code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.1.0
# language: julia
# name: julia-1.1
# ---
# # 1. Plain Vanilla Chebyshev
#
# In this exercise you should write some code that approximates the function $f(x) = x + 2x^2 - exp(-x)$ for $x\in[-3,3]$. You should define a function `q1(n)`, where `n` is the number of interpolation points. You should use an approximation of degree `deg=n-1`, and you should set `n=15` Chebyshev interpolation nodes.
#
# * predict `n_new=100` new equally spaced points in $[-3,3]$ using your interpolator.
# * make a plot with 2 panels. panel 1 shows true function values and your approximation and panel 2 shows the deviation in your approximation from the true $f$.
# * Write an automated test that passes if the maximal deviation in your approximation from the true $f$ is smaller than `1e-9`.
#
# # 2. Question 1. with `ApproxFun.jl`
#
# Redo exercise 1 with the `ApproxFun.jl` package. This should go into function `q2(n::Number)`, where now `n` is a positive number that will be used to define a symmetric interval around zero. So, you give `n=4`, you create $[-4,4]$ with `ApproxFun`.
#
# # 3. More fun with `ApproxFun.jl`
#
# Let's use the same setup as in question 2, `q3(n::Number)`. Now however, we want to combine 2 functions into a third one:
#
# $$
# f(x) = sin(x^2) \\
# g(x) = cos(x) \\
# h(x) = f(x) - g(x)
# $$
#
# Use `ApproxFun.jl` to define this approximation. then get all the roots of $h$, ie. all $x s.t. h(x) = 0$. Make a plot of $h(x),x\in[-10,10]$, and plot the roots of h onto the same plot. Finally, compute the definite integral
#
# $$
# \int_{-10}^0 h(z) dz
# $$
# and return it's value.
#
# Notice that by looking at the [readme](https://github.com/JuliaApproximation/ApproxFun.jl) of that package, you will go pretty far here.
#
#
# # 4. Plotting the Chebyshev Basis (Optional)
#
# plot the first nine Chebyshev basis functions in function `q4()`. You could take some inspiration for the plot from yesterday's slides.
#
# # 5. Importance of node placement: uniform vs chebyshev nodes (Optional)
#
# In this exercise we want to investigate one of the dangers with polynomial approximation: placement of knots. We will focus on the classic example of Runge's function:
# $$ f(x) = \frac{1}{1+25x^2} ,x\in[-5,5]$$
#
# Here we want to approximate $f$ with the Chebyshev polynomial. We want to learn the impact of evaluating the polynomial at an equidistant set of points as opposed to something else.
#
# * Produce a plot with 2 panels, *uniformly spaced* and *chebyshev nodes*:
# * Panel 1 should show approximations using a uniformly spaced grid of interpolation points.
# * Panel 2 should show approximations using chebyshev interpolation points.
# * Each panel should show 4 lines:
# 1. The true function
# 2. the resulting approximations from a Chebyshev Polynomial Interpolation of degrees $k=5,9,15$. Are we going to get a better approximation as we increase $k$?
# * For each approximation, you should choose $n=k+1$ interpolation points.
# * The code contains a custom `ChebyType` that I found useful to produce these plots. You may want to use it as well.
#
#
# # 6. Spline Interpolation: where to place knots? (Optional)
#
# * Produce a plot with 2 panels. Panel 1 (*Runge's function*) shows $f(x),x\in[-5,5]$, panel 2 (*Error in Runge's function*) shows deviations of your approximation to it (see below).
# * Use 2 versions of a cubic spline to approximate the function:
# * version 1: use 13 equally spaced knots when you set up the `BSpline` object. Remember, that this is
# ```julia
# using ApproXD
# b = BSpline(nknots,deg,lb,ub) # knot vector is chosen for you
# ```
# * version 2: use 13 knots that are concentrated towards 0.
# ```julia
# using ApproXD
# b = BSpline(my_knots,deg) # you choose my_knots
# ```
# * To estimate your approximating coefficients, evaluate the basis and $f$ at `nevals=5*13=65` equidistant points.
# * So, panel 2 of your plot should show 2 lines (`f - approx1` and `f - approx2`), as well as the placement of your concentrated knots.
#
# # 7. Splines and Kinks
#
# Interpolate the function $f(x) = |x|^{0.5},x\in[-1,1]$ with a cubic spline, with 13 knots, and 65 evaluation points to estimate it's coefficients.
#
# Produce a plot with 3 panels:
#
# 1. Plot the true function. It has a kink at 0.
# 1. plot two spline approximations.
# 1. a spline with a uniform knot vector.
# 1. a spline with a knot vector that has a knot multiplicity at the kink $x=0$. How many knots do you have to set equal to zero to produce a kink? note that the total number of (interior) knots should not change (i.e. 13).
# 1. In panel three plot the errors of both approximations.
| HWfuncappQuestions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="meczB1CRSQ8E" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 442} outputId="83422f8c-58b9-4df0-cbce-b82d794a3e74" executionInfo={"status": "ok", "timestamp": 1583371998950, "user_tz": -60, "elapsed": 12464, "user": {"displayName": "Pawe\u0142 C", "photoUrl": "", "userId": "15369492993684325952"}}
# !pip install --upgrade tables
# !pip install eli5
# + id="zh5J-eA-RLvv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 168} outputId="e33380ef-c1ba-4419-95be-6532c52d6705" executionInfo={"status": "ok", "timestamp": 1583372005023, "user_tz": -60, "elapsed": 3497, "user": {"displayName": "Pawe\u0142 C", "photoUrl": "", "userId": "15369492993684325952"}}
import pandas as pd
import numpy as np
from sklearn.dummy import DummyRegressor
from sklearn.tree import DecisionTreeRegressor
from sklearn.metrics import mean_absolute_error as mae
from sklearn.metrics import mean_squared_error
from sklearn.model_selection import cross_val_score
import eli5
from eli5.sklearn import PermutationImportance
# + id="XHewA41nVWKI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="337d7a1d-ca93-4605-f816-b2442d42615b" executionInfo={"status": "ok", "timestamp": 1583372148031, "user_tz": -60, "elapsed": 1723, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15369492993684325952"}}
# ls
# + id="YkJNnWXTT40r" colab_type="code" outputId="27adb684-c5d0-4ef2-8b73-b171eb3c42da" executionInfo={"status": "ok", "timestamp": 1583372298022, "user_tz": -60, "elapsed": 599, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15369492993684325952"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# cd "dw_matrix_car"
# + id="3P28HnZ7Unuz" colab_type="code" outputId="15d30b1d-16f6-47a0-eb48-ca63bd64f42c" executionInfo={"status": "ok", "timestamp": 1583372306031, "user_tz": -60, "elapsed": 3911, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15369492993684325952"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
df = pd.read_hdf('data/car.h5')
df.shape
# + id="VX_scl41Vjd7" colab_type="code" outputId="488b9854-ac41-42ff-acd4-b48e529d9f30" executionInfo={"status": "ok", "timestamp": 1583372308903, "user_tz": -60, "elapsed": 607, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15369492993684325952"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
df.select_dtypes(np.number).columns
# + id="LTOs4fixV6JA" colab_type="code" outputId="cde3755d-7957-4e1c-b094-e5cba981e574" executionInfo={"status": "ok", "timestamp": 1583372316597, "user_tz": -60, "elapsed": 599, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15369492993684325952"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
feats = ['car_id']
X = df[ feats ].values
y = df['price_value'].values
model = DummyRegressor()
model.fit(X,y)
y_pred = model.predict(X)
mae(y, y_pred)
# + id="raFa-vhkXu80" colab_type="code" outputId="21d09be4-1faf-474f-e8a7-7bb3ddf78df3" executionInfo={"status": "ok", "timestamp": 1583372318362, "user_tz": -60, "elapsed": 610, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15369492993684325952"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
[x for x in df.columns if 'price' in x]
# + id="gqOTKOneZGam" colab_type="code" outputId="35523280-2560-4e96-d526-54bb75226540" executionInfo={"status": "ok", "timestamp": 1583372320638, "user_tz": -60, "elapsed": 589, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15369492993684325952"}} colab={"base_uri": "https://localhost:8080/", "height": 68}
df['price_currency'].value_counts()
# + id="JiNg7840ZdO-" colab_type="code" outputId="84a96497-a818-44c6-bc02-c3b755aa7bc3" executionInfo={"status": "ok", "timestamp": 1583372323664, "user_tz": -60, "elapsed": 637, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15369492993684325952"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
df = df[ df['price_currency'] != 'EUR']
df.shape
# + id="pWdi-9Yua6b6" colab_type="code" outputId="91f3d921-fec6-4f2b-c343-8f4e12f7b336" executionInfo={"status": "ok", "timestamp": 1583372339666, "user_tz": -60, "elapsed": 578, "user": {"displayName": "Pawe\u014<NAME>", "photoUrl": "", "userId": "15369492993684325952"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
df['param_color'].factorize()[0]
# + id="fb0WyBhla2XU" colab_type="code" colab={}
SUFFIX_CAT = '__cat'
for feat in df.columns:
if isinstance(df[feat][0],list):continue
factorized_values = df[feat].factorize()[0]
if SUFFIX_CAT in feat:
df [feat] = factorized_values
else:
df [feat + SUFFIX_CAT] = factorized_values
# + id="i4-Tc-ZfdnUu" colab_type="code" outputId="44316672-18cf-477d-b5f7-06e8890b36f8" executionInfo={"status": "ok", "timestamp": 1583372359305, "user_tz": -60, "elapsed": 570, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15369492993684325952"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
cat_feats = [x for x in df.columns if SUFFIX_CAT in x ]
cat_feats = [x for x in cat_feats if 'price' not in x ]
len(cat_feats)
# + id="7KTSE1HPg-AS" colab_type="code" outputId="86b3e9c6-bb3d-46b7-dfba-115281f0a7c4" executionInfo={"status": "ok", "timestamp": 1583372384539, "user_tz": -60, "elapsed": 3800, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15369492993684325952"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
X = df[ cat_feats ].values
y = df['price_value'].values
model= DecisionTreeRegressor(max_depth=5)
scores = cross_val_score(model, X, y, cv=3, scoring='neg_mean_absolute_error')
np.mean(scores)
# + id="0QCcdOwohdHW" colab_type="code" outputId="dfa0c398-d37d-4623-91f1-937e3fc7594e" executionInfo={"status": "ok", "timestamp": 1583372433211, "user_tz": -60, "elapsed": 45682, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15369492993684325952"}} colab={"base_uri": "https://localhost:8080/", "height": 391}
m = DecisionTreeRegressor(max_depth=5)
m.fit(X,y)
imp = PermutationImportance(m, random_state=0).fit(X,y)
eli5.show_weights(imp, feature_names=cat_feats)
# + id="isbd2PDWh6ww" colab_type="code" colab={}
| day3_simple_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Integration by parts is another technique for simplifying integrands. As we saw in previous posts, each differentiation rule has a corresponding integration rule. In the case of integration by parts, the corresponding differentiation rule is the Product Rule. The technique of integration by parts allows us to simplify integrands of the form:
# $$ \int f(x) g(x) dx $$
# Examples of this form include:
# $$ \int x \cos{x} \space dx, \qquad \int e^x \cos{x} \space dx, \qquad \int x^2 e^x \space dx $$
#
# As integration by parts is the product rule applied to integrals, it helps to state the Product Rule again. The Product Rule is defined as:
# $$ \frac{d}{dx} \big[ f(x)g(x) \big] = f^{\prime}(x) g(x) + f(x) g^{\prime}(x) $$
#
# When we apply the product rule to indefinite integrals, we can restate the rule as:
#
# $$ \int \frac{d}{dx} \big[f(x)g(x)\big] \space dx = \int \big[f^{\prime} g(x) + f(x) g^{\prime}(x) \big] \space dx $$
#
# Then, rearranging so we get $f(x)g^{\prime}(x) \space dx$ on the left side of the equation:
#
# $$ \int f(x)g^{\prime}(x) \space dx = \int \frac{d}{dx} \big[f(x)g(x)\big] \space dx - \int f^{\prime}(x)g(x) \space dx $$
#
# Which gives us the integration by parts formula! The formula is typically written in differential form:
#
# $$ \int u \space dv = uv - \int v \space du $$
# ## Examples
#
# The following examples walkthrough several problems that can be solved using integration by parts. We also employ the wonderful [SymPy](https://www.sympy.org/en/index.html) package for symbolic computation to confirm our answers. To use SymPy later to verify our answers, we load the modules we will require and initialize several variables for use with the SymPy library.
# +
from sympy import symbols, limit, diff, sin, cos, log, tan, sqrt, init_printing, plot, integrate
from mpmath import ln, e, pi, cosh, sinh
init_printing()
x = symbols('x')
y = symbols('y')
# -
# Example 1: Evaluate the integrand $ \int x \sin{\frac{x}{2}} \space dx $
#
# Recalling the differential form of the integration by parts formula, $ \int u \space dv = uv - \int v \space du $, we set $u = x$ and $dv = \sin{\frac{x}{2}}$
#
# Solving for the derivative of $u$, we arrive at $du = 1 \space dx = dx$. Next, we find the antiderivative of $dv$. To find this antiderivative, we employ the Substitution Rule.
#
# $$ u = \frac{1}{2}x, \qquad du = {1}{2} \space dx, \qquad \frac{du}{dx} = 2 $$
# $$ y = \sin{u}, \qquad dy = -\cos{u} \space du, \qquad \frac{dy}{du} = -\cos{u} $$
#
# Therefore, $v = -2 \cos{\frac{x}{2}}$
#
# Entering these into the integration by parts formula:
#
# $$ -2x\cos{\frac{x}{2}} - (-2)\int \cos{\frac{x}{2}} $$
#
# Then, solving for the integrand $\int \cos{\frac{x}{2}}$, we employ the Substitution Rule again as before to arrive at $2\sin{\frac{x}{2}}$ (the steps in solving this integrand are the same as before when we solved for $\int \sin{\frac{x}{2}}$). Thus, the integral is evaluated as:
#
# $$ -2x\cos{\frac{x}{2}} + 4\sin{\frac{x}{2}} + C $$
#
# Using SymPy's [`integrate`](https://docs.sympy.org/latest/modules/integrals/integrals.html), we can verify our answer is correct (SymPy does not include the constant of integration $C$).
integrate(x * sin(x / 2), x)
# Example 2: Evaluate $\int t^2 \cos{t} \space dt$
#
# We start by setting $u = t^2$ and $dv = \cos{t}$. The derivative of $t^2$ is $2t$, thus $du = 2t \space dt$, or $\frac{du}{dt} = 2t$. Integrating $dv = \cos{t}$ gives us $v = \sin{t} \space du$. Entering these into the integration by parts formula:
#
# $$ t^2 \sin{t} - 2\int t \sin{t} $$
#
# Therefore, we must do another round of integration by parts to solve $\int t \sin{t}$.
#
# $$ u = t, \qquad du = dt $$
# $$ dv = \sin{t}, \qquad v = -\cos{t} \space du $$
#
# Putting these together into the integration by parts formula with the above:
#
# $$ t^2 \sin{t} - 2 \big(-t \cos{t} + \int \cos{t} \space dt \big) $$
#
# Which gives us the solution:
#
# $$ t^2 \sin{t} + 2t \cos{t} - 2 \sin{t} + C$$
#
# As before, we can verify that our answer is correct by leveraging SymPy.
t = symbols('t')
integrate(t ** 2 * cos(t), t)
# Example 3: $\int x e^x \space dx$
#
#
# Here, we set $u = x$ and $dv = e^x$. Therefore, $du = dx$ and $v = e^x \space dx$. Putting these together in the integration by parts formula:
#
# $$ xe^x - \int e^x $$
#
# As the integral of $e^x$ is just $e^x$, our answer is:
#
# $$ xe^x - e^x + C $$
#
# We can again verify our answer is accurate using SymPy.
integrate(x * e ** x, x)
# ## References
# <NAME>. and <NAME>. (n.d.). Thomas' calculus. 13th ed.
#
# <NAME>. (2007). Essential calculus: Early transcendentals. Belmont, CA: Thomson Higher Education.
| content/draft/Integration by Parts.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/", "height": 79} colab_type="code" id="wYtuKeK0dImp" outputId="d31adfc3-f94b-4b18-f09a-0095a4b80e49"
import csv
import numpy as np
import tensorflow as tf
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from google.colab import files
# + [markdown] colab_type="text" id="EmMyh9_mkDHF"
# The data for this exercise is available at: https://www.kaggle.com/datamunge/sign-language-mnist/home
#
# Sign up and download to find 2 CSV files: sign_mnist_test.csv and sign_mnist_train.csv -- You will upload both of them using this button before you can continue.
#
# + colab={"base_uri": "https://localhost:8080/", "height": 104, "resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY> "headers": [["content-type", "application/javascript"]], "ok": true, "status": 200, "status_text": ""}}} colab_type="code" id="IcLOZlnnc_N7" outputId="96ae5f29-d1ed-4036-de8b-3ba722d9d3f0"
uploaded=files.upload()
# + colab={"base_uri": "https://localhost:8080/", "height": 84} colab_type="code" id="4kxw-_rmcnVu" outputId="18ffd5e8-6130-4f87-a42f-a6280a85f05e"
def get_data(filename):
# You will need to write code that will read the file passed
# into this function. The first line contains the column headers
# so you should ignore it
# Each successive line contians 785 comma separated values between 0 and 255
# The first value is the label
# The rest are the pixel values for that picture
# The function will return 2 np.array types. One with all the labels
# One with all the images
#
# Tips:
# If you read a full line (as 'row') then row[0] has the label
# and row[1:785] has the 784 pixel values
# Take a look at np.array_split to turn the 784 pixels into 28x28
# You are reading in strings, but need the values to be floats
# Check out np.array().astype for a conversion
with open(filename) as training_file:
csv_rows = csv.reader(training_file, delimiter=',')
next(csv_rows) #skip csv header
images_list = []
labels_list = []
for row in csv_rows:
temp_image = row[1:785]
temp_img_array = np.array_split(temp_image, 28)
images_list.append(temp_img_array)
labels_list.append(row[0])
images = np.array(images_list).astype('float')
labels = np.array(labels_list).astype('float')
return images, labels
training_images, training_labels = get_data('sign_mnist_train.csv')
testing_images, testing_labels = get_data('sign_mnist_test.csv')
# Keep these
print(training_images.shape)
print(training_labels.shape)
print(testing_images.shape)
print(testing_labels.shape)
# Their output should be:
# (27455, 28, 28)
# (27455,)
# (7172, 28, 28)
# (7172,)
# + colab={"base_uri": "https://localhost:8080/", "height": 50} colab_type="code" id="awoqRpyZdQkD" outputId="5f7fb471-bf8b-4dff-8d84-633a68c97149"
# In this section you will have to add another dimension to the data
# So, for example, if your array is (10000, 28, 28)
# You will need to make it (10000, 28, 28, 1)
# Hint: np.expand_dims
training_images = np.expand_dims(training_images, axis=3)
testing_images = np.expand_dims(testing_images, axis=3)
# Create an ImageDataGenerator and do Image Augmentation
train_datagen = ImageDataGenerator(
rescale=1./255,
rotation_range=40,
height_shift_range=0.2,
width_shift_range=0.2,
shear_range=0.2,
zoom_range=0.2,
horizontal_flip=True,
fill_mode='nearest')
validation_datagen = ImageDataGenerator(
rescale=1./255)
# Keep These
print(training_images.shape)
print(testing_images.shape)
# Their output should be:
# (27455, 28, 28, 1)
# (7172, 28, 28, 1)
# + colab={"base_uri": "https://localhost:8080/", "height": 826} colab_type="code" id="Rmb7S32cgRqS" outputId="e4437f1e-b267-48a7-89b4-49c716460eea"
# Define the model
# Use no more than 2 Conv2D and 2 MaxPooling2D
model = tf.keras.models.Sequential([
tf.keras.layers.Conv2D(32, (3,3), activation='relu', input_shape=(28,28,1)),
tf.keras.layers.MaxPooling2D(2,2),
tf.keras.layers.Conv2D(64, (3,3), activation='relu'),
tf.keras.layers.MaxPooling2D(2,2),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(512, activation='relu'),
tf.keras.layers.Dense(26, activation='softmax')])
# Compile Model.
model.compile(optimizer = tf.train.AdamOptimizer(),
loss = 'sparse_categorical_crossentropy',
metrics=['accuracy'])
# Train the Model
history = model.fit_generator(train_datagen.flow(training_images, training_labels, batch_size=32),
steps_per_epoch=len(training_images) / 32,
epochs=15,
validation_data=validation_datagen.flow(testing_images, testing_labels, batch_size=32),
validation_steps=len(testing_images) / 32)
model.evaluate(testing_images, testing_labels)
# The output from model.evaluate should be close to:
[6.92426086682151, 0.56609035]
# + colab={"base_uri": "https://localhost:8080/", "height": 545} colab_type="code" id="_Q3Zpr46dsij" outputId="643549e5-a06b-4c6a-bc99-2ea6003e17fa"
# Plot the chart for accuracy and loss on both training and validation
import matplotlib.pyplot as plt
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(len(acc))
plt.plot(epochs, acc, 'r', label='Training accuracy')
plt.plot(epochs, val_acc, 'b', label='Validation accuracy')
plt.title('Training and validation accuracy')
plt.legend()
plt.figure()
plt.plot(epochs, loss, 'r', label='Training Loss')
plt.plot(epochs, val_loss, 'b', label='Validation Loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
# + colab={} colab_type="code" id="0UNPboS21jEO"
| Course 2 - CNNs in Tensorflow/.ipynb_checkpoints/Exercise_8_sign_language_mnist-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/marilynle/DS-Unit-1-Sprint-1-Dealing-With-Data/blob/master/Marilyn_Esko_LS_DSPT3_113_Making_Data_backed_Assertions_Assignment_3.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="Okfr_uhwhS1X" colab_type="text"
# # Lambda School Data Science - Making Data-backed Assertions
#
# This is, for many, the main point of data science - to create and support reasoned arguments based on evidence. It's not a topic to master in a day, but it is worth some focused time thinking about and structuring your approach to it.
# + [markdown] id="lOqaPds9huME" colab_type="text"
# ## Assignment - what's going on here?
#
# Consider the data in `persons.csv` (already prepared for you, in the repo for the week). It has four columns - a unique id, followed by age (in years), weight (in lbs), and exercise time (in minutes/week) of 1200 (hypothetical) people.
#
# Try to figure out which variables are possibly related to each other, and which may be confounding relationships.
#
# Try and isolate the main relationships and then communicate them using crosstabs and graphs. Share any cool graphs that you make with the rest of the class in Slack!
# + id="TGUS79cOhPWj" colab_type="code" colab={}
# TODO - your code here
# Use what we did live in lecture as an example
# HINT - you can find the raw URL on GitHub and potentially use that
# to load the data with read_csv, or you can upload it yourself
# + id="p2RpCH_zn93j" colab_type="code" outputId="26a6f772-f393-423e-dbd1-d09c103975d5" colab={"base_uri": "https://localhost:8080/", "height": 102}
# !pip install pandas==0.23.4
# + id="Z7KuhGy0oEUc" colab_type="code" colab={}
import pandas as pd
# + id="zwCyoTsAqeer" colab_type="code" outputId="d790e730-64d9-4242-ac2b-81bf86952172" colab={"base_uri": "https://localhost:8080/", "height": 359}
# Before I loaded the data, I looked at the raw text at: https://https://github.com/marilynle/DS-Unit-1-Sprint-1-Dealing-With-Data/blob/master/module3-databackedassertions/persons.csv
persons_data = pd.read_csv('https://raw.githubusercontent.com/LambdaSchool/DS-Unit-1-Sprint-1-Dealing-With-Data/master/module3-databackedassertions/persons.csv')
persons_data.head(10)
# + id="BHvEqjWE0HiH" colab_type="code" outputId="4ad6ce33-1b2e-4705-81ea-2d997d241c1b" colab={"base_uri": "https://localhost:8080/", "height": 102}
persons_data.dtypes
# + id="Dii8lWHN0iZW" colab_type="code" outputId="9ed3518d-2515-4a42-9744-ca99ed42e85c" colab={"base_uri": "https://localhost:8080/", "height": 297}
persons_data.describe()
# + id="2K8lXIhkvjxx" colab_type="code" outputId="fc8724ee-7a92-4f14-caa6-c5fd327ed691" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# Using Cross-tabulation to observe the relationship between the variables.
pd.crosstab(persons_data['weight'], persons_data['exercise_time'])
# + id="ZtxEmdCdF4gk" colab_type="code" outputId="b73c9e12-88f5-4937-d265-7b419ed87353" colab={"base_uri": "https://localhost:8080/", "height": 303}
# Making 5 equal-sized bins for each variable, using pandas.cut to make them
# IntervalIndex defines the exact bins to be used, then we can set the intervals of bins manually.
# The bins makes easier to observe the relations between the variables at the Cross-tabulation
bins_1 = pd.IntervalIndex.from_tuples([(0,60), (60,120),(120,180), (180,240),(240,300)])
exercise_time_bins = pd.cut(persons_data['exercise_time'], bins_1)
bins_2 = pd.IntervalIndex.from_tuples([(100,130), (130, 158),(158, 188), (188, 217),(217,246)])
weight_bins = pd.cut(persons_data['weight'], bins_2)
bins_3 = pd.IntervalIndex.from_tuples([(18,30), (30,43),(43,55), (55,68),(68,80)])
age_bins = pd.cut(persons_data['age'], bins_3)
pd.crosstab(exercise_time_bins, [weight_bins, age_bins])
# + id="Wu4D1PqwGg6o" colab_type="code" outputId="c1908b60-3ac1-4f95-b033-0bd0b087ef18" colab={"base_uri": "https://localhost:8080/", "height": 303}
# Using normalize to get a cross-tabulation with percentages instead of frequencies
pd.crosstab(exercise_time_bins, [weight_bins, age_bins], normalize='columns')
# + id="TmY-YhW0F4u0" colab_type="code" outputId="c2d13078-8081-4804-cd79-0d0db51e4f3c" colab={"base_uri": "https://localhost:8080/", "height": 303}
# Using normalize to get a cross-tabulation with percentages instead of frequencies
ct = pd.crosstab(age_bins, [weight_bins, exercise_time_bins ], normalize='columns')
ct
# + id="RaOq5rKtF46I" colab_type="code" outputId="579c8c89-f34a-4a42-d02f-f179019ec2ad" colab={"base_uri": "https://localhost:8080/", "height": 235}
a_w = pd.crosstab(age_bins, weight_bins)
a_w
# + id="0ezcrT4jIPYm" colab_type="code" outputId="c7910db8-3f05-4e00-d994-6a29bcb792cd" colab={"base_uri": "https://localhost:8080/", "height": 235}
a_w_percentage = pd.crosstab(age_bins, weight_bins,normalize='columns')
a_w_percentage
# + id="xcvpD1HvJwoz" colab_type="code" outputId="31161fd1-0dcb-48d9-dff6-ec662b30fc14" colab={"base_uri": "https://localhost:8080/", "height": 290}
# plot 1
a_w_percentage.plot()
# + id="97C8FshfM-1b" colab_type="code" outputId="c61cc7a7-4bbb-45a9-fc12-4801b28dc075" colab={"base_uri": "https://localhost:8080/", "height": 329}
# plot 2
a_w_percentage.plot(kind='bar')
# + id="QPuljgMUF5Ev" colab_type="code" outputId="906fe783-94ac-4b71-fe5d-46463179fe07" colab={"base_uri": "https://localhost:8080/", "height": 235}
a_t = pd.crosstab(age_bins, exercise_time_bins)
a_t
# + id="76hjeiRuIlZ4" colab_type="code" outputId="320debfc-5bc5-4d9a-87e0-99c86999781b" colab={"base_uri": "https://localhost:8080/", "height": 235}
a_t_percentage = pd.crosstab(age_bins, exercise_time_bins,normalize='columns')
a_t_percentage
# + id="kYQFUc5aOgTu" colab_type="code" outputId="f18ff885-1a26-40d6-ade8-6f67d47cfcef" colab={"base_uri": "https://localhost:8080/", "height": 290}
# plot 3
a_t_percentage.plot()
# + id="5IEoZUa9QOtr" colab_type="code" outputId="ddcb9377-3c69-4961-fba1-0555266f2030" colab={"base_uri": "https://localhost:8080/", "height": 330}
# plot 4
a_t_percentage.plot(kind='bar')
# + id="8jpYRJ0z1d0S" colab_type="code" outputId="e2dfd808-ef23-4418-a765-1513eaeeaf40" colab={"base_uri": "https://localhost:8080/", "height": 235}
t_w = pd.crosstab(exercise_time_bins, weight_bins)
t_w
# + id="Td14ehazI0Wv" colab_type="code" outputId="ecd99b8f-967d-4389-c603-ddfff4f9bf33" colab={"base_uri": "https://localhost:8080/", "height": 235}
t_w_percentage = pd.crosstab(exercise_time_bins, weight_bins,normalize='columns')
t_w_percentage
# + id="UFY-3bfR0MWv" colab_type="code" outputId="33418485-ae93-4f8a-916e-24843c102744" colab={"base_uri": "https://localhost:8080/", "height": 291}
# plot 5
t_w_percentage.plot()
# + id="PPzJAJbURK6Q" colab_type="code" outputId="3e8dd932-b715-408c-a71c-7f6b406da4c5" colab={"base_uri": "https://localhost:8080/", "height": 343}
# plot 6
t_w_percentage.plot(kind='bar')
# + [markdown] id="BT9gdS7viJZa" colab_type="text"
# ### Assignment questions
#
# After you've worked on some code, answer the following questions in this text block:
#
# 1. What are the variable types in the data?
# 2. What are the relationships between the variables?
# 3. Which relationships are "real", and which spurious?
#
# + [markdown] id="YAU11uyrbG-6" colab_type="text"
# 1. In general age, weight and exercise time are continuous numerical data, but the data in persons.csv are discrete variables and integers.
# 2. As we can see at the plot 1, the weight curves are correlated until certain age, but as the age advances, the weight curves show different behavior of each other.
# The weight curves declines as the age advances for all intervals, except intervals (158, 188] and (217, 246], wich have ascending curves for advanced ages.
# At the plot 3, the exercise time curves in general declines as the age advances. For the intervals of exercise time (0, 60], (60, 120], (120, 180], (180, 240], the curves have similar shapes but they change after middle age. The curve for interval (0, 60] ascends when the age advances, meaning that older people make less exercises. The curves for intervals (180, 240] and (240, 300] have a sharp decrease from middle age, reinforcing the previous reasoning.
# The curve for the interval (240, 300] is ascending for young people and declines sharply from middle age, meaning that younger people are more active.
# Looking at the plot 6, the weight intervals (100, 130] and (130, 158] , the less heavy group of people, are in general well distributed between all the exercise time intervals, but the bars are slightly smaller in the exercise time interval (0, 60], showing that they are more active. On the other hand, the heavier group of people, that belongs to the weight intervals (188, 217] and (217, 246] are more concentrated in the exercise time interval (0, 60], indicating that these people make less exercises. Then, the data shows a relation between exercise time and weight: heavier the person, less exercise, and the opposite is true. One of the things that is not clear is the causality: if more exercises
#
# 3. The relationships between age and weight, and then age and exercise time seems real, even though for certain groups the relationship is different. The relationship between weight and exercise time is spurious because we don't have information about these people height (are they the same height?). Perhaps other variables are important too. And the causality: they are heavier because they make less exercises or they make less exercises because they are heavier?
# + [markdown] id="_XXg2crAipwP" colab_type="text"
# ## Stretch goals and resources
#
# Following are *optional* things for you to take a look at. Focus on the above assignment first, and make sure to commit and push your changes to GitHub.
#
# - [Spurious Correlations](http://tylervigen.com/spurious-correlations)
# - [NIH on controlling for confounding variables](https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4017459/)
#
# Stretch goals:
#
# - Produce your own plot inspired by the Spurious Correlation visualizations (and consider writing a blog post about it - both the content and how you made it)
# - Pick one of the techniques that NIH highlights for confounding variables - we'll be going into many of them later, but see if you can find which Python modules may help (hint - check scikit-learn)
| Marilyn_Esko_LS_DSPT3_113_Making_Data_backed_Assertions_Assignment_3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Amazon EC2 & EBS
# > Introduction to AWS EC2 & EBS
#
# - toc: true
# - comments: true
# - author: <NAME>
# - categories: [aws,EC2,EBS]
# ### Amazon Elastic Compute Cloud (Amazon EC2)
#
# #### Compute Basics
# Instance Types
# Sample Instance Family
# c4 Compute optimized—For workloads requiring significant processing
# r3 Memory optimized—For memory-intensive workloads
# i2 Storage optimized—For workloads requiring high amounts of fast SSD storage
# g2 GPU-based instances—Intended for graphics and general-purpose GPU compute workloads
#
# Enhanced Networking
# For workloads requiring greater network performance, many instance types support enhanced networking.
# Enhanced networking reduces the impact of virtualization on network performance by enabling a
# capability called Single Root I/O Virtualization (SR-IOV).
# This results in more Packets Per Second (PPS), lower latency, and less jitter.
#
# Four sources of AMIs
# Published by AWS
# The AWS Marketplace
# Generated from Existing Instances
# Uploaded Virtual Servers
# #### Securely Using an Instance
# Addressing an Instance
# Public Domain Name System (DNS) Name
# Public IP
# Elastic IP
#
# Initial Access
# Virtual Firewall Protection
# Type of Security Group Capabilities
# EC2-Classic Security Groups Control outgoing instance traffic
# VPC Security Groups Control outgoing and incoming instance traffic
# #### The Lifecycle of Instances
#
# Bootstraping
# One of the parameters when an instance is launched is a string value called UserData.
# This string is passed to the operating system to be executed as part of the launch process
# the first time the instance is booted. On Linux instances this can be shell script,
# and on Windows instances this can be a batch style script or a PowerShell script.
#
# VM Import/Export
# Instance Metadata
# http://169.254.169.254/latest/meta-data/
# Managing Instances
# Tags can help you manage not just your Amazon EC2 instances
# Monitoring Instances
# AWS offers a service called Amazon CloudWatch that provides monitoring and alerting
# Modifying an Instance
# Instance Type
# Instances can be resized using the AWS Management Console, CLI, or API
# Security Groups
# If an instance is running in an Amazon VPC, you can change which security groups
# are associated with an instance while the instance is running.
# For instances outside of an Amazon VPC (called EC2-Classic), the association of
# the security groups cannot be changed after launch.
#
# Termination Protection
# In order to prevent termination via the AWS Management Console, CLI, or API,
# termination protection can be enabled for an instance. While enabled, calls
# to terminate the instance will fail until termination protection is disabled.
# #### Options
# Pricing Options
# On-Demand Instances
# The price per hour for each instance type
# Reserved Instances
# The Reserved Instance pricing option enables customers to make capacity
# reservations for predictable workloads.
# Spot Instances
# For workloads that are not time critical and are tolerant of interruption,
# Spot Instances offer the greatest discount.
#
# Tenancy Options
# Shared Tenancy
# Shared tenancy is the default tenancy model for all Amazon EC2 instances, regardless
# of instance type, pricing model, and so forth. Shared tenancy means that a single
# host machine may house instances from different customers. As AWS does not use
# overprovisioning and fully isolates instances from other instances on the same host,
# this is a secure tenancy model.
#
# Dedicated Instances
# Dedicated Instances run on hardware that’s dedicated to a single customer.
# As a customer runs more Dedicated Instances, more underlying hardware may be dedicated
# to their account. Other instances in the account (those not designated as dedicated)
# will run on shared tenancy and will be isolated at the hardware level from the
# Dedicated Instances in the account.
#
# Dedicated Host
# An Amazon EC2 Dedicated Host is a physical server with Amazon EC2 instance capacity fully
# dedicated to a single customer’s use. Dedicated Hosts can help you address licensing
# requirements and reduce costs by allowing you to use your existing server-bound
# software licenses. The customer has complete control over which specific host runs
# an instance at launch. This differs from Dedicated Instances in that a Dedicated
# Instance can launch on any hardware that has been dedicated to the account.
#
# Placement Groups
# A placement group is a logical grouping of instances within a single Availability Zone
#
# Instance Stores
# An instance store (sometimes referred to as ephemeral storage) provides temporary block-level
# storage for your instance.
# ### Amazon Elastic Block Store (Amazon EBS)
#
# Elastic Block Store Basics
# Each Amazon EBS volume is automatically replicated within its Availability Zone to protect
# you from component failure, offering high availability and durability
# Multiple Amazon EBS volumes can be attached to a single Amazon EC2 instance,
# although a volume can only be attached to a single instance at a time.
#
# Types of Amazon EBS Volumes
# Magnetic Volumes
# Magnetic volumes have the lowest performance characteristics of all Amazon EBS volume types.
# As such, they cost the lowest per gigabyte.
# They are an excellent, cost-effective solution for appropriate workloads.
# A magnetic Amazon EBS volume can range in size from 1 GB to 1 TB and will average 100 IOPS,
# but has the ability to burst to hundreds of IOPS.
# Cold workloads where data is infrequently accessed.
#
# General-Purpose SSD
# General-purpose SSD volumes offer cost-effective storage that is ideal for a broad range
# of workloads.
# They deliver strong performance at a moderate price point that is suitable for a wide
# range of workloads.
# A general-purpose SSD volume can range in size from 1 GB to 16 TB and provides a
# baseline performance of three IOPS per gigabyte provisioned, capping at 10,000 IOPS
#
# Provisioned IOPS SSD
# Provisioned IOPS SSD volumes are designed to meet the needs of I/O-intensive workloads,
# particularly database workloads that are sensitive to storage performance and
# consistency in random access I/O throughput.
# While they are the most expensive Amazon EBS volume type per gigabyte, they provide the
# highest performance of any Amazon EBS volume type in a predictable manner.
# A Provisioned IOPS SSD volume can range in size from 4 GB to 16 TB.
# When you provision a Provisioned IOPS SSD volume, you specify not just the size,
# but also the desired number of IOPS, up to the lower of the maximum of 30 times
# the number of GB of the volume, or 20,000 IOPS
#
# Throughput-Optimized
# HDD volumes are low-cost HDD volumes designed for frequent-access, throughput-intensive
# workloads such as big data, data warehouses, and log processing.
# Volumes can be up to 16 TB with a maximum IOPS of 500 and maximum throughput of 500 MB/s.
# These volumes are significantly less expensive than general-purpose SSD volumes.
#
# Cold HDD
# Volumes are designed for less frequently accessed workloads, such as colder data requiring
# fewer scans per day.
# Volumes can be up to 16 TB with a maximum IOPS of 250 and maximum throughput of 250 MB/s.
# These volumes are significantly less expensive than Throughput-Optimized HDD volumes.
# #### Protecting Data
# Backup/Recovery (Snapshots)
# Snapshots are incremental backups, which means that only the blocks on the device that have
# changed since your most recent snapshot are saved.
#
# Taking Snapshots
# Through the AWS Management Console
# Through the CLI
# Through the API
# By setting up a schedule of regular snapshots
#
# Snapshots are constrained to the region in which they are created, meaning you can use them to
# create new volumes only in the same region.
# If you need to restore a snapshot in a different region, you can copy a snapshot to another region.
#
# Creating a Volume from a Snapshot
# Recovering Volumes
# Encryption Options
| _notebooks/2020-06-21-Amazon Elastic Compute Cloud (EC2) and Amazon Elastic Block Store (EBS).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import tensorflow as tf
import tensorboard_jupyter as tb
import matplotlib.pyplot as plt
sess = tf.Session()
# 0を目的値として-1から1までの値を作成する
x_vals = tf.linspace(-1., 1., 500)
target = tf.constant(0.)
## 回帰のための損失関数 ##
# L2ノルムの損失関数。二乗誤差。
l2_y_vals = tf.square(target - x_vals)
l2_y_out = sess.run(l2_y_vals)
# tf.nn.l2_lossは上記の半分の値となる
# L1ノルムの損失関数。差の絶対値。
# 外れ値にはうまく対応するが、目的値に対してなめらかでないためアルゴリズムが収束しないことがある
l1_y_vals = tf.abs(target - x_vals)
l1_y_out = sess.run(l1_y_vals)
# Pseudo-Huber損失関数。L1とL2の長所を活かした関数。
delta1 = tf.constant(0.25)
phuber1_y_vals = tf.multiply(tf.square(delta1),
tf.sqrt(1. + tf.square((target - x_vals)/delta1)) - 1.)
phuber1_y_out = sess.run(phuber1_y_vals)
delta2 = tf.constant(5.)
phuber2_y_vals = tf.multiply(tf.square(delta2),
tf.sqrt(1. + tf.square((target - x_vals)/delta2)) - 1.)
phuber2_y_out = sess.run(phuber2_y_vals)
## 分類のための損失関数 ##
# 1を目的値として-3から5までの値を作成する
x_vals = tf.linspace(-3., 5., 500)
target = tf.constant(1.)
targets = tf.fill([500], 1.)
# ヒンジ損失関数。主にSVMで使用される
# 正解値からの距離を表す
hinge_y_vals = tf.maximum(0., 1. - tf.multiply(target, x_vals))
hinge_y_out = sess.run(hinge_y_vals)
# 交差エントロピー損失関数(ロジスティック関数)
xentropy_y_vals = - tf.multiply(target, tf.log(x_vals)) - tf.multiply((1. - target), tf.log(1. - x_vals))
xentropy_y_out = sess.run(xentropy_y_vals)
# シグモイド交差エントロピー損失関数
xentropy_sigmoid_y_vals = tf.nn.sigmoid_cross_entropy_with_logits(logits=x_vals, labels=targets)
xentropy_sigmoid_y_out = sess.run(xentropy_sigmoid_y_vals)
# 重み付き交差エントロピー誤差関数
weight = tf.constant(0.5)
xentropy_weighted_y_vals = tf.nn.weighted_cross_entropy_with_logits(x_vals, targets, weight)
xentropy_weighted_y_out = sess.run(xentropy_weighted_y_vals)
# ソフトマックス交差エントロピー誤差関数
unscaled_logits = tf.constant([[1., -3., 10.]])
target_dist = tf.constant([[0.1, 0.02, 0.88]])
softmax_entropy = tf.nn.softmax_cross_entropy_with_logits(logits=unscaled_logits, labels=target_dist)
print('ソフトマックス交差エントロピー誤差関数:', sess.run(softmax_entropy))
x_array = sess.run(x_vals)
plt.plot(x_array, l2_y_out, 'b-', label='L2 Loss')
plt.plot(x_array, l1_y_out, 'r--', label='L1 Loss')
plt.plot(x_array, phuber1_y_out, 'k-', label='phuber Loss(0.25)')
plt.plot(x_array, phuber2_y_out, 'g:', label='phuber Loss(0.5)')
plt.legend(loc='lower right', prop={'size': 11})
plt.show()
plt.plot(x_array, hinge_y_out, 'b-', label='Hinge Loss')
plt.plot(x_array, xentropy_y_out, 'r--', label='Cross Entropy Loss')
plt.plot(x_array, xentropy_sigmoid_y_out, 'k-', label='Cross Entropy Sigmoid Loss')
plt.plot(x_array, xentropy_weighted_y_out, 'g:', label='Weighted Cross Entropy Loss')
plt.legend(loc='lower right', prop={'size': 11})
plt.show()
tf.summary.FileWriter('./log/', sess.graph)
tb.show_graph(tf.get_default_graph().as_graph_def())
# -
| tensorflow-cookbook-keras/ch02/2.5_loss_function.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.4.1
# language: julia
# name: julia-1.4
# ---
import Plots: plot
function expr_to_function_and_string(e::Expr)::Tuple{Expr, String}
(
:(x -> $e),
String(Symbol(e)),
)
end
let e = :(x + sin(x^2))
expr_to_function_and_string(e)
end
macro curve(e, from, to)
@assert isa(from, Number)
@assert isa(to, Number)
func_expr, expr_str = expr_to_function_and_string(e)
quote
plot(
range($from, $to, length = 1000),
$func_expr,
xlabel="x",
ylabel=$expr_str,
leg=false,
)
end
end
@curve(x + sin(x^2), -5.0, 5.0)
@curve(sin(x^2) + x, -5.0, 5.0)
| Curve Macro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R (system-wide)
# language: r
# metadata:
# cocalc:
# description: R statistical programming language
# priority: 10
# url: https://www.r-project.org/
# name: ir
# resource_dir: /ext/jupyter/kernels/ir
# ---
# # Section 4 Practice
#
# Here are some additional practice and examples for you to try out!
| _ipynb_checkpoints/Section4Practice-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# +
import string
from random import *
import os
import glob
import face_recognition
# +
import speech_recognition as sr
from google.cloud import speech
import io
import os
#######################
GOOGLE_CLOUD_SPEECH_CREDENTIALS_PATH = '../files/TFM project-287dc6d9869a.json'
#######################
def transcript_audio(filepath, language, use_cloud):
transcript = '##NONE##'
# The name of the audio file to transcribe
file_name = os.path.join(os.path.dirname(''), filepath)
if use_cloud:
try:
# Instantiates a client
speech_client = speech.Client.from_service_account_json(GOOGLE_CLOUD_SPEECH_CREDENTIALS_PATH)
# Loads the audio into memory
with io.open(file_name, 'rb') as audio_file:
content = audio_file.read()
sample = speech_client.sample(
content,
source_uri=None,
encoding='LINEAR16',
sample_rate_hertz=16000)
# Detects speech in the audio file
alternatives = sample.recognize(language)
if (len(alternatives)>0):
transcript = alternatives[0].transcript
except Exception as e:
print(e)
if (transcript == '##NONE##'):
try:
r = sr.Recognizer()
with sr.AudioFile(file_name) as source:
audio = r.record(source)
# for testing purposes, we're just using the default API key
# to use another API key, use `r.recognize_google(audio, key="GOOGLE_SPEECH_RECOGNITION_API_KEY", show_all=True)`
# instead of `r.recognize_google(audio, show_all=True)`
alternatives = r.recognize_google(audio, show_all=False)
if (len(alternatives)>0):
transcript = alternatives
except sr.UnknownValueError:
print("Google Speech Recognition could not understand audio")
except sr.RequestError as e:
print("Could not request results from Google Speech Recognition service; {0}".format(e))
return transcript
# +
# Audio Play
import pyaudio
import wave
import time
import sys
import pygame as pg
def play_music(music_file, volume=0.8):
'''
stream music with mixer.music module in a blocking manner
this will stream the sound from disk while playing
'''
# set up the mixer
freq = 44100 # audio CD quality
bitsize = -16 # unsigned 16 bit
channels = 2 # 1 is mono, 2 is stereo
buffer = 2048 # number of samples (experiment to get best sound)
pg.mixer.init()
# volume value 0.0 to 1.0
pg.mixer.music.set_volume(volume)
clock = pg.time.Clock()
try:
pg.mixer.music.load(music_file)
print("Music file {} loaded!".format(music_file))
except pg.error:
print("File {} not found! ({})".format(music_file, pg.get_error()))
return
pg.mixer.music.play()
while pg.mixer.music.get_busy():
# check if playback has finished
clock.tick(30)
def play_any_audio(filename):
pg.mixer.init()
pg.mixer.music.load(filename)
pg.mixer.music.play()
def play_audio(filename):
WAVE_FILENAME = filename
if len(sys.argv) < 2:
print("Plays a wave file.\n\nUsage: %s filename.wav" % WAVE_FILENAME)
sys.exit(-1)
wf = wave.open(WAVE_FILENAME, 'rb')
p = pyaudio.PyAudio()
def callback(in_data, frame_count, time_info, status):
data = wf.readframes(frame_count)
return (data, pyaudio.paContinue)
stream = p.open(format=p.get_format_from_width(wf.getsampwidth()),
channels=wf.getnchannels(),
rate=wf.getframerate(),
output=True,
stream_callback=callback)
stream.start_stream()
while stream.is_active():
time.sleep(0.1)
stream.stop_stream()
stream.close()
wf.close()
p.terminate()
def record_audio(filename, seconds):
CHUNK = 1024
FORMAT = pyaudio.paInt16
#CHANNELS = 2
CHANNELS = 1
#RATE = 44100
RATE = 16000
p = pyaudio.PyAudio()
stream = p.open(format=FORMAT,
channels=CHANNELS,
rate=RATE,
input=True,
frames_per_buffer=CHUNK)
print("* recording")
frames = []
for i in range(0, int(RATE / CHUNK * seconds)):
data = stream.read(CHUNK)
frames.append(data)
print("* done recording")
stream.stop_stream()
stream.close()
p.terminate()
wf = wave.open(filename, 'wb')
wf.setnchannels(CHANNELS)
wf.setsampwidth(p.get_sample_size(FORMAT))
wf.setframerate(RATE)
wf.writeframes(b''.join(frames))
wf.close()
# +
import gettext
_ = lambda s: s
es_man = gettext.translation('text_man', localedir=LANGUAGE_PATH, languages=['es'])
es_man.install()
print(_('surprise'))
es_woman = gettext.translation('text_woman', localedir=LANGUAGE_PATH, languages=['es'])
es_woman.install()
print(_('surprise'))
es_woman = gettext.translation('text_', localedir=LANGUAGE_PATH, languages=['en'])
es_woman.install()
print(_('surprise'))
# +
import gettext
es_man = gettext.translation('text_man', localedir=LANGUAGE_PATH, languages=['es'])
es_man.install()
print(_('surprise'))
es_woman = gettext.translation('text_woman', localedir=LANGUAGE_PATH, languages=['es'])
es_woman.install()
print(_('surprise'))
es_woman = gettext.translation('text_', localedir=LANGUAGE_PATH, languages=['en'])
es_woman.install()
print(_('surprise'))
LANGUAGE_PATH = '../lang/'
LANG = 'es'
LANGUAGE = LANG + '/'
AUDIO_PATH = '../audio/'
KNOWN = 'known/'
IMAGE_PATH = '../images/'
def get_language_audios(path, audios, preds):
lang_audios = []
for audio in audios:
audio_path = path + audio
for pred in preds:
audio_path = audio_path.replace('['+pred+']', preds[pred])
lang_audios.append(audio_path)
return lang_audios
def get_formatted_language_audios(path, audios, predictions):
lang_audios = []
try:
for prediction in predictions:
print(prediction)
for audio in audios:
print(audio)
key = audio.split(':')[0]
if (key == 'GENDER' and prediction['NAME_AUDIO'] != ''):
audio_path = AUDIO_PATH + KNOWN + prediction['NAME_AUDIO']
lang_audios.append(audio_path)
else:
audio_path = path + audio.split(':')[1]
for key in prediction:
audio_path = audio_path.replace('['+key+']', prediction[key])
lang_audios.append(audio_path)
except Exception as e:
print('*a******')
print(e)
print('*a******')
return lang_audios
def get_formatted_language_text(language, prediction):
print(prediction)
lang_text = ''
try:
text_config = ''
with open(LANGUAGE_PATH + language + '/text_config.txt') as f:
for line in f:
text_config += line.rstrip()
print('1')
g = text_config.split(':')[0]
lang_text = text_config.split(':')[1]
for key in prediction:
g = g.replace('['+key+']', prediction[key])
print('2')
l = gettext.translation('text_' + g, localedir=LANGUAGE_PATH, languages=[language])
l.install()
print('****')
print('text_' + g)
print(LANGUAGE_PATH)
print(language)
print('****')
print('3')
t = ''
if (prediction['NAME'] != ''):
print('3a')
t = prediction['NAME']
else:
print('3b')
if(prediction['GENDER'] != ''):
print('3c')
print(prediction['GENDER'])
print('3d')
t = _(prediction['GENDER'])
print('3e')
print('4')
lang_text = lang_text.replace('[GENDER]', t)
print('5')
t = ''
if(prediction['EMOTION'] != ''):
t = _(prediction['EMOTION'])
lang_text = lang_text.replace('[EMOTION]', t)
print('6')
except Exception as e:
print('*t******')
print(e)
print('*t******')
return lang_text
config_audios = []
with open(LANGUAGE_PATH+LANGUAGE+'audio_config.txt') as f:
for line in f:
config_audios.append(line.rstrip())
#print(line)
label_dict = {'EMOTION': '', 'GENDER': '', 'NAME': '', 'NAME_AUDIO': ''}
pred_test = label_dict.copy();
pred_test['EMOTION'] = 'angry'
pred_test['GENDER'] = 'man'
pred_test['NAME'] = ''
pred_test['NAME_AUDIO'] = ''
text = get_formatted_language_text('es', pred_test)
print(text)
# +
import cv2
from keras.models import load_model
import numpy as np
from statistics import mode
from utils import preprocess_input
from utils import get_labels
# parameters
detection_model_path = '../models/face/haarcascade_frontalface_default.xml'
emotion_model_path = '../models/emotion/simple_CNN.530-0.65.hdf5'
gender_model_path = '../models/gender/simple_CNN.81-0.96.hdf5'
emotion_labels = get_labels('fer2013')
gender_labels = get_labels('imdb')
frame_window = 10
x_offset_emotion = 20
y_offset_emotion = 40
x_offset = 30
y_offset = 60
# loading models
face_detection = cv2.CascadeClassifier(detection_model_path)
emotion_classifier = load_model(emotion_model_path)
gender_classifier = load_model(gender_model_path)
# +
known_faces = []
for filepath in glob.iglob('../images/known/*.*', recursive=True):
filename = os.path.splitext(os.path.basename(filepath))[0]+'.mp3'
name = os.path.splitext(filename)[0].split('-')[0]
picture = face_recognition.load_image_file(filepath)
encoding = face_recognition.face_encodings(picture)[0]
known_faces.append([name, filename, encoding])
for i in range(len(known_faces)):
print(known_faces[i][0])
print(known_faces[i][1])
#print(known_faces[i][2])
# +
from gtts import gTTS
import os
from unidecode import unidecode
def capture_face_and_name(face):
rand = "".join(choice(string.ascii_letters) for x in range(randint(8, 8)))
name = "".join(choice(string.ascii_letters) for x in range(randint(6, 6)))
temp_wav = AUDIO_PATH + KNOWN + name + '-' + rand + '.wav'
#Play beep
play_music(AUDIO_PATH + 'beep.mp3')
#Record audio
record_audio(temp_wav, 2)
play_music(LANGUAGE_PATH + LANGUAGE + 'speech/one_moment.mp3')
#Transcript audio
transcript = transcript_audio(temp_wav, LANG, True)
print('***'+transcript+'***')
#if transcript didn't capture anything then exit
if (transcript == '' or transcript == '##NONE##'):
#Delete wav file
os.remove(temp_wav)
return None, transcript, None
#if transcript captures cancelation then cancel
elif (transcript.lower() == 'cancel' or transcript.lower() =='cancelar'):
#TODO: make it generic for any language
play_music(LANGUAGE_PATH + LANGUAGE + 'speech/canceled.mp3')
os.remove(temp_wav)
return None, 'canceled', None
#if transcript ok then proceed
else:
plain_transcript = unidecode(transcript)
mp3_name = plain_transcript + '-' + rand + '.mp3'
temp_mp3 = AUDIO_PATH + KNOWN + mp3_name
#Convert transcript to standard audio
tts = gTTS(text=transcript, lang=LANG, slow=False)
tts.save(temp_mp3)
#Play audio back
play_music(temp_mp3)
play_music(LANGUAGE_PATH + LANGUAGE + 'speech/saved.mp3')
#Delete wav file
os.remove(temp_wav)
#Save face image
face_img = IMAGE_PATH + KNOWN + plain_transcript + '-' + rand + '.jpg'
print(face_img)
cv2.imshow('image',face)
cv2.imwrite(face_img, face)
#Get face encoding
picture = face_recognition.load_image_file(face_img)
face_encoding = face_recognition.face_encodings(picture)[0]
print('---')
print (face_encoding)
print (plain_transcript)
print (mp3_name)
print('---')
return face_encoding, plain_transcript, mp3_name
# +
# video
video_capture = cv2.VideoCapture(0)
font = cv2.FONT_HERSHEY_SIMPLEX
cv2.namedWindow('window_frame')
emotion_label_window = []
gender_label_window = []
last_faces = []
ENCODING_FREQ = 10
encoding_count = 0
last_faces_count = 0
face_encodings = []
predictions = []
while True:
predictions = []
encoding_count += 1
last_faces_count = len(last_faces)
last_faces = []
_, frame = video_capture.read()
frame_ = frame.copy()
gray = cv2.cvtColor(frame, cv2.COLOR_BGR2GRAY)
frame = cv2.cvtColor(frame, cv2.COLOR_BGR2RGB)
faces = face_detection.detectMultiScale(gray, 1.3, 5)
do_encode = encoding_count>=ENCODING_FREQ | last_faces_count!=len(faces)
if (do_encode):
face_encodings = []
face_index = 0
for (x,y,w,h) in faces:
pred_dict = label_dict.copy();
face_index +=1
face = frame[(y - y_offset):(y + h + y_offset),
(x - x_offset):(x + w + x_offset)]
if (do_encode):
print('re-encoding')
face_encodings.append(face_recognition.face_encodings(frame, [tuple([int(y), int(x+w), int(y+h), int(x)])])[0])
encoding_count = 0
try:
if (len(face_encodings)>0 & face_index -1 < len(face_encodings)):
for i in range(len(known_faces)):
match = face_recognition.compare_faces([known_faces[i][2]], face_encodings[face_index-1])
if match[0]:
pred_dict['NAME'] = known_faces[i][0]
pred_dict['NAME_AUDIO'] = known_faces[i][1]
break;
except Exception as e:
print('*******')
print(e)
print(len(face_encodings))
print(face_index)
print('*******')
continue
#print('-----')
last_faces.append(cv2.cvtColor(face.copy(), cv2.COLOR_RGB2BGR))
gray_face = gray[(y - y_offset_emotion):(y + h + y_offset_emotion),
(x - x_offset_emotion):(x + w + x_offset_emotion)]
try:
face = cv2.resize(face, (48, 48))
gray_face = cv2.resize(gray_face, (48, 48))
except:
continue
face = np.expand_dims(face, 0)
face = preprocess_input(face)
gender_label_arg = np.argmax(gender_classifier.predict(face))
gender = gender_labels[gender_label_arg]
gender_label_window.append(gender)
gray_face = preprocess_input(gray_face)
gray_face = np.expand_dims(gray_face, 0)
gray_face = np.expand_dims(gray_face, -1)
emotion_label_arg = np.argmax(emotion_classifier.predict(gray_face))
emotion = emotion_labels[emotion_label_arg]
emotion_label_window.append(emotion)
if len(gender_label_window) >= frame_window:
emotion_label_window.pop(0)
gender_label_window.pop(0)
try:
emotion_mode = mode(emotion_label_window)
gender_mode = mode(gender_label_window)
except:
continue
if gender_mode == gender_labels[0]:
gender_color = (255, 0, 0)
else:
gender_color = (0, 255, 0)
pred_dict['EMOTION'] = emotion_mode
pred_dict['GENDER'] = gender_mode
display_text = get_formatted_language_text(LANG, pred_dict)
cv2.rectangle(frame, (x, y), (x + w, y + h), gender_color, 2)
cv2.putText(frame, display_text, (x, y - 30), font,
.7, gender_color, 1, cv2.LINE_AA)
#cv2.putText(frame, display_name, (x + 90, y - 30), font,
# .7, gender_color, 1, cv2.LINE_AA)
predictions.append(pred_dict)
try:
frame = cv2.cvtColor(frame, cv2.COLOR_RGB2BGR)
cv2.imshow('window_frame', frame)
except:
continue
c = chr(cv2.waitKey(2)& 255)
if (c!= 'ÿ'):
print(c + " pressed")
if (c=='l' or c=='L'):
print('*** Language change *** ')
if (LANG == 'es'):
LANG = 'en'
else:
LANG = 'es'
if (c=='a' or c=='A'):
print('*** Output predictions selected *** ')
lang_audios = get_formatted_language_audios(LANGUAGE_PATH + LANGUAGE, config_audios, predictions)
for lang_audio in lang_audios:
print(lang_audio)
play_music(lang_audio)
if (c == 's' or c=='S'):
print('*** Save person selected *** ')
try:
if (len(last_faces)==1):
name = '##NONE##'
while name == '##NONE##':
play_music(LANGUAGE_PATH + LANGUAGE + 'speech/who.mp3')
if cv2.waitKey(1) & 0xFF == ord('q'):
break
else:
print('START')
face_encoding, name, audio_file_name = capture_face_and_name(last_faces[0])
print('END')
print(name)
print(audio_file_name)
print(face_encoding)
print('END2')
if (name=='##NONE##'):
play_music(LANGUAGE_PATH + LANGUAGE + 'speech/not_understand.mp3')
elif (name == 'canceled'):
break
else:
print('appending')
known_faces.append([name, audio_file_name, face_encoding])
for i in range(len(known_faces)):
print(known_faces[i][0])
print(known_faces[i][1])
print(known_faces[i][2])
break
else:
play_music(LANGUAGE_PATH + LANGUAGE + 'speech/more_than_one_face.mp3')
except:
continue
if c == 'q':
break
video_capture.release()
cv2.destroyAllWindows()
# -
| src/.ipynb_checkpoints/recog name with audio-checkpoint-Josu-PC.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### 1. What protocols are required for Internet communication?
# ##### Ans: TCP or UDP/IP
# #### 2. Which network device copies packets onto all ports?
# ##### Ans: hub
# #### 3. What is the protocol associated with the world wide web?
# ##### Ans: HTTP
# #### 4. A MANET protocol is likely to differ from a typical LAN protocol because a MANET protocol will
# ##### Ans: consume less power than a typical LAN protocol.
# #### 5. The function of a packet header is to
# ##### Ans: contain packet information generated by a protocol layer.
# #### 6. A packet sniffer is a tool that can be used to record all local traffic on a network.
# ##### Ans: True
# #### 7. A hub can be used to communicate between two LANs with different protocols.
# ##### Ans: False
# #### 8. In the HTTP protocol, a request message is sent by a web client to a web server.
# ##### Ans: True
| Coursera/Introduction to the Internet of Things and Embedded Systems/Week-4/Quiz/Module-4-Quiz.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:new]
# language: python
# name: conda-env-new-py
# ---
import pylab
import pymoc
import xidplus
import numpy as np
# %matplotlib inline
# This notebook uses all the raw data from the masterlist, maps, PSF and relevant MOCs to create XID+ prior object and relevant tiling scheme
# ## Read in MOCs
# The selection functions required are the main MOC associated with the masterlist. As the prior for XID+ is based on IRAC detected sources coming from two different surveys at different depths (SERVS and SWIRE) I will split the XID+ run into two different runs. Here we use the SERVS depth.
Sel_func=pymoc.MOC()
Sel_func.read('../data/Lockman-SWIRE/holes_Lockman-SWIRE_irac1_O16_MOC.fits')
# ## Read in Masterlist
# Next step is to read in Masterlist and select only sources that are detected in mid-infrared and at least one other wavelength domain (i.e. optical or nir). This will remove most of the objects in the catalogue that are artefacts. We can do this by using the `flag_optnir_det` flag and selecting sources that have a binary value of $>= 5$
# +
from astropy.io import fits
masterlist=fits.open('../data/Lockman-SWIRE/master_catalogue_lockman-swire_20171129.fits')
# -
good=masterlist[1].data['flag_optnir_det']>=5
# ### Create uninformative (i.e. conservative) upper and lower limits based on IRAC fluxes
# As the default flux prior for XID+ is a uniform distribution, it makes sense to set reasonable upper and lower 24 micron flux limits based on the longest wavelength IRAC flux available. For a lower limit I take IRAC/500.0 and for upper limit I take IRACx500.
MIPS_lower=np.full(good.sum(),0.0)
MIPS_upper=np.full(good.sum(),1E5)
for i in range(250000,good.sum()):
if masterlist[1].data['f_irac_i4'][good][i]>0:
MIPS_lower[i]=masterlist[1].data['f_irac_i4'][good][i]/500.0
MIPS_upper[i]=masterlist[1].data['f_irac_i4'][good][i]*500.0
elif masterlist[1].data['f_irac_i3'][good][i]>0:
MIPS_lower[i]=masterlist[1].data['f_irac_i3'][good][i]/500.0
MIPS_upper[i]=masterlist[1].data['f_irac_i3'][good][i]*500.0
elif masterlist[1].data['f_irac_i2'][good][i]>0:
MIPS_lower[i]=masterlist[1].data['f_irac_i2'][good][i]/500.0
MIPS_upper[i]=masterlist[1].data['f_irac_i2'][good][i]*500.0
elif masterlist[1].data['f_irac_i1'][good][i]>0:
MIPS_lower[i]=masterlist[1].data['f_irac_i1'][good][i]/500.0
MIPS_upper[i]=masterlist[1].data['f_irac_i1'][good][i]*500.0
MIPS_Map=fits.open('../data/Lockman-SWIRE/wp4_lockman-swire_mips24_map_v1.0.fits.gz')
# ## Read in PSF
MIPS_psf=fits.open('../../dmu17/dmu17_Lockman-SWIRE/dmu17_MIPS_Lockman-SWIRE_20171122.fits')
centre=np.long((MIPS_psf[1].header['NAXIS1']-1)/2)
radius=20
import pylab as plt
plt.imshow(np.log10(MIPS_psf[1].data[centre-radius:centre+radius+1,centre-radius:centre+radius+1]/np.max(MIPS_psf[1].data[centre-radius:centre+radius+1,centre-radius:centre+radius+1])))
plt.colorbar()
# ## Set XID+ prior class
prior_MIPS=xidplus.prior(MIPS_Map[1].data,MIPS_Map[2].data,MIPS_Map[0].header,MIPS_Map[1].header,moc=Sel_func)
prior_MIPS.prior_cat(masterlist[1].data['ra'][good],masterlist[1].data['dec'][good],'master_catalogue_lockman-swire_20170817.fits',flux_lower=MIPS_lower,
flux_upper=MIPS_upper,ID=masterlist[1].data['help_id'][good])
prior_MIPS.set_prf(MIPS_psf[1].data[centre-radius:centre+radius+1,centre-radius:centre+radius+1]/1.0E6,np.arange(0,41/2.0,0.5),np.arange(0,41/2.0,0.5))
# ## Calculate tiles
# As fitting the whole map would be too computationally expensive, I split based on HEALPix pixels. For MIPS, the optimum order is 11. So that I don't have to read the master prior based on the whole map into memory each time (which requires a lot more memory) I also create another layer of HEALPix pixels based at the lower order of 7.
import pickle
#from moc, get healpix pixels at a given order
from xidplus import moc_routines
order=11
tiles=moc_routines.get_HEALPix_pixels(order,prior_MIPS.sra,prior_MIPS.sdec,unique=True)
order_large=7
tiles_large=moc_routines.get_HEALPix_pixels(order_large,prior_MIPS.sra,prior_MIPS.sdec,unique=True)
print('----- There are '+str(len(tiles))+' tiles required for input catalogue and '+str(len(tiles_large))+' large tiles')
output_folder=''
outfile=output_folder+'Master_prior.pkl'
with open(outfile, 'wb') as f:
pickle.dump({'priors':[prior_MIPS],'tiles':tiles,'order':order,'version':xidplus.io.git_version()},f)
outfile=output_folder+'Tiles.pkl'
with open(outfile, 'wb') as f:
pickle.dump({'tiles':tiles,'order':order,'tiles_large':tiles_large,'order_large':order_large,'version':xidplus.io.git_version()},f)
raise SystemExit()
with open('Master_prior.pkl', 'wb') as f:
pickle.dump({'priors':prior_MIPS,'tiles':tiles,'order':order,'version':xidplus.io.git_version()},f)
xidplus.io.git_version()
# ## Large pickle problem
# It would appear that if the file trying to be saved is too big, then pickle will fail. To get round this, use the following fix:
import copy
# +
class MacOSFile(object):
def __init__(self, f):
self.f = f
def __getattr__(self, item):
return getattr(self.f, item)
def read(self, n):
# print("reading total_bytes=%s" % n, flush=True)
if n >= (1 << 31):
buffer = bytearray(n)
idx = 0
while idx < n:
batch_size = min(n - idx, 1 << 31 - 1)
# print("reading bytes [%s,%s)..." % (idx, idx + batch_size), end="", flush=True)
buffer[idx:idx + batch_size] = self.f.read(batch_size)
# print("done.", flush=True)
idx += batch_size
return buffer
return self.f.read(n)
def write(self, buffer):
n = len(buffer)
print("writing total_bytes=%s..." % n, flush=True)
idx = 0
while idx < n:
batch_size = min(n - idx, 1 << 31 - 1)
print("writing bytes [%s, %s)... " % (idx, idx + batch_size), end="", flush=True)
self.f.write(buffer[idx:idx + batch_size])
print("done.", flush=True)
idx += batch_size
def pickle_dump(obj, file_path):
with open(file_path, "wb") as f:
return pickle.dump(obj, MacOSFile(f), protocol=pickle.HIGHEST_PROTOCOL)
def pickle_load(file_path):
with open(file_path, "rb") as f:
return pickle.load(MacOSFile(f))
# -
pickle_dump({'priors':[prior_MIPS],'tiles':tiles,'order':order,'version':xidplus.io.git_version()},'Master_prior.pkl')
| dmu26/dmu26_XID+MIPS_Lockman-SWIRE/XID+MIPS_prior.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Dmitri9149/TensorFlow_PyTorch_CNN/blob/main/PyTorch_LeNet_AvgPooling%2BSigmoid.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="MG1qzYNQNQKk" outputId="02598239-d105-421e-8308-1408d2223cd3"
# -U: Upgrade all packages to the newest available version
# !pip install -U d2l
from d2l import torch as d2l
import torch
from torch import nn
import numpy as np
import torch.nn.functional as F
# + [markdown] id="iXKcpvT0-wzm"
# ### Convolutional Neural Network LeNet.
#
# This is implementation of classical LeNet convolutional neural network , originally designed for handwritten digit recignition.
#
# The basic architecture is used for some experimentation:
# we may change AveragePooling to MaxPooling and Sigmoid to ReLu activations. It is interesting to check, how it will change the results.
#
# I use some code from d2l.ai : http://d2l.ai/
#
# There is also some intermediate code with experimentation with PyTorch objects.
# + [markdown] id="OAUUcE93H87E"
#
# + [markdown] id="T32WFBjZYnKt"
# ## In this version we use Average Pooling and Sigmoid activation.
# + id="wD8d2Mtcp2lo" colab={"base_uri": "https://localhost:8080/"} outputId="a1df3fcc-d5cc-4334-c580-45374dca6a54"
## assume the input image is (n,m) matrixes
class Reshape(torch.nn.Module):
def __init__(self,rows,cols):
super().__init__()
self.rows=torch.tensor(rows)
self.cols=torch.tensor(cols)
def forward(self,x):
return x.view(-1,1,self.rows,self.cols)
reshape = Reshape(rows=28,cols=28)
print(reshape.rows,reshape.cols)
# + colab={"base_uri": "https://localhost:8080/"} id="YF48taM8xdhI" outputId="24eae152-f6ed-40bc-ee42-d106a11ba06b"
class LN(nn.Module):
def __init__(self):
super(LN, self).__init__()
self.reshape = Reshape(rows=28,cols=28)
self.conv1 = nn.Conv2d(1,6,kernel_size=5, padding=2)
self.sigmoid = nn.Sigmoid()
self.relu=nn.ReLU()
self.conv2 = nn.Conv2d(6,16,kernel_size=5)
self.avgpool1=nn.AvgPool2d(kernel_size=2, stride=2)
self.maxpool1=nn.MaxPool2d(kernel_size = 2, stride = 2)
self.avgpool2=nn.AvgPool2d(kernel_size=2, stride=2)
self.maxpool2=nn.MaxPool2d(kernel_size=2, stride=2)
self.flatten=nn.Flatten()
self.fc1 = nn.Linear(16 * 5 * 5, 120)
self.fc2 = nn.Linear(120, 84)
self.fc3 = nn.Linear(84, 10)
def forward(self, x):
x = self.conv1(self.reshape(x))
x = F.avg_pool2d(F.sigmoid(x), kernel_size = (2,2), stride=2)
x = F.avg_pool2d(F.sigmoid(self.conv2(x)), kernel_size = (2,2), stride=2)
x = self.flatten(x)
x = F.sigmoid(self.fc1(x))
x = F.sigmoid(self.fc2(x))
# x = F.relu(self.fc1(x))
# x = F.relu(self.fc2(x))
x = self.fc3(x)
return x
net = LN()
print(net)
# + colab={"base_uri": "https://localhost:8080/"} id="OZtYR8gq4KlE" outputId="062cfc9f-2c57-4a84-981d-050f4b8f1947"
params = list(net.parameters())
print(len(params))
print(params[0].size()) ### print lernable parameters shape : Conv2d -> first
# + colab={"base_uri": "https://localhost:8080/"} id="QUXK85O-4gFs" outputId="d86435d7-5699-41d1-a940-3a50fa4c3737"
for i in range(len(params)):
print(params[i].size())
# + id="mmwdPdWcSl5I"
# torch.Size([6, 1, 5, 5]) conv1 OK
# torch.Size([6]) 1 st pooling layer ??
# torch.Size([16, 6, 5, 5]) conv2 OK
# torch.Size([16]) 2 nd pooling layer ??
# torch.Size([120, 400]) linear OK
# torch.Size([120]) sigmoid (or relu) ??
# torch.Size([84, 120]) linear OK
# torch.Size([84]) sigmoid (or relu)
# torch.Size([10, 84]) linear OK
# torch.Size([10]) ?? what is the layer ??
# + [markdown] id="r0Ak1NK2MNkD"
# #### Training
# + [markdown] id="JolY4OuAudZZ"
# Data loading
# + id="YZD_zMurPCQI"
# from d2l.ai book
def load_data_fashion_mnist(batch_size, resize=None):
"""Download the Fashion-MNIST dataset and then load it into memory."""
## list of transformations
trans = [transforms.ToTensor()]
if resize:
trans.insert(0, transforms.Resize(resize))
trans = transforms.Compose(trans)
mnist_train = torchvision.datasets.FashionMNIST(
root="../data", train=True, transform=trans, download=True)
mnist_test = torchvision.datasets.FashionMNIST(
root="../data", train=False, transform=trans, download=True)
return (data.DataLoader(mnist_train, batch_size, shuffle=True,
num_workers=get_dataloader_workers()),
data.DataLoader(mnist_test, batch_size, shuffle=False,
num_workers=get_dataloader_workers()))
# + id="gOKeVSxFj9dh" colab={"base_uri": "https://localhost:8080/", "height": 369, "referenced_widgets": ["ca99fd97474d4036a3bf58f5d11eeb64", "b7a559e071ef408d93399dc73c8e7724", "45e53c7abfc14ec49689b1ea3321082a", "617f844fc90d46269403c679d850f450", "e8e12a49350e403f8e8722c0bdcdac7d", "0e586d2973634c43bcd5d8eded385392", "<KEY>", "<KEY>", "5615c795b3ea4f2da9683f93ad94b309", "3c34b8757e634aff82c71138f31563e1", "<KEY>", "<KEY>", "<KEY>", "57d53655e9ae4866bb5eefded4260c60", "0536caec9d3a4a568f016b0d23be4a8d", "984585ad5a2d4cb49e1eed0c1201f63d", "fa48f4a5938c487dae7f6177572c8225", "<KEY>", "<KEY>", "<KEY>", "14b58ed48da04cb481a07078a735d15e", "0e869cd9059c4cb4ba813b32396b3cb6", "9a92b5a4fa5441c28d058042415d889b", "2335e8fc201049cd8e174e5e74e87d14", "e94a2dfd21fc49af95ef2b2e40ab084a", "<KEY>", "<KEY>", "d3a95bae43cc42cbad741fbc33c465ea", "545c5ced84f9493297520205aff42005", "1cef2cbcab5644ec9b528946789e968b", "b0cca72124684ad6a4dfa549a09ce0de", "9e04764ffa3b41908bd31dfd7c9a5c59"]} outputId="2b2a1dd9-25ec-4c58-8886-973671b66714"
batch_size = 256
train_iter, test_iter = d2l.load_data_fashion_mnist(batch_size=batch_size)
# + [markdown] id="Vc-wc7pNugxV"
# Evaluate accuracy on GPU (if we have GPU)
# + id="8aJF-ETX5uUg"
## y_hat is assumed to be a matrix, and we assume that the second
## dimension stores prediction scores for each class.
## we work in paradigm: first dim -> batch ; second dim -> probabilities
## of values
def accuracy(y_hat, y):
"""Compute the number of correct predictions."""
if len(y_hat.shape) > 1 and y_hat.shape[1] > 1:
## choose the best value for every sample in batch
y_hat = y_hat.argmax(dim=1)
## y_hat is transformed to the same dtype as y and compared with y to produce
## bool tensor of True / False
## y is label
y_hat_cast = y_hat.type(y.dtype) ## -> y_hat_cast now have dtype of y
cmp = (y_hat_cast == y) ## -> cmp is tensor of False and True, have to convert to float
else: raise Exception("Input is not a matrix. Exit.")
return torch.sum(cmp).float()
# + id="mdgalkYhAOBA" colab={"base_uri": "https://localhost:8080/"} outputId="c4eeb5b9-4317-448c-bffa-f14dd81457c4"
class Accumulator:
"""For accumulating sums over `n` variables."""
def __init__(self, n):
self.data = torch.zeros(n)
def add(self, *args):
self.data = torch.as_tensor([a + float(b) for a, b in zip(self.data, args)])
def reset(self):
self.data = torch.zeros(n)
def __getitem__(self, idx):
return self.data[idx]
# + id="A5YqOwlF2BV4"
def evaluate_accuracy_gpu(net, data_iter, device=None):
"""Compute the accuracy for a model on a dataset using a GPU."""
net.eval() # Set the model to evaluation mode
if not device:
device = next(iter(net.parameters())).device
# No. of correct predictions, no. of predictions
metric = Accumulator(2)
for X, y in data_iter:
X, y = X.to(device), y.to(device)
metric.add(accuracy(net(X), y), y.numel())
return metric[0] / metric[1]
# + id="i58COZ9ljdLS"
## from d2l.ai
def train_ch6(net, train_iter, test_iter, num_epochs, lr,
device=d2l.try_gpu()):
"""Train a model with a GPU (defined in Chapter 6)."""
def init_weights(m):
if type(m) == nn.Linear or type(m) == nn.Conv2d:
torch.nn.init.xavier_uniform_(m.weight)
net.apply(init_weights)
print('training on', device)
net.to(device)
optimizer = torch.optim.SGD(net.parameters(), lr=lr)
loss = nn.CrossEntropyLoss()
animator = d2l.Animator(xlabel='epoch', xlim=[1, num_epochs],
legend=['train loss', 'train acc', 'test acc'])
timer, num_batches = d2l.Timer(), len(train_iter)
for epoch in range(num_epochs):
# Sum of training loss, sum of training accuracy, no. of examples
metric = Accumulator(3)
for i, (X, y) in enumerate(train_iter):
timer.start()
net.train()
optimizer.zero_grad()
X, y = X.to(device), y.to(device)
y_hat = net(X)
l = loss(y_hat, y)
l.backward()
optimizer.step()
with torch.no_grad():
metric.add(l * X.shape[0], d2l.accuracy(y_hat, y), X.shape[0])
timer.stop()
train_l = metric[0]/metric[2]
train_acc = metric[1]/metric[2]
if (i + 1) % (num_batches // 5) == 0 or i == num_batches - 1:
animator.add(epoch + (i + 1) / num_batches,
(train_l, train_acc, None))
test_acc = evaluate_accuracy_gpu(net, test_iter)
animator.add(epoch + 1, (None, None, test_acc))
print(f'loss {train_l:.3f}, train acc {train_acc:.3f}, '
f'test acc {test_acc:.3f}')
print(f'{metric[2] * num_epochs / timer.sum():.1f} examples/sec '
f'on {str(device)}')
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="WuoHd_tujhev" outputId="578843c7-8b61-440b-a7a9-ef7e0ff15de7"
lr, num_epochs = 0.9, 10
train_ch6(net, train_iter, test_iter, num_epochs, lr)
| PyTorch_LeNet_AvgPooling+Sigmoid.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import tqdm
import wandb
import torch
from torch import nn, optim
import torch.nn.functional as F
from torchvision import datasets, transforms
# +
mode = 'disabled'
# mode = 'online'
wandb.init(project='leafsnap', entity='dianna-ai', mode=mode)
# -
config = wandb.config
config.img_size = 128
config.batch_size = 32
config.epochs = 10
config.learning_rate = 0.0005
config.weight_decay = 1e-5
config.kernel_size = 5 # must be an odd number to keep the image dimensions the same after convolution
config.pooling = 2
config.channels1 = 12
config.channels2 = 12
config.channels3 = 24
config.channels4 = 24
config.dropout = .4
if torch.cuda.is_available():
device = torch.device('cuda')
else:
device = torch.device('cpu')
# +
dataset_root = os.path.expanduser('~/nlesc/DIANNA/datasets/leafsnap/leafsnap-dataset-30subset')
metadata = pd.read_csv(os.path.join(dataset_root, 'leafsnap-dataset-30subset-images.txt'), delimiter='\t')
metadata.head()
# +
def valid_file(fname):
# dataset consists of .jpg images
# converted to .bmp for faster loading
return fname.endswith('.bmp')
transform = transforms.Compose([transforms.Resize(config.img_size),
transforms.CenterCrop(config.img_size),
transforms.ToTensor()])
train_data = datasets.ImageFolder(os.path.join(dataset_root, 'dataset/split/train'), transform=transform, is_valid_file=valid_file)
val_data = datasets.ImageFolder(os.path.join(dataset_root, 'dataset/split/validation'), transform=transform, is_valid_file=valid_file)
test_data = datasets.ImageFolder(os.path.join(dataset_root, 'dataset/split/test'), transform=transform, is_valid_file=valid_file)
num_classes = len(train_data.classes)
num_samples = len(train_data) + len(val_data) + len(test_data)
print(f'Number of classes: {num_classes}')
print(f'Total number of samples: {num_samples}')
print(f'Number of training samples: {len(train_data)}')
print(f'Number of validation samples: {len(val_data)}')
print(f'Number of test samples: {len(test_data)}')
# +
workers = min(12, os.cpu_count() - 1)
train_dataloader = torch.utils.data.DataLoader(train_data, batch_size=config.batch_size, shuffle=True, num_workers=workers, pin_memory=True)
val_dataloader = torch.utils.data.DataLoader(val_data, batch_size=config.batch_size, shuffle=True, num_workers=workers, pin_memory=True)
test_dataloader = torch.utils.data.DataLoader(test_data, batch_size=config.batch_size, shuffle=True, num_workers=workers, pin_memory=True)
images, labels = next(iter(train_dataloader))
im = np.array(images[1].permute(1,2,0))
plt.imshow(im)
plt.title(train_data.classes[labels[0]])
# +
class Network(nn.Module):
def __init__(self, img_size, num_classes, kernel_size, pooling_size, num_channels, dropout):
super().__init__()
self.num = len(num_channels)
self.conv_layers = nn.ModuleList()
self.bn_layers = nn.ModuleList()
self.pool_layers = nn.ModuleList()
for i, out_channels in enumerate(num_channels):
if i == 0:
in_channels = 3
else:
in_channels = num_channels[i-1]
padding = int((kernel_size - 1) / 2)
self.conv_layers.append(nn.Conv2d(in_channels=in_channels, out_channels=out_channels, kernel_size=kernel_size, stride=1, padding=padding))
self.bn_layers.append(nn.BatchNorm2d(out_channels))
# no pooling layer for the last step
if i < self.num - 1:
self.pool_layers.append(nn.MaxPool2d(pooling_size, pooling_size))
# image size has been reduced by a factor pooling_size as many times as there are pooling layres
reduced_img_size = int(img_size / pooling_size ** len(self.pool_layers))
self.final_layer_size = num_channels[-1] * reduced_img_size ** 2
self.drop = nn.Dropout(dropout)
self.fc1 = nn.Linear(self.final_layer_size, num_classes)
def forward(self, data):
for i in range(self.num):
conv = self.conv_layers[i]
bn = self.bn_layers[i]
data = F.relu(bn(conv(data)))
# apply pooling except in the last step
if i < self.num - 1:
pool = self.pool_layers[i]
data = pool(data)
data = data.view(-1, self.final_layer_size)
data = self.fc1(self.drop(data))
return data
# Instantiate a neural network model
channels = [config.channels1, config.channels2, config.channels3, config.channels4]
model = Network(config.img_size, num_classes, config.kernel_size, config.pooling, channels, config.dropout).to(device)
print(f'Number of trainable parameters: {sum([p.numel() for p in model.parameters()])}')
# +
optimizer = optim.Adam(model.parameters(), lr=config.learning_rate, weight_decay=config.weight_decay)
loss_func = nn.CrossEntropyLoss().to(device)
def accuracy(model_output, y_true):
_, y_pred = torch.max(model_output, dim=1)
return (y_pred == y_true).sum() / len(y_pred)
def train(model, train_data, optimizer, loss_func):
epoch_loss = 0
epoch_acc = 0
model.train()
for batch in tqdm.notebook.tqdm(train_data):
images, labels = batch
images = images.to(device)
labels = labels.to(device)
optimizer.zero_grad()
predictions = model(images)
loss = loss_func(predictions, labels)
acc = accuracy(predictions, labels)
loss.backward()
optimizer.step()
epoch_loss += loss.item()
epoch_acc += acc.item()
wandb.log({'train_loss': loss.item(), 'train_acc': acc.item()})
epoch_loss /= len(train_data)
epoch_acc /= len(train_data)
return epoch_loss, epoch_acc
@torch.no_grad()
def evaluate(model, data, loss_func, train_mode=False):
avg_loss = 0
avg_acc = 0
if train_mode:
model.train()
else:
model.eval()
for batch in data:
images, labels = batch
images = images.to(device)
labels = labels.to(device)
predictions = model(images)
loss = loss_func(predictions, labels)
acc = accuracy(predictions, labels)
avg_loss += loss.item()
avg_acc += acc.item()
avg_loss /= len(data)
avg_acc /= len(data)
return avg_loss, avg_acc
# +
best_val_loss = np.inf
best_val_acc = None
for epoch in range(config.epochs):
# run training loop
train_loss, train_acc = train(model, train_dataloader, optimizer, loss_func)
# evalute on validation data, with same model settings as during training (i.e. including dropout)
val_loss, val_acc = evaluate(model, val_dataloader, loss_func)
wandb.log({'train_epoch_loss': train_loss, 'train_epoch_acc': train_acc})
wandb.log({'val_epoch_loss': val_loss, 'val_epoch_acc': val_acc})
print(f'train loss: {train_loss:.2f} | train acc: {train_acc:.2f}')
print(f'val loss: {val_loss:.2f} | val acc: {val_acc:.2f}')
if val_loss < best_val_loss:
best_val_loss = val_loss
best_val_acc = val_acc
# ensure to store the model in eval mode
model.eval()
torch.save(model, 'leafsnap_model.pytorch')
print(f'best val loss: {best_val_loss:.2f} | best val acc: {best_val_acc:.2f}')
# -
test_loss, test_acc = evaluate(model, test_dataloader, loss_func, train_mode=False)
print(f'test loss: {test_loss:.2f} | test acc: {test_acc:.2f}')
# +
# load model from disk an convert to onnx
loaded_model = torch.load('leafsnap_model.pytorch')
loaded_model.eval()
# export needs example input
x = next(iter(train_dataloader))[0].to(device)
torch.onnx.export(loaded_model, x, 'leafsnap_model.onnx', opset_version=11,
export_params=True, input_names=['input'], output_names=['output'],
dynamic_axes={'input': {0: 'batch_size'},
'output': {0: 'batch_size'}})
| example_data/model_generation/LeafSnap/generate_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# #### Data source :https://archive.ics.uci.edu/ml/machine-learning-databases/00228/
import numpy as np
import pandas as pd
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.cross_validation import train_test_split
from sklearn.naive_bayes import MultinomialNB
df=pd.read_csv('smsspam',sep='\t',names=['Status','Message'])
df.head()
len(df)
len(df[df.Status=='spam'])
len(df[df.Status=='ham'])
df.loc[df["Status"]=='ham',"Status",]=1
df.loc[df["Status"]=='spam',"Status",]=0
df.head()
df_x=df["Message"]
df_y=df["Status"]
cv = CountVectorizer()
x_train, x_test, y_train, y_test = train_test_split(df_x, df_y, test_size=0.2, random_state=4)
x_train.head()
cv = CountVectorizer()
x_traincv = cv.fit_transform(["Hi How are you How are you doing","Hi what's up","Wow that's awesome"])
x_traincv.toarray()
cv.get_feature_names()
cv1 = CountVectorizer()
x_traincv=cv1.fit_transform(x_train)
a=x_traincv.toarray()
a[0]
cv1.inverse_transform(a[0])
x_train.iloc[0]
x_testcv=cv1.transform(x_test)
x_testcv.toarray()
mnb = MultinomialNB()
y_train=y_train.astype('int')
y_train
mnb.fit(x_traincv,y_train)
testmessage=x_test.iloc[0]
predictions=mnb.predict(x_testcv)
predictions
a=np.array(y_test)
a
count=0
for i in range (len(predictions)):
if predictions[i]==a[i]:
count=count+1
count
len(predictions)
1092/1115.0
| Data_Science/Text Analytics CV.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # A Network Tour of Data Science
# ### <NAME>, Winter 2016/17
# ## Assignment 1 : Unsupervised Clustering with the Normalized Association
# +
# Load libraries
# Math
import numpy as np
# Visualization
# %matplotlib notebook
import matplotlib.pyplot as plt
plt.rcParams.update({'figure.max_open_warning': 0})
from mpl_toolkits.axes_grid1 import make_axes_locatable
from scipy import ndimage
# Print output of LFR code
import subprocess
# Sparse matrix
import scipy.sparse
import scipy.sparse.linalg
# 3D visualization
import pylab
from mpl_toolkits.mplot3d import Axes3D
from matplotlib import pyplot
# Import data
import scipy.io
# Import functions in lib folder
import sys
sys.path.insert(1, 'lib')
# Import helper functions
# %load_ext autoreload
# %autoreload 2
from lib.utils import construct_kernel
from lib.utils import compute_kernel_kmeans_EM
from lib.utils import compute_purity
# Import distance function
import sklearn.metrics.pairwise
# Remove warnings
import warnings
warnings.filterwarnings("ignore")
# -
# **Question 1:** Write down the mathematical relationship between Normalized Cut (NCut) and Normalized Association (NAssoc) for K clusters. It is not necessary to provide details.
#
# The Normalized Cut problem is defined as:<br><br>
# $$
# \min_{\{S_k\}}\ NCut(\{S_k\}) := \sum_{k=1}^K \frac{Cut(S_k,S_k^c)}{Vol(S_k)} \ \textrm{ s.t. } \ \cup_{k=1}^{K} S_k = V, \ S_k \cap S_{k'}=\emptyset, \ \forall k \not= k' \quad\quad\quad(1)
# $$
#
# and the Normalized Association problem is defined as:<br><br>
# $$
# \max_{\{S_k\}}\ NAssoc(\{S_k\}):= \sum_{k=1}^K \frac{Assoc(S_k,S_k)}{Vol(S_k)} \ \textrm{ s.t. } \ \cup_{k=1}^{K} S_k = V, \ S_k \cap S_{k'}=\emptyset, \ \forall k \not= k' .
# $$
#
#
# We may rewrite the Cut operator and the Volume operator with the Assoc operator as:<br><br>
# $$
# Vol(S_k) = \sum_{i\in S_k, j\in V} W_{ij} \\
# Assoc(S_k,S_k) = \sum_{i\in S_k, j\in S_k} W_{ij} \\
# Cut(S_k,S_k^c) = \sum_{i\in S_k, j\in S_k^c=V\setminus S_k} W_{ij} = \sum_{i\in S_k, j\in V} W_{ij} - \sum_{i\in S_k, j\in S_k} W_{ij} = Vol(S_k) - Assoc(S_k,S_k)
# $$
#
#
# **Answer to Q1:** Your answer here.
#
# We have<br><br>
# $$
# \frac{Cut(S_k,S_k^c)}{Vol(S_k)} = \frac{Vol(S_k) - Assoc(S_k,S_k)}{Vol(S_k)} = 1- \frac{Assoc(S_k,S_k)}{Vol(S_k)}
# $$
#
# and<br><br>
#
# $$
# \sum_{k=1}^K \frac{Cut(S_k,S_k^c)}{Vol(S_k)} = K - \sum_{k=1}^K \frac{Assoc(S_k,S_k)}{Vol(S_k)}
# $$
#
# The relationship between Normalized Cut (NCut) and Normalized Association (NAssoc) for K clusters is thus<br><br>
#
# $$
# NCut(\{S_k\}) = K - NAssoc(\{S_k\}).
# $$
#
# **Question 2:** Using the relationship between NCut and NAssoc from Q1, it is therefore equivalent to maximize NAssoc by minimizing or maximizing NCut? That is
#
# $$
# \max_{\{S_k\}}\ NAssoc(\{S_k\}) \ \textrm{ s.t. } \cup_{k=1}^{K} S_k = V, \quad S_k \cap S_{k'}=\emptyset, \ \forall k \not= k'
# $$
#
# $$
# \Updownarrow
# $$
#
# $$
# \min_{\{S_k\}}\ NCut(\{S_k\}) \ \textrm{ s.t. } \cup_{k=1}^{K} S_k = V, \quad S_k \cap S_{k'}=\emptyset, \ \forall k \not= k'
# $$
#
# or
#
# $$
# \max_{\{S_k\}}\ NCut(\{S_k\}) \ \textrm{ s.t. } \cup_{k=1}^{K} S_k = V, \quad S_k \cap S_{k'}=\emptyset, \ \forall k \not= k'
# $$
#
# It is not necessary to provide details.
# **Answer to Q2:** Your answer here.
#
# As $\min F \Leftrightarrow \max -F$, we have equivalence between the max NAssoc problem:
#
# $$
# \max_{\{S_k\}}\ NAssoc(\{S_k\}) \ \textrm{ s.t. } \cup_{k=1}^{K} S_k = V, \quad S_k \cap S_{k'}=\emptyset, \ \forall k \not= k'
# $$
#
# and the min NCut problem:
#
# $$
# \min_{\{S_k\}}\ NCut(\{S_k\}) \ \textrm{ s.t. } \cup_{k=1}^{K} S_k = V, \quad S_k \cap S_{k'}=\emptyset, \ \forall k \not= k'
# $$
# **Question 3:** Solving the NCut problem in Q2 is NP-hard => let us consider a spectral relaxation of NCut. Write down the Spectral Matrix A of NCut that satisfies the equivalent functional optimization problem of Q2:
#
# $$
# \min_{Y}\ tr( Y^\top A Y) \ \textrm{ s.t. } \ Y^\top Y = I_K \textrm{ and } Y \in Ind_S, \quad\quad\quad(3)
# $$
#
# where
#
# $$
# Y \in Ind_S \ \textrm{ reads as } \ Y_{ik} =
# \left\{
# \begin{array}{ll}
# \big(\frac{D_{ii}}{Vol(S_k)}\big)^{1/2} & \textrm{if} \ i \in S_k\\
# 0 & \textrm{otherwise}
# \end{array}
# \right..
# $$
#
# and
#
# $$
# A=???
# $$
#
# It is not necessary to provide details.
#
# *Hint:* Let us introduce the indicator matrix $F$ of the clusters $S_k$ such that:
#
# $$
# F_{ik} =
# \left\{
# \begin{array}{ll}
# 1 & \textrm{if} \ i \in S_k\\
# 0 & \textrm{otherwise}
# \end{array}
# \right..
# $$
#
# We may rewrite the Cut operator and the Volume operator with $F$ as:
#
# $$
# Vol(S_k) = \sum_{i\in S_k, j\in V} W_{ij} = F_{\cdot,k}^\top D F_{\cdot,k}\\
# Cut(S_k,S_k^c) = \sum_{i\in S_k, j\in V} W_{ij} - \sum_{i\in S_k, j\in S_k} W_{ij} = F_{\cdot,k}^\top D F_{\cdot,k} - F_{\cdot,k}^\top W F_{\cdot,k} = F_{\cdot,k}^\top (D - W) F_{\cdot,k} \quad
# $$
#
# We thus have
#
# $$
# \frac{Cut(S_k,S_k^c)}{Vol(S_k)} = \frac{ F_{\cdot,k}^\top (D - W) F_{\cdot,k} }{ F_{\cdot,k}^\top D F_{\cdot,k} }
# $$
#
#
# Set $\hat{F}_{\cdot,k}=D^{1/2}F_{\cdot,k}$ and observe that
#
# $$
# \frac{ F_{\cdot,k}^\top (D - W) F_{\cdot,k} }{ F_{\cdot,k}^\top D F_{\cdot,k} } = \frac{ \hat{F}_{\cdot,k}^\top D^{-1/2}(D - W)D^{-1/2} \hat{F}_{\cdot,k} }{ \hat{F}_{\cdot,k}^\top \hat{F}_{\cdot,k} } = \frac{ \hat{F}_{\cdot,k}^\top (I - D^{-1/2}WD^{-1/2}) \hat{F}_{\cdot,k} }{ \hat{F}_{\cdot,k}^\top \hat{F}_{\cdot,k} } ,
# $$
#
# with $L_N=I - D^{-1/2}WD^{-1/2}$ is the normalized graph Laplacian. Set $Y_{\cdot,k}=\frac{\hat{F}_{\cdot,k}}{\|\hat{F}_{\cdot,k}\|_2}$:
#
# $$
# \frac{ \hat{F}_{\cdot,k}^\top L_N \hat{F}_{\cdot,k} }{ \hat{F}_{\cdot,k}^\top \hat{F}_{\cdot,k} } = Y_{\cdot,k}^\top L_N Y_{\cdot,k} \quad\quad\quad(2)
# $$
#
#
# Using (2), we can rewrite (1) as a functional optimization problem:
#
# $$
# \min_{Y}\ tr( Y^\top A Y) \ \textrm{ s.t. } \ Y^\top Y = I_K \textrm{ and } Y \in Ind_S,
# $$
#
# where
#
#
# $$
# Y \in Ind_S \ \textrm{ reads as } \ Y_{ik} =
# \left\{
# \begin{array}{ll}
# \big(\frac{D_{ii}}{Vol(S_k)}\big)^{1/2} & \textrm{if} \ i \in S_k\\
# 0 & \textrm{otherwise}
# \end{array}
# \right..
# $$
#
# and
#
# $$
# A=???
# $$
# **Answer to Q3:** Let us introduce the indicator matrix $F$ of the clusters $S_k$ such that:
#
# $$
# F_{ik} =
# \left\{
# \begin{array}{ll}
# 1 & \textrm{if} \ i \in S_k\\
# 0 & \textrm{otherwise}
# \end{array}
# \right..
# $$
#
# We may rewrite the Cut operator and the Volume operator with $F$ as:
#
# $$
# Vol(S_k) = \sum_{i\in S_k, j\in V} W_{ij} = F_{\cdot,k}^\top D F_{\cdot,k}\\
# Cut(S_k,S_k^c) = \sum_{i\in S_k, j\in V} W_{ij} - \sum_{i\in S_k, j\in S_k} W_{ij} = F_{\cdot,k}^\top D F_{\cdot,k} - F_{\cdot,k}^\top W F_{\cdot,k} = F_{\cdot,k}^\top (D - W) F_{\cdot,k} \quad
# $$
#
# We thus have
#
# $$
# \frac{Cut(S_k,S_k^c)}{Vol(S_k)} = \frac{ F_{\cdot,k}^\top (D - W) F_{\cdot,k} }{ F_{\cdot,k}^\top D F_{\cdot,k} }
# $$
#
#
# Set $\hat{F}_{\cdot,k}=D^{1/2}F_{\cdot,k}$ and observe that
#
# $$
# \frac{ F_{\cdot,k}^\top (D - W) F_{\cdot,k} }{ F_{\cdot,k}^\top D F_{\cdot,k} } = \frac{ \hat{F}_{\cdot,k}^\top D^{-1/2}(D - W)D^{-1/2} \hat{F}_{\cdot,k} }{ \hat{F}_{\cdot,k}^\top \hat{F}_{\cdot,k} } = \frac{ \hat{F}_{\cdot,k}^\top (I - D^{-1/2}WD^{-1/2}) \hat{F}_{\cdot,k} }{ \hat{F}_{\cdot,k}^\top \hat{F}_{\cdot,k} } ,
# $$
#
# where $L_N=I - D^{-1/2}WD^{-1/2}$ is the normalized graph Laplacian. Set $Y_{\cdot,k}=\frac{\hat{F}_{\cdot,k}}{\|\hat{F}_{\cdot,k}\|_2}$, we have:
#
# $$
# \frac{ \hat{F}_{\cdot,k}^\top L_N \hat{F}_{\cdot,k} }{ \hat{F}_{\cdot,k}^\top \hat{F}_{\cdot,k} } = Y_{\cdot,k}^\top L_N Y_{\cdot,k} \quad\quad\quad(2)
# $$
#
#
# Using (2), we can rewrite (1) as a functional optimization problem:
#
# $$
# \min_{Y}\ tr( Y^\top A Y) \ \textrm{ s.t. } \ Y^\top Y = I_K \textrm{ and } Y \in Ind_S,
# $$
#
# where
#
# $$
# Y \in Ind_S \ \textrm{ reads as } \ Y_{ik} =
# \left\{
# \begin{array}{ll}
# \big(\frac{D_{ii}}{Vol(S_k)}\big)^{1/2} & \textrm{if} \ i \in S_k\\
# 0 & \textrm{otherwise}
# \end{array}
# \right..
# $$
#
# and
#
# $$
# A=L_N=I-D^{-1/2}WD^{-1/2}.
# $$
# **Question 4:** Drop the cluster indicator constraint $Y\in Ind_S$ in Q3, how do you compute the solution $Y^\star$ of (3)? Why the first column of $Y^\star$ is not relevant for clustering?
# **Answer to Q4:** Your answer here.
#
# Dropping the constraint $Y\in Ind_S$ in (3) leads to a standard spectral relaxation problem:
#
# $$
# \min_{Y}\ tr( Y^\top A Y) \ \textrm{ s.t. } \ Y^\top Y = I_K,
# $$
#
# which solution $Y^\star$ is given by the $K$ smallest eigenvectors/eigenvalues of $A=L_N=I-D^{-1/2}WD^{-1/2}$. Note that the first column of $Y^\star$ is the constant signal $y_1=\frac{1}{\sqrt{n}}1_{n\times 1}$ associated to the smallest eigenvalue of $L_N$, which has value $\lambda_1=0$.
# **Question 5:** Plot in 3D the 2nd, 3rd, 4th columns of $Y^\star$. <br>
# Hint: Compute the degree matrix $D$.<br>
# Hint: You may use function *D_sqrt_inv = scipy.sparse.diags(d_sqrt_inv.A.squeeze(), 0)* for creating $D^{-1/2}$.<br>
# Hint: You may use function *I = scipy.sparse.identity(d.size, dtype=W.dtype)* for creating a sparse identity matrix.<br>
# Hint: You may use function *lamb, U = scipy.sparse.linalg.eigsh(A, k=4, which='SM')* to perform the eigenvalue decomposition of A.<br>
# Hint: You may use function *ax.scatter(Xdisp, Ydisp, Zdisp, c=Cgt)* for 3D visualization.
# Load dataset: W is the Adjacency Matrix and Cgt is the ground truth clusters
mat = scipy.io.loadmat('datasets/mnist_2000_graph.mat')
W = mat['W']
n = W.shape[0]
Cgt = mat['Cgt'] - 1; Cgt = Cgt.squeeze()
nc = len(np.unique(Cgt))
print('Number of nodes =',n)
print('Number of classes =',nc);
# +
# Your code here
# Construct Spectal Matrix A
d = W.sum(axis=0) + 1e-6 # degree vector
d = 1.0 / np.sqrt(d)
Dinv = scipy.sparse.diags(d.A.squeeze(), 0)
I = scipy.sparse.identity(d.size, dtype=W.dtype)
A = I - Dinv* (W* Dinv)
# Compute K smallest eigenvectors/eigenvalues of A
lamb, U = scipy.sparse.linalg.eigsh(A, k=4, which='SM')
# Sort eigenvalue from smallest to largest values
idx = lamb.argsort() # increasing order
lamb, U = lamb[idx], U[:,idx]
print(lamb)
# Y*
Y = U
# Plot in 3D the 2nd, 3rd, 4th columns of Y*
Xdisp = Y[:,1]
Ydisp = Y[:,2]
Zdisp = Y[:,3]
# 2D Visualization
plt.figure(14)
size_vertex_plot = 10
plt.scatter(Xdisp, Ydisp, s=size_vertex_plot*np.ones(n), c=Cgt)
plt.title('2D Visualization')
plt.show()
# 3D Visualization
fig = pylab.figure(15)
ax = Axes3D(fig)
ax.scatter(Xdisp, Ydisp, Zdisp, c=Cgt)
pylab.title('3D Visualization')
pyplot.show()
# -
# **Question 6:** Solve the unsupervised clustering problem for MNIST following the popular technique of [<NAME>, “On Spectral Clustering: Analysis and an algorithm”, 2002], i.e. <br>
# (1) Compute $Y^\star$? solution of Q4. <br>
# (2) Normalize the rows of $Y^\star$? with the L2-norm. <br>
# Hint: You may use function X = ( X.T / np.sqrt(np.sum(X**2,axis=1)+1e-10) ).T for the L2-normalization of the rows of X.<br>
# (3) Run standard K-Means on normalized $Y^\star$? to get the clusters, and compute the clustering accuracy. You should get more than 50% accuracy.
# Your code here
# Normalize the rows of Y* with the L2 norm, i.e. ||y_i||_2 = 1
Y = ( Y.T / np.sqrt(np.sum(Y**2,axis=1)+1e-10) ).T
# Your code here
# Run standard K-Means
Ker = construct_kernel(Y,'linear') # Compute linear Kernel for standard K-Means
Theta = np.ones(n) # Equal weight for each data
[C_kmeans,En_kmeans] = compute_kernel_kmeans_EM(nc,Ker,Theta,10)
acc= compute_purity(C_kmeans,Cgt,nc)
print('accuracy standard kmeans=',acc)
| algorithms/02_sol_assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # 超参调节、批量标准化、编程框架
# ## 1. 超参调节
# ### 1.1 调参过程
#
# 调参的优先级:
#
# 1)学习速率 $\alpha$
#
# 2)如果使用动量梯度下降 $\beta$;隐藏神经元数量;微批大小。
#
# 3)神经网络的层数;学习速率衰减
#
# 4)如果使用Adam,几乎不太会去调 $\beta_1=0.9, \beta_2=0.999, \epsilon=10^{-8}$
#
# 在机器学习的早期阶段,通常会使用网格搜索。当超参较少的情况下,网格搜索是没问题的。对于神经网络这样超参众多的模型,更推荐随机挑选。因为通常很难预先知道哪一个超参会更重要,实际上超参发挥的作用,会有很大的差距,会重要的超参尝试更多的值搜索效果会更好。
#
# 
#
# 另一个推荐的搜索策略,是采用由粗到细的搜索(coarse to fine),在表现较好的超参区域,放大,进一步搜索。
#
# 
# ### 1.2 使用合适的刻度来搜索超参
#
# 上一节提到,随机挑选超参可以提高超参搜索的效率,但与此同时,随机抽样并不意味着随机均匀抽样。当超参搜索的范围量级变动较大时,使用合适的刻度来随机抽样十分重要。
#
# 
#
# 
# ### 1.3 超参调节实践:熊猫策略 vs 鱼子酱策略
#
# 不同类型的问题,超参通常都是重新调节的;即使对于同一个问题,随着时间推移,新的数据进入训练集,时不时对模型表现进行评估并重新搜索超参也是十分必要的。
#
# 取决于所拥有的计算资料,调节超参也包含串行和并行两种策略。
#
# 
# ## 2. 批量标准化
#
# ### 2.1 对神经网络中的激活值进行标准化
#
# 批量标准化可以使超参搜索更容易,使神经网络模型更健壮,也使得训练更深的神经网络更为容易。批量标准化的起源,是当我们对输入进行标准化后,训练过程会更快。基于同样的原因,如果对每个隐藏层的激活值也进行标准化,那么训练过程会不会进一步加快?但在技术实现中,**实际情况更多是对线性组合的Z值进行标准化,而非激活值**。
#
# 
#
# 批量标准化,并不像对输入的处理,将其都标准化为均值0,标准差1,而是允许不同的均值和标准差。这个是通过 $\gamma$ 和 $\beta$ 这两个值来控制的,这两个值像权重一样,是模型可以学习得到的。
#
# 
# ### 2.2 将批量标准化应用到神经网络中
#
# 
#
# 批量标准化通常和微批梯度下降一起使用。另外,在使用了批量标准化之后,参数 $b$ 实际上可以去除了。
#
# 
# ### 2.3 理解批量标准化
#
# 1. 和对输入进行标准化一样,可以加快模型训练速度。
#
# 2. 能更好地应对数据集输入分布的偏移。这种偏移,术语叫做covariate shift。它描述的是,算法已经学到了从X到Y的映射关系,当输入X的分布发生变化,我们可能需要重新训练模型。不论X->Y的真实函数没有发生变化,重新训练模型都是必要的。
# 
# 
#
# 3. 类似于Dropout,批量标准化同时也起到正则化的效果。(但不要将其作为正则化的手段来用)
# 
# ### 2.4 针对测试集的批量标准化
#
# 批量标准化在训练过程中,每次处理一个微批。而在针对测试集进行预测的时候,可能会需要对单个样本进行计算,这时会缺少微批的均值和方差。我们需要单独预测均值和方差。可以使用各个批次的均值和方差,使用指数加权移动平均的方式,来进行预测。
#
# 
# ## 3. 多元分类
# ### 3.1 Softmax回归
#
# Softmax回归是Logistic回归的推广形式,可以用来处理多元分类问题。将神经网络最后一层,替换为Softmax回归层,即可进行多元分类。
#
# 
#
# Softmax回归本身的决策边界是线性的。
#
# 
# ### 3.2 训练Softmax分类器
#
# Softmax的概念是和Hardmax对应的,Hardmax会产出一个类似[1, 0, 0, 0]的向量,而Softmax产生的向量,是和为1的概率表述向量。
#
# Softmax的损失函数,反向传播,略。
# ## 4. 编程框架简介
# ### 4.1 深度学习框架
#
# 
# ### 4.2 Tensorflow
#
# 略
| Hyperparameter tuning, Batch Normalization and Programming Frameworks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **Importar bibliotecas**
import os, flopy
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import flopy.utils.binaryfile as bf
import shutil
# **MODELO DE FLUJO**
# **Asignación de nombre de modelo**
model_ws='../4_OUT/R_SIM'
modelname='Modelo_Flujo'
exe_name= '../1_SOLVER/mf2005.exe'
mf = flopy.modflow.Modflow(modelname, exe_name=exe_name, model_ws=model_ws)
# **Discretización Espacial**
# +
nlay = 30
nrow = 1
ncol = 100
delr=10.
delc=1.
top = 510.
botm = np.ones((nlay), dtype=np.int32)
for i in range (nlay):
botm[i]= 500 -(i*10)
nper = 3
nstp = [1]+[1]*2
perlen = [1] + [864000]*1 + [2523743999]*1
steady = [True] + [False]*2
dis=flopy.modflow.ModflowDis(mf, nlay=nlay, nrow=nrow, ncol=ncol, delr=delr, delc=delc, top=top,
botm=botm, nper=nper, perlen=perlen, steady=steady, itmuni=1, lenuni=2)
# +
#botm
#dis.plot()
# -
# **Definición de celdas activas/paquete BAS**
#
#
ibound = np.ones((nlay, nrow, ncol), dtype=np.int32)
ibound[1:,:,0] = -1
ibound[21:,:,-1] = -1
#ibound
# **Altura inicial - strt**
bas = flopy.modflow.ModflowBas(mf, ibound=ibound, strt=top)
#bas.plot()
# **Nivel constante - CHD**
# +
chdspd={}
chdspd[0]=[[i, 0, 0, 500., 500.] for i in range(1,30)]
for i in range (21,30):
chdspd[0].append(([i, 0, 99, 300.,300.]))
chdspd[1]=[[i, 0, 0, 500., 500.] for i in range(1,30)]
for i in range (21,30):
chdspd[1].append(([i, 0, 99, 300.,300.]))
chdspd[2]=[[i, 0, 0, 500., 500.] for i in range(1,30)]
for i in range (21,30):
chdspd[2].append(([i, 0, 99, 300.,300.]))
chd = flopy.modflow.ModflowChd(mf, stress_period_data=chdspd)
# -
# **Conductividad Hidráulica**
kx=pd.read_excel('../3_IN/SIMULACION_K.xlsx', sheet_name=None, header=None)
#kx
# **Paquete solucionador mf2005**
pcg =flopy.modflow.ModflowPcg(mf, mxiter=85, iter1=57, npcond=1, nbpol=1, hclose=1e-05, rclose=1e-05, relax=1,
iprpcg=0, mutpcg=0, ihcofadd=0)
#numero de simulaciones estocasticas
n_estocasticas=['00','01','02','03','04','05','06','07','08','09','10','11','12','13','14','15','16','17','18','19',
'20','21','22','23','24','25','26','27','28','29','30','31','32','33','34','35','36','37','38','39',
'40','41','42','43','44','45','46','47','48','49','50','51','52','53','54','55','56','57','58','59',
'60','61','62','63','64','65','66','67','68','69','70','71','72','73','74','75','76','77','78','79',
'80','81','82','83','84','85','86','87','88','89','90','91','92','93','94','95','96','97','98','99']
for i in n_estocasticas:
hk=kx[i].to_numpy()
lpf=flopy.modflow.ModflowLpf(mf, laytyp=1, ipakcb=53, hk=hk, vka=hk, hdry=-2e+20, chani=-1.0, ss=1e-05, sy=0.15)
#laywet=1, wetfct=1, iwetit=4, ihdwet=0)
#**Output Control - Paquete OC**
spd_oc = {}
for kper in range(nper):
for kstp in range(nstp[kper]):
spd_oc[(kper, kstp)] = ['save head','save budget']
oc= flopy.modflow.ModflowOc(mf,stress_period_data=spd_oc, compact=True)
#**LMT Linkage with MT3DMS for multi-species mass transport modeling**
lmt = flopy.modflow.ModflowLmt(mf, output_file_name='mt3d.ftl')
#**Escribir los archivos de salida mf2005**
mf.write_input()
#**Correr Modelo de Flujo**
mf.run_model(silent=True)
#Copiar y Guardar archivos de simulaciones
shutil.copy(model_ws +'/'+'Modelo_flujo.hds', model_ws+'/'+'hds'+'/'+"Modelo_Flujo_{}.hds".format(i))
shutil.copy(model_ws +'/'+'Modelo_flujo.cbc', model_ws+'/'+'cbc'+'/'+"Modelo_Flujo_{}.cbc".format(i))
shutil.copy(model_ws +'/'+'Modelo_flujo.list', model_ws+'/'+'list'+'/'+"Modelo_Flujo_{}.list".format(i))
shutil.copy(model_ws +'/'+'mt3d.ftl', model_ws+'/'+'mt3d'+'/'+"mt3d_{}.ftl".format(i))
# **MODELO DE TRANSPORTE**
# **Definicion del modelo**
for j in n_estocasticas:
ftlfilename='mt3d'+'/'+'/mt3d_{}.ftl'.format(j)
namemt3d='transModel'
mt_model = flopy.mt3d.Mt3dms(modelname=namemt3d, model_ws=model_ws, ftlfilename=ftlfilename, version='mt3d-usgs',
exe_name='../1_SOLVER/mt3d-usgs_1.1.0_64.exe', modflowmodel=mf)
#BTN file
timprs=[10800., 1728000., 4320000., 7776000., 15552000., 31536000., 94672800., 157788000., 220903200., 946728000.,
1105000000., 1578000000.]
icbund = np.ones((nlay, nrow, ncol), dtype=np.int32)
icbund[1:,:,0] = -1
nprs=len(timprs)
nper = 3
perlen = [1] + [864000]*1 + [2523800000]*1
btn = flopy.mt3d.Mt3dBtn(mt_model, ncomp=1, nper=nper, mcomp=1, tunit='sec', lunit='m', munit='g', prsity=0.25,
sconc=0.0, cinact=-1e+30, thkmin=0.01, ifmtcn=12, ifmtnp=5, ifmtrf=12, ifmtdp=12,
perlen=perlen, savucn=True, nprs=nprs, timprs=timprs, dt0=0, mxstrn=50000,ttsmult=1.,
ttsmax=0, icbund=icbund)
#Paquete de Advencion - ADV
adv = flopy.mt3d.Mt3dAdv(mt_model, mixelm=3, percel=1, mxpart=75000, nadvfd=2, itrack=2, wd=1, dceps=1e-05,
nplane=0, npl=0, nph=10, npmin=2, npmax=20, npsink=10, dchmoc=0.01)
#Solucionador - GCG
gcg = flopy.mt3d.Mt3dGcg(mt_model, mxiter=1, iter1=200, isolve=3, ncrs=0, cclose=1e-06, iprgcg=1)
#Paquete de Dispersion - DSP
dsp = flopy.mt3d.Mt3dDsp(mt_model, al=0.0, multiDiff=True, dmcoef=0, trpt=0.1, trpv=0.01)
itype = flopy.mt3d.Mt3dSsm.itype_dict()
#itype
#Concentracion del contamiante - SSM
ssm_data = {}
ssm_data[1] = [(k, 0, 0, 100, 1) for k in range (1,30)]
for k in range (1,30):
ssm_data[1].append((k, 0, 0, 100, -1))
ssm_data[2] = [(k, 0, 0, 0, 1) for k in range (1,30)]
for k in range (1,30):
ssm_data[2].append((k, 0, 0, 0, -1))
ssm = flopy.mt3d.Mt3dSsm(mt_model, stress_period_data=ssm_data)
#Escribir archivos de Salida del modelo de transporte - MT3D
mt_model.write_input()
#Correr de modelo de Transporte - MT3D
mt_model.run_model(silent=True)
shutil.copy(model_ws +'/'+'MT3D001.UCN', model_ws+'/'+'ucn'+'/'+"MT3D001_{}.UCN".format(j))
shutil.copy(model_ws +'/'+'MT3D001.MAS', model_ws+'/'+'t_mas'+'/'+"MT3D001_{}.MAS".format(j))
shutil.copy(model_ws +'/'+'transModel.list', model_ws+'/'+'t_list'+'/'+"Trans_{}.list".format(j))
| 2_Simulacion_Estocastica.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:RoboND]
# language: python
# name: conda-env-RoboND-py
# ---
# +
def get_line(start, end):
"""Bresenham's Line Algorithm
Produces a list of tuples from start and end
>>> points1 = get_line((0, 0), (3, 4))
>>> points2 = get_line((3, 4), (0, 0))
>>> assert(set(points1) == set(points2))
>>> print points1
[(0, 0), (1, 1), (1, 2), (2, 3), (3, 4)]
>>> print points2
[(3, 4), (2, 3), (1, 2), (1, 1), (0, 0)]
"""
# Setup initial conditions
x1, y1 = start
x2, y2 = end
dx = x2 - x1
dy = y2 - y1
# Determine how steep the line is
is_steep = abs(dy) > abs(dx)
# Rotate line
if is_steep:
x1, y1 = y1, x1
x2, y2 = y2, x2
# Swap start and end points if necessary and store swap state
swapped = False
if x1 > x2:
x1, x2 = x2, x1
y1, y2 = y2, y1
swapped = True
# Recalculate differentials
dx = x2 - x1
dy = y2 - y1
# Calculate error
error = int(dx / 2.0)
ystep = 1 if y1 < y2 else -1
# Iterate over bounding box generating points between start and end
y = y1
points = []
for x in range(x1, x2 + 1):
coord = (y, x) if is_steep else (x, y)
points.append(coord)
error -= abs(dy)
if error < 0:
y += ystep
error += dx
# Reverse the list if the coordinates were swapped
if swapped:
points.reverse()
return points
points1 = get_line((0, 0), (50, 40))
points2 = get_line((50, 40), (0, 0))
assert(set(points1) == set(points2))
print (points1)
print (points2)
# -
# Rover transformed vision image is y=160, x=320
# +
from math import sin, cos, radians, pi
def point_pos(x0, y0, d, theta_degrees):
""" """
theta_rad = pi/2 - radians(theta_degrees)
return x0 + d*cos(theta_rad), y0 + d*sin(theta_rad)
top_point = point_pos(160, 160, 150, 15)
print(top_point)
# +
import numpy as np
driving_mask = np.zeros((20, 20))
driving_mask[:10,:10] = 1 # assign ones to a range of rows and columns
# driving_mask[:,[1,2]] = 1 # select all rows and specific columns
print (driving_mask)
# +
import numpy as np
driving_mask = np.zeros((24, 24)) # initialize matrix of zeros
driving_mask.shape[1], driving_mask.shape[0] # .shape = (rows, col)
H_start_percent = 0 # percent value
H_end_percent = 90 # percent value
V_start_percent = 10 # percent value
V_end_percent = 20 # percent value
def remap_values(value, inMin, inMax, outMin, outMax):
# Figure out how 'wide' each range is
inSpan = inMax - inMin
outSpan = outMax - outMin
# Convert the left range into a 0-1 range (float)
valueScaled = float(value - inMin) / float(inSpan)
# Convert the 0-1 range into a value in the right range.
return outMin + (valueScaled * outSpan)
H_start_col = int(round(remap_values(H_start_percent, 0, 100, 0, driving_mask.shape[1])))
# print(H_start_col)
H_end_col = int(round(remap_values(H_end_percent, 0, 100, 0, driving_mask.shape[1])))
# print(H_end_col)
V_start_col = int(round(remap_values(V_start_percent, 0, 100, 0, driving_mask.shape[0])))
# print(V_start_col)
V_end_col = int(round(remap_values(V_end_percent, 0, 100, 0, driving_mask.shape[0])))
# print(V_end_col)
driving_mask[V_start_col:V_end_col,H_start_col:H_end_col] = 1 # select range of rows and columns
# driving_mask[:,[1,2]] = 1 # select all rows and specific columns
print (driving_mask)
# -
| code/Heading Line.ipynb |
# ---
# jupyter:
# jupytext:
# cell_metadata_filter: -all
# formats: ipynb
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction #
#
# Most of the techniques we've seen in this course have been for numerical features. The technique we'll look at in this lesson, *target encoding*, is instead meant for categorical features. It's a method of encoding categories as numbers, like one-hot or label encoding, with the difference that it also uses the *target* to create the encoding. This makes it what we call a **supervised** feature engineering technique.
# +
import pandas as pd
autos = pd.read_csv("../input/fe-course-data/autos.csv")
# -
# # Target Encoding #
#
# A **target encoding** is any kind of encoding that replaces a feature's categories with some number derived from the target.
#
# A simple and effective version is to apply a group aggregation from Lesson 3, like the mean. Using the *Automobiles* dataset, this computes the average price of each vehicle's make:
# +
autos["make_encoded"] = autos.groupby("make")["price"].transform("mean")
autos[["make", "price", "make_encoded"]].head(10)
# -
# This kind of target encoding is sometimes called a **mean encoding**. Applied to a binary target, it's also called **bin counting**. (Other names you might come across include: likelihood encoding, impact encoding, and leave-one-out encoding.)
#
# # Smoothing #
#
# An encoding like this presents a couple of problems, however. First are *unknown categories*. Target encodings create a special risk of overfitting, which means they need to be trained on an independent "encoding" split. When you join the encoding to future splits, Pandas will fill in missing values for any categories not present in the encoding split. These missing values you would have to impute somehow.
#
# Second are *rare categories*. When a category only occurs a few times in the dataset, any statistics calculated on its group are unlikely to be very accurate. In the *Automobiles* dataset, the `mercurcy` make only occurs once. The "mean" price we calculated is just the price of that one vehicle, which might not be very representative of any Mercuries we might see in the future. Target encoding rare categories can make overfitting more likely.
#
# A solution to these problems is to add **smoothing**. The idea is to blend the *in-category* average with the *overall* average. Rare categories get less weight on their category average, while missing categories just get the overall average.
#
# In pseudocode:
# ```
# encoding = weight * in_category + (1 - weight) * overall
# ```
# where `weight` is a value between 0 and 1 calculated from the category frequency.
#
# An easy way to determine the value for `weight` is to compute an **m-estimate**:
# ```
# weight = n / (n + m)
# ```
# where `n` is the total number of times that category occurs in the data. The parameter `m` determines the "smoothing factor". Larger values of `m` put more weight on the overall estimate.
#
# <figure style="padding: 1em;">
# <img src="https://i.imgur.com/1uVtQEz.png" width=500, alt="">
# <figcaption style="textalign: center; font-style: italic"><center>
# </center></figcaption>
# </figure>
#
# In the *Automobiles* dataset there are three cars with the make `chevrolet`. If you chose `m=2.0`, then the `chevrolet` category would be encoded with 60% of the average Chevrolet price plus 40% of the overall average price.
# ```
# chevrolet = 0.6 * 6000.00 + 0.4 * 13285.03
# ```
#
# When choosing a value for `m`, consider how noisy you expect the categories to be. Does the price of a vehicle vary a great deal within each make? Would you need a lot of data to get good estimates? If so, it could be better to choose a larger value for `m`; if the average price for each make were relatively stable, a smaller value could be okay.
#
# <blockquote style="margin-right:auto; margin-left:auto; background-color: #ebf9ff; padding: 1em; margin:24px;">
# <strong>Use Cases for Target Encoding</strong><br>
# Target encoding is great for:
# <ul>
# <li><strong>High-cardinality features</strong>: A feature with a large number of categories can be troublesome to encode: a one-hot encoding would generate too many features and alternatives, like a label encoding, might not be appropriate for that feature. A target encoding derives numbers for the categories using the feature's most important property: its relationship with the target.
# <li><strong>Domain-motivated features</strong>: From prior experience, you might suspect that a categorical feature should be important even if it scored poorly with a feature metric. A target encoding can help reveal a feature's true informativeness.
# </ul>
# </blockquote>
#
# # Example - MovieLens1M #
#
# The [*MovieLens1M*](https://www.kaggle.com/grouplens/movielens-20m-dataset) dataset contains one-million movie ratings by users of the MovieLens website, with features describing each user and movie. This hidden cell sets everything up:
# +
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
import warnings
plt.style.use("seaborn-whitegrid")
plt.rc("figure", autolayout=True)
plt.rc(
"axes",
labelweight="bold",
labelsize="large",
titleweight="bold",
titlesize=14,
titlepad=10,
)
warnings.filterwarnings('ignore')
df = pd.read_csv("../input/fe-course-data/movielens1m.csv")
df = df.astype(np.uint8, errors='ignore') # reduce memory footprint
print("Number of Unique Zipcodes: {}".format(df["Zipcode"].nunique()))
# -
# With over 3000 categories, the `Zipcode` feature makes a good candidate for target encoding, and the size of this dataset (over one-million rows) means we can spare some data to create the encoding.
#
# We'll start by creating a 25% split to train the target encoder.
# +
X = df.copy()
y = X.pop('Rating')
X_encode = X.sample(frac=0.25)
y_encode = y[X_encode.index]
X_pretrain = X.drop(X_encode.index)
y_train = y[X_pretrain.index]
# -
# The `category_encoders` package in `scikit-learn-contrib` implements an m-estimate encoder, which we'll use to encode our `Zipcode` feature.
# +
from category_encoders import MEstimateEncoder
# Create the encoder instance. Choose m to control noise.
encoder = MEstimateEncoder(cols=["Zipcode"], m=5.0)
# Fit the encoder on the encoding split.
encoder.fit(X_encode, y_encode)
# Encode the Zipcode column to create the final training data
X_train = encoder.transform(X_pretrain)
# -
# Let's compare the encoded values to the target to see how informative our encoding might be.
plt.figure(dpi=90)
ax = sns.distplot(y, kde=False, norm_hist=True)
ax = sns.kdeplot(X_train.Zipcode, color='r', ax=ax)
ax.set_xlabel("Rating")
ax.legend(labels=['Zipcode', 'Rating']);
# The distribution of the encoded `Zipcode` feature roughly follows the distribution of the actual ratings, meaning that movie-watchers differed enough in their ratings from zipcode to zipcode that our target encoding was able to capture useful information.
#
# # Your Turn #
#
# [**Apply target encoding**](#$NEXT_NOTEBOOK_URL$) to features in *Ames* and investigate a surprising way that target encoding can lead to overfitting.
| notebooks/feature_engineering_new/raw/tut6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# [](https://www.pythonista.io)
# # *IPython* y *Jupyter*.
# ## El proyecto *IPython*.
# El proyecto [*IPython*](https://ipython.org/) es un entorno (shell) interactivo de Python que extiende las funcionalidades del intérprete interactivo de Python, permitiendo entre otras cosas:
#
# * Coloreado del código.
# * Autocompletado de comandos.
# * Historial de comandos.
# * Cómputo paralelo y concurrente.
# * Capacidad de desplegar los gráficos generados por [*Matplotlib*](https://matplotlib.org/) en la misma ventana.
#
# *IPython* puede ser accedido desde 3 interfaces:
#
# * Un shell dentro de una terminal de texto.
# * Un shell dentro de una aplicación de escritorio basada en la biblioteca [*QT*](https://www.qt.io/).
# * Una notebook de *Jupyter*.
# ## El proyecto *Jupyter*.
#
# En el proceso de desarrollo de *IPython* se creó una interfaz basada en web capaz de ofrecer las funcionalidades de *IPython* de forma más amigable.
#
# La interfaz web de *IPython* evolucionó de tal forma que la matéfora de una libreta o "notebook" no sólo podía conectarse a un intérprete de *IPython*, sino que podía adapatarse a otros lenguajes de programación mediante "kernels". Es así que a partir de *IPython* surgió el proyecto [*Jupyter*](https://jupyter.org/) (su nombre es una abreviación de [*JUlia*](https://julialang.org/), *PYThon* y [*R*](https://www.r-project.org/).
#
# *Jupyter* es servidor web capaz de desplegar una interfaz que incluye:
#
# * Un gestor de archivos.
# * Editor de texto.
# * Un emulador de terminal.
# * Entornos interactivos basados en notebooks.
# * Un gestor de notebooks.
#
# ### Las notebooks.
#
# Las notebooks de *Jupyter* pueden combinar dos tipos primordiales de celdas:
# * Celdas de texto capaces de desplegar código basado en la sintaxis de [*Markdown*](https://markdown.es/).
# * Celdas de código ejecutable concapacidad de desplegar gráficos e imágenes, entre otras muchas funcionalidades.
#
# *Jupyter* puede mantener múltiples notebooks y terminales ejecutándose simultáneamente e incluso generar clusters de servidores para cómputo intensivo.
#
# Actualmente las notebooks de *Jupyter* soportan [kernels de diversos lenguajes](https://github.com/jupyter/jupyter/wiki/Jupyter-kernels).
#
# Las notebooks de *Jupyter* pueden ser exportadas en múltiples formatos, tales como *HTML*, *PDF*, [*reStructuredText*](http://docutils.sourceforge.net/rst.html), [*LaTeX*](https://www.latex-project.org/), [*asciidocs*](http://asciidoc.org/), e incluso [*Reveal.js*](https://revealjs.com).
# ## Los comandos mágicos de *IPython*.
#
# Todos los apuntes de este curso son notebooks de *Jupyter* que utilizan el kernel de *IPython*.
#
# Las notebooks de *Jupyter* son interfaces basada en web capaces de ejecutar comandos por medio de un "kernel", el cual permite ligar a la interfaz con un intérprete específico.
#
# El "kernel" de [*IPython*](https://ipython.org/) es el kernel que utiliza *Jupyter* por defecto y corresponde a un intérprete avanzado de Python el cual además de ejecutar expresiones y delcaraciones de Python puede ejecutar sus propios "comandos mágicos" o "magics". Para conocer más al respecto puede consultar la siguiente liga: http://ipython.readthedocs.io/en/stable/interactive/magics.html
#
# Los comandos mágicos de *IPython* vienen precedidos generalmente por uno o dos signos de porcentaje ```%```.
# ### Ejecución de un script desde una notebook de *Jupyter*.
#
# Para ejecutar un archivo de Python desde una notebook de *Jupyter* se utiliza la instrucción de *IPython* ```%run```, la cual permite ejecutar el script localizado en la ruta que se añade posteriormente.
#
# La sintaxis es la siguiente:
#
# ```
# # %run <ruta del script>
# ```
# **Ejemplos:**
# * La siguiente celda ejecutará el script localizado en [```src/holamundo.py```](src/holamundo.py).
# %run src/holamundo.py
# * La siguiente celda ejecutará el script localizado en [```src/holamundo2.py```](src/holamundo2.py), el cual desplegará un mensaje y abrirá una ventana que aceptará todos los caracteres que se escriban hasta que se presione la tecla <kbd>Intro</kbd>.
# %run src/holamundo2.py
# ## Instrucciones a la línea de comandos desde *Jupyter*.
#
# Además de los comandos mágicos, *IPython* permite ejecutar comandos directamente al sistema operativo desde el que está corriendo el servidor de *Jupyter* mediante la siguiente sintaxis:
#
# ```
# !<comando>
# ```
# Donde:
#
# * ```<comando``` es un comando válido para el sistema operativo desde el cual se ejecuta *Jupyter*.
#
# **Advertencia:** Sólo es posible ejecutar comandos que se ejecuten sin necesidad de mayor interacción con el sistema. Si existe algún tipo de interacción, la celda de ejecución se quedará bloqueada.
# **Ejemplo:**
# Se ejecutará el comando del sistema que enlista los contenidos de un directorio.
# * Para *GNU/Linux* o *MacOS X*:
# !ls
# * Para *Windows*:
# !dir
# <p style="text-align: center"><a rel="license" href="http://creativecommons.org/licenses/by/4.0/"><img alt="Licencia Creative Commons" style="border-width:0" src="https://i.creativecommons.org/l/by/4.0/80x15.png" /></a><br />Esta obra está bajo una <a rel="license" href="http://creativecommons.org/licenses/by/4.0/">Licencia Creative Commons Atribución 4.0 Internacional</a>.</p>
# <p style="text-align: center">© <NAME>. 2021.</p>
| 02_ipython_y_jupyter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import argparse
import logging
import time
import cv2
import numpy as np
from tf_pose.estimator import TfPoseEstimator
from tf_pose.networks import get_graph_path, model_wh
logger = logging.getLogger('TfPoseEstimator-WebCam')
logger.setLevel(logging.DEBUG)
ch = logging.StreamHandler()
ch.setLevel(logging.DEBUG)
formatter = logging.Formatter('[%(asctime)s] [%(name)s] [%(levelname)s] %(message)s')
ch.setFormatter(formatter)
logger.addHandler(ch)
fps_time = 0
if __name__ == '__main__':
parser = argparse.ArgumentParser(description='tf-pose-estimation realtime webcam')
parser.add_argument('--camera', type=int, default=0)
parser.add_argument('--resize', type=str, default='432x368',
help='if provided, resize images before they are processed. default=0x0, Recommends : 432x368 or 656x368 or 1312x736 ')
parser.add_argument('--resize-out-ratio', type=float, default=4.0,
help='if provided, resize heatmaps before they are post-processed. default=1.0')
parser.add_argument('--model', type=str, default='mobilenet_thin', help='cmu / mobilenet_thin')
parser.add_argument('--show-process', type=bool, default=False,
help='for debug purpose, if enabled, speed for inference is dropped.')
args = parser.parse_args()
logger.debug('initialization %s : %s' % (args.model, get_graph_path(args.model)))
w, h = model_wh(args.resize)
if w > 0 and h > 0:
e = TfPoseEstimator(get_graph_path(args.model), target_size=(w, h))
else:
e = TfPoseEstimator(get_graph_path(args.model), target_size=(432, 368))
logger.debug('cam read+')
cam = cv2.VideoCapture(args.camera)
ret_val, image = cam.read()
logger.info('cam image=%dx%d' % (image.shape[1], image.shape[0]))
idx = 0
while True:
idx = idx + 1
ret_val, image = cam.read()
cv2.imwrite('output_images/tf_pose_' + str(idx) + '_raw.jpg', image)
logger.debug('image process+')
humans = e.inference(image, resize_to_default=(w > 0 and h > 0), upsample_size=args.resize_out_ratio)
logger.debug('postprocess+')
image = TfPoseEstimator.draw_humans(image, humans, imgcopy=False)
logger.debug('show+')
cv2.putText(image,
"FPS: %f" % (1.0 / (time.time() - fps_time)),
(10, 10), cv2.FONT_HERSHEY_SIMPLEX, 0.5,
(0, 255, 0), 2)
cv2.imwrite('output_images/tf_pose_' + str(idx) + '_processed.jpg', image)
cv2.imshow('tf-pose-estimation result', image)
fps_time = time.time()
if cv2.waitKey(1) == 27:
break
logger.debug('finished+')
cv2.destroyAllWindows()
# -
| run.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tutorial notebook to run vis_cpu
# In this notebook we will set up different input parameters required for **vis_cpu**, and finally run the **vis_cpu** to generate visibilities for point sources. Also, we will plot the generated visibility amplitude and phase as a function of frequency and LST.
# +
import numpy as np
import matplotlib.pyplot as plt
from astropy.time import Time
from astropy.coordinates import EarthLocation
from vis_cpu import conversions, simulate_vis
from pyuvsim.analyticbeam import AnalyticBeam
# -
# # Initialize Observation parameters
# Set the antenna dictionary which contains the antenna index and co-ordinates. We also set different primary beam (PB) of each antennas using a list of UVBeam object (here Gaussian and Uniform PB are used for 2 antennas).
# +
nants = 2
ants = {}
for i in range(nants):
ants.update([(i, (-(i % 3), i, i % 4))])
#set different UVBeam for each antenna
beam = [AnalyticBeam("gaussian", sigma=0.5), AnalyticBeam("uniform")]
# -
# Set the input parameters (e.g. no. of frequencies, start freq (in Hz), end fequency (in Hz), no. of time samples, start lst (in rad), end lst (in rad).
Nfreqs = 10
start_freq = 100000000.0 # in Hz
end_freq = 120000000.0 # in Hz
freqs = np.linspace(start_freq , end_freq, Nfreqs) # freqs in Hz
Ntimes = 20
start_lst = np.pi / 1.5 # in rad
end_lst = np.pi / 1.3 # in rad
lsts = np.linspace(start_lst, end_lst, Ntimes) # lsts in rad
# # Initialize source parameters
# Set the source parameters (total no. of point sources, ra (rad), dec (rad), flux (in Jy) and sp. index
# +
nsource = 2
ra = np.deg2rad(np.linspace(0. , 30., nsource)) # ra of each source (in rad)
dec = np.deg2rad(np.linspace(-25. , -35., nsource)) # dec of each source (in rad)
flux = np.ones(nsource) # flux of each sources (in Jy)
sp_index = np.zeros(nsource) # sp. index of each source
flux_allfreq = ((freqs[:, np.newaxis] / freqs[0]) ** sp_index.T * flux.T).T
# -
# # Correct source locations so that vis_cpu uses the right frame
# Set telescope parameters (longitude (in deg), latitude (in deg), height (in meter). It converts the ra, dec into the ECI (Earth-Centered Inertial) system used by vis_cpu.
# +
hera_lon = 21.428305555555557
hera_lat = -30.72152777777791
hera_height = 1073.0000000093132
location = EarthLocation.from_geodetic(lat=hera_lat, lon=hera_lon, height=hera_height)
# +
obstime = Time("2018-08-31T04:02:30.11", format="isot", scale="utc")
ra_new, dec_new = conversions.equatorial_to_eci_coords(
ra, dec, obstime, location, unit="rad", frame="icrs"
)
# -
# # Run vis_cpu
# Use all the paramters above to run vis_cpu, which ouputs a complex visibility array. If `polarized = True`, the output will have shape (NAXES, NFEED, NFREQS, NTIMES, NANTS, NANTS), otherwise it will have
# shape (NFREQS, NTIMES, NANTS, NANTS)
# simulate visibilities
vis_vc = simulate_vis(
ants=ants,
fluxes=flux_allfreq,
ra=ra_new,
dec=dec_new,
freqs=freqs,
lsts=lsts,
beams=beam,
pixel_beams=False,
polarized=False,
precision=2,
)
vis_vc.shape
# # Plot auto and cross visibilities
# Here, we plot the visibility amplitude and phase as a function of frequency and LSTs. For auto-correlation (e.g. antenna pair (0,0)) we expect the phase would be zero. Here, the amplitude is constant with frequency as we assumed the spectral of the sources are zero.
# +
fig, ax = plt.subplots(1, 2, figsize=(16, 6))
ax[0].plot(freqs/1.e6, np.abs(vis_vc[:,0,0,0]),'k', label=r'Auto (ant_pair : 0,0)')
ax[0].set_ylabel('Amp', fontsize=15,labelpad=10)
ax[0].set_xlabel(r'freqs [MHz]', fontsize=15)
ax[0].legend(fontsize=15)
ax[1].plot(freqs/1.e6, np.angle(vis_vc[:,0,0,0]),'k', label=r'Auto (ant_pair : 0,0)')
ax[1].set_ylabel('Phase', fontsize=15,labelpad=10)
ax[1].set_xlabel(r'freqs [MHz]', fontsize=15)
ax[1].legend(fontsize=15)
plt.show()
# +
fig, ax = plt.subplots(1, 2, figsize=(16, 6))
ax[0].plot(freqs/1.e6, np.abs(vis_vc[:,0,0,1]),'r', label=r'Cross (ant_pair : 0,1)')
ax[0].set_ylabel('Amp', fontsize=15)
ax[0].set_xlabel(r'freqs [MHz]', fontsize=15)
ax[0].legend(fontsize=15)
ax[1].plot(freqs/1.e6, np.angle(vis_vc[:,0,0,1]),'r', label=r'Cross (ant_pair : 0,1)')
ax[1].set_ylabel('Phase', fontsize=15)
ax[1].set_xlabel(r'freqs [MHz]', fontsize=15)
ax[1].legend(fontsize=15)
plt.show()
# +
fig, ax = plt.subplots(1, 2, figsize=(16, 6))
ax[0].plot(lsts, np.abs(vis_vc[0,:,0,0]),'k', label=r'Auto (ant_pair : 0,0)')
ax[0].set_ylabel('Amp', fontsize=15,labelpad=10)
ax[0].set_xlabel(r'LST [rad]', fontsize=15)
ax[0].legend(fontsize=15)
ax[1].plot(lsts, np.angle(vis_vc[0,:,0,0]),'k', label=r'Auto (ant_pair : 0,0)')
ax[1].set_ylabel('Phase', fontsize=15,labelpad=10)
ax[1].set_xlabel(r'LST [rad]', fontsize=15)
ax[1].legend(fontsize=15)
plt.show()
# +
fig, ax = plt.subplots(1, 2, figsize=(16, 6))
ax[0].plot(lsts, np.abs(vis_vc[0,:,0,1]),'r', label=r'Cross (ant_pair : 0,1)')
ax[0].set_ylabel('Amp', fontsize=15)
ax[0].set_xlabel(r'LST [rad]', fontsize=15)
ax[0].legend(fontsize=15)
ax[1].plot(lsts, np.angle(vis_vc[0,:,0,1]),'r', label=r'Cross (ant_pair : 0,1)')
ax[1].set_ylabel('Phase', fontsize=15)
ax[1].set_xlabel(r'LST [rad]', fontsize=15)
ax[1].legend(fontsize=15)
plt.show()
# -
# In the **hera_sim** repo there is wrapper which can be used to set up these operations easily, and also allows you to run vis_cpu based on pyuvsim-compatible configuration.
| docs/tutorials/vis_cpu_tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
import numpy as np
from pandas.plotting import register_matplotlib_converters
register_matplotlib_converters()
df=pd.read_csv('fcc-forum-pageviews.csv',parse_dates=True, index_col='date')
df
df=df[(df.value >= df.value.quantile(0.025)) & (df.value <= df.value.quantile(0.975))]
df
# +
def draw_line_plot():
fig,ax = plt.subplots(figsize=(14,6))
ax.plot(df, color="red")
ax.set_xlabel('Date')
ax.set_ylabel('Page Views')
ax.set_title("Daily freeCodeCamp Forum Page Views 5/2016-12/2019")
fig.savefig('LinePlot.png')
draw_line_plot()
# -
df_bar=df.copy()
df_bar
df_bar['Year']=df_bar.index.year
df_bar['Month']=df_bar.index.month
df_bar['Month'] = pd.to_datetime(df_bar['Month'], format='%m').dt.month_name()
df_bar
pd.DataFrame(df_bar.groupby(['Year','Month'])['value'].mean())
# +
month=['January','February','March','April','May','June','July','August','September','October','November','December']
bar=sns.catplot(data=df_bar,x='Year',y='value',kind='bar',hue='Month',ci=None,palette="tab10", legend=False, hue_order=month)
fig=bar.fig
ax=bar.ax
ax.set_xlabel('Years')
ax.set_ylabel('Average Page Views')
ax.legend(loc='upper left', title='Months', fontsize=8)
plt.xticks(rotation=90)
plt.tight_layout()
fig.savefig('Bargraph.png')
# -
df_box = df.copy()
df_box.reset_index(inplace=True)
df_box['year'] = [d.year for d in df_box.date]
df_box['month'] = [d.strftime('%b') for d in df_box.date]
df_box
df_box.sort_values(by=['year','date'], ascending=[False, True], inplace=True)
df_box
# +
fig, ax=plt.subplots(1,2 , figsize=(20,6))
sns.boxplot(data=df_box, y="value", x="year",ax=ax[0])
sns.boxplot(data=df_box, y="value", x="month",ax=ax[1])
ax[0].set_ylabel('Page Views')
ax[1].set_ylabel('Page Views')
ax[0].set_xlabel('Year')
ax[1].set_xlabel('Month')
ax[0].set_title('Year-wise Box Plot (Trend)')
ax[1].set_title('Month-wise Box Plot (Seasonality)')
plt.tight_layout()
# -
fig.savefig('Boxplot.png')
| fcc-time-series-forum/time-series-visualiser-fcc.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ###### Content under Creative Commons Attribution license CC-BY 4.0, code under MIT license (c)2014 <NAME>, <NAME>, <NAME>.
# # Spreading out
# Welcome to the fifth, and last, notebook of Module 4 "_Spreading out: diffusion problems,"_ of our fabulous course **"Practical Numerical Methods with Python."**
#
# In this course module, we have learned about explicit and implicit methods for parabolic equations in 1 and 2 dimensions. So far, all schemes have been first-order in time and second-order in space. _Can we do any better?_ We certainly can: this notebook presents the Crank-Nicolson scheme, which is a second-order method in both time and space! We will continue to use the heat equation to guide the discussion, as we've done throughout this module.
# ## Crank-Nicolson scheme
# The [Crank Nicolson scheme](http://en.wikipedia.org/wiki/Crank–Nicolson_method) is a popular second-order, implicit method used with parabolic PDEs in particular. It was developed by <NAME> and [<NAME>](http://en.wikipedia.org/wiki/Phyllis_Nicolson). The main idea is to take the average between the solutions at $t^n$ and $t^{n+1}$ in the evaluation of the spatial derivative. Why bother doing that? Because the time derivative will then be discretized with a centered scheme, giving second-order accuracy!
#
# Remember the 1D heat equation from the [first notebook](https://nbviewer.jupyter.org/github/numerical-mooc/numerical-mooc/blob/master/lessons/04_spreadout/04_01_Heat_Equation_1D_Explicit.ipynb)? Just to refresh your memory, here it is:
#
# $$
# \begin{equation}
# \frac{\partial T}{\partial t} = \alpha \frac{\partial^2 T}{\partial x^2}.
# \end{equation}
# $$
#
# In this case, the Crank-Nicolson scheme leads to the following discretized equation:
#
# $$
# \begin{equation}
# \begin{split}
# & \frac{T^{n+1}_i - T^n_i}{\Delta t} = \\
# & \quad \alpha \cdot \frac{1}{2} \left( \frac{T^{n+1}_{i+1} - 2 T^{n+1}_i + T^{n+1}_{i-1}}{\Delta x^2} + \frac{T^n_{i+1} - 2 T^n_i + T^n_{i-1}}{\Delta x^2} \right) \\
# \end{split}
# \end{equation}
# $$
#
# Notice how the both time indices $n$ and $n+1$ appear on the right-hand side. You know we'll have to rearrange this equation, right? Now look at the stencil and notice that we are using more information than before in the update.
# 
# #### Figure 2. Stencil of the Crank-Nicolson scheme.
# Rearranging terms so that everything that we don't know is on the left side and what we do know on the right side, we get
#
# $$
# \begin{equation}
# \begin{split}
# & -T^{n+1}_{i-1} + 2 \left( \frac{\Delta x^2}{\alpha \Delta t} + 1 \right) T^{n+1}_i - T^{n+1}_{i+1} \\
# & \qquad = T^{n}_{i-1} + 2 \left( \frac{\Delta x^2}{\alpha \Delta t} - 1 \right) T^{n}_i + T^{n}_{i+1} \\
# \end{split}
# \end{equation}
# $$
#
# Again, we are left with a linear system of equations. Check out the left side of that equation: it looks a lot like the matrix from [notebook 2](https://nbviewer.jupyter.org/github/numerical-mooc/numerical-mooc/blob/master/lessons/04_spreadout/04_02_Heat_Equation_1D_Implicit.ipynb), doesn't it? Apart from the slight modification in the $T_i^{n+1}$ term, the left side of the equation is pretty much the same. What about the right-hand side? Sure, it looks quite different, but that is not a problem, we know all those terms!
#
# Things don't change much for boundary conditions, either. We've seen all the cases already. Say $T_0^{n+1}$ is a Dirichlet boundary. Then the equation for $i=1$ becomes
#
# $$
# \begin{equation}
# \begin{split}
# & 2 \left( \frac{\Delta x^2}{\alpha \Delta t} + 1 \right) T^{n+1}_1 - T^{n+1}_{2} \\
# & \qquad = T^{n}_{0} + 2 \left( \frac{\Delta x^2}{\alpha \Delta t} - 1 \right) T^{n}_1 + T^{n}_{2} + T^{n+1}_{0} \\
# \end{split}
# \end{equation}
# $$
#
# And if we have a Neumann boundary $\left(\left.\frac{\partial T}{\partial x}\right|_{x=L} = q\right)$ at $T_{n_x-1}^{n+1}$? We know this stuff, right? For $i=n_x-2$ we get
#
# $$
# \begin{equation}
# \begin{split}
# & -T^{n+1}_{n_x-3} + \left( 2 \frac{\Delta x^2}{\alpha \Delta t} + 1 \right) T^{n+1}_{n_x-2} \\
# & \qquad = T^{n}_{n_x-3} + 2 \left( \frac{\Delta x^2}{\alpha \Delta t} - 1 \right) T^{n}_{n_x-2} + T^{n}_{n_x-1} + q\Delta x \\
# \end{split}
# \end{equation}
# $$
#
# The code will look a lot like the implicit method from the [second notebook](https://nbviewer.jupyter.org/github/numerical-mooc/numerical-mooc/blob/master/lessons/04_spreadout/04_02_Heat_Equation_1D_Implicit.ipynb). Only some terms of the matrix and right-hand-side vector will be different, which changes some of our custom functions.
# ### The linear system
# Just like in [notebook 2](https://nbviewer.jupyter.org/github/numerical-mooc/numerical-mooc/blob/master/lessons/04_spreadout/04_02_Heat_Equation_1D_Implicit.ipynb), we need to solve a linear system on every time step of the form:
#
# $$
# [A][T^{n+1}_\text{int}] = [b]+[b]_{b.c.}
# $$
#
# The coefficient matrix is very similar to the previous case, but the right-hand side changes a lot:
# $$
# \begin{align}
# \left[
# \begin{array}{cccccc}
# 2 \left( \frac{1}{\sigma} + 1 \right) & -1 & 0 & \cdots & & 0 \\
# -1 & 2 \left( \frac{1}{\sigma} + 1\right) & -1 & 0 & \cdots & 0 \\
# 0 & & \ddots & & & \vdots \\
# \vdots & & & & 2 \left( \frac{1}{\sigma} + 1\right) & \\
# 0 & \cdots & & & -1 & \left( 2 \frac{1}{\sigma} + 1\right) \\
# \end{array}
# \right] \cdot
# \left[
# \begin{array}{c}
# T_1^{n+1} \\
# T_2^{n+1} \\
# \vdots \\
# \\
# T_{N-2}^{n+1} \\
# \end{array}
# \right] =
# \left[
# \begin{array}{c}
# T_0^n + 2 \left( \frac{1}{\sigma} - 1 \right) T_1^n + T_2^n \\
# T_1^n + 2 \left( \frac{1}{\sigma} - 1 \right) T_2^n + T_3^n \\
# \vdots \\
# \\
# T_{n_x-3}^n + 2 \left( \frac{1}{\sigma} - 1 \right) T_{n_x-2}^n + T_{n_x-1}^n \\
# \end{array}
# \right] +
# \begin{bmatrix}
# T_0^{n+1} \\
# 0\\
# \vdots \\
# 0 \\
# q \Delta x \\
# \end{bmatrix}
# \end{align}
# $$
# Let's write a function that will create the coefficient matrix and right-hand-side vectors for the heat conduction problem from [notebook 2](https://nbviewer.jupyter.org/github/numerical-mooc/numerical-mooc/blob/master/lessons/04_spreadout/04_02_Heat_Equation_1D_Implicit.ipynb): with Dirichlet boundary at $x=0$ and zero-flux boundary $(q=0)$ at $x=L$.
import numpy
from scipy import linalg
def lhs_operator(N, sigma):
"""
Computes and returns the implicit operator
of the system for the 1D diffusion equation.
We use Crank-Nicolson method, Dirichlet condition
on the left side of the domain and zero-gradient
Neumann condition on the right side.
Parameters
----------
N : integer
Number of interior points.
sigma : float
Value of alpha * dt / dx**2.
Returns
-------
A : numpy.ndarray
The implicit operator as a 2D array of floats
of size N by N.
"""
# Setup the diagonal of the operator.
D = numpy.diag(2.0 * (1.0 + 1.0 / sigma) * numpy.ones(N))
# Setup the Neumann condition for the last element.
D[-1, -1] = 1.0 + 2.0 / sigma
# Setup the upper diagonal of the operator.
U = numpy.diag(-1.0 * numpy.ones(N - 1), k=1)
# Setup the lower diagonal of the operator.
L = numpy.diag(-1.0 * numpy.ones(N - 1), k=-1)
# Assemble the operator.
A = D + U + L
return A
def rhs_vector(T, sigma, qdx):
"""
Computes and returns the right-hand side of the system
for the 1D diffusion equation, using a Dirichlet condition
on the left side and a Neumann condition on the right side.
Parameters
----------
T : numpy.ndarray
The temperature distribution as a 1D array of floats.
sigma : float
Value of alpha * dt / dx**2.
qdx : float
Value of the temperature flux at the right side.
Returns
-------
b : numpy.ndarray
The right-hand side of the system as a 1D array of floats.
"""
b = T[:-2] + 2.0 * (1.0 / sigma - 1.0) * T[1:-1] + T[2:]
# Set Dirichlet condition.
b[0] += T[0]
# Set Neumann condition.
b[-1] += qdx
return b
# We will solve the linear system at every time step. Let's define a function to step in time:
def crank_nicolson(T0, nt, dt, dx, alpha, q):
"""
Computes and returns the temperature along the rod
after a given number of time steps.
The function uses Crank-Nicolson method in time,
central differencing in space, a Dirichlet condition
on the left side, and a Neumann condition on the
right side.
Parameters
----------
T0 : numpy.ndarray
The initial temperature distribution as a 1D array of floats.
nt : integer
Number of time steps to compute.
dt : float
Time-step size.
dx : float
Distance between two consecutive locations.
alpha : float
Thermal diffusivity of the rod.
q : float
Value of the temperature gradient on the right side.
Returns
-------
T : numpy.ndarray
The temperature distribution as a 1D array of floats.
"""
sigma = alpha * dt / dx**2
# Create the implicit operator of the system.
A = lhs_operator(len(T0) - 2, sigma)
# Integrate in time.
T = T0.copy()
for n in range(nt):
# Generate the right-hand side of the system.
b = rhs_vector(T, sigma, q * dx)
# Solve the system with scipy.linalg.solve.
T[1:-1] = linalg.solve(A, b)
# Apply the Neumann boundary condition.
T[-1] = T[-2] + q * dx
return T
# And we are good to go! First, let's setup our initial conditions, and the matrix
# +
# Set parameters.
L = 1.0 # length of the rod
nx = 21 # number of points on the rod
dx = L / (nx - 1) # grid spacing
alpha = 1.22e-3 # thermal diffusivity of the rod
q = 0.0 # temperature gradient at the extremity
# Define the locations on the rod.
x = numpy.linspace(0.0, L, num=nx)
# Set the initial temperature distribution.
T0 = numpy.zeros(nx)
T0[0] = 100.0
# -
# Check the matrix...
A = lhs_operator(nx - 1, 0.5)
print(A)
# Looks okay! Now, step in time
# +
# Set the time-step size based on CFL limit.
sigma = 0.5
dt = sigma * dx**2 / alpha # time-step size
nt = 10 # number of time steps to compute
# Compute the temperature distribution.
T = crank_nicolson(T0, nt, dt, dx, alpha, q)
# -
# And plot,
from matplotlib import pyplot
# %matplotlib inline
# Set the font family and size to use for Matplotlib figures.
pyplot.rcParams['font.family'] = 'serif'
pyplot.rcParams['font.size'] = 16
# Plot the temperature along the rod.
pyplot.figure(figsize=(6.0, 4.0))
pyplot.xlabel('Distance [m]')
pyplot.ylabel('Temperature [C]')
pyplot.grid()
pyplot.plot(x, T, color='C0', linestyle='-', linewidth=2)
pyplot.xlim(0.0, L)
pyplot.ylim(0.0, 100.0);
# Works nicely. But wait! This method has elements of explicit and implicit discretizations. Is it *conditionally stable* like forward Euler, or *unconditionally stable* like backward Euler? Try out different values of `sigma`. You'll see Crank-Nicolson is an *unconditionally stable scheme* for the diffusion equation!
# ## Accuracy & convergence
# Using some techniques you might have learned in your PDE class, such as separation of variables, you can get a closed expression for the rod problem. It looks like this:
#
# $$
# \begin{eqnarray}
# T(x,t) = & \nonumber \\
# 100 - \sum_{n=1}^{\infty} & \frac{400}{(2n-1)\pi}\sin\left(\frac{(2n-1)\pi}{2L}x\right) \exp\left[-\alpha\left(\frac{(2n-1)\pi}{2L}\right)^2t\right]
# \end{eqnarray}
# $$
#
# Unfortunately, the analytical solution is a bit messy, but at least it gives a good approximation if we evaluate it for large $n$. Let's define a function that will calculate this for us:
def analytical_temperature(x, t, alpha, L, N):
"""
Computes and returns a truncated approximation
of the exact temperature distribution along the rod.
Parameters
----------
x : numpy.ndarray
Locations at which to calculate the temperature
as a 1D array of floats.
t : float
Time.
alpha : float
Thermal diffusivity of the rod.
L : float
Length of the rod.
N : integer
Number of terms to use in the expansion.
Returns
-------
T : numpy.ndarray
The truncated analytical temperature distribution
as a 1D array of floats.
"""
T = 100.0 * numpy.ones_like(x)
for n in range(1, N + 1):
k = (2 * n - 1) * numpy.pi / (2.0 * L)
T -= (400.0 / (2.0 * L * k) *
numpy.sin(k * x) * numpy.exp(- alpha * k**2 * t))
return T
# And let's see how that expression looks for the time where we left the numerical solution
# +
# Compute the analytical temperature distribution.
T_exact = analytical_temperature(x, nt * dt, alpha, L, 100)
# Plot the numerical and analytical temperatures.
pyplot.figure(figsize=(6.0, 4.0))
pyplot.xlabel('Distance [m]')
pyplot.ylabel('Temperature [C]')
pyplot.grid()
pyplot.plot(x, T, label='numerical',
color='C0', linestyle='-', linewidth=2)
pyplot.plot(x, T_exact, label='analytical',
color='C1', linestyle='--', linewidth=2)
pyplot.legend()
pyplot.xlim(0.0, L)
pyplot.ylim(0.0, 100.0);
# -
T1 = analytical_temperature(x, 0.2, alpha, L, 100)
T2 = analytical_temperature(x, 0.2, alpha, L, 200)
numpy.sqrt(numpy.sum((T1 - T2)**2) / numpy.sum(T2**2))
# That looks like it should. We'll now use this result to study the convergence of the Crank-Nicolson scheme.
# ### Time convergence
# We said this method was second-order accurate in time, remember? That's in theory, but we should test that the numerical solution indeed behaves like the theory says.
#
# Leaving $\Delta x$ constant, we'll run the code for different values of $\Delta t$ and compare the result at the same physical time, say $t=n_t\cdot\Delta t=10$, with the analytical expression above.
#
# The initial condition of the rod problem has a very sharp gradient: it suddenly jumps from $0{\rm C}$ to $100{\rm C}$ at the boundary. To resolve that gradient to the point that it doesn't affect time convergence, we would need a very fine mesh, and computations would be very slow. To avoid this issue, we will start from $t=1$ rather than starting from $t=0$.
#
# First, let's define a function that will compute the $L_2$-norm of the error:
def l2_error(T, T_exact):
"""
Computes and returns the relative L2-norm
of the difference between the numerical solution
and the exact solution.
Parameters
----------
T : numpy.ndarray
The numerical solution as an array of floats.
T_exact : numpy.ndarray
The exact solution as an array of floats.
Returns
-------
error : float
The relative L2-norm of the difference.
"""
error = numpy.sqrt(numpy.sum((T - T_exact)**2) /
numpy.sum(T_exact**2))
return error
# For fun, let's compare the Crank-Nicolson scheme with the implicit (a.k.a., backward) Euler scheme. We'll borrow some functions from [notebook 2](https://nbviewer.jupyter.org/github/numerical-mooc/numerical-mooc/blob/master/lessons/04_spreadout/04_02_Heat_Equation_1D_Implicit.ipynb) to do this.
def lhs_operator_btcs(N, sigma):
"""
Computes and returns the implicit operator
of the system for the 1D diffusion equation.
We use backward Euler method, Dirichlet condition
on the left side of the domain and zero-gradient
Neumann condition on the right side.
Parameters
----------
N : integer
Number of interior points.
sigma : float
Value of alpha * dt / dx**2.
Returns
-------
A : numpy.ndarray
The implicit operator as a 2D array of floats
of size N by N.
"""
# Setup the diagonal of the operator.
D = numpy.diag((2.0 + 1.0 / sigma) * numpy.ones(N))
# Setup the Neumann condition for the last element.
D[-1, -1] = 1.0 + 1.0 / sigma
# Setup the upper diagonal of the operator.
U = numpy.diag(-1.0 * numpy.ones(N - 1), k=1)
# Setup the lower diagonal of the operator.
L = numpy.diag(-1.0 * numpy.ones(N - 1), k=-1)
# Assemble the operator.
A = D + U + L
return A
def rhs_vector_btcs(T, sigma, qdx):
"""
Computes and returns the right-hand side of the system
for the 1D diffusion equation, using a Dirichlet condition
on the left side and a Neumann condition on the right side.
Parameters
----------
T : numpy.ndarray
The temperature distribution as a 1D array of floats.
sigma : float
Value of alpha * dt / dx**2.
qdx : float
Value of the temperature flux at the right side.
Returns
-------
b : numpy.ndarray
The right-hand side of the system as a 1D array of floats.
"""
b = T[1:-1] / sigma
# Set Dirichlet condition.
b[0] += T[0]
# Set Neumann condition.
b[-1] += qdx
return b
def btcs_implicit(T0, nt, dt, dx, alpha, q):
"""
Computes and returns the temperature along the rod
after a given number of time steps.
The function uses Euler implicit in time,
central differencing in space, a Dirichlet condition
on the left side, and a Neumann condition on the
right side.
Parameters
----------
T0 : numpy.ndarray
The initial temperature distribution
as a 1D array of floats.
nt : integer
Number of time steps to compute.
dt : float
Time-step size.
dx : float
Distance between two consecutive locations.
alpha : float
Thermal diffusivity of the rod.
q : float
Value of the temperature gradient on the right side.
Returns
-------
T : numpy.ndarray
The temperature distribution as a 1D array of floats.
"""
sigma = alpha * dt / dx**2
# Create the implicit operator of the system.
A = lhs_operator_btcs(len(T0) - 2, sigma)
# Integrate in time.
T = T0.copy()
for n in range(nt):
# Generate the right-hand side of the system.
b = rhs_vector_btcs(T, sigma, q * dx)
# Solve the system with scipy.linalg.solve.
T[1:-1] = linalg.solve(A, b)
# Apply the Neumann boundary condition.
T[-1] = T[-2] + q * dx
return T
# Now, let's do the runs!
# +
# Update parameters.
nx = 1001 # number of points on the rod
dx = L / (nx - 1) # grid spacing
# Define the locations on the rod.
x = numpy.linspace(0.0, L, num=nx)
# Create a list with the time-step sizes to use.
dt_values = [1.0, 0.5, 0.25, 0.125]
# Create empty lists to hold the errors for both schemes.
errors = []
errors_btcs = []
# Compute the initial temperature distribution at t=1.0.
t0 = 1.0
T0 = analytical_temperature(x, t0, alpha, L, 100)
# Compute the final analytical temperature at t=10.0.
t = 10.0
T_exact = analytical_temperature(x, t, alpha, L, 100)
# Compute the numerical solutions and errors.
for dt in dt_values:
nt = int((t - t0) / dt) # number of time steps
# Compute the solution using Crank-Nicolson scheme.
T = crank_nicolson(T0, nt, dt, dx, alpha, q)
# Compute and record the L2-norm of the error.
errors.append(l2_error(T, T_exact))
# Compute the solution using implicit BTCS scheme.
T = btcs_implicit(T0, nt, dt, dx, alpha, q)
# Compute and record the L2-norm of the error.
errors_btcs.append(l2_error(T, T_exact))
# -
# And plot,
# Plot the error versus the time-step size.
pyplot.figure(figsize=(6.0, 6.0))
pyplot.grid()
pyplot.xlabel(r'$\Delta t$')
pyplot.ylabel('Relative $L_2$-norm\nof the error')
pyplot.loglog(dt_values, errors, label='Crank-Nicolson',
color='black', linestyle='--', linewidth=2, marker='o')
pyplot.loglog(dt_values, errors_btcs, label='BTCS (implicit)',
color='black', linestyle='--', linewidth=2, marker='s')
pyplot.legend()
pyplot.axis('equal');
errors
# See how the error drops four times when the time step is halved? This method is second order in time!
#
# Clearly, Crank-Nicolson (circles) converges faster than backward Euler (squares)! Not only that, but also the error curve is shifted down: Crank-Nicolson is more accurate.
#
# If you look closely, you'll realize that the error in Crank-Nicolson decays about twice as fast than backward Euler: it's a second versus first order method!
# ### Spatial convergence
# To study spatial convergence, we will run the code for meshes with 21, 41, 81 and 161 points, and compare them at the same non-dimensional time, say $t=20$.
#
# Let's start by defining a function that will do everything for us
# +
# Set parameters.
dt = 0.1 # time-step size
t = 20.0 # final time
nt = int(t / dt) # number of time steps to compute
# Create a list with the grid-spacing sizes to use.
nx_values = [11, 21, 41, 81, 161]
# Create an empty list to store the errors.
errors = []
# Compute the numerical solutions and errors.
for nx in nx_values:
dx = L / (nx - 1) # grid spacing
x = numpy.linspace(0.0, L, num=nx) # grid points
# Set the initial conditions for the grid.
T0 = numpy.zeros(nx)
T0[0] = 100.0
# Compute the solution using Crank-Nicolson scheme.
T = crank_nicolson(T0, nt, dt, dx, alpha, q)
# Compute the analytical solution.
T_exact = analytical_temperature(x, t, alpha, L, 100)
# Compute and record the L2-norm of the error.
errors.append(l2_error(T, T_exact))
# -
# And plot!
# Plot the error versus the grid-spacing size.
pyplot.figure(figsize=(6.0, 6.0))
pyplot.grid()
pyplot.xlabel(r'$\Delta x$')
pyplot.ylabel('Relative $L_2$-norm\nof the error')
dx_values = L / (numpy.array(nx_values) - 1)
pyplot.loglog(dx_values, errors,
color='black', linestyle='--', linewidth=2, marker='o')
pyplot.axis('equal');
# That looks good! See how for each quadrant we go right, the error drops two quadrants going down (and even a bit better!).
# ##### Dig deeper
# Let's re-do the spatial convergence, but comparing at a much later time, say $t=1000$.
# +
# Set parameters.
dt = 0.1 # time-step size
t = 1000.0 # final time
nt = int(t / dt) # number of time steps to compute
# Create a list with the grid-spacing sizes to use.
nx_values = [11, 21, 41, 81, 161]
# Create an empty list to store the errors.
errors = []
# Compute the numerical solutions and errors.
for nx in nx_values:
dx = L / (nx - 1) # grid spacing
x = numpy.linspace(0.0, L, num=nx) # grid points
# Set the initial conditions for the grid.
T0 = numpy.zeros(nx)
T0[0] = 100.0
# Compute the solution using Crank-Nicolson scheme.
T = crank_nicolson(T0, nt, dt, dx, alpha, q)
# Compute the analytical solution.
T_exact = analytical_temperature(x, t, alpha, L, 100)
# Compute and record the L2-norm of the error.
errors.append(l2_error(T, T_exact))
# -
# Plot the error versus the grid-spacing size.
pyplot.figure(figsize=(6.0, 6.0))
pyplot.grid()
pyplot.xlabel(r'$\Delta x$')
pyplot.ylabel('Relative $L_2$-norm\nof the error')
dx_values = L / (numpy.array(nx_values) - 1)
pyplot.loglog(dx_values, errors,
color='black', linestyle='--', linewidth=2, marker='o')
pyplot.axis('equal');
errors
# Wait, convergence is not that great now! It's not as good as second order, but not as bad as first order. *What is going on?*
#
# Remember our implementation of the boundary conditions? We used
#
# $$
# \begin{equation}
# \frac{T^{n}_{N-1} - T^{n}_{N-2}}{\Delta x} = q
# \end{equation}
# $$
#
# Well, that is a **first-order** approximation!
#
# But, why doesn't this affect our solution at an earlier time? Initially, temperature on the right side of the rod is zero and the gradient is very small in that region; at that point in time, errors there were negligible. Once temperature starts picking up, we start having problems.
#
# **Boundary conditions can affect the convergence and accuracy of your solution!**
# ---
# ###### The cell below loads the style of the notebook
from IPython.core.display import HTML
css_file = '../../styles/numericalmoocstyle.css'
HTML(open(css_file, 'r').read())
| lessons/04_spreadout/04_05_Crank-Nicolson.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Project Start
#
from tqdm import tqdm
# +
import matplotlib.image as mpimg
import matplotlib.pyplot as plt
import numpy as np
import cv2
import glob
import os
#from skimage.feature import hog
#from skimage import color, exposure
# images are divided up into vehicles and non-vehicles
def get_image_list(path):
images_folder = path
image_array = []
image_folders = os.listdir(images_folder)
#print(image_folders)
for folder in image_folders :
image_array.extend(glob.glob(path+folder+'/*'))
#print ( path+folder)
print ("Images : " ,len(image_array))
return image_array
# Define a function to return some characteristics of the dataset
def data_look(car_list, notcar_list):
data_dict = {}
# Define a key in data_dict "n_cars" and store the number of car images
data_dict["n_cars"] = len(car_list)
# Define a key "n_notcars" and store the number of notcar images
data_dict["n_notcars"] = len(notcar_list)
# Read in a test image, either car or notcar
example_img = mpimg.imread(car_list[0])
# Define a key "image_shape" and store the test image shape 3-tuple
data_dict["image_shape"] = example_img.shape
# Define a key "data_type" and store the data type of the test image.
data_dict["data_type"] = example_img.dtype
# Return data_dict
return data_dict
# +
#cars = get_image_list('training_data/vehicles_smallset/')
#notcars = get_image_list('training_data/non-vehicles_smallset/')
cars = get_image_list('training_data/vehicles/')
notcars = get_image_list('training_data/non-vehicles/')
data_info = data_look(cars, notcars)
print('Your function returned a count of',
data_info["n_cars"], ' cars and',
data_info["n_notcars"], ' non-cars')
print('of size: ',data_info["image_shape"], ' and data type:',
data_info["data_type"])
# -
def print_random_car_notcar(cars, notcars):
# Just for fun choose random car / not-car indices and plot example images
car_ind = np.random.randint(0, len(cars))
notcar_ind = np.random.randint(0, len(notcars))
# Read in car / not-car images
car_image = mpimg.imread(cars[car_ind])
notcar_image = mpimg.imread(notcars[notcar_ind])
# Plot the examples
fig = plt.figure()
plt.subplot(121)
plt.imshow(car_image)
plt.title('Example Car Image')
plt.subplot(122)
plt.imshow(notcar_image)
plt.title('Example Not-car Image')
print_random_car_notcar(cars, notcars)
# +
from skimage.feature import hog
def get_hog_features(img, orient, pix_per_cell, cell_per_block, vis=True,
feature_vec=True):
"""
Function accepts params and returns HOG features (optionally flattened) and an optional matrix for
visualization. Features will always be the first return (flattened if feature_vector= True).
A visualization matrix will be the second return if visualize = True.
"""
return_list = hog(img, orientations=orient, pixels_per_cell=(pix_per_cell, pix_per_cell),
cells_per_block=(cell_per_block, cell_per_block),
block_norm= 'L2-Hys', transform_sqrt=False,
visualise= vis, feature_vector= feature_vec)
if vis:
hog_features = return_list[0]
hog_image = return_list[1]
return hog_features, hog_image
else:
hog_features = return_list
return hog_features
# +
def random_hog(car_images):
# Generate a random index to look at a car image
ind = np.random.randint(0, len(car_images))
# Read in the image
image = mpimg.imread(car_images[ind])
#RGB to HLS
YCrCb_image = cv2.cvtColor(image, cv2.COLOR_RGB2YCrCb)
Y_img = YCrCb_image[:,:,0]
Cr_img = YCrCb_image[:,:,1]
Cb_img = YCrCb_image[:,:,2]
hls_image = cv2.cvtColor(image, cv2.COLOR_RGB2HLS)
S_img = hls_image[:,:,2]
gray = cv2.cvtColor(image, cv2.COLOR_RGB2GRAY)
# Call our function with vis=True to see an image output
features, hog_image = get_hog_features(gray, orient= 5,
pix_per_cell= 8, cell_per_block= 2,
vis=True, feature_vec=False)
features1, hog_image1 = get_hog_features(gray, orient= 9,
pix_per_cell= 8, cell_per_block= 2,
vis=True, feature_vec=False)
features2, hog_image2 = get_hog_features(gray, orient= 14,
pix_per_cell= 8, cell_per_block= 3,
vis=True, feature_vec=False)
#================= next 4 images =================================#
features3, hog_image3 = get_hog_features(Y_img, orient= 9,
pix_per_cell= 8, cell_per_block= 2,
vis=True, feature_vec=False)
features4, hog_image4 = get_hog_features(S_img, orient= 9,
pix_per_cell= 8, cell_per_block= 5,
vis=True, feature_vec=False)
features5, hog_image5 = get_hog_features(Cb_img, orient= 9,
pix_per_cell= 8, cell_per_block= 2,
vis=True, feature_vec=False)
features6, hog_image6 = get_hog_features(S_img, orient= 9,
pix_per_cell= 8, cell_per_block= 4,
vis=True, feature_vec=False)
f, (ax1, ax2, ax3, ax4) = plt.subplots(1, 4, figsize=(30, 30))
f.tight_layout()
ax1.imshow(gray,cmap='gray')
ax1.set_title('Example Car Image', fontsize=35)
ax2.imshow(hog_image,cmap='gray')
ax2.set_title('HOG Visualization 1', fontsize=35)
ax3.imshow(hog_image1,cmap='gray')
ax3.set_title('HOG Visualization 2', fontsize=35)
ax4.imshow(hog_image2,cmap='gray')
ax4.set_title('HOG Visualization 3', fontsize=35)
f, (ax5, ax6, ax7, ax8) = plt.subplots(1, 4, figsize=(30, 30))
f.tight_layout()
ax5.imshow(hog_image3,cmap='gray')
ax5.set_title('HOG Visualization 5', fontsize=35)
ax6.imshow(hog_image4,cmap='gray')
ax6.set_title('HOG Visualization 6', fontsize=35)
ax7.imshow(hog_image5,cmap='gray')
ax7.set_title('HOG Visualization 7', fontsize=35)
ax8.imshow(hog_image6,cmap='gray')
ax8.set_title('HOG Visualization 8', fontsize=35)
# f, (ax9, ax10, ax11, ax12) = plt.subplots(1, 4, figsize=(30, 30))
# f.tight_layout()
# ax9.imshow(Y_img,cmap='gray')
# ax9.set_title('HOG Visualization 5', fontsize=35)
# ax10.imshow(Cr_img,cmap='gray')
# ax10.set_title('HOG Visualization 6', fontsize=35)
# ax11.imshow(Cb_img,cmap='gray')
# ax11.set_title('HOG Visualization 7', fontsize=35)
# ax12.imshow(S_img,cmap='gray')
# ax12.set_title('HOG Visualization 8', fontsize=35)
# plt.subplots_adjust(left=0., right=1, top=0.9, bottom=0.)
# -
random_hog(cars)
# +
def hog_feature_extract(cars, notcars):
# Generate a random index to look at a car image
features_car = []
features_notcar = []
hog_images_car = []
hog_images_notcar = []
count =0
for images in tqdm(cars):
# Read in the image
image = mpimg.imread(images)
gray = cv2.cvtColor(image, cv2.COLOR_RGB2GRAY)
features = get_hog_features(gray, orient= 9,
pix_per_cell= 8, cell_per_block= 2,
vis=False, feature_vec=True)
features_car.append(features)
#hog_images_car.append(hog_image)
count = count+1
print(count)
for images in tqdm(notcars):
# Read in the image
image = mpimg.imread(images)
gray = cv2.cvtColor(image, cv2.COLOR_RGB2GRAY)
features = get_hog_features(gray, orient= 9,
pix_per_cell= 8, cell_per_block= 2,
vis=False, feature_vec=True)
features_notcar.append(features)
#hog_images_notcar.append(hog_image)
#return features_car, hog_images_car, features_notcar, hog_images_notcar
return features_car, features_notcar
def extract_features(imgs, cspace='RGB', orient=9,
pix_per_cell=8, cell_per_block=2, hog_channel=0):
# Create a list to append feature vectors to
features = []
# Iterate through the list of images
for file in imgs:
# Read in each one by one
image = mpimg.imread(file)
# apply color conversion if other than 'RGB'
if cspace != 'RGB':
if cspace == 'HSV':
feature_image = cv2.cvtColor(image, cv2.COLOR_RGB2HSV)
elif cspace == 'LUV':
feature_image = cv2.cvtColor(image, cv2.COLOR_RGB2LUV)
elif cspace == 'HLS':
feature_image = cv2.cvtColor(image, cv2.COLOR_RGB2HLS)
elif cspace == 'YUV':
feature_image = cv2.cvtColor(image, cv2.COLOR_RGB2YUV)
elif cspace == 'YCrCb':
feature_image = cv2.cvtColor(image, cv2.COLOR_RGB2YCrCb)
else: feature_image = np.copy(image)
# Call get_hog_features() with vis=False, feature_vec=True
if hog_channel == 'ALL':
hog_features = []
for channel in range(feature_image.shape[2]):
hog_features.append(get_hog_features(feature_image[:,:,channel],
orient, pix_per_cell, cell_per_block,
vis=False, feature_vec=True))
hog_features = np.ravel(hog_features)
else:
hog_features = get_hog_features(feature_image[:,:,hog_channel], orient,
pix_per_cell, cell_per_block, vis=False, feature_vec=True)
# Append the new feature vector to the features list
features.append(hog_features)
# Return list of feature vectors
return features
print('Done')
# -
# +
print("cars :", len(cars))
print("notcars :", len(notcars))
#features_car, hog_images_car, features_notcar, hog_images_notcar = hog_feature_extract(cars,notcars)
#features_car, features_notcar = hog_feature_extract(cars,notcars)
#features_car, features_notcar = hog_feature_extract(cars,notcars)
colorspace = 'YCrCb' # Can be RGB, HSV, LUV, HLS, YUV, YCrCb
orient = 9
pix_per_cell = 8
cell_per_block = 2
hog_channel = 'ALL'
features_car = extract_features(cars, cspace=colorspace, orient=orient,
pix_per_cell=pix_per_cell, cell_per_block=cell_per_block,
hog_channel=hog_channel)
features_notcar = extract_features(notcars, cspace=colorspace, orient=orient,
pix_per_cell=pix_per_cell, cell_per_block=cell_per_block,
hog_channel=hog_channel)
# -
print("hog_images_car :", len(features_car))
print("hog_images_notcar :", len(features_notcar))
print(features_car[0].shape)
print(features_notcar[0].shape)
# +
def combine_dataset(features_car, features_notcar):
# dataset = np.copy(features_car)
# dataset = np.append(dataset,features_notcar, axis = 0)
# dataset_labels = np.ones_like(features_car)
# dataset_labels = np.append(dataset_labels,np.zeros_like(features_notcar))
dataset = np.vstack((features_car, features_notcar)).astype(np.float64)
dataset_labels = np.hstack(( np.ones(len(features_car)), np.zeros(len(features_notcar))))
print( len(dataset))
return dataset, dataset_labels
dataset, dataset_labels = combine_dataset(features_car, features_notcar)
# -
print(len(dataset), " ", len(dataset_labels))
print(len(features_car), " ", len(features_notcar))
print(np.ones_like(1196))
# +
from sklearn.model_selection import train_test_split
X_train , X_test, y_train, y_test = train_test_split(dataset, dataset_labels, test_size = 0.2)
# -
print ("cars_X_train : ", len(X_train))
print ("cars_X_test : ", len(X_test))
from sklearn.preprocessing import StandardScaler
X_scaler = StandardScaler().fit(X_train)
# Apply the scaler to X
X_train = X_scaler.transform(X_train)
X_test = X_scaler.transform(X_test)
# ## Train the svm
# +
from sklearn.svm import SVC
def train_svm(X_train , X_test, y_train, y_test):
clf = SVC(kernel = 'linear' , C = 1)
clf.fit(X_train, y_train)
acc = clf.score(X_test, y_test)
print(acc)
return clf
svc = train_svm(X_train , X_test, y_train, y_test)
# +
def convert_color(img, conv='RGB2YCrCb'):
if conv == 'RGB2YCrCb':
return cv2.cvtColor(img, cv2.COLOR_RGB2YCrCb)
if conv == 'BGR2YCrCb':
return cv2.cvtColor(img, cv2.COLOR_BGR2YCrCb)
if conv == 'RGB2LUV':
return cv2.cvtColor(img, cv2.COLOR_RGB2LUV)
def get_hog_features(img, orient, pix_per_cell, cell_per_block,
vis=False, feature_vec=True):
# Call with two outputs if vis==True
if vis == True:
features, hog_image = hog(img, orientations=orient,
pixels_per_cell=(pix_per_cell, pix_per_cell),
cells_per_block=(cell_per_block, cell_per_block),
block_norm= 'L2-Hys',
transform_sqrt=False,
visualise=vis, feature_vector=feature_vec)
return features, hog_image
# Otherwise call with one output
else:
features = hog(img, orientations=orient,
pixels_per_cell=(pix_per_cell, pix_per_cell),
cells_per_block=(cell_per_block, cell_per_block),
block_norm= 'L2-Hys',
transform_sqrt=False,
visualise=vis, feature_vector=feature_vec)
return features
def bin_spatial(img, size=(32, 32)):
color1 = cv2.resize(img[:,:,0], size).ravel()
color2 = cv2.resize(img[:,:,1], size).ravel()
color3 = cv2.resize(img[:,:,2], size).ravel()
return np.hstack((color1, color2, color3))
def color_hist(img, nbins=32): #bins_range=(0, 256)
# Compute the histogram of the color channels separately
channel1_hist = np.histogram(img[:,:,0], bins=nbins)
channel2_hist = np.histogram(img[:,:,1], bins=nbins)
channel3_hist = np.histogram(img[:,:,2], bins=nbins)
# Concatenate the histograms into a single feature vector
hist_features = np.concatenate((channel1_hist[0], channel2_hist[0], channel3_hist[0]))
# Return the individual histograms, bin_centers and feature vector
return hist_features
# -
def find_cars(img, ystarts, ystops, scales, svc, X_scaler, orient, pix_per_cell, cell_per_block, spatial_size, hist_bins):
draw_img = np.copy(img)
img = img.astype(np.float32)/255
bboxes = []
for scale, ystart, ystop in tqdm(zip(scales, ystarts, ystops)) :
img_tosearch = img[ystart:ystop,:,:]
ctrans_tosearch = convert_color(img_tosearch, conv='RGB2YCrCb')
if scale != 1:
imshape = ctrans_tosearch.shape
ctrans_tosearch = cv2.resize(ctrans_tosearch, (np.int(imshape[1]/scale), np.int(imshape[0]/scale)))
ch1 = ctrans_tosearch[:,:,0]
ch2 = ctrans_tosearch[:,:,1]
ch3 = ctrans_tosearch[:,:,2]
# Define blocks and steps as above
nxblocks = (ch1.shape[1] // pix_per_cell) - cell_per_block + 1
nyblocks = (ch1.shape[0] // pix_per_cell) - cell_per_block + 1
nfeat_per_block = orient*cell_per_block**2
# 64 was the orginal sampling rate, with 8 cells and 8 pix per cell
window = 64
nblocks_per_window = (window // pix_per_cell) - cell_per_block + 1
cells_per_step = 2 # Instead of overlap, define how many cells to step
nxsteps = (nxblocks - nblocks_per_window) // cells_per_step + 1
nysteps = (nyblocks - nblocks_per_window) // cells_per_step + 1
# Compute individual channel HOG features for the entire image
hog1 = get_hog_features(ch1, orient, pix_per_cell, cell_per_block, feature_vec=False)
hog2 = get_hog_features(ch2, orient, pix_per_cell, cell_per_block, feature_vec=False)
hog3 = get_hog_features(ch3, orient, pix_per_cell, cell_per_block, feature_vec=False)
for xb in range(nxsteps):
for yb in range(nysteps):
ypos = yb*cells_per_step
xpos = xb*cells_per_step
# Extract HOG for this patch
hog_feat1 = hog1[ypos:ypos+nblocks_per_window, xpos:xpos+nblocks_per_window].ravel()
hog_feat2 = hog2[ypos:ypos+nblocks_per_window, xpos:xpos+nblocks_per_window].ravel()
hog_feat3 = hog3[ypos:ypos+nblocks_per_window, xpos:xpos+nblocks_per_window].ravel()
hog_features = np.hstack((hog_feat1, hog_feat2, hog_feat3))
xleft = xpos*pix_per_cell
ytop = ypos*pix_per_cell
# Extract the image patch
subimg = cv2.resize(ctrans_tosearch[ytop:ytop+window, xleft:xleft+window], (64,64))
# Get color features
#spatial_features = bin_spatial(subimg, size=spatial_size)
#hist_features = color_hist(subimg, nbins=hist_bins)
# Scale features and make a prediction
#test_features = X_scaler.transform(np.hstack((spatial_features, hist_features, hog_features)).reshape(1, -1))
test_features = X_scaler.transform(np.array(hog_features).reshape(1, -1))
#test_features = X_scaler.transform(np.hstack((shape_feat, hist_feat)).reshape(1, -1))
test_prediction = svc.predict(test_features)
if test_prediction == 1:
xbox_left = np.int(xleft*scale)
ytop_draw = np.int(ytop*scale)
win_draw = np.int(window*scale)
#cv2.rectangle(draw_img,(xbox_left, ytop_draw+ystart),(xbox_left+win_draw,ytop_draw+win_draw+ystart),(0,0,255),6)
return draw_img
# +
img = mpimg.imread('test_images/test1.jpg')
ystart = [400,400,400,464]
ystop = [496,560,592,656]
#ystart = 400
#ystop = 656
scale = [1,2,3,4] #4 #1.5
color_space = 'YCrCb' # Can be RGB, HSV, LUV, HLS, YUV, YCrCb
orient = 9 # HOG orientations
pix_per_cell = 8 # HOG pixels per cell
cell_per_block = 2 # HOG cells per block
hog_channel = 'ALL' # Can be 0, 1, 2, or "ALL"
spatial_size = (16, 16) # Spatial binning dimensions
hist_bins = 16 # Number of histogram bins
spatial_feat = False # Spatial features on or off
hist_feat = False # Histogram features on or off
hog_feat = True # HOG features on or off
images = glob.glob('test_images/test*.jpg')
for image in tqdm(images) :
img = mpimg.imread(image)
out_img = find_cars(img, ystart, ystop, scale, svc, X_scaler, orient, pix_per_cell, cell_per_block, spatial_size, hist_bins)
plt.figure()
plt.imshow(out_img)
# -
# Here is your draw_boxes function from the previous exercise
def draw_boxes(img, bboxes, color=(0, 0, 255), thick=6):
# Make a copy of the image
imcopy = np.copy(img)
# Iterate through the bounding boxes
for bbox in bboxes:
# Draw a rectangle given bbox coordinates
cv2.rectangle(imcopy, bbox[0], bbox[1], color, thick)
# Return the image copy with boxes drawn
return imcopy
# +
# Define a function that takes an image,
# start and stop positions in both x and y,
# window size (x and y dimensions),
# and overlap fraction (for both x and y)
def slide_window(img, x_start_stop=[None, None], y_start_stop=[None, None],
xy_window=(64, 64), xy_overlap=(0.5, 0.5)):
# If x and/or y start/stop positions not defined, set to image size
if x_start_stop[0] == None:
x_start_stop[0] = 0
if x_start_stop[1] == None:
x_start_stop[1] = img.shape[1]
if y_start_stop[0] == None:
y_start_stop[0] = 0
if y_start_stop[1] == None:
y_start_stop[1] = img.shape[0]
# Compute the span of the region to be searched
xspan = x_start_stop[1] - x_start_stop[0]
yspan = y_start_stop[1] - y_start_stop[0]
# Compute the number of pixels per step in x/y
nx_pix_per_step = np.int(xy_window[0]*(1 - xy_overlap[0]))
ny_pix_per_step = np.int(xy_window[1]*(1 - xy_overlap[1]))
# Compute the number of windows in x/y
nx_buffer = np.int(xy_window[0]*(xy_overlap[0]))
ny_buffer = np.int(xy_window[1]*(xy_overlap[1]))
nx_windows = np.int((xspan-nx_buffer)/nx_pix_per_step)
ny_windows = np.int((yspan-ny_buffer)/ny_pix_per_step)
# Initialize a list to append window positions to
window_list = []
# Loop through finding x and y window positions
# Note: you could vectorize this step, but in practice
# you'll be considering windows one by one with your
# classifier, so looping makes sense
for ys in range(ny_windows):
for xs in range(nx_windows):
# Calculate window position
startx = xs*nx_pix_per_step + x_start_stop[0]
endx = startx + xy_window[0]
starty = ys*ny_pix_per_step + y_start_stop[0]
endy = starty + xy_window[1]
# Append window position to list
window_list.append(((startx, starty), (endx, endy)))
# Return the list of windows
return window_list
# +
image = mpimg.imread('test_images/test4.jpg')
windows = slide_window(image, x_start_stop=[None, None], y_start_stop=[None, None],
xy_window=(128, 128), xy_overlap=(0.5, 0.5))
window_img = draw_boxes(image, windows, color=(0, 0, 255), thick=6)
plt.imshow(window_img)
# +
import matplotlib.image as mpimg
import matplotlib.pyplot as plt
import numpy as np
import cv2
import glob
import time
from sklearn.svm import LinearSVC
from sklearn.preprocessing import StandardScaler
from skimage.feature import hog
# NOTE: the next import is only valid for scikit-learn version <= 0.17
# for scikit-learn >= 0.18 use:
# from sklearn.model_selection import train_test_split
# -
def extract_features1(imgs, cspace='RGB', orient=9,
pix_per_cell=8, cell_per_block=2, hog_channel=0):
# Create a list to append feature vectors to
features = []
# Iterate through the list of images
for file in imgs:
# Read in each one by one
image = imgs
# apply color conversion if other than 'RGB'
if cspace != 'RGB':
if cspace == 'HSV':
feature_image = cv2.cvtColor(image, cv2.COLOR_RGB2HSV)
elif cspace == 'LUV':
feature_image = cv2.cvtColor(image, cv2.COLOR_RGB2LUV)
elif cspace == 'HLS':
feature_image = cv2.cvtColor(image, cv2.COLOR_RGB2HLS)
elif cspace == 'YUV':
feature_image = cv2.cvtColor(image, cv2.COLOR_RGB2YUV)
elif cspace == 'YCrCb':
feature_image = cv2.cvtColor(image, cv2.COLOR_RGB2YCrCb)
else: feature_image = np.copy(image)
# Call get_hog_features() with vis=False, feature_vec=True
if hog_channel == 'ALL':
hog_features = []
for channel in range(feature_image.shape[2]):
hog_features.append(get_hog_features(feature_image[:,:,channel],
orient, pix_per_cell, cell_per_block,
vis=False, feature_vec=True))
hog_features = np.ravel(hog_features)
else:
hog_features = get_hog_features(feature_image[:,:,hog_channel], orient,
pix_per_cell, cell_per_block, vis=False, feature_vec=True)
# Append the new feature vector to the features list
features.append(hog_features)
# Return list of feature vectors
return features
# Define a function you will pass an image
# and the list of windows to be searched (output of slide_windows())
def search_windows(img, windows, clf, scaler, color_space='RGB',
spatial_size=(32, 32), hist_bins=32,
hist_range=(0, 256), orient=9,
pix_per_cell=8, cell_per_block=2,
hog_channel=0, spatial_feat=True,
hist_feat=True, hog_feat=True):
#1) Create an empty list to receive positive detection windows
on_windows = []
#2) Iterate over all windows in the list
for window in windows:
#3) Extract the test window from original image
test_img = cv2.resize(img[window[0][1]:window[1][1], window[0][0]:window[1][0]], (64, 64))
#4) Extract features for that window using single_img_features()
features = extract_features1(test_img, cspace=colorspace, orient=orient,
pix_per_cell=pix_per_cell, cell_per_block=cell_per_block,
hog_channel=hog_channel)
#5) Scale extracted features to be fed to classifier
test_features = scaler.transform(np.array(features).reshape(1, -1))
#6) Predict using your classifier
prediction = clf.predict(test_features)
#7) If positive (prediction == 1) then save the window
if prediction == 1:
on_windows.append(window)
#8) Return windows for positive detections
return on_windows
# +
sample_size = 500
cars = cars[0:sample_size]
notcars = notcars[0:sample_size]
### TODO: Tweak these parameters and see how the results change.
color_space = 'RGB' # Can be RGB, HSV, LUV, HLS, YUV, YCrCb
orient = 9 # HOG orientations
pix_per_cell = 16 # HOG pixels per cell
cell_per_block = 2 # HOG cells per block
hog_channel = 'ALL' # Can be 0, 1, 2, or "ALL"
y_start_stop = [400, 656] # Min and max in y to search in slide_window()
# Fit a per-column scaler
X_scaler = StandardScaler().fit(X_train)
# Apply the scaler to X
X_train = X_scaler.transform(X_train)
X_test = X_scaler.transform(X_test)
print('Using:',orient,'orientations',pix_per_cell,
'pixels per cell and', cell_per_block,'cells per block')
print('Feature vector length:', len(X_train[0]))
# Use a linear SVC
svc = LinearSVC()
# Check the training time for the SVC
t=time.time()
svc.fit(X_train, y_train)
t2 = time.time()
print(round(t2-t, 2), 'Seconds to train SVC...')
# Check the score of the SVC
print('Test Accuracy of SVC = ', round(svc.score(X_test, y_test), 4))
# Check the prediction time for a single sample
t=time.time()
image = mpimg.imread('test_images/test4.jpg')
draw_image = np.copy(image)
# Uncomment the following line if you extracted training
# data from .png images (scaled 0 to 1 by mpimg) and the
# image you are searching is a .jpg (scaled 0 to 255)
#image = image.astype(np.float32)/255
windows = slide_window(image, x_start_stop=[None, None], y_start_stop=y_start_stop,
xy_window=(96, 96), xy_overlap=(0.5, 0.5))
hot_windows = search_windows(image, windows, svc, X_scaler, color_space=color_space,
spatial_size=spatial_size, hist_bins=hist_bins,
orient=orient, pix_per_cell=pix_per_cell,
cell_per_block=cell_per_block,
hog_channel=hog_channel, spatial_feat=spatial_feat,
hist_feat=False, hog_feat=hog_feat)
window_img = draw_boxes(draw_image, hot_windows, color=(0, 0, 255), thick=6)
plt.imshow(window_img)
# -
# +
a = [np.array([[1,1],[1,1]]), np.array([[2,2],[2,2]])]
b = [np.array([[3,3],[3,3]]), np.array([[4,4],[4,4]])]
c = np.hstack((a, b))
print(a)
print()
print(b)
print()
print(c)
print()
print( np.vstack((a, b)))
print()
print( np.dstack((a, b)))
x = [1,23,2,4,5,6]
y = [1,23,1,1,1,1]
print(np.hstack((x,y)))
| Notebook1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# LIST BASICS
# ===========
#
#
# QUESTION 1
# ----------
#
#
# - make replicate_me 10 times longer i.e. [1] >> [1, 1, 1, 1, 1, 1, 1, 1, 1, 1]
#
# ```python
# replicate_me = [1]
#
# - combine the newly extended replicate_me with replicate_me
#
# ```python
# combine_me = [2,2,2,2,2,2]
# ```
# - replace the 4th number in the list with the variable substitute_me
#
# - do it again with a different method
#
# ```python
# substitute_me = ['surprise!']
# ```
#
# - replace every other element in the list with the number 5
# - HINT: you can do this in a single command
#
# - delete elements 5-8 from the list
# - add the numbers 8, 13 and the string 'hello' to the end of your list
#
#
#
#
# - do it again with a different method
# - reverse the order of the list
# - in one statement, return the 5th element of your list and remove it from the list
# - assign your list to a new variable called 'copy_of_list'
# - change all the elements in this new list to zeros
# - print out the original list (not the copy)
# - what happens? did you expect that? find a way to copy the list properly
# - think about or read about why this is the case in python
# +
replicate_me = [1]
x = 10
replicate_me = [i for i in replicate_me for _ in range(x)] #this is a list comprehension
print(replicate_me)
#TAS SOLUTION:
replicate_me2 = [1]
replicate_me2 = replicate_me2 * 10
print(replicate_me2)
#OR
replicate_me3 = [1]
replicate_me3 *= 10 #it's a bit like the += sign=, it will increment by whatever is behind
print(replicate_me3)
combine_me = [2,2,2,2,2,2]
combine_me += replicate_me
print (combine_me)
substitute_me = ['surprise!']
combine_me = [2, 2, 2, 2, 2, 2] + replicate_me
combine_me[3] = substitute_me # note 0-indexing
print(combine_me)
for i in range(len(combine_me)): #len(combine_me) will give you/calculate the length of that particluar string
if i == 3:
i = substitute_me
print (combine_me)
combine_me[1::2] = [5] * int(len(combine_me) / 2) #int = integer. This way it means Integer(length(list)/2)
#by adding len(something) this will return the length of the list/tuple/dictionary , i.e 45 if yiu have 45 characters so it's an easy way to find out how many items you have
print(combine_me)
del combine_me[4:7]
print(combine_me)
combine_me.extend([8, 13, 'hello'])
print(combine_me)
#OR
for element in (8, 13, 'hello'):
combine_me.append(element) #APPEND is adding a list unless you define that you want to add an element. Good for adding lists in lists
print(combine_me)
# Reverse the order with step=-1
combine_me = [2, 2, 2, 2, 2, 2] + replicate_me
combine_me = combine_me[::-1] #Adding :-1 to the list will reverse the order. The previous : is equivalent to start:end
print(combine_me)
# return (and remove) 5th item
print(combine_me.pop(4)) #pop will return the letter and remove it from the list
print(combine_me)
# +
# assign your list to a new variable called 'copy_of_list'
# change all the elements in this new list to zeros
# print out the original list (not the copy)
# what happens? did you expect that? find a way to copy the list properly
# think about or read about why this is the case in python
"""assign your list to a new variable called 'copy_of_list'"""
"""
WARNING: something of the form:
copy_of_list = combined
is incorrect because it only returns a new reference to the same list
and does not copy it. Any changes to the first will also affect the second
"""
# To actually do a copy:
from copy import copy # this would normally be at the top of your file! SOS: https://github.com/python/cpython/blob/3.6/Lib/copy.py
copy_of_list = copy(combine_me) #SOS: YOU NEED TO IMPORT THE COPY MODULE AND FROM THATONLY THE COPY FUNCTION AS THE DEEPCOPY IS SOMETHING DIFFERENT
# change all the elements in this new list to zeros
copy_of_list[:] = [0] * (len(copy_of_list))
# For numpy which we will see soon ( I have seen answers using it),
# it would be np.zeros(len(copy_of_list))
# Print out the original list (not the copy)
print(combine_me)
print(copy_of_list)
# What happens? did you expect that? find a way to copy the list properly
""" If you did not do a real copy, combined has been updated with zeros """
# Think about or read about why this is the case in python
""" The assignment just copies the reference to the list,
not the actual list, so both new_list and my_list refer to the same
list after the assignment."""
"""Is it safe to write from copy import copy in Python? – <NAME> Aug 21 '12 at 13:42
2
It is perfectly safe.
The copy module contains two functions (copy and deepcopy), and one exception (error).
While it is perfectly fine (even from a pep8 perspective) to import them into your namespace either singularly:
from copy import copy or more than one: from copy import copy, deepcopy.
With generic names (like copy) it can quickly become confusing to read your code and understand
where a function is coming from, it is therefore often good practice to import the module name only (
and not the functions etc. inside): from xml.dom import minidom then using minidom.parse(..)
– thebjorn Aug 21 '12 at 18:28"""
# -
# NESTED LISTS
# ============
#
# QUESTION 2
# ----------
#
# You have some experimental data (trials_1 and trials_2) which contain a sub-list for each experimental trial. Each number is the number of spikes for a single cell, an empty sublist means no spikes were detected during the trial
#
#
# ```python
#
# experiment_1_trials = [[1, 2, 3, 6],
# [],
# [],
# [4, 5, 6, 7, 8, 3, 12, 2],
# [2],
# [9, 10, 11, 63],
# [123, 2, 2],
# [199, 2, 24, 5, 66],
# [],
# [2, 3, 5, 6],
# [4]
# ]
#
# experiment_2_trials = [[12, 43, 2, 2],
# [2],
# [],
# [],
# [1, 2, 3, 5]
# ]
# ```
#
# - make one list called all_trials that contains all of experiment_1_trials and experiment_2_trials
# - see if you can do this in three different ways
# - calculate the sum of the 2nd and 3rd cells of the 1st and 4th trials
# - if you did this with a loop, do it again by indexing
# - print the number of cells firing in each trial
# - how many of the trials contain the number 2?
#
#
# Because you, the experimenter, used an outdated form of note taking, consisting of writing things down on tissues instead of a lab book or database you forgot that you actually did a trial in between experiment_1_trials and experiment_2_trials:
#
# ```python
# new_trial = [1, 3, 5, None, 5, 6, float('NaN')]
# ```
#
# - clean the new_trial to remove any non-numeric elements
# - insert new_trial into the main list in between experiment_1_trials and experiment_2_trials
# - ideally do this on the list you have already, don't rebuild the list from the original components
# - something went wrong on your last trial, but you want to keep the data somewhere in case you need it again
# - obtain and remove it from the list
# - do the same for the 4th trial
# - do so using a built-in list method
#
# - find out the number of trials where no cells fired
# - remove any empty trials from your list
# - if you do this with a simple for loop, do you get the result you expect? if not, why not?
# - if you didn't already, try these again using built-in list methods (where applicable)
#
# - remove all trials that contain less than 3 elements
# - create a list containing the number of spikes of the second cell of each of the remaining trials
#
# - flatten the list (i.e. remove the nesting, but keep the values) to make one list of all spike counts
# - put the numbers in descending order - with and without changing the original list
# +
#You have some experimental data (trials_1 and trials_2) which contain a sub-list for each experimental trial.
#Each number is the number of spikes for a single cell, an empty sublist means no spikes were detected during the trial
experiment_1_trials = [[1, 2, 3, 6],
[],
[],
[4, 5, 6, 7, 8, 3, 12, 2],
[2],
[9, 10, 11, 63],
[123, 2, 2],
[199, 2, 24, 5, 66],
[],
[2, 3, 5, 6],
[4]
]
experiment_2_trials = [[12, 43, 2, 2],
[2],
[],
[],
[1, 2, 3, 5]
]
# Make one list called all_trials that contains all of experiment_1_trials and experiment_2_trials
# from copy import copy (SOS: this would have been on top of the list if it wasn't imported in the previous cell)
all_trials = experiment_1_trials + experiment_2_trials
print("All trials: \n{}".format(all_trials))
# See if you can do this in three different ways
all_trials = copy(experiment_1_trials)
all_trials.extend(experiment_2_trials)
print("All trials: \n{}".format(all_trials))
""" or """
all_trials = copy(experiment_1_trials) #make sure you have imported copy (see 'from copy import copy' above)
for trial in experiment_2_trials:
all_trials.append(trial)
print("All trials: \n{}".format(all_trials))
"""" Do not do the following:this is not correct, see comment"""
all_trials = copy(experiment_1_trials)
all_trials.append(experiment_2_trials)
print("All trials: \n{}".format(all_trials)) #if you don't add each element, with append you will just add the list at the end creating a list of lists
# append can only add 1 element!!
""" Do not do the following:
all_trials = [experiment_1_trials, experiment_2_trials]
This is matlab syntax and in python will give you a list of lists
"""
all_trials = [experiment_1_trials, experiment_2_trials]
print("All trials: \n{}".format(all_trials))
# -
all_trials2 = experiment_1_trials[:] #instead of copy u can use this [:]
all_trials2.extend(experiment_2_trials)
print("All trials2: \n{}".format(all_trials))
all_trials2[2] = 2
print(all_trials2)
print (experiment_1_trials)
# +
all_trials = experiment_1_trials + experiment_2_trials
# Calculate the sum of the 2nd and 3rd cells of the 1st and 4th trials
print("Cell sums: \n{}".format(all_trials[0][1:3] + all_trials[3][1:3]))
# Print the number of cells firing in each trial
print("Firing cells per trial")
for i, trial in enumerate(all_trials): #this will give them numbers
print("Trial {}: {}".format(i, len(trial)))
# How many of the trials contain the number 2?
print("Trials with number 2")
n_trials_with_2 = 0
for trial in all_trials:
if 2 in trial:
n_trials_with_2 += 1
print(n_trials_with_2)
"""One liner"""
print(len([t for t in all_trials if 2 in t])) # Use of single letter var is OK only for listcomprehension
""" or """
print(sum([2 in t for t in all_trials]))
# +
n1 = 4
n2 = 5
s3 = "2**10 = {0} and {1} * {2} = {3:f}".format(2**10, n1, n2, n1 * n2)
print(s3)
# -
# CASTING AND LIST COMPREHENSIONS
# ===============================
#
# QUESTION 3
# ----------
#
# - use `range()` to create a list of numbers 1-100
# - check the type of your result
# - make a list of all numbers up to 100 squared
# - make a list of all the numbers in the list of squared numbers that are multiples of 9
# - reverse the order of each sublist in the list_of_x_y_coordinates (i.e. [1,2], [3,5] >> [2,1], [5,3])
#
# ```python
# list_of_x_y_coordinates = [[1, 2], [3, 5], [5, 6], [10, 20]]
# +
# Use 'range()' to create a list of numbers 1-100
numbers_to_100 = range(1,101)
print(type(numbers_to_100))
"""It is not a list but an iterator
You can cast it to a list however"""
numbers_to_100 = list(range(1,101))
print(type(numbers_to_100))
# make a list of all numbers up to 100 squared
numbers_to_100_squared = [x*x for x in numbers_to_100]
print(numbers_to_100_squared)
# make a list of all the numbers in the list of squared numbers that are multiples of 9
multiples_of_nine = [x*x for x in numbers_to_100 if x*x % 9 is 0]
print(multiples_of_nine)
#or
multiples_of_9 = [x**2 for x in numbers_to_100 if (x % 9) == 0]
print(multiples_of_9)
# reverse the order of each sublist in the list_of_x_y_coordinates (i.e. [1,2], [3,5] >> [2,1], [5,3])
list_of_x_y_coordinates = [[1, 2], [3, 5], [5, 6], [10, 20]]
rev_list_of_x_y_coordinates = [coords[::-1] for coords in list_of_x_y_coordinates] #: means all list and :-1 means to reverse it
print(rev_list_of_x_y_coordinates)
# -
# ```
#
# TUPLE BASICS
# ============
#
# QUESTION 4
# ----------
#
# ```python
# x = 10
# y = 10
# z = 5
# coordinates = x, y, z
# ```
#
# - check if the number 5 is in the coordinates tuple
# - try to change the y coordinate to 15
# - note that you can't do this - tuples are immutable
# - think about why/when that might be useful
# - think about when it might not be useful
# - if you first cast the tuple into a different type (one that you know is mutable) then you can change the coordinate - try this now
#
# - in this example, you only care about x and y together, but not z
# - try to use something called tuple unpacking to separate coordinates into the variables loc_2d and z
#
# ```python
# example_tuple = 'this', 'is', 'quite', 'stringy', {'dictionary': 100, 'stuff': 340, 'here': 23424}
# ```
#
# - use the same principle to put all the first 4 string variables into a single variable, and the dictionary in its own variable such that:
# ```python
#
# variable_1 = 'this', 'is', 'quite', 'stringy'
# variable_2 = {'dictionary': 100, 'stuff': 340, 'here': 23424}
# +
x = 10
y = 10
z = 5
coordinates = x, y, z
if 5 in coordinates:
print('ok')
#or
print(5 in coordinates)
#Try to change the y coordinate to 15
"""coordinates [1] = 15
THEN YOU RECEIVE THE FOLLOWING ERROR:
---------------------------------------------------------------------------
TypeError Traceback (most recent call last)
<ipython-input-18-14b3122ac4b8> in <module>()
9 #Try to change the y coordinate to 15
10
---> 11 coordinates [1] = 15
12
13 #Note that you can't do this - tuples are immutable
TypeError: 'tuple' object does not support item assignment"""
#Note that you can't do this - tuples are immutable
#Think about why/when that might be useful
"""
Immutability is good to make sure that you do not accidentally (nobody is perfect) change
the values inside your list. This is especially important when you pass your list to a
function because since lists are passed by reference, you could end up accidentally modifying
the values in your list if the function you use has a bug in it. Therefore, it is best
practice to pass it as immutable to the function."""
coordslist = list(coordinates) # you cant change elements in tuples, need to turn them into lists first
coordslist[1] = 15
print(coordslist)
# -
# ```
#
#
# STRINGS LISTS TUPLES AND DICTIONARIES COMBINED
# ==============================================
#
# QUESTION 5
# ----------
#
#
# - read the stimulus_starts and stimulus_ends from the stimulus_parameters.txt file
# - put them into a dictionary such that anything to the left of '=' is a key, and anything to the right is a value
# - save your dictionary to a file for later on
# - load your dictionary from earlier
# - loop over and print the following:
# - all the keys in the dictionary
# - all values in the dictionary
# - both
#
# - get the stimulus starts and ends and put them into the following variables:
# - do this using the dictionary key
# - do this in a way that both returns the values, and also removes them from the dictionary (HINT: look at built-in dictionary methods)
# - make sure that each entry in the list is an integer type, not a string
#
# ```python
# stimulus_starts =
# stimulus_ends =
# ```
#
# - loop through the starts and ends simultaneously and print them out
#
# - if you didn't already, use the built in function zip
# - to understand how this works, cast your zipped lists into a list and print it to the console
# - what happens if the two lists are of different lengths?
# - use zip and print out the start and end of the third stimulus
#
# - HINT/BONUS (recommended): you could try using a package called configobj for a 1 line solution to the first 4 parts of this question
# DICTIONARY COMPREHENSION
# ========================
#
# QUESTION 6
# ----------
#
# Imagine a (quite stupid because the keys don't do much) hypothetical dictionary like the one below to store postsynaptic events that contains the name of the event as a key and the time start, time end and amplitude as a tuple.
#
# - Using dictionary comprehension, return a dictionary (of the same not very smart format) that contains only events with
# - an amplitude > 20
# - a duration of 1
# - a duration between 2 and 4
#
# ```python
# synaptic_events = {
# 'e1': (10, 12, 22),
# 'e2': (15, 17, 12),
# 'e3': (20, 21, 25),
# 'e4': (25, 27, 32),
# 'e5': (30, 32, 5),
# 'e6': (35, 38, 7),
# 'e7': (40, 45, 48),
# 'e8': (46, 47, 12),
# 'e9': (49, 50, 47),
# 'e10': (51, 53, 22)
# }
# ```
#
# +
horsemen = ["war", "famine", "pestilence", "death"]
for h in horsemen:
print(h)
for i in [0, 1, 2, 3]:
print(horsemen[i])
for i in range(len(horsemen)):
print(horsemen[i])
"""All of these give the same results, i.e. the elements of the list"""
xs = [1, 2, 3, 4, 5]
for i in range(len(xs)):
xs[i] = xs[i]**2
print(xs[i]) #this will print every element to the power of 2
print(xs) """this will print the list with the calculations for each individual number (as it's a for list it )
it will update each and every one of them in the power of 2 every time it executes the for command
"""
| .ipynb_checkpoints/lists_tuples_and_dictionaries-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: ir
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# + [markdown] slideshow={"slide_type": "slide"}
#
# # Econ5121 B&C (Fall 2018)
#
# ### <NAME>
#
# * Time Series
# * Panel data
# * Generalized method of moments
# + [markdown] slideshow={"slide_type": "slide"}
# ### Useful R Packages
#
# * `quantmod`: financial and US macro data
# * `Quandl`: many data resources
#
# * `dynlm`: single-equation dynamic model
# * `tsDyn`: multiple-equation dynamic models
# + slideshow={"slide_type": "subslide"}
library(quantmod, quietly = TRUE)
getSymbols("^HSI") # S&P 500
tail(HSI)
plot(HSI$HSI.Close, type = "l")
# + slideshow={"slide_type": "subslide"}
library(Quandl)
CNH=Quandl("UNAE/GDPCD_CHN") #https://www.quandl.com/data/UNAE/GDPCD_CHN-GDP-Current-Prices-US-Dollars-China
HKG=Quandl("UNAE/GDPCD_HKG") #https://www.quandl.com/data/UNAE/GDPCD_HKG-GDP-Current-Prices-US-Dollars-China-Hong-Kong-SAR
head(CNH)
# + [markdown] slideshow={"slide_type": "slide"}
# # Dynamic regression model
#
# $$
# y_t = \beta_1 + \beta_2 x_t + \beta_3 x_{t-1} + \gamma y_{t-1} + e_t
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# ## Motivations
#
# * temporal lags of effect
# * expectation formed from the past
# * explicitly depends on history
# + slideshow={"slide_type": "subslide"}
library(quantmod)
getSymbols.FRED(Symbols = "POILBREUSDQ", env = .GlobalEnv) # Brent Oil price
x = POILBREUSDQ; T = length(x)
getSymbols.FRED(Symbols = "IPB50001SQ", env = .GlobalEnv) # Instrial Index (quaterly data)
Ty = length(IPB50001SQ); y = IPB50001SQ[(Ty - T + 1):Ty]
x = ts(x); y = ts(y)
plot(cbind(y,x), main = "")
# + [markdown] slideshow={"slide_type": "subslide"}
# ### ARDL(1,1) regression example
# + slideshow={"slide_type": "-"}
library(dynlm)
reg = dynlm( y ~ L(y, c(1) ) + L(x,c(1:2) ) )
print(summary(reg))
# + [markdown] slideshow={"slide_type": "slide"}
# # Lagged Effect
#
# $$y_t = \alpha + \sum_{i=0}^{\infty} \beta_i x_{t-i} + e_t$$
#
# ## Interpretation as a generative model
#
# * Impact multiplier: $\beta_0$
# * Cumulated effect (of $\tau$ periods): $\sum_{i=0}^{\tau} \beta_i $
# * Equilibrium multiplier: $\sum_{i=0}^{\infty} \beta_i $
# + [markdown] slideshow={"slide_type": "slide"}
# # Lag Operator
#
# $$L x_t = x_{t-1}$$
# $$L^{\tau} x_t = x_{t-\tau}$$
#
# ## Difference operator
# $$\Delta x_t = x_t - x_{t-1} = (1-L) x_t$$
# + [markdown] slideshow={"slide_type": "slide"}
# # Stationary time series
#
# For a univariate time series $(y_t)_{t= - \infty} ^ {\infty}$,
# * **Strictly stationary**: joint distribution of any finite coordinate only depends on their relative position.
# * **Weakly stationary**: the first two moments of any pair $y_t$ and $y_s$ only depends on their relative position.
# * $E[y_t] = \mu$ for all $t$
# * $\mathrm{var}[y_t] = \sigma^2$ for all $t$
# * $\mathrm{cov}[y_t, y_{t+\tau} ] $ only depends on $\tau$ independent of $t$
#
# + [markdown] slideshow={"slide_type": "subslide"}
#
# This notion can be extended to multiple-variate time series, for example $(y_t, x_t, e_t)$.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Dynamic regression model
#
# $$y_t = \alpha + \sum_{i=0}^{\infty} \beta_i x_{t-i} + e_t = \alpha + B(L) x_t + e_t $$
# where $$B(L) = \sum_{i=0}^{\infty} \beta_i L^i$$ is a polynomial of the lag operators.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Autoregressive model
# $$y_t = \alpha + \sum_{i=1}^p \gamma_p y_{t-p} + e_t $$ can be written as
# $$ C(L) y_t = \alpha + e_t $$ where $$C(L) = 1 -\gamma_1 L - \cdots - \gamma_p L^p $$ is a polynomial of the lag operators.
# + [markdown] slideshow={"slide_type": "slide"}
# # Invertibility
#
# If the roots of the polynomial equation $C(z) = 0$ **all** lies **outside** of the unit circle, we say the autoregressive model is invertible.
# + [markdown] slideshow={"slide_type": "subslide"}
# More generally, in the polynomial equation $C(z) = 0$, the root with the smallest module determines the trend of the time series.
#
# If $e_t$ is stationary with finite variance and $\alpha=0$
# * If the module of the smallest root is bigger than 1, $y_t$ is a stationary time series
# * If the module of the smallest root is equal to 1, $y_t$ is a **unit root** process
# * If the module of the smallest root is smaller than 1, $y_t$ is an **explosive** process
# + [markdown] slideshow={"slide_type": "slide"}
#
# ### Numerical Example
#
# * $C(L) = 1 - 0.5L$ is invertible.
# * $C(L) = 1 - L$ is non-invertible.
# * $C(L) = 1 - 1.1L$ is non-invertible.
# + slideshow={"slide_type": "fragment"}
AR = function(b,T){
y = rep(0,T)
for (t in 1:T){
if (t > 1) {
y[t] = b * y[t - 1] + rnorm(1)
}
}
return(ts(y) )
}
# + slideshow={"slide_type": "subslide"}
T = 1000; plot( x = 1:T, y = AR(0.9, T), type = "l")
# + slideshow={"slide_type": "subslide"}
T = 200; plot( x = 1:T, y = AR(1.0, T), type = "l")
# + slideshow={"slide_type": "subslide"}
T = 200; plot( x = 1:T, y = AR(1.05,T), type = "l")
# + [markdown] slideshow={"slide_type": "slide"}
# # Autoregressive Distributed Lag Models
#
#
# ARDL(p,r) model:
# $$C(L) y_t = \mu + B(L) x_t + e_t $$
# where
# $$C(L) = 1 -\gamma_1 L - \cdots - \gamma_p L^p$$
# and
# $$B(L) = \beta_0 + \beta_1 L + \cdots + \beta_r L^r.$$
#
# **Granger causality**: $\beta_0 = \beta_1 = \cdots = \beta_r = 0$.
# + [markdown] slideshow={"slide_type": "slide"}
# # Error Correction Model
#
# <NAME> (Nobel prize 2001)
#
# Subtract $y_{t-1}$ from both sides of the ARDL(1,1) model
# $$
# \begin{align*}
# \Delta y_t & = \mu + \beta_0 x_t + \beta_1 x_{t-1} + (\gamma_1 -1 ) y_{t-1} + e_t \\
# & = \mu + \beta_0 \Delta x_t + (\beta_1 + \beta_0) x_{t-1} + (\gamma_1 -1 ) y_{t-1} + e_t \\
# & = \mu + \beta_0 \Delta x_t + (\gamma_1 -1 )( y_{t-1} - \theta x_{t-1} ) + e_t
# \end{align*}
# $$
# where $\theta = (\beta_1 + \beta_0)/(1 - \gamma_1)$.
#
# * An equilibrium relationship $\Delta y_t = \mu + \beta_0 \Delta x_t + e_t$.
# * An equilibrium error $(\gamma_1 - 1 ) (y_{t-1} - \theta x_{t-1} ) $.
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
# * Useful to identify spurious regression
# * First difference recovers stationarity
#
# * Can be estimated either by OLS or by NLS or by MLE
# + [markdown] slideshow={"slide_type": "slide"}
#
# # Spurious Regression
#
#
# * The two time series $\{y_t\}$ and $\{x_t\}$ are generated independently, so that $E[y_t|x_t] = 0$.
# * However, we observe a high $R^2$ and large t-value if we regression $y_t$ against $x_t$.
#
# + slideshow={"slide_type": "subslide"}
T = 50
a = 1
y <- AR(a, T)
x <- AR(a, T)
matplot( cbind(y, x), type = "l", ylab = "" )
# + slideshow={"slide_type": "subslide"}
reg <- lm(y ~ x)
summary(reg)
# + [markdown] slideshow={"slide_type": "slide"}
# <NAME> Newbold (1974)
#
# run a regression to check that if we naively use 1.96 as the critical value for the $t$-ratio, how often we would reject the null hypothesis that $\beta = 0$.
#
# * The nominal asymptotic test size is $5\%$ according to the standard asymptotic theory
# * The empirical size is about 0.80 in this simulation
# * The drastic deviation suggests that the standard asymptotic theory fails in the nonstationary environment.
#
# + slideshow={"slide_type": "subslide"}
spurious <- function(i, a, T){
y <- AR(a, T)
x <- AR(a, T)
reg <- lm(y ~ x)
p.val <- summary(reg)[[4]][2,4]
# save the p-value of the estimate of x's coefficient
return(p.val)
}
library("plyr")
out <- ldply(.data = 1:1000, .fun = spurious, a = 1, T = 100)
print( mean(out < 0.05) )
# + [markdown] slideshow={"slide_type": "slide"}
# # Model Specification
#
# Information criterion.
#
# Let $k$ be the total number of slope coefficient in the model.
#
# * Akaike information criterion: $\log( \hat{\sigma}^2 ) + 2\times (k / T )$.
# * Tend to overfit, but better for prediction
# * Bayesian information criterion: $\log( \hat{\sigma}^2 ) + \log(T) \times (k / T )$
# * Model selection consistent
# + [markdown] slideshow={"slide_type": "slide"}
# # Seasonality
#
# * Generated due to sampling frequency
# * Add dummies to control seasonality
# * How many dummies?
# + [markdown] slideshow={"slide_type": "slide"}
# # Vector Autoregression (VAR)
#
# <NAME> (<NAME> 2011)
#
# An $m$-equation system
# $$ y_t = \mu + \Gamma_1 y_{t-1} + \cdots + \Gamma_p y_{t-p} + v_t $$
# where $E[ v_t v_t'] = \Omega$.
#
# For prediction purpose, as a reduced-form of structural simultaneous equations.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Estimation
#
# * For consistency and asymptotic normality, use OLS equation by equation
# * For asymptotic efficient, use multiple-equation GLS
#
#
# + [markdown] slideshow={"slide_type": "skip"}
# # Invertibility
#
# Write the VAR(p) as
# $$ (I_m - \Gamma (L) ) y_t = \mu + v_t $$
# where $\Gamma(z) = \Gamma_1 z + \cdots + \Gamma_p z^p$.
#
# Stable means that all roots of the $p$th order polynomial equation $$ I_m - \Gamma(z) = 0_m $$ lies out of the unit circle.
# + [markdown] slideshow={"slide_type": "slide"}
# # Impulse Response Function
#
# IRF characterizes the diffusion of an exogenous shock with the dynamic system.
#
# $$
# \begin{align*}
# y_t & = (I_m - \Gamma(L) )^{-1} (\mu + v_t) \\
# & = \bar{y} + v_t + \sum_{i=1}^{\infty} A_i v_{t-i}
# \end{align*}
# $$ where $\bar{y} = (I_m - \Gamma(L) )^{-1} \mu = ( I_m + \sum_{i=1}^{\infty} A_i ) \mu $.
# + slideshow={"slide_type": "slide"}
library(tsDyn)
data(barry)
plot(barry)
# + slideshow={"slide_type": "subslide"}
## For VAR
mod_var <- lineVar(barry, lag = 2)
print(mod_var)
# + slideshow={"slide_type": "subslide"}
irf_var = irf(mod_var, impulse = "dolcan", response = c("dolcan", "cpiUSA", "cpiCAN"), boot = FALSE)
print(irf_var)
# + slideshow={"slide_type": "subslide"}
plot(irf_var)
# + slideshow={"slide_type": "subslide"}
## For VECM
mod_VECM <- VECM(barry, lag = 2, estim="ML", r=2)
print(mod_VECM)
irf_vecm = irf(mod_VECM, impulse = "dolcan", response = c("dolcan", "cpiUSA", "cpiCAN"), boot = FALSE)
print(irf_vecm)
# + slideshow={"slide_type": "subslide"}
plot(irf_vecm)
# + [markdown] slideshow={"slide_type": "slide"}
# # Structural VAR
#
# * Unrestricted VAR: too many parameters? $m+p\cdot m^2 + m(m+1)/2$
# * Use economic theory to reduce the number of unknown parameters
| Msc/time_series.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
os.chdir("../webapp")
from webapp_utility import Loader
import warnings
warnings.filterwarnings('ignore')
# -
l = Loader()
# ### Word frequency
l.get_freq_distribution(["gun", "robbery"], interval=10)
l.get_freq_distribution(["cocaine"], interval=20)
# ### Embeddings
l.get_n_similar(word="gun", n=3, model_type="full")
l.get_n_similar(word="gun", n=3, model_type="one", year=2000)
l.get_n_similar(word=["gun", "cocaine"], n=3, model_type="ten", year=2000)
# ### Topics
l.get_topic_dist(["gun"], model="small")
l.get_topic_dist(["gun", "cocaine"], model="big")
l.get_topics_words(n=5, model="big")
# l.get_topics_words(n=5, model="small")
l.get_topics_date_distribution(interval=1000)
l.get_topics_date_distribution(interval=1000, model="small")
# l.get_topics_court_distribution()
l.get_topics_court_distribution(model="small")
l.get_topics_description(topic_id=13, category="Generic"), l.get_topics_description(topic_id=0, category="Specific")
# ### Semantic
c = l.get_semantic_data(["cocaine", "cannabis"], base_year=2010)
print(c.keys())
c["one_year"]
c["ten_year"]
| notebooks/08_webapp_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
drinks=pd.read_csv("http://bit.ly/drinksbycountry")
pd.set_option("display.max_rows",None)
drinks
pd.reset_option("display.max_rows")
drinks
train=pd.read_csv("http://bit.ly/kaggletrain")
train
pd.set_option("display.max_colwidth",1000)
train
pd.set_option("display.precision",2)
train
drinks['x']=drinks.wine_servings*2000
drinks.head()
drinks['y']=drinks['total_litres_of_pure_alcohol']*2000
drinks.head()
pd.set_option('display.float_format','{:,}'.format)
drinks.head()
pd.describe_option()
pd.reset_option('all')
| Lecture Assignments/Lec28.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Caso de uso: Asistente virtual
# <div class="alert alert-warning">
#
# <i class="fa fa-bug"></i>
# Si no queréis perder los resultados de ejemplo, **no ejecutéis** el cuaderno de jupyter sin tener una clave propia de la API de OpenAI. Estos cuadernos solo cumplen una función ilustrativa de los códigos y formas de utilizar el modelo GPT-3 con Python. En caso de disponer de una **clave**, guardadla en un archivo **.env** para mayor seguridad como **un texto entrecomillado asignado a la variable OPENAI_API_KEY**.
# </div>
#
# ## Autentificación
#
# La API de OpenAI utiliza claves de API para la autentificación.
#
# <div class="alert alert-danger">
#
# <i class="fa fa-exclamation-circle"></i>
# Recuerda que tu clave de API es un secreto. No la compartas con otros ni la expongas en ningún código del lado del cliente (navegadores, aplicaciones). Las solicitudes de producción deben dirigirse a través de su propio servidor *backend*, donde su clave de API puede cargarse de forma segura desde una variable de entorno o un servicio de gestión de claves.
#
# </div>
#
# Todas las solicitudes de API deben incluir su clave de API, es importante almacenar en un documento seguro la llave. Para ello, crea un archivo nuevo `.env` para almacenarla en su interior de la siguiente forma:
#
# `OPENAI_API_KEY = "MI_API_KEY"`
#
# Con esto, para recuperar la clave de la API, tendremos que usar el **getenv** de `os`.
# +
import openai
import os
from dotenv import load_dotenv
load_dotenv()
openai.api_key = os.getenv("OPENAI_API_KEY")
# -
# ## Introducción
#
# Uno de los posibles usos que se le puede dar a un modelo del lenguaje es la generación de textos y la resolución de preguntas, que es lo que suele conocerse como *chatbot* o asistente virtual.
#
# Para poder crear uno, OpenAi tiene el módulo *Answer* incorporado y, construyéndolo correctamente, nos permitirá generar una herramienta que buscará información en un documento y nos la mostrará.
#
# Para este supuesto, además, he querido introducir una herramienta de extracción de textos en pdf: **pdfminer**, que nos permite acceder al texto de estos documentos. He preparado, para ello, una extracción de texto de la página de Wikipedia en español que corresponde a la información del Covid-19 (la enfermedad), con la intención de generar un asistente que sea capaz de responder con lo que encuentre en ese documento.
#
# La elección del Covid-19 es intencionada, ya que GPT-3 fue congelado en octubre de 2019, cuando aún no existía y, por lo tanto, *no conoce* lo que realmente es.
#
# Para utilizar correctamente este cuaderno necesitas tener instalado **pdfminer** y **jsonlines**.
# +
#Instalar pdfminer si no lo tienes instalado
# #!pip3 install pdfminer.six
# #!pip3 install jsonlines
# -
# Con **extract_text** de pdfminer se extraerá el texto del documento pdf.
#
# Con la función que tenéis abajo: **extractor_texto** podemos, no solo extraer el texto, sino que nos permite eliminar artefactos y elementos que no nos interesa mantener en el documento.
#
# Para ello, después de extraer el texto, reemplaza elementos indeseados (como la codificación del salto de página —\n\n\x0c—, puntos suspensivos, dobles saltos de línea, espacios en blanco y otros elementos de formato. Luego unirá todo el texto en una única cadena de texto, separando los párrafos con *\n*.
#
# El extractor, además, hará una copia y guardará una copia del documento en txt.
# +
from pdfminer.high_level import extract_text
import codecs
ruta = 'documentos/Covid'
def extractor_texto(ruta):
txt = extract_text(ruta + '.pdf')
txt = txt.replace('\n\n\x0c', ' ').replace('...', ' ').split(sep='\n\n')
txt = [paragraph for paragraph in txt if paragraph!=' ']
txt = [paragraph.replace('\n',' ').replace(' ', ' ').replace("\f", ' ').replace("-", ' ') for paragraph in txt if len(paragraph)>30]
txt = '\n'.join(txt)
return txt
archivo = codecs.open(ruta + '.txt', "w", "utf-16")
archivo.write(txt)
archivo.close()
covid = extractor_texto(ruta)
covid
# -
# ### Carga el archivo en OpenAi
#
# Una de las cosas que podemos hacer es cargar archivos a OpenAi para que GPT-3 pueda extraer información controlada. Hay que tener en cuenta las limitaciones que impone para hacerlo: **máximo de un 1 Gb de archivos**.
#
# Para poder subirlos, debemos tratarlos antes y subirlos formateados en **jsonlines**; lo siguiente que vamos a hacer es generar este archivo *.jsonl* y escribir lo mínimo que necesita para admitirlo.
#
# Dentro del **jsonlines**, debemos guardar, en la clave **text** todo el texto que queremos introducir para que GPT-3 lo utilice para búsquedas. Dentro de este jsonlines también podemos introducir metadatos (los metadatos, sobre todo, son necesarios cuando nos estemos enfrentando a una clasificación junto con la clave *label*).
# +
import jsonlines
# ------------------------------------------- Crea el archivo jsonlines para poderlo subir a OpenAi
archivo=jsonlines.open("documentos/covid.jsonl","w")
jsonlines.Writer.write(archivo, {"text": covid, "metadata": "Información extraida de Wikipedia sobre el Covid-19."})
archivo.close()
# ------------------------------------------- Carga el documento en OpenAi
openai.File.create(file=open('documentos/covid.jsonl'), purpose='answers')
# -
# ### Asistente virtual: preguntas y respuestas
#
# Una vez tenemos el archivo de referencia subido a OpenAi, podemos comenzar a genera la petición del modelo de pregunta y respuesta; armarlo es un poco más complejo que una petición de generar texto. Para hacerlo correctamente, necesitamos generar la petición teniendo en cuenta lo que necesita para responder correctamente.
# 1. Definir el modelo de búsqueda en el documento: **search_model**.
# 2. Definir el modelo que generará la respuesta: **model**.
# 3. Definir la pregunta que queremos que resuelva: **question**.
# 4. Archivo o documento donde deberá encontrar la respuesta: **file**. En este caso, la única referencia que necesita es especificar el *id* que le ha asignado OpenAi al archivo.
# 5. Para hacerse una idea, necesitamos crearle un ejemplo de contexto en el que aportando un texto determinado en **examples_context**, tendrá una plantilla de pregunta/respuesta en **examples**; es bueno proporcionarle varios ejemplos de pregunta/respuesta que puede esperar.
# 6. Número máximo de palabras que va a generar como respuesta: **max_tokens**.
# 7. Elementos que harán que finalice en **stop**; en este caso, quiero que dé respuestas cortas, por lo que voy a evitar que genere saltos de línea y más de una frase especificándole que finalice con puntos.
# +
pregunta = '¿Cuáles son los síntomas del COVID-19?'
respuesta = openai.Answer.create(
search_model = "ada", #-------------- Modelo que utiliza para la búsqueda de información en el documento
model = "curie", #------------------- Modelo que utiliza para generar la respuesta
question = pregunta,
file = 'file-aujfELqAJSovNnxsHVn456cJ',
examples_context = "La pandemia de COVID-19 (conocida popularmente como pandemia de coronavirus) es una pandemia derivada de la enfermedad ocasionada por el virus SARS-CoV-2. Su primer caso fue identificado en diciembre de 2019 en la ciudad de Wuhan, capital de la provincia de Hubei, en la República Popular China, al reportarse casos de un grupo de personas enfermas con un tipo de neumonía desconocida. La mayoría de los individuos afectados tenían vinculación con trabajadores del Mercado mayorista de mariscos de Huanan. La Organización Mundial de la Salud (OMS) la reconoció como una pandemia el 11 de marzo de 2020 (cuando informó que había 4291 muertos y 118 000 casos en 114 países).",
examples = [["¿Qué virus originó la pandemia?","Fue ocasionada por el virus SARS-CoV-2."],
["¿Dónde se detectó por primera vez?", "Se detectó por primera vez en Wuhan, en China"],
["¿Cuándo reconoció la OMS la situación de pandemia?", "El 11 de marzo de 2020"],
["Me gustaría saber las medidas para evitar el contagio", "Es muy importante el uso de la mascarilla, tanto en exteriores como en interiores, también respetar la distancia de seguridad de 2 metros y lavarse habitualmente las manos."]],
max_tokens = 70,
stop = ["\n", "<|endoftext|>", "."])
respuesta["answers"]
# -
# Uno de los problemas que podemos encontrarnos, teniendo en cuenta que serán usuarios humanos los que interactuarán con el asistente, es que muchas veces nos gusta a todos buscarle las cosquillas y los límites al asistente saliéndonos del patrón de preguntas.
#
# Por eso, podemos hacer un poco más complejo el sistema haciendo que el asistente intente generar una respuesta según la petición de pregunta/respuesta y, en el caso de que no sea capaz de generarlo porque la entrada del usuario sea insuficiente, salte a generar texto nuevo actuando, en este caso, como un virólogo.
#
# **¿Por qué le doy la información de que responda como si tuviese el rol de virólogo?** Como estamos trabajando con un asistente virtual que busca información sobre el Covid-19 y sabemos que GPT-3 no tiene datos de esta enfermedad en su entrenamiento; no podemos pedirle ni facilitarle información de este virus ni de lo que ha ocurrido en los últimos dos años; lo que sí podemos hacer es guiarlo para que tenga un rol general de virólogo y que pueda ajustar la respuesta según eso.
#
# Hay otros condicionantes que no estoy teniendo en cuenta, por supuesto, a la hora de trabajar este asistente, porque es un ejemplo simple de cómo crear una base de asistente virtual con GPT-3; podemos buscar y encontrar una estructura más compleja y segura para que interactue de forma más eficiente.
# +
def asistente_virtual(entrada):
try:
respuesta = openai.Answer.create(
search_model = "babbage", #---------- Modelo que utiliza para la búsqueda de información en el documento
model = "curie", #------------------- Modelo que utiliza para generar la respuesta
question = entrada,
file = 'file-aujfELqAJSovNnxsHVn456cJ',
examples_context = "La pandemia de COVID-19 (conocida popularmente como pandemia de coronavirus) es una pandemia derivada de la enfermedad ocasionada por el virus SARS-CoV-2. Su primer caso fue identificado en diciembre de 2019 en la ciudad de Wuhan, capital de la provincia de Hubei, en la República Popular China, al reportarse casos de un grupo de personas enfermas con un tipo de neumonía desconocida. La mayoría de los individuos afectados tenían vinculación con trabajadores del Mercado mayorista de mariscos de Huanan. La Organización Mundial de la Salud (OMS) la reconoció como una pandemia el 11 de marzo de 2020 (cuando informó que había 4291 muertos y 118 000 casos en 114 países). Medidas para evitar el contagio: uso de mascarilla en exteriores e interiores, también cuando no pueda garantizarse la distancia mínima de seguridad, lavarse habitualmente las manos y distanciamiento social de al menos 2 metros.",
examples = [["¿Qué virus originó la pandemia?","Fue ocasionada por el virus SARS-CoV-2."],
["¿Dónde se detectó por primera vez?", "Se detectó por primera vez en Wuhan, en China"],
["¿Cuándo reconoció la OMS la situación de pandemia?", "El 11 de marzo de 2020"],
["Me gustaría saber las medidas para evitar el contagio", "Es muy importante el uso de la mascarilla, tanto en exteriores como en interiores, también respetar la distancia de seguridad de 2 metros y lavarse habitualmente las manos."]],
max_tokens = 70,
stop = ["\n", "<|endoftext|>", "."],
temperature= 0.73
)
return str(respuesta["answers"])
except:
respuesta = openai.Completion.create(engine="curie",
prompt='Usuario: ' + entrada + '\n\nVirólogo:',
max_tokens=70,
temperature= 0.73,
stop = ["\n", "<|endoftext|>", "Usuario:", "Virólogo:", ":"],)
return str(respuesta['choices'][0]['text'])
entrada = '¿Qué medidas tengo para evitar contagios por Covid?'
print(entrada)
print(asistente_virtual(entrada))
entrada = 'Hoy parece que va a llover…'
print(entrada)
print(asistente_virtual(entrada))
# -
entrada = 'La tierra es plana'
print(entrada)
print(asistente_virtual(entrada))
# ### Eliminar el archivo de OpenAi
#
# Dado que tenemos limitaciones de espacio para almacenar información en la nube a la que GPT-3 puede tener acceso, es conveniente mantener solo los documentos que sean útiles; lo que quiere decir que necesitamos saber cómo eliminar documentos.
#
# Con **openai.File.list()** podemos identificar los archivos que tenemos subidos; luego, con especificar el *id* en la instrucción **openai.File("id_objeto_a_eliminar").delete()**, podemos liberar el espacio que ocupa ese archivo.
openai.File.list()
openai.File("file-xiUg3HpWjA9idRbPBiZv4lpA").delete()
openai.File.list()
# ### Conclusiones
#
# GPT-3 nos permite generar un asistente virtual capaz de responder preguntas de forma bastante rápido y con pocas líneas de código. Hay que tener en cuenta que el modelo trabaja mejor cuantos más ejemplos tenga para interpretar; habría entonces que hacer un trabajo de contextualización del problema específico que va a solucionar el asistente y, dotarlo, también de recursos auxiliares que le permitan emular mejor la naturalidad de una conversación.
# <div class="alert alert-info">
#
# <i class="fa fa-code"></i> **Este cuaderno ha sido creado con la ayuda de GPT-3.**
# <hr/>
#
# **Si tienes alguna duda relacionada con estos cuadernos, puedes contactar conmigo:**
# Mª <NAME>. (Erebyel). **[web](https://www.erebyel.es/) • [Twitter](https://twitter.com/erebyel) • [Linkedin](https://www.linkedin.com/in/erebyel/)**.
#
# <hr/>
#
# <i class="fa fa-plus-circle"></i> **Fuentes:**
#
# * ***Documentación de la Beta de OpenAI***: https://beta.openai.com/docs/introduction
# * ***Wikipedia: Covid-19***: https://es.wikipedia.org/wiki/COVID-19
# </div>
| 2.3_Ejemplo de un caso de uso_Asistente virtual.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 用 GraphScope 像 NetworkX 一样进行图分析
#
# GraphScope 在大规模图分析基础上,提供了一套兼容 NetworkX 的图分析接口。
# 本文中我们将介绍如何用 GraphScope 像 NetworkX 一样进行图分析。
# ### NetworkX 是如何进行图分析的
#
# NetworkX 的图分析过程一般首先进行图的构建,示例中我们首先建立一个空图,然后通过 NetworkX 的接口逐步扩充图的数据。
# +
# Install graphscope package if you are NOT in the Playground
# !pip3 install graphscope
# -
import networkx
# 初始化一个空的无向图
G = networkx.Graph()
# 通过 add_edges_from 接口添加边列表,此处添加了两条边(1, 2)和(1, 3)
G.add_edges_from([(1, 2), (1, 3)])
# 通过 add_node 添加点4
G.add_node(4)
# 接着使用 NetworkX 来查询一些图的信息
# 使用 G.number_of_nodes 查询图G目前点的数目
G.number_of_nodes()
# 类似地,G.number_of_edges 可以查询图G中边的数量
G.number_of_edges()
# 通过 G.degree 来查看图G中每个点的度数
sorted(d for n, d in G.degree())
# 最后通过调用 NetworkX 内置的算法对图G进行分析
# 调用 connected_components 算法分析图G的联通分量
list(networkx.connected_components(G))
# 调用 clustering 算法分析图G的聚类情况
networkx.clustering(G)
# ### 如何使用 GraphScope 的 NetworkX 接口
# 图的构建
#
# 使用 GraphScope 的 NetworkX 兼容接口,我们只需要简单地将教程中的`import netwokx as nx`替换为`import graphscope.nx as nx`即可。
# GraphScope 支持与 NetworkX 完全相同的载图语法,这里我们使用`nx.Graph()`来建立一个空的无向图。
import graphscope.nx as nx
# 我们可以建立一个空的无向图
G = nx.Graph()
# 增加节点和边
#
# GraphScope 的图操作接口也保持了与 NetworkX 的兼容,用户可以通过`add_node`和 `add_nodes_from`来添加节点,通过`add_edge`和`add_edges_from`来添加边。
# +
# 通过 add_node 一次添加一个节点
G.add_node(1)
# 或从任何 iterable 容器中添加节点,如列表
G.add_nodes_from([2, 3])
# 如果容器中是元组的形式,还可以在添加节点的同时,添加节点属性
G.add_nodes_from([(4, {"color": "red"}), (5, {"color": "green"})])
# 对于边,可以通过 add_edge 的一次添加一条边
G.add_edge(1, 2)
e = (2, 3)
G.add_edge(*e)
# 通过 add_edges_from 添加边列表
G.add_edges_from([(1, 2), (1, 3)])
# 或者通过边元组的方式,在添加边的同时,添加边的属性
G.add_edges_from([(1, 2), (2, 3, {'weight': 3.1415})])
# -
# 查询图的元素
#
# GraphScope 支持兼容 NetworkX 的图查询接口。用户可以通过`number_of_nodes`和`number_of_edges`来获取图点和边的数量,通过`nodes`, `edges`,`adj`和`degree`等接口来获取图当前的点和边,以及点的邻居和度数等信息。
# 查询目前图中点和边的数目
G.number_of_nodes()
G.number_of_edges()
# 列出目前图中的点和边
list(G.nodes)
list(G.edges)
# 查询某个点的邻居
list(G.adj[1])
# 查询某个点的度
G.degree(1)
# 从图中删除元素
#
# 像 NetworkX 一样, GraphScope 也可以使用与添加元素相类似的方式从图中删除点和边,对图进行修改。例如可以通过`remove_node`和`remove_nodes_from`来删除图中的节点,通过`remove_edge`和`remove_edges_from`来删除图中的边。
# 通过 remove_node 删除一个点
G.remove_node(5)
list(G.nodes)
G.remove_nodes_from([4, 5])
list(G.nodes)
# 通过 remove_edge 删除一条边
G.remove_edge(1, 2)
list(G.edges)
# 通过 remove_edges_from 删除多条边
G.remove_edges_from([(1, 3), (2, 3)])
list(G.edges)
# 我们再来看一下现在的点和边的数目
G.number_of_nodes()
G.number_of_edges()
# 图分析
#
# GraphScope 可以通过兼容 NetworkX 的接口来对图进行各种算法的分析,示例里我们构建了一个简单图,然后分别使用`connected_components`分析图的联通分量,使用`clustering`来得到图中每个点的聚类系数,以及使用`all_pairs_shortest_path`来获取节点两两之间的最短路径。
# 首先构建图
G = nx.Graph()
G.add_edges_from([(1, 2), (1, 3)])
G.add_node(4)
# 通过 connected_components 算法找出图中的联通分量
list(nx.connected_components(G))
# 通过 clustering 算法计算每个点的聚类系数
nx.clustering(G)
sp = dict(nx.all_pairs_shortest_path(G))
sp[3]
# 图的简单绘制
#
# 同 NetworkX 一样,GraphScope支持通过`draw`将图进行简单地绘制出来,底层依赖的是`matplotlib`的绘图功能。
# 如果系统未安装`matplotlib`, 我们首先需要安装一下`matplotlib`包
# !pip3 install matplotlib
# 使用 GraphScope 来进行简单地绘制图
# 创建一个5个点的 star graph
G = nx.star_graph(5)
# 使用 nx.draw 绘制图
nx.draw(G, with_labels=True, font_weight='bold')
# ### GraphScope 相对 NetworkX 算法性能上有着数量级的提升
# 我们通过一个简单的实验来看一下 GraphScope 对比 NetworkX 在算法性能上到底提升多少。
#
# 实验使用来自 snap 的 [twitter] 图数据(https://snap.stanford.edu/data/ego-Twitter.html), 测试算法是 NetworkX 内置的 [clustering](https://networkx.org/documentation/stable/reference/algorithms/generated/networkx.algorithms.cluster.clustering.html#networkx.algorithms.cluster.clustering) 算法。
# 我们首先准备下数据,使用 wget 将数据集下载到本地
# !wget https://raw.githubusercontent.com/GraphScope/gstest/master/twitter.e -P /tmp
# 接着我们分别使用 GraphScope 和 NetworkX 载入 snap-twitter 数据
import os
import graphscope.nx as gs_nx
import networkx as nx
# 使用 NetworkX 载入 snap-twitter 图数据
g1 = nx.read_edgelist(
os.path.expandvars('/tmp/twitter.e'), nodetype=int, data=False, create_using=nx.Graph
)
type(g1)
# 使用 GraphScope 载入相同的 snap-twitter 图数据
g2 = gs_nx.read_edgelist(
os.path.expandvars('/tmp/twitter.e'), nodetype=int, data=False, create_using=gs_nx.Graph
)
type(g2)
# 最后我们使用 clustering 算法来对图进行聚类分析,来看一下 GraphScope 对比 NetworkX 在算法性能上有多少提升
# %%time
# 使用 GraphScope 计算图中每个点的聚类系数
ret_gs = gs_nx.clustering(g2)
# %%time
# 使用 NetworkX 计算图中每个点的聚类系数
ret_nx = nx.clustering(g1)
# 对比下两者的结果是否一致
ret_gs == ret_nx
| tutorials/zh/03_run_graphscope_like_networkx.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
import matplotlib.cm as cm
# +
pageSpeeds = np.random.normal(3.0, 1.0, 1000)
purchaseAmount = 100 - (pageSpeeds + np.random.normal(0, 0.1, 1000)) * 3
plt.scatter(pageSpeeds, purchaseAmount)
# +
from scipy import stats
slope, intercept, r_value, p_value, std_err = stats.linregress(pageSpeeds, purchaseAmount)
# -
r_value ** 2
# +
def predict(x):
return slope * x + intercept
fitLine = predict(pageSpeeds)
plt.scatter(pageSpeeds, purchaseAmount)
plt.plot(pageSpeeds, fitLine, c='r')
plt.show()
# -
| 3.1_Linear_Regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## 学习笔记
#
#
# ### 学习资料
#
# #### Computer Science
#
# [MIT线性代数](https://ocw.mit.edu/courses/find-by-topic/#cat=mathematics&subcat=linearalgebra)
#
# [MIT概率论](https://ocw.mit.edu/courses/find-by-topic/#cat=mathematics&subcat=probabilityandstatistics)
#
# https://ocw.mit.edu/courses/find-by-topic/#cat=engineering&subcat=computerscience
#
#
# #### tensorflow statar
#
# [Introduction to Machine Learning by Google Developer ](https://developers.google.com/machine-learning/crash-course/)
#
# [Get Started with TensorFlow](https://www.tensorflow.org/tutorials/)
#
# #### Convolutional Neural Networks for Visual Recognition
#
# [TensorFlow and deep learning, without a PhD](https://codelabs.developers.google.com/codelabs/cloud-tensorflow-mnist/#0)
#
# [UFLDL Tutorial](http://ufldl.stanford.edu/tutorial/)
#
# [CS231n: Convolutional Neural Networks for Visual Recognition](http://cs231n.stanford.edu/)
#
# [Convolutional Neural Networks for Visual Recognition (Spring 2017)](https://www.youtube.com/playlist?list=PL3FW7Lu3i5JvHM8ljYj-zLfQRF3EO8sYv)
| study_notes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import os
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
from keras.models import load_model
from preprocessing.image import extract_features, extract_feature_from_image
from preprocessing.text import create_tokenizer
from NIC import greedy_inference_model, image_dense_lstm, text_emb_lstm
from evaluate import decoder, beam_search
# -
# use training token set to create vocabulary
train_dir = './datasets/Flickr8k_text/Flickr_8k.trainImages.txt'
token_dir = './datasets/Flickr8k_text/Flickr8k.token.txt'
# the current best trained model
model_dir = './model-params-his/current_best.h5'
# +
# load vocabulary
tokenizer = create_tokenizer(train_dir, token_dir, start_end = True, use_all=True)
# set relevent parameters
vocab_size = tokenizer.num_words or (len(tokenizer.word_index)+1)
max_len = 24 # use 24 as maximum sentence's length when training the model
# -
# ## Greedy inference
NIC_inference = greedy_inference_model(vocab_size, max_len)
NIC_inference.load_weights(model_dir, by_name = True, skip_mismatch=True)
# +
def generate_caption_from_file(file_dir):
# Encoder
img_feature = extract_feature_from_image(file_dir)
# Decoder
caption = decoder(NIC_inference, tokenizer, img_feature, True)
return caption
def generate_caption_from_directory(file_directory):
# Encoder
img_features_dict = extract_features(file_directory)
# Decoder
captions = decoder(NIC_inference, tokenizer, img_features_dict['features'], True)
return img_features_dict['ids'], captions
# +
image_file_dir = './put-your-image-here/example.jpg'
# display image
img = mpimg.imread(image_file_dir)
plt.imshow(img)
#generate caption
caption = generate_caption_from_file(image_file_dir)
plt.show()
print(caption)
# +
image_dir = './put-your-image-here'
img_names, captions = generate_caption_from_directory(image_dir)
for img_file in os.listdir(image_dir):
img = mpimg.imread(image_dir + '/' + img_file)
plt.imshow(img)
img_name = os.path.splitext(img_file)[0]
idx = img_names.index(img_name)
plt.show()
print(captions[idx])
# -
# ## Beam search inference
# prepare inference model
NIC_text_emb_lstm = text_emb_lstm(vocab_size)
NIC_text_emb_lstm.load_weights(model_dir, by_name = True, skip_mismatch=True)
NIC_image_dense_lstm = image_dense_lstm()
NIC_image_dense_lstm.load_weights(model_dir, by_name = True, skip_mismatch=True)
# +
def generate_caption_from_file(file_dir, beam_width = 5, alpha = 0.7):
# Encoder
img_feature = extract_feature_from_image(file_dir)
# Decoder
a0, c0 = NIC_image_dense_lstm.predict([img_feature, np.zeros([1, 512]), np.zeros([1, 512])])
res = beam_search(NIC_text_emb_lstm, a0, c0, tokenizer, beam_width, max_len, alpha)
best_idx = np.argmax(res['scores'])
caption = tokenizer.sequences_to_texts([res['routes'][best_idx]])[0]
return caption
def generate_caption_from_directory(file_directory, beam_width = 5, alpha = 0.7):
# Encoder
img_features_dict = extract_features(file_directory)
# Decoder
N = img_features_dict['features'].shape[0]
a0, c0 = NIC_image_dense_lstm.predict([img_features_dict['features'], np.zeros([N, 512]), np.zeros([N, 512])])
captions = []
for i in range(N):
res = beam_search(NIC_text_emb_lstm, a0[i, :].reshape(1,-1), c0[i, :].reshape(1,-1), tokenizer, beam_width, max_len, alpha)
best_idx = np.argmax(res['scores'])
captions.append(tokenizer.sequences_to_texts([res['routes'][best_idx]])[0])
return img_features_dict['ids'], captions
# +
image_file_dir = './put-your-image-here/example.jpg'
# display image
img = mpimg.imread(image_file_dir)
plt.imshow(img)
#generate caption
caption = generate_caption_from_file(image_file_dir)
plt.show()
print(caption)
# +
image_dir = './put-your-image-here'
img_names, captions = generate_caption_from_directory(image_dir, 10, 1)
for img_file in os.listdir(image_dir):
img = mpimg.imread(image_dir + '/' + img_file)
plt.imshow(img)
img_name = os.path.splitext(img_file)[0]
idx = img_names.index(img_name)
plt.show()
print(captions[idx])
| NIC Interactive.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# <NAME>
# 20180723
# A script that recommends stock based on data and conditions given.
# Import Statement
from selenium import webdriver
from bs4 import BeautifulSoup
from decimal import Decimal
from selenium.common.exceptions import ElementNotVisibleException
import turicreate as tc
import datetime
fileRoot = './SelectedData'
# 执行UI
var_list= []
# 新建精选数据库
SELECTED_DATA = tc.SFrame({'code': ['000000'], 'name': ['数据不存在'],'bankuai': ['二次元'],
'close': [0.0], 'percent_chg': [0.0],'change': [0.0],
'volume': [0.0], 'turn_volume': [0.0],
'amplitude': [0.0],'volume_rate': [0.0],
'turnover_rate': [0.0],
'news_url': ['http://www.bilibili.com'],
'income_increase': [0.0], 'profit_increase': [0.0],
'PERation': [0.0]})
code_dict = {'a': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK04471',
'b': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK04931',
'c': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK05231',
'd': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK06991',
'e': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK07271',
'f': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK04741',
'g': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK05381',
'h': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK07311',
'i': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK04781',
'j': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK04791',
'k': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK04561',
'l': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK07331',
'm': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK04241',
'n': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK07321',
'o': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK07361',
'p': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK04801',
'q': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK04761',
'r': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK07381',
's': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK04381',
't': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK05381',
'u': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK04241',
'v': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK07361',
'w': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK04561',
'x': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK09101',
'y': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK07401',
'z': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK04221',
'aa': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK04541',
'bb': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK07391',
'cc': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK04571',
'dd': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK04641',
'ee': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK05451',
'ff': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK07281',
'gg': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK04851',
'hh': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK07291',
'ii': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK07351',
'jj': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK04511',
'kk': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK04751',
'll': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK04811',
'mm': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK07251',
'nn': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK07391',
'oo': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK07261',
'pp': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK04731',
'qq': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK04501',
'rr': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK04281',
'ss': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK04701',
'tt': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK04571',
'uu': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK07311',
'vv': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK04291',
'ww': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK07301',
'xx': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK05391',
'yy': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK05371',
'zz': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK04861',
'aaa': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK04841',
'bbb': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK07371',
'ccc': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK04471',
'ddd': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK04591',
'eee': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK04651',
'fff': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK07331',
'ggg': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK04331',
'hhh': 'http://quote.eastmoney.com/center/boardlist.html#boards-BK04771'}
name_dict = {'电子信息': 'a',
'新能源':'b',
'新材料':'c',
'全息技术':'d',
'医疗行业':'e',
'保险':'f',
'化工行业':'g',
'化肥行业':'h',
'有色金属':'i',
'钢铁行业':'j',
'家电行业':'k',
'包装材料':'l',
'水泥建材':'m',
'贵金属':'n',
'电信运营':'o',
'航天航空':'p',
'木业家具':'q',
'多元金融':'r',
'食品饮料':'s',
'化工行业':'t',
'水泥建材':'u',
'电信运营':'v',
'家电行业':'w',
'专用设备':'x',
'文教休闲':'y',
'交运物流':'z',
'塑胶制品':'aa',
'金属制品':'bb',
'输配电气':'cc',
'石油行业':'dd',
'机械行业':'ee',
'环保工程':'ff',
'旅游酒店':'gg',
'船舶制造':'hh',
'安防设备':'ii',
'房地产':'jj',
'银行':'kk',
'汽车行业':'ll',
'装修装饰':'mm',
'金属制品':'nn',
'园林工程':'oo',
'券商信托':'pp',
'港口水运':'qq',
'电力行业':'rr',
'造纸印刷':'ss',
'输配电气':'tt',
'化肥行业':'uu',
'交运设备':'vv',
'农药兽药':'ww',
'综合行业':'xx',
'材料行业':'yy',
'文化传媒':'zz',
'国际贸易':'aaa',
'软件服务':'bbb',
'电子信息':'ccc',
'电子元件':'ddd',
'医药制造':'eee',
'包装材料':'fff',
'农牧饲渔':'ggg',
'酿酒行业':'hhh'}
codename_dict = {'a': '电子信息',
'b': '新能源',
'c': '新材料',
'd': '全息技术',
'e': '医疗行业',
'f': '保险',
'g': '化工行业',
'h': '化肥行业',
'i': '有色金属',
'j': '钢铁行业',
'k': '家电行业',
'l': '包装材料',
'm': '水泥建材',
'n': '贵金属',
'o': '电信运营',
'p': '航天航空',
'q': '木业家具',
'r': '多元金融',
's': '食品饮料',
't':'化工行业',
'u':'水泥建材',
'v':'电信运营',
'w':'家电行业',
'x':'专用设备',
'y':'文教休闲',
'z':'交运物流',
'aa':'塑胶制品',
'bb':'金属制品',
'cc':'输配电气',
'dd':'石油行业',
'ee':'机械行业',
'ff':'环保工程',
'gg':'旅游酒店',
'hh':'船舶制造',
'ii':'安防设备',
'jj':'房地产',
'kk':'银行',
'll':'汽车行业',
'mm':'装修装饰',
'nn':'金属制品',
'oo':'园林工程',
'pp':'券商信托',
'qq':'港口水运',
'rr':'电力行业',
'ss':'造纸印刷',
'tt':'输配电气',
'uu':'化肥行业',
'vv':'交运设备',
'ww':'农药兽药',
'xx':'综合行业',
'yy':'材料行业',
'zz':'文化传媒',
'aaa':'国际贸易',
'bbb':'软件服务',
'ccc':'电子信息',
'ddd':'电子元件',
'eee':'医药制造',
'fff':'包装材料',
'ggg':'农牧饲渔',
'hhh':'酿酒行业'}
bankuai_weight_dict = {'软件服务': 10,
'航天航空': 10, # 再议
'新能源': 9, # 再议
'新材料': 9,
'材料行业': 9, # 再议
'电子信息': 8,
'有色金属': 4,
'?': 3, # 这啥
'钢铁行业': 2}
# +
# 一个从页面获取页数的函数
def getPageNumber(bs):
all_buttons = bs.findAll(class_="paginate_button")
if len(all_buttons) == 2:
return 1 # 处理只有一页的情况
else:
return len(all_buttons) - 2 # 下一页和Go按钮
# 一个自动判断量词的函数
def smartMultiply(string):
if string[len(string) - 1:len(string)] == '万':
string = Decimal(string[0:len(string) - 1])
string = float(string) * 10000
elif string[len(string) - 1:len(string)] == '亿':
string = Decimal(string[0:len(string) - 1])
string = float(string) * 100000000
elif string[len(string) - 1:len(string)] == '%':
string = Decimal(string[0:len(string) - 1])
string = float(string) * 0.01
else:
string = float(string)
return string
# 把数据中的 - 改为 0
def noSlash(str):
if str == '-':
return '0'
else:
return str
# 从一个静态BeautifulSoup页面解析表格并存储进SFrame
def grabData(bs, SFrame):
# 解出表格
table = bs.findAll(role='row')
table = table[7: len(table) - 1]
# 分析每个表格
counter = 0
while counter < len(table):
row_sframe = tc.SFrame({'code': [str(table[counter].find(class_=' listview-col-Code').string)],
'name': [str(table[counter].find(class_=' listview-col-Name').string)],
'close': [smartMultiply(noSlash(table[counter].find(class_=' listview-col-Close').string))],
'percent_chg': [smartMultiply(noSlash(table[counter].find(class_='listview-col-ChangePercent sorting_1').string))],
'change': [smartMultiply(noSlash(table[counter].find(class_=' listview-col-Change').string))],
'volume': [smartMultiply(noSlash(table[counter].find(class_=' listview-col-Volume').string))],
'turn_volume': [
smartMultiply(noSlash(table[counter].find(class_=' listview-col-Amount').string))],
'amplitude': [
smartMultiply(noSlash(table[counter].find(class_=' listview-col-Amplitude').string))],
'high': [smartMultiply(noSlash(table[counter].find(class_=' listview-col-High').string))],
'low': [smartMultiply(noSlash(table[counter].find(class_=' listview-col-Low').string))],
'now_open': [smartMultiply(noSlash(table[counter].find(class_=' listview-col-Open').string))],
'previous_close': [
smartMultiply(noSlash(table[counter].find(class_=' listview-col-PreviousClose').string))],
'volume_rate': [
smartMultiply(noSlash(table[counter].find(class_=' listview-col-VolumeRate').string))],
'turnover_rate': [
smartMultiply(noSlash(table[counter].find(class_=' listview-col-TurnoverRate').string))],
'report_url': [
'http://emweb.securities.eastmoney.com/f10_v2/FinanceAnalysis.aspx?type=web&code=sz' +
table[counter].find(class_=' listview-col-Code').string + '#lrb-0'],
'PERation':[
smartMultiply(noSlash(table[counter].find(class_=' listview-col-PERation').string))]
})
counter += 1
SFrame = SFrame.append(row_sframe)
return SFrame
# 自动处理数据的主程序
def makeData(url, SFrame):
browser = webdriver.Chrome() # Get local session of chrome
# url = search_area[topic] # Example: '电子信息'
browser.get(url) # Load page
browser.implicitly_wait(20) # 智能等待20秒
# 第一次访问时判定菜单数量来决定浏览多少次表格
bs = BeautifulSoup(browser.page_source, "lxml")
page_number = getPageNumber(bs)
# 循环浏览页面直到搜集完毕所有table
counter = 0
while counter < page_number:
SFrame = grabData(bs, SFrame)
try:
browser.find_element_by_id('main-table_next').click()
except ElementNotVisibleException:
print('Warning: 无法获得某些破损的数据.')
bs = BeautifulSoup(browser.page_source, "lxml")
counter += 1
SFrame = SFrame[1:len(SFrame)] # 删掉占位符
SFrame = SFrame.unique()
return SFrame
# 初步筛选分析程序
def analyze_stock(SFrame):
SFrame = analysis_turnover_rate(SFrame, var_list[0])
SFrame = analysis_volume_rate(SFrame, var_list[1])
return SFrame
# 返回所有换手率大于5%的行
def analysis_turnover_rate(SFrame, turnover_rate):
return SFrame[SFrame['turnover_rate'] > turnover_rate]
# 返回所有量比大于30%的行
def analysis_volume_rate(SFrame, volume_rate):
return SFrame[ SFrame['volume_rate'] > volume_rate]
# 查找报表
def getReport(xuangu, SFrame, bankuai, row, income_limit, profit_limit):
# 爬取网站源代码
url = row['report_url']
browser = webdriver.Chrome() # Get local session of chrome
browser.get(url) # Load page
soup = BeautifulSoup(browser.page_source, "lxml")
browser.close()
# 检查是否报表模板错误
check_report = soup.findAll(id='stock_full_name123')
if len(check_report) != 0:
# print(check_report)
# print(check_report[0])
# print(check_report[0]['value'])
keyword = check_report[0]['value']
if keyword == '- - - -': # 纠正SFrame
# print(keyword)
temp_row = row # 解决无法显示上证股票报表的问题
temp_row['report_url'] = temp_row['report_url'][0:80] + 'sh' + temp_row['report_url'][82:94]
# print(temp_row['report_url'])
return getReport(xuangu, SFrame, bankuai, temp_row, income_limit, profit_limit)
# 粗加工数据
ulist = []
trs = soup.find_all('tr')
for tr in trs:
ui = []
for td in tr:
ui.append(td.string)
ulist.append(ui)
# 提取营业额和净利润
income_increase = 0
profit_increase = 0
for element in ulist:
if ('营业总收入' in element):
income_data_list = element
now_data = smartMultiply(income_data_list[3])
past_data = smartMultiply(income_data_list[11])
income_increase = (now_data - past_data) / past_data
elif ('净利润' in element):
profit_data_list = element
now_data = smartMultiply(profit_data_list[3])
past_data = smartMultiply(profit_data_list[11])
profit_increase = (now_data - past_data) / past_data
if income_increase > income_limit and profit_increase > profit_limit:
print('营业总收入增长', income_increase)
print('净利润增长', profit_increase)
new_row = tc.SFrame({'code': [row['code']], 'name': [row['name']], 'bankuai': [bankuai],
'close': [row['close']], 'percent_chg': [row['percent_chg']], 'change': [row['change']],
'volume': [row['volume']], 'turn_volume': [row['turn_volume']],
'amplitude': [row['amplitude']], 'volume_rate': [row['volume_rate']], 'turnover_rate': [row['turnover_rate']],
'news_url': [''], 'income_increase': [income_increase],
'profit_increase': [profit_increase],'PERation': [row['PERation']]})
# print(new_row)
# 不好使啊
# xuangu.append(new_row)
# print(SELECTED_DATA)
#xuangu.append(new_row)
#print(xuangu)
result = [income_increase > income_limit and profit_increase > profit_limit, xuangu]
# print(bankuai,row['name'], result) # Debug
return result
def recommendStock(xuangu, SFrame, bankuai):
income_limit = var_list[2]
profit_limit = var_list[3]
counter = 0
while counter < len(SFrame):
result_list = getReport(xuangu, SFrame[counter], bankuai, SFrame[counter], income_limit, profit_limit)
if result_list[0]:
print('股票名称:' + SFrame[counter]['name'])
print('股票代码:' + SFrame[counter]['code'])
print('成交量:' + str(SFrame[counter]['volume']))
print('成交额:' + str(SFrame[counter]['turn_volume']))
print('成交量比增幅:' + str(SFrame[counter]['amplitude']))
print('换手率:' + str(SFrame[counter]['turnover_rate']))
print('-------------------------------------------------')
return result_list[1]
# appendSFrame(selected_data, SFrame[counter]['name'], bankuai, result_list[1], result_list[2])
counter += 1
def user_interface(xuangu):
choice = greetings()
if choice == 'a':
xuangu = inputCode(xuangu)
return xuangu
elif choice == 's':
searchCode()
elif choice == 'l':
print('----------------以下是所有板块代码----------------')
element_list = [(k, codename_dict[k]) for k in sorted(codename_dict.keys())]
counter = 0
while counter < len(element_list):
print(element_list[counter][0] + ' - ' + element_list[counter][1])
counter += 1
print('-----------------------------------------------')
user_interface(SELECTED_DATA)
elif choice == 'x':
print(SELECTED_DATA)
if SELECTED_DATA[0]['name'] != '数据不存在':
SELECTED_DATA.show()
else:
print('没有分析完成的数据!')
user_interface(SELECTED_DATA)
elif choice == 'j':
fileName = input('输入文件名:')
parseSFrame(fileName)
user_interface(SELECTED_DATA)
else:
user_interface(SELECTED_DATA)
return xuangu
def greetings():
print('+-------------------欢迎界面-------------------+')
print('| 请选择您要执行的操作: |')
print('| 1. 输入a来分析指定板块 |')
print('| 2. 输入s来搜寻板块代码 |')
print('| 3. 输入l来显示所有代码 |')
print('| 4. 输入x来显示所有选股 |')
print('| 5. 输入j来加载以前分析完成股票数据 |')
print('| 6. 输入n来加载以前分析完成新闻数据 |')
print('+---------------------------------------------+')
choice = str(input('命令:'))
return choice
def searchCode():
isDone = False
print('+--------------输入想要查找的板块名--------------+')
while isDone == False:
user_input = str(input('搜索板块名:'))
if user_input == 'quit':
print('用户已取消操作!')
break
try:
print(user_input + '代码是:' + name_dict[user_input])
isDone = True
except KeyError:
print('板块不存在!')
user_interface(SELECTED_DATA)
# 分析板块
def inputCode(xuangu):
print('+-------------------分析板块--------------------+')
print('|输入变量: |')
print('-----------------------------------------------')
turnover_rate = input('换手率大于(默认0.05对应5%):')
if turnover_rate == '':
print('换手率设为默认0.05!')
turnover_rate = 0.05
var_list.append(float(turnover_rate))
volume_rate= input('量比大于(默认0.3对应30%):')
if volume_rate == '':
print('量比设为默认0.3!')
volume_rate = 0.3
var_list.append(float(volume_rate))
income_rate = input('营业收入大于(默认0.3对应30%):')
if income_rate == '':
print('营业收入设为默认0.3!')
income_rate = 0.3
var_list.append(float(income_rate))
benefit_rate = input('净利润大于(默认0.3对应30%):')
if benefit_rate == '':
print('净利润设为默认0.3!')
benefit_rate = 0.3
var_list.append(float(benefit_rate))
print('+----------------------------------------------+')
print('|输入板块代号(输入quit退出至主页面, all分析所有板块)|')
print('-----------------------------------------------')
code_name = str(input('代号:'))
if code_name == 'all':
for each_bk in code_dict:
bk_name = codename_dict[each_bk]
xuangu = makeRecommend(xuangu, code_dict[each_bk], bk_name)
elif code_name == 'quit':
user_interface(xuangu)
else:
xuangu = makeRecommend(xuangu, code_dict[code_name], codename_dict[code_name])
saveSFrame(xuangu, fileRoot)
# showSFrame(xuangu)
return xuangu
# 目前很简陋
def showSFrame(selected_data):
print(selected_data)
# 用于加载SFrame
def parseSFrame(fileName):
filePath = './SelectedData/' + fileName + '/'
SELECTED_DATA = tc.SFrame(data=filePath)
# 用于保存SFrame
def saveSFrame(SFrame, fileRoot):
# 保存数据
date = '20' + str(datetime.datetime.now().strftime("%y%m%d-%H%M"))
filepath = fileRoot + '/' + str(date) + '/'
SFrame.save(filepath)
print('成功保存数据文件!数据路径:' + filepath)
# 打印时间戳
print('程序运行时间戳:20'
+ str(datetime.datetime.now().strftime("%y")) + '年'
+ str(datetime.datetime.now().strftime("%m")) + '月'
+ str(datetime.datetime.now().strftime("%d")) + '日'
+ str(datetime.datetime.now().strftime("%H")) + '时'
+ str(datetime.datetime.now().strftime("%M")) + '分'
+ str(datetime.datetime.now().strftime("%S")) + '秒')
# 用于移除第一个占位符
def removeFront(SFrame):
return SFrame[1:len(SFrame)]
# 推荐股票
def makeRecommend(xuangu, url, bk_name):
# 创建四个空SFrame,以占位行开头
all_data = tc.SFrame({'code': ['000000'], 'name': ['哔哩哔哩'],
'close': [0.0], 'percent_chg': [0.0],
'change': [0.0], 'volume': [0.0], 'turn_volume': [0.0], 'amplitude': [0.0],
'high': [0.0], 'low': [0.0],
'now_open': [0.0], 'previous_close': [0.0], 'volume_rate': [0.0],
'turnover_rate': [0.0], 'report_url': ['http://www.bilibili.com'], 'PERation': [0.0]})
# 获取信息
all_data = makeData(url, all_data)
# 初步筛选
analyze_data = analyze_stock(all_data)
# 最终推荐
print('---------------------' + bk_name + '---------------------')
return recommendStock(xuangu, analyze_data, bk_name)
# selected_data = removeFront(selected_data)
# user_interface(SELECTED_DATA)
# return selected_data
# +
# ===================================主执行部分===================================
# UI
SELECTED_DATA = user_interface(SELECTED_DATA)
# -
# # NEW
# +
# 功能性函数
# searchCode() 用于搜寻股票代码
def searchCode():
isDone = False
print('+--------------输入想要查找的板块名--------------+')
while isDone == False:
user_input = str(input('搜索板块名:'))
if user_input == 'quit':
print('用户已取消操作!')
break
try:
print(user_input + '代码是:' + name_dict[user_input])
isDone = True
except KeyError:
print('板块不存在!')
# 重启UI (待议)
user_interface(SELECTED_DATA)
# printAllBankuai()
def printAllBankuai():
print('----------------以下是所有板块代码----------------')
element_list = [(k, codename_dict[k]) for k in sorted(codename_dict.keys())]
counter = 0
while counter < len(element_list):
print(element_list[counter][0] + ' - ' + element_list[counter][1])
counter += 1
print('-----------------------------------------------')
# 重启UI
user_interface(SELECTED_DATA)
def showAllSelected():
print(SELECTED_DATA)
if SELECTED_DATA[0]['name'] != '数据不存在':
SELECTED_DATA.show()
else:
print('没有分析完成的数据!')
user_interface(SELECTED_DATA)
# 用于加载SFrame (也许需要写一个return 待议)
def parseSFrame():
fileName = input('输入文件名:')
filePath = './SelectedData/' + fileName + '/'
SELECTED_DATA = tc.SFrame(data=filePath)
user_interface(SELECTED_DATA)
# +
def user_interface(xuangu):
choice = greetings()
if choice == 'a':
xuangu = inputCode(xuangu)
return xuangu
elif choice == 's':
searchCode()
elif choice == 'l':
printAllBankuai()
elif choice == 'x':
showAllSelected()
elif choice == 'j':
parseSFrame()
else:
user_interface(SELECTED_DATA)
return xuangu
| Recommender.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: ipykernel_py2
# ---
import numpy as np
from pandas_datareader import data as wb
import matplotlib.pyplot as plt
PG = wb.DataReader('PG', data_source='morningstar', start='2002-1-1')
PG.head()
PG.tail()
# ## Simple Rate of Return
# $$
# \frac{P_1 - P_0}{P_0} = \frac{P_1}{P_0} - 1
# $$
PG['simple_return'] = (PG['Close'] / PG['Close'].shift(1)) - 1
print PG['simple_return']
| Python for Finance - Code Files/63 Simple Returns - Part I/Online Financial Data (APIs)/Python 2 APIs/Simple Returns - Part I - Lecture_Morn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Universal Geocoder Sample
#
# ### For simple profiling of the geocoder
import universal_geocoder as ugc
import pandas as pd
# This was some lat/long taken from geocoder file
output = ugc.geocode(47.534185, -122.371273)
# First few entries for check at location
output[:6]
# Time a single call of the geocoder
# %timeit ugc.geocode(47.534185, -122.371273)
# %load_ext line_profiler
# Profile the call to the geocoder
# %lprun -T out.txt -f ugc.geocode ugc.geocode(47.534185, -122.371273)
# !cat out.txt
| notebooks/profile_geocoder.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import openpathsampling as paths
# +
# always the most complicated bit
import openpathsampling.engines.toy as toys
import numpy as np
pes = (toys.OuterWalls([1.0,1.0], [0.0,0.0]) +
toys.Gaussian(-0.7, [12.0, 0.5], [-0.5, 0.0]) +
toys.Gaussian(-0.7, [12.0, 0.5], [0.5, 0.0]))
topology = toys.Topology(n_spatial=2, masses=[1.0, 1.0], pes=pes)
template = toys.Snapshot(coordinates=np.array([[0.0, 0.0]]),
velocities=np.array([[0.0, 0.0]]),
topology=topology)
engine = toys.Engine(options={'integ': toys.LangevinBAOABIntegrator(dt=0.02, temperature=0.1, gamma=2.5),
'n_frames_max': 5000,
'n_steps_per_frame': 10},
template=template)
# +
# states are volumes in a CV space: define the CV
def xval(snapshot):
return snapshot.xyz[0][0]
cv = paths.FunctionCV("xval", xval)
# -
stateA = paths.CVDefinedVolume(cv, float("-inf"), -0.2).named("A")
stateB = paths.CVDefinedVolume(cv, 0.2, float("inf")).named("B")
network = paths.FixedLengthTPSNetwork(stateA, stateB, length=5000)
scheme = paths.OneWayShootingMoveScheme(network, engine=engine)
# I'll fake an initial trajectory
delta = 2.0 / 5000
minimum = -1.0 - delta
trajectory = paths.Trajectory([
toys.Snapshot(coordinates=np.array([[minimum+k*delta, 0.0]]),
velocities=np.array([[0.1, 0.0]]),
topology=topology)
for k in range(5000)
])
initial_conditions = paths.SampleSet.map_trajectory_to_ensembles(trajectory, network.sampling_ensembles)
storage = paths.Storage("simple_fixedlen.nc", "w", template=template)
simulation = paths.PathSampling(storage, scheme, initial_conditions)
simulation.run(200)
| examples/simple_tps/simple_fixed_length_tps.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] tags=["include"]
# # POLSCI 3 Fall 2020
#
# ## Notebook for Week 3 Lecture video: Variables, Expected Utility, Plotting
#
# Estimated time: 20 minutes
# -
# Lets do some of the calculations for expected utility from the week 3 lectures.
#
# To get warmed up, we can just use Python as a calculator.
#
# Expected utility of a coin flip where H means losing 2 and T means winning 1
.5*(-2) + .5*1
# Expected utility of a coin flip where H means losing 2 and T means winning 5
.5*(-2) + .5*5
# In problem sets, we will often ask you to fill in or edit a code cell. We will give instructions for what you need to do to earn points in <font color="blue"> <b> bold blue font </b> </font>, and put additional pointers for what to do in the comments.
#
# <font color="blue"> <b> Put a new line of code in the cell below which computes the requested expected utility </font> </b>
# +
# Expected utility of a coin flip where H means losing 3 and T means winning 4
# -
# We can also move towards more general analysis by treating these as variables
#Creating variables for the value of H and T flips
uh = -2
ut = 1
.5*uh + .5*ut
# Or, we can be even more general and let the probability that the coin flip is H be a variable p
# Creating variables for all of the EU calculation
uh=-2
ut=1
p=.5
p*uh + (1-p)*ut
# Now remember the protest example, where the utility for getting arrested is -2 and for a fun protest is 1
#
# <font color="blue"> <b> Fill in the value of p which will make the expected utility 0 </font> </b>
# +
# meaning indifference between protesting and staying home
ua = -2
uf = 1
p = ...
p*ua + (1-p)*uf
# -
# One advantage to using variables is that we can also plot the utility of protest as a function of p, which will tell us when it is above zero (meaning protest is the better choice) and below zero (meaning staying home is the better choice)
#
# Don't worry about understanding all of the code for this one!
# Plotting the utility of protest as a function of p with a blue line
# The vertical line indicates where the utility crosses zero
import numpy as np
import matplotlib.pyplot as plt
p = np.arange(0,1, step=.01)
uprotest = -2*p + 1*(1-p)
plt.plot(p,uprotest)
plt.hlines(0,0,1)
plt.vlines(1/3,-2,1)
# Now let's do the same thing, but make getting arrested even worse. To do this, we change the line of code which computes the expected utility to assign a value of -5 to getting arrested rather than -2
# Plotting the utility of protest as a function of p with a blue line
# The vertical line indicates where the utility crosses zero
import numpy as np
import matplotlib.pyplot as plt
p = np.arange(0,1, step=.01)
#This is the line of code we changed!
uprotest = -5*p + 1*(1-p)
plt.plot(p,uprotest)
plt.hlines(0,0,1)
plt.vlines(1/6,-5,1)
# On some problem sets we will ask you questions about interpretation, and tell you to fill them in the cell below.
#
# <font color="blue"> <b> What does this graph tell you about how the probability and fear of getting arrested affects protest decisions? </font> </b>
# YOUR ANSWER HERE
| PS 3 2020 - Discussion Notebook 1, Introduction to Python, Expected Utility.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
# # one_to_one_joins
df1=pd.DataFrame({'employee':['Raju','Rani','Ramesh','Ram'],
'group':['Accounting','Engineering','Engineering','HR']})
df2=pd.DataFrame({'employee':['Raju','Rani','Ramesh','Ram'],
'hire_date':[2004,2008,2012,2014]})
df1
df2
df3=pd.merge(df1,df2)
df3
df4=pd.concat([df1,df2],axis=0)
df4
# # Many_to_one_joins
df5=pd.DataFrame({'group':['Accounting','Engineering','HR'],
'supervisor':['Carly','Guido','Steve']})
df5
# # many_to_many_joins
df6=pd.DataFrame({'group':['Accounting','Accounting','Engineering','Engineering','HR','HR'],
'skills':['math','spreadsheets','coading','linux','spreadsheet','organization']})
df6
pd.merge(df1,df2,on='employee')
data1=pd.DataFrame({'name':['mani','kanta','jyothi','asha','durga'],
'DOB':[1994,1994,1995,1992,1996],'age':[27,27,26,26,25]})
data2=pd.DataFrame({'name':['mani','kanta','jyothi','asha','durga'],
'Gender':['M','M','F','F','F'],'age':[27,27,26,26,25]})
data=pd.merge(data1,data2)
data
da=pd.merge(data1,data2,on=['name','age'])
da
# # The left_on and right_on keywords
data1=pd.DataFrame({'employee':['mani','kanta','jyothi','asha','durga'],
'DOB':[1994,1994,1995,1992,1996],'age':[27,27,26,26,25]})
data2=pd.DataFrame({'name':['mani','kanta','jyothi','asha','durga'],
'Gender':['M','M','F','F','F'],'rank':[27,27,26,26,25]})
x=pd.merge(data1,data2, left_on='employee',right_on='name')
x
x1=pd.merge(data1,data2, left_on='employee',right_on='name').drop('name',axis=1)
x1
# # left_index and right_index
df1a=df1.set_index('employee')
df2a=df2.set_index('employee')
pd.merge(df1a,df2a,left_index=True,right_index=True)
# # join
df1a.join(df2a)
# # set arthmetic for joins
df6=pd.DataFrame({'name':['peter','paul','mary'],
'food':['fish','beans','bread']},columns=['name','food'])
df7=pd.DataFrame({'name':['mary','joseph','paul'],
'drink':['win','beer','water']},columns=['name','drink'])
pd.merge(df6,df7)
pd.merge(df6,df7,how='inner')
pd.merge(df6,df7,how='outer')
pd.merge(df6,df7,how='left')
pd.merge(df6,df7,how='right')
# # overlapping columns names:the suffixes keyword
df8=pd.DataFrame({'name':['Bob','Jake','Lisa','Sue'],'rank':[1,2,3,4]})
df9=pd.DataFrame({'name':['Bob','Jake','Lisa','Sue'],'rank':[3,1,4,2]})
pd.merge(df8,df9,on='name')
pd.merge(df8,df9,on='name',suffixes=['_L','_R'])
| merge_and_joins.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: py3bI
# language: python
# name: py3bi
# ---
# # 6 - Data Compression
# **This short Notebook will introduce you to how to efficiently compress your data within SampleData datasets.**
#
# <div class="alert alert-info">
#
# **Note**
#
# Throughout this notebook, it will be assumed that the reader is familiar with the overview of the SampleData file format and data model presented in the [first notebook of this User Guide](./SampleData_Introduction.ipynb) of this User Guide.
#
# </div>
# ## Data compression with HDF5 and Pytables
# ### HDF5 and data compression
# HDF5 allows compression filters to be applied to datasets in a file to minimize the amount of space it consumes. These compression features allow to drastically improve the storage space required for you datasets, as well as the speed of I/O access to datasets, and can differ from one data item to another within the same HDF5 file. A detailed presentation of HDF5 compression possibilities is provided [here](https://support.hdfgroup.org/HDF5/faq/compression.html).
#
# The two main ingredients that control compression performances for HDF5 datasets are the compression filters used (which compression algorithm, with which parameters), and the using a chunked layout for the data. This two features are briefly developed hereafter.
# ### Pytables and data compression
# The application of compression filters to HDF5 files with the *SampleData* class is handled by the *Pytable* package, on which is built the *SampleData* HDF5 interface. *Pytables* implements a specific containers class, the `Filters` class, to gather the various settings of the compression filters to apply to the datasets in a HDF5 file.
#
# When using the *SampleData* class, you will have to specify this compression filter settings to class methods dedicated to data compression. These settings are the parameters of the *Pytables* `Filters` class. These settings and their possible values are detailed in the next subsection.
# #### Available filter settings
# The description given below of compression options available with *SampleData*/*Pytables* is exctracted from [the *Pytables* documentation of the *Filter* class](https://www.pytables.org/usersguide/libref/helper_classes.html#the-filters-class).
# * **complevel** (int) – Specifies a compression level for data. The allowed range is 0-9. A value of 0 (the default) disables compression.
#
# * **complib** (str) – Specifies the compression library to be used. Right now, `zlib` (the default), `lzo`, `bzip2` and `blosc` are supported. Additional compressors for Blosc like `blosc:blosclz` (‘blosclz’ is the default in case the additional compressor is not specified), `blosc:lz4`, `blosc:lz4hc`, `blosc:snappy`, `blosc:zlib` and `blosc:zstd` are supported too. Specifying a compression library which is not available in the system issues a FiltersWarning and sets the library to the default one.
#
# * **shuffle** (bool) – Whether or not to use the Shuffle filter in the HDF5 library. This is normally used to improve the compression ratio. A false value disables shuffling and a true one enables it. The default value depends on whether compression is enabled or not; if compression is enabled, shuffling defaults to be enabled, else shuffling is disabled. Shuffling can only be used when compression is enabled.
#
# * **bitshuffle** (bool) – Whether or not to use the BitShuffle filter in the Blosc library. This is normally used to improve the compression ratio. A false value disables bitshuffling and a true one enables it. The default value is disabled.
#
# * **fletcher32** (bool) – Whether or not to use the Fletcher32 filter in the HDF5 library. This is used to add a checksum on each data chunk. A false value (the default) disables the checksum.
#
# * **least_significant_digit** (int) – If specified, data will be truncated (quantized). In conjunction with enabling compression, this produces ‘lossy’, but significantly more efficient compression. For example, if least_significant_digit=1, data will be quantized using around(scale*data)/scale, where scale = 2^bits, and bits is determined so that a precision of 0.1 is retained (in this case bits=4). Default is None, or no quantization.
# ### Chunked storage layout
# Compressed data is stored in a data array of an HDF5 dataset using a chunked storage mechanism. When chunked storage is used, the data array is split into equally sized chunks each of which is stored separately in the file, as illustred on the diagram below. Compression is applied to each individual chunk. When an I/O operation is performed on a subset of the data array, only chunks that include data from the subset participate in I/O and need to be uncompressed or compressed.
#
# Chunking data allows to:
#
# * Generally improve, sometimes drastically, the I/O performance of datasets. This comes from the fact that the chunked layout removes the reading speed anisotropy for data array that depends along which dimension its elements are read (*i.e* the same number of disk access are required when reading data in rows or columns).
# * Chunked storage also enables adding more data to a dataset without rewriting the whole dataset.
#
# <img src="./Images/Tutorial_5/chuncked_layout.png" width="50%">
#
# **By default, data arrays are stored with a chunked layout in SampleData datasets.** The size of chunks is the key parameter that controls the impact on I/O performances for chunked datasets. **The shape of chunks is computed automatically by the Pytable package, providing a value yielding generally good I/O performances.** if you need to go further in the I/O optimization, you may consult the *Pytables* [documentation page](https://www.pytables.org/usersguide/optimization.html) dedicated to compression optimization for I/O speed and storage space. In addition, it is highly recommended to read [this document](https://support.hdfgroup.org/HDF5/doc/TechNotes/TechNote-HDF5-ImprovingIOPerformanceCompressedDatasets.pdf) in order to be able to efficiently optimize I/O and storage performances for your chunked datasets. These performance issues will not be discussed in this tutorial.
# ## Compressing your datasets with SampleData
# Within *Sampledata*, data compression can be applied to:
#
# * data arrays
# * structured data arrays
# * field data arrays
#
# There are two ways to control the compression settings of your *SampleData* data arrays:
#
# 1. Providing compression settings to data item creation methods
# 2. Using the `set_chunkshape_and_compression` and `set_nodes_compression_chunkshape` methods
# ### The compression options dictionary
# In both cases, you will have to pass the various settings of the compression filter you want to apply to your data to the appropriate *SampleData* method. All of these methods accept for that purpose a `compression_options` argument, which must be a dictionary. Its keys and associated values can be chosen among the ones listed in the **Available filter settings** subsection above.
# ### Compress already existing data arrays
# We will start by looking at how we can change compression settings of already existing data in *SampleData* datasets.
# For that, we will use a material science dataset that is part of the *Pymicro* example datasets.
from config import PYMICRO_EXAMPLES_DATA_DIR # import file directory path
import os
dataset_file = os.path.join(PYMICRO_EXAMPLES_DATA_DIR, 'example_microstructure') # test dataset file path
tar_file = os.path.join(PYMICRO_EXAMPLES_DATA_DIR, 'example_microstructure.tar.gz') # dataset archive path
# This file is zipped in the package to reduce its size. We will have to unzip it to use it and learn how to reduce its size with the *SampleData* methods. If you are just reading the documentation and not executing it, you may just skip this cell and the next one.
# Save current directory
cwd = os.getcwd()
# move to example data directory
os.chdir(PYMICRO_EXAMPLES_DATA_DIR)
# unarchive the dataset
os.system(f'tar -xvf {tar_file}')
# get back to UserGuide directory
os.chdir(cwd)
# #### Dataset presentation
# In this tutorial, we will work on a copy of this dataset, to leave the original data unaltered.
# We will start by creating an autodeleting copy of the file, and print its content to discover its content.
# import SampleData class
from pymicro.core.samples import SampleData as SD
# import Numpy
import numpy as np
# Create a copy of the existing dataset
data = SD.copy_sample(src_sample_file=dataset_file, dst_sample_file='Test_compression', autodelete=True,
get_object=True, overwrite=True)
print(data)
data.get_file_disk_size()
# As you can see, this dataset already contains a rich content. It is a digital twin of a real polycristalline microstructure of a grade 2 Titanium sample, gathering both experimental and numerical data obtained through Diffraction Contrast Tomography imaging, and FFT-based mechanical simulation.
#
# This dataset has actually been constructed using the `Microstructure` class of the pymicro package, which is based on the *SampleData* class. The link between these classes will be discussed in the next tutorial.
# This dataset contains only uncompressed data. We will try to reduce its size by using various compression methods on the large data items that it contains. You can see that most of them are stored in the *3DImage Group* `CellData`.
# ### Apply compression settings for a specific array
# We will start by compressing the `grain_map` *Field data array* of the `CellData` image. Let us look more closely on this data item:
data.print_node_info('grain_map')
# We can see above that this data item is not compressed (`complevel=0`), and has a disk size of almost 2 Mb.
#
# To apply a set of compression settings to this data item, you need to:
#
# 1. create a dictionary specifying the compression settings:
compression_options = {'complib':'zlib', 'complevel':1}
# 2. use the *SampleData* `set_chunkshape_and_compression` method with the dictionary and the name of the data item as arguments
data.set_chunkshape_and_compression(nodename='grain_map', compression_options=compression_options)
data.get_node_disk_size('grain_map')
data.print_node_compression_info('grain_map')
# As you can see, the storage size of the data item has been greatly reduced, by more than 10 times (126 Kb vs 1.945 Mb), using this compression settings. Let us see what will change if we use different settings :
# +
# No `shuffle` option:
print('\nUsing the shuffle option, with the zlib compressor and a compression level of 1:')
compression_options = {'complib':'zlib', 'complevel':1, 'shuffle':True}
data.set_chunkshape_and_compression(nodename='grain_map', compression_options=compression_options)
data.get_node_disk_size('grain_map')
# No `shuffle` option:
print('\nUsing no shuffle option, with the zlib compressor and a compression level of 9:')
compression_options = {'complib':'zlib', 'complevel':9, 'shuffle':False}
data.set_chunkshape_and_compression(nodename='grain_map', compression_options=compression_options)
data.get_node_disk_size('grain_map')
# No `shuffle` option:
print('\nUsing the shuffle option, with the lzo compressor and a compression level of 1:')
compression_options = {'complib':'lzo', 'complevel':1, 'shuffle':True}
data.set_chunkshape_and_compression(nodename='grain_map', compression_options=compression_options)
data.get_node_disk_size('grain_map')
# No `shuffle` option:
print('\nUsing no shuffle option, with the lzo compressor and a compression level of 1:')
compression_options = {'complib':'lzo', 'complevel':1, 'shuffle':False}
data.set_chunkshape_and_compression(nodename='grain_map', compression_options=compression_options)
data.get_node_disk_size('grain_map')
# -
# As you may observe, is significantly affected by the choice of the compression level. The higher the compression level, the higher the compression ratio, but also the lower the I/O speed. On the other hand, you can also remark that, in the present case, using the `shuffle` filter deteriorates the compression ratio.
#
# Let us try to with another data item:
# +
data.print_node_info('Amitex_stress_1')
# No `shuffle` option:
print('\nUsing the shuffle option, with the zlib compressor and a compression level of 1:')
compression_options = {'complib':'zlib', 'complevel':1, 'shuffle':True}
data.set_chunkshape_and_compression(nodename='Amitex_stress_1', compression_options=compression_options)
data.get_node_disk_size('Amitex_stress_1')
# No `shuffle` option:
print('\nUsing no shuffle option, with the zlib compressor and a compression level of 1:')
compression_options = {'complib':'zlib', 'complevel':1, 'shuffle':False}
data.set_chunkshape_and_compression(nodename='Amitex_stress_1', compression_options=compression_options)
data.get_node_disk_size('Amitex_stress_1')
# -
# On the opposite, for this second array, the shuffle filter improves significantly the compression ratio. However, in this case, you can see that the compression ratio achieved is much lower than for the `grain_map` array.
# <div class="alert alert-warning">
#
# **Warning 1**
#
# The efficiency of compression algorithms in terms of compression ratio is strongly affected by the data itself (variety, value and position of the stored values in the array). Compression filters will not have the same behavior with all data arrays, as you have observed just above. Be aware of this fact, and do not hesitate to conduct tests to find the best settings for you datasets !
#
# </div>
# <div class="alert alert-warning">
#
# **Warning 2**
#
# Whenever you change the compression or chunkshape settings of your datasets, the data item is re-created into the *SampleData* dataset, which may be costly in computational time. Be careful if you are dealing with very large data arrays and want to try out several settings to find the best I/O speed / compression ratio compromise, with the `set_chunkshape_and_compression` method. You may want to try on a subset of your large array to speed up the process.
# </div>
# ### Apply same compression settings for a serie of nodes
# If you need to apply the same compression settings to a list of data items, you may use the `set_nodes_compression_chunkshape`. This method works exactly like `set_chunkshape_and_compression`, but take a list of nodenames as arguments instead of just one. The inputted compression settings are then applied to all the nodes in the list:
# +
# Print current size of disks and their compression settings
data.get_node_disk_size('grain_map_raw')
data.print_node_compression_info('grain_map_raw')
data.get_node_disk_size('uncertainty_map')
data.print_node_compression_info('uncertainty_map')
data.get_node_disk_size('mask')
data.print_node_compression_info('mask')
# Compress datasets
compression_options = {'complib':'zlib', 'complevel':9, 'shuffle':True}
data.set_nodes_compression_chunkshape(node_list=['grain_map_raw', 'uncertainty_map', 'mask'],
compression_options=compression_options)
# Print new size of disks and their compression settings
data.get_node_disk_size('grain_map_raw')
data.print_node_compression_info('grain_map_raw')
data.get_node_disk_size('uncertainty_map')
data.print_node_compression_info('uncertainty_map')
data.get_node_disk_size('mask')
data.print_node_compression_info('mask')
# -
# ### Lossy compression and data normalization
# The compression filters used above preserve exactly the original values of the stored data. However, it is also possible with specific filters **a lossy compression**, which remove non relevant part of the data. As a result, data compression ratio is usually strongly increased, at the cost that stored data is no longer exactly equal to the inputed data array.
# One of the most important feature of data array that increase their compressibility, is the presence of patterns in the data. If a value or a serie of values is repeated multiple times throughout the data array, data compression can be very efficient (the pattern can be stored only once).
#
# Numerical simulation and measurement tools usually output data in a standard simple or double precision floating point numbers, yiedling data arrays with values that have a lot of digits. Typically, these values are all different, and hence these arrays cannot be efficiently compressed.
# The `Amitex_stress_1` and `Amitex_strain_1` data array are two tensor fields outputed by a continuum mechanics FFT-based solver, and typical fall into this category: they have almost no equal value or clear data pattern.
#
# As you can see above, the best achieved compression ratio is 60% while for the dataset `grain_map`, the compression is way more efficient, with a best ratio that climbs up to 97% (62 Kb with *zlib* compressor and compression level of 9, versus an initial data array of 1.945 Mb). This is due to the nature of the `grain_map` data array, which is a tridimensional map of grains identification number in microstructure of the Titanium sample represented by the dataset. It is hence an array containing a few integer values that are repeated many times.
#
# Let us analyze these two data arrays values to illustrate this difference:
import numpy as np
print(f"Data array `grain_map` has {data['grain_map'].size} elements,"
f"and {np.unique(data['grain_map']).size} different values.\n")
print(f"Data array `Amitex_stress_1` has {data['Amitex_stress_1'].size} elements,"
f"and {np.unique(data['Amitex_stress_1']).size} different values.\n")
# #### Lossy compression
# Usually, the relevant precision of data is only of a few digits, so that many values of the array should be considered equal. The idea of lossy compression is to truncate values up to a desired precision, which increases the number of equal values in a dataset and hence increases its compressibility.
# Lossy compression can be applied to floating point data arrays in *SampleData* datasets using the `least_significant_digit` compression setting. If you set the value of this option to $N$, the data will be truncated after the $N^{th}$ siginificant digit after the decimal point. Let us see an example.
# +
# We will store a value of an array to verify how it evolves after compression
original_value = data['Amitex_stress_1'][20,20,20]
# Apply lossy compression
data.get_node_disk_size('Amitex_stress_1')
# Set up compression settings with lossy compression: truncate after third digit adter decimal point
compression_options = {'complib':'zlib', 'complevel':9, 'shuffle':True, 'least_significant_digit':3}
data.set_chunkshape_and_compression(nodename='Amitex_stress_1', compression_options=compression_options)
data.get_node_disk_size('Amitex_stress_1')
# Get same value after lossy compression
new_value = data['Amitex_stress_1'][20,20,20]
print(f'Original array value: {original_value} \n'
f'Array value after lossy compression: {new_value}')
# -
# As you may observe, the compression ratio has been improved, and the retrieved values after lossy compression are effectively equal to the original array up to the third digit after the decimal point.
#
# We will now try to increase the compression ratio by reducing the number of conserved digits to 2:
# +
# Set up compression settings with lossy compression: truncate after third digit adter decimal point
compression_options = {'complib':'zlib', 'complevel':9, 'shuffle':True, 'least_significant_digit':2}
data.set_chunkshape_and_compression(nodename='Amitex_stress_1', compression_options=compression_options)
data.get_node_disk_size('Amitex_stress_1')
# Get same value after lossy compression
new_value = data['Amitex_stress_1'][20,20,20]
print(f'Original array value: {original_value} \n'
f'Array value after lossy compression 2 digits: {new_value}')
# -
# As you can see, the compression ratio has again been improved, now close to 75%. Know, you know how to do to choose the best compromise between lost precision and compression ratio.
# #### Normalization to improve compression ratio
# If you look more closely to the `Amitex_stress_1` array values, you can observe that the value of this array have been outputed within a certain scale of values, which in particular impact the number of significant digits that come before the decimal point. Sometimes precision of the data would require less significant digits than its scale of representation.
#
# In that case, storing the complete data array at its original scale is not necessary, and very inefficient in terms of data size. To optimize storage of such datasets, one can normalize them to a form with very few digits before the decimal point (1 or 2), and stored separately their scale to be able to revert the normalization operation when retrieiving data.
#
# This allows to reduce the total number of significant digits of the data, and hence further improve the achievable compression ratio with lossy compression.
# The *SampleData* class allows you to aplly automatically this operation when applying compression settings to your dataset. All you have to do is add to the `compression_option` dictionary the key `normalization` with one of its possible values.
#
# To try it, we will close (and delete) our test dataset and recopy the original file, to apply normalization and lossy compression on the original raw data:
# removing dataset to recreate a copy
del data
# creating a copy of the dataset to try out lossy compression methods
data = SD.copy_sample(src_sample_file=dataset_file, dst_sample_file='Test_compression', autodelete=True,
get_object=True, overwrite=True)
# ##### Standard Normalization
# The `standard` normalization setting will center and reduce the data of an array $X$ by storing a new array $Y$ that is:
#
# $Y = \frac{X - \bar{X}}{\sigma(X)}$
#
# where $\bar{X}$ and $\sigma(X)$ are respectively the mean and the standard deviation of the data array $X$.
#
# This operation reduces the number of significant digits before the decimal point to 1 or 2 for the large majority of the data array values. After *standard normalization*, lossy compression will yield much higher compression ratios for data array that have a non normalized scale.
#
# The SampleData class ensures that when data array are retrieved, or visualized, the user gets or sees the original data, with the normalization reverted.
#
# Let us try to apply it to our stress field `Amitex_stress_1`.
# +
# Set up compression settings with lossy compression: truncate after third digit adter decimal point
compression_options = {'complib':'zlib', 'complevel':9, 'shuffle':True, 'least_significant_digit':2,
'normalization':'standard'}
data.set_chunkshape_and_compression(nodename='Amitex_stress_1', compression_options=compression_options)
data.get_node_disk_size('Amitex_stress_1')
# Get same value after lossy compression
new_value = data['Amitex_stress_1'][20,20,20,:]
# Get in memory value of the node
memory_value = data.get_node('Amitex_stress_1', as_numpy=False)[20,20,20,:]
print(f'Original array value: {original_value} \n'
f'Array value after normalization and lossy compression 2 digits: {new_value}',
f'Value in memory: {memory_value}')
# -
# As you can see, the compression ratio has been strongly improved by this normalization operation, reaching 90%.
# When looking at the retrieved value after compression, you can see that depending on the field component that is observed, the relative precision loss varies. The third large component value error is less than 1%, which is consistent with the truncation to 2 significant digits. However, it is not the other components, that have smaller values by two or three orders of magnitude, and that are retrieved with larger errors.
#
# This is explained by the fact that the `standard` normalization option scales the array as a whole. As a result, if there are large differencies in the scale of different components of a vector or tensor field, the precision of the smaller components will be less preserved.
# ##### Standard Normalization per components for vector/tensor fields
# Another normalization option is available for *SampleData* field arrays, that allows to apply standard normalization individually to each component of a field in order to keep a constant relative precision for each component when applying lossy compression to the field data array.
#
# To use this option, you will need to set the `normalization` value to `standard_per_component`:
del data
data = SD.copy_sample(src_sample_file=dataset_file, dst_sample_file='Test_compression', autodelete=True,
get_object=True, overwrite=True)
# +
# Set up compression settings with lossy compression: truncate after third digit adter decimal point
compression_options = {'complib':'zlib', 'complevel':9, 'shuffle':True, 'least_significant_digit':2,
'normalization':'standard_per_component'}
data.set_chunkshape_and_compression(nodename='Amitex_stress_1', compression_options=compression_options)
data.get_node_disk_size('Amitex_stress_1')
# Get same value after lossy compression
new_value = data['Amitex_stress_1'][20,20,20,:]
# Get in memory value of the node
memory_value = data.get_node('Amitex_stress_1', as_numpy=False)[20,20,20,:]
print(f'Original array value: {original_value} \n'
f'Array value after normalization per component and lossy compression 2 digits: {new_value}\n',
f'Value in memory: {memory_value}')
# -
# As you can see, the error in the retrieved array is now less than 1% for each component of the field value. However, the cost was a reduced improvement of the compression ratio.
# ##### Visualization of normalized data
data.print_xdmf()
# As you can see, the `Amitex_stress_1` *Attribute* node data in the dataset XDMF file is now provided by a `Function` item type, involving three data array with the original field shape. This function computes:
#
# $X' = Y*\sigma(X) + \bar{X}$
#
# where $*$ and $+$ are element-wise product and addition operators for multidimensional arrays. This operation allows to revert the component-wise normalization of data. The Paraview software is able to interpret this syntax of the XDMF format and hence, when visualizing data, you will see the values with the original scaling.
# This operation required the creation of two large arrays in the dataset, that the store the mean and standard deviation of each component of the field, repeted for each spatial dimensions of the field data array. It is mandatory to allow visualization of the data with the right scaling in Paraview. However, as these array contain a very low amount of data ($2*N_c$: two times de number of components of the field), they can be very easily compressed and hence do not significantly affect the storage size of the data item, as you may see below:
data.get_node_disk_size('Amitex_stress_1')
data.get_node_disk_size('Amitex_stress_1_norm_std')
data.get_node_disk_size('Amitex_stress_1_norm_mean')
# ### Changing the chunksize of a node
# #### Compressing all fields when adding Image or Mesh Groups
# Changing the chunksize of a data array with *SampleData* is very simple. You just have to pass as a *tuple* the news shape of the chunks you want for your data array, and pass it as an argument to the `set_chunkshape_and_compression` or `set_nodes_compression_chunkshape`:
data.print_node_compression_info('Amitex_stress_1')
data.get_node_disk_size('Amitex_stress_1')
# Change chunkshape of the array
compression_options = {'complib':'zlib', 'complevel':9, 'shuffle':True, 'least_significant_digit':2,
'normalization':'standard_per_component'}
data.set_chunkshape_and_compression(nodename='Amitex_stress_1', chunkshape=(10,10,10,6),
compression_options=compression_options)
data.get_node_disk_size('Amitex_stress_1')
data.print_node_compression_info('Amitex_stress_1')
# As you can see, the chunkshape has been changed, which has also affected the memory size of the compressed data array. We have indeed reduced the number of chunks in the dataset, which reduces the number of data to store. This modification can also improve or deteriorate the I/O speed of access to your data array in the dataset. The reader is once again refered to dedicated documents to know more ion this matter: [here](https://support.hdfgroup.org/HDF5/doc/TechNotes/TechNote-HDF5-ImprovingIOPerformanceCompressedDatasets.pdf) and [here](https://www.pytables.org/usersguide/optimization.html).
# ### Compression data and setting chunkshape upon creation of data items
# Until here we have only modified the compression settings of already existing data items. In this process, the data items are replaced by the new compressed version of the data, which is a costly operation. For this reason, if they are known in advance, it is best to apply the compression filters and appropriate chunkshape when creating the data item.
#
# If you have read through all the tutorials of this user guide, you should know all the method that allow to create data items in your datasets, like `add_data_array`, `add_field`, `ædd_mesh`... All of these methods accept the two arguments `chunkshape` and `compression_options`, that work exaclty as for the `set_chunkshape_and_compression` or `set_nodes_compression_chunkshape` methods. You hence use them to create your data items directly with the appropriate compression settings.
#
# Let us see an example. We will get an array from our dataset, and try to recreate it with a new name and some data compression:
# removing dataset to recreate a copy
del data
# creating a copy of the dataset to try out lossy compression methods
data = SD.copy_sample(src_sample_file=dataset_file, dst_sample_file='Test_compression', autodelete=True,
get_object=True, overwrite=True)
# getting the `orientation_map` array
array = data['Amitex_stress_1']
# +
# create a new field for the CellData image group with the `orientation_map` array and add compression settings
compression_options = {'complib':'zlib', 'complevel':1, 'shuffle':True, 'least_significant_digit':2,
'normalization':'standard'}
new_cshape = (10,10,10,3)
# Add data array as field of the CellData Image Group
data.add_field(gridname='CellData', fieldname='test_compression', indexname='testC', array=array,
chunkshape=new_cshape, compression_options=compression_options, replace=True)
# -
# Check size and settings of new field
data.print_node_info('testC')
data.get_node_disk_size('testC')
data.print_node_compression_info('testC')
# The node has been created with the desired chunkshape and compression filters.
# ### Repacking files
# We now recreate a new copy of the original dataset, and try to reduce the size of oll heavy data item, to reduce as much as possible the size of our dataset.
# removing dataset to recreate a copy
del data
# creating a copy of the dataset to try out lossy compression methods
data = SD.copy_sample(src_sample_file=dataset_file, dst_sample_file='Test_compression', autodelete=True,
get_object=True, overwrite=True)
compression_options1 = {'complib':'zlib', 'complevel':9, 'shuffle':True, 'least_significant_digit':2,
'normalization':'standard'}
compression_options2 = {'complib':'zlib', 'complevel':9, 'shuffle':True}
data.set_chunkshape_and_compression(nodename='Amitex_stress_1', compression_options=compression_options1)
data.set_nodes_compression_chunkshape(node_list=['grain_map', 'grain_map_raw','mask'],
compression_options=compression_options2)
# Now that we have compressed a few of the items of our dataset, the disk size of its HDF5 file should have diminished. Let us check again the size of its data items, and of the file:
data.print_dataset_content(short=True)
data.get_file_disk_size()
# The file size has not changed, surprisingly, even if the large `Amitex_stress_1` array has been shrinked from almost 50 Mo to roughly 5 Mo. This is due to a specific feature of HDF5 files: they do not free up the memory space that they have used in the past. The memory space remains associated to the file, and is used in priority when new data is written into the dataset.
#
# After changing the compression settings of one or several nodes in your dataset, if that induced a reduction of your actual data memory size, and that you want your file to be smaller on disk. To retrieve the fried up memory spacae, you may **repack** your file (overwrite it with a copy of itself, that has just the size require to store all actual data).
#
# To do that, you may use the *SampleData* method `repack_h5file`:
data.repack_h5file()
data.get_file_disk_size()
# You see that repacking the file has allowed to free some memory space and reduced its size.
# <div class="alert alert-info">
#
# **Note**
#
# Note that the size of the file is larger than the size of data items printed by `print_dataset_content`. This extra size is the memory size occupied by the data array storing *Element Tags* for the mesh `grains_mesh`. Element tags are not printed by the printing methods as they can be very numerous and pollute the lecture of the printed information.
# </div>
# Once again, you should repack your file at carefully chosen times, as is it a very costly operation for large datasets. The *SampleData* class constructor has an **autorepack** option. If it is set to `True`, the file is automatically repacked when closing the dataset.
# We can now close our dataset, and remove the original unarchived file:
# remove SampleData instance
del data
os.remove(dataset_file+'.h5')
os.remove(dataset_file+'.xdmf')
| examples/SampleDataUserGuide/5_SampleData_data_compression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# <div class='bar_title'></div>
#
# *Practical Data Science*
#
# # Introduction
#
# <NAME> / <NAME><br>
# Chair of Information Systems and Management
#
# Winter Semester 19/20
# + [markdown] slideshow={"slide_type": "subslide"} toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#Chair-of-Information-Systems-and-Management" data-toc-modified-id="Chair-of-Information-Systems-and-Management-1"><span class="toc-item-num">1 </span>Chair of Information Systems and Management</a></span><ul class="toc-item"><li><span><a href="#Team" data-toc-modified-id="Team-1.1"><span class="toc-item-num">1.1 </span>Team</a></span></li><li><span><a href="#Research-Areas" data-toc-modified-id="Research-Areas-1.2"><span class="toc-item-num">1.2 </span>Research Areas</a></span></li><li><span><a href="#Master-Teaching-Program" data-toc-modified-id="Master-Teaching-Program-1.3"><span class="toc-item-num">1.3 </span>Master Teaching Program</a></span></li></ul></li><li><span><a href="#Course-Organization" data-toc-modified-id="Course-Organization-2"><span class="toc-item-num">2 </span>Course Organization</a></span><ul class="toc-item"><li><span><a href="#Learning-Objective" data-toc-modified-id="Learning-Objective-2.1"><span class="toc-item-num">2.1 </span>Learning Objective</a></span></li><li><span><a href="#Tentative-Schedule-of-Topics" data-toc-modified-id="Tentative-Schedule-of-Topics-2.2"><span class="toc-item-num">2.2 </span>Tentative Schedule of Topics</a></span></li></ul></li><li><span><a href="#Programming-Language-(Python)" data-toc-modified-id="Programming-Language-(Python)-3"><span class="toc-item-num">3 </span>Programming Language (Python)</a></span><ul class="toc-item"><li><span><a href="#Why-Python?" data-toc-modified-id="Why-Python?-3.1"><span class="toc-item-num">3.1 </span>Why Python?</a></span></li><li><span><a href="#Job-Opportunities-with-Python" data-toc-modified-id="Job-Opportunities-with-Python-3.2"><span class="toc-item-num">3.2 </span>Job Opportunities with Python</a></span></li></ul></li><li><span><a href="#Next-Steps" data-toc-modified-id="Next-Steps-4"><span class="toc-item-num">4 </span>Next Steps</a></span></li></ul></div>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Chair of Information Systems and Management
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Team
# + [markdown] slideshow={"slide_type": "fragment"}
# <img src="images/01/team.png" style="width:80%">
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Research Areas
# + [markdown] hide_input=true slideshow={"slide_type": "fragment"}
# <img src="images/01/research.png" style="width:80%">
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Master Teaching Program
# + [markdown] hide_input=true slideshow={"slide_type": "fragment"}
# <img src="images/01/courses_master.png" style="width:100%">
# + [markdown] slideshow={"slide_type": "slide"}
# ## Course Organization
# + [markdown] cell_style="split" slideshow={"slide_type": "fragment"}
# - Classes
# - Lecture: Tuesday 10:15 – 11:45, Room 409 - Neue Universität
# - Tutorial: Office Hours (after lecture?)
#
#
# - Self-study and Tutorial
# - Work on weekly assignments and programming examples
# - Worksheet submissions are mandatory
#
# + [markdown] cell_style="split" slideshow={"slide_type": "fragment"}
# - Graded capstone group project
# - Work on a real-world problem set
# - Groups of 3 students
# - Regular submissions
#
# - WueCampus
# - Forum and information
# - Access Code: PDS1920
#
# - Github
# - Slides and worksheets
# - Submission of solutions
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Learning Objective
# + [markdown] slideshow={"slide_type": "fragment"}
# * The foundations, frameworks and applications of the emerging field of data science
#
#
# * Design, implement, and evaluate the core algorithms underlying an end-to-end data science workflow, including data import, analysis, and presentation of information
#
#
# * Leverage the Python application programming interface (API) ecosystem and data infrastructure that supports data acquisition, storage, retrieval and analysis
#
#
# * The application of a data-based analytical approach to identify and solve problems
#
#
# * Implementation and execution skills for data-driven business analytics
# + [markdown] slideshow={"slide_type": "slide"}
# ### Tentative Schedule of Topics
# + [markdown] slideshow={"slide_type": "fragment"}
# 1. Common data science environments (Markdown, Git, Jupyter, Colab)
#
# 2. Python fundamentals and EDA
#
# 3. tbd
# + [markdown] slideshow={"slide_type": "slide"}
# ## Programming Language (Python)
# + [markdown] slideshow={"slide_type": "fragment"}
# __Python is becoming the world’s most popular coding language__
# <img src="images/01/programming_economist.png" style="width:70%">
#
# Source: [The Economist](https://www.economist.com/graphic-detail/2018/07/26/python-is-becoming-the-worlds-most-popular-coding-language)
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Why Python?
#
# While there are several great languages for data analysis (such as R, Julia, Scala, ...) we decided to use python for this class because of the following reasons:
#
# * Excellent interactive shell and a large collection of open source packages
#
# * Very simple syntax
#
# * Simple enough for things to happen quickly and powerful enough to allow the implementation of the most complex ideas
#
# * Several highly optimized machine learning libraries that you can drag and drop into your code even if you know a minimum of Python
#
# * Good visualization capabilities (but not as great as R)
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Job Opportunities with Python
# + [markdown] hide_input=true slideshow={"slide_type": "fragment"}
# <img src="images/01/McKinsey.png" style="width:100%">
# + [markdown] hide_input=true slideshow={"slide_type": "subslide"}
# <img src="images/01/Stellen.png" style="width:100%">
# + [markdown] slideshow={"slide_type": "slide"}
# ## Next Steps
# + [markdown] slideshow={"slide_type": "fragment"}
# What projects are you interested in?
#
# 1. Deep learning on image data
# 2. Deep learning on text data
# 2. Advanced machine learning on tabular data
# + [markdown] slideshow={"slide_type": "fragment"}
# What prior programming experience do you have?
# + [markdown] slideshow={"slide_type": "fragment"}
# Register for a capstone project group in the WueCampus class room until Monday 21.10.2019 12.00 PM
# - At most 3 students per group
# - If groups are not full we will try to merge them in order to fill the gaps
| Lecture/01_Introduction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.10.2 64-bit
# language: python
# name: python3
# ---
stringA = 'joão'
stringB = 'maria'
stringA > stringB
stringA < stringB
len(stringA)
len(stringB)
stringc = 'nnnnnn'
stringd = 'B'
print(f"{stringc == stringd}")
print(f"{stringc > stringd}")
print(f"{stringc < stringd}")
print(f"{stringc != stringd}")
i = 0
while i >= len(stringc):
print('Aqui é Galo!!!')
i += 1
# +
A = int(input('Informe um numero: '))
B = int(input('Informe um numero: '))
X = A + B
print(f'X = {X}.')
# -
| 001-Curso-De-Python/002-IGTI/Modulo-01/Aulas ao Vivo/03022022.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import torch.nn as nn
import torch
from models_multiview import FrontaliseModelMasks_wider
from PIL import Image, ImageDraw
import matplotlib.pyplot as plt
# %matplotlib inline
from torchvision.transforms import CenterCrop, Resize, Compose, ToTensor
# -
MODEL_PATH = '/scratch/shared/slow/ow/faces_bmvc/models_to_share/' # Update with your path to the model files
# Predicting landmarks for the given images on MAFL. Here we give a demo for a selection of random images.
# +
# Parameters:
inner_nc = 256; num_classes = 10; num_additional_ids=32
classifier_model = MODEL_PATH + '/release/aflw_4views.pth'
# Load the model and classifier weights
classifier = nn.Sequential(nn.BatchNorm1d(inner_nc), nn.Linear(inner_nc, num_classes, bias=False))
model = FrontaliseModelMasks_wider(3, inner_nc=inner_nc, num_additional_ids=num_additional_ids)
classifier.load_state_dict(torch.load(classifier_model)['state_dict'])
model.load_state_dict(torch.load(classifier_model)['state_dict_model'])
classifier.eval()
model.eval()
# Load the image from MAFL and perform cropping to make more like VoxCeleb
img = Image.open('./examples/mafl/031184.jpg').convert('RGB') # More in the examples folder, or download the full dataset
orig_image = img
OFFSET = 35
CROP = 35
IMAGE_SIZE = (256,256)
img = img.crop((0,CROP,img.size[0], img.size[1]))
size = (img.size[0]-OFFSET*2, img.size[1]-OFFSET*2)
transform = Compose([CenterCrop((size[1], size[0])), Resize(IMAGE_SIZE), ToTensor()])
# Predicted landmarks
xc = model.encoder(transform(img).unsqueeze(0))
landmarks = classifier(xc.squeeze().unsqueeze(0))
landmarks = landmarks.view(-1,2)
# Convert to image locations
offsetx = size[0]
offsety = size[1]
addx = OFFSET
addy = OFFSET+CROP
landmarks[:,0] = (landmarks[:,0] + 1) * offsetx / 2. + addx
landmarks[:,1] = (landmarks[:,1] + 1) * offsety / 2. + addy
# Visualise landmarks
draw = ImageDraw.Draw(orig_image)
colors = [(0,256,0), (256,0,0), (0,0,256), (256,256,0), (0,0,0)]
for img_index_i in range(0, 5):
draw.line([landmarks[img_index_i,0], landmarks[img_index_i,1]-4, landmarks[img_index_i,0], landmarks[img_index_i,1]+4], fill=colors[img_index_i % 5])
draw.line([landmarks[img_index_i,0]-4, landmarks[img_index_i,1], landmarks[img_index_i,0]+4, landmarks[img_index_i,1]], fill=colors[img_index_i % 5])
plt.imshow(orig_image)
# -
# Predicting landmarks for the given images on 300W. Here we give a demo for a selection of random images.
# +
# Parameters:
import numpy as np
inner_nc = 256; num_classes = 136; num_additional_ids=32
classifier_model = MODEL_PATH + '/release/300w_4views.pt'
# Load the model and classifier weights
classifier = nn.Sequential(nn.BatchNorm1d(inner_nc), nn.Linear(inner_nc, num_classes, bias=False))
model = FrontaliseModelMasks_wider(3, inner_nc=inner_nc, num_additional_ids=num_additional_ids)
classifier.load_state_dict(torch.load(classifier_model)['state_dict'])
model.load_state_dict(torch.load(classifier_model)['state_dict_model'])
classifier.eval()
model.eval()
# Load the image from 300W and perform cropping to make more like VoxCeleb
img = Image.open('./examples/300w/ibug_image_99.jpg').convert('RGB') # Download the full dataset for more examples
bbox = np.array([312.4117, 57.3576, 416.5690, 166.9425])
# Crop and resize again to make more VoxCeleb
orig_image = img
OFFSET = (bbox[2:] - bbox[0:2])*0.1
OFFSET = [-int(OFFSET[0]+0.5), -int(OFFSET[1] + 0.5)]
img = img.crop((bbox[0]+OFFSET[0], bbox[1]+OFFSET[1], bbox[2]-OFFSET[0], bbox[3]-OFFSET[1]))
size = (img.size[0], img.size[1])
transform = Compose([Resize((256, 256)), ToTensor()])
OFFSET = bbox[0:2] + OFFSET # OFFSET 10+5 w/ 1000 examples = 8.88; 10+10=8.2, 10+15=7.5
size = (img.size[0], img.size[1])
# Predicted landmarks
xc = model.encoder(transform(img).unsqueeze(0))
landmarks = classifier(xc.squeeze().unsqueeze(0))
landmarks = landmarks.view(-1,2)
# Convert to image locations
offsetx = size[0]
offsety = size[1]
addx = OFFSET[0]
addy = OFFSET[1]
landmarks[:,0] = (landmarks[:,0] + 1) * offsetx / 2. + addx
landmarks[:,1] = (landmarks[:,1] + 1) * offsety / 2. + addy
# Visualise landmarks
draw = ImageDraw.Draw(orig_image)
colors = [(0,0,256)]
for img_index_i in range(0, 68):
draw.line([landmarks[img_index_i,0], landmarks[img_index_i,1]-4, landmarks[img_index_i,0], landmarks[img_index_i,1]+4], fill=colors[0 % 5])
draw.line([landmarks[img_index_i,0]-4, landmarks[img_index_i,1], landmarks[img_index_i,0]+4, landmarks[img_index_i,1]], fill=colors[0 % 5])
OFFSET = (bbox[2:] - bbox[0:2])*0.1
OFFSET = [-int(OFFSET[0]+0.5), -int(OFFSET[1] + 0.5)]
orig_image = orig_image.crop((bbox[0]+OFFSET[0], bbox[1]+OFFSET[1], bbox[2]-OFFSET[0], bbox[3]-OFFSET[1]))
plt.imshow(orig_image)
# -
# And now show how we can predict emotion.
# +
# Parameters:
import numpy as np
inner_nc = 256; num_classes = 8; num_additional_ids=32
classifier_model = MODEL_PATH + '/release/affectnet_4views.pth'
# Load the model and classifier weights
classifier = nn.Sequential(nn.BatchNorm1d(inner_nc), nn.Linear(inner_nc, num_classes, bias=False))
model = FrontaliseModelMasks_wider(3, inner_nc=inner_nc, num_additional_ids=num_additional_ids)
classifier.load_state_dict(torch.load(classifier_model)['state_dict'])
model.load_state_dict(torch.load(classifier_model)['state_dict_model'])
classifier.eval()
model.eval()
# Load image from affectnet
img = './examples/affectnet/991_7803ea5b854f9d35db61a9afe2a3fc834894ca2a861af2ce151b206e.jpg'
img = Image.open(img).convert('RGB')
transform = Compose([Resize((256, 256)) ,ToTensor()])
# Predicted emotion
xc = model.encoder(transform(img).unsqueeze(0))
probabilities = nn.Sigmoid()(classifier(xc.squeeze().unsqueeze(0)))
probabilities = probabilities[0] / probabilities.sum()
prediction = probabilities.max(0)
emotions = ['Neutral', 'Happy', 'Sad', 'Surprise', 'Fear', 'Disgust', 'Anger', 'Contempt']
plt.imshow(img)
print("Probabilities: \n {} {:0.4f} \n {} {:0.4f} \n {} {:0.4f} \n {} {:0.4f} \n {} {:0.4f} \n {} {:0.4f} \n {} {:0.4f}".format(
emotions[0], probabilities[0].cpu().item(),
emotions[1], probabilities[1].cpu().item(),
emotions[2], probabilities[2].cpu().item(),
emotions[3], probabilities[3].cpu().item(),
emotions[4], probabilities[4].cpu().item(),
emotions[5], probabilities[5].cpu().item(),
emotions[6], probabilities[6].cpu().item(),
emotions[7], probabilities[7].cpu().item()))
print('\n With probability %f, predicted emotion %s.' % (prediction[0].cpu().item(), emotions[prediction[1].item()]))
# +
# Parameters:
import numpy as np
inner_nc = 256; num_classes = 3; num_additional_ids=32
classifier_model = MODEL_PATH + '/release/aflwpose_nv4.pth'
# Load the model and classifier weights
classifier = nn.Sequential(nn.BatchNorm1d(inner_nc), nn.Linear(inner_nc, num_classes, bias=False))
model = FrontaliseModelMasks_wider(3, inner_nc=inner_nc, num_additional_ids=num_additional_ids)
classifier.load_state_dict(torch.load(classifier_model)['state_dict'])
model.load_state_dict(torch.load(classifier_model)['state_dict_model'])
classifier.eval()
model.eval()
# Load the image from the AFLW test set and perform cropping
img = Image.open('./examples/aflw/image66922.jpg').convert('RGB')
img = img.crop([237,228,332,323])
transform = Compose([Resize((256, 256)), ToTensor()])
# Predicted pose
xc = model.encoder(transform(img).unsqueeze(0))
pose = 180/np.pi * classifier(xc.squeeze().unsqueeze(0)).cpu().data.numpy()[0]
gtpose = 180/np.pi * np.array([0.05019387, -0.11107627, -0.6923566])
plt.imshow(img)
print('Predicted head pose: roll %.2f, pitch %.2f, yaw %.2f' %(pose[0], pose[1], pose[2]))
print('Ground truth head pose: roll %.2f, pitch %.2f, yaw %.2f' %(gtpose[0], gtpose[1], gtpose[2]))
# -
| embedders/FAb-Net/FAb-Net/code/demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: covid-19
# language: python
# name: covid-19
# ---
# # Estimating COVID-19's $R_t$ for x nations
#
# <NAME> - May 02
#
# This notebook is an adaptation of [Kevin Systrom's notebook](https://github.com/k-sys/covid-19/blob/master/Realtime%20Rt%20mcmc.ipynb) to calculate $R_t$, however, instead of US states this does the calculation for worldwide nations.
# ## Estimating COVID-19's $R_t$ in Real-Time with PYMC3
#
# <NAME> - April 22
#
# Model originally built by [<NAME>](https://github.com/tvladeck) in Stan, parts inspired by the work over at https://epiforecasts.io/, lots of help from [<NAME>](https://twitter.com/twiecki). Thank you to everyone who helped.
#
# This notebook is a WIP - I'll add more context and commentary over the coming week.
# +
# setup conda environment
# https://docs.conda.io/projects/conda/en/latest/user-guide/tasks/manage-environments.html
# how to add a conda environment to jupyter notebooks
# https://medium.com/@nrk25693/how-to-add-your-conda-environment-to-your-jupyter-notebook-in-just-4-steps-abeab8b8d084
# check if python executable matches environment
import sys
print(sys.executable)
# +
# For some reason Theano is unhappy when I run the GP, need to disable future warnings
import warnings
warnings.simplefilter(action='ignore', category=FutureWarning)
import os
import requests
import pymc3 as pm
import pandas as pd
import numpy as np
import theano
import theano.tensor as tt
from matplotlib import pyplot as plt
from matplotlib import dates as mdates
from matplotlib import ticker
from datetime import date
from datetime import datetime
from IPython.display import clear_output
# %config InlineBackend.figure_format = 'retina'
# -
# ## Load State Information
# #### Load
file = '../data/covid19-feature.csv'
states_AT = pd.read_csv(file)
states_AT['date'] = pd.to_datetime(states_AT['date'], format='%Y-%m-%d')
states_AT.head()
# date, state, positive
states_AT_melt = states_AT[['date', 'location', 'total_cases']]
states_AT_melt.columns = ['date', 'state', 'positive']
states_AT_melt.head()
# #### Clean data with known modifications
# +
# Errors in Covidtracking.com
states_AT_melt_i = states_AT_melt.set_index(['state', 'date'])
states_AT_melt_i.loc[('Spain','2020-04-19'), 'positive'] = 190585
# states_AT_melt_i.loc[('Thailand','2020-03-04'), 'positive'] = 1
# states_AT_melt_i.loc[('Thailand','2020-03-06'), 'positive'] = 1
# states_AT_melt_i.loc[('Thailand','2020-03-08'), 'positive'] = 1
# states_AT_melt_i.loc[('Thailand','2020-03-09'), 'positive'] = 1
# states_AT_melt_i.loc[('Thailand','2020-03-10'), 'positive'] = 1
# states_AT_melt_i.loc[('Thailand','2020-03-13'), 'positive'] = 1
states_AT_melt_i.head()
# -
# #### Integrity Check
# +
# Make sure that all the states have current data
states = states_AT_melt_i
today = datetime.combine(date.today(), datetime.min.time())
last_updated = states.reset_index('date').groupby('state')['date'].max()
is_current = last_updated < today
try:
assert is_current.sum() == 0
except AssertionError:
print("Not all states have updated")
display(last_updated[is_current])
# Ensure all case diffs are greater than zero
for state, grp in states.groupby('state'):
new_cases = grp.positive.diff().dropna()
is_positive = new_cases.ge(0)
try:
assert is_positive.all()
except AssertionError:
print(f"Warning: {state} has date with negative case counts")
display(new_cases[~is_positive])
# Let's make sure that states have added cases
idx = pd.IndexSlice
assert not states.loc[idx[:, '2020-04-22':'2020-04-23'], 'positive'].groupby('state').diff().dropna().eq(0).any()
# -
# ## Load Patient Information
# #### Download
# ~100mb download (be ... patient!)
# +
def download_file(url, local_filename):
"""From https://stackoverflow.com/questions/16694907/"""
with requests.get(url, stream=True) as r:
r.raise_for_status()
with open(local_filename, 'wb') as f:
for chunk in r.iter_content(chunk_size=8192):
if chunk: # filter out keep-alive new chunks
f.write(chunk)
return local_filename
URL = "https://raw.githubusercontent.com/beoutbreakprepared/nCoV2019/master/latest_data/latestdata.csv"
LINELIST_PATH = 'data/linelist.csv'
if not os.path.exists(LINELIST_PATH):
print('Downloading file, this will take a while ~100mb')
try:
download_file(URL, LINELIST_PATH)
clear_output(wait=True)
print('Done downloading.')
except:
print('Something went wrong. Try again.')
else:
print('Already downloaded CSV')
# -
# #### Parse & Clean Patient Info
# +
# Load the patient CSV
patients = pd.read_csv(
'data/linelist.csv',
parse_dates=False,
usecols=[
'date_confirmation',
'date_onset_symptoms'],
low_memory=False)
patients.columns = ['Onset', 'Confirmed']
# There's an errant reversed date
patients = patients.replace('01.31.2020', '31.01.2020')
# Only keep if both values are present
patients = patients.dropna()
# Must have strings that look like individual dates
# "2020.03.09" is 10 chars long
is_ten_char = lambda x: x.str.len().eq(10)
patients = patients[is_ten_char(patients.Confirmed) &
is_ten_char(patients.Onset)]
# Convert both to datetimes
patients.Confirmed = pd.to_datetime(
patients.Confirmed, format='%d.%m.%Y')
patients.Onset = pd.to_datetime(
patients.Onset, format='%d.%m.%Y')
# Only keep records where confirmed > onset
patients = patients[patients.Confirmed >= patients.Onset]
# -
# #### Show Relationship between Onset of Symptoms and Confirmation
# +
ax = patients.plot.scatter(
title='Onset vs. Confirmed Dates - COVID19',
x='Onset',
y='Confirmed',
alpha=.1,
lw=0,
s=10,
figsize=(6,6))
formatter = mdates.DateFormatter('%m/%d')
locator = mdates.WeekdayLocator(interval=2)
for axis in [ax.xaxis, ax.yaxis]:
axis.set_major_formatter(formatter)
axis.set_major_locator(locator)
# -
# #### Calculate the Probability Distribution of Delay
# +
# Calculate the delta in days between onset and confirmation
delay = (patients.Confirmed - patients.Onset).dt.days
# Convert samples to an empirical distribution
p_delay = delay.value_counts().sort_index()
new_range = np.arange(0, p_delay.index.max()+1)
p_delay = p_delay.reindex(new_range, fill_value=0)
p_delay /= p_delay.sum()
# Show our work
fig, axes = plt.subplots(ncols=2, figsize=(9,3))
p_delay.plot(title='P(Delay)', ax=axes[0])
p_delay.cumsum().plot(title='P(Delay <= x)', ax=axes[1])
for ax in axes:
ax.set_xlabel('days')
# -
# ## A Single State
#
# #### Select State Data
state = 'Sweden'
confirmed = states.xs(state).positive.diff().dropna()
confirmed.tail()
# ### Translate Confirmation Dates to Onset Dates
#
# Our goal is to translate positive test counts to the dates where they likely occured. Since we have the distribution, we can distribute case counts back in time according to that distribution. To accomplish this, we reverse the case time series, and convolve it using the distribution of delay from onset to confirmation. Then we reverse the series again to obtain the onset curve. Note that this means the data will be 'right censored' which means there are onset cases that have yet to be reported so it looks as if the count has gone down.
# +
def confirmed_to_onset(confirmed, p_delay):
assert not confirmed.isna().any()
# Reverse cases so that we convolve into the past
convolved = np.convolve(confirmed[::-1].values, p_delay)
# Calculate the new date range
dr = pd.date_range(end=confirmed.index[-1],
periods=len(convolved))
# Flip the values and assign the date range
onset = pd.Series(np.flip(convolved), index=dr)
return onset
onset = confirmed_to_onset(confirmed, p_delay)
# -
# ### Adjust for Right-Censoring
#
# Since we distributed observed cases into the past to recreate the onset curve, we now have a right-censored time series. We can correct for that by asking what % of people have a delay less than or equal to the time between the day in question and the current day.
#
# For example, 5 days ago, there might have been 100 cases onset. Over the course of the next 5 days some portion of those cases will be reported. This portion is equal to the cumulative distribution function of our delay distribution. If we know that portion is say, 60%, then our current count of onset on that day represents 60% of the total. This implies that the total is 166% higher. We apply this correction to get an idea of what actual onset cases are likely, thus removing the right censoring.
# +
def adjust_onset_for_right_censorship(onset, p_delay):
cumulative_p_delay = p_delay.cumsum()
# Calculate the additional ones needed so shapes match
ones_needed = len(onset) - len(cumulative_p_delay)
padding_shape = (0, ones_needed)
# Add ones and flip back
cumulative_p_delay = np.pad(
cumulative_p_delay,
padding_shape,
constant_values=1)
cumulative_p_delay = np.flip(cumulative_p_delay)
# Adjusts observed onset values to expected terminal onset values
adjusted = onset / cumulative_p_delay
return adjusted, cumulative_p_delay
adjusted, cumulative_p_delay = adjust_onset_for_right_censorship(onset, p_delay)
# -
# Take a look at all three series: confirmed, onset and onset adjusted for right censoring.
# +
fig, ax = plt.subplots(figsize=(5,3))
confirmed.plot(
ax=ax,
label='Confirmed',
title=state,
c='k',
alpha=.25,
lw=1)
onset.plot(
ax=ax,
label='Onset',
c='k',
lw=1)
adjusted.plot(
ax=ax,
label='Adjusted Onset',
c='k',
linestyle='--',
lw=1)
ax.legend();
# -
# Let's have the model run on days where we have enough data ~last 60 or so
# ### Sample the Posterior with PyMC3
# We assume a poisson likelihood function and feed it what we believe is the onset curve based on reported data. We model this onset curve based on the same math in the previous notebook:
#
# $$ I^\prime = Ie^{\gamma(R_t-1)} $$
#
# We define $\theta = \gamma(R_t-1)$ and model $ I^\prime = Ie^{\theta} $ where $\theta$ observes a random walk. We let $\gamma$ vary independently based on known parameters for the serial interval. Therefore, we can recover $R_t$ easily by $R_t = \frac{\theta}{\gamma}+1$
#
# The only tricky part is understanding that we're feeding in _onset_ cases to the likelihood. So $\mu$ of the poisson is the positive, non-zero, expected onset cases we think we'd see today.
#
# We calculate this by figuring out how many cases we'd expect there to be yesterday total when adjusted for bias and plugging it into the first equation above. We then have to re-bias this number back down to get the expected amount of onset cases observed that day.
class MCMCModel(object):
def __init__(self, region, onset, cumulative_p_delay, window=90):
# Just for identification purposes
self.region = region
# For the model, we'll only look at the last N
self.onset = onset.iloc[-window:]
self.cumulative_p_delay = cumulative_p_delay[-window:]
# Where we store the results
self.trace = None
self.trace_index = self.onset.index[1:]
def run(self, chains=1, tune=3000, draws=1000, target_accept=.95):
with pm.Model() as model:
# Random walk magnitude
step_size = pm.HalfNormal('step_size', sigma=.03)
# Theta random walk
theta_raw_init = pm.Normal('theta_raw_init', 0.1, 0.1)
theta_raw_steps = pm.Normal('theta_raw_steps', shape=len(self.onset)-2) * step_size
theta_raw = tt.concatenate([[theta_raw_init], theta_raw_steps])
theta = pm.Deterministic('theta', theta_raw.cumsum())
# Let the serial interval be a random variable and calculate r_t
serial_interval = pm.Gamma('serial_interval', alpha=6, beta=1.5)
gamma = 1.0 / serial_interval
r_t = pm.Deterministic('r_t', theta/gamma + 1)
inferred_yesterday = self.onset.values[:-1] / self.cumulative_p_delay[:-1]
expected_today = inferred_yesterday * self.cumulative_p_delay[1:] * pm.math.exp(theta)
# Ensure cases stay above zero for poisson
mu = pm.math.maximum(.1, expected_today)
observed = self.onset.round().values[1:]
cases = pm.Poisson('cases', mu=mu, observed=observed)
self.trace = pm.sample(
chains=chains,
tune=tune,
draws=draws,
target_accept=target_accept)
return self
def run_gp(self):
with pm.Model() as model:
gp_shape = len(self.onset) - 1
length_scale = pm.Gamma("length_scale", alpha=3, beta=.4)
eta = .05
cov_func = eta**2 * pm.gp.cov.ExpQuad(1, length_scale)
gp = pm.gp.Latent(mean_func=pm.gp.mean.Constant(c=0),
cov_func=cov_func)
# Place a GP prior over the function f.
theta = gp.prior("theta", X=np.arange(gp_shape)[:, None])
# Let the serial interval be a random variable and calculate r_t
serial_interval = pm.Gamma('serial_interval', alpha=6, beta=1.5)
gamma = 1.0 / serial_interval
r_t = pm.Deterministic('r_t', theta / gamma + 1)
inferred_yesterday = self.onset.values[:-1] / self.cumulative_p_delay[:-1]
expected_today = inferred_yesterday * self.cumulative_p_delay[1:] * pm.math.exp(theta)
# Ensure cases stay above zero for poisson
mu = pm.math.maximum(.1, expected_today)
observed = self.onset.round().values[1:]
cases = pm.Poisson('cases', mu=mu, observed=observed)
self.trace = pm.sample(chains=1, tune=1000, draws=1000, target_accept=.8)
return self
# ### Run Pymc3 Model
# +
def df_from_model(model):
r_t = model.trace['r_t']
mean = np.mean(r_t, axis=0)
median = np.median(r_t, axis=0)
hpd_90 = pm.stats.hpd(r_t, credible_interval=.9)
hpd_50 = pm.stats.hpd(r_t, credible_interval=.5)
idx = pd.MultiIndex.from_product([
[model.region],
model.trace_index
], names=['region', 'date'])
df = pd.DataFrame(data=np.c_[mean, median, hpd_90, hpd_50], index=idx,
columns=['mean', 'median', 'lower_90', 'upper_90', 'lower_50','upper_50'])
return df
def create_and_run_model(name, state):
confirmed = state.positive.diff().dropna()
onset = confirmed_to_onset(confirmed, p_delay)
adjusted, cumulative_p_delay = adjust_onset_for_right_censorship(onset, p_delay)
return MCMCModel(name, onset, cumulative_p_delay).run()
# -
# df.loc[~df.index.str.endswith('_test')]
states_model = states.iloc[states.index.get_level_values('state').isin([
'United States',
'Spain',
'Italy',
'United Kingdom',
'France',
'Germany',
'Turkey',
'Russia',
'Brazil',
'Iran',
'China',
'Canada',
'Belgium',
'Peru',
'Netherlands',
'India',
'Switzerland',
'Ecuador',
'Saudi Arabia',
'Portugal',
'Sweden',
'Ireland',
'Mexico',
'Pakistan',
'Chile',
'Singapore',
'Israel',
'Belarus',
'Austria',
'Qatar',
'Japan',
'South Korea',
'Norway'
])]
states_model.head()
# +
models = {}
for state, grp in states_model.groupby('state'):
print(state)
if state in models:
print(f'Skipping {state}, already in cache')
continue
models[state] = create_and_run_model(state, grp.droplevel(0))
# -
# ### Handle Divergences
# +
# Check to see if there were divergences
n_diverging = lambda x: x.trace['diverging'].nonzero()[0].size
divergences = pd.Series([n_diverging(m) for m in models.values()], index=models.keys())
has_divergences = divergences.gt(0)
print('Diverging states:')
display(divergences[has_divergences])
# Rerun states with divergences
for state, n_divergences in divergences[has_divergences].items():
models[state].run()
# -
# ## Compile Results
# +
results = None
for state, model in models.items():
df = df_from_model(model)
if results is None:
results = df
else:
results = pd.concat([results, df], axis=0)
# -
# ### Render to CSV
# Uncomment if you'd like
results.to_csv('../data/world_rt.csv')
# ### Render Charts
def plot_rt(name, result, ax, c=(.3,.3,.3,1), ci=(0,0,0,.05)):
ax.set_ylim(0, 2)
ax.set_title(name)
ax.plot(result['median'],
marker='o',
markersize=4,
markerfacecolor='w',
lw=1,
c=c,
markevery=2)
ax.fill_between(
result.index,
result['lower_50'].values,
result['upper_50'].values,
color=ci,
lw=0)
ax.axhline(1.0, linestyle=':', lw=1)
ax.xaxis.set_major_formatter(mdates.DateFormatter('%m/%d'))
ax.xaxis.set_major_locator(mdates.WeekdayLocator(interval=2))
# +
ncols = 3
nrows = int(np.ceil(results.index.levels[0].shape[0] / ncols))
fig, axes = plt.subplots(
nrows=nrows,
ncols=ncols,
figsize=(10, nrows*3),
sharey='row')
for ax, (state, result) in zip(axes.flat, results.groupby('region')):
plot_rt(state, result.droplevel(0), ax)
fig.tight_layout()
fig.set_facecolor('w')
| jupyter/covid19-world-rt.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="Ocb1x0zeP7Fe"
# # Compile and deploy the TFX pipeline to Kubeflow Pipelines
#
# This notebook is the second of two notebooks that guide you through automating the [Real-time Item-to-item Recommendation with BigQuery ML Matrix Factorization and ScaNN](https://github.com/GoogleCloudPlatform/analytics-componentized-patterns/tree/master/retail/recommendation-system/bqml-scann) solution with a pipeline.
#
# Use this notebook to compile the TFX pipeline to a Kubeflow Pipelines (KFP) package. This process creates an Argo YAML file in a .tar.gz package, and is accomplished through the following steps:
#
# 1. Build a custom container image that includes the solution modules.
# 2. Compile the TFX Pipeline using the TFX command-line interface (CLI).
# 3. Deploy the compiled pipeline to KFP.
#
# The pipeline workflow is implemented in the [pipeline.py](tfx_pipeline/pipeline.py) module. The [runner.py](tfx_pipeline/runner.py) module reads the configuration settings from the [config.py](tfx_pipeline/config.py) module, defines the runtime parameters of the pipeline, and creates a KFP format that is executable on AI Platform pipelines.
#
# Before starting this notebook, you must run the [tfx01_interactive](tfx01_interactive.ipynb) notebook to create the TFX pipeline.
#
# + [markdown] id="qdYregr9P7Fl"
# ## Install required libraries
# + id="oFAdKJSdP7Fm"
# %load_ext autoreload
# %autoreload 2
# + id="KfrHaJmyP7Fm"
# !pip install -q -U kfp
# + [markdown] id="C-Ibd8heP7Fn"
# ## Set environment variables
#
# Update the following variables to reflect the values for your GCP environment:
#
# # + `PROJECT_ID`: The ID of the Google Cloud project you are using to implement this solution.
# # + `BUCKET`: The name of the Cloud Storage bucket you created to use with this solution. The `BUCKET` value should be just the bucket name, so `myBucket` rather than `gs://myBucket`.
# # + `GKE_CLUSTER_NAME`: The name of the Kubernetes Engine cluster used by the AI Platform pipeline. You can find this by looking at the **Cluster** column of the `kubeflow-pipelines` pipeline instance on the AI Platform Pipelines page.
# # + `GKE_CLUSTER_ZONE`: The zone of the Kubernetes Engine cluster used by the AI Platform pipeline. You can find this by looking at the **Zone** column of the `kubeflow-pipelines` pipeline instance on the AI Platform Pipelines page.
# + id="VHES7S4iP7Fn"
import os
os.environ["PROJECT_ID"] = "yourProject" # Set your project.
os.environ["BUCKET"] = "yourBucket" # Set your bucket.
os.environ["GKE_CLUSTER_NAME"] = "yourCluster" # Set your GKE cluster name.
os.environ["GKE_CLUSTER_ZONE"] = "yourClusterZone" # Set your GKE cluster zone.
os.environ["IMAGE_NAME"] = "tfx-ml"
os.environ["TAG"] = "tfx0.25.0"
os.environ[
"ML_IMAGE_URI"
] = f'gcr.io/{os.environ.get("PROJECT_ID")}/{os.environ.get("IMAGE_NAME")}:{os.environ.get("TAG")}'
os.environ["NAMESPACE"] = "kubeflow-pipelines"
os.environ["ARTIFACT_STORE_URI"] = f'gs://{os.environ.get("BUCKET")}/tfx_artifact_store'
os.environ["GCS_STAGING_PATH"] = f'{os.environ.get("ARTIFACT_STORE_URI")}/staging'
os.environ["RUNTIME_VERSION"] = "2.2"
os.environ["PYTHON_VERSION"] = "3.7"
os.environ["BEAM_RUNNER"] = "DirectRunner"
os.environ[
"MODEL_REGISTRY_URI"
] = f'{os.environ.get("ARTIFACT_STORE_URI")}/model_registry'
os.environ["PIPELINE_NAME"] = "tfx_bqml_scann"
# + id="OQua8zUuP7Fo"
from tfx_pipeline import config
for key, value in config.__dict__.items():
if key.isupper():
print(f"{key}: {value}")
# + [markdown] id="jiOIUg-fP7Fo"
# ## Run the Pipeline locally by using the Beam runner
# + id="zuheADbDP7Fp"
import logging
import kfp
import ml_metadata as mlmd
import tensorflow as tf
import tfx
from ml_metadata.proto import metadata_store_pb2
from tfx.orchestration.beam.beam_dag_runner import BeamDagRunner
from tfx_pipeline import pipeline as pipeline_module
logging.getLogger().setLevel(logging.INFO)
print("TFX Version:", tfx.__version__)
# + id="K5UbLvJ8P7Fp"
pipeline_root = f"{config.ARTIFACT_STORE_URI}/{config.PIPELINE_NAME}_beamrunner"
model_regisrty_uri = f"{config.MODEL_REGISTRY_URI}_beamrunner"
local_mlmd_sqllite = "mlmd/mlmd.sqllite"
print(f"Pipeline artifacts root: {pipeline_root}")
print(f"Model registry location: {model_regisrty_uri}")
if tf.io.gfile.exists(pipeline_root):
print("Removing previous artifacts...")
tf.io.gfile.rmtree(pipeline_root)
if tf.io.gfile.exists("mlmd"):
print("Removing local mlmd SQLite...")
tf.io.gfile.rmtree("mlmd")
print("Creating mlmd directory...")
tf.io.gfile.mkdir("mlmd")
metadata_connection_config = metadata_store_pb2.ConnectionConfig()
metadata_connection_config.sqlite.filename_uri = local_mlmd_sqllite
metadata_connection_config.sqlite.connection_mode = 3
print("ML metadata store is ready.")
beam_pipeline_args = [
f"--runner=DirectRunner",
f"--project={config.PROJECT_ID}",
f"--temp_location={config.ARTIFACT_STORE_URI}/beam/tmp",
]
pipeline_module.SCHEMA_DIR = "tfx_pipeline/schema"
pipeline_module.LOOKUP_CREATOR_MODULE = "tfx_pipeline/lookup_creator.py"
pipeline_module.SCANN_INDEXER_MODULE = "tfx_pipeline/scann_indexer.py"
# + id="BA3IbMZZP7Fq"
runner = BeamDagRunner()
pipeline = pipeline_module.create_pipeline(
pipeline_name=config.PIPELINE_NAME,
pipeline_root=pipeline_root,
project_id=config.PROJECT_ID,
bq_dataset_name=config.BQ_DATASET_NAME,
min_item_frequency=15,
max_group_size=10,
dimensions=50,
num_leaves=500,
eval_min_recall=0.8,
eval_max_latency=0.001,
ai_platform_training_args=None,
beam_pipeline_args=beam_pipeline_args,
model_regisrty_uri=model_regisrty_uri,
metadata_connection_config=metadata_connection_config,
enable_cache=True,
)
runner.run(pipeline)
# + [markdown] id="sDHQOTlzP7Fr"
# ## Build the container image
#
# The pipeline uses a custom container image, which is a derivative of the [tensorflow/tfx:0.25.0](https://hub.docker.com/r/tensorflow/tfx) image, as a runtime execution environment for the pipeline's components. The container image is defined in a [Dockerfile](tfx_pipeline/Dockerfile).
#
# The container image installs the required libraries and copies over the modules from the solution's [tfx_pipeline](tfx_pipeline) directory, where the custom components are implemented. The container image is also used by AI Platform Training for executing the training jobs.
#
# Build the container image using Cloud Build and then store it in Cloud Container Registry:
#
# + id="PW_sUGUtP7Fr"
# !gcloud builds submit --tag $ML_IMAGE_URI tfx_pipeline
# + [markdown] id="-_bmd8YmP7Fr"
# ## Compile the TFX pipeline using the TFX CLI
#
# Use the TFX CLI to compile the TFX pipeline to the KFP format, which allows the pipeline to be deployed and executed on AI Platform Pipelines. The output is a .tar.gz package containing an Argo definition of your pipeline.
#
# + id="n5QGIAclP7Fs"
# !rm ${PIPELINE_NAME}.tar.gz
# !tfx pipeline compile \
# --engine=kubeflow \
# --pipeline_path=tfx_pipeline/runner.py
# + [markdown] id="qo6z0fQcP7Fs"
# ## Deploy the compiled pipeline to KFP
#
# Use the KFP CLI to deploy the pipeline to a hosted instance of KFP on AI Platform Pipelines:
#
# + id="baYgjczHP7Fs" language="bash"
#
# gcloud container clusters get-credentials ${GKE_CLUSTER_NAME} --zone ${GKE_CLUSTER_ZONE}
# export KFP_ENDPOINT=$(kubectl describe configmap inverse-proxy-config -n ${NAMESPACE} | grep "googleusercontent.com")
#
# kfp --namespace=${NAMESPACE} --endpoint=${KFP_ENDPOINT} \
# pipeline upload \
# --pipeline-name=${PIPELINE_NAME} \
# ${PIPELINE_NAME}.tar.gz
# + [markdown] id="xuSbXIvdbYqL"
# After deploying the pipeline, you can browse it by following these steps:
#
# 1. Open the [AI Platform Pipelines page](https://pantheon.corp.google.com/ai-platform/pipelines/clusters).
# 1. For the `kubeflow-pipelines` instance, click **Open Pipelines Dashboard**.
# 1. Click **Pipelines** and confirm that `tfx_bqml_scann` appears on the list of pipelines.
# + [markdown] id="YlNx68crP7Ft"
# ## Run the deployed pipeline
#
# Run the pipeline by using the KFP UI:
#
# 1. Open the [AI Platform Pipelines page](https://pantheon.corp.google.com/ai-platform/pipelines/clusters).
# 1. For the `kubeflow-pipelines` instance, click **Open Pipelines Dashboard**.
# 1. Click **Experiments**.
# 1. Click **Create Run**.
# 1. For **Pipeline**, choose **tfx_bqml_scann** and then click **Use this pipeline**.
# 1. For **Pipeline Version**, choose **tfx_bqml_scann**.
# 1. For **Run name**, type `run of tfx_bqml_scann`.
# 1. For **Experiment**, choose **Default** and then click **Use this experiment**.
# 1. Click **Start**.
#
#
# + [markdown] id="HmudthxFlnTm"
# The pipelines dashboard displays a list of pipeline runs. In the list, click the name of your run to see a graph of the run displayed. While your run is still in progress, the graph changes as each step executes. Click any step to explore the run's inputs, outputs, logs, etc.
# + [markdown] id="b9lcrRA_P7Fu"
# ## License
#
# Copyright 2020 Google LLC
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License. You may obtain a copy of the License at: http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
#
# See the License for the specific language governing permissions and limitations under the License.
#
# **This is not an official Google product but sample code provided for an educational purpose**
| notebooks/community/analytics-componetized-patterns/retail/recommendation-system/bqml-scann/tfx02_deploy_run.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: U4S1DS10 (Python 3.7)
# language: python
# name: u4-s1-nlp-ds10
# ---
# ### NLP Med Cab Part ii
# - cont. from 1.2 Notebook
# - Perform stemming and lemmatization on tokens
# +
"""
Import Statements
"""
# Classics
import numpy as np
import pandas as pd
# Base
from collections import Counter
# Plotting
import squarify
import seaborn as sns
import matplotlib.pyplot as plt
# NLP Libraries
import spacy
from spacy.tokenizer import Tokenizer
nlp = spacy.load("en_core_web_md")
# -
pwd
# ls
df = pd.read_csv('/Users/jorge/Med-Cabinet-2/Data/medcab1_2.csv')
df = df.fillna('none')
df = df.drop('Unnamed: 0', 1)
df = df.replace([True, False],
[1.0, 0.0])
print(df.shape)
df.head(3)
# ### Search Results by quality strains:
good_stuff = df[df['Rating'] >= 4.0]
good_stuff = df.replace(np.nan, '', regex=True)
# ### Standaridize our string input:
df.columns
# Combine labels into one string, for vectorization:
import re, string
# df['tokens'].head()
df['tokens'] = df['tokens'].str.replace(r'\\(x|n)[a-z0-9]{0,2}', ' ')
df['tokens'] = df['tokens'].str.replace(r'<[^<]+?>', '').str.replace(r'^b.', '')
df['tokens'] = df['tokens'].str.replace('/', ' ')
def punct_cleaning(column):
processed_data = []
for i in column:
text = re.sub('[%s]' % string.punctuation, '', i).lower()
processed_data.append(text)
return pd.DataFrame(processed_data)
df['tokens'] = punct_cleaning(df['tokens'])
df['tokens'][0]
df['lemmas'] = df['tokens'].apply(lambda text:
[token.lemma_ for token in nlp(text) if (token.is_stop != True) and
(token.is_punct != True)])
df['lemmas'] = df['lemmas'].str.join(' ')
df['lemmas']
sns.distplot(doc_len)
plt.show()
def count(docs):
word_counts = Counter()
appears_in = Counter()
total_docs = len(docs)
for doc in docs:
word_counts.update(doc)
appears_in.update(set(doc))
temp = zip(word_counts.keys(), word_counts.values())
wc = pd.DataFrame(temp, columns = ['word', 'count'])
wc['rank'] = wc['count'].rank(method='first', ascending=False)
total = wc['count'].sum()
wc['pct_total'] = wc['count'].apply(lambda x: x / total)
wc = wc.sort_values(by='rank')
wc['cul_pct_total'] = wc['pct_total'].cumsum()
t2 = zip(appears_in.keys(), appears_in.values())
ac = pd.DataFrame(t2, columns=['word', 'appears_in'])
wc = ac.merge(wc, on='word')
wc['appears_in_pct'] = wc['appears_in'].apply(lambda x: x / total_docs)
return wc.sort_values(by='rank')
wc = count(df['tokens'])
wc.head()
# cumlative distribution plot
sns.lineplot(x='rank', y='cul_pct_total', data=wc)
# Frequency of apperances
sns.distplot(wc['appears_in_pct'])
plt.show()
| Med-Cabinet-2/DS-ML-Engineering-/Notebooks/Med_Cab_Text_Data_1.3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
os.environ['TF_CPP_MIN_LOG_LEVEL']='2'
import numpy as np
import tensorflow as tf
import time
import utils
# -
# Define paramaters for the model
learning_rate = 0.01
batch_size = 128
n_epochs = 30
n_train = 60000
n_test = 10000
# Step 1: Read in data
mnist_folder = 'data/mnist'
utils.download_mnist(mnist_folder)
train, val, test = utils.read_mnist(mnist_folder, flatten=True)
# Step 2: Create datasets and iterator
# create training Dataset and batch it
train_data = tf.data.Dataset.from_tensor_slices(train)
train_data = train_data.shuffle(10000) # if you want to shuffle your data
train_data = train_data.batch(batch_size)
# create testing Dataset and batch it
test_data = tf.data.Dataset.from_tensor_slices(test)
test_data = test_data.batch(batch_size)
# +
# create one iterator and initialize it with different datasets
iterator = tf.data.Iterator.from_structure(train_data.output_types,
train_data.output_shapes)
img, label = iterator.get_next()
train_init = iterator.make_initializer(train_data) # initializer for train_data
test_init = iterator.make_initializer(test_data) # initializer for train_data
# -
# x: inputs
x = tf.placeholder(tf.float32, name='x', shape=[None, 784])
# y: labels
y = tf.placeholder(tf.int32, name='y', shape=[None])
# Step 3: create weights and bias
# w is initialized to random variables with mean of 0, stddev of 0.01
# b is initialized to 0
# shape of w depends on the dimension of X and Y so that Y = tf.matmul(X, w)
# shape of b depends on Y
w = tf.get_variable(name='w', shape =[784, 10], initializer= tf.random_normal_initializer())
b = tf.get_variable(name='b', shape=[10], initializer = tf.zeros_initializer())
# Step 4: build model
# the model that returns the logits.
# this logits will be later passed through softmax layer
logits = tf.matmul(x, w) +b
# Step 5: define loss function
# use cross entropy of softmax of logits as the loss function
entropy = tf.nn.softmax_cross_entropy_with_logits(labels=label, logits=logits)
loss= tf.reduce_mean(entropy)
# Step 6: define optimizer
# using Adamn Optimizer with pre-defined learning rate to minimize loss
optimizer = tf.train.AdamOptimizer(learning_rate=0.01).minimize(loss)
# +
# Step 7: calculate accuracy with test set
preds = tf.nn.softmax(logits)
correct_preds = tf.equal(tf.argmax(preds, 1), tf.argmax(label, 1))
accuracy = tf.reduce_sum(tf.cast(correct_preds, tf.float32))
writer = tf.summary.FileWriter('./graphs/logreg', tf.get_default_graph())
with tf.Session() as sess:
start_time = time.time()
sess.run(tf.global_variables_initializer())
# train the model n_epochs times
for i in range(n_epochs):
sess.run(train_init) # drawing samples from train_data
total_loss = 0
n_batches = 0
try:
while True:
_, l = sess.run([optimizer, loss])
total_loss += l
n_batches += 1
except tf.errors.OutOfRangeError:
pass
print('Average loss epoch {0}: {1}'.format(i, total_loss/n_batches))
print('Total time: {0} seconds'.format(time.time() - start_time))
# test the model
sess.run(test_init) # drawing samples from test_data
total_correct_preds = 0
try:
while True:
accuracy_batch = sess.run(accuracy)
total_correct_preds += accuracy_batch
except tf.errors.OutOfRangeError:
pass
print('Accuracy {0}'.format(total_correct_preds/n_test))
writer.close()
# -
| 03.basic_model/03. Logistic Regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## Enriching Data with moving statistics
# The bigquery cell magic allows for a rapid explorative development, but clutters the notebook with large result sets.
# %load_ext google.cloud.bigquery
# %%bigquery
SELECT
DEP_T, AIRLINE, ARR, DEP_DELAY, ARR_DELAY
from `going-tfx.examples.ATL_JUNE_SIGNATURE`
where date='2006-06-12'
order by dep_t limit 2
# ---
# Doing it with ```datalab.bigquery``` doesn't display the large result set
import google.datalab.bigquery as bq
samples=bq.Query("""
SELECT
DEP_T, AIRLINE, ARR, DEP_DELAY, ARR_DELAY
FROM
`going-tfx.examples.ATL_JUNE_SIGNATURE`
WHERE
date='2006-06-12'
ORDER BY
dep_t
""").execute().result().to_dataframe()
two_records = samples[:2].to_dict(orient='records')
print("%s samples, showing first 2:" % len(samples))
two_records
# ---
# ### Beam Transform ```DoFn```s
import apache_beam as beam
from apache_beam import window
from apache_beam.options.pipeline_options import PipelineOptions
# #### Add Timestamp
class AddTimeStampDoFn(beam.DoFn):
def __init__(self, offset, *args, **kwargs):
self.offset = offset
super(beam.DoFn, self).__init__(*args, **kwargs)
def process(self, element):
timestamp = (self.offset +
(element['DEP_T'] // 100) * 3600 +
(element['DEP_T'] % 100) * 60)
time_stamped = beam.window.TimestampedValue(element, timestamp)
yield time_stamped
# #### Add hour of day
class Add_HOD(beam.DoFn):
def process(self, element):
element=element.copy()
dep_t = element['DEP_T']
element['HOD'] = dep_t // 100
yield element
# #### Key out DEP_DELAY for averaging
class DEP_DELAY_by_HOD(beam.DoFn):
def process(self, element):
element=element.copy()
yield element['HOD'], element['DEP_DELAY']
# #### Key the whole records
# Keying the records allows as to CoGroupByKey after the windowed statistics are available
class Record_by_HOD(beam.DoFn):
def process(self, element):
element=element.copy()
yield element['HOD'], element
# #### Unnest
# Unnest the resulting structure coming from ```CoGroupByKey``` to simple records
class Flatten_EnrichedFN(beam.DoFn):
def process(self, element):
hod=element[0]
avg=element[1]['avg'][0]
cnt=element[1]['cnt'][0]
records=element[1]['rec'][0]
for record in records:
record['CNT_BTH']=cnt
record['AVG_DEP_DELAY_BTH']=avg
yield record
# We'll add a timestamp as if the records were all today's records
# ### Creating the pipeline
import time
import datetime
OFFSET = int(time.time() // 86400 * 86400)
OFFSET
data = samples.to_dict(orient='records')
windowed = (
data
| "Add_timestamp" >> beam.ParDo(AddTimeStampDoFn(OFFSET))
| "Add_HOD" >> beam.ParDo(Add_HOD())
| "Window_1h" >> beam.WindowInto(window.FixedWindows(3600))
)
len(windowed)
# #### Counts by the hour
records_by_hod = (
windowed
| "Record_by_HOD" >> beam.ParDo(Record_by_HOD())
| "Group_by_HOD" >> beam.GroupByKey()
)
counts_by_hod = (
records_by_hod
| "Count" >> beam.CombineValues(beam.combiners.CountCombineFn())
)
counts_by_hod[:5]
# +
#records_by_hod[2]
# -
# #### Averages by the hour
avgs_by_hod = (
windowed
| "Make_HOD" >> beam.ParDo(DEP_DELAY_by_HOD())
| "Avg_by_HOD" >> beam.CombinePerKey(beam.combiners.MeanCombineFn())
)
avgs_by_hod[:3]
# #### Co-Group and Flatten
combined = ( {'cnt': counts_by_hod, 'avg': avgs_by_hod, 'rec': records_by_hod }
| "Co_Group_HOD" >> beam.CoGroupByKey()
| "Flatten" >> beam.ParDo(Flatten_EnrichedFN())
)
combined[:3]
| kovalevskyi/Windowing_Tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# +
import os
import numpy as np
import pandas as pd
from sklearn import preprocessing
from sklearn.pipeline import Pipeline
from sklearn.linear_model import LogisticRegression
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
from sklearn.svm import SVC
from sklearn.tree import DecisionTreeClassifier
from sklearn.tree import ExtraTreeClassifier
from sklearn.naive_bayes import GaussianNB
from sklearn.neighbors import KNeighborsClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import ExtraTreesClassifier
from sklearn.model_selection import train_test_split
from sklearn.metrics import classification_report
from sklearn.metrics import f1_score
from sklearn.metrics import accuracy_score
from sklearn.metrics import confusion_matrix
import matplotlib.pyplot as plt
# %matplotlib inline
import sys
sys.path.append("..")
from gcforest.gcforest import GCForest
import cmaes as cma
# -
# # 数据以及参数
random_seed = 42
# 其余基本模型的参数都是sklearn默认的
#
# 设置深度森林参数:最大层数20,连续5层没有效果(WF1)提升则停止,每一层:4个随机森林,3个决策树,1个逻辑回归
def get_toy_config():
config = {}
ca_config = {}
ca_config["random_state"] = random_seed
ca_config["max_layers"] = 20
ca_config["early_stopping_rounds"] = 5
ca_config["n_classes"] = 6
ca_config["estimators"] = []
ca_config["estimators"].append({"n_folds": 5, "type": "RandomForestClassifier", "random_state" : random_seed})
ca_config["estimators"].append({"n_folds": 5, "type": "RandomForestClassifier", "random_state" : random_seed})
ca_config["estimators"].append({"n_folds": 5, "type": "RandomForestClassifier", "random_state" : random_seed})
ca_config["estimators"].append({"n_folds": 5, "type": "RandomForestClassifier", "random_state" : random_seed})
ca_config["estimators"].append({"n_folds": 5, "type": "DecisionTreeClassifier"})
ca_config["estimators"].append({"n_folds": 5, "type": "DecisionTreeClassifier"})
ca_config["estimators"].append({"n_folds": 5, "type": "DecisionTreeClassifier"})
ca_config["estimators"].append({"n_folds": 5, "type": "LogisticRegression"})
config["cascade"] = ca_config
return config
path = os.getcwd()+'/../data/20122018freshwater_four_feature.csv'
data = pd.read_csv(path, na_values = np.nan)
# training/valid/test: 0.6/0.2/0.2, 各数据集划分的时候要注意
X = data.drop(['本周水质'], axis=1).values # Series
y = data['本周水质'].values
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2,
stratify = y, random_state = random_seed)
# Z-score
clean_pipeline = Pipeline([('imputer', preprocessing.Imputer(missing_values='NaN',strategy="median")),
('std_scaler', preprocessing.StandardScaler()),])
X_train = clean_pipeline.fit_transform(X_train)
X_test = clean_pipeline.fit_transform(X_test)
X_train2, X_valid, y_train2, y_valid = train_test_split(X_train, y_train, test_size=0.25,
stratify = y_train, random_state = random_seed)
X_train = X_train2
y_train = y_train2
# # Base Model Pre-train
# +
config = get_toy_config()
models = [
LogisticRegression(),
LinearDiscriminantAnalysis(),
SVC(probability=True),
DecisionTreeClassifier(),
ExtraTreeClassifier(),
GaussianNB(),
KNeighborsClassifier(),
RandomForestClassifier(random_state=random_seed),
ExtraTreesClassifier(random_state=random_seed),
GCForest(config)
]
# # 除去所有表现优异的树模型,保留3个基础的机器学习模型
# models = [
# LogisticRegression(),
# LinearDiscriminantAnalysis(),
# SVC(probability=True),
# GaussianNB(),
# KNeighborsClassifier(),
# ]
# +
y_pred_proba_all = [] # 各模型权重参数
test_entries = [] # 各基模型测试集结果
train_entries = [] # 各基模型训练集结果
for model in models:
model_name = model.__class__.__name__
if model_name == 'GCForest':
model.fit_transform(X_train, y_train, X_test, y_test)
else:
model.fit(X_train, y_train)
y_train_pred = model.predict(X_train)
y_test_pred = model.predict(X_test)
y_pred_proba = model.predict_proba(X_valid) # 20%验证集上的概率向量,为训练权重做准备
f1_train = f1_score(y_train, y_train_pred, average='weighted')
f1_test = f1_score(y_test, y_test_pred, average='weighted')
acc_train = accuracy_score(y_train, y_train_pred)
acc_test = accuracy_score(y_test, y_test_pred)
train_entries.append((model_name, f1_train, acc_train))
test_entries.append((model_name, f1_test, acc_test))
y_pred_proba_all.append(y_pred_proba)
train_df = pd.DataFrame(train_entries, columns=['model_name', 'train_f1_weighted', 'train_accuracy'])
test_df = pd.DataFrame(test_entries, columns=['model_name', 'test_f1_weighted', 'test_accuracy'])
# -
print("Results on training data")
train_df
print("Results on test data")
test_df
# 深度森林训练结果:4层不再有效果提升
#
# opt_layer_num=4, weighted_f1_train=99.45%, weighted_f1_test=97.66%
#
# 60%训练集,20%的测试集仅用做展示
# # 训练权重
y_pred_proba_all = np.asarray(y_pred_proba_all)
np.save("../npy/NCE_proba_of_10models.npy", y_pred_proba_all)
# +
# 直接载入各个模型在验证集的概率向量,仅在懒得pre train的时候使用
# y_pred_proba_all = np.load("../npy/NCE_proba_of_10models.npy")
# -
# ## Train weights via NCE Ensemble on validation set
def trainNCE(X_valid, y_valid, y_pred_proba_all):
number_of_exp = 100
classifier_num = 10
population_num = 1000
retain_population_num = 100
max_iteration = 50
population_weights = np.zeros((population_num, classifier_num))
population_retain_weights = np.zeros((retain_population_num, classifier_num))
population_score = []
population_retain_score = []
all_best_weights = np.zeros((max_iteration, classifier_num)) # 某次训练时,所有迭代步骤中最好的种群的权重
all_best_f1s = np.zeros(max_iteration) # 某次训练时,每次迭代都取精英种群中最高的f1,构成这个“最高f1数组”
all_mean_f1s = np.zeros(max_iteration) # 某次训练时,每次迭代都取精英种群f1的均值,构成这个“平均f1数组”
all_best_f1s_mean = np.zeros(number_of_exp) # 每次训练最高f1数组的均值, 即 np.mean(all_best_f1s)
all_best_f1s_std = np.zeros(number_of_exp) # 每次训练最高f1数组的标准差, 即 np.std(all_best_f1s)
all_mean_f1s_mean = np.zeros(number_of_exp)
all_mean_f1s_std = np.zeros(number_of_exp)
mu = np.zeros(classifier_num)
sigma = np.ones(classifier_num)
# 在验证集集上: 训练每个基学习器的投票参数
for i in range(max_iteration):
print("Iteration: %d" %(i))
# 该次迭代的所有种群们
population_score = np.zeros(population_num)
population_weights = np.zeros((population_num, classifier_num))
# 该次迭代的优势种群们
population_retain_score = np.zeros(retain_population_num)
population_retain_weights = np.zeros((retain_population_num, classifier_num))
# 生成所有种群
for j in range(classifier_num):
w = np.random.normal(mu[j], sigma[j]+700/(i+1), population_num)
# w = np.random.normal(mu[j], sigma[j], population_num)
population_weights[:,j] = w
# 映射所有种群的权重至[0:1]
for j in range(population_num):
w2 = np.zeros(classifier_num)
for k in range(classifier_num):
w2[k] = np.exp(-population_weights[j][k]*population_weights[j][k])
# w2[k] = np.exp(population_weights[j][k])/np.sum(np.exp(population_weights[j]))
population_weights[j] = w2
# 计算所有种群得分
for j in range(population_num):
y_pred_ensemble_proba = np.zeros((len(y_valid), 6)) # 集成器概率向量
# 为每一个基学习器乘上权重
for k in range(classifier_num):
y_pred_ensemble_proba += y_pred_proba_all[k] * population_weights[j][k]
y_pred_ensemble = np.argmax(y_pred_ensemble_proba, axis=1)
f1 = f1_score(y_valid, y_pred_ensemble, average="weighted")
population_score[j] = f1
# 所有种群得分按降序排列
retain_index = np.argsort(-np.array(population_score))[:retain_population_num]
# 记录该次迭代中的优势种群们
population_retain_weights = population_weights[retain_index] # 精英样本权重
population_retain_score = np.array(population_score)[retain_index] # 精英样本得分
# 记录每次迭代最好的种群和value
all_best_weights[i] = population_retain_weights[0]
all_best_f1s[i] = population_retain_score[0] # 最高得分
all_mean_f1s[i] = np.mean(population_retain_score) # 精英样本平均得分
# 更新mu,sigma为优势种群们的分布
mu = np.mean(population_retain_weights, axis = 0)
sigma = np.std(population_retain_weights, axis = 0) #default: ddof = 0, The divisor used in calculations is N - ddof
# print("mu\n",mu)
# print("sigma\n", sigma)
# print("Weighted F1 Score after rank")
# print(population_retain_score)
# print("Weights")
# print(population_retain_weights)
# 权重训练完毕
print("==================Finish training==================")
last_weight = population_retain_weights[0] # 最后一轮迭代精英样本中具有最高F1的权重
last_f1 = all_best_f1s[-1] # 最后一轮迭代中精英样本中的最高F1
best_f1 = all_best_f1s[np.argmax(all_best_f1s)] # 所有轮迭代中最高的F1
best_weight = all_best_weights[np.argmax(all_best_f1s)] # 所有轮迭代中具有最高F1
print("Best F1 in last iteration: %f" % (last_f1))
print("Last weight in last iteration: %s" %(last_weight))
print("Best F1 in all iterations: %f" % (best_f1))
print("Best weight in all iterations: %s" %(best_weight))
print("Last mu\n", mu)
print("Last sigma\n", sigma)
plt.figure(figsize=(10,6))
plt.plot(all_best_f1s[:25], 'b', label = 'Best weighted F1 score of elite samples')
plt.plot(all_mean_f1s[:25], 'r', label = 'Mean weighted F1 score of elite samples')
plt.xlabel('Iteration')
plt.ylabel('Weighted F1 score')
plt.legend(frameon=False)
plt.grid(True)
# plt.savefig('../img/weighed_F1_iteration(1-10).eps',format='eps')
return last_weight
# ## Train weights via NCE Ensemble on validation set
def trainCMAES(X_valid, y_valid, y_pred_proba_all):
es = cma.CMAEvolutionStrategy(10 * [0], 0.5)
# help(cma)
# help(es)
score = "f1_weighted"
# score = "accuracy"
while not es.stop():
solutions = es.ask()
es.tell(solutions, [cma.ff.water_ensemble(x, y_pred_proba_all, y_valid, metric=score) for x in solutions])
es.logger.add() # write data to disc to be plotted
es.disp()
# 训练完毕
es.result_pretty() # pretty print result
cma.plot()
weights = np.exp(es.result.xbest)/np.sum(np.exp(es.result.xbest))
return weights
# ## 得到10个基模型的权重
# +
weights = np.exp(es.result.xbest)/np.sum(np.exp(es.result.xbest))
# weights = np.load("../npy/cmaes_weights.npy") # 载入保存的权重,直接载入仅在懒得训练CMAES时使用或者想得到固定的结果
# weights of each base models
print("CMAES weights", weights)
# np.save("../npy/cmaes_weights.npy", weights) # 保存结果
# -
# # 测试
# 代入权重,在3个集合上计算ACC和F1
#
# - 训练集(总样本数量的60%)X_train
# - 验证集(总样本数量的20%,调参数,用CMAES得到权重)X_valid
# - 测试集(总样本数的20%)X_test
# - 各模型稳定性基于交叉验证,在除测试集外的80%上做5折 X_train
# ### ACC
# +
# 取训练好的模型
for model in models:
model_name = model.__class__.__name__
train_pred = model.predict(X_train)
valid_pred = model.predict(X_valid)
test_pred = model.predict(X_test)
print("=================" + model_name + "=================")
train_cm = confusion_matrix(y_train, train_pred)
valid_cm = confusion_matrix(y_valid, valid_pred)
test_cm = confusion_matrix(y_test, test_pred)
i=0
train_acc_all = np.zeros(6)
for c in train_cm:
train_acc_all[i] = c[i]/np.sum(c)
print("%d train_acc: %.2f" %(i+1, 100*train_acc_all[i]))
i=i+1
print("average: %.2f" % (100*np.mean(train_acc_all)))
i=0
valid_acc_all = np.zeros(6)
for c in valid_cm:
valid_acc_all[i] = c[i]/np.sum(c)
print("%d valid_acc: %.2f" %(i+1, 100*valid_acc_all[i]))
i=i+1
print("average: %.2f" % (100*np.mean(valid_acc_all)))
i=0
test_acc_all = np.zeros(6)
for c in test_cm:
test_acc_all[i] = c[i]/np.sum(c)
print("%d test_acc: %.2f" %(i+1, 100*test_acc_all[i]))
i=i+1
print("average: %.2f" % (100*np.mean(test_acc_all)))
# - 计算CE(1-10)
population_best_weight = weights
classifier_num = 10
# 所有学习器都输出概率向量,最后投票
y_train_pred_proba_all = []
y_valid_pred_proba_all = []
y_test_pred_proba_all = []
# 取训练好的模型,计算各模型”验证集“上输出概率向量
for model in models:
train_pred_proba = model.predict_proba(X_train)
valid_pred_proba = model.predict_proba(X_valid)
test_pred_proba = model.predict_proba(X_test)
y_train_pred_proba_all.append(train_pred_proba)
y_valid_pred_proba_all.append(valid_pred_proba)
y_test_pred_proba_all.append(test_pred_proba)
y_train_pred_ensemble_proba = np.zeros((len(y_train), 6)) # 初始化集成器概率向量
y_valid_pred_ensemble_proba = np.zeros((len(y_valid), 6)) # 初始化集成器概率向量
y_test_pred_ensemble_proba = np.zeros((len(y_test), 6)) # 初始化集成器概率向量
# 为每一个基学习器乘上权重
for k in range(classifier_num):
y_train_pred_ensemble_proba += y_train_pred_proba_all[k] * population_best_weight[k]
y_valid_pred_ensemble_proba += y_valid_pred_proba_all[k] * population_best_weight[k]
y_test_pred_ensemble_proba += y_test_pred_proba_all[k] * population_best_weight[k]
y_train_pred_ensemble = np.argmax(y_train_pred_ensemble_proba, axis=1)
y_valid_pred_ensemble = np.argmax(y_valid_pred_ensemble_proba, axis=1)
y_test_pred_ensemble = np.argmax(y_test_pred_ensemble_proba, axis=1)
# 计算各水质等级的得分
print("=================CMAES=================")
train_cm = confusion_matrix(y_train, y_train_pred_ensemble)
valid_cm = confusion_matrix(y_valid, y_valid_pred_ensemble)
test_cm = confusion_matrix(y_test, y_test_pred_ensemble)
i=0
train_acc_all = np.zeros(6)
for c in train_cm:
train_acc_all[i] = c[i]/np.sum(c)
print("%d train_acc: %.2f" %(i+1, 100*train_acc_all[i]))
i=i+1
print("average: %.2f" % (100*np.mean(train_acc_all)))
i=0
valid_acc_all = np.zeros(6)
for c in valid_cm:
valid_acc_all[i] = c[i]/np.sum(c)
print("%d valid_acc: %.2f" %(i+1, 100*valid_acc_all[i]))
i=i+1
print("average: %.2f" % (100*np.mean(valid_acc_all)))
i=0
test_acc_all = np.zeros(6)
for c in test_cm:
test_acc_all[i] = c[i]/np.sum(c)
print("%d test_acc: %.2f" %(i+1, 100*test_acc_all[i]))
i=i+1
print("average: %.2f" % (100*np.mean(test_acc_all)))
# -
# ### F1
# +
# 所有学习器都输出概率向量,最后投票
y_train_pred_proba_all = []
y_valid_pred_proba_all = []
y_test_pred_proba_all = []
# 取训练好的模型,计算各模型”验证集“上输出概率向量
for model in models:
model_name = model.__class__.__name__
train_pred_proba = model.predict_proba(X_train)
valid_pred_proba = model.predict_proba(X_valid)
test_pred_proba = model.predict_proba(X_test)
train_pred = model.predict(X_train)
valid_pred = model.predict(X_valid)
test_pred = model.predict(X_test)
print("=================" + model_name + "=================")
print(classification_report(y_train, train_pred, digits=4))
print(classification_report(y_valid, valid_pred, digits=4))
print(classification_report(y_test, test_pred, digits=4))
y_train_pred_proba_all.append(train_pred_proba)
y_valid_pred_proba_all.append(valid_pred_proba)
y_test_pred_proba_all.append(test_pred_proba)
y_train_pred_ensemble_proba = np.zeros((len(y_train), 6)) # 初始化集成器概率向量
y_valid_pred_ensemble_proba = np.zeros((len(y_valid), 6)) # 初始化集成器概率向量
y_test_pred_ensemble_proba = np.zeros((len(y_test), 6)) # 初始化集成器概率向量
# 为每一个基学习器乘上权重
for k in range(classifier_num):
y_train_pred_ensemble_proba += y_train_pred_proba_all[k] * population_best_weight[k]
y_valid_pred_ensemble_proba += y_valid_pred_proba_all[k] * population_best_weight[k]
y_test_pred_ensemble_proba += y_test_pred_proba_all[k] * population_best_weight[k]
y_train_pred_ensemble = np.argmax(y_train_pred_ensemble_proba, axis=1)
y_valid_pred_ensemble = np.argmax(y_valid_pred_ensemble_proba, axis=1)
y_test_pred_ensemble = np.argmax(y_test_pred_ensemble_proba, axis=1)
# 计算各水质等级的得分
print("=================CMAES=================")
print(classification_report(y_train, y_train_pred_ensemble, digits=4))
print(classification_report(y_valid, y_valid_pred_ensemble, digits=4))
print(classification_report(y_test, y_test_pred_ensemble, digits=4))
# -
# 人为设定了10个参数的初值x0=10*[0],sigma0=0.5, 算法包中对sigma0的描述为:
#
# http://cma.gforge.inria.fr/apidocs-pycma/cma.evolution_strategy.CMAEvolutionStrategy.html
#
# > initial standard deviation. The problem variables should have been scaled, such that a single standard deviation on all variables is useful and the optimum is expected to lie within about x0 +- 3*sigma0. See also options scaling_of_variables. Often one wants to check for solutions close to the initial point. This allows, for example, for an easier check of consistency of the objective function and its interfacing with the optimizer. In this case, a much smaller sigma0 is advisable.
#
# CMAES 知乎算法专栏
# https://zhuanlan.zhihu.com/p/31193293
# +
# 算法默认参数
cma.CMAOptions.defaults()
| notebook/NCE_10.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.5 ('siena_eeg_ecg')
# language: python
# name: python3
# ---
# # Exploração de carcterísticas
# +
import os
from os.path import join, dirname
from dotenv import load_dotenv
dotenv_path = join(dirname('__file__'), '.env')
load_dotenv(dotenv_path)
ROOT_PATH = os.environ.get("ROOT_PATH")
# -
import pandas as pd
import matplotlib.pyplot as plt
from matplotlib.gridspec import GridSpec
import seaborn as sns
import numpy as np
df = pd.read_parquet(f"{ROOT_PATH}/features/features.parquet")
df = df.astype({
'var': float,
'skew': float,
'kur': float,
'label': str
})
# ## Agrupamento Geral
# +
fig = plt.figure(figsize=(7, 14))
gs = GridSpec(nrows=3, ncols=1)
# Boxplots
ax0 = fig.add_subplot(gs[0])
g1 = sns.boxplot(data=df, x='label',y='var',ax=ax0, showfliers=False)
g1.set(xticklabels=['Ictal','Pré-ictal','Pós-ictal','Normal','Recuperação'])
g1.set_ylabel('Variance + Log Entropy', size=15)
g1.set_xlabel('')
ax1 = fig.add_subplot(gs[1])
g2 = sns.boxplot(data=df, x='label',y='kur',ax=ax1, showfliers=False)
g2.set(xticklabels=['Ictal','Pré-ictal','Pós-ictal','Normal','Recuperação'])
g2.set_ylabel('Kurtosis + Log Entropy', size=15)
g2.set_xlabel('')
ax2 = fig.add_subplot(gs[2])
g3 = sns.boxplot(data=df, x='label',y='skew',ax=ax2, showfliers=False)
g3.set(xticklabels=['Ictal','Pré-ictal','Pós-ictal','Normal','Recuperação'])
g3.set_ylabel('Skewness + Log Entropy', size=15)
g3.set_xlabel('')
# -
# ## Ampliando Grupos
# +
df1 = df[df['label'].isin(['normal','rep'])]
df2 = df[df['label'].isin(['pre','pos'])]
df3 = df[df['label']=='ictal']
df1['type'] = np.repeat('normal_rep', len(df1))
df2['type'] = np.repeat('pre_pos', len(df2))
df3['type'] = np.repeat('ictal', len(df3))
df = pd.concat([df1,df2,df3])
# +
fig = plt.figure(figsize=(7, 14))
gs = GridSpec(nrows=3, ncols=1)
# Boxplots
ax0 = fig.add_subplot(gs[0])
g1 = sns.boxplot(data=df, x='type',y='var',ax=ax0)
g1.set_xlabel('')
ax1 = fig.add_subplot(gs[1])
sns.kdeplot(df[df['type']=='ictal']['var'].to_numpy(),label='Ictal', ax=ax1)
sns.kdeplot(df[df['type']== 'normal_rep']['var'].to_numpy(), label='pre-pos', ax=ax1)
sns.kdeplot(df[df['type']== 'pre_pos']['var'].to_numpy(), label='normal-rep', ax=ax1)
ax1.legend()
| notebooks/04_feature_analytics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# [View in Colaboratory](https://colab.research.google.com/github/abhigoogol/Planet-understanding-the-amazon-from-space-Multilabel-classification/blob/master/Planet_Understanding_Multi_label_classification.ipynb)
# + [markdown] id="wr6StYO5NWzB" colab_type="text"
# # Installation
# + id="thEf4l5AvOD4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 2903} outputId="83435c6c-12f0-4891-cc4f-5ab389a21baa"
# !apt update && apt install -y libsm6 libxext6
# !pip3 install http://download.pytorch.org/whl/cu80/torch-0.3.0.post4-cp36-cp36m-linux_x86_64.whl
# !pip install opencv-python
# !pip install fastai
# !pip3 install torchvision
# + id="lpxgaCVPDMRd" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 317} outputId="d7c36473-4915-4196-ccb7-97823c55a948"
# !pip install kaggle
# !mkdir .kaggle
# !ls .kaggle
# !echo '{"username": "abhisheksambyal", "key": "c6959ce81f26c06e28938ae290088cbe", "path": "/content/"}' > .kaggle/kaggle.json
# !cat .kaggle/kaggle.json
# !chmod 600 /content/.kaggle/kaggle.json
# + id="mqYoxjXWE0KI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 517} outputId="3a6f4bd3-0430-4a17-8c38-b92d312d7837"
# !kaggle competitions download -c planet-understanding-the-amazon-from-space
# + id="s9WwefC8BOuS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 417} outputId="583d3d9e-7dde-4a6c-cbd5-617fb4a8f309"
# !apt-get install p7zip-full
# + [markdown] id="-qZ8eS8HPa6z" colab_type="text"
# # Extracting Dataset
# + id="YIPdrv8rGmhr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 167} outputId="d1f0821f-e067-4589-d265-3af2ee4c88d7"
# # !ls competitions/planet-understanding-the-amazon-from-space
# # !ls competitions/planet-understanding-the-amazon-from-space/*.zip
# !unzip -q competitions/planet-understanding-the-amazon-from-space/\*.zip -d competitions/planet-understanding-the-amazon-from-space
# !ls competitions/planet-understanding-the-amazon-from-space
# + id="TX3By3F3JS4V" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 50} outputId="887596f4-0837-4982-ba76-8a3456f25f36"
# # !ls competitions/planet-understanding-the-amazon-from-space/*.7z
# # !ls competitions/planet-understanding-the-amazon-from-space/*jpg*.7z
# !ls competitions/planet-understanding-the-amazon-from-space/*tif*.7z
# + id="XT0SPW4hCcZq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 967} outputId="e15f204a-725a-45aa-a2eb-8820685d291b"
!7za x competitions/planet-understanding-the-amazon-from-space/\*jpg*.7z -ocompetitions/planet-understanding-the-amazon-from-space
# !ls competitions/planet-understanding-the-amazon-from-space
# + id="_o_c8b8cLJsu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 67} outputId="994ca14a-a361-4f9c-d533-1d2f9bb2b7f6"
# # !ls competitions/planet-understanding-the-amazon-from-space/*.tar
# !find competitions/planet-understanding-the-amazon-from-space/ -type f -name "*.tar"
# + id="IRwKwwIICgTD" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 183} outputId="760cda21-9cbc-4940-9d57-b2e8f80bc4de"
# !find competitions/planet-understanding-the-amazon-from-space/ -type f -name "*.tar" -exec tar xvf {} -C competitions/planet-understanding-the-amazon-from-space/ \;
# !ls competitions/planet-understanding-the-amazon-from-space
# + id="79Z-DHQ6IDoA" colab_type="code" colab={}
# !cp -rv competitions/planetunderstandingtheamazonfromspace/test-jpg-additional/. competitions/planetunderstandingtheamazonfromspace/test-jpg/
# + id="ZzPQykInGw4Y" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 33} outputId="1fd3d751-a06a-46cb-e276-b6182b52f172"
# !ls competitions/planetunderstandingtheamazonfromspace/test-jpg
# + id="R_rsSZHfHF1R" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 50} outputId="bb9ceafa-b5c3-443a-ac47-5fa1db1f76b8"
# !find competitions/planetunderstandingtheamazonfromspace/ -iname 'test-jpg-additional'
# + id="zgU8Acl6F0EU" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 33} outputId="26129293-c6f3-40db-f94b-439783f65b68"
# !ls competitions/planetunderstandingtheamazonfromspace/test-jpg/ | wc -l
# + id="6N8s_dbQGG1m" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 33} outputId="454f18ce-4476-4af8-b818-def2a9998769"
# !ls competitions/planetunderstandingtheamazonfromspace/test-jpg-additional/ | wc -l
# + id="3XdjVKzUGGuV" colab_type="code" colab={}
# + id="1yaWGy0nGGnx" colab_type="code" colab={}
# + [markdown] id="MvH20g4jPqE5" colab_type="text"
# # Training
# + id="kA1-wWjuf40_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 200} outputId="ddd08e08-bd8f-4eac-d426-16a04377afb9"
# # !touch __init__.py
# # !touch competitions/__init__.py
# # !touch competitions/planet-understanding-the-amazon-from-space/__init__.py
# !wget https://raw.githubusercontent.com/fastai/fastai/master/courses/dl1/planet.py
# # !mv competitions/planet-understanding-the-amazon-from-space/ competitions/planetunderstandingtheamazonfromspace/
# + id="eRlTILlZljkY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 533} outputId="9277021b-70dc-47b4-c3d1-9db64ad54825"
# !cat planet.py
# + id="DRFmUaWLhLir" colab_type="code" colab={}
# !mv planet.py competitions/planetunderstandingtheamazonfromspace/
# + id="0VELW_w7Pi0y" colab_type="code" colab={}
from fastai.conv_learner import *
# + id="CFgZANDjP-Fb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 233} outputId="e987dad8-b4d5-4b32-9856-cd0ed03003b6"
# !ls competitions/planetunderstandingtheamazonfromspace/
PATH = 'competitions/planetunderstandingtheamazonfromspace/'
# + id="M25AIKUIP-CQ" colab_type="code" colab={}
from competitions.planetunderstandingtheamazonfromspace.planet import f2
metris = [f2]
f_model = resnet34
# + id="cAUxg5rAP94g" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 33} outputId="9970a06d-1a5a-4ee1-a9d9-312e5a9e5f85"
label_csv = f'{PATH}/train_v2.csv'
n = len(list(open(label_csv))) - 1
val_idxs = get_cv_idxs(n)
len(val_idxs)
# + id="JdHzl_OfP91Q" colab_type="code" colab={}
def get_data(sz):
tfms = tfms_from_model(f_model, sz, aug_tfms=transforms_top_down, max_zoom=1.05)
return ImageClassifierData.from_csv(PATH, 'train-jpg', label_csv, tfms=tfms,
suffix='.jpg', val_idxs=val_idxs, test_name='test-jpg')
# + id="VDi_ohF2WGsH" colab_type="code" colab={}
sz=64
data = get_data(sz)
# + id="70H82WOyes7W" colab_type="code" colab={}
learn = ConvLearner.pretrained(f_model, data)
# + id="JuhPtpysFP3v" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 33} outputId="ab23c9bc-3bba-498d-e175-348f88f2eeed"
len(data.test_ds.fnames)
# + id="c94IHVBrVCWR" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 615} outputId="998cc8ca-ae68-406e-d638-f3a458b72835"
lrf=learn.lr_find()
learn.sched.plot()
lr = 0.2
learn.fit(lr, 3, cycle_len=1, cycle_mult=2)
# + id="csb1gYn_VCTq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 200} outputId="325f84e7-36c7-46bd-839e-ae0cd78c39de"
lrs = np.array([lr/9, lr/3, lr])
learn.unfreeze()
learn.fit(lrs, 3, cycle_len=1, cycle_mult=2)
# + id="A8HOoni7VCRS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 347} outputId="ef144b4d-71c3-49e5-97f3-6ca7cfe58b30"
learn.save(f'{sz}')
learn.sched.plot_loss()
# + id="NiFv4Eo8VCOB" colab_type="code" colab={}
# + id="N7H2QO71VCMF" colab_type="code" colab={}
sz = 128
# + id="q8Tez3V1VCJw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 200} outputId="11561b66-bf2b-4542-ecc6-fa4b44430414"
learn.set_data(get_data(sz))
learn.freeze()
learn.fit(lr, 3, cycle_len=1, cycle_mult=2)
# + id="P9K79pgaVCHH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 200} outputId="96f14936-4491-42dc-dae3-bf2de683a262"
learn.unfreeze()
learn.fit(lrs, 3, cycle_len=1, cycle_mult=2)
# + id="aioQEtuyVCEX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 347} outputId="ef8612f4-5d86-48fb-d4fc-8ac56c7a4c18"
learn.save(f'{sz}')
learn.sched.plot_loss()
# + id="3pEiMBj_P9pb" colab_type="code" colab={}
# + id="7FoJls5oP9nE" colab_type="code" colab={}
sz = 256
# + id="j7blUuFRP9kU" colab_type="code" colab={}
learn.set_data(get_data(sz))
learn.freeze()
learn.fit(lr, 3, cycle_len=1, cycle_mult=2)
# + id="tKMIk8VEnfFr" colab_type="code" colab={}
learn.unfreeze()
learn.fit(lrs, 1, cycle_len=1, cycle_mult=1)
learn.save(f'{sz}')
learn.load(f'{sz}')
# + [markdown] id="05oiCMqKn0Kp" colab_type="text"
# # Prediction
# + id="-aqzo4d1n2AR" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 17} outputId="f7addfa1-b9bd-46ec-d14d-ca8ba0deb4e4"
multi_preds, y = learn.TTA(is_test=True)
preds = np.mean(multi_preds, 0)
# + id="jQwdqquqsmPp" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 67} outputId="64826921-7334-4cbb-f86e-72dafcd969bc"
print(multi_preds.shape)
print(y.shape)
print(preds.shape)
# + id="ejvoqJATn19Q" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 33} outputId="ab04aa58-b5ab-454c-8662-1b4e63971e33"
f2(preds,y)
# + id="XFbtj09Fn17E" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 50} outputId="0160e462-76d2-4eba-e42e-1064e30a6828"
preds[0]
# + id="sf0BEPMsn14m" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 33} outputId="915180c8-2dfe-49e7-e6df-2a59e8d995d8"
len(data.test_ds.fnames)
# + id="bdAF7oE7D2B-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 33} outputId="70de1cb5-e7f3-48e7-fb45-667be00f440b"
# !ls competitions/planetunderstandingtheamazonfromspace/test-jpg/ | wc -l
# + id="9mrzjUE5D1cO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 33} outputId="86a8aeb2-9bc5-4c71-8843-069fcb8992f7"
# !ls competitions/planetunderstandingtheamazonfromspace/test-jpg-additional/ | wc -l
# + [markdown] id="WeapfCUUUv0a" colab_type="text"
# # Submission
# + id="OKHBzh687t3p" colab_type="code" colab={}
classes = np.array(data.classes, dtype=str)
res = [" ".join(classes[np.where(pp > 0.5)]) for pp in preds]
test_fnames = [os.path.basename(f).split(".")[0] for f in data.test_ds.fnames]
test_df = pd.DataFrame(res, index=test_fnames, columns=['tags'])
test_df.to_csv('planet-amazon-from-the-space1.csv', index_label='image_name')
# + id="Dr0ZHSkoXImN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 183} outputId="21606341-4aa4-4ce3-af48-91dfcb26002d"
# !head planet-amazon-from-the-space1.csv
# + id="SEVp0DQoYLFf" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 33} outputId="38c47155-6787-4fd2-fada-95d2e6451e65"
# !kaggle competitions submit -c planet-understanding-the-amazon-from-space -f planet-amazon-from-the-space1.csv -m "First Submission"
# + id="ZrwF5cOVPfja" colab_type="code" colab={}
| Planet_Understanding_Multi_label_classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.6 64-bit (''base'': conda)'
# name: python3
# ---
# # Chapter 3: Probability
# ### Guided Practice 3.6
#
# Another random process can be random selecting a winner from a set of partecipants of a TV show. If we suppose the process if purely random, the probability of selecting a generic individual numbered $k$ is $P(k) = \frac{1}{n}$ where $n$ is the number of partecipants.
# ### Guided Practice 3.7
#
# * (a) They are disjoint cause when we roll a dice we only get one face and not multiple faces at the same time.
# * (b) The addition rule tells us that $P(1) + P(4) + P(5) = \frac{1}{3}$.
# ### Guided Practice 3.8
#
# To answer these questions it is best if we use a script.
# +
import pandas as pd
from pathlib import Path
loans_path = Path("../datasets/loans_full_schema.csv")
loans_df = pd.read_csv(loans_path)
# -
# Let's now see the variable `homeownership`.
loans_df["homeownership"].unique()
# * (a) Of course we can say they are disjoint (they can't happen at the same time).
#
# Now let's define the proportions.
print(loans_df[loans_df["homeownership"] == "MORTGAGE"].shape[0] / loans_df.shape[0])
print(loans_df[loans_df["homeownership"] == "OWN"].shape[0] / loans_df.shape[0])
print(loans_df[loans_df["homeownership"] == "MORTGAGE"].shape[0] / loans_df.shape[0] + loans_df[loans_df["homeownership"] == "OWN"].shape[0] / loans_df.shape[0])
# * (b) We have `MORTGAGE` proportion equal to 0.4789 and `OWN` proportion equal to 0.1353.
# * (c) For the addition rule we have the probability of random selecting one of the `homeownership` above as 0.6142.
#
# Let's test point (c) by sampling 1000 random loans and see the proportion of those who are either one or the other.
#
rand_sample = loans_df.sample(n=1000)
print(rand_sample[rand_sample["homeownership"].isin(["MORTGAGE", "OWN"])].shape[0] / rand_sample.shape[0])
# The result is pretty coherent with the computed probability.
# ### Guided Practice 3.9
#
# * (a) Since $P(A) = P(1\ or\ 2)$ we have $P(A) = P(1) + P(2) = \frac{1}{6} + \frac{1}{6} = \frac{1}{3}$.
# * (b) For the same reason as above, $P(B) = P(4) + P(6) = \frac{1}{3}$.
# ### Guided Practice 3.10
#
# * (a) Event D represent the outcome of getting either 2 or 3.
# * (b) Yes.
# * (c) No.
# ### Guided Practice 3.11
#
# We can compute the probability of A or B occurring as:
# $$
# P(A\ or\ B) = P(A) + P(B) = P(1) + P(2) + P(4) + P(6) = \frac{4}{6} = 0.67
# $$
# ### Guided Practice 3.12
#
# * (a) $\frac{13}{52} = \frac{1}{4} = 0.25$
# * (b) $\frac{12}{52} = 0.231$
# ### Guided Practice 3.13
#
# * (a) There would not be overlapping between the two Venn Diagrams representing A and B.
# * (b) If $P(A\ and\ B) = 0$ then $P(A\ or\ B) = P(A) + P(B)$.
# ### Guided Practice 3.14
# +
from matplotlib_venn import venn2
joint = loans_df[loans_df["application_type"] == "joint"]
mortgage = loans_df[loans_df["homeownership"] == "MORTGAGE"]
j_set = set(joint.index.values.tolist())
m_set = set(mortgage.index.values.tolist())
ax = venn2([j_set, m_set], ["joint", "MORTGAGE"])
# -
# ### Guided Practice 3.16
#
# Distribution (c) seems the correct one. Distribution (a) does not sum to 1, while distribution (b) has a negative value (-0.27) which does not represent any probability.
# ### Guided Practice 3.17
#
# * (a) $P(D^{c}) = P(1\ or\ 4\ or\ 5\ or\ 6) = 0.667$
# * (b) $P(D^{c}) + P(D) = 1$
# ### Guided Practice 3.18
#
# * (a) $A^{c} = \{3, 4, 5, 6\}$ and $B^{c} = \{1, 2, 3, 5\}$.
# * (b) $P(A^{c}) = P(B^{c}) = 0.667$.
# * (c) They both are 1.
# ### Guided Practice 3.19
#
# * (a) Event $A^{c}$ is the event that both dice give 6 ($6 + 6 = 12$). It is $P(6\ and\ 6)$.
# * (b) $P(A^{c}) = 0.028$.
# * (c) $P(A) = 1 - P(A^{c}) = 0.972$.
# ### Guided Practice 3.22
#
# * (a) The probability follows the definition of independent probability $0.09 * 0.09 = 0.081$.
# * (b) $(1 - 0.09) * (1 - 0.09) = 0.83$.
# ### Guided Practice 3.23
#
# * (a) $(1 - 0.09)^{5} = 0.62$
# * (b) $0.09^{5} = 0.000009$
# * (c) $1 - (1 - 0.09)^{5} = 0.38$
# ### Guided Practice 3.24
#
# * (a) It is still $0.455$.
# * (b) It is $0.455^{2} = 0.207$
# * (c) It is $0.045$.
# * (d) It is $0.045 * 0.207 = 0.009$
# ### Exercise 3.1 - True or false.
#
# * (a) The probability won't change even if the sequence of events seen is very unlike.
# * (b) They're not mutually exclusive since a face card can be red.
# * (c) They are indeed disjoint since ace is a type, like the three face cards are different types.
# ### Exercise 3.2 - Roulette wheel.
#
# * (a) The probability is still $\frac{18}{38} = 0.474$.
# * (b) Same as above.
# * (c) I am very confident since each spin is independent from the other spins, and therefore in way probability will change.
# ### Exercise 3.3 - Four games, one winner.
#
# * (a) In this version, I would flip a coin 10 times, since I don't want to reach the stationary proportion of heads (around 0.5), which will be reached when we flip a coin 100 times. At 10 flips the proportion of heads is still fluctuating and if we get 6 heads and 4 tails, we have reached the minimum amount to win. It's clearly easier than reaching 60 heads and 40 tails.
# * (b) Here I will flip the coin 100 times, since we know that the probability of tossing a head is 0.5. At 100 flips, we will definitely reach a value greater than 0.40.
# * (c) As above, I would flip 100 coins, so I will reach a proportion very close to 0.5, and thus in our range.
# * (d) Here I will flip only 10 coins and if I am lucky I will only get 3 or less heads, which will make me win.
# ### Exercise 3.4 - Backgammon.
#
# This comes from the probability of two disjoint events happening. Since the single outcome of each dice in independent from the outcome of the other, we have two disjoint events. Any combination is equally possible, with a probability of $\frac{1}{36}$, so the combinations gotten by the two players are equally possible, and purely random.
# ### Exercise 3.5 - Coin flips.
#
# * (a) The probability of getting all tails is $(\frac{1}{2})^{10} = 0.000976$
# * (b) Same as above.
# * (c) The probability of getting at least one tail is $1 - (\frac{1}{2})^{10} = 0.9990$
# ### Exercise 3.6 - Dice rolls.
#
# * (a) Since the minimum value in a dice is 1, rolling two dices will always give a number from 2 to 12, so the probability of getting the sum of the two to 1 is 0. However let's create a script to automate the calculation.
# +
from itertools import product
def calculate_prob_sum(s):
comb = list(product(range(1, 7), repeat=2))
return len([p for p in comb if p[0] + p[1] == s]) / len(comb)
for num in [1, 5, 12]:
print(f"The probability of getting a sum of {num} is {calculate_prob_sum(num)}!")
# -
# * (b) 0.111.
# * (c) 0.028.
# ### Exercise 3.7 - Swing voters.
#
# * (a) They are not mutually exclusive events, since there is overlapping between the two categories.
# * (b) To answer this question, we better do the analysis ourself with the original dataset which can be found [here](https://www.pewresearch.org/politics/dataset/april-2012-values-survey/).
# +
from matplotlib_venn import venn2
values12_path = Path("../datasets/values12public.sav")
values12_df = pd.read_spss(values12_path)
registered = values12_df[values12_df["RV"] == "YES"]
registered = registered[["int_date", "swing", "swingdet", "party"]]
registered["swinger"] = ~registered["swingdet"].str.contains("no chance")
independent = set(registered[registered["party"] == "Independent"].index.values)
swingers = set(registered[registered["swinger"]].index.values)
ax = venn2([independent, swingers], ["Independent", "Swing Voter"])
# -
# * (c) Since 35% of voters identify themselves as independent and only 11% are both swing voters and independent, then 24% are independent but not swing voters. Let's see if this is true by querying directly into the data.
registered[(~registered["swinger"]) & (registered["party"] == "Independent")].shape[0] / registered.shape[0]
# * (d) We can calculate the probability of either one of the two events and/or the other happening using the following formula.
# $$
# P(swing\ voter\ or\ independent) = P(swing\ voter) + P(independent) - P(swing\ voter\ and\ independent) = 0.35 + 0.23 - 0.11 = 0.47 = 47\%
# $$
#
# We can test this result directly with the data.
registered[(registered["swinger"]) | (registered["party"] == "Independent")].shape[0] / registered.shape[0]
# * (e) This probability is the complement of the above calculated probability, therefore 53% of the people are neither independent nor swing voters.
# * (f) Let's calculate the probability of both the two events happening in the hypothesis they are independent.
# $$
# P(swing\ voter) * P(independent) = 0.35 * 0.23 = 0.08
# $$
# This does not equal the probability of the two events happening together, which is 0.11, therefore they are dependent. As a generale rule if $P(A) * P(B) \neq P(A\ and\ B)$ then the two events are dependent.
# ### Exercise 3.8 - Poverty and language.
#
# * (a) No, both events can happen together.
# * (b) We use synthetic data which replicates the true proportions. The probabilities are 0.146 for being below the poverty line, 0.207 for speaking a language other than English and 0.042 for both the two events happening at the same time.
# +
sample = set([i for i in range(1, 1001)])
poverty_line = set([i for i in range(1, 146)])
foreign_language = set([i for i in range(1, 43)] + [i for i in range(400, 566)])
ax = venn2([poverty_line, foreign_language], ["Below Poverty Line", "Foreign Language"], )
# -
# * (c) We can derive it from $20.7\% - 4.2\% = 16.5\%$.
# * (d) $P(poverty\ or\ foreign) = P(poverty) + P(foreign) - P(poverty\ and\ foreign) = 31.1\%$.
# * (e) $20.7\% - 4.2\% = 16.5\%$.
# * (f) We can verify it by calculating $0.146 * 0.207 = 0.030 \neq 0.402$. Therefore they're not independent.
# ### Exercise 3.9 - Disjoint vs. independent.
#
# * (a) Independent. We can both get A, but our outcome won't influence the other in any way.
# * (b) Neither, we worked together, so it makes it likely that we influence each other grade.
# * (c) No. If events are disjoint, they are dependent. If they can occur at the same time, they can be either dependent or independent.
# ### Exercise 3.10 - Guessing on an exam.
#
# * (a) $(\frac{3}{4})^{4} * \frac{1}{4} = 0.00293$.
# * (b) $(\frac{1}{4})^{5} = 0.000977$.
# * (c) $1 - (\frac{3}{4})^{5} = 0.997$.
# ### Exercise 3.11 - Educational attainment of couples.
#
# * (a) $0.16 + 0.09 = 0.25$.
# * (b) $0.26$.
# * (c) Assuming that the two events are independent (i.e. couples are randomly made), $0.25 * 0.26 = 0.065$.
# * (d) For the sake of simulation yes, although people with the same educations most likely make couples.
# ### Exercise 3.12 - School absences.
#
# * (a) $1 - 0.68 = 0.32$.
# * (b) $0.32 + 0.25 = 0.57$.
# * (c) $0.68$.
# * (d) Assuming there's no correlation between the two kids' health, $0.32 * 0.32 = 0.1024$.
# * (e) Same assumptions as above, $0.68 * 0.68 = 0.4624$.
# * (f) In point (d) assumption is legit since if one kid is healthy, the other will be healthy too. In point (e), no, since if one kid gets flu, the other has a good change of getting it too and so he has a good chance of missing school, too. Therefore, the events are not independent.
# ### Guided Practice 3.28
#
# Events are disjoint as it can also be deducted from the probability table, probabilities are non negative and sum to 1.
# ### Guided Practice 3.29
#
# * (a) $P(\texttt{mach\_learn is fashion} \mid \texttt{truth is fashion})$.
# * (b) $P(\texttt{mach\_learn is fashion} \mid \texttt{truth is fashion}) = \frac{P(\texttt{mach\_learn is fashion}\ and\ \texttt{truth is fashion})}{P(\texttt{truth is fashion})} = 0.6374$.
# ### Guided Pracrice 3.30
#
# * (a) $P(\texttt{mach\_learn is not fashion} \mid \texttt{truth is fashion}) = \frac{P(\texttt{mach\_learn is not fashion}\ and\ \texttt{truth is fashion})}{P(\texttt{truth is fashion})} = 0.3626$. Note that it is the complementary (as guessed) of the above calculated conditional probability.
# * (b) For the above reason, the sum is 1.
# * (c) I can say that the probability that the photo is fashion, in the conditional probability setting, becomes the new probability space, and therefore all the outcomes conditioned by this probability must sum to 1.
# ### Guided Practice 3.31
# $$
# P(died \mid not\ inoculated) = \frac{P(died,\ not\ inoculated)}{P(not\ inoculated)} = 0.1411
# $$
# ### Guided Practice 3.32
# $$
# P(died \mid inoculated) = \frac{P(died,\ inoculated)}{P(inoculated)} = 0.0255
# $$
# ### Guided Practice 3.33
#
# * (a) Observational since we do not have randomization.
# * (b) We can't because there might be confounding variables.
# * (c) Those who have been inoculated might be healthier to start with, and then they might also conduct a better lifestyle.
# ### Guided Practice 3.35
#
# The probability is obtained by multiplying the two probabilities: $0.03824$.
# ### Guided Practice 3.36
#
# Here it is simply a matter of subtracting the proportion from one, so we get $2.46\%$.
# ### Guided Practice 3.37
#
# It seems very effective given the abyssal difference, __but purely based on the probabilities__.
# ### Guided Practice 3.38
#
# * (a) $\frac{1}{6}$.
# * (b) $\frac{1}{6} * \frac{1}{6} = \frac{1}{36}$.
# * (c) $P(Y = 1 \mid X = 1) = \frac{P(Y = 1, X = 1)}{P(X = 1)}$, but since the outcome of Y does not influence the outcome of X in any way, $P(X = 1, Y = 1)$ is simply $P(X = 1) * P(Y = 1)$, and so $P(X = 1 \mid Y = 1) = P(X = 1) = \frac{1}{6}$.
# * (d) It is the same probability, given by the same considerations. The two outcomes are independent so the probability of both happening together is the product of the single probabilities.
# ### Guided Practice 3.39
#
# The wrong thing lies on the fact that he believes that probability decreases over time, while each spin is independent from the previous one thus the probability is still the same.
# ### Guided Practice 3.41
#
# * (a) Let's build a tree diagram.
# +
import graphviz
g = graphviz.Digraph("G")
g.node("A", "Tree")
g.node("B", "Yes")
g.node("C", "No")
g.node("D", "Passed")
g.node("E", "Failed")
g.node("F", "Passed")
g.node("G", "Failed")
g.edge("A", "B", label="78%")
g.edge("A", "C", label="22%")
g.edge("B", "D", label="97%")
g.edge("B", "E", label="3%")
g.edge("C", "F", label="57%")
g.edge("C", "G", label="43%")
g
# -
# * (b) We can compute this probability as $P(passed) = P(passed \mid tree) * P(tree) + P(passed \mid no\ tree) * P(no\ tree) = 0.882 = 88.20\%$.
# * (c) $P(tree \mid passed) = \frac{P(tree,\ passed)}{P(passed)} = \frac{0.97 * 0.78}{0.882} = \frac{0.7566}{0.882} = 0.8578$.
# ### Guided Practice 3.43
#
# Let's plot the tree diagram.
# +
g = graphviz.Digraph("G")
g.node("A", "Event")
g.node("B", "Academic")
g.node("C", "Sporting")
g.node("D", "No Event")
g.node("E", "Fill up")
g.node("F", "No fill up")
g.node("G", "Fill up")
g.node("H", "No fill up")
g.node("I", "Fill up")
g.node("L", "No fill up")
g.edge("A", "B", label="35%")
g.edge("A", "C", label="20%")
g.edge("A", "D", label="45%")
g.edge("B", "E", label="25%")
g.edge("B", "F", label="75%")
g.edge("C", "G", label="70%")
g.edge("C", "H", label="30%")
g.edge("D", "I", label="5%")
g.edge("D", "L", label="95%")
g
# -
# The probability being asked in the problem is $P(sport \mid fill\ up)$.
# $$
# P(sport \mid fillup) = \frac{P(sport,\ fill\ up)}{P(fill\ up)} = \frac{P(sport,\ fill\ up)}{P(fill\ up \mid academic)P(academic) + P(fill\ up \mid sport)P(sport) + P(fill\ up \mid no\ event)P(no\ event)} = \frac{0.20 * 0.70}{0.35 * 0.25 + 0.20 * 0.70 + 0.45 * 0.05} = 0.56 = 56.00\%
# $$
# ### Guided Practice 3.45
#
# $$
# P(A_{2},\ B) = \frac{P(B \mid A_{2}) * P(A_{2})}{P(B \mid A_{1}) * P(A_{1}) + P(B \mid A_{2}) * P(A_{2}) + P(B \mid A_{3}) * P(A_{3})} = \frac{0.0875}{0.2499} = 0.35 = 35.00\%
# $$
# ### Guided Practice 3.46
#
# $$
# P(A_{3},\ B) = \frac{P(B \mid A_{3}) * P(A_{3})}{P(B \mid A_{1}) * P(A_{1}) + P(B \mid A_{2}) * P(A_{2}) + P(B \mid A_{3}) * P(A_{3})} = \frac{0.0225}{0.2499} = 0.09 = 9.00\%
# $$
# ### Exercise 3.13 - Joint and conditional probabilities.
#
# * (a) If and only if they are independent, since the joint probability is the product of the marginal probabilities. Otherwise no.
# * (b)
# * i. $P(A\ and\ B) = P(A) * P(B) = 0.21$.
# * ii. $P(A\ or\ B) = P(A) + P(B) - P(A\ and\ B) = 1 - 0.21 = 0.79$.
# * iii. $P(A \mid B) = P(A) = 0.3$.
# * (c) No, if they were, the joint probability would be as above.
# * (d) $\frac{0.1}{0.7} = 0.1429$.
# ### Exercise 3.14 - PB & J.
#
# $$
# P(jelly \mid peanut\ butter) = \frac{P(jelly,\ peanut\ butter)}{P(peanut\ butter)} = \frac{0.78}{0.80} = 0.975
# $$
# ### Exercise 3.15 - Global warming.
#
# * (a) $P(Earth\ is\ warming,\ Liberal\ Democrat) = 0.18 \neq P(Earth\ is\ warming) * P(Liberal\ Democrat) = 0.12$, so they're not mutually exclusive.
# * (b) $P(Earth\ is\ warming\ or\ Liberal\ Democrat) = P(Earth\ is\ warming) + P(Liberal\ Democrat) - P(Earth\ is\ warming,\ Liberal\ Democrat) = 0.60 + 0.20 - 0.18 = 0.62$.
# * (c) $P(Earth\ is\ warming \mid Liberal\ Democrat) = \frac{P(Earth\ is\ warming,\ Liberal\ Democrat)}{P(Liberal\ Democrat)} = \frac{0.18}{0.20} = 0.89$.
# * (d) $P(Earth\ is\ warming \mid Conservative\ Republican) = \frac{P(Earth\ is\ warming,\ Conservative\ Republican)}{P(Conservative\ Republican)} = \frac{0.11}{0.33} = 0.33$.
# * (e) Given the huge difference between the Democrats and Republicans believing in Global Warming, I would say ideology influences this opinion.
# * (f) $P(Mod/Lib\ Republican \mid Not\ warming) = \frac{P(Mod/Lib\ Republican,\ Not\ warming)}{P(Not\ warming)} = \frac{0.06}{0.34} = 0.18$.
# ### Exercise 3.16 - Health coverage, relative frequencies.
#
# * (a) Since $P(Yes, Excellent) = 0.2099 \approx 0.2035 = P(Yes) * P(Excellent)$, I would say the two are independent events.
# * (b) $0.2329$.
# * (c) $P(Excellent \mid Yes) = \frac{P(Excellent,\ Yes)}{P(Yes)} = 0.2402$.
# * (d) $P(Excellent \mid No) = \frac{P(Excellent,\ No)}{P(No)} = 0.1823$.
# * (e) Since $P(No, Excellent) = 0.023 \approx 0.029 = P(No) * P(Excellent)$, I would say the two are independent events.
# ### Exercise 3.17 - Burger preferences.
#
# * (a) Since $\frac{6}{500} = 0.012$, they're not mutually exclusive.
# * (b) $\frac{162}{248} = 0.65$.
# * (c) $\frac{181}{252} = 0.72$.
# * (d) Assuming they don't date for the shared likings, i.e. their likings are independent (which is quite reasonable), $0.468$ is the probability.
# * (e) $0.012 + 0.504 - 0.002 = 0.514$.
# ### Exercise 3.18 - Assortative mating.
#
# * (a) Let $A_1$ the probability of the male partner having blue eyes and $A_2$ the probability of the female partner having blue eys. Then, $P(A_1\ or\ B_1) = P(A_1) + P(B_1) - P(A_1\ and\ B_1) = \frac{114}{204} + \frac{108}{204} - \frac{78}{204} = 0.7059$.
# * (b) This can be deducted from the table $\frac{78}{114} = 0.6842$.
# * (c) Same as above, respectively $\frac{19}{54} = 0.3519$ and $\frac{11}{36} = 0.3056$.
# * (d) By looking at the contingency table, it seems that most couple have the same eyes color. Therefore, they're not independent.
# ### Exercise 3.19 - Drawing box plots.
#
# * (a) Let's draw the tree.
# +
g = graphviz.Digraph("G")
g.node("A", "Box Plot")
g.node("B", "Yes")
g.node("C", "No")
g.node("D", "Passed")
g.node("E", "Failed")
g.node("F", "Passed")
g.node("G", "Failed")
g.edge("A", "B", label="80%")
g.edge("A", "C", label="20%")
g.edge("B", "D", label="86%")
g.edge("B", "E", label="14%")
g.edge("C", "F", label="65%")
g.edge("C", "G", label="35%")
g
# -
# * (b) From the graph, $P(Box\ Plot \mid Passed) = \frac{0.80 * 0.86}{0.80 * 0.86 + 0.20 * 0.60} = 0.8411 = 84.11\%$.
# ### Exercise 3.20 - Predisposition for thrombosis.
#
# Let's denote having the predisposition as event $A$, and resulting positive as event $B$. We know that $P(A) = 3\% = 0.03$, $P(B \mid A) = 99\% = 0.99$ and $P(B \mid A^C) = 1 - P(B^C \mid A^C) = 1 - 0.98 = 0.02$. Let's apply Bayes Theorem:
# $$
# P(A \mid B) = \frac{P(B \mid A) * P(A)}{P(B \mid A) * P(A) + P(B \mid A^C) * P(A^C)} = \frac{0.99 * 0.03}{0.99 * 0.03 + 0.02 * 0.97} = 0.6049
# $$
# ### Exercise 3.21 - It’s never lupus.
#
# Let's get directly to the numbers, by calculating the probability that a patient has lupus given that he/she resulted positive. Let's denote these two events respectively $A$ and $B$.
# $$
# P(A \mid B) = \frac{P(B \mid A) * P(A)}{P(B \mid A) * P(A) + P(B \mid A^C) * P(A^C)} = \frac{0.98 * 0.02}{0.98 * 0.02 + 0.26 * 0.98} = 0.0714.
# $$
# Given the low probability we got, the show is right to say it's never lupus.
# ### Exercise 3.22 - Exit poll.
# Let $A$ be the event of sampling a person with a college degree and $B$ the probability of sampling a person who voted for <NAME>.
# $$
# P(B \mid A) = \frac{P(A \mid B) * P(B)}{P(A \mid B) * P(B) + P(A \mid B^C) * P(B^C)} = \frac{0.37 * 0.53}{0.37 * 0.53 + 0.44 * 0.47} = 0.4867 = 48.67\%
# $$
# ### Guided Practice 3.49
#
# The General Multiplication Rule on the probability of not being picked on $Q_1$, then two conditional probabilities which chains on this first probability and builds a prior probability on it/them.
# ### Guided Practice 3.51
#
# $(\frac{1}{15})^3 = 0.000296$.
# ### Guided Practice 3.52
#
# * (a) First let's calculate the probability of not winning (let the probability of winning be $A$) as $P(A^C) = \frac{29}{30} * \frac{28}{29} * ...\ * \frac{23}{24} = \frac{23}{30} = 0.767$. Now $P(A) = 1 - P(A^C) = 1 - 0.767 = 0.233$.
# * (b) Same as above, but since we are sampling with replacement, each time we sample from all the 30 numbers, and so $P(A^C) = (\frac{29}{30})^7 = 0.789$. Now $P(A) = 1 - 0.789 = 0.211$.
# ### Guided Practice 3.53
#
# There's more chance of winning if we sample without replacement.
# ### Exercise 3.23 - Marbles in an urn.
#
# * (a) The probability to get the first marble blue is $P(Blue) = \frac{3}{5 + 3 + 2} = \frac{3}{10} = 0.3$.
# * (b) The probability is still $0.3$.
# * (c) The probability is still $0.3$.
# * (d) $\frac{3}{10} * \frac{3}{10} = 0.3 * 0.3 = 0.09$.
# * (e) With replacement we have independence among draws as the current draw is not affected by the previous in any way.
# ### Exercise 3.24 - Socks in a drawer.
#
# * (a) Here we are sampling without replacement, so $\frac{4}{12} * \frac{3}{11} = 0.091$.
# * (b) $\frac{7}{12} * \frac{6}{11} = 0.318$.
# * (c) $1 - (\frac{9}{12} * \frac{8}{11}) = 0.454$
# * (d) 0.
# * (e) $0.091 + 0.151 + 0.045 = 0.287$.
# ### Exercise 3.25 - Chips in a bag.
#
# * (a) $\frac{2}{9} = 0.222$.
# * (b) $\frac{3}{9} = 0.333$.
# * (c) $\frac{3}{10} * \frac{2}{9} = 0.067$.
# * (d) The draws are not independent anymore since each time we are decreasing the number of chips, so the probabilities change.
# ### Exercise 3.26 - Books on a bookshelf.
#
# * (a) $\frac{28}{95} * \frac{59}{94} = 0.185$.
# * (b) To answer this question, we have to calculate the total probability considering whether or not the fiction book drawn before is hardcover or paperback: $P(second = hardcover \mid first = fiction) = P(second = hardcover \mid first = fiction,\ hardcover) * P(first = fiction,\ hardcover) + P(second = hardcover \mid first = fiction,\ paperback) * P(first = fiction,\ paperback) = \frac{27}{94} * \frac{13}{95} + \frac{28}{94} * \frac{59}{95} = 0.224$.
# * (c) In this case, the two extractions are independent, therefore we calculate such a probability as: $P(second = hardcover \mid first = fiction) = \frac{72}{95} * \frac{28}{95} = 0.223$.
# * (d) The probability of picking a hardcover fiction book ($0.181$) is much less than picking a paperback fiction book ($0.819$), so the first is pretty insignificant relative to the second. Therefore, it is very likely that the fiction book drawn at first is also paperback.
#
# ### Exercise 3.27 - Student outfits.
#
# $$
# \frac{5}{24} * \frac{7}{23} * \frac{6}{22} = 0.017
# $$
# ### Exercise 3.28 - The birthday problem.
#
# * (a) $\frac{1}{365} * \frac{1}{365} = 0.0000007511$.
# * (b) $1 - \frac{362}{365} = 0.0082$.
# ### Guided Practice 3.55
#
# I wouldn't be surprised as that is just an expected value.
# ### Guided Practice 3.59
#
# * (a) $100\% - 25\% - 60\% = 15\%$.
# * (b) Let's render the table.
#
# |_Y_| 0 | 159 | 200 |
# |- |--- |--- |--- |
# |P| 0.15 | 0.25 | 0.60 |
#
# * (c) $159.75$.
# * (d) $69.28$.
# ### Guided Practice 3.62
#
# Such equation would simply be the difference of what she gained vs. what she spent, so $NetChange = Earned - Spent$.
# ### Guided Practice 3.63
#
# Pretty straightforwardly, $175 - 23 = 152$.
# ### Guided Practice 3.64
#
# Given that these are expected values, I do not see anything wrong if they show variability around what we calculated.
# ### Guided Practice 3.66
#
# The values is given by $6000 * 0.02 + 2000 * 0.0002 = 120.4$.
# ### Guided Practice 3.67
#
# Not at all, also because that value is relatively small.
# ### Guided Practice 3.69
#
# I think the assumption is valid since there's no correlation between each single day's commute time.
# ### Guided Practice 3.70
#
# Elena net gain equation is $\sqrt{25^2 - 8^2} = 23.69$.
# ### Exercise 3.29 - College smokers.
#
# * (a) In a random sample of 100 people, we have approximately 13 people smoking.
# * (b) People going to the gym are usually the health-aware students so they do not represent a random sample.
# ### Exercise 3.30 - Ace of clubs wins.
#
# * (a) We can use a table, and we denote with _X_ the random variable representing the card draw.
#
# | X | 0$ | $5 | $10 | $30 |
# |:---: |:---: |:---: |:---: |:---: |
# | __P__ | 0.5 | 0.25 | 0.231 | 0.019 |
#
# Expected winning is given by $E(X) = 0.5 * 0 + 0.25 * 5 + 0.231 * 10 + 30 * 0.019 = 4.13$, while standard deviation is $5.45$.
#
# * (b) Maybe $4.00, since that it how I would expect to win.
# ### Exercise 3.31 - Hearts win.
#
# * (a) As before, let's make a table to show the distribution of the _X_ random variable which represent each possible outcome of the game.
#
# | X | 0$ | $25 | $50 |
# |:---: |:---: |:---: |:---: |
# | P | 0.8621 | 0.1180 | 0.0129 |
#
# Expected winning is $3.60$ and standard deviation is $9.62$.
#
# * (b) Let _Y_ be the profit, then $E(Y) = 3.60 - 5.00 = -1.40$ and $\sqrt{VAR(Y)} = 10.84$.
# * (c) It would be very risky still, since chances to win are very low.
# ### Exercise 3.32 - Is it worth it?
#
# * (a) The distribution of Andy's profit per game is as follows.
#
# | X | -$2 | $1 | $3 | $23 |
# |:---: |:---: |:---: |:---: |:---: |
# | P | 0.6923 | 0.2308 | 0.0577 | 0.0192 |
#
# * (b) To answer this question, let's calculate the expected value and standard deviation which are $-\$0.54$ and $\$3.81$. Therefore, gain margins are very low therefore it is not an advisable way to play.
# ### Exercise 3.33 - Portfolio return.
#
# The expected return is $5\%$.
# ### Exercise 3.34 - Baggage fees.
#
# * (a) If we just don't consider those who checks more than 2 luggages, we can build the probability table.
#
# | X | $0 | $25 | $60 |
# |:---: |:---: |:---: |:---: |
# | P | 0.54 | 0.34 | 0.12 |
#
# $\mu = \$15.70$ and $\sigma = \$19.95$.
#
# * (b) If we assume that each passenger's luggage number is independent of any other passenger's luggage number, then we can simply calculate $\mu = 15.70 * 120 = \$1884$ and $\sigma = \$218.54$.
# ### Exercise 3.35 - American roulette.
#
# If it lands on red, I will get \$2, otherwise, I am going to lose (-\$1). So, the expected win is $\mu = 0.47 * 1 - 0.53 * 1 = -\$0.00563$ and $\sigma = \$0.9986$.
# ### Exercise 3.36 - European roulette.
#
# * (a) $\mu = \$1.38$ and $\sigma = \$4.50$.
# * (b) $\mu = \$0.28$ and $\sigma = \$2.28$.
# * (c) The more you play, the less is your earning margin.
# ### Guided Practice 3.73
#
# * (a) This probability, given the random sampling process, is $0.1157^3 = 0.0016$.
# * (b) This probability is $(1 - 0.1157)^3 = 0.6915$.
# ### Guided Practice 3.75
#
# By rounding to the nearest centimiter, the distribution will become discrete and so there will be some chances someone will measure that height.
# ### Exercise 3.37 - Cat weights.
#
# * (a) 41.67%.
# * (b) 13.89%.
# * (c) 38.19%.
# ### Exercise 3.38 - Income and gender.
#
# * (a) The distribution seems symmetric, maybe slightly right-skewed.
# * (b) 62.2%.
# * (c) Assuming that salary and gender are not associated, 30.48%.
# * (d) At this point, my assumption is not valid anymore since most women make less than that salary.
# ### Exercise 3.39 - Grade distributions.
# * (a) Invalid: total probability is more than 1.
# * (b) Valid: each probability is a number between 0 and 1, with the total sum of 1.
# * (c) Invalid: total probability less than 1.
# * (d) Invalid: there's a negative probability which is not possible.
# * (e) Valid, for the same reason of answer (b).
# * (f) Invalid: there's a negative probability and a single probability greater than 1.
# ### Exercise 3.40 - Health coverage, frequencies.
#
# * (a) 0.02745.
# * (b) 0.3316.
# ### Exercise 3.41 - HIV in Swaziland.
#
# Using __Bayes' Theorem__, we get a probability of 0.82.
# ### Exercise 3.42 - Twins.
#
# Applying again Bayes' Theorem we get 0.4615.
# ### Exercise 3.43 - Cost of breakfast.
#
# * (a) $\mu = \$1.40 + \$2.50 = \$3.90$ and $\sigma = \$0.34$.
# * (b) $\mu = \$3.90 * 7 = \$27.3$ and $\sigma = \$0.90$.
# ### Exercise 3.44 - Scooping ice cream.
#
# * (a) A total of $48 + 6 = 54$ ice creams are served, with a standard deviation of 1.
# * (b) $\mu = 46$ and $\sigma = 1$.
# * (c) Because, for how variances are defined, they are meant to be positive.
# ### Exercise 3.45 - Variance of a mean, Part I.
#
# $$
# (\frac{1}{2})^2 * Var(X_1) + (\frac{1}{2})^2 * Var(X_2)
# $$
# ### Exercise 3.46 - Variance of a mean, Part II.
#
# $$
# (\frac{1}{3})^2 * Var(X_1) + (\frac{1}{3})^2 * Var(X_2) + (\frac{1}{3})^2 * Var(X_3)
# $$
# ### Exercise 3.47 - Variance of a mean, Part III.
#
# $$
# (\frac{1}{n})^2 * Var(X_1) + (\frac{1}{n})^2 * Var(X_2) +\ ...\ + (\frac{1}{n})^2 * Var(X_n)
# $$
#
| chapter3/chapter3_walkthrough.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Dropping New York and '0-17' age group
# ### Requires reconstructing dataframes
#
# This is a copy of the similarly-named notebook.
# +
import pandas as pd
import numpy as np
import datetime
from datetime import timedelta
import statsmodels.api as sm
import scipy.stats as scs
import matplotlib.pyplot as plt
import plotly.express as px
import warnings
warnings.filterwarnings('ignore')
# -
# ## COVID By Age Analysis
# Define a global variable list for the age groupings to be used throughout this notebook.
age_groups_list = ['0_17', '18_49', '50_64', '65_plus']
short_age_groups_list = ['18_49', '50_64', '65_plus']
# Let's import seroprevalence by age for each Round.
# +
sp_df = pd.read_csv('../data/Nationwide_Commercial_Laboratory_Seroprevalence_Survey_December.csv')[
['Site','Date Range of Specimen Collection', 'Round', 'Catchment population',
'n [0-17 Years Prevalence]', 'n [18-49 Years Prevalence]',
'n [50-64 Years Prevalence]', 'n [65+ Years Prevalence]',
'Rate (%) [0-17 Years Prevalence]', 'Rate (%) [18-49 Years Prevalence]',
'Rate (%) [50-64 Years Prevalence]', 'Rate (%) [65+ Years Prevalence]',
'Rate (%) [Cumulative Prevalence]', 'Estimated cumulative infections count']]
# Recode 777 and 666 (samples were too small) to nan.
sp_df.replace(to_replace=777,value=np.nan,inplace=True)
sp_df.replace(to_replace=666,value=np.nan,inplace=True)
# Drop NY from the seroprevalence dataframe
sp_df = sp_df[sp_df['Site']!='NY']
sp_df.rename(columns={'Catchment population':'N_catchment',
'n [0-17 Years Prevalence]':'n_sample_0_17',
'n [18-49 Years Prevalence]':'n_sample_18_49',
'n [50-64 Years Prevalence]':'n_sample_50_64',
'n [65+ Years Prevalence]':'n_sample_65_plus',
'Rate (%) [0-17 Years Prevalence]':'pct_sample_0_17',
'Rate (%) [18-49 Years Prevalence]':'pct_sample_18_49',
'Rate (%) [50-64 Years Prevalence]':'pct_sample_50_64',
'Rate (%) [65+ Years Prevalence]':'pct_sample_65_plus',
'Estimated cumulative infections count':'n_est_infections_from_table'
},inplace=True)
# -
print('Percent state/time period insufficient data\n')
for i in age_groups_list:
print(f"Age group '{i}': \t", 100-round(sp_df[f"pct_sample_{i}"].count()/len(sp_df)*100), f'%')
# The seroprevalence DataFrame contains the total population for the state, but it doesn't provide the population by age group for each state. If we know the population by age for each state then we can calculate the total estimated infections by age and state from the infection rate by age that is provided in the seroprevalence DataFrame. We can use these estimates to calculate summary statistics such as the overall mean undercount rate with confidence intervals.
#
# We can obtain population by age and state from Census data and then merge the population data with the seroprevalence DataFrame to create a comprehensive DataFrame. Prior to merging the tables, we need to sum up the populations to create census age groupings that match the seroprevalence age groupings.
#
# Source: https://www.census.gov/data/tables/time-series/demo/popest/2010s-state-detail.html
# ### Loading in state census by age data
# +
state_census = pd.read_csv('../data/sc-est2019-agesex-civ.csv')
state_census = state_census[state_census['SEX']==0][['STATE','NAME','AGE','POPEST2019_CIV']]
pop_grouped = state_census.groupby('STATE').agg(st_fips = ('STATE', 'max'),
st_name = ('NAME', 'max')).reset_index()
# -
# Delete New York from the census count
state_census = state_census[state_census['NAME'] != 'New York']
del pop_grouped['STATE']
pop_grouped['pop_0_17'] = (state_census[state_census['AGE'].isin(range(0,18))].
groupby('STATE').sum()['POPEST2019_CIV'].reset_index())['POPEST2019_CIV']
pop_grouped['pop_18_49'] = (state_census[state_census['AGE'].isin(range(18,50))].
groupby('STATE')['POPEST2019_CIV'].sum().reset_index())['POPEST2019_CIV']
pop_grouped['pop_50_64'] = (state_census[state_census['AGE'].isin(range(50,65))].
groupby('STATE')['POPEST2019_CIV'].sum().reset_index())['POPEST2019_CIV']
pop_grouped['pop_65_plus'] = (state_census[state_census['AGE'].isin(range(65,100))].
groupby('STATE')['POPEST2019_CIV'].sum().reset_index())['POPEST2019_CIV']
# We need to merge (join) the seroprevalence DataFrame with the census table DataFrame ('pop_grouped'), but the state field in the seroprevalence table ('Site') does not match the format of the state fields in the census table ('st+abbr' or 'st_name'). We are going to need to upload a table which contains state values which are common to both, and use that table to join the other two together. One of many tables that will work comes from the COVID Tracking Project. We will use that one here.
NY_df = pd.read_csv("https://api.covidtracking.com/v1/states/daily.csv")[['date', 'state', 'positiveIncrease']]
NY_df = NY_df[NY_df['state']=='NY']
NY_df['date'] = pd.to_datetime(NY_df['date'], format='%Y%m%d')
NY_df['date'].dt.year
# Step 1. Load in the COVID Tracking Project dataset to facilitate merging seroprevalence and census DataFrames.
state_merge_df = pd.read_csv("https://api.covidtracking.com/v1/states/daily.csv")[
['state', 'fips']]
# +
# Delete NY from COVID case counts, but first, capture the percent of cases per month that NY contributes
# because we'll need to those numbers from the age-related COVID counts which aren't organized by state.
state_merge_df
# -
state_merge_df = state_merge_df[state_merge_df['state']!='NY']
# Step 2. Merge COVID Tracking Project DataFrame with census DataFrame by fips code which will populate census DataFrame with state two-letter abbreviation.
pop_grouped = pop_grouped.merge(state_merge_df.groupby('state').max().reset_index(),
left_on = 'st_fips', right_on = 'fips')[
['st_name', 'state', 'fips', 'pop_0_17', 'pop_18_49',
'pop_50_64', 'pop_65_plus']]
# Step 3. Finally, merge census DataFrame from step 2 to populate seroprevalence DataFrame with census data by age and state.
sp_and_census_df = sp_df.merge(pop_grouped, left_on = 'Site', right_on = 'state')
def create_full_month_df(df, start_round, end_round):
'''
Create an abbreviated seroprevalence DataFrame consisting of rounds of the same month
Parameters
----------
df: pandas DataFrame
start_round: number indicating first round to include
end_round: number indicating last round to include
Returns
-------
month_df: Pandas DataFrame
'''
month_df = df[(df['Round'] >= start_round) & (df['Round'] <= end_round)]
month_df = month_df.groupby('Site').agg(
N_catchment = ('N_catchment', 'max'),
n_sample_0_17 = ('n_sample_0_17', 'sum'),
n_sample_18_49 = ('n_sample_18_49', 'sum'),
n_sample_50_64 = ('n_sample_50_64', 'sum'),
n_sample_65_plus = ('n_sample_65_plus', 'sum'),
pct_sample_0_17 = ('pct_sample_0_17', 'mean'),
pct_sample_18_49 = ('pct_sample_18_49', 'mean'),
pct_sample_50_64 = ('pct_sample_50_64', 'mean'),
pct_sample_65_plus = ('pct_sample_65_plus', 'mean'),
n_est_infections_from_table = ('n_est_infections_from_table', 'mean'),
pop_0_17 = ('pop_0_17', 'max'),
pop_18_49 = ('pop_18_49', 'max'),
pop_50_64 = ('pop_50_64', 'max'),
pop_65_plus = ('pop_65_plus', 'max'),
)
return month_df
def point_and_var_calcs(df):
'''
Calculates the estimated number of infections and the std error contribution for each stratum in a seroprevalence DataFrame
Parameters
----------
df: pandas DataFrame
Returns
-------
df: same Pandas DataFrame with new columns added
'''
for ages in age_groups_list:
df[f'n_est_infections_{ages}'] = (df[f'pct_sample_{ages}'] *
df[f'pop_{ages}']) / 100
df[f'stratum_std_err_contribution_{ages}'] = (df[f'pop_{ages}']**2 *
(1- df[f'n_sample_{ages}'] /
df[f'pop_{ages}']) *
(df[f'n_sample_{ages}'] /
(df[f'n_sample_{ages}'] - 1)) *
df[f'pct_sample_{ages}']/100 *
(1 - df[f'pct_sample_{ages}']/100) /
df[f'n_sample_{ages}'])
return df
def missing_data_adj(df):
'''
Adjusts a seroprevalence DataFrame to account for missing data
Parameters
----------
df: pandas DataFrame
Returns
df: pandas DataFrame
'''
# Slight upward adjustment to counts to compensate for missing data
df['pct_age_data_missing'] = (df['n_est_infections_from_table'] -
df['n_est_infections_0_17'] -
df['n_est_infections_18_49'] -
df['n_est_infections_50_64'] -
df['n_est_infections_65_plus'] ) / df['n_est_infections_from_table']
for ages in age_groups_list:
df[f'Est infections (from sp), {ages}'] = (df[f'n_est_infections_{ages}'] /
df[f'pop_{ages}'] / (1-df['pct_age_data_missing']))
return df
def bar_chart_with_yerr(x_pos, means, std_devs, colors, suptitle, sub_title, tick_labels, sources, ylabel, chartname= 'misc'):
fig, ax = plt.subplots(figsize = (10,7))
ax.bar(x_pos,
means,
yerr=[i * critical_value for i in std_devs],
color=colors,
align='center', alpha=0.5, ecolor='black', capsize=10)
ax.set_xticks(x_pos)
ax.set_xticklabels(tick_labels)
ax.set_ylabel(ylabel)
ax.set_title(sub_title)
fig.suptitle(suptitle, size=15, y=0.95)
plt.figtext(0.9, 0, sources, horizontalalignment='right')
ax.yaxis.grid(True)
plt.savefig(f'img/{chartname}.png');
august_df = create_full_month_df(sp_and_census_df, 0, 1)
november_df = create_full_month_df(sp_and_census_df, 7, 8)
august_df = point_and_var_calcs(august_df)
november_df = point_and_var_calcs(november_df)
sp_and_census_df = point_and_var_calcs(sp_and_census_df)
august_df = missing_data_adj(august_df)
november_df = missing_data_adj(november_df)
# +
august_means, august_std = [], []
for ages in age_groups_list:
august_means.append(august_df.sum()[f'n_est_infections_{ages}'] / august_df.sum()[f'pop_{ages}'])
august_std.append(np.sqrt(august_df.sum()[f'stratum_std_err_contribution_{ages}']) / august_df.sum()[f'pop_{ages}'])
sup_title = ' Seroprevalence-Derived Infection Prevalence by Age Group'
sources_ = 'Data sources: CDC Nationwide Commercial Laboratory Seroprevalence Survey, US Census 2019 Population Estimate'
y_label = 'Infection prevalence derived from seroprevalence testing'
bar_chart_with_yerr(x_pos=range(len(age_groups_list)), means=august_means, std_devs=august_std, colors = ['C0', 'C1', 'C2', 'C3'],
suptitle=sup_title, sub_title='Average of August 2020',
tick_labels=age_groups_list, sources=sources_, ylabel=y_label)
# +
november_means, november_std = [], []
for ages in age_groups_list:
november_means.append(november_df.sum()[f'n_est_infections_{ages}'] / november_df.sum()[f'pop_{ages}'])
november_std.append(np.sqrt(november_df.sum()[f'stratum_std_err_contribution_{ages}']) / november_df.sum()[f'pop_{ages}'])
sup_title = ' Seroprevalence-Derived Infection Prevalence by Age Group'
sources_ = 'Data sources: CDC Nationwide Commercial Laboratory Seroprevalence Survey, US Census 2019 Population Estimate'
y_label = 'Infection prevalence derived from seroprevalence testing'
bar_chart_with_yerr(x_pos=range(len(age_groups_list)), means=november_means, std_devs=november_std,
suptitle=sup_title, sub_title='Average of November 2020', colors = ['C0', 'C1', 'C2', 'C3'],
tick_labels=age_groups_list, sources=sources_, ylabel=y_label)
# +
delta_means, delta_std = [], []
delta_means = [november_means[i] - august_means[i] for i in range(len(november_means))]
delta_std = [np.sqrt(november_std[i]**2 + august_std[i]**2) for i in range(len(november_means))]
sup_title = ' Change in Seroprevalence-Derived Infection Prevalence by Age Group'
sources_ = 'Data sources: CDC Nationwide Commercial Laboratory Seroprevalence Survey, US Census 2019 Population Estimate'
y_label = 'Infection prevalence derived from seroprevalence testing'
bar_chart_with_yerr(x_pos=range(len(age_groups_list)), means=delta_means, std_devs=delta_std,
suptitle=sup_title, sub_title='Growth Between November and August 2020',
colors = ['C0', 'C1', 'C2', 'C3'],
tick_labels=age_groups_list, sources=sources_, ylabel=y_label)
# +
aug_nov_mean, aug_nov_std = [], []
aug_nov_mean.append(august_df.sum()[f'n_est_infections_from_table'] / august_df.sum()[f'N_catchment'])
aug_nov_std.append(np.sqrt(august_df.sum()[f'stratum_std_err_contribution_0_17'] +
august_df.sum()[f'stratum_std_err_contribution_18_49'] +
august_df.sum()[f'stratum_std_err_contribution_50_64'] +
august_df.sum()[f'stratum_std_err_contribution_65_plus'])
/ august_df.sum()[f'N_catchment'])
aug_nov_mean.append(november_df.sum()[f'n_est_infections_from_table'] / november_df.sum()[f'N_catchment'] -
august_df.sum()[f'n_est_infections_from_table'] / august_df.sum()[f'N_catchment'])
aug_nov_std.append(np.sqrt((np.sqrt(august_df.sum()[f'stratum_std_err_contribution_0_17'] +
august_df.sum()[f'stratum_std_err_contribution_18_49'] +
august_df.sum()[f'stratum_std_err_contribution_50_64'] +
august_df.sum()[f'stratum_std_err_contribution_65_plus'])
/ august_df.sum()[f'N_catchment'])**2 +
(np.sqrt(november_df.sum()[f'stratum_std_err_contribution_0_17'] +
november_df.sum()[f'stratum_std_err_contribution_18_49'] +
november_df.sum()[f'stratum_std_err_contribution_50_64'] +
november_df.sum()[f'stratum_std_err_contribution_65_plus'])
/ november_df.sum()[f'N_catchment'])**2))
labels = ['August','August to November']
sup_title = ' Seroprevalence-Derived Infection Prevalence'
sources_ = 'Data sources: CDC Nationwide Commercial Laboratory Seroprevalence Survey, US Census 2019 Population Estimate'
y_label = 'Infection prevalence derived from seroprevalence testing'
bar_chart_with_yerr(x_pos=range(len(labels)), means=aug_nov_mean, std_devs=aug_nov_std,
suptitle=sup_title, sub_title='All Age Groups, August and November 2020',
colors = ['C4', 'C8'],
tick_labels=labels, sources=sources_, ylabel=y_label)
# +
delta_aug_nov_mean, delta_aug_nov_std = [], []
delta_aug_nov_mean.append(august_df.sum()[f'n_est_infections_from_table'] / august_df.sum()[f'N_catchment'])
delta_aug_nov_std.append(np.sqrt(august_df.sum()[f'stratum_std_err_contribution_0_17'] +
august_df.sum()[f'stratum_std_err_contribution_18_49'] +
august_df.sum()[f'stratum_std_err_contribution_50_64'] +
august_df.sum()[f'stratum_std_err_contribution_65_plus'])
/ august_df.sum()[f'N_catchment'])
delta_aug_nov_mean.append(november_df.sum()[f'n_est_infections_from_table'] / november_df.sum()[f'N_catchment'])
delta_aug_nov_std.append(np.sqrt(november_df.sum()[f'stratum_std_err_contribution_0_17'] +
november_df.sum()[f'stratum_std_err_contribution_18_49'] +
november_df.sum()[f'stratum_std_err_contribution_50_64'] +
november_df.sum()[f'stratum_std_err_contribution_65_plus'])
/ november_df.sum()[f'N_catchment'])
labels = ['August','November']
sup_title = ' Seroprevalence-Derived Infection Prevalence'
sources_ = 'Data sources: CDC Nationwide Commercial Laboratory Seroprevalence Survey, US Census 2019 Population Estimate'
y_label = 'Infection prevalence derived from seroprevalence testing'
bar_chart_with_yerr(x_pos=range(len(labels)), means=delta_aug_nov_mean, std_devs=delta_aug_nov_std,
suptitle=sup_title, sub_title='All Age Groups, August and November 2020',
colors = ['C4', 'C8'],
tick_labels=labels, sources=sources_, ylabel=y_label)
# -
# ### Creating a usable date range
# We need to convert the date range for each round to a usable format. The specimen collection windows differ slightly by state. Let's find the latest closing date for a given round to use as our marker.
sp_and_census_df['date_range_close'] = pd.to_datetime(sp_and_census_df['Date Range of Specimen Collection'].
str[-12:].str.strip())
round_dates = (sp_and_census_df.groupby('Round').agg(
date_range_close = ('date_range_close', 'max'))
)
round_dates.head(2)
# ### Aggregating by seroprevalence round
# Let's now aggregate the data by seroprevalence rounds (i.e. batches). Once this is done we will adjust the counts upwards to compensate for missing data.
sp_by_round_df = sp_and_census_df.groupby('Round').agg(
pop_0_17 = ('pop_0_17', 'sum'),
pop_18_49 = ('pop_18_49', 'sum'),
pop_50_64 = ('pop_50_64', 'sum'),
pop_65_plus = ('pop_65_plus', 'sum'),
n_est_infections_0_17 = ('n_est_infections_0_17', 'sum'),
n_est_infections_18_49 = ('n_est_infections_18_49', 'sum'),
n_est_infections_50_64 = ('n_est_infections_50_64', 'sum'),
n_est_infections_65_plus = ('n_est_infections_65_plus', 'sum'),
n_est_infections_from_table = ('n_est_infections_from_table', 'sum'),
last_date_of_round = ('date_range_close', 'max')
).reset_index()
# +
# Slight upward adjustment to counts to compensate for missing data
sp_by_round_df['pct_age_data_missing'] = (sp_by_round_df['n_est_infections_from_table'] -
sp_by_round_df['n_est_infections_0_17'] -
sp_by_round_df['n_est_infections_18_49'] -
sp_by_round_df['n_est_infections_50_64'] -
sp_by_round_df['n_est_infections_65_plus'] ) / sp_by_round_df['n_est_infections_from_table']
for ages in age_groups_list:
sp_by_round_df[f'Est infections (from sp), {ages}'] = (sp_by_round_df[f'n_est_infections_{ages}'] /
sp_by_round_df[f'pop_{ages}'] / (1-sp_by_round_df['pct_age_data_missing']))
# +
# Let's see what we have now.
fig, ax = plt.subplots(figsize = (16, 8))
for ages in age_groups_list:
col_name = f'Est infections (from sp), {ages}'
ax.plot(sp_by_round_df['last_date_of_round'],
sp_by_round_df[col_name],
label = col_name,
marker = '.')
ax.set_ylim(0,0.16)
ax.tick_params(axis='x', which='major', labelsize=8)
ax.set_xlabel('End date for specimen collection round')
ax.set_ylabel('Infection rate derived from seroprevalence testing')
ax.set_title('Derived From Antibody Seroprevalence Testing')
ax.legend()
fig.suptitle(f' Seroprevalence-Derived Infection Rate by Age Group', size=15, y=0.95)
plt.figtext(0.9, 0, 'Data sources: CDC Nationwide Commercial Laboratory Seroprevalence Survey, US Census 2019 Population Estimate', horizontalalignment='right');
plt.savefig(f'img/seroprev_by_age_and_period.png')
# -
# ### Loading and preparing case diagnosis data for comparison
#
# Now we need to do the same thing with the case data--calculate the case rate by age group. Let's start by summing up cumulative cases for each age group in the CDC data. We'll loop through the data 8 times--once for each of the specimen collection rounds. For each loop, we'll cut off the count at the date corresponding to that particular specimen collection round's cutoff date. Finally, we will transpose our table and then convert the age groups from the CDC data to match the age groups for the seroprevalence data.
#
# We're also going to need to pull in estimated population from the US Census so we can calculate case diagnosis infection rates.
rows_ = 10000000 #dataset is large. Need to limit rows and/or columns to load.
CDC_case_df1 = pd.read_csv('../data/COVID-19_Case_Surveillance_Public_Use_Data_Feb_2021.csv',
usecols=[0,5,6], encoding='latin-1', nrows=rows_, skiprows=1, header=None)
CDC_case_df2 = pd.read_csv('../data/COVID-19_Case_Surveillance_Public_Use_Data_Feb_2021.csv',
usecols=[0,5,6], encoding='latin-1', nrows=rows_, skiprows=10000001, header=None)
# Source: https://data.cdc.gov/Case-Surveillance/COVID-19-Case-Surveillance-Public-Use-Data/vbim-akqf/data
CDC_case_df = pd.concat([CDC_case_df1, CDC_case_df2], ignore_index=True)
CDC_case_df.columns = ['cdc_case_earliest_dt ', 'sex', 'age_group']
CDC_case_df = CDC_case_df[CDC_case_df['age_group']!='Missing'] # less than 1% unknowns, so drop them.
# +
# Create a new DataFrame with each cases by age_groups in rows and by round in columns. Transpose to orient properly.
age_group_df = pd.DataFrame(CDC_case_df['age_group'].unique(), columns=['age_group'])
round_ = 1
for date_ in sp_by_round_df['last_date_of_round']:
curr_case_df = (CDC_case_df[pd.to_datetime(CDC_case_df['cdc_case_earliest_dt ']) < date_].
groupby('age_group').count()
)
curr_case_df = curr_case_df.rename(columns={'sex': round_})[round_].reset_index()
round_ +=1
age_group_df = age_group_df.merge(curr_case_df, on='age_group')
age_group_df = age_group_df.T
age_group_df.columns = age_group_df.iloc[0]
age_group_df = age_group_df[1:]
age_group_df = age_group_df.reset_index().rename(columns={'index':'Round'})
# -
# Aligning case count age groups with seroprevalence age groups.
age_group_df['cases_0_17'] = (age_group_df['0 - 9 Years'] +
4/5 * age_group_df['10 - 19 Years'])
age_group_df['cases_18_49'] = (1/5 * age_group_df['10 - 19 Years'] +
age_group_df['20 - 29 Years'] +
age_group_df['30 - 39 Years'] +
age_group_df['40 - 49 Years'])
age_group_df['cases_50_64'] = (age_group_df['50 - 59 Years'] +
1/2 * age_group_df['60 - 69 Years'])
age_group_df['cases_65_plus'] = (1/2* age_group_df['60 - 69 Years'] +
age_group_df['70 - 79 Years'] +
age_group_df['80+ Years'])
# ### Comparing antibody seroprevalence counts to antigen diagnostic case counts
# Merge the two distinct DataFrames
sp_and_case_df = sp_by_round_df.merge(age_group_df, on='Round')
sp_and_case_df
# Calculating the case undercount rates
for ages in age_groups_list:
sp_and_case_df[f'Est infections (from cases), {ages}'] = (sp_and_case_df[f'cases_{ages}'] / sp_and_case_df[f'pop_{ages}'] /
(1-pct_unknowns))
sp_and_case_df[f'Undercount rate, {ages}'] = (sp_and_case_df[f'n_est_infections_{ages}'] /
sp_and_case_df[f'cases_{ages}'])
# +
# Chart case-derived estimate of infection rate by age group
fig, ax = plt.subplots(figsize = (16, 8))
for ages in age_groups_list:
col_name = f'Est infections (from cases), {ages}'
ax.plot(sp_and_case_df['last_date_of_round'],
sp_and_case_df[col_name],
label = col_name,
marker = '.')
ax.tick_params(axis='x', which='major', labelsize=8)
ax.set_ylim(0,0.16)
ax.set_xlabel('Specimen collection round number')
ax.set_ylabel('Infection rate derived from cases')
ax.set_title('Derived From Antigen Case Diagnostic Testing')
ax.legend()
fig.suptitle(f' Case-Derived Infection Rate by Age Group', size=15, y=0.95)
plt.figtext(0.9, 0, 'Data sources: CDC COVID-19 Case Surveillance Public Data', horizontalalignment='right')
plt.savefig(f'img/cases_by_age_and_period.png');
# -
# ## Dropping New York and '0-17' age group
# ### Requires reconstructing dataframes
# +
fig, ax = plt.subplots(figsize = (16, 8))
for ages in short_age_groups_list:
col_name = f'Undercount rate, {ages}'
ax.plot(sp_and_case_df['last_date_of_round'],
sp_and_case_df[col_name],
label = col_name,
marker = '.')
ax.tick_params(axis='x', which='major', labelsize=8)
ax.set_ylim(0,5)
ax.set_xlabel('Specimen collection round number')
ax.set_ylabel('Undercount Rate')
ax.set_title('Seroprevalence Estimate Divided by Cumulative Cases')
ax.legend()
fig.suptitle(f'Case Undercount Rate by Age Group', size=15, y=0.95)
plt.figtext(0.9, 0, 'Data sources: CDC COVID-19 Case Surveillance Public Data, CDC Nationwide Commercial Laboratory Seroprevalence Survey, US Census 2019 Population Estimate', horizontalalignment='right')
plt.savefig(f'img/undercount_by_age_and_period.png');
# -
# Add in confidence intervals, statistical tests to describe how likely it is that the observed trends reflect actual trends.
#
# Try to come up with an estimate of how undercount rates by age have changed over time. Will need to estimate the undercount before August?
sp_and_case_df.columns
# +
cols =['n_est_infections_0_17', 'n_est_infections_18_49', 'n_est_infections_50_64',
'n_est_infections_65_plus', 'n_est_infections_from_table', 'cases_0_17', 'cases_18_49', 'cases_50_64',
'cases_65_plus']
for ages in age_groups_list:
col = f'n_est_infections_{ages}'
baseline_new_name = col + '_baseline'
sp_and_case_df[baseline_new_name] = (sp_and_case_df[col][0] + sp_and_case_df[col][1]) / 2
change_new_name = col + '_change_from_base'
sp_and_case_df[change_new_name] = sp_and_case_df[col] - sp_and_case_df[baseline_new_name]
for ages in age_groups_list:
col = f'cases_{ages}'
baseline_new_name = col + '_baseline'
sp_and_case_df[baseline_new_name] = (sp_and_case_df[col][0] + sp_and_case_df[col][1]) / 2
change_new_name = col + '_change_from_base'
sp_and_case_df[change_new_name] = sp_and_case_df[col] - sp_and_case_df[baseline_new_name]
sp_and_case_df['Undercount rate (before August), 0_17'] = (sp_and_case_df['n_est_infections_0_17_baseline'] /
sp_and_case_df['cases_0_17_baseline'])
sp_and_case_df['Undercount rate (before August), 18_49'] = (sp_and_case_df['n_est_infections_18_49_baseline'] /
sp_and_case_df['cases_18_49_baseline'])
sp_and_case_df['Undercount rate (before August), 50_64'] = (sp_and_case_df['n_est_infections_50_64_baseline'] /
sp_and_case_df['cases_50_64_baseline'])
sp_and_case_df['Undercount rate (before August), 65_plus'] = (sp_and_case_df['n_est_infections_65_plus_baseline'] /
sp_and_case_df['cases_65_plus_baseline'])
sp_and_case_df['Undercount rate (from August), 0_17'] = (sp_and_case_df['n_est_infections_0_17_change_from_base'] /
sp_and_case_df['cases_0_17_change_from_base'])
sp_and_case_df['Undercount rate (from August), 18_49'] = (sp_and_case_df['n_est_infections_18_49_change_from_base'] /
sp_and_case_df['cases_18_49_change_from_base'])
sp_and_case_df['Undercount rate (from August), 50_64'] = (sp_and_case_df['n_est_infections_50_64_change_from_base'] /
sp_and_case_df['cases_50_64_change_from_base'])
sp_and_case_df['Undercount rate (from August), 65_plus'] = (sp_and_case_df['n_est_infections_65_plus_change_from_base'] /
sp_and_case_df['cases_65_plus_change_from_base'])
# -
sp_and_case_df['cases_0_17_change_from_base']
# +
fig, ax = plt.subplots(figsize = (12, 8))
for ages in age_groups_list:
col_name = f'Undercount rate (from August), {ages}'
ax.plot(sp_and_case_df['last_date_of_round'],
sp_and_case_df[col_name],
label = col_name,
marker = '.')
ax.set_xlabel('Specimen collection round number')
ax.set_ylabel('Undercount Rate')
ax.set_title('Cumulative Cases Compared to Seroprevalence Estimate')
ax.set_xlim(left=sp_and_case_df['last_date_of_round'][4])
ax.set_ylim(0,12)
ax.legend()
fig.suptitle(f'Case Undercount Rate From August 2020 by Age Group', size=15, y=0.95)
plt.figtext(0.9, 0, 'Data sources: CDC COVID-19 Case Surveillance Public Data, CDC Nationwide Commercial Laboratory Seroprevalence Survey, US Census 2019 Population Estimate', horizontalalignment='right');
# +
summary_df = sp_and_case_df[sp_and_case_df['last_date_of_round']>'2020-10-01']
for col_name in ['Undercount rate (before August), 0_17', 'Undercount rate (before August), 18_49',
'Undercount rate (before August), 50_64', 'Undercount rate (before August), 65_plus',
'Undercount rate (from August), 0_17', 'Undercount rate (from August), 18_49',
'Undercount rate (from August), 50_64', 'Undercount rate (from August), 65_plus']:
print(col_name[-7:], ' ', summary_df[col_name].mean())
# -
# ## New section comparing August to difference between November and August
aug_case_rate_by_age, nov_minus_aug_case_rate_by_age = [], []
aug_ttl, nov_minus_aug_ttl, pop_ttl = 0, 0, 0
for ages in age_groups_list:
aug_ttl += sp_and_case_df[f'cases_{ages}'].iloc[0]
nov_minus_aug_ttl += sp_and_case_df[f'cases_{ages}'].iloc[6] - sp_and_case_df[f'cases_{ages}'].iloc[0]
pop_ttl += sp_and_case_df[f'pop_{ages}'][0]
aug_rate = sp_and_case_df[f'Est infections (from cases), {ages}'].iloc[:1].mean()
nov_rate = sp_and_case_df[f'Est infections (from cases), {ages}'].iloc[6:7].mean()
aug_case_rate_by_age.append(aug_rate)
nov_minus_aug_case_rate_by_age.append(nov_rate - aug_rate)
case_rate_ttls = [aug_ttl / pop_ttl, nov_minus_aug_ttl / pop_ttl]
# +
august_undercount_by_age = [august_means[i] / aug_case_rate_by_age[i] for i in range(len(august_means))]
august_undercount_by_age_std = [august_std[i] / august_means[i] * august_undercount_by_age[i]
for i in range(len(august_std))]
nov_minus_aug_undercount_by_age = [delta_means[i] / nov_minus_aug_case_rate_by_age[i]
for i in range(len(delta_means))]
nov_minus_aug_undercount_by_age_std = [delta_std[i] / delta_means[i] * nov_minus_aug_undercount_by_age[i]
for i in range(len(delta_means))]
august_delta_nov_undercount_ttl = [aug_nov_mean[i] / case_rate_ttls[i] for i in range(len(aug_nov_mean))]
august_delta_nov_undercount_std = [aug_nov_std[i] / aug_nov_mean[i] * august_delta_nov_undercount_ttl[i]
for i in range(len(aug_nov_mean))]
# +
labels = ['Up to August 2020','September to November 2020']
sup_title = ' Diagnostic Case Undercount Rate'
sources_ = 'Data sources: CDC COVID-19 Case Surveillance Public Data, CDC Nationwide Commercial Laboratory Seroprevalence Survey, US Census 2019 Population Estimate'
y_label = 'Antigen Seroprevalence Tests Divided By Cumulative Antibody Case Diagnoses'
bar_chart_with_yerr(x_pos=range(len(labels)), means=august_delta_nov_undercount_ttl, std_devs=august_delta_nov_undercount_std,
suptitle=sup_title, sub_title='All Age Groups', colors = ['C4', 'C8'],
tick_labels=labels, sources=sources_, ylabel=y_label, chartname='all_august_before_after')
# -
labels = age_groups_list
sup_title = ' Diagnostic Case Undercount Rate'
sources_ = 'Data sources: CDC COVID-19 Case Surveillance Public Data, CDC Nationwide Commercial Laboratory Seroprevalence Survey, US Census 2019 Population Estimate'
y_label = 'Antigen Seroprevalence Tests Divided By Cumulative Antibody Case Diagnoses'
bar_chart_with_yerr(x_pos=range(len(labels)), means=august_undercount_by_age, std_devs=august_undercount_by_age_std,
suptitle=sup_title, sub_title='Up to August 2020', colors = ['C0', 'C1', 'C2', 'C3'],
tick_labels=labels, sources=sources_, ylabel=y_label)
labels = age_groups_list
sup_title = ' Diagnostic Case Undercount Rate'
sources_ = 'Data sources: CDC COVID-19 Case Surveillance Public Data, CDC Nationwide Commercial Laboratory Seroprevalence Survey, US Census 2019 Population Estimate'
y_label = 'Antigen Seroprevalence Tests Divided By Cumulative Antibody Case Diagnoses'
bar_chart_with_yerr(x_pos=range(len(labels)), means=nov_minus_aug_undercount_by_age, std_devs=nov_minus_aug_undercount_by_age_std,
suptitle=sup_title, sub_title='September to November 2020', colors = ['C0', 'C1', 'C2', 'C3'],
tick_labels=labels, sources=sources_, ylabel=y_label)
| No_NY_covid-age-case-vs-seroprev-Copy1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import re
from tf_rnn_classifier import TfRNNClassifier
import tweepy
from collections import Counter
from sklearn.feature_extraction import DictVectorizer
from sklearn.metrics import classification_report, accuracy_score, f1_score
from sklearn.model_selection import train_test_split, GridSearchCV
from sklearn.naive_bayes import MultinomialNB
from sklearn.svm import LinearSVC
from textblob import TextBlob
auth = tweepy.AppAuthHandler('aiA4EZjqeZ13it3Qvsgy9nHcv', '<KEY>')
# auth.set_access_token('<KEY>', '<KEY>')
api = tweepy.API(auth, wait_on_rate_limit=True,wait_on_rate_limit_notify=True)
# -
def create_training_data():
num_tweets = 1000
files = ['data/positive_tweets.txt', 'data/negative_tweets.txt']
for file in files:
query = ''
if file == 'data/positive_tweets.txt':
query = ':) OR (: OR :-) OR <3 OR :D -:('
else:
query = ':( OR ): OR >:( -:)'
query +=' -filter:retweets' # ignore retweets
with open(file,'w') as f:
for tweet in tweepy.Cursor(api.search,q=query,tweet_mode="extended").items(num_tweets):
if tweet.lang == 'en':
txt = tweet.full_text.replace('\n', '')
f.write(txt)
f.write('\n')
create_training_data()
# +
class NBAData(object):
def __init__(self):
self.path = 'data/'
# Call GATE POS tagger to label words accordingly
def tag_tweets(self, file):
# files in the tagger expect to be referenced from within tagger folder
curr_path = os.getcwd()
path = '{}{}'.format(curr_path, '/twitie-tagger')
os.chdir(path)
# change tweets_pos/new_rigs
c = 'java -jar ./twitie_tag.jar ./models/gate-EN-twitter.model ../data/{} > ../data/tagged_{}'.format(file, file)
os.system(c)
os.chdir(curr_path)
def bigrams_unigrams_phi(self, text):
words = ['<S>'] + text.split() + ['</S>']
bigrams = []
unigrams = []
for i in range(len(words) - 1):
if i != 0:
unigrams.append((words[i],))
bigrams.append((words[i], words[i+1]))
return Counter(unigrams + bigrams)
# Featurizes data into POS bigrams after data has been cleaned, stripped, etc
def transform_data(self, data, vectorizer=None):
dicts = {}
feat_matrix = None
feat_dicts = []
for i in range(len(data)):
feat_dicts.append(self.bigrams_unigrams_phi(data[i]))
vectorizer = DictVectorizer()
feat_matrix = vectorizer.fit_transform(feat_dicts)
return {'X': feat_matrix, 'vectorizer': vectorizer}
# remove links, regularize capitalization, etc on all files passed in
def preprocess_data(self, file):
pos_emoji_pattern = r'(:-?(?:\)+|D+))|((?:\(+)-?:<?)|(<3+)'
neg_emoji_pattern = r'(>?:-?(?:\(+))|((?:D+|\)+)-?:<?)'
lines = None
with open('{}{}'.format(self.path, file), 'r') as f:
lines = f.readlines() # old contents with all the extra shit
with open('{}{}'.format(self.path, file), 'w') as f:
for line in lines:
new_line = re.sub(r'https?:\/\/.*[\r\n]*', '', line, flags=re.MULTILINE).lower()
new_line = re.sub(pos_emoji_pattern, '', new_line)
new_line = re.sub(neg_emoji_pattern, '', new_line)
f.write(new_line)
# def get_labelled_data(self, file):
# data = []
# with open('{}{}'.format(self.path, dataset), 'r') as f:
# for line in f.readlines():
# line = line.strip()
# idx = line.rfind(',')
# data.append((line[1:idx-1], int(line[idx+1:])))
# return data
def get_processed_data(self, file):
data = []
with open('{}{}'.format(self.path, file), 'r') as f:
for line in f.readlines():
line = line.strip()
data.append(line)
return data
def label_data(self, process_data):
pos_emoji_pattern = r'(:-?(?:\)+|D+))|((?:\(+)-?:<?)|(<3+)'
neg_emoji_pattern = r'(>?:-?(?:\(+))|((?:D+|\)+)-?:<?)'
data = []
for entry in process_data:
entry = re.sub(r'https?:\/\/.*[\r\n]*', '', entry, flags=re.MULTILINE)
sentiment = 0
if re.findall(pos_emoji_pattern, entry):
sentiment += 1
entry = re.sub(pos_emoji_pattern, '', entry)
if re.findall(neg_emoji_pattern, entry):
sentiment -= 1
entry = re.sub(neg_emoji_pattern, '', entry)
data.append((entry, sentiment))
return data
def predict_baseline(self, data, labels):
correct = 0
attempts = 0
incorrect = [('tweet', 'exp_sentiment', 'sentiment')]
for i in range(len(data)):
tweet = data[i]
sentiment = labels[i]
exp_sentiment = self.get_tweet_sentiment_baseline(tweet)
if exp_sentiment == sentiment:
correct += 1
else:
incorrect.append((tweet, exp_sentiment, sentiment))
attempts += 1
# print ('{}% successful: {} correct, {} attempts'.format(correct*100//attempts, correct, attempts))
print(float(correct)/attempts)
def get_tweet_sentiment_baseline(self, tweet):
analysis = TextBlob(self.clean_tweet(tweet))
# set sentiment
if analysis.sentiment.polarity > 0:
return 1
elif analysis.sentiment.polarity <= 0:
return -1
def predict(self, X, y, clf):
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=42)
clf.fit(X_train, y_train)
predictions = clf.predict(X_test)
print('Accuracy: %0.03f' % accuracy_score(y_test, predictions))
print(classification_report(y_test, predictions, digits=3))
misclassified_samples = X_test[y_test != predictions]
print(misclassified_samples)
return clf
def clean_tweet(self, tweet):
return ' '.join(re.sub("(@[A-Za-z0-9]+)|([^0-9A-Za-z \t])|(\w+:\/\/\S+)", " ", tweet).split())
def get_vocab(self, data):
wc = Counter([w for d in data for w in d.split()])
wc = wc.items()
vocab = {w for w, c in wc}
vocab.add("$UNK")
return sorted(vocab)
# -
model = NBAData()
files = ['positive_tweets.txt', 'negative_tweets.txt']
for file in files: # probably want to go ahead and do this for all tweets about NBA players too...
model.preprocess_data(file)
model.tag_tweets(file) # new files w/ GATE POS labels for all relevant tweets
# +
# textblob shitty ass fuckin baseline you piece of shit rig
model = NBAData()
pos_data = model.get_processed_data('positive_tweets.txt') # list of processed tweets
neg_data = model.get_processed_data('negative_tweets.txt')
pos_labels = [1] * len(pos_data)
neg_labels = [-1] * len(neg_data)
model.predict_baseline(pos_data + neg_data, pos_labels + neg_labels)
# -
model = NBAData()
# +
import numpy as np
tagged_files = ['tagged_positive_tweets.txt', 'tagged_negative_tweets.txt']
pos_data = model.get_processed_data('tagged_positive_tweets.txt') # list of processed tweets
neg_data = model.get_processed_data('tagged_negative_tweets.txt')
res = model.transform_data(pos_data + neg_data)
X = res['X']
pos_labels = [1] * len(pos_data)
neg_labels = [-1] * len(neg_data)
y = pos_labels + neg_labels
print("Number of training samples: {}".format(len(y)))
nb_clf = model.predict(X, y, MultinomialNB(alpha=0.5))
svc_clf = model.predict(X, y, LinearSVC())
# tf_rnn = TfRNNClassifier(
# model.get_vocab(pos_data + neg_data),
# embed_dim=50,
# hidden_dim=50,
# max_length=200, # idk how long a tweet is
# hidden_activation=tf.nn.tanh,
# cell_class=tf.nn.rnn_cell.LSTMCell,
# train_embedding=True,
# max_iter=100,
# eta=0.05)
# rnn_X = [[w for w in d.split()] for d in pos_data + neg_data]
# X_train, X_test, y_train, y_test = train_test_split(rnn_X, y, test_size=0.2, random_state=42)
# tf_rnn.fit(X_train, y_train)
# tf_rnn_dev_predictions = tf_rnn.predict(X_test)
# print(classification_report(y_test, tf_rnn_dev_predictions))
# print(accuracy_score(y_test, tf_rnn_dev_predictions))
# -
| MVPy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Working with JAX numpy and calculating perplexity: Ungraded Lecture Notebook
# Normally you would import `numpy` and rename it as `np`.
#
# However in this week's assignment you will notice that this convention has been changed.
#
# Now standard `numpy` is not renamed and `trax.fastmath.numpy` is renamed as `np`.
#
# The rationale behind this change is that you will be using Trax's numpy (which is compatible with JAX) far more often. Trax's numpy supports most of the same functions as the regular numpy so the change won't be noticeable in most cases.
#
# +
import numpy
import trax
import trax.fastmath.numpy as np
# Setting random seeds
trax.supervised.trainer_lib.init_random_number_generators(32)
numpy.random.seed(32)
# -
# One important change to take into consideration is that the types of the resulting objects will be different depending on the version of numpy. With regular numpy you get `numpy.ndarray` but with Trax's numpy you will get `jax.interpreters.xla.DeviceArray`. These two types map to each other. So if you find some error logs mentioning DeviceArray type, don't worry about it, treat it like you would treat an ndarray and march ahead.
#
# You can get a randomized numpy array by using the `numpy.random.random()` function.
#
# This is one of the functionalities that Trax's numpy does not currently support in the same way as the regular numpy.
numpy_array = numpy.random.random((5,10))
print(f"The regular numpy array looks like this:\n\n {numpy_array}\n")
print(f"It is of type: {type(numpy_array)}")
# You can easily cast regular numpy arrays or lists into trax numpy arrays using the `trax.fastmath.numpy.array()` function:
# +
trax_numpy_array = np.array(numpy_array)
print(f"The trax numpy array looks like this:\n\n {trax_numpy_array}\n")
print(f"It is of type: {type(trax_numpy_array)}")
# -
# Hope you now understand the differences (and similarities) between these two versions and numpy. **Great!**
#
# The previous section was a quick look at Trax's numpy. However this notebook also aims to teach you how you can calculate the perplexity of a trained model.
#
# ## Calculating Perplexity
# The perplexity is a metric that measures how well a probability model predicts a sample and it is commonly used to evaluate language models. It is defined as:
#
# $$P(W) = \sqrt[N]{\prod_{i=1}^{N} \frac{1}{P(w_i| w_1,...,w_{n-1})}}$$
#
# As an implementation hack, you would usually take the log of that formula (to enable us to use the log probabilities we get as output of our `RNN`, convert exponents to products, and products into sums which makes computations less complicated and computationally more efficient). You should also take care of the padding, since you do not want to include the padding when calculating the perplexity (because we do not want to have a perplexity measure artificially good). The algebra behind this process is explained next:
#
#
# $$log P(W) = {log\big(\sqrt[N]{\prod_{i=1}^{N} \frac{1}{P(w_i| w_1,...,w_{n-1})}}\big)}$$
#
# $$ = {log\big({\prod_{i=1}^{N} \frac{1}{P(w_i| w_1,...,w_{n-1})}}\big)^{\frac{1}{N}}}$$
#
# $$ = {log\big({\prod_{i=1}^{N}{P(w_i| w_1,...,w_{n-1})}}\big)^{-\frac{1}{N}}} $$
# $$ = -\frac{1}{N}{log\big({\prod_{i=1}^{N}{P(w_i| w_1,...,w_{n-1})}}\big)} $$
# $$ = -\frac{1}{N}{\big({\sum_{i=1}^{N}{logP(w_i| w_1,...,w_{n-1})}}\big)} $$
# You will be working with a real example from this week's assignment. The example is made up of:
# - `predictions` : batch of tensors corresponding to lines of text predicted by the model.
# - `targets` : batch of actual tensors corresponding to lines of text.
# +
from trax import layers as tl
# Load from .npy files
predictions = numpy.load('predictions.npy')
targets = numpy.load('targets.npy')
# Cast to jax.interpreters.xla.DeviceArray
predictions = np.array(predictions)
targets = np.array(targets)
# Print shapes
print(f'predictions has shape: {predictions.shape}')
print(f'targets has shape: {targets.shape}')
# -
# Notice that the predictions have an extra dimension with the same length as the size of the vocabulary used.
#
# Because of this you will need a way of reshaping `targets` to match this shape. For this you can use `trax.layers.one_hot()`.
#
# Notice that `predictions.shape[-1]` will return the size of the last dimension of `predictions`.
reshaped_targets = tl.one_hot(targets, predictions.shape[-1]) #trax's one_hot function takes the input as one_hot(x, n_categories, dtype=optional)
print(f'reshaped_targets has shape: {reshaped_targets.shape}')
# By calculating the product of the predictions and the reshaped targets and summing across the last dimension, the total log perplexity can be computed:
total_log_ppx = np.sum(predictions * reshaped_targets, axis= -1)
# Now you will need to account for the padding so this metric is not artificially deflated (since a lower perplexity means a better model). For identifying which elements are padding and which are not, you can use `np.equal()` and get a tensor with `1s` in the positions of actual values and `0s` where there are paddings.
non_pad = 1.0 - np.equal(targets, 0)
print(f'non_pad has shape: {non_pad.shape}\n')
print(f'non_pad looks like this: \n\n {non_pad}')
# By computing the product of the total log perplexity and the non_pad tensor we remove the effect of padding on the metric:
real_log_ppx = total_log_ppx * non_pad
print(f'real perplexity still has shape: {real_log_ppx.shape}')
# You can check the effect of filtering out the padding by looking at the two log perplexity tensors:
print(f'log perplexity tensor before filtering padding: \n\n {total_log_ppx}\n')
print(f'log perplexity tensor after filtering padding: \n\n {real_log_ppx}')
# To get a single average log perplexity across all the elements in the batch you can sum across both dimensions and divide by the number of elements. Notice that the result will be the negative of the real log perplexity of the model:
log_ppx = np.sum(real_log_ppx) / np.sum(non_pad)
log_ppx = -log_ppx
print(f'The log perplexity and perplexity of the model are respectively: {log_ppx} and {np.exp(log_ppx)}')
# **Congratulations on finishing this lecture notebook!** Now you should have a clear understanding of how to work with Trax's numpy and how to compute the perplexity to evaluate your language models. **Keep it up!**
| Natural Language Processing Specialization/n-gram_vs_sequence_models/C3_W2_lecture_notebook_perplexity.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="YeI6DQjrAxV6"
# Extract Survival curves
# + id="wdfbeCUZBOCT"
# Use DeepSurv from the repo
import sys
sys.path.append('../deepsurv')
from sklearn.preprocessing import StandardScaler
from sklearn_pandas import DataFrameMapper
import torch
import torchtuples as tt
from pycox.datasets import metabric
from pycox.models import CoxPH
from pycox.evaluation import EvalSurv
from sklearn_pandas import DataFrameMapper
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from pysurvival.models.semi_parametric import NonLinearCoxPHModel
from pysurvival.utils.metrics import concordance_index
from pysurvival.utils.display import integrated_brier_score
from sklearn.model_selection import KFold
# + id="Qqpo2oFtydn0"
# #!pip install pysurvival
# + colab={"base_uri": "https://localhost:8080/"} id="33lUJI7Es79p" outputId="7700b2bc-b97f-4fff-d5e5-fc2490ca083e"
# %cd /content/drive/MyDrive/Colab Notebooks/research_2021/
# + colab={"base_uri": "https://localhost:8080/"} id="TSp9YmUtgbLL" outputId="369f0504-9be3-4f51-d70f-1ad7abbbec0e"
from google.colab import drive
drive.mount('/content/drive')
# + [markdown] id="5CDnGHZqBdbA"
# Load datasets
# + id="YNIjI954BU8e"
# Loading and preparing the Datasets
train_dataset_fp1 = 'New1996Chad.csv'
train_dataset_fp2 = 'New2004Chad.csv'
train_dataset_fp3 = 'New2014Chad.csv'
train_df1 = pd.read_csv(train_dataset_fp1)
train_df2 = pd.read_csv(train_dataset_fp2)
train_df3 = pd.read_csv(train_dataset_fp3)
#train_df4 = pd.read_csv(train_dataset_fp4)
# + [markdown] id="XSIdw4nRBtEv"
#
#
# Split the data into train, and test dataset
#
# + id="7SqonIkDF0Oh"
data_ds = train_df3.copy()
df_train = data_ds.copy()
df_test = df_train.sample(frac=0.2)
df_train = df_train.drop(df_test.index)
cols_leave = ['Data.Time', 'Data.status','Data.B4', 'Data.V025', 'Data.V149', 'Data.V190']
#cols_leave = ['Data.Time', 'Data.status','Data.b4', 'Data.v025', 'Data.v149']
leave = [(col, None) for col in cols_leave]
x_mapper = DataFrameMapper(leave)
x_train = x_mapper.fit_transform(df_train).astype('float32')
x_test = x_mapper.transform(df_test).astype('float32')
get_target = lambda df: (df['Data.Time'].values, df['Data.status'].values)
y_train = get_target(df_train)
durations_test, events_test = get_target(df_test)
# + [markdown] id="ZvmqOu79F9KO"
# Run the deep surv model
# + id="OzlZlAZ7F8ff" colab={"base_uri": "https://localhost:8080/"} outputId="6281d130-bdee-4f51-b2c1-00a08d6f9ac8"
## Deepsurv Input
dropout = 0.2
epochs = 1000
structure = [ {'activation': 'ReLU', 'num_units': 128}, {'activation': 'ReLU', 'num_units': 128},{'activation': 'ReLU', 'num_units': 128},{'activation': 'ReLU', 'num_units': 128}]
model_ =NonLinearCoxPHModel(structure=structure)
model_.fit(x_train, y_train[0],y_train[1], lr=1e-8,num_epochs = epochs, dropout = dropout,init_method='xav_uniform',l2_reg=1e-4, batch_normalization=True)
# + [markdown] id="IhTuH6pSGN92"
# Predicted survival probabilities
# + id="jGAz5hgXGPzs"
# Computing the Survival function for all times t on the test dataset
predicted = model_.predict_survival(x_test)
surv_pred= predicted.mean(axis=0)
t = model_.times
pairs = {'duration':t, '0': surv_pred}
df = pd.DataFrame.from_dict(pairs)
df.to_csv('Deep_surv_survival_Chad2014New.csv')
# + [markdown] id="EJhc5nq4Ggq-"
# Plot the survival curves
#
# + id="03KwHRKJGe51" colab={"base_uri": "https://localhost:8080/", "height": 748} outputId="8b74abe0-4f1f-4a90-de34-3fea19be1d24"
# One country from each region
from numpy import *
import math
import matplotlib.pyplot as plt
# plot all the curves together
fig = plt.figure()
fig, axs = plt.subplots(2, 2, sharex=True, sharey=True)
fig.suptitle('Survival curves extracted from the deepsurv model')
# Zimbabwe
Zim_2006 = pd.read_csv('Deep_surv_survival_zim2006New.csv')
Zim_2011 = pd.read_csv('Deep_surv_survival_zim2011New.csv')
Zim_2015 = pd.read_csv('Deep_surv_survival_zim2015New.csv')
t1z = Zim_2006["duration"]
t2z = Zim_2011["duration"]
t3z = Zim_2015["duration"]
axs[0,0].plot(t1z, Zim_2006["0"], t2z, Zim_2011["0"], t3z,Zim_2015["0"], linewidth=3.0)
axs[0,0].legend(["2006", "2011", "2015"], title="Zimbabwe")
axs[0,0].axhline(y=0.5, color='black', linestyle='-')
# Uganda
Uganda_2006 = pd.read_csv('Deep_surv_survival_Uganda2006New.csv')
Uganda_2011 = pd.read_csv('Deep_surv_survival_Uganda2011New.csv')
Uganda_2016 = pd.read_csv('Deep_surv_survival_Uganda2016New.csv')
t1 = Uganda_2006["duration"]
t2 = Uganda_2011["duration"]
t3 = Uganda_2016["duration"]
axs[0,1].plot(t1, Uganda_2006["0"], t2, Uganda_2011["0"], t3,Uganda_2016["0"], linewidth=3.0)
axs[0,1].legend([ "2006", "2011", "2016"], title="Uganda")
axs[0,1].axhline(y=0.5, color='black', linestyle='-')
# Ghana
Ghana_2003 = pd.read_csv('Deep_surv_survival_Ghana2003New.csv')
Ghana_2008 = pd.read_csv('Deep_surv_survival_Ghana2008New.csv')
Ghana_2014 = pd.read_csv('Deep_surv_survival_Ghana2014New.csv')
t1G = Ghana_2003["duration"]
t2G = Ghana_2008["duration"]
t3G = Ghana_2014["duration"]
axs[1,0].plot( t1G, Ghana_2003["0"],t2G, Ghana_2008["0"], t3G,Ghana_2014["0"], linewidth=3.0)
axs[1,0].legend(["2003", "2008", "2014"], title="Ghana")
axs[1,0].axhline(y=0.5, color='black', linestyle='-')
# Chad
Chad_2004 = pd.read_csv('Deep_surv_survival_Chad2004New.csv')
Chad_2014 = pd.read_csv('Deep_surv_survival_Chad2014New.csv')
t1C = Chad_2004["duration"]
t2C = Chad_2014["duration"]
axs[1,1].plot( t1C, Chad_2004["0"], t2C, Chad_2014["0"], linewidth=3.0)
axs[1,1].legend(["2004", "2014"], title="Chad")
axs[1,1].set_ylim(ymin=0)
axs[1,1].axhline(y=0.5, color='black', linestyle='-')
fig.text(0.5, 0.04, 'Survival time in months', ha='center', va='center')
fig.text(0.02, 0.5, 'Survival probabilities', ha='center', va='center', rotation='vertical')
# + [markdown] id="_RBLqd-qGyRY"
# 10-fold cross validation
# + [markdown] id="ipNzVnxGLS8Y"
# Load datasets
#
# + id="KZcgRFTkLfOS"
train_dataset_fp1 = 'New1996Chad.csv'
train_dataset_fp2 = 'New2004Chad.csv'
train_dataset_fp3 = 'New2014Chad.csv'
#train_dataset_fp4 = 'New2014Ghana.csv'
train_df1 = pd.read_csv(train_dataset_fp1)
train_df2 = pd.read_csv(train_dataset_fp2)
train_df3 = pd.read_csv(train_dataset_fp3)
#train_df4 = pd.read_csv(train_dataset_fp4)
# + [markdown] id="oKjDEemjSaMb"
# Data splitting
# + id="CgsSgYl2NWfj" colab={"base_uri": "https://localhost:8080/"} outputId="38f16539-b017-49fb-8c15-bb45e7358f53"
cv =KFold(n_splits=10,random_state=42, shuffle=False)
train_index1= list()
train_index2= list()
train_index3= list()
train_index4= list()
test_index1= list()
test_index2= list()
test_index3= list()
test_index4= list()
for i in cv.split(train_df1):
train_index1.append(i[0])
test_index1.append(i[1])
for j in cv.split(train_df2):
train_index2.append(j[0])
test_index2.append(j[1])
for k in cv.split(train_df3):
train_index3.append(k[0])
test_index3.append(k[1])
#for b in cv.split(train_df4):
# train_index4.append(b[0])
# test_index4.append(b[1])
# + [markdown] id="Ew96q2rjPKKs"
# Run the model
# + id="P0VnCxEzPIDo" colab={"base_uri": "https://localhost:8080/"} outputId="28d7b17f-3ec0-4dd0-de59-0345e10128ff"
scores1 = []
scores2 = []
scores3 = []
scores4 = []
l=0
while l<10:
#1999
df1 = train_df1.iloc[test_index1[l],:]
dft1=train_df1.iloc[test_index1[l],:]
df_train1 = df1
df_test1 = dft1
#cols_leave = ['Data.Time', 'Data.status','Data.B4', 'Data.V025', 'Data.V149', 'Data.V190']
cols_leave = ['Data.Time', 'Data.status','Data.b4', 'Data.v025', 'Data.v149']
leave = [(col, None) for col in cols_leave]
x_mapper = DataFrameMapper(leave)
x_train1 = x_mapper.fit_transform(df_train1).astype('float32')
x_test1 = x_mapper.transform(df_test1).astype('float32')
get_target = lambda df: (df['Data.Time'].values, df['Data.status'].values)
y_train1 = get_target(df_train1)
durations_test1, events_test1 = get_target(df_test1)
##2006
df2 = train_df2.iloc[test_index2[l],:]
dft2=train_df2.iloc[test_index2[l],:]
df_train2 = df2
df_test2 = dft2
cols_leave = ['Data.Time', 'Data.status','Data.B4', 'Data.V025', 'Data.V149', 'Data.V190']
leave = [(col, None) for col in cols_leave]
x_mapper = DataFrameMapper(leave)
x_train2 = x_mapper.fit_transform(df_train2).astype('float32')
x_test2 = x_mapper.transform(df_test2).astype('float32')
get_target = lambda df: (df['Data.Time'].values, df['Data.status'].values)
y_train2 = get_target(df_train2)
durations_test2, events_test2 = get_target(df_test2)
##2011
df3 = train_df3.iloc[test_index3[l],:]
dft3=train_df3.iloc[test_index3[l],:]
df_train3 = df3
df_test3 = dft3
cols_leave = ['Data.Time', 'Data.status','Data.B4', 'Data.V025', 'Data.V149', 'Data.V190']
leave = [(col, None) for col in cols_leave]
x_mapper = DataFrameMapper(leave)
x_train3 = x_mapper.fit_transform(df_train3).astype('float32')
x_test3 = x_mapper.transform(df_test3).astype('float32')
get_target = lambda df: (df['Data.Time'].values, df['Data.status'].values)
y_train3 = get_target(df_train3)
durations_test3, events_test3 = get_target(df_test3)
##2015
#df4 = train_df4.iloc[test_index4[l],:]
#dft4=train_df4.iloc[test_index4[l],:]
#df_train4 = df4
#df_test4 = dft4
#cols_leave = ['Data.Time', 'Data.status','Data.B4', 'Data.V025', 'Data.V149', 'Data.V190']
#leave = [(col, None) for col in cols_leave]
#x_mapper = DataFrameMapper(leave)
#x_train4 = x_mapper.fit_transform(df_train4).astype('float32')
#x_test4 = x_mapper.transform(df_test4).astype('float32')
#get_target = lambda df: (df['Data.Time'].values, df['Data.status'].values)
#y_train4 = get_target(df_train4)
#durations_test4, events_test4 = get_target(df_test4)
# model parameters
dropout = 0.2
epochs = 1000
structure = [ {'activation': 'ReLU', 'num_units': 128}, {'activation': 'ReLU', 'num_units': 128},{'activation': 'ReLU', 'num_units': 128},{'activation': 'ReLU', 'num_units': 128}]
# model 1
model1 =NonLinearCoxPHModel(structure=structure)
model1.fit(x_train1, y_train1[0],y_train1[1], lr=1e-8,num_epochs = epochs, dropout = dropout,init_method='xav_uniform',l2_reg=1e-4, batch_normalization=True)
# model 2
model2 =NonLinearCoxPHModel(structure=structure)
model2.fit(x_train2, y_train2[0],y_train2[1], lr=1e-8,num_epochs = epochs, dropout = dropout,init_method='xav_uniform',l2_reg=1e-4,batch_normalization=True)
# model 3
model3 =NonLinearCoxPHModel(structure=structure)
model3.fit(x_train3, y_train3[0],y_train3[1], lr=1e-8,num_epochs = epochs, dropout = dropout,init_method='xav_uniform',l2_reg=1e-4,batch_normalization=True)
# model 4
#model4 =NonLinearCoxPHModel(structure=structure)
#model4.fit(x_train4, y_train4[0],y_train4[1], lr=1e-8,num_epochs = epochs, dropout = dropout,init_method='xav_uniform',l2_reg=1e-4, batch_normalization=True)
# C-index
#1999
scores1.append(concordance_index(model1,x_test1,durations_test1,events_test1))
#2006
scores2.append(concordance_index(model2,x_test2,durations_test2,events_test2))
#2011
scores3.append(concordance_index(model3,x_test3,durations_test3,events_test3))
#2015
#scores4.append(concordance_index(model4,x_test4,durations_test4,events_test4))
l+=1
#plt.boxplot(scores)
#deepsurv.plot_log(metrics)
# + [markdown] id="9C0pFf07PSm8"
# Store results
# + id="zps2pf2-PWGd"
ChadResultsNew = pd.DataFrame()
ChadResultsNew['1996'] = scores1
ChadResultsNew['2004'] = scores2
ChadResultsNew['2014'] = scores3
#GhanaResultsNew['2014'] = scores4
ChadResultsNew
ChadResultsNew.to_csv('ChadResultsNew.csv')
# + [markdown] id="OphXar9_PXMJ"
# Plot results
# + id="wk29UfkyP5MU"
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sns
import itertools
import pandas as pd
import plotly.graph_objects as go
import matplotlib.pyplot as plt
import seaborn as sns, numpy as np
from pylab import *
# + id="4_3FAdfxQHXj" colab={"base_uri": "https://localhost:8080/", "height": 683} outputId="9d984d8e-10f2-4d54-f4d1-eccd399df2f2"
# One country from each region
sns.set(rc={"figure.figsize": (16, 10)})
plt.suptitle("10-fold Crossvalidated C-index")
subplot(2,2,1)
# Zimbabwe
RSF_zim = 'Zim_RSF.csv'
RSF_zim = pd.read_csv(RSF_zim)
New = pd.DataFrame(RSF_zim)
ZimbabweResults = 'ZimbabweResultsNew.csv'
ZimbabweResults = pd.read_csv(ZimbabweResults)
ZimbabweResults= ZimbabweResults.drop(['1999'], axis=1)
df = pd.concat([ZimbabweResults, New])
g = ['Deepsurv','RSF']
K =10
model = list(itertools.chain.from_iterable(itertools.repeat(i, K) for i in g))
df["model"] = model
df = pd.melt(df, id_vars=['model'], value_vars=[ '2006','2011','2015'])
df.columns = ['Model', 'Year', 'C-index']
ax= sns.boxplot(y='C-index', x='Year',
data=df,
palette="colorblind",
hue='Model',showmeans=True,meanprops={"marker":"o",
"markerfacecolor":"white",
"markeredgecolor":"black",
"markersize":"10"})
ax.set(xlabel=None)
ax.set(title='Zimbabwe')
ax.set_ylim([0.2,1])
subplot(2,2,2)
# Uganda
RSF_Uganda = 'Uganda_RSF.csv'
RSF_Uganda = pd.read_csv(RSF_Uganda)
New = pd.DataFrame(RSF_Uganda)
UgandaResults = 'UgandaResultsNew.csv'
UgandaResults = pd.read_csv(UgandaResults)
UgandaResults = UgandaResults.drop(['2001'], axis=1)
df = pd.concat([UgandaResults, New])
g = ['Deepsurv','RSF']
K =10
model = list(itertools.chain.from_iterable(itertools.repeat(i, K) for i in g))
df["model"] = model
df = pd.melt(df, id_vars=['model'], value_vars=['2006','2011','2016'])
df.columns = ['Model', 'Year', 'C-index']
ax= sns.boxplot(y='C-index', x='Year',
data=df,
palette="colorblind",
hue='Model',showmeans=True,meanprops={"marker":"o",
"markerfacecolor":"white",
"markeredgecolor":"black",
"markersize":"10"})
ax.set(ylabel=None)
ax.set(xlabel=None)
ax.set(title='Uganda')
ax.set_ylim([0.2,1])
subplot(2,2,3)
# Ghana
RSF_Ghana = 'Ghana_RSF.csv'
RSF_Ghana = pd.read_csv(RSF_Ghana)
New = pd.DataFrame(RSF_Ghana)
GhanaResults = 'GhanaResultsNew.csv'
GhanaResults = pd.read_csv(GhanaResults)
GhanaResults = GhanaResults.drop(['1998'], axis=1)
df = pd.concat([GhanaResults, New])
g = ['Deepsurv','RSF']
K =10
model = list(itertools.chain.from_iterable(itertools.repeat(i, K) for i in g))
df["model"] = model
df = pd.melt(df, id_vars=['model'], value_vars=['2003','2008','2014'])
df.columns = ['Model', 'Year', 'C-index']
ax = sns.boxplot(y='C-index', x='Year',
data=df,
palette="colorblind",
hue='Model',showmeans=True,meanprops={"marker":"o",
"markerfacecolor":"white",
"markeredgecolor":"black",
"markersize":"10"})
ax.set(title='Ghana')
ax.set_ylim([0.2,1])
subplot(2,2,4)
# Chad
RSF_Chad = 'Chad_RSF.csv'
RSF_Chad = pd.read_csv(RSF_Chad)
New = pd.DataFrame(RSF_Chad)
ChadResults = 'ChadResultsNew.csv'
ChadResults = pd.read_csv(ChadResults)
ChadResults = ChadResults.drop(['1996'], axis=1)
df = pd.concat([ChadResults, New])
g = ['Deepsurv','RSF']
K =10
model = list(itertools.chain.from_iterable(itertools.repeat(i, K) for i in g))
df["model"] = model
df = pd.melt(df, id_vars=['model'], value_vars=['2004','2014'])
df.columns = ['Model', 'Year', 'C-index']
ax = sns.boxplot(y='C-index', x='Year',
data=df,
palette="colorblind",
hue='Model',showmeans=True,meanprops={"marker":"o",
"markerfacecolor":"white",
"markeredgecolor":"black",
"markersize":"10"})
ax.set(ylabel=None)
ax.set(title='Chad')
ax.set_ylim([0.2,1])
#plt.show()
# save the plot as a file
plt.savefig('CVCINDEX.tif', transparent=True, dpi=350, bbox_inches="tight", pad_inches=0.0)
# + id="EDsCG_98ykPD"
train_dataset_fp1 = 'New2001Uganda.csv'
train_dataset_fp2 = 'New2006Uganda.csv'
train_dataset_fp3 = 'New2011Uganda.csv'
train_dataset_fp4 = 'New2016Uganda.csv'
train_dataset_fp1 = 'New1999Zim.csv'
train_dataset_fp2 = 'New2006Zim.csv'
train_dataset_fp3 = 'New2011Zim.csv'
train_dataset_fp4 = 'New2015Zim.csv'
train_dataset_fp1 = 'New1998Ghana.csv'
train_dataset_fp2 = 'New2003Ghana.csv'
train_dataset_fp3 = 'New2008Ghana.csv'
train_dataset_fp4 = 'New2014Ghana.csv'
train_dataset_fp1 = 'New1996Chad.csv'
train_dataset_fp2 = 'New2004Chad.csv'
train_dataset_fp3 = 'New2014Chad.csv'
Results
ZimbabweResultsNew = pd.DataFrame()
ZimbabweResultsNew['1999'] = scores1
ZimbabweResultsNew['2006'] = scores2
ZimbabweResultsNew['2011'] = scores3
ZimbabweResultsNew['2015'] = scores4
ZimbabweResultsNew
#ZimbabweResultsNew.to_csv('ZimbabweResultsNew.csv')
UgandaResultsNew = pd.DataFrame()
UgandaResultsNew['2001'] = scores1
UgandaResultsNew['2006'] = scores2
UgandaResultsNew['2011'] = scores3
UgandaResultsNew['2016'] = scores4
UgandaResultsNew
#UgandaResultsNew.to_csv('UgandaResultsNew.csv')
GhanaResultsNew = pd.DataFrame()
GhanaResultsNew['1998'] = scores1
GhanaResultsNew['2003'] = scores2
GhanaResultsNew['2008'] = scores3
GhanaResultsNew['2014'] = scores4
GhanaResultsNew
#GhanaResultsNew.to_csv('GhanaResultsNew.csv')
ChadResultsNew = pd.DataFrame()
ChadResultsNew['1996'] = scores1
ChadResultsNew['2004'] = scores2
ChadResultsNew['2014'] = scores3
#GhanaResultsNew['2014'] = scores4
ChadResultsNew
#ChadResultsNew.to_csv('ChadResultsNew.csv')
# + colab={"base_uri": "https://localhost:8080/", "height": 417} id="lfQ9E5km83yL" outputId="ee98d88c-1c33-4bb0-8cb0-ba1da640ad07"
train_df2
| PYTHONCODE.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import numpy as np
# Read in the image
image = mpimg.imread('imagedata/test.jpg')
# Grab the x and y size and make a copy of the image
ysize = image.shape[0]
xsize = image.shape[1]
color_select = np.copy(image)
line_image = np.copy(image)
# Define color selection criteria
# MODIFY THESE VARIABLES TO MAKE YOUR COLOR SELECTION
red_threshold = 200
green_threshold = 200
blue_threshold = 200
rgb_threshold = [red_threshold, green_threshold, blue_threshold]
# Define the vertices of a triangular mask.
# Keep in mind the origin (x=0, y=0) is in the upper left
# MODIFY THESE VALUES TO ISOLATE THE REGION
# WHERE THE LANE LINES ARE IN THE IMAGE
left_bottom = [0, 539]
right_bottom = [900, 539]
apex = [450,300]
# Perform a linear fit (y=Ax+B) to each of the three sides of the triangle
# np.polyfit returns the coefficients [A, B] of the fit
fit_left = np.polyfit((left_bottom[0], apex[0]), (left_bottom[1], apex[1]), 1)
fit_right = np.polyfit((right_bottom[0], apex[0]), (right_bottom[1], apex[1]), 1)
fit_bottom = np.polyfit((left_bottom[0], right_bottom[0]), (left_bottom[1], right_bottom[1]), 1)
# Mask pixels below the threshold
color_thresholds = (image[:,:,0] < rgb_threshold[0]) | \
(image[:,:,1] < rgb_threshold[1]) | \
(image[:,:,2] < rgb_threshold[2])
# Find the region inside the lines
XX, YY = np.meshgrid(np.arange(0, xsize), np.arange(0, ysize))
region_thresholds = (YY > (XX*fit_left[0] + fit_left[1])) & \
(YY > (XX*fit_right[0] + fit_right[1])) & \
(YY < (XX*fit_bottom[0] + fit_bottom[1]))
# Mask color and region selection
color_select[color_thresholds | ~region_thresholds] = [0, 0, 0]
# Color pixels red where both color and region selections met
line_image[~color_thresholds & region_thresholds] = [255, 0, 0]
# Display the image and show region and color selections
plt.imshow(image)
x = [left_bottom[0], right_bottom[0], apex[0], left_bottom[0]]
y = [left_bottom[1], right_bottom[1], apex[1], left_bottom[1]]
plt.plot(x, y, 'b--', lw=4)
plt.imshow(color_select)
plt.imshow(line_image)
plt.show()
| Term1/Color Region.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pyvista as pv
mesh = pv.read("pdata_C1048.vtk")
mesh
cc = mesh.cell_centers()
cc
# +
# %%timeit
mesh.cell_centers()
# -
| poc-3/data/test/synthetic/benchmark_cell_centers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Observations and Insights
#
# #1. The study was conducted with an equal sampling of mice based on gender dwhich could be beneficial if any of the tumors
# #2. Mice treated with the drug Capomulin had the lowest/least growth of tumors over the 45 day study.
# #3. Capomulin was tested almost 2 to 1 as the drug Propriva
# +
# Dependencies and Setup
import matplotlib.pyplot as plt
import pandas as pd
from pandas import DataFrame, Series
import scipy.stats as st
import numpy as numpy
# Study data files
mouse_metadata_path = "data/Mouse_metadata.csv"
study_results_path = "data/Study_results.csv"
# Read the mouse data and the study results
mouse_metadata = pd.read_csv(mouse_metadata_path)
study_results = pd.read_csv(study_results_path)
# Combine the data into a single dataset
test_df=pd.merge(mouse_metadata,study_results,on="Mouse ID")
# Display the data table for preview
test_df.head()
# -
# Checking the number of mice.
total_mice = len(test_df['Mouse ID'].unique())
total_mice
# Getting the duplicate mice by ID number that shows up for Mouse ID and Timepoint.
test_df.loc[test_df.duplicated(subset=['Mouse ID', 'Timepoint'], keep='first')]
# Optional: Get all the data for the duplicate mouse ID.
test_df[test_df["Mouse ID"]=="g989"]
# Create a clean DataFrame by dropping the duplicate mouse by its ID.
clean_df = test_df[test_df["Mouse ID"]!="g989"]
clean_df
# Checking the number of mice in the clean DataFrame.
clean_mice = len(clean_df['Mouse ID'].unique())
clean_mice
# ## Summary Statistics
# Generate a summary statistics table of mean, median, variance, standard deviation, and SEM of the tumor volume for each regimen
# Use groupby and summary statistical methods to calculate the following properties of each drug regimen:
# mean, median, variance, standard deviation, and SEM of the tumor volume.
# Assemble the resulting series into a single summary dataframe.
mean = clean_df.groupby('Drug Regimen')['Tumor Volume (mm3)'].mean()
median = clean_df.groupby('Drug Regimen')['Tumor Volume (mm3)'].median()
variance = clean_df.groupby('Drug Regimen')['Tumor Volume (mm3)'].var()
stdev = clean_df.groupby('Drug Regimen')['Tumor Volume (mm3)'].std()
sem = clean_df.groupby('Drug Regimen')['Tumor Volume (mm3)'].sem()
# Generate a summary statistics table of mean, median, variance, standard deviation, and SEM of the tumor volume for each regimen
sum_stat = pd.DataFrame({
"Mean": mean,
"Median": median,
"Variance": variance,
"Standard Deviation": stdev,
"SEM": sem
}
)
sum_stat
# Using the aggregation method, produce the same summary statistics in a single line
sum_stat.aggregate(['sum'])
# +
# Generate a bar plot showing the total number of measurements taken on each drug regimen using pandas.
# Find the total number of drug measurements
counts = clean_df['Drug Regimen'].value_counts()
counts
# +
# Plot the measurements on a bar plot using Pandas
counts.plot(kind="bar")
plt.xlabel("Test Drugs")
plt.xticks(rotation=90)
plt.ylabel("Number of Measurements")
plt.title("Measurements per Drug Regimine")
plt.show()
# -
# Plot measurements taken on each drug regimen using pyplot.
counts = clean_df['Drug Regimen'].value_counts()
plt.bar(counts.index.values,counts.values)
plt.xlabel("Study Drugs")
plt.title("Measurments by Drug")
plt.xticks(rotation=90)
plt.ylabel("Total Number of Measurements")
plt.show()
# +
# Get totals of Mice in test study
gendergp = clean_df.groupby('Sex')
genders = gendergp["Mouse ID"].nunique()
percentgp = (genders / clean_mice)*100
genderdf = pd.DataFrame(
{'Percentage of Mice': percentgp,
'Total Counts': genders
}
)
genderdf['Percentage of Mice'] = genderdf['Percentage of Mice'].map('{:.2f}%'.format)
genderdf
# +
# Generate a pie plot showing the distribution of female versus male mice using pandas
genderdf = pd.DataFrame(
{'Percentage of Mice': percentgp,
'Total Counts': genders
}
)
plot = genderdf.plot.pie(y='Percentage of Mice')
plt.legend
plt.show()
# +
# Generate a pie plot showing the distribution of female versus male mice using pyplot
# Generate a pie plot showing the distribution of female versus male mice using pyplot
labels = ["Males", "Females"]
colors = ["red", "green"]
explode = [0, 0.05]
plt.pie(genders, labels=labels, autopct="%1.1f%%", startangle=45, colors=colors, explode=explode)
plt.title ("Gender Ratio of Study Mice")
plt.show()
# -
# ## Quartiles, Outliers and Boxplots
# +
# Calculate the final tumor volume of each mouse across four of the treatment regimens:
# Capomulin, Ramicane, Infubinol, and Ceftamin
# Start by getting the last (greatest) timepoint for each mouse
drug_tv = clean_df.sort_values(["Drug Regimen", "Mouse ID", "Timepoint"], ascending=True)
# Select final volume of each mouse
final_tv_df = clean_df.loc[clean_df["Timepoint"] == 45]
final_tv_df.head()
# -
max_mouse_timepoint = clean_df.groupby('Mouse ID')['Timepoint'].max()
max_mouse_timepoint.reset_index()
# +
# Merge this group df with the original dataframe to get the tumor volume at the last timepoint
merged_df = pd.merge(max_mouse_timepoint, clean_df, how = 'left', on= ['Mouse ID','Timepoint'])
merged_df.tail(25)
# +
# Put treatments into a list for for loop (and later for plot labels)
treatments = ["Capomulin", "Ramicane", "Infubinol", "Ceftamin"]
# Create empty list to fill with tumor vol data (for plotting)
tumor_vol_data = []
# Calculate the IQR and quantitatively determine if there are any potential outliers.
for treat in treatments:
final_vol = merged_df.loc[merged_df['Drug Regimen']==treat, 'Tumor Volume (mm3)']
tumor_vol_data.append(final_vol)
# Find the Quartiles
quartiles = final_vol.quantile([.25,.5,.75])
lowerq = quartiles[0.25]
upperq = quartiles[0.75]
iqr = upperq-lowerq
print(f"The lower quartile of treatment is: {lowerq}")
print(f"The upper quartile of treatment is: {upperq}")
print(f"The interquartile range of treatment is: {iqr}")
print(f"The the median of treatment is: {quartiles[0.5]} ")
lower_bound = lowerq - (1.5*iqr)
upper_bound = upperq + (1.5*iqr)
print(f"Values below {lower_bound} could be outliers.")
print(f"Values above {upper_bound} could be outliers.")
# -
# Generate a box plot of the final tumor volume of each mouse across four regimens of interest
fig1, ax1 = plt.subplots()
ax1.set_title('Final Tumor Volume')
ax1.set_ylabel('Drug Regimen')
ax1.boxplot(tumor_vol_data)
plt.show()
# ## Line and Scatter Plots
# +
# Generate a line plot of tumor volume vs. time point for a mouse treated with Capomulin
t_vol = clean_df.loc[clean_df['Drug Regimen'] =="Capomulin"]
t_point = t_vol.loc[t_vol['Mouse ID']=='m601']
plt.plot (t_point['Timepoint'],t_point['Tumor Volume (mm3)'])
plt.xlabel('Timepoint (days)')
plt.ylabel('Tumor Volume (mm3)')
plt.title('Capomulin data of Mouse m601')
plt.show()
# +
# Generate a scatter plot of average tumor volume vs. mouse weight for the Capomulin regimen
#avg_t_vol = clean_df.loc[clean_df['Drug Regimen'] =="Capomulin"]
#cap_weight = avg_t_vol.loc[avg_t_vol['Mouse ID']=='m601']
plt.scatter(clean_df.iloc[:,5],clean_df.iloc[:,2])
plt.xlabel('Income Per Capita')
plt.ylabel('Average Alcohol Consumed Per Person Per Year (L)')
plt.show()
# -
# ## Correlation and Regression
# Calculate the correlation coefficient and linear regression model for mouse weight and average tumor volume for the Capomulin regimen
weight = clean_df.iloc[:,4]
tumor = clean_df.iloc[:,5]
correlation = st.pearsonr(weight,tumor)
print(f"The correlation between both factors is {round(correlation[0],2)}")
| Pymaceuticals/pymaceuticals.ipynb |
# ---
# jupyter:
# jupytext:
# formats: ipynb,md:myst
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] tags=[]
# # Transformation from/to IR
#
# $$
# \newcommand{\iv}{{\mathrm{i}\nu}}
# \newcommand{\wmax}{{\omega_\mathrm{max}}}
# \newcommand{\dd}{{\mathrm{d}}}
# $$
#
# In this section, we explain how to transform numerical data to IR.
# -
# ## Poles
# We consider a Green's function genereated by poles:
#
# $$
# G(\mathrm{i}\nu) = \sum_{p=1}^{N_\mathrm{P}} \frac{c_p}{\mathrm{i}\nu - \omega_p},
# $$
#
# where $\nu$ is a fermionic or bosonic Matsubara frequency.
# The corresponding specral function $A(\omega)$ is given by
#
# $$
# A(\omega) = \sum_{p=1}^{N_\mathrm{P}} c_p \delta(\omega - \omega_p).
# $$
#
# The modified (regularized) spectral function reads
#
# $$
# \rho(\omega) =
# \begin{cases}
# \sum_{p=1}^{N_\mathrm{P}} c_p \delta(\omega - \omega_p) & \mathrm{(fermion)},\\
# \sum_{p=1}^{N_\mathrm{P}} (c_p/\tanh(\beta \omega_p/2)) \delta(\omega - \omega_p) & \mathrm{(boson)}.
# \end{cases}
# $$
#
# for the logistic kernel.
# We immediately obtain
#
# $$
# \rho_l =
# \begin{cases}
# \sum_{p=1}^{N_\mathrm{P}} c_p V_l(\omega_p) & \mathrm{(fermion)},\\
# \sum_{p=1}^{N_\mathrm{P}} c_p V_l(\omega_p)/\tanh(\beta \omega_p/2))& \mathrm{(boson)}.
# \end{cases}
# $$
#
# The following code demostrates this transformation for bosons.
# +
import sparse_ir
import numpy as np
# %matplotlib inline
import matplotlib.pyplot as plt
plt.rcParams['font.size'] = 15
beta = 15
wmax = 10
basis_b = sparse_ir.FiniteTempBasis("B", beta, wmax, eps=1e-10)
coeff = np.array([1])
omega_p = np.array([0.1])
rhol_pole = np.einsum('lp,p->l', basis_b.v(omega_p), coeff/np.tanh(0.5*beta*omega_p))
gl_pole = - basis_b.s * rhol_pole
plt.semilogy(np.abs(rhol_pole), marker="o", label=r"$|\rho_l|$")
plt.semilogy(np.abs(gl_pole), marker="x", label=r"$|g_l|$")
plt.xlabel(r"$l$")
plt.ylim([1e-5, 1e+1])
plt.legend(frameon=False)
plt.show()
# -
# Alternatively, we can use ``spr`` (sparse pole presentation) module.
# +
from sparse_ir.spr import SparsePoleRepresentation
sp = SparsePoleRepresentation(basis_b, omega_p)
gl_pole2 = sp.to_IR(coeff)
plt.semilogy(np.abs(gl_pole2), marker="x", label=r"$|g_l|$ from SPR")
plt.semilogy(np.abs(gl_pole), marker="x", label=r"$|g_l|$")
plt.xlabel(r"$l$")
plt.ylim([1e-5, 1e+1])
plt.legend(frameon=False)
plt.show()
# -
# ## From smooth spectral function
#
# For a smooth spectral function $\rho(\omega)$, the expansion coefficients can be evaluated by computing the integral
#
# $$
# \rho_l = \int_{-\omega_\mathrm{max}}^{\omega_\mathrm{max}} \mathrm{d} \omega V_l(\omega) \rho(\omega).
# $$
#
# One might consider to use the Gauss-Legendre quadrature.
# As seen in previous sections, the distribution of $V_l(\omega)$ is much denser than Legendre polynomial $P_l(x(\tau))$ around $\tau=0, \beta$.
# Thus, evaluating the integral precisely requires the use of composite Gauss–Legendre quadrature,
# where the whole inteval $[-\omega_\mathrm{max}, \omega_\mathrm{max}]$ is divided to subintervals and the normal Gauss-Legendre quadrature is
# applied to each interval.
# The roots of $V_l(\omega)$ for the highest $l$ used in the expansion
# is a reasonable choice of the division points.
# If $\rho(\omega)$ is smooth enough within each subinterval,
# the result converges exponentially with increasing the degree of the Gauss-Legendre quadrature.
#
# Below, we demonstrate how to compute $\rho_l$ for a spectral function consisting of of three Gausssian peaks using the composite Gauss-Legendre quadrature.
# Then, $\rho_l$ can be transformed to $g_l$ by multiplying it with $- S_l$.
# +
# Three Gaussian peaks (normalized to 1)
gaussian = lambda x, mu, sigma:\
np.exp(-((x-mu)/sigma)**2)/(np.sqrt(np.pi)*sigma)
rho = lambda omega: 0.2*gaussian(omega, 0.0, 0.15) + \
0.4*gaussian(omega, 1.0, 0.8) + 0.4*gaussian(omega, -1.0, 0.8)
omegas = np.linspace(-5, 5, 1000)
plt.xlabel(r"$\omega$")
plt.ylabel(r"$\rho(\omega)$")
plt.plot(omegas, rho(omegas))
plt.show()
# +
beta = 10
wmax = 10
basis = sparse_ir.FiniteTempBasis("F", beta, wmax, eps=1e-10)
rhol = basis.v.overlap(rho)
gl = - basis.s * rhol
plt.semilogy(np.abs(rhol), marker="o", ls="", label=r"$|\rho_l|$")
plt.semilogy(np.abs(gl), marker="s", ls="", label=r"$|g_l|$")
plt.semilogy(np.abs(basis.s), marker="", ls="--", label=r"$S_l$")
plt.xlabel(r"$l$")
plt.ylim([1e-5, 10])
plt.legend(frameon=False)
plt.show()
#plt.savefig("coeff.pdf")
# -
# $\rho_l$ is evaluated on arbitrary real frequencies as follows.
# +
rho_omgea_reconst = basis.v(omegas).T @ rhol
plt.xlabel(r"$\omega$")
plt.ylabel(r"$\rho(\omega)$")
plt.plot(omegas, rho_omgea_reconst)
plt.show()
# -
# ## From IR to imaginary time
#
# We are now ready to evaluate $g_l$ on arbitrary $\tau$ points.
# A naive way is as follows.
taus = np.linspace(0, beta, 1000)
gtau1 = basis.u(taus).T @ gl
plt.plot(taus, gtau1)
plt.xlabel(r"$\tau$")
plt.ylabel(r"$G(\tau)$")
plt.show()
# Alternatively, we can use ``TauSampling`` as follows.
smpl = sparse_ir.TauSampling(basis, taus)
gtau2 = smpl.evaluate(gl)
plt.plot(taus, gtau1)
plt.xlabel(r"$\tau$")
plt.ylabel(r"$G(\tau)$")
plt.show()
# ## From full imaginary-time data
#
# A numerically stable way to expand $G(\tau)$ in IR
# is evaluating the integral
#
# $$
# G_l = \int_0^\beta \mathrm{d} \tau G(\tau) U_l(\tau).
# $$
#
# You can use `overlap` function as well.
# +
def eval_gtau(taus):
uval = basis.u(taus) #(nl, ntau)
if isinstance(taus, np.ndarray):
print(uval.shape, gl.shape)
return uval.T @ gl
else:
return uval.T @ gl
gl_reconst = basis.u.overlap(eval_gtau)
ls = np.arange(basis.size)
plt.semilogy(ls[::2], np.abs(gl_reconst)[::2], label="reconstructed", marker="+", ls="")
plt.semilogy(ls[::2], np.abs(gl)[::2], label="exact", marker="x", ls="")
plt.semilogy(ls[::2], np.abs(gl_reconst - gl)[::2], label="error", marker="p")
plt.xlabel(r"$l$")
plt.xlabel(r"$|g_l|$")
plt.ylim([1e-20, 1])
plt.legend(frameon=False)
plt.show()
# -
# ## Remark: What happens if $\omega_\mathrm{max}$ is too small?
#
# If $G_l$ do not decay like $S_l$, $\omega_\mathrm{max}$ may be too small.
# Below, we numerically demonstrate it.
# +
from scipy.integrate import quad
beta = 10
wmax = 0.5
basis_bad = sparse_ir.FiniteTempBasis("F", beta, wmax, eps=1e-10)
# We expand G(τ).
gl_bad = [quad(lambda x: eval_gtau(x) * basis_bad.u[l](x), 0, beta)[0] for l in range(basis_bad.size)]
plt.semilogy(np.abs(gl_bad), marker="s", ls="", label=r"$|g_l|$")
plt.semilogy(np.abs(basis_bad.s), marker="x", ls="--", label=r"$S_l$")
plt.xlabel(r"$l$")
plt.ylim([1e-5, 10])
#plt.xlim([0, basis.size])
plt.legend(frameon=False)
plt.show()
#plt.savefig("coeff_bad.pdf")
# -
# ## Matrix-valued object
#
# `evaluate` and `fit` accept a matrix-valued object as an input.
# The axis to which the transformation applied can be specified by using the keyword augment `axis`.
np.random.seed(100)
shape = (1,2,3)
gl_tensor = np.random.randn(*shape)[..., np.newaxis] * gl[np.newaxis, :]
print("gl: ", gl.shape)
print("gl_tensor: ", gl_tensor.shape)
smpl_matsu = sparse_ir.MatsubaraSampling(basis)
gtau_tensor = smpl_matsu.evaluate(gl_tensor, axis=3)
print("gtau_tensor: ", gtau_tensor.shape)
gl_tensor_reconst = smpl_matsu.fit(gtau_tensor, axis=3)
assert np.allclose(gl_tensor, gl_tensor_reconst)
| src/transformation_py.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Simple D3 example
#
# The following demonstration of D3 usage comes from
# [Custom D3.js Visualization in a Jupyter Notebook](https://www.stefaanlippens.net/jupyter-custom-d3-visualization.html).
# To cover the basics, let's start simple with just inline `%%javascript` snippet cells.
#
# First, we tell the RequireJS environment where to find the version of D3.js we want. Note that the .js extension is omitted in the URL.
# + language="javascript"
# require.config({
# paths: {
# d3: 'https://d3js.org/d3.v5.min'
# }
# });
# -
# We can now create a D3.js powered SVG drawing, for example as follows:
# + language="javascript"
# (function(element) {
# require(['d3'], function(d3) {
# var data = [1, 2, 4, 8, 16, 8, 4, 2, 1]
#
# var svg = d3.select(element.get(0)).append('svg')
# .attr('width', 400)
# .attr('height', 200);
# svg.selectAll('circle')
# .data(data)
# .enter()
# .append('circle')
# .attr("cx", function(d, i) {return 40 * (i + 1);})
# .attr("cy", function(d, i) {return 100 + 30 * (i % 3 - 1);})
# .style("fill", "#1570a4")
# .transition().duration(2000)
# .attr("r", function(d) {return 2*d;})
# ;
# })
# })(element);
# -
| notebooks/misc/D3-simple-example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from opentree import OT
import dendropy as dp
import re, os, copy, shutil
from ipywidgets import interactive, FloatSlider
from IPython.display import display, Javascript
import ipywidgets as widgets
import functools
from ipywidgets import VBox, HBox
import pandas as pd
wdir=0
inputfile=0
outputfile=0
os.getcwd()
# +
#modified from https://www.geeksforgeeks.org/remove-invalid-parentheses/
# Python3 program to remove invalid parenthesis
# Method checks if character is parenthesis(open
# or closed)
def isParenthesis(c):
return ((c == '(') or (c == ')'))
# method returns true if contains valid
# parenthesis
def isValidString(str):
cnt = 0
for i in range(len(str)):
if (str[i] == '('):
cnt += 1
elif (str[i] == ')'):
cnt -= 1
if (cnt < 0):
return False
return (cnt == 0)
# method to remove invalid parenthesis
def removeInvalidParenthesis(str):
if (len(str) == 0):
return
# visit set to ignore already visited
visit = set()
# queue to maintain BFS
q = []
temp = 0
level = 0
# pushing given as starting node into queu
q.append(str)
visit.add(str)
while(len(q)):
str = q[0]
q.pop()
if (isValidString(str)):
#print(str)
# If answer is found, make level true
# so that valid of only that level
# are processed.
level = True
if (level):
continue
for i in range(len(str)):
if (not isParenthesis(str[i])):
continue
# Removing parenthesis from str and
# pushing into queue,if not visited already
temp = str[0:i] + str[i + 1:]
if temp not in visit:
q.append(temp)
visit.add(temp)
return str
# +
#speciesfile- excel
def make_tree():
os.chdir(wdir)
otts=[]
names_df=pd.read_csv(inputfile)
names=names_df['Scientific name'].tolist()
print(names)
for i in range(len(names)):
otts.append(OT.get_ottid_from_name(names[i]))
ret = OT.synth_induced_tree(ott_ids = otts,
label_format="name")
treefile = "unstripped.txt"
ret.tree.write(path = treefile, schema = "newick")
ret.tree.print_plot(width=100)
stripped_tree = dp.Tree.get(file=open("unstripped.txt"), schema="newick")
taxon_names = [s.label for s in stripped_tree.taxon_namespace]
# Extract tree with labels. This will remove all of the empty singleton
# entries that otl brings down.
final_tree = stripped_tree.extract_tree_with_taxa_labels(taxon_names)
file1 = open("cypriniformes_speciestree.txt","w")#write mode
file1.write(str(final_tree))
file1.close()
str_tree=str(final_tree)
#print('string tree: \n',str_tree)
for_brackets=re.split('(\W+)',str_tree)
count=-1
for i in for_brackets:
count=count+1
if i=='':
for_brackets.pop(count)
if 'mrcaott' in i:
for_brackets.pop(count)
count=-1
for i in for_brackets:
count=count+1
#print(i[0].isupper())
if i[0].isupper()==False and '_' in i:
for_brackets.pop(count)
#print(for_brackets)
count=-1
for i in names:
count=count+1
j=i.replace(' ','_')
names[count]=j
#print('new names:{} \n'.format(len(names)),names)
count=-1
good_tree=[]
for i in for_brackets:
count=count+1
if '_' in i:
for j in names:
if i[0:10] in j:
good_tree.append(j)
#print('appending {}'.format(j))
else:
if 'Pan' in i:
good_tree.append('Pan_Troglo')
if '(' not in i and ')' not in i and ',' not in i:
print ('deleting {}'.format(i))
else:
good_tree.append(i)
#print('appending {}'.format(i))
good_tree=''.join(good_tree)
good_tree=good_tree.replace(',)','')
good_tree=good_tree.replace('()','')
good_tree=removeInvalidParenthesis(good_tree)
print(good_tree)
with open(outputfile,'w') as text_file:
text_file.write(good_tree)
return good_tree
# +
def run_next(ev,button):
global wdir, inputfile, outputfile
wdir=wdir_w.value
inputfile=inputfile_w.value
outputfile=outputfile_w.value
display(Javascript('IPython.notebook.execute_cells([IPython.notebook.get_selected_index()+1])'))
wdir_w=widgets.Text(value='/Users/ivan/Documents/Raleigh_lab/dnds', description= 'directory with species file')
inputfile_w=widgets.Text(value='IAPP_orthologs.csv', description= 'species csv file')
outputfile_w=widgets.Text(value='species_tree_IAPP_procedural', description= 'output file name')
button_js=widgets.Button(description="Build tree")
button_js.on_click(functools.partial(run_next,button=button_js))
box=HBox([wdir_w,inputfile_w,outputfile_w,button_js])
#print'''Please input the path to the directory containing the allignment
#Note that it must be as /Users/name/directory rather than ~/directory
#Output file can have any name, python will creatte it for you
#Currently only takes .phy files, 10 char trimmed (not padded)
#Press 'Convert file' and the output .txt will appear in your specified directory '''
display(box)
# -
make_tree()
| updated_treegrabber.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # "YouTube Talk: AdaHessian Optimizer, With The Authors"
#
# > I recently hosted a paper presentation of the AdaHessian optimizer from its authors
#
# - badges: false
# - categories: [papers, research, optimization, optimizer, youtube, nlp, vision]
# - image: images/adahessian.png
# I recently hosted a presentation with the authors of the [AdaHessian optimizer paper](https://arxiv.org/pdf/2006.00719.pdf), <NAME> and <NAME> and, to the fastai community.
#
# AdaHessian is a promising new optimizer and one of the first second-order optimizers that is becoming practical to use. I've been working on a port of it to fastai which I hope to release shortly, I;ll post up here once I have it released. Until then, I hope you enjoy the presentation and Q&A afterwards!
#
# ([**Click here**](https://www.youtube.com/watch?v=S87ancnZ0MM) of the embedded link below doesn't work)
#
# > youtube: https://www.youtube.com/watch?v=S87ancnZ0MM
#
# ## Useful links
#
# - [Paper link](https://arxiv.org/pdf/2006.00719.pdf)
# - [AdaHessian Paper repo](https://github.com/amirgholami/adahessian)
# - [My Fastai Implementation](https://github.com/morganmcg1/AdahessianFastai), basic working version, mixed precision coming
# - [Fastai discussion](https://forums.fast.ai/t/adahessian/76214/15)
# - [@lessw2020, Best-Deep-Learning-Optimizers Code](https://github.com/lessw2020/Best-Deep-Learning-Optimizers/tree/master/adahessian)
# - [@davda54 pytorch implementation](https://github.com/davda54/ada-hessian)
#
# ## Paper Citation
@article{yao2020adahessian,
title={ADAHESSIAN: An Adaptive Second Order Optimizer for Machine Learning},
author={<NAME> <NAME> <NAME> and Keutzer, Kurt and <NAME>},
journal={arXiv preprint arXiv:2006.00719},
year={2020}
}
# ## Thanks for Reading 😃
# As always, I would love to hear if you have any comments, thoughts or criticisms, you can find me on Twitter at [@mcgenergy](www.twitter.com/mcgenergy)
| _notebooks/2020-09-27-adahessian_with_the_authors.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: pyjax
# language: python
# name: pyjax
# ---
# +
import itertools
import math
from functools import partial
import numpy as onp
import jax
print("jax version: ", jax.__version__)
import jax.experimental.optimizers as optimizers
import jax.experimental.stax as stax
import jax.numpy as np
from jax import jit
import matplotlib.pyplot as plt
import IMNN
print("IMNN version: ", IMNN.__version__)
from IMNN.experimental.jax.imnn import (
AggregatedGradientIMNN,
AggregatedNumericalGradientIMNN,
AggregatedSimulatorIMNN,
GradientIMNN,
NumericalGradientIMNN,
SimulatorIMNN,
)
from IMNN.experimental.jax.lfi import (
ApproximateBayesianComputation,
GaussianApproximation,
)
from IMNN.experimental.jax.utils import value_and_jacrev, value_and_jacfwd
rng = jax.random.PRNGKey(0)
# -
np.less_equal(1, 0)
# +
# Define a log_normal field class
class LogNormalField:
@staticmethod
def compute_rsquared(nside):
"""
Compute the correlation function of the underlying gaussian field
Parameters:
nside : int
Image is nside x nside pixels
"""
import jax.numpy as np
from scipy.linalg import toeplitz
_Di = np.tile(toeplitz(np.arange(nside)),(nside,nside))
_Dj = np.concatenate(
[np.concatenate(
[np.tile(np.abs(i-j),(nside,nside)) for i in range(nside)],
axis=0)
for j in range(nside)],axis=1)
_distance_squared = _Di*_Di+_Dj*_Dj
return _distance_squared
# The lognormal correlation function where the gaussian field has a gaussian power spectrum,
# and the gaussian correlation function xi_G.
@staticmethod
def xi_G(rsq, beta):
"""
Calculates the two-point correlation function of a gaussian field with gaussian power spectrum
Parameters:
rsq : float
separation^2
beta : float
Gaussian smoothing width of gaussian field
"""
import jax.numpy as np
# ADD IN SIGMA PARAM HERE
xi = np.exp(-0.25*rsq/(beta**2))
return xi
@staticmethod
def fill_zeros(k, value):
from functools import partial
def fnk(k):
return jax.lax.cond(np.less_equal(k, 1e-5), lambda _: 0., lambda k: k+value, operand=k)
if len(k.shape) == 1:
return jax.vmap(fnk)(k)
else:
return jax.vmap(partial(fill_zeros, value=value))(k)
@staticmethod
def xi_LN(r, beta, alpha, PixelNoise):
"""
Calculates the lognormal two-point correlation function
Parameters:
r : float
Pair separation
beta : float
Gaussian smoothing width of underlying gaussian field
alpha : float
Nongaussianity parameter in lognormal transformation
PixelNoise : float
Standard deviation of added noise per pixel
"""
import jax.numpy as np
xi = 1/(np.power(alpha+1e-12,2)) * (np.exp(np.power(alpha,2)*np.exp(-0.25*np.power(r/beta,2))) - 1)
# Add pixel noise at zero separation:
xi = self.fill_zeros(xi, PixelNoise**2)
#xi[np.where(r<1e-5)] += PixelNoise**2
return xi
@staticmethod
def dxi_LN_dalpha(r, beta, alpha, PixelNoise):
import jax.numpy as np
return 2/(alpha+1e-12) * np.exp(-0.25*np.power(r/beta,2)) * np.exp(np.power(alpha,2)*np.exp(-0.25*np.power(r/beta,2))) - 2/np.power(alpha+1e-12,3) * (np.exp(np.power(alpha,2)*np.exp(-0.25*np.power(r/beta,2))) - 1)
@staticmethod
def dxi_LN_dbeta(r, beta, alpha, PixelNoise):
import jax.numpy as np
return (-0.5*r/np.power(beta,2)) * np.exp(-0.25*np.power(r/beta,2)) * np.exp(np.power(alpha,2)*np.exp(-0.25*np.power(r/beta,2)))
def __init__(self,Lside,rmax,nbin):
"""
Parameters:
rmax : float
Maximum pair separation considered
nbin : int
Number of bins for shell-averaged correlation function
"""
import jax.numpy as np
self.rmax = rmax
self.nbin = nbin
self.Lside = Lside
# compute the separations and indices on a grid
self.rsq = self.compute_rsquared(Lside)
self.r = np.sqrt(self.rsq)
self.bins = np.arange(nbin)*rmax/nbin
self.index = np.digitize(self.r,self.bins)
self.average_r = np.array([self.r[self.index == n].mean() for n in range(nbin) if np.sum(self.index == n)>0])
@staticmethod
def G_to_LN(gaussian, alpha):
import jax.numpy as np
# Make lognormal (variance of gaussian field is unity by construction)
# Divide by 1/alpha so that the signal-to-noise ratio is independent of alpha
return 1./alpha * (np.exp(alpha * gaussian-0.5*alpha**2)-1)
def run_simulation(self, key, alpha, beta, PixelNoise):
"""
Create a lognormal field from a gaussian field with a Gaussian correlation function
"""
import jax.numpy as np
# split keys, one for field and one for noise
key1,key2 = jax.random.split(key)
Lside = self.Lside
rsq = self.rsq
# Compute the Gaussian correlation function
xiG = self.xi_G(rsq,beta)
print('xiG shape: ', xiG.shape)
# Compute the Gaussian random field
#field = (jax.random.multivariate_normal(key1, np.zeros(Lside*Lside), xiG)).reshape(Lside,Lside)
print('field shape: ', field.shape)
# Make lognormal (variance of gaussian field is unity by construction)
field = self.G_to_LN(field, alpha)
# Add noise
field += jax.random.normal(key2, shape=(Lside,Lside))*np.sqrt(PixelNoise)
return field
def pymc3_model(self, field_data, alphamin, alphamax, betamin, betamax, PixelNoise):
import numpy as np
import pymc3 as pm
LN_model = pm.Model()
Lside = self.Lside
rsq = self.rsq
zero = np.zeros(Lside*Lside)
PixelNoiseVector = PixelNoise*np.ones(Lside*Lside)
InvNoiseCovariance = np.diag(1/(PixelNoiseVector**2))
field_data = field_data.reshape(Lside*Lside)
with LN_model:
# (TLM) TODO: add in μ,σ for full BHM
# Uniform priors for unknown model parameters (alpha,beta):
alpha_p = pm.Uniform("alpha", lower=alphamin, upper=alphamax)
beta_p = pm.Uniform("beta", lower=betamin, upper=betamax)
# Compute (beta-dependent) gaussian field correlation function:
xi = pm.math.exp(-0.25*rsq/(beta_p*beta_p))
# Gaussian field values are latent variables:
gaussian = pm.MvNormal("gaussian",mu=zero,cov=xi,shape=Lside*Lside)
# Expected value of lognormal field, for given (alpha, beta, gaussian):
muLN = 1/alpha_p * (pm.math.exp(alpha_p * gaussian-0.5*alpha_p*alpha_p)-1)
# Likelihood (sampling distribution) of observations, given the mean lognormal field:
Y_obs = pm.MvNormal("Y_obs", mu=muLN, tau=InvNoiseCovariance, observed=field_data)
return LN_model
def run_diff_simulation(self, alpha, beta, PixelNoise, step, seed):
"""
Run simulations for finite differencing
"""
import numpy as np
from scipy.stats import multivariate_normal
Lside = self.Lside
rsq = self.rsq
alphap = alpha*(1+step)
alpham = alpha*(1-step)
betap = beta*(1+step)
betam = beta*(1-step)
# Compute the gaussian correlation function
xiG = self.xi_G(rsq,beta)
xiG_betap = self.xi_G(rsq,betap)
xiG_betam = self.xi_G(rsq,betam)
# Compute Gaussian random fields with the same phases
Gfield = multivariate_normal(mean=np.zeros(Lside*Lside), cov=xiG).rvs(random_state=seed).reshape(Lside,Lside)
Gfield_betap = multivariate_normal(mean=np.zeros(Lside*Lside), cov=xiG_betap).rvs(random_state=seed).reshape(Lside,Lside)
Gfield_betam = multivariate_normal(mean=np.zeros(Lside*Lside), cov=xiG_betam).rvs(random_state=seed).reshape(Lside,Lside)
# Make lognormal (variance of gaussian field is unity by construction)
field = self.G_to_LN(Gfield, alpha)
field_betap = self.G_to_LN(Gfield_betap, alpha)
field_betam = self.G_to_LN(Gfield_betam, alpha)
field_alphap = self.G_to_LN(Gfield, alphap)
field_alpham = self.G_to_LN(Gfield, alpham)
# Add noise
noise = np.random.normal(loc=0.0,scale=PixelNoise,size=(Lside,Lside))
field += noise
field_betap += noise
field_betam += noise
field_alphap += noise
field_alpham += noise
return field, field_alphap, field_alpham, field_betap, field_betam
def compute_corrfn(self,field):
"""
Compute two-point correlation function
"""
import numpy as np
index = self.index
nbin = self.nbin
# compute the correlations
correlations = np.outer(field,field)
corrfn = np.array([correlations[index==n].mean() for n in range(nbin) if len(correlations[index==n])>0])
return corrfn
def compute_corrfn_derivatives(self, field, field_alphap, field_alpham, field_betap, field_betam, step):
"""
Compute derivatives of the two-point correlation function
"""
# Compute correlation functions
corrfn = self.compute_corrfn(field)
corrfn_dalphap = self.compute_corrfn(field_alphap)
corrfn_dalpham = self.compute_corrfn(field_alpham)
corrfn_dbetap = self.compute_corrfn(field_betap)
corrfn_dbetam = self.compute_corrfn(field_betam)
# Compute derivatives by second-order central finite differences
dcorrfn_dalpha = (corrfn_dalpham - 2*corrfn + corrfn_dalphap)/(step**2)
dcorrfn_dbeta = (corrfn_dbetam - 2*corrfn + corrfn_dbetap )/(step**2)
return dcorrfn_dalpha, dcorrfn_dbeta
def covariance(self,fields):
"""
Compute covariance from a number of fields
Parameter:
fields : int
lognormal field objects contributing to the covariance matrix
"""
import numpy as np
nsims = len(fields)
nbins = self.nonzerobins
print('Number of simulations',nsims)
print('Number of non-zero pair bins',nbins)
corrfns = np.array([fields[i]['corrfn'] for i in range(nsims)])
meanxi = np.mean(corrfns,axis=0)
covxi = np.cov(corrfns.T)
return meanxi, covxi
# Utility properties
@staticmethod
def var_th(alpha, PixelNoise):
import numpy as np
return 1/np.power(alpha+1e-12,2)*(np.exp(alpha**2)-1)+PixelNoise**2
@staticmethod
def skew_th(alpha):
import numpy as np
return (np.exp(alpha**2)+2)*np.sqrt(np.exp(alpha**2)-1)
@staticmethod
def dskew_dalpha(alpha):
import numpy as np
return 2*alpha*np.exp(alpha**2) * ( np.sqrt(np.exp(alpha**2)-1) - 0.5*(np.exp(alpha**2)+2)/(np.sqrt(np.exp(alpha**2)-1)) )
@staticmethod
def kurtosis_th(alpha):
import numpy as np
return np.exp(4*alpha**2)+2*np.exp(3*alpha**2)+3*np.exp(2*alpha**2)-6
@staticmethod
def dkurtosis_dalpha(alpha):
import numpy as np
return 8*alpha*np.exp(4*alpha**2)+6*alpha*np.exp(3*alpha**2)+6*alpha*np.exp(2*alpha**2)
@staticmethod
def max(field):
import numpy as np
return np.max(field)
@staticmethod
def min(field):
import numpy as np
return np.min(field)
@staticmethod
def var(field):
import numpy as np
return np.var(field)
@staticmethod
def mean(field):
import numpy as np
return np.mean(field)
@staticmethod
def skew(field):
from scipy.stats import skew
return skew(field.flatten())
@staticmethod
def kurtosis(field):
from scipy.stats import kurtosis
return kurtosis(field.flatten())
# xi has empty bins removed. Note the number of non-empty elements
@property
def nonzerobins(self):
return len(self.average_r)
@property
def dt(self):
import numpy as np
return np.dtype([('field', np.float, (self.Lside,self.Lside)), ('corrfn', np.float, (self.nonzerobins))])
# end class LogNormalField
# +
Lside = 16
alpha = 1.0
beta = 0.5
PixelNoise = 0.01
# Setup for correlation function
nbin = 4*Lside
ndata = 4*Lside
rmax = Lside*np.sqrt(2)
# -
LN=LogNormalField(Lside,rmax,nbin)
rng,key = jax.random.split(rng)
field = LN.run_simulation(key, 1.0, 0.5, 0.01)
LN.rsq.shape
plt.imshow(field)
plt.colorbar()
LN.average_r.shape
simulator_args = {'N': 32, 'squeeze': False}
def simulator(rng, θ, simulator_args=simulator_args):
A,B = θ
noise = 0.01
def fn(key, A,B):
if simulator_args['squeeze']:
return np.expand_dims(LN.run_simulation(key,A,B,noise), 0)
else:
return (np.expand_dims(np.expand_dims(LN.run_simulation(key,A,B,noise), 0), 0))
if A.shape == B.shape:
if len(A.shape) == 0:
return fn(rng, A, B)
else:
keys = jax.random.split(rng, num=A.shape[0] + 1)
rng = keys[0]
keys = keys[1:]
return jax.vmap(
lambda key, A, B: simulator(key, (A, B), simulator_args=simulator_args)
)(keys, A, B)
else:
if len(A.shape) > 0:
keys = jax.random.split(rng, num=A.shape[0] + 1)
rng = keys[0]
keys = keys[1:]
return jax.vmap(
lambda key, A: simulator(key, (A, B), simulator_args=simulator_args)
)(keys, A)
elif len(B.shape) > 0:
keys = jax.random.split(rng, num=B.shape[0])
return jax.vmap(
lambda key, B: simulator(key, (A, B), simulator_args=simulator_args)
)(keys, B)
def simulator_gradient(rng, θ, simulator_args=simulator_args):
return value_and_jacrev(simulator, argnums=1, allow_int=True, holomorphic=True)(rng, θ, simulator_args=simulator_args)
simulation.shape
# +
rng,key = jax.random.split(rng)
θ_fid = np.array([1.0, 0.5], dtype=np.float32)
simulation, simulation_gradient = value_and_jacfwd(simulator, argnums=1)(rng, θ_fid, simulator_args=simulator_args)
plt.imshow(np.squeeze(simulation[0]), extent=(0,1,0,1))
plt.colorbar()
plt.title('example simulation')
plt.show()
plt.imshow(np.squeeze(simulation_gradient[0].T[0].T), extent=(0,1,0,1))
plt.title('gradient of simulation')
plt.colorbar()
plt.show()
# -
# # set up model
# +
n_params = 2
n_summaries = 2
θ_fid = np.array([1.0, 0.5], dtype=np.float32)
N = Lside
input_shape = (1,1, N,N)
# IMNN params
n_s = 5000
n_d = 5000
λ = 100.0
ϵ = 0.1
# -
LN.rsq.shape
rng, initial_model_key = jax.random.split(rng)
rng, fitting_key = jax.random.split(rng)
def InceptBlock2(filters, strides, do_5x5=True, do_3x3=True):
"""InceptNet convolutional striding block.
filters: tuple: (f1,f2,f3)
filters1: for conv1x1
filters2: for conv1x1,conv3x3
filters3L for conv1x1,conv5x5"""
filters1, filters2, filters3 = filters
conv1x1 = stax.serial(stax.Conv(filters1, (1,1), strides, padding="SAME"))
filters4 = filters2
conv3x3 = stax.serial(stax.Conv(filters2, (1,1), strides=None, padding="SAME"),
stax.Conv(filters4, (3,3), strides, padding="SAME"))
filters5 = filters3
conv5x5 = stax.serial(stax.Conv(filters3, (1,1), strides=None, padding="SAME"),
stax.Conv(filters5, (5,5), strides, padding="SAME"))
maxpool = stax.serial(stax.MaxPool((3,3), padding="SAME"),
stax.Conv(filters4, (1,1), strides, padding="SAME"))
if do_3x3:
if do_5x5:
return stax.serial(
stax.FanOut(4), # should num=3 or 2 here ?
stax.parallel(conv1x1, conv3x3, conv5x5, maxpool),
stax.FanInConcat(),
stax.LeakyRelu)
else:
return stax.serial(
stax.FanOut(3), # should num=3 or 2 here ?
stax.parallel(conv1x1, conv3x3, maxpool),
stax.FanInConcat(),
stax.LeakyRelu)
else:
return stax.serial(
stax.FanOut(2), # should num=3 or 2 here ?
stax.parallel(conv1x1, maxpool),
stax.FanInConcat(),
stax.LeakyRelu)
fs = 32
model = stax.serial(
#InceptBlock2((fs,fs,fs), strides=(4,4)),
#InceptBlock2((fs,fs,fs), strides=(4,4)),
InceptBlock2((fs,fs,fs), strides=(4,4)),
InceptBlock2((fs,fs,fs), strides=(2,2), do_5x5=False, do_3x3=False),
InceptBlock2((fs,fs,fs), strides=(2,2), do_5x5=False, do_3x3=False),
stax.Conv(n_summaries, (1,1), strides=(1,1), padding="SAME"),
stax.Flatten
)
optimiser = optimizers.adam(step_size=1e-3)
# +
from IMNN.experimental.jax.imnn._imnn import _IMNN
from IMNN.experimental.jax.utils import check_simulator, value_and_jacrev
class SimIMNN(_IMNN):
def __init__(self, n_s, n_d, n_params, n_summaries, input_shape, θ_fid,
model, optimiser, key_or_state, simulator, verbose=True):
super().__init__(
n_s=n_s,
n_d=n_d,
n_params=n_params,
n_summaries=n_summaries,
input_shape=input_shape,
θ_fid=θ_fid,
model=model,
key_or_state=key_or_state,
optimiser=optimiser,
verbose=verbose)
self.simulator = check_simulator(simulator)
self.simulate = True
def get_fitting_keys(self, rng):
return jax.random.split(rng, num=3)
def get_summaries(self, w, key, validate=False):
def get_summary(key, θ):
return self.model(w, self.simulator(key, θ))
def get_derivatives(key):
return value_and_jacrev(get_summary, argnums=1)(key, self.θ_fid)
keys = np.array(jax.random.split(key, num=self.n_s))
summaries, derivatives = jax.vmap(get_derivatives)(keys[:self.n_d])
if self.n_s > self.n_d:
summaries = np.vstack([
summaries,
jax.vmap(partial(get_summary, θ=self.θ_fid))(keys[self.n_d:])])
return np.squeeze(summaries), np.squeeze(derivatives)
import jax
import jax.numpy as np
from IMNN.experimental.jax.imnn import SimulatorIMNN
from IMNN.experimental.jax.utils import value_and_jacrev, check_devices, \
check_type, check_splitting
class AggregatedSimulatorIMNN(SimulatorIMNN):
def __init__(self, n_s, n_d, n_params, n_summaries, input_shape, θ_fid,
model, optimiser, key_or_state, simulator, devices,
n_per_device, verbose=True):
super().__init__(
n_s=n_s,
n_d=n_d,
n_params=n_params,
n_summaries=n_summaries,
input_shape=input_shape,
θ_fid=θ_fid,
model=model,
key_or_state=key_or_state,
optimiser=optimiser,
simulator=simulator,
verbose=verbose)
self.devices = check_devices(devices)
self.n_devices = len(self.devices)
self.n_per_device = check_type(n_per_device, int, "n_per_device")
if self.n_s == self.n_d:
check_splitting(self.n_s, "n_s and n_d", self.n_devices,
self.n_per_device)
else:
check_splitting(self.n_s, "n_s", self.n_devices, self.n_per_device)
check_splitting(self.n_d, "n_d", self.n_devices, self.n_per_device)
def get_summaries(self, w, key=None, validate=False):
def derivative_scan(counter, rng):
def get_device_summaries(rng):
def get_summary(key, θ):
return self.model(w, self.simulator(key, θ))
def get_derivatives(rng):
return value_and_jacrev(get_summary, argnums=1)(
rng, self.θ_fid)
keys = np.array(jax.random.split(rng, num=self.n_per_device))
return jax.vmap(get_derivatives)(keys)
keys = np.array(jax.random.split(rng, num=self.n_devices))
summaries, derivatives = jax.pmap(
get_device_summaries, devices=self.devices)(keys)
return counter, (summaries, derivatives)
def summary_scan(counter, rng):
def get_device_summaries(rng):
def get_summary(key):
return self.model(w, self.simulator(key, self.θ_fid))
keys = np.array(jax.random.split(rng, num=self.n_per_device))
return jax.vmap(get_summary)(keys)
keys = np.array(jax.random.split(rng, num=self.n_devices))
summaries = jax.pmap(
get_device_summaries, devices=self.devices)(keys)
return counter, summaries
n = self.n_d // (self.n_devices * self.n_per_device)
if self.n_s > self.n_d:
n_r = (self.n_s - self.n_d) // (self.n_devices * self.n_per_device)
key, *keys = jax.random.split(key, num=n_r + 1)
counter, remaining_summaries = jax.lax.scan(
summary_scan, n_r, np.array(keys))
keys = np.array(jax.random.split(key, num=n))
counter, results = jax.lax.scan(
derivative_scan, 0, keys)
summaries, derivatives = results
if self.n_s > self.n_d:
summaries = np.vstack([summaries, remaining_summaries])
return (summaries.reshape((-1, self.n_summaries)),
derivatives.reshape((-1, self.n_summaries, self.n_params)))
# -
IMNN = SimIMNN(
n_s=5000,
n_d=5000,
n_params=n_params,
n_summaries=n_summaries,
input_shape=input_shape,
θ_fid=θ_fid,
model=model,
optimiser=optimiser,
key_or_state=initial_model_key,
simulator=lambda rng, θ: simulator(rng, θ, simulator_args=simulator_args),
# devices=[jax.devices()[0]],
# n_per_device=1000
)
IMNN_rngs = 1 * [fitting_key] #+ 12 * [None]
labels = [
"Simulator, InceptNet\n"
]
# %%time
for i in range(1):
rng,fit_rng = jax.random.split(rng)
IMNN.fit(λ=10., ϵ=ϵ, rng=fit_rng, min_iterations=500) #for IMNN, IMNN_rng in zip(IMNNs, IMNN_rngs);
IMNNs = [IMNN]
for i, (IMNN, label) in enumerate(zip(IMNNs, labels)):
if i == 0:
ax = IMNN.training_plot(expected_detF=None, colour="C{}".format(i), label=label)
elif i == 10:
other_ax = IMNN.training_plot(
expected_detF=None, colour="C{}".format(i), label=label
)
elif i == 11:
IMNN.training_plot(
ax=other_ax,
expected_detF=50, colour="C{}".format(i), label=label
)
other_ax[0].set_yscale("log")
other_ax[2].set_yscale("log")
else:
IMNN.training_plot(
ax=ax, expected_detF=None, colour="C{}".format(i), label=label, ncol=5
);
ax[0].set_yscale("log")
IMNN.F
# # ABC inference
# +
class uniform:
def __init__(self, low, high):
self.low = np.array(low)
self.high = np.array(high)
self.event_shape = [[] for i in range(self.low.shape[0])]
def sample(self, n=None, seed=None):
if n is None:
n = 1
keys = np.array(jax.random.split(
seed,
num=len(self.event_shape)))
return jax.vmap(
lambda key, low, high : jax.random.uniform(
key,
shape=(n,),
minval=low,
maxval=high))(
keys, self.low, self.high)
prior = uniform([0.1, 0.1], [1.6, 1.6])
#prior = uniform([0.1, 0.1], [5.0, 3.0])
# -
rng, key = jax.random.split(rng)
θ_target = np.array([0.9, 0.6])
target_data = simulator(
key,
θ_target,
simulator_args={**simulator_args, **{'squeeze':False}})
@jit #partial(jax.jit, static_argnums=0)
def get_estimate(d):
if len(d.shape) == 1:
return IMNN.θ_fid + np.einsum(
"ij,kj,kl,l->i",
IMNN.invF,
IMNN.dμ_dθ,
IMNN.invC,
IMNN.model(IMNN.best_w, d, rng=rng) - IMNN.μ)
else:
return IMNN.θ_fid + np.einsum(
"ij,kj,kl,ml->mi",
IMNN.invF,
IMNN.dμ_dθ,
IMNN.invC,
IMNN.model(IMNN.best_w, d, rng=rng) - IMNN.μ)
estimates = get_estimate(target_data) #[i.get_estimate(target_data) for i in IMNNs];
GAs = [GaussianApproximation(get_estimate(target_data), IMNN.invF, prior)]
# %matplotlib inline
for i, (GA, label) in enumerate(zip(GAs, labels)):
if i == 0:
ax = GA.marginal_plot(
axis_labels=[r"$\alpha$", r"$\beta$"], label='on-the-fly IMNN', colours="C{}".format(i)
)
else:
GA.marginal_plot(ax=ax, label=label, colours="C{}".format(i), ncol=8)
ABC = ApproximateBayesianComputation(
target_data, prior,
lambda A,B : simulator(A,B, simulator_args={**simulator_args, **{'squeeze':True}}),
get_estimate, F=IMNN.F, gridsize=50
)
target_data.shape
# %%time
rng,abc_key = jax.random.split(rng)
ABC(rng=abc_key,
n_samples=int(1e3),
min_accepted=15000,
max_iterations=20000,
ϵ=0.05,
smoothing=0.);
ABC.parameters.accepted[0].shape
α
# +
# %matplotlib inline
#plt.style.use('default')
new_colors = [ '#2c0342', '#286d87', '#4fb49d', '#9af486']
fig,ax = plt.subplots(nrows=2, ncols=2, figsize=(3.37*2, 3.37*2))
#latexify(fig_width=3.37, fig_height=3.37)
ABC.scatter_plot(ax=ax,
colours=new_colors[0],
axis_labels=[r"$\alpha$", r"$\beta$"],
s=8,
label='ABC estimate')
# ABC.marginal_plot(ax=ax,
# axis_labels=[r"$A$", r"$B$"], colours='green',
# label='ABC marginal plot')
GAs[0].marginal_plot(ax=ax, colours=new_colors[2], axis_labels=[r"$\alpha$", r"$\beta$"], label=None, ncol=1)
ax[0,1].imshow(target_data[0, 0])
ax[0,1].set_yticks([])
#ax[0,1].set_title(r'$\theta_{\rm target} = A,B = (%.2f,%.2f)$'%(θ_target[0], θ_target[1]))
ax[0,0].axvline(θ_target[0], linestyle='--', c='k')
ax[1,0].axvline(θ_target[0], linestyle='--', c='k')
ax[1,0].axhline(θ_target[1], linestyle='--', c='k')
ax[1,1].axhline(θ_target[1], linestyle='--', c='k', label=r'$\theta_{\rm target}$')
ax[1,0].set_xlabel(r'$\alpha$')
ax[1,0].set_ylabel(r'$\beta$')
ax[0,0].axvline(θ_fid[0], linestyle='--', c='k', alpha=0.4)
ax[0,0].set_yticks([])
#ax[1,0].contourf(A_range, B_range, L1.reshape((size, size)))
#ax[0, 0].plot(A_range, np.real(loglikeA), color='g', label='loglikeA')
ax[1,0].axvline(θ_fid[0], linestyle='--', c='k', alpha=0.4)
ax[1,0].axhline(θ_fid[1], linestyle='--', c='k', alpha=0.4)
ax[1,1].axhline(θ_fid[1], linestyle='--', c='k', alpha=0.4, label=r'$\theta_{\rm fid}$')
ax[1,1].set_yticks([])
#ax[1,1].plot(np.real(loglikeB), B_range, color='g', label='loglikeB')
ax[1,1].legend(framealpha=0.)
# add in the likelihood estimate
# ax[0, 0].plot(A_range, likelihoodA, color='#FF8D33', label=None)
# ax[0, 1].axis("off")
# ax[1, 0].contour(A_range, B_range, np.real(likelihood), levels=value, colors='#FF8D33')
# ax[1, 1].plot(likelihoodB, B_range, color='#FF8D33', label='loglike')
ax[0,0].legend(framealpha=0.)
#plt.savefig('/mnt/home/tmakinen/repositories/field-plots/128x128-contours.png', dpi=400)
plt.subplots_adjust(wspace=0, hspace=0)
plt.show()
# -
import powerbox as pbox
# +
α = 1.0
β = 0.5
#mf = hmf.MassFunction(z=0)
# Generate a callable function that returns the cosmological power spectrum.
power = lambda k : ((1 / α**2) * np.exp(α**2 * np.exp(-0.25 * (k**2 / β**2)) - 1))
# Create the power-box instance. The boxlength is in inverse units of the k of which pk is a function, i.e.
# Mpc/h in this case.
pb = pbox.LogNormalPowerBox(N=128, dim=2, pk = power, boxlength= 128., seed=123)
# -
plt.imshow(pb.delta_x(), extent=(0,100,0,100))
plt.colorbar()
plt.show()
| fields/.ipynb_checkpoints/florent-LN-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import dask.async
import dask.array as da
import sys
sys.path.insert(0, '../..')
# %load_ext autoreload
# %autoreload 1
# %aimport allel
# %aimport allel.model
# %aimport allel.model.dask
allel.__version__
# ## Example 1
g = allel.GenotypeArray([[[0, 0], [0, 1], [0, 1]],
[[0, 1], [1, 1], [1, 1]],
[[1, 1], [1, 1], [0, 0]],
[[1, 1], [1, 1], [0, 0]],
[[1, 1], [0, 1], [-1, -1]]], dtype='i1')
g
# the mapping array is effectively a look-up table, telling how to transform
# integer values in each row
mapping = np.array([[0, 1], # no transformation
[1, 0], # 0->1, 1->0
[0, 0], # 0->0, 1->0
[1, 0], # 0->1, 1->0
[0, 1]], # no transformation
dtype='i1')
# the only shape constraint is that size of first dimension must match
assert g.shape[0] == mapping.shape[0]
# this is the pure numpy implementation
expect = g.map_alleles(mapping)
expect
chunks_dim0 = 2
gd = da.from_array(g, chunks=(chunks_dim0, 2, None))
gd
md = da.from_array(mapping, chunks=(chunks_dim0, None)) # N.B., first dimension chunk size matches gd1
md
def f(block, bmapping):
return allel.GenotypeArray(block).map_alleles(bmapping[:, 0, :])
gmapped1 = da.map_blocks(f, gd, md[:, None, :], chunks=gd.chunks)
gmapped1
actual = allel.GenotypeArray(gmapped1.compute())
actual
assert np.array_equal(expect, actual)
# ## Example 2
g = allel.GenotypeArray([
[[0, 0], [0, 1], [0, 0], [0, 1], [-1, -1]],
[[0, 2], [1, 1], [0, 2], [1, 1], [-1, -1]],
[[0, 0], [0, 1], [0, 0], [0, 1], [-1, -1]],
[[0, 2], [1, 1], [0, 2], [1, 1], [-1, -1]],
[[1, 0], [2, 1], [1, 0], [2, 1], [-1, -1]],
[[2, 2], [-1, -1], [2, 2], [-1, -1], [-1, -1]],
[[-1, -1], [-1, -1], [-1, -1], [-1, -1], [-1, -1]]
], dtype='i1')
g
mapping = np.array([[0, 1, 2],
[2, 0, 1],
[1, 2, 0],
[2, 1, 0],
[1, 2, 0],
[2, 1, 0],
[2, 0, 1]], dtype=np.int8)
expect = g.map_alleles(mapping)
expect
gd = da.from_array(g, chunks=(chunks_dim0, 2, None))
gd
md = da.from_array(mapping, chunks=(chunks_dim0, None))
md
def f(block, bmapping):
return allel.GenotypeArray(block).map_alleles(bmapping[:, 0, :])
res = da.map_blocks(f, gd, md[:, None, :], chunks=gd.chunks)
res
actual = allel.GenotypeArray(res.compute())
actual
assert np.array_equal(expect, actual)
# ### Alternative solution
def ff(block, bmapping):
return allel.GenotypeArray(block[:, :, :, 0]).map_alleles(bmapping[:, 0, 0, :])
res2 = da.map_blocks(ff, gd[:, :, :, None], md[:, None, None, :], chunks=gd.chunks, drop_dims=3)
res2
actual2 = res2.compute()
actual2.shape
assert np.array_equal(expect, actual2)
# ## Example 3
g = allel.GenotypeArray([
[[0, 0], [0, 1], [-1, -1]],
[[0, 2], [1, 1], [-1, -1]],
[[1, 0], [2, 1], [-1, -1]],
[[2, 2], [-1, -1], [-1, -1]],
[[-1, -1], [-1, -1], [-1, -1]]
], dtype='i1')
g
mapping = np.array([[0, 1, 2],
[2, 0, 1],
[1, 2, 0],
[2, 1, 0],
[2, 0, 1]], dtype=np.int8)
expect = [[[0, 0], [0, 1], [-1, -1]],
[[2, 1], [0, 0], [-1, -1]],
[[2, 1], [0, 2], [-1, -1]],
[[0, 0], [-1, -1], [-1, -1]],
[[-1, -1], [-1, -1], [-1, -1]]]
assert np.array_equal(expect, g.map_alleles(mapping))
gd = da.from_array(g, chunks=(2, 2, None))
gd
md = da.from_array(mapping, chunks=(2, None))
md
res = da.map_blocks(f, gd, md[:, None, :], chunks=gd.chunks)
res
actual = allel.GenotypeArray(res.compute())
actual
assert np.array_equal(expect, actual)
| notebooks/sandbox/dask_map_alleles.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Efficiency Notebooks: Exploring the emissions density of power used by homes
# We'll be using data from the Texas ISO, ERCOT, to calculate the emissions amount and emission sources from the generated power used by the electrical grid home usage for 50 homes.
#
# Entirely solar equipped homes will be used to demonstrate the interplay between solar generation and emissions during this time period (March - August 2018)
#
# ERCOT emissions/generation data pulled from http://www.ercot.com/content/wcm/lists/181766/FuelMixReport_PreviousYears.zip
# which you can find at http://www.ercot.com/gridinfo/generation
#import packages
import pandas as pd
import matplotlib.pyplot as plt
import psycopg2
import sqlalchemy as sqla
import os
import sys
sys.path.insert(0,'..')
from config.read_config import get_database_config
import numpy as np
import statistics
# %matplotlib inline
sys.executable # shows you your path to the python you're using
# set constants for lbs of CO2 / kWh
_gas_cc_lbs = 0.75
_gas_lbs = 1.0
_coal_lbs = 2.21
# +
# read in db credentials from ../config/config.txt
# * make sure you add those to the ../config/config.txt file! *
## Uncomment the following line to use the live database queries
database_config = get_database_config("../config/config.txt")
#
# +
# get our DB connection
# uncomment if you want to use the live queries to the database instead of the prepared data
engine = sqla.create_engine('postgresql://{}:{}@{}:{}/{}'.format(database_config['username'],
database_config['password'],
database_config['hostname'],
database_config['port'],
database_config['database']
))
# +
#Select a list of Austin homes from dataport metadata with good data availability for grid
query = """select distinct dataid, egauge_1min_data_availability, grid, solar from other_datasets.metadata
where grid = 'yes'
and solar = 'yes'
and egauge_1min_min_time <= '2018-03-01'
and egauge_1min_max_time > '2018-09-01'
and city='Austin'
and (egauge_1min_data_availability like '100%'
or
egauge_1min_data_availability like '99%'
or
egauge_1min_data_availability like '98%'
or
egauge_1min_data_availability like '97%'
)
and gas_ert_min_time <= '2018-03-01'
and gas_ert_max_time > '2018-09-01'
limit 50
;
"""
df = pd.read_sql_query(sqla.text(query), engine)
df
# -
# grab dataids and convert them to a string to put into the SQL query
dataids_list = df['dataid'].tolist()
print("{} dataids selected listed here:".format(len(dataids_list)))
dataids_str = ','.join(list(map(str, dataids_list)))
dataids_str
# +
#Pull electricity data for selected homes.
data = """select dataid,localminute::timestamp,grid
from electricity.eg_realpower_1min
where localminute >= '2018-03-01' and localminute < '2018-09-01' """
data = data + """AND dataid in ({})""".format(dataids_str)
# create a dataframe with the data from the sql query
grid_df = pd.read_sql_query(sqla.text(data), engine)
grid_df
# +
# read in 2018 ERCOT emissions data
ercot = pd.read_csv('ercot_emissions_2018.csv')
# index by Energy, GWh
ercot = ercot.set_index('Energy, GWh')
# remove the commas from the numbers
ercot.replace(',','', regex=True, inplace=True)
# convert to a float from a string
ercot = ercot.astype('float64')
ercot
# -
# Calc just one of them months and sources for a sanity check
perc_coal_mar = ercot.loc['Coal','Mar'] / ercot.loc['Total','Mar']
perc_coal_mar
# find the percentages for coal, gas, and gas-cc of the total blend of generation sources from ERCOT for our months and the emissions-producing sources
sources = ['Coal', 'Gas', 'Gas-CC']
months = ['Mar', 'Apr', 'May', 'Jun', 'Jul', 'Aug']
percs = {}
for source in sources:
for month in months:
percs[source + '' + month] = ercot.loc[source, month] / ercot.loc['Total', month]
percs
# +
# take the mean across the months for each source
coal_ave = statistics.mean([percs['CoalMar'], percs['CoalApr'], percs['CoalMay'], percs['CoalJun'], percs['CoalJul'], percs['CoalAug']])
gas_ave = statistics.mean([percs['GasMar'], percs['GasApr'], percs['GasMay'], percs['GasJun'], percs['GasJul'], percs['GasAug']])
gascc_ave = statistics.mean([percs['Gas-CCMar'], percs['Gas-CCApr'], percs['Gas-CCMay'], percs['Gas-CCJun'], percs['Gas-CCJul'], percs['Gas-CCAug']])
print ('Coal = {}%'.format(coal_ave * 100))
print ('Gas = {}%'.format(gas_ave * 100))
print ('Gas-CC = {}%'.format(gascc_ave * 100))
# +
# complete the full percentage, fill with the rest of the sources that are largely non-emissions producing
the_rest = 1.0 - coal_ave - gas_ave - gascc_ave
# pie chart
pie_data = [coal_ave, gas_ave, gascc_ave, the_rest]
pie_labels = ['Coal', 'Gas', 'Gas-CC', 'Other']
explode = [.05, .05, .05, .05] # separates the slices a little bit
plt.pie(pie_data, labels=pie_labels, autopct='%1.1f%%', startangle=15, shadow = True, explode=explode)
plt.title('ERCOT Generation Percentages')
plt.axis('equal')
plt.show()
# +
# convert ercot table to percentages:
def add_percentages(column):
return column / column['Total']
ercot_perc = ercot.apply(add_percentages)
ercot_perc
# -
ercot_perc.index.name = "% of Generation"
ercot_perc
# clean up that percentage table
ercot_perc = ercot_perc.drop(index=['Biomass', 'Hydro', 'Nuclear', 'Other', 'Solar', 'Wind', 'Total'], columns=['Jan', 'Feb', 'Sep', 'Oct', 'Nov', 'Dec', 'Total'])
ercot_perc
# +
# index by localminute
grid_df = grid_df.set_index('localminute')
# bring to central timezone
grid_df = grid_df.tz_localize(tz='US/Central')
grid_df
# -
# drop any rows that have blank grid
grid_df = grid_df.dropna(how='any')
grid_df
# calculate the average grid usage of the homes over this time period
grouped_grid = grid_df.groupby([pd.Grouper(freq='D')]).mean()
grouped_grid['grid'] = grouped_grid['grid'] * 24 # converts daily average grid use/generation to kWh
grouped_grid
# the above was using the monthly averages from Mar - Aug from ercot all averaged together for each source
# let's use the actual monthy averages for each point instead
grid_more = grouped_grid
# extract and addd the month to the dataframe
grid_more['Month'] = grid_more.index.strftime('%B')
grid_more['Month'] = grid_more['Month'].astype(str)
grid_more
# +
# convert the month to the same 3 letter abbreviation as in the ERCOT table
def shorten_month(col):
col['Month'] = col['Month'][0:3]
return col
grid_more = grid_more.apply(shorten_month, axis=1)
grid_more
# +
# Assign based on the monthly percentage breakdown
def assign_lbs(row):
row['Gas-CC lbs'] = (ercot_perc.loc['Gas-CC', row.Month] * row['grid']) * _gas_cc_lbs
row['Gas lbs'] = (ercot_perc.loc['Gas', row.Month] * row['grid']) * _gas_lbs
row['Coal lbs'] = (ercot_perc.loc['Coal', row.Month] * row['grid']) * _coal_lbs
return row
grid_more = grid_more.apply(assign_lbs, axis=1)
grid_more
# -
# don't need these anymore
grid_more = grid_more.drop(columns=['dataid', 'Month'])
grid_more
# Add a total CO2 column
grid_more['Total CO2'] = grid_more['Gas-CC lbs'] + grid_more['Gas lbs'] + grid_more['Coal lbs']
grid_more
grid_more = grid_more.rename({'grid':'Grid Use (kWh)'} , axis='columns')
grid_more.plot(figsize=(25,15), title='Daily Grid (kWh) and Emissions in lbs of CO2', grid=True, xlabel='Day', ylabel='kWh or lbs CO2')
# # Observations:
#
# - These all solar homes have the capacity to offset some of the neighbors' emissions in the "shoulder months" by putting energy back on the grid
# - Total CO2 as measured in lbs/kWh tracks at nearly 1-to-1
#
| Efficiency/Efficiency-Emissions-Density.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # Build a product recommendation engine
#
# 
#
# This notebook contains steps and code to create a recommendation engine based on shopping history, deploy that model to Watson Machine Learning, and create an app with PixieApps. This notebook runs on Python 3.x with Apache Spark 2.1.
#
# ## Learning Goals
#
# The learning goals of this notebook are:
#
# * Load a CSV file into the Object Storage service linked to your Watson Studio
# * Use the *k*-means algorithm, which is useful for cluster analysis in data mining, to segment customers into clusters for the purpose of making an in-store purchase recommendation
# * Deploy the model to the IBM Watson Machine Learning service in IBM Cloud to create your recommendation application
# ## Table of contents
#
# 1. [Setup](#setup)<br>
# 2. [Load and explore data](#load)<br>
# 3. [Create a KMeans model](#kmeans)<br>
# 3.1. [Prepare data](#prepare_data)<br>
# 3.2. [Create clusters and define the model](#build_model)<br>
# 4. [Persist the model](#persist)<br>
# 5. [Deploy the model to the cloud](#deploy)<br>
# 5.1. [Create deployment for the model](#create_deploy)<br>
# 5.2. [Test model deployment](#test_deploy)<br>
# 6. [Create product recommendations](#create_recomm)<br>
# 6.1. [Test product recommendations model](#test_recomm)<br>
# 7. [Summary and next steps](#summary)<br>
# ## 1. Setup
#
#
# Before you use the sample code in this notebook, you must perform the following setup tasks:
#
# * Create a Watson Machine Learning service instance (a free plan is offered) and associate it with your project
#
# We'll be using a few libraries for this exercise:
#
# 1. [Watson Machine Learning Client](http://wml-api-pyclient.mybluemix.net/): Client library to work with the Watson Machine Learning service on IBM Cloud. Library available on [pypi](https://pypi.org/project/watson-machine-learning-client/). Service available on [IBM Cloud](https://cloud.ibm.com/catalog/services/machine-learning).
# 1. [Pixiedust](https://github.com/pixiedust/pixiedust): Python Helper library for Jupyter Notebooks. Available on [pypi](https://pypi.org/project/pixiedust/).
# 1. [ibmos2spark](https://github.com/ibm-watson-data-lab/ibmos2spark): Facilitates Data I/O between Spark and IBM Object Storage services
# !pip install --upgrade ibmos2spark
# !pip install --upgrade pixiedust
# !pip install --upgrade watson-machine-learning-client
import pixiedust
# <a id="load"></a>
# ## 2. Load and explore data
#
# In this section you will load the data file that contains the customer shopping data using PixieDust's [`sampleData`](https://pixiedust.github.io/pixiedust/loaddata.html) method:
df = pixiedust.sampleData('https://raw.githubusercontent.com/IBM/product-recommendation-with-watson-ml/master/data/customers_orders1_opt.csv')
# To better examine and visualize the data, run the following cell to view it in a table format. Note that Pixiedust's `display` method can also render data using various chart types, such as pie charts, line graphs, and scatter plots.
# + pixiedust={"displayParams": {"handlerId": "tableView"}}
display(df)
# -
# <a id="kmeans"></a>
# ## 3. Create a *k*-means model with Spark
#
# In this section of the notebook you use the *k*-means implementation to associate every customer to a cluster based on their shopping history.
#
# First, import the Apache Spark Machine Learning packages ([MLlib](http://spark.apache.org/docs/2.2.0/api/python/pyspark.ml.html)) that you need in the subsequent steps:
#
from pyspark.ml import Pipeline
from pyspark.ml.clustering import KMeans
from pyspark.ml.clustering import KMeansModel
from pyspark.ml.feature import VectorAssembler
from pyspark.ml.linalg import Vectors
# <a id="prepare_data"></a>
# ### 3.1 Prepare data
#
# Create a new data set with just the data that you need. Filter the columns that you want, in this case the customer ID column and the product-related columns. Remove the columns that you don't need for aggregating the data and training the model:
# Here are the product cols. In a real world scenario we would query a product table, or similar.
product_cols = ['Baby Food', 'Diapers', 'Formula', 'Lotion', 'Baby wash', 'Wipes', 'Fresh Fruits', 'Fresh Vegetables', 'Beer', 'Wine', 'Club Soda', 'Sports Drink', 'Chips', 'Popcorn', 'Oatmeal', 'Medicines', 'Canned Foods', 'Cigarettes', 'Cheese', 'Cleaning Products', 'Condiments', 'Frozen Foods', 'Kitchen Items', 'Meat', 'Office Supplies', 'Personal Care', 'Pet Supplies', 'Sea Food', 'Spices']
# Here we get the customer ID and the products they purchased
df_filtered = df.select(['CUST_ID'] + product_cols)
# Run the `display()` command again, this time to view the filtered information:
# + pixiedust={"displayParams": {"handlerId": "tableView"}}
display(df_filtered)
# -
# Now, aggregate the individual transactions for each customer to get a single score per product, per customer.
# + pixiedust={"displayParams": {"handlerId": "tableView"}}
df_customer_products = df_filtered.groupby('CUST_ID').sum() # Use customer IDs to group transactions by customer and sum them up
df_customer_products = df_customer_products.drop('sum(CUST_ID)')
display(df_customer_products)
# -
# <a id="build_model"></a>
# ### 3.2 Create clusters and define the model
#
# Create 100 clusters with a *k*-means model based on the number of times a specific customer purchased a product.
#
# | No Clustering | Clustering |
# |------|------|
# |  |  |
#
# First, create a feature vector by combining the product and quantity columns:
# + pixiedust={"displayParams": {"handlerId": "dataframe"}}
assembler = VectorAssembler(inputCols=["sum({})".format(x) for x in product_cols],outputCol="features") # Assemble vectors using product fields
# -
# Next, create the *k*-means clusters and the pipeline to define the model:
kmeans = KMeans(maxIter=50, predictionCol="cluster").setK(100).setSeed(1) # Initialize model
pipeline = Pipeline(stages=[assembler, kmeans])
model = pipeline.fit(df_customer_products)
# Finally, calculate the cluster for each customer by running the original dataset against the *k*-means model:
# + pixiedust={"displayParams": {"handlerId": "tableView"}}
df_customer_products_cluster = model.transform(df_customer_products)
display(df_customer_products_cluster)
# -
# <a id="persist"></a>
# ## 4. Persist the model
#
# In this section you will learn how to store your pipeline and model in Watson Machine Learning repository by using Python client libraries.
# ### 4.1 Configure IBM Watson Machine Learning credentials
#
# To access your machine learning repository programmatically, you need to copy in your credentials, which you can see in your **IBM Watson Machine Learning** service details in IBM Cloud.
#
# > **IMPORTANT**: Update `apikey` and `instance_id` below. Credentials can be found on _Service Credentials_ tab of the Watson Machine Learning service instance created on the IBM Cloud.
# +
# @hidden_cell
wml_credentials = {
"apikey": "***",
"iam_apikey_description": "Auto-generated for key ***",
"iam_apikey_name": "Service credentials-1",
"iam_role_crn": "crn:v1:bluemix:public:iam::::serviceRole:Writer",
"iam_serviceid_crn": "crn:v1:bluemix:public:iam-identity::a/***",
"instance_id": "***",
"url": "https://us-south.ml.cloud.ibm.com"
}
print(wml_credentials)
# -
# Connect to the Watson Machine Learning service using the provided credentials.
from watson_machine_learning_client import WatsonMachineLearningAPIClient
client = WatsonMachineLearningAPIClient(wml_credentials)
print(client.version)
# ### 4.2 Save the model
#
# #### Save the model to the Watson Machine Learning repository
#
# You use the Watson Machine Learning client's [Repository class](http://wml-api-pyclient.mybluemix.net/#repository) to store and manage models in the Watson Machine Learning service.
#
# > **NOTE**: You can also use Watson Studio to manage models. In this notebook we are using the client library instead.
train_data = df_customer_products.withColumnRenamed('CUST_ID', 'label')
# > **TIP**: Update the cell below with your name, email, and name you wish to give to your model.
model_props = {client.repository.ModelMetaNames.AUTHOR_NAME: "IBM",
client.repository.ModelMetaNames.NAME: "Shopping Recommendation Engine"}
published_model = client.repository.store_model(model=model, pipeline=pipeline, meta_props=model_props, training_data=train_data)
# > **NOTE**: You can delete a model from the repository by calling `client.repository.delete`.
# #### Display list of existing models in the Watson Machine Learning repository
client.repository.list_models()
# #### Display information about the saved model
import json
saved_model_uid = client.repository.get_model_uid(published_model)
model_details = client.repository.get_details(saved_model_uid)
print(json.dumps(model_details, indent=2))
# <a id="deploy"></a>
# ## 5. Deploy model to the IBM cloud
#
# You use the Watson Machine Learning client's [Deployments class](http://wml-api-pyclient.mybluemix.net/#deployments) to deploy and score models.
#
# ### 5.1 Create an online deployment for the model
#
created_deployment = client.deployments.create(saved_model_uid, 'Shopping Recommendation Engine Deployment')
# ### 5.2 Retrieve the scoring endpoint for this model
scoring_endpoint = client.deployments.get_scoring_url(created_deployment)
print(scoring_endpoint)
# <a id="test_deploy"></a>
# ### 5.3 Test the deployed model
#
# To verify that the model was successfully deployed to the cloud, you'll specify a customer ID, for example customer 12027, to predict this customer's cluster against the Watson Machine Learning deployment, and see if it matches the cluster that was previously associated this customer ID.
customer = df_customer_products_cluster.filter('CUST_ID = 12027').collect()
print("Previously calculated cluster = {}".format(customer[0].cluster))
# To determine the customer's cluster using Watson Machine Learning, you need to load the customer's purchase history. This function uses the local data frame to select every product field and the number of times that customer 12027 purchased a product.
from six import iteritems
def get_product_counts_for_customer(cust_id):
cust = df_customer_products.filter('CUST_ID = {}'.format(cust_id)).collect()
fields = []
values = []
for row in cust:
for product_col in product_cols:
field = 'sum({})'.format(product_col)
value = row[field]
fields.append(field)
values.append(value)
return (fields, values)
# This function takes the customer's purchase history and calls the scoring endpoint:
def get_cluster_from_watson_ml(fields, values):
scoring_payload = {'fields': fields, 'values': [values]}
predictions = client.deployments.score(scoring_endpoint, scoring_payload)
return predictions['values'][0][len(product_cols)+1]
# Finally, call the functions defined above to get the product history, call the scoring endpoint, and get the cluster associated to customer 12027:
product_counts = get_product_counts_for_customer(12027)
fields = product_counts[0]
values = product_counts[1]
print("Cluster calculated by Watson ML = {}".format(get_cluster_from_watson_ml(fields, values)))
# <a id="create_recomm"></a>
# ## 6. Create product recommendations
#
# Now you can create some product recommendations.
#
# First, run this cell to create a function that queries the database and finds the most popular items for a cluster. In this case, the **df_customer_products_cluster** dataframe is the database.
# This function gets the most popular clusters in the cell by grouping by the cluster column
def get_popular_products_in_cluster(cluster):
df_cluster_products = df_customer_products_cluster.filter('cluster = {}'.format(cluster))
df_cluster_products_agg = df_cluster_products.groupby('cluster').sum()
row = df_cluster_products_agg.rdd.collect()[0]
items = []
for product_col in product_cols:
field = 'sum(sum({}))'.format(product_col)
items.append((product_col, row[field]))
sortedItems = sorted(items, key=lambda x: x[1], reverse=True) # Sort by score
popular = [x for x in sortedItems if x[1] > 0]
return popular
# Now, run this cell to create a function that will calculate the recommendations based on a given cluster. This function finds the most popular products in the cluster, filters out products already purchased by the customer or currently in the customer's shopping cart, and finally produces a list of recommended products.
# This function takes a cluster and the quantity of every product already purchased or in the user's cart
from pyspark.sql.functions import desc
def get_recommendations_by_cluster(cluster, purchased_quantities):
# Existing customer products
print('PRODUCTS ALREADY PURCHASED/IN CART:')
customer_products = []
for i in range(0, len(product_cols)):
if purchased_quantities[i] > 0:
customer_products.append((product_cols[i], purchased_quantities[i]))
df_customer_products = sc.parallelize(customer_products).toDF(["PRODUCT","COUNT"])
df_customer_products.show()
# Get popular products in the cluster
print('POPULAR PRODUCTS IN CLUSTER:')
cluster_products = get_popular_products_in_cluster(cluster)
df_cluster_products = sc.parallelize(cluster_products).toDF(["PRODUCT","COUNT"])
df_cluster_products.show()
# Filter out products the user has already purchased
print('RECOMMENDED PRODUCTS:')
df_recommended_products = df_cluster_products.alias('cl').join(df_customer_products.alias('cu'), df_cluster_products['PRODUCT'] == df_customer_products['PRODUCT'], 'leftouter')
df_recommended_products = df_recommended_products.filter('cu.PRODUCT IS NULL').select('cl.PRODUCT','cl.COUNT').sort(desc('cl.COUNT'))
df_recommended_products.show(10)
# Next, run this cell to create a function that produces a list of recommended items based on the products and quantities in a user's cart. This function uses Watson Machine Learning to calculate the cluster based on the shopping cart contents and then calls the **get_recommendations_by_cluster** function.
# This function would be used to find recommendations based on the products and quantities in a user's cart
def get_recommendations_for_shopping_cart(products, quantities):
fields = []
values = []
for product_col in product_cols:
field = 'sum({})'.format(product_col)
if product_col in products:
value = quantities[products.index(product_col)]
else:
value = 0
fields.append(field)
values.append(value)
return get_recommendations_by_cluster(get_cluster_from_watson_ml(fields, values), values)
# Run this cell to create a function that produces a list of recommended items based on the purchase history of a customer. This function uses Watson Machine Learning to calculate the cluster based on the customer's purchase history and then calls the **get_recommendations_by_cluster** function.
# This function is used to find recommendations based on the purchase history of a customer
def get_recommendations_for_customer_purchase_history(customer_id):
product_counts = get_product_counts_for_customer(customer_id)
fields = product_counts[0]
values = product_counts[1]
return get_recommendations_by_cluster(get_cluster_from_watson_ml(fields, values), values)
# Now you can take customer 12027 and produce a recommendation based on that customer's purchase history:
get_recommendations_for_customer_purchase_history(12027)
# Now, take a sample shopping cart and produce a recommendation based on the items in the cart:
get_recommendations_for_shopping_cart(['Diapers','Baby wash','Oatmeal'],[1,2,1])
# The next optional section outlines how you can easily expose recommendations to notebook users, for example for test purposes.
#
# <a id="test_recomm"></a>
# ### 6.1 Test product recommendations model
#
# You can interactively test your product recommendations model using a simple PixieApp. [PixieApps](https://pixiedust.github.io/pixiedust/pixieapps.html) encapsulate business logic and data visualizations, making it easy for notebook users to explore data without having to write any code. Typically these applications are pre-packaged and imported into a notebook. However, for illustrative purposes we've embedded the product recommendation source code in this notebook.
#
# 
#
# Run the cell below, add items to the shopping cart and click the _Refresh_ button to review the recommendation results.
# + pixiedust={"displayParams": {}}
# This function takes a cluster and the quantity of every product already purchased or in the user's cart & returns the data frame of recommendations for the PixieApp
from pyspark.sql.functions import desc
def get_recommendations_by_cluster_app(cluster, purchased_quantities):
# Existing customer products
customer_products = []
for i in range(0, len(product_cols)):
if purchased_quantities[i] > 0:
customer_products.append((product_cols[i], purchased_quantities[i]))
df_customer_products = sc.parallelize(customer_products).toDF(["PRODUCT","COUNT"])
# Get popular products in the cluster
cluster_products = get_popular_products_in_cluster(cluster)
df_cluster_products = sc.parallelize(cluster_products).toDF(["PRODUCT","COUNT"])
# Filter out products the user has already purchased
df_recommended_products = df_cluster_products.alias('cl').join(df_customer_products.alias('cu'), df_cluster_products['PRODUCT'] == df_customer_products['PRODUCT'], 'leftouter')
df_recommended_products = df_recommended_products.filter('cu.PRODUCT IS NULL').select('cl.PRODUCT','cl.COUNT').sort(desc('cl.COUNT'))
return df_recommended_products
# PixieDust sample application
from pixiedust.display.app import *
@PixieApp
class RecommenderPixieApp:
def setup(self):
self.product_cols = product_cols
def computeUserRecs(self, shoppingcart):
#format products and quantities from shopping cart display data
lst = list(zip(*[(item.split(":")[0],int(item.split(":")[1])) for item in shoppingcart.split(",")]))
products = list(lst[0])
quantities = list(lst[1])
#format for the Model function
lst = list(zip(*[('sum({})'.format(item),quantities[products.index(item)] if item in products else 0) for item in self.product_cols]))
fields = list(lst[0])
values = list(lst[1])
#call the run Model function
recommendations_df = get_recommendations_by_cluster_app(get_cluster_from_watson_ml(fields, values), values)
recs = [row["PRODUCT"] for row in recommendations_df.rdd.collect()]
return recs[:5]
@route(shoppingCart="*")
def _recommendation(self, shoppingCart):
recommendation = self.computeUserRecs(shoppingCart)
self._addHTMLTemplateString(
"""
<table style="width:100%"> {% for item in recommendation %} <tr> <td type="text" style="text-align:left">{{item}}</td> </tr> {% endfor %} </table>
""", recommendation = recommendation)
@route()
def main(self):
return """
<script>
function getValuesRec(){
return $( "input[id^='prod']" )
.filter(function( index ) {
return parseInt($(this).val()) > 0;})
.map(function(i, product) {
return $(product).attr("name") + ":" + $(product).val();
}).toArray().join(",");}
function getValuesCart(){
return $( "input[id^='prod']" )
.filter(function( index ) {
return parseInt($(this).val()) > 0; })
.map(function(i, product) {
return $(product).attr("name") + ":" + $(product).val();
}).toArray(); }
function populateCart(field) {
user_cart = getValuesCart();
$("#user_cart{{prefix}}").html("");
if (user_cart.length > 0) {
for (var i in user_cart) {
var item = user_cart[i];
var item_arr = item.split(":")
$("#user_cart{{prefix}}").append('<tr><td style="text-align:left">'+item_arr[1]+" "+item_arr[0]+"</td></tr>"); } }
else { $("#user_cart{{prefix}}").append('<tr><td style="text-align:left">'+ "Cart Empty" +"</td></tr>"); } }
function increase_by_one(field) {
nr = parseInt(document.getElementById(field).value);
document.getElementById(field).value = nr + 1;
populateCart(field); }
function decrease_by_one(field) {
nr = parseInt(document.getElementById(field).value);
if (nr > 0) { if( (nr - 1) >= 0) { document.getElementById(field).value = nr - 1; } }
populateCart(field); }
</script>
<table> Products: {% for item in this.product_cols %}
{% if loop.index0 is divisibleby 4 %} <tr> {% endif %}
<div class="prod-quantity">
<td class="col-md-3">{{item}}:</td><td><input size="2" id="prod{{loop.index}}{{prefix}}" class="prods" type="text"
style="text-align:center" value="0" name="{{item}}" /></td>
<td><button onclick="increase_by_one('prod{{loop.index}}{{prefix}}');">+</button></td>
<td><button onclick="decrease_by_one('prod{{loop.index}}{{prefix}}');">-</button></td>
</div>
{% if ((not loop.first) and (loop.index0 % 4 == 3)) or (loop.last) %} </tr> {% endif %}
{% endfor %} </table>
<div class="row">
<div class="col-sm-6"> Your Cart: </div>
<div class="col-sm-6"> Your Recommendations: <button pd_options="shoppingCart=$val(getValuesRec)" pd_target="recs{{prefix}}">
<pd_script type="preRun"> if (getValuesRec()==""){alert("Your cart is empty");return false;} return true;
</pd_script>Refresh </button>
</div>
</div>
<div class="row">
<div class="col-sm-3"> <table style="width:100%" id="user_cart{{prefix}}"> </table> </div> <div class="col-sm-3"> </div>
<div class="col-sm-3" id="recs{{prefix}}" pd_loading_msg="Calling your model in Watson ML"></div> <div class="col-sm-3"> </div>
</div>
"""
#run the app
RecommenderPixieApp().run(runInDialog='false')
# -
# ## <font color=green>Congratulations</font>, you've sucessfully created a recommendation engine, deployed it to the Watson Machine Learning service, and created a PixieApp!
#
# You can now switch to the Watson Machine Learning console to deploy the model and then test it in application, or continue within the notebook to deploy the model using the APIs.
| notebooks/wml-product-recommendation-engine.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (Data Science)
# language: python
# name: python3__SAGEMAKER_INTERNAL__arn:aws:sagemaker:us-east-1:081325390199:image/datascience-1.0
# ---
# # Task 1: Import Dataset
# +
import pandas as pd
# read the csv file
insurance_df = pd.read_csv('insurance_data.csv')
# -
insurance_df.head()
# # Task 2: Upload the data to S3 bucket
# +
import sagemaker
# Creating Sagemaker session using Sagemaker SDK
sess = sagemaker.Session()
#Prefix that we want to use
prefix = 'modern-ai-zero-coding/input'
#Uploading the data to S3 bucket
uri = sess.upload_data(path = 'insurance_data.csv', key_prefix = prefix)
print(uri)
# +
# location to store the output
output = "s3://sagemaker-us-east-1-126821927778/modern-ai-zero-coding/output"
print(output)
# -
| ML_projects/6. Insurance Premium with AWS SageMaker AutoPilot/Insurance_Prediction_AutoPilot_Startup.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import sys
import os
import argparse
import time
import numpy as np
sys.path.append('../')
import torch
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
import torch.backends.cudnn as cudnn
import torch.optim.lr_scheduler as lr_scheduler
import torchvision
import torchvision.transforms as transforms
import matplotlib.pyplot as plt
import easydict as edict
from lib import models, datasets
import math
# +
# parameters
args = edict
# imagenet
args.cache = '../checkpoint/train_features_labels_cache/instance_imagenet_train_feature_resnet50.pth.tar'
args.val_cache = '../checkpoint/train_features_labels_cache/instance_imagenet_val_feature_resnet50.pth.tar'
args.save_path = '../checkpoint/pseudos/unsupervised_imagenet32x32_nc_wrn-28-2'
os.makedirs(args.save_path, exist_ok=True)
args.low_dim = 128
args.num_class = 1000
args.rng_seed = 0
# +
ckpt = torch.load(args.cache)
train_labels, train_features = ckpt['labels'], ckpt['features']
ckpt = torch.load(args.val_cache)
val_labels, val_features = ckpt['val_labels'], ckpt['val_features']
train_features = torch.cat([val_features, train_features], dim=0)
train_labels = torch.cat([val_labels, train_labels], dim=0)
print(train_features.dtype, train_labels.dtype)
print(train_features.shape, train_labels.shape)
# -
# # use cpu because the following computation need a lot of memory
device = 'cpu'
train_features, train_labels = train_features.to(device), train_labels.to(device)
# +
num_train_data = train_labels.shape[0]
num_class = torch.max(train_labels) + 1
torch.manual_seed(args.rng_seed)
torch.cuda.manual_seed_all(args.rng_seed)
perm = torch.randperm(num_train_data).to(device)
print(perm)
# -
# # soft label
class AverageMeter(object):
"""Computes and stores the average and current value"""
def __init__(self):
self.reset()
def reset(self):
self.val = 0
self.avg = 0
self.sum = 0
self.count = 0
def update(self, val, n=1):
self.val = val
self.sum += val * n
self.count += n
self.avg = self.sum / self.count
# +
n_chunks = 100
n_val = val_features.shape[0]
prec_top5 = AverageMeter()
prec_top1 = AverageMeter()
index_labeled = torch.arange(n_val, train_features.shape[0])
index_unlabeled = torch.arange(n_val)
num_labeled_data = index_labeled.shape[0]
for i_chunks, index_unlabeled_chunk in enumerate(index_unlabeled.chunk(n_chunks)):
# calculate similarity matrix
dist = torch.mm(train_features[index_unlabeled_chunk], train_features[index_labeled].t())
K = min(num_labeled_data, 200)
bs = index_unlabeled_chunk.shape[0]
yd, yi = dist.topk(K, dim=1, largest=True, sorted=True)
candidates = train_labels.view(1,-1).expand(bs, -1)
retrieval = torch.gather(candidates, 1, index_labeled[yi])
retrieval_one_hot = torch.zeros(bs * K, num_class).to(device)
retrieval_one_hot.scatter_(1, retrieval.view(-1, 1), 1)
temperature = 0.1
yd_transform = (yd / temperature).exp_()
probs = torch.sum(torch.mul(retrieval_one_hot.view(bs, -1 , num_class), yd_transform.view(bs, -1, 1)), 1)
probs.div_(probs.sum(dim=1, keepdim=True))
probs_sorted, predictions = probs.sort(1, True)
correct = predictions.eq(train_labels[index_unlabeled_chunk].data.view(-1,1))
top5 = torch.any(correct[:, :5], dim=1).float().mean()
top1 = correct[:, 0].float().mean()
prec_top5.update(top5, bs)
prec_top1.update(top1, bs)
print('[{}]/[{}] top5={:.2%}({:.2%}) top1={:.2%}({:.2%})'.format(
i_chunks, n_chunks, prec_top5.val, prec_top5.avg, prec_top1.val, prec_top1.avg))
# +
# n_chunks = 100
# prec_top5 = AverageMeter()
# for num_labeled_data in [10000]:
# index_labeled = []
# index_unlabeled = []
# data_per_class = num_labeled_data // args.num_class
# for c in range(args.num_class):
# indexes_c = perm[train_labels[perm] == c]
# index_labeled.append(indexes_c[:data_per_class])
# index_unlabeled.append(indexes_c[data_per_class:])
# index_labeled = torch.cat(index_labeled)
# index_unlabeled = torch.cat(index_unlabeled)
# for i_chunks, index_unlabeled_chunk in enumerate(index_unlabeled.chunk(n_chunks)):
# # calculate similarity matrix
# dist = torch.mm(train_features[index_unlabeled_chunk], train_features[index_labeled].t())
# K = min(num_labeled_data, 5000)
# bs = index_unlabeled_chunk.shape[0]
# yd, yi = dist.topk(K, dim=1, largest=True, sorted=True)
# candidates = train_labels.view(1,-1).expand(bs, -1)
# retrieval = torch.gather(candidates, 1, index_labeled[yi])
# retrieval_one_hot = torch.zeros(bs * K, num_class).to(device)
# retrieval_one_hot.scatter_(1, retrieval.view(-1, 1), 1)
# temperature = 0.1
# yd_transform = (yd / temperature).exp_()
# probs = torch.sum(torch.mul(retrieval_one_hot.view(bs, -1 , num_class), yd_transform.view(bs, -1, 1)), 1)
# probs.div_(probs.sum(dim=1, keepdim=True))
# probs_sorted, predictions = probs.sort(1, True)
# correct = predictions.eq(train_labels[index_unlabeled_chunk].data.view(-1,1))
# top5 = torch.any(correct[:, :5], dim=1).float().mean()
# prec_top5.update(top5, bs)
# print('[{}]/[{}] {:.2%} {:.2%}'.format(i_chunks, n_chunks, prec_top5.val, prec_top5.avg))
# -
| notebooks/knn-imagenet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
from tpot import TPOTClassifier
from sklearn.model_selection import train_test_split
import pandas as pd
import numpy as np
from fancyimpute import IterativeSVD, SoftImpute
train = pd.read_csv('train.csv')
train
# ## Data janitor
# We need a class named 'class' for TPOT, and I want to have a title (Mr, Mrs, Dr.) by splitting the names by dots, the title is a good predictor.
# +
train.rename(columns={'Survived':'class'}, inplace=True)
train['Last_name'], train['First_names'] =train['Name'].str.split(',',1).str
train['Title'] = train['First_names'].str.split('.',).str[0].map(lambda x:x.strip())
# used to do Deck, but too many missing values
# train['Deck'] = train['Cabin'].str[0]
# used to replace NA by -999, for now using imputation
#train = train.fillna(-999)
#title_translate_dict = {-999:-999}
#for counter, name in enumerate(train['Title'].unique()):
# if name in title_translate_dict: continue
# title_translate_dict[name] = counter
#title_translate_dict[]
# -
# Replace sex with numbers
train['Sex'] = train['Sex'].map({'male':0,'female':1})
#train['Embarked'] = train['Embarked'].map({'S':0,'C':1,'Q':2})
#train['Title'] = train['Title'].map(title_translate_dict)
#train['Deck'] = train['Cabin'].str[0]
#train = train.fillna(-999)
#train.head(5)
# Take three classes and sorta one-hot encode them
# +
from sklearn.preprocessing import MultiLabelBinarizer
mlb = MultiLabelBinarizer()
#CabinTrans = mlb.fit_transform([{str(val)} for val in train['Cabin'].values])
TitleTrans = mlb.fit_transform([{str(val)} for val in train['Title'].values])
PClassTrans = mlb.fit_transform([{str(val)} for val in train['Pclass'].values])
EmbarkedTrans = mlb.fit_transform([{str(val)} for val in train['Embarked'].values])
# -
PClassTrans
TitleTrans
EmbarkedTrans
train_new = train.drop(['Name','Pclass','Ticket','Cabin','class', 'Last_name', 'First_names','Title', 'Embarked'], axis=1)
train_new.head(5)
# Now plop all the tables together and impute
# +
#train_new = np.hstack((train_new.values,CabinTrans))
train_new = np.hstack((train_new.values, TitleTrans))
train_new = np.hstack((train_new, PClassTrans))
train_new = np.hstack((train_new, EmbarkedTrans))
train_new = SoftImpute().complete(train_new)
#train_new = train_new.values
# -
train_class = train['class'].values
training_indices, validation_indices = training_indices, testing_indices = train_test_split(train.index, stratify = train_class, train_size=0.75, test_size=0.25)
training_indices.size, validation_indices.size
tpot = TPOTClassifier(verbosity=2, generations=5)
tpot.fit(train_new[training_indices], train_class[training_indices])
tpot.score(train_new[validation_indices], train.loc[validation_indices, 'class'].values)
tpot.export('tpot_titanic_pipeline.py')
# +
titanic_sub = pd.read_csv('test.csv')
titanic_sub['Last_name'], titanic_sub['First_names'] = titanic_sub['Name'].str.split(',',1).str
titanic_sub['Title'] = titanic_sub['First_names'].str.split('.',).str[0].map(lambda x:x.strip())
# There is one wonderfully named Oliva y Ocana, Dona. Fermina , which I assume is equivalent to 'Lady'
# we don't have it in the training set so replace it
titanic_sub['Title'].replace('Dona', 'Lady', inplace=True)
for var in ['Cabin', 'Title']:
new = list(set(titanic_sub[var]) - set(train[var]))
titanic_sub.ix[titanic_sub[var].isin(new), var] = np.nan
titanic_sub['Sex'] = titanic_sub['Sex'].map({'male':0,'female':1})
#titanic_sub['Embarked'] = titanic_sub['Embarked'].map({'S':0,'C':1,'Q':2})
#titanic_sub['Title'] = titanic_sub['Title'].map(title_translate_dict)
#print(train['Title'].unique())
#titanic_sub = titanic_sub.fillna(-999)
mlb = MultiLabelBinarizer()
TitleTrans = mlb.fit([{str(val)} for val in train['Title'].values]).transform([{str(val)} for val in titanic_sub['Title'].values])
PClassTrans = mlb.fit([{str(val)} for val in train['Pclass'].values]).transform([{str(val)} for val in titanic_sub['Pclass'].values])
EmbarkedTrans = mlb.fit([{str(val)} for val in train['Embarked'].values]).transform([{str(val)} for val in titanic_sub['Embarked'].values])
titanic_sub = titanic_sub.drop(['Name','Pclass','Ticket','Cabin', 'Last_name', 'First_names','Title', 'Embarked'], axis=1)
titanic_sub_new = np.hstack((titanic_sub.values, TitleTrans))
titanic_sub_new = np.hstack((titanic_sub_new, PClassTrans))
titanic_sub_new = np.hstack((titanic_sub_new, EmbarkedTrans))
titanic_sub_new = SoftImpute().complete(titanic_sub_new)
#assert (train_new.shape[1] == titanic_sub_new.shape[1]), "Not Equal"
submission = tpot.predict(titanic_sub_new)
final = pd.DataFrame({'PassengerId': titanic_sub['PassengerId'], 'Survived': submission})
final.to_csv('submission.csv', index = False)
# -
# After submission:
# # Your submission scored 0.78947, which is not an improvement of your best score. Keep trying!
| Titanic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/hrai/pytorch-tutorials/blob/master/PyTorch_tutorial.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="xi9zSUhtNl-j" colab_type="text"
# ## PyTorch Basics
#
# + id="KPfaajzmEdTP" colab_type="code" colab={}
from __future__ import print_function
import torch
# + id="qL7AJYfkEj6s" colab_type="code" outputId="c55bd897-f4cd-49fa-ecd8-c54e7cd8b083" colab={"base_uri": "https://localhost:8080/", "height": 104}
x=torch.empty(5,3)
print(x)
# + id="PKC382VtEtD8" colab_type="code" outputId="52a8eb9c-80e9-4b0b-8ac0-c55ede538627" colab={"base_uri": "https://localhost:8080/", "height": 104}
x=torch.rand(5,3)
print(x)
# + id="6AF99p7xEzEP" colab_type="code" outputId="ab01ff84-5b44-42ec-f4ea-3f27c7c19e02" colab={"base_uri": "https://localhost:8080/", "height": 104}
x=torch.zeros(5,3, dtype=torch.long)
x
# + id="XuSvM0BsFUu6" colab_type="code" outputId="91768060-a195-4524-e1b8-b81ecc15ac25" colab={"base_uri": "https://localhost:8080/", "height": 34}
x = torch.tensor([5.5, 3])
print(x)
# + id="3FW6viMJFh0q" colab_type="code" outputId="cb9da5fa-263f-45ff-b35d-f45b8afc49e8" colab={"base_uri": "https://localhost:8080/", "height": 191}
x=x.new_ones(5,3,dtype=torch.double)
print(x)
x=torch.randn_like(x, dtype=torch.float)
print(x)
# + id="G5xgAV4HF6ov" colab_type="code" outputId="c1cfeb30-171e-400c-d8b1-bc9659acebed" colab={"base_uri": "https://localhost:8080/", "height": 34}
print(x.size())
# + id="JZvrDP4XGDD8" colab_type="code" outputId="283dca62-8df7-44bf-9da8-1085ea3c74ed" colab={"base_uri": "https://localhost:8080/", "height": 104}
y=torch.rand(5,3)
print(x+y)
# + id="1fKO4n_fGPGA" colab_type="code" outputId="0e60ed0c-1ca7-4221-bfc9-4f7c21f50fc4" colab={"base_uri": "https://localhost:8080/", "height": 104}
print(torch.add(x,y))
# + id="2KgKMwj0GVcB" colab_type="code" outputId="74ff0909-0a3d-4c00-e561-aa2075fb44c1" colab={"base_uri": "https://localhost:8080/", "height": 104}
result=torch.empty(5,3)
torch.add(x,y,out=result)
print(result)
# + id="lfPCmFDxGocO" colab_type="code" outputId="e9eabaee-f72d-4a60-d3e7-d6d3b576f30a" colab={"base_uri": "https://localhost:8080/", "height": 104}
y.add_(x)
print(y)
# + id="VHBTqUv4ICab" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 139} outputId="019ee6ea-eabf-4d2c-c907-4d007547ad95"
print(x)
print(x[:,2])
print(x[2,:])
# + id="OdTa0G-3IGI1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="dd3ca99b-c562-4048-f229-446bd90dc631"
x=torch.randn(4,4)
y=x.view(16)
z=x.view(-1,8)
print(x.size(),y.size(),z.size())
# + id="CMSYsuNxNDwK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 141} outputId="8fecb5e7-3049-4f06-ab5b-0c902370679f"
print(x)
print(y)
# + id="MXlF7DsFNVMz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="f470b554-919e-40b8-c755-5ff90fbd6902"
x=torch.randn(1)
print(x)
print(x.item())
# + [markdown] id="3x8mTnT4NvCh" colab_type="text"
# ## NumPy bridge
# + id="ylXi4pV5OAft" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="e99dd133-e4ab-4699-af23-7dac4966c5eb"
a=torch.ones(5)
print(a)
# + id="EWQWfd4eOFgT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="65a121f5-54f7-4764-d817-836ed9ae182f"
b=a.numpy()
print(b)
# + id="u-i3iNgKOSKs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="5ffeb131-a088-4c35-a223-f4731a5d3cb7"
a.add_(1)
print(a)
print(b)
# + id="aucD_sabOskS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="71e3196e-e6df-4d0c-bae1-f40b02e5a84d"
import numpy as np
a=np.ones(5)
b=torch.from_numpy(a)
np.add(a,1,out=a)
print(a)
print(b)
# + id="J_9LZhoOPIGk" colab_type="code" colab={}
# let us run this cell only if CUDA is available
# We will use ``torch.device`` objects to move tensors in and out of GPU
if torch.cuda.is_available():
device = torch.device("cuda") # a CUDA device object
y = torch.ones_like(x, device=device) # directly create a tensor on GPU
x = x.to(device) # or just use strings ``.to("cuda")``
z = x + y
print(z)
print(z.to("cpu", torch.double)) # ``.to`` can also change dtype together!
| PyTorch_tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction to Version Control with GIT
# + [markdown] slideshow={"slide_type": "slide"}
# ## Introduction
#
# **Version control** is a system which:
#
# - records all files that make up a project (down to the line) over time
# - tracks their development
# - provides the ability to recall previous versions of files.
# - facilitates collaborative editing of files by different parties.
#
# This type of system is essential for ensuring reproducibility of scientific research
#
# Why should I care obout version control? Probably most of us have at some point in their career (some earlier, some later) come across as situation where they had to write document that evolved over time.It starts out simple, than more and more content comes in, often content needs to be reorganized, certain parts will be deleted or moved to other places. Changes introduced are reverted and changed in a different way etc. More often than not one wishes to travel back in time to an earlier stage in the history of the project.
# -
# There are a number of popular tools for version control, the particular tool we will use is **git**.
# + [markdown] slideshow={"slide_type": "slide"}
# A "poor man's version control" system is shown in this directory listing:
# -
# ```shell
# # ls ~/poor_VC
#
# paper_final.tex paper_v2_richard.tex
# paper_really_final.tex paper_v2.tex
# paper.tex paper_v3_with_richards_comments.tex
# paper_v1.tex
# ```
# GIT does away with different file names for different versions. You will always see the same filename in your projects directory. Git will save snapshots of all (tracked) files on request and you will be able to navigate back an forth through the history that any tracked file went through. Moreover, you *branch* off
# to test a new idea, switch back to the original branch (called the *master*) and merge branches that you or a collaborator created. One such git workflow is depicted below:
#
# 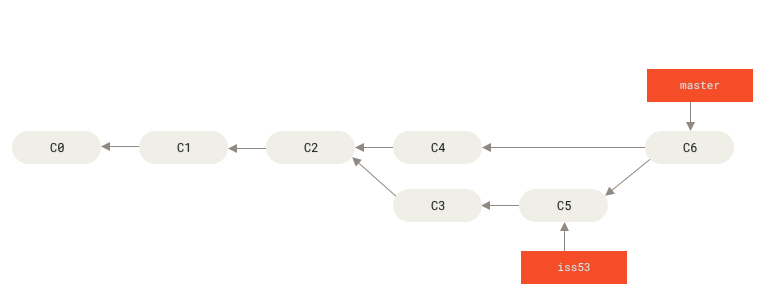
#
# Source: https://git-scm.com/book/en/v2
# In this example, the developer started to work on a file and created three snapshots (*commits* in git-speak): C0, C1, C2. She then wants to try out a new, highly experimental feature that she is not sure if it will break the code. So she creates a new branch (iss53) and commits C3 to it.
# Then, she reckognizes that something must be changed in the main line of development, so she switches back to the master branch and commits C4.
# Again back to the feature branch *iss53* and commit C5. At this point, she wants to integrate the new feature from branch *iss53* into the *master* branch. The merge
# point becomes a new *commit*, C6. And so forth. At no point did she have to rename or backup her files, everything is handled by git.
# The git version control system can be thought of as a iterative workflow where paper documents move between various places:
#
# 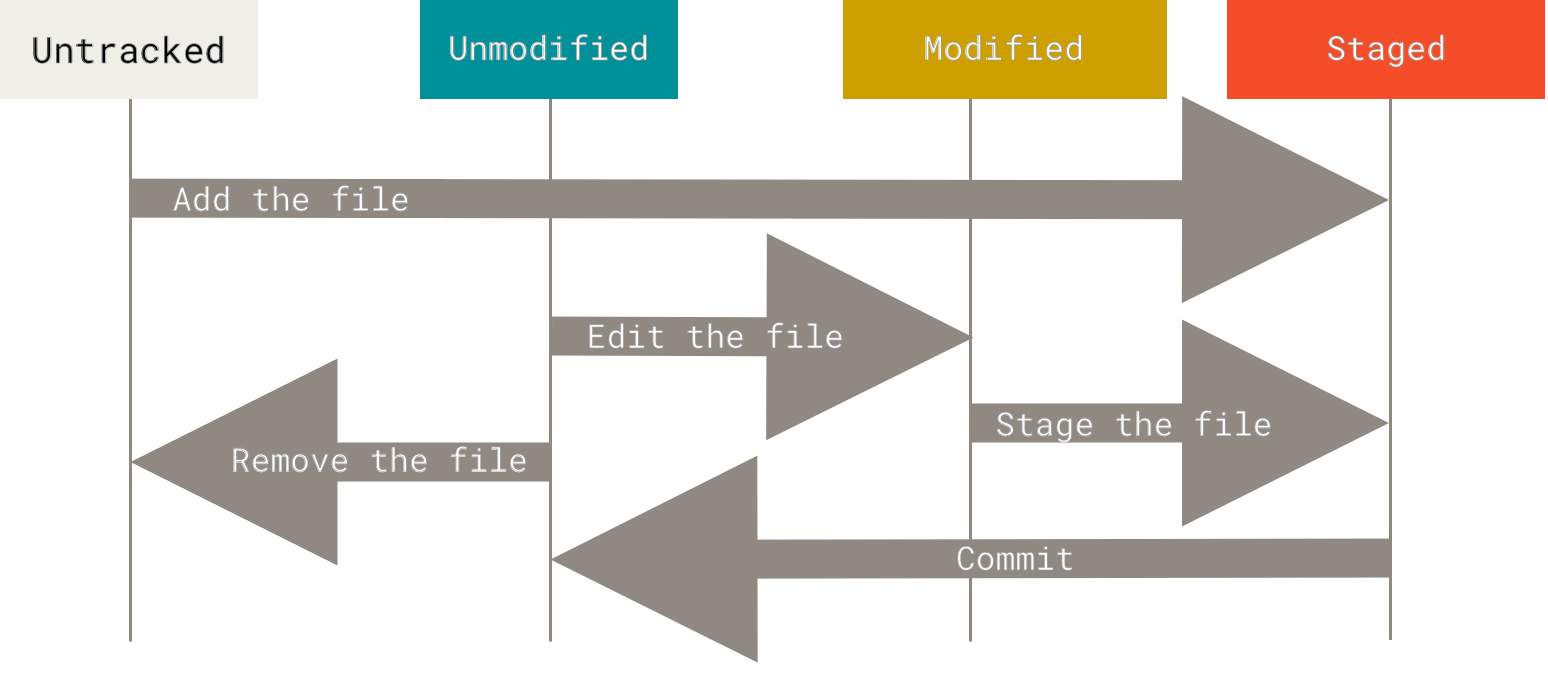
#
# - The modified state: The document is on our desk before us and we edit (write) it.
# - Staged: When our task on the document is done or we have to work on sth else, we put the file into a shelf that has a label "Keep" on it. It can be thought of as a stash of documents that we need to keep within reach but are not actively working on it. From that stash, we can easily pull the document back to our desk and continue working on it.
# - Committed: At some point, we would like to take a snapshot of the document, so we can re-open precisely the current state at a later point in time. In a paper world, we would take the stash of documents to the copy machine, make a copy from each document and put the copy into a file cabinet, tagged with a date, our name, and a remark telling us what the change of this copy with respect to the previous copy is. The original document is now in a "unmodified" state (with respect to the last backup copy).
#
# We then take the document back to our desk and continue working on it: It is then again in the "modified state".
#
# 
# # Setting up git
#
# Initially we need to set up git.
#
# git keeps track of the entire history of a project. This does not only mean keeping track of what was done but also who did it. So we start by telling git who we are by running the following two commands:
#
# ```shell
# $ git config --global user.name "<NAME>"
# $ git config --global user.email "Your Email"
# ```
#
# **Note** this is not data that is being collected by any cloud service or similar. It just stays with your project.
#
# **Windows**
# Note that all these commands work on the anaconda prompt but if you want to use tab completion you can use the git bash command line specifically for git.
#
#
# Moreover, we are going to set [Nano](https://www.nano-editor.org) as the default editor for git. For unix you can use the following command:
#
# ```shell
# $ git config --global core.editor "nano"
# ```
#
# We are going to use Nano later in order to write a commit message.
#
#
# ## Exercise:
# 1. Configure your `git` installation as shown above. Set your name, email, and preferred editor.
# 1. Lookup the documentation for the `git config`. Find out how to list the current global git configuration.
# 1. List the current global git configuration and confirm that your user name, email and preferred editor are setup correctly.
# ```
#
# ## Solutions:
# 1. Configure git
# ```shell
# $ git config --global user.name "<NAME>"
# $ git config --global user.email "<EMAIL>"
# ```
#
# 1. `git-config` documentation
# ```shell
# $ git config -h
# ```
# prints the usage instructions to the terminal:
# ```
# usage: git config [<options>]
#
# Config file location
# --global use global config file
# --system use system config file
# --local use repository config file
# --worktree use per-worktree config file
# -f, --file <file> use given config file
# --blob <blob-id> read config from given blob object
#
# Action
# --get get value: name [value-regex]
# --get-all get all values: key [value-regex]
# --get-regexp get values for regexp: name-regex [value-regex]
# --get-urlmatch get value specific for the URL: section[.var] URL
# --replace-all replace all matching variables: name value [value_regex]
# --add add a new variable: name value
# --unset remove a variable: name [value-regex]
# --unset-all remove all matches: name [value-regex]
# --rename-section rename section: old-name new-name
# --remove-section remove a section: name
# -l, --list list all
# -e, --edit open an editor
# --get-color find the color configured: slot [default]
# --get-colorbool find the color setting: slot [stdout-is-tty]
#
# Type
# -t, --type <> value is given this type
# --bool value is "true" or "false"
# --int value is decimal number
# --bool-or-int value is --bool or --int
# --path value is a path (file or directory name)
# --expiry-date value is an expiry date
#
# Other
# -z, --null terminate values with NUL byte
# --name-only show variable names only
# --includes respect include directives on lookup
# --show-origin show origin of config (file, standard input, blob, command line)
# --default <value> with --get, use default value when missing entry
# ```
#
# 1. List the git configuration
# ```shell
# $ git config --global --list
# user.name=<NAME>
# user.email=<EMAIL>
# filter.lfs.clean=git-lfs clean -- %f
# filter.lfs.smudge=git-lfs smudge -- %f
# filter.lfs.process=git-lfs filter-process
# filter.lfs.required=true
# diff.tool=vimdiff
# core.editor=vim
# ```
#
#
#
#
# # Initialising a git repository
#
# In order to demonstrate how version control with git works we are going to use the `rsd-workshop` folder we created before.
#
# We need tell git to start keeping an eye on this repository (folder/project). While in the `rsd-workshop` directory type:
#
# ```shell
# $ git init
# ```
#
# You should then see a message saying that you have successfully initialized a git repository.
#
# ## Exercise
# 1. Change into your directory `rsd-workshop` and initialize it as a git repository.
# # Staging and committing changes
#
# To see the status of the repository we just initialized type:
#
# ```shell
# $ git status
# ```
#
# We should see something like:
#
# 
# There are various pieces of useful information here, first of all that `addition.py`, `if-statement.py` and `while-loops.py` are not currently tracked files.
#
# We are now going to track the `addition.py` file:
#
# ```shell
# $ git add addition.py
# ```
#
# If we run git status again we see:
#
# 
# We have propagated our file from the "Untracked" to the "Staged" status.
# 
# So the `addition.py` file is now ready to be "committed".
#
# ```shell
# $ git commit
# ```
#
# When doing this, a text editor should open up prompting you to write what is called a commit message. In our case the text editor that opens is Nano.
#
# For the purposes of using git Nano is more than a sufficient editor, all you need to know how to do is:
#
# - Write in Nano: just type;
# - Save in Nano: Ctrl + O;
# - Quit Nano: Ctrl + X.
#
# Type the following as the first commit message:
#
# ```shell
# Add addition script
#
# Addition script contains a function which adds two numbers.
# ```
#
# save and exit.
#
# git should confirm that you have successfully made your first commit.
#
#
# 
#
# **Note** A commit message is made up of 2 main components:
#
# ```shell
# <Title of the commit>
#
# <Description of what was done>
# ```
#
# - The title should be a description in the form of "if this commit is applied `<title of the commit>` will happen". The convention is for this to be rather short and to the point.
# - The description can be as long as needed and should be a helpful explanation of what is happening.
#
# A commit is a snapshot that git makes of your project, you should use this at meaningful steps of the progress of a project.
# Now, our file is in the "Unmodified" state, because it is identical to the _copy_ that we filed away to our "file cabinet" (the location where git stores the snapshots).
#
# 
# ## Exercise:
# 1. Add and commit the file `addition.py` to git. Write a short but telling commit message.
# # Ignoring files
#
# There are still two files in the repository that are currently not being tracted. These are `if-statement.py` and `while-loops.py`.
#
# We do not want to keep tract of those files as they are not related to our project.
#
# To tell git to ignore these files we will add them to a blank file entitled `.gitignore`.
#
# Open your editor and open a new file (`File > New file`) and type:
#
# ```shell
# if-statements.py
# while-loops.py
# ```
#
# Save that file as `.gitignore` and then run:
#
# ```shell
# $ git status
# ```
#
# We see now that `git` is ignoring those 2 files but is aware of the `.gitignore` file.
# ## Exercise:
# 1. Add and commit the file `.gitignore` to git. Give a good commit message.
# 1. Confirm that you have a clean working directory.
# + [markdown] jupyter={"source_hidden": true}
# ## Solution:
# ```shell
# $ git add .gitgignore
# $ git commit
# $ git status
# ```
# -
# Now if we run `git status`, we see a message saying that everything in our repository is tracked and up to date.
# # Tracking changes to files
#
#
# Let's assume that we want to refactor (a fancy way of saying "change") the function `add_two_numbers` to `add_two_even_numbers` such that the function adds two even numbers but prints a warning if not both numbers are even.
#
# ## Exercise:
# 1. Change the file `addition.py` to look like this:
#
# ```python
# # addition.py
# def add_two_even_numbers(a, b):
# if a % 2 == 0 and b % 2 == 0:
# return a + b
# else:
# print("Please use even numbers.")
#
# print(add_two_even_numbers(4, 6))
# ```
#
# 1. Save your file and confirm that git detects the change in one file.
#
# ```shell
# $ git status
# ```
#
# ## Solution:
# 
# Our file is now in the modified state, as `git status` tells us.
#
# 
# To see what has been modified you need to type:
#
# ```shell
# $ git diff addition.py
# ```
# and press `q` to exit.
#
#
# To "stage" the file for a commit we use `git add` again:
#
# ```shell
# $ git add addition.py
# ```
#
# Now let us commit:
#
# ```shell
# $ git commit
# ```
#
# With the following commit message:
#
# ```
# Change add two numbers function to add two even numbers
# ```
#
# Finally, we can check the status:
# ```shell
# $ git status
# ```
# to confirm that everything has been done correctly.
# # Exploring history
# `git` allows us to visualize the history of a project and even to change it. To view the history of a repository type:
#
# ```shell
# $ git log
# ```
#
# This displays the full log of the project:
#
# 
#
#
# We see that there are 3 commits there, each with a seemingly random set of numbers and characters. This set of characters is called a "hash".
#
# The first commit with title `adds addition script` has hash: aab73629642568b9be5ca5faa5e091ea9a629d67.
#
# **Note** that on your machines this hash will be different, in fact every hash is mathematically guaranteed to be unique, thus it is uniquely assigned to the changes made.
#
# Hashes can be very useful, e.g. to re-create an earlier state of the project.
# ## Time travel: Checking out a (previous) commit
# The `git checkout` commands allows us to revert our file(s) to a state from an earlier commit.
# Let's try this out and check out the commit just before we made the last change.
# Find the commit hash to revert to from `git log` first. Highlight the hash and copy it (Ctrl-C). Then paste into the `git checkout` command. It's actually sufficient to take only the first 8 characters from the hash. Remember that your hash will look different from the example.
# ```shell
# $ git checkout fbb2cd03
# ```
# The message about "detached HEAD" means that our file(s) have now reverted to a state from an earlier commit. Any changes to our files would not be commited directy
# back to our branch (more about branches further below).
# The history reported by `git log` now contains only the commits before the checked out commit as you can see by running
#
# ```shell
# $ git log
# ```
#
# Open the file `addition.py` and confirm that it is again in the state before we changed `add_two_numbers()` to `add_two_even_numbers()`.
# ## Check out the tip of the branch (aka HEAD)
# To go back to the "tip of the branch" (the state where we left from before checking out an earlier commit) we run
# ```shell
# $ git checkout master
# ```
# Confirm we are back on the master branch:
#
# ```shell
# $ git status
# ```
# Check the history:
#
# ```shell
# $ git log
# ```
# ## Exercise:
# 1. Checkout the previous commit.
# 1. Confirm that you are on the targeted commit.
# 1. Check that the file `addition.py` contains the ancient code, i.e. before we refactored the code to handle odd and even numbers seperately.
# 1. Go back to the tip of the `master` branch.
# # Creating branches
#
# Branches allow us to work on different versions of the same file in parallel, which is very important when developing software, but also when we do research.
# When typing `git status` we have seen that one piece of information regularly given was:
#
# ```shell
# On branch master
# ```
#
# This is telling us which branch of "history" we are currently on. We can view all branches with the command:
#
# ```shell
# $ git branch
# ```
#
# This shows:
#
# ```
# * master
# ```
#
#
# So currently there is only one branch called master.
#
# ## Exercise:
# 1. Create a new branch called `implement-add-odd-numbers`.
# 1. Confirm that the new branch has been created.
# 1. Switch ("checkout") to the new branch.
# 1. Confirm that you are on the new branch.
#
# ## Solutions:
# 1. Create branch
# ```shell
# $ git branch implement-add-odd-numbers
# ```
#
# 1. Check branches:
# ```
# $ git branch
# implement-add-odd-numbers
# * master
# ```
#
# 1. Switch to the new branch
# ```shell
# $ git checkout implement-add-odd-numbers
# ```
#
# 1. Check branch
# ```shell
# git branch
# * implement-add-odd-numbers
# master
# $ git status
# ```
# # Implement a feature in the new branch
# While we are on this branch we are going to add a new function in the `addition.py` and that is a function that adds two odd numbers.
#
# ## Exercise:
# 1. Add the following code to `addition.py`:
#
# ```python
# def add_two_odd_numbers(a, b):
# if a % 2 != 0 and b % 2 != 0:
# return a + b
# print("Please use odd numbers.")
#
# print(add_two_odd_numbers(1, 3))
#
# ```
#
# 1. Add and commit the changes to git.
# 1. Checkout the master branch and confirm thate "addition.py" is in the old state.
# 1. Switch back to the brach `implement-add-odd-numbers` and confirm the new change has been played back.
# ## Solutions:
# 1. Open "addition.py" in your editor and apply the changes.
#
# 1. Add and commit.
# ```shell
# $ git add addition.py
# $ git commit
# ```
#
# 1. Checkout master
# ```
# $ git checkout master
# ```
#
# 1. List the code in a pager.
# ```shell
# $ less addition.py
# ```
#
# 1. Checkout new branch.
# ```shell
# $ git checkout implement-add-odd-numbers
# $ less addition.py
# ```
#
#
# ### Merging locally
# The `git merge` command integrates changes from a branch into the currently checked out branch. Before merging, make sure to have checked out
# the target branch. In our example we merge changes from the "implement-add-odd-numbers" branch into the "master" branch.
# Before merging, always make sure you are on the *target* branch:
#
# ```shell
# $ git checkout master
#
# Switched to branch 'master'
# ```
# Now use the `git merge` command to bring in the changes from the "implement-add-odd-numbers" branch.
# It is good practice to always append the `--no-ff` flag to `git merge` to create a dedicated merge commit. In this way one can later identify changes in the code that are a result from a merge.
# ```shell
# $ git merge implement-add-odd-numbers --no-ff
# ```
# In your editor, the commit message will already be present, you can leave it as is or add more detailed information. Save and close the editor.
# ## Exercise:
# 1. Confirm that the changes from the branch `implement-add-odd-numbers` are properly merged into the master branch.
#
# ## Solutions:
# 1.
# ```
# $ git log
# ```
#
# Pro tip: To visualize the tree structure of branches and commits, use the following `git log` command:
# ```shell
# $ git log --pretty=oneline --graph --abbrev-commit
#
# * c1ba40d (HEAD -> master) Merge branch 'implement-add-odd-numbers'
# |\
# | * fadd60e (implement-add-odd-numbers) implement function for adding odd numbers
# |/
# * d80f1c9 change add two numbers function to add two even numbers
# * 3fca9d1 Add .gitignore
# * 8487ec4 Add addition script
# ```
# ### Delete feature branch
# If we are certain that the feature branch `implement-add-odd-numbers` will no longer be used, we can delete it:
# ```shell
# $ git branch -D implement-add-odd-numbers
# ```
# ## Exercise:
# 1. Confirm that the feature branch is deleted.
# ## Solution:
# ```shell
# $ git branch
# ```
# # Conclusions
#
# This is the end of Day 1 of our workshop. Here's what you learned today:
#
# * Using the command line to navigate the directory tree and create directories.
# * First steps in python programming
# * Using git for version control:
# - The git cycle: `add`, `commit`, `push`
# - Going back and forth in history
# - Branching: list, create, merge, delete
# # Outlook
# On day 2 of the workshop, we will cover:
# * Best practices for software development: Documentation and testing
# * Git in the cloud: GitLab, Merge requests, `git push` and `git pull`
# * Continuous integration: Automated testing
# * Gitlab pages: Publish your online reference manual
| Day I - Part III - Introduction to git.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Refine the Data
# Create the items, users and interaction matrix
#
# - **Items**: item id + metadata features
# - **Users**: User id + metadata features
# - **Interaction**: Explicit Matrix, Implicit Matrix
# +
import sys
sys.path.append("../")
import warnings
warnings.filterwarnings("ignore")
# -
import numpy as np
import pandas as pd
import re
# ## Items
#
# For the items, we will keep the following:
# - From `items_raw` dataframe
# - movie_id
# - title
# - year (release)
# - genre categories
# - From `item_features` dataframe
# - overview
# - language (original_language)
# - runtime
# - vote_average
# - vote_count
items_raw = pd.read_csv("data/items_raw.csv")
item_features = pd.read_csv("data/item_features.csv")
# *1. Get `year` from `release_title`*
items_raw["release_date"] = pd.to_datetime(items_raw.release_date, infer_datetime_format=True)
items_raw["year"] = items_raw.release_date.apply(lambda x: str(x.year))
# *2. Drop `imdb_url`, `video_release_date` & `release_date`*
items_main = items_raw.drop(['video_release_date', 'release_date', 'imdb_url'], axis=1).copy()
# +
# Match Whitespace + ( + YEAR + )
# regex_year = re.compile(r'\s\(\d{4}\)')
# items["movie"] = items.title.str.replace(regex_year, "")
# items["movie"] = items.movie.str.strip()
# -
# *3. Get the additional features from the item_features*
items_addtl = item_features[['overview', 'original_language', 'runtime', 'vote_average', 'vote_count', "movie_id"]].copy()
# *4. Merge the two dataframes*
items = pd.merge(left=items_main, right=items_addtl, on="movie_id", how="left")
items.head()
items.overview.isna().sum()
items.overview.fillna("None", inplace=True)
# # Image Features
from keras.models import Model
from keras.applications.inception_v3 import InceptionV3
from keras.models import Model
from os import listdir
from keras.preprocessing.image import load_img
from keras.preprocessing.image import img_to_array
base_modelv3 = InceptionV3(weights='imagenet', include_top=False)
model= Model(inputs=base_modelv3.input, outputs=base_modelv3.output)
# +
# model.summary()
# -
def preprocess_input(x):
x /= 255.
x -= 0.5
x *= 2.
return x
images = []
for name in listdir("data/posters"):
images.append(name)
images_1 = images[:1000]
def load_photos_predict(directory):
images = []
for name in images_1:
filename = directory + '/' + name
image = load_img(filename, target_size=(299, 299))
# convert the image pixels to a numpy array
image = img_to_array(image)
# reshape data for the model
image = np.expand_dims(image, axis=0)
# prepare the image for the model
image = preprocess_input(image)
# get image id
image_id = name.split('.')[0]
feature = model.predict(image).ravel()
images.append(feature)
return images
# %%time
poster_features = load_photos_predict("/tf/notebooks/data/data/posters/")
poster_features[:2]
# # Build nearest neighbor model
from reco.recommend import get_similar
from sklearn.neighbors import NearestNeighbors, VALID_METRICS
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import PIL
from sklearn.neighbors import NearestNeighbors
from sklearn.metrics.pairwise import cosine_distances, cosine_similarity
def get_similar(embedding, k):
model_similar_items = NearestNeighbors(n_neighbors=k, metric="cosine",
algorithm="brute").fit(embedding)
distances, indices = model_similar_items.kneighbors(embedding)
return distances, indices
# %%time
item_distances, item_similar_indices = get_similar(poster_features[:100], 5)
item_similar_indices[:5]
def show_similar(item_index, item_similar_indices):
movie_ids = item_similar_indices[item_index]
#movie_ids = item_encoder.inverse_transform(s)
images = []
for movie_id in movie_ids:
img_path = 'data/posters/' + str(movie_id+1) + '.jpg'
images.append(mpimg.imread(img_path))
plt.figure(figsize=(20,10))
columns = 5
for i, image in enumerate(images):
plt.subplot(len(images) / columns + 1, columns, i + 1)
plt.axis('off')
plt.imshow(image)
show_similar(0, item_similar_indices)
| 14-Image-Features.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from ipyleaflet import Map, Heatmap
from random import uniform
m = Map(center=[0, 0], zoom=2)
m
locations = [[uniform(-80, 80), uniform(-180, 180), uniform(0, 1000)] for i in range(1000)] # lat, lng, intensity
heat = Heatmap(locations=locations, radius=20, blur=10)
m.add_layer(heat)
heat.radius = 30
heat.blur = 50
heat.max = 0.5
heat.gradient = {0.4: 'red', 0.6: 'yellow', 0.7: 'lime', 0.8: 'cyan', 1.0: 'blue'}
heat.latlngs = [[uniform(-80, 80), uniform(-180, 180), uniform(0, 1000)] for i in range(1000)]
| examples/ipyleaflet/Heatmap.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="JA6penX4qyKt" colab_type="text"
#
# <img width="60" src="https://drive.google.com/uc?export=view&id=1JQRWCUpJNAvselJbC_K5xa5mcKl1gBQe">
#
#
# + id="5TGU-sXQqzUn" colab_type="code" outputId="99824b29-aa73-4957-eafe-0f401ddb30d7" executionInfo={"status": "ok", "timestamp": 1543878756350, "user_tz": 180, "elapsed": 12832, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-AGuXHmDxlt4/AAAAAAAAAAI/AAAAAAAAL4U/UNG5600HVJI/s64/photo.jpg", "userId": "03342558297964642149"}} colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY> "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": "OK"}}, "base_uri": "https://localhost:8080/", "height": 98}
# Uploading files from your local file system
from google.colab import files
uploaded = files.upload()
for fn in uploaded.keys():
print('User uploaded file "{name}" with length {length} bytes'.format(
name=fn, length=len(uploaded[fn])))
# + [markdown] id="molRlwPZPt78" colab_type="text"
# # 1.0 Comparing Frequency Distributions
# + [markdown] id="s8I4cyo2QAxC" colab_type="text"
# ## 1.1 Comparing Frequency Distributions
# + [markdown] id="sFIzYqg_QJdC" colab_type="text"
# In the previous mission, we learned **what graphs we can use to visualize the frequency distribution of any kind of variable**. In this mission, we'll learn about the graphs we can use to **compare** multiple frequency distributions at once.
#
# We'll continue to work with the WNBA data set. Below are the first five rows to help you recollect its structure:
#
# | _ | Name | Team | Pos | Height | Weight | BMI | Birth_Place | Birthdate | Exp_ordinal |
# |---|-----------------|------|-----|--------|--------|-----------|-------------|-------------------|-------------------|
# | 0 | Aerial Powers | DAL | F | 183 | 71.0 | 21.200991 | US | January 17, 1994 | Little experience |
# | 1 | <NAME> | LA | G/F | 185 | 73.0 | 21.329438 | US | May 14, 1982 | Veteran |
# | 2 | <NAME> | CON | G | 170 | 69.0 | 23.875433 | US | October 27, 1990 | Experienced |
# | 3 | <NAME> | SAN | G/F | 185 | 84.0 | 24.543462 | US | December 11, 1988 | Very experienced |
# | 4 | <NAME> | MIN | G | 175 | 78.0 | 25.469388 | US | August 5, 1994 | Rookie |
#
#
# Notice in the table above that we've kept the **Exp_ordinal** variable we created in the previous mission. To remind you, this variable is measured on an **ordinal scale** and describes the level of experience of a player according to the following labeling convention:
#
#
# | Years in WNBA | Label |
# |---------------|-------------------|
# | 0 | Rookie |
# | 1-3 | Little experience |
# | 4-5 | Experienced |
# | 5-10 | Very experienced |
# | >10 | Veteran |
#
#
# + id="zoiBp_j4WkjK" colab_type="code" outputId="6bd0a9f6-fd61-42b5-c39a-ada33eab77fe" executionInfo={"status": "ok", "timestamp": 1543878761245, "user_tz": 180, "elapsed": 665, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-AGuXHmDxlt4/AAAAAAAAAAI/AAAAAAAAL4U/UNG5600HVJI/s64/photo.jpg", "userId": "03342558297964642149"}} colab={"base_uri": "https://localhost:8080/", "height": 121}
import pandas as pd
import matplotlib.pyplot as plt
# read the dataset
wnba = pd.read_csv("wnba.csv")
# cleaning the experience column
wnba.loc[wnba.Experience == 'R',"Experience"] = 0
wnba["Experience"] = wnba["Experience"].astype(int)
# create exp_ordinal column
wnba["Exp_ordinal"] = pd.cut(wnba.Experience,
bins=[-1,0,3,5,10,100],
labels=["rookie","Little experience","Experienced",
"Very experienced","Veteran"])
# verify the results
wnba.Exp_ordinal.value_counts()
# + [markdown] id="GK6qbEZWWjs8" colab_type="text"
#
# Let's say we're interested in analyzing how the distribution of the **Pos** variable (**player position**) varies with the level of experience. In other words, we want to determine, for instance, what are the positions on the court that rookies play most as, and how do rookies compare to veterans with respect to positions on the field.
#
# Here's a series of steps we can take to achieve that:
#
# - Segment the players in the data set by level of experience.
# - For each segment, generate a frequency distribution table for the **Pos** variable.
# - Analyze the frequency distributions comparatively.
#
# In the cell below, we've already done the first step for you and segmented the players in the data set by level of experience. The next two steps are left for you as an exercise.
#
# **Exercise**
#
# <img width="100" src="https://drive.google.com/uc?export=view&id=1E8tR7B9YYUXsU_rddJAyq0FrM0MSelxZ">
#
# - For each segment, generate a frequency distribution table for the **Pos** variable.
# - For the **rookie** segment, assign the frequency distribution table to a variable named **rookie_distro**. tip: use df.value_counts() function.
# - For the **Little experience** segment, assign the table to **little_xp_distro.**
# - For the **Experienced** segment, assign the table to **experienced_distro.**
# - For the **Very experienced** segment, assign the table to **very_xp_distro**.
# - For the **Veteran** segment, assign the table to **veteran_distro.**
# - Print all the tables and analyze them comparatively to determine whether there are any clear patterns in the distribution of player position depending on the level of experience.
# + id="6D5-HZRuWYoV" colab_type="code" outputId="8c252f26-7326-4e96-ae8e-a3f9c808fa6f" executionInfo={"status": "ok", "timestamp": 1543878773407, "user_tz": 180, "elapsed": 703, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-AGuXHmDxlt4/AAAAAAAAAAI/AAAAAAAAL4U/UNG5600HVJI/s64/photo.jpg", "userId": "03342558297964642149"}} colab={"base_uri": "https://localhost:8080/", "height": 357}
rookies = wnba[wnba['Exp_ordinal'] == 'Rookie']
little_xp = wnba[wnba['Exp_ordinal'] == 'Little experience']
experienced = wnba[wnba['Exp_ordinal'] == 'Experienced']
very_xp = wnba[wnba['Exp_ordinal'] == 'Very experienced']
veterans = wnba[wnba['Exp_ordinal'] == 'Veteran']
# put your code here
little_xp_distro = little_xp.Pos.value_counts().plot.pie(figsize=(6,6))
plt.show()
# + [markdown] id="2FCO4tS1YHhj" colab_type="text"
# ## 1.2 Grouped Bar Plots
# + [markdown] id="FUraFtkBYSnJ" colab_type="text"
# The purpose of the previous exercise was to give us a sense about how cumbersome really is to compare multiple distributions at once using frequency tables. Fortunately, we can make the comparison much quicker and more efficiently using graphs.
#
# All the five frequency tables we wanted to compare were for the **Pos** variable, which is measured on a nominal scale. Remember that one kind of graph we can use to visualize the distribution of a nominal variable is a bar plot. A simple solution to our problem is to generate a bar plot for each table, and then group all the bar plots on a single figure.
#
# This is where we'd like to arrive:
#
# <img width="600" src="https://drive.google.com/uc?export=view&id=1xO1mMfvHCMhgglqAI0FrCBE-sE9gt9PZ">
#
#
# Because we grouped all the bar plots together, the graph above is called a **grouped bar plot**. We can generate a grouped bar plot just like the one above using the [seaborn.countplot()](https://seaborn.pydata.org/generated/seaborn.countplot.html) function from the seaborn module, which you might already be familiar with from our visualization lessons. In the code snippet below, we will:
#
# - Import the **seaborn** module with the alias **sns**.
# - Generate the plot with **sns.countplot()**. We'll use the following parameters for this function:
# - **x** — specifies as a string the name of the column we want on the x-axis. We'll place the **Exp_ordinal** column on the x-axis.
# - **hue** — specifies as a string the name of the column we want the bar plots generated for. We want to generate the bar plots for the **Pos** column.
# - **data** - specifies the name of the variable which stores the data set. We stored the data in a variable named **wnba**.
# + id="4v0P7J-NY3kx" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 433} outputId="4b165e0c-2121-4dce-e1e4-eda9e6ca133c" executionInfo={"status": "ok", "timestamp": 1543878786426, "user_tz": 180, "elapsed": 662, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-AGuXHmDxlt4/AAAAAAAAAAI/AAAAAAAAL4U/UNG5600HVJI/s64/photo.jpg", "userId": "03342558297964642149"}}
import seaborn as sns
sns.countplot(x = 'Exp_ordinal', hue = 'Pos', data = wnba)
# + [markdown] id="R8LC7UkfjF6W" colab_type="text"
# Comparing the **five distributions** is now easier, and we can make a couple of observations:
#
# - There's only one **rookie** playing on a combined position **(F/C)**. This is significantly less compared to more experienced players, which suggests that combined positions (**F/C** and **G/F**) may require more complex skills on the field that rookies rarely have.
# - Rookies are the only category where we don't find players on all positions. We can see there are no rookies who play on a G/F position.
# - Guards predominate for every level of experience. This probably means that most players in a basketball team are guards. It's worth examining the distributions of a couple of teams to find whether this is true. If it's true, it might be interesting to find out why teams need so many guards.
#
#
# **Exercise**
#
# <img width="100" src="https://drive.google.com/uc?export=view&id=1E8tR7B9YYUXsU_rddJAyq0FrM0MSelxZ">
#
#
# - Usng **sns.countplot()**, generate a grouped bar plot similar to the one above.
#
# - Place the **Exp_ordinal** variable on the x-axis.
# - Generate the bar plots for the **Pos** variable. The data set is stored in **wnba** variable.
# - Using the **order** parameter of **sns.countplot()**, order the values on the x-axis in **descending** order. The **order** parameter takes in a list of strings, so you should use **order = ['Veteran', 'Very experienced', ..........]**.
# - Using the **hue_order** parameter, order the bars of each bar plot in ascending alphabetic order. **hue_order** takes in a list of strings, so you can use **hue_order = ['C', 'F', ......].**
# + id="S_fKRx8HjzNs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 433} outputId="2dca3a9b-b9ae-439c-f4f1-5d2b746eb085" executionInfo={"status": "ok", "timestamp": 1543878792614, "user_tz": 180, "elapsed": 1016, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-AGuXHmDxlt4/AAAAAAAAAAI/AAAAAAAAL4U/UNG5600HVJI/s64/photo.jpg", "userId": "03342558297964642149"}}
# put your code here
sns.countplot(x='Exp_ordinal', hue='Pos', data=wnba, order=['Veteran','Very experienced','Experienced','Little experience','rookie'],
hue_order=['C','F','F/C','G','G/F'])
# + [markdown] id="4WXoWBidk3e9" colab_type="text"
# ## 1.3 Challenge: Do Older Players Play Less?
#
# + [markdown] id="YoFzcAMEpHe3" colab_type="text"
# When players get past a certain age, they become less and less physically fit as they get older. Intuitively, the fitness level of a player should directly affect how much she plays in a season. On average, a WNBA player played approximately 497 minutes in the 2016-2017 season:
#
# + id="o8V9Ucd3pQMO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="6af5e017-468a-4c9a-d83c-8cceeb364f28" executionInfo={"status": "ok", "timestamp": 1543878795823, "user_tz": 180, "elapsed": 590, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-AGuXHmDxlt4/AAAAAAAAAAI/AAAAAAAAL4U/UNG5600HVJI/s64/photo.jpg", "userId": "03342558297964642149"}}
wnba['MIN'].mean()
# + [markdown] id="Glt-LLCJpSLi" colab_type="text"
# Let's hypothesize that older players generally play less than this average of 497 minutes, while younger players generally play more. As a benchmark to distinguish between younger and older players, we'll take the mean age of players in our sample, which is approximately 27:
# + id="Rl37jGIapau8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="127744b9-26e7-407d-999c-9f818e29822b" executionInfo={"status": "ok", "timestamp": 1543878797880, "user_tz": 180, "elapsed": 603, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-AGuXHmDxlt4/AAAAAAAAAAI/AAAAAAAAL4U/UNG5600HVJI/s64/photo.jpg", "userId": "03342558297964642149"}}
wnba['Age'].mean()
# + [markdown] id="37-K5Vrepghb" colab_type="text"
# To test our hypothesis, we can generate a grouped bar plot to examine the frequency distribution of younger and older players that played under the average or as much as the average or above. Our hypothesis predicts that we should see a grouped bar plot that looks similar to this:
#
# <img width="600" src="https://drive.google.com/uc?export=view&id=11S_m6RQAGChN_iOy7mS1qoEjCOWSEBDV">
#
#
# To generate a graph like the one above, we'll first need to create two new variables:
#
# - An ordinal variable which labels each player as "young" or "old". If the player is 27 or over, we'll label her "old", otherwise the label is "young".
# - An ordinal variable which describes whether the minutes played is below or above average (or equal to the average). If a player played 497 minutes or more, we'll assign her the label "average or above", otherwise we'll assign "below average".
#
# In the code below, we'll use **lambda** functions to describe quickly the labeling logic above and **Series.apply()** to apply the **lambda** functions on the **Age** and **MIN** columns. We'll name the two resulting columns **age_mean_relative** and **min_mean_relative**.
#
# + id="Jf0Bew4Pp7EU" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 206} outputId="ccb07ba6-de9a-4324-a21b-6c6b27568f9b" executionInfo={"status": "ok", "timestamp": 1543878801061, "user_tz": 180, "elapsed": 580, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-AGuXHmDxlt4/AAAAAAAAAAI/AAAAAAAAL4U/UNG5600HVJI/s64/photo.jpg", "userId": "03342558297964642149"}}
wnba['age_mean_relative'] = wnba['Age'].apply(lambda x: 'old' if x >= 27 else 'young')
wnba['min_mean_relative'] = wnba['MIN'].apply(lambda x: 'average or above' if x >= 497 else 'below average')
cols = ["Name","Age","age_mean_relative","MIN","min_mean_relative"]
wnba[cols].head()
# + [markdown] id="ifhDhs76rp_k" colab_type="text"
# **Exercise**
#
# <img width="100" src="https://drive.google.com/uc?export=view&id=1E8tR7B9YYUXsU_rddJAyq0FrM0MSelxZ">
#
#
# - Generate a grouped bar plot to confirm or reject our hypothesis. Using **sns.countplot()**:
# - Place the **age_mean_relative** variable on the x-axis. The **age_mean_relative** and **min_mean_relative** are already defined.
# - Generate the frequency distributions for the **min_mean_relative variable.**
# - Analyze the graph and determine whether the data confirms or rejects our hypothesis. If it's a confirmation assign the string **'confirmation'** to a variable named **result**. If it's a rejection, assign the string **'rejection'** to the variable **result.**
# + id="gesr-86PvZxu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 416} outputId="804bc977-211e-41bc-b0c6-716d68afc69d" executionInfo={"status": "ok", "timestamp": 1543878804165, "user_tz": 180, "elapsed": 626, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-AGuXHmDxlt4/AAAAAAAAAAI/AAAAAAAAL4U/UNG5600HVJI/s64/photo.jpg", "userId": "03342558297964642149"}}
# put your code here
sns.countplot(x='age_mean_relative', hue='min_mean_relative', data=wnba)
result='rejection'
# + [markdown] id="zQNpeRmKvfa7" colab_type="text"
# ## 1.4 Comparing Histograms
# + [markdown] id="3wbt6EU4wBm8" colab_type="text"
# Contrary to what our hypothesis predicted, the grouped bar plot we built showed that among old players the "average or above" category is the most numerous. Among young players we saw an opposite pattern: there are more players who played below the average number of minutes.
#
# A shortcoming of our analysis so far is that the **min_mean_relative** variable doesn't show much granularity. We can see that more **old players** belong to the **"average or above"** category than to **"below average"**, but we can't tell, for instance, whether **old players** generally play much more than the average. For all we know, they could have all played exactly 497 minutes (which is the average).
#
# The **min_mean_relative** variable is ordinal, and it was derived from the **MIN** variable, which is measured on a ratio scale. The information provided by the **MIN** variable is much more granular, and we can plot the distribution of this variable instead. Because the **MIN** variable is measured on a ratio scale, we'll need to use histograms instead of bar plots.
#
# The easiest way to **compare two histograms** is to superimpose one on top of the other. We can do that by using the pandas visualization methods we learned in the previous mission:
# + id="TMnltjWRw7qn" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 347} outputId="d9a1e322-13db-4112-aff6-988535a1b895" executionInfo={"status": "ok", "timestamp": 1543878807709, "user_tz": 180, "elapsed": 576, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-AGuXHmDxlt4/AAAAAAAAAAI/AAAAAAAAL4U/UNG5600HVJI/s64/photo.jpg", "userId": "03342558297964642149"}}
wnba[wnba.Age >= 27]['MIN'].plot.hist(label = 'Old', legend = True)
wnba[wnba.Age < 27]['MIN'].plot.hist(label = 'Young', legend = True)
plt.show()
# + [markdown] id="QnQWaVZGxB1U" colab_type="text"
# We can now see that most of the **old players** that belong to the **"average or above"** category play significantly more than average. The main downside of the visualization above is that the histogram for **young players** covers a large part of the other histogram. We can fix this easily by plotting only the shape of the histograms. We can do this using the **histtype** parameter and choose the **'step'** type:
#
#
# + id="tbee7KFnxnHE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 347} outputId="74cedf8e-7927-4085-f633-13102fd694f2" executionInfo={"status": "ok", "timestamp": 1543878810506, "user_tz": 180, "elapsed": 612, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-AGuXHmDxlt4/AAAAAAAAAAI/AAAAAAAAL4U/UNG5600HVJI/s64/photo.jpg", "userId": "03342558297964642149"}}
sns.set_style("white")
wnba[wnba.Age >= 27]['MIN'].plot.hist(histtype = 'step', label = 'Old', legend = True,color="red")
wnba[wnba.Age < 27]['MIN'].plot.hist(histtype = 'step', label = 'Young', legend = True,color="blue")
plt.show()
# + [markdown] id="1co9KIQKxp8Y" colab_type="text"
# **Exercise**
#
# <img width="100" src="https://drive.google.com/uc?export=view&id=1E8tR7B9YYUXsU_rddJAyq0FrM0MSelxZ">
#
#
# - Looking on our graph above, it's not easy to visualize where the average number of minutes is. Using the [plt.axvline()](https://matplotlib.org/api/_as_gen/matplotlib.pyplot.axvline.html) function, add a vertical line to demarcate the average point:
# - The vertical line should be at point 497 on the x-axis.
# - Use the label parameter of **plt.axvline()** to label it **'Average'**. Display the label by running **plt.legend()**.
# + id="9UD1sMoGyZ-v" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 347} outputId="24466c07-dfbe-4a45-c4ca-87327393ed84" executionInfo={"status": "ok", "timestamp": 1543878815090, "user_tz": 180, "elapsed": 714, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-AGuXHmDxlt4/AAAAAAAAAAI/AAAAAAAAL4U/UNG5600HVJI/s64/photo.jpg", "userId": "03342558297964642149"}}
# put your code here
sns.set_style("white")
wnba[wnba.Age >= 27]['MIN'].plot.hist(histtype = 'step', label = 'Old', legend = True,color="red")
wnba[wnba.Age < 27]['MIN'].plot.hist(histtype = 'step', label = 'Young', legend = True,color="blue")
plt.axvline(497, label='Average')
plt.legend()
plt.show()
# + [markdown] id="qkGRfrn5yhgZ" colab_type="text"
# ## 1.5 Kernel Density Estimate Plots
#
# + [markdown] id="-549n8Jc01yJ" colab_type="text"
# The step-type histograms we built made it possible to see clearly both distributions. The graph looked a bit overcrowded though, and the legend was not ideally positioned.
#
# <img width="500" src="https://drive.google.com/uc?export=view&id=1Oogmu0kyhTgtK-N1zqsdzYGisTM2D6bE">
#
# If we added more histograms to the graph above, it would become highly unreadable, and it'd be difficult to see any clear patterns. One solution to this problem is to smooth out the shape of the histograms to make them look less dense on the graph. This is how a single histogram would look smoothed out:
#
#
# <img width="500" src="https://drive.google.com/uc?export=view&id=1us30ptyKArBL7GvemVRzYvBlGOD3KuMR">
#
# We can smooth out our two histograms above for old and young players using the [Series.plot.kde()](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.Series.plot.kde.html) function:
#
# + id="GsVemyup09BU" colab_type="code" colab={}
wnba[wnba.Age >= 27]['MIN'].plot.kde(label = 'Old', legend = True)
wnba[wnba.Age < 27]['MIN'].plot.kde(label = 'Young', legend = True)
plt.show()
# + [markdown] id="mv7RT9Gc1w04" colab_type="text"
# Each of the smoothed histograms above is called a **kernel density estimate** plot or, shorter, **kernel density plot**. Unlike histograms, **kernel density** plots display densities on the y-axis instead of frequencies. The density values are actually probability values — which we'll be able to understand more about after the probability courses. All you need to know for now is that we can use kernel density plots to get a much clear picture about the shape of a distribution.
#
#
# **Exercise**
#
# <img width="100" src="https://drive.google.com/uc?export=view&id=1E8tR7B9YYUXsU_rddJAyq0FrM0MSelxZ">
#
#
# - Reproduce the kernel density plots above, and add a vertical line to demarcate the average point.
# - The vertical line should be at point 497 on the x-axis.
# - Label the vertical line **'Average'** and make sure the label is displayed in the legend.
# - Can we still see that most of the old players that belong to the **"average or above"** category play significantly more than average? If so, is the pattern more obvious (faster to observe) than in the case of the step-type histograms?
# + id="4aeD2UhB2HXZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 347} outputId="ce74dadc-3f7b-4baa-bca0-c7205f24d17a" executionInfo={"status": "ok", "timestamp": 1543878821257, "user_tz": 180, "elapsed": 765, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-AGuXHmDxlt4/AAAAAAAAAAI/AAAAAAAAL4U/UNG5600HVJI/s64/photo.jpg", "userId": "03342558297964642149"}}
# put your code here
wnba[wnba.Age >= 27]['MIN'].plot.kde(label = 'Old', legend = True)
wnba[wnba.Age < 27]['MIN'].plot.kde(label = 'Young', legend = True)
plt.axvline(497, label='Average', color='red')
plt.legend()
plt.show()
# + [markdown] id="5aQBIQSN2mWU" colab_type="text"
# ## 1.6 Drawbacks of Kernel Density Plots
# + [markdown] id="mw-p3Uh1293i" colab_type="text"
# As data scientists, we'll often need to compare more than two distributions. In fact, previously in this mission we compared five distributions on a grouped bar plot:
#
#
# <img width="400" src="https://drive.google.com/uc?export=view&id=1nSTEDf8EAAE8fQSqxs5dwrlqoU7PF9Gx">
#
# Grouped bar plots are ideal for variables measured on nominal and ordinal scales. For variables measured on a ratio or interval scale, we learned that kernel density plots are a good solution when we have many distributions to compare. However, kernel density plots tend to become unreadable as we reach five distributions or more.
#
# Let's say we're interested in analyzing the distribution of player height as a function of player position. In other words, we want to figure out, for instance, whether centers are generally taller than forwards, whether forwards are generally shorter than guards, and so on. In the code below, we'll segment the data set by player position, and for each segment we'll generate a kernel density plot for the distribution of the **Height** variable:
# + id="-C6bbEnh3EDZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 347} outputId="8a5bbf6a-6995-4d29-fe7f-71ae160eec7f" executionInfo={"status": "ok", "timestamp": 1543878844411, "user_tz": 180, "elapsed": 691, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-AGuXHmDxlt4/AAAAAAAAAAI/AAAAAAAAL4U/UNG5600HVJI/s64/photo.jpg", "userId": "03342558297964642149"}}
wnba[wnba.Pos == 'F']['Height'].plot.kde(label = 'F', legend = True)
wnba[wnba.Pos == 'C']['Height'].plot.kde(label = 'C', legend = True)
wnba[wnba.Pos == 'G']['Height'].plot.kde(label = 'G', legend = True)
wnba[wnba.Pos == 'G/F']['Height'].plot.kde(label = 'G/F', legend = True)
wnba[wnba.Pos == 'F/C']['Height'].plot.kde(label = 'F/C', legend = True)
plt.show()
# + [markdown] id="fdeh9aOX3uqM" colab_type="text"
# If we look very closely, we can see a couple of clear patterns: the shortest players are generally guards, the tallest players are generally centers, mid-height players are generally forwards or play in a combined position, etc.
#
# Having to look very closely to a graph to identify obvious patterns is far from ideal. If there's any pattern, we want to see it immediately. To overcome this problem, we can use other kinds of graphs, which present the same information in a more readable way. For the rest of this mission, we'll explore two such alternatives.
#
# + [markdown] id="5d4hHHgb4Qnm" colab_type="text"
# ## 1.7 Strip Plots
# + [markdown] id="YJWvOLd34Uom" colab_type="text"
# This is one alternative we can use to visualize the distribution of **heights** as a function of **player** position:
#
#
# <img width="400" src="https://drive.google.com/uc?export=view&id=1aQYZGHJg1IDb0C5dUxaSE1coWb3ZI6Dp">
#
#
# The **Pos** variable is represented on the x-axis, while **Height** is on the y-axis. Each of the five vertical lines made of distinctly colored bullets represents a distribution. These are the logical steps we'd take to build a plot like the one above:
#
# - Segment the data set by player position.
# - For every segment:
# - List all the values in the **Height** variable.
# - For every value in that list, draw a bullet point on a graph. The x-coordinate of the bullet point is given by the player position, and the y-coordinate by the player's height.
#
#
# <img width="500" src="https://drive.google.com/uc?export=view&id=1Cp_Pd3uSY-9nE7mPw9oGz788ZPsrMUpJ">
#
#
# Because we segment by player position, for every segment the player position values will be identical for every player while their heights will vary more or less. Because of the segmentation, the player position is also guaranteed to be different from segment to segment. After drawing all the bullet points for all the segments, we'll inevitably end up with five narrow vertical strips, one above each unique value on the x-axis. Because of this, each of the five plots is called a **strip plot**.
#
# To generate the first graph above with five strip plots, we can use the [sns.stripplot()](https://seaborn.pydata.org/generated/seaborn.stripplot.html?highlight=stripplot#seaborn.stripplot) function from the seaborn module. We place the **Pos** variable on the x-axis and **Height** on the y-axis:
#
#
#
#
# + id="vcVqYdrz4YQ1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 361} outputId="052d5b34-0304-4a6c-a2b5-cb2a851635fa" executionInfo={"status": "ok", "timestamp": 1543879157212, "user_tz": 180, "elapsed": 678, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-AGuXHmDxlt4/AAAAAAAAAAI/AAAAAAAAL4U/UNG5600HVJI/s64/photo.jpg", "userId": "03342558297964642149"}}
sns.stripplot(x = 'Pos', y = 'Height', data = wnba)
plt.show()
# + [markdown] id="BPum6AHx5yZ4" colab_type="text"
# Patterns are now immediately visible. We can see on the graph that the **shortest players are guards** — in fact, all players under 180 cm are guards. The **tallest players are centers** — this is the only category with players above 2 meters. Among combined positions, we can see that **F/C has slightly taller representatives** — most likely because it requires center qualities (and we've seen that the tallest players are generally centers).
#
# A **big downside** of strip plots is that the bullet **points overlap**. We can **fix** this by adding a bit of **jitter** to each distribution. We can do this by setting the jitter parameter to **True:**
# + id="S97zw2FS6R44" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 361} outputId="3830413c-acc4-4a6f-9289-095cee15a7ca" executionInfo={"status": "ok", "timestamp": 1543879174199, "user_tz": 180, "elapsed": 697, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-AGuXHmDxlt4/AAAAAAAAAAI/AAAAAAAAL4U/UNG5600HVJI/s64/photo.jpg", "userId": "03342558297964642149"}}
sns.stripplot(x = 'Pos', y = 'Height', data = wnba, jitter = True)
plt.show()
# + [markdown] id="jRG4rpvz6WKo" colab_type="text"
# On a side note, you might have noticed that strip plots are similar to the scatter plots we learned about in the visualization courses. **In fact, strip plots are actually scatter plots.** When one of the variables is nominal or ordinal, a scatter plot will generally take the form of a series of narrow strips (the number of narrow strips will be the same as the number of unique values in the nominal or ordinal variable).
#
# **Exercise**
#
# <img width="100" src="https://drive.google.com/uc?export=view&id=1E8tR7B9YYUXsU_rddJAyq0FrM0MSelxZ">
#
#
# - Using strip plots, examine the distribution of player **weight** (not height) as a function of player **position**. The graph should have the following properties:
# - The **Pos** variable in on the x-axis and the **Weight** variable on the y-axis.
# - Each **strip** plot has **jitter** added to it. The amount of jitter to apply is the one specific to **jitter = True.**
# - Do you see any similarity with the distributions of the **Height** variable? If so, how could this be explained?
# + id="Ai9Nv3Um6p2a" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 361} outputId="81721fa4-26a9-4b03-cbf5-a3fde6bb24c7" executionInfo={"status": "ok", "timestamp": 1543879504463, "user_tz": 180, "elapsed": 642, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-AGuXHmDxlt4/AAAAAAAAAAI/AAAAAAAAL4U/UNG5600HVJI/s64/photo.jpg", "userId": "03342558297964642149"}}
# put your code here
sns.stripplot(x='Pos', y='Weight', data=wnba, jitter=True)
plt.show()
# + [markdown] id="_-16bb-77Omv" colab_type="text"
# ## 1.8 Box plots
# + [markdown] id="tRpQfNku8RP4" colab_type="text"
# Besides strip plots, there's another kind of graph we can use to display many distributions at once and make sure everything is still readable. Below, we use this kind of graph to plot again the distribution of player height as a function of player position:
#
#
# <img width="400" src="https://drive.google.com/uc?export=view&id=1YM9kJ-0f8eMvYEvI1C11TpvwQlqSuyRa">
#
# Each individual plot above shows a distribution. Let's isolate the height distribution of guards and understand it by comparing it with a histogram showing the same distribution:
#
# <img width="800" src="https://drive.google.com/uc?export=view&id=1ubAeLqYPthw2jJpN_ApMc30qM1BjSXpN">
#
# In a nutshell, the graph on the right shows the range of the distribution and its three quartiles (the 25th, the 50th and the 75th percentile). This allows us to get a good visual intuition about the proportion of values that fall under a certain quartile, between any two quartiles, or between a quartile and the minimum or the maximum value in the distribution:
#
#
# <img width="800" src="https://drive.google.com/uc?export=view&id=1Y0H3DLjHVbZOZSzOq8htlinzCxEAQW_R">
#
#
# The two lines extending upwards and downwards out of the box in the middle look a bit like two whiskers, reason for which we call this plot a **box-and-whisker** plot, or, more convenient, just **box plot.**
#
# We can generate the five box plots above using the [sns.boxplot()](https://seaborn.pydata.org/generated/seaborn.boxplot.html) function. On the x-axis we want the **Pos** variable, and on the y-axis the **Height** variable.
#
#
# + id="06YEoa0W8W42" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 396} outputId="f5dfae4d-376e-4fe3-bef0-68ff3a0b795d" executionInfo={"status": "ok", "timestamp": 1543879748061, "user_tz": 180, "elapsed": 677, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-AGuXHmDxlt4/AAAAAAAAAAI/AAAAAAAAL4U/UNG5600HVJI/s64/photo.jpg", "userId": "03342558297964642149"}}
sns.boxplot(x = 'Pos', y = 'Height', data = wnba)
plt.show()
# + [markdown] id="ODb44RO8-Qc9" colab_type="text"
# You might wonder what is the meaning of those few dots for the box plots of centers and guards/forwards (G/F), and **why some box plots seem to lack some of the quartiles**. We'll discuss this in the next screen. Now, let's practice generating box plots.
#
# **Exercise**
#
# <img width="100" src="https://drive.google.com/uc?export=view&id=1E8tR7B9YYUXsU_rddJAyq0FrM0MSelxZ">
#
# - Using **sns.boxplot()**, generate a series of box plots to examine the distribution of player weight as a function of player position. Place the **Pos** variable on the x-axis and the **Weight** variable on the y-axis.
# + id="CV_jILuT-hXh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 396} outputId="9464da08-cf4a-42e3-8ad8-41b4897f137f" executionInfo={"status": "ok", "timestamp": 1543879838165, "user_tz": 180, "elapsed": 923, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-AGuXHmDxlt4/AAAAAAAAAAI/AAAAAAAAL4U/UNG5600HVJI/s64/photo.jpg", "userId": "03342558297964642149"}}
# put your code here
sns.boxplot(x='Pos', y='Weight', data=wnba)
plt.show()
# + [markdown] id="pCXUOoFP-7me" colab_type="text"
# ## 1.9 Outliers
# + [markdown] id="TgQBx0Hn_F7e" colab_type="text"
# The few dots we see for the box plots of centers and guards/forwards (G/F) represent values in the distribution that are much larger or much lower than the rest of the values. A value that is much lower or much larger than the rest of the values in a distribution is called an **outlier.**
#
# <img width="400" src="https://drive.google.com/uc?export=view&id=1WoZ6rZWu8bIFhJaPwQ5mDvoo2pU0dzLa">
#
#
# A value is an **outlier** if:
#
# - It's larger than the upper quartile by 1.5 times the difference between the upper quartile and the lower quartile (the difference is also called the interquartile range).
# - It's lower than the lower quartile by 1.5 times the difference between the upper quartile and the lower quartile (the difference is also called the interquartile range).
#
# <img width="600" src="https://drive.google.com/uc?export=view&id=1NminuWMq8htgOFD5TeAyiLFhXdCtHQjJ">
#
# Probably this is not yet crystal clear, so let's walk through an example. Let's consider the box plot for centers:
#
# <img width="800" src="https://drive.google.com/uc?export=view&id=1krahzq8cce3FQmzXVekQM0UumStTIq8l">
#
#
# From the output of **wnba[wnba['Pos'] == 'C']['Height'].describe()**, we can see that the upper quartile (the 75th percentile) is 196 and the lower quartile (the 25th percentile) is 193. Hence, the interquartile range is 3.
#
#
# $$
# \text{interquartile range} = \text{upper quartile} - \text{lower quartil}
# $$
#
#
# Every value that is $3 \times 1.5$ bigger than the upper quartile is considered an outlier. $3 \times 1.5 = 4.5$, and the upper quartile is 196. This means that any value greater than $196 + 4.5 = 200.5$ is considered an outlier.
#
# Similarly, every value that is $3 \times 1.5$ lower that the lower quartile is an outlier. $3 \times 1.5 = 4.5$, and the upper quartile is 193. This means that any value less than $193 - 4.5 = 188.5$ is an outlier.
#
#
# <img width="500" src="https://drive.google.com/uc?export=view&id=18HtBhxsTPDtuhq4W0YoXfCs8Rx1gz-yQ">
#
#
# This formal definition of an outlier is arbitrary, and it could be changed if we wanted to. For any given distribution, the upper and lower quartiles, and the interquartile range remain constant. However, the 1.5 factor can vary. If the factor is increased, then the range outside which values are considered outliers increases as well. If the factor is decreased, the range outside which values are considered outlier decreases as well.
#
# When we generate boxplots, we can increase or decrease this factor by using the **whis** parameter of the **sns.boxplot()** function. This is the same height distribution for centers without any outliers:
#
#
#
# + id="GEU9c7vH_LUw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 365} outputId="84705787-98e3-4eba-8b7c-4326b7b5ac4c" executionInfo={"status": "ok", "timestamp": 1543880049737, "user_tz": 180, "elapsed": 673, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-AGuXHmDxlt4/AAAAAAAAAAI/AAAAAAAAL4U/UNG5600HVJI/s64/photo.jpg", "userId": "03342558297964642149"}}
sns.boxplot(wnba[wnba['Pos'] == 'C']['Height'], whis = 4,
orient = 'vertical', width = .15,)
plt.show()
# + [markdown] id="M0g8HA1yDpos" colab_type="text"
# **Exercise**
#
# <img width="100" src="https://drive.google.com/uc?export=view&id=1E8tR7B9YYUXsU_rddJAyq0FrM0MSelxZ">
#
# - Consider the quartiles of the **Games Played** variable:
#
# ```python
# >> wnba['Games Played'].describe()
# count 143.000000
# mean 24.356643
# std 7.104259
# min 2.000000
# 25% 22.000000
# 50% 27.000000
# 75% 29.000000
# max 32.000000
# Name: Games Played, dtype: float64
# ```
#
# - Find the interquartile range, and assign the result to a variable named **iqr**.
# - Using a factor of 1.5, calculate the lower and upper bound outside which values are considered outliers.
# - Assign the value of the lower bound to a variable named **lower_bound**.
# - Assign the upper bound to a variable named **upper_bound.**
# - Find how many values in the distribution are outliers.
# - Assign the number of outliers below the lower bound to a variable named **outliers_low.**
# - Assign the number of outliers below the upper bound to a variable named **outliers_high.**
# - Plot a boxplot to check whether your answers are sensible.
#
#
#
# + id="gzs1kjoLE0kX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 396} outputId="f538dd38-7ec2-46dc-8697-31dc35754e68" executionInfo={"status": "ok", "timestamp": 1543882053819, "user_tz": 180, "elapsed": 561, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-AGuXHmDxlt4/AAAAAAAAAAI/AAAAAAAAL4U/UNG5600HVJI/s64/photo.jpg", "userId": "03342558297964642149"}}
# put your code here
iqr = wnba['Games Played'].describe()['75%'] - wnba['Games Played'].describe()['25%']
lower_bound = wnba['Games Played'].describe()['max'] - iqr*1.5
upper_bound = wnba['Games Played'].describe()['max'] + iqr*1.5
outliers_low = lower_bound - wnba['Games Played'].describe()['min']
outliers_high = upper_bound - wnba['Games Played'].describe()['min']
sns.boxplot(x='Pos', y='Games Played', data=wnba)
plt.show()
# + [markdown] id="GfyIiVn0F1A8" colab_type="text"
# ## 1.10 Next Steps
# + [markdown] id="MzxEpz9YGAq-" colab_type="text"
# In this mission, we learned how to compare frequency distributions using graphs. Grouped bar plots are ideal to compare the frequency distributions of nominal or ordinal variables. For variables measured on an interval or ratio scale, we can use step-type histograms, kernel density plots, or, for better readability, strip plots or box plots.
#
# <img width="400" src="https://drive.google.com/uc?export=view&id=1J7n1gvx8sQpJ-WNZF5do8VPQk_vf2ORb">
#
#
# We've come a long way in this course from learning about sampling to visualizing multiple frequency distributions. We've made great progress so far and completed the workflow we set out to do in the first mission.
#
#
# <img width="600" src="https://drive.google.com/uc?export=view&id=1XQ_nPiVB1pMBaS0ikBE6IPeifOYbDG11">
#
| unidade_3/Aula 22/Lesson 22.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # Sequence Models
#
# Imagine that you're watching movies on Netflix. As a good Netflix user you decide to rate each of the movies religiously. After all, a good movie is a good movie, and you want to watch more of them, right? As it turns out, things are not quite so simple. People's opinions on movies can change quite significantly over time. In fact, psychologists even have names for some of the effects:
#
# * There's [anchoring](https://en.wikipedia.org/wiki/Anchoring), based on someone else's opinion. For instance after the Oscar awards, ratings for the corresponding movie go up, even though it's still the same movie. This effect persists for a few months until the award is forgotten. [Wu et al., 2017](https://dl.acm.org/citation.cfm?id=3018689) showed that the effect lifts rating by over half a point.
# * There's the [Hedonic adaptation](https://en.wikipedia.org/wiki/Hedonic_treadmill), where humans quickly adapt to accept an improved (or a bad) situation as the new normal. For instance, after watching many good movies, the expectations that the next movie be equally good or better are high, and even an average movie might be considered a bad movie after many great ones.
# * There's seasonality. Very few viewers like to watch a Santa Claus movie in August.
# * In some cases movies become unpopular due to the misbehaviors of directors or actors in the production.
# * Some movies become cult movies, because they were almost comically bad. *Plan 9 from Outer Space* and *Troll 2* achieved a high degree of notoriety for this reason.
#
# In short, ratings are anything but stationary. Using temporal dynamics helped [<NAME>, 2009](https://dl.acm.org/citation.cfm?id=1557072) to recommend movies more accurately. But it isn't just about movies.
#
# * Many users have highly particular behavior when it comes to the time when they open apps. For instance, social media apps are much more popular after school with students. Stock market trading apps are more commonly used when the markets are open.
# * It is much harder to predict tomorrow's stock prices than to fill in the blanks for a stock price we missed yesterday, even though both are just a matter of estimating one number. After all, hindsight is so much easier than foresight. In statistics the former is called *prediction* whereas the latter is called *filtering*.
# * Music, speech, text, movies, steps, etc. are all sequential in nature. If we were to permute them they would make little sense. The headline *dog bites man* is much less surprising than *man bites dog*, even though the words are identical.
# * Earthquakes are strongly correlated, i.e. after a massive earthquake there are very likely several smaller aftershocks, much more so than without the strong quake. In fact, earthquakes are spatiotemporally correlated, i.e. the aftershocks typically occur within a short time span and in close proximity.
# * Humans interact with each other in a sequential nature, as can be seen in Twitter fights, dance patterns and debates.
#
# ## Statistical Tools
#
# In short, we need statistical tools and new deep networks architectures to deal with sequence data. To keep things simple, we use the stock price as an example.
#
# 
#
# Let's denote the prices by $x_t \geq 0$, i.e. at time $t \in \mathbb{N}$ we observe some price $x_t$. For a trader to do well in the stock market on day $t$ he should want to predict $x_t$ via
#
# $$x_t \sim p(x_t|x_{t-1}, \ldots x_1).$$
#
# ### Autoregressive Models
#
# In order to achieve this, our trader could use a regressor such as the one we trained in the [section on regression](../chapter_deep-learning-basics/linear-regression-gluon.md). There's just a major problem - the number of inputs, $x_{t-1}, \ldots x_1$ varies, depending on $t$. That is, the number increases with the amount of data that we encounter, and we will need an approximation to make this computationally tractable. Much of what follows in this chapter will revolve around how to estimate $p(x_t|x_{t-1}, \ldots x_1)$ efficiently. In a nutshell it boils down to two strategies:
#
# 1. Assume that the potentially rather long sequence $x_{t-1}, \ldots x_1$ isn't really necessary. In this case we might content ourselves with some timespan $\tau$ and only use $x_{t-1}, \ldots x_{t-\tau}$ observations. The immediate benefit is that now the number of arguments is always the same, at least for $t > \tau$. This allows us to train a deep network as indicated above. Such models will be called *autoregressive* models, as they quite literally perform regression on themselves.
# 1. Another strategy is to try and keep some summary $h_t$ of the past observations around and update that in addition to the actual prediction. This leads to models that estimate $x_t|x_{t-1}, h_{t-1}$ and moreover updates of the form $h_t = g(h_t, x_t)$. Since $h_t$ is never observed, these models are also called *latent autoregressive models*. LSTMs and GRUs are examples of this.
#
# Both cases raise the obvious question how to generate training data. One typically uses historical observations to predict the next observation given the ones up to right now. Obviously we do not expect time to stand still. However, a common assumption is that while the specific values of $x_t$ might change, at least the dynamics of the time series itself won't. This is reasonable, since novel dynamics are just that, novel and thus not predictable using data we have so far. Statisticians call dynamics that don't change *stationary*. Regardless of what we do, we will thus get an estimate of the entire time series via
#
# $$p(x_1, \ldots x_T) = \prod_{t=1}^T p(x_t|x_{t-1}, \ldots x_1).$$
#
# Note that the above considerations still hold if we deal with discrete objects, such as words, rather than numbers. The only difference is that in such a situation we need to use a classifier rather than a regressor to estimate $p(x_t| x_{t-1}, \ldots x_1)$.
#
# ### Markov Model
#
# Recall the approximation that in an autoregressive model we use only $(x_{t-1}, \ldots x_{t-\tau})$ instead of $(x_{t-1}, \ldots x_1)$ to estimate $x_t$. Whenever this approximation is accurate we say that the sequence satisfies a Markov condition. In particular, if $\tau = 1$, we have a *first order* Markov model and $p(x)$ is given by
#
# $$p(x_1, \ldots x_T) = \prod_{t=1}^T p(x_t|x_{t-1}).$$
#
# Such models are particularly nice whenever $x_t$ assumes only discrete values, since in this case dynamic programming can be used to compute values along the chain exactly. For instance, we can compute $x_{t+1}|x_{t-1}$ efficiently using the fact that we only need to take into account a very short history of past observations.
#
# $$p(x_{t+1}|x_{t-1}) = \sum_{x_t} p(x_{t+1}|x_t) p(x_t|x_{t-1})$$
#
# Going into details of dynamic programming is beyond the scope of this section. Control and reinforcement learning algorithms use such tools extensively.
#
# ### Causality
#
# In principle, there's nothing wrong with unfolding $p(x_1, \ldots x_T)$ in reverse order. After all, by conditioning we can always write it via
#
# $$p(x_1, \ldots x_T) = \prod_{t=T}^1 p(x_t|x_{t+1}, \ldots x_T).$$
#
# In fact, if we have a Markov model we can obtain a reverse conditional probability distribution, too.
# In many cases, however, there exists a natural direction for the data, namely going forward in time. It is clear that future events cannot influence the past. Hence, if we change $x_t$, we may be able to influence what happens for $x_{t+1}$ going forward but not the converse. That is, if we change $x_t$, the distribution over past events will not change. Consequently, it ought to be easier to explain $x_{t+1}|x_t$ rather than $x_t|x_{t+1}$. For instance, [Hoyer et al., 2008](https://papers.nips.cc/paper/3548-nonlinear-causal-discovery-with-additive-noise-models) show that in some cases we can find $x_{t+1} = f(x_t) + \epsilon$ for some additive noise, whereas the converse is not true. This is great news, since it is typically the forward direction that we're interested in estimating. For more on this topic see e.g. the book by [<NAME>, 2015](https://mitpress.mit.edu/books/elements-causal-inference). We are barely scratching the surface of it.
#
# ## Toy Example
#
# After so much theory, let's try this out in practice. Since much of the modeling is identical to when we built regression estimators in Gluon, we will not delve into much detail regarding the choice of architecture besides the fact that we will use several layers of a fully connected network. Let's begin by generating some data. To keep things simple we generate our 'time series' by using a sine function with some additive noise.
# +
# %matplotlib inline
from IPython import display
from matplotlib import pyplot as plt
from mxnet import autograd, nd, gluon, init
display.set_matplotlib_formats('svg')
embedding = 4 # Embedding dimension for autoregressive model
T = 1000 # Generate a total of 1000 points
time = nd.arange(0,T)
x = nd.sin(0.01 * time) + 0.2 * nd.random.normal(shape=(T))
plt.plot(time.asnumpy(), x.asnumpy());
# -
# Next we need to turn this 'time series' into data the network can train on. Based on the embedding dimension $\tau$ we map the data into pairs $y_t = x_t$ and $\mathbf{z}_t = (x_{t-1}, \ldots x_{t-\tau})$. The astute reader might have noticed that this gives us $\tau$ fewer datapoints, since we don't have sufficient history for the first $\tau$ of them. A simple fix, in particular if the time series is long is to discard those few terms. Alternatively we could pad the time series with zeros. The code below is essentially identical to the training code in previous sections.
# +
features = nd.zeros((T-embedding, embedding))
for i in range(embedding):
features[:,i] = x[i:T-embedding+i]
labels = x[embedding:]
ntrain = 600
train_data = gluon.data.ArrayDataset(features[:ntrain,:], labels[:ntrain])
test_data = gluon.data.ArrayDataset(features[ntrain:,:], labels[ntrain:])
# Vanilla MLP architecture
def get_net():
net = gluon.nn.Sequential()
net.add(gluon.nn.Dense(10, activation='relu'))
net.add(gluon.nn.Dense(10, activation='relu'))
net.add(gluon.nn.Dense(1))
net.initialize(init.Xavier())
return net
# Least mean squares loss
loss = gluon.loss.L2Loss()
# -
# We kept the architecture fairly simple. A few layers of a fully connected network, ReLu activation and $\ell_2$ loss. Now we are ready to train.
# +
# Simple optimizer using adam, random shuffle and minibatch size 16
def train_net(net, data, loss, epochs, learningrate):
batch_size = 16
trainer = gluon.Trainer(net.collect_params(), 'adam',
{'learning_rate': learningrate})
data_iter = gluon.data.DataLoader(data, batch_size, shuffle=True)
for epoch in range(1, epochs + 1):
for X, y in data_iter:
with autograd.record():
l = loss(net(X), y)
l.backward()
trainer.step(batch_size)
l = loss(net(data[:][0]), nd.array(data[:][1]))
print('epoch %d, loss: %f' % (epoch, l.mean().asnumpy()))
return net
net = get_net()
net = train_net(net, train_data, loss, 10, 0.01)
l = loss(net(test_data[:][0]), nd.array(test_data[:][1]))
print('test loss: %f' % l.mean().asnumpy())
# -
# The both training and test loss are small and we would expect our model to work well. Let's see what this means in practice. The first thing to check is how well the model is able to predict what happens in the next timestep.
estimates = net(features)
plt.plot(time.asnumpy(), x.asnumpy(), label='data');
plt.plot(time[embedding:].asnumpy(), estimates.asnumpy(), label='estimate');
plt.legend();
# ## Predictions
#
# This looks nice, just as we expected it. Even beyond 600 observations the estimates still look rather trustworthy. There's just one little problem to this - if we observe data only until time step 600, we cannot hope to receive the ground truth for all future predictions. Instead, we need to work our way forward one step at a time:
#
# $$\begin{aligned}
# x_{601} & = f(x_{600}, \ldots, x_{597}) \\
# x_{602} & = f(x_{601}, \ldots, x_{598}) \\
# x_{603} & = f(x_{602}, \ldots, x_{599})
# \end{aligned}$$
#
# In other words, very quickly will we have to use our own predictions to make future predictions. Let's see how well this goes.
# +
predictions = nd.zeros_like(estimates)
predictions[:(ntrain-embedding)] = estimates[:(ntrain-embedding)]
for i in range(ntrain-embedding, T-embedding):
predictions[i] = net(
predictions[(i-embedding):i].reshape(1,-1)).reshape(1)
plt.plot(time.asnumpy(), x.asnumpy(), label='data');
plt.plot(time[embedding:].asnumpy(), estimates.asnumpy(), label='estimate');
plt.plot(time[embedding:].asnumpy(), predictions.asnumpy(),
label='multistep');
plt.legend();
# -
# As the above example shows, this is a spectacular failure. The estimates decay to 0 pretty quickly after a few prediction steps. Why did the algorithm work so poorly? This is ultimately due to the fact that errors build up. Let's say that after step 1 we have some error $\epsilon_1 = \bar\epsilon$. Now the *input* for step 2 is perturbed by $\epsilon_1$, hence we suffer some error in the order of $\epsilon_2 = \bar\epsilon + L \epsilon_1$, and so on. The error can diverge rather rapidly from the true observations. This is a common phenomenon - for instance weather forecasts for the next 24 hours tend to be pretty accurate but beyond that their accuracy declines rapidly. We will discuss methods for improvig this throughout this chapter and beyond.
#
# Let's verify this observation by computing the $k$-step predictions on the entire sequence.
# +
k = 33 # Look up to k - embedding steps ahead
features = nd.zeros((T-k, k))
for i in range(embedding):
features[:,i] = x[i:T-k+i]
for i in range(embedding, k):
features[:,i] = net(features[:,(i-embedding):i]).reshape((-1))
for i in (4, 8, 16, 32):
plt.plot(time[i:T-k+i].asnumpy(), features[:,i].asnumpy(),
label=('step ' + str(i)))
plt.legend();
# -
# This clearly illustrates how the quality of the estimates changes as we try to predict further into the future. While the 8-step predictions are still pretty good, anything beyond that is pretty useless.
#
# ## Summary
#
# * Sequence models require specialized statistical tools for estimation. Two popular choices are autoregressive models and latent-variable autoregressive models.
# * As we predict further in time, the errors accumulate and the quality of the estimates degrades, often dramatically.
# * There's quite a difference in difficulty between filling in the blanks in a sequence (smoothing) and forecasting. Consequently, if you have a time series, always respect the temporal order of the data when training, i.e. never train on future data.
# * For causal models (e.g. time going forward), estimating the forward direction is typically a lot easier than the reverse direction, i.e. we can get by with simpler networks.
#
# ## Exercises
#
# 1. Improve the above model.
# * Incorporate more than the past 4 observations? How many do you really need?
# * How many would you need if there were no noise? Hint - you can write $\sin$ and $\cos$ as a differential equation.
# * Can you incorporate older features while keeping the total number of features constant? Does this improve accuracy? Why?
# * Change the architecture and see what happens.
# 1. An investor wants to find a good security to buy. She looks at past returns to decide which one is likely to do well. What could possibly go wrong with this strategy?
# 1. Does causality also apply to text? To which extent?
# 1. Give an example for when a latent variable autoregressive model might be needed to capture the dynamic of the data.
#
# ## Scan the QR Code to [Discuss](https://discuss.mxnet.io/t/2860)
#
# 
| book-d2l-en/chapter_recurrent-neural-networks/sequence.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: RL
# language: python
# name: rl
# ---
# +
import os
import random
import math
import time
import numpy as np
import tensorflow as tf
import matplotlib.pyplot as plt
from collections import deque
from tensorflow.keras.optimizers import Adam
from tensorflow.keras import Sequential
from tensorflow.keras.layers import Dense, Conv1D, MaxPooling1D, Flatten, concatenate, Conv2D, MaxPooling2D
import tensorflow.keras.losses as kls
import tensorflow_probability as tfp
from libs.utils import *
from libs.generate_boxes import *
| 3DBPP/ActorCritic/ppo_bpp_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # NumPy
#
# NumPy (or Numpy) is a Linear Algebra Library for Python, the reason it is so important for Data Science with Python is that almost all of the libraries in the PyData Ecosystem rely on NumPy as one of their main building blocks.
#
# Numpy is also incredibly fast, as it has bindings to C libraries. For more info on why you would want to use Arrays instead of lists, check out this great [StackOverflow post](http://stackoverflow.com/questions/993984/why-numpy-instead-of-python-lists).
#
# We will only learn the basics of NumPy, to get started we need to install it!
# ## Installation Instructions
#
# **It is highly recommended you install Python using the Anaconda distribution to make sure all underlying dependencies (such as Linear Algebra libraries) all sync up with the use of a conda install. If you have Anaconda, install NumPy by going to your terminal or command prompt and typing:**
#
# conda install numpy
#
# **If you do not have Anaconda and can not install it, please refer to [Numpy's official documentation on various installation instructions.](http://docs.scipy.org/doc/numpy-1.10.1/user/install.html)**
# ## Using NumPy
#
# Once you've installed NumPy you can import it as a library:
import numpy as np
# Numpy has many built-in functions and capabilities. We won't cover them all but instead we will focus on some of the most important aspects of Numpy: vectors,arrays,matrices, and number generation. Let's start by discussing arrays.
#
# # Numpy Arrays
#
# NumPy arrays are the main way we will use Numpy throughout the course. Numpy arrays essentially come in two flavors: vectors and matrices. Vectors are strictly 1-d arrays and matrices are 2-d (but you should note a matrix can still have only one row or one column).
#
# Let's begin our introduction by exploring how to create NumPy arrays.
#
# ## Creating NumPy Arrays
#
# ### From a Python List
#
# We can create an array by directly converting a list or list of lists:
my_list = [1,2,3] # a normal python list
my_list
np.array(my_list) # casting a normal python 1D list into a vector, deonoted by the sinfle set of square brackets
my_matrix = [[1,2,3],[4,5,6],[7,8,9]] # a normal python list of lists / 2D python list
my_matrix
np.array(my_matrix) # cast a 2D python list into a matrix (2D numpy array), denoted by the 2 sets of square brackets
# # Built-in Methods
#
# There are lots of built-in ways to generate Arrays, more efficiently than the above methods:
# ### ARANGE()
#
# Return evenly spaced values within a given interval. Similar to the python range(start, stop, step)
np.arange(0,9) # Works similar to range()
np.arange(0,11,2) # with a step value
# ### ZEROS() & ONES()
#
# Generate arrays of zeros or ones
np.zeros(5) # Creates a 1D array of zeros i.e 1 row
np.zeros((5,5)) # Creates a 2D array of zeros i.e (5 rows, 5 colums). Note the arg for 2D array must be a tuple
np.ones(3) # Works the same as the zeros method.
np.ones((3,3))
# ### LINSPACE()
# Return a vector of evenly spaced numbers over a specified interval.
np.linspace(0,10,3) # Return 3 evenly spaced numbers between 0 and 10
np.linspace(2,9,100) # Return 100 evenly spaced numbers between 2 and 9
np.linspace(0,1,50) # Return 50 evenly spaced numbers between 0 and 1
# ## EYE()
#
# Creates an identity matrix (a square matrix with every element in the leading diagonal having 1s and all other elements having zeros) with the number of rows and columns equal to the argument passed to it.
np.eye(3)
# ## RANDOM
#
# Numpy also has lots of ways to create random number arrays:
#
# ### RAND()
#
# Creates an array (either vector or matrix) of the given shape and populates it with random samples from a uniform distribution over ``[0, 1]``.
np.random.rand(4) # Create a 4-element vector
np.random.rand(5,5) # Create a 5*5 matrix
# ### RANDN()
#
# Return a sample (or samples) from the "standard normal" distribution. Unlike rand which is uniform:
np.random.randn(2)
### RANF()
np.random.ranf(5)
np.random.randn(5,5)
# ### RANDINT()
# Return random integers from `low` (inclusive) to `high` (exclusive) i.e 'low' has a higher chance of selection if 'high' is not included.
np.random.randint(1,100) # returns one random integer between 1 and 100
np.random.randint(1,100,10) # returns a vector of 10 random integer between 1 and 100
# # Array Attributes and Methods
arr = np.arange(25) # create a vector of 25 elements between 0 - 24
ranarr = np.random.randint(0,50,9) # create a vector of 10 random elements between 0 and 50
arr
ranarr
# ## RESHAPE()
# Returns an array containing the same data with a new shape. Note the number of elements in the input must be enough to exacly build the new shape else a ValueError execption will be raised, NO MORE, NO LESS.
arr.reshape(5,5) # Since arr has 25 elements, these are enough to create a 5*5 matrix
arr.reshape(6,5) # Raises ValueError
arr.reshape(4,4) # Raises ValueError
# ### MAX(), MIN(), ARGMAX(), ARGMIN()
#
# These are useful methods for finding max() or min() values in an array, or to find their index locations using argmin() or argmax()
rarray = ranarr.reshape(3,3)
rarray
rarray.max()
rarray.argmax()
rarray.min()
rarray.argmin()
# ## SHAPE
#
# Shape is an array ATTRIBUTE (not a method), which returns the shape of the array
t_arry = np.random.rand(5,5) # This is 5*5 matrix
t_arry
t_arry.shape # Returns a tuple denoting the dimensions size of the array
t_arry.shape[0] # Access 1st dimension size from shape tuple
t_arry.shape[1] # Access 2nd dimension size from shape tuple
arr.reshape(1,25) # Re-shape above 5*5 matrix into a 1*25 vector
arr.reshape(1,25).shape
arr.reshape(25,1) # Re-shape above 1*25 vector into a 25*1 vector
arr.reshape(25,1).shape
arr.reshape(5,5).shape
# ### DTYPE
#
# You can also grab the data type of the object in the array. This an array ATTRIBUTE.
arr.dtype
arr2 = np.random.rand(5)
arr2.dtype
# # Great Job!
| Python/data_science/data_analysis/02-Python-for-Data-Analysis-NumPy/01-NumPy Arrays.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: tmp
# language: python
# name: tmp
# ---
run -i './ismir2020_make_datasets.py'
run -i './ismir2020_featsubsets_segmentation.py'
run -i './ismir2020_featuresubsets.py'
run -i './ismir2020_vizpgrams.py'
from MTCFeatures import MTCFeatureLoader
import music21 as m21
import pickle
from collections import Counter
# # Data Preparation
# The construction of melody sequences and the extraction of the 5-grams is in notebook ismir2020_extract_grams
# Load the melody sequences
# +
seqs_mtc = MTCFeatureLoader('ismir2020_seqs_mtc_sel.jsonl.gz').sequences()
seqs_mtc = list(seqs_mtc)
seqs_essen = MTCFeatureLoader('ismir2020_seqs_essen_sel.jsonl.gz').sequences()
seqs_essen = list(seqs_essen)
seqs_chor = MTCFeatureLoader('ismir2020_seqs_chor_sel.jsonl.gz').sequences()
seqs_chor = list(seqs_chor)
# -
# Load the pgrams
# +
pgrams_mtc = pd.read_pickle("ismir2020_pgrams_mtc_sel.pkl")
arfftype_mtc = pickle.load( open( "ismir2020_arfftype_mtc_sel.pkl", "rb" ) )
pgrams_essen = pd.read_pickle("ismir2020_pgrams_essen_sel.pkl")
arfftype_essen = pickle.load( open( "ismir2020_arfftype_essen_sel.pkl", "rb" ) )
pgrams_chor = pd.read_pickle("ismir2020_pgrams_chor_sel.pkl")
arfftype_chor = pickle.load( open( "ismir2020_arfftype_chor_sel.pkl", "rb" ) )
# -
# Replace missing values (for random forest classifier)
# +
#replace missing values
#pitchproximity -> 13
pps = [
'pitchproximityfirst',
'pitchproximitysecond',
'pitchproximitythird',
'pitchproximityfourth',
'pitchproximityfifth'
]
pgrams_mtc[pps] = pgrams_mtc[pps].fillna(value=13)
pgrams_essen[pps] = pgrams_essen[pps].fillna(value=13)
pgrams_chor[pps] = pgrams_chor[pps].fillna(value=13)
#pitchreversal -> -2
prs = [
'pitchreversalfirst',
'pitchreversalsecond',
'pitchreversalthird',
'pitchreversalfourth',
'pitchreversalfifth'
]
pgrams_mtc[prs] = pgrams_mtc[prs].fillna(value=-2)
pgrams_essen[prs] = pgrams_essen[prs].fillna(value=-2)
pgrams_chor[prs] = pgrams_chor[prs].fillna(value=-2)
#intervalfirst -> 0
pgrams_mtc[['intervalfirst']] = pgrams_mtc[['intervalfirst']].fillna(value=0)
pgrams_essen[['intervalfirst']] = pgrams_essen[['intervalfirst']].fillna(value=0)
pgrams_chor[['intervalfirst']] = pgrams_chor[['intervalfirst']].fillna(value=0)
#intervalfirst -> 0
pgrams_mtc[['intervalfirst']] = pgrams_mtc[['intervalfirst']].fillna(value=0)
pgrams_essen[['intervalfirst']] = pgrams_essen[['intervalfirst']].fillna(value=0)
pgrams_chor[['intervalfirst']] = pgrams_chor[['intervalfirst']].fillna(value=0)
#derived from intervalfirst -> '='
for featname,replacement in [
('intervalsizefirstsecond','='),
('intervalsizefirstthird','='),
('intervalsizefirstfourth','='),
('intervalsizefirstfifth','='),
('intervaldirfirstsecond','='),
('intervaldirfirstthird','='),
('intervaldirfirstfourth','='),
('intervaldirfirstfifth','='),
('VosCenterGravityfirst',False)
]:
pgrams_mtc[[featname]] = pgrams_mtc[[featname]].fillna(value=replacement)
pgrams_essen[[featname]] = pgrams_essen[[featname]].fillna(value=replacement)
pgrams_chor[[featname]] = pgrams_chor[[featname]].fillna(value=replacement)
#nextisrestthird -> True
pgrams_mtc[['nextisrestthird']] = pgrams_mtc[['nextisrestthird']].fillna(value=True) #n.b. affects all final cadences
pgrams_essen[['nextisrestthird']] = pgrams_essen[['nextisrestthird']].fillna(value=True) #n.b. affects all final cadences
pgrams_chor[['nextisrestthird']] = pgrams_chor[['nextisrestthird']].fillna(value=True) #n.b. affects all final cadences
#localboundary first note -> 0
pgrams_mtc[['lbdmfirst']] = pgrams_mtc[['lbdmfirst']].fillna(value=0.0)
pgrams_essen[['lbdmfirst']] = pgrams_essen[['lbdmfirst']].fillna(value=0.0)
pgrams_chor[['lbdmfirst']] = pgrams_chor[['lbdmfirst']].fillna(value=0.0)
#for the rest just remove the rows with missing values
# -
# Remove all final cadences
# +
#remove all final cadences
pgrams_mtc = pgrams_mtc.loc[pgrams_mtc['cadence_class']!='finalcadence']
arfftype_mtc['cadence_class'] = '{midcadence, nocadence}'
pgrams_essen = pgrams_essen.loc[pgrams_essen['cadence_class']!='finalcadence']
arfftype_essen['cadence_class'] = '{midcadence, nocadence}'
pgrams_chor = pgrams_chor.loc[pgrams_chor['cadence_class']!='finalcadence']
arfftype_chor['cadence_class'] = '{midcadence, nocadence}'
# -
# Selections already have been made:
pgrams_sel_mtc = pgrams_mtc
pgrams_sel_essen = pgrams_essen
pgrams_sel_chor = pgrams_chor
# Write separate arffs for the ismir2020 featsets
# +
writeISMIR2020ARFF = False
if writeISMIR2020ARFF:
for featlist in ismir2020featsets.keys():
toarff = {}
for featname in ismir2020featsets[featlist]:
toarff[featname] = arfftype[featname]
toarff['cadence_class'] = arfftype['cadence_class']
mtcfname = f'ismir2020_arff/ismir2020_pgram_mtc_{featlist}.arff'
essenfname = f'ismir2020_arff/ismir2020_pgram_essen_{featlist}.arff'
chorfname = f'ismir2020_arff/ismir2020_pgram_chor_{featlist}.arff'
pgrams2arff(pgrams_sel_mtc, toarff, mtcfname, classfeat='cadence_class')
if not 'lyr' in featlist and featlist != 'ismir2020_elementaryall':
pgrams2arff(pgrams_sel_essen, toarff, essenfname, classfeat='cadence_class')
pgrams2arff(pgrams_sel_chor, toarff, chorfname, classfeat='cadence_class')
# -
# Prepare the datasets for classifcation
# +
# This is the dataset to work with
# Retain only the features
# keep class labels separate
dataset_mtc_raw = pgrams_sel_mtc
#which feature to use as class label
classfeat = 'cadence_class'
#get class labels
y_mtc_str = dataset_mtc_raw.loc[:,[classfeat]].values.reshape(-1)
#replace all cadence types with the same 'cadence' label
y_mtc_str = ['nocadence' if lab == 'nocadence' else 'cadence' for lab in y_mtc_str]
#select a subset of features
#featset = ismir2020featsets['ismir2020_elementarypitch']
#featset = ismir2020featsets['ismir2020_elementaryrhythm']
#featset = ismir2020featsets['ismir2020_elementarylyrics']
featset = ismir2020featsets['ismir2020_elementarypitchrhythm']
#featset = ismir2020featsets['ismir2020_elementaryall']
#featset = ismir2020featsets['ismir2020_othermodels']
#featset = ismir2020featsets['ismir2020_all']
#featset = ismir2020featsets['ismir2020_all_lyr']
#featset = ismir2020featsets['ismir2020_all_gt']
#featset = ismir2020featsets['ismir2020_all_lyr_gt']
dataset_mtc_raw = dataset_mtc_raw.loc[:,list(featset)]
cnt = Counter(y_mtc_str)
print(str(cnt))
#remove rows with missing values
#after feature selection!
#also remove those from labels y_mtc_str
dataset_mtc_raw.loc[:,'tmp0'] = y_mtc_str
dataset_mtc_raw = dataset_mtc_raw.dropna(axis=0)
y_mtc_str = dataset_mtc_raw.loc[:,['tmp0']].values.reshape(-1)
del dataset_mtc_raw['tmp0']
cnt = Counter(y_mtc_str)
print(str(cnt))
#make onehot features for all nominal features (dtype=object)
#but not for class feature
onehotcolumns = []
#for dt, name in zip(dataset_mtc_raw.dtypes, dataset_mtc_raw.columns):
# if dt=='object' and name != classfeat:
# onehotcolumns.append(name)
for dt, name in zip(dataset_mtc_raw.dtypes, dataset_mtc_raw.columns):
if arfftype_mtc[name]!='numeric' and arfftype_mtc[name]!="{True, False}":
onehotcolumns.append(name)
#These columns have bool as dt:
#if 'grouperfirst' in dataset_mtc_raw.columns: onehotcolumns.append('grouperfirst')
#if 'groupersecond' in dataset_mtc_raw.columns: onehotcolumns.append('groupersecond')
#if 'grouperthird' in dataset_mtc_raw.columns: onehotcolumns.append('grouperthird')
dataset_mtc_raw = pd.get_dummies(dataset_mtc_raw, columns=onehotcolumns)
dataset_mtc = dataset_mtc_raw
# +
# This is the dataset to work with
# Retain only the features
# keep class labels separate
dataset_essen_raw = pgrams_sel_essen
#which feature to use as class label
essen_classfeat = 'cadence_class'
#get class labels
y_essen_str = dataset_essen_raw.loc[:,[classfeat]].values.reshape(-1)
#replace all cadence types with the same 'cadence' label
y_essen_str = ['nocadence' if lab == 'nocadence' else 'cadence' for lab in y_essen_str]
#use same featset as for MTC
essen_featset = featset
dataset_essen_raw = dataset_essen_raw.loc[:,list(essen_featset)]
cnt = Counter(y_essen_str)
print(str(cnt))
#remove rows with missing values
#after feature selection!
#also remove from labels y_essen_str
dataset_essen_raw.loc[:,'tmp0'] = y_essen_str
dataset_essen_raw = dataset_essen_raw.dropna(axis=0)
y_essen_str = dataset_essen_raw.loc[:,['tmp0']].values.reshape(-1)
del dataset_essen_raw['tmp0']
cnt = Counter(y_essen_str)
print(str(cnt))
#make onehot features for all nominal features (dtype=object)
#but not for class feature
onehotcolumns = []
#for dt, name in zip(dataset_essen_raw.dtypes, dataset_essen_raw.columns):
# if dt=='object' and name != classfeat:
# onehotcolumns.append(name)
for dt, name in zip(dataset_essen_raw.dtypes, dataset_essen_raw.columns):
if arfftype_essen[name]!='numeric' and arfftype_mtc[name]!="{True, False}":
onehotcolumns.append(name)
#These columns have bool as dt:
#if 'grouperfirst' in dataset_essen_raw.columns: onehotcolumns.append('grouperfirst')
#if 'groupersecond' in dataset_essen_raw.columns: onehotcolumns.append('groupersecond')
#if 'grouperthird' in dataset_essen_raw.columns: onehotcolumns.append('grouperthird')
dataset_essen_raw = pd.get_dummies(dataset_essen_raw, columns=onehotcolumns)
dataset_essen = dataset_essen_raw
# +
# This is the dataset to work with
# Retain only the features
# keep class labels separate
dataset_chor_raw = pgrams_sel_chor
#which feature to use as class label
chor_classfeat = 'cadence_class'
#get class labels
y_chor_str = dataset_chor_raw.loc[:,[classfeat]].values.reshape(-1)
#replace all cadence types with the same 'cadence' label
if chor_classfeat == 'cadence_class':
y_chor_str = ['nocadence' if lab == 'nocadence' else 'cadence' for lab in y_chor_str]
#use same featset as for MTC
chor_featset = featset
dataset_chor_raw = dataset_chor_raw.loc[:,list(chor_featset)]
cnt = Counter(y_chor_str)
print(str(cnt))
#just remove rows with missing values
#after feature selection!
#also remove from labels y_chor_str
dataset_chor_raw.loc[:,'tmp0'] = y_chor_str
dataset_chor_raw = dataset_chor_raw.dropna(axis=0)
y_chor_str = dataset_chor_raw.loc[:,['tmp0']].values.reshape(-1)
del dataset_chor_raw['tmp0']
cnt = Counter(y_chor_str)
print(str(cnt))
#make onehot features for all nominal features (dtype=object)
#but not for class feature
onehotcolumns = []
#for dt, name in zip(dataset_chor_raw.dtypes, dataset_chor_raw.columns):
# if dt=='object' and name != classfeat:
# onehotcolumns.append(name)
for dt, name in zip(dataset_chor_raw.dtypes, dataset_chor_raw.columns):
if arfftype_chor[name]!='numeric' and arfftype_mtc[name]!="{True, False}":
onehotcolumns.append(name)
#These columns have bool as dt:
#if 'grouperfirst' in dataset_chor_raw.columns: onehotcolumns.append('grouperfirst')
#if 'groupersecond' in dataset_chor_raw.columns: onehotcolumns.append('groupersecond')
#if 'grouperthird' in dataset_chor_raw.columns: onehotcolumns.append('grouperthird')
dataset_chor_raw = pd.get_dummies(dataset_chor_raw, columns=onehotcolumns)
dataset_chor = dataset_chor_raw
# -
from sklearn.model_selection import GroupKFold
from sklearn.model_selection import StratifiedKFold
from sklearn.preprocessing import LabelEncoder
from sklearn.utils import shuffle
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import classification_report
# Prepare the groups for cross validation
# +
songids_mtc = [sid[:12] for sid in dataset_mtc.index]
pgramids_mtc = dataset_mtc.index.values
group_encoder_mtc = LabelEncoder()
groups_mtc = group_encoder_mtc.fit_transform(songids_mtc)
pgramids_essen = dataset_essen.index.values
songids_essen = [essen_id[:-4] for essen_id in pgramids_essen]
group_encoder_essen = LabelEncoder()
groups_essen = group_encoder_essen.fit_transform(songids_essen)
pgramids_chor = dataset_chor.index.values
songids_chor = [chor_id[:-4] for chor_id in pgramids_chor]
group_encoder_chor = LabelEncoder()
groups_chor = group_encoder_chor.fit_transform(songids_chor)
# -
# Prepare X and Y
# +
y_encoder = LabelEncoder()
y_encoder.fit(y_mtc_str)
X_mtc = dataset_mtc.values
y_mtc = y_encoder.transform(y_mtc_str)
X_essen = dataset_essen.values
y_essen = y_encoder.transform(y_essen_str)
X_chor = dataset_chor.values
y_chor = y_encoder.transform(y_chor_str)
# -
# # Classification with Random Forest
# Do classification of MTC
# +
X_mtc_shuffled, y_mtc_shuffled, groups_mtc_shuffled = shuffle(X_mtc, y_mtc, groups_mtc)
group_kfold = GroupKFold(n_splits=5)
y_mtc_shuffled_pred = np.ones(len(y_mtc), dtype=np.int8)
fold=0
for train_index, test_index in group_kfold.split(X_mtc_shuffled, y_mtc_shuffled, groups_mtc_shuffled):
print(f"Fold {fold}")
clf = RandomForestClassifier(n_jobs=12, n_estimators=40)
clf.fit(X_mtc_shuffled[train_index],y_mtc_shuffled[train_index])
y_mtc_shuffled_pred[test_index] = clf.predict(X_mtc_shuffled[test_index])
fold += 1
print(classification_report(y_mtc_shuffled,y_mtc_shuffled_pred))
# -
# Do classification for ESSEN
# +
X_essen_shuffled, y_essen_shuffled, groups_essen_shuffled = shuffle(X_essen, y_essen, groups_essen)
group_kfold = GroupKFold(n_splits=5)
y_essen_shuffled_pred = np.ones(len(y_essen), dtype=np.int8)
fold=0
for train_index, test_index in group_kfold.split(X_essen_shuffled, y_essen_shuffled, groups_essen_shuffled):
print(f"Fold {fold}")
clf = RandomForestClassifier(n_jobs=12, n_estimators=40)
clf.fit(X_essen_shuffled[train_index],y_essen_shuffled[train_index])
y_essen_shuffled_pred[test_index] = clf.predict(X_essen_shuffled[test_index])
fold += 1
print("5-fold CV on Essen:")
print(classification_report(y_essen_shuffled,y_essen_shuffled_pred))
#build classifier for entire dataset
clf_essen = RandomForestClassifier(n_jobs=12, n_estimators=40)
clf_essen.fit(X_essen, y_essen)
if featset == essen_featset:
#build classifier for mtc
clf_mtc = RandomForestClassifier(n_jobs=12, n_estimators=40)
clf_mtc.fit(X_mtc, y_mtc)
y_essen_mtc_pred = clf_mtc.predict(X_essen)
print("Essen classified with MTC training:")
print(classification_report(y_essen,y_essen_mtc_pred))
y_mtc_essen_pred = clf_essen.predict(X_mtc)
print("MTC classified with Essen training:")
print(classification_report(y_mtc, y_mtc_essen_pred))
if essen_featset == chor_featset:
y_chor_essen_pred = clf_essen.predict(X_chor)
print("Chorales classified with Essen training:")
print(classification_report(y_chor, y_chor_essen_pred))
# -
# Do classification for CHOR
# +
X_chor_shuffled, y_chor_shuffled, groups_chor_shuffled = shuffle(X_chor, y_chor, groups_chor)
group_kfold = GroupKFold(n_splits=5)
y_chor_shuffled_pred = np.ones(len(y_chor), dtype=np.int8)
fold=0
for train_index, test_index in group_kfold.split(X_chor_shuffled, y_chor_shuffled, groups_chor_shuffled):
print(f"Fold {fold}")
clf = RandomForestClassifier(n_jobs=12, n_estimators=40)
clf.fit(X_chor_shuffled[train_index],y_chor_shuffled[train_index])
y_chor_shuffled_pred[test_index] = clf.predict(X_chor_shuffled[test_index])
fold += 1
print("5-fold CV on Chor:")
print(classification_report(y_chor_shuffled,y_chor_shuffled_pred))
#build classifier for entire set
clf_chor = RandomForestClassifier(n_jobs=12, n_estimators=40)
clf_chor.fit(X_chor, y_chor)
if featset == chor_featset:
clf_mtc = RandomForestClassifier(n_jobs=12, n_estimators=40)
clf_mtc.fit(X_mtc, y_mtc)
y_chor_mtc_pred = clf_mtc.predict(X_chor)
print("Chor classified with MTC training:")
print(classification_report(y_chor,y_chor_mtc_pred))
y_mtc_chor_pred = clf_chor.predict(X_mtc)
print("MTC classified with Chor training:")
print(classification_report(y_mtc, y_mtc_chor_pred))
if chor_featset == essen_featset:
y_essen_chor_pred = clf_chor.predict(X_essen)
print("Essen classified with Chorale training:")
print(classification_report(y_essen, y_essen_chor_pred))
# -
# # Visualise discovered rules and annotate melodies.
# Functions for converting the rules to pandas queries.
# +
ordvals = [
'-',
'+',
'=',
'start',
'in',
'end',
]
def transformOne(relation):
res = relation.strip(" ()")
for ov in ordvals:
if res.endswith(ov):
res = res[:-len(ov)] + '"' + ov + '"'
res = res.replace(" = "," == ",1)
return res
def JRIP2pandaquery(jrip_query, invert=False):
if jrip_query.find('=>') > 0:
jrip_query = jrip_query[:jrip_query.find('=>')]
jrip_query = jrip_query.split('and')
jrip_query = [transformOne(el) for el in jrip_query]
jrip_query = ' & '.join(jrip_query)
if invert:
jrip_query = '~( ' + jrip_query + ')'
return jrip_query
# -
# Function for pretty printing the rule set.
def pprint_rule(rule):
rule_elms = rule.split(' and ')
print(' and\n '.join(rule_elms))
# The rule sets.
jrip_rules_mtc_pitchrhythm = """(IOIbeatfractionthirdfourth = -) and (completesmeasuresong = True) and (IOIbeatfractionthird >= 1.25) and (meternumerator >= 4) and (IOIbeatfractionfirst <= 0.666667) => cadence_class=midcadence (739.0/54.0)
(IOIbeatfractionthirdfourth = -) and (completesmeasuresong = True) and (IOIbeatfractionthird >= 1) and (IOIbeatfractionsecondthird = +) and (beatstrengthfourth >= 1) => cadence_class=midcadence (705.0/88.0)
(IOIbeatfractionthirdfourth = -) and (completesmeasuresong = True) and (IOIbeatfractionthird >= 1.25) and (IOIbeatfractionfifth <= 1.5) and (VosHarmonyfourth >= 4) and (intervalsecond <= 0) and (diatonicpitchthird <= 30) => cadence_class=midcadence (272.0/15.0)
(IOIbeatfractionthirdfourth = -) and (completesmeasuresong = True) and (beatstrengthfirst <= 0.5) and (IOIbeatfractionthird >= 1.333333) and (meternumerator >= 4) and (beatstrengthsecond <= 0.25) => cadence_class=midcadence (136.0/14.0)
(IOIbeatfractionthirdfourth = -) and (completesmeasuresong = True) and (VosHarmonyfourth >= 4) and (intervalfifth >= 0) and (IOIbeatfractionfifth <= 0.333333) and (midipitchfourth <= 67) and (beatduration >= 1.5) => cadence_class=midcadence (102.0/12.0)
(IOIbeatfractionthirdfourth = -) and (completesmeasuresong = True) and (intervaldirthirdfourth = +) and (intervalfifth >= 0) and (diatonicpitchthirdfourth = =) => cadence_class=midcadence (436.0/92.0)
(IOIbeatfractionthirdfourth = -) and (intervaldirthirdfourth = +) and (IOIbeatfractionthird >= 1.666667) and (completesbeatsong = False) => cadence_class=midcadence (206.0/17.0)
(IOIbeatfractionthirdfourth = -) and (completesmeasuresong = True) and (VosHarmonyfourth >= 4) and (contourthird = -) and (intervalfifth >= 0) and (contourfirst = -) and (VosHarmonythird <= 3) => cadence_class=midcadence (128.0/20.0)
(IOIbeatfractionthirdfourth = -) and (completesmeasuresong = True) and (VosHarmonyfourth >= 4) and (IOIbeatfractionfifth <= 0.5) and (intervalfifth >= 2) and (midipitchfourth <= 68) and (VosHarmonyfirst <= 4) => cadence_class=midcadence (87.0/10.0)
(IOIbeatfractionthirdfourth = -) and (completesmeasuresong = True) and (IOIbeatfractionfifth <= 0.5) and (IOIbeatfractionsecondthird = +) and (intervaldirthirdfourth = +) and (VosCenterGravitysecond = True) => cadence_class=midcadence (113.0/19.0)
(IOIbeatfractionthirdfourth = -) and (completesmeasuresong = True) and (IOIbeatfractionthird >= 1.333333) and (IOIbeatfractionfirst <= 1) and (diatonicpitchthirdfourth = =) => cadence_class=midcadence (154.0/47.0)
(IOIbeatfractionthirdfourth = -) and (completesmeasuresong = True) and (VosHarmonyfourth >= 4) and (beatstrengthfirst <= 0.5) and (IOIbeatfractionsecondthird = +) and (contourfirst = +) => cadence_class=midcadence (124.0/42.0)
(IOIbeatfractionthirdfourth = -) and (IOIbeatfractionthird >= 0.833333) and (beatstrengththird <= 0.5) and (intervalthird <= 0) and (IOIbeatfractionsecondthird = +) and (beatstrengthfourth >= 0.5) => cadence_class=midcadence (293.0/77.0)
(IOIbeatfractionthirdfourth = -) and (IOIbeatfractionthird >= 0.833333) and (intervalthird <= 1) and (beatstrengthfifth >= 1) and (IOIbeatfractionfirst <= 1) and (VosHarmonyfourth >= 6) => cadence_class=midcadence (99.0/18.0)
(IOIbeatfractionthirdfourth = -) and (IOIbeatfractionthird >= 1.666667) and (IOIbeatfractionfifth <= 0.75) and (intervalfourth >= 0) => cadence_class=midcadence (230.0/80.0)
(IOIbeatfractionthirdfourth = -) and (IOIbeatfractionthird >= 0.833333) and (completesmeasuresong = True) and (IOIbeatfractionsecond <= 0.666667) and (contoursecond = -) and (VosHarmonythird <= 2) and (intervalfourth >= 0) => cadence_class=midcadence (37.0/6.0)
(IOIbeatfractionthirdfourth = -) and (IOIbeatfractionthird >= 1.25) and (beatcount <= 2) and (intervalfifth >= 0) and (diatonicpitchfifth <= 32) => cadence_class=midcadence (214.0/65.0)
(IOIbeatfractionthirdfourth = -) and (completesmeasuresong = True) and (VosHarmonyfourth >= 4) and (contourfirst = -) and (intervalfourth <= -4) => cadence_class=midcadence (75.0/29.0)
(IOIbeatfractionthirdfourth = -) and (IOIbeatfractionthird >= 1) and (beatstrengththird <= 0.5) and (intervalthird <= 0) and (onthebeatfifth = True) and (IOIbeatfractionfifth <= 0.666667) => cadence_class=midcadence (138.0/49.0)
(IOIbeatfractionthirdfourth = -) and (completesmeasuresong = True) and (IOIbeatfractionfifth <= 0.666667) and (intervalthird <= 1) and (IOIbeatfractionfirst <= 0.333333) and (beatstrengthfifth >= 1) and (VosHarmonysecond >= 3) => cadence_class=midcadence (78.0/22.0)
(IOIbeatfractionthirdfourth = -) and (IOIbeatfractionthird >= 2.5) and (intervalfourth >= 0) and (IOIbeatfractionsecond <= 2) => cadence_class=midcadence (86.0/14.0)
(IOIbeatfractionthirdfourth = -) and (IOIbeatfractionthird >= 1) and (completesmeasuresong = True) and (intervalthird <= 1) and (diatonicpitchfirst >= 33) and (IOIbeatfractionfirstsecond = +) => cadence_class=midcadence (73.0/21.0)
(IOIbeatfractionthirdfourth = -) and (completesmeasuresong = True) and (IOIbeatfractionfifth <= 0.75) and (intervalthird <= 1) and (VosHarmonyfourth <= 0) and (intervalfirst <= 1) => cadence_class=midcadence (180.0/84.0)
(IOIbeatfractionsecondthird = +) and (VosHarmonyfourth >= 4) and (IOIbeatfractionthird >= 2.5) => cadence_class=midcadence (59.0/17.0)
(IOIbeatfractionthird >= 0.666667) and (completesmeasuresong = True) and (IOIbeatfractionfifth <= 0.75) and (intervaldirthirdfourth = +) and (contoursecond = -) and (intervalfifth >= -1) => cadence_class=midcadence (198.0/79.0)
(completesmeasuresong = True) and (VosHarmonyfourth >= 4) and (diatonicpitchsecondthird = -) and (contourfirst = -) and (midipitchsecond <= 66) and (IOIbeatfractionthirdfourth = =) and (intervalfourth >= 7) => cadence_class=midcadence (46.0/9.0)
(completesmeasuresong = True) and (VosHarmonyfourth >= 4) and (onthebeatthird = True) and (IOIbeatfractionfifth <= 0.75) and (beatcount <= 2) and (VosHarmonyfirst <= 3) and (scaledegreesecond <= 3) => cadence_class=midcadence (187.0/75.0)
(completesmeasuresong = True) and (beatstrengththirdfourth = -) and (IOIbeatfractionfourthfifth = =) and (intervaldirthirdfourth = +) and (contourfirst = -) => cadence_class=midcadence (177.0/81.0)
(IOIbeatfractionthird >= 0.666667) and (completesmeasuresong = True) and (IOIbeatfractionfifth <= 0.75) and (intervalthird <= 0) and (VosHarmonyfourth >= 4) and (midipitchsecond >= 72) => cadence_class=midcadence (118.0/55.0)
(IOIbeatfractionthird >= 1.25) and (beatstrengththird <= 0.5) and (IOIbeatfractionfifth <= 1.5) and (VosHarmonyfourth >= 4) and (VosHarmonyfirst <= 3) => cadence_class=midcadence (114.0/48.0)
(IOIbeatfractionthird >= 0.666667) and (IOIbeatfractionfifth <= 0.75) and (beatstrengththird <= 0.5) and (IOIbeatfractionsecondthird = +) and (intervaldirthirdfourth = +) and (VosHarmonyfirst <= 3) and (intervalsecond <= -2) => cadence_class=midcadence (143.0/67.0)"""
jrip_rules_essen_pitchrhythm = """(completesmeasuresong = True) and (IOIbeatfractionthirdfourth = -) and (contourthird = -) and (beatstrengthfirst <= 0.5) and (IOIbeatfractionthird >= 1.333333) and (IOIbeatfractionfifth <= 1) => cadence_class=midcadence (1284.0/41.0)
(completesmeasuresong = True) and (IOIbeatfractionthirdfourth = -) and (contourthird = -) and (intervalfifth >= 0) and (scaledegreefourth <= 5) and (meternumerator >= 4) and (beatstrengthfifth >= 0.25) => cadence_class=midcadence (316.0/32.0)
(completesmeasuresong = True) and (IOIbeatfractionsecondthird = +) and (IOIbeatfractionfourthfifth = =) and (IOIbeatfractionthird >= 1.5) and (contoursecond = -) => cadence_class=midcadence (446.0/44.0)
(completesmeasuresong = True) and (IOIbeatfractionthirdfourth = -) and (contourthird = -) and (intervaldirthirdfourth = +) and (intervalfifth >= 0) and (beatcount <= 2) and (VosCenterGravitysecond = True) => cadence_class=midcadence (100.0/7.0)
(completesmeasuresong = True) and (IOIbeatfractionsecondthird = +) and (IOIbeatfractionfifth <= 1) and (IOIbeatfractionthird >= 1.333333) and (beatstrengthfourthfifth = -) => cadence_class=midcadence (318.0/36.0)
(completesmeasuresong = True) and (IOIbeatfractionsecondthird = +) and (IOIbeatfractionfirstsecond = =) and (VosHarmonyfourth >= 6) and (intervalfifth >= 4) => cadence_class=midcadence (113.0/14.0)
(completesmeasuresong = True) and (IOIbeatfractionthirdfourth = -) and (intervalfifth >= 0) and (beatstrengthfirst <= 0.5) and (beatstrengthfifth >= 0.5) and (beatstrengthfourth <= 0.25) and (scaledegreefourth <= 2) => cadence_class=midcadence (68.0/5.0)
(completesmeasuresong = True) and (IOIbeatfractionsecondthird = +) and (IOIbeatfractionfifth <= 1) and (IOIbeatfractionthird >= 1.333333) and (beatstrengthfirst <= 0.5) and (beatcount <= 2) => cadence_class=midcadence (90.0/5.0)
(completesmeasuresong = True) and (IOIbeatfractionthirdfourth = -) and (VosHarmonyfourth >= 4) and (contourfirst = -) and (scaledegreefourth <= 1) => cadence_class=midcadence (96.0/11.0)
(completesmeasuresong = True) and (IOIbeatfractionsecondthird = +) and (IOIbeatfractionfourthfifth = =) and (intervaldirthirdfourth = +) and (beatstrengthfifth >= 1) and (VosHarmonysecond >= 2) and (intervalfourth <= 1) => cadence_class=midcadence (53.0/4.0)
(completesmeasuresong = True) and (beatstrengththird >= 0.5) and (IOIbeatfractionfourthfifth = =) and (contourfirst = -) and (intervaldirthirdfourth = +) and (VosCenterGravitythird = True) => cadence_class=midcadence (304.0/67.0)
(completesmeasuresong = True) and (IOIbeatfractionsecondthird = +) and (IOIbeatfractionfifth <= 1) and (contourfirst = -) and (intervalfifth >= -1) and (scaledegreefirst >= 4) and (intervaldirsecondthird = =) => cadence_class=midcadence (133.0/24.0)
(completesmeasuresong = True) and (IOIbeatfractionthirdfourth = -) and (intervalthird <= -2) and (contourfirst = -) and (beatstrengthfirstsecond = +) => cadence_class=midcadence (97.0/15.0)
(completesmeasuresong = True) and (IOIbeatfractionsecondthird = +) and (IOIbeatfractionfifth <= 1) and (VosHarmonyfirst <= 3) and (intervalfourth >= 0) and (VosHarmonyfirst >= 2) => cadence_class=midcadence (577.0/215.0)
(completesmeasuresong = True) and (IOIbeatfractionsecondthird = +) and (IOIbeatfractionfourthfifth = =) and (midipitchthird <= 69) and (VosHarmonyfirst <= 3) and (VosHarmonyfourth >= 5) => cadence_class=midcadence (90.0/22.0)
(completesmeasuresong = True) and (beatstrengththird >= 0.5) and (IOIbeatfractionfourthfifth = =) and (VosHarmonyfourth >= 4) and (beatstrengthfirst <= 0.25) and (IOIbeatfractionsecondthird = -) and (meternumerator >= 4) => cadence_class=midcadence (55.0/1.0)
(completesmeasuresong = True) and (beatstrengththird >= 0.5) and (IOIbeatfractionfourthfifth = =) and (contourthird = -) and (IOIbeatfractionfirstsecond = +) and (beatstrengthfourth <= 0.25) and (IOIbeatfractionthird >= 0.666667) => cadence_class=midcadence (65.0/16.0)
(completesmeasuresong = True) and (beatstrengththird >= 0.5) and (IOIbeatfractionfourthfifth = =) and (VosHarmonyfourth >= 4) and (IOIbeatfractionfirst <= 0.666667) and (contoursecond = -) and (midipitchthird <= 66) and (VosCenterGravityfifth = False) => cadence_class=midcadence (47.0/8.0)
(completesmeasuresong = True) and (beatstrengththird >= 0.5) and (IOIbeatfractionfourthfifth = =) and (contoursecond = -) and (VosHarmonyfourth <= 0) and (scaledegreesecond >= 5) and (beatstrengthfourth <= 0.25) => cadence_class=midcadence (54.0/2.0)
(completesmeasuresong = True) and (IOIbeatfractionthirdfourth = -) and (intervalthird <= 1) and (intervalfifth >= 0) and (intervaldirsecondthird = +) and (IOIbeatfractionfirst <= 0.5) and (scaledegreefirst >= 4) => cadence_class=midcadence (63.0/8.0)
(completesmeasuresong = True) and (beatstrengththird >= 0.5) and (IOIbeatfractionfifth <= 0.75) and (contourthird = -) and (beatstrengthsecond >= 1) and (beatstrengthfourth <= 0.25) and (contourreversal = False) => cadence_class=midcadence (89.0/17.0)
(completesmeasuresong = True) and (IOIbeatfractionthirdfourth = -) and (intervalfifth >= 0) and (VosHarmonyfourth >= 4) and (contourfirst = -) and (IOIbeatfractionfirstsecond = =) and (midipitchsecond <= 69) => cadence_class=midcadence (38.0/6.0)
(completesmeasuresong = True) and (IOIbeatfractionthirdfourth = -) and (intervalfifth >= 0) and (scaledegreesecond >= 3) and (intervaldirthirdfourth = +) and (beatstrengthsecond >= 1) => cadence_class=midcadence (77.0/27.0)
(completesmeasuresong = True) and (beatstrengththird >= 0.5) and (IOIbeatfractionfourthfifth = =) and (VosHarmonyfourth >= 4) and (contourfirst = -) and (intervaldirsecondthird = =) and (beatstrengthfifth >= 1) => cadence_class=midcadence (74.0/22.0)
(completesmeasuresong = True) and (beatstrengththird >= 0.5) and (IOIbeatfractionfifth <= 0.75) and (VosHarmonyfourth >= 4) and (midipitchsecond >= 71) and (beatstrengthfourth >= 0.5) and (scaledegreefifth <= 3) and (diatonicpitchfirst >= 31) => cadence_class=midcadence (52.0/9.0)
(completesmeasuresong = True) and (beatstrengththird >= 0.5) and (IOIbeatfractionfifth <= 0.75) and (beatcount <= 2) and (beatstrengthfirst <= 0.25) and (beatstrengthfourthfifth = +) => cadence_class=midcadence (170.0/54.0)
(completesmeasuresong = True) and (IOIbeatfractionsecondthird = +) and (IOIbeatfractionfifth <= 1.5) and (VosHarmonyfirstsecond = +) and (ambitus <= 1) => cadence_class=midcadence (160.0/59.0)
(completesmeasuresong = True) and (IOIbeatfractionsecondthird = +) and (VosHarmonyfourth >= 4) and (IOIbeatfractionfirstsecond = =) and (beatstrengthfourth >= 1) => cadence_class=midcadence (77.0/29.0)
(completesmeasuresong = True) and (beatstrengththird >= 0.5) and (IOIbeatfractionfourthfifth = =) and (VosHarmonyfirst <= 3) and (diatonicpitchfourth <= 28) and (ambitus <= 1) and (VosHarmonyfirstsecond = +) => cadence_class=midcadence (45.0/13.0)
(completesmeasuresong = True) and (beatstrengththird >= 0.5) and (IOIbeatfractionfourthfifth = =) and (VosHarmonyfirst <= 4) and (diatonicpitchthird <= 30) and (diatonicpitchsecond >= 30) and (beatstrengthsecond <= 0.25) and (VosHarmonyfirstsecond = +) and (diatonicpitchthird >= 30) and (scaledegreefourth >= 3) => cadence_class=midcadence (45.0/8.0)
(completesmeasuresong = True) and (beatstrengththird >= 0.5) and (IOIbeatfractionfourthfifth = =) and (VosHarmonyfirst <= 4) and (VosHarmonyfourth >= 5) and (scaledegreefirst <= 4) and (ambitus <= 2) => cadence_class=midcadence (170.0/67.0)
(completesmeasuresong = True) and (beatstrengththird >= 0.5) and (IOIbeatfractionfifth <= 0.75) and (intervalthird <= -2) and (beatcount <= 2) and (scaledegreethird >= 5) => cadence_class=midcadence (68.0/28.0)
(completesmeasuresong = True) and (IOIbeatfractionthirdfourth = -) and (intervalthird <= -2) and (scaledegreefourth <= 1) => cadence_class=midcadence (77.0/31.0)
(completesmeasuresong = True) and (beatstrengththird >= 0.5) and (VosHarmonyfourth >= 4) and (intervalfourth <= -4) and (diatonicpitchfourth <= 25) and (onthebeatfourth = False) => cadence_class=midcadence (65.0/19.0)
(IOIbeatfractionsecondthird = +) and (IOIbeatfractionthird >= 1.5) and (beatstrengththird <= 0.5) and (intervalfourth >= 0) => cadence_class=midcadence (379.0/122.0)
(completesmeasuresong = True) and (IOIbeatfractionthird >= 0.666667) and (IOIbeatfractionfifth <= 1.5) and (contourfirst = -) and (VosHarmonyfourth >= 4) and (beatstrengthfirst <= 0.25) => cadence_class=midcadence (133.0/47.0)
(completesmeasuresong = True) and (beatstrengththird >= 0.5) and (IOIbeatfractionfourthfifth = =) and (midipitchfirst >= 68) and (scaledegreefourth <= 3) and (scaledegreefourth >= 3) and (VosHarmonyfirst <= 2) => cadence_class=midcadence (126.0/60.0)"""
jrip_rules_chor_pitchrhythm = """(completesmeasuresong = True) and (intervaldirthirdfourth = +) and (IOIbeatfractionfourthfifth = =) and (IOIbeatfractionfirstsecond = +) and (beatstrengththirdfourth = -) => cadence_class=midcadence (257.0/10.0)
(completesmeasuresong = True) and (IOIbeatfractionthird >= 2) and (IOIbeatfractionfourthfifth = =) and (beatstrengthsecond <= 0.25) => cadence_class=midcadence (368.0/4.0)
(completesmeasuresong = True) and (VosHarmonyfourth >= 4) and (contourfourth = +) and (contourfirst = -) => cadence_class=midcadence (255.0/42.0)
(completesmeasuresong = True) and (VosHarmonyfourth <= 0) and (IOIbeatfractionfifth <= 1) and (beatstrengthfourth <= 0.25) => cadence_class=midcadence (320.0/79.0)
(completesmeasuresong = True) and (VosHarmonyfourth >= 4) and (IOIbeatfractionfirstsecond = +) and (diatonicpitchthird <= 30) and (contoursecond = -) => cadence_class=midcadence (47.0/3.0)
(completesmeasuresong = True) and (intervalfourth >= 0) and (beatstrengththird >= 0.5) and (IOIbeatfractionfourthfifth = =) and (VosHarmonyfifth <= 0) and (VosCenterGravityfourth = False) => cadence_class=midcadence (40.0/3.0)
(IOIbeatfractionthird >= 2) and (intervaldirthirdfourth = +) and (beatstrengthfirstsecond = +) and (beatcount >= 4) => cadence_class=midcadence (137.0/5.0)
(IOIbeatfractionthird >= 2) and (intervaldirthirdfourth = +) and (VosHarmonyfourth >= 4) => cadence_class=midcadence (50.0/6.0)
(completesmeasuresong = True) and (VosHarmonyfourth >= 4) and (intervaldirfirstsecond = -) and (intervalthird <= 1) => cadence_class=midcadence (144.0/67.0)
(IOIbeatfractionthird >= 3) and (IOIbeatfractionfourthfifth = =) => cadence_class=midcadence (40.0/4.0)
(IOIbeatfractionthird >= 3) and (intervalfourth >= 0) => cadence_class=midcadence (21.0/6.0)
(completesmeasuresong = True) and (beatstrengththirdfourth = -) and (IOIbeatfractionfirstsecond = +) and (beatstrengthsecond >= 0.5) => cadence_class=midcadence (73.0/20.0)"""
# Chose a rule set and accompanying dataset
# +
jrip_rules = jrip_rules_mtc_pitchrhythm
dataset = pgrams_sel_mtc.copy()
#jrip_rules = jrip_rules_essen_pitchrhythm
#dataset = pgrams_sel_essen.copy()
#jrip_rules = jrip_rules_chor_pitchrhythm
#dataset = pgrams_sel_chor.copy()
# -
# Show the rules
jrip_rules = jrip_rules.split('\n')
for ix, rule in enumerate(jrip_rules):
print(f"Rule {ix}:")
pprint_rule(' '+rule)
# Function to remove all objects that obey to any rule in the rule set, leaving all objects not covered by any rule.
def apply_all_jrip_rules(df, rules):
for rule in rules:
df = df.query(JRIP2pandaquery(rule, invert=True))
return df
# Function to annotate the objects in case a rule applies
#annotates whether a rule applies for an object
#rule -1 means false negative
#NB does annotation IN PLACE
#TODO: Only annotate a rule if current rule == -1 (or solution for now: do it in reverse order)
def annotate_jrip_rules(df, rules, name):
df.loc[:,name] = -1 #erase previous or create new series
for ix, rule in reversed(list(enumerate(rules))):
df.loc[df.eval(JRIP2pandaquery(rule)).values,name] = ix
return df
# Do it
# +
#add the numbers of the rules
annotate_jrip_rules(dataset, jrip_rules, 'rule')
#select all objects not covered by a rule:
notcovered = apply_all_jrip_rules(dataset, jrip_rules)
#select all cadences
cad = dataset.query('cadence_class == "midcadence"')
#cad = notcovered.query('cadence_class == "midcadence"')
# -
# Let's have a look
dataset
# We compute precicion, recall, and F1 (check)
def posneg(c_class, rule):
return ('FP' if (rule!=-1 and c_class == 'nocadence') else #FP
'TP' if (rule!=-1 and c_class != 'nocadence') else #TP
'TN' if (rule==-1 and c_class == 'nocadence') else #TN
'FN' if (rule==-1 and c_class != 'nocadence') else #FN
'UNK')
# +
#to compute FP, FN, TP, TN
#select all cadences and all objects labeled as cadence
hits_misses = dataset.query('rule != -1 | cadence_class != "nocadence"')
#find all errors
errors = [posneg(c_class, rule) for c_class, rule in zip(hits_misses['cadence_class'].values, hits_misses['rule'].values)]
hits_misses.insert(1, 'posneg', errors)
# -
c = Counter(errors)
TP = c['TP']
FP = c['FP']
FN = c['FN']
TN = len(dataset) - TP - FP - FN
total = len(dataset)
prec = TP / (TP + FP)
rec = TP / (TP + FN)
print("TP: ", TP)
print("FP: ", FP)
print("FN: ", FN)
print("TN: ", TN)
print("Total: ", total)
print(prec, rec)
# Store the annotations to disc in a temporary file
hits_misses[['songid','ix0_0','ix2_1','rule','cadence_class']].to_csv('tmp.csv', index=False)
# Any special wishes?
wanted_songids = [
'chor047-sop',
]
# Generate visualisations of rule predictions
viztrigrams(
'tmp.csv',
'ismir2020_mtc_pitchrhythm',
#'ismir2020_essen_pitchrhythm',
#'ismir2020_chor_pitchrhythm',
max_number=50,
random_song_selection=True,
#nlbids=wanted_songids
)
# Do some adjustments and repairs to the generated lilypond files
# +
from pathlib import Path
path = Path('.')
# add staff size
for directory in ['ismir2020_mtc_pitchrhythm','ismir2020_chor_pitchrhythm','ismir2020_essen_pitchrhythm']:
for e in (path/directory).rglob('*.ly'):
with open(e, 'a') as f:
f.write("\n#(set-global-staff-size 14)\n")
f.write("\\layout { indent = 0\\cm}\n")
#For chor music21 puts \autoBeamOff in ly source
#This causes lyrics to shift in case of beamed notes!
#Remove it
for e in (path/'ismir2020_chor_pitchrhythm').rglob('*.ly'):
with open(e, 'r') as f:
lines = f.readlines()
with open(e, 'w') as f:
for line in lines:
if 'autoBeamOff' in line:
line = '% removed \\autoBeamOff\n'
f.write(line)
# -
| ismir2020.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# name: python2
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/google/patents-public-data/blob/master/examples/patent_set_expansion.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="MfeUM-QyOWDN" colab_type="text"
# # Patent Set Expansion
#
# Copyright 2017 Google Inc.
#
# Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
#
# ## Overview
#
# Patent landscaping is an analytical approach commonly used by corporations, patent offices, and academics to better understand the potential technical coverage of a large number of patents where manual review (i.e., actually reading the patents) is not feasible due to time or cost constraints.
#
# Landscaping methods generally fall into one of two categories:
#
# 1. Unsupervised: Given a portfolio of patents about which the user knows no prior information, utilize an unsupervised algorithm to generate topic clusters to provide users a better high-level overview of what that portfolio contains.
# 2. Supervised: Given a seed set of patents about which the user is confident covers a specific technology, identify other patents among a given set that are likely to relate to the same technology.
#
# This notebook creates an example of performing the supervised landscaping method using Python and BigQuery. It performs this by expanding a starting set of patents expected to cover some topic and ranking the results. The methodology overcomes the shortcomings of other landscaping or expansion methods, namely: speed, cost and transparency.
#
# The patent expansion performed here proceeds through the following steps:
#
# 1. Select a seed set of patents.
# 2. Organize and cluster the seed.
# 3. Run searches against the clusters.
# 4. Apply confidence functions to rank the search results.
#
# ## Pre-requisites
# - A google cloud account with access to BigQuery (if you don't have an account yet, this [page](https://cloud.google.com/free/) outlines the process)
# - SQL knowledge
# - Python programming
# - Knowledge of some often used python packages: Numpy, Pandas, sklearn, matplotlib
# + [markdown] id="SMpnczPqQqwB" colab_type="text"
# ## Import Libraries and Authenticate Colab
# + id="q-frA0oZh0Jj" colab_type="code" cellView="both" colab={}
#@markdown Import all the required python libraries and authenticate colab user.
# imports for clustering algorithm
import bisect
import collections
import math
import numpy as np
from sklearn.metrics import silhouette_samples
from sklearn.metrics import silhouette_score
from sklearn.neighbors import NearestNeighbors
# Charting
import matplotlib.pyplot as plt
import seaborn as sns
# General
import pandas as pd
from scipy import spatial
import time
import random
from sklearn import manifold
# BigQuery
from google.cloud import bigquery
from oauth2client.client import GoogleCredentials
from googleapiclient import discovery
# Set BigQuery application credentials
import os
os.environ["GOOGLE_APPLICATION_CREDENTIALS"] = "path_to_file.json"
# + id="tSBGTz-o9XUJ" colab_type="code" cellView="both" colab={}
#@markdown Insert bigquery project id.
bq_project_id = "your_bq_project_id" #@param {type:"string"}
client = bigquery.Client(project=bq_project_id)
# + [markdown] id="bVTXD2sGiLaG" colab_type="text"
# ## Selecting a Seed Set
#
# Performing a landscape search on a set of patents first requires a seed set for the search to be performed against. In order to produce a high-quality search, the input patents should themselves be closely related. The more closely related the input set is, the more confidence you can have in the results. A set of completely random patents will likely yield noisy and more uncertain results.
#
# The input set could span a CPC code, a technology, an assignee, an inventor, etc., or a specific list of patents covering some known technological area. In this walkthrough a term (word) is used to find a seed set. In the [Google Patents Public Datasets](https://console.cloud.google.com/marketplace/details/google_patents_public_datasets/google-patents-public-data), there is a “top terms” field available for all patents in the “google_patents_research.publications” table. The field contains 10 of the most important terms used in a patent. The terms can be unigrams (ex. “aeroelastic”, “genotyping”, “engine”) or bi-grams (ex. “electrical circuit”, “background noise”, “thermal conductivity”).
#
# With an input set selected, you’ll next need a representation of a patent. Rather than using the entire text of a patent or discrete features of a patent, it’s more consumable to use an embedding for each patent. Embeddings are a learned representation of a data input through some type of model, often with a neural network architecture. They reduce the dimensionality of an input set by mapping the most important features of the inputs to a vector of continuous numbers. A benefit of using embeddings is the ability to calculate distances between them, since several distance measures between vectors exist.
#
# You can find a set of patent embeddings on BigQuery in the same Google Patents Public Datasets. The patent embeddings were built using a machine learning model that predicted a patent's CPC code from its text. Therefore the learned embeddings are a vector of 64 continuous numbers intended to encode the information in a patent's text. Distances between the embeddings can then be calculated and used as a measure of similarity between two patents.
#
# In the following example query, we’ve selected a random set of US patents (and collected their embeddings) granted after Jan 1, 2005 with a top term of "neural network".
#
# + id="SrD07oAFnrZb" colab_type="code" cellView="form" outputId="c8b42577-8bd7-4391-a777-211cd0f81d27" colab={"base_uri": "https://localhost:8080/", "height": 210}
search_term = "neural <PASSWORD>" #@param {type:"string"}
return_count = 250 #@param {type:"integer"}
def create_query(search_term):
q = r'''
WITH
pubs as (
SELECT DISTINCT
pub.publication_number
FROM `patents-public-data.patents.publications` pub
INNER JOIN `patents-public-data.google_patents_research.publications` gpr ON
pub.publication_number = gpr.publication_number
WHERE
pub.country_code = 'US'
AND "''' + search_term + '''" IN UNNEST(gpr.top_terms)
AND pub.grant_date >= 20050101 AND pub.grant_date < 20100101
)
SELECT
publication_number, url,
embedding_v1
FROM
`patents-public-data.google_patents_research.publications`
WHERE
publication_number in (SELECT publication_number from pubs)
AND RAND() <= '''+str(return_count)+'''/(SELECT COUNT(*) FROM pubs)
'''
return q
df = client.query(create_query(search_term)).to_dataframe()
if len(df) == 0:
raise ValueError('No results for your search term. Retry with another term.')
else:
print('Search complete for search term: \"{}\". {} random assets selected.'
.format(search_term, len(df)))
embedding_dict = dict(zip(df.publication_number.tolist(),
df.embedding_v1.tolist()))
df.head()
# + [markdown] id="DLQ1drWQiUsI" colab_type="text"
# ## Organizing the Seed Set
#
# With the input set determined and the embedding representations retrieved, you have a few options for determining similarity to the seed set of patents.
#
# Let’s go through each of the options in more detail.
#
# #### Centroid
#
# Calculating an overall embedding point—centroid, medoid, etc.— for the entire input set and performing similarity to that value. Under this method, one metric is calculated to represent the entire input set. That means that the input set of embeddings, which could contain information on hundreds or thousands of patents, ends up pared down to a single point.
#
# There are drawbacks to any methodology that is dependent on one point. If the value itself is not well selected, all results from the search will be poor. Furthermore, even if the point is well selected, the search depends on only that one embedding point, meaning all search results may represent the same area of a topic, technology, etc. By reducing the entire set of inputs to one point, you’ll lose significant information about the input set.
#
# #### Seed Set Size x N
#
# Seed set x N similarity, i.e., calculating similarity to all patents in the input set to all other patents. Doing it this way means you apply the vector distance metric used between each patent in the input set and all other patents in existence. This method presents a few issues:
# - Lack of tractability. Calculating similarity for (seed_set_size x all_patents) is an expensive solution in terms of time and compute.
# - Outliers in the input set are treated as equals to highly representative patents.
# - Dense areas around a single point could be overrepresented in the results.
# - Reusing the input points for similarity may fail to expand the input space.
#
# #### Clustering
#
# Clustering the input set and performing similarity to a cluster. We recommend clustering as the preferred approach to this problem, as it will overcome many of the issues presented by the other two methods. Using clustering, information about the seed set will be condensed into multiple representative points, with no point being an exact replica of its input. With multiple representative points, you can capture various parts of the input technology, features, etc.
# + [markdown] id="Q-2V0IHuTEyY" colab_type="text"
# ## Clustering the Seed
#
# A couple of notes about the embeddings on bigquery:
# - The embeddings are a vector of 64 values, meaning that data is high dimensional.
# - As mentioned previously, the embeddings were trained in a prediction task, not explicitly trained to capture the "distance" between patents.
#
# Based on the embedding training, the clustering algorithm needs to be able to effectively handle clusters of varying density. Since the embeddings were not trained to separate patents evenly, there will be areas of the embedding space that are more or less dense than others, yet represent similar information between documents.
#
# Furthermore, with high-dimensional data, similarity measures can degrade rapidly. One possible approach to overcoming the dimensionality is to use a secondary metric to represent the notion of distance. Rather than using absolute distance values, it’s been shown that a ranking of data points from their distances (and removing the importance of the distance magnitudes) will produce more stable results with higher dimensional data. So our clustering algorithm should remove sole dependence on absolute distance.
#
# It’s also important that a clustering method be able to detect outliers. When providing a large set of input patents, you can expect that not all documents in the set will be reduced to a clear sub-grouping. When the clustering algorithm is unable to group data in a space, it should be capable of ignoring those documents and spaces.
#
# Several algorithms exist that provide these characteristics, any of which can be applied to this problem in place of the algorithm used here. In this application, the shared nearest neighbor (SNN) clustering method is implemented for determining the patent grouping.
#
# SNN is a clustering method that evaluates the neighbors for each point in a dataset and compares the neighbors shared between points to find clusters. SNN is a useful clustering algorithm for determining clusters of varying density. It is good for high-dimensional data, since the explicit distance value is not used in its calculation; rather, it uses a ranking of neighborhood density.
#
# The complete clustering algorithm code is built in the following code block. It is implemented in a manner similar to a scikit-learn model with a fit method.
# + id="0CCX3N0YiWkj" colab_type="code" cellView="both" colab={}
#@markdown Create the shared nearest neighbor clustering algorithm.
"""Implementation of sharest nearest neighbor clustering.
Based on the following algorithm:
http://mlwiki.org/index.php/SNN_Clustering
Following paper:
www.dbs.ifi.lmu.de/~zimek/publications/SSDBM2010/SNN-SSDBM2010-preprint.pdf
Shared Nearest neighbor clustering is a clustering method that evaluates the
neighbors for each point in a dataset and compares those neighbors shared
between points to find clusters. SNN is a useful clustering algorithm for
determining clusters of varying density. It isgood for high dimensional data
since in those spaces the idea of a distance measurement to determine density
is not ideal and with snn distance density is replaced with neighborhood
density.
"""
import bisect
import collections
import math
import numpy as np
from sklearn.metrics import silhouette_samples
from sklearn.metrics import silhouette_score
from sklearn.neighbors import NearestNeighbors
def shared_nearest_neighbor_graph(indices):
"""Create the shared nearest neighbor graph.
Function will create the shared nearest neighbor graph from the nearest
neighbor indices. Function uses the Jarvis-Patrick algorithm specified by the
model.
Args:
indices: A list of lists wrepresenting the nearest neighbor indices of each
point.
Returns:
snn_graph: List representing shared neighbors of each point.
"""
count = len(indices)
snn_graph = []
for _ in range(count):
snn_graph.append([0] * count)
for i, i_neighbors in enumerate(indices):
for j, j_neighbors in enumerate(indices):
if j < i:
continue
if j in i_neighbors and i in j_neighbors:
intersect = len(set(i_neighbors).intersection(j_neighbors))
snn_graph[i][j] = intersect
snn_graph[j][i] = intersect
return snn_graph
def calculate_density(x, eps):
"""Calculate density of a point.
Calculates density of a point based on a required level of epsilon density.
Args:
x: A list of integers representing the shared nearest neighbor counts.
eps: An integer representing the required density of a point.
Returns:
density: An integer representing the density of a point.
"""
# Bisect appears to be fastest, so used it versus others.
density = len(x) - bisect.bisect(sorted(x), eps-1)
return density
def snn_density_graph(shared_nearest_neighbors, eps):
"""Function to iterate through all points in graph and calculate density."""
snn_density = [calculate_density(x, eps) for x in shared_nearest_neighbors]
return snn_density
def find_core_points(snn_density, min_points):
"""Find core points from SNN density list and minimum points requirement."""
return [i for i, density in enumerate(snn_density) if density >= min_points]
def find_core_neighbors(p, core_points_list, shared_nearest_neighbors, eps):
"""Find core point neighbors for a given point.
For a specified point, p, check the SNN density values between p and all other
core points.
Args:
p: A core point represented by an int index value.
core_points_list: List of core points in the dataset.
shared_nearest_neighbors: An SNN graph for a dataset.
eps: The threshold level of density for between points.
Returns:
list: A list of core points which share greater than the epsilon threshold
level of similar neighbors to the provided point p.
"""
return [
core_point for core_point in core_points_list
if shared_nearest_neighbors[p][core_point] >= eps
]
def expand_cluster(labels, neighbor_core, core_points_list, c,
shared_nearest_neighbors, eps, visited):
"""Expand the cluster from the core neighbors.
Function to take the cluster labels (that may be at some intermediate state)
and expand a current set of core point neighbors to additional core point
neighbors and update its cluster label.
Args:
labels: A list of cluster labels for each point in the data.
neighbor_core: A set of core point neighbors.
core_points_list: A list of core points in the data set.
c: An int representing the current cluster label that is being expanded.
shared_nearest_neighbors: A shared nearest neighbor graph.
eps: The threshold level of density for between points.
visited: A set of points already visited for cluster labeling.
Returns:
labels: An updated list of cluster labels for each point in the data.
"""
while neighbor_core:
p = neighbor_core.pop()
if p in visited:
continue
labels[p] = c
visited.add(p)
neighbor_core.update(find_core_neighbors(p, core_points_list,
shared_nearest_neighbors, eps))
return labels
def core_points_from_clusters(core_points_list, shared_nearest_neighbors, eps):
"""Get core points from a cluster.
Function computes the initial cluster labels for the core points in a dataset.
Args:
core_points_list: A list of core points.
shared_nearest_neighbors: A shared nearest neighbor graph.
eps: The threshold level of density for between points.
Returns:
labels: A list of cluster labels for each point in the data.
"""
# Set to hold visited points.
visited = set()
# Cluster label for each point initialized to 0.
labels = [0 for i in range(len(shared_nearest_neighbors))]
# Used to denote the current cluster label.
c = 0
for i in range(len(core_points_list)):
# Skip already visitied points, else add to visited set.
p = core_points_list[i]
if p in visited:
continue
visited.add(p)
# Update cluster label and apply to current point.
c = c + 1
labels[p] = c
# Expand labels from the core neighbors.
neighbor_core = set(find_core_neighbors(p, core_points_list,
shared_nearest_neighbors, eps))
labels = expand_cluster(labels, neighbor_core, core_points_list, c,
shared_nearest_neighbors, eps, visited)
return labels
def label_vote(matched_list):
"""Return most frequently occurring value in list (lowest index if tie)."""
counted = collections.Counter(matched_list)
return max(matched_list, key=counted.get)
def compute_final_labels(labels, core_points, shared_nearest_neighbors, eps):
"""Get the final cluster labels.
Function evaluates the cluster status of non-core data points and tries to
assign them to a cluster label. If no applicable cluster can be found, the
cluster label is left as 0, representing an outlier. Assigning is done by
a "vote" of the label of all core points that have greater than eps value to
the point currently under consideration. We could also implement the label
with the max neighbors instead of voting, though this nearly always ends in
the same result.
Args:
labels: A list of cluster labels for each point in the data.
core_points: List of core points.
shared_nearest_neighbors: A shared nearest neighbor graph.
eps: The threshold level of density for between points.
Returns:
labels: A list of cluster labels for each point in the data.
"""
for i in range(len(labels)):
# If a point is a core point, its cluster has been assigned and we continue.
if i in core_points:
continue
non_core_neighbor_labels = [
labels[k]
for k, j in enumerate(shared_nearest_neighbors[i])
if j >= eps and k in core_points
]
if non_core_neighbor_labels:
updated_label = label_vote(non_core_neighbor_labels)
labels[i] = updated_label
else:
labels[i] = 0
return labels
def calculate_cluster_centers(x, cluster_labels):
"""Calculated center point of each cluster."""
cluster_set = set(cluster_labels)
cluster_set.discard(0) # Remove outliers.
cluster_centers = {}
for cluster in cluster_set:
mask = np.in1d(cluster_labels, cluster)
center = np.mean([x[i] for i in range(len(x)) if mask[i]], axis=0)
cluster_centers[cluster] = list(center)
return cluster_centers
def run_snn(x, metric, n_neighbors, min_points, eps):
"""Run shared nearest neighbor algorithm.
Function takes the input data x and proceeds by running the shared nearest
neighbor algorithm (http://mlwiki.org/index.php/SNN_Clustering). The algorithm
follows these steps:
Step 1 - Find Nearest Neighbors. Nearest neighbor data held in "indices".
Step 2 - Construct SNN graph.
Step 3 - Find the SNN density of each point.
Step 4 - Find the core points.
Step 5a - Find clusters from the core points.
Step 5b - Align non-noise non-core points to clusters.
Step 6 - Calculate cluster centroid.
Step 5, the cluster assignment requires two steps. The first assigns clusters
by determining if two core points are within the eps radius, in which case
they belong to the same cluster. Afer the core points are examined, all points
that are not within a radius of eps of a core point are discarded and labeled
as noise.
Args:
x: Input data, a list of numeric (int or float) lists.
metric: String value of the distance metric requested.
n_neighbors: An integer for the number of neighbors to calculate for each
data point in the dataset.
min_points: Integer for minimum required points to determine a core point.
eps: Float value representing the required neighbor density for forming
clusters.
Returns:
labels: Cluster label for each data point.
cluster_centers: Centroids for each cluster.
indices: K nearest neighbors list for each data point.
shared_nearest_neighbors: Shared nearest neighbor graph.
core_points_list: List of core points.
"""
if not n_neighbors:
# If n_neighbors not set, fall to default values.
n_neighbors = int(math.sqrt(len(x))/2)
min_points = int(n_neighbors/2)
eps = min_points
else:
# Set some default behavior for min_points and eps.
if not min_points:
min_points = int(n_neighbors/2)
if not eps:
eps = min_points
# Step 1.
# Add 1 since NearestNeighbors returns itself as a nearest point.
nbrs = NearestNeighbors(n_neighbors + 1, metric=metric).fit(x)
_, indices = nbrs.kneighbors(x)
# Remove self as similar and convert to list (for speed in graph calc).
indices = indices[:, 1:].tolist() # converting from np array to list
# Step 2.
shared_nearest_neighbors = shared_nearest_neighbor_graph(indices)
# Step 3.
snn_density = snn_density_graph(shared_nearest_neighbors, eps)
# Step 4.
core_points_list = find_core_points(snn_density, min_points)
# Step 5a.
labels_init = core_points_from_clusters(core_points_list,
shared_nearest_neighbors, eps)
# Step 5b.
labels = compute_final_labels(labels_init, core_points_list,
shared_nearest_neighbors, eps)
# Step 6.
cluster_centers = calculate_cluster_centers(x, labels)
return labels, cluster_centers
class SharedNearestNeighbors(object):
"""Shared Nearest Neighbor clustering object."""
def __init__(self, n_neighbors=None, min_points=None, eps=None):
# Attributes
self.labels = None
self.cluster_centers = None
self.neighbor_indices = None
self.snn_graph = None
self.core_points = None
# Parameters
# Keep distance to manhattan or cosine for now.
self.metric = 'cosine' #'manhattan'
self.n_neighbors = n_neighbors
self.min_points = min_points
self.eps = eps
def fit(self, x):
"""Compute the shared nearest neighbor clustering."""
self.labels, self.cluster_centers = run_snn(
x, self.metric, self.n_neighbors, self.min_points, self.eps)
return self
def fit_predict(self, x):
"""Compute the clusters and return predicted cluster labels."""
return self.fit(x).labels
def silhouette(self, x):
"""Find silhouette scores and samples from the input dataset."""
return (silhouette_score(x, self.labels, metric=self.metric),
silhouette_samples(x, self.labels, metric=self.metric))
# + [markdown] id="ObKvpa3w2VHE" colab_type="text"
# For each cluster found through SNN, a representative point for each cluster is found in order to perform a search against it. Two common approaches for representing geometric centers are centroids and medoids. The centroid simply takes the mean value from each of the 64 embedding dimensions. A medoid is the point in a cluster whose average dissimilarity to all objects in a cluster is minimized.
#
# In the next code block we run the clustering and calculate several cluster characteristics, including its centroid. Then we plot a visualization of the clustering result, using TSNE to reduce the dimensions of the inputs.
# + id="QiOxzFyuSROW" colab_type="code" cellView="both" outputId="c63b507d-2b3c-48b4-aeda-dbce8d659612" colab={"base_uri": "https://localhost:8080/", "height": 1000}
#@markdown Run the clustering algorithm, calculate cluster characteristics and visualize.
patents = embedding_dict.keys()
embeddings = embedding_dict.values()
snn = SharedNearestNeighbors()
snn.fit(embeddings)
cluster_labels = snn.labels
cluster_centers = snn.cluster_centers
cluster_lengths = collections.Counter(cluster_labels)
cluster_dict = {}
cluster_set = set(cluster_labels)
# Outliers in clustering will be labeled with 0 so no cluster calculated.
cluster_set.discard(0)
# For each cluster we calculate various characteristics and organize data.
for i in cluster_set:
mask = np.in1d(cluster_labels, i)
masked_embeddings = np.array(embeddings)[mask]
centroid = cluster_centers[i]
cluster_length = cluster_lengths[i]
# Now from cluster center we calculate distance of all belonging points.
centroid_sim = [spatial.distance.cosine(masked_embeddings[j], centroid)
for j in range(len(masked_embeddings))]
cluster_dict[i] = {
'centroid': centroid,
'mean_sim': np.mean(centroid_sim),
'std_sim': np.std(centroid_sim),
'max_sim': np.max(centroid_sim),
'min_sim': np.min(centroid_sim),
'cluster_length': cluster_length,
}
print('Cluster {} \n\tSize: {}. Mean sim: {}. Standard deviation: {}.'.format(
i,
str(cluster_dict[i]['cluster_length']),
str(round(cluster_dict[i]['mean_sim'], 3)),
str(round(cluster_dict[i]['std_sim'], 3)),
))
print('\tMax distance: {}. Min distance: {}.'.format(
str(round(cluster_dict[i]['max_sim'], 3)),
str(round(cluster_dict[i]['min_sim'], 3))
))
print('')
# + id="1qu837oRE8Py" colab_type="code" cellView="both" outputId="0c472ae3-c2a0-493c-eafe-1b803f6d8fa8" colab={"base_uri": "https://localhost:8080/", "height": 575}
#@markdown Cluster Visualization.
palette = {
i: "#"+''.join([random.choice('0123456789ABCDEF') for j in range(6)])
for i in range(len(cluster_set)+1)
}
palette[0] = '#EEEEEE' # Set outlier color to gray
labels_colors = [palette[col] for col in cluster_labels]
n_neighbors = int(math.sqrt(len(embeddings)))
coordinates = manifold.TSNE(n_components=2).fit_transform(embeddings)
with plt.style.context("seaborn-whitegrid"):
fig = plt.figure(figsize=(10,10))
plt.scatter(coordinates.T[0], coordinates.T[1], c=labels_colors, s=75,
edgecolors='black', linewidths=0.4)
plt.xticks([])
plt.yticks([])
plt.show()
# + [markdown] id="Fnbtr9cBiW-6" colab_type="text"
# ## Similarity Search
#
# Once the cluster groups and their centers have been determined, you’ll need a measure of similarity between vectors. Several measures exist, and you can implement any preferred measure. In this example, we used cosine distances to find the similarity between two vectors. Cosine distance is calculated as:
#
# > cosine distance($x$,$y$) $ = 1 - \frac{\sum_{i=1}^n x_i y_i}{\sqrt{\sum_{i=1}^n x_i^2}\sqrt{\sum_{i=1}^n y_i^2}}$
#
# Using the cosine distance, the similarity between a cluster center is compared to all other patents using each of their embeddings. Distance values close to zero mean that the patent is very similar to the cluster point, whereas distances close to one are very far from the cluster point.
#
# Below we iterate through each of the clusters and, using a bigquery udf, calculate the similarity between each centroid and all other patents. We return a dataframe that contains top $n$ results for each similarity search.
#
# + id="wEPGQh8ViYrX" colab_type="code" outputId="eab13556-c20b-4d69-c005-2a0864c4ffa5" cellView="both" colab={"base_uri": "https://localhost:8080/", "height": 465}
#@markdown Run Similarity Search.
similarity_result = pd.DataFrame()
for label, cluster_info in cluster_dict.items():
start = time.time()
query_string = r'''
#standardSQL
CREATE TEMPORARY FUNCTION cosine_distance(patent ARRAY<FLOAT64>)
RETURNS FLOAT64
LANGUAGE js AS """
var cluster_center = [cluster_center];
var dotproduct = 0;
var A = 0;
var B = 0;
for (i = 0; i < patent.length; i++){
dotproduct += (patent[i] * cluster_center[i]);
A += (patent[i]*patent[i]);
B += (cluster_center[i]*cluster_center[i]);
}
A = Math.sqrt(A);
B = Math.sqrt(B);
var cosine_distance = 1 - (dotproduct)/(A)*(B);
return cosine_distance;
""";
CREATE TEMPORARY FUNCTION manhattan_distance(patent ARRAY<FLOAT64>)
RETURNS FLOAT64
LANGUAGE js AS """
var cluster_center = [cluster_center];
var mdist = 0;
for (i = 0; i < patent.length; i++){
mdist += Math.abs(patent[i] - cluster_center[i]);
}
return mdist;
""";
SELECT DISTINCT
[cluster_label] as cluster,
gpr.publication_number,
cosine_distance(gpr.embedding_v1) AS cosine_distance
FROM `patents-public-data.google_patents_research.publications` gpr
WHERE
gpr.country = 'United States' AND
gpr.publication_number not in [cluster_input_list] AND
cosine_distance(gpr.embedding_v1) < [max_distance]
ORDER BY
cosine_distance
LIMIT [max_results]
'''
query_string = query_string.replace('[cluster_center]',
str(cluster_info['centroid']))
query_string = query_string.replace('[cluster_label]', str(label))
# Just set a wide max search distance.
max_distance = cluster_info['mean_sim'] + cluster_info['std_sim'] * 8
query_string = query_string.replace('[max_distance]', str(max_distance))
# Set a max number of results per cluster similarity search.
max_results = cluster_info['cluster_length'] * 20
query_string = query_string.replace('[max_results]', str(max_results))
# Remove input list from results
input_list = str([x.encode('utf-8') for x in df.publication_number.tolist()])
input_list = input_list.replace('[', '(').replace(']', ')')
query_string = query_string.replace('[cluster_input_list]', input_list)
temp = client.query(query_string).to_dataframe()
similarity_result = similarity_result.append(temp, ignore_index=True)
end = time.time()
print('Search cluster {}: {} secs.'.format(str(label), round(end-start, 2)))
# Deduplicate if multiple publications are in a couple of clusters.
agg_dict = {'cosine_distance': 'min'}
temp_similarity = similarity_result.groupby('publication_number',
as_index=False).agg(agg_dict)
similarity_result = pd.merge(similarity_result, temp_similarity, how='inner',
on=['publication_number','cosine_distance'])
print('')
similarity_result.head()
# + [markdown] id="j73TtmMviaUQ" colab_type="text"
# ## Confidence Application
#
# The previous step returns the most similar results to each cluster along with its cosine distance values. From here, the final step takes properties of the cluster and the distance measure from the similarity results to create a confidence level for each result. There are multiple ways to construct a confidence function, and each method may have benefits to certain datasets.
#
# In this walkthrough, we do the confidence scoring using a half squash function. The half squash function is formulated as follows:
#
# >confidence($x$) $ = \frac{x^{power}}{x^{power} + hal\!f^{power}} $
#
# The function takes as input the cosine distance value found between a patent and a cluster center ($x$). Furthermore, the function requires two parameters that affect how the distances of the results are fit onto the confidence scale:
#
# 1. $power$. Defines the properties of the distribution that the distance results are placed onto, effectively the slope of the curve. In this version a power of 2 is used.
# 2. $hal\!f$. Represents the midpoint of the curve returned and defines the saturation on either side of the curve. In this implementation, each cluster uses its own half value.
#
# The half value for each cluster, $i$, is formulated as follows:
#
# >half_value$(i)$ = mean_cluster$(i)$ + ( stddev_cluster$(i)$ x 2 )
#
# >mean_cluster$(i)$ = The mean cosine distance value of all patents in cluster $i$ to the center of cluster $i$
#
# >stddev_cluster$(i)$ = The standard deviation of the cosine distance values of all patents in cluster $i$ to the center of cluster $i$
#
# The confidence scoring function effectively re-saturates the returned distance values to a scale between [0,1] with an exponentially decreasing value as the distance between a patent and the cluster center grows.
# + id="7fCOrRGMib8m" colab_type="code" cellView="form" colab={}
#@markdown Squashing Functions and Half Value Calculations.
# Squash half function
def squash_half(value, half, power):
"""Half squashing function to smooth and compress values."""
if value < 0:
return 0
elif half < 0:
return 1
if power == 1:
return value / (value + half)
value = value / half
value = math.pow(value, power)
return 1 - (value / (value + 1))
# Half value calculations by cluster
halfs = {}
for label, cluster_info in cluster_dict.items():
# If cluster not big, adjust half value scoring.
if cluster_info['cluster_length'] >= 5:
half = cluster_info['mean_sim'] + (cluster_info['std_sim'] * 2)
else:
half = cluster_info['max_sim']
halfs[label] = half
# The half squash power value.
power = 2
# + id="MDkDLFxH8cGP" colab_type="code" cellView="form" colab={}
#@markdown Apply Confidence.
# Confidence application function.
def apply_confidence_to_result(row):
squashed_value = squash_half(row['cosine_distance'],
halfs[row['cluster']],
power)
return int(squashed_value * 100)
similarity_result['confidence'] = similarity_result.apply(
lambda x : apply_confidence_to_result(x), axis=1
)
# + id="PK-HBsMFFnif" colab_type="code" cellView="form" outputId="dacee465-390d-476f-9045-d6780538adb8" colab={"base_uri": "https://localhost:8080/", "height": 455}
#@markdown View Results.
density_data = {}
for i in range(100, 0, -1):
if i == 100:
density_data[i] = len(df)
else:
density_data[i] = len(similarity_result.loc[similarity_result.confidence == i]) + density_data[i+1]
temp = density_data.copy()
min_value = temp[1]
max_value = temp[100]
for i in range(1, 101):
if temp[i] == min_value:
temp.pop(i)
elif temp[i] == max_value:
temp.pop(i)
else:
temp[i] = temp[i] - max_value
y_pos = temp.keys()
performance = temp.values()
with plt.style.context("seaborn-whitegrid"):
plt.figure(figsize=(20,7))
plt.bar(y_pos, performance, align='center', alpha=0.5)
plt.xticks(y_pos, y_pos)
plt.ylabel('Count')
plt.xlabel('Confidence level (%)')
plt.title('Similarity Results')
plt.xticks(rotation=90)
plt.show()
# + [markdown] id="9GWvSC6SXy0C" colab_type="text"
# ## Results and Overview
#
# Applying the confidence function for all of the similarity search results yields a distribution of patents by confidence score across all of the clusters found. At the highest levels of confidence less results will appear. As you move down the confidence distribution the number of results increases exponentially.
#
# Not all results returned are guaranteed to be "hits", however the higher the confidence level the more likely a result is positive. Depending on the input set, the confidence levels will not necessarily begin at 99%. From the results above, using our “neural network” random patent set, the highest confidence results sit in the 60-70% range. From our own experimentation, the more tightly related the input set, the higher the confidence level in the results will be, since the clusters will be more compact.
#
# This walkthrough provides one method for expanding a set of patents using the Google Patents Public Datasets, python and sql. Several changes and adjustments can be made to the queries, cluster algorithm, distance calculations and confidence functions to suit any dataset or experiment.
#
#
#
| examples/patent_set_expansion.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:py35]
# language: python
# name: conda-env-py35-py
# ---
# +
# Read the data with tags
# Plot citations vs time to normalise for time
# Dissolve data by region
# Adjusted citations per paper by region
# Metrics by region:
# (n_papers_ai_tag / n_papers_total) timeseries by (year)
# (n_papers_ai_tag / n_papers_total) 200X vs (n_papers_ai_tag / n_papers_total) 2015 vs [size](change in normalised citations per paper)
# +
#
# -
# %matplotlib inline
import pandas as pd
import seaborn as sns
import numpy as np
from collections import Counter
from matplotlib import pyplot as plt
from mpl_toolkits.basemap import Basemap
import matplotlib.cm as cm
import matplotlib as mpl
# +
# Read the data and apply a is_AI flag
df = pd.read_csv("/Users/hep/Downloads/compsci_stats_with_tag.csv")
df["is_ai"] = df.tags.apply(lambda x : any(tag in x for tag in ['cs.CV','cs.CL','stat.ML','cs.LG']))
# Year by year denomiator
denominator = {}
denom_cites = {}
for year,grouped in df.groupby("year"):
N_ai = grouped.is_ai.sum()
N_total = len(grouped)
denominator[year] = N_ai/N_total
C_ai = grouped.loc[grouped.is_ai].citations.sum()
C_total = grouped.citations.sum()
denom_cites[year] = C_ai/C_total
# Year by year numerator for each institute
data = []
for [inst,year],grouped in df.groupby(["institute","year"]):
if denominator[year] == 0:
continue
n_ai = grouped.is_ai.sum()
n_total = len(grouped)
numerator = n_ai/n_total
rca = numerator/denominator[year]
rca_cites = 0
if denom_cites[year] > 0 and grouped.citations.sum() > 0:
c_ai = grouped.loc[grouped.is_ai].citations.sum()
c_total = grouped.citations.sum()
num_cites = c_ai/c_total
rca_cites = num_cites/denom_cites[year]
lat = grouped.sample().lat.values[0]
lon = grouped.sample().lon.values[0]
data.append(dict(institute=inst,year=year,rca_cites=rca_cites,
rca=rca,lat=lat,lon=lon))
_df = pd.DataFrame(data)
# +
def get_changers(df,yr_min,yr_max):
changes = {}
for inst,grouped in df.groupby("institute"):
# Look at greatest changes in RCA, for RCA > 0 and between yr_min and yr_max
in_range = (grouped.year > yr_min) & (grouped.year <= yr_max) & (grouped.rca > 1)
if in_range.sum() > 1:
_min = grouped.loc[in_range,"rca"].min()
_max = grouped.loc[in_range,"rca"].max()
changes[inst] = np.fabs(_max - _min)
return Counter(changes).most_common()[0:2]
def get_best(df,yr_min,yr_max):
condition = (df.year > yr_min) & (df.year <= yr_max) & (df.rca > 0)
df_sorted = df.loc[condition].sort_values(by="rca",ascending=False)
return df_sorted[["institute","rca"]].head(2).values
changes_before = get_changers(_df,2005,2010)
changes_after = get_changers(_df,2010,2015)
best_before = get_best(_df,2005,2010)
best_after = get_best(_df,2010,2015)
# -
insts = []
insts += [inst for inst,_ in changes_before]
insts += [inst for inst,_ in changes_after]
insts += [inst for inst,_ in best_after]
insts += [inst for inst,_ in best_before]
insts = set(insts)
condition = _df.institute.apply(lambda x : x in insts)
fig,ax = plt.subplots(figsize=(10,5))
for inst,grouped in _df.loc[condition].groupby("institute"):
ax = grouped.plot.line("year","rca",label=inst,ax=ax)
ax.legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.)
fig,ax = plt.subplots(figsize=(10,5))
sns.boxplot(data=_df.loc[(_df.year > 2000) & (_df.year < 2017) & (_df.rca > 0.1)],x="year",y="rca",ax=ax)
# +
condition = (_df.year >= 2000) & (_df.year < 2017) & (_df.rca > 0)
lens = {}
for inst,grouped in _df.loc[condition].groupby("institute"):
lens[inst] = len(grouped)
#indexes = _df.loc[condition].sort_values(by="rca",ascending=False).index
most_common = Counter(lens).most_common()[0:100]
most_common = [inst for inst,_ in most_common]
condition = condition & (_df.institute.apply(lambda x : x in most_common))
fig,ax = plt.subplots(figsize=(10,5))
sns.boxplot(data=_df.loc[condition],x="year",y="rca",ax=ax)
ax.set_ylabel("RCA (top 100 institutes by AI publications)")
ax.set_xlabel("Year")
for year,grouped in _df.loc[condition].groupby("year"):
print(year,max(grouped.rca)-min(grouped.rca),len(grouped))
# +
'''Function to plot lat lon coordinates on a map'''
def make_map(df,ax=None,alpha=0.8,s=2.5,embolden=None,text="",
norm_var=None,cmap=cm.cool,rca_type="rca",**kwargs):
# Convert lat lon to lists
lats = list(df.lat)
lons = list(df.lon)
# Hack to brighten up markers: plot the marker <embolden> times
if embolden:
lats = lats*embolden
lons = lons*embolden
# If an axis hasn't been passed, create one
if ax == None:
fig = plt.figure(figsize=(10,5))
ax_cb = fig.add_axes([0.94, 0.097, 0.03, 0.77])
ax = fig.add_axes([0.05, 0.05, 0.9, 0.9])
#fig,ax = plt.subplots(figsize=(10,5))
# Make the basemap
m = Basemap(projection='merc',llcrnrlat=-60,urcrnrlat=65,
llcrnrlon=-170,urcrnrlon=190,lat_ts=2,resolution='c',)
m.drawcoastlines(linewidth=0.2,color="white")
m.drawcountries(linewidth=0.25,color="white")
m.drawmapboundary(fill_color='black',linewidth=0)
# Convert the coordiates to this projection
lons,lats = m(lons,lats)
# rca color
df[rca_type].loc[df[rca_type] > norm_var] = norm_var
norm = mpl.colors.Normalize(vmin=df[rca_type].min(), vmax=norm_var)
rca = df[rca_type]/norm_var
# Scatter the data
m.scatter(x=lons,y=lats,ax=ax,
latlon=False,alpha=alpha,s=s,
color=cmap(rca),**kwargs)
cb = mpl.colorbar.ColorbarBase(ax_cb, cmap=cmap,norm=norm,orientation='vertical')
# Top title
ax.text(5e5, 17e6, text, color="plum", fontsize=12, weight="bold")
# # Summary of data
# ax.text(5e5, 2.5e6, "Total papers on topic: "+str(condition.sum()), color="white")
# ax.text(5e5, 1.5e6, "Expected: "+str(int(expected))+" ("+sign+str(int(pc_diff))+"%)", color="white")
# # 'Legend'
# ax.text(1.6e7, 3e6, "Institutes involved in the highly cited paper", color="white")
# ax.text(1.6e7, 2e6, "Institutes with new papers on topic", color="white")
# ax.scatter([1.55e7]*100,[3.2e6]*100,color="lime",marker=(5,2))
# ax.scatter([1.55e7]*100,[2.2e6]*100,color="orange",marker="+")
# ax.add_patch(patches.Rectangle((1.5e7,1.1e6),1.5e7,3.1e6,
# fill=False,edgecolor="white"))
fig.savefig(text.replace("/","_")+".png")
return ax
condition = (_df.year >= 2000) & (_df.year < 2017) & (_df.rca > 0)
norm_var = np.percentile(_df.rca,92)
cmap = cm.afmhot
ax_16 = make_map(_df.loc[condition & (_df.year >= 2015)],text="2015/16",norm_var=norm_var,cmap=cmap)
ax_11 = make_map(_df.loc[condition & (_df.year < 2012) & (_df.year >= 2010)],text="2010/11",norm_var=norm_var,cmap=cmap)
# +
condition = (_df.year >= 2000) & (_df.year < 2017) & (_df.rca_cites > 0)
lens = {}
for inst,grouped in _df.loc[condition].groupby("institute"):
lens[inst] = len(grouped)
#indexes = _df.loc[condition].sort_values(by="rca",ascending=False).index
most_common = Counter(lens).most_common()[0:100]
most_common = [inst for inst,_ in most_common]
condition = condition & (_df.institute.apply(lambda x : x in most_common))
fig,ax = plt.subplots(figsize=(10,5))
sns.boxplot(data=_df.loc[condition],x="year",y="rca_cites",ax=ax)
ax.set_ylabel("RCA [citations] (top 100 institutes by AI publications)")
ax.set_xlabel("Year")
# +
condition = (_df.year >= 2000) & (_df.year < 2017) & (_df.rca_cites > 0)
norm_var = np.percentile(_df.rca_cites,95)
cmap = cm.afmhot
make_map(_df.loc[condition & (_df.year >= 2015)],text="2015/16",norm_var=norm_var,cmap=cmap)
make_map(_df.loc[condition & (_df.year < 2012) & (_df.year >= 2010)],text="2010/11",
norm_var=norm_var,cmap=cmap,rca_type="rca_cites")
# -
((_df.year >= 2010) & (_df.year < 2012)).sum()
_df.columns
(4950-3270)/3270
| notebooks/jaklinger/topic_modelling_analysis/rca_boxplot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text"
# # English speaker accent recognition using Transfer Learning
#
# **Author:** [<NAME>](https://twitter.com/fadibadine)<br>
# **Date created:** 2022/04/16<br>
# **Last modified:** 2022/04/16<br>
# **Description:** Training a model to classify UK & Ireland accents using feature extraction from Yamnet.
# + [markdown] colab_type="text"
# ## Introduction
#
# The following example shows how to use feature extraction in order to
# train a model to classify the English accent spoken in an audio wave.
#
# Instead of training a model from scratch, transfer learning enables us to
# take advantage of existing state-of-the-art deep learning models and use them as feature extractors.
#
# Our process:
#
# * Use a TF Hub pre-trained model (Yamnet) and apply it as part of the tf.data pipeline which transforms
# the audio files into feature vectors.
# * Train a dense model on the feature vectors.
# * Use the trained model for inference on a new audio file.
#
# Note:
#
# * We need to install TensorFlow IO in order to resample audio files to 16 kHz as required by Yamnet model.
# * In the test section, ffmpeg is used to convert the mp3 file to wav.
#
# You can install TensorFlow IO with the following command:
# + colab_type="code"
# !pip install -U -q tensorflow_io
# + [markdown] colab_type="text"
# ## Configuration
# + colab_type="code"
SEED = 1337
EPOCHS = 100
BATCH_SIZE = 64
VALIDATION_RATIO = 0.1
MODEL_NAME = "uk_irish_accent_recognition"
# Location where the dataset will be downloaded.
# By default (None), keras.utils.get_file will use ~/.keras/ as the CACHE_DIR
CACHE_DIR = None
# The location of the dataset
URL_PATH = "https://www.openslr.org/resources/83/"
# List of datasets compressed files that contain the audio files
zip_files = {
0: "irish_english_male.zip",
1: "midlands_english_female.zip",
2: "midlands_english_male.zip",
3: "northern_english_female.zip",
4: "northern_english_male.zip",
5: "scottish_english_female.zip",
6: "scottish_english_male.zip",
7: "southern_english_female.zip",
8: "southern_english_male.zip",
9: "welsh_english_female.zip",
10: "welsh_english_male.zip",
}
# We see that there are 2 compressed files for each accent (except Irish):
# - One for male speakers
# - One for female speakers
# However, we will using a gender agnostic dataset.
# List of gender agnostic categories
gender_agnostic_categories = [
"ir", # Irish
"mi", # Midlands
"no", # Northern
"sc", # Scottish
"so", # Southern
"we", # Welsh
]
class_names = [
"Irish",
"Midlands",
"Northern",
"Scottish",
"Southern",
"Welsh",
"Not a speech",
]
# + [markdown] colab_type="text"
# ## Imports
# + colab_type="code"
import os
import io
import csv
import random
import numpy as np
import pandas as pd
import tensorflow as tf
import tensorflow_hub as hub
import tensorflow_io as tfio
from tensorflow import keras
import matplotlib.pyplot as plt
import seaborn as sns
from scipy import stats
from IPython.display import Audio
# Set all random seeds in order to get reproducible results
keras.utils.set_random_seed(SEED)
# Where to download the dataset
DATASET_DESTINATION = os.path.join(CACHE_DIR if CACHE_DIR else "~/.keras/", "datasets")
# + [markdown] colab_type="text"
# ## Yamnet Model
#
# Yamnet is an audio event classifier trained on the AudioSet dataset to predict audio
# events from the AudioSet ontology. It is available on TensorFlow Hub.
#
# Yamnet accepts a 1-D tensor of audio samples with a sample rate of 16 kHz.
# As output, the model returns a 3-tuple:
#
# * Scores of shape `(N, 521)` representing the scores of the 521 classes.
# * Embeddings of shape `(N, 1024)`.
# * The log-mel spectrogram of the entire audio frame.
#
# We will use the embeddings, which are the features extracted from the audio samples, as the input to our dense model.
#
# For more detailed information about Yamnet, please refer to its [TensorFlow Hub](https://tfhub.dev/google/yamnet/1) page.
# + colab_type="code"
yamnet_model = hub.load("https://tfhub.dev/google/yamnet/1")
# + [markdown] colab_type="text"
# ## Dataset
#
# The dataset used is the
# [Crowdsourced high-quality UK and Ireland English Dialect speech data set](https://openslr.org/83/)
# which consists of a total of 17,877 high-quality audio wav files.
#
# This dataset includes over 31 hours of recording from 120 vounteers who self-identify as
# native speakers of Southern England, Midlands, Northern England, Wales, Scotland and Ireland.
#
# For more info, please refer to the above link or to the following paper:
# [Open-source Multi-speaker Corpora of the English Accents in the British Isles](https://aclanthology.org/2020.lrec-1.804.pdf)
# + [markdown] colab_type="text"
# ## Download the data
# + colab_type="code"
# CSV file that contains information about the dataset. For each entry, we have:
# - ID
# - wav file name
# - transcript
line_index_file = keras.utils.get_file(
fname="line_index_file", origin=URL_PATH + "line_index_all.csv"
)
# Download the list of compressed files that contains the audio wav files
for i in zip_files:
fname = zip_files[i].split(".")[0]
url = URL_PATH + zip_files[i]
zip_file = keras.utils.get_file(fname=fname, origin=url, extract=True)
os.remove(zip_file)
# + [markdown] colab_type="text"
# ## Load the data in a Dataframe
#
# Of the 3 columns (ID, filename and transcript), we are only interested in the filename column in order to read the audio file.
# We will ignore the other two.
# + colab_type="code"
dataframe = pd.read_csv(
line_index_file, names=["id", "filename", "transcript"], usecols=["filename"]
)
dataframe.head()
# + [markdown] colab_type="text"
# Let's now preprocess the dataset by:
#
# * Adjusting the filename (removing a leading space & adding ".wav" extension to the
# filename).
# * Creating a label using the first 2 characters of the filename which indicate the
# accent.
# * Shuffling the samples.
# + colab_type="code"
# The purpose of this function is to preprocess the dataframe by applying the following:
# - Cleaning the filename from a leading space
# - Generating a label column that is gender agnostic i.e.
# welsh english male and welsh english female for example are both labeled as
# welsh english
# - Add extension .wav to the filename
# - Shuffle samples
def preprocess_dataframe(dataframe):
# Remove leading space in filename column
dataframe["filename"] = dataframe.apply(lambda row: row["filename"].strip(), axis=1)
# Create gender agnostic labels based on the filename first 2 letters
dataframe["label"] = dataframe.apply(
lambda row: gender_agnostic_categories.index(row["filename"][:2]), axis=1
)
# Add the file path to the name
dataframe["filename"] = dataframe.apply(
lambda row: os.path.join(DATASET_DESTINATION, row["filename"] + ".wav"), axis=1
)
# Shuffle the samples
dataframe = dataframe.sample(frac=1, random_state=SEED).reset_index(drop=True)
return dataframe
dataframe = preprocess_dataframe(dataframe)
dataframe.head()
# + [markdown] colab_type="text"
# ## Prepare training & validation sets
#
# Let's split the samples creating training and validation sets.
# + colab_type="code"
split = int(len(dataframe) * (1 - VALIDATION_RATIO))
train_df = dataframe[:split]
valid_df = dataframe[split:]
print(
f"We have {train_df.shape[0]} training samples & {valid_df.shape[0]} validation ones"
)
# + [markdown] colab_type="text"
# ## Prepare a TensorFlow Dataset
#
# Next, we need to create a `tf.data.Dataset`.
# This is done by creating a `dataframe_to_dataset` function that does the following:
#
# * Create a dataset using filenames and labels.
# * Get the Yamnet embeddings by calling another function `filepath_to_embeddings`.
# * Apply caching, reshuffling and setting batch size.
#
# The `filepath_to_embeddings` does the following:
#
# * Load audio file.
# * Resample audio to 16 kHz.
# * Generate scores and embeddings from Yamnet model.
# * Since Yamnet generates multiple samples for each audio file,
# this function also duplicates the label for all the generated samples
# that have `score=0` (speech) whereas sets the label for the others as
# 'other' indicating that this audio segment is not a speech and we won't label it as one of the accents.
#
# The below `load_16k_audio_file` is copied from the following tutorial
# [Transfer learning with YAMNet for environmental sound classification](https://www.tensorflow.org/tutorials/audio/transfer_learning_audio)
# + colab_type="code"
@tf.function
def load_16k_audio_wav(filename):
# Read file content
file_content = tf.io.read_file(filename)
# Decode audio wave
audio_wav, sample_rate = tf.audio.decode_wav(file_content, desired_channels=1)
audio_wav = tf.squeeze(audio_wav, axis=-1)
sample_rate = tf.cast(sample_rate, dtype=tf.int64)
# Resample to 16k
audio_wav = tfio.audio.resample(audio_wav, rate_in=sample_rate, rate_out=16000)
return audio_wav
def filepath_to_embeddings(filename, label):
# Load 16k audio wave
audio_wav = load_16k_audio_wav(filename)
# Get audio embeddings & scores.
# The embeddings are the audio features extracted using transfer learning
# while scores will be used to identify time slots that are not speech
# which will then be gathered into a specific new category 'other'
scores, embeddings, _ = yamnet_model(audio_wav)
# Number of embeddings in order to know how many times to repeat the label
embeddings_num = tf.shape(embeddings)[0]
labels = tf.repeat(label, embeddings_num)
# Change labels for time-slots that are not speech into a new category 'other'
labels = tf.where(tf.argmax(scores, axis=1) == 0, label, len(class_names) - 1)
# Using one-hot in order to use AUC
return (embeddings, tf.one_hot(labels, len(class_names)))
def dataframe_to_dataset(dataframe, batch_size=64):
dataset = tf.data.Dataset.from_tensor_slices(
(dataframe["filename"], dataframe["label"])
)
dataset = dataset.map(
lambda x, y: filepath_to_embeddings(x, y),
num_parallel_calls=tf.data.experimental.AUTOTUNE,
).unbatch()
return dataset.cache().batch(batch_size).prefetch(tf.data.AUTOTUNE)
train_ds = dataframe_to_dataset(train_df)
valid_ds = dataframe_to_dataset(valid_df)
# + [markdown] colab_type="text"
# ## Build the model
#
# The model that we use consists of:
#
# * An input layer which is the embedding output of the Yamnet classifier.
# * 4 dense hidden layers and 4 dropout layers.
# * An output dense layer.
#
# The model's hyperparameters were selected using
# [KerasTuner](https://keras.io/keras_tuner/).
# + colab_type="code"
keras.backend.clear_session()
def build_and_compile_model():
inputs = keras.layers.Input(shape=(1024), name="embedding")
x = keras.layers.Dense(256, activation="relu", name="dense_1")(inputs)
x = keras.layers.Dropout(0.15, name="dropout_1")(x)
x = keras.layers.Dense(384, activation="relu", name="dense_2")(x)
x = keras.layers.Dropout(0.2, name="dropout_2")(x)
x = keras.layers.Dense(192, activation="relu", name="dense_3")(x)
x = keras.layers.Dropout(0.25, name="dropout_3")(x)
x = keras.layers.Dense(384, activation="relu", name="dense_4")(x)
x = keras.layers.Dropout(0.2, name="dropout_4")(x)
outputs = keras.layers.Dense(len(class_names), activation="softmax", name="ouput")(
x
)
model = keras.Model(inputs=inputs, outputs=outputs, name="accent_recognition")
model.compile(
optimizer=keras.optimizers.Adam(learning_rate=1.9644e-5),
loss=keras.losses.CategoricalCrossentropy(),
metrics=["accuracy", keras.metrics.AUC(name="auc")],
)
return model
model = build_and_compile_model()
model.summary()
# + [markdown] colab_type="text"
# ## Class weights calculation
#
# Since the dataset is quite unbalanced, we wil use `class_weight` argument during training.
#
# Getting the class weights is a little tricky because even though we know the number of
# audio files for each class, it does not represent the number of samples for that class
# since Yamnet transforms each audio file into multiple audio samples of 0.96 seconds each.
# So every audio file will be split into a number of samples that is proportional to its length.
#
# Therefore, to get those weights, we have to calculate the number of samples for each class
# after preprocessing through Yamnet.
# + colab_type="code"
class_counts = tf.zeros(shape=(len(class_names),), dtype=tf.int32)
for x, y in iter(train_ds):
class_counts = class_counts + tf.math.bincount(
tf.cast(tf.math.argmax(y, axis=1), tf.int32), minlength=len(class_names)
)
class_weight = {
i: tf.math.reduce_sum(class_counts).numpy() / class_counts[i].numpy()
for i in range(len(class_counts))
}
print(class_weight)
# + [markdown] colab_type="text"
# ## Callbacks
#
# We use Keras callbacks in order to:
#
# * Stop whenever the validation AUC stops improving.
# * Save the best model.
# * Call TensorBoard in order to later view the training and validation logs.
# + colab_type="code"
early_stopping_cb = keras.callbacks.EarlyStopping(
monitor="val_auc", patience=10, restore_best_weights=True
)
model_checkpoint_cb = keras.callbacks.ModelCheckpoint(
MODEL_NAME + ".h5", monitor="val_auc", save_best_only=True
)
tensorboard_cb = keras.callbacks.TensorBoard(
os.path.join(os.curdir, "logs", model.name)
)
callbacks = [early_stopping_cb, model_checkpoint_cb, tensorboard_cb]
# + [markdown] colab_type="text"
# ## Training
# + colab_type="code"
history = model.fit(
train_ds,
epochs=EPOCHS,
validation_data=valid_ds,
class_weight=class_weight,
callbacks=callbacks,
verbose=2,
)
# + [markdown] colab_type="text"
# ## Results
#
# Let's plot the training and validation AUC and accuracy.
# + colab_type="code"
fig, axs = plt.subplots(nrows=1, ncols=2, figsize=(14, 5))
axs[0].plot(range(EPOCHS), history.history["accuracy"], label="Training")
axs[0].plot(range(EPOCHS), history.history["val_accuracy"], label="Validation")
axs[0].set_xlabel("Epochs")
axs[0].set_title("Training & Validation Accuracy")
axs[0].legend()
axs[0].grid(True)
axs[1].plot(range(EPOCHS), history.history["auc"], label="Training")
axs[1].plot(range(EPOCHS), history.history["val_auc"], label="Validation")
axs[1].set_xlabel("Epochs")
axs[1].set_title("Training & Validation AUC")
axs[1].legend()
axs[1].grid(True)
plt.show()
# + [markdown] colab_type="text"
# ## Evaluation
# + colab_type="code"
train_loss, train_acc, train_auc = model.evaluate(train_ds)
valid_loss, valid_acc, valid_auc = model.evaluate(valid_ds)
# + [markdown] colab_type="text"
# Let's try to compare our model's performance to Yamnet's using one of Yamnet metrics (d-prime)
# Yamnet achieved a d-prime value of 2.318.
# Let's check our model's performance.
# + colab_type="code"
# The following function calculates the d-prime score from the AUC
def d_prime(auc):
standard_normal = stats.norm()
d_prime = standard_normal.ppf(auc) * np.sqrt(2.0)
return d_prime
print(
"train d-prime: {0:.3f}, validation d-prime: {1:.3f}".format(
d_prime(train_auc), d_prime(valid_auc)
)
)
# + [markdown] colab_type="text"
# We can see that the model achieves the following results:
#
# Results | Training | Validation
# -----------|-----------|------------
# Accuracy | 54% | 51%
# AUC | 0.91 | 0.89
# d-prime | 1.882 | 1.740
# + [markdown] colab_type="text"
# ## Confusion Matrix
#
# Let's now plot the confusion matrix for the validation dataset.
#
# The confusion matrix lets us see, for every class, not only how many samples were correctly classified,
# but also which other classes were the samples confused with.
#
# It allows us to calculate the precision and recall for every class.
# + colab_type="code"
# Create x and y tensors
x_valid = None
y_valid = None
for x, y in iter(valid_ds):
if x_valid is None:
x_valid = x.numpy()
y_valid = y.numpy()
else:
x_valid = np.concatenate((x_valid, x.numpy()), axis=0)
y_valid = np.concatenate((y_valid, y.numpy()), axis=0)
# Generate predictions
y_pred = model.predict(x_valid)
# Calculate confusion matrix
confusion_mtx = tf.math.confusion_matrix(
np.argmax(y_valid, axis=1), np.argmax(y_pred, axis=1)
)
# Plot the confusion matrix
plt.figure(figsize=(10, 8))
sns.heatmap(
confusion_mtx, xticklabels=class_names, yticklabels=class_names, annot=True, fmt="g"
)
plt.xlabel("Prediction")
plt.ylabel("Label")
plt.title("Validation Confusion Matrix")
plt.show()
# + [markdown] colab_type="text"
# ## Precision & recall
#
# For every class:
#
# * Recall is the ratio of correctly classified samples i.e. it shows how many samples
# of this specific class, the model is able to detect.
# It is the ratio of diagonal elements to the sum of all elements in the row.
# * Precision shows the accuracy of the classifier. It is the ratio of correctly predicted
# samples among the ones classified as belonging to this class.
# It is the ratio of diagonal elements to the sum of all elements in the column.
# + colab_type="code"
for i, label in enumerate(class_names):
precision = confusion_mtx[i, i] / np.sum(confusion_mtx[:, i])
recall = confusion_mtx[i, i] / np.sum(confusion_mtx[i, :])
print(
"{0:15} Precision:{1:.2f}%; Recall:{2:.2f}%".format(
label, precision * 100, recall * 100
)
)
# + [markdown] colab_type="text"
# ## Run inference on test data
#
# Let's now run a test on a single audio file.
# Let's check this example from [The Scottish Voice](https://www.thescottishvoice.org.uk/home/)
#
# We will:
#
# * Download the mp3 file.
# * Convert it to a 16k wav file.
# * Run the model on the wav file.
# * Plot the results.
# + colab_type="code"
filename = "audio-sample-Stuart"
url = "https://www.thescottishvoice.org.uk/files/cm/files/"
if os.path.exists(filename + ".wav") == False:
print(f"Downloading {filename}.mp3 from {url}")
command = f"wget {url}{filename}.mp3"
os.system(command)
print(f"Converting mp3 to wav and resampling to 16 kHZ")
command = (
f"ffmpeg -hide_banner -loglevel panic -y -i {filename}.mp3 -acodec "
f"pcm_s16le -ac 1 -ar 16000 {filename}.wav"
)
os.system(command)
filename = filename + ".wav"
# + [markdown] colab_type="text"
# The below function `yamnet_class_names_from_csv` was copied and very slightly changed
# from this [Yamnet Notebook](https://colab.research.google.com/github/tensorflow/hub/blob/master/examples/colab/yamnet.ipynb).
# + colab_type="code"
def yamnet_class_names_from_csv(yamnet_class_map_csv_text):
"""Returns list of class names corresponding to score vector."""
yamnet_class_map_csv = io.StringIO(yamnet_class_map_csv_text)
yamnet_class_names = [
name for (class_index, mid, name) in csv.reader(yamnet_class_map_csv)
]
yamnet_class_names = yamnet_class_names[1:] # Skip CSV header
return yamnet_class_names
yamnet_class_map_path = yamnet_model.class_map_path().numpy()
yamnet_class_names = yamnet_class_names_from_csv(
tf.io.read_file(yamnet_class_map_path).numpy().decode("utf-8")
)
def calculate_number_of_non_speech(scores):
number_of_non_speech = tf.math.reduce_sum(
tf.where(tf.math.argmax(scores, axis=1, output_type=tf.int32) != 0, 1, 0)
)
return number_of_non_speech
def filename_to_predictions(filename):
# Load 16k audio wave
audio_wav = load_16k_audio_wav(filename)
# Get audio embeddings & scores.
scores, embeddings, mel_spectrogram = yamnet_model(audio_wav)
print(
"Out of {} samples, {} are not speech".format(
scores.shape[0], calculate_number_of_non_speech(scores)
)
)
# Predict the output of the accent recognition model with embeddings as input
predictions = model.predict(embeddings)
return audio_wav, predictions, mel_spectrogram
# + [markdown] colab_type="text"
# Let's run the model on the audio file:
# + colab_type="code"
audio_wav, predictions, mel_spectrogram = filename_to_predictions(filename)
infered_class = class_names[predictions.mean(axis=0).argmax()]
print(f"The main accent is: {infered_class} English")
# + [markdown] colab_type="text"
# Listen to the audio
# + colab_type="code"
Audio(audio_wav, rate=16000)
# + [markdown] colab_type="text"
# The below function was copied from this [Yamnet notebook](tinyurl.com/4a8xn7at) and adjusted to our need.
#
# This function plots the following:
#
# * Audio waveform
# * Mel spectrogram
# * Predictions for every time step
# + colab_type="code"
plt.figure(figsize=(10, 6))
# Plot the waveform.
plt.subplot(3, 1, 1)
plt.plot(audio_wav)
plt.xlim([0, len(audio_wav)])
# Plot the log-mel spectrogram (returned by the model).
plt.subplot(3, 1, 2)
plt.imshow(
mel_spectrogram.numpy().T, aspect="auto", interpolation="nearest", origin="lower"
)
# Plot and label the model output scores for the top-scoring classes.
mean_predictions = np.mean(predictions, axis=0)
top_class_indices = np.argsort(mean_predictions)[::-1]
plt.subplot(3, 1, 3)
plt.imshow(
predictions[:, top_class_indices].T,
aspect="auto",
interpolation="nearest",
cmap="gray_r",
)
# patch_padding = (PATCH_WINDOW_SECONDS / 2) / PATCH_HOP_SECONDS
# values from the model documentation
patch_padding = (0.025 / 2) / 0.01
plt.xlim([-patch_padding - 0.5, predictions.shape[0] + patch_padding - 0.5])
# Label the top_N classes.
yticks = range(0, len(class_names), 1)
plt.yticks(yticks, [class_names[top_class_indices[x]] for x in yticks])
_ = plt.ylim(-0.5 + np.array([len(class_names), 0]))
| examples/audio/ipynb/uk_ireland_accent_recognition.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (TensorFlow 2 CPU Optimized)
# language: python
# name: python3__SAGEMAKER_INTERNAL__arn:aws:sagemaker:us-west-2:236514542706:image/tensorflow-2.1-cpu-py36
# ---
# # Amazon SageMaker inference deployment to CPUs, GPUs, and EI
# This example demonstrates Amazon SageMaker inference deployment using SageMaker SDK
#
# This example was tested on Amazon SageMaker Studio Notebook
# Run this notebook using the following Amazon SageMaker Studio conda environment:
# `TensorFlow 2 CPU Optimized`
# +
# # !pip install --upgrade pip -q
# # !pip install --upgrade sagemaker -q
# +
import tarfile
import sagemaker
import tensorflow as tf
import tensorflow.keras as keras
import shutil
import os
import time
from tensorflow.keras.applications.resnet50 import ResNet50
role = sagemaker.get_execution_role()
sess = sagemaker.Session()
region = sess.boto_region_name
bucket = sess.default_bucket()
print('sagemaker version: '+sagemaker.__version__)
print('tensorflow version: '+tf.__version__)
# +
def load_save_resnet50_model(model_path):
model = ResNet50(weights='imagenet')
shutil.rmtree(model_path, ignore_errors=True)
model.save(model_path, include_optimizer=False, save_format='tf')
saved_model_dir = 'resnet50_saved_model'
model_ver = '1'
model_path = os.path.join(saved_model_dir, model_ver)
# load_save_resnet50_model(model_path)
# -
shutil.rmtree('model.tar.gz', ignore_errors=True)
# !tar cvfz model.tar.gz -C resnet50_saved_model .
# +
from sagemaker.tensorflow.model import TensorFlowModel, TensorFlowPredictor
prefix = 'keras_models'
s3_model_path = sess.upload_data(path='model.tar.gz', key_prefix=prefix)
model = TensorFlowModel(model_data=s3_model_path,
framework_version='1.15',
role=role,
predictor_cls = TensorFlowPredictor,
sagemaker_session=sess)
# -
# ### Deploy to CPU instance
predictor_cpu = model.deploy(initial_instance_count=1,
instance_type='ml.c5.xlarge')
# ### Deploy using EI
predictor_ei = model.deploy(initial_instance_count=1,
instance_type='ml.c5.xlarge',
accelerator_type='ml.eia2.large')
# ### Deploy to GPU instance
predictor_gpu = model.deploy(initial_instance_count=1,
instance_type='ml.g4dn.xlarge')
# ### Test endpoint
# +
## If you have an existing endpoint, create a predictor using the endpoint name
# from sagemaker.tensorflow.model import TensorFlowPredictor
# predictor = TensorFlowPredictor('ENDPOINT_NAME',
# sagemaker_session=sess)
# -
def image_preprocess(img, reps=1):
img = np.asarray(img.resize((224, 224)))
img = np.stack([img]*reps)
img = tf.keras.applications.resnet50.preprocess_input(img)
return img
# +
from PIL import Image
import numpy as np
import json
img= Image.open('kitten.jpg')
img = image_preprocess(img, 5)
# -
# ### Invoke CPU endpoint
response = predictor_cpu.predict(data=img)
probs = np.array(response['predictions'][0])
tf.keras.applications.resnet.decode_predictions(np.expand_dims(probs, axis=0), top=5)
# ### Invoke CPU Instance + EI endpoint
response = predictor_ei.predict(data=img)
probs = np.array(response['predictions'][0])
tf.keras.applications.resnet.decode_predictions(np.expand_dims(probs, axis=0), top=5)
# ### Invoke G4 GPU Instance with NVIDIA T4 endpoint
response = predictor_gpu.predict(data=img)
probs = np.array(response['predictions'][0])
tf.keras.applications.resnet.decode_predictions(np.expand_dims(probs, axis=0), top=5)
| sagemaker-tf-cpu-gpu-ei-resnet50.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from IPython.display import Image
import numpy as np
np.random.seed(1)
# -
Image(filename='and-or.png')
or_input = np.array([[0,0], [0,1], [1,0], [1,1]])
or_output = np.array([[0,1,1,1]]).T
or_input
or_output
# +
def sigmoid(x): # Returns values that range between -1 and 1
# BTW, this is pretty fun stuff: https://www.google.com.sg/#q=1/(1%2Bexp(-x))
return 1/(1+np.exp(-x))
def sigmoid_derivative(x): # We don't really care what it outputs, lolz...
return x*(1-x)
# -
sigmoid(np.array([2.5, 0.32, -1.42]))
sigmoid_derivaive(np.array([2.5, 0.32, -1.42]))
sigmoid_derivative(np.array([2.5, 0.32, -2.42]))
def cost(predicted, truth):
return truth - predicted
gold = np.array([0.5, 1.2, 9.8])
pred = np.array([0.6, 1.0, 10.0])
cost(pred, gold)
gold = np.array([0.5, 1.2, 9.8])
pred = np.array([9.3, 4.0, 99.0])
cost(pred, gold)
# +
num_data, input_dim = or_input.shape
output_dim = len(or_output.T)
# Initialize weights for the input layer, aka "syn0", syn is short for synapse.
syn0 = np.random.random((input_dim, output_dim))
# +
num_epochs = 10000
learning_rate = 1.0
X = or_input
y = or_output
for _ in range(num_epochs):
# forward propagation.
l0 = X
l1 = sigmoid(np.dot(l0, syn0))
# how much did we miss?
l1_error = cost(l1, y)
# back propagation.
# multiply how much we missed by the
# slope of the sigmoid at the values in l1
l1_delta = l1_error * sigmoid_derivative(l1)
# update weights
syn0 += learning_rate * np.dot(l0.T, l1_delta)
# -
l1
[int(l > 0.5) for l in l1]
or_output
# Do the same but with 1 more hidden layer
# ====
# +
def sigmoid(x): # Returns values that range between -1 and 1
# BTW, this is pretty fun stuff: https://www.google.com.sg/#q=1/(1%2Bexp(-x))
return 1/(1+np.exp(-x)) # ... # YOUR CODE HERE
def sigmoid_derivative(x): # We don't really care what it outputs, lolz...
return x*(1-x) # ... # YOUR CODE HERE
# Cost functions.
def cost(predicted, truth):
return truth - predicted
X = or_input = np.array([[0,0], [0,1], [1,0], [1,1]])
y = or_output = np.array([[0,1,1,1]]).T
# Initialize weights for the input layer, aka "syn0", syn is short for synapse.
num_data, input_dim = or_input.shape
hidden_dim = 5 #... # YOUR CODE HERE
syn0 = np.random.random((input_dim, hidden_dim))#... # YOUR CODE HERE
# Initialize weights for the first hidden layer, aka "syn1".
output_dim = len(or_output.T) #... # YOUR CODE HERE
syn1 = np.random.random((hidden_dim, output_dim)) #... # YOUR CODE HERE
num_epochs = 10000
learning_rate = 1.0
cost = cost
for _ in range(num_epochs):
# forward propagation.
l0 = X
l1 = sigmoid(np.dot(l0, syn0))
l2 = sigmoid(np.dot(l1, syn1))
# how much did we miss?
l2_error = cost(l2, y)
# Now we back propagate...
# in what direction is the target value?
# were we really sure? if so, don't change too much.
l2_delta = l2_error * sigmoid_derivative(l2)
# how much did each l1 value contribute to the l2 error (according to the weights)?
l1_error = l2_delta.dot(syn1.T)
# in what direction is the target l1?
# were we really sure? if so, don't change too much.
l1_delta = l1_error * sigmoid_derivative(l1)
syn1 += l1.T.dot(l2_delta)
syn0 += l0.T.dot(l1_delta)
# -
l2 # output layer.
l1 # hidden layer.
# Now let's make it even more challenging,
# ====
# we'll drop one data point and use 3 layers
# ====
# +
def sigmoid(x): # Returns values that range between -1 and 1
# BTW, this is pretty fun stuff: https://www.google.com.sg/#q=1/(1%2Bexp(-x))
return 1/(1+np.exp(-x)) # ... # YOUR CODE HERE
def sigmoid_derivative(x): # We don't really care what it outputs, lolz...
return x*(1-x) # ... # YOUR CODE HERE
# Cost functions.
def cost(predicted, truth):
return truth - predicted
X = or_input = np.array([[0,0], [0,1], [1,0]])
y = or_output = np.array([[0,1,1]]).T
# Initialize weights for the input layer, aka "syn0", syn is short for synapse.
num_data, input_dim = or_input.shape
hidden_dim_1 = 5 #... # YOUR CODE HERE
syn0 = np.random.random((input_dim, hidden_dim_1))#... # YOUR CODE HERE
# Initialize weights for the first hidden layer, aka "syn1".
hidden_dim_2 = 3 #... # YOUR CODE HERE
syn1 = np.random.random((hidden_dim_1, hidden_dim_2)) #... # YOUR CODE HERE
# Initialize weights for the first hidden layer, aka "syn2".
output_dim = len(or_output.T) #... # YOUR CODE HERE
syn2 = np.random.random((hidden_dim_2, output_dim)) #... # YOUR CODE HERE
num_epochs = 10000
learning_rate = 1.0
cost = cost
for _ in range(num_epochs):
# forward propagation.
l0 = X
l1 = sigmoid(np.dot(l0, syn0))
l2 = sigmoid(np.dot(l1, syn1))
l3 = sigmoid(np.dot(l2, syn2))
# how much did we miss?
l3_error = cost(l3, y)
# Now we back propagate...
# in what direction is the target value?
# were we really sure? if so, don't change too much.
l3_delta = l3_error * sigmoid_derivative(l3)
# how much did each l2 value contribute to the l3 error (according to the weights)?
l2_error = l3_delta.dot(syn2.T)
# in what direction is the l2 weights changing?
l2_delta = l2_error * sigmoid_derivative(l2)
# how much did each l1 value contribute to the l2 error (according to the weights)?
l1_error = l2_delta.dot(syn1.T)
# in what direction is the target l1?
# were we really sure? if so, don't change too much.
l1_delta = l1_error * sigmoid_derivative(l1)
syn2 += l2.T.dot(l3_delta)
syn1 += l1.T.dot(l2_delta)
syn0 += l0.T.dot(l1_delta)
# -
l3
# +
new_input = np.array([[1,1]])
# apply l1 to new input
_l1 = sigmoid(np.dot(new_input, syn0))
# apply l2 to new input
_l2 = sigmoid(np.dot(_l1, syn1))
# apply l3 (output layer) to new input
prediction = _l3 = sigmoid(np.dot(_l2, syn2))
prediction
# -
# Now let's do the same in tensorflow!!!
# ====
import tensorflow as tf
# +
# Parameters
learning_rate = 1.0
num_epochs = 10000
# Network Parameters
hidden_dim_1 = 5 # 1st layer number of features
hidden_dim_2 = 3 # 2nd layer number of features
input_dim = 2 # Input dimensions.
output_dim = 1 # Output dimensions.
# tf Graph input
x = tf.placeholder("float", [None, input_dim])
y = tf.placeholder("float", [hidden_dim_2, output_dim])
# +
# Without biases.
weights = {
'syn0': tf.Variable(tf.random_normal([input_dim, hidden_dim_1])),
'syn1': tf.Variable(tf.random_normal([hidden_dim_1, hidden_dim_2])),
'syn2': tf.Variable(tf.random_normal([hidden_dim_2, output_dim]))
}
# Create a model
def multilayer_perceptron(X, weights, biases):
# Hidden layer 1
layer_1 = tf.matmul(X, weights['syn0'])
# Hidden layer 2
layer_2 = tf.matmul(layer_1, weights['syn1'])
# Output layer
out_layer = tf.matmul(layer_2, weights['syn2'])
return out_layer
# +
# With biases.
weights = {
'syn0': tf.Variable(tf.random_normal([input_dim, hidden_dim_1])),
'syn1': tf.Variable(tf.random_normal([hidden_dim_1, hidden_dim_2])),
'syn2': tf.Variable(tf.random_normal([hidden_dim_2, output_dim]))
}
biases = {
'b0': tf.Variable(tf.random_normal([hidden_dim_1])),
'b1': tf.Variable(tf.random_normal([hidden_dim_2])),
'b2': tf.Variable(tf.random_normal([output_dim]))
}
# Create a model
def multilayer_perceptron(X, weights, biases):
# Hidden layer 1
layer_1 = tf.add(tf.matmul(X, weights['syn0']), biases['b0'])
# Hidden layer 2
layer_2 = tf.add(tf.matmul(layer_1, weights['syn1']), biases['b1'])
# Output layer
out_layer = tf.matmul(layer_2, weights['syn2']) + biases['b2']
return out_layer
# +
# With biases.
weights = {
'syn0': tf.Variable(tf.random_normal([input_dim, hidden_dim_1])),
'syn1': tf.Variable(tf.random_normal([hidden_dim_1, hidden_dim_2])),
'syn2': tf.Variable(tf.random_normal([hidden_dim_2, output_dim]))
}
biases = {
'b0': tf.Variable(tf.random_normal([hidden_dim_1])),
'b1': tf.Variable(tf.random_normal([hidden_dim_2])),
'b2': tf.Variable(tf.random_normal([output_dim]))
}
# Create a model
def multilayer_perceptron(X, weights, biases):
# Hidden layer 1 + sigmoid activation function
layer_1 = tf.add(tf.matmul(X, weights['syn0']), biases['b0'])
layer_1 = tf.nn.sigmoid(layer_1)
# Hidden layer 2 + sigmoid activation function
layer_2 = tf.add(tf.matmul(layer_1, weights['syn1']), biases['b1'])
layer_2 = tf.nn.sigmoid(layer_2)
# Output layer
out_layer = tf.matmul(layer_2, weights['syn2']) + biases['b2']
out_layer = tf.nn.sigmoid(out_layer)
return out_layer
# +
# Construct model
pred = multilayer_perceptron(x, weights, biases)
# Define loss and optimizer
cost = tf.sub(y, pred)
# Or you can use fancy cost like:
##tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(pred, y))
optimizer = tf.train.GradientDescentOptimizer(learning_rate).minimize(cost)
init = tf.initialize_all_variables()
# -
np.array([[0,1,1]]).T
# +
or_input = np.array([[0.,0.], [0.,1.], [1.,0.]])
or_output = np.array([[0.,1.,1.]]).T
# Launch the graph
with tf.Session() as sess:
sess.run(init)
# Training cycle
for epoch in range(num_epochs):
batch_x, batch_y = or_input, or_output # Loop over all data points.
# Run optimization op (backprop) and cost op (to get loss value)
_, c = sess.run([optimizer, cost], feed_dict={x: batch_x, y: batch_y})
#print (c)
# Now let's test it on the unknown dataset.
new_inputs = np.array([[1.,1.], [1.,0.]])
feed_dict = {x: new_inputs}
predictions = sess.run(pred, feed_dict)
print (predictions)
# -
# Now let's do the same in tensorflow learn (aka skflow)!!!
# ====
from tensorflow.contrib import learn
classifier = learn.DNNClassifier(hidden_units=[5, 3], n_classes=2)
# +
or_input = np.array([[0.,0.], [0.,1.], [1.,0.]])
or_output = np.array([[0,1,1]]).T
classifier.fit(or_input, or_output, steps=0.05, batch_size=3)
# -
classifier.predict(np.array([ [1., 1.], [1., 0.] , [0., 0.] , [0., 1.]]))
# +
from tensorflow.contrib import learn
classifier = learn.DNNClassifier(hidden_units=[5, 3], n_classes=2)
or_input = np.array([[0.,0.], [0.,1.], [1.,0.]])
or_output = np.array([[0,1,1]]).T
classifier.fit(or_input, or_output, steps=10000, batch_size=3)
classifier.predict(np.array([ [1., 1.], [1., 0.] , [0., 0.] , [0., 1.]]))
# -
| Bosch 800 Series.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Run all notebooks
#
# Run all notebooks in correct order.
# %run ./00_setup_shapes.ipynb
# %run ./0_process_bathymetry.ipynb
# %run ./1a_gather_data_stations.ipynb
# %run ./1b_gather_data_rain.ipynb
# %run ./2_data_description.ipynb
# %run ./3_calculation.ipynb
| notebooks/run.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:pyosim_aws]
# language: python
# name: conda-env-pyosim_aws-py
# ---
# <!--BOOK_INFORMATION-->
# <img style="float: right; width: 100px" src="https://raw.github.com/pyomeca/design/master/logo/logo_cropped.svg?sanitize=true">
#
# <font size="+2">Pyosim in the cloud</font>
#
# <font size="+1">with [pyomeca](https://github.com/pyomeca/pyom</font>a)
#
# <NAME> (<EMAIL> | [GitHub](https://github.com/romainmartinez))
#
# # Thorax rotation
# +
from pathlib import Path
import yaml
import numpy as np
import pandas as pd
import altair as alt
# from pyosim import Conf
from pyomeca import Markers3d, Analogs3d, FrameDependentNpArray
import spm1d
import matplotlib.pyplot as plt
from src.util import (
parse_conditions,
condition_counter,
random_balanced_design,
get_spm_cluster,
describe_clusters,
)
# -
# %load_ext autoreload
# %autoreload 2
# %load_ext lab_black
alt.data_transformers.enable("data_server")
# to make this notebook's output stable across runs
RANDOM_SEED = 42
np.random.seed(RANDOM_SEED)
def parse_conditions(d):
d["hand"] = d["filename"].str[17]
d["filename"] = d["filename"].str.replace("wu_expertsnovices[D|G]_", "")
return d.merge(
pd.read_csv(project_path / "_conf.csv")[["participant", "group"]].rename(
columns={"group": "expert"}
),
on="participant",
).assign(
fatigue=lambda x: x["filename"].str[0].astype(int) - 1,
height=lambda x: x["filename"].str[2].astype("category"),
mass=lambda x: x["filename"].str[3:5].astype(int),
filename=lambda x: x["participant"] + "_" + x["filename"],
participant=lambda x: x["participant"].astype("category"),
n_trial=lambda x: x["filename"].str.split("_").str[-1],
)
project_path = Path("/home/romain/Downloads/experts-novices/opensim/")
def correct_path(x):
return Path(
x.replace(
"home/laboratoire/mnt/E/Projet_ExpertsNovices/",
"/home/romain/Downloads/experts-novices/",
)
)
# +
mot = []
f = []
onsets = pd.read_csv(project_path / "onsets.csv")
for _, ifile in onsets.iterrows():
filename = correct_path(ifile["filepathK"])
if filename in f:
continue
else:
f.append(filename)
try:
e = Analogs3d.from_mot(filename)
except FileNotFoundError:
print(f"{filename} not found")
continue
idx = np.logical_and(
e.get_time_frames < ifile["offset"], e.get_time_frames > ifile["onset"]
)
e = e[..., idx]
mot.append(
e.center()
.time_normalization()
.to_dataframe()[["thorax_rotation"]]
.assign(filename=filename.stem, participant=filename.parent.parent.stem)
)
# +
data = (
pd.concat(mot)
.reset_index()
.assign(index=lambda x: x["index"] / 100)
.pipe(parse_conditions)
.query('fatigue == 0 & height == "r"')
.drop("fatigue", axis=1)
)
data.sample(5)
# -
# ## Correct for left handed
data.loc[data["hand"] == "G", "thorax_rotation"] = (
data.loc[data["hand"] == "G", "thorax_rotation"] * -1
)
# ## Correct rotation
data["thorax_rotation"] = data.groupby("filename")["thorax_rotation"].apply(
lambda x: (x if x.iloc[0] < 0 else -x) + 100
)
# data["thorax_rotation"] = data.groupby("filename")["thorax_rotation"].apply(
# lambda x: x - x.iloc[0]
# )
# data["thorax_rotation"] = data["thorax_rotation"].replace(to_replace=0, method="bfill")
alt.Chart(data).mark_line(opacity=0.1).encode(
alt.X("index"),
alt.Y("thorax_rotation"),
alt.Detail("filename"),
alt.Color("expert:N"),
)
# ### Balance dataset
#
# By randomly taking the minimum number of trials for each condition
data.drop_duplicates(["filename", "participant"]).groupby(["expert", "mass"]).size()
balanced_trials = random_balanced_design(
data, ["expert", "mass"], random_state=RANDOM_SEED, participant=True
)
_filename, _participant = (
balanced_trials["filename"].to_list(),
balanced_trials["participant"].to_list(),
)
data = data.query("filename == @_filename & participant == @_participant")
data.drop_duplicates(["filename", "participant"]).groupby(["expert", "mass"]).size()
# ### Stats
id_vars = ["filename", "participant", "expert", "height", "mass", "n_trial", "index"]
value = "thorax_rotation"
stats_df = (
data[id_vars]
.assign(value=data[value])
.assign(participant=lambda x: x["participant"].astype("category").cat.codes)
.set_index(id_vars)
.unstack()
)
stats_df = stats_df.fillna(stats_df.mean())
stats_df
# +
α = 0.05
n_iter = 1000
spm = spm1d.stats.nonparam.anova2(
stats_df.to_numpy(),
A=stats_df.index.get_level_values("expert"),
B=stats_df.index.get_level_values("mass"),
)
spmi = spm.inference(alpha=α, iterations=n_iter)
plt.figure(figsize=(11, 8))
spmi.plot()
plt.tight_layout()
# -
for i in spmi:
print(i)
clusters = pd.DataFrame(
{
"effect": ["main sex", "main mass"],
"p": [0, 0],
"start": [0, 0],
"end": [100, 100],
}
)
effect = {"main sex": "expert", "main mass": "mass", "interaction sex-mass": "expert"}
describe_clusters(
clusters, data[id_vars + [value]].rename(columns={value: "value"}).dropna(), effect
)
base = alt.Chart(data[id_vars + [value]]).encode(
alt.X(
"index", title="Normalized trial", axis=alt.Axis(format="%", labelFlush=False)
)
)
mu = base.mark_line().encode(
alt.Y(f"mean({value})", title="box-thorax distance (% height)",)
)
sigma = base.mark_errorband(extent="stdev").encode(alt.Y(value, title=None))
men_scale = alt.Scale(scheme="set1")
(mu + sigma)
(mu + sigma).encode(alt.Color("expert:N", scale=men_scale))
(mu + sigma).encode(alt.Color("mass:N"))
(mu + sigma).encode(alt.Color("expert:N", scale=men_scale)).facet(column="mass")
| notebooks/03.06-thorax-rotation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Extraction of cluster geometries from a trajectory
from exa.util import isotopes
import exatomic
from exatomic.core.two import compute_atom_two_out_of_core # If we need to compute atom_two out of core (low RAM)
from exatomic.algorithms.neighbors import periodic_nearest_neighbors_by_atom # Only valid for simple cubic periodic cells
from exatomic.base import resource # Helper function to find the path to an example file
xyzpath = resource("rh2.traj.xyz")
xyz = exatomic.XYZ(xyzpath) # Editor that can parse the XYZ file and build a universe!
u = xyz.to_universe() # Parse the XYZ data to a universe
u.info() # Current tables
# Add the unit cell dimensions to the frame table of the universe
a = 37.08946302
for i, q in enumerate(("x", "y", "z")):
for j, r in enumerate(("i", "j", "k")):
if i == j:
u.frame[q+r] = a
else:
u.frame[q+r] = 0.0
u.frame["o"+q] = 0.0
u.frame['periodic'] = True
len(u) # Number of steps in this trajectory (small for example purposes)
isotopes.Rh.radius # Default Rh (covalent) radius
# ### To extract clusters from the trajectory, we don't need to pre-compute atom_two
u.atom.head()
# %%time
dct = periodic_nearest_neighbors_by_atom(u, # Universe with all frames from which we want to extract clusters
"Rh", # Source atom from which we will search for neighbors
a, # Cubic cell dimension
# Cluster sizes we want
[0, 1, 2, 3, 4, 8, 12, 16],
# Additional arguments to be passed to the atomic two body calculation
# Note that it is critical to increase dmax (in compute_atom_two)!!!
Rh=2.6, dmax=a/2)
dct.keys() # 'dct' is a dictionary containing our clustered universes as well as the sorted neighbor list (per frame)
dct['nearest'].head() # Table of sorted indexes of nearest molecules to "Rh" in each frame
exatomic.UniverseWidget(dct[0]) # Just the analyte
exatomic.UniverseWidget(dct[16]) # View the universe with 16 nearest molecules
# ### If we had wanted to view the entire universe...
# %time u.compute_atom_two(Rh=2.6) # Compute atom two body data with slightly larger Rh radius
# OR
# #%time compute_atom_two_out_of_core("atom_two.hdf", u, a, Rh=2.6) # Writes the two body data to "atom_two.hdf" with keys "frame_{index}/atom_two"
u.memory_usage().sum()/1024**2 # RAM usage (MB)
exatomic.UniverseWidget(u)
| docs/source/notebooks/04_cluster_extraction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
#MPI and OPENMP Strong Scaling STRONG SCALING
#input n_x=2000 n_y=2000
T_1 = 440.25
def speedup(t_1,T_p):
return t_1/T_p
# (num_threads, speedup)
MPIStrongScaling = [(2, speedup(T_1,247.91)), (4, speedup(T_1,84.69)),
(6, speedup(T_1,52.70)), (8, speedup(T_1,33.88)),
(12, speedup(T_1,26.41)), (16, speedup(T_1,19.38)),
(20, speedup(T_1,16.21))]
OpenMpStrongScaling = [(2, speedup(T_1,121.32)), (4, speedup(T_1,62.31)),
(6, speedup(T_1,43.39)), (8, speedup(T_1,33.02)),
(12, speedup(T_1,23.13)), (16, speedup(T_1,17.54)),
(20, speedup(T_1,14.04))]
#PLOTTING
plt.scatter(*zip(*MPIStrongScaling), label= "MPI n_x,n_y=2000")
plt.plot(*zip(*MPIStrongScaling))
plt.scatter(*zip(*OpenMpStrongScaling), label= "OPENMP n_x,n_y=2000")
plt.plot(*zip(*OpenMpStrongScaling))
plt.legend()
plt.xlabel("# MPI processes / threads")
plt.ylabel("Speedup")
plt.rcParams["figure.figsize"] = [16,9]
# +
#WEAK SCALING MPI and OPENMP
#n_x and n_y are ranges in 500, 707, 1000, 1225, 1421, 1579
T_p_MPI = [15.49, 10.66,10.52,10.49,9.82,10.15]
T_p_OPENMP = [7.69, 7.84,8.36,8.74,9.02,8.82]
#times measured with the serial program with n increasing
T_1 = [27.58,55.03,110.17,165.20,222.26,274.44]
n_procs_or_threads= [2,4,8,12,16,20]
speedup_MPI = []
speedup_OPENMP = []
res_MPI_ws = []
res_OPENMP_ws = []
for i in range(0,6):
speedup_MPI.append(T_1[i]/T_p_MPI[i])
speedup_OPENMP.append(T_1[i]/T_p_OPENMP[i])
res_MPI_ws.append((n_procs_or_threads[i],speedup_MPI[i]))
res_OPENMP_ws.append((n_procs_or_threads[i], speedup_OPENMP[i]))
plt.scatter(*zip(*res_MPI_ws), label="MPI weak scaling")
plt.plot(*zip(*res_MPI_ws))
plt.scatter(*zip(*res_OPENMP_ws), label="OPENMP weak scaling")
plt.plot(*zip(*res_OPENMP_ws))
plt.legend()
plt.xlabel("MPI processes / threads")
plt.ylabel("speedup")
# -
# +
T_1 = 440.25
def speedup(t_1,T_p):
return t_1/T_p
# (num_threads, speedup)
hybridStrongScal = [(1, speedup(T_1,14.09)), (4, speedup(T_1,19.18)),
(10, speedup(T_1,16.01)), (20, speedup(T_1,16.25))]
#PLOTTING
plt.scatter(*zip(*hybridStrongScal), label= "hybrid n_x,n_y=2000")
plt.plot(*zip(*hybridStrongScal))
plt.xticks((1, 4, 10, 20), ("(1,20)", "(4,6)", "(10,2)", "(20,1)"))
plt.legend()
plt.xlabel("(MPI processes, threads)")
plt.ylabel("Speedup")
plt.rcParams["figure.figsize"] = [16,9]
# +
#MPI strong scaling Ulysses v2- n_x=n_y=4000
T_1 = 1947.85
def speedup(t_1,T_p):
return t_1/T_p
# (num_threads, speedup)
MPISCULv2 = [(2, speedup(T_1,1024.04)), (4, speedup(T_1,331.37)),
(8, speedup(T_1,157.91)), (16, speedup(T_1,75.51)),
(32, speedup(T_1,36.91))]
#PLOTTING
plt.scatter(*zip(*MPISCULv2), label= "Strong scaling")
plt.plot(*zip(*MPISCULv2))
plt.xticks((2, 4, 8, 16, 32), ())
plt.legend()
plt.xlabel("(MPI processes, threads)")
plt.ylabel("Speedup")
plt.rcParams["figure.figsize"] = [16,9]
| .ipynb_checkpoints/AssignmentHPC_03-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# # SEIR model
from covid19be import load_data
data = load_data()
# +
opts = {
"frac_dh": 3594 / 7844, # deaths in hospitals / total deaths
"hh": 0.05, # fraction of hospitalized
"gamma": 1 / 12.4, # inverse recovery time"
"epsilon": 1 / 5.2, # inverse incubation time
"dea": 0.5, # fatality rate in icu
"n0": 11000000, # population size
"n0_MRS": 400000, # population en MR/MRS + personnel soignant
"e0_factor": 37, # e0 = i0 * factor
"e0_MRS_factor": 20, # e0_MRS = i0_MRS * factor
"window": 7, # size of the window for fitting Re's"
}
def smoothen(v, n_pts=opts["window"]):
box = np.ones(n_pts) / n_pts
return np.convolve(v, box, mode="same")
def SEIR(r0, i0=3, gg=0.75, n_futures=0, opts=opts):
# Load hyper-parameters
hh = opts["hh"]
gamma = opts["gamma"]
epsilon = opts["epsilon"]
dea = opts["dea"]
n0 = opts["n0"]
n0_MRS = opts["n0_MRS"]
e0_factor = opts["e0_factor"]
window = opts["window"]
# Smoothen and extend R0s
r0 = smoothen(r0)
r0 = np.concatenate((r0, np.array([r0[-1]] * (window + n_futures))))
# Initial conditions
drea = dea * 1 / 5
rrea = (1 - dea) * 1 / 20
hospi = 0.0
n = [n0-n0_MRS]
i = [i0]
e = [i[-1] * e0_factor]
h = [0.0]
l = [0.0]
r = [0.0]
m = [0.0]
s = [n[-1] - e[-1] - i[-1] - r[-1]]
# Simulate forward
n_days = len(r0)
for day in range(n_days):
lam = gamma * r0[day]
if day == 14:
hospi = hh / 7
ds = -lam * (i[-1] / 2 + e[-1]) * s[-1] / n[-1]
de = lam * (i[-1] / 2 + e[-1]) * s[-1] / n[-1] - epsilon * e[-1]
di = epsilon * e[-1] - gamma * i[-1] - hospi * i[-1]
dh = hospi * i[-1] - gg * h[-1] / 7 - (1 - gg) * h[-1] / (4 + 2 * np.tanh((l[-1]-500)/300)) + rrea * l[-1]
dl = (1 - gg) * h[-1] / (4 + 2 * np.tanh((l[-1]-500)/300)) - drea * l[-1] - rrea * l[-1]
dr = gamma * i[-1] + gg * h[-1] / 7
dm = drea * l[-1]
s.append(s[-1] + ds)
e.append(e[-1] + de)
i.append(i[-1] + di)
h.append(h[-1] + dh)
l.append(l[-1] + dl)
if l[-1] > 1895:
dm = dm + (l[-1] - 1895)
l[-1] = 1895
r.append(r[-1] + dr)
m.append(m[-1] + dm)
n.append(s[-1] + e[-1] + i[-1] + h[-1] + l[-1] + r[-1])
return np.array(s), np.array(e), np.array(i), np.array(h), np.array(l), np.array(m), np.array(r)
def SEIR_MRS(r0_mrs, n_futures=0, opts=opts):
# Load hyper-parameters
gamma = opts["gamma"]
epsilon = opts["epsilon"]
n0_MRS = opts["n0_MRS"]
e0_MRS_factor = opts["e0_MRS_factor"]
window = opts["window"]
# Smoothen and extend R0s
r0_mrs = smoothen(r0_mrs)
r0_mrs = np.concatenate((r0_mrs, np.array([r0_mrs[-1]] * (window + n_futures))))
# Initial conditions
alpha = 0.15 / 10
lam = gamma * 4.3
n = [n0_MRS]
i = [1]
e = [i[-1] * e0_MRS_factor]
r = [0.0]
s = [n[-1] - e[-1] - i[-1] - r[-1]]
m = [0.0]
# Simulate forward
n_days = len(r0_mrs)
for day in range(n_days):
lam = gamma * r0_mrs[day]
ds = -lam * (i[-1] / 2 + e[-1]) * s[-1] / n[-1]
de = lam * (i[-1] / 2 + e[-1]) * s[-1] / n[-1] - epsilon * e[-1]
di = epsilon * e[-1] - (gamma + alpha) * i[-1]
dr = gamma * i[-1]
dm = alpha * i[-1]
s.append(s[-1] + ds)
e.append(e[-1] + de)
i.append(i[-1] + di)
r.append(r[-1] + dr)
m.append(m[-1] + dm)
n.append(s[-1] + e[-1] + i[-1] + r[-1])
return np.array(s), np.array(e), np.array(i), np.array(m), np.array(r)
def simulate(theta, n_futures=0):
# Unpack parameters
r0, r0_mrs, i0, gg = theta
# Simulate
s, e, i, h, l, m, r = SEIR(r0, i0=i0, gg=gg, n_futures=n_futures)
_, _, _, m_mrs, _ = SEIR_MRS(r0_mrs, n_futures=n_futures)
return s, e, i, h, l, m, m_mrs, r
# +
from distributions import poisson_logpdf
from scipy.optimize import minimize
# Pack-unpack helpers for passing parameters around
def pack(r0, r0_rms, i0, gg):
v = np.zeros(len(r0) + len(r0_rms) + 2)
v[:len(r0)] = r0
v[len(r0):len(r0)+len(r0_rms)] = r0_rms
v[-2] = i0
v[-1] = gg
return v
def unpack(v):
return v[:(len(v)-2)//2], v[(len(v)-2)//2:len(v)-2], v[-2], v[-1]
# Fit
def fit(data, logpdf=poisson_logpdf, window=opts["window"], frac_dh=opts["frac_dh"]):
def cost(x):
# parameters
r0, r0_mrs, i0, gg = unpack(x)
# cost
c = 0.0
s, e, i, h, l, m, r = SEIR(r0, i0=i0, gg=gg)
c -= logpdf(data["n_hospitalized"].values[9:], mu=h[10:]+l[10:])
c -= logpdf(data["n_icu"].values[9:], mu=l[10:])
c -= logpdf(frac_dh * data["n_deaths"].values[9:-2], mu=m[10:-2])
# fit on deaths with SEIR_MRS
s, e, i, m_mrs, r = SEIR_MRS(r0_mrs)
c -= logpdf(data["n_deaths"].values[:-2], mu=m[1:-2] + m_mrs[1:-2])
# ^ we omit the last two death data points, because not consolidated yet
return c
# x0
r0 = [4.3] * (len(data) - window)
r0_rms = [4.3] * (len(data) - window)
i0 = 3.0
gg = 0.75
x0 = pack(r0, r0_rms, i0, gg)
# bounds
bounds = []
for _ in range(len(r0)):
bounds.append((0.25, 7.5))
for _ in range(len(r0_rms)):
bounds.append((0.25, 7.5))
bounds.append((1, 500))
bounds.append((0.65, 0.85))
# fit
res = minimize(cost, x0=x0, bounds=bounds, options={"maxfun": 25000})
return unpack(res.x)
# -
# # Parametric bootstrap
# +
from distributions import poisson_rvs
from copy import deepcopy
from joblib import Parallel, delayed
def _fit(data, h, l, m, m_mrs, seed, logpdf=poisson_logpdf, rvs=poisson_rvs):
data_i = deepcopy(data)
rs = np.random.RandomState(seed=seed)
# resample
data_i["n_hospitalized"] = rvs(h[1:] + l[1:], random_state=rs)
data_i["n_icu"] = rvs(l[1:], random_state=rs)
data_i["n_deaths"] = rvs(m[1:] + m_mrs[1:], random_state=rs)
# fit on bootstrap
theta_i = fit(data_i, logpdf=logpdf)
return theta_i
def parametric_bootstrap(data, n_replicas=1, logpdf=poisson_logpdf, n_jobs=-1):
# Best fit on original data
theta_mle = fit(data, logpdf=logpdf)
_, _, _, h, l, m, m_mrs, _ = simulate(theta_mle)
# Bootstrap
thetas_bootstrap = Parallel(n_jobs=n_jobs, verbose=10, backend="multiprocessing")(
delayed(_fit)(data, h, l, m, m_mrs, i) for i in range(n_replicas))
return theta_mle, thetas_bootstrap
# -
# this will take a while...
theta_mle, thetas_bootstrap = parametric_bootstrap(data, n_replicas=100, n_jobs=-1)
# +
# Plots
n_futures = 365
n_days = len(data)
window = opts["window"]
bootstraps = {
"h": [],
"l": [],
"m": [],
"m_mrs": [],
"r0": [],
"r0_mrs": []
}
for theta_i in thetas_bootstrap:
_, _, _, h, l, m, m_mrs, _ = simulate(theta_i, n_futures=n_futures)
bootstraps["h"].append(h)
bootstraps["l"].append(l)
bootstraps["m"].append(m)
bootstraps["m_mrs"].append(m_mrs)
r0, r0_mrs, _, _ = theta_i
_r0 = smoothen(r0)
_r0_mrs = smoothen(r0_mrs)
_r0 = np.concatenate((_r0, np.array([_r0[-1]] * (window + n_futures))))
_r0_mrs = np.concatenate((_r0_mrs, np.array([_r0_mrs[-1]] * (window + n_futures))))
bootstraps["r0"].append(_r0)
bootstraps["r0_mrs"].append(_r0_mrs)
for k, v in bootstraps.items():
bootstraps[k] = np.array(v)
# +
hl_50 = np.percentile(bootstraps["h"]+bootstraps["l"], 50, axis=0)
hl_5 = np.percentile(bootstraps["h"]+bootstraps["l"], 5, axis=0)
hl_95 = np.percentile(bootstraps["h"]+bootstraps["l"], 95, axis=0)
l_50 = np.percentile(bootstraps["l"], 50, axis=0)
l_5 = np.percentile(bootstraps["l"], 5, axis=0)
l_95 = np.percentile(bootstraps["l"], 95, axis=0)
m_50 = np.percentile(bootstraps["m"]+bootstraps["m_mrs"], 50, axis=0)
m_5 = np.percentile(bootstraps["m"]+bootstraps["m_mrs"], 5, axis=0)
m_95 = np.percentile(bootstraps["m"]+bootstraps["m_mrs"], 95, axis=0)
r0_50 = np.percentile(bootstraps["r0"], 50, axis=0)
r0_5 = np.percentile(bootstraps["r0"], 5, axis=0)
r0_95 = np.percentile(bootstraps["r0"], 95, axis=0)
r0_mrs_50 = np.percentile(bootstraps["r0_mrs"], 50, axis=0)
r0_mrs_5 = np.percentile(bootstraps["r0_mrs"], 5, axis=0)
r0_mrs_95 = np.percentile(bootstraps["r0_mrs"], 95, axis=0)
# +
fig, ax = plt.subplots(2, 1, figsize=(10, 10), gridspec_kw={"height_ratios": (4, 1)})
# Plot hospitalizations, icus and deaths
ax[0].plot(range(1, n_days+1 + n_futures), hl_50[1:], c="b")
ax[0].fill_between(range(1, n_days+1 + n_futures), hl_5[1:], hl_95[1:], color="b", alpha=0.2)
ax[0].plot(range(1, n_days+1), data["n_hospitalized"].values, ".", c="b", label="hospitalized")
ax[0].plot(range(1, n_days+1 + n_futures), l_50[1:], c="r")
ax[0].fill_between(range(1, n_days+1 + n_futures), l_5[1:], l_95[1:], color="r", alpha=0.2)
ax[0].plot(range(1, n_days+1), data["n_icu"].values, ".", c="r", label="icu")
ax[0].plot(range(1, n_days+1 + n_futures), m_50[1:], c="k")
ax[0].fill_between(range(1, n_days+1 + n_futures), m_5[1:], m_95[1:], color="k", alpha=0.2)
ax[0].plot(range(1, n_days+1), data["n_deaths"].values, ".", c="k", label="deaths")
ax[0].grid()
ax[0].set_ylim(0, 10000)
ax[0].set_xlim(0, 100)
ax[0].set_xticks(np.arange(1, 100, 5))
ax[0].set_xticklabels([d.strftime('%Y-%m-%d') for d in pd.date_range(start="2020-02-28", end="2020-12-31")[:100:5]], rotation=90)
ax[0].legend()
# Plots R0s
ax[1].plot(range(1, n_days+1 + n_futures), r0_50, c="orange", label="R0 in population")
ax[1].fill_between(range(1, n_days+1 + n_futures), r0_5, r0_95, color="orange", alpha=0.2)
ax[1].plot(range(1, n_days+1 + n_futures), r0_mrs_50, c="brown", label="R0 in MRS")
ax[1].fill_between(range(1, n_days+1 + n_futures), r0_mrs_5, r0_mrs_95, color="brown", alpha=0.2)
ax[1].grid(which="both")
ax[1].set_ylim(0, 5)
ax[1].set_yticks(np.arange(0, 5, step=0.5))
for j, label in enumerate(ax[1].get_yticklabels()):
if j % 2 != 0:
label.set_visible(False)
ax[1].set_xlim(0, 100)
ax[1].set_xticks(np.arange(1, 100, 5))
ax[1].set_xticklabels([d.strftime('%Y-%m-%d') for d in pd.date_range(start="2020-02-28", end="2020-12-31")[:100:5]], rotation=90)
ax[1].legend()
plt.subplots_adjust(hspace=0.5)
#plt.savefig("plot-bootstrap.png")
plt.show()
# -
# The parametric bootstrap uncertainty estimates only account for the variability of the best fit, had the data been resampled following the distribution assumed at the best fit parameter estimates.
#
# Importantly, this does *not* account for the epistemic uncertainty in the model hyper-parameters. Accounting for those would produce much larger uncertainty estimates.
| seir-vdwnico-bootstrap.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (oat-use)
# language: python
# name: oat-use
# ---
# %run "Common setup.ipynb"
from SALib.analyze.rbd_fast import analyze as rbd_analyze
numeric_samples = pd.read_csv(f'{DATA_DIR}moat_10_numeric_samples.csv', index_col=0)
numeric_vals = numeric_samples[perturbed_cols].values
# +
# Coupling disabled
moat_10_no_irrigation_results = pd.read_csv(f'{DATA_DIR}moat_no_irrigation_10_results.csv', index_col=0)
moat_10_no_irrigation_results['Avg. $/ML'].fillna(moat_10_no_irrigation_results["Avg. Annual Profit ($M)"], inplace=True)
np_res = moat_10_no_irrigation_results.values
res, idx = [], []
for reps in range(2, np_res.shape[0]+1):
rbd_fast_results = rbd_analyze(CIM_SPEC, numeric_vals[:reps, :],
np_res[:reps],
M=4, seed=101)
disabled = rbd_fast_results.to_df()
tmp = disabled.loc[tgt_param, 'S1']
res.append(tmp)
idx.append(reps)
# End for
disabled = pd.DataFrame({'S1':res}, index=idx)
# +
# Coupling enabled
moat_10_with_irrigation_results = pd.read_csv(f'{DATA_DIR}moat_with_irrigation_10_results.csv', index_col=0)
moat_10_with_irrigation_results['Avg. $/ML'].fillna(moat_10_with_irrigation_results["Avg. Annual Profit ($M)"], inplace=True)
np_res = moat_10_with_irrigation_results.values
res, idx = [], []
for reps in range(2, np_res.shape[0]+1):
rbd_fast_results = rbd_analyze(CIM_SPEC, numeric_vals[:reps, :],
np_res[:reps],
M=4, seed=101)
enabled = rbd_fast_results.to_df()
tmp = enabled.loc[tgt_param, 'S1']
res.append(tmp)
idx.append(reps)
# End for
enabled = pd.DataFrame({'S1':res}, index=idx)
# +
fig, axes = plt.subplots(1,2, figsize=(12,4), sharey=True, sharex=True)
labels = [str(i) if i % 54 == 0 else '' for i in idx]
disabled.loc[:, 'S1'].plot(kind='bar',
legend=None,
title='Disabled',
ax=axes[0],
rot=45,
width=0.7,
edgecolor='C0', logy=True)
enabled.loc[:, 'S1'].plot(kind='bar',
legend=None,
title='Enabled',
ax=axes[1],
rot=45,
width=0.7,
edgecolor='C0', logy=True)
axes[1].legend(bbox_to_anchor=(1.35, 0.65))
axes[0].set_xticklabels(labels)
axes[1].set_xticklabels(labels)
fig.suptitle("EASI Analysis\non Morris Samples", x=0.5, y=1.15, fontsize=20)
plt.xlabel("$N$", x=-0.1, labelpad=15);
# -
fig.savefig(FIG_DIR+'EASI-morris_larger_sample.png', dpi=300, bbox_inches='tight')
# +
diff = (disabled.loc[:, 'S1'] - enabled.loc[:, 'S1']).to_frame()
ax = diff.plot(title="EASI\nDisabled - Enabled",
legend=None,
marker='o',
markersize=4)
ax.set_xticks(diff.index)
ax.set_xticklabels(labels, rotation=45)
ax.set_xlabel("$N$");
plt.savefig(FIG_DIR+"EASI_diff_morris.png", dpi=300, bbox_inches='tight')
# -
| notebooks/5b EASI on Morris sampling results.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import seaborn as sns
from tqdm import tqdm
import matplotlib
import matplotlib.pyplot as plt
from wordcloud import WordCloud
import json
import codecs
import re
% matplotlib inline
matplotlib.rcParams['font.sans-serif'] = ["Whitney Book","simhei"]
data = []
with codecs.open('data/chinese_only.json', 'rb',encoding='utf-8') as f:
for line in f:
data.append(json.loads(line))
data[0]['genres']
# 返回指定director/cast 相关的电影(片名[0],评分)tuples
def getRatings(name=None,person_type=None):
'''
Return all of specified person's {MovieName:Rating} dict.
-----------------------------------------------------------
Input:
name: (str) director/cast name.
person_type: (str) 'directors'/'casts'
___________________________________________________________
Output:
PersonRating: (dict) {MovieName1: Rating1, MovieName2: Rating2, ...}
-----------------------------------------------------------
Example:
getRatings(name='黄渤',person_type='casts')
'''
PersonRating = {}
for rec in data:
if(rec['subtype']=='movie' and rec['ratings_count']>=10000 ):
for iperson in rec[person_type]:
if(iperson['name']==name):
PersonRating[rec['title']]=rec['rating']['average']
return PersonRating
# Distribution of the year of movie
for mtype in ['剧情','科幻','爱情','悬疑','动作']:
years = np.array([])
for rec in data:
if(rec['subtype']=='movie' and mtype in rec['genres']):
iyear = re.sub(r'\D','',str(rec['year']))[:4] # regex: filter first 4 numbers only (for case '2000年';'2008-2010年')
if(not iyear.strip()==''): # for case '不详'
rec['year'] = int(iyear)
years = np.append(years,int(iyear))
else:
rec['year'] = None
plt.figure(figsize=(8,6))
sns.distplot(years,kde=False)
plt.xlabel('电影年份',fontname='simhei',fontsize=12)
plt.ylabel('电影数量',fontname='simhei',fontsize=12)
plt.xticks(fontsize=12)
plt.yticks(fontsize=12)
plt.title('{0}片年份分布'.format(mtype),fontname='simhei',fontsize=15)
plt.xlim(xmax=2025)
plt.show()
# Distribution of the year of movie
years = np.array([])
for rec in data:
if(rec['subtype']=='movie'):
iyear = re.sub(r'\D','',str(rec['year']))[:4] # regex: filter first 4 numbers only (for case '2000年';'2008-2010年')
if(not iyear.strip()==''): # for case '不详'
rec['year'] = int(iyear)
years = np.append(years,int(iyear))
else:
rec['year'] = None
plt.figure(figsize=(8,6))
sns.distplot(years,kde=False)
plt.xlabel('电影年份',fontname='simhei',fontsize=12)
plt.ylabel('电影数量',fontname='simhei',fontsize=12)
plt.xticks(fontsize=12)
plt.yticks(fontsize=12)
plt.title('豆瓣电影数据:电影年份分布',fontname='simhei',fontsize=15)
plt.xlim(xmax=2025)
plt.savefig('豆瓣电影数据:电影年份分布.png',dpi=600,transparent=True)
plt.figure(figsize=(10,6))
sns.distplot(years[years>=1990],kde=False) # only for movie no sooner than 1990
plt.xlabel('电影年份',fontname='simhei',fontsize=12)
plt.ylabel('电影数量',fontname='simhei',fontsize=12)
plt.xticks(fontsize=12)
plt.yticks(fontsize=12)
plt.title('1990年及以后电影年份分布',fontname='simhei',fontsize=15)
plt.xlim(xmax=2025)
plt.savefig('1990年及以后电影年份分布.png',dpi=600,transparent=True)
# Distribution of movie ratings_count
ratecount = np.array([])
for rec in data:
if(rec['subtype']=='movie'):
ratecount = np.append(ratecount,rec['ratings_count'])
plt.figure(figsize=(4,3))
sns.distplot(ratecount[ratecount<10000],kde=False)
plt.xlabel('评分人数(人)',fontname='simhei',fontsize=10)
plt.ylabel('电影数量',fontname='simhei',fontsize=10)
plt.title('其中低于万人评分的电影',fontname='simhei',fontsize=12)
plt.savefig('低于万人评分人数分布.png',dpi=600,transparent=True)
# +
plt.figure(figsize=(8,6))
sns.distplot(ratecount[ratecount>10000],kde=False)
plt.xlabel('评分人数(人)',fontname='simhei',fontsize=12)
plt.ylabel('电影数量',fontname='simhei',fontsize=12)
plt.title('豆瓣电影数据:评分人数分布',fontname='simhei',fontsize=15)
plt.xticks(fontsize=12)
plt.yticks(fontsize=12)
plt.savefig('豆瓣电影数据:评分人数分布.png',dpi=600,transparent=True)
# -
# Distribution of movie ratings
rates = np.array([])
for rec in data:
if(rec['subtype']=='movie'):# and rec['ratings_count']>=10000):
rates = np.append(rates,rec['rating']['average'])
# +
plt.figure(figsize=(8,6))
sns.distplot(rates,kde=False)
plt.xlabel('用户评分',fontname='simhei',fontsize=12)
plt.ylabel('电影数量',fontname='simhei',fontsize=12)
plt.title('豆瓣电影数据:用户评分分布',fontname='simhei',fontsize=15)
plt.xticks(np.arange(0,11,1),fontsize=12)
plt.yticks(fontsize=12)
plt.xlim(-0.1,10.1)
plt.savefig('豆瓣电影数据:用户评分分布.png',dpi=600,transparent=True)
# +
plt.figure(figsize=(8,6))
sns.distplot(rates[rates>0],kde=False)
plt.xlabel('用户评分',fontname='simhei',fontsize=12)
plt.ylabel('电影数量',fontname='simhei',fontsize=12)
plt.title('豆瓣电影数据:用户评分分布',fontname='simhei',fontsize=15)
plt.xticks(np.arange(0,11,1),fontsize=12)
plt.yticks(fontsize=12)
plt.xlim(-0.1,10.1)
plt.savefig('用户评分(大于0分)分布.png',dpi=600,transparent=True)
# -
numcount = {}
totalrate = {}
for rec in data:
if(rec['subtype']=='movie'):
for mvtype in rec['genres']:
if numcount.get(mvtype) is None:
# Add a new type into dict
numcount[mvtype] = 1
totalrate[mvtype] = rec['rating']['average']
else:
# Count of this type ++
numcount[mvtype] += 1
totalrate[mvtype] += rec['rating']['average']
popcount = {}
totalpoprate = {}
for rec in data:
if(rec['subtype']=='movie' and rec['ratings_count']>=10000):
for mvtype in rec['genres']:
if popcount.get(mvtype) is None:
popcount[mvtype] = 1
totalpoprate[mvtype] = rec['rating']['average']
else:
popcount[mvtype] += 1
totalpoprate[mvtype] += rec['rating']['average']
df = pd.DataFrame.from_dict(numcount,orient='index')
df.columns = ['count']
tmp = pd.DataFrame.from_dict(totalrate,orient='index')
df.insert(1,'totalrate',tmp)
df.insert(2,'avgrate',df['totalrate']/df['count'])
NumTop10Type = df.sort_values('count',ascending=False)[:10]
RateTop10Type = df.sort_values('avgrate',ascending=False)[:10]
pop = pd.DataFrame.from_dict(popcount,orient='index')
pop.columns = ['count']
tmp = pd.DataFrame.from_dict(totalpoprate,orient='index')
pop.insert(1,'totalpoprate',tmp)
pop.insert(2,'avgpoprate',pop['totalpoprate']/pop['count'])
PopNumTop10Type = pop.sort_values('count',ascending=False)[:10]
PopRateTop10Type = pop.sort_values('avgpoprate',ascending=False)[:10]
plt.figure(figsize=(12,6))
sns.barplot(x=NumTop10Type.index,y='count',data=NumTop10Type,palette='coolwarm_r')
plt.xticks(fontname='simhei',fontsize=12)
plt.yticks(fontsize=12)
#plt.xlabel('电影类型',fontname='simhei',fontsize=15)
plt.ylabel('电影数量',fontname='simhei',fontsize=15)
plt.title('数量排名前十的电影类型',fontname='simhei',fontsize=16)
plt.tight_layout()
plt.savefig('数量排名前十的电影类型.png',dpi=600,transparent=True)
plt.figure(figsize=(12,6))
sns.barplot(x=RateTop10Type.index,y='avgrate',data=RateTop10Type,palette='coolwarm_r')
plt.xticks(fontname='simhei',fontsize=12)
plt.yticks(fontsize=12)
#plt.xlabel('电影类型',fontname='simhei',fontsize=15)
plt.ylabel('平均用户评分',fontname='simhei',fontsize=15)
plt.title('评分排名前十的电影类型',fontname='simhei',fontsize=16)
plt.tight_layout()
plt.savefig('全部电影中评分排名前十的电影类型.png',dpi=600,transparent=True)
plt.figure(figsize=(12,6))
sns.barplot(x=PopRateTop10Type.index,y='avgpoprate',data=PopRateTop10Type,palette='coolwarm_r')
plt.xticks(fontname='simhei',fontsize=12)
plt.yticks(fontsize=12)
#plt.xlabel('电影类型',fontname='simhei',fontsize=15)
plt.ylabel('平均用户评分',fontname='simhei',fontsize=15)
plt.title('逾万人评分电影中评分排名前十的电影类型',fontname='simhei',fontsize=16)
plt.tight_layout()
plt.savefig('逾万人评分电影中评分排名前十的电影类型.png',dpi=600,transparent=True)
total_mv_haverate = len(rates[rates>0])
sorted_rates = sorted(rates[rates>0])
top20perc_rate = sorted_rates[int(0.8*total_mv_haverate)]
bot40perc_rate = sorted_rates[int(0.4*total_mv_haverate)]
def getPeriodRatings(startyr = None, endyr = None):
'''
Get the ratings of movies sorted by casts/directors
FILTERED BY YEAR PERIOD
---------------------------------------------------
INPUT:
startyr: int, start year of filter period
endyr: int, end year of filter period
(filtered year is in [startyr,endyr])
---------------------------------------------------
OUTPUT: cast_rating_df; dir_rating_df
cast_rating_df: pandas DataFrame
field: rating_mean;
rating_std;
great_count;
bad_count
dir_rating_df: pandas DataFrame
field: rating_mean;
rating_std;
great_count;
bad_count
'''
cast_ratings = {}
director_ratings = {}
for rec in data:
if(rec['subtype']=='movie' and rec['ratings_count']>=10000):
if(rec['year'] in range(startyr,endyr+1)):
for icast in rec['casts']:
if(cast_ratings.get(icast['name']) is None):
cast_ratings[icast['name']] = np.array([])
cast_ratings[icast['name']] = np.append(cast_ratings[icast['name']],rec['rating']['average'])
for idir in rec['directors']:
if(director_ratings.get(idir['name']) is None):
director_ratings[idir['name']] = np.array([])
director_ratings[idir['name']] = np.append(director_ratings[idir['name']],rec['rating']['average'])
# Find the variability of casts'/directors' performance (in rating of movies)
cast_rating_mean = {}
cast_rating_std = {}
for icast in cast_ratings:
cast_rating_mean[icast] = np.mean(cast_ratings[icast])
cast_rating_std[icast] = np.std(cast_ratings[icast])
dir_rating_mean = {}
dir_rating_std = {}
for idir in director_ratings:
dir_rating_mean[idir] = np.mean(director_ratings[idir])
dir_rating_std[idir] = np.std(director_ratings[idir])
# stat of casts/directors involved in great/bad movies
# definition of a great movie:rating above top 10% (top10perc_rate = 8.0)
# definition of a bad movie:rating below bottom 40% (bot40perc_rate = 6.0)
cast_great_count = {}
cast_bad_count = {}
dir_great_count = {}
dir_bad_count = {}
for rec in data:
if(rec['subtype']=='movie' and rec['ratings_count']>=10000 and (rec['year'] in range(startyr,endyr+1))):
if(rec['rating']['average']>=top20perc_rate):
for icast in rec['casts']:
if(cast_great_count.get(icast['name']) is None):
cast_great_count[icast['name']] = 1
else:
cast_great_count[icast['name']] += 1
for idir in rec['directors']:
if(dir_great_count.get(idir['name']) is None):
dir_great_count[idir['name']] = 1
else:
dir_great_count[idir['name']] += 1
if(rec['rating']['average']<=bot40perc_rate):
for icast in rec['casts']:
if(cast_bad_count.get(icast['name']) is None):
cast_bad_count[icast['name']] = 1
else:
cast_bad_count[icast['name']] += 1
for idir in rec['directors']:
if(dir_bad_count.get(idir['name']) is None):
dir_bad_count[idir['name']] = 1
else:
dir_bad_count[idir['name']] += 1
cast_rating_df = pd.DataFrame.from_dict(cast_rating_mean,orient='index')
cast_rating_df.columns=['rating_mean']
tmp = pd.DataFrame.from_dict(cast_rating_std,orient='index')
cast_rating_df.insert(1,'rating_std',tmp)
tmp = pd.DataFrame.from_dict(cast_great_count,orient='index')
cast_rating_df.insert(2,'great_count',tmp)
tmp = pd.DataFrame.from_dict(cast_bad_count,orient='index')
cast_rating_df.insert(3,'bad_count',tmp)
no_bad = cast_rating_df['bad_count'].isnull()
cast_rating_df.loc[no_bad,'bad_count']=0
no_great = cast_rating_df['great_count'].isnull()
cast_rating_df.loc[no_great,'great_count']=0
cast_compound = (cast_rating_df.rating_mean*cast_rating_df.great_count
- cast_rating_df.rating_std*cast_rating_df.bad_count/(cast_rating_df.great_count+cast_rating_df.bad_count))
cast_rating_df.insert(4,'compound',cast_compound)
dir_rating_df = pd.DataFrame.from_dict(dir_rating_mean,orient='index')
dir_rating_df.columns=['rating_mean']
tmp = pd.DataFrame.from_dict(dir_rating_std,orient='index')
dir_rating_df.insert(1,'rating_std',tmp)
tmp = pd.DataFrame.from_dict(dir_great_count,orient='index')
dir_rating_df.insert(2,'great_count',tmp)
tmp = pd.DataFrame.from_dict(dir_bad_count,orient='index')
dir_rating_df.insert(3,'bad_count',tmp)
no_bad = dir_rating_df['bad_count'].isnull()
dir_rating_df.loc[no_bad,'bad_count']=0
no_great = dir_rating_df['great_count'].isnull()
dir_rating_df.loc[no_great,'great_count']=0
dir_compound = (dir_rating_df.rating_mean*dir_rating_df.great_count
- dir_rating_df.rating_std*dir_rating_df.bad_count/(dir_rating_df.great_count+dir_rating_df.bad_count))
dir_rating_df.insert(4,'compound',dir_compound)
return cast_rating_df, dir_rating_df
# +
cast_rating_df_90_00, dir_rating_df_90_00 = getPeriodRatings(startyr=1990,endyr=2000)
cast_rating_df_01_10, dir_rating_df_01_10 = getPeriodRatings(startyr=2001,endyr=2010)
cast_rating_df_11_18, dir_rating_df_11_18 = getPeriodRatings(startyr=2011,endyr=2018)
cast_top_rating_90_00 = cast_rating_df_90_00.sort_values(['compound','rating_mean'],ascending=False)
cast_top_rating_01_10 = cast_rating_df_01_10.sort_values(['compound','rating_mean'],ascending=False)
cast_top_rating_11_18 = cast_rating_df_11_18.sort_values(['compound','rating_mean'],ascending=False)
dir_top_rating_90_00 = dir_rating_df_90_00.sort_values(['compound','rating_mean'],ascending=False)
dir_top_rating_01_10 = dir_rating_df_01_10.sort_values(['compound','rating_mean'],ascending=False)
dir_top_rating_11_18 = dir_rating_df_11_18.sort_values(['compound','rating_mean'],ascending=False)
# -
# Draw Top 10 casts/directors
def DrawTop10(title=None,data=None):
plt.figure(figsize=(6,8))
sns.barplot(y=data.index[:10],x='compound',palette='summer',data=data[:10],orient='h')
plt.yticks(fontname='simhei',fontsize=15)
plt.xlabel('综合得分',fontname='simhei',fontsize='15')
plt.xticks(fontsize=15)
plt.savefig(title+'.png',dpi=600,transparent=True)
plt.close()
DrawTop10(title='90年代Top10演员',data=cast_top_rating_90_00)
DrawTop10(title='00年代Top10演员',data=cast_top_rating_01_10)
DrawTop10(title='10年代Top10演员',data=cast_top_rating_11_18)
DrawTop10(title='90年代Top10导演',data=dir_top_rating_90_00)
DrawTop10(title='00年代Top10导演',data=dir_top_rating_01_10)
DrawTop10(title='10年代Top10导演',data=dir_top_rating_11_18)
from wordcloud import WordCloud
def GenerateWordCloud(title=None,data=None,key=None,N_person=20):
text=''
for idf in data.index[:N_person]:
text += int(np.ceil(data.loc[idf,key]))*(idf+' ')
font = r'C:\Windows\Fonts\simhei.ttf'
wc = WordCloud(collocations=False, colormap= 'summer', min_font_size=8, background_color = 'white',font_path=font, width=3500, height=2000, margin=50).generate(text.lower())
plt.axis("off")
wc.to_file(title+'.png')
GenerateWordCloud(title='90年代Top20演员',data=cast_top_rating_90_00,key='compound',N_person=20)
GenerateWordCloud(title='00年代Top20演员',data=cast_top_rating_01_10,key='compound',N_person=20)
GenerateWordCloud(title='10年代Top20演员',data=cast_top_rating_11_18,key='compound',N_person=20)
GenerateWordCloud(title='90年代Top20导演',data=dir_top_rating_90_00,key='compound',N_person=20)
GenerateWordCloud(title='00年代Top20导演',data=dir_top_rating_01_10,key='compound',N_person=20)
GenerateWordCloud(title='10年代Top20导演',data=dir_top_rating_11_18,key='compound',N_person=20)
| 1_DoubanMovieRanking(Enrollment)/GoodActors_Directors.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction
# State notebook purpose here
# ### Imports
# Import libraries and write settings here.
import datashader as ds
# +
from sqlalchemy import create_engine
engine = create_engine('postgres://localhost:5432/global_fishing_watch')
engine.table_names()
# +
df = pd.read_sql("SELECT * FROM fishing_effort LIMIT 100000",
engine, parse_dates=['date'])
df['lon'] = df['lon_bin'] / 100
df['lat'] = df['lat_bin'] / 100
df.head()
# +
from datashader.utils import lnglat_to_meters
import datashader.transfer_functions as tf
df['x'], df['y'] = lnglat_to_meters(df['lon'], df['lat'])
df.head()
# -
cvs = ds.Canvas(plot_width=1000, plot_height=500)
agg = cvs.points(df, 'x', 'y', ds.mean('fishing_hours'))
img = tf.shade(agg, cmap=['lightblue', 'darkblue'], how='log')
img
agg
# +
bound = 20026376.39
bounds = dict(x_range = (-bound, bound), y_range = (int(-bound*0.4), int(bound*0.6)))
plot_width = 1000
plot_height = int(plot_width*0.5)
cvs = ds.Canvas(plot_width=plot_width, plot_height=plot_height, **bounds)
# -
tf.shade(agg, cmap=['red'])
s = 'string'
# +
# Data manipulation
import pandas as pd
import numpy as np
# Options for pandas
pd.options.display.max_columns = 50
pd.options.display.max_rows = 30
# Display all cell outputs
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = 'all'
from IPython import get_ipython
ipython = get_ipython()
# autoreload extension
if 'autoreload' not in ipython.extension_manager.loaded:
# %load_ext autoreload
# %autoreload 2
# Visualizations
import plotly.plotly as py
import plotly.graph_objs as go
from plotly.offline import iplot, init_notebook_mode
init_notebook_mode(connected=True)
import cufflinks as cf
cf.go_offline(connected=True)
cf.set_config_file(theme='white')
# -
# # Analysis/Modeling
# Do work here
# # Results
# Show graphs and stats here
# # Conclusions and Next Steps
# Summarize findings here
| datashader-work/datashader-tryout.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import matplotlib
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import math
from sklearn.preprocessing import MinMaxScaler
import seaborn as sns
import datetime
linestyles = ['-', '--', '-.', ':']
# -
def visualizeCorrelationPlot(dc, title=""):
fig = plt.figure()
plt.figure(figsize=(20, 20))
ax = fig.add_subplot(111)
cax = ax.matshow(dc, vmin=-1, vmax=1)
fig.colorbar(cax)
ticks = np.arange(0,len(dc.columns),1)
ax.set_yticks(ticks)
ticks = np.arange(0,len(dc.columns),1)
ax.set_xticks(ticks)
ax.set_yticklabels(dc.columns)
ax.set_xticklabels(dc.columns, rotation="vertical")
ax.set_title(title)
plt.show()
# <br><br><br>
# # Raw Traffic and Rolling & Expanding Traffic Visualization
rolling_expanding_bar = []
def getNeededFeatures(columns, arrFeaturesNeed=[], featureEngineering="Original"):
to_remove = []
if len(arrFeaturesNeed) == 0: #all features aren't needed
return []
else:
if featureEngineering == "Original":
compareTo = " "
elif featureEngineering == "Rolling" or featureEngineering == "Expanding":
compareTo = "_"
for f in arrFeaturesNeed:
for c in range(0, len(columns)):
if (len(columns[c].split(compareTo)) <= 1 and featureEngineering != "Original"):
to_remove.append(c)
if f not in columns[c].split(compareTo)[0] and columns[c].split(compareTo)[0] not in arrFeaturesNeed:
to_remove.append(c)
if len(columns[c].split(compareTo)) > 1:
if "Esum" in columns[c].split(compareTo)[1]: #Removing all Expanding Sum
to_remove.append(c)
if "min" in columns[c].split(compareTo)[1]: #Removing all Expanding Sum
to_remove.append(c)
if "max" in columns[c].split(compareTo)[1]: #Removing all Expanding Sum
to_remove.append(c)
return to_remove
# +
ROAD = "<NAME>"
YEAR = "2015"
EXT = ".csv"
DATASET_DIVISION = "seasonWet"
DIR = "../../../../datasets/Thesis Datasets/"
ROLLING_WINDOW = 8
EXPANDING_WINDOW = 8
####################################### ORIG #######################################
TRAFFIC_WINDOWSIZE = 3
TRAFFIC_DIR = DIR + "mmda/"
TRAFFIC_FILENAME = "mmda_" + ROAD + "_" + YEAR + "_" + DATASET_DIVISION
orig_traffic = pd.read_csv(TRAFFIC_DIR + TRAFFIC_FILENAME + EXT, skipinitialspace=True)
orig_traffic = orig_traffic.fillna(0)
#Converting index to date and time, and removing 'dt' column
orig_traffic.index = pd.to_datetime(orig_traffic.dt, format='%d/%m/%Y %H:%M')
cols_to_remove = [0]
cols_to_remove = getNeededFeatures(orig_traffic.columns, ["statusN"])
orig_traffic.drop(orig_traffic.columns[[cols_to_remove]], axis=1, inplace=True)
orig_traffic.head()
####################################################################################
##################################### ROLLING #####################################
TRAFFIC_DIR = DIR + "mmda/Rolling/" + DATASET_DIVISION + "/"
TRAFFIC_FILENAME = "eng_win" + str(ROLLING_WINDOW) + "_mmda_" + ROAD + "_" + YEAR + "_" + DATASET_DIVISION
rolling_traffic = pd.read_csv(TRAFFIC_DIR + TRAFFIC_FILENAME + EXT, skipinitialspace=True)
cols_to_remove = [0, 1, 2]
cols_to_remove += getNeededFeatures(rolling_traffic.columns, ["statusN"], "Rolling")
rolling_traffic.index = pd.to_datetime(rolling_traffic.dt, format='%Y-%m-%d %H:%M')
rolling_traffic.drop(rolling_traffic.columns[[cols_to_remove]], axis=1, inplace=True)
rolling_traffic.head()
####################################################################################
##################################### EXPANDING #####################################
TRAFFIC_DIR = DIR + "mmda/Expanding/" + DATASET_DIVISION + "/"
TRAFFIC_FILENAME = "eng_win" + str(EXPANDING_WINDOW) + "_mmda_" + ROAD + "_" + YEAR + "_" + DATASET_DIVISION
expanding_traffic = pd.read_csv(TRAFFIC_DIR + TRAFFIC_FILENAME + EXT, skipinitialspace=True)
cols_to_remove = [0, 1, 2, 5]
cols_to_remove += getNeededFeatures(expanding_traffic.columns, ["statusN"], "Rolling")
expanding_traffic.index = pd.to_datetime(expanding_traffic.dt, format='%d/%m/%Y %H:%M')
expanding_traffic.drop(expanding_traffic.columns[[cols_to_remove]], axis=1, inplace=True)
####################################################################################
linestyles = ['-', '--', '-.', ':']
orig = orig_traffic[orig_traffic.columns[0]]
rolling = rolling_traffic[rolling_traffic.columns[0]]
expanding = expanding_traffic[expanding_traffic.columns[0]]
startIndex = 0
endIndex = 192
plt.plot(orig, color='gray', label="Original Traffic", linestyle=':')
plt.plot(rolling, color='r', alpha=0.7, label="Rolling \nTraffic \nat window " + str(ROLLING_WINDOW))
plt.plot(expanding, color='b', alpha=0.7, label='Expanding \nTraffic \nat window' + str(EXPANDING_WINDOW))
plt.legend(loc=1)
datemin = datetime.date(2015, 10, 12)
datemax = datetime.date(2015, 10, 14)
plt.xlim([datemin, datemax])
plt.xticks(rotation='45')
plt.xlabel("Date")
plt.yticks([0, 0.625, 1], ["H", "M", "L"])
plt.ylabel("Congestion Level")
plt.grid()
plt.show()
# -
# <br><br><br>
# # Correlation 'growth' of Rolling with Current Traffic
# +
merged_traffic = pd.concat([orig_traffic,
orig_traffic.shift(1).rolling(2).mean(),
orig_traffic.shift(2).rolling(3).mean(),
orig_traffic.shift(3).rolling(4).mean(),
orig_traffic.shift(7).rolling(8).mean(),
orig_traffic.shift(11).rolling(12).mean(),
orig_traffic.shift(23).rolling(24).mean(),
orig_traffic.shift(47).rolling(48).mean(),
orig_traffic.shift(95).rolling(96).mean(),
], axis=1)
merged = merged_traffic.corr(method='spearman').iloc[:, 0].values
rolling_expanding_bar.append(merged)
plt.bar(np.arange(len(merged)), merged, align='center', alpha=0.7, color=['r', 'g', 'b'])
windows = [1, 2, 3, 4, 8, 12, 24, 48, 96]
plt.xticks(np.arange(len(windows)), windows)
plt.xlabel("Windows")
plt.ylabel("Correlation Coefficient")
plt.title("Correlation between Rolling with Current Northbound Traffic")
# -
# <br><br><br>
# # Correlation 'growth' of Expanding with Current Traffic
# +
windows = [2, 3, 4, 8, 12, 24, 48, 96]
expanding_traffic_list= []
for w in windows:
EXPANDING_WINDOW = w
##################################### EXPANDING #####################################
TRAFFIC_DIR = DIR + "mmda/Expanding/" + DATASET_DIVISION + "/"
TRAFFIC_FILENAME = "eng_win" + str(EXPANDING_WINDOW) + "_mmda_" + ROAD + "_" + YEAR + "_" + DATASET_DIVISION
expanding_traffic = pd.read_csv(TRAFFIC_DIR + TRAFFIC_FILENAME + EXT, skipinitialspace=True)
cols_to_remove = [0, 1, 2, 5]
cols_to_remove += getNeededFeatures(expanding_traffic.columns, ["statusN"], "Rolling")
expanding_traffic.index = pd.to_datetime(expanding_traffic.dt, format='%d/%m/%Y %H:%M')
expanding_traffic.drop(expanding_traffic.columns[[cols_to_remove]], axis=1, inplace=True)
####################################################################################
temp = pd.DataFrame(data=expanding_traffic.iloc[:,0].values, columns=['E'+str(w)])
expanding_traffic_list.append(temp)
#Adding orig traffic
expanding_traffic_list.insert(0, pd.DataFrame(data=orig_traffic.iloc[:, 0].values, columns=['orig']))
expanding_traffic_list
merged = pd.concat(expanding_traffic_list, axis=1)
merged_expanding = merged
merged_corr = merged.corr(method='spearman').iloc[:, 0].values
rolling_expanding_bar.append(merged_corr)
plt.bar(np.arange(len(merged_corr)), merged_corr, align='center', alpha=0.7, color=['r', 'g', 'b'])
plt.xticks(np.arange(len(windows)), windows)
plt.xlabel("Windows")
plt.ylabel("Correlation Coefficient")
plt.title("Correlation between Expanding with Current Northbound Traffic")
visualizeCorrelationPlot(merged.corr(method='spearman'), title="")
merged.corr(method='spearman')
# +
windows = [1, 2, 3, 4, 8, 12, 24, 48, 96]
rolling = rolling_expanding_bar[0]
expanding = rolling_expanding_bar[1]
ind = np.arange(len(rolling)) # the x locations for the groups
width = 0.35 # the width of the bars
fig, ax = plt.subplots()
rects1 = ax.bar(ind, rolling, width, color='g')
rects2 = ax.bar(ind + width, expanding, width, color='y')
# add some text for labels, title and axes ticks
ax.set_ylabel('Correlation')
ax.set_xlabel('Windows')
# ax.set_title('Correlation of Expanding and Rolling Traffic to Current Traffic by window')
ax.set_xticks(ind + width / 2)
ax.set_xticklabels((windows))
ax.legend((rects1[0], rects2[0]),
('Rolling',
'Expanding '))
plt.show()
print(rolling_expanding_bar[0])
print(rolling_expanding_bar[1])
# -
# <br><br><br>
# # Correlating Rolling and Expanding Traffic
# +
windows = [2, 3, 4, 8, 12, 24, 48, 96]
expanding_traffic_list= []
rolling_traffic_list= []
for w in windows:
EXPANDING_WINDOW = w
##################################### EXPANDING #####################################
TRAFFIC_DIR = DIR + "mmda/Expanding/" + DATASET_DIVISION + "/"
TRAFFIC_FILENAME = "eng_win" + str(EXPANDING_WINDOW) + "_mmda_" + ROAD + "_" + YEAR + "_" + DATASET_DIVISION
expanding_traffic = pd.read_csv(TRAFFIC_DIR + TRAFFIC_FILENAME + EXT, skipinitialspace=True)
cols_to_remove = [0, 1, 2, 5]
cols_to_remove += getNeededFeatures(expanding_traffic.columns, ["statusN"], "Rolling")
expanding_traffic.index = pd.to_datetime(expanding_traffic.dt, format='%d/%m/%Y %H:%M')
expanding_traffic.drop(expanding_traffic.columns[[cols_to_remove]], axis=1, inplace=True)
####################################################################################
temp = pd.DataFrame(data=expanding_traffic.iloc[:,0].values, columns=['E'+str(w)])
expanding_traffic_list.append(temp)
ROLLING_WINDOW = w
##################################### ROLLING #####################################
TRAFFIC_DIR = DIR + "mmda/Rolling/" + DATASET_DIVISION + "/"
TRAFFIC_FILENAME = "eng_win" + str(ROLLING_WINDOW) + "_mmda_" + ROAD + "_" + YEAR + "_" + DATASET_DIVISION
rolling_traffic = pd.read_csv(TRAFFIC_DIR + TRAFFIC_FILENAME + EXT, skipinitialspace=True)
cols_to_remove = [0, 1, 2]
cols_to_remove += getNeededFeatures(rolling_traffic.columns, ["statusN"], "Rolling")
rolling_traffic.index = pd.to_datetime(rolling_traffic.dt, format='%Y-%m-%d %H:%M')
rolling_traffic.drop(rolling_traffic.columns[[cols_to_remove]], axis=1, inplace=True)
####################################################################################
temp = pd.DataFrame(data=rolling_traffic.iloc[:,0].values, columns=['R'+str(w)])
rolling_traffic_list.append(temp)
merged_expanding_corr = pd.concat(expanding_traffic_list, axis=1)
merged_rolling_corr = pd.concat(rolling_traffic_list, axis=1)
merged_corr = pd.concat([merged_expanding_corr, merged_rolling_corr], axis=1)
#Array to Dataframe
merged_corr = merged_corr.corr(method='spearman')
visualizeCorrelationPlot(merged_corr, title="Correlation between Expanding and Rolling windows of Traffic at different windows\n\n")
merged_corr
# -
# <br><br><br>
# # PM1 Yesterday - Today FE + Connected Roads
# +
#Experiments Comparison for all windows
line1 = [0.084, 0.110, 0.138, 0.218, 0.227, 0.259, 0.260, 0.249] #PM1 - Yesterday - Weekday
line2 = [0.066, 0.097, 0.125, 0.205, 0.229, 0.247, 0.260, 0.244] #PM1 - Yesterday - WeekdayWeekends
line3 = [0.045, 0.036, 0.043, 0.049, 0.036, 0.040, 0.043, 0.041] #PM1 - Today - Weekday
line4 = [0.038, 0.032, 0.038, 0.039, 0.036, 0.036, 0.034, 0.032] #PM1 - Today - WeekdayWeekends
line5 = [0.069, 0.096, 0.124, 0.202, 0.225, 0.267, 0.266, 0.242] #PM1 - Yesterday - Connected Roads - Weekday
line6 = [0.069, 0.091, 0.111, 0.168, 0.186, 0.202, 0.194, 0.200] #PM1 - Yesterday - Connected Roads - WeekdayWeekends
windows = [2, 3, 4, 8, 12, 24, 48, 96]
plt.figure(figsize=(15, 4))
plt.title("PM1 Current - W/WO WeekEnds - With R&E")
plt.plot(line3, label="PM1-Today-Day")
plt.plot(line4, label="PM1-Today-DayEnds")
plt.legend()
plt.xticks(range(len(windows)), (windows))
plt.figure(figsize=(15, 4))
plt.title("PM1 Past - W/WO WeekEnds - With R&E")
plt.plot(line1, label="PM1-Yesterday-Day")
plt.plot(line2, label="PM1-Yesterday-DayEnds")
plt.legend()
plt.xticks(range(len(windows)), (windows))
plt.figure(figsize=(15, 4))
plt.title("PM1 Current/Past - W/WO WeekEnds - With R&E - Connected")
plt.plot(line5, label="PM1-Yesterday-DayEnds-Connected")
plt.plot(line6, label="PM1-Today-DayEnds-Connected")
plt.legend()
plt.xticks(range(len(windows)), (windows))
plt.figure(figsize=(15, 4))
plt.title("All PM1 Runs")
plt.plot(line3, label="PM1-Today-Day")
plt.plot(line4, label="PM1-Today-DayEnds")
plt.plot(line1, label="PM1-Yesterday-Day")
plt.plot(line2, label="PM1-Yesterday-DayEnds")
plt.plot(line5, label="PM1-Yesterday-DayEnds-Connected")
plt.plot(line6, label="PM1-Today-DayEnds-Connected")
plt.legend()
plt.xticks(range(len(windows)), (windows))
plt.show()
# -
# <br><br><br>
# # PM1 Windows
# ### PM1 Windows without WP
# +
windows = [2, 3, 4, 8, 12, 24, 48, 96]
pm1_rolling = [0.056, 0.105, 0.145, 0.211, 0.23, 0.26, 0.27, 0.253]
pm1_expanding = [0.013, 0.024, 0.034, 0.067, 0.088, 0.124, 0.176, 0.229]
pm1_rolling_wp = [0.063, 0.11, 0.145, 0.21, 0.217, 0.245, 0.257, 0.242]
pm1_expanding_wp = [0.023, 0.03, 0.039, 0.069, 0.087, 0.121, 0.172, 0.205]
ind = np.arange(len(pm1_rolling)) # the x locations for the groups
width = 0.35 # the width of the bars
fig, ax = plt.subplots(figsize=(17,4))
rects1 = ax.bar(ind, pm1_rolling, width, color='r')
ax.plot(pm1_rolling, color='r', alpha=0.4)
rects2 = ax.bar(ind + width, pm1_expanding, width, color='y')
ax.plot(pm1_expanding, color='y', alpha=0.4)
# add some text for labels, title and axes ticks
ax.set_ylabel('RMSE')
ax.set_xlabel('Windows')
ax.set_title('Error comparison of TOM for Rolling and Expanding Window Combinations with consideration of Work Day and Peak Hours')
ax.set_xticks(ind + width / 2)
ax.set_xticklabels((windows))
ax.legend((rects1[0], rects2[0]),
('Rolling',
'Expanding '))
plt.show()
# -
# ### PM1 Windows with WP and Rolling + Expanding
# +
windows = [2, 3, 4, 8, 12, 24, 48, 96]
pm1_rolling = [0.056, 0.105, 0.145, 0.211, 0.23, 0.26, 0.27, 0.253]
pm1_expanding = [0.013, 0.024, 0.034, 0.067, 0.088, 0.124, 0.176, 0.229]
pm1_roll_expanding = [0.038, 0.031, 0.034, 0.066, 0.082, 0.120, 0.177, 0.245]
ind = np.arange(len(pm1_rolling_wp)) # the x locations for the groups
width = 0.20 # the width of the bars
fig, ax = plt.subplots(figsize=(7,4))
rects1 = ax.bar(ind, pm1_rolling, width, color='r')
# ax.plot(pm1_rolling, color='r', alpha=0.4)
rects2 = ax.bar(ind + width, pm1_expanding, width, color='y')
# ax.plot(pm1_expanding, color='y', alpha=0.4)
rects3 = ax.bar(ind + width + width, pm1_roll_expanding, width, color='orange')
# ax.plot(pm1_roll_expanding, color='orange', alpha=0.4)
# add some text for labels, title and axes ticks
ax.set_ylabel('RMSE')
ax.set_xlabel('Windows')
ax.set_title('Error comparison of TOM for Rolling and Expanding Window Combinations\n with consideration of Work Day and Peak Hours')
ax.set_xticks(ind + width / 2)
ax.set_xticklabels((windows))
ax.legend((rects1[0], rects2[0], rects3[0]), ('Rolling', 'Expanding', 'Rolling and Expanding'))
plt.show()
# -
# <br><br><br>
# # PM1 Best Features
# +
#PM1 Best Features (Original, WP, WP Roll, WP Exp, WP Roll & Exp)
rmse = [0.260, 0.248, 0.257, 0.172, 0.177]
datasets = ['Original',
'Work day \n + Peak hour',
'Work day \n + Peak hour \n+ Rolling',
'Work day \n + Peak hour \n+ Expanding',
'Work day \n + Peak hour \n+ Rolling \n+ Expanding']
plt.figure(figsize=(10, 5))
plt.margins(0.05, 0.25)
plt.bar(datasets, rmse, align='center', alpha=0.7, color=['r', 'g', 'b'])
for a, b in zip(datasets, rmse):
lbl = str(b)
if str(b * 100).split(".")[1] == '0':
lbl = lbl + "0"
plt.text(a, b, lbl+"\n")
# plt.title("Error Comparison of TOM for Different Setups")
plt.xlabel("\nFeature Combinations")
plt.ylabel("RMSE")
fig.tight_layout()
plt.show()
# -
# <br><br><br>
# # PM2 Windows
# +
windows = [2, 3, 4, 8, 12, 24, 48, 96]
pm2_rolling = [0.251, 0.257, 0.257, 0.257, 0.262, 0.265, 0.263, 0.266]
pm2_expanding = [0.253, 0.251, 0.259, 0.263, 0.256, 0.259, 0.269, 0.254]
ind = np.arange(len(pm1_rolling_wp)) # the x locations for the groups
width = 0.35 # the width of the bars
fig, ax = plt.subplots(figsize=(10,4))
rects1 = ax.bar(ind, pm2_rolling, width, color='r')
ax.plot(pm2_rolling, color='r', alpha=0.4)
rects2 = ax.bar(ind + width, pm2_expanding, width, color='y')
ax.plot(pm2_expanding, color='y', alpha=0.4)
# add some text for labels, title and axes ticks
ax.set_ylabel('RMSE')
ax.set_xlabel('Windows')
ax.set_title('Performance of WOM for Rolling and Expanding window combinations')
ax.set_xticks(ind + width / 2)
ax.set_xticklabels((windows))
ax.legend((rects1[0], rects2[0]), ('Rolling', 'Expanding'))
plt.margins(0.03, 0.3)
plt.show()
# -
# <br><br><br>
# # PM2 Best Features
# +
#PM2 Best Features (Original, Correlated, Rolling, Expanding, Correlated with Rollign and Expanding)
rmse = [0.238, 0.279, 0.257, 0.259, 0.267]
datasets = ['Original',
'Correlated \n Variables',
'Correlated \n Variables \n + Rolling',
'Correlated \n Variables \n + Expanding',
'Correlated \n Variables \n + Rolling \n + Expanding']
plt.figure(figsize=(10, 5))
plt.margins(0.05, 0.25)
plt.bar(datasets, rmse, align='center', alpha=0.7, color=['r', 'g', 'b'], width=0.7)
for a, b in zip(datasets, rmse):
lbl = str(b)
if str(b * 100).split(".")[1] == '0':
lbl = lbl + "0"
plt.text(a, b, lbl+"\n")
plt.xlabel("\nFeature Combinations")
plt.ylabel("RMSE")
fig.tight_layout()
plt.show()
# -
# <br><br><br>
# # Fusion: Feature vs Decision for All - Best Datasets
# +
datasets = ['Original', 'With Feature Selection \nand Engineering']
ff_rmse = [0.246, 0.186]
df_rmse = [0.238, 0.175]
# improvement
# FF = (0.246 - 0.238)/.246
ind = np.arange(len(ff_rmse)) # the x locations for the groups
width = 0.35 # the width of the bars
fig, ax = plt.subplots(figsize=(7,6))
rects1 = ax.bar(ind, ff_rmse, width, color='r')
# ax.plot(ff_rmse, color='r', alpha=0.4)
rects2 = ax.bar(ind + width, df_rmse, width, color='y')
# ax.plot(df_rmse, color='y', alpha=0.4)
# add some text for labels, title and axes ticks
ax.set_ylabel('RMSE')
ax.set_xlabel('Features')
ax.set_title('Error Comparison of Weather-Traffic Fusion in feature and decision level')
ax.set_xticks(ind + width / 2)
ax.set_xticklabels((datasets))
ax.legend((rects1[0], rects2[0]), (datasets))
plt.show()
# -
# +
#PM1 Analyzation
rmse = [0.270, 0.264, 0.274]
datasets = ['With Feature \nEngineering', 'No Feature Engineering + \nRedundant Variables',
'With Feature \nEngineering \nRedundant \nVariables']
plt.ylim(0, 0.35)
plt.xlabel("\nFeature Combinations")
plt.bar(datasets, rmse, align='center', alpha=0.7)
for a, b in zip(datasets, rmse):
plt.text(a, b, str(b) + "\n")
plt.ylabel("RMSE")
plt.show()
# +
#PM1 Analyzation
rmse = [0.284, 0.276, 0.270]
datasets = ['All Weather', 'Correlated \nWeather', 'With Feature \nEngineering']
plt.ylim(0, 0.35)
plt.xlabel("\nFeature Combinations")
plt.bar(datasets, rmse, align='center', alpha=0.7)
for a, b in zip(datasets, rmse):
plt.text(a, b, str(b) + "\n")
plt.ylabel("RMSE")
plt.show()
# +
line1 = [0.101, 0.123, 0.120, 0.105, 0.104] #Original Traffic
line2 = [0.191, 0.098, 0.087, 0.073, 0.116] #Transformedraffic
x = np.array(list(range(1, 6)))
plt.plot(x, line1)
plt.plot(x, line2)
plt.xticks(x)
plt.legend(["Original Traffic", "Transformed Traffic"])
plt.xlabel("Window Size")
plt.ylabel("RMSE")
plt.show()
# +
abc = 0.375
ab = 0.372
ac = 0.373
ab9 = 0.326
datasets = ["A+B+C", "A+B", "A+C", "A+B+W9"]
scores = [abc, ab, ac, ab9]
plt.bar(datasets, scores, align='center', alpha=0.5)
for a, b in zip(datasets, scores):
plt.text(a, b, str(b))
plt.xlabel("Feature Combinations")
plt.ylabel("RMSE")
plt.show()
# +
abc = 0.335
ab = 0.183
ac = 0.326
ab9 = 0.175
datasets = ["T+AW", "T'+AW", "T+BW", "T'+BW"]
scores = [abc, ab, ac, ab9]
plt.bar(datasets, scores, align='center', alpha=0.5)
for a, b in zip(datasets, scores):
plt.text(a, b, str(b))
plt.xlabel("Feature Combinations")
plt.ylabel("RMSE")
plt.show()
# +
output = pd.read_csv("logs/3-TAFT/fc_output_Taft Ave._2015.csv", skipinitialspace=True)
output.head()
startIndex = 5844
endIndex = 6899
#1 whole month
dt = output.dt[startIndex-2:endIndex-1]
actual = output.Actual[startIndex-2:endIndex-1]
predicted = output.Predicted[startIndex-2:endIndex-1]
plt.plot(predicted)
plt.plot(actual)
plt.legend(["Predicted Traffic", "Actual Traffic"])
plt.show()
# +
output = pd.read_csv("logs/7-TAFT (Orig Traffic, temp-windspd,temp4,press4)/fc_output_Taft Ave._2015.csv", skipinitialspace=True)
output.head()
startIndex = 5844
endIndex = 6899
#1 whole month
dt = output.dt[startIndex-2:endIndex-1]
actual = output.Actual[startIndex-2:endIndex-1]
predicted = output.Predicted[startIndex-2:endIndex-1]
plt.plot(predicted)
plt.plot(actual)
plt.legend(["Predicted Traffic", "Actual Traffic"])
plt.show()
# +
ROAD = "<NAME>"
YEAR = "2015"
EXT = ".csv"
fc_results = pd.read_csv("../output/fc_" + ROAD + "_" + YEAR + EXT, skipinitialspace=True)
line1 = fc_results.Actual[1117:1788]
line2 = fc_results.Predicted[1117:1788]
font = {'family' : 'normal',
'weight': 'normal',
'size' : 12}
matplotlib.rc('font', **font)
fig, ax = plt.subplots(figsize=(10,5))
linestyles = ['-', '--', '-.', ':']
ax.plot(line1, color='red', alpha=0.7, label='Actual Traffic')
fig.canvas.draw()
labels = [""]
for d in fc_results.dt[1110:1789]:
if "00:00:00" in d:
temp = d.split(" ")[0].split("-")
temp = "Oct" + temp[2]
labels.append(temp)
print(temp)
ax.set_xticklabels(labels)
ax.set_yticks([0, 0.5, 1.0])
ax.set_yticklabels(["Light \nTraffic", "Moderate \nTraffic", "Heavy \nTraffic"])
ax.plot(line2, linestyle=linestyles[1], color='blue', alpha=0.7, label='Predicted Traffic')
ax.legend(loc='upper right')
# -
| analysis/DataVisualization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# -*- coding: utf-8 -*-
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or
# implied.
# See the License for the specific language governing permissions and
# limitations under the License.
#
# -
# # Hyperparameter tuning with Random Forest
#
# Based on https://towardsdatascience.com/optimizing-hyperparameters-in-random-forest-classification-ec7741f9d3f6
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn
plt.rcParams['figure.figsize'] = 9, 6
from sklearn.model_selection import train_test_split
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import accuracy_score
from sklearn.model_selection import validation_curve
# -
# ## Plotting function
def plot_valid_curve(param_range, train_scores_mean, train_scores_std, test_scores_mean, test_scores_std):
plt.title("Validation Curve")
plt.xlabel(r"$\gamma$")
plt.ylabel("Score")
plt.ylim(0.991, 1.001)
lw = 2
plt.semilogx(param_range,
train_scores_mean,
label="Training score",
color="darkorange",
lw=lw
)
plt.fill_between(param_range,
train_scores_mean - train_scores_std,
train_scores_mean + train_scores_std,
alpha=0.2,
color="darkorange",
lw=lw
)
plt.semilogx(param_range,
test_scores_mean,
label="Cross-validation score",
color="navy",
lw=lw
)
plt.fill_between(param_range,
test_scores_mean - test_scores_std,
test_scores_mean + test_scores_std,
alpha=0.2,
color="navy",
lw=lw
)
plt.legend(loc="best")
return
# ## Wine dataset with split ratio 65:35
# URL https://archive.ics.uci.edu/ml/datasets/wine+quality
# +
red = pd.read_csv('data/winequality-red.csv', delimiter=';')
red['color'] = 1
white = pd.read_csv('data/winequality-white.csv', delimiter=';')
white['color'] = 0
data = pd.concat([red, white], ignore_index=True, sort=False)
n_samples, n_features = data.shape
n_samples, n_features
# -
# #### add noisy columns (simulate real data)
# +
# you can try a change to 100 to see computational effect
n_cols = 20 * n_features
random_state = np.random.RandomState(0)
df_cols = pd.DataFrame(data=random_state.randn(n_samples, n_cols),
columns=range(1, n_cols+1)
)
print(df_cols.shape)
data = pd.concat([data, df_cols], axis=1)
data.shape
# -
# #### split dataset
# +
train, test = train_test_split(data, test_size=0.35, shuffle=True, stratify=None)
print(len(train), len(test))
x_train, y_train = train.loc[:, train.columns != 'color'], train['color']
x_test, y_test = test.loc[:, test.columns != 'color'], test['color']
# -
# ## Random Forest classifier
# +
# default n_estimators=100
forest = RandomForestClassifier(random_state=1)
model = forest.fit(x_train, y_train)
y_pred = model.predict(x_test)
print(accuracy_score(y_test, y_pred))
# -
# ## Hyperparameter *n_estimators* tuning
# **Note: long running time - a few minutes**
# +
param_range = [100, 200, 300, 400, 500]
train_scores, test_scores = validation_curve(RandomForestClassifier(),
X = x_train,
y = y_train,
param_name = 'n_estimators',
param_range = param_range,
scoring="accuracy",
cv = 3
)
train_scores_mean = np.mean(train_scores, axis=1)
train_scores_std = np.std(train_scores, axis=1)
test_scores_mean = np.mean(test_scores, axis=1)
test_scores_std = np.std(test_scores, axis=1)
print(train_scores)
print(test_scores)
# -
# ## Plotting
plot_valid_curve(param_range, train_scores_mean, train_scores_std, test_scores_mean, test_scores_std)
# ## Random Forest with hyperparameter setting
# +
forest = RandomForestClassifier(random_state = 1,
n_estimators = 200,
max_depth = None,
min_samples_split = 2,
min_samples_leaf = 1
)
model = forest.fit(x_train, y_train)
y_pred = model.predict(x_test)
print(accuracy_score(y_test, y_pred))
# -
# ## Grid Search
# **Note: long running time cca 30m**
# +
n_estimators = [100, 200, 300, 400, 500]
max_depth = [5, 10, 20, 30, 40, 50]
min_samples_split = [2, 5, 10, 15, 20]
min_samples_leaf = [1, 2, 5, 10]
hyper = dict(n_estimators = n_estimators,
max_depth = max_depth,
min_samples_split = min_samples_split,
min_samples_leaf = min_samples_leaf
)
gs = GridSearchCV(forest, hyper, cv=3, verbose=1, n_jobs=-1)
best = gs.fit(x_train, y_train)
print(best)
# -
# ## The best model
# +
forest = RandomForestClassifier(n_estimators=200, random_state=1)
model = forest.fit(x_train, y_train)
y_pred = model.predict(x_test)
print(accuracy_score(y_test, y_pred))
# -
# # DÚ (len) na rozmýšľanie:
# https://scikit-learn.org/stable/modules/generated/sklearn.ensemble.RandomForestClassifier.html
# - different split ratio 65:35, 70:30, 75:26, 80:20
# - different hyperpamater setting
# - different criterion{“gini”, “entropy”}
# - different classifiers
# - different metrics
# - different datasets
#
# **The quest**
# - What is the best model?
# - Is the score good enough?
# - Do we need more tuning for this concrete case?
#
# **Auxiliary**
# - Nice visualizations of data and/or results
| cvicenia/tyzden-10/IAU_103_random-forest_hyperparameter-tuning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import turicreate as tc
# # Load train and test data
train_data = tc.SFrame.read_csv("Iris_train.csv")
train_data.show()
test_data = tc.SFrame.read_csv("Iris_test.csv")
test_data
# # Create a model
model = tc.decision_tree_classifier.create(train_data, "Species")
# # Save predictions to an SArray
predictions = model.predict(test_data)
predictions
# # Evaluate the model and save the results into a dictionary
results = model.evaluate(test_data)
results
# # Save the resulting SFrame to a file
model.save("Iris.model")
# # Export an activity classifier model in Core ML format
model.export_coreml("Iris.mlmodel")
| ml/Iris.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.ticker import (MultipleLocator, FormatStrFormatter,
AutoMinorLocator)
lin_range = np.linspace(start=0.01,stop=100,num=10000)
log_normalized = -(np.log10(lin_range)-2)
fig, ax = plt.subplots(figsize=(20, 20))
plt.scatter(lin_range, log_normalized, c=log_normalized, cmap = 'brg_r')
#plt.plot(lin_range, log_normalized )
plt.colorbar()
plt.title('Scatter plot: Logarithmic vs. Normalized linear AQ')
plt.xlabel('normalized linear AQ [%]')
plt.ylabel('logarithmic AQ')
plt.show()
# -
| AsksinPP_developments/sketches/HB-UNI-Sensor1-AQ-BME680_KF/Kalman_Filter/log_mapping.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Intro
# ¡Hola!
#
# Este curso sale derivado de la necesidad de poder enseñar a programar de manera asíncrona en los cursos de [Biología Matemática II](http://www.fciencias.unam.mx/docencia/horarios/presentacion/322261) y el [Seminario Matemáticas Aplicadas II](http://www.fciencias.unam.mx/docencia/horarios/presentacion/322339) en donde el uso de la computadora se hará fundamental.
# Si bien esta es una introducción somera a Python trataremos de cubrir las bases para que de manera autónoma se pueda extender el estudio del lenguaje hacia cualquiera de [_sus propósitos_](https://www.python.org/about/apps/). En Python es posible hacer [desarrollo web-fullstack](https://www.fullstackpython.com/web-development.html) y al mismo tiempo nos permite hacer cómputo científico. Incluso, junto con [R](https://www.r-project.org/about.html), es uno los lenguajes principales para desarrollo de machine learning, ingeniería de datos, visualización, entre otros tópicos bien interesantes.
# **NOTA:** Este curso está bien inspirado y guiado en el [curso de kaggle de Python](https://www.kaggle.com/learn/python) el cual es gratuito. Seguiré la misma estructura con ejemplos y ejercicios similares pero en esta serie iré introduciendo ejercicios que competen a un curso en biología matemática o matemáticas aplicadas.
# # ¡Hola, Python! (¿En Jupyter Notebook?)
# Antes de comenzar de lleno a hablar de Python me gustaría que abordaramos un poco dónde es que estamos. Y no me refiero _a la [Wittgenstein](https://en.wikipedia.org/wiki/Ludwig_Wittgenstein)_ sino más referente a donde vamos a programar. Una de dos, o estás corriendo este notebook en local en Jupyter [Notebook](https://jupyterlab.readthedocs.io/en/stable/getting_started/installation.html)/[Lab](https://jupyterlab.readthedocs.io/en/stable/getting_started/installation.html) o bien lo estás corriendo en [Google Colab](https://colab.research.google.com/) o un similar([Cocalc](https://cocalc.com/), [Kaggle Notebooks](https://www.kaggle.com/notebooks), [Binder](https://mybinder.org/), etc.). Si estás leyendo esto en GitHub te recomiendo que descarges este repositorio completo, lo subas a drive y te vayas directo a Colab. Aquí te explico cómo hacerlo.
# Ahora, ya que estamos aquí de verdad vamos a hablar de manera breve de los [Jupyter notebooks](https://jupyter.org/):
# >Un notebook de manera general es un formato con extentension `.ipynb` con un formato [JSON](https://en.wikipedia.org/wiki/JSON) que guarda un registro completo de la sesión de usuario (los cambios que hace o no hace en el archivo) e incluye código, texto narrativo, ecuaciones y una salida erriquecedora. El notebook se comunica con _kernels_ mediante el _protocolo de cómputo interactivo_. Por último, los kernels son procesos que corren código de manera interactiva en un lenguaje de programación específico y regresa el output a la persona que ejecuta el código.
#
# Lo que esto quiere decir es que vamos a poder ejecutar código en el lenguaje de nuestra preferencia y además en un ambiente que sea agradable e intereactivo no en una terminal plana. Además vamos a poder añadir texto con formato, latex y usar la salida del código, mostrar gráficas e incluso interactuar con ellas.
# Ahora sí, vamos con Python.
#
# Vamos a comenzar haciendo un programa corto pero no sencillo.
# +
# Conteo inicial de especies
numero_de_especies = 1
print(numero_de_especies)
# Nuevo conteo de especies
numero_de_especies = numero_de_especies + 4
if numero_de_especies > 3:
print("Hay más de tres en el conteo de la espcie")
canción_de_especímenes = '¡'+ "Espécimen " * numero_de_especies + '!'
print(canción_de_especímenes)
# -
#
# Con el pequeño bloque de código anterior vamos a poder demostrar los aspectos más importantes de cómo luce códifo de Python y además cómo trabaja. Vayamos a analizar el código desde las primeras líneas hasta la última.
numero_de_especies = 1
# **Asignamiento de variables:** Lo que hicimos en esta línea es crear una variable llamada `numero_de_especies` y le asigna un valor de 0 usando `=` el cual es llamado el operador de asignación.
# > **NOTA:** Si has programado alguna vez en un lenguaje como C/C#, Java o C++ notarás que en Python hay algunas cosas que no tienes que hacer:
# > * No necesitas _declarar_ las variables antes de usarla
# > * No necesitas decirle a Python qué tipo de valor le vas a asignar a una variable. Incluso puedes asignarle otro tipo de valor a la variable sin tener que declararla, por ejemplo puede ser un booleano o un flotante.
print(numero_de_especies)
# **Llamada de funciones:** `print` es una función de Python que muestra el valor que se le pasa en la pantalla. Llamamos a las funciones colocando paréntesis después del nombre y poniendo las entradas o argumentos de la función en esos paréntesis.
# Nuevo conteo de especies
numero_de_especies = numero_de_especies + 4
# La primer línea del block anterior es llamado un **comentario**. En Python los comentarios comienzan con el símbolo `#` y los comentarios son líneas de código que no se ejecutan, podemos escribir lo que queramos y no afectara a nuestro código.
# En la siguiente línea vemos una operación de _reasignamiento_. Únicamente se realiza una reasginación cuando la variable ya existía antes y cambiamos ahora el valor usando el operador de asignación `=`, si la variable no existía antes entonces es simplemente una asignación.
# En este caso estamos asignando a `numero_de_especies` el valor resultante de la operación aritmética `numero_de_especies + 4`... Raro ¿Cierto?
# En Python, y en la gran mayoría de leguajes, el programa se ejecuta línea por línea, i.e. primero la línea de código 0, luego la 1, hasta la $n$. Luego, en cada una de las líneas se ejecuta la operación que esté o bien a la derecha del operador de asignación, o bien dentro de parentesís.
#
# Por lo anterior es que entonces `numero_de_especies` tiene un valor de `5` pues primero se ejecuta la operación `numero_de_especies + 4` y el valor de `numero_de_especies` para ese momento era `1` por lo que `numero_de_especies + 4` es `5`, luego de que se resuelve esa operación se ejecuta la reasignación. `numero_de_especies` toma ahora el valor de `5`.
#
# ¿Cuál crees que sea ahora el output de la siguiente línea de código?
# ```python
# print(numero_de_especies + 1)
# ```
#
# +
if numero_de_especies > 3:
print("Hay más de tres en el conteo de la especie")
canción_de_especímenes = '¡'+ "Espécimen " * numero_de_especies + '!'
print(canción_de_especímenes)
# -
# Justo ahora no vamos a hablar de **condicionales** sin embargo aquí tenemos uno: `if`. Una cosa fantástica de Python es su legibilidad y simplicidad, podemos intuir el uso del `if`; solamente pasará algo si se cumple una condición, eso hará el `if`.
# En el caso anterior `"Hay más de tres en el conteo de la especie"` solamente aparecerá en la pantalla si `numero_de_especies` es mayor que `3`. Sin embargo el último `print(canción_de_especímenes)` se ejecutará sin importar el valor de `numero_de_especies` pero ¿Cómo sabemos nosotros (y Python) que se va ejecutar un bloque de código o no otro según una condición?
# Los dos puntos `:` al final del `if` indican que un _nuevo bloque de código_ es empezando. Las líneas de código subsecuentes que estén identadas son parte del nuevo bloque de código. Otros lenguajes usan `{}` para denotar el inicio de un nuevo código de bloque:
# ```javascript
# conteo_especies = 5
#
# if(conteo_especies == 5){
# console.log('Son 5 especies contadas')
# }
# ```
# El uso del espaciado o identación tiende a ser una cosa de gran sorpresa -en algunos casos de disgusto- para personas que han programado en otros lenguaje antes sin embargo con la práctica tiende a ser más consistente y limpio (no tienes que estar cazando un `}` perdido, por ejemplo).
# ¿Qué output crees que tenga el siguiente bloque de código?
# ```python
# conteo_especies = 5
# if conteo_especies > 2:
# if conteo_especies < 6:
# print("Hay más de 2 especies pero menos de 6")
# else:
# print("Hay contadas más de 6 especies")
# ```
#
# Las dos últimas líneas en nuestro código original no están identadas por lo que no pertenecen al bloque del `if`. Estas últimas líneas trabajan con la variable `canción_de_especímenes`. A lo largo del código trabjamos con **strings** en Python, por ejemplo
# ```python
# "Hay más de tres en el conteo de la especie"
# ```
# Las cadenas o strings pueden ser marcadas o denotadas por pares de comillas sencillas `"."` o por comidas simples `'.'` pero no combinaciones de ellas.
canción_de_especímenes = '¡'+ "Espécimen " * numero_de_especies + '!'
print(canción_de_especímenes)
# El operador `*` puede ser usado para multiplicar dos números (`3*3` no devolvería un `9`) pero también para multiplicar strings por números, para obtener esa string repetida el número del veces por el que se le multiplicó.
# Por otro lado tenemos el operador `+` que puede ser usado para sumar dos números (`3 + 3` nos devolvería un `6`) pero también para **concatenar** strings. Este operador es muy últil pues podemos realizar operaciones que en otros lenguajes es un poco engorrosa simplemente como `'¡' + 'Excelente' + '!'` lo cual nos devolvería la string `'¡Excelente!'`
#
# ¿Qué crees que nos deolvería el siguiente bloque de código?
# ```python
# a = 'azul'
# b = 'verde'
# amarillo = a + '+' + b
# print(amarillo)
# ```
# Por último, al hecho de que un operador pueda realizar una operación en función de qué tipo de variables esté operando se le llama **sobrecarga de operadores** y es bastante usado. Con la práctica se volverá muy natural usar los opereadores sin pensar en el tipo de variables.
# # Números y arimética en Python
# Hasta este momento ya revisamos un ejemplo de una variable que contiene un número:
numero_de_especies = 1
# Si nos referimos a _número_ realmente nos refiremos a un ente más complicado que dentro de Python, la forma de poder definirlo y pedirle a Python que la describe es como sigue:
type(numero_de_especies)
# Notemos que es un `int`, una abreviación para _integer_ (entero). Hay otro tipo de número que usaremos de manera común en Python:
type(3.1416)
# Un `float` es un número con dígitos decimales y son muy útiles para denotar pesos o proporciones.
# La función `type()` es la segunda función _built-in_ que vemos (la primera fue `print()`) y de verdad que es bien importante ser capaz de preguntarle a Python _¿Qué tipo de varible es esta?_
#
# La cosa más natural de querer hacer con los números son operaciones aritméticas. Hasta el momento ya usamos y visitamos `*` y `+`. En Python también tenemos el resto de operaciones básicas de operaciones de calculadora:
#
# | Operador |Nombre | Descripción |
# |:--------:|----------------------------------|--------------------------------------------------------------------------------------|
# | a+b | Suma | Suma de a y b |
# | a - b | Resta | Diferencia entre a y b |
# | a * b | Multiplicación | Producto de a y b |
# | a / b | División (_True Division_) | División entre a y b |
# | a // b | División entera (Floor division) | División entera entre a y b removiendo la parte decimal |
# | a % b | Módulo | Entero restante después de la división de a y b. Operación del módulo entre enteros. |
# | a ** b | Exponenciación | a elevado a la potencia b |
# | -a | Negación | El inverso o negativo de a |
#
# Quizás una de las cosas más interesantes es que mientras que otros lenguajes tiene únicamente una operación de división en Python hay dos: _True Division_ y _Floor Division_.
#
# Mientras que el operador `/` siempre nos da un `float`:
print('2/5 = {}'.format(2/5))
print('3/2 = {}'.format(3/2))
# El operador `//` siempre nos da un número entero: el entero más cercano al resultado de la división:
print('2//5 = {}'.format(2//5))
print('3//2 = {}'.format(3//2))
#
#
# ## Precedencia de operadores
# A lo largo de primaria, secundaria y preparatoria fuimos prendiendo nuevos tipos de operaciones y con ellas fuimos aprendiendo en qué orden ejecutarlas dentro de una expresión. El orden de ejecución de operaciones es Patentesís, Exponentes, Multiplicaciones/Divisiones y Adición/Substracción.
#
# En python las operaciones siguen un orden similar:
-3 + 7 + 8
-4 + 10*2
# Sin embargo el orden de las operaciones no es necesariamente lo que siempre queremos
largo_lengua_rana = 3 #cm
largo_rana = 5 #cm
# ¿Cuánto mide de largo una rana en metros con la lengua estirada?
largo_rana_total = largo_lengua_rana + largo_rana / 100
print("El largo total de la rana en metros es =",largo_rana_total,"(?)")
# Para corregir lo anterior neceistamos la ayuda de los parentesís. En cuanto nostros los añadimos creamos _sub-expresiones_ y con esto forzamos a Python a evaluarlas en el orden que queramos:
largo_rana_total = (largo_lengua_rana + largo_rana) / 100
print("El largo total de la rana en metros es =",largo_rana_total)
#
# ## Una par más de funciones built-in.
# Hasta ahora se presentaron ya dos funciones built-in `print()` y `type()`. Estas funciones pueden recibir cualquier tipo de variable y nos darán un output. Ahora vamos a presentar unas funciones que son específicas para poder trabajar con variables numéricas.
# Las funciones `min` y `max` nos dan como output el mínimo y el máximo de sus argumentos,
print(min(3,1,4))
print(max(1,1,2,3,5))
# Por otro lado la función `abs` nos regresa el valor absoluto de su argumento:
print(abs(42))
print(abs(-42))
# Por último, en adición a las funciones anteriores tenemos también las funciones `int` y `float` que además de ser tipos númericos en Python también pueden ser llamadas como funciones. Dichas funciones conviertes sus argumentos al tipo del nombre correspondiente:
print(float(42))
print(int(3.1416))
# Una cosa poderosa de éstas funciones es que pueden convertir strings en tipos numéricos:
print(int('41')+1)
print(float('3')+0.1416)
#
# ## Tu turno
# Ahora es tu oportunidad de probar lo que aprendimos en esta breve clase. Inténtalo en tu [primer notebook de ejercicios](./Ejercicios_1.ipynb).
| Clase_1/Clase_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
fuel = pd.read_csv("./Data/fuel.csv",
usecols=['Year','Make','Model','Range (FT1)','City Range (FT1)'])
fuel.head(10)
fuel.drop_duplicates(subset=['Year','Make','Model'],inplace=True)
fuel.dropna(how='any',inplace=True)
fuel.set_index(['Year','Make','Model'],inplace=True)
fuel.head()
fuel.stack().head()
fuel.stack().to_frame().head()
fuel.unstack().head()
fuel.unstack(level=0).head(5)
fuel.stack().unstack(level=0).head()
fuel.stack().unstack(level=1).head()
fuel.stack().unstack(level=['Year','Make']).head()
fuel.stack().unstack(level=['Make','Year']).sort_index(axis=1).head(20
)
fuel.stack().unstack(level=['Make','Year']).sort_index(axis=0).head(20)
| Multiindex_tabele_przestawne/Stack_Unstack_lab.ipynb |