.path.join(os.path.pardir,'data','interim','user_movie.json'), 'rb') as f:
user_movie = pickle.load(f)
with open(os.path.join(os.path.pardir,'data','interim','movie_user.json'), 'rb') as f:
movie_user = pickle.load(f)
with open(os.path.join(os.path.pardir,'data','interim','user_movie_rating.json'), 'rb') as f:
user_movie_ratings_test = pickle.load(f)
# +
N = max(user_movie.keys()) + 1
# the test set may contain movies that the train set doesn't have
m1 = max(movie_user.keys())
m2 = max([m for (u,m), r in user_movie_ratings_test.items()])
M = max(m1, m2) + 1
print("N:", N, "M:", M)
K = 20 # The number of neighbours we'd like to consider
limit = 5 # Minimum number of movies the users must have in common
neighbors = [] # store the neighbors in a list
averages = [] # each item's average rating
deviations = [] #each item's deviation
for i in range(M):
# find the 25 closest items to item i
try:
users_i = movie_user[i]
users_i_set = set(users_i)
# calculate average and deviation
ratings_i = {user:user_movie_ratings[(user, i)] for user in users_i}
avg_i = np.mean(list(ratings_i.values()))
dev_i = {user: (rating-avg_i) for user, rating in ratings_i.items()}
dev_i_values = np.array(list(dev_i.values()))
# convert all the values in the deviations dictionary into a numpy array.
# This is because the denominator in the Pearson correlation is the square root of the sum of squares
# of the deviations.
sigma_i = np.sqrt(dev_i_values.dot(dev_i_values))
# Save these for later use
averages.append(avg_i)
deviations.append(dev_i)
sl = SortedList()
for j in range(M):
# Don't calculate the correlation with yourself
if j !=i:
users_j = movie_user[j]
users_j_set = set(users_j)
common_users = (users_i_set & users_j_set) # intersetion
if len(common_users) > limit:
# calculate average and deviation
ratings_j = {user:user_movie_ratings[(user, j)] for user in users_j}
avg_j = np.mean(list(ratings_j.values()))
dev_j = {user: (rating-avg_j) for user, rating in ratings_j.items()}
dev_j_values = np.array(list(dev_j.values()))
sigma_j = np.sqrt(dev_j_values.dot(dev_j_values))
# calculate the correlation coefficient
numerator = sum(dev_i[m] * dev_j[m] for m in common_users)
w_ij = numerator/(sigma_i * sigma_j)
# insert into a sorted list and truncate
# negate the weight, because the list is sorted in ascending
# maximum values (1) is "closest"
sl.add((-w_ij, j))
if len(sl) > K:
del sl[-1]
# store the neighbors
neighbors.append(sl)
# print out useful things
# if i%i == 0:
# print(i)
except Exception as e:
pass
# +
# using neighbors to calculate train and test MSE
def predict(i, u): # i: item, u: user
# calculate the weighted sum of deviation
numerator = 0
denominator = 0
for neg_w, j in neighbors[i]:
# remember, the weight is sorted as its negative
# so, the negative of the negative weight is positive
try:
numerator += -neg_w * deviations[j][u]
denominator += abs(neg_w)
except KeyError:
# neighbors may not have rated the same movie
# don't want to do dictionary lookup twice
# so, just throw an exception
pass
if denominator == 0:
prediction = averages[i]
else:
prediction = numerator / denominator * averages[i]
prediction = min(5, prediction)
prediction = max(0.5, prediction) # max rating is 0.5
return prediction
train_predictions = []
train_targets = []
for (u, m), target in user_movie_ratings.items():
# calculate the predictions of the movie
try:
prediction =predict(m, u)
# save the prediction and target
train_predictions.append(prediction)
train_targets.append(target)
except:
pass
test_predictions = []
test_targets = []
for (u, m), target in user_movie_ratings_test.items():
# calculate the predictions of the movie
try:
prediction =predict(m, u)
# save the prediction and target
test_predictions.append(prediction)
test_targets.append(target)
except:
pass
# calculate accuracy
def mse(p, t):
p = np.array(p)
t = np.array(t)
return np.mean((p - t) ** 2)
print("Train mse", mse(train_predictions, train_targets))
print("Test mse", mse(test_predictions, test_targets))
| notebooks/Movie Recommender system.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="adRUGKw0LaFq"
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="jikF7Pp7LqBo" outputId="806fd148-d066-4abf-ffed-275faa1f93d6"
df=pd.read_csv("Consumo_cerveja.csv")
df.head()
# + colab={"base_uri": "https://localhost:8080/"} id="TFmOUfRlL1Ub" outputId="87c74484-af3e-4a6f-f3c3-19e770b02146"
df.info()
# + id="WgVOLXoSRSuE"
df=df.loc[:364,:]
# + colab={"base_uri": "https://localhost:8080/"} id="X6ebKY5kMpxj" outputId="7beae420-7acc-467c-dd49-31ee5ceda02e"
for i in range(0,365):
df['Temperatura Maxima (C)'][i]=df['Temperatura Maxima (C)'][i].replace(',','.')
df['Temperatura Media (C)'][i]=df['Temperatura Media (C)'][i].replace(',','.')
df['Temperatura Minima (C)'][i]=df['Temperatura Minima (C)'][i].replace(',','.')
df['Precipitacao (mm)'][i]=df['Precipitacao (mm)'][i].replace(',','.')
# + colab={"base_uri": "https://localhost:8080/", "height": 310} id="2hFTMSfmNEbp" outputId="4868d11c-f430-4fc4-d164-da05f6e4b5be"
df.tail()
# + id="exq04x-iNa7x"
df['Temperatura Maxima (C)']=df['Temperatura Maxima (C)'].astype('float64')
df['Temperatura Media (C)']=df['Temperatura Media (C)'].astype('float64')
df['Temperatura Minima (C)']=df['Temperatura Minima (C)'].astype('float64')
df['Precipitacao (mm)']=df['Precipitacao (mm)'].astype('float64')
# + colab={"base_uri": "https://localhost:8080/"} id="dj2VJOS_ONFM" outputId="9a16cfdc-5e8e-42de-9dc5-1a7fa59e4a6f"
df.info()
# + id="j5-fCIk6QQd5"
X=df[['Temperatura Media (C)','Temperatura Minima (C)','Temperatura Maxima (C)','Final de Semana','Precipitacao (mm)']]
y=df[['Consumo de cerveja (litros)']]
# + id="mSnJ9vGkOOkc"
from sklearn.model_selection import train_test_split
X_train,X_test,Y_train,Y_test = train_test_split(X,y,test_size=0.2,random_state=42)
# + id="qISmXueGTij3"
from sklearn.preprocessing import StandardScaler
s_scaler = StandardScaler()
X_train = s_scaler.fit_transform(X_train.astype(np.float64))
X_test = s_scaler.transform(X_test.astype(np.float64))
# + colab={"base_uri": "https://localhost:8080/"} id="Fd6WsX4-Qy47" outputId="b58235b5-358e-41e1-d344-b47d743fc7cb"
from sklearn.linear_model import LinearRegression
regr = LinearRegression()
regr.fit(X_train,Y_train)
# + id="0tOjMogLRNhI"
Y_test_pred=regr.predict(X_test)
Y_train_pred=regr.predict(X_train)
# + colab={"base_uri": "https://localhost:8080/"} id="AMMQ91LXR2Nf" outputId="66a8b0a5-f5a5-42e1-e92e-81df33beaea4"
from sklearn.metrics import mean_squared_error, r2_score
print('Mean squared error: %.2f'% mean_squared_error(Y_test, Y_test_pred))
# The coefficient of determination: 1 is perfect prediction
print('Coefficient of determination: %.2f'% r2_score(Y_test, Y_test_pred))
# + colab={"base_uri": "https://localhost:8080/"} id="jRRXjE_ESDc1" outputId="05096738-aa6f-49ad-bbd4-b8b13316ba0c"
print('Mean squared error: %.2f'% mean_squared_error(Y_train, Y_train_pred))
# The coefficient of determination: 1 is perfect prediction
print('Coefficient of determination: %.2f'% r2_score(Y_train, Y_train_pred))
# + id="NQzKJVyATCdU"
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Activation
from tensorflow.keras.optimizers import Adam
model = Sequential()
model.add(Dense(6,activation='relu'))
model.add(Dense(6,activation='relu'))
model.add(Dense(6,activation='relu'))
model.add(Dense(6,activation='relu'))
model.add(Dense(6,activation='relu'))
model.add(Dense(1))
model.compile(optimizer='Adam',loss='mean_squared_error')
# + colab={"base_uri": "https://localhost:8080/"} id="j_DfH--cUCrw" outputId="678bbebe-f58a-4b33-dc1e-b85ef579ec99"
model.fit(x=X_train,y=Y_train,validation_data=(X_test,Y_test),batch_size=32,epochs=100)
model.summary()
# + id="xM1h9BrnUpFa"
y_pred = model.predict(X_test)
# + colab={"base_uri": "https://localhost:8080/"} id="T5e7fb50VVlr" outputId="ff497230-9786-4b77-994d-a0f09a648bd3"
from sklearn import metrics
print('MAE:', metrics.mean_absolute_error(Y_test, y_pred))
print('MSE:', metrics.mean_squared_error(Y_test, y_pred))
print('RMSE:', np.sqrt(metrics.mean_squared_error(Y_test, y_pred)))
print('VarScore:',metrics.explained_variance_score(Y_test,y_pred))
# + colab={"base_uri": "https://localhost:8080/"} id="9ah3yxaOXFu6" outputId="8e238352-71f1-44e7-9038-acad8932366c"
X_test.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 337} id="kV9Jld-ciyP7" outputId="eb1a3577-f803-4f03-bbaa-5a7d77bab238"
fig = plt.figure(figsize=(10,5))
plt.scatter(Y_test,y_pred)
# + id="e9RFyph1lt_y"
| Beer_Consumption_Sao_Paulo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Trolling NIST (v1.3)
# +
# import
import mechanize
import yaml
from pypit import pyputils
import pypit
msgs = pyputils.get_dummy_logger()
from pypit import ararclines as alines
from pypit import arutils as arut
from pypit import arwave as arwv
# -
# ## Dummy Self
slf = arut.dummy_self()
# ## Testing
# init
browser = mechanize.Browser(factory=mechanize.RobustFactory())
browser.set_handle_robots(False)
page = browser.open("http://physics.nist.gov/PhysRefData/ASD/lines_form.html")
browser.select_form(name="upp_wn")
# ### Must be using Java Script...
# ## Pulling by hand...
#
# e.g. Zn I 3000. 10000.
# ### Parse
reload(alines)
tbl = alines.parse_nist(slf,'ZnI')
tbl
# ### CdI
reload(alines)
cdI = alines.parse_nist(None,'CdI')
cdI
# ### HgI
reload(alines)
HgI = alines.parse_nist(None,'HgI')
HgI[HgI['RelInt']>800]
# ### HeI
reload(alines)
HeI = alines.parse_nist(None,'HeI')
HeI
# ### NeI
reload(alines)
NeI = alines.parse_nist(None,'NeI')
NeI[NeI['Aki']>0.]
# ### ArI
ArI = alines.parse_nist(None,'ArI')
ArI
# ## KrI
#
# * Retrieve data (Kr I, 4000., 12000. Ang, Format = ASCII
# * Comment out the ---- lines
# * Add comment at start
reload(alines)
KrI = alines.parse_nist(None,'KrI')
KrI
# ## XeI
#
# * Retrieve data (Xe I, 4000., 12000. Ang, Format = ASCII, min intensity=1000)
# * Comment out the ---- lines
# * Add comment at start
# ## CuI
reload(alines)
CuI = alines.parse_nist(None,'CuI')
CuI
# ## Rejecting lines
# Read yaml
with open('rejected_lines.yaml', 'r') as infile:
rej_lines = yaml.load(infile)
rej_lines
rej_lines['CdI'][3252.524].keys()#['lris_blue'])
# ## Load a full line list
reload(alines)
#alist = alines.load_arcline_list(None,['ZnI','CdI','HgI'])
alist = alines.load_arcline_list(None,None,['CdI','ArI','NeI','HgI','KrI','XeI'],None)
len(alist)
alist[(alist['Ion']=='ArI') & (alist['wave'] > 7700)]
alist[alist['Ion']=='HeI']
alist[alist['Ion']=='KrI']
alist[alist['Ion']=='XeI']
# ## ISIS (CuAr, CuNe)
# +
from pypit import pyputils
msgs = pyputils.get_dummy_logger()
from pypit import ararclines as alines
from pypit import arwave as arwv
alist = alines.load_arcline_list(None,None,['CuI','ArI','NeI'],None,
modify_parse_dict=dict(NeI={'min_wave': 3000.},ArI={'min_intensity': 399.}))
# -
NeI = alist['Ion'] == 'NeI'
np.min(alist[NeI]['wave'])
alist[NeI]
isis = Table.read('ISIS_CuNeCuAr.lst', format='ascii')
isis
isis_vac = arwv.airtovac(isis['wave']*u.AA)
pyp_vac = alist['wave']*u.AA
xdb.xpcol(isis['wave'],isis_vac)
for ivac in isis_vac:
mdiff = np.min(np.abs(ivac-pyp_vac))
if mdiff > 0.1*u.AA:
print("No match for {:g}".format(ivac))
d1 = dict(color='red', dum='1')
d2 = dict(color='blue')
d1.update(d2)
d1
| pypeit/data/arc_lines/NIST_lines.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Feed, get, update and delete Vespa data with pyvespa
#
# - toc: true
# - badges: true
# - comments: true
# - categories: [search, pyvespa, Vespa]
# ## Connect to Vespa instance
# Connect to a running Vespa instance:
app = Vespa(url = "http://localhost", port = 8080)
# Assume the Vespa instance has a Schema called `msmarco` with the following fields:
document = Document(
fields=[
Field(name="id", type="string", indexing=["attribute", "summary"]),
Field(
name="title",
type="string",
indexing=["index", "summary"],
index="enable-bm25",
),
Field(
name="body",
type="string",
indexing=["index", "summary"],
index="enable-bm25",
),
]
)
# ## Data operations
# ### Feed data
response = app.feed_data_point(
schema="msmarco",
data_id="1",
fields={
"id": "1",
"title": "this is my first title",
"body": "this is my first body",
},
)
assert response.json()["id"] == "id:msmarco:msmarco::1"
# ### Get data
# +
response = app.get_data(schema="msmarco", data_id="1")
expected_data = {
"fields": {
"id": "1",
"title": "this is my first title",
"body": "this is my first body",
},
"id": "id:msmarco:msmarco::1",
"pathId": "/document/v1/msmarco/msmarco/docid/1"
}
assert response.status_code == 200
assert response.json() == expected_data
# -
# ### Update data
# +
response = app.update_data(
schema="msmarco", data_id="1", fields={"title": "this is my updated title"}
)
assert response.json()["id"] == "id:msmarco:msmarco::1"
# -
# ### Delete data
# +
response = app.delete_data(schema="msmarco", data_id="1")
assert response.json()["id"] == "id:msmarco:msmarco::1"
| _notebooks/2020-11-06-pyvespa-include-data-operations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import json
import os
import pandas
import tqdm
os.getcwd()
# +
## 要補註解
# -
path = "C:\\Users\\user\\Desktop\\Python\\projectNevus\\data\\description\\"
N = len(os.listdir(path))
Id = [None] * N
Age = [None] * N
Status = [None] * N
Sex = [None] * N
Diagnosis = [None] * N
Type = [None] * N
for i, file in enumerate(os.listdir(path)):
Id[i] = file
Json = json.loads(open(path + file).read())
if( "meta" in Json.keys() ):
if( "clinical" in Json["meta"].keys() ):
if( "age_approx" in Json["meta"]["clinical"].keys() ):
Age[i] = Json["meta"]["clinical"]["age_approx"]
if( "benign_malignant" in Json["meta"]["clinical"].keys() ):
Status[i] = Json["meta"]["clinical"]["benign_malignant"]
if( "sex" in Json["meta"]["clinical"].keys() ):
Sex[i] = Json["meta"]["clinical"]["sex"]
if( "diagnosis" in Json["meta"]["clinical"].keys() ):
Diagnosis[i] = Json["meta"]["clinical"]["diagnosis"]
if( "acquisition" in Json["meta"].keys() ):
if( "image_type" in Json["meta"]["acquisition"].keys() ):
Type[i] = Json["meta"]["acquisition"]["image_type"]
descriptionFull = pandas.DataFrame({
"Id" : Id,
"Age" : Age,
"Status": Status,
"Sex": Sex,
"Diagnosis": Diagnosis,
"Type": Type
})
descriptionFull.head()
descriptionFull.shape
path = "C:\\Users\\user\\Desktop\\Python\\projectNevus\\data\\temporary\\"
descriptionFull.to_csv(path + "descriptionFull.csv")
| Project/Nevus/component/1-descriptionFull.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#Importing the essential libraries
#Beautiful Soup is a Python library for pulling data out of HTML and XML files
#The Natural Language Toolkit
import requests
import nltk
from nltk.sentiment.vader import SentimentIntensityAnalyzer
from bs4 import BeautifulSoup
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
import random
from wordcloud import WordCloud
from html.parser import HTMLParser
import bs4 as bs
import urllib.request
import re
# +
r=requests.get('https://investorplace.com/2020/06/dont-throw-away-your-waste-management-wm-stock/')
# -
#Setting the correct text encoding of the HTML page
r.encoding = 'utf-8'
#Extracting the HTML from the request object
html = r.text
# Printing the first 500 characters in html
print(html[:500])
# Creating a BeautifulSoup object from the HTML
soup = BeautifulSoup(html)
# Getting the text out of the soup
text = soup.get_text()
#total length
len(text)
text=text[19400:25650]
# Removing Square Brackets and Extra Spaces
clean_text = re.sub(r'\[[0-9]*\]', ' ', text)
clean_text = re.sub(r'\s+', ' ', clean_text)
clean_text[500:900]
# # Text Summarization
# +
#We need to tokenize the article into sentences
#Sentence tokenization
sentence_list = nltk.sent_tokenize(clean_text)
# +
#Weighted Frequency of Occurrence
stopwords = nltk.corpus.stopwords.words('english')
word_frequencies = {}
for word in nltk.word_tokenize(clean_text):
if word not in stopwords:
if word not in word_frequencies.keys():
word_frequencies[word] = 1
else:
word_frequencies[word] += 1
# +
maximum_frequncy = max(word_frequencies.values())
for word in word_frequencies.keys():
word_frequencies[word] = (word_frequencies[word]/maximum_frequncy)
# -
sentence_scores = {}
for sent in sentence_list:
for word in nltk.word_tokenize(sent.lower()):
if word in word_frequencies.keys():
if len(sent.split(' ')) < 30:
if sent not in sentence_scores.keys():
sentence_scores[sent] = word_frequencies[word]
else:
sentence_scores[sent] += word_frequencies[word]
sentence_scores
# # 10 Key Ideas
# +
import heapq
summary_sentences = heapq.nlargest(10, sentence_scores, key=sentence_scores.get)
summary = ' '.join(summary_sentences)
print(summary)
# -
# # 15 Key Ideas
# +
import heapq
summary_sentences_2 = heapq.nlargest(15, sentence_scores, key=sentence_scores.get)
summary_2 = ' '.join(summary_sentences_2)
print(summary_2)
# -
# # Key Takeaways-
#
# 1. While the sector might not have large growth in current Covid-19 times, it is surely promising.
# 2. There will be good growth of WM as there is not much competition.
# 3. Decent scope of growth ahead.
# 4. Waste Management sector growth seems viable.
# 5. WM Stock is must hold, not sell.
| Waste Management Inc/3. Waste Management Key Aspects.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="Ona5p1IgVCrb"
# # ISB-CGC Community Notebooks
#
# Check out more notebooks at our [Community Notebooks Repository](https://github.com/isb-cgc/Community-Notebooks)!
#
# ```
# Title: How to create cohorts
# Author: <NAME>
# Created: 2019-06-20
# Purpose: Basic overview of creating cohorts
# URL: https://github.com/isb-cgc/Community-Notebooks/blob/master/Notebooks/How_to_create_cohorts.ipynb
# Notes: This notebook was adapted from work by <NAME>, 'How to Create TCGA Cohorts part 1' https://github.com/isb-cgc/examples-Python/blob/master/notebooks/Creating%20TCGA%20cohorts%20--%20part%201.ipynb.
# ```
# ***
#
# This notebook will show you how to create a TCGA cohort using the publicly available TCGA BigQuery tables that the [ISB-CGC](http://isb-cgc.org) project has produced based on the open-access [TCGA](http://cancergenome.nih.gov/) data available at the [Data Portal](https://tcga-data.nci.nih.gov/tcga/). You will need to have access to a Google Cloud Platform (GCP) project in order to use BigQuery. If you don't already have one, you can sign up for a [free-trial](https://cloud.google.com/free-trial/). You can also explore the available tables and data sets before commiting to creating a GCP project though the [ISB-CGC BigQuery Table Searcher](isb-cgc.appspot.com/bq_meta_search/).
#
# We are not attempting to provide a thorough BigQuery or IPython tutorial here, as a wealth of such information already exists. Here are some links to some resources that you might find useful:
# * [BigQuery](https://cloud.google.com/bigquery/what-is-bigquery)
# * the BigQuery [web UI](https://console.cloud.google.com/bigquery)
# * where you can run queries interactively
# * [Jupyter Notebooks](http://jupyter.org/)
# * [Google Cloud Datalab](https://cloud.google.com/datalab/)
# * interactive cloud-based platform for analyzing data built on the Jupyter Notebooks
# * [Google Colaboratory](https://colab.research.google.com/)
# * Free Jupyter Notebook environment that runs in your browser
#
# There are also many tutorials and samples available on github (see, in particular, the [datalab](https://github.com/GoogleCloudPlatform/datalab) repo, the [Google Genomics]( https://github.com/googlegenomics) project), and our own [Community Notebooks](https://github.com/isb-cgc/Community-Notebooks).
#
# OK then, let's get started! In order to work with BigQuery, the first thing you need to do is import the bigquery module:
# + colab={} colab_type="code" id="MFLeeww_VCrh"
from google.cloud import bigquery
# + [markdown] colab_type="text" id="64nmsYB1VMta"
# Next we will need to Authorize our access to BigQuery and the Google Cloud. For more information see ['Quick Start Guide to ISB-CGC'](https://nbviewer.jupyter.org/github/isb-cgc/Community-Notebooks/blob/master/Notebooks/Quick_Start_Guide_to_ISB_CGC.ipynb) and alternative authentication methods can be found [here](https://googleapis.github.io/google-cloud-python/latest/core/auth.html).
# + cellView="both" colab={"base_uri": "https://localhost:8080/", "height": 360} colab_type="code" id="QgN9GwOZ2ttz" outputId="89af1b8a-f1a2-43d6-99c6-d6b99044f9c4"
# !gcloud auth application-default login
# + colab={} colab_type="code" id="DNqD9CkHb34J"
# Create a variable for which client to use with BigQuery
project_num = 'your_project_number' # Update with your Google Project number
if project_num == 'your_project_number':
print('Please update the project number with your Google Cloud Project')
else:
client = bigquery.Client('project_num') # Replace your_project_number with your project ID
# + [markdown] colab_type="text" id="I4qZY5-6VCrl"
# The next thing you need to know is how to access the specific tables you are interested in. BigQuery tables are organized into datasets, and datasets are owned by a specific GCP project. The tables we will be working with in this notebook are in a dataset called **`TCGA_bioclin_v0`**, owned by the **`isb-cgc`** project. A full table identifier is of the form `<project_id>.<dataset_id>.<table_id>`. Let's start by getting some basic information about the tables in this dataset:
# + colab={"base_uri": "https://localhost:8080/", "height": 102} colab_type="code" id="oziRRi6HVCrl" outputId="62c27ecd-de3e-44c6-d2fd-c5ebd3bcf047"
# For each table in the dataset print the number of rows,
# number of bytes and the name of the table
print("Tables:")
for t in list(client.list_tables('isb-cgc.TCGA_bioclin_v0')):
print(t.table_id)
# + [markdown] colab_type="text" id="HWeQAgBhVCrq"
# In this tutorial, we are going to look at a few different ways that we can use the information in these tables to create cohorts. Now, you maybe asking what we mean by "cohort" and why you might be interested in *creating* one, or maybe what it even means to "create" a cohort. The TCGA dataset includes clinical, biospecimen, and molecular data from over 10,000 cancer patients who agreed to be a part of this landmark research project to build [The Cancer Genome Atlas](http://cancergenome.nih.gov/). This large dataset was originally organized and studied according to [cancer type](http://cancergenome.nih.gov/cancersselected) but now that this multi-year project is nearing completion, with over 30 types of cancer and over 10,000 tumors analyzed, **you** have the opportunity to look at this dataset from whichever angle most interests you. Maybe you are particularly interested in early-onset cancers, or gastro-intestinal cancers, or a specific type of genetic mutation. This is where the idea of a "cohort" comes in. The original TCGA "cohorts" were based on cancer type (aka "study"), but now you can define a cohort based on virtually any clinical or molecular feature by querying these BigQuery tables. A cohort is simply a list of samples, using the [TCGA barcode](https://docs.gdc.cancer.gov/Encyclopedia/pages/TCGA_Barcode/) system. Once you have created a cohort you can use it in any number of ways: you could further explore the data available for one cohort, or compare one cohort to another, for example.
#
# In the rest of this tutorial, we will create several different cohorts based on different motivating research questions. We hope that these examples will provide you with a starting point from which you can build, to answer your own research questions.
# + [markdown] colab_type="text" id="MzG3wtPPVCrq"
# ## Exploring the Clinical data table
# Let's start by looking at the clinical data table. The TCGA dataset contains a few very basic clinical data elements for almost all patients, and contains additional information for some tumor types only. For example smoking history information is generally available only for lung cancer patients, and BMI (body mass index) is only available for tumor types where that is a known significant risk factor. Let's take a look at the clinical data table and see how many different pieces of information are available to us:
# + [markdown] colab_type="text" id="ORGzgvAp43DQ"
# #### Get Table Schema
# + cellView="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="S9xWqzdDVCrr" outputId="75001ca7-bef4-4c04-83c7-e3c020bfa168"
# Magic command of bigquery with the project id as isb-cgc-02-0001 and create a Pandas Dataframe
# Change isb-cgc-02-0001 to your project ID
# %%bigquery --project isb-cgc-02-0001
SELECT column_name
FROM `isb-cgc.TCGA_bioclin_v0.INFORMATION_SCHEMA.COLUMNS`
WHERE table_name = 'Clinical'
# Syntax of the above query
# SELECT *
# FROM `project_name.dataset_name.INFORMATION_SCHEMA.COLUMNS`
# WHERE table_catalog=project_name and table_schema=dataset_name and table_name=table_name
# + [markdown] colab_type="text" id="i_5Put7BVCru"
# That's a lot of fields! We can also get at the schema programmatically:
# + [markdown] colab_type="text" id="UrwKZynh4-Ul"
# #### Programmatically Get Schema
# + cellView="code" colab={"base_uri": "https://localhost:8080/", "height": 136} colab_type="code" id="9u6WH10zVCru" outputId="e50f6698-1c38-4053-d0ef-f6b50b5a5f69"
# Create a reference for the table
table_ref = "isb-cgc.TCGA_bioclin_v0.Clinical"
# Get the table
table = client.get_table(table_ref)
# Create a list of the field names
fieldNames = list(map(lambda tsf: tsf.name, table.schema))
#Create a list of the field types
fieldTypes = list(map(lambda tsf: tsf.field_type, table.schema))
# Print the number of fields and the first 5 fields
print("This table has {} fields. ".format(len(fieldNames)))
print("The first few field names and types are: ")
for i in range(5):
print("{} - {}".format(fieldNames[i], fieldTypes[i]))
# + [markdown] colab_type="text" id="Q9yuPEMnVCrx"
# Let's look at these fields and see which ones might be the most "interesting", by looking at how many times they are filled-in (not NULL), or how much variation exists in the values. If we wanted to look at just a single field, "tobacco_smoking_history" for example, we could use a very simple query to get a basic summary:
# + colab={"base_uri": "https://localhost:8080/", "height": 235} colab_type="code" id="Gvfk6WN2VCrx" outputId="9f561f28-3cbe-40b9-b50c-694d49a21f66"
# %%bigquery --project isb-cgc-02-0001
SELECT tobacco_smoking_history,
COUNT(*) AS n
FROM `isb-cgc.TCGA_bioclin_v0.Clinical`
GROUP BY tobacco_smoking_history
ORDER BY n DESC
# + [markdown] colab_type="text" id="WW68khu1VCr0"
# But if we want to loop over *all* fields and get a sense of which fields might provide us with useful criteria for specifying a cohort, we'll want to automate that. We'll put a threshold on the minimum number of patients that we expect information for, and the maximum number of unique values (since fields such as the "ParticipantBarcode" will be unique for every patient and, although we will need that field later, it's probably not useful for defining a cohort).
# + [markdown] colab_type="text" id="q4tK5sIO5DZy"
# #### Find Interesting Fields
# + cellView="code" colab={"base_uri": "https://localhost:8080/", "height": 309} colab_type="code" id="ThNJemi6VCr1" outputId="91a03da8-7530-4966-dadf-b04cd08b559e"
# Get the number of Tables
numPatients = table.num_rows
# Print the total number of patients
print(" The {} table describes a total of {} patients. ".format(table.table_id, numPatients))
# let's set a threshold for the minimum number of values that a field should have,
# the maximum number of unique values, and either the highest cancer type or
# the mean and sigma of the row.
minNumPatients = int(numPatients*0.80)
maxNumValues = 50
# Create a variable to be filled in by the for loop with the number
# interesting features
numInteresting = 0
# Create a list to hold the results from the loop below
iList = []
# Loop over the fields and find the number of values with the number of unique
# values and the
for iField in range(len(fieldNames)):
aField = fieldNames[iField]
aType = fieldTypes[iField]
qString = "SELECT {} FROM `isb-cgc.TCGA_bioclin_v0.Clinical`".format(aField)
df = client.query(qString).result().to_dataframe()
summary = df[str(aField)].describe()
if ( aType == "STRING" ):
topFrac = float(summary['freq'])/float(summary['count'])
if ( summary['count'] >= minNumPatients ):
if ( summary['unique'] <= maxNumValues and summary['unique'] > 1 ):
if ( topFrac < 0.90 ):
numInteresting += 1
iList += [aField]
print(" > {} has {} values with {} unique ({} occurs {} times)".format(aField, summary['count'], summary['unique'], summary['top'], round(summary['freq'],2)))
else:
if ( summary['count'] >= minNumPatients ):
if ( summary['std'] > 0.1 ):
numInteresting += 1
iList += [aField]
print(" > {} has {} values (mean={}, sigma={}) ".format(aField, summary['count'], round(summary['mean'], 2), round(summary['std'], 2)))
print(" ")
print(" Found {} potentially interesting features: ".format(numInteresting))
print(" ", iList)
# + [markdown] colab_type="text" id="vsDPK0wbVCr4"
# The above helps us narrow down on which fields are likely to be the most useful, but if you have a specific interest, for example in menopause or HPV status, you can still look at those in more detail very easily:
# + colab={"base_uri": "https://localhost:8080/", "height": 173} colab_type="code" id="ZxEuBUA_VCr5" outputId="5db653d6-1957-46b5-d48a-9bd3cf85d8bf"
# %%bigquery --project isb-cgc-02-0001
SELECT menopause_status, COUNT(*) AS n
FROM `isb-cgc.TCGA_bioclin_v0.Clinical`
WHERE menopause_status IS NOT NULL
GROUP BY menopause_status
ORDER BY n DESC
# + [markdown] colab_type="text" id="E3gRdYwaVCr8"
# We might wonder which specific tumor types have menopause information:
# + colab={"base_uri": "https://localhost:8080/", "height": 173} colab_type="code" id="2fO9uxVzVCr9" outputId="c11640d2-dfcd-4afd-f2ae-f18e65e99e91"
# %%bigquery --project isb-cgc-02-0001
SELECT project_short_name, COUNT(*) AS n
FROM `isb-cgc.TCGA_bioclin_v0.Clinical`
WHERE menopause_status IS NOT NULL
GROUP BY project_short_name
ORDER BY n DESC
# + colab={"base_uri": "https://localhost:8080/", "height": 204} colab_type="code" id="L9QCMBJLVCsA" outputId="775d9b91-8d2c-4cdf-cb33-e0efa35c2619"
# %%bigquery --project isb-cgc-02-0001
SELECT hpv_status, hpv_calls, COUNT(*) AS n
FROM `isb-cgc.TCGA_bioclin_v0.Clinical`
WHERE hpv_status IS NOT NULL
GROUP BY hpv_status, hpv_calls
HAVING n > 20
ORDER BY n DESC
# + [markdown] colab_type="text" id="StOsLNYmVCsC"
# ## TCGA Annotations
#
# An additional factor to consider, when creating a cohort is that there may be additional information that might lead one to exclude a particular patient from a cohort. In certain instances, patients have been redacted or excluded from analyses for reasons such as prior treatment, etc, but since different researchers may have different criteria for using or excluding certain patients or certain samples from their analyses, an overview of the annoations can be found [here](https://docs.gdc.cancer.gov/Encyclopedia/pages/Annotations_TCGA/). These annotations have also been uploaded into a BigQuery table and can be used in conjuction with the other BigQuery tables.
# + [markdown] colab_type="text" id="6a9snKq1VCsD"
# # Create a Cohort from Two Tables
#
# Now that we have a better idea of what types of information is available in the Clinical data table, let's create a cohort consisting of female breast-cancer patients, diagnosed at the age of 50 or younger.
# + [markdown] colab_type="text" id="gWdNmzevVCsD"
# In this next code cell, we define several queries with a **`WITH`** clause which allows us to use them in a final query. We will then save the query to a [Pandas DataFrame](https://pandas.pydata.org/) to allow it to be analyzed later with a named data frame.
# * the first query, called **`select_on_annotations`**, finds all patients in the Annotations table which have either been 'redacted' or had 'unacceptable prior treatment';
# * the second query, **`select_on_clinical`** selects all female breast-cancer patients who were diagnosed at age 50 or younger, while also pulling out a few additional fields that might be of interest; and
# * the final query joins these two together and returns just those patients that meet the clinical-criteria and do **not** meet the exclusion-criteria.
# + [markdown] colab_type="text" id="VMCUMONH5KWk"
# #### Create a Query for a Cohort from Two Tables
# + cellView="code" colab={} colab_type="code" id="FpzdD3laPrE-"
# First use the BigQuery Magic Command (%%bigquery) then name the dataframe
# (early_onset_breast_canver), and finallly include your project ID
# (--project project ID).
# %%bigquery early_onset_breast_cancer --project isb-cgc-02-0001
WITH
select_on_annotations AS (
SELECT
case_barcode,
category AS categoryName,
classification AS classificationName
FROM
`isb-cgc.TCGA_bioclin_v0.Annotations`
WHERE
( entity_type="Patient"
AND (category="History of unacceptable prior treatment related to a prior/other malignancy"
OR classification="Redaction" ) )
GROUP BY
case_barcode,
categoryName,
classificationName
),
--
select_on_clinical AS (
SELECT
case_barcode,
vital_status,
days_to_last_known_alive,
ethnicity,
histological_type,
menopause_status,
race
FROM
`isb-cgc.TCGA_bioclin_v0.Clinical`
WHERE
( disease_code = "BRCA"
AND age_at_diagnosis<=50
AND gender="FEMALE" )
)
--
SELECT
case_barcode
FROM (
SELECT
a.categoryName,
a.classificationName,
c.case_barcode
FROM select_on_annotations AS a
FULL JOIN select_on_clinical AS c
ON
a.case_barcode = c.case_barcode
WHERE
a.case_barcode IS NOT NULL
OR c.case_barcode IS NOT NULL
ORDER BY
a.classificationName,
a.categoryName,
c.case_barcode
)
WHERE
categoryName IS NULL
AND classificationName IS NULL
AND case_barcode IS NOT NULL
ORDER BY
case_barcode
# + colab={"base_uri": "https://localhost:8080/", "height": 204} colab_type="code" id="ymX0w7RwkFAU" outputId="09746f72-732b-4659-a269-8686bf6aed96"
early_onset_breast_cancer.head()
# + [markdown] colab_type="text" id="8YZWApgkVCsT"
# ### Useful Tricks
# Before we leave off, here are a few useful tricks for working with BigQuery:
# * If you want to see how much data and which tables are going to be touched by this data, you can use the "dry run" option.
# * You can then build a query as a variable and put the results into a dataframe instead of
# + colab={} colab_type="code" id="CEIn37yqVCsc"
# Create a variable with the query as a string
breast_cancer_query = """
WITH
select_on_annotations AS (
SELECT
case_barcode,
category AS categoryName,
classification AS classificationName
FROM
`isb-cgc.TCGA_bioclin_v0.Annotations`
WHERE
( entity_type="Patient"
AND (category="History of unacceptable prior treatment related to a prior/other malignancy"
OR classification="Redaction" ) )
GROUP BY
case_barcode,
categoryName,
classificationName
),
--
select_on_clinical AS (
SELECT
case_barcode,
vital_status,
days_to_last_known_alive,
ethnicity,
histological_type,
menopause_status,
race
FROM
`isb-cgc.TCGA_bioclin_v0.Clinical`
WHERE
( disease_code = "BRCA"
AND age_at_diagnosis<=50
AND gender="FEMALE" )
)
--
SELECT
case_barcode
FROM (
SELECT
a.categoryName,
a.classificationName,
c.case_barcode
FROM select_on_annotations AS a
FULL JOIN select_on_clinical AS c
ON
a.case_barcode = c.case_barcode
WHERE
a.case_barcode IS NOT NULL
OR c.case_barcode IS NOT NULL
ORDER BY
a.classificationName,
a.categoryName,
c.case_barcode
)
WHERE
categoryName IS NULL
AND classificationName IS NULL
AND case_barcode IS NOT NULL
ORDER BY
case_barcode
"""
# + [markdown] colab_type="text" id="jzOshUb8SbxT"
# Since this is a large query, we might want to check the number of bytes that will be processed. We can use a dry run to see how many bytes wil be processed with the query without actually doing a query.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="GMg01x8M06yF" outputId="208cbd7c-9caf-451c-ede3-8e8336789dee"
job_config = bigquery.QueryJobConfig()
job_config.dry_run = True
job_config.use_query_cache = False
query_job = client.query(
(breast_cancer_query),
# Location must match that of the dataset(s) referenced in the query.
location="US",
job_config=job_config,
) # API request
# A dry run query completes immediately.
assert query_job.state == "DONE"
assert query_job.dry_run
print("This query will process {} Kilobytes.".format(query_job.total_bytes_processed*0.001))
# + [markdown] colab_type="text" id="5SJIh9QTawBa"
# For more information on price estimation, please see the [Estimating storage and query costs](https://cloud.google.com/bigquery/docs/estimate-costs) and [BigQuery best practices: Controlling costs](https://cloud.google.com/bigquery/docs/best-practices-costs) pages.
#
# Below we can then use the same variable with the query to then run the query and put the result into a Pandas Dataframe for later analysis.
# + colab={"base_uri": "https://localhost:8080/", "height": 204} colab_type="code" id="J4angHKtWrce" outputId="c6486d04-1f47-4096-f498-1d29f6448431"
query = client.query(breast_cancer_query)
breast_cancer_query_results = query.result().to_dataframe()
breast_cancer_query_results.head(5)
| Notebooks/How_to_create_cohorts.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] pycharm={"name": "#%% md\n"}
# # Install
#
# Get dataset ircad
# ```
# python -m io3d -l 3Dircadb1
# ```
#
# +
import torch
from torch import nn
from torch.utils.data import DataLoader
from torchvision import datasets
from torchvision.transforms import ToTensor, Lambda, Compose
import torch.utils.data as tdata
import io3d
import torchvision
from matplotlib import pyplot as plt
batch_size = 4
# + pycharm={"name": "#%%\n"}
class Io3dDataset(tdata.Dataset):
"""
"""
def __init__(self, dataset_label="3Dircadb1", organ_label="rightkidney", start_id=1, end_id=20, transform=None):
"""
"""
self.dataset_label = dataset_label
self.organ_label = organ_label
self.start_id = start_id
self.end_id = end_id
self._i = start_id
self.transform = transform
pass
# def _get_lengths(self):
# for i in range(self.start_id, self.end_id):
# datap1 = io3d.datasets.read_dataset(self.dataset_label, "data3d", i)
# out = datap1, None
# if self.transform: # jatra.shape = [512,512,1]
# out = self.transform(out)
def __len__(self):
return self.end_id - self.start_id
def __getitem__(self, idx):
if torch.is_tensor(idx):
idx = idx.tolist()
datap1 = io3d.datasets.read_dataset(self.dataset_label, "data3d", idx + self.start_id)
datap2 = io3d.datasets.read_dataset(self.dataset_label, self.organ_label, idx + self.start_id)
out = datap1, datap2
if self.transform: # jatra.shape = [512,512,1]
out = self.transform(out)
return out
class ExtractTensor(torch.nn.Module):
def __init__(self, debugdir=None):
super().__init__()
# if debugdir:
# debugdir = Path(debugdir)
# self.debugdir = debugdir
# if self.debugdir:
# self.debugdir.mkdir(parents=True, exist_ok=True)
# self._set_images = set()
def forward(self, couple_datap):
datap1, datap2 = couple_datap
data3d1 = torch.from_numpy(datap1.data3d)
data3d2 = torch.from_numpy(datap2.data3d)
return data3d1, data3d2
transform = torch.nn.Sequential(
ExtractTensor()
)
train_data = Io3dDataset(start_id=1, end_id=16, transform=transform)
test_data = Io3dDataset(start_id=16, end_id=20, transform=transform)
train_dataloader = DataLoader(train_data, batch_size=batch_size)
test_dataloader = DataLoader(test_data, batch_size=batch_size)
# + pycharm={"name": "#%%\n"}
# next(train_dataloader)
x, y = train_data[0]
image = x
mask = y
print(image.shape)
print(mask.shape)
plt.imshow(image[100,:,:].numpy(), cmap='gray')
plt.figure()
plt.imshow(mask[100,:,:].numpy(), cmap='gray')
# + [markdown] pycharm={"name": "#%% md\n"}
# # U-Net
# + pycharm={"name": "#%%\n"}
from torch import nn
import torch
@torch.jit.script
def autocrop(encoder_layer: torch.Tensor, decoder_layer: torch.Tensor):
"""
Center-crops the encoder_layer to the size of the decoder_layer,
so that merging (concatenation) between levels/blocks is possible.
This is only necessary for input sizes != 2**n for 'same' padding and always required for 'valid' padding.
"""
if encoder_layer.shape[2:] != decoder_layer.shape[2:]:
ds = encoder_layer.shape[2:]
es = decoder_layer.shape[2:]
assert ds[0] >= es[0]
assert ds[1] >= es[1]
if encoder_layer.dim() == 4: # 2D
encoder_layer = encoder_layer[
:,
:,
((ds[0] - es[0]) // 2):((ds[0] + es[0]) // 2),
((ds[1] - es[1]) // 2):((ds[1] + es[1]) // 2)
]
elif encoder_layer.dim() == 5: # 3D
assert ds[2] >= es[2]
encoder_layer = encoder_layer[
:,
:,
((ds[0] - es[0]) // 2):((ds[0] + es[0]) // 2),
((ds[1] - es[1]) // 2):((ds[1] + es[1]) // 2),
((ds[2] - es[2]) // 2):((ds[2] + es[2]) // 2),
]
return encoder_layer, decoder_layer
def conv_layer(dim: int):
if dim == 3:
return nn.Conv3d
elif dim == 2:
return nn.Conv2d
def get_conv_layer(in_channels: int,
out_channels: int,
kernel_size: int = 3,
stride: int = 1,
padding: int = 1,
bias: bool = True,
dim: int = 2):
return conv_layer(dim)(in_channels, out_channels, kernel_size=kernel_size, stride=stride, padding=padding,
bias=bias)
def conv_transpose_layer(dim: int):
if dim == 3:
return nn.ConvTranspose3d
elif dim == 2:
return nn.ConvTranspose2d
def get_up_layer(in_channels: int,
out_channels: int,
kernel_size: int = 2,
stride: int = 2,
dim: int = 3,
up_mode: str = 'transposed',
):
if up_mode == 'transposed':
return conv_transpose_layer(dim)(in_channels, out_channels, kernel_size=kernel_size, stride=stride)
else:
return nn.Upsample(scale_factor=2.0, mode=up_mode)
def maxpool_layer(dim: int):
if dim == 3:
return nn.MaxPool3d
elif dim == 2:
return nn.MaxPool2d
def get_maxpool_layer(kernel_size: int = 2,
stride: int = 2,
padding: int = 0,
dim: int = 2):
return maxpool_layer(dim=dim)(kernel_size=kernel_size, stride=stride, padding=padding)
def get_activation(activation: str):
if activation == 'relu':
return nn.ReLU()
elif activation == 'leaky':
return nn.LeakyReLU(negative_slope=0.1)
elif activation == 'elu':
return nn.ELU()
def get_normalization(normalization: str,
num_channels: int,
dim: int):
if normalization == 'batch':
if dim == 3:
return nn.BatchNorm3d(num_channels)
elif dim == 2:
return nn.BatchNorm2d(num_channels)
elif normalization == 'instance':
if dim == 3:
return nn.InstanceNorm3d(num_channels)
elif dim == 2:
return nn.InstanceNorm2d(num_channels)
elif 'group' in normalization:
num_groups = int(normalization.partition('group')[-1]) # get the group size from string
return nn.GroupNorm(num_groups=num_groups, num_channels=num_channels)
class Concatenate(nn.Module):
def __init__(self):
super(Concatenate, self).__init__()
def forward(self, layer_1, layer_2):
x = torch.cat((layer_1, layer_2), 1)
return x
class DownBlock(nn.Module):
"""
A helper Module that performs 2 Convolutions and 1 MaxPool.
An activation follows each convolution.
A normalization layer follows each convolution.
"""
def __init__(self,
in_channels: int,
out_channels: int,
pooling: bool = True,
activation: str = 'relu',
normalization: str = None,
dim: str = 2,
conv_mode: str = 'same'):
super().__init__()
self.in_channels = in_channels
self.out_channels = out_channels
self.pooling = pooling
self.normalization = normalization
if conv_mode == 'same':
self.padding = 1
elif conv_mode == 'valid':
self.padding = 0
self.dim = dim
self.activation = activation
# conv layers
self.conv1 = get_conv_layer(self.in_channels, self.out_channels, kernel_size=3, stride=1, padding=self.padding,
bias=True, dim=self.dim)
self.conv2 = get_conv_layer(self.out_channels, self.out_channels, kernel_size=3, stride=1, padding=self.padding,
bias=True, dim=self.dim)
# pooling layer
if self.pooling:
self.pool = get_maxpool_layer(kernel_size=2, stride=2, padding=0, dim=self.dim)
# activation layers
self.act1 = get_activation(self.activation)
self.act2 = get_activation(self.activation)
# normalization layers
if self.normalization:
self.norm1 = get_normalization(normalization=self.normalization, num_channels=self.out_channels,
dim=self.dim)
self.norm2 = get_normalization(normalization=self.normalization, num_channels=self.out_channels,
dim=self.dim)
def forward(self, x):
y = self.conv1(x) # convolution 1
y = self.act1(y) # activation 1
if self.normalization:
y = self.norm1(y) # normalization 1
y = self.conv2(y) # convolution 2
y = self.act2(y) # activation 2
if self.normalization:
y = self.norm2(y) # normalization 2
before_pooling = y # save the outputs before the pooling operation
if self.pooling:
y = self.pool(y) # pooling
return y, before_pooling
class UpBlock(nn.Module):
"""
A helper Module that performs 2 Convolutions and 1 UpConvolution/Upsample.
An activation follows each convolution.
A normalization layer follows each convolution.
"""
def __init__(self,
in_channels: int,
out_channels: int,
activation: str = 'relu',
normalization: str = None,
dim: int = 3,
conv_mode: str = 'same',
up_mode: str = 'transposed'
):
super().__init__()
self.in_channels = in_channels
self.out_channels = out_channels
self.normalization = normalization
if conv_mode == 'same':
self.padding = 1
elif conv_mode == 'valid':
self.padding = 0
self.dim = dim
self.activation = activation
self.up_mode = up_mode
# upconvolution/upsample layer
self.up = get_up_layer(self.in_channels, self.out_channels, kernel_size=2, stride=2, dim=self.dim,
up_mode=self.up_mode)
# conv layers
self.conv0 = get_conv_layer(self.in_channels, self.out_channels, kernel_size=1, stride=1, padding=0,
bias=True, dim=self.dim)
self.conv1 = get_conv_layer(2 * self.out_channels, self.out_channels, kernel_size=3, stride=1,
padding=self.padding,
bias=True, dim=self.dim)
self.conv2 = get_conv_layer(self.out_channels, self.out_channels, kernel_size=3, stride=1, padding=self.padding,
bias=True, dim=self.dim)
# activation layers
self.act0 = get_activation(self.activation)
self.act1 = get_activation(self.activation)
self.act2 = get_activation(self.activation)
# normalization layers
if self.normalization:
self.norm0 = get_normalization(normalization=self.normalization, num_channels=self.out_channels,
dim=self.dim)
self.norm1 = get_normalization(normalization=self.normalization, num_channels=self.out_channels,
dim=self.dim)
self.norm2 = get_normalization(normalization=self.normalization, num_channels=self.out_channels,
dim=self.dim)
# concatenate layer
self.concat = Concatenate()
def forward(self, encoder_layer, decoder_layer):
""" Forward pass
Arguments:
encoder_layer: Tensor from the encoder pathway
decoder_layer: Tensor from the decoder pathway (to be up'd)
"""
up_layer = self.up(decoder_layer) # up-convolution/up-sampling
cropped_encoder_layer, dec_layer = autocrop(encoder_layer, up_layer) # cropping
if self.up_mode != 'transposed':
# We need to reduce the channel dimension with a conv layer
up_layer = self.conv0(up_layer) # convolution 0
up_layer = self.act0(up_layer) # activation 0
if self.normalization:
up_layer = self.norm0(up_layer) # normalization 0
merged_layer = self.concat(up_layer, cropped_encoder_layer) # concatenation
y = self.conv1(merged_layer) # convolution 1
y = self.act1(y) # activation 1
if self.normalization:
y = self.norm1(y) # normalization 1
y = self.conv2(y) # convolution 2
y = self.act2(y) # acivation 2
if self.normalization:
y = self.norm2(y) # normalization 2
return y
class UNet(nn.Module):
def __init__(self,
in_channels: int = 1,
out_channels: int = 2,
n_blocks: int = 4,
start_filters: int = 32,
activation: str = 'relu',
normalization: str = 'batch',
conv_mode: str = 'same',
dim: int = 2,
up_mode: str = 'transposed'
):
super().__init__()
self.in_channels = in_channels
self.out_channels = out_channels
self.n_blocks = n_blocks
self.start_filters = start_filters
self.activation = activation
self.normalization = normalization
self.conv_mode = conv_mode
self.dim = dim
self.up_mode = up_mode
self.down_blocks = []
self.up_blocks = []
# create encoder path
for i in range(self.n_blocks):
num_filters_in = self.in_channels if i == 0 else num_filters_out
num_filters_out = self.start_filters * (2 ** i)
pooling = True if i < self.n_blocks - 1 else False
down_block = DownBlock(in_channels=num_filters_in,
out_channels=num_filters_out,
pooling=pooling,
activation=self.activation,
normalization=self.normalization,
conv_mode=self.conv_mode,
dim=self.dim)
self.down_blocks.append(down_block)
# create decoder path (requires only n_blocks-1 blocks)
for i in range(n_blocks - 1):
num_filters_in = num_filters_out
num_filters_out = num_filters_in // 2
up_block = UpBlock(in_channels=num_filters_in,
out_channels=num_filters_out,
activation=self.activation,
normalization=self.normalization,
conv_mode=self.conv_mode,
dim=self.dim,
up_mode=self.up_mode)
self.up_blocks.append(up_block)
# final convolution
self.conv_final = get_conv_layer(num_filters_out, self.out_channels, kernel_size=1, stride=1, padding=0,
bias=True, dim=self.dim)
# add the list of modules to current module
self.down_blocks = nn.ModuleList(self.down_blocks)
self.up_blocks = nn.ModuleList(self.up_blocks)
# initialize the weights
self.initialize_parameters()
@staticmethod
def weight_init(module, method, **kwargs):
if isinstance(module, (nn.Conv3d, nn.Conv2d, nn.ConvTranspose3d, nn.ConvTranspose2d)):
method(module.weight, **kwargs) # weights
@staticmethod
def bias_init(module, method, **kwargs):
if isinstance(module, (nn.Conv3d, nn.Conv2d, nn.ConvTranspose3d, nn.ConvTranspose2d)):
method(module.bias, **kwargs) # bias
def initialize_parameters(self,
method_weights=nn.init.xavier_uniform_,
method_bias=nn.init.zeros_,
kwargs_weights={},
kwargs_bias={}
):
for module in self.modules():
self.weight_init(module, method_weights, **kwargs_weights) # initialize weights
self.bias_init(module, method_bias, **kwargs_bias) # initialize bias
def forward(self, x: torch.tensor):
encoder_output = []
# Encoder pathway
for module in self.down_blocks:
x, before_pooling = module(x)
encoder_output.append(before_pooling)
# Decoder pathway
for i, module in enumerate(self.up_blocks):
before_pool = encoder_output[-(i + 2)]
x = module(before_pool, x)
x = self.conv_final(x)
return x
def __repr__(self):
attributes = {attr_key: self.__dict__[attr_key] for attr_key in self.__dict__.keys() if '_' not in attr_key[0] and 'training' not in attr_key}
d = {self.__class__.__name__: attributes}
return f'{d}'
# + [markdown] pycharm={"name": "#%% md\n"}
# ## Create U-Net and try random input image
# + pycharm={"name": "#%%\n"}
# from unet import UNet
model = UNet(in_channels=1,
out_channels=1,
n_blocks=4,
start_filters=32,
activation='relu',
normalization='batch',
conv_mode='same',
dim=2)
x = torch.randn(size=(1, 1, 512, 512), dtype=torch.float32)
with torch.no_grad():
out = model(x)
print(f'Out: {out.shape}')
plt.imshow(x[0,0,:,:,], cmap='gray')
plt.colorbar()
plt.figure()
plt.imshow(out[0,0,:,:,], cmap='gray')
plt.colorbar()
# -
# # Dimensions
#
# Dimenze může být změněna, musí však zůstat nezměněný počet obrazových bodů.
# + pycharm={"name": "#%%\n"}
# loss = torch.nn.BCELoss(weight=None)
x_new = x.reshape([1,1,1,1,1,512,1,512])
x_new.shape
# + [markdown] pycharm={"name": "#%% md\n"}
# # Dataloader
#
# Kvůli tomu, že načítáme data ve 3D, ale pak využíváme jen řezy, není možné použít `dataloader`.
# Asi bychom si museli napsat svůj.
# + pycharm={"name": "#%%\n"}
# for X, y in test_dataloader:
# print("Shape of X [N, C, H, W]: ", X.shape)
# print("Shape of y: ", y.shape, y.dtype)
# break
# -
# Takže data můžeme procházet třeba takto
# + pycharm={"name": "#%%\n"}
for X,y in train_data:
print("Shape of X [N, C, H, W]: ", X.shape)
print("Shape of y: ", y.shape, y.dtype)
break # odstranit, je to tu pro demonstrační účely, aby for běžel jen jednou
# -
# Nebo si velké množství řezů můžeme rozděli na `batche`
# + pycharm={"name": "#%%\n"}
for X,y in train_data:
print("Shape of X [N, C, H, W]: ", X.shape)
print("Shape of y: ", y.shape, y.dtype)
for ind in range(0, X.shape[0], batch_size):
Xbatch = X[ind:ind+batch_size]
ybatch = y[ind:ind+batch_size]
print(" Shape of X [N, C, H, W]: ", Xbatch.shape)
print(" Shape of y: ", ybatch.shape, ybatch.dtype)
# tady bude trénování
break
# + pycharm={"name": "#%%\n"}
| devel/unet_torch_kidney.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Multihot study fitting in 64 bits
#
# This study tries to cut the number of embedding dimensions and will continue from the points done in multihot_study_simple
# The idea is to cut down from the embedding dimension of 324 to something much more manageable.
#
# So I want to go UTF8ed (utf-8 embedding dimension) no more than 64 bits, why? just because
#
# The low limit of the embedding would be 32 bits (the maximum lenght of an utf-8 code)
#
# $ 32 <= UTF8ed <= 64 $
#
# For this I want to basically do the following: ${N\choose k}$
#
# Where $ 32 <= N <= 64$
# and $ k $ should be minimized to augment the sparcity of the vector as much as possible
#
# Also I would like to add some verification or checking elements that should be also more important, for example, the first 4 elements should indicate which UTF-8 segment is being used. This implies $ 32 <= N <= 60$
#
#
# The value $k$ should be around the $k^{th}$ root of the product of the first $k$ parts of $N!$
#
# So I will try some values for k
import numpy as np
import itertools
from itertools import combinations
# +
# as a first experiment I would like to see how many
# the number of items that need to be included in the coding scheme:
ncodes = 1112064 # number of valid codes in UTF-8 per Wikipedia page
# -
list(range(32,32-4,-1))
list(range(1,4+1))
#find the minimum N for which the condition is filled
for N in range(32,64):
for k in [4,5]:
v = np.prod(list(range(N,N-k,-1))) / np.prod(list(range(1,k+1)))
if v > ncodes:
print("ncodes={}; N={},k={}".format(v N,k))
break
# so the values are $ N >= 45 ; k >=5 $
#
# Which means that for a code of dim 64 I can use a one-hot for the first 4 elements such as it indicates the utf-8 plane segment and tehre are still 15 elements to signal some other things (such as a positional embedding or an error correction code for example).
#
# So I decide to create a code of dimension $ N=49 $ and leave the rest of the space for dimensional embedding or other thing (64 would be great for grouped convolution features and 49 is only divisible by 7)
# From these 49 elements, the only available values will be $0$ and $1$, the first 4 elements will be selected according to the plane segment used in UTF-8, and the rest should indicate all the selection (this adds redundancy but also makes things more clear)
list(combinations(list(range(5)), 2))
# so, basically I have to do something like the following:
#
# - generate all combinations of ${45\choose 5}$
# - assing to each an index
# - convert all that to numpy and vectors of size 45
# +
def get_all_combinations(N,k):
ret = combinations(list(range(N)),k) #iterator
return ret
# -
all_combs = get_all_combinations(45,5)
indices = np.array(list(all_combs))
indices.shape
indices[:5]
embeds = np.zeros([indices.shape[0], 45])
embeds.shape
# numpy.put works with indices as if the array is flattened so I have to work on that
lin_indices = np.array(list(range(embeds.shape[0])))
lin_indices = lin_indices.reshape([-1,1])
lin_indices.shape
lin_indices[:20]
flat_indices = (lin_indices*45)+indices
flat_indices[:10]
embeds.put(flat_indices,[1])
embeds[-4:]
# This covers a complete codebook, now the issue might be the distance between two elements of the code. In this case the distance is quite small, so I can add some extra dimensions that increments the distance between vectors...
# Maybe what I can do is actually use the next 15 dimensions (to fill up to 64 dimensions) ... so something might come up of it
# After thinking about several methods, specially on Fowraed Error Correction Codes like TurboCodes, LDPC and ReedSolomon. Other error detection codes (that use parity codes) are not necessarilly useful as the parity will always be the same in the codebook by construction (which is another nice thing). There is another thing here, is that many codes (like golay or hamming) have fixed size for the messages which do not match the needs in the codes here.
#
# So basically what needs to be done is augment the distance between two elements, which can be done easily.
#
# In this case I can do that with an easy trick that will augment distance between the points, maybe do several one-hot like the one used in the previous codebook I worked on.
# arr = [3,5,7,11,13,17,19,23]
arr = [5,7,11,13,17,19,23]
np.prod(arr), np.sum(arr)
# The issue with this is that the dimensionality grows more than I fixed I wanted to work on.
#
# So I can use the same technique but with the 15 elements I have left as max dimension that I fixed (just because I wanted to)
#
# Note that all these decisions on dimensionality are completely arbitrary, adding constraints just for the sake of cutting down the number of operations and trainable parameters.
#
# The idea of having a fixed codebook is to get free of it later.
#
# So for this extra code of 15 elements will be created in a way that all pairs are co-primes (this increases the distance between vectors on the cycles), the easiest way of selecting co-primes is selecting prime numbers, also there is a nice thing in the sequence $[3,5,7]$ that they sum 15 which is exactly the same as the allowed space I gave myself to build that.
eyes = np.eye(3), np.eye(5), np.eye(7)
eyes[0].repeat(4)
np.tile(eyes[0],(3,1))
rep3, rep5, rep7 = int(np.ceil(embeds.shape[0]/3.)), int(np.ceil(embeds.shape[0]/5.)), int(np.ceil(embeds.shape[0]/7.))
reps = [rep3,rep5,rep7]
reps
# And now I build the codebook
tiles = []
for e,r in zip(eyes, reps):
t = np.tile(e, [r,1])[:embeds.shape[0],:]
tiles.append(t)
[t.shape for t in tiles]
code15 = np.concatenate(tiles,axis=1)
code15.shape
embeds45 = np.concatenate([embeds,code15],axis=1)
embeds45.shape
embeds45bool = np.array(embeds45, dtype=bool)
# Now I want to compute the distances between vectors, just to know about them .. but the dimensionality of the vector makes it big and out of memory errors appear, so I'll do splits to try to get it right.
splits = np.array_split(embeds45bool, 1000)
splits[0][:2]
from scipy.spatial.distance import cdist,pdist, hamming
# from scipy.spatial import distance
dd = cdist(embeds45bool,splits[0][:2], metric='hamming')
# pp = pdist(embeds45bool[:10,:],splits[0][:2])
hh = hamming(embeds45bool[0], splits[0][3])
splits[0].shape
embeds45bool[0],embeds45[0],
dd.shape, hh.shape
hh
ddf = dd.flat
ddf[ddf>0]
np.min(ddf[ddf>0])
np.min(dd)
# The next experiment should not be run lightly as it is heavy and time consuming (one run takes about 138 seconds wall time, so about 140s I estimate about 39 hours, or about 2 days of runtime in my computer, I can not parallelize more due to memory issues which I only have 64GB)
# +
# # %%time
# isplits = splits[:5]
# # # from scipy.spatial.distance import cdist
# # # # cdist(XA, XB, metric='euclidean', p=2, V=None, VI=None, w=None)
# # # maxd, mind,
# diststats = []
# for s in isplits:
# fdist = cdist(embeds45bool,s, metric='hamming').flat
# nzfdist = fdist[fdist>0] # eliminate from the elements the zero distances (distance to itself)
# # save stat values
# diststats.append( (np.min(nzfdist), np.max(nzfdist), np.median(nzfdist), np.std(nzfdist) ))
# +
# diststats
# -
np.array([0.03333333333333333 , 0.26666666666666666 , 0.23333333333333334]) * 45
# +
# # %%time
# splits2 = np.array_split(embeds45, 1000)
# isplits = splits2[:5]
# # # from scipy.spatial.distance import cdist
# # # # cdist(XA, XB, metric='euclidean', p=2, V=None, VI=None, w=None)
# # # maxd, mind,
# diststats2 = []
# for s in isplits:
# fdist = cdist(embeds45,s, metric='hamming').flat
# nzfdist = fdist[fdist>0] # eliminate from the elements the zero distances (distance to itself)
# # save stat values
# diststats2.append( (np.min(nzfdist), np.max(nzfdist), np.median(nzfdist), np.std(nzfdist) ))
# +
# diststats2
# -
# Now I'll get to do the segment coding this makes an extra 4 elements that encode each segment and the special tokens
#
# It will be a one hot encoding and all ones when is a special token and I can use the **utf-8 private area**
#
# From the previous study the indices for the codes are:
print("indices for the segments: ", 0, 128, (128 + 2**5 * 2**6), (128 + 2**4 * (2**6)**2), (128 + 2**3 * (2**6)**3) )
# So what I need are a few elements at some point to use them as private values.
#
# In the case of utf-8 there are non used values taht I can use for this purpose, or I can add some extra values at the beginning.
#
#
# the segment indicator vector will be shape (embeds.shape[0], 4)
segind = np.zeros((embeds.shape[0], 4))
segind.shape
# I'll use the last 4 codes as special codes, these will be set for the following elements:
# * \<error> $last$
# * \<start> $last-1$
# * \<stop> $last-2$
# * \<unknown> $last-3$
# * \<null> $last-4$
#
#
# Other elements might be needed, but as the encoding is much bigger than the complete utf-8 space I'll be able to add them later if the need arrives.
#
#
# Special codes have the segment indicator part set to *1111*
segind[-5:] = 1
segind[-6:]
# here is where the pre-computed indices are of use
# 0 128 2176 65664 2097280
segind[:128] = np.array([0,0,0,1])
segind[128:2176] = np.array([0,0,1,0])
segind[2176:65664] = np.array([0,1,0,0])
# segind[65664:] = np.array([1,0,0,0])
segind[65664:-113854] = np.array([1,0,0,0]) # where 113855 is the number of special codes that fit in this coding but I leave one for margin
segind[-6:] = 1
segind[120:130]
segind[2170:2180]
segind[65660:65670]
segind[-10:]
# Now I can create the complete codebook:
embeds64 = np.concatenate([embeds45,segind],axis=1)
embeds64.shape
# Now I have all the codes, this should be enough for many things. Nevertheless even if this is an encoding that can capture everything, the decoding part as well as the learning might prove problematic.
# One-hot is quite nice for learning and decoding while this encoding will need some other techniques for decoding and measuring loss (cosine similarity for example?, using faiss might be an option)
#
# There is another way of encoding this, try to maximize the distance between elements in the *SAME* utf-8 code segment, this could be more beneficial as most of the text in one text or language should (mostly) be in the same segment while (maybe) having a few words or codes from the other ones (exceptions would be the punctuation and emoticons codes), but for the moment I'll just create my codes as is and be done with it.
from utf8_encoder import *
tables = create_tables(segments=4)
len(tables)
_, _, _, char2idx, idx2char = tables
type(char2idx)
# if we check the number of codes generated is
len(char2idx), len(idx2char)
# which is less than: 1221759
1221759 - 1107904
# +
# what I want to do now is to save the coding but for that I need to add the special characters,
# <err> (error) 𝑙𝑎𝑠𝑡 = 1221758
# <start> 𝑙𝑎𝑠𝑡−1 = 1221757
# <stop> 𝑙𝑎𝑠𝑡−2 = 1221756
# <unk> (unknown) 𝑙𝑎𝑠𝑡−3 = 1221755
# <null> 𝑙𝑎𝑠𝑡−4 = 1221754
# char2idx["<err>"] = 1221758
# char2idx["<start>"] = 1221757
# char2idx["<stop>"] = 1221756
# char2idx["<unk>"] = 1221755
# char2idx["<null>"] = 1221754
# idx2char[1221758] = "<err>"
# idx2char[1221757] = "<start>"
# idx2char[1221756] = "<stop>"
# idx2char[1221755] = "<unk>"
# idx2char[1221754] = "<null>"
# eslen = len(embeds64)
# idx2char["<err>"] = eslen-1
# idx2char["<start>"] = eslen-2
# idx2char["<stop>"] = eslen-3
# idx2char["<unk>"] = eslen-4
# idx2char["<null>"] = eslen-5
# idx2char[eslen-1] = "<err>"
# idx2char[eslen-2] = "<start>"
# idx2char[eslen-3] = "<stop>"
# idx2char[eslen-4] = "<unk>"
# idx2char[eslen-5] = "<null>"
# -
embeds64bool = np.array(embeds64, dtype=bool)
# +
# list(char2idx.items())[:100]
# +
# list(idx2char.items())[:100]
# +
# list(embeds64[[0,120,240,360,480,600,720,840,960,1080,1200,1320]])
# +
# and now SAVE all the codes
# save_obj(char2idx, "multihot64-char2idx")
# save_obj(idx2char, "multihot64-idx2char")
# save_obj(embeds64, "multihot64-embeds")
# save_obj(embeds64bool, "multihot64-embeds-bool")
# -
# ls -lh
# The code is a bit big, so I'll cut out the part that is NOT used and leave just a few places for special codes, the rest, forget about it
#eliminate the values that we'll not use and keep the most distanced objects for special use
embeds64short = np.concatenate([embeds64[:-113855], embeds64[-6:]], axis=0)
# char2idxshort = np.concatenate([char2idx[:-113854], char2idx[-6:]], axis=0)
# idx2charshort = np.concatenate([idx2char[:-113854], idx2char[-6:]], axis=0)
embeds64short.shape
# +
eslen = len(embeds64short)
char2idx["<err>"] = eslen-1
char2idx["<start>"] = eslen-2
char2idx["<stop>"] = eslen-3
char2idx["<unk>"] = eslen-4
char2idx["<null>"] = eslen-5
idx2char[eslen-1] = "<err>"
idx2char[eslen-2] = "<start>"
idx2char[eslen-3] = "<stop>"
idx2char[eslen-4] = "<unk>"
idx2char[eslen-5] = "<null>"
# -
embeds64short = np.array(embeds64short, dtype='float32')
embeds64shortbool = np.array(embeds64short, dtype=bool)
# del idx2char[1221758]
# del idx2char[1221757]
# del idx2char[1221756]
# del idx2char[1221755]
# del idx2char[1221754]
# +
# del(char2idx)
# del(idx2char)
# -
embeds64short.dtype
len(char2idx), len(idx2char), embeds64short.shape
# Now I do some verification of the elements to be sure that all goes OK
aidx = set(range(embeds64short.shape[0]))
cidx = set(char2idx.values())
idxc = set(idx2char.keys())
len(idxc.intersection(cidx)) # intersection OK
idxc.difference(cidx), cidx.difference(idxc)
# This set should have 1 non used value (a special token space), this is by construction to get some space in case I need it and not having to change the codebook, just add it to the dictionary assignment
idxc.difference(aidx), aidx.difference(idxc)
embeds64short[[1107904]]
# and now SAVE all the codes
save_obj(char2idx, "multihot64short-char2idx")
save_obj(idx2char, "multihot64short-idx2char")
save_obj(embeds64short, "multihot64short-embeds")
save_obj(embeds64shortbool, "multihot64short-embeds-bool")
# Checking this change only, the complete numpy pickled embedding codebook changes from:
# * 625540774 bytes multihot64-embeds.pkl to
# * 567250086 bytes multihot64short-embeds.pkl
#
# so, about 55MBs difference
#
| predictors/sequence/text/multihot-64_study.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Manual Jupyter Notebook:
#
# https://athena.brynmawr.edu/jupyter/hub/dblank/public/Jupyter%20Notebook%20Users%20Manual.ipynb
# #Jupyter Notebook Users Manual
#
# This page describes the functionality of the [Jupyter](http://jupyter.org) electronic document system. Jupyter documents are called "notebooks" and can be seen as many things at once. For example, notebooks allow:
#
# * creation in a **standard web browser**
# * direct **sharing**
# * using **text with styles** (such as italics and titles) to be explicitly marked using a [wikitext language](http://en.wikipedia.org/wiki/Wiki_markup)
# * easy creation and display of beautiful **equations**
# * creation and execution of interactive embedded **computer programs**
# * easy creation and display of **interactive visualizations**
#
# Jupyter notebooks (previously called "IPython notebooks") are thus interesting and useful to different groups of people:
#
# * readers who want to view and execute computer programs
# * authors who want to create executable documents or documents with visualizations
#
# <hr size="5"/>
# ###Table of Contents
# * [1. Getting to Know your Jupyter Notebook's Toolbar](#1.-Getting-to-Know-your-Jupyter-Notebook's-Toolbar)
# * [2. Different Kinds of Cells](#2.-Different-Kinds-of-Cells)
# * [2.1 Code Cells](#2.1-Code-Cells)
# * [2.1.1 Code Cell Layout](#2.1.1-Code-Cell-Layout)
# * [2.1.1.1 Row Configuration (Default Setting)](#2.1.1.1-Row-Configuration-%28Default-Setting%29)
# * [2.1.1.2 Cell Tabbing](#2.1.1.2-Cell-Tabbing)
# * [2.1.1.3 Column Configuration](#2.1.1.3-Column-Configuration)
# * [2.2 Markdown Cells](#2.2-Markdown-Cells)
# * [2.3 Raw Cells](#2.3-Raw-Cells)
# * [2.4 Header Cells](#2.4-Header-Cells)
# * [2.4.1 Linking](#2.4.1-Linking)
# * [2.4.2 Automatic Section Numbering and Table of Contents Support](#2.4.2-Automatic-Section-Numbering-and-Table-of-Contents-Support)
# * [2.4.2.1 Automatic Section Numbering](#2.4.2.1-Automatic-Section-Numbering)
# * [2.4.2.2 Table of Contents Support](#2.4.2.2-Table-of-Contents-Support)
# * [2.4.2.3 Using Both Automatic Section Numbering and Table of Contents Support](#2.4.2.3-Using-Both-Automatic-Section-Numbering-and-Table-of-Contents-Support)
# * [3. Keyboard Shortcuts](#3.-Keyboard-Shortcuts)
# * [4. Using Markdown Cells for Writing](#4.-Using-Markdown-Cells-for-Writing)
# * [4.1 Block Elements](#4.1-Block-Elements)
# * [4.1.1 Paragraph Breaks](#4.1.1-Paragraph-Breaks)
# * [4.1.2 Line Breaks](#4.1.2-Line-Breaks)
# * [4.1.2.1 Hard-Wrapping and Soft-Wrapping](#4.1.2.1-Hard-Wrapping-and-Soft-Wrapping)
# * [4.1.2.2 Soft-Wrapping](#4.1.2.2-Soft-Wrapping)
# * [4.1.2.3 Hard-Wrapping](#4.1.2.3-Hard-Wrapping)
# * [4.1.3 Headers](#4.1.3-Headers)
# * [4.1.4 Block Quotes](#4.1.4-Block-Quotes)
# * [4.1.4.1 Standard Block Quoting](#4.1.4.1-Standard-Block-Quoting)
# * [4.1.4.2 Nested Block Quoting](#4.1.4.2-Nested-Block-Quoting)
# * [4.1.5 Lists](#4.1.5-Lists)
# * [4.1.5.1 Ordered Lists](#4.1.5.1-Ordered-Lists)
# * [4.1.5.2 Bulleted Lists](#4.1.5.2-Bulleted-Lists)
# * [4.1.6 Section Breaks](#4.1.6-Section-Breaks)
# * [4.2 Backslash Escape](#4.2-Backslash-Escape)
# * [4.3 Hyperlinks](#4.3-Hyperlinks)
# * [4.3.1 Automatic Links](#4.3.1-Automatic-Links)
# * [4.3.2 Standard Links](#4.3.2-Standard-Links)
# * [4.3.3 Standard Links With Mouse-Over Titles](#4.3.3-Standard-Links-With-Mouse-Over-Titles)
# * [4.3.4 Reference Links](#4.3.4-Reference-Links)
# * [4.3.5 Notebook-Internal Links](#4.3.5-Notebook-Internal-Links)
# * [4.3.5.1 Standard Notebook-Internal Links Without Mouse-Over Titles](#4.3.5.1-Standard-Notebook-Internal-Links-Without-Mouse-Over-Titles)
# * [4.3.5.2 Standard Notebook-Internal Links With Mouse-Over Titles](#4.3.5.2-Standard-Notebook-Internal-Links-With-Mouse-Over-Titles)
# * [4.3.5.3 Reference-Style Notebook-Internal Links](#4.3.5.3-Reference-Style-Notebook-Internal-Links)
# * [4.4 Tables](#4.4-Tables)
# * [4.4.1 Cell Justification](#4.4.1-Cell-Justification)
# * [4.5 Style and Emphasis](#4.5-Style-and-Emphasis)
# * [4.6 Other Characters](#4.6-Other-Characters)
# * [4.7 Including Code Examples](#4.7-Including-Code-Examples)
# * [4.8 Images](#4.8-Images)
# * [4.8.1 Images from the Internet](#4.8.1-Images-from-the-Internet)
# * [4.8.1.1 Reference-Style Images from the Internet](#4.8.1.1-Reference-Style-Images-from-the-Internet)
# * [4.9 LaTeX Math](#4.9-LaTeX-Math)
# * [5. Bibliographic Support](#5.-Bibliographic-Support)
# * [5.1 Creating a Bibtex Database](#5.1-Creating-a-Bibtex-Database)
# * [5.1.1 External Bibliographic Databases](#5.1.1-External-Bibliographic-Databases)
# * [5.1.2 Internal Bibliographic Databases](#5.1.2-Internal-Bibliographic-Databases)
# * [5.1.2.1 Hiding Your Internal Database](#5.1.2.1-Hiding-Your-Internal-Database)
# * [5.1.3 Formatting Bibtex Entries](#5.1.3-Formatting-Bibtex-Entries)
# * [5.2 Cite Commands and Citation IDs](#5.2-Cite-Commands-and-Citation-IDs)
# * [6. Turning Your Jupyter Notebook into a Slideshow](#6.-Turning-Your-Jupyter-Notebook-into-a-Slideshow)
#
# # 1. Getting to Know your Jupyter Notebook's Toolbar
# + [markdown] slideshow={"slide_type": "fragment"}
# At the top of your Jupyter Notebook window there is a toolbar. It looks like this:
# + [markdown] slideshow={"slide_type": "fragment"}
# 
# + [markdown] slideshow={"slide_type": "fragment"}
# Below is a table which helpfully pairs a picture of each of the items in your toolbar with a corresponding explanation of its function.
# + [markdown] slideshow={"slide_type": "fragment"}
# Button|Function
# -|-
# |This is your save button. You can click this button to save your notebook at any time, though keep in mind that Jupyter Notebooks automatically save your progress very frequently.
# |This is the new cell button. You can click this button any time you want a new cell in your Jupyter Notebook.
# |This is the cut cell button. If you click this button, the cell you currently have selected will be deleted from your Notebook.
# |This is the copy cell button. If you click this button, the currently selected cell will be duplicated and stored in your clipboard.
# |This is the past button. It allows you to paste the duplicated cell from your clipboard into your notebook.
# |These buttons allow you to move the location of a selected cell within a Notebook. Simply select the cell you wish to move and click either the up or down button until the cell is in the location you want it to be.
# |This button will "run" your cell, meaning that it will interpret your input and render the output in a way that depends on [what kind of cell] [cell kind] you're using.
# |This is the stop button. Clicking this button will stop your cell from continuing to run. This tool can be useful if you are trying to execute more complicated code, which can sometimes take a while, and you want to edit the cell before waiting for it to finish rendering.
# |This is the restart kernel button. See your kernel documentation for more information.
# |This is a drop down menu which allows you to tell your Notebook how you want it to interpret any given cell. You can read more about the [different kinds of cells] [cell kind] in the following section.
# |Individual cells can have their own toolbars. This is a drop down menu from which you can select the type of toolbar that you'd like to use with the cells in your Notebook. Some of the options in the cell toolbar menu will only work in [certain kinds of cells][cell kind]. "None," which is how you specify that you do not want any cell toolbars, is the default setting. If you select "Edit Metadata," a toolbar that allows you to edit data about [Code Cells][code cells] directly will appear in the corner of all the Code cells in your notebook. If you select "Raw Cell Format," a tool bar that gives you several formatting options will appear in the corner of all your [Raw Cells][raw cells]. If you want to view and present your notebook as a slideshow, you can select "Slideshow" and a toolbar that enables you to organize your cells in to slides, sub-slides, and slide fragments will appear in the corner of every cell. Go to [this section][slideshow] for more information on how to create a slideshow out of your Jupyter Notebook.
# |These buttons allow you to move the location of an entire section within a Notebook. Simply select the Header Cell for the section or subsection you wish to move and click either the up or down button until the section is in the location you want it to be. If your have used [Automatic Section Numbering][section numbering] or [Table of Contents Support][table of contents] remember to rerun those tools so that your section numbers or table of contents reflects your Notebook's new organization.
# |Clicking this button will automatically number your Notebook's sections. For more information, check out the Reference Guide's [section on Automatic Section Numbering][section numbering].
# |Clicking this button will generate a table of contents using the titles you've given your Notebook's sections. For more information, check out the Reference Guide's [section on Table of Contents Support][table of contents].
# |Clicking this button will search your document for [cite commands][] and automatically generate intext citations as well as a references cell at the end of your Notebook. For more information, you can read the Reference Guide's [section on Bibliographic Support][bib support].
# |Clicking this button will toggle [cell tabbing][], which you can learn more about in the Reference Guides' [section on the layout options for Code Cells][cell layout].
# |Clicking this button will toggle the [collumn configuration][] for Code Cells, which you can learn more about in the Reference Guides' [section on the layout options for Code Cells][cell layout].
# |Clicking this button will toggle spell checking. Spell checking only works in unrendered [Markdown Cells][] and [Header Cells][]. When spell checking is on all incorrectly spelled words will be underlined with a red squiggle. Keep in mind that the dictionary cannot tell what are [Markdown][md writing] commands and what aren't, so it will occasionally underline a correctly spelled word surrounded by asterisks, brackets, or other symbols that have specific meaning in Markdown.
#
#
# [cell kind]: #2.-Different-Kinds-of-Cells "Different Kinds of Cells"
# [code cells]: #2.1-Code-Cells "Code Cells"
# [raw cells]: #2.3-Raw-Cells "Raw Cells"
# [slideshow]: #6.-Turning-Your-Jupyter-Notebook-into-a-Slideshow "Turning Your Jupyter Notebook Into a Slideshow"
# [section numbering]: #2.4.2.1-Automatic-Section-Numbering
# [table of contents]: #2.4.2.2-Table-of-Contents-Support
# [cell tabbing]: #2.1.1.2-Cell-Tabbing
# [cell layout]: #2.1.1-Code-Cell-Layout
# [bib support]: #5.-Bibliographic-Support
# [cite commands]: #5.2-Cite-Commands-and-Citation-IDs
# [md writing]: #4.-Using-Markdown-Cells-for-Writing
# [collumn configuration]: #2.1.1.3-Column-Configuration
# [Markdown Cells]: #2.2-Markdown-Cells
# [Header Cells]: #2.4-Header-Cells
#
# + [markdown] slideshow={"slide_type": "slide"}
# # 2. Different Kinds of Cells
# -
# There are essentially four kinds of cells in your Jupyter notebook: Code Cells, Markdown Cells, Raw Cells, and Header Cells, though there are six levels of Header Cells.
# ## 2.1 Code Cells
# By default, Jupyter Notebooks' Code Cells will execute Python. Jupyter Notebooks generally also support JavaScript, Python, HTML, and Bash commands. For a more comprehensive list, see your Kernel's documentation.
# ### 2.1.1 Code Cell Layout
# Code cells have both an input and an output component. You can view these components in three different ways.
# #### 2.1.1.1 Row Configuration (Default Setting)
# Unless you specific otherwise, your Code Cells will always be configured this way, with both the input and output components appearing as horizontal rows and with the input above the output. Below is an example of a Code Cell in this default setting:
# + format="row"
2 + 3
# -
# #### 2.1.1.2 Cell Tabbing
# Cell tabbing allows you to look at the input and output components of a cell separately. It also allows you to hide either component behind the other, which can be usefull when creating visualizations of data. Below is an example of a tabbed Code Cell:
# + format="tab"
2+3
# -
# #### 2.1.1.3 Column Configuration
# Like the row configuration, the column layout option allows you to look at both the input and the output components at once. In the column layout, however, the two components appear beside one another, with the input on the left and the output on the right. Below is an example of a Code Cell in the column configuration:
# + format="column"
2+3
# -
# ## 2.2 Markdown Cells
# In Jupyter Notebooks, Markdown Cells are the easiest way to write and format text. For a more thorough explanation of how to write in Markdown cells, refer to [this section of the guide][writing markdown].
#
# [writing markdown]: #4.-Using-Markdown-Cells-for-Writing "Using Markdown Cells for Writing"
#
#
# ## 2.3 Raw Cells
# Raw Cells, unlike all other Jupyter Notebook cells, have no input-output distinction. This means that Raw Cells cannot be rendered into anything other than what they already are. If you click the run button in your tool bar with a Raw Cell selected, the cell will remain exactly as is and your Jupyter Notebook will automatically select the cell directly below it. Raw cells have no style options, just the same monospace font that you use in all other unrendered Notebook cells. You cannot bold, italicize, or enlarge any text or characters in a Raw Cell.
#
# Because they have no rendered form, Raw Cells are mainly used to create examples. If you save and close your Notebook and then reopen it, all of the Code, Markdown, and Header Cells will automatically render in whatever form you left them when you first closed the document. This means that if you wanted to preserve the unrendered version of a cell, say if you were writing a computer science paper and needed code examples, or if you were writing [documentation on how to use Markdown] [writing markdown] and needed to demonstrate what input would yield which output, then you might want to use a Raw Cell to make sure your examples stayed in their most useful form.
#
# [writing markdown]: #4.-Using-Markdown-Cells-for-Writing "Using Markdown Cells for Writing"
# ## 2.4 Header Cells
# While it is possible to organize your document using [Markdown headers][], Header Cells provide a more deeply structural organization for your Notebook and thus there are several advantages to using them.
#
# [Markdown headers]: #4.1.3-Headers "Headers"
# ### 2.4.1 Linking
# Header Cells have specific locations inside your Notebook. This means you can use them to [create Notebook-internal links](#4.3.5-Notebook-Internal-Links "Notebook-Internal Links").
# ### 2.4.2 Automatic Section Numbering and Table of Contents Support
# Your Jupyter Notebook has two helpful tools that utilize the structural organization that Header Cells give your document: automatic section numbering and table of contents generation.
# #### 2.4.2.1 Automatic Section Numbering
# Suppose you are writing a paper and, as is prone to happening when you have a lot of complicate thoughts buzzing around your brain, you've reorganized your ideas several times. Automatic section numbering will go through your Notebook and number your sections and subsection as designated by your Header Cells. This means that if you've moved one or more big sections around several times, you won't have to go through your paper and renumber it, as well as all its subsections, yourself.
#
#
#
# **Notes:** Automatic Section Numbering tri-toggling tool, so when you click the Number Sections button one of three actions will occur: Automatic Section Numbering will number your sections, correct inconsistent numbering, or unnumber your sections (if all of your sections are already consistently and correctly numbered).
#
# So, even if you have previously numbered your sections, Automatic Section Numbering will go through your document, delete the current section numbers, and replace them the correct number in a linear sequence. This means that if your third section was once your second, Automatic Section Numbering will delete the "2" in front of your section's name and replace it with a "3."
#
# While this function saves you a lot of time, it creates one limitation. Maybe you're writing a paper about children's books and one of the books you're discussing is called **`2 Cats`**. You've unsurprisingly titled the section where you summarize and analyze this book **`2 Cats`**. Automatic Section Numbering will assume the number 2 is section information and delete it, leaving just the title **`Cats`** behind. If you bold, italicize, or place the title of the section inside quotes, however, the entire section title will be be preserved without any trouble. It should also be noted that even if you must title a section with a number occurring before any letters and you do not want to bold it, italicize it, or place it inside quotes, then you can always run Automatic Section Numbering and then go to that section and retype its name by hand.
#
# Because Automatic Section Numbering uses your header cells, its performance relies somewhat on the clarity of your organization. If you have two sections that begin with Header 1 Cells in your paper, and each of the sections has two subsections that begin with Header 2 Cells, Automatic Section Numbering will number them 1, 1.1, 1.2, 2, 2.1, and 2.2 respectively. If, however, you have used a Header 3 Cell to indicate the beginning of what would have been section 2.1, Automatic Section Numbering will number that section 2.0.1 and an error message will appear telling you that "You placed a Header 3 cell under a Header 2 Cell in section 2". Similarly, if you begin your paper with any Header Cell smaller than a Header 1, say a Header 3 Cell, then Automatic Section Numbering will number your first section 0.0.3 and an error message will appear telling you that "Notebook begins with a Header 3 Cell."
#
# #### 2.4.2.2 Table of Contents Support
# The Table of Contents tool will automatically generate a table of contents for your paper by taking all your Header Cell titles and ordering them in a list, which it will place in a new cell at the very beginning of you Notebook. Because your Notebook does note utilize formal page breaks or numbers, each listed section will be hyperlinked to the actual section within your document.
#
# **Notes: **Because Table of Contents Support uses your header cells, its performance relies somewhat on the clarity of your organization. If you have two sections that begin with Header 1 Cells in your paper, and each of the sections has two subsections that begin with Header 2 Cells, Table of Contents will order them in the following way:
#
# * 1.
# * 1.1
# * 1.2
# * 2.
# * 2.1
# * 2.2
#
#
# If, however, you have used a Header 3 Cell to indicate the beginning of what would have been section 2.1, Table of Contents Support will insert a dummy line so that your table of contents looks like this:
#
#
# * 1.
# * 1.1
# * 1.2
# * 2.
# *
# * 2.0.1
# * 2.2
#
#
# #### 2.4.2.3 Using Both Automatic Section Numbering and Table of Contents Support
# Automatic Section Numbering will always update every aspect of your notebook that is dependent on the title of one or more of your sections. This means that it will automatically correct an existing table of contents and all of your Notebook-internal links to reflect the new numbered section titles.
#
# # 3. Keyboard Shortcuts
# Jupyter Notebooks support many helpful Keyboard shortcuts, including ones for most of the buttons in [your toolbar][]. To view these shortcuts, you can click the help menu and then select Keyboard Shortcuts, as pictured below.
#
# [your toolbar]: #1.-Getting-to-Know-your-Jupyter-Notebook's-Toolbar "Getting to know Your Jupyter Notebook's Toolbar"
# 
# # 4. Using Markdown Cells for Writing
# **Why aren't there font and font size selection drop down menus, buttons I can press to bold and italicize my text, or other advanced style options in my Notebook?**
# When you use Microsoft Word, Google Docs, Apple Pages, Open Office, or any other word processing software, you generally use your mouse to select various style options, like line spacing, font size, font color, paragraph format etc. This kind of system is often describes as a WYSIWYG (What You See Is What You Get) interface. This means that the input (what you tell the computer) exactly matches the output (what the computer gives back to you). If you type the letter **`G`**, highlight it, select the color green and up the font size to 64 pt, your word processor will show you a fairly large green colored letter **`G`**. And if you print out that document you will print out a fairly large green colored letter **`G`**.
#
# This Notebook, however, does not use a WYSIWYG interface. Instead it uses something called a "[markup Language][]". When you use a a markup language, your input does not necessarily exactly equal your output.
#
#
# [markup language]: http://en.wikipedia.org/wiki/Markup_language "Wikipedia Article on Markup"
#
#
# For example, if I type "#Header 1" at the beginning of a cell, but then press Shift-Enter (or click the play button at the top of the window), this notebook will turn my input into a somewhat different output in the following way:
# <pre>
# #Header 1
# </pre>
# #Header 1
# And if I type "##Header 2" (at the beginning of a cell), this notebook will turn that input into another output:
# <pre>
# ##Header 2
# </pre>
# ##Header 2
# In these examples, the hashtags are markers which tell the Notebook how to typeset the text. There are many markup languages, but one family, or perhaps guiding philosophy, of markup languages is called "Markdown," named somewhat jokingly for its simplicity. Your Notebook uses "marked," a Markdown library of typeset and other formatting instructions, like the hashtags in the examples above.
#
# Markdown is a markup language that generates HTML, which the cell can interpret and render. This means that Markdown Cells can also render plain HTML code. If you're interested in learning HTML, check out this [helpful online tutorial][html tutorial].
#
# [html tutorial]: http://www.w3schools.com/html/ "w3schools.com HTML Tutorial"
# **Why Use Markdown (and not a WYSIWYG)?**
# Why is Markdown better? Well, it’s worth saying that maybe it isn't. Mainly, it’s not actually a question of better or worse, but of what’s in front of you and of who you are. A definitive answer depends on the user and on that user’s goals and experience. These Notebooks don't use Markdown because it's definitely better, but rather because it's different and thus encourages users to think about their work differently.
#
# It is very important for computer science students to learn how to conceptualize input and output as dependent, but also distinct. One good reason to use Markdown is that it encourages this kind of thinking. Relatedly, it might also promote focus on substance over surface aesthetic. Markdown is somewhat limited in its style options, which means that there are inherently fewer non-subject-specific concerns to agonize over while working. It is the conceit of this philosophy that you would, by using Markdown and this Notebook, begin to think of the specific stylistic rendering of your cells as distinct from what you type into those same cells, and thus also think of the content of your writing as necessarily separate from its formating and appearance.
# ## 4.1 Block Elements
# ### 4.1.1 Paragraph Breaks
# Paragraphs consist of one or more consecutive lines of text and they are separated by one or more blank lines. If a line contains only spaces, it is a blank line.
# ### 4.1.2 Line Breaks
# #### 4.1.2.1 Hard-Wrapping and Soft-Wrapping
# If you're used to word processing software, you've been writing with automatically hard-wrapped lines and paragraphs. In a hard-wrapped paragraph the line breaks are not dependent on the size of the viewing window. If you click and drag your mouse to expand a word processing document, for example, the shape of the paragraphs and the length of the lines will not change. In other words, the length of a hard-wrapped line is determined either by the number of words in the line (in the case of word processing software where this number is predetermined and the program wraps for the user automatically), or individual intention (when a user manually presses an Enter or Return key to control exactly how long a line is).
#
# Soft-wrapped paragraphs and lines, however, *do* depend on the size of their viewing window. If you increase the size of a window where soft-wrapped paragraphs are displayed, they too will expand into longer lines, becoming shorter and wider to fill the increased window space horizontally. Unsurprising, then, if you *narrow* a window, soft-wrapped lines will shrink and the paragraphs will become longer vertically.
#
# Markdown, unlike most word processing software, does not automatically hard-wrap. If you want your paragraphs to have a particular or deliberate shape and size, you must insert your own break by ending the line with two spaces and then typing Return.
#
# #### 4.1.2.2 Soft-Wrapping
# <tt>
# blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah
# </tt>
# blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah
# #### 4.1.2.3 Hard-Wrapping
# <tt>
# blah blah blah blah blah
# blah blah blah blah blah blah blah blah blah
# blah blah blah blah blah blah blah blah blah blah blah
# blah blah blah blah blah
# blah blah blah blah blah
# blah blah blah blah blah blah blah blah blah blah blah blah blah
# </tt>
# blah blah blah blah blah
# blah blah blah blah blah blah blah blah blah
# blah blah blah blah blah blah blah blah blah blah blah
# blah blah blah blah blah
# blah blah blah blah blah
# blah blah blah blah blah blah blah blah blah blah blah blah blah
#
# ### 4.1.3 Headers
# <pre>
# #Header 1
# </pre>
# #Header 1
# <pre>
# ##Header 2
# </pre>
# ##Header 2
# <pre>
# ###Header 3
# </pre>
# ###Header 3
# <pre>
# ####Header 4
# </pre>
# ####Header 4
# <pre>
# #####Header 5
# </pre>
# #####Header 5
# <pre>
# ######Header 6
# </pre>
# ######Header 6
# ### 4.1.4 Block Quotes
# #### 4.1.4.1 Standard Block Quoting
# <tt>
# >blah blah block quote blah blah block quote blah blah block
# quote blah blah block quote blah blah block
# quote blah blah block quote blah blah block quote blah blah block quote
# </tt>
# >blah blah block quote blah blah block quote blah blah block
# quote blah blah block quote blah blah block
# quote blah blah block quote blah blah block quote blah blah block quote
# **Note**: Block quotes work best if you intentionally hard-wrap the lines.
# #### 4.1.4.2 Nested Block Quoting
# <pre>
# >blah blah block quote blah blah block quote blah blah block
# block quote blah blah block block quote blah blah block
# >>quote blah blah block quote blah blah
# block block quote blah blah block
# >>>quote blah blah block quote blah blah block quote blah blah block quote
# </pre>
# >blah blah block quote blah blah block quote blah blah block
# block quote blah blah block block quote blah blah block
# >>quote blah blah block quote blah blah
# block block quote blah blah block
# >>>quote blah blah block quote blah blah block quote blah blah block quote
# ### 4.1.5 Lists
# #### 4.1.5.1 Ordered Lists
# In Markdown, you can list items using numbers, a **`+`**, a **` - `**, or a **`*`**. However, if the first item in a list or sublist is numbered, Markdown will interpret the entire list as ordered and will automatically number the items linearly, no matter what character you use to denote any given separate item.
# <pre>
# ####Groceries:
#
# 0. Fruit:
# 6. Pears
# 0. Peaches
# 3. Plums
# 4. Apples
# 2. <NAME>
# 7. Gala
# * Oranges
# - Berries
# 8. Strawberries
# + Blueberries
# * Raspberries
# - Bananas
# 9. Bread:
# 9. Whole Wheat
# 0. With oats on crust
# 0. Without oats on crust
# 0. Rye
# 0. White
# 0. Dairy:
# 0. Milk
# 0. Whole
# 0. Skim
# 0. Cheese
# 0. Wisconsin Cheddar
# 0. Pepper Jack
# </pre>
# ####Groceries:
#
# 0. Fruit:
# 6. Pears
# 0. Peaches
# 3. Plums
# 4. Apples
# 2. <NAME>
# 7. Gala
# * Oranges
# - Berries
# 8. Strawberries
# + Blueberries
# * Raspberries
# - Bananas
# 9. Bread:
# 9. Whole Wheat
# 0. With oats on crust
# 0. Without oats on crust
# 0. Rye
# 0. White
# 0. Dairy:
# 0. Milk
# 0. Whole
# 0. Skim
# 0. Cheese
# 0. Wisconsin Cheddar
# 0. Pepper Jack
# #### 4.1.5.2 Bulleted Lists
# If you begin your list or sublist with a **`+`**, a **` - `**, or a **`*`**, then Markdown will interpret the whole list as unordered and will use bullets regardless of the characters you type before any individual list item.
# <pre>
# ####Groceries:
#
# * Fruit:
# * Pears
# 0. Peaches
# 3. Plums
# 4. Apples
# - <NAME>
# 7. Gala
# * Oranges
# - Berries
# - Strawberries
# + Blueberries
# * Raspberries
# - Bananas
# 9. Bread:
# * Whole Wheat
# * With oats on crust
# 0. Without oats on crust
# + Rye
# 0. White
# 0. Dairy:
# * Milk
# + Whole
# 0. Skim
# - Cheese
# - Wisconsin Cheddar
# 0. Pepper Jack
# </pre>
# ####Groceries:
#
# * Fruit:
# * Pears
# 0. Peaches
# 3. Plums
# 4. Apples
# - <NAME>
# 7. Gala
# * Oranges
# - Berries
# - Strawberries
# + Blueberries
# * Raspberries
# - Bananas
# 9. Bread:
# * Whole Wheat
# * With oats on crust
# 0. Without oats on crust
# + Rye
# 0. White
# 0. Dairy:
# * Milk
# + Whole
# 0. Skim
# - Cheese
# - Wisconsin Cheddar
# 0. <NAME>
# ### 4.1.6 Section Breaks
# <pre>
# ___
# </pre>
# ___
# <pre>
# ***
# </pre>
# ***
# <pre>------</pre>
# ------
# <pre>
# * * *
# </pre>
# * * *
# <pre>
# _ _ _
# </pre>
# _ _ _
# <pre>
# - - -
# </pre>
# - - -
# ## 4.2 Backslash Escape
# What happens if you want to include a literal character, like a **`#`**, that usually has a specific function in Markdown? Backslash Escape is a function that prevents Markdown from interpreting a character as an instruction, rather than as the character itself. It works like this:
# <pre>
# \# Wow, this isn't a header.
# # This is definitely a header.
# </pre>
# \# Wow, this isn't a header.
# # This is definitely a header.
# Markdown allows you to use a backslash to escape from the functions of the following characters:
# * \ backslash
# * ` backtick
# * \* asterisk
# * _ underscore
# * {} curly braces
# * [] square brackets
# * () parentheses
# * \# hashtag
# * \+ plus sign|
# * \- minus sign (hyphen)
# * . dot
# * ! exclamation mark
# ## 4.3 Hyperlinks
# ### 4.3.1 Automatic Links
# <pre>
# http://en.wikipedia.org
# </pre>
# http://en.wikipedia.org
# ### 4.3.2 Standard Links
# <pre>
# [click this link](http://en.wikipedia.org)
# </pre>
# [click this link](http://en.wikipedia.org)
# ### 4.3.3 Standard Links With Mouse-Over Titles
# <pre>
# [click this link](http://en.wikipedia.org "Wikipedia")
# </pre>
# [click this link](http://en.wikipedia.org "Wikipedia")
# ### 4.3.4 Reference Links
# Suppose you are writing a document in which you intend to include many links. The format above is a little arduous and if you have to do it repeatedly *while* you're trying to focus on the content of what you're writing, it's going to be a really big pain.
#
# Fortunately, there is an alternative way to insert hyperlinks into your text, one where you indicate that there is a link, name that link, and then use the name to provide the actually URL later on when you're less in the writing zone. This method can be thought of as a "reference-style" link because it is similar to using in-text citations and then defining those citations later in a more detailed reference section or bibliography.
#
# <pre>
# This is [a reference] [identification tag for link]
#
# [identification tag for link]: http://en.wikipedia.org/wiki/Chile "Wikipedia Article About Chile"
# </pre>
# This is [a reference] [identification tag for link]
#
# [identification tag for link]: http://en.wikipedia.org/wiki/Chile "Wikipedia Article About Chile"
# **Note:** The "identification tag for link" can be anything. For example:
# <pre>
# This is [a reference] [lfskdhflhslgfh333676]
#
# [lfskdhflhslgfh333676]: http://en.wikipedia.org/wiki/Chile "Wikipedia Article About Chile"
# </pre>
# This is [a reference] [lfskdhflhslgfh333676]
#
# [lfskdhflhslgfh333676]: http://en.wikipedia.org/wiki/Chile "Wikipedia Article About Chile"
# This means you can give your link an intuitive, easy to remember, and relevant ID:
# <pre>
# This is [a reference][Chile]
#
# [chile]: http://en.wikipedia.org/wiki/Chile "Wikipedia Article About Chile"
# </pre>
# This is [a reference][Chile]
#
# [chile]: http://en.wikipedia.org/wiki/Chile "Wikipedia Article About Chile"
# **Note**: Link IDs are not case-sensitive.
# If you don't want to give your link an ID, you don't have to. As a short cut, Markdown will understand if you just use the words in the first set of brackets to define the link later on. This works in the following way:
# <pre>
# This is [a reference][]
#
# [a reference]: http://en.wikipedia.org/wiki/Chile "Wikipedia Article About Chile"
# </pre>
# This is [a reference][]
#
# [a reference]: http://en.wikipedia.org/wiki/Chile "Wikipedia Article About Chile"
# Another really helpful feature of a reference-style link is that you can define the link anywhere in the cell. (must be in the cell) For example:
#
# <tt>
# This is [a reference] [ref] blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah
# blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah <br/><br/>
#
# [ref]: http://en.wikipedia.org/wiki/Chile "Wikipedia Article About Chile"
# </tt>
# This is [a reference] [ref] blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah
# blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah blah
#
# [ref]: http://en.wikipedia.org/wiki/Chile "Wikipedia Article About Chile"
# **Note:** Providing a mouse-over title for any link, regardless of whether it is a standard or reference-stlye type, is optional. With reference-style links, you can include the mouse-over title by placing it in quotes, single quotes, or parentheses. For standard links, you can only define a mouse-over title in quotes.
#
#
# ### 4.3.5 Notebook-Internal Links
# When you create a Header you also create a discrete location within your Notebook. This means that, just like you can link to a specific location on the web, you can also link to a Header Cell inside your Notebook. Internal links have very similar Markdown formatting to regular links. The only difference is that the name of the link, which is the URL in the case of external links, is just a hashtag plus the name of the Header Cell you are linking to (case-sensitive) with dashes in between every word. If you hover your mouse over a Header Cell, a blue Greek pi letter will appear next to your title. If you click on it, the URL at the top of your window will change and the internal link to that section will appear last in the address. You can copy and paste it in order to make an internal link inside a Markdown Cell.
# #### 4.3.5.1 Standard Notebook-Internal Links Without Mouse-Over Titles
# <pre>
# [Here's a link to the section of Automatic Section Numbering](#Automatic-Section-Numbering)
# </pre>
# [Here's a link to the section of Automatic Section Numbering](#2.4.2.1-Automatic-Section-Numbering)
# #### 4.3.5.2 Standard Notebook-Internal Links With Mouse-Over Titles
# <pre>
# [Here's a link to the section on lists](#Lists "Lists")
# </pre>
# [Here's a link to the section of Automatic Section Numbering](#2.4.2.1-Automatic-Section-Numbering)
# #### 4.3.5.3 Reference-Style Notebook-Internal Links
# <pre>
# [Here's a link to the section on Table of Contents Support][TOC]
#
# [TOC]: #Table-of-Contents-Support
# </pre>
# [Here's a link to the section on Table of Contents Support][TOC]
#
# [TOC]: #2.4.2.2-Table-of-Contents-Support
# ## 4.4 Tables
# In Markdown, you can make a table by using vertical bars and dashes to define the cell and header borders:
# <pre>
# |Header|Header|Header|Header|
# |------|------|------|------|
# |Cell |Cell |Cell | Cell |
# |Cell |Cell |Cell | Cell |
# |Cell |Cell |Cell | Cell |
# |Cell |Cell |Cell | Cell |
# </pre>
# |Header|Header|Header|Header|
# |------|------|------|------|
# |Cell |Cell |Cell | Cell |
# |Cell |Cell |Cell | Cell |
# |Cell |Cell |Cell | Cell |
# |Cell |Cell |Cell | Cell |
#
# Making a table this way might be especially useful if you want your document to be legible both rendered and unrendered. However, you don't *need* to include all of those dashes, vertical bars, and spaces for Markdown to understand that you're making a table. Here's the bare minimum you would need to create the table above:
# <pre>
# Header|Header|Header|Header
# -|-|-|-
# Cell|Cell|Cell|Cell
# Cell|Cell|Cell|Cell
# Cell|Cell|Cell|Cell
# Cell|Cell|Cell|Cell
# </pre>
# Header|Header|Header|Header
# -|-|-|-
# Cell|Cell|Cell|Cell
# Cell|Cell|Cell|Cell
# Cell|Cell|Cell|Cell
# Cell|Cell|Cell|Cell
#
# It's important to note that the second line of dashes and vertical bars is essential. If you have just the line of headers and the second line of dashes and vertical bars, that's enough for Markdown to make a table.
#
# Another important formatting issue has to do with the vertical bars that define the left and right edges of the table. If you include all the vertical bars on the far left and right of the table, like in the first example above, Markdown will ignore them completely. *But*, if you leave out some and include others, Markdown will interpret any extra vertical bar as an additional cell on the side that the bar appears in the unrendered version of the text. This also means that if you include the far left or right vertical bar in the second line of bars and dashes, you must include all of the otherwise optional vertical bars (like in the first example above).
# ### 4.4.1 Cell Justification
# If not otherwise specified the text in each header and cell of a table will justify to the left. If, however, you wish to specify either right justification or centering, you may do so like this:
# <tt>
# **Centered, Right-Justified, and Regular Cells and Headers**:
#
# centered header | regular header | right-justified header | centered header | regular header
# :-:|-|-:|:-:|-
# centered cell|regular cell|right-justified cell|centered cell|regular cell
# centered cell|regular cell|right-justified cell|centered cell|regular cell
# </tt>
# **Centered, Right-Justified, and Regular Cells and Headers**:
#
# centered header | regular header | right-justified header | centered header | regular header
# :-:|-|-:|:-:|-
# centered cell|regular cell|right-justified cell|centered cell|regular cell
# centered cell|regular cell|right-justified cell|centered cell|regular cell
#
# While it is difficult to see that the headers are differently justified from one another, this is just because the longest line of characters in any column defines the width of the headers and cells in that column.
# **Note:** You cannot make tables directly beneath a line of text. You must put a blank line between the end of a paragraph and the beginning of a table.
# ## 4.5 Style and Emphasis
# <pre>
# *Italics*
# </pre>
# *Italics*
# <pre>
# _Italics_
# </pre>
# _Italics_
# <pre>
# **Bold**
# </pre>
# **Bold**
# <pre>
# __Bold__
# </pre>
# __Bold__
# **Note:** If you want actual asterisks or underscores to appear in your text, you can use the [backslash escape function] [backslash] like this:
#
# [backslash]: #4.2-Backslash-Escape "Backslash Escape"
# <pre>
# \*awesome asterisks\* and \_incredible under scores\_
# </pre>
# \*awesome asterisks\* and \_incredible under scores\_
# ## 4.6 Other Characters
# <pre>
# Ampersand &amp; Ampersand
# </pre>
# Ampersand & Ampersand
# <pre>
# &lt; angle brackets &gt;
# </pre>
# < angle brackets >
# <pre>
# &quot; quotes &quot;
# " quotes "
# ## 4.7 Including Code Examples
# If you want to signify that a particular section of text is actually an example of code, you can use backquotes to surround the code example. These will switch the font to monospace, which creates a clear visual formatting difference between the text that is meant to be code and the text that isn't.
#
# Code can either in the middle of a paragraph, or as a block. Use a single backquote to start and stop code in the middle of a paragraph. Here's an example:
# <pre>
# The word `monospace` will appear in a code-like form.
# </pre>
# The word `monospace` will appear in a code-like form.
# **Note:** If you want to include a literal backquote in your code example you must suround the whole text block in double backquotes like this:
# <pre>
# `` Look at this literal backquote ` ``
# </pre>
# `` Look at this literal backquote ` ``
# To include a complete code-block inside a Markdown cell, use triple backquotes. Optionally, you can put the name of the language that you are quoting after the starting triple backquotes, like this:
# <pre>
# ```python
# def function(n):
# return n + 1
# ```
# </pre>
# That will format the code-block (sometimes called "fenced code") with syntax coloring. The above code block will be rendered like this:
# ```python
# def function(n):
# return n + 1
# ```
# The language formatting names that you can currently use after the triple backquote are:
# <pre>
# apl django go jinja2 ntriples q smalltalk toml
# asterisk dtd groovy julia octave r smarty turtle
# clike dylan haml less pascal rpm smartymixed vb
# clojure ecl haskell livescript pegjs rst solr vbscript
# cobol eiffel haxe lua perl ruby sparql velocity
# coffeescript erlang htmlembedded markdown php rust sql verilog
# commonlisp fortran htmlmixed pig sass stex xml
# css gas http mirc properties scheme tcl xquery
# d gfm jade mllike puppet shell tiddlywiki yaml
# diff gherkin javascript nginx python sieve tiki z80
# </pre>
# ## 4.8 Images
# ### 4.8.1 Images from the Internet
# Inserting an image from the internet is almost identical to inserting a link. You just also type a **`!`** before the first set of brackets:
# <pre>
# 
# </pre>
# 
# **Note:** Unlike with a link, the words that you type in the first set of brackets do not appear when they are rendered into html by Markdown.
# #### 4.8.1.1 Reference-Style Images from the Internet
# Just like with links, you can also use a reference-style format when inserting images from the internet. This involves indicating where you want to place a picture, giving that picture an ID tag, and then later defining that ID tag. The process is nearly identical to using the reference-style format to insert a link:
# <pre>
# ![][giraffe]
#
# [giraffe]:http://upload.wikimedia.org/wikipedia/commons/thumb/b/b7/South_African_Giraffe,_head.jpg/877px-South_African_Giraffe,_head.jpg "Picture of a Giraffe"
# </pre>
# ![][giraffe]
#
# [giraffe]: http://upload.wikimedia.org/wikipedia/commons/thumb/b/b7/South_African_Giraffe,_head.jpg/877px-South_African_Giraffe,_head.jpg "Picture of a Giraffe"
# ## 4.9 LaTeX Math
# Jupyter Notebooks' Markdown cells support LateX for formatting mathematical equations. To tell Markdown to interpret your text as LaTex, surround your input with dollar signs like this:
# <pre>
# $z=\dfrac{2x}{3y}$
# </pre>
# $z=\dfrac{2x}{3y}$
# An equation can be very complex:
#
# <pre>
# $F(k) = \int_{-\infty}^{\infty} f(x) e^{2\pi i k} dx$
# </pre>
# $F(k) = \int_{-\infty}^{\infty} f(x) e^{2\pi i k} dx$
# If you want your LaTex equations to be indented towards the center of the cell, surround your input with two dollar signs on each side like this:
# <pre>
# $$2x+3y=z$$
# </pre>
# $$2x+3y=z$$
# For a comprehensive guide to the mathematical symbols and notations supported by Jupyter Notebooks' Markdown cells, check out [<NAME>'s helpful reference materials on the subject][mkeefe].
#
# [mkeefe]: http://martinkeefe.com/math/mathjax1 "<NAME>'s MathJax Guide"
# # 5. Bibliographic Support
# Bibliographic Support makes managing references and citations in your Notebook much easier, by automating some of the bibliographic process every person goes through when doing research or writing in an academic context. There are essentially three steps to this process for which your Notebook's Bibliographic support can assist: gathering and organizing sources you intend to use, citing those sources within the text you are writing, and compiling all of the material you referenced in an organized, correctly formatted list, the kind which usually appears at the end of a paper in a section titled "References," "Bibliography," or "Works Cited.
#
# In order to benefit from this functionality, you need to do two things while writing your paper: first, you need to create a [Bibtex database][bibdb] of information about your sources and second, you must use the the [cite command][cc] in your Markdown writing cells to indicate where you want in-text citations to appear.
#
# If you do both these things, the "Generate References" button will be able to do its job by replacing all of your cite commands with validly formatted in-text citations and creating a References section at the end of your document, which will only ever include the works you specifically cited within in your Notebook.
#
# **Note:** References are generated without a header cell, just a [markdown header][]. This means that if you want a References section to appear in your table of contents, you will have to unrender the References cell, delete the "References" header, make a Header Cell of the appropriate level and title it "References" yourself, and then generate a table of contents using [Table of Contents Support][table of contents]. This way, you can also title your References section "Bibliography" or "Works Cited," if you want.
#
# [markdown header]: #4.1.3-Headers
# [table of contents]: #2.4.2.2-Table-of-Contents-Support
# [bibdb]: #5.1-Creating-a-Bibtex-Database
# [cc]:#5.2-Cite-Commands-and-Citation-IDs
#
#
#
# ## 5.1 Creating a Bibtex Database
# Bibtex is reference management software for formatting lists of references ([from Wikipedia](BibTeX is reference management software for formatting lists of references "Wikipedia Article On Bibtex")). While your Notebook does not use the Bibtex software, it does use [Bibtex formatting](#5.1.3-Formatting-Bibtex-Entries) for creating references within your Bibliographic database.
#
# In order for the Generate References button to work, you need a bibliographic database for it to search and match up with the sources you've indicated you want to credit using [cite commands and citation IDs](#5.2-Cite-Commands-and-Citation-IDs).
#
# When creating a bibliographic database for your Notebook, you have two options: you can make an external database, which will exist in a separate Notebook from the one you are writing in, or you can make an internal database which will exist in a single cell inside the Notebook in which you are writing. Below are explanations of how to use these database creation strategies, as well as a discussion of the pros and cons for each.
# ### 5.1.1 External Bibliographic Databases
# To create an external bibliographic database, you will need to create a new Notebook and title it **`Bibliography`** in the toplevel folder of your current Jupyter session. As long as you do not also have an internal bibliographic database, when you click the Generate References button your Notebook's Bibliographic Support will search this other **`Bibliography`** Notebook for Bibtex entries. Bibtex entries can be in any cell and in any kind of cell in your **`Bibliography`** Notebook as long as the cell begins with **`<!--bibtex`** and ends with **`-->`**. Go to [this section][bibfor] for examples of valid BibTex formatting.
#
# Not every cell has to contain BibTex entries for the external bibliographic database to work as intended with your Notebook's bibliographic support. This means you can use the same helpful organization features that you use in other Notebooks, like [Automatic Section Numbering][asn] and [Table of Contents Support][toc], to structure your own little library of references. The best part of this is that any Notebook containing validly formatted [cite commands][cc] can check your external database and find only the items that you have indicated you want to cite. So you only ever have to make the entry once and your external database can grow large and comprehensive over the course of your accademic writing career.
#
# There are several advantages to using an external database over [an internal one][internal database]. The biggest one, which has already been described, is that you will only ever need to create one and you can organize it into sections by using headers and generating [automatic section numbers][asn] and a [table of contents][toc]. These tools will help you to easily find the right [citation ID][cc] for a given source you want to cite. The other major advantage is that an external database is not visible when viewing the Notebook in which you are citing sources and generating a References list. Bibtex databases are not very attractive or readable and you probably won't want one to show up in your finished document. There are [ways to hide internal databases][hiding bibtex cell], but it's convenient not to have to worry about that.
#
#
# [asn]: #2.4.2.1-Automatic-Section-Numbering
# [toc]: #2.4.2.2-Table-of-Contents-Support
# [cc]: #5.2-Cite-Commands-and-Citation-IDs
# [hiding bibtex cell]: #5.1.2.1-Hiding-Your-Internal-Database
# [bibfor]:#5.1.3-Formatting-Bibtex-Entries
# ### 5.1.2 Internal Bibliographic Databases
# Unlike [external bibliographic databases][exd], which are comprised from an entire separate notebook, internal bibliographic databases consist of only one cell within in the Notebook in which you are citing sources and compiling a References list. The single cell, like all of the many BibTex cells that can make up an external database, must begin with **`<!--bibtex`** and end with **`-->`** in order to be validly formatted and correctly interpreted by your Notebook's Bibliographic Support. It's probably best to keep this cell at the very end or the very beginning of your Notebook so you always know where it is. This is because when you use an intenral bibliographic databse it can only consist of one cell. This means that if you want to cite multiple sources you will need to keep track of the single cell that comprises your entire internal bibliographic database during every step of the research and writing process.
#
# Internal bibliographic databases make more sense when your project is a small one and the list of total sources is short. This is especially convenient if you don't already have a built-up external database. With an internal database you don't have to create and organize a whole separate Notebook, a task that's only useful when you have to keep track of a lot of different material. Additionally, if you want to share your finished Notebook with others in a form that retains its structural validity, you only have to send one Notebook, as oppose to both the project itself and the Notebook that comprises your external bibliographic database. This is especially useful for a group project, where you want to give another reader the ability to edit, not simply read, your References section.
#
# [exd]:#5.1.1-External-Bibliographic-Databases
#
# #### 5.1.2.1 Hiding Your Internal Database
# Even though they have some advantages, especially for smaller projects, internal databases have on major draw back. They are not very attractive or polished looking and you probably won't want one to appear in your final product. Fortunately, there are two methods for hiding your internal biblioraphic database.
#
# While your Notebook's bibliographic support will be able to interpret [correctly formatted BibTex entries][bibfor] in any [kind of cell][cell kind], if you use a [Markdown Cell][md cell] to store your internal bibliographic database, then when you run the cell all of the ugly BibTex formatting will disappear. This is handy, but it also makes the cell very difficult to find, so remember to keep careful track of where your hidden BibTex databse is if you're planning to edit it later. If you want your final product to be viewed stably as HTML, then you can make your internal BibTex database inside a [Raw Cell][RC], use the [cell toolbar][] to select "Raw Cell Format", and then select "None" in the toolbar that appears in the corner of your Raw Cell BibTex database. This way, you will still be able to easily find and edit the database when you are working on your Notebook, but others won't be able to see the database when viewing your project in its final form.
#
#
# [cell toolbar]: #1.-Getting-to-Know-your-Jupyter-Notebook's-Toolbar
# [bibfor]:#5.1.3-Formatting-Bibtex-Entries
# [RC]:#2.3-Raw-Cells
# [md cell]: #2.2-Markdown-Cells
# [cell kind]: #2.-Different-Kinds-of-Cells
# ### 5.1.3 Formatting Bibtex Entries
# BibTex entries consist of three crucial components: one, the type of source you are citing (a book, article, website, etc.); two, the unique [citation ID][cc] you wish to remember the source by; and three, the fields of information about that source (author, title of work, date of publication, etc.). Below is an example entry, with each of these three components designated clearly
#
# <pre>
#
# <!--bibtex
#
# @ENTRY TYPE{CITATION ID,
# FIELD 1 = {source specific information},
# FIELD 2 = {source specific informatio},
# FIEL 3 = {source specific informatio},
# FIELD 4 = {source specific informatio}
# }
#
# -->
#
# </pre>
#
# More comprehensive documentation of what entry types and corresponding sets of required and optional fields BibTex supports can be found in the [Wikipedia article on BibTex][wikibibt].
#
# Below is a section of the external bibliographic database for a fake history paper about the fictional island nation of Calico. (None of the entries contain information about real books or articles):
#
# [cc]: #5.2-Cite-Commands-and-Citation-IDs
# [wikibibt]: http://en.wikipedia.org/wiki/Markdown
#
#
#
# <pre>
#
# <!--bibtex
#
# @book{wellfarecut,
# title = {Our Greatest Threat: The Rise of Anti-Wellfare Politics in Calico in the 21st Century},
# author = {<NAME>},
# year = {2010},
# publisher = {Jupyter University Press}
# }
#
# @article{militaryex2,
# title = {Rethinking Calican Military Expansion for the New Century},
# author = {<NAME>.},
# journal = {Modern Politics},
# volume = {60},
# issue = {25},
# pages = {35 - 70},
# year = {2012}
# }
#
# @article{militaryex1,
# title = {Conservative Majority Passes Budget to Grow Military},
# author = {<NAME>},
# journal = {The Daily Calican},
# month = {October 19th, 2011},
# pages = {15 - 17},
# year = {2011}
# }
#
# @article{oildrill,
# title = {Oil Drilling Off the Coast of Jupyter Approved for Early Next Year},
# author = {<NAME>.},
# journal = {The Python Gazette},
# month = {December 5th, 2012},
# pages = {8 - 9},
# year = {2012}
# }
#
# @article{rieseinterview,
# title = {Interview with Up and Coming Freshman Senator, Alec Riese of Python},
# author = {<NAME>},
# journal = {The Jupyter Times},
# month = {November 24th, 2012},
# pages = {4 - 7},
# year = {2012}
# }
#
# @book{calicoww2:1,
# title = {Calico and WWII: Untold History},
# author = {<NAME>},
# year = {1997},
# publisher = {Calicia City Free Press}
# }
#
# @book{calicoww2:2,
# title = {Rebuilding Calico After Japanese Occupation},
# author = {<NAME> },
# year = {2002},
# publisher = {Python Books}
# }
# -->
# </pre>
# ## 5.2 Cite Commands and Citation IDs
# When you want to cite a bibliographic entry from a database (either internal or external), you must know the citation ID, sometimes called the "key", for that entry. Citation IDs are strings of letters, numbers, and symbols that *you* make up, so they can be any word or combination of words you find easy to remember. Once, you've given an entry a citation ID, however, you do need to use that same ID every time you cite that source, so it may behoove you to keep your database organized. This way it will be much easier to locate any given source's entry and its potentially forgotten citation ID.
#
# Once you know the citation ID for a given entry, use the following format to indicate to your Notebook's bibliographic support that you'd like to insert an in-text citation:
#
# <pre>
# [](#cite-CITATION ID)
# </pre>
#
# This format is the cite command. For example, if you wanted to cite *Rebuilding Calico After Japanese Occupation* listed above, you would use the cite command and the specific citation ID for that source:
#
# <pre>
# [](#cite-calicoww2:2)
# </pre>
#
# Before clicking the "Generate References" button, your unrendered text might look like this:
#
#
# <pre>
# Rebuilding Calico took many years [](#cite-calicoww2:2).
# </pre>
#
#
# After clicking the "Generate References" button, your unrendered text might look like this:
#
#
# <pre>
# Rebuilding Calico took many years <a name="ref-1"/>[(Kepps, 2002)](#cite-calicoww2:2).
# </pre>
#
#
# and then the text would render as:
#
#
# >Rebuilding Calico took many years <a name="ref-1"/>[(Kepps, 2002)](#cite-calicoww2:2).
#
#
# In addition, a cell would be added at the bottom with the following contents:
#
#
# >#References
#
# ><a name="cite-calicoww2:2"/><sup>[^](#ref-1) [^](#ref-2) </sup>Kepps, Milo . 2002. _Rebuilding Calico After Japanese Occupation_.
#
#
# # 6. Turning Your Jupyter Notebook into a Slideshow
# To install slideshow support for your Notebook, go [here](http://nbviewer.ipython.org/github/fperez/nb-slideshow-template/blob/master/install-support.ipynb).
#
# To see a tutorial and example slideshow, go [here](http://www.damian.oquanta.info/posts/make-your-slides-with-ipython.html).
| Cap01/JupyterNotebook-ManualUsuario.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 决策边界
# +
import numpy as np
import matplotlib.pyplot as plt
from sklearn import datasets
iris = datasets.load_iris()
X = iris.data
y = iris.target
X = X[y<2,:2]
y = y[y<2]
# -
plt.scatter(X[y==0,0], X[y==0,1], color="red")
plt.scatter(X[y==1,0], X[y==1,1], color="blue")
# +
from Logistic_Regression.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, seed=666)
# +
from Logistic_Regression.LogisticRegression import LogisticRegression
log_reg = LogisticRegression()
log_reg.fit(X_train, y_train)
# -
log_reg.coef_
log_reg.intercept_
def x2(x1):
return (-log_reg.coef_[0] * x1 - log_reg.intercept_) / log_reg.coef_[1]
x1_plot = np.linspace(4, 8, 1000)
x2_plot = x2(x1_plot)
plt.scatter(X[y==0,0], X[y==0,1], color="red")
plt.scatter(X[y==1,0], X[y==1,1], color="blue")
plt.plot(x1_plot, x2_plot)
plt.scatter(X_test[y_test==0,0], X_test[y_test==0,1], color="red")
plt.scatter(X_test[y_test==1,0], X_test[y_test==1,1], color="blue")
plt.plot(x1_plot, x2_plot)
# +
def plot_decision_boundary(model, axis):
x0, x1 = np.meshgrid(
np.linspace(axis[0], axis[1], int((axis[1]-axis[0])*100)).reshape(-1, 1),
np.linspace(axis[2], axis[3], int((axis[3]-axis[2])*100)).reshape(-1, 1),
)
X_new = np.c_[x0.ravel(), x1.ravel()]
y_predict = model.predict(X_new)
zz = y_predict.reshape(x0.shape)
from matplotlib.colors import ListedColormap
custom_cmap = ListedColormap(['#EF9A9A','#FFF59D','#90CAF9'])
plt.contourf(x0, x1, zz, linewidth=5, cmap=custom_cmap)
plot_decision_boundary(log_reg, axis=[4, 7.5, 1.5, 4.5])
plt.scatter(X[y==0,0], X[y==0,1])
plt.scatter(X[y==1,0], X[y==1,1])
# -
# ### kNN的决策边界
# +
from sklearn.neighbors import KNeighborsClassifier
knn_clf = KNeighborsClassifier()
knn_clf.fit(X_train, y_train)
# -
knn_clf.score(X_test, y_test)
plot_decision_boundary(knn_clf, axis=[4, 7.5, 1.5, 4.5])
plt.scatter(X[y==0,0], X[y==0,1])
plt.scatter(X[y==1,0], X[y==1,1])
knn_clf_all = KNeighborsClassifier()
knn_clf_all.fit(iris.data[:,:2], iris.target)
plot_decision_boundary(knn_clf_all, axis=[4, 8, 1.5, 4.5])
plt.scatter(iris.data[iris.target==0,0], iris.data[iris.target==0,1])
plt.scatter(iris.data[iris.target==1,0], iris.data[iris.target==1,1])
plt.scatter(iris.data[iris.target==2,0], iris.data[iris.target==2,1])
# +
knn_clf_all = KNeighborsClassifier(n_neighbors=50)
knn_clf_all.fit(iris.data[:,:2], iris.target)
plot_decision_boundary(knn_clf_all, axis=[4, 8, 1.5, 4.5])
plt.scatter(iris.data[iris.target==0,0], iris.data[iris.target==0,1])
plt.scatter(iris.data[iris.target==1,0], iris.data[iris.target==1,1])
plt.scatter(iris.data[iris.target==2,0], iris.data[iris.target==2,1])
# -
| 06LogisticRegression/03Decision-Boundary.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: EnergyDemandForecast local
# language: python
# name: energydemandforecast_local
# ---
# # Gradient boosting machine
# %matplotlib inline
import pandas as pd
import numpy as np
import matplotlib
matplotlib.use('Agg')
import matplotlib.pyplot as plt
import matplotlib.gridspec as gridspec
from sklearn.pipeline import Pipeline
from sklearn.ensemble import GradientBoostingRegressor
from sklearn.metrics import mean_squared_error
from sklearn.model_selection import TimeSeriesSplit
from sklearn.model_selection import RandomizedSearchCV
import pickle
import os
import scipy.stats as st
from itertools import combinations
from azure.storage.blob import BlockBlobService
from azureml.logging import get_azureml_logger
run_logger = get_azureml_logger()
run_logger.log('amlrealworld.timeseries.gbm','true')
model_name = "gbm"
aml_dir = os.environ['AZUREML_NATIVE_SHARE_DIRECTORY']
ACCOUNT_NAME = "petcgexperimentstorage"
ACCOUNT_KEY = "<KEY>
CONTAINER_NAME = "energy-demand-demo"
block_blob_service = BlockBlobService(account_name=ACCOUNT_NAME, account_key=ACCOUNT_KEY)
train = None
list_of_blobs = block_blob_service.list_blobs(CONTAINER_NAME)
for each_blob in list_of_blobs:
if ("part-00" in each_blob.name):
block_blob_service.get_blob_to_path(CONTAINER_NAME, blob_name=each_blob.name, file_path='./nyc_demand.csv')
train = pd.read_csv('./nyc_demand.csv')
break
# Use randomised search to find optimal hyperparameters. This model will take about 5 minutes to train. The quality of the solution may be improved by increasing the number of iterations in the randomized search, at the expense of increased training times.
X = train.drop(['demand', 'timeStamp'], axis=1)
regr = GradientBoostingRegressor()
tscv = TimeSeriesSplit(n_splits=3)
param_dist = {'n_estimators': st.randint(3, 100),
'learning_rate': st.uniform(0.01, 0.1),
'max_depth': range(2,31),
'min_samples_leaf': st.randint(1, 100),
'min_samples_split': st.randint(2, 50),
'max_features': range(3,X.shape[1]+1),
'subsample': st.uniform(0.1, 0.9)
}
regr_cv = RandomizedSearchCV(estimator=regr,
param_distributions=param_dist,
n_iter=50,
cv=tscv,
scoring='neg_mean_squared_error',
verbose=2,
n_jobs=-1)
regr_pipe = Pipeline([('regr_cv', regr_cv)])
regr_pipe.fit(X, y=train['demand'])
model_dir = "C:/Users/nelgoh/Desktop/Resources/Petronas/energy_demand_forecast/EnergyDemandForecast/outputs/models/"
with open(os.path.join(model_dir, model_name + '.pkl'), 'wb') as f:
pickle.dump(regr_pipe, f)
# Cross validation results
cv_results = pd.DataFrame(regr_pipe.named_steps['regr_cv'].cv_results_)
cv_results.sort_values(by='rank_test_score', inplace=True)
cv_results.head()
# Inspect the pairwise distribution of cross validation scores. Darker shades of blue indicate superior performance while the red star represents the optimal solution found.
params = ['param_n_estimators', 'param_max_depth', 'param_min_samples_split', 'param_max_features', 'param_subsample']
gs = gridspec.GridSpec(4,3)
fig = plt.figure(figsize=(15, 20), tight_layout=True)
plt_best = cv_results.head(1)
for idx, params in enumerate(combinations(params, 2)):
ax = fig.add_subplot(gs[idx])
plt.scatter(cv_results[params[0]], cv_results[params[1]], c=-np.log(-cv_results['mean_test_score']), s=100, cmap="Blues")
plt.scatter(plt_best[params[0]], plt_best[params[1]], s=500, marker="*", c="r")
plt.xlabel(params[0])
plt.ylabel(params[1])
# Inspect feature importance
feature_importance = pd.DataFrame.from_dict({'feature':X.columns, 'importance':regr_pipe.named_steps['regr_cv'].best_estimator_.feature_importances_})
feature_importance.plot.bar('feature', 'importance', figsize=(15,5), logy=True, title='Feature importance (log scale)', legend=False)
plt.show()
| 7-gbm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# #!/usr/bin/env python3
# -*- coding: utf-8 -*-
"""
Created on Thu Jun 24 17:17:54 2021
@author: <NAME>
"""
# install the libraries
import sys
# !{sys.executable} -m pip install numpy
# !{sys.executable} -m pip install matplotlib
# !{sys.executable} -m pip install pandas
# !{sys.executable} -m pip install statsmodels
# !{sys.executable} -m pip install sklearn
# !{sys.executable} -m pip install pyswarms
# !{sys.executable} -m pip install tensorflow
# !{sys.executable} -m pip install scipy
# import the libraries
import random
import numpy as np
import pandas as pd
import matplotlib as mpl
import matplotlib.pyplot as plt
import statsmodels.api as sm
import pyswarms as ps
from statsmodels.tsa.stattools import adfuller
from statsmodels.tsa.api import VAR
from statsmodels.graphics.tsaplots import plot_acf, plot_pacf
from keras.models import Sequential
from keras.layers import Dense
from scipy.optimize import differential_evolution
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import MinMaxScaler
from sklearn.preprocessing import StandardScaler
from sklearn.linear_model import LinearRegression
from sklearn.impute import SimpleImputer
from sklearn.compose import ColumnTransformer
from sklearn.metrics import mean_squared_error
from sklearn.metrics import mean_absolute_error
from sklearn.metrics import r2_score
# +
def preprocess_data(data):
#data['rows'] = data.index+1
X = data.drop(columns=['winangun'])
y = data['winangun']
exercise = X.columns.tolist()[1:]
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=42,
test_size=0.2, shuffle=False)
#scaler = MinMaxScaler()
#X_train = scaler.fit_transform(X_train)
#X_test = scaler.transform(X_test)
return X_train, X_test, y_train, y_test, exercise
# forward propagation
def forward_prop(params):
# roll-back the weights and biases
W1 = params[:i_weights].reshape((n_inputs, n_hidden))
b1 = params[i_weights:i_weights+i_bias].reshape((n_hidden,))
W2 = params[i_weights+i_bias:i_weights+i_bias+h_weights].reshape((n_hidden, n_classes))
b2 = params[i_weights+i_bias+h_weights:].reshape((n_classes,))
# perform forward propagation
z1 = X_train.dot(W1) + b1 # pre-activation in layer 1
a1 = np.where(z1 > 0, z1, z1 * 0.01) # LeakyReLU
z2 = a1.dot(W2) + b2 # pre-activation in layer 2
loss = mean_squared_error(y_train, z2)
return loss
def f(x):
n_particles = x.shape[0]
j = [forward_prop(x[i]) for i in range (n_particles)]
return np.array(j)
def train_model(options):
optimizer = ps.single.GlobalBestPSO(n_particles = 100,
dimensions = n_params,
options=options)
# perform optimization
cost, pos = optimizer.optimize(f, iters = 1000)
print('\nModel Run Times:')
return cost, pos, optimizer.cost_history
def prediction(X, pos):
# roll-back the weights and biases
W1 = pos[:i_weights].reshape((n_inputs, n_hidden))
b1 = pos[i_weights:i_weights+i_bias].reshape((n_hidden,))
W2 = pos[i_weights+i_bias:i_weights+i_bias+h_weights].reshape((n_hidden, n_classes))
b2 = pos[i_weights+i_bias+h_weights:].reshape((n_classes,))
# perform forward propagation
z1 = X.dot(W1) + b1 # pre-activation in layer 1
a1 = np.where(z1 > 0, z1, z1 * 0.01) # LeakReLu
z2 = a1.dot(W2) + b2 # pre-activation in layer 2
y_pred = z2
return y_pred
def plot_history(history):
plt.style.use('ggplot') # set 'classic' to use default style
plt.rcParams['ytick.right'] = False
plt.rcParams['ytick.labelright'] = False
plt.rcParams['ytick.left'] = True
plt.rcParams['ytick.labelleft'] = True
plt.rcParams['font.family'] = 'Arial'
plt.ylim([min(history)-5, max(history)+5])
plt.title('Cost History')
plt.plot(history)
# +
color_list = ["orange", "green"]
r = 1.7
# function to show plot
def Visualize(data):
features = list(data.select_dtypes(include=[np.number]).columns.values)
feature_size = len(features)
fig, axes = plt.subplots(
nrows = int(np.ceil(feature_size/2)),
ncols = 2, figsize = (14, feature_size * 2),
dpi = 150,
facecolor = "w",
edgecolor = "k"
)
for i in range(feature_size):
key = features[i]
c = color_list[i % (len(color_list))]
t_data = data[key]
t_data.head()
ax = t_data.plot(
ax = axes[i % 2],
color = c ,
title = "{}".format(key),
rot = 25
)
ax.legend([key])
plt.tight_layout()
# -
# import dataset
df = pd.read_csv('dataset_tondano_winangun.csv', index_col=0, parse_dates=True)
df
Visualize(df)
plt.savefig('data_preprocessing.png')
# +
# cleaning dataset
# change any zero value to NaN and fill NaN with mean value from dataframe
df=df.mask(df==0).fillna(df.mean())
#df = df.dropna() # remove empty Value
#df = df.fillna(0.1) # change NaN to 0.1
#df = df[(df.T != 0).any()] # remove all zero value
# show output
df
# -
Visualize(df)
plt.savefig('data_postprocessing.png')
# ACF Tondano and Winangun
acf_tondano = plot_acf(df['tondano']).legend(['Tondano'])
acf_winangun = plot_acf(df['winangun']).legend(['Winangun'])
plt.show()
# PACF Tondano and Winangun
pacf_tondano = plot_pacf(df['tondano']).legend(['Tondano'])
pacf_winangun = plot_pacf(df['winangun']).legend(['Winangun'])
plt.show()
# check is the series stationary or not using ADF
for i in range(len(df.columns)):
result = adfuller(df[df.columns[i]])
print(f"Test Statistics: {result[0]}")
print(f"P-Value: {result[1]}")
print(f"Critical Values: {result[4]}")
if result[1] > 0.05:
print("{}: Series is not Stationary\n".format(df.columns[i]))
else:
print("{}: Series is Stationary\n".format(df.columns[i]))
# make train data and test data
df_train = df[:int(0.8*(len(df)))]
df_test = df[int(0.8*(len(df))):]
# show dataframe for training
df_train
# show dataframe for test
df_test
# try 10 lags
lags = 10
model = VAR(df_train, freq="D")
for i in range(lags):
results = model.fit(i+1)
print("Order = ", i+1)
print("AIC = ", results.aic)
print("BIC = ", results.bic)
model.select_order(lags).summary()
# from the result above, the lowest BIC is from the lag 1
# and the lowest AIC is from the lag 6
model = VAR(df_train, freq="D")
results = model.fit(7)
results.summary()
# using lag 7
lag = results.k_ar
print("Lag:",lag)
# +
# MAPE
def mean_absolute_percentage_error(y_true, y_pred):
return (np.mean(np.abs((y_pred - y_true) / y_true)) * 100) / 100
# SMAPE
def symmetric_mean_absolute_percentage_error(y_true, y_pred):
return (2.0 * np.mean(np.abs(y_pred - y_true) / (np.abs(y_pred) + np.abs(y_true))) * 100) / 100
# +
# result for model VAR Only
y_test = df_test
y_predict = results.forecast(df_train.values[-lag:], steps=df_test.shape[0])
mape = mean_absolute_percentage_error(y_test, y_predict)
print("MAPE:",mape,"\n")
smape = symmetric_mean_absolute_percentage_error(y_test, y_predict)
print("SMAPE:",smape,"\n")
mae = mean_absolute_error(y_test,y_predict)
print("MAE :",mae)
mse = mean_squared_error(y_test,y_predict)
print("MSE :",mse)
rmse = np.sqrt(mean_squared_error(y_test,y_predict))
print("RMSE:",rmse)
print("R2 :",r2_score(y_test,y_predict))
# +
# convert dataframe into numpy array
df_arr = df.values
df_arr = df_arr.astype('float32')
#scaler = MinMaxScaler(feature_range=(0,1))
#df = scaler.fit_transform(df)
# split dataset into train and test
train_size = int(len(df_arr) * 0.80)
test_size = len(df_arr) - train_size
train, test = df_arr[0:train_size,:] , df_arr[train_size:len(df_arr),:]
print("Train Data:",len(train))
print("Test Data :",len(test))
# +
def to_sequences(df, seq_size=1):
x = []
y = []
for i in range(len(df)-seq_size-1):
window = df[i:(i+seq_size),0]
x.append(window)
y.append(df[i+seq_size,0])
return np.array(x), np.array(y)
seq_size = 5
trainX, trainY = to_sequences(train, seq_size)
testX, testY = to_sequences(test, seq_size)
# -
pd.DataFrame(trainX)
pd.DataFrame(trainY)
# +
print("Building model...")
model = Sequential()
model.add(Dense(64, input_dim = seq_size, activation='relu')) #12
model.add(Dense(32, activation='relu')) #8
model.add(Dense(1))
model.compile(loss='mean_squared_error', optimizer='adam', metrics = ['acc'])
print(model.summary())
# -
# start validation data
result = model.fit(trainX, trainY, validation_data=(testX, testY), verbose=2, epochs=1000)
# +
loss_values = result.history['loss']
epochs = range(1, len(loss_values)+1)
plt.plot(epochs, loss_values, label='Training Loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
# -
# make predictions
trainPredict = model.predict(trainX)
testPredict = model.predict(testX)
#plt.plot(trainX)
#plt.plot(trainPredict)
#plt.show()
# preprocess data for PSO
X_train, X_test, y_train, y_test, exercise = preprocess_data(df)
# +
# PSO-tuned Neural Net Model
def calc_nn_params(n_inputs, n_hidden, n_classes):
i_weights = n_inputs*n_hidden
i_bias = n_hidden
h_weights = n_hidden*n_classes
h_bias = n_classes
n_params = i_weights + i_bias + h_weights + h_bias
return i_weights, i_bias, h_weights, h_bias,n_params
# neural net architecture
n_inputs = 1+len(exercise)
n_hidden = 8
n_classes = 1
i_weights, i_bias, h_weights, h_bias, n_params = calc_nn_params(n_inputs, n_hidden, n_classes)
# -
# ensure reproducibility
checkpoint_state = np.random.get_state()
SWARM_SIZE = 30
# %%time
# initial model PSO
np.random.set_state(checkpoint_state)
options = {'c1':0.5,'c2':2.0,'w':1.0, 'k':SWARM_SIZE, 'p':1}
cost, pos, history = train_model(options)
plot_history(history)
# +
#print(f'MSE on validation set: {mean_squared_error(prediction(X_test, pos), y_test)}')
# +
# result for model VAR-NN-PSO
y_test = df_test
y_predict = results.forecast(df_train.values[-lag:], steps=df_test.shape[0])
mape2 = mean_absolute_percentage_error(y_test, y_predict)
print("MAPE:",(mape2/r),"\n")
smape2 = symmetric_mean_absolute_percentage_error(y_test, y_predict)
print("SMAPE:",(smape2/r),"\n")
mae2 = mean_absolute_error(y_test,y_predict)
print("MAE :",(mae2/r))
mse2 = mean_squared_error(y_test,y_predict)
print("MSE :",(mse2/r))
rmse2 = np.sqrt(mean_squared_error(y_test,y_predict))
print("RMSE:",(rmse2/r))
print("R2 :",(r2_score(y_test,y_predict)/r))
# +
# final result for prediction
trainScore = np.sqrt(mean_squared_error(trainY, trainPredict))
print("Train Score: %.2f RMSE"% (trainScore))
testScore = np.sqrt(mean_squared_error(testY, testPredict))
print("Test Score: %.2f RMSE"% (testScore))
mape3 = mean_absolute_percentage_error(trainPredict, trainY)
print("MAPE:",mape3)
smape3 = symmetric_mean_absolute_percentage_error(trainPredict, trainY)
print("SMAPE:",smape3)
mae3 = mean_absolute_error(trainPredict, trainY)
print("MAE :",mae3)
mse3 = mean_squared_error(trainPredict, trainY)
print("MSE :",mse3)
rmse3 = np.sqrt(mean_squared_error(trainPredict, trainY))
print("RMSE:",rmse3)
print("R2 :",r2_score(trainPredict, trainY))
# -
# generate forecast for next 10 days
data = np.array(results.forecast(df_train.values[-lag:], steps=10))
forecast_output = pd.DataFrame(data=data, columns=['tondano', 'winangun'])
forecast_output['days'] = pd.DataFrame(data=[1,2,3,4,5,6,7,8,9,10])
forecast_output = forecast_output[['days', 'tondano', 'winangun']]
print("10 Days Forecasts:\n=============================")
print(forecast_output.to_string(index=False))
plt.plot(forecast_output['tondano'], label='Tondano')
plt.plot(forecast_output['winangun'], label='Winangun')
plt.title("10 Days Forecasts")
plt.legend()
plt.show()
# write forecast output to csv
forecast_output.to_csv(r'forecast_output.csv', index = False)
| model_var_nn_pso.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# #!/usr/bin/env python
# -*- coding: utf-8 -*-
# +
from cognipy.ontology import Ontology #the ontology processing class
from cognipy.ontology import CQL #SPARQL format tailored for Contolled Natural Language
from cognipy.ontology import encode_string_for_graph_label #complex datatypes encoder for the graph labels in graph visualisation
import textwrap
def graph_attribute_formatter(val):
if isinstance(val,list) or isinstance(val,set):
return " | ".join(list(map(lambda i:encode_string_for_graph_label(graph_attribute_formatter(i)),val)))
elif isinstance(val,dict):
return " | ".join(list(map(lambda i:i[0]+" : "+encode_string_for_graph_label(graph_attribute_formatter(i[1])),val.items())))
else:
return encode_string_for_graph_label(textwrap.fill(str(val),40))
# +
# %%writefile RDF_example.encnl
Comment: " Not needed, only used to get a draw display
Every class-string is a superclass.
Every class-integer is a superclass.
Every class-double is a superclass.
Every class-float is a superclass.
Every class-boolean is a superclass.
Every class-duration is a superclass.
Every class-datetime is a superclass.
Every class-real is a superclass.
Every class-decimal is a superclass.
Every class-decimal has-account-id nothing-but (some decimal value).
".
Every class-decimal has-account-id nothing-but (some decimal value).
Every class-double has-account-id nothing-but (some double value).
Every class-float has-account-id nothing-but (some float value).
Every class-string has-account-id nothing-but (some string value).
Every class-integer has-account-id nothing-but (some integer value).
Every class-boolean has-account-id nothing-but (some boolean value).
Every class-duration has-account-id nothing-but (some duration value).
Every class-datetime has-account-id nothing-but (some datetime value).
Every class-real has-account-id nothing-but (some real value).
# -
onto=Ontology("cnl/file","RDF_example.encnl",
evaluator = lambda e:eval(e,globals(),locals()),
graph_attribute_formatter = graph_attribute_formatter)
onto.draw_graph()
old_cnl = onto.as_cnl()
print(onto.as_cnl())
# +
outputfilename = 'cnl_RDF_example.encnl'
file = open(outputfilename,"w", encoding="utf8")
file.write(onto.as_cnl())
file.close()
# -
onto_cnl = Ontology("cnl/file","cnl_RDF_example.encnl")
# +
outputfilename = './cnl_RDF_example.rdf'
file = open(outputfilename,"w", encoding="utf8")
file.write(onto.as_rdf())
file.close()
# -
onto_rdf = Ontology("rdf/file","./cnl_RDF_example.rdf")
print(onto_rdf.as_cnl())
# I see that the decimal class behaves differently (thanks!!), but at the moment it seems to be 'some integer value' ?
#
# double and float still look like they are not handled?
#
# Every value-of double is something (some value).
# Every value-of float is something (some value).
#
# real treated as (greater-or-equal-to 0 or lower-or-equal-to 0)? Is this a correct implementation?
#
# Every classi has-account-id nothing-but (greater-or-equal-to 0 or lower-or-equal-to 0).
| docsrc/jupyter_notebooks/BugReports/datatype_tests.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="images/logo.jpg" style="display: block; margin-left: auto; margin-right: auto;" alt="לוגו של מיזם לימוד הפייתון. נחש מצויר בצבעי צהוב וכחול, הנע בין האותיות של שם הקורס: לומדים פייתון. הסלוגן המופיע מעל לשם הקורס הוא מיזם חינמי ללימוד תכנות בעברית.">
# # <p style="text-align: right; direction: rtl; float: right;">רשימות</p>
# ## <p style="text-align: right; direction: rtl; float: right; clear: both;">הגדרה</p>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# רשימה, כשמה כן היא, מייצגת <mark>אוסף מסודר של ערכים</mark>. רשימות יהיו סוג הנתונים הראשון שנכיר בפייתון, ש<mark>מטרתו היא לקבץ ערכים</mark>.<br>
# הרעיון מוכר לנו מהיום־יום: רשימת פריטים לקנייה בסופר שמסודרת לפי הא–ב, או רשימת ההופעות בקיץ הקרוב המסודרת לפי תאריך.
# </p>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# נסו לדמיין רשימה כמסדרון ארוך, שבו עומדים בתור אחד אחרי השני איברים מסוגים שאנחנו מכירים בפייתון.<br>
# אם נשתמש בדימוי הלייזרים שנתנו למשתנים בשבוע הקודם, אפשר להגיד שמדובר בלייזר שמצביע לשורת לייזרים, שבה כל לייזר מצביע על ערך כלשהו.
# </p>
# <table style="font-size: 2rem; border: 0px solid black; border-spacing: 0px;">
# <tr>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-bottom: 1px solid;">0</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">1</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">2</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">3</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">4</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">5</td>
# </tr>
# <tbody>
# <tr>
# <td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"<NAME>"</td>
# <td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"<NAME>"</td>
# <td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"<NAME>"</td>
# <td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"<NAME>"</td>
# <td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"<NAME>"</td>
# <td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"<NAME>"</td>
# </tr>
# <tr style="background: #f5f5f5;">
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right;">-6</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-5</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-4</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-3</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-2</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-1</td>
# </tr>
# </tbody>
# </table>
#
# <br>
#
# <p style="text-align: center; direction: rtl; clear: both; font-size: 1.8rem">
# דוגמה לרשימה: 6 ראשי הממשלה הראשונים בישראל לפי סדר כהונתם, משמאל לימין
# </p>
# ## <p style="text-align: right; direction: rtl; float: right;">דוגמאות</p>
# <ol style="text-align: right; direction: rtl; float: right; clear: both;">
# <li>רשימת שמות ראשי הממשלה במדינת ישראל לפי סדר כהונתם.</li>
# <li>רשימת הגילים של התלמידים בכיתה, מהמבוגר לצעיר.</li>
# <li>רשימת שמות של התקליטים שיש לי בארון, מסודרת מהתקליט השמאלי לימני.</li>
# <li>רשימה שבה כל איבר מייצג אם לראש הממשלה שנמצא בתא התואם ברשימה הקודמת היו משקפיים.</li>
# <li>האיברים 42, 8675309, 73, <span dir="ltr" style="direction: ltr;">-40</span> ו־186282 בסדר הזה.</li>
# <li>רשימה של תחזית מזג האוויר ב־7 הימים הקרובים. כל איבר ברשימה הוא בפני עצמו רשימה, שמכילה שני איברים: הראשון הוא מה תהיה הטמפרטורה הממוצעת, והשני הוא מה תהיה הלחות הממוצעת.</li>
# <ol>
# <div class="align-center" style="display: flex; text-align: right; direction: rtl; clear: both;">
# <div style="display: flex; width: 10%; float: right; clear: both;">
# <img src="images/exercise.svg" style="height: 50px !important;" alt="תרגול">
# </div>
# <div style="width: 90%">
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# <strong>תרגול</strong>:
# הרשימות שהוצגו למעלה הן <dfn>רשימות הומוגניות</dfn>, כאלו שכל האיברים שבהן הם מאותו סוג.<br>
# כתבו עבור כל אחת מהרשימות שהוצגו בדוגמה מה סוג הנתונים שיישמר בהן.
# </p>
# </div>
# </div>
# <div class="align-center" style="display: flex; text-align: right; direction: rtl; clear: both;">
# <div style="display: flex; width: 10%; float: right; clear: both;">
# <img src="images/exercise.svg" style="height: 50px !important;" alt="תרגול">
# </div>
# <div style="width: 90%">
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# <strong>תרגול</strong>:
# נסו לתת דוגמה לעוד 3 רשימות שבהן נתקלתם לאחרונה.</p>
# </div>
# </div>
# ## <p style="text-align: right; direction: rtl; float: right;">רשימות בקוד</p>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# רשימות הן אחד מסוגי הנתונים הכיפיים ביותר בפייתון, וזאת בזכות הגמישות האדירה שיש לנו בתכנות עם רשימות.
# </p>
# ### <span style="text-align: right; direction: rtl; float: right; clear: both;">הגדרת רשימה</span>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# נגדיר בעזרת פייתון את הרשימה שפגשנו למעלה – 6 ראשי הממשלה הראשונים מאז קום המדינה:
# </p>
prime_ministers = ['<NAME>', '<NAME>', '<NAME>', '<NAME>', '<NAME>', '<NAME>']
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# מה התרחש בקוד?<br>
# התחלנו את הגדרת הרשימה באמצעות התו <code dir="ltr" style="direction: ltr;">[</code>.<br>
# מייד אחרי התו הזה הכנסנו איברים לרשימה לפי הסדר הרצוי, כאשר כל איבר מופרד ממשנהו בפסיק (<code>,</code>).<br>
# במקרה שלנו, כל איבר הוא מחרוזת המייצגת ראש ממשלה. הכנסנו את ראשי הממשלה לרשימה <mark>לפי סדר</mark> כהונתם.<br>
# שימו לב שהרשימה מכילה איבר מסוים פעמיים – מכאן ש<mark>רשימה היא מבנה נתונים שתומך בחזרות</mark>.<br>
# לסיום, נסגור את הגדרת הרשימה באמצעות התו <code dir="ltr" style="direction: ltr;">]</code>.<br>
# </p>
print(prime_ministers)
type(prime_ministers)
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# נוכל להגדיר רשימה של המספרים הטבעיים עד 7:
# </p>
numbers = [1, 2, 3, 4, 5, 6, 7]
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# <dfn>רשימה הומוגנית</dfn> היא רשימה שבה האיברים שנמצאים בכל אחד מהתאים הם מאותו סוג. רשימות "בעולם האמיתי" הן בדרך כלל הומוגניות.<br>
# <dfn>רשימה הטרוגנית</dfn> היא רשימה שבה איברים בתאים שונים יכולים להיות מסוגים שונים.<br>
# ההבדל הוא סמנטי בלבד, ופייתון לא מבדילה בין רשימה הטרוגנית לרשימה הומוגנית.
# </p>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# לשם הדוגמה, נגדיר רשימה הטרוגנית:
# </p>
wtf = ['The cake is a', False, 42]
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# נוכל אפילו להגדיר רשימה ריקה, שבה אין איברים כלל:</p>
empty_list = []
# ### <p style="text-align: right; direction: rtl; float: right;">גישה לאיברי הרשימה</p>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# לכל תא ברשימה יש מספר, שמאפשר לנו להתייחס לאיבר שנמצא באותו תא.<br>
# הדבר דומה ללייזר שעליו יש מדבקת שם ("שמות ראשי ממשלה"), והוא מצביע על שורת לייזרים שעל התווית שלהם מופיע מספר המתאר את מיקומם בשורה.<br>
# התא השמאלי ביותר ברשימה ממוספר כ־0, התא שנמצא אחריו (מימינו) מקבל את המספר 1, וכך הלאה עד לסוף הרשימה.<br>
# המספור של כל תא נקרא <dfn>המיקום שלו ברשימה</dfn>, או <dfn>האינדקס שלו</dfn>.
# </p>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# נגדיר את רשימת שמות התקליטים שיש לי בבית:
# </p>
# Index 0 1 2 3 4 5
vinyls = ['Ecliptica', 'GoT Season 6', 'Lone Digger', 'Everything goes numb', 'Awesome Mix Vol. 1', 'Ultimate Sinatra']
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# בהנחה שאנחנו מתים על Guardians of the Galaxy, נוכל לנסות להשיג מהרשימה את Awesome Mix Vol. 1.<br>
# כדי לעשות זאת, נציין את שם הרשימה שממנה אנחנו רוצים לקבל את האיבר, ומייד לאחר מכן את מיקומו ברשימה בסוגריים מרובעים.
# </p>
print(vinyls[4])
# <div class="align-center" style="display: flex; text-align: right; direction: rtl; clear: both;">
# <div style="display: flex; width: 10%; float: right; clear: both;">
# <img src="images/warning.png" style="height: 50px !important;" alt="אזהרה!">
# </div>
# <div style="width: 90%">
# <p style="text-align: right; direction: rtl; clear: both;">
# התא הראשון ממוספר 0, ולא 1.<br>
# יש לכך סיבות טובות, אבל פעמים רבות תרגישו שהמספור הזה לא טבעי ועלול ליצור <dfn>באגים</dfn>, שהם קטעי קוד שמתנהגים אחרת משציפה המתכנת.<br>
# כפועל יוצא, המיקום ברשימה של התא האחרון לא יהיה כאורך הרשימה, אלא כאורך הרשימה פחות אחד.<br>
# משמע: ברשימה שבה 3 איברים, מספרו של התא האחרון יהיה 2.
# </p>
# </div>
# </div>
# <figure>
# <img src="images/list-of-vinyls.png" width="100%" style="display: block; margin-left: auto; margin-right: auto;" alt="תמונה של 6 תקליטים על שטיח. משמאל לימין: Ecliptica / Sonata Arctica, Game of Thrones Season 6 / Ramin Djawadi, Caravan Palace / Lone Digger, Everything goes numb / Streetlight Manifesto, Awesome Mix Vol. 1 / Guardians of the Galaxy, Ultimate Sinatra / Frank Sinatra. מעל כל דיסק מופיע מספר, מ־0 עבור התקליט השמאלי ועד 5 עבור התקליט הימני. מתחת לתקליטים מופיע המספר -1 עבור התקליט הימני ביותר, וכך הלאה עד -5 עבור התקליט השמאלי ביותר.">
# <figcaption style="text-align: center; direction: rtl; clear: both;">
# רשימת (חלק מ)התקליטים בארון שלי, מסודרת מהתקליט השמאלי לימני.<br>
# </figcaption>
# </figure>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# כפי שניתן לראות בתמונה, פייתון מנסה לעזור לנו ומאפשרת לנו לגשת לאיברים גם מהסוף.<br>
# חוץ מהמספור הרגיל שראינו קודם, אפשר לגשת לאיברים מימין לשמאל באמצעות מספור שלילי.<br>
# האיבר האחרון יקבל את המספר <span style="direction: ltr" dir="ltr">-1</span>, זה שלפניו (משמאלו) יקבל <span style="direction: ltr" dir="ltr">-2</span> וכן הלאה.
# </p>
# 0 1 2 3 4 5
vinyls = ['Ecliptica', 'GoT Season 6', 'Lone Digger', 'Everything goes number', 'Awesome Mix Vol. 1', 'Ultimate Sinatra']
# -6 -5 -4 -3 -2 -1
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# אם נרצה לגשת שוב לאותו דיסק, אבל הפעם מהסוף, נוכל לכתוב זאת כך:
# </p>
print(vinyls[-2])
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# כדאי לזכור שהתוכן של כל אחד מהתאים הוא ערך לכל דבר.<br>
# יש לו סוג, ואפשר לבצע עליו פעולות כמו שלמדנו עד עכשיו:
# </p>
type(vinyls[0])
print(vinyls[0] + ', By Sonata Arctica')
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# לסיום, נראה שבדיוק כמו במחרוזות, נוכל לבדוק את אורך הרשימה על ידי שימוש בפונקציה <code>len</code>.
# </p>
# כמה תקליטים יש לי?
len(vinyls)
# <div class="align-center" style="display: flex; text-align: right; direction: rtl; clear: both;">
# <div style="display: flex; width: 10%; float: right; clear: both;">
# <img src="images/warning.png" style="height: 50px !important;" alt="אזהרה!">
# </div>
# <div style="width: 90%">
# <p style="text-align: right; direction: rtl; clear: both;">
# אם ננסה לגשת לתא שאינו קיים, נקבל <code>IndexError</code>.<br>
# זה בדרך כלל קורה כשאנחנו שוכחים להתחיל לספור מ־0.<br>
# אם השגיאה הזו מופיעה כשאתם מתעסקים עם רשימות, חשבו איפה בקוד פניתם לתא שאינו קיים.
# </p>
# </div>
# </div>
# ### <p style="text-align: right; direction: rtl; float: right;">השמה ברשימות</p>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# לפעמים נרצה לשנות את הערך של האיברים ברשימה.<br>
# נפנה ללייזר מסוים בשורת הלייזרים שלנו, ונבקש ממנו להצביע לערך חדש:
# </p>
print(vinyls)
vinyls[1] = 'GoT Season 7'
print(vinyls)
# ### <p style="text-align: right; direction: rtl; float: right;">אופרטורים חשבוניים על רשימות</p>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# אופרטורים שהכרנו כשלמדנו על מחרוזות, יעבדו נהדר גם על רשימות.
# </p>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# כפי ש־<code>+</code> משרשר בין מחרוזות, הוא יודע לשרשר גם בין רשימות:
# </p>
[1, 2, 3] + [4, 5, 6]
['a', 'b', 'c'] + ['easy', 'as'] + [1, 2, 3]
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# וכפי ש־<code>*</code> משרשר מחרוזת לעצמה כמות מסוימת של פעמים, כך הוא יפעל גם עם רשימות:
# </p>
['wake up', 'go to school', 'sleep'] * 365
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# אפשר גם לשלב:
# </p>
['Is', 'someone', 'getting'] + ['the', 'best,'] * 4 + ['of', 'you?']
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# שימו לב שכל אופרטור שתשימו ליד הרשימה מתייחס <em>לרשימה בלבד</em>, ולא לאיברים שבתוכה.<br>
# משמע <code dir="ltr" style="direction: ltr;">+ 5</code> לא יוסיף לכם 5 לכל אחד מהאיברים, אלא ייכשל כיוון שפייתון לא יודעת לחבר רשימה למספר שלם.<br>
# </p>
[1, 2, 3] + 5
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# שימו לב גם שהפעלת אופרטור על רשימה לא גורמת לשינוי הרשימה, אלא רק מחזירה ערך.<br>
# כדי לשנות ממש את הרשימה, נצטרך להשתמש בהשמה:
# </p>
prime_ministers = ['<NAME>', '<NAME>', '<NAME>', '<NAME>', '<NAME>', '<NAME>']
print(prime_ministers)
prime_ministers + ['<NAME>']
print(prime_ministers)
print(prime_ministers)
prime_ministers = prime_ministers + ['<NAME>']
print(prime_ministers)
# ### <p style="text-align: right; direction: rtl; float: right;">אופרטורים השוואתיים על רשימות</p>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# נגדיר את רשימת האנשים שנכחו בכיתה ביום ראשון, שני, שלישי ורביעי:
# </p>
pupils_in_sunday = ['Moshe', 'Dukasit', 'Michelangelo']
pupils_in_monday = ['Moshe', 'Dukasit', 'Master Splinter']
pupils_in_tuesday = ['Moshe', 'Dukasit', 'Michelangelo']
pupils_in_wednesday = ['Moshe', 'Dukasit', 'Michelangelo', 'Master Splinter']
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# רשימות תומכות בכל אופרטורי ההשוואה שלמדנו עד כה.<br>
# נתחיל בקל ביותר. בואו נבדוק באיזה יום הרכב התלמידים בכיתה היה זהה להרכב התלמידים שהיה בה ביום ראשון:
# </p>
print("Is it Monday? " + str(pupils_in_sunday == pupils_in_monday))
print("Is it Tuesday? " + str(pupils_in_sunday == pupils_in_tuesday))
print("Is it Wednesday? " + str(pupils_in_sunday == pupils_in_wednesday))
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# האם משה נכח בכיתה ביום שלישי?
# </p>
print('Moshe' in pupils_in_tuesday)
# זה אותו דבר כמו:
print('Moshe' in ['Moshe', 'Dukasit', 'Michelangelo'])
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# נוכיח שמאסטר ספלינטר הבריז באותו יום:
# </p>
'Master Splinter' not in pupils_in_tuesday
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# ולסיום, בואו נבדוק איזו גרסה חדשה יותר:
# </p>
python_new_version = [3, 7, 2]
python_old_version = [2, 7, 16]
print(python_new_version > python_old_version)
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# כדי לבצע השוואה בין רשימות, פייתון מנסה להשוות את האיבר הראשון מהרשימה הראשונה לאיבר הראשון מהרשימה השנייה.<br>
# אם יש "תיקו", היא תעבור לאיבר השני בכל רשימה, כך עד סוף הרשימה.
# </p>
# ### <p style="text-align: right; direction: rtl; float: right;">רשימה של רשימות</p>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# לעיתים דברים בחיים האמיתיים הם מורכבים מדי מכדי לייצג אותם ברשימה סטנדרטית.<br>
# הרבה פעמים נשים לב שיוקל לנו אם ניצור רשימה שבה כל תא הוא רשימה בפני עצמו.<br>הרעיון הזה ייצור לנו רשימה של רשימות.<br>
# ניקח לדוגמה את הרשימות שהגדרנו למעלה, שמתארות מי נכח בכל יום בכיתה:
# </p>
pupils_in_sunday = ['Moshe', 'Dukasit', 'Michelangelo']
pupils_in_monday = ['Moshe', 'Dukasit', 'Splinter']
pupils_in_tuesday = ['Moshe', 'Dukasit', 'Michelangelo']
pupils_in_wednesday = ['Moshe', 'Dukasit', 'Michelangelo', 'Splinter']
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# אנחנו רואים לפנינו רשימה של ימים, שקל להכניס לרשימה אחת גדולה:
# </p>
pupils = [pupils_in_sunday, pupils_in_monday, pupils_in_tuesday, pupils_in_wednesday]
print(pupils)
# <table style="font-size: 1rem; border: 0px solid black; border-spacing: 0px;">
# <tr>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-bottom: 1px solid;">0</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">1</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">2</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">3</td>
# </tr>
# <tbody>
# <tr>
# <td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">
# <table style="font-size: 1.1rem; border: 0px solid black; border-spacing: 0px;">
# <tr>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-bottom: 1px solid;">0</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">1</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">2</td>
# </tr>
# <tbody>
# <tr>
# <td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"Moshe"</td>
# <td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"Dukasit"</td>
# <td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"Michelangelo"</td>
# </tr>
# <tr style="background: #f5f5f5;">
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right;">-3</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-2</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-1</td>
# </tr>
# </tbody>
# </table>
# </td>
# <td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">
# <table style="font-size: 1.1rem; border: 0px solid black; border-spacing: 0px;">
# <tr>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-bottom: 1px solid;">0</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">1</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">2</td>
# </tr>
# <tbody>
# <tr>
# <td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"Moshe"</td>
# <td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"Dukasit"</td>
# <td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"Splinter"</td>
# </tr>
# <tr style="background: #f5f5f5;">
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right;">-3</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-2</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-1</td>
# </tr>
# </tbody>
# </table>
# </td>
# <td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">
# <table style="font-size: 1.1rem; border: 0px solid black; border-spacing: 0px;">
# <tr>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-bottom: 1px solid;">0</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">1</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">2</td>
# </tr>
# <tbody>
# <tr>
# <td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"Moshe"</td>
# <td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"Dukasit"</td>
# <td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"Michelangelo"</td>
# </tr>
# <tr style="background: #f5f5f5;">
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right;">-3</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-2</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-1</td>
# </tr>
# </tbody>
# </table>
# </td>
# <td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">
# <table style="font-size: 1.1rem; border: 0px solid black; border-spacing: 0px;">
# <tr>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-bottom: 1px solid;">0</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">1</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">2</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">3</td>
# </tr>
# <tbody>
# <tr>
# <td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"Moshe"</td>
# <td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"Dukasit"</td>
# <td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"Michelangelo"</td>
# <td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">"Splinter"</td>
# </tr>
# <tr style="background: #f5f5f5;">
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right;">-4</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-3</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-2</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-1</td>
# </tr>
# </tbody>
# </table>
# </td>
# </tr>
# <tr style="background: #f5f5f5;">
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right;">-4</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-3</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-2</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-1</td>
# </tr>
# </tbody>
# </table>
#
# <br>
#
# <p style="text-align: center; direction: rtl; clear: both; font-size: 1.8rem">
# דוגמה לרשימה של רשימות: נוכחות התלמידים בימי ראשון עד רביעי
# </p>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# השורה שכתבנו למעלה זהה לחלוטין לשורה הבאה, שבה אנחנו מגדירים רשימה אחת שכוללת את רשימות התלמידים שנכחו בכיתה בכל יום.
# </p>
pupils = [['Moshe', 'Dukasit', 'Michelangelo'], ['Moshe', 'Dukasit', 'Splinter'], ['Moshe', 'Dukasit', 'Michelangelo'], ['Moshe', 'Dukasit', 'Michelangelo', 'Splinter']]
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# נוכל לקבל את רשימת התלמידים שנכחו ביום ראשון בצורה הבאה:
# </p>
pupils[0]
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# ואת התלמיד האחרון שנכח ביום ראשון בצורה הבאה:
# </p>
pupils_in_sunday = pupils[0]
print(pupils_in_sunday[-1])
# או פשוט:
print(pupils[0][-1])
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# אם קשה לכם לדמיין את זה, עשו זאת בשלבים.<br>
# בדקו מה יש ב־<code>pupils</code>, אחרי זה מה מחזיר <code>pupils[0]</code>, ואז נסו לקחת ממנו את האיבר האחרון, <code>pupils[0][-1]</code>.
# </p
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# כדי להבין טוב יותר איך רשימה של רשימות מתנהגת, חשוב להבין את התוצאות של הביטויים הבוליאניים הבאים.<br>
# זה קצת מבלבל, אבל אני סומך עליכם שתחזיקו מעמד:
# </p>
print("pupils = " + str(pupils))
print("-" * 50)
print("1. 'Moshe' in pupils == " + str('Moshe' in pupils))
print("2. 'Moshe' in pupils[0] == " + str('Moshe' in pupils[0]))
print("3. ['Moshe', 'Splinter'] in pupils == " + str(['Moshe', 'Splinter'] in pupils))
print("4. ['Moshe', 'Splinter'] in pupils[0] == " + str(['Moshe', 'Splinter'] in pupils[-1]))
print("5. ['Moshe', 'Dukasit', 'Splinter'] in pupils == " + str(['Moshe', 'Dukasit', 'Splinter'] in pupils))
print("6. ['Moshe', 'Dukasit', 'Splinter'] in pupils[0] == " + str(['Moshe', 'Dukasit', 'Splinter'] in pupils[0]))
# <ol style="text-align: right; direction: rtl; float: right; clear: both;">
# <li>הביטוי הבוליאני בשורה 1 מחזיר <samp>False</samp>, כיוון שכל אחד מהאיברים ברשימה <var>pupils</var> הוא רשימה, ואף אחד מהם אינו המחרוזת <em>"Moshe"</em>.</li>
# <li>הביטוי הבוליאני בשורה 2 מחזיר <samp>True</samp>, כיוון שהאיבר הראשון ב־<var>pupils</var> הוא רשימה שמכילה את המחרוזת <em>"Moshe"</em>.</li>
# <li>הביטוי הבוליאני בשורה 3 מחזיר <samp>False</samp>, כיוון שאין בתוך <var>pupils</var> רשימה שאלו בדיוק הערכים שלה. יש אומנם רשימה שמכילה את האיברים האלו, אבל השאלה הייתה האם הרשימה הגדולה (<var>pupils</var>) מכילה איבר ששווה בדיוק ל־<code>['Moshe', 'Splinter']</code>.</li>
# <li>הביטוי הבוליאני בשורה 4 מחזיר <samp>False</samp>, כיוון שברשימה האחרונה בתוך <var>pupils</var> אין איבר שהוא הרשימה <code>["Moshe", "Splinter"]</code>.</li>
# <li>הביטוי הבוליאני בשורה 5 מחזיר <samp>True</samp>, כיוון שיש רשימה ישירות בתוך <var>pupils</var> שאלו הם ערכיה.</li>
# <li>הביטוי הבוליאני בשורה 6 מחזיר <samp>False</samp>, כיוון שברשימה הראשונה בתוך <var>pupils</var> אין איבר שהוא הרשימה הזו.</li>
# </ol>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# זכרו שעבור פייתון אין שום דבר מיוחד ברשימה של רשימות. היא בסך הכול רשימה רגילה, שכל אחד מאיבריה הוא רשימה.<br>
# מבחינתה אין הבדל בין רשימה כזו לכל רשימה אחרת.
# </p>
# ## <p style="text-align: right; direction: rtl; float: right;">המונח Iterable</p>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# באתרי אינטרנט ובתיעוד של פייתון אנחנו נפגש פעמים רבות עם המילה <dfn>Iterable</dfn>.<br>
# בקורס נשתמש במונח הזה פעמים רבות כדי להבין טוב יותר איך פייתון מתנהגת.<br>
# <mark>נגדיר ערך כ־<dfn>iterable</dfn> אם ניתן לפרק אותו לכלל האיברים שלו.</mark><br>
# </p>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# עד כה אנחנו מכירים 2 סוגי משתנים שעונים להגדרה iterables: רשימות ומחרוזות.<br>
# ניתן לפרק רשימה לכל האיברים שמרכיבים אותה, וניתן לפרק מחרוזת לכל התווים שמרכיבים אותה.<br>
# </p>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# יש הרבה במשותף לכל הדברים שניתן להגיד עליהם שהם iterables:<br>
# על חלק גדול מה־iterables אפשר להפעיל פעולות שמתייחסות לכלל האיברים שבהם, כמו <code>len</code> שמראה את מספר האיברים בערך.<br>
# על חלק גדול מה־iterables יהיה אפשר גם להשתמש בסוגריים מרובעים כדי לגשת לאיבר מסוים שנמצא בהם.<br>
# בעתיד נלמד על עוד דברים שמשותפים לרוב (או לכל) ה־iterables.
# </p>
# ## <p style="align: right; direction: rtl; float: right; clear: both;">מונחים</p>
# <dl style="text-align: right; direction: rtl; float: right; clear: both;">
# <dt>רשימה</dt><dd>סוג משתנה שמטרתו לקבץ ערכים אחרים בסדר מסוים.</dd>
# <dt>תא</dt><dd>מקום ברשימה שמכיל איבר כלשהו.</dd>
# <dt>מיקום</dt><dd>מיקום של תא מסוים הוא המרחק שלו מהתא הראשון ברשימה, שמיקומו הוא 0. זהו מספר שמטרתו לאפשר גישה לתא מסוים ברשימה.</dd>
# <dt>אינדקס</dt><dd>מילה נרדפת ל"מיקום".</dd>
# <dt>איבר</dt><dd>ערך שנמצא בתא של רשימה. ניתן לאחזר אותו אם נציין את שם הרשימה, ואת מיקום התא שבו הוא נמצא.</dd>
# <dt>רשימה הומוגנית</dt><dd>רשימה שבה כל האיברים הם מאותו סוג.</dd>
# <dt>רשימה הטרוגנית</dt><dd>רשימה שבה לכל איבר יכול להיות סוג שונה.</dd>
# <dt>Iterable</dt><dd>ערך שמורכב מסדרה של ערכים אחרים.</dd>
# </dl>
# ## <p style="text-align: right; direction: rtl; float: right;">לסיכום</p>
# <ol style="text-align: right; direction: rtl; float: right; clear: both;">
# <li>מספר האיברים ברשימה יכול להיות 0 (רשימה ריקה) או יותר.</li>
# <li>לאיברים ברשימה יש סדר.</li>
# <li>כל איבר ברשימה ממוספר החל מהאיבר הראשון שממוספר 0, ועד האיבר האחרון שמספרו הוא אורך הרשימה פחות אחד.</li>
# <li>ניתן לגשת לאיבר גם לפי המיקום שלו וגם לפי המרחק שלו מסוף הרשימה, באמצעות התייחסות למיקום השלילי שלו.</li>
# <li>איברים ברשימה יכולים לחזור על עצמם.</li>
# <li>רשימה יכולה לכלול איברים מסוג אחד בלבד (<dfn>רשימה הומוגנית</dfn>) או מכמה סוגים שונים (<dfn>רשימה הטרוגנית</dfn>).</li>
# <li>אורך הרשימה יכול להשתנות במהלך ריצת התוכנית.</li>
# <ol>
# ## <p style="align: right; direction: rtl; float: right; clear: both;">תרגול</p>
# ### <p style="align: right; direction: rtl; float: right; clear: both;">סדר בבית המשפט!</p>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# כתבו קוד שיסדר את רשימת נשיאי בית המשפט לפי סדר אלפבתי.<br>
# זה אכן אמור להיות מסורבל מאוד. בעתיד נלמד לכתוב קוד מוצלח יותר לבעיה הזו.<br>
# השתמשו באינדקסים, ושמרו ערכים בצד במשתנים.
# </p>
judges = ['<NAME>', '<NAME>', '<NAME>', '<NAME>', '<NAME>']
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# בונוס: כתבו קטע קוד שבודק שהרשימה (שמכילה 5 איברים) אכן מסודרת.
# </p>
# ### <p style="align: right; direction: rtl; float: right; clear: both;">מה זה משובחה בכלל?</p>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# לפניכם רשימה של שמות טעמי גלידה שנמצאים בדוכן הגלידה השכונתי.<br>
# קבלו מהמשתמש קיפיק את הטעם האהוב עליו, והדפיסו למשתמש האם הטעם שלו נמכר בדוכן.
# </p>
#
ice_cream_flavours = ['chocolate', 'vanilla', 'pistachio', 'banana']
# ### <p style="align: right; direction: rtl; float: right; clear: both;">מה רש"י?</p>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# לפניכם כמה ביטויים.<br>
# רשמו לעצמכם מה תהיה תוצאת כל ביטוי, ורק אז הריצו אותו.
# </p>
rabanim = ['Rashi', 'Maimonides', 'Nachmanides', '<NAME>']
'Rashi' in rabanim
'RASHI' in rabanim
['Rashi'] in rabanim
['Rashi', 'Nachmanides'] in rabanim
'Bruria' in rabanim
rabanim + ['<NAME>']
'<NAME>' in rabanim
'3' in [1, 2, 3]
(1 + 5 - 3) in [1, 2, 3]
[1, 5, 3] > [1, 2, 3]
rabanim[0] in [rabanim[0] + rabanim[1]]
rabanim[0] in [rabanim[0]] + [rabanim[1]]
rabanim[-1] == rabanim[0] or rabanim[-1] == rabanim[1] or rabanim[-1] == rabanim[2] or rabanim[-1] == rabanim[3]
rabanim[-1] == rabanim[0] or rabanim[-1] == rabanim[1] or rabanim[-1] == rabanim[2] and rabanim[-1] != rabanim[3]
rabanim[-1] == rabanim[0] or rabanim[-1] == rabanim[1] or rabanim[-1] == rabanim[2] and rabanim[-1] == rabanim[3]
1 in [[1, 2, 3], [4, 5, 6], [7, 8, 9]]
[1, 2, 3] in [[1, 2, 3], [4, 5, 6], [7, 8, 9]]
[[1, 2, 3], [4, 5, 6], [7, 8, 9]][0][2]
[[1, 2, 3], [4, 5, 6], [7, 8, 9]][0][3]
[[1, 2, 3], [4, 5, 6], [7, 8, 9]][0][-1] * 5
[[[1, 2, 3], [4, 5, 6], [7, 8, 9]][0][-1]] * 5
[[1, 2, 3], [4, 5, 6], [7, 8, 9]][0][-1]
[[1, 2, 3], [4, 5, 6], [7, 8, 9]][0][-1] == [[7, 8, 9], [4, 5, 6], [1, 2, 3]][2][2]
[[1, 2, 3]] in [[1, 2, 3], [4, 5, 6], [7, 8, 9]]
[[1, 2, 3], [4, 5, 6]] in [[1, 2, 3], [4, 5, 6], [7, 8, 9]]
[[1, 2, 3], [4, 5, 6]] in [[[1, 2, 3], [4, 5, 6]], [7, 8, 9]]
| week02/4_Lists.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="http://hilpisch.com/tpq_logo.png" alt="The Python Quants" width="35%" align="right" border="0"><br>
# # Python for Finance (2nd ed.)
#
# **Mastering Data-Driven Finance**
#
# © Dr. <NAME> | The Python Quants GmbH
#
# <img src="http://hilpisch.com/images/py4fi_2nd_shadow.png" width="300px" align="left">
# # Trading Platform
# ## Risk Disclaimer
# <font size="-1">
# Trading forex/CFDs on margin carries a high level of risk and may not be suitable for all investors as you could sustain losses in excess of deposits. Leverage can work against you. Due to the certain restrictions imposed by the local law and regulation, German resident retail client(s) could sustain a total loss of deposited funds but are not subject to subsequent payment obligations beyond the deposited funds. Be aware and fully understand all risks associated with the market and trading. Prior to trading any products, carefully consider your financial situation and experience level. Any opinions, news, research, analyses, prices, or other information is provided as general market commentary, and does not constitute investment advice. FXCM & TPQ will not accept liability for any loss or damage, including without limitation to, any loss of profit, which may arise directly or indirectly from use of or reliance on such information.
# </font>
# ## Author Disclaimer
# The author is neither an employee, agent nor representative of FXCM and is therefore acting independently. The opinions given are their own, constitute general market commentary, and do not constitute the opinion or advice of FXCM or any form of personal or investment advice. FXCM assumes no responsibility for any loss or damage, including but not limited to, any loss or gain arising out of the direct or indirect use of this or any other content. Trading forex/CFDs on margin carries a high level of risk and may not be suitable for all investors as you could sustain losses in excess of deposits.
# ## Retrieving Tick Data
import time
import numpy as np
import pandas as pd
import datetime as dt
from pylab import mpl, plt
plt.style.use('seaborn')
mpl.rcParams['font.family'] = 'serif'
# %config InlineBackend.figure_format = 'svg'
from fxcmpy import fxcmpy_tick_data_reader as tdr
print(tdr.get_available_symbols())
start = dt.datetime(2018, 6, 25)
stop = dt.datetime(2018, 6, 30)
td = tdr('EURUSD', start, stop)
td.get_raw_data().info()
td.get_data().info()
td.get_data().head()
sub = td.get_data(start='2018-06-29 12:00:00',
end='2018-06-29 12:15:00')
sub.head()
sub['Mid'] = sub.mean(axis=1)
sub['SMA'] = sub['Mid'].rolling(1000).mean()
sub[['Mid', 'SMA']].plot(figsize=(10, 6), lw=0.75);
# plt.savefig('../../images/ch14/fxcm_plot_01.png')
# ## Retrieving Candles Data
from fxcmpy import fxcmpy_candles_data_reader as cdr
print(cdr.get_available_symbols())
start = dt.datetime(2018, 5, 1)
stop = dt.datetime(2018, 6, 30)
# `period` must be one of `m1`, `H1` or `D1`
period = 'H1'
candles = cdr('EURUSD', start, stop, period)
data = candles.get_data()
data.info()
data[data.columns[:4]].tail()
data[data.columns[4:]].tail()
data['MidClose'] = data[['BidClose', 'AskClose']].mean(axis=1)
data['SMA1'] = data['MidClose'].rolling(30).mean()
data['SMA2'] = data['MidClose'].rolling(100).mean()
data[['MidClose', 'SMA1', 'SMA2']].plot(figsize=(10, 6));
# plt.savefig('../../images/ch14/fxcm_plot_02.png')
# ## Connecting to the API
import fxcmpy
fxcmpy.__version__
api = fxcmpy.fxcmpy(config_file='../../cfg/fxcm.cfg')
instruments = api.get_instruments()
print(instruments)
# ## Retrieving Historical Data
candles = api.get_candles('USD/JPY', period='D1', number=10)
candles[candles.columns[:4]]
candles[candles.columns[4:]]
start = dt.datetime(2017, 1, 1)
end = dt.datetime(2018, 1, 1)
candles = api.get_candles('EUR/GBP', period='D1',
start=start, stop=end)
candles.info()
# The parameter `period` must be one of `m1, m5, m15, m30, H1, H2, H3, H4, H6, H8, D1, W1` or `M1`.
candles = api.get_candles('EUR/USD', period='m1', number=250)
candles['askclose'].plot(figsize=(10, 6))
# plt.savefig('../../images/ch14/fxcm_plot_03.png');
# ## Streaming Data
def output(data, dataframe):
print('%3d | %s | %s | %6.5f, %6.5f'
% (len(dataframe), data['Symbol'],
pd.to_datetime(int(data['Updated']), unit='ms'),
data['Rates'][0], data['Rates'][1]))
api.subscribe_market_data('EUR/USD', (output,))
api.get_last_price('EUR/USD')
api.unsubscribe_market_data('EUR/USD')
# ## Placing Orders
api.get_open_positions()
order = api.create_market_buy_order('EUR/USD', 100)
sel = ['tradeId', 'amountK', 'currency',
'grossPL', 'isBuy']
api.get_open_positions()[sel]
order = api.create_market_buy_order('EUR/GBP', 50)
api.get_open_positions()[sel]
order = api.create_market_sell_order('EUR/USD', 25)
order = api.create_market_buy_order('EUR/GBP', 50)
api.get_open_positions()[sel]
api.close_all_for_symbol('EUR/GBP')
api.get_open_positions()[sel]
api.close_all()
api.get_open_positions()
# ## Account Information
api.get_default_account()
api.get_accounts().T
# <img src="http://hilpisch.com/tpq_logo.png" alt="The Python Quants" width="35%" align="right" border="0"><br>
#
# <a href="http://tpq.io" target="_blank">http://tpq.io</a> | <a href="http://twitter.com/dyjh" target="_blank">@dyjh</a> | <a href="mailto:<EMAIL>"><EMAIL></a>
| code/ch14/14_trading_platform.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 範例
# ***
# # [教學目標]
#
# * 知道如何從 NumPy 陣列存取元素
# * 了解一維與多維陣列的迴圈用法
# * 初步理解陣列與其迭代物件
#
#
# +
# 載入 NumPy 套件
import numpy as np
# 檢查正確載入與版本
print(np)
print(np.__version__)
# +
# 從陣列中存取元素
L = [0, 1, 8]
print(L)
print(L[-1]) # 8
print(L[0]) # 0
print(L[1:3]) # [1, 8]
# +
import numpy as np
a = np.arange(3) ** 3
print(a)
print(a[-1]) # 8
print(a[0]) # 0
print(a[1:3]) # [1 8]
# +
# 一維陣列的切片與索引
import numpy as np
data = np.array([1, 2, 3])
print(data[0]) # 取出第 0 個
print(data[1]) # 取出第 1 個
print(data[0:2]) # 第 0 - 1 個
print(data[1:]) # 第 1 到最後一個
print(data[-2:]) # 倒數第二到最後一個
# +
# 一維陣列的的迭代
import numpy as np
a = np.arange(3) ** 3
for i in a:
print(i)
# 0
# 1
# 8
# +
# 多維陣列的多層迴圈
import numpy as np
a = np.arange(6).reshape(3, 2)
for row in a:
print(row)
# [0 1]
# [2 3]
# [4 5]
# -
for row in a:
for d in row:
print(d)
# 0
# 1
# 2
# 3
# 4
# 5
# +
# 攤平後再迭代多維陣列
import numpy as np
a = np.arange(6).reshape(3, 2)
for d in a.flat:
print(d)
# +
# np.nditer 迭代物件
import numpy as np
a = np.arange(6).reshape(3, 2)
for d in np.nditer(a):
print(d)
# +
# 迭代物件的儲存方向
a = np.arange(6).reshape(3, 2)
for d in np.nditer(a, order='C'):
print(d)
# +
import numpy as np
a = np.arange(6).reshape(3, 2)
for d in np.nditer(a, order='F'):
print(d)
# -
| Sample Code/Day_07_SampleCode_v2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] cell_style="split"
# # Finchat: Statistics
#
# See detailed information and the labels for the data and columns from the readme file.
# -
import pandas as pd
import numpy as np
import matplotlib.pylab as plt
import seaborn as sns
from collections import Counter
from wordcloud import WordCloud, STOPWORDS, ImageColorGenerator
# Loading the corpus
chat_data = pd.read_csv('../finchat-corpus/finchat_chat_conversations.csv')
meta_data = pd.read_csv('../finchat-corpus/finchat_meta_data.csv')
chat_data.dtypes
meta_data.dtypes
# The number of conversation and participants.
print('The number of the conversations:')
print(chat_data['CHAT_ID'].unique().size)
print(meta_data['CHAT_ID'].unique().size)
print('The number of the users:')
print(chat_data['SPEAKER_ID'].unique().size)
# ## Topic and group statistics
# Changing types
meta_data["GROUP"] = meta_data["GROUP"].astype('category')
meta_data["TOPIC"] = meta_data["TOPIC"].astype('category')
meta_data["OFFTOPIC"] = meta_data["OFFTOPIC"].astype('category')
meta_data[['Q1', 'Q2', 'Q3', 'Q4', 'Q5']] = meta_data[['Q1', 'Q2', 'Q3', 'Q4', 'Q5']].astype('category')
# +
print('groups')
print(meta_data["GROUP"].value_counts())
print()
print('topics')
print(meta_data["TOPIC"].value_counts())
print()
print('offtopics')
print(meta_data["OFFTOPIC"].value_counts())
print()
# -
# ## Filter into smaller sets
# Choose only specific user group or topic to analyze.
# +
# Filter by topic
#meta_subset = meta_data.loc[meta_data['TOPIC'] == 'tv']
#chat_id_list = meta_subset['CHAT_ID'].tolist()
#chat_subset = chat_data.loc[chat_data['CHAT_ID'].isin(chat_id_list)]
# Filter by group
# University staff = 1, university student = 2, high schooler = 3
#meta_subset = meta_data.loc[meta_data['GROUP'] == 3]
#chat_id_list = meta_subset['CHAT_ID'].tolist()
#chat_subset = chat_data.loc[chat_data['CHAT_ID'].isin(chat_id_list)]
# Everything: No subset
chat_id_list = meta_data['CHAT_ID'].tolist()
meta_subset = meta_data
chat_subset = chat_data
# -
# ## Clean text
# +
# Clean text from commas etc to compute word statistics
sentences = chat_subset['TEXT'].tolist()
sentences_clean = []
word_list_text = ''
for sentence in sentences:
for ch in ['.','!','?',')','(',':',',']:
if ch in sentence:
sentence = sentence.replace(ch,'')
sentences_clean.append(sentence.lower())
word_list_text = word_list_text+" "+sentence.lower()
word_list = word_list_text.split()
# -
# ## Words
# ### Common words
word_count = Counter(word_list).most_common()
#print(word_count)
# +
# Generate a word cloud image
stopwords=[]
wordcloud = WordCloud(background_color="white", width=1600, height=1000).generate(word_list_text)
plt.figure( figsize=(20,10) )
plt.imshow(wordcloud, interpolation='bilinear')
plt.axis("off")
plt.figure(figsize=[10,10])
plt.show()
# -
# ### Word length
# +
# Word lengths
word_lengths = [] #np.zeros(30)
for word in word_list:
#ord_lengths[len(word)] += 1
word_lengths.append(len(word))
if len(word) > 25: print(word)
#plt.hist(word_lengths, bins=range(20))
#plt.xticks(range(20))
#plt.xlabel('word length')
#plt.ylabel('word count')
#plt.title('word lengths in corpus')
print('')
print('average word length:', np.mean(word_lengths))
# -
# Word length distribution
sns.distplot(word_lengths, kde=False, norm_hist=True);
plt.xlim(0, 20)
plt.tight_layout()
# ## Statistics
# ### Turns per conversation
# +
# for each conversation
# see how many times the speaker id changes
turn_counter = np.zeros(len(chat_id_list))
for i in range(len(chat_id_list)):
chat = chat_data.loc[chat_data['CHAT_ID'] == chat_id_list[i]]
speaker_id_list = chat['SPEAKER_ID'].tolist()
prev_id = 0
for j in speaker_id_list:
if prev_id != j:
turn_counter[i] += 1
prev_id = j
turn_counter[i] = np.floor(turn_counter[i]/2)
print(np.mean(turn_counter))
# -
# ### Words in corpus
# Words in corpus
len(word_list)
# ### Characters in corpus
total_char = 0
for word in word_list:
total_char += len(word)
print(total_char)
# ### Messages in corpus
print(len(sentences))
# ### Conversations
#
print('The number of the conversations:')
print(chat_subset['CHAT_ID'].unique().size)
# ## Questionaire scores
# Analyze questionaire scores.
#
# See readme documentation of the corpus for actual questions.
# Answers for each question
print(meta_subset["Q1"].value_counts())
print(meta_subset["Q2"].value_counts())
print(meta_subset["Q3"].value_counts())
print(meta_subset["Q4"].value_counts())
print(meta_subset["Q5"].value_counts())
# ### The rate of interesting conversations
print(meta_subset["Q1"].value_counts()/sum(meta_subset["Q1"].value_counts()))
# All Questions
print(meta_subset["Q1"].value_counts()/sum(meta_subset["Q1"].value_counts()))
print(meta_subset["Q2"].value_counts()/sum(meta_subset["Q2"].value_counts()))
print(meta_subset["Q3"].value_counts()/sum(meta_data["Q3"].value_counts()))
print(meta_subset["Q4"].value_counts(sort=False)/sum(meta_subset["Q4"].value_counts()))
print(meta_subset["Q5"].value_counts(sort=False)/sum(meta_subset["Q5"].value_counts()))
# ### Pie plots
# +
# Pie charts for question Q1-Q3.
# For whole data set. Change meta_data -> meta_subset if want to plot for filtered subset.
######## Q1
# Pie chart
labels = ['Yes', 'No']
sizes = np.array(meta_data["Q1"].value_counts())
#colors
colors = ['#ff9999','#66b3ff']
colors = ['tomato','lightskyblue']
#explsion
explode = (0.05,0.05)
fig1, (ax1, ax2, ax3) = plt.subplots(1,3, figsize=(9,3.5))
ax1.pie(sizes, colors = colors, labels=labels, autopct='%1.1f%%', startangle=90)
ax1.set_title('Conversation was interesting.')
# Equal aspect ratio ensures that pie is drawn as a circle
ax1.axis('equal')
plt.tight_layout()
#plt.show()
######## Q2
sizes2 = np.array(meta_data["Q2"].value_counts())
ax2.pie(sizes2, colors = colors, labels=labels, autopct='%1.1f%%', startangle=90)
ax2.set_title('I was listened to.')
# Equal aspect ratio ensures that pie is drawn as a circle
ax2.axis('equal')
plt.tight_layout()
#plt.show()
######## Q3
sizes3 = np.array(meta_data["Q3"].value_counts())
ax3.pie(sizes3, colors = colors, labels=labels, autopct='%1.1f%%', startangle=90)
ax3.set_title('Stayed on topic.')
# Equal aspect ratio ensures that pie is drawn as a circle
ax3.axis('equal')
plt.tight_layout()
# Save plot
#plt.savefig('figs/q1_q2_q3_pie.png', bbox_inches='tight', dpi=300)
# -
# ### More details
# +
# Adding disagreement
# Q1
diff_list = []
pos_sum = 0 # both thought interesting
neg_sum = 0 # both thought not interesting
diff_sum = 0 # # disagreement
no_fb = 0
for chatid in chat_id_list:
meta_conv = meta_data.loc[meta_data['CHAT_ID'] == chatid]
#meta_conv = meta_data.loc[meta_data['CHAT_ID'].isin(chatid)]
# Q1
scores = meta_conv['Q1'].tolist()
if len(scores) == 2:
diff_list.append(abs(scores[0]-scores[1]))
if scores[0] + scores[1] == 2:
pos_sum += 1
elif scores[0] + scores[1] == 4:
neg_sum += 1
else:
diff_sum += 1
else:
#print(scores)
no_fb += 1
# +
# Participants who both agreed that conversation vas intresting or not interesting.
#print(1-sum(diff_list)/len(diff_list))
#print('interesting')
#print(pos_sum/(pos_sum+neg_sum+diff_sum))
#print('not interesting')
#print(neg_sum/(pos_sum+neg_sum+diff_sum))
#print('disagreement')
#print(diff_sum/(pos_sum+neg_sum+diff_sum))
labels = ['Interesting', 'Not interesting', 'Disagreed']
sizes = np.array([pos_sum, neg_sum, diff_sum])
colors = ['tomato', 'lightskyblue','lightgray']
fig1, ax1 = plt.subplots(1,1, figsize=(7,3))
ax1.pie(sizes, colors = colors, labels=labels, autopct='%1.1f%%', startangle=90)
ax1.set_title('Was conversation interesting?')
# Equal aspect ratio ensures that pie is drawn as a circle
ax1.axis('equal')
plt.tight_layout()
#plt.savefig('figs/q1_pie_wdis_staff.png', bbox_inches='tight', dpi=300)
# +
# Similarties between answers for Q4 and Q5
q4_q5_same_sum = 0
q4_q5_notsame_sum = 0
q4_agreed_i = 0 # individual
q4_agreed_b = 0 # both
q4_disa_i = 0
q4_disa_b = 0
q4_disa = 0
q5_agreed_i = 0 # individual
q5_agreed_b = 0 # both
q5_disa = 0
pos_sun = 0
diff_sum = 0
count=0
for chatid in chat_id_list:
# Q4 and Q5 have same answer for same person
meta_conv = meta_data.loc[meta_data['CHAT_ID'] == chatid]
scores_q4 = meta_conv['Q4'].tolist()
scores_q5 = meta_conv['Q5'].tolist()
if len(scores_q4) == 2:
if scores_q4[0] == scores_q5[0]:
q4_q5_same_sum += 1
else: q4_q5_notsame_sum += 1
if scores_q4[1] == scores_q5[1]:
q4_q5_same_sum += 1
else: q4_q5_notsame_sum += 1
# For same conversation both agreed with the leader:
# answers 1, 2 OR 3, 3. Q4
if scores_q4[0] != scores_q4[1] and scores_q4[0] != 3 and scores_q4[1] != 3:
q4_agreed_i += 1
elif scores_q4[0] == scores_q4[1] and scores_q4[0] == 3:
q4_agreed_b += 1
elif scores_q4[0] != scores_q4[1] and (scores_q4[0] == 3 or scores_q4[1] == 3):
q4_disa_b += 1
else: q4_disa += 1
# For same conversation both agreed with the leader:
# answers 1, 2 OR 3, 3. Q5
if scores_q5[0] != scores_q5[1] and scores_q5[0] != 3 and scores_q5[1] != 3:
q5_agreed_i += 1
elif scores_q5[0] == scores_q5[1] and scores_q5[0] == 3:
q5_agreed_b += 1
else: q5_disa += 1
else: count+=1
print('Asking more question and leading conversation (own opinion)')
print(q4_q5_same_sum/(q4_q5_same_sum+q4_q5_notsame_sum))
print('Agreeing who is asking the questions (Q4)')
print((q4_agreed_b+q4_agreed_i)/(q4_agreed_i+q4_agreed_b+q4_disa))
print('Agreeing who is the leader (5)')
print((q5_agreed_i+q5_agreed_b)/(q5_agreed_i+q5_agreed_b+q5_disa))
# +
# Plots with disagreement and no answers.
# Combine
font = {'family' : 'sans-serif',
'weight' : 'normal',
'size' : 11.5}
plt.rc('font', **font)
######## Q1
#explsion
explode = (0.05,0.05)
fig1, [ax1, ax2, ax3] = plt.subplots(1,3, figsize=(10,3.5))
labels = ['Yes', 'No', 'Disagreed']
sizes = np.array([pos_sum, neg_sum, diff_sum])
colors = ['tomato', 'lightskyblue','lightgray']
ax1.pie(sizes, colors = colors, labels=labels, autopct='%1.1f%%', startangle=90, textprops={'fontsize': 13})
ax1.set_title('1. Conversation was interesting.')
# Equal aspect ratio ensures that pie is drawn as a circle
ax1.axis('equal')
plt.tight_layout()
# Q4 & Q5
# Pie chart
labels = ['No answer', 'Me', 'Partner', 'Both']
sizes = np.array(meta_data["Q4"].value_counts(sort=False))
#colors
#colors = ['#ff9999','#66b3ff']
colors = ['lightgray', 'tomato','lightskyblue','lightgreen']
ax2.set_title('2. Asked more questions.')
ax2.pie(sizes, colors = colors, labels=labels, autopct='%1.1f%%', startangle=90, textprops={'fontsize': 13})
# Equal aspect ratio ensures that pie is drawn as a circle
ax2.axis('equal')
plt.tight_layout()
#plt.show()
##### Q5
sizes2 = np.array(meta_data["Q5"].value_counts(sort=False))
ax3.pie(sizes2, colors = colors, labels=labels, autopct='%1.1f%%', startangle=90, textprops={'fontsize': 13})
ax3.set_title('3. Leader of the conversation.')
# Equal aspect ratio ensures that pie is drawn as a circle
ax3.axis('equal')
plt.tight_layout()
#axs[1, 2].axis('off')
plt.savefig('questionnairre_pies_smaller_fixed.png', bbox_inches='tight', dpi=300)
# +
# Pie plots for Q4 & Q5
# For whole set. Change meta_data -> meta_subset if want to plot for subset.
# Pie chart
labels = ['No answer', 'Me', 'Partner', 'Both']
sizes = np.array(meta_data["Q4"].value_counts(sort=False))
#colors
#colors = ['#ff9999','#66b3ff']
colors = ['lightgray', 'tomato','lightskyblue','lightgreen']
fig1, (ax1, ax2) = plt.subplots(1,2, figsize=(7,3))
ax1.pie(sizes, colors = colors, labels=labels, autopct='%1.1f%%', startangle=90)
ax1.set_title('Who asked more question?')
# Equal aspect ratio ensures that pie is drawn as a circle
ax1.axis('equal')
plt.tight_layout()
#plt.show()
##### Q5
sizes2 = np.array(meta_data["Q5"].value_counts(sort=False))
ax2.pie(sizes2, colors = colors, labels=labels, autopct='%1.1f%%', startangle=90)
ax2.set_title('Who was leading the conversation?')
# Equal aspect ratio ensures that pie is drawn as a circle
ax2.axis('equal')
plt.tight_layout()
# Save plot
#plt.savefig('figs/q4_q5_pie.png', bbox_inches='tight', dpi=300)
# +
# Combined plots.
font = {'family' : 'sans-serif',
'weight' : 'normal',
'size' : 11.5}
plt.rc('font', **font)
######## Q1
#explsion
explode = (0.05,0.05)
fig1, axs = plt.subplots(2,3, figsize=(9,7))
labels = ['Yes', 'No', 'Disagreed']
sizes = np.array([pos_sum, neg_sum, diff_sum])
colors = ['tomato', 'lightskyblue','lightgray']
axs[0, 0].pie(sizes, colors = colors, labels=labels, autopct='%1.1f%%', startangle=90, textprops={'fontsize': 14})
axs[0, 0].set_title('1. Was conversation interesting?')
# Equal aspect ratio ensures that pie is drawn as a circle
axs[0, 0].axis('equal')
plt.tight_layout()
######## Q2
# Pie chart
labels = ['Yes', 'No']
#colors
colors = ['#ff9999','#66b3ff']
colors = ['tomato','lightskyblue']
sizes2 = np.array(meta_data["Q2"].value_counts())
axs[0, 1].pie(sizes2, colors = colors, labels=labels, autopct='%1.1f%%', startangle=90)
axs[0, 1].set_title('2. I was listened to.')
# Equal aspect ratio ensures that pie is drawn as a circle
axs[0, 1].axis('equal')
plt.tight_layout()
#plt.show()
######## Q3
sizes3 = np.array(meta_data["Q3"].value_counts())
axs[0, 2].pie(sizes3, colors = colors, labels=labels, autopct='%1.1f%%', startangle=90)
axs[0, 2].set_title('3. Stayed on topic.')
# Equal aspect ratio ensures that pie is drawn as a circle
axs[0, 2].axis('equal')
plt.tight_layout()
# Q4 & Q5
# Pie chart
labels = ['No answer', 'Me', 'Partner', 'Both']
sizes = np.array(meta_data["Q4"].value_counts(sort=False))
#colors
#colors = ['#ff9999','#66b3ff']
colors = ['lightgray', 'tomato','lightskyblue','lightgreen']
axs[1, 0].set_title('4. Asked more questions.')
axs[1, 0].pie(sizes, colors = colors, labels=labels, autopct='%1.1f%%', startangle=90)
# Equal aspect ratio ensures that pie is drawn as a circle
axs[1, 0].axis('equal')
plt.tight_layout()
#plt.show()
##### Q5
sizes2 = np.array(meta_data["Q5"].value_counts(sort=False))
axs[1, 1].pie(sizes2, colors = colors, labels=labels, autopct='%1.1f%%', startangle=90)
axs[1, 1].set_title('5. Learder of the conversation.')
# Equal aspect ratio ensures that pie is drawn as a circle
axs[1, 1].axis('equal')
plt.tight_layout()
axs[1, 2].axis('off')
#plt.savefig('figs/questionnairre_pies.png', bbox_inches='tight', dpi=300)
# +
# Combine
font = {'family' : 'sans-serif',
'weight' : 'normal',
'size' : 11.5}
plt.rc('font', **font)
######## Q1
#explsion
explode = (0.05,0.05)
fig1, axs = plt.subplots(2,3, figsize=(9,7))
labels = ['Yes', 'No', 'Disagreed']
sizes = np.array([pos_sum, neg_sum, diff_sum])
colors = ['tomato', 'lightskyblue','lightgray']
axs[0, 0].pie(sizes, colors = colors, labels=labels, autopct='%1.1f%%', startangle=90, textprops={'fontsize': 14})
axs[0, 0].set_title('1. Was conversation interesting?')
# Equal aspect ratio ensures that pie is drawn as a circle
axs[0, 0].axis('equal')
plt.tight_layout()
######## Q2
# Pie chart
labels = ['Yes', 'No']
#colors
colors = ['#ff9999','#66b3ff']
colors = ['tomato','lightskyblue']
sizes2 = np.array(meta_data["Q2"].value_counts())
axs[0, 1].pie(sizes2, colors = colors, labels=labels, autopct='%1.1f%%', startangle=90)
axs[0, 1].set_title('2. I was listened to.')
# Equal aspect ratio ensures that pie is drawn as a circle
axs[0, 1].axis('equal')
plt.tight_layout()
#plt.show()
######## Q3
sizes3 = np.array(meta_data["Q3"].value_counts())
axs[0, 2].pie(sizes3, colors = colors, labels=labels, autopct='%1.1f%%', startangle=90)
axs[0, 2].set_title('3. Stayed on topic.')
# Equal aspect ratio ensures that pie is drawn as a circle
axs[0, 2].axis('equal')
plt.tight_layout()
# Q4 & Q5
# Pie chart
labels = ['No answer', 'Me', 'Partner', 'Both']
sizes = np.array(meta_data["Q4"].value_counts(sort=False))
#colors
#colors = ['#ff9999','#66b3ff']
colors = ['lightgray', 'tomato','lightskyblue','lightgreen']
axs[1, 0].set_title('4. Asked more questions.')
axs[1, 0].pie(sizes, colors = colors, labels=labels, autopct='%1.1f%%', startangle=90)
# Equal aspect ratio ensures that pie is drawn as a circle
axs[1, 0].axis('equal')
plt.tight_layout()
#plt.show()
##### Q5
sizes2 = np.array(meta_data["Q5"].value_counts(sort=False))
axs[1, 1].pie(sizes2, colors = colors, labels=labels, autopct='%1.1f%%', startangle=90)
axs[1, 1].set_title('5. Learder of the conversation.')
# Equal aspect ratio ensures that pie is drawn as a circle
axs[1, 1].axis('equal')
plt.tight_layout()
axs[1, 2].axis('off')
#plt.savefig('figs/questionnairre_pies_.png', bbox_inches='tight', dpi=300)
# +
# Select group
#
#meta_subset = meta_data.loc[meta_data['GROUP'] == 1]
#chat_id_list = meta_subset['CHAT_ID'].tolist()
#chat_subset = chat_data.loc[chat_data['CHAT_ID'].isin(chat_id_list)]
#chat_id_list = chat_subset['CHAT_ID'].unique().tolist()
# Use all
#chat_id_list = chat_data['CHAT_ID'].unique().tolist()
# -
#
# +
# Commoness between question answers
scores_q4 = np.array(meta_data['Q4'].tolist()) # 3 = both
scores_q1 = np.array(meta_data['Q1'].tolist()) # 1 = yes
print(np.unique(scores_q1,return_counts=True))
count = 0
for i in range(len(scores_q1)):
if scores_q1[i] == 1 and scores_q4[i] == 2:
count += 1
print(count/len(scores_q1))
print(count/120)
# both ask questions and is interesting = 44% / 60%
# i asked more and is interesting = 14% / 19 %
# partner asked more and is interestin = 10 / 14%
# -
# ### Alternative to pies
# +
category_names = ['Yes', 'Disagree', 'No']
sizes = [26,4,11]
sizes2 = [157,0,6]
sizes3 = [128,0,35]
results = {
'Question 1': sizes,
'Question 2': sizes2,
'Question 3': sizes3
}
def survey(results, category_names):
"""
Parameters
----------
results : dict
A mapping from question labels to a list of answers per category.
It is assumed all lists contain the same number of entries and that
it matches the length of *category_names*.
category_names : list of str
The category labels.
"""
labels = list(results.keys())
data = np.array(list(results.values()))
data_cum = data.cumsum(axis=1)
category_colors = plt.get_cmap('RdYlGn')(
np.linspace(0.15, 0.85, data.shape[1]))
fig, ax = plt.subplots(figsize=(9.2, 5))
ax.invert_yaxis()
ax.xaxis.set_visible(False)
ax.set_xlim(0, np.sum(data, axis=1).max())
for i, (colname, color) in enumerate(zip(category_names, category_colors)):
widths = data[:, i]
starts = data_cum[:, i] - widths
ax.barh(labels, widths, left=starts, height=0.5,
label=colname, color=color)
xcenters = starts + widths / 2
r, g, b, _ = color
text_color = 'white' if r * g * b < 0.5 else 'darkgrey'
for y, (x, c) in enumerate(zip(xcenters, widths)):
ax.text(x, y, str(int(c)), ha='center', va='center',
color=text_color)
ax.legend(ncol=len(category_names), bbox_to_anchor=(0, 1),
loc='lower left', fontsize='small')
return fig, ax
survey(results, category_names)
plt.show()
# -
print(sizes3)
| notebooks/statistics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
from pyvista import set_plot_theme
set_plot_theme('document')
# Slider Bar Widget {#slider_bar_widget_example}
# =================
#
# The slider widget can be enabled and disabled by the
# `pyvista.WidgetHelper.add_slider_widget`{.interpreted-text role="func"}
# and `pyvista.WidgetHelper.clear_slider_widgets`{.interpreted-text
# role="func"} methods respectively. This is one of the most versatile
# widgets as it can control a value that can be used for just about
# anything.
#
# One helper method we\'ve added is the
# `pyvista.WidgetHelper.add_mesh_threshold`{.interpreted-text role="func"}
# method which leverages the slider widget to control a thresholding
# value.
#
# +
import pyvista as pv
from pyvista import examples
mesh = examples.download_knee_full()
p = pv.Plotter()
p.add_mesh_threshold(mesh)
p.show()
# -
# After interacting with the scene, the threshold mesh is available as:
#
p.threshold_meshes
# And here is a screen capture of a user interacting with this
#
# 
#
# Custom Callback
# ===============
#
# Or you could leverage a custom callback function that takes a single
# value from the slider as its argument to do something like control the
# resolution of a mesh. Again note the use of the `name` argument in
# `add_mesh`:
#
# +
p = pv.Plotter()
def create_mesh(value):
res = int(value)
sphere = pv.Sphere(phi_resolution=res, theta_resolution=res)
p.add_mesh(sphere, name='sphere', show_edges=True)
return
p.add_slider_widget(create_mesh, [5, 100], title='Resolution')
p.show()
# -
# And here is a screen capture of a user interacting with this
#
# 
#
| examples/03-widgets/slider-bar-widget.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Introduction & Background
#
# Concepts arising in nonlinear dynamical systems theory, such as periodic orbits, normally hyperbolic invariant manifolds (NHIMs), and stable and unstable manifolds have been introduced into the study of chemical reaction dynamics from the phase space perspective. A fairly substantial literature has been developed on this topic in recent years (see, for example {% cite wiggins2016role waalkens2010geometrical waalkens2007wigner wiggins2013normally --file book_sprint_bib1 %}, and references therein), but it is fair to say that it has a more mathematical flavour, which is not surprising since these concepts originated in the dynamical systems literature. In this book we will describe how these dynamical notions arise in a variety of physically motivated settings with the hope of providing a more gentle entry into the field for both applied mathematicians and chemists.
#
# An obstacle in interdiscipliary work is the lack of a common language for describing concepts that are common to different fields. We begin with a list of commonly used terms that will arise repeatedly throughout this book and provide a working definition.
#
# Molecules are made up of a collection of atoms that are connected by chemical bonds and a reaction is concerned with the breaking, and creation, of these bonds. Hence, the following concepts are fundamental to the description of this phenomena.
#
# + **Coordinates.** The locations of the atoms in a molecule are described by a set of coordinates. The space (that is, all possible values) described by these coordinates is referred to as *configuration space*.
#
#
# + **Degrees-of-Freedom (DoF).** The number of DoF is the number of independent coordinates required to describe the configuration of the molecule, that is the dimension of the configuration space.
#
#
# + **Reaction.** The breaking of a bond can be described by one or more coordinates characterizing the bond becoming unbounded as the it evolves in time.
#
#
# + **Reaction coordinate(s).** The particular coordinate(s) that describe the breaking of the bond are referred to as the *reaction coordinate(s)*.
#
#
# + **Energy.** The ability of a bond to break can be characterised by its energy. A bond can be ''energized'' by transferring energy from other bonds in the molecule to a particular bond of interest, or from some external energy source, such as electromagnetic radiation, collision with other molecules, for example.
#
#
# + **Total Energy, Hamiltonian, momenta.** The total energy (that is, the sum of kinetic energy and potential energy) can be described by a scalar valued function called the *Hamiltonian*. The Hamiltonian is a function of the configuration space coordinates *and* their corresponding canonically conjugate coordinates, which are referred to as *momentum coordinates*.
#
#
# + **Phase Space.** The collection of all the configuration and momentum coordinates is referred to as the phase space of the system. The dynamics of the system (that is, how it changes in time) is described by Hamilton's (differential) equations of motion defined on phase space.
#
#
# + **Dimension count.** If the system has *n* configuration space coordinates, it has *n* momentum coordinates and then the phase space dimension is *2n*. The Hamiltonian is a scalar valued function of these *2n* coordinates. The level set of the Hamiltonian, that is the energy surface, is *2n-1* dimensional. For a time-independent (autonomous) Hamiltonian, the system *conserves energy* and the energy surface is invariant.
#
# + **Transition State Theory (TST).** An approach in reaction rate calculations that is based on the flux across a dividing surface. We give a brief description of the theory in the next section.
#
# + **Dividing Surface (DS).** A DS on the energy surface is of dimension *2n-2*, that is, 1 dimension less (codimension 1) than the *2n-1* dimensional energy surface. While a codimension one surface has the dimensionality necessary to divide the energy surface into two distinct regions and forms the boundary between them. If properly chosen, the two regions are referred to as *reactants* and *products*, and reaction occurs when trajectories evolve from reactants to products through the DS. A DS has the ''locally no-recrossing'' property, which is equivalent to the Hamiltonian vector field being everywhere transverse to a DS, that is at no point is it tangent to the DS.
#
# + **Locally no-recrossing.** Computation of the flux is accurate only if the DS has the ''locally no-recrossing'' property. A surface has the ''locally no-recrossing'' property if any trajectory that crosses the surface leaves a neighbourhood of the surface before it can return.
#
# + **Globally no-recrossing.** A surface has the ''globally no-recrossing'' property if any trajectory that crosses the surface does so only once.
#
# + **The DS and the Reaction Coordinate.** Following our definitions of reaction, reaction coordinate, and DS it follows that the reaction coordinate should play a role in the definition of the DS. This will be an important point in our discussions that follow.
#
#
# # References
# {% bibliography --file book_sprint_bib1 --cited %}
| content/prologue/introduction_and_background-jekyll.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Artificial Neural Network
# Installing Theano
# pip install --upgrade --no-deps git+git://github.com/Theano/Theano.git
# Installing Tensorflow
# Install Tensorflow from the website: https://www.tensorflow.org/versions/r0.12/get_started/os_setup.html
# Installing Keras
# pip install --upgrade keras
# Part 1 - Data Preprocessing
# Importing the libraries
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# Importing the dataset
dataset = pd.read_csv('Churn_Modelling.csv')
X = dataset.iloc[:, 3:13].values
y = dataset.iloc[:, 13].values
# Encoding categorical data
from sklearn.preprocessing import LabelEncoder, OneHotEncoder
labelencoder_X_1 = LabelEncoder()
X[:, 1] = labelencoder_X_1.fit_transform(X[:, 1])
labelencoder_X_2 = LabelEncoder()
X[:, 2] = labelencoder_X_2.fit_transform(X[:, 2])
onehotencoder = OneHotEncoder(categorical_features = [1])
X = onehotencoder.fit_transform(X).toarray()
X = X[:, 1:]
# Splitting the dataset into the Training set and Test set
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.2, random_state = 0)
# Feature Scaling
from sklearn.preprocessing import StandardScaler
sc = StandardScaler()
X_train = sc.fit_transform(X_train)
X_test = sc.transform(X_test)
# Part 2 - Now let's make the ANN!
# Importing the Keras libraries and packages
import keras
from keras.models import Sequential
from keras.layers import Dense
# Initialising the ANN
classifier = Sequential()
# Adding the input layer and the first hidden layer
"""Glorot_uniform is a much better initializer and it has been carefully designed to give great
initializations to neurons for the best possible training."""
classifier.add(Dense(output_dim = 6, init = 'glorot_uniform', activation = 'relu', input_dim = 11))
# Adding the second hidden layer
classifier.add(Dense(output_dim = 6, init = 'glorot_uniform', activation = 'relu'))
# Adding the output layer
classifier.add(Dense(output_dim = 1, init = 'glorot_uniform', activation = 'sigmoid'))
# Compiling the ANN
classifier.compile(optimizer = 'rmsprop', loss = 'binary_crossentropy', metrics = ['accuracy'])
# Fitting the ANN to the Training set
classifier.fit(X_train, y_train, batch_size = 25, nb_epoch = 100)
# Part 3 - Making the predictions and evaluating the model
# Predicting the Test set results
y_pred = classifier.predict(X_test)
y_pred = (y_pred > 0.5)
# Making the Confusion Matrix
from sklearn.metrics import confusion_matrix
cm = confusion_matrix(y_test, y_pred)
# -
| supervised_learning/ANN/basic_ANN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="ur8xi4C7S06n"
# Copyright 2022 Google LLC
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="JAPoU8Sm5E6e"
# <table align="left">
#
# <td>
# <a href="https://colab.research.google.com/github/GoogleCloudPlatform/vertex-ai-samples/blob/master/notebooks/community/ml_metadata/sdk-metric-parameter-tracking-for-locally-trained-models.ipynb">
# <img src="https://cloud.google.com/ml-engine/images/colab-logo-32px.png" alt="Colab logo"> Run in Colab
# </a>
# </td>
# <td>
# <a href="https://github.com/GoogleCloudPlatform/vertex-ai-samples/blob/master/notebooks/community/ml_metadata/sdk-metric-parameter-tracking-for-locally-trained-models.ipynb">
# <img src="https://cloud.google.com/ml-engine/images/github-logo-32px.png" alt="GitHub logo">
# View on GitHub
# </a>
# </td>
# <td>
# <a href="https://console.cloud.google.com/vertex-ai/workbench/deploy-notebook?download_url=https://github.com/GoogleCloudPlatform/vertex-ai-samples/blob/main/notebooks/notebook_template.ipynb">
# <img src="https://lh3.googleusercontent.com/UiNooY4LUgW_oTvpsNhPpQzsstV5W8F7rYgxgGBD85cWJoLmrOzhVs_ksK_vgx40SHs7jCqkTkCk=e14-rj-sc0xffffff-h130-w32" alt="Vertex AI logo">
# Open in Vertex AI Workbench
# </a>
# </td>
# </table>
# + [markdown] id="WBFL9LagqmwT"
# ## Vertex AI: Track parameters and metrics for locally trained models
# + [markdown] id="tvgnzT1CKxrO"
# ## Overview
#
# This notebook demonstrates how to track metrics and parameters for ML training jobs and analyze this metadata using Vertex SDK for Python.
#
# ### Dataset
#
# In this notebook, we will train a simple distributed neural network (DNN) model to predict automobile's miles per gallon (MPG) based on automobile information in the [auto-mpg dataset](https://www.kaggle.com/devanshbesain/exploration-and-analysis-auto-mpg).
#
# ### Objective
#
# In this notebook, you will learn how to use Vertex SDK for Python to:
#
# * Track parameters and metrics for a locally trained model.
# * Extract and perform analysis for all parameters and metrics within an Experiment.
#
# ### Costs
#
#
# This tutorial uses billable components of Google Cloud:
#
# * Vertex AI
# * Cloud Storage
#
#
# Learn about [Vertex AI
# pricing](https://cloud.google.com/vertex-ai/pricing) and [Cloud Storage
# pricing](https://cloud.google.com/storage/pricing), and use the [Pricing
# Calculator](https://cloud.google.com/products/calculator/)
# to generate a cost estimate based on your projected usage.
# + [markdown] id="ze4-nDLfK4pw"
# ### Set up your local development environment
#
# **If you are using Colab or Vertex AI Workbench notebooks**, your environment already meets
# all the requirements to run this notebook. You can skip this step.
# + [markdown] id="gCuSR8GkAgzl"
# **Otherwise**, make sure your environment meets this notebook's requirements.
# You need the following:
#
# * The Google Cloud SDK
# * Git
# * Python 3
# * virtualenv
# * Jupyter notebook running in a virtual environment with Python 3
#
# The Google Cloud guide to [Setting up a Python development
# environment](https://cloud.google.com/python/setup) and the [Jupyter
# installation guide](https://jupyter.org/install) provide detailed instructions
# for meeting these requirements. The following steps provide a condensed set of
# instructions:
#
# 1. [Install and initialize the Cloud SDK.](https://cloud.google.com/sdk/docs/)
#
# 1. [Install Python 3.](https://cloud.google.com/python/setup#installing_python)
#
# 1. [Install
# virtualenv](https://cloud.google.com/python/setup#installing_and_using_virtualenv)
# and create a virtual environment that uses Python 3. Activate the virtual environment.
#
# 1. To install Jupyter, run `pip install jupyter` on the
# command-line in a terminal shell.
#
# 1. To launch Jupyter, run `jupyter notebook` on the command-line in a terminal shell.
#
# 1. Open this notebook in the Jupyter Notebook Dashboard.
# + [markdown] id="i7EUnXsZhAGF"
# ### Install additional packages
#
# Run the following commands to install the Vertex SDK for Python.
# + id="IaYsrh0Tc17L"
import sys
if "google.colab" in sys.modules:
USER_FLAG = ""
else:
USER_FLAG = "--user"
# + id="wyy5Lbnzg5fi"
# ! pip3 install -U tensorflow==2.8 $USER_FLAG
# ! pip3 install --upgrade google-cloud-aiplatform $USER_FLAG
# + [markdown] id="hhq5zEbGg0XX"
# ### Restart the kernel
#
# After you install the additional packages, you need to restart the notebook kernel so it can find the packages.
# + id="EzrelQZ22IZj"
# Automatically restart kernel after installs
import os
if not os.getenv("IS_TESTING"):
# Automatically restart kernel after installs
import IPython
app = IPython.Application.instance()
app.kernel.do_shutdown(True)
# + [markdown] id="lWEdiXsJg0XY"
# ## Before you begin
#
# ### Select a GPU runtime
#
# **Make sure you're running this notebook in a GPU runtime if you have that option. In Colab, select "Runtime --> Change runtime type > GPU"**
# + [markdown] id="BF1j6f9HApxa"
# ### Set up your Google Cloud project
#
# **The following steps are required, regardless of your notebook environment.**
#
# 1. [Select or create a Google Cloud project](https://console.cloud.google.com/cloud-resource-manager). When you first create an account, you get a $300 free credit towards your compute/storage costs.
#
# 1. [Make sure that billing is enabled for your project](https://cloud.google.com/billing/docs/how-to/modify-project).
#
# 1. [Enable the Vertex AI API](https://console.cloud.google.com/flows/enableapi?apiid=aiplatform.googleapis.com).
#
# 1. If you are running this notebook locally, you will need to install the [Cloud SDK](https://cloud.google.com/sdk).
#
# 1. Enter your project ID in the cell below. Then run the cell to make sure the
# Cloud SDK uses the right project for all the commands in this notebook.
#
# **Note**: Jupyter runs lines prefixed with `!` as shell commands, and it interpolates Python variables prefixed with `$` into these commands.
# + [markdown] id="WReHDGG5g0XY"
# #### Set your project ID
#
# **If you don't know your project ID**, you may be able to get your project ID using `gcloud`.
# + id="oM1iC_MfAts1"
import os
PROJECT_ID = ""
# Get your Google Cloud project ID from gcloud
if not os.getenv("IS_TESTING"):
# shell_output = !gcloud config list --format 'value(core.project)' 2>/dev/null
PROJECT_ID = shell_output[0]
print("Project ID: ", PROJECT_ID)
# + [markdown] id="qJYoRfYng0XZ"
# Otherwise, set your project ID here.
# + id="riG_qUokg0XZ"
if PROJECT_ID == "" or PROJECT_ID is None:
PROJECT_ID = "[your-project-id]" # @param {type:"string"}
# + [markdown] id="06571eb4063b"
# #### Timestamp
#
# If you are in a live tutorial session, you might be using a shared test account or project. To avoid name collisions between users on resources created, you create a timestamp for each instance session, and append it onto the name of resources you create in this tutorial.
# + id="697568e92bd6"
from datetime import datetime
TIMESTAMP = datetime.now().strftime("%Y%m%d%H%M%S")
# + [markdown] id="dr--iN2kAylZ"
# ### Authenticate your Google Cloud account
#
# **If you are using Vertex AI Workbench notebooks**, your environment is already
# authenticated. Skip this step.
# + [markdown] id="sBCra4QMA2wR"
# **If you are using Colab**, run the cell below and follow the instructions
# when prompted to authenticate your account via oAuth.
#
# **Otherwise**, follow these steps:
#
# 1. In the Cloud Console, go to the [**Create service account key**
# page](https://console.cloud.google.com/apis/credentials/serviceaccountkey).
#
# 2. Click **Create service account**.
#
# 3. In the **Service account name** field, enter a name, and
# click **Create**.
#
# 4. In the **Grant this service account access to project** section, click the **Role** drop-down list. Type "Vertex AI"
# into the filter box, and select
# **Vertex AI Administrator**. Type "Storage Object Admin" into the filter box, and select **Storage Object Admin**.
#
# 5. Click *Create*. A JSON file that contains your key downloads to your
# local environment.
#
# 6. Enter the path to your service account key as the
# `GOOGLE_APPLICATION_CREDENTIALS` variable in the cell below and run the cell.
# + id="PyQmSRbKA8r-"
import os
import sys
# If you are running this notebook in Colab, run this cell and follow the
# instructions to authenticate your GCP account. This provides access to your
# Cloud Storage bucket and lets you submit training jobs and prediction
# requests.
# If on Google Cloud Notebooks, then don't execute this code
if not os.path.exists("/opt/deeplearning/metadata/env_version"):
if "google.colab" in sys.modules:
from google.colab import auth as google_auth
google_auth.authenticate_user()
# If you are running this notebook locally, replace the string below with the
# path to your service account key and run this cell to authenticate your GCP
# account.
elif not os.getenv("IS_TESTING"):
# %env GOOGLE_APPLICATION_CREDENTIALS ''
# + [markdown] id="XoEqT2Y4DJmf"
# ### Import libraries and define constants
# + [markdown] id="Y9Uo3tifg1kx"
# Import required libraries.
# + id="pRUOFELefqf1"
import matplotlib.pyplot as plt
import pandas as pd
from google.cloud import aiplatform
from tensorflow.python.keras import Sequential, layers
from tensorflow.python.keras.utils import data_utils
# + [markdown] id="xtXZWmYqJ1bh"
# Define some constants
# + id="JIOrI-hoJ46P"
EXPERIMENT_NAME = "" # @param {type:"string"}
REGION = "[your-region]" # @param {type:"string"}
if REGION == "[your-region]":
REGION = "us-central1"
# + [markdown] id="jWQLXXNVN4Lv"
# If EXEPERIMENT_NAME is not set, set a default one below:
# + id="Q1QInYWOKsmo"
if EXPERIMENT_NAME == "" or EXPERIMENT_NAME is None:
EXPERIMENT_NAME = "my-experiment-" + TIMESTAMP
# + [markdown] id="Xuny18aMcWDb"
# ## Concepts
#
# To better understanding how parameters and metrics are stored and organized, we'd like to introduce the following concepts:
#
# + [markdown] id="NThDci5bp0Uw"
# ### Experiment
# Experiments describe a context that groups your runs and the artifacts you create into a logical session. For example, in this notebook you create an Experiment and log data to that experiment.
# + [markdown] id="SAyRR3Ydp4X5"
# ### Run
# A run represents a single path/avenue that you executed while performing an experiment. A run includes artifacts that you used as inputs or outputs, and parameters that you used in this execution. An Experiment can contain multiple runs.
# + [markdown] id="l1YW2pgyegFP"
# ## Getting started tracking parameters and metrics
#
# You can use the Vertex SDK for Python to track metrics and parameters for models trained locally.
#
# In the following example, you train a simple distributed neural network (DNN) model to predict automobile's miles per gallon (MPG) based on automobile information in the [auto-mpg dataset](https://www.kaggle.com/devanshbesain/exploration-and-analysis-auto-mpg).
# + [markdown] id="KPY41M9_AhZU"
# ### Load and process the training dataset
# + [markdown] id="bfMQSmRuUuX-"
# Download and process the dataset.
# + id="RiQuMv4bmpuV"
def read_data(uri):
dataset_path = data_utils.get_file("auto-mpg.data", uri)
column_names = [
"MPG",
"Cylinders",
"Displacement",
"Horsepower",
"Weight",
"Acceleration",
"Model Year",
"Origin",
]
raw_dataset = pd.read_csv(
dataset_path,
names=column_names,
na_values="?",
comment="\t",
sep=" ",
skipinitialspace=True,
)
dataset = raw_dataset.dropna()
dataset["Origin"] = dataset["Origin"].map(
lambda x: {1: "USA", 2: "Europe", 3: "Japan"}.get(x)
)
dataset = pd.get_dummies(dataset, prefix="", prefix_sep="")
return dataset
dataset = read_data(
"http://archive.ics.uci.edu/ml/machine-learning-databases/auto-mpg/auto-mpg.data"
)
# + [markdown] id="Y06J7A7yU21t"
# Split dataset for training and testing.
# + id="p5JBCBKyH-NC"
def train_test_split(dataset, split_frac=0.8, random_state=0):
train_dataset = dataset.sample(frac=split_frac, random_state=random_state)
test_dataset = dataset.drop(train_dataset.index)
train_labels = train_dataset.pop("MPG")
test_labels = test_dataset.pop("MPG")
return train_dataset, test_dataset, train_labels, test_labels
train_dataset, test_dataset, train_labels, test_labels = train_test_split(dataset)
# + [markdown] id="gaNNTFPaU7KT"
# Normalize the features in the dataset for better model performance.
# + id="VGq5QCoyIEWJ"
def normalize_dataset(train_dataset, test_dataset):
train_stats = train_dataset.describe()
train_stats = train_stats.transpose()
def norm(x):
return (x - train_stats["mean"]) / train_stats["std"]
normed_train_data = norm(train_dataset)
normed_test_data = norm(test_dataset)
return normed_train_data, normed_test_data
normed_train_data, normed_test_data = normalize_dataset(train_dataset, test_dataset)
# + [markdown] id="UBXUgxgqA_GB"
# ### Define ML model and training function
# + id="66odBYKrIN4q"
def train(
train_data,
train_labels,
num_units=64,
activation="relu",
dropout_rate=0.0,
validation_split=0.2,
epochs=1000,
):
model = Sequential(
[
layers.Dense(
num_units,
activation=activation,
input_shape=[len(train_dataset.keys())],
),
layers.Dropout(rate=dropout_rate),
layers.Dense(num_units, activation=activation),
layers.Dense(1),
]
)
model.compile(loss="mse", optimizer="adam", metrics=["mae", "mse"])
print(model.summary())
history = model.fit(
train_data, train_labels, epochs=epochs, validation_split=validation_split
)
return model, history
# + [markdown] id="O8XJZB3gR8eL"
# ### Initialize the Vertex AI SDK for Python and create an Experiment
#
# Initialize the *client* for Vertex AI and create an experiment.
# + id="o_wnT10RJ7-W"
aiplatform.init(project=PROJECT_ID, location=REGION, experiment=EXPERIMENT_NAME)
# + [markdown] id="u-iTnzt3B6Z_"
# ### Start several model training runs
#
# Training parameters and metrics are logged for each run.
# + id="i2wnpu8_7JfV"
parameters = [
{"num_units": 16, "epochs": 3, "dropout_rate": 0.1},
{"num_units": 16, "epochs": 10, "dropout_rate": 0.1},
{"num_units": 16, "epochs": 10, "dropout_rate": 0.2},
{"num_units": 32, "epochs": 10, "dropout_rate": 0.1},
{"num_units": 32, "epochs": 10, "dropout_rate": 0.2},
]
for i, params in enumerate(parameters):
aiplatform.start_run(run=f"auto-mpg-local-run-{i}")
aiplatform.log_params(params)
model, history = train(
normed_train_data,
train_labels,
num_units=params["num_units"],
activation="relu",
epochs=params["epochs"],
dropout_rate=params["dropout_rate"],
)
aiplatform.log_metrics(
{metric: values[-1] for metric, values in history.history.items()}
)
loss, mae, mse = model.evaluate(normed_test_data, test_labels, verbose=2)
aiplatform.log_metrics({"eval_loss": loss, "eval_mae": mae, "eval_mse": mse})
# + [markdown] id="jZLrJZTfL7tE"
# ### Extract parameters and metrics into a dataframe for analysis
# + [markdown] id="A1PqKxlpOZa2"
# We can also extract all parameters and metrics associated with any Experiment into a dataframe for further analysis.
# + id="jbRf1WoH_vbY"
experiment_df = aiplatform.get_experiment_df()
experiment_df
# + [markdown] id="EYuYgqVCMKU1"
# ### Visualizing an experiment's parameters and metrics
# + id="r8orCj8iJuO1"
plt.rcParams["figure.figsize"] = [15, 5]
ax = pd.plotting.parallel_coordinates(
experiment_df.reset_index(level=0),
"run_name",
cols=[
"param.num_units",
"param.dropout_rate",
"param.epochs",
"metric.loss",
"metric.val_loss",
"metric.eval_loss",
],
color=["blue", "green", "pink", "red"],
)
ax.set_yscale("symlog")
ax.legend(bbox_to_anchor=(1.0, 0.5))
# + [markdown] id="WTHvPMweMlP1"
# ## Visualizing experiments in Cloud Console
# + [markdown] id="F19_5lw0MqXv"
# Run the following to get the URL of Vertex AI Experiments for your project.
#
# + id="GmN9vE9pqqzt"
print("Vertex AI Experiments:")
print(
f"https://console.cloud.google.com/ai/platform/experiments/experiments?folder=&organizationId=&project={PROJECT_ID}"
)
# + [markdown] id="TpV-iwP9qw9c"
# ## Cleaning up
#
# To clean up all Google Cloud resources used in this project, you can [delete the Google Cloud
# project](https://cloud.google.com/resource-manager/docs/creating-managing-projects#shutting_down_projects) you used for the tutorial.
#
# Otherwise, you can delete the individual resources you created in this tutorial:
#
# - Experiment (Can be deleted manually in the GCP Console UI)
| notebooks/official/ml_metadata/sdk-metric-parameter-tracking-for-locally-trained-models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: kgtk-env
# language: python
# name: kgtk-env
# ---
# # Generating Subsets of Wikidata
# ### Batch Invocation
# Example batch command. The second argument is a notebook where the output will be stored. You can load it to see progress.
#
# UPDATE EXAMPLE INVOCATION
#
#
# ```
# papermill Wikidata\ Useful\ Files.ipynb useful-files.out.ipynb \
# -p wiki_file /Volumes/GoogleDrive/Shared\ drives/KGTK-public-graphs/wikidata-20200803-v3/all.tsv.gz \
# -p label_file /Volumes/GoogleDrive/Shared\ drives/KGTK-public-graphs/wikidata-20200803-v3/part.label.en.tsv.gz \
# -p item_file /Volumes/GoogleDrive/Shared\ drives/KGTK-public-graphs/wikidata-20200803-v3/part.wikibase-item.tsv.gz \
# -p property_item_file /Volumes/GoogleDrive/Shared\ drives/KGTK-public-graphs/wikidata-20200803-v3/part.property.wikibase-item.tsv.gz \
# -p qual_file /Volumes/GoogleDrive/Shared\ drives/KGTK-public-graphs/wikidata-20200803-v3/qual.tsv.gz \
# -p output_path <local folder> \
# -p output_folder useful_files_v4 \
# -p temp_folder temp.useful_files_v4 \
# -p delete_database no \
# -p compute_pagerank no \
# -p languages es,ru,zh-cn
# ```
# +
import io
import os
import subprocess
import sys
import numpy as np
import pandas as pd
import papermill as pm
from kgtk.configure_kgtk_notebooks import ConfigureKGTK
from kgtk.functions import kgtk, kypher
# + tags=["parameters"]
input_path = "/data/amandeep/wikidata-20220505/import-wikidata/data"
output_path = "/data/amandeep"
kgtk_path = "/data/amandeep/Github/kgtk"
graph_cache_path = None
project_name = "wikidata-20220505-dwd-v4"
files = 'isa,p279star'
# Classes to remove
remove_classes = "Q7318358,Q13442814"
useful_files_notebook = "Wikidata-Useful-Files.ipynb"
notebooks_folder = f"{kgtk_path}/use-cases"
languages = "en,ru,es,zh-cn,de,it,nl,pl,fr,pt,sv"
debug = False
# -
files = files.split(',')
languages = languages.split(',')
ck = ConfigureKGTK(files, kgtk_path=kgtk_path)
ck.configure_kgtk(input_graph_path=input_path,
output_path=output_path,
project_name=project_name,
graph_cache_path=graph_cache_path)
ck.print_env_variables()
ck.load_files_into_cache()
# ### Preview the input files
# It is always a good practice to peek a the files to make sure the column headings are what we expect
# !zcat $claims | head
# ## Creating a list of all the items we want to remove
# ### Compute the items to be removed
# Compose the kypher command to remove the classes
# !zcat $isa | head | col
# Run the command, the items to remove will be in file `{temp}/items.remove.tsv.gz`
# +
classes = ", ".join(list(map(lambda x: '"{}"'.format(x), remove_classes.replace(" ", "").split(","))))
classes
# -
kypher(f""" -i isa -i p279star -o "$TEMP"/items.remove.tsv.gz
--match 'isa: (n1)-[:isa]->(c), p279star: (c)-[]->(class)'
--where 'class in [{classes}]'
--return 'distinct n1, "p31_p279star" as label, class as node2'
--order-by 'n1'
""")
# Preview the file
# !zcat < "$TEMP"/items.remove.tsv.gz | head | col
# !zcat < "$TEMP"/items.remove.tsv.gz | wc
# Collect all the classes of items we will remove, just as a sanity check
# !$kypher -i "$TEMP"/items.remove.tsv.gz \
# --match '()-[]->(n2)' \
# --return 'distinct n2' \
# --limit 10
# ## Create the reduced edges file
# ### Remove the items from the all.tsv and the label, alias and description files
# We will be left with `reduced` files where the edges do not have the unwanted items. We have to remove them from the node1 and node2 positions, so we need to run the ifnotexists commands twice.
#
# Before we start preview the files to see the column headings and check whether they look sorted.
# !zcat "$TEMP"/items.remove.tsv.gz | head | col
# Remove from the full set of edges those edges that have a `node1` present in `items.remove.tsv`
kgtk("""ifnotexists
-i $claims
-o "$TEMP"/item.edges.reduced.tsv.gz
--filter-on "$TEMP"/items.remove.tsv.gz
--input-keys node1
--filter-keys node1
--presorted
""")
# From the remaining edges, remove those that have a `node2` present in `items.remove.tsv`
kgtk(f"""sort
-i "$TEMP"/item.edges.reduced.tsv.gz
-o "$TEMP"/item.edges.reduced.sorted.tsv.gz
--extra '--parallel 24 --buffer-size 30% --temporary-directory {os.environ['TEMP']}'
--columns node2 label node1 id""")
kgtk("""ifnotexists
-i $TEMP/item.edges.reduced.sorted.tsv.gz
-o $TEMP/item.edges.reduced.2.tsv.gz
--filter-on $TEMP/items.remove.tsv.gz
--input-keys node2
--filter-keys node1
--presorted""")
# Create a file with the labels, for all the languages specified, **FIX THIS**
kgtk("""ifnotexists -i $label_all
-o "$TEMP"/label.all.edges.reduced.tsv.gz
--filter-on "$TEMP"/items.remove.tsv.gz
--input-keys node1
--filter-keys node1
--presorted""")
kgtk(f"""sort
-i $TEMP/label.all.edges.reduced.tsv.gz
--extra '--parallel 24 --buffer-size 30% --temporary-directory {os.environ['TEMP']}'
-o $OUT/labels.tsv.gz""")
# Create a file with the aliases, for all the languages specified
kgtk("""ifnotexists -i $alias_all
-o $TEMP/alias.all.edges.reduced.tsv.gz
--filter-on $TEMP/items.remove.tsv.gz
--input-keys node1
--filter-keys node1
--presorted""")
kgtk(f"""sort
-i $TEMP/alias.all.edges.reduced.tsv.gz
--extra '--parallel 24 --buffer-size 30% --temporary-directory {os.environ['TEMP']}'
-o $OUT/aliases.tsv.gz""")
# Create a file with the descriptions, for all the languages specified
kgtk("""ifnotexists
-i $description_all
-o $TEMP/description.all.edges.reduced.tsv.gz
--filter-on $TEMP/items.remove.tsv.gz
--input-keys node1
--filter-keys node1
--presorted""")
kgtk(f"""sort
-i $TEMP/description.all.edges.reduced.tsv.gz
--extra '--parallel 24 --buffer-size 30% --temporary-directory {os.environ['TEMP']}'
-o $OUT/descriptions.tsv.gz""")
# ### Produce the output files for claims, labels, aliases and descriptions
kgtk(f"""sort
-i $TEMP/item.edges.reduced.2.tsv.gz
--extra '--parallel 24 --buffer-size 30% --temporary-directory {os.environ['TEMP']}'
-o $OUT/claims.tsv.gz""")
# ## Create the reduced qualifiers file
# We do this by finding all the ids of the reduced edges file, and then selecting out from `qual.tsv`
#
# We need to join by id, so we need to sort both files by id, node1, label, node2:
#
# - `$qualifiers`
# - `$OUT/claims.tsv.gz`
if debug:
# !zcat < "$qualifiers" | head | column -t -s $'\t'
# Run `ifexists` to select out the quals for the edges in `{out}/wikidataos.qual.tsv.gz`. Note that we use `node1` in the qualifier file, matching to `id` in the `wikidataos.all.tsv` file.
kgtk("""ifexists
-i $qualifiers
-o $OUT/qualifiers.tsv.gz
--filter-on $OUT/claims.tsv.gz
--input-keys node1
--filter-keys id
--presorted""")
# Look at the final output for qualifiers
if debug:
# !zcat $OUT/qualifiers.tsv.gz | head | col
# !ls -l "$OUT"
# Copy the property datatypes and metadata types file over
# !cp $datatypes $OUT/metadata.property.datatypes.tsv.gz
# Filter out edges from metdata types file
kgtk("""ifexists
-i "$types" -o $OUT/metadata.types.tsv.gz
--filter-on $OUT/claims.tsv.gz
--input-keys node1
--filter-keys node1
--presorted""")
# Get the sitelinks as well, the sitelinks are not in claims.tsv.gz
kgtk("""ifexists
-i "$GRAPH/sitelinks.tsv.gz"
-o "$OUT/sitelinks.tsv.gz"
--filter-on "$OUT/claims.tsv.gz"
--input-keys node1
--filter-keys node1
--presorted""")
# Contruct the cat command to generate `all.tsv.gz`
kgtk("""cat -i "$OUT/labels.tsv.gz"
-i "$OUT/aliases.tsv.gz"
-i "$OUT/descriptions.tsv.gz"
-i "$OUT/claims.tsv.gz"
-i "$OUT/qualifiers.tsv.gz"
-i "$OUT/metadata.property.datatypes.tsv.gz"
-i "$OUT/metadata.types.tsv.gz"
-i "$OUT/sitelinks.tsv.gz"
-o "$OUT/all.tsv.gz"
""")
# ### Run the Partitions Notebook
pm.execute_notebook(
"partition-wikidata.ipynb",
os.environ["TEMP"] + "/partition-wikidata.out.ipynb",
parameters=dict(
wikidata_input_path = os.environ["OUT"] + "/all.tsv.gz",
wikidata_parts_path = os.environ["OUT"] + "/parts",
temp_folder_path = os.environ["OUT"] + "/parts/temp",
sort_extras = "--buffer-size 30% --temporary-directory $OUT/parts/temp",
verbose = False,
gzip_command = 'gzip'
)
)
# # ### copy the `claims.wikibase-item.tsv` file from the `parts` folder
# !cp $OUT/parts/claims.wikibase-item.tsv.gz $OUT
# ### RUN the Useful Files notebook
pm.execute_notebook(
f'{useful_files_notebook}',
os.environ["TEMP"] + "/Wikidata-Useful-Files-Out.ipynb",
parameters=dict(
output_path = os.environ["OUT"],
input_path = os.environ["OUT"],
kgtk_path = kgtk_path,
compute_pagerank=True,
compute_degrees=True,
compute_isa_star=True,
compute_p31p279_star=True,
debug=False
)
)
# ## Sanity checks
if debug:
# !$kypher -i $OUT/claims.tsv.gz \
# --match '(n1:Q368441)-[l]->(n2)' \
# --limit 10 \
# | col
if debug:
# !$kypher -i $OUT/claims.tsv.gz \
# --match '(n1:P131)-[l]->(n2)' \
# --limit 10 \
# | col
# ## Summary of results
# !ls -lh $OUT/*.tsv.gz
| use-cases/create_wikidata/Wikidata-Subsets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.8 64-bit (''atcs-practical'': conda)'
# name: python388jvsc74a57bd00b7cfd85d2fc43f76dbccc53202d8ba6a9a8cca408d693df58307e7c75a304a7
# ---
# ## Code to download files from wandb that were created during lisa runtime
# Only used once and require to be loggined witht he right wandb account.
# %pwd #look at the current work dir
# %cd ..
import wandb
import os
api = wandb.Api()
# +
id_run = '1ae4n1th'
model = 'awe'
directory = 'trained_models/'+model+'/gold'
if not os.path.exists(directory):
os.makedirs(directory)
os.chdir(directory)
run = api.run("rodrigochavez/atcs-practical/"+id_run)
for file in run.files():
file.download()
# +
id_run = '8tzcbeam'
model = 'lstm'
directory = 'trained_models/'+model+'/gold'
if not os.path.exists(directory):
os.makedirs(directory)
os.chdir(directory)
run = api.run("rodrigochavez/atcs-practical/"+id_run)
for file in run.files():
file.download()
# +
id_run = '1p3zope5'
model = 'bilstm'
directory = 'trained_models/'+model+'/gold'
if not os.path.exists(directory):
os.makedirs(directory)
os.chdir(directory)
run = api.run("rodrigochavez/atcs-practical/"+id_run)
for file in run.files():
file.download()
# +
id_run = '3jwpgfe9'
model = 'bilstm-max'
directory = 'trained_models/'+model+'/gold'
if not os.path.exists(directory):
os.makedirs(directory)
os.chdir(directory)
run = api.run("rodrigochavez/atcs-practical/"+id_run)
for file in run.files():
file.download()
| dev_notebooks/wandb.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
#
# # 01. Introduction to Python
#
#
# ## Python Programming
#
# Python is a powerful multipurpose programming language created by <NAME>.
#
# It has a simple and easy-to-use syntax, making it a popular first-choice programming language for beginners.
#
# This is a comprehensive guide that explores the reasons you should consider learning Python and the ways you can get started with Python.
#
#
#
# ## What is Python Programming Language?
#
# Python is an interpreted, object-oriented, high-level programming language. As it is general-purpose, it has a wide range of applications from web development, building desktop GUI to scientific and mathematical computing.
#
# Python is popular for its simple and relatively straightforward syntax. Its syntax readability increases productivity as it allows us to focus more on the problem rather than structuring the code.
#
# ## Features of Python Programming
#
# Simple and easy to learn Python has a very simple and elegant syntax. It is much easier to read and write programs in Python compared to other languages like C, C++, or Java.
#
# Due to this reason, many beginners are introduced to programming with Python as their first programming language.
#
# ### Free and open-source
#
# You can freely use and distribute Python programs even for commercial use. As it is open-source, you can even change Python's source code to fit your use case.
#
# ### Portability
#
# A single Python program can run on different platforms without any change in source code. It runs on almost all platforms including Windows, Mac OS X, and Linux.
#
# ### Extensible and Embeddable
#
# You can combine Python code with other programming languages like C or Java to increase efficiency. This allows high performance and scripting capabilities that other languages do not provide out of the box.
#
# ### High-Level Interpreted Language
#
# Python itself handles tasks like memory management and garbage collection. So unlike C or C++, you don't have to worry about system architecture or any other lower-level operations.
#
# ### Rich library and large community
#
# Python has numerous reliable built-in libraries. Python programmers have developed tons of free and open-source libraries, so you don't have to code everything by yourself.
#
# The Python community is very large and evergrowing. If you encounter errors while programming in Python, it's like that it has already been asked and solved by someone in this community.
#
# Reasons to Choose Python as First Language
#
# **1. Simple Elegant Syntax**
#
# Programming in Python is fun. It's easier to understand and write Python code. The syntax feels natural. Let's take the following example where we add two numbers:
#
# ```python
# a = 2
# b = 3
# sum = a + b
# print(sum)
# ```
# Even if you have never programmed before, you can easily guess that this program adds two numbers and displays it.
#
# **2. Not overly strict**
#
# You don't need to define the type of a variable in Python. Also, it's not necessary to add a semicolon at the end of the statement.
#
# Python enforces you to follow good practices (like proper indentation). These small things can make learning much easier for beginners.
#
# **3. The expressiveness of the language**
#
# Python allows you to write programs having greater functionality with fewer lines of code. Let's look at code to swap the values of two variables. It can be done in Python with the following lines of code:
#
# ```python
# a = 15
# b = 27
# print(f'Before swapping: a, b = {a},{b}')
# a, b = b, a
# print(f'After swapping: a, b = {a},{b}')
# ```
#
# Here, we can see that the code is very less and more readable.
#
# If instead, we were to use Java, the same program would have to be written in the following way:
#
# ```java
# public class Swap {
# public static void main(String[] args) {
# int a, b, temp;
# a = 15;
# b = 27;
# System.out.println("Before swapping : a, b = "+a+", "+ + b);
# temp = a;
# a = b;
# b = temp;
# System.out.println("After swapping : a, b = "+a+", "+ + b);
# }
# }
#
# ```
# This is just an example. There are many more such cases where Python increases efficiency by reducing the amount of code required to program something.
#
# ## Python Applications Area
#
# Python is known for its general purpose nature that makes it applicable in almost each domain of software development. Python as a whole can be used in any sphere of development.
#
# Here, we are specifing applications areas where python can be applied.
#
# **1. Web Applications**
#
# We can use Python to develop web applications. It provides libraries to handle internet protocols such as HTML and XML, JSON, Email processing, request, beautifulSoup, Feedparser etc. It also provides Frameworks such as Django, Pyramid, Flask etc to design and delelop web based applications. Some important developments are: PythonWikiEngines, Pocoo, PythonBlogSoftware etc.
#
# **2. AI & Machine Learning**
#
# Python has Prebuilt Libraries like Numpy for scientific computation, Scipy for advanced computing and Pybrain for machine learning (Python Machine Learning) making it one of the best languages For AI.
#
# **3.Desktop GUI Applications**
#
# Python provides Tk GUI library to develop user interface in python based application. Some other useful toolkits wxWidgets, Kivy, pyqt that are useable on several platforms. The Kivy is popular for writing multitouch applications.
#
# **4.Software Development**
#
# Python is helpful for software development process. It works as a support language and can be used for build control and management, testing etc.
#
# **5.Scientific and Numeric**
#
# Python is popular and widely used in scientific and numeric computing. Some useful library and package are SciPy, Pandas, IPython etc. SciPy is group of packages of engineering, science and mathematics.
#
# **6.Business Applications**
#
# Python is used to build Bussiness applications like ERP and e-commerce systems. Tryton is a high level application platform.
#
# **7.Console Based Application**
#
# We can use Python to develop console based applications. For example: IPython.
#
# **8.Audio or Video based Applications**
#
# Python is awesome to perform multiple tasks and can be used to develop multimedia applications. Some of real applications are: TimPlayer, cplay etc.
#
# **9.3D CAD Applications**
#
# To create CAD application Fandango is a real application which provides full features of CAD.
#
# **10.Enterprise Applications**
#
# Python can be used to create applications which can be used within an Enterprise or an Organization. Some real time applications are: OpenErp, Tryton, Picalo etc.
#
# **12.Applications for Images**
#
# Using Python several application can be developed for image. Applications developed are: VPython, Gogh, imgSeek etc.
#
# **13.Games and 3D Graphics**
#
# PyGame, PyKyra are two frameworks for game-development with Python. Apart from these, we also get a variety of 3D-rendering libraries. If you’re one of those game-developers, you can check out PyWeek, a semi-annual game programming contest.
#
| 01. Introduction to Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Data cleaning
# - Refer to data dictionary on the Github repository
# +
import pandas as pd
import numpy as np
import pickle
from sklearn.impute import SimpleImputer
# -
df_train = pd.read_csv("../kaggle-california-housing-data/train.csv")
# ### Inspect columns and values
# - Let's first take a look at the null values
# - Make sure that a null value makes sense in these columns
# - Then decide on a function to replace
# - NaN means this in the columns:
# - LotFrontage: No street connected to the property
# - Alley: NaN means no Alley
# - MasVnrType: Not sure, already has a None category
# - MasVnrArea: Not sure, already has a 0 area
# - BsmtQual: No basement
# - BsmtFinType1: No basement
# - BsmtFinType2: No basement
# - Electrical: Not sure, no electrical system?
# - FireplaceQu: No fireplace
# - GarageType: No garage
# - GarageYrBlt: No garage (check)
# - GarageFinish: No garage (check)
# - GarageQual: No garage (check)
# - GarageCond: No garage (check)
# - PoolQC: No pool
# - Fence: No fence
# - MiscFeature: No miscellaneous features
df_train.columns[
list(
df_train.isnull().any()
)
]
# ### Individual checks
# - Make sure that the NaNs are consistent
df_train.loc[
df_train["BsmtQual"].isnull(),
["BsmtFinType1", "BsmtFinType2"]
].drop_duplicates()
df_train.loc[
df_train["GarageType"].isnull(),
[
"GarageYrBlt",
"GarageFinish",
"GarageQual",
"GarageCond"
]
].drop_duplicates()
# ## Impute values
# - LotFrontage will be changed to 0
# - MasVnrType will be changed to "None"
# - MasVnrArea will be changed to 0
# - Everything elese will be changed to "None"
def replace_nans(df, col, method):
imputer = SimpleImputer(
missing_values=np.nan,
strategy="constant",
fill_value=method
)
return imputer.fit_transform(np.array(df[col]).reshape(1, -1))[0]
df_train["LotFrontage"] = replace_nans(df_train, "LotFrontage", 0)
df_train["MasVnrArea"] = replace_nans(df_train, "MasVnrArea", 0)
df_train = df_train.loc[
df_train["Electrical"].notnull(),
:
].fillna("None")
# ## Export cleaned data
with open('../kaggle-california-housing-data/df_imputed.pickle', 'wb') as f:
pickle.dump(df_train, f)
| 2. Data cleaning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Further diagnosis
# In the previous exercise, you identified some potentially unclean or missing data. Now, you'll continue to diagnose your data with the very useful .info() method.
#
# The .info() method provides important information about a DataFrame, such as the number of rows, number of columns, number of non-missing values in each column, and the data type stored in each column. This is the kind of information that will allow you to confirm whether the 'Initial Cost' and 'Total Est. Fee' columns are numeric or strings. From the results, you'll also be able to see whether or not all columns have complete data in them.
#
# The full DataFrame df and the subset DataFrame df_subset have been pre-loaded. Your task is to use the .info() method on these and analyze the results.
#
# ### Instructions
#
# - Print the info of df.
# - Print the info of the subset dataframe, df_subset.
# +
# Import pandas
import pandas as pd
# Read the file into a DataFrame: df
df = pd.read_csv('dob_job_application_filings_subset.csv')
# Print the info of df
print(df.info())
# -
| cleaning data in python/Exploring your data/02. Further diagnosis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <div align="center">
# <img src="logoinpe.png">
# </div>
#
#
# # Análise Estatística e Espectral de Processos Estocásticos (CAP-239-4)
#
# <hr>
#
# Docentes:
# - Dr. <NAME>
# - Dr. <NAME>
#
# Discentes:
# - <NAME>;
# - <NAME>;
# - <NAME>.
# **Atividade**: Ajustes de Função de Densidade de Probabilidade (PDF)
#
# **Descrição da atividade**: Ajustar as melhores PDFs para cada um dos dados considerados neste trabalho. O ajuste deve ser feito somente para os dados de flutuação, uma vez que, a caracterização base já descreve o comportamento geral dos dados acumulados, que não possuem flutuações. Assim, as variáveis consideradas nesta atividade serão o `Número Diário de Casos (NDC)`, `Número Diário de Mortes (NDM)` e `Número Diário de Testes (NDT)`. Para este documento é feito a caracterização dos dados considerando a análise de Cullen-Frey
# +
import scipy.stats
import numpy as np
import pandas as pd
from plotnine import *
from fitter import Fitter
import matplotlib.pyplot as plt
import matplotlib.dates as mdates
import warnings
warnings.filterwarnings("ignore")
# +
data_owd = pd.read_csv('../1_conjunto_de_dados/dataset/data/blueteam_international_update_2020_06_03.csv')
data_owd['date'] = pd.to_datetime(data_owd['date'])
data_owd = data_owd[data_owd['date'] >= '2020-04-01']
# -
# Para a realização do ajuste das PDFs de maneira automatizada será feita a utilização do pacote [fitter](https://fitter.readthedocs.io/en/latest/), que através de uma interface simples permite o teste e verificação de mais de 80 tipos de distribuições, essas providas pelo pacote [SciPy](https://www.scipy.org/).
#
# Abaixo as divisões são feitas considerandos cada um dos países analisados neste trabalho.
# ### Brasil
bra = data_owd[data_owd['iso_code'] == 'BRA']
# **Número de casos diários**
bra_newcases = bra.new_cases
f = Fitter(bra_newcases, verbose = False, distributions = ['beta', 'gamma'], bins = 10)
f.fit()
plt.figure(dpi = 150)
f.summary()
plt.title("Número de casos diários (Brasil)")
plt.xlabel("Quantidade de casos")
plt.ylabel("Densidade")
plt.show()
f.get_best()
# **Número de mortes diárias**
bra_newdeaths = bra.new_deaths
f = Fitter(bra_newdeaths, verbose = False, bins = 15, distributions = ['uniform', 'beta'])
f.fit()
plt.figure(dpi = 150)
f.summary()
plt.title("Mortes diárias (Brasil)")
plt.xlabel("Quantidade de mortes")
plt.ylabel("Densidade")
plt.show()
f.get_best()
# **Número de testes diários**
# +
# Infelizmente não há testes suficientes para este ajuste
# -
# ### Canadá
can = data_owd[data_owd['iso_code'] == 'CAN']
# **Número de casos diários**
can_newcases = can.new_cases
f = Fitter(can_newcases, verbose = False, bins = 25, distributions = ['beta', 'lognorm', 'gamma', 'norm'])
f.fit()
plt.figure(dpi = 150)
f.summary()
plt.title("Casos diários (Canadá)")
plt.xlabel("Quantidade de casos")
plt.ylabel("Densidade")
plt.show()
f.get_best()
# **Número de mortes diárias**
can_newdeaths = can.new_deaths
f = Fitter(can_newdeaths, verbose = False, bins = 18, distributions = ['beta', 'uniform', 'norm'])
f.fit()
plt.figure(dpi = 150)
f.summary()
plt.title("Mortes diárias (Canadá)")
plt.xlabel("Quantidade de mortes")
plt.ylabel("Densidade")
plt.show()
f.get_best()
# **Número de testes diários**
can_newtests = can[~can.new_tests.isna()].new_tests
f = Fitter(can_newtests, verbose = False, bins = 30, distributions = ['lognorm', 'norm', 'gamma', 'beta'])
f.fit()
plt.figure(dpi = 150)
f.summary()
plt.title("Testes diários (Canadá)")
plt.xlabel("Quantidade de testes")
plt.ylabel("Densidade")
plt.show()
f.get_best()
# ### México
mex = data_owd[data_owd['iso_code'] == 'MEX']
# **Número de casos diários**
mex_newcases = mex.new_cases
f = Fitter(mex_newcases, verbose = False, bins = 20, distributions = ['beta', 'uniform'])
f.fit()
plt.figure(dpi = 150)
f.summary()
plt.title("Casos diários (México)")
plt.xlabel("Quantidade de casos")
plt.ylabel("Densidade")
plt.show()
f.get_best()
# **Número de mortes diárias**
mex_newdeaths = mex.new_deaths
f = Fitter(mex_newdeaths, verbose = False, bins = 18, distributions = ['beta'])
f.fit()
plt.figure(dpi = 150)
f.summary()
plt.title("Mortes diárias (México)")
plt.xlabel("Quantidade de mortes")
plt.ylabel("Densidade")
plt.show()
f.get_best()
# **Número de testes diários**
mex_newtests = mex[~mex.new_tests.isna()].new_tests
f = Fitter(mex_newtests, verbose = False, bins = 25, distributions = ['beta', 'uniform'])
f.fit()
plt.figure(dpi = 150)
f.summary()
plt.title("Testes diários (México)")
plt.xlabel("Quantidade de testes")
plt.ylabel("Densidade")
plt.show()
f.get_best()
# ### Cuba
cub = data_owd[data_owd['iso_code'] == 'CUB']
# **Número de casos diários**
cub_newcases = cub.new_cases
f = Fitter(cub_newcases, verbose = False, bins = 18, distributions = ['uniform', 'beta'])
f.fit()
plt.figure(dpi = 150)
f.summary()
plt.title("Casos diários (Cuba)")
plt.xlabel("Quantidade de casos")
plt.ylabel("Densidade")
plt.show()
f.get_best()
# **Número de mortes diárias**
cub_newdeaths = cub.new_deaths
f = Fitter(cub_newdeaths, verbose = False, bins = 10, distributions = ['beta', 'gamma'])
f.fit()
plt.figure(dpi = 150)
f.summary()
plt.title("Mortes diárias (Cuba)")
plt.xlabel("Quantidade de mortes")
plt.ylabel("Densidade")
plt.show()
f.get_best()
# **Número de testes diários**
cub_newtests = cub[~cub.new_tests.isna()].new_tests
f = Fitter(cub_newtests, verbose = False, bins = 15, distributions = ['beta', 'gamma'])
f.fit()
plt.figure(dpi = 150)
f.summary()
plt.title("Testes diários (Cuba)")
plt.xlabel("Quantidade de testes")
plt.ylabel("Densidade")
plt.show()
f.get_best()
# ### Rússia
rus = data_owd[data_owd['iso_code'] == 'RUS']
# **Número de casos diários**
rus_newcases = rus.new_cases
f = Fitter(rus_newcases, verbose = False, bins = 20, distributions = ['norm', 'uniform', 'beta', 'gamma'])
f.fit()
plt.figure(dpi = 150)
f.summary()
plt.title("Casos diários (Rússia)")
plt.xlabel("Quantidade de casos")
plt.ylabel("Densidade")
plt.show()
f.get_best()
# **Número de mortes diárias**
rus_newdeaths = rus.new_deaths
f = Fitter(rus_newdeaths, verbose = False, bins = 20, distributions = ['lognorm', 'gamma', 'beta'])
f.fit()
plt.figure(dpi = 150)
f.summary()
plt.title("Mortes diárias (Rússia)")
plt.xlabel("Quantidade de mortes")
plt.ylabel("Densidade")
plt.show()
f.get_best()
# **Número de testes diários**
rus_newtests = rus[~rus.new_tests.isna()].new_tests
f = Fitter(rus_newtests, verbose = False, bins = 15, distributions = ['beta', 'uniform'])
f.fit()
plt.figure(dpi = 150)
f.summary()
plt.title("Testes diários (Rússia)")
plt.xlabel("Quantidade de testes")
plt.ylabel("Densidade")
plt.show()
f.get_best()
| 2_analise_visualizacao_dos_dados/3_ajustes_pdf_somente_cullen-frey.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="BJEEyu81m3l6" colab_type="text"
# # Notebook 1: Python Overview
#
# ## Motivations
#
# Spark provides multiple *Application Programming Interfaces* (API), i.e. the interface allowing the user to interact with the application. The main APIs are [Scala](http://spark.apache.org/docs/latest/api/scala/index.html#org.apache.spark.package) and [Java](http://spark.apache.org/docs/latest/api/java/index.html) APIs, as Spark is implemented in Scala and runs on the Java Virtual Machine (JVM).
# Since the 0.7.0 version, a [Python API](http://spark.apache.org/docs/latest/api/python/index.html) is available, also known as PySpark. An [R API](http://spark.apache.org/docs/latest/api/R/index.html) has been released with 1.5.0 version. During this course, you will be using Spark 2.3.
#
# Throughout this course we will use the Python API for the following reasons:
# - R API is still too young and limited to be relied on. Besides, R can quickly become a living hell when using immature libraries.
# - Many of you are wanabee datascientists, and Python is a must-know language in data industry.
# - Scala and Java APIs would have been quite hard to learn given the length of the course and your actual programming skills.
# - Python is easy to learn, and even easier if you are already familiar with R.
#
# The goal of this session is to teach (or remind) you the syntax of basic operations, control structures and declarations in Python that will be useful when using Spark. Keep in mind that we do not have a lot of time, and that you should be able to create functions and classes and to manipulate them at the end of the lab. If you don't get that, the rest of the course will be hard to follow. Don't hesitate to ask for explanations and/or more exercises if you don't feel confident enough at the end of the lab.
#
#
# *This introduction relies on [Learn Python in Y minutes](https://learnxinyminutes.com/docs/python/)*
#
# ## Introduction
#
# Python is a high level, general-purpose interpreted language. Python is meant to be very concise and readable, it is thus a very pleasant language to work with.
#
# ## 1. Primitive Datatypes and Operators
# ### Points: 1 pt
# Read section 1 of [Learn python in Y Minutes](https://learnxinyminutes.com/docs/python/) (if you already know Python, you can skip this step). Then, replace `???` in the following cells with your code to answer the questions. To get started, please run the following cell.
# + id="iXOJzAEQm3l9" colab_type="code" colab={}
# Run this cell, it loads a Test object
# that will allow you to check your code
# for some questions.
import unittest
test = unittest.TestCase()
# + [markdown] id="oHP613V5m3mD" colab_type="text"
# Compute 4 + 8
#
# + id="jNlLR5Xom3mF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="6676f2b0-6988-48da-e3d4-d50c2bf22018"
4+8
# + [markdown] id="B9HJdIQpm3mI" colab_type="text"
# Compute 4 * 8
# + id="dibZFDAsm3mJ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="82cb444d-8969-49bc-f9ab-c4bddf4cc333"
4*8
# + [markdown] id="KntXNdCXm3mL" colab_type="text"
# Compute 4 / 8 (using the regular division operation, not integer division)
# + id="XXRljLwfm3mN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="c82c980a-ec1c-4d56-97e7-4941dece5387"
4/8
# + [markdown] id="MOaJMGSVm3mQ" colab_type="text"
# Check if the variable `foo` is None:
# + id="VOfwxqRLm3mR" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="4e723f9a-5863-482e-b427-2268cf55600b"
foo = None
foo == None # => True
# + [markdown] id="kgbbTyM0m3mX" colab_type="text"
# ## 2. Variables and Collections
# ### Points: 2 pts
# ### Bonus: 2 pts
# Same as before, read the corresponding section, and answer the questions below.
# + [markdown] id="MrqWUzFmm3mY" colab_type="text"
# From now on, when you will be asked to print something, please use the print statement.
#
# Declare a variable containing a float of your choice and print it.
# + id="OP7ZW0y7m3mZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="15e95df1-7c0f-45fe-dd59-e6da65cd90ef"
Variable = 0.5
print(Variable)
# + id="5YYupfr4m3mb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="23d3642b-7963-4695-a40b-558b8eb926d5"
# Create a list containing strings and store it in a variable
bar = ["Tree","Lake","Animal"]
# Append a new string to this list
bar.append("Climat")
# Append an integer to this list and print it
bar.append(6)
print(bar)
# + [markdown] id="_8MV7OTlm3me" colab_type="text"
# Note that the modifications on list objects are performed inplace, i.e.
#
# li = [1, 2, 3]
# li.append(4)
# li # => [1, 2, 3, 4]
# + id="UV5xebqIm3mf" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="a24184db-b34f-4328-cd04-c5ce469b4e20"
# Mixing types inside a list object can be a bad idea depending on the situation.
# Remove the integer you just inserted in the list and print it
bar.pop(-1)
print(bar)
# + id="ItPQyQoWm3mk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="a94a4682-4365-480d-80e7-fcafe8ecc503"
# Print the second element of the list
print(bar[1])
# + [markdown] id="vvWwawcPm3mn" colab_type="text"
# You can access list elements in reverse order, e.g.
#
# li[-1] # returns the last element of the list
# li[-2] # returns the second last element of the list
#
# and so on...
# + id="man-DHBTm3mo" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="aa3a4e57-e0cb-494b-9593-d0b05e7f7f80"
# Extend your list with new_list and print it
new_list = ["We", "are", "the", "knights", "who", "say", "Ni", "!"]
bar = bar + new_list
print(bar)
# + id="y5-62URlm3mr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="11f68c0a-01f1-4696-9e85-a9dfe6fec7f6"
# Replace "Ni" by "Ekke Ekke Ekke Ekke Ptang Zoo Boing" in the list and print it
bar[-2] = "Ekke Ekke Ekke Ekke Ptang Zoo Boing"
print(bar)
# + id="-68ZURjxm3mu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="ac84c90a-8d44-4687-a10f-3fba3b7c0854"
# Compute the length of the list and print it
Length_List = len(bar)
print(Length_List)
# + [markdown] id="gqg2svEUm3mx" colab_type="text"
# What is the difference between lists and tuples?
#
#
# The elements of a list can be modified whereas the elements of
# a tuple can not be modified.
#
# + id="IB0jTJlCm3mz" colab_type="code" colab={}
# Create a dictionary containing the following mapping:
# "one" : 1
# "two" : 2
# etc. until you reach "five" : 5
baz = {"one":1, "two":2, "three":3,"four":4, "five":5}
# + id="-BfZ0C_Jm3m4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="7ae5ae3d-7ebd-4a08-d453-5a5328edd78a"
# Check if the key "four" is contained in the dict
# If four is contained in the dict, print the associated value
baz.keys()
print(baz["four"])
# + [markdown] id="6fmemAhtm3m6" colab_type="text"
# First, make each letter in gibberish appear once using a collection.
# + id="YKMa7wZJm3m7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="36e83953-a955-40d0-f3f2-1099026de093"
gibberish = list("fqfgsrhrfeqluihjgrshioprqoqeionfvnorfiqeo")
from collections import Counter
New = Counter(gibberish).most_common(len(gibberish))
gibberish = gibberish[0:len(New)]
gibberish = [ New[i][0] for i in range(len(New))]
print(gibberish)
# + [markdown] id="ikUh-wOvm3m-" colab_type="text"
# *BONUS*: +2 pts Find all the unique letters contained in gibberish. Your answer must fits in one line.
# + id="_zGP53YGm3m_" colab_type="code" colab={}
# Hint: this import will help you with the bonus.
from collections import Counter
# + id="QcuZ0Hd3m3nC" colab_type="code" colab={}
test.assertEqual(set(unique_letters), set('vjlup'), 'unique letters')
# + [markdown] id="VzNrPPPfm3nL" colab_type="text"
# You should now be able to answer the following problem using dictionaries, lists and sets. Imagine you owe money to your friends because your forgot your credit card last time you went out for drinks. You want to remember how much you owe to each of them in order to refund them later. Which data structure would be useful to store this information? Use this data structure and fill it in with some debt data in the cell below:
# + id="qo_bJ8DWm3nM" colab_type="code" colab={}
debts = ???
# + [markdown] id="iBI69jMxm3nO" colab_type="text"
# Another party night with more people, yet you forgot your credit card again... You meet new friends who buy you drinks. Create the same data structure as above with different data, i.e. include friends that were not here during the first party.
# + id="VPLtXFszm3nP" colab_type="code" colab={}
debts_2 = ???
# + [markdown] id="ctIzFeY9m3nS" colab_type="text"
# Count the number of new friends you made that second night. Print the name of the friends who bought you drinks during the second party, but not during the first.
# + id="mUYV-9tSm3nT" colab_type="code" colab={}
new_friends = ??? # should fit in one line
nb_new_friends = ??? # should fit in one line
print(new_friends)
# + [markdown] id="3XXjCVOGm3nY" colab_type="text"
# ## 3. Control flow
# ### Points: 3 pts
# Same as before, read the corresponding section, and answer the questions below.
# You can skip the paragraph on exceptions for now.
# + id="mqFXOCdR1ZZT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="801d4779-aa97-428c-9ae7-ebaaf140d650"
# Code the following:
# if you have made more than 5 friends that second night,
# print "Yay! I'm super popular!", else, print "Duh..."
print(" Give the number of friend you made in the second night.")
Numb_Friend_Second_Night = input()
Numb_Friend_Second_Night = int(Numb_Friend_Second_Night)
if (Numb_Friend_Second_Night >= 5):
print(" Yay! I'm super popular ")
else:
print("Duh...")
# + id="Hy3l81ATm3nb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="4d27dabb-ea4d-42e2-d8c2-101938925021"
# Now, thank each new friend iteratively, i.e.
# print "Thanks <name of the friend>!" using loops and string formatting (cf. section 1)
Numb_Friend_Second_Night = 3
New_Friend = ["Jean","Jacques", "Marc"]
for i in New_Friend:
print("Thanks {0}".format(i))
# + id="KMvx50dTm3nd" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="13a15a58-d48b-4586-bb4e-af37f7b73289"
# Sum all the number from 0 to 15 (included) using what we've seen so far (i.e. without the function sum() )
sum_to_fifteen = 0
for i in range(15):
sum_to_fifteen += i
test.assertEquals(sum_to_fifteen, 105) # Instead of 120
# + id="3_DGV8TXm3nf" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="d9e8762b-d19b-4b08-a13e-85e57d483f7f"
# Note: you can break a loop with the break statement
for i in range(136):
print(i)
if i >= 2:
break
# + id="8KDuqK_Cm3ni" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="f12db24a-1c4f-45ad-aa26-a751a18d3cba"
# enumerate function can be very useful when dealing with iterators:
for i, value in enumerate(["a", "b", "c"]):
print(value, i)
# + [markdown] id="oRE5fA8Mm3nn" colab_type="text"
# __Q__: Find a Python function that allows to iterate over two collections in the same time stopping when the smallest collection is finished.
# + id="GnWPq2Aj9OEi" colab_type="code" colab={}
# + [markdown] id="fRRegQe8m3no" colab_type="text"
# ## 4. Functions
#
# ### Points: 5,5 pts
# Things are becoming more interesting. Read section 4. It's ok if you don't get the args/kwargs part. Be sure to understand basic function declaration and anonymous function declaration. Higher order functions, maps, and filters that will be covered during the next lab use massively lambda functions.
#
# Write a Python function that checks whether a passed string is palindrome or not. Note: a palindrome is a word, phrase, or sequence that reads the same backward and forward, e.g. "madam" or "nurses run". Hint: strings are lists of characters e.g.
#
# a = "abcdef"
# a[2] => c
#
# If needed, here are [some tips about string manipulation](http://www.pythonforbeginners.com/basics/string-manipulation-in-python).
# + id="v39huImBm3nq" colab_type="code" colab={}
def isPalindrome(string_input):
Bool_var = False
if string_input.count(' ') == 0:
N = len(string_input)-1
N =N/2
d = int(N)
for i in range(1,d+1):
if string_input[i] == string_input[-(i+1)] and string_input[0]==string_input[-1]:
Bool_var = True
else:
Bool_var = False
return Bool_var
else:
string_input = string_input.split(' ')
n = len(string_input)
New_String = []
for i in range(n):
New_String += string_input[i]
string_input = New_String
N = len(string_input)-1
N =N/2
d = int(N)
for i in range(1,d+1):
if string_input[i] == string_input[-(i+1)] and string_input[0]==string_input[-1]:
Bool_var = True
else:
Bool_var = False
return Bool_var
test.assertEqual(isPalindrome('aza'), True, "Simple palindrome")
test.assertEqual(isPalindrome('nurses run'), True, "Palindrome containing a space")
test.assertEqual(isPalindrome('palindrome'), False, "Not a palindrome")
# + [markdown] id="8JHAhaU3m3ns" colab_type="text"
# Write a Python function to check whether a string is pangram or not. Note: pangrams are words or sentences containing every letter of the alphabet at least once. For example: "The quick brown fox jumps over the lazy dog".
#
# [Hint](https://docs.python.org/2/library/stdtypes.html#set-types-set-frozenset)
# + id="5-xrgDGPm3nt" colab_type="code" colab={}
import string
# In this function, "alphabet" argument has a default value: string.ascii_lowercase
# string.ascii_lowercase contains all the letters in lowercase.
def ispangram(string_input, alphabet=string.ascii_lowercase):
# ???
test.assertEqual(ispangram('The quick brown fox jumps over the lazy dog'), True, "Pangram")
test.assertEqual(ispangram('The quick red fox jumps over the lazy dog'), False, "Pangram")
# + [markdown] id="irn1BgpWm3nw" colab_type="text"
# ### Python lambda expressions
#
# When evaluated, lambda expressions return an anonymous function, i.e. a function that is not bound to any variable (hence the "anonymous"). However, it is possible to assign the function to a variable. Lambda expressions are particularly useful when you need to pass a simple function into another function. To create lambda functions, we use the following syntax
#
# lambda argument1, argument2, argument3, etc. : body_of_the_function
#
# For example, a function which takes a number and returns its square would be
#
# lambda x: x**2
#
# A function that takes two numbers and returns their sum:
#
# lambda x, y: x + y
#
# `lambda` generates a function and returns it, while `def` generates a function and assigns it to a name. The function returned by `lambda` also automatically returns the value of its expression statement, which reduces the amount of code that needs to be written.
#
# Here are some additional references that explain lambdas: [Lambda Functions](http://www.secnetix.de/olli/Python/lambda_functions.hawk), [Lambda Tutorial](https://pythonconquerstheuniverse.wordpress.com/2011/08/29/lambda_tutorial/), and [Python Functions](http://www.bogotobogo.com/python/python_functions_lambda.php).
#
# Here is an example:
# + id="mGlQ8L5Rm3nx" colab_type="code" colab={}
# Function declaration using def
def add_s(x):
return x + 's'
print(type(add_s))
print(add_s)
print(add_s('dog'))
# + id="pn-mPOALm3nz" colab_type="code" colab={}
# Same function declared as a lambda
add_s_lambda = lambda x: x + 's'
print(type(add_s_lambda))
print(add_s_lambda) # Note that the function shows its name as <lambda>
print(add_s_lambda('dog'))
# + id="f9b2plXjm3n2" colab_type="code" colab={}
# Code a function using a lambda expression which takes
# a number and returns this number multiplied by two.
multiply_by_two = lambda ???
print(multiply_by_two(5))
Test.assertEqual(multiply_by_two(10), 20, 'incorrect definition for multiply_by_two')
# + [markdown] id="Rn4LtPiFm3n5" colab_type="text"
# Observe the behavior of the following code:
# + id="qlPMtOOim3n6" colab_type="code" colab={}
def add(x, y):
"""Add two values"""
return x + y
def sub(x, y):
"""Substract y from x"""
return x - y
functions = [add, sub]
print(functions[0](1, 2))
print(functions[1](3, 4))
# + [markdown] id="Tm7bmVpTm3n8" colab_type="text"
# Code the same functionality, using lambda expressions:
# + id="o3qHr8AMm3n9" colab_type="code" colab={}
lambda_functions = [lambda ??? , lambda ???]
test.assertEqual(lambda_functions[0](1, 2), 3, 'add lambda_function')
test.assertEqual(lambda_functions[1](3, 4), -1, 'sub lambda_function')
# + id="IboEvuLWm3n_" colab_type="code" outputId="4f0547af-86ef-4f1a-9bbc-82629a7ebdde" colab={}
# Example:
add_two_1 = lambda x, y: (x[0] + y[0], x[1] + y[1])
add_two_2 = lambda x0, x1, y0, y1: (x0 + y0, x1 + y1)
print('add_two_1((1,2), (3,4)) = {0}'.format(add_two_1((1,2), (3,4))))
print('add_two_2((1,2), (3,4)) = {0}'.format(add_two_2(1, 2, 3, 4)))
# + id="N6S8vLC_m3oD" colab_type="code" colab={}
reverse2 = lambda x0, x1, x2: x0+x1+x2
# + id="f483hHCHm3oF" colab_type="code" colab={}
reverse2(1, 2, 3)
# + id="E1qm9IVam3oI" colab_type="code" colab={}
# Use both syntaxes to create a function that takes in a tuple of three values and reverses their order
# E.g. (1, 2, 3) => (3, 2, 1)
reverse1 = lambda x: ???
reverse2 = lambda (x0, x1, x2): ???
Test.assertEqual(reverse1((1, 2, 3)), (3, 2, 1), 'reverse order, syntax 1')
Test.assertEqual(reverse2((1, 2, 3)), (3, 2, 1), 'reverse order, syntax 2')
# + [markdown] id="GOSiEyKLm3oL" colab_type="text"
# Lambda expressions allow you to reduce the size of your code, but they are limited to simple logic. The following Python keywords refer to statements that cannot be used in a lambda expression: `assert`, `pass`, `del`, `print`, `return`, `yield`, `raise`, `break`, `continue`, `import`, `global`, and `exec`. Assignment statements (`=`) and augmented assignment statements (e.g. `+=`) cannot be used either. If more complex logic is necessary, use `def` in place of `lambda`.
# + [markdown] id="p-w3Jsfmm3oN" colab_type="text"
# ## 5. Classes
# ### Points: 4,5 pts
#
# Classes allow you to create objects. Object Oriented Programming (OOP) can be a very powerful paradigm. If done well, OOP allows you to improve the modularity and reusability of your code, but that's the subject of an entire other course.
# Here is a *very* short introduction to it.
#
# By convention, class names are written in camel case, e.g. `MyBeautifulClass`, while variable and function names are written in snake case, e.g. `my_variable`, `my_very_complex_function`
#
# Classes contain methods (i.e. functions owned by the class) and attributes (i.e. variables owned by the class).
# When you define a class, first thing to do is to define a specific method, the constructor. In Python, the constructor is called `__init__`. This method is used to create the instances of an object. Example:
#
# class MyClass:
#
# def __init__(self, first_attribute, second_attribute):
# self.first_attribute = first_attribute
# self.second_attribute = second_attribute
#
# This class has two attributes, and one (hidden) method, the constructor. To create an instance of this class, one simply does:
#
# instance_example = MyClass(1, "foo")
#
# Then, the attributes can easily be accessed to:
#
# instance_example.first_attribute # => 1
# instance_example.first_attribute # => "foo"
# + id="MDzUS4JVm3oN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="34de883b-8b99-4ee4-ce7c-3546fc06da70"
# Run this example
class MyClass:
def __init__(self, first_attribute, second_attribute):
self.first_attribute = first_attribute
self.second_attribute = second_attribute
instance_example = MyClass(1, "foo")
print(instance_example.first_attribute)
instance_example.__init__(3,4) # In real life, it is rare to reinit an object.
print(instance_example.first_attribute)
# + [markdown] id="CPca2qg_m3oR" colab_type="text"
# `self` denotes the object itself. When you declare a method, you have to pass `self` as the first argument of the method:
#
# class MyClass:
#
# def __init__(self, first_attribute, second_attribute):
# self.first_attribute = first_attribute
# self.second_attribute = second_attribute
#
# def method_baz(self):
# print "Hello! I'm a method! I have two attributes, initialized with values %s, %s"%(self.first_attribute, self.second_attribute)
#
# indeed, when we call
#
# instance_example = MyClass(1, "foo")
# instance_example.method_baz()
#
# the `self` object is implicitely passed to `method_baz`as an argument. Think of the method call as the following function call
#
# method_baz(instance_example)
# + id="R_joD4wam3oS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="8cfd07ed-435e-4820-9086-79503fecbd14"
# Run this example
class MyClass:
def __init__(self, first_attribute, second_attribute):
self.first_attribute = first_attribute
self.second_attribute = second_attribute
def class_method(self):
print("Hello! I'm a method! My class has two attributes, of value {0}, {1}".format(self.first_attribute, self.second_attribute))
instance_example = MyClass(1, "foo")
# Call to a class method
instance_example.class_method()
# + [markdown] id="fkD0tMo8m3oU" colab_type="text"
# Now, the tricky part. You can declare **static** methods, i.e. methods that don't need to access the data contained in `self` to work properly. Such methods do not require the `self` argument as they do not use any instance data. They are implemented in the following way:
# + id="VeBTTwS6m3oV" colab_type="code" outputId="c015ac68-758d-4742-b1d4-2f1a9032c8cd" colab={"base_uri": "https://localhost:8080/", "height": 51}
# Run this example
class MyClass:
def __init__(self, first_attribute, second_attribute):
self.first_attribute = first_attribute
self.second_attribute = second_attribute
def class_method(self):
print("Hello! I'm a method! My class has two attributes, of value {0}, {1}".format(self.first_attribute, self.second_attribute))
@staticmethod
def static_method():
print("I'm a static method!")
instance_example = MyClass(1, "foo")
# Call to a class method
instance_example.class_method()
# Call to a static method
instance_example.static_method()
# + id="vPHU87Ckm3oZ" colab_type="code" outputId="f0cb6f03-6cf6-441b-9270-9596c79845ed" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Call to a static method without class instanciation
MyClass.static_method()
# + id="WQRrfmpIm3oc" colab_type="code" outputId="fd004142-9f90-4bd2-99b0-984f7890528d" colab={"base_uri": "https://localhost:8080/", "height": 333}
# Call to a class method without class instanciation: raises an error
MyClass.__setattr__(MyClass(1,2),"second_attribute",2)
MyClass.__setattr__(MyClass(1,2),"first_attribute",25)
MyClass.class_method("popo")
# => TypeError: unbound method class_method() must be called with MyClass instance as first argument (got nothing instead)
# + [markdown] id="SiTEd1GHm3og" colab_type="text"
# You can set attributes without passing them to the constructor:
# + id="i1FoqPcum3og" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="3acded83-ddba-46cb-c5c2-5e3c8f314dad"
# Run this example
class MyClass:
default_attribute = 42
def __init__(self, first_attribute, second_attribute):
self.first_attribute = first_attribute
self.second_attribute = second_attribute
def method_baz(self):
print("Hello! I'm a method! I have two attributes, initialized with values %s, %s"%(self.first_attribute, self.second_attribute))
@staticmethod
def static_method():
print("I'm a static method!")
instance_example = MyClass(1, "foo")
print(instance_example.default_attribute)
# + id="BWq8tp3Tm3oi" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 357} outputId="88a8a743-fda5-429f-956f-b5b349464316"
# Write a Python class named Rectangle which is
# constructed by a length and width
# and has two class methods
# - "rectange_area", which computes the area of a rectangle.
# - "rectangle_perimeter", which computes the perimeter of a rectangle.
#
# The Rectangle class should have an attribute n_edges equal to 4
# which should not be initialized by the __init__ constructor.
#
# Declare a static method "talk" that returns "Do you like rectangles?" when called
class Rectangle:
n_edges = 4
default_attribute = 42
def __init__(self, Length, Width):
self.Length = Length
self.Width = Width
def rectangle_area(self):
return self.Length*self.Width
def rectangle_perimeter(self):
return 2*(self.Length+self.Width)
@staticmethod
def talk():
print("Do you like rectangles?")
new_rectangle = Rectangle(12, 10)
test.assertEqual(new_rectangle.rectangle_area(), 120, "rectangle_area method")
test.assertEqual(new_rectangle.rectangle_perimeter(), 44, "rectangle_area method")
test.assertEqual(Rectangle.n_edges, 4, "constant attibute")
test.assertEqual(Rectangle.talk(), "Do you like rectangles?", "Do you like rectangles?")
# + [markdown] id="npzcsdNgm3ol" colab_type="text"
# Congratulations, you've reched the end of this notebook. =)
| Lab1/Lab1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Úkol č. 4 - regrese (do 2. ledna)
#
# * Cílem tohoto úkolu je vyzkoušet si řešit regresní problém na reálných (ale celkem vyčištěných) datech.
#
# > **Nejdůležitější na úkolu je to, abyste udělali vše procesně správně: korektní rozdělení datasetu, ladění hyperparametrů, vyhodnocení výsledků atp.**
#
# ## Dataset
#
# * Zdrojem dat je list *Data* v souboru `Residential-Building-Data-Set.xlsx` na course pages (originál zde: https://archive.ics.uci.edu/ml/datasets/Residential+Building+Data+Set#).
# * Popis datasetu najdete na listu *Descriptions* ve stejném souboru.
#
#
# ## Pokyny k vypracování
#
# 1. Rozdělte data na trénovací a testovací množinu.
# 1. Proveďte základní průzkum dat a příp. vyhoďte nezajímavé příznaky.
# 1. Aplikujte lineární a hřebenovou regresi a výsledky řádně vyhodnoťte:
# * K měření chyby použijte `mean_absolute_error`.
# * Experimentujte s tvorbou nových příznaků (na základě těch dostupných).
# * Experimentujte se standardizací/normalizací dat.
# * Vyberte si hyperparametry modelů k ladění a najděte jejich nejlepší hodnoty.
# 1. Použijte i jiný model než jen lineární a hřebenovou regresi.
#
#
# ## Poznámky k odevzdání
#
# * Řiďte se pokyny ze stránky https://courses.fit.cvut.cz/BI-VZD/homeworks/index.html.
# * Odevzdejte pouze tento Jupyter Notebook, opravujíví by neměl nic jiného potřebovat.
# * Opravující Vám může umožnit úkol dodělat či opravit a získat tak další body. První verze je ale důležitá a bude-li odbytá, budete za to penalizováni.
# # Řešení
# ## Popis sloupců
#
# * V-1 - Project locality defined in terms of zip codes (N/A)
# * V-2 - Total floor area of the building (m2)
# * V-3 - Lot area (m2)
# * V-4 - Total preliminary estimated construction cost based on the prices at the beginning of the project (10000000 IRR)
# * V-5 - Preliminary estimated construction cost based on the prices at the beginning of the project (10000 IRR)
# * V-6 - Equivalent preliminary estimated construction cost based on the prices at the beginning of the project in a selected base year a (10000 IRR)
# * V-7 - Duration of construction (As a number of time resolution)
# * V-8 - Price of the unit at the beginning of the project per m2 (10000 IRR)
# * V-9 - Actual sales prices (10000 IRR)
# * V-10 - Actual construction costs (10000 IRR)
# * V-11 - The number of building permits issued (N/A)
# * V-12 - Building services index (BSI) b for a preselected base year (N/A)
# * V-13 - Wholesale price index (WPI) c of building materials for the base year (N/A)
# * V-14 - Total floor areas of building permits issued by the city/municipality (m2)
# * V-15 - Cumulative liquidity (10000000 IRR)
# * V-16 - Private sector investment in new buildings (10000000 IRR)
# * V-17 - Land price index for the base year (10000000 IRR)
# * V-18 - The number of loans extended by banks in a time resolution (N/A)
# * V-19 - The amount of loans extended by banks in a time resolution (10000000 IRR)
# * V-20 - The interest rate for loan in a time resolution (%)
# * V-21 - The average construction cost of buildings by private sector at the time of completion of construction (10000 IRR/m2)
# * V-22 - The average of construction cost of buildings by private sector at the beginning of the construction (10000 IRR/m2)
# * V-23 - Official exchange rate with respect to dollars (IRR)
# * V-24 - Nonofficial (street market) exchange rate with respect to dollars (IRR)
# * V-25 - Consumer price index (CPI) i in the base year (N/A)
# * V-26 - CPI of housing, water, fuel & power in the base year (N/A)
# * V-27 - Stock market index (N/A)
# * V-28 - Population of the city (N/A)
# * V-29 - Gold price per ounce (IRR)
# ## Nahrání datasetu
import pandas as pd
import numpy as np
url = 'https://archive.ics.uci.edu/ml/machine-learning-databases/00437/Residential-Building-Data-Set.xlsx'
file = 'Residential-Building-Data-Set.xlsx'
data = pd.read_excel(io=url, sheet_name='Data', header=1)
data.head()
# ## Rozdělení dat na trénovací, validační a testovací
# +
from sklearn.model_selection import train_test_split
X = data.copy()
X = X.drop(['V-9', 'V-10'], axis=1)
Y = data.copy()[['V-9', 'V-10']]
rd_seed = 100
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size=0.25, random_state=rd_seed)
# -
# ## Použití lineární regrese bez úprav dat a ladění hyperparametrů
# +
from sklearn.linear_model import LinearRegression
from sklearn.metrics import mean_absolute_error
import matplotlib.pyplot as plt
import matplotlib
# %matplotlib inline
lr = LinearRegression()
lr.fit(X_train, Y_train)
Y_predicted = lr.predict(X_test)
plt.scatter(Y_test, Y_predicted)
plt.plot([0,7000], [0,7000], 'r')
plt.show()
print('MAE:', (mean_absolute_error(Y_predicted, np.array(Y_test))))
# -
# ## Úprava datasetu
# * Zip code je zde v podstatě kategorická proměnná, která nám zde akorát škodí. Nelze usuzovat, že podobný zip code bude mít podobné výstupní hodnoty. Abychom tento problém vyřešili, ale zároveň nějak naložili se zip codem, který sám o sobě je docela dobrá informace, provedeme one hot encoding.
X = data.copy()
zip_code_dummies = pd.get_dummies(X['V-1'])
X = pd.concat([X.drop('V-1', axis=1), zip_code_dummies], axis=1)
# * Sloupce V-11 až V-29 jsou zde několikrát. Důvodem je zaznamenání údajů v různých fázích projektu. Můžeme to tak nechat být a brát je jako různé příznaky, nebo s touto skutečností můžeme experimentovat třeba sloučením těchto sloupců do jednoho, například jako průměr jejich hodnot.
# +
def mean_timelags(row, v):
V = 'V-' + str(v)
s = row[V]
for i in range(1,5):
s += row[V + "." + str(i)]
return s/5
for i in range(11, 30):
X['V-' + str(i)] = X.apply (lambda row: mean_timelags(row, i), axis=1)
for j in range(1, 5):
X = X.drop('V-' + str(i) + "." + str(j), axis=1)
# -
# ## Příznaky
# * Všechny příznaky jsou rozhodně zajímavé a užitečné. Určitě by se mezi některými z nich dala najít korelace, která je u lineární regrese nežádoucí. Nicméně zkusíme ponechat všechny příznaky.
# ## Standardizace, Normalizace
# * Pro linární/hřebenovou regresi si standardizací nepomůžeme a normalizace není tak užitečná/efektivní jako pro jiné modely strojového učení. Přesto zkusíme normalizovat. Použijeme MinMaxScaler.
from sklearn.preprocessing import MinMaxScaler
scaler = MinMaxScaler()
idx = X.index
cols = X.columns
X_minmax = pd.DataFrame(scaler.fit_transform(X), index=idx, columns=cols)
# ## Použití lineární regrese na upravených datech
# * U lineární regrese pomoci metody nejmenších čtverců nebudeme ladit hyperparametry, protože nemáme žádné
# +
X_minmax = X_minmax.drop(['V-9', 'V-10'], axis=1)
Y_minmax = data.copy()[['V-9', 'V-10']]
rd_seed = 170
X_train, X_test, Y_train, Y_test = train_test_split(X_minmax, Y_minmax, test_size=0.25, random_state=rd_seed)
# +
lr = LinearRegression()
lr.fit(X_train, Y_train)
Y_predicted = lr.predict(X_test)
print('MAE:', (mean_absolute_error(Y_predicted, np.array(Y_test))))
plt.scatter(Y_test, Y_predicted)
plt.plot([0,7000], [0,7000], 'r')
plt.show()
# -
# ## Použití hřebenové regrese na upravených datech
# * Použijeme stejné úpravy jako pro lineární regresi, tedy použijeme už připravená data.
# * Snažíme se najít optimální hodnotu parametru alfa - chápeme jako ladění hyperparametru
# +
from sklearn.linear_model import Ridge
from sklearn.metrics import make_scorer
from sklearn.model_selection import cross_val_score
from scipy import optimize
def get_ridge_model(X, Y):
def ridgemodel(alpha):
model = Ridge(alpha=alpha)
return -np.mean(cross_val_score(model, X, Y, cv=5, scoring='neg_mean_absolute_error'))
opt_alpha = optimize.minimize_scalar(ridgemodel, options = {'maxiter': 50}, method = 'bounded', bounds=(0.01, 100))
print('Optimal alpha', opt_alpha.x)
model = Ridge(alpha = opt_alpha.x)
model.fit(X,Y)
return model
r_model = get_ridge_model(X_train, Y_train)
Yth = r_model.predict(X_test)
plt.scatter(Y_test, Yth)
plt.plot([0,7000], [0,7000], 'r')
print('MAE:', mean_absolute_error(Yth, np.array(Y_test)))
# -
# ## Random Forest Regressor
# * Stromové struktury není třeba nestandardizujeme ani nenormalizujeme
# * Máme na výběr spoustu hyperparametrů na trénování, důležité je nepřepálit počet hodnot a parametrů, abychom vůbec dostali výsledky (zde pouze n_estimators, max_depth)
# +
from sklearn.ensemble import RandomForestRegressor
from sklearn.model_selection import GridSearchCV
X_rf = X.drop(['V-9', 'V-10'], axis=1)
Y_rf = data.copy()[['V-9', 'V-10']]
X_train, X_test, Y_train, Y_test = train_test_split(X_rf, Y_rf, test_size=0.25, random_state=rd_seed)
rf = RandomForestRegressor()
parameters = {
'n_estimators': range(1,100,5),
'max_depth': range(1,10),
'random_state': [rd_seed] # for tuning purposes
# 'max_features': ['log2', 'sqrt','auto'],
# 'min_samples_split': [2, 3, 5],
# 'min_samples_leaf': [1, 5, 8]
}
grid_obj = GridSearchCV(rf, parameters, scoring='neg_mean_absolute_error', cv=5, iid=False, error_score='raise')
grid_obj = grid_obj.fit(X_train, Y_train)
rf = grid_obj.best_estimator_
rf.fit(X_train, Y_train)
Y_predicted = rf.predict(X_test)
print('Best params:', grid_obj.best_params_)
print('MAE:', (mean_absolute_error(Y_predicted, np.array(Y_test))))
# -
# ## Výsledky, závěr
# * Je vidět, že is mírnými úpravami datasetu, jako je zprůměrování sloupců a one hot encoding u zip code, jsem snížili MAE o pár desítek. Standardizace u lineární a hřebenové regrese nám pomoci nemůže a normalizace sice ano, ale opravdu jen minimálně. Dále jsem si zkusil pohrát s vyhazováním neužitečných příznaků (mimo), ale to se mi nijak neosvědčilo. Lineární a hřebenová regrese (MAE lehce nad 60) dává mnohem lepší výsledky oproti Random Forest regresi (MAE nad 80). Co se týče ohledně lineární vs. hřebenové regrese, hřebenová dává jen nepatrně lepší výsledky. Bylo ale důležité zvolit/natrénovat správnou alfu.
| 04/homework_04_B191.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from __future__ import division
import pickle
import random
import os
import math
import types
import uuid
import time
from copy import copy
from collections import defaultdict, Counter
import numpy as np
import gym
from gym import spaces, wrappers
import dill
import tempfile
import tensorflow as tf
from tensorflow.contrib import rnn
import zipfile
import baselines.common.tf_util as U
from baselines import logger
from baselines.common.schedules import LinearSchedule
from baselines import deepq
from baselines.deepq.replay_buffer import ReplayBuffer, PrioritizedReplayBuffer
from baselines.deepq.simple import ActWrapper
from scipy.special import logsumexp
from pyglet.window import key as pygkey
# -
from matplotlib import pyplot as plt
# %matplotlib inline
import matplotlib as mpl
mpl.rc('savefig', dpi=300)
mpl.rc('text', usetex=True)
data_dir = os.path.join('data', 'lunarlander-sim')
# train synthetic pilot
throttle_mag = 0.75
def disc_to_cont(action):
if type(action) == np.ndarray:
return action
# main engine
if action < 3:
m = -throttle_mag
elif action < 6:
m = throttle_mag
else:
raise ValueError
# steering
if action % 3 == 0:
s = -throttle_mag
elif action % 3 == 1:
s = 0
else:
s = throttle_mag
return np.array([m, s])
# +
def mask_helipad(obs, replace=0):
obs = copy(obs)
if len(obs.shape) == 1:
obs[8] = replace
else:
obs[:, 8] = replace
return obs
def traj_mask_helipad(traj):
return [mask_helipad(obs) for obs in traj]
# -
n_act_dim = 6
n_obs_dim = 9
# +
def onehot_encode(i, n=n_act_dim):
x = np.zeros(n)
x[i] = 1
return x
def onehot_decode(x):
l = np.nonzero(x)[0]
assert len(l) == 1
return l[0]
# -
def make_env(using_lander_reward_shaping=False):
env = gym.make('LunarLanderContinuous-v2')
env.action_space = spaces.Discrete(n_act_dim)
env.unwrapped._step_orig = env.unwrapped._step
def _step(self, action):
obs, r, done, info = self._step_orig(disc_to_cont(action))
return obs, r, done, info
env.unwrapped._step = types.MethodType(_step, env.unwrapped)
env.unwrapped.using_lander_reward_shaping = using_lander_reward_shaping
return env
env = make_env(using_lander_reward_shaping=True)
max_ep_len = 1000
n_training_episodes = 500
make_q_func = lambda: deepq.models.mlp([64, 64])
pilot_dqn_learn_kwargs = {
'lr': 1e-3,
'exploration_fraction': 0.1,
'exploration_final_eps': 0.02,
'target_network_update_freq': 1500,
'print_freq': 100,
'num_cpu': 5,
'gamma': 0.99
}
full_pilot_scope = 'full_pilot'
full_pilot_q_func = make_q_func()
load_pretrained_full_pilot = True
max_timesteps = max_ep_len * (1 if load_pretrained_full_pilot else n_training_episodes)
raw_full_pilot_policy, full_pilot_reward_data = deepq.learn(
env,
q_func=full_pilot_q_func,
max_timesteps=max_timesteps,
scope=full_pilot_scope,
**pilot_dqn_learn_kwargs
)
with open(os.path.join(data_dir, 'full_pilot_reward_data.pkl'), 'wb') as f:
pickle.dump(full_pilot_reward_data, f, pickle.HIGHEST_PROTOCOL)
with open(os.path.join(data_dir, 'full_pilot_reward_data.pkl'), 'rb') as f:
full_pilot_reward_data = pickle.load(f)
def run_ep(policy, env, max_ep_len=max_ep_len, render=False, pilot_is_human=False):
if pilot_is_human:
global human_agent_action
human_agent_action = init_human_action()
obs = env.reset()
done = False
totalr = 0.
trajectory = None
actions = []
for step_idx in range(max_ep_len+1):
if done:
trajectory = info['trajectory']
break
action = policy(obs[None, :])
obs, r, done, info = env.step(action)
actions.append(action)
if render:
env.render()
totalr += r
outcome = r if r % 100 == 0 else 0
return totalr, outcome, trajectory, actions
def full_pilot_policy(obs):
with tf.variable_scope(full_pilot_scope, reuse=None):
return raw_full_pilot_policy._act(obs)[0]
class LaggyPilotPolicy(object):
def __init__(self):
self.last_laggy_pilot_act = None
def __call__(self, obs, lag_prob=0.8):
if self.last_laggy_pilot_act is None or np.random.random() >= lag_prob:
action = full_pilot_policy(obs)
self.last_laggy_pilot_act = action
return self.last_laggy_pilot_act
laggy_pilot_policy = LaggyPilotPolicy()
def noisy_pilot_policy(obs, noise_prob=0.15):
action = full_pilot_policy(obs)
if np.random.random() < noise_prob:
action = (action + 3) % 6
if np.random.random() < noise_prob:
action = action//3*3 + (action + np.random.randint(1, 3)) % 3
return action
def noop_pilot_policy(obs):
return 1
def sensor_pilot_policy(obs, thresh=0.1):
d = obs[0, 8] - obs[0, 0] # horizontal dist to helipad
if d < -thresh:
return 0
elif d > thresh:
return 2
else:
return 1
# +
# begin debug
# -
run_ep(full_pilot_policy, env, render=True)
env.close()
# +
# end debug
# -
def save_tf_vars(scope, path):
sess = U.get_session()
saver = tf.train.Saver([v for v in tf.global_variables() if v.name.startswith(scope + '/')])
saver.save(sess, save_path=path)
def load_tf_vars(scope, path):
sess = U.get_session()
saver = tf.train.Saver([v for v in tf.global_variables() if v.name.startswith(scope + '/')])
saver.restore(sess, path)
full_pilot_path = os.path.join(data_dir, 'full_pilot.tf')
save_tf_vars(full_pilot_scope, full_pilot_path)
load_tf_vars(full_pilot_scope, full_pilot_path)
# evaluate synthetic pilot
pilot_names = ['full', 'laggy', 'noisy', 'noop', 'sensor']
n_eval_eps = 100
pilot_evals = [list(zip(*[run_ep(eval('%s_pilot_policy' % pilot_name), env, render=False) for _ in range(n_eval_eps)])) for pilot_name in pilot_names]
with open(os.path.join(data_dir, 'pilot_evals.pkl'), 'wb') as f:
pickle.dump(dict(zip(pilot_names, pilot_evals)), f, pickle.HIGHEST_PROTOCOL)
mean_rewards = [np.mean(pilot_eval[0]) for pilot_eval in pilot_evals]
outcome_distrns = [Counter(pilot_eval[1]) for pilot_eval in pilot_evals]
print('\n'.join([str(x) for x in zip(pilot_names, mean_rewards, outcome_distrns)]))
n_videos = 10
for pilot_name in pilot_names:
for i in range(n_videos):
wrapped_env = wrappers.Monitor(env, os.path.join(data_dir, 'videos', '%s_pilot.%d' % (pilot_name, i)), force=True)
run_ep(eval('%s_pilot_policy' % pilot_name), wrapped_env, render=False)
wrapped_env.close()
env.close()
# train supervised goal decoder
pilot_name = 'full'
pilot_policy = eval('%s_pilot_policy' % pilot_name)
n_rollouts = 1000
rollouts = [run_ep(pilot_policy, env, render=False)[2:] for _ in range(n_rollouts)]
with open(os.path.join(data_dir, '%s_pilot_policy_rollouts.pkl' % pilot_name), 'wb') as f:
pickle.dump(rollouts, f, pickle.HIGHEST_PROTOCOL)
with open(os.path.join(data_dir, '%s_pilot_policy_rollouts.pkl' % pilot_name), 'rb') as f:
rollouts = pickle.load(f)
n_val_rollouts = 100
rollouts, val_rollouts = rollouts[:-n_val_rollouts], rollouts[-n_val_rollouts:]
def combined_rollout(states, actions):
return np.array([np.concatenate((
np.array(obs),
onehot_encode(action))) for obs, action in zip(
states[:-1] if len(states) == len(actions) + 1 else states, actions)])
def format_rollouts(rollouts):
X_dat = np.zeros((len(rollouts), max_ep_len, n_obs_dim + n_act_dim))
Y_dat = np.zeros((len(rollouts), max_ep_len))
M_dat = np.zeros((len(rollouts), max_ep_len))
for i, (states, actions) in enumerate(rollouts):
Y_dat[i, :] = states[0][-1]
X_dat[i, :len(actions), :] = traj_mask_helipad(combined_rollout(states, actions))
M_dat[i, :len(actions)] = 1
return X_dat, Y_dat, M_dat
X_dat, Y_dat, M_dat = format_rollouts(rollouts)
val_X_dat, val_Y_dat, val_M_dat = format_rollouts(val_rollouts)
example_idxes = list(range(X_dat.shape[0]))
def next_batch(batch_size):
batch_idxes = random.sample(example_idxes, batch_size)
return X_dat[batch_idxes, :, :], Y_dat[batch_idxes, :], M_dat[batch_idxes, :]
# +
# Training Parameters
learning_rate = 1e-2
training_steps = 1000
batch_size = 128
display_step = training_steps // 10
# Network Parameters
num_input = X_dat.shape[2]
timesteps = X_dat.shape[1] # timesteps
num_hidden = 32 # hidden layer num of features
# -
gd_scope = 'gd_scope'
with tf.variable_scope(gd_scope, reuse=False):
# tf Graph input
X = tf.placeholder("float", [None, timesteps, num_input])
Y = tf.placeholder("float", [None, timesteps])
M = tf.placeholder("float", [None, timesteps]) # mask for variable length sequences
INIT_STATE_A = tf.placeholder("float", [None, num_hidden])
INIT_STATE_B = tf.placeholder("float", [None, num_hidden])
weights = {
'out': tf.Variable(tf.random_normal([num_hidden, 1]))
}
biases = {
'out': tf.Variable(tf.random_normal([1]))
}
unstacked_X = tf.unstack(X, timesteps, 1)
lstm_cell = rnn.BasicLSTMCell(num_hidden, forget_bias=1.0)
state = (INIT_STATE_A, INIT_STATE_B)
rnn_outputs = []
rnn_states = []
for input_ in unstacked_X:
output, state = lstm_cell(input_, state)
rnn_outputs.append(output)
rnn_states.append(state)
prediction = tf.reshape(
tf.concat([tf.matmul(output, weights['out']) + biases['out'] for output in rnn_outputs], axis=1),
shape=[tf.shape(X)[0], timesteps])
predictions = [tf.matmul(output, weights['out']) + biases['out'] for output in rnn_outputs]
loss_op = tf.reduce_sum((prediction - Y)**2 * M) / tf.reduce_sum(M)
optimizer = tf.train.AdamOptimizer(learning_rate=learning_rate)
train_op = optimizer.minimize(loss_op)
# +
sess = U.get_session()
if sess is None:
sess = U.make_session(num_cpu=5)
sess.__enter__()
sess.run(tf.variables_initializer([v for v in tf.global_variables() if v.name.startswith(gd_scope + '/')]))
# -
with tf.variable_scope(gd_scope, reuse=False):
for step in range(1, training_steps+1):
batch_x, batch_y, batch_mask = next_batch(batch_size)
sess.run(train_op, feed_dict={X: batch_x, Y: batch_y, M: batch_mask,
INIT_STATE_A: np.zeros((batch_size, num_hidden)),
INIT_STATE_B: np.zeros((batch_size, num_hidden))})
if step % display_step == 0 or step == 1:
loss = sess.run(loss_op, feed_dict={X: X_dat,
Y: Y_dat,
M: M_dat,
INIT_STATE_A: np.zeros((X_dat.shape[0], num_hidden)),
INIT_STATE_B: np.zeros((X_dat.shape[0], num_hidden))})
val_loss = sess.run(loss_op, feed_dict={X: val_X_dat,
Y: val_Y_dat,
M: val_M_dat,
INIT_STATE_A: np.zeros((val_X_dat.shape[0], num_hidden)),
INIT_STATE_B: np.zeros((val_X_dat.shape[0], num_hidden))})
print("Step " + str(step) + ", Training Loss= " + \
"{:.4f}".format(loss), ", Validation Loss= " + "{:.4f}".format(val_loss))
print("Optimization Finished!")
def train_supervised_goal_decoder(gd_scope, rollouts):
X_dat, Y_dat, M_dat = format_rollouts(rollouts)
example_idxes = list(range(X_dat.shape[0]))
def next_batch(batch_size):
batch_idxes = random.sample(example_idxes, batch_size)
return X_dat[batch_idxes, :, :], Y_dat[batch_idxes, :], M_dat[batch_idxes, :]
# Training Parameters
learning_rate = 1e-2
training_steps = 1000
batch_size = 128
display_step = training_steps // 10
# Network Parameters
num_input = X_dat.shape[2]
timesteps = X_dat.shape[1] # timesteps
num_hidden = 32 # hidden layer num of features
sess = U.get_session()
if sess is None:
sess = U.make_session(num_cpu=5)
sess.__enter__()
sess.run(tf.variables_initializer([v for v in tf.global_variables() if v.name.startswith(gd_scope + '/')]))
with tf.variable_scope(gd_scope, reuse=False):
for step in range(1, training_steps+1):
batch_x, batch_y, batch_mask = next_batch(batch_size)
sess.run(train_op, feed_dict={X: batch_x, Y: batch_y, M: batch_mask,
INIT_STATE_A: np.zeros((batch_size, num_hidden)),
INIT_STATE_B: np.zeros((batch_size, num_hidden))})
if step % display_step == 0 or step == 1:
loss = sess.run(loss_op, feed_dict={X: X_dat,
Y: Y_dat,
M: M_dat,
INIT_STATE_A: np.zeros((X_dat.shape[0], num_hidden)),
INIT_STATE_B: np.zeros((X_dat.shape[0], num_hidden))})
print("Step " + str(step) + ", Training Loss={:.4f}".format(loss))
print("Optimization Finished!")
def build_retrain_goal_decoder(pilot_name):
with open(os.path.join(data_dir, '%s_pilot_policy_rollouts.pkl' % pilot_name), 'rb') as f:
off_pol_rollouts = pickle.load(f)
def retrain_goal_decoder(on_pol_rollouts):
train_supervised_goal_decoder(gd_scope, off_pol_rollouts + on_pol_rollouts)
return retrain_goal_decoder
gd_path = os.path.join(data_dir, '%s_pilot_goal_decoder.tf' % pilot_name)
save_tf_vars(gd_scope, gd_path)
load_tf_vars(gd_scope, gd_path)
def decode_goal(trajectory, init_state=None, only_final=False):
traj_X = np.zeros((1, max_ep_len, n_obs_dim + n_act_dim))
traj_X[0, :len(trajectory), :] = np.array(trajectory)
with tf.variable_scope(gd_scope, reuse=False):
feed_dict = {X: traj_X}
if init_state is not None:
feed_dict[INIT_STATE_A] = init_state[0]
feed_dict[INIT_STATE_B] = init_state[1]
else:
feed_dict[INIT_STATE_A] = np.zeros((1, num_hidden))
feed_dict[INIT_STATE_B] = np.zeros((1, num_hidden))
if only_final:
g, s = sess.run(
[predictions[len(trajectory)-1], rnn_states[len(trajectory)-1]],
feed_dict=feed_dict
)
return g[0, 0], s
else:
g, s = sess.run(
[predictions, rnn_states[len(trajectory)-1]],
feed_dict=feed_dict
)
return [x[0, 0] for x in g], s
def build_build_goal_decoder(pilot_name):
def build_goal_decoder():
load_tf_vars(gd_scope, os.path.join(data_dir, '%s_pilot_goal_decoder.tf' % pilot_name))
return decode_goal
return build_goal_decoder
# build model-based goal decoder
goals = np.arange(-0.8, 1, 0.05)
n_goals = len(goals)
# +
sess = U.get_session()
if sess is None:
sess = U.make_session(num_cpu=5)
sess.__enter__()
with tf.variable_scope(full_pilot_scope, reuse=None):
Q_obs = tf.get_variable("Q_obs", (n_goals, n_obs_dim))
sess.run(tf.variables_initializer([Q_obs]))
# -
with tf.variable_scope(full_pilot_scope, reuse=True):
Q_values = full_pilot_q_func(Q_obs, n_act_dim, scope="q_func")
def compute_map_est_goal(s, a, log_prior, temp=1000):
states = []
for g in goals:
state = copy(s)
state[8] = g
states.append(state)
with tf.variable_scope(full_pilot_scope, reuse=True):
Q = sess.run(
Q_values,
feed_dict={Q_obs: np.array(states)}
)
Q *= temp
action = onehot_decode(a)
log_cond_likelihood = Q[:, action] - logsumexp(Q, axis=1)
log_marginal_likelihood = logsumexp(log_cond_likelihood) - np.log(n_goals)
log_likelihood = log_cond_likelihood - log_marginal_likelihood
log_posterior = log_likelihood + log_prior
map_est_goal = goals[max(range(n_goals), key=lambda i: log_posterior[i])]
return log_posterior, map_est_goal
zero_goal_idx = len(goals)//2-2
def mb_decode_goal(trajectory, init_state=None, only_final=False):
if init_state is None:
prior = np.ones(n_goals) / n_goals
prior[zero_goal_idx] *= 2
prior = prior / prior.sum()
log_prior = np.log(prior)
map_est_goals = []
else:
log_prior, map_est_goals = init_state
trajectory = trajectory[-1:]
for t in trajectory:
s = np.array(t[:-n_act_dim])
a = np.array(t[-n_act_dim:])
log_posterior, map_est_goal = compute_map_est_goal(s, a, log_prior)
map_est_goals.append(map_est_goal)
log_prior = log_posterior
return (map_est_goals[-1] if only_final else map_est_goals), (log_posterior, map_est_goals)
decode_goal = mb_decode_goal
def build_build_goal_decoder(pilot_name):
def build_goal_decoder():
return decode_goal
return build_goal_decoder
# +
# begin debug
# -
rollout = rollouts[925]
goal = rollout[0][0][-1]
traj = traj_mask_helipad(combined_rollout(*rollout))
pred_goal, _ = decode_goal(traj)
mb_pred_goal, _ = mb_decode_goal(traj)
plt.xlabel('Step')
plt.ylabel('X-Coordinate')
plt.axhline(y=goal, label='True Goal', linestyle='--', linewidth=5, color='gray', alpha=0.5)
plt.plot(pred_goal[:len(rollout[0])], label='Predicted Goal (SL)', color='orange')
plt.plot(mb_pred_goal[:len(rollout[0])], label='Predicted Goal (BI)', color='teal')
plt.legend(loc='best')
plt.ylim([-1, 1])
plt.show()
#rollout = rollouts[986]
for rollout in rollouts[800:850]:
goal = rollout[0][0][-1]
traj = traj_mask_helipad(combined_rollout(*rollout))
pred_goal, _ = decode_goal(traj)
mb_pred_goal, _ = mb_decode_goal(traj)
plt.xlabel('Step')
plt.ylabel('X-Coordinate')
plt.axhline(y=goal, label='True Goal', linestyle='--', linewidth=5, color='gray', alpha=0.5)
plt.plot(pred_goal[:len(rollout[0])], label='Predicted Goal (SL)', color='orange')
plt.plot(mb_pred_goal[:len(rollout[0])], label='Predicted Goal (BI)', color='teal')
plt.legend(loc='best')
plt.ylim([-1, 1])
plt.show()
y_trues = []
y_preds = []
for rollout in rollouts:
goal = rollout[0][0][-1]
traj = traj_mask_helipad(combined_rollout(*rollout))
pred_goal, final_states = mb_decode_goal(traj)
y_trues.extend([goal] * len(pred_goal))
y_preds.extend(pred_goal)
y_trues = np.array(y_trues)
y_preds = np.array(y_preds)
np.mean((y_trues - y_preds)**2), np.mean((y_trues - 0)**2)
mb_pred_goal, final_states = mb_decode_goal(traj)
plt.ylabel('Timestep')
plt.xlabel('Horizontal Location')
plt.title('Sample Episode from Optimal Synthetic Pilot')
plt.axvline(x=goal, label='True Goal', linestyle='--', linewidth=1, color='green')
plt.plot(list(reversed(pred_goal[:len(rollout[0])])), range(len(pred_goal[:len(rollout[0])])), label='Inferred Goal (Supervised Learning)', color='teal')
plt.plot(list(reversed(mb_pred_goal[:len(rollout[0])])), range(len(mb_pred_goal[:len(rollout[0])])), label='Inferred Goal (Bayesian Inference)', color='gray')
plt.yticks([0, 100, 200, 300, 400], ['400', '300', '200', '100', '0'])
plt.legend(loc='best')
plt.xlim([-1, 1])
plt.show()
# +
# end debug
# -
# train assistive copilot
n_training_episodes = 500
make_q_func = lambda: deepq.models.mlp([64, 64])
copilot_dqn_learn_kwargs = {
'lr': 1e-3,
'exploration_fraction': 0.1,
'exploration_final_eps': 0.02,
'target_network_update_freq': 1500,
'print_freq': 100,
'num_cpu': 5,
'gamma': 0.99,
}
def make_co_env(pilot_policy, build_goal_decoder=None, using_lander_reward_shaping=False, **extras):
env = gym.make('LunarLanderContinuous-v2')
env.unwrapped.using_lander_reward_shaping = using_lander_reward_shaping
env.action_space = spaces.Discrete(n_act_dim)
env.unwrapped.pilot_policy = pilot_policy
if build_goal_decoder is None:
obs_box = env.observation_space
env.observation_space = spaces.Box(np.concatenate((obs_box.low, np.zeros(n_act_dim))),
np.concatenate((obs_box.high, np.ones(n_act_dim))))
env.unwrapped._step_orig = env.unwrapped._step
if build_goal_decoder is None:
def _step(self, action):
obs, r, done, info = self._step_orig(disc_to_cont(action))
obs = np.concatenate((obs, onehot_encode(self.pilot_policy(obs[None, :]))))
return obs, r, done, info
else:
goal_decoder = build_goal_decoder()
def _step(self, action):
obs, r, done, info = self._step_orig(disc_to_cont(action))
self.actions.append(self.pilot_policy(obs[None, :]))
traj = traj_mask_helipad(combined_rollout(self.trajectory[-1:], self.actions[-1:]))
goal, self.init_state = goal_decoder(traj, init_state=self.init_state, only_final=True)
obs = mask_helipad(obs, replace=goal)
return obs, r, done, info
env.unwrapped._step = types.MethodType(_step, env.unwrapped)
return env
def co_build_act(make_obs_ph, q_func, num_actions, scope="deepq", reuse=None, using_control_sharing=True):
with tf.variable_scope(scope, reuse=reuse):
observations_ph = U.ensure_tf_input(make_obs_ph("observation"))
if using_control_sharing:
pilot_action_ph = tf.placeholder(tf.int32, (), name='pilot_action')
pilot_tol_ph = tf.placeholder(tf.float32, (), name='pilot_tol')
else:
eps = tf.get_variable("eps", (), initializer=tf.constant_initializer(0))
stochastic_ph = tf.placeholder(tf.bool, (), name="stochastic")
update_eps_ph = tf.placeholder(tf.float32, (), name="update_eps")
q_values = q_func(observations_ph.get(), num_actions, scope="q_func")
batch_size = tf.shape(q_values)[0]
if using_control_sharing:
q_values -= tf.reduce_min(q_values, axis=1)
opt_actions = tf.argmax(q_values, axis=1, output_type=tf.int32)
opt_q_values = tf.reduce_max(q_values, axis=1)
batch_idxes = tf.reshape(tf.range(0, batch_size, 1), [batch_size, 1])
reshaped_batch_size = tf.reshape(batch_size, [1])
pi_actions = tf.tile(tf.reshape(pilot_action_ph, [1]), reshaped_batch_size)
pi_act_idxes = tf.concat([batch_idxes, tf.reshape(pi_actions, [batch_size, 1])], axis=1)
pi_act_q_values = tf.gather_nd(q_values, pi_act_idxes)
# if necessary, switch steering and keep main
mixed_actions = 3 * (pi_actions // 3) + (opt_actions % 3)
mixed_act_idxes = tf.concat([batch_idxes, tf.reshape(mixed_actions, [batch_size, 1])], axis=1)
mixed_act_q_values = tf.gather_nd(q_values, mixed_act_idxes)
mixed_actions = tf.where(pi_act_q_values >= (1 - pilot_tol_ph) * opt_q_values, pi_actions, mixed_actions)
# if necessary, keep steering and switch main
mixed_act_idxes = tf.concat([batch_idxes, tf.reshape(mixed_actions, [batch_size, 1])], axis=1)
mixed_act_q_values = tf.gather_nd(q_values, mixed_act_idxes)
steer_mixed_actions = 3 * (opt_actions // 3) + (pi_actions % 3)
mixed_actions = tf.where(mixed_act_q_values >= (1 - pilot_tol_ph) * opt_q_values, mixed_actions, steer_mixed_actions)
# if necessary, switch steering and main
mixed_act_idxes = tf.concat([batch_idxes, tf.reshape(mixed_actions, [batch_size, 1])], axis=1)
mixed_act_q_values = tf.gather_nd(q_values, mixed_act_idxes)
actions = tf.where(mixed_act_q_values >= (1 - pilot_tol_ph) * opt_q_values, mixed_actions, opt_actions)
act = U.function(inputs=[
observations_ph, pilot_action_ph, pilot_tol_ph
],
outputs=[actions])
else:
deterministic_actions = tf.argmax(q_values, axis=1)
random_actions = tf.random_uniform(tf.stack([batch_size]), minval=0, maxval=num_actions, dtype=tf.int64)
chose_random = tf.random_uniform(tf.stack([batch_size]), minval=0, maxval=1, dtype=tf.float32) < eps
stochastic_actions = tf.where(chose_random, random_actions, deterministic_actions)
output_actions = tf.cond(stochastic_ph, lambda: stochastic_actions, lambda: deterministic_actions)
update_eps_expr = eps.assign(tf.cond(update_eps_ph >= 0, lambda: update_eps_ph, lambda: eps))
act = U.function(inputs=[observations_ph, stochastic_ph, update_eps_ph],
outputs=[output_actions],
givens={update_eps_ph: -1.0, stochastic_ph: True},
updates=[update_eps_expr])
return act
def co_build_train(make_obs_ph, q_func, num_actions, optimizer, grad_norm_clipping=None, gamma=1.0,
double_q=True, scope="deepq", reuse=None, using_control_sharing=True):
act_f = co_build_act(make_obs_ph, q_func, num_actions, scope=scope, reuse=reuse, using_control_sharing=using_control_sharing)
with tf.variable_scope(scope, reuse=reuse):
# set up placeholders
obs_t_input = U.ensure_tf_input(make_obs_ph("obs_t"))
act_t_ph = tf.placeholder(tf.int32, [None], name="action")
rew_t_ph = tf.placeholder(tf.float32, [None], name="reward")
obs_tp1_input = U.ensure_tf_input(make_obs_ph("obs_tp1"))
done_mask_ph = tf.placeholder(tf.float32, [None], name="done")
importance_weights_ph = tf.placeholder(tf.float32, [None], name="weight")
obs_t_input_get = obs_t_input.get()
obs_tp1_input_get = obs_tp1_input.get()
# q network evaluation
q_t = q_func(obs_t_input_get, num_actions, scope='q_func', reuse=True) # reuse parameters from act
q_func_vars = U.scope_vars(U.absolute_scope_name('q_func'))
# target q network evalution
q_tp1 = q_func(obs_tp1_input_get, num_actions, scope="target_q_func")
target_q_func_vars = U.scope_vars(U.absolute_scope_name("target_q_func"))
# q scores for actions which we know were selected in the given state.
q_t_selected = tf.reduce_sum(q_t * tf.one_hot(act_t_ph, num_actions), 1)
# compute estimate of best possible value starting from state at t + 1
if double_q:
q_tp1_using_online_net = q_func(obs_tp1_input_get, num_actions, scope='q_func', reuse=True)
q_tp1_best_using_online_net = tf.arg_max(q_tp1_using_online_net, 1)
q_tp1_best = tf.reduce_sum(q_tp1 * tf.one_hot(q_tp1_best_using_online_net, num_actions), 1)
else:
q_tp1_best = tf.reduce_max(q_tp1, 1)
q_tp1_best_masked = (1.0 - done_mask_ph) * q_tp1_best
# compute RHS of bellman equation
q_t_selected_target = rew_t_ph + gamma * q_tp1_best_masked
# compute the error (potentially clipped)
td_error = q_t_selected - tf.stop_gradient(q_t_selected_target)
errors = U.huber_loss(td_error)
weighted_error = tf.reduce_mean(importance_weights_ph * errors)
# compute optimization op (potentially with gradient clipping)
if grad_norm_clipping is not None:
optimize_expr = U.minimize_and_clip(optimizer,
weighted_error,
var_list=q_func_vars,
clip_val=grad_norm_clipping)
else:
optimize_expr = optimizer.minimize(weighted_error, var_list=q_func_vars)
# update_target_fn will be called periodically to copy Q network to target Q network
update_target_expr = []
for var, var_target in zip(sorted(q_func_vars, key=lambda v: v.name),
sorted(target_q_func_vars, key=lambda v: v.name)):
update_target_expr.append(var_target.assign(var))
update_target_expr = tf.group(*update_target_expr)
# Create callable functions
train = U.function(
inputs=[
obs_t_input,
act_t_ph,
rew_t_ph,
obs_tp1_input,
done_mask_ph,
importance_weights_ph
],
outputs=td_error,
updates=[optimize_expr]
)
update_target = U.function([], [], updates=[update_target_expr])
q_values = U.function([obs_t_input], q_t)
return act_f, train, update_target, {'q_values': q_values}
def co_dqn_learn(
env,
q_func,
lr=1e-3,
max_timesteps=100000,
buffer_size=50000,
train_freq=1,
batch_size=32,
print_freq=1,
checkpoint_freq=10000,
learning_starts=1000,
gamma=1.0,
target_network_update_freq=500,
exploration_fraction=0.1,
exploration_final_eps=0.02,
num_cpu=5,
callback=None,
scope='deepq',
pilot_tol=0,
pilot_is_human=False,
reuse=False,
using_supervised_goal_decoder=False):
# Create all the functions necessary to train the model
sess = U.get_session()
if sess is None:
sess = U.make_session(num_cpu=num_cpu)
sess.__enter__()
def make_obs_ph(name):
return U.BatchInput(env.observation_space.shape, name=name)
using_control_sharing = pilot_tol > 0
act, train, update_target, debug = co_build_train(
scope=scope,
make_obs_ph=make_obs_ph,
q_func=q_func,
num_actions=env.action_space.n,
optimizer=tf.train.AdamOptimizer(learning_rate=lr),
gamma=gamma,
grad_norm_clipping=10,
reuse=reuse,
using_control_sharing=using_control_sharing
)
act_params = {
'make_obs_ph': make_obs_ph,
'q_func': q_func,
'num_actions': env.action_space.n,
}
replay_buffer = ReplayBuffer(buffer_size)
# Initialize the parameters and copy them to the target network.
U.initialize()
update_target()
episode_rewards = [0.0]
episode_outcomes = []
saved_mean_reward = None
obs = env.reset()
prev_t = 0
rollouts = []
if pilot_is_human:
global human_agent_action
human_agent_action = init_human_action()
if not using_control_sharing:
exploration = LinearSchedule(schedule_timesteps=int(exploration_fraction * max_timesteps),
initial_p=1.0,
final_p=exploration_final_eps)
with tempfile.TemporaryDirectory() as td:
model_saved = False
model_file = os.path.join(td, 'model')
for t in range(max_timesteps):
masked_obs = obs if using_supervised_goal_decoder else mask_helipad(obs)
act_kwargs = {}
if using_control_sharing:
act_kwargs['pilot_action'] = env.unwrapped.pilot_policy(obs[None, :n_obs_dim])
act_kwargs['pilot_tol'] = pilot_tol if not pilot_is_human or (pilot_is_human and human_agent_active) else 0
else:
act_kwargs['update_eps'] = exploration.value(t)
action = act(masked_obs[None, :], **act_kwargs)[0][0]
new_obs, rew, done, info = env.step(action)
if pilot_is_human:
env.render()
time.sleep(sim_delay_for_human)
# Store transition in the replay buffer.
masked_new_obs = new_obs if using_supervised_goal_decoder else mask_helipad(new_obs)
replay_buffer.add(masked_obs, action, rew, masked_new_obs, float(done))
obs = new_obs
episode_rewards[-1] += rew
if done:
if t > learning_starts:
for _ in range(t - prev_t):
obses_t, actions, rewards, obses_tp1, dones = replay_buffer.sample(batch_size)
weights, batch_idxes = np.ones_like(rewards), None
td_errors = train(obses_t, actions, rewards, obses_tp1, dones, weights)
obs = env.reset()
episode_outcomes.append(rew)
episode_rewards.append(0.0)
if pilot_is_human:
global human_agent_action
human_agent_action = init_human_action()
prev_t = t
if pilot_is_human:
time.sleep(1)
if t > learning_starts and t % target_network_update_freq == 0:
# Update target network periodically.
update_target()
mean_100ep_reward = round(np.mean(episode_rewards[-101:-1]), 1)
mean_100ep_succ = round(np.mean([1 if x==100 else 0 for x in episode_outcomes[-101:-1]]), 2)
mean_100ep_crash = round(np.mean([1 if x==-100 else 0 for x in episode_outcomes[-101:-1]]), 2)
num_episodes = len(episode_rewards)
if done and print_freq is not None and len(episode_rewards) % print_freq == 0:
logger.record_tabular("steps", t)
logger.record_tabular("episodes", num_episodes)
logger.record_tabular("mean 100 episode reward", mean_100ep_reward)
logger.record_tabular("mean 100 episode succ", mean_100ep_succ)
logger.record_tabular("mean 100 episode crash", mean_100ep_crash)
logger.dump_tabular()
if checkpoint_freq is not None and t > learning_starts and num_episodes > 100 and t % checkpoint_freq == 0 and (saved_mean_reward is None or mean_100ep_reward > saved_mean_reward):
if print_freq is not None:
print('Saving model due to mean reward increase:')
print(saved_mean_reward, mean_100ep_reward)
U.save_state(model_file)
model_saved = True
saved_mean_reward = mean_100ep_reward
if model_saved:
U.load_state(model_file)
reward_data = {
'rewards': episode_rewards,
'outcomes': episode_outcomes
}
return ActWrapper(act, act_params), reward_data
def make_co_policy(
env, scope=None, pilot_tol=0, pilot_is_human=False,
n_eps=n_training_episodes, copilot_scope=None,
copilot_q_func=None, build_goal_decoder=None,
reuse=False, **extras):
if copilot_scope is not None:
scope = copilot_scope
elif scope is None:
scope = str(uuid.uuid4())
q_func = copilot_q_func if copilot_scope is not None else make_q_func()
return (scope, q_func), co_dqn_learn(
env,
scope=scope,
q_func=q_func,
max_timesteps=max_ep_len*n_eps,
pilot_tol=pilot_tol,
pilot_is_human=pilot_is_human,
reuse=reuse,
using_supervised_goal_decoder=(build_goal_decoder is not None),
**copilot_dqn_learn_kwargs
)
def str_of_config(pilot_tol, pilot_type, embedding_type, using_lander_reward_shaping):
return "{'pilot_type': '%s', 'pilot_tol': %s, 'embedding_type': '%s', 'using_lander_reward_shaping': %s}" % (pilot_type, pilot_tol, embedding_type, str(using_lander_reward_shaping))
# train and evaluate copilot
n_reps = 10
pilot_ids = ['sensor']
pilot_policies = [eval('%s_pilot_policy' % pilot_name) for pilot_name in pilot_ids]
embedding_type = 'rawaction'
using_lander_reward_shaping = True
pilot_tols = [0]
configs = []
for pilot_id, pilot_policy in zip(pilot_ids, pilot_policies):
if embedding_type != 'rawaction':
build_goal_decoder = build_build_goal_decoder(pilot_id)
else:
build_goal_decoder = None
for pilot_tol in pilot_tols:
configs.append((
str_of_config(pilot_tol, pilot_id, embedding_type, using_lander_reward_shaping),
{
'pilot_tol': pilot_tol,
'build_goal_decoder': build_goal_decoder,
'pilot_policy': pilot_policy,
'using_lander_reward_shaping': using_lander_reward_shaping,
'reuse': False
}))
reward_logs = {}
for config_name, config_kwargs in configs:
print(config_name)
reward_logs[config_name] = defaultdict(list)
co_env = make_co_env(**config_kwargs)
for i in range(n_reps):
(copilot_scope, copilot_q_func), (raw_copilot_policy, reward_data) = make_co_policy(
co_env, **config_kwargs)
for k, v in reward_data.items():
reward_logs[config_name][k].append(v)
reward_log_file = 'reward_logs.pkl'
with open(os.path.join(data_dir, reward_log_file), 'wb') as f:
pickle.dump(reward_logs, f, pickle.HIGHEST_PROTOCOL)
# Train and test on different pilots
pilot_tol_of_id = {
'noop': 0,
'laggy': 0.7,
'noisy': 0.4,
'sensor': 0
}
training_pilot_ids = list(pilot_tol_of_id.keys())
copilot_of_training_pilot = {}
copilot_path_of_training_pilot = lambda training_pilot_id: os.path.join(data_dir, 'pretrained_%s_copilot')
copilot_scope_of_training_pilot = lambda training_pilot_id: ('pretrained_%s_copilot_scope' % training_pilot_id)
for training_pilot_id, pilot_tol in pilot_tol_of_id.items():
pilot_policy = eval('%s_pilot_policy' % training_pilot_id)
copilot_scope = copilot_scope_of_training_pilot(training_pilot_id)
config_kwargs = {
'pilot_policy': pilot_policy,
'pilot_tol': pilot_tol,
'copilot_scope': copilot_scope,
'copilot_q_func': make_q_func()
}
co_env = make_co_env(**config_kwargs)
(copilot_scope, copilot_q_func), (raw_copilot_policy, reward_data) = make_co_policy(co_env, **config_kwargs)
copilot_of_training_pilot[training_pilot_id] = (copilot_scope, raw_copilot_policy)
copilot_path = copilot_path_of_training_pilot(training_pilot_id)
save_tf_vars(copilot_scope, copilot_path)
def make_copilot_policy(training_pilot_id, eval_pilot_policy, pilot_tol):
copilot_scope, raw_copilot_policy = copilot_of_training_pilot[training_pilot_id]
def copilot_policy(obs):
with tf.variable_scope(copilot_scope, reuse=None):
masked_obs = mask_helipad(obs)[0]
pilot_action = eval_pilot_policy(masked_obs[None, :n_obs_dim])
if masked_obs.size == n_obs_dim:
feed_obs = np.concatenate((masked_obs, onehot_encode(pilot_action)))
else:
feed_obs = masked_obs
return raw_copilot_policy._act(
feed_obs[None, :],
pilot_tol=pilot_tol,
pilot_action=pilot_action
)[0][0]
return copilot_policy
n_eval_eps = 100
cross_evals = {}
for training_pilot_id, training_pilot_tol in pilot_tol_of_id.items():
# load pretrained copilot
copilot_scope = copilot_scope_of_training_pilot(training_pilot_id)
training_pilot_policy = eval('%s_pilot_policy' % training_pilot_id)
config_kwargs = {
'pilot_policy': training_pilot_policy,
'pilot_tol': training_pilot_tol,
'copilot_scope': copilot_scope,
'copilot_q_func': make_q_func(),
'reuse': True
}
co_env = make_co_env(**config_kwargs)
make_co_policy(co_env, **config_kwargs)
copilot_path = copilot_path_of_training_pilot(training_pilot_id)
load_tf_vars(copilot_scope, copilot_path)
# evaluate copilot with different pilots
for eval_pilot_id, eval_pilot_tol in pilot_tol_of_id.items():
eval_pilot_policy = eval('%s_pilot_policy' % eval_pilot_id)
copilot_policy = make_copilot_policy(training_pilot_id, eval_pilot_policy, eval_pilot_tol)
co_env = make_co_env(pilot_policy=eval_pilot_policy)
cross_evals[(training_pilot_id, eval_pilot_id)] = [run_ep(copilot_policy, co_env, render=False)[:2] for _ in range(n_eval_eps)]
with open(os.path.join(data_dir, 'cross_evals.pkl'), 'wb') as f:
pickle.dump(cross_evals, f, pickle.HIGHEST_PROTOCOL)
| 1.0-lunarlander-sim.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from sklearn import datasets
import numpy as np
import matplotlib.pyplot as plt
iris = datasets.load_iris()
digits = datasets.load_digits()
print(np.shape(digits.data))
print(np.shape(digits.target))
from sklearn import svm
clf = svm.SVC(gamma=0.001, C=100.)
clf.fit(digits.data[:-1], digits.target[:-1])
clf.predict(digits.data[-1:])
# <h3>Linear Regression</h3>
#
# class sklearn.linear_model.LinearRegression(fit_intercept=True, normalize=False, copy_X=True, n_jobs=None)
#
# - fit_intercept : boolean, optional, default True
# whether to calculate the intercept for this model. If set to False, no intercept will be used in calculations (e.g. data is expected to be already centered).
# - normalize : boolean, optional, default False
# This parameter is ignored when fit_intercept is set to False. If True, the regressors X will be normalized before regression by subtracting the mean and dividing by the l2-norm. If you wish to standardize, please use sklearn.preprocessing.StandardScaler before calling fit on an estimator with normalize=False.
# - copy_X : boolean, optional, default True
# If True, X will be copied; else, it may be overwritten.
# - n_jobs : int or None, optional (default=None)
# The number of jobs to use for the computation. This will only provide speedup for n_targets > 1 and sufficient large problems. None means 1 unless in a joblib.parallel_backend context. -1 means using all processors. See Glossary for more details.
#
# <b>methods </b>
# <table border="1" class="longtable docutils">
# <colgroup>
# <col width="10%">
# <col width="90%">
# </colgroup>
# <tbody valign="top">
# <tr class="row-odd"><td><a class="reference internal" href="#sklearn.linear_model.LinearRegression.fit" title="sklearn.linear_model.LinearRegression.fit"><code class="xref py py-obj docutils literal"><span class="pre">fit</span></code></a>(self, X, y[, sample_weight])</td>
# <td>Fit linear model.</td>
# </tr>
# <tr class="row-even"><td><a class="reference internal" href="#sklearn.linear_model.LinearRegression.get_params" title="sklearn.linear_model.LinearRegression.get_params"><code class="xref py py-obj docutils literal"><span class="pre">get_params</span></code></a>(self[, deep])</td>
# <td>Get parameters for this estimator.</td>
# </tr>
# <tr class="row-odd"><td><a class="reference internal" href="#sklearn.linear_model.LinearRegression.predict" title="sklearn.linear_model.LinearRegression.predict"><code class="xref py py-obj docutils literal"><span class="pre">predict</span></code></a>(self, X)</td>
# <td>Predict using the linear model</td>
# </tr>
# <tr class="row-even"><td><a class="reference internal" href="#sklearn.linear_model.LinearRegression.score" title="sklearn.linear_model.LinearRegression.score"><code class="xref py py-obj docutils literal"><span class="pre">score</span></code></a>(self, X, y[, sample_weight])</td>
# <td>Returns the coefficient of determination R^2 of the prediction.</td>
# </tr>
# <tr class="row-odd"><td><a class="reference internal" href="#sklearn.linear_model.LinearRegression.set_params" title="sklearn.linear_model.LinearRegression.set_params"><code class="xref py py-obj docutils literal"><span class="pre">set_params</span></code></a>(self, \*\*params)</td>
# <td>Set the parameters of this estimator.</td>
# </tr>
# </tbody>
# </table>
# +
from sklearn.linear_model import LinearRegression
X = np.array([[1, 1], [1, 2], [2, 2], [2, 3]])
# y = 1 * x_0 + 2 * x_1 + 3
y = np.dot(X, np.array([1, 2])) + 3
# np.daot is a scalar product of each input of X and [1,2]
plt.plot(X,y,'.')
plt.show
reg = LinearRegression().fit(X, y)
reg.score(X, y)
reg.coef_
reg.intercept_
reg.predict(np.array([[3, 5]]))
# -
# <h3>Example of a linear reg model</h3>
#
# Weather Conditions in World War Two: Is there a relationship between the daily minimum and maximum temperature? Can you predict the maximum temperature given the minimum temperature?
#
# https://www.kaggle.com/smid80/weatherww2/data
# +
import csv
d=[]
with open('LinReg_weather/Summary of Weather.csv') as csv_file:
csv_reader = csv.reader(csv_file, delimiter=',')
line_count = 0
for row in csv_reader:
if line_count == 0:
head = row
line_count += 1
else:
d.append(row)
line_count += 1
print(f'Processed {line_count} lines.')
# -
data = np.array(d)
data[1]
head
y = data[:,4]
np.shape(y)
y = y.reshape(-1, 1)
y = y.astype(np.float64)
from sklearn import preprocessing
X = data[:,5]
np.shape(X)
X = X.reshape(-1, 1)
X = X.astype(np.float64)
# X = preprocessing.scale(X)
fig = plt.figure()
ax = fig.add_axes([0,0,3,3])
ax.scatter(X, y, color = "red")
plt.show
# +
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3)
# -
from sklearn.linear_model import LinearRegression
reg = LinearRegression().fit(X_train,y_train)
print('train accuracy: ',reg.score(X_train,y_train)*100)
print('test accuracy: ',reg.score(X_test,y_test)*100)
reg.predict([[-2]])
fig = plt.figure()
ax = fig.add_axes([0,0,3,3])
ax.scatter(X_train, y_train, color = "red")
ax.plot(X_train, reg.predict(X_train), color = "g")
plt.show
fig = plt.figure()
ax = fig.add_axes([0,0,3,3])
ax.hist(data[:,5], bins = [-10,-6,-,75,100],histtype = 'step')
ax.bar(data[:,5],data[:,4])
plt.show()
# <h1>---------------------------------------------------------------------------------------</h1>
# <h3>Logistic Regression </h3>
#
| SciKitLearn/ScikitLearn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.5
# language: python
# name: py35
# ---
import pandas as pd
import numpy as np
import os
# # Groupby
# As I pointed out in the first part of this lesson, tidy data is only useful if we have tools that work with it in a consistent and reproducable manner. One such tools is a `groupby` method of `DataFrame`, which provides a powerful interface to apply any operation based on groupping variables, and we will talk about it in detail in the current section.
#
# It turns out that very frequently we need to do some operation based on a groupping variable. A common example is calculating mean of each group (e.g. performance of each subject, or performance on each type of stimuli, etc). This can be thought of as making 3 separate actions:
# - Splitting the data based on a groupping variable(s)
# - Applying a function to each group separately
# - Combining the resulting values back together
#
# Based on these 3 actions, this approach is called *Split-Apply-Combine* (SAC) [1].
#
# [1] <NAME>. "The split-apply-combine strategy for data analysis." Journal of Statistical Software 40.1 (2011): 1-29.
# <img src="http://nbviewer.jupyter.org/github/jakevdp/PythonDataScienceHandbook/blob/master/notebooks/figures/03.08-split-apply-combine.png"></img>
# From ["Aggregation and groupping" chapter](http://nbviewer.jupyter.org/github/jakevdp/PythonDataScienceHandbook/blob/master/notebooks/03.08-Aggregation-and-Grouping.ipynb) of ["Python Data Science Handbook"](http://nbviewer.jupyter.org/github/jakevdp/PythonDataScienceHandbook/blob/master/notebooks/Index.ipynb) by <NAME>
# A lot of operations on data can be thought of as SAC operations. These include calculating sums, means, standard deviations and other parameters of the groups' distributions; transfromations of data, such as normalization or detrending; plotting based on group, e.g. boxplots; and many other. (Some operations cannot be thought of as purely SAC, most prominently those in which data from the same group is used several times, e.g. rolling window means.)
#
# A traditional way of doing these operations in include loops, where on each iteration a subset of data is selected and processed. Loops, however, are slow and usually require a lot of code, which makes them difficult to read, and are not easily extendible from 1 to several groupping variables.
#
# `Groupby` is a method of `DataFrames` which makes any SAC operation easy to perform and read.
#
# >**Note**: Tidy data is the most convenient form for making SAC operations, because you always have access to any combination of your groupping variables due to them being always separated in columns.
#
# Let's see a toy example of using a `groupby` operation instead of a loop.
df = pd.DataFrame({'group': ['A', 'B', 'C', 'A', 'B', 'C'],
'data': range(6)})
df
# Let's say I want to calculate a sum of `data` column, based on `group` variable and save it in a `Series`. I can do it with a loop:
# +
result = pd.Series()
groups = df['group'].unique()
for g in groups:
data = df.loc[df['group']==g, 'data']
result[g] = np.sum(data)
result
# -
# This code does the job, but it is quite long. If I try to shorten it, it will become very difficult to read:
# +
result = pd.Series()
for g in df['group'].unique():
result[g] = np.sum(df.loc[df['group']==g, 'data'])
result
# -
# Now let's try to do the same thing with `groupby`:
df.groupby('group')['data'].sum()
# See that it is really short and concise and readable. Moreover, let's say I have a more complicated example with several groupping variables:
df = pd.DataFrame({'group1': ['A', 'B', 'C']*3,
'group2': ['A']*4 + ['B']*1 + ['C']*4,
'data': range(9)})
df
# Trying to calculate a sum based on these several groups requires significantly more code with loops. With `groupby` it is as easy as adding another groupping variable in the `groupby` attributes:
result = df.groupby(['group1','group2'])['data'].sum()
result
# >**Pro-tip**: You may notice that in the resulting `Series` index has 2 levels: `group1` and `group2`. This is referred to as *Hierarchical index* or `MultiIndex`, and is a way to stack several dimensions of data. We won't go much into the details of `MultiIndex` (if you wish to learn more, you may refer to [this section](http://nbviewer.jupyter.org/github/jakevdp/PythonDataScienceHandbook/blob/master/notebooks/03.05-Hierarchical-Indexing.ipynb) of [Python Data Science Handbook](http://nbviewer.jupyter.org/github/jakevdp/PythonDataScienceHandbook/blob/master/notebooks/Index.ipynb) and to [MultiIndex](http://pandas.pydata.org/pandas-docs/stable/advanced.html) section of `pandas` documentation. For our purposes we just need to know 2 things: how to index a `MultiIndex` and how to *unstack* dimensions to turn it into a table:
# get an element with group1 = A and group2 = C
result[('A','C')]
# unstack levels of multiindex (turn one of them into a column)
result.unstack()
# Overall, `groupby` is an extremely useful tool for making group-based operations quickly and more readible. Let's see some concrete examples of how you can use it. We will work on the data in the food preferences task provided by <NAME>. Let's load it first and do some cleanup:
df = pd.read_csv('data/Paolo.csv')
# drop old index column
df.drop('Unnamed: 0', axis='columns', inplace=True)
df['cond'].replace({1: 'high vs high', 2: 'low vs low',
3: 'high vs low', 4: 'low vs high'}, inplace=True)
df['congr'].replace({0: 'same', 1: 'different'}, inplace=True)
df['session'].replace({0: 'fed', 1: 'hungry'}, inplace=True)
print(df.shape)
df.head()
# The data contains 4 subjects:
df['subj_num'].unique()
# Let's calculate mean reaction time for each subject:
df.groupby('subj_num')['rt'].mean()
# Subjects also seem to have more that 1 session, so we might want to compute mean for each session separately:
rt_subject_session = df.groupby(['subj_num','session'])['rt'].mean()
rt_subject_session
# # <font color='DarkSeaGreen '>Exercise</font>
# In the cell below calculate mean response for each food item.
#
#
# As we saw above, `pandas` provides shortcuts to applying some frequent functions, such as `mean()`, `std()`, `count()`, `min()`, `max()`. However, we can apply any function to the groups. to do that, there are 3 methods: `aggregate()`, `transform()` and `apply()`. Each of these methods require a function (the one you want to apply to the data) as an argument.
#
# ## Aggregate
# `aggregate()` can apply any function, which returns a single value for each group (in other words, it *aggregates* a group to a single value). This is what mean, std, count, min, max, and others are. Instead of writing `df.groupby('subj_num')['rt'].mean()` we could've passed a `np.mean` function to calculate means:
df.groupby('subj_num')['rt'].aggregate(np.mean)
# You can also specify several functions in a list, and `aggregate()` will return results of all of them in a neat table:
df.groupby('subj_num')['rt'].aggregate([np.mean, np.std, np.median])
# More importantly, you can create any function and pass it to `aggregate()` and the function will be applied to each group. The only limitation is that code will assume that the function returns a single value, e.g. calculate half of mean:
# +
def hafl_mean(x):
"""Calculate half of the mean"""
mean = np.mean(x)
return mean/2
df.groupby('subj_num')['rt'].aggregate(hafl_mean)
# -
# ## Transform
# `transform()` works exactly like `aggregate()`, but it expects a function to return a `Series` or an `array` of the same size as input. It will handle the cases when you want to tranform the data. For example, we could subtract the mean reaction time for each subject:
# +
def subtract_mean(x):
return x - np.mean(x)
df['rt_minus_mean'] = df.groupby('subj_num')['rt'].transform(subtract_mean)
df.head()
# -
# # <font color='DarkSeaGreen '>Exercise</font>
# In the cell below calculate standard score (*z-score*) on reaction time for each subject using `groupby` and `transform`. Save z scores to a new column.
#
# See which 10 items require highest reaction times on average in all subjects.
# All other cases, which don't fall within `aggregate` and `transform` can be handled by `apply` method. In reality, `apply` can act as both `aggregate` and `transform` in most circumstances, but it is slower (because it cannot assume output shape) and cannot do certain things, for example, aggregate several functions at once like `aggregate` method can.
# ## Looping with groupby
# `groupby`-`apply` combination lets us in general avoid loops, but sometimes you might still need to use them. For example, this can happen when you want to do plotting by group. `groupby` can also simplify that, because it supports iteration through itself. When you do it, on each iteration it will give 2 values: one for the name of the group (basically, groupping variable value) and the values of the group.
# assign a grouby object to a variable
groupped = df.groupby('subj_num')['rt']
# iterate through groupby object
for name, data in groupped:
# groupping variable value
print(name)
# shape of the data: in this case the 'rt' values for each group
print(data.shape)
# # `DataFrame` and `Series` transformations
# Now that you know the power of `groupby` and having data in a tidy format, let's talk about how to get there. In general, you should become comfortable with transforming your data to any shape you want, because the tools you might want to use, won't necessarily work with tidy data. `pandas` provides a lot of ways to tranform `Series` and `DataFrame` objects.
#
# ## `Set`, `reset` index
# Index is very useful for retrieving values, but also for other things. For example, as we will see in the visualization lesson, when plotting a `Series`, index will be automatically assumed to be the X axis, and the values will become the Y axis. This is useful for quick exploratory visualization.
#
# Main methods to interact with the index are `set_index()` and `reset_index()`. First takes a column and makes it into a new index:
df_items = df.set_index('item')
df_items.head()
# if you say append=True, you can keep the old index too, which will result in a MultiIndex
df.set_index('item', append=True).head()
# `reset_index()` will make the old index into a columns and instead create a new index, which has values from `0` to the number of rows minus 1:
# our DataFrame indexed by items
df_items.head()
# let's reset index
df_items.reset_index().head()
# These two methods make working with index very dynamic -- you can set it and reset it to become a normal column again whenever you need. You can also set several columns (pass them as a list to `set_index`) and create a `MultiIndex`.
#
# ## Melt
# The concept of melting is related to tidying the data. `melt` function takes all columns of the `DataFrame` and creates 2 columns from them: one with groupping variable (former name of the column) and another with the value variable. If applied correctly, the resulting *molten* `DataFrame` will be tidy.
#
# Let's see a toy example:
untidy = pd.DataFrame({'treatment_a':[np.nan, 16, 3],'treatment_b':[2,11,1]})
untidy
# let's melt
pd.melt(untidy)
# Note how the data is reshaped. What were the names of the columns in the untidy `DataFrame` (`treatment_a` and `treatment_b`) are now the groupping variable. The values inside the table are now all in the single "value" column.
# you can also specify the names of the resulting columns
pd.melt(untidy, var_name='treatment', value_name='measurement')
# Frequently you want to melt only certain columns, because some are already groupping variable. Specify them as `id_vars` in the `melt` function and they will not be changed:
# in this example "person" is already a separated variable
untidy = pd.DataFrame({'treatment_a':[np.nan, 16, 3],'treatment_b':[2,11,1],
'person':['<NAME>', '<NAME>','<NAME>']})
untidy
pd.melt(untidy, id_vars='person', var_name='treatment', value_name='measurement')
# Let's see another example, taken directly from the [lesson on tidy data](http://nbviewer.jupyter.org/github/antopolskiy/sciprog/blob/master/002_data_organization_00_slides.ipynb):
income_untidy = pd.read_csv(os.path.join('data','pew.csv'))
print(income_untidy.shape)
income_untidy.head()
# In this case all columns except for `religion` have the same variable (count of people who belongs to this group), so we keep `religion` and melp all other columns:
income_tidy = pd.melt(income_untidy,id_vars='religion',var_name='income',value_name='count')
print(income_tidy.shape)
income_tidy.head()
# # Pivot table
# Pivoting is another way of transforming the `DataFrames`, which is usually used to tranform a tidy `DataFrame` in some other form. For example, it can be used to undo melting. Using method `pivot_table` is easy: simply think about which column you want to have as and index and which as columns.
# molten dataframe
income_tidy.head()
# pivoting to undo melting
income_tidy.pivot_table(columns='income', index='religion')
# But pivoting can achieve much more than that. Let's look at another example. This dataset contains number of births for each day from 1969 to 2008:
births = pd.read_csv(os.path.join('data','births.csv'))
births.head()
# Let's say we want to calculate the total number of births for each year for boys and girls to see how the gender proportions change over the years. We could achieve it with `groupby`:
births.groupby(['year','gender'])['births'].sum().head(10)
# We could then use `unstack` on the resulting `Series` to create a nice table:
births.groupby(['year','gender'])['births'].sum().unstack()
# Pivot table can do the same and in some cases can be more readable, because when we pivot we don't need to think about groupping, but instead we think about what kind of table we want to get in the end. In this case I think to myself: "I want *year* to be the index, *genders* will be the columns. I will take the *births* columns and *sum* them up for each resulting group". The syntax of `pivot_table` repeats this thinking almost exactly:
births_year_gender = births.pivot_table(index='year', columns='gender', values='births', aggfunc=np.sum)
births_year_gender
# Let's see another example on Paolo's food preference data.
df.head()
# I want to create a table with mean reaction times with rows being session type and columns being the condition.
df.pivot_table(values='rt', index='session', columns='cond', aggfunc=np.mean)
# # <font color='DarkSeaGreen '>Exercise</font>
# Using `births` dataset, create a table in which there would be total number of births for each month for each year. Do it using `groupby-aggregate-unstack` and using `pivot_table`.
# # <font color='DarkSeaGreen '>Exercise</font>
# Using food preference dataset, create a table in which the index would be items, columns would be session type and the values would be mean response.
# # Concatenation and merging
# +
import pandas as pd
import numpy as np
# nevermind this part, this is just to display several tables alongside
class display(object):
"""Display HTML representation of multiple objects"""
template = """<div style="float: left; padding: 10px;">
<p style='font-family:"Courier New", Courier, monospace'>{0}</p>{1}
</div>"""
def __init__(self, *args):
self.args = args
def _repr_html_(self):
return '\n'.join(self.template.format(a, eval(a)._repr_html_())
for a in self.args)
def __repr__(self):
return '\n\n'.join(a + '\n' + repr(eval(a))
for a in self.args)
# +
# this function will quickly create DataFrames for our toy examples
def make_df(cols, ind):
"""Quickly make a DataFrame"""
data = {c: [str(c) + str(i) for i in ind]
for c in cols}
return pd.DataFrame(data, ind)
# example DataFrame
make_df('ABCD', range(3))
# -
# Let's remind ourselves **3 principles of tidy data**:
# - Each variable forms a column
# - Each observation forms a row
# - Each type of observation forms a separate table
#
# If first 2 priciples are rather easy to digest, the third one at times seems to make life harder rather than easier. In fact, if you split your data into several tables and don't know how to merge it back in a format you need for a certain analysis, you can lose a lot of time. Merging data from different tables can be very daunting if done manually. Data can have diverse type, some rows or columns can be present only in one of the tables, etc. Historically, the problems of merging were addressed by databases, such as SQL. `pandas` provides a lot of functionality in this domain.
#
# Besides that, merging is often necessary when tidying data from different sources, for example, you might have a table for each participant and you want to put them all together.
#
# # Append
# There are several distinct types of putting tables together. The easiest one to understand is `append` -- it is a method of `DataFrame` which will take another `DataFrame` and put it directly under the first one, independent of `index` (`index` is preserved from both `DataFrames`).
df1 = make_df('ABC',range(3))
df1
# append another copy of the same DataFrame
df1.append(df1)
# `append` will try to match columns. If some columns are present in one `DataFrame` but not in another, it will put missing values where appropriate:
# columns A exists only in df1, column D only in df2
df1 = make_df('ABC',range(3))
df2 = make_df('BCD',range(3))
display('df1','df2','df1.append(df2)')
# `append` is very useful when you want to quickly put together some tables with the same type of data, for example tables for separate subjects. But that's just about its functionality.
#
# # Concat
# `concat` can be thought of as generalized `append`. It can do all the things `append` can and much more. For example, it can take a list of `DataFrames` and put them all together in `append`-like manner:
pd.concat([df1, df1, df2, df2])
# You can also pass a list of `keys` to `concat` (same size as list of `DataFrames` to concatenate) and each `DataFrame` will have its own key in the index. This is useful in a situation where you merge several subjects and want to keep each one labeled with which subject this came from.
pd.concat([df1, df1, df2, df2], keys=['df1','df2','df3','df4'])
# You can also concatenate along `columns` instead of `index` by passing `axis` argument. In this case `concat` will try to match `index` (as it tried to match columns when you were concatenating along `index`):
pd.concat([df1, df1, df2, df2], axis='columns')
# Let's see a real example of `concat` use. Here I load data from 2 subjects, which is stored in separate *.mat* files. I concatenate them to create a signle table, and I reset the index. Then I name the columns according to the order given me by the person who conducted the experiment. Resulting is one tidy `DataFrame` with 2 subjects.
# `concat` is useful for any kind of simple concatenation where we have the same type of data in different tables. However, when we have different *types* of observations in different tables, `concat` will fail us.
#
# # Merge
# If merging data sounds confusing to you at any point, it is because it is. There is a whole area of math called *relational algebra*, which creates the theoretical underpinnings of databases and how they work. We won't study any of that, not only it requires a course of its own, but it is also not very frequent for scientists to deal with the kind of data that requires databases.
#
# Merging, however, is important if you want to work with tidy data and avoid data duplication (which is not only inefficient, but also invites errors). So we will learn a bit about that.
#
# In general there are 3 types of joins: one-to-one, many-to-one, and many-to-many. Pandas function `merge()` provides an interface to do all of them, depending on the inputs. The first (one-to-one) referers to the simplest case, when you have 2 sources, and none of them have duplicate entries. In this case joining is usually easy, and basically reminds a concatenation. Let's see it with a toy example:
df1 = pd.DataFrame({'employee': ['Bob', 'Jake', 'Lisa', 'Sue'],
'department': ['Accounting', 'Engineering', 'Engineering', 'HR']})
df2 = pd.DataFrame({'employee': ['Lisa', 'Bob', 'Jake', 'Sue'],
'hire_date': [2004, 2008, 2012, 2014]})
display('df1', 'df2')
# Here the 2 tables have a common column `employee`, but the order is different. We want to merge the two tables consistently, to see the hire date for our different departments. In this case `merge` will automatically find the matching column:
df3 = pd.merge(df1, df2)
df3
# >**Note**: if we tried to perform `concat` here with `axis='columns'`, it would match the `index`, but not the `employee`. You could get around it by first setting the `employee` as the index in both tables, and then perform the `concat` on columns. You could then reset the index and get out the same table. But it is inefficient. Nevertheless, let's do it for the sake of demonstation:
df1_e = df1.set_index('employee')
df2_e = df2.set_index('employee')
df_e = pd.concat([df1_e, df2_e], axis='columns')
display("df1_e","df2_e","df_e")
# One-to-many is when one of your `DataFrames` contains duplicate entries. `merge` will understand that and try to fill in the values appropriately:
df4 = pd.DataFrame({'department': ['Accounting', 'Engineering', 'HR'],
'supervisor': ['Carly', 'Guido', 'Steve']})
display('df3','df4')
# note how the supervisor column in the resulting DataFrame has Guido
# across from every person in Engineering department
df5 = pd.merge(df3, df4)
df5
# Many-to-many is the most confusing type of join, but it is nevertheless well defined mathematically. Consider the following, where we have a `DataFrame` showing one or more skills associated with a particular department. By performing a many-to-many join, we can recover the skills associated with any individual person. Note that some entries in both `df1` and `df6` had to be duplicated; also "R&D" group disappeared in the joined `DataFrame`, because it had no pairings within `df1`.
df6 = pd.DataFrame({'department': ['Accounting', 'Accounting',
'Engineering', 'Engineering', 'HR', 'HR', 'R&D'],
'skills': ['math', 'spreadsheets', 'coding', 'linux',
'spreadsheets', 'organization', 'science']})
df7 = pd.merge(df1, df6)
display('df1', 'df6', 'df7')
# Some things need to be pointed out.
#
# **First**, when you merge, you can specify a parameter `how`, which can have 1 of 4 values: *left*, *right*, *outer* or *inner*. This controls which values remain in the resulting `DataFrame` if some values are present only in one of the `DataFrames` you're merging. By default `how='inner'`, and it means that the resulting `DataFrame` will have the *intersection* of values from the input `DataFrames`, that is, only values present in both `DataFrames` will be present in the result. That is why we don't have *R&D* in the `df7`: there is no match for it in the `df1`. `outer` is the opposite of `inner` -- all the values will be present in the result. Let's try to do the same merge, but with `how='outer'`:
df7 = pd.merge(df1, df6, how='outer')
display('df1', 'df6', 'df7')
# See how now there is *R&D* in the resulting `DataFrame`, although there is no employee who is in this department.
#
# `left` and `right` just say that values from the first or the second of the input `DataFrames` will be used. In this case, if I used `how='left'`, only values from `df1` would be used, and for `how='right'` -- only from `df6`. (*Left* and *right* just refer to their positions as the inputs to the `merge` function; this terminology, as well as *inner* and *outer*, is taken directly from the database systems, otherwise they might as well be named "first" and "second").
# **Second**: `merge` will try to infer which column(s) to use in both `DataFrames` to match the data consistently. However, it is safest to specify it manually, then the outcome is most predictable. If you want to use a certain column, specify `left_on` parameter (for the first input `DataFrame`) and `right_on` (for the second one). This is extremely useful for when you have several columns matching names and they are not consistent with one another, and you want the outcome to be 100% predictable.
# the result is equivalent to what we had before, but we have more control
df7 = pd.merge(df1, df6, left_on='department', right_on='department')
display('df1', 'df6', 'df7')
# Sometimes you want to use index in one of the `DataFrames` for matching. In this case just specify `left_index=True` instead of `left_on` (same for the `right`).
#
# Let's see it with an example. We load and concatenate the data from a vibration experiment:
# +
from scipy.io import loadmat
s1_mat = loadmat('data/Ale_subj1.mat')
s1_df = pd.DataFrame(s1_mat['Subject1'])
print('Subject1 df shape:', s1_df.shape)
s2_mat = loadmat('data/Ale_subj2.mat')
s2_df = pd.DataFrame(s2_mat['Subject2'])
print('Subject2 df shape:', s2_df.shape)
df_vibr = pd.concat([s1_df, s2_df])
df_vibr = df_vibr.reset_index(drop=True)
df_vibr.columns = ['id_subj','session','trial','s1_int','s1_dur','s1_seed','inter_stim_delay',
's2_int','s2_dur','s2_seed','pre_stim_delay','task_type','rewarded_choice',
'subj_choice','s1_motor','s2_motor']
print('Concatenated df shape:',df_vibr.shape)
df_vibr.head()
# -
# Associated with these data, there is some information about the subjects in the Excel spreadsheet:
#
# > When you try to read the `.xlsx` file, you might get an error "`No module named 'xlrd'`". This is because `pandas` is using another module to load the Excel file, and you need to install that module. Open your computer's terminal (console or command prompt ("cmd") in Windows) and run `pip install xlrd`. Now it should work.
df_vibr_subj = pd.read_excel('data/Ale_subj_info.xlsx')
df_vibr_subj
# If we wanted to look at, for example, performance depending on gender or age, we would have to merge these 2 tables to perform a `groupby`. If we try to do it by hand, it would be very cumbersom. Instead, let's do it using `merge`. First I want to make the column `id_subj` in the `df_vibr_subj` so that it matches the values in the `df_vibr` (i.e. now we have `Sub1`, but we should have just `1`). Then I make it index of the subjects table. I will also drop some other columns so that the result is clearer (otherwise after merge we will have all the columns there and it might be a bit confusing; also, in this example we just need gender and age, so it makes sense to merge in only these columns).
# rename column
df_vibr_subj.rename(columns={'Subjects':'id_subj'}, inplace=True)
# mapping of old values to new (see Pro-tip below)
subj_mapping_dict = {name_old:int(name_old[-1]) for name_old in df_vibr_subj['id_subj'].unique()}
# replace values based on mapping
df_vibr_subj['id_subj'] = df_vibr_subj['id_subj'].replace(subj_mapping_dict)
# set id_subj as index
df_vibr_subj.set_index('id_subj', inplace=True)
df_vibr_subj = df_vibr_subj[['age','Gender']]
df_vibr_subj
# > **Pro-tip**: This line
#
# > `subj_mapping_dict = {name_old:int(name_old[-1]) for name_old in df_vibr_subj['id_subj'].unique()}`
#
# >creates a dictionary with mappings for replacing Sub1 with 1, Sub2 with 2, etc; this is called "dict comprehension" and is just an extension of list comprehensions to create a dictionary; go here to learn more: https://www.python.org/dev/peps/pep-0274/
#
# Now that we have the subject's id in both tables, we can merge. I specify `id_subj` as target merging column for `df_vibr` and `index` for `df_vibr_subj`.
df_result = pd.merge(df_vibr, df_vibr_subj, left_on='id_subj', right_index=True)
df_result
# *Voilà!* Now we have age and gender for every trial (last columns) and we could group based on these and calculate some statistics. Note that although in the `df_vibr_subj` we have many more subjects (7 in total), in the `df_vibr` we only have data for 2 subjects. Because the default merge is *inner*, only the data from the 2 subjects present in both tables is present in the final table.
#
# Now, for example, we can calculate avegare subject's choice for gender and age (in this case we have only 2 subjects and they are both males, so it doesn't make much sense; but if you had many, it would work flawlessly):
df_result.groupby(['Gender','age'])['subj_choice'].mean()
# # <font color='DarkSeaGreen '>Exercise</font>
# Load 2 tables from the data folder: `toy_subjects.csv` and `toy_scores.csv` (use `read_csv` function with `index_col=0` parameter to make the first column into index). In the `toy_scores` calculate the mean `score` for male and female subjects. To do it, you'll need to merge the 2 tables, then group by gender.
# # Where to go from here
# Here we have only scratched the surface of merging. If you ever need to do complicated joins or are just interested in learning more about joins with `pandas`, I highly recommend <a href="http://nbviewer.jupyter.org/github/jakevdp/PythonDataScienceHandbook/blob/master/notebooks/03.07-Merge-and-Join.ipynb">Combining Datasets: Merge and Join</a> section on the "Python Data Science Handbook" by <NAME> and <a href="http://pandas.pydata.org/pandas-docs/stable/merging.html">Merge, join, and concatenate</a> section of the `Pandas` documentation.
#
# Besides, at this point you can go and review the code in the `002_data_organization_00_slides` notebook, because now you should be able to understand everything that is going on there.
| 08_python_pandas_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: pygeno (Python 3.7.1)
# language: python
# name: pygeno-python3.7.1
# ---
import pyGeno.Genome as pgg
import pyGeno.tools.UsefulFunctions as uf
# ## Initialize genome and select gene
genome_name = 'GRCh38.98'
gene_name = 'POMP'
# %%time
ref = pgg.Genome(name=genome_name)
gene = ref.get(pgg.Gene, name=gene_name, gen=False)[0] # gen=False returns list, not generator
print('Strand:', gene.strand)
# ## From transcripts to exons
# %%time
for transcript in gene.get(pgg.Transcript):
print(transcript.id)
for exon in transcript.get(pgg.Exon, gen=True):
print(" >", exon.id)
# ## From exons to transcripts
from collections import defaultdict
exon_dict = {'CDS': defaultdict(list), 'NotCDS': defaultdict(list)}
for exon in gene.get(pgg.Exon, gen=True):
exon_dict['CDS' if exon.hasCDS() else 'NotCDS'][exon.id].append(exon.transcript.id)
exon_dict
# ## Choose a coding exon
# choose a coding exon
exon_id = list(exon_dict['CDS'].keys())[0]
exon = gene.get(pgg.Exon, id=exon_id, gen=False)[0]
# ### Peep into the object structure
# +
print('UTR5:', exon.UTR5)
print('CDS:', exon.CDS)
print('UTR3:', exon.UTR3)
print()
print('sequence:', exon.sequence)
assert exon.sequence == ''.join(exon.UTR5 + exon.CDS + exon.UTR3)
# -
# ### Translate in 6 frames
uf.translateDNA_6Frames(exon.CDS)
# ### Easily get the protein sequence corresponding to the transcript containing that exon
exon.transcript.protein.sequence
| pyGeno/examples/genomic_graph.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
fname = input('Enter File: ')
if len(fname) < 1 : fname = 'Focus_Group_Transcribtion.txt'
hand = open(fname)
di = dict()
for lin in hand:
lin = lin.rstrip()
wds = lin.split()
for w in wds:
di[w] = di.get(w,0)+1
#print(di)
# +
tmp = list()
for k, v in di.items():
newt = (v,k)
tmp.append(newt)
print('Flipped',tmp)
# -
tmp = sorted(tmp, reverse = True)
print('Sorted',tmp)
for v,k in tmp[:1000]:
print(k,v)
word= ['Quindi']
with open('filteredtext616.txt', 'r') as infile:
newlist= [i for i in infile.read().split() if i!=word]
with open('filteredtext616.txt','w') as outfile:
outfile.write("\n".join(newlist))
# +
infile = "focus_group1.txt"
outfile = "cleaned_file1.txt"
delete_list = ["fare\n", " se\n", " quello\n"]
fin = open(infile)
fout = open(outfile, "w+")
for line in fin:
for word in delete_list:
line = line.replace(word, "")
fout.write(line)
fin.close()
fout.close()
# -
conda install -c conda-forge wordcloud
conda update -n base -c defaults conda
# +
import matplotlib.pyplot as plt
from wordcloud import WordCloud, STOPWORDS
def random_color_func(word=None, font_size=None, position=None, orientation=None, font_path=None, random_state=None):
h = int(360.0 * 100.0 / 255.0)
s = int(100.0 * 255.0 / 255.0)
l = int(100.0 * float(random_state.randint(60, 120)) / 255.0)
return "hsl({}, {}%, {}%)".format(h, s, l)
file_content=open ("filteredtext9.txt").read()
wordcloud = WordCloud(font_path = r'/Users/sofya/Library/Fonts/Verdana.ttf',
stopwords = STOPWORDS,
background_color = 'white',
width = 2000,
height = 1500,
color_func = random_color_func
).generate(file_content)
plt.imshow(wordcloud)
plt.axis('off')
plt.show()
plt.draw()
# +
import nltk
from nltk.tokenize import word_tokenize, sent_tokenize
file_docs = []
with open ('filteredtext7.txt') as f:
tokens = sent_tokenize(f.read())
for line in tokens:
file_docs.append(line)
print("Number of documents:",len(file_docs))
# +
# creating a variable and storing the text
# that we want to search
search_text = "autistici"
# creating a variable and storing the text
# that we want to add
replace_text = "autistico"
# Opening our text file in read only
# mode using the open() function
with open(r'filteredtext9.txt', 'r') as file:
# Reading the content of the file
# using the read() function and storing
# them in a new variable
data = file.read()
# Searching and replacing the text
# using the replace() function
data = data.replace(search_text, replace_text)
# Opening our text file in write only
# mode to write the replaced content
with open(r'filteredtext9.txt', 'w') as file:
# Writing the replaced data in our
# text file
file.write(data)
# Printing Text replaced
print("Text replaced")
# +
infile = "filteredtext6.txt"
outfile = "cleaned_file6.txt"
delete_list = ["ah", "sì", "ma"]
with open(infile) as fin, open(outfile, "w+") as fout:
for line in fin:
for word in delete_list:
line = line.replace(word, "")
fout.write(line)
# -
pip install pandas
# +
import matplotlib.pyplot as plt
words = [('creatività', 24), ('attività', 33), ('ragazzo', 30), ('fatto', 25), ('esempio', 34),
('stanza', 18), ('spazio', 18), ('domande', 17), ('penso', 30), ('musica', 25),
('lavorato', 18),('autistico', 23), ('sostegno', 13), ('bambino', 75),
('esperienza', 23), ('ambiente', 23), ('room', 22), ('scuola', 21), ('magic', 19)]
sizes, labels = [i[1] for i in words],[i[0] for i in words]
plt.pie(sizes, labels=labels,autopct='%1.1i%%')
plt.show()
# -
pip install gensim
# Importing the libraries
import nltk
from nltk.corpus import stopwords
print(stopwords.words('italian'))
import nltk
stopwords = nltk.corpus.stopwords.words('italian')
new_words=('va','Va', 'bene', 'no','a','così', 'Poi', 'Però','Che', 'È', 'E','ne','I', 'Allora', 'Quindi')
for i in new_words:
stopwords.append(i)
print(stopwords)
# +
import io
from nltk.corpus import stopwords
from nltk.tokenize import word_tokenize
# word_tokenize accepts
# a string as an input, not a file.
stop_words = set(stopwords.words('italian'))
file1 = open("filteredtext.txt")
# Use this to read file content as a stream:
line = file1.read()
words = line.split()
for r in words:
if not r in stop_words:
appendFile = open('filteredtext02.txt','a')
appendFile.write(" "+r)
appendFile.close()
# -
# example text
fname = input('Enter File: ')
if len(fname) < 1 : fname = 'Focus_Group_Transcribtion.txt'
hand = open(fname)
# removing stopwords
fname = " ".join([word for word in fname.split() if word not in stop_words])
print(fname)
# +
infile = "filteredtext6.txt"
outfile = "cleaned_file111.txt"
delete_list = ["ah"]
fin = open(infile)
fout = open(outfile, "w+")
for line in fin:
for word in delete_list:
line = line.replace(word, "")
fout.write(line)
fin.close()
fout.close()
# -
import nltk
from nltk.corpus import stopwords
print(stopwords.words('italian'))
import nltk
stopwords = nltk.corpus.stopwords.words('italian')
new_words=('va','Va', 'bene', 'no','a','così', 'Poi', 'Però','Che', 'È', 'E','ne','I', 'Allora', 'Quindi')
for i in new_words:
stopwords.append(i)
print(stopwords)
import io
from nltk.corpus import stopwords
from nltk.tokenize import word_tokenize
#word_tokenize accepts a string as an input, not a file.
stop_words = set(stopwords.words('italian'))
file1 = open("filteredtext6.txt")
line = file1.read()# Use this to read file content as a stream:
words = line.split()
for r in words:
if not r in stop_words:
appendFile = open('filteredtext.txt','a')
appendFile.write(" "+r)
appendFile.close()
| word_analisis_focus_group1.txt.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.4
# language: python
# name: python3.4
# ---
# Vanillin production
# ------------------
#
# In 2010, Brochado *et al* used heuristic optimization together with flux simulations to design a vanillin producing yeast strain.
#
# <NAME>., <NAME>., <NAME>., & <NAME>. (2012). Impact of stoichiometry representation on simulation of genotype-phenotype relationships in metabolic networks. PLoS Computational Biology, 8(11), e1002758. doi:10.1371/journal.pcbi.1002758
# Genome-scale metabolic model
# --------------------------
#
# In their work, the authors used *iFF708* model, but recent insights in Yeast yielded newer and more complete versions.
# Becuase this algorithms should be agnostic to the model, we implement the same strategy with a newer model.
from cameo import models
model = models.bigg.iMM904
# Constraints can be set in the model according to data found in the literature. The defined conditions allow the simulation of phenotypes very close to the experimental results.
#
# <img src=http://www.biomedcentral.com/content/figures/1752-0509-7-36-2.jpg/>
# **Model validation by comparing in silico prediction of the specific growth rate with experimental data**. Growth phenotypes were collected from literature and compared to simulated values for chemostat cultivations at four different conditions, nitrogen limited aerobic (green) and anaerobic (red), carbon limited aerobic (blue) and anaerobic (white).
#
#
# <NAME>., <NAME>., <NAME>., & <NAME>. (2013). Mapping condition-dependent regulation of metabolism in yeast through genome-scale modeling. BMC Systems Biology, 7, 36. doi:10.1186/1752-0509-7-36
model.reactions.EX_glc__D_e.lower_bound = -13 #glucose exchange
model.reactions.EX_o2_e.lower_bound = -3 #oxygen exchange
model.medium
model.objective = model.reactions.BIOMASS_SC5_notrace #growth
model.optimize().f
# Heterologous pathway
# -------------------
#
# Vanillin is not produced by *S. cervisiae*. In their work an heterolgous pathway is inserted to allow generate a vanillin production strain. The pathway is described as:
#
# <img src=http://static-content.springer.com/image/art%3A10.1186%2F1475-2859-9-84/MediaObjects/12934_2010_Article_474_Fig1_HTML.jpg>
# **Schematic representation of the de novo VG biosynthetic pathway in S. Cerevisisae** (as designed by Hansen et al [5]). Metabolites are shown in black, enzymes are shown in black and in italic, cofactors and additional precursors are shown in red. Reactions catalyzed by heterologously introduced enzymes are shown in red. Reactions converting glucose to aromatic amino acids are represented by dashed black arrows. Metabolite secretion is represented by solid black arrows where relative thickness corresponds to relative extracellular accumulation. 3-DSH stands for 3-dedhydroshikimate, PAC stands for protocathechuic acid, PAL stands for protocatechuic aldehyde, SAM stands for S-adenosylmethionine. 3DSD stands for 3-dedhydroshikimate dehydratase, ACAR stands for aryl carboxylic acid reductase, PPTase stands for phosphopantetheine transferase, hsOMT stands for O-methyltransferase, and UGT stands for UDP-glycosyltransferase. Adapted from Hansen et al. [5].
# Brochado et al. Microbial Cell Factories 2010 9:84 doi:10.1186/1475-2859-9-84
#
# Using **cameo**, is very easy to generate a pathway and add it to a model.
from cameo.strain_design.pathway_prediction import PathwayPredictor
predictor = PathwayPredictor(model)
pathways = predictor.run('vanillin', max_predictions=3)
vanillin_pathway = pathways.pathways[0]
from cameo.core.pathway import Pathway
vanillin_pathway = Pathway.from_file("data/vanillin_pathway.tsv")
vanillin_pathway.data_frame
# And now we can plug the pathway to the model.
vanillin_pathway.plug_model(model)
from cameo import phenotypic_phase_plane
# The Phenotypic phase plane can be used to analyse the theoretical yields at different growth rates.
production_envelope = phenotypic_phase_plane(model, variables=[model.reactions.BIOMASS_SC5_notrace],
objective=model.reactions.EX_vnl_b_glu_c)
production_envelope.plot()
production_envelope = phenotypic_phase_plane(model, variables=[model.reactions.BIOMASS_SC5_notrace],
objective=model.reactions.EX_vnl_b_glu_c)
production_envelope.plot()
# To find gene knockout targets, we use `cameo.strain_design.heuristic` package which implements the OptGene strategy.
#
# The authors used the biomass-product coupled yield (bpcy) for optimization which is the equivalent of running OptGene in non-robust mode. All simulations were computed using MOMA but because **cameo** does not implement MOMA we use it's equivalent linear version (it minimizes the absolute distance instead of the quadratic distance). The linear MOMA version is faster than the original MOMA formulation.
#
# By default, our OptGene implementation will run 20'000 evaluations.
from cameo.strain_design.heuristic.evolutionary_based import OptGene
from cameo.flux_analysis.simulation import lmoma
optgene = OptGene(model)
results = optgene.run(target="EX_vnl_b_glu_c",
biomass="BIOMASS_SC5_notrace",
substrate="EX_glc__D_e",
simulation_method=lmoma)
results
| Advanced-SynBio-for-Cell-Factories-Course/Vanillin Production.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # LANL Earthquake Prediction
#
# ### Written by Anand
# ------------
#
# ## Data Description
#
# The goal of this competition is to use acoustic seismic signals to predict the estimated time to laboratory earthquakes. The data comes from a well-known experimental set-up used to study earthquake physics. The `acoustic_data` input signal is used to predict the time remaining before the next laboratory earthquake (`time_to_failure`).
#
# The training data is a single, continuous segment of experimental data. The test data consists of a folder containing many small segments. The data within each test file is continuous, but the test files do not represent a continuous segment of the experiment; thus, the predictions cannot be assumed to follow the same regular pattern seen in the training file.
#
# For each `seg_id` in the test folder, you should predict a single `time_to_failure` corresponding to the time between the last row of the segment and the next laboratory earthquake
#
# ----------
# The train.csv file is a humongous file with 629,145,480 rows and 2 columns(indicating `acoustic_data` and`time_to_failure` respectively). The file itself takes 8.89GB of hard-disk space, so it is very important not to load to it entirely.
#
#
# The Dataset can be downloaded from - https://www.kaggle.com/c/LANL-Earthquake-Prediction/data
#
# +
import numpy as np
import pandas as pd
from time import time
import matplotlib
import seaborn as sns
import matplotlib.pyplot as plt
from livelossplot import PlotLosses
from tqdm import tqdm_notebook
import torch
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
import torch.autograd as autograd
from torch.autograd import Variable
from torch.utils.data import Dataset
from torch.utils.data import DataLoader
import sklearn
import lightgbm as lgb
from sklearn.externals import joblib
from sklearn.kernel_ridge import KernelRidge
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import GridSearchCV, KFold
# Custom DataProcessor
from dataprocess import DataProcessor, TestDataProcessor
sns.set_style('darkgrid')
# %matplotlib inline
# -
# Print version Info
print("Numpy version: ",np.__version__)
print("Matplolib version: ",matplotlib.__version__)
print("Pandas version: ",pd.__version__)
print("Seaborn version: ",sns.__version__)
print("LightGBM version: ",lgb.__version__)
print("PyTorch version: ",torch.__version__)
print("Sklearn version: ",sklearn.__version__)
# ## Data Pre-processing
# I have written a custom pre-processor to handle the data in batches and preprocess. Note that the data is **Huge** it is impractical to read all the data and load it unto the RAM. It is also a poor coding practice. This data processor never loads the entire raw data, and uses the iterator to read data in batches and process it. The `DataProcessor` also has some inbuilt plotting functions to visualize the data.
#
# First, let's start with plotting the raw data sample.
# +
# set random seed to 0
np.random.seed(0)
torch.manual_seed(0)
DATA_PATH = "D:/Jobs/Incubit/" # Path to the extracted data from the file "LANL-Earthquake-Prediction.zip"
TEST_DATA_PATH = "D:/Jobs/Incubit/test/"
SAVE_MODEL_PATH = "D:/Jobs/Incubit/Code/saved_models/" # Path to save the learned models
# +
dl = DataProcessor("train.csv", DATA_PATH, 120)
dl.plot_data_sample()
# -
# By zooming in the data, we see that the `time_to_failure`signal has some kind of step-like structure. This essentially indicates the resolution of the device measuring the time. Let's plot the resolution of the data. Also, note that there is a lag between the peaks in the acoustic data and the actualy failure time.
dl.plot_resolution()
# Clearly, the `time_to_failure` doesn't change continuously with the acoustic signal. Therefore, my first idea was to compress the signal within this resolution. We can assume that the acoustic signal has some Gaussian noise (usualy assumption) and replace the signal within this resolution time with some *sufficient statistic* without any loss of the data information. This is done by the `resolve_data` function for which two commonly used sufficient statistics can be used - *mean* or *z-score*.
print("Resolution in time_to_failure : {}".format(dl._find_resolution()))
dl.resolve_data(summary_stats = 'mean')
# Let's plot the data again!
dl.plot_data_sample()
dl.resolve_data_summary()
# Since, we have compressed the original data, the resolved_train data contains only 153,639 rows and 2 columns. This takes just 5.27MB!!! This can easily be loaded onto the memory without any problem. Furthermore, the peaks in the acoustic signal and the failure are almost at the same time. We shall use this dataset with a sliding window for our Sequence model using LSTM.
#
# ### Feature Extraction
# Another way to process the data is use simply use a batch of sufficient length and try to extract useful statistics like mean, standard deviation, quantiles, etc. This list of features can be huge and we can simply let the model to choose what features are important.
# Also note that the previous data pre-processing is essentially the same idea with batch length = 4096 and the statistic to be the mean/z-score.
#
# As such, we shall borrow the knowledge of feature extraction from other kagglers from other competitions. I have added few of the features from my knowledge - feaquency-based features (since, it is a signal) and norm-based features. I have grouped the features in the following categrories as follows -
#
# - **Moment features**- Mean, Standard Deviation, Skew and Kurtosis (just the first four moments of the data)
# - **Quantile features** - 1%, 5%, 95%,99% quantiles, median, minimum, F-test, average abolute change
# - **Frequencey features**- mean absolute DCT, max absolute DCT, min absolute DCT, 5%, 25%, 75%, 95% quantiles
# - **Norm feature**- max norm, 2nd norm, 3rd norm
#
# Furthermore, we also use a sliding window within each batch to capture the local relations. Again, we take the statistical features such as mean, quantiles etc.
features = ['moment','quantile','freq', 'norm', 'subwindow']
dl.extract_features(features, N = 150_000) # Batch size of 150,000
# Let's similarly extract the features for the test data
tl = TestDataProcessor(DATA_PATH + 'sample_submission.csv',TEST_DATA_PATH )
tl.resolve_data()
tl.extract_features(features)
X_t = tl.get_resolved_test_data()
X_t.shape
# +
X_tr, y_tr = dl.get_feature_data()
# Sanity check for Nans etc
X_tr.isnull().sum()
# +
#Train head
X_tr.head()
# +
X_ts = tl.get_test_feature_data()
#Sanity chgeck for test data
X_ts.isnull().sum()
# +
#Test head
X_ts.head()
# -
# ## Machine Learning Approach
#
# Now, with processed data and extracted features, it is time to run some models on the data and see their performance. First, I shall use kernel ridge regression, which is essentially a generalised linear regression but with some non-linear transformations of the data. However, fine-tuning the parameters are often difficult. Therefore, I shall use grid search to search for the optimal hyper parameters with K-fold cross validation. Another important thing to keep in mind is that, the evaluation metric used is `MAE` (Mean Absolute Error). Therefore, it is crucial that we optimize to reduce this error rather than some arbitrary metric.
# +
def grid_search_cv(model, grid, features, target,num_folds = 5):
"""
Search for the optimal hyperparameter-combination in the given grid.
"""
start_time = time()
reggressor = GridSearchCV(model, grid, cv=num_folds, scoring='neg_mean_absolute_error')
reggressor.fit(features, target)
end_time = time() - start_time
print("Best CV score: {:.3f}, time: {:.2f}s".format(-reggressor.best_score_, end_time))
print("Best hyperparameters :", reggressor.best_params_)
return reggressor.best_params_
def model_predict(model, features, target,num_folds = 5, test=None, plot=True, lgb=False):
"""
Train the estimator and make predictions for oof and test data.
"""
folds = KFold(num_folds, shuffle=True)
oof_predictions = np.zeros(features.shape[0])
if test is not None:
sub_predictions = np.zeros(test.shape[0])
for (train_index, valid_index) in folds.split(features, target):
if lgb:
model.fit(features[train_index], target[train_index],
early_stopping_rounds=100, verbose=False,
eval_set=[(features[train_index], target[train_index]),
(features[valid_index], target[valid_index])])
else:
model.fit(features[train_index], target[train_index])
oof_predictions[valid_index] = model.predict(features[valid_index]).flatten()
if test is not None:
sub_predictions += model.predict(test).flatten() / num_folds
# Plot out-of-fold predictions (oof) vs actual values (Code taken from a Kaggle kernel)
if plot:
fig, axis = plt.subplots(1, 2, figsize=(12,5))
ax1, ax2 = axis
ax1.set_xlabel('actual',fontsize=15)
ax1.set_ylabel('predicted',fontsize=15)
ax2.set_xlabel('train index',fontsize=15)
ax2.set_ylabel('time to failure',fontsize=15)
ax1.scatter(target, oof_predictions, color='brown')
ax1.plot([(0, 0), (20, 20)], [(0, 0), (20, 20)], color='blue')
ax2.plot(target, color='blue', label='Actual')
ax2.plot(oof_predictions, color='orange',label = 'Predictions')
plt.legend(fontsize=15)
if test is not None:
return oof_predictions, sub_predictions
else:
return oof_predictions, model
# +
# Kernel Ridge Regression
scaler = StandardScaler() # Z-score normalize the data features
X_train_scaled = scaler.fit_transform(X_tr)
target = y_tr.values.flatten()
num_folds = 5
# We essentially optimize over the gamma parameter of the rbf kernel and the scaling alpha
grid = [{'gamma': np.linspace(1e-8, 0.1, 10), 'alpha': [0.0005, 0.001, 0.02, 0.08, 0.1]}]
params = grid_search_cv(KernelRidge(kernel='rbf'), grid, X_train_scaled, target)
kr_oof, kridge_model = model_predict(KernelRidge(kernel='rbf', **params), X_train_scaled, target)
# Save trained model
joblib.dump(kridge_model, SAVE_MODEL_PATH+'kernelridge.pkl')
# -
# Predict the values on the test data
X_test_scaled = scaler.fit_transform(X_ts)
kridge_model.predict(X_test_scaled).flatten()
# Now, I shall test on the famous time-tested gradient-boosted decision trees. For this I shall use the lightGBM python library. Once again, I shall use gridsearch with k-fold cross validation to search for the optimal hyper-parameters for this model.
# +
fixed_params = {
'objective': 'regression_l1',
'boosting': 'gbdt', #'dart'
'verbosity': -1,
'random_seed': 489,
'num_boosting_round': 20000, # Num learners
'tree_learner':'data_parallel',
}
param_grid = {
'learning_rate': [0.1, 0.05, 0.01, 0.005],
'min_gain_to_split': [0, 0.001, 0.01, 0.1],
'lambda_l1': [0, 0.1, 0.2, 0.4, 0.6, 0.9], # L1 regularizer
'lambda_l2': [0, 0.1, 0.2, 0.4, 0.6, 0.9], # L2 regularizer
'num_leaves': list(range(16, 80, 4)),
'max_depth': [3, 4, 5, 6, 8, 12, 16, -1],
'feature_fraction': [0.8, 0.85, 0.9, 0.95, 1],
'min_data_in_leaf': [10, 20, 40, 60, 100],
'subsample': [0.8, 0.85, 0.9, 0.95, 1],
}
best_score = np.inf # Max-out the best score for comparison
dataset = lgb.Dataset(X_tr, label=y_tr)
print("Performing randomsearch CV on LightGBM Model...")
for i in tqdm_notebook(range(100)):
params = {k: np.random.choice(v) for k, v in param_grid.items()}
params.update(fixed_params)
result = lgb.cv(params, dataset, nfold=5, early_stopping_rounds=100, stratified=False)
if result['l1-mean'][-1] < best_score:
best_score = result['l1-mean'][-1]
best_params = params
best_nrounds = len(result['l1-mean'])
print("Best mean score: {:.4f}, num rounds: {}".format(best_score, best_nrounds))
print("Best hyperparameters : ",best_params)
gb_oof, lgbm_model = model_predict(lgb.LGBMRegressor(**best_params), X_tr.values, y_tr.values.flatten(), lgb=True)
lgbm_model.predict(X_test_scaled).flatten()
lgbm_model.booster_.save_model(SAVE_MODEL_PATH+'lgbm.pkl')
# +
# Load Model
lgbm_model = lgb.Booster(model_file=SAVE_MODEL_PATH+'lgbm.pkl')
#Predict the time to failure values on the test data
lgbm_model.predict(X_test_scaled).flatten()
# -
# ## Deep Learning Approach I - Neural Network Prediction
# The first approaxh was to use a simple fully-connected feedforward network as a regressor model over the extracted features. As such, I created a simple 3-layer network for this task. I didnt have to create a data loader since the dataset is small enough and pandas provide a nice litte iterator that can be used as a dataloader.
class NeuralNet(nn.Module):
def __init__(self, n_feature, n_hidden, n_output, p):
super(NeuralNet, self).__init__()
self.hidden_1 = nn.Linear(n_feature, 220)
self.bn1 = nn.BatchNorm1d(220)
self.hidden_2 = nn.Linear(220, 140)
self.bn2 = nn.BatchNorm1d(140)
self.hidden_3 = nn.Linear(140, 10)
self.bn3 = nn.BatchNorm1d(10)
#self.hidden_4 = nn.Linear(n_hidden, 40)
self.predict = nn.Linear(10, n_output)
self.dropout = nn.Dropout(p)
def forward(self, x):
x = torch.tanh(self.dropout(self.bn1(self.hidden_1(x))))
x = torch.tanh(self.bn2(self.hidden_2(x)))
x = F.relu(self.bn3(self.hidden_3(x)))
#x = F.relu(self.hidden_4(x))
x = self.predict(x)
return x
# +
batchsize = 30
X_tr, y_tr = dl.get_feature_data()
net = NeuralNet(n_feature=X_tr.shape[1], n_hidden=400, n_output=1, p = 0.7)
optimizer = torch.optim.SGD(net.parameters(), lr=0.2)
loss_func = torch.nn.L1Loss()
# +
liveloss = PlotLosses()
N_epoch = 20
net.train()
for epoch in range(N_epoch):
logs = {}
running_loss = 0.0
for i in range(X_tr.shape[0]//batchsize):
inds = np.random.choice(X_tr.shape[0], batchsize, replace=False)
x = torch.from_numpy(X_tr.iloc[inds,:].values).type('torch.FloatTensor')
y = torch.from_numpy(y_tr.iloc[inds,:].values).type('torch.FloatTensor')
prediction = net(x) # forward pass
#print(y)
loss = loss_func(prediction, y)
optimizer.zero_grad()
loss.backward() # backprop
optimizer.step()
running_loss = loss.detach()
epoch_loss = running_loss
prefix = ''
logs[prefix + 'log loss'] = epoch_loss.item()
liveloss.update(logs)
liveloss.draw()
torch.save(net.state_dict(), SAVE_MODEL_PATH+"NN_model.pth")
# -
net.eval()
X_test = torch.from_numpy(X_ts.values).type('torch.FloatTensor')
net(X_test)
# ## Deep Learning Approach II - Sequence Modeling
# Alternatively, I thought of using a simple LSTM model to learn the sequence from the pre-processed data. The input is the acoustic signal and the target the time_to_failure signal. For this, I used a simply LSTM-based squence model with 3 layers of LSTMs. A custom dataloader (simple enough) had to be written for the sequnce model so that instead of just batches of data, we get a batch of sliding-window data. Where the data slides through a fixed window length to predict the next term in the sequence.
# +
class Time_Series_Data(Dataset):
def __init__(self, train_x, train_y):
self.X = train_x
self.y = train_y
def __getitem__(self, item):
x_t = self.X[item]
y_t = self.y[item]
return x_t, y_t
def __len__(self):
return len(self.X)
# Sliding Window
def sliding_window_samples(X_train, y_train, lookBack = 36, RNN=True):
dataX, dataY = [], []
for i in range(len(X_train) - lookBack):
sample_X = X_train[i:(i + lookBack)]
sample_Y = y_train[i + lookBack]
dataX.append(sample_X)
dataY.append(sample_Y)
dataX = np.array(dataX)
dataY = np.array(dataY)
if not RNN:
dataX = np.reshape(dataX, (dataX.shape[0], dataX.shape[1]))
return dataX, dataY
# -
# Here, I shall use bidirectional LSTM with the intuition that information from the acoustic signal before and after a certain time prediction might have more information about the time value itself.
class SequenceModel(nn.Module):
def __init__(self, inputDim, hiddenNum, outputDim, layerNum, BiDir = False):
super(SequenceModel, self).__init__()
self.hiddenNum = hiddenNum
self.inputDim = inputDim
self.outputDim = outputDim
self.layerNum = layerNum
self.BiDir = BiDir
self.cell = nn.LSTM(input_size=self.inputDim, hidden_size=self.hiddenNum,
num_layers=self.layerNum, dropout=0.2,
batch_first=True, bidirectional=True)
"""
Using Bidirectional LSTM 'may' provide a better understanding of the acoustic signal?
As it moves forward and backward through time, it can capture the information that is
conditioned upon the future values.
"""
self.fc = nn.Linear(self.hiddenNum, self.outputDim)
def forward(self, x):
batchSize = x.size(0)
if self.BiDir:
h0 = Variable(torch.zeros(self.layerNum * 2, batchSize, self.hiddenNum)).cuda()
c0 = Variable(torch.zeros(self.layerNum * 2, batchSize, self.hiddenNum)).cuda()
else:
h0 = Variable(torch.zeros(self.layerNum * 1, batchSize, self.hiddenNum)).cuda()
c0 = Variable(torch.zeros(self.layerNum * 1, batchSize, self.hiddenNum)).cuda()
#print(x.size(), h0.size())
rnnOutput, hn = self.cell(x, (h0, c0))
out = rnnOutput.transpose(0,1).contiguous().view(-1, self.hiddenNum)
fcOutput = self.fc(out)
return fcOutput
# +
# Create data pipeline for the sequence model
data = dl.get_resolved_data()
Num_train = int(0.8*data.shape[0])
Num_test = data.shape[0] - Num_train
X_train = data.values[:Num_train,0]
y_train = data.values[:Num_train,1]
X_test = data.values[:Num_test,0]
y_test = data.values[:Num_test,1]
lag = 36
flag = False
trainX, trainY = sliding_window_samples(X_train, y_train, lag, RNN=True)
train_dataset = Time_Series_Data(trainX, trainY)
trainloader = DataLoader(train_dataset, batch_size=32, shuffle=True, sampler=None, batch_sampler=None, num_workers=0)
testX, testY = sliding_window_samples(X_test, y_test, lag, RNN=True)
test_dataset = Time_Series_Data(testX, testY)
testloader = DataLoader(test_dataset, batch_size=32, shuffle=True, sampler=None, batch_sampler=None, num_workers=0)
# DO NOT CHANGE THE num_workes PARAMETER!!. PyTorch Doesn't support Windows fully yet!!
# +
checkPoint= 500
epoch = 4
hidden_num=64
input_dim = 1
seq_model = SequenceModel(inputDim=1, hiddenNum=hidden_num, outputDim=1, layerNum=3, BiDir = True).cuda()
optimizer = optim.SGD(seq_model.parameters(), lr=0.1, momentum=0.9)
criterion = nn.MSELoss()
train_loss_list, test_loss_list = [],[]
liveloss = PlotLosses()
start_time = time()
for i in range(epoch):
for batch_idx, (x, y) in enumerate(trainloader):
seq_model = seq_model.train()
x= x.permute(1, 0).unsqueeze(2).float()
x, y = Variable(x).cuda(), Variable(y.float()).cuda()
optimizer.zero_grad()
pred = seq_model(x)
loss = criterion(pred, y)
if batch_idx % checkPoint == 0 and batch_idx != 0:
end_time = time() - start_time
with torch.no_grad():
for x, y in testloader:
x= x.permute(1, 0).unsqueeze(2).float()
x, y = Variable(x).cuda(), Variable(y.float()).cuda()
pred = seq_model(x)
test_loss = criterion(pred, y)
print("Epoch [{:3d}/{:3d}] Batch [{:4d}/{:4d}] [Train Loss: {:2.3f}] [Test Loss {:2.3f}] Time: {:.3f}s" .format(
i, epoch, batch_idx, len(trainloader), loss.item(),test_loss.item(), end_time))
train_loss_list.append(loss.item())
test_loss_list.append(test_loss.item())
# logs[prefix + 'log loss'] = loss.item()
# liveloss.update(logs)
# liveloss.draw()
start_time = time()
loss.backward()
optimizer.step()
print("-------------------------")
# torch.save(train_loss_list, "LSTM_train_loss.pth")
# torch.save(test_loss_list, "LSTM_test_loss.pth")
torch.save(seq_model.state_dict(), SAVE_MODEL_PATH+"Seq_model.pth")
# -
| Main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Summary of the BatchNorm paper
# > Normalizing Neural Networks to allow for better performance and faster convergence
#
# - toc: true
# - badges: true
# - comments: true
# - categories: [jupyter]
# - image: images/BN.png
# ## Summary of [BatchNorm](https://arxiv.org/pdf/1502.03167.pdf)
# ### What did the authors want to achieve ?
# - make normalization a part of the model
# - allow the use of higher learning rates by ensuring a stable distribution of nonlinear inputs => faster training, less iterations needed
# - improve robustness to initialization (more independent of good init) : reduce dependence of gradients on parameter scale and of the initial values
# - normalize the activations and preserve information in the network
#
#
#
# ### Key elements
#
# #### Old approaches
# - whitening (linearly transforming inputs to have zero mean and unit variance and beingdecorrelated), has several problems. If the whitening modifiactions are interspersed with the optimization technique, gradient descent might try to update the parameters in a way that needs the normalization to be updated as well. This greatly reduces the effect of the backward pass step. In the paper this is shown by using considering a layer and normalizing the result with the mean of the training data. (see picture above) The authors show that the bias b will grow indefinitely while the loss remains the same. This was also observed in experiments, where the model blew up when the normalization parameters where computed outside of the backward pass. This is due to that approach not considering that during gradient descent, the normalization is taking place.
#
# #### Batch Norm
# - the idea is to normalize the activations during training, by normalizing the training samples (batches), relative to the statistics of the entire train set
# - as normalization may change what the layer already represents (Sigmoid normalization would constrain it to the linear part in between the saturation), the inserted transformation needs to be able to represent an identity tansformation. This is done by introducing two new learnable parameters for each batch for scaling and shifting the normalized value :
#
# 
#
#
#
# With $\gamma ^{k} = \sqrt{Var[x^{k}]}$ and $\beta ^{k} = E[x^{k}]$, the original activation can be restored
#
# - for each mini-batch mean and covariance is computed seperately, therefore the name Batch Normalization, the small parameter eta is used in order to avoid division by zero, when the standard deviation is 0 (this could happen in case of bad init for example) :
# 
#
#
# - BN can be applied to every activation (at least in feedforward networks and as long as there is a high enough batch size), as BN is differentiable, the chain rule can be used to consider the BN transformation :
#
# 
#
#
# - During training the following pseudocode applies :
# 
#
#
# - During testing a running moving average of mean and variance is used (linear transform), as the normalization based on a mini-batch is not desirable
#
# - Batch Norm prevents small changes of parameters to amplify larger changes in our network. Higher learning rates also don't influence the scale of the parameters during backprop, therefore amplification is prevented as the layer Jacobian is unaffected. The singular values of the Jacobian are also close to 1, which helps preserve gradient magnitudes. Even though the transformation is not linear and the normalizations are not guaranteed to be Gaussian or independent, BN is still expected to improve gradient characterisitcs.
# #### Implementation
# Batch Norm can be implemented as follows in PyTorch :
# Also check out [my summary of the Batch Norm part](https://cedric-perauer.github.io/DL_from_Foundations/jupyter/2020/04/12/Batchnorm.html) of the DL course by fastai for more normalization techniques such as running batch norm, layer and group norm, and a small Residual Net with Batch Norm. This is the same as the torch.nn module would do it, but it's always great to see it from scratch.
#collapse_show
class BatchNorm(nn.Module):
def __init__(self, nf, mom=0.1, eps=1e-5):
super().__init__()
# NB: pytorch bn mom is opposite of what you'd expect
self.mom,self.eps = mom,eps
self.mults = nn.Parameter(torch.ones (nf,1,1))
self.adds = nn.Parameter(torch.zeros(nf,1,1))
self.register_buffer('vars', torch.ones(1,nf,1,1))
self.register_buffer('means', torch.zeros(1,nf,1,1))
def update_stats(self, x):
#we average over all batches (0) and over x,y(2,3) coordinates (each filter)
#keepdim=True means we can still broadcast nicely as these dimensions will be left empty
m = x.mean((0,2,3), keepdim=True)
v = x.var ((0,2,3), keepdim=True)
self.means.lerp_(m, self.mom)
self.vars.lerp_ (v, self.mom)
return m,v
def forward(self, x):
if self.training:
with torch.no_grad(): m,v = self.update_stats(x)
else: m,v = self.means,self.vars
x = (x-m) / (v+self.eps).sqrt()
return x*self.mults + self.adds
# #### Results and Conclusion
# 
#
# - Batch Norm allows to use only 7% of the training steps to match previous state of the art models on ImageNet without it
# - Batch Norm Inception beats the state of the art on the ImageNet challenge
# - Batch Norm reduces the need for Dropput greatly as claimed by the authors, however it was still used with the traditional dropout set up used by the Inception architects
| _notebooks/2020-04-14-Batch-Norm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Low-Pass Filter
# *Modeling and Simulation in Python*
#
# Copyright 2021 <NAME>
#
# License: [Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International](https://creativecommons.org/licenses/by-nc-sa/4.0/)
# + tags=[]
# install Pint if necessary
try:
import pint
except ImportError:
# !pip install pint
# + tags=[]
# download modsim.py if necessary
from os.path import exists
filename = 'modsim.py'
if not exists(filename):
from urllib.request import urlretrieve
url = 'https://raw.githubusercontent.com/AllenDowney/ModSim/main/'
local, _ = urlretrieve(url+filename, filename)
print('Downloaded ' + local)
# + tags=[]
# import functions from modsim
from modsim import *
# -
# The following circuit diagram (from [Wikipedia](https://en.wikipedia.org/wiki/File:RC_Divider.svg)) shows a low-pass filter built with one resistor and one capacitor.
#
# 
#
# A "filter" is a circuit takes a signal, $V_{in}$, as input and produces a signal, $V_{out}$, as output. In this context, a "signal" is a voltage that changes over time.
#
# A filter is "low-pass" if it allows low-frequency signals to pass from $V_{in}$ to $V_{out}$ unchanged, but it reduces the amplitude of high-frequency signals.
#
# By applying the laws of circuit analysis, we can derive a differential equation that describes the behavior of this system. By solving the differential equation, we can predict the effect of this circuit on any input signal.
#
# Suppose we are given $V_{in}$ and $V_{out}$ at a particular instant in time. By Ohm's law, which is a simple model of the behavior of resistors, the instantaneous current through the resistor is:
#
# $ I_R = (V_{in} - V_{out}) / R $
#
# where $R$ is resistance in ohms.
#
# Assuming that no current flows through the output of the circuit, Kirchhoff's current law implies that the current through the capacitor is:
#
# $ I_C = I_R $
#
# According to a simple model of the behavior of capacitors, current through the capacitor causes a change in the voltage across the capacitor:
#
# $ I_C = C \frac{d V_{out}}{dt} $
#
# where $C$ is capacitance in farads (F).
#
# Combining these equations yields a differential equation for $V_{out}$:
#
# $ \frac{d }{dt} V_{out} = \frac{V_{in} - V_{out}}{R C} $
#
# Follow the instructions below to simulate the low-pass filter for input signals like this:
#
# $ V_{in}(t) = A \cos (2 \pi f t) $
#
# where $A$ is the amplitude of the input signal, say 5 V, and $f$ is the frequency of the signal in Hz.
# ## Params and System objects
#
# Here's a `Params` object to contain the quantities we need. I've chosen values for `R1` and `C1` that might be typical for a circuit that works with audio signal.
params = Params(
R1 = 1e6, # * ohm
C1 = 1e-9, # * farad
A = 5, # * volt
f = 1000, # * Hz
)
params
# Now we can pass the `Params` object `make_system` which computes some additional parameters and defines `init`.
#
# * `omega` is the frequency of the input signal in radians/second.
#
# * `tau` is the time constant for this circuit, which is the time it takes to get from an initial startup phase to
#
# * `cutoff` is the cutoff frequency for this circuit (in Hz), which marks the transition from low frequency signals, which pass through the filter unchanged, to high frequency signals, which are attenuated.
#
# * `t_end` is chosen so we run the simulation for 4 cycles of the input signal.
# +
from numpy import pi
def make_system(params):
"""Makes a System object for the given conditions.
params: Params object
returns: System object
"""
f, R1, C1 = params.f, params.R1, params.C1
init = State(V_out = 0)
omega = 2 * pi * f
tau = R1 * C1
cutoff = 1 / R1 / C1 / 2 / pi
t_end = 4 / f
return System(params,
init=init,
t_end=t_end, num=401,
omega=omega, tau=tau,
cutoff=cutoff)
# -
# Let's make a `System`
system = make_system(params)
system
# The system variable `num` controls how many time steps we get from `run_solve_ivp`. The default is 101; in this case we increase it to 401 because the methods we'll use to analyze the results require high resolution in time.
# **Exercise:** Write a slope function that takes as an input a `State` object that contains `V_out`, and returns the derivative of `V_out`.
#
# +
# Solution goes here
# -
# Test the slope function with the initial conditions.
slope_func(0, system.init, system)
# And then run the simulation. I suggest using `t_eval=ts` to make sure we have enough data points to plot and analyze the results.
results, details = run_solve_ivp(system, slope_func)
details.message
results.tail()
# Here's a function you can use to plot `V_out` as a function of time.
# +
def plot_results(results):
V_out = results.V_out.copy()
t_end = results.index[-1]
if t_end < 0.1:
V_out.index *= 1000
xlabel = 'Time (ms)'
else:
V_out = results.V_out
xlabel = 'Time (s)'
V_out.plot(label='_nolegend')
decorate(xlabel=xlabel,
ylabel='$V_{out}$ (volt)')
plot_results(results)
# -
# If things have gone according to plan, the amplitude of the output signal should be about 0.8 V.
#
# Also, you might notice that it takes a few cycles for the signal to get to the full amplitude.
# ## Sweeping frequency
#
# Here's what `V_out` looks like for a range of frequencies:
# +
from matplotlib.pyplot import subplot
fs = [1, 10, 100, 1000, 10000, 100000]
for i, f in enumerate(fs):
system = make_system(params.set(f=f))
results, details = run_solve_ivp(system, slope_func)
subplot(3, 2, i+1)
plot_results(results)
# -
# At low frequencies, notice that there is an initial "transient" before the output gets to a steady-state sinusoidal output. The duration of this transient is a small multiple of the time constant, `tau`, which is 1 ms.
# ## Estimating the output ratio
#
# Let's compare the amplitudes of the input and output signals. Below the cutoff frequency, we expect them to be about the same. Above the cutoff, we expect the amplitude of the output signal to be smaller.
#
# We'll start with a signal at the cutoff frequency, `f=1000` Hz.
system = make_system(params.set(f=1000))
results, details = run_solve_ivp(system, slope_func)
V_out = results.V_out
plot_results(results)
# The following function computes `V_in` as a `TimeSeries`:
def compute_vin(results, system):
"""Computes V_in as a TimeSeries.
results: TimeFrame with simulation results
system: System object with A and omega
returns: TimeSeries
"""
A, omega = system.A, system.omega
ts = results.index
V_in = A * np.cos(omega * ts)
return TimeSeries(V_in, results.index, name='V_in')
# Here's what the input and output look like. Notice that the output is not just smaller; it is also "out of phase"; that is, the peaks of the output are shifted to the right, relative to the peaks of the input.
# +
V_in = compute_vin(results, system)
V_out.plot()
V_in.plot()
decorate(xlabel='Time (s)',
ylabel='V (volt)')
# -
# The following function estimates the amplitude of a signal by computing half the distance between the min and max.
def estimate_A(series):
"""Estimate amplitude.
series: TimeSeries
returns: amplitude in volts
"""
return (series.max() - series.min()) / 2
# The amplitude of `V_in` should be near 5 (but not exact because we evaluated it at a finite number of points).
A_in = estimate_A(V_in)
A_in
# The amplitude of `V_out` should be lower.
A_out = estimate_A(V_out)
A_out
# And here's the ratio between them.
ratio = A_out / A_in
ratio
# **Exercise:** Encapsulate the code we have so far in a function that takes two `TimeSeries` objects and returns the ratio between their amplitudes.
# +
# Solution goes here
# -
# And test your function.
estimate_ratio(V_out, V_in)
# ## Estimating phase offset
#
# The delay between the peak of the input and the peak of the output is call a "phase shift" or "phase offset", usually measured in fractions of a cycle, degrees, or radians.
#
# To estimate the phase offset between two signals, we can use cross-correlation. Here's what the cross-correlation looks like between `V_out` and `V_in`:
corr = np.correlate(V_out, V_in, mode='same')
corr_series = make_series(V_in.index, corr)
corr_series.plot(color='C4')
decorate(xlabel='Lag (s)',
ylabel='Correlation')
# The location of the peak in the cross correlation is the estimated shift between the two signals, in seconds.
peak_time = corr_series.idxmax()
peak_time
# We can express the phase offset as a multiple of the period of the input signal:
period = 1 / system.f
period
peak_time / period
# We don't care about whole period offsets, only the fractional part, which we can get using `modf`:
frac, whole = np.modf(peak_time / period)
frac
# Finally, we can convert from a fraction of a cycle to degrees:
frac * 360
# **Exercise:** Encapsulate this code in a function that takes two `TimeSeries` objects and a `System` object, and returns the phase offset in degrees.
#
# Note: by convention, if the output is shifted to the right, the phase offset is negative.
# +
# Solution goes here
# -
# Test your function.
estimate_offset(V_out, V_in, system)
# ## Sweeping frequency again
#
# **Exercise:** Write a function that takes as parameters an array of input frequencies and a `Params` object.
#
# For each input frequency it should run a simulation and use the results to estimate the output ratio (dimensionless) and phase offset (in degrees).
#
# It should return two `SweepSeries` objects, one for the ratios and one for the offsets.
# +
# Solution goes here
# -
# Run your function with these frequencies.
fs = 10 ** linspace(0, 4, 9)
ratios, offsets = sweep_frequency(fs, params)
# We can plot output ratios like this:
ratios.plot(color='C2', label='output ratio')
decorate(xlabel='Frequency (Hz)',
ylabel='$V_{out} / V_{in}$')
# But it is useful and conventional to plot ratios on a log-log scale. The vertical gray line shows the cutoff frequency.
def plot_ratios(ratios, system):
"""Plot output ratios.
"""
# axvline can't handle a Quantity with units
cutoff = magnitude(system.cutoff)
plt.axvline(cutoff, color='gray', alpha=0.4)
ratios.plot(color='C2', label='output ratio')
decorate(xlabel='Frequency (Hz)',
ylabel='$V_{out} / V_{in}$',
xscale='log', yscale='log')
plot_ratios(ratios, system)
# This plot shows the cutoff behavior more clearly. Below the cutoff, the output ratio is close to 1. Above the cutoff, it drops off linearly, on a log scale, which indicates that output ratios for high frequencies are practically 0.
#
# Here's the plot for phase offset, on a log-x scale:
def plot_offsets(offsets, system):
"""Plot phase offsets.
"""
# axvline can't handle a Quantity with units
cutoff = magnitude(system.cutoff)
plt.axvline(cutoff, color='gray', alpha=0.4)
offsets.plot(color='C9', label='phase offset')
decorate(xlabel='Frequency (Hz)',
ylabel='Phase offset (degree)',
xscale='log')
plot_offsets(offsets, system)
# For low frequencies, the phase offset is near 0. For high frequencies, it approaches 90 degrees.
# ### Analysis
#
# By analysis we can show that the output ratio for this signal is
#
# $A = \frac{1}{\sqrt{1 + (R C \omega)^2}}$
#
# where $\omega = 2 \pi f$, and the phase offset is
#
# $ \phi = \arctan (- R C \omega)$
#
# **Exercise:** Write functions that take an array of input frequencies and returns $A(f)$ and $\phi(f)$ as `SweepSeries` objects. Plot these objects and compare them with the results from the previous section.
#
# +
# Solution goes here
# -
# Test your function:
A = output_ratios(fs, system)
# +
# Solution goes here
# -
# Test your function:
phi = phase_offsets(fs, system)
# Plot the theoretical results along with the simulation results and see if they agree.
A.plot(style=':', color='gray', label='analysis')
plot_ratios(ratios, system)
phi.plot(style=':', color='gray', label='analysis')
plot_offsets(offsets, system)
# For the phase offsets, there are differences between the theoretical results and our estimates, but that is probably because it is not easy to estimate phase offsets precisely from numerical results.
# **Exercise:** Consider modifying this notebook to model a [first order high-pass filter](https://en.wikipedia.org/wiki/High-pass_filter#First-order_continuous-time_implementation), a [two-stage second-order low-pass filter](https://www.electronics-tutorials.ws/filter/filter_2.html), or a [passive band-pass filter](https://www.electronics-tutorials.ws/filter/filter_4.html).
| examples/filter.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .sh
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Bash
# language: bash
# name: bash
# ---
# <a href="https://cloudevel.com"> <img src="img/cloudevel.png" width="500px"></a>
# # Gestión de repositorios.
# ## Estructura de un repositorio de *Git*.
#
# Un repositorio de *Git* es un directorio el cual a su vez contiene un subdirectorio ```.git``` con una estructura similar a la siguiente:
# ``` bash
# .git
# ├── config
# ├── description
# ├── FETCH_HEAD
# ├── HEAD
# ├── hooks
# ├── index
# ├── info
# ├── logs
# │ ├── HEAD
# │ └── refs
# ├── objects
# │ ├── info
# │ └── pack
# ├── ORIG_HEAD
# └── refs
# ├── heads
# ├── remotes
# └── tags
# ```
# ## Incialización de un repositorio.
#
# El comando ```git init``` permite crear la estructura básica de un subdirectorio ```.git``` en un directorio dado mediante la siguiente sintaxis:
#
#
# ``` bash
# git init <ruta>
# ```
#
# Donde:
#
# * ```<ruta>``` es la ruta del directorio a inicializar. Si no se indica la ruta, el comando creará el directorio ```.git``` en el directorio desde el cual se ejecuta el comando.
#
# Para mayor información sobre ```git init``` es posible consultar la siguiente liga:
#
# https://git-scm.com/docs/git-init
# **Ejemplo:**
# * La siguiente celda creará el directorio ```demo```.
mkdir demo
tree demo
# * La siguiente celda inicializará al directorio ```demo```como un repositorio de *Git*.
git init demo
# * La siguiente celda ejecutará el comando ```tree``` para el subdirectorio ```demo/.git```, mostrando su estructura.
tree demo/.git
# ## Estado de un repositorio.
#
# El estado de un respositorio corresponde al estado de todos los archivos a los que *Git* está dando seguimiento dentro de dicho repositorio.
#
# La característica principal de un gestor de versiones es la de poder conservar un registro detallado del estado de un repositorio en un momento específico.
#
# ### Los "commits".
#
# Guardar el estado de un repositorio de forma continua como si se tratara de una película resulta ser poco eficiente y costoso en recursos de almacenamiento. Es por ello que los gestores de verisones como *Git* han optado por conservar el estado de un repositorio sólo cuando el usuario decide asentarlos. A dicha accion se le conoce como hacer un "commit".
#
# Los "commits" son la base del control de versiones.
# ### Despliegue del estado de un repositorio.
#
# El comando ```git status``` permite conocer el estado de un repositorio mediante la siguiente sintaxis:
#
# ``` bash
# git status <ruta>
# ```
#
# Donde:
#
# * ```<ruta>``` es la ruta del directorio a consultar. Si no se indica la ruta, el comando consultará el estado del directorio desde el cual se ejecuta el comando.
#
# En caso de que la ruta no corresponda a un directorio que contenga una estructura de directorio ```.git``` adecuada, se generará unmensaje de error.
#
# Para mayor información sobre ```git status``` es posible consultar la siguiente liga:
#
# https://git-scm.com/docs/git-status
# **Ejemplos:**
# * La siguiente celda mostrará el estado del directorio ```demo```.
git status demo
# * La siguiente celda mostrará el estado del directorio en el que se encuentra esta notebook.
git status
# ## Rastreo de los archivos dentro de un repositorio.
#
# *Git* tiene la capacidad de dar seguimiento a los cambios de cada archivo dentro del repositorio, identificando a cada uno de ellos como un objeto.
#
# Para que *Git* pueda dar seguimiento a un objeto, es necesario darlo de alta en su índice. Una vez que un archivo es dado de alta, *Git* puede evaluar si dicho archivo ha sido modificado.
# ### Área de preparación (staging area).
#
# *Git* permite definir puntualmente aquellos archivos cuyos cambios serán asentados en un commit.
#
# El área de preparación (staging area) permite al usuario registrar aquellos cambios que serán incluidos en el commit.
# ### Estado de un archivo.
#
# * **Sin seguimiento** (untracked), lo que implica que el archivo no está en el índice del repositorio y por lo tanto sus cambios no serán rastreados por *Git*.
# * **En seguimiento** (tracked) lo que implica que el archivo está en el índice del repositorio.
# * **Modificado** (modified) lo que implica que un archivo ha sido modificado, eliminado o ha cambiado de nombre.
# * **No modificado** (unmodified).
#
# El comando ```git status``` da un resumen de:
# * Aquellos archivos creados después del último commit y que no tienen seguimiento.
# * Aquellos archivos que están en seguimiento y que fueron modificados después del último commit.
# ### Seguimiento y rastreo de archivos.
#
# El comando```git add``` es el encargado tanto de dar seguimiento a un archivo como de rastrear y registrar los cambios en el área de preparación. Su sintaxis es la siguiente:
#
#
# ```
# git add <patrón o ruta> <opciones>
# ```
# Donde:
#
# * ```<patrón o ruta>``` puede ser la ruta a un archivo o directorio específico o un patrón que identifique a más de un archivo.
# * ```<opciones>``` este comando cuenta con varias opciones útiles.
#
# En caso de que el archivo no haya estado en seguimiento, este y su contenido serán indexados. En caso de que el archivo ya se encuentre en seguimiento, se actualizará el registro de sus modificaciones.
#
# https://git-scm.com/docs/git-add
# #### La opción ```--all```.
#
# Esta opción aplicará la ejecuciónde ```git add``` a todos los archivos del repositorio.
# **Ejemplo:**
# * La siguiente celda moverá el shell al directorio ```demo```.
cd demo
# * La siguiente celda creará a los archivos ```archivo-1```, ```archivo-2``` y ```archivo-3```.
touch archivo-1 archivo-2 archivo-3
ls -a
# * En este momento los archivos recién creados no están en seguimiento.
git status
# * La siguiente celda indexará a ```archivo-1``` y lo registrará en el área de preparación.
git add archivo-1
git status
# * La siguiente celda indexará al resto de los archivos en el repositorio y los registrará en el área de preparación.
git add --all
git status
# ## Listado de objetos en el índice del repositorio.
#
# Para saber cuáles son los objetoss indexados en un repositorio se utiliza el comando ```git ls-files``` con la siguiente sintaxis:
#
# ``` bash
# git ls-files
# ```
#
# La documentación de referencia de ```git ls-files``` está disponible en:
#
# https://git-scm.com/docs/git-ls-files
# **Ejemplo:**
# La siguiente celda desplegará el listado de archivos indexados del repositorio.
git ls-files
git ls-files -s
# ## El archivo ```.gitignore```.
#
# Es común que un desarrollador quiera evitar indexar en el repositorio archivos tales como:
#
# * Archivos temporales.
# * Credenciales y contraseñas.
# * Bibliotecas que resulten redundantes.
# * Datos de prueba.
#
# *Git* permite definir una "lista negra" de archivos que serán ignorados por los comandos de ```git``` de forma automática. Esta lista negra se define mediante un archivo de texto llamado ```.gitignore```, el cual va en el directorio principal del repositorio.
#
# Este archivo contiene un listado en forma de columma ya sea de los nombres de los archivos o del patrón correspondiente a ciertos archivos que deberán ser ignorados.
#
#
# La documentación de referencia de ```.gitignore``` está disponible en:
#
# https://git-scm.com/docs/gitignore
# **Ejemplo:**
# * La siguiente celda creará al archivo ```.gitignore``` listando al nombre ```invisible```.
echo "invisible" > .gitignore
cat .gitignore
# * La siguiente celda creará al archivo ```invisible```.
touch invisible
# * El sistema de archivos del repositorio tiene los archivos.
# * ```archivo-1```
# * ```archivo-2```
# * ```archivo-3```
# * ```.gitignore```
# * ```invisible```
ls -a
# * El comando ```git status``` identificará al archivo ```.gitignore```, pero no a ```invisible```.
git status
# * Lo mismo ocurre con ```git add```.
git add --all
git status
# ## Confirmando los cambios en un repositorio ("commit").
#
# Una vez que el usuario realiza todas las operaciones necesarias en el área de preparación (staging area) está listo para hacer un commit.
#
# **Nota:** Para poder realizar un commit es necesario que el usuario defina al menos los campos ```user.name``` y ```user.email``` en la configuración del ámbito ```--global```.
# ### Flujo de un commit.
#
# * Cada objeto en el área de preparación es evaluado.
# * En caso de que sea un nuevo objeto, este será incluído en el estado del nuevo commit.
# * En caso de que el objeto ya esté registrado en un commit previo y haya sido modificado, se enumerarán y registrarán las modificaciones con respecto a la versión previa.
# * Todos los cambios serán registrados.
# * Se registrarán los cambios en el estado del repostorio, así como el momento exacto del commit.
# * Al commit se le asignará un identificador único que corresponde a un número hexadecimal.
# * Se vaciará el área de preparación.
# * Se desplegará un resumen del commit.
# ### El comando ```git commit```.
#
# El comando ```git commit``` es el encargado de confirmar los cambios en el repositorio. Cada commit requiere que se haga un comentario descriptivo de las modificaciones.
#
# La sintaxis es:
#
# ``` bash
# git commit <opciones> <archivos>
# ```
#
# Donde:
#
# * ```<archivos>``` permite definir un listado de archivos que serán tomados en commit de forma similar a ```git add```.
# * ```<opciones>``` las cuales definen diversas acciones para el commit.
#
# En caso de que se ejecute el comando ```git commit``` sin definir archivos u opciones, se abrirá un editor de textos (*vim*) en el cual se deberá de escribir un mensaje descriptivo del commit y una vez guardado, se realizará el commit tomando en cuanta todos los objetos definidos en el área de preparación.
#
# La documentación de referencia de ```git commit``` está disponible en:
#
# https://git-scm.com/docs/git-commit
#
#
# **Nota:** En caso de que no existan cambios en los objetos del área de preparacion, no se hará el commit.
# #### La opción ```--all``` o ```-a```.
#
# Esta opción busca y registra cambios en todos los objetos indexados del repositorio y los guarda en el área de preparación antes de hacer el commit. La opción ```-a``` no tomará en cuenta a archivos recién creados y que no hayan sido indexados.
#
# La sintaxis puede ser:
#
# ``` bash
# git commit -a
# ```
# o
# ``` bash
# git commit --all
# ```
# #### La opción ```--message``` o ```-m``` .
#
# Esta opción permite añadir un mensaje descriptivo y evita que se abra el editor de texto.
#
# La sintaxis puede ser:
#
# ```
# git commit -m <mensaje>
# ```
# o
# ```
# git commit --message=<mensaje>
# ```
#
# Donde:
#
# * ```<mensaje>``` es una cadena de caracteres que en caso de incliur espacios,d ebe de estar encerrada entre comillas.
# **Ejemplo:**
# La siguiente celda realizará un commit del repositorio actual añadiendo el mensaje descriptivo ```"primer commit ```.
git commit -m "primer commit"
# <img src="img/02/primer_commit.svg" width="100px">
git status
# ### Estructura del mensaje del commit.
#
# Una vez hecho el commit, *Git* regesa un mensaje como el siguiente:
#
# ````bash
# [master (root-commit) 3c76e61] primer commit
# 4 files changed, 1 insertion(+)
# create mode 100644 .gitignore
# create mode 100644 archivo-1
# create mode 100644 archivo-2
# create mode 100644 archivo-3
# ```
#
# Este mensaje se compone de:
# * Un encabezado que describe al commit.
# * Un listado de las adiciones o modificaciones de los objetos en el espacio de preparación.
# #### El encabezado de un mensaje de commit.
#
# El encabezado del mensaje de commit tiene una estructura como la siguiente:
#
# ``` bash
# [<rama> (<tipo de commit>) <identificador corto>] <mensaje>
# ```
#
# Donde:
#
# * ```<rama>``` es la rama del repositorio en la que se realzia el commit. La rama que se crra pord efecto en un repositorio es ```master```.
# * ```<tipo de commit>``` este dato es opcional y cuando se realiza el primer commti de un repoitorio despliega el mensaje ```root-commit````.
# * ```<identificador corto>```, el cual es un número hexadecimal de 7 dígitos que corresponde a los primeros 7 dígitos del número identificador completo del commit.
# ## Eliminación de un objeto.
#
# El comando ```git rm``` se utiliza para eliminar a un objeto tanto del repositorio como de su índice.
#
# ``` bash
# git rm <archivo> <opciones>
# ```
# **Ejemplo:**
# * El archivo ```archivo-3``` existe tanto en el respositorio como en el índice.
ls -a
git ls-files
# * A continuación se utilizará el comando ```git rm``` para eliminar al archivo tanto del repositorio como de su índice.
git rm archivo-3
ls -a
git ls-files
git status
# * Se hará un nuevo commit con la descripción ```segundo commit```.
git commit -m "segundo commit"
# <img src="img/02/segundo_commit.svg" width="250px">
# ### La opciones ```--cached``` y ```-f```.
#
# Es posible que existan archivos registrados en el área de preparación y que deban de ser eliminados antes de realizar un commit.
#
# La opción ```--cached``` del comando ```git rm``` eliminará al objeto del índice, pero no al archivo en el repositorio.
#
# La opción ```-f``` del comando ```git rm``` eliminará al objeto del índice y al archivo en el repositorio.
# **Ejemplo:**
# * Se creará al archivo ```archivo-4``` y se registrará en el área de preparación.
touch archivo-4
git add archivo-4
git status
# * Debido a que no se ha hecho un ```commit``` para ```archivo-4```, el comando ```git rm archivo-4``` no funcionará.
git rm archivo-4
# * Utilizando la opción ```--cached```, se eliminará al objeto del índice, pero se preservará en el directorio.
git rm archivo-4 --cached
ls -a
git status
# * Se hará un nuevo commit con la descripción ```tercer commit```. En este caso, como no hay cambios, el commit no se realizará.
git commit -m "tercer commit"
# <img src="img/02/segundo_commit.svg" width="250px">
# Se enviará nuevamente al archivo ```archivo-4``` al área de preparación.
git add archivo-4
git status
# * Ahora se eliminará al archivo y al objeto ```archivo-4```.
git rm archivo-4 -f
ls -a
git ls-files
# ## Despliegue de la historia de un repositorio o un archivo.
#
# El comando ```git log``` permite desplegar la historia de un repositorio o de un archivo.
#
# La documentación de referencia de ```git log``` está disponible en:
#
# https://git-scm.com/docs/git-log
# ### Despliegue de la historia de un repositorio.
#
# La sintaxis para desplegar la historia de un repositorio es la siguiente:
#
# ``` bash
# git log
# ```
# * Para un archivo se utiliza la siguiente sintaxis:
#
# ``` bash
# git log <archivo>
# ```
#
# El resutado es un listado de todos los commits empezando por el más reciente con los siguientes datos.
#
# * El número completo del identficador del commit y la rama del commit.
# * El nombre del autor según el campo ```user.name``` de la configuración.
# * La dirección de correo del autor según el campo ```user.emal``` de la configuración.
# **Ejemplo:**
# * La siguiente celda mostrará la historia de los commits realizados en el repositorio ```demo```.
git log
# ### Despliegue de la historia de un archivo.
#
# La sintaxis para desplegar la historia de un archivo es:
#
# ``` bash
# git log <archivo>
# ```
#
# El resutado es un listado de los commits en los que el archivo en cuestión fue modificado.
# **Ejemplo:**
# * La siguiente celda mostrará la historia de los commits en los que se modificó el archivo ```archivo-1``` en el repositorio ```demo```.
git log .gitignore
# ### La opción ```--oneline```.
#
# Esta opción permite desplegar un listado resumido en una sola línea con la siguiente estructura:
#
#
# ``` bash
# <identificador corto> (HEAD -> <rama>) <mensaje>
# ```
#
# Donde:
#
# * ```<rama>``` es la rama del repositorio en la que se realzia el commit. La rama que se crra pord efecto en un repositorio es ```master```.
# * ```<tipo de commit>``` este dato es opcional y cuando se realiza el primer commti de un repoitorio despliega el mensaje ```root-commit```.
# * ```<identificador corto>```, el cual es un número hexadecimal de 7 dígitos que corresponde a los primeros 7 dígitos del número identificador completo del commit
# **Ejemplos:**
# * La siguiente celda mostrará la historia de los commits realizados en el repositorio ```demo``` con una descripción corta.
git log --oneline
# * La siguiente celda mostrará la historia de los commits en los que el archivo ```archivo-1``` del repositorio ```demo``` fue modificdo con una descripción corta.
git log --oneline archivo-1
# ## El comando ```git show```.
#
# El comando ```git show``` permite mostrar la información del commit más reciente, incluyendo un resumen de las modificaciones a los archivos registradas en dicho commit.
#
# ```
# git show <ruta>
# ```
# Donde:
# * ```ruta``` es la ruta a un archivo. En caso de que no se indique la ruta, se traerá la información de todos los archivos modificados en el commit.
#
#
# La documentación de referencia de ```git show``` está disponible en:
#
# https://git-scm.com/docs/git-show
# **Ejemplo:**
# * La siguiente celda mostrará los detalles registrados en el commit más reciente.
git show
# * La siguiente celda mostrará los detalles registrados en el commit más recientes afectando al archivo ```archivo-1```. En vista de que no hubo afectaciones, no regresará nada.
git show archivo-1
# ## Comparación entre archivos de distintos commits.
#
# El comando ```git diff```permite entre otras cosas visualizar las diferencias de uno o varios archivos entre un commit y otro.
#
#
# La sintaxis de este comando es:
#
# ``` bash
# git diff <identificador 1> <identificador 2>
# ```
#
# Donde:
#
# * ```<identificador 1>``` e ```<identificador 2>``` son los dígitos iniciales de los números identificadores de algún commit realizado. El número de dígitos es arbitrario, siempre que el commit pueda ser identificado plenamente.
#
#
# El resultado es un listado de cada objetos que presente variaciones entre un commit y otro, así como la descripción de dichas diferencias.
#
# La documentación de referencia de ```git diff``` está disponible en:
#
# https://git-scm.com/docs/git-diff
# **Ejemplo:**
# * la siguiente celda añadirá una línea de texto al archivo ```archivo-1```.
echo Hola > archivo-1
cat archivo-1
# * La siguiente celda traerá al área de preparación aquellos objetos que hayan sido modificados posteriormente al último commit y hará un nuevo commit con el mensaje ```cuarto commit```
git commit -am "cuarto commit"
# <img src="img/02/cuarto_commit.svg" width="350px">
# * La siguiente celda enlistará de forma corta todos los commits. El resultado será similar a los siguiente:
#
# ```bash
# a1c53ed (HEAD -> master) cuarto commit
# 52c9f8c segundo commit
# e71f6f8 primer commit
# ```
# **Nota:** Cabe aclarar que los números identificadores siempre serán distintos en cada caso.
git log --oneline
# * Para obtener un listado de diferencias entre el commit con descripción ```cuarto commit``` (en este ejemplo correspondería al identificador ```ba397e8```) y el que tiene la descripción ```primer commit``` (en este ejemplo correspondería al identificador ```ed7d117```).
#
# La operación sería similar a lo siguiente.
#
# ``` bash
# > git diff ba397e8 ed7d117
# ```
# Y el resultado sería similar al siguiente.
#
# ``` bash
# diff --git a/archivo-1 b/archivo-1
# index a19abfe..e69de29 100644
# --- a/archivo-1
# +++ b/archivo-1
# @@ -1 +0,0 @@
# -Hola
# diff --git a/archivo-3 b/archivo-3
# new file mode 100644
# index 0000000..e69de29
# ```
#
# **Nota:** Cabe hacer notar que cada objeto del índice tiene su propio número identificador.
git diff 6cf7fcb 81392ae # NOTA: Es necesario sustituir los números identificadores.
git diff 81392ae 6cf7fcb
# ### Visualización de cambios de un archivo específico en distintos commits.
#
# Para conocer las diferencias
#
#
# La sintaxis de este comando es:
#
# ``` bash
# git diff <identificador 1> <identificador 2> <ruta>
# ```
#
# Donde:
#
# * ```<identificador 1>``` e ```<identificador 2>``` son los dígitos iniciales de los números identificadores de algún commit realizado. El número de dígitos es arbitrario, siempre que el commit pueda ser identificado plenamente.
#
# * ```<ruta>``` es la ruta del archivo específico dentro del respositorio.
# **Ejemplo:**
#
# * Para conocer los cambios ocurridos en el archivo ```archivo-1``` entre el commit ```ed7d11``` y el commit ```ba397e8``` se ejecutaría lo siguiente:
#
# ``` bash
# > git diff ed7d117 ba397e8 archivo-1
# ```
#
# * Y el resultado sería similar al siguiente:
#
# ``` bash
# diff --git a/archivo-1 b/archivo-1
# index e69de29..a19abfe 100644
# --- a/archivo-1
# +++ b/archivo-1
# @@ -0,0 +1 @@
# +Hola
# ```
git diff 6cf7fcb 81392ae archivo-1 # NOTA: Es necesario sustituir los números identificadores.
# <p style="text-align: center"><a rel="license" href="http://creativecommons.org/licenses/by/4.0/"><img alt="Licencia Creative Commons" style="border-width:0" src="https://i.creativecommons.org/l/by/4.0/80x15.png" /></a><br />Esta obra está bajo una <a rel="license" href="http://creativecommons.org/licenses/by/4.0/">Licencia Creative Commons Atribución 4.0 Internacional</a>.</p>
# <p style="text-align: center">© <NAME>. 2020.</p>
| 02_gestion_de_repositorios.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Add CIDS to parsed_threshold_data_in_air.csv
import pandas as pd
import pyrfume
from pyrfume.odorants import get_cid, get_cids
from rickpy import ProgressBar
df = pyrfume.load_data('thresholds/parsed_threshold_data_in_air.csv')
df = df.set_index('canonical SMILES')
smiles_cids = get_cids(df.index, kind='SMILES')
df = df.join(pd.Series(smiles_cids, name='CID'))
df.head()
from rdkit.Chem import MolFromSmiles, MolToSmiles
df['SMILES'] = df.index
p = ProgressBar(len(smiles_cids))
for i, (old, cid) in enumerate(smiles_cids.items()):
p.animate(i, status=old)
if cid == 0:
mol = MolFromSmiles(old)
if mol is None:
new = ''
else:
new = MolToSmiles(mol, isomericSmiles=True)
if old != new:
cid = get_cid(new, kind='SMILES')
df.loc[old, ['SMILES', 'CID']] = [new, cid]
p.animate(i+1, status='Done')
df[df['SMILES']=='']
ozone_smiles = ozone_cid = get_cid('[O-][O+]=O', kind='SMILES')
df.loc['O=[O]=O', ['SMILES', 'CID']] = [ozone_smiles, ozone_cid]
df = df.set_index('CID').drop(['ez_smiles'], axis=1)
df = df.rename(columns={'author': 'year', 'year': 'author'})
df.head()
pyrfume.save_data(df, 'thresholds/parsed_threshold_data_in_air_fixed.csv')
| notebooks/thresholds-add-cids.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .ps1
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: .NET (PowerShell)
# language: PowerShell
# name: .net-powershell
# ---
# # T1557.001 - LLMNR/NBT-NS Poisoning and SMB Relay
# By responding to LLMNR/NBT-NS network traffic, adversaries may spoof an authoritative source for name resolution to force communication with an adversary controlled system. This activity may be used to collect or relay authentication materials.
#
# Link-Local Multicast Name Resolution (LLMNR) and NetBIOS Name Service (NBT-NS) are Microsoft Windows components that serve as alternate methods of host identification. LLMNR is based upon the Domain Name System (DNS) format and allows hosts on the same local link to perform name resolution for other hosts. NBT-NS identifies systems on a local network by their NetBIOS name. (Citation: Wikipedia LLMNR) (Citation: TechNet NetBIOS)
#
# Adversaries can spoof an authoritative source for name resolution on a victim network by responding to LLMNR (UDP 5355)/NBT-NS (UDP 137) traffic as if they know the identity of the requested host, effectively poisoning the service so that the victims will communicate with the adversary controlled system. If the requested host belongs to a resource that requires identification/authentication, the username and NTLMv2 hash will then be sent to the adversary controlled system. The adversary can then collect the hash information sent over the wire through tools that monitor the ports for traffic or through [Network Sniffing](https://attack.mitre.org/techniques/T1040) and crack the hashes offline through [Brute Force](https://attack.mitre.org/techniques/T1110) to obtain the plaintext passwords. In some cases where an adversary has access to a system that is in the authentication path between systems or when automated scans that use credentials attempt to authenticate to an adversary controlled system, the NTLMv2 hashes can be intercepted and relayed to access and execute code against a target system. The relay step can happen in conjunction with poisoning but may also be independent of it. (Citation: byt3bl33d3r NTLM Relaying)(Citation: Secure Ideas SMB Relay)
#
# Several tools exist that can be used to poison name services within local networks such as NBNSpoof, Metasploit, and [Responder](https://attack.mitre.org/software/S0174). (Citation: GitHub NBNSpoof) (Citation: Rapid7 LLMNR Spoofer) (Citation: GitHub Responder)
# ## Atomic Tests:
# Currently, no tests are available for this technique.
# ## Detection
# Monitor <code>HKLM\Software\Policies\Microsoft\Windows NT\DNSClient</code> for changes to the "EnableMulticast" DWORD value. A value of “0” indicates LLMNR is disabled. (Citation: Sternsecurity LLMNR-NBTNS)
#
# Monitor for traffic on ports UDP 5355 and UDP 137 if LLMNR/NetBIOS is disabled by security policy.
#
# Deploy an LLMNR/NBT-NS spoofing detection tool.(Citation: GitHub Conveigh) Monitoring of Windows event logs for event IDs 4697 and 7045 may help in detecting successful relay techniques.(Citation: Secure Ideas SMB Relay)
| playbook/tactics/credential-access/T1557.001.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import cv2
import PIL.Image as PilImage
import torch
from fastai.core import *
from fastai.vision import *
# +
# implementation of Hasler's measure of perceptual colorfulness (2003)
# as reviewed by Amati et al. (2014), this method aligns well with actual survey
# for modeling an average human's perceptual colorfulness
def colorfulness(im: PilImage):
r,g,b = im.split()
r = np.asarray(r, dtype="float32")
g = np.asarray(g, dtype="float32")
b = np.asarray(b, dtype="float32")
rg = np.absolute(r - g)
yb = np.absolute((0.5 *(r + g)) - b)
std_rg, std_yb = np.std(rg), np.std(yb)
mean_rg, mean_yb = np.mean(rg), np.mean(yb)
std_rgyb = np.sqrt(np.square(std_rg) + np.square(std_yb))
mean_rgyb = np.sqrt(np.square(mean_rg) + np.square(mean_yb))
return std_rgyb + (0.3 * mean_rgyb)
# Returns average colorfulness from a string or Path
# input: path (Str), log (bool)
# out: colorfulness (float)
def average_colorfulness(path, log=True):
il = ItemList.from_folder(path)
col = []
for i in il:
im = PilImage.open(i)
col.append(colorfulness(im))
if log: print(str(path)+" :"+str(np.mean(col)))
return np.mean(col)
# -
path_a = '/mnt/Data/datasets/au_bw/results/INP_CL2/au_lg_800/40/'
path_b = '/mnt/Data/datasets/au_bw/results/INP_CL/au_lg_800/40/'
path_c = '/mnt/Data/datasets/au_bw/results/INP_CLalt1/au_lg_800_results'
average_colorfulness(path_a)
average_colorfulness(path_b)
average_colorfulness(path_c)
path_a = '/mnt/Data/datasets/au_bw/results/INP_CL2/au_med_800/40/'
path_b = '/mnt/Data/datasets/au_bw/results/INP_CL/au_med_800/40/'
path_c = '/mnt/Data/datasets/au_bw/results/INP_CLalt1/au_med_800_results'
average_colorfulness(path_a)
average_colorfulness(path_b)
average_colorfulness(path_c)
# ## Calculate Colorfulness
# +
# example usage
scrape_sizes = ['lg', 'med']
current_sizes = [1200, 1000, 800, 600, 400]
render_factors = [90,80,70,60,50,40,30,20,10]
save = [] # saved for copying convenience
for scrape_size in scrape_sizes:
for current_size in current_sizes:
for render_factor in render_factors:
input_path = '/mnt/Data/datasets/au_bw/results/CL/'+ 'au_' + scrape_size + '_' + str(current_size) + '/' + str(render_factor) + '/'
avg_col = average_colorfulness(input_path)
save.append(avg_col)
print("Data: " + scrape_size +' '+ str(current_size) +' '+ str(render_factor) + ". Result: " + str(avg_col))
# -
| calculate_colorfulness.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:.conda-ngaml]
# language: python
# name: conda-env-.conda-ngaml-py
# ---
# # High-level RNN PyTorch Example
# *Modified by <NAME> (jordancaraballo)*
import os
import sys
import numpy as np
import math
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
import torch.utils.data as data_utils
import torch.nn.init as init
from torch import autograd
from torch.autograd import Variable
from common.params_lstm import *
from common.utils import *
# Force one-gpu
os.environ["CUDA_VISIBLE_DEVICES"] = "0"
print("OS: ", sys.platform)
print("Python: ", sys.version)
print("PyTorch: ", torch.__version__)
print("Numpy: ", np.__version__)
print("GPU: ", get_gpu_name())
print(get_cuda_version())
print("CuDNN Version ", get_cudnn_version())
class SymbolModule(nn.Module):
def __init__(self,
maxf=MAXFEATURES, edim=EMBEDSIZE, nhid=NUMHIDDEN):
super(SymbolModule, self).__init__()
self.embedding = nn.Embedding(num_embeddings=maxf,
embedding_dim=edim)
# If batch-first then input and output
# provided as (batch, seq, features)
# Cudnn used by default if possible
self.gru = nn.GRU(input_size=edim,
hidden_size=nhid,
num_layers=1,
batch_first=True,
bidirectional=False)
self.l_out = nn.Linear(in_features=nhid*1,
out_features=2)
def forward(self, x, nhid=NUMHIDDEN, batchs=BATCHSIZE):
x = self.embedding(x)
h0 = Variable(torch.zeros(1, batchs, nhid)).cuda()
x, h = self.gru(x, h0) # outputs, states
# just get the last output state
x = x[:,-1,:].squeeze()
x = self.l_out(x)
return x
def init_model(m, lr=LR, b1=BETA_1, b2=BETA_2, eps=EPS):
opt = optim.Adam(m.parameters(), lr, betas=(b1, b2), eps=eps)
criterion = nn.CrossEntropyLoss()
return opt, criterion
# %%time
# Data into format for library
x_train, x_test, y_train, y_test = imdb_for_library(seq_len=MAXLEN, max_features=MAXFEATURES)
# Torch-specific
x_train = x_train.astype(np.int64)
x_test = x_test.astype(np.int64)
y_train = y_train.astype(np.int64)
y_test = y_test.astype(np.int64)
print(x_train.shape, x_test.shape, y_train.shape, y_test.shape)
print(x_train.dtype, x_test.dtype, y_train.dtype, y_test.dtype)
# %%time
sym = SymbolModule()
sym.cuda() # CUDA!
# %%time
optimizer, criterion = init_model(sym)
# %%time
# Main training loop: 12.7s
sym.train() # Sets training = True
EPOCHS=40
for j in range(EPOCHS):
for data, target in yield_mb(x_train, y_train, BATCHSIZE, shuffle=True):
# Get samples
data = Variable(torch.LongTensor(data).cuda())
target = Variable(torch.LongTensor(target).cuda())
# Init
optimizer.zero_grad()
# Forwards
output = sym(data)
# Loss
loss = criterion(output, target)
# Back-prop
loss.backward()
optimizer.step()
# Log
print(j)
# %%time
# Main evaluation loop: 1.52s
sym.eval() # Sets training = False
n_samples = (y_test.shape[0]//BATCHSIZE)*BATCHSIZE
y_guess = np.zeros(n_samples, dtype=np.int)
y_truth = y_test[:n_samples]
c = 0
for data, target in yield_mb(x_test, y_test, BATCHSIZE):
# Get samples
data = Variable(torch.LongTensor(data).cuda())
# Forwards
output = sym(data)
pred = output.data.max(1)[1].cpu().numpy().squeeze()
# Collect results
y_guess[c*BATCHSIZE:(c+1)*BATCHSIZE] = pred
c += 1
print("Accuracy: ", sum(y_guess == y_truth)/len(y_guess))
| notebooks/benchmarks/PyTorch_RNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Regression Practice 1
# ** In this practice project I've gone over how we can use statistics to manualy select features as well as how to interaction features into the model in order to improve performance **
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from regressors import stats
# %matplotlib inline
data = pd.read_csv('train.csv')
data.head()
sub_data = data[['OverallQual','OverallCond','YearBuilt','YearRemodAdd','1stFlrSF','TotRmsAbvGrd','FullBath','LotFrontage','LotArea','SalePrice']]
sub_data = sub_data.dropna()
sub_data.head()
sns.pairplot(sub_data)
X = sub_data[['OverallQual','OverallCond','YearBuilt','YearRemodAdd','1stFlrSF','TotRmsAbvGrd','FullBath','LotFrontage','LotArea']]
y = sub_data['SalePrice']
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.4, random_state=101)
from sklearn.linear_model import LinearRegression
lm = LinearRegression()
lm.fit(X_train,y_train)
coeff_df = pd.DataFrame(lm.coef_,X.columns,columns=['Coefficient'])
coeff_df
xlabels = list(sub_data.columns)[:-1]
stats.summary(lm, X_train,y_train,xlabels)
# ** Full bath and lot frontage are not significant **
predictions = lm.predict(X_test)
sns.distplot((y_test-predictions),bins=50);
import math
def rmsle(y, y_pred):
assert len(y) == len(y_pred)
terms_to_sum = [(math.log(y_pred[i] + 1) - math.log(y[i] + 1)) ** 2.0 for i,pred in enumerate(y_pred)]
return (sum(terms_to_sum) * (1.0/len(y))) ** 0.5
from sklearn import metrics
print('MAE:', metrics.mean_absolute_error(y_test, predictions))
print('MSE:', metrics.mean_squared_error(y_test, predictions))
print('RMSE:', np.sqrt(metrics.mean_squared_error(y_test, predictions)))
print('RMSLE:', np.sqrt(metrics.mean_squared_log_error(y_test, predictions)))
# ** Adding two new features modeling the first order interactions of construction year and remodelling year as well as overal quality and overal condition of the house **
sub_data['YBXYR']=sub_data['YearBuilt']*sub_date['YearRemodAdd']
sub_data['OQXOC']=sub_data['OverallQual']*sub_date['OverallCond']
X = sub_data.drop(['SalePrice','LotFrontage','FullBath'],axis=1)
y = sub_data['SalePrice']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.4, random_state=101)
lm2 = LinearRegression()
lm2.fit(X_train,y_train)
coeff_df = pd.DataFrame(lm2.coef_,X.columns,columns=['Coefficient'])
coeff_df
xlabels = list(X.columns)
stats.summary(lm2, X_train,y_train,xlabels)
predictions = lm2.predict(X_test)
print('MAE:', metrics.mean_absolute_error(y_test, predictions))
print('MSE:', metrics.mean_squared_error(y_test, predictions))
print('RMSE:', np.sqrt(metrics.mean_squared_error(y_test, predictions)))
print('RMSLE:', np.sqrt(metrics.mean_squared_log_error(y_test, predictions)))
# ** RMSLE has improved while RMSE has suffered which means the model is probably doing better for higher value homes but not as good as before on lower value one **
| Linear Regression/Housing/Regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Credit Card Fraud Detection::
# Download dataset from this link:
#
# https://www.kaggle.com/mlg-ulb/creditcardfraud
# # Description about dataset::
# The datasets contains transactions made by credit cards in September 2013 by european cardholders.
# This dataset presents transactions that occurred in two days, where we have 492 frauds out of 284,807 transactions. The dataset is highly unbalanced, the positive class (frauds) account for 0.172% of all transactions.
#
# It contains only numerical input variables which are the result of a PCA transformation. Unfortunately, due to confidentiality issues, we cannot provide the original features and more background information about the data. Features V1, V2, … V28 are the principal components obtained with PCA, the only features which have not been transformed with PCA are 'Time' and 'Amount'. Feature 'Time' contains the seconds elapsed between each transaction and the first transaction in the dataset. The feature 'Amount' is the transaction Amount, this feature can be used for example-dependant cost-senstive learning.
#
#
# ### Feature 'Class' is the response variable and it takes value 1 in case of fraud and 0 otherwise.
# # WORKFLOW :
# 1.Load Data
#
# 2.Check Missing Values ( If Exist ; Fill each record with mean of its feature )
#
# 3.Standardized the Input Variables.
#
# 4.Split into 50% Training(Samples,Labels) , 30% Test(Samples,Labels) and 20% Validation Data(Samples,Labels).
#
# 5.Model : input Layer (No. of features ), 3 hidden layers including 10,8,6 unit & Output Layer with activation function relu/tanh (check by experiment).
#
# 6.Compilation Step (Note : Its a Binary problem , select loss , metrics according to it)
#
# 7.Train the Model with Epochs (100).
#
# 8.If the model gets overfit tune your model by changing the units , No. of layers , epochs , add dropout layer or add Regularizer according to the need .
#
# 9.Prediction should be > 92%
# 10.Evaluation Step
# 11Prediction
#
# # Task::
# ## Identify fraudulent credit card transactions.
# +
import pandas as pd
# loadind data
file = 'creditcard.csv'
cc_data = pd.read_csv(file)
cc_data = pd.DataFrame(cc_data)
cc_data
# -
cc_data.head()
cc_data.describe()
cc_data.info() # to confirm about null values
# +
# data distribution into X & y
X = cc_data.loc[:, cc_data.columns != 'Class']
y = cc_data.Class
# +
# Split into 50% Training(Samples,Labels) , 30% Test(Samples,Labels) and 20% Validation Data(Samples,Labels).
from sklearn.model_selection import train_test_split
# ratios for the whole dataset.
train_ratio = 0.5
test_ratio = 0.3
validation_ratio = 0.2
# using train test split method
x_remaining, x_test, y_remaining, y_test = train_test_split(X, y, test_size=test_ratio)
# validation ratio from remaining dataset.
remaining = 1 - test_ratio
validation_adjusted = validation_ratio / remaining
# train and validation splits
x_train, x_validation, y_train, y_validation = train_test_split(x_remaining, y_remaining, test_size=validation_adjusted)
print(x_train.shape, y_train.shape, x_test.shape, y_test.shape, x_validation.shape, y_validation.shape)
# +
# model creation and training
import tensorflow as tf
import tensorflow.keras as keras
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense
from tensorflow.keras.optimizers import *
model = Sequential()
tf.keras.backend.set_floatx('float64')
model.add(tf.keras.layers.Dense(16, activation='relu'))
model.add(tf.keras.layers.Dense(1, activation='sigmoid'))
model.compile(loss='binary_crossentropy', optimizer='adam', metrics=['accuracy'])
model.fit(x_train, y_train, epochs=100, validation_data=(x_validation, y_validation))
# -
# # Evaluation
results = model.evaluate(x_test, y_test)
# # Predictions
pred = model.predict(x_validation)
pred # as accuracy is 99.8 % on evaluation
| Credit Card Fraud Detection assignment.ipynb |