code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Saving, Loading Qiskit Machine Learning Models and Continuous Training
#
# In this tutorial we will show how to save and load Qiskit machine learning models. Ability to save a model is very important, especially when a significant amount of time is invested in training a model on a real hardware. Also, we will show how to resume training of the previously saved model.
#
# In this tutorial we will cover how to:
#
# * Generate a simple dataset, split it into training/test datasets and plot them
# * Train and save a model
# * Load a saved model and resume training
# * Evaluate performance of models
# * PyTorch hybrid models
# First off, we start from the required imports. We'll heavily use SciKit-Learn on the data preparation step. In the next cell we also fix a random seed for reproducibility purposes.
# +
import matplotlib.pyplot as plt
import numpy as np
from qiskit import Aer
from qiskit.algorithms.optimizers import COBYLA
from qiskit.circuit.library import RealAmplitudes
from qiskit.utils import QuantumInstance, algorithm_globals
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import OneHotEncoder, MinMaxScaler
from qiskit_machine_learning.algorithms.classifiers import VQC
from IPython.display import clear_output
algorithm_globals.random_seed = 42
# -
# We will be using two quantum simulators. We'll start training on the QASM simulator then will resume training on the statevector simulator. The approach shown in this tutorial can be used to train a model on a real hardware available on the cloud and then re-use the model for inference on a local simulator.
# +
qi_qasm = QuantumInstance(
Aer.get_backend("aer_simulator"),
shots=1024,
seed_simulator=algorithm_globals.random_seed,
seed_transpiler=algorithm_globals.random_seed,
)
qi_sv = QuantumInstance(
Aer.get_backend("aer_simulator_statevector"),
seed_simulator=algorithm_globals.random_seed,
seed_transpiler=algorithm_globals.random_seed,
)
# -
# ## 1. Prepare a dataset
#
# Next step is to prepare a dataset. Here, we generate some data in the same way as in other tutorials. The difference is that we apply some transformations to the generated data. We generates `40` samples, each sample has `2` features, so our features is an array of shape `(40, 2)`. Labels are obtained by summing up features by columns and if the sum is more than `1` then this sample is labeled as `1` and `0` otherwise.
num_samples = 40
num_features = 2
features = 2 * algorithm_globals.random.random([num_samples, num_features]) - 1
labels = 1 * (np.sum(features, axis=1) >= 0) # in { 0, 1}
# Then, we scale down our features into a range of `[0, 1]` by applying `MinMaxScaler` from SciKit-Learn. Model training convergence is better when this transformation is applied.
features = MinMaxScaler().fit_transform(features)
features.shape
# Let's take a look at the features of the first `5` samples of our dataset after the transformation.
features[0:5, :]
# We choose `VQC` or Variational Quantum Classifier as a model we will train. This model, by default, takes one-hot encoded labels, so we have to transform the labels that are in the set of `{0, 1}` into one-hot representation. We employ SciKit-Learn for this transformation as well. Please note that the input array must be reshaped to `(num_samples, 1)` first. The `OneHotEncoder` encoder does not work with 1D arrays and our labels is a 1D array. In this case a user must decide either an array has only one feature(our case!) or has one sample. Also, by default the encoder returns sparse arrays, but for dataset plotting it is easier to have dense arrays, so we set `sparse` to `False`.
labels = OneHotEncoder(sparse=False).fit_transform(labels.reshape(-1, 1))
labels.shape
# Let's take a look at the labels of the first `5` labels of the dataset. The labels should be one-hot encoded.
labels[0:5, :]
# Now we split our dataset into two parts: a training dataset and a test one. As a rule of thumb, 80% of a full dataset should go into a training part and 20% into a test one. Our training dataset has `30` samples. The test dataset should be used only once, when a model is trained to verify how well the model behaves on unseen data. We employ `train_test_split` from SciKit-Learn.
train_features, test_features, train_labels, test_labels = train_test_split(
features, labels, train_size=30, random_state=algorithm_globals.random_seed
)
train_features.shape
# Now it is time to see how our dataset looks like. Let's plot it.
# +
def plot_dataset():
plt.scatter(
train_features[np.where(train_labels[:, 0] == 0), 0],
train_features[np.where(train_labels[:, 0] == 0), 1],
marker="o",
color="b",
label="Label 0 train",
)
plt.scatter(
train_features[np.where(train_labels[:, 0] == 1), 0],
train_features[np.where(train_labels[:, 0] == 1), 1],
marker="o",
color="g",
label="Label 1 train",
)
plt.scatter(
test_features[np.where(test_labels[:, 0] == 0), 0],
test_features[np.where(test_labels[:, 0] == 0), 1],
marker="o",
facecolors="w",
edgecolors="b",
label="Label 0 test",
)
plt.scatter(
test_features[np.where(test_labels[:, 0] == 1), 0],
test_features[np.where(test_labels[:, 0] == 1), 1],
marker="o",
facecolors="w",
edgecolors="g",
label="Label 1 test",
)
plt.legend(bbox_to_anchor=(1.05, 1), loc="upper left", borderaxespad=0.0)
plt.plot([1, 0], [0, 1], "--", color="black")
plot_dataset()
plt.show()
# -
# On the plot above we see:
#
# * Solid <span style="color:blue">blue</span> dots are the samples from the training dataset labeled as `0`
# * Empty <span style="color:blue">blue</span> dots are the samples from the test dataset labeled as `0`
# * Solid <span style="color:green">green</span> dots are the samples from the training dataset labeled as `1`
# * Empty <span style="color:green">green</span> dots are the samples from the test dataset labeled as `1`
#
# We'll train our model using solid dots and verify it using empty dots.
# ## 2. Train a model and save it
#
# We'll train our model in two steps. On the first step we train our model in `20` iterations.
maxiter = 20
# Create an empty array for callback to store values of the objective function.
objective_values = []
# We re-use a callback function from the Neural Network Classifier & Regressor tutorial to plot iteration versus objective function value with some minor tweaks to plot objective values at each step.
# +
# callback function that draws a live plot when the .fit() method is called
def callback_graph(_, objective_value):
clear_output(wait=True)
objective_values.append(objective_value)
plt.title("Objective function value against iteration")
plt.xlabel("Iteration")
plt.ylabel("Objective function value")
stage1_len = np.min((len(objective_values), maxiter))
stage1_x = np.linspace(1, stage1_len, stage1_len)
stage1_y = objective_values[:stage1_len]
stage2_len = np.max((0, len(objective_values) - maxiter))
stage2_x = np.linspace(maxiter, maxiter + stage2_len - 1, stage2_len)
stage2_y = objective_values[maxiter : maxiter + stage2_len]
plt.plot(stage1_x, stage1_y, color="orange")
plt.plot(stage2_x, stage2_y, color="purple")
plt.show()
plt.rcParams["figure.figsize"] = (12, 6)
# -
# As mentioned above we train a `VQC` model and set `COBYLA` as an optimizer with a chosen value of the `maxiter` parameter. Then we evaluate performance of the model to see how well it was trained. Then we save this model for a file. On the second step we load this model and will continue to work with it.
#
# Here, we manually construct an ansatz to fix an initial point where to start optimization from.
# +
original_optimizer = COBYLA(maxiter=maxiter)
ansatz = RealAmplitudes(num_features)
initial_point = np.asarray([0.5] * ansatz.num_parameters)
# -
# We create a model and set a quantum instance to the QASM simulator we created earlier.
original_classifier = VQC(
ansatz=ansatz, optimizer=original_optimizer, callback=callback_graph, quantum_instance=qi_qasm
)
# Now it is time to train the model.
original_classifier.fit(train_features, train_labels)
# Let's see how well our model performs after the first step of training.
print("Train score", original_classifier.score(train_features, train_labels))
print("Test score ", original_classifier.score(test_features, test_labels))
# Next, we save the model. You may choose any file name you want. Please note that the `save` method does not append an extension if it is not specified in the file name.
original_classifier.save("vqc_classifier.model")
# ## 3. Load a model and continue training
#
# To load a model a user have to call a class method `load` of the corresponding model class. In our case it is `VQC`. We pass the same file name we used in the previous section where we saved our model.
loaded_classifier = VQC.load("vqc_classifier.model")
# Next, we want to alter the model in a way it can be trained further and on another simulator. To do so, we set the `warm_start` property. When it is set to `True` and `fit()` is called again the model uses weights from previous fit to start a new fit. We also set quantum instance of the underlying network to the statevector simulator we created in the beginning of the tutorial. Finally, we create and set a new optimizer with `maxiter` is set to `80`, so the total number of iterations is `100`.
loaded_classifier.warm_start = True
loaded_classifier.neural_network.quantum_instance = qi_sv
loaded_classifier.optimizer = COBYLA(maxiter=80)
# Now we continue training our model from the state we finished in the previous section.
# + nbsphinx-thumbnail={"output-index": 0}
loaded_classifier.fit(train_features, train_labels)
# -
print("Train score", loaded_classifier.score(train_features, train_labels))
print("Test score", loaded_classifier.score(test_features, test_labels))
# Let's see which data points were misclassified. First, we call `predict` to infer predicted values from the training and test features.
train_predicts = loaded_classifier.predict(train_features)
test_predicts = loaded_classifier.predict(test_features)
# Plot the whole dataset and the highlight the points that were classified incorrectly.
# +
# return plot to default figsize
plt.rcParams["figure.figsize"] = (6, 4)
plot_dataset()
# plot misclassified data points
plt.scatter(
train_features[np.all(train_labels != train_predicts, axis=1), 0],
train_features[np.all(train_labels != train_predicts, axis=1), 1],
s=200,
facecolors="none",
edgecolors="r",
linewidths=2,
)
plt.scatter(
test_features[np.all(test_labels != test_predicts, axis=1), 0],
test_features[np.all(test_labels != test_predicts, axis=1), 1],
s=200,
facecolors="none",
edgecolors="r",
linewidths=2,
)
# -
# So, if you have a large dataset or a large model you can train it in multiple steps as shown in this tutorial.
# ## 4. PyTorch hybrid models
#
# To save and load hybrid models, when using the TorchConnector, follow the PyTorch recommendations of saving and loading the models. For more details please refer to the PyTorch Connector tutorial [here](https://qiskit.org/documentation/machine-learning/tutorials/05_torch_connector.html) where a short snippet shows how to do it.
#
# Take a look at this pseudo-like code to get the idea:
# ```python
# # create a QNN and a hybrid model
# qnn = create_qnn()
# model = Net(qnn)
# # ... train the model ...
#
# # save the model
# torch.save(model.state_dict(), "model.pt")
#
# # create a new model
# new_qnn = create_qnn()
# loaded_model = Net(new_qnn)
# loaded_model.load_state_dict(torch.load("model.pt"))
# ```
# +
import qiskit.tools.jupyter
# %qiskit_version_table
# %qiskit_copyright
| docs/tutorials/09_saving_and_loading models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: feml
# language: python
# name: feml
# ---
# ## Desafío:
#
# Mejora el desempeño de nuestra regressión Lasso **SOLO usando diferentes pasos de ingenería de variables!!**.
#
# El desempeño de nuestro modelo, como puedes ver en este notebook, es el siguiente:
# - rmse en el set de prueba: 44798.497576784845
# - r2 en el set de prueba: 0.7079639526659389
#
# Para mejorar el desempeño de nuestro modelo debes lograr un r2 en el set de prueba mayor que 0.71 y un rmse en el set de prueba menor que 44798.
#
#
# ### Condiciones:
#
# - No PUEDES cambiar los parámetros del modelo Lasso
# - DEBES usar las mismas semillas para la clase Lasso y el método train_test_split como mostramos en el notebook (no cambies los valores del parámetro random_state)
# - DEBES usar TODAS las variables del conjunto de datos (a excepción de Id)- NO PUEDES seleccionar variables
#
#
# ### Si mejoras el desempeño de nuestro modelo:
#
# Crea una solicitud de extracción -'pull request'- con tu notebook en este repositorio de github:
# https://github.com/solegalli/udemy-ivml-desafio
#
# Recuerda que necesitas primero bifurcar -'fork'- el repositorio, luego cargar tu notebook ganador a tu repositorio y finalmente crear una solicitudes de extracción (PR 'pull request') a nuestro repositorio. Luego revisaremos y fusionaremos tu solicitud, la cual aparecerá en nuestro repo y estará disponible para todos los estudiantes de este curso. De esta forma, otros estudiantes podrán aprender de tu creatividad e ingenio al transformar las variables de este conjunto de datos!
# ## House Prices dataset
# +
from math import sqrt
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# para el modelo
from sklearn.model_selection import train_test_split
from sklearn.linear_model import Lasso
from sklearn.pipeline import Pipeline
from sklearn.metrics import mean_squared_error, r2_score
# para la ingeniería de variables
from sklearn.preprocessing import StandardScaler
from feature_engine import missing_data_imputers as mdi
from feature_engine import discretisers as dsc
from feature_engine import categorical_encoders as ce
# -
# ### Cargar los datos
# +
# cargar los datos
data = pd.read_csv('../houseprice.csv')
# +
# crea una lista de los tipos de variables
categorical = [var for var in data.columns if data[var].dtype == 'O']
year_vars = [var for var in data.columns if 'Yr' in var or 'Year' in var]
discrete = [
var for var in data.columns if data[var].dtype != 'O'
and len(data[var].unique()) < 20 and var not in year_vars
]
numerical = [
var for var in data.columns if data[var].dtype != 'O'
if var not in discrete and var not in ['Id', 'SalePrice']
and var not in year_vars
]
print('Hay {} variables continuas '.format(len(numerical)))
print('Hay {} variables discretas'.format(len(discrete)))
print('Hay {} variables temporales'.format(len(year_vars)))
print('Hay {} variables categóricas'.format(len(categorical)))
# -
# ### Separar sets de entrenamiento y prueba
# +
# IMPORTANTE:
# manten el random_state igual a zero para poder reproducir los resultados
# Separemos los datos en sets de entrenamiento y prueba
X_train, X_test, y_train, y_test = train_test_split(data.drop(
['Id', 'SalePrice'], axis=1),
data['SalePrice'],
test_size=0.1,
random_state=0)
# +
# calculemos el tiempo transcurrido
def elapsed_years(df, var):
# captura la diferencia entre la variable año var
# y el año de construcción de la casa
df[var] = df['YrSold'] - df[var]
return df
for var in ['YearBuilt', 'YearRemodAdd', 'GarageYrBlt']:
X_train = elapsed_years(X_train, var)
X_test = elapsed_years(X_test, var)
# -
# elimina la variable YrSold
X_train.drop('YrSold', axis=1, inplace=True)
X_test.drop('YrSold', axis=1, inplace=True)
# captura el nombre de las columnas para usarlas más
# adelante en el notebook
final_columns = X_train.columns
# ## Pipeline de Ingeniería de Variables
# +
# Vamos a manipular las variables discretas como
# si fueran categóricas
# para lograrlo usando Feature-engine
# necesitamos modificar el tipo de variables a
# una tipo 'object'
X_train[discrete] = X_train[discrete].astype('O')
X_test[discrete] = X_test[discrete].astype('O')
# -
house_pipe = Pipeline([
# imputación de dato ausentes - sección 4
('missing_ind',
mdi.AddNaNBinaryImputer(
variables=['LotFrontage', 'MasVnrArea', 'GarageYrBlt'])),
('imputer_num',
mdi.MeanMedianImputer(
imputation_method='median',
variables=['LotFrontage', 'MasVnrArea', 'GarageYrBlt'])),
('imputer_cat', mdi.CategoricalVariableImputer(variables=categorical)),
# codificación de variables categóricas - sección 6
('rare_label_enc',
ce.RareLabelCategoricalEncoder(tol=0.05,
n_categories=6,
variables=categorical + discrete)),
('categorical_enc',
ce.OrdinalCategoricalEncoder(encoding_method='ordered',
variables=categorical + discrete)),
# discretiazación + codificación - sección 8
('discretisation',
dsc.EqualFrequencyDiscretiser(q=5,
return_object=True,
variables=numerical)),
('encoding',
ce.OrdinalCategoricalEncoder(encoding_method='ordered',
variables=numerical)),
# escalamiento de variables - sección 10
('scaler', StandardScaler()),
# regresión
('lasso', Lasso(random_state=0))
])
# +
# ajustemos el pipeline - conjunto de pasos de
# para el pre-procesamiento y modelamiento
house_pipe.fit(X_train, y_train)
# obtengamos las predicciones
X_train_preds = house_pipe.predict(X_train)
X_test_preds = house_pipe.predict(X_test)
# +
# revisemos desempeño del modelo:
print('mse entrenamiento: {}'.format(mean_squared_error(y_train, X_train_preds)))
print('rmse entrenamiento: {}'.format(sqrt(mean_squared_error(y_train, X_train_preds))))
print('r2 entrenamiento: {}'.format(r2_score(y_train, X_train_preds)))
print()
print('mse prueba: {}'.format(mean_squared_error(y_test, X_test_preds)))
print('rmse prueba: {}'.format(sqrt(mean_squared_error(y_test, X_test_preds))))
print('r2 prueba: {}'.format(r2_score(y_test, X_test_preds)))
# +
# grafiquemos las predicciones vs. los valores reales
plt.scatter(y_test,X_test_preds)
plt.xlabel('Precio real de las casas')
plt.ylabel('Predicciones de precio de las casas')
# +
# exploremos la importancia de las variables
# en este caso importancia es el valor
# absoluto del coeficiente asignado
# por Lasso
importance = pd.Series(np.abs(house_pipe.named_steps['lasso'].coef_))
importance.index = list(final_columns)+['LotFrontage_na', 'MasVnrArea_na', 'GarageYrBlt_na']
importance.sort_values(inplace=True, ascending=False)
importance.plot.bar(figsize=(18,6))
# -
| 13-Desafio.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Dataset
# +
from hana_ml import dataframe
import json
import time
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
from data_load_utils import DataSets, Settings
import plotting_utils
url, port, user, pwd = Settings.load_config("../../config/e2edata.ini")
conn = dataframe.ConnectionContext(url, port, user, pwd)
from scipy.stats import norm
from scipy.linalg import cholesky
import numpy as np
from numpy.random import rand
num_samples = 600
S1 = 12
S2 = 100
np.random.seed(seed=2334)
x1 = norm.rvs(loc=0, scale=1, size=(1, num_samples))[0]
x2 = norm.rvs(loc=0, scale=1, size=(1, num_samples))[0]
x3 = norm.rvs(loc=0, scale=1, size=(1, num_samples))[0]
x4 = norm.rvs(loc=0, scale=1, size=(1, num_samples))[0]
std_m = np.array([
[6.8, 0, 0, 0],
[0, 1.4, 0, 0],
[0, 0, 1.4, 0],
[0, 0, 0, 2.9]
])
# specify desired correlation
corr_m = np.array([
[1, .35, 0.33, 0.78],
[.35, 1, 0.90, 0.28],
[.33, 0.90, 1, 0.27],
[.78, 0.28, 0.27, 1]
])
# calc desired covariance (vc matrix)
cov_m = np.dot(std_m, np.dot(corr_m, std_m))
L = cholesky(cov_m, lower=True)
corr_data = np.dot(L, [x1, x2, x3, x4]).T
beta=np.array([-3.49, 13, 13, 0.0056])
omega1 = 2*np.pi/S1
omega2 = 2*np.pi/S2
timestamp = np.array([i for i in range(num_samples)])
y1 = np.multiply(50*rand(num_samples), 20*rand(1)*np.cos(omega1*timestamp)) \
+ np.multiply(32*rand(num_samples), 30*rand(1)*np.cos(3*omega1*timestamp)) \
+ np.multiply(rand(num_samples), rand(1)*np.sin(omega2*timestamp))
y2 = np.multiply(rand(num_samples), timestamp)
y3 = corr_data.dot(beta.T)
y = y1 + y2 + y3
plt.plot(y)
# -
# ### ARIMA Expaliner
timestamp = [i for i in range(len(y))]
raw = {'ID':timestamp, 'Y':y, 'X1':corr_data[:,0], 'X2':corr_data[:,1], 'X3':corr_data[:,2], 'X4':corr_data[:,3]}
rdata = pd.DataFrame(raw)
cutoff = (int)(rdata.shape[0]*0.9)
#print(len(rdata.iloc[:cutoff:,:]))
df_fit = dataframe.create_dataframe_from_pandas(conn, rdata.iloc[:cutoff,:], table_name='PAL_ARIMA_FIT_TBL', force=True)
print(df_fit.head(5).collect())
from hana_ml.algorithms.pal.tsa.arima import ARIMA
from hana_ml.algorithms.pal.tsa.auto_arima import AutoARIMA
tic = time.perf_counter()
arima= AutoARIMA(background_size=-1)
arima.fit(df_fit, key='ID', endog='Y', exog=['X1', 'X2', 'X3', 'X4'])
toc = time.perf_counter()
print('time is %.4f s' % (toc-tic))
print(arima.fitted_.collect())
print(arima.model_.collect())
#print(len(rdata.iloc[cutoff:,:]))
df_predict = dataframe.create_dataframe_from_pandas(conn, rdata.iloc[cutoff:,:], table_name='PAL_ARIMA_PREDICT_TBL', force=True)
#print(df_predict.head(5).collect())
tic = time.perf_counter()
res = arima.predict(df_predict, top_k_attributions=30, seasonal_width=0.035, trend_width=0.035, show_explainer=True)
toc = time.perf_counter()
print('time is %.4f s' % (toc-tic))
print(res.head(5).collect())
print(arima.explainer_.head(5).collect())
# +
def plotComp(explainer, s = 0, ranges=[]):
name = ["TREND","SEASONAL","TRANSITORY","IRREGULAR"]
plt.figure(figsize=(8,8))
for i in range(1, 5, 1):
plt.subplot(4,1,i)
plt.plot(explainer.iloc[:,i].to_numpy())
ax = plt.gca()
if i == 1 and s > 1:
for j in range(explainer.shape[0] // s):
ax.axvline(x=(j+1)*s, color='red', ls='--')
plt.title(name[i-1])
if i != 4:
ax.get_xaxis().set_visible(False)
plotComp(arima.explainer_.collect())
# +
def readRScode(explainer):
dicts = {}
for j in range(explainer.shape[0]):
if len(explainer["EXOGENOUS"].values[j]) == 0:
return dicts
fattr = json.loads(explainer["EXOGENOUS"].values[j])
for i in range(len(fattr)):
if(fattr[i]['attr'] in dicts):
dicts[fattr[i]['attr']].append(fattr[i]['val'])
else:
dicts[fattr[i]['attr']] = [fattr[i]['val']]
return dicts
def plotExt(explainer):
dicts = readRScode(explainer)
if len(dicts) == 0:
return
ax = pd.DataFrame(dicts).plot(figsize=(12,6), kind='bar', stacked=True)
ax.legend(bbox_to_anchor=(1.0, 1.0))
ax.plot()
plotExt(arima.explainer_.collect())
# -
decompose_result = arima.explainer_.collect()
#print(decompose_result)
plt.plot(decompose_result['TREND']+decompose_result["SEASONAL"]+decompose_result["IRREGULAR"])
plt.plot(res.collect()['FORECAST'])
plt.plot(df_predict.collect()['Y'])
# ### Additive Model Forecast Explainer
# +
from hana_ml.algorithms.pal.tsa import additive_model_forecast
dates = pd.date_range('2018-01-01', '2019-08-23',freq='D')
data_additive = {'ID':dates, 'Y':y, 'X1':corr_data[:,0], 'X2':corr_data[:,1], 'X3':corr_data[:,2], 'X4':corr_data[:,3]}
data = pd.DataFrame(data_additive)
cutoff = (int)(data.shape[0]*0.9)
print(len(data.iloc[:cutoff:,:]))
df_fit_additive = dataframe.create_dataframe_from_pandas(conn, data.iloc[:cutoff,:], table_name='PAL_ADDITIVE_FIT_TBL', force=True)
print(df_fit_additive.head(5).collect())
print(len(data.iloc[cutoff:,:]))
df_predict_additive= dataframe.create_dataframe_from_pandas(conn, data.iloc[cutoff:,:], table_name='PAL_ADDITIVE_PREDICT_TBL', force=True)
print(df_predict_additive.head(5).collect())
# -
holiday_dic={"Date":['2018-01-01','2018-01-04','2018-01-05','2019-06-25','2019-06-29'],
"Name":['A', 'A', 'B', 'A', 'D']}
df=pd.DataFrame(holiday_dic)
df_holiday= dataframe.create_dataframe_from_pandas(conn, df, table_name='PAL_HOLIDAY_TBL', force=True)
df_holiday=df_holiday.cast('Date', 'TIMESTAMP')
#print(df_holiday.dtypes())
# +
amf = additive_model_forecast.AdditiveModelForecast(growth='linear',
regressor = ['{"NAME": "X1", "PRIOR_SCALE":4, "MODE": "additive" }'],
seasonality=['{ "NAME": "yearly", "PERIOD":365.25, "FOURIER_ORDER":10 }',
'{ "NAME": "weekly", "PERIOD":7, "FOURIER_ORDER":3 }',
'{ "NAME": "daily", "PERIOD":1, "FOURIER_ORDER":4 }'])
#amf = additive_model_forecast.AdditiveModelForecast(growth='linear',
# regressor = ['{"NAME": "X1", "PRIOR_SCALE":4, "MODE": "multiplicative"}',
# '{"NAME": "X2", "PRIOR_SCALE":4, "MODE": "multiplicative"}'],
# seasonality=['{ "NAME": "yearly", "PERIOD":365.25, "FOURIER_ORDER":10 }',
# '{ "NAME": "weekly", "PERIOD":7, "FOURIER_ORDER":3 }',
# '{ "NAME": "daily", "PERIOD":1, "FOURIER_ORDER":4 }'])
amf.fit(df_fit_additive, key='ID', endog='Y', exog=['X1','X2','X3','X4'], holiday=df_holiday)
model_content = amf.model_.collect()['MODEL_CONTENT']
# +
#MODEL_CONTENT
#"regressor_mode":[[1,0,0,0]], 1 is multiplicative, 0 is additive.
#{"GROWTH":"linear","FLOOR":0.0,"SEASONALITY_MODE":"additive","start":"2018-01-01 00:00:00.0000000","y_scale":1446.4012732304484,"t_scale":46569600.0,"cat_field_moments":"","regressor_name":[["X1","X2","X3","X4"]],"regressor_mu":[[-0.15352821733097303,-0.02059063426228545,-0.002622304887415057,-0.10345704770210677]],"regressor_std":[[6.791693022193103,1.3353406148220398,1.3372264170094813,2.9259028871764767]],"regressor_mode":[[1,0,0,0]],"regressor_prior_scale":[[4.0,10.0,10.0,10.0]],"seasonality_name":[["DAILY","weekly"]],"seasonality_period":[[30.0,7.0]],"seasonality_fourier_order":[[10.0,3.0]],"seasonality_prior_scale":[[10.0,10.0]],"seasonality_mode":[[0,0]],"holiday_name":[["A","A","B","A","D"]],"holiday_ts":[["2018-01-01 00:00:00.0000000","2018-01-04 00:00:00.0000000","2018-01-05 00:00:00.0000000","2019-06-25 00:00:00.0000000","2019-06-29 00:00:00.0000000"]],"holiday_lw":[[0,0,0,0,0]],"holiday_uw":[[0,0,0,0,0]],"k":0.15606952574757727,"m":0.025836199662412366,"delta":[[-6.292126065062676e-9,4.01672269727309e-9,2.1199765509020952e-10,1.1932762503249709e-8,-1.047803486376465e-9,1.8065798418999697e-9,-2.8338697798904576e-9,2.2734610420403154e-9,-9.36221620884908e-10,-3.096382176225027e-10,5.653371984077469e-10,0.000031771424734759676,-1.5103667461775418e-9,-6.952915812686268e-10,5.433302104204915e-10,1.4341194324681475e-8,-3.574888497386598e-9,-9.234203104205178e-9,1.4041591463701508e-9,-4.00089518484598e-10,8.627030356665826e-10,-9.464287117143478e-10,4.888208823553357e-9,3.401487456236214e-9,-4.233127346960964e-9]],"sigma_obs":0.29438644660697607,"beta":[[0.015268230138996909,-0.003723949172106288,-0.0012454511299372015,-0.014667147419791627,-0.002224602367867149,-0.011283684269001934,-0.007317937252829013,-0.0004909591674430246,0.015828377538860389,0.010889381438311124,0.013112690781791187,0.004675841012075908,-0.004266746836168113,-0.004887592550691263,-0.007487924296738099,-0.015804821532827714,-0.004811506922077433,0.0014824729740768239,0.001687420100693144,-0.022634540897335136,0.005235189349347453,0.010465205938117489,0.001449572788947729,-0.017951886315960669,0.009509697995534044,0.006325292067880513,0.01758779609226314,-0.003368025414656735,0.0,0.03176160107239431,-0.019135738672922624,0.041173876634595849,-0.01850148374778878]],"changepoints_t":[[0.03153988868274583,0.06307977736549166,0.09647495361781076,0.1280148423005566,0.15955473098330243,0.19109461966604825,0.22448979591836736,0.2560296846011132,0.287569573283859,0.31910946196660486,0.3525046382189239,0.38404452690166976,0.4155844155844156,0.44712430426716145,0.4805194805194805,0.5120593692022264,0.5435992578849722,0.575139146567718,0.608534322820037,0.640074211502783,0.6716141001855288,0.7031539888682746,0.7365491651205937,0.7680890538033395,0.7996289424860853]],"holidays_prior_scale":10.0}
# +
#t= dataframe.create_dataframe_from_pandas(conn, amf.model_.collect(), table_name='AAAA', force=True)
# -
res = amf.predict(data=df_predict_additive, key= 'ID', show_explainer=True, decompose_seasonality=True, decompose_holiday=True)
print(res.head(5).collect())
print(amf.explainer_.head(5).collect())
print(amf.explainer_.head(15).collect()['SEASONAL'])
print(amf.explainer_.head(5).collect()['EXOGENOUS'][0])
res = amf.predict(data=df_predict_additive, show_explainer=True, decompose_seasonality=False, decompose_holiday=False)
print(res.head(10).collect())
print(amf.explainer_.head(10).collect())
# +
explainer= amf.explainer_.collect()
def readExog(explainer):
dicts = {}
for j in range(explainer.shape[0]):
if len(explainer["EXOGENOUS"].values[j]) == 2:
return dicts
exog = json.loads(explainer["EXOGENOUS"].values[j])
if j == 0:
dicts['X1']= [exog['X1']]
dicts['X2']= [exog['X2']]
dicts['X3']= [exog['X3']]
dicts['X4']= [exog['X4']]
else:
dicts['X1'].append(exog['X1'])
dicts['X2'].append(exog['X2'])
dicts['X3'].append(exog['X3'])
dicts['X4'].append(exog['X4'])
return dicts
def plotExt(explainer):
dicts = readExog(explainer)
if len(dicts) == 0:
return
#print(dicts)
ax = pd.DataFrame(dicts).plot(figsize=(12,6), kind='bar', stacked=True)
ax.legend(bbox_to_anchor=(1.0, 1.0))
ax.plot()
plotExt(explainer)
# -
| Python-API/pal/notebooks/Time_Series_Explainer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Archive Streaming
# This notebook demonstrates:
#
# 1. how to visualize an archive (downloaded with [busdata.streambus](example_busdata.ipynb)) as a live stream; and,
# 2. how to infer bus operation information from the stream
#
# with the `simulator` module in the `buskit` package.
#
# ## NOTE: For inferring, reset the route (step 2.1) before each attempt to clear existing info in objects.
# (will be incorporated into functions or methods in the future)
# # 1. Setup
# +
import csv
import time
import dateutil
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import buskit as bk
from buskit import busdata as bdata
from buskit import dashboard as bdash
from buskit import simulator as bsim
from IPython.display import display, clear_output
# %pylab inline
# -
# # 2. Streaming Environment
# ## 2.1 Route Setup
# +
# specify path to the archive AVL file
archive_path = 'MTA_data/B15-180625-235941-44650-Mon.csv'
# parse stops and set links
stops, links, stop_pos = bsim.set_route(archive_path, 1)
# create empty dictionary storage for bus objects
live_bus = {}
# -
# ## 2.2 Archive Stream Visualization
# +
# bsim.plot_stream?
# -
bsim.plot_stream(archive_path, 1, live_bus, stops, links, stop_pos)
# ## 2.3 Archive Stream Inferring
# Trying to infer the following:
# 1. Link traveling speed (LTS)
# 2. Dwelling time (DT)
#
# The inferred data is stored in all stop and link objects
# +
# bsim.infer?
# -
bsim.infer(archive_path, 1, live_bus, stops, links, stop_pos, runtime=90)
# plot TSD for cross-validation
df = pd.read_csv(archive_path)
df = bdata.split_trips(df)
bdata.plot_tsd(df, 1, 0, 90)
# ### 2.3.1 Stop
# stop name
stops[37].name
# buses that passed or stoped at the stop
stops[37].log_bus_ref
# the assumed bus arrival times at the stop
stops[37].log_arr_t
# dwell time records (5 is the default if the bus is passing by without pinging at the stop)
stops[37].log_dwell_t
# headways OR max pax waiting time between each bus
stops[37].log_wait_t
# ### 2.3.2 Link
# the calculated average traveling speed
# for each pair of consecutive ping that covers this link
links[37].log_speed
| example_stream-archive.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
er_data = pd.read_csv('employee_retention_data.csv')
er_data['join_date'] = pd.to_datetime(er_data['join_date'])
er_data['quit_date'] = pd.to_datetime(er_data['quit_date'])
# # DATA EXPLORATION
# First, calculating the employee retention time based on thier joining and leaving dates.
er_data['retention_time'] = er_data['quit_date'].subtract(er_data['join_date']).astype('timedelta64[D]') / 365
er_data.corr()
# +
fig, axes = plt.subplots(nrows=1, ncols=2, sharey=True)
axes[0].scatter(er_data['salary'], er_data['retention_time'])
axes[0].set_xlabel("Salary ($)")
axes[0].set_ylabel("Retention Time (days)")
axes[1].scatter(er_data['seniority'], er_data['retention_time'])
axes[1].set_xlabel("Seniority (years)")
axes[1].set_xlim([0,30])
plt.tight_layout(pad=0.4, w_pad=0.5, h_pad=1.0)
plt.show()
# -
# It can be seen from the above correlation table and scatter plot that there is low correlation between the employee retention time with the employee salary and seniority.
plt.scatter(er_data['salary'], er_data['seniority'])
plt.xlabel("Salary ($)")
plt.ylabel("Seniority (years)")
plt.show()
# As one would expect, employees with higher seniority have higher salaries. Note we have outliers in the data: there are people who have seniority of 100 years! (do companies also hire zombies?)
er_data.sort_values(['join_date', 'quit_date']).describe()
fig, axes = plt.subplots(nrows=1, ncols=3)
er_data.boxplot('seniority', ax=axes[0])
er_data.boxplot('salary', ax=axes[1])
er_data.boxplot('retention_time', ax=axes[2])
plt.tight_layout(pad=0.4, w_pad=0.5, h_pad=1.0)
plt.show()
# Usually employees retain for around a little over a year (1.14 years). There are few outliers which retain for around 3-5 years.
sns.set(style="ticks")
sns.pairplot(er_data.fillna(0)[['employee_id','company_id', 'seniority', 'salary', 'retention_time']], hue='company_id')
plt.show()
# From the bottom left histogram for retention time, it can be seen that most people stay at a company for either 1, 1.5 or 2 years for all companies.
# ### Time Series Table
# +
# table for number of employees quit the job grouped by company id and quit date
outflow = er_data.groupby(['company_id', 'quit_date'], as_index=False)['employee_id'].count()
outflow['flow'] = -outflow['employee_id']
outflow['date'] = outflow['quit_date']
del outflow['employee_id']
del outflow['quit_date']
# table for number of employees joined the job grouped by company id and join date
inflow = er_data.groupby(['company_id', 'join_date'], as_index=False)['employee_id'].count()
inflow['flow'] = inflow['employee_id']
inflow['date'] = inflow['join_date']
del inflow['employee_id']
del inflow['join_date']
# employee flow table
flow_data = pd.concat([outflow, inflow]).sort_values(['company_id', 'date'])
flow_data['head_count'] = flow_data.groupby('company_id', as_index=False)['flow'].cumsum()
# -
company_1 = flow_data[flow_data.company_id == 1]
company_2 = flow_data[flow_data.company_id == 2]
company_3 = flow_data[flow_data.company_id == 3]
plt.plot(company_1['date'], company_1['flow'], company_2['date'], company_2['flow'], company_3['date'], company_3['flow'])
plt.xlabel('Time')
plt.ylabel('Flow')
plt.show()
plt.plot(company_1['date'], company_1['head_count'], company_2['date'], company_2['head_count'], company_3['date'], company_3['head_count'])
plt.xlabel('Time')
plt.ylabel('Head Count')
plt.show()
# As one would expect, different companies have different churns.
# +
from lifelines import KaplanMeierFitter
from lifelines.utils import datetimes_to_durations
kmf = KaplanMeierFitter()
T, E = datetimes_to_durations(er_data['join_date'], er_data['quit_date'])
kmf.fit(T, event_observed=E)
kmf.plot()
plt.show()
# -
# The KM_estimate indicates that people mostly leave in 365 days, and around the 700 days.
# # Conclusion
# THere are poor predictive features in the data. On can predict retention by the tenure period of an employee. Most employees leave in one year or the second year of thier employement.
| Practice/Data_Challenge/data_challenge_1/Solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import warnings
warnings.filterwarnings('ignore')
import numpy as np
import matplotlib.pylab as plt
import copy
import sys
sys.path.append('../')
from angler import Simulation, Optimization
from angler.structures import ortho_port
from angler.plot import *
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
# -
lambda0 = 2e-6 # free space wavelength (m)
c0 = 3e8 # speed of light in vacuum (m/s)
omega = 2*np.pi*c0/lambda0 # angular frequency (2pi/s)
dl = 0.4e-1 # grid size (L0)
NPML = [25, 25] # number of pml grid points on x and y borders
pol = 'Ez' # polarization (either 'Hz' or 'Ez')
source_amp = 6 # amplitude of modal source (A/L0^2?)
# material constants
n_index = 2.44 # refractive index
eps_m = n_index**2 # relative permittivity
chi3 = 4.1*1e-19 # Al2S3 from Boyd (m^2/V^2)
# max_ind_shift = 5.8e-3 # maximum allowed nonlinear refractive index shift (computed from damage threshold)
# geometric parameters
L1 = 6 # length waveguides in design region (L0)
L2 = 6 # width of box (L0)
H1 = 6 # height waveguides in design region (L0)
H2 = 6 # height of box (L0)
w = .3 # width of waveguides (L0)
l = 3 # length of waveguide from PML to box (L0)
spc = 3 # space between box and PML (L0)
# +
# define permittivity of three port system
eps_r, design_region = ortho_port(L1, L2, H1, H2, w, l, dl, NPML, eps_m)
(Nx, Ny) = eps_r.shape
nx, ny = int(Nx/2), int(Ny/2) # halfway grid points
simulation = Simulation(omega,eps_r,dl,NPML,pol)
# compute the grid size the total grid size
print("Computed a domain with {} grids in x and {} grids in y".format(Nx,Ny))
print("The simulation has {} grids per free space wavelength".format(int(lambda0/dl/simulation.L0)))
simulation.plt_eps()
plt.show()
print(np.sum(simulation.eps_r[1,:]>1))
print(np.sum(simulation.eps_r[:,1]>1))
print(np.sum(simulation.eps_r[:,-1]>1))
# +
# set the modal source and probes
simulation = Simulation(omega, eps_r, dl, NPML, 'Ez')
simulation.add_mode(np.sqrt(eps_m), 'x', [NPML[0]+int(l/2/dl), ny], int(H1/2/dl), scale=source_amp)
simulation.setup_modes()
# left modal profile
right = Simulation(omega, eps_r, dl, NPML, 'Ez')
right.add_mode(np.sqrt(eps_m), 'x', [-NPML[0]-int(l/2/dl), ny], int(H1/2/dl))
right.setup_modes()
J_right = np.abs(right.src)
# top modal profile
top = Simulation(omega, eps_r, dl, NPML, 'Ez')
top.add_mode(np.sqrt(eps_m), 'y', [nx, -NPML[1]-int(l/2/dl)], int(L1/2/dl))
top.setup_modes()
J_top = np.abs(top.src)
# -
# set source and solve for electromagnetic fields
(_, _, Ez) = simulation.solve_fields()
simulation.plt_abs(outline=True, cbar=True);
# +
# compute straight line simulation
eps_r_wg, _ = two_port(L1, H1, w, l, spc, dl, NPML, eps_start=eps_m)
(Nx_wg, Ny_wg) = eps_r_wg.shape
nx_wg, ny_wg = int(Nx_wg/2), int(Ny_wg/2) # halfway grid points
simulation_wg = Simulation(omega, eps_r_wg, dl, NPML, 'Ez')
simulation_wg.add_mode(np.sqrt(eps_m), 'x', [NPML[0]+int(l/2/dl), ny_wg], int(Ny/3), scale=source_amp)
simulation_wg.setup_modes()
# compute normalization
sim_out = Simulation(omega, eps_r_wg, dl, NPML, 'Ez')
sim_out.add_mode(np.sqrt(eps_m), 'x', [-NPML[0]-int(l/2/dl), ny], int(Ny/3))
sim_out.setup_modes()
J_out = np.abs(sim_out.src)
(_, _, Ez_wg) = simulation_wg.solve_fields()
SCALE = np.sum(np.square(np.abs(Ez_wg))*J_out)
J_out = J_out
print('computed a scale of {} in units of E^2 J_out'.format(SCALE))
J_right = J_right / SCALE
J_top = J_top / SCALE
# +
# set source and solve for electromagnetic fields
f, (ax1, ax2, ax3) = plt.subplots(1,3, figsize=(20,5))
_ = simulation.solve_fields()
_ = top.solve_fields()
_ = right.solve_fields()
simulation.plt_abs(outline=True, cbar=True, ax=ax1)
top.plt_abs(outline=True, cbar=True, ax=ax2)
right.plt_abs(outline=True, cbar=True, ax=ax3)
ax1.set_title('left sourced')
ax2.set_title('top sourced')
ax3.set_title('right sourced')
plt.show()
# -
# changes design region. 'style' can be in {'full', 'empty', 'halfway', 'random'}
simulation.init_design_region(design_region, eps_m, style='random_sym')
simulation.plt_eps()
plt.show()
# +
# add nonlinearity
nl_region = copy.deepcopy(design_region)
simulation.nonlinearity = [] # This is needed in case you re-run this cell, for example (or you can re-initialize simulation every time)
simulation.add_nl(chi3, nl_region, eps_scale=True, eps_max=eps_m)
# +
# define objective function
import autograd.numpy as npa
from angler.objective import Objective, obj_arg
arg1 = obj_arg('ez', component='Ez', nl=False)
arg2 = obj_arg('ez_nl', component='Ez', nl=True)
def J(e, e_nl):
linear_right = 1*npa.sum(npa.square(npa.abs(e))*J_right)
linear_top = -1*npa.sum(npa.square(npa.abs(e))*J_top)
nonlinear_right = -1*npa.sum(npa.square(npa.abs(e_nl))*J_right)
nonlinear_top = 1*npa.sum(npa.square(npa.abs(e_nl))*J_top)
objfn = (linear_right + linear_top + nonlinear_right + nonlinear_top)/2
return objfn
objective = Objective(J, arg_list=[arg1, arg2])
# -
# make optimization object and check derivatives
R = 5 # filter radius of curvature (pixels) (takes a while to set up as R > 5-10)
beta = 500
eta= 0.50
simulation.init_design_region(design_region, eps_m, style='halfway')
optimization = Optimization(objective=objective, simulation=simulation, design_region=design_region, eps_m=eps_m, R=R, beta=beta, eta=eta)
plt.imshow(simulation.rho.T)
plt.colorbar()
plt.show()
plt.imshow(simulation.eps_r.T)
plt.colorbar()
plt.show()
# check the derivatives (note, full derivatives are checked, linear and nonlinear no longer separate)
(grad_avm, grad_num) = optimization.check_deriv(Npts=5, d_rho=5e-4)
print('adjoint gradient = {}\nnumerical gradient = {}'.format(grad_avm, grad_num))
new_eps = optimization.run(method='lbfgs', Nsteps=1000, step_size=1e-1)
f, (ax1, ax2) = plt.subplots(1, 2, figsize=(16,5))
simulation.plt_eps(ax=ax1, outline=False)
ax1.set_title('final permittivity distribution')
optimization.plt_objs(ax=ax2)
ax2.set_yscale('linear')
plt.show()
# +
# compare the linear and nonlinear fields
# setup subplots
f, (ax1, ax2, ax3) = plt.subplots(1,3, figsize=(20,5))
# linear fields
(Hx,Hy,Ez) = simulation.solve_fields()
simulation.plt_abs(ax=ax1, vmax=20)
ax1.set_title('linear field')
# nonlinear fields
(Hx_nl,Hy_nl,Ez_nl,_) = simulation.solve_fields_nl()
simulation.plt_abs(ax=ax2, vmax=20, nl=True)
ax2.set_title('nonlinear field')
# difference
simulation.plt_diff(ax=ax3)
ax3.set_title('|Ez| for linear - nonlinear')
plt.show()
# -
# compute the refractive index shift
index_shift = simulation.compute_index_shift()
print('maximum refractive index shift of {}'.format(np.max(index_shift)))
plt.imshow(index_shift.T, cmap='magma', origin='lower')
plt.colorbar()
plt.title('refractive index shift')
plt.show()
# +
# input power
W_in = simulation.W_in
print("W_in = {}".format(W_in))
# linear powers
(Hx,Hy,Ez) = simulation.solve_fields()
W_right_lin = simulation.flux_probe('x', [-NPML[0]-int(l/2/dl), ny], int(H1/2/dl))
W_top_lin = simulation.flux_probe('y', [nx, -NPML[1]-int(l/2/dl)], int(H1/2/dl))
# nonlinear powers
(Hx_nl,Hy_nl,Ez_nl,_) = simulation.solve_fields_nl()
W_right_nl = simulation.flux_probe('x', [-NPML[0]-int(l/2/dl), ny], int(H1/2/dl), nl=True)
W_top_nl = simulation.flux_probe('y', [nx, -NPML[1]-int(l/2/dl)], int(H1/2/dl), nl=True)
print('linear transmission (right) = {:.4f}'.format(W_right_lin / W_in))
print('linear transmission (top) = {:.4f}'.format(W_top_lin / W_in))
print('nonlinear transmission (right) = {:.4f}'.format(W_right_nl / W_in))
print('nonlinear transmission (top) = {:.4f}'.format(W_top_nl / W_in))
S = [[W_top_lin / W_in, W_right_lin / W_in],
[W_top_nl / W_in, W_right_nl / W_in]]
plt.imshow(S, cmap='magma')
plt.colorbar()
plt.title('power matrix')
plt.show()
| Notebooks/T_port.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Mining SMS data from the UK, classifying spam messages
#
# # Table of Contents
# 1. [Imports](#imports)
# 2. [Exploratory Data Analysis](#eda)
# 1. [Text Preprocessing](#preprocessing)
# 2. [Feature Engineering](#features)
# 3. [Sentiment Analysis](#sentiment)
# 3. [Classification](#classification)
# # 1. Imports <a class='anchor' id='imports'></a>
import numpy as np
from scipy import sparse
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from collections import Counter
from nltk.corpus import stopwords
stop_words = set(stopwords.words('english'))
from nltk import word_tokenize
from nltk.stem.porter import PorterStemmer
import re
from vaderSentiment.vaderSentiment import SentimentIntensityAnalyzer
from wordcloud import WordCloud
from sklearn.model_selection import train_test_split, cross_val_score
from sklearn.metrics import accuracy_score, confusion_matrix
from sklearn.ensemble import RandomForestClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.preprocessing import LabelEncoder
from sklearn.decomposition import TruncatedSVD
# # 2. Exploratory Data Analysis <a class='anchor' id='eda'></a>
# We needed to specify names for the columns in order to be able to read the dataset correctly otherwise parsing text containing delimiter symbol would raise error
#
# Also there were two null objects in texts so we dropped them
# +
df = pd.read_csv(
'../data/SmsCollection.csv',
sep=';',
names=['label', 'text'],
skiprows=1,
encoding='utf8').dropna()
df.head()
# -
df.info()
df.groupby('label').describe()
sns.set(
style='whitegrid',
palette='Set2',
font_scale=1.4,
rc={"figure.figsize": [10, 6]})
sns.palplot(sns.color_palette('Set2'))
sns.countplot(df['label'])
plt.title('Distribution of spam vs ham')
plt.xlabel('')
plt.ylabel('Count')
plt.show()
# ## A. Text Preprocessing <a class='anchor' id='preprocessing'></a>
# We clean the original textual data by removing non-alphanumeric characters, then we use the original Porter Stemmer to simplify the vocabulary variations, exclude stop words and numbers
# remove all non-alphanumeric characters
df['text_processed'] = df['text'].apply(
lambda s: ' '.join(
PorterStemmer().stem(word) for word in word_tokenize(re.sub(r'[^A-Za-z0-9 ]', ' ', s).lower())
if not word in stop_words and not word.isdigit()
)
)
print('Before:\n' + df['text'][42] + '\n\nAfter:\n' + str(df['text_processed'][42]))
# ## B. Feature Engineering <a class='anchor' id='features'></a>
print('Words:\n', word_tokenize(df['text_processed'][42]))
print('Count:\n', len(word_tokenize(df['text_processed'][42])))
# +
# number of words in a message
df['words_count'] = df.apply(
lambda row: len([word for word in word_tokenize(row['text_processed'])
if not word.isdigit()]), axis=1)
df.head()
# -
# We can see the most common text size in a message
print('ham:\n',
Counter(df.loc[df['label'] == 'ham']['words_count']).most_common(25))
print('spam:\n',
Counter(df.loc[df['label'] == 'spam']['words_count']).most_common(25))
# +
# number of characters in a message, i.e. message length (excluding spaces)
df['char_count'] = df['text_processed'].apply(
lambda s: len(re.sub(r'[ ]', '', s)))
df.head()
# -
# We can see the most common text length (number of characters) in a message for both labels
print('ham:\n',
Counter(df.loc[df['label'] == 'ham']['char_count']).most_common(25))
print('spam:\n',
Counter(df.loc[df['label'] == 'spam']['char_count']).most_common(25))
# We drop some of the rows in the data frame as their number of words/characters is less than 1 and the total number of such occurancies is 12 which will not significantly affect our dataset
df.loc[df['words_count'] < 1]
df.loc[df['char_count'] < 2]
# +
df.drop(df.loc[df['char_count'] < 2].index, inplace=True)
df.drop(df.loc[df['words_count'] < 1].index, inplace=True)
df.info()
# +
sns.distplot(
df.loc[df['label'] == 'ham']['words_count'],
bins=np.arange(0, 30),
hist_kws={'alpha': .75},
label='ham')
sns.distplot(
df.loc[df['label'] == 'spam']['words_count'],
bins=np.arange(0, 30),
hist_kws={'alpha': .75},
label='spam')
plt.xlim(0, 30)
plt.xlabel('Number of words')
plt.title('Words count distribution')
plt.legend()
plt.show()
# -
# In principle the KDEs have to be similar to the previous ones
# +
sns.distplot(
df.loc[df['label'] == 'ham']['char_count'],
bins=np.arange(0, 120, 5),
hist_kws={'alpha': .75},
label='ham')
sns.distplot(
df.loc[df['label'] == 'spam']['char_count'],
bins=np.arange(0, 120, 5),
hist_kws={'alpha': .75},
label='spam')
plt.xlim(0, 120)
plt.xlabel('Number of characters')
plt.title('Text length distribution')
plt.legend()
plt.show()
# -
# We can see some outliers after 140 characters and about 180 words
# +
sns.scatterplot(
x=list(Counter(df['words_count']).keys()),
y=list(Counter(df['words_count']).values()),
label='Words')
sns.scatterplot(
x=list(Counter(df['char_count']).keys()),
y=list(Counter(df['char_count']).values()),
label='Chars')
plt.xlim(-15, 500)
plt.ylim(-15, 500)
plt.title('Outliers for the number of words/characters')
plt.ylabel('Count')
plt.xlabel('Quantity')
plt.legend()
plt.show()
# -
# We now can calculate the average word length per text message as a feature and see the distribution of it
# +
# average word length in a text
df['avg_word_length'] = df.char_count / df.words_count
df.head()
# +
sns.distplot(
df.loc[df['label'] == 'ham']['avg_word_length'],
hist_kws={'alpha': .75},
label='ham')
sns.distplot(
df.loc[df['label'] == 'spam']['avg_word_length'],
hist_kws={'alpha': .75},
label='spam')
plt.xlim(0, 10)
plt.xlabel('Average word length')
plt.title('Average word length distribution')
plt.legend()
plt.show()
# -
# ### C. Sentiment Analysis <a class='anchor' id='sentiment'></a>
# For the sentiment analysis we use the original data without cleaning as the VADER sentiment analysis tool already takes care of the preprocessing and it is especially tailored to use for internet text lexicon as it is seen to be used in the text messages
# %%time
df = pd.concat([
df, df['text'].apply(
lambda s: SentimentIntensityAnalyzer().polarity_scores(s)).apply(
pd.Series)
],
axis=1)
# After we calculate the sentiment intensity we then mark the messages as positive/negative/neutral based on the compound component
# +
df.loc[(df['compound'] > -.05) &
(df['compound'] < .05), 'sentiment'] = 'neutral'
df.loc[df['compound'] <= -.05, 'sentiment'] = 'negative'
df.loc[df['compound'] >= .05, 'sentiment'] = 'positive'
df.head()
# -
# Now we can plot the results of the sentiment analysis and see the distribution for each label
print('overall:', Counter(df['sentiment']))
print('ham:', Counter(df.loc[df['label'] == 'ham']['sentiment']))
print('spam:', Counter(df.loc[df['label'] == 'spam']['sentiment']))
# +
sns.countplot(
x='label',
hue='sentiment',
data=df,
hue_order=['positive', 'negative', 'neutral'])
plt.title('Sentiment distribution per label')
plt.xlabel('Label')
plt.ylabel('Count')
plt.show()
# -
# Let's now see the most common words for both labels. For this we exclude stop words and plot a word cloud.
# +
df_plot = df.loc[df['label'] == 'ham']['text_processed']
word_cloud = WordCloud(
max_words=25, stopwords=stop_words, margin=10, width=1920,
height=1080).generate(str(df_plot.values))
plt.imshow(word_cloud, interpolation='bilinear')
plt.axis('off')
plt.tight_layout(pad=0)
plt.title('Most common words in ham texts')
plt.show()
# +
df_plot = df.loc[df['label'] == 'spam']['text_processed']
word_cloud = WordCloud(
max_words=25, stopwords=stop_words, margin=10, width=1920,
height=1080).generate(str(df_plot.values))
plt.imshow(word_cloud, interpolation='bilinear')
plt.axis('off')
plt.tight_layout(pad=0)
plt.title('Most common words in spam texts')
plt.show()
# -
# Finally we will add a new feature - first we create a list of most common words in spam messages and then for each text we count a number of words which fall in this list
Counter(word for word in word_tokenize(str(df_plot.values))).most_common(25)
spam_list = [
s for s, _ in Counter(
word for word in word_tokenize(str(df_plot.values))).most_common(25)
]
# +
df['spam_words'] = df['text_processed'].apply(
lambda s: len([word for word in word_tokenize(s) if word in spam_list]))
df.head()
# -
# # 3. Classification <a class='anchor' id='classification'></a>
# +
# encode labels
df['label'] = LabelEncoder().fit_transform(df['label'])
# vectorize the texts into bag of words feature
bow = CountVectorizer().fit_transform(df['text_processed'])
# drop textual columns for classification
df.drop(['text', 'sentiment'], axis=1, inplace=True)
df.head()
# -
# stack bag of words with other numerical features in a numpy array
supervised_features = sparse.hstack((bow, df.iloc[:, 2:].values))
print([x for x in supervised_features.toarray()[42] if x > 0])
# +
# split the dataset
X_train, X_test, y_train, y_test = train_test_split(
supervised_features, df['label'], test_size=.33)
print('X_train:', X_train.shape[0])
print('X_test:', X_test.shape[0])
print('y_train:', y_train.shape[0])
print('y_test:', y_test.shape[0])
# +
# K-Neighbours classsification
# fit the model
model = KNeighborsClassifier()
model.fit(X_train, y_train)
# run on the test set
y_knc = model.predict(X_test)
# -
# cross validation
scores = cross_val_score(model, supervised_features, df['label'], cv=5)
print('KNeighbors accuracy score:', accuracy_score(y_test, y_knc))
print('KNeighbors confusion matrix:\n', confusion_matrix(y_test, y_knc))
print('KNeighbors cross-validation scores:', scores)
print('KNeighbors final score after cross-validation: %0.2f (+/- %0.2f)' % (scores.mean(), scores.std() * 2))
# +
# Random Forest classification
# fit the model
model = RandomForestClassifier(n_estimators=42)
model.fit(X_train, y_train)
# run on the test set
y_rfc = model.predict(X_test)
# -
# cross validation
scores = cross_val_score(model, supervised_features, df['label'], cv=5)
print('Random Forest accuracy score:', accuracy_score(y_test, y_rfc))
print('Random Forest confusion matrix:\n', confusion_matrix(y_test, y_rfc))
print('Random Forest cross-validation scores:', scores)
print('Random Forest final score after cross-validation: %0.2f (+/- %0.2f)' % (scores.mean(), scores.std() * 2))
# In theory these results could be improved by using TFIDF vectors instead of BoW
| SMS-mining.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 说明
#
# ```
# "车辆位置数据":{
# "定位状态":"1",
# "经度":"4",
# "纬度":"4",
# }
# ```
# ## 导入工具包和公共函数
import Ipynb_importer
from public_fun import *
# ## 类主体
class fun_07_02_05(object):
def __init__(self, data):
self.cf = [1, 4, 4]
self.cf_a = hexlist2(self.cf)
data = data[2:]
self.o = data[0:self.cf_a[-1]]
self.list_o = [
"定位状态",
"经度",
"纬度",
]
self.oj = list2dict(self.o, self.list_o, self.cf_a)
self.oj2 = {'车辆位置数据':self.oj}
self.ol = pd.DataFrame([self.oj]).reindex(columns=self.list_o)
self.pj = {
'定位状态' : fun_07_02_05.fun_01(self.oj['定位状态']),
'经度' : hex2dec(self.oj['经度'], k=0.000001),
'纬度' : hex2dec(self.oj['纬度'], k=0.000001),
}
self.pj2 = {'车辆位置数据':self.pj}
self.pl = pd.DataFrame([self.pj]).reindex(columns=self.list_o)
self.next = data[len(self.o):]
self.nextMark = data[len(self.o):len(self.o)+2]
def fun_01(data):
n = '{:08b}'.format(int(data, 16))
state = n[-1]
longitude = n[-2]
latitude = n[-3]
if state == '0':
state_s = "定位有效"
else :
state_s = "定位无效"
if longitude == '0':
longitude_s = "北纬"
else :
longitude_s = "南纬"
if latitude == '0':
latitude_s = "东经"
else :
latitude_s = "西经"
output = [n, state_s, longitude_s, latitude_s]
return output
# ## 执行
# +
data = "0500072035A801EAC74A060179100E010D0FC801013C01043A0700000000000000000009010100483C3B3C3A3B3B3C3A3B3B3B3A3B3B3B3A3B3B3B3B3B3A3B3C3B3B3C3B3C3A3B3C3B3A3B3B3B3B3B3A3C3B3C3A3B3B3B3A3B3B3B3A3B3B3B3B3B3A3B3C3B3A3B3B3C3A3B3B3B3A3B3B0801011706273C00900001900FFA10040FFA0FFA0FFA0FFA10040FFA1004100410040FFA0FC8100410041004100410041004100410040FFA10041004100410041004100410040FFA10040FFA0FFA0FFA10040FFA0FFA0FFA0FFA100410040FFA0FFA100410040FFA0FFA10040FFA0FFA0FFA10040FFA0FFA0FFA0FFA0FFA0FFA0FF00FFA0FFA0FFA10040FFA0FFA100410040FF010040FFA0FFA0FFA0FFA0FFA10040FFA0FFA0FFA100410040FFA0FF00FFA0FFA0FFA0FFA0FFA0FFA0FFA10040FFA0FFA10041004100410040FFA0FFA0FFA0FFA0FFA0FFA0FFA0FFA0FFA100410040FFA0FFA1004100410040FFA0FFA0FFA0FFA0FFA0FFA0FFA0FFA100E0FF00FFA0FFA0FFA0FFA100410040FFA100410040FFA10040FFA10040FFA100410040FFA10041004100410040FFAAB"
fun_07_02_05 = fun_07_02_05(data)
# -
# ## 结果
fun_07_02_05.o
fun_07_02_05.oj
fun_07_02_05.ol
fun_07_02_05.pj
fun_07_02_05.pl
fun_07_02_05.next
fun_07_02_05.nextMark
| notebook/.ipynb_checkpoints/fun_07_02_05-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Point source extraction using CNN
# This is a tutorial about how you can train a convolutional filter to increase signal to noise ratio.
# +
import sys
import glob
import numpy as np
# Add pydeepkat directory
import deepsource9 as ds9
import matplotlib.pylab as plt
from matplotlib import gridspec
from matplotlib.colors import LogNorm
# %matplotlib inline
# -
# The data we want to increase its signal to noise are radio images. "PreProcessDataProvider" class makes random patches from them by specifiying container directories and names conditions of images and coresponding models files. Image and model file names will be fetched using glob. Model files includes gounrd truth of point source positions. You can make an instance like:
# +
def kernel1(cat):
return ds9.horn_kernel(cat,radius=15,step_height=2)
def kernel2(cat):
return ds9.gaussian_kernel(cat,sigma=10)
images_path=['../data/image'+str(i+1)+'.fits' for i in range(3)]
models_path=['../data/image'+str(i+1)+'.txt' for i in range(3)]
ppdp = ds9.GeneralDataProvider(images_path=images_path,models_path=models_path,margin=0,
nx=300, ny=300,kernels=[kernel1,kernel2], b_n=0.05, alpha=0.5)
# -
# nx and nx are patch size. You can specify the number of required patches and call the innstance to get a random patch. It returns a patch of image and corresponding demand. They look like:
# +
image,demand = ppdp(1)
gs = gridspec.GridSpec(1, 2)
plt.figure(figsize=(6,3))
ax = plt.subplot(gs[0, 0])
ax.imshow(image[0,:,:,0])
ax.set_title('Image')
plt.axis('off');
ax = plt.subplot(gs[0, 1])
ax.imshow(demand[0,:,:,0])
ax.set_title('Demand')
plt.axis('off');
plt.subplots_adjust(wspace=0.05)
# -
# Above, we do not use any kernel to produce demand. It is good to using a sequense of kernels to make more significant contribution for point sources in loss function. We can use horn shape and Gaussian kernels. These kernels operate like:
# +
d_example = np.zeros((41,41))
d_example[20,20] = 1
d_horn = ds9.horn_kernel(d_example,radius=15,step_height=2)
d_horn_gauss = ds9.gaussian_kernel(d_horn,sigma=7)
gs = gridspec.GridSpec(1, 3)
plt.figure(figsize=(9,3))
ax = plt.subplot(gs[0, 0])
ax.imshow(d_example)
ax.set_title('point source')
plt.axis('off');
ax = plt.subplot(gs[0, 1])
ax.imshow(d_horn)
ax.set_title('Horn')
plt.axis('off');
ax = plt.subplot(gs[0, 2])
ax.imshow(d_horn_gauss)
ax.set_title('Horn-gaussian')
plt.axis('off');
plt.subplots_adjust(wspace=0.05)
# -
# You can add any desired kernel to "PreProcessDataProvider":
# +
rb = 0
restore=0
patch_size = 400
def kernel1(cat):
return ds9.horn_kernel(cat,radius=10,step_height=2)
def kernel2(cat):
return ds9.gaussian_kernel(cat,sigma=7)
images_path=['../data/image'+str(i+1)+'.fits' for i in range(6)]
models_path=['../data/image'+str(i+1)+'.txt' for i in range(6)]
ppdp = ds9.GeneralDataProvider(images_path=images_path,models_path=models_path,margin=0,
nx=300, ny=300,kernels=[kernel1,kernel2], b_n=0.1, alpha=0.3)
x,y = ppdp(1)
gs = gridspec.GridSpec(1, 2)
plt.figure(figsize=(6,3))
ax = plt.subplot(gs[0, 0])
ax.imshow(x[0,:,:,0])
ax.set_title('Image')
plt.axis('off')
ax = plt.subplot(gs[0, 1])
ax.imshow(y[0,:,:,0])
ax.set_title('Demand')
plt.axis('off');
plt.subplots_adjust(wspace=0.05)
# -
# Then you can use "ppdp" to feed convolutional neural network (CNN). CNN will be trained to transform an image to corresponding demand. "ConvolutionalLayers" class provides a CNN for this purpose. You can easily specify any desired architecture and train/save/restore it. In bellow example there is an architecture defined in file "arch.py" placed in current directory. You can train you CNN using train method:
# +
rb = 0
restore=0
patch_size = 300
cnn = ds9.ConvolutionalLayers(nl=patch_size,learning_rate = 0.001,n_channel=1,
restore=1,model_add='./cmodel',arch_file_name='arch')
cnn.train(ppdp,training_epochs=100,n_s=1, dropout=0.2,verbose=5)
# -
# If you want to restore your trained model to use or tue up, you can set the "restore" argument to 1.
# Then "conv" methos can transform any image to demand:
# +
x,y = ppdp(1)
xf = cnn.conv(x)
gs = gridspec.GridSpec(1, 3)
plt.figure(figsize=(7.5,3.5))
ax = plt.subplot(gs[0, 0])
plt.axis('off')
ax.set_title('Data')
ax.imshow(x[0,10:-10,10:-10,0])
ax = plt.subplot(gs[0, 1])
plt.axis('off')
ax.set_title('CNN-output')
ax.imshow(xf[10:-10,10:-10])
ax = plt.subplot(gs[0, 2])
plt.axis('off')
ax.set_title('Demand')
ax.imshow(y[0,10:-10,10:-10,0])
plt.subplots_adjust(wspace=0.05,bottom=0.0, top=0.92, left=0.02, right=0.98)
# -
# Now let's transform a whole image and extract point sources from demand map. Since operating on a large image may consume big amount of memory, there is a method to do this job by tiling.
# +
image_file = '../data/image7.fits'
model_file = '../data/image7.txt'
catalog = ds9.ps_extract(image_file,model_file,cnn,ds9.fetch_data,loc_det='mean',
ignore_border=0,jump_lim=3,area_lim=10,lw=300,pad=10)
# -
ds9.visualize_cross_match(image_file,model_file,catalog)
catalog.shape
# +
#_,number_of_sources = ds9.full_completeness_purity(image_file,model_file,catalog,output_csv_file,
# ignore_border=500, quality_threshold=0.9)
for i in range(7,11):
tp_df, fp_df, fn_df = ds9.do_crossmatch_2(catalog[:,0].astype(int),
catalog[:,1].astype(int),
1.,
'../data/',
'image'+str(i)+'.fits',
'../data/image'+str(i)+'.txt',
'',
input_format='xy')
# calculate completeness & purity
# cp_path = os.path.join(cp_output_dir_path, image_names[ii]+"_cp.csv")
snr_edges, snr_centers, purity, completeness = ds9.calc_completeness_purity(tp_df,
fp_df,
fn_df,
save_path='./catalogs/catalog'+str(i)+'.csv')
# print number_of_sources
# mean 394
# peak 378
# com 388
# +
csv_list = glob.glob('./catalogs/*.csv')
import matplotlib.pylab as plt
from matplotlib import gridspec
gs = gridspec.GridSpec(1, 2)
plt.figure(figsize =(15,5))
ax0 = plt.subplot(gs[0, 0])
ax1 = plt.subplot(gs[0, 1])
print ds9.PC_mean_plot(ax0,ax1,csv_list,do_labels=True,s2n=4.)
ax0.set_xscale('log')
ax0.set_xlim(1,8)
ax0.set_ylim(0,1.1)
ax0.set_xticks([3,5,8])
ax0.set_xticklabels([3,5,8])
ax1.set_xscale('log')
ax1.set_xlim(1,8)
ax1.set_ylim(0,1.1)
ax1.set_xticks([3,5,8])
ax1.set_xticklabels([3,5,8])
plt.legend(loc='best')
# mean (0.6566051116677564, 4.6390907497292)
# peak (0.9341329225922596, 3.9212680362675005)
# com (0.8750727917088663, 3.926884174631457)
# -
def maker(d,radius,lst,original_value,set_value):
nn = []
for i,j in lst:
rr, cc = draw.circle(i, j, radius=radius, shape=d.shape)
nn.append(d[rr, cc].sum())
un = ds9.circle(d,radius=radius,original_value=original_value,set_value=set_value)\
+ds9.circle(d,radius=radius+1,original_value=original_value,set_value=set_value)
return un,np.array(nn)
# +
image_file = '../data/image7.fits'
model_file = '../data/image7.txt'
catalog = ds9.ps_extract(image_file,model_file,cnn,ds9.fetch_data,loc_det='mean',
ignore_border=0,jump_lim=3,area_lim=10,lw=300,pad=10)
image, x_true, y_true = ds9.fetch_data(image_file,model_file)
image_size = image.shape[2]
cnn_output = cnn.conv_large_image(image,lw=300)
# -
from skimage import draw
nn = []
mag = np.zeros(cnn_output.shape)
for i,j in catalog[:,:2].astype(int):
rr, cc = draw.circle_perimeter(j, i, radius=5, shape=mag.shape)
nn.append(cnn_output[rr, cc].sum())
mag[rr, cc] = 1
nn = np.array(nn)
lst = catalog[:,:2]
n_ps = catalog.shape[0]
# +
prob = []
true_fraction = []
for radius in range(10):
un,nn = maker(cnn_output,radius,lst,dmax,dmax)
nn = 0.00002041*nn
prob.append(nn)
n_in = 0
for i in range(n_ps):
if np.sqrt((lst[i][0]-x_true[i])**2+(lst[i][1]-y_true[i])**2)<radius:
n_in += 1
n_in = 1.*n_in/n_ps
true_fraction.append(n_in)
true_fraction = np.array(true_fraction)
prob = np.array(prob)
# -
prob.shape
# +
plt.plot(np.mean(prob,axis=1),true_fraction,marker='o')
plt.ylabel(r'$Fraction\ of\ true\ sources\ within\ CR_P$')
plt.xlabel(r'$Credible\ level\ P$')
plt.savefig('CR_P.jpg',dpi=300)
# -
gs = gridspec.GridSpec(1, 1)
plt.figure(figsize =(15,15))
ax = plt.subplot(gs[0, 0])
ax.imshow(mag+cnn_output)
# +
d = np.zeros((image_size,image_size))
for i,j in catalog[:,:2].astype(int):
d[i,j] = 1
d = ds9.gaussian_kernel(d,sigma=10)
dmax = d.max()
d /= dmax
dmax = d.max()
plt.imshow(d)
# +
prob = []
true_fraction = []
for radius in range(20):
un,nn = maker(d,radius,lst,dmax,dmax)
nn = 0.00002041*nn
prob.append(nn)
n_in = 0
for i in range(n_ps):
if np.sqrt((lst[i][0]-x_true[i])**2+(lst[i][1]-y_true[i])**2)<radius:
n_in += 1
n_in = 1.*n_in/n_ps
true_fraction.append(n_in)
true_fraction = np.array(true_fraction)
prob = np.array(prob)
# +
plt.plot(np.mean(prob,axis=1),true_fraction,marker='o')
plt.ylabel(r'$Fraction\ of\ true\ sources\ within\ CR_P$')
plt.xlabel(r'$Credible\ level\ P$')
plt.savefig('CR_P2.jpg',dpi=300)
# -
# +
import matplotlib.pylab as plt
from matplotlib import gridspec
gs = gridspec.GridSpec(1, 2)
plt.figure(figsize =(8,3))
ax0 = plt.subplot(gs[0, 0])
ax1 = plt.subplot(gs[0, 1])
rb = '0'
csv_list = glob.glob('/home/gf/work/forsat/AIMS_works/radio_source/results/different/res/ps_cat'+rb+'/'+rb+'_*.csv')
csv_list = ['/home/gf/work/forsat/AIMS_works/radio_source/results/different/res/ps_cat'+rb+'/'+rb+'_'+str(i)+
'.csv' for i in range(101,152)]
pc5_cnn,pcc_cnn = pk.PC_mean_plot(ax0,ax1,csv_list,do_labels=True,s2n=4.,clr='b')
# csv_list = glob.glob('/home/gf/work/forsat/AIMS_works/radio_source/results/different/\
# different_field_2sig-isl_1sig-peak_cropped-500-3600_robust0_cm-9-2p5-2_cp/\
# stimela-image_image*_25SEP_robust-'+rb+'-image_cp.csv')
# csv_list = ['/home/gf/work/forsat/AIMS_works/radio_source/results/different/\
# different_field_2sig-isl_1sig-peak_cropped-500-3600_robust'+rb+'_cm-9-2p5-2_cp/\
# stimela-image_image'+str(i)+'_25SEP_robust-'+rb+'-image_cp.csv' for i in range(101,152)]
csv_list = ['/home/gf/work/forsat/AIMS_works/radio_source/results/samefield/res_same/ps_cat'+rb+'/'+rb+'_'+str(i)+
'.csv' for i in range(101,152)]
pc5_bdsf,pcc_bdsf = pk.PC_mean_plot(ax0,ax1,csv_list,do_labels=False,s2n=4.,clr='r')
ax0.set_xscale('log')
ax0.set_xlim(1,8)
ax0.set_ylim(0,1.1)
ax0.set_xticks([3,5,8])
ax0.set_xticklabels([3,5,8])
ax1.set_xscale('log')
ax1.set_xlim(1,8)
ax1.set_ylim(0,1.1)
ax1.set_xticks([3,5,8])
ax1.set_xticklabels([3,5,8])
plt.legend(loc='best')
plt.suptitle('robust {}: PC4-CNN-diff={:2.2f} (blue), PC4-CNN-same={:2.2f} (red)'.format(rb,pc5_cnn,pc5_bdsf))
plt.subplots_adjust(wspace=0.2,bottom=0.14, top=0.92, left=0.07, right=0.98)
plt.savefig('diff_pybdsf-robust-'+rb+'.jpg')
# -
print pc5_bdsf,pc5_cnn
print pcc_bdsf,pcc_cnn
# +
np.round(np.mean(np.argwhere(filt),axis=0)).astype(int)
# +
d = np.zeros((100,100))
for i in range(100):
for j in range(100):
d[i,j]=np.exp(-0.0001*((i-50)**2+(j-50)**2))
d[59,59]=11
d[60:80,60:80]=10
plt.imshow(d)
filt = d>0.9
# plt.imshow(filt)
print np.round(np.mean(np.argwhere(filt),axis=0)).astype(int)
print np.array([np.where(d==np.max(d[filt]))[0][0],np.where(d==np.max(d[filt]))[1][0]]).astype(int)
print cent_of_mass(d,filt)
# -
def cent_of_mass(d,filt):
indx = np.where(filt)[0]
indy = np.where(filt)[1]
li = 0
lj = 0
mass = 0
for i,j in zip(indx,indy):
val = d[i,j]
mass += val
li += val*i
lj += val*j
li /= mass
lj /= mass
return np.round(np.array([li,lj])).astype(int)
| notebooks/Convolutional_Layers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# In this notebook we implement preprocessing for the next step, blob detection. Since we need the images in many formats, it's probably easiest to build a class from which we can access them.
# +
import cv2 as cv
import numpy as np
import matplotlib.pyplot as plt
# %load_ext autoreload
# %autoreload 2
# -
# # Building class
class Frame:
"""Class for each frame of the movie containing active area mask
and can return grayscale, apply mask etc. A lot easier to have a single
object instead of frame_grayscale etc."""
def __init__(self, image, mask):
self.mask = mask
self.image = image
def __call__(self, gray=True, masked=True):
""" Returns grayscale masked image."""
image = self.image
if gray is True:
image = self.grayscale(image)
if (masked is True):
image = self.apply_mask(image, self.mask)
return image
def grayscale(self, image):
"""Returns grayscale of image."""
grayscale_image = cv.cvtColor(image, cv.COLOR_BGR2GRAY)
return grayscale_image
def apply_mask(self, image, mask):
"""Returns masked image, only works on grayscale image"""
masked_image = image
np.putmask(masked_image, ~mask, 255)
return masked_image
# # testing
video_path = '../../data/movies/4arenas_QR.h264'
capture = cv.VideoCapture(video_path)
im = capture.read()[1]
# +
# Building mask, this will be better later
mask = np.ones(im.shape[:2], dtype=np.bool)
mask[:180, :300] = 0 # upper left corner
mask[-230:, :300] = 0 # lower left corner
mask[-230:, -300:] = 0 # lower right corner
mask[:180, -300:] = 0 # lower right corner
mask[:70, :] = 0
mask[-120:, :] = 0
mask[:, :180] = 0
mask[:, -200:] = 0
# -
frame = Frame(image=im, mask=mask)
plt.imshow(frame(gray=True, masked=False), cmap='gray')
plt.imshow(frame.image, cmap='gray')
im.shape
| dev/Implementing/preprocessing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="Qv8_Q9knv3om"
# 
#
# __Autor__: <NAME> (<EMAIL>)
# + [markdown] colab_type="text" id="FC9NvpcO7XjW"
# # Pensamento estatístico em Python
#
# Neste módulo falaremos sobre testes de hipóteses.
#
# 
# + colab={} colab_type="code" id="HK2aSUXU9I_s"
from math import sqrt
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import statsmodels.api as sm
import scipy.stats as sct
import seaborn as sns
# + colab={} colab_type="code" id="GsoiY5gw9KJ_"
# Algumas configurações para o matplotlib.
# %matplotlib inline
from IPython.core.pylabtools import figsize
figsize(12, 8)
sns.set()
# + [markdown] colab_type="text" id="9U2fS1OAKHCB"
# ## Testes de hipóteses
# + [markdown] colab_type="text" id="jXgrQ3Ef9FJ5"
# ### Introdução
#
# Já vimos antes como podemos fazer estimativas para parâmetros populacionais a partir de amostras da população. Agora veremos como testar hipóteses sobre parâmetros populacionais, incluindo sua distribuição. Testes de hipóteses são as principais ferramentas da inferência estatística para isso.
#
# Em um teste de hipóteses, formulamos duas hipóteses __complementares__ a respeito de um parâmetro populacional de interesse chamadas hipótese nula (_null hypothesis_) e hipótese alternativa (_alternative hypothesis_). Denotamos a hipótese nula por $H_{0}$ e a hipótese alternativa por $H_{1}$.
#
# __Exemplo__:
#
# Podemos estar interessados na média dos pesos de determinada população ($\mu$) e queremos testar se seu valor verdadeiro é 70 kg a partir de uma amostra coletada. Para isso, geramos as seguintes hipóteses:
#
# $$
# \begin{cases}
# H_{0}: \mu = 70 & \quad \text{(Hipótese nula)} \\
# H_{1}: \mu \neq 70 & \quad \text{(Hipótese alternativa)}
# \end{cases}
# $$
#
# Então seguimos um procedimento para avaliar se devemos:
#
# 1. Rejeitar a hipótese nula em favor da hipótese alternativa. Nesse caso, diríamos que a verdadeira média populacional _não_ é 70 kg, mas não conseguiríamos afimar qual é o seu verdadadeiro valor.
# 2. Não rejeitar a hipótese nula, mantendo-a. Nesse caso, diríamos que não temos evidências o suficiente para rejeitar a hipótese de que a verdadeira média populacional é 70 kg. No entanto, isso não significa que a média deva ser de fato 70 kg, mas sim que a nossa amostra parece sustentar essa ideia.
#
# > Apesar de comum, é incorreto dizer que "aceitamos a hipótese nula". Na verdade, simplesmente __não__ a rejeitamos por falta de evidências.
#
# Algumas observações sobre as hipóteses acima:
#
# * Notem como as duas hipóteses são complementares.
# * As hipóteses são feitas sobre o parâmetro populacional ($\mu$) e não sobre o estimador amostral (que poderia ser $\bar{X}$).
# * Só existem duas hipóteses. Não podemos gerar múltiplas hipóteses simultaneamente.
# * O resultado do teste não nos diz nada sobre nossa teoria, e sim sobre o que os dados indicam.
# + [markdown] colab_type="text" id="ixmJD-GOH3OX"
# ### Funcionamento
#
# Para seguir com o nosso teste de hipóteses (TH), devemos coletar uma amostra da população e trabalhar com algum estimador do parâmetro populacional sob estudo. No caso acima, podemos utilizar a média amostral ($\bar{X}$) que é o melhor estimador para a média populacional.
#
# Imagine que a média da amostral foi 74 kg. Note como isso pode ocorrer mesmo quando a verdadeira média populacional é 70 kg, pois a amostra é aleatória, e para cada amostra aleatória, obteríamos um valor diferente para a média.
#
# A questão é: essa diferença de 4 kg foi devido ao acaso (devido a aleatoriedade da amostra) ou porque a média populacional não é mesmo 70 kg (talvez 73 kg)? É para responder a esse tipo de questão que usamos o TH.
# + [markdown] colab_type="text" id="gh85LB7jKEWQ"
# __Todo procedimento de um TH é feito considerando a $H_{0}$ verdadeira__.
#
# Podemos considerar inicialmente que a verdadeira média populacional é de fato $\mu = 70$ e que os pesos são normalmente distribuídos (depois veremos como relaxar essa suposição).
#
# Sendo isso verdade, as médias de pesos das amostras devem se distribuir normalmente em torno de 70 kg. O que devemos então fazer é definir uma região onde ainda consideramos aceitável presumir que a verdadeira média é 70 kg. Tudo que estiver fora dessa região é considerado "muito longe" para que a verdadeira média seja 70 kg.
#
# A primeira região (a que sustenta a hipótese da média real 70 kg) é chamada __região de aceitação__ (RA), e tudo que estiver fora dela é chamado __região crítica__ (RC).
#
# Por exemplo, podemos definir RA como sendo o intervalo de 68 kg a 72 kg, ou seja, consideramos que qualquer diferença de 2kg ou menos de 70 kg é devido ao acaso. Qualquer valor fora desse intervalo já é longe demais de 70 kg para que esta seja a verdadeira média. Nesse cenário, a nossa média amostral de 74 kg cai na RC e portanto rejeitaríamos a hipótese nula.
#
# O que precisamos agora é de um meio formal de definir essas regiões. Para isso, utilizaremos a informação de que, sob a hipótese nula, a média amostral $\bar{X}$ tem distribuição normal em torno de $\mu = 70$. E em vez de definirmos o tamanho da região de aceitação, definimos o tamanho da região crítica.
# + [markdown] colab_type="text" id="TQigeNcbTF9u"
# Todo TH conta com uma estatística de teste (vamos chamá-la de $T$), que é gerada a partir da amostra. A partir dessa estatística de teste e de sua distribuição, podemos definir RA e RC em termos de probabilidade.
#
# Por exemplo, podemos construir essas regiões de forma que, se $H_{0}$ for verdadeira, então $T$ tem 5% de probabilidade de cair na RC. Essa probabilidade de cair na região crítica, sendo $H_{0}$ verdadeira, é uma probabilidade de erro. Esse erro é chamado de Erro Tipo I e sua probabilidade é chamada __nível de significância__ e denotada por $\alpha$.
#
# Podemos cometer outro tipo de erro ao não rejeitarmos $H_{0}$ quando ela é realmente falsa. Esse é o Erro Tipo II e sua probabilidade é denotada por $\beta$.
#
# Em resumo:
#
# $$\alpha = P(\text{Erro Tipo I}) = P(\text{rejeitar } H_{0} | H_{0} \text{ verdadeira})$$
# $$\beta = P(\text{Erro Tipo II}) = P(\text{não rejeitar } H_{0} | H_{0} \text{ falsa})$$
#
# > $\alpha$ e $\beta$ não possuem relação matemática.
#
# __Quando criamos um TH, devemos decidir *a priori* o valor de $\alpha$__. Ele será nossa base de comparação para rejeitarmos ou não a $H_{0}$. Não fazer isso é chamado _p-value hacking_.
#
# Valores típicos de $\alpha$ são 0.025, 0.05 e 0.10.
# + [markdown] colab_type="text" id="ttPP2_m7gBzX"
# ### Classificação do teste de hipóteses
#
# Os TH podem ser classificados em:
#
# * Bilaterais: quando a região crítica encontra-se dos dois lados da distribuição de $T$ sob $H_{0}$.
# * Unilaterais: quando a região crítica encontra-se somente de um dos lados da distribuição de $T$ sob $H_{0}$.
#
# Quando o TH é bilateral, a probabilidade $\alpha$ geralmente é dividida em duas partes iguais, uma em cada lado da distribuição. Quando o TH é unilateral, toda probabilidade acumula-se em um dos lados.
#
# Também podemos falar em hipóteses alternativas simples e compostas:
#
# * Simples: quando não desigualdade.
# * Composta: quando há desigualdade.
#
# Na figura a seguir, consideramos que $H_{0}: \mu = \mu_{0}$ e mostramos o caso bilateral e dois unilaterais:
#
# 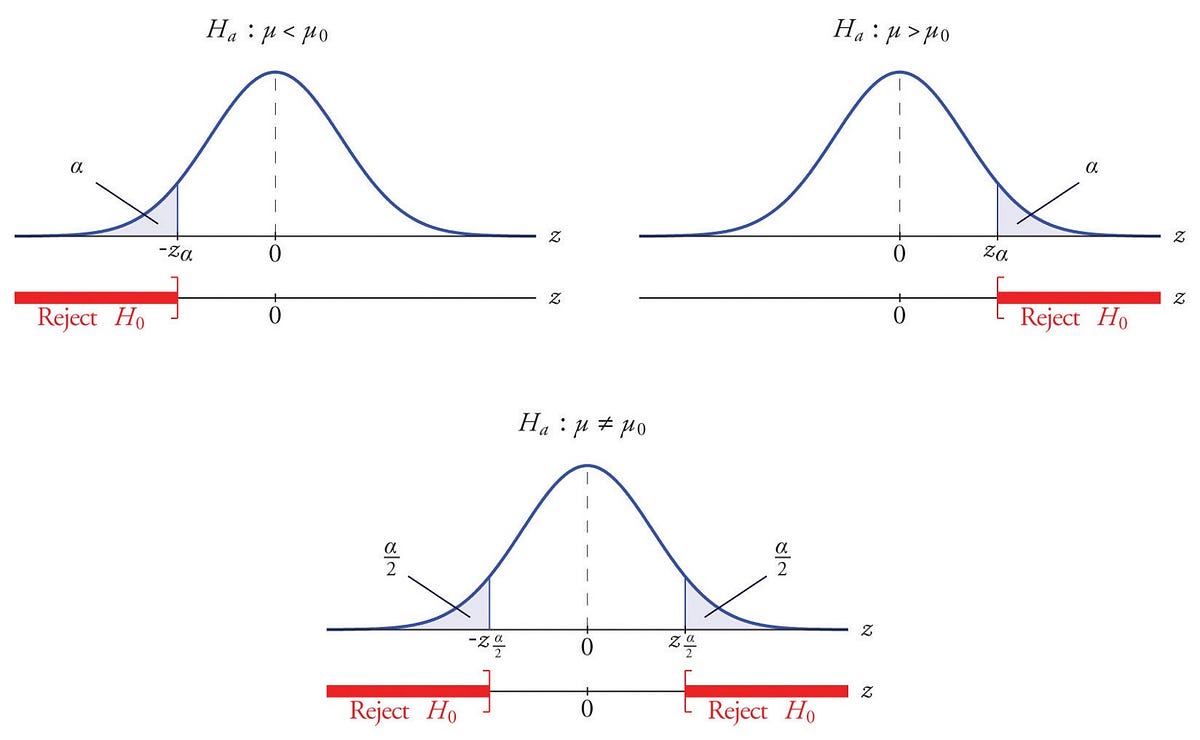
# + [markdown] colab_type="text" id="UZAHGAvvWJJx"
# ### Valor-p
#
# O valor-p (do Inglês, _p-value_), também chamado de _nível descritivo_, de um TH é um valor associado ao resultado, $t^{*}$, da estatística de teste $T$ sob $H_{0}$.
#
# __O valor-p é a probabilidade de $T$ tomar um valor igual ou mais extremo que $t^{*}$, sendo $H_{0}$ verdadeira__.
#
# Obviamente, se essa probabilidade for muito baixa, podemos interpretar que é muito raro encontrarmos $t^{*}$ se $H_{0}$ for realmente verdadeira. Por outro lado, se ela for alta, podemos concluir que deve ser razoável encontrarmos esse valor de $t^{*}$ quando $H_{0}$ é verdadeira.
#
# Mas qual o limiar? O que é considerada uma probabilidade baixa ou alta? É aí que entra o $\alpha$ novamente!
#
# O nosso limiar é o valor de $\alpha$ estabelecido:
#
# * Se o valor-p for menor que $\alpha$, então $t^{*}$ caiu dentro da região crítica, e portanto devemos rejeitar $H_{0}$.
# * Se o valor-p for maior que $\alpha$, então $t^{*}$ caiu na região de aceitação e devemos não rejeitar $H_{0}$.
#
# Essa é a importância de estabelecermos $\alpha$ antes do experimento. Do contrário, poderíamos ajustar o valor de $\alpha$ para atender nossas expectativas sobre o resultado.
#
# É importante notar que o valor-p faz sentido no contexto da estatística frequentista, ou seja, considerando a probabilidade no "longo prazo". Além disso, ele nada nos afirma sobre a teoria sendo testada, apenas o que os dados dizem.
#
# Também temos outra interpretação para o valor-p: __O valor-p é o menor nível de significância, $\alpha$, para o qual rejeitaríamos $H_{0}$__.
#
# 
#
# É importante notar que o valor-p é usado extensivamente na estatística frequentista, mas a estatística Bayesiana possui outra abordagem que dispensa o valor-p.
#
# O valor-p é considerado perigoso, pois muitas pessoas não sabem utilizá-lo adequadamente, nem interpretá-lo corretamente, levando a uma série de conclusões duvidosas.
#
# __Lembre-se: o valor-p nos permite fazer afimarções sobre os dados, não sobre a teoria sendo testada__.
#
# Mas como o próprio valor-p se distribui?
#
# De forma geral:
#
# * Quando $H_{0}$ é realmente falsa, a distribuição do valor-p depende do poder do teste, ou seja, da capacidade do teste de detectar uma $H_{0}$ falsa. Quanto maior o poder do teste, maior a chance de obtermos um $\alpha$ pequeno ($< 0.05$).
#
# 
#
# * Quando $H_{0}$ é realmente verdadeira, o valor-p tem distribuição uniforme, com $100\alpha\%$ dos valores-p sendo menores que $\alpha$. Em outras palavras, temos $100\alpha\%$ de chance de cometermos um Erro Tipo I.
#
# 
#
# Vamos fazer simulações de Monte-Carlo para mostrar isso:
# + colab={"base_uri": "https://localhost:8080/", "height": 600} colab_type="code" id="reI10ijK_yEc" outputId="151a3018-2486-49dd-ff4d-0ea90588aaf4"
# H_0 é falsa.
pvalues1 = []
for i in range(1000):
pvalues1.append(sct.ttest_1samp(sct.norm.rvs(loc=10, scale=5, size=100), popmean=12).pvalue) # Menor poder.
pvalues2 = []
for i in range(1000):
pvalues2.append(sct.ttest_1samp(sct.norm.rvs(loc=10, scale=5, size=100), popmean=8).pvalue) # Maior poder.
fig, axs = plt.subplots(1, 2, figsize=(20, 10))
sns.distplot(pvalues1, kde=False, bins=20, hist_kws={"density": True}, ax=axs[0])
sns.distplot(pvalues2, kde=False, bins=20, hist_kws={"density": True}, ax=axs[1]);
# + colab={"base_uri": "https://localhost:8080/", "height": 491} colab_type="code" id="W4q6OtvwCHnT" outputId="14fb01f3-7288-4ba8-8f1b-26edb91e8d72"
# H_0 é verdadeira.
pvalues = []
for i in range(1000):
pvalues.append(sct.ttest_1samp(sct.norm.rvs(loc=10, scale=5, size=100), popmean=10).pvalue)
sns.distplot(pvalues, kde=False, bins=20, hist_kws={"density": True});
# + [markdown] colab_type="text" id="AdvPFicghiVP"
# ### Poder
#
# Definimos o poder de um teste como a probabilidade de rejeitarmos $H_{0}$ quando $H_{0}$ é realmente falsa. Em outras palavras, o poder é a probabilidade de não cometermos um Erro Tipo II:
#
# $$\text{Poder} = \pi(\mu) = P(\text{rejeitar } H_{0} | H_{0} \text{ falsa}) = 1 - \beta$$
#
# O poder é bastante influenciado pelo tamanho da amostra, então cuidado com interpretações sobre ele.
# + [markdown] colab_type="text" id="uJWHMRWjOGfQ"
# ## _Q-Q plot_
#
# O _q-q plot_ é um gráfico para comparação de distribuições de probabilidades. Geralmente, uma das distribuições é teórica e com distribuição bem conhecida. Essa distribuição teórica é convencionalmente posta no eixo x. No eixo y, colocamos os quantis da nossa distribuição amostral, que gostaríamos de comparar com a teórica.
#
# 
#
# Para o caso mais comum onde em y temos a amostra e em x temos a distribuição teórica com a qual queremos comparar, podemos pensar da seguinte forma:
#
# Sendo a amostra $S = \{s_{1}, s_{2}, \dots, s_{n}\}$, um ponto $(x, y)$ de um _q-q plot_ é tal que:
#
# $$
# \begin{cases}
# x = F^{-1}(G(s)) \\
# y = s
# \end{cases}
# $$
#
# sendo $F^{-1}$ a função quantil (inversa da CDF) da distribuição teórica, $G$ a CDF empírica da amostra e $s = s_{1}, s_{2}, \dots, s_{n}$.
#
# De qualquer forma, a cada ponto $(x_{i}, y_{i})$ do _q-q plot_ vale a relação $F(x_{i}) = G(y_{i})$.
#
# Se as duas distribuições a serem comparadas são de amostras, o raciocínio permanece muito parecido.
# + [markdown] colab_type="text" id="utv4UirwXWjS"
# Se as duas distribuições forem iguais, ou seja, $F = G$, então os pontos ficarão em cima da reta $y = x$ (inclinada 45º). Quanto mais alinhados os pontos estiverem em cima dessa reta, mais a distribuição da amostra se aproxima da distribuição teórica. Se os pontos ficarem em cima de uma outra reta, mas alinhados, pode ser que as distribuições estejam somente fora de escala. Nesse caso, pode ser interessante transformar uma das distribuições, por exemplo, padronizando a amostra (para ter média 0 e variância 1).
#
# O _q-q plot_ é uma boa forma gráfica de sabermos se as duas distribuições são iguais ou parecidas. A sua intuição é também utilizada em alguns testes de hipóteses para aderência à distribuições teóricas como o teste de normalidade de Jarque-Bera.
# + [markdown] colab_type="text" id="6LS4avxFK02T"
# __Q-Q plot para dados normais__:
# + colab={"base_uri": "https://localhost:8080/", "height": 508} colab_type="code" id="JZq5F2apKXeC" outputId="be32059e-b508-4475-d913-6d3f45012856"
normal_data = sct.norm.rvs(loc=10, scale=4, size=1000)
sm.qqplot(normal_data, fit=True, line="45");
# + [markdown] colab_type="text" id="xp0bvdpfLJF3"
# __Q-Q plot para dados não normais__:
# + colab={"base_uri": "https://localhost:8080/", "height": 508} colab_type="code" id="scSaAzuqLMhH" outputId="f9f8d593-b171-4cb2-9d21-8b08448d94de"
non_normal_data = sct.expon.rvs(size=1000)
sm.qqplot(non_normal_data, fit=True, line="45");
# + [markdown] colab_type="text" id="-6mP3jQoZK0z"
# ## Testes de hipóteses clássicos
#
# Diversos testes de hipóteses para uma gama de tarefas já foram desenvolvidos pela comunidade estatística. Por ora, nosso trabalho é somente entender e saber aplicar os mais usuais. A teoria formal sobre a construção de TH pode ser encontrada facilmente em qualquer literatura sobre inferência estatística.
#
# Dois tipos bem comuns de TH são os testes de comparação e os testes de aderência (_goodness-of-fit_). Veremos aqui dois testes para comparação de médias e dois testes de normalidade: Shapiro-Wilk e Jarque-Bera.
#
# Antes de prosseguirmos, um aviso por <NAME> Pearson:
#
# > _Statistical tests should be used with discretion and understanding, and not as instruments which themeselves give the final verdict_.
# + [markdown] colab_type="text" id="Bs8KSKJsg2zE"
# ### Teste de média de uma amostra
#
# O teste de hipóteses mais conhecido certamente é sobre a média de uma amostra: o famoso teste-$t$ da média.
#
# Nosso objetivo com esse teste é avaliar se uma dada amostra $S$ foi coletada de uma distribuição cuja média $\mu$ é igual a $\mu_{0}$.
#
# Podemos formular nossas hipóteses da seguinte forma:
#
# $$
# \begin{cases}
# H_{0}: \mu = \mu_{0} \\
# H_{1}: \mu \neq \mu_{0}
# \end{cases}
# $$
#
# Partimos de uma suposição inicial: nossa distribuição original é normal sob $H_{0}$ (depois veremos que é possível relaxar isso).
#
# A partir dessa suposição, podemos afimar que nossa média amostral $\bar{X}$ também tem distribuição simétrica de média $\mu_{0}$ e desvio-padrão $s/\sqrt{n}$, chamado erro-padrão.
#
# Para construir nossa estatística de teste $t$, fazemos:
#
# $$t = \frac{\bar{x} - \mu_{0}}{s/\sqrt{n}}$$
#
# onde $\bar{x}$ é a média obtida a partir da amostra, $s$ é o desvio-padrão calculado a partir da amostra e $n$ é o tamanho da amostra.
#
# Essa estatística $t$ tem distribuição $t$-Student com $n-1$ graus de liberdade, que é bem próxima da distribuição normal. Ela é utilizada em vez da normal, pois suas caudas mais pesadas compensam a aproximação feita de $s$ para o desvio padrão.
#
# Como sempre devemos estabelecer _a priori_ o valor de $\alpha$, nosso nível de significância, a fim de compararmos com valor numérico obtido de $t$. Se $t$ cair na região de aceitação (que depende de $\alpha$), então não rejeitamos a hipótese de que a verdadeira média é $\mu_{0}$. Do contrário, podemos dizer que temos evidências o suficiente para rejeitar tal hipótese, e portanto $\mu$ não deve ser igual a $\mu_{0}$.
#
# Para relaxar a suposição de distribuição normal dos dados, apelamos para o Teorema Central do Limite (TCL) que nos afirma que:
#
# $$\bar{X} \xrightarrow{d} N(\mu, \frac{\sigma^{2}}{n})$$
#
# quando $n \rightarrow \infty$.
#
# Ou seja, se tivermos uma amostra grande o suficiente, podemos usar o TCL para justificar os cálculos anteriores, incluindo a fórmula da estatística de teste $t$, sem precisar presumir normalidade dos dados.
#
# Para isso, sob $H_{0}$, substituímos $\mu$ por $\mu_{0}$ e estimamos $\sigma$ como $s$ (desvio-padrão amostral), chegando à mesma fórmula de $t$. Além disso, a distribuição $t$-Student se aproxima de uma distribuição normal quando $n \rightarrow \infty$.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="c20v0EEuMsdl" outputId="d2c9298e-5676-4409-f887-642bcc5dcd9f"
data = sct.expon.rvs(scale=10, size=1000) # Mean = scale = 1/lambda = 10.
sct.ttest_1samp(data, popmean=10) # Deveria não rejeitar H_0.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="ln4Z3Iy3OlpI" outputId="6f01f421-3ba0-460a-e197-30bdba035b6e"
data = sct.expon.rvs(scale=8, size=1000) # Mean = scale = 1/lambda = 8.
sct.ttest_1samp(data, popmean=10) # Deveria rejeitar H_0.
# + [markdown] colab_type="text" id="x_fccwEBlvHZ"
# ### Teste de comparação das médias de duas amostras
#
# Outro cenário bastante comum é querermos comparar as médias de duas amostras diferentes para descobrirmos se as duas amostras vêm de distribuições de mesma média.
#
# Entendido o teste anterior, o racional do presente teste é bem direto: usamos a diferença entre as duas médias amostrais e os respectivos desvios-padrão no cálculo da estatística de teste $t$. Ou seja,
#
# $$t = \frac{\bar{x}_{1} - \bar{x}_{2}}{\sqrt{s_{1}^{2} + s_{2}^{2}}}$$
#
# onde $\bar{x}_{1}$ e $\bar{x}_{2}$ são as médias da primeira e segunda amostras e $s_{1}$ e $s_{2}$ são os desvios-padrão das duas amostras.
#
# Sob $H_{0}$, é possível mostrar que $t$ tem distribuição $t$-Student com $n_{1} + n_{2} - 2$ graus de liberdade, onde $n_{1}$ e $n_{2}$ são os tamanhos das amostras.
#
# A interpretação do resultado de $t$ com relação ao nível de significância e consequente rejeição (ou não) de $H_{0}$ é análoga ao teste anterior.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="fr4Y2bzdMLVD" outputId="aa85e2b2-3ba8-4561-90ad-36a97f046417"
data_one = sct.expon.rvs(scale=8, size=100) # Mesmas distribuições.
data_two = sct.expon.rvs(scale=8, size=100)
sct.ttest_ind(data_one, data_two) # Não deveria rejeitar H_0.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="ejx3ybHxPU7w" outputId="002b27e6-bc86-4fba-8b4b-8cf9d8ab2566"
data_one = sct.expon.rvs(scale=8, size=100) # Distribuições diferentes.
data_two = sct.expon.rvs(scale=12, size=100)
sct.ttest_ind(data_one, data_two) # Deveria rejeitar H_0.
# + [markdown] colab_type="text" id="C2Xf3GlJsDbp"
# ### Shapiro-Wilk
#
# O teste de Shapiro-Wilk é um teste de aderência à distribuição normal, que abreviamos para teste de normalidade. Nosso intuito é verificar se uma dada amostra veio ou não de uma distribuição normal.
#
# Não precisamos e não entraremos nos detalhes da sua estatística de teste. Tudo que precisamos saber por ora é:
#
# * A hipótese nula, $H_{0}$, é a normalidade dos dados.
# * Se o valor-p for menor que o nível de significância $\alpha$, então temos evidências de que os dados não vêm de uma distribuição normal.
# * Se o valor-p for maior que $\alpha$, então não podemos afimar que os dados não vêm de uma distribuição normal (o que é sutilmente diferente de afirmar que eles _vêm_ de uma distribuição normal. Cuidado!).
# * Apesar de ter tendência a melhores resultados quanto maior a amostra, a maior parte das implementações não suporta computações com amostras muito grandes.
# * A implementação do SciPy por exemplo só suporta até 5000 observações.
# * É altamente aconselhado fazer o teste em conjunto com uma análise gráfica de um _q-q plot_.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="EG-ltCuQPc1W" outputId="32ec836d-404e-4b98-cc65-459add84f44d"
normal_data = sct.norm.rvs(loc=10, scale=4, size=100)
sct.shapiro(normal_data)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="8tvWUYX4Przx" outputId="83e01ed0-78b9-420b-8572-7d37db3026b3"
normal_data = sct.expon.rvs(scale=4, size=100)
sct.shapiro(normal_data)
# + [markdown] colab_type="text" id="adIEM13XuRlN"
# ### Jarque-Bera
#
# Assim como Shapiro-Wilk, o teste de Jarque-Bera é um teste de aderência à distribuição normal com $H_{0}$ sendo a normalidade dos dados. A diferença reside na estatística de teste utilizada.
#
# A estatística de teste é baseada na assimetria (_skewness_) e curtose (_kurtosis_) excessiva da amostra. Se a amostra vem de uma distribuição normal, então esses valores devem ser muito próximos de zero. Se isso acontecer, então a estatística de teste tem distribuição $\chi^{2}$ com dois graus de liberdade.
#
# No entanto, se a amostra for pequena, podemos ter muitos falsos negativos (Erro Tipo I) ao utilizarmos a distribuição $\chi^{2}$, ou seja, rejeitamos $H_{0}$ quando ela é verdadeira.
#
# Para evitar isso, as implementações costumam utilizar aproximações por Monte-Carlo quando $n$ é pequeno, reservando a aproximação $\chi^{2}$ para amostras grandes.
#
# Novamente, é altamente aconselhado complementar o resultado desse teste com um _q-q plot_.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="LcUvjC4APy5D" outputId="117aa7b6-c8c0-475d-ff72-ef93ea386b1d"
normal_data = sct.norm.rvs(loc=10, scale=4, size=100)
sct.jarque_bera(normal_data)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="xzU7eZBqP4Jj" outputId="da5fbf20-db39-419d-a8cc-118e666be6ca"
normal_data = sct.expon.rvs(scale=4, size=100)
sct.jarque_bera(normal_data)
# + [markdown] colab_type="text" id="cDpdgnjvwFD7"
# ## Referências
#
# * [A Gentle Introduction to Statistical Hypothesis Testing](https://machinelearningmastery.com/statistical-hypothesis-tests/)
#
# * [How to Correctly Interpret P Values](https://blog.minitab.com/blog/adventures-in-statistics-2/how-to-correctly-interpret-p-values)
#
# * [A Dirty Dozen: Twelve P-Value Misconceptions](http://www.perfendo.org/docs/BayesProbability/twelvePvaluemisconceptions.pdf)
#
# * [An investigation of the false discovery rate and the misinterpretation of p-values](https://royalsocietypublishing.org/doi/pdf/10.1098/rsos.140216)
#
# * [Statistical tests, P values, confidence intervals, and power: a guide to misinterpretations](https://link.springer.com/content/pdf/10.1007%2Fs10654-016-0149-3.pdf)
#
# * [Why Are P Values Misinterpreted So Frequently?](https://statisticsbyjim.com/hypothesis-testing/p-values-misinterpreted/)
#
# * [Statistical Significance Explained](https://towardsdatascience.com/statistical-significance-hypothesis-testing-the-normal-curve-and-p-values-93274fa32687)
#
# * [Definition of Power](https://newonlinecourses.science.psu.edu/stat414/node/304/)
#
# * [The Math Behind A/B Testing with Example Python Code](https://towardsdatascience.com/the-math-behind-a-b-testing-with-example-code-part-1-of-2-7be752e1d06f)
#
# * [Handy Functions for A/B Testing in Python](https://medium.com/@henryfeng/handy-functions-for-a-b-testing-in-python-f6fdff892a90)
| Modulo 5/pensamentoEstatistico.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
machine_name = 'EAST'
divertor_name='EAST_divertor.dat'
equili_name='73999_030400ms.mat'
from os import sys, path; sys.path.append(path.dirname(path.abspath(path.abspath(''))))
import ergospy.file, ergospy.IPyUI
import matplotlib.path as mplPath
path_machine = ergospy.file.PathMachine(machine_name)
folder_equilibrium = ergospy.file._path_ergos / 'equilibrium_preparation'
div_RZ = ergospy.file.DivertorRead(machine_name, divertor_name)
div_path = mplPath.Path(div_RZ)
# -
# ## FLTs Initiate from SOL with Equilibrium Field
# +
import numpy as np
import pandas as pd
FLT_df = pd.DataFrame(
columns =
["Len", "Min s",
"Init R", "Init Z", "Init Phi",
"Up End R", "Up End Z", "Up End Phi",
"Down End R", "Down End Z", "Down End Phi",
"Up End AOI", "Down End AOI"]) # AOI for Angle of Incidence
equili_dict = ergospy.file.EquiliDictRead(machine_name, equili_name)
S, TET = equili_dict['S'][:,0], equili_dict['TET'][:,0]
Q = equili_dict['Q'][:,0]
r_mesh,z_mesh = equili_dict['r_mesh'], equili_dict['z_mesh']
TET_halfnum, repeat_times = 15, 1
TET_indrange = range(-TET_halfnum, TET_halfnum+1)
start_points_rzphi = np.stack(
(np.repeat(r_mesh[-1,TET_indrange]+7e-2,repeat_times),
np.repeat(z_mesh[-1,TET_indrange], repeat_times),
np.random.uniform(0.0,2*np.pi,len(r_mesh[-1,TET_indrange])*repeat_times)), axis=-1)
start_points_rzphi+= np.random.rand(*start_points_rzphi.shape)*1e-5
flt_num = start_points_rzphi.shape[0]
FLT_df['Init R'] = start_points_rzphi[:,0]
FLT_df['Init Z'] = start_points_rzphi[:,1]
FLT_df['Init Phi']=start_points_rzphi[:,2]
print(f"Totally {flt_num} lines to trace.")
# -
from importlib import reload
reload(ergospy.measure.FLT)
# +
from multiprocessing import Pool
from ergospy.field.Field import Field2D
# Generate 2D RZ-mesh
folder_equilibrium = '../equilibrium_preparation/'
nR, nZ = np.loadtxt(folder_equilibrium+'nR_nZ.dat', dtype=np.uint32)
RZlimit = np.loadtxt(folder_equilibrium+'R_Z_min_max.dat', dtype=np.float32)
R = np.linspace(RZlimit[0],RZlimit[1], nR) # [Rmin, Rmin+dR, ..., Rmax] totally nR elements.
Z = np.linspace(RZlimit[2],RZlimit[3], nZ) # [Zmin, Zmin+dZ, ..., Zmax] totally nZ elements.
# Reading scalar field and calculate their components in cylindrical coordinate
BR = np.loadtxt(folder_equilibrium+'BR.dat', dtype=np.float32).T
BZ = np.loadtxt(folder_equilibrium+'BZ.dat', dtype=np.float32).T
BPhi=np.loadtxt(folder_equilibrium+'Bt.dat', dtype=np.float32).T
EQU_2D = Field2D(R,Z, BR, BZ, BPhi)
B = EQU_2D.field_mag()
print("Prepare linear B field interpolator ")
from scipy.interpolate import RegularGridInterpolator
BR_interp = RegularGridInterpolator((R, Z), BR/B)
BZ_interp = RegularGridInterpolator((R, Z), BZ/B)
BPhi_interp = RegularGridInterpolator((R,Z), BPhi/B)
print("Finished linear B field interpolator ")
def field_interp(y):
return np.asarray([ BR_interp(y[:-1])[0], BZ_interp(y[:-1])[0], BPhi_interp(y[:-1])[0]/y[0] ])
# +
from ergospy.field.field_line_tracing import FLT3D
from ergospy.measure.FLT import FLT_length, FLT_reach_min_s, FLT_endpoint
def FLT3D_params(*args):
strmline = FLT3D(*args)
up_end, down_end = FLT_endpoint(strmline)
return {'Len':FLT_length(strmline),
'Min s':FLT_reach_min_s(strmline, folder_equilibrium),
'Up End':up_end, 'Down End':down_end}
import timeit; start_time = timeit.default_timer()
# If you want to test the parallel speedup, just add your preferred number to the Pool arg `processes`
proc_pool, proc_list = Pool(), []
for j in range(flt_num):
proc_list.append(proc_pool.apply_async(
FLT3D_params,
args=(j, start_points_rzphi[j,:], div_path, div_RZ, field_interp, True, True, 20.0e1, 4e-1),
# kwds={'report_progress':True}
))
for i, proc in enumerate(proc_list):
FLT_param = proc.get()
FLT_df.loc[i, 'Len'] = FLT_param['Len']
FLT_df.loc[i, 'Min s'] = FLT_param['Min s']
FLT_df.loc[i, 'Up End R'], FLT_df.loc[i, 'Up End Z'], FLT_df.loc[i, 'Up End Phi'] = FLT_param['Up End']
FLT_df.loc[i, 'Down End R'], FLT_df.loc[i, 'Down End Z'], FLT_df.loc[i, 'Down End Phi'] = FLT_param['Down End']
# for k,v in proc.get().items():
# FLT_df.loc[i,k] = v
proc_pool.close(); proc_pool.join()
elapsed = timeit.default_timer() - start_time
print(elapsed)
print("All field line traces ODE done...")
# print(len_params_list)
# -
FLT_df
SP_rzphi = np.empty((len(FLT_df)*2, 3))
SP_rzphi[:len(FLT_df),:] = FLT_df[['Up End R','Up End Z','Up End Phi']].to_numpy()
SP_rzphi[len(FLT_df):,:] = FLT_df[['Down End R','Down End Z','Down End Phi']].to_numpy()
# for i in range(len(FLT_df)):
# SP_rzphi[i,:] = FLT_df.at[i, 'Up End']
# SP_rzphi[i+len(FLT_df),:] = FLT_df.at[i, 'Down End']
# +
import matplotlib.pyplot as plt
from matplotlib import colors
from matplotlib.ticker import PercentFormatter, MaxNLocator
fig, axs = plt.subplots(1, 2, tight_layout=True)
for i in range(2):
# ax = axs[i]
# N is the count in each bin, bins is the lower-limit of the bin
N, bins, patches = axs[i].hist(SP_rzphi[:,i], bins=40, range=[np.min(div_RZ[:,i]), np.max(div_RZ[:,i])])
# N, bins, patches = axs[i].hist(SP_rzphi[:len(FLT_df),i], bins=10,)
# N, bins, patches = axs[i].hist(SP_rzphi[len(FLT_df):,i], bins=10,)
axs[i].yaxis.set_major_locator(MaxNLocator(integer=True))
# We'll color code by height, but you could use any scalar
fracs = N / N.max()
# we need to normalize the data to 0..1 for the full range of the colormap
norm = colors.Normalize(fracs.min(), fracs.max())
# Now, we'll loop through our objects and set the color of each accordingly
for thisfrac, thispatch in zip(fracs, patches):
color = plt.cm.viridis(norm(thisfrac))
thispatch.set_facecolor(color)
# ax = axs[1]
# # We can also normalize our inputs by the total number of counts
# axs[1].hist(UP_SP_rzphi[:,1], bins=n_bins, density=True)
# # Now we format the y-axis to display percentage
# axs[1].yaxis.set_major_formatter(PercentFormatter(xmax=1))
# +
from ergospy.measure import divertor
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib notebook
fig, ax = plt.subplots(1, 1)
ax.plot(div_RZ[:,0], div_RZ[:,1])
point = np.array([1.4371,0.7983])
ax.scatter(FLT_df['Init R'],FLT_df['Init Z'], s=10, c='grey', label='Init')
ax.scatter(SP_rzphi[:len(FLT_df),0], SP_rzphi[:len(FLT_df),1], s=10, c='purple', label='SP UP')
ax.scatter(SP_rzphi[len(FLT_df):,0], SP_rzphi[len(FLT_df):,1], s=10, c='red', label='SP DOWN')
ax.set_aspect('equal')
ax.legend(loc="center left", fontsize='x-small')
plt.show()
# tile_ind, tile_mu = divertor.nearest_tile_ind_mu(point, div_edge)
# print(divertor.div_len_from_origin(tile_ind, tile_mu, div_edge))
# -
# ## Potential Reference:
# - [Plot an histogram with y-axis as percentage (using FuncFormatter?)](https://stackoverflow.com/questions/51473993/plot-an-histogram-with-y-axis-as-percentage-using-funcformatter)
# - [Add Matplotlib Percentage Ticks to a Histogram](https://malithjayaweera.com/2018/09/add-matplotlib-percentage-ticks-histogram/)
from importlib import reload
reload(ergospy.measure.FLT)
reload(ergospy.field.field_line_tracing)
del FLT_df
from importlib import reload
reload(measure_div)
# +
import ergospy.measure.divertor as measure_div
equili_SP_RZ = {
'UP': [1.72373, 1.08829],
'DOWN':[1.76893,-1.1537]
}
equili_SP_accu_len = {
'UP': measure_div.nearest_point_accu_len(equili_SP_RZ['UP'], div_RZ),
'DOWN': measure_div.nearest_point_accu_len(equili_SP_RZ['DOWN'], div_RZ),
}
# -
# ## FLTs Initiate from a Grid on Divertor near to the Original Strike Points with Perturbant Field
path_RMP = path_machine / 'HCFs' / 'field_computation' / 'c'
# +
Dist_N, Phi_N = 64, 64
DIR = 'UP' # 'DOWN'
Dist_shift_a = np.linspace(-6e-2, 14e-2, Dist_N) if DIR=='UP' else np.linspace(-6e-2, 2e-2, Dist_N)
Phi_a = np.linspace(0, 2*np.pi, Phi_N)
Dist_g, Phi_g = np.meshgrid(
Dist_shift_a + equili_SP_accu_len[DIR],
Phi_a, indexing='ij')
R_g, Z_g = np.empty_like(Dist_g), np.empty_like(Dist_g)
# Failed parallel code
# vec_accu_len_to_RZ = np.vectorize(measure_div.accu_len_to_RZ)
# vec_accu_len_to_RZ(Dist_g, div_RZ)
for i in range(Dist_N):
R_g[i,:] = measure_div.accu_len_to_RZ(Dist_g[i,0], div_RZ)[0,None]
Z_g[i,:] = measure_div.accu_len_to_RZ(Dist_g[i,0], div_RZ)[1,None]
start_points_rzphi = np.empty((Dist_N*Phi_N, 3))
start_points_rzphi[:,0] = R_g.flatten()
start_points_rzphi[:,1] = Z_g.flatten()
start_points_rzphi[:,2]=Phi_g.flatten()
while not np.all(div_path.contains_points(start_points_rzphi[:,0:2])):
points_maybe_outside = div_path.contains_points(start_points_rzphi[:,0:2])#.astype(int)
print("Let points be inside, how many outside now:", np.count_nonzero(~points_maybe_outside))
# start_points_rzphi[~points_maybe_outside] = start_points_rzphi[~points_maybe_outside] * (1-1e-7)
start_points_rzphi = start_points_rzphi * (1-5e-7)
flt_num = start_points_rzphi.shape[0]
print(f"Totally {flt_num} lines to trace.")
# +
import pandas as pd
FLT_df = pd.DataFrame(
columns =
["Len", "Min s",
"Init R", "Init Z", "Init Phi",
"Up End R", "Up End Z", "Up End Phi",
"Down End R", "Down End Z", "Down End Phi",
"Up End AOI", "Down End AOI"]) # AOI for Angle of Incidence
FLT_df['Init R'] = start_points_rzphi[:,0]
FLT_df['Init Z'] = start_points_rzphi[:,1]
FLT_df['Init Phi']=start_points_rzphi[:,2]
FLT_df['Dist'] = Dist_g.flatten()
# -
FLT_df
# +
from ergospy.field.Field import Field3D
RMP_field_raw = 5.0* ergospy.file.FieldRead(path_RMP, xcor='RZPhi', fcor='RZPhi')
RMP_param = ergospy.file.FieldMeshRead(path_RMP, xcor='RZPhi', fcor='RZPhi')
RMP_3D = Field3D(RMP_param['R'], RMP_param['Z'], RMP_param['Phi'], RMP_field_raw[...,0], RMP_field_raw[...,1], RMP_field_raw[...,2])
B_3D = RMP_3D + EQU_2D
R,Z,Phi = B_3D.R, B_3D.Z, B_3D.Phi
BR,BZ,BPhi = B_3D.BR, B_3D.BZ, B_3D.BPhi
B = B_3D.field_mag()
# Extend [2pi-dPhi, 2pi]
Phi = np.append(Phi, 2*np.pi+Phi[...,0:2], axis=-1)
BR = np.append(BR, BR[...,0:2], axis=-1)
BZ = np.append(BZ, BZ[...,0:2], axis=-1)
BPhi = np.append(BPhi, BPhi[...,0:2], axis=-1)
B = np.append(B, B[...,0:2], axis=-1)
print("Prepare linear B field interpolator ")
BR_interp = RegularGridInterpolator((R, Z, Phi), BR/B)
BZ_interp = RegularGridInterpolator((R, Z, Phi), BZ/B)
BPhi_interp = RegularGridInterpolator((R, Z, Phi),BPhi/B)
print("Finished linear B field interpolator ")
def field_interp(y):
ymod = y.copy(); ymod[-1] %= (2*np.pi)
return np.asarray([ BR_interp(ymod)[0], BZ_interp(ymod)[0], BPhi_interp(ymod)[0]/y[0] ])
# FLT_direciton, Co-field and Counter-field direction
FLT_direction = [True, False] # [False, True] if the magnetic field is counterclock when viewing the torus from the top.
# -
print(R.min(),R.max(),Z.min(),Z.max())
# +
import progressbar, sys
from multiprocessing import Pool
from multiprocessing.sharedctypes import Value
def FLT3D_params(*args, **kwargs):
strmline = FLT3D(*args, **kwargs)
up_end, down_end = FLT_endpoint(strmline)
return {'Len':FLT_length(strmline),
'Min s':FLT_reach_min_s(strmline, folder_equilibrium),
'Up End':up_end, 'Down End':down_end}
import timeit; start_time = timeit.default_timer()
# If you want to test the parallel speedup, just add your preferred number to the Pool arg `processes`
proc_pool, proc_list = Pool(), []
flt_todo = FLT_df[FLT_df['Len'].isnull()].index
for j in flt_todo:
proc_list.append(proc_pool.apply_async(
FLT3D_params,
args=(j, start_points_rzphi[j,:], div_path, div_RZ, field_interp, FLT_direction[0], FLT_direction[1], 20.0e1, 4e-1),
kwds={'report_progress':False}
))
with progressbar.ProgressBar(max_value=len(flt_todo)) as bar:
progress_num = Value('i', -1)
for i, proc in enumerate(proc_list):
try:
FLT_param = proc.get()
FLT_df.loc[i, 'Len'] = FLT_param['Len']
FLT_df.loc[i, 'Min s'] = FLT_param['Min s']
FLT_df.loc[i, 'Up End R'], FLT_df.loc[i, 'Up End Z'], FLT_df.loc[i, 'Up End Phi'] = FLT_param['Up End']
FLT_df.loc[i, 'Down End R'], FLT_df.loc[i, 'Down End Z'], FLT_df.loc[i, 'Down End Phi'] = FLT_param['Down End']
except Exception as err:
print(f"The {i} field line met error {err}.")
with progress_num.get_lock():
progress_num.value += 1
bar.update(progress_num.value)
proc_pool.close(); proc_pool.join()
elapsed = timeit.default_timer() - start_time; print(elapsed)
print("All field line traces ODE done...")
# print(len_params_list)
# -
SP_rzphi = np.empty((len(FLT_df)*2, 3))
SP_rzphi[:len(FLT_df),:] = FLT_df[['Up End R','Up End Z','Up End Phi']].to_numpy()
SP_rzphi[len(FLT_df):,:] = FLT_df[['Down End R','Down End Z','Down End Phi']].to_numpy()
# +
import matplotlib.pyplot as plt
from matplotlib import colors
from matplotlib.ticker import PercentFormatter, MaxNLocator
fig, axs = plt.subplots(1, 2, tight_layout=True)
for i, param in enumerate(['Len', 'Min s']):
# ax = axs[i]
# N is the count in each bin, bins is the lower-limit of the bin
x_range = [0.7, 1.00] if param=='Min s' else None
N, bins, patches = axs[i].hist(FLT_df[param], bins=40, range=x_range)
axs[i].yaxis.set_major_locator(MaxNLocator(integer=True))
# We'll color code by height, but you could use any scalar
fracs = N / N.max()
# we need to normalize the data to 0..1 for the full range of the colormap
norm = colors.Normalize(fracs.min(), fracs.max())
# Now, we'll loop through our objects and set the color of each accordingly
for thisfrac, thispatch in zip(fracs, patches):
color = plt.cm.viridis(norm(thisfrac))
thispatch.set_facecolor(color)
# ax = axs[1]
# # We can also normalize our inputs by the total number of counts
# axs[1].hist(UP_SP_rzphi[:,1], bins=n_bins, density=True)
# # Now we format the y-axis to display percentage
# axs[1].yaxis.set_major_formatter(PercentFormatter(xmax=1))
# +
from ergospy.measure import divertor
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib notebook
fig, ax = plt.subplots(1, 1)
ax.plot(div_RZ[:,0], div_RZ[:,1])
point = np.array([1.4371,0.7983])
ax.scatter(FLT_df['Init R'],FLT_df['Init Z'], s=80, c='grey', label='Init')
ax.scatter(SP_rzphi[:len(FLT_df),0], SP_rzphi[:len(FLT_df),1], s=10, c='purple', label='SP UP')
ax.scatter(SP_rzphi[len(FLT_df):,0], SP_rzphi[len(FLT_df):,1], s=10, c='red', label='SP DOWN')
ax.set_aspect('equal')
ax.legend(loc="center left", fontsize='x-small')
plt.show()
# tile_ind, tile_mu = divertor.nearest_tile_ind_mu(point, div_edge)
# print(divertor.div_len_from_origin(tile_ind, tile_mu, div_edge))
# +
import matplotlib.pyplot as plt
fig, ax = plt.subplots(1,1,figsize=(10,4.5))
ax.set_xlabel('Toroidal Angle $\phi$', fontsize=10)
ax.set_ylabel('Length along the Divertor (m)', fontsize=10)
ax.set_title('Colour = length of field lines (m)', fontsize=15)
pc = ax.pcolormesh(
FLT_df['Init Phi'].to_numpy(dtype='float').reshape(Dist_N, Phi_N),
FLT_df['Dist'].to_numpy(dtype='float').reshape(Dist_N, Phi_N),
FLT_df['Len'].to_numpy(dtype='float').reshape(Dist_N, Phi_N),
cmap='plasma', vmin=0.)
fig.colorbar(pc, ax=ax)
plt.show()
# -
print(FLT_df['Len'].to_numpy().reshape(Dist_N, Phi_N).shape)
# +
import matplotlib.pyplot as plt
import matplotlib.colors as mcolors
fig, ax = plt.subplots(1,1, figsize=(10,4.5))
ax.set_xlabel('Toroidal Angle $\phi$', fontsize=10)
ax.set_ylabel('Length along the Divertor (m)', fontsize=10)
ax.set_title('Colour = $\min(s)$ that the field line can reach', fontsize=15)
pc = ax.pcolormesh(
FLT_df['Init Phi'].to_numpy(dtype='float').reshape(Dist_N, Phi_N),
FLT_df['Dist'].to_numpy(dtype='float').reshape(Dist_N, Phi_N),
FLT_df['Min s'].to_numpy(dtype='float').reshape(Dist_N, Phi_N),
cmap='RdGy', norm=mcolors.TwoSlopeNorm(vcenter=1., ))
fig.colorbar(pc, ax=ax)
plt.show()
# -
| nb/poincare_footprint/footprint_plot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] papermill={"duration": 0.016234, "end_time": "2021-05-21T00:48:50.202473", "exception": false, "start_time": "2021-05-21T00:48:50.186239", "status": "completed"} tags=[]
# ## Dependencies
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _kg_hide-input=true _kg_hide-output=true _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" papermill={"duration": 9.226181, "end_time": "2021-05-21T00:48:59.443776", "exception": false, "start_time": "2021-05-21T00:48:50.217595", "status": "completed"} tags=[]
import warnings, math, json
import pandas as pd
from sklearn.model_selection import KFold
from sklearn.metrics import mean_squared_error
import tensorflow_addons as tfa
import tensorflow.keras.layers as L
import tensorflow.keras.backend as K
from tensorflow.keras import optimizers, losses, metrics, Model
from tensorflow.keras.callbacks import EarlyStopping, ModelCheckpoint, LearningRateScheduler
from transformers import TFAutoModelForSequenceClassification, TFAutoModel, AutoTokenizer
from commonlit_scripts import *
seed = 0
seed_everything(seed)
sns.set(style='whitegrid')
warnings.filterwarnings('ignore')
pd.set_option('display.max_colwidth', 150)
# + [markdown] papermill={"duration": 0.014885, "end_time": "2021-05-21T00:48:59.473916", "exception": false, "start_time": "2021-05-21T00:48:59.459031", "status": "completed"} tags=[]
# ### Hardware configuration
# + _kg_hide-input=true _kg_hide-output=false papermill={"duration": 6.144878, "end_time": "2021-05-21T00:49:05.633763", "exception": false, "start_time": "2021-05-21T00:48:59.488885", "status": "completed"} tags=[]
strategy, tpu = get_strategy()
AUTO = tf.data.AUTOTUNE
REPLICAS = strategy.num_replicas_in_sync
print(f'REPLICAS: {REPLICAS}')
# + [markdown] papermill={"duration": 0.015228, "end_time": "2021-05-21T00:49:05.665183", "exception": false, "start_time": "2021-05-21T00:49:05.649955", "status": "completed"} tags=[]
# # Load data
# + _kg_hide-input=true papermill={"duration": 0.183488, "end_time": "2021-05-21T00:49:05.864423", "exception": false, "start_time": "2021-05-21T00:49:05.680935", "status": "completed"} tags=[]
train_filepath = '/kaggle/input/commonlitreadabilityprize/train.csv'
train = pd.read_csv(train_filepath)
print(f'Train samples: {len(train)}')
# removing unused columns
train.drop(['url_legal', 'license'], axis=1, inplace=True)
display(train.head())
# + [markdown] papermill={"duration": 0.016516, "end_time": "2021-05-21T00:49:05.897695", "exception": false, "start_time": "2021-05-21T00:49:05.881179", "status": "completed"} tags=[]
# # Model parameters
# + papermill={"duration": 0.027813, "end_time": "2021-05-21T00:49:05.942276", "exception": false, "start_time": "2021-05-21T00:49:05.914463", "status": "completed"} tags=[]
config = {
"BATCH_SIZE": 8 * REPLICAS,
"LEARNING_RATE": 1e-5 * REPLICAS,
"EPOCHS": 100,
"ES_PATIENCE": 30,
"N_FOLDS": 5,
"N_USED_FOLDS": 1,
"SEQ_LEN": 256,
"BASE_MODEL": '/kaggle/input/huggingface-roberta/roberta-base/',
"SEED": seed,
}
with open('config.json', 'w') as json_file:
json.dump(json.loads(json.dumps(config)), json_file)
# + [markdown] papermill={"duration": 0.017495, "end_time": "2021-05-21T00:49:05.977385", "exception": false, "start_time": "2021-05-21T00:49:05.959890", "status": "completed"} tags=[]
# ## Auxiliary functions
# + _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _kg_hide-input=true _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a" papermill={"duration": 0.037718, "end_time": "2021-05-21T00:49:06.031602", "exception": false, "start_time": "2021-05-21T00:49:05.993884", "status": "completed"} tags=[]
# Datasets utility functions
def custom_standardization(text, is_lower=True):
if is_lower:
text = text.lower() # if encoder is uncased
text = text.strip()
return text
def sample_target(features, target):
mean, _, stddev = target
sampled_target = tf.random.normal([], mean=tf.cast(mean, dtype=tf.float32),
stddev=tf.cast(stddev, dtype=tf.float32), dtype=tf.float32)
return (features, (mean, sampled_target))
def aux_target(features, label):
aux_label = [1, 0, 0, 0, 0]
if label[0] >= 1.: # target
aux_label = [0, 0, 0, 0, 1]
elif label[0] > -1.: # target
aux_label = [0, 0, 0, 1, 0]
elif label[0] > -2.: # target
aux_label = [0, 0, 1, 0, 0]
elif label[0] > -3.: # target
aux_label = [0, 1, 0, 0, 0]
return (features, (label[0], label[1], aux_label))
def get_dataset(pandas_df, tokenizer, labeled=True, ordered=False, repeated=False,
is_sampled=False, batch_size=32, seq_len=128, is_lower=True):
"""
Return a Tensorflow dataset ready for training or inference.
"""
text = [custom_standardization(text, is_lower) for text in pandas_df['excerpt']]
# Tokenize inputs
tokenized_inputs = tokenizer(text, max_length=seq_len, truncation=True,
padding='max_length', return_tensors='tf')
if labeled:
dataset = tf.data.Dataset.from_tensor_slices(({'input_ids': tokenized_inputs['input_ids'],
'attention_mask': tokenized_inputs['attention_mask']},
(pandas_df['target'], pandas_df['target'], pandas_df['standard_error'])))
if is_sampled:
dataset = dataset.map(sample_target, num_parallel_calls=tf.data.AUTOTUNE)
dataset = dataset.map(aux_target, num_parallel_calls=tf.data.AUTOTUNE)
else:
dataset = tf.data.Dataset.from_tensor_slices({'input_ids': tokenized_inputs['input_ids'],
'attention_mask': tokenized_inputs['attention_mask']})
if repeated:
dataset = dataset.repeat()
if not ordered:
dataset = dataset.shuffle(2048)
dataset = dataset.batch(batch_size)
dataset = dataset.cache()
dataset = dataset.prefetch(tf.data.AUTOTUNE)
return dataset
# + [markdown] papermill={"duration": 0.017061, "end_time": "2021-05-21T00:49:06.065434", "exception": false, "start_time": "2021-05-21T00:49:06.048373", "status": "completed"} tags=[]
# # Model
# + _kg_hide-output=true papermill={"duration": 31.523035, "end_time": "2021-05-21T00:49:37.605185", "exception": false, "start_time": "2021-05-21T00:49:06.082150", "status": "completed"} tags=[]
def model_fn(encoder, seq_len=256):
input_ids = L.Input(shape=(seq_len,), dtype=tf.int32, name='input_ids')
input_attention_mask = L.Input(shape=(seq_len,), dtype=tf.int32, name='attention_mask')
outputs = encoder({'input_ids': input_ids,
'attention_mask': input_attention_mask})
last_hidden_state = outputs['last_hidden_state']
cls_token = last_hidden_state[:, 0, :]
output = L.Dense(1, name='output')(cls_token)
output_sample = L.Dense(1, name='output_sample')(cls_token)
output_aux = L.Dense(5, activation='softmax', name='output_aux')(cls_token)
model = Model(inputs=[input_ids, input_attention_mask],
outputs=[output, output_sample, output_aux])
optimizer = optimizers.Adam(lr=config['LEARNING_RATE'])
model.compile(optimizer=optimizer,
loss={'output': losses.MeanSquaredError(),
'output_sample': losses.MeanSquaredError(),
'output_aux': losses.CategoricalCrossentropy(label_smoothing=0.1)},
loss_weights={'output': 1.,
'output_sample': 1.,
'output_aux': 1.},
metrics={'output': metrics.RootMeanSquaredError(name='rmse'),
'output_sample': metrics.RootMeanSquaredError(name='rmse'),
'output_aux': metrics.CategoricalAccuracy(name='acc')})
return model
with strategy.scope():
encoder = TFAutoModel.from_pretrained(config['BASE_MODEL'])
# Freeze embeddings
encoder.layers[0].embeddings.trainable = False
model = model_fn(encoder, config['SEQ_LEN'])
model.summary()
# + [markdown] papermill={"duration": 0.018105, "end_time": "2021-05-21T00:49:37.641547", "exception": false, "start_time": "2021-05-21T00:49:37.623442", "status": "completed"} tags=[]
# # Training
# + _kg_hide-input=true _kg_hide-output=true papermill={"duration": 298.951773, "end_time": "2021-05-21T00:54:36.612994", "exception": false, "start_time": "2021-05-21T00:49:37.661221", "status": "completed"} tags=[]
tokenizer = AutoTokenizer.from_pretrained(config['BASE_MODEL'])
skf = KFold(n_splits=config['N_FOLDS'], shuffle=True, random_state=seed)
oof_pred = []; oof_labels = []; history_list = []; oof_pred2 = []
for fold,(idxT, idxV) in enumerate(skf.split(train)):
if fold >= config['N_USED_FOLDS']:
break
if tpu: tf.tpu.experimental.initialize_tpu_system(tpu)
print(f'\nFOLD: {fold+1}')
print(f'TRAIN: {len(idxT)} VALID: {len(idxV)}')
# Model
K.clear_session()
with strategy.scope():
encoder = TFAutoModel.from_pretrained(config['BASE_MODEL'])
# Freeze embeddings
encoder.layers[0].embeddings.trainable = False
model = model_fn(encoder, seq_len=config['SEQ_LEN'])
model_path = f'model_{fold}.h5'
es = EarlyStopping(monitor='val_output_rmse', mode='min',
patience=config['ES_PATIENCE'], restore_best_weights=True, verbose=1)
checkpoint = ModelCheckpoint(model_path, monitor='val_output_rmse', mode='min',
save_best_only=True, save_weights_only=True)
# Train
history = model.fit(x=get_dataset(train.loc[idxT], tokenizer, repeated=True, is_sampled=True,
batch_size=config['BATCH_SIZE'], seq_len=config['SEQ_LEN']),
validation_data=get_dataset(train.loc[idxV], tokenizer, ordered=True,
batch_size=config['BATCH_SIZE'], seq_len=config['SEQ_LEN']),
steps_per_epoch=10,
callbacks=[es, checkpoint],
epochs=config['EPOCHS'],
verbose=2).history
history_list.append(history)
# Save last model weights
model.load_weights(model_path)
# Results
print(f"#### FOLD {fold+1} OOF RMSE = {np.min(history['val_output_rmse']):.4f}")
print(f"#### FOLD {fold+1} OOF RMSE = {np.min(history['val_output_sample_rmse']):.4f} (sampled)")
print(f"#### FOLD {fold+1} OOF ACC = {np.max(history['val_output_aux_acc']):.4f}")
# OOF predictions
valid_ds = get_dataset(train.loc[idxV], tokenizer, ordered=True,
batch_size=config['BATCH_SIZE'], seq_len=config['SEQ_LEN'])
oof_labels.append([target[0].numpy() for sample, target in iter(valid_ds.unbatch())])
x_oof = valid_ds.map(lambda sample, target: sample)
preds = model.predict(x_oof)
oof_pred.append(preds[0])
oof_pred2.append(preds[1])
# + [markdown] papermill={"duration": 0.053207, "end_time": "2021-05-21T00:54:36.720975", "exception": false, "start_time": "2021-05-21T00:54:36.667768", "status": "completed"} tags=[]
# ## Model loss and metrics graph
# + _kg_hide-input=true papermill={"duration": 2.194935, "end_time": "2021-05-21T00:54:38.969810", "exception": false, "start_time": "2021-05-21T00:54:36.774875", "status": "completed"} tags=[]
for fold, history in enumerate(history_list):
print(f'\nFOLD: {fold+1}')
plot_metrics(history)
# + [markdown] papermill={"duration": 0.066192, "end_time": "2021-05-21T00:54:39.101454", "exception": false, "start_time": "2021-05-21T00:54:39.035262", "status": "completed"} tags=[]
# # Model evaluation
#
# We are evaluating the model on the `OOF` predictions, it stands for `Out Of Fold`, since we are training using `K-Fold` our model will see all the data, and the correct way to evaluate each fold is by looking at the predictions that are not from that fold.
#
# ## OOF metrics
# + _kg_hide-input=true papermill={"duration": 0.079506, "end_time": "2021-05-21T00:54:39.246730", "exception": false, "start_time": "2021-05-21T00:54:39.167224", "status": "completed"} tags=[]
y_true = np.concatenate(oof_labels)
y_preds = np.concatenate(oof_pred)
y_preds2 = np.concatenate(oof_pred2)
y_preds_merge = np.mean([y_preds, y_preds2], axis=0)
for fold, history in enumerate(history_list):
print(f"#### FOLD {fold+1} OOF RMSE = {np.min(history['val_output_rmse']):.4f}")
print(f"#### FOLD {fold+1} OOF RMSE = {np.min(history['val_output_sample_rmse']):.4f} (sampled)")
print(f"#### FOLD {fold+1} OOF ACC = {np.max(history['val_output_aux_acc']):.4f}")
print(f'OOF RMSE: {mean_squared_error(y_true, y_preds, squared=False):.4f}')
print(f'OOF RMSE: {mean_squared_error(y_true, y_preds2, squared=False):.4f} (sampled)')
print(f'OOF RMSE: {mean_squared_error(y_true, y_preds_merge, squared=False):.4f} (merged)')
# + [markdown] papermill={"duration": 0.066348, "end_time": "2021-05-21T00:54:39.378070", "exception": false, "start_time": "2021-05-21T00:54:39.311722", "status": "completed"} tags=[]
# ### **Error analysis**, label x prediction distribution
#
# Here we can compare the distribution from the labels and the predicted values, in a perfect scenario they should align.
# + _kg_hide-input=true papermill={"duration": 0.56856, "end_time": "2021-05-21T00:54:40.011800", "exception": false, "start_time": "2021-05-21T00:54:39.443240", "status": "completed"} tags=[]
preds_df = pd.DataFrame({'Label': y_true, 'Pred': y_preds[:,0], 'Pred2': y_preds2[:,0]})
fig, ax = plt.subplots(1, 1, figsize=(20, 6))
sns.distplot(preds_df['Label'], ax=ax, label='Label')
sns.distplot(preds_df['Pred'], ax=ax, label='Prediction')
sns.distplot(preds_df['Pred2'], ax=ax, label='Prediction_samp')
ax.legend()
plt.show()
# + _kg_hide-input=true papermill={"duration": 1.574683, "end_time": "2021-05-21T00:54:41.655445", "exception": false, "start_time": "2021-05-21T00:54:40.080762", "status": "completed"} tags=[]
sns.jointplot(data=preds_df, x='Label', y='Pred', kind='reg', height=8)
plt.show()
sns.jointplot(data=preds_df, x='Label', y='Pred2', kind='reg', height=8)
plt.show()
| Model backlog/Train/36-commonlit-roberta-base-aux-5-cats-smoothing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from os import path
import matplotlib.pyplot as plt
import numpy as np
from enmspring.graphs_bigtraj import BackboneMeanModeAgent
from enmspring.eigenvector_backbone import NresidEigenvaluePlot
rootfolder = '/home/ytcdata/bigtraj_fluctmatch/500ns'
drawzone_folder = '/home/yizaochen/Desktop/drawzone_temp'
# ### Part 1: Initialize
# +
host = 'a_tract_21mer'
interval_time = 500
s_agent = BackboneMeanModeAgent(host, rootfolder, interval_time)
s_agent.load_mean_mode_laplacian_from_npy()
s_agent.eigen_decompose()
s_agent.process_first_small_agent()
s_agent.set_benchmark_array()
s_agent.set_strand_array()
s_agent.initialize_nodes_information()
s_agent.split_node_list_into_two_strand()
# -
# ### Part 2: Observe Eigenvector
# +
strand_id = 'STRAND2' # STRAND1, STRAND2
mode_id = 1 # Start from 1
show_xticklabel = False
figsize = (8,4)
s_agent.plot_sele_eigenvector(figsize, strand_id, mode_id, show_xticklabel)
plt.tight_layout()
plt.show()
# -
# ### Part 3: Number of Resid and Eigenvalue Plot
strand_id = 'STRAND2' # STRAND1, STRAND2
plot_agent = NresidEigenvaluePlot(host, strand_id, s_agent)
criteria = 0.01
ylim1 = (0, 65)
ylim2 = (1, 21)
assist_hlines = range(1, 21, 2)
figsize = (8, 4)
fig, ax1, ax2 = plot_agent.plot(figsize, criteria, ylim1, ylim2, assist_hlines)
plt.tight_layout()
plt.show()
# ### Part 4: Batch Generate
hosts = ['a_tract_21mer', 'atat_21mer', 'g_tract_21mer', 'gcgc_21mer']
strandids = ['STRAND1', 'STRAND2']
for host in hosts:
s_agent = BackboneMeanModeAgent(host, rootfolder, interval_time)
s_agent.load_mean_mode_laplacian_from_npy()
s_agent.eigen_decompose()
s_agent.process_first_small_agent()
s_agent.set_benchmark_array()
s_agent.set_strand_array()
s_agent.initialize_nodes_information()
s_agent.split_node_list_into_two_strand()
for strand_id in strandids:
plot_agent = NresidEigenvaluePlot(host, strand_id, s_agent)
criteria = 0.01
ylim1 = (0, 850)
ylim2 = (1, 13)
assist_hlines = range(1, 14)
figsize = (8, 4)
fig, ax1, ax2 = plot_agent.plot(figsize, criteria, ylim1, ylim2, assist_hlines)
plt.tight_layout()
plt.savefig(path.join(drawzone_folder, f'{host}.{strand_id}.png'), dpi=100)
plt.show()
| notebooks/prominent_modes_backbone_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Clustering con Python
# ### Importar el dataset
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.cluster import AgglomerativeClustering
from sklearn.cluster import KMeans
from scipy.cluster.hierarchy import dendrogram, linkage
df = pd.read_csv("../datasets/wine/winequality-red.csv",sep=";")
df.shape
df.head()
plt.hist(df["quality"])
df.groupby("quality").mean()
# ### Normaliación de los datos
df_norm = (df-df.min())/(df.max()-df.min())
df_norm.head()
# ## Clustering jerárquico con scikit-learn
def dendrogram_tune(*args,**kwargs):
max_d = kwargs.pop("max_d", None)
if (max_d and 'color_threshold' not in kwargs):
kwargs['color_threshold'] = max_d
annotate_above = kwargs.pop('annotate_above',0)
ddata = dendrogram(*args,**kwargs)
if (not kwargs.get('no_plot', False)):
plt.title("Clustering jerárquico con Dendrograma truncado")
plt.xlabel("Índice del dataset (o tamaño del cluster)")
plt.ylabel("Distancia")
for index, distance, color in zip(ddata['icoord'], ddata['dcoord'], ddata['color_list']):
x = 0.5 * sum(index[1:3])
y = distance[1]
if (y > annotate_above):
plt.plot(x,y,'o',c=color)
plt.annotate("%.3g"%y,(x,y),xytext=(0,-5),
textcoords = "offset points", va="top", ha="center")
if (max_d):
plt.axhline(y=max_d, c='k')
return ddata
Z = linkage(df_norm,"ward")
plt.figure(figsize=(25,10))
plt.title("Dendrograma de los vinos")
plt.xlabel("ID del vino")
plt.ylabel("Distancia")
dendrogram_tune(Z,truncate_mode="lastp",p=12,show_contracted=True,annotate_above=10,max_d=4.5)
plt.show()
# +
last = Z[-10:,2]
last_rev = last[::-1]
print(last_rev)
idx = np.arange(1,len(last)+1)
plt.plot(idx,last_rev)
acc = np.diff(last,2)
acc_rev = acc[::-1]
plt.plot(idx[:-2]+1,acc_rev)
plt.show()
k = acc_rev.argmax()+2
print(f"El número óptimo de clusters es {k}")
# -
clus = AgglomerativeClustering(n_clusters=6,linkage="ward").fit(df_norm)
md_h = pd.Series(clus.labels_)
md_h
plt.hist(md_h)
plt.title("Histograma de los clusters")
plt.xlabel("Cluster")
plt.ylabel("Número de vinos del cluster")
clus.children_
# ## K-means
model = KMeans(n_clusters=6)
model.fit(df_norm)
model.labels_
md_k = pd.Series(model.labels_)
df_norm["clust_h"] = md_h
df_norm["clust_k"] = md_k
df_norm.head()
plt.hist(md_k)
model.cluster_centers_
model.inertia_
# ## Interpretación final
df_norm.groupby("clust_k").mean()
| code/T6 - 4 - Clustering completo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Linear Regression
# +
from statistics import mean
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import style
import random
style.use('fivethirtyeight')
# +
# xs = np.array([1,2,3,4,5,6], dtype=np.float64)
# ys = np.array([5,4,6,5,6,7], dtype=np.float64)
# plt.scatter(xs,ys)
# plt.show()
# -
def create_dataset(hm, varience, step=2, correlation=False):
val = 1
ys = []
for i in range(hm):
y = val + random.randrange(-varience, varience)
ys.append(y)
if correlation and correlation == 'pos':
val += step
elif correlation and correlation == 'neg':
val -= step
xs = [i for i in range(len(ys))]
return np.array(xs, dtype=np.float64), np.array(ys, dtype=np.float64)
# We can test our assumption using varience. If varience decreases, r-square increases
xs, ys = create_dataset(40, 40, 2, correlation='pos')
plt.scatter(xs,ys)
plt.show()
def best_fit_slope_and_intercept(xs, ys):
m = ((mean(xs) * mean(ys)) - mean(xs*ys)) / ((mean(xs)**2) - mean(xs**2))
b = mean(ys) - m * mean(xs)
return m, b
m, b = best_fit_slope_and_intercept(xs, ys)
print(m)
print(b)
regression_line = [(m*x)+b for x in xs]
print(regression_line)
plt.scatter(xs,ys)
plt.plot(regression_line)
plt.show()
predict_x = 8
predict_y = (m*predict_x) + b
plt.scatter(xs,ys)
plt.plot(regression_line)
plt.scatter(predict_x,predict_y, color='red', s=100)
plt.show()
# This line is good fit line but not the best fit line.
def squared_error(ys_orig,ys_line):
return sum((ys_line-ys_orig)**2)
def coeff_of_determination(ys_orig, ys_line):
# y_mean_line = [mean(y) for y in ys_orig]
# y_mean_line = mean(ys_orig)
y_mean_line = [mean(ys_orig)] * len(ys_orig)
squared_error_regr = squared_error(ys_orig,ys_line)
squared_error_regr_y_mean = squared_error(ys_orig,y_mean_line)
return 1 - (squared_error_regr/squared_error_regr_y_mean)
r_squared = coeff_of_determination(ys, regression_line)
print(r_squared)
| Sentdex/12 - Testing Assumptions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Building a Model To Predict Diabetes
# The given dataset lists the glucose level reading of several pregnant women taken either during a survey examination or routine medical care. It specifies if the 2-hour post-load plasma glucose was at least 200 mg/dl.
# ### I'll perform the following tasks here:
# 1. I'll find the features of the dataset
# 2. Find the response label of the dataset
# 3. I'll create a model to predict the diabetes outcome
# 4. I'll use training and testing datasets to train the model
# 5. Finally, I'll check the accuracy of the model.
# #### Importing the dataset
#Importing the required libraries
import pandas as pd
#Importing the diabetes dataset
df=pd.read_csv('pima-indians-diabetes.data',header=None)
# #### Analyzing the dataset
#Viewing the first five observations of the dataset
df.head()
# #### Finding the features of the dataset
#Using the .NAMES file to view and set the features of the dataset
df_names=['Number of times pregnant','Plasma glucose concentration','Diastolic blood pressure','Triceps skin fold thickness','2-Hour serum insulin','Body mass index','Diabetes pedigree function','Age','Class variable']
#Using the feature names set earlier and fix it as the column headers of the dataset
df=pd.read_csv('pima-indians-diabetes.data',header=None,names=df_names)
#Verifing if the dataset is updated with the new headers
df.head()
#Viewing the number of observations and features of the dataset
df.shape
# #### Finding the response of the dataset
#Selecting features from the dataset to create the model
features=['Number of times pregnant','2-Hour serum insulin','Body mass index','Age']
#Creating the feature object
x_features=df[features]
#Creating the reponse object
y_target=df['Class variable']
#Viewing the shape of the feature object
x_features.shape
#Viewing the shape of the target object
y_target.shape
# #### Using training and testing datasets to train the model
#Splitting the dataset to test and train the model
from sklearn.model_selection import train_test_split
x_train, x_test, y_train, y_test=train_test_split(x_features, y_target, random_state=1)
# #### Creating a model to predict the diabetes outcome
# Creating a logistic regression model using the training set
from sklearn.linear_model import LogisticRegression
logreg=LogisticRegression()
logreg.fit(x_train, y_train)
#Making predictions using the testing set
y_predict=logreg.predict(x_test)
# #### Checking the accuracy of the model
#Evaluating the accuracy of your model
from sklearn import metrics
print(metrics.accuracy_score(y_test, y_predict))
#Printing the first 30 actual and predicted responses
print('Actual: ', y_test.values[0:30])
print('Predicted: ', y_predict[0:30])
# The task is done. Thank you for checking out my notebook. Regards,
# * <NAME> :)
| Model to predict diabetes using logistic regression.ipynb |
# ---
# jupyter:
# jupytext:
# formats: ipynb,py:light
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ------
# <div>
# <center><h5>Higher Order Tutorial on Deep Learning</h5></center>
# <center><strong><h2>Graph Convolution Networks</h2></strong></center>
# <center><strong><h3>1.0.1 - Node Classification</h3></strong></center>
# <div>
# ------
# ### Keras DGL - Node Classification:
# ## `tl;dr: GraphCNN(output_dim, num_filters, graph_conv_filters)`
#
# Importing:
# ```python
# from keras_dgl.layers import GraphCNN
# ```
#
# Just like any keras model:
# ```python
# model = Sequential()
# model.add(GraphCNN(16, 2, graph_conv_filters, input_shape=(X.shape[1],)))
# model.add(GraphCNN(Y.shape[1], 2, graph_conv_filters))
# model.add(Activation('softmax'))
# ```
# ------
# + [markdown] slideshow={"slide_type": "slide"}
# # Graph Node Classification
# + [markdown] slideshow={"slide_type": "slide"}
# ### Motivation :
#
# There is a lot of data out there that can be represented in the form of a graph
# in real-world applications like in Citation Networks, Social Networks (Followers
# graph, Friends network, … ), Biological Networks or Telecommunications. <br>
# Using Graph extracted features can boost the performance of predictive models by
# relying of information flow between close nodes. However, representing graph
# data is not straightforward especially if we don’t intend to implement
# hand-crafted features.<br> In this post we will explore some ways to deal with
# generic graphs to do node classification based on graph representations learned
# directly from data.
#
# ### Dataset :
#
# The [Cora](https://linqs-data.soe.ucsc.edu/public/lbc/cora.tgz) citation network
# data set will serve as the base to the implementations and experiments
# throughout this post. Each node represents a scientific paper and edges between
# nodes represent a citation relation between the two papers.<br> Each node is
# represented by a set of binary features ( Bag of words ) as well as by a set of
# edges that link it to other nodes.<br> The dataset has **2708** nodes classified
# into one of seven classes. The network has **5429** links. Each Node is also
# represented by a binary word features indicating the presence of the
# corresponding word. Overall there is **1433** binary (Sparse) features for each
# node. In what follows we *only* use **140** samples for training and the rest
# for validation/test.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Problem Setting :
#
# 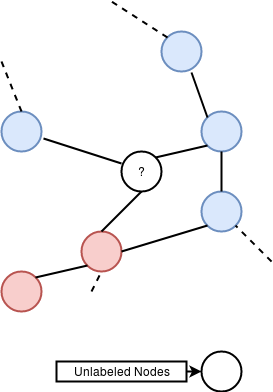
#
# **Problem** : Assigning a class label to nodes in a graph while having few
# training samples.<br> **Intuition**/**Hypothesis** : Nodes that are close in the
# graph are more likely to have similar labels.<br> **Solution** : Find a way to
# extract features from the graph to help classify new nodes.
#
# ### Proposed Approach :
#
# <br>
#
# **Baseline Model :**
#
# 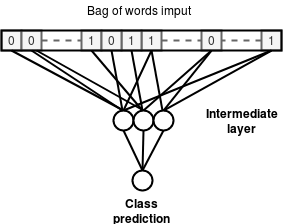
#
# We first experiment with the simplest model that learn to predict node classes
# using only the binary features and discarding all graph information.<br> This
# model is a fully-connected Neural Network that takes as input the binary
# features and outputs the class probabilities for each node.
#
# #### **Baseline model Accuracy : 53.28%**
# Source: https://github.com/CVxTz/graph_classification
# + [markdown] slideshow={"slide_type": "slide"}
# **Adding Graph features :**
#
# One way to automatically learn graph features by embedding each node into a
# vector by training a network on the auxiliary task of predicting the inverse of
# the shortest path length between two input nodes like detailed on the figure and
# code snippet below :
#
# 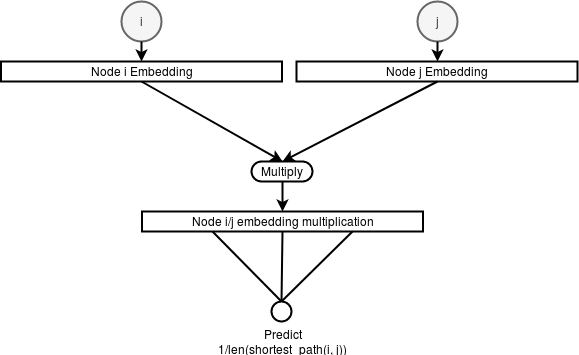
# + [markdown] slideshow={"slide_type": "slide"}
# The next step is to use the pre-trained node embedding as input to the
# classification model. We also add the an additional input which is the average
# binary features of the neighboring nodes using distance of learned embedding
# vectors.
#
# The resulting classification network is described in the following figure :
#
# 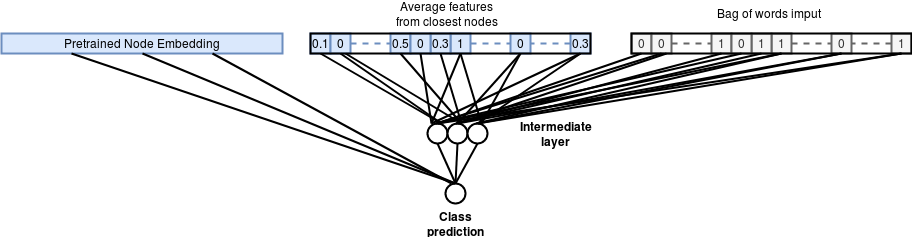
#
# <span class="figcaption_hack">Using pretrained embeddings to do node classification</span>
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# **Improving Graph feature learning :**
#
# We can look to further improve the previous model by pushing the pre-training
# further and using the binary features in the node embedding network and reusing
# the pre-trained weights from the binary features in addition to the node
# embedding vector. This results in a model that relies on more useful
# representations of the binary features learned from the graph structure.
#
# 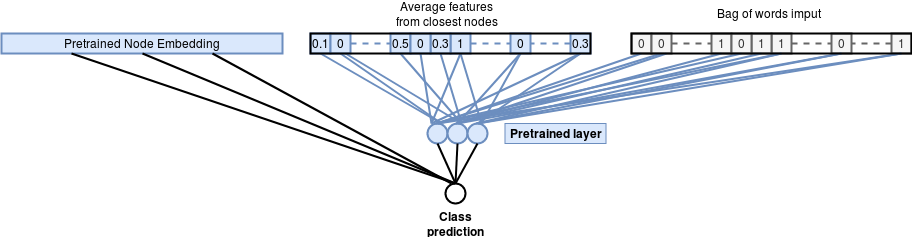
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Graph Neural Networks
# + [markdown] colab={"base_uri": "https://localhost:8080/", "height": 1295} colab_type="code" id="3lOBizVa4rVt" outputId="a3142dd2-4ff0-4bb6-a833-a7046f4e0596"
#
# Mathematically, the GCN model follows this formula:
#
# $H^{(l+1)} = \sigma(\tilde{D}^{-\frac{1}{2}}\tilde{A}\tilde{D}^{-\frac{1}{2}}H^{(l)}W^{(l)})$
#
# Here, $H^{(l)}$ denotes the $l^{th}$ layer in the network,
# $\sigma$ is the non-linearity, and $W$ is the weight matrix for
# this layer. $D$ and $A$, as commonly seen, represent degree
# matrix and adjacency matrix, respectively. The ~ is a renormalization trick
# in which we add a self-connection to each node of the graph, and build the
# corresponding degree and adjacency matrix. The shape of the input
# $H^{(0)}$ is $N \times D$, where $N$ is the number of nodes
# and $D$ is the number of input features. We can chain up multiple
# layers as such to produce a node-level representation output with shape
# $N \times F$, where $F$ is the dimension of the output node
# feature vector.
#
# The equation can be efficiently implemented using sparse matrix
# multiplication kernels (such as Kipf's
# `https://github.com/tkipf/pygcn`). The above DGL implementation
# in fact has already used this trick due to the use of builtin functions. To
# understand what is under the hood, please read the tutorial on page rank specified in this repository.
#
# __References__: <br />
# [1] Kipf, <NAME>., and <NAME>. "Semi-supervised classification with graph convolutional networks." arXiv preprint arXiv:1609.02907 (2016). <br />
# [2] <NAME>, <NAME>, and <NAME>. "Convolutional neural networks on graphs with fast localized spectral filtering." In Advances in Neural Information Processing Systems, pp. 3844-3852. 2016. <br />
# [3] Simonovsky, Martin, and <NAME>. "Dynamic edge-conditioned filters in convolutional neural networks on graphs." In Proc. CVPR. 2017. <br />
# + colab={"base_uri": "https://localhost:8080/", "height": 1295} colab_type="code" id="3lOBizVa4rVt" outputId="a3142dd2-4ff0-4bb6-a833-a7046f4e0596" language="bash"
# if [ ! -d "keras-deep-graph-learning" ] ; then git clone https://github.com/ypeleg/keras-deep-graph-learning; fi
# + colab={"base_uri": "https://localhost:8080/", "height": 1295} colab_type="code" id="3lOBizVa4rVt" outputId="a3142dd2-4ff0-4bb6-a833-a7046f4e0596"
from tachles import fix_gcn_paths, load_cora
# + colab={"base_uri": "https://localhost:8080/", "height": 1295} colab_type="code" id="3lOBizVa4rVt" outputId="a3142dd2-4ff0-4bb6-a833-a7046f4e0596"
fix_gcn_paths()
import keras_dgl
from keras_dgl.layers import GraphCNN, GraphAttentionCNN
from examples.utils import normalize_adj_numpy, evaluate_preds
# + [markdown] colab={"base_uri": "https://localhost:8080/", "height": 1295} colab_type="code" id="3lOBizVa4rVt" outputId="a3142dd2-4ff0-4bb6-a833-a7046f4e0596"
# ## The CORA Dataset
# The dataset used in this demo can be downloaded from https://linqs-data.soe.ucsc.edu/public/lbc/cora.tgz
#
# The following is the description of the dataset:
# > The Cora dataset consists of 2708 scientific publications classified into one of seven classes.
# > The citation network consists of 5429 links. Each publication in the dataset is described by a
# > 0/1-valued word vector indicating the absence/presence of the corresponding word from the dictionary.
# > The dictionary consists of 1433 unique words. The README file in the dataset provides more details.
#
# Download and unzip the cora.tgz file to a location on your computer and set the `data_dir` variable to
# point to the location of the dataset (the directory containing "cora.cites" and "cora.content").
# + colab={"base_uri": "https://localhost:8080/", "height": 1295} colab_type="code" id="3lOBizVa4rVt" outputId="a3142dd2-4ff0-4bb6-a833-a7046f4e0596"
X, Y_train, Y_test, A, train_idx, val_idx, test_idx, train_mask = load_cora()
print X.shape, Y_train.shape, Y_test.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 1295} colab_type="code" id="3lOBizVa4rVt" outputId="a3142dd2-4ff0-4bb6-a833-a7046f4e0596"
import keras.backend as K
import numpy as np
import networkx as nx
import matplotlib.pyplot as plt
from keras.layers import Dense, Activation, Dropout, Input
from keras.models import Model, Sequential
from keras.callbacks import Callback
from keras.regularizers import l2
from keras.optimizers import Adam
# + colab={"base_uri": "https://localhost:8080/", "height": 1295} colab_type="code" id="3lOBizVa4rVt" outputId="a3142dd2-4ff0-4bb6-a833-a7046f4e0596"
def plot_graph(adjacency_matrix):
rows, cols = np.where(adjacency_matrix == 1)
edges = zip(rows.tolist(), cols.tolist())
gr = nx.Graph()
gr.add_edges_from(edges)
fig, ax = plt.subplots(1, 1, figsize=(6, 6))
nx.draw_networkx(gr, ax=ax, with_labels=False, node_size=5, width=.5)
ax.set_axis_off()
plt.show()
plt.close()
# + colab={"base_uri": "https://localhost:8080/", "height": 1295} colab_type="code" id="3lOBizVa4rVt" outputId="a3142dd2-4ff0-4bb6-a833-a7046f4e0596"
print X[0]
plot_graph(A)
# + [markdown] colab={"base_uri": "https://localhost:8080/", "height": 1295} colab_type="code" id="3lOBizVa4rVt" outputId="a3142dd2-4ff0-4bb6-a833-a7046f4e0596"
# ## GraphCNN
#
# ```python
# GraphCNN(output_dim, num_filters, graph_conv_filters, activation=None, use_bias=True, kernel_initializer='glorot_uniform', bias_initializer='zeros', kernel_regularizer=None, bias_regularizer=None, activity_regularizer=None, kernel_constraint=None, bias_constraint=None)
# ```
#
# GraphCNN layer assumes a fixed input graph structure which is passed as a layer argument. As a result, the input order of graph nodes are fixed for the model and should match the nodes order in inputs. Also, graph structure can not be changed once the model is compiled. This choice enable us to use Keras Sequential API but comes with some constraints (for instance shuffling is not possible anymore in-or-after each epoch).<br />
#
#
# __Arguments__
#
# - __output_dim__: Positive integer, dimensionality of each graph node feature output space (or also referred dimension of graph node embedding).
# - __num_filters__: Positive integer, number of graph filters used for constructing __graph_conv_filters__ input.
# - __graph_conv_filters__ input as a 2D tensor with shape: `(num_filters*num_graph_nodes, num_graph_nodes)`<br />
# `num_filters` is different number of graph convolution filters to be applied on graph. For instance `num_filters` could be power of graph Laplacian. Here list of graph convolutional matrices are stacked along second-last axis.<br />
# - __activation__: Activation function to use
# (see [activations](https://keras.io/activations)).
# If you don't specify anything, no activation is applied
# (ie. "linear" activation: `a(x) = x`).
# - __use_bias__: Boolean, whether the layer uses a bias vector.
# - __kernel_initializer__: Initializer for the `kernel` weights matrix
# (see [initializers](https://keras.io/initializers)).
# - __bias_initializer__: Initializer for the bias vector
# (see [initializers](https://keras.io/initializers)).
# - __kernel_regularizer__: Regularizer function applied to
# the `kernel` weights matrix
# (see [regularizer](https://keras.io/regularizers)).
# - __bias_regularizer__: Regularizer function applied to the bias vector
# (see [regularizer](https://keras.io/regularizers)).
# - __activity_regularizer__: Regularizer function applied to
# the output of the layer (its "activation").
# (see [regularizer](https://keras.io/regularizers)).
# - __kernel_constraint__: Constraint function applied to the kernel matrix
# (see [constraints](https://keras.io/constraints/)).
# - __bias_constraint__: Constraint function applied to the bias vector
# (see [constraints](https://keras.io/constraints/)).
#
#
#
# __Input shapes__
#
# * 2D tensor with shape: `(num_graph_nodes, input_dim)` representing graph node input feature matrix.<br />
#
#
# __Output shape__
#
# * 2D tensor with shape: `(num_graph_nodes, output_dim)` representing convoluted output graph node embedding (or signal) matrix.<br />
#
#
#
# ----
# ## Remarks
#
# __Why pass graph_conv_filters as a layer argument and not as an input in GraphCNN?__<br />
# The problem lies with keras multi-input functional API. It requires --- all input arrays (x) should have the same number of samples i.e., all inputs first dimension axis should be same. In special cases the first dimension of inputs could be same, for example check out Kipf .et.al. keras implementation [[source]](https://github.com/tkipf/keras-gcn/blob/master/kegra/train.py). But in cases such as a graph recurrent neural networks this does not hold true.
#
#
# __Why pass graph_conv_filters as 2D tensor of this specific format?__<br />
# Passing graph_conv_filters input as a 2D tensor with shape: `(K*num_graph_nodes, num_graph_nodes)` cut down few number of tensor computation operations.
#
# __References__: <br />
# [1] Kipf, <NAME>., and <NAME>. "Semi-supervised classification with graph convolutional networks." arXiv preprint arXiv:1609.02907 (2016). <br />
# [2] Defferrard, Michaël, <NAME>, and <NAME>. "Convolutional neural networks on graphs with fast localized spectral filtering." In Advances in Neural Information Processing Systems, pp. 3844-3852. 2016. <br />
# [3] Simonovsky, Martin, and <NAME>. "Dynamic edge-conditioned filters in convolutional neural networks on graphs." In Proc. CVPR. 2017. <br />
#
#
# <span style="float:right;">[[source]](https://github.com/ypeleg/keras-deep-graph-learning/blob/master/examples/gcnn_node_classification_example.py)</span>
# + colab={"base_uri": "https://localhost:8080/", "height": 1295} colab_type="code" id="3lOBizVa4rVt" outputId="a3142dd2-4ff0-4bb6-a833-a7046f4e0596"
class EdgeEval(Callback):
def __init__(self):
super(EdgeEval, self).__init__()
def on_epoch_end(self, epoch, logs=None):
Y_pred = model.predict(X, batch_size=A.shape[0])
_, train_acc = evaluate_preds(Y_pred, [Y_train], [train_idx])
_, test_acc = evaluate_preds(Y_pred, [Y_test], [test_idx])
print("Epoch: {:04d}".format(epoch), "train_acc= {:.4f}".format(train_acc[0]), "test_acc= {:.4f}".format(test_acc[0]))
# + [markdown] colab={"base_uri": "https://localhost:8080/", "height": 1295} colab_type="code" id="3lOBizVa4rVt" outputId="a3142dd2-4ff0-4bb6-a833-a7046f4e0596"
# ## The model itself
# + colab={"base_uri": "https://localhost:8080/", "height": 1295} colab_type="code" id="3lOBizVa4rVt" outputId="a3142dd2-4ff0-4bb6-a833-a7046f4e0596"
A_norm = normalize_adj_numpy(A, True)
num_filters = 2
graph_conv_filters = np.concatenate([A_norm, np.matmul(A_norm, A_norm)], axis=0)
print graph_conv_filters.shape
graph_conv_filters = K.constant(graph_conv_filters)
# Build Model
inp = Input(shape=(X.shape[1],))
x = GraphCNN(16, num_filters, graph_conv_filters, activation='elu', kernel_regularizer=l2(5e-4))(inp)
x = Dropout(0.2)(x)
x = GraphCNN(Y_train.shape[1], num_filters, graph_conv_filters, activation='elu', kernel_regularizer=l2(5e-4))(x)
x = Activation('softmax')(x)
model = Model(inputs = inp, outputs = x)
model.compile(loss='categorical_crossentropy', optimizer=Adam(lr=0.01), metrics=['acc'])
model.summary()
# + colab={"base_uri": "https://localhost:8080/", "height": 1295} colab_type="code" id="3lOBizVa4rVt" outputId="a3142dd2-4ff0-4bb6-a833-a7046f4e0596"
model.fit(X, Y_train, sample_weight=train_mask, batch_size=A.shape[0], epochs=500, shuffle=False, callbacks=[EdgeEval()], verbose=1)
# + [markdown] colab={"base_uri": "https://localhost:8080/", "height": 1295} colab_type="code" id="3lOBizVa4rVt" outputId="a3142dd2-4ff0-4bb6-a833-a7046f4e0596"
# <span style="float:right;">[[source]](https://github.com/vermaMachineLearning/keras-deep-graph-learning/blob/master/keras_dgl/layers/graph_attention_cnn_layer.py#L10)</span>
# ## GraphAttentionCNN
#
# ```python
# GraphAttentionCNN(output_dim, adjacency_matrix, num_filters=None, graph_conv_filters=None, activation=None, use_bias=False, kernel_initializer='glorot_uniform', bias_initializer='zeros', kernel_regularizer=None, bias_regularizer=None, activity_regularizer=None, kernel_constraint=None, bias_constraint=None)
# ```
#
# GraphAttention layer assumes a fixed input graph structure which is passed as a layer argument. As a result, the input order of graph nodes are fixed for the model and should match the nodes order in inputs. Also, graph structure can not be changed once the model is compiled. This choice enable us to use Keras Sequential API but comes with some constraints (for instance shuffling is not possible anymore in-or-after each epoch). See further [remarks below](http://127.0.0.1:8000/Layers/Convolution/graph_conv_layer/#remarks) about this specific choice.<br />
#
#
# __Arguments__
#
# - __output_dim__: Positive integer, dimensionality of each graph node feature output space (or also referred dimension of graph node embedding).
# - __adjacency_matrix__: input as a 2D tensor with shape: `(num_graph_nodes, num_graph_nodes)` with __diagonal values__ equal to 1.<br />
# - __num_filters__: None or Positive integer, number of graph filters used for constructing __graph_conv_filters__ input.
# - __graph_conv_filters__: None or input as a 2D tensor with shape: `(num_filters*num_graph_nodes, num_graph_nodes)`<br />
# `num_filters` is different number of graph convolution filters to be applied on graph. For instance `num_filters` could be power of graph Laplacian. Here list of graph convolutional matrices are stacked along second-last axis.<br />
# - __activation__: Activation function to use
# (see [activations](../activations.md)).
# If you don't specify anything, no activation is applied
# (ie. "linear" activation: `a(x) = x`).
# - __use_bias__: Boolean, whether the layer uses a bias vector (recommended setting is False for this layer).
# - __kernel_initializer__: Initializer for the `kernel` weights matrix
# (see [initializers](../initializers.md)).
# - __bias_initializer__: Initializer for the bias vector
# (see [initializers](../initializers.md)).
# - __kernel_regularizer__: Regularizer function applied to
# the `kernel` weights matrix
# (see [regularizer](../regularizers.md)).
# - __bias_regularizer__: Regularizer function applied to the bias vector
# (see [regularizer](../regularizers.md)).
# - __activity_regularizer__: Regularizer function applied to
# the output of the layer (its "activation").
# (see [regularizer](../regularizers.md)).
# - __kernel_constraint__: Constraint function applied to the kernel matrix
# (see [constraints](https://keras.io/constraints/)).
# - __bias_constraint__: Constraint function applied to the bias vector
# (see [constraints](https://keras.io/constraints/)).
#
#
#
# __Input shapes__
#
# * 2D tensor with shape: `(num_graph_nodes, input_dim)` representing graph node input feature matrix.<br />
#
#
# __Output shape__
#
# * 2D tensor with shape: `(num_graph_nodes, output_dim)` representing convoluted output graph node embedding (or signal) matrix.<br />
#
#
# <span style="float:right;">[[source]](https://github.com/vermaMachineLearning/keras-deep-graph-learning/blob/master/examples/graph_attention_cnn_node_classification_example.py)</span>
# + colab={"base_uri": "https://localhost:8080/", "height": 1295} colab_type="code" id="3lOBizVa4rVt" outputId="a3142dd2-4ff0-4bb6-a833-a7046f4e0596"
A_norm = normalize_adj_numpy(A, True)
num_filters = 2
graph_conv_filters = np.concatenate([A_norm, np.matmul(A_norm, A_norm)], axis=0)
print graph_conv_filters.shape
graph_conv_filters = K.constant(graph_conv_filters)
# Build Model
inp = Input(shape=(X.shape[1],))
x = GraphAttentionCNN(8, A, num_filters, graph_conv_filters, num_attention_heads=8, attention_combine='concat', attention_dropout=0.6, activation='elu', kernel_regularizer=l2(5e-4))(inp)
x = Dropout(0.6)(x)
x = GraphAttentionCNN(Y_train.shape[1], A, num_filters, graph_conv_filters, num_attention_heads=1, attention_combine='average', attention_dropout=0.6, activation='elu', kernel_regularizer=l2(5e-4))(x)
x = Activation('softmax')(x)
model = Model(inputs = inp, outputs = x)
model.compile(loss='categorical_crossentropy', optimizer=Adam(lr=0.01), metrics=['acc'])
model.summary()
# + colab={"base_uri": "https://localhost:8080/", "height": 1295} colab_type="code" id="3lOBizVa4rVt" outputId="a3142dd2-4ff0-4bb6-a833-a7046f4e0596"
model.fit(X, Y_train, sample_weight=train_mask, batch_size=A.shape[0], epochs=500, shuffle=False, callbacks=[EdgeEval()], verbose=1)
| 1.0.1 - Graph Convolution Networks - Node Classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Shapelets and the Shapelet Transform with sktime
#
# [Github weblink](https://github.com/alan-turing-institute/sktime/blob/master/examples/shapelet_transform.ipynb)
#
# Introduced in [1], a shapelet is a time series subsequences that is identified as being representative of class membership. Shapelets are a powerful approach for measuring _phase-independent_ similarity between time series; they can occur at any point within a series and offer _interpretable_ results for how matches occur. The original research extracted shapelets to build a decision tree classifier. As an example, the top shapelet from the <a href="http://timeseriesclassification.com/description.php?Dataset=GunPoint">GunPoint</a> problem found in [1] is shown below (the shapelet is highlighted in red):
#
# 
#
# The GunPoint problem contains univariate tracking data of actor's wrists as they either take a prop gun from a holster, point it, and return it to the holster, or they simply mime the action without the prop. The classification problem is to detect whether a motion trace is a case of _gun_ or _no gun_. The most discriminatory shapelet shown above offers an interpretable result into how classification decisions can be made; the highlighted red section of the series above is the shapelet and it occurs when an actor returns the gun to the holster. As explained in [1], this is discriminatory because if the gun prop is present then the actor's hand stops suddenly, but if the prop is not present, the actor's hand will not stop abruptly and will subtly continue past their waist due to inertia.
#
# ## The Shapelet Transform
#
# Much research emphasis has been placed on shapelet-based approaches for time series classification (TSC) since the original research was proposed. The current state-of-the-art for shapelets is the **shapelet transform** (ST) [2, 3]. The transform improves upon the original use of shapelets by separating shapelet extraction from the classification algorithm, allowing interpretable phase-independent classification of time series with any standard classification algorithm (such as random/rotation forest, neural networks, nearest neighbour classifications, ensembles of all, etc.). To facilitate this, rather than recursively assessing data for the best shapelet, the transform evaluates candidate shapelets in a single procedure to rank them based on information gain. Then, given a set of _k_ shapelets, a time series can be transformed into _k_ features by calculating the distance from the series to each shapelet. By transforming a dataset in this manner any vector-based classification algorithm can be applied to a shapelet-transformed time series problem while the interpretability of shapelets is maintained through the ranked list of the _best_ shapelets during transformation. An example of the interpretability offered from ST is shown below with the top 10 shapelets extracted from the GunPoint problem in [2]:
#
# 
#
# The image above demonstrates that there are two clear groupings of shapelets: one where the gun is placed into the holster, as in [1], but also one where the gun is removed from the holster. It would be a reasonable assumption that if placing the prob back into the holster is discriminatory then removing it from the holster should also be; this was not detected in the original research but is explicitly identified by using the shapelet transform.
#
# #### References
# [1] <NAME>, and <NAME>. "Time series shapelets: a novel technique that allows accurate, interpretable and fast classification." Data mining and knowledge discovery 22, no. 1-2 (2011): 149-182.
#
# [2] <NAME>, <NAME>, <NAME>, and <NAME>. "A shapelet transform for time series classification." In Proceedings of the 18th ACM SIGKDD international conference on Knowledge discovery and data mining, pp. 289-297. ACM, 2012.
#
# [3] Hills, Jon, <NAME>, <NAME>, <NAME>, and <NAME>. "Classification of time series by shapelet transformation." Data Mining and Knowledge Discovery 28, no. 4 (2014): 851-881.
#
# [4] Bostrom, Aaron, and <NAME>. "Binary shapelet transform for multiclass time series classification." In Transactions on Large-Scale Data-and Knowledge-Centered Systems XXXII, pp. 24-46. Springer, Berlin, Heidelberg, 2017.
#
# ## Example: The Shapelet Transform in sktime
#
# The following workbook demonstrates a full workflow of using the shapelet transform in `sktime` with a `scikit-learn` classifier with the GunPoint problem.
#
#
# +
from sktime.datasets import load_gunpoint
from sktime.transformers.shapelets import ContractedRandomShapeletTransform
train_x, train_y = load_gunpoint(split='TRAIN', return_X_y=True)
test_x, test_y = load_gunpoint(split='TEST', return_X_y=True)
# +
# How long (in minutes) to extract shapelets for.
# This is a simple lower-bound initially; once time is up, no further shapelets will be assessed
time_limit_in_mins = 0.1
# The initial number of shapelet candidates to assess per training series. If all series are visited
# and time remains on the contract then another pass of the data will occur
initial_num_shapelets_per_case = 10
# Whether or not to print on-going information about shapelet extraction. Useful for demo/debugging
verbose = True
st = ContractedRandomShapeletTransform(
time_limit_in_mins=time_limit_in_mins,
initial_num_shapelets_per_case=initial_num_shapelets_per_case,
verbose=verbose)
st.fit(train_x, train_y)
# +
# %matplotlib inline
import matplotlib.pyplot as plt
# for each extracted shapelet (in descending order of quality/information gain)
for s in st.shapelets[0:5]:
# summary info about the shapelet
print(s)
# plot the series that the shapelet was extracted from
plt.plot(
train_x.iloc[s.series_id,0],
'gray'
)
# overlay the shapelet onto the full series
plt.plot(
list(range(s.start_pos,(s.start_pos+s.length))),
train_x.iloc[s.series_id,0][s.start_pos:s.start_pos+s.length],
'r',
linewidth=3.0
)
plt.show()
# +
# for each extracted shapelet (in descending order of quality/information gain)
for i in range(0,5):
s = st.shapelets[i]
# summary info about the shapelet
print("#"+str(i)+": "+str(s))
# overlay shapelets
plt.plot(
list(range(s.start_pos,(s.start_pos+s.length))),
train_x.iloc[s.series_id,0][s.start_pos:s.start_pos+s.length]
)
plt.show()
# +
import time
import numpy as np
from sktime.datasets import load_gunpoint
from sktime.transformers.shapelets import ContractedRandomShapeletTransform
from sklearn.ensemble.forest import RandomForestClassifier
from sklearn.pipeline import Pipeline
np.random.seed(seed=0)
train_x, train_y = load_gunpoint(split='TRAIN', return_X_y=True)
test_x, test_y = load_gunpoint(split='TEST', return_X_y=True)
# example pipleine with 1 minute time limit
pipeline = Pipeline([
('st', ContractedRandomShapeletTransform(time_limit_in_mins=0.1,
initial_num_shapelets_per_case=10,
verbose=False)),
('rf', RandomForestClassifier(n_estimators=100)),
])
start = time.time()
pipeline.fit(train_x, train_y)
end_build = time.time()
preds = pipeline.predict(test_x)
end_test = time.time()
print("Results:")
print("Correct:")
correct = sum(preds == test_y)
print("\t"+str(correct)+"/"+str(len(test_y)))
print("\t"+str(correct/len(test_y)))
print("\nTiming:")
print("\tTo build: "+str(end_build-start)+" secs")
print("\tTo predict: "+str(end_test-end_build)+" secs")
| examples/shapelet_transform.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import cv2
from os import path as osp
from glob import glob
import os
import sys
module_path = os.path.abspath(os.path.join('..'))
if module_path not in sys.path:
sys.path.append(module_path)
from superpoint.settings import EXPER_PATH
from superpoint.evaluations.evaluation_tool import keep_true_keypoints, select_k_best, warp_keypoints, filter_keypoints
from superpoint.evaluations.detector_evaluation import div0
import superpoint.evaluations.descriptor_evaluation as ev
from utils import plot_imgs
def keep_shared_points(data, keypoint_map, inv, keep_k_points=1000):
"""
Compute a list of keypoints from the map, filter the list of points by keeping
only the points that once mapped by H are still inside the shape of the map
and keep at most 'keep_k_points' keypoints in the image.
"""
keypoints = np.where(keypoint_map > 0)
prob = keypoint_map[keypoints[0], keypoints[1]]
#print(prob)
keypoints = np.stack([keypoints[0], keypoints[1], prob], axis=-1)
#print(keypoints)
keypoints = keep_true_keypoints(data, keypoints, inv)
#print(keypoints)
#print("soretd:")
idx = np.argsort(keypoints[:,2])[::-1]
#print(idx)
#print(keypoints[idx])
keypoints = select_k_best(keypoints, keep_k_points)
#print(keypoints)
return keypoints.astype(int)
def get_ground_truth(data, keypoints, warped_keypoints, shape, correctness_thresh, inv):
"""
Compute the ground truth keypoints matchings from image to image' where image' in the result
of warping image with H_matrix.
"""
#keypoints = np.stack([keypoints[0], keypoints[1]], axis=-1)
# Warp the original keypoints with the true homography
true_warped_keypoints = warp_keypoints(data, keypoints[:, [1, 0]], inv)
true_warped_keypoints = np.stack([true_warped_keypoints[:, 1],
true_warped_keypoints[:, 0]], axis=-1)
true_warped_keypoints = filter_keypoints(true_warped_keypoints, shape)
print(warped_keypoints)
print(true_warped_keypoints)
#print(true_warped_keypoints.shape)
#print(warped_keypoints.shape)
diff = np.expand_dims(warped_keypoints, axis=1) - np.expand_dims(true_warped_keypoints, axis=0)
dist = np.linalg.norm(diff, axis=-1)
#print(dist)
matches = np.less_equal(dist, correctness_thresh)
return matches, len(true_warped_keypoints)
def compute_pr_rec(prob, gt, n_gt, total_n, remove_zero=1e-4, simplified=False):
"""
computes precison and recall of the image
return: precision and recall
"""
matches = gt
#print(gt.shape)
tp = 0
fp = 0
tp_points = []
matched = np.zeros(len(gt))
for m in matches:
correct = np.any(m)
if correct:
gt_idx = np.argmax(m)
#tp +=1
#at most one tp should be considerd for each ground turth point
if gt_idx not in tp_points:
tp_points.append(gt_idx)
tp += 1
else:
fp += 1
else:
#tp.append(False)
fp += 1
#compute precison and recall
matching_score = tp / total_n if total_n!=0 else 0
prec = tp / (tp+fp) if (tp+fp)!=0 else 0
recall = tp / (n_gt) if n_gt!= 0 else 0
#print(n_gt)
#print(tp)
#print(fp)
return prec, recall, matching_score
def draw_matches(data):
keypoints1 = [cv2.KeyPoint(p[1], p[0], 1) for p in data['keypoints1']]
keypoints2 = [cv2.KeyPoint(p[1], p[0], 1) for p in data['keypoints2']]
inliers = data['inliers'].astype(bool)
matches = np.array(data['matches'])[inliers].tolist()
img1 = np.concatenate([output['image1'], output['image1'], output['image1']], axis=2) * 255
img2 = np.concatenate([output['image2'], output['image2'], output['image2']], axis=2) * 255
return cv2.drawMatches( np.uint8(img1), keypoints1, np.uint8(img2), keypoints2, matches,
None, matchColor=(0,255,0), singlePointColor=(0, 0, 255))
# +
data = np.load("0_.npz")
#prob = np.load("prob.npy")
#warped_prob = np.load("warped_prob.npy")
prob = data['prob']
warped_prob = data['warped_prob']
shape = prob.shape
warped_shape = warped_prob.shape
# Keeps only the points shared between the two views
keypoints = keep_shared_points(data, prob, False, 1000)
warped_keypoints = keep_shared_points(data, warped_prob, True, 1000)
desc = data['desc'][keypoints[:, 0], keypoints[:, 1]]
warped_desc = data['warped_desc'][warped_keypoints[:, 0],
warped_keypoints[:, 1]]
orb = False
# Match the keypoints with the warped_keypoints with nearest neighbor search
if orb:
desc = desc.astype(np.uint8)
warped_desc = warped_desc.astype(np.uint8)
bf = cv2.BFMatcher(cv2.NORM_HAMMING, crossCheck=True)
else:
bf = cv2.BFMatcher(cv2.NORM_L2, crossCheck=True)
matches = bf.match(desc, warped_desc)
matches_idx = np.array([m.queryIdx for m in matches]).astype(int)
m_keypoints = keypoints[matches_idx, :]
matches_idx = np.array([m.trainIdx for m in matches]).astype(int)
m_warped_keypoints = warped_keypoints[matches_idx, :]
print(m_keypoints.shape)
print(m_warped_keypoints.shape)
precisions = []
recalls = []
ms = []
for t in range(3,4):
#find ground truth
true_keypoints, n_gt1 = get_ground_truth(data, m_warped_keypoints, m_keypoints, warped_shape, t, inv=True)
true_warped_keypoints, n_gt2 = get_ground_truth(data, m_keypoints, m_warped_keypoints, shape, t, inv=False)
#calculate precison and recall
print(len(warped_keypoints))
prec1, recall1, ms1 = compute_pr_rec(m_warped_keypoints, true_warped_keypoints, n_gt2, len(warped_keypoints))
prec2, recall2, ms2 = compute_pr_rec(m_keypoints, true_keypoints, n_gt1, len(keypoints))
#average precison and recall for two images
prec = (prec1 + prec2)/2
recall = (recall1 + recall2)/2
matching_score = (ms1 + ms2)/2
precisions.append(prec)
recalls.append(recall)
ms.append(matching_score)
sum_ = 0
for i in range(1,len(precisions)):
sum_ += precisions[i]*np.abs((recalls[i]-recalls[i-1]))
mean_AP = np.sum(precisions[1:] * np.abs((np.array(recalls[1:]) - np.array(recalls[:-1]))))
print(precisions)
print(recalls)
#print(sum_)
print(mean_AP)
m = []
m = m + (ms)
m = m + (ms)
print(np.mean(m))
output = ev.compute_homography(data, 1000, 3, False)
output['image1'] = data['image']
output['image2'] = data['warped_image']
img = np.float32(draw_matches(output) / 255.)
plot_imgs([img], titles=["test"], dpi=200)
# -
| notebooks/debug_descriptor-evaluation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] heading_collapsed=true
# # Question 3: Object Identification
# + hidden=true
# Libraries
import numpy as np
from PIL import Image
import matplotlib.pyplot as plt
import histogram_module # functions for histograms
import gauss_module # functions for gaussian filters
import dist_module # functions for compute distances
import match_module # functions for finding matches
import rpc_module # functions for precision recall plots
import pandas as pd
from tqdm.notebook import tqdm
import warnings
warnings.filterwarnings("ignore")
from sklearn.model_selection import ParameterGrid # useful in implementig the grid search
# + hidden=true
# Load data
with open('model.txt') as fp:
model_images = fp.readlines()
model_images = [x.strip() for x in model_images]
with open('query.txt') as fp:
query_images = fp.readlines()
query_images = [x.strip() for x in query_images]
# Histogram and distance specifications
dist_type = 'intersect'
hist_type = 'rg'
num_bins = 30
# + [markdown] heading_collapsed=true
# # a)
# Having implemented different distance functions and image histograms, we can now test how suitable
# they are for retrieving images in query-by-example scenario. Implement a function find best match, in
# match module.py, which takes a list of model images and a list of query images and for each query im-
# age returns the index of closest model image. The function should take string parameters, which identify
# distance function, histogram function and number of histogram bins. See comments in the beginning of
# find best match for more details. Additionally to the indices of the best matching images your implemen-
# tation should also return a matrix which contains distances between all pairs of model and query images.
# + hidden=true
## Find best match (Question 3.a)
[best_match, D] = match_module.find_best_match(model_images, query_images, dist_type, hist_type, num_bins)
# + hidden=true
print(D)
# + hidden=true
print(best_match)
# + [markdown] heading_collapsed=true
# # b)
# Implement a function show neighbors (in match module.py) which takes a list of model images and a list of
# query images and for each query image visualizes several model images which are closest to the query image
# according to the specified distance metric. Use the function find best match in your implementation.
# + hidden=true
## visualize nearest neighbors (Question 3.b)
query_images_vis = [query_images[i] for i in np.array([0,4,9])]
match_module.show_neighbors(model_images, query_images_vis, dist_type, hist_type, num_bins)
# -
# # c)
# Use the function find best match to compute recognition rate for different combinations of distance and
# histogram functions. The recognition rate is given by a ratio between number of correct matches and total
# number of query images. Experiment with different functions and numbers of histogram bins, try to find
# combination that works best. Submit the summary of your experiments in a report as part of your solution.
# +
## compute recognition percentage (Question 3.c)
num_correct = sum(best_match == range(len(query_images)))
print('number of correct matches: %d (%f)'% (num_correct, 1.0 * num_correct / len(query_images)))
# +
# Compute recognition rate using a function that can take also slices of query images
# Using the same specification as before, we get same result
match_module.recognition_rate(model_images, query_images, query_images, dist_type, hist_type, num_bins)
# +
# Different combinations used for distance functions, histogram functions and number of bins
hyper_grid_cv = {'dist_type':['intersect', 'chi2', 'l2'], 'hist_type':['grayvalue', 'rgb', 'rg', 'dxdy'],
'num_bins':[5, 10, 30, 50]}
# Find recognition rate for each combination
match_module.grid_search(hyper_grid_cv, model_images, query_images, query_images)
# Show results
rec_rate = pd.read_csv('recognition_rates.csv')
rec_rate.sort_values('recognition_rate', ascending=False)
# -
# # Question 4: Performance Evaluation (bonus of 10 points)
#
# +
with open('model.txt') as fp:
model_images = fp.readlines()
model_images = [x.strip() for x in model_images]
with open('query.txt') as fp:
query_images = fp.readlines()
query_images = [x.strip() for x in query_images]
# -
for num_bins in [5,10,20,50]:
plt.figure(figsize=(8, 6))
rpc_module.compare_dist_rpc(model_images, query_images, ['chi2', 'intersect', 'l2'], 'rg', num_bins, ['r', 'g', 'b'])
plt.title('RG histograms'+str(num_bins))
plt.savefig('.\plots\RG_histograms'+str(num_bins))
plt.figure(figsize=(8, 6))
rpc_module.compare_dist_rpc(model_images, query_images, ['chi2', 'intersect', 'l2'], 'rgb', num_bins, ['r', 'g', 'b'])
plt.title('RGB histograms'+str(num_bins))
plt.savefig('.\plots\RGB_histograms'+str(num_bins))
plt.figure(figsize=(8, 6))
rpc_module.compare_dist_rpc(model_images, query_images, ['chi2', 'intersect', 'l2'], 'dxdy', num_bins, ['r', 'g', 'b'])
plt.title('dx/dy histograms'+str(num_bins))
plt.savefig('.\plots\dxdy_histograms'+str(num_bins))
plt.figure(figsize=(8, 6))
rpc_module.compare_dist_rpc(model_images, query_images, ['chi2', 'intersect', 'l2'], 'grayvalue', num_bins, ['r', 'g', 'b'])
plt.title('grayvalue histograms'+str(num_bins))
plt.savefig('.\plots\grayvalue_histograms'+str(num_bins))
| Assignment/Identification/Identification_Q3_Q4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %cd ~/NetBeansProjects/ExpLosion/
from notebooks.common_imports import *
from gui.user_code import get_demsar_diagram, pairwise_significance_exp_ids
from gui.output_utils import *
sns.timeseries.algo.bootstrap = my_bootstrap
sns.categorical.bootstrap = my_bootstrap
# -
# # word2vec vs glove vs count @ wikipedia
# +
d = {'expansions__noise': 0.0,
'expansions__decode_handler': 'SignifiedOnlyFeatureHandler',
'expansions__k': 3,
'expansions__vectors__rep': 0,
'expansions__vectors__unlabelled_percentage': 100.0,
'expansions__vectors__unlabelled' : 'wiki',
'expansions__allow_overlap': 0,
'labelled': 'amazon_grouped-tagged',
'expansions__vectors__algorithm': 'word2vec',
'document_features_tr': 'J+N+AN+NN',
'document_features_ev': 'AN+NN',
'expansions__vectors__unlabelled_percentage':100,
'expansions__entries_of':None,
'expansions__vectors__composer__in': ['Add', 'Mult', 'Right', 'Left']
}
def _get(d):
return [x.id for x in Experiment.objects.filter(**d).\
order_by('expansions__vectors__algorithm', 'expansions__vectors__composer')]
w2v = _get(d)
d['expansions__vectors__algorithm'] = 'glove'
glove = _get(d)
d['expansions__vectors__algorithm'] = 'count_windows'
wins = _get(d)
d['expansions__vectors__algorithm'] = 'count_dependencies'
deps = _get(d)
print('wins', wins, '\nglove', glove, '\nw2v', w2v)
horder = ['deps', 'wins', 'glove', 'word2vec']
xorder = ['Add', 'Mult', 'Left', 'Right']
with sns.color_palette("cubehelix", 4):
diff_plot_bar( [deps, wins, glove, w2v], horder,
[Experiment.objects.get(id=id).expansions.vectors.composer for id in w2v],
xlabel='', hline_at=random_vect_baseline(),
hue_order=horder, order=xorder)
plt.savefig('plot-w2v-vs-glove.pdf', format='pdf', dpi=300, bbox_inches='tight', pad_inches=0.1)
# sanity check
from itertools import chain
for id in chain.from_iterable([wins, glove, w2v]):
print('id %d, score %.2f'%(id, Results.objects.get(id=id, classifier=CLASSIFIER).accuracy_mean))
# -
# # Are differences significant?
pairwise_significance_exp_ids(zip(wins,deps), ['expansions__vectors__composer',
'expansions__vectors__algorithm'])
pairwise_significance_exp_ids(zip(wins,glove), ['expansions__vectors__composer',
'expansions__vectors__algorithm'])
| notebooks/w2v_vs_glove_vs_count_NPs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # **Review of AlphaGo algorithm by Google DeepMind**
# by <NAME>
# **Summary of paper goals and techiniques**
#
# The board Go is viewed as the most challenging board games to be tackled by artificial intelligence. Since the search space is enourmous, it is unfeasible to train traditional search algorithms to build a game-playing agent. Instead, such artificial agent must, somehow, reduce the search space and intelligently evaluate board positions, without compromising performance.
#
# The novel framework proposed by Google DeepMind [1], the AlphaGo, combines several techniques that to predict and estimate of current and future moves and, therefore, is feasible to be deployed at runtime. The team come up with a pipeline of techniques that takes advantage of human expertise as well as self-play simulations evaluated by machine learning techniques.
#
# The pipeline consists of a Supervised Learning (SL) policy network, taken from expert human moves. Such step provides fast and efficient learning to begin. Afterwards, a Reinforcement Learning (RL) policy is trained to update the results from the SL policy, maximizing it towards winnings of games of self-play. This policy network combined outputs a probability distribution over legal moves at the current state. A value network is then trained by regression to predict the value of each position, i.e. whether the current player wins, from games of self-play of the RL network. A Monte Carlo Tree Search (MCTS) effectively evaluates the value of each position in the tree and outputs the most promising one.
# **Results**
# AlphaGo outperformed other Go programs 99.8% of the time and defeated, for the first time, the European Go champion by 5 games to 0.
# **References**
# [1] <NAME>, et al. "Mastering the game of Go with deep neural networks and tree search." nature 529.7587 (2016): 484.
| projects/isolation/research_review.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.10 64-bit (windows store)
# language: python
# name: python3
# ---
# !nvidia-smi --help
# gpu_info = !nvidia-smi --list-gpus
print(gpu_info)
# +
# gpu_detail = !nvidia-smi --query --id=0 --xml-format
detailmatcher = re.compile('^<([a-z_]+)>(.+)<\/[a-z_]+>')
for line in gpu_detail:
cleanline = line.strip()
keyvalue = detailmatcher.findall(cleanline)
if len(keyvalue) == 0:
continue
if keyvalue[0][0] == "driver_version":
print(f' Driver Version: {keyvalue[0][1]}')
if keyvalue[0][0] == "cuda_version":
print(f' CUDA Version: {keyvalue[0][1]}')
if keyvalue[0][0] == "product_brand":
print(f' Brand: {keyvalue[0][1]}')
if keyvalue[0][0] == "product_architecture":
print(f' Architecture: {keyvalue[0][1]}')
if keyvalue[0][0] == "total":
print(f' Memory: {keyvalue[0][1]}')
# +
import re
regex = r"^GPU ([0-9+]): (.+) \(UUID: ([a-zA-Z0-9-]+)\)$"
matcher = re.compile(regex)
# gpus_infos = !nvidia-smi --list-gpus
for gpu in gpus_info:
gpu_info = matcher.findall(gpu)
gpu_id = gpu_info[0][0]
gpu_name = gpu_info[0][1]
print(f'GPU {gpu_id}: {gpu_name}')
# gpu_detail = !nvidia-smi --query --id=$gpu_id --xml-format
detailmatcher = re.compile('^<([a-z_]+)>(.+)<\/[a-z_]+>')
for line in gpu_detail:
cleanline = line.strip()
keyvalue = detailmatcher.findall(cleanline)
if len(keyvalue) == 0:
continue
if keyvalue[0][0] == "driver_version":
print(f' Driver Version: {keyvalue[0][1]}')
if keyvalue[0][0] == "cuda_version":
print(f' CUDA Version: {keyvalue[0][1]}')
if keyvalue[0][0] == "product_brand":
print(f' Brand: {keyvalue[0][1]}')
if keyvalue[0][0] == "product_architecture":
print(f' Architecture: {keyvalue[0][1]}')
if keyvalue[0][0] == "total":
print(f' Memory: {keyvalue[0][1]}')
# -
# !nvidia-smi --query --xml-format
# +
import xml.etree.ElementTree as ET
# gpu_detail = !nvidia-smi --query --xml-format
root = ET.ElementTree(ET.fromstringlist(gpu_detail)).getroot()
for child in root:
if child.tag == 'driver_version':
print(f'NVIDIA Driver Version: {child.text}')
elif child.tag == 'cuda_version':
print(f'CUDA Version: {child.text}')
elif child.tag == 'gpu':
print(f'{child.attrib["id"]}:')
for gpu_property in child:
if gpu_property.tag == 'product_name':
print(f' Product Name : {gpu_property.text}')
elif gpu_property.tag == 'product_brand':
print(f' Brand : {gpu_property.text}')
elif gpu_property.tag == 'product_architecture':
print(f' Architecture : {gpu_property.text}')
elif gpu_property.tag == 'uuid':
print(f' UUID : {gpu_property.text}')
elif gpu_property.tag == 'fb_memory_usage':
for mem in gpu_property:
if mem.tag == 'total':
print(f' Total Memory : {mem.text}')
| get_gpu.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # DSND Project - Communicating with Stakeholders
# ## Business understanding
# This notebook is analyses of Airbnb data for seattle city. This analysis will be based on the CRISP-DM process.
#
# The analysis is used to answer the following questions about the data:
# - What are the top contributing features for pricing?
# - How are Airbnb prices distributed across top 3 features?
# - What are the most expensive zipcodes in Seattle?
#
# ## Data understanding
# ### Setup - Load the required python libraries
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
# -
# ### Get the data and verify it
calendar = pd.read_csv('calendar.csv')
listings = pd.read_csv('listings.csv')
reviews = pd.read_csv('reviews.csv')
# #### Verify if data is loaded properly
calendar.head(5)
listings.head(5)
reviews.head(5)
# ### Do basic analysis of data
# #### The most useful data for our analysis is available in the listings file and hence our focus from now on, will be on analysing the data within this dataframe
listings.shape
# We have 3818 listings (rows) and each listing is made up of 92 features (columns)
# #### Now lets look at the features of each row
listings.info();
# #### Analysing the features from an Airbnb host perspective, i think i would use a sub-set of these features to price and list my property.
# #### Similarly, analysing the features from an Airbnb customer perspective, i identified another sub-set of these features which would help me decide if i would select and stay in a given property.
# #### I combined both sub-sets to come up with a list of features that would be useful in answering the questions listed above.
#
# #### The features that are most relevant for our analysis are:
# - host_response_time
# - host_response_rate
# - zipcode
# - property_type
# - room_type
# - accommodates
# - bathrooms
# - bedrooms
# - beds
# - price
# - minimum_nights
# - number_of_reviews
# - review_scores_rating
# - reviews_per_month
# ## Data wrangling
# ### To speed up processing, lets remove unwanted features from the data
# +
needed_features = [ 'id', 'host_response_time' , 'host_response_rate' , 'zipcode' , 'property_type',
'room_type' , 'accommodates' , 'bathrooms' , 'bedrooms' , 'beds' , 'price' ,
'minimum_nights' , 'number_of_reviews' , 'review_scores_rating' , 'reviews_per_month']
new_listings = listings[needed_features].copy()
new_listings.shape
# -
new_listings.info();
# ### Now let's wrangle each feature to ensure that they contain valid data and in a format useful for modeling
# #### host_response_time
# host_response_time is a categorical feature with text values such as within an hour, within a few hours, within a day and a few days or more. This feature also contains missing values. We will conver this field into numeric values as below
#
# 0 - within an hour
#
# 1 - within a few hours
#
# 2 - within a day
#
# 3 - a few days or more
#
new_listings.host_response_time
new_listings.host_response_time.unique()
new_listings.replace({'within an hour': 0,
'within a few hours': 1,
'within a day': 2,
'a few days or more': 3}, inplace=True)
new_listings.host_response_time.unique()
new_listings.host_response_time
# #### Since host_response_time is a categorical feature, it makes sence to replace missing values with Mode of existing data i.e most frequently occuring data
new_listings['host_response_time'].fillna(new_listings['host_response_time'].mode()[0], inplace=True)
new_listings.host_response_time
# #### host_response_rate
# host_response_rate is a percentage value and contains many missing values. We'll replace missing values with Mean value
new_listings.head()
new_listings['host_response_rate'].fillna(0, inplace=True)
new_listings['host_response_rate'] = new_listings['host_response_rate'].astype(str)
new_listings['host_response_rate'] = new_listings['host_response_rate'].str.replace(r'%', '')
new_listings['host_response_rate'] = new_listings['host_response_rate'].astype(int)
new_listings['host_response_rate']=new_listings['host_response_rate'].mask(new_listings['host_response_rate']==0).fillna(new_listings['host_response_rate'].mean())
new_listings.host_response_rate.unique()
new_listings.head()
# #### Zipcodes
# zipcode is a numerical value and contains some missing values. We'll drop those rows since it's not practical to assume a zip code for a listing. This impacts only 7 rows which would not make any big difference to overall analysis
new_listings = new_listings.dropna(subset=['zipcode'])
new_listings.zipcode.unique()
# #### Also one of the zipcode is wrongly input. We'll correct the value manully
new_listings['zipcode'] = new_listings['zipcode'].astype(str)
new_listings['zipcode'] = new_listings['zipcode'].str.replace(r'99\n', '')
new_listings.zipcode.unique()
# #### property_type
# property_type is a categorical feature with text values. We will conver this field into numeric values as below
#
# 0 - Apartment
#
# 1 - House
#
# 2 - Cabin
#
# 3 - Condominium
#
# 4 - Camper/RV
#
# 5 - Bungalow
#
# 6 - Townhouse
#
# 7 - Loft
#
# 8 - Boat
#
# 9 - Dorm
#
# 10 - Bed & Breakfast
#
# 11 - Yurt
#
# 12 - Chalet
#
# 13 - Tent
#
# 14 - Treehouse
#
# 15 - Other
new_listings[new_listings['property_type'].isnull()]
# #### Since there is only one row with missing info, we'll replace this data with 'other' category
new_listings['property_type'].fillna(15, inplace=True)
new_listings.property_type.unique()
new_listings.replace({ 'Apartment': 0,
'House':1,
'Cabin':2,
'Condominium':3,
'Camper/RV':4,
'Bungalow':5,
'Townhouse':6,
'Loft':7,
'Boat':8,
'Dorm':9,
'Bed & Breakfast':10,
'Yurt':11,
'Chalet':12,
'Tent':13,
'Treehouse':14,
'Other':15
}, inplace=True)
new_listings.property_type.unique()
# #### room_type
# room_type is a categorical feature with text values. We will conver this field into numeric values as below
#
# 0 - Entire home/apt
#
# 1 - Entire home/apt
#
# 2 - Shared room
new_listings[new_listings['room_type'].isnull()]
# #### There are no missing values in this feature
new_listings.room_type.unique()
new_listings.replace({ 'Entire home/apt': 0,
'Private room':1,
'Shared room':2,
}, inplace=True)
new_listings.room_type.unique()
# #### accommodates
# Accommodates is already a numeric column with data in correct format
new_listings[new_listings['accommodates'].isnull()]
# #### There are no missing values in this feature
new_listings.accommodates.unique()
# #### bathrooms
# The number of bathrooms in an Airbnb property in Seattle is rightly formated as float64 so their no need to wrangle them
new_listings[new_listings['bathrooms'].isnull()]
# #### Since it is dificult to assume a value for number of bathrooms, we'll use the mode of the column i.e most occuring number
new_listings['bathrooms'].fillna(new_listings['bathrooms'].mode()[0], inplace=True)
new_listings[new_listings['bathrooms'].isnull()]
# #### bedrooms
# The number of bedrooms in an Airbnb property in Seattle is rightly formated as float64 so their no need to wrangle them
new_listings[new_listings['bedrooms'].isnull()]
# #### Since it is dificult to assume a value for number of bedrooms, we'll use the mode of the column i.e most occuring number
new_listings['bedrooms'].fillna(new_listings['bedrooms'].mode()[0], inplace=True)
new_listings[new_listings['bedrooms'].isnull()]
# #### beds
# The number of beds in an Airbnb property in Seattle is rightly formated as float64 so their no need to wrangle them
new_listings[new_listings['beds'].isnull()]
# #### Since it is dificult to assume a value for number of beds, we'll use the mode of the column i.e most occuring number
#
new_listings['beds'].fillna(new_listings['beds'].mode()[0], inplace=True)
new_listings[new_listings['beds'].isnull()]
# #### price
# Price column values are strings. We need to convert them to integers. Before we can do that, we need to remove the '$' sign from tghe price dsta
new_listings[new_listings['price'].isnull()]
new_listings.price
def valid_prices(x):
"""Make the price colums valid by removing $ signs and converting to float"""
x = float(x[1:].replace(',',''))
return x
new_listings['price'] = new_listings['price'].apply(lambda x: valid_prices(x))
new_listings.price
# #### minimum_nights
# The minimum_nights feature of the listing is rightly formatted in float format and doesn't need any wrangling.
# ##### The max value of this column seems a bit extreme at 1000 but for now i'll ignore this to see how it impacts the analysis.
new_listings[new_listings['minimum_nights'].isnull()]
new_listings.minimum_nights.describe()
# #### number_of_reviews
# This column too is rightly formatted for our needs and doesn't need any wrangling
new_listings[new_listings['number_of_reviews'].isnull()]
new_listings.number_of_reviews.unique()
# #### review_scores_rating
# This column too is rightly formatted for our needs and doesn't need any wrangling
# But there are lots of rows with missing values for this feature. We'll replace the missing values with median of existing review ratings
new_listings[new_listings['review_scores_rating'].isnull()]
new_listings['review_scores_rating'].fillna(new_listings['review_scores_rating'].median(), inplace=True)
new_listings[new_listings['review_scores_rating'].isnull()]
new_listings.review_scores_rating.describe()
# #### reviews_per_month
# This column too is rightly formatted for our needs and doesn't need any wrangling. But there are a lot of missing values and we'll replace them with median value
new_listings[new_listings['reviews_per_month'].isnull()]
new_listings['reviews_per_month'].fillna(new_listings['reviews_per_month'].median(), inplace=True)
new_listings[new_listings['reviews_per_month'].isnull()]
new_listings.reviews_per_month.describe()
# ## Data Modeling
# #### Before modeling the data, let's verify that the dataset is in required format and doesn't contain any missing values
new_listings.isnull().sum()[new_listings.isnull().sum()>0]
new_listings.head()
# ### Question 1 - What are the top contributing features for pricing?
#
# To look at the features that most influence the price, we'll use pearson correlation of all the features. This needs all features to be in numerical format.
new_listings = new_listings.apply(pd.to_numeric)
new_listings.head()
# +
features = ['host_response_time' , 'host_response_rate' , 'zipcode' , 'property_type',
'room_type' , 'accommodates' , 'bathrooms' , 'bedrooms' , 'beds' , 'price' ,
'minimum_nights' , 'number_of_reviews' , 'review_scores_rating' , 'reviews_per_month']
pcorr = np.corrcoef(new_listings[features].values.T)
fig, ax = plt.subplots(figsize=(14,14))
sns.set(font_scale=1)
heatmap = sns.heatmap(pcorr, cbar = True, annot=True, square = True, fmt = '.2f',yticklabels = features, xticklabels = features)
# -
# ### From the model above, we can determine that the following features have the most influence on price, based on Airbnb Seattle dataset
# 1. Accommodates
# 2. Bedrooms
# 3. Beds
# 4. Bathrooms
# ### Question 2 - How are Airbnb prices distributed across top 3 features?
# #### 2.1 To look at the correlation of price against Accommodates, Let's look at a scatter plot for price vs accommodate
plt.scatter(new_listings['price'],new_listings['accommodates'])
plt.ylabel('Accommodates')
plt.xlabel('Price in $')
plt.title('Accommodates vs price')
# #### From the above plot, it is clear that the price increases based on the Number of people a property can accommodate.
# #### 2.2 To look at the correlation of price against Bedrooms, Let's look at a scatter plot for price vs bedrooms
plt.scatter(new_listings['price'],new_listings['bedrooms'])
plt.ylabel('Number of Bedrooms')
plt.xlabel('Price in $')
plt.title('Number of Bedrooms vs price')
# #### From the above plot, it is clear that the price increases based on the Number of bedrooms a property contains. Also most of the properties in Seattle seems to offer upto 3 bedrooms and number of data points above that is very little
# #### 2.3 To look at the correlation of price against Number of Beds, Let's look at a scatter plot for price vs beds
plt.scatter(new_listings['price'],new_listings['beds'])
plt.ylabel('Number of Beds')
plt.xlabel('Price in $')
plt.title('Number of Beds vs price')
# #### From the above plot, it is clear that the price increases based on the Number of beds a property offers. Also most of the properties in Seattle seems to offer upto 6 beds and number of data points above that is very little
# ### Question 3 - What are the most expensive zipcodes in Seattle?
# Now to test our model, lets look the correlation of price agains another feature - Zipcode
plt.scatter(new_listings['price'],new_listings['zipcode'])
plt.ylabel('Zipcode')
plt.xlabel('Price in $')
plt.title('Zipcode vs price')
# #### From the above plot, it is clear that the price is not influenced by the zipcode of a property.
# #### Most of Airbnb properies in Seattle seems to be priced below 400 USD irrespective of their zipcode
# ## Conclusion
# The 3 features that have the highest correlation with price are:
#
# 1 - Accomodates
#
# 2- Bedrooms
#
# 3 - Beds
#
# Zipcodes or the location of the property has no influence on the price. Most of the properties in Seattle are priced below USD 400.
#
# Other features such as host reponse time or rate and overall reviews of the property also don't have any influence on the price.
| DSND-Airbnb-Seattle.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import matplotlib
import matplotlib.pyplot as plt
from datasets import get_dataset
import numpy as np
np.random.seed(123)
import random
random.seed(123)
import tensorflow as tf
from keras.backend.tensorflow_backend import set_session
config = tf.ConfigProto()
config.gpu_options.allow_growth=True
sess = tf.Session(config=config)
set_session(tf.Session(config=config))
from keras.utils import plot_model
from keras.models import Sequential, load_model
from keras.layers import Dense, Activation, Flatten, BatchNormalization, Dropout, Reshape
from keras.optimizers import Adadelta, SGD
from keras.callbacks import TensorBoard, EarlyStopping, ModelCheckpoint
from sklearn.cluster import KMeans
from sklearn.preprocessing import label_binarize
import cv2
import pdb
import progressbar
import os
from scipy import stats
from sklearn.metrics import accuracy_score
# -
n_views = 5
datasets = ['australian', 'bupa', 'colic', 'diabetes', 'german', 'ionosphere', 'kr-vs-kp', 'tic-tac-toe', 'vote', 'wdbc']
dataset = datasets[9]
# +
# Algorithm:
# Input: L, U, Learn
# Output: H
# 1. Partition L into (v_1, v_2, ..., v_n)
# 2. Learn h_i on v_i using Learn
# 3. while (one of h_i changes):
# 3.1 Q = []
# 3.2 for i in range(n):
# if majority on classifiers other than h_i has better accuracy on L:
# for u in U:
# if more than 50% of classifiers other than h_i agree on u:
# Q_i = Q_i Union {u, prediction(u)}
# 3.3 for i in range(n):
# Learn h_i on L Union q_i
# 4. Output H = Majority Over h_i
# -
def find_optimal_n(L_x, lower_cap=3, upper_cap=6):
min_counts = []
clusters = []
for find_n in range(lower_cap, upper_cap+1):
kmeans = KMeans(n_clusters=find_n, random_state=0).fit_predict(L_x)
clusters.append(kmeans)
_, counts = np.unique(kmeans, return_counts=True)
min_counts.append(min(counts))
return lower_cap + np.argmax(min_counts), clusters[np.argmax(min_counts)]
# +
# 1. Partition L into (v_1, v_2, ..., v_n)
ds = get_dataset(dataset, 0.7, 0.25)
[L_x, L_y], U, [test_x, test_y] = ds.get_data()
n_views, kmeans = find_optimal_n(L_x)
V = []
for ind in range(n_views):
left = int(ind * L_x.shape[0] / n_views)
right = int((ind+1) * L_x.shape[0] / n_views)
indices = np.where(kmeans == ind)
print L_x[indices].shape
# V.append([L_x[left:right], L_y[left:right]])
V.append([L_x[indices], L_y[indices]])
# +
# 2. Learn h_i on v_i using Learn
H = []
n_attr = V[ind][0].shape[1]
for ind in range(n_views):
h = Sequential()
h.add(Dense(input_shape=(n_attr,), units=n_attr / 2))
h.add(Activation('relu'))
h.add(BatchNormalization())
h.add(Dense(units=n_attr/5))
h.add(Activation('relu'))
h.add(BatchNormalization())
h.add(Dropout(0.5))
h.add(Dense(units=V[ind][1].shape[1]))
h.add(Activation('softmax'))
h.compile(loss='categorical_crossentropy', optimizer=Adadelta(), metrics=['accuracy'])
H.append(h)
for ind in range(n_views):
H[ind].fit(V[ind][0], V[ind][1], epochs=50, batch_size=32, validation_split=0.2, verbose=True)
print H[ind].evaluate(test_x,test_y)
# +
# 3
changed = True
to_plot = []
num_runs = 0
while (changed and num_runs <= 10):
preds_L = []
for ind in range(n_views):
preds_L.append(H[ind].predict(L_x))
preds_L = np.array(preds_L)
preds_U = []
for ind in range(n_views):
preds_U.append(H[ind].predict(U))
preds_U = np.array(preds_U)
perfs = []
test_preds = []
for ind in range(n_views):
test_preds.append(H[ind].predict(test_x))
test_preds = np.array(test_preds)
for ind in range(n_views):
perf = accuracy_score(np.argmax(test_y, axis=1), np.argmax(test_preds[ind], axis=1))
perfs.append(perf)
perfs.append(accuracy_score(np.argmax(test_y, axis=1),
np.argmax(np.sum(test_preds, axis=0), axis=1)))
to_plot.append(perfs)
Q = []
update = [False for _ in range(n_views)]
for cur in range(n_views):
elems_take = [view_ind for view_ind in range(n_views) if view_ind != cur]
preds_others_L = preds_L[elems_take]
preds_others_U = preds_U[elems_take]
acc_others_L = accuracy_score(np.argmax(L_y, axis=1),
np.argmax(np.sum(preds_others_L, axis=0), axis=1))
acc_cur_L = accuracy_score(np.argmax(L_y, axis=1), np.argmax(preds_L[cur], axis=1))
q_cur = [[], []]
if acc_others_L > acc_cur_L:
update[cur] = True
for u_ind in range(preds_U.shape[1]):
sum_prediction = np.argmax(np.sum(preds_others_U[:, u_ind], axis=0))
if np.sum(np.argmax(preds_others_U[:, u_ind], axis=1) == sum_prediction) >= 0.5 * (n_views - 1):
q_cur[0].append(U[u_ind])
label_temp = [0, 0]; label_temp[sum_prediction] = 1
q_cur[1].append(label_temp)
Q.append([np.array(q_cur[0]), np.array(q_cur[1])])
for cur in range(n_views):
if update[cur]:
comb_x = np.concatenate([L_x, Q[cur][0]], axis=0)
comb_y = np.concatenate([L_y, Q[cur][1]], axis=0)
H[cur].fit(comb_x, comb_y, epochs=20, batch_size=32, validation_split=0.2, verbose=False)
preds_L_new = []
for ind in range(n_views):
preds_L_new.append(H[ind].predict(L_x))
preds_U_new = []
for ind in range(n_views):
preds_U_new.append(H[ind].predict(U))
preds_L_red = np.argmax(np.sum(np.array(preds_L),axis=0),axis=1)
preds_L_new_red = np.argmax(np.sum(np.array(preds_L_new),axis=0),axis=1)
preds_U_red = np.argmax(np.sum(np.array(preds_U),axis=0),axis=1)
preds_U_new_red = np.argmax(np.sum(np.array(preds_U_new),axis=0),axis=1)
same = np.all(preds_L_red == preds_L_new_red) and np.all(preds_U_red == preds_U_new_red)
changed = not same
num_runs += 1
plt.figure(figsize=(7, 7))
plt.clf()
handles = []
labels = []
for ind in range(n_views):
ys = [x[ind] for x in to_plot]
handle, = plt.plot(range(len(to_plot)), ys, marker='o', label = str(ind))
handles.append(handle)
labels.append('Classifier %d' % ind)
ys = [x[n_views] for x in to_plot]
handle, = plt.plot(range(len(to_plot)), ys, marker='o', label = 'Overall')
handles.append(handle)
labels.append('Sum based')
plt.legend(handles, labels, bbox_to_anchor=(1.04,1), loc="upper left")
# plt.legend(handles, labels)
plt.xlabel('Iterations')
plt.ylabel('Accuracy')
plt.title('Iterations vs accuracy for Dataset: %s' % dataset)
plt.show()
# -
| n-view-alg5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="wl_e-LwlWi9-" colab_type="code" colab={}
import tensorflow as tf
# + id="tUvOUUksWr0D" colab_type="code" colab={}
from tensorflow.keras.models import Sequential
# + id="noONFo_xW2Kb" colab_type="code" colab={}
model = Sequential()
# + id="kTL06zpRW5mk" colab_type="code" colab={}
from tensorflow.keras.layers import Conv2D
# + id="VzsfC1ZvXOiY" colab_type="code" colab={}
conv1 = Conv2D(32, 8, (4,4), activation='relu', padding='valid', input_shape=(84, 84, 1))
# + id="rJieA7BLXR7j" colab_type="code" colab={}
conv2 = Conv2D(64, 4, (2,2), activation='relu', padding='valid')
# + id="IwMsEZ8JXUHl" colab_type="code" colab={}
conv3 = Conv2D(64, 3, (1,1), activation='relu', padding='valid')
# + id="qOlyRHhDXMuO" colab_type="code" colab={}
model.add(conv1)
model.add(conv2)
model.add(conv3)
# + id="7YCm9nCiW7eg" colab_type="code" colab={}
from tensorflow.keras.layers import Flatten
# + id="AxkT-keGW_YB" colab_type="code" colab={}
model.add(Flatten())
# + id="ndOHDu7ZW_fk" colab_type="code" colab={}
from tensorflow.keras.layers import Dense
# + id="CW42s7gJXebS" colab_type="code" colab={}
fc1 = Dense(256, activation='relu')
# + id="EAAfBLWQXhmx" colab_type="code" colab={}
fc2 = Dense(4)
# + id="qj6Y03J7W0dK" colab_type="code" colab={}
model.add(fc1)
model.add(fc2)
# + id="5DgPiGANW0jz" colab_type="code" colab={}
from tensorflow.keras.optimizers import RMSprop
# + id="JoPKdR0CWr40" colab_type="code" colab={}
optimizer=RMSprop(lr=0.00025)
# + id="YFrcdqRfWr7V" colab_type="code" colab={}
model.compile(loss='mse', optimizer=optimizer, metrics=['accuracy'])
# + id="lEO1N-x2XIb3" colab_type="code" outputId="3d70bb34-0b98-43ca-8b7e-59c2e628d7f9" executionInfo={"status": "ok", "timestamp": 1588506589068, "user_tz": -600, "elapsed": 946, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiYepcJQjaVrQ1i01LEROgsJ738vi03JrR51Ryb3w=s64", "userId": "11809607246124237079"}} colab={"base_uri": "https://localhost:8080/", "height": 357}
model.summary()
| Exercise02/Exercise02.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] nbpresent={"id": "aab7fd3c-11a1-4073-a957-c10735d7afae"} slideshow={"slide_type": "slide"}
# # COMP 562 – Lecture 12
# $$
# \newcommand{\xx}{\mathbf{x}}
# \newcommand{\yy}{\mathbf{y}}
# \newcommand{\zz}{\mathbf{z}}
# \newcommand{\vv}{\mathbf{v}}
# \newcommand{\bbeta}{\boldsymbol{\mathbf{\beta}}}
# \newcommand{\mmu}{\boldsymbol{\mathbf{\mu}}}
# \newcommand{\ssigma}{\boldsymbol{\mathbf{\sigma}}}
# \newcommand{\reals}{\mathbb{R}}
# \newcommand{\loglik}{\mathcal{LL}}
# \newcommand{\penloglik}{\mathcal{PLL}}
# \newcommand{\likelihood}{\mathcal{L}}
# \newcommand{\Data}{\textrm{Data}}
# \newcommand{\given}{ \big| }
# \newcommand{\MLE}{\textrm{MLE}}
# \newcommand{\EE}{\mathbb{E}}
# \newcommand{\KL}{\textrm{KL}}
# \newcommand{\Bound}{\mathcal{B}}
# \newcommand{\tth}{\textrm{th}}
# \newcommand{\Gaussian}[2]{\mathcal{N}\left(#1,#2\right)}
# \newcommand{\norm}[1]{\left\lVert#1\right\rVert}
# \newcommand{\ones}{\mathbf{1}}
# \newcommand{\diag}[1]{\textrm{diag}\left( #1 \right)}
# \newcommand{\sigmoid}[1]{\sigma\left(#1\right)}
# \newcommand{\myexp}[1]{\exp\left\{#1\right\}}
# \newcommand{\mylog}[1]{\log\left\{#1\right\}}
# \newcommand{\argmax}{\mathop{\textrm{argmax}}}
# \newcommand{\new}{\textrm{new}}
# \newcommand{\old}{\textrm{old}}
# $$
# + [markdown] nbpresent={"id": "7d3a96df-84a2-481a-a4e4-49ac4837dfbf"} slideshow={"slide_type": "slide"}
# # EM Algorithm for Mixture of Gaussians without Covariance
#
# The model
# $$
# \begin{aligned}
# p(h\mid \alpha) &= \alpha_h \\
# p(\xx \mid h,\mu) &= (2\pi)^{-\frac{d}{2}} \myexp{-\frac{1}{2}(\xx - \mu_{h_t})^T(\xx - \mu_{h_t})} \\
# \end{aligned}
# $$
# is a variant of **Mixture of Gaussians**
#
# $\alpha_c$ is an a-priori probability that a sample comes from class $c$ -- also called **mixing proportion**
#
# The bound
# $$
# \begin{aligned}
# \Bound(\Theta,q) &= \sum_{i=1}^N \sum_{h_i} q_i(h_i) \log \frac{ p(\xx_i,h_i\mid \Theta) }{ q_i(h_i) } \\
# &= \sum_{i=1}^T \sum_{h_i} q_i(h_i) \left[ \log \alpha_{h_i} -\frac{d}{2} \log (2\pi) -\frac{1}{2}(\xx - \mu_{h_i})^T(\xx - \mu_{h_i}) \right] \\
# &- \sum_{i=1}^T \sum_{h_i} q_i(h_i) \log q_i(h_i)
# \end{aligned}
# $$
# In this case $\Theta = (\alpha_1,...,\alpha_K,\mu_1, ...,\mu_K)$
# + [markdown] slideshow={"slide_type": "slide"}
#
# # Mixture of Gaussians without Covariance -- E-step
#
# In E-step we optimize $q$s given $\Theta$
#
# $$
# \begin{aligned}
# q^{\new}_i &= \argmax_{q_i} \mathcal{B}(\Theta^{\old},q) \\
# \end{aligned}
# $$
#
# In general, we can take derivatives, equate them to zero, and solve:
#
# $$
# \nabla_{q_i} \mathcal{B}(\Theta^{\old},q) = 0
# $$
#
# We can show that in our case, the E-step updates are:
#
# $$
# \begin{aligned}
# q_i(h_i = k) &= p(h_i =k \mid \xx_i, \mu) = \frac{p(\xx_i,h_i = k \mid \mu)}{\underbrace{\sum_c p(\xx_i,h_i=c \mid \mu)}_{\textrm{same for all values of } k}}\\
# &\propto p(\xx_i,h_i = k\mid \mu) = p(h_i = k \mid \alpha) p(\xx \mid h_i=k,\mu) \\
# &= \alpha_{h_i} (2\pi)^{-\frac{d}{2}} \myexp{-\frac{1}{2}(\xx - \mu_h)^T(\xx - \mu_h)}
# \end{aligned}
# $$
# + [markdown] slideshow={"slide_type": "slide"}
# # Mixture of Gaussians without Covariance -- M-step
#
# In M-step we optimize $\Theta$ given $q$s
# $$
# \Theta^{\new} = \argmax_{\Theta} \mathcal{B}(\Theta,q^{\new})
# $$
#
# In general, we can take derivatives, equate them to zero, and solve:
# $$
# \nabla_{\Theta} \mathcal{B}(\Theta,q^{\new}) = 0
# $$
#
# We can show that in our case, the M-step updates are:
#
# $$
# \begin{aligned}
# \mu_k^* &= \frac{\sum_i q_i(h_i = k) \xx_i}{\sum_i q_i(h_i = k)} \\
# \alpha^*_k &= \frac{\sum_i q_i(h_i = k)}{N}
# \end{aligned}
# $$
# + [markdown] slideshow={"slide_type": "slide"}
# # EM Algorithm for Mixture of Gaussians with Covariance
#
# The model
#
# $$
# \begin{aligned}
# p(h\mid \alpha) &= \alpha_h \\
# p(\xx \mid h,\mu) &= (2\pi)^{-\frac{d}{2}}\color{red}{|\Sigma_h|^{-\frac{1}{2}}} \myexp{-\frac{1}{2}(\xx - \mu_{h_i})^T\color{red}{\Sigma_h^{-1}}(\xx - \mu_{h_i})} \\
# \end{aligned}
# $$
#
# This is also a variant of **Mixture of Gaussians** Note that we introduced a covariance matrix per cluster
#
# * Hidden variables: $h_i$ -- cluster membership for sample $i$
# * Parameters: $\Theta = (\underbrace{\alpha_1,...,\alpha_K}_{\textrm{proportions}},\underbrace{\mu_1, ...,\mu_K}_{\textrm{means}},\underbrace{\Sigma_1,...,\Sigma_K}_{\textrm{covariances}})$
# + [markdown] slideshow={"slide_type": "slide"}
# # EM Algorithm for Mixture of Gaussians with Covariance
#
# We plug-in probabilities $p(\xx_i\mid h_i,\Theta)$ and $p(h_i\mid \alpha)$ in the bound
#
# $$
# \begin{aligned}
# \Bound(\Theta,q) &&= \sum_{i=1}^N \sum_{h_i} q_i(h_i)&\log \frac{ p(\xx_i,h_i\mid \Theta) }{ q_i(h_i) } \\
# &&= \sum_{i=1}^N \sum_{h_i} q_i(h_i) &\left[ \log \alpha_{h_i} -\frac{d}{2} \log (2\pi)-\color{red}{\frac{1}{2}\log|\Sigma_{h_i}|}\right. \\
# &&& \left. -\frac{1}{2}(\xx_i - \mu_{h_i})^T\color{red}{\Sigma_{h_i}^{-1}}(\xx_i - \mu_{h_i}) \right] \\
# &&- \sum_{i=1}^N \sum_{h_i} q_i(h_i)& \log q_i(h_i)
# \end{aligned}
# $$
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Mixture of Gaussians with Covariance -- E-step
#
# $$
# \begin{aligned}
# q_i(h_i = k) &= p(h_i =k \mid \xx_i, \mu) = \frac{p(\xx_i,h_i = k \mid \mu)}{\underbrace{\sum_c p(\xx_i,h_i=c \mid \mu)}_{\textrm{same for all values of } k}}\\
# &\propto p(\xx_i,h_i = k\mid \mu) = p(h_i = k \mid \alpha) p(\xx \mid h_i=k,\mu) \\\\
# &= \alpha_{h_i} (2\pi)^{-\frac{d}{2}}\color{red}{|\Sigma_{h_i}^{-1}|} \myexp{-\frac{1}{2}(\xx - \mu_{h_i})^T\color{red}{\Sigma_{h_i}^{-1}}(\xx - \mu_{h_i})}
# \end{aligned}
# $$
# Implementation:
# ```
# q = numpy.zeros((K,N)) # clusters x samples
# q = logjointp(x,Theta) # compute all joints at once
# loglik = numpy.sum(logsumexp(q)) # compute loglikelihood
# q = q - logsumexp(q) # normalizing across clusters
# ```
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Mixture of Gaussians with Covariance -- M-step
#
# Updates for parameters of prior probability $p(h \mid \alpha)$
# $$
# \alpha^*_k = \frac{\sum_i q_i(h_i = k)}{N}
# $$
# Updates for means of clusters
# $$
# \mu_k^* = \frac{\sum_i q_i(h_i = k) \xx_i}{\sum_i q_i(h_i = k)}
# $$
# Updates for covariances of clusters
# $$
# \Sigma_k^* = \frac{\sum_i q_i(h_i = k) (\xx_i - \mu^*_k)(\xx_i-\mu^*_k)^T}{\sum_i q_i(h_i = k)}
# $$
# + [markdown] slideshow={"slide_type": "slide"}
# # Debugging EM Algorithm
#
# 1. Log-likelihood should always go up!
# 2. Synthetic data is your friend, if you generate data from your model you get samples and cluster membership
# 3. E-step computes cluster membership based on parameters. Use this!
# * Synthesize data from ground truth parameters
# * Start your EM from ground truth parameters, not random initialization
# * Does your E step associate samples with correct clusters?
# * Select one sample and look at its posterior probability for the cluster it came from
# 4. M-step updates parameters based on cluster membership. Use this!
# * Using synthetic data, set ```q``` to be one-hot according to ground truth
# * Start your M-step with this ```q```
# * If you don't get parameters back that are close to the ground truth
# * To isolate a broken update, let M-step update just one parameter (for example mus)
# 5. Starting your EM with ground truth parameters should not budge too much
#
# Between these tricks you should be able to isolate source of your problem
| CourseMaterial/COMP562_Lect12/12.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="P1Lzs2Icj4K5"
# !pip install fastai --upgrade
# + [markdown] id="aAa84Er1kS3X"
# # Stochastic Gradient Descent
# + [markdown] id="IqoS9BFVr40b"
# Stochastic Gradient Descent (SGD) is an optimisation algorithm which can be used to minimise a loss function in order to fit a model.
# This notebook illustrates the process of SGD by using it to train an image classifier from scratch. This example is taken from the [fast.ai course](https://course.fast.ai/).
#
# The MNIST dataset is used which contains images of handwritten numbers from 0 to 9. The problem is to create a model that can correctly predict which digit a given image is. For this example the dataset is restricted to the digits **3** and **7**.
#
#
# + id="5hjFItdMr5Z0"
from fastai.vision.all import *
import seaborn as sns
import pandas as pd
# + [markdown] id="8s5KPD5aj0Ma"
# ## Downloading and preparing the dataset
# The dataset contains 'train' and 'valid' folders each of which contain a folder for each of the two classes: **3** and **7**. Create a list of tensors from the MNIST data. We have 6,131 **3s**, and 6,265 **7s**.
# + id="4kQUtsXgkRuA"
path = untar_data(URLs.MNIST_SAMPLE)
Path.BASE_PATH = path
# + id="k-O6Ga0mlAPL" outputId="6505613b-e2bb-46f9-9811-8b16aecd6c16" colab={"base_uri": "https://localhost:8080/"}
threes = (path/'train'/'3').ls().sorted()
sevens = (path/'train'/'7').ls().sorted()
three_tensors = [tensor(Image.open(o)) for o in threes]
seven_tensors = [tensor(Image.open(o)) for o in sevens]
len(three_tensors),len(seven_tensors)
# + [markdown] id="olHTeLUekVM4"
# Combine all the images of each class into a single three-dimensional tensor and convert the pixel values to floats between 0 and 1. Looking at the shape of the tensor, we have 6,131 images of 3s that are 28 by 28 pixels.
# + id="UoXbT070n0D5" outputId="58db7f1a-d2a7-41be-d480-e18f0f7b77a5" colab={"base_uri": "https://localhost:8080/"}
stacked_sevens = torch.stack(seven_tensors).float()/255
stacked_threes = torch.stack(three_tensors).float()/255
stacked_threes.shape
# + [markdown] id="JDqrphLCmYhJ"
# Concatenate the separate class tensors into a single tensor ```train_x``` to hold all the images for training and create the tensor ```train_y``` to hold the training labels or targets (1 for **3s** and 0 for **7s**). The training images and labels are zipped together in ```dset``` because a PyTorch ```Dataset``` is required to return a tuple when indexed.
# ***Note***: the images have been changed from a 28 by 28 matrix to a 28*28 vector (784 column-long row) where each column represents a pixel value. This is important as it enables us to use matrix multiplication between the input data (pixel values) and parameters to produce an output.
# + id="wEUZhr20rK-k" outputId="798d429a-3d55-42b4-ea1b-2d57dad5a486" colab={"base_uri": "https://localhost:8080/"}
train_x = torch.cat([stacked_threes, stacked_sevens]).view(-1, 28*28)
train_y = tensor([1]*len(threes) + [0]*len(sevens)).unsqueeze(1)
dset = list(zip(train_x,train_y))
dset[0][0].shape, dset[0][1].shape
# + [markdown] id="-bYF1p2yqqqP"
# Create the validation dataset in the same manner
# + id="JTlSr_zItv-e"
valid_3_tens = torch.stack([tensor(Image.open(o))
for o in (path/'valid'/'3').ls()])
valid_3_tens = valid_3_tens.float()/255
valid_7_tens = torch.stack([tensor(Image.open(o))
for o in (path/'valid'/'7').ls()])
valid_7_tens = valid_7_tens.float()/255
valid_x = torch.cat([valid_3_tens, valid_7_tens]).view(-1, 28*28)
valid_y = tensor([1]*len(valid_3_tens) + [0]*len(valid_7_tens)).unsqueeze(1)
valid_dset = list(zip(valid_x,valid_y))
# + [markdown] id="Q0NoUCM-q7fM"
# We now have two datasets: ```dset``` which contains all the training images and their associated labels, and ```valid_dset``` which contains all the validation images and their labels. The datasets are a list of tuple pairs where the first item is an image represented as a vector of magnitude 784 and the second item is a label, either 0 or 1.
# + [markdown] id="Wlbf9jkitYH3"
# ## Building the architecture
# The first step is to initialize the parameters. These are the weights or coefficients which will be applied to the data to make class predictions. It is common to initialize the parameters randomly. Create ```weights```, a 784 row vector corresponding to the pixel length of the images, each row has a value randomly initialized using a normal random number distribution which will be applied to the pixel values to predict the class value. ```bias``` is a randomly initialised scalar variable which increases flexibility by allowing the output of the linear equations to be non-zero when the input values are 0.
# + id="W-dCDpyp5g1s" outputId="0acf2fb2-cf8f-49f4-953b-6e5eb4ea4443" colab={"base_uri": "https://localhost:8080/"}
def init_params(size, std=1.0): return (torch.randn(size)*std).requires_grad_()
weights = init_params((28*28,1))
bias = init_params(1)
weights.shape, bias.shape
# + [markdown] id="Vj7zfve6Dreb"
# The graph below shows that each of the 784 weights has been given a random value from a standard normal distribution.
# + id="6jiW-rRkWEC_" outputId="a1975f09-9edb-4859-fe68-aab6b8ea5760" colab={"base_uri": "https://localhost:8080/", "height": 390}
wdf = pd.DataFrame(weights)
wdf.rename(columns={0 :'weight value'}, inplace=True )
g = sns.catplot(x=wdf.index, y='weight value', data=wdf, kind='bar')
(g.set_axis_labels("Weights", "Weight Value")
.set_xticklabels([]))
# + [markdown] id="4ZstFol0y1xX"
# ### Define the model
# Create a function that takes the image vectors as input and multiplies them by the parameters to produce an output which will be used to predict the class value. This simple function is actually the model and since the parameters are randomly intitialised its output won't actually have anything to do with what we want. However, once we compare the outputs with our targets and use some *loss function* to evaluate its performance, we can use SGD to minimise the loss and improve the model.
# + id="siuBZIEG5zzZ" outputId="deb22d73-6d0e-4255-a420-cd9aad50dc14" colab={"base_uri": "https://localhost:8080/"}
def linear1(xb): return xb@weights + bias
preds = linear1(train_x)
preds
# + [markdown] id="ginrffc0zjpX"
# Next we set an arbitrary threshold to predict whether a given image is a **3** or a **7**.
# + id="WL9Y6XYz54hf" outputId="fb176570-4e86-4a4e-b1e2-b961968ad6f8" colab={"base_uri": "https://localhost:8080/"}
thresh = 0.0
corrects = (preds>thresh).float() == train_y
corrects.float().mean().item()
# + [markdown] id="w0ETktln0Ehq"
# The graph below shows the distribution of the prediction values for each class value. We can see that our arbitrary threshold of 0 in fact does a very poor job of predicting the class values because there is a healthy mix of both **3**s and **7**s on either end of 0. These two lovely shaped distributions are to be expected from intialising the weights randomly using the standard normal distribution.
# + id="3jpFUK8Oz6zp" outputId="a3ee6131-194a-494b-9cd3-46d761049f83" colab={"base_uri": "https://localhost:8080/", "height": 296}
df = pd.DataFrame(preds)
df['threes'] = df[:6131]
df['sevens'] = df[0][6131:]
sns.histplot(df['threes'], color="skyblue")
sns.histplot(df['sevens'], color="lightsalmon")
# + [markdown] id="l0SsU-M_90pA"
# ### Define a *loss function*
# The loss function is a means of assessing the the current weight assignments in terms of actual performance.
#
# This loss function takes the outputs from the ```linear1``` model as inputs and compares them to the targets. It measures the distance of each prediction from its target label (i.e. 1 for **3**s and 0 for **7**s) for a given batch containing prediction values and target labels and returns an average of these distances. It is our goal to minimise this function by changing the values of the weights. A score of 0 would mean that for some combination of weights applied to a batch of inputs, the model produced outputs that corresponded exactly to the batch's target labels.
#
# One problem with ```mnist_loss``` as currently defined is that it assumes that prediction values are always between 0 and 1 (the above graph shows they are actually ranging from around -20 to 20). The sigmoid method in the loss function transforms the prediction values into values that fall in the domain of 0 to 1 to allow our loss function to work.
#
# 
#
#
#
# + id="ti829BzB7Do-"
def mnist_loss(predictions, targets):
predictions = predictions.sigmoid()
return torch.where(targets==1, 1-predictions, predictions).mean()
# + [markdown] id="JLA_qqcwDLdB"
# Creater the ```dl``` ```DataLoader``` which takes our dataset and turns it into an iterator over many batches. The batchs are tuples of tensors representing batches of images and targets. Here we specify a batch size of 256. Processing the dataset in batches like this is the difference between *Gradient Descent* and *Stochastic Gradient Descent*. *Gradient Descent* requires much more computational resources as the entire dataset needs to be processed for one epoch.
# + id="wjBsuxZr_iPm"
dl = DataLoader(dset, batch_size=256)
valid_dl = DataLoader(valid_dset, batch_size=256)
# + [markdown] id="o6KO5crZLedK"
# ### Optimisation
# Now that we have a loss function to measure the performance of our model, we need a method that updates the paramaters (weights and bias) to minimise the loss function and improve the model's performance, this is often referred to as stepping. The goal is to adjust the weights until we find the bottom of the loss function and an efficient method for this is to use the gradient of the loss function to tell us which direction to adjust the weights.
#
# 
#
# + [markdown] id="3kCODiN6SMy4"
# ```calc_grad``` will process our batches through our model and assess performance using the loss function while tracking the gradient for each weight.
# + id="eOY8272QDEV-"
def calc_grad(x_batch, y_batch, model):
preds = model(x_batch)
loss = mnist_loss(preds, y_batch)
loss.backward()
# + [markdown] id="5v-j7w3QTvjw"
# Now all that is left is to update the parameters based on the gradients. It is common to multiply the gradient by a small number called the learning rate to control the size of gradient steps. If it is too large, the algorithm could step beyond the minimum and jump around the function a lot. If it is too small, it will take longer to reach the minimum. Often times the learning rate is set through some trial and error.
#
# The full process is represented in the flow chart below and shows how it is an iterative process.
# We initialise the weights and measure the predictions from a model using a loss function, record the gradients and step the parameters toward the bottom of the loss function. We repeat the process with the stepped parameters and continue iterating to improve the predictions until we decide to stop the process.
#
# 
#
# + [markdown] id="wJWhjYYIYF5Q"
# The ```train_epoch``` function includes a loop to upgrade the parameters based on the gradients and a learning rate: ```lr```.
# + id="A5uNfdWLDhiz"
def train_epoch(model, lr, params):
for x_batch, y_batch in dl:
calc_grad(x_batch, y_batch, model)
for p in params:
p.data -= p.grad*lr
p.grad.zero_()
# + [markdown] id="oUZAbMVjZEaL"
# We also want to check how we're doing, by looking at the accuracy of the validation set. To decide if an output represents a **3** or a **7**, we can just check whether it's greater than 0. The ```batch_accuracy``` function does this and returns an accuracy score. ```validate_epoch``` then puts all the batches together and returns an overall score for each epoch.
# + id="Qt1cWbNgY7nA"
def batch_accuracy(x_batch, y_batch):
preds = x_batch.sigmoid()
correct = (preds>0.5) == y_batch
return correct.float().mean()
def validate_epoch(model):
accs = [batch_accuracy(model(xb), yb) for xb,yb in valid_dl]
return round(torch.stack(accs).mean().item(), 4)
# + [markdown] id="l8JXh9O8Z5Kg"
# Now all thats left is to train the model. First we'll run three epochs and see how the accuracy improves and look at how the distribution of class values (targets) changes. The accuracy improves rapidly in just three epochs and we receive accuracy scores of 52.28%, 65.23% and 85%.
# + id="7dCh1NERD0c8" outputId="21261314-4899-4a65-fb9c-0dcbb2263c7d" colab={"base_uri": "https://localhost:8080/"}
lr = 1.
params = weights, bias
for i in range(3):
train_epoch(linear1, lr, params)
print(validate_epoch(linear1), end=' ')
# + [markdown] id="jYrH_qjebVa9"
# The graph below shows the distributions of the predictions of the class values being pulled away from each other.
# + id="RU_SuFijYLMo" outputId="3f6bca69-82d1-43c6-e478-5c184d78755b" colab={"base_uri": "https://localhost:8080/", "height": 299}
preds = linear1(train_x)
df = pd.DataFrame(preds)
df['threes'] = df[:6131]
df['sevens'] = df[0][6131:]
sns.histplot(df['threes'], color="skyblue")
sns.histplot(df['sevens'], color="lightsalmon")
# + [markdown] id="yYhdqPt2b75Y"
# After 15 more epochs the accuracy improves to over 97%
# + id="cS9xkJ9-GpDu" outputId="2b51f3df-21f5-4dd2-b257-d99c7a9ce345" colab={"base_uri": "https://localhost:8080/"}
for i in range(15):
train_epoch(linear1, lr, params)
print(validate_epoch(linear1), end=' ')
# + [markdown] id="dW_F5dfjdaTF"
# We can also see the how the current combination of weights produces predictions which divide the class values into two distinct groups compared to the starting randomly initialised weights. The predictions of images of **7**s which intially had a greater mean than the predictions of **3**s has been pulled to the left into larger negative values. This is because large negative values are converted to 0 or very close to 0 by the sigmoid function. Since we set the target for **7**s to 0, the weights have been optimised to the point where the model will produce large negative prediction values for images of **7**s which corresponds to its target label of 0. Likewise the model will produce large positive prediction values for **3**s which corresponds to its target label of 1 once the sigmoid function is applied.
#
#
# + id="lzAZlny7INLm" outputId="964c5866-a3fd-450f-e3a3-03cd0b68e18c" colab={"base_uri": "https://localhost:8080/", "height": 297}
preds = linear1(train_x)
df = pd.DataFrame(preds)
df['threes'] = df[:6131]
df['sevens'] = df[0][6131:]
sns.histplot(df['threes'], color="skyblue")
sns.histplot(df['sevens'], color="lightsalmon")
# + [markdown] id="9u1Sdrule6_u"
# ### Distribution of predictions using randomly initialised weights
# 
| _notebooks/2020-11-15-SGD.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Environment (conda_tensorflow_p36)
# language: python
# name: conda_tensorflow_p36
# ---
# # 使用自定义Image进行推理
# 需要用到Tensorflow 和Keras , kernel 选择conda_tensorflow_p36
#
# ## 把训练好的模型存放到指定路径下
# `output/model/tf_server/`
#
# 目录结构如下
# ```
# └── tf_server
# └── 1
# ├── saved_model.pb
# └── variables
# ├── variables.data-00000-of-00001
# └── variables.index
# ```
# ## Build容器
# 对外提供服务时,需要使用容器方式。
# 图片分类本身可不使用自定义容器,这里只是演示如何自定义容器。
# + language="sh"
# docker build -t sagemaker-cat-vs-dog .
# -
# ## 本地运行docker
# + language="sh"
# docker run -p 8501:8501 -d sagemaker-cat-vs-dog
# -
# ## 用本地Docker测试
# +
import requests
import numpy as np
import json
import boto3
IMAGE_WIDTH = 150
IMAGE_HEIGHT = 150
# 修改测试图片地址
image_paths = 'test/cat.681.jpg'
#image_paths = 'test/dog.592.jpg'
model_server_url = 'http://127.0.0.1:8501/v1/models/sagemaker-demo:predict'
# +
from keras.preprocessing import image
images = image.load_img(image_paths, target_size=(IMAGE_WIDTH, IMAGE_HEIGHT))
input_image = image.img_to_array(images)
input_image = np.expand_dims(input_image, axis=0)
input_image /= 255.
input_images = input_image.tolist()
data = json.dumps({"inputs":input_images})
headers = {"content-type": "application/json"}
json_response = requests.post(model_server_url, data=data, headers=headers)
json_result = json.loads(json_response.text)
print(json_result)
# -
# 根据需要修改class name,按字典序
class_name=['cat','dog']
class_name[np.argmax(json_result["outputs"])]
# ## 推送docker镜像到ECR
# + language="sh"
# algorithm_name=sagemaker-cat-vs-dog
# REGION=$(aws configure get region)
# ACCOUNT=$(aws sts get-caller-identity --query Account --output text)
#
# # If the ECS repository doesn't exist, creates it.
# aws ecr create-repository --repository-name ${algorithm_name} > /dev/null 2>&1
#
# # ECR requires the image name to be in this format:
# REPOSITORY_NAME=${ACCOUNT}.dkr.ecr.${REGION}.amazonaws.com.cn/${algorithm_name}:latest
#
# # Tags the image with the expect ECR format
# docker tag ${algorithm_name} ${REPOSITORY_NAME}
#
# # Allows docker access to ECR
# $(aws ecr get-login --no-include-email)
#
# # pushes the image to ECR
# docker push ${REPOSITORY_NAME}
# -
# ## 部署模型到SageMaker
# +
from sagemaker.model import Model
import sagemaker
from sagemaker import get_execution_role
#role = get_execution_role()
role="arn:aws-cn:iam::315505707008:role/service-role/AmazonSageMaker-ExecutionRole-20200430T124235"
image_uri = "315505707008.dkr.ecr.cn-northwest-1.amazonaws.com.cn/sagemaker-cat-vs-dog"
endpoint_name = "sagemaker-cat-vs-dog"
my_model = Model(
role=role,
image_uri=image_uri)
#该步骤大概需要10分钟
xgb_predictor = my_model.deploy(initial_instance_count=1,
endpoint_name=endpoint_name,
instance_type='ml.t2.medium'
)
# -
# ## 推理
# ### 读取数据
# +
from keras.preprocessing import image
import json
import numpy as np
IMAGE_WIDTH = 150
IMAGE_HEIGHT = 150
# 修改测试图片地址
image_paths = 'test/cat.681.jpg'
#image_paths = 'test/dog.592.jpg'
images = image.load_img(image_paths, target_size=(IMAGE_WIDTH, IMAGE_HEIGHT))
input_image = image.img_to_array(images)
input_image = np.expand_dims(input_image, axis=0)
input_image /= 255.
input_images = input_image.tolist()
data = json.dumps({"name": 'tensorflow/serving/predict',"signature_name":'predict',"inputs":input_images})
# -
input_image.shape
# ### 方式一、使用boto3
# +
import boto3
endpoint_name = "sagemaker-cat-vs-dog"
client = boto3.client('runtime.sagemaker')
response = client.invoke_endpoint(EndpointName=endpoint_name,
Body=data)
print(response)
# -
response_body = response['Body']
body= response_body.read()
results = json.loads(body.decode('utf-8'))
print(results)
# ### 方式二、使用SageMaker SDK
import sagemaker
xgb_predictor = sagemaker.predictor.Predictor(
endpoint_name=endpoint_name,
sagemaker_session=sagemaker.Session())
result = xgb_predictor.predict(data)
results = json.loads(result.decode('utf-8'))
print(results)
| sagemaker/encapsulation/inference-custom-image.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=true editable=true
# Ensure that the path to `opengrid` is in your PYTHONPATH. If you installed opengrid with `pip install opengrid`, this is done automatically.
# + deletable=true editable=true
from opengrid_dev import config
# + deletable=true editable=true
c = config.Config()
for s in c.sections():
print("Section: {}".format(s))
for i in c.items(s):
print("\tItem: {}".format(i))
# + deletable=true editable=true
c.get('opengrid_server', 'host')
# + deletable=true editable=true
c.get('data', 'folder')
# + deletable=true editable=true
| opengrid_dev/notebooks/Demo/Demo_Config.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Data Analysis Tools
# # Assignment: Running an analysis of variance
#
# Following is the Python program I wrote to fulfill the first assignment of the [Data Analysis Tools online course](https://www.coursera.org/learn/data-analysis-tools/home/welcome).
#
# I decided to use [Jupyter Notebook](http://nbviewer.jupyter.org/github/ipython/ipython/blob/3.x/examples/Notebook/Index.ipynb) as it is a pretty way to write code and present results.
#
# ## Research question
#
# Using the [Gapminder database](http://www.gapminder.org/), I would like to see if an increasing Internet usage results in an increasing suicide rate. A study shows that other factors like unemployment could have a great impact.
#
# So for this assignment, the three following variables will be analyzed:
#
# - Internet Usage Rate (per 100 people)
# - Suicide Rate (per 100 000 people)
# - Unemployment Rate (% of the population of age 15+)
#
#
# ## Data management
#
# For the question I'm interested in, the countries for which data are missing will be discarded. As missing data in Gapminder database are replace directly by `NaN` no special data treatment is needed.
# + hide_input=false
# Magic command to insert the graph directly in the notebook
# %matplotlib inline
# Load a useful Python libraries for handling data
import pandas as pd
import numpy as np
import statsmodels.formula.api as smf
import seaborn as sns
import matplotlib.pyplot as plt
from IPython.display import Markdown, display
# -
# Read the data
data_filename = r'gapminder.csv'
data = pd.read_csv(data_filename, low_memory=False)
data = data.set_index('country')
# General information on the Gapminder data
# + variables={"len(data)": "<p><strong>NameError</strong>: name 'data' is not defined</p>\n", "len(data.columns)": "<p><strong>NameError</strong>: name 'data' is not defined</p>\n"}
display(Markdown("Number of countries: {}".format(len(data))))
display(Markdown("Number of variables: {}".format(len(data.columns))))
# -
# Convert interesting variables in numeric format
for variable in ('internetuserate', 'suicideper100th', 'employrate'):
data[variable] = pd.to_numeric(data[variable], errors='coerce')
#
# But the unemployment rate is not provided directly. In the database, the employment rate (% of the popluation) is available. So the unemployement rate will be computed as `100 - employment rate`:
data['unemployrate'] = 100. - data['employrate']
# The first records of the data restricted to the three analyzed variables are:
subdata = data[['internetuserate', 'suicideper100th', 'unemployrate']]
subdata.head(10)
# ## Data analysis
#
# The distribution of the three variables have been analyzed [previously](Visualizing_Data.ipynb).
#
#
# ## Variance analysis
#
# Now that the univariate distribution as be plotted and described, the bivariate graphics will be plotted in order to test our research hypothesis.
#
# Let's first focus on the primary research question;
#
# - The explanatory variable is the internet use rate (quantitative variable)
# - The response variable is the suicide per 100,000 people (quantitative variable)
#
# From the scatter plot, a slope slightly positive has been seen. And as most of the countries have no or very low internet use rate, an effect is maybe seen only on the countries having the higher internet use rate.
subdata2 = subdata.assign(internet_grp4 = pd.qcut(subdata.internetuserate, 4,
labels=["1=25th%tile", "2=50th%tile",
"3=75th%tile", "4=100th%tile"]))
sns.factorplot(x='internet_grp4', y='suicideper100th', data=subdata2,
kind="bar", ci=None)
plt.xlabel('Internet use rate (%)')
plt.ylabel('Suicide per 100 000 people (-)')
_ = plt.title('Average suicide per 100,000 people per internet use rate quartile')
# This case falls under the Categorical to Quantitative case of interest for this assignement. So ANOVA analysis can be performed here.
#
#
# - The null hypothesis is: There is no relationship between the Internet use rate and suicide
# - The alternate hypothesis is: There is a relationship between the Internet use rate and suicide
model1 = smf.ols(formula='suicideper100th ~ C(internet_grp4)', data=subdata2).fit()
model1.summary()
# The p-value found is 0.143 > 0.05. Therefore the null hypothesis cannot be rejected. There is no relationship between the internet use rate and the suicide rate.
# ## Test case to fulfill the assignment
#
# In order to fulfill the assignment, I will switch to the NESARC database.
nesarc = pd.read_csv('nesarc_pds.csv', low_memory=False)
races = {1 : 'White',
2 : 'Black',
3 : 'American India/Alaska',
4 : 'Asian/Native Hawaiian/Pacific',
5 : 'Hispanic or Latino'}
subnesarc = (nesarc[['S3BQ4', 'ETHRACE2A']]
.assign(ethnicity=lambda x: pd.Categorical(x['ETHRACE2A'].map(races)),
nb_joints_day=lambda x: (pd.to_numeric(x['S3BQ4'], errors='coerce')
.replace(99, np.nan)))
.dropna())
g = sns.factorplot(x='ethnicity', y='nb_joints_day', data=subnesarc,
kind="bar", ci=None)
g.set_xticklabels(rotation=90)
plt.ylabel('Number of cannabis joints per day')
_ = plt.title('Average number of cannabis joints smoked per day depending on the ethnicity')
# The null hypothesis is *There is no relationship between the number of joints smoked per day and the ethnicity*.
#
# The alternate hypothesis is *There is a relationship between the number of joints smoked per day and the ethnicity*.
model2 = smf.ols(formula='nb_joints_day ~ C(ethnicity)', data=subnesarc).fit()
model2.summary()
# The p-value is much smaller than 5%. Therefore the null hypothesis is rejected. We can now look at which group are really different from the other.
# +
import statsmodels.stats.multicomp as multi
multi1 = multi.MultiComparison(subnesarc['nb_joints_day'], subnesarc['ethnicity'])
result1 = multi1.tukeyhsd()
result1.summary()
# -
# From the Tukey's Honestly Significant Difference Test, we can conclude there are 3 relationship presenting a real difference:
#
# - American India/Alaska group smokes more than the Hispanic or Latino group
# - American India/Alaska group smokes more than the White group
# - Black group smokes more than White group
# ## Summary
#
# Using the ANOVA test on the research question *do countries with a high internet use rate have a higher number of suicides?* brought me to the conclusion that there is no relationship between the internet use rate and the number of suicide.
#
# So in order to fulfill this assignment, I switch to the NESARC database. My interest focus on a possible relationship between ethnicity and the number of cannabis joints smoked per day. After verifying that there is a significant relationship, I applied the Tukey HSD method to figure out which groups were really different one from the other.
#
# There are 3 relationship presenting a real difference:
#
# - American India/Alaska group smokes more than the Hispanic or Latino group
# - American India/Alaska group smokes more than the White group
# - Black group smokes more than White group
#
#
# > If you are interested into data sciences, follow me on [Tumblr](http://fcollonval.tumblr.com/).
| Analysis_Variance.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# __Hydrograph Development Notebooks__
#
#
# __Breach Hydrographs, Deposit, NY__
#
#
# PYTHON
#
#
# Overview: This notebook was created to document the development of breach hydrographs using historical flow data for two locations along the levee at [Deposit, NY](https://www.google.com/maps/place/Deposit,+NY+13754/@42.0669205,-75.4170064,403m/data=!3m1!1e3!4m5!3m4!1s0x89db121636b095eb:0x8831b5a6c812e9f7!8m2!3d42.0600834!4d-75.4276769).
#
# Updated 1.10.2017
# +
import os
from glob import glob
from importlib import reload
import utils; reload(utils)
from utils import *
import pandas as pd
import numpy as np
from scipy import interpolate
import matplotlib.pyplot as plt
import warnings
warnings.filterwarnings("ignore")
# %matplotlib inline
# -
# ### Read in Base Hydrograph
# 1. Read in Base Hydrograph from TR-20 Hydrologic model (Unsteady HEC-RAS file)
hdf_plan_file = r'p:\02\NY\Broome_Co_36007C\LAMP2\TECH\Analysis\Modeling\WorkingModels\TOQC\Deposit\Deposit.p06.hdf'
df = GetRasUnsteadyFlow(hdf_plan_file)
df.plot()
df.max()
# # Develop a discharge hydrograph of the 1% storm for the main flooding source
#
# ## Exploratory Analysis
# [Notebook](FirstLook_GageData.ipynb) developed to evaluate available gage data in the vicinity, plot available time series & qualitatively assess differences in hydrograph shapes.
#
# ## Discharge Hydrograph
# In house, detailed hydrologic models created using the TR-20 model exist for both flooding sources. 1% Discharge hydrographs copied from model output.
#
# ## Develop of a breach hydrograph using the flow hydrograph created in step 1.
#
# In order to convert the flow hydrograph to a stage hydrograph at any given location, a hydraulic analysis is necessary to properly account for differences in the cross-sectional area at different locations along the reach. For this study a 1D, Steady State model was used to simulate a Natural Valley scenario in the levee impact area.
#
# The geometry from this model was used to compute flows ranging from 1oo cfs to 2,500 cfs in increments of 100 cfs. The results of these simulations were used to develop a rating curve at each area of interest to translate flow to stage. The image below is an example of the results at a cross section, illustrating how geometric differences at different flow levels may impact the resultant stage for a given reach.
#
# Note that the change in water surface elevation when the flow is constrained by the channel and the levee during overbank flow rises at a greater rate when compared with the unconstrained flow when conveyance occurs on both sides of the levee (natural valley).
#
# <img src="https://raw.githubusercontent.com/Dewberry-RSG/HydrologyTools/master/images/XS_Example.png" , width=1000,height=600/>
#
#
# ### Procedure to create Breach Hydrograph
#
# A. Read in HEC-RAS data for the XS of interest & create a stage/discharge rating curve using computed flows.
#
# B. Using the data from the rating curve in Part A, create a function (nth degree polynomial interpolation equation) to convert flow to stage.
#
# C. Convert the 1% flow hydrograph created in Step 1 to a stage hydrograph using the rating curve function created in Part B.
#
# D. Normalize the stage to 'feet above the breach point' using the stage hydrograph created in Part C and the breach elevation (head = 0 at breach point).
#
# E. Using the head above breach hydrograph created in Part D, calculate weir flow for (use the Standard Weir Equation, below) each timestep & write to file.
#
# F. Input weir flow hydrograph created in Part E into HEC-RAS unsteady flow file. END.
#
# #### The Standard Weir Equation:
# ## $\qquad$ $Q = CLH^{2/3}$
#
# Where:
#
# $\qquad$ __Q__ = Discharge (cfs)
# $\qquad$ __C__ = Weir coefficient (unitless)
# $\qquad$ __L__ = Weir crest length (ft)
# $\qquad$ __H__ = Energy head over the weir crest (ft)
#
#
# **From HEC-RAS Lateral Weir Coefficients, use the default Weir Coefficient of 2.0 (range is 1.5-2.6, given on page 3-50 of the [2D Users Manual](http://www.hec.usace.army.mil/software/hec-ras/documentation/HEC-RAS%205.0%202D%20Modeling%20Users%20Manual.pdf))*
# ### Breach Location # 1: Big Hollow (upstream)
# <img src="https://raw.githubusercontent.com/Dewberry-RSG/HydrologyTools/master/images/938_location_1_B_Hollow.PNG", width=900,height=800/>
#
# +
# Enter Ras Plan File
rasdata = r'p:\02\NY\Broome_Co_36007C\LAMP2\TECH\Analysis\Modeling\WorkingModels\TOQC\Deposit\Deposit.p03.hdf'
input_hydro = df['Big_Hollow']
station = str(938)
breach_point = 1
breach_height = 1008.89
data_dir = r'C:\Users\slawler\Repos\HydrologyTools\sample_data'
GetBreachFlow(input_hydro,'Deposit', rasdata, station, breach_point, breach_height, data_dir, date_int = 2)
# -
# ### Breach Location # 2: Big Hollow (downstream)
#
# <img src="https://raw.githubusercontent.com/Dewberry-RSG/HydrologyTools/master/images/1_location_2_B_Hollow.PNG", width=900,height=800/>
#
# +
input_hydro = df['Big_Hollow']
station = str(1)
breach_point = 2
breach_height = 996.19
GetBreachFlow(input_hydro, 'Deposit', rasdata, station, breach_point, breach_height, data_dir, date_int = 2)
# -
# ### Breach Location # 1: Butler Brook (upstream)
# <img src="https://raw.githubusercontent.com/Dewberry-RSG/HydrologyTools/master/images/938_location_1_B_Hollow.PNG", width=900,height=800/>
# +
input_hydro = df['Butler_Brook']
station = str(2300)
breach_point = 1
breach_height = 1007
GetBreachFlow(input_hydro, 'Deposit', rasdata, station, breach_point, breach_height, data_dir, date_int =4)
# -
# ### Breach Location # 2:
# <img src="https://raw.githubusercontent.com/Dewberry-RSG/HydrologyTools/master/images/1_location_2_B_Hollow.PNG", width=900,height=800/>
# +
input_hydro = df['Butler_Brook']
station = str(1)
breach_point = 2
breach_height = 996.42
GetBreachFlow(input_hydro, 'deposit', rasdata, station, breach_point, breach_height, data_dir, date_int = 1)
| nbs/Deposit_BreachHydro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import torch
import torch.nn as nn
import torch.optim as optim
import torchvision
import torchvision.transforms as transforms
from icap import data, models, task, utils
import pytorch_lightning as pl
from pytorch_lightning import loggers as pl_loggers
from pytorch_lightning.metrics import Accuracy
# +
torch.backends.cudnn.benchmark = True
# device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
# Hyperparameters
BATCH_SIZE = 128
NUM_WORKERS = 8
SPLIT_VAL = 0.2
EMBED_SIZE = 256
HIDDEN_SIZE = 256
NUM_LAYERS = 1
LRATE = 3e-5
MAX_EPOCHS = 100
NUM_GPU=1
# +
train_transform = transforms.Compose([
transforms.Resize((356, 356)),
transforms.RandomCrop((224, 224)),
transforms.RandomHorizontalFlip(),
transforms.ToTensor(),
transforms.Normalize((0.485, 0.456, 0.406),(0.229, 0.224, 0.225)),
])
valid_transform = transforms.Compose([
transforms.Resize((224, 224)),
transforms.ToTensor(),
transforms.Normalize((0.485, 0.456, 0.406),(0.229, 0.224, 0.225)),
])
root_dir = "data/flickr8k/images/"
caption_file = "data/flickr8k/captions.txt"
train_loader, trainset = data.flickr_dataloader(root_dir, caption_file, transform=train_transform,
num_workers=NUM_WORKERS, shuffle=True, train=True)
valid_loader, validset = data.flickr_dataloader(root_dir, caption_file, transform=valid_transform,
num_workers=NUM_WORKERS, shuffle=False, train=False)
# +
# initialize model, loss etc
PAD_INDEX = trainset.vocab.stoi["<PAD>"]
VOCAB_SIZE = len(trainset.vocab)
print(f'VOCAB_SIZE : {VOCAB_SIZE}')
model = models.ImageCaptionNet(EMBED_SIZE, HIDDEN_SIZE, VOCAB_SIZE, NUM_LAYERS)
criterion = nn.CrossEntropyLoss(ignore_index=PAD_INDEX)
optimizer = optim.Adam(model.parameters(), lr=LRATE)
trainer_task = task.ImageCaptionTask(model, optimizer, criterion, vocab_size=VOCAB_SIZE)
# +
checkpoint_path = '../saved_model/flickr'
# DEFAULTS used by the Trainer
checkpoint_callback = pl.callbacks.ModelCheckpoint(
filepath=checkpoint_path,
save_top_k=1,
verbose=True,
monitor='checkpoint_on',
mode='min',
prefix='flickr30k_net_'
)
tensorboard_logger = pl_loggers.TensorBoardLogger('../logs/test')
trainer = pl.Trainer(max_epochs=MAX_EPOCHS, gpus=NUM_GPU,
logger=tensorboard_logger, checkpoint_callback=checkpoint_callback)
# -
trainer.fit(trainer_task, train_loader, valid_loader)
# +
img, text = validset[201]
img = img.unsqueeze(0)
imtest = utils.ImageCaptionTest(model, trainset.vocab)
imtest.show_result(img, text)
# -
| notebook/train_v2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Linear Regression implemenation
# +
from statistics import mean
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import style
import random
style.use('fivethirtyeight')
def create_dataset(hm, variance, step = 2, correlation=False):
val = 1
ys = []
for i in range(hm):
y = val + random.randrange(-variance, variance)
ys.append(y)
if(correlation and correlation == 'pos'):
val += step
elif (correlation and correlation == 'neg'):
val -= step
xs = [i for i in range(len(ys))]
return np.array(xs, dtype=np.float64), np.array(ys, dtype=np.float64)
#xs = [1, 2, 3, 4, 5, 6]
#ys = [5, 4, 6, 5, 6, 7]
xs, ys = create_dataset(100, 60, 2, 'pos')
plt.scatter(xs, ys)
plt.show()
# +
# creating numpy array
xs = np.array(xs, dtype=np.float64)
ys = np.array(ys, dtype=np.float64)
def best_fit_slop_and_intercept(xs, ys):
m = ( ((mean(xs) * mean(ys)) - mean(xs*ys)) /
((mean(xs)**2) - mean(xs**2))
)
b = mean(ys) - m*mean(xs)
return m, b
m, b = best_fit_slop_and_intercept(xs, ys)
regression_line = [(m*x) + b for x in xs]
predict_x = 110
predict_y = (m*predict_x) + b
plt.scatter(xs, ys)
plt.scatter(predict_x, predict_y, color='red')
plt.plot(xs, regression_line)
plt.show()
# +
def squared_error(ys_orig, ys_line):
return sum((ys_line - ys_orig)**2)
def coefficient_of_determination(ys_orig, ys_line):
squared_error_regression = squared_error(ys_orig, ys_line);
squared_error_mean = squared_error(ys_orig, mean(ys_orig));
return 1 - (squared_error_regression/squared_error_mean)
r_squared = coefficient_of_determination(ys, regression_line)
print('r_squared = '+str(r_squared))
| linear_regression/linear_regression_from_scratch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import mysql.connector
import os
import time
# discover which candidates need data still
mydb = mysql.connector.connect(
host="localhost",
port=3308,
user="root",
password="<PASSWORD>",
database="ftvBackEnd"
)
cursor = mydb.cursor()
query = """
SELECT
cid
FROM
candidate
where
cycle = 2020
AND id not in (
SELECT candidate_id FROM `candidate-industry`
)
"""
cursor.execute(query)
still_needs_data = cursor.fetchall()
print(len(still_needs_data))
mydb.close()
url = ("https://www.opensecrets.org/api/?method=candIndustry&cid="
"{cid}"
"&cycle="
"{cycle}"
"&apikey="
"{apikey}"
"&output=json"
)
cycle=2020
apikey="wouldn't you like to know"
json_list = []
no_resource_found = []
for candidate in still_needs_data:
open_secret_request = requests.get(url.format(cid=candidate[0], cycle=cycle, apikey=apikey))
time.sleep(1)
print(open_secret_request)
if open_secret_request.ok:
json_list.append(open_secret_request.json())
elif open_secret_request.status_code == 404:
no_resource_found.append(candidate)
else:
break
# +
def insert_new_industry(code, name, cursor):
query = """
INSERT IGNORE INTO
industry(`code`, name)
VALUES
(%s, %s)
"""
cursor.execute(query, (code, name))
def populate_ftv(cid, cycle, last_updated, industry_array, cursor):
clean_cid_cycle_query = """
DELETE FROM
`candidate-industry`
WHERE
candidate_id = (SELECT id FROM candidate WHERE cid=%s AND cycle=%s);
"""
insert_funding_query = """
INSERT INTO
`candidate-industry`(candidate_id, industry_id, indivs, pacs, total, last_updated)
VALUES
(
(SELECT id FROM candidate WHERE cid=%s AND cycle=%s),
(SELECT id FROM industry WHERE `code`=%s),
%s,
%s,
%s,
%s
)
"""
cursor.execute(clean_cid_cycle_query, (cid, cycle))
if len(industry_array) == 1:
insert_new_industry(
industry_array["@attributes"]["industry_code"],
industry_array["@attributes"]["industry_name"],
cursor)
cursor.execute(insert_funding_query, (
cid,
cycle,
industry_array["@attributes"]["industry_code"],
industry_array["@attributes"]["indivs"],
industry_array["@attributes"]["pacs"],
industry_array["@attributes"]["total"],
last_updated
))
else:
for industry in industry_array:
insert_new_industry(
industry["@attributes"]["industry_code"],
industry["@attributes"]["industry_name"],
cursor)
params = (
cid,
cycle,
industry["@attributes"]["industry_code"],
industry["@attributes"]["indivs"],
industry["@attributes"]["pacs"],
industry["@attributes"]["total"],
last_updated
)
cursor.execute(insert_funding_query, params)
mydb = mysql.connector.connect(
host="localhost",
port=3308,
user="root",
password="<PASSWORD>",
database="ftvBackEnd"
)
cursor = mydb.cursor()
for candidate in json_list:
cid = candidate['response']['industries']['@attributes']['cid']
cycle = candidate['response']['industries']['@attributes']['cycle']
last_updated = candidate['response']['industries']['@attributes']['last_updated']
industries = candidate['response']['industries']['industry']
populate_ftv(cid, cycle, last_updated, industries, cursor)
mydb.commit()
mydb.close()
# -
json_list
| serverside/database/etl/scrape_opensecrets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import re
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
train = pd.read_csv('./dataset/train.csv', index_col=0)
print('Train data shape:', train.shape)
train.head()
train.info()
# +
Categorical = ['Survived', 'Sex', 'Pclass', 'SibSp', 'Parch', 'Embarked']
Real_valued = ['Age', 'Fare']
Uncat = ['Name', 'Ticket', 'Cabin']
# -
sns.heatmap(train.corr(), vmin=-1, vmax=1, )
plt.show()
# # Categorical
# ## Target
plt.figure(figsize=(8,4))
sns.barplot(data=train, x='Survived', color='skyblue')
# ## Features
fig, axs = plt.subplots(3, 2, figsize=(12,16))
for ax, cat in zip(axs.flatten(), Categorical[1:]):
sns.barplot(data=train, x=cat, y='Survived', ax=ax)
# # Real Valued
# +
fig, axs = plt.subplots(2, 1, figsize=(12,12))
for ax, var in zip(axs, Real_valued):
sns.distplot(train[train.Survived == 1][var].dropna(), color='darkgreen', ax=ax, label='Survived')
sns.distplot(train[train.Survived == 0][var].dropna(), color='darkred', ax=ax, label='Not-Survived')
ax.legend()
# -
# # Feature Engineering
# ## Passenger Name
train.Name.head()
train['Name'] = train.Name.map(lambda x:re.findall('([A-Za-z]+\.)' ,x)[0])
train.Name.head()
plt.figure(figsize=(12,6))
sns.barplot(x=train.Name, y=train.Survived)
plt.xticks(rotation='vertical')
plt.show()
title, count = np.unique(train.Name.values, return_counts=True)
pd.DataFrame({'title':title, 'count':count}).T
def group_titles(titles):
for i, each in enumerate(titles):
if any(each == ele for ele in ['Mr.', 'Miss.', 'Mrs.', 'Master.']):
continue
elif any(each == ele for ele in ['Sir.', 'Ms.', 'Mme.', 'Mlle.', 'Lady.', 'Countess.']):
titles[i] = 'grp1'
else:
titles[i] = 'grp2'
group_titles(train.Name.values)
plt.figure(figsize=(12,6))
sns.barplot(x=train.Name, y=train.Survived)
plt.xticks(rotation='vertical')
plt.show()
| 01-Titanic_Machine_Learning_from_Disaster/01_exploratory_data_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: CUDAtorch
# language: python
# name: cudatorch
# ---
# + id="resistant-artist"
import pandas as pd
import torch
import pytorch_lightning as pl
from tqdm import tqdm
import torchmetrics
import math
from urllib.request import urlretrieve
from zipfile import ZipFile
import os
import torch.nn as nn
import numpy as np
# + id="radio-effectiveness"
users = pd.read_csv(
"data/users.csv",
sep=",",
)
ratings = pd.read_csv(
"data/ratings.csv",
sep=",",
)
movies = pd.read_csv(
"data/movies.csv", sep=","
)
# + [markdown] id="constitutional-lyric"
# ## Pytorch dataset
# + id="terminal-aspect"
import pandas as pd
import torch
import torch.utils.data as data
from torchvision import transforms
import ast
from torch.nn.utils.rnn import pad_sequence
class MovieDataset(data.Dataset):
"""Movie dataset."""
def __init__(
self, ratings_file,test=False
):
"""
Args:
csv_file (string): Path to the csv file with user,past,future.
"""
self.ratings_frame = pd.read_csv(
ratings_file,
delimiter=",",
# iterator=True,
)
self.test = test
def __len__(self):
return len(self.ratings_frame)
def __getitem__(self, idx):
data = self.ratings_frame.iloc[idx]
user_id = data.user_id
movie_history = eval(data.sequence_movie_ids)
movie_history_ratings = eval(data.sequence_ratings)
target_movie_id = movie_history[-1:][0]
target_movie_rating = movie_history_ratings[-1:][0]
movie_history = torch.LongTensor(movie_history[:-1])
movie_history_ratings = torch.LongTensor(movie_history_ratings[:-1])
sex = data.sex
age_group = data.age_group
occupation = data.occupation
return user_id, movie_history, target_movie_id, movie_history_ratings, target_movie_rating, sex, age_group, occupation
# + id="dominican-canberra"
genres = [
"Action",
"Adventure",
"Animation",
"Children's",
"Comedy",
"Crime",
"Documentary",
"Drama",
"Fantasy",
"Film-Noir",
"Horror",
"Musical",
"Mystery",
"Romance",
"Sci-Fi",
"Thriller",
"War",
"Western",
]
for genre in genres:
movies[genre] = movies["genres"].apply(
lambda values: int(genre in values.split("|"))
)
sequence_length = 8
# + colab={"referenced_widgets": ["", "83a4c8aa292f42309167fd2b3df8ff9f"]} id="damaged-soldier" outputId="4327a927-29e4-4fdc-9769-d79c3bb64daa"
class BST(pl.LightningModule):
def __init__(
self, args=None,
):
super().__init__()
super(BST, self).__init__()
self.save_hyperparameters()
self.args = args
#-------------------
# Embedding layers
##Users
self.embeddings_user_id = nn.Embedding(
int(users.user_id.max())+1, int(math.sqrt(users.user_id.max()))+1
)
###Users features embeddings
self.embeddings_user_sex = nn.Embedding(
len(users.sex.unique()), int(math.sqrt(len(users.sex.unique())))
)
self.embeddings_age_group = nn.Embedding(
len(users.age_group.unique()), int(math.sqrt(len(users.age_group.unique())))
)
self.embeddings_user_occupation = nn.Embedding(
len(users.occupation.unique()), int(math.sqrt(len(users.occupation.unique())))
)
self.embeddings_user_zip_code = nn.Embedding(
len(users.zip_code.unique()), int(math.sqrt(len(users.sex.unique())))
)
##Movies
self.embeddings_movie_id = nn.Embedding(
int(movies.movie_id.max())+1, int(math.sqrt(movies.movie_id.max()))+1
)
self.embeddings_position = nn.Embedding(
sequence_length, int(math.sqrt(len(movies.movie_id.unique())))+1
)
###Movies features embeddings
genre_vectors = movies[genres].to_numpy()
self.embeddings_movie_genre = nn.Embedding(
genre_vectors.shape[0], genre_vectors.shape[1]
)
self.embeddings_movie_genre.weight.requires_grad = False #Not training genres
self.embeddings_movie_year = nn.Embedding(
len(movies.year.unique()), int(math.sqrt(len(movies.year.unique())))
)
# Network
self.transfomerlayer = nn.TransformerEncoderLayer(63, 3, dropout=0.2)
self.linear = nn.Sequential(
nn.Linear(
589,
1024,
),
nn.LeakyReLU(),
nn.Linear(1024, 512),
nn.LeakyReLU(),
nn.Linear(512, 256),
nn.LeakyReLU(),
nn.Linear(256, 1),
)
self.criterion = torch.nn.MSELoss()
self.mae = torchmetrics.MeanAbsoluteError()
self.mse = torchmetrics.MeanSquaredError()
def encode_input(self,inputs):
user_id, movie_history, target_movie_id, movie_history_ratings, target_movie_rating, sex, age_group, occupation = inputs
#MOVIES
movie_history = self.embeddings_movie_id(movie_history)
target_movie = self.embeddings_movie_id(target_movie_id)
positions = torch.arange(0,sequence_length-1,1,dtype=int,device=self.device)
positions = self.embeddings_position(positions)
encoded_sequence_movies_with_poistion_and_rating = (movie_history + positions) #Yet to multiply by rating
target_movie = torch.unsqueeze(target_movie, 1)
transfomer_features = torch.cat((encoded_sequence_movies_with_poistion_and_rating, target_movie),dim=1)
#USERS
user_id = self.embeddings_user_id(user_id)
sex = self.embeddings_user_sex(sex)
age_group = self.embeddings_age_group(age_group)
occupation = self.embeddings_user_occupation(occupation)
user_features = torch.cat((user_id, sex, age_group,occupation), 1)
return transfomer_features, user_features, target_movie_rating.float()
def forward(self, batch):
transfomer_features, user_features, target_movie_rating = self.encode_input(batch)
transformer_output = self.transfomerlayer(transfomer_features)
transformer_output = torch.flatten(transformer_output,start_dim=1)
#Concat with other features
features = torch.cat((transformer_output,user_features),dim=1)
output = self.linear(features)
return output, target_movie_rating
def training_step(self, batch, batch_idx):
out, target_movie_rating = self(batch)
out = out.flatten()
loss = self.criterion(out, target_movie_rating)
mae = self.mae(out, target_movie_rating)
mse = self.mse(out, target_movie_rating)
rmse =torch.sqrt(mse)
self.log(
"train/mae", mae, on_step=True, on_epoch=False, prog_bar=False
)
self.log(
"train/rmse", rmse, on_step=True, on_epoch=False, prog_bar=False
)
self.log("train/step_loss", loss, on_step=True, on_epoch=False, prog_bar=False)
return loss
def validation_step(self, batch, batch_idx):
out, target_movie_rating = self(batch)
out = out.flatten()
loss = self.criterion(out, target_movie_rating)
mae = self.mae(out, target_movie_rating)
mse = self.mse(out, target_movie_rating)
rmse =torch.sqrt(mse)
return {"val_loss": loss, "mae": mae.detach(), "rmse":rmse.detach()}
def validation_epoch_end(self, outputs):
avg_loss = torch.stack([x["val_loss"] for x in outputs]).mean()
avg_mae = torch.stack([x["mae"] for x in outputs]).mean()
avg_rmse = torch.stack([x["rmse"] for x in outputs]).mean()
self.log("val/loss", avg_loss, on_step=False, on_epoch=True, prog_bar=False)
self.log("val/mae", avg_mae, on_step=False, on_epoch=True, prog_bar=False)
self.log("val/rmse", avg_rmse, on_step=False, on_epoch=True, prog_bar=False)
def test_epoch_end(self, outputs):
users = torch.cat([x["users"] for x in outputs])
y_hat = torch.cat([x["top14"] for x in outputs])
users = users.tolist()
y_hat = y_hat.tolist()
data = {"users": users, "top14": y_hat}
df = pd.DataFrame.from_dict(data)
print(len(df))
df.to_csv("lightning_logs/predict.csv", index=False)
def configure_optimizers(self):
return torch.optim.AdamW(self.parameters(), lr=0.0005)
@staticmethod
def add_model_specific_args(parent_parser):
parser = ArgumentParser(parents=[parent_parser], add_help=False)
parser.add_argument("--learning_rate", type=float, default=0.01)
return parser
####################
# DATA RELATED HOOKS
####################
def setup(self, stage=None):
print("Loading datasets")
self.train_dataset = MovieDataset("data/train_data.csv")
self.val_dataset = MovieDataset("data/test_data.csv")
self.test_dataset = MovieDataset("data/test_data.csv")
print("Done")
def train_dataloader(self):
return torch.utils.data.DataLoader(
self.train_dataset,
batch_size=128,
shuffle=False,
num_workers=os.cpu_count(),
)
def val_dataloader(self):
return torch.utils.data.DataLoader(
self.val_dataset,
batch_size=128,
shuffle=False,
num_workers=os.cpu_count(),
)
def test_dataloader(self):
return torch.utils.data.DataLoader(
self.test_dataset,
batch_size=128,
shuffle=False,
num_workers=os.cpu_count(),
)
model = BST()
trainer = pl.Trainer(gpus=1,max_epochs=50)
trainer.fit(model)
# + id="quick-router"
| pytorch_bst.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # hipspy issue 88
#
# This notebook is to debug the following issue: https://github.com/hipspy/hips/issues/88
# and to find a good example of single channel grayscale JPEG HiPS on the CDS servers.
# %matplotlib inline
import matplotlib.pyplot as plt
import hips
from urllib.error import HTTPError
from skimage.io import imread
# ## Example tile that is single channel JPEG
url = 'https://github.com/hipspy/hips-extra/raw/master/datasets/samples/IRAC4/Norder3/Dir0/Npix299.jpg'
data = imread(url)
print(data.shape)
print(data.dtype)
# ## List HiPS that are single channel JPEG
# +
def get_tile_info(survey):
"""Fetch one tile and check it's data shape and dtype"""
meta = hips.HipsTileMeta(order=3, ipix=0, file_format='jpg')
url = survey.base_url + '/' + meta.tile_default_url
data = imread(url)
return dict(shape=data.shape, dtype=data.dtype, url=url)
def collect_all_info():
surveys = hips.HipsSurveyPropertiesList.fetch()
infos = []
for survey in surveys.data:
if 'hips_tile_format' not in survey.data:
print('Missing hips_tile_format info for:', survey.data['ID'])
continue
if 'jpeg' in survey.tile_format:
try:
info = get_tile_info(survey)
info['id'] = survey.data['ID']
print(info)
infos.append(info)
except HTTPError:
print('HTTP error:', survey.data['ID'])
return infos
infos = collect_all_info()
# -
from pprint import pprint
# pprint(surveys.data[0].data)
for info in infos:
if len(info['shape']) < 3:
print(info)
# +
# Apparently there's no public single-channel JPEG HiPS tiles?
# -
# ## Example of drawing a single-channel JPEG
#
# TODO: This is not a single-channel JPEG example at the moment!
# +
from astropy.coordinates import SkyCoord
from hips import WCSGeometry
geometry = WCSGeometry.create(
skydir=SkyCoord(0, 0, unit='deg', frame='galactic'),
width=2000, height=1000, fov="3 deg",
coordsys='galactic', projection='AIT',
)
# -
from hips import HipsSurveyProperties
url = 'http://alasky.u-strasbg.fr/2MASS/H/properties'
hips_survey = HipsSurveyProperties.fetch(url)
from hips import make_sky_image, SimpleTilePainter
# data = make_sky_image(geometry, hips_survey, 'jpg')
painter = SimpleTilePainter(geometry, hips_survey, 'jpg')
painter.run()
tile = painter.tiles[0]
print(tile.data.shape)
print(tile.data.dtype)
print(painter.image.shape)
print(painter.image.dtype)
| issue-88/issue-88.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **This notebook is an exercise in the [Geospatial Analysis](https://www.kaggle.com/learn/geospatial-analysis) course. You can reference the tutorial at [this link](https://www.kaggle.com/alexisbcook/your-first-map).**
#
# ---
#
# # Introduction
#
# [Kiva.org](https://www.kiva.org/) is an online crowdfunding platform extending financial services to poor people around the world. Kiva lenders have provided over $1 billion dollars in loans to over 2 million people.
#
# <center>
# <img src="https://i.imgur.com/2G8C53X.png" width="500"><br/>
# </center>
#
# Kiva reaches some of the most remote places in the world through their global network of "Field Partners". These partners are local organizations working in communities to vet borrowers, provide services, and administer loans.
#
# In this exercise, you'll investigate Kiva loans in the Philippines. Can you identify regions that might be outside of Kiva's current network, in order to identify opportunities for recruiting new Field Partners?
#
# To get started, run the code cell below to set up our feedback system.
# +
import geopandas as gpd
from learntools.core import binder
binder.bind(globals())
from learntools.geospatial.ex1 import *
# -
# ### 1) Get the data.
#
# 다음 셀을 사용하여 `loans_filepath`에 있는 shapefile을 로드하여 GeoDataFrame `world_loans`를 생성합니다.
# +
loans_filepath = "../input/geospatial-learn-course-data/kiva_loans/kiva_loans/kiva_loans.shp"
# Your code here: Load the data
world_loans = gpd.read_file(loans_filepath)
# Check your answer
#q_1.check()
# Uncomment to view the first five rows of the data
#world_loans.head()
# +
# Lines below will give you a hint or solution code
#q_1.hint()
#q_1.solution()
# -
# ### 2) Plot the data.
#
# #### 변경 없이 다음 코드 셀을 실행하여 국가 경계가 포함된 GeoDataFrame 'world'를 로드합니다.
# This dataset is provided in GeoPandas
world_filepath = gpd.datasets.get_path('naturalearth_lowres')
world = gpd.read_file(world_filepath)
world.head()
# `world' 및 `world_loans` GeoDataFrames를 사용하여 전 세계의 Kiva 대출 위치를 시각화합니다.
# +
# Your code here
ax = world.plot(figsize=(20,20), color='white', linestyle=':', edgecolor='red')
world_loans.plot(ax=ax, markersize=2)
# Uncomment to see a hint
#q_2.hint()
# +
# Get credit for your work after you have created a map
#q_2.check()
# Uncomment to see our solution (your code may look different!)
#q_2.solution()
# -
# ### 3) Select loans based in the Philippines.
#
# 다음으로 필리핀에 기반을 둔 대출에 중점을 둘 것입니다. 다음 코드 셀을 사용하여 필리핀에 기반을 둔 대출이 있는 `world_loans`의 모든 행을 포함하는 GeoDataFrame `PHL_loans`를 만듭니다.
# Your code here
PHL_loans = PHL_loans = world_loans.loc[world_loans.country=="Philippines"].copy()
# Check your answer
q_3.check()
# +
# Lines below will give you a hint or solution code
#q_3.hint()
#q_3.solution()
# -
# ### 4) Understand loans in the Philippines.
#
# 필리핀의 모든 섬에 대한 경계를 포함하는 GeoDataFrame `PHL`을 로드하려면 변경 없이 다음 코드 셀을 실행하십시오.
# Load a KML file containing island boundaries
gpd.io.file.fiona.drvsupport.supported_drivers['KML'] = 'rw'
PHL = gpd.read_file("../input/geospatial-learn-course-data/Philippines_AL258.kml", driver='KML')
PHL.head()
# 'PHL' 및 'PHL_loans' GeoDataFrames를 사용하여 필리핀의 대출을 시각화합니다.
# +
# Your code here
ax1 = PHL.plot(figsize=(20,20), color='white', linestyle=':', edgecolor='red')
PHL_loans.plot(ax=ax, markersize=2)
# Uncomment to see a hint
#q_4.a.hint()
# +
# Get credit for your work after you have created a map
#q_4.a.check()
# Uncomment to see our solution (your code may look different!)
#q_4.a.solution()
# -
# 새로운 필드 파트너를 모집하는 데 유용할 수 있는 섬을 식별할 수 있습니까? 현재 Kiva의 손이 닿지 않는 곳에 보이는 섬이 있습니까?
#
# [이 지도](https://bit.ly/2U2G7x7)가 질문에 답하는 데 유용할 수 있습니다.
# +
# View the solution (Run this code cell to receive credit!)
#q_4.b.solution()
#잠재적인 섬이 많이 있지만 필리핀 중부는 현재 데이터 세트에서 대출이 없는 비교적 큰 섬으로 두드러집니다. 이 섬은 잠재적으로 새로운 필드 파트너를 모집하기에 좋은 위치입니다!
# -
# # Keep going
#
# Continue to learn about **[coordinate reference systems](https://www.kaggle.com/alexisbcook/coordinate-reference-systems)**.
# ---
#
#
#
#
# *Have questions or comments? Visit the [course discussion forum](https://www.kaggle.com/learn/geospatial-analysis/discussion) to chat with other learners.*
| _notebooks/exercise-your-first-map (1).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import os
from plotnine import *
# ## Sanity check: read coverage file into a dataframe
# +
cov = []
for strand in ['fwd', 'rev']:
df = pd.read_csv('../results/yoon01/lncRNA_locus.{}.cov.gz'.format(strand),
usecols=[1,2], sep='\t', header=None, names=['position', 'coverage'])
df['strand'] = '+' if strand=='fwd' else '-'
cov.append(df)
covdf = pd.concat(cov)
covdf
# -
# ## Build `GTF` dataframe from the `lncRNA` annotation file
def gtf_df(filename):
res = []
with open(filename, 'rt') as fi:
for line in fi:
fields = line.strip().split('\t')
if fields[2] == 'exon':
rec = {}
idfields = fields[8].strip().split(';')
for idfield in idfields:
if idfield:
key, val = idfield.split()
if key == 'transcript_id' or key == 'exon_number':
rec.update({key: val.strip('"')})
rec.update({'chr': fields[0],
'start': int(fields[3]),
'end': int(fields[4])})
res.append(rec)
return pd.DataFrame.from_records(res)
gtf = gtf_df('../ref/lncRNA.gtf')
gtf
gtf['exon'] = ['exon {}'.format(x.exon_number) for _,x in gtf.iterrows()]
gtf['strand'] = '-'
gtf['track'] = gtf['transcript_id']
gtf
# ## Second annotation dataframe will only have `lnrCXCR4` and `UBBP1`
# +
# `exon 1` and `exon 3` are swapped in the .gtf file
# we correct it here
gtf_ = gtf.loc[4:,:]
gtf_.loc[4,'exon'] = 'exon 3'
gtf_.loc[6,'exon'] = 'exon 1'
gtf_.loc[7, 'track'] = 'UBBP1'
gtf_
# -
# ## Plotting functions
# +
# Need to convert .bam coords to genomic coords compatible with .gtf
OFFSET = gtf.start.min() - 501
# +
def feature_df(gff, features, feature_col='gene'):
if isinstance(features, str):
features = [features]
g = gff[gff[feature_col].isin(features)].copy()
g['length'] = np.abs(g.end - g.start)
return g
def cond_df(region, sample, res_dir='../results', strands=('fwd','rev')):
'''
For condition coverage calculated in
xx - Genome coverage by condition
'''
cov = []
if strands is None:
df = pd.read_csv(os.path.join(res_dir, sample, 'lncRNA_locus.cov.gz'),
usecols=[1,2], sep='\t', header=None,
names=['position', 'coverage'])
else:
for strand in strands:
df = pd.read_csv(os.path.join(res_dir, sample, 'lncRNA_locus.{strand}.cov.gz'.format(
strand=strand)),
usecols=[1,2], sep='\t', header=None,
names=['position', 'coverage'])
df['strand'] = '+' if strand==strands[0] else '-'
df['position'] += OFFSET
start,end = region
df = df[(df['position']>start) & (df['position']<end)]
cov.append(df)
data = pd.concat(cov)
return data
def region_plot(region, sample, strand='+', features=None, padding=None):
if features is not None:
region = (np.min(features['start']), np.max(features['end']))
if strand is not None:
strand = list(set(features['strand']))[0]
if padding is not None:
region = region[0] - padding, region[1] + padding
if strand is None:
data = cond_df(region, sample, strands=None)
data.loc[data['coverage']<0, 'coverage'] = 0
else:
data = cond_df(region, sample)
data = data[data['strand']==strand]
fmark = '>' if strand == '+' else '<'
g = ggplot(data, aes('position', 'coverage'))
g += geom_area(alpha=0.25)
if features is not None:
anno_list = []
ymax = -(np.max(data['coverage']))
features['center'] = features['start'] + features['length'] // 2
for i,track in enumerate(set(features['track'])):
features.loc[features['track']==track, 'fy'] = (i+1)*0.125*ymax
g += geom_tile(features, aes(x='center', y='fy', width='length', fill='track', height=0.1*ymax),
alpha=0.15)
g += geom_text(features, aes(x='center', y='fy', label='exon'), alpha=0.5, size=24)
g += geom_text(features, aes(x='start + length // 5', y='fy'), label=fmark, size=20, alpha=0.5)
if strand =='-':
g += scale_y_reverse()
g += theme_bw()
g += theme(figure_size=(18,12),
axis_title_x=element_text(size=28),
axis_title_y=element_text(size=28),
axis_text_x=element_text(size=22),
axis_text_y=element_text(size=22),
legend_text=element_text(size=24),
legend_title=element_text(size=28),
legend_entry_spacing_y=15
)
return g
# -
# ## Locus coverage plots
region_plot(None, 'yoon01', padding=100, strand=None,
features=feature_df(gtf_, list(gtf_['exon'].values), feature_col='exon'))
region_plot(None, 'yoon02', padding=100, strand=None,
features=feature_df(gtf_, list(gtf_['exon'].values), feature_col='exon'))
region_plot(None, 'yoon03', padding=100, strand=None,
features=feature_df(gtf_, list(gtf_['exon'].values), feature_col='exon'))
| notebooks/02 - Locus coverage plots.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# name: ir
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/nestorsgarzonc/Data-Science-Proyects/blob/master/Credit_card_approval_R.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="UO-VaLr0EtFq"
# #Importar librerias necesarias
# + [markdown] colab_type="text" id="uMqtkYmWE3K8"
# ## Descargar paquetes
# + [markdown] colab_type="text" id="Xy1rDKDBOXyv"
# Al instalar los paquetes puede tardar aproximadamente 5 minutos
#
# Nota: mnormt es una dependencia del paquete psych, se intala una version anterior por compatibilidad de version con la 2.0 de mnormt
#
#
# + colab={} colab_type="code" id="9qeBa8tjQhxY"
packageurl <- "https://cran.r-project.org/src/contrib/Archive/mnormt/mnormt_1.5-7.tar.gz"
install.packages(packageurl, repos=NULL, type="source")
# + colab={} colab_type="code" id="V1xBjU5qD9Qu"
install.packages("psych", repos='https://www.icesi.edu.co/CRAN/')
install.packages("dplyr") #Resumenes estadisticos
install.packages("ggplot2") #Graficos en general
install.packages('RCurl') #Paquete para obtener archivos desde internet
install.packages("modeest") #Calcular moda
install.packages("dplyr") #Agrupar datos
install.packages("fdth") #Graficos descriptivos
install.packages("ggpubr") #Grafico de medias
# + [markdown] colab_type="text" id="S0lp1Ge_E7Ub"
# ##Cargar paquetes
# + colab={} colab_type="code" id="lM6qVfc7EzbM"
library("ggplot2")
library("psych")
library("dplyr")
library ("RCurl")
library ("modeest")
library("fdth")
library("ggpubr")
# + [markdown] colab_type="text" id="YtB2c83tNyq_"
# # Cargar dataset
# + [markdown] colab_type="text" id="akoe1v2pN-yk"
# Nota: el dataset original tenia un problema de interpretacion en colab, por ello se modifico la codificacion y esta disponible ambas versiones en el siguiente repositorio: https://github.com/nestorsgarzonc/Data-analysis-R
# + colab={} colab_type="code" id="g8vx6PFsFfQ0"
#download <- getURL("https://raw.githubusercontent.com/nestorsgarzonc/Data-analysis-R/master/Base_Banco.csv")
#data <- read.table (text = download, sep = ';', fileEncoding = "UCS-2LE")
# + colab={} colab_type="code" id="FUmNxFC5Fw8B"
#head(data)
# + colab={} colab_type="code" id="NkA0NvK8IZRO"
#dim(data)
# + colab={} colab_type="code" id="ERTTxL2xLlOD"
#colnames(data) <- as.character(unlist(data[1,]))
#data = data[-1, ]
# + [markdown] colab_type="text" id="QfwHNG_8Dl7i"
# Importar dataframe modificado, se descarga el dataset alojado en GitHub usando el metodo:
#
# ```
# getURL('URL')
# ```
#
# Como la primera columna del dataset (X) es el indice del dataset, se define como el index del dataframe
# ```
# read.csv(row.names='X')
# ```
#
#
#
#
# + colab={} colab_type="code" id="-zr8pGBmNl65"
download <- getURL("https://raw.githubusercontent.com/nestorsgarzonc/Data-analysis-R/master/banco_dataframe.csv")
BaseBanco <- read.csv(text = download, row.names = 'X')
# + [markdown] colab_type="text" id="kpZRaE3dRncv"
# #Exploracion de datos
# + [markdown] colab_type="text" id="AN_73olrCF6v"
# Las variables estan definidas de la siguiente manera
#
#
# 1. **Edad**: Edad en años del cliente.
# 2. **Ocupación**: Ocupación del cliente.
# 3. **Est_civil**: Estado civil del cliente.
# 4. **Nivel_educativo**: Máximo nivel escolar alcanzado por el cliente.
# 5. **Contrato_meses**: Duración del último o actual contrato de trabajo del cliente.
# 6. **Cred_hipotecario**: Si el cliente tiene o no crédito hipotecario vigente con el banco.
# 7. **Cred_personal**: Si el cliente tiene o no crédito personal vigente con el banco.
# 8. **Medio_contacto**: Medio por el cual se contactó al cliente.
# 9. **Día**: Día del mes en el que se realizó el último contacto con el cliente.
# 10. **Mes**: Mes del año en el que se realizó el último contacto con el cliente.
# 11. **Duración_seg**: Duración en segundos del último contacto con el cliente.
# 12. **Cont_campaña_actual**: Número de veces que se contactó al cliente en la campaña mencionada.
# 13. **Campañas_previas**: Número de campañas de promoción en las que se ha incluido al cliente previamente.
# 14. **Aceptación_TC**: Si el cliente aceptó o no la tarjeta de crédito en la campaña mencionada.
# + [markdown] colab_type="text" id="eU305SYy2KyU"
# A continuacion se realiza la inspeccion de las 5 primeras y 5 ultimas columnas del dataset
# + colab={"base_uri": "https://localhost:8080/", "height": 176} colab_type="code" id="MSKotsHiB0lq" outputId="b48d59d6-eb6d-401f-b818-9288194a53ad"
head(BaseBanco)
# + colab={"base_uri": "https://localhost:8080/", "height": 176} colab_type="code" id="NYr9KrgjHwYE" outputId="e5ad1676-9333-4fe0-8f46-8ed4a987f901"
tail(BaseBanco)
# + colab={"base_uri": "https://localhost:8080/", "height": 33} colab_type="code" id="g8VIy8M-B2my" outputId="6945f418-04c3-4112-d9ac-4e8673e15d25"
dim(BaseBanco)
# + [markdown] colab_type="text" id="TMgFcRnt2eBX"
# Nuestro dataset esta conformado de 4521 filas y 14 columnas las cuales son estan distribuidas de la siguiente manera
# + colab={"base_uri": "https://localhost:8080/", "height": 303} colab_type="code" id="x51qh5SyRqkc" outputId="3d4b1b24-72f7-4ec2-b47f-431b9810252c"
str(BaseBanco)
# + [markdown] colab_type="text" id="L0Cd3-nI2uuw"
# El dataset esta conformado por 14 variables las cuales 6 son cuantitativas y 8 son cualitativas.
# Las variables cualitativas estan definidas como factores y tienen las siguientes clases:
# + colab={"base_uri": "https://localhost:8080/", "height": 66} colab_type="code" id="wcLLe2Lc5ejU" outputId="78cae567-c404-4467-8459-1cd9c0f6fb2d"
#Ocupacion
unique(BaseBanco$`ocupación`)
# + colab={"base_uri": "https://localhost:8080/", "height": 50} colab_type="code" id="EGZOCMoa5qs1" outputId="b0549295-809e-4a65-bb1a-c806669280e4"
#Estado civil
unique(BaseBanco$est_civil)
# + colab={"base_uri": "https://localhost:8080/", "height": 50} colab_type="code" id="8wu9Lx7J5qwi" outputId="7e648594-022e-4e7e-9160-80917a28fa49"
#Nivel educativo
unique(BaseBanco$nivel_educativo)
# + colab={"base_uri": "https://localhost:8080/", "height": 50} colab_type="code" id="uh1Eurw47UGr" outputId="36796c38-162e-44a9-e670-13dd68d8a0b9"
#Medio de contacto
unique(BaseBanco$Medio_contacto)
# + [markdown] colab_type="text" id="xJWBSid48Mja"
# Las siguientes variables categoricas tienen salida binaria, si o no
# + colab={"base_uri": "https://localhost:8080/", "height": 116} colab_type="code" id="7GXdbXkZ7KTg" outputId="f61dbf96-6235-4f48-c9db-ccd01549a950"
#Credito hipotecario
unique(BaseBanco$Cred_hipotecario)
#Credito personal
unique(BaseBanco$Cred_personal)
#Aceptacion de la tarjeta de credito
unique(BaseBanco$aceptacion_TC)
# + [markdown] colab_type="text" id="hTWE3cfM8dg1"
# A continuacion vamos a desestructurar el dataframe en variables para mayor comodidad y renombrar algunas variables que contienen simbolos como tildes, ñ y demas para seguir una misma convencion 👨💻👩💻
# + colab={} colab_type="code" id="fTvFvFcK7UfH"
attach(BaseBanco)
# + colab={"base_uri": "https://localhost:8080/", "height": 66} colab_type="code" id="D-FonN_U7Kb3" outputId="4aeb4e57-58e2-4dc5-a72a-551532d24b45"
colnames(BaseBanco)
# + colab={} colab_type="code" id="lHzuGIZG7KW6"
ocupacion <- `ocupación`
cred_hipotecario <- Cred_hipotecario
cred_personal <- Cred_personal
medio_contacto <- Medio_contacto
dia <- Dia
mes <- Mes
cont_campana_actual <- `cont_campaña_actual`
campanas_previas <- `campañas_previas`
# + [markdown] colab_type="text" id="ID1-xWoHRrS7"
# #Analisis datos
# + [markdown] colab_type="text" id="15c0z3bf-FQa"
# ## Analisis variables cuantitativas
# + [markdown] colab_type="text" id="OjT2CP7BD0DI"
# Las variables cuantitativas son:
#
#
# 1. edad
# 2. contrato_meses
# 3. dia
# 4. duracion_seg
# 5. cont_campana_actual
# 6. campanas_previas
#
# Las variables cuantitativas a analizar individualmente son:
#
#
# 1. edad
# 2. contrato_meses
# 3. duracion_seg
# 4. cont_campana_actual
#
# + [markdown] colab_type="text" id="Mhu0U1nXYCxs"
# ###Analisis general
#
# Haciendo el summary de nuestro dataset se puede ver que para la edad media de los clientes es de 41 años con al parecer presencia de datos atipicos.
#
# La mayoria de clientes son personas con un nivel educativo de bachillerato y universitario, son personas casadas y solteras, administradores e informales.
#
# Generalmente hacen un contrato de 36 meses (la media y la mediana estan en un valor muy cercano, se prevee que sea simetrico), la mayoria de personas no tiene un credito vigente con el banco
#
# El medio de contacto es fuertemente por celular siendo generalmente a mitad de mes (notese que la media y la mediana son muy cercanas), siendo fuertemente para casi desde mitad de año (mayo) con una duración de contacto medio de 185 segundos.
#
# El numero de campañas que se realizo tiene una media de 0.54 y el numero de campañas en las que se contacto al cliente fue de 2, para estas dos variables seria interesante un analisis con media recortada ya que las colas estan fuertementes inclinadas en ambos casos. Finalmente la proporcion de personas que aceptan la tarjeta de credito es de 0.1152
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 585} colab_type="code" id="UNqRv0pSYIBn" outputId="03658c7f-766a-41f5-aee5-2736bf9bf3d6"
summary(BaseBanco)
# + [markdown] colab_type="text" id="pHHKRIWfYc0e"
# ###Analisis especifico (por cada variable)
# + [markdown] colab_type="text" id="KeCy8IrfqDzE"
# ####Edad
# + [markdown] colab_type="text" id="h8PyQAWkWOYY"
# La media de la edad al ser sensible a datos atipicos se procede a analizar la media con media recortada recortada para reducir la inestabilidad de la media concluyendo que esta mucho mas cercano a la mediana que es una medida robusta.
#
# La moda de la edad es unica (unimodal), la edad media de los clientes es de 39-40 años
#
# + colab={"base_uri": "https://localhost:8080/", "height": 101} colab_type="code" id="tjqQDiHE7Khp" outputId="e330a18a-d39c-4edf-9c55-5c3cb10bc1ec"
#Tendencia central y posicion
paste('Media:', mean(edad))
paste('Mediana:', median(edad))
paste('Moda:', mfv(edad))
print('Quantiles:')
quantile(edad)
# + colab={"base_uri": "https://localhost:8080/", "height": 33} colab_type="code" id="fOH9aiiP7KnW" outputId="6aee0c8f-fd1c-4a20-971b-4dbf1fb3fccf"
paste('Media con media recortada al 20%: ', mean(edad, trim=0.2))
# + [markdown] colab_type="text" id="2QM3xFOCjbM8"
# Podemos ver que la edad va desde los 19 años hasta los 87 años, con un rango intercuantilico de 16 años
# + colab={"base_uri": "https://localhost:8080/", "height": 66} colab_type="code" id="y2-1MzbPiwM2" outputId="2662288e-fd7d-4f42-d47d-c8e2474602ab"
#Dispersion:
paste('Rango de la edad', range(edad))
paste('Rango intercuartilico de la edad', IQR(edad))
paste('Varianza de la edad', var(edad))
# + colab={"base_uri": "https://localhost:8080/", "height": 33} colab_type="code" id="Hcipp63N7Kym" outputId="f7414174-d594-401a-f2ba-8a334423296a"
paste('La edad de los clientes estan desviados de la media un: ', sd(edad), 'de la muestra')
# + colab={"base_uri": "https://localhost:8080/", "height": 66} colab_type="code" id="LZGhNzYG7Kwj" outputId="9d74c1a0-f3b1-4681-82c8-715a35d3dc24"
limiteSup<-mean(edad)+sd(edad)
limiteInf<-mean(edad)-sd(edad)
paste('La edad de los clientes que estan a una desviacion estandar de la media son:')
paste('X-s: ', limiteInf)
paste('X+s: ', limiteSup)
# + colab={"base_uri": "https://localhost:8080/", "height": 33} colab_type="code" id="b-6Rfy4J7Kqa" outputId="8feca056-c20c-492c-e25f-6fffe2f55f04"
numClientesEdad<-sum(edad>limiteInf & edad<limiteSup)
paste('La cantidad de clientes que esta a una desviacion estandar son de:',
numClientesEdad)
# + [markdown] colab_type="text" id="JNm1_I76me8P"
# El coeficiente de variacion al ser un valor bajo significa que no hay mucha dispersion respecto la media
# + colab={"base_uri": "https://localhost:8080/", "height": 33} colab_type="code" id="BjrhYaQ07KlY" outputId="24c070f1-60aa-4d19-9103-1a33641d09ef"
cv <- sd(edad)/mean(edad)*100
paste('El coeficiente de variacion es de: ', cv)
# + [markdown] colab_type="text" id="m_xiLHyGnrvT"
# La simetria al ser positiva cercana a cero significa que la cola esta levemente distribuida a la derecha
# Por curtosis se sabe que la curva es mesocurtica tiende a cero
# + colab={"base_uri": "https://localhost:8080/", "height": 50} colab_type="code" id="hYIoq2Hn7KfY" outputId="2e75afa2-0dd6-45d3-890f-4aca8e5be765"
paste('Asimetria de la edad: ', skew(edad))
paste('Curtosis de la edad: ', kurtosi(edad))
# + colab={"base_uri": "https://localhost:8080/", "height": 436} colab_type="code" id="obKebW4R7KZs" outputId="f9bb915e-1718-4bbc-f194-f2997dc14d89"
hist(edad, freq = F, main = "Histograma edad de los clientes", xlab = "edad", ylab="Frecuencia relativa")
curve(dnorm(x,mean=mean(edad),sd=sd(edad)), col="darkred",lwd=2, add=TRUE)
# + [markdown] colab_type="text" id="CAzwwUQht6u7"
# ####Contrato por meses
#
# El contrato a meses es la duracion del actual o ultimo contrato de trabajo del cliente.
#
# La media y la mediana estan muy cercanas.
# El contrato por meses sigue una moda del tipo unimodal
# + colab={"base_uri": "https://localhost:8080/", "height": 101} colab_type="code" id="powTxcWwRsoW" outputId="1f65c3d9-1651-4aae-af43-7ae3f2a404df"
#Tendencia central y posicion
paste('Media:', mean(contrato_meses))
paste('Mediana:', median(contrato_meses))
paste('Moda:', mfv(contrato_meses))
print('Quantiles:')
quantile(contrato_meses)
# + [markdown] colab_type="text" id="c3OLQNRfvbFN"
# El contrato por meses va desde 1 hasta 72 meses con un rango intercuantilico de 36
# + colab={"base_uri": "https://localhost:8080/", "height": 66} colab_type="code" id="rBAyFVlNuYk_" outputId="75ac2582-15b8-4455-a686-049b6e3aebf5"
#Dispersion:
paste('Rango de la contrato_meses', range(contrato_meses))
paste('Rango intercuartilico de la contrato_meses', IQR(contrato_meses))
paste('Varianza de la contrato_meses', var(contrato_meses))
# + colab={"base_uri": "https://localhost:8080/", "height": 50} colab_type="code" id="zvfAkvgrvY9_" outputId="0b20330c-414f-4e08-ee9f-5dceeb961cc9"
paste('El contrato por meses de los clientes estan desviados de la media un: ',
sd(contrato_meses), 'de la muestra')
# + colab={"base_uri": "https://localhost:8080/", "height": 66} colab_type="code" id="ZjaPyr3hvoL3" outputId="24311fb1-95dd-4dc6-c5b4-a2cf0ff583ba"
limiteSup<-mean(contrato_meses)+sd(contrato_meses)
limiteInf<-mean(contrato_meses)-sd(contrato_meses)
paste('El contrato por meses de los clientes que estan a una desviacion estandar de la media son:')
paste('X-s: ', limiteInf)
paste('X+s: ', limiteSup)
# + [markdown] colab_type="text" id="rlUN57WVwEoM"
# El coeficiende de variacion al ser un valor alto significa que hay dspersion de los datos respecto la media
# + colab={"base_uri": "https://localhost:8080/", "height": 33} colab_type="code" id="ijWRJkPrv5Aa" outputId="de3dee12-3171-4ffb-c213-f14def1d3149"
cv <- sd(contrato_meses)/mean(contrato_meses)*100
paste('El coeficiente de variacion es de: ', cv)
# + [markdown] colab_type="text" id="N8WmrjnUwdM9"
# La asimetria por ser cercana a cero significa que esta levemente inclinada a la derecha y la curtosis es negativa por ende es platicurtica
# + colab={"base_uri": "https://localhost:8080/", "height": 50} colab_type="code" id="-T203rCXwTtz" outputId="1d82e959-a8b0-48cb-f6e6-9cda24f512be"
paste('Asimetria: ', skew(contrato_meses))
paste('Curtosis: ', kurtosi(contrato_meses))
# + colab={"base_uri": "https://localhost:8080/", "height": 436} colab_type="code" id="XyWr8pyMwpGo" outputId="80e77a25-29b4-4f04-c313-4f261f2ef018"
hist(contrato_meses, freq = F, main = "Histograma contrato por meses de los clientes", xlab = "contrato_meses", ylab="Frecuencia relativa", xlim=range(-10, 90), ylim=range(0, 0.02) )
curve(dnorm(x,mean=mean(contrato_meses),sd=sd(contrato_meses)), col="darkred",lwd=2, add=TRUE)
# + [markdown] colab_type="text" id="8phzOf6bywqf"
# ####Duracion segundos contacto cliente
# + [markdown] colab_type="text" id="EFIpr_Cv00sZ"
# La duracion en segundos del ultimo contacto con el cliente tiene una media y mediana alejada por presencia de datos atipicos, por ello vamos a tomar la media recortada al 20% para evitar los posibles datos atipicos, como se ve en el rango intercuantilico pasamos de tener 329 en el 75% a tener 3025 en el 100%. La media recortada ahora esta mas cercana a la mediana
# + colab={"base_uri": "https://localhost:8080/", "height": 101} colab_type="code" id="svgOixfTw7Dv" outputId="78cff834-5a01-4c16-d545-6c5f828b697e"
#Tendencia central y posicion
paste('Media:', mean(duracion_seg))
paste('Mediana:', median(duracion_seg))
paste('Moda:', mfv(duracion_seg))
print('Quantiles:')
quantile(duracion_seg)
# + colab={"base_uri": "https://localhost:8080/", "height": 33} colab_type="code" id="wFrAM9fZywbs" outputId="199929ae-6753-454a-f552-e2dae4d89390"
paste('Media con media recortada al 20%: ', mean(duracion_seg, trim=0.2))
# + [markdown] colab_type="text" id="a-RqvmM-16cm"
# En el anterior punto se analizo la presencia de grandes datos atipicos, en el rango intercuantilico va desde 3 hasta 3025
# + colab={"base_uri": "https://localhost:8080/", "height": 66} colab_type="code" id="s4UzXIjT1QwJ" outputId="08409133-9e1b-40c4-d7b3-e2dc662206db"
#Dispersion:
paste('Rango de la duracion_seg', range(duracion_seg))
paste('Rango intercuartilico de la duracion_seg', IQR(duracion_seg))
paste('Varianza de la duracion_seg', var(duracion_seg))
# + colab={"base_uri": "https://localhost:8080/", "height": 50} colab_type="code" id="kk_SnHH413gO" outputId="94553768-4612-47c5-f31a-7456e2272f84"
paste('El ultimo contacto con los clientes estan desviados de la media un: ',
sd(duracion_seg), 'de la muestra')
# + colab={"base_uri": "https://localhost:8080/", "height": 66} colab_type="code" id="tddRm8l8hnhf" outputId="2f4ab046-570d-4431-b507-ed80234e7b8a"
limiteSup<-mean(duracion_seg)+sd(duracion_seg)
limiteInf<-mean(duracion_seg)-sd(duracion_seg)
paste('El ultimo contacto con los clientes que estan a una desviacion estandar de la media son:')
paste('X-s: ', limiteInf)
paste('X+s: ', limiteSup)
# + [markdown] colab_type="text" id="3cMKDgSkhyg_"
# El coeficiente de variacion de lo clientes al ser un valor alto signiifica que hay una alta dispersion de los datos respecto la media
# + colab={"base_uri": "https://localhost:8080/", "height": 33} colab_type="code" id="gSWNZdh8hsKK" outputId="a31aef96-4c1b-47ee-9c70-74a95dd4d6ad"
cv <- sd(duracion_seg)/mean(duracion_seg)*100
paste('El coeficiente de variacion es de: ', cv)
# + [markdown] colab_type="text" id="o9jEdTYEiARt"
# La asimetria al ser positiva significa que la cola esta a la derecha con un nivel de cursosis leptocurtico
# + colab={"base_uri": "https://localhost:8080/", "height": 50} colab_type="code" id="fyoeWUhDh10G" outputId="b1e9b9c8-177f-472c-a5c0-b1843aa79744"
paste('Asimetria: ', skew(duracion_seg))
paste('Curtosis: ', kurtosi(duracion_seg))
# + colab={"base_uri": "https://localhost:8080/", "height": 436} colab_type="code" id="BKx8Qc2NiA0U" outputId="959ba020-abaf-40c4-94c2-dd761155c27d"
hist(duracion_seg, freq = F, main = "Histograma contrato por meses de los clientes", xlab = "duracion_seg", ylab="Frecuencia relativa")
curve(dnorm(x,mean=mean(duracion_seg),sd=sd(duracion_seg)), col="darkred",lwd=2, add=TRUE)
# + [markdown] colab_type="text" id="5vMQoH6fjOZx"
# ###Analisis varias varibles
#
# En esta sesion vamos a comparar graficos las siguientes variables y hacer su respectivo analisis:
# 1. Edad vs aceptacion tarjeta de credito
# 2. Medio de contacto vs contrato meses
# 3. Contador camapaña actual vs aceptacion tarjeta credito
# 4. Medio de contacto vs Dia
# 5. Dia vs aceptacion TC
# 6. Nivel educativo vs contrato meses
# 7. Estado civil vs contrato meses
# + [markdown] colab_type="text" id="rphMim2Yk1gS"
# ####1. Edad vs aceptacion tarjeta de credito
#
# Segun la grafica comparando si las personas aceptaron o no una tarjeta de credito segun las edades se puede concluir que tiene una asimetria positiva en ambos casos significando que las personas mas joven sean el publico objetivo para los bancos, no obstante son muchas mas las personas que rechazan que las que aceptan la tarjeta.
# En personas de edad mayor (vease la cola) se puede ver que el banco son las que menos acuden
# + colab={"base_uri": "https://localhost:8080/", "height": 436} colab_type="code" id="t2jnCEYRiY-X" outputId="0b4a20a4-cb63-4051-d443-d65d85982aac"
par(mfrow=c(2,1))
hist(edad[aceptacion_TC=="si"], main="Edad en la que acepta tarjeta credito", xlab="edad", ylab="")
hist(edad[aceptacion_TC=="no"], main="Edad en la que no acepta tarjeta credito", xlab="edad", ylab="")
# + [markdown] colab_type="text" id="Fic_1-comxDv"
# ####2. Medio de contacto vs contrato meses
#
# Por las graficas se puede ver que se comporta de manera casi uniforme excepto por la cola los ultimos meses.
# Se puede ver que la mayoria de los clientes fueron por celular siendo el canal mas efectivo seguido por el chat. Para cuando son una gran cantidad de meses se puede ver que hay una menor cantidad de clientes.
# + colab={"base_uri": "https://localhost:8080/", "height": 436} colab_type="code" id="a0QjwddKlUlo" outputId="a25f8a2f-128d-4445-e164-052e56f94cdb"
par(mfrow=c(3,1))
hist(contrato_meses[medio_contacto=="celular"], main="Tiempo de contrato por canal de adquisicion (Celular)", xlab="meses", ylab="")
hist(contrato_meses[medio_contacto=="chat"], main="Tiempo de contrato por canal de adquisicion (Chat)", xlab="meses", ylab="")
hist(contrato_meses[medio_contacto=="telefono"], main="Tiempo de contrato por canal de adquisicion (Telefono)", xlab="meses", ylab="")
# + [markdown] colab_type="text" id="gN4HN39BpY8P"
# ####3. Contador campaña actual vs aceptacion tarjeta credito
#
# Segun los histogramas se puede ver que los datos siguen una alta asimetria positiva mostrando que la mayoria de veces en que se tiene una respuesta son en las primeras iteraciones.
# + colab={"base_uri": "https://localhost:8080/", "height": 436} colab_type="code" id="a2ZdBG60nMoB" outputId="6c1f32be-9cda-4f48-a33b-688b1987147e"
par(mfrow=c(2,1))
hist(cont_campana_actual[aceptacion_TC=="si"], main="Numero de veces que se contacto al cliente y acepto la tarjeta de credito", xlab="# contactos", ylab="")
hist(cont_campana_actual[aceptacion_TC=="no"], main="Numero de veces que se contacto al cliente y no acepto la tarjeta de credito", xlab="# contactos", ylab="")
# + [markdown] colab_type="text" id="DB5Hq0eXrKxd"
# ####4. Medio de contacto vs dia
#
# Como se puede ver en las graficas las solicitudes de las tarjetas de credito al principio y a final de mes se comporta muy similar en el caso de el celular y chat como canal de adquisicion
# + colab={"base_uri": "https://localhost:8080/", "height": 436} colab_type="code" id="6ese2qCwpunw" outputId="88c9f49e-734b-4567-9f17-7f2043fc099e"
par(mfrow=c(3,1))
hist(dia[medio_contacto=="celular"], main="Dia en que se le solicito una tarjeta por celular", xlab="Dia del mes", ylab="")
hist(dia[medio_contacto=="chat"], main="Dia en que se le solicito una tarjeta por chat", xlab="Dia del mes", ylab="")
hist(dia[medio_contacto=="telefono"], main="Dia en que se le solicito una tarjeta telefono", xlab="Dia del mes", ylab="")
# + [markdown] colab_type="text" id="jFJNwgFMtYcn"
# ####5. Dia vs aceptacion
#
# Segun los dos histogramas los clientes que aceptan o rechazan la tarjeta mantiene un comportamiento similar
# + colab={"base_uri": "https://localhost:8080/", "height": 436} colab_type="code" id="j_SBEwtfrX9-" outputId="bf4c7190-e746-4bb3-e383-75bb49ee7803"
par(mfrow=c(2,1))
hist(dia[aceptacion_TC=="si"], main="Dia en que se le solicito una tarjeta y acepto tarjeta de credito", xlab="Dia del mes", ylab="")
hist(dia[aceptacion_TC=="no"], main="Dia en que se le solicito una tarjeta y no acepto tarjeta de credito", xlab="Dia del mes", ylab="")
# + [markdown] colab_type="text" id="x6T2-xokxaJR"
# ####6. Nivel educativo vs contrato por meses
#
# Se puede ver que hay mas personas en contratos de bachilleratos y menos en tecnicos.
# En todos los niveles educativos se ve que hay una menor cantidad de contratos para una cantidad de meses larga, de resto se comporta de manera casi uniforme.
# + colab={"base_uri": "https://localhost:8080/", "height": 436} colab_type="code" id="eLLLnmJovBIf" outputId="b1d612c5-af80-4004-87a9-c278a4b59455"
par(mfrow=c(2,2))
hist(contrato_meses[nivel_educativo=="primaria"], main="contrato por meses primaria", xlab="contrato meses", ylab="")
hist(contrato_meses[nivel_educativo=="bachillerato"], main="contrato por meses bachillerato", xlab="contrato meses", ylab="")
hist(contrato_meses[nivel_educativo=="universitario"], main="contrato por meses universitario", xlab="contrato meses", ylab="")
hist(contrato_meses[nivel_educativo=="técnico"], main="contrato por meses tecnico", xlab="contrato meses", ylab="")
# + [markdown] colab_type="text" id="8MVL-85i0_OM"
# ####7. Estado civil vs contrato meses
#
# Bonus: Segun este grafico las personas solteras, casadas o divorciadas mantienen un contrato con un comportamiento muy similar, pero las muestras para las personas casadas son grandes mientras que las de las personas divorsiadas son aproximadamente 1/4
# + colab={"base_uri": "https://localhost:8080/", "height": 436} colab_type="code" id="BUJ_27FPyyWo" outputId="1a37952d-0002-4bbf-8e49-97ebed526f98"
par(mfrow=c(2,2))
hist(contrato_meses[est_civil=="casado"], main="contrato por meses casado", xlab="contrato meses", ylab="")
hist(contrato_meses[est_civil=="soltero"], main="contrato por meses soltero", xlab="contrato meses", ylab="")
hist(contrato_meses[est_civil=="divorciado"], main="contrato por meses divorciado", xlab="contrato meses", ylab="")
# + [markdown] colab_type="text" id="9f682Z_6EBMu"
# ##Analisis variables cualitativas
#
# Las variables cualitativas son:
#
# * ocupación
# * est_civil
# * nivel_educativo
# * Cred_hipotecario
# * Cred_personal
# * Medio_contacto
# * Mes
# * aceptacion_TC
#
# Las variables a analizar individualmente son:
#
# * ocupación
# * est_civil
# * nivel_educativo
# * Cred_hipotecario
# * aceptacion_TC
# + [markdown] colab_type="text" id="s8nmSPf6Jg8o"
# ###Analisis por cada variable
# + [markdown] colab_type="text" id="Ps4OiY7BJrER"
# ####Ocupacion
#
# Por ocupacion, la mayor proporcion de clientes esta dada por administradores, empresarios y profesores
# + colab={"base_uri": "https://localhost:8080/", "height": 141} colab_type="code" id="NA0sj2wh1nvt" outputId="4f060753-2465-4fa9-ed82-8a514a22ef93"
tabla_ocupacion <- table(ocupacion)
tabla_ocupacion
# + colab={"base_uri": "https://localhost:8080/", "height": 141} colab_type="code" id="YSZllefrJ-pj" outputId="09b5c856-ef5c-4dbe-ef21-52ec9b837c60"
prop.table(tabla_ocupacion)
# + [markdown] colab_type="text" id="hCn2026qSW3d"
# ####Estado civil
#
# Por estado civil la mayor proporcion de clientes esta dada por personas casadas
# + colab={"base_uri": "https://localhost:8080/", "height": 70} colab_type="code" id="huLDg8V5KFBi" outputId="fa08d708-9547-4861-c307-0507ee470d60"
tabla_est_civil <- table(est_civil)
tabla_est_civil
# + colab={"base_uri": "https://localhost:8080/", "height": 70} colab_type="code" id="ucld20evR9xl" outputId="3095e1d1-f6e4-4f80-a2fc-122fc278680b"
prop.table(tabla_est_civil)
# + [markdown] colab_type="text" id="CpGwwMMDTFpn"
# ####Nivel educativo
#
# Para el nivel educativo la mayor proporcion de personas esta dada por personas de bachilleres y universitarios
# + colab={"base_uri": "https://localhost:8080/", "height": 70} colab_type="code" id="1pk98tw8SlVd" outputId="c2a7f349-ea56-4cf9-f383-b7de843201d0"
tabla_nivel_educativo <- table(nivel_educativo)
tabla_nivel_educativo
# + colab={"base_uri": "https://localhost:8080/", "height": 70} colab_type="code" id="CVSUwYcCTL-4" outputId="535b0c33-683a-44ee-e957-de92038178fe"
prop.table(tabla_nivel_educativo)
# + [markdown] colab_type="text" id="vs3yo1TjTgew"
# ####Credito hipotecario
#
# El 56.60% de los clientes tiene un credito hipotecario vigente con el banco
# + colab={"base_uri": "https://localhost:8080/", "height": 70} colab_type="code" id="t5v_Hs-5TPMg" outputId="332b5fbb-0824-448c-ee0e-f5f582e21f8f"
tabla_cred_hipotecario <- table(cred_hipotecario)
tabla_cred_hipotecario
# + colab={"base_uri": "https://localhost:8080/", "height": 70} colab_type="code" id="uDXL7y5wT9x6" outputId="174e4ebe-773c-4de7-aeab-4a03e1bd8959"
prop.table(tabla_cred_hipotecario)
# + [markdown] colab_type="text" id="Zu7CnKm4UWtR"
# ####Aceptacion tarjeta de credito
#
# El 88.476% de los clientes no aceptaron la tarjeta de credito
# + colab={"base_uri": "https://localhost:8080/", "height": 70} colab_type="code" id="MR6S9SoBUBw4" outputId="76fc4c1b-d4c7-4a8f-9a0a-152224445fe5"
tabla_aceptacion_TC <- table(aceptacion_TC)
tabla_aceptacion_TC
# + colab={"base_uri": "https://localhost:8080/", "height": 70} colab_type="code" id="EY2wBcQ7UdjF" outputId="0fbc09de-d48c-4d87-c0db-9c0cebbdc240"
prop.table(tabla_aceptacion_TC)
# + colab={} colab_type="code" id="fSfQUKIqUfs8"
| Credit_card_approval_R.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
s = pd.Series([1, 2, 3, 5, np.nan, 7, 8])
s
dates = pd.date_range('20170208', periods=6)
dates
df = pd.DataFrame(np.random.randn(6,4), index=dates, columns=list('ABCD'))
df
np.random.randn(1000, 4)
| 1-neural-networks/jupyter-demo/pandas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="fya744P_lD2_"
# # Objective of Course
#
# * Apply Python concepts to real world application development
# + [markdown] colab_type="text" id="HDegPGoilFuo"
# # Application Level Components
#
# * User interface
# * Data storage
# * Connectivity
# * Main logic
# * Testing frameworks
# + [markdown] colab_type="text" id="33pHS_ChE0Vx"
# <br><br><font color="#B81590">$$\large-\infty-$$</font><br><br>
# + [markdown] colab_type="text" id="3rMgpdjRlts5"
# ## User Interface
#
# An application doesn't necessarily need a graphical user interface (GUI). It can be run automatically at certain times or under certain conditions. For example this can run at 3 AM.
#
# `cleanup_tmp_files.py --location /tmp`
#
# Alternatively, it can be a command line interface similar to the Python interpreter.
#
# ```
# todo.py
# Enter your task: Clean the dishes
# Task created
# ```
#
# For graphical interfaces, Python can use `Tkniter`, `wxPython`, or many other graphical interface frameworks.
#
# This presentation will be based on the first example, which may be run automatically.
# + [markdown] colab_type="text" id="Lwwqe6ESEyK7"
# <br><br><font color="#B81590">$$\large-\infty-$$</font><br><br>
# + [markdown] colab_type="text" id="mIj6mq_ODoR8"
# ## Data Storage
#
# Data storage can be in a database, flat files, or even an Excel file.
#
# ### <font color="#D21087">Excel Files</font>
#
# <font color='#544640'>We'll open an Excel (.xlsx) containing some stock ticker symbols with prices.
#
# We're going to use `pandas`, a very powerful data science and data manipulation library that can handle large amounts of multidimensional "panel data" (hence the name) efficiently. It is generally used for data science and computing applications.
#
# It's also convenient for accessing and handling tabular data. There are *many* libraries that can handle Excel files, `pandas` is only one. `pandas` comes with some caveats (and limitations) that we won't go into here, related to it's original intended use; i.e. it is definitely not just a 'excel reader' library!
#
# Side note: `pandas` objects are called `dataframes`, much like R's native data structure and are very similar.
#
# For our purposes, here is the relevant doc:
#
# https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.read_excel.html</font>
# + colab={} colab_type="code" id="2UlfPzsB2eH6"
# environment setup
import os
import pandas as pd
# + [markdown] colab_type="text" id="NNRQ2boB4NCL"
# First we will import the data from the spreadsheet into Python and Pandas.
# + colab={} colab_type="code" id="sbYfR0tt4TtI"
data = pd.read_excel('/content/Ticker Symbols.xlsx', sheet_name = None)
# + [markdown] colab_type="text" id="tmaxYjhp4y3o"
# <font color='#544640'>That was easy. When we specify sheet_name, we can do so using `pd.read_excel` returns a `dict` whose key values are the names of the sheets in the `xlsx` document.</font>
#
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="L6zYnmAE44Aj" outputId="6b26409f-fd0d-42b0-8598-76727bbc4c27"
list(data.keys())
# + [markdown] colab_type="text" id="K6gvjwsl5HDJ"
# Notice the list above shows the worksheet from the spreadsheet. Now we can list the stocks we want to pull the price for.
# + colab={"base_uri": "https://localhost:8080/", "height": 104} colab_type="code" id="84APntb_5Zik" outputId="9c00ab94-c810-4c2a-d3c4-238c43c47aa8"
dataframe = data['Stocks']
dataframe['Ticker']
# + [markdown] colab_type="text" id="kxj5vvWq5wsG"
# Very cool. We now have a list of ticker symbols we want to retrieve stock prices for.
#
# ---
#
#
# + [markdown] colab_type="text" id="ZuYh9pg0pqWG"
# <br><br><font color="#B81590">$$\large-\infty-$$</font><br><br>
# + [markdown] colab_type="text" id="whJddCSJdJGh"
# ## Screen Scraping
#
# This example is not relevant to our stocks application but it is good to know how to do.
#
# <font color='#544640'>In this simple example we're going to do some very basic web scraping.
#
# Please note the random pause/wait time introduced in the loops in this example. It's important not to get yourself into trouble by sending too many requests too frequently to the site you are accessing. Your computer will basically try to (mini-)DoS a target host if you aren't careful.
# -
# ### <font color="#D21087">Libraries Used:</font><font color='#544640'>
#
# * `BeautifulSoup4`: https://readthedocs.org/projects/beautiful-soup-4/
# * `requests`: http://docs.python-requests.org/en/master/
# * `re`: https://docs.python.org/3/howto/regex.html
# +
# set up environment
import bs4 as bs
from bs4 import BeautifulSoup
from bs4.element import Comment
import re
# these two packages do almost the same thing
# used one for one example and one for another
import urllib.request
import requests
# -
# ### <font color="#D21087">Get All Links on a Page:</font><font color='#544640'>
# <font color='#544640'>You can easily write a custom web-crawler/scraper by traversing links one by one through a domain. Use at your own risk - don't say I didn't warn you. :)</font>
# +
target_page = 'https://en.wikipedia.org/wiki/Insight_Enterprises'
page_data = requests.get(target_page)
# use this regular expression to strip out HTML tags, if needed
# re.sub('<[^<]+?>', '', page_data.text)
soup = bs.BeautifulSoup(page_data.text, 'html.parser')
links = soup.find_all('a', attrs={'href': re.compile('^http://')})
for link in links[:10]:
print(link.get('href'))
# -
# <font color='#544640'>Doesn't Wikipedia have a lot more clickable links on any given page than the ones in the article references? What's going on?</font>
# ### <font color="#D21087">Find Specific Text on a Page:</font><font color='#544640'>
# <font color='#544640'>Sometimes, instead, you want to find or check for the presence of specific text.</font>
# +
# wonderful example from https://stackoverflow.com/questions/1936466/beautifulsoup-grab-visible-webpage-text
def tag_visible(element):
if element.parent.name in ['style', 'script', 'head', 'title', 'meta', '[document]']:
return False
if isinstance(element, Comment):
return False
return True
def text_from_html(body):
soup = BeautifulSoup(body, 'html.parser')
texts = soup.findAll(text=True)
visible_texts = filter(tag_visible, texts)
return u" ".join(t.strip() for t in visible_texts)
html = urllib.request.urlopen('https://en.wikipedia.org/wiki/Insight_Enterprises').read()
text_content = text_from_html(html)
# +
lines = text_content.split('. ')
for l in lines:
print(l)
print('\n')
# + [markdown] colab_type="text" id="MwgAeZ0n6c9I"
# ## Connectivity
#
# Connectivity can mean a lot of things. It can be connecting to a database for your data storage retrieval. Today's example will require us to retrieve the stock prices for the tickers in the Excel spreadsheet.
#
# `requests` is a very common library for HTTP requests in Python. `requests` will communicate with Alpha Vantage's API (https://www.alphavantage.co/) for stock price lookups.
#
# First, we need to import requests.
# + [markdown] colab_type="text" id="OHowDPZi7Kb5"
# First, we need to construct our URL based on the API documentation (https://www.alphavantage.co/documentation/).
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="yc2yDSjt7PbW" outputId="6d4d55d5-24be-488e-fed9-c6227b8a6a5a"
import requests
API_KEY=""
params = {'function': 'TIME_SERIES_DAILY',
'symbol': 'MSFT',
'apikey': API_KEY}
response = requests.get('https://www.alphavantage.co/query', params=params)
response.url
# + [markdown] colab_type="text" id="gFwGRWb376nq"
# Notice the URL has the keys and values in the dictionary I specified. What did Alpha Advantage return to us? Lets see in JSON format.
# + colab={"base_uri": "https://localhost:8080/", "height": 8771} colab_type="code" id="Bg3Ib6Ua8Ds9" outputId="6aa2fe81-b459-497b-b839-693fe073ca40"
response.json()
# + [markdown] colab_type="text" id="9e_8oIXr8QR4"
# That's a lot. 100 days of data to be precise. What about today's opening price?
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="B0eOys1F8U3C" outputId="667c0774-3fa1-400d-f4e8-f88e912b6d2d"
from datetime import datetime
current = datetime.today().strftime('%Y-%m-%d')
prices = response.json()
prices['Time Series (Daily)'][current]['1. open']
# + [markdown] colab_type="text" id="e-ufKeFroTl8"
# We have the proper components ready to go for this example. Data storage via Excel. Connectivity over a RESTful API using requests. Lets tie it together.
# + [markdown] colab_type="text" id="Teea5iIXpsH6"
# <br><br><font color="#B81590">$$\large-\infty-$$</font><br><br>
# + [markdown] colab_type="text" id="Cz0vLrKRoznG"
# ## Main Logic
#
# The main logic of the program is really what controls everything. It's the glue that brings your Lego pieces together. Don't be that guy. Don't use glue on your Lego pieces.
#
# First, we should move the components into functions.
# + colab={} colab_type="code" id="lhzHyVicpiF-"
def get_stock_opening_price(symbol):
params = {'function': 'TIME_SERIES_DAILY',
'symbol': symbol,
'apikey': API_KEY}
response = requests.get('https://www.alphavantage.co/query', params=params)
response_json = response.json()
return response_json['Time Series (Daily)'][current]['1. open']
def read_from_excel(filename):
return pd.read_excel(filename, sheet_name=None)
# + [markdown] colab_type="text" id="SmYPBT7jqW98"
# Next, we will call `read_from_excel()` to get a `list`
# of stock symbols to query.
#
# + colab={"base_uri": "https://localhost:8080/", "height": 104} colab_type="code" id="4vFAClxBqmOo" outputId="4d5622d3-e10b-41d4-f8ba-f127fb453c1f"
data = read_from_excel('/content/Ticker Symbols.xlsx')
dataframe = data['Stocks']
for index, row in dataframe.iterrows():
dataframe.loc[index, "Price"] = get_stock_opening_price(row['Ticker'])
print(dataframe)
# + [markdown] colab_type="text" id="AjE5MFrVtSMo"
# Finally, we should write the information back to the Excel file.
# + colab={} colab_type="code" id="-_I-63l-0nIF"
with pd.ExcelWriter('/content/Ticker Symbols.xlsx') as writer:
dataframe.to_excel(writer, sheet_name='Stocks', index=False)
# + [markdown] colab_type="text" id="JkGmjztPps_2"
# <br><br><font color="#B81590">$$\large-\infty-$$</font><br><br>
# + [markdown] colab_type="text" id="jl0d_TnCyY2S"
# ## Testing Frameworks
#
# Any application that is more than just a simple script should have some automatible testing associated to it.
#
# There are three main types of tests - unit, integration, and validation.
#
# > In computer programming, **unit testing** is a software testing method by which individual units of source code, sets of one or more computer program modules together with associated control data, usage procedures, and operating procedures, are tested to determine whether they are fit for use. - https://en.wikipedia.org/wiki/Unit_testing
#
# > **Integration testing** (sometimes called integration and testing, abbreviated I&T) is the phase in software testing in which individual software modules are combined and tested as a group. Integration testing is conducted to evaluate the compliance of a system or component with specified functional requirements. It occurs after unit testing and before validation testing. Integration testing takes as its input modules that have been unit tested, groups them in larger aggregates, applies tests defined in an integration test plan to those aggregates, and delivers as its output the integrated system ready for system testing. - https://en.wikipedia.org/wiki/Integration_testing
#
# > In software project management, software testing, and software engineering, **verification and validation** (V&V) is the process of checking that a software system meets specifications and that it fulfills its intended purpose. It may also be referred to as software quality control. It is normally the responsibility of software testers as part of the software development lifecycle. In simple terms, software verification is: "Assuming we should build X, does our software achieve its goals without any bugs or gaps?" On the other hand, software validation is: "Was X what we should have built? Does X meet the high level requirements?"- https://en.wikipedia.org/wiki/Software_verification_and_validation
#
# Today we will mostly discuss unit testing and touch on integration tests.
# + [markdown] colab_type="text" id="Zv-4MaXpzgJ6"
# ### Unit Testing
#
# Unit tests perform a test against a single piece of code. Each test case should be tested independently from other test cases.
# + colab={} colab_type="code" id="j71srmahz9DN"
def add(a, b):
return a + b
def subtract(a, b):
return a - b
def multiply(a, b):
return a * b
def divide(a, b):
return a / b
# + [markdown] colab_type="text" id="7PdSTsej0ENK"
# The following class is using the built-in Python `unittest` module to perform unit tests against the functions.
# + colab={"base_uri": "https://localhost:8080/", "height": 121} colab_type="code" id="kbX7cIzY00bP" outputId="f9be84b9-7489-4562-9c11-49fb9bae4bce"
import unittest
class MasteryNotebook(unittest.TestCase):
def test_add(self):
self.assertEqual(add(1,2), 3)
self.assertEqual(add(0,1), 1)
self.assertEqual(add(-1,-1), -2)
def test_subtract(self):
self.assertEqual(subtract(2, 1), 1)
self.assertEqual(subtract(0, 1), -1)
def test_multiply(self):
self.assertEqual(multiply(2, 1), 2)
self.assertEqual(multiply(-1, 1), -1)
def test_divide(self):
with self.assertRaises(ZeroDivisionError):
divide(1, 0)
unittest.main(argv=[''], verbosity=1, exit=False)
# + [markdown] colab_type="text" id="UMUo1OT85vuu"
# #### Mocks
#
# Remember, a unit test is meant to test an isolated piece of code. What if your unit test requires another source, such as a local database or network connection?
#
# > In object-oriented programming, **mock objects** are simulated objects that mimic the behavior of real objects in controlled ways, most often as part of a software testing initiative. - https://en.wikipedia.org/wiki/Mock_object
#
# Mocks go beyond the content of this course. But they allow you to simulate an external response in a controlled manner. In other words, the mock pretends to be what you want it to be (ex. database call).
#
# *Note:* [Some people](http://arlobelshee.com/tag/no-mocks/) don't like mocks and think it means there is room for improvement with code structure. I'm not opinionated here. Do what accomplishes your task.
# + [markdown] colab_type="text" id="YzXfo2NN7WjI"
# ### Integration Tests
#
# Integration testing is frequently accomplished using `tox`, which is a Python testing tool. You can also use a full Continuous Integration (CI) system such as TravisCI or Jenkins to run your tests. This is outside the scope of this presentation.
# + [markdown] colab_type="text" id="2sTNsPitTJyw"
# ### Coverage
#
# Code coverage means how many lines of code are actually tested. But does that mean that as long as a line of code is tested it is properly tested? No. In the unit test example above, the `test_divide()` test suite only tests for the exception. I'd argue the test suite doesn't fully cover all the cases it should test for. Design your unit tests as well as you can and build-out over time. They won't be perfect day one.
# + [markdown] colab_type="text" id="z0dOlXf5puBN"
# <br><br><font color="#B81590">$$\large-\infty-$$</font><br><br>
# + [markdown] colab_type="text" id="dyovVB8dlsPi"
# # Resources
#
#
#
# [wxPython](https://www.wxpython.org/)
#
# [Pandas](https://pandas.pydata.org/)
#
# [Python Context Managers and the "with" Statement](https://realpython.com/courses/python-context-managers-and-with-statement/)
#
# [Requests Library](https://2.python-requests.org/en/master/)
#
# [Getting Started With Testing in Python - Real Python](https://realpython.com/python-testing/)
#
# [Demystifying the Patch Function - Video](https://www.youtube.com/watch?v=ww1UsGZV8fQ)
#
# [Reading and Writing Files in Python](https://realpython.com/read-write-files-python/)
#
# [Automate the Boring Stuff with Python](https://automatetheboringstuff.com/)
| Mastery_Intermediate_Programming_with_Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Iterators Exercises
#
# A string is iterable because you can go through each letter that composes the string.
# It's not an iterator. Running iter("string") returns an iterator.
# The iterator will call next() on the iterator until it has completely cycled through.
#
# when the iterator is exhausted and there is nothing more that comes from next(), Python throws a StopIterator error.
#
# 1. Create our own for loop
# +
class Myiterator():
def __init__(self, iterable):
self.iterable = iterable
self.idx = 0
self.limit = len(iterable)
def __iter__(self):
return self
def __next__(self):
r = self.iterable[self.idx]
self.idx += 1
if self.idx >= self.limit:
raise StopIteration
return r
def my_for(iterable, function):
I = iter(Myiterator(iterable))
for i in I:
function(i)
def square(x):
print(x*x)
my_for("hello", print)
my_for([1, 2, 3, 4, 5], square)
# -
# 2. Write a custom iterator.
# +
class Counter:
def __init__(self, low, high, inc=1):
self.current = low
self.high = high
self.it = low
self.inc = inc
# this needs to return an iterator
def __iter__(self):
return self
def __next__(self):
r = self.it
self.it += self.inc
if r >= self.high:
raise StopIteration
return r
for x in Counter(0, 10,2):
print(x)
# -
| exercises/03 iterators.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
os.chdir('../..')
import sys
sys.path.append("/workspace/src")
# ## Imports
# +
from util.misc import load, save
from databases.datasets import PersonStackedMuPoTsDataset, PersonStackedMucoTempDataset, Mpi3dTestDataset, Mpi3dTrainDataset
from databases.joint_sets import CocoExJoints, MuPoTSJoints, JointSet
from util.pose import _calc_limb_length
from databases import mpii_3dhp, mupots_3d
import numpy as np
# %matplotlib notebook
# -
def plfig(lines=1, full_width=False):
if lines==1 and not full_width:
plt.figure()
elif full_width:
plt.figure(figsize=(9.5, 3*lines))
else:
plt.figure(figsize=(4.5, 3*lines))
# ## Bone stability
# Visualize the length of a bone in training/test set.
#
# It seems in Muco-Temp (and therefore in MPII-3DHP), bones are normalized except for:
# left/right hip-knee (30-40)
# spine-neck (~10)
# neck-shoulder (5-10)
#
# hip is also the average of lefthip/right, which is not the same as in the captury videos. It seems left/right hip joints are synthetic and are not coming directly from captury.
class FullMpiiSet(JointSet):
NAMES = np.array(['spine3', 'spine4', 'spine2', 'spine', 'pelvis',
'neck', 'head', 'head_top', 'left_clavicle', 'left_shoulder', 'left_elbow',
'left_wrist', 'left_hand', 'right_clavicle', 'right_shoulder', 'right_elbow', 'right_wrist',
'right_hand', 'left_hip', 'left_knee', 'left_ankle', 'left_foot', 'left_toe',
'right_hip' , 'right_knee', 'right_ankle', 'right_foot', 'right_toe'])
NUM_JOINTS=28
NAMES.flags.writeable = False
# +
full_names = ['spine3', 'spine4', 'spine2', 'spine', 'pelvis',
'neck', 'head', 'head_top', 'left_clavicle', 'left_shoulder', 'left_elbow',
'left_wrist', 'left_hand', 'right_clavicle', 'right_shoulder', 'right_elbow', 'right_wrist',
'right_hand', 'left_hip', 'left_knee', 'left_ankle', 'left_foot', 'left_toe',
'right_hip' , 'right_knee', 'right_ankle', 'right_foot', 'right_toe']
parent1 = np.array([3, 1, 4, 5, 5, 2, 6, 7, 6, 9, 10, 11, 12, 6, 14, 15, 16, 17, 5, 19, 20, 21, 22, 5, 24, 25, 26, 27 ])-1
parent2 = np.array([4, 3, 5, 5, 5, 1, 2, 6, 2, 6, 9, 10, 11, 2, 6, 14, 15, 16, 4, 5, 19, 20, 21, 4, 5, 24, 25, 26]) -1
for i in range(len(full_names)):
print('%15s %15s %s' % (full_names[i], full_names[parent1[i]], full_names[parent2[i]]))
# -
val_data = PersonStackedMucoTempDataset('hrnet', 'normal')
test_data = PersonStackedMuPoTsDataset('hrnet', 'normal', 'all')
refine_results = load('results_smoothed_83-2955.pkl')
class RefResults:
index= refine_results['index']
poses3d= refine_results['pred']
refine_data = RefResults()
mpi_train_data = Mpi3dTrainDataset('hrnet', 'normal', 'megadepth_at_hrnet', True, 2)
mpi_test_data = Mpi3dTestDataset('hrnet', 'normal', 'megadepth_at_hrnet')
gt = mupots_3d.load_gt_annotations(16)
validFrame = gt['isValidFrame']
# +
BONES = [['left_ankle', 'left_knee'], ['left_hip', 'left_knee'], ['left_hip', 'hip'],
['hip', 'spine'], ['spine', 'head/nose'], ['left_shoulder', 'left_elbow']]
# BONES = [['right_ankle', 'right_knee'], ['right_hip', 'right_knee'], ['right_hip', 'hip'],
# ['hip', 'spine'], ['spine', 'neck'], ['right_shoulder', 'right_elbow']]
# BONES = [['neck', 'right_shoulder'], ['right_shoulder', 'right_elbow'], ['right_elbow', 'right_wrist']]
# BONES = [['spine', 'neck'], ['neck', 'head/nose'], ['head/nose', 'head_top']]
# BONES=[['left_ankle', 'left_knee'], ['left_hip', 'left_knee'],['left_shoulder', 'left_elbow']]
joint_set = MuPoTSJoints()
data = mpi_train_data
seqs = np.unique(data.index.seq)
seq = np.random.choice(seqs)
# seq='16/2'
print(seq)
inds = data.index.seq==seq
plfig(1, False)
# plt.subplot(1,3,1)
names=['ankle-knee', 'knee-hip', 'elbow-shoulder']
for i,bone in enumerate(BONES):
lens = _calc_limb_length(data.poses3d[inds], joint_set, [bone])
plt.plot(lens, label=bone[0])
print(np.std(lens))
# ax2=plt.gca().twinx()
# ax2.plot(gt['occlusions'][:,2, joint_set.index_of('left_shoulder')], color='black')
plt.legend()
# -
# mupots: 16/2 - jumps, all frames are valid
# +
# Mupots gt vs pred
joint_set = MuPoTSJoints()
seqs = np.unique(test_data.index.seq)
seq = np.random.choice(seqs)
print(seq)
inds = test_data.index.seq==seq
assert np.all(refine_data.index.seq[inds]==seq)
bones = [['left_ankle', 'left_knee'], ['left_knee', 'left_hip', ], ['left_hip', 'hip'],
['right_wrist', 'right_elbow'], ['right_elbow', 'right_shoulder', ], ['right_shoulder', 'neck']]
plfig(2, True)
for i, bone in enumerate(bones):
plt.subplot(2,3,i+1)
lens = _calc_limb_length(test_data.poses3d[inds], joint_set, [bone])
plt.plot(lens, label='gt')
lens = _calc_limb_length(refine_data.poses3d[inds], joint_set, [bone])
plt.plot(lens, label='pred')
plt.title('%s %s' % (bone[0], bone[1]))
plt.tight_layout()
# + [markdown] heading_collapsed=true
# ### Mpii-train data full joints
# + hidden=true
sub=7
seq=1
annot = load(os.path.join(mpii_3dhp.MPII_3DHP_PATH, 'S%d' % sub, 'Seq%d' % seq, 'annot.mat'))
annot3 = list([x[0].reshape((-1, 28, 3)).astype('float32') for x in annot['annot3']])
# + hidden=true
lhip = joint_set.index_of('left_hip')
rhip = joint_set.index_of('right_hip')
np.std(np.linalg.norm((p[:,rhip]+p[:,lhip])/2-p[:,joint_set.index_of('pelvis')], axis=1))
# + hidden=true
# is there any joint the knee is normalized to? answer: No
lhip = joint_set.index_of('left_knee')
p = annot3[7]
for i in range(28):
print(i, FullMpiiSet.NAMES[i], np.std(np.linalg.norm(p[:,lhip]-p[:,i], axis=1)))
# + hidden=true
#
BONES = [['left_hip', 'pelvis'], ['neck', 'left_clavicle'], ['spine4', 'neck']]
joint_set = FullMpiiSet()
plfig()
for bone in BONES:
lens = _calc_limb_length(annot3[9], joint_set, [bone])
plt.plot(lens, label=bone[0])
print(np.std(lens))
plt.legend()
| src/notebooks/analysis/inspect_results.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lab 2 - Comparison with Diffusion Theory
# The purpose of this lab is to compare flux and reaction rates computed using MCNP (or OpenMC) with those that you compute using Diffusion Theory.
#
# A mono-energtic point source is located at the center of a sphere of lead. The spatial flux distribution and reaction rates computed with OpenMC will be compared with the same results from Diffusion Theory.
#
# ## Diffusion Theory Results
# It is useful to have a good idea of what the expected result will be even before you create your model and begin computation.
#
# For this lab we will compute and plot diffusion theory results to which we will compare OpenMC tally outputs.
#
# The given data will be the macroscopic cross section for scattering and absorption: $\Sigma_a$ and $\Sigma_s$ respectively; and the average cosine of neutron scattering angle: $\bar{\mu}$.
#
# From these, the transport mean free path $(\lambda_{tr})$ is computed:
# $$\lambda_{tr}=\frac{1}{\Sigma_{tr}}=\frac{1}{\Sigma_s(1-\bar{\mu})}$$
#
# The diffusion coefficient $(D)$:
# $$D = \frac{\lambda_{tr}}{3}$$
# and, for diffusion theory with vacuum boundary conditions, the extrapolation distance $(d)$ is needed:
# $$d=0.71 \lambda_{tr}$$
#
# The diffusion length $(L)$ is also needed which is computed:
# $$L = \sqrt{\frac{D}{\Sigma_a}}$$
#
# ### Parameters
lead_density = 11.35; # g/cm3
Sigma_a_lead = 5.03e-3; # 1/cm, macroscopic absorption cross section for lead
Sigma_s_lead = 0.3757; # 1/cm
mu_bar = 3.2e-3;
# +
import numpy as np
lam_tr = 1./(Sigma_s_lead*(1.-mu_bar))
D = lam_tr/3.
d = 0.71*lam_tr
L = np.sqrt(D/Sigma_a_lead)
# -
# ### Analytic Results
# For a point source in spherical coordinates, the flux as a functio of position is:
# $$\phi(r)=\frac{S \sinh{\left(\frac{R+d-r}{L}\right)}}{4 \pi D r \sinh{\left(\frac{R+d}{L} \right)}}$$
# Where $S$ is the source strength (neutrons/s) and $R$ is the radius of the diffusive medium; all other variables as previously defined.
#
# The diffusion theory solution for flux $(\phi(r))$ for this problem is plotted below.
# +
# %matplotlib inline
import matplotlib.pyplot as plt
S = 1000; #n/sec, source strength
R = 10; # cm, lead diffusive medium radius
def analytic_flux(r):
return S*np.sinh((R+d-r)/(L))/(4.*np.pi*D*r*np.sinh((R+d)/L))
r = np.linspace(1e-1,R,1000)
phi = analytic_flux(r)
analytic_sol = plt.semilogy(r,phi);
plt.setp(analytic_sol,'color','b','linewidth',3);
plt.xlabel('r [cm]',fontsize=14,fontweight='bold');
plt.ylabel('$\phi(r)$ [1/cm^2-s]',fontsize=14,fontweight='bold');
plt.grid(True);
plt.title('Analytic Flux',fontsize=16,fontweight='bold');
# -
# The lab also asks the students to compute the rate of neutron absorption. Accounting for angular symmetry and assuming constant material properties, the absorption rate should be:
# $$\text{Absorption Rate } = 4 \pi \Sigma_a \int_0^R \phi(r) r^2 \ dr$$
# +
import scipy.integrate as integrate
abs_rate,err_bnd = integrate.quad(lambda r: analytic_flux(r)*(r**2),
0,R)
abs_rate = 4.*np.pi*Sigma_a_lead*abs_rate
print(f'Diffusion Theory Absorption rate: {abs_rate:4.3f} per second')
# -
# ## OpenMC Model
import openmc
# ## Materials
#
# For this lab, the only material we need is lead. We will enter this by nuclide so students can get practice with the interface.
lead = openmc.Material(name='lead')
lead.add_nuclide('Pb204',0.014)
lead.add_nuclide('Pb206',0.241)
lead.add_nuclide('Pb207',0.221)
lead.add_nuclide('Pb208',0.524)
lead.set_density('g/cm3',11.35)
# Alternatively, the lead could have been added elementally
# +
lead = openmc.Material(name='lead')
lead.add_element('Pb',1.0)
lead.set_density('g/cm3',11.35)
# if I had the correct atom densities in the box above, these *should*
# be equivalent.
mf = openmc.Materials([lead])
mf.export_to_xml()
# -
# ## Geometry
# For a simple geometric domain, I will have a sphere of lead surrounded by a void.
# +
sphere_rad = 10.; # cm, radius of sphere
sphere = openmc.Sphere(r=sphere_rad)
outside = openmc.Sphere(r=1.5*sphere_rad,boundary_type='vacuum')
lead_sphere = openmc.Cell()
lead_sphere.fill = lead
lead_sphere.region = -sphere
outside_void = openmc.Cell()
outside_void.fill = None
outside_void.region = +sphere & -outside
root = openmc.Universe()
root.add_cells([lead_sphere,outside_void])
g = openmc.Geometry()
g.root_universe = root
g.export_to_xml()
# -
# ## Tallies
# For this lab, tallies are needed for the following values:
# <ol>
# <li>neutron absorption rate in the lead sphere </li>
# <li>flux as a function of radius (out to the extrapolated radius) </li>
# </ol>
# +
tallies = openmc.Tallies() # object to hold all tallies
sphere_filter = openmc.CellFilter(lead_sphere.id)
t = openmc.Tally(name='abs_tally')
t.filters = [sphere_filter]
t.scores = ['absorption']
#t.nuclides = ['all'] #<-- if you want absorption broken out by nuclide
tallies.append(t)
regMesh = openmc.RegularMesh()
meshDim = 31
regMesh.dimension = (meshDim,meshDim)
regMesh.lower_left = (0,0)
regMesh.upper_right = (R,R)
mesh_filt = openmc.MeshFilter(regMesh)
t = openmc.Tally(name='flux_tally')
t.filters = [mesh_filt]
t.scores = ['flux']
tallies.append(t)
tallies.export_to_xml()
# -
# ## Settings
# This is another fixed-source problem much like lab 1
# +
settings = openmc.Settings()
settings.run_mode = 'fixed source'
settings.batches = 50
settings.particles = 1000000
source = openmc.Source()
source.particle = 'neutron'
source.space = openmc.stats.Point(xyz=(0.,0.,0.))
source.angle = openmc.stats.Isotropic();
source.energy = openmc.stats.Discrete([0.0253],[1.0]) #0.0253 eV source
source.strength = S
settings.source = source;
settings.export_to_xml()
# -
openmc.run()
# ## Check Results
# Now we will open the statepoint file and analyze the results
sp = openmc.StatePoint('statepoint.50.h5')
abs_tally = sp.get_tally(name='abs_tally')
df = abs_tally.get_pandas_dataframe()
df.head()
abs_rate_omc = df['mean'][0] # there must be a better way
print(f'Absorption Rate from OpenMC: {abs_rate_omc:6.3f} per second.')
sp.tallies
flux_tally = sp.get_tally(name='flux_tally')
df_flux = flux_tally.get_pandas_dataframe()
df_flux.head()
df_flux.max()
df_flux['mean'].max()
# +
from matplotlib.colors import LogNorm # so I can use log of tally value
fluxes = df_flux['mean'].values
rel_err = df_flux['std. dev.'].values / fluxes
Xs = df_flux['mesh 1']['x'].values
Ys = df_flux['mesh 1']['y'].values
fluxes.shape = (meshDim,meshDim)
rel_err.shape = (meshDim,meshDim)
#fluxes = fluxes.T
#rel_err = rel_err.T
Xs.shape = (meshDim,meshDim)
Ys.shape = (meshDim,meshDim)
fig = plt.subplot(121)
plt.imshow(fluxes,interpolation='none',norm=LogNorm(),cmap='jet');
plt.grid(False);
plt.ylim(plt.ylim()[::-1]) #invert the y-axis
plt.title('Flux Tally - mean values',fontsize=14,fontweight='bold');
#plt.colorbar();
fig2 = plt.subplot(122)
plt.imshow(rel_err,interpolation='none',cmap='jet');
plt.ylim(plt.ylim()[::-1]);
plt.title('Rel. Unc.',fontsize=14,fontweight='bold');
plt.grid(False);
#plt.colorbar();
# -
# Note that the relative uncertainty increases as you move further from the source. *(take a moment to consider how weird this is in the context of other numerical algorithms in scientific computing)* This is a statistical reality when using Monte Carlo Methods. The issue is that fewer particles travel that far from the source and, as a consequence, there are fewer "scoring" opportunities with those tally sites. A large part of the theory and practical toolset of a Monte Carlo code has the aim of addressing this problem.
fluxes[int(meshDim/2),int(meshDim/2)]
fluxes.max()
fluxes.min()
flux_profile = fluxes[0,:]
X_ord = Xs[0,:]
X_ord = X_ord*(R/(meshDim+1))
plt.semilogy(X_ord,flux_profile,label='OpenMC',linestyle='--',
linewidth=4);
plt.semilogy(r,phi,label='Diffusion Theory',linestyle='-.',
linewidth=4);
plt.xlabel('R [cm]',fontsize=12,fontweight='bold')
plt.ylabel('$\phi$ [$n/cm^2-s$]',fontsize=12,fontweight='bold')
plt.legend();
plt.grid(True);
plt.title('Flux Comparison',fontsize=14,fontweight='bold');
| lab2/.ipynb_checkpoints/lab2_notebook-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + nbsphinx="hidden"
# %matplotlib inline
# %config InlineBackend.figure_format = 'svg'
import itertools
import numpy as np
import matplotlib.pyplot as plt
# -
# # Python examples
#
# This is a short description on how to use the DLR representation and the algorithms using this representation implemented in the python module `pydlr`.
#
# If you find this useful in your work please cite the companion paper to the `pydlr` module, see the [citation information](background.rst).
# ## DLR Grids
#
# The DLR representation sets up near optimal grids in
#
# - real frequency $\omega$,
# - imaginary time $\tau$, and
# - Matsubara frequency $i\omega$,
#
# for a given maximal energy scale $E_{max}$.
#
# Combined with the inverse temperature $\beta$ this gives the unitless cutoff parameter
#
# $$ \Lambda = \beta E_{max} $$
#
# that has to be supplied when constructing the grids, in combination with the required accuracy $\epsilon$.
# +
from pydlr import dlr
E_max, beta = 2., 10.
d = dlr(lamb=beta*E_max, eps=1e-10)
w_x = d.get_dlr_frequencies()
tau_l = d.get_tau(beta)
iw_q = d.get_matsubara_frequencies(beta)
pts = [(w_x, r'Real frequency DLR points $\omega_x$'),
(tau_l, r'Imaginary time DLR points $\tau_l$'),
(iw_q.imag, r'Matsubara frequency DLR points $i\omega_q$')]
subp = [3, 1, 1]
for val, label in pts:
plt.subplot(*subp); subp[-1] += 1;
plt.plot(val, 0*val, 'o'); plt.xlabel(label)
plt.tight_layout()
# -
# ## Dimer example
#
# A simple example that can be solved trivially as a two state system is the hybridized fermionic dimer system.
#
# The second quantized Hamiltonian $H$ is given by
#
# $$
# H = e_1 c_1^\dagger c_1 + e_2 c_2^\dagger c_2 + V ( c_1^\dagger c_2 + c_2^\dagger c_1) \, ,
# $$
#
# where $e_1$ and $e_2$ are the energies of the two levels and $V$ is their hybridization. In matrix form the Hamiltonian takes the form
#
# $$
# H =
# \left[\begin{array}{c} c_1^\dagger c_2^\dagger \end{array}\right]
# \cdot
# \left[\begin{array}{cc} e_1 & V \\ V & e_2 \end{array}\right]
# \cdot
# \left[\begin{array}{c} c_1 \\ c_2 \end{array}\right]
# \, .
# $$
#
# The single particle Green's function $G_{ab}(\tau)$ is a $2 \times 2$ matrix in imaginary time $\tau \in [0, \beta]$
#
# $$
# G_{ab}(\tau) \equiv - \langle c_a(\tau) c_b^\dagger(0) \rangle
# $$
#
# and can be computed with `pydlr` as:
# +
from pydlr import dlr
beta = 800.
e_1, e_2, V = 0.2, -0.5, 1.
h = np.array([
[e_1, V],
[V, e_2],])
d = dlr(lamb=2.*beta)
G_lab = d.free_greens_function_tau(h, beta)
tau_l = d.get_tau(beta)
# -
# In imaginary time the Green's function $G(\tau)$ is here represented using a carefully selected set of imaginary time nodes $\tau_l$.
#
# Equivalently there is a set of carefully selected Matsubara frequency points $i\omega_w$ on which the Green's function can be represented.
#
# The transform between these two spaces is done through the DLR coefficient representation at real-frequencies $\omega_x$.
#
# $$
# \text{Imaginary time } G(\tau_l) \leftrightarrow \text{DLR coefficients } G_x \leftrightarrow \text{Matsubara frequency } G(i\omega_w)
# $$
#
#
# +
# Transform from imaginary time tau to DLR coefficients
G_xab = d.dlr_from_tau(G_lab)
# Transform DLR to imaginary time tau
G_lab_ref = d.tau_from_dlr(G_xab)
np.testing.assert_array_almost_equal(G_lab, G_lab_ref)
# Trasform from DLR to Matsubara frequencies
G_wab = d.matsubara_from_dlr(G_xab, beta)
w = d.get_matsubara_frequencies(beta)
# -
# Since the DLR coefficient representation gives the imaginary time representation as
#
# $$
# G(\tau) = \sum_{x} K_\beta (\tau, \omega_x) G_x
# $$
#
# where $K_\beta$ is the analytical continuation kernel at inverse temperature $\beta$.
#
# Thus the DLR coefficients can be used to evaluate $G(\tau)$ on any grid in imaginary time $\tau \in [0, \beta]$.
# +
# Evaluate on arbitrary tau grid
tau_i = np.linspace(0, beta, num=1000)
G_iab = d.eval_dlr_tau(G_xab, tau_i, beta)
# +
# Imaginary-time plot
subp = [2, 2, 1]
for a, b in itertools.product(range(2), repeat=2):
plt.subplot(*subp); subp[-1] += 1
plt.plot(tau_i, G_iab[:, a, b], '-', label=r'Arb $\tau$-grid', alpha=0.5)
plt.plot(tau_l, G_lab[:, a, b], '.', label='DRL points', alpha=0.5)
plt.ylabel(r'$G_{' + f'{a+1},{b+1}' + r'}(\tau)$')
plt.xlabel(r'$\tau$')
plt.legend(loc='best');
plt.tight_layout()
plt.show()
# -
# Equivalently the DLR representation can be evaluated at arbitrary points in frequency.
w_W = 1.j * np.linspace(-20, 20, num=400)
G_Wab = d.eval_dlr_freq(G_xab, w_W, beta)
# +
# Matsubara frequency plot
subp = [2, 2, 1]
for a, b in itertools.product(range(2), repeat=2):
plt.subplot(*subp); subp[-1] += 1
plt.plot(w.imag, G_wab[:, a, b].real, '.', label='DLR Re', alpha=0.5)
plt.plot(w.imag, G_wab[:, a, b].imag, '.', label='DLR Im', alpha=0.5)
plt.plot(w_W.imag, G_Wab[:, a, b].real, '-', label='Interp Re', alpha=0.5)
plt.plot(w_W.imag, G_Wab[:, a, b].imag, '-', label='Interp Im', alpha=0.5)
plt.ylabel(r'$G_{' + f'{a+1},{b+1}' + r'}(i\omega_n)$')
plt.xlabel(r'$\omega_n$')
plt.legend(loc='best', fontsize=7);
plt.tight_layout()
plt.show()
# -
# ## Dyson equation
#
# The dimer example can also be used as a simple application of the Dyson equation.
#
# For example the 1,1 component of the Green's function
#
# $$G_{1,1}(\tau) = G(\tau)$$
#
# can also be obtained from a system with single particle Hamiltonian
#
# $$h_1 = [e_1]$$
#
# and the self energy
#
# $$\Sigma(\tau) = V^2 \, g_{e_{2}}(\tau)$$
#
# where $g_{e_{2}}(\tau)$ is the free Green's function for a single energy level $h_2 = [e_2]$.
# +
h1 = np.array([[e_1]])
sigma_l = V**2 * d.free_greens_function_tau(np.array([[e_2]]), beta)
# -
# ### Matsubara Dyson Equation
#
# The Dyson equation for the single-particle Green's function $G$ is given by
#
# $$
# G(i\omega_n) = \left[ i\omega_n - h - \Sigma(i\omega_n) \right]^{-1}
# $$
#
# where $\omega_n$ are the Matsubara frequencies, $h$ is the single particle Hamiltonian and $\Sigma$ is the self-energy.
# +
sigma_x = d.dlr_from_tau(sigma_l)
sigma_w = d.matsubara_from_dlr(sigma_x, beta)
G_w = d.dyson_matsubara(h1, sigma_w, beta)
G_l_matsubara = d.tau_from_dlr(d.dlr_from_matsubara(G_w, beta))
np.testing.assert_array_almost_equal(G_l_matsubara, G_lab[:, 0:1, 0:1])
# -
# ### Imaginary time Dyson equation
#
# An equivalent formulation in imaginary time is given by the integro-differential form
#
# $$
# (-\partial_\tau - h - \Sigma \, \ast ) G(\tau) = 0
# $$
#
# with the boundary condition $G(0) - \xi G(\beta) = -1$ and the imaginary time convolution
#
# $$
# \Sigma \ast G \equiv \int_{0}^\beta d\bar{\tau} \,
# \Sigma(\tau - \bar{\tau}) G(\bar{\tau})
# $$
# +
G_x = d.dyson_dlr_integrodiff(h1, sigma_x, beta)
G_l_integrodiff = d.tau_from_dlr(G_x)
np.testing.assert_array_almost_equal(G_l_integrodiff, G_lab[:, 0:1, 0:1])
# -
# This can further be rewritten using the free Green's function $g$ defined as
#
# $$
# (-\partial_\tau - h ) g(\tau) = 0
# $$
#
# multiplying the integro-differential form from the left gives the Dyson equation in the integral formulation
#
# $$
# (1 + g \ast \Sigma \, \ast) \, G = g
# $$
# +
G_l_integro = d.tau_from_dlr(d.dyson_dlr(h1, sigma_x, beta))
np.testing.assert_array_almost_equal(G_l_integro, G_lab[:, 0:1, 0:1])
# -
# While it is possible to solve the Dyson equation in all these ways the numerically most accurate is the last integral formulation in DLR space, as can be seen when comparing the errors.
# +
error = lambda G_l : np.max(np.abs(G_l - G_lab[:, 0:1, 0:1]))
print(f'Error {error(G_l_matsubara):2.2E} Matsubara')
print(f'Error {error(G_l_integrodiff):2.2E} integrodiff')
print(f'Error {error(G_l_integro):2.2E} integro')
# -
# ## Non-linear problems
#
# While the dimer system can be used to test the Dyson equation, it is more common to use it in problems where the self-energy $\Sigma$ is a functional of the Green's function itself
#
# $$
# \Sigma = \Sigma[G]
# $$
#
# Combined with the Dyson equation this gives the non-linear integral equation problem
#
# $$
# (1 + g \ast \Sigma[G] \, \ast ) \, G = g
# $$
#
# Here follows a few simple examples of non-linear problems.
# ### The semi infinite chain (or the Bethe graph)
#
# The Green's function $G$ of the last site in the semi-infinite chain with onsite energy $e$ and nearest-neightbour hopping $t$ can be determined recursively to obey the Dyson equation
#
# $$
# \left( -\partial_\tau - h - \frac{t^2}{4} G \, \ast \right) \, G = 0
# $$
#
# This equation can be solved analytically and be shown to correspond to a system with the semi-circular density of states
#
# $$
# \rho(\omega) = \frac{2}{\pi t^2} \sqrt{\left(\omega + t - h\right)\left(t + h - \omega\right)}
# $$
#
# whoose imaginary time Green's function is given by
#
# $$
# G(\tau) = -\int_{-\infty}^\infty K_\beta(\tau, \omega) \rho(\omega) \, d\omega
# $$
#
# +
def dos(x, h, t):
a = -t + h
b = +t + h
return 2./np.pi/t**2 * np.sqrt((x - a) * (b - x))
h = 0.0
t = 1.
x = np.linspace(-t + h, +t + h, num=1000)
rho = dos(x, h, t)
norm = np.trapz(rho, x=x)
print(f'norm = {norm}')
plt.figure()
plt.plot(x, rho)
plt.xlabel(r'$\omega$')
plt.ylabel(r'$\rho(\omega)$')
plt.show()
# -
# Since the density of states is known the imaginary time Green's function can be computed semi-analytically through the evaluation of the real-frequency integral.
#
# The integral can be computed to machine prescision using `scipu.integrate.quad` when accounting for the square root singularities using the `alg` weight function, see
#
# <https://docs.scipy.org/doc/scipy/reference/generated/scipy.integrate.quad.html>
# +
from pydlr import kernel
from scipy.integrate import quad
def eval_semi_circ_tau(tau, beta, h, t):
I = lambda x : -2 / np.pi / t**2 * kernel(np.array([tau])/beta, beta*np.array([x]))[0,0]
g, res = quad(I, -t+h, t+h, weight='alg', wvar=(0.5, 0.5))
return g
eval_semi_circ_tau = np.vectorize(eval_semi_circ_tau)
beta = 100.
tau_l = d.get_tau(beta)
G_l = eval_semi_circ_tau(tau_l, beta, h, t)
# -
# Finally we solve the non-linear integrodifferential problem
#
# $$
# \left( -\partial_\tau - h - \frac{t^2}{4} G \, \ast \right) \, G = 0
# $$
#
# using forward iteration and compare to the semi-analytical result.
# +
tol = 1e-15
max_iter = 1000
G_x = np.zeros((len(tau_l), 1, 1))
G_l_iter = np.zeros_like(G_x)
for iter in range(max_iter):
G_x = d.dyson_dlr(np.array([[h]]), 0.25 * t**2 * G_x, beta)
G_l_iter_new = d.tau_from_dlr(G_x)
diff = np.max(np.abs(G_l_iter - G_l_iter_new))
G_l_iter = G_l_iter_new
if diff < tol:
print(f'Converged in {iter+1} iterations.')
break
print(f'Error {np.max(np.abs(G_l_iter[:, 0, 0] - G_l)):2.2E} rel diff {diff:2.2E}')
plt.figure()
plt.plot(tau_l, -G_l, label='semi-analytic')
plt.plot(tau_l, -np.squeeze(G_l_iter), '.', label='dyson')
plt.grid(True)
plt.legend(loc='best')
plt.ylabel(r'$-G(\tau)$')
plt.xlabel(r'$\tau$')
plt.show()
# -
# ### SYK-model example
#
# The SYK model is given by the self-consistent second-order self energy
#
# $$
# \Sigma(\tau) = J^2 (G(\tau))^2 G(\beta - \tau)
# \, ,
# $$
#
# here the evaluation at $\beta - \tau$ can be obtained using the evaluation of DLR coefficients at arbitrary points in imaginary time.
def sigma_x_syk(g_x, J, d, beta):
tau_l = d.get_tau(beta)
tau_l_rev = beta - tau_l
g_l = d.tau_from_dlr(g_x)
g_l_rev = d.eval_dlr_tau(g_x, tau_l_rev, beta)
sigma_l = J**2 * g_l**2 * g_l_rev
sigma_x = d.dlr_from_tau(sigma_l)
return sigma_x
# The asymptotic conformal solution $G_c(\tau)$ is given by
#
# $$
# G_c(\tau) = - \frac{\pi^{1/4}}{\sqrt{2\beta}} \left( \sin \left( \frac{\pi \tau}{\beta} \right)\right)^{-1/2}
# $$
def conformal_tau(tau, beta):
return -np.pi**0.25 / np.sqrt(2 * beta) * 1./np.sqrt(np.sin(np.pi * tau/beta))
# Finally we solve the non-linear integral equation
#
# $$
# (1 + g \ast \Sigma[G] \, \ast ) \, G = g
# $$
#
# using a root solver and the `df-sane` algorithm from `scipy.optimize.root`.
def solve_syk_root(d, mu, beta=1., J=1.0, g0_l=None, tol=1e-16, verbose=False):
if verbose:
print('='*72)
print('SYK root DLR solver')
print('-'*72)
print(f'mu = {mu}, J = {J}, beta = {beta}')
print(f'lamb = {lamb}, n_dlr = {len(d)}')
print(f'tol = {tol}')
print('='*72)
if g0_l is not None: g_l = g0_l[:, 0, 0]
else: g_l = d.free_greens_function_tau(np.array([[mu]]), beta)[:, 0, 0]
def target_function(g_l):
g_l = g_l.reshape((len(g_l), 1, 1))
sigma_x = sigma_x_syk(d.dlr_from_tau(g_l), J, d, beta)
g_x_new = d.dyson_dlr(np.array([[mu]]), sigma_x, beta)
g_l_new = d.tau_from_dlr(g_x_new)
return np.squeeze((g_l - g_l_new).real)
from scipy.optimize import root
sol = root(target_function, g_l*0, method='df-sane', tol=tol, options={'maxfev':10000})
diff = np.max(np.abs(target_function(sol.x)))
if verbose: print(f'nfev = {sol.nfev}, diff = {diff}')
g_l = sol.x.reshape((len(g_l), 1, 1))
return g_l
# +
J = 1.
mu0 = 0.
beta = 1.e4
lamb = beta * 5
tol = 1e-13
d = dlr(lamb=lamb)
tau_l = d.get_tau(beta)
g_l_root = solve_syk_root(d, mu0, beta=beta, J=J, tol=tol, verbose=True)
g_l_conformal = conformal_tau(tau_l, beta)
# -
plt.plot(tau_l, -np.squeeze(g_l_root), label='SYK')
plt.plot(tau_l, -g_l_conformal, '--', label='conformal')
plt.semilogy([], [])
plt.ylim([5e-3, 1.])
plt.ylabel(r'$-G(\tau)$')
plt.xlabel(r'$\tau$')
plt.legend(loc='best')
plt.grid(True)
| doc/python_examples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This script trains and select models for this problem
# load the packages
import pandas as pd
import numpy as np
from sklearn.preprocessing import OneHotEncoder
from sklearn.model_selection import GridSearchCV
from sklearn.preprocessing import StandardScaler
from sklearn.preprocessing import LabelEncoder
from sklearn.model_selection import train_test_split
from sklearn.model_selection import StratifiedKFold
from sklearn.metrics import accuracy_score
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import RandomForestClassifier
from sklearn.svm import SVC
from sklearn.compose import ColumnTransformer
from sklearn.pipeline import Pipeline
from sklearn.pipeline import make_pipeline
from sklearn.metrics import make_scorer
from sklearn.impute import SimpleImputer
import matplotlib.pyplot as plt
import matplotlib
df = pd.read_csv('../data/processed_sleep_fft.csv')
cat_ftrs = ['channel_name']
scalar_ftrs = ['alpha', 'theta', 'slowwave', 'sigma']
le = LabelEncoder()
y = le.fit_transform(df['label'])
subject_ID = df['subject']
nap_ID = df['NAP']
dropc = ['Unnamed: 0', 'label', 'subject', 'NAP']
X = df.drop(columns= dropc)
# check balance
classes, counts = np.unique(y, return_counts=True)
for i in range(len(classes)):
print ('balance', i, counts[i]/ len(y))
# encode groups 4 subject ID * 2 naps
import itertools
n_subject = np.unique(subject_ID)
n_nap = np.unique(nap_ID)
iterset = (list(itertools.product(n_subject, n_nap)))
group = np.zeros((len(subject_ID), 1))
i = 0
for sbj, nap in iterset:
idx = np.logical_and(subject_ID == sbj, nap_ID== nap)
group[idx] = i
i+= 1
# +
# from sklearn.model_selection import GroupKFold
# from sklearn.model_selection import GroupShuffleSplit
# def ML_pipeline_groups_GridSearchCV_SVC(X,y,groups,random_state,n_folds):
# # create a test set based on groups
# splitter = GroupShuffleSplit(n_splits=1,test_size=0.2,random_state=random_state)
# for i_other,i_test in splitter.split(X, y, groups):
# X_other, y_other, groups_other = X.iloc[i_other], y[i_other], groups[i_other]
# X_test, y_test, groups_test = X.iloc[i_test], y[i_test], groups[i_test]
# # splitter for _other
# kf = GroupKFold(n_splits=n_folds)
# # create the pipeline: preprocessor + supervised ML method
# cat_ftrs = ['channel_name']
# cont_ftrs = ['alpha', 'theta', 'slowwave', 'sigma']
# cat_transformer = Pipeline(steps = [
# ('imputer1', SimpleImputer(missing_values='0.0', strategy='constant',fill_value='missing')),
# ('onehot', OneHotEncoder(sparse = False, categories = 'auto'))])
# cont_transformer = Pipeline(steps = [
# ('imputer2', SimpleImputer(missing_values = np.nan,strategy = 'mean')),
# ('scaler', StandardScaler())])
# preprocessor = ColumnTransformer(remainder='passthrough',
# transformers=[
# ('num', cont_transformer, cont_ftrs),
# ('cat', cat_transformer, cat_ftrs)])
# # make overall pipeline
# pipe = make_pipeline(
# preprocessor,
# SVC(probability = True, max_iter = 1000))
# # the parameter(s) we want to tune
# param_grid = {'svc__C': np.logspace(-2,2,num=5),'svc__gamma': np.logspace(-2,2,num=5)}
# # prepare gridsearch
# grid = GridSearchCV(pipe, param_grid=param_grid,scoring = make_scorer(accuracy_score),
# cv=kf, return_train_score = True,iid=True, n_jobs = -1)
# # do kfold CV on _other
# grid.fit(X_other, y_other, groups_other)
# return grid, grid.score(X_test, y_test)
# +
# test_scores_SVC = []
# for i in range(5):
# print('Starting:', i, 'process')
# grid, test_score = ML_pipeline_groups_GridSearchCV_SVC(X,y,group,i*42,2)
# print(grid.best_params_)
# print('best CV score:',grid.best_score_)
# print('test score:',test_score)
# test_scores_SVC.append(test_score)
# print('test accuracy:',np.around(np.mean(test_scores_SVC),2),'+/-',np.around(np.std(test_scores_SVC),2))
# -
from sklearn.model_selection import GroupKFold
from sklearn.model_selection import GroupShuffleSplit
def ML_pipeline_groups_GridSearchCV_Logistic(X,y,groups,random_state,n_folds):
# create a test set based on groups
splitter = GroupShuffleSplit(n_splits=1,test_size=0.2,random_state=random_state)
for i_other,i_test in splitter.split(X, y, groups):
X_other, y_other, groups_other = X.iloc[i_other], y[i_other], groups[i_other]
X_test, y_test, groups_test = X.iloc[i_test], y[i_test], groups[i_test]
# splitter for _other
kf = GroupKFold(n_splits=n_folds)
# create the pipeline: preprocessor + supervised ML method
cat_ftrs = ['channel_name']
cont_ftrs = ['alpha', 'theta', 'slowwave', 'sigma']
cat_transformer = Pipeline(steps = [
('imputer1', SimpleImputer(missing_values='0.0', strategy='constant',fill_value='missing')),
('onehot', OneHotEncoder(sparse = False, categories = 'auto'))])
cont_transformer = Pipeline(steps = [
('imputer2', SimpleImputer(missing_values = np.nan,strategy = 'mean')),
('scaler', StandardScaler())])
preprocessor = ColumnTransformer(remainder='passthrough',
transformers=[
('num', cont_transformer, cont_ftrs),
('cat', cat_transformer, cat_ftrs)])
# make overall pipeline
pipe = make_pipeline(
preprocessor,
LogisticRegression(penalty = 'l1', solver = 'saga', max_iter = 1000, multi_class = 'multinomial'))
# the parameter(s) we want to tune
param_grid = {'logisticregression__C': np.logspace(-2,2,num=5)}
# prepare gridsearch
grid = GridSearchCV(pipe, param_grid=param_grid,scoring = make_scorer(accuracy_score),
cv=kf, return_train_score = True,iid=True, n_jobs = -1)
# do kfold CV on _other
grid.fit(X_other, y_other, groups_other)
return grid, grid.score(X_test, y_test)
test_scores_logistic = []
for i in range(5):
print('Starting:', i, 'process')
grid, test_score = ML_pipeline_groups_GridSearchCV_Logistic(X,y,group,i*42,2)
print(grid.best_params_)
print('best CV score:',grid.best_score_)
print('test score:',test_score)
test_scores_logistic.append(test_score)
print('test accuracy:',np.around(np.mean(test_scores_logistic),2),'+/-',np.around(np.std(test_scores_logistic),2))
from sklearn.model_selection import GroupKFold
from sklearn.model_selection import GroupShuffleSplit
def ML_pipeline_groups_GridSearchCV_RandomForest(X,y,groups,random_state,n_folds):
# create a test set based on groups
splitter = GroupShuffleSplit(n_splits=1,test_size=0.2,random_state=random_state)
for i_other,i_test in splitter.split(X, y, groups):
X_other, y_other, groups_other = X.iloc[i_other], y[i_other], groups[i_other]
X_test, y_test, groups_test = X.iloc[i_test], y[i_test], groups[i_test]
# splitter for _other
kf = GroupKFold(n_splits=n_folds)
# create the pipeline: preprocessor + supervised ML method
cat_ftrs = ['channel_name']
cont_ftrs = ['alpha', 'theta', 'slowwave', 'sigma']
cat_transformer = Pipeline(steps = [
('imputer1', SimpleImputer(missing_values='0.0', strategy='constant',fill_value='missing')),
('onehot', OneHotEncoder(sparse = False, categories = 'auto'))])
cont_transformer = Pipeline(steps = [
('imputer2', SimpleImputer(missing_values = np.nan,strategy = 'mean')),
('scaler', StandardScaler())])
preprocessor = ColumnTransformer(remainder='passthrough',
transformers=[
('num', cont_transformer, cont_ftrs),
('cat', cat_transformer, cat_ftrs)])
# make overall pipeline
pipe = make_pipeline(
preprocessor,
RandomForestClassifier(random_state= random_state))
# specify parameters
param_grid = {'randomforestclassifier__max_depth' : np.logspace(0, 3, num=5),
'randomforestclassifier__min_samples_split' : np.linspace(2, 100, num = 5, dtype = int)}
# prepare gridsearch
grid = GridSearchCV(pipe, param_grid=param_grid,scoring = make_scorer(accuracy_score),
cv=kf, return_train_score = True,iid=True, n_jobs = -1)
# do kfold CV on _other
grid.fit(X_other, y_other, groups_other)
return grid, grid.score(X_test, y_test)
test_scores_randomForest = []
for i in range(5):
print('Starting:', i, 'process')
grid, test_score = ML_pipeline_groups_GridSearchCV_RandomForest(X,y,group,i*42,2)
print(grid.best_params_)
print('best CV score:',grid.best_score_)
print('test score:',test_score)
test_scores_randomForest.append(test_score)
print('test accuracy:',np.around(np.mean(test_scores_randomForest),2),'+/-',np.around(np.std(test_scores_randomForest),2))
from sklearn.model_selection import GroupKFold
from sklearn.model_selection import GroupShuffleSplit
from sklearn.model_selection import ParameterGrid
from xgboost import XGBClassifier
def ML_pipeline_groups_GridSearchCV_XGboost(X,y,groups,random_state,n_folds):
# create a test set based on groups
splitter = GroupShuffleSplit(n_splits=1,test_size=0.2,random_state=random_state)
for i_other,i_test in splitter.split(X, y, groups):
X_other, y_other, groups_other = X.iloc[i_other], y[i_other], groups[i_other]
X_test, y_test, groups_test = X.iloc[i_test], y[i_test], groups[i_test]
# splitter for _other
kf = GroupKFold(n_splits=n_folds)
# create the pipeline: preprocessor + supervised ML method
cat_ftrs = ['channel_name']
cont_ftrs = ['alpha', 'theta', 'slowwave', 'sigma']
cat_transformer = Pipeline(steps = [
('imputer1', SimpleImputer(missing_values='0.0', strategy='constant',fill_value='missing')),
('onehot', OneHotEncoder(sparse = False, categories = 'auto'))])
cont_transformer = Pipeline(steps = [
('scaler', StandardScaler())])
preprocessor = ColumnTransformer(remainder='passthrough',
transformers=[
('num', cont_transformer, cont_ftrs),
('cat', cat_transformer, cat_ftrs)])
# make overall pipeline
pipe = make_pipeline(
preprocessor,
XGBClassifier(seed = random_state))
# specify parameters
param_grid = {"xgbclassifier__reg_alpha":np.logspace(-2,2,num=5) }
# prepare gridsearch
grid = GridSearchCV(pipe, param_grid=(param_grid),scoring = make_scorer(accuracy_score),
cv=kf, return_train_score = True,iid=True, n_jobs = -1)
# do kfold CV on _other
grid.fit(X_other, y_other, groups_other)
return grid, grid.score(X_test, y_test)
test_scores_xgboost = []
for i in range(5):
print('Starting:', i, 'process')
grid, test_score = ML_pipeline_groups_GridSearchCV_XGboost(X,y,group,i*42,2)
print(grid.best_params_)
print('best CV score:',grid.best_score_)
print('test score:',test_score)
test_scores_xgboost.append(test_score)
print('test accuracy:',np.around(np.mean(test_scores_xgboost),2),'+/-',np.around(np.std(test_scores_xgboost),2))
test_scores_xgboost_2 = []
for i in range(5):
print('Starting:', i, 'process')
grid, test_score = ML_pipeline_groups_GridSearchCV_XGboost(X,y,group,i*42,5)
print(grid.best_params_)
print('best CV score:',grid.best_score_)
print('test score:',test_score)
test_scores_xgboost_2.append(test_score)
print('test accuracy:',np.around(np.mean(test_scores_xgboost_2),2),'+/-',np.around(np.std(test_scores_xgboost_2),2))
| src/ML_pipeline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import csv
import pandas as pd
import requests
from bs4 import BeautifulSoup, SoupStrainer
import re
import pandas as pd
import json
import time
bussiness_licences = "business-licences-hackathon.csv"
file = open(bussiness_licences, "r")
reader = pd.read_csv(file, sep=';')
# -
URL = 'https://beta.canadasbusinessregistries.ca/search/results?search=%7Baaa%7D&location=BC&status=Active'
#tree = BeautifulSoup(URL)
#good_html = tree.prettify()
page = requests.get(URL)
# +
import urllib.request as urllib2
import json
# new url
url = 'https://beta.canadasbusinessregistries.ca/search/results?search=%7Baaa%7D&location=BC&status=Active'
# read all data
#page = urllib2.urlopen(url).read()
data = json.loads(url)
connection = urllib2.urlopen(url)
js = connection.read()
#requests.get(url).json()
# convert json text to python dictionary
#data = json.loads(page)
info = json.loads(js.decode("utf-8"))
print(info)
# -
from itertools import product
from string import ascii_lowercase
keywords = [''.join(i) for i in product(ascii_lowercase, repeat = 3)]
keywords[0:5]
url = 'https://searchapi.mrasservice.ca/Search/api/v1/search?fq=keyword:%7B%22aaa%22&location=BC&lang=en&queryaction=fieldquery&sortfield=score&sortorder=desc'
response = requests.get(url)
content_json = response.json()
for ngram in keywords[0:5]:
url = 'https://searchapi.mrasservice.ca/Search/api/v1/search?fq=keyword:%7B%22' + ngram + '%22&location=BC&lang=en&queryaction=fieldquery&sortfield=score&sortorder=desc'
response = requests.get(url)
content_json = response.json()
time.sleep(3)
content_json['docs']
df
df = pd.DataFrame(content_json['docs'])
df.columns
df.drop(columns=['Registry_Source','version', 'Juri_ID','Data_Source',
'text','Alternate_Name', 'id', '_version_', 'hierarchy',
'parent_id', 'Reg_date_XPR_juri',
'HJ_country', 'HJ_ID','HJ_entity_ID','HJ_entity_name'])#.dropna()
content_json['docs']
soup = BeautifulSoup(page.content, 'html.parser')
#results = soup.find('list-unstyled organization__info')
results = soup.find('div', class_="primary-card__info")
#results = soup.find(id='wb-cont')
print(results.prettify())
#business_names = results.find_all('span', class_='info info--business-number')
for business in business_names:
print(business)
| scraping_rachel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Analysis for the Modeling Game
# +
# %matplotlib inline
import copy
import lmfit
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from scipy.integrate import odeint
import constants as cn
import tellurium as te
import gene_network as gn
import modeling_game as mg
import model_fitting as mf
import gene_analyzer as ga
import run_game as rg
import util
# -
# ## Stress Tests
#
# - Decrease Vm8 by 50% and increase Vm3 by 50%
# - Show P1
# +
def simulate(parameters=None):
"""
Runs the simulation for the default model file.
:return pd.Series, pd.Series, RoadRunner: time, P1, road runner instance
"""
model = rg.getModel()
result = mf.runSimulation(model=model,
parameters=parameters,
sim_time=1200, num_points=120)
df_protein = mg.makeDF(result.data, is_mrna=False)
return df_protein.index, df_protein["P1"], result.road_runner
def getP1Data(is_adjust=False):
"""
Obtains P1 data and time information.
:param bool is_adjust: if True, perturb Vm8, Vm3
:return pd.Series, pd.Series: time, P1
"""
_, _, road_runner = simulate()
if is_adjust:
perturb = 0.5
else:
perturb = 0.0
vm8 = perturb*road_runner.Vm8
vm3 = (1 + perturb)*road_runner.Vm3
parameters = mg.makeParameters(['Vm8', 'Vm3'], [vm8, vm3])
ser_time, ser_p1, road_runner = simulate(parameters=parameters)
print("Vm3: %f\nVm8: %f" % (road_runner.Vm3, road_runner.Vm8))
return ser_time, ser_p1
# -
ser_time, ser_p1 = getP1Data()
plt.plot(ser_time, ser_p1)
# Fails the stress test since no shift in the peak. See "Final_stress_tests.pptx"
ser_time, ser_p1 = getP1Data(is_adjust=True)
plt.plot(ser_time, ser_p1)
# ## Model Identification
#
analyzers = []
analyzers.append(evaluate(["1+4"], max_iteration=20, start_time=0, end_time=1200))
analyzers.append(evaluate(["2+4"], max_iteration=20, start_time=0, end_time=1200))
analyzers.append(evaluate(["3+6"], max_iteration=20, start_time=0, end_time=1200))
analyzers.append(evaluate(["4-2A-5"], max_iteration=20, start_time=0, end_time=1200))
analyzers.append(evaluate(["5+6"], max_iteration=20, start_time=0, end_time=1200))
analyzers.append(evaluate(["6+7A-1"], max_iteration=20, start_time=0, end_time=1200))
analyzers.append(evaluate(["7-8"], max_iteration=20, start_time=0, end_time=1200))
analyzers.append(evaluate(["8-1"], max_iteration=30, start_time=0, end_time=1200))
# ## Saving an Identified Simulation Model
# Builds a complete simulation model from the separate analyzers.
# Saves the models and parameters to files
df_params, model = rg.saveAnalysisResults(analyzers)
df_params
print(model)
# ## Running a Saved Model
# Reads the model and parameters from a file. Fits the parameters. Plots the results.
rg.runModel()
# ## Correlation Analysis
# The goal here is to gain intuition about possible TF configurations for genes.
#
# 1. Construct cross correlatons between mRNA and proteins at different time lags. Plt as heatmap.
df_mrna = pd.read_csv("wild.csv")
df_mrna = df_mrna.set_index("time")
df_protein = pd.read_csv("wild_protein.csv")
df_protein = df_protein.set_index("time")
def correlate(df1, df2, lag=0):
"""
Constructs the correlation with the specified lag
for df2. Assumes that df1 and df2 have different columns.
"""
def truncate(df, is_front=True):
if is_front:
df_trunc = df.loc[df.index[lag:], :]
else:
if lag == 0:
df_trunc = df
else:
df_trunc = df.loc[df.index[:-lag], :]
df_trunc.index = range(len(df_trunc))
return df_trunc
# Adjust lengths
indices = range(len(df1) - lag)
df1_trunc = truncate(df1, is_front=False)
df2_trunc = truncate(df2, is_front=True)
df_tot = pd.concat([df1_trunc, df2_trunc], axis=1)
# Correlate and select columns
df_corr = df_tot.corr()
columns2 = df2.columns
df_result = df_corr[columns2].copy()
df_result = df_result.drop(columns2)
return df_result
correlate(df_protein, df_mrna, lag=0)
correlate(df_protein, df_mrna, lag=1)
correlate(df_protein, df_mrna, lag=2)
correlate(df_protein, df_mrna, lag=3)
| archived_lectures/ModelingGame/game_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/DSNortsev/CSE590-PythonAndDataAnalytics/blob/main/HW2/HW2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="QYAkwgbsHGKl"
from collections import OrderedDict
import numpy as np
import pandas as pd
import re
# + [markdown] id="Rx1Bmt9vKILf"
# <b>This assignment deals with loading a simple text file into a Python structure, lists, arrays, and dataframes.</b>
#
# <b>a. Locate a movie script, play script, poem, or book of your choice in .txt format*. Project Gutenburg is a great resource for this if you're not sure where to start.</b>
#
# <b>b. Load the words of this structure, one-by-one, into a one-dimensional, sequential Python list (i.e. the first word should be the first element in the list, while the last word should be the last element). It's up to you how to deal with special chacters -- you can remove them manually, ignore them during the loading process, or even count them as words, for example.</b>
# + colab={"base_uri": "https://localhost:8080/"} id="egu22DuCPct0" outputId="b71bf33d-6c6a-45bc-c35d-85d4e2fa0c58"
def load_data(data_file):
"""
Reads txt file and returns a list of words
"""
# Compile regex pattern
regex_pattern = re.compile('[^A-Za-z0-9.:/-]+')
# Read txt file
with open(data_file) as f:
# Find all special characters in the word and replace it with empty string
# Remove leading and trailing special characters
return [re.sub(regex_pattern, '', word).lower().strip('.:/') for line in f for word in line.split()]
words_list = load_data('the_martian_circe.txt')
print(words_list[:100])
# + [markdown] id="78Bzx91XKQJv"
# <b>c. Use your list to create and print a two-column pandas data-frame with the following properties: i. Each index should mark the first occurrence of a unique word (independent of case) in the text. ii. The first column for each index should represent the word in question at that index iii. The second column should represent the number of times that particular word appears in the text.</b>
# + colab={"base_uri": "https://localhost:8080/", "height": 424} id="KxfjI1C2KUcq" outputId="7b8fc181-475a-4e6d-a934-8f2034e3a96a"
def count_elements(list_of_elements):
"""
Count words occurance with preserved possition
"""
data = OrderedDict()
for element in list_of_elements:
if element not in data:
data[element] = 1
else:
data[element] += 1
return data
counted_occurence = count_elements(words_list)
df = pd.DataFrame(counted_occurence.items(), columns=['Word', 'Count'])
df
# + [markdown] id="gIE3pwXVKg1E"
# <b> d. The co-occurrence of two events represents the likelihood of the two occurring together. A simple example of co-occurrence in texts is a predecessor-successor relationship -- that is, the frequency with which one word immediately follows another. The word "cellar," for example, is commonly followed by "door." </b>
#
# For this task, you are to construct a 2-dimensional predecessor-successor co-occurrence array as follows**: i. The row index corresponds to the word from the same index in part c.'s data-frame. ii. The column index likewise corresponds to the word in the same index in the data-frame. iii. The value in each array location represents the count of the number of times the word corresponding to the row index immediately precedes the word correponding to the column index in the text.
# + colab={"base_uri": "https://localhost:8080/"} id="wvXSaFH3T05q" outputId="f22ff637-f423-42a0-c899-b4a31fcfe340"
def generate_co_occurance_matrix(words_list, columns):
"""
Generates co-occurance matrix:
word_list: a list of words in the text
columns: unique list of words
"""
# Convert words list to numpy array
words_array = np.array(words_list)
# Convert columns list to numpy array
columns_array = np.array(columns)
# Generate zero value matrix with integer value
matrix = np.zeros((len(columns), len(columns)), dtype=np.int)
count = 0
# Iterate over unique words
for word in columns_array:
# find row position in matrix for the word
row_position = np.where(columns_array == word)[0][0]
# Find all occurances of this word in the list and iterate over it
for word_position in np.where(words_array == word)[0]:
if word_position < len(words_array) - 1:
# Find position of the successor word
col_position = np.where(columns_array == str(words_array[word_position + 1]))[0][0]
# Incremenet predecessor-successor co-occurrence by one
matrix[row_position, col_position] += 1
return matrix
matrix = generate_co_occurance_matrix(words_list, list(counted_occurence.keys()))
matrix
# + [markdown] id="Ya8ljAYHLE43"
# <b>e. Based on the data-frame derived in part c. and array derived in part d., determine and print the following information:<br></br>
# i. The first occurring word in the text. </b>
# + colab={"base_uri": "https://localhost:8080/", "height": 36} id="xXXg1c8GLdxr" outputId="a24d171d-8f12-4d68-eda5-1e83805c3b8f"
df['Word'].iloc[0]
# + [markdown] id="kD5jeilZLRB3"
# <b>ii. The unique word that first occurs last within the text. </b>
# + colab={"base_uri": "https://localhost:8080/", "height": 36} id="5BlExGSpMkgE" outputId="279c1bfb-1c08-4040-afb7-09ba929d4a12"
df['Word'].iloc[-1]
# + [markdown] id="rnb2WyNALlUB"
# <b>iii. The most common word </b>
# + colab={"base_uri": "https://localhost:8080/", "height": 36} id="zo60FyyYM5yS" outputId="bc0873e5-0b6c-45eb-cd88-626d680b716e"
df[df['Count'] == df['Count'].max()]['Word'].iloc[0]
# + [markdown] id="psgz2QERM09H"
# <b> v. Words A and B such that B follows A more than any other combination of words.</b>
# + colab={"base_uri": "https://localhost:8080/"} id="4XCfzXs5M3lD" outputId="1440e916-0fcd-40b5-92fa-5832df130916"
# Find max value in matrix
max_occurence = np.amax(matrix)
# Find positions of max value
position = np.where(matrix == max_occurence)
print(df['Word'].iloc[position[0][0]], df['Word'].iloc[position[1][0]])
# + [markdown] id="Cs4lIl93MtLL"
# <b>vi. The word that most commonly follows the least common word </b>
# + colab={"base_uri": "https://localhost:8080/", "height": 424} id="RVJ_svJIMlfK" outputId="a4cf9496-a585-4146-dfa0-6b5588d7230a"
def find_most_common_word_follows_least(df, matrix):
# Find least common words
least_common_words_index = df.index[df['Count'] == df['Count'].min()].tolist()
# Create empty PandaFrame
result = pd.DataFrame(data = [], columns=['Predecessor', 'Successor',
'Occurence_predecessor','Occurence_successor' ])
# Iterate over least common words
for column_pos in least_common_words_index:
# Get one dimension matrix for that word
tmp_array = matrix[:,column_pos]
# Find all occurence with greater than 0
positive_occurence_list = tmp_array[tmp_array>0]
# Iterate over all occurence
for occurence in positive_occurence_list:
# Find index of the element
row_pos = np.where(tmp_array == occurence)[0][0]
# Generate new row
new_row = [df.iloc[row_pos]['Word'], df.iloc[column_pos]['Word'],
df.iloc[row_pos]['Count'], df.iloc[column_pos]['Count']]
# Append it to result
result.loc[len(result)] = new_row
return result
df_task4 = find_most_common_word_follows_least(df, matrix)
df_task4
# + colab={"base_uri": "https://localhost:8080/", "height": 424} id="3gZal80JlEIu" outputId="337f5f85-d5e7-4de1-e3ec-5d0303a0e978"
df_task4[df_task4['Occurence_predecessor'] == df_task4['Occurence_predecessor'].max()]
| HW2/HW2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ER+ breast cancer effective graph
#
# > <NAME>, <NAME>, <NAME>, & <NAME> [2021]. "The effective graph reveals redundancy, canalization, and control pathways in biochemical regulation and signaling". Proceedings of the National Academy of Sciences (PNAS), 118 (12).
import numpy as np
import pandas as pd
from collections import Counter
import graphviz
# Cana
import cana
from cana.boolean_network import BooleanNetwork
from cana.datasets.bio import BREAST_CANCER
# Matplotlib
import matplotlib as mpl
import matplotlib.pyplot as plt
from matplotlib.colors import LinearSegmentedColormap
# Networkx
import networkx as nx
def calculates_path_length(G, path, weight='weight'):
path_weight_sum = 0.0
weakest_link = np.inf
for source, target in zip(path[:-1], path[1:]):
edge_weight = G.edges[(source, target)][weight]
path_weight_sum += edge_weight
if edge_weight < weakest_link:
weakest_link = edge_weight
return path_weight_sum, weakest_link
C = BREAST_CANCER()
SG = C.structural_graph()
EG = C.effective_graph(threshold=-1)
#
EG0 = C.effective_graph(threshold=0)
EG0p2 = C.effective_graph(threshold=.2)
EG0p4 = C.effective_graph(threshold=.4)
pd.options.display.float_format = '{:.2g}'.format
#
df = pd.DataFrame({
'node':[n.name for n in C.nodes],
'k':[n.k for n in C.nodes],
'k_r':[n.input_redundancy(norm=False) for n in C.nodes],
'k_e':[n.effective_connectivity(norm=False) for n in C.nodes],
'k_r*':[n.input_redundancy(norm=True) for n in C.nodes],
'k_e*':[n.effective_connectivity(norm=True) for n in C.nodes],
'k^{out}':[v for n,v in EG.out_degree()],
'k_e^{out}':[v for n,v in EG.out_degree(weight='weight')],
})
#df = df[['k','k_r','k_e','k_r*','k_e*','k^{out}','k_e^{out}']]
df['k_e^{out}/k^{out}'] = df['k_e^{out}'] / df['k^{out}']
df.sort_values('k',ascending=False,inplace=True)
#print(df.to_latex(escape=False))
#
drugs = [3, 4, 5, 6, 7, 8, 9]
dfd = df.loc[drugs, ['node', 'k^{out}', 'k_e^{out}', 'k_e^{out}/k^{out}']]
dfd.sort_values(['k^{out}', 'k_e^{out}'], ascending=[False, False], inplace=True)
display(dfd)
def number_of_input_nodes(G):
count = 0
for n, d in G.nodes(data=True):
inputs = [True if i == j else False for i, j in G.in_edges(n)]
if inputs == [] or inputs == [True]:
count += 1
return count
#
def number_of_nodes_with_self_loop(G):
count = 0
for n, d in G.nodes(data=True):
inputs = [True if i == j else False for i, j in G.in_edges(n)]
if any(inputs):
count += 1
return count
#
def number_of_input_nodes_with_self_loop(G):
count = 0
for n, d in G.nodes(data=True):
inputs = [True if i == j else False for i, j in G.in_edges(n)]
if inputs == [True]:
count += 1
return count
for G, graph in [(SG, 'IG'), (EG0, 'EG'), (EG0p2, 'EG (0.2)'), (EG0p4, 'EG (0.4)')]:
n_nodes = G.number_of_nodes()
print('{g:s}: Number of nodes: {n:d}'.format(g=graph, n=n_nodes))
n_nodes_with_self_loop = number_of_nodes_with_self_loop(G)
print('{g:s}: Number nodes with a self-loop: {n:d}'.format(g=graph, n=n_nodes_with_self_loop))
n_input_nodes = number_of_input_nodes(G)
print('{g:s}: Number of input nodes: {n:d}'.format(g=graph, n=n_input_nodes))
n_input_nodes_with_self_loop = number_of_input_nodes_with_self_loop(G)
print('{g:s}: Number of input nodes with self-loop: {n:d}'.format(g=graph, n=n_input_nodes_with_self_loop))
print('--')
def weakly_connected_components(G):
wcc = [len(cc) for cc in nx.weakly_connected_components(G)]
return len(wcc), wcc
#
def strongly_connected_components(G):
scc = [len(cc) for cc in nx.strongly_connected_components(G)]
return len(scc), scc
#
def SortedCounter(l):
c = dict(sorted(Counter(l).most_common(), reverse=True))
return c
for G, graph in [(SG, 'IG'), (EG0, 'EG'), (EG0p2, 'EG (0.2)'), (EG0p4, 'EG (0.4)')]:
n_wcc, wcc = weakly_connected_components(G)
print('{g:s}: {n:d} weakly connected components, sizes {l:}'.format(g=graph, n=n_wcc, l=SortedCounter(wcc)))
#
n_scc, scc = strongly_connected_components(G)
print('{g:s}: {n:d} strongly connected components, sizes {l:}'.format(g=graph, n=n_scc, l=SortedCounter(scc)))
print('--')
wcc = nx.weakly_connected_components(EG0p2)
for id,comp in enumerate(wcc):
print('Component id={id:d} with size: {size:d}'.format(id=id, size=len(comp)))
names = []
for node in comp:
name = G.nodes[node]['label']
names.append(name)
print(names)
# Print node names
for i,d in SG.nodes(data=True):
print(i,d)
net_redundancy = 0
#
for nid, node in enumerate(C.nodes,start=0):
net_redundancy += node.input_redundancy(norm=False)
#
net_redundancy_norm = net_redundancy / C.Nnodes
#
print("Total Network Redundancy: {:.4} (norm: {:.4})".format(net_redundancy, net_redundancy_norm))
dict_effconn = {nid: node.effective_connectivity(norm=False)
for nid, node in enumerate(C.nodes, start=0) }
#
nx.set_node_attributes(EG, dict_effconn, 'effective_connectivity')
# +
# Node Manual Positioning
idmap = {d['label']:i for i,d in SG.nodes(data=True)}
att = {}
colors = {
'Apoptosis': '#b3dc66',
'Proliferation': '#bfbcd9',
'Drugs': '#d299ff',
'ER signaling': '#d3d3d3',
'mTORC1 pathway': '#ffb96c',
'AKT pathway': '#8fd3c4',
'MAPK pathway': '#ffffae',
'PI3K pathway': '#7aa6cb',
'RTK signaling': '#f18568'}
#Line 11
att['HER3_T'] = {'pos':'.25,11', 'type':'RTK signaling'}
att['HER3'] = {'pos':'1.5,11', 'type':'RTK signaling'}
att['HER3_2'] = {'pos':'2.7,11', 'type':'RTK signaling'}
att['IGF1R_T'] = {'pos':'3.9,11', 'type':'RTK signaling'}
# Line 10
att['Neratinib'] = {'pos':'0,10', 'type':'Drugs'}
att['Alpelisib'] = {'pos':'4.3,9', 'type':'Drugs'}
att['Fulvestrant'] = {'pos':'7.4,11', 'type':'Drugs', 'width':'0.9'}
att['HER2_3_2'] = {'pos':'1.25,10', 'type':'RTK signaling'}
att['HER2_3'] = {'pos':'2.5,10', 'type':'RTK signaling'}
att['HER2'] = {'pos':'3.8,10', 'type':'RTK signaling'}
att['IGF1R'] = {'pos':'5,11', 'type':'RTK signaling'}
att['IGF1R_2'] = {'pos':'5,10', 'type':'RTK signaling'}
att['ESR1'] = {'pos':'6.8,10', 'type':'ER signaling'}
att['ESR1_2'] = {'pos':'8,10', 'type':'ER signaling'}
att['FOXA1'] = {'pos':'9.2,10', 'type':'ER signaling'}
att['PBX1'] = {'pos':'10.7,8', 'type':'ER signaling'}
# Line 9
att['Trametinib'] = {'pos':'0,9', 'type':'Drugs', 'width':'0.9'}
att['PI3K'] = {'pos':'3,9', 'type':'PI3K pathway'}
att['Everolimus'] = {'pos':'8,6', 'type':'Drugs', 'width':'0.9'}
att['Ipatasertib'] = {'pos':'6.0,9', 'type':'Drugs', 'width':'0.9'}
att['ER'] = {'pos':'8.5,9', 'type':'ER signaling'}
#Line 8
att['MAPK'] = {'pos':'0.2,8', 'type':'MAPK pathway'}
att['PI3K_2'] = {'pos':'3.0,8', 'type':'PI3K pathway'}
att['PIP3'] = {'pos':'4.3,8', 'type':'PI3K pathway'}
att['PIP3_2'] = {'pos':'4.3,7', 'type':'PI3K pathway'}
att['PTEN'] = {'pos':'3.0,7', 'type':'PI3K pathway'}
att['ER_transcription'] = {'pos':'7.6,8', 'type':'ER signaling', 'width':'1.3'}
att['ER_transcription_2'] = {'pos':'9.4,8', 'type':'ER signaling', 'width':'1.5'}
# Line 7
att['MAPK_2'] = {'pos':'0,7', 'type':'MAPK pathway'}
att['RAS'] = {'pos':'1.5,9', 'type':'MAPK pathway'}
att['RAS_2'] = {'pos':'1.5,7', 'type':'MAPK pathway'}
att['RAS_3'] = {'pos':'1.5,8', 'type':'MAPK pathway'}
att['PDK1_pm'] = {'pos':'4.5,6', 'type':'AKT pathway'}
att['mTORC2_pm'] = {'pos':'6.25,7', 'type':'AKT pathway', 'width':'1.2'}
att['KMT2D'] = {'pos':'9,7', 'type':'ER signaling'}
att['MYC'] = {'pos':'10,7', 'type':'ER signaling'}
att['MYC_2'] = {'pos':'11.2,7', 'type':'ER signaling'}
# Line 6
att['PDK1'] = {'pos':'0.5,6', 'type':'AKT pathway'}
att['mTORC2'] = {'pos':'1.75,6', 'type':'AKT pathway'}
att['PIM'] = {'pos':'3,6', 'type':'AKT pathway'}
att['AKT'] = {'pos':'6,6', 'type':'AKT pathway'}
att['Palbociclib'] = {'pos':'9.7,6', 'type':'Drugs', 'width':'0.9'}
# Line 5
att['SGK1_T'] = {'pos':'0.25,5', 'type':'AKT pathway'}
att['SGK1'] = {'pos':'1.5,5', 'type':'AKT pathway'}
att['TSC'] = {'pos':'5.5,5', 'type':'mTORC1 pathway'}
att['p21_p27'] = {'pos':'7,5', 'type':'Proliferation'}
att['p21_p27_T'] = {'pos':'8,5', 'type':'Proliferation'}
att['CDK46'] = {'pos':'9,5', 'type':'Proliferation'}
att['cyclinD'] = {'pos':'10.25,5', 'type':'Proliferation'}
# Line 4
att['FOXO3_Ub'] = {'pos':'1.3,4', 'type':'AKT pathway', 'width':'0.9'}
att['FOXO3'] = {'pos':'2.75,4', 'type':'AKT pathway'}
att['cycE_CDK2'] = {'pos':'6.5,4', 'type':'Proliferation', 'width':'1.0'}
att['cycE_CDK2_T'] = {'pos':'7.8,4', 'type':'Proliferation', 'width':'1.2'}
att['PRAS40'] = {'pos':'4,4', 'type':'mTORC1 pathway'}
att['mTORC1'] = {'pos':'5,4', 'type':'mTORC1 pathway'}
att['cycD_CDK46'] = {'pos':'9.2,4', 'type':'Proliferation', 'width': '1.0'}
att['cycD_CDK46_2'] = {'pos':'11.0,4', 'type':'Proliferation', 'width': '1.4'}
# Line 3
att['BIM_T'] = {'pos':'.5,3', 'type':'Apoptosis'}
att['BCL2_T'] = {'pos':'2,3', 'type':'Apoptosis'}
att['EIF4F'] = {'pos':'4,3', 'type':'mTORC1 pathway'}
att['S6K'] = {'pos':'5,3', 'type':'mTORC1 pathway'}
att['pRb'] = {'pos':'7.0,3', 'type':'Proliferation'}
att['pRb_2'] = {'pos':'8.25,3', 'type':'Proliferation'}
att['pRb_3'] = {'pos':'9.5,3', 'type':'Proliferation'}
att['cyclinD_2'] = {'pos':'10.5,3', 'type':'Proliferation'}
# Line 2
att['BIM'] = {'pos':'0,2', 'type':'Apoptosis'}
att['BAD'] = {'pos':'1,2', 'type':'Apoptosis'}
att['BCL2'] = {'pos':'2,2', 'type':'Apoptosis'}
att['MCL1'] = {'pos':'3,2', 'type':'Apoptosis'}
att['Translation'] = {'pos':'4.5,2', 'type':'mTORC1 pathway', 'width':'0.9'}
att['E2F'] = {'pos':'7.0,2', 'type':'Proliferation'}
att['E2F_2'] = {'pos':'8.25,2', 'type':'Proliferation'}
att['E2F_3'] = {'pos':'9.5,2', 'type':'Proliferation'}
# Line 1
att['Apoptosis'] = {'pos':'0.2,1', 'type':'Apoptosis', 'width':'1.0'}
att['Apoptosis_2'] = {'pos':'1.5,1', 'type':'Apoptosis', 'width':'1.0'}
att['Apoptosis_3'] = {'pos':'2.8,1', 'type':'Apoptosis', 'width':'1.0'}
att['Proliferation'] = {'pos':'6.5,1', 'type':'Proliferation', 'width':'1.1'}
att['Proliferation_2'] = {'pos':'7.9,1', 'type':'Proliferation', 'width':'1.1'}
att['Proliferation_3'] = {'pos':'9.3,1', 'type':'Proliferation', 'width':'1.1'}
att['Proliferation_4'] = {'pos':'10.7,1', 'type':'Proliferation', 'width':'1.1'}
# Check if overlapping nodes
poschk = set()
for k,v in att.items():
if v['pos'] in poschk:
raise TypeError("Overlapping nodes: {k:s}".format(k=k))
else:
poschk.add(v['pos'])
# Inverse Map
att = {idmap[k]:v for k,v in att.items()}
# -
# ## Interaction Graph
# +
pSG = graphviz.Digraph(name='Structural Graph', engine='neato')
pSG.attr('graph', size='7,7', concentrate='false', simplify='false', overlap='true',splines='true',ratio='.7',outputorder="edgesfirst",nodesep='.25',mindist='.20')
pSG.attr('node', pin='true', shape='box', height='0.4', fixedsize='false',margin='.05', color='black', style='filled', fillcolor='#515660', penwidth='1', fontname='Helvetica', fontcolor='black',fontsize='10')
pSG.attr('edge', arrowhead='normal', arrowsize='.5', penwidth='4')
for nid,SGatt in SG.nodes(data=True):
label = SGatt['label']
if nid in att:
pos = att[nid].get('pos', '')
shape = att[nid].get('shape', 'box')
fillcolor = colors[att[nid].get('type')]
#width = ''att[nid].get('width', '0.7')
pSG.node(str(nid), label=label, pos=pos, shape=shape, fillcolor=fillcolor, )
max_penwidth = 4
for uid,vid,d in SG.edges(data=True):
uid = str(uid)
vid = str(vid)
weight = '%d' % (d['weight']*100)
# self loop color
if uid == vid:
color = '#bdbdbd'
uid = uid + ':w'
vid = vid + ':c'
else:
color = '#636363'
pSG.edge(uid, vid, weight=weight, color=color)
display(pSG)
# Export
#pSG.render("BreastCancer-IG", cleanup=True)
# -
# ## Effective Graph
# +
pEG = graphviz.Digraph(name='Structural Graph', engine='neato')
pEG.attr('graph', size='7,7', concentrate='false', simplify='false', overlap='true',splines='true',ratio='.7',outputorder="edgesfirst",nodesep='.25',mindist='.20')
pEG.attr('node', pin='true', shape='box', height='0.4', fixedsize='false',margin='.05', color='black', style='filled', fillcolor='#515660', penwidth='1', fontname='Helvetica', fontcolor='black',fontsize='10')
pEG.attr('edge', arrowhead='normal', arrowsize='.5', penwidth='4')
max_effoutdegree = max([v for n,v in EG.out_degree()])
list_effconn = [d['effective_connectivity'] for n,d in EG.nodes(data=True)]
min_effconn, max_effconn = min(list_effconn), max(list_effconn)
#
for nid,d in EG.nodes(data=True):
label = d['label']
ntype = att[nid].get('type')
pos = att[nid].get('pos', '')
shape = att[nid].get('shape')
fillcolor = colors[ntype]
pEG.node(str(nid), label=label, pos=pos, shape=shape, fillcolor=fillcolor,)
max_penwidth = 4
for uid,vid,d in EG.edges(data=True):
uid = str(uid)
vid = str(vid)
weight = '%d' % (d['weight']*100)
penwidth = '%.2f' % ( d['weight']*max_penwidth )
if d['weight'] > 0:
if uid == vid:
color = '#bdbdbd'
uid = uid + ':w'
vid = vid + ':c'
else:
color = '#636363'
pEG.edge(uid,vid, weight=weight, penwidth=penwidth, color=color)
else:
pEG.edge(uid,vid, style='dashed', color='#A90533')
display(pEG)
# Export
#pEG.render("BreastCancer-EG", cleanup=True)
# +
pEG = graphviz.Digraph(name='Structural Graph', engine='neato')
pEG.attr('graph', size='7,7', concentrate='false', simplify='false', overlap='true',splines='true',ratio='.7',outputorder="edgesfirst",nodesep='.25',mindist='.20')
pEG.attr('node', pin='true', shape='box', height='0.4', fixedsize='false',margin='.05', color='black', style='filled', fillcolor='#515660', penwidth='1', fontname='Helvetica', fontcolor='black',fontsize='10')
pEG.attr('edge', arrowhead='normal', arrowsize='.5', penwidth='4')
max_effoutdegree = max([v for n,v in EG0p2.out_degree()])
list_effconn = [d['effective_connectivity'] for n,d in EG.nodes(data=True)]
min_effconn, max_effconn = min(list_effconn), max(list_effconn)
#
for nid,d in EG0p4.nodes(data=True):
label = d['label']
ntype = att[nid].get('type')
pos = att[nid].get('pos', '')
shape = att[nid].get('shape')
fillcolor = colors[ntype]
pEG.node(str(nid), label=label, pos=pos, shape=shape, fillcolor=fillcolor,)
max_penwidth = 4
for uid,vid,d in EG0p4.edges(data=True):
uid = str(uid)
vid = str(vid)
weight = '%d' % (d['weight']*100)
penwidth = '%.2f' % ( d['weight']*max_penwidth )
if d['weight'] > 0.4:
if uid == vid:
color = '#bdbdbd'
uid = uid + ':w'
vid = vid + ':c'
else:
color = '#636363'
pEG.edge(uid,vid, weight=weight, penwidth=penwidth, color=color)
else:
pEG.edge(uid,vid, style='dashed', color='#A90533')
display(pEG)
# Export
#pEG.render("BreastCancer-EGt0p4", cleanup=True)
# +
pEG = graphviz.Digraph(name='Structural Graph', engine='dot')
pEG.attr('graph', size='8.5,9', concentrate='false', simplify='false', overlap='false',splines='true',ratio='compress',outputorder="edgesfirst",nodesep='.25',ranksep='.25')
pEG.attr('node', pin='true', shape='box', height='0.4', fixedsize='false', color='black', style='filled', fillcolor='#515660', penwidth='0.5', fontname='Helvetica', fontcolor='black',fontsize='12')
pEG.attr('edge', arrowhead='normal', arrowsize='.5', color='black', penwidth='3')
max_effoutdegree = max([v for n,v in EG.out_degree()])
list_effconn = [d['effective_connectivity'] for n,d in EG.nodes(data=True)]
min_effconn, max_effconn = min(list_effconn), max(list_effconn)
#
# ColorBar
interval = np.linspace(0.0, 0.70, 256)
cmap = LinearSegmentedColormap.from_list('custom', ['white','#d62728'])
#norm = mpl.colors.Normalize(vmin=0, vmax=1)
norm = mpl.colors.Normalize(vmin=1, vmax=max_effoutdegree)
#
colornodes = ['Apoptosis', 'Apoptosis_2', 'Apoptosis_3', 'Proliferation', 'Proliferation_2', 'Proliferation_3', 'Proliferation_4']
#
# Manually position nodes using the DotLayout
dotpos = nx.drawing.nx_agraph.graphviz_layout(EG, prog='dot', args='-Gconcentrate=false -Gsimplify=false -Goverlap=false -Gsplines=True -Gratio=compress -Goutputorder=edgesfirst -Gnodesep=.25 -Granksep=.20')
#
for nid,d in EG.nodes(data=True):
label = d['label']
ntype = att[nid].get('type')
x,y = dotpos[nid][0], dotpos[nid][1]
pos = '{x:.2f},{y:.2f}'.format(x=x, y=y) #''#att[nid].get('pos', '')
shape = att[nid].get('shape')
fillcolor = colors[ntype]
pEG.node(str(nid), label=label, pos=pos, shape=shape, fillcolor=fillcolor)
max_penwidth = 4
for uid,vid,d in EG.edges(data=True):
uid = str(uid)
vid = str(vid)
weight = '%d' % (d['weight']*100)
penwidth = '%.2f' % ( d['weight']*max_penwidth )
if d['weight'] >= 0.2:
if uid == vid:
color = '#bdbdbd'
uid = uid + ':w'
vid = vid + ':c'
else:
color = '#636363'
pEG.edge(uid,vid, weight=weight, penwidth=penwidth, color=color)
else:
pass
#pEG.edge(uid,vid, style='dashed', color='#A90533')
# Export
display(pEG)
#pEG.render("BreastCancer-EGt0p2-dotlayout", cleanup=True)
# +
# Legend
from matplotlib.patches import Patch
fig, ax = plt.subplots(nrows=1, ncols=1, figsize=(2,2))
labels = [
'RTK signaling',
'PI3K pathway',
'MAPK pathway',
'AKT pathway',
'mTORC1 pathway',
'ER signaling',
'Apoptosis',
'Proliferation',
'Drugs']
handles = []
for label in labels:
facecolor = colors[label]
patch = Patch(facecolor=facecolor, edgecolor='black', label=label)
handles.append(patch)
leg = ax.legend(handles=handles, labels=labels, loc='upper left', ncol=3)
plt.axis('off')
plt.show()
| tutorials/PNAS 2021 - ER+ Breast Cancer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import os
import rabbitpy
import time
from ch6 import utils
# Open the channel and connection
connection = rabbitpy.Connection()
channel = connection.channel()
exchange = rabbitpy.DirectExchange(channel, 'rpc-replies')
exchange.declare()
# +
# Create the response queue that will automatically delete, is not durable and
# is exclusive to this publisher
queue_name = 'response-queue-%s' % os.getpid()
response_queue = rabbitpy.Queue(channel,
queue_name,
auto_delete=True,
durable=False,
exclusive=True)
# Declare the response queue
if response_queue.declare():
print('Response queue declared')
# Bind the response queue
if response_queue.bind('rpc-replies', queue_name):
print('Response queue bound')
# +
# Iterate through the images to send RPC requests for
for img_id, filename in enumerate(utils.get_images()):
print('Sending request for image #%s: %s' % (img_id, filename))
# Create the message
message = rabbitpy.Message(channel,
utils.read_image(filename),
{'content_type': utils.mime_type(filename),
'correlation_id': str(img_id),
'reply_to': queue_name},
opinionated=True)
# Pubish the message
message.publish('direct-rpc-requests', 'detect-faces')
# Loop until there is a response message
message = None
while not message:
time.sleep(0.5)
message = response_queue.get()
# Ack the response message
message.ack()
# Caculate how long it took from publish to response
duration = (time.time() -
time.mktime(message.properties['headers']['first_publish']))
print('Facial detection RPC call for image %s total duration: %s' %
(message.properties['correlation_id'], duration))
# Display the image in the IPython notebook interface
utils.display_image(message.body, message.properties['content_type'])
print('RPC requests processed')
# Close the channel and connection
channel.close()
connection.close()
| notebooks/6.1.3 RPC Publisher.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import seaborn as sns
from matplotlib import pyplot as plt
import math
data = pd.read_csv("nces-ed-attainment.csv")
data
# # scikit-learn
# Below are a list of functions/features you will most likely use on this assignment. This is not the only functions possible to use and these functions can actually be used in a lot more cool and complicated ways, but we are going to focus on the basics in this wordbank. For these examples, we will use the iris dataset provided by `seaborn`
#
# To run this document, you must first run the following cell(s).
# +
import pandas as pd
import seaborn as sns
sns.set()
iris = pd.read_csv('/course/lessons/iris.csv')
iris.head()
# -
# Commonly people call the features X and the labels y
X = iris.loc[:, iris.columns != 'species']
y = iris['species']
# ## `sklearn.model_selection.train_test_split`
# This function can split your dataset into train and test sets with sizes of the given ration. Returns a 4-tuple of `(train_data, test_data, train_label, test_label)`.
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.33)
print(f'Train size: {len(X_train)} ({len(X_train) * 100 / len(X):0.2f}%)')
print(f'Test size: {len(X_test)} ({len(X_test) * 100 / len(X):0.2f}%)')
# ## `sklearn.tree.DecisionTreeClassifier`
# A tree-based model to solve classification task (e.g. predicting a label like "spam" or "not spam"). See sections below for functions that can be used on any model.
# +
from sklearn.tree import DecisionTreeClassifier
model = DecisionTreeClassifier()
# -
# ## `sklearn.tree.DecisionTreeRegressor`
# A tree-based model to solve a regression task (e.g. predicting numerical quantity). See sections below for functions that can be used on any model.
# +
from sklearn.tree import DecisionTreeRegressor
model = DecisionTreeRegressor()
# -
# ## Model: `fit`
# Every model has a `fit` function that takes a dataset (features and labels) and trains the model using that data. For this example, since we will be using the iris dataset which is predicting the class of iris from information about its petals, we will be using a classifier.
# +
from sklearn.tree import DecisionTreeClassifier
model = DecisionTreeClassifier()
model.fit(X_train, y_train)
# -
# ## Model: `predict`
# Every model has a `predict` function that takes a dataset (features) and predicts all the labels for that dataset. You must `fit` the model before you may `predict` with it. For this example, since we will be using the iris dataset which is predicting the class of iris from information about its petals, we will be using a classifier.
#
# We assume the previous cell weas the last to run.
y_train_pred = model.predict(X_train)
y_test_pred = model.predict(X_test)
#pd.Series(y_train_pred)
d = X_train.copy()
d['actual'] = pd.Series(y_train_pred)
print(pd.Series(y_train_pred))
y_train_pred
# ## `sklearn.metrics.accuracy_score`
# If you are solving a classification problem, a common metric for the quality of the model is the accuracy in terms of the ratio of examples it predicted the correct label. A higher accuracy means the model more closely fit the data.
#
# Mathematically, this is defined as
#
# $$Accuracy(y_{true}, y_{pred}) = \sum_{i=1}^n \textbf{1}\left(y_{true}(i) = y_{pred}(i)\right)$$
#
# where $y_{true}$ are the true labels, $y_{pred}$ are the predicted labels, $n$, is the number of examples, and $\textbf{1}$ takes the value 1 if the condition inside is true, and 0 otherwise.
#
# Alternatively, you could write this in code as
#
# ```python
# def accuraccy(y_true, y_pred):
# correct = 0
# for i in range(len(y_true)):
# if y_true[i] == y_pred[i]:
# correct += 1
# return correct / len(y_true)
# ```
#
# It's much simpler to have scikit-learn compute this for you like below (assumes the cells above have been run):
# +
from sklearn.metrics import accuracy_score
print('Train accuracy:', accuracy_score(y_train, y_train_pred))
print('Test accuracy:', accuracy_score(y_test, y_test_pred))
# -
# ## `sklearn.metrics.mean_square_error`
# If you are solving a regression problem, a common metric for the quality of the model is the average value of squares of taking the difference between a prediction and the true value. This is called mean squared error or MSE. A lower MSE means the model more closely fit the data.
#
# Mathematically, this is defined as
#
# $$MSE(y_{true}, y_{pred})= \frac{1}{n}\sum_{i=1}^n \left( y_{true}(i)-y_{pred}(i)\right)^2$$
#
# where $y_{true}$ are the true values, $y_{pred}$ are the predicted values, $n$, is the number of examples.
#
# Alternatively, you could write this in code as
#
# ```python
# def mse(y_true, y_pred):
# total_error = 0
# for i in range(len(y_true)):
# total_error += (y_true[i] - y_pred[i]) ** 2
# return total_error / len(y_true)
# ```
#
# It's much simpler to have scikit-learn compute this for you like below. **You can't actually run these cells** since the data and predictions were made for a classification problem, but if they were used for regression you could then run.
# +
from sklearn.metrics import mean_squared_error
print('Train MSE:', mean_squared_error(y_train, y_train_pred))
print('Test MSE:', mean_square_error(y_test, y_test_pred))
| ScikitLearnWordbank.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Binary Predictors in a Logistic Regression
# Using the same code as in the previous exercise, find the odds of 'duration'.
#
# What do they tell you?
# ## Import the relevant libraries
# +
import numpy as np
import pandas as pd
import statsmodels.api as sm
import matplotlib.pyplot as plt
import seaborn as sns
sns.set()
from scipy import stats
stats.chisqprob = lambda chisq, df: stats.chi2.sf(chisq, df)
# -
# ## Load the data
# Load the ‘Bank_data.csv’ dataset.
raw_data = pd.read_csv('Bank_data.csv')
raw_data
# We make sure to create a copy of the data before we start altering it. Note that we don't change the original data we loaded.
data = raw_data.copy()
# Removes the index column thata comes with the data
data = data.drop(['Unnamed: 0'], axis = 1)
# We use the map function to change any 'yes' values to 1 and 'no'values to 0.
data['y'] = data['y'].map({'yes':1, 'no':0})
data
data.describe()
# ### Declare the dependent and independent variables
# Use 'duration' as the independet variable.
y = data['y']
x1 = data['duration']
# ### Simple Logistic Regression
# Run the regression.
x = sm.add_constant(x1)
reg_log = sm.Logit(y,x)
results_log = reg_log.fit()
# Get the regression summary
results_log.summary()
# Create a scatter plot of x1 (Duration, no constant) and y (Subscribed)
plt.scatter(x1,y,color = 'C0')
# Don't forget to label your axes!
plt.xlabel('Duration', fontsize = 20)
plt.ylabel('Subscription', fontsize = 20)
plt.show()
# ### Find the odds of duration
# the odds of duration are the exponential of the log odds from the summary table
np.exp(0.0051)
# The odds of duration are pretty close to 1. This tells us that although duration is a significant predictor, a change in 1 day would barely affect the regression.
#
# Note that we could have inferred that from the coefficient itself.
#
# Finally, note that the data is not standardized (scaled) and duration is a feature of a relatively big order of magnitude.
| 9_LogisticRegression_BinaryVariables_S36_L245/Binary Predictors in a Logistic Regression - Solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Feature Selection Mini-Project - Find signature
#
# Katie explained in a video a problem that arose in preparing Chris and Sara’s email for the author identification project; it had to do with a feature that was a little too powerful (effectively acting like a signature, which gives an arguably unfair advantage to an algorithm). You’ll work through that discovery process here.
#
#
# ## Overfitting a Decision Tree 1
#
# This bug was found when Katie was trying to make an overfit decision tree to use as an example in the decision tree mini-project. A decision tree is classically an algorithm that can be easy to overfit; one of the easiest ways to get an overfit decision tree is to use a small training set and lots of features.
#
# #### If a decision tree is overfit, would you expect the accuracy on a test set to be very high or pretty low?
# Ans : Pretty low
#
# #### If a decision tree is overfit, would you expect high or low accuracy on the training set?
# Ans : High
#
# The accuracy would be very high on the training set, but would plummet once it was actually tested.
#
# ## Number of Features and Overfitting
#
# A classic way to overfit an algorithm is by using lots of features and not a lot of training data.
#
# You can find the starter code in ```feature_selection/find_signature.py```.
# Get a decision tree up and training on the training data, and print out the accuracy.
# +
# Starter code
import pickle
import numpy
numpy.random.seed(42)
# +
# Starter code
### The words (features) and authors (labels), already largely processed.
### These files should have been created from the previous (Lesson 10)
### mini-project.
words_file = "../text_learning/your_word_data.pkl"
authors_file = "../text_learning/your_email_authors.pkl"
word_data = pickle.load( open(words_file, "rb"))
authors = pickle.load( open(authors_file, "rb") )
# +
# Starter code
### test_size is the percentage of events assigned to the test set (the remainder go into training)
### feature matrices changed to dense representations for compatibility with classifier functions in versions 0.15.2 and earlier
#from sklearn import cross_validation ==> verouderd
from sklearn.model_selection import train_test_split
features_train, features_test, labels_train, labels_test = train_test_split(word_data, authors, test_size=0.1, random_state=42)
from sklearn.feature_extraction.text import TfidfVectorizer
vectorizer = TfidfVectorizer(sublinear_tf=True, max_df=0.5, stop_words='english') # max_df = skip a word if it exists in >50% of documents
# the vectorizer is being fitted on the training features. This will allow it to build its list of
# vocabulary to generate features and also get feature names.
features_train = vectorizer.fit_transform(features_train)
features_test = vectorizer.transform(features_test).toarray()
# +
# Starter code
### a classic way to overfit is to use a small number of data points and a large number of features;
### train on only 150 events to put ourselves in this regime
features_train = features_train[:150].toarray()
labels_train = labels_train[:150]
# -
# #### How many training points are there, according to the starter code?
len(features_train)
# Yup! We've limited our training data quite a bit, so we should be expecting our models to potentially overfit.
# ### Accuracy of Your Overfit Decision Tree
#
# What’s the accuracy of the decision tree you just made? (Remember, we're setting up our decision tree to overfit -- ideally, we want to see the test accuracy as relatively low.)
from sklearn import tree
from sklearn.metrics import accuracy_score
clf = tree.DecisionTreeClassifier()
clf = clf.fit(features_train, labels_train)
pred = clf.predict(features_test)
accuracy = accuracy_score(labels_test, pred)
print ("Accuracy of DT classifier is: ", accuracy)
# Yes, the test performance has an accuracy much higher than it is expected to be - if we are overfitting, then the test performance should be relatively low.
# ### Identify the Most Powerful Features
#
# Take your (overfit) decision tree and use the ```feature_importances_``` attribute to get a list of the relative importance of all the features being used.
#
# We suggest iterating through this list (it’s long, since this is text data) and only printing out the feature importance if it’s above some threshold (say, 0.2--remember, if all words were equally important, each one would give an importance of far less than 0.01). What’s the importance of the most important feature? What is the number of this feature?
#
# __Guidance:__ The object of this exercise is to analyze which features are most predictive or most important. The output should be all features above a certain threshold and their feature importance and feature number. The feature importance of the most important feature and the number of that feature are what should be entered into the quiz.
#
# One way to proceed would be to run the code and look at the output (the printed output). You may consider playing with the threshold and seeing how this impacts the output.
#
# The number we get from "identifying the most powerful features" is actually the index of for that feature
importances = clf.feature_importances_
type(importances)
importances.dtype
# Answer
for i in range(len(importances)):
if importances[i] > 0.2:
print ("Most Important feature : ",importances[i])
print ("Feature Number : ",i)
importances
import numpy as np
importances = np.sort(importances) # nb: bij numpy array kun je niet ascending sorteren
importances = importances[::-1]
importances[:10]
for i in importances:
if i > 0.1:
print (i)
# - __Deciding on a threshold :__ This will vary from model to model. The feature importance is somewhat of a measure of how much information we gain from using that feature as measured by the impact the split has on overall system purity. That is feature splits that decrease the impurity of the system more are more important. Often we look at several features and choose a threshold based on using a reasonable number of features with reasonably high scores relative to other features. For example in the plot below we would likely choose the first three features, after which there is a drop off in importance.
#
# from IPython.display import Image
# Image(filename='Embarcadero.png')
#
# 
#
#
#
# - To get the most important feature we don't necessarily need to set a threshold as we do in the code above. You could also return all feature importances and sort. Setting a threshold is a good idea because it filters out features we wouldn't consider important and gives us a smaller list to work with.
#
# - To get the feature number we can use the index of importances. If we determine the index of the highest scoring feature, this can be used to determine the feature number. There are other approaches as well, such as counting through the feature iteration.
# ## Use TfIdf to Get the Most Important Word
#
# In order to figure out what words are causing the problem, you need to go back to the TfIdf and use the feature numbers that you obtained in the previous part of the mini-project to get the associated words. You can return a list of all the words in the TfIdf by calling ```get_feature_names()``` on it; pull out the word that’s causing most of the discrimination of the decision tree. What is it? Does it make sense as a word that’s uniquely tied to either __<NAME>__ or __<NAME>__, a signature of sorts?
words_list = vectorizer.get_feature_names()
words_list[21323]
type(words_list)
word_list.dtype
len(word_list)
word_list = np.sort(words_list)
word_list
import pandas as pd
data_file = pd.DataFrame(word_list)
data_file
da
data_file[0].value_counts()
words_list[21323:21330]
# This is the most powerful word when the decision tree is making its classification decision.
#
# Even though our training data is limited, we still have a word that is highly indicative of author.
# ## Remove, Repeat
#
# This word seems like an outlier in a certain sense, so let’s remove it and refit. Go back to ```text_learning/vectorize_text.py```, and remove this word from the emails using the same method you used to remove “sara”, “chris”, etc. Rerun ```vectorize_text.py```, and once that finishes, rerun find_signature.py. Any other outliers pop up? What word is it? Seem like a signature-type word? (Define an outlier as a feature with importance >0.2, as before).
#
# After removing the first signature word, another powerful signature word arises.
#
# __cgermannsf__
#
# ## Checking Important Features Again
#
# Update ```vectorize_test.py``` one more time, and rerun. Then run ```find_signature.py``` again. Any other important features (importance>0.2) arise? How many? Do any of them look like “signature words”, or are they more “email content” words, that look like they legitimately come from the text of the messages?
#
# __houectect__
#
# Yes, there is one more word ("houectect"). Your guess about what this word means is as good as ours, but it doesn't look like an obvious signature word so let's keep moving without removing it.
# ## Accuracy of the Overfit Tree
#
# What’s the accuracy of the decision tree now? We've removed two "signature words", so it will be more difficult for the algorithm to fit to our limited training set without overfitting. Remember, the whole point was to see if we could get the algorithm to overfit--a sensible result is one where the accuracy isn't that great!
accuracy = accuracy_score(labels_test, pred)
print ("Accuracy of DT classifier is : ", accuracy)
# Now that we've removed the outlier "signature words", the training data is starting to overfit to the words that remain.
# + jupyter={"outputs_hidden": true}
# -
| 12_Find_signature-1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.0 64-bit (''pytorch'': conda)'
# name: python3
# ---
import numpy as np
import torch
torch.set_printoptions(edgeitems=2, threshold=50)
with open('../data/p1ch4/jane-austen/1342-0.txt', encoding='utf8') as f:
text = f.read()
lines = text.split('\n')
line = lines[200]
line
letter_t = torch.zeros(len(line), 128) # <1>
letter_t.shape
for i, letter in enumerate(line.lower().strip()):
letter_index = ord(letter) if ord(letter) < 128 else 0 # <1>
letter_t[i][letter_index] = 1
# +
def clean_words(input_str):
punctuation = '.,;:"!?”“_-'
word_list = input_str.lower().replace('\n',' ').split()
word_list = [word.strip(punctuation) for word in word_list]
return word_list
words_in_line = clean_words(line)
line, words_in_line
# +
word_list = sorted(set(clean_words(text)))
word2index_dict = {word: i for (i, word) in enumerate(word_list)}
len(word2index_dict), word2index_dict['impossible']
# +
word_t = torch.zeros(len(words_in_line), len(word2index_dict))
for i, word in enumerate(words_in_line):
word_index = word2index_dict[word]
word_t[i][word_index] = 1
print('{:2} {:4} {}'.format(i, word_index, word))
print(word_t.shape)
# -
word_t = word_t.unsqueeze(1)
word_t.shape
[(c, ord(c)) for c in sorted(set(text))]
ord('l'
)
| _notebooks/2021-09-07-text-jane-austen.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 1° passo: extrair as características da imagens da base de treinamento
# Para cada imagem da base de dados, iremos extrair os Momentos de Hu (que são 7), e no final do vetor, colocaremos a classe que pertence a imagem
# + active=""
# Exemplo: [2.6, 3.9, 7.1, -14.4, -20.4, -18.5, 13, 1]
# + active=""
# Exemplo: [2.6, 3.9, 7.1, -14.4, -20.4, -18.5, 13, 2]
# + active=""
# Exemplo: [2.6, 3.9, 7.1, -14.4, -20.4, -18.5, 13, 1]
# +
import cv2
import numpy as np
import matplotlib.pyplot as plt
import os
files_path = [os.path.abspath(x) for x in os.listdir('./') if x.endswith('.png')]
def extrair_caracteristica(img):
gray = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
_, imgBinaria = cv2.threshold(gray, 250, 255, cv2.THRESH_BINARY) #seleciona apenas os pixels dentro do intervalo [250,255]
momentos = cv2.moments(imgBinaria)
momentosDeHu = cv2.HuMoments(momentos)
feature = (-np.sign(momentosDeHu) * np.log10(np.abs(momentosDeHu)))
return feature
base_teste = []
#extraindo as características das imagens na base de dados
for i in files_path:
diretorio, arquivo = os.path.split(i)
imagem = cv2.imread(arquivo)
carac = extrair_caracteristica(imagem)
classe = arquivo.split('-')
base_teste.append((carac, classe[0]))
# -
print(base_teste)
# 2° passo: calcular o vetor de característica da imagem consulta
img = cv2.imread('2-1.png')
vetor_consulta = extrair_caracteristica(img)
print(vetor_consulta)
# 3° passo: definir uma função de distância. Abaixo temos a distância Euclidiana
def distancia(a, b):
M = len(a)
soma = 0
for i in range(M):
soma = soma + ((a[i]-b[i])**2)
return np.sqrt(soma)
# 4° passo: calcular a distancia do vetor_consulta com todos os vetores das imagens da base_teste
# OBSERVAÇÃO: após calcular distância, incluir a classe da imagem que foi calculada a distância
# +
#calculando a distancia do vetor de características da imagem consulta com todos
# os vetores de características extraidos das imagens que estão na base de dados
d = []
for feat in base_teste:
vetor = feat[0]
dist = distancia(vetor, vetor_consulta)
d.append((dist, feat[1]))
# -
print(d)
# 5° passo: ordenar as distâncias em ordem crescente (menor para o maior)
e = sorted(d)
print(e)
# 6° passo: iremos contar qual é a classe que mais se repete considerando o top-k
# Neste exemplo, iremos usar k = 3, ou seja, as 3 imagens mais similares
# A classe que mais se repetir, será a classe da imagem consulta
k1 = e[0][1]
k2 = e[1][1]
k3 = e[2][1]
print(k1, k2, k3)
from statistics import mode
a = mode([k1,k2,k3])
print("classe final: ", a)
# ## Código final, juntando todos os passos
# +
import cv2
import numpy as np
import matplotlib.pyplot as plt
import os
from statistics import mode
def distancia(a, b):
M = len(a)
soma = 0
for i in range(M):
soma = soma + ((a[i]-b[i])**2)
return np.sqrt(soma)
def extrair_caracteristica(img):
gray = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
_, imgBinaria = cv2.threshold(gray, 250, 255, cv2.THRESH_BINARY) #seleciona apenas os pixels dentro do intervalo [250,255]
momentos = cv2.moments(imgBinaria)
momentosDeHu = cv2.HuMoments(momentos)
feature = (-np.sign(momentosDeHu) * np.log10(np.abs(momentosDeHu)))
return feature
files_path = [os.path.abspath(x) for x in os.listdir('./') if x.endswith('.png')]
base_teste = []
#extraindo as características das imagens na base de dados
for i in files_path:
diretorio, arquivo = os.path.split(i)
imagem = cv2.imread(arquivo)
carac = extrair_caracteristica(imagem)
classe = arquivo.split('-')
base_teste.append((carac, classe[0]))
img = cv2.imread('img_consulta.jpg')
vetor_consulta = extrair_caracteristica(img)
#calculando a distancia do vetor de características da imagem consulta com todos
# os vetores de características extraidos das imagens que estão na base de dados
d = []
for feat in base_teste:
vetor = feat[0]
dist = distancia(vetor, vetor_consulta)
d.append((dist, feat[1]))
e = sorted(d)
k1 = e[0][1]
k2 = e[1][1]
k3 = e[2][1]
a = mode([k1,k2,k3])
print(k1, k2, k3)
print("classe final: ", a)
# -
| 1 - Aulas/aula10/classificacao.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# ! nvidia-smi
# ! cat /proc/cpuinfo
# ! pip install fastcore --upgrade -qq
# ! pip install fastai --upgrade -qq
# ! pip install transformers --upgrade -qq
# ! pip install datasets --upgrade -qq
# ! pip install pytorch_lightning --upgrade -qq
# ! pip install wandb --upgrade -qq
# ! pip install ohmeow-blurr --upgrade -qq
# ! pip install timm --upgrade -qq
# ! pip install git+https://github.com/warner-benjamin/fastai_snippets.git -qq
import torch
torch.__version__
# %env WANDB_SILENT=true
import wandb
wandb.login()
import os
os.environ["TOKENIZERS_PARALLELISM"] = "false"
# # Imports
# +
import gc
# import fastai
from fastai.vision.all import *
from fastai.text.all import *
from fastai.callback.wandb import WandbCallback
from fastai_snippets.callback import simpleprofiler
from fastai_snippets.utils import simpleprofiler_wandb
# import blurr/huggingface
from blurr.data.all import *
from blurr.modeling.all import *
from datasets import load_dataset
from transformers import AutoTokenizer, AutoConfig, AutoModelForSequenceClassification
#import pytorch lightning
import timm
import pytorch_lightning as pl
from pytorch_lightning import LightningDataModule, LightningModule, Trainer
from torch.optim.lr_scheduler import OneCycleLR
from torch.optim import AdamW
import torchvision.transforms as tvt
from torchvision.datasets import ImageFolder
from pytorch_lightning.loggers import WandbLogger
import torchmetrics
# -
# # Setup
plat = 'Colab Pro High RAM'
GPU = 'T4'
img_bs = 64
nlp_bs = 16
source = untar_data(URLs.IMAGENETTE_320)
source = untar_data(URLs.IMAGENETTE)
# ## Modify SimpleProfiler to Log Results to wandb
# +
from pytorch_lightning.profiler.simple import SimpleProfiler
@patch
def summary(self:SimpleProfiler):
output_table = wandb.Table(columns=["Action", "Mean duration (s)", "Duration StDev (s)", "Num calls", "Total time (s)", "Percentage %"])
if len(self.recorded_durations) > 0:
max_key = max(len(k) for k in self.recorded_durations.keys())
def log_row(action, mean, num_calls, total, per):
row = f"{sep}{action:<{max_key}s}\t| {mean:<15}\t|"
row += f"{num_calls:<15}\t| {total:<15}\t| {per:<15}\t|"
return row
report, total_duration = self._make_report()
output_table.add_data("Total", "-", "-", "_", f"{total_duration:.5}", "100 %")
for action, durations, duration_per in report:
output_table.add_data(
action,
f"{np.mean(durations):.5}",
f"{np.std(durations):.5}",
f"{len(durations):}",
f"{np.sum(durations):.5}",
f"{duration_per:.5}",
)
wandb.log({"simple_profiler": output_table})
# -
# ## PyTorch Lightning Imagenette
class ImagenetteDataModule(LightningDataModule):
def __init__(self, size, woof, bs, train_transform=None, valid_transform=None):
super().__init__()
self.size, self.woof, self.bs = size, woof, bs
imagewoof_stats = ([0.496,0.461,0.399],[0.257,0.249,0.258])
imagenette_stats = ([0.465,0.458,0.429],[0.285,0.28,0.301])
if train_transform == None:
self.train_transform = tvt.Compose([
tvt.RandomResizedCrop(size, scale=(0.35, 1)),
tvt.RandomHorizontalFlip(),
tvt.ToTensor(),
tvt.Normalize(*imagewoof_stats) if woof else tvt.Normalize(*imagenette_stats)])
else:
self.train_transform = transforms.Compose(train_transform)
if valid_transform == None:
self.valid_transform = tvt.Compose([
tvt.CenterCrop(size),
tvt.ToTensor(),
tvt.Normalize(*imagewoof_stats) if woof else tvt.Normalize(*imagenette_stats)])
else:
self.valid_transform = transforms.Compose(valid_transform)
def prepare_data(self):
if self.size<=224: path = URLs.IMAGEWOOF_320 if self.woof else URLs.IMAGENETTE_320
else : path = URLs.IMAGEWOOF if self.woof else URLs.IMAGENETTE
self.source = untar_data(path)
def setup(self, stage=None):
if stage == "fit" or stage is None:
self.train = ImageFolder(self.source/'train', self.train_transform)
self.val = ImageFolder(self.source/'val', self.valid_transform)
def train_dataloader(self):
return DataLoader(self.train, batch_size=self.bs, shuffle=True, pin_memory=True, num_workers=min(8, num_cpus()))
def val_dataloader(self):
return DataLoader(self.val, batch_size=self.bs, pin_memory=True, num_workers=min(8, num_cpus()))
def teardown(self, stage=None):
self.train = None
self.val = None
class ResNet(LightningModule):
def __init__(self, model, lr=3e-3, mom=0.9, wd=1e-2):
super().__init__()
self.save_hyperparameters(ignore='model')
self.model = model()
self.loss_fn = LabelSmoothingCrossEntropy()
self.accuracy = torchmetrics.Accuracy()
def forward(self, x):
return self.model(x)
def training_step(self, batch, batch_idx):
x, y = batch
x = self(x)
loss = self.loss_fn(x, y)
self.log("train_loss", loss)
return loss
def evaluate(self, batch, stage=None):
x, y = batch
x = self(x)
loss = self.loss_fn(x, y)
preds = F.softmax(x, dim=-1).argmax(dim=-1)
self.accuracy(preds, y)
if stage:
self.log(f"{stage}_loss", loss, prog_bar=True, on_epoch=True)
self.log(f"{stage}_acc", self.accuracy, prog_bar=True, on_epoch=True)
def validation_step(self, batch, batch_idx):
self.evaluate(batch, "val")
def test_step(self, batch, batch_idx):
self.evaluate(batch, "test")
def configure_optimizers(self):
optimizer = AdamW(self.parameters(),lr=self.hparams.lr, eps=1e-5)
scheduler_dict = {
"scheduler": OneCycleLR(
optimizer=optimizer,
max_lr =self.hparams.lr,
total_steps=self.num_training_steps
),
"interval": "step",
"frequency": 1
}
return {"optimizer": optimizer, "lr_scheduler": scheduler_dict}
# lightly modified from rohitgr7 https://github.com/PyTorchLightning/pytorch-lightning/issues/10760
@property
def num_training_steps(self) -> int:
"""Total training steps inferred from datamodule and devices."""
if self.trainer.num_training_batches != float('inf'):
dataset_size = self.trainer.num_training_batches
else:
print('Requesting dataloader...')
dataset_size = len(self.trainer._data_connector._train_dataloader_source.dataloader())
if isinstance(self.trainer.limit_train_batches, int):
dataset_size = min(dataset_size, self.trainer.limit_train_batches)
else:
dataset_size = int(dataset_size * self.trainer.limit_train_batches)
accelerator_connector = self.trainer._accelerator_connector
if accelerator_connector.use_ddp2 or accelerator_connector.use_dp:
effective_devices = 1
else:
effective_devices = self.trainer.devices
effective_devices = effective_devices * self.trainer.num_nodes
effective_batch_size = self.trainer.accumulate_grad_batches * effective_devices
max_estimated_steps = math.ceil(dataset_size // effective_batch_size) * self.trainer.max_epochs
max_estimated_steps = min(max_estimated_steps, self.trainer.max_steps) if self.trainer.max_steps != -1 else max_estimated_steps
return max_estimated_steps
def train_pl(model, epochs, name, size):
resnet = ResNet(model)
imagenette = ImagenetteDataModule(size, False, img_bs)
wandb_logger = WandbLogger(project="sagecolab", name=f'{name} {plat} {GPU} fp16', log_model=False)
trainer = Trainer(gpus=1, precision=16, max_epochs=epochs, num_sanity_val_steps=0,
benchmark=True, profiler="simple", logger=wandb_logger, enable_checkpointing=False)
trainer.fit(resnet, imagenette)
wandb.log({}) # ensure sync of last step
wandb.finish()
trainer, resnet, imagenette= None, None, None
gc.collect()
torch.cuda.empty_cache()
# ## Fastai Imagenette
# +
imagewoof_stats = ([0.496,0.461,0.399],[0.257,0.249,0.258])
imagenette_stats = ([0.465,0.458,0.429],[0.285,0.28,0.301])
def get_imagenette_dls(size, woof, bs, sh=0., augs=None, workers=None, stats=True):
if size<=224: path = URLs.IMAGEWOOF_320 if woof else URLs.IMAGENETTE_320
else : path = URLs.IMAGEWOOF if woof else URLs.IMAGENETTE
source = untar_data(path)
if workers is None: workers = min(8, num_cpus())
batch_tfms = []
if stats:
if woof:
batch_tfms += [Normalize.from_stats(*imagewoof_stats)]
else:
batch_tfms += [Normalize.from_stats(*imagenette_stats)]
if augs: batch_tfms += augs
if sh: batch_tfms.append(RandomErasing(p=0.3, max_count=4, sh=sh))
dblock = DataBlock(blocks=(ImageBlock, CategoryBlock),
splitter=GrandparentSplitter(valid_name='val'),
get_items=get_image_files, get_y=parent_label,
item_tfms=[RandomResizedCrop(size, min_scale=0.35), FlipItem(0.5)],
batch_tfms=batch_tfms)
return dblock.dataloaders(source, path=source, bs=bs, num_workers=workers)
# -
def train_imagenette_fastai(model, epochs, name, size, precision=[16, 32], augs=None):
for fp in precision:
dls = get_imagenette_dls(size, False, img_bs, augs=augs)
train_fastai(dls, model(), epochs, name, precision=fp)
# ## Fastai-Blurr IMDB
def get_imdb_dls(model_name, bs):
dataset = load_dataset('imdb')
df = pd.DataFrame(dataset['train'])
df['is_valid'] = False
df = df.sample(frac=0.2, random_state=42)
df2 = pd.DataFrame(dataset['test'])
df2['is_valid'] = True
df2 = df2.sample(frac=0.2, random_state=42)
df = df.append(df2, ignore_index=True)
config = AutoConfig.from_pretrained(model_name)
config.num_labels = len(df['label'].unique())
hf_arch, hf_config, hf_tokenizer, hf_model = BLURR.get_hf_objects(model_name, model_cls=AutoModelForSequenceClassification, config=config)
dblock = DataBlock(blocks=(HF_TextBlock(hf_arch, hf_config, hf_tokenizer, hf_model), CategoryBlock),
get_x=ColReader('text'),
get_y=ColReader('label'),
splitter=ColSplitter())
return hf_model, dblock.dataloaders(df, bs=bs, workers=min(8, num_cpus()))
def train_imdb_fastai(model_name, epochs, name, precision=[16, 32]):
for fp in precision:
hf_model, dls = get_imdb_dls(model_name, nlp_bs if fp==16 else int(nlp_bs/2))
model = HF_BaseModelWrapper(hf_model)
train_fastai(dls, model, epochs, name, splitter=hf_splitter, cbs=[HF_BaseModelCallback], precision=fp)
# ## Generic Fastai Training
def train_fastai(dls, model, epochs, name, splitter=trainable_params, cbs=[], precision=16):
run = wandb.init(project="sagecolab", name=f'{name} {plat} {GPU} fp{precision}')
learn = Learner(dls, model, cbs=cbs, splitter=splitter).profile()
if precision==16:
learn.to_fp16()
learn.fit_one_cycle(epochs, 3e-3, cbs=[WandbCallback(log=None, log_preds=False, log_model=False)])
run.finish()
learn.dls, learn = None, None
gc.collect()
torch.cuda.empty_cache()
# # Training
# ## Train Imagenette Fastai
model = partial(xse_resnet50, n_out=10)
train_imagenette_fastai(model, 4, name='XSEResNet50', size=224)
model = partial(xresnet18, n_out=10, act_cls=nn.Mish)
train_imagenette_fastai(model, 4, name='XResNet18 128', size=128, precision=[16])
model = partial(xse_resnext50, n_out=10, act_cls=nn.Mish)
train_imagenette_fastai(model, 4, name='XSEResNeXt50', size=256, precision=[16])
# ## Train IMDB
train_imdb_fastai('roberta-base', 4, 'Roberta')
# ## Train Imagenette PyTorch Lightning
model = partial(timm.create_model, model_name='resnet50', pretrained=False, num_classes=10)
train_pl(model, 4, name='ResNet50', size=256)
| 2021/sagemaker_colab/Colab_Pro_High_RAM_Benchmark_T4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # **Calculated Features**
# Real datasets often have underlying complex business rules. YData's platform allows a seamless combination of manually defined business rules with powerful synthesis technologies via a feature called **Calculated Features**, improving the quality of the generated datasets.
#
# ## Outline
# * What are **Calculated Features**?
# * An example dataset: loans
# * Exploring the dataset
# * Synthesizing data with and without **Calculated Features**
# * Evaluating the synthetic data
#
# ## What are **Calculated Features**?
# Features that are based on business rules and typically have a deterministic nature, through a combination of pre-existing features.
#
# These calculated features, regardless of complexity, can be directly synthesized. However, the quality of the synthetic features can be improved if we infuse our models with some of the business logic we know right from the start.
#
# In this notebook, we will exemplify how business rules for feature calculation can be passed to YData's `RegularSynthesizer` and applied to the synthesis process as a way to improve fidelity.
#
# ## An example dataset: loans
# For this example we have chosen a loans dataset from *LendingClub* loans. The original file can be found [here](https://data.world/lpetrocelli/lendingclub-loan-data-2017-q-1).
#
# Loans, as a heavily procedural financial instrument, have of course underlying business logic. We expect our dataset to reflect some of these, like for instance: calculation of revolving credit utilization rate, monthly installment values and total payments.
#
# ## Exploring the dataset
# First, let's read the dataset using a Google Cloud Connector, inspect the data and check, if, as expected, the business logic we mentioned is verified.
# +
from warnings import filterwarnings
filterwarnings('ignore')
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
from ydata.connectors import GCSConnector
from ydata.synthesizers.regular import RegularSynthesizer
from ydata.metadata.metadata import Metadata
from ydata.utils.data_types import DataType
from utils import viz_main, viz_side_by_side # a couple of visualization utilities
# Reading our data file from the GCS cloud
connector = GCSConnector(project_id='ydatasynthetic')
real_ds = connector.read_file("gs://ydata_testdata/tabular/loan_data/loan_samples.csv.zip", file_type='csv')
# Inspecting head of the dataset
real_ds._train.head().T
# -
# Before verifying the business logic, let's first detail it:
# * The expected revolving balance (product of revolving balance utilization rate by total credit limit) should match the actual revolving balance
# * For any given record, the average payment components should add up to the total payment
# * For any given record, the cumulative payments should add up to the expected payment
#
# The following visualizations will help us us validate these assumptions.
viz_main(real_ds)
# The 3 aspects mentioned are indeed verified in the real data and any synthetic equivalent should exhibit these properties.
#
# Let's check if we can leverage YData's `RegularSynthesizer`'s to achieve just that!
# ## Synthesizing data with and without **Calculated Features**
# ### Setting up
# +
# Creating a metadata object
metadata = Metadata()
metadata(real_ds)
print({k: v.datatype for k, v in metadata.columns.items()})
# Only term feature datatype should be categorical
updated_dtypes = []
for column in metadata.columns.values():
if column.datatype == DataType.CATEGORICAL and column.name != 'term':
metadata.columns[column.name].datatype = DataType.NUMERICAL
updated_dtypes.append(column.name)
print("Updated column dtypes:", {k: v.datatype for k, v in metadata.columns.items() if k in updated_dtypes})
# -
# ### Regular Synthesis
# Direct synthesis is obtained by passing the dataset and metadata objects to the `RegularSynthesizer`'s fit method.
# +
SAMPLE_SIZE = 1000
# Fit a synthesizer without using Calculated Features
synth_reg = RegularSynthesizer()
synth_reg.fit(real_ds, metadata=metadata)
# Obtaining samples
samples_reg = synth_reg.sample(SAMPLE_SIZE)
# -
# ### Synthesis with **Calculated Features**
# To produce the desired **Calculated Features** we will first translate each business rule into a function (either a regular method or a `lambda`) and infuse them into the `RegularSynthesizer`.
# Revolving credit utilization is computed as the quotient of revolving credit balance and the total limit of credit:
# Computes the revolving credit utilization based on the current revolving balance and the credit limit.
get_revolving_util = lambda revol_bal, total_rev_hi_lim : (revol_bal/total_rev_hi_lim).values
# The installment (to be paid each month) is calculated according to the [formula for amortization](https://en.wikipedia.org/wiki/Amortization_calculator):
def get_installment(int_rate, loan_amnt, term):
"Computes the installment values due monthly based on an amortization loan schedule."
n = term.str.rstrip('m').astype('int') # The total number of periods
period_int = int_rate/12 # The adjusted annual interest for the monthly installment periods
return loan_amnt*((period_int*(1+period_int)**n)/((1+period_int)**n-1)).values
# The total payment is computed as the sum of all payment parcels:
def get_total_payment(total_rec_int, total_rec_late_fee, total_rec_prncp):
"Computes total payment as the sum of all payment parcels."
return (total_rec_int + total_rec_late_fee + total_rec_prncp).values
# We can now train and sample from a synthesizer with this additional information:
calculated_features = [
{
'calculated_features': 'revol_util',
'function': get_revolving_util,
'calculated_from': ["revol_bal", "total_rev_hi_lim"]
},
{
'calculated_features': 'installment',
'function': get_installment,
'calculated_from': ["int_rate", "loan_amnt", "term"]
},
{
'calculated_features': 'total_pymnt',
'function': get_total_payment,
'calculated_from': ["total_rec_int", "total_rec_late_fee", "total_rec_prncp"]
},
]
# +
# Fit a synthesizer using Calculated Features
synth_calcft = RegularSynthesizer()
synth_calcft.fit(real_ds, metadata=metadata, calculated_features=calculated_features)
# Obtaining samples
samples_calcft = synth_calcft.sample(SAMPLE_SIZE)
# -
# ## Evaluating the synthetic data
# Let's evaluate the effectiveness of our **Calculated Features** by redoing our EDA visualizations for both cases (with and without the manually defined business logic).
viz_side_by_side({'Regular Synthesis': samples_reg, 'Synthetic data with Calculated Features': samples_calcft})
# The addition of the **Calculated Features** guarantees that the generated data tighthly adheres to our business constraints. Without it, instead of deterministic behaviours, we obtain close approximations. Combining data-driven modelling processes with domain knowledge guarantees we play to the strength's of each: learning the continuous distributions of the underlying features while maintaining the exact nature of business logic when they need to be combined.
# ### Wrapping up
# As the synthetic data sample visualizations showed, by using **Calculated Features** we were able to verify the business rules found in the real data.
#
# Defining **Calculated Features** is easy and allows us to guarantee that the produced samples respect intricate deterministic relationships. You can leverage this feature to make the synthetization models adhere to the specific constraints of your domain, simply by specifying them as functions.
| 5 - synthetic-data-applications/regular-tabular/loans-calculated_features/calculated_features.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Normalization
# The data used in this notebook is lymphocyte data for one patient's B cells and T cells. Here we plot the runs to determine the need for normalization and test normalization techniques.
# First, we import our loader module. This brings the functions defined there (in our repository at ~/load_data.py) into scope so we can use them in this script. Then we can load our data and store it as <code>data_raw</code>.
import load_data
#data_raw = load_data.load_FragPipe()
data_raw=load_data.load_max_quant()
# +
import matplotlib.pyplot as plt
import seaborn as sns
plt.title("Raw data")
colors = ["#0066ff","#0066ff","#0066ff","#0066ff","#0066ff","orange","orange","orange","orange", "orange"]
sns.set(font_scale=1.5)
figure = sns.boxplot(data=data_raw, width=.5, palette=colors)
figure.set_ylabel("Intensity")
figure.set_yscale("log")
plt.setp(figure.get_xticklabels(), rotation=45,horizontalalignment='right')
plt.show()
# +
def dist(ser, log=False):
f=sns.distplot(ser, hist=False)
f.set_xlabel("Intensity")
f.set_ylabel("Frequency")
if log: f.set_xscale("log", basex=2)
plt.title("Raw data")
data_raw.apply(dist)
plt.show()
plt.title("Raw data on log scale")
data_raw.apply(dist, log=True)
plt.show()
# -
# Global median normalization
# https://datascienceplus.com/proteomics-data-analysis-2-3-data-filtering-and-missing-value-imputation/
from numpy import nan
from numpy import log2
from statistics import median
data_log2 = log2(data_raw.replace(0,nan))
data_log2_medNorm = data_log2.apply(lambda series: series-median(series.dropna()))
# +
plt.title("Normalized data")
figure = sns.boxplot(data=data_log2_medNorm, width=.5, palette=colors)
figure.set_ylabel("Intensity")
plt.setp(figure.get_xticklabels(), rotation=45,horizontalalignment='right')
plt.show()
# +
plt.title("Normalized data")
sns.set_style("white")
data_log2_medNorm.apply(dist)
plt.show()
# -
# The global median normalization works well with this data. The dataset is now log2 and scaled to zero.
medians = data_log2.apply(lambda series: median(series.dropna()))
global_median = median(medians.dropna())
data_normalized = data_log2.apply(lambda series: series-median(series.dropna())+global_median)
data_normalized.apply(dist)
plt.show()
# Here, the data is centered at the global median instead of 0.
# #### Normalization reveals biologial grouping
#
# PCA plots to compare the raw and normalized data clustering. This shows the benefit of normalizing to reveal biological differences. Prior to normalization, the batch effects or differences between runs due to differences in number of cells or instrument sensitivity obscure clustering.
# +
from sklearn.decomposition import PCA
import matplotlib.pyplot as plt
import math
from numpy import isnan
import numpy as np
import pandas as pd
sns.set_style("white")
pca = PCA(n_components=5)
alist=data_raw.values.flatten()
alist= [a for a in alist if not isnan(a)]
nan_appoximate = float(alist[math.ceil(float(len(alist))*.01)])
pca_result = pca.fit_transform(np.nan_to_num(data_raw.transpose(), nan=nan_appoximate))
cell_types = [" B_"," T_"] #Cell types; these strings are in the sample names of those types
samples=np.array(data_raw.columns.values)
for cell_type in cell_types:
cells_of_type = list(i for i,s in enumerate(samples) if cell_type in s)
plt.scatter(pca_result[cells_of_type,0],pca_result[cells_of_type,1])
plt.title("Raw Data")
plt.legend(['B cells', "T cells"])
plt.show()
# +
pca = PCA(n_components=5)
alist=data_log2_medNorm.values.flatten()
alist= [a for a in alist if not isnan(a)]
nan_appoximate = float(alist[math.ceil(float(len(alist))*.01)])
pca_result = pca.fit_transform(np.nan_to_num(data_log2_medNorm.transpose(), nan=nan_appoximate))
cell_types = [" B_"," T_"] #Cell types; these strings are in the sample names of those types
for cell_type in cell_types:
cells_of_type = list(i for i,s in enumerate(samples) if cell_type in s)
plt.scatter(pca_result[cells_of_type,0],pca_result[cells_of_type,1])
plt.title("Normalized data")
plt.legend(['B cells', "T cells"], loc='upper right', bbox_to_anchor=(1.5, 1))
plt.show()
# -
| Normalization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# name: python2
# ---
# + [markdown] id="AC8adBmw-5m3" colab_type="text"
# This is an Earth Engine <> TensorFlow demonstration notebook. The default public runtime already has the tensorflow libraries we need installed. The first step is to verify that by importing the TensorFlow library. Run the code in the cell by clicking the run button on the left (hover on the `[ ]`).
#
# **NOTE: The cloud storage buckets used here are no longer writable. (Temporary access was for the Earth Engine Users Summit 2018 only). Please substitute your own cloud storage bucket.**
#
#
# + id="i1PrYRLaVw_g" colab_type="code" colab={} cellView="code"
#@title Import tensorflow library
import tensorflow as tf
# + [markdown] id="46iNFM8lV6kP" colab_type="text"
# Check the TensorFlow install by running a hello world operation (from the [Cloud ML example](https://cloud.google.com/ml-engine/docs/tensorflow/getting-started-training-prediction#run_a_simple_tensorflow_python_program)):
# + id="R27Tc9zqerkG" colab_type="code" colab={} cellView="code"
#@title Hello World TensorFlow
hello = tf.constant('Hello world!')
with tf.Session() as sess:
print sess.run(hello)
# + [markdown] id="--JtxLXsUpVA" colab_type="text"
# Note that you can use "magic" commands by prepending an `!` to a bash command. For example, here we will install a python library to enable us to connect to Google Drive. Learn more about magic functions from **Code snippets** to the left. The objective here is to enable access to thinhs in Drive that you may have exported from Earth Engine.
# + id="sYyTIPLsvMWl" colab_type="code" colab={} cellView="code"
#@title Install the PyDrive library
# This only needs to be done once per notebook.
# !pip install -U PyDrive
# + [markdown] id="qptAXhKmXo_J" colab_type="text"
# We need to import some authentication APIs so that we can read from Drive and/or a cloud storage bucket. See the **Code snippets** to the left.
# + id="5qMKG1hEXuML" colab_type="code" colab={} cellView="code"
#@title Import authentication libraries
from google.colab import auth
from pydrive.auth import GoogleAuth
from pydrive.drive import GoogleDrive
from oauth2client.client import GoogleCredentials
# + [markdown] id="dEM3FP4YakJg" colab_type="text"
# **Authentication**. The following will trigger the browser dance to authenticate. Follow the link, copy the code from another browser window to the indicated field, then press return. You should use the same account to authenticate here that you used to join the training group (which is hopefully the same account you use to login to Earth Engine, otherwise the exports will end up in the Drive of another account.)
# + id="CHsEU90Xyjmo" colab_type="code" colab={} cellView="code"
#@title Authenticate
auth.authenticate_user()
gauth = GoogleAuth()
gauth.credentials = GoogleCredentials.get_application_default()
drive = GoogleDrive(gauth)
# + [markdown] id="ZcjQnHH8zT4q" colab_type="text"
# We've already generated some training data in Earth Engine. Specifically, these are exported testing and training data from a very simple classification demo. The script exports a training dataset, a testing dataset and the image data (in TFRecord format) on which to make predictions:
#
# https://code.earthengine.google.com/a7ed957f3034825a54b6b546b8c5ce83
#
# RUN THE EXPORTS
#
# ---
#
# Note that the script exports to two places: Your drive account and a public cloud storage bucket. You can grab the files you need from either place, as demonstrated in the following sections. Here we'll use Drive, but note that code to use Cloud Storage is also here in case you need it.
# + id="EazQzf8lzLF3" colab_type="code" colab={} cellView="code"
#@title Load training/testing data from Earth Engine exports
# Specify the training file exported from EE.
# If you wish to use your own data, then
# replace the file ID, below, with your own file.
trainFileId = '1bLHhjGjKYXtdK_XAwC9636ZuxAKuGlmO' # nclinton version!
trainDownload = drive.CreateFile({'id': trainFileId})
# Create a local file of the specified name.
tfrTrainFile = 'training.tfrecord.gz'
trainDownload.GetContentFile(tfrTrainFile)
print 'Successfully downloaded training file?'
print tf.gfile.Exists(tfrTrainFile)
# Specify the test file.
# If you wish to use your own data, then
# replace the file ID, below, with your own file.
testFileId = '1PWakg7ygx-vRm5O_QKup6GIJup8LIvLy' # nclinton version!
testDownload = drive.CreateFile({'id': testFileId})
# Creates a local file of the specified name.
tfrTestFile = 'testing.tfrecord.gz'
testDownload.GetContentFile(tfrTestFile)
print 'Successfully downloaded testing file?'
print tf.gfile.Exists(tfrTestFile)
print 'Content of the working directory:'
# !ls
# + [markdown] id="LS4jGTrEfz-1" colab_type="text"
# Here we are going to read from the Drive file into a `tf.data.Dataset`. ([Slide](https://docs.google.com/presentation/d/1fEf-oScgbC9zjbzI3K3jUlHf4JdmoSLFiG_H491FUmk/edit#slide=id.g3b76860e75_0_63)). Check that you can read examples from the file. The purpose here is to ensure that we can read from the file without an error. The actual content is not necessarily human readable.
# + id="T3PKyDQW8Vpx" colab_type="code" colab={} cellView="code"
#@title Inspect the TFRecord dataset
driveDataset = tf.data.TFRecordDataset(tfrTrainFile, compression_type='GZIP')
iterator = driveDataset.make_one_shot_iterator()
foo = iterator.get_next()
with tf.Session() as sess:
print sess.run([foo])
# + [markdown] id="BrDYm-ibKR6t" colab_type="text"
# Define the structure of your data. This includes the names of the bands you originally exported from Earth Engine and the name of the class property. Unfortunately, these are called *features* in the TensorFlow context (not to be confused with an `ee.Feature`). ([Slide](https://docs.google.com/presentation/d/1fEf-oScgbC9zjbzI3K3jUlHf4JdmoSLFiG_H491FUmk/edit#slide=id.g3b76860e75_0_67)). Think of `columns` as a placeholder for the data that you're going to read in.
# + id="-6JVQV5HKHMZ" colab_type="code" colab={} cellView="code"
#@title Define the structure of the training/testing data
# Names of the features.
bands = ['B2', 'B3', 'B4', 'B5', 'B6', 'B7']
label = 'landcover'
featureNames = list(bands)
featureNames.append(label)
# Feature columns
columns = [
tf.FixedLenFeature(shape=[1], dtype=tf.float32) for k in featureNames
]
# Dictionary with names as keys, features as values.
featuresDict = dict(zip(featureNames, columns))
print featuresDict
# + [markdown] id="QNfaUPbcjuCO" colab_type="text"
# Now we need to make a parsing function. The parsing function reads data from a serialized example proto into a dictionary in which the keys are the feature names and the values are the tensors storing the value of the feature for that example. ([TF reference](https://www.tensorflow.org/programmers_guide/datasets#parsing_tfexample_protocol_buffer_messages), [Cloud ML reference](https://github.com/GoogleCloudPlatform/cloudml-samples/blob/master/cloudml-template/template/trainer/input.py#L61)).
#
# Here we make a parsing function for the TFRecord files we've been generating. The check at the end is to print a single parsed example.
# + id="x2Q0g3fBj2kD" colab_type="code" colab={} cellView="code"
#@title Make and test a parsing function
def parse_tfrecord(example_proto):
parsed_features = tf.parse_single_example(example_proto, featuresDict)
labels = parsed_features.pop(label)
return parsed_features, tf.cast(labels, tf.int32)
# Map the function over the dataset
parsedDataset = driveDataset.map(parse_tfrecord, num_parallel_calls=5)
iterator = parsedDataset.make_one_shot_iterator()
foo = iterator.get_next()
with tf.Session() as sess:
print sess.run([foo])
# + [markdown] id="xLCsxWOuEBmE" colab_type="text"
# Another thing we might want to do as part of the input process is to create new features, for example NDVI. Here are some helper functions for that. Note that a and b are expected to be shape=[1] tensors and features is s dictionary of input tensors keyed by feature name.
# + id="lT6v2RM_EB1E" colab_type="code" colab={} cellView="code"
#@title Make functions to add additional features
# Compute normalized difference of two inputs. If denomenator is zero, add a small delta.
def normalizedDifference(a, b):
nd = (a - b) / (a + b)
nd_inf = (a - b) / (a + b + 0.000001)
return tf.where(tf.is_finite(nd), nd, nd_inf)
# Add normalized differences and 3-D coordinates to the dataset. Shift the label to zero.
def addFeatures(features, label):
features['NDVI'] = normalizedDifference(features['B5'], features['B4'])
return features, label
# + [markdown] id="sz7kX5alFi4K" colab_type="text"
# Now we need to define an input function that will feed data from a file into a TensorFlow model. Putting together what we've done so far, here is the complete function for input, parsing and feature engineering:
# + id="rktz1DqqFlh_" colab_type="code" colab={} cellView="code"
#@title Make an input function
def tfrecord_input_fn(fileName,
numEpochs=None,
shuffle=True,
batchSize=None):
dataset = tf.data.TFRecordDataset(fileName, compression_type='GZIP')
# Map the parsing function over the dataset
dataset = dataset.map(parse_tfrecord, num_parallel_calls=5)
# Add additional features.
dataset = dataset.map(addFeatures)
# Shuffle, batch, and repeat.
if shuffle:
dataset = dataset.shuffle(buffer_size=batchSize * 10)
dataset = dataset.batch(batchSize)
dataset = dataset.repeat(numEpochs)
# Make a one-shot iterator.
iterator = dataset.make_one_shot_iterator()
features, labels = iterator.get_next()
return features, labels
# + [markdown] id="t9pWa54oG-xl" colab_type="text"
# The classifier we will use is a deep neural network (DNN) from the [`tf.estimator` package](https://www.tensorflow.org/api_docs/python/tf/estimator). ([Slide](https://docs.google.com/presentation/d/1fEf-oScgbC9zjbzI3K3jUlHf4JdmoSLFiG_H491FUmk/edit#slide=id.g3b76860e75_0_59)). First, define the input features, including the newly created NDVI column. Here we specify an optimizer so that we can also set the learning rate. Specify 7 nodes in the first hidden layer, 7 in the second and 5 in the third. These are arbitrary demonstration numbers.
#
# Lastly, train the classifier. In order to pass the classifier a single argument input function, use a lambda function to specify the number of epochs and batch size. You could also specify the number of training steps here where steps = N / batchSize for a single epoch ([reference](https://developers.google.com/machine-learning/glossary/#epoch)).
# + id="OCZq3VNpG--G" colab_type="code" colab={} cellView="code"
#@title Make and train a classifier
inputColumns = {tf.feature_column.numeric_column(k) for k in ['B2', 'B3', 'B4', 'B5', 'B6', 'B7', 'NDVI']}
learning_rate = 0.05
optimizer = tf.train.AdagradOptimizer(learning_rate)
classifier = tf.estimator.DNNClassifier(feature_columns=inputColumns,
hidden_units=[5, 7, 5],
n_classes=3,
model_dir='output',
optimizer=optimizer)
classifier.train(input_fn=lambda: tfrecord_input_fn(fileName=tfrTrainFile, numEpochs=8, batchSize=1))
# + [markdown] id="Pa4ex_4eKiyb" colab_type="text"
# Now that we have a trained classifier, we can evaluate it using the test set. To do that, use the same input function on a different file. Since this is the test set, just use one epoch and don't shuffle. Here we just print the overall accuracy.
# + id="tE6d7FsrMa1p" colab_type="code" colab={} cellView="code"
#@title Evaluate the classifier
accuracy_score = classifier.evaluate(
input_fn=lambda: tfrecord_input_fn(fileName=tfrTestFile, numEpochs=1, batchSize=1, shuffle=False)
)['accuracy']
# + [markdown] id="kUYADjdrc4Ie" colab_type="text"
# Training an estimator triggers storage of the state of the final model. Unless you want subsequent runs to update previous model state, you may want to run the following (you will have to uncomment it first) to get rid of old model output. Use with caution!
# + id="WSEnOAdGcYxL" colab_type="code" colab={} cellView="code"
#@title Optionally delete model output
# # !rm -rf output
# + [markdown] id="lGD4hzhAb4PR" colab_type="text"
# **Optional**. The following code cell checks that the classifier can work by training it and testing it on the files stored in the cloud storage bucket. To see the code, toggle the form with the control to the right.
# + id="9nBrxvvCUDbi" colab_type="code" colab={} cellView="form"
#@title Optional Cloud Storage way (No need to run)
inputColumns = {tf.feature_column.numeric_column(k) for k in ['B2', 'B3', 'B4', 'B5', 'B6', 'B7', 'NDVI']}
learning_rate = 0.05
optimizer = tf.train.AdagradOptimizer(learning_rate)
classifier = tf.estimator.DNNClassifier(feature_columns=inputColumns,
hidden_units=[5, 7, 5],
n_classes=3,
model_dir='output',
optimizer=optimizer)
# TensorFlow can read directly from a cloud storage location, so all we need to do is specify the path.
tfrTrainFileCloud = 'gs://nclinton-training-temp/tf_demo_train_9a26cef21ab34f6257d0a250882124fcee_export.tfrecord.gz'
tfrTestFileCloud = 'gs://nclinton-training-temp/tf_demo_test_9a26cef21ab34f6257d0a250882124fcee_export.tfrecord.gz'
# Just check that you can see file(s):
print tf.gfile.Exists(tfrTrainFileCloud)
print tf.gfile.Exists(tfrTestFileCloud)
# Train and test, passing the cloud storage path into the input function.
classifier.train(input_fn=lambda: tfrecord_input_fn(fileName=tfrTrainFileCloud, numEpochs=8, batchSize=1))
accuracy_score = classifier.evaluate(
input_fn=lambda: tfrecord_input_fn(fileName=tfrTestFileCloud, numEpochs=1, batchSize=1, shuffle=False)
)['accuracy']
# + [markdown] id="Nej2HDZM-G0H" colab_type="text"
# Get predictions on the evaluation dataset. Note that we're going to make two iterators for this dataset. The first one is just to see what's in there, to do a sanity check on the output. We'll use the second one, below, to write the predictions to a TFRecord file.
#
# Note that you can get both the predicted class and support probabilities for that classification.
# + id="12jRDVG4BpJ8" colab_type="code" colab={} cellView="code"
#@title Make predictions on the test data
import itertools
# Do the prediction from the trained classifier.
checkPredictions = classifier.predict(
input_fn=lambda: tfrecord_input_fn(fileName=tfrTestFile, numEpochs=1, batchSize=1, shuffle=False)
)
# Make a couple iterators.
iterator1, iterator2 = itertools.tee(checkPredictions, 2)
# Iterate over the predictions, printing the class_ids and posteriors.
for pred_dict in iterator1:
class_id = pred_dict['class_ids']
probability = pred_dict['probabilities']
print class_id, probability
# + [markdown] id="YRJsMMq_h7ZM" colab_type="text"
# **Optional**. To write into a TFRecord file, it helps to have alittle understanding of how the records are stored. This next example is to practice building a single record and writing it. Specifically, define a `tf.train.Example` [protocol buffer](https://developers.google.com/protocol-buffers/) and write it to a file.
# + id="66V-P88AjKRb" colab_type="code" colab={} cellView="code"
#@title Demonstration of writing an Example
checkFilename = 'check.TFRecord'
writer = tf.python_io.TFRecordWriter(checkFilename)
example = tf.train.Example(
features=tf.train.Features(
feature={
'prediction': tf.train.Feature(
int64_list=tf.train.Int64List(
value=[1])),
'posteriors': tf.train.Feature(
float_list=tf.train.FloatList(
value=[0.1, 0.2, 0.3]))
}
))
writer.write(example.SerializeToString())
writer.flush()
writer.close()
# + [markdown] id="_BtyFe6Oc7jo" colab_type="text"
# Now let's check that we can read our example back out of the file.
# + id="No8107mzrpbc" colab_type="code" colab={} cellView="code"
#@title Demonstration of reading an Example
checkDataset = tf.data.TFRecordDataset('check.TFRecord')
checkDict = {
'prediction': tf.FixedLenFeature(shape=[1], dtype=tf.int64),
'posteriors': tf.FixedLenFeature(shape=[3], dtype=tf.float32),
}
checkParsed = checkDataset.map(
lambda example_proto: tf.parse_single_example(example_proto, checkDict))
iterator = checkParsed.make_one_shot_iterator()
foo = iterator.get_next()
with tf.Session() as sess:
print sess.run([foo])
# + [markdown] id="8iW4iPSkrrHb" colab_type="text"
# Now iterate over the predictions on the test data and try writing all those into a file. For each prediction, we make a new `tf.Example` proto out of the prediction data, then write it. Finally execute a shell command to see if we've successfully written the file.
#
# See:
#
# https://github.com/tensorflow/tensorflow/blob/r1.8/tensorflow/core/example/feature.proto
#
# https://github.com/tensorflow/tensorflow/blob/master/tensorflow/examples/how_tos/reading_data/convert_to_records.py
# + id="maMPR8yurm-j" colab_type="code" colab={} cellView="code"
#@title Demonstration of writing predictions to a file
outputFilename = 'checkPredictions.TFRecord'
writer = tf.python_io.TFRecordWriter(outputFilename)
for pred_dict in iterator2:
example = tf.train.Example(
features=tf.train.Features(
feature={
'prediction': tf.train.Feature(
int64_list=tf.train.Int64List(
value=pred_dict['class_ids'])),
'probabilities': tf.train.Feature(
float_list=tf.train.FloatList(
value=pred_dict['probabilities']))
}
))
writer.write(example.SerializeToString())
writer.close()
# !ls -Al
# + [markdown] id="nhHrnv3VR0DU" colab_type="text"
# Now it's time to classify the image from Earth Engine. The way this happens is by exporting an image as a TFRecord file [announcement doc](https://docs.google.com/document/d/1njY_MKvXELEWvDaXmA56TFSteeiSytkziHBD9Pr8Q9I/edit?usp=sharing). [The script for exporting the training and testing data](https://code.earthengine.google.com/a7ed957f3034825a54b6b546b8c5ce83) also exports a piece of the composite for classification. Specifically, `Export.image` now accepts `'TFRecord'` for `fileFormat`.
#
# Theres some other new stuff in that export. Specifically, note that we're exporting pixels in 256x256 patches for efficiency. Also note that the image gets split into multiple TFRecord files in its destination folder.
# + [markdown] id="BdQrQIzKvEqp" colab_type="text"
# Because there are multiple files that make up the image, use the Google PyDrive library to search for the files that match a particular prefix string. Specifically, this is the name you specified in the JavaScript for the exported files. Download all the Drive files that match that field, one of which is the JSON that we don't need as input to the model (but will need for import to Earth Engine after we've made predictions). Lastly, print the list of filenames for a reality check.
#
# See https://pythonhosted.org/PyDrive/filelist.html for pyDrive docs. Here's where you can find the info on that query expression: https://developers.google.com/drive/api/v2/search-parameters#file_fields
# + id="R_7a_r9tuo1U" colab_type="code" colab={} cellView="code"
#@title Find the exported image and JSON files in Drive
file_list = drive.ListFile({
# You have to know this base filename from wherever you did the export.
'q': 'title contains "tf_demo_image_9a26cef21ab34f6257d0a250882124fc"'
}).GetList()
fileNames = []
jsonFile = None
for gDriveFile in file_list:
title = gDriveFile['title']
# Download to the notebook server VM.
gDriveFile.GetContentFile(title)
# If the filename contains .gz, it's part of the image.
if (title.find('gz') > 0):
fileNames.append(gDriveFile['title'])
if (title.find('json') > 0):
jsonFile = title
# Make sure the files are in the right order.
fileNames.sort()
# Check the list of filenames to ensure there's nothing unintentional in there.
print(fileNames)
# + id="PCnHyu53rQs4" colab_type="code" colab={} cellView="form"
#@title Optional Cloud Storage way (No need to run)
# We'll need this for importing the classified image back to Earth Engine.
jsonFile = 'gs://nclinton-training-temp/tf_demo_image_9a26cef21ab34f6257d0a250882124fcmixer.json'
# TensorFlow can read directly from a cloud storage bucket.
# Ensure that the files are in order.
fileNames = [
'gs://nclinton-training-temp/tf_demo_image_9a26cef21ab34f6257d0a250882124fc00000.tfrecord.gz',
'gs://nclinton-training-temp/tf_demo_image_9a26cef21ab34f6257d0a250882124fc00001.tfrecord.gz'
]
print(fileNames)
# + [markdown] id="6xyzyPPJwpVI" colab_type="text"
# We can feed this list of files directly to the Dataset constructor to make a combined dataset. However, the the input function is slightly different from the previous ones. Mainly, this is because the pixels are written into records as patches, we need to read the patches in as one big tensor (one patch for each band), then flatten them into lots of little tensors. Once the input function is defined that can handle the shape of the image data, all you need to do is feed it directly to the trained model to make predictions.
# + id="tn8Kj3VfwpiJ" colab_type="code" colab={} cellView="code"
#@title Make an input function for exported image data
# You have to know the following from your export.
PATCH_WIDTH = 256
PATCH_HEIGHT = 256
PATCH_DIMENSIONS_FLAT = [PATCH_WIDTH * PATCH_HEIGHT, 1]
bands = ['B2', 'B3', 'B4', 'B5', 'B6', 'B7']
# Note that the tensors are in the shape of a patch, one patch for each band.
columns = [
tf.FixedLenFeature(shape=PATCH_DIMENSIONS_FLAT, dtype=tf.float32) for k in bands
]
featuresDict = dict(zip(bands, columns))
# This function adds NDVI to a feature that doesn't have a label.
def addServerFeatures(features):
return addFeatures(features, None)[0]
# This input function reads in the TFRecord files exported from an image.
# Note that because the pixels are arranged in patches, we need some additional
# code to reshape the tensors.
def predict_input_fn(fileNames):
# Note that you can make one dataset from many files by specifying a list.
dataset = tf.data.TFRecordDataset(fileNames, compression_type='GZIP')
def parse_image(example_proto):
parsed_features = tf.parse_single_example(example_proto, featuresDict)
return parsed_features
dataset = dataset.map(parse_image, num_parallel_calls=5)
# Break our long tensors into many littler ones
dataset = dataset.flat_map(lambda features: tf.data.Dataset.from_tensor_slices(features))
# Add additional features (NDVI).
dataset = dataset.map(addServerFeatures)
# Read in batches corresponding to patch size.
dataset = dataset.batch(PATCH_WIDTH * PATCH_HEIGHT)
# Make a one-shot iterator.
iterator = dataset.make_one_shot_iterator()
return iterator.get_next()
# Do the prediction from the trained classifier.
predictions = classifier.predict(
input_fn=lambda: predict_input_fn(fileNames)
)
# + [markdown] id="RNwwKeGtkhh_" colab_type="text"
# Name the TFRecord file you're going to create with a unique identifier for you (like your username). We'll write this file directly into a temporary cloud storage bucket created for this training. *The bucket will be deleted daily, so don't store anything in there*.
# + id="xuQ1s6YJhwOP" colab_type="code" colab={} cellView="code"
#@title Define output names
# INSERT YOUR USERNAME HERE (e.g. nclinton):
username = ''
baseName = 'gs://nclinton-training-temp/' + username
outputImageFile = baseName + '_predictions.TFRecord'
outputJsonFile = baseName + '_predictions.json'
print 'Writing to: ' + outputImageFile
# + [markdown] id="bPU2VlPOikAy" colab_type="text"
# We already have the predictions as a list. Iterate over them as we did previously, except with some additional code to handle the shape. Specifically, we need to write the pixels into the file as patches in the same order they came out. (Note: 5,620,989 pixels)
# + id="kATMknHc0qeR" colab_type="code" colab={} cellView="code"
#@title Make predictions on the image data, write to a file
iter1, iter2 = itertools.tee(predictions, 2)
# Iterate over the predictions, printing the class_ids and posteriors.
# This is just to examine the first prediction.
for pred_dict in iter1:
print pred_dict
break # OK
# Instantiate the writer.
writer = tf.python_io.TFRecordWriter(outputImageFile)
# Every patch-worth of predictions we'll dump an example into the output
# file with a single feature that holds our predictions. Since are predictions
# are already in the order of the exported data, our patches we create here
# will also be in the right order.
patch = [[], [], [], []]
curPatch = 1
for pred_dict in iter2:
patch[0].append(pred_dict['class_ids'])
patch[1].append(pred_dict['probabilities'][0])
patch[2].append(pred_dict['probabilities'][1])
patch[3].append(pred_dict['probabilities'][2])
# Once we've seen a patches-worth of class_ids...
if (len(patch[0]) == PATCH_WIDTH * PATCH_HEIGHT):
print('Done with patch ' + str(curPatch) + '...')
# Create an example
example = tf.train.Example(
features=tf.train.Features(
feature={
'prediction': tf.train.Feature(
int64_list=tf.train.Int64List(
value=patch[0])),
'bareProb': tf.train.Feature(
float_list=tf.train.FloatList(
value=patch[1])),
'vegProb': tf.train.Feature(
float_list=tf.train.FloatList(
value=patch[2])),
'waterProb': tf.train.Feature(
float_list=tf.train.FloatList(
value=patch[3])),
}
)
)
# Write the example to the file and clear our patch array so it's ready for
# another batch of class ids
writer.write(example.SerializeToString())
patch = [[], [], [], []]
curPatch += 1
writer.close()
# + [markdown] id="M6sNZXWOSa82" colab_type="text"
# Note that you should also move the JSON file downloaded earlier and give it the same base name as the TFRecord file with the predictions in it. It's not necessary to do this, but will be helpful in the upload to Earth Engine command.
#
#
# + id="q6yLaut2UYPp" colab_type="code" colab={} cellView="code"
#@title Copy the JSON file to a cloud storage bucket
# Copy the JSON file so it has the same base name as the image.
# !gsutil cp {jsonFile} {outputJsonFile}
# !gsutil ls gs://nclinton-training-temp
# + [markdown] id="ScjihzmEn1EV" colab_type="text"
# Almost there! Now we have a predictions image, sitting in a cloud storage bucket. The purpose of doing it this way is to enable us to upload the image to Earth Engine from the cloud storage bucket. This can be accomplished with the [Earth Engine command line tool](https://developers.google.com/earth-engine/command_line#upload). But first we need to install the Earth Engine API and authenticate.
# + id="8TLqch_Bjz92" colab_type="code" colab={} cellView="code"
#@title Install the Earth Engine API
# !pip install earthengine-api
# !earthengine authenticate --quiet
# + [markdown] id="Ejxa1MQjEGv9" colab_type="text"
# Follow the link in the output above, copy the authorization link into the code cell below and run it to authenticate Earth Engine.
# + id="LtLqeBZMljga" colab_type="code" colab={} cellView="code"
#@title Authentication for Earth Engine
# !earthengine authenticate --authorization-code=<YOUR CODE HERE>
# + [markdown] id="cgyvGbBxD9-Z" colab_type="text"
# Let's just test the `earthengine` command by looking for help on the upload command.
# + id="HzwiVqbcmJIX" colab_type="code" colab={} cellView="code"
#@title Get earthengine upload help
# !earthengine upload image -h
# + [markdown] id="2ZyCo297Clcx" colab_type="text"
# Now we're ready to move the image file back to Earth Engine. Note that we give both the image TFRecord file and the JSON file as arguments to `earthengine upload`. Here's where it's useful to copy the JSON file to have a consistent basename with the image.
# + id="NXulMNl9lTDv" colab_type="code" colab={} cellView="code"
#@title Upload the classified image to Earth Engine
# Change the filenames to match your personal user folder in Earth Engine.
# e.g. users/nclinton/TF_nclinton_predictions
outputAssetID = 'users/nclinton/TF_foobar_predictions'
# !earthengine upload image --asset_id={outputAssetID} {outputImageFile} {outputJsonFile}
# + id="_vB-gwGhl_3C" colab_type="code" colab={} cellView="code"
#@title Check the status of the asset ingestion
import ee
ee.Initialize()
tasks = ee.batch.Task.list()
print tasks
# + [markdown] id="qMaPrK6OCxxA" colab_type="text"
# Check the output in Earth Engine (nclinton version): https://code.earthengine.google.com/47ba19eedba20fad5d3df28fa2c4be1c
| TF_demo1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import h5py
from PIL import Image
import pandas as pd
from tqdm.notebook import tqdm
from pathlib import Path
import json
fashiongen_h5 = h5py.File('/net/acadia10a/data/zkhan/fashion-gen/fashiongen_256_256_validation.h5', mode='r')
fashiongen_h5.keys()
Image.fromarray(fashiongen_h5['input_image'][0])
fashiongen_h5['input_description'][0]
fashiongen_h5['input_concat_description'][0]
fashiongen_h5['input_name'][0]
Image.fromarray(fashiongen_h5['input_image'][30])
fashiongen_h5['input_description'][30]
fashiongen_h5['input_name'][30]
# # Creating the test set
num_rows = len(fashiongen_h5['input_pose'])
rows = []
for h5_idx in tqdm(range(num_rows)):
row = dict(
h5_index=h5_idx,
pose=fashiongen_h5['input_pose'][h5_idx][0].decode('utf-8'),
product_id=fashiongen_h5['input_productID'][h5_idx][0]
)
rows.append(row)
df = pd.DataFrame(rows)
df
test_set = df.groupby('product_id').first().head(2000).reset_index()
test_set
root = Path('/net/acadia10a/data/zkhan/fashion-gen/')
save_path = root / 'test_images'
test_json = []
save_path.mkdir(exist_ok=True)
pairs_saved = 0
for row in tqdm(test_set.itertuples()):
if pairs_saved >= 1000:
break
try:
image = fashiongen_h5['input_image'][row.h5_index]
image_as_pil = Image.fromarray(image)
image_path = str(save_path / f'{row.h5_index}-{row.product_id}_{row.pose}.jpg')
image_description = fashiongen_h5['input_description'][row.h5_index][0].decode('utf-8')
image_name = fashiongen_h5['input_name'][row.h5_index][0].decode('utf-8')
image_as_pil.save(image_path)
test_json.append(
{
'image': f'test_images/{row.h5_index}-{row.product_id}_{row.pose}.jpg',
'caption': [
image_name
]
}
)
pairs_saved +=1
except UnicodeDecodeError:
continue
with open(save_path / 'test_pairs.json', 'w') as f:
json.dump(test_json, f)
# # Creating the training set
fashiongen_h5 = h5py.File('/net/acadia10a/data/zkhan/fashion-gen/fashiongen_256_256_train.h5', mode='r')
num_images = len(fashiongen_h5['input_productID'])
root = Path('/net/acadia10a/data/zkhan/fashion-gen/')
save_path = root / 'train_images'
train_json = []
save_path.mkdir(exist_ok=True)
pairs_saved = 0
for idx in tqdm(range(num_images)):
try:
#image = fashiongen_h5['input_image'][idx]
#image_as_pil = Image.fromarray(image)
image_path = str(save_path / f'{idx}.jpg')
image_description = fashiongen_h5['input_description'][idx][0].decode('utf-8')
image_name = fashiongen_h5['input_name'][idx][0].decode('utf-8')
#image_as_pil.save(image_path)
train_json.extend(
[
{
'image': f'train_images/{idx}.jpg',
'caption': image_name,
},
{
'image': f'train_images/{idx}.jpg',
'caption': image_description
}
]
)
pairs_saved +=1
except UnicodeDecodeError:
continue
print(pairs_saved)
Path(train_json[0]['image']).name
with open('/net/acadia10a/data/zkhan/fashion-gen/')
# I fucked up earlier, fixing my mistake here.
for idx, pair in tqdm(enumerate(train_json)):
train_json[idx]['image_id'] = f"fashiongen_{Path(pair['image']).stem}"
train_json[0]
with open(save_path / 'train_pairs.json', 'w') as f:
json.dump(train_json, f)
| notebooks/fashion-gen-retrieval.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# + deletable=true editable=true
import numpy as np
from bqplot import *
# + deletable=true editable=true
size = 100
scale = 100.
np.random.seed(0)
x_data = np.arange(size)
y_data = np.cumsum(np.random.randn(size) * scale)
# + [markdown] deletable=true editable=true
# ## Line Chart
# + deletable=true editable=true
x_sc = LinearScale()
y_sc = LinearScale()
ax_x = Axis(label='X', scale=x_sc, grid_lines='solid')
ax_y = Axis(label='Y', scale=y_sc, orientation='vertical', grid_lines='solid')
line = Lines(x=x_data, y=x_data, scales={'x': x_sc, 'y': y_sc})
fig = Figure(axes=[ax_x, ax_y], marks=[line], title='First Example')
fig
# + [markdown] deletable=true editable=true
# This image can be saved by calling the `save_png` function of the `Figure` object:
# + deletable=true editable=true
fig.save_png()
# + [markdown] deletable=true editable=true
# ## Line Chart with dates as x data
# + deletable=true editable=true
dates = np.arange('2005-02', '2005-03', dtype='datetime64[D]')
size = len(dates)
prices = scale + 5 * np.cumsum(np.random.randn(size))
# + deletable=true editable=true
dt_x = DateScale()
lin_y = LinearScale()
x_ax = Axis(label='Date', scale=dt_x, tick_format='%b-%d', grid_lines='solid')
x_ay = Axis(label=('Price'), scale=lin_y, orientation='vertical', tick_format='0.0f', grid_lines='solid')
lc = Lines(x=dates, y=prices, scales={'x': dt_x, 'y': lin_y}, colors=['blue'])
fig = Figure(marks=[lc], axes=[x_ax, x_ay], background_style={'fill': 'lightgreen'},
title_style={'font-size': '20px','fill': 'DarkOrange'}, title='Changing Styles')
fig
# + deletable=true editable=true
fig.background_style = {'fill': 'Black'}
# + [markdown] deletable=true editable=true
# ## Scatter Chart
# + deletable=true editable=true
sc_x = LinearScale()
sc_y = LinearScale()
scatter = Scatter(x=x_data, y=y_data, scales={'x': sc_x, 'y': sc_y}, colors=['blue'])
ax_x = Axis(label='Test X', scale=sc_x)
ax_y = Axis(label='Test Y', scale=sc_y, orientation='vertical', tick_format='0.2f')
Figure(axes=[ax_x, ax_y], marks=[scatter])
# + [markdown] deletable=true editable=true
# ## Histogram
# + deletable=true editable=true
scale_x = LinearScale()
scale_y = LinearScale()
hist = Hist(sample=y_data, scales={'sample': scale_x, 'count': scale_y})
ax_x = Axis(label='X', scale=scale_x, tick_format='0.2f')
ax_y = Axis(label='Y', scale=scale_y, orientation='vertical', grid_lines='solid')
Figure(axes=[ax_x, ax_y], marks=[hist])
# + [markdown] deletable=true editable=true
# ## Bar Chart
# + deletable=true editable=true
sc_x1 = OrdinalScale()
sc_y1 = LinearScale()
bar_x = Axis(label='X', scale=sc_x1)
bar_y = Axis(label='Y', scale=sc_y1, orientation='vertical', tick_format='0.0f', grid_lines='solid')
bar_chart = Bars(x=['A', 'B', 'C', 'D', 'E', 'F', 'G', 'H', 'I', 'J', 'K', 'L', 'M', 'N', 'P', 'Q', 'R', 'S', 'T', 'U'],
y=np.abs(y_data[:20]), scales={'x': sc_x1, 'y': sc_y1})
Figure(axes=[bar_x, bar_y], marks=[bar_chart], padding_x=0)
| examples/Basic Plotting/Basic Plotting.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# ## Analyze A/B Test Results
#
# This project will assure you have mastered the subjects covered in the statistics lessons. The hope is to have this project be as comprehensive of these topics as possible. Good luck!
#
# ## Table of Contents
# - [Introduction](#intro)
# - [Part I - Probability](#probability)
# - [Part II - A/B Test](#ab_test)
# - [Part III - Regression](#regression)
#
#
# <a id='intro'></a>
# ### Introduction
#
# A/B tests are very commonly performed by data analysts and data scientists. It is important that you get some practice working with the difficulties of these
#
# For this project, you will be working to understand the results of an A/B test run by an e-commerce website. Your goal is to work through this notebook to help the company understand if they should implement the new page, keep the old page, or perhaps run the experiment longer to make their decision.
#
# **As you work through this notebook, follow along in the classroom and answer the corresponding quiz questions associated with each question.** The labels for each classroom concept are provided for each question. This will assure you are on the right track as you work through the project, and you can feel more confident in your final submission meeting the criteria. As a final check, assure you meet all the criteria on the [RUBRIC](https://review.udacity.com/#!/projects/37e27304-ad47-4eb0-a1ab-8c12f60e43d0/rubric).
#
# <a id='probability'></a>
# #### Part I - Probability
#
# To get started, let's import our libraries.
import pandas as pd
import numpy as np
import random
import statsmodels.api as sm
import statsmodels.formula.api as smf
from statsmodels.stats.outliers_influence import variance_inflation_factor
from scipy.stats import norm
from patsy import dmatrices
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# We are setting the seed to assure you get the same answers on quizzes as we set up
random.seed(42)
# `1.` Now, read in the `ab_data.csv` data. Store it in `df`. **Use your dataframe to answer the questions in Quiz 1 of the classroom.**
#
# a. Read in the dataset and take a look at the top few rows here:
# read in the dataset
df=pd.read_csv('ab_data.csv')
df.head()
# b. Use the below cell to find the number of rows in the dataset.
df.shape[0]
# c. The number of unique users in the dataset.
df.user_id.nunique()
# d. The proportion of users converted.
proportion_converted=(df['converted']==1).mean()*100
print("The proportion of users converted is {} %.".format(proportion_converted))
# e. The number of times the `new_page` and `treatment` don't line up.
# mismatch where group is treatment and landing page is not new page
not_equal1=(df[(df['group']=='treatment') & (df['landing_page']!='new_page')]).shape[0]
# mismatch where landing page is new page and group is not treatment
not_equal2=(df[(df['landing_page']=='new_page') & (df['group']!='treatment')]).shape[0]
total_unequal= not_equal1+not_equal2
print("No. of times new_page and treatment don't line up is {}.".format(total_unequal))
# f. Do any of the rows have missing values?
# do all columns have same number of data
df.info()
# find the rows with any missing values and add them
(df.isnull().sum(axis=1)).sum()
# **There is no row with missing values.**
# `2.` For the rows where **treatment** is not aligned with **new_page** or **control** is not aligned with **old_page**, we cannot be sure if this row truly received the new or old page. Use **Quiz 2** in the classroom to provide how we should handle these rows.
#
# a. Now use the answer to the quiz to create a new dataset that meets the specifications from the quiz. Store your new dataframe in **df2**.
# drop rows where treatment and new_page don't line up
df2=df.drop(df[((df['group'] == 'treatment') == (df['landing_page'] == 'new_page')) == False].index)
# Double Check all of the correct rows were removed - this should be 0
df2[((df2['group'] == 'treatment') == (df2['landing_page'] == 'new_page')) == False].shape[0]
# `3.` Use **df2** and the cells below to answer questions for **Quiz3** in the classroom.
# a. How many unique **user_id**s are in **df2**?
df2.user_id.nunique()
# b. There is one **user_id** repeated in **df2**. What is it?
# the ids that are duplicate, includes the first one as well
df2[df2.user_id.duplicated(keep=False) == True]
# c. What is the row information for the repeat **user_id**?
# **user_id 773192 has duplicate information.**
# d. Remove **one** of the rows with a duplicate **user_id**, but keep your dataframe as **df2**.
# find the index of the duplicated id and drop it
df2=df2.drop(df2[df2.user_id.duplicated() == True].index)
df2.shape
# `4.` Use **df2** in the below cells to answer the quiz questions related to **Quiz 4** in the classroom.
#
# a. What is the probability of an individual converting regardless of the page they receive?
# probability of an individual having 1 in converted column
converted_prob=(df2['converted']==1).mean()
print("Probability of an individual converting regardless of the page they receive is {}.".format(converted_prob))
# b. Given that an individual was in the `control` group, what is the probability they converted?
# probability of conversion being in the control group
control_prob=df2.query('group=="control"').converted.mean()
print("Probability of an individual converting from control group is {}.".format(control_prob))
# c. Given that an individual was in the `treatment` group, what is the probability they converted?
# probability of conversion being in the treatment group
treatment_prob=df2.query('group=="treatment"').converted.mean()
print("Probability of an individual converting from treatment group is {}.".format(treatment_prob))
# difference in the probability of an individual converting, belonging to the two groups.
diff=treatment_prob-control_prob
diff
# d. What is the probability that an individual received the new page?
# the probability that an individual received the new page
received_new=df2[df2['landing_page']=="new_page"].landing_page.count()/df2.shape[0]
print("The probability that an individual received the new page is {}.".format(received_new))
# e. Consider your results from a. through d. above, and explain below whether you think there is sufficient evidence to say that the new treatment page leads to more conversions.
# **I don't think there is enough evidence to suggest that the new treatment leads to more conversions. We know that there is a 50-50 chance that an individual will receive the new page. The control group has slightly more conversion than the treatment group. We need to test if the difference is significant or not.**
# <a id='ab_test'></a>
# ### Part II - A/B Test
#
# Notice that because of the time stamp associated with each event, you could technically run a hypothesis test continuously as each observation was observed.
#
# However, then the hard question is do you stop as soon as one page is considered significantly better than another or does it need to happen consistently for a certain amount of time? How long do you run to render a decision that neither page is better than another?
#
# These questions are the difficult parts associated with A/B tests in general.
#
#
# `1.` For now, consider you need to make the decision just based on all the data provided. If you want to assume that the old page is better unless the new page proves to be definitely better at a Type I error rate of 5%, what should your null and alternative hypotheses be? You can state your hypothesis in terms of words or in terms of **$p_{old}$** and **$p_{new}$**, which are the converted rates for the old and new pages.
# $$H_0: p_{new}<=p_{old}$$
# $$H_1: p_{new}>p_{old}$$
# `2.` Assume under the null hypothesis, $p_{new}$ and $p_{old}$ both have "true" success rates equal to the **converted** success rate regardless of page - that is $p_{new}$ and $p_{old}$ are equal. Furthermore, assume they are equal to the **converted** rate in **ab_data.csv** regardless of the page. <br><br>
#
# Use a sample size for each page equal to the ones in **ab_data.csv**. <br><br>
#
# Perform the sampling distribution for the difference in **converted** between the two pages over 10,000 iterations of calculating an estimate from the null. <br><br>
#
# Use the cells below to provide the necessary parts of this simulation. If this doesn't make complete sense right now, don't worry - you are going to work through the problems below to complete this problem. You can use **Quiz 5** in the classroom to make sure you are on the right track.<br><br>
# a. What is the **convert rate** for $p_{new}$ under the null?
pnew=converted_prob
pnew
# b. What is the **convert rate** for $p_{old}$ under the null? <br><br>
pold=converted_prob
pold
# c. What is $n_{new}$?
# no of samples shown new_page
nnew=df2[df2['landing_page']=="new_page"]['user_id'].count()
# d. What is $n_{old}$?
# no of samples shown old_page
nold=df2[df2['landing_page']=="old_page"]['user_id'].count()
# e. Simulate $n_{new}$ transactions with a convert rate of $p_{new}$ under the null. Store these $n_{new}$ 1's and 0's in **new_page_converted**.
new_page_converted=np.random.binomial(1, pnew, nnew)
new_page_converted
# f. Simulate $n_{old}$ transactions with a convert rate of $p_{old}$ under the null. Store these $n_{old}$ 1's and 0's in **old_page_converted**.
old_page_converted=np.random.binomial(1, pold, nold)
old_page_converted
# g. Find $p_{new}$ - $p_{old}$ for your simulated values from part (e) and (f).
diff_page=new_page_converted.mean()-old_page_converted.mean()
diff_page
# h. Simulate 10,000 $p_{new}$ - $p_{old}$ values using this same process similarly to the one you calculated in parts **a. through g.** above. Store all 10,000 values in a numpy array called **p_diffs**.
# sampling distribution of the difference of the means of conversions for old page and new page
new_converted_simulation = np.random.binomial(nnew, pnew, 10000)/nnew
old_converted_simulation = np.random.binomial(nold, pold, 10000)/nold
p_diffs = new_converted_simulation - old_converted_simulation
# i. Plot a histogram of the **p_diffs**. Does this plot look like what you expected? Use the matching problem in the classroom to assure you fully understand what was computed here.
# Convert to numpy array
p_diffs = np.array(p_diffs)
# Plot sampling distribution with null mean of 0 as the center
plt.hist(p_diffs)
plt.axvline(0, color='red')
plt.axvline(diff, color='red');
# j. What proportion of the **p_diffs** are greater than the actual difference observed in **ab_data.csv**?
# proportion of p_diffs greater than the actual difference observed in part 1
(p_diffs>diff).mean()
# k. In words, explain what you just computed in part **j.** What is this value called in scientific studies? What does this value mean in terms of whether or not there is a difference between the new and old pages?
# **In part j we computed the p-value based on the null hypothesis (question 2) that pnew-pold=0.This was calculated from the sampling distribution of the difference of means which in this case is same as the sampling distribution under the null hypothesis as it is centered at 0 (which is the null mean, since the null hypothesis is pnew-pold=0 and the alternative hypothesis is pnew-pold>0) and has the same spread.**
#
# **The p-value is 0.903 which is greater than alpha=0.05 thus we fail to reject the null hypothesis. So we see that we fail to reject the null hupothesis. So, there is no difference between the new and old page and hence no reason to move away from the old page.**
# l. We could also use a built-in to achieve similar results. Though using the built-in might be easier to code, the above portions are a walkthrough of the ideas that are critical to correctly thinking about statistical significance. Fill in the below to calculate the number of conversions for each page, as well as the number of individuals who received each page. Let `n_old` and `n_new` refer the the number of rows associated with the old page and new pages, respectively.
# the number of conversions for each pag
convert_old = df2[df2['landing_page']=='old_page'].converted.sum()
convert_new = df2[df2['landing_page']=='new_page'].converted.sum()
#the number of individuals who received each page
n_old = nold
n_new = nnew
# m. Now use `stats.proportions_ztest` to compute your test statistic and p-value. [Here](http://knowledgetack.com/python/statsmodels/proportions_ztest/) is a helpful link on using the built in.
# compute z-score and p-value for one tailed test using stats.proportions_ztest
z_score, p_value = sm.stats.proportions_ztest([convert_new, convert_old], [n_new, n_old], alternative='larger')
z_score, p_value
# +
# how significant our z-score is
print("Significance of z-score is {}.".format(norm.cdf(z_score)))
# our critical value at 95% confidence is
print("The critical value is {}".format(norm.ppf(1-(0.05/2))))
# -
# n. What do the z-score and p-value you computed in the previous question mean for the conversion rates of the old and new pages? Do they agree with the findings in parts **j.** and **k.**?
# **Since the z-score of -1.31 is less than the critical value of 1.959963984540054, we fail to reject the null hypothesis that there is no difference between the two proportions and they are equal to the converted rate in ab_data.csv regardless of the page.**
#
# **This is perfectly in line with our previous findings in parts j and k.**
# <a id='regression'></a>
# ### Part III - A regression approach
#
# `1.` In this final part, you will see that the result you acheived in the previous A/B test can also be acheived by performing regression.<br><br>
#
# a. Since each row is either a conversion or no conversion, what type of regression should you be performing in this case?
# **Since we have a nominal(true or false for conversion) variable as the dependent variable, we should use Logistic Regression.**
# b. The goal is to use **statsmodels** to fit the regression model you specified in part **a.** to see if there is a significant difference in conversion based on which page a customer receives. However, you first need to create a column for the intercept, and create a dummy variable column for which page each user received. Add an **intercept** column, as well as an **ab_page** column, which is 1 when an individual receives the **treatment** and 0 if **control**.
# create dummy variables for the group variable
df2[['page','ab_page']]=pd.get_dummies(df2['group'])
# dropping the page dummy variables as ab_page gives the same information
df2=df2.drop('page', axis=1)
# add intercept to the data
df2['intercept']=1
# c. Use **statsmodels** to import your regression model. Instantiate the model, and fit the model using the two columns you created in part **b.** to predict whether or not an individual converts.
# instantiate the logistic regression model
logit_model_df2=sm.Logit(df2['converted'], df2[['intercept', 'ab_page']])
#fit the model
results_df2=logit_model_df2.fit()
# d. Provide the summary of your model below, and use it as necessary to answer the following questions.
# get the summary
results_df2.summary()
# The estimated coefficients are the log odds. By exponentiating these values,we can calculate
# the odds. And since the coefficient is negative, we invert the odds ratio to find the relation
# between conversion rate and the ab_page
1/np.exp(-0.0150)
# e. What is the p-value associated with **ab_page**? Why does it differ from the value you found in **Part II**?<br><br> **Hint**: What are the null and alternative hypotheses associated with your regression model, and how do they compare to the null and alternative hypotheses in the **Part II**?
# **The P value associated with the ab_page is 0.190, which is higher than the alpha value of 0.05. So we fail to reject the null hypothesis.**
#
# **For the logistic regression the null hypothesis is there is no relationship between the landing page and conversion rate. So there should be no difference in conversion rate based on the page or $$pnew-pold=0$$ and the alternate hypothesis is there is relation between the two so $$pnew-pold\neq0$$ . Here it is a two tail test and hence we see a difference in the p value obtained from part 2 where we had the alternate hypothesis pnew-pold >0.**
#
# **Also from the coefficient of ab_page we can say that the conversion rate for the old page is 1.015 times of the new page. So they are almost same which aligns with our proposition of failing to reject the null hypothesis.**
# f. Now, you are considering other things that might influence whether or not an individual converts. Discuss why it is a good idea to consider other factors to add into your regression model. Are there any disadvantages to adding additional terms into your regression model?
# **Adding more than one explanatory variable to the regression model help us determine the relative influence that more than one predictor have on the response. This multiple logistic regression might help to get insights that is not possible just from single logistic regression. Such as we can see if certain landing pages lead to more conversion for certain countries or certain times of the day or week that would not be possible with just the landing page variable. It also helps identify outliers in the sample data.**
#
# **Adding additional terms to the model has the disadvantage that instead of increasing the quality of the model it could decrease it.Incomplete data can leads to concluding of an incorrect relationship between the variables.Also it could lead to falsely concluding that correlation is a causation.**
#
# **https://sciencing.com/advantages-disadvantages-multiple-regression-model-12070171.html**
# g. Now along with testing if the conversion rate changes for different pages, also add an effect based on which country a user lives. You will need to read in the **countries.csv** dataset and merge together your datasets on the approporiate rows. [Here](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.join.html) are the docs for joining tables.
#
# Does it appear that country had an impact on conversion? Don't forget to create dummy variables for these country columns - **Hint: You will need two columns for the three dummy variables.** Provide the statistical output as well as a written response to answer this question.
# read from the dataset and merge the two datasets based on the user_id column
countries_df = pd.read_csv('./countries.csv')
df_new = countries_df.set_index('user_id').join(df2.set_index('user_id'), how='inner')
df_new.head(5)
# Create the necessary dummy variables
df_new[['ca', 'uk', 'us']]=pd.get_dummies(df_new['country'])
# logistic regression with the ab_page and countries (ca as the baseline)
logit_model_df_page_country=sm.Logit(df_new['converted'], df_new[['intercept', 'ab_page', 'us', 'uk']])
results_df_page_country=logit_model_df_page_country.fit()
results_df_page_country.summary()
# odds ratio
np.exp(results_df_page_country.params)
# **We find conversion rate is not effected by countries or different landing page.
# From the coefficient of US we can say that the conversion rate for US is 1.04 times of CA holding all other variables constant.
# From the coefficient of UK we can say that the conversion rate of UK is 1.05 times of CA holding all other variables constant.
# None of the predictors are statistically significant**
# h. Though you have now looked at the individual factors of country and page on conversion, we would now like to look at an interaction between page and country to see if there significant effects on conversion. Create the necessary additional columns, and fit the new model.
#
# Provide the summary results, and your conclusions based on the results.
# +
# Fit Your Linear Model And Obtain the Results
# Find the interaction terms
df_new['page_us'] = df_new.ab_page * df_new.us
df_new['page_uk'] = df_new.ab_page * df_new.uk
# Relation between converted and the pages, countries and pages*countries
model_multiply = smf.ols(formula='converted ~ ab_page + us + page_us+ uk+ page_uk', data=df_new).fit()
summary_multiply = model_multiply.summary()
summary_multiply
# -
# **From the p values we can see that none of the tested variables are less than 0.05. So we fail to reject the null hypothesis that there is no relation between the tested X variables, country and page and y variable, conversion. The interactions also had P values more than 0.05 so no particular landing page had any effect on the country. Also we have a R-squared value of 0, suggesting no relationship at all**
logit_model_multiply = smf.logit(formula='converted ~ ab_page + us + page_us+ uk+ page_uk', data=df_new).fit()
summary_logit = logit_model_multiply.summary()
summary_logit
# **For logistic regression we also see that the p values are greater than 0.05. So the tested X variables (landing_page and country) or their interation terms didn't have any effect on the model. So we fail to reject the null hypothesis that there is no relationship between the response and predictor variables.**
# ### Additional Analysis:
#
# **Effect of day of the week and time of the day on regression.**
#
# We will create the necessary dummy varaiables.
# convert timestamp column to series and convert the series to day of week(Moday=0,...)
df_new['weekday'] = pd.to_datetime(df_new['timestamp']).dt.dayofweek
# convert weekday to 1 and weekend to 0
df_new['weekday']=(df_new['weekday'] < 5).astype(int)
# convert the hours and minutes to decimals for categorising the day based on time in decimals
df_new['time'] = pd.to_datetime(df_new['timestamp']).dt.hour+pd.to_datetime(df_new['timestamp']).dt.minute/60
# Categorise the day based on the decimal time
df_new['time_aspect']=df_new.apply(lambda x: 'morning' if x.time>=7 and x.time<=12 else('afternoon' if x.time>12 and x.time<=18 else 'evening'), axis=1)
# Create the necessary dummy variables for time_aspect
df_new[['afternoon','evening', 'morning']]=pd.get_dummies(df_new['time_aspect'])
# Since there are so many X variables now, we will check for multicollinearity
sns.pairplot(df_new[['ab_page', 'us', 'uk', 'weekday', 'afternoon', 'evening']]);
# We don't see any linear correlation between the X variables. We can also check the variance inflation factor.
# Define y and X matrix
y,X=dmatrices('converted ~ ab_page+us+uk+weekday+afternoon+evening', data=df_new, return_type='dataframe')
# For each X, calculate VIF and save in dataframe
vif = pd.DataFrame()
vif["VIF Factor"] = [variance_inflation_factor(X.values, i) for i in range(X.shape[1])]
vif["features"] = X.columns
vif
# **Since the vif factors are less than 10 the predictor variables are not correlated with one another (No multicollinearity).**
# We will create the interaction terms between page and weekday, page and afternoon, evening only as we are aiming to find out which page leads to better conversion and if other factors along with the page version can influence the conversion.
df_new.head()
# +
# Create the interaction terms
df_new['page_weekday'] = df_new.ab_page * df_new.weekday
df_new['page_afternoon'] = df_new.us * df_new.afternoon
df_new['page_evening'] = df_new.us * df_new.evening
# Create the model and fit it.
multiple_mode= smf.logit(formula='converted ~ ab_page + us + page_us+ uk+ page_uk+weekday+page_weekday+afternoon+page_afternoon+evening+page_evening', data=df_new).fit()
multiple_mode.summary()
# -
# odds ratios and 95% Confidence interval
params = multiple_mode.params
conf = multiple_mode.conf_int()
conf['OR'] = params
conf.columns = ['2.5%', '97.5%', 'OR']
np.exp(conf)
# **From our results we see that our p values are greater than 0.05. So we fail to reject the null hypothesis that there is no relationship between the version of page or country or time (in terms of weekday or weekend, or time of the day) or their interaction terms with the conversion rate.**
#
# **From the coefficients also we see that the effect of these predictor variables on the conversion is somewhat similar(So from 0.91-1.05).Which means that each of the page version, country where the consumers live, the day time, the interaction terms has no effect on the conversion rate while other variables remain constant.**
#
# **We have a negative number for log-likelihood. A log-likelihood of 0 indicates a good overall fit of the model while a negative infinity means a poor fit. So from our results we see that the model didn't fit the data well. Although, in this last logistic regression model the pseudo R squared value is greater than all the other logistic regression models we had. So adding multiple variables has actually improved the quality of the model.**
# <a id='conclusions'></a>
# ## Conclusions
#
# Congratulations on completing the project!
#
# ### Gather Submission Materials
#
# Once you are satisfied with the status of your Notebook, you should save it in a format that will make it easy for others to read. You can use the __File -> Download as -> HTML (.html)__ menu to save your notebook as an .html file. If you are working locally and get an error about "No module name", then open a terminal and try installing the missing module using `pip install <module_name>` (don't include the "<" or ">" or any words following a period in the module name).
#
# You will submit both your original Notebook and an HTML or PDF copy of the Notebook for review. There is no need for you to include any data files with your submission. If you made reference to other websites, books, and other resources to help you in solving tasks in the project, make sure that you document them. It is recommended that you either add a "Resources" section in a Markdown cell at the end of the Notebook report, or you can include a `readme.txt` file documenting your sources.
#
# ### Submit the Project
#
# When you're ready, click on the "Submit Project" button to go to the project submission page. You can submit your files as a .zip archive or you can link to a GitHub repository containing your project files. If you go with GitHub, note that your submission will be a snapshot of the linked repository at time of submission. It is recommended that you keep each project in a separate repository to avoid any potential confusion: if a reviewer gets multiple folders representing multiple projects, there might be confusion regarding what project is to be evaluated.
#
# It can take us up to a week to grade the project, but in most cases it is much faster. You will get an email once your submission has been reviewed. If you are having any problems submitting your project or wish to check on the status of your submission, please email us at <EMAIL>. In the meantime, you should feel free to continue on with your learning journey by beginning the next module in the program.
from subprocess import call
call(['python', '-m', 'nbconvert', 'Analyze_ab_test_results_notebook.ipynb'])
# ### Resources:
# https://datascience.stackexchange.com/questions/12645/how-to-count-the-number-of-missing-values-in-each-row-in-pandas-dataframe
# https://stackoverflow.com/questions/14657241/how-do-i-get-a-list-of-all-the-duplicate-items-using-pandas-in-python
# http://joelcarlson.github.io/2016/05/10/Exploring-Interactions/
# http://gael-varoquaux.info/stats_in_python_tutorial/#testing-for-interactions
# http://songhuiming.github.io/pages/2016/07/12/statsmodels-regression-examples/
# https://www.statsmodels.org/dev/example_formulas.html
# http://blog.yhat.com/posts/logistic-regression-and-python.html
# https://stackoverflow.com/questions/32278728/convert-dataframe-date-row-to-a-weekend-not-weekend-value
# https://stackoverflow.com/questions/35595710/splitting-timestamp-column-into-seperate-date-and-time-columns
# https://stackoverflow.com/questions/36083857/pandas-way-convert-time-of-the-day-valid-datetime-time-to-float-variables
# https://stackoverflow.com/questions/44991438/lambda-including-if-elif-else
# https://www.statisticssolutions.com/assumptions-of-logistic-regression/
# https://stackoverflow.com/questions/50591982/importerror-cannot-import-name-timestamp
# http://knowledgetack.com/python/statsmodels/proportions_ztest/
# https://softwareengineering.stackexchange.com/questions/254475/how-do-i-move-away-from-the-for-loop-school-of-thought
# https://www.juanshishido.com/logisticcoefficients.html
# https://stats.idre.ucla.edu/other/mult-pkg/faq/pvalue-htm/
| Analyze_ab_test_results_notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
sns.set(rc={"figure.figsize": (6, 6)})
# color_palette() without arg = default color
# set_palette() set all panel color
# -
current_palette = sns.color_palette()
sns.palplot(current_palette)
# deep, muted, pastel, bright, dark, colorblind
sns.palplot(sns.color_palette("hls", 8))
# example:
data = np.random.normal(size=(20, 8)) + np.arange(8) / 2
sns.boxplot(data=data,palette=sns.color_palette("hls", 8))
# l- lightness
# s-saturation
sns.palplot(sns.hls_palette(8, l=.7, s=.9))
sns.palplot(sns.color_palette("Paired",8))
plt.plot([0, 1], [0, 1], sns.xkcd_rgb["pale red"], lw=3)
plt.plot([0, 1], [0, 2], sns.xkcd_rgb["medium green"], lw=3)
plt.plot([0, 1], [0, 3], sns.xkcd_rgb["denim blue"], lw=3)
colors = ["windows blue", "amber", "greyish", "faded green", "dusty purple"]
sns.palplot(sns.xkcd_palette(colors))
# Contiunes colors
sns.palplot(sns.color_palette("BuGn_r"))
# cubehelix_palette() linear change color
sns.palplot(sns.color_palette("cubehelix", 8))
sns.palplot(sns.cubehelix_palette(8, start=.5, rot=-.75))
sns.palplot(sns.cubehelix_palette(8, start=.75, rot=-.150))
sns.palplot(sns.light_palette("green"))
sns.palplot(sns.light_palette("navy", reverse=True))
x, y = np.random.multivariate_normal([0, 0], [[1, -.5], [-.5, 1]], size=300).T
pal = sns.dark_palette("green", as_cmap=True)
sns.kdeplot(x, y, cmap=pal);
sns.palplot(sns.light_palette((210, 90, 60), input="husl"))
| Yudi TANG/Packages/Seaborn/Seaborn color .ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import math
import os
def printCategory(category_dict):
for category in category_dict:
print(category)
if category_dict[category] is not None:
sub_dic = category_dict[category]
printCategory(sub_dic)
def createLevelColumns(df, levels, new_columns):
for i in range(len(df)):
df_level1, df_level2, df_level3 = "NULL","NULL","NULL"
df_level1, nextLevels = findNotNullLevel(df, i, levels)
if nextLevels is not None:
df_level2, nextLevels = findNotNullLevel(df, i, nextLevels)
if nextLevels is not None:
df_level3, nextLevel = findNotNullLevel(df, i, nextLevels)
most_specific_category = "NULL"
if df_level3 != "NULL":
df_level3 = df_level3 + "__" + df_level2 + "__" + df_level1
if df_level2 != "NULL":
df_level2 = df_level2 + "__" + df_level1
if df_level3 != "NULL":
most_specific_category = df_level3
elif df_level2 != "NULL":
most_specific_category = df_level2
elif df_level1 != "NULL":
most_specific_category = df_level1
df.iloc[i, df.columns.get_loc('level1')] = df_level1
df.iloc[i, df.columns.get_loc('level2')] = df_level2
df.iloc[i, df.columns.get_loc('level3')] = df_level3
df.iloc[i, df.columns.get_loc('mostSpecificCategory')] = most_specific_category
def findNotNullLevel(df, i, levels):
for level in levels:
if df.iloc[i][level] == "NULL":
continue
else:
return df.iloc[i][level], levels[level]
return "NULL", None
def unionTwoLists(list1, list2):
for category in list1:
if category not in list2:
list2.append(category)
return list2
def checkNULL(checked_list):
for item in checked_list:
if item == "NULL":
print("Contains NULL")
return
print("Not contains NULL")
# -
# #### 导入数据 yearly and range
# #### process为designed schema
# 1. GazetteerEconomy table
#
# 1.1 Index - Id
#
# 1.2 Gazetteer Code - gazetteerId
#
# 1.3 Category - categoryId
#
# 1.4 yearly - startYear = endYear
#
# 1.5 2010 data - data
#
# 1.6 Unit - unitId
#
# 2. EconomyCategory table
# 2.1 Category Division1 Subdivision - Agri. Subdivision - Misc. Subdivision - Service Division2 Subdivision - Househould
#
# 3. UnitCategory
#
# 4. Gazetteer
path = os.path.abspath(os.getcwd())
df = pd.read_csv(path + "/Data2/Economy - Yearly.csv")
df2 = pd.read_csv(path + "/Data2/Economy - Range.csv")
# +
# create new columns at df and df2
new_columns = ['level1', 'level2', 'level3', 'mostSpecificCategory', 'categoryId']
for column in new_columns:
df[column] = None
df = df.where(df.notnull(), "NULL")
for column in new_columns:
df2[column] = None
df2 = df2.where(df2.notnull(), "NULL")
# create dictionary records category levles
heading = {'Category':
{'Division1':{'Subdivision - Agri.': None, 'Subdivision - Misc.': None, 'Subdivision - Service': None},
'Division2':{'Subdivision - Household': None}
}
}
printCategory(heading)
createLevelColumns(df, heading, new_columns)
createLevelColumns(df2, heading, new_columns)
level1 = unionTwoLists([cat for cat in df['level1'].astype('category').unique()],
[cat for cat in df2['level1'].astype('category').unique()] )
level2 = unionTwoLists([cat for cat in df['level2'].astype('category').unique()],
[cat for cat in df2['level2'].astype('category').unique()] )
level3 = unionTwoLists([cat for cat in df['level3'].astype('category').unique()],
[cat for cat in df2['level3'].astype('category').unique()] )
most_specific_category = unionTwoLists([cat for cat in df['mostSpecificCategory'].astype('category').unique()],
[cat for cat in df2['mostSpecificCategory'].astype('category').unique()])
total_categories = unionTwoLists(level1, level2)
total_categories = unionTwoLists(level3, total_categories)
# get total categories
total_categories.sort()
checkNULL(total_categories)
print("total number of categories are " + str(len(total_categories)))
# temp = [ item.split('__', 1) for item in total_categories]
# get most specific category
most_specific_category.sort()
checkNULL(most_specific_category)
print("total number of recorded categories are " + str(len(most_specific_category)))
# create dict "dic_category_id" store { category_name : id}
dic_category_id = {}
count = 1
for category in total_categories:
if category != "NULL" and category not in dic_category_id:
dic_category_id[category] = count
count = count + 1
# create dict "dic_for_unitId" stores {unit: id}
dic_for_unitId = {}
count = 1
units_from_df = unionTwoLists([cat for cat in df['Unit'].astype('category').unique()], [cat for cat in df2['Unit'].astype('category').unique()])
for unit in units_from_df:
if unit not in dic_for_unitId:
dic_for_unitId[unit] = count
count = count + 1
# +
# creat categoryId column at dataframe
df_categoryId = []
for i in range(len(df)):
category = df.iloc[i]['mostSpecificCategory']
if category in dic_category_id:
df_categoryId.append(dic_category_id[category])
else:
print("Not recorded category for entity " + i)
break;
df['categoryId'] = df_categoryId
df_categoryId = []
for i in range(len(df2)):
category = df2.iloc[i]['mostSpecificCategory']
if category in dic_category_id:
df_categoryId.append(dic_category_id[category])
else:
print("Not recorded category for entity " + i)
break;
df2['categoryId'] = df_categoryId
# -
df2.columns
# +
# create economy_df
economy_df = pd.DataFrame(columns = ['gazetteerId', 'categoryId', 'startYear', 'endYear', 'data', 'unitId'])
years = [str(i) for i in range(1949,2020)]
dic_for_economy_df = {'gazetteerId':[], 'categoryId':[], 'startYear':[], 'endYear':[], 'data':[], 'unitId':[]}
# Process yearly data
for i in range(len(df)):# each row
for year in years: # 1949 - 2019
if df.iloc[i][year] != "NULL":
dic_for_economy_df['gazetteerId'].append(df.iloc[i]['村志代码 Gazetteer Code'])
dic_for_economy_df['categoryId'].append(df.iloc[i]['categoryId'])
dic_for_economy_df['startYear'].append(int(year))
dic_for_economy_df['endYear'].append(int(year))
dic_for_economy_df['data'].append(df.iloc[i][year])
dic_for_economy_df['unitId'].append(dic_for_unitId[df.iloc[i]['Unit']])
# Process range data
for i in range(len(df2)):
dic_for_economy_df['gazetteerId'].append(df2.iloc[i]['村志代码 Gazetteer Code'])
dic_for_economy_df['categoryId'].append(df2.iloc[i]['categoryId'])
dic_for_economy_df['startYear'].append(df2.iloc[i]['Start Year'])
dic_for_economy_df['endYear'].append(df2.iloc[i]['End Year'])
dic_for_economy_df['data'].append(df2.iloc[i]['Data'])
dic_for_economy_df['unitId'].append(dic_for_unitId[df.iloc[i]['Unit']])
for attribute in economy_df.columns:
economy_df[attribute] = dic_for_economy_df[attribute]
economy_df.head()
# +
# create economyCategory_df
economyCategory_df = pd.DataFrame(columns = ['id', 'name', 'parentId'])
dic_for_ecoCategorydf = {'id':[], 'name':[], 'parentId':[]}
for category in dic_category_id:
child_parent = category.split('__', 1)
name = child_parent[0]
if len(child_parent) == 1:
dic_for_ecoCategorydf['id'].append(dic_category_id[category])
dic_for_ecoCategorydf['name'].append(name)
dic_for_ecoCategorydf['parentId'].append("NULL")
else:
parentId = dic_category_id[child_parent[1]]
dic_for_ecoCategorydf['id'].append(dic_category_id[category])
dic_for_ecoCategorydf['name'].append(name)
dic_for_ecoCategorydf['parentId'].append(parentId)
for attribute in economyCategory_df.columns:
economyCategory_df[attribute] = dic_for_ecoCategorydf[attribute]
len(economyCategory_df)
# +
# creat economyUnitCategory_df
economyUnitCategory_df = pd.DataFrame(columns = ['id', 'name'])
dic_for_economyUnitCategory_df = {'id':[], 'name':[]}
for unit_name in dic_for_unitId:
dic_for_economyUnitCategory_df['id'].append(dic_for_unitId[unit_name])
dic_for_economyUnitCategory_df['name'].append(unit_name)
for attribute in economyUnitCategory_df.columns:
economyUnitCategory_df[attribute] = dic_for_economyUnitCategory_df[attribute]
len(economyUnitCategory_df)
# -
count = 0
for i in range(len(economy_df)):
if economy_df.iloc[i]['data'] == 0:
count = count + 1
count
# +
# economy_df.to_csv('economy_df.csv', index = False, na_rep = "NULL")
# -
len(economy_df)
# +
# economyCategory_df.to_csv('economyCategory.csv', index = False, na_rep = "NULL")
# +
# economyUnitCategory_df.to_csv('economyUnitCategory_df.csv', index = False, na_rep = "NULL")
# -
print(economyCategory_df.head(2))
print(economyUnitCategory_df.head(2))
print(economy_df.head(10))
len(economy_df)
| process_data/process_data_economy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# +
import warnings
warnings.filterwarnings('ignore')
# %load_ext rpy2.ipython
# %run ../notebook-init.py
# load R libraries
# %R invisible(library(ggplot2))
# %R invisible(library(fastcluster))
# %R invisible(library(reshape))
# %R invisible(library(reshape2))
# %R invisible(library(gplots))
# %R invisible(library(RSQLite))
#set up ggplot2 defaults
# %R theme_set(theme_gray(base_size=18))
# %pylab inline
pylab.rcParams['figure.figsize'] = (20, 20)
# + language="R"
#
# pGeneIsAR <- 1 / (10)
# nGenes <- 100
#
# step <- 1
# points <- 1:100
#
# plot(dbinom(step*points,size=nGenes,prob=pGeneIsAR))
# nGenes
#
#
# + language="R"
#
# pGeneIsAR <- 1 / (100*1000*1000)
# nGenes <- 100*(1000*1000*1000*1000)*1000
#
# step <- 100*1000
# start <- 900*1000*1000
# end <- 1100*1000*1000
# X <- seq(start,end,by=step)
# print(range(X))
# plot(pbinom(X,size=nGenes,prob=pGeneIsAR))
# nGenes
#
# +
import numpy as np
import pandas as pd
from glob import glob
sbres = glob('../shortbred/FMTchrons2/*results.txt')
tbls = {}
for fname in sbres:
tbl = pd.read_table(fname,header=0,index_col=0)
tblname = fname.split('/')[-1]
tblname = tblname.split('.')[0]
rep = tblname.split('-')[2]
time = tblname.split('-')[1]
if time.lower() == 'pre_fmt':
time = 0
elif '4' in time:
time = 4
elif '8' in time:
time = 8
if rep not in tbls:
tbls[rep] = {}
tbls[rep][time] = tbl[['Hits']]
subtbls={}
for rep, tblset in tbls.items():
subtbls[rep] = pd.concat(tblset,axis=1)
pnl = pd.Panel(subtbls)
# +
s = []
for i in pnl.major_axis:
a = pnl.major_xs(i).values.flatten()
assert len(a) == 18*3
s.append(a)
pnl.major_xs(pnl.major_axis[0])
# + magic_args="-i s" language="R"
#
#
#
# print( apply(s,2,function(r){
# return( sum(r[!is.na(r)] > 0))
# }))
# + magic_args="-i s" language="R"
#
#
#
# # print( apply(s,1,function(r){
# # return( sum(r > 0))
# # }))
#
#
# k <- s[,104]
# k <- k[!is.na(k)]
# print(length(k))
# print(sum(k > 0))
# print(sum(k == 0))
# print(head(k))
# # print(max(k))
# #k <- k[k<100]
# # print(length(hinfo$breaks))
# # print(table(k))
# t <- table(k)
# # print(as.numeric(names(t)))
# df <- data.frame(t)
#
#
# N <- 40*1000*1000
# df <- df
# df$p <- as.numeric(df$k) / N
# df$bin <- sum(df$Freq) * dbinom(as.numeric(df$k),N, weighted.mean(df$p,df$Freq))
#
#
# plot(df$p,(df$Freq/sum(df$Freq)))
# abline(v=weighted.mean(df$p,df$Freq))
# df
# + language="R"
#
# library(VGAM)
#
# Y <- (df$Freq / sum(df$Freq))
# X <- df$p
#
# fit <- vglm( cbind(df$Freq,df$k) ~ 1, betabinomial, trace=TRUE)
#
# Coef(fit)
#
#
# + language="R"
#
# library(VGAM)
#
# successes <- print(rep(df$k,df$Freq))
# trials <- rep(N,length(successes))
# fit <- vglm( cbind(trials,successes) ~ 1, betabinomial, trace=TRUE)
#
# Coef(fit)
#
#
#
# -
| notebooks/FMT/jnotebooks/Gene Prevalence Model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] tags=["remove_cell"]
# # Quantum Phase Estimation
# -
# ## Contents
#
# 1. [Overview](#overview)
# 1.1 [Intuition](#intuition)
# 1.2 [Mathematical Basis](#maths)
# 2. [Example: T-gate](#example_t_gate)
# 2.1 [Creating the Circuit](#creating_the_circuit)
# 2.2 [Results](#results)
# 3. [Getting More Precision](#getting_more_precision)
# 3.1 [The Problem](#the_problem)
# 3.2 [The Solution](#the_solution)
# 4. [Experimenting on Real Devices](#real_devices)
# 4.1 [With the Circuit from 2.1](#circuit_2.1)
# 5. [Exercises](#exercises)
# 6. [Looking Forward](#looking_forward)
# 7. [References](#references)
# 8. [Contributors](#contributors)
# Quantum phase estimation is one of the most important subroutines in quantum computation. It serves as a central building block for many quantum algorithms. The objective of the algorithm is the following:
#
# Given a unitary operator $U$, the algorithm estimates $\theta$ in $U\vert\psi \rangle =e^{\boldsymbol{2\pi i} \theta }|\psi \rangle$. Here $|\psi\rangle$ is an eigenvector and $e^{\boldsymbol{2\pi i}\theta}$ is the corresponding eigenvalue. Since $U$ is unitary, all of its eigenvalues have a norm of 1.
# ## 1. Overview <a id='overview'></a>
# The general quantum circuit for phase estimation is shown below. The top register contains $t$ 'counting' qubits, and the bottom contains qubits in the state $|\psi\rangle$:
# 
#
# ### 1.1 Intuition <a id='intuition'></a>
# The quantum phase estimation algorithm uses phase kickback to write the phase of $U$ (in the Fourier basis) to the $t$ qubits in the counting register. We then use the inverse QFT to translate this from the Fourier basis into the computational basis, which we can measure.
#
# We remember (from the QFT chapter) that in the Fourier basis the topmost qubit completes one full rotation when counting between $0$ and $2^t$. To count to a number, $x$ between $0$ and $2^t$, we rotate this qubit by $\tfrac{x}{2^t}$ around the z-axis. For the next qubit we rotate by $\tfrac{2x}{2^t}$, then $\tfrac{4x}{2^t}$ for the third qubit.
#
# 
#
# When we use a qubit to control the $U$-gate, the qubit will turn (due to kickback) proportionally to the phase $e^{2i\pi\theta}$. We can use successive $CU$-gates to repeat this rotation an appropriate number of times until we have encoded the phase theta as a number between $0$ and $2^t$ in the Fourier basis.
#
# Then we simply use $QFT^\dagger$ to convert this into the computational basis.
#
#
# ### 1.2 Mathematical Foundation <a id='maths'></a>
#
# As mentioned above, this circuit estimates the phase of a unitary operator $U$. It estimates $\theta$ in $U\vert\psi \rangle =e^{\boldsymbol{2\pi i} \theta }|\psi \rangle$, where $|\psi\rangle$ is an eigenvector and $e^{\boldsymbol{2\pi i}\theta}$ is the corresponding eigenvalue. The circuit operates in the following steps:
#
# i. **Setup**: $\vert\psi\rangle$ is in one set of qubit registers. An additional set of $n$ qubits form the counting register on which we will store the value $2^n\theta$:
#
#
#
# $$ \psi_0 = \lvert 0 \rangle^{\otimes n} \lvert \psi \rangle$$
#
#
#
# ii. **Superposition**: Apply a $n$-bit Hadamard gate operation $H^{\otimes n}$ on the counting register:
#
#
#
# $$ \psi_1 = {\frac {1}{2^{\frac {n}{2}}}}\left(|0\rangle +|1\rangle \right)^{\otimes n} \lvert \psi \rangle$$
#
#
#
# iii. **Controlled Unitary Operations**: We need to introduce the controlled unitary $C-U$ that applies the unitary operator $U$ on the target register only if its corresponding control bit is $|1\rangle$. Since $U$ is a unitary operatory with eigenvector $|\psi\rangle$ such that $U|\psi \rangle =e^{\boldsymbol{2\pi i} \theta }|\psi \rangle$, this means:
#
#
#
# $$U^{2^{j}}|\psi \rangle =U^{2^{j}-1}U|\psi \rangle =U^{2^{j}-1}e^{2\pi i\theta }|\psi \rangle =\cdots =e^{2\pi i2^{j}\theta }|\psi \rangle$$
#
#
#
# Applying all the $n$ controlled operations $C − U^{2^j}$ with $0\leq j\leq n-1$, and using the relation $|0\rangle \otimes |\psi \rangle +|1\rangle \otimes e^{2\pi i\theta }|\psi \rangle =\left(|0\rangle +e^{2\pi i\theta }|1\rangle \right)\otimes |\psi \rangle$:
#
# \begin{aligned}
# \psi_{2} & =\frac {1}{2^{\frac {n}{2}}} \left(|0\rangle+{e^{\boldsymbol{2\pi i} \theta 2^{n-1}}}|1\rangle \right) \otimes \cdots \otimes \left(|0\rangle+{e^{\boldsymbol{2\pi i} \theta 2^{1}}}\vert1\rangle \right) \otimes \left(|0\rangle+{e^{\boldsymbol{2\pi i} \theta 2^{0}}}\vert1\rangle \right) \otimes |\psi\rangle\\\\
# & = \frac{1}{2^{\frac {n}{2}}}\sum _{k=0}^{2^{n}-1}e^{\boldsymbol{2\pi i} \theta k}|k\rangle \otimes \vert\psi\rangle
# \end{aligned}
# where $k$ denotes the integer representation of n-bit binary numbers.
#
# iv. **Inverse Fourier Transform**: Notice that the above expression is exactly the result of applying a quantum Fourier transform as we derived in the notebook on [Quantum Fourier Transform and its Qiskit Implementation](qft.ipynb). Recall that QFT maps an n-qubit input state $\vert x\rangle$ into an output as
#
# $$
# QFT\vert x \rangle = \frac{1}{2^\frac{n}{2}}
# \left(\vert0\rangle + e^{\frac{2\pi i}{2}x} \vert1\rangle\right)
# \otimes
# \left(\vert0\rangle + e^{\frac{2\pi i}{2^2}x} \vert1\rangle\right)
# \otimes
# \ldots
# \otimes
# \left(\vert0\rangle + e^{\frac{2\pi i}{2^{n-1}}x} \vert1\rangle\right)
# \otimes
# \left(\vert0\rangle + e^{\frac{2\pi i}{2^n}x} \vert1\rangle\right)
# $$
#
# Replacing $x$ by $2^n\theta$ in the above expression gives exactly the expression derived in step 2 above. Therefore, to recover the state $\vert2^n\theta\rangle$, apply an inverse Fourier transform on the ancilla register. Doing so, we find
#
# $$
# \vert\psi_3\rangle = \frac {1}{2^{\frac {n}{2}}}\sum _{k=0}^{2^{n}-1}e^{\boldsymbol{2\pi i} \theta k}|k\rangle \otimes | \psi \rangle \xrightarrow{\mathcal{QFT}_n^{-1}} \frac {1}{2^n}\sum _{x=0}^{2^{n}-1}\sum _{k=0}^{2^{n}-1} e^{-\frac{2\pi i k}{2^n}(x - 2^n \theta)} |x\rangle \otimes |\psi\rangle
# $$
#
# v. **Measurement**:
# The above expression peaks near $x = 2^n\theta$. For the case when $2^n\theta$ is an integer, measuring in the computational basis gives the phase in the ancilla register with high probability:
#
#
#
# $$ |\psi_4\rangle = | 2^n \theta \rangle \otimes | \psi \rangle$$
#
#
#
# For the case when $2^n\theta$ is not an integer, it can be shown that the above expression still peaks near $x = 2^n\theta$ with probability better than $4/\pi^2 \approx 40\%$ [1].
# ## 2. Example: T-gate <a id='example_t_gate'></a>
# Let’s take a gate we know well, the $T$-gate, and use Quantum Phase Estimation to estimate its phase. You will remember that the $T$-gate adds a phase of $e^\frac{i\pi}{4}$ to the state $|1\rangle$:
#
# $$ T|1\rangle =
# \begin{bmatrix}
# 1 & 0\\
# 0 & e^\frac{i\pi}{4}\\
# \end{bmatrix}
# \begin{bmatrix}
# 0\\
# 1\\
# \end{bmatrix}
# = e^\frac{i\pi}{4}|1\rangle $$
#
# Since QPE will give us $\theta$ where:
#
#
#
# $$ T|1\rangle = e^{2i\pi\theta}|1\rangle $$
#
#
#
# We expect to find:
#
#
#
# $$\theta = \frac{1}{8}$$
#
#
#
# In this example we will use three qubits and obtain an _exact_ result (not an estimation!)
# ### 2.1 Creating the Circuit <a id='creating_the_circuit'></a>
# Let's first prepare our environment:
# +
#initialization
import matplotlib.pyplot as plt
import numpy as np
import math
# importing Qiskit
from qiskit import IBMQ, Aer
from qiskit import QuantumCircuit, ClassicalRegister, QuantumRegister, execute
# import basic plot tools
from qiskit.visualization import plot_histogram
# -
# Now, set up the quantum circuit. We will use four qubits -- qubits 0 to 2 as counting qubits, and qubit 3 as the eigenstate of the unitary operator ($T$).
#
# We initialize $\vert\psi\rangle = \vert1\rangle$ by applying an $X$ gate:
qpe = QuantumCircuit(4, 3)
qpe.x(3)
qpe.draw()
# Next, we apply Hadamard gates to the counting qubits:
for qubit in range(3):
qpe.h(qubit)
qpe.draw()
# Next we perform the controlled unitary operations. **Remember:** Qiskit orders its qubits the opposite way round to the image above.
repetitions = 1
for counting_qubit in range(3):
for i in range(repetitions):
qpe.cu1(math.pi/4, counting_qubit, 3); # This is C-U
repetitions *= 2
qpe.draw()
# We apply the inverse quantum Fourier transformation to convert the state of the counting register. Here we provide the code for $QFT^\dagger$:
def qft_dagger(circ, n):
"""n-qubit QFTdagger the first n qubits in circ"""
# Don't forget the Swaps!
for qubit in range(n//2):
circ.swap(qubit, n-qubit-1)
for j in range(n):
for m in range(j):
circ.cu1(-math.pi/float(2**(j-m)), m, j)
circ.h(j)
# We then measure the counting register:
qpe.barrier()
# Apply inverse QFT
qft_dagger(qpe, 3)
# Measure
qpe.barrier()
for n in range(3):
qpe.measure(n,n)
qpe.draw()
# ### 2.2 Results <a id='results'></a>
# +
backend = Aer.get_backend('qasm_simulator')
shots = 2048
results = execute(qpe, backend=backend, shots=shots).result()
answer = results.get_counts()
plot_histogram(answer)
# -
# We see we get one result (`001`) with certainty, which translates to the decimal: `1`. We now need to divide our result (`1`) by $2^n$ to get $\theta$:
#
#
#
# $$ \theta = \frac{1}{2^3} = \frac{1}{8} $$
#
#
#
# This is exactly the result we expected!
# ## 3. Example: Getting More Precision <a id='getting_more_precision'></a>
# ### 3.1 The Problem <a id='the_problem'></a>
#
# Instead of a $T$-gate, let’s use a gate with $\theta = \frac{1}{3}$. We set up our circuit as with the last example:
# +
# Create and set up circuit
qpe2 = QuantumCircuit(4, 3)
# Apply H-Gates to counting qubits:
for qubit in range(3):
qpe2.h(qubit)
# Prepare our eigenstate |psi>:
qpe2.x(3)
# Do the controlled-U operations:
angle = 2*math.pi/3
repetitions = 1
for counting_qubit in range(3):
for i in range(repetitions):
qpe2.cu1(angle, counting_qubit, 3);
repetitions *= 2
# Do the inverse QFT:
qft_dagger(qpe2, 3)
# Measure of course!
for n in range(3):
qpe2.measure(n,n)
qpe2.draw()
# +
# Let's see the results!
backend = Aer.get_backend('qasm_simulator')
shots = 4096
results = execute(qpe2, backend=backend, shots=shots).result()
answer = results.get_counts()
plot_histogram(answer)
# -
# We are expecting the result $\theta = 0.3333\dots$, and we see our most likely results are `010(bin) = 2(dec)` and `011(bin) = 3(dec)`. These two results would tell us that $\theta = 0.25$ (off by 25%) and $\theta = 0.375$ (off by 13%) respectively. The true value of $\theta$ lies between the values we can get from our counting bits, and this gives us uncertainty and imprecision.
#
# ### 3.2 The Solution <a id='the_solution'></a>
# To get more precision we simply add more counting qubits. We are going to add two more counting qubits:
# +
# Create and set up circuit
qpe3 = QuantumCircuit(6, 5)
# Apply H-Gates to counting qubits:
for qubit in range(5):
qpe3.h(qubit)
# Prepare our eigenstate |psi>:
qpe3.x(5)
# Do the controlled-U operations:
angle = 2*math.pi/3
repetitions = 1
for counting_qubit in range(5):
for i in range(repetitions):
qpe3.cu1(angle, counting_qubit, 5);
repetitions *= 2
# Do the inverse QFT:
qft_dagger(qpe3, 5)
# Measure of course!
qpe3.barrier()
for n in range(5):
qpe3.measure(n,n)
qpe3.draw()
# +
### Let's see the results!
backend = Aer.get_backend('qasm_simulator')
shots = 4096
results = execute(qpe3, backend=backend, shots=shots).result()
answer = results.get_counts()
plot_histogram(answer)
# -
# The two most likely measurements are now `01011` (decimal 11) and `01010` (decimal 10). Measuring these results would tell us $\theta$ is:
#
# $$
# \theta = \frac{11}{2^5} = 0.344,\;\text{ or }\;\; \theta = \frac{10}{2^5} = 0.313
# $$
#
# These two results differ from $\frac{1}{3}$ by 3% and 6% respectively. A much better precision!
# ## 4. Experiment with Real Devices <a id='real_devices'></a>
# ### 4.1 Circuit from 2.1 <a id='circuit_2.1'></a>
#
# We can run the circuit in section 2.1 on a real device, let's remind ourselves of the circuit:
qpe.draw()
# + tags=["uses-hardware"]
# Load our saved IBMQ accounts and get the least busy backend device with less than or equal to n qubits
IBMQ.load_account()
from qiskit.providers.ibmq import least_busy
from qiskit.tools.monitor import job_monitor
provider = IBMQ.get_provider(hub='ibm-q')
backend = provider.get_backend('ibmq_vigo')
# Run with 2048 shots
shots = 2048
job = execute(qpe, backend=backend, shots=2048, optimization_level=3)
job_monitor(job)
# + tags=["uses-hardware"]
# get the results from the computation
results = job.result()
answer = results.get_counts(qpe)
plot_histogram(answer)
# -
# We can hopefully see that the most likely result is `001` which is the result we would expect from the simulator. Unlike the simulator, there is a probability of measuring something other than `001`, this is due to noise and gate errors in the quantum computer.
# ## 5. Exercises <a id='exercises'></a>
# 1. Try the experiments above with different gates ($\text{CNOT}$, $S$, $T^\dagger$), what results do you expect? What results do you get?
#
# 2. Try the experiment with a $Y$-gate, do you get the correct result? (Hint: Remember to make sure $|\psi\rangle$ is an eigenstate of $Y$!)
#
# ## 6. Looking Forward <a id='looking_forward'></a>
#
# The quantum phase estimation algorithm may seem pointless, since we have to know $\theta$ to perform the controlled-$U$ operations on our quantum computer. We will see in later chapters that it is possible to create circuits for which we don’t know $\theta$, and for which learning theta can tell us something very useful (most famously how to factor a number!)
# ## 7. References <a id='references'></a>
#
# [1] <NAME> and <NAME>. 2011. Quantum Computation and Quantum Information: 10th Anniversary Edition (10th ed.). Cambridge University Press, New York, NY, USA.
# ## 8. Contributors <a id='contributors'></a>
# 03/20/2020 — <NAME> (@HwajungKang) — Fixed inconsistencies with qubit ordering
import qiskit
qiskit.__qiskit_version__
| 4.b Quantum-phase-estimation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# prerequisite package imports
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sb
# %matplotlib inline
from solutions_explain import polishing_solution_1
# -
# In this workspace, you'll perform some polishing and cleanup of a plot created using the Pokémon creature dataset.
pokemon = pd.read_csv('./data/pokemon.csv')
pokemon.head()
# **Task**: Perform polishing and cleaning steps for the following multivariate plot, where the relationship between height and weight are depicted for the Fairy and Dragon Pokémon types. You'll need to do more than just add meaningful labels and titles to the plot. Pay attention to the interpretability of tick marks and the ability to associate each point to a type of Pokemon.
#
# For reference, height is in terms of meters and weight is in terms of kilograms. One other thing you might try is to change the colors mapped to each type to match those given by the game series. Fairy is depicted with a light pink (hex code #ee99ac) while Dragon is associated with a medium purple (hex code #7038f8). (Type colors taken from [Pokémon Wiki Bulbapedia](https://bulbapedia.bulbagarden.net/wiki/Category:Type_color_templates).)
# +
# data processing: isolate only fairy and dragon type Pokemon
type_cols = ['type_1','type_2']
non_type_cols = pokemon.columns.difference(type_cols)
pkmn_types = pokemon.melt(id_vars = non_type_cols, value_vars = type_cols,
var_name = 'type_level', value_name = 'type').dropna()
pokemon_sub = pkmn_types.loc[pkmn_types['type'].isin(['fairy','dragon'])]
# -
# MODIFY THIS CODE
g = sb.FacetGrid(data = pokemon_sub, hue = 'type', size = 5)
g.map(plt.scatter, 'weight','height')
g.set(xscale = 'log')
# run this cell to check your work against ours
polishing_solution_1()
| Matplotlib_multivariate/Polishing_Plots_Practice.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="TfGjejTgVm9c"
# # 1. while문
# + id="08cARjCLVm9e"
while 조건 :
수행할 문장1
수행할 문장2
# + id="9AXgpXBhVnAU" outputId="40d5fbf0-6705-4943-8d38-9f333dc47d0d"
# 10이하의 짝수 프린트하기
i = 1
while i <= 10 :
if i % 2 == 0 :
print(i)
i += 1
# + [markdown] id="zLJk0i7xxF9d"
# ## break
# + id="Fp3oqutHVnAa" outputId="93ec3bb4-2fc8-469a-cc79-e111d4378d71"
# 100번째 방문자 찾기
i = 90
while i :
i += 1
if i == 100 :
print("축하합니다. %d번째 방문자입니다." % i)
break
print("감사합니다. 이벤트가 종료되었습니다.")
# + [markdown] id="ba6TllU_xHgf"
# ## continue
# + id="GTXmxI3hVnAe" outputId="ca9beef5-6b64-4e4b-a497-624728d45afd"
i = 0
while i < 11 :
i += 1
if i == 6 :
continue
if i % 2 == 0 :
print(i)
# + [markdown] id="A6xcfQN0Vm7M"
# # 2. for문
# + id="f_291YetVm7c"
for 변수 in range(변수가 속한 자료형, 혹은 변수의 범위) :
수행해야할 문장1
수행해야할 문장2
# + [markdown] id="lnLCmlMXSvt0"
# ## 변수에 범위를 지정하기
# ### 1. range
# + id="WgQt89R9Vm72" outputId="6b3e8134-5ae0-4a0b-f253-f6329ba4df57"
for x in range(0,5) :
print(x)
# + [markdown] id="CTMs6r4RQvr6"
# ## 연습문제 7
# 카운트 다운 만들어보기
# + colab={"base_uri": "https://localhost:8080/", "height": 119} id="KbpbBXVVVm8V" executionInfo={"elapsed": 654, "status": "ok", "timestamp": 1593354107527, "user": {"displayName": "pam S", "photoUrl": "", "userId": "14096643837279750054"}, "user_tz": -540} outputId="30c681a3-a8cd-4d31-c392-c26278c46667"
# 정답을 작성해주세요.
# + [markdown] id="netH8iM8TDc2"
# ### 2. 자료형으로 범위주기
# + id="cRwbBTrDVm86" outputId="bf19068a-43f8-45be-c21b-4eaa9b7acb13"
word = 'Hello!'
for w in word:
print(w)
# + id="LKo-KX3NVm9F" outputId="6056c562-26fa-4a45-d0da-4dfce18f0578"
for a, b in [(2,1), (2,2), (2,3), (2,4)] :
print(a*b)
# + [markdown] id="qMHHji5pRBLo"
# ## 연습문제 8
# 구구단 2단과 3단을 출력하기
# + colab={"base_uri": "https://localhost:8080/", "height": 357} id="QQGa8qMeVnAi" executionInfo={"elapsed": 1162, "status": "ok", "timestamp": 1593354198456, "user": {"displayName": "pam S", "photoUrl": "", "userId": "14096643837279750054"}, "user_tz": -540} outputId="cb3c0cf6-ad58-4732-e380-aefca1031320"
# 정답을 작성해주세요.
# + id="bQNXp8ZrT5iO"
| python/practice/1-4_loops.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# <center> <h1> <b>Access Modifiers
# <b>Access modifiers</b> (or <i>access specifiers</i>) are keywords in object-oriented languages that set the accessibility of classes, methods, and other members. Generally there are three types of Access modifiers, <b><i>Public</i></b> , <b><i>Protected</i></b> , <b><i>Private</i></b>. Private access modifier is the most secure access modifier.
# <img src="https://miro.medium.com/max/1400/1*AsKRlD4xL50sqSDAOYvhMA.jpeg">
# <h1><b>Public Access Modifier:</b></h1>
# The members of a class that are declared public are easily accessible from any part of the program. All data members and member functions of a class are public by default.
# <h2><b>Python 3</b></h2>
# +
#program to illustrate public access modifier in a class
class Acc:
# constructor
def __init__(self, name, age):
#public data members (instance variable)
self.name = name
self.age = age
# public member function
def displayAge(self):
#accessing public data member
print("Age : ",self.age)
# creating object of the class
obj = Acc("John",22)
# accessing public data member
print("Name : ",obj.name)
#calling the public member function of the class
obj.displayAge()
# -
# <h1><b>Protected Access Modifier:</b></h1>
# The members of a class that are declared protected are only accessible to a class derived from it (child class). Data members of a class are declared protected by adding a single underscore '_' in python before the data member of that class.
# <h2><b>Python 3</b></h2>
# +
#program to illustrate protected access modifier in a class
# super class or parent class
class Student:
#protected data members or class variable
_name = None
_roll = None
_branch = None
#constructor (initializing instance variable)
def __init__(self, name, roll , branch):
self._name = name
self._roll = roll
self._branch = branch
#protected member function
def _display(self):
#accessing protected data members
print("Roll : ",self._roll)
# derived or child class
class Pupil(Student):
#constructor
def __init__(self, name, roll, branch):
super().__init__(name,roll,branch)
#public member function
def display(self):
#accessing protected data members of super class
print("Name : ",self._name)
#accessing protected member function of super class
self._display()
# creating objects of the derived class
obj = Pupil("Nathan",123456,"CSE")
#calling public member function functions of the class
obj.display()
# -
# <h1><b>Private Access Modifier:</b></h1>
# The members of a class that are declared private are accessible within the class only, private access modifier is the most secure access modifier. Data members of a class are declared private by adding a double underscore ‘__’ symbol before the data member of that class.
# <h2><b>Python3</b></h2>
# +
# program to illustrate private access modifier in a class
class Geek:
# private members
__name = None
__roll = None
__branch = None
# constructor
def __init__(self, name, roll, branch):
self.__name = name
self.__roll = roll
self.__branch = branch
# private member function
def __displayDetails(self):
# accessing private data members
print("Name: ", self.__name)
print("Roll: ", self.__roll)
print("Branch: ", self.__branch)
# public member function
def accessPrivateFunction(self):
# accessing private member function
self.__displayDetails()
# creating object
obj = Geek("R2J", 1706256, "Information Technology")
# calling public member function of the class
obj.accessPrivateFunction()
# -
| Access Modifiers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # IFRS9 portfolio analytics
# In this notebook I aim to provide a practical implementation to address daily credit portfolio monitoring needs, in particular to track and explain **expected credit losses per IFRS 9 standard**, perform vintage analysis, drill-down to loan-level data, analyze changes between periods and determine main drivers behind the portfolio risk KPIs.
#
# I hope this example can serve as inspiration for your work in the IFRS field.
# As a quick reminder, as a result of the Great Financial Crisis of 2008, the accounting standards were redesigned, which culminated with the International Financial Reporting Standard number 9 (IFRS 9) and Current Expected Credit Loss (CECL). They key innovation of the new standard was the requirement to recognize in financial statements not only the incurred losses for a credit portfolio, but compute a forward looking measure - expected credit losses for a portfolio.
#
# By common practice, the expected credit losses computation relies on the three ingredients: probability of default (PD), loss given default (LGD), and exposure at default (EAD) framework and the three stage classification. Continue reading to learn how loan-level credit risk data can be consolidated in an analytical app - IFRS 9 app - enriched with static attributes and analized interactively in the UI.
#
# <img src=./ifrs9-app.png/ width = 70%>
#
# We'll start by creating an atoti app, then we'll load sample data and implement chains of calculations for ECL and other analytical measures.
# <div style="text-align: center;" ><a href="https://www.atoti.io/?utm_source=gallery&utm_content=ifrs9" target="_blank" rel="noopener noreferrer"><img src="https://data.atoti.io/notebooks/banners/discover.png" alt="Try atoti"></a></div>
# # Launch atoti
# Let's create an atoti session to launch the aggregation engine and the UI.
# + tags=[]
import atoti as tt
session = tt.create_session(
config={"user_content_storage": "content", "java_options": ["-Xmx8G"]}
)
session.link()
# -
# ## Input Data overview
#
# In this section we will load the data used by the ECL calculation into in-memory datastores.
#
# The sample data will be fetched from csv files hosted on s3, and you can replace them with your own sources.
#
# I’ve created a sample data set based on the [Lending Club dataset available on Kaggle](https://www.kaggle.com/janiobachmann/datasets) and mocked-up the risk engine outputs - PDs, LGDs, EADs and stages - to illustrate how the IFRS 9 metrics can be computed and visualized in an analytical app.
#
# We'll use these three datastores:
#
# - `risk_engine_data` keeps information on the individual loans, their explosures, stages, credit risk parameters for ECL calculation and other attributes by date,
# - `lending_club_data` keeps additional data - loan attributes and amounts
# - `loans_at_inception` is storing loan opening information, such as opening date and opening risk characteristics.
# ### Credit Risk Engine Inputs
# +
# # !conda install -c conda-forge python-wget -y
# +
from zipfile import ZipFile
import wget
from IPython.display import clear_output, display
# +
def bar_custom(current, total, width=80):
clear_output(wait=True)
print("Downloading: %d%% [%d / %d] bytes" % (current / total * 100, current, total))
url = "https://s3.eu-west-3.amazonaws.com/data.atoti.io/notebooks/ifrs9/lending-club-data.zip"
filename = wget.download(url, bar=bar_custom)
# -
# unzipping the file
with ZipFile("lending-club-data.zip", "r") as zipObj:
# Extract all the contents of zip file in current directory
zipObj.extractall()
src = "./"
risk_engine_data = session.read_csv(
src + "lending-club-data/risk-engine/*.csv",
keys=["Reporting Date", "id"],
table_name="Credit Risk",
types={
"EAD": tt.type.NULLABLE_DOUBLE,
"Stage": tt.type.NULLABLE_INT,
"Previous Stage": tt.type.NULLABLE_INT,
"DaysPastDue": tt.type.NULLABLE_DOUBLE,
},
)
risk_engine_data.head(3)
# ### Creating a cube
# As a next step, I'm creating the cube.
cube = session.create_cube(risk_engine_data, "IFRS9")
# ### Adding LendingClub data
# The [LendingClub](https://en.wikipedia.org/wiki/LendingClub) data I'm using for illustrative purposes can be downloaded from: [<NAME> on Kaggle](https://www.kaggle.com/janiobachmann/datasets). Here's a description of the dataset and field description: [A Hitchhiker's Guide to Lending Club Loan Data
# ](https://www.kaggle.com/pragyanbo/a-hitchhiker-s-guide-to-lending-club-loan-data).
lending_club_data = session.read_csv(
src + "lending-club-data/loans.csv",
keys=["id"],
table_name="Lending Club Data",
process_quotes=True,
)
risk_engine_data.join(lending_club_data)
lending_club_data.head(3)
# See later how to create measures aggregating Lending Club data.
# ### Adding loans opening parameters
loans_at_inception = session.read_csv(
"lending-club-data/static.csv",
keys=["id"],
table_name="Parameters at inception",
)
loans_at_inception.head(3)
# Linking contracts and their opening information
risk_engine_data.join(loans_at_inception)
# ### Cube schema
# Based on the datastores we've loaded and linked together, this is the view of our current cube schema.
cube.schema
# # Getting started with cube configuration
#
# In this section I'm demonstrating some of the typical actions that you can apply to your newly created cube.
# These are the variables for faster access to the cube data elements:
l, m, h = cube.levels, cube.measures, cube.hierarchies
# By setting the sort on the reporting date to DESC, we make sure that the latest date appears by default.
l["Reporting Date"].comparator = tt.comparator.DESCENDING
# Multi-level hierarchies come handy when you have a typical way to expand data:
h["Opening Date (detailed)"] = {
"Year": lending_club_data["Opening Year"],
"Month": lending_club_data["Opening Month"],
"Day": lending_club_data["Opening Day"],
}
# +
# with 0.6.0, int and long table columns, unless they are key columns, automatically become measures instead of levels.
# With this change, all the numeric columns behave the same.
h["Stage"] = [risk_engine_data["Stage"]]
h["Months Since Inception"] = [risk_engine_data["Months Since Inception"]]
# -
# # Measures visualizing credit risk inputs
# In this section you will find various measures that we create to visualize risk engine output in the UI.
# ## Stages and their variations
#
# In this section, we define measures to visualize Stages and their variations.
# As a reminder, **Impairment** of financial assets is recognised in stages:
#
# - **Stage 1** Low credit risk or no increase since initial recognition: one-year ECL
# - **Stage 2** If the credit risk increases significantly and is not considered low: lifetime ECL
# - **Stage 3** impaired assets
# +
m["Stage"] = tt.value(risk_engine_data["Stage"])
m["Stage"].folder = "Stage"
# This is how we can create measures to display the previous value and daily changes side by side with the value:
m["Previous Stage"] = tt.shift(m["Stage"], on=l["Reporting Date"], offset=1)
m["Previous Stage"].folder = "Stage"
m["Stage Variation"] = tt.where(
m["Previous Stage"] != None, m["Stage"] - m["Previous Stage"]
)
m["Stage Variation"].folder = "Stage"
m["Stage Variation"].formatter = "DOUBLE[+#,###;-#,###]"
# -
# ## EAD aggregation
# The **Exposure At Default** is the estimated amount of loss a bank may be exposed to when a debtor defaults on a loan.
# +
m["EAD"] = tt.agg.sum(risk_engine_data["EAD"])
m["EAD"].folder = "EAD"
# This is how we can create measures to display the previous value and daily changes side by side with the value:
m["Previous EAD"] = tt.shift(m["EAD"], on=l["Reporting Date"], offset=1)
m["Previous EAD"].folder = "EAD"
m["EAD (Chg)"] = tt.where(m["Previous EAD"] != None, m["EAD"] - m["Previous EAD"])
m["EAD (Chg)"].folder = "EAD"
m["EAD (Chg)"].formatter = "DOUBLE[+#,###.##;-#,###.##]"
m["EAD (Chg %)"] = tt.where(m["Previous EAD"] != 0, m["EAD (Chg)"] / m["Previous EAD"])
m["EAD (Chg %)"].folder = "EAD"
m["EAD (Chg %)"].formatter = "DOUBLE[+#,###.##%;-#,###.##%]"
# -
# Here's a query for the EAD and variations:
cube.query(
m["EAD"],
m["Previous EAD"],
m["EAD (Chg)"],
m["EAD (Chg %)"],
levels=[l["Reporting Date"]],
)
# ## Visualizing PD
# The **Probability Of Default** is the likelihood that your debtor will default on its debts (goes bankrupt or so) within certain period (12 months for loans in Stage 1 and life-time for other loans).
#
# PD (12) - is 12 month probability of default and PD (LT) - is the lifetime probability of default.
# ### 12 months PD and variations
# +
m["PD (12)"] = tt.agg.mean(risk_engine_data["PD12"])
m["PD (12)"].folder = "PD"
m["PD (12)"].formatter = "DOUBLE[#,###.##%]"
m["Previous PD (12)"] = tt.shift(m["PD (12)"], on=l["Reporting Date"], offset=1)
m["Previous PD (12)"].folder = "PD"
m["Previous PD (12)"].formatter = "DOUBLE[#,###.##%]"
m["PD (12) (Chg)"] = tt.where(
m["Previous PD (12)"] != None, m["PD (12)"] - m["Previous PD (12)"]
)
m["PD (12) (Chg)"].folder = "PD"
m["PD (12) (Chg)"].formatter = "DOUBLE[+#,###.##%;-#,###.##%]"
# -
# The measures display average across contracts, but require having the slicing hierarchies in the view.
# ### Lifetime PD and variations
# +
m["PD (LT)"] = tt.agg.mean(risk_engine_data["PDLT"])
m["PD (LT)"].folder = "PD"
m["PD (LT)"].formatter = "DOUBLE[#,###.##%]"
m["Opening PD (LT)"] = tt.agg.mean(loans_at_inception["Opening PDLT"])
m["Opening PD (LT)"].folder = "PD"
m["Previous PD (LT)"] = tt.shift(m["PD (LT)"], on=l["Reporting Date"], offset=1)
m["Previous PD (LT)"].folder = "PD"
m["PD (LT) (Chg)"] = tt.where(
m["Previous PD (LT)"] != None, m["PD (LT)"] - m["Previous PD (LT)"]
)
m["PD (LT) (Chg)"].folder = "PD"
# Variation from opening
m["PD (LT) Variation"] = (m["PD (LT)"] - m["Opening PD (LT)"]) / m["Opening PD (LT)"]
m["PD (LT) Variation"].folder = "PD"
m["PD (LT) Variation"].formatter = "DOUBLE[+#,###.##%;-#,###.##%]"
# -
# ## Visualizing LGD
# The **Lost Given Default** is the percentage that you can lose when the debtor defaults.
# +
m["LGD"] = tt.agg.mean(risk_engine_data["LGD"])
m["LGD"].folder = "LGD"
m["LGD"].formatter = "DOUBLE[#,###.##%]"
m["Previous LGD"] = tt.shift(m["LGD"], on=l["Reporting Date"], offset=1)
m["Previous LGD"].folder = "LGD"
m["Previous LGD"].formatter = "DOUBLE[#,###.##%]"
# -
# ## Summary
#
# In this section we have created measures for credit risk inputs - EAD, PD, LGD, stages and their variations.
#
# Other measures that might be useful for visualization include - curing time, time weighted by EAD, statistics on the number of days past due and other.
# # ECL computation
# The **Expected Credit Loss** is a probability-weighted estimate of credit loss. Depending on the "Stage" of a loan, the ECL is computed per one of the following formulae:
#
# - IFRS Stage 1: $ECL=EAD \cdot PD^{12M} \cdot LGD$
# - IFRS Stage 2: $ECL=EAD \cdot PD^{LT} \cdot LGD$
# - IFRS Stage 3: $ECL=EAD \cdot LGD$
#
# Let's define each of these expressions:
ecl_stage_1 = tt.agg.sum_product(
risk_engine_data["LGD"], risk_engine_data["EAD"], risk_engine_data["PD12"]
)
ecl_stage_2 = tt.agg.sum_product(
risk_engine_data["LGD"], risk_engine_data["EAD"], risk_engine_data["PDLT"]
)
ecl_stage_3 = tt.agg.sum_product(risk_engine_data["LGD"], risk_engine_data["EAD"])
# Now, the measure visible in the UI will pick the correct formula depending on the stage:
m["ECL"] = (
tt.filter(ecl_stage_1, l["Stage"] == 1)
+ tt.filter(ecl_stage_2, l["Stage"] == 2)
+ tt.filter(ecl_stage_3, l["Stage"] == 3)
)
m["ECL"].folder = "ECL"
cube.query(m["ECL"], levels=[l["Reporting Date"]])
# As usual, let's create measures to visualize the previous reporting date values and changes:
# +
m["Previous ECL"] = tt.shift(m["ECL"], on=l["Reporting Date"], offset=1)
m["Previous ECL"].folder = "ECL"
m["ECL (Chg)"] = tt.where(m["Previous ECL"] != None, m["ECL"] - m["Previous ECL"])
m["ECL (Chg)"].folder = "ECL"
m["ECL (Chg %)"] = tt.where(m["Previous ECL"] != 0, m["ECL (Chg)"] / m["Previous ECL"])
m["ECL (Chg %)"].folder = "ECL"
m["ECL (Chg %)"].formatter = "DOUBLE[+#,###.##%;-#,###.##%]"
m["ECL of old contracts"] = tt.where(l["Reporting Date"] != l["issue_d"], m["ECL"])
m["ECL of old contracts"].folder = "ECL"
m["ECL (Chg without new contracts)"] = tt.where(
m["Previous ECL"] != None, m["ECL of old contracts"] - m["Previous ECL"]
)
m["ECL (Chg without new contracts)"].folder = "ECL"
m["ECL (Chg % without new contracts)"] = tt.where(
m["Previous ECL"] != 0, m["ECL (Chg without new contracts)"] / m["Previous ECL"]
)
m["ECL (Chg % without new contracts)"].folder = "ECL"
m["ECL (Chg % without new contracts)"].formatter = "DOUBLE[+#,###.##%;-#,###.##%]"
# -
# Now the ECL measures are computed from the EAD, PD and LGD every time a user displays ECL. This allows to manipulate the inputs - see in the measure simulations section. Now let's explore the ECL analytics in more detail and introduce the conpect of ECL explainers.
# # ECL change explainers
# Lets introduce **ECL Explainer** as measures attributing the change in ECL to the underlying factors - PD, LGD and EAD. To compute them, we will apply the same formula as for the ECL measure, but will keep each of the three factors at the level of the previous reporting date, so it has no effect on the ECL variation.
#
# For the simplicity of this example we don't take into account the stage migrations. The methodology could be improved to reflect the effect of stage changes.
# ## ECL variation due to PD changes
# +
ecl_pd_explain_stage_1 = tt.agg.sum_product(
risk_engine_data["LGD"],
risk_engine_data["EAD"],
risk_engine_data["Previous PD12"],
)
ecl_pd_explain_stage_2 = tt.agg.sum_product(
risk_engine_data["LGD"],
risk_engine_data["EAD"],
risk_engine_data["Previous PDLT"],
)
m["ECL with previous PD"] = (
tt.filter(ecl_pd_explain_stage_1, l["Stage"] == 1)
+ tt.filter(ecl_pd_explain_stage_2, l["Stage"] == 2)
+ tt.filter(ecl_stage_3, l["Stage"] == 3)
)
m["ECL variation due to PD changes"] = m["ECL"] - m["ECL with previous PD"]
m["ECL variation due to PD changes"].folder = "ECL"
# -
cube.query(
m["ECL (Chg)"],
m["ECL variation due to PD changes"],
levels=[l["Reporting Date"]],
)
# ## ECL variation due to LGD changes
# +
ecl_lgd_explain_stage_1 = tt.agg.sum_product(
risk_engine_data["Previous LGD"],
risk_engine_data["EAD"],
risk_engine_data["PD12"],
)
ecl_lgd_explain_stage_2 = tt.agg.sum_product(
risk_engine_data["Previous LGD"],
risk_engine_data["EAD"],
risk_engine_data["PDLT"],
)
ecl_lgd_explain_stage_3 = tt.agg.sum_product(
risk_engine_data["Previous LGD"], risk_engine_data["EAD"]
)
m["ECL with previous LGD"] = (
tt.filter(ecl_lgd_explain_stage_1, l["Stage"] == 1)
+ tt.filter(ecl_lgd_explain_stage_2, l["Stage"] == 2)
+ tt.filter(ecl_lgd_explain_stage_3, l["Stage"] == 3)
)
m["ECL variation due to LGD changes"] = m["ECL"] - m["ECL with previous LGD"]
m["ECL variation due to LGD changes"].folder = "ECL"
# -
m["ecl_lgd_explain_stage_1"] = ecl_lgd_explain_stage_1
cube.query(
m["ECL (Chg)"],
m["ECL variation due to PD changes"],
m["ECL variation due to LGD changes"],
levels=[l["Reporting Date"]],
)
session.link()
# ## ECL variation due to EAD changes
# +
ecl_ead_explain_stage_1 = tt.agg.sum_product(
risk_engine_data["LGD"],
risk_engine_data["Previous EAD"],
risk_engine_data["PD12"],
)
ecl_ead_explain_stage_2 = tt.agg.sum_product(
risk_engine_data["LGD"],
risk_engine_data["Previous EAD"],
risk_engine_data["PDLT"],
)
ecl_ead_explain_stage_3 = tt.agg.sum_product(
risk_engine_data["LGD"], risk_engine_data["Previous EAD"]
)
m["ECL with previous EAD"] = (
tt.filter(ecl_ead_explain_stage_1, l["Stage"] == 1)
+ tt.filter(ecl_ead_explain_stage_2, l["Stage"] == 2)
+ tt.filter(ecl_ead_explain_stage_3, l["Stage"] == 3)
)
m["ECL variation due to EAD changes"] = m["ECL"] - m["ECL with previous EAD"]
m["ECL variation due to EAD changes"].folder = "ECL"
# -
cube.query(
m["ECL (Chg)"],
m["ECL variation due to PD changes"],
m["ECL variation due to LGD changes"],
m["ECL variation due to EAD changes"],
levels=[l["Reporting Date"]],
)
# ## Unexplained variation
# You can notice that the unexplained ECL is quite huge - because our approach doesn't account for trade transitions.
m["ECL unexplained variation"] = (
m["ECL (Chg)"]
- m["ECL variation due to PD changes"]
- m["ECL variation due to LGD changes"]
- m["ECL variation due to EAD changes"]
)
cube.query(
m["ECL (Chg)"],
m["ECL variation due to PD changes"],
m["ECL variation due to LGD changes"],
m["ECL variation due to EAD changes"],
m["ECL unexplained variation"],
levels=[l["Reporting Date"]],
)
# # Vintage analysis
# In vintage analysis, the portfolio is broken down into “vintages” based on the origination month. This is because credit portfolios follow a certain lifecycle pattern with more loans defaulting in the first months of inception and stabilizing after a certain period.
#
# The Vintage analysis is simple in our IFRS 9 app:
#
# - to break the portfolio into vintages, we will use "Opening date" hierarchy and put it on the rows if a pivot table
# - then to observe the evolution of portfolio through time - we will put the "Reporting Date" hierarchy onto the columns of a pivot table.
#
# Then inside the pivot table, we can display any measure you want, for example, ECL, EAD, % of customers past due, % of loans past due.
# Let's take an example of the % of loans past due and create a special measure to visualize it.
#
# Total number of loans is available in the native cube measure "contributors.COUNT":
cube.query(m["contributors.COUNT"], levels=[l["Reporting Date"]]).head(5)
# The number of contracts having more than 30 days past due:
# Below I'm creating various possibilities for the days past due threshold parameter. I will be able to choose them from a hierarchy:
PastDueDaysThresholds = cube.create_parameter_simulation(
"PastDueDaysThresholds",
measures={"PastDueDaysThreshold": 30.0},
base_scenario_name=">30 days",
)
PastDueDaysThresholds += (">60 days", 60.0)
PastDueDaysThresholds += (">90 days", 90.0)
# +
# Indicator - let's assume that a loan is classified as a past due due after 30 days of being "past due".
m["DaysPastDue"] = tt.value(risk_engine_data["DaysPastDue"])
# Number of contracts past due:
m["Num_Contracts_Past_due"] = tt.agg.sum(
tt.where(m["DaysPastDue"] > m["PastDueDaysThreshold"], 1.0, 0.0),
scope=tt.scope.origin(l["id"]),
)
m["% past due"] = m["Num_Contracts_Past_due"] / m["contributors.COUNT"]
m["% past due"].formatter = "DOUBLE[#,###.##%]"
# -
cube.query(m["% past due"], levels=[l["Reporting Date"]])
# # Annex I: LendingClub data viz
#
# In this section we will create measures visualizing the LendingClub data. I'm creating a second cube to aggregate lending_club_data - so that there's no linkage with "reporting dates" and no duplication of data.
#
# Please note, that I'm operating with a subset of data - to save up time on loading data from s3 (see input data section).
cube2 = session.create_cube(lending_club_data, "Lending Club EDA")
m2 = cube2.measures
# ## Loan amount
# The listed amount of the loan applied for by the borrower.
m2["loan_amount"] = tt.agg.sum(lending_club_data["loan_amnt"])
m2["loan_amount"].folder = "LendingClub"
# Average loan amount:
m2["loan_amount.MEAN"] = tt.agg.mean(lending_club_data["loan_amnt"])
m2["loan_amount.MEAN"].folder = "LendingClub"
# + atoti={"widget": {"mapping": {"columns": ["ALL_MEASURES"], "measures": ["[Measures].[loan_amnt.SUM]"], "rows": ["[Lending Club Data].[issue_d].[issue_d]"]}, "query": {"mdx": "SELECT NON EMPTY {[Measures].[loan_amnt.SUM]} ON COLUMNS, NON EMPTY Hierarchize(Descendants({[Lending Club Data].[issue_d].[AllMember]}, 1, SELF_AND_BEFORE)) ON ROWS FROM [Lending Club EDA]", "updateMode": "once"}, "serverKey": "default", "switchedTo": "plotly-stacked-column-chart", "widgetKey": "pivot-table"}}
session.visualize("Total loan amount")
# + atoti={"widget": {"mapping": {"horizontalSubplots": [], "splitBy": ["[Lending Club Data].[grade].[grade]", "ALL_MEASURES"], "values": ["[Measures].[loan_amnt.MEAN]"], "verticalSubplots": [], "xAxis": ["[Lending Club Data].[issue_d].[issue_d]"]}, "query": {"mdx": "SELECT NON EMPTY Crossjoin(Hierarchize(Descendants({[Lending Club Data].[grade].[AllMember]}, 1, SELF_AND_BEFORE)), {[Measures].[loan_amnt.MEAN]}) ON COLUMNS, NON EMPTY Hierarchize(Descendants({[Lending Club Data].[issue_d].[AllMember]}, 1, SELF_AND_BEFORE)) ON ROWS FROM [Lending Club EDA]", "updateMode": "once"}, "serverKey": "default", "widgetKey": "plotly-line-chart"}}
session.visualize("Average loan size by Credit Score")
# -
# ## Interest rate
# + atoti={"widget": {"mapping": {"horizontalSubplots": [], "splitBy": ["[Lending Club Data].[grade].[grade]", "ALL_MEASURES"], "values": ["[Measures].[int_rate.MEAN]"], "verticalSubplots": [], "xAxis": ["[Lending Club Data].[issue_d].[issue_d]"]}, "query": {"mdx": "SELECT NON EMPTY Crossjoin(Hierarchize(Descendants({[Lending Club Data].[grade].[AllMember]}, 1, SELF_AND_BEFORE)), {[Measures].[int_rate.MEAN]}) ON COLUMNS, NON EMPTY Hierarchize(Descendants({[Lending Club Data].[issue_d].[AllMember]}, 1, SELF_AND_BEFORE)) ON ROWS FROM [Lending Club EDA]", "updateMode": "once"}, "serverKey": "default", "widgetKey": "plotly-line-chart"}}
session.visualize("Interest rates by credit score")
# -
# ## Count loans
# + atoti={"widget": {"mapping": {"columns": ["ALL_MEASURES"], "measures": ["[Measures].[contributors.COUNT]"], "rows": ["[Lending Club Data].[loan_status].[loan_status]"]}, "query": {"mdx": "SELECT NON EMPTY {[Measures].[contributors.COUNT]} ON COLUMNS, NON EMPTY Hierarchize(Descendants({[Lending Club Data].[loan_status].[AllMember]}, 1, SELF_AND_BEFORE)) ON ROWS FROM [Lending Club EDA]", "updateMode": "once"}, "serverKey": "default", "switchedTo": "plotly-donut-chart", "widgetKey": "pivot-table"}}
session.visualize("Count loans")
# + atoti={"widget": {"mapping": {"horizontalSubplots": [], "stackBy": ["[Lending Club Data].[loan_status].[loan_status]", "ALL_MEASURES"], "values": ["[Measures].[contributors.COUNT]"], "verticalSubplots": [], "xAxis": ["[Lending Club Data].[purpose].[purpose]"]}, "query": {"mdx": "SELECT NON EMPTY Crossjoin(Hierarchize(Descendants({[Lending Club Data].[loan_status].[AllMember]}, 1, SELF_AND_BEFORE)), {[Measures].[contributors.COUNT]}) ON COLUMNS, NON EMPTY Hierarchize(Descendants({[Lending Club Data].[purpose].[AllMember]}, 1, SELF_AND_BEFORE)) ON ROWS FROM [Lending Club EDA]", "updateMode": "once"}, "serverKey": "default", "widgetKey": "plotly-100-stacked-column-chart"}}
session.visualize("Proportion of loans purpose and by status")
# -
session.link()
# <div style="text-align: center;" ><a href="https://www.atoti.io/?utm_source=gallery&utm_content=ifrs9" target="_blank" rel="noopener noreferrer"><img src="https://data.atoti.io/notebooks/banners/discover-try.png" alt="Try atoti"></a></div>
| notebooks/ifrs9/main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# %load_ext autoreload
# %autoreload 2
# +
import os
import igraph as ig
import arw
import utils
import random
import time
random.seed(time.time())
# -
# ### Load network dataset and extract ARW input data
# +
path = './datasets/acl.pkl'
network = ig.Graph.Read_Pickle(path)
print network.summary()
attr = 'single_attr' if network['attributed'] else None
input_data = utils.extract_arw_input_data(network, 'time', 0.00, 0.01, debug=False, attrs=attr)
# -
# ### Generate ARW graph with fitted parameters
# +
params = dict(p_diff=0.08, p_same=0.06, jump=0.42, out=1)
arw_graph = arw.RandomWalkSingleAttribute(params['p_diff'], params['p_same'],
params['jump'], params['out'],
input_data['gpre'], attr_name=attr)
arw_graph.add_nodes(input_data['chunk_sizes'], input_data['mean_outdegs'],
chunk_attr_sampler=input_data['chunk_sampler'] if attr else None)
arw_graphs[network] = arw_graph
# -
# ### Compare graph statistics
utils.plot_deg_and_cc_and_deg_cc([arw_graph.g, network], ['ARW', 'Dataset'], get_atty=network['attributed'])
| example.ipynb |
# ---
# title: "Rock Paper Scissor"
# author: "TACT"
# date: 2019-04-20
# description: "-"
# type: technical_note
# draft: false
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import random
chose = ["rock","paper","scissor"]
def play(youChose):
computerChose = random.choice(chose)
print('you:',youChose,',computer:',computerChose)
if youChose == computerChose:
return 'Draw'
elif youChose == 'rock' and computerChose == 'paper':
return "computer won"
elif youChose == 'rock' and computerChose == 'scissor':
return "you won"
elif youChose == 'paper' and computerChose == 'rock':
return 'you won'
elif youChose == 'paper' and computerChose == 'scissor':
return 'computer won'
elif youChose == 'scissor' and computerChose == 'rock':
return 'computer won'
elif youChose == 'scissor' and computerChose == 'paper':
return 'you won'
play('rock')
play('paper')
play('scissor')
play('scissor')
play('rock')
play('rock')
| docs/python/basics/rock-paper-scissor.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Image Classification using LeNet CNN
# ## CIFAR-10 Dataset - 10 classes of animals and objects
# import the necessary packages
from LeNet import LeNet
from sklearn.model_selection import train_test_split
from keras.datasets import cifar10
from keras.optimizers import RMSprop
from keras.utils import np_utils
from keras import backend as K
import numpy as np
import argparse
import cv2
# ## Load the data
# grab the CIFAR-10 dataset (may take time the first time)
print("[INFO] downloading CIFAR-10...")
((trainData, trainLabels), (testData, testLabels)) = cifar10.load_data()
# ## Prepare the data
# parameters for CIFAR-10 data set
num_classes = 10
image_width = 32
image_height = 32
image_channels = 3
# shape the input data using "channels last" ordering
# num_samples x rows x columns x depth
trainData = trainData.reshape(
(trainData.shape[0], image_height, image_width, image_channels))
testData = testData.reshape(
(testData.shape[0], image_height, image_width, image_channels))
# scale data to the range of [0.0, 1.0]
trainData = trainData.astype("float32") / 255.0
testData = testData.astype("float32") / 255.0
# transform the training and testing labels into vectors in the
# range [0, classes] -- this generates a vector for each label,
# where the index of the label is set to `1` and all other entries
# to `0`; in the case of CIFAR-10, there are 10 class labels
trainLabels = np_utils.to_categorical(trainLabels, num_classes) # one hot encoding
testLabels = np_utils.to_categorical(testLabels, num_classes)
# ## Train Model
# +
# initialize the model
print("[INFO] compiling model...")
model = LeNet.build(numChannels=image_channels,
imgRows=image_height, imgCols=image_width,
numClasses=num_classes,
weightsPath=None)
# initialize the optimizer
opt = RMSprop(lr=0.0001, decay=1e-6) # RMS Prop
# build the model
model.compile(loss="categorical_crossentropy", # Soft-Max
optimizer=opt, metrics=["accuracy"])
# +
# initialize hyper parameters
batch_size = 128
epochs = 1
print("[INFO] training...")
model.fit(trainData, trainLabels, batch_size=batch_size,
epochs=epochs, verbose=1)
# -
# show the accuracy on the testing set
print("[INFO] evaluating...")
(loss, accuracy) = model.evaluate(testData, testLabels,
batch_size=batch_size, verbose=1)
print("[INFO] accuracy: {:.2f}%".format(accuracy * 100))
model.save_weights("lenet_cifar_test.hdf5", overwrite=True)
# ## Evaluate Pre-trained Model
# +
# load the model weights
print("[INFO] compiling model...")
model = LeNet.build(numChannels=image_channels,
imgRows=image_height, imgCols=image_width,
numClasses=num_classes,
weightsPath="weights/lenet_weights_cifar.hdf5")
# initialize the optimizer
opt = RMSprop(lr=0.0001, decay=1e-6) # RMS Prop
# build the model
model.compile(loss="categorical_crossentropy", # Soft-Max
optimizer=opt, metrics=["accuracy"])
# -
# show the accuracy on the testing set
print("[INFO] evaluating...")
(loss, accuracy) = model.evaluate(testData, testLabels,
batch_size=batch_size, verbose=1)
print("[INFO] accuracy: {:.2f}%".format(accuracy * 100))
# ## Model Predictions
# +
# set prediction parameters
num_predictions = 10
# randomly select a few testing digits
for i in np.random.choice(np.arange(0, len(testLabels)), size=(num_predictions,)):
# classify the digit
probs = model.predict(testData[np.newaxis, i])
prediction = probs.argmax(axis=1)
# extract the image from the testData
chR = (testData[i][:, :, 0] * 255).astype("uint8")
chG = (testData[i][:, :, 1] * 255).astype("uint8")
chB = (testData[i][:, :, 2] * 255).astype("uint8")
# merge the channels into one image
image = cv2.merge((chB, chG, chR))
# resize the image from a 32 x 32 image to a 96 x 96 image so we can better see it
image = cv2.resize(image, (96, 96), interpolation=cv2.INTER_LINEAR)
print("[INFO] Predicted: {}, Actual: {}".format(
prediction[0], np.argmax(testLabels[i])))
# show the image and prediction
classLabels = ['airplane', 'automobile', 'bird', 'cat',
'deer', 'dog', 'frog', 'horse', 'ship', 'truck']
cv2.putText(image, classLabels[prediction[0]], (5, 20),
cv2.FONT_HERSHEY_SIMPLEX, 0.75, (0, 255, 0), 2)
cv2.imshow("Object", image)
cv2.waitKey(0)
# close the display window
cv2.destroyAllWindows()
# -
| ImageClassification/LeNet_CIFAR.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a href="https://githubtocolab.com/giswqs/geemap/blob/master/examples/notebooks/14_legends.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab"/></a>
#
# Uncomment the following line to install [geemap](https://geemap.org) if needed.
# +
# # !pip install geemap
# -
import ee
import geemap
geemap.show_youtube('NwnW_qOkNRw')
# ## Add builtin legends from geemap Python package
#
# https://github.com/giswqs/geemap/blob/master/geemap/legends.py
#
# ### Available builtin legends:
legends = geemap.builtin_legends
for legend in legends:
print(legend)
# ### Available Land Cover Datasets in Earth Engine
#
# https://developers.google.com/earth-engine/datasets/tags/landcover
# ### National Land Cover Database (NLCD)
#
# https://developers.google.com/earth-engine/datasets/catalog/USGS_NLCD_RELEASES_2019_REL_NLCD
Map = geemap.Map()
Map.add_basemap('HYBRID')
landcover = ee.Image('USGS/NLCD_RELEASES/2019_REL/NLCD/2019').select('landcover')
Map.addLayer(landcover, {}, 'NLCD Land Cover')
Map.add_legend(builtin_legend='NLCD')
Map
# ### National Wetlands Inventory (NWI)
#
# https://www.fws.gov/wetlands/data/mapper.html
Map = geemap.Map()
Map.add_basemap('HYBRID')
Map.add_basemap('FWS NWI Wetlands')
Map.add_legend(builtin_legend='NWI')
Map
# ### MODIS Land Cover Type Yearly Global 500m
#
# https://developers.google.com/earth-engine/datasets/catalog/MODIS_051_MCD12Q1
# +
Map = geemap.Map()
Map.add_basemap('HYBRID')
landcover = ee.Image('MODIS/051/MCD12Q1/2013_01_01').select('Land_Cover_Type_1')
Map.setCenter(6.746, 46.529, 2)
Map.addLayer(landcover, {}, 'MODIS Land Cover')
Map.add_legend(builtin_legend='MODIS/051/MCD12Q1')
Map
# -
# ## Add customized legends for Earth Engine data
#
# There are three ways you can add customized legends for Earth Engine data
#
# 1. Define legend keys and colors
# 2. Define legend dictionary
# 3. Convert Earth Engine class table to legend dictionary
#
# ### Define legend keys and colors
# +
Map = geemap.Map()
legend_keys = ['One', 'Two', 'Three', 'Four', 'ect']
# colorS can be defined using either hex code or RGB (0-255, 0-255, 0-255)
legend_colors = ['#8DD3C7', '#FFFFB3', '#BEBADA', '#FB8072', '#80B1D3']
# legend_colors = [(255, 0, 0), (127, 255, 0), (127, 18, 25), (36, 70, 180), (96, 68 123)]
Map.add_legend(
legend_keys=legend_keys, legend_colors=legend_colors, position='bottomleft'
)
Map
# -
# ### Define a legend dictionary
# +
Map = geemap.Map()
legend_dict = {
'11 Open Water': '466b9f',
'12 Perennial Ice/Snow': 'd1def8',
'21 Developed, Open Space': 'dec5c5',
'22 Developed, Low Intensity': 'd99282',
'23 Developed, Medium Intensity': 'eb0000',
'24 Developed High Intensity': 'ab0000',
'31 Barren Land (Rock/Sand/Clay)': 'b3ac9f',
'41 Deciduous Forest': '68ab5f',
'42 Evergreen Forest': '1c5f2c',
'43 Mixed Forest': 'b5c58f',
'51 Dwarf Scrub': 'af963c',
'52 Shrub/Scrub': 'ccb879',
'71 Grassland/Herbaceous': 'dfdfc2',
'72 Sedge/Herbaceous': 'd1d182',
'73 Lichens': 'a3cc51',
'74 Moss': '82ba9e',
'81 Pasture/Hay': 'dcd939',
'82 Cultivated Crops': 'ab6c28',
'90 Woody Wetlands': 'b8d9eb',
'95 Emergent Herbaceous Wetlands': '6c9fb8',
}
landcover = ee.Image('USGS/NLCD/NLCD2016').select('landcover')
Map.addLayer(landcover, {}, 'NLCD Land Cover')
Map.add_legend(legend_title="NLCD Land Cover Classification", legend_dict=legend_dict)
Map
# -
# ### Convert an Earth Engine class table to legend
#
# For example: MCD12Q1.051 Land Cover Type Yearly Global 500m
#
# https://developers.google.com/earth-engine/datasets/catalog/MODIS_051_MCD12Q1
# +
Map = geemap.Map()
ee_class_table = """
Value Color Description
0 1c0dff Water
1 05450a Evergreen needleleaf forest
2 086a10 Evergreen broadleaf forest
3 54a708 Deciduous needleleaf forest
4 78d203 Deciduous broadleaf forest
5 009900 Mixed forest
6 c6b044 Closed shrublands
7 dcd159 Open shrublands
8 dade48 Woody savannas
9 fbff13 Savannas
10 b6ff05 Grasslands
11 27ff87 Permanent wetlands
12 c24f44 Croplands
13 a5a5a5 Urban and built-up
14 ff6d4c Cropland/natural vegetation mosaic
15 69fff8 Snow and ice
16 f9ffa4 Barren or sparsely vegetated
254 ffffff Unclassified
"""
landcover = ee.Image('MODIS/051/MCD12Q1/2013_01_01').select('Land_Cover_Type_1')
Map.setCenter(6.746, 46.529, 2)
Map.addLayer(landcover, {}, 'MODIS Land Cover')
legend_dict = geemap.legend_from_ee(ee_class_table)
Map.add_legend(legend_title="MODIS Global Land Cover", legend_dict=legend_dict)
Map
| examples/notebooks/14_legends.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="img/hp-logo-3.png" width="70">
#
# # HP Prime Virtual Calculator Emulator
# Emuladores de la calculadora HP Prime
# __Enlaces de descarga del emulador de la calculadora:__
# [>> HP Prime Virtual Calculator Emulator for Windows (32-bit) <<](https://www.hpcalc.org/details/7468)
# [>> HP Prime Virtual Calculator Emulator for Windows (64-bit) <<](https://www.hpcalc.org/details/8939)
# [>> HP Prime Virtual Calculator for MacOS <<](https://www.hpcalc.org/details/7799)
# [>> HP Prime Virtual Calculator for Android <<](https://mega.nz/file/kxB3xC7R#n86mVIVgNltHru1pn0qIAYmXMvClj3i83xiEaE_DdEY)
# __Nota:__ las versiones windows de la calculadora, también se pueden instalar y ejecutar en Linux mediante Wine / PlayOnLinux.
# ## Video promocional de la calculadora
from IPython.display import YouTubeVideo
YouTubeVideo('WF8tZP0uKu0', width=800, height=450)
# ## Manual de la calculadora:
from IPython.display import HTML
HTML('<iframe src=pdf/User_Guide_ESP_2018_01_12_1.pdf width=90% height=500</iframe>')
# ## Video introducción
YouTubeVideo('DFnlOq4iH4E', width=800, height=450)
# ## Imágenes
# <img src="img/c04427073_1750x1285.jpg">
# <img src="img/c03708410_1750x1285.jpg">
| HP_Prime.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Multi-options model
from prayas import *
# The experiment consists of multiple variants and in each variant the visitor has one ore more options to choose. A detailed explanation of the methodology is available in *[Bayesian A/B Testing for Business Decisions](https://arxiv.org/abs/2003.02769)* by <NAME> and <NAME> (2020).
# In this example, the experiment consists of two variants with each variant having `9` different options from which the visitor can choose:
m = MultiOptionsModel(variants=["Original", "Progressive"],
options=[9, 9],
baseline="Original")
# In addition to the *conversion* measure, we are also interested in measuring the *revenue* and the *gain* both in Euro:
# +
rev_a = [27.95, 47.95, 63.95,
35.95, 63.95, 79.95,
79.95, 151.95, 223.95]
rev_b = [34.95, 59.95, 79.95,
37.95, 67.95, 84.95,
69.95, 132.95, 195.95]
m.add_measure("revenue",
success_value=[rev_a, rev_b])
m.add_measure("gain",
success_value=[rev_a, rev_b],
nonsuccess_value=[np.repeat(-0.06, 9),
np.repeat(-0.04, 9)])
# -
# The full model specification for this experiment is:
print(m)
# Set the result of the experiment:
m.set_result(successes=[[50, 5, 5, 28, 7, 5, 20, 1, 6],
[28, 3, 6, 30, 6, 5, 27, 6, 3]],
trials=[8067, 8082])
# Investigate the result:
m.plot();
# The plot provides the posteriors of the measures 'conversion rate', 'revenue', and 'gain'. We can already see that for different measures the location of the posteriors to each other is different.
#
# Get details on the result:
m.score_baseline()
# The progressive variant has lower conversion rates than the original variant (-10% on average) but the revenue per visitor is almost the same for both variants since the probability to be best is almost 50% for both variants; both variants are equally likely to be the best. The progressive variant has less conversions for the lower priced option compared to the original, but makes up for the lost revenue with higher conversions on the premium product. For the metric gain per visitor, progressive discount had the highest probability to be best with a low expected loss of 2%.
| docs/notebooks/02-multi-options-model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import nltk
# +
#nltk.download_shell()
# -
message = [line.rstrip() for line in open('SMSSpamCollection')]
len(message)
import pandas as pd
import numpy as np
messages = pd.read_csv('SMSSpamCollection', sep='\t', names=['label', 'message'])
messages.head()
messages['length']= messages['message'].apply(len)
messages.head()
import seaborn as sns
messages['length'].hist()
# # part two (2)
import string
mess = 'simple message! notice: it has punctuations.'
nopunc1 = [c for c in mess if c not in string.punctuation]
nopunc2 = ''.join(nopunc1)
nopunc2
from nltk.corpus import stopwords
stopwords.words('english')
nopunc = ''.join(nopunc)
nopunc
x = ['acdd', 'b','c','d','e']
' '.join(x)
nopunc.split()
clean_mess = [word for word in nopunc.split() if word.lower() not in stopwords.words('english')]
clean_mess
# +
#checking for stopwords.word('english')
nopunc2.split()
# -
clean_msg =[word for word in nopunc2.split() if word.lower() not in stopwords.words('english')]
clean_msg
# # tokenisation
def text_process(mess):
"""
1. remove punc
2. remove stop words
3. return list of clean text words
"""
# ***
nopunc = [char for char in mess if char not in string.punctuation]
nopunc = ''.join(nopunc)
return [word for word in nopunc.split() if word.lower() not in stopwords.words('english')]
# messages.head()
messages['message'].head(5).apply(text_process)
messages['message'].head(5).apply(text_process)
# # next step is vectorization
from sklearn.feature_extraction.text import CountVectorizer
bow_transformer = CountVectorizer(analyzer = text_process)
#.fit(messages['message'])
bow_transformer.fit(messages['message'])
print(len(bow_transformer.vocabulary_))
mess4 = messages['message'][3]
print(mess4)
bow4 = bow_transformer.transform([mess4])
print(bow4)
bow_transformer.get_feature_names()[9554]
bow_transformer.get_feature_names()[4068]
# # PART 3
#transforming the messages
message_bow = bow_transformer.transform(messages['message'])
print('shape of Sparse Matrix: ', message_bow.shape)
message_bow.nnz
sparsity = (100.0*message_bow.nnz / (message_bow.shape[0] * message_bow.shape[1]))
print('sparsity: {} '.format((sparsity)))
# # TF-IDF : term frequency-inverse document frequency
from sklearn.feature_extraction.text import TfidfTransformer
tfidf_transformer = TfidfTransformer().fit(message_bow)
tfidf4 = tfidf_transformer.transform(bow4)
print(tfidf4)
tfidf_transformer.idf_[bow_transformer.vocabulary_['university']]
from sklearn.feature_extraction.text import TfidfTransformer
tfidf_transform = TfidfTransformer().fit(message_bow)
tfidf44 = tfidf_transform.transform(bow4)
print(tfidf44)
print(tfidf_transform.idf_[bow_transformer.vocabulary_['u']])
print(tfidf_transform.idf_[bow_transformer.vocabulary_['university']])
# # training of our model
# +
#transforming the entire message
#in to erm frequency-inverse document frequency
message_tfidf = tfidf_transformer.transform(message_bow)
# -
print(message_tfidf.shape)
from sklearn.naive_bayes import MultinomialNB
spam_detect_model = MultinomialNB().fit(message_tfidf, messages['label'])
#testing predicting
spam_detect_model.predict(tfidf44)[0]
messages['label'][3]
message_tfidf
all_pred = spam_detect_model.predict(message_tfidf)
all_pred
# # using train_test_split(
from sklearn.model_selection import train_test_split
msg_train, msg_test, label_train, label_test = train_test_split(messages['message'], messages['label'], test_size =0.3)
# +
#msg_test
# -
# # using pipeline to summarise all the steps
from sklearn.pipeline import Pipeline
#pipeline helps us to shorten all previous steps
pipeline = Pipeline([
('bow', CountVectorizer(analyzer = text_process)),
('tfidf', TfidfTransformer()),
('classifier', MultinomialNB())
])
pipeline.fit(msg_train, label_train)
# +
#predictions = pipeline.predict(msg_test)
# +
#pipeline.predict(msg_test)
# -
len(msg_test)
| 02 Introduction to NLP 2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:analytics] *
# language: python
# name: conda-env-analytics-py
# ---
# ### Problem Statment
# Use Netflix Movies and TV Shows dataset from Kaggle and perform following operations :
# - Make a visualization showing the total number of movies watched by children
# - Make a visualization showing the total number of standup comedies
# - Make a visualization showing most watched shows.
# - Make a visualization showing highest rated show
# - Make a dashboard containing all of these above visualizations.
# %config Completer.use_jedi = False
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from wordcloud import WordCloud
import re
# -
df = pd.read_csv('netflix_titles.csv')
df.head()
df.info()
df.isnull().sum()
df.drop(index=df[df['date_added'].isnull()].index,inplace=True)
df.drop(index=df[df['rating'].isnull()].index,inplace=True)
df.drop(columns=['director','cast'],inplace=True)
df['country'] = df['country'].fillna(value=df['country'].mode()[0])
df.isnull().sum()
df['year_added'] = df['date_added'].apply(lambda x: x.split(' ')[-1])
df['year_added'] = pd.to_numeric(df['year_added'])
df['type'].value_counts()
df['rating'].value_counts()
target_audience = {
'TV-PG': 'Kids',
'TV-MA': 'Adults',
'TV-Y7-FV': 'Kids',
'TV-Y7': 'Kids',
'TV-14': 'Teens',
'R': 'Adults',
'TV-Y': 'Kids',
'NR': 'Adults',
'PG-13': 'Teens',
'TV-G': 'Kids',
'PG': 'Kids',
'G': 'Kids',
'UR': 'Adults',
'NC-17': 'Adults'
}
df['target_audience'] = df['rating'].apply(lambda x: target_audience.get(x))
df['target_audience'].head()
df['target_audience'].value_counts()
df['origin_country'] = df['country'].apply(lambda x: x.split(',')[0])
df['origin_country'].unique()
df.rename(columns={'listed_in':'genres'},inplace=True)
df['genres'].head(10)
df['genres'] = df['genres'].apply(lambda x: re.split(', | ,| & |,',x))
df['genres'].head(10)
df.info()
plt.figure(figsize=(5,5))
plt.pie(df['target_audience'].value_counts(),labels=['Adults','Teens','Kids'],colors=['#fcbe86','#95cf95','#8ebad9'],autopct='%.2f')
print(df['target_audience'].value_counts())
plt.figure(figsize=(5,5))
plt.pie(df[df['type']=='Movie']['target_audience'].value_counts(),labels=['Adults','Teens','Kids'],colors=['#fcbe86','#95cf95','#8ebad9'],autopct='%.2f')
print(df[df['type']=='Movie']['target_audience'].value_counts())
plt.title('Movies Audience Distribution')
plt.figure(figsize=(5,5))
plt.pie(df[df['type']=='TV Show']['target_audience'].value_counts(),labels=['Adults','Teens','Kids'],colors=['#fcbe86','#95cf95','#8ebad9'],autopct='%.2f')
print(df[df['type']=='TV Show']['target_audience'].value_counts())
plt.title('TV Shows Audience Distribution')
plt.figure(figsize=(15,7))
sns.histplot(data=df, x='rating',hue='target_audience',alpha=0.5)
plt.title('Content Ratings Histogram')
plt.figure(figsize=(16,8))
sns.scatterplot(x=df['release_year'],y=df['year_added'],hue=df['type'])
plt.title('Content Production')
plt.grid('on')
plt.figure(figsize=(5,5))
plt.pie(df['type'].value_counts(),labels=['Movies','TV Shows'],colors=['#fcbe86','#8ebad9'],autopct='%.2f')
print(df['type'].value_counts())
# +
movie_genres_dict = {}
for i in df[df['type']=='Movie'].index:
genres_list = df.loc[i]['genres']
for genre in genres_list:
genre=genre.lower()
if movie_genres_dict.get(genre):
movie_genres_dict[genre] = movie_genres_dict.get(genre) + 1
else:
movie_genres_dict[genre] = 1
movie_genres_df = pd.DataFrame(data={'genre':movie_genres_dict.keys(),'count':movie_genres_dict.values()})
movie_genres_df.head()
# -
plt.figure(figsize=(10,10))
sns.barplot(data=movie_genres_df.sort_values(by='count',ascending=False),x='count',y='genre')
plt.title('Movie Genres')
# +
tv_genres_dict = {}
for i in df[df['type']=='TV Show'].index:
genres_list = df.loc[i]['genres']
for genre in genres_list:
genre=genre.lower()
if tv_genres_dict.get(genre):
tv_genres_dict[genre] = tv_genres_dict.get(genre) + 1
else:
tv_genres_dict[genre] = 1
tv_genres_df = pd.DataFrame(data={'genre':tv_genres_dict.keys(),'count':tv_genres_dict.values()})
tv_genres_df.head()
# -
plt.figure(figsize=(10,10))
sns.barplot(data=tv_genres_df.sort_values(by='count',ascending=False),x='count',y='genre')
plt.title('TV Show Genres')
# +
text = ""
for i in df.index:
genres_list = df.loc[i]['genres']
for genre in genres_list:
genre=genre.lower()
text += genre + ' '
wordcloud = WordCloud(background_color='white', width=1680, height=1680, max_words=150).generate(text)
plt.figure(figsize=(16,16))
plt.imshow(wordcloud)
plt.axis('off')
| A06/notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# # Single effect model sanity check
#
# Check for agreement of different regression methods in VEM updates.
library(mvsusieR)
set.seed(2)
attach(mvsusie_sim1(r=1))
data = mvsusieR:::DenseData$new(X,y)
head(data$XtY)
head(data$XtR)
# ## Univariate single effect regression
prior_var = 0.2 * as.numeric(var(y))
residual_var = as.numeric(var(y))
m1 = mvsusieR:::SingleEffectModel(mvsusieR:::BayesianSimpleRegression)$new(ncol(X), residual_var, prior_var)
# ## Multivariate single effect regression
prior_covar = mvsusieR:::MashInitializer$new(list(0.2 * cov(y)), 1, alpha=0)
residual_covar = cov(y)
m2 = mvsusieR:::SingleEffectModel(mvsusieR:::MashRegression)$new(ncol(X), residual_covar, prior_covar)
# ## Predictions after one fit
pred01 = m1$predict(data)
m1$fit(data)
pred1 = m1$predict(data)
pred02 = m2$predict(data)
m2$fit(data)
pred2 = m2$predict(data)
head(pred1)
head(pred2)
head(pred01)
head(pred02)
head(m1$pip)
head(m2$pip)
head(m1$posterior_b1 / m1$pip)
head(m2$posterior_b1 / m2$pip)
m1$lbf
m2$lbf
data$add_to_fitted(pred1)
data$compute_residual()
head(data$XtY)
head(data$XtR)
# ## Predictions after two fits
m1$fit(data)
pred1 = m1$predict(data)
m2$fit(data)
pred2 = m2$predict(data)
head(pred1)
head(pred2)
head(m1$pip)
head(m2$pip)
head(m1$posterior_b1)
head(m2$posterior_b1)
m1$lbf
m2$lbf
data$add_to_fitted(pred1)
data$compute_residual()
head(data$XtR)
# ## Predictions after three fits
m1$fit(data)
pred1 = m1$predict(data)
m2$fit(data)
pred2 = m2$predict(data)
head(m1$posterior_b1)
head(m2$posterior_b1)
m1$lbf
m2$lbf
# ## Check on main interface call
L = 10
residual_var = as.numeric(var(y))
scaled_prior_var = V[1,1] / residual_var
A = susie(X,y,L=L,prior_variance=scaled_prior_var,
compute_objective=FALSE)
residual_cov = cov(y)
m_init = mvsusieR:::MashInitializer$new(list(V), 1, alpha = 0)
B = susie(X,y,L=L,prior_variance=m_init,compute_objective=FALSE)
max(abs(A$lbf - B$lbf))
| inst/prototypes/univariate_algorithm_agreement.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#Import modules and libraries
import pandas as pd
from glob import glob
import datetime
stock_files =sorted(glob('data/JC-*-citibike-tripdata.csv'))
stock_files
df=pd.concat((pd.read_csv(file)
for file in stock_files), ignore_index = True)
df.dtypes
df['starttime']=pd.to_datetime(df['starttime'])
df['Age']=(df['starttime'].dt.year)-df['birth year']
df.head()
df.to_csv("JC-201801-202104-citibike-tripdata.csv")
# +
#age_df=df.loc[:,['starttime','birth year','Age']]
# +
#age_df=age_df.loc[df['Age']>100]
# +
#age_df['birth year'].value_counts()
# +
#age_df=age_df.set_index('Age')
#age_df
# -
| join_citibike_csv_files.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tutorial 4: Trajectory aggregation (flow maps)
#
# This tutorial covers trajectory generalization and aggregation using flow maps.
# %matplotlib inline
# +
import urllib
import os
import numpy as np
import pandas as pd
from geopandas import GeoDataFrame, read_file
from shapely.geometry import Point, LineString, Polygon, MultiPoint
from datetime import datetime, timedelta
import sys
sys.path.append("..")
import movingpandas as mpd
import warnings
warnings.simplefilter("ignore")
# -
# ## Ship movements (AIS data)
df = read_file('data/demodata_ais.gpkg')
df['t'] = pd.to_datetime(df['Timestamp'], format='%d/%m/%Y %H:%M:%S')
df = df.set_index('t')
df = df[df.SOG>0]
df.size
MIN_LENGTH = 100 # meters
traj_collection = mpd.TrajectoryCollection(df, 'MMSI', min_length=MIN_LENGTH)
print("Finished creating {} trajectories".format(len(traj_collection)))
trips = traj_collection.split_by_observation_gap(timedelta(minutes=5))
print("Extracted {} individual trips from {} continuous vessel tracks".format(len(trips), len(traj_collection)))
# Generalizing the trip trajectories significantly speeds up the following aggregation step.
# %%time
generalized = trips.generalize(mode='min-distance', tolerance=100)
#generalized = trips.generalize(mode='min-time-delta', tolerance=timedelta(minutes=1))
#generalized = trips.generalize(mode='douglas-peucker', tolerance=100)
# %%time
aggregator = mpd.TrajectoryCollectionAggregator(generalized, max_distance=1000, min_distance=100, min_stop_duration=timedelta(minutes=5))
pts = aggregator.get_significant_points_gdf()
clusters = aggregator.get_clusters_gdf()
( pts.hvplot(geo=True, tiles='OSM', frame_width=800) *
clusters.hvplot(geo=True, color='red') )
flows = aggregator.get_flows_gdf()
( trips.hvplot(color='gray') *
flows.hvplot(geo=True, hover_cols=['weight'], line_width='weight', alpha=0.5, color='#1f77b3') *
clusters.hvplot(geo=True, color='red', size='n') )
# ### Comparison of generalized vs. original trajectories
# %%time
aggregator_original = mpd.TrajectoryCollectionAggregator(trips, max_distance=1000, min_distance=100, min_stop_duration=timedelta(minutes=5))
( aggregator_original.get_flows_gdf().hvplot(title='Original', geo=True, tiles='OSM', hover_cols=['weight'], line_width='weight', alpha=0.5, color='#1f77b3', frame_height=400, frame_width=400) *
aggregator_original.get_clusters_gdf().hvplot(geo=True, color='red', size='n') +
flows.hvplot(title='Generalized', geo=True, tiles='OSM', hover_cols=['weight'], line_width='weight', alpha=0.5, color='#1f77b3', frame_height=400, frame_width=400) *
clusters.hvplot(geo=True, color='red', size='n')
)
# ## Bird migration data
df = read_file('data/demodata_gulls.gpkg')
df['t'] = pd.to_datetime(df['timestamp'])
df = df.set_index('t')
df.size
traj_collection = mpd.TrajectoryCollection(df, 'individual-local-identifier', min_length=MIN_LENGTH)
print("Finished creating {} trajectories".format(len(traj_collection)))
trips = traj_collection.split_by_date(mode='month')
print("Extracted {} individual trips from {} continuous tracks".format(len(trips), len(traj_collection)))
generalized = trips.generalize(mode='min-time-delta', tolerance=timedelta(days=1))
# %%time
aggregator = mpd.TrajectoryCollectionAggregator(generalized, max_distance=1000000, min_distance=100000, min_stop_duration=timedelta(minutes=5))
# +
flows = aggregator.get_flows_gdf()
clusters = aggregator.get_clusters_gdf()
( flows.hvplot(geo=True, hover_cols=['weight'], line_width='weight', alpha=0.5, color='#1f77b3', tiles='OSM') *
clusters.hvplot(geo=True, color='red', size='n') )
# -
# %%time
aggregator_original = mpd.TrajectoryCollectionAggregator(trips, max_distance=1000000, min_distance=100000, min_stop_duration=timedelta(minutes=5))
( aggregator_original.get_flows_gdf().hvplot(title='Original', geo=True, tiles='OSM', hover_cols=['weight'], line_width='weight', alpha=0.5, color='#1f77b3', frame_height=600, frame_width=400) *
aggregator_original.get_clusters_gdf().hvplot(geo=True, color='red', size='n') +
flows.hvplot(title='Generalized', geo=True, tiles='OSM', hover_cols=['weight'], line_width='weight', alpha=0.5, color='#1f77b3', frame_height=600, frame_width=400) *
clusters.hvplot(geo=True, color='red', size='n')
)
| tutorials/4_generalization_and_aggregation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.2 64-bit ('.tgy_veribilimi')
# metadata:
# interpreter:
# hash: ba55969ab8b665a0d7ebc430170b2908b5d222710046023b08b5a17088334349
# name: Python 3.8.2 64-bit ('.tgy_veribilimi')
# ---
# # LOGISTIK REGRESYON
import numpy as np
import pandas as pd
import statsmodels.api as sm
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn.preprocessing import scale, StandardScaler
from sklearn.model_selection import train_test_split, GridSearchCV, cross_val_score
from sklearn.metrics import confusion_matrix, accuracy_score, mean_squared_error, r2_score, roc_auc_score, roc_curve, classification_report
from sklearn.linear_model import LogisticRegression
import warnings
warnings.filterwarnings("ignore", category=DeprecationWarning)
warnings.filterwarnings("ignore", category=FutureWarning)
# ## Veri Seti Hikayesi ve Problem : Şeker Hastalığı Tahmini
df = pd.read_csv("verisetleri\diabetes.csv")
df.head()
# +
# Hasta bilgileri ve tahlil bilgileri üzerinden tahmin yapılacak.
# +
# Preganacies-> hamilelik sayısı, Glucose-> gilikoz, BlookPressure-> kan basıncı, BMI-> vücut kitle endesi
# Outcome -> şekir hastalığını ifade eder
# -
# ## Lojistic Regresyon
# ## Model & Tahmin
# +
# TEST VE TRAIN OLARAK AYRIM YAPILMADIĞINA DİKKAT EDİNİZ. SONRAKİ BÖLÜMLERDE YAPILACAKTIR.
# -
df["Outcome"].value_counts()
df.describe().T
# Hamilelik max 17 aykırı gözlem olabilir. ( Diğer keşifçi veri analizine ilişkin bazı değerlendirmeler yapılmalıdır. Ve veri önişlemle adımları uygulanmalıdır. )
y = df["Outcome"]
X = df.drop(["Outcome"], axis=1)
y.head()
X.head()
# "liblinear" --> Lojistik regresyonda katsayıları bulabilmek adına kullanılan birden fazla minimizasyon yaklaşımı var. Yani gerçek değerler ile tahmin edilen değerler arasındaki farkların karelerinin toplamını ifade etmek adına bu sefer loglost gibi veya klasik regresyondan alışık olduğumuz yöntemler gibi bazı birbirinden farklı yöntemler var. Hatta ritch ve lasso dan alışık olduğumuz düzenlileştirme yöntemlerinin de işin içine katıldığı bazı katsayı bulma yöntemleri var. Bu sebeple bunu liblinear olarak ifade ettik.
loj_model = LogisticRegression(solver="liblinear").fit(X, y)
# B0 katsayısı
loj_model.intercept_
# Bağımısız değişkenlerin katsayıları
loj_model.coef_
# ### TAHMİN İŞLEMLERİ
# Tahmin edilen değerler
loj_model.predict(X)[0:10]
# Gerçek değerler
y[0:10]
# Başarısının değerlendirilmesi
y_pred = loj_model.predict(X)
# Karmaşıklık matrisi
confusion_matrix(y, y_pred)
# Karmaşıklık matrisinin değerlerini yorumlamak kolay olmayacağızndan accuracy_score kullanacağız.
# accuracy_score() --> doğruluk oranı, (başarılı işler) / (tüm durum)
accuracy_score(y, y_pred)
# ### Detaylı sınıflandırma raporu
# DAHA DETAYLI BİR SINIFLANDIRMA RAPORU ALMAK İÇİN classification_report KULLANILIR
print(classification_report(y, y_pred))
# ### Olasılık Değerlerinin Bulunması
# Bazı durmlarda sonuçları 1-bir ve 0-sıfır olarak değilde, direk olasılık değerlerinin kendisinden almak istersek - ki logistik fonksiyonu bize birinci sınıfın yani ilgilenmiş oluduğumuz 1-bir sınıfının gerçekleşme olasılığını veriyordu. - predict_proba() fonksiyonunu kullanırız.
loj_model.predict_proba(X)[0:10]
# +
# predict() fonksiyonu arka planda bu değerleri 0-sıfır ve 1-dönüştürmektedir.
# Bu sonuçlarda ilgileneceğimiz yani 1-bir ve 0-sıfır değerlerinin gösterildiği bölüm ikinci sutün dur.
# -
# ### ROC eğrisinin oluşturulması
# +
# ROC EGRİSİ : False positive rate ve true positive rate ler üzerinden oluşturulan bir grafik aracılığı ile bize bilgi sunan bir değerdir. Eğri altındaki alan ile ilgileniyorduk. Bu ne kadar büyük ise modelimiz o kadar başarılı şeklinde yorumlar yapabiliriz.
# -
logit_roc_auc = roc_auc_score(y, loj_model.predict(X))
fpr, tpr, thresholds = roc_curve(y, loj_model.predict_proba(X)[:,1])
plt.figure()
plt.plot(fpr, tpr, label='AUC (area=%0.2f)' % logit_roc_auc)
plt.plot([0, 1], [0, 1], 'r--')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Receiver operating characteristic')
plt.legend(loc="lower right")
plt.savefig('Log_ROC')
plt.show()
# ### MODEL TUNNING ( MODEL VALIDATION - MODEL DOĞRULAMA )
# +
# Regression bölümünde olduğu gibi bir model tunning işlemi ele alınmayacaktır. Model doğrulama, model validation işlemi yapılacaktır. Konu bütünlüğü olması açısından model tunning şeklinde isimlendirdik.
# +
# Fakat yinede logjistik regresyon yöntemlerinde tune edilmesi gereken hipeparametreler bulunmaktadır. Örneğin ritch, lasso ve elasticnet ten alışık olduğumuz l1 normu yani bir düzenlileştirme katsayısı modelde bulunduğunda buna yönelik olarak bir lojistik regresyon kullanıldığında parametre tahminleri için bu durumda aslına bakarsanız optimize edilmesi gereken bir hiperparametre elimizde olmuş oluyor. Fakat bizim kullanmış olduğumuz linear lojistic regresyon modeli klasik lojistik regresyon modeli bu sebeple extra bir hiperparametremiz yok.
# -
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.30, random_state=42)
loj_model = LogisticRegression(solver="liblinear").fit(X_train, y_train)
y_pred = loj_model.predict(X_test)
print(accuracy_score(y_test, y_pred))
cross_val_score(loj_model, X_test, y_test)
# +
# Birden fazla hata hesaplama işlemi gerçekleştirmektedir. Biz bunu dilediğimiz katlı örneğin 10 katlı olacak şekilde gerçekleştirebiliriz. Öntanımlı değeri itibariyle 5 tane gelmiş oldu.
# -
cross_val_score(loj_model, X_test, y_test, cv=10)
# +
# mean ile bunların ortalamasını alabiliriz. Böylece test setine ilişkin daha doğru bir test hatası değerine erişmiş oluruz.
# -
cross_val_score(loj_model, X_test, y_test, cv=10).mean()
# +
# SONRAKİ UYGULAMALARDA SADECE accuracy_score() DEĞERLENDİRMESİ YAPACAĞIZ. Hiper parametrelerin optimum değerlerine ulaşmak için de cross validation yöntemini kullanmış olacağız.
# Yani modelleri train seti ile eğitirken cross validation yöntemini kullanacağız. Ve dışarda bıraktığımız test setini performansı test etmek için kullanmış olacağız.
| 8 4 1 LOJISTIK REGRESYON.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Taxonomic analysis
#
# Preambule to Multiple Sequence alignment and Phylogenetic tree building.
# ### Import necessary modules
# +
from Bio import (
SeqIO as seqio,
SearchIO as searchio,
Entrez as entrez
)
from Bio.Seq import Seq as seq
import toml
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from functools import partial
from annotathon.utils.customobjs import (
Path as path,
objdict as odict
)
from annotathon.parsing.blast import parse_alignment_descriptions as parse_ncbi
from annotathon.annotation.helper_functions import *
# -
# ### Configuration to access NCBI's servers :
# Load configuration to access NCBI :
with open("../creds/entrezpy.toml", "r") as f:
ncbi = toml.load(f, _dict=odict)
# set credentials :
entrez.api_key = ncbi.credentials.apikey
entrez.email = ncbi.credentials.email
entrez.tool = ncbi.credentials.tool
# set plotting params :
# %matplotlib inline
plt.style.use('seaborn')
plt.rcParams['figure.figsize'] = (15, 8)
with open("../config/locations.toml", "r") as f:
_config = toml.load(f, _dict=odict)
_config
locations = odict({
key: path(value) for key, value in _config.locations.items()
})
locations
blast = odict({
"locations": locations,
"data": odict({})
})
blast
description_glob = "*Alignment-Descriptions*"
# ### Load blast alignment descriptions
blast.data.update(odict({
"default": parse_ncbi(blast.locations.default.lglob(description_glob)[0]),
"cinqk": parse_ncbi(blast.locations.cinqk.lglob(description_glob)[0]),
"landmark": parse_ncbi(blast.locations.landmark.lglob(description_glob)[0]),
"sp": parse_ncbi(blast.locations.sp.lglob(description_glob)[0]),
"taxo": parse_ncbi(blast.locations.anthony.lglob(description_glob)[0]),
"hypo": parse_ncbi(blast.locations.anthony.lglob("*.csv")[1])
}))
# This is probably unnecessary given that we now have all the information from the genbank !
#blast.data.taxo.loc[:, "Description"] = add_function(blast.data.taxo.Description)
blast.data.taxo.loc[:, "function"] = add_function(blast.data.taxo.Description)
blast.data.taxo.loc[:, "species"] = add_species(blast.data.taxo.Description)
#blast.data.cinqk.loc[:, "Description"] = add_function(blast.data.cinqk.Description)
blast.data.cinqk.loc[:, "function"] = add_function(blast.data.cinqk.Description)
blast.data.cinqk.loc[:, "species"] = add_species(blast.data.cinqk.Description)
tests = pd.read_csv("efetch-tests.csv")
with entrez.efetch(db="protein", id=tests.Accession.to_list(), rettype="gb", retmode="text") as finallyyeah:
y = [ entry for entry in seqio.parse(finallyyeah, format="gb") ]
y[0].annotations
for i in y:
print(len(i.annotations["accessions"]))
for i in y:
print(i.annotations["taxonomy"])
# SAVE AS GENEBANK !
# FASTA LOSES LOTS OF INFORMATION !
blast.data.cinqk.head()
with entrez.efetch(db="protein", id=blast.data.cinqk.Accession.to_list(), rettype="gb", retmode="text") as in_handle:
with open("5k-info.gb", "w") as out_handle:
sequences = seqio.parse(in_handle, format="gb")
seqio.write(sequences, out_handle, format="gb")
# some dummy test :
with entrez.efetch(db="nucleotide", id="EU490707", rettype="gb", retmode="text") as wow:
print(wow.read())
help(entrez.efetch)
| workflow/Entrez.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/weg92/Final/blob/master/William_Gleason_HW2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="O2P3Ez5gDb1K" colab_type="code" outputId="32486b79-42c5-4dfe-cc4d-4fd1bc9919de" colab={"base_uri": "https://localhost:8080/", "height": 938}
# Author: <NAME> <<EMAIL>>
# <NAME> <<EMAIL>>
# Class: IS 465-002
# The following program extracts themes or subjects from a set of documents for
# readability. The end reader can then find out the topic of an article before
# reading it. Frobenius norm and generalized Kullback-Leibler divergence are
# utilized in this code. Each assist in Non-negative Matrix Factorization or
# NMF, a concept (tool) that analyzes a data set and extracts information of
# importance. Within the code, there are multiple ways to categorize the
# importance of some text in any document and reference this text to a specific
# theme based on the given text.
# The beginning section to the code will fetch the newsgroups or datasets.
# Then, it will extract data from the sets. It will conclude by displaying the
# Topic and the amount of time the code took to run for each set.
from time import time
from sklearn.feature_extraction.text import TfidfVectorizer, CountVectorizer
from sklearn.decomposition import NMF, LatentDirichletAllocation
from sklearn.datasets import fetch_20newsgroups
n_samples = 2000
n_features = 1000
n_components = 10
n_top_words = 20
def print_top_words(model, feature_names, n_top_words):
for topic_idx, topic in enumerate(model.components_):
message = "Topic #%d: " % topic_idx
message += " ".join([feature_names[i]
for i in topic.argsort()[:-n_top_words - 1:-1]])
print(message)
print()
# The code of this program mines for data that a user would be looking for.
# The program goes through newsgroups and strips the articles of headers,
# footers and quoted replies. This allows for the program to sift through
# the data and bring what is important to the user. Below is the code in
# which the program executes in-order to give users clear information.
print("Loading dataset...")
t0 = time()
data, _ = fetch_20newsgroups(shuffle=True, random_state=1,
remove=('headers', 'footers', 'quotes'),
return_X_y=True)
data_samples = data[:n_samples]
print("done in %0.3fs." % (time() - t0))
# Use tf-idf (term frequency-inverse document frequency) features for NMF.
print("Extracting tf-idf features for NMF...")
tfidf_vectorizer = TfidfVectorizer(max_df=0.95, min_df=2,
max_features=n_features,
stop_words='english')
t0 = time()
tfidf = tfidf_vectorizer.fit_transform(data_samples)
print("done in %0.3fs." % (time() - t0))
# Use tf (term frequency) features for LDA (Latent Dirichlet Allocation).
print("Extracting tf features for LDA...")
tf_vectorizer = CountVectorizer(max_df=0.95, min_df=2,
max_features=n_features,
stop_words='english')
t0 = time()
tf = tf_vectorizer.fit_transform(data_samples)
print("done in %0.3fs." % (time() - t0))
print()
# Fit the NMF (Non-Negative Matrix Factorization) model.
# This section of code utilizes Frobenius norm to output the data and print the
# time it took in secondsprint
("Fitting the NMF model (Frobenius norm) with tf-idf features, "
"n_samples=%d and n_features=%d..."
% (n_samples, n_features))
t0 = time()
nmf = NMF(n_components=n_components, random_state=1,
alpha=.1, l1_ratio=.5).fit(tfidf)
print("done in %0.3fs." % (time() - t0))
print("\nTopics in NMF model (Frobenius norm):")
tfidf_feature_names = tfidf_vectorizer.get_feature_names()
print_top_words(nmf, tfidf_feature_names, n_top_words)
# Fit the NMF (Non-Negative Matrix Factorization) model.
# This code uses generalized Kullback-Leibler divergence
# to output the data and print the time it took in seconds.
print("Fitting the NMF model (generalized Kullback-Leibler divergence) with "
"tf-idf features, n_samples=%d and n_features=%d..."
% (n_samples, n_features))
t0 = time()
nmf = NMF(n_components=n_components, random_state=1,
beta_loss='kullback-leibler', solver='mu', max_iter=1000, alpha=.1,
l1_ratio=.5).fit(tfidf)
print("done in %0.3fs." % (time() - t0))
# Fit the LDA (Latent Dirichlet Allocation) model.
print("\nTopics in NMF model (generalized Kullback-Leibler divergence):")
tfidf_feature_names = tfidf_vectorizer.get_feature_names()
print_top_words(nmf, tfidf_feature_names, n_top_words)
print("Fitting LDA models with tf features, "
"n_samples=%d and n_features=%d..."
% (n_samples, n_features))
lda = LatentDirichletAllocation(n_components=n_components, max_iter=5,
learning_method='online',
learning_offset=50.,
random_state=0)
t0 = time()
lda.fit(tf)
print("done in %0.3fs." % (time() - t0))
print("\nTopics in LDA model:")
tf_feature_names = tf_vectorizer.get_feature_names()
print_top_words(lda, tf_feature_names, n_top_words)
# + id="6X9vhMbmckLE" colab_type="code" outputId="7395d505-d84a-47ac-cabd-500dd1d6a059" colab={"base_uri": "https://localhost:8080/", "height": 54}
wordlist = data_samples[0].split()
BigString = ' '.join(data_samples)
wordfreq = []
for w in BigString:
wordfreq.append(wordlist.count(w))
str(list(zip(wordlist,wordfreq)))
| William_Gleason_HW2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from sklearn.preprocessing import LabelEncoder
from sklearn.preprocessing import MinMaxScaler
import numpy as np
import mahotas
import glob
import cv2
import os
import h5py
# fixed-sizes for image
fixed_size = tuple((500, 500))
# path to training data
train_path = "dataset/train"
# no.of.trees for Random Forests
num_trees = 100
# bins for histogram
bins = 8
# train_test_split size
test_size = 0.10
# seed for reproducing same results
seed = 9
# feature-descriptor-1: Hu Moments
def fd_hu_moments(image):
image = cv2.cvtColor(image, cv2.COLOR_BGR2GRAY)
feature = cv2.HuMoments(cv2.moments(image)).flatten()
return feature
# feature-descriptor-2: Haralick Texture
def fd_haralick(image):
# convert the image to grayscale
gray = cv2.cvtColor(image, cv2.COLOR_BGR2GRAY)
# compute the haralick texture feature vector
haralick = mahotas.features.haralick(gray).mean(axis=0)
# return the result
return haralick
# feature-descriptor-3: Color Histogram
def fd_histogram(image, mask=None):
# convert the image to HSV color-space
image = cv2.cvtColor(image, cv2.COLOR_BGR2HSV)
# compute the color histogram
hist = cv2.calcHist([image], [0, 1, 2], None, [bins, bins, bins], [0, 256, 0, 256, 0, 256])
# normalize the histogram
cv2.normalize(hist, hist)
# return the histogram
return hist.flatten()
# get the training labels
train_labels = os.listdir(train_path)
# sort the training labels
train_labels.sort()
print(train_labels)
# empty lists to hold feature vectors and labels
global_features = []
labels = []
i, j = 0, 0
k = 0
# num of images per class
images_per_class = 80
# +
# loop over the training data sub-folders
for training_name in train_labels:
# join the training data path and each species training folder
dir = os.path.join(train_path, training_name)
# get the current training label
current_label = training_name
k = 1
# loop over the images in each sub-folder
for x in range(1,images_per_class+1):
# get the image file name
file = dir + "/" + str(x) + ".jpg"
# read the image and resize it to a fixed-size
image = cv2.imread(file)
image = cv2.resize(image, fixed_size)
####################################
# Global Feature extraction
####################################
fv_hu_moments = fd_hu_moments(image)
fv_haralick = fd_haralick(image)
fv_histogram = fd_histogram(image)
###################################
# Concatenate global features
###################################
global_feature = np.hstack([fv_histogram, fv_haralick, fv_hu_moments])
# update the list of labels and feature vectors
labels.append(current_label)
global_features.append(global_feature)
i += 1
k += 1
print ("[STATUS] processed folder: {}".format(current_label))
j += 1
print( "[STATUS] completed Global Feature Extraction...")
# +
# get the overall feature vector size
print ("[STATUS] feature vector size {}".format(np.array(global_features).shape))
# get the overall training label size
print ("[STATUS] training Labels {}".format(np.array(labels).shape))
# encode the target labels
targetNames = np.unique(labels)
le = LabelEncoder()
target = le.fit_transform(labels)
print ("[STATUS] training labels encoded...")
# normalize the feature vector in the range (0-1)
scaler = MinMaxScaler(feature_range=(0, 1))
rescaled_features = scaler.fit_transform(global_features)
print ("[STATUS] feature vector normalized...")
print ("[STATUS] target labels: {}".format(target))
print ("[STATUS] target labels shape: {}".format(target.shape))
# save the feature vector using HDF5
h5f_data = h5py.File('output/data.h5', 'w')
h5f_data.create_dataset('dataset_1', data=np.array(rescaled_features))
h5f_label = h5py.File('output/labels.h5', 'w')
h5f_label.create_dataset('dataset_1', data=np.array(target))
h5f_data.close()
h5f_label.close()
print ("[STATUS] end of training..")
| .ipynb_checkpoints/glo-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Continuous Integration
# + [markdown] slideshow={"slide_type": "slide"}
# ## Overview:
# - **Teaching:** 10 min
# - **Exercises:** 0 min
#
# **Questions**
# - How can I automate running the tests on more platforms than my own?
#
# **Objectives**
# - Understand how continuous integration speeds software development
# - Understand the benefits of continuous integration
# - Implement a continuous integration server
# - Identify a few options for hosting a continuous integration server
# + [markdown] slideshow={"slide_type": "slide"}
# To make running the tests as easy as possible, many software development teams implement a strategy called continuous integration. As its name implies, continuous integration integrates the test suite into the development process. Every time a change is made to the repository, the continuous integration system builds and checks that code.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Information: Thought Experiment: Does your software work on your colleague’s computer?
# Imagine you developed software on a MacOSX computer. Last week, you helped your office mate build and run it on their Linux computer. You’ve made some changes since then.
#
# 1. How can you be sure it will still work if they update their repository when they come back from vacation?
# 2. How long will that process take?
# + [markdown] slideshow={"slide_type": "slide"}
# How long will that process take?
# The typical story in a research lab is that, well, you don’t know whether it will work on your colleagues’ machine until you try rebuilding it on their machine. If you have a build system, it might take a few minutes to update the repository, rebuild the code, and run the tests. If you don’t have a build system, it could take all afternoon just to see if your new changes are compatible.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Let the computers do the work
# Scientists are good at creative insights, conceptual understanding, critical analysis, and consuming espresso. Computers are good at following instructions. Science would be more fun if the scientists could just give the computers the instructions and go grab an espresso.
#
# Continuous integration servers allow just that. Based on your instructions, a continuous integration server can:
#
# - check out new code from a repository
# - spin up instances of supported operating systems (i.e. various versions of OSX, Linux, Windows, etc.).
# - spin up those instances with different software versions (i.e. Python 2.7 and Python 3.0)
# - run the build and test scripts
# - check for errors
# - and report the results.
# + [markdown] slideshow={"slide_type": "slide"}
# Since the first step the server conducts is to check out the code from a repository, we’ll need to put our code online to make use of this kind of server (unless we are able/willing to set up our own CI server).
# + [markdown] slideshow={"slide_type": "slide"}
# ## Exercise: Set up a mean git repository on GitHub
# Your `mean.py` and `test_mean.py` files can be the contents of a repository on GitHub.
#
# Go to GitHub and create a repository called mean.
# Clone that repository (git clone https://github.com:yourusername/mean)
# Copy the `mean.py` and `test_mean.py` files into the repository directory.
# Use git to add, commit, and push the two files to GitHub.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Giving Instructions
# Your work on the mean function has both code and tests. Let’s copy that code into its own repository and add continuous integration to that repository.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Information: What is required?
# It doesn’t need a build system, because Python does not need to be compiled.
#
# 1. What does it need?
# 2. Write the names of the software dependencies in a file called `requirements.txt` and save the file.
# 3. In fact, why don’t you go ahead and version control it?
# + [markdown] slideshow={"slide_type": "slide"}
# ## Travis-CI
# Travis is a continous integration server hosting platform. It’s commonly used in Ruby development circles as well as in the scientific Python community.
#
# To use Travis, all you need is an account. It’s free so someone in your group should sign up for a Travis account. Then follow the instructions on the Travis website to connect your Travis account with GitHub.
#
# A file called `.travis.yml` in your repository will signal to Travis that you want to build and test this repository on Travis-CI. Such a file, for our purposes, is very simple:
# ```yaml
# language: python
# python:
# - "2.6"
# - "2.7"
# - "3.2"
# - "3.3"
# - "3.4"
# - "nightly"
# # command to install dependencies
# install:
# - "pip install -r requirements.txt"
# # command to run tests
# script: py.test
# ```
# + [markdown] slideshow={"slide_type": "slide"}
# However, the exact syntax is very important, https://lint.travis-ci.org/ can be used to check for typographic errors. You can see how the Python package manager, `pip`, will use your `requirements.txt` file from the previous exercise. That `requirements.txt` file is a conventional way to list all of the python packages that we need. If we needed pytest, numpy, and pymol, the `requirements.txt` file would look like this:
#
# ```bash
# numpy
# pymol
# pytest
# ```
# + [markdown] slideshow={"slide_type": "slide"}
# ## Exercise: Last steps
# 1. Add `.travis.yml` to your repository
# 2. Commit and push it.
# 3. Check the situation at your server
# + [markdown] slideshow={"slide_type": "slide"}
# ## Continuous Integration Hosting
# We gave the example of Travis because it’s very very simple to spin up. While it is able to run many flavors of Linux, it currently doesn’t support other platforms as well. Depending on your needs, you may consider other services such as:
#
# - buildbot
# - CDash
# - Jenkins
# + [markdown] slideshow={"slide_type": "slide"}
# ## Key Points:
# - Servers exist for automatically running your tests
# - Running the tests can be triggered by a GitHub pull request
# - CI allows cross-platform build testing
# - A `.travis.yml` file configures a build on the travis-ci servers
# - Many free CI servers are available
| nbplain/10_episode.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# ## Minimum discernable reflectivty of CPOL
import pyart
import numpy as np
import time
from matplotlib import pyplot as plt
# %matplotlib inline
import math
# ## Load radar data and grid (no filter)
# +
radar_file = '/home/rjackson/data/radar/Gunn_pt_20070101000000ppi.nc'
Radar = pyart.io.read(radar_file)
display = pyart.graph.RadarMapDisplay(Radar)
display.plot_ppi('reflectivity', 0)
plt.xlim([-150, 150])
plt.ylim([-150, 150])
# -
# def grid_radar(radar, grid_shape=(20, 301, 301), xlim=(-150000, 150000),
# ylim=(-150000, 150000), zlim=(1000, 20000), bsp=1.0,
# min_radius=750, h_factor=4.0, nb=1.5,
# fields=['DT', 'VT'], origin=None, gatefilter=None):
# bt = time.time()
# radar_list = [radar]
# if origin is None:
# origin = (radar.latitude['data'][0],
# radar.longitude['data'][0])
# grid = pyart.map.grid_from_radars(
# radar_list, grid_shape=grid_shape,
# grid_limits=(zlim, ylim, xlim),
# grid_origin=origin, fields=fields,
# weighting_function='Cressman',
# gridding_algo='map_gates_to_grid',
# h_factor=h_factor,
# min_radius=min_radius,
# bsp=bsp,
# nb=nb,
# gatefilter=gatefilter)
# print(time.time() - bt, 'seconds to grid radar')
# return grid
Grid = grid_radar(Radar,
origin=(Radar.latitude['data'][0], Radar.longitude['data'][0]),
xlim=(-150000, 150000), ylim=(-150000, 150000),
fields=['reflectivity'], min_radius=500.0, bsp=1.0, nb=1.5,
h_factor=2.0,
zlim=(500, 20000), grid_shape=(40, 121, 121))
plt.figure(figsize=(10,10))
plt.subplot(221)
CPOLGridDisplay = pyart.graph.GridMapDisplay(Grid)
CPOLGridDisplay.plot_basemap(min_lat=-13.0, max_lat=-11.8,
min_lon=130.3, max_lon=131.7,
lat_lines=[-13.1, -12.7, -12.3, -11.9, -11.5],
lon_lines=[130.0, 130.5, 131.0, 131.5, 132.0],
resolution='i')
CPOLGridDisplay.plot_grid('reflectivity', 1, vmin=-20, vmax=10,
cmap=pyart.graph.cm.NWSRef)
plt.subplot(222)
CPOLGridDisplay = pyart.graph.GridMapDisplay(Grid)
CPOLGridDisplay.plot_basemap(min_lat=-13.0, max_lat=-11.8,
min_lon=130.3, max_lon=131.7,
lat_lines=[-13.1, -12.7, -12.3, -11.9, -11.5],
lon_lines=[130.0, 130.5, 131.0, 131.5, 132.0],
resolution='i')
CPOLGridDisplay.plot_grid('reflectivity', 5, vmin=-20, vmax=10,
cmap=pyart.graph.cm.NWSRef)
plt.subplot(223)
CPOLGridDisplay = pyart.graph.GridMapDisplay(Grid)
CPOLGridDisplay.plot_basemap(min_lat=-13.0, max_lat=-11.8,
min_lon=130.3, max_lon=131.7,
lat_lines=[-13.1, -12.7, -12.3, -11.9, -11.5],
lon_lines=[130.0, 130.5, 131.0, 131.5, 132.0],
resolution='i')
CPOLGridDisplay.plot_grid('reflectivity', 20, vmin=-20, vmax=10,
cmap=pyart.graph.cm.NWSRef)
plt.subplot(224)
CPOLGridDisplay = pyart.graph.GridMapDisplay(Grid)
CPOLGridDisplay.plot_basemap(min_lat=-13.0, max_lat=-11.8,
min_lon=130.3, max_lon=131.7,
lat_lines=[-13.1, -12.7, -12.3, -11.9, -11.5],
lon_lines=[130.0, 130.5, 131.0, 131.5, 132.0],
resolution='i')
CPOLGridDisplay.plot_grid('reflectivity', 25, vmin=-20, vmax=10,
cmap=pyart.graph.cm.NWSRef)
# ## Calculate reflectivity vs. distance from radar
x = Grid.point_x['data']
y = Grid.point_y['data']
dist_from_center = np.sqrt(np.square(x)+np.square(y))/1e3
dist_bins = np.arange(0, 150, 1)
reflectivity = np.nan*np.ones((40, len(dist_bins)))
std = np.nan*np.ones((40, len(dist_bins)))
for levels in range(0, 40):
for i in range(0, len(dist_bins)-1):
which = np.where(np.logical_and(dist_from_center[levels] >= dist_bins[i],
dist_from_center[levels] < dist_bins[i+1]))
reflectivity[levels,i] = np.ma.mean(Grid.fields['reflectivity']['data'][levels][which])
std[levels,i] = np.ma.std(Grid.fields['reflectivity']['data'][levels][which])
dist_bins_grid, z_grid = np.meshgrid(dist_bins, np.arange(0,20,0.5))
plt.pcolormesh(dist_bins_grid, z_grid, reflectivity, vmin=-30, vmax=10)
ax = plt.colorbar()
ax.set_label('Reflectivity [dBZ]')
plt.xlabel('Horizontal Distance from CPOL [km]')
plt.ylabel('Level')
# +
plt.figure(figsize=(6,12))
plt.subplot(311)
plt.plot(dist_bins_grid[20], reflectivity[20])
plt.plot(dist_bins_grid[30], reflectivity[30])
plt.plot(dist_bins_grid[39], reflectivity[39])
plt.plot([60,60], [-15,5], linestyle='--')
plt.legend(['10 km', '15 km', '20 km'], loc='best')
plt.ylim([-15, 5])
plt.xlabel('Horizontal distance from Radar [km]')
plt.ylabel('Noise floor [dBZ]')
plt.subplot(312)
plt.plot(dist_bins_grid[20], reflectivity[20]+std[20])
plt.plot(dist_bins_grid[30], reflectivity[30]+std[20])
plt.plot(dist_bins_grid[39], reflectivity[39]+std[20])
plt.plot([60,60], [-15,5], linestyle='--')
plt.legend(['10 km', '15 km', '20 km'], loc='best')
plt.ylim([-15, 5])
plt.xlabel('Horizontal distance from Radar [km]')
plt.ylabel('Noise floor + 1 std. dev. [dBZ]')
plt.subplot(313)
plt.plot(dist_bins_grid[20], reflectivity[20]+3*std[20])
plt.plot(dist_bins_grid[30], reflectivity[30]+3*std[20])
plt.plot(dist_bins_grid[39], reflectivity[39]+3*std[20])
plt.plot([60,60], [-15,5], linestyle='--')
plt.legend(['10 km', '15 km', '20 km'], loc='best')
plt.ylim([-15, 5])
plt.xlabel('Horizontal distance from Radar [km]')
plt.ylabel('Noise floor + 3 std. dev. [dBZ]')
| notebooks/Minimum discernable reflectivity.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="Tce3stUlHN0L"
# ##### Copyright 2018 The TensorFlow Authors.
#
# + cellView="form" colab={} colab_type="code" id="tuOe1ymfHZPu"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="MfBg1C5NB3X0"
# # 텐서플로로 분산 훈련하기
# + [markdown] colab_type="text" id="r6P32iYYV27b"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/guide/distributed_training"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />TensorFlow.org에서 보기</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/ko/guide/distributed_training.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />구글 코랩(Colab)에서 실행하기</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/ko/guide/distributed_training.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />깃허브(GitHub) 소스 보기</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/docs-l10n/site/ko/guide/distributed_training.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="vqrz9ZBdzQ9C"
# Note: 이 문서는 텐서플로 커뮤니티에서 번역했습니다. 커뮤니티 번역 활동의 특성상 정확한 번역과 최신 내용을 반영하기 위해 노력함에도
# 불구하고 [공식 영문 문서](https://www.tensorflow.org/?hl=en)의 내용과 일치하지 않을 수 있습니다.
# 이 번역에 개선할 부분이 있다면
# [tensorflow/docs-l10n](https://github.com/tensorflow/docs-l10n/) 깃헙 저장소로 풀 리퀘스트를 보내주시기 바랍니다.
# 문서 번역이나 리뷰에 참여하려면
# [<EMAIL>](https://groups.google.com/a/tensorflow.org/forum/#!forum/docs-ko)로
# 메일을 보내주시기 바랍니다.
# + [markdown] colab_type="text" id="xHxb-dlhMIzW"
# ## 개요
#
# `tf.distribute.Strategy`는 훈련을 여러 GPU 또는 여러 장비, 여러 TPU로 나누어 처리하기 위한 텐서플로 API입니다. 이 API를 사용하면 기존의 모델이나 훈련 코드를 조금만 고쳐서 분산처리를 할 수 있습니다.
#
# `tf.distribute.Strategy`는 다음을 주요 목표로 설계하였습니다.
#
# * 사용하기 쉽고, 연구원, 기계 학습 엔지니어 등 여러 사용자 층을 지원할 것.
# * 그대로 적용하기만 하면 좋은 성능을 보일 것.
# * 전략들을 쉽게 갈아 끼울 수 있을 것.
#
# `tf.distribute.Strategy`는 텐서플로의 고수준 API인 [tf.keras](https://www.tensorflow.org/guide/keras) 및 [tf.estimator](https://www.tensorflow.org/guide/estimator)와 함께 사용할 수 있습니다. 코드 한두 줄만 추가하면 됩니다. 사용자 정의 훈련 루프(그리고 텐서플로를 사용한 모든 계산 작업)에 함께 사용할 수 있는 API도 제공합니다.
# 텐서플로 2.0에서는 사용자가 프로그램을 즉시 실행(eager execution)할 수도 있고, [`tf.function`](../tutorials/eager/tf_function.ipynb)을 사용하여 그래프에서 실행할 수도 있습니다. `tf.distribute.Strategy`는 두 가지 실행 방식을 모두 지원하려고 합니다. 이 가이드에서는 대부분의 경우 훈련에 대하여 이야기하겠지만, 이 API 자체는 여러 환경에서 평가나 예측을 분산 처리하기 위하여 사용할 수도 있다는 점을 참고하십시오.
#
# 잠시 후 보시겠지만 코드를 약간만 바꾸면 `tf.distribute.Strategy`를 사용할 수 있습니다. 변수, 층, 모델, 옵티마이저, 지표, 서머리(summary), 체크포인트 등 텐서플로를 구성하고 있는 기반 요소들을 전략(Strategy)을 이해하고 처리할 수 있도록 수정했기 때문입니다.
#
# 이 가이드에서는 다양한 형식의 전략에 대해서, 그리고 여러 가지 상황에서 이들을 어떻게 사용해야 하는지 알아보겠습니다.
# + colab={} colab_type="code" id="EVOZFbNgXghB"
# 텐서플로 패키지 가져오기
# !pip install tensorflow-gpu==2.0.0-rc1
import tensorflow as tf
# + [markdown] colab_type="text" id="eQ1QESxxEbCh"
# ## 전략의 종류
# `tf.distribute.Strategy`는 서로 다른 다양한 사용 형태를 아우르려고 합니다. 몇 가지 조합은 현재 지원하지만, 추후에 추가될 전략들도 있습니다. 이들 중 몇 가지를 살펴보겠습니다.
#
# * 동기 훈련 대 비동기 훈련: 분산 훈련을 할 때 데이터를 병렬로 처리하는 방법은 크게 두 가지가 있습니다. 동기 훈련을 할 때는 모든 워커(worker)가 입력 데이터를 나누어 갖고 동시에 훈련합니다. 그리고 각 단계마다 그래디언트(gradient)를 모읍니다. 비동기 훈련에서는 모든 워커가 독립적으로 입력 데이터를 사용해 훈련하고 각각 비동기적으로 변수들을 갱신합니다. 일반적으로 동기 훈련은 올 리듀스(all-reduce)방식으로 구현하고, 비동기 훈련은 파라미터 서버 구조를 사용합니다.
# * 하드웨어 플랫폼: 한 장비에 있는 다중 GPU로 나누어 훈련할 수도 있고, 네트워크로 연결된 (GPU가 없거나 여러 개의 GPU를 가진) 여러 장비로 나누어서, 또 혹은 클라우드 TPU에서 훈련할 수도 있습니다.
#
# 이런 사용 형태들을 위하여, 현재 5가지 전략을 사용할 수 있습니다. 이후 내용에서 현재 TF 2.0 베타에서 상황마다 어떤 전략을 지원하는지 이야기하겠습니다. 일단 간단한 개요는 다음과 같습니다.
#
# | 훈련 API | MirroredStrategy | TPUStrategy | MultiWorkerMirroredStrategy | CentralStorageStrategy | ParameterServerStrategy |
# |:----------------------- |:------------------- |:--------------------- |:--------------------------------- |:--------------------------------- |:-------------------------- |
# | **Keras API** | 지원 | 2.0 RC 지원 예정 | 실험 기능으로 지원 | 실험 기능으로 지원 | 2.0 이후 지원 예정 |
# | **사용자 정의 훈련 루프** | 실험 기능으로 지원 | 실험 기능으로 지원 | 2.0 이후 지원 예정 | 2.0 RC 지원 예정 | 아직 미지원 |
# | **Estimator API** | 제한적으로 지원 | 제한적으로 지원 | 제한적으로 지원 | 제한적으로 지원 | 제한적으로 지원 |
# + [markdown] colab_type="text" id="DoQKKK8dtfg6"
# ### MirroredStrategy
# `tf.distribute.MirroredStrategy`는 장비 하나에서 다중 GPU를 이용한 동기 분산 훈련을 지원합니다. 각각의 GPU 장치마다 복제본이 만들어집니다. 모델의 모든 변수가 복제본마다 미러링 됩니다. 이 미러링된 변수들은 하나의 가상의 변수에 대응되는데, 이를 `MirroredVariable`라고 합니다. 이 변수들은 동일한 변경사항이 함께 적용되므로 모두 같은 값을 유지합니다.
#
# 여러 장치에 변수의 변경사항을 전달하기 위하여 효율적인 올 리듀스 알고리즘을 사용합니다. 올 리듀스 알고리즘은 모든 장치에 걸쳐 텐서를 모은 다음, 그 합을 구하여 다시 각 장비에 제공합니다. 이 통합된 알고리즘은 매우 효율적이어서 동기화의 부담을 많이 덜어낼 수 있습니다. 장치 간에 사용 가능한 통신 방법에 따라 다양한 올 리듀스 알고리즘과 구현이 있습니다. 기본값으로는 NVIDIA NCCL을 올 리듀스 구현으로 사용합니다. 또한 제공되는 다른 몇 가지 방법 중에 선택하거나, 직접 만들 수도 있습니다.
#
# `MirroredStrategy`를 만드는 가장 쉬운 방법은 다음과 같습니다.
# + colab={} colab_type="code" id="9Z4FMAY9ADxK"
mirrored_strategy = tf.distribute.MirroredStrategy()
# + [markdown] colab_type="text" id="wldY4aFCAH4r"
# `MirroredStrategy` 인스턴스가 생겼습니다. 텐서플로가 인식한 모든 GPU를 사용하고, 장치 간 통신에는 NCCL을 사용할 것입니다.
#
# 장비의 GPU 중 일부만 사용하고 싶다면, 다음과 같이 하면 됩니다.
# + colab={} colab_type="code" id="nbGleskCACv_"
mirrored_strategy = tf.distribute.MirroredStrategy(devices=["/gpu:0", "/gpu:1"])
# + [markdown] colab_type="text" id="8-KDnrJLAhav"
# 장치 간 통신 방법을 바꾸고 싶다면, `cross_device_ops` 인자에 `tf.distribute.CrossDeviceOps` 타입의 인스턴스를 넘기면 됩니다. 현재 기본값인 `tf.distribute.NcclAllReduce` 이외에 `tf.distribute.HierarchicalCopyAllReduce`와 `tf.distribute.ReductionToOneDevice` 두 가지 추가 옵션을 제공합니다.
# + colab={} colab_type="code" id="6-xIOIpgBItn"
mirrored_strategy = tf.distribute.MirroredStrategy(
cross_device_ops=tf.distribute.HierarchicalCopyAllReduce())
# + [markdown] colab_type="text" id="45H0Wa8WKI8z"
# ### CentralStorageStrategy
# `tf.distribute.experimental.CentralStorageStrategy`도 동기 훈련을 합니다. 하지만 변수를 미러링하지 않고, CPU에서 관리합니다. 작업은 모든 로컬 GPU들로 복제됩니다. 단, 만약 GPU가 하나밖에 없다면 모든 변수와 작업이 그 GPU에 배치됩니다.
#
# 다음과 같이 `CentralStorageStrategy` 인스턴스를 만드십시오.
# + colab={} colab_type="code" id="rtjZOyaoMWrP"
central_storage_strategy = tf.distribute.experimental.CentralStorageStrategy()
# + [markdown] colab_type="text" id="KY1nJHNkMl7b"
# `CentralStorageStrategy` 인스턴스가 만들어졌습니다. 인식한 모든 GPU와 CPU를 사용합니다. 각 복제본의 변수 변경사항은 모두 수집된 후 변수에 적용됩니다.
# + [markdown] colab_type="text" id="aAFycYUiNCUb"
# Note: 이 전략은 아직 개선 중이고 더 많은 경우에 쓸 수 있도록 만들고 있기 때문에, [`실험 기능`](https://www.tensorflow.org/guide/versions#what_is_not_covered)으로 지원됩니다. 따라서 다음에 API가 바뀔 수 있음에 유념하십시오.
# + [markdown] colab_type="text" id="8Xc3gyo0Bejd"
# ### MultiWorkerMirroredStrategy
#
# `tf.distribute.experimental.MultiWorkerMirroredStrategy`은 `MirroredStrategy`와 매우 비슷합니다. 다중 워커를 이용하여 동기 분산 훈련을 합니다. 각 워커는 여러 개의 GPU를 사용할 수 있습니다. `MirroredStrategy`처럼 모델에 있는 모든 변수의 복사본을 모든 워커의 각 장치에 만듭니다.
#
# 다중 워커(multi-worker)들 사이에서는 올 리듀스(all-reduce) 통신 방법으로 [CollectiveOps](https://github.com/tensorflow/tensorflow/blob/master/tensorflow/python/ops/collective_ops.py)를 사용하여 변수들을 같은 값으로 유지합니다. 수집 연산(collective op)은 텐서플로 그래프에 속하는 연산 중 하나입니다. 이 연산은 하드웨어나 네트워크 구성, 텐서 크기에 따라 텐서플로 런타임이 지원하는 올 리듀스 알고리즘을 자동으로 선택합니다.
#
# 여기에 추가 성능 최적화도 구현하고 있습니다. 예를 들어 작은 텐서들의 여러 올 리듀스 작업을 큰 텐서들의 더 적은 올 리듀스 작업으로 바꾸는 정적 최적화 기능이 있습니다. 뿐만아니라 플러그인 구조를 갖도록 설계하였습니다. 따라서 추후에는 사용자가 자신의 하드웨어에 더 최적화된 알고리즘을 사용할 수도 있을 것입니다. 참고로 이 수집 연산은 올 리듀스 외에 브로드캐스트(broadcast)나 전체 수집(all-gather)도 구현하고 있습니다.
#
# `MultiWorkerMirroredStrategy`를 만드는 가장 쉬운 방법은 다음과 같습니다.
# + colab={} colab_type="code" id="m3a_6ebbEjre"
multiworker_strategy = tf.distribute.experimental.MultiWorkerMirroredStrategy()
# + [markdown] colab_type="text" id="bt94JBvhEr4s"
# `MultiWorkerMirroredStrategy`에 사용할 수 있는 수집 연산 구현은 현재 두 가지입니다. `CollectiveCommunication.RING`는 gRPC를 사용한 링 네트워크 기반의 수집 연산입니다. `CollectiveCommunication.NCCL`는 [Nvidia의 NCCL](https://developer.nvidia.com/nccl)을 사용하여 수집 연산을 구현한 것입니다. `CollectiveCommunication.AUTO`로 설정하면 런타임이 알아서 구현을 고릅니다. 최적의 수집 연산 구현은 GPU의 수와 종류, 클러스터의 네트워크 연결 등에 따라 다를 수 있습니다. 예를 들어 다음과 같이 지정할 수 있습니다.
# + colab={} colab_type="code" id="QGX_QAEtFQSv"
multiworker_strategy = tf.distribute.experimental.MultiWorkerMirroredStrategy(
tf.distribute.experimental.CollectiveCommunication.NCCL)
# + [markdown] colab_type="text" id="0JiImlw3F77E"
# 다중 GPU를 사용하는 것과 비교해서 다중 워커를 사용하는 것의 가장 큰 차이점은 다중 워커에 대한 설정 부분입니다. 클러스터를 구성하는 각 워커에 "TF_CONFIG" 환경변수를 사용하여 클러스터 설정을 하는 것이 텐서플로의 표준적인 방법입니다. [아래쪽 "TF_CONFIG"](#TF_CONFIG) 항목에서 어떻게 하는지 자세히 살펴보겠습니다.
# + [markdown] colab_type="text" id="E20tG21LFfv1"
# Note: 이 전략은 아직 개선 중이고 더 많은 경우에 쓸 수 있도록 만들고 있기 때문에, [`실험 기능`](https://www.tensorflow.org/guide/versions#what_is_not_covered)으로 지원됩니다. 따라서 나중에 API가 바뀔 수 있음에 유념하십시오.
# + [markdown] colab_type="text" id="kPEBCMzsGaO5"
# ### TPUStrategy
# `tf.distribute.experimental.TPUStrategy`는 텐서플로 훈련을 텐서처리장치(Tensor Processing Unit, TPU)에서 수행하는 전략입니다. TPU는 구글의 특별한 주문형 반도체(ASIC)로서, 기계 학습 작업을 극적으로 가속하기 위하여 설계되었습니다. TPU는 구글 코랩, [Tensorflow Research Cloud](https://www.tensorflow.org/tfrc), [Google Compute Engine](https://cloud.google.com/tpu)에서 사용할 수 있습니다.
#
# 분산 훈련 구조의 측면에서, TPUStrategy는 `MirroredStrategy`와 동일합니다. 동기 분산 훈련 방식을 사용합니다. TPU는 자체적으로 여러 TPU 코어들에 걸친 올 리듀스 및 기타 수집 연산을 효율적으로 구현하고 있습니다. 이 구현이 `TPUStrategy`에 사용됩니다.
#
# `TPUStrategy`를 사용하는 방법은 다음과 같습니다.
#
# Note: 코랩에서 이 코드를 사용하려면, 코랩 런타임으로 TPU를 선택해야 합니다. TPUStrategy를 사용하는 방법에 대한 튜토리얼을 곧 추가하겠습니다.
#
# ```
# cluster_resolver = tf.distribute.cluster_resolver.TPUClusterResolver(
# tpu=tpu_address)
# tf.config.experimental_connect_to_host(cluster_resolver.master())
# tf.tpu.experimental.initialize_tpu_system(cluster_resolver)
# tpu_strategy = tf.distribute.experimental.TPUStrategy(cluster_resolver)
# ```
# + [markdown] colab_type="text" id="oQ7EqjpmK6DU"
# `TPUClusterResolver` 인스턴스는 TPU를 찾도록 도와줍니다. 코랩에서는 아무런 인자를 주지 않아도 됩니다. 클라우드 TPU에서 사용하려면, TPU 자원의 이름을 `tpu` 매개변수에 지정해야 합니다. 또한 TPU는 계산하기 전 초기화(initialize)가 필요합니다. 초기화 중 TPU 메모리가 지워져서 모든 상태 정보가 사라지므로, 프로그램 시작시에 명시적으로 TPU 시스템을 초기화(initialize)해 주어야 합니다.
# + [markdown] colab_type="text" id="jARHpraJMbRa"
# Note: 이 전략은 아직 개선 중이고 더 많은 경우에 쓸 수 있도록 만들고 있기 때문에, [`실험 기능`](https://www.tensorflow.org/guide/versions#what_is_not_covered)으로 지원됩니다. 따라서 나중에 API가 바뀔 수 있음에 유념하십시오.
# + [markdown] colab_type="text" id="3ZLBhaP9NUNr"
# ### ParameterServerStrategy
# `tf.distribute.experimental.ParameterServerStrategy`은 여러 장비에서 훈련할 때 파라미터 서버를 사용합니다. 이 전략을 사용하면 몇 대의 장비는 워커 역할을 하고, 몇 대는 파라미터 서버 역할을 하게 됩니다. 모델의 각 변수는 한 파라미터 서버에 할당됩니다. 계산 작업은 모든 워커의 GPU들에 복사됩니다.
#
# 코드만 놓고 보았을 때는 다른 전략들과 비슷합니다.
# ```
# ps_strategy = tf.distribute.experimental.ParameterServerStrategy()
# ```
# + [markdown] colab_type="text" id="zr1wPHYvOH0N"
# 다중 워커 환경에서 훈련하려면, 클러스터에 속한 파라미터 서버와 워커를 "TF_CONFIG" 환경변수를 이용하여 설정해야 합니다. 자세한 내용은 [아래쪽 "TF_CONFIG"](#TF_CONFIG)에서 설명하겠습니다.
# + [markdown] colab_type="text" id="hQv1lm9UPDFy"
# 여기까지 여러 가지 전략들이 어떻게 다르고, 어떻게 사용하는지 살펴보았습니다. 이어지는 절들에서는 훈련을 분산시키기 위하여 이들을 어떻게 사용해야 하는지 살펴보겠습니다. 이 문서에서는 간단한 코드 조각만 보여드리겠지만, 처음부터 끝까지 전체 코드를 실행할 수 있는 더 긴 튜토리얼의 링크도 함께 안내해드리겠습니다.
# + [markdown] colab_type="text" id="_mcuy3UhPcen"
# ## 케라스와 함께 `tf.distribute.Strategy` 사용하기
# `tf.distribute.Strategy`는 텐서플로의 [케라스 API 명세](https://keras.io) 구현인 `tf.keras`와 함께 사용할 수 있습니다. `tf.keras`는 모델을 만들고 훈련하는 고수준 API입니다. 분산 전략을 `tf.keras` 백엔드와 함께 쓸 수 있으므로, 케라스 사용자들도 케라스 훈련 프레임워크로 작성한 훈련 작업을 쉽게 분산 처리할 수 있게 되었습니다. 훈련 프로그램에서 고쳐야하는 부분은 거의 없습니다. (1) 적절한 `tf.distribute.Strategy` 인스턴스를 만든 다음 (2)
# 케라스 모델의 생성과 컴파일을 `strategy.scope` 안으로 옮겨주기만 하면 됩니다. `Sequential` , 함수형 API, 클래스 상속 등 모든 방식으로 만든 케라스 모델을 다 지원합니다.
#
# 다음은 한 개의 밀집 층(dense layer)을 가진 매우 간단한 케라스 모델에 분산 전략을 사용하는 코드의 일부입니다.
# + colab={} colab_type="code" id="gbbcpzRnPZ6V"
mirrored_strategy = tf.distribute.MirroredStrategy()
with mirrored_strategy.scope():
model = tf.keras.Sequential([tf.keras.layers.Dense(1, input_shape=(1,))])
model.compile(loss='mse', optimizer='sgd')
# + [markdown] colab_type="text" id="773EOxCRVlTg"
# 위 예에서는 `MirroredStrategy`를 사용했기 때문에, 하나의 장비가 다중 GPU를 가진 경우에 사용할 수 있습니다. `strategy.scope()`로 분산 처리할 부분을 코드에 지정할 수 있습니다. 이 범위(scope) 안에서 모델을 만들면, 일반적인 변수가 아니라 미러링된 변수가 만들어집니다. 이 범위 안에서 컴파일을 한다는 것은 작성자가 이 전략을 사용하여 모델을 훈련하려고 한다는 의미입니다. 이렇게 구성하고 나서, 일반적으로 실행하는 것처럼 모델의 `fit` 함수를 호출합니다.
# `MirroredStrategy`가 모델의 훈련을 사용 가능한 GPU들로 복제하고, 그래디언트들을 수집하는 것 등을 알아서 처리합니다.
# + colab={} colab_type="code" id="ZMmxEFRTEjH5"
dataset = tf.data.Dataset.from_tensors(([1.], [1.])).repeat(100).batch(10)
model.fit(dataset, epochs=2)
model.evaluate(dataset)
# + [markdown] colab_type="text" id="nofTLwyXWHK8"
# 위에서는 훈련과 평가 입력을 위해 `tf.data.Dataset`을 사용했습니다. 넘파이(numpy) 배열도 사용할 수 있습니다.
# + colab={} colab_type="code" id="Lqgd9SdxW5OW"
import numpy as np
inputs, targets = np.ones((100, 1)), np.ones((100, 1))
model.fit(inputs, targets, epochs=2, batch_size=10)
# + [markdown] colab_type="text" id="IKqaj7QwX0Zb"
# 데이터셋이나 넘파이를 사용하는 두 경우 모두 입력 배치가 동일한 크기로 나누어져서 여러 개로 복제된 작업에 전달됩니다. 예를 들어, `MirroredStrategy`를 2개의 GPU에서 사용한다면, 크기가 10개인 배치(batch)가 두 개의 GPU로 배분됩니다. 즉, 각 GPU는 한 단계마다 5개의 입력을 받게 됩니다. 따라서 GPU가 추가될수록 각 에포크(epoch) 당 훈련 시간은 줄어들게 됩니다. 일반적으로는 가속기를 더 추가할 때마다 배치 사이즈도 더 키웁니다. 추가한 컴퓨팅 자원을 더 효과적으로 사용하기 위해서입니다. 모델에 따라서는 학습률(learning rate)을 재조정해야 할 수도 있을 것입니다. 복제본의 수는 `strategy.num_replicas_in_sync`로 얻을 수 있습니다.
# + colab={} colab_type="code" id="quNNTytWdGBf"
# 복제본의 수로 전체 배치 크기를 계산.
BATCH_SIZE_PER_REPLICA = 5
global_batch_size = (BATCH_SIZE_PER_REPLICA *
mirrored_strategy.num_replicas_in_sync)
dataset = tf.data.Dataset.from_tensors(([1.], [1.])).repeat(100)
dataset = dataset.batch(global_batch_size)
LEARNING_RATES_BY_BATCH_SIZE = {5: 0.1, 10: 0.15}
learning_rate = LEARNING_RATES_BY_BATCH_SIZE[global_batch_size]
# + [markdown] colab_type="text" id="z1Muy0gDZwO5"
# ### 현재 어떤 것이 지원됩니까?
#
# TF 2.0 베타 버전에서는 케라스와 함께 `MirroredStrategy`와 `CentralStorageStrategy`, `MultiWorkerMirroredStrategy`를 사용하여 훈련할 수 있습니다. `CentralStorageStrategy`와 `MultiWorkerMirroredStrategy`는 아직 실험 기능이므로 추후 바뀔 수 있습니다.
# 다른 전략도 조만간 지원될 것입니다. API와 사용 방법은 위에 설명한 것과 동일할 것입니다.
#
# | 훈련 API | MirroredStrategy | TPUStrategy | MultiWorkerMirroredStrategy | CentralStorageStrategy | ParameterServerStrategy |
# |---------------- |--------------------- |----------------------- |----------------------------------- |----------------------------------- |--------------------------- |
# | Keras API | 지원 | 2.0 RC 지원 예정 | 실험 기능으로 지원 | 실험 기능으로 지원 | 2.0 RC 지원 예정 |
#
# ### 예제와 튜토리얼
#
# 위에서 설명한 케라스 분산 훈련 방법에 대한 튜토리얼과 예제들의 목록입니다.
#
# 1. `MirroredStrategy`를 사용한 [MNIST](../tutorials/distribute/keras.ipynb) 훈련 튜토리얼.
# 2. ImageNet 데이터와 `MirroredStrategy`를 사용한 공식 [ResNet50](https://github.com/tensorflow/models/blob/master/official/vision/image_classification/resnet_imagenet_main.py) 훈련.
# 3. 클라우드 TPU에서 ImageNet 데이터와 `TPUStrategy`를 사용한 [ResNet50](https://github.com/tensorflow/tpu/blob/master/models/experimental/resnet50_keras/resnet50.py) 훈련. 이 예제는 현재 텐서플로 1.x 버전에서만 동작합니다.
# 4. `MultiWorkerMirroredStrategy`를 사용한 [MNIST](../tutorials/distribute/multi_worker_with_keras.ipynb) 훈련 튜토리얼.
# 5. `MirroredStrategy`를 사용한 [NCF](https://github.com/tensorflow/models/blob/master/official/recommendation/ncf_keras_main.py) 훈련.
# 6. `MirroredStrategy`를 사용한 [Transformer](https://github.com/tensorflow/models/blob/master/official/nlp/transformer/transformer_main.py) 훈련.
# + [markdown] colab_type="text" id="IlYVC0goepdk"
# ## 사용자 정의 훈련 루프와 함께 `tf.distribute.Strategy` 사용하기
# 지금까지 살펴본 것처럼 고수준 API와 함께 `tf.distribute.Strategy`를 사용하려면 코드 몇 줄만 바꾸면 되었습니다. 조금만 더 노력을 들이면 이런 프레임워크를 사용하지 않는 사용자도 `tf.distribute.Strategy`를 사용할 수 있습니다.
#
# 텐서플로는 다양한 용도로 사용됩니다. 연구자들 같은 일부 사용자들은 더 높은 자유도와 훈련 루프에 대한 제어를 원합니다. 이 때문에 추정기나 케라스 같은 고수준 API를 사용하기 힘든 경우가 있습니다. 예를 들어, GAN을 사용하는데 매번 생성자(generator)와 판별자(discriminator) 단계의 수를 바꾸고 싶을 수 있습니다. 비슷하게, 고수준 API는 강화 학습(Reinforcement learning)에는 그다지 적절하지 않습니다. 그래서 이런 사용자들은 보통 자신만의 훈련 루프를 작성하게 됩니다.
#
# 이 사용자들을 위하여, `tf.distribute.Strategy` 클래스들은 일련의 주요 메서드들을 제공합니다. 이 메서드들을 사용하려면 처음에는 코드를 이리저리 조금 옮겨야 할 수 있겠지만, 한번 작업해 놓으면 전략 인스턴스만 바꿔서 GPU, TPU, 여러 장비로 쉽게 바꿔가며 훈련을 할 수 있습니다.
#
# 앞에서 살펴본 케라스 모델을 사용한 훈련 예제를 통하여 사용하는 모습을 간단하게 살펴보겠습니다.
# + [markdown] colab_type="text" id="XNHvSY32nVBi"
# 먼저, 전략의 범위(scope) 안에서 모델과 옵티마이저를 만듭니다. 이는 모델이나 옵티마이저로 만들어진 변수가 미러링 되도록 만듭니다.
# + colab={} colab_type="code" id="W-3Bn-CaiPKD"
with mirrored_strategy.scope():
model = tf.keras.Sequential([tf.keras.layers.Dense(1, input_shape=(1,))])
optimizer = tf.keras.optimizers.SGD()
# + [markdown] colab_type="text" id="mYkAyPeYnlXk"
# 다음으로는 입력 데이터셋을 만든 다음, `tf.distribute.Strategy.experimental_distribute_dataset` 메서드를 호출하여 전략에 맞게 데이터셋을 분배합니다.
# + colab={} colab_type="code" id="94BkvkLInkKd"
with mirrored_strategy.scope():
dataset = tf.data.Dataset.from_tensors(([1.], [1.])).repeat(1000).batch(
global_batch_size)
dist_dataset = mirrored_strategy.experimental_distribute_dataset(dataset)
# + [markdown] colab_type="text" id="grzmTlSvn2j8"
# 그리고 나서는 한 단계의 훈련을 정의합니다. 그래디언트를 계산하기 위해 `tf.GradientTape`를 사용합니다. 이 그래디언트를 적용하여 우리 모델의 변수를 갱신하기 위해서는 옵티마이저를 사용합니다. 분산 훈련을 위하여 이 훈련 작업을 `step_fn` 함수 안에 구현합니다. 그리고 `step_fn`을 앞에서 만든 `dist_dataset`에서 얻은 입력 데이터와 함께 `tf.distrbute.Strategy.experimental_run_v2`메서드로 전달합니다.
# + colab={} colab_type="code" id="NJxL5YrVniDe"
@tf.function
def train_step(dist_inputs):
def step_fn(inputs):
features, labels = inputs
with tf.GradientTape() as tape:
logits = model(features)
cross_entropy = tf.nn.softmax_cross_entropy_with_logits(
logits=logits, labels=labels)
loss = tf.reduce_sum(cross_entropy) * (1.0 / global_batch_size)
grads = tape.gradient(loss, model.trainable_variables)
optimizer.apply_gradients(list(zip(grads, model.trainable_variables)))
return cross_entropy
per_example_losses = mirrored_strategy.experimental_run_v2(
step_fn, args=(dist_inputs,))
mean_loss = mirrored_strategy.reduce(
tf.distribute.ReduceOp.MEAN, per_example_losses, axis=0)
return mean_loss
# + [markdown] colab_type="text" id="yRL5u_NLoTvq"
# 위 코드에서 몇 가지 더 짚어볼 점이 있습니다.
#
# 1. 손실(loss)을 계산하기 위하여 `tf.nn.softmax_cross_entropy_with_logits`를 사용하였습니다. 그리고 손실의 합을 전체 배치 크기로 나누는 부분이 중요합니다. 이는 모든 복제된 훈련이 동시에 이루어지고 있고, 각 단계에 훈련이 이루어지는 입력의 수는 전체 배치 크기와 같기 때문입니다. 따라서 손실 값은 각 복제된 작업 내의 배치 크기가 아니라 전체 배치 크기로 나누어야 맞습니다.
# 2. `tf.distribute.Strategy.experimental_run_v2`에서 반환된 결과를 모으기 위하여 `tf.distribute.Strategy.reduce` API를 사용하였습니다. `tf.distribute.Strategy.experimental_run_v2`는 전략의 각 복제본에서 얻은 결과를 반환합니다. 그리고 이 결과를 사용하는 방법은 여러 가지가 있습니다. 종합한 결과를 얻기 위하여 `reduce` 함수를 사용할 수 있습니다. `tf.distribute.Strategy.experimental_local_results` 메서드로 각 복제본에서 얻은 결과의 값들 목록을 얻을 수도 있습니다.
# 3. 분산 전략 범위 안에서 `apply_gradients` 메서드가 호출되면, 평소와는 동작이 다릅니다. 구체적으로는 동기화된 훈련 중 병렬화된 각 작업에서 그래디언트를 적용하기 전에, 모든 복제본의 그래디언트를 더해집니다.
# + [markdown] colab_type="text" id="o9k_6-6vpQ-P"
# 훈련 단계를 정의했으므로, 마지막으로는 `dist_dataset`에 대하여 훈련을 반복합니다.
# + colab={} colab_type="code" id="Egq9eufToRf6"
with mirrored_strategy.scope():
for inputs in dist_dataset:
print(train_step(inputs))
# + [markdown] colab_type="text" id="jK8eQXF_q1Zs"
# 위 예에서는 `dist_dataset`을 차례대로 처리하며 훈련 입력 데이터를 얻었습니다. `tf.distribute.Strategy.make_experimental_numpy_dataset`를 사용하면 넘파이 입력도 쓸 수 있습니다. `tf.distribute.Strategy.experimental_distribute_dataset` 함수를 호출하기 전에 이 API로 데이터셋을 만들면 됩니다.
#
# 데이터를 차례대로 처리하는 또 다른 방법은 명시적으로 반복자(iterator)를 사용하는 것입니다. 전체 데이터를 모두 사용하지 않고, 정해진 횟수만큼만 훈련을 하고 싶을 때 유용합니다. 반복자를 만들고 명시적으로 `next`를 호출하여 다음 입력 데이터를 얻도록 하면 됩니다. 위 루프 코드를 바꿔보면 다음과 같습니다.
# + colab={} colab_type="code" id="e5BEvR0-LJAc"
with mirrored_strategy.scope():
iterator = iter(dist_dataset)
for _ in range(10):
print(train_step(next(iterator)))
# + [markdown] colab_type="text" id="vDJO8mnypqBA"
# `tf.distribute.Strategy` API를 사용하여 사용자 정의 훈련 루프를 분산 처리 하는 가장 단순한 경우를 살펴보았습니다. 현재 API를 개선하는 과정 중에 있습니다. 이 API를 사용하려면 사용자 쪽에서 꽤 많은 작업을 해야 하므로, 나중에 별도의 더 자세한 가이드로 설명하도록 하겠습니다.
# + [markdown] colab_type="text" id="BZjNwCt1qBdw"
# ### 현재 어떤 것이 지원됩니까?
# TF 2.0 베타 버전에서는 사용자 정의 훈련 루프와 함께 위에서 설명한 `MirroredStrategy`, 그리고 `TPUStrategy`를 사용할 수 있습니다. 또한 `MultiWorkerMirorredStrategy`도 추후 지원될 것입니다.
#
# | 훈련 API | MirroredStrategy | TPUStrategy | MultiWorkerMirroredStrategy | CentralStorageStrategy | ParameterServerStrategy |
# |:----------------------- |:------------------- |:------------------- |:----------------------------- |:------------------------ |:------------------------- |
# | 사용자 정의 훈련 루프 | 지원 | 지원 | 2.0 RC 지원 예정 | 2.0 RC 지원 예정 | 아직 미지원 |
#
# ### 예제와 튜토리얼
# 사용자 정의 훈련 루프와 함께 분산 전략을 사용하는 예제들입니다.
#
# 1. `MirroredStrategy`로 MNIST를 훈련하는 [튜토리얼](../tutorials/distribute/training_loops.ipynb).
# 2. `MirroredStrategy`를 사용하는 [DenseNet](https://github.com/tensorflow/examples/blob/master/tensorflow_examples/models/densenet/distributed_train.py) 예제.
# 3. `MirroredStrategy`와 `TPUStrategy`를 사용하여 훈련하는 [BERT](https://github.com/tensorflow/models/blob/master/official/bert/run_classifier.py) 예제.
# 이 예제는 분산 훈련 도중 체크포인트로부터 불러오거나 주기적인 체크포인트를 만드는 방법을 이해하는 데 매우 유용합니다.
# 4. `keras_use_ctl` 플래그를 켜서 활성화할 수 있는 `MirroredStrategy`로 훈련한 [NCF](https://github.com/tensorflow/models/blob/master/official/recommendation/ncf_keras_main.py) 예제.
# 5. `MirroredStrategy`를 사용하여 훈련하는 [NMT](https://github.com/tensorflow/examples/blob/master/tensorflow_examples/models/nmt_with_attention/distributed_train.py) 예제.
# + [markdown] colab_type="text" id="nO0hmFCRoIll"
# ## 추정기(Estimator)와 함께 `tf.distribute.Strategy` 사용하기
# `tf.estimator`는 원래부터 비동기 파라미터 서버 방식을 지원했던 분산 훈련 텐서플로 API입니다. 케라스와 마찬가지로 `tf.distribute.Strategy`를 `tf.estimator`와 함께 쓸 수 있습니다. 추정기 사용자는 아주 조금만 코드를 변경하면, 훈련이 분산되는 방식을 쉽게 바꿀 수 있습니다. 따라서 이제는 추정기 사용자들도 다중 GPU나 다중 워커뿐 아니라 다중 TPU에서 동기 방식으로 분산 훈련을 할 수 있습니다. 하지만 추정기는 제한적으로 지원하는 것입니다. 자세한 내용은 아래 [현재 어떤 것이 지원됩니까?](#estimator_support) 부분을 참고하십시오.
#
# 추정기와 함께 `tf.distribute.Strategy`를 사용하는 방법은 케라스와는 살짝 다릅니다. `strategy.scope`를 사용하는 대신에, 전략 객체를 추정기의 [`RunConfig`](https://www.tensorflow.org/api_docs/python/tf/estimator/RunConfig)(실행 설정)에 넣어서 전달해야합니다.
#
# 다음은 기본으로 제공되는 `LinearRegressor`와 `MirroredStrategy`를 함께 사용하는 방법을 보여주는 코드입니다.
# + colab={} colab_type="code" id="oGFY5nW_B3YU"
mirrored_strategy = tf.distribute.MirroredStrategy()
config = tf.estimator.RunConfig(
train_distribute=mirrored_strategy, eval_distribute=mirrored_strategy)
regressor = tf.estimator.LinearRegressor(
feature_columns=[tf.feature_column.numeric_column('feats')],
optimizer='SGD',
config=config)
# + [markdown] colab_type="text" id="n6eSfLN5RGY8"
# 위 예제에서는 기본으로 제공되는 추정기를 사용하였지만, 직접 만든 추정기도 동일한 코드로 사용할 수 있습니다. `train_distribute`가 훈련을 어떻게 분산시킬지를 지정하고, `eval_distribute`가 평가를 어떻게 분산시킬지를 지정합니다. 케라스와 함께 사용할 때 훈련과 평가에 동일한 분산 전략을 사용했던 것과는 차이가 있습니다.
#
# 다음과 같이 입력 함수를 지정하면 추정기의 훈련과 평가를 할 수 있습니다.
# + colab={} colab_type="code" id="2ky2ve2PB3YP"
def input_fn():
dataset = tf.data.Dataset.from_tensors(({"feats":[1.]}, [1.]))
return dataset.repeat(1000).batch(10)
regressor.train(input_fn=input_fn, steps=10)
regressor.evaluate(input_fn=input_fn, steps=10)
# + [markdown] colab_type="text" id="hgaU9xQSSk2x"
# 추정기와 케라스의 또 다른 점인 입력 처리 방식을 살펴봅시다. 케라스에서는 각 배치의 데이터가 여러 개의 복제된 작업으로 나누어진다고 설명했습니다. 하지만 추정기에서는 사용자가 `input_fn` 입력 함수를 제공하고, 데이터를 워커나 장비들에 어떻게 나누어 처리할지를 온전히 제어할 수 있습니다. 텐서플로는 배치의 데이터를 자동으로 나누지도 않고, 각 워커에 자동으로 분배하지도 않습니다. 제공된 `input_fn` 함수는 워커마다 한 번씩 호출됩니다. 따라서 워커마다 데이터셋을 받게 됩니다. 한 데이터셋의 배치 하나가 워커의 복제된 작업 하나에 들어가고, 따라서 워커 하나에 N개의 복제된 작업이 있으면 N개의 배치가 수행됩니다. 다시 말하자면 `input_fn`이 반환하는 데이터셋은 `PER_REPLICA_BATCH_SIZE` 즉 복제 작업 하나가 배치 하나에서 처리할 크기여야 합니다. 한 단계에서 처리하는 전체 배치 크기는 `PER_REPLICA_BATCH_SIZE * strategy.num_replicas_in_sync`가 됩니다. 다중 워커를 사용하여 훈련할 때는 데이터를 워커별로 쪼개거나, 아니면 각자 다른 임의의 순서로 섞는 것이 좋을 수도 있습니다. 이렇게 처리하는 예제는 [추정기로 다중 워커를 써서 훈련하기](../tutorials/distribute/multi_worker_with_estimator.ipynb)에서 볼 수 있습니다.
# + [markdown] colab_type="text" id="_098zB3vVhuV"
# 추정기와 함께 `MirroredStrategy`를 사용하는 예를 보았습니다. `TPUStrategy`도 같은 방법으로 추정기와 함께 사용할 수 있습니다.
# ```
# config = tf.estimator.RunConfig(
# train_distribute=tpu_strategy, eval_distribute=tpu_strategy)
# ```
# + [markdown] colab_type="text" id="G3ieQKfWZhhL"
# 비슷하게 다중 워커나 파라미터 서버를 사용한 전략도 사용할 수 있습니다. 코드는 거의 같지만, `tf.estimator.train_and_evaluate`를 사용해야 합니다. 그리고 클러스터에서 프로그램을 실행할 때 "TF_CONFIG" 환경변수를 설정해야 합니다.
# + [markdown] colab_type="text" id="A_lvUsSLZzVg"
# ### 현재 어떤 것이 지원됩니까?
#
# TF 2.0 베타 버전에서는 추정기와 함께 모든 전략을 제한적으로 지원합니다. 기본적인 훈련과 평가는 동작합니다. 하지만 스캐폴드(scaffold) 같은 고급 기능은 아직 동작하지 않습니다. 또한 다소 버그가 있을 수 있습니다. 현재로써는 추정기와 함께 사용하는 것을 활발히 개선할 계획은 없습니다. 대신 케라스나 사용자 정의 훈련 루프 지원에 집중할 계획입니다. 만약 가능하다면 `tf.distribute` 사용시 이 API들을 먼저 고려하여 주십시오.
#
# | 훈련 API | MirroredStrategy | TPUStrategy | MultiWorkerMirroredStrategy | CentralStorageStrategy | ParameterServerStrategy |
# |:--------------- |:------------------ |:------------- |:----------------------------- |:------------------------ |:------------------------- |
# | 추정기 API | 제한적으로 지원 | 제한적으로 지원 | 제한적으로 지원 | 제한적으로 지원 | 제한적으로 지원 |
#
# ### 예제와 튜토리얼
# 다음은 추정기와 함께 다양한 전략을 사용하는 방법을 처음부터 끝까지 보여주는 예제들입니다.
#
# 1. [추정기로 다중 워커를 써서 훈련하기](../tutorials/distribute/multi_worker_with_estimator.ipynb)에서는 `MultiWorkerMirroredStrategy`로 다중 워커를 써서 MNIST를 훈련합니다.
# 2. [처음부터 끝까지 살펴보는 예제](https://github.com/tensorflow/ecosystem/tree/master/distribution_strategy)에서는 tensorflow/ecosystem의 쿠버네티스(Kubernetes) 템플릿을 이용하여 다중 워커를 사용하여 훈련합니다. 이 예제에서는 케라스 모델로 시작해서 `tf.keras.estimator.model_to_estimator` API를 이용하여 추정기 모델로 변환합니다.
# 3. `MirroredStrategy`나 `MultiWorkerMirroredStrategy`로 훈련할 수 있는 공식 [ResNet50](https://github.com/tensorflow/models/blob/master/official/r1/resnet/imagenet_main.py) 모델.
# 4. `TPUStrategy`를 사용한 [ResNet50](https://github.com/tensorflow/tpu/blob/master/models/experimental/distribution_strategy/resnet_estimator.py) 예제.
# + [markdown] colab_type="text" id="Xk0JdsTHyUnE"
# ## 그 밖의 주제
# 이번 절에서는 다양한 사용 방식에 관련한 몇 가지 주제들을 다룹니다.
# + [markdown] colab_type="text" id="cP6BUIBtudRk"
# <a id="TF_CONFIG">
# ### TF\_CONFIG 환경변수 설정하기
# </a>
#
# 다중 워커를 사용하여 훈련할 때는, 앞서 설명했듯이 클러스터의 각 실행 프로그램마다 "TF\_CONFIG" 환경변수를 설정해야합니다. "TF\_CONFIG" 환경변수는 JSON 형식입니다. 그 안에는 클러스터를 구성하는 작업과 작업의 주소 및 각 작업의 역할을 기술합니다. [tensorflow/ecosystem](https://github.com/tensorflow/ecosystem) 저장소에서 훈련 작업에 맞게 "TF\_CONFIG"를 설정하는 쿠버네티스(Kubernetes) 템플릿을 제공합니다.
#
# "TF\_CONFIG" 예를 하나 보면 다음과 같습니다.
# ```
# os.environ["TF_CONFIG"] = json.dumps({
# "cluster": {
# "worker": ["host1:port", "host2:port", "host3:port"],
# "ps": ["host4:port", "host5:port"]
# },
# "task": {"type": "worker", "index": 1}
# })
# ```
#
# + [markdown] colab_type="text" id="fezd3aF8wj9r"
# 이 "TF\_CONFIG"는 세 개의 워커와 두 개의 파라미터 서버(ps) 작업을 각각의 호스트 및 포트와 함께 지정하고 있습니다. "task" 부분은 클러스터 내에서 현재 작업이 담당한 역할을 지정합니다. 여기서는 워커(worker) 1번, 즉 두 번째 워커라는 뜻입니다. 클러스터 내에서 가질 수 있는 역할은 "chief"(지휘자), "worker"(워커), "ps"(파라미터 서버), "evaluator"(평가자) 중 하나입니다. 단, "ps" 역할은 `tf.distribute.experimental.ParameterServerStrategy` 전략을 사용할 때만 쓸 수 있습니다.
# + [markdown] colab_type="text" id="GXIbqSW-sFVg"
# ## 다음으로는...
#
# `tf.distribute.Strategy`는 활발하게 개발 중입니다. 한 번 써보시고 [깃허브 이슈](https://github.com/tensorflow/tensorflow/issues/new)를 통하여 피드백을 주시면 감사하겠습니다.
| site/ko/guide/distributed_training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1> Simple Single Layer RNN with Monthly dataset</h1>
import os
import numpy as np
import math
import pandas as pd
import seaborn as sns
import tensorflow as tf
import matplotlib.pyplot as plt
from tensorflow.keras.optimizers import SGD
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, LSTM, Dropout, GRU, SimpleRNN
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score
from sklearn.preprocessing import MinMaxScaler
from sklearn.metrics import mean_absolute_error
from sklearn.metrics import mean_squared_error
plt.style.use('fivethirtyeight')
# salmon_data = pd.read_csv(r"/Users/ismaelcastro/Documents/Computer Science/CS Classes/CS230/project/data.csv")
# salmon_data.head()
# salmon_copy = salmon_data # Create a copy for us to work with
def load_data(pathname):
salmon_data = pd.read_csv(pathname)
salmon_data.head()
salmon_copy = salmon_data
salmon_copy.rename(columns = {"mo": "month", "da" : "day", "fc" : "king"}, inplace = True)
salmon_copy['date']=pd.to_datetime(salmon_copy[['year','month','day']])
king_data = salmon_copy.filter(["date","king"], axis=1)
king_greater = king_data['date'].apply(pd.Timestamp) >= pd.Timestamp('01/01/1939')
greater_than = king_data[king_greater]
king_all = greater_than[greater_than['date'].apply(pd.Timestamp) <= pd.Timestamp('12/31/2020')]
king_all_copy = king_all
king_all_copy = king_all_copy.reset_index()
king_all_copy = king_all_copy.drop('index', axis=1)
return king_all_copy, king_data
chris_path = '/Users/chrisshell/Desktop/Stanford/SalmonData/Use Data/passBonCS.csv'
ismael_path = '/Users/ismaelcastro/Documents/Computer Science/CS Classes/CS230/project/data.csv'
abdul_path = '/Users/abdul/Downloads/SalmonNet/passBonCS.csv'
king_all_copy, king_data= load_data(ismael_path)
print(king_all_copy)
data_copy = king_all_copy
data_copy['date']
data_copy.set_index('date', inplace=True)
data_copy.index = pd.to_datetime(data_copy.index)
data_copy = data_copy.resample('1M').sum()
data_copy
print(data_copy)
data_copy.shape
data_copy.reset_index(inplace=True)
data_copy = data_copy.rename(columns = {'index':'date'})
print(data_copy)
def create_train_test(king_all):
king_training_parse = king_all['date'].apply(pd.Timestamp) <= pd.Timestamp('12/31/2015')
king_training = king_all[king_training_parse]
king_training = king_training.reset_index()
king_training = king_training.drop('index', axis=1)
king_test_parse = king_all['date'].apply(pd.Timestamp) > pd.Timestamp('12/31/2015')
king_test = king_all[king_test_parse]
king_test = king_test.reset_index()
king_test = king_test.drop('index', axis=1)
print(king_test.shape)
# Normalizing Data
king_training[king_training["king"] < 0] = 0
king_test[king_test["king"] < 0] = 0
king_train_pre = king_training["king"].to_frame()
king_test_pre = king_test["king"].to_frame()
scaler = MinMaxScaler(feature_range=(0, 1))
king_train_norm = scaler.fit_transform(king_train_pre)
king_test_norm = scaler.fit_transform(king_test_pre)
x_train = []
y_train = []
x_test = []
y_test = []
y_test_not_norm = []
y_train_not_norm = []
for i in range(6,924): # 30
x_train.append(king_train_norm[i-6:i])
y_train.append(king_train_norm[i])
for i in range(6, 60):
x_test.append(king_test_norm[i-6:i])
y_test.append(king_test_norm[i])
# make y_test_not_norm
for i in range(6, 60):
y_test_not_norm.append(king_test['king'][i])
for i in range(6,924): # 30
y_train_not_norm.append(king_training['king'][i])
return x_train, y_train, x_test, y_test, scaler, y_test_not_norm, y_train_not_norm
x_train, y_train, x_test, y_test, scaler, y_test_not_norm, y_train_not_norm = create_train_test(data_copy)
x_train = np.array(x_train)
x_test = np.array(x_test)
x_train = np.reshape(x_train, (x_train.shape[0],x_train.shape[1],1)).astype(np.float32)
x_test = np.reshape(x_test, (x_test.shape[0],x_test.shape[1],1))
y_train = np.array(y_train)
y_test = np.array(y_test)
y_test_not_norm = np.array(y_test_not_norm)
print(y_test.shape)
y_test_not_norm = y_test_not_norm.reshape((y_test_not_norm.shape[0], 1))
print(y_test_not_norm.shape)
y_train_not_norm = np.array(y_train_not_norm)
y_train_not_norm = y_train_not_norm.reshape((y_train_not_norm.shape[0], 1))
print(y_train_not_norm.shape)
print(y_train.shape)
# +
def plot_predictions(test,predicted):
plt.plot(test, color='red',label='Real Chinook Count')
plt.plot(predicted, color='blue',label='Predicted Chinook Count')
plt.title('Chinook Population Prediction')
plt.xlabel('Time')
plt.ylabel('Chinook Count')
plt.legend()
plt.show()
def plot_loss(history):
plt.plot(history.history['loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.show()
def return_rmse(test, predicted):
rmse = math.sqrt(mean_squared_error(test, predicted))
print("The root mean squared error is {}.".format(rmse))
def month_to_year(month_preds):
month_preds = month_preds[5:]
print(len(month_preds))
year_preds = []
for i in range(12, len(month_preds), 12):
salmon_count = np.sum(month_preds[i - 12:i])
year_preds.append(salmon_count)
year_preds = pd.DataFrame(year_preds, columns = ["Count"])
return year_preds
# -
def create_single_layer_rnn_model(x_train, y_train, x_test, y_test, scaler):
'''
create single layer rnn model trained on x_train and y_train
and make predictions on the x_test data
'''
# create a model
model = Sequential()
model.add(SimpleRNN(32))
#model.add(SimpleRNN(32, return_sequences=True))
#model.add(SimpleRNN(32, return_sequences=True))
#model.add(SimpleRNN(1))
model.add(Dense(1))
model.compile(optimizer='adam', loss='mean_squared_error')
# fit the RNN model
history = model.fit(x_train, y_train, epochs=300, batch_size=64)
print("predicting")
# Finalizing predictions
RNN_train_preds = model.predict(x_train)
RNN_test_preds = model.predict(x_test)
#Descale
RNN_train_preds = scaler.inverse_transform(RNN_train_preds)
y_train = scaler.inverse_transform(y_train)
RNN_test_preds = scaler.inverse_transform(RNN_test_preds)
RNN_test_preds = RNN_test_preds.astype(np.int64)
y_test = scaler.inverse_transform(y_test)
return model, RNN_train_preds, RNN_test_preds, history, y_train, y_test
model, RNN_train_preds, RNN_test_preds, history_RNN, y_train, y_test = create_single_layer_rnn_model(x_train, y_train, x_test, y_test, scaler)
# plot single_layer_rnn_model
plot_predictions(y_train, RNN_train_preds)
return_rmse(y_train, RNN_train_preds)
print(RNN_train_preds.shape)
plot_predictions(y_test, RNN_test_preds)
return_rmse(y_test, RNN_test_preds)
plot_loss(history_RNN)
# global var for baseline
y_test_year = month_to_year(y_test)
len(y_test)
len(y_test_year)
y_test_year = month_to_year(y_test)
bs_chris_path = '/Users/chrisshell/Desktop/Stanford/SalmonData/Use Data/Forecast Data Update.csv'
bs_ismael_path = '/Users/ismaelcastro/Documents/Computer Science/CS Classes/CS230/project/forecast_data_17_20.csv'
bs_abdul_path = '/Users/abdul/Downloads/SalmonNet/Forecast Data Update.csv'
baseline_data = pd.read_csv(bs_ismael_path)
traditional = pd.DataFrame(baseline_data["Count"])
print(traditional)
y_test_year = y_test_year.astype(np.int64)
print(y_test_year)
# print(GRU_test_year)
RNN_test_year = month_to_year(RNN_test_preds)
RNN_test_year
# test RMSE with baseline and RNN
return_rmse(y_test_year, traditional)
return_rmse(y_test_year, RNN_test_year)
| .ipynb_checkpoints/monthly_robust_rnn-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # QNN on Pynq
#
# This notebook covers how to use low quantized neural networks on Pynq.
# It shows an example of webcam based Cifar-10 recognition using CNV network inspired at VGG-16, featuring 6 convolutional layers, 3 max pool layers and 3 fully connected layers. There are 3 different precision available:
#
# - CNVW1A1 using 1 bit weights and 1 bit activation,
# - CNVW1A2 using 1 bit weights and 2 bit activation and
# - CNVW2A2 using 2 bit weights and 2 activation
#
# All of them can be performed in pure software and hardware accelerated environment.
# In order to reproduce this notebook, you will need an external USB Camera connected to the PYNQ Board.
import bnn
# ## 1. Load image from the camera
#
# The image is captured from the external USB camera and shown:
# +
import cv2
from PIL import Image as PIL_Image
from PIL import ImageEnhance
from PIL import ImageOps
# says we capture an image from a webcam
cap = cv2.VideoCapture(0)
_ , cv2_im = cap.read()
cv2_im = cv2.cvtColor(cv2_im,cv2.COLOR_BGR2RGB)
img = PIL_Image.fromarray(cv2_im)
#original captured image
#orig_img_path = '/home/xilinx/jupyter_notebooks/bnn/pictures/webcam_cifar-10.jpg'
#img = PIL_Image.open(orig_img_path)
# The enhancement values (contrast and brightness) depend on backgroud, external lights etc
bright = ImageEnhance.Brightness(img)
img = bright.enhance(0.95)
img
# -
# ## 2. Hardware Inference
#
# The inference can be performed with different precision for weights and activation. Creating a specific Classifier will automatically download the correct bitstream onto PL and load the specified parameters.
# ### Case 1:
#
# #### W1A1 - 1 bit weights and 1 activation
hw_classifier = bnn.CnvClassifier(bnn.NETWORK_CNVW1A1,"cifar10",bnn.RUNTIME_HW)
class_ranksW1A1=hw_classifier.classify_image_details(img)
inferred_class=class_ranksW1A1.argmax()
print("Inferred class: {0}".format(inferred_class))
print("Class name: {0}".format(hw_classifier.class_name(inferred_class)))
# ### Case 2:
#
# #### W1A2 - 1 bit weight and 2 activation
hw_classifier = bnn.CnvClassifier(bnn.NETWORK_CNVW1A2,"cifar10",bnn.RUNTIME_HW)
class_ranksW1A2=hw_classifier.classify_image_details(img)
inferred_class=class_ranksW1A2.argmax()
print("Inferred class: {0}".format(inferred_class))
print("Class name: {0}".format(hw_classifier.class_name(inferred_class)))
# ### Case 3:
#
# #### W2A2 - 2 bit weights and 2 activation
hw_classifier = bnn.CnvClassifier(bnn.NETWORK_CNVW2A2,"cifar10",bnn.RUNTIME_HW)
class_ranksW2A2=hw_classifier.classify_image_details(img)
inferred_class=class_ranksW2A2.argmax()
print("Inferred class: {0}".format(inferred_class))
print("Class name: {0}".format(hw_classifier.class_name(inferred_class)))
# ## 3. Summary
#
# ### Rankings
#
# The rankings can be visualized using `matplotlib`:
#
# #### W1A1:
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
x_pos = np.arange(len(class_ranksW1A1))
fig, ax = plt.subplots()
ax.bar(x_pos, (class_ranksW1A1), 0.7)
ax.set_xticklabels(hw_classifier.classes, rotation='vertical')
ax.set_xticks(x_pos)
ax.set
plt.show()
# -
# #### W1A2:
#
x_pos = np.arange(len(class_ranksW1A2))
fig, ax = plt.subplots()
ax.bar(x_pos, (class_ranksW1A2), 0.7)
ax.set_xticklabels(hw_classifier.classes, rotation='vertical')
ax.set_xticks(x_pos)
ax.set
plt.show()
# #### W2A2:
x_pos = np.arange(len(class_ranksW2A2))
fig, ax = plt.subplots()
ax.bar(x_pos, (class_ranksW2A2), 0.7)
ax.set_xticklabels(hw_classifier.classes, rotation='vertical')
ax.set_xticks(x_pos)
ax.set
plt.show()
# ## 4. Reset the device
#
# +
from pynq import Xlnk
xlnk = Xlnk()
xlnk.xlnk_reset()
| notebooks/CNV-QNN_Cifar10_Webcam.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="ON4WbEBvk9Z9"
# # Multivariate Linear Regression
# ## Predicting Healthcare COVID Insurance Price from age, race, ethnicity, gender, latitude, longitude
# + [markdown] id="RKxoZ5lknWll"
# ### Import Libraries
# + id="Lq6qDjhHnbak"
# %matplotlib inline
from mpl_toolkits.mplot3d import Axes3D
import numpy as np
import matplotlib.pyplot as plt
from sklearn.preprocessing import normalize
# + [markdown] id="W5b_-H-zn3_O"
# ### Read Data
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="2y7aFAEzmcRX" outputId="21d09e32-aac7-4797-ca64-cc958ee8d751"
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
# %matplotlib inline
#importing dataset using panda
#dataset
data = pd.read_csv('https://storage.googleapis.com/dataprep-staging-b4c6b8ff-9afc-4b23-a0d8-d480526baaa4/yz1268%40nyu.edu/jobrun/Untitled%20recipe%20%E2%80%93%204.csv/2021-08-16_23-54-42_00000000')
#to see what my dataset is comprised of
data.head()
#healthcare_coverage,age,race, ethnicity, gender, latitude, longitude,healthcare expense.
#the first column is y, the rest are all x1, x2,...xn.
# + id="Ip2EQ_PGoQJO" colab={"base_uri": "https://localhost:8080/"} outputId="4e1f87dd-4fa4-4a84-e623-7081f5e5025d"
data.shape
# + colab={"base_uri": "https://localhost:8080/"} id="N0Ju4Pg3BX_q" outputId="6aafe253-1b42-42b7-cf1b-f293f0505571"
print(type(data))
#print(data.keys())
data = data.to_numpy()
print(type(data))
# + colab={"base_uri": "https://localhost:8080/"} id="WHmV7yfL_PGR" outputId="702c7b09-0fc2-4ecd-b936-53bdc98cad26"
print(data)
# + colab={"base_uri": "https://localhost:8080/"} id="XHJv8viwAwk0" outputId="0ebaf85a-f0e8-4641-afa6-e8f79a3052e9"
data.shape
# + [markdown] id="ydBEcRFeiDtS"
# ### Normalize Data
# + id="y1BwuHufiHJ_"
data = normalize(data, axis=0)
#data = data.values.tolist()
# + colab={"base_uri": "https://localhost:8080/"} id="186dPWxPu2z0" outputId="9aa6e987-9a67-4bcb-9717-79517c1206e0"
#print(data[0])
print('the Y value in regression model',data[:,0:1])
print('the X1 & X2 values in regression model',data[:,1:3])
# + [markdown] id="8rMv-L53odN5"
# ### Seperate Data into X and Y
# + id="uop_AFQSolHl"
X = data[:, 1:6]
#X = data[:, 1:7] The last column is the medical expense, it is not used in the first train.
Y = data[:, 0:1]
#https://www.earthdatascience.org/courses/intro-to-earth-data-science/scientific-data-structures-python/pandas-dataframes/indexing-filtering-data-pandas-dataframes/
# + colab={"base_uri": "https://localhost:8080/"} id="oAY6-vomtQqX" outputId="5f3bae8c-6ca4-4f41-8eb7-a14bb18bde3f"
#X[:, 6]
data[:, ]
# + colab={"base_uri": "https://localhost:8080/"} id="mI-AHQ84o7_o" outputId="7f420d5a-6138-4e4d-9dce-684864caf28b"
X.size
# + colab={"base_uri": "https://localhost:8080/"} id="Zszwk6QIx-Uv" outputId="38b37db2-b7c3-4ef2-a61b-2e0448b92a0a"
print(X)
# + [markdown] id="wPSTKDAXpLAN"
# ### Visualize the Data
# + id="eqSVKtKBWv9-" colab={"base_uri": "https://localhost:8080/", "height": 248} outputId="b64b377d-739b-4b7d-eb04-a371bb7d573d"
# Fixing random state for reproducibility
np.random.seed(19680801)
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
xs = X[:, 0]
ys = X[:, 1]
zs = Y
ax.scatter(xs, ys, zs)
ax.set_xlabel('size')
ax.set_ylabel('bedroom')
ax.set_zlabel('price')
plt.show()
# + [markdown] id="Yrjucdifq_mt"
# ### Hyperparameters
# + id="OwVfkk3RrC_I"
learning_rate = 0.09
max_iteration = 500
# + id="RSXPA9L25UGl"
s_learning_rate = 0.06
s_max_iteration = 500
# + id="IeY-_RqF5V5v"
mb_learning_rate = 0.09
mb_max_iteration = 500
batch_size = 16
# + [markdown] id="EzXaG70OrZtn"
# ### Parameters
# + id="2INJM9AIrcBu"
theta = np.zeros((X.shape[1]+1, 1))
#theta = np.zeros(data.shape[1])
s_theta = np.zeros((X.shape[1]+1, 1))
mb_theta = np.zeros((X.shape[1]+1, 1))
# + colab={"base_uri": "https://localhost:8080/"} id="MsZHYL52l1UK" outputId="36035281-c164-4a2b-d54e-777a620d9f6a"
#np.zeros(data.shape[1])
X.shape[1]
# + colab={"base_uri": "https://localhost:8080/"} id="I8qVGCVpiEjP" outputId="e5e9602d-d56c-4835-ccda-004f0b06616e"
theta.size
# + colab={"base_uri": "https://localhost:8080/"} id="b6U7xvjahnXg" outputId="6613470c-1738-4b71-98a4-45e191d0ad42"
data.shape[0]
# + colab={"base_uri": "https://localhost:8080/"} id="24hMQ0i9iMt3" outputId="31c254c4-8dd4-4d52-f229-18c67c3182b9"
tempX = np.ones((X.shape[0], X.shape[1] + 1))
print(tempX)
# + colab={"base_uri": "https://localhost:8080/"} id="lo0FH10_iOY8" outputId="dfc2657c-b613-4505-a591-e5edcfe04999"
tempX[:,1:].size
# + colab={"base_uri": "https://localhost:8080/"} id="0ybLjmWAi0Wv" outputId="00166c4b-b36f-4329-ab04-1861b0ce131d"
tempX.size
# + colab={"base_uri": "https://localhost:8080/"} id="roBskY-Ziu6U" outputId="08246581-96f0-4cdb-da50-4572969cb32f"
np.matmul(tempX, theta)
# + [markdown] id="FPt2eKjBr1Lx"
# ### Hypothesis
# + id="enVE4tHWsBBF"
def h (theta, X) :
tempX = np.ones((X.shape[0], X.shape[1] + 1))
tempX[:,1:] = X
return np.matmul(tempX, theta)
# + [markdown] id="IA9I6vaXs7nb"
# ### Loss Function
# + id="3KOQVUjBs-Ub"
def loss (theta, X, Y) :
return np.average(np.square(Y - h(theta, X))) / 2
# + [markdown] id="7TE79NG-tyLG"
# ### Calculate Gradients
# + id="yy9yxye4t0BJ"
def gradient (theta, X, Y) :
tempX = np.ones((X.shape[0], X.shape[1] + 1))
tempX[:,1:] = X
d_theta = - np.average((Y - h(theta, X)) * tempX, axis= 0)
d_theta = d_theta.reshape((d_theta.shape[0], 1))
return d_theta
# + [markdown] id="jVo5TbKyu9KL"
# ### Batch Gradient Descent
# + id="fprg4-tcu-4v"
def gradient_descent (theta, X, Y, learning_rate, max_iteration, gap) :
cost = np.zeros(max_iteration)
for i in range(max_iteration) :
d_theta = gradient (theta, X, Y)
theta = theta - learning_rate * d_theta
cost[i] = loss(theta, X, Y)
if i % gap == 0 :
print ('iteration : ', i, ' loss : ', loss(theta, X, Y))
return theta, cost
# + [markdown] id="Si5f2m-9-krz"
# ### Mini-Batch Gradient Descent
# + id="kCGUklWXyov-"
def minibatch_gradient_descent (theta, X, Y, learning_rate, max_iteration, batch_size, gap) :
cost = np.zeros(max_iteration)
for i in range(max_iteration) :
for j in range(0, X.shape[0], batch_size):
d_theta = gradient (theta, X[j:j+batch_size,:], Y[j:j+batch_size,:])
theta = theta - learning_rate * d_theta
cost[i] = loss(theta, X, Y)
if i % gap == 0 :
print ('iteration : ', i, ' loss : ', loss(theta, X, Y))
return theta, cost
# + [markdown] id="epWfjT9R-sk3"
# ### Stochastic Gradient Descent
# + id="cWDNv4BkvpvQ"
def stochastic_gradient_descent (theta, X, Y, learning_rate, max_iteration, gap) :
cost = np.zeros(max_iteration)
for i in range(max_iteration) :
for j in range(X.shape[0]):
d_theta = gradient (theta, X[j,:].reshape(1, X.shape[1]), Y[j,:].reshape(1, 1))
theta = theta - learning_rate * d_theta
cost[i] = loss(theta, X, Y)
if i % gap == 0 :
print ('iteration : ', i, ' loss : ', loss(theta, X, Y))
return theta, cost
# + [markdown] id="uMIZFxUhwQZO"
# ### Train Model
# + id="-0okDGSCwUC7" colab={"base_uri": "https://localhost:8080/"} outputId="31ab4134-e443-4e5e-8a99-d4ddeedf61a3"
theta, cost = gradient_descent (theta, X, Y, learning_rate, max_iteration, 100)
# + id="YBJreRDKwtvI" colab={"base_uri": "https://localhost:8080/"} outputId="ab519634-b3fd-4169-e25d-d17c55ea7af6"
s_theta, s_cost = stochastic_gradient_descent (s_theta, X, Y, s_learning_rate, s_max_iteration, 100)
# + id="viSuvODWza-1" colab={"base_uri": "https://localhost:8080/"} outputId="c372e206-3a48-4bec-f4c7-e9ee8e845bde"
mb_theta, mb_cost = minibatch_gradient_descent (mb_theta, X, Y, mb_learning_rate, mb_max_iteration, batch_size, 100)
# + [markdown] id="MFp2MUlexVd5"
# ### Optimal values of Parameters using Trained Model
# + id="tcMQWkuvwYu7" colab={"base_uri": "https://localhost:8080/"} outputId="b32e2e5b-a0bf-4851-f81d-c663f05aff1a"
theta
# + id="Orub1Wgdx0yc" colab={"base_uri": "https://localhost:8080/"} outputId="fa127ab6-6163-4c53-d748-0a5ab71d4f25"
s_theta
# + id="TBrs3crW55Ka" colab={"base_uri": "https://localhost:8080/"} outputId="e26547d4-3aa8-4471-92dc-946a9054508d"
mb_theta
# + [markdown] id="Pi99RMws736c"
# ### Cost vs Iteration Plots
# + id="SvzRaqIQ78Pl" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="aea61aa3-c541-42af-d249-856f6c71d57e"
#plot the cost
fig, ax = plt.subplots()
ax.plot(np.arange(max_iteration), cost, 'r')
ax.plot(np.arange(max_iteration), s_cost, 'b')
ax.plot(np.arange(max_iteration), mb_cost, 'g')
ax.legend(loc='upper right', labels=['batch gradient descent', 'stochastic gradient descent', 'mini-batch gradient descent'])
ax.set_xlabel('Iterations')
ax.set_ylabel('Cost')
ax.set_title('Error vs. Training Epoch')
plt.show()
# + id="vxLme6Nz6BOd"
| SGD_db_Multivariate_Linear_Regression_Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/bala-codes/Sentiment_Analysis_on_Amazon_Product_Reviews_Using_Machine_and_Deep_Learning/blob/master/codes%20(DL%20-%20RNN%20%26%20LSTM)/Part-3.1%20Sentiment%20Analysis%20-%20Prediction%20-%20Long%20Short%20Term%20Memory%20-%20GLOVE%20EMBEDDING%20-%20Single%20Input%20Edition.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="HyEV8MJe4Ek9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 125} outputId="74f5e3db-0645-4225-e949-cfeb08c1960b"
from google.colab import drive
drive.mount('/content/drive')
# + id="COvo7tDT4FxA" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 152} outputId="5b368e64-eb9f-4c1a-b098-f3291b1a3449"
#Importing the libraries which are required.
import pandas as pd
import nltk
import re
import seaborn as sns
import matplotlib.pyplot as plt
plt.style.use('ggplot')
import numpy as np
import pickle
from nltk.corpus import stopwords
import string
nltk.download('stopwords')
nltk.download('wordnet')
from keras.models import model_from_json
import os
os.environ['TF_CPP_MIN_LOG_LEVEL'] = '3'
import tensorflow as tf
try:
from tensorflow.python.util import module_wrapper as deprecation
except ImportError:
from tensorflow.python.util import deprecation_wrapper as deprecation
deprecation._PER_MODULE_WARNING_LIMIT = 0
from tensorflow.python.util import deprecation
deprecation._PRINT_DEPRECATION_WARNINGS = False
# + id="mVTGMtj54RMW" colab_type="code" colab={}
all_punctuations = string.punctuation + '‘’,:”][],'
def punc_remover(raw_text):
no_punct = "".join([i for i in raw_text if i not in all_punctuations])
return no_punct
'''def stopword_remover(no_punc_text):
words = no_punc_text.split()
no_stp_words = " ".join([i for i in words if i not in stopwords.words('english')])
return no_stp_words'''
lemmer = nltk.stem.WordNetLemmatizer()
def lem(words):
return " ".join([lemmer.lemmatize(word,'v') for word in words.split()])
def text_cleaner(raw):
cleaned_text = (punc_remover(raw.lower()))
return lem(cleaned_text.lower())
def plot_preds1(val1, val2):
class_labels = ['POSITIVE', 'NEGATIVE']
j = [val1, val2]
y_pos = np.arange(len(class_labels))
colors = ['g','r']
plt.barh(y_pos,j, color = colors, alpha = 0.6)
plt.yticks(y_pos,class_labels)
plt.title('PREDICTION FOR BEING POSITIVE VS NEGATIVE SENTIMENTS')
plt.xlabel('Percentage')
plt.ylabel('Labels')
plt.show()
def plot_preds2(val1, val2):
class_labels = ['NEGATIVE','POSITIVE']
j = [val1, val2]
y_pos = np.arange(len(class_labels))
colors = ['g', 'r']
plt.barh(y_pos,j, color = colors, alpha = 0.6)
plt.yticks(y_pos,class_labels)
plt.title('PREDICTION FOR BEING POSITIVE VS NEGATIVE SENTIMENTS')
plt.xlabel('Percentage')
plt.ylabel('Labels')
plt.show()
# + id="ljDmhp8y4c7p" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="01c3f2ce-f478-4ac6-dd3e-36100ba4e89b"
# Use the below function to load the Saved Tokenizer
with open('/content/drive/My Drive/Machine Learning Projects/SENTIMENT ANALYSIS - AMAZON CUSTOMER REVIEWS/SOURCE CODES AND DATASETS/PACKAGE 3 -SOURCE CODES AND FILES/Pretrained Models/Deep Learning Models - With Glove Embedding/LSTM_tokenizer_GLOVE.pkl', 'rb') as handle:
tokenizer = pickle.load(handle)
import tensorflow as tf
from keras.preprocessing.sequence import pad_sequences
maxlen = 300
# nn_emb_model = tf.keras.models.load_model('/content/drive/My Drive/Machine Learning Projects/SENTIMENT ANALYSIS - AMAZON CUSTOMER REVIEWS/SOURCE CODES AND DATASETS/PACKAGE 3 -SOURCE CODES AND FILES/Pretrained Models/Deep Learning Models - With Keras Embeddings/SENTIMENT_ANALYSIS_LSTM_KERAS_EMBED_MODEL.h5')
json_file = open('/content/drive/My Drive/Machine Learning Projects/SENTIMENT ANALYSIS - AMAZON CUSTOMER REVIEWS/SOURCE CODES AND DATASETS/PACKAGE 3 -SOURCE CODES AND FILES/Pretrained Models/Deep Learning Models - With Glove Embedding/LSTM_sentiment_GLOVE.json', 'r')
loaded_model_json = json_file.read()
json_file.close()
loaded_model = model_from_json(loaded_model_json)
# load weights into new model
loaded_model.load_weights("/content/drive/My Drive/Machine Learning Projects/SENTIMENT ANALYSIS - AMAZON CUSTOMER REVIEWS/SOURCE CODES AND DATASETS/PACKAGE 3 -SOURCE CODES AND FILES/Pretrained Models/Deep Learning Models - With Glove Embedding/LSTM_sentiment_GLOVE.h5")
print("Loaded model from disk successfully")
def prediction(x):
process1 = tokenizer.texts_to_sequences(x)
#print("P1",process1)
process2 = pad_sequences(process1, padding='post', maxlen=maxlen)
#print("P3",process2)
prediction = loaded_model.predict(process2)
prediction = np.squeeze(prediction)
val1 = np.squeeze(prediction)
val2 = 1 - np.squeeze(prediction)
value1 = "The Given Review is a Positive Sentiment"
value0 = "The Given Review is a Negative Sentiment"
if prediction >= 0.5:
plot_preds1(val1 , val2)
return value1
else:
plot_preds1(val1 , val2)
return value0
# + id="Eh5EFMEr4Uhm" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 387} outputId="c4900a33-16ce-4dba-f853-946f2d3c1618"
x = input("ENTER THE URL HERE : ")
x = text_cleaner(x)
print(x)
x=[x,]
print("\n\n",prediction(x))
# + id="SwBYBx7-2lyX" colab_type="code" colab={}
| codes (DL - RNN & LSTM)/Part-3.1 Sentiment Analysis - Prediction - Long Short Term Memory - GLOVE EMBEDDING - Single Input Edition.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### This notebook is a submission for the hackathon
#
# https://www.kaggle.com/c/santander-customer-transaction-prediction
import sys
# !{sys.executable} -m pip install --user modAL
import pandas as pd
import numpy as np
from sklearn.ensemble import GradientBoostingClassifier,RandomForestClassifier,ExtraTreesClassifier
#import xgboost as xgb
#import lightgbm as lgbm
import seaborn as sns
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
from sklearn.naive_bayes import GaussianNB
from sklearn import metrics
from sklearn.model_selection import KFold
import time
train = pd.read_csv('train.csv')
test = pd.read_csv('test.csv')
def train_model(model,train,target,test,samples=-1,**params):
kford=KFold(n_splits=5,random_state=2,shuffle=True,)
start_time=time.time()
aucs=[]
test_preds=[]
# for early stopping
# it takes a long time if using all the samples.
if samples<=-1:
samples=train.shape[0]
else:
samples=min(train.shape[0],samples)
print("Started to fit model")
print("fit on {} samples".format(samples))
for ford,(train_idx,val_idx) in enumerate(kford.split(train[:samples],target[:samples])):
print("ford:",ford)
sample_x=train.iloc[train_idx].values
sample_y=target.iloc[train_idx].values
sample_val_x=train.iloc[val_idx].values
sample_val_y=target.iloc[val_idx].values
ford_time=time.time()
model.fit(sample_x,sample_y)
print("epoch cost time {:1}s".format(time.time()-ford_time))
y_pred_prob=model.predict_proba(sample_x)[:,1]
y_val_pred_prob=model.predict_proba(sample_val_x)[:,1]
train_auc=metrics.roc_auc_score(sample_y,y_pred_prob)
val_auc=metrics.roc_auc_score(sample_val_y,y_val_pred_prob)
print("train auc:{:4},val auc:{:4}".format(train_auc,val_auc))
aucs.append([train_auc,val_auc])
test_preds.append(model.predict_proba(test)[:,1])
end_time=time.time()
val_aucs=[auc[1] for auc in aucs]
print("using {} samples,total time:{:1}s,mean val auc:{:4}".format(samples,end_time-start_time,np.mean(val_aucs)))
test_preds=pd.DataFrame(test_preds).T
test_preds.index=test.index
return test_preds
train.shape
target = train['target']
train.info()
train_df = train[0:]
train_df.head(5)
from imblearn.under_sampling import CondensedNearestNeighbour
cnn = CondensedNearestNeighbour(n_jobs=-1)
del train['ID_code']
del train['target']
train.tail(5)
from imblearn.under_sampling import CondensedNearestNeighbour,RandomUnderSampler,NearMiss # doctest: +SKIP
from modAL.models import ActiveLearner
from modAL.uncertainty import uncertainty_sampling
from sklearn.ensemble import RandomForestClassifier
learner = ActiveLearner(
estimator=RandomForestClassifier(),
query_strategy=uncertainty_sampling,
X_training=train[:10000], y_training=target[:10000]
)
import numpy as np
import matplotlib.pyplot as plt
from sklearn.decomposition import PCA
from sklearn.datasets import load_iris
from sklearn.neighbors import KNeighborsClassifier
from modAL.models import ActiveLearner
train.values[0:10000].shape
# initial training data
X_train = train.values[0:10000]
y_train = target.values[0:10000]
# generating the pool
X_pool = np.delete(train.values, (10000, 200), axis=0)
y_pool = np.delete(target.values,(10000, 200))
# visualizing the classes
with plt.style.context('seaborn-white'):
pca = PCA(n_components=2).fit_transform(X_train)
plt.figure(figsize=(7, 7))
plt.scatter(x=pca[:, 0], y=pca[:, 1], c=y_train, cmap='viridis', s=50)
plt.title('The iris dataset')
plt.show()
y_pool.shape
# initializing the active learner
learner = ActiveLearner(
estimator=KNeighborsClassifier(n_neighbors=2),
X_training=X_train, y_training=y_train
)
# +
# visualizing initial prediction
with plt.style.context('seaborn-white'):
plt.figure(figsize=(7, 7))
prediction = learner.predict(X_train)
plt.scatter(x=pca[:, 0], y=pca[:, 1], c=prediction, cmap='viridis', s=10000)
plt.title('Initial accuracy: %f' % learner.score(X_train,y_train))
plt.show()
print('Accuracy before active learning: %f' % learner.score(X_train,y_train))
# -
rus = RandomUnderSampler(random_state=42) # doctest: +SKIP
X_res, y_res = rus.fit_resample(train, target) #doctest: +SKIP
nm = NearMiss(random_state=42) # doctest: +SKIP
X_res_nm, y_res_nm = nm.fit_resample(train, target) #doctest: +SKIP
X_res = pd.DataFrame(X_res,columns=train.columns)
y_res = pd.DataFrame(y_res,columns={'target'})
X_res.shape
test.head(2)
del test['ID_code']
pd.Series(y_res).value_counts()
pd.Series(target).value_counts()
model=RandomForestClassifier()
test_preds=train_model(model,train,target,test,samples=-1)
test_preds_rusample=train_model(model,X_res,y_res,test,samples=-1)
from imblearn.under_sampling import NearMiss
# +
def run(trn_ds, tst_ds, lbr, model, qs, quota):
E_in, E_out = [], []
for _ in range(quota):
# Standard usage of libact objects
ask_id = qs.make_query()
X, _ = zip(*trn_ds.data)
lb = lbr.label(X[ask_id])
trn_ds.update(ask_id, lb)
model.train(trn_ds)
E_in = np.append(E_in, 1 - model.score(trn_ds))
E_out = np.append(E_out, 1 - model.score(tst_ds))
return E_in, E_out
def split_train_test(dataset_filepath, test_size, n_labeled):
X, y = import_libsvm_sparse(dataset_filepath).format_sklearn()
X_train, X_test, y_train, y_test = \
train_test_split(X, y, test_size=test_size)
trn_ds = Dataset(X_train, np.concatenate(
[y_train[:n_labeled], [None] * (len(y_train) - n_labeled)]))
tst_ds = Dataset(X_test, y_test)
fully_labeled_trn_ds = Dataset(X_train, y_train)
return trn_ds, tst_ds, y_train, fully_labeled_trn_ds
def main():
# Specifiy the parameters here:
# path to your binary classification dataset
dataset_filepath = os.path.join(
os.path.dirname(os.path.realpath(__file__)), 'diabetes.txt')
test_size = 0.33 # the percentage of samples in the dataset that will be
# randomly selected and assigned to the test set
n_labeled = 10 # number of samples that are initially labeled
# Load dataset
trn_ds, tst_ds, y_train, fully_labeled_trn_ds = \
split_train_test(dataset_filepath, test_size, n_labeled)
trn_ds2 = copy.deepcopy(trn_ds)
lbr = IdealLabeler(fully_labeled_trn_ds)
quota = len(y_train) - n_labeled # number of samples to query
# Comparing UncertaintySampling strategy with RandomSampling.
# model is the base learner, e.g. LogisticRegression, SVM ... etc.
qs = UncertaintySampling(trn_ds, method='lc', model=LogisticRegression())
model = LogisticRegression()
E_in_1, E_out_1 = run(trn_ds, tst_ds, lbr, model, qs, quota)
qs2 = RandomSampling(trn_ds2)
model = LogisticRegression()
E_in_2, E_out_2 = run(trn_ds2, tst_ds, lbr, model, qs2, quota)
# Plot the learning curve of UncertaintySampling to RandomSampling
# The x-axis is the number of queries, and the y-axis is the corresponding
# error rate.
query_num = np.arange(1, quota + 1)
plt.plot(query_num, E_in_1, 'b', label='qs Ein')
plt.plot(query_num, E_in_2, 'r', label='random Ein')
plt.plot(query_num, E_out_1, 'g', label='qs Eout')
plt.plot(query_num, E_out_2, 'k', label='random Eout')
plt.xlabel('Number of Queries')
plt.ylabel('Error')
plt.title('Experiment Result')
plt.legend(loc='upper center', bbox_to_anchor=(0.5, -0.05),
fancybox=True, shadow=True, ncol=5)
plt.show()
if __name__ == '__main__':
main()
# -
from sklearn.metrics import mean_squared_error
from sklearn.model_selection import GridSearchCV
model=lgbm.LGBMClassifier(n_estimators= 500, learning_rate= 0.1,num_leaves=31)
test_preds=train_model(model,train,target,test,samples=-1)
submission=pd.DataFrame(test_preds.mean(axis=1),columns=["target"])
#test_tar = pd.DataFrame(prediction,columns=['target'])
pd.concat([test_id,submission],axis=1).to_csv('GNB.csv',index=False)
submission
# Create a lgb dataset
train_set = lgb.Dataset(features, label = labels)
stdc = StandardScaler()
train_normalized = stdc.fit_transform(train)
X_train, X_test, y_train, y_test = train_test_split(train,target,test_size=0.2)
rfc = RandomForestClassifier()
rfc.fit(X_train,y_train)
y_pred = rfc.predict(X_test)
from sklearn.metrics import accuracy_score
accuracy_score(y_test,y_pred)
xgbc = xgb.XGBClassifier()
xgbc.fit(X_train,y_train)
y_pred_xgb = xgbc.predict(X_test)
accuracy_score(y_test,y_pred_xgb)
# +
prediction = xgbc.predict(test_normalized)
test_tar = pd.DataFrame(prediction,columns=['target'])
pd.concat([test_id,test_tar],axis=1).to_csv('xgb.csv',index=False)
# -
lgbmc = lgbm.LGBMClassifier()
lgbmc.fit(X_train,y_train)
y_pred_lgbm = lgbmc.predict(X_test)
accuracy_score(y_test,y_pred_lgbm)
from keras.models import Sequential
from keras.layers import Dense, Activation,Dropout
import keras
from keras import regularizers
X=np.array(train).astype(float)
Y=np.array(target).astype(np.int32)
X.shape
Y.shape
sgd = keras.optimizers.SGD(lr=0.005, momentum=0.0, decay=0.0, nesterov=False)
model = Sequential()
model.add(Dense(100, activation='relu', input_dim=200))
model.add(Dropout(0.5))
model.add(Dense(100,kernel_regularizer=regularizers.l2(0.01),
activity_regularizer=regularizers.l1(0.01)))
model.add(Dropout(0.4))
model.add(Dense(2, activation='softmax'))
model.compile(optimizer=sgd,
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
model.fit(X,Y,verbose=1,shuffle=True, nb_epoch=10,batch_size=100,validation_split=0.2)
model.predict(np.array(test).astype(float) )
test_tar = pd.DataFrame(model.predict_classes(np.array(test).astype(float)),columns=['target'])
pd.concat([test_id,test_tar],axis=1).to_csv('dnn.csv',index=False)
pd.DataFrame(model.predict_classes(np.array(test).astype(float)),columns=['target']).target.value_counts()
test
train_df['target'].value_counts()
test_id = test['ID_code']
del test['ID_code']
test_normalized = stdc.fit_transform(test)
prediction = lgbmc.predict_proba(test)
predic = prediction[:,1]
test_tar = pd.DataFrame(predic,columns=['target'])
pd.concat([test_id,test_tar],axis=1).to_csv('lgbm.csv',index=False)
test_tar.target.value_counts()
| Kaggle/santander-customer-transaction-prediction/Customer_transaction_santander_Undersampling_submissions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from IPython.display import HTML
# # Explorative analysis and data visualization
# #### <NAME> - SDS 2018, S3-4 - 7 September 2018
#
# This is a Jupyter notebook which you are going to see and use a lot as an aspiring data scientist. Notebooks can be run with different kernels (Python, R, F# etc.). This one runs Python 3.
#
# Notebooks combine Markdown cells for content with computational cells. This allows to
#
# In this session we will explore a **real** dataset from the Stanford Open Policing Project.
# The project is collecting and standardizing data on vehicle and pedestrian stops from law enforcement departments across the US — and making that information freely available. They have already gathered 130 million records from 31 state police agencies and have begun collecting data on stops from law enforcement agencies in major cities, as well.
#
# You can read more about the project [here](https://openpolicing.stanford.edu)
#
# 
#
#
# #### Exploratory data analysis
#
# You can read more about EDA with pandas [here](https://www.datacamp.com/community/tutorials/exploratory-data-analysis-python)
HTML('<iframe width="800" height="500" src="https://www.youtube-nocookie.com/embed/PelSGxTPlXM?rel=0&controls=0&showinfo=0&start=435" frameborder="0" allow="autoplay; encrypted-media" allowfullscreen></iframe>')
# This session will also be an introduction to the Python data science stack, specifically the __pandas__ and __seaborn__ packages.
#
# 
#
# - Pandas is Python's main library for managing and analysing dataframes.
# - Seaborn is a high-level library for statistical visualisation.
#
# Everything that we do today can be easily transleted in R (using dplys, ggplot etc.) Thus, it is more about the concepts than the particular language.
#
# ---
#
# ## Agenda
#
# - Import and examine the data for one US state
# - Preprocess the data (cleaning, adjusting datatypes)
# - Calculate some simple statistics on different levels of aggregation
# - Make informative plots
# - Come up and test *hypotheses*
# ## Let's get started
# First we'll download the latest dataset directly from the Stanford server. To do that we use Jupyter's **!** command line magic. Passing a `!` in a code cell, will send the command to shell rather than Python or R. `!wget` will open the GNU Wget Unix/Linux program that downloads content from web servers.
# Download the semi-raw data
# #!wget https://stacks.stanford.edu/file/druid:py883nd2578/RI-clean.csv.gz
# #!wget https://storage.googleapis.com/sds-file-transfer/CT-clean.csv.gz
# !wget https://storage.googleapis.com/sds-file-transfer/RI-clean.csv.gz
# Unzip the file to get a csv
# !gunzip RI-clean.csv.gz
# !ls -lh
# Let's import pandas as pd. The appbriviation pd is a convention. We also load the data using pandas's `read_csv` command. Pandas will try to infer the appropriate datatypes dor all columns. We set the `low_memory` argument to `False`, which is often done in cases with possible mixed datatypes. By the way: You if you place the cursor after the '(' after a command and use string+tab, you can read the docummentation for the particular command or function.
import pandas as pd
data = pd.read_csv('RI-clean.csv', low_memory=False)
# We can examine the dataset in several ways, for instance, by checking the first couple of rows or by printing the "info".
#Display the first 5 (or more/less rows) of each column
data.head()
# - Each row is one traffic stop
# - NaN are missing values
#Display overview information for the dataframe
data.info()
# print the shape of the DF as an easy alternative to looking up this in info.
data.shape
# ### Missing values
# Show missing values (mask), cut to the first 10 rows. Note, that python index alsways starts with 0!
data.isnull()[:10]
# The isnull command generates a dataframe with bool (True/False) outputs that you can apply commands on.
data.isnull().sum()
# We can see that the county_name column is all missing. And thus we can drop it. Also, we probably should drop all cases, where we don't know the time and date of the stop, the gender of the driver
# We drop the fine_grained_location, county_fips, county_name columns.
data.drop(['county_name', 'county_fips', 'fine_grained_location'], axis='columns', inplace=True)
# Drop all columns where stop_data, time, and driver_gender are missing
data.dropna(subset=['stop_date', 'stop_time', 'driver_gender'], inplace=True)
data.info()
# ### Adjusting data types
# let's check this potential candidate column
data.is_arrested.head()
# This column should be boolian but for some reason ended up being a string (or object). This is not efficient and limits our options in terms of what we can do with it. Therefore, we will change the datatype.
#
# #### On bracket vs. dot notation:
#
# In Python you will find 2 notation types. [ ] and .
#
# ``` ri['is_arrested'] is the same as ri.is_arrested````
#
# However, if you assign something and have it on the left side of = you should always use [ ] notation
#
#
# Assign the is_arrested column a new datatype
data['is_arrested'] = data.is_arrested.astype('bool')
# Is there another column that may suffer from this problem?
# ### Adjusting dates, times and index
#
# As you can see, stop_date and time are objects. That's not very useful. Let's transform them into a handy date-time-index.
#
# First, we will concatenate the two columns into one. Second, we will ask pandas to parse it and set the DF's index as the date and time of the stop. This makes lots of sense, given that each row is an *event*.
# +
# We start by concatenating the two string columns into one that we call combined, using str.cat
combined = data.stop_date.str.cat(data.stop_time, sep=' ')
print(combined.head())
# -
# Pandas provides many really nice string options that you definetely should explore. Just set your cursor after str and press tab for a list of options.
# You can for example get dummies from a categorical string variable (here just for the first 10)
data.violation[:10].str.get_dummies()
# Now we can create a date_and_time column form our combined (Series - basically a DF with only one column). Finally we will set the index of the dateframe to be the column (instead of a normal index). This will open up for many options, e.g. resampling.
# Parse the combined series to datetime-format and assign it to a new column
data['date_and_time'] = pd.to_datetime(combined)
# Replace the standard index by the new column (which will in turn disappear)
data.set_index('date_and_time', inplace=True)
# +
# data['district'] = data.county_name
# -
data.info()
# ## Exploring the data step by step
#
# We will start with simple countrs, proportions, averages etc. and move from there to more advanced concepts
# We can explore unique values for a column (even if it's a string)
data.stop_outcome.unique()
# count the distinct values
data.stop_outcome.value_counts()
# there are many ways to do the same thing
data.groupby('stop_outcome').size()
# Value counts provides a nice proportions option
data.stop_outcome.value_counts(normalize=True)
# Let's check distribution by race
data.driver_race.value_counts(normalize=True)
# ### Let's try out some hypotheses
#
# One hypothesis could be that the stop_outcome is different for different races. Discrimination?
# create 3 series for different races
black = data[data.driver_race == 'Black']
white = data[data.driver_race == 'White']
hispanic = data[data.driver_race == 'Hispanic']
black.stop_outcome.value_counts(normalize=True)
white.stop_outcome.value_counts(normalize=True)
hispanic.stop_outcome.value_counts(normalize=True)
# #### Let's try out to come up with some interesting hypotheses and find answers using the methods that we learned so far.
# Now you are probably asking: Can't we speed this up somehow?
# +
#You probably will have to
# #!pip install pandas_profiling
# -
import pandas_profiling
pandas_profiling.ProfileReport(data)
# ### Filtering my multiple conditions
#
# We can of cause chain filter conditions (you probably learned to do that in tha last session using R)
# note the ( ) around the conditions
hispanic_and_arrested = data[(data.driver_race == 'Hispanic')
& (data.is_arrested == True)]
# in case you are bored by the shape command
len(hispanic_and_arrested)
# We can also ask for hispanic OR arrested (not sure how much that tells us)
hispanic_or_arrested = data[(data.driver_race == 'Hispanic')
| (data.is_arrested == True)]
hispanic_or_arrested.shape
# ##### Rules for filtering
#
# - & AND
# - | OR
# - Each condition must be surrounded by () and many are possible
# - == Equality
# - != Inequality
#
# ##### Remember, that we are not making any statement about causation. This is purely a correlation exercise (so far!)
#
# #### A bit on boolean series
#
# True = 1 and False = 0
# Which means that you can perform calculations on them:
#
# +
# Create a DataFrame of male and female drivers stopped for speeding
female_and_speeding = data[(data.driver_gender == 'F') & (data.violation == 'Speeding')]
male_and_speeding = data[(data.driver_gender == 'M') & (data.violation == 'Speeding')]
# Compute the stop outcomes for drivers (as proportions)
print(female_and_speeding.stop_outcome.value_counts(normalize=True))
print(male_and_speeding.stop_outcome.value_counts(normalize=True))
# -
# If you have multiple things that should be shown from one cell's output, it's better to print it out
print(data.is_arrested.dtype)
print(data.is_arrested.mean())
# ### Using groupby to compare groups
#
# Rememebre when we compared stop outcome rates by race? Well: That was not very elegant. We can certainly do better using the groupby function
# Here we first group by the race and then calculate the mean for the arrested column
data.groupby('driver_race').is_arrested.mean()
# We can see that black and Hispanic drivers end up more than twice as often arrested than white drivers. But perhaps geography plays a role and perhaps there are some outlier "bad neighborhoods"
# Group data by district and race and calculate the mean of a third factor
data.groupby(['district','driver_race']).is_arrested.mean()
# Which violations lead for the different genders to what rates of arrest?
data.groupby(['violation','driver_gender']).is_arrested.mean()
# ### "protective frisk"
# Sometimes during stops if a search is conducted, the officer also checks the driver if they have a weapon. This is called a "protective frisk".
# Let's try to figure out if men are frisked more than women.
# Look at the different search types performed
data.search_type.value_counts()
# #### Extracting a string
# As you can see, search type is a multiple choice object/string column. *Incident to Arrest* and *Pribable Cause* are the most commont but combinations are possible. We can use the `str.contains` method to filter to filter out cases of interest. This will return a boolean series, which we can assign to a new varioable 'frisk' in our dataframe.
# +
# We ask pandas to find out if the string in the search_type column contatins
# the sequence "Protective Frisk"
# We assign the result to a new column that we call "frisk"
data['frisk'] = data.search_type.str.contains('Protective Frisk', na = False)
# +
# backup code in case you are not convinced that it worked
# this snippet shows the source and the boolean result side by side
# generate a data frame by concatenating source and target of the str.contains method. Note, that this concat
# of columns or rows is different from the str.cat that we learned before
frisks = pd.concat([data.search_type, data.search_type.str.contains('Protective Frisk', na = False)], axis=1)
# display the second column using the iloc selector (here useful since we have two columns with the same name)
frisks[frisks.iloc[:,1] == True]
# -
data['frisk'].sum()
# Do men get frisked more often?
data.groupby('driver_gender').frisk.mean()
# ## Using the datetime index to select data
#
# What if you assume that things got better or worse over the years? Or perhaps the time of the day plays a role? 📈
# Remember we assigned a datetime column to our index?
# In case you need a recap:
#
# ```python
# combined = data.stop_date.str.cat(data.stop_time, sep=' ')
# data['date_and_time'] = pd.to_datetime(combined)
# data.set_index('date_and_time', inplace=True)
# ```
#
# That allows us now to access the time dimension at various levles in our index.
#
# ```python
# data.index.day
# data.index.month
# data.index.day_name()
# data.index.month_name()
# ```
#
# We can now use that for groupby etc.
# Are things getting better or worse over the years?
data.groupby(data.index.year).frisk.mean()
data.groupby(data.index.year).is_arrested.mean()
data.groupby(data.index.month_name()).is_arrested.mean()
# ### Let's start plotting some of these stats
#
# Pandas provides a very easy plotting interface for standard visualisations. For more complex plots, we will be using Seaborn later on.
#
# But for now, you can access plotting simply by adding ```.plot()```` after applying some calculation to a dataframe or series.
# +
# don't forget to tell Jupyter to activate inline plots
# %matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sns
sns.set(style="darkgrid")
plt.rcParams["figure.figsize"] =(12,10)
# +
# e.g. Show the different months that can be accessed
figure = data.groupby(data.index.month).is_arrested.mean().plot(kind='bar')
plt.xlabel('month')
plt.ylabel('arrest rate')
plt.title('Monthly mean arrest rate in traffic stops')
# saving the plot is as easy as
plt.savefig('is_arrested_bymonth.pdf', format='pdf')
# -
# #### Using resampling to aggregate data rather than group it.
#
# But isn't that the same what we just did before? Well, actually it's not the same. Instead of grouping by month (12 months), we will re-aggregate the data for the individual months: Jan 05 - Dec 15.
#
# ``` python
# data.is_arrested.resample('M').mean()````
#
# More on that here: [Resampling time series data with pandas](http://benalexkeen.com/resampling-time-series-data-with-pandas/)
#
data.is_arrested.resample('M').mean().plot()
data.search_conducted.resample('M').mean()
data['search_conducted'] = data.search_conducted.astype('bool')
# +
# Save the monthly rate of drug-related stops
monthly_drug_rate = data.drugs_related_stop.resample('M').mean()
# Calculate and save the monthly search rate
monthly_search_rate = data.search_conducted.resample('M').mean()
# Concatenate the two
monthly = pd.concat([monthly_drug_rate,monthly_search_rate], axis='columns')
# cut of a few years in the beginning
monthly = monthly[monthly.index>=pd.to_datetime('2007-1-1')]
# Create subplots from 'annual'
monthly.plot(subplots=True)
# Display the subplots
plt.show()
# -
# #### Crosstabs
#
# Crosstabs allow us to explore categorical data considering varous dimensions. Actually, it is the same as grouping by several columns, counting it up with .size() and then bringing the data in wide format (using .unstack())
#
# You'll agree that crosstab is easier...however for some reason computationally more expensive 🤔
# Sidenote:
#
# ```%time``` infront of a line of code to measure execution time
# ```%%time``` before executing a code-chunk (for instance a loop or a function which we will cover eventully)
#
# ```%timeit``` will perform the same 10, 100, 1000 times and give you an average (you never know what your CPU and memory are up to at any moment)
#
# %time data.groupby([data.driver_race, data.driver_gender]).size().unstack()
# %time pd.crosstab(data.driver_race, data.driver_gender)
# ### Transforming categorical in to nummerical data
#
# You may have noted the ```stop_duration``` column in our dataset and that it is an ```object``` variable. That means, we can use it as a dimension but not to perform any calculations. What we can do, is map the categories to a reasonable nummerical value using a mapping dictionary the ```map``` command.
# +
# First, we should inspect the unique values of the column:
# you could use the unique() command but value_counts() is helpful here,
# as it can help us identify outlier cases
data.stop_duration.value_counts()
# -
# As you can see, there are 6 rows that clearly don't fit and should probably be eliminated
# We need to filter the dataframe for observations where the ```stop_duration``` hase one of the three values: '0-15 Min', '16-30 Min', '30+ Min'
#
# We can use pandas' ```isin``` function here. By the way: If you are doing the opposite "not in" query, you can set a ```~``` after the ```[``` and it will inverse your selection.
# I'm lazy: First we can ask pandas to give us a list of possible values that we can copy-paste below into our query
data.stop_duration.unique()
# +
# This step should be pretty familiar
data = data[data.stop_duration.isin(['0-15 Min', '16-30 Min', '30+ Min'])]
# +
# Let's see if the inverse also works:
data[~data.stop_duration.isin(['0-15 Min', '16-30 Min', '30+ Min'])].stop_duration.unique()
# -
# As you can see, challenges pop up all the time and that's why working with data is not a linear or clean process but messy, at times confusing and will require you to look up things on the interenet all the time.
#
# Now we can create a mapping dictionary. But wait, we never covered what a dictionary is.
#
# A dictionary is one of Python's fundamental data structures. You already met ints, floats, strings. There are also lists, sets, tuples and dictionaries (these are the most common types).
#
# For now: A dictionary maps ```keys```to ```values```
#
# 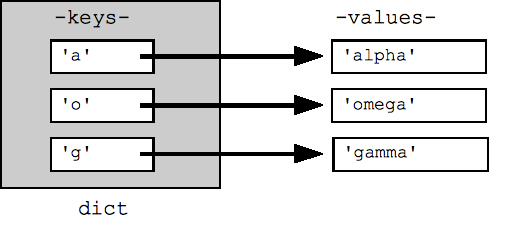
#
# You input a key and you get a value, disregarding the order. A dictionary has a slightly weird syntax with curly brackets and : but you'll get used to it.
# +
# Let's create our mapping
mapping = {'0-15 Min':7.5, '16-30 Min':23, '30+ Min':45}
# And use it right away to create a new column
data['stop_duration_num'] = data.stop_duration.map(mapping)
# a quick check of what we achieved
print(data.stop_duration_num[:5])
# -
# #### A quick intro to loops
#
# The great advantage with computers is that they are happy to do boring repetitive stuff for us without getting tired. If we think a bit about the ```map``` function, what it did was to take every observation of ```stop_duration``` in our dataframe and translate it to the corresponding value in our mapping dictionary.
#
# Let's try some made up pseudocode:
#
# ```
#
# stop_duration_num = [] #creating an empty list
#
# for every observation in data.stop_duration do:
# look up corresponding value in the mapping dictionary
# append the value to the stop_duration_num list
#
# finally:
#
# data['stop_duration_num'] = stop_duration_num list
#
# ```
#
#
# Actually we can write exactly that as a loop
#
# ```python
# stop_duration_num_list = []
#
# for x in data.stop_duration:
# value = mapping[x]
# stop_duration_num_list.append(value)
#
# # we skip the creation of the new column since we already have it
#
# ```
#
# Aside form for-loop as that one there are also while loops that will do something while some condition is met
#
# You can find more on datatypes and loops (iteration) in [this cheat sheet](https://www.theredhillacademy.org.uk/pluginfile.php?file=/15673/block_html/content/Python%20Cheat%20Sheet.pdf)
# and many other places. Iteration is a core concept in computer science and will be important in later modules. If you code for the first time in your life, it's a slightly strange concept to get your head around but after some time it becomes second nature. For now, it's good to get some initial feeling for the concept.
#
# #### Let's return to our ```stop_duration``` problem
#
# Since, we created a nummerical value, we can for instance check if average stop duration is different for different violations:
# +
# Let's do everything in one line
data.groupby([data.violation_raw]).stop_duration_num.mean().sort_values().plot(kind='barh')
# -
# #### Cutting intervals
#
# ```data.driver_age``` contains the (duh) the age of the driver. ```data.driver_age.dtype``` will tell us that it's a continuous nummerical value and thus good for more advanced analysis but perhaps a bit to detailed for exploration (?)
#
# More instrumental in that context would be to slice that variable up into ordered categries corresponding to age-populations of interest, say "teen", "20s", "30s" etc.
# Let's check the variable
data.driver_age.describe()
# We can slice tha data into some numebr of bins (sometimes useful)
data['age_cat'] = pd.cut(data.driver_age,bins=6)
# We can also provide labels
labels=["teen", "20s", "30s", "40s", "50+"]
data['age_cat'] = pd.cut(data.driver_age, bins=5, labels=labels)
# We can also provide custom intervals
# Unfortunately (not sure why) there is no way to do both together
bins = pd.IntervalIndex.from_tuples([(10, 20), (20, 30), (30, 40), (40,50), (50,100)])
data['age_cat'] = pd.cut(data.driver_age, bins = bins)
data.dropna(subset=['age_cat'], inplace=True)
# +
# We can plot that but perhaps let's modify a bit first
table = pd.crosstab(data.driver_race, data.age_cat)
# keep only minority races
table = table.iloc[:3,]
# let's plot some stacked bars
table.plot(kind='bar', stacked=True)
# -
# ### Enters weather data
#
# To be precise: Local climatological data from https://www.ncdc.noaa.gov/
#
# I put the data in a Google Cloud bucket: https://storage.googleapis.com/sds-file-transfer/RI-weather.csv
weather = pd.read_csv('weather.csv')
weather.head()
weather[['AWND', 'WSF2']].head()
weather[['AWND', 'WSF2']].plot(kind='box')
weather['WDIFF'] = weather.WSF2 - weather.AWND
weather.WDIFF.plot(kind='hist')
weather.WDIFF.plot(kind='hist', bins=50)
plt.savefig('fig1.pdf') # would like to save it for later? Pass this line in the same cell and you'll get the pdf
# #### Introducing Seaborn
#
# Seaborn is a great project with the mission to make statistical plots easier in Python. You can find more on their [Homepage](https://seaborn.pydata.org/index.html).
#
# Datacamp created [this cheat sheet](https://www.datacamp.com/community/blog/seaborn-cheat-sheet-python) that summarizes most important functions.
# +
# We already imported seaborn (without talking a lot about it). Let's do it again, which is not a problem
import seaborn as sns # sns is the conventional abbriviation
sns.set(style="darkgrid") # Darkgrid is a nice ggplot (R) - like style that looks
# -
# Let's do the same what we just did but now using Searborn
# We replace the standard histogram by a Distribution plot (same same)
sns.distplot(weather.WDIFF)
# You can read much more about all the different things that you can do with distplots [here](https://seaborn.pydata.org/generated/seaborn.distplot.html)
# +
fig = sns.distplot(weather.WDIFF, hist=False,
rug=True,
kde_kws={'shade':True})
fig.figure.savefig('fig.pdf') # In case you would like to keep it :-)
# -
# Further examining the weather data
weather.shape
weather.columns
#Let's select just a sequence of columns 'from-to'
temp = weather.loc[:, 'TAVG':'TMAX']
temp.head()
# +
# Can we say something about average temperature and average wind?
sns.jointplot(weather.TAVG, weather.AWND, kind="hex", color="#4CB391")
# -
# #### Data types advanced
#
# We already used the map function to transform strings to nummerical values.
# Now, let's try to create categories. This is not always necessary but nice to know.
mapping = {'0-15 Min':'short', '16-30 Min':'medium', '30+ Min':'long'}
data['stop_length'] = data.stop_duration.map(mapping)
data.stop_length.memory_usage(deep=True)
# +
# We need to initiate this new data format first importing it
from pandas.api.types import CategoricalDtype
# Define our categories
cats = ['short', 'medium', 'long']
# And define the specific cateogory type (this is useful for survey data that is ordered)
cat_type = CategoricalDtype(categories=cats, ordered=True)
# Finally let's asign it
data['stop_length'] = data.stop_length.astype(cat_type)
# +
data.stop_length.memory_usage(deep=True)
# The first thing we notice --> this type is much more memory friendly. This is good when moving towards big data
# -
# Also now we see that pandas knows that short is short < medium < long
data.stop_length.head()
# Any relation between stop duration and likelihood to get arrested?
data.groupby('stop_length').is_arrested.mean()
# #### Back to our weather data
#
# Do you think that arrest rates or certain violations are related to weather?
# For that we need to connect our weather data with the stops data
#
# We will merge the two dataframes on the date index, as weather data is available daily
# First let's prepare the weather data first, by parsing the date-column
weather.DATE = pd.to_datetime(weather.DATE)
# We need to do the same with our stops data and its stop_date column
data.stop_date = pd.to_datetime(data.stop_date)
# +
# now let's merge
data_weather = pd.merge(data, weather, left_on='stop_date', right_on='DATE', how='left')
# -
# Pandas merge command is fairly simple:
#
# We start with the left dataframe "data" in our case, then the right "weather", then pass the left and the right key and finally the merger type: Here "left", meaning that we would like to keep the left as it is and multiply the right on top of it (if that makes sense)
#
# 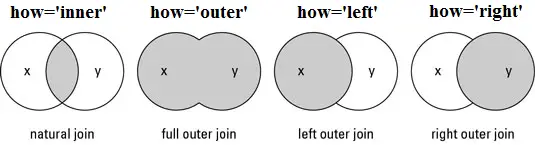
#
# And if you still think merging is mysterious then you should check out this [youtube tutorial](https://youtu.be/h4hOPGo4UVU)
# data_weather.set_index('date_and_time', inplace=True)
data_weather.index = data.index
# It is a not totally crazy assumption that police stops are shorter if it's cold...maybe. Let's check that.
#
# The syntax is a bit more advanced but I'll do my best to explain. We will control for the violation type, assuming that different violations lead to different durations by their nature.
# We need to aggregate the data first on 2 levels: Time and Violation.
# For this we need to use pandas Grouper module (I also had to look it up)
violation_duration = data_weather.groupby([pd.Grouper(key='stop_date', freq='D'), 'violation']).stop_duration_num.mean()
# Such groupby commands deliver a multi-indexed series. That's not useful for further work.
# But we can transform them into Dataframes with various index levels turning into columns.
violation_duration = pd.DataFrame(violation_duration).reset_index()
# Same for the temperature data
avg_temp = data_weather.resample('D').TAVG.mean()
avg_temp = pd.DataFrame(avg_temp).reset_index()
# Now let's merge both DFs
search_temp = pd.merge(violation_duration, avg_temp, left_on='stop_date', right_on='date_and_time', how='left')
# We plot our resulting DF using Seaborn using the col argument to create subplots for the different violation types
sns.lmplot('stop_duration_num', 'TAVG', data=search_temp[search_temp.stop_duration_num < 20], col = 'violation')
# #### Heatmaps are another useful visualization
# You should by now be able to read this without problems
sns.heatmap(pd.crosstab(data.driver_race, data.driver_gender, values=data.is_arrested, aggfunc='mean'))
# +
# We can make this heatmap more informative using some of Seaborn's functionality
sns.heatmap(pd.crosstab(data.driver_race, data.driver_gender, values=data.is_arrested, aggfunc='mean'),
annot=True, cmap="YlGnBu", cbar=False, linewidths=.5)
# -
# ### Search discrimination?
#
# Finally we would like to find out if searches are conducted more often for some races.
# First we need to create a boolean variable for searches
data['search_conducted'] = data.search_conducted.astype('bool')
# +
# Calculate the search rate by race
search_rates = pd.crosstab(data.district, data.driver_race, data.search_conducted, aggfunc='mean')
### Exactly the same
# search_rates = data.groupby(['district','driver_race']).search_conducted.mean().unstack()
# -
searches_per_district = data.groupby('district').size()
search_rates_per_district = pd.concat([search_rates, searches_per_district], axis = 1)
search_rates_per_district['stops'] = search_rates_per_district[0]
search_rates_per_district
# Now some more advanced plotting:
#
# Can you understand what's going on here?
plt.figure(figsize=(9,9))
ax = plt.axes()
ax.set_title('White vs. Hispanic Search Rate')
plot = sns.scatterplot(x="White", y="Hispanic", size='stops', hue="stops", data=search_rates_per_district)
plt.axis([0,0.085,0,0.085])
plot.plot(plot.get_xlim(), plot.get_ylim(), ls="--", c=".3")
plt.figure(figsize=(9,9))
ax = plt.axes()
ax.set_title('White vs. Black Search Rate')
plot = sns.scatterplot(x="White", y="Black", size='stops', hue="stops", data=search_rates_per_district)
plt.axis([0,0.085,0,0.085])
plot.plot(plot.get_xlim(), plot.get_ylim(), ls="--", c=".3")
found_rates = data.groupby(['district','driver_race']).contraband_found.sum() / data.groupby(['district','driver_race']).search_conducted.sum()
found_rates_per_district = pd.concat([found_rates.unstack(),data.groupby('district').search_conducted.sum()], axis = 1)
found_rates_per_district
plt.figure(figsize=(9,9))
ax = plt.axes()
ax.set_title('White vs. Black Find Rate')
plot = sns.scatterplot(x="White", y="Black", size='search_conducted', hue="search_conducted", data=found_rates_per_district)
plt.axis([0,0.5,0,0.5])
plot.plot(plot.get_xlim(), plot.get_ylim(), ls="--", c=".3")
plt.figure(figsize=(9,9))
ax = plt.axes()
ax.set_title('White vs. Hispanic Find Rate')
plot = sns.scatterplot(x="White", y="Hispanic", size='search_conducted', hue="search_conducted", data=found_rates_per_district)
plt.axis([0,0.5,0,0.5])
plot.plot(plot.get_xlim(), plot.get_ylim(), ls="--", c=".3")
| M1_S3-4 (part one).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sudoku
import numpy as np
import functools
# ## List of Lists to save the indexes of non-zero values in sudoku boards
# ## MIght be lighter on memory
# ## SHould be faster also{
#
normal_sudoku = [[8,0,0,0,0,0,0,0,0],
[0,0,3,6,0,0,0,0,0],
[0,7,0,0,9,0,2,0,0],
[0,5,0,0,0,7,0,0,0],
[0,0,0,0,4,5,7,0,0],
[0,0,0,1,0,0,0,3,0],
[0,0,1,0,0,0,0,6,8],
[0,0,8,5,0,0,0,1,0],
[0,9,0,0,0,0,4,0,0]]
a = sudoku.Sudoku(normal_sudoku, kind = "adj_list")
a.next_move()
len(a.avail_moves)
a.board[1]
a.blocked_dict[0]
a.avail_moves
# # Sudoku.next_move can return a couple different things:
#
# # While there are single moves it calls sudoku.make_move(singlemove)
#
# # If there are no possible moves:
# # Return: No possible moves
# # If there are no single moves:
# # Return: list of guesses with minimum lenght
a.next_move()
a.make_move(21, 3)
a = {1:3, 4:5}
len()
len(a)
a.set_visual_board()
a.next_move()
a.avail_moves
# %%timeit
for x,row in enumerate(normal_sudoku):
for y,num in enumerate(row):
aoa[num]
# %%timeit
for x,row in enumerate(normal_sudoku):
for y,num in enumerate(row):
testboard[num][x][y] = True
tzero = testboard[0]
testboard
def available_moves(sudoku):
'returns the available guess_list of every square'
guess_list = all_moves()
for i in range(9):
for j in range(9):
if (not (sudoku[i][j])) == False:
numago = sudoku[i][j]
guess_list[(9*i + j)] = []
for k in range(9):
if numago in guess_list[9*i +k]:
guess_list[(9*i + k)].remove(numago) #remove da linha
if numago in guess_list[(9*k + j)]:
guess_list[(9*k + j)].remove(numago) #remove da coluna
a,b = (i//3)*3,(j//3)*3
for m in range(a,a+3):
for n in range(b,b+3):
if numago in guess_list[9*m+n]:
guess_list[(9*m+n)].remove(numago)
return guess_list
| jupyter notebooks/Some Tests.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.10 64-bit (''tf_mac'': conda)'
# name: python3
# ---
# +
import pandas as pd
df = pd.read_csv('./datasets/adult.csv',delimiter=',', skipinitialspace=True)
del df['fnlwgt']
del df['education-num']
for col in df.columns:
if '?' in df[col].unique():
df[col][df[col] == '?'] = df[col].value_counts().index[0]
# -
columns = df.columns.tolist()
columns = columns[-1:] + columns[:-1]
df = df[columns]
class_name = 'class'
possible_outcomes = list(df[class_name].unique())
df.head(5)
possible_outcomes
integer_features = list(df.select_dtypes(include=['int64']).columns)
float_features = list(df.select_dtypes(include=['float64']).columns)
string_features = list(df.select_dtypes(include=['object']).columns)
type_features = {
'integer': integer_features,
'float': float_features,
'string': string_features,
}
type_features
### Seperate to numerical categorical
numerical_cols = type_features['integer'] + type_features['float']
categorical_cols = type_features['string']
numerical_cols
categorical_cols
# +
from sklearn.preprocessing import LabelEncoder
def label_encode(df, columns, encoder_dict=None):
df_temp = df.copy(deep=True)
if encoder_dict:
for col in columns:
col_encoder = encoder_dict[col]
df_temp[col] = col_encoder.transform(df_temp[col])
else:
encoder_dict = {}
for col in columns:
encoder = LabelEncoder()
df_temp[col] = encoder.fit_transform(df_temp[col])
encoder_dict[col] = encoder
return df_temp, encoder_dict
# -
def label_decode(df, columns, encoder_dict):
temp_df = df.copy(deep=True)
for col in columns:
encoder = encoder_dict[col]
temp_df[col] = encoder.inverse_transform(temp_df[col])
return temp_df
df, encode_dict = label_encode(df, categorical_cols)
df
categorical_cols
pd.get_dummies(df['workclass'], drop_first=True)
def transform_to_dummy(df, columns):
for col in columns:
df = pd.concat([df,pd.get_dummies(df[col], prefix=col)],axis=1)
df.drop([col],axis=1, inplace=True)
return df
df = transform_to_dummy(df, categorical_cols)
df.head(5)
from sklearn.tree import DecisionTreeClassifier
| deprecated/dataset_preprocessing.ipynb |