code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/JohnCrl/CPEN-21A-ECE-2-1/blob/main/Midterm_Exam.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="36-LXXQzwZRT"
# **Midterm Exam**
# + [markdown] id="EVA0Gn7Hx_Lf"
# ***Problem Statement 1***
# + colab={"base_uri": "https://localhost:8080/"} id="tIe5qHyHwPJP" outputId="f261551c-9767-4a1c-ebc3-2d55e257c6aa"
a= "<NAME>"
b= "202012942"
c= "21 years old"
d= "November 18, 2000"
e= "Lot 1373 C-2 Marseilla St. Bagbag II, Rosario Cavite"
f= "Bachelor of Science in Electronics Communication Engineering"
g= "1.67"
print(a)
print(b)
print(c)
print(d)
print(e)
print(f)
print(g)
# + [markdown] id="tAwMZKNayMRH"
# ***Problem Statement 2***
# + [markdown] id="LrS4x3Au2XLB"
# **A.**
# + colab={"base_uri": "https://localhost:8080/"} id="STZX2RbDyPL4" outputId="1b2050bd-2318-4d3b-ece9-defeee389fcb"
n=4
a=(2<n) and (n<6)
print(a)
# + [markdown] id="esfbbacG2Z9l"
# **B.**
# + colab={"base_uri": "https://localhost:8080/"} id="0xiFpz6M2bNJ" outputId="0fc37436-cb08-4c33-bf17-f01a3fa0f33c"
n=4
b=(2<n) or (n==6)
print(b)
# + [markdown] id="9kwg5L5C2-gf"
# **C.**
# + colab={"base_uri": "https://localhost:8080/"} id="1EKNwz-x3AKH" outputId="d7ffa178-3b6d-4725-ee4b-421f4bf7f28d"
n=4
c=(not(2<n or n==6))
print(c)
# + [markdown] id="LXYGN_Rf3gIQ"
# **D.**
# + colab={"base_uri": "https://localhost:8080/"} id="u0FzMNNy3hmo" outputId="1ee9d732-a8ae-4331-ead1-41842f501681"
n=4
d=(not(n<6))
print(d)
# + [markdown] id="Mons48lP4iVA"
# **E.**
# + colab={"base_uri": "https://localhost:8080/"} id="nu6ssWtL4kcf" outputId="bbd778cd-f503-4166-8e97-081fff6ac69b"
n=4
answ="Y"
e=(answ=="Y") or (answ=="y")
print(e)
# + [markdown] id="HRWz05L-6sVA"
# **F.**
# + colab={"base_uri": "https://localhost:8080/"} id="Zji5Z-Gi6u1f" outputId="1cace002-1951-4824-b932-7ac978a11245"
n=4
answ="Y"
f=(answ=="Y") and (answ=="y")
print(f)
# + [markdown] id="47YG0VaR70pH"
# **G.**
# + colab={"base_uri": "https://localhost:8080/"} id="cDI-VYhB75pI" outputId="b6563723-fb60-4f18-bb43-12391c4a43ce"
n=4
answ="Y"
g=not(answ=="y")
print(g)
# + [markdown] id="S78useXG8vLL"
# **H.**
# + colab={"base_uri": "https://localhost:8080/"} id="MrM7TSOv8wlz" outputId="ae8d548d-d9b6-4284-9168-e02d95f23a6e"
n=4
answ="Y"
h=(((2<n)and(n==5+1)) or (answ=="No"))
print(h)
# + [markdown] id="Oac4IA28-3o3"
# **I.**
# + colab={"base_uri": "https://localhost:8080/"} id="yrZ3coRn-5s5" outputId="320fbb2e-8ab5-40d6-a420-38a3a018d023"
n=4
answ="Y"
i=(((n==2)and(n==7)) or (answ=="Y"))
print(i)
# + [markdown] id="BuFFbHb3_Nam"
# **J.**
# + colab={"base_uri": "https://localhost:8080/"} id="Yqq5O5X4_O-P" outputId="26f1aef9-a335-4256-94dd-253f68f7ad72"
n=4
answ="Y"
j=((n==2)and((n==7)or(answ=="Y")))
print(j)
# + [markdown] id="zbu8FSKIygqv"
# ***Problem Statement 3***
# + colab={"base_uri": "https://localhost:8080/"} id="NaHFzeduyi-X" outputId="3070cc9f-2f2e-4146-b454-f64df9020e8f"
x= 2
y= -3
w= 7
z= -10
print(x/y)
print(w/y/x)
print(z/y%x)
print(x%-y*w)
print(x%y)
print(z%w-y/x*5+5)
print(9-x%(2+y))
print(z//w)
print((2+y)**2)
print(w/x*2)
| Midterm_Exam.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.11 64-bit (''base'': conda)'
# name: python3
# ---
# +
# quick calculations on population and country area
# imports
import pandas as pd
import sys
# set up file paths and other data globals
import config
import modify
sys.path.append(config.CURRENT_DIR_STR)
sys.path.append(config.COVID_PACKAGE_STR)
sys.path.append(config.UPDATE_FILE_STR)
# local imports
from covid_package.data_funcs.store_data import read_json_data, convert_owid_data
from covid_package.libs.valid_keys import fetch_l0_keys
# + tags=[]
# read the updated(?) data file from the data dir
data = read_json_data(config.COUNTRY_FILE_STR)
# convert the OWID_ keys
#data = convert_owid_data(data)
#data.pop('CYN')
# repopulate the keys
key_list = fetch_l0_keys(data)
#print(data)
# + tags=[]
# organize the pandas column data
loc_list = []
pop_list = []
dens_list = []
for val in data.values():
loc_list.append(val['location'])
pop_list.append(val['population'])
dens_list.append(val['population_density'])
#print([loc_list.append(val['location']) for val in data.values()])
pop_dict = {
'iso': key_list,
'location': loc_list,
'population': pop_list,
'population_density': dens_list
}
# -
# create the dataframe
df = pd.DataFrame(pop_dict)
pd.options.display.max_rows = 999
df.drop(df[df['population_density'] == 0].index, inplace = True)
df.sort_values(by = 'location', inplace=True)
df
df['area km^2'] = df['population'] / df['population_density']
df['population'] = df['population'].astype(int)
df['area km^2'] = df['area km^2'].astype(int)
df
| pop_calcs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: pumpitup
# language: python
# name: pumpitup
# ---
import os
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
#import geopandas as gpd
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import RandomizedSearchCV
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
from sklearn.compose import ColumnTransformer
from sklearn.preprocessing import OneHotEncoder
from sklearn.pipeline import Pipeline
from sklearn.impute import SimpleImputer
from sklearn.linear_model import LogisticRegression
submit_X = pd.read_csv('Source_data/testset_values.csv')
data_test = test_transformer(submit_X)
data_test.columns == data.drop(columns=['label']).columns
num_cols = ['gps_height', 'population']
cat_cols = ['basin', 'region_code', 'district_code', 'extraction_type', 'payment_type', 'water_quality', 'quantity',
'source', 'management', 'management_group', 'waterpoint_type', 'funder', 'installer', 'subvillage',
'ward', 'scheme_management', 'scheme_name', 'amount_tsh']
scaler = StandardScaler()
ohe = OneHotEncoder(handle_unknown='ignore')
data2.columns
scaler = StandardScaler()
ohe = OneHotEncoder(drop='first')
train_num = data2.loc[:,]
for c in train_num.columns:
train_num[c] = scaler.fit_transform(train_num.loc[:,c].to_numpy().reshape(-1,1))#
train_num
colt = ColumnTransformer([('basin', ohe, ['basin']),
('r_code', ohe, ['region_code']),
('d_code', ohe, ['district_code']),
('ex', ohe, ['extraction_type']),
('pay', ohe, ['payment_type']),
('qual', ohe, ['water_quality']),
('quant', ohe, ['quantity']),
('s_type', ohe, ['source']),
('mang', ohe, ['management']),
('mang_g', ohe, ['management_group']),
('wp_type', ohe, ['waterpoint_type']),
('fund', ohe, ['funder']),
('inst', ohe, ['installer']),
('subv', ohe, ['subvillage']),
('ward', ohe, ['ward']),
('sch_man', ohe, ['scheme_management']),
('sch', ohe, ['scheme_name'])])
train_cat = data2.loc[:,]
train_df = colt.fit_transform(train_cat).toarray()
train_cat = pd.DataFrame(train_df, columns=colt.get_feature_names())
train = train_num.merge(train_cat, left_index=True, right_index=True)
train
train.shape
train = train.merge(data2[['lga_Njombe', 'year', 'month', 'tsh_bin', 'years_old', 'popbins', 'longitude', 'latitude',
'public_meeting', 'permit']], left_index=True, right_index=True)
train_X, test_X, train_y, test_y = train_test_split(train, data2['label'], test_size=0.25, random_state=42)
rf = RandomForestClassifier()
params = {'n_estimators':[10, 100, 200, 500], 'criterion':['gini', 'entropy']}
rs = RandomizedSearchCV(rf, params, n_iter=5)
rf.fit(train_X, train_y)
rf.score(test_X, test_y)
rs.fit(train_X, train_y)
rs.best_estimator_
rs.cv_results_
rf.classes_
features = pd.Series(rf.feature_importances_, index=train.columns)
features.sort_values(ascending=False).head(20)
data2['label2'] = data2['label'].replace({'functional needs repair':'functional'})
train_X, test_X, train_y, test_y = train_test_split(train, data2['label'], test_size=0.25, random_state=42)
rf.fit(train_X, train_y)
rf.score(test_X, test_y)
rs.score(test_X, test_y)
y_pred = rs.predict(test_X)
from sklearn.metrics import confusion_matrix, ConfusionMatrixDisplay
cm=confusion_matrix(test_y, y_pred, labels=rs.classes_)
disp = ConfusionMatrixDisplay(cm, display_labels=rs.classes_)
plt.figure(figsize=(40,8))
disp.plot()
test_X.insert(2,value=y_pred, column='prediction')
test_X
y_pred = rs.predict(train)
rs.score(train, data2['label'])
data2.insert(2, value=y_pred, column='prediction')
data2.head()
errors = data2.loc[data2['label']!=data2['prediction']]
errors
correct = data2.loc[data2['label']==data2['prediction']]
correct
errors.describe()
sns.scatterplot(data=errors, x='longitude', y='latitude')
plt.show()
errors['amount_tsh'].plot(c='red')
data2['amount_tsh'].plot(c='blue')
plt.show()
submit_X = pd.read_csv('Source_data/testset_values.csv')
train_num = submit_X.loc[:,['amount_tsh', 'gps_height', 'population',]]
for c in train_num.columns:
train_num[c] = scaler.fit_transform(train_num.loc[:,c].to_numpy().reshape(-1,1))#
train_num
colt = ColumnTransformer([('basin', ohe, ['basin']),
('r_code', ohe, ['region_code']),
('d_code', ohe, ['district_code']),
('ex', ohe, ['extraction_type']),
('pay', ohe, ['payment_type']),
('qual', ohe, ['water_quality']),
('quant', ohe, ['quantity']),
('s_type', ohe, ['source']),
('mang', ohe, ['management']),
('mang_g', ohe, ['management_group']),
('wp_type', ohe, ['waterpoint_type']),
('fund', ohe, ['funder']),
('inst', ohe, ['installer']),
('subv', ohe, ['subvillage']),
('ward', ohe, ['ward']),
('sch_man', ohe, ['scheme_management']),
('sch', ohe, ['scheme_name'])])
train_cat = submit_X.loc[:,['basin', 'region_code', 'district_code', 'extraction_type', 'payment_type', 'water_quality',
'quantity', 'source', 'management', 'management_group', 'waterpoint_type',
'funder', 'installer', 'subvillage', 'ward', 'scheme_management', 'scheme_name']]
train_df = colt.fit_transform(train_cat).toarray()
train_df
train_cat = pd.DataFrame(train_df, columns=colt.get_feature_names())
train_cat
train = train_num.merge(train_cat, left_index=True, right_index=True)
train.shape
train.shape
train = train.merge(data2[['lga_Njombe', 'year', 'month', 'tsh_bin', 'years_old', 'popbins', 'longitude', 'latitude',
'public_meeting', 'permit']], left_index=True, right_index=True)
train_X, test_X, train_y, test_y = train_test_split(train, data2['label'], test_size=0.25, random_state=42)
rf = RandomForestClassifier()
params = {'n_estimators':[10, 100, 200, 500], 'criterion':['gini', 'entropy']}
rs = RandomizedSearchCV(rf, params, n_iter=5)
rf.fit(train_X, train_y)
rf.score(test_X, test_y)
submition = rf.predict(sub_train)
submission1 = pd.DataFrame(submition, columns=['status_group'])
submission1['id'] = submit_X['id']
submission1.to_csv('submission1_BW.csv')
def test_transformer(data):
data = data.drop(columns=['source_type', 'source_class'])
data = data.drop(columns=['extraction_type_group', 'extraction_type_class'])
data['extraction_type'].replace({'other - swn 81':'other-handpump',
'other - play pump':'other-handpump',
'walimi':'other-handpump',
'other - mkulima/shinyanga':'other-handpump',
'swn 80':'swn_80',
'nira/tanira':'nira-tanira',
'india mark ii':'india_mark_ii',
'india mark iii':'india_mark_iii',
'other - rope pump':'other-rope_pump',}, inplace=True)
data['source'].replace({'shallow well':'shallow_well',
'machine dbh':'machine_dbh',
'rainwater harvesting':'rainwater_harvesting',
'hand dtw':'hand_dtw'}, inplace=True)
counts = data['subvillage'].value_counts()
counts = counts.loc[counts >=200]
counts = list(counts.index)
data.loc[~data['subvillage'].isin(counts), 'subvillage'] = 'other'
data.drop(columns=['region'], inplace=True)
data.fillna(inplace=True, value={'installer':'unknown','permit':False, 'funder':'unknown', 'public_meeting':False,
'scheme_management':'unknown', 'scheme_name':'unknown'})
data['lga_Njombe'] = data['lga'].replace({'Njombe':1})
data.loc[data['lga_Njombe']!=1, 'lga_Njombe'] = 0
data.drop(columns=['payment'], inplace=True)
data['basin'].replace({'Ruvuma / Southern Coast':'Ruvuma-Southern_Coast',
'Wami / Ruvu':'Wami-Ruvu'}, inplace=True)
data['date_recorded']= pd.to_datetime(data['date_recorded'])
data['date_recorded'].describe(datetime_is_numeric=True)
data['year']=data['date_recorded'].dt.year
data['month']=data['date_recorded'].dt.month
data.drop(columns=['wpt_name', 'num_private', 'recorded_by'], inplace=True)
data['amount_tsh'] = data['amount_tsh'].astype('int')
data['tsh_bin'] = pd.cut(data['amount_tsh'], [-1, 1, 20.0, 30.0, 50.0, 100.0, 250, 500.0, 1000.0, 2200.0, 70000.0, 500000],
labels=list(range(1,12)))
data['public_meeting'] = data['public_meeting'].map({True:1, False:0})
data['permit'] = data['permit'].map({True:1, False:0})
data['construction_year'].replace({0:1999}, inplace=True)
data['construction_year'] = pd.to_datetime(data['construction_year'], format='%Y')
data['years_old'] = data['date_recorded'].dt.year - data['construction_year'].dt.year
data.drop(columns=['quality_group', 'quantity_group'], inplace=True)
data.drop(columns=['waterpoint_type_group'],inplace=True)
counts2 = data['scheme_name'].value_counts()
counts2 = counts2.loc[counts2 >=200]
counts2 = list(counts2.index)
data.loc[~data['scheme_name'].isin(counts2), 'scheme_name'] = 'other'
counts3 = data['funder'].value_counts()
counts3 = counts3.loc[counts3 >=500]
counts3 = list(counts3.index)
data.loc[~data['funder'].isin(counts3), 'funder'] = 'other'
counts4 = data['installer'].value_counts()
counts4 = counts4.loc[counts4 >=500]
counts4 = list(counts4.index)
data.loc[~data['installer'].isin(counts4), 'installer'] = 'other'
data['popbins'] = pd.cut(data['population'], [-1,2,250,500,1000,2500,10000,40000], labels=list(range(1,8)))
counts5 = data['ward'].value_counts()
verybig = counts5.loc[counts5.between(200,400)].index
big = counts5.loc[counts5.between(100,200)].index
medium = counts5.loc[counts5.between(50,100)].index
small = counts5.loc[counts5.between(25,50)].index
verysmall = counts5.loc[counts5 <=25].index
data.loc[data['ward'].isin(verybig), 'ward'] = 'verybig'
data.loc[data['ward'].isin(big), 'ward'] = 'big'
data.loc[data['ward'].isin(medium), 'ward'] = 'medium'
data.loc[data['ward'].isin(small), 'ward'] = 'small'
data.loc[data['ward'].isin(verysmall), 'ward'] = 'verysmall'
data['longitude'] = data['longitude'].replace({0:np.random.choice(range(31,33))})
test_transformer(submit_X)
submit_X.shape, submit_X.columns
data2.shape, data2.columns
| Submissions/B/2D-Submission.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from itertools import combinations
import networkx as nx
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib inline
G = nx.complete_graph(3)
G.add_edges_from([(2, 3), (3, 4), (3, 6), (3,5), (4, 6), (5,6)])
nx.draw_networkx(G)
maximal_cliques_of_G = list(nx.find_cliques(G))
maximal_cliques_of_G
def make_first_structure_vector(list_of_maximal_cliques):
list_of_q_connected_components_at_qth_levels = [] #qth entry is the list of q-connected components
q_max = np.max([len(x) for x in list_of_maximal_cliques])-1
for q in range(q_max+1):
temp_list_of_max_cliques = list_of_maximal_cliques.copy()
q_connected_at_this_level = []
temp_list_of_max_cliques = [x for x in temp_list_of_max_cliques if len(x)>q]
while temp_list_of_max_cliques != []:
temp_list = []
temp_list.append(temp_list_of_max_cliques[0])
temp_list_of_max_cliques.remove(temp_list_of_max_cliques[0])
counter_for_check = -1
while len(temp_list)>counter_for_check:
counter_for_check = len(temp_list)
for clique in temp_list_of_max_cliques:
if len(clique) == q:
temp_list_of_max_cliques.remove(clique)
if any([len(list(set(clique_for_eval)&set(clique))) == q+1 for clique_for_eval in temp_list]):
temp_list.append(clique)
temp_list_of_max_cliques.remove(clique)
q_connected_at_this_level.append(temp_list)
list_of_q_connected_components_at_qth_levels.append(q_connected_at_this_level)
Q_vector = [len(x) for x in list_of_q_connected_components_at_qth_levels]
return Q_vector, list_of_q_connected_components_at_qth_levels
# +
Q_vector, q_connected_components = make_first_structure_vector(maximal_cliques_of_G)
print("The Q_vector is",Q_vector)
for index in range(len(q_connected_components)):
print("The {}^th connected components at the {}^th level is {}".format(index, index, q_connected_components[index]), end="\n" )
# -
G1 = G.copy()
G1.add_edge(4, 5)
maximal_cliques_of_G1 = list(nx.find_cliques(G1))
nx.draw_networkx(G1)
# +
Q_vector, q_connected_components = make_first_structure_vector(maximal_cliques_of_G1)
print("The Q_vector is",Q_vector)
for index in range(len(q_connected_components)):
print("The {}^th connected components at the {}^th level is {}".format(index, index, q_connected_components[index]), end="\n" )
# -
| .ipynb_checkpoints/Algebraic_topological_measures-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1 style="font-family:Georgia;font-size:2em"> Object-Oriented Programming</h1>
# <font style="font-family:Georgia;font-size:17px">
# <a href="https://pythoninternal.wordpress.com/2014/08/11/everythings-an-object/"><p>Everything in python is an object.</a>
# <p>Object-oriented programming in python refers to creation, modification, and use of custom classes,
# or rather, custom data types.
# <p>Objects are entities that have a set of data and several methods defined for that specific object. They are instances of
# a class. A class is a template for what an object is--a sort of abstraction. For example, the definition of a chair is
# the 'class' while an actual tangible chair would be an 'object'.
# Such a chair has data such as
# number_of_legs,
# size_of_seat,
# presence_of_cushion, etc.
# It could have functions such as
# increase_height(),
# recline(), etc.
# </font>
class Chair:
def __init__(self, name):
self.name = name # self. notation preceeding a variable indicates that the variable is local to the object
jack_chair = Chair("Jack's Chair")
jack_chair.name
# >Additional notes:
# The first argument that we pass into a class function is always an instance of the class, i.e., an object. It is a common convention to name the object `self`. The variables associated with the object are referenced using the notation `self.variable_name`.
# +
class Chair:
all_chairs = [] # static class variable
def __init__(self, name):
self.name = name
self.all_chairs.append(name)
jack_chair = Chair("Jack's Chair")
pat_chair = Chair("Pat's Chair")
pat_chair.all_chairs # by definition, prints all chairs' names available
# +
class Chair:
def __init__(self, num_legs, can_recl, price):
self.number_of_legs = int(num_legs)
self.can_recline = int(can_recl)
self.price = float(price)
self.is_reclined = False
def recline(self): # the self argument is used in each of the member methods.
if(self.can_recline):
self.is_reclined = True
print("Reclined!")
else:
self.is_reclined = False
print("Not a reclining chair. Can't recline.")
# File handling
x = open("chairs.csv","r")
office_chairs = list()
for line in x:
values = line.split(',')
number_of_legs, can_recline, price = values[0],values[1],values[2]
office_chairs.append(Chair(number_of_legs,can_recline,price))
for chair in office_chairs:
print(chair.is_reclined)
chair.recline()
print(chair.is_reclined)
| Session1/2. Object-Oriented Programming.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ------------- User's settings -------------
# +
# Location of digested data
input_directory = '/digested/'
# Location of saved trained model
model_directory = '/model_directory/'
# Desired location for outputs
output_directory = '/output_directory/'
# -
# # ------------- (semi)-Automatic -------------
# +
# %matplotlib inline
import keras
import pickle
from keras.layers import *
from keras.models import Sequential
import numpy
import os
import os.path
import matplotlib.pyplot
import pandas
import seaborn
import sklearn.metrics
import tensorflow
from tensorflow.contrib.tensorboard.plugins import projector
# -
# Configure GPU/CPU devices:
# +
# -------- If using Tensorflow-GPU: -------- #
configuration = tensorflow.ConfigProto()
configuration.gpu_options.allow_growth = True
configuration.gpu_options.visible_device_list = "0"
session = tensorflow.Session(config=configuration)
keras.backend.set_session(session)
# -------- If using Tensorflow (CPU) : -------- #
# configuration = tensorflow.ConfigProto()
# session = tensorflow.Session(config=configuration)
# keras.backend.set_session(session)
# -
if not os.path.exists(output_directory):
os.makedirs(output_directory)
# # Data queueing
# +
def training_data_generator(input_x, input_y, batch_size):
num_examples, num_labels = input_y.shape
label_indices = []
for i in range(num_labels):
indices = [j for j in range(num_examples) if input_y[j,i] > 0]
label_indices.append(indices)
print("Label",i,":",len(indices),"examples")
samples_per_label = int(batch_size / num_labels)
def generator():
while True:
x_samples = []
y_samples = []
for i in range(num_labels):
random.shuffle(label_indices[i])
indices = label_indices[i][0:samples_per_label]
x_samples.append( input_x[indices, ...] )
y_samples.append( input_y[indices, ...] )
x_samples = numpy.concatenate( x_samples )
y_samples = numpy.concatenate( y_samples )
batch_indices = numpy.arange(x_samples.shape[0])
numpy.random.shuffle(batch_indices)
x_samples = x_samples[batch_indices, ...]
y_samples = y_samples[batch_indices, ...]
yield (x_samples, y_samples)
return generator()
def prediction_data_generator(input_x, input_y, batch_size):
num_examples, num_labels = input_y.shape
steps = int(num_examples / batch_size)
def generator():
i = 0
while True:
start = i*batch_size
end = (i+1)*batch_size
x_sample = input_x[start:end, ...]
y_sample = input_y[start:end, ...]
yield (x_sample, y_sample)
i = i + 1 if i < steps else 0
print("Prediction steps:",steps)
return generator(), steps
# +
# This function to normalize illumination discrepancy across images
def min_max_norm(x, minimum=None, maximum=None):
channels = x.shape[-1]
if minimum is None and maximum is None:
minimum = []
maximum = []
for channel in range(channels):
minimum.append( x[..., channel].min() )
maximum.append( x[..., channel].max() )
result = numpy.zeros_like(x)
for ch in range(channels):
result[..., ch] = 100.0*( (numpy.ndarray.astype(x[..., ch], numpy.float32) - minimum[ch])/(maximum[ch] - minimum[ch]) )
return (result, minimum, maximum)
# -
# # Load data:
# +
training_x = numpy.load(os.path.join(input_directory, "training_x.npy"))
training_y = numpy.load(os.path.join(input_directory, "training_y.npy"))
# input_directory = "/path/to/other/input_directory/if/needed"
testing_x = numpy.load(os.path.join(input_directory, "testing_x.npy"))
testing_y = numpy.load(os.path.join(input_directory, "testing_y.npy"))
# +
print("Loading training data")
# Use this function to normalize signal intensities across images
training_x, pix_min, pix_max = min_max_norm(training_x)
training_generator = training_data_generator(training_x, training_y, 32)
print(training_x.shape, training_y.shape)
# +
print("Loading test data")
# Use this function to normalize signal intensities across images
testing_x, _, _ = min_max_norm(testing_x, pix_min, pix_max)
testing_generator, testing_steps = prediction_data_generator(testing_x, testing_y, 32)
print(testing_x.shape)
# -
# # Load trained model:
# (can also load checkpoints)
model = keras.models.load_model( os.path.join(model_directory, 'model.h5') )
model.load_weights(os.path.join(model_directory, 'model.h5'))
# # Evaluate testing set
model.evaluate_generator(
generator=testing_generator,
steps=256
)
# # Extract the most crucial layer
layers = model.layers
model.summary()
# Look for the densely/fully connected layer nearest to the classier, which is the one that has the shape of (None, number-of-classes)
#
# ==================================================================
#
# Example 1: in case of classification of 7 classes, the last few layers are:
#
# _________________________________________________________________
# dense_1 (Dense) (None, 1024) 943820
# _________________________________________________________________
# dropout_1 (Dropout) (None, 1024) 0
# _________________________________________________________________
# dense_2 (Dense) (None, 7) 7175
# _________________________________________________________________
# activation_1 (Activation) (None, 7) 0
#
#
# then look for the layer dense_1 , which has a shape of (None, 1024)
#
# ==================================================================
#
# Example 2: in case of classification of 5 classes, the last few layers are:
#
# activation_49 (Activation) (None, 8, 8, 2048) 0
# _________________________________________________________________
# avg_pool (AveragePooling2D) (None, 1, 1, 2048) 0
# _________________________________________________________________
# global_average_pooling2d_1 (Glob (None, 2048) 0
# _________________________________________________________________
# dense_2 (Dense) (None, 5) 10245
#
# then look for the layer global_average_pooling2d_1 , which has a shape of (None, 2048)
print(layers[-4])
abstract_model = None # Clear cached abstract_model
abstract_model = Sequential([layers[-4]])
extracted_features = abstract_model.predict_generator(
generator=testing_generator,
steps=256)
# # Metadata for embeddings
# +
print('Converting numeric labels into class names...')
class_names = pickle.load(open(os.path.join(input_directory, "class_names.sav"), 'rb'))
def save_metadata(file):
with open(file, 'w') as f:
for i in range(test_y.shape[0]):
f.write('{}\n'.format( class_names[test_y[i]] ))
save_metadata( os.path.join(output_directory, 'metadata.tsv') )
print('Done.')
# -
# # Predicted values in .TXT
# To be uploaded and viewed on http://projector.tensorflow.org
numpy.savetxt( os.path.join(output_directory, 'table_of_features.txt' ), extracted_features, delimiter='\t')
# # Note:
#
# Once finished, open http://projector.tensorflow.org on web-browser.
#
# Click "Load data" on the left panel.
#
# - Step 1: Load a TSV file of vectors >> Choose file: 'table_of_features.txt'
#
# - Step 2: Load a TSV file of metadata >> Choose file: 'metadata.tsv'
#
# Hit ESC or click outside the load data window to dismiss.
# # Predicted values in .NPY
# Used for generating Tensorboard embeddings to be viewed locally on http://localhost:6006
numpy.save( os.path.join(output_directory, 'table_of_features.npy' ), extracted_features )
# +
extracted_features = numpy.load( 'table_of_features.npy' )
embedding_var = tensorflow.Variable(extracted_features)
embedSess = tensorflow.Session()
# save variable in session
embedSess.run(embedding_var.initializer)
# save session (only used variable) to file
saver = tensorflow.train.Saver([embedding_var])
saver.save(embedSess, 'tf.ckpt')
summary_writer = tensorflow.summary.FileWriter('./')
config = tensorflow.contrib.tensorboard.plugins.projector.ProjectorConfig()
embedding = config.embeddings.add()
embedding.tensor_name = embedding_var.name
embedding.metadata_path = 'metadata.tsv' # this metadata_path need to be modified later. See note.
tensorflow.contrib.tensorboard.plugins.projector.visualize_embeddings(summary_writer, config)
embedSess.close()
# -
# # Note:
# Tensorboard embeddings files will be saved in the same location with this script.
#
# Collect the following files into one folder:
#
# - metadata.tsv
# - checkpoint
# - projector_config.pbtxt
# - tf.ckpt.index
# - tf.ckpt.meta
# - tf.ckpt.data-00000-of-00001
#
# Open with any text editor : "projector_config.pbtxt"
#
# "/path/to/logdir/metadata.tsv" has to be specified, CANNOT be relative path "./metadata.tsv", nor "~/metadata.tsv"
#
# Then type command in terminal: tensorboard --logdir="/path/to/logdir"
#
# Next, open web-browser, connect to http://localhost:6006
# # Plot categorical accuracy and loss
metrics = pandas.read_csv(os.path.join(model_directory, 'training.csv') )
print(metrics)
matplotlib.pyplot.plot(metrics["acc"])
matplotlib.pyplot.plot(metrics["val_acc"])
matplotlib.pyplot.plot(metrics["loss"])
matplotlib.pyplot.plot(metrics["val_loss"])
# # Confusion matrix
# +
predicted = model.predict(
batch_size=50,
x=testing_x
)
predicted = numpy.argmax(predicted, -1)
expected = numpy.argmax(testing_y[:, :], -1)
# +
confusion = sklearn.metrics.confusion_matrix(expected, predicted)
confusion = pandas.DataFrame(confusion)
matplotlib.pyplot.figure(figsize=(12, 8))
seaborn.heatmap(confusion, annot=True)
matplotlib.pyplot.savefig( os.path.join(output_directory, 'confusion_matrix.eps') , format='eps', dpi=600)
| STEP_4_Test_and_Visualization_built-in_CNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Generate circles using evolutionary algorithm
#
# The code is based on
# https://github.com/tjwei/play_nsfw,
# which is based on https://scturtle.me/posts/2014-04-18-ga.html
#
# It depends on DEAP https://deap.readthedocs.io/en/master/
import multiprocessing
from deap import base, creator, tools, algorithms
from PIL import Image, ImageDraw
from random import randint
import skimage
SIZE_X, SIZE_Y = 256, 256
NUMBER_OF_TRIANGLES = 50
POPULATION = 50
NGEN = 4000
POLY = 3
def gen_one_triangle():
return (tuple([(randint(0, SIZE_X), randint(0, SIZE_Y)) for i in range(POLY)]),
randint(0,255), randint(0,255), randint(0,255), randint(0,128))
# +
creator.create("Fitness", base.Fitness, weights=(1.0,)) # maximize fitness
creator.create("Individual", list, fitness=creator.Fitness) # individual class
toolbox = base.Toolbox()
toolbox.register("attr", gen_one_triangle) # the above function
toolbox.register("individual", tools.initRepeat, # initialization of individual
creator.Individual, toolbox.attr, NUMBER_OF_TRIANGLES)
toolbox.register("population", tools.initRepeat, # initialization of population
list, toolbox.individual)
# -
import numpy as np
target_im = Image.open('sample_images/800px-Meisje_met_de_parel.jpg').crop((0,100,800,900)).resize((256,256), Image.LANCZOS)
target_array = np.array(target_im, dtype='float')/255.
target_im
# +
def triangles_to_image(triangles):
im = Image.new('RGB', (SIZE_X, SIZE_Y), (0, 0, 0))
for tri in triangles:
mask = Image.new('RGBA', (SIZE_X, SIZE_Y))
draw = ImageDraw.Draw(mask)
draw.polygon(tri[0], fill=tri[1:])
im.paste(mask, mask=mask)
del mask, draw
return im
def evaluate(t2):
output_im = triangles_to_image(t2)
output_array = np.array(output_im, dtype='float')/255.
score = skimage.measure.compare_psnr(output_array, target_array, 1.)
#score = skimage.measure.compare_ssim(output_array, target_array, data_range=1., multichannel=True)
return (score,)
# -
def mutate(triangles):
e0 = triangles.fitness.values
for i in range(10):
tid = randint(0, NUMBER_OF_TRIANGLES - 1)
oldt = triangles[tid]
p = randint(-1, 2 * POLY + 4 - 1)
if p == -1:
tid2 = randint(0, NUMBER_OF_TRIANGLES - 1)
triangles[tid], triangles[tid2] = triangles[tid2], oldt
else:
t = list(oldt)
if p < 2 * POLY:
points = list(t[0])
pnt = list(points[p // 2])
if p%2 == 0:
pnt[0] = randint(0, SIZE_X)
else:
pnt[1] = randint(0, SIZE_Y)
points[p // 2] = tuple(pnt)
t[0] = tuple(points)
else:
p -= 2 * POLY - 1
t[p] = randint(0, 255)
triangles[tid] = tuple(t)
if evaluate(triangles) > e0:
break
else:
if p == -1:
triangles[tid], triangles[tid2] = oldt, triangles[tid]
else:
triangles[tid] = oldt
return [triangles]
toolbox.register("evaluate", evaluate)
toolbox.register("mate", tools.cxTwoPoint) # crossover
toolbox.register("mutate", mutate) # mutation
toolbox.register("select", tools.selTournament, tournsize=3)
from IPython.display import display, clear_output
import numpy as np
class ipyHOF(tools.HallOfFame):
def __init__(self, *args, **kwargs):
super().__init__(*args, **kwargs)
self.last_displayed_fitness = -1
def insert(self, item):
if item.fitness.values[0] >= self.last_displayed_fitness+0.001:
self.last_displayed_fitness = item.fitness.values[0]
clear_output(True)
display(triangles_to_image(item))
super().insert(item)
# +
pop = toolbox.population(n=POPULATION)
hof = ipyHOF(3)
stats = tools.Statistics(lambda ind: ind.fitness.values)
stats.register("std", np.std)
stats.register("max", np.max)
stats.register("avg", np.mean)
stats.register("min", np.min)
# For multiprocessing
pool = multiprocessing.Pool()
toolbox.register("map", pool.map)
# -
try:
pop, log = algorithms.eaSimple(pop, toolbox, cxpb=0.5, mutpb=0.1, ngen=NGEN,
stats=stats, halloffame=hof, verbose=True)
except KeyboardInterrupt:
display(triangles_to_image(hof[0]))
# +
#triangles_to_image(hof[0]).save('sample_results/triangles-50-ssim.png')
# -
img1 = triangles_to_image(hof[0])
img2 = Image.open('sample_results/circle-30.png')
img3 = np.clip((np.array(img1, dtype='float')+np.array(img2, dtype='float'))/2, 0, 255)
img3 = Image.fromarray(img3.astype('uint8'))
img3
| deap/Triangles.ipynb |
// ---
// title: "Mapping A Function To A Collection"
// author: "<NAME>"
// date: 2017-12-20T11:53:49-07:00
// description: "Mapping a function to a collection using Scala."
// type: technical_note
// draft: false
// jupyter:
// jupytext:
// text_representation:
// extension: .scala
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Apache Toree - Scala
// language: scala
// name: apache_toree_scala
// ---
// ## Preliminaries
import scala.collection.mutable.ArrayBuffer
// ## Create Collection
// Create an array of strings
var birds = ArrayBuffer("Hawk", "Condor", "Eagle", "Pigeon")
// ## Create Function
// Create a function that returns the length of a string
val getLength = (i: String) => i.length
// ## Map The Function To The Collection
// Map the function to the array
birds.map(getLength)
// ## Map An Anonymous Function To The Collection
// Map the anonymous function to the collection
birds.map(_.toUpperCase)
| docs/scala/basics/mapping_a_function_to_a_collection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.8 64-bit (''flowing'': conda)'
# name: python3
# ---
# # SiouxFalls - (Parametric) Price of Anarchy
# ## 1. Imports and data readin
# First we load the paminco package and read the SiouxFalls data.
# The data was taken from [here](https://github.com/bstabler/TransportationNetworks) and converted to our format of choice: ``XML``.
# The data for SiouxFalls comes with paminco and can be easily loaded:
# +
import paminco
sioux = paminco.load_sioux()
# -
# By default the edge cost equal the link travel time: $F_e = l_e$.
# The link travel time is defined as
#
# $$
# \begin{equation*}
# l_e(x_e) = \text{fft}_e \cdot \left( 1 + B_e \cdot \left(\frac{x}{\text{cap}_e}\right) ^ {p_e} \right)
# \end{equation*}
# $$
# ## 2. Fixed Price of Anarchy with Frank-Wolfe
# We find minimum cost flows that coincide with user equilibrium and system optimum if we transform the edge cost by:
#
# $$
# \begin{align}
# \text{User equilibrium:} \quad F_e &= \int_0^{x_e} l_e(s) ds \\
# \text{System optimum:} \quad F_e &= x_e \cdot l_e \\
# \end{align}
# $$
# +
import copy
# Calculate user equilibrium -> F_e = integral_0^(x_e) l_e(s)ds
sioux_ue = copy.deepcopy(sioux)
sioux_ue.cost.integrate(inplace=True)
fw_ue = paminco.NetworkFW(sioux_ue)
fw_ue.run(max_iter=500)
# Calculate system optimum -> F_e = x_e * l_e
sioux_so = copy.deepcopy(sioux)
sioux_so.cost.times_x(inplace=True)
fw_so = paminco.NetworkFW(sioux_so)
fw_so.run(max_iter=500)
# -
# The [Price of Annarchy](https://en.wikipedia.org/wiki/Price_of_anarchy) (PoA) measures the inefficency of the network utilization due to selfish behaviour of the participants.
# It is calculated as the ratio of the total (or equivalently average) travel time for all particpants in the user equilbrium to that of the system optimum:
#
# $$
# \begin{equation*}
# \text{PoA} = \frac{C(\text{UE})}{C(\text{SO})},
# \end{equation*}
# $$
#
# where $C$ is the total system travel time (TSTT):
#
# $$
# \begin{equation*}
# \text{TTST}(\mathbf{x}) = x_e \cdot l_e,
# \end{equation*}
# $$
#
# i.e., the objective function of the system optimal minimum cost flow.
def TTST(x):
return (x * sioux.cost(x)).sum()
# The Price of Anarchy for SiouxFalls is about ``3.8 percent``:
poa = TTST(fw_ue.x) / TTST(fw_so.x)
poa
# ## 3. Paramtric Price of Anarchy with MCA
# The above example allowed to compute system inefficiency for a fixed demand $\mathbf{B}$.
#
# However, it is of interest how the POA varies by the demand multiplier $\lambda$.
# We can achieve this with the ``MCA`` algorithm by calculating parametric user equilibira flows and parametric system optima:
# +
sioux2 = copy.deepcopy(sioux)
sioux2.set_demand(("20", "3", 100000))
sioux_ue = copy.deepcopy(sioux2)
sioux_ue.cost.integrate(inplace=True)
mca_ue = paminco.MCA(sioux_ue)
mca_ue.run()
sioux_so = copy.deepcopy(sioux2)
sioux_so.cost.times_x(inplace=True)
mca_so = paminco.MCA(sioux_so)
mca_so.run()
# -
# For any demand multiplier in $\lambda \in [0, 1]$, we can now compute a price of anarchy:
# +
import numpy as np
lambdas = np.linspace(1e-5, 1, 100)
cost_user_equilibira = np.array([TTST(mca_ue.flow_at(l)) for l in lambdas])
cost_system_optima = np.array([TTST(mca_so.flow_at(l)) for l in lambdas])
paramtric_poa = cost_user_equilibira / cost_system_optima
# +
import matplotlib.pyplot as plt
fig, ax = plt.subplots(figsize=(10, 6))
ax.plot(lambdas, paramtric_poa, color="black")
ax.set_xlabel("$\lambda$", fontsize=16)
ax.set_ylabel("PoA", fontsize=16)
ax.set_title("Parametric Price of Anarchy", fontsize=16)
| docs/source/user_guide/applications/traffic/sioux_par_poa.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import torch
dtype = torch.float
device = torch.device('cpu')
N = 64
D_in = 1000
H = 100
D_out = 10
x = torch.randn((N,D_in),device=device,dtype=dtype)
y = torch.randn((N,D_out),device=device,dtype=dtype)
model = torch.nn.Sequential(
torch.nn.Linear(D_in,H),
torch.nn.ReLU(),
torch.nn.Linear(H, D_out)
)
loss_fn = torch.nn.MSELoss(reduction="sum")
LR = 1e-4
T = 10
for t in range(T):
y_pred = model(x)
loss = loss_fn(y_pred,y)
print(t,loss.item())
model.zero_grad() #zero gradient, why?
loss.backward() #computes w1.grad and w2.grad
with torch.no_grad():
for param in model.parameters():
param -= LR*param.grad
| tutorial3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <b>Se a reta que passa pelos pontos $A(-2,5,1)$, $B(1,3,0)$ é paralela à reta determinada por $C(3,-1,-1)$, $D(0,m,n)$, podemos afirmar que os valores de $m$ e $n$ são, respectivamente: </b>
# <b>Se as retas são pararalelas, elas possuem o mesmo vetor $\vec{AB}$</b>
# $\vec {AB} = B - A$
# $\vec{AB}(1,3,0) - (-2, 5, 1)$<br><br>
# $\vec{AB}(1+2, 3-5, 0 -1)$<br><br>
# $\vec{AB}(3, -2, -1)$
# <b>Equação vetorial da reta que passa pelos pontos $A(-2,5,1)$, $B(1,3,0)$ </b>
# $(-2,5,1) = (1, 3, 0) + t(3,-2,-1)$
# <b>Equação vetorial da reta que passa pelos pontos $C(3,-1,-1)$, $D(0,m,n)$ </b>
# $(3, -1, -1) = (0, m, n) + t(3, -2, -1)$
# <b>Montando a equação paramétrica da reta que passa pelos pontos $C(3,-1,-1)$, $D(0,m,n)$</b>
# $\, 3 = 0 + 3t$<br>
# $-1 = m -2t$<br>
# $-1 = n - t$<br>
# <b>Achando o valor de $t$</b>
# $3 = 3t$<br>
# $\frac{3}{3} = t$<br>
# $t = 1$
# <b>Substituindo o valor de $t$ para achar $m$</b>
# $-1 = m - 2\cdot 1$<br>
# $-1 = m - 2$<br>
# $-1 + 2 = m $<br>
# $1 = m$
# <b>Substituindo o valor de $t$ para achar $n$</b>
# $-1 = n - 1$<br>
# $-1 + 1 = n$<br>
# $n = 0$
# $m = 1$ e $n = 0$
| Lista de retas/02.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from ImageData import ScanAssets
a = ScanAssets("../images/")
a.do(None)
a.data[212].data[0]
import numpy as np
opencvImage = np.array(a.data[212].data[0])
import cv2
ret, threshed_img = cv2.threshold(opencvImage, 40, 255, cv2.THRESH_BINARY)
contours, hier = cv2.findContours(threshed_img, cv2.RETR_TREE, cv2.CHAIN_APPROX_SIMPLE)
# +
import math
maxarea = 0
max2 = 0
#cv2.cvtColor(img, cv2.COLOR_GRAY2RGB, img)
(imWidth, imHeight) = opencvImage.shape
imArea = imWidth * imHeight
#framemax
print("Contours: "+str(len(contours)))
for c in contours:
# get the bounding rect
x, y, w, h = cv2.boundingRect(c)
# draw a green rectangle to visualize the bounding rect
cv2.rectangle(opencvImage, (x, y), (x+w, y+h), (255, 255, 255), 2)
area = w * h
print (area)
if area > maxarea:
max2 = maxarea
maxarea = area
framemax = (x,y, w, h)
elif area > max2:
max2 = area
frame2 = (x, y, w, h)
threshold = 0.05
spaceA = (1.0 / imArea * maxarea)
spaceB = (1.0 / imArea * max2)
if spaceA >= threshold:
frame2 = framemax
print("Framemax has"+str(spaceA)+" percent space!")
elif spaceB >= threshold:
framemax = frame2
print("Frame2 has"+str(spaceB)+" percent space!")
if framemax[1] > frame2[1]:
uborder = cv2.line(opencvImage, (framemax[0], framemax[1]+framemax[3]), (framemax[0]+framemax[2], framemax[1]+framemax[3]), (255, 0, 0), 1)
oborder = cv2.line(opencvImage, (frame2[0], frame2[1]),(frame2[0]+frame2[2], frame2[1]), (255, 0, 0), 1)
size = opencvImage.shape
cropX = 0
cropWidth = size[1]
cropY = frame2[1]
cropHeight = 0 + (framemax[1]+framemax[3])
if cropY >= imHeight:
print("Fehler 1")
if cropHeight >= imHeight:
print("Fehler 2")
if cropWidth >= imWidth:
#cropWidth = imWidth - 1
print("Fehler 3! KORRIGIERT")
crop_img = opencvImage[cropY : cropHeight, cropX : cropWidth]
partHeight, partWidth = crop_img.shape
part_img = crop_img[math.floor(partHeight * 0.20): math.floor(partHeight * 0.85), math.floor(partWidth * 0.15): math.floor(partWidth * 0.80)]
# -
from PIL import Image
plt = Image.fromarray(crop_img)
plt
plt = Image.fromarray(part_img)
plt
| notebooks/Working with OpenCV and Pillow.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Issues:
#
# - [ ] inflated \# units (e.g. parkmerced) due to double counting or counting non-residential units
# - [x] properties in some assessor roll years and not others
# - **solution**: take record from closest year, only consider evictions >= 2007
# - [x] properties with multiple rent control eligibility use codes:
# - **solution**: count how many of these there are, take the max use code
# - [ ] zero-unit buildings in new construction
# - **solution**: year_built < 2010, or units > 0
# - [ ] properties with multiple year-built's
# - **solution**: take the max year built, which will give a conservative estimate w/r/t rent control
# - [x] year-built = 0
# - **solution**: year_built > 1800
# - [ ] condo conversion or parcel splits after eviction but before earliest assessor record
# - SRES --> MRES would overcount MRES evictions
# - MRES --> SRES would undercount MRES evictions
# - **solution**: only count evictions after 2007
# - many of these are the "0000 0000 000000000" values in assessor rolls
# - [ ] rent controlled properties more likely to be in state of disrepair and therefore require demolition/capital improvements
# - **solution**: fit hedonic and control for stddev above/below predicted value
# - [ ] petition applies to multiple units
# - **solution**: use new eviction .csv with unit counts
import pandas as pd
import numpy as np
from matplotlib import pyplot as plt
import seaborn as sns
import statsmodels.formula.api as smf
# %matplotlib inline
# ### Load assessor universe
# +
# asr_all = pd.read_csv('../data/assessor_2007-2018_clean_w_none_sttyps.csv')
# -
asr = pd.read_csv('../data/asr_grouped_by_yr.csv')
asr['any_ev'] = (asr['ev_count'] > 0).astype(int)
asr['any_ev_07'] = (asr['ev_count_post_07'] > 0).astype(int)
asr['pre_1980'] = (asr['year_built_max'] < 1980)
asr['built_1980'] = None
asr.loc[asr['pre_1980'], 'built_1980'] = 'before'
asr.loc[~asr['pre_1980'], 'built_1980'] = 'after'
asr['ev_per_unit'] = asr['ev_count'] / asr['total_units']
asr['ev_per_unit_since_07'] = asr['ev_count_post_07'] / asr['total_units']
asr
asr.columns
# ### Load eviction data
ev = pd.read_csv('../data/ev_matched.csv')
# ### Eviction type counts by built year
# Retain only evictions since 2007
ev = ev[ev['year'] >= 2007]
# % evictions matched to assessor records
len(ev[~pd.isnull(ev['asr_index'])]) / len(ev)
ev = ev.merge(asr, left_on='asr_index', right_on='index', suffixes=('_ev', '_asr'))
ev = ev[ev['any_rc_eligibility'] == 1]
ev.loc[pd.isnull(ev['type']), 'type'] = 'unknown'
type_counts = ev.groupby(['built_1980', 'type']).agg(count=('index_ev', 'nunique')).reset_index()
pre_sums = type_counts.groupby('built_1980')['count'].sum()
# +
type_counts = type_counts.pivot(index='type', columns='built_1980', values='count')
type_counts['pct_after'] = type_counts['after'] / pre_sums['after']
type_counts['pct_before'] = type_counts['before'] / pre_sums['before']
# -
# #### 8x rate of OMI's, but this is prob due to structural differences
type_counts.sort_values('pct_before', ascending=False)
ev['ev_type_cat'] = 'breach of lease'
ev.loc[ev['type'].isin([
'OMI', 'Capital Improvement', 'ELLIS', 'Condo Conversion', 'Substantial Rehabilitation',
'Lead Remediation', 'Good Samaritan Tenancy Ends',
'Development Agreement', 'Demolition']), 'ev_type_cat'] = 'no fault'
ev.loc[ev['type'].isin(['unknown', 'Other']), 'ev_type_cat'] = 'unknown/Other'
cat_counts = ev.groupby(['built_1980', 'ev_type_cat']).agg(count=('index_ev', 'nunique')).reset_index()
cat_counts
cat_counts = cat_counts.pivot(index='ev_type_cat', columns='built_1980', values='count')
cat_counts['pct_after'] = cat_counts['after'] / pre_sums['after']
cat_counts['pct_before'] = cat_counts['before'] / pre_sums['before']
cat_counts
# ### Mean differences
# #### Evictions post-2007:
# +
mean_diffs = asr[
(asr['year_built_max'] < 2007) &
(asr['year_built_min'] > 0)].groupby(['any_rc_eligibility', 'pre_1980']).agg(
mean_any_ev=('any_ev_07', 'mean'),
total_addresses=('index', 'count'),
total_units=('total_units', 'sum'),
total_evictions=('ev_count_post_07', 'sum'),
)
mean_diffs['units_per_address'] = mean_diffs['total_units'] / mean_diffs['total_addresses']
mean_diffs['evictions_per_address'] = mean_diffs['total_evictions'] / mean_diffs['total_addresses']
mean_diffs['evictions_per_unit'] = mean_diffs['total_evictions'] / mean_diffs['total_units']
mean_diffs
# -
# #### All Evictions
# +
mean_diffs = asr[
(asr['year_built_max'] < 2007) &
(asr['year_built_min'] > 0)].groupby(['any_rc_eligibility', 'pre_1980']).agg(
mean_any_ev=('any_ev_07', 'mean'),
total_addresses=('index', 'count'),
total_units=('total_units', 'sum'),
total_evictions=('ev_count', 'sum'),
)
mean_diffs['units_per_address'] = mean_diffs['total_units'] / mean_diffs['total_addresses']
mean_diffs['evictions_per_address'] = mean_diffs['total_evictions'] / mean_diffs['total_addresses']
mean_diffs['evictions_per_unit'] = mean_diffs['total_evictions'] / mean_diffs['total_units']
mean_diffs
# -
# ### Plots
rc_pop = asr[
(asr['any_rc_eligibility'] == 1) & (asr['year_built_max'] > 1500) &
(asr['year_built_max'] < 2500) & (asr['total_units'] > 0)]
# +
yr_vs_ev = rc_pop.groupby('year_built_max').agg({
'ev_per_unit':'mean',
'ev_per_unit_since_07':'mean'
}).reset_index()
yr_vs_ev1 = yr_vs_ev[(yr_vs_ev['year_built_max'] < 1980) &
(yr_vs_ev['year_built_max'] >= 1953)]
yr_vs_ev2 = yr_vs_ev[(yr_vs_ev['year_built_max'] >= 1980) &
(yr_vs_ev['year_built_max'] <= 2007)]
# -
fig, ax = plt.subplots(figsize=(13,7))
sns.regplot('year_built_max', 'ev_per_unit_since_07', yr_vs_ev1, ax=ax, truncate=True, label='rent controlled')
sns.regplot('year_built_max', 'ev_per_unit_since_07', yr_vs_ev2, ax=ax, truncate=True, label='non-rent controlled')
ax.axvline(1979.5, ls=':', c='r')
ax.legend()
_ = ax.set_xlabel("property built-year", fontsize=16)
_ = ax.set_ylabel("avg.\nevictions/unit\nper year", fontsize=16, rotation=0, labelpad=70)
_ = ax.set_title("SF Eviction Rates (2007-2017)\nfor Multi-family Residential Properties\n"
"(incl. SRO's, excl. TIC's)", fontsize=20)
ax.set_ylim((-0.005, 0.05))
ax.annotate('rent control \nbuilt-year threshold', xy=(1979, 0.04), xycoords='data',
xytext=(0.3, 0.8), textcoords='axes fraction',
arrowprops=dict(facecolor='black',frac=0.05, width=0.5, headwidth=10),
horizontalalignment='center', verticalalignment='center', fontsize=12
)
# ### Fit Hedonic regression
asr_all = pd.read_csv('./evictions/data/assessor_2007-2018_clean_w_none_sttyps.csv')
asr_all['total_value'] = asr_all['RP1LNDVAL'] + asr_all['RP1IMPVAL']
asr_all.loc[pd.isnull(asr_all['RP1NBRCDE']), 'RP1NBRCDE'] = 'unknown'
asr_grouped_by_yr = asr_all.groupby(['asr_yr', 'house_1', 'house_2', 'street_name', 'street_type']).agg(
total_units=('UNITS', 'sum'),
diff_unit_counts=('UNITS', 'nunique'),
min_units=('UNITS', 'min'),
diff_bldg_types=('bldg_type', 'nunique'),
bldg_type_min=('bldg_type', 'min'),
bldg_type_max=('bldg_type', 'max'),
diff_rc_eligibility=('rc_eligible', 'nunique'),
any_rc_eligibility=('rc_eligible', 'max'),
diff_years_built=('YRBLT', 'nunique'),
year_built_min=('YRBLT', 'min'),
year_built_max=('YRBLT', 'max'),
total_value=('total_value', 'sum'),
total_beds=('BEDS', 'sum'),
total_baths=('BATHS', 'sum'),
mean_stories=('STOREYNO', 'mean'),
total_sqft=('SQFT', 'sum'),
nbd=('RP1NBRCDE', pd.Series.mode),
total_rooms=('ROOMS', 'sum'),
total_area=('LAREA', 'sum')
).reset_index()
asr_grouped_by_yr['nbd'] = asr_grouped_by_yr['nbd'].apply(lambda x: list(x)[0] if type(x) == np.ndarray else x)
asr_grouped_by_yr['yr_built_since_1900'] = asr_grouped_by_yr['year_built_max'] - 1900
df_hed = asr_grouped_by_yr[
(asr_grouped_by_yr['any_rc_eligibility'] == 1) &
(asr_grouped_by_yr['total_units'] > 0) &
(asr_grouped_by_yr['year_built_max'] >= 1950) &
(asr_grouped_by_yr['year_built_max'] <= 2010) &
(asr_grouped_by_yr['total_sqft'] > 0) &
# (asr_grouped_by_yr['total_beds'] > 0)
(asr_grouped_by_yr['total_baths'] > 0) &
(asr_grouped_by_yr['total_rooms'] > 0) &
(asr_grouped_by_yr['mean_stories'] > 0) &
(asr_grouped_by_yr['total_area'] > 0)
]
hedonic = smf.ols(
'total_value ~ total_sqft + np.log1p(total_beds) + np.log1p(total_baths) + np.log1p(total_units) + mean_stories + total_area + '
'total_rooms + yr_built_since_1900 + C(asr_yr) + nbd', data=df_hed
).fit()
df_hed['hedonic_resid'] = hedonic.resid
print(hedonic.summary())
# ### Fitting the sharp RD
# Control variables to add:
# - rent burden?
# - stddev prop value
bandwidth = 27
df = asr[
(asr['any_rc_eligibility'] == 1) & (asr['year_built_max'] > 1980 - bandwidth) &
(asr['year_built_max'] < 1980 + bandwidth) & (asr['total_units'] > 0)]
df['rent_control'] = False
df.loc[df['pre_1980'] == True, 'rent_control'] = True
df['year_built_centered'] = df['year_built_max'] - 1980
df.groupby('pre_1980').agg(
mean_any_ev=('any_ev', 'mean'),
total_addresses=('index', 'count'),
total_units=('total_units', 'sum'),
total_evictions=('ev_count', 'sum'),
ev_per_unit=('ev_per_unit', 'mean')
)
df.columns
df = pd.merge(
df,
df_hed[[
'asr_yr', 'house_1', 'house_2', 'street_name', 'street_type', 'total_rooms',
'total_value', 'total_area', 'total_sqft', 'nbd', 'total_baths', 'hedonic_resid']],
on=['asr_yr', 'house_1', 'house_2', 'street_name', 'street_type'])
rd = smf.ols(
"ev_per_unit_since_07 ~ rent_control + year_built_centered*rent_control + "
"np.log1p(total_value):np.log(total_sqft) + np.log(total_units)",
data=df)
fitted = rd.fit()
print(fitted.summary())
fitted.params[1]
# ### Potential evictions
# +
units_by_yr = asr[
(asr['any_rc_eligibility'] == 1) &
(asr['year_built_max'] > 1900) &
(asr['year_built_max'] < 2100)].groupby('year_built_max').agg({'total_units': 'sum'}).reset_index()
fig, ax = plt.subplots(figsize=(13,8))
ax.scatter(units_by_yr['year_built_max'], units_by_yr['total_units'], s=25, facecolors='none', edgecolors='r')
ax.plot(units_by_yr['year_built_max'], units_by_yr['total_units'], lw=1, c='k', )
_ = ax.set_xlabel("year built", fontsize=16)
_ = ax.set_ylabel("# new units", fontsize=16)
_ = ax.set_title("SF New Construction: Rent-control eligible use-codes", fontsize=20)
# -
rc_pop = asr[(asr['any_rc_eligibility'] == 1) & (asr['year_built_max'] > 1979)]
rc_pop = rc_pop.groupby('year_built_max').agg({'total_units': 'sum'})
rc_pop.index.name = "new rent control year-built cutoff"
rc_pop['cumulative_units'] = rc_pop['total_units'].cumsum()
rc_pop['potential_evictions'] = rc_pop['cumulative_units'] * fitted.params[1]
rc_pop['pct_growth'] = rc_pop['potential_evictions'] / ev_per_year
rc_pop
# ### RDD package
import rdd
| notebooks/analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# <img style="float: left;" alt="Drawing" src="./figures/3Di_beeldmerk_RGB.png" width="100"/>
# ## From starting a 3Di-simulation to downloading and analysing the results in a jupyter notebook
# Welcome! In this notebook we will show you how to start a <a href="https://3diwatermanagement.com/">3Di</a>-simulation in a jupyter notebook by using the API-v3. In addition, we will show you how to download, visualize and analyse the results of the 3Di-simulation.
#
# The following steps will be taken according to an example of an 3Di model:
# - **step 1:** Creating a 3Di-simulation by using the threedi-api
# - **step 2:** Adding events to this 3Di-simulation by using the threedi-api
# - **step 3:** Running the 3Di-simulation by using the threedi-api
# - **step 4:** Downloading the results of the 3Di-simulation
# - **step 5:** Analysing the results of the simulation
# **Step 1: Starting a 3Di-simulation by using the threedi-api**
# Importing all required packages:
from datetime import datetime
from getpass import getpass
import pandas as pd
import requests
import json
from openapi_client import ApiException, SimulationsApi, OrganisationsApi
from openapi_client.api import AuthApi
from openapi_client.api import ThreedimodelsApi
from openapi_client.models import RasterEdit
from threedi_api_client.threedi_api_client import ThreediApiClient
from pandas.io.json import json_normalize
from pathlib import Path
import matplotlib.pyplot as plt
import matplotlib.pylab as pylab
import numpy as np
# Set some figures plot parameters:
# +
plt.style.use('bmh')
params = {'legend.fontsize': 'x-large',
'figure.figsize': (15, 15),
'axes.labelsize': 'x-large',
'axes.titlesize':'x-large',
'xtick.labelsize':'x-large',
'ytick.labelsize':'x-large'}
pylab.rcParams.update(params)
# -
# Provide your credentials to connect to the threedi-api:
# +
api_host = "https://api.3di.live/v3.0"
username = input("Username: ")
password = <PASSWORD>("Password: ")
config = {
"API_HOST": api_host,
"API_USERNAME": username,
"API_PASSWORD": password
}
api_client = ThreediApiClient(config=config)
# -
# Check the connection with your provided credentials:
# +
auth_api = AuthApi(api_client)
try:
user = auth_api.auth_profile_list()
except ApiException as e:
print("Oops, something went wrong. Maybe you made a typo?")
else:
print(f"Successfully logged in as {user.username}!")
# -
# 1.3 In order to run a simulation you need a threedi-model. Let's see which threedi-models are available:
# +
models_api = ThreedimodelsApi(api_client)
models = models_api.threedimodels_list(limit=10) # limit to the first 10 results
for model in models.results:
print(f"{model.name}")
# -
# In this notebook we will use one revision of the 3Di-model "BWN Schermer", and we can look up this model by using the following query:
models = models_api.threedimodels_list(name__icontains='bwn_schermer_1d2d_glg')
my_model =models.results[0]
my_model
# The 3Di-model of BWN Schermer:
#
# <img style="float: left;" alt="Drawing" src="./figures/3di_model_bwn_schermer.PNG" width="700"/>
# Now that we have a model we are almost ready to create the simulation. However, first we'll need to get an organisation under which's name we will run the simulation.
#
# Let's see which organisations are available within my user account:
# +
organisation_api = OrganisationsApi(api_client)
organisations = organisation_api.organisations_list()
for organisation in organisations.results:
print(f"{organisation.name}: {organisation.unique_id}")
# -
# In this example we use the organisation from N&S Demo:
#
# organisation_uuid = "a1993f6e13564e9687ae03a3604463f9"
organisation_uuid = "a1993f6e13564e9687ae03a3604463f9"
# 1.5 Let's create the simulation of the chosen model now, with this organisation uuid. Note that it will not run yet.
# +
simulation_api = SimulationsApi(api_client)
my_simulation = simulation_api.simulations_create(
data={
"name": "demo_simulation_3dinotebook_schermer",
"threedimodel": my_model.id,
"organisation": organisation_uuid,
"start_datetime": datetime.now(),
"duration": 3600 # in seconds, so we simulate for 1 hour
}
)
#print an overview of the simulation
my_simulation
# -
# You can check the status of the simulation with the following api call:
#check the status of the simulation with:
status = simulation_api.simulations_status_list(my_simulation.id)
print(status)
# We can see the simulation has not started yet. The options at the name of the status can be: "created", "started" and "finished".
# **Step 2: Adding events to this 3Di-simulation by using the threedi-api**
# In the previous step we created a simulation for the 3Di model of rockflow. Several events can be added to this 3Di-simulation:
#
# * initial waterlevels
# * rain
# * breaches
# * laterals
#
# In this step we create a simple constant rain event of 30 minutes with an offset of 5 minutes:
# +
from openapi_client.models import ConstantRain
constant_rain = ConstantRain(
simulation=my_simulation.id, # the ID we got from our create call above
offset=300, # let the rain start after 5 minutes
duration=1800, # let the rain last for half an hour
value=0.0006, # not too extreme after all...;-)
units="m/s" # the only unit supported for now
)
print (constant_rain)
# -
# Now we add this constant rain event to our created simulation
simulation_api.simulations_events_rain_constant_create(my_simulation.id, constant_rain)
# 2.3 We can get an overview of the added events to our 3Di-simulation by the following api-call:
events = simulation_api.simulations_events(my_simulation.id)
print(events)
# So, we can indeed see here that we have only added the constant rain event to our 3di-simulation.
# **Step 3: Running the 3Di-simulation by using the threedi-api**
# We will now start our simulation with the constant rain event:
simulation_api.simulations_actions_create(my_simulation.id, data={"name": "start"})
# We can check the status of the 3Di-simulation with:
#check the status of the simulation with:
status = simulation_api.simulations_status_list(my_simulation.id)
print(status)
# In the end we must see that our simulation has finished:
#check the status of the simulation with:
status = simulation_api.simulations_status_list(my_simulation.id)
print(status)
# **Step 4: Downloading the results of the 3Di-simulation**
# When our simulation has finished, it is time to analyse the results of the 3Di-model. Before that, we have to download all the results of the simulation.
# We select again the model and check if the status is finished:
# +
status = simulation_api.simulations_status_list(my_simulation.id)
print(my_simulation)
print(f"status: {status}")
assert status.name == 'finished'
# -
# If you began this script at step 4, after previously running a simulation, and you want to find and define this simulation. You can run the following code.
#
# Otherwise, you can skipt this part.
# +
my_simulation_name = 'demo_simulation_3dinotebook_schermer'
username = 'valerie.demetriades'
my_simulation = simulation_api.simulations_list(
name=my_simulation_name, user__username=username
).results[0]
print(my_simulation)
print(f"status: {status}")
assert status.name == 'finished'
# -
# Important note: results[0] returns the last simulation that matches the simulation name and username. Please check if this is the right simulation. If you mean to select an older simulation, change the number to the corresponding simulation. .results[1], .results[2], etc.
#
# Let's see which result files are available:
# +
result_files = simulation_api.simulations_results_files_list(my_simulation.id)
for result in result_files.results:
print(result)
# -
# And let's download all above-mentioned result files and put them in a local folder:
# +
download_folder = Path(f'Results {my_simulation.name}')
download_folder.mkdir(exist_ok=True)
for file in result_files.results:
download_url = simulation_api.simulations_results_files_download(
id=file.id, simulation_pk=my_simulation.id
)
file_path = download_folder / file.filename
r = requests.get(download_url.get_url)
with open(file_path, 'wb') as f:
for chunk in r.iter_content(chunk_size=8192):
f.write(chunk)
print(f"Finished downloading {file.filename}")
# -
# We have downloaded all the results of the 3Di-simulation, however, the ThreediToolbox and threedigrid also require the "gridadmin.h5" file of the simulation. This is a model specific file so it's under the threedimodels-api. We'll also download this file:
# +
model_api = ThreedimodelsApi(api_client)
threedi_model_id = my_simulation.threedimodel_id
download_url = model_api.threedimodels_gridadmin_download(threedi_model_id)
file_path = download_folder / "gridadmin.h5"
r = requests.get(download_url.get_url)
with open(file_path, "wb") as f:
for chunk in r.iter_content(chunk_size=8192):
f.write(chunk)
print(f"Finished downloading gridadmin.h5")
# -
# **Step 5: Analysing the results of the simulation**
# In this step we will visualize and analyse the results of the 3Di-simulation. We will do this by making use of the package ThreedimodelsApi and threedigrid.
#Import libraries
from threedigrid.admin.gridresultadmin import GridH5ResultAdmin
from threedigrid.admin.gridadmin import GridH5Admin
import os
# +
#Define paths and files
result_path = download_folder
nc = os.path.join(result_path,'results_3di.nc')
f = os.path.join(result_path,'gridadmin.h5')
#Load files into gr object
ga = GridH5Admin(f)
gr = GridH5ResultAdmin(f,nc)
# +
plt.figure()
#plotting only your 2d open water grid
xyc = ga.nodes.subset('2d_open_water').coordinates
plt.plot(xyc[0], xyc[1], '.', label='2D open water')
#plotting your 1D grid
xyc = ga.nodes.subset('1D_ALL').coordinates
plt.plot(xyc[0], xyc[1], '.', label='1D all', color='green', markersize=4)
# Setting the axis right
plt.legend()
plt.axis('equal')
plt.axis('tight')
plt.title("2D Open water and 1D grid - 3Di simulation BWN Schermer", fontsize=20)
# -
# Let's plot the waterlevels at some points of the grid:
# +
#see which nodes id's are available
print (ga.nodes.id)
# Select some nodes
sselect = [10000,20000, 30000, 40000]
sset = ga.nodes.filter(id__in=sselect)
# +
# Show in a plot where these nodes are in the model
plt.figure()
plt.plot(ga.nodes.coordinates[0], ga.nodes.coordinates[1], '.', markersize=2)
# and than your selection in red with extra large markers
plt.plot(sset.coordinates[0], sset.coordinates[1], '.', markersize=20, color='orange')
plt.text(sset.coordinates[0][0]+100, sset.coordinates[1][0]+200, '1', fontsize=15)
plt.text(sset.coordinates[0][1]+100, sset.coordinates[1][1]+200, '2', fontsize=15)
plt.text(sset.coordinates[0][2]+100, sset.coordinates[1][2]+200, '3', fontsize=15)
plt.text(sset.coordinates[0][3]+100, sset.coordinates[1][3]+200, '4', fontsize=15)
plt.axis('tight')
plt.axis('equal')
plt.xlim(110000, 122500)
plt.ylim(506000, 518000)
plt.title('Overview of grid points where we would like to extract the waterlevel timeseries of the simulation', fontweight='bold')
# -
# Defining the time axis, within a time domain
ts = gr.nodes.timestamps[np.where((gr.nodes.timestamps >=0) & (gr.nodes.timestamps <= 7200))]
#Selecting the water levels based on your choice for domain and period.
s1_select = gr.nodes.filter(id__in=sselect).timeseries(start_time=0, end_time=7200).s1
# +
plt.figure(1)
# If you want to change the size and shape of only this figure
plt.figure(figsize=(20,7))
# Plotting the timeseries of your selection
plt.plot(ts[3:], s1_select[3:])
# Defining the legend based on the nodes of your selection
plt.legend(['1', '2', '3', '4'], frameon=True, fontsize=15, title='Grid points')
# Naming your axis
plt.xlabel('Time [seconds]')
plt.ylabel('Waterlevel [mNAP]')
# Title of your Figure
plt.title('Waterlevels at the different grid nodes - BWN Schermer')
# -
# Let's zoom in on a part of the model:
nodes_bbox = ga.nodes.filter(coordinates__in_bbox=[116005, 512000, 116010, 512250])
# +
# Show in a plot where these nodes are in the model
plt.style.use('seaborn')
plt.figure(figsize=(20,7))
plt.plot(ga.nodes.coordinates[0], ga.nodes.coordinates[1], '.', markersize=5)
# and than your selection in red with extra large markers
plt.plot(nodes_bbox.coordinates[0], nodes_bbox.coordinates[1], '.', markersize=20)
plt.axis('tight')
plt.axis('equal')
plt.xlim(115980, 116000)
plt.ylim(512000, 512400)
plt.title('Zooming in on a part of the model - a certain cross section', fontweight='bold')
# -
# We have selected a certain cross section and now we can calcute the following parameters at the points within this cross-section. Note that there are a lot more parameters to calculate, but they wont be used in this analysis.
#
# * velocity
# * discharge
# * ......
#
#
# +
# Selecting the lines in this specific area
lines_bbox = ga.lines.filter(line_coords__in_bbox=[116005, 512000, 116010, 512250])
# Selecting only the highest indexes, to select only the velocities in the y-direction
yline_bbox = lines_bbox.filter(id__gt=4000)
# +
# By selecting the coordinates of the lines, they give you the start and end location. One can for example compute the line centre
ycoor_line = lines_bbox.filter(id__gt=4000).line_coords[1]
# -
# showing some results, for example variations of velocity over the width
line_select = yline_bbox.id
# checking number of times a result is saved
gr.lines.timestamps.shape
# There are 13 timestamps and the last timestamp is the one we are going to use for this analysis. Ofcourse, it is also possible to use other timestamps.
# Selecting the set of velocities and dischargers at final timestamp
uset = gr.lines.filter(id__in=line_select).timeseries(indexes=[12]).u1 #calculate velocity at last timestamp
qset = gr.lines.filter(id__in=line_select).timeseries(indexes=[12]).q #calculate discharge at last timestamp
# +
#Plotting the combined results of velocity, discharge and wet cross-sectional area
plt.figure(1, figsize=(20,15))
plt.subplot(311)
plt.plot(uset[0], ycoor_line, 'yv',markersize=15)
plt.xlabel('velocity [m/s]')
plt.ylabel('Y-coordinate')
plt.title('Various variables at cross-section')
plt.figure(2, figsize=(20,15))
plt.subplot(312)
plt.plot(qset[0], ycoor_line, 'b.',markersize=15)
plt.xlabel('discharge [m3/s]')
plt.ylabel('Y-coordinate')
# -
#
# **-----------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------**
# The end
#
# In this tutorial we showed you:
# - how to start a simulation of a 3Di-model,
# - how to add a simple rain event to your simulation,
# - how to download the results of the model simulation,
# - and in the end how to visualise your model and how to use threedigrid to do any analysis on the results.
#
# And all possible within a Jupyter notebook by making use of the API-v3. Ofcourse there are a lot more options for the analysis possible, by using threedigrid. Or for the events you can add to your simulation. Do you want to learn more or are you interested in some more possible analysis in a jupyter notebook?
#
# **Please contact**:
#
# <NAME> | <EMAIL>
#
# <NAME> | <EMAIL>
#
| Notebooks 3Di - API v3 - VD/notebook1 - from starting your simulation to analysing your results/notebook1-simulation to results.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # 8. Geography
#
# Datashader contains a `geo` module which contains helper functions which should be familiar to the geospatial community.
#
# Some of the functions available include
# * [Generate Terrain](#ds.geo---generate-terrain)
# * [Hillshade](#ds.geo---hillshade-function)
# * [Slope](#ds.geo---slope-function)
# * [Aspect](#ds.geo---aspect-function)
# * [Bump](#ds.geo---bump-function)
# * [NDVI](#ds.geo---ndvi-function)
# * [Mean](#ds.geo---mean-function)
#
from datashader.transfer_functions import shade, stack
import numpy as np
# ## Generate Terrain Data
#
# To demonstrate using these functions, let's generate some fake terrain...
# +
from datashader import Canvas
from datashader.geo import generate_terrain
from datashader.colors import Elevation
W = 1000
H = 750
canvas = Canvas(plot_width=W, plot_height=H, x_range=(-20e6, 20e6), y_range=(-20e6, 20e6))
terrain = generate_terrain(canvas)
# -
# The grayscale value above shows the elevation linearly in intensity (with the large black areas indicating low elevation), but it will look more like a landscape if we map the lowest values to colors representing water, and the highest to colors representing mountaintops:
shade(terrain, cmap=Elevation, how='linear')
# ## Hillshade
#
# [Hillshade](https://en.wikipedia.org/wiki/Terrain_cartography) is a technique used to visualize terrain as shaded relief, illuminating it with a hypothetical light source. The illumination value for each cell is determined by its orientation to the light source, which is based on slope and aspect.
# +
from datashader.geo import hillshade
illuminated = hillshade(terrain)
shade(illuminated, cmap=['gray', 'white'], alpha=255, how='linear')
# -
# You can combine hillshading with elevation colormapping to indicate terrain types:
stack(shade(illuminated, cmap=['gray', 'white'], alpha=255, how='linear'),
shade(terrain, cmap=Elevation, how='linear', alpha=128))
# ## Slope
# [Slope](https://en.wikipedia.org/wiki/Slope) is the inclination of a surface.
# In geography, *slope* is amount of change in elevation of a terrain regarding its surroundings.
#
# Datashader's slope function returns slope in degrees. Below we highlight areas at risk for avalanche by looking at [slopes around 38 degrees](http://wenatcheeoutdoors.org/2016/04/07/avalanche-abcs-for-snowshoers/).
# +
from datashader.geo import slope
avalanche_slope_risk = slope(terrain)
avalanche_slope_risk.data = np.where(np.logical_and(avalanche_slope_risk.data > 25,
avalanche_slope_risk.data < 50),
1, np.nan)
stack(
shade(terrain, cmap=['black', 'white'], how='linear'),
shade(illuminated, cmap=['black', 'white'], alpha=128, how='linear'),
shade(avalanche_slope_risk, cmap='red', alpha=100),
)
# -
# ## Aspect
#
# [Aspect](https://en.wikipedia.org/wiki/Aspect_(geography)) is the orientation of slope, measured clockwise in degrees from 0 to 360, where 0 is north-facing, 90 is east-facing, 180 is south-facing, and 270 is west-facing.
#
# Below, we look to find slopes which face close to north.
# +
from datashader.geo import aspect
north_faces = aspect(terrain)
north_faces.data = np.where(np.logical_or(north_faces.data > 350 ,
north_faces.data < 10), 1, np.nan)
stack(
shade(terrain, cmap=['black', 'white'], how='linear'),
shade(illuminated, cmap=['black', 'white'], alpha=128, how='linear'),
shade(north_faces, cmap=['aqua'], alpha=50),
)
# -
# ## NDVI
#
# The Normalized Difference Vegetation Index (NDVI) quantifies vegetation by measuring the difference between near-infrared (which vegetation strongly reflects) and red light (which vegetation absorbs).
#
# For example, when you have negative values, it’s highly likely that it’s water. On the other hand, if you have a NDVI value close to +1, there’s a high possibility that it’s dense green leaves.
# But when NDVI is close to zero, there isn’t green leaves and it could even be an urbanized area.
# The output of *NDVI* ranges from [-1,+1], where `-1` means more "Red" radiation while `+1` means more "NIR" radiation.
#
# Below, we simulate the red and near-infrared bands using `datashader.perlin` random noise with different seeds and frequencies. Green areas should be those > 0, where higher NDVI values would indicate vegetation.
# +
from datashader.geo import ndvi
from datashader.geo import perlin
near_infrared_band = perlin(W, H, freq=(4, 3), seed=1)
red_band = perlin(W, H, freq=(32, 32), seed=2)
vegetation_index = ndvi(near_infrared_band, red_band)
shade(vegetation_index, cmap=['purple','black','green'], how='linear')
# -
# ## Bump
# Bump mapping is a cartographic technique often used to create the appearance of trees or other land features.
#
# The `datashader.bump` will produce a bump aggregate that can then used to add detail to the terrain. In this case, I will pretend the bumps are trees and shade them with green.
# +
from datashader.geo import bump
from functools import partial
def tree_heights(locations, min_val, max_val, height):
out = np.zeros(len(locations))
for i, (x, y) in enumerate(locations):
val = terrain.data[y, x]
if val > min_val and val < max_val:
out[i] = height
else:
out[i] = 0
return out
TREE_COUNT = 200000
trees = bump(W, H, count=TREE_COUNT // 3,
height_func=partial(tree_heights, min_val=50, max_val=500, height=10))
trees += bump(W, H, count=TREE_COUNT,
height_func=partial(tree_heights, min_val=500, max_val=2000, height=20))
trees += bump(W, H, count=TREE_COUNT // 3,
height_func=partial(tree_heights, min_val=2000, max_val=3000, height=10))
tree_colorize = trees.copy()
tree_colorize.data[tree_colorize.data == 0] = np.nan
stack(shade(terrain + trees, cmap=['black', 'white'], how='linear'),
shade(hillshade(terrain + trees), cmap=['black', 'white'], alpha=128, how='linear'),
shade(tree_colorize, cmap='limegreen', how='linear'))
# -
# ## Mean
# The `datashader.mean` function will smooth a given aggregate by using a 3x3 mean convolution filter. Optional parameters include `passes`, which is used to run the mean filter multiple times, and also `excludes` which are values that will not be modified by the mean filter.
#
# Just for fun, let's add a coastal vignette to give out terrain scene a bit more character. Notice the water below now has a nice coastal gradient which adds some realism to our scene.
# +
from datashader.geo import mean
LAND_CONSTANT = 50.
water = terrain.copy()
water.data = np.where(water.data > 0, LAND_CONSTANT, 0)
water = mean(water, passes=10, excludes=[LAND_CONSTANT])
water.data[water.data == LAND_CONSTANT] = np.nan
stack(
shade(terrain, cmap=['black', 'white'], how='linear'),
shade(water, cmap=['aqua','white']),
shade(hillshade(terrain), cmap=['black', 'white'], alpha=128, how='linear'),
)
# -
# ## Conclusion
#
# We've now seen a bunch of datashader's `geo` helper functions.
#
# Let's make our final archipelago scene by stacking `terrain`, `water`, `hillshade`, and `tree_highlights` together into one output image:
stack(shade(terrain + trees, cmap=Elevation, how='linear'),
shade(water, cmap=['aqua','white']),
shade(hillshade(terrain + trees), cmap=['black', 'white'], alpha=128, how='linear'),
shade(tree_colorize, cmap='limegreen', how='linear'))
# ### References
# - <NAME>., and <NAME>., 1998. Principles of Geographical Information Systems (Oxford University Press, New York), pp 406
# - Making Maps with Noise Functions: https://www.redblobgames.com/maps/terrain-from-noise/
# - How Aspect Works: http://desktop.arcgis.com/en/arcmap/10.3/tools/spatial-analyst-toolbox/how-aspect-works.htm#ESRI_SECTION1_4198691F8852475A9F4BC71246579FAA
| datashader-work/datashader-examples/user_guide/8_Geography.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Homework 1
# ### <NAME>, 03/2021
# ## Question 1
import torch as pt
import random
# Defining more sophisticated `print` function (the one used in the laboratory).
def pretty_print(obj, title=None):
if title is not None:
print(title)
print(obj)
print("\n")
# Creating class for multi-layer perceptrons:
# +
class MultiLayerPerceptron(pt.nn.Module):
def __init__(self):
super().__init__()
# Create members to simulate layers
self._h_layer_1 = pt.nn.Linear(in_features = 5, out_features = 11, bias = False)
self._h_layer_2 = pt.nn.Linear(in_features = 11, out_features = 16, bias = False)
self._h_layer_3 = pt.nn.Linear(in_features = 16, out_features = 13, bias = False)
self._h_layer_4 = pt.nn.Linear(in_features = 13, out_features = 8, bias = False)
self._o_layer = pt.nn.Linear(in_features = 8, out_features = 4, bias = False)
def forward(self, X):
out = self._h_layer_1(X)
out = pt.nn.functional.relu(out)
out = self._h_layer_2(out)
out = pt.nn.functional.relu(out)
out = self._h_layer_3(out)
out = pt.nn.functional.relu(out)
out = self._h_layer_4(out)
out = pt.nn.functional.relu(out)
out = self._o_layer(out)
out = pt.nn.functional.softmax(out, dim=1)
return out
# -
# ## Question 2
# Creating instance of `MultiLayerPerceptron` class:
mlp = MultiLayerPerceptron()
# #### Print summary with standard method
pretty_print(mlp, "Multi-Layer Perceptron")
# #### Print summary with `torchsummary.summary`
import sys
from torchsummary import summary
summary(mlp)
# ## Question 3
#
# ### *No bias* case
# Since the network has a total of six layers, input and output layers included, we need five matrices to store the weights.
#
# A first matrix $W^{(1)}$ is needed between the input and the first hidden layer. Since we have a "5-noded" input layer and an "11-noded" hidden layer, $W^{(1)}$ will be of the form
# $$
# W^{(1)}=
# \begin{pmatrix}
# w^{(1)}_{1,1} & \dotsm & w^{(1)}_{1,5}\\
# \dotsm & \ddots & \dotsm\\
# w^{(1)}_{11,1} & \dotsm & w^{(1)}_{11,5}
# \end{pmatrix}
# $$
# that is a $11\times5$ order matrix.
#
# A second matrix of weights is needed between the first and the second hidden layers: since the former belongs to $\mathbb{R}^{11}$ and the latter to $\mathbb{R}^{16}$, we will need a $16\times 11$ matrix this time:
# $$
# W^{(2)}=
# \begin{pmatrix}
# w^{(2)}_{1,1} & \dotsm & w^{(2)}_{1,11}\\
# \dotsm & \ddots & \dotsm\\
# w^{(2)}_{16,1} & \dotsm & w^{(2)}_{16,11}
# \end{pmatrix}
# $$
# that is a $11\times5$ order matrix.
#
# Since hidden layers two and three belongs respectively to $\mathbb{R}^{16}$ and $\mathbb{R}^{13}$, now we need a $13\times16$ order matrix:
# $$
# W^{(3)}=
# \begin{pmatrix}
# w^{(3)}_{1,1} & \dotsm & w^{(3)}_{1,16}\\
# \dotsm & \ddots & \dotsm\\
# w^{(3)}_{13,1} & \dotsm & w^{(3)}_{13,16}
# \end{pmatrix}.
# $$
#
# Similar reasoning brings us to deduce that, in order to deal with the rest of the layers, we need
# $$
# W^{(4)}=
# \begin{pmatrix}
# w^{(4)}_{1,1} & \dotsm & w^{(4)}_{1,13}\\
# \dotsm & \ddots & \dotsm\\
# w^{(4)}_{8,1} & \dotsm & w^{(4)}_{8,13}
# \end{pmatrix}
# $$
# $$
# W^{(5)}=
# \begin{pmatrix}
# w^{(5)}_{1,1} & \dotsm & w^{(5)}_{1,8}\\
# \dotsm & \ddots & \dotsm\\
# w^{(5)}_{4,1} & \dotsm & w^{(5)}_{4,8}
# \end{pmatrix},
# $$
# that are of order $8\times13$ and $4\times8$, respectively.
#
# This leads to the need of a number of parameters corresponding
# $$
# N_p=11\cdot5+16\cdot11+13\cdot16+8\cdot13+4\cdot8=575.
# $$
#
# ### *Bias* case
# In this case, to the number of parameters computed for the previous case, one must add the number of biases needed for each hidden layer layer.
#
# Since the first hidden layer belongs to $\mathbb{R}^{11}$, here we need a bias vector
# $$
# b^{(1)}=
# \begin{pmatrix}
# b^{(1)}_{1} \\
# \vdots \\
# b^{(1)}_{11}
# \end{pmatrix},
# $$
# which is a $11\times1$ vector. With a similar anrgument, the bias vectors for the rest of the hidden layers are
# $$
# b^{(2)}=
# \begin{pmatrix}
# b^{(2)}_{1} \\
# \vdots \\
# b^{(2)}_{16}
# \end{pmatrix},\,\,\,\,\,
# b^{(3)}=
# \begin{pmatrix}
# b^{(3)}_{1} \\
# \vdots \\
# b^{(3)}_{13}
# \end{pmatrix},\,\,\,\,\,
# b^{(4)}=
# \begin{pmatrix}
# b^{(4)}_{1} \\
# \vdots \\
# b^{(4)}_{8}
# \end{pmatrix},\,\,\,\,\,
# b^{(5)}=
# \begin{pmatrix}
# b^{(5)}_{1} \\
# \vdots \\
# b^{(5)}_{4}
# \end{pmatrix},
# $$
# that are $16\times1$, $13\times1$, $8\times1$ and $4\times 1$ vectors, respectively.
#
# The total number of parameters needed goes to
# $$
# N_p=575+11\cdot1+16\cdot1+13\cdot1+8\cdot1+4\cdot1=575+52=627.
# $$
# ## Question 4
for par_name, par in mlp.state_dict().items():
print(par_name, par)
print('\n')
norm_1 = par.norm(1).item()
pretty_print(norm_1, "1-norm of tensor {}".format(par_name))
norm_2 = par.norm(2).item()
pretty_print(norm_2, "2-norm of tensor {}".format(par_name))
print('-------------------------------------------------------')
print('\n')
| homework_1/homework_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
plt.rcParams.update({'font.size': 24,'lines.linewidth':3})
from astropy.io import fits # /!\ You'll have to install the package astropy to use this!
ddir='' #your data directory, like '/Users/ayep/research/data'; starting with "/Users" makes code easily portable to other computers :)
pdir='' #your plot directory, like '/Users/ayep/research/plot'
#My everyday functions:
#open data files line-by-line but split up by tab ('\t')
def opendat(dir,filename): #dir,'filename'. For opening a data file. Can then send through roundtable.
f=open(dir+filename,'r')
dat=f.readlines()
f.close()
labels=dat[0][0:-1].split()
dat2=[[a.strip('\n') for a in d.split('\t')] for d in dat if d[0]!='#']
dat3=[['nan' if a.strip()=='' else a for a in d] for d in dat2]
return [dat3,labels]
#open data files into well-named variables:
def opendat2(dirr,filename,params): #Use as var,var,var...=opendat2(dir,'filename',['keys']).
dat,label=opendat(dirr,filename) #Get keys by first leaving ['keys'] blank: opendat2(dirr,filename,[])
print(label)
varrs=[]
for i in range(len(params)):
j=label.index(params[i])
try:
var=np.array([float(d[j]) for d in dat]) #works for float.
varrs.append(var)
except ValueError:
var=[d[j].strip() for d in dat] #works for strings.
varrs.append(var)
return varrs
# -
# ## Spectra from .fits Files
# Fortunately Python offers packages (especially astropy) to deal with spectra .fits files. :) Some spectra have wavelength and flux, some only have flux, some split them into orders.... Every telescope spits out data a bit differently, but the first step is just to open it. I happen to have CHIRON data, which has wavelength and flux and several orders (that is, separate wavelength-range regions).
# +
#a few functions
#wavelength, flux, because I'm so lazy that I have a funciton for this.... Set up for CHIRON's format
def wf(dat,o): #whichord,dat. Setup: w,f=wf(#,dat)
w=[d[0] for d in dat[o]]
f=[d[1] for d in dat[o]]
return w,f
#normalize
#Blaze file:
bdat=opendat2(ddir,'blaze.dat',['0', '1', '2', '3', '4', '5', '6', '7', '8', '9', '10', '11', '12', '13', '14', '15', '16', '17', '18', '19', '20', '21', '22', '23', '24', '25', '26', '27', '28', '29', '30', '31', '32', '33', '34', '35', '36', '37', '38', '39', '40', '41', '42', '43', '44', '45', '46', '47', '48', '49', '50', '51', '52', '53', '54', '55', '56', '57', '58', '59', '60', '61'])
#Orders of interest:
os=[0,1,2,3,5,6,7,10,11,12,13,14,17,18,19,20,21,22,23,24,25,28,31,32,33,34,36,37,40,52] #of interest; dodge pressure-broadened (wide wings) and telluric: skip any with sharp lines in B star! ex: 26,29,35,38,39(butHa),43-48,51(best!),53-7 have tel. lines.
def useblaze(fraw,head,o): #flux of object, order of interest
fb=bdat[o]
#factor=np.max(fb)/(np.max(f[350:375])-np.std(f[350:375]))
#fn=np.array(f)/(np.array(fb)*(np.max(f[350:375])-np.std(f[350:375]))/np.max(fb))
gain=float(head['GAIN'])
RN=float(head['RON'])
K=2.5
mean=np.mean(fraw[325:425])
std=np.std(fraw[325:425])
#kill cosmic rays, particularly in peak region
f=[fraw[0],]+[fraw[i] if fraw[i]<mean+2.5*std else np.mean((fraw[i-1],fraw[i+1])) for i in range(1,len(fraw)-1)]+[fraw[-1],]
SNR=np.array(f)*gain/np.sqrt(np.array(f)*gain+K*RN**2.)
SNRb=np.array(fb)*gain/np.sqrt(np.array(fb)*gain+K*RN**2.)
#SNR=signal/noise --> noise=signal/SNR
#f=signal+noise=signal+signal/SNR=signal(1+1/SNR)
#signal=f/(1+1/SNR)
#scale max signal of blaze to max signal of target.
signalmax=sorted(f[325:425])[-6]/(1.+1./np.mean(SNR[325:425])) #cut the highest
signalbmax=sorted(fb[325:425])[-6]/(1.+1./np.mean(SNRb[325:425])) #cut the highest
scaleblaze=signalmax/signalbmax
#fn=np.array(f)/(np.array(fb)*np.max(f[350:375])/np.max(fb)) #low-noise standards work better like this.
fn=np.array(fraw)/(np.array(fb)*scaleblaze)
return fn
#convenient plotting functions
def plott(w,fn,o):
plt.figure(figsize=(20,10))
plt.plot(w,fn,lw=2)
plt.plot((w[0],w[-1]),(1,1),lw=1,ls='--',color='gray')
plt.ylim(0,4)
plt.title('Order '+str(o))
# -
#open .fits files
hdulist=fits.open('SampleSpectrum_G3V.fits') # opens the fits file
#hdulist.info() #can toggle to view some file info
dat=hdulist[0].data #puts the spectrum data itself into a variable called dat
head=hdulist[0].header #puts header information (RA, Dec, time of observation, etc.) into head
hdulist.close() #closes .fits file now that you've taken the data you want
# +
#View an order:
plt.plot(dat[0])
# +
#I've made a function, so view it this way:
w,f=wf(dat,1)
#normalize:
fn=useblaze(f,head,1)
#plot:
plott(w,fn,1)
# -
| 2020_Workshop/Alex_Python/SpecialData_Spectra_and_Dictionaries.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # YUSAG Linear Regression Model
# - <NAME>, Yale Undergraduate Sports Analytics Group
# This notebook is one of several notebooks exploring ratings systems often used in sports. All of the notebooks can be found in [this repo](https://github.com/mc-robinson/Ratings_Methods).
#
# Specifically, this notebook attempts to both explain and implement our own YUSAG ratings model. Much of this model is derived from the work of Professor [<NAME>](http://www.stat.yale.edu/~jay/), and we thank him very much for his guidance.
# ## The Basic Overview ##
#
# The material in this notebook follows nicely from the [notebook on Massey's method](https://github.com/mc-robinson/Ratings_Methods/blob/master/massey_ratings.ipynb), in which we explain the basics of least squares and linear regression. But in case you are starting with this notebook, I'll repeat the very, very brief review of linear regression. Please see the references if you want more detail.
#
# (Note: for a discussion of where the term "regression" comes from, see my notebook [here](https://github.com/mc-robinson/random_tutorials/blob/master/why_we_call_it_regression.ipynb))
#
# ### The Form of Multiple Linear Regression ###
#
# In linear regression, we simply attempt to predict a scalar variable $\hat y$ as the weighted sum of a bunch of input features (explanatory variables) and a bias term (the interecept). The general form is:
#
# $$
# \hat y = w_0 + w_1 x_1 + w_2 x_2 + \cdots + w_n x_n
# $$
# where:
# * $\hat y$ is the predicted response variable
# * $x_i$ is the i'th feature value
# * $w_j$ is the j'th model parameter
# * $w_0$ is the bias term
# * $w_1, w_2,...,w_n$ are the feature weights
# * $n$ is the number of features
#
# So how do we figure out the values of the model coefficients $w_0,...,w_n$? The answer is that we learn these parameters when we train the linear model on the data. In fact, for linear regression, the fit is determined using the least squares criterion. That is, we seek the parameters that minimize the sum of squared errors. Once the model is trained, and the best parameters are learned, we can use the model for prediction.
# ## Working Through an Example ##
#
# In order to work through the implementation of this method and understand how it works, I'm going to use an extremely simplified example. Let's imagine the Ivy league is instead the IV league and consists of only four teams who each play each other once: Harvard, Princeton, Yale, and Columbia.
#
# Here are the results from the 2016 IV season that I scraped from the NCAA stats website:
import numpy as np
import pandas as pd
IV_df = pd.read_csv('IV_league_2016_YUSAG_lin_reg_data.csv')
IV_df
# Now you probably noticed that every game is repeated in the above csv file. This is the way we get the data when we scrape the NCAA stats websites. The logical thing to do would be to only keep one copy of each game. But we don't! We've actually found it's quite fine to keep two copies of each game with the `team` and `opponent` variables switched (more on this later)
#
# Note that the `location` variable refers to the location of the `team` variable:
# * Home = 1
# * Neutral = 0
# * Away = -1
# ### The Model ###
#
# The goal of our linear regression model is quite simple; we are trying to explain the score differential of each game based on the strength of the `team`, the strength of the `opponent`, and the `location`.
#
# Thus we need to create a `score_diff` response variable for each game:
IV_df['score_diff'] = IV_df['team_score']-IV_df['opponent_score']
IV_df.head()
# You may notice that the `team` and `opponent` features are categorical, and thus are not currently ripe for use with linear regression. However, we can use what is called 'one hot encoding' in order to transform these features into a usable form. One hot encoding works by taking the `team` feature, for example, and transforming it into many features such as `team_Yale` and `team_Harvard`. This `team_Yale` feature will usally equal zero, except when the `team` is actually Yale, then `team_Yale` will equal 1. In this way, it's a binary encoding (which is actually very useful for us as we'll see later).
# One can use sklearn.preprocessing.OneHotEncoder for this task, but I am going to use Pandas instead:
# +
# create dummy variables, need to do this in python b/c does not handle automatically like R
team_dummies = pd.get_dummies(IV_df.team, prefix='team')
opponent_dummies = pd.get_dummies(IV_df.opponent, prefix='opponent')
IV_df = pd.concat([IV_df, team_dummies, opponent_dummies], axis=1)
# -
IV_df.head()
# Now let's make our training data, so that we can construct the model. For now, I am going to ignore the `location` feature.
# make the training data
X = IV_df.drop(['year','month','day','team','opponent','team_score','opponent_score','score_diff','location'], axis=1)
y = IV_df['score_diff']
X.head()
y.head()
# Now let's train the linear regression model. I am going to force the bias term (intercept) of the model to be 0, just to make the interpretation of the model slightly easier.
from sklearn.linear_model import LinearRegression
lin_reg = LinearRegression(fit_intercept=False)
lin_reg.fit(X, y)
# print the coefficients
print(lin_reg.intercept_)
print(lin_reg.coef_)
# get the R^2 value on the training data
r_squared = lin_reg.score(X, y)
print('R^2 on the training data:')
print(r_squared)
# Now that the model is trained, let's look at the model coefficients for each team.
# get the coefficients for each feature
coef_data = list(zip(X.columns,lin_reg.coef_))
coef_df = pd.DataFrame(coef_data,columns=['feature','feature_coef'])
coef_df
# Now let's get our ratings for each team. Note that in this model, a team's rating is simply defined as its linear regression coefficient for the `team_name` variable, which we call the ***YUSAG coefficient***. Let's eliminate the `opponent_name` variables so we have the true ratings.
# +
# first get rid of opponent_ variables
team_df = coef_df[~coef_df['feature'].str.contains("opponent")]
# rank them by coef, not alphabetical order
ranked_team_df = team_df.sort_values(['feature_coef'],ascending=False)
# reset the indices at 0
ranked_team_df = ranked_team_df.reset_index(drop=True);
# rename 'feature_coef' column
ranked_team_df = ranked_team_df.rename(columns={'feature_coef': 'YUSAG_coefficient'})
# -
ranked_team_df.head()
# Note: this is exactly the answer we got from Massey's method, as explained in [this notebook](https://github.com/mc-robinson/Ratings_Methods/blob/master/massey_ratings.ipynb). If you read the end of that notebook, the connection should be relatively clear.
#
# Now let's re-train the model, while also including the `location` feature:
# make the training data (now with location)
X = IV_df.drop(['year','month','day','team','opponent','team_score','opponent_score','score_diff'], axis=1)
y = IV_df['score_diff']
from sklearn.linear_model import LinearRegression
lin_reg = LinearRegression(fit_intercept=False)
lin_reg.fit(X, y)
# print the coefficients
print(lin_reg.intercept_)
print(lin_reg.coef_)
# get the R^2 value on the training data
r_squared = lin_reg.score(X, y)
print('R^2 on the training data:')
print(r_squared)
# get the coefficients for each feature
coef_data = list(zip(X.columns,lin_reg.coef_))
coef_df = pd.DataFrame(coef_data,columns=['feature','feature_coef'])
coef_df
# +
# first get rid of opponent_ variables
team_df = coef_df[~coef_df['feature'].str.contains("opponent")]
# get rid of the location variable
team_df = team_df.iloc[1:]
# rank them by coef, not alphabetical order
ranked_team_df = team_df.sort_values(['feature_coef'],ascending=False)
# reset the indices at 0
ranked_team_df = ranked_team_df.reset_index(drop=True);
# rename 'feature_coef' column
ranked_team_df = ranked_team_df.rename(columns={'feature_coef': 'YUSAG_coefficient'})
# -
ranked_team_df
# Now let's go through the details of the model.
# ### How The Model Actually Works ###
#
# You may notice that the coefficients for `team_Yale` and `opponent_Yale` are just the negatives of each other. This is true for every team and is precisely why we include two copies of every game in our training data. The coefficient for the `team_Yale` feature is what we have called the YUSAG coefficient.
#
# When predicting a game's score differential on a **neutral field**, the predicted score differential (`score_diff`) is just the difference in YUSAG coefficients. The reason this works is the binary encoding we did earlier.
#
# So let's think about what we are doing when we predict the score differential for the Princeton-Harvard game with `team` = Princeton and `opponent` = Harvard on a neutral field.
#
# In our model, the coefficients are as follows:
# * team_Princeton_coef = 12.475
# * opponent_Princeton_coef = -12.475
# * team_Harvard_coef = 3.275
# * opponent_Harvard_coef = -3.275
#
# when we go to use the model for this game, it looks like this:
#
# `score_diff` = (location_coef $*$ `location`) + (team_Princeton_coef $*$ `team_Princeton`) + (opponent_Princeton_coef $*$ `opponent_Princeton`) + (team_Harvard_coef $*$ `team_Harvard`) + (opponent_Harvard_coef $*$ `opponent_Harvard`) + (team_Yale_coef $*$ `team_Yale`) + (opponent_Yale_coef $*$ `opponent_Yale`) + (team_Columbia_coef $*$ `team_Columbia`) + (opponent_Columbia_coef $*$ `opponent_Columbia`)
#
#
# To put numbers in for the variables, the model looks like this:
#
#
# `score_diff` = (location_coef $*$ 0) + (team_Princeton_coef $*$ 1) + (opponent_Princeton_coef $*$ 0) + (team_Harvard_coef $*$ 0) + (opponent_Harvard_coef $*$ 1) + $\cdots \\$
#
# where are the other terms are simply $0$.
#
# Which can also be written as:
#
# `score_diff` = (location_coef $*$ 0) + (12.475 $*$ 1) + (-3.275 $*$ 1) = 12.475 - 3.275 = Princeton_YUSAG_coef - Harvard_YUSAG_coef
#
# Thus showing how the difference in YUSAG coefficients is the same as the predicted score differential. Furthermore, the higher YUSAG coefficient a team has, the better they are.
#
# Lastly, if the Princeton-Harvard game was to be home at Princeton, we would just add the location_coef:
#
# `score_diff` = (location_coef $*$ $1$) + (team_Princeton_coef $*$ $1$) + (opponent_Harvard_coef $*$ $1$) = $-10.1 + 12.475 - 3.275$ = location_coef + Princeton_YUSAG_coef - Harvard_YUSAG_coef
#
# Note: With this small sample size of games, we have somehow selected 6 games with the visiting team winning 5 of them. Therefore, our location coefficient has surprisingly come out to be strongly negative. Over the course of a whole season, it will surely become positive (usually around 2-3 points).
# When we actually run our YUSAG model, we use ridge regression (adds an l2 penalty with alpha = 1.0) because that prevents the model from overfitting and also limits the values of the coefficients to not be huge (this sometimes happens when running on whole season of data).
from sklearn.linear_model import Ridge
ridge_reg = Ridge(fit_intercept=False)
ridge_reg.fit(X, y)
# print the coefficients
print(ridge_reg.intercept_)
print(ridge_reg.coef_)
# get the R^2 value
r_squared = ridge_reg.score(X, y)
print('R^2 on the training data:')
print(r_squared)
# get the coefficients for each feature
coef_data = list(zip(X.columns,ridge_reg.coef_))
coef_df = pd.DataFrame(coef_data,columns=['feature','feature_coef'])
coef_df
# +
# first get rid of opponent_ variables
team_df = coef_df[~coef_df['feature'].str.contains("opponent")]
# get rid of the location variable
team_df = team_df.iloc[1:]
# rank them by coef, not alphabetical order
ranked_team_df = team_df.sort_values(['feature_coef'],ascending=False)
# reset the indices at 0
ranked_team_df = ranked_team_df.reset_index(drop=True);
# rename 'feature_coef' column
ranked_team_df = ranked_team_df.rename(columns={'feature_coef': 'YUSAG_coefficient'})
# -
ranked_team_df
| YUSAG_linear_regression_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:ds] *
# language: python
# name: conda-env-ds-py
# ---
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:80% !important; }</style>"))
# +
import sys
sys.path.append('../../arxiv_cs/')
# %load_ext autoreload
# %autoreload 2
# +
import json
import pandas as pd
import gzip
from pathlib import Path
from utils import raw2json, clean_df
# -
DATA_RAW_PATH = Path("../../data/raw")
textfile = DATA_RAW_PATH / "arxiv_cs.txt.gz"
raw2json(textfile, DATA_RAW_PATH, compressed=True)
# +
df = pd.read_json(DATA_RAW_PATH / "arxiv_cs.json.gz")
df.head()
# -
df.abstract[0]
| notebooks/experimental/04_raw2json.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Homework: Mass Email Marketing
#
# ## The Problem
#
# You have been contracted by former energy company **Enron** to create a database of email addresses for "mass marketing" *ahem* ,*cough*: SPAMMING. You will get the emails for this "mass marketing" campaign from the email inboxes of the sales team, provided here.
#
# - `enron-allen-inbox.txt`
# - `enron-donohoe-inbox.txt`
# - `enron-lay-inbox.txt`
# - `enron-williams-inbox.txt`
# - `enron-onemail-inbox.txt` ( a sample with just one email in it - helpful for testing)
#
# ### Run this cell to download the files.
#
# ! curl https://raw.githubusercontent.com/mafudge/datasets/master/ist256/07-Files/enron-allen-inbox.txt -o enron-allen-inbox.txt
# ! curl https://raw.githubusercontent.com/mafudge/datasets/master/ist256/07-Files/enron-donohoe-inbox.txt -o enron-donohoe-inbox.txt
# ! curl https://raw.githubusercontent.com/mafudge/datasets/master/ist256/07-Files/enron-lay-inbox.txt -o enron-lay-inbox.txt
# ! curl https://raw.githubusercontent.com/mafudge/datasets/master/ist256/07-Files/enron-williams-inbox.txt -o enron-williams-inbox.txt
# ! curl https://raw.githubusercontent.com/mafudge/datasets/master/ist256/07-Files/enron-onemail-inbox.txt -o enron-onemail-inbox.txt
# Your task is to provide a drop-down so the user of the program can select one of the 5 mailboxes. Upon running the interaction the program will:
#
# - read the selected mailbox file a line at a time
# - find any lines beginning with `From:`.
# - extract out the email address from the `From:` line.
# - use the `isEmail()` function (provided below) to ensure its a valid email address.
# - print the email address
# - write the email to the emails file. (for example `enron-allen-inbox.txt` would write the emails to `enron-allen-emails.txt`.
# - NOTE: any emails from the `enron.com` domain are internal and should be omitted from the list. We don't need to "mass market" to ourselves!
#
#
# HINTS:
#
# - We saw how to extract emails in the Lab. This approach should work here.
# - The **problem simplification** approach is a good approach to this problem. Start with a simpler problem and add one more piece of complexity with each iteration:
# - First solve a simpler problem which you extract the single email from the `enron-onemail-inbox.txt` file.
# - Then re-write your program to use another mailbox like `enron-allen-inbox.txt` to make sure it prints multiple emails.
# - Next re-write your program to omit any emails with the domain `enron.com`
# - Next not only print the emails, but write them back out to the file: `enron-allen-emails.txt`
# - Finally add the ipywidget drop-down where you can select a mailbox and save to a different `emails` file based on the `inbox` file name.
#
# + [markdown] label="problem_analysis_cell"
# ## Part 1: Problem Analysis
#
# Inputs:
#
# ```
# TODO: Inputs
# ```
#
# Outputs:
#
# ```
# TODO: Outputs
# ```
#
# Algorithm (Steps in Program):
#
# ```
# TODO:Steps Here
#
# ```
# -
# ## Part 2: Code Solution
#
# You may write your code in several cells, but place the complete, final working copy of your code solution within this single cell below. Only the within this cell will be considered your solution. Any imports or user-defined functions should be copied into this cell.
# + label="code_solution_cell"
# Step 2: Write code here
import re
def isemail(text):
# + [markdown] label="homework_questions_cell"
# ## Part 3: Questions
#
# 1. Did you write your own user-defined function? For what purpose?
#
# `--== Double-Click and Write Your Answer Below This Line ==--`
#
#
# 2. Explain how you might re-write this program to create one large file from all the mailboxes. No code, just explain it.
#
# `--== Double-Click and Write Your Answer Below This Line ==--`
#
#
# 3. Devise an approach to remove duplicate emails from the output file. You don't have to write as code, just explain it.
#
# `--== Double-Click and Write Your Answer Below This Line ==--`
#
#
#
# + [markdown] label="reflection_cell"
# ## Part 4: Reflection
#
# Reflect upon your experience completing this assignment. This should be a personal narrative, in your own voice, and cite specifics relevant to the activity as to help the grader understand how you arrived at the code you submitted. Things to consider touching upon: Elaborate on the process itself. Did your original problem analysis work as designed? How many iterations did you go through before you arrived at the solution? Where did you struggle along the way and how did you overcome it? What did you learn from completing the assignment? What do you need to work on to get better? What was most valuable and least valuable about this exercise? Do you have any suggestions for improvements?
#
# To make a good reflection, you should journal your thoughts, questions and comments while you complete the exercise.
#
# Keep your response to between 100 and 250 words.
#
# `--== Double-Click and Write Your Reflection Below Here ==--`
#
# -
# run this code to turn in your work!
from coursetools.submission import Submission
Submission().submit()
| lessons/07-Files/HW-Files.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# ### Plot structure and dispersion curve of modes 0, 1, 2 AST
# +
import cmocean as cmo
import matplotlib.pyplot as plt
import numpy as np
import scipy as sc
import scipy.io as sio
import pandas as pd
# %matplotlib inline
# +
from IPython.display import HTML
HTML('''<script>
code_show=true;
function code_toggle() {
if (code_show){
$('div.input').hide();
} else {
$('div.input').show();
}
code_show = !code_show
}
$( document ).ready(code_toggle);
</script>
<form action="javascript:code_toggle()"><input type="submit" value="Click here to toggle on/off the raw code."></form>''')
# +
def get_Brink(file_fig, file_ratio):
# Brink mode
file = sio.loadmat(file_fig)
z, xpl, xxx, zzz = file['z'][0,:], file['xpl'][0,:], file['xxx'][0,:], file['zzz'][0,:]
k, f = file['wavenumber'][0][0], file['frequency'][0][0]
# (u is cross-shore and v is alongshore in Brink. I'm flipping them back right here)
p0, v0, u0, w0, r0 = file['p_profile'], file['u_profile'],file['v_profile'], file['w_profile'], file['r_profile']
file_rfw = sio.loadmat(file_ratio)
R = file_rfw['ratio_for_wave'][0][0]
print('ratio for wave', R)
print('k (rad/cm) ', k)
print('lambda (km) ', (2*np.pi/(k*100E3)))
print('omega (rad/s) ', f)
print('c (m/s)', (f/(k*100)))
scale=0.2
w = w0 * 0.01 * scale # cms-1 to ms-1 and normalization (?)
u = u0 * 0.01 * scale # cms-1 to ms-1 and normalization
v = v0 * 0.01 * scale # cms-1 to ms-1 and normalization
r = r0 * 1.0 * scale # mg/cm³ to kg/m³ and normalization
p = p0 * 0.1 * scale # dyn/cm² to 0.1 Pa (or kg m-1 s-2) and normalization
return(u,v,w,r,p,z,k,f,xpl, xxx, zzz)
def plot_Brink(ax2,ax3,ax4,u,v,p,z,xpl,xxx,zzz,minp,maxp,nlev=15):
landc='#8b7765'
levels=np.linspace(minp,maxp,nlev)
p3 = ax4.contourf(xpl, z, p, levels=levels, cmap=cmo.cm.delta, vmin=minp,
vmax=maxp, zorder=1)
ax4.contour(xpl, z, p, levels=[0], linewidths=2, linestyles='-', colors='k', zorder=2)
ax4.contour(xpl, z, p, levels=levels, linewidths=1, linestyles='-', colors='k', zorder=2)
ax4.fill_between(xxx, zzz.min(), zzz, facecolor=landc, zorder=3)
ax4.set_title('p')
levels=np.linspace(np.nanmin(v),np.nanmax(v),nlev)
p4 = ax2.contourf(xpl, z, v, levels=levels, cmap=cmo.cm.balance, vmin=-np.nanmax(v),
vmax=np.nanmax(v), zorder=1)
ax2.contour(xpl, z, v, levels=[0], linewidths=2, linestyles='-', colors='k', zorder=2)
ax2.contour(xpl, z, v, levels=levels, linewidths=1, linestyles='-', colors='k', zorder=2)
ax2.fill_between(xxx, zzz.min(), zzz, facecolor=landc, zorder=3)
ax2.set_title('v, cross-shelf')
levels=np.linspace(np.nanmin(u),np.nanmax(u),nlev)
p4 = ax3.contourf(xpl, z, u, levels=levels, cmap=cmo.cm.balance, vmin=np.nanmin(u),
vmax=-np.nanmin(u), zorder=1)
ax3.contour(xpl, z, u, levels=[0], linewidths=2, linestyles='-', colors='k', zorder=2)
ax3.contour(xpl, z, u, levels=levels, linewidths=1, linestyles='-', colors='k', zorder=2)
ax3.fill_between(xxx, zzz.min(), zzz, facecolor=landc, zorder=3)
ax3.set_title('u, along-shelf')
def plot_surface(ax0,ax1,v,p,xpl):
ax0.plot(xpl,p,'-', color='navy', label='surface pressure')
ax1.plot(xpl,v,'-', color='navy', label='surface cross-shore vel.')
ax0.axhline(0, color='0.5')
ax1.axhline(0, color='0.5')
ax0.set_ylabel('P')
ax1.set_ylabel('v')
ax0.legend()
ax1.legend()
ax1.set_xlabel('Cross-shelf distance / km')
# -
# ### Shelf profile
fig, (ax2,ax3,ax4) = plt.subplots(1,3,figsize=(12,4),sharey=True)
u,v,w,r,p,z,k,f,xpl,xxx,zzz = get_Brink('../AST/figures_mode0_AST.mat' ,
'../AST/ratio_for_wave_mode0_AST.mat')
minp = np.nanmin(p)
maxp = np.nanmax(p)
plot_Brink(ax2, ax3, ax4, u, v, p, z, xpl, xxx, zzz, minp, maxp)
ax2.set_ylabel('Depth / m')
ax2.set_xlabel('Cross-shelf distance / km')
ax3.set_xlabel('Cross-shelf distance / km')
ax4.set_xlabel('Cross-shelf distance / km')
fig, (ax0,ax1) = plt.subplots(2,1,figsize=(8,3),sharex=True)
plot_surface(ax0,ax1,v[-1,:],p[-1,:],xpl)
fig, (ax2,ax3,ax4) = plt.subplots(1,3,figsize=(12,4),sharey=True)
u,v,w,r,p,z,k,f,xpl,xxx,zzz = get_Brink('../AST/figures_mode1_AST.mat' ,
'../AST/ratio_for_wave_mode1_AST.mat')
minp = - np.nanmax(p)
maxp = np.nanmax(p)
nlevels = 70
plot_Brink(ax2, ax3, ax4, u, v, p, z, xpl, xxx, zzz, minp, maxp, nlev=nlevels)
ax2.set_ylabel('Depth / m')
ax2.set_xlabel('Cross-shelf distance / km')
ax3.set_xlabel('Cross-shelf distance / km')
ax4.set_xlabel('Cross-shelf distance / km')
fig, (ax0,ax1) = plt.subplots(2,1,figsize=(8,3),sharex=True)
plot_surface(ax0,ax1,v[-1,:],p[-1,:],xpl)
fig, (ax2,ax3,ax4) = plt.subplots(1,3,figsize=(12,4),sharey=True)
u,v,w,r,p,z,k,f,xpl,xxx,zzz = get_Brink('../AST/figures_mode2_AST.mat' ,
'../AST/ratio_for_wave_mode2_AST.mat')
minp = np.nanmin(p)
maxp = -np.nanmin(p)
plot_Brink(ax2, ax3, ax4, u, v, p, z, xpl, xxx, zzz, minp, maxp)
ax2.set_ylabel('Depth / m')
ax2.set_xlabel('Cross-shelf distance / km')
ax3.set_xlabel('Cross-shelf distance / km')
ax4.set_xlabel('Cross-shelf distance / km')
fig, (ax0,ax1) = plt.subplots(2,1,figsize=(8,3),sharex=True)
plot_surface(ax0,ax1,v[-1,:],p[-1,:],xpl)
# ### Axis profile
fig, (ax2,ax3,ax4) = plt.subplots(1,3,figsize=(12,4),sharey=True)
u,v,w,r,p,z,k,f,xpl,xxx,zzz = get_Brink('../AST/figures_mode0_ASTAX.mat' ,
'../AST/ratio_for_wave_mode0_ASTAX.mat')
minp = np.nanmin(p)
maxp = np.nanmax(p)
plot_Brink(ax2, ax3, ax4, u, v, p, z, xpl, xxx, zzz, minp, maxp)
ax2.set_ylabel('Depth / m')
ax2.set_xlabel('Cross-shelf distance / km')
ax3.set_xlabel('Cross-shelf distance / km')
ax4.set_xlabel('Cross-shelf distance / km')
fig, (ax0,ax1) = plt.subplots(2,1,figsize=(8,3),sharex=True)
plot_surface(ax0,ax1,v[-1,:],p[-1,:],xpl)
fig, (ax2,ax3,ax4) = plt.subplots(1,3,figsize=(12,4),sharey=True)
u,v,w,r,p,z,k,f,xpl,xxx,zzz = get_Brink('../AST/figures_mode1_ASTAX.mat' ,
'../AST/ratio_for_wave_mode1_ASTAX.mat')
minp = - np.nanmax(p)
maxp = np.nanmax(p)
nlevels = 70
plot_Brink(ax2, ax3, ax4, u, v, p, z, xpl, xxx, zzz, minp, maxp, nlev=nlevels)
ax2.set_ylabel('Depth / m')
ax2.set_xlabel('Cross-shelf distance / km')
ax3.set_xlabel('Cross-shelf distance / km')
ax4.set_xlabel('Cross-shelf distance / km')
fig, (ax0,ax1) = plt.subplots(2,1,figsize=(8,3),sharex=True)
plot_surface(ax0,ax1,v[-1,:],p[-1,:],xpl)
fig, (ax2,ax3,ax4) = plt.subplots(1,3,figsize=(12,4),sharey=True)
u,v,w,r,p,z,k,f,xpl,xxx,zzz = get_Brink('../AST/figures_mode2_ASTAX.mat' ,
'../AST/ratio_for_wave_mode2_ASTAX.mat')
minp = np.nanmin(p)
maxp = -np.nanmin(p)
plot_Brink(ax2, ax3, ax4, u, v, p, z, xpl, xxx, zzz, minp, maxp)
ax2.set_ylabel('Depth / m')
ax2.set_xlabel('Cross-shelf distance / km')
ax3.set_xlabel('Cross-shelf distance / km')
ax4.set_xlabel('Cross-shelf distance / km')
fig, (ax0,ax1) = plt.subplots(2,1,figsize=(8,3),sharex=True)
plot_surface(ax0,ax1,v[-1,:],p[-1,:],xpl)
# +
g = 9.81 # gravitational accel. m/s^2
Hs = 150 # m shelf break depth
f = 1.0E-4 # inertial frequency
files = ['../AST/dispc_mode0_AST.dat',
'../AST/dispc_mode1_AST.dat',
'../AST/dispc_mode2_AST.dat',
'../BAR/dispc_mode0_BAR.dat',
'../BAR/dispc_mode1_BAR.dat',
'../BAR/dispc_mode2_BAR.dat',
'../AST/dispc_ASTAX_cnywave.dat',
'../ARGO/dispc_ARGOAX_cnywave.dat',
'../BAR/dispc_BAR_canyon_wave.dat',
'../PATH/dispc_PATHAX_cnywave.dat',
]
colors = ['navy',
'blue',
'lightskyblue',
'forestgreen',
'limegreen',
'lightgreen',
'red',
'orange',
'lightcoral',
'pink']
labels = ['$c_0$','$c_1$','$c_2$','Bar $c_0$','Bar $c_1$','Bar $c_2$','AST canyon','ARGO canyon','BAR canyon','PATH canyon']
fig1, ax0 = plt.subplots(1, 1, figsize=(10, 6.5))
for file, col, lab in zip(files, colors, labels):
data_mode = pd.read_csv(file, delim_whitespace=True, header=None, names=['wavenum', 'freq', 'perturbation'])
omega = data_mode['freq'][:-3]
k = data_mode['wavenum'][:-3]*100
ax0.plot(k*1E5, omega*1E4, '-',
color=col,linewidth=2,
label=lab+'=%1.2f m/s' % (np.mean(omega/k)))
ax0.plot(20,0.73066615,'o',color='red', label='AST-AX wave $\lambda=2W_m$')
ax0.plot(23.9,0.805,'o',color='lightcoral', label='BAR-AX wave $\lambda=2W_m$')
ax0.plot(20,0.58396825,'o',color='orange',label='ARGO-AX wave $\lambda=2W_m$')
ax0.plot(23.9,0.74,'o',color='pink',label='PATH-AX wave $\lambda=2W_m$')
k_vec = np.linspace(1E-10,1E-5,20)
ax0.plot(k_vec*1E5, (k_vec*(g*Hs)**0.5)*1E4, '-',color='k', label=r'$\omega=k(gH_s)^{1/2}$')
ax0.plot(k_vec*1E5, (k_vec*(g*200)**0.5)*1E4, '-',color='0.5', label=r'$\omega=k(200g)^{1/2}$')
ax0.axhline(f*1E4, linestyle='--', color='0.4', label='$f$')
ax0.axvline(2.24, linestyle=':', color='0.7', label='domain length')
ax0.axvline(20.0, linestyle=':', color='0.2', label='AST $2W_m$')
ax0.axvline(24.0, linestyle='-.', color='0.2', label='BAR $2W_m$')
ax0.set_xlabel(r'$k$ / $10^{-5}$ rad m$^{-1}$', labelpad=0.1)
ax0.set_ylabel(r'$\omega$ / $10^{-4}$ rad s$^{-1}$', labelpad=0.1)
ax0.set_ylim(0, f*1.2*1E4)
ax0.legend(bbox_to_anchor=(1,1))
plt.tight_layout()
plt.savefig('../figures/disp_curve_all.png', format='png', bbox_inches='tight')
# -
# Short wave of wavelength $\approx 2W$
#
# Astoria width at mouth = 15.7 km
#
# Astoria wave length 31.4 km, wave number $2\times10^{-4}$ m$^{-1}$
#
# Barkely width at mouth = 13.0 km
#
# Barkley wave length = 26 km wave numbers $2.4\times10^{-4}$ m$^{-1}$
#
# +
D = 280E3 # m, Domain length
T_model = 9*24*3600 # s, duration of simulation
min_c = D/T_model # min speed for recirculation
# short wave speeds (from cells below this one)
c_ast = 0.36533307500000006
c_bar = 0.3359149931109348
c_argo = 0.2919695265236738
c_path = 0.30920920629618803
# Mean Incoming flow
cf_ast = 0.300 #$\pm$ 0.002 ms$^-1$
cf_argo = 0.329 # $\pm$ 0.004 ms$^-1$
cf_bar = 0.300 #$\pm$ 0.001 ms$^-1$
cf_path = 0.288 #$\pm$ 0.004 ms$^-1$
# Time of recircualtion
Tast = D/c_ast
Tbar = D/c_bar
Targo = D/c_argo
Tpath= D/c_path
# Time of recircualtion against mean incoming flow
Tast_f = D/(c_ast-cf_ast)
Tbar_f = D/(c_bar-cf_bar)
Targo_f = D/(c_argo-cf_argo)
Tpath_f= D/(c_path-cf_path)
print('Velocity above which recirculation occurs: %1.2f m/s' %(min_c))
print('Astoria canyon wave takes %1.2f days to recirculate' %(Tast/(3600*24)))
print('Barkley canyon wave takes %1.2f days to recirculate' %(Tbar/(3600*24)))
print('ARGO canyon wave takes %1.2f days to recirculate' %(Targo/(3600*24)))
print('PATH canyon wave takes %1.2f days to recirculate' %(Tpath/(3600*24)))
print(' ')
print('Against the flow:')
print('Astoria canyon wave takes %1.2f days to recirculate' %(Tast_f/(3600*24)))
print('Barkley canyon wave takes %1.2f days to recirculate' %(Tbar_f/(3600*24)))
print('ARGO canyon wave takes %1.2f days to recirculate' %(Targo_f/(3600*24)))
print('PATH canyon wave takes %1.2f days to recirculate' %(Tpath_f/(3600*24)))
# -
print('speed against mean incoming flow')
print('AST %1.2f m/s' %(c_ast-cf_ast))
print('BAR %1.2f m/s' %(c_bar-cf_bar))
print('ARGO %1.2f m/s ' %(c_argo-cf_argo))
print('PATH %1.2f m/s ' %(c_path-cf_path))
# ### Astoria short wave (2Wm)
#
# - Found using canyon axis profile because there is where it is generated
# Astoria mid length wave
fig, (ax2,ax3,ax4) = plt.subplots(1,3,figsize=(12,4),sharey=True)
u,v,w,r,p,z,k,f,xpl,xxx,zzz = get_Brink('../AST/figures_ASTAX_canyon_wave.mat' ,
'../AST/ratio_for_wave_ASTAX_canyon_wave.mat')
minp = np.nanmin(p)
maxp = -np.nanmin(p)
plot_Brink(ax2, ax3, ax4, u, v, p, z, xpl, xxx, zzz, minp, maxp)
ax2.set_ylabel('Depth / m')
ax2.set_xlabel('Cross-shelf distance / km')
ax3.set_xlabel('Cross-shelf distance / km')
ax4.set_xlabel('Cross-shelf distance / km')
fig, (ax0,ax1) = plt.subplots(2,1,figsize=(8,3),sharex=True)
plot_surface(ax0,ax1,v[-1,:],p[-1,:],xpl)
# ### Barkley short wave (2Wm)
# Barkley mid length wave
fig, (ax2,ax3,ax4) = plt.subplots(1,3,figsize=(12,4),sharey=True)
u,v,w,r,p,z,k,f,xpl,xxx,zzz = get_Brink('../BAR/figures_BARAX_canyon_wave.mat' ,
'../BAR/ratio_for_wave_BARAX_canyon_wave.mat')
minp = np.nanmin(p)
maxp = -np.nanmin(p)
plot_Brink(ax2, ax3, ax4, u, v, p, z, xpl, xxx, zzz, minp, maxp)
ax2.set_ylabel('Depth / m')
ax2.set_xlabel('Cross-shelf distance / km')
ax3.set_xlabel('Cross-shelf distance / km')
ax4.set_xlabel('Cross-shelf distance / km')
fig, (ax0,ax1) = plt.subplots(2,1,figsize=(8,3),sharex=True)
plot_surface(ax0,ax1,v[-1,:],p[-1,:],xpl)
# ### ARGO short wave (2Wm)
# mid length wave
fig, (ax2,ax3,ax4) = plt.subplots(1,3,figsize=(12,4),sharey=True)
u,v,w,r,p,z,k,f,xpl,xxx,zzz = get_Brink('../ARGO/figures_ARGOAX_canyon_wave.mat' ,
'../ARGO/ratio_for_wave_ARGOAX_canyon_wave.mat')
minp = np.nanmin(p)
maxp = -np.nanmin(p)
plot_Brink(ax2, ax3, ax4, u, v, p, z, xpl, xxx, zzz, minp, maxp)
ax2.set_ylabel('Depth / m')
ax2.set_xlabel('Cross-shelf distance / km')
ax3.set_xlabel('Cross-shelf distance / km')
ax4.set_xlabel('Cross-shelf distance / km')
fig, (ax0,ax1) = plt.subplots(2,1,figsize=(8,3),sharex=True)
plot_surface(ax0,ax1,v[-1,:],p[-1,:],xpl)
# ### PATH short wave (2Wm)
# mid length wave
fig, (ax2,ax3,ax4) = plt.subplots(1,3,figsize=(12,4),sharey=True)
u,v,w,r,p,z,k,f,xpl,xxx,zzz = get_Brink('../PATH/figures_PATHAX_canyon_wave.mat' ,
'../PATH/ratio_for_wave_PATHAX_canyon_wave.mat')
minp = np.nanmin(p)
maxp = -np.nanmin(p)
plot_Brink(ax2, ax3, ax4, u, v, p, z, xpl, xxx, zzz, minp, maxp)
ax2.set_ylabel('Depth / m')
ax2.set_xlabel('Cross-shelf distance / km')
ax3.set_xlabel('Cross-shelf distance / km')
ax4.set_xlabel('Cross-shelf distance / km')
fig, (ax0,ax1) = plt.subplots(2,1,figsize=(8,3),sharex=True)
plot_surface(ax0,ax1,v[-1,:],p[-1,:],xpl)
| forPaper2/shelfWaves/CTW_Paper2/notebooks/AST_modes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # License Plate Detection with OpenCV
#
# In this project we demonstrate how to use OpenCV only, with traditional computer vision approaches, to perform License Plate Detection (LPD).
#
# We follow two approaches:
#
# 1- __Morphology based approach__: where only morphological transforms are used, along with some rules to detect the LP.
#
# 2- __Charater based approach__: in addition to basic morphological approaches, basic char detection, also based on morphology, is used as an extra characteristic of the LP.
#
# Further, the problem of Licence Plate Recognition (LPR), by recognizing the number and digits written, can be addressed by the second approach.
#
# In both approaches, we load HD videos (1080p). Due to the camera position, this is the most effective resolution to detect LP patterns.
#
# In both approaches we merge car detection, using background subtraction, to narrow the search space.
# # Pre-requisites
#
# You need to install the packages in `requirements.txt`:
#
# `pip install -r requirements.txt`
from utils import *
from MOD import detect_moving_objects
import numpy as np
import cv2
import imutils
import os
from tqdm import tqdm
import matplotlib.pyplot as plt
from video import process_video
# # Moving object detection (MOD)
#
# In this part, we show how to detect and isolate the car box.
#
# We use background subtraction. [See this reference](https://www.pyimagesearch.com/2015/05/25/basic-motion-detection-and-tracking-with-python-and-opencv/). This is possible due to the fixed camera position.
#
# We can detect bounding rectangle or oriented one. The oriented bbox is not very accurate, and later it turns to be not important for LPD.
# ## Video processing
# The `process_video` function takes car of frame processing of the given `video_file`. The output is saved in the location of the output `video_output_file`.
#
# This function can be used to:
# - Detect Moving cars.
# - Detect LPs within car frames, and plot it back in the original frame.
# - Detect LPs in the big frame directly.
from video import process_video
video_file = 'dat/detection_test.mp4'
video_output_file = 'dat/cars_detection.mp4'
process_video(video_file, video_output_file, show_cars_bbox=1)
# From command line
# !python main.py --video_file dat/detection_test.mp4 --video_output_file dat/cars_detection.mp4 --show_cars_bbox 1
# # Morphology based approach
#
# This approach is based on applying morphological operations to emphasasize the LP pattern. Mainly, two main patters:
# - Edge of bright area
# - Rectangular shape
#
# As usual with rule based approaches, we suffer sensitivity to parameters settings. To make it less critical we perform two simple tricks:
# - Apply the rules only on the car patches, thanks to the car detection step.
# - Resize into standard size makes it easier to set global rules, with less sensitivity to scale. This is also possible thanks to the car detection step.
#
# In this approach we follow the following steps:
#
# - Resize frame to standard size.
# - Transform frame into gray scale.
# - Adaptive thresholding.
# - Canny edge detection.
# - Dilation loop (3 iteration, 3x3 kernel).
# - Contours on dialted image.
# - Get candidate plates by fitting oriented bbox around contours.
# - Filter the candidate LPs with rules on L,W of the oriented bbox.
# - Resize the frame back into the original size
#
#
from morpho import detect_LP
img = cv2.imread("imgs/char_frame_180_car_no_lp1.png")
plot_img(img)
detected_img, LPs = detect_LP(img, debug=False)
plot_img(detected_img)
# ## Calibrating the rules
# In this section we use sample images captured from the test video in order to calibrate the min and max L and W of the plate detection.
#
# We set `debug=True` in order to see the intermediate results (thresholding, edges, dilation).
import cv2
from utils import plot_img
from morpho import detect_LP_morpho
img = cv2.imread("imgs/char_frame_180_car_no_lp1.png")
plot_img(img)
detected_img, LPs = detect_LP_morpho(cv2.resize(img, (500,500)), L_min=35, L_max=60, W_min=55, W_max=90, debug=True)
plot_img(detected_img)
video_file = 'dat/detection_test.mp4'
video_output_file = 'dat/morpho_LP_detection.mp4'
process_video(video_file, video_output_file, detect_LP_fn=detect_LP)
# From command line:
# !python main.py --video_file dat/detection_test.mp4 --video_output_file dat/cars_detection.mp4 --detect_LP_fn 2
# We notice the following problems:
# - Many false positives
# - Rules apply to disoriented false contours
# - Aggeessive dilation make bigger rectangles in some cases
#
#
# It is recommended:
# - Integrate a tracker to smooth the false positives (TBD).
# - Add more features of LP, like characters, which we will do next.
# # Character based approach
#
# The main approach in this part is imported from this nice git [repo](https://github.com/MicrocontrollersAndMore/OpenCV_3_License_Plate_Recognition_Python.git) Code is copied here just for self contained repo, with minor changes.
#
#
# The approach shares the same preprocessing steps as in the morphological approach above.
#
# However, we integrate extra features, which is char detection. This facilitates the filtering out operation, instead of only relying on the L, W rules.
#
# The downside is that, we now depend on the language of the sign. For different languages, we need different char detector, which is not a bi issue.
from char import detect_LP_char
img = cv2.imread("imgs/char_frame_180_car_no_lp1.png")
plot_img(img)
detected_img, LPs = detect_LP_char(cv2.resize(img, (700,700)), L_min=0, L_max=50, W_min=0, W_max=150, debug=True)
plot_img(detected_img)
from char import detect_LP
video_file = 'dat/detection_test.mp4'
video_output_file = 'dat/char_LP_detection.mp4'
process_video(video_file, video_output_file, detect_LP_fn=detect_LP)
# From command line:
# !python main.py --video_file dat/detection_test.mp4 --video_output_file dat/cars_detection.mp4 --detect_LP_fn 1
# The effect of adding the characters detection feature is clear in filtering out false positive.
# # Effect of MOD
# Now we will run the same approach, but on the whole frame, instead of detecting the car first
video_file = 'dat/detection_test.mp4'
video_output_file = 'dat/char_LP_detection_without_car_detection.mp4'
process_video(video_file, video_output_file, detect_LP_fn=detect_LP, cars_detection=False)
# !python main.py --video_file dat/detection_test.mp4 --video_output_file dat/cars_detection.mp4 --detect_LP_fn 1 --cars_detection False
# Again, lots of false positives detected. This shows the effect of detecting the moving cars as a preprocessing step.
#
# In the final video, you might now see any detected plates, since they are all filtered out by the internal rules.
# # Conclusion
#
# In this project we used native OpenCV, with traditional CV transformations, to detect the license plates. Morphological operations can do the job, however, it requires tuning, and is sensitive to calibration. Adding some features like characters matching improves the performance, however, it might require tuning on different languages.
# # References
# - https://www.pyimagesearch.com/2015/05/25/basic-motion-detection-and-tracking-with-python-and-opencv/
# - https://sod.pixlab.io/articles/license-plate-detection.html
# - https://github.com/MicrocontrollersAndMore/OpenCV_3_License_Plate_Recognition_Python.git
| LPD.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Pro-Machina/LSTM-Covid-Predictions/blob/main/Kshitij_EPI_LSTM_Final_Code_14_Day_Lead.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="LrBqvNsCIUP6"
# # Using LSTM to predict COVID-19 Cases in India
# ## Abstract
# The COVID-19 pandemic has resulted in a significant loss of economic output and human life. A means by which to accurately forecast the spread of the disease in relation to government policy is of critical importance when determining how to minimize both the spread of disease and economic impact. The objectives of this study is to investigate the dependence of COVID-19-related deaths on the mobility habits of individuals and the government response data. The project will investigate if there are differences in the effects of incorporating the mobility and government policy data for regions of varying population density. Using the Google's mobility dataset in conjunction with The WHO dataset for COVID-19 cases and deaths as well as government response data from Oxford University to train an LSTM model, the project aims to evaluate its performance using the root mean squared error between the predicted number of cases and the actual number of cases, and compare it to an ARIMA model.
# + colab={"base_uri": "https://localhost:8080/"} id="R6vPfP3eD6-A" outputId="24d6622a-0840-4cd2-9d12-85aa6f7e839b"
# !pip install --upgrade cython
# !pip install --upgrade git+https://github.com/statsmodels/statsmodels
# + id="N40GANKxv_fV" colab={"base_uri": "https://localhost:8080/"} outputId="b8d455b2-1907-4f8c-d881-fc28d862c057"
# Import packages
from sklearn.preprocessing import StandardScaler
import statsmodels.api as sm
from statsmodels.tsa.arima.model import ARIMA
from sklearn.metrics import mean_squared_error
import statsmodels.api as sm
import torch
from torch import nn
from torch.utils.data import Dataset
from torch.utils.data import DataLoader
from torch.utils.data import SequentialSampler
from torch import nn
import datetime
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from google.colab import drive
import statsmodels.api as sm
import scipy
import warnings
warnings.filterwarnings("ignore")
drive.mount('/content/drive')
# + [markdown] id="KH_TBSSTQCd8"
# ## Functions for Data Engineering
# + id="DQFlKSe0wSgG"
###
def filter_dates_df(start_date, end_date, df):
""" Extract the data corresponding to specific dates """
if 'date' in df:
filtered_data = df[(df['date'] >= start_date) & (df['date'] <= end_date)]
elif 'Date' in df:
filtered_data = df[(df['Date'] >= start_date) & (df['Date'] <= end_date)]
elif 'Date_reported' in df:
filtered_data = df[(df['Date_reported'] >= start_date) & (df['Date_reported'] <= end_date)]
return filtered_data
###
def filter_covid_data(covid_df):
""" Outputs the required dataset with required columns """
covid_df = covid_df.fillna(0)
covid_df = covid_df[covid_df['Country'] == 'India']
covid_df['Date_reported'] = pd.to_datetime(covid_df['Date_reported']).dt.date
covid_df = covid_df.drop_duplicates(subset = ['Date_reported'], keep = 'last')
covid_df = covid_df.drop(['Country_code', 'WHO_region'], axis=1)
covid_df = filter_dates_df(datetime.date(2020, 1, 15), datetime.date(2021, 10, 24), covid_df)
return covid_df
def filter_mobility_data(mobility_df):
""" Outputs the required dataset with required columns """
# mobility_df = mobility_df.fillna(0)
mobility_df = mobility_df[(mobility_df['place_id'] == 'ChIJkbeSa_BfYzARphNChaFPjNc')]
mobility_df = mobility_df.fillna(method='ffill')
mobility_df['date'] = pd.to_datetime(mobility_df['date']).dt.date
mobility_df = filter_dates_df(datetime.date(2020, 1, 15), datetime.date(2021, 10, 24), mobility_df)
return mobility_df
def filter_policy_data(policy_df):
""" Outputs the required dataset with required columns """
policy_df = policy_df.fillna(0)
policy_df = policy_df[(policy_df['CountryName'] == 'India')]
policy_df['Date'] = pd.to_datetime(policy_df['Date'], format='%Y%m%d').dt.date
policy_df = filter_dates_df(datetime.date(2020, 1, 15), datetime.date(2021, 10, 24), policy_df)
return policy_df
###
def standardize_df (df_input, target):
""" Returns standardized data """
df = df_input.copy()
target_mean = df[target].mean()
target_sd = df[target].std()
for c in df.columns:
mean = df[c].mean()
sd = df[c].std()
df[c] = (df[c] - mean)/sd
df = df.fillna(0)
return target_mean, target_sd, df
###
def scale_test_data(df_test_input, df_train_input):
""" Standardizes the test data according to the training data """
df_test = df_test_input.copy()
df_train = df_train_input.copy()
for c in df_test.columns:
df_test[c] = (df_test[c] - df_train.mean(axis=0)[c])/df_train.std(axis=0)[c]
df_test = df_test.fillna(0)
return df_test
###
def LSTM_df (covid_df, mobility_df, policy_df, use_data = 'ALL'):
""" Returns the data frame that can be used for LSTM input, use_data from 'ALL', 'C&D', 'MOB', 'POL' """
reindex_df = np.linspace(1, int(covid_df.shape[0]), int(covid_df.shape[0]), dtype=int)
covid_df = covid_df[['Date_reported', 'New_cases', 'Cumulative_cases', 'New_deaths', 'Cumulative_deaths']]
covid_df = covid_df.set_index('Date_reported')
mobility_df = mobility_df[['date', 'retail_and_recreation_percent_change_from_baseline', 'grocery_and_pharmacy_percent_change_from_baseline', 'parks_percent_change_from_baseline', 'transit_stations_percent_change_from_baseline', 'workplaces_percent_change_from_baseline', 'residential_percent_change_from_baseline']]
mobility_df = mobility_df.set_index('date')
policy_df = policy_df[['Date', 'C1_School closing', 'C2_Workplace closing', 'C3_Cancel public events', 'C4_Restrictions on gatherings', 'C5_Close public transport', 'C6_Stay at home requirements', 'C7_Restrictions on internal movement', 'C8_International travel controls', 'E1_Income support', 'E2_Debt/contract relief', 'E3_Fiscal measures', 'E4_International support', 'H1_Public information campaigns', 'H2_Testing policy', 'H3_Contact tracing', 'H4_Emergency investment in healthcare', 'H5_Investment in vaccines', 'H6_Facial Coverings', 'H7_Vaccination policy', 'H8_Protection of elderly people']]
policy_df = policy_df.set_index('Date')
if use_data == 'ALL':
lstm_df = pd.concat([covid_df, mobility_df, policy_df], axis=1)
elif use_data == 'C&D':
lstm_df = covid_df
elif use_data == 'MOB':
lstm_df = pd.concat([covid_df, mobility_df], axis=1)
elif use_data == 'POL':
lstm_df = pd.concat([covid_df, policy_df], axis=1)
return lstm_df
###
def train_test_data (df, start_date, end_date, factor=0.9):
""" Splits the data into test and train according to the input factor """
no_of_days = (end_date-start_date).days
test_data_start_date = start_date + datetime.timedelta(days=int(no_of_days*factor))
df_train = df.loc[:test_data_start_date].copy()
df_test = df.loc[test_data_start_date:].copy()
return df_train, df_test
###
def get_target_features(df_input, target_col, forecast_lead):
""" Gets the target and features from the data frame """
df = df_input.copy()
features = list(df.columns.difference([target_col]))
target = f"{target_col}_lead{forecast_lead}"
df[target] = df[target_col].shift(-forecast_lead)
df = df.iloc[:-forecast_lead]
return target, features, df
# + id="7e4W9wz1yQW0"
# Creating data-loader compatible dataset
class SequenceDataset(Dataset):
def __init__(self, dataframe, target, features, sequence_length=5):
self.features = features
self.target = target
self.sequence_length = sequence_length
self.y = torch.tensor(dataframe[target].values).float()
self.X = torch.tensor(dataframe[features].values).float()
def __len__(self):
return self.X.shape[0]
def __getitem__(self, i):
if i >= self.sequence_length - 1:
i_start = i - self.sequence_length + 1
x = self.X[i_start:(i + 1), :]
else:
padding = self.X[i].repeat(self.sequence_length - i - 1, 1)
x = self.X[0:(i + 1), :]
x = torch.cat((x, padding), 0)
return x, self.y[i]
# Creating data-loader compatible dataset
class SequenceDatasetTest(Dataset):
def __init__(self, dataframe, df2, target, features, sequence_length=5):
self.features = features
self.target = target
self.sequence_length = sequence_length
self.df2 = df2
self.y = torch.tensor(dataframe[target].values).float()
self.X = torch.tensor(dataframe[features].values).float()
self.X_train = torch.tensor(df2[features].values).float()
def __len__(self):
return self.X.shape[0]
def __getitem__(self, i):
if i >= self.sequence_length - 1:
i_start = i - self.sequence_length + 1
x = self.X[i_start:(i + 1), :]
else:
start = self.sequence_length - i - 1
padding = self.X_train[-start:]
x = self.X[0:(i + 1), :]
x = torch.cat((padding, x), 0)
return x, self.y[i]
# Creating LSTM model
class ShallowRegressionLSTM(nn.Module):
def __init__(self, num_features, hidden_units):
super().__init__()
self.num_features = num_features # this is the number of features
self.hidden_units = hidden_units
self.num_layers = 1
self.lstm = nn.LSTM(
input_size = num_features,
hidden_size=hidden_units,
batch_first=True,
num_layers=self.num_layers,
)
self.linear = nn.Linear(in_features=self.hidden_units, out_features=1)
def forward(self, x):
batch_size = x.shape[0]
h0 = torch.zeros(self.num_layers, batch_size, self.hidden_units).requires_grad_()
c0 = torch.zeros(self.num_layers, batch_size, self.hidden_units).requires_grad_()
_, (hn, _) = self.lstm(x, (h0, c0))
out = self.linear(hn[0]).flatten()
return out
###
def train_model(data_loader, model, loss_function, optimizer):
""" Funtion for training the model """
num_batches = len(data_loader)
total_loss = 0
model.train()
for X, y in data_loader:
output = model(X)
loss = loss_function(output, y)
optimizer.zero_grad()
loss.backward()
optimizer.step()
total_loss += loss.item()
avg_loss = total_loss / num_batches
print(f"Train loss: {avg_loss}")
###
def test_model(data_loader, model, loss_function):
""" Function for testing the model """
num_batches = len(data_loader)
total_loss = 0
model.eval()
with torch.no_grad():
for X, y in data_loader:
output = model(X)
total_loss += loss_function(output, y).item()
avg_loss = total_loss / num_batches
print(f"Test loss: {avg_loss}")
###
def predict(data_loader, model):
""" Uses model to predict """
output = torch.tensor([])
model.eval()
with torch.no_grad():
for X, _ in data_loader:
y_star = model(X)
output = torch.cat((output, y_star), 0)
return output
# + id="krdkb0o33DZ0"
def mainProgram (covid_df, mobility_df, policy_df, target_prediction = 'deaths', batch_size = 4, sequence_length = 500, lead = 14, learning_rate = 5e-5, num_hidden_units = 40, epochs = 100, ar_order = 14, ma_order = 0, d = 1):
""" The program puts everything together to give outputs of the LSTM model """
output_LSTM_w_all = {}
output_LSTM_w_mob = {}
output_LSTM_w_pol = {}
output_ARIMA = {}
true_target = {}
### Block 1
city_covid_df = filter_covid_data(covid_df)
city_len = len(city_covid_df)
city_mobility_data = filter_mobility_data(mobility_df)
city_policy_data = filter_policy_data(policy_df)
# options = ['ALL', 'MOB', 'POL']
options = ['ALL']
city = 'India'
for use_data in options:
city_lstm_df = LSTM_df(city_covid_df, city_mobility_data, city_policy_data, use_data)
#
city_target, city_features, city_lstm_df = get_target_features(city_lstm_df, target_prediction, forecast_lead=lead)
split = 0.8
city_train, city_test = train_test_data(city_lstm_df, datetime.date(2020, 1, 15), datetime.date(2021, 10, 24), split)
city_target_mean, city_target_sd, city_train_scaled = standardize_df(city_train, city_target)
city_test_scaled = scale_test_data(city_test, city_train)
#
torch.manual_seed(0)
city_train_dataset = SequenceDataset(city_train_scaled, city_target, city_features, sequence_length)
city_test_dataset = SequenceDatasetTest(city_test_scaled, city_train_scaled, city_target, city_features, sequence_length)
city_train_loader = DataLoader(city_train_dataset, batch_size=batch_size)
city_test_loader = DataLoader(city_test_dataset, batch_size=batch_size)
##
X, y = next(iter(city_train_loader))
model = ShallowRegressionLSTM(num_features = X.shape[2] , hidden_units=num_hidden_units)
loss_function = nn.MSELoss()
optimizer = torch.optim.Adam(model.parameters(), lr=learning_rate)
##
print("Untrained test\n--------")
test_model(city_test_loader, model, loss_function)
print()
for e in range(epochs):
print(f"Epoch {e+1}\n---------")
train_model(city_train_loader, model, loss_function, optimizer=optimizer)
test_model(city_test_loader, model, loss_function)
print()
city_train_eval_loader = DataLoader(city_train_dataset, batch_size=batch_size, shuffle=False)
ystar_col = "Model forecast"
city_train_scaled[ystar_col] = predict(city_train_eval_loader, model).numpy()
city_test_scaled[ystar_col] = predict(city_test_loader, model).numpy()
df_out = pd.concat((city_train_scaled, city_test_scaled))[[city_target, ystar_col]]
for c in df_out.columns:
df_out[c] = df_out[c] * city_target_sd + city_target_mean
n = len(df_out['Model forecast'])
y_pred_LSTM = list(df_out['Model forecast'][int(split*n):].values)
print("Test RMSE for LSTM-", use_data, "for", city, '=', round(mean_squared_error(y_pred_LSTM,df_out[city_target][int(split*n):])**0.5, 3))
print('\n')
if use_data == 'ALL':
output_LSTM_w_all[city] = y_pred_LSTM
elif use_data == 'MOB':
output_LSTM_w_mob[city] = y_pred_LSTM
elif use_data == 'POL':
output_LSTM_w_pol[city] = y_pred_LSTM
#### ARIMA
y_pred_arima = []
new_df = city_covid_df.copy()
# target_series is the cases/deaths for the particular state
target_series = list(new_df[target_prediction].values)
# splitting into history and test data
history = list(target_series[:int(split*n)])
test = list(target_series[int(split*n):])
for i in range(len(test)):
model = ARIMA(history, order=(lead,d,ma_order))
model_fit = model.fit()
yhat = model_fit.forecast()[0]
y_pred_arima.append(yhat)
history.append(yhat)
rmse_arima_prev = (mean_squared_error(test, y_pred_arima))**0.5
print("Test RMSE for ARIMA for ", city," = ", round(rmse_arima_prev,3))
print("\n")
output_ARIMA[city] = y_pred_arima
true_target[city] = test
return output_LSTM_w_all, output_LSTM_w_mob, output_LSTM_w_pol, output_ARIMA, true_target
# + [markdown] id="Fw3qsMRUKco1"
# ## Dataset
# Three data-sets are used as input feature to LSTM. These are:
#
#
# * [COVID-19 Cases and Deaths data from WHO.](https://data.humdata.org/dataset/coronavirus-covid-19-cases-and-deaths)
# * [Google's mobility trend data.](https://www.google.com/covid19/mobility/)
# * [Oxford's COVID-19 goverment response tracker.](https://github.com/OxCGRT/covid-policy-tracker)
#
#
#
#
# + id="bZPBx8dI3Gdo"
# Covid Cases and Deaths data
covid_df = pd.read_csv("/content/drive/MyDrive/CSE 8803 EPI/WHO-COVID-19-global-data.csv")
# Mobility Data
mobility_df = pd.read_csv("/content/drive/MyDrive/CSE 8803 EPI/Global_Mobility_Report.csv")
# Policy data
policy_df = pd.read_csv("/content/drive/MyDrive/CSE 8803 EPI/OxCGRT_latest.csv")
# + colab={"base_uri": "https://localhost:8080/"} id="c_rrp6X7hILJ" outputId="8d5773b7-c19b-4c04-fce2-648509b6514e"
output_LSTM_w_all, output_LSTM_w_mob, output_LSTM_w_pol, output_ARIMA, true_target = mainProgram (covid_df, mobility_df, policy_df, 'New_cases', batch_size = 50, sequence_length = 500, learning_rate = 5e-5, lead = 1, num_hidden_units = 10, epochs = 300)
# + colab={"base_uri": "https://localhost:8080/", "height": 531} id="HwrOLuDuLYGH" outputId="d710160d-ebd2-40cf-a84b-09d228d0620e"
# Plotting for all cities
region_list = ['India']
for city in region_list:
y_pred_LSTM_w_all = output_LSTM_w_all[city]
y_pred_arima = output_ARIMA[city]
test = true_target[city]
plt.figure(figsize=(15,8))
plt.title("Deaths forecasted for " + str(city), fontsize = 20)
plt.xticks(fontsize=15)
plt.yticks(fontsize=15)
plt.xlabel("Time step", fontsize=18)
plt.ylabel("Deaths", fontsize=18)
plt.plot(range(len(y_pred_LSTM_w_all)), y_pred_LSTM_w_all)
plt.plot(range(len(y_pred_arima)), y_pred_arima)
plt.plot(range(len(test)), test)
legend = ['LSTM with mob. and pol. data', 'ARIMA', 'Truth']
plt.legend(legend, fontsize=14)
plt.show()
# + id="IqfnrUdk2qqh"
| Kshitij_EPI_LSTM_Final_Code_14_Day_Lead.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1 - 自动化特征工程
#
# ```
# 描述:自动化机器学习的前提 --- 自动化特征工程。
# 作者:Chenyyx
# 时间:2020-01-09
# ```
#
# ```
# 目录:
# 1 - 特征工程简要介绍
# 2 - 3种自动化特征工程的生成方法
# 3 - Featuretools 库使用规则
# - 3.1 - 实体和实体集
# - 3.2 - DFS
# - 3.3 - 特征基元
# 4 - Featuretools 操作小实例
# 5 - 总结
# 6 - 参考链接
# ```
#
# ## 1 - 特征工程简要介绍
#
# **features(特征)** 是从数据中抽取出来对最终结果的预测有帮助的信息,**feature engineering(特征工程)** 则是特征在机器学习问题中使其算法和模型可以发挥更好的过程,该过程通常需要数据科学家根据经验找出最佳的特征组合形式,因为人的能力有限,所以找到的特征组合往往也不够全面,造成了效果和效率的局限性。而自动化特征工程可以根据数据特征进行自动组合,有效地解决了认为组合特征不全面和耗时的问题。
#
# 特征工程是一个与具体场景绑定的事情,因此自动化特征工程应该是一件根据模型选择数据类型等背景信息并进行自动化的工作。如果把自动化理解为不需要人工参与设计,那么实现自动化的方式多种多样,最简单的方式为遍历搜索,通过计算机遍历所有的可能组合也是一种自动化;通过模型的方法去完成同样是一种自动化,如通过神经网络自动完成图像与文本等的特征工程。
#
# **特征工程也被称为特征构造,是从现有数据中构造新的特征从而训练机器学习模型的过程**。这一步可能比实际上使用的模型更重要,因为一个机器学习算法只能从我们给定的数据中学习,所以构造一个和任务相关的特征是至关重要的。
#
# 因此,我们可以把自动化特征工程定义为如何根据具体场景去自动构建流程,而无需人工参与完成特征工程的一种方法。
#
# ## 2 - 3种自动化特征工程的生成方法
#
# 自动化特征工程的生成方法,分别是深度特征合成算法,`FeatureTools` 自动特征提取,基于时序特征的自动化特征工程。
#
# ### 2.1 - 深度特征合成算法
#
# 深度特征合成(Deep Feature Synthesis,DFS)是一种用于对关系数据和时间数据执行特征工程的自动方法。深度特征合成并不是通过深度学习生成特征,而是通过多重的特征叠加,一个特征的深度是构建这个特征所用到的基元的个数。关于 DFS 有如下 3 个关键的概念:
#
# - 特征主要来源是数据集中数据点之间的关系。
# - 对于不同的数据集,许多特征的构建是通过相似的数学运算得到的。
# - 新的特征是基于先前获取的特征构建的。
#
# ### 2.2 - Featuretools 自动特征提取
#
# Featuretools 是一个自动执行特征工程的开源库,它可以提高特征工程的效率,采用客观的方法来创建新的特征并将其用于机器学习的任务。Featuretools 将深度特征合成算法作为库的核心内容,以特征基元作为基本操作,通过叠加基元操作得到新的特征。
#
# ### 2.3 - 基于时序数据的自动化特征工程
#
# 时序数据即时间序列数据,是同一指标按时间顺序记录的数据列。
#
# 时序数据在数据分析和机器学习中具有很重要的意义,从时序数据中提取特征也是数据预处理中的关键一步。 TSFRESH 是一种兼容于 Pandas 和 scikit-learn 接口的 Python 包,用于提取时序数据特征。
#
# 下面,我会主要介绍 `FeatureTools` 自动特征提取。其余两个简要介绍一下。下面会有对应的 `Python` 代码,记得结合使用规则一起查看。
#
# ## 3 - Featuretools 库使用规则
#
# `Featuretools` 是基于数据实体和实体时间的关系,基于 `DFS` 算法使用特征基元等操作来实现自动化的特征提取。
#
# ### 3.1 - 实体和实体集
#
# 一个实体集(entity set)是实体(entity)和实体之间关系(relationship)的集合,在 Featuretools 中,实体集有利于准备和结构化数据集用作特征工程。
#
# 原始的数据表一般以 DataFrame 的形式保存,每一张数据表都是一个实体,而每个实体都必须含有唯一元素的索引,用来区分不同的数据。我们需要为这些实体构建有利于 Featuretools 使用的实体集。
#
# - 1 - 创建实体集
# 实体集的初始化可以使用 `ft.EntitySet(id="name")` 函数来构建一个新的实体集,如果需要对实体集命名,修改 `id` 参数即可对实体集命名。
#
# - 2 - 添加实体
# 实体集创建成功后需要像实体集内添加实体,使用 `ft.entity_from_dataframe()` 函数卡伊通过修改以下参数,来实现对函数功能的选择:
# - `entity_id`:设置添加实体的名字。
# - `dataframe`:表。
# - `index` 和 `time_index`:实体的索引。
# - `variable_types`:字典可以对实体中的数据类型进行定义。
# - 3 - 添加实体间的关系
# 最后我们需要添加实体集中各个实体之间的关系,通过 `ft.relationship()` 函数添加实体之间的关系,也可以通过 `EntitySet.add_relationship(new_relationship)` 来添加新的实体关系。
# 创建好实体集后就可以将实体集用于后续的特征提取。
#
# ### 3.2 - DFS
#
# 如果不使用自动化特征工程,则可以直接运行 `DFS` 来生成特征,通过 `ft.dfs()` 函数来生成特征矩阵,`dfs` 函数具有以下参数:
#
# - `Entityset`:实体集名称。
# - `target_entity`:实体集中的目标实体。
# - `primitives`:选择使用的特征基元操作列表,这里的基元操作有两种,一种是聚合基元,一种是转换基元。
# - `max_depth`:特征叠加深度。通过叠加产生的特征比通过单个特征基元产生的特征更加具有表现力,这就能够为机器学习模型提供更为复杂有效的特征。
#
# ### 3.3 - 特征基元
#
# 特征基元是 `Featuretools` 用来自动构建特征的基础操作,通过单独使用或者叠加使用特征基元构造新的特征。使用特征基元的意义在于,只要限制输入和输出的数据类型,就可以在不同的数据集中采用相同的特征基元操作。
#
# 特征基元有如下两种:
#
# - 聚合基元:根据父表与子表的关联,在不同的实体间完成对子表的统计操作。
# - 转换基元:转换基元是对单个实体进行的操作,对实体的一个或者多个变量进行操作,并为该实体统计出一个新的变量。
#
# 如果需要自动提取特征,只需要调用相同的 `ft.dfs()` 函数,但是不传入 `agg_primitives` 选择特征基元,就可以让 `Featuretools` 自动生成特征。
#
# 除此之外,我们还可以通过 API 来定义自己的特征基元,确定特征基元的类型,定义输入和输出的数据类型,在 Python 中编写该特征基元的功能函数,就可以实现特征基元,并可以在和其他基元的叠加中使用。
#
# `Featuretools` 库以 `DFS` 为核心,通过叠加使用特征基元操作,能够构建大量有效的特征,为自动化特征工程提供了很大的帮助。
#
# ## 4 - Featuretools 操作小实例
#
# 下面我们看一下怎么使用 `Featuretools` 这个工具库。
# 导入对应的库
import pandas as pd
import numpy as np
import featuretools as ft
# 导入了对应的科学计算库,接下来我们加载数据,实验数据是我根据网上的一些数据,将大数据文件,只截取一部分变成小数据文件而来的。
# 加载数据
clients = pd.read_csv('data/clients.csv', parse_dates=['joined'])
loans = pd.read_csv('data/loans.csv', parse_dates=['loan_start', 'loan_end'])
payments = pd.read_csv('data/payments.csv', parse_dates=['payment_date'])
# 查看一下 clients 的前5行数据
clients.head()
# 查看一下 loans 的前5行数据
loans.head()
# 查看一下 payments 的前5行数据
payments.head()
# 构造特征是一个非常耗时的过程,因为每个新的特征通常需要几步才能构造,特别是当使用多张表的信息时。我们可以将特征构造的操作分为两类:【转换】 和 【聚合】。后面我们通过例子来看看这些概念的实际应用。
#
# 通过从一或多列中构造新的特征,【转换】作用于单张表(在 Python 中,表是一个 Pandas DataFrame)。举个例子,如有以上的 clients 表数据。我们可以通过查找 joined 列中的月份或是自然对数化 income 列的数据来构造新的特征。这些都是转换操作,因为它只使用了一张表的信息。
#
# 另一方面,【聚合】是跨表实现的,并使用一对多的关联来对观测值分组,然后计算统计量。例如,仍然看上面的结果,若我们有另外一张包含客户贷款信息的表格,其中每个客户可能有多项贷款,我们便可以计算每个客户贷款的平均值,最大值和最小值等统计量。
#
# 这个过程包括根据不同客户对贷款表进行分组并计算聚合后的统计量,然后将结果整合到客户数据中。实际上,这些操作本身并不困难,但是如果有数百个变量分布在数十张表中,这个过程将无法通过人工完成。理想情况下,我们希望有一个解决方案能够在不同表间自动执行转换和聚合操作,并将结果整合到一张表中。
#
# --------------------------------------------------------------------------------------
#
# 幸运的是,Featuretools 正是我们正在寻找的解决方案。我们首先明确一下接下来我们要进行操作的数据信息:
#
# - clients:关于信用社客户的基本信息。每个客户只对应数据框中的一行。
# - loans:向用户提供的贷款。每项贷款只对应数据框中的一行,但是客户可能有多项贷款。
# - payments:贷款还本的支付。每笔支付只对应一行,但是每项贷款可以有多笔支付。
#
# 如果我们有一个机器学习任务,例如预测客户未来是否会偿还一项贷款,我们希望将所有关于客户的信息整合到一张表中。这些表是相关的(通过 client_id 和 loan_id 变量),并且我们可以通过一系列转换和聚合操作来人工实现这个过程。然而,我们很快就可以使用特征工具来自动实现这个过程。
#
# 使用特征工具的前两个概念是 【实体】 和 【实体集】。一个实体就是一张表(或是 Pandas 中的一个 DataFrame(数据框))。一个实体集是一组表以及它们之间的关联。将一个实体集看成另一种 Python 数据结构,并带有自己的方法和属性。
#
# 现在我们需要整合两个实体。每个实体都必须带有一个索引,它是一个包含所有唯一元素的列。就是说,索引中的每个值只能在表中出现一次。在 clients 数据框中的索引是 client_id,因为每个客户在该数据框中只对应一行。我们使用 `entity_from_dataframe` 将一个带有索引的实体添加一个实体集中:
# +
# 创建实体集,名称叫 clients
es = ft.EntitySet(id='clients')
# 添加 clients 实体
es = es.entity_from_dataframe(entity_id = 'clients', dataframe = clients,
index = 'client_id', time_index = 'joined')
# -
# loans 数据框中还有另外一个唯一的索引,loan_id,同时将其添加到实体集的语法与 clients 一样。然而,payments 数据框不存在唯一索引。当我们把 payments 数据框添加到实体集中时,我们需要传入参数 make_index=True,同时指定索引的名字。另外,尽管特征工具能自动推断实体中每列的数据类型,但是我们可以通过将列数据类型的字典传递给参数 variable_types 来覆盖它。
# +
# 添加 loans 实体
es = es.entity_from_dataframe(entity_id = 'loans', dataframe = loans,
variable_types = {'repaid': ft.variable_types.Categorical},
index = 'loan_id',
time_index = 'loan_start')
# 添加 payments 实体
es = es.entity_from_dataframe(entity_id = 'payments', dataframe = payments,
variable_types = {'missed': ft.variable_types.Categorical},
make_index = True,
index = 'payment_id',
time_index = 'payment_date')
# 添加完成了实体集,将实体集打印出来看看
es
# -
# 对于此数据框,尽管 missed 是一个整数,但是它不是一个数值变量,因为它只能取 2 个离散的数值,所以在特征工具中,将其看成一个分类变量。
#
# 接下来,我们考虑两张表之间 【关联】的最好方法是类比父子之间的关联。这是一种一对多的关联:每个父亲可以有多个儿子。对表来说,每个父亲对应一张父表中的一行,但是子表中可能有多行对应于同一张父表中的多个儿子。
#
# 例如,在我们的数据集中,clients 数据框是 loans 数据框的一张父表。每个客户只对应 clients 表中的一行,但是可能对应 loans 表中的多行。同样,loans 表是 payments 表的一张父表,因为每项贷款可以有多项支付。父亲通过共享变量与儿子相关联。当我们执行聚合操作的时候,我们根据父变量对子表进行分组,并计算每个父亲的儿子的统计量。
#
# 为了形式化特征工具中的关联规则,我们仅需指定链接两张表的变量。clients 表和 loans 表通过 client_id 变量连接,同时 loans 和 payments 表通过 loan_id 变量连接。创建关联并将其添加到实体集中的语法如下所示:
# +
# 添加实体关系
# 通过 client_id 关联 clients 和 loans 实体
r_client_previous = ft.Relationship(es['clients']['client_id'],
es['loans']['client_id'])
# 将 relationship 加入到实体集中
es = es.add_relationship(r_client_previous)
# 通过 loan_id 关联 payments 和 loans 实体
r_payments = ft.Relationship(es['loans']['loan_id'],
es['payments']['loan_id'])
es = es.add_relationship(r_payments)
# 为实体集中的实体添加了关联,将实体集打印出来看看
es
# -
# 该实体集现在包含三个实体(表),以及将这些表连接在一起的关联规则。在添加实体和形式化关联规则之后,实体集就完整了并准备好从中构造新的特征。
#
# **特征基元**
#
# 在我们深入了解深度特征合成之前,我们需要了解特征基元的概念。我们其实早就知道是什么了,只是我们刚刚用不同的名字来称呼它们!它们只是我们用来构造新特征的操作:
#
# - 聚合:根据父与子(一对多)的关联完成的操作,也就是根据父亲分组并计算儿子的统计量。一个例子就是根据 client_id 对 loan 表分组并找到每个客户的最大贷款额。
# - 转换:对一张表中一或多列完成的操作。一个例子就是取一张表中两列之间的差值或者取一列的绝对值。
#
# 在 featuretools 中单独使用这些基元或者叠加使用这些基元可以构造新的特征。以下是特征工具中的一些特征基元的列表,也可以自定义特征基元。
#
# 
#
# 这些基元可以单独使用或者是组合使用以构造新的特征。为了使用特定的基元构造新的特征,我们使用 `ft.dfs` 函数(代表深度特征合成)。我们传入 `entityset` 和 `target_entity`,这是我们想要在其中添加特征的表,被选参数 `trans_primitives(转换)` 和 `agg_primitives(聚合)`。
# 聚合特征,并生成新的特征
features, feature_name = ft.dfs(entityset=es, target_entity='clients')
features.head()
# +
# 查看一下所有 primitives 的操作,以便对下面的操作有个简单的了解
primitives = ft.list_primitives()
pd.options.display.max_colwidth = 100
primitives[primitives['type'] == 'aggregation'].head(30)
# -
primitives[primitives['type'] == 'transform'].head(60)
# +
# 聚合特征,通过制定聚合 agg_primitives 和转换 trans_primitives 生成新特征
features_2, feature_names_2 = ft.dfs(entityset=es, target_entity='clients',
agg_primitives=['mean', 'max', 'percent_true', 'last'],
trans_primitives=['year', 'month', 'subtract_numeric', 'divide_numeric'])
features_2.head()
# -
# 可以看到上边我们进行了简单的 `转换` 和 `聚合` 操作。
#
# 特征工具构造了很多特征供我们使用。尽管这个过程确实能自动构造新的特征,但是它不会取代数据科学家,因为我们仍然需要弄清楚如何处理这些特征。例如,我们的目的是预测一位客户是否会偿还贷款,我们可以寻找与特定结果最相关的特征。此外,如果我们具有领域知识,我们可以用这些知识来选择指定的特征基元或候选特征的种子深度特征合成。
#
# 特征工程自动化解决了一个问题,但是却带来了另外的一个问题:**特征太多了**。尽管在拟合一个模型之前很难说哪些特征是重要的,但很可能不是所有这些特征都与我们想要训练的模型的任务相关。此外,拥有太多特征(参见《Irrelevant Features and the Subset Selection Problem》)可能会导致模型性能不佳,因为较无益的特征会淹没那些更重要的特征。
#
# **特征过多问题以维度灾难著称。随着特征数量的上升(数据维度增长),模型越来越难以学习特征与目标之间的映射关系。事实上,让模型表现良好所需的数据量与特征数量成指数关系**。
#
# 维度灾难与特征降维(也叫特征选择,去除不相关特征的过程)相对。这可以采用多种形式:`主成分分析(PCA)`、`SelectKBest`、使用模型中特征的重要性或使用深度神经网络进行自编码。但是,特征降维是另一篇文章的不同主题。到目前为止,我们知道我们可以使用特征工具以最小的努力从许多表中构造大量的特征!
#
# ## 5 - 总结
#
# 与机器学习中的许多主题一样,使用特征工具进行特征工程自动化是一个基于简单想法的复杂概念。使用实体集、实体和关联的概念,`featuretools` 可以执行深度特征合成操作来构造新的特征。深度特征合成可以依次叠加特征基元:【聚合】,它们在多张表间的一对多关联中起作用,以及【转换】,是应用于单张表中一或多列以从多张表中构造新的特征的函数。
#
# 下面是更具体的一些小总结:
#
# - 实体(二维表):每个实体必须要有一个唯一的索引,若没有索引,则需要设置 `make_index=True` 参数。
# - time_index(时间索引):表示该行数据(该条数据)的信息被记录(知晓)的时间。
# - 创建实体集,需要设置一个实体集的 id。
# - 变量类型,Variable Types:ft 会根据变量的不同类型施加不同的特征衍生操作,虽然 ft 可以推测变量类型,但是有时还是尽量指明变量类型,如 boolean 变量,以避免特征衍生过程中出现对 boolean 变量的求平均值,最大最小值等无意义的合成特征。
# - 对于无意义的变量(如子表中的 ID),需要先删除这些字段,以避免在根据这些无实际意义的变量(字段)生成更多无意义的新的特征。
# - 在创建实体之间关系的时候,若存在多种可以使用的关联关系(键),一定要根据数据表之间的实际业务逻辑场景关系确定合适的连接方案,特别是涉及到三级表之间的关联时候:A-B-C(A为B的父表,B为C的父表,A也可以与C通过特定的键联系),这一点非常重要。对于最终确定的表之间联系的逻辑图后,最好可以将表之间的关系画出来,防止混乱。同时,需要将表C中没有使用的键(没有实际业务意义),需要做删除处理,对应上文第5条所解释的要求。
# - Feature Primitives:特征基元,一个 FP 是对几张表或者是表的一组子集进行的一个操作,目的是创建一个新的特征。这些操作本质也非常简单,但是这些简单的操作可以相互叠加,进而创造出非常复杂的特征。特征基元主要分为两类:
# - 聚合:对于每一张父表,对子表的数据进行统计量的计算,如 min,max,mean,std,var 等。
# - 转换:计算单一一张表中的一列或者几列。如计算两列之间的差值等。
# - 查看特征基元的方法:`ft.list_primitives()`。
#
# ## 6 - 参考链接
#
# - https://blog.csdn.net/hellozhxy/article/details/80772872
# - https://docs.featuretools.com/en/stable/
# - https://www.jianshu.com/p/71782dbe2e1e
# - https://blog.csdn.net/qq_40802887/article/details/88765543
# - https://github.com/Featuretools/Automated-Manual-Comparison/blob/master/Loan%20Repayment/notebooks/Automated%20Loan%20Repayment.ipynb
| 1_auto_feature.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import re
import numpy as np
import tensorflow as tf
np.random.seed(1)
tf.random.set_seed(2)
import pandas as pd
import keras
# from tqdm import tqdm
from tqdm import tqdm_notebook as tqdm
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import LabelEncoder
from sklearn.utils import class_weight
from sklearn.metrics import f1_score, classification_report, log_loss
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
from keras.utils import to_categorical
from keras.models import Sequential
from keras.layers import Embedding, LSTM, Dense, SpatialDropout1D, Bidirectional, Flatten
from keras.layers import Dropout, Conv1D, GlobalMaxPool1D, GRU, GlobalAvgPool1D
from keras.optimizers import Adam
from keras.callbacks import ModelCheckpoint, EarlyStopping
#print(os.listdir('../LSTM'))
# +
data = pd.read_csv('SB-multi-LSTM.csv')
data= data[data['hour']!='dummy']
data['A1']=data['A1'].astype("category")
data['A2']=data['A2'].astype("category")
data['A3']=data['A3'].astype("category")
data['sex']=data['sex'].astype("category")
data['age']=data['age'].astype("category")
data['org']=data['org'].astype("category")
data['weekday']=data['weekday'].astype("category")
data['RMV']=data['RMV'].astype("category")
data['hour']=data['hour'].astype("int")
data['month']=data['month'].astype("int")
data['truth']=data['truth'].astype("category")
data.head()
# #data['truth']=np.where(data['truth'] ==1, 'YY', data['truth'])
# data['truth']=np.where(data['truth'] ==2, '0', data['truth'])
# data['truth']=np.where(data['truth'] =='3', '0', data['truth'])
# data['truth']=np.where(data['truth'] =='4', '0', data['truth'])
# data['truth']=np.where(data['truth'] =='5', '0', data['truth'])
# #data['A1']=np.where(data['A1'] ==1, 'YY', data['A1'])
# data['A1']=np.where(data['A1'] ==2, '0', data['A1'])
# data['A1']=np.where(data['A1'] =='3', '0', data['A1'])
# data['A1']=np.where(data['A1'] =='4', '0', data['A1'])
# data['A1']=np.where(data['A1'] =='5', '0', data['A1'])
# #data['A2']=np.where(data['A2'] ==1, 'YY', data['A2'])
# data['A2']=np.where(data['A2'] ==2, '0', data['A2'])
# data['A2']=np.where(data['A2'] =='3', '0', data['A2'])
# data['A2']=np.where(data['A2'] =='4', '0', data['A2'])
# data['A2']=np.where(data['A2'] =='5', '0', data['A2'])
# #data['A3']=np.where(data['A3'] ==1, 'YY', data['A3'])
# data['A3']=np.where(data['A3'] ==2, '0', data['A3'])
# data['A3']=np.where(data['A3'] =='3', '0', data['A3'])
# data['A3']=np.where(data['A3'] =='4', '0', data['A3'])
# data['A3']=np.where(data['A3'] =='5', '0', data['A3'])
data.columns
# -
data= data[data['truth']!=5]
data= data[data['A1']!=5]
data= data[data['A2']!=5]
data= data[data['A3']!=5]
data= data[data['RMV']!=5]
data= data[data['REM']!=5]
data= data[data['RGLAD']!=5]
data= data[data['RLFC']!=5]
# +
data = pd.get_dummies(data, prefix=['A1', 'A2', 'A3', 'weekday', 'month', 'age', 'sex',
'org', 'RMV', 'REM', 'RGLAD', 'RLFC'], columns=['A1', 'A2', 'A3', 'weekday', 'month', 'age', 'sex',
'org', 'RMV', 'REM', 'RGLAD', 'RLFC'], drop_first=False)
data.head()
# -
data.columns
# +
X = data.drop('truth', axis=1)
y = data['truth']
# +
from sklearn import preprocessing
from keras.utils import np_utils
from keras.utils import to_categorical
# label_encoder object knows how to understand word labels.
# label_encoder = preprocessing.LabelEncoder()
# # Encode labels in column 'species'.
# y = label_encoder.fit_transform(y)
# encode class values as integers
# encoder = LabelEncoder()
# encoder.fit(y)
# encoded_Y = encoder.transform(y)
# # convert integers to dummy variables (i.e. one hot encoded)
# dummy_y = np_utils.to_categorical(encoded_Y)
# label_encoder object knows how to understand word labels.
label_encoder = preprocessing.LabelEncoder()
# Encode labels in column 'species'.
y = label_encoder.fit_transform(y)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.30, random_state=42)
from keras.utils import to_categorical
y_train = to_categorical(y_train)
y_test = to_categorical(y_test)
# make the model and load the training dataset.
#y_train = to_categorical(y_train)
# -
y_train
# +
dic = {"`":"'", "’":"'", "‘":"'", "´":"'","ain't": "is not", "aren't": "are not","can't": "cannot", "'cause": "because", "could've": "could have",
"couldn't": "could not", "didn't": "did not", "doesn't": "does not", "don't": "do not", "hadn't": "had not",
"hasn't": "has not", "haven't": "have not", "he'd": "he would","he'll": "he will", "he's": "he is", "how'd": "how did",
"how'd'y": "how do you", "how'll": "how will", "how's": "how is", "I'd": "I would", "I'd've": "I would have",
"I'll": "I will", "I'll've": "I will have","I'm": "I am", "I've": "I have", "i'd": "i would", "i'd've": "i would have",
"i'll": "i will", "i'll've": "i will have","i'm": "i am", "i've": "i have", "isn't": "is not", "it'd": "it would",
"it'd've": "it would have", "it'll": "it will", "it'll've": "it will have","it's": "it is", "let's": "let us",
"ma'am": "madam", "mayn't": "may not", "might've": "might have","mightn't": "might not","mightn't've": "might not have",
"must've": "must have", "mustn't": "must not", "mustn't've": "must not have", "needn't": "need not",
"needn't've": "need not have","o'clock": "of the clock", "oughtn't": "ought not", "oughtn't've": "ought not have",
"shan't": "shall not", "sha'n't": "shall not", "shan't've": "shall not have", "she'd": "she would",
"she'd've": "she would have", "she'll": "she will", "she'll've": "she will have", "she's": "she is",
"should've": "should have", "shouldn't": "should not", "shouldn't've": "should not have", "so've": "so have",
"so's": "so as", "this's": "this is","that'd": "that would", "that'd've": "that would have", "that's": "that is",
"there'd": "there would", "there'd've": "there would have", "there's": "there is", "here's": "here is","they'd": "they would",
"they'd've": "they would have", "they'll": "they will", "they'll've": "they will have", "they're": "they are",
"they've": "they have", "to've": "to have", "wasn't": "was not", "we'd": "we would", "we'd've": "we would have",
"we'll": "we will", "we'll've": "we will have", "we're": "we are", "we've": "we have", "weren't": "were not",
"what'll": "what will", "what'll've": "what will have", "what're": "what are", "what's": "what is", "what've": "what have",
"when's": "when is", "when've": "when have", "where'd": "where did", "where's": "where is", "where've": "where have",
"who'll": "who will", "who'll've": "who will have", "who's": "who is", "who've": "who have", "why's": "why is",
"why've": "why have", "will've": "will have", "won't": "will not", "won't've": "will not have", "would've": "would have",
"wouldn't": "would not", "wouldn't've": "would not have", "y'all": "you all", "y'all'd": "you all would",
"y'all'd've": "you all would have","y'all're": "you all are","y'all've": "you all have","you'd": "you would",
"you'd've": "you would have", "you'll": "you will", "you'll've": "you will have", "you're": "you are", "you've": "you have",
"It's": 'It is', "Can't": 'Can not', "I've": 'I have', "\n":" ", "—":"", ".":"", "…":"", "!":"", ":":" ",
"-":" ","•":""}
#"😍":"lovely"
# -
def preprocess_text(sen):
# Remove punctuations and numbers
sentence = re.sub('[^a-zA-Z]', ' ', sen)
# Single character removal
sentence = re.sub(r"\s+[a-zA-Z]\s+", ' ', sentence)
# Removing multiple spaces
sentence = re.sub(r'\s+', ' ', sentence)
for i, j in dic.items():
sentence = sentence.lower().replace(i, j)
sentence = re.sub(r"http[s]?://t.co/[A-Za-z0-9]*"," ",sentence) #URLs
return sentence
X1_train = []
sentences = list(X_train["text"])
for sen in sentences:
X1_train.append(preprocess_text(sen))
X1_test = []
sentences = list(X_test["text"])
for sen in sentences:
X1_test.append(preprocess_text(sen))
# +
tokenizer = Tokenizer()
tokenizer.fit_on_texts(X1_train)
X1_train = tokenizer.texts_to_sequences(X1_train)
X1_test = tokenizer.texts_to_sequences(X1_test)
vocab_size = len(tokenizer.word_index) + 1
maxlen = 27
X1_train = pad_sequences(X1_train, padding='post', maxlen=maxlen)
X1_test = pad_sequences(X1_test, padding='post', maxlen=maxlen)
# +
from numpy import array
from numpy import asarray
from numpy import zeros
embeddings_dictionary = dict()
# glove_file = open('glove.twitter.27B.200d.txt', encoding="utf8")
# for line in glove_file:
# records = line.split()
# word = records[0]
# vector_dimensions = asarray(records[1:], dtype='float32')
# embeddings_dictionary[word] = vector_dimensions
# glove_file.close()
embedding_matrix = zeros((vocab_size, 200))
for word, index in tokenizer.word_index.items():
embedding_vector = embeddings_dictionary.get(word)
if embedding_vector is not None:
embedding_matrix[index] = embedding_vector
# -
X.columns
# +
X2_train = X_train[['A1_1', 'A1_2', 'A1_3', 'A1_4', 'A1_5', 'A2_1', 'A2_2',
'A2_3', 'A2_4', 'A2_5', 'A3_1', 'A3_2', 'A3_3', 'A3_4', 'A3_5',
'RMV_1', 'RMV_2', 'RMV_3', 'RMV_4', 'RMV_5', 'REM_1',
'REM_2', 'REM_3', 'REM_4', 'RGLAD_1', 'RGLAD_2', 'RGLAD_3', 'RGLAD_4',
'RLFC_1', 'RLFC_2', 'RLFC_3', 'RLFC_4']].values
X2_test = X_test[['A1_1', 'A1_2', 'A1_3', 'A1_4', 'A1_5', 'A2_1', 'A2_2',
'A2_3', 'A2_4', 'A2_5', 'A3_1', 'A3_2', 'A3_3', 'A3_4', 'A3_5',
'RMV_1', 'RMV_2', 'RMV_3', 'RMV_4', 'RMV_5', 'REM_1',
'REM_2', 'REM_3', 'REM_4', 'RGLAD_1', 'RGLAD_2', 'RGLAD_3', 'RGLAD_4',
'RLFC_1', 'RLFC_2', 'RLFC_3', 'RLFC_4']].values
ncols=X2_train.shape[1]
# -
ncols
# +
from keras.layers import Input
from keras.layers import Dropout, Conv1D, GlobalMaxPool1D, GRU, GlobalAvgPool1D
input_1 = Input(shape=(maxlen,))
input_2 = Input(shape=(ncols,))
# -
embedding_layer = Embedding(vocab_size, 200, weights=[embedding_matrix], trainable=False)(input_1)
#LSTM_Layer_1 = Bidirectional(LSTM(128, dropout=0.25, recurrent_dropout=0.25))(embedding_layer)
LSTM_Layer_1 = LSTM(128, dropout=0.25, recurrent_dropout=0.25)(embedding_layer)
LSTM_Layer_2 = Dense(64, activation='relu')(LSTM_Layer_1)
dense_layer_1 = Dense(10, activation='relu')(input_2)
dense_layer_2 = Dense(64, activation='relu')(dense_layer_1)
# +
from keras.layers import concatenate
from keras.models import Model
from keras.layers import Dropout, Conv1D, GlobalMaxPool1D, GRU, GlobalAvgPool1D
concat_layer = concatenate([LSTM_Layer_2, dense_layer_2])
dense_layer_3 = Dense(10, activation='relu')(dense_layer_2)
output = Dense(4, activation='softmax')(dense_layer_3)
model = Model(inputs=input_2, outputs=output)
# +
from keras import backend as K
import tensorflow as tf
from sklearn.metrics import roc_auc_score
def recall_m(y_true, y_pred):
true_positives = K.sum(K.round(K.clip(y_true * y_pred, 0, 1)))
possible_positives = K.sum(K.round(K.clip(y_true, 0, 1)))
recall = true_positives / (possible_positives + K.epsilon())
return recall
def precision_m(y_true, y_pred):
true_positives = K.sum(K.round(K.clip(y_true * y_pred, 0, 1)))
predicted_positives = K.sum(K.round(K.clip(y_pred, 0, 1)))
precision = true_positives / (predicted_positives + K.epsilon())
return precision
def f1_m(y_true, y_pred):
precision = precision_m(y_true, y_pred)
recall = recall_m(y_true, y_pred)
return 2*((precision*recall)/(precision+recall+K.epsilon()))
def auroc(y_true, y_pred):
return tf.py_function(roc_auc_score, (y_true, y_pred), tf.double)
model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['acc', f1_m,precision_m, recall_m])
print(model.summary())
# -
from keras.utils import plot_model
plot_model(model, to_file='model_plot3.png', show_shapes=True, show_layer_names=True)
history = model.fit(x=X2_train, y=y_train, batch_size=128, epochs=50, verbose=1, validation_split=0.2)
# +
loss, accuracy, f1_score, precision, recall = model.evaluate(x=X2_test, y=y_test, verbose=1)
# +
import matplotlib.pyplot as plt
plt.plot(history.history['acc'])
plt.plot(history.history['val_acc'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train','test'], loc='upper left')
plt.show()
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train','test'], loc='upper left')
plt.show()
# +
print(__doc__)
import matplotlib.pyplot as plt
import numpy as np
from sklearn import svm, datasets
from sklearn.metrics import precision_recall_curve
from sklearn.metrics import average_precision_score
from sklearn.preprocessing import label_binarize
from sklearn.multiclass import OneVsRestClassifier
from sklearn.model_selection import train_test_split
predictions = model.predict(X2_test)
y_pred = (predictions > 0.7)
y_test = label_binarize(y_test, classes=['YY','YN','NY','NN'])
y_pred = label_binarize(y_pred, classes=['YY','YN','NY','NN'])
n_classes = 4
# Compute Precision-Recall and plot curve
precision = dict()
recall = dict()
average_precision = dict()
for i in range(n_classes):
precision[i], recall[i], _ = precision_recall_curve(y_test[:, i],
y_pred[:, i])
average_precision[i] = average_precision_score(y_test[:, i], y_pred[:, i])
# Compute micro-average ROC curve and ROC area
precision["micro"], recall["micro"], _ = precision_recall_curve(y_test.ravel(),
y_pred.ravel())
average_precision["micro"] = average_precision_score(y_test, y_pred,
average="micro")
# Plot Precision-Recall curve
plt.clf()
plt.plot(recall[0], precision[0], label='Precision-Recall curve')
plt.xlabel('Recall')
plt.ylabel('Precision')
plt.ylim([0.0, 1.05])
plt.xlim([0.0, 1.0])
plt.title('Precision-Recall example: AUC={0:0.2f}'.format(average_precision[0]))
plt.legend(loc="lower left")
plt.show()
# Plot Precision-Recall curve for each class
plt.clf()
plt.plot(recall["micro"], precision["micro"],
label='micro-average Precision-recall curve (area = {0:0.2f})'
''.format(average_precision["micro"]))
for i in range(n_classes):
plt.plot(recall[i], precision[i],
label='Precision-recall curve of class {0} (area = {1:0.2f})'
''.format(i, average_precision[i]))
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('Recall')
plt.ylabel('Precision')
plt.title('Extension of Precision-Recall curve to multi-class')
plt.legend(loc="lower right")
plt.show()
# -
matrix = metrics.confusion_matrix(y_test.argmax(axis=1), y_pred.argmax(axis=1))
matrix
# +
import shap
explainer = shap.DeepExplainer(model, X1_train[:100])
shap_values = explainer.shap_values(X1_test[:10])
# -
f=shap.force_plot(explainer.expected_value, shap_values, X, show=False)
shap.save_html("index.htm", f)
auc_score
# +
from matplotlib import colors as plt_colors
import numpy as np
import shap
import matplotlib.pyplot as pl
shap.summary_plot(shap_values, X, show=False)
pl.savefig("ranks.pdf", dpi=800, bbox_inches = 'tight')
pl.show()
# -
# -------------------------------------------
#
# +
from cleanlab.latent_estimation import estimate_cv_predicted_probabilities
import cleanlab
s = np.array(y)
probabilities = estimate_cv_predicted_probabilities(
X,
s,
clf=model,
)
psx = cleanlab.latent_estimation.estimate_cv_predicted_probabilities(
X, s, clf=model)
# -
import pandas as pd
import numpy as np
df=data
for tweet, label in df.sample(10)[["text", "result"]].values:
print(label, tweet)
# +
import tensorflow_hub as hub
# embed = hub.load("https://tfhub.dev/google/universal-sentence-encoder-large/4")
# -
X_train_embeddings = embed(df.text.values)
# +
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import accuracy_score, f1_score
from sklearn.ensemble import RandomForestClassifier
logistic_clf = RandomForestClassifier(n_estimators=100, max_depth= 30, max_features=.75, criterion='entropy', min_samples_split=15, class_weight='balanced')
# -
logistic_clf.fit(X_train_embeddings['outputs'][:3000,:],
df.result.values[:3000])
RandomForestClassifier(n_estimators=100, max_depth= 30, max_features=.75, criterion='entropy', min_samples_split=15, class_weight='balanced')
# +
y_pred = logistic_clf.predict(X_train_embeddings['outputs'][3000:,:])
y_pred_proba = logistic_clf.predict_proba(X_train_embeddings['outputs'][3000:,:])
print("Accuracy: {:.1%}".format(accuracy_score(df.result.values[3000:], y_pred)))
print("F1: {:.1%}".format(f1_score(df.result.values[3000:], y_pred)))
# +
from cleanlab.pruning import get_noise_indices
ordered_label_errors = get_noise_indices(
s=df.result.values[3000:],
psx=y_pred_proba,
sorted_index_method='normalized_margin', # Orders label errors
)
# +
print("We found {} label errors.".format(len(ordered_label_errors)))
# -
ordered_label_errors
df.loc[220]
error_df = df.loc[df.index[3000:]].loc[df.index[3000+ordered_label_errors]]
for tweet, label in error_df[[ "text", "result"]][:30].values:
print(label, tweet)
| LSTM/LSTM-SB-Meta.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="b518b04cbfe0"
# ##### Copyright 2020 The TensorFlow Authors.
# + cellView="form" id="906e07f6e562"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="e2d97c7e31aa"
# # Making new Layers and Models via subclassing
# + [markdown] id="4e352274064f"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/guide/keras/custom_layers_and_models"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/snapshot-keras/site/en/guide/keras/custom_layers_and_models.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/keras-team/keras-io/blob/master/guides/making_new_layers_and_models_via_subclassing.py"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/guide/keras/custom_layers_and_models.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# </table>
# + [markdown] id="8d4ac441b1fc"
# ## Setup
# + id="4e7dce39dd1d"
import tensorflow as tf
from tensorflow import keras
# + [markdown] id="7b363673d96c"
# ## The `Layer` class: the combination of state (weights) and some computation
#
# One of the central abstraction in Keras is the `Layer` class. A layer
# encapsulates both a state (the layer's "weights") and a transformation from
# inputs to outputs (a "call", the layer's forward pass).
#
# Here's a densely-connected layer. It has a state: the variables `w` and `b`.
# + id="59b8317dbd3c"
class Linear(keras.layers.Layer):
def __init__(self, units=32, input_dim=32):
super(Linear, self).__init__()
w_init = tf.random_normal_initializer()
self.w = tf.Variable(
initial_value=w_init(shape=(input_dim, units), dtype="float32"),
trainable=True,
)
b_init = tf.zeros_initializer()
self.b = tf.Variable(
initial_value=b_init(shape=(units,), dtype="float32"), trainable=True
)
def call(self, inputs):
return tf.matmul(inputs, self.w) + self.b
# + [markdown] id="dac8fb03a642"
# You would use a layer by calling it on some tensor input(s), much like a Python
# function.
# + id="cdcd15d5e68a"
x = tf.ones((2, 2))
linear_layer = Linear(4, 2)
y = linear_layer(x)
print(y)
# + [markdown] id="382960020a56"
# Note that the weights `w` and `b` are automatically tracked by the layer upon
# being set as layer attributes:
# + id="d3d875af9465"
assert linear_layer.weights == [linear_layer.w, linear_layer.b]
# + [markdown] id="ec9d72aa7538"
# Note you also have access to a quicker shortcut for adding weight to a layer:
# the `add_weight()` method:
# + id="168548eba841"
class Linear(keras.layers.Layer):
def __init__(self, units=32, input_dim=32):
super(Linear, self).__init__()
self.w = self.add_weight(
shape=(input_dim, units), initializer="random_normal", trainable=True
)
self.b = self.add_weight(shape=(units,), initializer="zeros", trainable=True)
def call(self, inputs):
return tf.matmul(inputs, self.w) + self.b
x = tf.ones((2, 2))
linear_layer = Linear(4, 2)
y = linear_layer(x)
print(y)
# + [markdown] id="070ea9b4db6c"
# ## Layers can have non-trainable weights
#
# Besides trainable weights, you can add non-trainable weights to a layer as
# well. Such weights are meant not to be taken into account during
# backpropagation, when you are training the layer.
#
# Here's how to add and use a non-trainable weight:
# + id="7c4cb404145f"
class ComputeSum(keras.layers.Layer):
def __init__(self, input_dim):
super(ComputeSum, self).__init__()
self.total = tf.Variable(initial_value=tf.zeros((input_dim,)), trainable=False)
def call(self, inputs):
self.total.assign_add(tf.reduce_sum(inputs, axis=0))
return self.total
x = tf.ones((2, 2))
my_sum = ComputeSum(2)
y = my_sum(x)
print(y.numpy())
y = my_sum(x)
print(y.numpy())
# + [markdown] id="40f5b74d3d87"
# It's part of `layer.weights`, but it gets categorized as a non-trainable weight:
# + id="3d4db4ef4fa4"
print("weights:", len(my_sum.weights))
print("non-trainable weights:", len(my_sum.non_trainable_weights))
# It's not included in the trainable weights:
print("trainable_weights:", my_sum.trainable_weights)
# + [markdown] id="fe6942aff7c6"
# ## Best practice: deferring weight creation until the shape of the inputs is known
#
# Our `Linear` layer above took an `input_dim `argument that was used to compute
# the shape of the weights `w` and `b` in `__init__()`:
# + id="275b68d5ea9f"
class Linear(keras.layers.Layer):
def __init__(self, units=32, input_dim=32):
super(Linear, self).__init__()
self.w = self.add_weight(
shape=(input_dim, units), initializer="random_normal", trainable=True
)
self.b = self.add_weight(shape=(units,), initializer="zeros", trainable=True)
def call(self, inputs):
return tf.matmul(inputs, self.w) + self.b
# + [markdown] id="5ebcacebb348"
# In many cases, you may not know in advance the size of your inputs, and you
# would like to lazily create weights when that value becomes known, some time
# after instantiating the layer.
#
# In the Keras API, we recommend creating layer weights in the `build(self,
# inputs_shape)` method of your layer. Like this:
# + id="118c899f427e"
class Linear(keras.layers.Layer):
def __init__(self, units=32):
super(Linear, self).__init__()
self.units = units
def build(self, input_shape):
self.w = self.add_weight(
shape=(input_shape[-1], self.units),
initializer="random_normal",
trainable=True,
)
self.b = self.add_weight(
shape=(self.units,), initializer="random_normal", trainable=True
)
def call(self, inputs):
return tf.matmul(inputs, self.w) + self.b
# + [markdown] id="78061e0583c6"
# The `__call__()` method of your layer will automatically run build the first time
# it is called. You now have a layer that's lazy and thus easier to use:
# + id="0697afb97bc1"
# At instantiation, we don't know on what inputs this is going to get called
linear_layer = Linear(32)
# The layer's weights are created dynamically the first time the layer is called
y = linear_layer(x)
# + [markdown] id="51b81f42b466"
# Implementing `build()` separately as shown above nicely separates creating weights
# only once from using weights in every call. However, for some advanced custom
# layers, it can become impractical to separate the state creation and computation.
# Layer implementers are allowed to defer weight creation to the first `__call__()`,
# but need to take care that later calls use the same weights. In addition, since
# `__call__()` is likely to be executed for the first time inside a `tf.function`,
# any variable creation that takes place in `__call__()` should be wrapped in a`tf.init_scope`.
# + [markdown] id="0b7a45f57610"
# ## Layers are recursively composable
#
# If you assign a Layer instance as an attribute of another Layer, the outer layer
# will start tracking the weights created by the inner layer.
#
# We recommend creating such sublayers in the `__init__()` method and leave it to
# the first `__call__()` to trigger building their weights.
# + id="1aaaf82ab8ce"
class MLPBlock(keras.layers.Layer):
def __init__(self):
super(MLPBlock, self).__init__()
self.linear_1 = Linear(32)
self.linear_2 = Linear(32)
self.linear_3 = Linear(1)
def call(self, inputs):
x = self.linear_1(inputs)
x = tf.nn.relu(x)
x = self.linear_2(x)
x = tf.nn.relu(x)
return self.linear_3(x)
mlp = MLPBlock()
y = mlp(tf.ones(shape=(3, 64))) # The first call to the `mlp` will create the weights
print("weights:", len(mlp.weights))
print("trainable weights:", len(mlp.trainable_weights))
# + [markdown] id="2bf11b296bd2"
# ## The `add_loss()` method
#
# When writing the `call()` method of a layer, you can create loss tensors that
# you will want to use later, when writing your training loop. This is doable by
# calling `self.add_loss(value)`:
# + id="ba2782dc0879"
# A layer that creates an activity regularization loss
class ActivityRegularizationLayer(keras.layers.Layer):
def __init__(self, rate=1e-2):
super(ActivityRegularizationLayer, self).__init__()
self.rate = rate
def call(self, inputs):
self.add_loss(self.rate * tf.reduce_sum(inputs))
return inputs
# + [markdown] id="a883b230a9e9"
# These losses (including those created by any inner layer) can be retrieved via
# `layer.losses`. This property is reset at the start of every `__call__()` to
# the top-level layer, so that `layer.losses` always contains the loss values
# created during the last forward pass.
# + id="b56d223a30cd"
class OuterLayer(keras.layers.Layer):
def __init__(self):
super(OuterLayer, self).__init__()
self.activity_reg = ActivityRegularizationLayer(1e-2)
def call(self, inputs):
return self.activity_reg(inputs)
layer = OuterLayer()
assert len(layer.losses) == 0 # No losses yet since the layer has never been called
_ = layer(tf.zeros(1, 1))
assert len(layer.losses) == 1 # We created one loss value
# `layer.losses` gets reset at the start of each __call__
_ = layer(tf.zeros(1, 1))
assert len(layer.losses) == 1 # This is the loss created during the call above
# + [markdown] id="0809dec680ff"
# In addition, the `loss` property also contains regularization losses created
# for the weights of any inner layer:
# + id="41016153e983"
class OuterLayerWithKernelRegularizer(keras.layers.Layer):
def __init__(self):
super(OuterLayerWithKernelRegularizer, self).__init__()
self.dense = keras.layers.Dense(
32, kernel_regularizer=tf.keras.regularizers.l2(1e-3)
)
def call(self, inputs):
return self.dense(inputs)
layer = OuterLayerWithKernelRegularizer()
_ = layer(tf.zeros((1, 1)))
# This is `1e-3 * sum(layer.dense.kernel ** 2)`,
# created by the `kernel_regularizer` above.
print(layer.losses)
# + [markdown] id="589465e06e4f"
# These losses are meant to be taken into account when writing training loops,
# like this:
#
# ```python
# # Instantiate an optimizer.
# optimizer = tf.keras.optimizers.SGD(learning_rate=1e-3)
# loss_fn = keras.losses.SparseCategoricalCrossentropy(from_logits=True)
#
# # Iterate over the batches of a dataset.
# for x_batch_train, y_batch_train in train_dataset:
# with tf.GradientTape() as tape:
# logits = layer(x_batch_train) # Logits for this minibatch
# # Loss value for this minibatch
# loss_value = loss_fn(y_batch_train, logits)
# # Add extra losses created during this forward pass:
# loss_value += sum(model.losses)
#
# grads = tape.gradient(loss_value, model.trainable_weights)
# optimizer.apply_gradients(zip(grads, model.trainable_weights))
# ```
# + [markdown] id="7fb41ca8c3b0"
# For a detailed guide about writing training loops, see the
# [guide to writing a training loop from scratch](https://www.tensorflow.org/guide/keras/writing_a_training_loop_from_scratch/).
#
# These losses also work seamlessly with `fit()` (they get automatically summed
# and added to the main loss, if any):
# + id="769bc6612ebf"
import numpy as np
inputs = keras.Input(shape=(3,))
outputs = ActivityRegularizationLayer()(inputs)
model = keras.Model(inputs, outputs)
# If there is a loss passed in `compile`, the regularization
# losses get added to it
model.compile(optimizer="adam", loss="mse")
model.fit(np.random.random((2, 3)), np.random.random((2, 3)))
# It's also possible not to pass any loss in `compile`,
# since the model already has a loss to minimize, via the `add_loss`
# call during the forward pass!
model.compile(optimizer="adam")
model.fit(np.random.random((2, 3)), np.random.random((2, 3)))
# + [markdown] id="149c71e442bb"
# ## The `add_metric()` method
#
# Similarly to `add_loss()`, layers also have an `add_metric()` method
# for tracking the moving average of a quantity during training.
#
# Consider the following layer: a "logistic endpoint" layer.
# It takes as inputs predictions & targets, it computes a loss which it tracks
# via `add_loss()`, and it computes an accuracy scalar, which it tracks via
# `add_metric()`.
# + id="bfb2df515096"
class LogisticEndpoint(keras.layers.Layer):
def __init__(self, name=None):
super(LogisticEndpoint, self).__init__(name=name)
self.loss_fn = keras.losses.BinaryCrossentropy(from_logits=True)
self.accuracy_fn = keras.metrics.BinaryAccuracy()
def call(self, targets, logits, sample_weights=None):
# Compute the training-time loss value and add it
# to the layer using `self.add_loss()`.
loss = self.loss_fn(targets, logits, sample_weights)
self.add_loss(loss)
# Log accuracy as a metric and add it
# to the layer using `self.add_metric()`.
acc = self.accuracy_fn(targets, logits, sample_weights)
self.add_metric(acc, name="accuracy")
# Return the inference-time prediction tensor (for `.predict()`).
return tf.nn.softmax(logits)
# + [markdown] id="e68f88373800"
# Metrics tracked in this way are accessible via `layer.metrics`:
# + id="1834d74450b6"
layer = LogisticEndpoint()
targets = tf.ones((2, 2))
logits = tf.ones((2, 2))
y = layer(targets, logits)
print("layer.metrics:", layer.metrics)
print("current accuracy value:", float(layer.metrics[0].result()))
# + [markdown] id="546cfbd4ea05"
# Just like for `add_loss()`, these metrics are tracked by `fit()`:
# + id="f5e74cb4da34"
inputs = keras.Input(shape=(3,), name="inputs")
targets = keras.Input(shape=(10,), name="targets")
logits = keras.layers.Dense(10)(inputs)
predictions = LogisticEndpoint(name="predictions")(logits, targets)
model = keras.Model(inputs=[inputs, targets], outputs=predictions)
model.compile(optimizer="adam")
data = {
"inputs": np.random.random((3, 3)),
"targets": np.random.random((3, 10)),
}
model.fit(data)
# + [markdown] id="4012fa8683e5"
# ## You can optionally enable serialization on your layers
#
# If you need your custom layers to be serializable as part of a
# [Functional model](https://www.tensorflow.org/guide/keras/functional/), you can optionally implement a `get_config()`
# method:
# + id="0a720cbd5f54"
class Linear(keras.layers.Layer):
def __init__(self, units=32):
super(Linear, self).__init__()
self.units = units
def build(self, input_shape):
self.w = self.add_weight(
shape=(input_shape[-1], self.units),
initializer="random_normal",
trainable=True,
)
self.b = self.add_weight(
shape=(self.units,), initializer="random_normal", trainable=True
)
def call(self, inputs):
return tf.matmul(inputs, self.w) + self.b
def get_config(self):
return {"units": self.units}
# Now you can recreate the layer from its config:
layer = Linear(64)
config = layer.get_config()
print(config)
new_layer = Linear.from_config(config)
# + [markdown] id="1b43aad6c145"
# Note that the `__init__()` method of the base `Layer` class takes some keyword
# arguments, in particular a `name` and a `dtype`. It's good practice to pass
# these arguments to the parent class in `__init__()` and to include them in the
# layer config:
# + id="0cbad8a6e6cd"
class Linear(keras.layers.Layer):
def __init__(self, units=32, **kwargs):
super(Linear, self).__init__(**kwargs)
self.units = units
def build(self, input_shape):
self.w = self.add_weight(
shape=(input_shape[-1], self.units),
initializer="random_normal",
trainable=True,
)
self.b = self.add_weight(
shape=(self.units,), initializer="random_normal", trainable=True
)
def call(self, inputs):
return tf.matmul(inputs, self.w) + self.b
def get_config(self):
config = super(Linear, self).get_config()
config.update({"units": self.units})
return config
layer = Linear(64)
config = layer.get_config()
print(config)
new_layer = Linear.from_config(config)
# + [markdown] id="2421f80b5b86"
# If you need more flexibility when deserializing the layer from its config, you
# can also override the `from_config()` class method. This is the base
# implementation of `from_config()`:
#
# ```python
# def from_config(cls, config):
# return cls(**config)
# ```
#
# To learn more about serialization and saving, see the complete
# [guide to saving and serializing models](https://www.tensorflow.org/guide/keras/save_and_serialize/).
# + [markdown] id="3d7e2304a047"
# ## Privileged `training` argument in the `call()` method
#
# Some layers, in particular the `BatchNormalization` layer and the `Dropout`
# layer, have different behaviors during training and inference. For such
# layers, it is standard practice to expose a `training` (boolean) argument in
# the `call()` method.
#
# By exposing this argument in `call()`, you enable the built-in training and
# evaluation loops (e.g. `fit()`) to correctly use the layer in training and
# inference.
# + id="a169812c2c00"
class CustomDropout(keras.layers.Layer):
def __init__(self, rate, **kwargs):
super(CustomDropout, self).__init__(**kwargs)
self.rate = rate
def call(self, inputs, training=None):
if training:
return tf.nn.dropout(inputs, rate=self.rate)
return inputs
# + [markdown] id="9e1482c9f010"
# ## Privileged `mask` argument in the `call()` method
#
# The other privileged argument supported by `call()` is the `mask` argument.
#
# You will find it in all Keras RNN layers. A mask is a boolean tensor (one
# boolean value per timestep in the input) used to skip certain input timesteps
# when processing timeseries data.
#
# Keras will automatically pass the correct `mask` argument to `__call__()` for
# layers that support it, when a mask is generated by a prior layer.
# Mask-generating layers are the `Embedding`
# layer configured with `mask_zero=True`, and the `Masking` layer.
#
# To learn more about masking and how to write masking-enabled layers, please
# check out the guide
# ["understanding padding and masking"](https://www.tensorflow.org/guide/keras/masking_and_padding/).
# + [markdown] id="344110f9e134"
# ## The `Model` class
#
# In general, you will use the `Layer` class to define inner computation blocks,
# and will use the `Model` class to define the outer model -- the object you
# will train.
#
# For instance, in a ResNet50 model, you would have several ResNet blocks
# subclassing `Layer`, and a single `Model` encompassing the entire ResNet50
# network.
#
# The `Model` class has the same API as `Layer`, with the following differences:
#
# - It exposes built-in training, evaluation, and prediction loops
# (`model.fit()`, `model.evaluate()`, `model.predict()`).
# - It exposes the list of its inner layers, via the `model.layers` property.
# - It exposes saving and serialization APIs (`save()`, `save_weights()`...)
#
# Effectively, the `Layer` class corresponds to what we refer to in the
# literature as a "layer" (as in "convolution layer" or "recurrent layer") or as
# a "block" (as in "ResNet block" or "Inception block").
#
# Meanwhile, the `Model` class corresponds to what is referred to in the
# literature as a "model" (as in "deep learning model") or as a "network" (as in
# "deep neural network").
#
# So if you're wondering, "should I use the `Layer` class or the `Model` class?",
# ask yourself: will I need to call `fit()` on it? Will I need to call `save()`
# on it? If so, go with `Model`. If not (either because your class is just a block
# in a bigger system, or because you are writing training & saving code yourself),
# use `Layer`.
#
# For instance, we could take our mini-resnet example above, and use it to build
# a `Model` that we could train with `fit()`, and that we could save with
# `save_weights()`:
# + [markdown] id="09caa642b72e"
# ```python
# class ResNet(tf.keras.Model):
#
# def __init__(self, num_classes=1000):
# super(ResNet, self).__init__()
# self.block_1 = ResNetBlock()
# self.block_2 = ResNetBlock()
# self.global_pool = layers.GlobalAveragePooling2D()
# self.classifier = Dense(num_classes)
#
# def call(self, inputs):
# x = self.block_1(inputs)
# x = self.block_2(x)
# x = self.global_pool(x)
# return self.classifier(x)
#
#
# resnet = ResNet()
# dataset = ...
# resnet.fit(dataset, epochs=10)
# resnet.save(filepath)
# ```
# + [markdown] id="a2e32d225a1b"
# ## Putting it all together: an end-to-end example
#
# Here's what you've learned so far:
#
# - A `Layer` encapsulate a state (created in `__init__()` or `build()`) and some
# computation (defined in `call()`).
# - Layers can be recursively nested to create new, bigger computation blocks.
# - Layers can create and track losses (typically regularization losses) as well
# as metrics, via `add_loss()` and `add_metric()`
# - The outer container, the thing you want to train, is a `Model`. A `Model` is
# just like a `Layer`, but with added training and serialization utilities.
#
# Let's put all of these things together into an end-to-end example: we're going
# to implement a Variational AutoEncoder (VAE). We'll train it on MNIST digits.
#
# Our VAE will be a subclass of `Model`, built as a nested composition of layers
# that subclass `Layer`. It will feature a regularization loss (KL divergence).
# + id="56aaae7af872"
from tensorflow.keras import layers
class Sampling(layers.Layer):
"""Uses (z_mean, z_log_var) to sample z, the vector encoding a digit."""
def call(self, inputs):
z_mean, z_log_var = inputs
batch = tf.shape(z_mean)[0]
dim = tf.shape(z_mean)[1]
epsilon = tf.keras.backend.random_normal(shape=(batch, dim))
return z_mean + tf.exp(0.5 * z_log_var) * epsilon
class Encoder(layers.Layer):
"""Maps MNIST digits to a triplet (z_mean, z_log_var, z)."""
def __init__(self, latent_dim=32, intermediate_dim=64, name="encoder", **kwargs):
super(Encoder, self).__init__(name=name, **kwargs)
self.dense_proj = layers.Dense(intermediate_dim, activation="relu")
self.dense_mean = layers.Dense(latent_dim)
self.dense_log_var = layers.Dense(latent_dim)
self.sampling = Sampling()
def call(self, inputs):
x = self.dense_proj(inputs)
z_mean = self.dense_mean(x)
z_log_var = self.dense_log_var(x)
z = self.sampling((z_mean, z_log_var))
return z_mean, z_log_var, z
class Decoder(layers.Layer):
"""Converts z, the encoded digit vector, back into a readable digit."""
def __init__(self, original_dim, intermediate_dim=64, name="decoder", **kwargs):
super(Decoder, self).__init__(name=name, **kwargs)
self.dense_proj = layers.Dense(intermediate_dim, activation="relu")
self.dense_output = layers.Dense(original_dim, activation="sigmoid")
def call(self, inputs):
x = self.dense_proj(inputs)
return self.dense_output(x)
class VariationalAutoEncoder(keras.Model):
"""Combines the encoder and decoder into an end-to-end model for training."""
def __init__(
self,
original_dim,
intermediate_dim=64,
latent_dim=32,
name="autoencoder",
**kwargs
):
super(VariationalAutoEncoder, self).__init__(name=name, **kwargs)
self.original_dim = original_dim
self.encoder = Encoder(latent_dim=latent_dim, intermediate_dim=intermediate_dim)
self.decoder = Decoder(original_dim, intermediate_dim=intermediate_dim)
def call(self, inputs):
z_mean, z_log_var, z = self.encoder(inputs)
reconstructed = self.decoder(z)
# Add KL divergence regularization loss.
kl_loss = -0.5 * tf.reduce_mean(
z_log_var - tf.square(z_mean) - tf.exp(z_log_var) + 1
)
self.add_loss(kl_loss)
return reconstructed
# + [markdown] id="2f8ae035a7c9"
# Let's write a simple training loop on MNIST:
# + id="40f11d1ef3bc"
original_dim = 784
vae = VariationalAutoEncoder(original_dim, 64, 32)
optimizer = tf.keras.optimizers.Adam(learning_rate=1e-3)
mse_loss_fn = tf.keras.losses.MeanSquaredError()
loss_metric = tf.keras.metrics.Mean()
(x_train, _), _ = tf.keras.datasets.mnist.load_data()
x_train = x_train.reshape(60000, 784).astype("float32") / 255
train_dataset = tf.data.Dataset.from_tensor_slices(x_train)
train_dataset = train_dataset.shuffle(buffer_size=1024).batch(64)
epochs = 2
# Iterate over epochs.
for epoch in range(epochs):
print("Start of epoch %d" % (epoch,))
# Iterate over the batches of the dataset.
for step, x_batch_train in enumerate(train_dataset):
with tf.GradientTape() as tape:
reconstructed = vae(x_batch_train)
# Compute reconstruction loss
loss = mse_loss_fn(x_batch_train, reconstructed)
loss += sum(vae.losses) # Add KLD regularization loss
grads = tape.gradient(loss, vae.trainable_weights)
optimizer.apply_gradients(zip(grads, vae.trainable_weights))
loss_metric(loss)
if step % 100 == 0:
print("step %d: mean loss = %.4f" % (step, loss_metric.result()))
# + [markdown] id="f0d65fae5d3d"
# Note that since the VAE is subclassing `Model`, it features built-in training
# loops. So you could also have trained it like this:
# + id="5af13f70d528"
vae = VariationalAutoEncoder(784, 64, 32)
optimizer = tf.keras.optimizers.Adam(learning_rate=1e-3)
vae.compile(optimizer, loss=tf.keras.losses.MeanSquaredError())
vae.fit(x_train, x_train, epochs=2, batch_size=64)
# + [markdown] id="d34b7ba21662"
# ## Beyond object-oriented development: the Functional API
#
# Was this example too much object-oriented development for you? You can also
# build models using the [Functional API](https://www.tensorflow.org/guide/keras/functional/). Importantly,
# choosing one style or another does not prevent you from leveraging components
# written in the other style: you can always mix-and-match.
#
# For instance, the Functional API example below reuses the same `Sampling` layer
# we defined in the example above:
# + id="be77fc8f9b26"
original_dim = 784
intermediate_dim = 64
latent_dim = 32
# Define encoder model.
original_inputs = tf.keras.Input(shape=(original_dim,), name="encoder_input")
x = layers.Dense(intermediate_dim, activation="relu")(original_inputs)
z_mean = layers.Dense(latent_dim, name="z_mean")(x)
z_log_var = layers.Dense(latent_dim, name="z_log_var")(x)
z = Sampling()((z_mean, z_log_var))
encoder = tf.keras.Model(inputs=original_inputs, outputs=z, name="encoder")
# Define decoder model.
latent_inputs = tf.keras.Input(shape=(latent_dim,), name="z_sampling")
x = layers.Dense(intermediate_dim, activation="relu")(latent_inputs)
outputs = layers.Dense(original_dim, activation="sigmoid")(x)
decoder = tf.keras.Model(inputs=latent_inputs, outputs=outputs, name="decoder")
# Define VAE model.
outputs = decoder(z)
vae = tf.keras.Model(inputs=original_inputs, outputs=outputs, name="vae")
# Add KL divergence regularization loss.
kl_loss = -0.5 * tf.reduce_mean(z_log_var - tf.square(z_mean) - tf.exp(z_log_var) + 1)
vae.add_loss(kl_loss)
# Train.
optimizer = tf.keras.optimizers.Adam(learning_rate=1e-3)
vae.compile(optimizer, loss=tf.keras.losses.MeanSquaredError())
vae.fit(x_train, x_train, epochs=3, batch_size=64)
# + [markdown] id="e2f135ea7cf5"
# For more information, make sure to read the [Functional API guide](https://www.tensorflow.org/guide/keras/functional/).
| site/en-snapshot/guide/keras/custom_layers_and_models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# !conda install pytorch==1.9.1 -c pytorch
# !conda install pandas==1.3.3
# !conda install numpy==1.20.3
# +
'''
在虚拟环境python_torch中,启动jupyter notebook;
并运行如下代码,检查各个程序库的版本。
'''
import torch
#查看PyTorch的版本号。
torch.__version__
# +
import numpy as np
np.__version__
# +
import pandas as pd
pd.__version__
| Chapter_6/.ipynb_checkpoints/Section_6.1.1-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # ModelFlow, a toolkit
# Python is an incredible and versatile language embedded a powerful ecosystem. For data
# science
# the Pandas library is a powerful "Swiss Army Knife".
#
# In economic and for modeling banks we need **lagged variables**
# and **simultaneous formulas** (circular references in Excel speak).
#
# ModelFlow, a toolkit to enable lagged variables and simultaneous formulas.
#
# This notebook ModelFlow to extend dataframes. Other notebooks show ModelFlow as a class.
# + [markdown] slideshow={"slide_type": "slide"}
# # Jupyter
#
# This is a Jupyter notebook. Jupyter is a Python Shell
#
# You will notice **input cells** (marked:In\[\]) and **output cells** (marked: Out\[\])
#
# It is live, so you can try it out yourself, if you have access to the
# ModelFlow toolkit, else you just have to watch.
#
# This Jupyter notebook show how ModelFlow can extend pandas dataframes to run models.
#
# The notebook focus on a simple example and do not explore all the features and
# options. Also the models are toy models created to be small but still illustrative.
# + [markdown] slideshow={"slide_type": "slide"}
# # Import stuff
# + slideshow={"slide_type": "-"}
import pandas as pd # Python data science library
import sys
from IPython.display import SVG, display
sys.path.append('modelflow/')
import modelmf # This will extend pandas dataframes with ModelFlow
# + [markdown] slideshow={"slide_type": "slide"}
# # Create a Pandas Dataframe
# We make up some data.
#
# Pandas dataframes are tables with **row** and **column** names. Columns are variables, and rows are the time dimension.
# + slideshow={"slide_type": "-"}
df = pd.DataFrame({'LOAN': [100,100,100,100],'SECURITIES': [10,11,12,13],
'CASH': [4,4,4,4], 'DEPOSIT' : [100,100,100,100],
'BONDS':[1,2,3,10], 'NEW_LOAN' : [1,20,30,40] },
index=[2018,2019,2020,2021])
df
# + [markdown] slideshow={"slide_type": "slide"}
# # A model, where Pandas don't work out of the box
# A very small stylized dynamic model of the balance sheet of a bank is created.
# + slideshow={"slide_type": "-"}
fmodel = '''\
£ Stock
ASSETS = LOAN + SECURITIES + CASH
FUNDING = DEPOSIT + BONDS
EQUITY = ASSETS - FUNDING
LIABILITIES = FUNDING + EQUITY
£ stock flow
DEPOSIT = DEPOSIT(-1) + NEW_DEPOSIT
LOAN = LOAN(-1)+ NEW_LOAN
NEW_BONDS = (NEW_LOAN - NEW_DEPOSIT)
BONDS = BONDS(-1) + NEW_BONDS'''
# + [markdown] slideshow={"slide_type": "slide"}
# # Apply the model on the dataframe.
#
# To do this we use dataframe.mfcalc.
# + slideshow={"slide_type": "-"}
df.mfcalc(fmodel)
# + [markdown] slideshow={"slide_type": "slide"}
# # Notice:
# * The model is run from 2019. It cant run 2018 as as there is no values for laggged variables in 2018.
# * The model is calculated even when the formulas where not in the logical order.
# * Variables in the model missing from the dataframe are set to 0
# + [markdown] slideshow={"slide_type": "slide"}
# # There is more
# The result from a model run can be used straight in python programs.
#
# But, A model instance ```<dataframe>.mf``` contains
#
# * The first and last solution of the model
# * The directed graph of which variable contributes to which variable
# * All formulas in the model
#
# This makes it a powerful tool for model and result analysis.
# + [markdown] slideshow={"slide_type": "slide"}
# # Make another experiment
# First we update some exogenous variables (variables which are only on the right hand side of the model). Then we run the model again.
# -
df['NEW_LOAN']= [1,40,50,80]
df['NEW_DEPOSIT']= [1,30,25,50]
df.mfcalc(fmodel)
# + [markdown] slideshow={"slide_type": "slide"}
# # Visualizing
# The results can be compared and visualized.
#
# Wildcards can be used to select the variables to visualize.
#
# If this is not sufficient the whole suite of Python visualization (as Matplotlib, Seaborn, Plotly) can be used on top of the resulting dataframes.
# + [markdown] slideshow={"slide_type": "slide"}
# # Plot the last result
# -
_ = df.mf['*'].plot()
# + [markdown] slideshow={"slide_type": "slide"}
# # Plot the difference between the first and last run
# -
_ = df.mf['*'].dif.plot()
# + [markdown] slideshow={"slide_type": "slide"}
# # Or as heatmap
# -
_ = df.mf[['*']].dif.heat(title='All',annot=True)
# + [markdown] slideshow={"slide_type": "slide"}
# # The stucture of the model (dependency graph)
# -
df.mf.drawmodel()
# + slideshow={"slide_type": "subslide"}
df.mf.drawmodel(all =1)
# + [markdown] slideshow={"slide_type": "slide"}
# # What explains the difference for a variable
# Which of the input variables explains the difference of the results of a formula between two runs.
# + [markdown] slideshow={"slide_type": "subslide"}
# If we have:
#
# $y = f(a,b)$
#
# and we have two solutions where the variables differs by $\Delta y, \Delta a, \Delta b$
#
# How much of $\Delta y$ can be explained by $\Delta a$ and $\Delta b$ ?
#
# Analytical the attributions $\Omega a$ and $\Omega b$ can be calculated like this:
#
# $\Delta y = \underbrace{\Delta a \frac{\partial {f}}{\partial{a}}(a,b)}_{\Omega a} +
# \underbrace{\Delta b \frac{\partial {f}}{\partial{b}}(a,b)}_{\Omega b}+Residual$
# + [markdown] slideshow={"slide_type": "subslide"}
# If we have two experiments:
#
# \begin{eqnarray}
# y_0&=&𝑓(a_{0},b_{0}) \\
# y_1&=&𝑓(a_0+\Delta a,b_{0}+ \Delta b)
# \end{eqnarray}
#
# ModelFlow will do a numerical approximation of $\Omega a$ and $\Omega b$.
#
# \begin{eqnarray}
# \Omega f_a&=&f(a_1,b_1 )-f(a_1-\Delta a,b_1) \\
# \Omega f_b&=&f(a_1,b_1 )-f(a_1,b_1-\Delta b)
# \end{eqnarray}
#
#
#
# If the model is fairly linear, the residual will be small.
#
# \begin{eqnarray}
# residual = \Omega f_a + \Omega f_b -(y_1 - y_0)
# \end{eqnarray}
# -
# Now look at generations of attributions
# + slideshow={"slide_type": "slide"}
_= df.mf.bonds.explain(up=2,HR=0,pdf=0)
# + [markdown] slideshow={"slide_type": "slide"}
# # Beyond a simpel model
#
# This was a simple model. You don't even need a computer to solve it, or you could use Excel to handle the model. However sometime you want to:
#
# * Invert the model to solve for targets as function of instruments.
# * Use a a more rich business logic language
# * Let variables be matrices (dense or sparse)
# * Scale to large model (even millions of equations)
# * Handle simultaneous models
# * Make identical models for a lot of banks/sectors or whatever
# * Solve models fast
# * Grab a model from Matlab/Dynare/Excel/Latex
# * Attribute difference between scenarios to individual variables or groups of variables.
# * Speed up the model by compiling the model
# * Linearize the model using symbolic differentiation
# * Calculate stability of a linearized model.
| ModelFlow, extend DataFrame.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# Customizing Matplotlib with style sheets and rcParams
# =====================================================
#
# Tips for customizing the properties and default styles of Matplotlib.
#
# Using style sheets
# ------------------
#
# The ``style`` package adds support for easy-to-switch plotting "styles" with
# the same parameters as a
# `matplotlib rc <customizing-with-matplotlibrc-files>` file (which is read
# at startup to configure matplotlib).
#
# There are a number of pre-defined styles `provided by Matplotlib`_. For
# example, there's a pre-defined style called "ggplot", which emulates the
# aesthetics of ggplot_ (a popular plotting package for R_). To use this style,
# just add:
#
import numpy as np
import matplotlib.pyplot as plt
import matplotlib as mpl
from cycler import cycler
plt.style.use('ggplot')
data = np.random.randn(50)
# To list all available styles, use:
#
#
print(plt.style.available)
# Defining your own style
# -----------------------
#
# You can create custom styles and use them by calling ``style.use`` with the
# path or URL to the style sheet. Additionally, if you add your
# ``<style-name>.mplstyle`` file to ``mpl_configdir/stylelib``, you can reuse
# your custom style sheet with a call to ``style.use(<style-name>)``. By default
# ``mpl_configdir`` should be ``~/.config/matplotlib``, but you can check where
# yours is with ``matplotlib.get_configdir()``; you may need to create this
# directory. You also can change the directory where matplotlib looks for
# the stylelib/ folder by setting the MPLCONFIGDIR environment variable,
# see `locating-matplotlib-config-dir`.
#
# Note that a custom style sheet in ``mpl_configdir/stylelib`` will
# override a style sheet defined by matplotlib if the styles have the same name.
#
# For example, you might want to create
# ``mpl_configdir/stylelib/presentation.mplstyle`` with the following::
#
# axes.titlesize : 24
# axes.labelsize : 20
# lines.linewidth : 3
# lines.markersize : 10
# xtick.labelsize : 16
# ytick.labelsize : 16
#
# Then, when you want to adapt a plot designed for a paper to one that looks
# good in a presentation, you can just add::
#
# >>> import matplotlib.pyplot as plt
# >>> plt.style.use('presentation')
#
#
# Composing styles
# ----------------
#
# Style sheets are designed to be composed together. So you can have a style
# sheet that customizes colors and a separate style sheet that alters element
# sizes for presentations. These styles can easily be combined by passing
# a list of styles::
#
# >>> import matplotlib.pyplot as plt
# >>> plt.style.use(['dark_background', 'presentation'])
#
# Note that styles further to the right will overwrite values that are already
# defined by styles on the left.
#
#
# Temporary styling
# -----------------
#
# If you only want to use a style for a specific block of code but don't want
# to change the global styling, the style package provides a context manager
# for limiting your changes to a specific scope. To isolate your styling
# changes, you can write something like the following:
#
#
with plt.style.context('dark_background'):
plt.plot(np.sin(np.linspace(0, 2 * np.pi)), 'r-o')
plt.show()
#
# matplotlib rcParams
# ===================
#
#
# Dynamic rc settings
# -------------------
#
# You can also dynamically change the default rc settings in a python script or
# interactively from the python shell. All of the rc settings are stored in a
# dictionary-like variable called :data:`matplotlib.rcParams`, which is global to
# the matplotlib package. rcParams can be modified directly, for example:
#
#
mpl.rcParams['lines.linewidth'] = 2
mpl.rcParams['lines.linestyle'] = '--'
plt.plot(data)
# Note, that in order to change the usual `plot` color you have to change the
# *prop_cycle* property of *axes*:
#
#
mpl.rcParams['axes.prop_cycle'] = cycler(color=['r', 'g', 'b', 'y'])
plt.plot(data) # first color is red
# Matplotlib also provides a couple of convenience functions for modifying rc
# settings. The :func:`matplotlib.rc` command can be used to modify multiple
# settings in a single group at once, using keyword arguments:
#
#
mpl.rc('lines', linewidth=4, linestyle='-.')
plt.plot(data)
# The :func:`matplotlib.rcdefaults` command will restore the standard matplotlib
# default settings.
#
# There is some degree of validation when setting the values of rcParams, see
# :mod:`matplotlib.rcsetup` for details.
#
#
# The :file:`matplotlibrc` file
# -----------------------------
#
# Matplotlib uses :file:`matplotlibrc` configuration files to customize all
# kinds of properties, which we call 'rc settings' or 'rc parameters'. You can
# control the defaults of almost every property in Matplotlib: figure size and
# DPI, line width, color and style, axes, axis and grid properties, text and
# font properties and so on. Matplotlib looks for :file:`matplotlibrc` in four
# locations, in the following order:
#
# 1. :file:`matplotlibrc` in the current working directory, usually used for
# specific customizations that you do not want to apply elsewhere.
#
# 2. :file:`$MATPLOTLIBRC` if it is a file, else :file:`$MATPLOTLIBRC/matplotlibrc`.
#
# 3. It next looks in a user-specific place, depending on your platform:
#
# - On Linux and FreeBSD, it looks in
# :file:`.config/matplotlib/matplotlibrc` (or
# :file:`$XDG_CONFIG_HOME/matplotlib/matplotlibrc`) if you've customized
# your environment.
#
# - On other platforms, it looks in :file:`.matplotlib/matplotlibrc`.
#
# See `locating-matplotlib-config-dir`.
#
# 4. :file:`{INSTALL}/matplotlib/mpl-data/matplotlibrc`, where
# :file:`{INSTALL}` is something like
# :file:`/usr/lib/python3.7/site-packages` on Linux, and maybe
# :file:`C:\\Python37\\Lib\\site-packages` on Windows. Every time you
# install matplotlib, this file will be overwritten, so if you want
# your customizations to be saved, please move this file to your
# user-specific matplotlib directory.
#
# Once a :file:`matplotlibrc` file has been found, it will *not* search any of
# the other paths.
#
# To display where the currently active :file:`matplotlibrc` file was
# loaded from, one can do the following::
#
# >>> import matplotlib
# >>> matplotlib.matplotlib_fname()
# '/home/foo/.config/matplotlib/matplotlibrc'
#
# See below for a sample `matplotlibrc file<matplotlibrc-sample>`.
#
#
# A sample matplotlibrc file
# ~~~~~~~~~~~~~~~~~~~~~~~~~~
#
# .. literalinclude:: ../../../matplotlibrc.template
#
#
#
#
| matplotlib/tutorials_jupyter/introductory/customizing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="bOTLzSgyZcNi"
# # Library Import and Environment Setting
# + id="PvchdBUeZX6i" executionInfo={"status": "ok", "timestamp": 1641579689127, "user_tz": -540, "elapsed": 261, "user": {"displayName": "\uae40\ubcc4\ud76c", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06602448826503759202"}}
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
import matplotlib.cm as cm
plt.style.use('seaborn')
mpl.rcParams['xtick.labelsize'] = 10
mpl.rcParams['ytick.labelsize'] = 10
mpl.rcParams['axes.labelsize'] = 20
mpl.rcParams['axes.titlesize'] = 20
mpl.rcParams['legend.fontsize'] = 20
# + id="5gXR95a1ZwAk" executionInfo={"status": "ok", "timestamp": 1641576238029, "user_tz": -540, "elapsed": 4, "user": {"displayName": "\uae40\ubcc4\ud76c", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06602448826503759202"}}
# basic_nodes
class plus_node:
def __init__(self):
self._x, self._y = None, None
self._z = None
def forward(self,x,y):
self._x, self._y = x, y
self._z = self._x + self._y
return self._z
def backward(self, dz): # dz = dJ/dz
return dz*1,dz*1 #dJ/dx= dJ/dz * dz/dx, dJ/dy
class minus_node:
def __init__(self):
self._x, self._y = None, None
self._z = None
def forward(self, x, y):
self._x, self._y = x, y
self._z = self._x - self._y
return self._z
def backward(self, dz):
return 1*dz, -1*dz
class mul_node:
def __init__(self):
self._x, self._y = None, None
self._z = None
def forward(self, x, y):
self._x , self._y = x, y
self._z = self._x * self._y
return self._z
def backward(self, dz):
return dz*self._y, dz*self._x # dJ/dz * dz/dx, dJ/dy
class square_node:
def __init__(self):
self._x = None
self._z = None
def forward(self, x):
self._x = x
self._z = self._x * self._x
return self._z
def backward(self, dz):
return dz*(2*self._x)
class mean_node:
def __init__(self):
self._x = None
self._z = None
def forward(self, x):
self._x = x
self._z = np.mean(self._x) # z = 1/n *(x1+x2+...+xn)
return self._z
def backward(self, dz):
dx = dz*1/len(self._x)*np.ones_like(self._x) # dJ/dx1 = dJ/dz * dz/dx1 = dJ/dz * (1/n)
return dx
# + [markdown] id="UBD_FfpPlps8"
# ### Mini-batch Preparation
# + id="3y98ApBQkV-5" executionInfo={"status": "ok", "timestamp": 1641579462831, "user_tz": -540, "elapsed": 247, "user": {"displayName": "\uae40\ubcc4\ud76c", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06602448826503759202"}}
def dataset_generator(x_dict):
x_data = np.random.normal(x_dict['mean'], x_dict['std'], x_dict['n_sample'])
x_data_noise = x_data + x_dict['noise_factor']*np.random.normal(0,1, x_dict['n_sample'])
if x_dict['direction'] > 0:
y_data = (x_data_noise > x_dict['cutoff']).astype(np.int)
else:
y_data = (x_data_noise < x_dict['cutoff']).astype(np.int)
data = np.zeros(shape=(x_dict['n_sample'],1))
data = np.hstack((data, x_data.reshape(-1,1), y_data.reshape(-1,1)))
return data
def get_data_batch(data, batch_idx):
global n_batch, batch_size
if batch_idx is n_batch -1:
batch = data[batch_idx*batch_size:]
else:
batch = data[batch_idx*batch_size : (batch_idx+1)*batch_size]
return batch
# + [markdown] id="El1x38e_mhVF"
# # Affine Module
# + id="DjmW-3INmDmj" executionInfo={"status": "ok", "timestamp": 1641579623630, "user_tz": -540, "elapsed": 305, "user": {"displayName": "\uae40\ubcc4\ud76c", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06602448826503759202"}}
class Affine:
def __init__(self):
self._feature_dim = 1
self._Th = None
self.node_imp()
self.random_initialization()
def node_imp(self):
self._node1 = mul_node()
self._node2 = plus_node()
def random_initialization(self):
r_feature_dim = 1/self._feature_dim
self._Th = np.random.uniform(low = -1*r_feature_dim,
high = r_feature_dim,
size = (self._feature_dim+1, 1))
def forward(self, X):
self._Z1 = self._node1.forward(self._Th[1], X)
self._Z2 = self._node2.forward(self._Th[0], self._Z1)
return self._Z2
def backward(self, dZ, lr):
dTh0, dZ1 = self._node2.backward(dZ)
dTh1, dX = self._node1.backward(dZ1)
self._Th[1] = self._Th[1] - lr*np.sum(dTh1)
self._Th[0] = self._Th[0] - lr*np.sum(dTh0)
def get_Th(self):
return self._Th
# + [markdown] id="peWJUYgjpA0b"
# # Sigmoid Moudle
# + id="T_NBOMKUmivF" executionInfo={"status": "ok", "timestamp": 1641577701671, "user_tz": -540, "elapsed": 276, "user": {"displayName": "\uae40\ubcc4\ud76c", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06602448826503759202"}}
class Sigmoid:
def __init__(self):
self._Pred = None
def forward(self, Z):
self._Pred = 1/(1+np.exp(-1*Z))
return self._Pred
def backward(self, dPred):
Partial = self._Pred * (1- self._Pred)
dZ = dPred * Partial
return dZ
# + [markdown] id="HRgY_t7EqAa2"
# # Binary Cross Entropy Module
# + id="xtaxcmdqmjF9" executionInfo={"status": "ok", "timestamp": 1641578059552, "user_tz": -540, "elapsed": 271, "user": {"displayName": "\uae40\ubcc4\ud76c", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06602448826503759202"}}
class BinaryCrossEntropy_Cost:
def __init__(self):
self._Y, self._Pred = None, None
self._mean_node = mean_node()
def forward(self, Y, Pred):
self._Y, self._Pred = Y, Pred
Loss = -1*(Y*np.log(self._Pred) + (1-Y)*np.log(1-self._Pred))
J = self._mean_node.forward(Loss)
def backward(self):
dLoss = self._mean_node.backward(1)
dPred = dLoss * (self._Pred - self._Y)/(self._Pred*(1-self._Pred))
return dPred
# + [markdown] id="VtZ7_E3vqGpC"
# # SVLoR Module
# + id="-OjObZnNqGI0" executionInfo={"status": "ok", "timestamp": 1641578862239, "user_tz": -540, "elapsed": 252, "user": {"displayName": "\uae40\ubcc4\ud76c", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06602448826503759202"}}
class SVLoR:
def __init__(self):
self._feature_dim = 1
self._affine = Affine()
self._sigmoid = Sigmoid()
def forward(self, X):
Z = self._affine.forward(X)
Pred = self._sigmoid.forward(Z)
return Pred
def backward(self, dPred, lr):
dZ = self._sigmoid.backward(dPred)
self._affine.backward(dZ, lr)
def get_Th(self):
return self._affine.get_Th()
# + [markdown] id="dIt6lmNHvd76"
# # Utility Function
# + id="Iv-lPGLGvhbn" executionInfo={"status": "ok", "timestamp": 1641579658724, "user_tz": -540, "elapsed": 285, "user": {"displayName": "\uae40\ubcc4\ud76c", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06602448826503759202"}}
def result_tracker():
global iter_idx, check_freq
global th_accum, model
if iter_idx % check_freq == 0:
th_accum = np.hstack((th_accum, model.get_Th()))
cost_list.append(J)
iter_idx += 1
def result_visualizer():
global th_accum, cost_list
fig,ax = plt.subplots(2, 1, figsize = (20,5))
fig.subplots_adjust(hspace = 0.3)
ax[0].set_title(r'$\vec{\theta}$' + 'Update ')
ax[0].plot(th_accum[1,:], label = r'$\theta_{1}$')
ax[0].plot(th_accum[0,:], label = r'$\theta_{0}$')
ax[0].legend()
iter_ticks = np.linspace(0,th_accum.shape[1],10).astype(np.int)
ax[0].set_xticks(iter_ticks)
ax[1].set_title('Cost')
ax[1].plot(cost_list)
ax[1].set_xticks(iter_ticks)
n_pred = 1000
fig,ax = plt.subplots(figsize = (20,5))
ax.set_title('Predictor Update')
ax.scatter(data[:,1], data[:,-1])
ax_idx_arr = np.linspace(0,len(cost_list)-1,n_pred).astype(np.int)
cmap = cm.get_cmap('rainbow',lut = len(ax_idx_arr))
x_pred = np.linspace(np.min(data[:,1]),np.max(data[:,1]),1000)
for ax_cnt, ax_idx in enumerate(ax_idx_arr):
z = th_accum[1, ax_idx] * x_pred + th_accum[0,ax_idx]
a = 1/(1 + np.exp(-1 * z))
ax.plot(x_pred, a, color = cmap(ax_cnt),alpha = 0.2)
y_ticks = np.round(np.linspace(0, 1, 7),2)
ax.set_yticks(y_ticks)
# + [markdown] id="BgO_roZ4uszY"
# # Learning
# + colab={"base_uri": "https://localhost:8080/", "height": 444} id="m0vM9VynmQPK" executionInfo={"status": "ok", "timestamp": 1641579696962, "user_tz": -540, "elapsed": 4081, "user": {"displayName": "\uae40\ubcc4\ud76c", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06602448826503759202"}} outputId="cd3350e0-ae65-4ded-93be-71ffd99c9c6a"
# 데이터 생성
x_dict = {'mean':0, 'std':1, 'n_sample':100,
'noise_factor':0.5,
'cutoff':0, 'direction':1}
data = dataset_generator(x_dict)
# batch size 선언
batch_size = 8
n_batch = np.ceil(data.shape[0]/batch_size).astype(int)
# 모델 선언
model = SVLoR()
BCE_cost = BinaryCrossEntropy_Cost()
# parameter setting
th_accum = model.get_Th()
cost_list = []
epochs, lr = 500, 0.05
iter_idx, check_freq = 0, 5
# training
for epoch in range(epochs):
np.random.shuffle(data)
for batch_idx in range(n_batch):
batch_data = get_data_batch(data, batch_idx)
X, Y = batch_data[:,1], batch_data[:,-1]
# forwardpropagation
Pred = model.forward(X)
J = BCE_cost.forward(Y, Pred)
# backpropagation
dPred = BCE_cost.backward()
model.backward(dPred, lr)
result_tracker()
result_visualizer()
# + [markdown] id="yZJnn7bkyXfi"
# batch size를 크게 할수록 부드럽게 학습한다. <br/>
# Noise에 대한 Loss들도 평균적으로 반영하기 때문
# + [markdown] id="mvIJz0Ppxo2U"
# # Analysis
# + [markdown] id="2eCgMuKMxu6h"
# ### mean을 증가
# + colab={"base_uri": "https://localhost:8080/", "height": 444} id="MXNj5yNuwehH" executionInfo={"status": "ok", "timestamp": 1641579770181, "user_tz": -540, "elapsed": 4284, "user": {"displayName": "\uae40\ubcc4\ud76c", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06602448826503759202"}} outputId="7181a1d7-a962-469f-9477-6fd52838e482"
x_dict = {'mean':10, 'std':1, 'n_sample':100,
'noise_factor':0.5,
'cutoff':10, 'direction':1}
data = dataset_generator(x_dict)
batch_size = 8
n_batch = np.ceil(data.shape[0]/batch_size).astype(int)
model = SVLoR()
BCE_Cost = BinaryCrossEntropy_Cost()
th_accum = model.get_Th()
cost_list = []
epochs, lr = 500, 0.05
iter_idx, check_freq = 0, 5
for epoch in range(epochs):
np.random.shuffle(data)
for batch_idx in range(n_batch):
batch = get_data_batch(data, batch_idx)
X,Y = batch[:,1], batch[:,-1]
Pred = model.forward(X)
J = BCE_Cost.forward(Y, Pred)
dPred = BCE_Cost.backward()
model.backward(dPred, lr)
result_tracker()
result_visualizer()
# + [markdown] id="vxoQGmzJx5Rp"
# mean의 값을 증가시켜서 학습하게 되면
# x2와 x1이 불균형하게 학습
# + [markdown] id="5VbZ9XSSyBfP"
# ### std를 증가
# + colab={"base_uri": "https://localhost:8080/", "height": 574} id="KJUw2FxRx18I" executionInfo={"status": "ok", "timestamp": 1641579975882, "user_tz": -540, "elapsed": 3864, "user": {"displayName": "\uae40\ubcc4\ud76c", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06602448826503759202"}} outputId="98c01cec-56b8-479a-a127-81beac117faf"
x_dict = {'mean':1, 'std':7, 'n_sample':100,
'noise_factor':0.5,
'cutoff':1, 'direction':1}
data = dataset_generator(x_dict)
batch_size = 8
n_batch = np.ceil(data.shape[0]/batch_size).astype(int)
model = SVLoR()
BCE_Cost = BinaryCrossEntropy_Cost()
th_accum = model.get_Th()
cost_list = []
epochs, lr = 500, 0.005
iter_idx, check_freq = 0, 5
for epoch in range(epochs):
np.random.shuffle(data)
for batch_idx in range(n_batch):
batch = get_data_batch(data, batch_idx)
X,Y = batch[:,1], batch[:,-1]
Pred = model.forward(X)
J = BCE_Cost.forward(Y, Pred)
dPred = BCE_Cost.backward()
model.backward(dPred, lr)
result_tracker()
result_visualizer()
| Maths for Deep Learning/10_Single-variate Logistic Regression_for Several Sample.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/utkarsh512/Ad-hominem-fallacies/blob/master/unittest.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="_uPkLIlOsBRT"
# # Reproducing results of the paper _Before Name-calling: Dynamics and Triggers of Ad Hominem Fallacies in Web Argumentation_
# There are three models which are used in the paper:
# * `CNN` and `Stacked Bi-LSTM` model for classification of comments without context
# * `SSASE` model for classification with context
# + colab={"base_uri": "https://localhost:8080/"} id="vuvQ2wQYou73" outputId="ffcdd5eb-2f60-4b61-9433-5e7fb83bb01c"
# mounting Google Drive
from google.colab import drive
drive.mount('/content/gdrive')
# + colab={"base_uri": "https://localhost:8080/"} id="_q4VXdX1o5p7" outputId="e89ac442-5e11-49d4-bb9d-186afae55e2d"
# %%shell
# cd /content/gdrive/'My Drive'/
# rm -rf Ad-hominem-fallacies
git clone https://github.com/utkarsh512/Ad-hominem-fallacies.git
# + colab={"base_uri": "https://localhost:8080/"} id="8cH8F7I0pINZ" outputId="e666f55b-843b-4c67-fce6-4aca02fa02b8"
# %%shell
# cd /content/gdrive/'My Drive'/Ad-hominem-fallacies/experiments
pip install virtualenv
virtualenv env --python=python3
source env/bin/activate
pip install lda scipy==1.1.0 nltk==3.2.5
# + colab={"base_uri": "https://localhost:8080/"} id="ZoaT5qxTpQ-6" outputId="d5e8fa93-d894-4690-93ff-af6cd47a29c4"
# %%shell
# cd /content/gdrive/'My Drive'/Ad-hominem-fallacies/experiments
wget https://public.ukp.informatik.tu-darmstadt.de/ih/RedditChangeMyView2017/en-top100k.embeddings.pkl.gz
# + colab={"base_uri": "https://localhost:8080/"} id="yZfs6W5spwPw" outputId="e88b40bc-88a7-4030-9355-1667aabc2dc4"
# %%shell
# cd /content/gdrive/'My Drive'/Ad-hominem-fallacies/experiments
tar -xvf sampled-threads-ah-delta-context3.tar.bz2 -C data/
# + colab={"base_uri": "https://localhost:8080/"} id="8QrHlig_p4oh" outputId="923c6a7a-2069-43c9-f9d1-1dbf2e0d8a09"
# %%shell
# cd /content/gdrive/'My Drive'/Ad-hominem-fallacies/experiments
pip install lda
python classification_experiments.py --model cnn
# + colab={"base_uri": "https://localhost:8080/"} id="FzQ_M8vM3aen" outputId="e3a5aee1-6ecd-49f6-ffe9-74c97bd7768f"
# %%shell
# cd /content/gdrive/'My Drive'/Ad-hominem-fallacies/experiments
pip install lda
python classification_experiments.py --model bilstm
# + id="GahLy-de8W9c" colab={"base_uri": "https://localhost:8080/"} outputId="549aed1f-5837-47fe-c80f-107bbf476765"
# %%shell
# cd /content/gdrive/'My Drive'/Ad-hominem-fallacies/experiments
pip install lda
python classification_experiments.py --model ssase
# + id="UfGbpEH262w2"
| unittest.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a href="https://www.kaggle.com/satyabrataroy/simple-house-price-prediction-model-30-days-ml?scriptVersionId=88449377" target="_blank"><img align="left" alt="Kaggle" title="Open in Kaggle" src="https://kaggle.com/static/images/open-in-kaggle.svg"></a>
# + [markdown] papermill={"duration": 0.041266, "end_time": "2022-02-22T07:23:25.806191", "exception": false, "start_time": "2022-02-22T07:23:25.764925", "status": "completed"} tags=[]
# # _House Price Prediction_
# + [markdown] papermill={"duration": 0.036683, "end_time": "2022-02-22T07:23:25.880205", "exception": false, "start_time": "2022-02-22T07:23:25.843522", "status": "completed"} tags=[]
# !
# + [markdown] papermill={"duration": 0.03671, "end_time": "2022-02-22T07:23:25.95376", "exception": false, "start_time": "2022-02-22T07:23:25.91705", "status": "completed"} tags=[]
# # Objectives:
#
# > In this notebook we will create the best model to **predict** the prices of residential homes in Ames, Iowa, using different **Regression Algorithms**
# + [markdown] papermill={"duration": 0.037247, "end_time": "2022-02-22T07:23:26.028876", "exception": false, "start_time": "2022-02-22T07:23:25.991629", "status": "completed"} tags=[]
# # Contents:
#
# 1. [Data Exploration](#Data-Exploration)
#
# 1.1 [Import all the necessary libraries](#Import-all-the-necessary-libraries)
#
# 1.2 [Load the Train & Test Data into Dataframe](#Load-the-Train-&-Test-Data-into-Dataframe)
#
# 1.3 [Basic information about the datset](#Basic-information-about-the-datset)
#
# 2. [Model Parameters Defining](#Model-Parameters-Defining)
#
# 3. [Check Missing Data](#Check-Missing-Data)
#
# 4. [Handle Missing Data](#Handle-Missing-Data)
#
# 5. [Model Creation](#Model-Creation)
#
# 5.1 [Random Forest Model](#Random-Forest-Model)
#
# 5.2 [Gradient Boost Model](#Gradient-Boost-Model)
#
# 5.3 [Linear Regression Model](#Linear-Regression-Model)
#
# 6. [Build the Best Model on full dataset](#Build-the-Best-Model-on-full-dataset)
#
# 7. [Save the Model](#Save-the-Model)
# + [markdown] papermill={"duration": 0.036629, "end_time": "2022-02-22T07:23:26.103233", "exception": false, "start_time": "2022-02-22T07:23:26.066604", "status": "completed"} tags=[]
# # Data Exploration
# + [markdown] papermill={"duration": 0.036471, "end_time": "2022-02-22T07:23:26.177272", "exception": false, "start_time": "2022-02-22T07:23:26.140801", "status": "completed"} tags=[]
# ## Import all the necessary libraries
# + _kg_hide-input=true papermill={"duration": 1.240708, "end_time": "2022-02-22T07:23:27.454695", "exception": false, "start_time": "2022-02-22T07:23:26.213987", "status": "completed"} tags=[]
import numpy as np
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_squared_error
from sklearn.ensemble import RandomForestRegressor, GradientBoostingRegressor
from sklearn.linear_model import LinearRegression
import warnings
warnings.filterwarnings('ignore')
warnings.filterwarnings('ignore', category=DeprecationWarning)
# + [markdown] papermill={"duration": 0.036938, "end_time": "2022-02-22T07:23:27.529029", "exception": false, "start_time": "2022-02-22T07:23:27.492091", "status": "completed"} tags=[]
# ## Load the Train & Test Data into Dataframe
# + papermill={"duration": 0.132801, "end_time": "2022-02-22T07:23:27.699903", "exception": false, "start_time": "2022-02-22T07:23:27.567102", "status": "completed"} tags=[]
train_df = pd.read_csv('../input/home-data-for-ml-course/train.csv')
test_df = pd.read_csv('../input/home-data-for-ml-course/test.csv')
# + [markdown] papermill={"duration": 0.03654, "end_time": "2022-02-22T07:23:27.773488", "exception": false, "start_time": "2022-02-22T07:23:27.736948", "status": "completed"} tags=[]
# ## Basic information about the datset
# + papermill={"duration": 0.049845, "end_time": "2022-02-22T07:23:27.860766", "exception": false, "start_time": "2022-02-22T07:23:27.810921", "status": "completed"} tags=[]
train_df.shape
# + papermill={"duration": 0.046504, "end_time": "2022-02-22T07:23:27.944998", "exception": false, "start_time": "2022-02-22T07:23:27.898494", "status": "completed"} tags=[]
train_df.columns
# + papermill={"duration": 0.046782, "end_time": "2022-02-22T07:23:28.029773", "exception": false, "start_time": "2022-02-22T07:23:27.982991", "status": "completed"} tags=[]
test_df.shape
# + [markdown] papermill={"duration": 0.037634, "end_time": "2022-02-22T07:23:28.105416", "exception": false, "start_time": "2022-02-22T07:23:28.067782", "status": "completed"} tags=[]
# # Model Parameters Defining
# + papermill={"duration": 0.05703, "end_time": "2022-02-22T07:23:28.200555", "exception": false, "start_time": "2022-02-22T07:23:28.143525", "status": "completed"} tags=[]
numeric_features = train_df.select_dtypes(exclude=['object']).columns
# + papermill={"duration": 0.048271, "end_time": "2022-02-22T07:23:28.287071", "exception": false, "start_time": "2022-02-22T07:23:28.2388", "status": "completed"} tags=[]
numeric_features
# + papermill={"duration": 0.046629, "end_time": "2022-02-22T07:23:28.37204", "exception": false, "start_time": "2022-02-22T07:23:28.325411", "status": "completed"} tags=[]
numeric_features = list(numeric_features)[:-1]
# + papermill={"duration": 0.048826, "end_time": "2022-02-22T07:23:28.459139", "exception": false, "start_time": "2022-02-22T07:23:28.410313", "status": "completed"} tags=[]
numeric_df = train_df[numeric_features]
# + papermill={"duration": 0.047444, "end_time": "2022-02-22T07:23:28.545693", "exception": false, "start_time": "2022-02-22T07:23:28.498249", "status": "completed"} tags=[]
numeric_df.shape
# + [markdown] papermill={"duration": 0.044064, "end_time": "2022-02-22T07:23:28.630009", "exception": false, "start_time": "2022-02-22T07:23:28.585945", "status": "completed"} tags=[]
# # Check Missing Data
# + papermill={"duration": 0.051701, "end_time": "2022-02-22T07:23:28.721381", "exception": false, "start_time": "2022-02-22T07:23:28.66968", "status": "completed"} tags=[]
numeric_df.isnull().sum()
# + [markdown] papermill={"duration": 0.038994, "end_time": "2022-02-22T07:23:28.79962", "exception": false, "start_time": "2022-02-22T07:23:28.760626", "status": "completed"} tags=[]
# # Handle Missing Data
# + papermill={"duration": 0.047255, "end_time": "2022-02-22T07:23:28.885925", "exception": false, "start_time": "2022-02-22T07:23:28.83867", "status": "completed"} tags=[]
numeric_df = numeric_df.fillna(0)
# + papermill={"duration": 0.050875, "end_time": "2022-02-22T07:23:28.976715", "exception": false, "start_time": "2022-02-22T07:23:28.92584", "status": "completed"} tags=[]
numeric_df.isnull().sum()
# + papermill={"duration": 0.047116, "end_time": "2022-02-22T07:23:29.064068", "exception": false, "start_time": "2022-02-22T07:23:29.016952", "status": "completed"} tags=[]
X = numeric_df.copy()
y = train_df.SalePrice
# + papermill={"duration": 0.050362, "end_time": "2022-02-22T07:23:29.155118", "exception": false, "start_time": "2022-02-22T07:23:29.104756", "status": "completed"} tags=[]
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.25, random_state=1)
# + [markdown] papermill={"duration": 0.039784, "end_time": "2022-02-22T07:23:29.234351", "exception": false, "start_time": "2022-02-22T07:23:29.194567", "status": "completed"} tags=[]
# # Model Creation
# + [markdown] papermill={"duration": 0.039512, "end_time": "2022-02-22T07:23:29.313735", "exception": false, "start_time": "2022-02-22T07:23:29.274223", "status": "completed"} tags=[]
# ## Random Forest Model
# + papermill={"duration": 9.634177, "end_time": "2022-02-22T07:23:38.987498", "exception": false, "start_time": "2022-02-22T07:23:29.353321", "status": "completed"} tags=[]
rf_model = RandomForestRegressor(random_state=1, n_estimators=700)
rf_model.fit(X_train, y_train)
rf_val_prediction = rf_model.predict(X_test)
rf_val_rmse = np.sqrt(mean_squared_error(rf_val_prediction, y_test))
print(f'RMSE : {rf_val_rmse}')
# + [markdown] papermill={"duration": 0.04257, "end_time": "2022-02-22T07:23:39.073523", "exception": false, "start_time": "2022-02-22T07:23:39.030953", "status": "completed"} tags=[]
# ## Gradient Boost Model
# + papermill={"duration": 1.95265, "end_time": "2022-02-22T07:23:41.069143", "exception": false, "start_time": "2022-02-22T07:23:39.116493", "status": "completed"} tags=[]
gb_model = GradientBoostingRegressor(random_state=1, n_estimators=375, min_samples_split=3, min_samples_leaf=2)
gb_model.fit(X_train, y_train)
gb_val_prediction = gb_model.predict(X_test)
gb_val_rmse = round(np.sqrt(mean_squared_error(gb_val_prediction, y_test)), 2)
print(f'RMSE : {gb_val_rmse}')
# + [markdown] papermill={"duration": 0.039645, "end_time": "2022-02-22T07:23:41.14984", "exception": false, "start_time": "2022-02-22T07:23:41.110195", "status": "completed"} tags=[]
# ## Linear Regression Model
# + papermill={"duration": 0.237019, "end_time": "2022-02-22T07:23:41.427078", "exception": false, "start_time": "2022-02-22T07:23:41.190059", "status": "completed"} tags=[]
linreg_model = LinearRegression()
linreg_model.fit(X_train, y_train)
linreg_val_prediction = linreg_model.predict(X_test)
linreg_val_rmse = np.sqrt(mean_squared_error(linreg_val_prediction, y_test))
print(f'RMSE : {linreg_val_rmse}')
# + [markdown] papermill={"duration": 0.057254, "end_time": "2022-02-22T07:23:41.558849", "exception": false, "start_time": "2022-02-22T07:23:41.501595", "status": "completed"} tags=[]
# # Build the Best Model on full dataset
# + [markdown] papermill={"duration": 0.043009, "end_time": "2022-02-22T07:23:41.642602", "exception": false, "start_time": "2022-02-22T07:23:41.599593", "status": "completed"} tags=[]
# ## Create the Model
# + papermill={"duration": 2.517458, "end_time": "2022-02-22T07:23:44.202596", "exception": false, "start_time": "2022-02-22T07:23:41.685138", "status": "completed"} tags=[]
gb_model_on_full_data = GradientBoostingRegressor(random_state=1, n_estimators=375, min_samples_split=3, min_samples_leaf=2)
gb_model_on_full_data.fit(X, y)
# + [markdown] papermill={"duration": 0.042337, "end_time": "2022-02-22T07:23:44.286456", "exception": false, "start_time": "2022-02-22T07:23:44.244119", "status": "completed"} tags=[]
# ## Create the Test Dataframe with selected Numeric Features
# + papermill={"duration": 0.049857, "end_time": "2022-02-22T07:23:44.377074", "exception": false, "start_time": "2022-02-22T07:23:44.327217", "status": "completed"} tags=[]
test_X = test_df[numeric_features]
# + papermill={"duration": 0.050658, "end_time": "2022-02-22T07:23:44.469859", "exception": false, "start_time": "2022-02-22T07:23:44.419201", "status": "completed"} tags=[]
test_X.shape
# + [markdown] papermill={"duration": 0.040599, "end_time": "2022-02-22T07:23:44.551364", "exception": false, "start_time": "2022-02-22T07:23:44.510765", "status": "completed"} tags=[]
# ## Check the Missing Data
# + papermill={"duration": 0.053508, "end_time": "2022-02-22T07:23:44.647375", "exception": false, "start_time": "2022-02-22T07:23:44.593867", "status": "completed"} tags=[]
test_X.isnull().sum()
# + [markdown] papermill={"duration": 0.041523, "end_time": "2022-02-22T07:23:44.730446", "exception": false, "start_time": "2022-02-22T07:23:44.688923", "status": "completed"} tags=[]
# ## Handle the Missing Data
# + papermill={"duration": 0.052016, "end_time": "2022-02-22T07:23:44.825245", "exception": false, "start_time": "2022-02-22T07:23:44.773229", "status": "completed"} tags=[]
test_X = test_X.fillna(0)
# + papermill={"duration": 0.053311, "end_time": "2022-02-22T07:23:44.920306", "exception": false, "start_time": "2022-02-22T07:23:44.866995", "status": "completed"} tags=[]
test_X.isnull().sum()
# + [markdown] papermill={"duration": 0.042138, "end_time": "2022-02-22T07:23:45.004168", "exception": false, "start_time": "2022-02-22T07:23:44.96203", "status": "completed"} tags=[]
# ## Make predictions
# + papermill={"duration": 0.060656, "end_time": "2022-02-22T07:23:45.106351", "exception": false, "start_time": "2022-02-22T07:23:45.045695", "status": "completed"} tags=[]
# make predictions which we will submit.
test_preds = gb_model_on_full_data.predict(test_X)
# + papermill={"duration": 0.050143, "end_time": "2022-02-22T07:23:45.198108", "exception": false, "start_time": "2022-02-22T07:23:45.147965", "status": "completed"} tags=[]
test_preds
# + [markdown] papermill={"duration": 0.042138, "end_time": "2022-02-22T07:23:45.281895", "exception": false, "start_time": "2022-02-22T07:23:45.239757", "status": "completed"} tags=[]
# # Save the Model
# + papermill={"duration": 0.058488, "end_time": "2022-02-22T07:23:45.382537", "exception": false, "start_time": "2022-02-22T07:23:45.324049", "status": "completed"} tags=[]
# The lines below shows how to save predictions in format used for competition scoring
output = pd.DataFrame({'Id': test_df.Id,
'SalePrice': test_preds})
output.to_csv('submission.csv', index=False)
# + [markdown] papermill={"duration": 0.041793, "end_time": "2022-02-22T07:23:45.466747", "exception": false, "start_time": "2022-02-22T07:23:45.424954", "status": "completed"} tags=[]
# ##
| simple-house-price-prediction-model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Example LaTeX notation
# https://en.wikibooks.org/wiki/LaTeX/Mathematics
#
#
# |||||
# |-|-|-|-|
# |\alpha<br/>$\alpha$ | A<br/>$A$| \beta<br/>$\beta$ | B<br/>$B$ | \gamma<br/>$\gamma$ | \Gamma<br/>$\Gamma$ | \delta<br/>$\delta$ | \Delta<br/>$\Delta$ |
# | \epsilon<br/>$\epsilon$ | \varepsilon<br/>$\varepsilon$ | E<br/>$E$ | \zeta<br/>$\zeta$ | Z<br/>$Z$ | \eta<br/>$\eta$ | H<br/>$H$ | \theta<br/>$\theta$ |
# | \vartheta<br/>$\vartheta$ | \Theta<br/>$\Theta$ | \iota<br/>$\iota$ | I<br/>$I$ | \kappa<br/>$\kappa$ | \varkappa<br/>$\varkappa$ | K<br/>$K$ | \lambda<br/>$\lambda$ |
# | \lambda<br/>$\lambda$ | \mu<br/>$\mu$ | M<br/>$M$ | \nu<br/>$\nu$ | N<br/>$N$ | \xi<br/>$\xi$ | \Xi<br/>$\Xi$ | \omicron<br/>$\omicron$ |
# | O<br/>$O$ | \pi<br/>$\pi$ | \varpi<br/>$\varpi$ | \Pi<br/>$\Pi$ | \rho<br/>$\rho$ | \varrho<br/>$\varrho$ | P<br/>$P$ | \sigma<br/>$\sigma$ |
# | \varsigma<br/>$\varsigma$ | \Sigma<br/>$\Sigma$ | \tau<br/>$\tau$ | T<br/>$T$ | \upsilon<br/>$\upsilon$ | \Upsilon<br/>$\Upsilon$ | \phi<br/>$\phi$ | \varphi<br/>$\varphi$ |
# | \Phi<br/>$\Phi$ | \chi<br/>$\chi$ | X<br/>$X$ | \psi<br/>$\psi$ | \Psi<br/>$\Psi$ | \omega<br/>$\omega$ | \Omega<br/>$\Omega$ |
#
# |||
# |-|-|
# |( \big( \Big( \bigg( \Bigg( <br/>\Bigg) \bigg) \Big) \big) )<br/>$\displaystyle ( \big( \Big( \bigg( \Bigg( \Bigg) \bigg) \Big) \big) )$|\{ \big\{ \Big\{ \bigg\{ \Bigg\{ <br/>\Bigg\} \bigg\} \Big\} \big\} \} <br/>$\displaystyle \{ \big\{ \Big\{ \bigg\{ \Bigg\{ \Bigg\} \bigg\} \Big\} \big\} \}$|
# |[ \big[ \Big[ \bigg[ \Bigg[<br/>\Bigg] \bigg] \Big] \big] ] <br/>$\displaystyle [ \big[ \Big[ \bigg[ \Bigg[ \Bigg] \bigg] \Big] \big] ]$|\langle \big \langle \Big \langle \bigg \langle \Bigg \langle <br/> \Bigg \rangle \bigg \rangle \Big \rangle \big \rangle \rangle <br/>$\displaystyle \langle \big \langle \Big \langle \bigg \langle \Bigg \langle \Bigg \rangle \bigg \rangle \Big \rangle \big \rangle \rangle$|
#
#
# ### Symbols
#
# ||||||||||||
# |-|-|-|-|-|-|-|-|-|-|-|
# | Addition/Subtraction | add<br/>**a+b**<br/>$a+b$ | subtract<br/>**a-b**<br/>$a-b$ | plus or minus<br/>**\pm**<br/>$\pm$ | minus or plus<br/>**\mp**<br/>$\mp$ |
# | Multiply/Divide | multiply<br/>**ab**<br/>$ab$ | dot multiply<br/>**a\cdotb**<br/>$a\cdot b$ | times<br/>**a\times b**<br/>$a\times b$ | inline divide<br/>**^a/_b**<br/>$^a/_b$ | fraction<br/>**\frac{a}{b}**<br/>$\frac{a}{b}$ |
# | Power/Root | power<br/>**a^{n}**<br/>$a^{n}$ | power fraction<br/>**a^\frac{m}{n}**<br/>$a^\frac{m}{n}$ | power power<br/>**a^{b^{c}}**<br/>$a^{b^{c}}$ | square root<br/>**\sqrt{m}**<br/>$\sqrt{m}$ | root<br/>**\sqrt[n]{m}**<br/>$\sqrt[n]{m}$ | root fraction<br/>**\sqrt{\frac{a}{b}}**<br/>$\sqrt{\frac{a}{b}}$ |
# | Factorial | factorial<br/>**!**<br/>$!$ |
# | Modulo | modulo<br/>**a \bmod b**<br/>$a \bmod b$ | modulo<br/>**a \pmod b**<br/>$a \pmod b$ |
# |
# | Equality/Equivalence Relation | equal<br/>**a=b**<br/>$a=b$ | equivalent<br/>**a\equiv b**<br/>$a\equiv b$ | not equal<br/>**\neq**<br/>$\neq$ | proportional to<br/>**\propto**<br/>$\propto$ | approach to a limit<br/>**\doteq**<br/>$\doteq$ | approximate<br/>**\approx**<br/>$\approx$ | is isomorphic to<br/>**\cong**<br/>$\cong$ | similar<br/>**\sim**<br/>$\sim$ | similar or equal<br/>**\simeq**<br/>$\simeq$ |
# | Inequality Relation | less than<br/>**<**<br/>$<$ | less than or equal<br/>**\leq**<br/>$\leq$ | greater than<br/>**>**<br/>$>$ | greater than or equal<br/>**\geq**<br/>$\geq$ | ll<br/>**\ll**<br/>$\ll$ | gg<br/>**\gg**<br/>$\gg$ |
# |
# | Trigonometry | sine<br/>**\sin(\theta)**<br/>$\sin(\theta)$ | cosine<br/>**\cos(\theta)**<br/>$\cos(\theta)$ | tangent<br/>**\tan(\theta)**<br/>$\tan(\theta)$ | secant<br/>**\sec(\theta)**<br/>$\sec(\theta)$ | cosecant<br/>**\csc(\theta)**<br/>$\csc(\theta)$ | cotangent<br/>**\cot(\theta)**<br/>$\cot(\theta)$ |
# | Inverse Trigonometry | arc sine<br/>**\arcsin(\theta)**<br/>$\arcsin(\theta)$ | arc cosine<br/>**\arccos(\theta)**<br/>$\arccos(\theta)$ | arc tangent<br/>**\arctan(\theta)**<br/>$\arctan(\theta)$ | arc secant | arc cosecant | arc cotangent |
# | Hyperbolic | hyperbolic sine<br/>**\sinh(\theta)**<br/>$\sinh(\theta)$ | hyperbolic cosine<br/>**\cosh(\theta)**<br/>$\cosh(\theta)$ | hyperbolic tangent<br/>**\tanh(\theta)**<br/>$\tanh(\theta)$ | hyperbolic secant | hyperbolic cosecant | hyperbolic cotangent<br/>**\coth(\theta)**<br/>$\coth(\theta)$ |
# |
# | Sum/Product | sum<br/>**\displaystyle<br/>\sum_{i=n}^{10}t_i**<br/>$\displaystyle \sum_{i=n}^{10}t_i$ | product<br/>**\displaystyle<br/>\prod_{i=n}^{10}t_i**<br/>$\displaystyle \prod_{i=n}^{10}t_i$ | coproduct<br/>**\displaystyle<br/>\coprod_{j\in J} X_j**<br/>$\displaystyle \coprod_{j\in J}X_j$ | big-o-plus<br/>**\displaystyle<br/>\bigoplus_{j\in J} X_j**<br/>$\displaystyle \bigoplus_{j\in J}X_j$ | big-o-times<br/>**\displaystyle<br/>\bigotimes_{j\in J} X_j**<br/>$\displaystyle \bigotimes_{j\in J}X_j$ | big-o-dot<br/>**\displaystyle<br/>\bigodot_{j\in J} X_j**<br/>$\displaystyle \bigodot_{j\in J}X_j$ |
# |
# | Calculus | limit<br/>**\displaystyle<br/>\lim_{x\to\infty}**<br/>$\displaystyle \lim_{x\to\infty}$| integral<br/>**\displaystyle<br/>\int_0^\infty**<br/>$\displaystyle \int_0^\infty$ | 2nd integral<br/>**\displaystyle<br/>\iint_0^\infty**<br/>$\displaystyle \iint_0^\infty$ | 3rd integral<br/>**\displaystyle<br/>\iiint_0^\infty**<br/>$\displaystyle \iiint_0^\infty$ | o integral<br/>**\displaystyle<br/>\oint_0^\infty**<br/>$\displaystyle \oint_0^\infty$ |
# |
# | Set Relation | subset<br/>**\subset**<br/>$\subset$ | subset or equal<br/>**\subseteq**<br/>$\subseteq$ | n subset or equal<br/>**\nsubseteq**<br/>$\nsubseteq$ | superset<br/>**\supset**<br/>$\supset$ | superset or equal<br/>**\supseteq**<br/>$\supseteq$ | n superset or equal<br/>**\nsupseteq**<br/>$\nsupseteq$ | square subset<br/>**\sqsubset**<br/>$\sqsubset$ | square subset or equal<br/>**\sqsubseteq**<br/>$\sqsubseteq$ | square superset<br/>**\sqsupset**<br/>$\sqsupset$ | square superset or equal<br/>**\sqsupseteq**<br/>$\sqsupseteq$ |
# | Logic Notation | exists<br/>**\exists**<br/>$\exists$ | does not exist<br/>**\nexists**<br/>$\nexists$ | for all<br/>**\forall**<br/>$\forall$ | element of<br/>**\in**<br/>$\in$ | not element of<br/>**\notin**<br/>$\notin$ | not element of<br/>**\ni**<br/>$\ni$ |
# | Logic Operators | not<br/>**\neg**<br/>$\neg$ | logical and<br/>**\land**<br/>$\land$ | logical or<br/>**\lor**<br/>$\lor$ | implies<br/>**\implies**<br/>$\implies$ | if and only if<br/>**\iff**<br/>$\iff$ | turnstile<br/>**\vdash**<br/>$\vdash$ | reversed turnstile<br/>**\dashv**<br/>$\dashv$ |
# |
# | Other | tensor product<br/>**\otimes**<br/>$\otimes$ | direct sum sets<br/>**\oplus**<br/>$\oplus$ |
# | unamed | times<br/>**\times**<br/>$\times$ | cup<br/>**\cup**<br/>$\cup$ | cap<br/>**\cap**<br/>$\cap$ |
# |
# | functions | ker<br/>**\ker**<br/>$\ker$ | lim sup<br/>**\limsup**<br/>$\limsup$ | deg<br/>**\deg**<br/>$\deg$ | gcd<br/>**\gcd**<br/>$\gcd$ | lg<br/>**\lg**<br/>$\lg$ |
# | functions | sup<br/>**\sup**<br/>$\sup$ | det<br/>**\det**<br/>$\det$ |hom<br/>**\hom**<br/>$\hom$ |arg<br/>**\arg**<br/>$\arg$ |dim<br/>**\dim**<br/>$\dim$ |lim inf<br/>**\liminf**<br/>$\liminf$ |
# |
# | unamed | asymptotic<br/>**\asymp**<br/>$\asymp$ | bowtie<br/>**\bowtie**<br/>$\bowtie$ |
# |
# | unamed | smile<br/>**\smile**<br/>$\smile$ | frown<br/>**\frown**<br/>$\frown$ |models<br/>**\models**<br/>$\models$ |mid<br/>**\mid**<br/>$\mid$ |sphericalangle<br/>**\sphericalangle**<br/>$\sphericalangle$ |measuredangle<br/>**\measuredangle**<br/>$\measuredangle$ |
# |
# | order | precedes<br/>**\prec**<br/>$\prec$ | succeeds<br/>**\succ**<br/>$\succ$ | precedes / equals<br/>**\preceq**<br/>$\preceq$ | succeeds / equals<br/>**\succeq**<br/>$\succeq$ |
# | geometry | parallel<br/>**\parallel**<br/>$\parallel$ | not parallel<br/>**\nparallel**<br/>$\nparallel$ | perpendicular <br/>**\perp**<br/>$\perp$ |
# ( ) [ ]| ' :
#
# \parallel → $\parallel$
# \nparallel → $\nparallel$
#
# factorial<br/>**!**
#
# Binomial $\binom{x}{y}$
#
# Evaluation $f(n) = n^5 + 4n^2 + 2 |_{n=17}$
#
#
# \parallel → $\parallel$
# $\displaystyle \lim_{0->\infty}$
# radius = 1
# regular polygon inside a circle, cannot use pi in our calculations
#
# 2 sides is a double-sided line across the diameter
# c = 2r + 2r = 4r
#
# | Example Description | Example LaTeX Notation | Example LaTeX-Rendered Symbols |
# | ---- | ---- | ----- |
# | Complex Fraction | \frac{\frac{a}{b}+\frac{c}{d}}{e-f^g} | $\frac{\frac{a}{b}+\frac{c}{d}}{e-f^g}$ |
# | Factorial | \frac{n!}{k!(n-k)!} | $\frac{n!}{k!(n-k)!}$ |
# | Polynomial | a.x^2 b.x + c | $a x^2+b x+c$ |
# | Exponential | \exp(-x) = 0 | $\exp(-x) = 0$ |
# | Natural Logarithmic | \ln(y) = x | $\ln(y) = x$ |
# | Logarithmic | \log_{10}(y) = x | $\log_{10}(y) = x$ |
# | For all x from set X | \forall x \in X | $\forall x \in X$ |
# | There exists y less than epsilon | \exists y \leq \epsilon | $\exists y \leq \epsilon$ |
# | Quadratic Eq | \over{-b\pm\sqrt{4ac + b^2}}{2a} | ${-b\pm\sqrt{4ac + b^2}}\over{2a}$ |
| notebooks/.ipynb_checkpoints/latex-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # High-Level API
# ## Overview
# The High-Level API makes it easy to rapidly:
#
# * Prepare samples.
# * Hypertune/ train batches of models.
# * Feed appropriate data/ parameters into those models.
# * Evaluate model performance with metrics and plots.
#
# It does so by wrapping and bundling together the methods of the [Low-Level API](api_low_level.html). The table below demonstrates how the high-level entities abstract abstract the low-level entities. While this abstraction eliminates many steps to enable rapid model prototyping, it comes at a cost of customization.
# | High-level object | Groups together or creates the following objects |
# |:-----------------:|:-----------------------------------------------------------------------------------------------------------------:|
# | `Pipeline` | Dataset, File, Image, Tabular, Label, Featureset, Splitset, Foldset, Folds, Encoderset, Labelcoder, Featurecoder. |
# | `Algorithm` | Functions to build, train, predict, and evaluate a machine learning model. |
# | `Experiment` | Algorithm, Hyperparamset, Hyperparamcombos, Batch, Job, Jobset, Result. |
# ## Prerequisites
# If you've already completed the instructions on the [Installation](installation.html) page, then let's get started.
import aiqc
from aiqc import datum
# ## 1. Pipeline
# ### a) Tabular Dataset
# Tabular/ delimited/ flat-file `Dataset.Tabular` can be created from either Pandas DataFrames or flat files (CSV/ TSV or Parquet).
# Let's grab one of AIQC's built-in datasets from the `datum` module that we imported above. This module is described in the 'Built-In Examples - Datasets' section of the documentation.
df = datum.to_pandas(name='iris.tsv')
# The `Pipeline` process starts with raw data. A Dataset object is generated from that data and prepared for training based on the parameters the user provides to the `Pipeline.make` method. To get started, set the `dataFrame_or_filePath` equal to the dataframe we just fetched. It's the only argument that's actually required so
# Import any scikit-learn encoders that you want to use to encode labels and/ or features. Any encoders that you pass in will need to be instantiated with the attributes you want them to use.
# > Reference the `Encoderset` section of the low-level API for more detail on how to include/ exclude specific `Featureset` columns by name/dtype. The `feature_encoders` argument seen below takes a list of dictionaries as input, where each dictionary contains the `**kwargs` for a `Featurecoder`.
from sklearn.preprocessing import OneHotEncoder, PowerTransformer
# Rather than wrangling your data with many lines of data science code, just set the arguments below and AIQC takes care of the rest: stratification (including continuous dtypes), validation splits, cross-validation folds, and dtype/column specific encoders to be applied on-read.
# > Don't use `fold_count` unless your (total sample count / fold_count) still gives you an accurate representation of your sample population. You can try it with the 'iris_10x.tsv' datum.
splitset = aiqc.Pipeline.Tabular.make(
dataFrame_or_filePath = df
, dtype = None
, label_column = 'species'
, features_excluded = None
, size_test = 0.24
, size_validation = 0.12
, fold_count = None
, bin_count = None
, label_encoder = OneHotEncoder(sparse=False)
, feature_encoders = [{
"sklearn_preprocess": PowerTransformer(method='box-cox', copy=False)
, "dtypes": ['float64']
}]
)
# ### b) Image Dataset
# AIQC also supports image data and convolutional analysis.
#
# In order to perform *supervised learning* on image files, you'll need both a `Dataset.Image` and a `Dataset.Tabular`:
#
# * `Dataset.Image` can be created from either a folder of images or a list of urls. The Pillow library is used to normalize images ingested into AIQC. Each image must be the same size (dimensions) and mode (colorscale).
#
# * `Dataset.Tabular` is created as seen in the section above. It must contain 1 row per image.
#
# * Then a `Splitset` is constructed using:
# * The `Label` of the `Dataset.Tabular`.
# * The `Featureset` of the `Dataset.Image`.
# Again, we'll use the built-in data found in the `datum` module that we imported above.
df = datum.to_pandas(name='brain_tumor.csv')
image_urls = datum.get_remote_urls(manifest_name='brain_tumor.csv')
img_splitset = aiqc.Pipeline.Image.make(
folderPath_or_urls = image_urls
, pillow_save = {}
, tabularDF_or_path = df
, tabular_dtype = None
, label_column = 'status'
, label_encoder = None
, size_test = 0.30
, size_validation = None
, fold_count = 4
, bin_count = None
)
# ## 2. Experiment
# As seen in the [Compatibility Matrix](compatibility.html), the only library supported at this point in time is `Keras` as it is the most straightforward for entry-level users.
#
# > You can find great examples of machine learning cookbooks on this blog: [MachineLearningMastery.com "Multi-Label Classification"](https://machinelearningmastery.com/multi-label-classification-with-deep-learning/)
import keras
from keras.models import Sequential
from keras.layers import Dense, Dropout
from keras.callbacks import History
# ### Model Functions
# When we define our models, we'll do so by wrapping each phase in the following functions:
#
# * `function_model_build` contains the topology and parameters.
# * `function_model_train` specifies the samples and how the model should run.
# * `function_model_predict` and `function_model_loss` are automatically determined based on the `Algorithm.analysis_type`.
# Because these are full-blown functions, we can even play with the topology as a parameter! As demonstrated by the `if (hyperparameters['extra_layer'])` line below.
# > You can name the functions below whatever you want, but do not change their predetermined arugments (e.g. `features_shape`, `**hyperparameters`, `model`, etc.). These items are used behind the scenes to pass the appropriate data, parameters, and models into your training jobs.
#
# > Put a placeholder anywhere you want to try out different hyperparameters: `hyperparameters['<some_variable_name>']`. You'll get a chance to define the hyperparameters in a minute.
# #### `function_model_build()`
def function_model_build(features_shape, label_shape, **hyperparameters):
model = Sequential()
model.add(Dense(units=features_shape[0], activation='relu', kernel_initializer='he_uniform'))
model.add(Dropout(hyperparameters['dropout_size']))
if (hyperparameters['extra_layer']):
model.add(Dense(units=hyperparameters['neuron_count'], activation='relu', kernel_initializer='he_uniform'))
model.add(Dropout(hyperparameters['dropout_size']))
model.add(Dense(units=label_shape[0], activation='softmax', name='output'))
model.compile(optimizer='adamax', loss='categorical_crossentropy', metrics=['accuracy'])
return model
# #### `function_model_train()`
def function_model_train(model, samples_train, samples_evaluate, **hyperparameters):
model.fit(
samples_train["features"]
, samples_train["labels"]
, validation_data = (
samples_evaluate["features"]
, samples_evaluate["labels"]
)
, verbose = 0
, batch_size = 3
, epochs = hyperparameters['epoch_count']
, callbacks=[History()]
)
return model
# > Reference the [low-level API documentation](api_low_level.html#Optional,-callback-to-stop-training-early.) for information on the custom 'early stopping' callbacks AIQC makes available.
# ### Hyperparameters
# The `hyperparameters` below will be automatically fed into the functions above as `**kwargs` via the `**hyperparameters` argument we saw earlier.
#
# For example, wherever you see `hyperparameters['neuron_count']`, it will pull from the *key:value* pair `"neuron_count": [9, 12]` seen below. Where model A will have 9 neurons and model B will have 12 neurons.
hyperparameters = {
"neuron_count": [9, 12]
, "extra_layer": [True, False]
, "dropout_size": [0.10, 0.20]
, "epoch_count": [50]
}
# Then pass these functions into the `Algorithm`.
#
# The `library` and `analysis_type` help handle the model and its output behind the scenes. Current analysis types include: 'classification_multi', 'classification_binary', and 'regression'.
# ### `Experiment.make()`
# Now it's time to bring together the data and logic into an `Experiment`.
batch = aiqc.Experiment.make(
library = "keras"
, analysis_type = "classification_multi"
, function_model_build = function_model_build
, function_model_train = function_model_train
, splitset_id = splitset.id
, repeat_count = 2
, hide_test = False
, function_model_predict = None #automated
, function_model_loss = None #automated
, hyperparameters = hyperparameters
, foldset_id = None
, encoderset_id = splitset.encodersets[0]
)
batch.run_jobs()
# ---
# For more information on visualization of performance metrics, reference the [Visualization & Metrics](visualization.html) documentation.
| docs/_build/html/notebooks/api_high_level.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Convergence
#
# Description of the UCI protocol: https://ucichessengine.wordpress.com/2011/03/16/description-of-uci-protocol/
# Let us parse the logs first:
# %pylab inline
# ! grep "multipv 1" log8c.txt | grep -v lowerbound | grep -v upperbound > log8c_g.txt
def parse_info(l):
D = {}
k = l.split()
i = 0
assert k[i] == "info"
i += 1
while i < len(k):
if k[i] == "depth":
D[k[i]] = int(k[i+1])
i += 2
elif k[i] == "seldepth":
D[k[i]] = int(k[i+1])
i += 2
elif k[i] == "multipv":
D[k[i]] = int(k[i+1])
i += 2
elif k[i] == "score":
if k[i+1] == "cp":
D["score_p"] = int(k[i+2]) / 100. # score in pawns
i += 3
elif k[i] == "nodes":
D[k[i]] = int(k[i+1])
i += 2
elif k[i] == "nps":
D[k[i]] = int(k[i+1])
i += 2
elif k[i] == "hashfull":
D[k[i]] = int(k[i+1]) / 1000. # between 0 and 1
i += 2
elif k[i] == "tbhits":
D[k[i]] = int(k[i+1])
i += 2
elif k[i] == "time":
D[k[i]] = int(k[i+1]) / 1000. # elapsed time in [s]
i += 2
elif k[i] == "pv":
D[k[i]] = k[i+1:]
return D
else:
raise Exception("Unknown kw")
# +
# Convert to an array of lists
D = []
for l in open("log8c_g.txt").readlines():
D.append(parse_info(l))
# Convert to a list of arrays
data = {}
for key in D[-1].keys():
d = []
for x in D:
if key in x:
d.append(x[key])
else:
d.append(-1)
if key != "pv":
d = array(d)
data[key] = d
# -
# ## The Speed of Search
# The number of nodes searched depend linearly on time:
title("Number of nodes searched in time")
plot(data["time"] / 60., data["nodes"], "o")
xlabel("Time [min]")
ylabel("Nodes")
grid()
show()
# So nodes per second is roughly constant:
title("Positions per second in time")
plot(data["time"] / 60., data["nps"], "o")
xlabel("Time [min]")
ylabel("Positions / s")
grid()
show()
# The hashtable usage is at full capacity:
title("Hashtable usage")
hashfull = data["hashfull"]
hashfull[hashfull == -1] = 0
plot(data["time"] / 60., hashfull * 100, "o")
xlabel("Time [min]")
ylabel("Hashtable filled [%]")
grid()
show()
# Number of nodes needed for the given depth grows exponentially, except for moves that are forced, which require very little nodes to search (those show as a horizontal plateau):
title("Number of nodes vs. depth")
semilogy(data["depth"], data["nodes"], "o")
x = data["depth"]
y = exp(x/2.2)
y = y / y[-1] * data["nodes"][-1]
semilogy(x, y, "-")
xlabel("Depth [half moves]")
ylabel("Nodes")
grid()
show()
title("Number of time vs. depth")
semilogy(data["depth"], data["time"]/60., "o")
xlabel("Depth [half moves]")
ylabel("Time [min]")
grid()
show()
# ## Convergence wrt. Depth
title("Score")
plot(data["depth"], data["score_p"], "o")
xlabel("Depth [half moves]")
ylabel("Score [pawns]")
grid()
show()
# Convergence of the variations:
for i in range(len(data["depth"])):
print "%2i %s" % (data["depth"][i], " ".join(data["pv"][i])[:100])
| examples_manual/Convergence4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/emretezisci/Hello-World-/blob/master/UST_TRY_Basic.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="ZARiJVXW1p00" colab_type="code" colab={}
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import datetime
from fbprophet import Prophet
from fbprophet.diagnostics import performance_metrics
from fbprophet.diagnostics import cross_validation
from fbprophet.plot import plot_cross_validation_metric
from fbprophet.plot import add_changepoints_to_plot
from fbprophet.plot import plot_yearly
# + id="rLjyo5GZ15Ed" colab_type="code" colab={}
# Create a datetime object with today's value
today = datetime.datetime.today()
# Add one day to today's date
tomorrow = today + datetime.timedelta(1)
# I imported tomorrow's data because of timezome difference
startDate = '01-01-2005'
endDate = str(datetime.datetime.strftime(tomorrow,'%d-%m-%Y'))
key = ''
aggregationTypes = 'max'
frequency = '2'
# Import the data
url = "https://evds2.tcmb.gov.tr/service/evds/series=TP.DK.USD.A&startDate=" + startDate + '&endDate=' +endDate + '&type=csv' + '&key=' + key + '&aggregationTypes=' + aggregationTypes + '&frequency=' + frequency
df = pd.read_csv(url)
# + id="PtWHf1eV2Kwq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="2ac9b55c-77ac-4d8f-82a2-e37d6e7e397e"
# Sort the data in descending order
df = df.sort_index(ascending=False)
df.head()
# + id="NGgmOR814iY4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="edf688ee-be6d-4c54-cb2e-9aca0febcb59"
# Rename necessary columns and remove unnecessary one
df = df.rename(columns={"Tarih": "ds", "TP_DK_USD_A": "y"})
df.drop('UNIXTIME', axis=1, inplace=True)
df.head()
# + id="NPH-4Fj52UMO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="31d31a32-3759-4adf-f2f3-ff27cd5f01c8"
# Fit the mode
m = Prophet()
m.fit(df)
# + id="WrYC2otf6dcy" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="2a452ea1-cce2-4421-c9d1-204e3a240eba"
# Predict the future
# The predict method will assign each row in future a predicted value which it names yhat.
# If you pass in historical dates, it will provide an in-sample fit.
future = m.make_future_dataframe(periods=365)
future.tail()
# + id="XQEQJ6SV67Wz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="3e78adb4-dd87-4b1b-ce9e-ae24d5cf7ee6"
forecast = m.predict(future)
forecast[['ds', 'yhat', 'yhat_lower', 'yhat_upper']].tail()
# + id="0LFFS1pl-Hm2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="fdef2743-62a0-46b5-849a-04012f30a19b"
# Horizon of 365 days, starting with 730 days of training data in the first cutoff and then making predictions every 180 days.
df_cv = cross_validation(m, initial='730 days', period='180 days', horizon = '365 days')
df_cv.tail()
# + id="Wn857O6Y_67L" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="40281033-269b-4011-89c7-75c7c895b8ee"
df_p = performance_metrics(df_cv)
df_p.head()
# + id="k4IaBuYsXBIo" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="b98892f5-a7c5-4b67-9e9b-5989bac08029"
np.mean(df_p['mape'])
# + id="__2xOgVzeWxh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="d74de79a-b4ed-431a-98d0-48a01d519a44"
np.mean(df_p['rmse'])
# + id="vqcw4RpSAHkn" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 388} outputId="656ca3d7-db69-459c-c8a9-9d24a9758191"
fig = plot_cross_validation_metric(df_cv, metric='mape')
# + id="-Q__8U3u7Mys" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 437} outputId="31e5fd39-137c-4b06-890d-2efd43e5a59f"
# Plot the forecast
fig1 = m.plot(forecast)
# + id="pW3bcJ-W7Vq4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 653} outputId="4ea28f5b-640c-440a-ba80-f75f01cfed7b"
# Display trend, yearly seasonality, and weekly seasonality
fig1_1 = m.plot_components(forecast)
# + id="8gkPrOpYl3BH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 351} outputId="99cc536d-ce08-4f69-80e8-ab911972f10d"
forecastnew = forecast['ds']
forecastnew2 = forecast['yhat']
forecastnew = pd.concat([forecastnew,forecastnew2], axis=1)
mask = (forecastnew['ds'] > "2019-04-03") & (forecastnew['ds'] <= "2020-01-01")
forecastedvalues = forecastnew.loc[mask]
mask = (forecastnew['ds'] > "2005-01-03") & (forecastnew['ds'] <= "2019-04-03")
forecastnew = forecastnew.loc[mask]
fig, ax1 = plt.subplots(figsize=(21, 5))
ax1.plot(forecastnew.set_index('ds'), color='b')
ax1.plot(forecastedvalues.set_index('ds'), color='r')
ax1.set_ylabel('USD / TRY')
ax1.set_xlabel('Date')
print("Red = Predicted Values, Blue = Base Values")
# + id="vP8TK01_8ccW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 454} outputId="8bf5560f-0b5c-42ae-9051-27722088f7f2"
# Try without the weekend data
df2 = df.copy()
df2['ds'] = pd.to_datetime(df2['ds'])
df2 = df2[df2['ds'].dt.dayofweek < 5]
m2 = Prophet().fit(df2)
future2 = m2.make_future_dataframe(periods=365, freq='D')
forecast2 = m2.predict(future2)
fig2 = m.plot(forecast2)
# + id="o2TyqIWu9ijX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="657a87e2-acd0-48a4-80ae-3fcc3a8787f2"
forecast2[['ds', 'yhat', 'yhat_lower', 'yhat_upper']].tail()
# + id="yAuVGda5A8oB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="2669984c-e635-4ceb-9b9f-06e8492752f6"
df_cv2 = cross_validation(m2, initial='730 days', period='180 days', horizon = '365 days')
df_cv2.tail()
# + id="GtgQcWHsA-ZY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="7ddf9cf6-8907-47c1-e752-6b4b7fafbccb"
df_p2 = performance_metrics(df_cv2)
df_p2.head()
# + id="uyCOybGkXE3R" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="9fa386fe-fdf4-4919-d0e7-d99c53fdfd10"
np.mean(df_p2['mape'])
# + id="QNUp71swfG77" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="68a286c8-b97a-4916-d7b6-51fc0c117862"
np.mean(df_p2['rmse'])
# + id="4BC8xLaFDSgE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 437} outputId="83c225f9-adce-41da-fa97-35a7e1a37ac2"
fig2 = m2.plot(forecast2)
a = add_changepoints_to_plot(fig2.gca(), m2, forecast2)
# + id="-_RS5IkVllxE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 351} outputId="97c4cec2-7aab-4d0d-e212-124b5aee3469"
forecastnew = forecast2['ds']
forecastnew2 = forecast2['yhat']
forecastnew = pd.concat([forecastnew,forecastnew2], axis=1)
mask = (forecastnew['ds'] > "2019-04-03") & (forecastnew['ds'] <= "2020-01-01")
forecastedvalues = forecastnew.loc[mask]
mask = (forecastnew['ds'] > "2005-01-03") & (forecastnew['ds'] <= "2019-04-03")
forecastnew = forecastnew.loc[mask]
fig, ax1 = plt.subplots(figsize=(21, 5))
ax1.plot(forecastnew.set_index('ds'), color='b')
ax1.plot(forecastedvalues.set_index('ds'), color='r')
ax1.set_ylabel('USD / TRY')
ax1.set_xlabel('Date')
print("Red = Predicted Values, Blue = Base Values")
# + id="ysXgaizuHBA9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="6dba7168-34ab-4f23-d703-39eb3ff3561a"
# Try with different interval widt and seasonality
df3 = df.copy()
df3['ds'] = pd.to_datetime(df3['ds'])
df3 = df3[df3['ds'].dt.dayofweek < 5]
m3 = Prophet(interval_width=0.75, yearly_seasonality=False, weekly_seasonality=False, changepoint_range=1) .fit(df3)
future3 = m3.make_future_dataframe(freq='D', periods=5)
future.tail()
# + id="2K6b7kv4I6VC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 437} outputId="7ea8d50e-ddae-4ff3-de9a-46feb1cc69e4"
forecast3 = m3.predict(future3)
fig3 = m3.plot(forecast3)
a = add_changepoints_to_plot(fig3.gca(), m3, forecast3)
# + id="_ImKParfTAKy" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="1b505ad3-4735-48f2-fb66-0d897ff8d56e"
forecast3[['ds', 'yhat', 'yhat_lower', 'yhat_upper']].tail()
# + id="DfkphNStTQws" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="e5dbef54-e09d-4324-ca4f-f46606ef07b7"
from fbprophet.diagnostics import cross_validation, performance_metrics
df_cv3 = cross_validation(m3, initial = '730 days', period = '180 days', horizon = '365 days')
# + id="122oq2cbUW3O" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="52d51eec-457a-44a2-d7d7-0fe0a20b868d"
df_p3 = performance_metrics(df_cv3)
df_p3.head()
# + id="WHV_C51ZUtZs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="f7be496f-1be8-4369-fd80-aa48f56e0b3c"
np.mean(df_p3['mape'])
# + id="t4Bgfo4SfDOm" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="ffd7bdc9-6453-45ad-86d2-7241889d51be"
np.mean(df_p3['rmse'])
# + id="4uVDlCxcU2JK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 388} outputId="a5c23c9c-ee95-4761-849b-2db8ce57422b"
fig3_1 = plot_cross_validation_metric(df_cv3, metric='mape')
# + id="wu-qwdR1VNrG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 351} outputId="08e022b0-6807-45dc-d27c-5be69c218c3b"
forecastnew = forecast3['ds']
forecastnew2 = forecast3['yhat']
forecastnew = pd.concat([forecastnew,forecastnew2], axis=1)
mask = (forecastnew['ds'] > "2019-04-03") & (forecastnew['ds'] <= "2020-01-01")
forecastedvalues = forecastnew.loc[mask]
mask = (forecastnew['ds'] > "2005-01-03") & (forecastnew['ds'] <= "2019-04-03")
forecastnew = forecastnew.loc[mask]
fig, ax1 = plt.subplots(figsize=(21, 5))
ax1.plot(forecastnew.set_index('ds'), color='b')
ax1.plot(forecastedvalues.set_index('ds'), color='r')
ax1.set_ylabel('USD / TRY')
ax1.set_xlabel('Date')
print("Red = Predicted Values, Blue = Base Values")
# + id="4o994IziZztN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 454} outputId="78e98ed3-6e96-4aab-b642-456293477d76"
# Try taking log of values
df4 = df.copy()
df4['ds'] = pd.to_datetime(df3['ds'])
df4 = df4[df4['ds'].dt.dayofweek < 5]
df4['y'] = np.log(df4['y'])
m4 = Prophet(interval_width=0.77, yearly_seasonality=False, weekly_seasonality=False, changepoint_range=1) .fit(df4)
future4 = m4.make_future_dataframe(freq='D', periods=5)
forecast4 = m4.predict(future4)
fig4 = m4.plot(forecast4)
a = add_changepoints_to_plot(fig4.gca(), m4, forecast4)
# + id="DScBEsLgasCy" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="dd1fc16f-b47d-42c9-df64-998b265cacc2"
forecast4[['ds', 'yhat', 'yhat_lower', 'yhat_upper']].tail()
# + id="81SAC1n1ch-i" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="91ff4010-e97d-4c59-9c14-57178932aed3"
df_cv4 = cross_validation(m4, initial = '730 days', period = '180 days', horizon = '365 days')
# + id="r6fa0Nd5cni2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="ac875ae1-f61c-48a8-f4d4-0586d49dea92"
df_p4 = performance_metrics(df_cv4)
df_p4.head()
# + id="L9cVYml0crJS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="5ad01ce0-9d8d-41c7-871e-a199feb3492a"
np.mean(df_p4['mape'])
# + id="QkOwLy92e8cQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="6fcdc021-1ab6-44db-82f1-8d9ce2fdbffc"
np.mean(df_p4['rmse'])
# + id="XCty-IhzcuD0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 388} outputId="b95e5630-e781-444e-b01e-684571df8840"
fig4 = plot_cross_validation_metric(df_cv4, metric='mape')
# + id="d26FbkTRjQPD" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 351} outputId="3d8eda29-10c8-4f16-b0a4-f3167d07511c"
forecastnew = forecast4['ds']
forecastnew2 = forecast4['yhat']
forecastnew2 = np.exp(forecastnew2)
forecastnew = pd.concat([forecastnew,forecastnew2], axis=1)
mask = (forecastnew['ds'] > "2019-04-03") & (forecastnew['ds'] <= "2020-01-01")
forecastedvalues = forecastnew.loc[mask]
mask = (forecastnew['ds'] > "2005-01-03") & (forecastnew['ds'] <= "2019-04-03")
forecastnew = forecastnew.loc[mask]
fig, ax1 = plt.subplots(figsize=(21, 5))
ax1.plot(forecastnew.set_index('ds'), color='b')
ax1.plot(forecastedvalues.set_index('ds'), color='r')
ax1.set_ylabel('USD / TRY')
ax1.set_xlabel('Date')
print("Red = Predicted Values, Blue = Base Values")
| USD-TRY_Prophet_Forecasting.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#Page 1
'''
Read the dataset in as a list using the csv module.
Import the csv module.
Open the file using the open() function.
Use the csv.reader() function to load the opened file.
Call list() on the result to get a list of all the data in the file.
Assign the result to the variable data.
Display the first 5 rows of data to verify everything.
'''
import csv
f = open("guns.csv", 'r')
data = list(csv.reader(f))
print(data[:5])
#Page 2
'''
Extract the first row of data, and assign it to the variable headers.
Remove the first row from data.
Display headers.
Display the first 5 rows of data to verify that you removed the header row properly.
'''
headers = data[1]
data = data[1:len(data)]
print(headers)
print(data[:5])
#Page 3
'''
Use a list comprehension to extract the year column from data.
Because the year column is the second column in the data, you'll need to get the element at index 1 in each row.
Assign the result to the variable years.
Create an empty dictionary called year_counts.
Loop through each element in years.
If the element isn't a key in year_counts, create it, and set the value to 1.
If the element is a key in year_counts, increment the value by one.
Display year_counts to see how many gun deaths occur in each year.
'''
years = [row[1] for row in data]
year_counts ={}
for y in years:
if y in year_counts:
year_counts[y] += 1
else:
year_counts[y] = 1
year_counts
# +
#Page 4
'''
Use a list comprehension to create a datetime.datetime object for each row. Assign the result to dates.
The year column is in the second element in each row.
The month column is the third element in each row.
Make sure to convert year and month to integers using int().
Pass year, month, and day=1 into the datetime.datetime() function.
Display the first 5 rows in dates to verify everything worked.
Count up how many times each unique date occurs in dates. Assign the result to date_counts.
This follows a similar procedure to what we did in the last screen with year_counts.
Display date_counts.
'''
import datetime as dt
current_datetime = dt.datetime.utcnow()
current_datetime
for row in data:
#print(row[2])
row_dateTime = dt.datetime(year = int(row[1]), month = int(row[2]), day = 22)
##print(row_dateTime)
row.append(row_dateTime)
date_count = {}
for row in data:
if row[11] in date_count:
date_count[row[11]] = date_count[row[11]] + 1
else:
date_count[row[11]] = 1
date_count
# -
#Page 5
'''
Count up how many times each item in the sex column occurs.
Assign the result to sex_counts.
Count up how many times each item in the race column occurs.
Assign the result to race_counts.
Display race_counts and sex_counts to verify your work, and see if you can spot any patterns.
Write a markdown cell detailing what you've learned so far, and what you think might need further examination
'''
sex_counts = {}
race_counts = {}
for row in data:
if row[5] in sex_counts:
sex_counts[row[5]] += 1
else:
sex_counts[row[5]] = 1
if row[7] in race_counts:
race_counts[row[7]] += 1
else:
race_counts[row[7]] = 1
print(sex_counts)
print(race_counts)
#Page 6
'''
Read in census.csv, and convert to a list of lists. Assign the result to the census variable.
Display census to verify your work.
'''
census = list(csv.reader(open("census.csv", 'r')))
census
# +
#Page 7
'''
Manually create a dictionary, mapping that maps each key from race_counts to the population count of the race from census.
The keys in the dictionary should be Asian/Pacific Islander, Black, Native American/Native Alaskan, Hispanic, and White.
In the case of Asian/Pacific Islander, you'll need to add the counts from census for Race Alone - Asian, and Race Alone - Native Hawaiian and Other Pacific Islander.
Create an empty dictionary, race_per_hundredk.
Loop through each key in race_counts.
Divide the value associated with the key in race_counts by the value associated with the key in mapping.
Multiply by 100000.
Assign the result to the same key in race_per_hundredk.
When you're done, race_per_hundredk should contain the rate of gun deaths per 100000 people for each racial category.
Print race_per_hundredk to verify your work.
'''
mapping = {}
newCensus = list(census[1])
mapping["White"] = int(newCensus[10])
mapping["Hispanic"] = int(newCensus[11])
mapping["Black"] = int(newCensus[12])
mapping["Native American/Native Alaskan"] = int(newCensus[13])
mapping["Asian/Pacific Islander"] = int(newCensus[14]) + int(newCensus[15])
race_per_hundredk = {}
for row in race_counts:
race_per_hundredk[row] = (race_counts[row] / mapping[row]) * 100000
print(race_per_hundredk)
# +
#Page 8:
'''
Extract the intent column using a list comprehension. The intent column is the fourth column in data.
Assign the result to intents.
Extract the race column using a list comprehension. The race column is the eighth column in data.
Assign the result to races.
Create an empty dictionary called homicide_race_counts
Use the enumerate() function to loop through each item in races. The position should be assigned to the loop variable i, and the value to the loop variable race.
Check the value at position i in intents.
If the value at position i in intents is Homicide:
If the key race doesn't exist in homicide_race_counts, create it.
Add 1 to the value associated with race in homicide_race_counts.
When you're done, homicide_race_counts should have one key for each of the racial categories in data. The associated value should be the number of gun deaths by homicide for that race.
Perform the same procedure we did in the last screen using mapping on homicide_race_counts to get from raw numbers to rates per 100000.
Display homicide_race_counts to verify your work.
Write up your findings in a markdown cell.
Write up any next steps you want to pursue with the data in a markdown cell.
'''
intents = [row[3] for row in data]
races = [row[7] for row in data]
homicide_race_counts = {}
for i, race in enumerate(races):
if intents[i] =="Homicide":
if race in homicide_race_counts:
homicide_race_counts[race] += 1
else:
homicide_race_counts[race] = 1
race_per_hundredk2 = {}
for row in homicide_race_counts:
race_per_hundredk2[row] = (homicide_race_counts[row] / mapping[row]) * 100000
print(race_per_hundredk2)
## Look at the number of blacks who are killed by a gun in a homicide
# -
| Guided Project Exploring Gun Deaths in the US.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # TFRecords files
#
# * [`tf.slim site`](https://github.com/tensorflow/models/tree/master/research/slim)
# * 위의 링크에 나온 방법으로 TensorFlow에서 제공하는 `flower` 데이터 셋을 download 받고
# * 그 후 `TFRecords` format의 데이터로 만들어보자.
# ## Downloading and converting to TFRecord format
#
# ```shell
# $ DATA_DIR=/tmp/data/flowers
# $ python download_and_convert_data.py \
# --dataset_name=flowers \
# --dataset_dir="${DATA_DIR}"
# ```
# * script 파일을 그대로 이용하여도 된다.
# * `flower` 데이터 셋을 download 한 이후 `TFRecords` format으로 변환
# +
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import math
import os
import random
import sys
import tarfile
from six.moves import urllib
import tensorflow as tf
from datasets import dataset_utils
# +
# The URL where the Flowers data can be downloaded.
_DATA_URL = 'http://download.tensorflow.org/example_images/flower_photos.tgz'
# The number of images in the validation set.
_NUM_VALIDATION = 350
# Seed for repeatability.
_RANDOM_SEED = 0
# The number of shards per dataset split.
_NUM_SHARDS = 5
# The path where the Flowers dataset is
dataset_dir = '../data/flowers'
# -
# ### downlaod `flower` dataset
if not tf.gfile.Exists(dataset_dir):
tf.gfile.MakeDirs(dataset_dir)
# downlaod and uncompress dataset.tar.gz
if not tf.gfile.Exists(dataset_dir + '/flower_photos'):
dataset_utils.download_and_uncompress_tarball(_DATA_URL, dataset_dir)
def _get_filenames_and_classes(dataset_dir):
"""Returns a list of filenames and inferred class names.
Args:
dataset_dir: A directory containing a set of subdirectories representing
class names. Each subdirectory should contain PNG or JPG encoded images.
Returns:
A list of image file paths, relative to `dataset_dir` and the list of
subdirectories, representing class names.
"""
flower_root = os.path.join(dataset_dir, 'flower_photos')
directories = []
class_names = []
for filename in os.listdir(flower_root):
path = os.path.join(flower_root, filename)
if os.path.isdir(path):
directories.append(path)
class_names.append(filename)
photo_filenames = []
for directory in directories:
for filename in os.listdir(directory):
path = os.path.join(directory, filename)
photo_filenames.append(path)
return photo_filenames, sorted(class_names)
photo_filenames, class_names = _get_filenames_and_classes(dataset_dir)
print('dataset size: {}'.format(len(photo_filenames)))
print('class_names : {}'.format(class_names))
class_names_to_ids = dict(zip(class_names, range(len(class_names))))
# print class names to ids
for key, value in class_names_to_ids.items():
print('class name: {} -- index: {}'.format(key, value))
# ### Divide into train and test
random.seed(_RANDOM_SEED)
random.shuffle(photo_filenames)
training_filenames = photo_filenames[_NUM_VALIDATION:]
validation_filenames = photo_filenames[:_NUM_VALIDATION]
print('training dataset size: {}'.format(len(training_filenames)))
print('validation dataset size: {}'.format(len(validation_filenames)))
class ImageReader(object):
"""Helper class that provides TensorFlow image coding utilities."""
def __init__(self):
# Initializes function that decodes RGB JPEG data.
self._decode_jpeg_data = tf.placeholder(dtype=tf.string)
self._decode_jpeg = tf.image.decode_jpeg(self._decode_jpeg_data, channels=3)
def read_image_dims(self, sess, image_data):
image = self.decode_jpeg(sess, image_data)
return image.shape[0], image.shape[1] # image_height, image_width
def decode_jpeg(self, sess, image_data):
image = sess.run(self._decode_jpeg,
feed_dict={self._decode_jpeg_data: image_data})
assert len(image.shape) == 3
assert image.shape[2] == 3
return image
def _get_dataset_filename(dataset_dir, split_name, shard_id):
output_filename = 'flowers_%s_%05d-of-%05d.tfrecord' % (
split_name, shard_id, _NUM_SHARDS)
return os.path.join(dataset_dir, output_filename)
# +
def int64_feature(values):
"""Returns a TF-Feature of int64s.
Args:
values: A scalar or list of values.
Returns:
A TF-Feature.
"""
if not isinstance(values, (tuple, list)):
values = [values]
return tf.train.Feature(int64_list=tf.train.Int64List(value=values))
def bytes_feature(values):
"""Returns a TF-Feature of bytes.
Args:
values: A string.
Returns:
A TF-Feature.
"""
return tf.train.Feature(bytes_list=tf.train.BytesList(value=[values]))
def float_feature(values):
"""Returns a TF-Feature of floats.
Args:
values: A scalar of list of values.
Returns:
A TF-Feature.
"""
if not isinstance(values, (tuple, list)):
values = [values]
return tf.train.Feature(float_list=tf.train.FloatList(value=values))
# -
def _convert_dataset(split_name, filenames, class_names_to_ids, dataset_dir):
"""Converts the given filenames to a TFRecord dataset.
Args:
split_name: The name of the dataset, either 'train' or 'validation'.
filenames: A list of absolute paths to png or jpg images.
class_names_to_ids: A dictionary from class names (strings) to ids
(integers).
dataset_dir: The directory where the converted datasets are stored.
"""
assert split_name in ['train', 'validation']
num_per_shard = int(math.ceil(len(filenames) / float(_NUM_SHARDS)))
# num_per_shard = int( 3320 / 5.0 ) = 664 for training dataset
# num_per_shard = int( 350 / 5.0 ) = 70 for validation dataset
with tf.Graph().as_default():
image_reader = ImageReader()
with tf.Session('') as sess:
for shard_id in range(_NUM_SHARDS):
output_filename = _get_dataset_filename(
dataset_dir, split_name, shard_id)
# output_filename -> '../data/flowers/flowers_train_00000-of-00005.tfrecord'
# '../data/flowers/flowers_train_00001-of-00005.tfrecord'
# ...
# '../data/flowers/flowers_train_00004-of-00005.tfrecord'
# step 1
with tf.python_io.TFRecordWriter(output_filename) as tfrecord_writer:
start_ndx = shard_id * num_per_shard
end_ndx = min((shard_id+1) * num_per_shard, len(filenames))
for i in range(start_ndx, end_ndx):
sys.stdout.write('\r>> Converting image %d/%d shard %d' % (
i+1, len(filenames), shard_id))
sys.stdout.flush()
# Read the filename:
# step 2
image_data = tf.gfile.FastGFile(filenames[i], 'rb').read()
height, width = image_reader.read_image_dims(sess, image_data)
class_name = os.path.basename(os.path.dirname(filenames[i]))
class_id = class_names_to_ids[class_name]
#example = dataset_utils.image_to_tfexample(
# image_data, b'jpg', height, width, class_id)
# step 3
features = tf.train.Features(feature={'image/encoded': bytes_feature(image_data),
'image/format': bytes_feature(b'jpg'),
'image/class/label': int64_feature(class_id),
'image/height': int64_feature(height),
'image/width': int64_feature(width),
})
# step 4
example = tf.train.Example(features=features)
# step 5
tfrecord_writer.write(example.SerializeToString())
sys.stdout.write('\n')
sys.stdout.flush()
# ## `TFRecords` format 파일 만드는 방법
#
# ```python
# # Step 1: create a writer to write tfrecord to that file
# tfrecord_writer = tf.python_io.TFRecordWriter(output_filename)
#
# # Step 2: get serialized data (binary values and shape of image)
# # 한 개의 example(우리가 말하는 data 하나)을 만들기 위해 필요한 정보를 모은다.
# image_data = tf.gfile.FastGFile(filenames[i], 'rb').read()
# height, width = image_reader.read_image_dims(sess, image_data)
# class_name = os.path.basename(os.path.dirname(filenames[i]))
# class_id = class_names_to_ids[class_name]
#
# # Step 3: create a tf.train.Features object
# features = tf.train.Features(feature={'image/encoded': bytes_feature(image_data),
# 'image/format': bytes_feature(image_format),
# 'image/class/label': int64_feature(class_id),
# 'image/height': int64_feature(height),
# 'image/width': int64_feature(width),
# })
#
# # Step 4: create a sample containing of features defined above
# example = tf.train.Example(features=features)
#
# # Step 5: write the sample to the tfrecord file
# tfrecord_writer.write(sample.SerializeToString())
# tfrecord_writer.close()
# ```
# First, convert the training and validation sets.
_convert_dataset('train', training_filenames, class_names_to_ids, dataset_dir)
_convert_dataset('validation', validation_filenames, class_names_to_ids, dataset_dir)
# Finally, write the labels file:
labels_to_class_names = dict(zip(range(len(class_names)), class_names))
dataset_utils.write_label_file(labels_to_class_names, dataset_dir)
# print class names to ids
for key, value in labels_to_class_names.items():
print('label index: {} -- class name: {}'.format(key, value))
def _clean_up_temporary_files(dataset_dir):
"""Removes temporary files used to create the dataset.
Args:
dataset_dir: The directory where the temporary files are stored.
"""
filename = _DATA_URL.split('/')[-1]
filepath = os.path.join(dataset_dir, filename)
tf.gfile.Remove(filepath)
# left original jpg files
#tmp_dir = os.path.join(dataset_dir, 'flower_photos')
#tf.gfile.DeleteRecursively(tmp_dir)
_clean_up_temporary_files(dataset_dir)
print('Finished converting the Flowers dataset!')
| 10.tfrecords.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
from Functions import Cleaning_Functions
from sklearn import model_selection, linear_model, neighbors, preprocessing, metrics, ensemble
fun = Cleaning_Functions()
clean = pd.read_csv("../data/clean.csv")
clean = clean.drop(["continent"], axis =1)
clean.YEAR = clean.YEAR.astype('category')
# -
def standarize_data(df):
"""
Input: a dataset
action: returns numeric column values scaled by mean and standard deviation
"""
numeric_data = df.select_dtypes(include=['float64', 'int64'])
for i in numeric_data.columns:
df[i] = (df[i] - df[i].mean())/df[i].std()
return df
market_data = fun.delete_id_columns(clean) #1
market_data, pred_market = fun.drop_response_rows_with_NAs(market_data, "Market_Orientation", "PPI_Likelihood") #2
market_data = fun.replace_NAN_with_na(market_data) #3
market_data = fun.entry_to_lowercase(market_data) #4
market_data = fun.remove_underscores_spaces(market_data) #5
market_data = fun.convert_to_categorical(market_data) #6
market_data = fun.impute_data(market_data)
market_data = standarize_data(market_data)
#Ana's fuc
def get_dummyXs_y(df, y_var):
y = df[y_var]
X = df.drop(y_var, axis = 1)
X_cat = X.select_dtypes(include = ["category", "O"])
X_num = X.select_dtypes(include=['float64', 'int64'])
X_cat_dummy = pd.get_dummies(X_cat)
newX = pd.concat([X_num, X_cat_dummy], axis = 1)
return newX, y
X, y = get_dummyXs_y(market_data, "Market_Orientation")
X_tr, X_te, y_tr, y_te = model_selection.train_test_split(X,y, test_size = 0.3, random_state = 50)
# +
def fit_predict(clf, X_tr, X_te, y_tr, y_te):
clf.fit(X_tr,y_tr)
pred = clf.predict(X_te)
mse = metrics.mean_squared_error(y_te, pred)
return "MSE: {} ".format(mse)
def tune_parameters(X_train, y_train, clf, param_dict, cv=5):
best_model = model_selection.GridSearchCV(clf, param_dict, cv=cv, scoring = "neg_mean_squared_error", n_jobs =-1, verbose=3)
best_model.fit(X_train, y_train)
print("Best Parameters: {} \n Training MSE: {} \n Parameter Index: {}".format(best_model.best_params_,best_model.best_score_,best_model.best_index_) ) # best is alpha = 0
#uses gridsearch, prints best parameters, best model, its MSE on the training set
#returns classifer
return clf
test_mse_market = []
# -
lasso_cv_mark=linear_model.LassoCV(alphas=np.arange(0.01,2,0.001))
model_cv=lasso_cv_mark.fit(X_tr,y_tr)
model_cv.alpha_
best_lassom = linear_model.Lasso(alpha = model_cv.alpha_)
best_lassom.fit(X_tr,y_tr)
y_pred_lasso = best_lassom.predict(X_te)
MSE_lasso_market = metrics.mean_squared_error(y_te,y_pred_lasso)
MSE_lasso_market
import matplotlib.pyplot as plt
alpha_set=model_cv.alphas_.T
np.mean(model_cv.mse_path_, axis=1)
plt.plot(alpha_set,np.mean(model_cv.mse_path_, axis=1))
plt.xlabel("Alphas")
plt.ylabel("MSE")
plt.title("MO: Alpha Values Vs MSE")
# +
betas_market = best_lassom.coef_
max5_index_mark = [list(abs(betas_market)).index(x) for x in np.sort(abs(betas_market))[::-1][:20]]
min5_index_mark = [list(abs(betas_market)).index(x) for x in np.sort(abs(betas_market))[:5]]
betas_market[max5_index_mark]
market = pd.DataFrame([X_tr.columns[max5_index_mark], betas_market[max5_index_mark]]).T
market
# -
feature_imp_mark = pd.DataFrame([X_tr.columns, best_lassom.coef_]).T
feature_imp_mark = feature_imp_mark.sort_values(by = 1,ascending = False).reset_index(drop = True)
feature_imp_mark.iloc[0:15,:]
clean = pd.read_csv("../data/clean.csv")
clean = clean.drop(["Country"], axis =1)
clean.YEAR = clean.YEAR.astype('category')
ppi_data = fun.delete_id_columns(clean) #1
ppi_data, pred_ppi = fun.drop_response_rows_with_NAs(ppi_data, "PPI_Likelihood", "Market_Orientation") #2
ppi_data = fun.replace_NAN_with_na(ppi_data) #3
ppi_data = fun.entry_to_lowercase(ppi_data) #4
ppi_data = fun.remove_underscores_spaces(ppi_data) #5
ppi_data = fun.convert_to_categorical(ppi_data) #6
ppi_data = fun.impute_data(ppi_data)
ppi_data = standarize_data(ppi_data)
# +
X, y = get_dummyXs_y(ppi_data, "PPI_Likelihood")
X_tr, X_te, y_tr, y_te = model_selection.train_test_split(X,y, test_size = 0.3, random_state = 2021)
lasso_cv_ppi=linear_model.LassoCV(alphas=np.arange(0.01,2,0.1))
model_cv=lasso_cv_ppi.fit(X_tr,y_tr)
model_cv.alpha_
# -
model_cv.mse_path_
import matplotlib.pyplot as plt
alpha_set=model_cv.alphas_.T
np.mean(model_cv.mse_path_, axis=1)
plt.plot(alpha_set,np.mean(model_cv.mse_path_, axis=1))
plt.xlabel("Alphas")
plt.ylabel("MSE")
plt.title("PPI:Alpha Values Vs MSE")
best_lassop = linear_model.Lasso(alpha = model_cv.alpha_)
best_lassop.fit(X_tr,y_tr)
y_pred_lasso = best_lassop.predict(X_te)
MSE_lasso_ppi = metrics.mean_squared_error(y_te,y_pred_lasso)
MSE_lasso_ppi
feature_imp_ppi = pd.DataFrame([X_tr.columns, best_lassop.coef_] ).T
feature_imp_ppi = feature_imp_ppi.sort_values(by = 1,ascending = False).reset_index(drop = True)
feature_imp_ppi.iloc[0:15,:]
combined_imp = pd.concat([feature_imp_ppi, feature_imp_mark], axis = 1).iloc[0:20,:]
combined_imp.columns = ["PPI Likelihood","Abs Coefficent","Market Orientation","Abs Coefficent"]
combined_imp
# +
betas_ppi = best_lassop.coef_
max5_index_ppi = [list(abs(betas_ppi)).index(x) for x in np.sort(abs(betas_ppi))[::-1][:20]]
min5_index_ppi = [list(abs(betas_ppi)).index(x) for x in np.sort(abs(betas_ppi))[:5]]
betas_ppi[max5_index_ppi]
ppi = pd.DataFrame([X_tr.columns[max5_index_ppi], betas_ppi[max5_index_ppi]]).T
combine = pd.concat([ppi, market], axis=1)
combine.columns = ["PPI Likelihood","Coefficent","Market Orientation","Coefficent"]
combine
| finalized_code/Lasso_Reg.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## In this file we will evaluate whether, the extracted indexes (rep frames) are enough to use with deep learning in terms of training and evaluation
#
"""
Steps:
1. Create the indexes
2. 1 instance of the network is trained using the whole data with augmentation
3. 1 instance of the network is trained using rep frame data with augmentation
4. Compare training speed
5. Compare training loss
6. Compare mAP
"""
# +
import os
os.getcwd()
# %pylab inline
# %load_ext autoreload
# %autoreload 2
import os
import sys
sys.argv=['']
sys.path.append('../../')
import numpy as np
#import utils.helpers as helpers
import utils as helpers
from loaders.uadetrac_loader import UADetracLoader
from eva_storage.preprocessingModule import PreprocessingModule
from eva_storage.UNet import UNet
from eva_storage.clusterModule import ClusterModule
from filters.minimum_filter import FilterMinimum
from eva_storage.evaluation.evaluate_compression import *
# -
# load the data
loader = UADetracLoader()
images = loader.load_cached_images()
labels = loader.load_cached_labels()
video_start_indices = loader.get_video_start_indices()
# run background subtraction
pm = PreprocessingModule()
seg_images = pm.run(images,video_start_indices)
print(seg_images.shape)
# train the main network
unet = UNet()
unet.train(images, seg_images, epoch = 100)
# Generate the compressed / segmented images
unet_compressed_images, unet_segmented_images = unet.execute()
# Create clusters
cm = ClusterModule()
image_cluster_labels = cm.run(unet_compressed_images)
# +
# Generate binary labels
## within labels['vehicle'] there are ['car', 'others', 'van', 'bus']
car_labels = helpers.generateBinaryLabels(labels['vehicle'])
other_labels = helpers.generateBinaryLabels(labels['vehicle'], label_of_interest = 'others')
van_labels = helpers.generateBinaryLabels(labels['vehicle'], 'van')
bus_labels = helpers.generateBinaryLabels(labels['vehicle'], 'bus')
# -
| eva_storage/src/evaluation/evaluate_rep/.ipynb_checkpoints/eva_vs_nop__dl-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # pytorch で DL を行う最小限セット
# reference
# https://qiita.com/fukuit/items/215ef75113d97560e599#comments
# +
import torch.nn as nn
import torch.nn.functional as F
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
self.conv1 = nn.Conv2d(1, 32, 3)
self.conv2 = nn.Conv2d(32, 64, 3)
self.pool = nn.MaxPool2d(2, 2)
self.dropout1 = nn.Dropout2d()
self.fc1 = nn.Linear(9216, 128)
self.dropout2 = nn.Dropout2d()
self.fc2 = nn.Linear(128, 10)
def forward(self, x):
x = F.relu(self.conv1(x))
x = self.pool(F.relu(self.conv2(x)))
x = self.dropout1(x)
x = x.view(-1, 9216)
x = F.relu(self.fc1(x))
x = self.dropout2(x)
x = self.fc2(x)
return x
# +
'''
PyTorch MNIST sample
'''
import argparse
import time
import numpy as np
import torch
import torch.nn as nn
import torch.nn.functional as F
from torch.utils.data import DataLoader
import torchvision
import torchvision.transforms as transforms
from torchvision.datasets import MNIST
import torch.optim as optim
from net import Net
# +
def parser():
'''
argument
'''
parser = argparse.ArgumentParser(description='PyTorch MNIST')
parser.add_argument('--epochs', '-e', type=int, default=2,
help='number of epochs to train (default: 2)')
parser.add_argument('--lr', '-l', type=float, default=0.01,
help='learning rate (default: 0.01)')
args = parser.parse_args()
return args
# +
def main():
'''
main
'''
args = parser()
args.epochs=2
args.lr=0.01
transform = transforms.Compose(
[transforms.ToTensor(),
transforms.Normalize((0.5, ), (0.5, ))])
trainset = MNIST(root='./data',
train=True,
download=True,
transform=transform)
testset = MNIST(root='./data',
train=False,
download=True,
transform=transform)
trainloader = DataLoader(trainset,
batch_size=100,
shuffle=True,
num_workers=2)
testloader = DataLoader(testset,
batch_size=100,
shuffle=False,
num_workers=2)
classes = tuple(np.linspace(0, 9, 10, dtype=np.uint8))
# model
net = Net()
# define loss function and optimier
criterion = nn.CrossEntropyLoss()
optimizer = optim.SGD(net.parameters(),
lr=args.lr, momentum=0.99, nesterov=True)
# train
for epoch in range(args.epochs):
running_loss = 0.0
for i, (inputs, labels) in enumerate(trainloader, 0):
# zero the parameter gradients
optimizer.zero_grad()
# forward + backward + optimize
outputs = net(inputs)
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
# print statistics
running_loss += loss.item()
if i % 100 == 99:
print('[{:d}, {:5d}] loss: {:.3f}'
.format(epoch+1, i+1, running_loss/100))
running_loss = 0.0
print('Finished Training')
# test
correct = 0
total = 0
with torch.no_grad():
for (images, labels) in testloader:
outputs = net(images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy: {:.2f} %'.format(100 * float(correct/total)))
# -
#TODO notebookからだと、引数に余分なものを指定してエラーになる。
if __name__ == '__main__':
start_time = time.time()
main()
print('elapsed time: {:.3f} [sec]'.format(time.time() - start_time))
main()
| notebook/basic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Sparkify Project Workspace
import os
os.environ["PYSPARK_PYTHON"] = "python3"
os.environ["PYSPARK_DRIVER_PYTHON"] = "python3"
os.environ["JAVA_HOME"] = "/Library/java/JavaVirtualMachines/adoptopenjdk-8.jdk/contents/Home/"
import findspark
findspark.init()
# +
# import libraries
from pyspark.sql import SparkSession
from pyspark.sql import functions as F
from pyspark.sql import types as T
from pyspark.sql import Window
import datetime
from pyspark.ml import Pipeline
from pyspark.ml.feature import (
StringIndexer, OneHotEncoderEstimator, MinMaxScaler, VectorAssembler)
from pyspark.ml.classification import (
DecisionTreeClassifier, RandomForestClassifier, GBTClassifier, LogisticRegression)
from pyspark.ml.tuning import ParamGridBuilder, CrossValidator
from pyspark.ml.evaluation import BinaryClassificationEvaluator, MulticlassClassificationEvaluator
from sklearn.metrics import classification_report, confusion_matrix
from sklearn.metrics import roc_curve
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# -
# Create spark session
spark = SparkSession \
.builder \
.appName("Sparkify") \
.getOrCreate()
# # Load and Clean Dataset
# Load and clean the `mini_sparkify_event_data.json` dataset, checking for invalid or missing data - for example, records without user-ids or session-ids.
# Read in sparkify dataset
event_data = "mini_sparkify_event_data.json"
df = spark.read.json(event_data)
df.head()
df.printSchema()
# number of records
df.count()
# check number of missing values in each column
missing_values = [F.count(F.when(F.col(c).isNull(), c)).alias(c)
for c in df.columns]
df.select(missing_values).show()
# Users without registration are not valid for churn analysis, and thus will be dropped.
df_valid = df.dropna(subset=['registration'])
# check number of missing values for each column again
missing_values = [F.count(F.when(F.col(c).isNull(), c)).alias(c)
for c in df_valid.columns]
df_valid.select(missing_values).show()
# `artist`, `length`, and `song` will be null when a user is visiting pages other than `NextSong`.
df_valid.filter(F.col('artist').isNotNull()).select(
'page').dropDuplicates().show()
# # Exploratory Data Analysis
# EDA on a small subset of the data with basic manipulations within Spark.
# number of users
num_users = df_valid.select("userId").dropDuplicates().count()
print(num_users)
# number of artists
df_valid.select("artist").dropDuplicates().count()
# auth
df_valid.select("auth").dropDuplicates().show()
# itemInSession
df_valid.select("itemInSession").describe().show()
# length
df_valid.select("length").describe().show()
# histogram for length
df_valid.select("length").toPandas().hist(bins=100);
# level
df.select("level").dropDuplicates().show()
# location
df.select("location").dropDuplicates().show(10, False)
# method
df.select("method").dropDuplicates().show()
# page
df.select("page").dropDuplicates().show(22, False)
# registration
df.select("registration").show(5)
# songs
df.select("song").show(5, False)
# number of songs
df.select("song").drop_duplicates().count()
# status
df.select("status").drop_duplicates().show()
# ts
df.select("ts").show(5)
# userAgent
df_valid.select("userAgent").drop_duplicates().show(10, False)
# ## Define Churn
#
# Define churn events as`Cancellation Confirmation` events.
# Create`churn_event` and `churn_user` columns.
# +
# define churn event
flag_churn_event = F.udf(
lambda x: 1 if x == 'Cancellation Confirmation' else 0, T.IntegerType())
# create churn event column
df_valid = df_valid.withColumn('churn_event', flag_churn_event('page'))
# create churn user column
df_valid = df_valid.withColumn('churn_user', F.max(
'churn_event').over(Window.partitionBy('userId')))
# -
df_valid.groupBy('churn_user').count().show()
44864/(44864 + 233290) * 100
# ## Explore Data
# More exploratory data analysis to observe the behavior of users who stayed vs users who churned. The relations between users' attributes such as gender, level, location, and churn events are explored.
# We also explore aggregates on these two groups of users, observing how much of a specific action they experienced such as number of songs played or how many times they visited home page per week.
# number of churn and stayed users
num_churn_users = df_valid.filter(df_valid['churn_user'] == '1').select(
F.countDistinct('userId')).collect()[0][0]
num_stayed_users = num_users - num_churn_users
print("number of users that churn:", num_churn_users)
print("number of users that stayed:", num_stayed_users)
# +
d = {'churns': [num_stayed_users, num_churn_users]}
df_churns = pd.DataFrame(data=d, index=['Stayed', 'Churn'])
ax = df_churns.plot.bar(legend=False)
ax.grid(axis='y')
ax.set_ylabel('Count', fontsize=12)
ax.set_title('')
plt.xticks(rotation=0);
plt.savefig('churns')
# -
# create datetime columns for registration and timestamp
df_valid = df_valid.withColumn(
'registration_dt', (F.col('registration')/1000).cast(T.TimestampType()))
df_valid = df_valid.withColumn(
'dt', (F.col('ts')/1000).cast(T.TimestampType()))
# example of churn by `Cancellation Confirmation`
df_valid.select('userId', 'firstName', 'page', 'dt', 'level').where((df_valid['userId'] == 73) & (
df_valid['page'] == 'Cancellation Confirmation')).show(5, False)
# example of Submit Downgrade`
df_valid.select('userId', 'firstName', 'page', 'dt', 'level').where((df_valid['userId'] == 39) & (
df_valid['page'] == 'Submit Downgrade')).show(5, False)
# example of churn by `Submit Downgrade` followed by `Cancellation Confirmation`
df_valid.select('userId', 'firstName', 'page', 'dt', 'level').where((df_valid['userId'] == 103) & (
(df_valid['page'] == 'Submit Downgrade') | (df_valid['page'] == 'Cancellation Confirmation'))).show(5, False)
# ### Gender
num_females = df_valid.filter(df_valid['gender'] == 'F').select(
F.countDistinct('userId')).collect()[0][0]
num_males = num_users - num_females
print("number of females:", num_females)
print("number of males:", num_males)
df_gender = df_valid.groupby('gender').sum('churn_event').toPandas()
df_gender['count'] = [num_females, num_males]
df_gender['percent churn'] = df_gender['sum(churn_event)'] / \
df_gender['count']*100
df_gender.columns = ['gender', 'num_churns', 'count', 'percent churn']
df_gender.set_index('gender')
ax = df_gender.plot.bar(x='gender', y='percent churn', legend=False)
ax.grid(axis='y')
ax.set_xlabel('')
ax.set_ylabel('Percent churn', fontsize=12)
ax.set_title('Gender')
plt.xticks(rotation=0);
plt.savefig('gender')
# ### Item In Session
df_item_churn = df_valid.filter(df_valid['churn_user'] == 1).select('itemInSession').toPandas()
df_item_stayed = df_valid.filter(df_valid['churn_user'] == 0).select('itemInSession').toPandas()
df_item = pd.concat([df_item_stayed, df_item_churn], axis=1, ignore_index=True)
df_item.describe().transpose()
# ### Level
num_free = df_valid.filter(df_valid['level'] == 'free').select(
F.countDistinct('userId')).collect()[0][0]
num_paid = df_valid.filter(df_valid['level'] == 'paid').select(
F.countDistinct('userId')).collect()[0][0]
df_level = df_valid.groupby('level').sum('churn_event').toPandas()
df_level['count'] = [num_free, num_paid]
df_level['percent churn'] = df_level['sum(churn_event)'] / \
df_level['count'] * 100
df_level.columns = ['level', 'num_churns', 'count', 'percent churn']
df_level.set_index('level')
ax = df_level.plot.bar(x='level', y='percent churn', legend=False)
ax.grid(axis='y')
ax.set_ylabel('Percent churn', fontsize=12)
ax.set_xlabel('')
ax.set_title('Level')
plt.xticks(rotation=0);
plt.savefig('level')
# ### Location
df_valid.select("location").show(10, False)
# +
# get state from location
get_state = F.udf(lambda x: x[-2:])
# create state column
df_valid = df_valid.withColumn("state", get_state(df_valid['location']))
# -
df_state_counts = df_valid.select(
'userId', 'state').dropDuplicates().groupby('state').count().toPandas()
ax = df_state_counts.plot.bar(x='state', y='count',
figsize=(12, 6))
ax.set_ylabel('number of distinct users', fontsize=12);
df_state_churns = df_valid.groupby('state').sum('churn_event').toPandas()
df_state_churns.columns = ['state', 'num_churns']
# +
ax = df_state_churns.plot.bar(x='state', y='num_churns', figsize=(12, 6), legend=False)
ax.grid(axis='y')
ax.set_xlabel('')
ax.set_ylabel('Churn count', fontsize=16);
#ax.set_title('State', fontsize=18)
plt.xticks(fontsize=14)
plt.yticks(fontsize=14)
plt.tight_layout()
plt.savefig('state')
# -
df_state_churns['percent churn'] = df_state_churns['num_churns'] / \
df_state_counts['count'] * 100
df_state_churns.plot.bar(x='state', y='percent churn', figsize=(12, 6));
# ### Registration
# find earliest and latest registration dates
start_reg_date = df_valid.agg({'registration_dt': 'min'}).collect()
end_reg_date = df_valid.agg({'registration_dt': 'max'}).collect()
print('registration start date:', str(start_reg_date[0][0]))
print('registration end date:', str(end_reg_date[0][0]))
# find earliest and latest dates (from timestamp)
start_date = df_valid.agg({'dt': 'min'}).collect()
end_date = df_valid.agg({'dt': 'max'}).collect()
print('start date:', str(start_date[0][0]))
print('end date:', str(end_date[0][0]))
# number of days since registration
df_valid = df_valid.withColumn("days_since_reg", F.max(
F.datediff('dt', 'registration_dt')).over(Window.partitionBy('userId')))
df_valid.select('days_since_reg').describe().show()
df_valid.select('days_since_reg').toPandas().hist(bins=25)
df_days_stayed = df_valid.filter(
df_valid['churn_user'] == 0).select('days_since_reg').toPandas()
df_days_churn = df_valid.filter(
df_valid['churn_user'] == 1).select('days_since_reg').toPandas()
df_reg_days = pd.concat([df_days_stayed, df_days_churn], axis=1, ignore_index=True)
df_reg_days.columns = ['stayed', 'churn']
df_reg_days.describe().transpose()
ax = df_reg_days.boxplot()
ax.set_ylabel('Number of days', fontsize=12)
ax.set_title('Days since registration');
plt.savefig('days_since_registration')
# ### Timestamp
get_hour = F.udf(lambda x: datetime.datetime.fromtimestamp(
x / 1000.0).hour, T.IntegerType())
get_day = F.udf(lambda x: datetime.datetime.fromtimestamp(
x / 1000.0).day, T.IntegerType())
get_month = F.udf(lambda x: datetime.datetime.fromtimestamp(
x / 1000.0).month, T.IntegerType())
df_valid = df_valid.withColumn('hour', get_hour(F.col('ts')))\
.withColumn('day', get_day(F.col('ts')))\
.withColumn('month', get_month(F.col('ts')))
# +
ax = df_valid.groupBy('hour').sum('churn_event').sort('hour').toPandas(
).plot.bar(x='hour', y='sum(churn_event)', figsize=(12, 6), legend=False)
ax.grid(axis='y')
ax.set_xlabel('Hour', fontsize=16)
ax.set_ylabel('Count', fontsize=16)
ax.set_title('Hourly churn events', fontsize=18)
plt.xticks(fontsize=16)
plt.yticks(fontsize=16)
plt.tight_layout()
plt.savefig('hourly_churn_events')
# +
ax = df_valid.groupBy('day').sum('churn_event').sort('day').toPandas(
).plot.bar(x='day', y='sum(churn_event)', figsize=(12, 6), legend=False)
ax.grid(axis='y')
ax.set_xlabel('Day', fontsize=16)
ax.set_ylabel('Count', fontsize=16)
ax.set_title('Daily churn events', fontsize=18)
plt.xticks(fontsize=16)
plt.yticks(fontsize=16)
plt.tight_layout()
plt.savefig('daily_churn_events')
# -
ax = df_valid.groupBy('month').sum('churn_event').sort(
'month').toPandas().plot.bar(x='month', y='sum(churn_event)')
ax.grid(axis='y')
ax.set_xlabel('Month', fontsize=12)
ax.set_ylabel('Count', fontsize=12)
ax.set_title('Monthly churn events');
plt.savefig('montly_churn_events')
# ### Song
# +
# time window to find number of songs played in a week
def days(x):
"""Return time in seconds for the input time in days."""
time_in_seconds = x * 86400
return time_in_seconds
windowval = (
Window.partitionBy("userId")
.orderBy(F.col("ts")/1000)
.rangeBetween(-days(7), 0)
)
# -
# number of songs played in a week
df_valid = df_valid.withColumn("num_songs", F.count("song").over(windowval))
# check
df_valid.select('userId', 'dt', 'song', 'num_songs', 'churn_event',
'churn_user').where(df_valid['userId'] == 32).show(200)
def make_boxplot(df, col_name):
"""Return boxplot of stayed and churn for the attribute col_name of the dataframe df."""
df_stayed = df.filter(df['churn_user'] == 0).select(
col_name).toPandas()
df_churn = df.filter(df['churn_user'] == 1).select(
col_name).toPandas()
df = pd.concat([df_stayed, df_churn], axis=1, ignore_index=True)
df.columns = ['stayed', 'churn']
statistics = df.describe().transpose()
ax = df.boxplot()
ax.set_title(col_name + ' per week', fontsize=12)
ax.set_ylabel('Count', fontsize=12)
return statistics, ax
make_boxplot(df_valid, 'num_songs')
# ### Artist
# number of distinct artists played in a week
df_valid = df_valid.withColumn("num_artists", F.size(
F.collect_set("artist").over(windowval)))
make_boxplot(df_valid, 'num_artists')
# ### Session
# number of sessions in a week
df_valid = df_valid.withColumn(
'num_sessions', F.count('sessionId').over(windowval))
make_boxplot(df_valid, 'num_sessions')
# ### Status
df_valid.groupby('status').count().show()
# flag the status 'redirect' and 'not found'
flag_redirect = F.udf(lambda x: 1 if x == 307 else 0, T.IntegerType())
flag_not_found = F.udf(lambda x: 1 if x == 404 else 0, T.IntegerType())
# create redirect and page_not_found columns
df_valid = df_valid.withColumn(
"redirect", flag_redirect(df_valid['status']))
df_valid = df_valid.withColumn(
"page_not_found", flag_not_found(df_valid['status']))
df_valid.groupBy('churn_user').sum('redirect').show()
df_valid.groupBy('churn_user').sum('page_not_found').show()
df_valid.filter(df_valid['status'] == '307').select(
F.sum("churn_event")).show()
df_valid.filter(df_valid['status'] == '200').select(
F.sum("churn_event")).show()
# ### Page
df_valid.groupby('page').count().show(22, False)
# +
# create aggregate features from the column 'page'
pages = ['Thumbs Down', 'Thumbs Up', 'Home', 'Roll Advert',
'Logout', 'Add to Playlist', 'Add Friend', 'Error']
for page in pages:
flag_page = F.udf(lambda x: 1 if x == page else 0, T.IntegerType())
df_valid = df_valid.withColumn('flag', flag_page('page'))
df_valid = df_valid.withColumn(
'num_' + page, F.sum('flag').over(windowval)).drop('flag')
# -
make_boxplot(df_valid, 'num_Thumbs Down')
plt.savefig('num_Thumbs_Down')
make_boxplot(df_valid, 'num_Thumbs Up')
make_boxplot(df_valid, 'num_Home')
make_boxplot(df_valid, 'num_Roll Advert')
plt.savefig('num_Roll_Advert')
make_boxplot(df_valid, 'num_Logout')
make_boxplot(df_valid, 'num_Add to Playlist')
make_boxplot(df_valid, 'num_Add Friend')
plt.savefig('num_Add_Friend')
make_boxplot(df_valid, 'num_Error')
df_temp = df_valid.select(
'churn_user',
'itemInSession',
'length',
'days_since_reg',
'hour',
'day',
'month',
'num_songs',
'num_artists',
'num_sessions',
'num_Thumbs Down',
'num_Thumbs Up',
'num_Home',
'num_Roll Advert',
'num_Logout',
'num_Add to Playlist',
'num_Add Friend',
'num_Error'
).toPandas()
plt.figure(figsize=(15, 12))
sns.heatmap(df_temp.corr().abs(), annot=True, cmap='coolwarm');
#plt.ylim(12, 0)
# Highly correlated features will be dropped to avoid multicollinearity.
df_temp_2 = df_valid.select(
'churn_user',
'itemInSession',
'days_since_reg',
'hour',
'day',
'month',
'num_Thumbs Down',
'num_Roll Advert',
'num_Add Friend',
'num_Error'
).toPandas()
# +
plt.figure(figsize=(15, 12))
ax = sns.heatmap(df_temp_2.corr().abs(), annot=True, cmap='coolwarm');
plt.xticks(fontsize=16)
plt.yticks(fontsize=16)
plt.tight_layout()
plt.savefig('heatmap')
# -
# # Feature Engineering
# Data preprocessing and preparing machine learning pipelines for logistic regression, random forest and gradient boosted trees models. Stages for the pipelines include indexing and one-hot encoding categorical columns, transforming columns into a single vector column, and scaling the values of the features into the range from 0 to 1.
# +
def clean_data(df):
"""Return valid data by dropping missing values in registration."""
df_clean = df.dropna(subset=['registration']).na.fill(
0, subset=['length'])
return df_clean
def is_churn(df):
"""Return dataframe with labels for churn users."""
flag_churn_event = F.udf(
lambda x: 1 if x == 'Cancellation Confirmation' else 0, T.IntegerType())
df_new = df.withColumn(
'churn_event', flag_churn_event(F.col('page')))
df_new = df_new.withColumn('label', F.max(
'churn_event').over(Window.partitionBy('userId')))
return df_new
def registration_days(df):
"""Return the number of days since registration."""
df_new = df.withColumn('registration date', (F.col(
'registration')/1000).cast(T.TimestampType()))
df_new = df_new.withColumn(
'date', (F.col('ts')/1000).cast(T.TimestampType()))
df_new = df_new.withColumn('days_since_registration', (F.datediff(
'date', 'registration date'))).drop('date', 'registration date')
return df_new
def timestamp(df):
"""Return dataframe with hour, day, and month."""
get_hour = F.udf(lambda x: datetime.datetime.fromtimestamp(
x / 1000.0).hour, T.IntegerType())
get_day = F.udf(lambda x: datetime.datetime.fromtimestamp(
x / 1000.0).day, T.IntegerType())
get_month = F.udf(lambda x: datetime.datetime.fromtimestamp(
x / 1000.0).month, T.IntegerType())
df_new = df.withColumn('hour', get_hour(F.col('ts'))).withColumn(
'day', get_day(F.col('ts'))).withColumn('month', get_month(F.col('ts')))
return df_new
# time window to find aggregate values per day
def days(x):
"""Return time in seconds for the input time in days."""
time_in_seconds = x * 86400
return time_in_seconds
windowval = (
Window.partitionBy("userId")
.orderBy(F.col("ts")/1000)
.rangeBetween(-days(7), 0)
)
# create aggregate features from the column 'page'
pages = ['Thumbs Down', 'Roll Advert', 'Add Friend', 'Error']
def aggregate_features(df):
"""Return selected aggregate features from page."""
df_new = df
for page in pages:
flag_page = F.udf(lambda x: 1 if x ==
page else 0, T.IntegerType())
df_new = df_new.withColumn('flag', flag_page('page'))
df_new = df_new.withColumn(
'num_' + page, F.sum('flag').over(windowval)).drop('flag')
return df_new
def retrive_state(df):
"""Return dataframe with states retrived from locations."""
df_new = df
get_state = F.udf(lambda x: x[-2:])
df_new = df_new.withColumn("state", get_state(
df_new['location'])).drop('location')
return df_new
ignore_list = ['artist',
'auth',
'firstName',
'length',
'lastName',
'method',
'page',
'registration',
'sessionId',
'song',
'ts',
'userAgent',
'userId',
'churn_event',
]
def ignore(df, ignore_list):
"""Remove unnecessary feature columns."""
df_new = df.drop(*ignore_list)
return df_new
def preprocess_data(df):
"""
1. Remove invalid data.
2. Add churn event and churn user labels.
3. Add number of days since registration.
4. Add hour, day, and month.
5. Add aggregate features.
6. Get state from location.
7. Remove unnecessary feature columns.
"""
df_clean = clean_data(df)
df_churn = is_churn(df_clean)
df_reg_days = registration_days(df_churn)
df_timestamp = timestamp(df_reg_days)
df_aggregate = aggregate_features(df_timestamp)
df_state = retrive_state(df_aggregate)
df_processed = ignore(df_state, ignore_list)
return df_processed
df_processed = preprocess_data(df).cache()
df.unpersist()
# Prepare stages for ML Pipeline
stages = []
# Index and one-hot encode categorical columns
cat_columns = ['gender', 'level', 'status', 'state', 'month']
for column in cat_columns:
indexer = StringIndexer(inputCol=column, outputCol=column+'_index')
encoder = OneHotEncoderEstimator(
inputCols=[indexer.getOutputCol()], outputCols=[column + '_vect'])
stages += [indexer, encoder]
num_columns = [
'itemInSession',
'days_since_registration',
'hour',
'day',
'num_Thumbs Down',
'num_Roll Advert',
'num_Add Friend',
'num_Error'
]
# Transform columns into a single vector column.
vect_assembler_inputs = [c + '_vect' for c in cat_columns] + num_columns
assembler = VectorAssembler(
inputCols=vect_assembler_inputs, outputCol="features")
stages += [assembler]
# Rescale each feature into the range from 0 to 1.
scaler = MinMaxScaler(inputCol="features", outputCol="scaled_features")
stages += [scaler]
# Instantiate classifiers
lr = LogisticRegression(labelCol='label', featuresCol='scaled_features')
rfc = RandomForestClassifier(
labelCol='label', featuresCol='scaled_features')
gbt = GBTClassifier(labelCol='label', featuresCol='scaled_features')
# Set stages for classifiers
stages_lr = stages + [lr]
stages_rfc = stages + [rfc]
stages_gbt = stages + [gbt]
# -
print('df_processed shape: {}, {}'.format(df_processed.count(), len(df_processed.columns)))
df_processed.columns
# # Modeling
# Split the full dataset into train and test datasets, set stages for pipelines and train models.
# Split the full dataset into train and test sets
train, test = df_processed.randomSplit([0.8, 0.2], seed=42)
# set pipelines
pipeline_lr = Pipeline().setStages(stages_lr)
pipeline_rfc = Pipeline().setStages(stages_rfc)
pipeline_gbt = Pipeline().setStages(stages_gbt)
# train models
model_lr = pipeline_lr.fit(train)
model_rfc = pipeline_rfc.fit(train)
model_gbt = pipeline_gbt.fit(train)
# # Model Evaluation
# +
# Instantiate evaluators
f1_eval = MulticlassClassificationEvaluator(
labelCol='label', metricName='f1') # F1 score evaluator
AUC_eval = BinaryClassificationEvaluator(
labelCol='label', metricName="areaUnderROC") # AUC evaluator
# -
def evaluate_model(model):
""" Return f1 and AUC scores of the model evaluated with test data."""
preds = model.transform(test)
f1 = f1_eval.evaluate(preds)
AUC = AUC_eval.evaluate(preds)
return f1, AUC
lr_f1, lr_AUC = evaluate_model(model_lr)
print('Logistic Regression')
print('F1 =', lr_f1)
print('AUC =', lr_AUC)
rfc_f1, rfc_AUC = evaluate_model(model_rfc)
print('Random Forest Classifier')
print('F1 =', rfc_f1)
print('AUC =', rfc_AUC)
gbt_f1, gbt_AUC = evaluate_model(model_gbt)
print('Gradient-boosted Tree Classifier')
print('F1 =', gbt_f1)
print('AUC =', gbt_AUC)
# +
# create a dataframe for metrics
d = {'Classifier': ['LogisticRegression', 'RandomForestClassifier','GBTClassifier'],
'F1': [lr_f1, rfc_f1, gbt_f1],
'AUC': [lr_AUC, rfc_AUC, gbt_AUC]}
df_metrics = pd.DataFrame(data=d)
# -
df_metrics
# ### Model Selection
# Hyperparameter tuning for GBTClassifier using three-fold cross validation.
# Create param_grid for Cross Validation
param_grid = (ParamGridBuilder()
.addGrid(gbt.stepSize, [0.1, 0.5])
.addGrid(gbt.maxIter, [20, 32])
.build())
# +
pipeline = Pipeline().setStages(stages)
# Fit the pipeline with training data and transform train and test datasets
train_prep = pipeline.fit(train).transform(train).cache()
test_prep =pipeline.fit(train).transform(test).cache()
# +
cv = CrossValidator(
estimator=gbt,
estimatorParamMaps=param_grid,
evaluator=f1_eval,
numFolds=2)
cv_model = cv.fit(train_prep)
# -
# extract best model's parameters
cv_model.bestModel.extractParamMap()
cv_preds = cv_model.transform(test_prep)
cv_f1 = f1_eval.evaluate(cv_preds)
print('best_model F1 score:', cv_f1)
# #### Classification Report
cv_preds_pd = cv_preds.toPandas()
labels = cv_preds_pd['label']
predictions = cv_preds_pd['prediction']
probabilities = cv_preds_pd['probability'].apply(lambda x: x[1]).values
print(classification_report(labels, predictions))
confusion_matrix(labels, predictions)
# +
# Create ROC curve
fpr, tpr, _ = roc_curve(labels, probabilities)
plt.plot(fpr, tpr)
plt.xlabel('False positive rate')
plt.ylabel('True positive rate')
plt.title('ROC curve');
| Sparkify.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Particle In a Box
#
# The Schödinger equation can be solved analytically for simple systems, but more complex systems require numerical methods to compute approximate solutions. In this notebook, we will compare the analytical solution for a single-dimensional particle-in-a-box problem with the solution obtained by the variational method.
# ## Analytical Solution
#
# The derivation of the analytical solution for a 1-D particle in a box problem can be found in section 4.6 of Oxtoby (7th edition). See how the solution changes as you modify the quantum number $n$.
# +
import numpy as np
import matplotlib.pyplot as plt
L = 1 # box length
n = 1 # quantum number
x = np.linspace(0, L, 1000) # x position
psi_true = np.sqrt(2 / L) * np.sin(n * np.pi / L * x)
plt.figure(figsize=(10,3))
plt.subplot(1, 2, 1)
plt.plot(x, psi_true)
plt.title("Wavefunction")
plt.subplot(1, 2, 2)
plt.plot(psi_true ** 2)
plt.title("Probability Density")
plt.tight_layout()
plt.show()
# -
# ## Numerical Solution
#
# The variational method allows us to estimate the wavefunction by approximating it with a parameterized function that satisfies the boundary conditions, and then minimizing the total energy. You can learn more about the variational method [here](https://en.wikipedia.org/wiki/Variational_method_(quantum_mechanics)).
# +
# approximate the n=2 quantum state
psi_true = np.sqrt(2 / L) * np.sin(1 * np.pi / L * x)
# polynomial coefficients (highest order first)
p = np.array([0, 0, 0, 1, 0]) # <-- see what happens when you change these
# approximate wavefunction as a polynomial in x(1-x)
psi_trial = np.polyval(p, x*(1-x))
psi_trial *= np.max(psi_true) / np.max(psi_trial) # normalization
# compute energy from variational theorem
first_deriv = np.diff(psi_trial) / 0.001
second_deriv = np.diff(first_deriv) / 0.001
H = -0.5 / (2*np.pi)**2 * second_deriv
var_num = np.sum(psi_trial[:-2] * H * psi_trial[:-2])
var_denom = np.sum(psi_trial[:-2] * psi_trial[:-2])
E_trial = var_num / var_denom
print("Trial Energy:", E_trial)
plt.figure(figsize=(10,3))
plt.subplot(1, 2, 1)
plt.plot(x, psi_true, color='blue')
plt.gca().twinx()
plt.plot(x, psi_trial, color='red')
plt.title("Wavefunction")
plt.subplot(1, 2, 2)
plt.plot(psi_true ** 2, color='blue')
plt.gca().twinx()
plt.plot(psi_trial ** 2, c='red')
plt.title("Probability Density")
plt.tight_layout()
plt.show()
plt.figure(figsize=(10,3))
plt.subplot(1, 2, 1)
plt.plot(x, psi_trial - psi_true, color='blue')
plt.title("Wavefunction Error")
plt.subplot(1, 2, 2)
plt.plot(psi_trial ** 2 - psi_true ** 2, color='blue')
plt.title("Probability Density Error")
plt.tight_layout()
plt.show()
| hw7-particle-in-a-box.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Visualizing data with matplotlib
# Somtimes graphs provide the best way to visualize data
#
# The **matplotlib** library allows you to draw graphs to help with visualization
#
# If we want to visualize data, we will need to load some data into a DataFrame
import pandas as pd
# Load our data from the csv file
delays_df = pd.read_csv('Data/Lots_of_flight_data.csv')
# In order to display plots we need to import the **matplotlib** library
import matplotlib.pyplot as plt
# A common plot used in data science is the scatter plot for checking the relationship between two columns
# If you see dots scattered everywhere, there is no correlation between the two columns
# If you see somethign resembling a line, there is a correlation between the two columns
#
# You can use the plot method of the DataFrame to draw the scatter plot
# * kind - the type of graph to draw
# * x - value to plot as x
# * y - value to plot as y
# * color - color to use for the graph points
# * alpha - opacity - useful to show density of points in a scatter plot
# * title - title of the graph
#Check if there is a relationship between the distance of a flight and how late the flight arrives
delays_df.plot(
kind='scatter',
x='DISTANCE',
y='ARR_DELAY',
color='blue',
alpha=0.3,
title='Correlation of arrival and distance'
)
plt.show()
#Check if there is a relationship between the how late the flight leaves and how late the flight arrives
delays_df.plot(
kind='scatter',
x='DEP_DELAY',
y='ARR_DELAY',
color='blue',
alpha=0.3,
title='Correlation of arrival and departure delay'
)
plt.show()
# The scatter plot allows us to see there is no correlation between distance and arrival delay but there is a strong correlation between departure delay and arrival delay.
#
| Intro to Python for ML/15 - Visualizing data with Matplotlib/15 - Visualizing correlations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="N0_cP7SYTczU"
# ### **Mari Mengenal Python**
# + [markdown] colab_type="text" id="HHabukq-T0W_"
# Program pertama: "Hello World"
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 883, "status": "ok", "timestamp": 1595149352799, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiVlpSEE7CTptyyLOF0HBAyiJig8SaCaXCH-pm1kw=s64", "userId": "07646999100105069157"}, "user_tz": -420} id="4fkjTE7qTWyB" outputId="8a721e8e-dbf7-4497-b2fb-4ab132e98f11"
print("Hello World!")
# + [markdown] colab_type="text" id="6TU6j_QMT5-h"
# Program Pertamaku
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" executionInfo={"elapsed": 862, "status": "ok", "timestamp": 1595149368741, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiVlpSEE7CTptyyLOF0HBAyiJig8SaCaXCH-pm1kw=s64", "userId": "07646999100105069157"}, "user_tz": -420} id="Qk8ewzeWTXUR" outputId="e941af7a-f090-4241-e06d-737410a19e10"
print("Halo Dunia")
print("Riset Bahasa Python")
# + [markdown] colab_type="text" id="pbS6ypD_T9vA"
# Struktur Program Python - Part 1
# + colab={"base_uri": "https://localhost:8080/", "height": 85} colab_type="code" executionInfo={"elapsed": 818, "status": "ok", "timestamp": 1595149384179, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiVlpSEE7CTptyyLOF0HBAyiJig8SaCaXCH-pm1kw=s64", "userId": "07646999100105069157"}, "user_tz": -420} id="g1t3YNRQTXac" outputId="106bbf67-9a3a-4f24-d3c5-ce1f2ddf126b"
# Statement
print("Belajar Python menyenangkan")
print("Halo Dunia")
print("Hello World!")
# Variables & Literals
bilangan1 = 5
bilangan2 = 10
kalimat1 = "Belajar Bahasa Python"
# Operators
print(bilangan1 + bilangan2)
# + [markdown] colab_type="text" id="0Co_C4-XUBSm"
# Tugas Praktek
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 869, "status": "ok", "timestamp": 1595149397923, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiVlpSEE7CTptyyLOF0HBAyiJig8SaCaXCH-pm1kw=s64", "userId": "07646999100105069157"}, "user_tz": -420} id="6pqcCJX9TXc7" outputId="65f986d7-f97f-4338-84dc-3cf1a9d78d86"
bilangan1 = 20
bilangan2 = 10
print(bilangan1 - bilangan2)
# + [markdown] colab_type="text" id="zlITf6UVUGBK"
# Tugas Praktek
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 827, "status": "ok", "timestamp": 1595149419728, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiVlpSEE7CTptyyLOF0HBAyiJig8SaCaXCH-pm1kw=s64", "userId": "07646999100105069157"}, "user_tz": -420} id="eorkr77HTXfX" outputId="884e0d71-d528-49dd-8174-c31495313369"
harga_asli = 20000
potongan = 2000
harga_setelah_potongan = harga_asli - potongan
harga_final = harga_setelah_potongan * 1.1
print(harga_final)
# + [markdown] colab_type="text" id="K7bINgK7UOcH"
# ### **Python Variables & Data Types**
# + [markdown] colab_type="text" id="XeiPcJEoUVYp"
# Sequence Type – Part 1
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" executionInfo={"elapsed": 978, "status": "ok", "timestamp": 1595149481158, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiVlpSEE7CTptyyLOF0HBAyiJig8SaCaXCH-pm1kw=s64", "userId": "07646999100105069157"}, "user_tz": -420} id="q-D1ATHlTXhl" outputId="9486dcb1-e272-42ba-8825-1f6a0ddf5849"
contoh_list = [1,'dua',3,4.0,5]
print(contoh_list[0])
print(contoh_list[3])
contoh_list = [1,'dua',3,4.0,5]
contoh_list[3] = 'empat'
print(contoh_list[3])
# + [markdown] colab_type="text" id="MYgRlHSYUaS3"
# Sequence Type – Part 2
# + colab={"base_uri": "https://localhost:8080/", "height": 214} colab_type="code" executionInfo={"elapsed": 820, "status": "error", "timestamp": 1595149501310, "user": {"displayName": "<NAME>agoto", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiVlpSEE7CTptyyLOF0HBAyiJig8SaCaXCH-pm1kw=s64", "userId": "07646999100105069157"}, "user_tz": -420} id="z_5rs1NlTXjx" outputId="4ac46152-2fd3-4b29-a1ec-bb24b348b4f6"
contoh_tuple = ('Januari','Februari','Maret','April')
print(contoh_tuple[0])
contoh_tuple = ('Januari','Februari','Maret','April')
contoh_tuple[0] = "Desember"
# + [markdown] colab_type="text" id="LEZpgwnpUeK2"
# Set Type
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" executionInfo={"elapsed": 839, "status": "ok", "timestamp": 1595149521911, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiVlpSEE7CTptyyLOF0HBAyiJig8SaCaXCH-pm1kw=s64", "userId": "07646999100105069157"}, "user_tz": -420} id="3biG0V7jTXl_" outputId="cb82f4ad-a2c2-492a-b299-5c0e5b172882"
contoh_list = ['Dewi','Budi','Cici','Linda','Cici']
print(contoh_list)
contoh_set = {'Dewi','Budi','Cici','Linda','Cici'}
print(contoh_set)
contoh_frozen_set = ({'Dewi','Budi','Cici','Linda','Cici'})
print(contoh_frozen_set)
# + [markdown] colab_type="text" id="J4F_bBOzUjb4"
# Mapping Type
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" executionInfo={"elapsed": 850, "status": "ok", "timestamp": 1595149537605, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiVlpSEE7CTptyyLOF0HBAyiJig8SaCaXCH-pm1kw=s64", "userId": "07646999100105069157"}, "user_tz": -420} id="3_xXCKDQTXoA" outputId="5660e162-edce-4dfb-a049-53b4c74d6a7a"
person = {'nama': '<NAME>', 'pekerjaan': 'Programmer'}
print(person['nama'])
print(person['pekerjaan'])
# + [markdown] colab_type="text" id="Z-hyisGRUnNx"
# Tugas Praktek
# + colab={} colab_type="code" id="PhA6BOR0UoTQ"
sepatu = {"nama": "<NAME>", "harga": 150000, "diskon": 30000}
baju = {"nama": "<NAME>", "harga": 80000, "diskon": 8000}
celana = {"nama": "<NAME>", "harga": 200000, "diskon": 60000}
# + [markdown] colab_type="text" id="uolH3ZJzUqPV"
# Tugas Praktek
# + colab={} colab_type="code" id="vQdDmO0nUsYV"
sepatu = {"nama": "<NAME>", "harga": 150000, "diskon": 30000}
baju = {"nama": "<NAME>", "harga": 80000, "diskon": 8000}
celana = {"nama": "<NAME>", "harga": 200000, "diskon": 60000}
daftar_belanja = [sepatu, baju, celana]
# + [markdown] colab_type="text" id="tEPGL0DCUwXf"
# Tugas Praktek
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 854, "status": "ok", "timestamp": 1595149589689, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiVlpSEE7CTptyyLOF0HBAyiJig8SaCaXCH-pm1kw=s64", "userId": "07646999100105069157"}, "user_tz": -420} id="9CKAaHUkUwp9" outputId="0ff786b9-9ae1-46b9-aa57-b723cdf1611e"
# Data yang dinyatakan ke dalam dictionary
sepatu = {"nama": "<NAME>", "harga": 150000, "diskon": 30000}
baju = {"nama": "<NAME>", "harga": 80000, "diskon": 8000}
celana = {"nama": "<NAME>", "harga": 200000, "diskon": 60000}
# Hitunglah harga masing-masing data setelah dikurangi diskon
harga_sepatu = sepatu["harga"] - sepatu["diskon"]
harga_baju = baju["harga"] - baju["diskon"]
harga_celana = celana["harga"] - celana["diskon"]
# Hitung harga total
total_harga = harga_sepatu + harga_baju + harga_celana
# Hitung harga kena pajak
total_pajak = total_harga * 0.1
# Cetak total_harga + total_pajak
print(total_harga + total_pajak)
# + [markdown] colab_type="text" id="LI0KXAOiVXFM"
# ### **Python Operators**
# + [markdown] colab_type="text" id="vM-9nVi7Vg_B"
# Nilai Prioritas Operator dalam Python – Part 1
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" executionInfo={"elapsed": 881, "status": "ok", "timestamp": 1595149789944, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiVlpSEE7CTptyyLOF0HBAyiJig8SaCaXCH-pm1kw=s64", "userId": "07646999100105069157"}, "user_tz": -420} id="IC1CXttGVhQ1" outputId="1bbae88b-627d-4239-f594-69a1117a8617"
# Kode awal
total_harga = 150000
potongan_harga = 0.3
pajak = 0.1 # pajak dalam persen ~ 10%
harga_bayar = 1 - 0.3 # baris pertama
harga_bayar *= total_harga # baris kedua
pajak_bayar = pajak * harga_bayar # baris ketiga
harga_bayar += pajak_bayar # baris ke-4
print("Kode awal - harga_bayar=", harga_bayar)
# Penyederhanaan baris kode dengan menerapkan prioritas operator
total_harga = 150000
potongan_harga = 0.3
pajak = 0.1 # pajak dalam persen ~ 10%
harga_bayar = (1 - 0.3) * total_harga #baris pertama
harga_bayar += harga_bayar * pajak # baris kedua
print("Penyederhanaan kode - harga_bayar=", harga_bayar)
# + [markdown] colab_type="text" id="mBJvMChNVqSI"
# Tugas Praktek
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 850, "status": "ok", "timestamp": 1595149827891, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiVlpSEE7CTptyyLOF0HBAyiJig8SaCaXCH-pm1kw=s64", "userId": "07646999100105069157"}, "user_tz": -420} id="64b8I9cMVqp2" outputId="82f6a770-cbb4-4ad8-ac0c-ef581c12ef42"
sepatu = { "nama" : "<NAME>", "harga": 150000, "diskon": 30000 }
baju = { "nama" : "<NAME>", "harga": 80000, "diskon": 8000 }
celana = { "nama" : "<NAME>", "harga": 200000, "diskon": 60000 }
harga_sepatu = sepatu["harga"] - sepatu["diskon"]
harga_baju = baju["harga"] - baju["diskon"]
harga_celana = celana["harga"] - celana["diskon"]
total_harga = (harga_sepatu + harga_baju + harga_celana) * 1.1
print(total_harga)
# + [markdown] colab_type="text" id="yxRaoEJZVwE4"
# ### **Pythons Conditioning & Looping**
# + [markdown] colab_type="text" id="dOLJt0_fVzZn"
# Python Conditioning for Decision – Part 2
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" executionInfo={"elapsed": 812, "status": "ok", "timestamp": 1595149865508, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiVlpSEE7CTptyyLOF0HBAyiJig8SaCaXCH-pm1kw=s64", "userId": "07646999100105069157"}, "user_tz": -420} id="FkQyWh3LVz3e" outputId="909f91c6-17cb-4c03-f9c4-892209024dbe"
# Statement if
x = 4
if x % 2 == 0: # jika sisa bagi x dengan 2 sama dengan 0
print("x habis dibagi dua") # statemen aksi lebih menjorok ke dalam
# Statement if ... elif ... else
x = 7
if x % 2 == 0: # jika sisa bagi x dengan 2 sama dengan 0
print("x habis dibagi dua")
elif x % 3 == 0: # jika sisa bagi x dengan 3 sama dengan 0
print("x habis dibagi tiga")
elif x % 5 == 0: # jika sisa bagi x dengan 5 sama dengan 0
print("x habis dibagi lima")
else:
print("x tidak habis dibagi dua, tiga ataupun lima")
# + [markdown] colab_type="text" id="2s2NDaPeV3_V"
# Python Conditioning for Decision – Part 3
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 875, "status": "ok", "timestamp": 1595149884205, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiVlpSEE7CTptyyLOF0HBAyiJig8SaCaXCH-pm1kw=s64", "userId": "07646999100105069157"}, "user_tz": -420} id="hxBHcLYxV4RN" outputId="c91f7494-b2cf-400f-f384-040cba4c1fa7"
jam = 13
if jam >= 5 and jam < 12: # selama jam di antara 5 s.d. 12
print("Selamat pagi!")
elif jam >= 12 and jam < 17: # selama jam di antara 12 s.d. 17
print("Selamat siang!")
elif jam >= 17 and jam < 19: # selama jam di antara 17 s.d. 19
print("Selamat sore!")
else: # selain kondisi di atas
print("Selamat malam!")
# + [markdown] colab_type="text" id="zTV5kfU7V7sD"
# Tugas Praktek
# + colab={"base_uri": "https://localhost:8080/", "height": 85} colab_type="code" executionInfo={"elapsed": 860, "status": "ok", "timestamp": 1595149899038, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiVlpSEE7CTptyyLOF0HBAyiJig8SaCaXCH-pm1kw=s64", "userId": "07646999100105069157"}, "user_tz": -420} id="Mq4RlofRV8Fs" outputId="14c0eb92-b549-4011-f4cd-d613655af08d"
tagihan_ke = 'Mr. Yoyo'
warehousing = { 'harga_harian': 1000000, 'total_hari':15 }
cleansing = { 'harga_harian': 1500000, 'total_hari':10 }
integration = { 'harga_harian':2000000, 'total_hari':15 }
transform = { 'harga_harian':2500000, 'total_hari':10 }
sub_warehousing = warehousing['harga_harian'] * warehousing['total_hari']
sub_cleansing = cleansing['harga_harian'] * cleansing['total_hari']
sub_integration = integration['harga_harian'] * integration['total_hari']
sub_transform = transform['harga_harian'] * transform['total_hari']
total_harga = sub_warehousing + sub_cleansing + sub_integration + sub_transform
print("Tagihan kepada:")
print(tagihan_ke)
print("Selamat pagi, anda harus membayar tagihan sebesar:")
print(total_harga)
# + [markdown] colab_type="text" id="ML87mFnXV_xK"
# Tugas Praktek
# + colab={"base_uri": "https://localhost:8080/", "height": 85} colab_type="code" executionInfo={"elapsed": 822, "status": "ok", "timestamp": 1595149918164, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiVlpSEE7CTptyyLOF0HBAyiJig8SaCaXCH-pm1kw=s64", "userId": "07646999100105069157"}, "user_tz": -420} id="TZv51ZSeV_85" outputId="d8143e11-93de-4981-9a8a-1de08e566e34"
jam = 17
tagihan_ke = 'Mr. Yoyo'
warehousing = { 'harga_harian': 1000000, 'total_hari':15 }
cleansing = { 'harga_harian': 1500000, 'total_hari':10 }
integration = { 'harga_harian':2000000, 'total_hari':15 }
transform = { 'harga_harian':2500000, 'total_hari':10 }
sub_warehousing = warehousing['harga_harian']*warehousing['total_hari']
sub_cleansing = cleansing['harga_harian']*cleansing['total_hari']
sub_integration = integration['harga_harian']*integration['total_hari']
sub_transform = transform['harga_harian']*transform['total_hari']
total_harga = sub_warehousing+sub_cleansing+sub_integration+sub_transform
print("Tagihan kepada:")
print(tagihan_ke)
if jam > 19:
print("Selamat malam, anda harus membayar tagihan sebesar:")
elif jam > 17:
print("Selamat sore, anda harus membayar tagihan sebesar:")
elif jam > 12:
print("Selamat siang, anda harus membayar tagihan sebesar:")
else:
print("Selamat pagi, anda harus membayar tagihan sebesar:")
print(total_harga)
# + [markdown] colab_type="text" id="f63H9UYfWGMP"
# Python while loops – Part 1
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" executionInfo={"elapsed": 865, "status": "ok", "timestamp": 1595149941603, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiVlpSEE7CTptyyLOF0HBAyiJig8SaCaXCH-pm1kw=s64", "userId": "07646999100105069157"}, "user_tz": -420} id="qGfnUPkFWGYx" outputId="28fd49ce-3d1a-43f1-ecc4-c59d49c0a1be"
# Tagihan
tagihan = [50000, 75000, 125000, 300000, 200000]
# Tanpa menggunakan while loop
total_tagihan = tagihan[0] + tagihan[1] + tagihan[2] + tagihan[3] + tagihan[4]
print(total_tagihan)
# Dengan menggunakan while loop
i = 0 # sebuah variabel untuk mengakses setiap elemen tagihan satu per satu
jumlah_tagihan = len(tagihan) # panjang (jumlah elemen dalam) list tagihan
total_tagihan = 0 # mula-mula, set total_tagihan ke 0
while i < jumlah_tagihan: # selama nilai i kurang dari jumlah_tagihan
total_tagihan += tagihan[i] # tambahkan tagihan[i] ke total_tagihan
i += 1 # tambahkan nilai i dengan 1 untuk memproses tagihan selanjutnya.
print(total_tagihan)
# + [markdown] colab_type="text" id="OyB21mI6WKHP"
# Python while loops – Part 2
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" executionInfo={"elapsed": 878, "status": "ok", "timestamp": 1595149958137, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiVlpSEE7CTptyyLOF0HBAyiJig8SaCaXCH-pm1kw=s64", "userId": "07646999100105069157"}, "user_tz": -420} id="1KDF4MLIWKfj" outputId="e51ec065-a205-4d83-d7a5-e35ba71a166c"
tagihan = [50000, 75000, -150000, 125000, 300000, -50000, 200000]
i = 0
jumlah_tagihan = len(tagihan)
total_tagihan = 0
while i < jumlah_tagihan:
# jika terdapat tagihan ke-i yang bernilai minus (di bawah nol),
# pengulangan akan dihentikan
if tagihan[i] < 0:
total_tagihan = -1
print("terdapat angka minus dalam tagihan, perhitungan dihentikan!")
break
total_tagihan += tagihan[i]
i += 1
print(total_tagihan)
# + [markdown] colab_type="text" id="rRpUrjxcWOEx"
# Python while loops – Part 3
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 821, "status": "ok", "timestamp": 1595149974506, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiVlpSEE7CTptyyLOF0HBAyiJig8SaCaXCH-pm1kw=s64", "userId": "07646999100105069157"}, "user_tz": -420} id="l-Yfl-85WOSY" outputId="4465d540-2e2b-4c22-a00d-a940bea82aeb"
tagihan = [50000, 75000, -150000, 125000, 300000, -50000, 200000]
i = 0
jumlah_tagihan = len(tagihan)
total_tagihan = 0
while i < jumlah_tagihan:
# jika terdapat tagihan ke-i yang bernilai minus (di bawah nol),
# abaikan tagihan ke-i dan lanjutkan ke tagihan berikutnya
if tagihan[i] < 0:
i += 1
continue
total_tagihan += tagihan[i]
i += 1
print(total_tagihan)
# + [markdown] colab_type="text" id="Y_0Yx7YoWR_W"
# Python for loops – Part 1
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 808, "status": "ok", "timestamp": 1595149989900, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiVlpSEE7CTptyyLOF0HBAyiJig8SaCaXCH-pm1kw=s64", "userId": "07646999100105069157"}, "user_tz": -420} id="bszKS1_5WSMX" outputId="41129431-16af-41bf-8f2e-9506bc9b4da5"
list_tagihan = [50000, 75000, -150000, 125000, 300000, -50000, 200000]
total_tagihan = 0
for tagihan in list_tagihan: # untuk setiap tagihan dalam list_tagihan
total_tagihan += tagihan # tambahkan tagihan ke total_tagihan
print(total_tagihan)
# + [markdown] colab_type="text" id="y1jPRBO6WVor"
# Python for loops – Part 2
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" executionInfo={"elapsed": 1005, "status": "ok", "timestamp": 1595150005421, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiVlpSEE7CTptyyLOF0HBAyiJig8SaCaXCH-pm1kw=s64", "userId": "07646999100105069157"}, "user_tz": -420} id="6KIhfuJlWV7x" outputId="6a1a8800-50f3-47c7-b9ef-80ebe2bc6830"
list_tagihan = [50000, 75000, -150000, 125000, 300000, -50000, 200000]
total_tagihan = 0
for tagihan in list_tagihan:
if tagihan < 0:
print("terdapat angka minus dalam tagihan, perhitungan dihentikan!")
break
total_tagihan += tagihan
print(total_tagihan)
# + [markdown] colab_type="text" id="dIzO8TxBWZnm"
# Python for loops – Part 3
# + colab={"base_uri": "https://localhost:8080/", "height": 170} colab_type="code" executionInfo={"elapsed": 834, "status": "ok", "timestamp": 1595150021066, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiVlpSEE7CTptyyLOF0HBAyiJig8SaCaXCH-pm1kw=s64", "userId": "07646999100105069157"}, "user_tz": -420} id="DtqA3ah9WZ0H" outputId="e446109c-1ca4-45b1-8578-18991428bfa6"
list_daerah = ['Malang', 'Palembang', 'Medan']
list_buah = ['Apel', 'Duku', 'Jeruk']
for nama_daerah in list_daerah:
for nama_buah in list_buah:
print(nama_buah+" "+nama_daerah)
# + [markdown] colab_type="text" id="69spzHHqWdGi"
# Tugas Praktek
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" executionInfo={"elapsed": 844, "status": "ok", "timestamp": 1595150034916, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiVlpSEE7CTptyyLOF0HBAyiJig8SaCaXCH-pm1kw=s64", "userId": "07646999100105069157"}, "user_tz": -420} id="7KtpqFl2WdV4" outputId="a065c7d2-1f8e-488b-8bc6-5b9f69e4d9a2"
list_cash_flow = [
2500000, 5000000, -1000000, -2500000, 5000000, 10000000,
-5000000, 7500000, 10000000, -1500000, 25000000, -2500000
]
total_pengeluaran, total_pemasukan = 0, 0
for dana in list_cash_flow:
if dana > 0:
total_pemasukan += dana
else:
total_pengeluaran += dana
total_pengeluaran *= -1
print(total_pengeluaran)
print(total_pemasukan)
# + [markdown] colab_type="text" id="FYYE57woWh9_"
# ### **Mini Quiz**
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 850, "status": "ok", "timestamp": 1595150061380, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiVlpSEE7CTptyyLOF0HBAyiJig8SaCaXCH-pm1kw=s64", "userId": "07646999100105069157"}, "user_tz": -420} id="qawnM3tTWjPN" outputId="f6823c96-6599-4d55-e514-50a09eace7b0"
# Data
uang_jalan = 1500000
jumlah_hari = 31
list_plat_nomor = [8993, 2198, 2501, 2735, 3772, 4837, 9152]
# Pengecekan kendaraan dengan nomor pelat ganjil atau genap
# Deklarasikan kendaraan_genap dan kendaraan_ganjil = 0
kendaraan_genap = 0
kendaraan_ganjil = 0
for plat_nomor in list_plat_nomor:
if plat_nomor %2 == 0:
kendaraan_genap += 1
else:
kendaraan_ganjil += 1
# Total pengeluaran untuk kendaraan dengan nomor pelat ganjil
# dan genap dalam 1 bulan
i = 1
total_pengeluaran = 0
while i <= jumlah_hari:
if i % 2 == 0:
total_pengeluaran += (kendaraan_genap * uang_jalan)
else:
total_pengeluaran += (kendaraan_ganjil * uang_jalan)
i += 1
# Cetak total pengeluaran
print(total_pengeluaran)
# + [markdown] colab_type="text" id="tkjkyJP6E64p"
# Kesimpulan
# + [markdown] colab_type="text" id="2QccCzdtE9nh"
# 1. Alasan Python secara luas digunakan dalam komputasi saintifik, web, ranah data (data domain).
# 2. Konstruksi dari struktur bahasa pemrograman Python.
# 3. Teknik mempraktekkan penggunaan tipe data pada Python.
# 4. Teknik mempraktekkan penggunaan jenis-jenis operator pada Python.
# 5. Teknik mempraktekkan penggunaan pengkondisian untuk pengambilan keputusan dan perulangan pada Python.
# 6. Program Python untuk penyelesaian kasus bisnis sederhana.
| Python for Data Professional Beginner - Part 1/Python for Data Professional Beginner - Part 1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="ULx_xUCpR-yo" executionInfo={"status": "ok", "timestamp": 1612774549562, "user_tz": -180, "elapsed": 27458, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11793698916100822526"}} outputId="84ff8e5f-841e-4989-9993-6ab21b0a36b5"
# !sudo apt install -y libsndfile1
# !pip install numba==0.48
# !pip install git+https://github.com/fastaudio/fastaudio.git
# + colab={"base_uri": "https://localhost:8080/"} id="jnyma5fkuA6A" executionInfo={"status": "ok", "timestamp": 1612774556189, "user_tz": -180, "elapsed": 34076, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11793698916100822526"}} outputId="21480486-e192-4b13-dbc1-30901b0e9751"
# !pip install timm
# !pip install efficientnet_pytorch
# + id="E798_JhkR-y_" executionInfo={"status": "ok", "timestamp": 1612774842176, "user_tz": -180, "elapsed": 1286, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11793698916100822526"}}
# %reload_ext autoreload
# %autoreload 2
# %matplotlib inline
# + colab={"base_uri": "https://localhost:8080/"} id="puKPWfwVR-zA" executionInfo={"status": "ok", "timestamp": 1612774845034, "user_tz": -180, "elapsed": 4125, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11793698916100822526"}} outputId="2bc34fe3-5052-495f-f6c3-9fcbc2d50e65"
import torchaudio
torchaudio.set_audio_backend("sox_io")
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="tvpx5YenR-zB" executionInfo={"status": "ok", "timestamp": 1612774847022, "user_tz": -180, "elapsed": 6108, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11793698916100822526"}} outputId="fdddc0c6-9b27-48ac-9847-5f456a2e845d"
import numpy as np
import pandas as pd
import os
import sys
sys.path.append("..")
import librosa as lr
import librosa.display
import soundfile as sf
import io
from pathlib import Path
from tqdm.notebook import tqdm
from fastaudio.core.all import *
from fastaudio.augment.all import *
from fastai.torch_basics import *
from fastai.basics import *
from fastai.data.all import *
from fastai.callback.all import *
from fastai.vision.all import *
import fastai
fastai.__version__
# + colab={"base_uri": "https://localhost:8080/"} id="byHiKwd1J4-8" executionInfo={"status": "ok", "timestamp": 1612774867991, "user_tz": -180, "elapsed": 27071, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11793698916100822526"}} outputId="fab7099f-cbae-44f4-b97a-40ddb41a50dd"
from google.colab import drive
drive.mount('/content/drive')# You must grant COLAB access to your Google Drive
# + colab={"base_uri": "https://localhost:8080/", "height": 436} id="ZFkobKPoR-zB" executionInfo={"status": "ok", "timestamp": 1612774884407, "user_tz": -180, "elapsed": 43482, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11793698916100822526"}} outputId="94b247d1-f6e3-4f2a-d630-1bd4cfb02887"
DATA_ROOT = Path("/content/drive/MyDrive/Colab Notebooks/RainForestAudio/data")
AUDIO_ROOT = Path(DATA_ROOT/"train/")
TRAIN_AUDIO_ROOT = Path(DATA_ROOT/"samples_center")
#TRAIN_AUDIO_ROOT = Path("/content/samples_long")
#TRAIN_AUDIO_ROOT = Path(DATA_ROOT/"samples_mixed")
TEST_AUDIO_ROOT = Path(DATA_ROOT/"test")
VAL_AUDIO_ROOT = Path(DATA_ROOT/"val")
df_train = pd.DataFrame([path.stem for path in Path(TRAIN_AUDIO_ROOT).glob("*.flac")], columns=["recording_id"])
train_folds = np.load(Path(DATA_ROOT/"folds.npy"), allow_pickle=True)
df_test = pd.DataFrame([path.stem for path in Path(TEST_AUDIO_ROOT).glob("*.flac")], columns=["recording_id"])
df = pd.read_csv(Path(DATA_ROOT/"train_tp.csv"))
print(df_train.shape, df_test.shape)
df
# + [markdown] id="vedRqZlR7L-4"
# # Define multi class model
# + colab={"base_uri": "https://localhost:8080/"} id="XVWvGZv1LOE4" executionInfo={"status": "ok", "timestamp": 1612774884408, "user_tz": -180, "elapsed": 43477, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11793698916100822526"}} outputId="02a4b055-d735-43ea-a374-2916addbfcdc"
def get_y_fn(x):
y = str(x).split(".")[0].split('_')[-2]
return y
FOLD = 4
val_index = train_folds[FOLD]
FOLD = str(FOLD)
val_index
# + colab={"base_uri": "https://localhost:8080/"} id="h8-aYX03_IRV" executionInfo={"status": "ok", "timestamp": 1612774884409, "user_tz": -180, "elapsed": 43474, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11793698916100822526"}} outputId="e86f7199-fe86-44b2-d337-0661268b6cc2"
train_folds[0].shape
# + [markdown] id="WGGZLJuaTmd6"
# ## define mixup
# + colab={"base_uri": "https://localhost:8080/"} id="OJdJEY12Tl4-" executionInfo={"status": "ok", "timestamp": 1612774886291, "user_tz": -180, "elapsed": 45351, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11793698916100822526"}} outputId="c5104964-5761-4c68-a073-b33bb0f6ee11"
# !git clone https://github.com/nestordemeure/ManifoldMixupV2.git
# + colab={"base_uri": "https://localhost:8080/"} id="xxN6SwtbTsld" executionInfo={"status": "ok", "timestamp": 1612774888744, "user_tz": -180, "elapsed": 47800, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11793698916100822526"}} outputId="332c64a2-3c36-4e70-bbe3-43377b5e8a59"
# %run /content/ManifoldMixupV2/manifold_mixup.py
OutputMixup()
# + [markdown] id="CQc9Jo_wDCPv"
# ## Define db and dls
# + colab={"base_uri": "https://localhost:8080/"} id="HI_0-L4N3ldB" executionInfo={"status": "ok", "timestamp": 1612774890984, "user_tz": -180, "elapsed": 50037, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11793698916100822526"}} outputId="91f4e292-0b48-443c-dfb6-408075b83960"
# !git clone https://NadyaStrogankova:c12ca00be6ebdcb705be6f0e9fac559a684c3d3b@github.com/NadyaStrogankova/RainforestAudioKaggle.git
# + id="rf8Z7C_P3oID" executionInfo={"status": "ok", "timestamp": 1612774890985, "user_tz": -180, "elapsed": 50037, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11793698916100822526"}}
# %run /content/RainforestAudioKaggle/transforms.py
# + colab={"base_uri": "https://localhost:8080/"} id="OaxZGk-1R-zS" outputId="91d4acb3-a266-472a-814d-d838f139d8bf"
cfg = AudioConfig.BasicMelSpectrogram(
mel=True,
# to_db = False
f_min=df["f_min"].min(),
f_max=df["f_max"].max(),
# n_fft=1024,
n_mels=384,
hop_length=292,
# n_fft = 892,
#hop_length=245,
#n_mels = 224,
# normalized=True
)
item_tfms = [Resample(28000),
ResizeSignal(8000, pad_mode=AudioPadType.Repeat),
AddNoise(noise_level=0.05, color=NoiseColor.Pink),
AddNoise(noise_level=0.05, color=NoiseColor.White),
]
batch_tfms = [ AudioToSpec.from_cfg(cfg),
# Normalize_channel(),
# PowerSpec(), # увеличение контрастности
# TAmplitudeToDB(),
Normalize_channel_1(),
# PowerSpec(1.5, 0.7),
WhiteNoise(0.005, cfg),
PinkNoise(0.005, cfg),
# RowNoise(0.025, cfg),
LowerUpperFreq(cfg),
Normalize_channel_2(),
PowerSpec(2, 0.7), # увеличение контрастности
# Normalize_channel_3(),
SGRoll(),
Mask(),
# Mono2Color()
# Normalize(ch_mean, ch_std, axes=(0, 1, 3)),
]
AddNoise.split_idx = 0
Mask.split_idx = 0
SGRoll.split_idx = 0
auds = DataBlock(blocks = (AudioBlock, CategoryBlock),
get_items=get_audio_files,
item_tfms = item_tfms,
splitter = IndexSplitter(val_index.tolist()), #report unnesesary transform to list
#splitter = RandomSplitter(),
get_y=get_y_fn,
batch_tfms = batch_tfms
)
print(auds.summary(TRAIN_AUDIO_ROOT))
dls = auds.dataloaders(TRAIN_AUDIO_ROOT, bs=24, verbose=True, before_batch = batch_tfms,
num_workers=2
#, shuffle=True
)
# + id="O3e_04_DZKaq"
def _one_sample_positive_class_precisions(scores, truth):
num_classes = scores.shape[0]
pos_class_indices = np.flatnonzero(truth > 0)
if not len(pos_class_indices):
return pos_class_indices, np.zeros(0)
retrieved_classes = np.argsort(scores)[::-1]
class_rankings = np.zeros(num_classes, dtype=np.int)
class_rankings[retrieved_classes] = range(num_classes)
retrieved_class_true = np.zeros(num_classes, dtype=np.bool)
retrieved_class_true[class_rankings[pos_class_indices]] = True
retrieved_cumulative_hits = np.cumsum(retrieved_class_true)
precision_at_hits = (
retrieved_cumulative_hits[class_rankings[pos_class_indices]] /
(1 + class_rankings[pos_class_indices].astype(np.float)))
return pos_class_indices, precision_at_hits
def lwlrap(scores, truth):
#print(truth.shape, scores.shape)
num_samples, num_classes = scores.shape
scores = scores.cpu().numpy()
gt = np.zeros((num_samples, num_classes))
for n, i in enumerate(truth.cpu().numpy().astype(int)):
gt[n, i] = 1
assert gt.shape == scores.shape
precisions_for_samples_by_classes = np.zeros((num_samples, num_classes))
for sample_num in range(num_samples):
pos_class_indices, precision_at_hits = _one_sample_positive_class_precisions(scores[sample_num, :], gt[sample_num, :])
precisions_for_samples_by_classes[sample_num, pos_class_indices] = precision_at_hits
labels_per_class = np.sum(gt > 0, axis=0)
weight_per_class = labels_per_class / float(np.sum(labels_per_class))
per_class_lwlrap = (np.sum(precisions_for_samples_by_classes, axis=0) /
np.maximum(1, labels_per_class))
#return per_class_lwlrap, weight_per_class
return (per_class_lwlrap * weight_per_class).sum()
# + id="A-HyUhFNhvlw"
from efficientnet_pytorch import EfficientNet
from timm import create_model
# from https://colab.research.google.com/github/muellerzr/Practical-Deep-Learning-for-Coders-2.0/blob/master/Computer%20Vision/05_EfficientNet_and_Custom_Weights.ipynb#scrollTo=VXPjDVUlJgCU
# + id="ewC-e6U0uHDi"
def create_timm_body(arch:str, pretrained=True, cut=None):
model = create_model(arch, pretrained=pretrained, in_chans=1)
if cut is None:
ll = list(enumerate(model.children()))
cut = next(i for i,o in reversed(ll) if has_pool_type(o))
if isinstance(cut, int): return nn.Sequential(*list(model.children())[:cut])
elif callable(cut): return cut(model)
else: raise NamedError("cut must be either integer or function")
body = create_timm_body('resnest50_fast_1s1x64d', pretrained=True)
body = create_timm_body('resnet34', pretrained=True, cut=-2)
#head = create_head(num_features_model(nn.Sequential(*body.children())) * (2), dls.c)
head = create_head(512, dls.c, concat_pool=False, ps=0.4)
model = nn.Sequential(body, head)
apply_init(model[1], nn.init.kaiming_normal_)
len(model)
# + id="CMvV_yrz509E"
model
# + id="XeYflMxtR-zS"
learn = Learner(dls, model,
# n_in=1, #<- Only audio specific modification here
# loss_func=LabelSmoothingCrossEntropyFlat(),
cbs = OutputMixup(alpha=0.4),
metrics=[accuracy, lwlrap])
learn.to_fp16()
learn.loss_func
# + id="xb5wZcjKIIJ8"
dls.show_batch(), dls.vocab
# + id="2EEWc0D-R-zT"
#learn.lr_find()
# + id="7VAxjc5_O2nd"
#wdc wc
# + id="U5gS61f2fSYO"
EXP_NAME = "rn34_fold" + FOLD
SUFFIX ="step1"
learn.freeze()
learn.fit(3, 1e-3/2)
learn.unfreeze()
base_lr = 1e-3/2
learn.fit_one_cycle(25, slice(base_lr / 2, base_lr * 2), wd=1e-2, div=10, div_final=1e+2, pct_start=0.3,
cbs=[SaveModelCallback(fname = f'{EXP_NAME}_{SUFFIX}', with_opt=True)]
)
# + id="986JzSZPhJNF"
SUFFIX ="step2"
learn.load(EXP_NAME+"_step1")
base_lr = base_lr / 4
learn.fit_one_cycle(24, slice(base_lr / 4, base_lr * 4), wd=1e-2, div=10, div_final=1e+2, pct_start=0.3,
cbs=[SaveModelCallback(fname = f'{EXP_NAME}_{SUFFIX}', with_opt=True)]
)
# + id="y4-B1Shji-lx"
SUFFIX ="step3"
#learn.load(EXP_NAME+"_step2")
base_lr = 1e-3/2
base_lr = base_lr / 8
learn.fit_one_cycle(25, slice(base_lr / 2, base_lr * 2), wd=1e-2, div=10, div_final=1e+2, pct_start=0.3,
cbs=[SaveModelCallback(fname = f'{EXP_NAME}_{SUFFIX}', with_opt=True)]
)
# + id="R96dpPgZ_l-C"
# SUFFIX ="step4"
# #learn.load(EXP_NAME+"_step2")
# base_lr = 1e-3/2
# base_lr = base_lr / 16
# learn.fit_one_cycle(25, slice(base_lr / 2, base_lr * 2), wd=1e-2, div=10, div_final=1e+2, pct_start=0.3,
# cbs=[SaveModelCallback(fname = f'{EXP_NAME}_{SUFFIX}', with_opt=True)]
# )
# + id="F0mDWQ-eJTEe"
# SUFFIX ="step5"
# #learn.load(EXP_NAME+"_step2")
# base_lr = 1e-3/2
# base_lr = base_lr / 32
# learn.fit_one_cycle(25, slice(base_lr / 2, base_lr * 2), wd=1e-2, div=10, div_final=1e+2, pct_start=0.3,
# cbs=[SaveModelCallback(fname = f'{EXP_NAME}_{SUFFIX}', with_opt=True)])
# + [markdown] id="oYuAzydi7sNv"
# ## confusion matrix
# + [markdown] id="_EAhQY4p7YLx"
# # Predict all classes
# + id="lx9-pVnNUuT6"
learn.load("rn34_fold"+FOLD+"_step1", with_opt=False)
learn.remove_cbs(OutputMixup)
learn.cbs
# + id="_N8zoCoknR8h"
interp = ClassificationInterpretation.from_learner(learn)
interp.plot_confusion_matrix()
# + id="pFMZcAnefQSR"
# #!ln -s drive/MyDrive/Colab\ Notebooks/RainForestAudio/data/ol_samples.tar.gz ol_samples.tar.gz
# #!ln -s drive/MyDrive/Colab\ Notebooks/RainForestAudio/data/val val
# + id="OurWFYgnAMiG"
# #!tar -xf ol_samples.tar.gz
# + id="Mlgz3NW6x3Iz"
TEST_SAMPLES_AUDIO_ROOT = Path("/content/samples")
# #!ln -s drive/MyDrive/Colab\ Notebooks/RainForestAudio/data/val/ /content/val
VAL_AUDIO_ROOT = Path("/content/val")
test_ds = auds.new(VAL_AUDIO_ROOT)
test_dl = learn.dls.test_dl(get_audio_files(TEST_SAMPLES_AUDIO_ROOT))
#print(test_dl.summary(TEST_TARGET_AUDIO_ROOT))
#test_probas, *_ = learn.tta(dl=test_dl, n=2, use_max=True)
test_probas, *_ = learn.tta(dl=test_dl, n=6, use_max=False, beta=1/7)
# + id="6tGOuDrT5kjG"
result=[]
for probas, fname in zip(test_probas, test_dl.items):
print(fname)
result.append([fname.stem.split("_")[0], fname.stem.split("_")[1], probas.numpy()])
res = pd.DataFrame(result, columns =["recording_id", "part_id", "probas"])
res["part_id"] = res["part_id"].astype(int)
res.to_csv("all_class_pred_"+FOLD+".csv", columns =["recording_id", "part_id", "probas"])
# + id="hFMycFCou8Fj"
subm = []
for n, row in res.sort_values(by="recording_id").groupby(by="recording_id"):
a = np.stack(row["probas"].values)
pred = np.concatenate([np.array(n).reshape(1),
np.array(dls.vocab[np.argmax(a.max(axis=0))]).reshape(1),
#np.where(a>0.7, a, 0).sum(axis=0)
a.max(axis=0)
])
subm.append(pred)
#print(n)
if n == "047a7c4bf":
#print(row["probas"].max(axis=1))
cls = np.stack(row["probas"].values).max(axis=0).argsort()
#print(np.stack(row["probas"].values).argsort())
#print(dls.vocab[cls[-1]], dls.vocab[cls[-2]], dls.vocab[cls[-3]])
#print(subm)
submission = pd.DataFrame(subm, columns=["recording_id"] + ["top_cat"] + ["s" + i for i in dls.vocab])
submission.head(10)
# + id="hNaDxF9i9BxC"
submission.drop(["top_cat"], axis=1, inplace=True)
submission.to_csv("subm_29_fold_"+FOLD+"_0.csv", columns=["recording_id"] + ["s" + str(i) for i in range(24)], index=False)
# + id="eYoGVIKpkuYD"
from sklearn.metrics import accuracy_score
gt=[]
pred=[]
for n, row in submission.iterrows():
try:
gr_tr = df[df["recording_id"] == row["recording_id"]]["species_id"].to_numpy()
if gr_tr.shape[0] > 1:
if int(row["top_cat"]) in gr_tr:
gt.append(int(row["top_cat"]))
else:
gt.append(gr_tr[0])
else:
gt.append(gr_tr[0])
pred.append(int(row["top_cat"]))
except:
pass
accuracy_score(gt,pred), len(gt)
#gt, pred
# + [markdown] id="kWqKOZozzgqK"
# Усредненный сабмит
# + id="QJt4s3ATzfmP"
subm = []
#subm.append(submission.sort_values(by="recording_id").to_numpy()[:,1:].astype(float))
for i in range(4):
s = pd.read_csv("subm_29_fold_"+str(i)+"_0.csv")
subm.append(s.sort_values(by="recording_id").to_numpy()[:,1:])
subm = np.stack(subm)
subm.shape
fs = pd.DataFrame(np.concatenate([submission.sort_values(by="recording_id").to_numpy()[:,0].reshape(-1,1), subm.sum(axis=0)], axis=1), columns=s.columns)
fs.head()
# + id="snvvT4mb3UOj"
fs.to_csv("subm_29_rn34.csv", index=False)
# + id="ZQ5_0kSlzfpn"
# !tar -cf rn_34_weigth_2901.tar models
# + id="YuC4LH7HLFUp"
# !cp rn_34_weigth_2901.tar /content/drive/MyDrive/Colab\ Notebooks/RainForestAudio
# + [markdown] id="oAdoQTK_zJeH"
# ## Вспомогательное
#
# + id="va90ifjuZnBf"
class MaskFreq_fixed(SpectrogramTransform):
"""Google SpecAugment frequency masking from https://arxiv.org/abs/1904.08779."""
def __init__(self, num_masks=1, size=20, start=None, val=None):
self.num_masks = num_masks
self.size = size
self.start = start
self.val = val
def encodes(self, sg: AudioSpectrogram) -> AudioSpectrogram:
channel_mean = sg.contiguous().view(sg.size(0), -1).mean(-1)[:, None, None]
mask_val = ifnone(self.val, channel_mean)
if sg.ndim == 4:
b, c, y, x = sg.shape
# Position of the first mask
start = ifnone(self.start, random.randint(0, y - self.size))
for _ in range(self.num_masks):
mask = torch.ones(self.size, x).cuda() * mask_val.cuda()
mask = mask.view(b, c, self.size, x)
#print("sg, mask:", sg.shape, mask.shape)
if not 0 <= start <= y - self.size:
raise ValueError(
f"Start value '{start}' out of range for AudioSpectrogram of shape {sg.shape}"
)
sg[:, :, start : start + self.size, :] = mask
# Setting start position for next mask
start = random.randint(0, y - self.size)
else:
c, y, x = sg.shape
# Position of the first msk
start = ifnone(self.start, random.randint(0, y - self.size))
for _ in range(self.num_masks):
mask = torch.ones(self.size, x) * mask_val
if not 0 <= start <= y - self.size:
raise ValueError(
f"Start value '{start}' out of range for AudioSpectrogram of shape {sg.shape}"
)
sg[:, start : start + self.size, :] = mask
# Setting start position for next mask
start = random.randint(0, y - self.size)
return sg
| bag_of_notebooks/fastaudio_resnet34_moreaugs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/aguilarmajan/CPEN-21A-CPE-1-1/blob/main/Operations%20and%20Expressions.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="BwqggyOd1w0-"
# # Operations and Expressions
#
# + colab={"base_uri": "https://localhost:8080/"} id="KwYUq9ieRgnP" outputId="5f38192b-1ba4-44b7-d77d-42c21710880b"
print (10+5)
print (10-5)
print (10*5)
print (10/5)
print (10%5)
print (10//3)
print (10**2)
# + colab={"base_uri": "https://localhost:8080/"} id="vVCL9OOASbkv" outputId="25b74403-453d-4c44-f3f7-71f136d6ce7f"
print(bool("Hello"))
print (bool(15))
# + colab={"base_uri": "https://localhost:8080/"} id="-ezVoP0-StB6" outputId="22996c73-8af0-4418-929b-34751d6fb620"
print(bool(False))
print(bool(None))
print(bool(0))
print(bool(""))
print(bool())
print(bool([]))
print(bool({}))
# + id="7jBO0p_DUnNr"
def myFunction():
return True
print(myFunction())
# + id="QXxIM5nHUzck"
def myFunction():
return True
if myFunction():
print("YES!")
else:
print("NO!")
# + colab={"base_uri": "https://localhost:8080/"} id="rp30NqDNVd07" outputId="9ebd59e2-acdd-4eca-be00-805ab9652a53"
print (11>8)
a = 6
b = 7
print(a==b)
print(a!=a)
| Operations and Expressions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Pyolite
# language: python
# name: python
# ---
# + [markdown] tags=[]
# # `folium` Interactive Map Demo
#
# Simple demonstration of rendering a map in a `jupyterlite` notebook.
#
# Note that the `folium` package has several dependencies which themselves may have dependencies.
#
# The following code fragement, run in a fresh Python enviroment into which `folium` has already been installed, identifies the packages that are loaded in when `folium` is loaded:
#
# ```python
# #https://stackoverflow.com/a/40381601/454773
# import sys
# before = [str(m) for m in sys.modules]
# import folium
# after = [str(m) for m in sys.modules]
# set([m.split('.')[0] for m in after if not m in before and not m.startswith('_')])
# ```
#
# The loaded packages are:
#
# ```
# {'branca',
# 'certifi',
# 'chardet',
# 'cmath',
# 'csv',
# 'dateutil',
# 'encodings',
# 'folium',
# 'gzip',
# 'http',
# 'idna',
# 'importlib',
# 'jinja2',
# 'markupsafe',
# 'mmap',
# 'numpy',
# 'pandas',
# 'pkg_resources',
# 'pytz',
# 'requests',
# 'secrets',
# 'stringprep',
# 'urllib3',
# 'zipfile'}
# ```
#
# -
# The following packages seem to need installing in order load `folium`, along with folium itself:
#
# ```
# chardet, certifi, idna, branca, urllib3, Jinja2, requests, Markupsafe
# ```
#
# Universal wheels, with filenames of the form `PACKAGE-VERSION-py2.py3-none-any.whl` appearing in the *Download files* area of a PyPi package page ([example](https://pypi.org/project/requests/#files)] are required in order to install the package.
#
# One required package, [`Markupsafe`](https://pypi.org/project/Markupsafe/#files)) *did not* have a universal wheel available, so a wheel was manually built elsewhere (by hacking the [`setup.py` file](https://github.com/pallets/markupsafe/blob/main/setup.py) to force it to build the wheel in a platform and speedup free way) and pushed to a downloadable location in an [*ad hoc* wheelhouse](https://opencomputinglab.github.io/vce-wheelhouse/).
# +
# Install folium requirements
import micropip
await micropip.install("https://opencomputinglab.github.io/vce-wheelhouse/wheelhouse/MarkupSafe-2.0.1-py2.py3-none-any.whl")
await micropip.install('https://files.pythonhosted.org/packages/19/c7/fa589626997dd07bd87d9269342ccb74b1720384a4d739a1872bd84fbe68/chardet-4.0.0-py2.py3-none-any.whl')
await micropip.install('https://files.pythonhosted.org/packages/05/1b/0a0dece0e8aa492a6ec9e4ad2fe366b511558cdc73fd3abc82ba7348e875/certifi-2021.5.30-py2.py3-none-any.whl')
await micropip.install('https://files.pythonhosted.org/packages/d7/77/ff688d1504cdc4db2a938e2b7b9adee5dd52e34efbd2431051efc9984de9/idna-3.2-py3-none-any.whl')
await micropip.install("https://files.pythonhosted.org/packages/61/1f/570b0615c452265d57e4114e633231d6cd9b9d275256778a675681e4f711/branca-0.4.2-py3-none-any.whl")
await micropip.install('https://files.pythonhosted.org/packages/0c/cd/1e2ec680ec7b09846dc6e605f5a7709dfb9d7128e51a026e7154e18a234e/urllib3-1.26.5-py2.py3-none-any.whl')
await micropip.install('https://files.pythonhosted.org/packages/80/21/ae597efc7ed8caaa43fb35062288baaf99a7d43ff0cf66452ddf47604ee6/Jinja2-3.0.1-py3-none-any.whl')
await micropip.install('https://files.pythonhosted.org/packages/29/c1/24814557f1d22c56d50280771a17307e6bf87b70727d975fd6b2ce6b014a/requests-2.25.1-py2.py3-none-any.whl')
# Install folium
await micropip.install("https://files.pythonhosted.org/packages/c3/83/e8cb37afc2f016a1cf4caab8d22caf7fe4156c4c15230d8abc9c83547e0c/folium-0.12.1-py2.py3-none-any.whl")
# -
# ## Demo of `folium` Map
#
# Load in the `folium` package:
import folium
# And render a demo map:
m = folium.Map(location=[50.693848, -1.304734], zoom_start=11)
m
| examples/pyolite - folium.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import math
t = int(input())
for _ in range(t):
a, b = map(int, input().split())
a -= 1
b -= 1
c = 0
fac = abs(a-b)
i = 1
while i <= math.sqrt(fac):
if fac % i == 0:
if (fac // i == i):
c += 1
else:
c += 2
i += 1
print(c)
# -
t = int(input())
for _ in range(t):
a, b = map(int, input().split())
c = 0
for i in range(1, max(a, b)):
if a%i == b%i:
c += 1
print(c)
# +
a = 5
b = 12
lst = []
eq = []
for i in range(1, 100):
c = a%i
d = b%i
if c == d:
eq.append(i)
lst.append((i, c,d))
print(eq)
| CodeChef/EXAMCHT.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
pip install gspread_dataframe
import requests
url = "https://scholar.google.com/citations?hl=en&user=N5APA98AAAAJ"
r =requests.get(url)
text = r.text
text
from bs4 import BeautifulSoup
soup = BeautifulSoup(r.text, 'html.parser')
titles = []
authors = []
publications_date = []
cite = []
title = soup.find_all('td', {'class':'gsc_a_t'})
for title in title:
title = str(title).split('<td class="gsc_a_t"><a class="gsc_a_at"')
title = title[1]
title = title.split(' href="javascript:void(0)">')
title = title[1]
title = str(title).split('</a>')
title_f = title[0]
titles.append(title_f)
author = title[1]
author = author.split('<div class="gs_gray">')
author = author[1]
author = author.split('</div>')
author = author[0]
authors.append(author)
publication_date = soup.find_all('td', {'gsc_a_y'})
for year in publication_date:
year = str(year).split('<td class="gsc_a_y"><span class="gsc_a_h gsc_a_hc gs_ibl">')
year = year[1]
year = year.split('</span></td>')
year = year[0]
publications_date.append(year)
cite_by = soup.find_all('a', {'class':'gsc_a_ac gs_ibl'})
for cit in cite_by:
cit = str(cit).split('<a class="gsc_a_ac gs_ibl"')
cit =cit[1].split('>')
cit = cit[1].split('</a')
cit = cit[0]
cite.append(cit)
import pandas as pd
df = pd.DataFrame({
'title': titles,
'authors': authors,
'publication_date': publication_date,
'description':'',
'cite_by': cite_by
})
df
df.to_csv('Paper Table.csv')
# +
from selenium import webdriver
from selenium.webdriver.common.by import By
from selenium.webdriver.common.keys import Keys
from selenium.webdriver.support.ui import WebDriverWait
from selenium.webdriver.support import expected_conditions as EC
from selenium import webdriver
from webdriver_manager.chrome import ChromeDriverManager
PATH='./chromedriver'
driver = webdriver.Chrome(ChromeDriverManager().install())
#1 open homepage
driver.get("https://scholar.google.com/")
print(driver.title)
# -
#2 enter search box
search_box=driver.find_element_by_css_selector('input#gs_hdr_tsi.gs_in_txt.gs_in_ac')
search_box.send_keys("Thammasat University",Keys.ENTER)
#3 click thammasat university
driver.implicitly_wait(3)
driver.find_element_by_css_selector('div.gs_ob_inst_r')\
.find_element_by_css_selector('a').click()
#4 create dataframe
import pandas as pd
df=pd.DataFrame(
{
'user_ID':['N5APA98AAAAJ'],
'name':['<NAME>'],
'affiliation':['Thammasat university']
}
)
df
#5 for every author, get detail and save to df
while 1:
for i in driver.find_elements(By.CSS_SELECTOR,'div.gs_ai_t'):
author=i.find_element_by_css_selector('a')
aff=i.find_element_by_css_selector('div.gs_ai_aff')
print(
author.get_attribute('href').split('=')[-1],
author.text,
aff.text
)
df = df.append(
{
'user_ID': author.get_attribute('href').split('=')[-1],
'name': author.text,
'affiliation': aff.text
}
, ignore_index=True
)
driver.implicitly_wait(3)
x=driver.find_element_by_css_selector('#gsc_authors_bottom_pag > div > button.gs_btnPR.gs_in_ib.gs_btn_half.gs_btn_lsb.gs_btn_srt.gsc_pgn_pnx')
x.click()
df.tail(20)
#6 delete column Duplicate
df = df.drop_duplicates(subset="name")
df
#7 store df into csv file
df.to_csv('Authors Table.csv')
| data collecting process.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# (prototype) FX Graph Mode Quantization User Guide
# ===========================================================
#
# **Author**: `<NAME> <https://github.com/jerryzh168>`_
#
# FX Graph Mode Quantization requires a symbolically traceable model.
# We use the FX framework (TODO: link) to convert a symbolically traceable nn.Module instance to IR,
# and we operate on the IR to execute the quantization passes.
# Please post your question about symbolically tracing your model in `PyTorch Discussion Forum <https://discuss.pytorch.org/c/quantization/17>`_
#
# Quantization will only work on the symbolically traceable parts of your model.
# Data dependent control flow (if statements / for loops etc using symbolically traced values) are one common pattern which is not supported.
# If your model is not symbolically traceable end to end, you have a couple of options to enable FX Graph Mode Quantization only on a part of the model.
# You can use any combination of these options:
#
# 1. Non traceable code doesn’t need to be quantized
# a. Symbolically trace only the code that needs to be quantized
# b. Skip symbolic tracing the non-traceable code
#
# 2. Non traceable code needs to be quantized
# a. Refactor your code to make it symbolically traceable
# b. Write your own observed and quantized submodule
#
# If the code that is not symbolically traceable does not need to be quantized, we have the following two options
# to run FX Graph Mode Quantization:
#
# 1.a. Symbolically trace only the code that needs to be quantized
# -----------------------------------------------------------------
#
# When the whole model is not symbolically traceable but the submodule we want to quantize is
# symbolically traceable, we can run quantization only on that submodule.
#
#
# before:
#
# .. code:: python
#
# class M(nn.Module):
#
# def forward(self, x):
# x = non_traceable_code_1(x)
# x = traceable_code(x)
# x = non_traceable_code_2(x)
# return x
#
#
# after:
#
# .. code:: python
#
# class FP32Traceable(nn.Module):
#
# def forward(self, x):
# x = traceable_code(x)
# return x
#
# class M(nn.Module):
#
# def __init__(self):
# self.traceable_submodule = FP32Traceable(...)
#
# def forward(self, x):
# x = self.traceable_code_1(x)
# # We'll only symbolic trace/quantize this submodule
# x = self.traceable_submodule(x)
# x = self.traceable_code_2(x)
# return x
#
#
# quantization code:
#
# .. code:: python
#
# qconfig_dict = {"": qconfig}
# model_fp32.traceable_submodule = \
# prepare_fx(model_fp32.traceable_submodule, qconfig_dict)
#
# Note if original model needs to be preserved, you will have to
# # copy it yourself before calling the quantization APIs.
#
#
#
# 1.b. Skip symbolically trace the non-traceable code
# ---------------------------------------------------
# When we have some non-traceable code in the module, and this part of code doesn’t need to be quantized,
# we can factor out this part of the code into a submodule and skip symbolically trace that submodule.
#
#
# before
#
# .. code:: python
#
# class M(nn.Module):
#
# def forward(self, x):
# x = self.traceable_code_1(x)
# x = non_traceable_code(x)
# x = self.traceable_code_2(x)
# return x
#
#
# after, non-traceable parts moved to a module and marked as a leaf
#
# .. code:: python
#
# class FP32NonTraceable(nn.Module):
#
# def forward(self, x):
# x = non_traceable_code(x)
# return x
#
# class M(nn.Module):
#
# def __init__(self):
# ...
# self.non_traceable_submodule = FP32NonTraceable(...)
#
# def forward(self, x):
# x = self.traceable_code_1(x)
# # we will configure the quantization call to not trace through
# # this submodule
# x = self.non_traceable_submodule(x)
# x = self.traceable_code_2(x)
# return x
#
# quantization code:
#
# .. code:: python
#
# qconfig_dict = {"": qconfig}
#
# prepare_custom_config_dict = {
# # option 1
# "non_traceable_module_name": "non_traceable_submodule",
# # option 2
# "non_traceable_module_class": [MNonTraceable],
# }
# model_prepared = prepare_fx(
# model_fp32,
# qconfig_dict,
# prepare_custom_config_dict=prepare_custom_config_dict,
# )
#
# If the code that is not symbolically traceable needs to be quantized, we have the following two options:
#
#
# 2.a Refactor your code to make it symbolically traceable
# --------------------------------------------------------
# If it is easy to refactor the code and make the code symbolically traceable,
# we can refactor the code and remove the use of non-traceable constructs in python.
#
# More information about symbolic tracing support can be found in: (TODO: link)
#
# before:
#
# .. code:: python
#
# def transpose_for_scores(self, x):
# new_x_shape = x.size()[:-1] + (self.num_attention_heads, self.attention_head_size)
# x = x.view(*new_x_shape)
# return x.permute(0, 2, 1, 3)
#
#
# This is not symbolically traceable because in x.view(*new_x_shape)
# unpacking is not supported, however, it is easy to remove the unpacking
# since x.view also supports list input.
#
#
# after:
#
# .. code:: python
#
# def transpose_for_scores(self, x):
# new_x_shape = x.size()[:-1] + (self.num_attention_heads, self.attention_head_size)
# x = x.view(new_x_shape)
# return x.permute(0, 2, 1, 3)
#
#
# quantization code:
#
# This can be combined with other approaches and the quantization code
# depends on the model.
#
#
#
#
# 2.b. Write your own observed and quantized submodule
# -----------------------------------------------------
#
# If the non-traceable code can’t be refactored to be symbolically traceable,
# for example it has some loops that can’t be eliminated, like nn.LSTM,
# we’ll need to factor out the non-traceable code to a submodule (we call it CustomModule in fx graph mode quantization) and
# define the observed and quantized version of the submodule (in post training static quantization or quantization aware training for static quantization)
# or define the quantized version (in post training dynamic and weight only quantization)
#
#
# before:
#
# .. code:: python
#
# class M(nn.Module):
#
# def forward(self, x):
# x = traceable_code_1(x)
# x = non_traceable_code(x)
# x = traceable_code_1(x)
# return x
#
# after:
#
# 1. Factor out non_traceable_code to FP32NonTraceable
# non-traceable logic, wrapped in a module
#
# .. code:: python
#
# class FP32NonTraceable:
# ...
#
#
# 2. Define observed version of FP32NonTraceable
#
# .. code:: python
#
# class ObservedNonTraceable:
#
# @classmethod
# def from_float(cls, ...):
# ...
#
# 3. Define statically quantized version of FP32NonTraceable
# and a class method "from_observed" to convert from ObservedNonTraceable
# to StaticQuantNonTraceable
#
# .. code:: python
#
# class StaticQuantNonTraceable:
#
# @classmethod
# def from_observed(cls, ...):
# ...
#
#
# .. code:: python
#
# # refactor parent class to call FP32NonTraceable
# class M(nn.Module):
#
# def __init__(self):
# ...
# self.non_traceable_submodule = FP32NonTraceable(...)
#
# def forward(self, x):
# x = self.traceable_code_1(x)
# # this part will be quantized manually
# x = self.non_traceable_submodule(x)
# x = self.traceable_code_1(x)
# return x
#
#
# quantization code:
#
#
# .. code:: python
#
# # post training static quantization or
# # quantization aware training (that produces a statically quantized module)v
# prepare_custom_config_dict = {
# "float_to_observed_custom_module_class": {
# "static": {
# FP32NonTraceable: ObservedNonTraceable,
# }
# },
# }
#
# model_prepared = prepare_fx(
# model_fp32,
# qconfig_dict,
# prepare_custom_config_dict=prepare_custom_config_dict)
#
# calibrate / train (not shown)
#
# .. code:: python
#
# convert_custom_config_dict = {
# "observed_to_quantized_custom_module_class": {
# "static": {
# ObservedNonTraceable: StaticQuantNonTraceable,
# }
# },
# }
# model_quantized = convert_fx(
# model_prepared,
# convert_custom_config_dict)
#
# post training dynamic/weight only quantization
# in these two modes we don't need to observe the original model, so we
# only need to define thee quantized model
#
# .. code:: python
#
# class DynamicQuantNonTraceable: # or WeightOnlyQuantMNonTraceable
# ...
# @classmethod
# def from_observed(cls, ...):
# ...
#
# prepare_custom_config_dict = {
# "non_traceable_module_class": [
# FP32NonTraceable
# ]
# }
#
#
# .. code:: python
#
# # The example is for post training quantization
# model_fp32.eval()
# model_prepared = prepare_fx(
# model_fp32,
# qconfig_dict,
# prepare_custom_config_dict=prepare_custom_config_dict)
#
# convert_custom_config_dict = {
# "observed_to_quantized_custom_module_class": {
# "dynamic": {
# FP32NonTraceable: DynamicQuantNonTraceable,
# }
# },
# }
# model_quantized = convert_fx(
# model_prepared,
# convert_custom_config_dict)
#
# You can also find examples for custom modules in test ``test_custom_module_class`` in ``torch/test/quantization/test_quantize_fx.py``.
#
#
| docs/_downloads/85aac427463b59681a558c91a0017d72/fx_graph_mode_quant_guide.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import sys
import pandas
import matplotlib
import seaborn
import sklearn
print(sys.version)
print(pandas.__version__)
print(matplotlib.__version__)
print(seaborn.__version__)
print(sklearn.__version__)
# -
x = 4
print x
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.model_selection import train_test_split
games = pandas.read_csv("scrapers.csv")
print(games.columns)
print("------------------")
print(games.shape)
plt.hist(games["average_rating"])
plt.show()
#print the first row of all games with zero scores
print(games[games["average_rating"]==0].iloc[0])
#print the first row of games with scores > 0
print(games[games["average_rating"]>0].iloc[0])
#remove rows without any reviews
games = games[games["users_rated"] > 0]
#remove any rows with missing values
games = games.dropna(axis=0)
# make a histogram of all avg ratings
plt.hist(games["average_rating"])
plt.show()
print(games.columns)
# +
#correlation matrix
cormat = games.corr()
fig = plt.figure(figsize=(12,9))
sns.heatmap(cormat,vmax=.8,square =True)
plt.show()
# +
#get all columns from the dataframe
columns = games.columns.tolist()
#filter the columns to remove data we dont want
columns = [c for c in columns if c not in ["bayes_average_rating","average_rating", "type", "name", "id"]]
#store the var we ll be predicting on
target = "average_rating"
#generate training & test datasets
from sklearn.model_selection import train_test_split
train = games.sample(frac=0.8, random_state = 1)
#select anything not in the training set and put it in test
test = games.loc[~games.index.isin(train.index)]
#print shapes
print(train.shape)
print(test.shape)
# -
#import linear regression model first
from sklearn.linear_model import LinearRegression
from sklearn.metrics import mean_squared_error
#initlize model class
LR = LinearRegression()
LR.fit(train[columns], train[target])
#generate predictions for testing set
predictions = LR.predict(test[columns])
#compute error btw our test predictions and actual values
mean_squared_error(predictions,test[target])
# +
#import the random forest model
from sklearn.ensemble import RandomForestRegressor
#init model
RFR = RandomForestRegressor(n_estimators = 100,min_samples_leaf =10,random_state=1)
#fit to the data
RFR.fit(train[columns],train[target])
# -
predictions = RFR.predict(test[columns])
# +
#compute the error btw our test predictions and actual values
mean_squared_error(predictions, test[target])
#make prediction with both models
rating_lr = LR.predict(test[columns].iloc[0].values.reshape(1,-1))
rating_rfr = RFR.predict(test[columns].iloc[0].values.reshape(1,-1))
#print out the predictions
print(rating_lr)
print(rating_rfr)
print("actual value from the test dataset: " + str(test[target].iloc[0] ) )
# -
| board_game_review_prediction/board game review prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # K Closest Points to Origin
#
# We have a list of points on the plane. Find the K closest points to the origin (0, 0).
#
# (Here, the distance between two points on a plane is the Euclidean distance.)
#
# You may return the answer in any order. The answer is guaranteed to be unique (except for the order that it is in.)
#
#
# ```
# Example 1:
#
# Input: points = [[1,3],[-2,2]], K = 1
# Output: [[-2,2]]
# Explanation:
# The distance between (1, 3) and the origin is sqrt(10).
# The distance between (-2, 2) and the origin is sqrt(8).
# Since sqrt(8) < sqrt(10), (-2, 2) is closer to the origin.
# We only want the closest K = 1 points from the origin, so the answer is just [[-2,2]].
# Example 2:
#
# Input: points = [[3,3],[5,-1],[-2,4]], K = 2
# Output: [[3,3],[-2,4]]
# (The answer [[-2,4],[3,3]] would also be accepted.)
#
#
# Note:
#
# 1 <= K <= points.length <= 10000
# -10000 < points[i][0] < 10000
# -10000 < points[i][1] < 10000
# ```
#
# [K Closest Points to Origin](https://leetcode.com/problems/k-closest-points-to-origin/)
#
class Solution:
def kClosest(self, points, K):
"""
:type points: List[List[int]]
:type K: int
:rtype: List[List[int]]
"""
pointMap = {}
pointVals = []
for point in points:
x = abs(point[0])
y = abs(point[1])
value = x * x + y * y
if value in pointMap:
pointMap[value].append(point)
else:
pointMap[value] = [point]
pointVals.append(value)
pointVals.sort()
result = []
for value in pointVals:
if len(result) < K:
result.extend(pointMap[value])
else:
break
return result
| KClosestPointstoOrigin.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy
from matplotlib import pyplot
# %matplotlib inline
pyplot.rcParams['font.family'] = 'serif'
pyplot.rcParams['font.size'] = 16
# +
# parameters
L = 1.0
nx = 51
dx = L / (nx-1)
alpha = 1.22e-3 # thermal diffusivity
x = numpy.linspace(0.0, L, num=nx)
# initial condition
T0 = numpy.zeros(nx)
T0[0] = 100.0
# -
def ftcs(T0, nt, dt, dx, alpha):
T = T0.copy()
sigma = alpha * dt / dx**2
for n in range(nt):
T[1:-1] = (T[1:-1] +
sigma * (T[2:] - 2.0 * T[1:-1] + T[:-2]))
return T
# +
# set time steps based on CFL limit
nt = 100
sigma = 0.5
dt = sigma * dx**2 / alpha
# compute
T = ftcs(T0, nt, dt, dx, alpha)
# -
def plot_it(x, y, x_lim1, x_lim2, y_lim1, y_lim2):
pyplot.figure(figsize=(6.0, 4.0))
pyplot.xlabel('Distance [m]')
pyplot.ylabel('Temperature [C]')
pyplot.grid()
pyplot.plot(x, y, linewidth=2)
pyplot.xlim(x_lim1, x_lim2)
pyplot.ylim(y_lim1, y_lim2);
plot_it(x, T, 0.0, L, 0.0, 100.0)
# ## Boundary Conditions
# ### Dirichlet
# +
nt = 1000
T = ftcs(T0, nt, dt, dx, alpha)
# -
plot_it(x, T, 0.0, L, 0.0, 100.0)
def ftcs_mixed_bcs(T0, nt, dt, dx, alpha):
T = T0.copy()
sigma = alpha * dt / dx ** 2
for n in range(nt):
T[1:-1] = T[1:-1] + sigma * (T[2:] - 2.0 * T[1:-1] + T[:-2])
# Neumann condition
T[-1] = T[-2]
return T
# +
nt = 1000
T = ftcs_mixed_bcs(T0, nt, dt, dx, alpha)
plot_it(x, T, 0.0, L, 0.0, 100.0)
# -
| .ipynb_checkpoints/Week8 Part 1 Lesson 04_01-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # 1) Regression fundamentals
# ## 1) Data and model
# - x: input
# - y: output
# - f(x): functional relationship, expected relationship between x and y
#
# $y_{i} = f(x_{i}) + e_{i}$
# - $e_{i}$: error term
#
# You can easily imagine that there are errors in this model, because you can have two houses that have exactly the same number of square feet, but sell for very different prices because they could have sold at different times. They could have had different numbers of bedrooms, or bathrooms, or size of the yard, or specific location, neighborhoods, school districts. Lots of things that we might not have taken into account in our model.
# <img src="images/lec1_pic01.png">
#
# *Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-regression/lecture/M6cMZ/regression-fundamentals-data-model) 6:45*
#
# <!--TEASER_END-->
# ## 2) Block diagram
# - $\hat{y}$: predicted house sales price
# - $\hat{f}$: estimated function
# - y: actual sales price
#
# we're gonna compare the actual sales price to the predicted sales price using the any given $\hat{f}$. And the quality metric will tell us how well we do. So there's gonna be some error in our predicted values. And the machine learning algorithm we're gonna use to fit these regression models is gonna try to minimize that error. So it's gonna search over all these functions to reduce the error in these predicted values.
# <img src="images/lec1_pic02.png">
#
# *Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-regression/lecture/fKsPh/regression-ml-block-diagram) 3:48*
#
# <!--TEASER_END-->
# # 2) The simple linear regression model, its use, and interpretation
# ## 1) The simple linear regression model
# **What's the equation of a line? **
# - it's just (intercept + slope * our variable of interest) so that we're gonna say that's $f(x) = w_{0} + w_{1}x$
#
# And what this regression model then specifies is that each one of our observations $y_{i}$ is simply that function evaluated at $x_{i}$, so:
# $$y_{i} = w_{0} + w_{1}x + \epsilon_{i}$$
#
# - $\epsilon_{i}$: error term, the distance from our specific observation back down to the line
# - $w_{0}, w_{1}$: intercept and slope respectively, they are called regression coefficients.
# <img src="images/lec1_pic03.png">
#
# *Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-regression/lecture/N8p7w/the-simple-linear-regression-model) 1:12*
#
# <!--TEASER_END-->
# ## 2) The cost of using a given line
# **What is the difference between residual and error?**
#
# https://www.coursera.org/learn/ml-regression/lecture/WYPGc/the-cost-of-using-a-given-line/discussions/Lx0xn5j1EeW0dw6k4EUmPw
# - Residual is the difference between the observed value and the predicted value. Error is the difference between the observed value and the (often unknown) true value. As such, residuals refer to samples whereas errors refer to populations.
# - There is a true function f(x) that we want to learn, and the observed values $y_{i}$ we have are in fact: $y_{i} = f(x_{i}) + e_{i}$, because we need to assume our measures have some error (or noise if you prefer this term). This $e_{i}$ is the real error, because the real value is $f(x_{i})$. In the other hand, the residual is $y_{i} - \hat f(x_{i})$, where $\hat f$ is our approximation (estimation) of the real f(x)
# **Residual sum of squares (RSS)**
#
# The sum of all the differences between predicted values and actual values and then square the sum.
# <img src="images/lec1_pic04.png">
# <img src="images/lec1_pic05.png">
#
# *Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-regression/lecture/WYPGc/the-cost-of-using-a-given-line) 3:26*
#
# <!--TEASER_END-->
# ## 3) Using the fitted line
# - a model is in terms of sum parameters and a fitted line is a specific example within that model class
#
# **Why the hat notation?**
#
# https://www.coursera.org/learn/ml-regression/lecture/RjYbf/using-the-fitted-line/discussions/QOsWrZkGEeWKNwpBrKr_Fw
#
# - In statistics, the hat operator is used to denote the predicted value of the parameter.
#
# eg: Y-hat stands for the predicted values of Y (house-price).
#
# http://mathworld.wolfram.com/Hat.html
#
# https://en.wikipedia.org/wiki/Hat_operator
#
# - The hat denotes a predicted value, as contrasted with an observed value. For our purposes right now, I think of the hat value as the value that sits on the regression line, because that's the value our regression analysis would predict. So, for example, the residual for a particular observation is y_i minus y_ihat, where y_i is the actual observed outcome at a particular observed value of x and y_ihat is the value that our regression analysis predicts for y at that same x value.
# <img src="images/lec1_pic06.png">
#
# *Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-regression/lecture/RjYbf/using-the-fitted-line) 2:00*
#
# <!--TEASER_END-->
# ## 4) Interpreting the fitted line
# - $\hat{w}$: predicted changed in the output per unit change in the input.
#
# One thing I want to make very, very clear is that the magnitude of this slope, depends both on the units of our input, and on the units of our output. So, in this case, the slope, the units of slope, are dollars per square feet. And so, if I gave you a house that was measured in some other unit, then this coefficient would no longer be appropriate for that.
#
# For example, if the input is square feet and you have another house was measured in square meters instead of square feet, well, clearly I can't just plug into that equation.
# <img src="images/lec1_pic07.png">
#
# *Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-regression/lecture/x8ohF/interpreting-the-fitted-line) 3:54*
#
# <!--TEASER_END-->
# # 3) An aside on optimization: one dimensional objectives
# ## 1) Defining our least squares optimization objective
# $RSS(w_{0}, w_{1}) = g(w_{0}, w_{1})$
#
# Our objective here is to minimize over all possible combinations of $w_{0}$, and $w_{1}$
# <img src="images/lec1_pic08.png">
#
# *Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-regression/lecture/a1QCT/defining-our-least-squares-optimization-objective) 2:35*
#
# <!--TEASER_END-->
# ## 2) Finding maxima or minima analytically
# - Concave function: The way we can define a concave function is we can look at any two values of w: a, and b. Then we draw a line between a and b, that line lies below g(w) everywhere.
# - Convex function: Opposite properties of Concave function where the line connects g(a) and g(b) is above g(w) everywhere.
# - There a functions which are neither Concave nor Convex function where the line lies both below and above the g(w) function.
# <img src="images/lec1_pic09.png">
#
# *Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-regression/lecture/RUtxG/finding-maxima-or-minima-analytically) 3:49*
#
# <!--TEASER_END-->
# - In a concave function, at the point of max g(w), this is the point where the derivative = 0. Same thing for convex function, if we want to find minimum of all w over g(w), at the minimum point, the derivative = 0.
#
# Example: $g(w) = 5 - (w - 10)^{2}$
#
# $\frac{d g(w)}{d w} = 0 - 2(w-10)^{2} \cdot 1 = -2w + 20$
#
# When we draw this derivative, we can see it has a concave form. **How do I find this maximum?**
# - I take this derivative and set it equal to 0, and we can solve it for w = 10.
# <img src="images/lec1_pic10.png">
#
# *Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-regression/lecture/RUtxG/finding-maxima-or-minima-analytically) 6:45*
#
# <!--TEASER_END-->
# ## 3) Finding the max via hill climbing
# If we're looking at these concave situations and our interest is in finding the max over all w of g(w) one thing we can look at is something called a **hill-climbing algorithm**. Where it's going to be an integrative algorithm where we start somewhere in this space of possible w's and then we keep changing w hoping to get closer and closer to the optimum.
#
# Okay, so, let's say that we start w here. And a question is well **should I increase w, move w to the right or should I decrease w and move w to the left to get closer to the optimal. **
# - What I can do is I can look at the function at w and I can take the derivative and if the derivative is positive like it is here, this is the case where I want to increase w. If the derivative is negative, then I want to decrease w.
# - So, we can actually divide the space into two.
# - Where on the left of the optimal, we have that the derivative of g with respect to w is greater than 0. And these are the cases where we're gonna wanna increase w.
# - And on the right-hand side of the optimum we have that the derivative of g with respect to w is negative. And these are cases where we're gonna wanna decrease w.
# - If I'm exactly at the optimum, which maybe I'll call $w^{*}$. I do not want to move to the right or the left, because I want to stay at the optimum. The derivative at this point is 0.
#
# So, again, the derivative is telling me what I wanna do. We can write down this climbing algorithm:
# - While not converged, I'm gonna take my previous w, where I was at iteration t, so t is the iteration counter. And I'm gonna move in the direction indicated by the derivative. So, if the derivative of the function is positive, I'm going to be increasing w, and if the derivative is negative, I'm going to be decreasing w, and that's exactly what I want to be doing.
# - But instead of moving exactly the amount specified by the derivative at that point, we can introduce something, I'll call it eta ($\eta$). And $\eta$ is what's called a step size.
# - So, when I go to compute my next w value, I'm gonna take my previous w value and I'm going to move and amount based on the derivative as determined by the step size.
#
# Example: Let's say I happen to start on this left hand side at this w value here. And at this pointthe derivative is pretty large. This function's pretty steep. So, I'm going to be taking a big step. Then, I compute the derivative. I'm still taking a fairly big step. I keep stepping increasing. What I mean by each of these is I keep increasing w. Keep taking a step in w. Going, computing the derivative and as I get closer to the optimum, the size of the derivative has decreased.
# <img src="images/lec1_pic11.png">
#
# *Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-regression/lecture/O4j1e/finding-the-max-via-hill-climbing) 3:34*
#
# <!--TEASER_END-->
# ## 4) Finding the min via hill descent
# We can use the same type of algorithm to find the minimum of a function.
#
# **When the derivative is positive we want to decrease w and when the derivative is negative, you wanna increase w**
#
# The update of the hill descent algorithm is gonna look almost exactly the same as the hill climbing, except we have the minus sign, so we're going to move in the opposite direction.
# <img src="images/lec1_pic12.png">
#
# *Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-regression/lecture/zVcGn/finding-the-min-via-hill-descent) 2:40*
#
# <!--TEASER_END-->
# ## 5) Choosing stepsize and convergence criteria
# So in these algorithms, I said that there's a stepsize. Stepsize is denoted as $\eta$. This determines how much you're changing your W at every iteration of this algorithm.
#
# One choice you can look at is something called fixed stepsize or constant stepsize, where, as the name implies, you just set eta equal to some constant. So for example, maybe 0.1.
#
# But what can happen is that you can jump around quite a lot. I keep taking these big steps. And I end up jumping over the optimal to a point here and then I jump back and then I'm going back and forth, and back and forth. And I converge very slowly to the optimal itself.
# <img src="images/lec1_pic13.png">
#
# *Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-regression/lecture/3UvFZ/choosing-stepsize-and-convergence-criteria) 2:00*
#
# <!--TEASER_END-->
# **A common choice is to decrease the stepsize as the number of iterations increase. **
#
# One thing you have to be careful about is not decreasing the stepsize too rapidly. Because if you're doing that, you're gonna, again, take a while to converge. Because you're just gonna be taking very, very, very small steps. Okay, so in summary choosing your stepsize is just a bit of an art.
# <img src="images/lec1_pic14.png">
#
# *Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-regression/lecture/3UvFZ/choosing-stepsize-and-convergence-criteria) 4:30*
#
# <!--TEASER_END-->
# **How are we going to assess our convergence? **
#
# Well, we know that the optimum occurs when the derivative of the function is equal to 0.
# $$\frac{dg(w)}{dw} = 0$$
#
# But what we're gonna see in practice, is that the derivative, it's gonna get smaller and smaller, but it won't be exactly 0. At some point, we're going to want to say, okay, that's good enough. We're close enough to the optimum. I'm gonna terminate this algorithm.
#
# In practice, stop when
# $$\left|\frac{dg(w)}{dw}\right| < \epsilon$$
#
# $\epsilon$ is a threshold I'm setting. Then if this is satisfied, then I'm gonna terminate the algorithm and return whatever solution I have $w^{(t)}$. So in practice, we're just gonna choose epsilon to be very small. And I wanna emphasize that what
# very small means depends on the data that you're looking at, what the form of this function is.
# <img src="images/lec1_pic15.png">
#
# *Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-regression/lecture/3UvFZ/choosing-stepsize-and-convergence-criteria) 5:30*
#
# <!--TEASER_END-->
# # 4) An aside on optimization: multidimensional objectives
# ## 1) Gradients: derivatives in multiple dimensions
# So up until this point we've talked about functions of just one variable and finding there minimum or maximum. But remember when we were talking about residual sums of squares, we had two variables. Two parameters of our model, w0 and w1. And we wanted to minimize over both.
#
# Let's talk about how we're going to move functions defined over multiple variables. Moving to multiple dimensions here, where when we have these functions in higher dimensions we don't talk about derivatives any more we talk about gradients in their place.
#
# **What is a gradient?**
# - $\bigtriangledown g(w)$: notation of gradient of a function
# - w: vector of different w's $[w_{0},w_{1},...,w_{p}]$
#
# $$\bigtriangledown g(w) =
# \begin{bmatrix}
# \frac{\partial g}{\partial w_{0}}
# \\ \frac{\partial g}{\partial w_{1}}
# \\ ...
# \\ \frac{\partial g}{\partial w_{p}}
# \end{bmatrix}$$
#
# The definition of a gradient: it's gonna be a vector, where we're gonna look at what are called the partial derivatives of g. We're going to look at the partial with respect to W zero. The partial of G with respect to W one. W one all the way up to the partial of G with respect to some WP.
#
# It's exactly like a derivative where we're taking the derivative with respect to in this case W one. But what are we going to
# do with all the other W's? W zero, W two, W three all the up to WP? Well we're just going to treat them like constants.
#
# So the vector represents a (p + 1), cause we're indexing starting at zero. This is a (p+1) -dimensional vector.
# <img src="images/lec1_pic16.png">
#
# *Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-regression/lecture/ZwU5b/gradients-derivatives-in-multiple-dimensions) 3:05*
#
# <!--TEASER_END-->
# **Work through Example**
#
# We have a function:
# $$g(w) = 5w_{0} + 10w_{0}w_{1} + 2w_{1}^{2} $$
#
#
# Let's compute the gradient of w
# - partial of G with respect to W zero: $\frac{\partial g}{\partial w_{0}} = 5 + 10w_{1}$, in this case we treat $w_{1}$ like a constant.
# - partial of G with respect to W one: $\frac{\partial g}{\partial w_{1}} = 10w_{0} + 4w_{1}$, where $w_{0}$ is a constant in this case.
#
# This is my gradient:
# $$\bigtriangledown g(w) =
# \begin{bmatrix}
# 5 + 10w_{1}
# \\ 10w_{0} + 4w_{1}
# \end{bmatrix}$$
#
# If I want to look at the gradient at any point on this surface well I'm just going to plug in whatever the W one and W zero values are at this point, and I'm going to compute the gradient. And it'll be some vector. It's just some number in the first component, some number in the second component, and that forms some factor.
# <img src="images/lec1_pic17.png">
#
# *Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-regression/lecture/ZwU5b/gradients-derivatives-in-multiple-dimensions) 5:08*
#
# <!--TEASER_END-->
# ## 2) Gradient descent: multidimensional hill descent
# Instead of looking at these 3D mesh plots that we've been looking at, we can look at a contour plot, where we can kind of think of this as a bird's eye view.
# <img src="images/lec1_pic18.png">
#
# *Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-regression/lecture/6PJ3h/gradient-descent-multidimensional-hill-descent) 2:37
#
# <!--TEASER_END-->
# Let's talk about the gradient descent algorithm, which is the analogous algorithm to what I call the hill decent algorithm in 1D.
#
# But, in place of the derivative of the function, we've now specified the gradient of the function. And other than that, everything looks exactly the same. So what we're doing, is we're taking we now have a vector of parameters, and we're updating them all at once. We're taking our previous vector and we're updating with our sum, eta times our gradient which was also a vector. So, it's just the vector analog of the hill descent algorithm.
#
# If we take a look at the picture, we start at a point where the gradient is actually pointing in the direction of steepest assent (up hill). But we're moving in the negative gradient direction.
#
#
#
#
# <img src="images/lec1_pic19.png">
#
# *Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-regression/lecture/6PJ3h/gradient-descent-multidimensional-hill-descent) 5:30
#
# <!--TEASER_END-->
# # 5) Finding the least squares line
# ## 1) Computing the gradient of RSS
# Now, we can think about applying these optimization notions and optimization algorithms that we described to our specific scenario of interest. Which is searching over all possible lines and finding the line that best fits our data.
#
# So the first thing that's important to mention is the fact that our objective is Convex. And what this is implies is that the solution to this minimization problem is unique. We know there's a unique minimum to this function. And likewise, we know that our gradient descent algorithm will converge to this minimum.
# <img src="images/lec1_pic20.png">
#
# *Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-regression/lecture/PRcIZ/computing-the-gradient-of-rss) 1:20
#
# <!--TEASER_END-->
# Let's return to the definition of our cost, which is the residual sum of squares of our two parameters, (wo,w1), which I've written again right here.
#
# But before we get to taking the gradient, which is gonna be so crucial for our gradient descent algorithm, let's just note the following fact about derivatives, where if we take the derivative of the sum of functions over some parameter, some variable w. So N different functions, g1 to gN, the derivative distributes across the sum. And we can rewrite this as the sum of the derivative, okay? So the derivative of the sum of functions is the same as the sum of the derivative of the individual functions so in our case, we have that $g_{i}(w)$.
#
# The $g_{i}(w)$ that I'm writing here is equal to:
# $$ g_{i}(w) = (y_{i} - [w_{0} + w_{1} x_{i}])^{2}$$
#
# And we see that the residual sum of squares is indeed a sum over n different functions of w0 and w1. And so in our case when we're thinking about taking the partial of the residual sum of squares. With respect to, for example w0, this is going to be equal:
# $$\frac{\partial RSS(w)}{\partial w_{0}} = \sum_{i=1}^n \frac{\partial}{\partial w_{0}}(y_{i} - [w_{0} + w_{1} x_{i}])^{2}$$
#
# And the same holds for W1.
# <img src="images/lec1_pic21.png">
#
# *Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-regression/lecture/PRcIZ/computing-the-gradient-of-rss) 3:45
#
# <!--TEASER_END-->
# **Let's go ahead and actually compute this gradient**
#
# And the first thing we're gonna do is take the derivative or the partial with respect to the W0.
#
# Okay, so I'm gonna use this fact that, I showed on the previous slide to take the sum to the outside. And then I'm gonna take the partial with respect to the inside. So, I have a function raised to a power. So i'm gonna bring that power down. I'm gonna get a 2 here, rewrite the function. That's W1 XI, and now the power is just gonna be 1 here. But then, I have to take the derivative of the inside. And so what's the derivative of the inside when I'm taking this derivative with respect to W0? Well, what I have is I have a -1 multiplying W0, and everything else I'm just treating as a constant. So, I need to multiply by -1.
# <img src="images/lec1_pic22.png">
#
# *Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-regression/lecture/PRcIZ/computing-the-gradient-of-rss) 4:58
#
# <!--TEASER_END-->
# **Let's go ahead and now take the derivative or the partial with respect to W1**
#
# So in this case, again I'm pulling the sum out same thing happens where I'm gonna bring the 2 down. And I'm gonna rewrite the function here, the inside part of the function, raise it just to the 1 power. And then, when I take the derivative of this part, this inside part, with respect to W1, What do I have? Well all of these things are constants with respect to W1, but what's multiplying W1? I have a (-xi) so I'm going to need to multiply by (-xi).
# <img src="images/lec1_pic23.png">
#
# *Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-regression/lecture/PRcIZ/computing-the-gradient-of-rss) 5:58
#
# <!--TEASER_END-->
# **Let's put this all together**
#
# So this is the gradient of our residual sum of squares, and it's a vector of two dimensions because we have two variables, w0 and w1. Now what can w think about doing? Well of course we can think about doing the gradient descent algorithm. But let's hold off on that because what do we know is another way to solve for the minimum of this function?
#
# Well we know we can, just like we talked about in one D, taking the derivative and setting it equal to zero, that was the first approach for solving for the minimum. Well here we can take the gradient and set it equal to zero.
# <img src="images/lec1_pic24.png">
#
# *Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-regression/lecture/PRcIZ/computing-the-gradient-of-rss) 7:13
#
# <!--TEASER_END-->
# ## 2) Approach 1: closed-form solution
# So this is gonna be our Approach 1. And this is drawn here on this 3D mesh plot, where that green surface is the gradient at the minimum. And what we see is that's where the gradient = 0. And that red dot is the, the optimal point that we're gonna be looking at.
#
# Let's take this gradient, set it equal to zero and solve for W0 and W1. Those are gonna be our estimates of our two parameters of our model that define our fitted line.
#
# I'm gonna take the top line and I'm going to set it equal to 0. The reason I'm putting the hats on are now, these are our solutions. These are our estimated values of these parameters.
# ** Break down the math **
#
# https://www.coursera.org/learn/ml-regression/lecture/G9oBu/approach-1-closed-form-solution/discussions/Ywv6RZfxEeWNbBIwwhtGwQ
#
#
# First lets make our notation shorter:
#
# $\sum\limits_{i=1}^N=\Sigma$
#
# ** Top term (Row 1)**
#
# - 1) Break summation down: $$-2(\Sigma y_i-\Sigma w_0 - \Sigma w_1 x_i) = 0$$
# - 2) Divide both sides by -2: $$\Sigma y_i-\Sigma w_0 - \Sigma w_1 x_i = 0$$
# - 3) Summation of a constant replacement: $$\Sigma y_i - Nw_0 - \Sigma w_1 x_i = 0$$
# - 4) Solve for $w_0$: $$Nw_0 = \Sigma y_i - \Sigma w_1 x_i$$
# - 5) Divide by N: $$w_0 = \frac{\Sigma y_i}{N} - \frac{\Sigma w_1 x_i}{N}$$
# - 6) Move $w_1$ constant out: $$w_0 = \frac{\Sigma y_i}{N} - w_1\frac{\Sigma x_i}{N}$$
# - 7) Put party hats on $w_0$ and $w_1$: $$\hat{w_0} = \frac{\Sigma y_i}{N} - \hat{w_1}\frac{\Sigma x_i}{N}$$
#
# ** Bottom term (Row 2)**
# - 1) Factor in $x_i$: $$-2[\Sigma( y_i x_i - w_0 x_i - w_1 x_i^2)]= 0$$
# - 2) Break summation down: $$-2[\Sigma y_i x_i - \Sigma w_0 x_i - \Sigma w_1 x_i^2]= 0$$
# - 3) Divide both sides by -2: $$\Sigma y_i x_i - \Sigma w_0 x_i - \Sigma w_1 x_i^2 = 0$$
# - 4) Move constants $w_0$ and $w_1$ out: $$\Sigma y_i x_i - w_0\Sigma x_i - w_1\Sigma x_i^2 = 0$$
# - 5) Replace $w_0$ with equation from Row 1, Line 6: $$\Sigma y_i x_i - \frac{\Sigma y_i}{N} \Sigma x_i + w_1\frac{\Sigma x_i}{N}\Sigma x_i - w_1\Sigma x_i^2 = 0$$
# - 6) Multiply by 1 namely N/N to remove denominator of N: $$\frac{N}{N}\Sigma y_i x_i - \frac{\Sigma y_i}{N} \Sigma x_i + w_1\frac{\Sigma x_i}{N}\Sigma x_i - w_1\frac{N}{N}\Sigma x_i^2 = 0$$
# - 7) Group numerator and multiply By N to remove denominator: $$N\Sigma y_i x_i - \Sigma y_i \Sigma x_i + w_1\Sigma x_i\Sigma x_i - w_1N\Sigma x_i^2 = 0$$
# - 8) Group $w_1$ terms: $$w_1(\Sigma x_i\Sigma x_i - N\Sigma x_i^2) = \Sigma y_i \Sigma x_i - N\Sigma y_i x_i$$
# - 9) Solve for $w_1$: $$w_1 =\frac{(\Sigma y_i \Sigma x_i - N\Sigma y_i x_i)}{(\Sigma x_i\Sigma x_i - N\Sigma x_i^2)}$$
# - 10) Divide top and bottom by -N (Same as multiply by 1): $$w_1 = \frac{(\Sigma y_i x_i - \frac{\Sigma y_i \Sigma x_i}{N})}{( \Sigma x_i^2 - \frac{\Sigma x_i\Sigma x_i}{N})}$$
# - 11) Put party hat on $w_1$: $$\hat{w_1} = \frac{(\Sigma y_i x_i - \frac{\Sigma y_i \Sigma x_i}{N})}{( \Sigma x_i^2 - \frac{\Sigma x_i\Sigma x_i}{N})}$$
#
# <img src="images/lec1_pic25.png">
#
# *Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-regression/lecture/G9oBu/approach-1-closed-form-solution) 5:11
#
# <!--TEASER_END-->
# ## 3) Approach 2: gradient descent
# We discussed the other approach that we can take is to do Gradient descent where we're walking down this surface of residual sum of squares trying to get to the minimum. Of course we might over shoot it and go back and forth but that's the general idea that
# we're doing this iterative procedure. And in this case it's useful to reinterpret this gradient of the residual sum of squares that we computed previously.
#
# A couple notation:
# - $y_i$: actual house sales observation
# - $(w_0 + w_1 x_i)$: predicted value, but I am gonna write it as a function of $w_0$ and $w_1$, to make it clear that it's the prediction I'm forming when using $w_0$ and $w_1$, so: $\hat y_i(w_0,w_1)$
#
#
# <img src="images/lec1_pic26.png">
#
# *Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-regression/lecture/Ifx9C/approach-2-gradient-descent) 1:30
#
# <!--TEASER_END-->
# **Then we can write our gradient descent algorithm**
#
# while not converged, we're gonna take our previous vector of W0 at iteration T, W1 at iteration T and We're going to subtract eta times the gradient.
# <img src="images/lec1_pic27.png">
#
# *Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-regression/lecture/Ifx9C/approach-2-gradient-descent) 3:25
#
# <!--TEASER_END-->
# **We can also rewrite the algorithm. **
#
# I want it in this form to provide a little bit of intuition here. Because what happens if overall, we just tend to be underestimating our values y?
#
# So, if overall, we're under predicting $\hat y_i$, then we're gonna have that the sum of yi- y hat i is going to be positive ($\sum [y_i - \hat y_i]$ is positive). Because we're saying that $\hat y_i$ is always below, or in general, below the true value $y_i$, so this is going to be positive.
#
# And what's gonna happen? Well, this term here $\sum [y_i - \hat y_i]$ is positive. We're multiplying by a positive thing, and adding that to our vector W. So $w_0$ is going to increase. And that makes sense, because we have some current estimate of our regression fit. But if generally we're under predicting our observations that means probably that line is too low. So, we wanna shift it up. That means increasing $w_0$.
#
# So, there's a lot of intuition in this formula for what's going on in this gradient descent algorithm. And that's just talking about this first term $w_0$, but then there's this second term $w_1$, which is the slope of the line. And in this case there's
# a similar intuition. But we need to multiply by this $x_i$, accounting for the fact that this is a slope term.
#
# <img src="images/lec1_pic28.png">
#
# *Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-regression/lecture/Ifx9C/approach-2-gradient-descent) 6:00
#
# <!--TEASER_END-->
# ## 4) Comparing the approaches
# Let's take a moment to compare the two approaches that we've gone over, either setting the gradient equal to zero or doing gradient descent.
#
#
# Well, in the case of minimizing residual sum of squares, we showed that both were fairly straight forward to do. But in a lot of the machine learning method's that we're interested in taking the gradient and setting it equal to zero, well there's just no close form solution to that problem. So, often we have to turn to method's like gradient descent.
#
# And likewise, as we're gonna see in the next module, where we turn to having lots of different inputs, lots of different features in our regression. Even though there might be a close form solution to setting the gradient equal to zero, sometimes in practice it can be much more efficient computationally to implement the gradient descent approach.
#
# And likewise, as we're gonna see in the next module, where we turn to having lots of different inputs, lots of different features in our regression. Even though there might be a close form solution to setting the gradient equal to zero, sometimes in practice it can be much more efficient computationally to implement the gradient descent approach.
# <img src="images/lec1_pic29.png">
#
# *Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-regression/lecture/5oBEV/comparing-the-approaches) 1:20
#
# <!--TEASER_END-->
# # 6) Discussion and summary of simple linear regression
# ## 1) Asymmetric cost functions
# Let's discuss about the intuition of what happens if we use a different measure of error.
#
# Okay so this residual sum of squares that we've been looking at is something that's called a Symmetric cost function. And that's because what we're assuming when we look at this aerometric is the fact that if we over estimate the value of our house, that has the same cost as if we under estimate the value of the house.
# <img src="images/lec1_pic30.png">
#
# *Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-regression/lecture/JdRnN/asymmetric-cost-functions) 2:41
#
# <!--TEASER_END-->
# What happens if there might not be symmetric cost to these error?
#
# But what if the cost of listing my house sales price as too high is bigger than the cost if I listed it as too low?
# - If I list the price too high, there will be no offers.
# - If I list the price too low, I still get offer, but not as high as I could have if I had more accurately estimated the value of the house.
#
# So in this case it might be more appropriate to use an asymmetric cost function where the errors are not weighed equally between these two types of mistakes. So if we choose asymmetric cost, I prefer to underestimate the value than over.
# <img src="images/lec1_pic31.png">
#
# *Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-regression/lecture/JdRnN/asymmetric-cost-functions) 3:18
#
# <!--TEASER_END-->
# ## 2) A brief recap
# <img src="images/lec1_pic32.png">
#
# *Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-regression/lecture/UdYun/a-brief-recap) 0:45
#
# <!--TEASER_END-->
| machine_learning/2_regression/lecture/week1/.ipynb_checkpoints/Linear Regression-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Initial Analysis and Plots
# This section looks at the data and draws simple conclusions from it as used in the submitted paper
import pandas as pd
pd.set_option('display.max_columns', 50)
df = pd.read_csv("data.csv")
df.shape
df.head()
df.events.unique()
df['events'].value_counts()/sum(df['events'].value_counts())
# Only using subset as dataset is so large
df = df[(df['year']==2017) & ((df['month']==6)|(df['month']==7)|(df['month']==8))]
df.describe()
# %matplotlib inline
x= df['from_station_name'].value_counts()
x=x[x>10000]
x.plot(kind='bar')
import seaborn as sns
sns.distplot(df['tripduration'])
# 0 is Monday
df.groupby(['hour']).count()[['trip_id']].plot(kind='bar',figsize=(8,4))
# 0 is Monday
df.groupby(['day']).count()[['trip_id']].plot(kind='bar',figsize=(10,10))
df['Location'] = df[['latitude_start','longitude_start']].apply(tuple, axis=1)
# +
# link to reference for mapping
# https://towardsdatascience.com/exploring-and-visualizing-chicago-transit-data-using-pandas-and-bokeh-part-ii-intro-to-bokeh-5dca6c5ced10
import math
def merc(Coords):
lat = Coords[0]
lon = Coords[1]
r_major = 6378137.000
x = r_major * math.radians(lon)
scale = x/lon
y = 180.0/math.pi * math.log(math.tan(math.pi/4.0 +
lat * (math.pi/180.0)/2.0)) * scale
return (x, y)
df['coords_x'] = df['Location'].apply(lambda x: merc(x)[0])
df['coords_y'] = df['Location'].apply(lambda x: merc(x)[1])
# -
total_start = df.groupby(['coords_x','coords_y'], as_index = False).count()[['coords_x','coords_y','Location']]
total_start['circle_size'] = total_start['Location'] / 750
# Plot Locations of stops
from bokeh.plotting import figure, show, output_notebook
from bokeh.tile_providers import CARTODBPOSITRON
p = figure(x_range=(-9780000, -9745000), y_range=(5130000, 5160000))
p.add_tile(CARTODBPOSITRON)
p.circle(x = total_start['coords_x'],
y = total_start['coords_y'])
output_notebook()
show(p)
# Plot popularity of locations
from bokeh.plotting import figure, show, output_notebook
from bokeh.tile_providers import CARTODBPOSITRON
p = figure(x_range=(-9780000, -9745000), y_range=(5130000, 5160000))
p.add_tile(CARTODBPOSITRON)
p.circle(x = total_start['coords_x'],
y = total_start['coords_y'],
size=total_start['circle_size'],
line_color="#FF0000",
fill_color="#FF0000",
fill_alpha=0.05)
output_notebook()
show(p)
# ## Model
# This section will be for processing data to fit model and then evaluate the model
df_model = df[['month','week','day','hour','temperature','events','from_station_name','tripduration']]
df_model.head()
# creates binary outcome for duration of greater than 10 minutes.
df_model['duration'] = np.where(df['tripduration']>df['tripduration'].mean(),1,0)
# creates binary clear or not clear for events
df_model.loc[df_model['events']=='cloudy','events']='clear'
df_model.loc[df_model['events']=='rain or snow','events']='not clear'
df_model.loc[df_model['events']=='tstorms','events']='not clear'
df_model = df_model[df_model['events']!='unknown']
df_model.head()
counts = df_model['from_station_name'].value_counts()
repl = counts[counts <= 9000].index
dummy = pd.get_dummies(df.from_station_name.replace(repl,'other'))
df_model = df_model.drop(['from_station_name','tripduration'], axis = 1)
df_model.head()
# data for model leaves out one to avoid multicollinearity
data_model = df_model.join(dummy.loc[:,'Canal St & Madison St':])
#moves duration binary to last column
data_model = data_model[['month', 'week', 'day', 'hour', 'temperature', 'events',
'Canal St & Madison St', 'Clinton St & Madison St',
'Clinton St & Washington Blvd', 'Columbus Dr & Randolph St',
'Daley Center Plaza', 'Dearborn St & Erie St',
'Franklin St & Monroe St', 'Kingsbury St & Kinzie St',
'Lake Shore Dr & North Blvd', 'Michigan Ave & Washington St',
'Orleans St & Merchandise Mart Plaza', 'Streeter Dr & Grand Ave',
'Theater on the Lake', 'other', 'duration']]
data_model['events'] = np.where(data_model['events']=='clear',1,0)
# Head of processed data before splitting into train and test
data_model.head()
# +
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split
from sklearn import metrics
X = data_model.iloc[:,:-1]
y = data_model['duration']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3)
model = LogisticRegression()
model.fit(X_train, y_train)
# -
y_pred = model.predict(X_test)
model.score(X_test, y_test)
from sklearn.metrics import confusion_matrix
confusion_matrix = confusion_matrix(y_test, y_pred)
print(confusion_matrix)
# +
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
from sklearn import metrics
X = data_model.iloc[:,:-1]
y = data_model['duration']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3)
model = RandomForestClassifier()
model.fit(X_train, y_train)
y_pred = model.predict(X_test)
model.score(X_test,y_test)
# -
from sklearn.metrics import confusion_matrix
confusion_matrix = confusion_matrix(y_test, y_pred)
print(confusion_matrix)
| project1/Lang-project1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # DRUN Deep Q-learning driving network (regular observations)
# A Deep Q-Learning network which uses the Microsoft AirSim simulation wrapped in OpenAI gym enviourment class for training, practising navigation from point A on a map to point B without colliding.
# ### Library imports
# #### Custom Open AI gym
# Installing our custom "airsim_gym" gym enviourment package.
# !pip install -e airsim_gym
# #### Other libraries
# Importing all the libraries used in the project.
# +
from __future__ import absolute_import
from collections import namedtuple, deque
from math import exp
import gym
import numpy as np
import tensorflow as tf
from tensorflow.keras import Model
from tensorflow.keras.layers import Input, Conv2D, Flatten, Dense, Activation, Concatenate
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.losses import Huber
# -
# ### Connect to environment
env = gym.make("airsim_gym:airsim-regular-v0")
# ### Hyper-parameters
# +
# Model hyperparameters
STATE_SIZE = [256, 256, 4]
ACTION_SIZE = env.action_space.n
STACK_SIZE = 64
LEARNING_RATE = 0.0002
# Training parameters
TOTAL_EPISODES = 5000
MAX_STEPS = 1000
BATCH_SIZE = 64
PRETRAIN_LENGTH = BATCH_SIZE
MEMORY_SIZE = 1000000
UPDATE_AFTER_ACTIONS = 4
# Epsilon greedy
EXPLORE_START = 1.0
EXPLORE_STOP = 0.01
DECAY_RATE = 0.0001
# Q-learning hyperparameters
GAMMA = 0.95
# Script execution
TRAINING = True
ENV_PREVIEW = False
# -
# ### Environment preview
if ENV_PREVIEW:
env.reset()
for _ in range(10):
env.step(env.action_space.sample())
# ### Image processing utilities
# #### prepocess_frame
# Preprocessing in order to reduce the complexity of our states and consecutively to reduce the computation time needed for training.
#
def preprocess_frame(frame):
# Converts frame from RGB to grayscale
grayscale_frame = np.mean(frame, -1)
# Normalize Pixel Values
normalized_frame = grayscale_frame/255.0
return normalized_frame
# #### stack_frames
# Stacking frames in or to crate a sense of motion to our Neural Network.
def stack_frames(stacked_frames, state, is_new_episode: bool, stack_size: int = STACK_SIZE):
# Preprocess frame
frame = preprocess_frame(state)
if is_new_episode:
# Clear our stacked_frames
stacked_frames = [np.zeros(STATE_SIZE[:2], dtype=np.int) for i in range(stack_size)]
stacked_frames = deque(stacked_frames, maxlen=stack_size)
# In a new episode the deque is filled with the same frame
for _ in range(stack_size):
stacked_frames.append(frame)
else:
# Append frame to deque, pops the last
stacked_frames.append(frame)
# Build the stacked state (first dimension specifies different frames)
stacked_state = np.stack(stacked_frames, axis=2)
return stacked_state, stacked_frames
stacked_frames = deque([np.zeros(STATE_SIZE[:2], dtype=np.int) for i in range(STACK_SIZE)], maxlen=4)
# ### Replay memory
# Create the Memory object that contains a deque. A deque (double ended queue) is a data type that removes the oldest element each time that you add a new element over the size limit.
# #### Expiriance replay
Experience = namedtuple(
"Experience",
("observation", "position", "action", "next_observation", "next_position", "reward")
)
# #### Define replay memory class
class ReplayMemory():
def __init__(self, capacity):
self.buffer = deque(maxlen=capacity)
self.push_count = 0
def add(self, experience):
self.buffer.append(experience)
self.push_count += 1
def sample(self, batch_size):
buffer_size = len(self.buffer)
index = np.random.choice(
np.arange(buffer_size),
size=batch_size,
replace=False
)
return [self.buffer[i] for i in index]
def is_sample_available(self, batch_size):
return len(self.buffer) >= batch_size
# #### Agent class
# #### Initialize replay memory
# +
replay_memory = ReplayMemory(MEMORY_SIZE)
observation_stack = None
next_observation_stack = None
next_action = None
done = False
observation = None
for i in range(PRETRAIN_LENGTH):
if i == 0:
# If no state is available, we get one from the reset
start_observation, position = env.reset()
_, observation_stack = stack_frames(
observation_stack,
start_observation,
True,
)
_, next_observation_stack = stack_frames(
next_observation_stack,
start_observation,
True,
)
# Random action
if (observation is None):
action = env.action_space.sample()
observation, position, reward, done = env.step(action)
_, next_observation_stack = stack_frames(
next_observation_stack,
observation,
False,
)
_, observation_stack = stack_frames(
observation_stack,
observation,
False,
)
# Hit something
if done:
print("done")
# Empty frame on episode ending
next_observation = np.zeros(STATE_SIZE[:2], dtype=np.float32)
_, next_observation_stack = stack_frames(
next_observation_stack,
next_observation,
False,
)
next_position = position
# Add experience to memory
replay_memory.add(
Experience(
observation_stack,
position,
action,
next_observation_stack,
next_position,
reward,
),
)
# Start a new episode
start_observation, position = env.reset()
_, observation_stack = stack_frames(
observation_stack,
start_observation,
True,
)
_, next_observation_stack = stack_frames(
next_observation_stack,
start_observation,
True,
)
observation = None
position = None
done = False
else:
# Get the next state
next_observation, next_position, next_action, next_done = env.step(action)
_, next_observation_stack = stack_frames(
next_observation_stack,
observation,
False,
)
# Add experience to memory
replay_memory.add(
Experience(
observation_stack,
position,
action,
next_observation_stack,
next_position,
reward,
),
)
# Our state is now the next_observation
observation = next_observation
position = next_position
done = next_done
# -
# ### Epsilon greedy strategy
# $\epsilon$ select a random action $a_t$, otherwise select $a_t = \mathrm{argmax}_a Q(s_t,a)$. Over time the exploration probability decays in favour of the exploatation rate.
class EpsilonGreedy():
def __init__(self, start, stop, decay):
self.start = start
self.stop = stop
self.decay = decay
def get_exploration_rate(self, current_step):
rate = self.stop + (self.start - self.stop)
rate *= exp(-1 * current_step * self.decay)
return rate
def predict_action(self, current_step, observation, position, env, dqn):
# Randomizing a number
exp_exp_tradeoff = np.random.rand()
explore_probability = self.get_exploration_rate(current_step)
if explore_probability < exp_exp_tradeoff:
# A random action is sampled
action = env.action_space.sample()
else:
# Get action from Q-network (exploitation)
# Estimate the Qs values state
observation = np.array(observation)
position = np.array(position)
observation = observation.reshape(1, *observation.shape)
position = position.reshape(1, *position.shape)
print(observation.shape, position.shape)
prediction = dqn.predict([observation, position])
# Take the biggest Q value (= the best action)
action = np.argmax(prediction)
return action, explore_probability
epsilon = EpsilonGreedy(EXPLORE_START, EXPLORE_START, DECAY_RATE)
# ### Deep Q-learning network
# This is our Deep Q-learning model:
#
# We take a stack of 4 frames and two normalized coordinates as input:
# - Image is passed through 3 CNN layers
# - Then it is concatinated with the coordinates
# - Finally it passes through 3 FC layers
# - Outputs a Q value for each actions
def drun_dqn() -> Model:
image_input = Input(STATE_SIZE)
coords_input = Input(2)
img_net = Conv2D(32, (4, 4), strides=(4, 4), activation="relu", padding="same", input_shape=STATE_SIZE)(image_input)
img_net = Conv2D(64, (3, 3), strides=(2, 2), activation="relu", padding="same")(img_net)
img_net = Conv2D(64, (3, 3), strides=(2, 2), activation="relu", padding="same")(img_net)
img_net = Flatten()(img_net)
combined = Concatenate(axis=1)
combined = combined([img_net, coords_input])
dense_net = Dense(512, activation=tf.nn.relu)(combined)
dense_net = Dense(512, activation=tf.nn.relu)(dense_net)
dense_net = Dense(512, activation=tf.nn.relu)(dense_net)
output = Dense(ACTION_SIZE, activation=tf.nn.elu)(dense_net)
return Model(inputs=(image_input, coords_input), outputs=output)
# +
model = drun_dqn()
optimizer = Adam(learning_rate=LEARNING_RATE, clipnorm=1.0)
loss_function = Huber()
model.summary()
# -
# ### Network training
# Standart Q-learning algorithm:
#
# 1. Initialize replay memory capacity.
# 2. Initialize the policy network with random weights.
# 3. Clone the policy network, and call it the target network.
# 4. For each episode:
# 1. Initialize the starting state.
# 2. For each time step:
# 1. Select an action.
# - Via exploration or exploitation
# 2. Execute selected action in an emulator.
# 3. Observe reward and next state.
# 4. Store experience in replay memory.
# 5. Sample random batch from replay memory.
# 6. Preprocess states from batch.
# 7. Pass batch of preprocessed states to policy network.
# 8. Calculate loss between output Q-values and target Q-values.
# - Requires a pass to the target network for the next state
# 9. Gradient descent updates weights in the policy network to minimize loss.
# - After time steps, weights in the target network are updated to the weights in the policy network.
if TRAINING:
decay_step = 0
for episode in range(TOTAL_EPISODES):
episode_step = 0
episode_rewards = []
observation, position = env.reset()
observation, stacked_frames = stack_frames(stacked_frames, observation, True)
while episode_step < MAX_STEPS:
# Increase episode_decay/decay_steps
episode_step += 1
decay_step += 1
# Predict the action to take and take it
action, explore_probability = epsilon.predict_action(decay_step, observation, position, env, model)
# Do the action
observation, position, reward, done = env.step(action)
observation = preprocess_frame(observation)
# Add the reward to total reward
episode_rewards.append(reward)
# If the game is finished
if done:
# Empty frame on episode ending
next_observation = np.zeros(observation.shape)
next_position = [0.0, 0.0]
# Add experience to memory
replay_memory.add(Experience(stacked_frames, position, action, next_observation, next_position, reward, done))
# Start a new episode
observation, position = env.reset()
# Stack the frames
observation, stacked_frames = stack_frames(stacked_frames, observation, True)
# Set episode_step = max_steps to end the episode
episode_step = MAX_STEPS
# Get the total reward of the episode
total_reward = np.sum(episode_rewards)
print("Episode: {}".format(episode),
"Total reward: {}".format(total_reward),
"Explore probability: {:.4f}".format(explore_probability))
replay_memory.add(Experience(stacked_frames, position, action, next_observation, next_position, reward, done))
else:
# Get the next state
next_observation, next_position = env.get_state()
next_observation, stacked_frames = stack_frames(stacked_frames, next_observation, False)
# Add experience to memory
replay_memory.add(Experience(stacked_frames, position, action, next_observation, next_position, reward, done))
# st+1 is now our current state
observation = next_observation
# LEARNING PART
# Obtain random mini-batch from memory
if episode_step % UPDATE_AFTER_ACTIONS == 0 and replay_memory.is_sample_available(BATCH_SIZE):
batch = replay_memory.sample(BATCH_SIZE)
observation_mb = np.array([item.observation for item in batch])
observation_mb = np.rollaxis(observation_mb, 1, observation_mb.ndim)
position_mb = np.array([item.position for item in batch])
actions_mb = np.array([item.action for item in batch])
next_observations_mb = np.array([item.next_observation for item in batch])
next_positions_mb = np.array([item.next_position for item in batch])
rewards_mb = np.array([item.reward for item in batch])
dones_mb = np.array([item.done for item in batch])
print(observation.shape, position_mb.shape)
target_Qs_batch = []
# Build the updated Q-values for the sampled future states
# Use the target model for stability
future_rewards = model.predict([observation_mb, position_mb])
# Q value = reward + discount factor * expected future reward
updated_q_values = rewards_mb + GAMMA * tf.reduce_max(
future_rewards, axis=1
)
# If final frame set the last value to -1
updated_q_values = updated_q_values * (1 - dones_mb) - dones_mb
# Create a mask so we only calculate loss on the updated Q-values
masks = tf.one_hot(actions_mb, ACTION_SIZE)
# Train the model on the states and updated Q-values
q_values = model([observation_mb, position_mb])
# Apply the masks to the Q-values to get the Q-value for action taken
q_action = tf.reduce_sum(tf.multiply(q_values, masks), axis=1)
# Calculate loss between new Q-value and old Q-value
loss = loss_function(updated_q_values, q_action)
print("Training loss: {:.4f}".format(loss))
# Backpropagation
grads = tape.gradient(loss, model.trainable_variables)
optimizer.apply_gradients(zip(grads, model.trainable_variables))
# Save model every 10 episodes
if episode % 10 == 0:
model.save("model/")
| ai/train_regular.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 데이터 불러오기
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import datetime as dt
from sklearn.ensemble import RandomForestClassifier
from xgboost import XGBClassifier
from xgboost import plot_importance
import warnings
warnings.filterwarnings('ignore')
# %matplotlib inline
# +
def str_col(df):
col = []
for i in range(0,len(df.dtypes)):
if str(df.dtypes[i]) == 'object':
col.append(df.dtypes.index[i])
print(col)
return col
def int_col(df):
col = []
for i in range(0,len(df.dtypes)):
if str(df.dtypes[i]) != 'object':
col.append(df.dtypes.index[i])
print(col)
return col
def p_100(a, b):
print( round( (a/(a+b))*100,2), "%" )
def extraction_func(df, col_name, num_list):
temp = pd.DataFrame()
for i in num_list:
temp = pd.concat([ temp, df.loc[df[col_name] == i ] ],axis=0)
return temp
# + tags=[]
scaled_insurance = pd.read_csv('./temp_data/save_scaled_insurance.csv',encoding='utf-8')
print(scaled_insurance.shape)
print(scaled_insurance.dtypes)
print(scaled_insurance.isnull().sum())
scaled_insurance.tail(5)
# -
# ##### 데이터 복사
copy_insurance = scaled_insurance.copy()
# ## 훈련데이터와 검증데이터로 분리
# +
from sklearn.model_selection import train_test_split
label = 'SIU_CUST_YN'
x_train, x_test, y_train, y_test = train_test_split(copy_insurance[copy_insurance.columns.drop(label)], copy_insurance[label],\
test_size=0.2, random_state=42)
train_0 = y_train.to_frame().loc[y_train.to_frame()['SIU_CUST_YN'] == 'N'].count()
train_1 = y_train.to_frame().loc[y_train.to_frame()['SIU_CUST_YN'] == 'Y'].count()
test_0 = y_test.to_frame().loc[y_test.to_frame()['SIU_CUST_YN'] == 'N'].count()
test_1 = y_test.to_frame().loc[y_test.to_frame()['SIU_CUST_YN'] == 'Y'].count()
x_train.to_csv('./temp_data/save_x_train.csv',index = True)
y_train.to_csv('./temp_data/save_y_train.csv',index = True)
x_test.to_csv('./temp_data/save_x_test.csv',index = True)
y_test.to_csv('./temp_data/save_y_test.csv',index = True)
print(train_0)
print(train_1)
print(test_0)
print(test_1)
# +
y_train = y_train.replace('Y',1)
y_train = y_train.replace('N',0)
y_test = y_test.replace('Y',1)
y_test = y_test.replace('N',0)
plt.hist(y_train)
plt.xticks([0,1])
plt.show()
print(y_train.value_counts())
print(p_100(y_train.value_counts()[1],y_train.value_counts()[0]))
plt.hist(y_test)
plt.xticks([0,1])
plt.show()
print(y_test.value_counts())
print(p_100(y_test.value_counts()[1],y_test.value_counts()[0]))
# -
# ## 군집 전 의사결정에 영향이 매우 적은 변수들을 제외하기 위해 feature importance 확인
# - 현재 변수가 79개로 3,4차 군집을 해도 군집이 되자 않음
# +
f_extraction = pd.concat([x_train, y_train], axis=1)
x_train_2, x_test_2, y_train_2, y_test_2 = train_test_split(f_extraction[f_extraction.columns.drop('SIU_CUST_YN')], f_extraction['SIU_CUST_YN'], test_size=0.2, random_state=42)
# + tags=[]
# 원래 여기 데이터에는 검증 데이터를 넣어야함 Test 데이터 넣으면 안됨!
# 검증 데이터 넣어주어서 교차검증 해보도록하기
evals = [(x_test_2, y_test_2)]
xgb_wrapper = XGBClassifier(n_estimators=400, learning_rate=0.1,
max_depth=3, random_state=42)
# eval_metric넣어주면서 검증 데이터로 loss 측정할 때 사용할 metric 지정
xgb_wrapper.fit(x_train_2, y_train_2, early_stopping_rounds=200,
eval_set=evals, eval_metric='logloss')
preds = xgb_wrapper.predict(x_test_2)
preds_proba = xgb_wrapper.predict_proba(x_test_2)[:, 1]
print(preds_proba[:10])
# -
# feature별 중요도 시각화하기
fig, ax = plt.subplots(figsize=(15,15))
plot_importance(xgb_wrapper, ax)
# ##### 필요한 컬럼만 추출
x_train_FI = x_train[['DISTANCE','PAYM_AMT','CHLD_CNT','SUM_ORIG_PREM','TOTALPREM','HOUSE_HOSP_DIST','RCBASE_HSHD_INCM','AGE','CUST_RGST','CLAIM_NUM','CUST_INCM',\
'DMND_RESN_CODE_1', 'DMND_RESN_CODE_2','DMND_RESN_CODE_3', 'DMND_RESN_CODE_4', 'DMND_RESN_CODE_5','DMND_RESN_CODE_6',\
'DMND_RESN_CODE_7', 'DMND_RESN_CODE_9','ACCI_DVSN_1', 'ACCI_DVSN_2', 'ACCI_DVSN_3','SEX_남', 'SEX_여','CUST_ROLE_0',\
'CUST_ROLE_1', 'CUST_ROLE_2', 'CUST_ROLE_21', 'CUST_ROLE_3','CUST_ROLE_4', 'CUST_ROLE_5']]
x_train_FI.to_csv('./temp_data/save_feature_importance.csv',index = True)
| 3. feature_extraction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Before starting this notebook
# # !cd ..
# +
import base64
# Local
from image_net_labels import MAPPING
# 3rd Party
from PIL import Image
import requests
from io import BytesIO
from baybars.timber import get_logger
import requests
# +
# asyncs are removed due to https://github.com/jupyter/notebook/issues/3397
# +
logger = get_logger('resnet_client')
# The server URL specifies the endpoint of your server running the ResNet
# model with the name "resnet" and using the predict interface.
RESNET_STATUS_URL = 'http://localhost:8501/v1/models/resnet'
SERVER_URL = 'http://localhost:8501/v1/models/resnet:predict'
# The image URL is the location of the image we should send to the server
IMAGE_URL = 'https://tensorflow.org/images/blogs/serving/cat.jpg'
def get_as_base64(image_url):
out = None
response = requests.get(image_url)
if response.status_code == 200:
out = base64.b64encode(response.content)
return out
# -
def main():
out = []
response = requests.get(RESNET_STATUS_URL)
logger.info('resnet status: {}'.format(response.json()))
image_content = get_as_base64(IMAGE_URL)
predict_request = {
'instances': [{'b64': image_content.decode()}],
}
# Send few requests to warm-up the model.
for _ in range(3):
response = requests.post(SERVER_URL, json=predict_request)
response.raise_for_status()
# Send few actual requests and report average latency
total_time = 0
num_requests = 100
for _ in range(num_requests):
response = requests.post(SERVER_URL, json=predict_request)
response.raise_for_status()
total_time += response.elapsed.total_seconds()
prediction = response.json()['predictions'][0]
out.append(prediction)
logger.info('Prediction class: {}, avg latency: {} ms'.format(
prediction['classes'], (total_time*1000)/num_requests))
return out
predictions = main()
response = requests.get(IMAGE_URL)
Image.open(BytesIO(response.content))
MAPPING[predictions[0]['classes']]
| notebooks/2. Rest API Example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Форматирование в С++
# В лекции:
# * С-шный форматированный вывод
# * Потоки в С++
# * `boost::format`
# * fmtlib
# * `std::format` в C++20
# * сравнение билиотек
# <br />
# ##### naive C
# https://en.cppreference.com/w/cpp/io/c
#
# https://en.cppreference.com/w/cpp/io/c/fprintf
# В чистом С для форматирования есть семейство функций `printf`.
#
# Они поддерживают набор форматов для встроенных типов:
# * целые числа
# * вещественные числа
# * С-строки
# * указатели
#
# ```c++
# printf("%i < %s < %.2f", 3, "pi", 3.15f);
# ```
# **Вопрос**: какие преимущества и недостатки у этого способа?
#
# <details>
# <summary>преимущества</summary>
# <p>
#
# * быстрое исполнение (сравнительно)
# * быстрая скорость компиляции
# * маленький размер бинарного кода
# * мало зависимостей (требуется только libc)
#
# </p>
# </details>
#
# <details>
# <summary>недостатки</summary>
# <p>
#
# * обязанность программиста следить за соответствием типов и форматов (много ошибок)
#
# </p>
# </details>
# <br />
# C-шный вариант позволяет записывать результат не только в `FILE*`, но и в строковый буфер:
#
# ```c++
# char buf[256];
# std::sprintf(buf, "%i < %s < %.2f", 3, "pi", 3.15f);
# ```
#
# Это весьма удобный С-шный способ формировать форматированные строки.
#
# **Вопрос**: какой недостаток вы видите в этом способе?
#
# <details>
# <summary>ответ</summary>
# <p>
#
# Функция `sprintf` не может проконтроллировать выход за границы буфера, программист обязан умозрительно гарантировать, что буфера хватит. Конечно, программисты иногда ошибаются.
#
# </p>
# </details>
# <br />
# Чтобы справиться с проблемой, в С++11 добавили `snprintf` - форматированный вывод в строку с контролем длины:
#
# ```c++
# // форматируем строку с указанием размера буфера,
# // std::snprintf проконтроллирует выход за границы массива
# char buf[256];
# int rv = std::snprintf(buf, 256, "%i < %s < %.2f", 3, "pi", 3.15f);
# // теперь надо разобрать код возврата
#
# // отрицательные значения - индикатор ошибки внутри snprintf
# if (rv < 0)
# std::cout << "formatting error" << std::endl;
#
# // значения < 256 - значит null-terminated output успешно записано в буфер
# if (rv < 256)
# std::cout << "succeed, symbols written: " << rv << std::endl;
#
# // значение >= 256 - сколько символов без нуля нужно в буфере,
# // чтобы полностью записать строку
# if (rv >= 256)
# std::cout << "buffer is too small, required size is: " << rv << std::endl;
# ```
# Трюк с предварительным определением нужного размера:
#
# ```c++
# // оценим, сколько символов требуется, чтобы записать форматированную строку
# const int sz = std::snprintf(nullptr, 0, "sqrt(2) = %f", std::sqrt(2));
#
# // выделим на куче нужное кол-во символов +1 для нуль-терминатора
# std::vector<char> buf(sz + 1);
#
# // собственно, само форматирование
# std::snprintf(&buf[0], buf.size(), "sqrt(2) = %f", std::sqrt(2));
# ```
#
# Такой подход безопаснее, но, увы, требуется дважды выполнить полное форматирование строки:
# * сначала чтобы узнать размер
# * затем чтобы записать результат
# <br />
# ##### C++ streams
# https://en.cppreference.com/w/cpp/io/manip
# Стандартные С++ потоки позволяют задавать форматирование:
# Форматирование `bool`:
#
# ```c++
# std::cout << std::boolalpha
# << "true: " << true << std::endl
# << "false: " << false << std::endl;
# std::cout << std::noboolalpha
# << "true: " << true << std::endl
# << "false: " << false << std::endl;
# ```
#
# Вывод:
#
# ```sh
# true: true
# false: false
# true: 1
# false: 0
# ```
# Основа системы счисления для `oct/hex` чисел:
#
# ```c++
# std::cout << std::hex
# << "42 = " << std::showbase << 42 << std::endl
# << "42 = " << std::noshowbase << 42 << std::endl;
# ```
#
# Вывод:
#
# ```sh
# 42 = 0x2a
# 42 = 2a
# ```
# Форматирование `float`:
#
# ```c++
# const long double pi = std::acos(-1.L);
# std::cout << "default precision (6): " << pi << std::endl
# << "std::setprecision(10): " << std::setprecision(10) << pi << std::endl;
#
# std::cout << "fixed: " << std::fixed << 0.01 << std::endl
# << "scientific: " << std::scientific << 0.01 << std::endl
# << "hexfloat: " << std::hexfloat << 0.01 << std::endl
# << "default: " << std::defaultfloat << 0.01 << std::endl;
# ```
#
# Вывод:
#
# ```sh
# default precision (6): 3.14159
# std::setprecision(10): 3.141592654
#
# fixed: 0.010000
# scientific: 1.000000e-02
# hexfloat: 0x1.47ae147ae147bp-7
# default: 0.01
# ```
# И некоторые другие способы отформатировать значения, смотрите их в документации по ссылке.
#
# А лучше не смотрите, т.к. далее мы рассмотрим более продвинутые способы.
# <br />
# ##### boost::format
# https://www.boost.org/doc/libs/1_66_0/libs/format/doc/format.html
# Способы форматирования, поддерживаемые `boost::format`:
#
# * `%spec`, spec - [printf specification](https://www.boost.org/doc/libs/1_66_0/libs/format/doc/format.html#printf_directives)
#
# boost::format позволяет эмулировать обычный сишный `printf` [за небольшим числом хитрых исключений](https://www.boost.org/doc/libs/1_66_0/libs/format/doc/format.html#printf_differences)
#
# ```c++
# std::cout << boost::format("%i <= pi <= %f") % 3 % 3.15 << std::endl;
# ```
#
# * `%|spec|`, spec - [printf specification](https://www.boost.org/doc/libs/1_66_0/libs/format/doc/format.html#printf_directives)
#
# У этого способа есть две особенности:
# * авторы boost утверждают, что он улучшает читабельность (возможно, это так)
# * не требуется указание типа:
#
# ```c++
# // все результаты ниже имеют ширину 5 и выровнены влево
# boost::format("%|-5|") % 3.14;
# boost::format("%|-5|") % 3;
# boost::format("%|-5|") % "3.1"s;
# ```
#
# * `%N%` - позиционные аргументы
#
# ```c++
# boost::format("%1% <= pi <= %2%") % 3 % 3.14;
# ```
# <br />
# ##### поддержка пользовательских типов
# Чтобы вывести пользоветальский тип, достаточно определить для него `operator<<`, здесь всё естественно:
#
# ```c++
# struct Point
# {
# float x;
# float y;
# };
#
# std::ostream& operator << (std::ostream& os, const Point& p)
# {
# return os << '(' << p.x << ',' << p.y << ')';
# }
#
# Point p{3, 4};
# std::cout << boost::format("Point of interest is %1%") % p;
# ```
# <br />
# ##### что такое `boost::format`?
# `boost::format` - это объект-`formatter`:
#
# ```c++
# // создали объект-formatter
# boost::format f("%1% %2% %3% %1%");
#
# // скормили объекту аргументы
# f % 10 % 20 % 30;
#
# // после того как formatter насытили аргументами,
# // можно спрашивать с него результат, например, через <<
# std::cout << f; // "10 20 30 10"
#
# // можно спрашивать с него результат несколько раз
# std::cout << f; // "10 20 30 10"
# std::string s = f.str();
#
# // можно начать его насыщать аргументами заново
# f % 1001;
# try
# {
# std::cout << f;
# }
# catch (const boost::io::too_few_args&)
# {
# std::cout << "Formatter is not fed" << std::endl;
# }
#
# // насытив formatter, можно снова спрашивать с него результат
# std::cout << f % "abc" % "def";
#
# // можно через методы объекта модифицировать спецификацию формата
# f = boost::format("%1% %2% %3% %2% %1%");
# f.modify_item(4, boost::io::group(std::setfill('_'),
# std::hex,
# std::showbase,
# std::setw(5)));
# std::cout << f % 1 % 2 % 3; // "1 2 3 __0x2 1 \n"
#
# // задавать нумерованные аргументы по номеру явно
# f = boost::format("%1% %2% %3% %2% %1%");
# f.bind_arg(1, "x");
# f.bind_arg(2, "y");
# f.bind_arg(3, "z");
# std::cout << f;
#
# // поведение аргументов, заданных через bind и %, различно:
# // * через bind - привязанные (bounded) аргументы
# // * через % - регулярные (regular) аргументы
# f = boost::format("%1% %2% %3% %2% %1%");
# f.bind_arg(1, "10");
# f.bind_arg(1, "11"); // перезадали аргумент по номеру
# std::cout << f % 2 % 3; // "11 2 3 2 11"
#
# try
# {
# // бросит исключение, потому что аргумент N1 привязанный
# // значит через % насыщаются только "регулярные" N2 и N3
# std::cout << f % 6 % 7 % 8;
# }
# catch (const boost::io::too_many_args&)
# {
# std::cout << "Formatter is fed out";
# }
#
# // очистка регулярных аргументов
# f.clear();
#
# // очитска и регулярных, и связанных аргументов
# f.clear_binds();
# ```
# <br />
# `boost::format` - богатый и удобный инструмент. Код с его использованием становится короче, понятнее и читабельнее.
#
# Сравните:
#
# ```c++
# std::cout << "ERROR: failed to open " << filename << " at line " << line_number << ". OS response: " << reponse << '\n';
#
# std::cout << boost::format("ERROR: failed to open %1% at line %2%. OS response: %3%\n")
# % filename % line_number % response;
# ```
#
# Но `boost::format` обладает двумя сущестенными недостатками:
# * долгая компиляция
# * медленный
#
# (цифры ниже)
# <br />
# ```c++
# spdlog
# ```
# ##### fmtlib
# https://fmt.dev/latest/
#
# https://github.com/fmtlib/fmt
# Если вы всегда хотели так же элегантно форматировать строки как это сделано в python через метод str.format, то будущее уже здесь.
# ```c++
# // format to string
# std::string s = fmt::format("{} was {} when he became an epic hero", "Ilya", 33);
#
# // format to local buffer
# char[16] out = "";
# fmt::format_to(out, "{}", 42);
#
# // format to local buffer with restrictions
# char[16] out = "";
# fmt::format_to_n(out, 16, "{}", 42);
#
# // format to output iterator
# std::vector<char> out;
# fmt::format_to(std::back_inserter(out), "{}", 42);
#
# // format to std out
# fmt::print("{} was {} when he became an epic hero", "Ilya", 33);
#
# // format to FILE* stream
# FILE *f = ...;
# fmt::print(f, "{} was {} when he became an epic hero", "Ilya", 33);
# ```
# <br />
# Аналогично python-овскому `str.format`:
#
# * форматирование аргументов
#
# ```c++
# fmt::print('pi is approximately equal to {:.3f}', M_PI);
# ```
#
# * нумерация аргументов
#
# ```c++
# // заметьте, что в отличие от |boost::format| аргументы нумеруются с 0
# fmt::print("{0} was at war with {1} at least 12 times. {0} has won approximately 7 times.",
# "Russian Empire",
# "Ottomans Empire"s);
# ```
#
# * именование аргументов
#
# ```c++
# fmt::print("{ru} was at war with {ot} at least 12 times. {ru} has won approximately 7 times.",
# fmt::arg("ru", "Russian Empire"),
# fmt::arg("ot", "Ottomans Empire"s));
# ```
# <br />
# ##### compile-time проверки формата
# Но у нас есть С++, и в отличие от python, мы можем проверить корректность строки форматирования на этапе компиляции:
#
# ```c++
# // compile-time error: 'd' is an invalid specifier for strings.
# std::string s = format(FMT_STRING("{:d}"), "foo");
#
# // compile-time error: argument N2 is not set
# std::string s = format(FMT_STRING("{2}"), 42);
# ```
# <br />
# ##### Форматирование пользовательских типов
# ```c++
# // Пользовательский тип
# struct Point
# {
# float x;
# float y;
# float z;
# };
#
# // Специализация шаблона fmt::formatter для пользовательского типа.
# //
# // Она определяет:
# // * какие опции форматирования доступны
# // * как их обрабатывать
# //
# // В примере рассмотрим опции:
# // {} - формат по умолчанию
# // {:f} - формат с фиксированной точкой
# // {:e} - научный формат
# //
# // Нужно определить 2 метода:
# // |parse| - нужно разобрать строку формата
# // и сохранить результат как внутреннее
# // состояние структуры
# // |format| - имея заполненное внутреннее состояние, выполнить
# // форматирование для конкретного аргумента
# template<>
# struct fmt::formatter<Point>
# {
# enum class Form
# {
# def, // default form for {}
# fix, // fixed format for {:f}
# exp // exponential for {:e}
# };
# Form form = Form::def;
#
# // отметьте здесь constexpr
# constexpr auto parse(format_parse_context& ctx)
# {
# // [ctx.begin(), ctx.end()) - подстрока для парсинга.
# //
# // В таком вызове:
# //
# // fmt::format("{:f} - point of interest", point{1, 2});
# //
# // подстрока равна "f} - point of interest".
# //
# // Обязанность метода - разобрать строку формата до '}' и вернуть,
# // итератор, указывающий на '}', если не получилось - кинуть особое исключение.
# const char error_message_invalid_format[] =
# "invalid format for Point argument, expected {:f} or {:e} or {}";
#
# auto it = ctx.begin();
# const auto end = ctx.end();
#
# // случай "{}"
# if (it != end && *it == '}')
# // сразу возвращаем итератор на "}"
# return it;
#
# // считаем f или e из строки формата
# if (it != end)
# {
# if (*it == 'f')
# form = Form::fix;
# else if (*it == 'e')
# form = Form::exp;
# else
# throw format_error(error_message_invalid_format);
#
# ++it;
# }
#
# // убедимся, что в строке формата больше ничего нет,
# // и мы остановились именно на "}"
# if (it == end || *it != '}')
# throw format_error(error_message_invalid_format);
#
# // возвращаем итератор на "}"
# return it;
# }
#
# template <typename FormatContext>
# auto format(const Point& p, FormatContext& ctx)
# {
# const char* const fmt = form == Form::fix ? "({:.1f}, {:.1f}, {:.1f})" :
# form == Form::exp ? "({:.1e}, {:.1e}, {:.1e})" :
# "({}, {}, {})";
# return format_to(ctx.out(), fmt, p.x, p.y, p.z);
# }
# };
# ```
#
# Использование:
#
# ```c++
# int main()
# {
# std::cout << fmt::format(" {0:f}\n {0:e}\n {0}", Point{3.f, 4.f, 5.f}) << std::endl;
# }
# ```
#
# Вывод:
#
# ```sh
# (3.0, 4.0, 5.0)
# (3.0e+00, 4.0e+00, 5.0e+00)
# (3.0, 4.0, 5.0)
# ```
# <br />
# Трюк для красивого вывода enum-ов:
#
# ```c++
# enum class Color {
# red,
# green,
# blue
# };
#
# template <>
# struct fmt::formatter<Color> : formatter<string_view>
# {
# // метод |parse| отнаследован от форматера строк,
# // а значит наш форматер Color уже умеет понимать
# // все возможные способы отформатировать строку,
# // осталось только эту строку предоставить!
#
# template <typename FormatContext>
# auto format(Color c, FormatContext& ctx)
# {
# string_view name = "unknown";
# switch (c) {
# case color::red: name = "red"; break;
# case color::green: name = "green"; break;
# case color::blue: name = "blue"; break;
# }
# return formatter<string_view>::format(name, ctx);
# }
# };
#
# // usage:
# fmt::print("{:>20}", Color::red);
# ```
# <br />
# ###### несколько приятных утилит из fmtlib
# shortcut для конвертации в строку через формат по умолчанию (замена для `std::format("{}", x)`):
#
# ```c++
# std::string s1 = fmt::to_string(42);
# std::string s2 = fmt::to_string(3.14);
# ```
# `fmt::join`. Обратите внимание, что `fmt::join` принимает range произвольных типов:
#
# ```c++
# std::vector<int> v = {1, 2, 3};
# fmt::print("{}", fmt::join(v, ", ")); // Output: "1, 2, 3"
# ```
# Буфер - аналог small vector
# ```c++
# // fmt::memory_buffer out;
# fmt::memory_buffer<char, 256> out;
# format_to(out, "The answer is {}.", 42);
# ```
# Форматирование дат
#
# ```c++
# std::time_t t = std::time(nullptr);
# fmt::print("The date is {:%Y-%m-%d}.", *std::localtime(&t)); // Prints "The date is 2016-04-29."
# ```
# <br />
# ##### std::format (since C++20)
# https://en.cppreference.com/w/cpp/utility/format
#
# https://en.cppreference.com/w/cpp/utility/format/formatter
# `fmtlib` оказался настолько хорош, что его доработали и перенесли в стандартную библиотеку С++, где назвали `std::format`. С С++20 можно пользоваться продвинутыми способами форматирования без сторонних библиотек.
# <br />
# ##### сравнение библиотек
# Скорость исполнения, [подробности эксперимента](https://github.com/fmtlib/fmt#speed-tests)
#
# | Library | Method | Run Time, s
# |:------------------|:--------------|:-----------:
# | libc | printf | 1.04
# | libc++ | std::ostream | 3.05
# | {fmt} 6.1.1 | fmt::print | 0.75
# | Boost Format 1.67 | boost::format | 7.24
# | Folly Format | folly::format | 2.23
#
# Время компиляции, [подробности эксперимента](https://github.com/fmtlib/fmt#compile-time-and-code-bloat)
#
# | Method | Compile Time, s |
# |:--------------|:---------------:|
# | printf | 2.6 |
# | printf+string | 16.4 |
# | iostreams | 31.1 |
# | {fmt} | 19.0 |
# | Boost Format | 91.9 |
# | Folly Format | 115.7 |
#
# Размер исполняемого файла, [подробности эксперимента](https://github.com/fmtlib/fmt#compile-time-and-code-bloat)
#
# | Method | Executable size, KiB | Stripped size, KiB |
# |:--------------|:--------------------:|:------------------:|
# | printf | 29 | 26 |
# | printf+string | 29 | 26 |
# | iostreams | 59 | 55 |
# | {fmt} | 37 | 34 |
# | Boost Format | 226 | 203 |
# | Folly Format | 101 | 88 |
# <br />
# ##### Резюме
#
# * В обычном С++ приложении для форматирования строк используйте `std::format` с 20-го стандарта либо `fmtlib` до С++20.
#
| 2020/sem2/lecture_9_format/lecture.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Stationary Iterative Methods for Linear Systems
# +
import numpy as np
import scipy.linalg as la
import matplotlib.pyplot as pt
# -
# Let's solve $u''=-30x^2$ with $u(0)=1$ and $u(1)=-1$.
# +
n = 50
mesh = np.linspace(0, 1, n)
h = mesh[1] - mesh[0]
# -
# Set up the system matrix `A` to carry out centered finite differences
#
# $$
# u''(x)\approx \frac{u(x+h) - 2u(x) + u(x-h)}{h^2}.
# $$
#
# Use `np.eye(n, k=...)`. What needs to be in the first and last row?
A = (np.eye(n, k=1) + -2*np.eye(n) + np.eye(n, k=-1))/h**2
A[0] = 0
A[-1] = 0
A[0,0] = 1
A[-1,-1] = 1
# Next, fix the right hand side:
b = -30*mesh**2
b[0] = 1
b[-1] = -1
# Compute a reference solution `x_true` to the linear system:
x_true = la.solve(A, b)
pt.plot(mesh, x_true)
# Next, we'll try all the stationary iterative methods we have seen.
# ## Jacobi
x = np.zeros(n)
# Next, apply a Jacobi step:
# +
x_new = np.empty(n)
for i in range(n):
x_new[i] = b[i]
for j in range(n):
if i != j:
x_new[i] -= A[i,j]*x[j]
x_new[i] = x_new[i] / A[i,i]
x = x_new
# -
pt.plot(mesh, x)
pt.plot(mesh, x_true, label="true")
pt.legend()
# * Ideas to accelerate this?
# * Multigrid
# ## Gauss-Seidel
x = np.zeros(n)
# +
x_new = np.empty(n)
for i in range(n):
x_new[i] = b[i]
for j in range(i):
x_new[i] -= A[i,j]*x_new[j]
for j in range(i+1, n):
x_new[i] -= A[i,j]*x[j]
x_new[i] = x_new[i] / A[i,i]
x = x_new
pt.plot(mesh, x)
pt.plot(mesh, x_true, label="true")
pt.legend()
# -
# ### And now Successive Over-Relaxation ("SOR")
x = np.zeros(n)
# +
x_new = np.empty(n)
for i in range(n):
x_new[i] = b[i]
for j in range(i):
x_new[i] -= A[i,j]*x_new[j]
for j in range(i+1, n):
x_new[i] -= A[i,j]*x[j]
x_new[i] = x_new[i] / A[i,i]
direction = x_new - x
omega = 1.5
x = x + omega*direction
pt.plot(mesh, x)
pt.plot(mesh, x_true, label="true")
pt.legend()
pt.ylim([-1.3, 1.3])
# -
| pdes/Stationary Iterative Methods.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Bookworm
#
#
# ## Overview
#
# In this project, you will build a simple question-answering agent that is able to learn from any text data you provide, and answer queries posed in natural language. You will use IBM Watson's cloud-based services to process the input text data and find relevant responses.
#
# ## Learning Objectives
#
# By completing this project, you will learn how to:
#
# - Create a cloud-based NLP service instance and configure it.
# - Ingest a set of text documents using the service and analyze the results.
# - Accept questions in natural language and parse them.
# - Find relevant answers from the preprocessed text data.
#
# ## Getting Started
#
# In order to use Watson's cloud-based services, you first need to create an account on the [IBM Bluemix platform](https://console.ng.bluemix.net/).
#
# <div>
# <div style="display: table-cell; width: 50%;">
# <img src="images/watson-logo.png" alt="IBM Watson logo" width="200" />
# </div>
# <div style="display: table-cell; width: 50%;">
# <img src="images/bluemix-logo.png" alt="IBM Bluemix logo" width="400" />
# </div>
# </div>
#
# Then, for each service you want to use, you have to create an instance of that service. You can continue with the tasks below, and create a service instance when indicated.
# ## 1. Create and configure Discovery service
#
# Create an instance of the **Discovery** service. You will use this to process a set of text documents, and _discover_ relevant facts and relationships.
#
# - Go to the [IBM Bluemix Catalog](https://console.ng.bluemix.net/catalog/?taxonomyNavigation=services&category=watson).
# - Select the service you want, **Discovery**, under the **Watson** category.
# - Enter a Service Name for that instance, e.g. **Disco1** and a Credential Name, e.g. **Disco1-Creds** (these are just for you to be able to refer to later, they do not affect the functioning of the service).
# - You should be able to see your newly-created service in your [Services Dashboard](https://console.ng.bluemix.net/dashboard/services).
# - Open the service instance, click on the **Service credentials** tab, and then **View credentials** under Actions. This is where you will find the username and password to use when connecting to the service.
#
# <img src="images/discovery-creds.png" alt="Discovery Service - Credentials tab" width="800" />
#
# Save the credentials for the discovery service in a JSON file in the current directory named `service-credentials.json` with the following format:
#
# ```json
# {
# "discovery": {
# "username": "<your Discovery username here>",
# "password": "<<PASSWORD>>"
# },
# "conversation": {
# "username": "",
# "password": ""
# }
# }
#
# ```
#
# You will be filling out the Conversation service credentials later, when you create an instance for it. Note that you should keep these credentials secret. Please do not turn them in with your submission!
#
# ### Connect to the service instance
#
# Let's connect to the service instance you just created using IBM Watson's [Python SDK](https://github.com/watson-developer-cloud/python-sdk). You will first need to install the SDK:
# ```bash
# pip install watson-developer-cloud
# ```
#
# Now execute each code cell below using **`Shift+Enter`**, and complete any steps indicated by a **`TODO`** comment. For more information on the Discovery service, please read the [Documentation](https://www.ibm.com/watson/developercloud/doc/discovery/index.html) and look at the [API Reference](https://www.ibm.com/watson/developercloud/discovery/api/v1/?python) as needed.
# +
# Usual Python imports
import sys
import os
import glob
import json
# BeautifulSoup, for parsing HTML
from bs4 import BeautifulSoup
# Matplotlib, for plotting
import matplotlib.pyplot as plt
# %matplotlib inline
# Watson Python SDK
import watson_developer_cloud
# Utility functions
import helper
# -
# Connect to the Discovery service instance
# TODO: Ensure that your username and password from the Service Credentials tab are in service-credentials.json
# Note that these credentials are different from your IBM Bluemix login, and are specific to the service instance
discovery_creds = helper.fetch_credentials("discovery")
discovery = watson_developer_cloud.DiscoveryV1(
version='2016-11-07',
username=discovery_creds['username'],
password=<PASSWORD>_creds['password'])
# ### Create an environment
#
# The Discovery service organizes everything needed for a particular application in an _environment_. Let's create one called "Bookworm" for this project.
#
# > _**Note**: It is okay to run this block multiple times - it will not create duplicate environments with the same name._
# Prepare an environment to work in
env, env_id = helper.fetch_object(
discovery, "environment", "Bookworm",
create=True, create_args=dict(
description="A space to read and understand stories", # feel free to edit
size=0 # use 0 for free plan (see API reference for more on sizing)
))
print(json.dumps(env, indent=2))
# ### Verify configuration options
#
# A _configuration_ defines what natural language processing routines are applied to any documents that are submitted to the service. Each environment gets a default configuration when it is created.
#
# You can fetch the default configuration and view the different options using the following piece of code.
#
# View default configuration
cfg_id = discovery.get_default_configuration_id(environment_id=env_id)
cfg = discovery.get_configuration(environment_id=env_id, configuration_id=cfg_id)
print(json.dumps(cfg, indent=2))
# There are 3 main configuration blocks that affect how input documents are processed:
#
# 1. **`conversions`**: How to convert documents in various formats (Word, PDF, HTML) and extract elements that indicate some structure (e.g. headings).
# 2. **`enrichments`**: What NLP output results are we interested in (keywords, entities, sentiment, etc.).
# 3. **`normalizations`**: Post-processing steps to be applied to the output. This can be left empty in most cases, unless you need the output to be normalized into a very specific format.
#
# _**Note**: The default configuration for an environment cannot be modified. If you need to change any of the options, you will need to create a new one, and then edit it. The easiest way to do this is using the service dashboard, which is described later._
# ### Test your configuration
#
# It is a good idea to test your configuration on a small sample text before you apply it to a larger document collection.
#
# _**Note**: We have supplied a sample document (`data/sample.html`) containing the opening crawl text for Star Wars: Episode IV, but you are free to use a text of your choosing._
#
# **Q**: (optional) If you use your own sample text, provide a brief title and description below.
#
# **A**:
# Test configuration on some sample text
data_dir = "data"
filename = os.path.join(data_dir, "sample.html")
with open(filename, "r") as f:
res = discovery.test_document(environment_id=env_id, configuration_id=cfg_id, fileinfo=f)
print(json.dumps(res, indent=2))
# ### Analyze test output
#
# The results returned by the service contain a _snapshot_ of the information extracted at each step of processing - document conversions, enrichments and normalizations. We are interested in the output of applying enrichments (`"enrichments_output"`) or after normalizing them (`"normalizations_output"`). These should be identical if no post-processing/normalizations were specified in the configuration.
# Take a closer look at the results from the "enrichments_output" or "normalizations_output" step
output = next((s["snapshot"] for s in res["snapshots"] if s["step"] == "enrichments_output"), None)
print(json.dumps(output, indent=2))
# Answer the following questions based on the output above. Note that it contains the input HTML, extracted text and metadata as well as the actual enrichment results (`"enriched_text"` block).
#
# #### Sentiment
#
# **Q**: What is the overall sentiment detected in this text? Mention the `type` (positive/negative) and `score`.<br />
# (_Hint: Look for the `"docSentiment"` key in the output._)
#
# **A**:
#
#
# #### Concepts
#
# **Q**: List 3 concepts that have been identified with a relevance > 0.5. Note that not all concepts here may be present directly in the text, some may have been inferred by Watson.<br />
# (_Hint: Look for `"concepts"`._)
#
# **A**:
#
#
# #### Relations
#
# Each relation is essentially a deeper analysis of a sentence (or part of a sentence). Here is a sample relation:
# ```json
# {
# "sentence": " During the battle, Rebel spies managed to steal secret plans to the Empire's ultimate weapon, the DEATH STAR, an armored space station with enough power to destroy an entire planet.",
# "subject": {
# "text": "Rebel spies",
# "keywords": [
# {
# "text": "Rebel spies"
# }
# ]
# },
# "action": {
# "text": "managed to steal",
# "lemmatized": "manage to steal",
# "verb": {
# "text": "steal",
# "tense": "future"
# }
# },
# "object": {
# "text": "secret plans",
# "keywords": [
# {
# "text": "secret plans"
# }
# ]
# }
# }
# ```
#
# In this case, Watson seems to have done a pretty good job of extracting some meaning from the sentence.
#
# **Q**: Find a relation where the extracted meaning is not as accurate, or not what you would've expected. List the `sentence`, `subject`, `action` and `object` parts as identified, and what you would've marked instead.<br />
# (_Hint: Look for `"relations"`._)
#
# **A**:
#
# #### Keywords
#
# You may have noticed that Watson identifies some `"keywords"` in the relations, e.g. `"Rebel spies"` and `"secret plans"` in the Star Wars example above. The output also contains a list of all keywords at the top level, for your convenience, along with their relevance to the document and sentiment conveyed. Let's visualize these keywords as a word cloud!
#
# Note: We'll be using this handy [worldcoud library](https://github.com/amueller/word_cloud) to generate the visualization. So you will need to install it first:
# ```bash
# pip install wordcloud
# ```
# +
# Visualize keywords by relevance as a wordcloud
from wordcloud import WordCloud
wc_data = { w["text"]: w["relevance"] for w in output["enriched_text"]["keywords"] }
wc = WordCloud(width=400, height=300, scale=2, background_color="white", colormap="Vega10")
wc.generate_from_frequencies(wc_data) # use precomputed relevance instead of frequencies
plt.figure(figsize=(4, 3), dpi=200)
plt.imshow(wc, interpolation='bilinear')
plt.axis("off")
# -
# Feel free to play with this visualization and improve it. What about using a different metric instead of relevance, e.g. direct word frequencies that the wordcloud library computes by default?
#
# #### Other results
#
# Watson's output also includes processed results from other enrichments that were applied to the text, including entities and taxonomy (what topic or category does this text relate to).
#
# ```json
# "taxonomy": [
# {
# "label": "/art and entertainment/movies and tv/movies",
# "score": 0.584247,
# "confident": false
# },
# {
# "label": "/society/unrest and war",
# "score": 0.517031,
# "confident": false
# },
# {
# "confident": false,
# "label": "/law, govt and politics/armed forces/army",
# "score": 0.215561
# }
# ],
# ```
#
# Get a good sense of all the different pieces of information available in the results. Start thinking about which ones will be useful for looking up answers to questions, and how you might use them.
# ## 2. Ingest documents
#
# ### Create a collection
#
# A _collection_ is used to organize documents of the same kind. For instance, you may want to create a collection of book reviews, or a collection of Wikipedia articles, but it may not make much sense to mix the two groups. This allows Watson to make meaningful inferences over the set of documents, find commonalities and identify important concepts.
#
# Let's create one called "Story Chunks".
# Prepare a collection of documents to use
col, col_id = helper.fetch_object(discovery, "collection", "Story Chunks", environment_id=env_id,
create=True, create_args=dict(
environment_id=env_id, configuration_id=cfg_id,
description="Stories and plots split up into chunks suitable for answering"
))
print(json.dumps(col, indent=2))
# Once you have created a collection, you should be able to view it using the Discovery Service tool. To open, go to the **Manage** tab for your service instance, and click the **Launch tool** button.
#
# <img src="images/discovery-manage.png" alt="Discovery service - Manage tab" width="800" />
#
# Here you should see the "Story Chunks" collection you just created.
#
# <img src="images/discovery-tooling.png" alt="Discovery service - Tool showing collections" width="800" />
#
# You can open the collection to view more details about it. If you need to modify configuration options, click the **Switch** link and create a new configuration (the default one cannot be changed).
# ### Add documents
#
# Okay, now that we have everything set up, let's add a set of "documents" we want Watson to look up answers from, using the Python SDK. Note that Watson treats each "document" as a unit of text that is returned as the result of a query. But we want to retrieve a paragraph of text for each question. So, let's split each file up into individual paragraphs. We will use the [BeautifulSoup](https://www.crummy.com/software/BeautifulSoup/bs4/doc/) library for this purpose.
#
# _**Note**: You could also add and manage documents in the collection using the Discovery tool, but you would have to split paragraphs up into separate files._
#
# _**Note**: We have provided a set of files (`data/Star-Wars/*.html`) with summary plots for Star Wars movies, but you are free to use a collection of your choice. Open one of the files in a text editor to see how the paragraphs are delimited using `<p>...</p>` tags - this is how the code block below split paragraphs into separate "documents"._
# Add documents to collection
doc_ids = [] # to store the generated id for each document added
for filename in glob.glob(os.path.join(data_dir, "Star-Wars", "*.html")):
print("Adding file:", filename)
with open(filename, "r") as f:
# Split each individual <p> into its own "document"
doc = f.read()
soup = BeautifulSoup(doc, 'html.parser')
for i, p in enumerate(soup.find_all('p')):
doc_info = discovery.add_document(environment_id=env_id, collection_id=col_id,
file_data=json.dumps({"text": p.get_text(strip=True)}),
mime_type="application/json",
metadata={"title": soup.title.get_text(strip=True)})
doc_ids.append(doc_info["document_id"])
print("Total", len(doc_ids), "documents added.")
# If you look at the collection details, you may notice that the `"document_counts"` field now shows some documents as `available` or `processing`. Once processing is complete, you should see all the documents under the `available` count.
# View collection details to verify all documents have been processed
col, col_id = helper.fetch_object(discovery, "collection", "Story Chunks", environment_id=env_id)
print(json.dumps(col, indent=2))
# So, what did the Discovery service learn? If you list the fields extracted from the set of documents in the collection as part of the enrichment process, you'll see familiar fields like `concepts`, `entities` and `keywords` that were returned in the test analysis.
# List all fields extracted
discovery.list_collection_fields(environment_id=env_id, collection_id=col_id)
# ### Test query
#
# Let's perform a simple query to see if the service can fetch the proper document for us:
# > _Look for all paragraphs that have a relation (sentence) with "Jar Jar" as the subject, and return the title and text._
#
# A simple query
results = discovery.query(environment_id=env_id, collection_id=col_id,
query_options={
"query": "enriched_text.relations.subject.text:\"Jar Jar\"",
"return": "metadata.title,text"
})
print(json.dumps(results, indent=2))
# Change the above query and see what results you get! Try to find one that returns relevant results, and keep that (along with the output) for review.
#
# > See [Query building reference](https://www.ibm.com/watson/developercloud/doc/discovery/query-reference.html) for descriptions of all possible parameters, operators and aggregations. You can also choose to build the query using the web interface (click the "Story Chunks" collection to query it), and then reproduce the query here.
#
# Then answer the questions below:
#
# **Q**: What query did you try? Express it in plain words below.
#
# **A**:
#
#
# **Q**: What answer did you get back from Watson? You only need to mention the relevant snippet of text fro mthe paragraph(s) returned.
#
# **A**:
#
# ## 3. Parse natural language questions
#
# In order to understand questions posed in natural language, we'll use another Watson service called [Conversation](https://www.ibm.com/watson/developercloud/doc/conversation/index.html). It can be used to design conversational agents or _chatbots_ that exhibit complex behavior, but for the purpose of this project, we'll only use it to parse certain kinds of queries.
#
# ### Create a Conversation service instance
#
# Just like you did for the Discovery service, create an instance of the Conversation service. Then launch the associated tool from the service dashboard.
#
# <img src="images/conversation-tooling.png" alt="Conversation service - Tool homepage" width="800" />
#
# A _workspace_ allows you to keep all the items you need for a particular application in one place, just like an _environment_ in case of the Discovery service. Create one called "Bookworm" with a suitable description, such as "I know a lot of stories. Ask me a question!"
#
# <img src="images/conversation-workspace.png" alt="Conversation service - Blank workspace" width="800" />
#
# This should open up a blank workspace, where you can add intents, define the entities you want the agent to idenitfy and structure the overall dialog.
#
# ### Add intents
#
# An _intent_ is the goal or purpose of a user's input. Create a set of intents (at least 3) that capture the different kinds of questions that you want the system to answer, e.g. _who_, _what_ and _where_. Along with each intent, add a list of user examples or _utterances_ that map to that intent.
#
# For instance, you could enter the following examples for the _where_ intent:
#
# - Where is the Jedi temple located?
# - Where was Luke born?
#
# The Conversation service recommends at least 5 examples for each intent so that Watson learns how to recognize it. These don't have to be very precise, but more examples the better.
#
# <img src="images/conversation-intents.png" alt="Conversation service - Intents listed" width="800" />
#
# Feel free to create your own intents, based on the kinds of questions you want the system to answer, e.g. "How many ...", "What are the most common ..." etc. Each intent will need to be mapped to an appropriate query.
#
# > See [**Defining intents**](https://www.ibm.com/watson/developercloud/doc/conversation/intents.html) for a helpful video and further instructions.
#
# **Q**: What intents did you add to the Conversation service instance?
#
# **A**:
#
#
#
# **Q**: Pick one of these intents, and list at least 5 examples for the intent that you entered.
#
# **A**:
#
#
#
# ### Add entities
#
# Once you have your intents set, let's tell the service what entities we want it to identify. One way to do this is using the tool interface, and entering them one-by-one.
#
# > Go to [**Defining entities**](https://www.ibm.com/watson/developercloud/doc/conversation/entities.html) to see how that is done.
#
# But that can be tedious! So let's refer back to the entities that the Discovery service identified, and load them in programmatically.
#
# As before, let's connect to the Conversation service first. Remember to enter your service credentials below.
# Connect to the Conversation service instance
# TODO: Enter your username and password from the Service Credentials tab in service-credentials.json
conversation_creds = helper.fetch_credentials('conversation')
conversation = watson_developer_cloud.ConversationV1(
version='2017-02-03',
username=conversation_creds['username'],
password=conversation_creds['password'])
# Fetch the workspace you just created called "Bookworm".
wrk, wrk_id = helper.fetch_object(conversation, "workspace", "Bookworm")
print(json.dumps(wrk, indent=2))
# Collect all the entities from the Discovery service collection.
# +
# Get all the entities from the collection and group them by type
response = discovery.query(environment_id=env_id, collection_id=col_id,
query_options={
"return": "enriched_text.entities.type,enriched_text.entities.text"
})
# Group individual entities by type ("Person", "Location", etc.)
entities_by_type = {}
for document in response["results"]:
for entity in document["enriched_text"]["entities"]:
if entity["type"] not in entities_by_type:
entities_by_type[entity["type"]] = set()
entities_by_type[entity["type"]].add(entity["text"])
# Ignore case to avoid duplicates
for entity_type in entities_by_type:
entities_by_type[entity_type] = {
e.lower(): e for e in entities_by_type[entity_type]
}.values()
# Restructure for loading into Conversation workspace
entities_grouped = [{
"entity": entity_type,
"values": [{"value": entity} for entity in entities]}
for entity_type, entities in entities_by_type.items()]
entities_grouped
# -
# Update the workspace with these entities and verify that have been added correctly.
# +
# Add these grouped entities to the Conversation workspace
conversation.update_workspace(workspace_id=wrk_id, entities=entities_grouped)
workspace_details = conversation.get_workspace(workspace_id=wrk_id, export=True)
print(json.dumps(workspace_details["entities"], indent=2))
# -
# _**Note**: Ensure that at least 3 entity types, with at least 1 example entity each have been added._
#
# Here is what the list of entities should look like through the Conversation tool.
#
# <img src="images/conversation-entities.png" alt="Conversation service - Entities listed" width="800" />
#
# **Q**: Name 3 entity types that were added, with at least 1 example entity each (e.g. entity type: _City_, example: _Los Angeles_).
#
# **A**:
# ### Design dialog flow
#
# As a final step in creating the Conversation interface, let's design a typical dialog with a user. The most intuitive way to do this is to use the Dialog tab in the tool. Here, you can add _nodes_ that capture different stages in the dialog flow, and connect them in a meaningful way.
#
# Go ahead and add at least 3 dialog nodes. Specify the triggers in terms of the intents and entities that you'd like to match, and an optional intermediate response like "Let me find that out for you." The actual response will be fetched by querying the Discovery service.
#
# Here is what the dialog nodes should look like.
#
# <img src="images/conversation-dialog_nodes.png" alt="Conversation service - Dialog nodes" width="640" />
#
# **Q**: Specify 3 dialog nodes you added, along with the trigger (intent and/or entities) for each.
#
# **A**:
# ### Test dialog
#
# Let's run through a test dialog to demonstrate how the system transitions to one of the nodes you defined above.
# +
# Testing the dialog flow
# Start conversation with a blank message
results = conversation.message(workspace_id=wrk_id, message_input={})
context = results["context"]
# Then ask a sample question
question= "Who is Luke's father?"
results = conversation.message(workspace_id=wrk_id, message_input={
"text": question,
"context": context
})
print(json.dumps(results, indent=2))
# -
# ## 4. Query document collection to fetch answers
#
# The Discovery service includes a simple mechanism to make queries against your enriched collection of documents. But you have a lot of control over what fields are searched, how results are aggregated and values are returned.
#
# ### Process sample question
#
# Choose a sample nautal language question to ask, and run it through the Conversation service, just like you did above when testing dialog flow.
# +
# TODO: Run a sample question through Conversation service
question= "Who is Luke's father?"
results = conversation.message(workspace_id=wrk_id, message_input={
"text": question,
"context": context
})
print(json.dumps(results, indent=2))
# Run a sample question through Conversation service
question= "Who is Luke's father?"
results = conversation.message(workspace_id=wrk_id, message_input={
"text": question,
"context": context
})
print(json.dumps(results, indent=2))
# -
# Now extract the intent and entities identified in the question, and optionally what dialog node was triggered (in case you need it later to customize your response). Some sample code is provided below, but you may need to modify it.
# +
# TODO: Identify the intent(s) the user expressed (typically a single one)
query_intents = [intent["intent"] for intent in results["intents"]]
print("Intent(s):", query_intents)
# TODO: Extract the entities found in the question text
query_entities = [entity["value"] for entity in results["entities"]]
print("Entities:", query_entities)
# TODO: (optional) Find out what dialog node was triggered
query_nodes = results["input"]["context"]["system"]["dialog_stack"][0]["dialog_node"]
print("Queries:", query_nodes)
# -
# ### Query the collection
#
# Design a query based on the information extracted above, and run it against the document collection. The sample query provided below simple looks for all the entities in the raw `text` field. Modify it to suit your needs.
#
# Take a look at the [API Reference](https://www.ibm.com/watson/developercloud/discovery/api/v1/?python#query-collection) to learn more about the query options available, and for more guidance see this [documentation page](https://www.ibm.com/watson/developercloud/doc/discovery/using.html).
#
# _**Note**: You may want to design different queries based on the intent / dialog node that was triggered._
# TODO: Query the Discovery service based on the intent and entities
query_results = discovery.query(environment_id=env_id, collection_id=col_id,
query_options={
"query": "text:{}".format(",".join("\"{}\"".format(e) for e in query_entities)),
"return": "text"
})
print(json.dumps(query_results, indent=2))
# ### Process returned results
#
# If you properly structure the query, Watson is able to do a pretty good job of finding the relevant information. But the result returned is a JSON object. Now your task is to convert that result into an appropriate response that best addresses the original natural language question that was asked.
#
# E.g. if the question was "Who saved Han Solo from Jabba the Hutt?" the answer should ideally just be "The Rebels" and not the entire paragraph describing Han Solo's rescue. But that can be a backup response if you cannot be more specific.
#
# _**Note**: You may have to go back to the previous step and modify the query, especially what you want the Discovery service to return, and this may depend on the intent / dialog node triggered. E.g. study the different parts of a "relation" structure to see how you might construct queries to match them._
# +
# TODO: Process returned results and compose an appropriate response
from nltk.collocations import *
import nltk
# Process returned results and compose an appropriate response\
obj = "father"
raw = ''
for result in query_results['results']:
answers = [potent_ans for potent_ans in result['text'].split('.') if obj in potent_ans]
if answers:
answer = [potent_ans for potent_ans in result['text'].split('.') if obj in potent_ans][0]
print(answer+'\n')
raw += (' '+answer)
# -
# ## 5. Reflections
#
# **Q**: Now that you have gone through this exercise of designing a system that uses two IBM Watson services, what did you learn? What were some of the strengths and weaknesses of this approach?
#
# **A**:
#
#
# ## (Optional) Extensions
#
# We have provided a set of sample data files containing Star Wars plot summaries. But as mentioned before, you are free to use your own dataset. In fact, a larger dataset maybe more suitable for use with IBM Watson's NLP services. If you used your own dataset, answer the following questions.
#
# **Q**: What dataset did you use, and in what ways is it different from the sample files provided?
#
# **A**:
#
#
# **Q**: Either include your dataset in the .zip file or repository you submit, or provide clear instructions on how to obtain the dataset, so that your reviewer can run your notebook or inspect the data to verify your results.
#
# **A**:
#
#
# _You can also design a web-based application that utilizes these services and deploy that on Bluemix! If you do, please share with your instructors and peers._
| bookworm/bookworm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Measured 190 GHz Active 2-Port
# The Vector Fitting feature is demonstrated using a 2-port S-matrix of an active circuit measured from 140 GHz to 220 GHz. Additional explanations and background information can be found in the [Vector Fitting tutorial](../../tutorials/VectorFitting.ipynb).
import skrf
import numpy as np
# This example is a lot more tricky to fit, because the responses contain a few "bumps" and noise from the measurement. In such a case, finding a good number of initial poles can take a few iterations.
# Load the Network from a Touchstone file and create the Vector Fitting instance:
nw2 = skrf.network.Network('./190ghz_tx_measured.S2P')
vf2 = skrf.VectorFitting(nw2)
# **First attempt:** Perform the fit using 4 real poles and 3 complex-conjugate poles with *linear* spacing (default):
vf2.vector_fit(n_poles_real=4, n_poles_cmplx=3)
# The function `plot_convergence()` can be helpful to examine the convergence and see if something was going wrong. In this case, it took quite a while (84 iteration steps), but the results converged nevertheless.
vf2.plot_convergence()
# Checking the results by comparing the model responses to the original sampled data indicates a successful fit:
vf2.plot_s_mag(0, 0) # s11
vf2.plot_s_mag(1, 0) # s21
# It is a good idea to also check the model response well outside the original frequency range. This reveals a strong resonance at higher frequencies (at approx. 330 GHz), which is not ideal:
freqs2 = np.linspace(0, 500e9, 501)
vf2.plot_s_mag(0, 0, freqs2)
vf2.plot_s_mag(1, 0, freqs2)
# **Second attempt:** Maybe an even better fit can be achieved, so let's try again. It sometimes helps to change the initial pole spacing from *linear* to *logarithmic*:
vf2.vector_fit(n_poles_real=4, n_poles_cmplx=3, init_pole_spacing='log')
vf2.plot_convergence()
# This fit converged slightly quicker (66 iteration steps) and also matches the network data very well inside the fitting band. Interestingly, the strong resonance from before in the outside band is replaced with a much weaker one at 263 GHz:
vf2.plot_s_mag(0, 0)
vf2.plot_s_mag(1, 0)
vf2.plot_s_mag(0, 0, freqs2)
vf2.plot_s_mag(1, 0, freqs2)
# This looks good, so let's export the model as a SPICE subcircuit. For example:
#
# `vf2.write_spice_subcircuit_s('/home/vinc/Desktop/190ghz_tx.sp')`
#
# The subcircuit can then be simulated in SPICE with the same AC simulation setup as in the [ring slot example](./vectorfitting_ringslot.ipynb):
# <img src="./ngspice_190ghz_tx_sp_mag.svg" />
# <img src="./ngspice_190ghz_tx_sp_smith.svg" />
| doc/source/examples/vectorfitting/vectorfitting_190ghz_active.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Approximation
# $\hat{y} = wx + b$, this is the linear function we want to approximate a set of data $\{ x_i, y_i\}, i \in [1, n]$
#
# ## Cost Function
# MSE = $J(w, b) = \frac{1}{N} \sum_{i=1}^n (y_i - (wx_i + b)) ^ 2$
#
# We want to find the right weight $w$ and bias $b$ to minimun the cost fuction, one way to do so is **Gradient Descent**.
#
# $$\frac{\partial J(w, b)}{\partial w} = - \frac{2}{N} \sum_{i=1}^n (y_i - (w x_i + b)) x_i$$
# $$\frac{\partial J(w, b)}{\partial b} = - \frac{2}{N} \sum_{i=1}^n (y_i - (w x_i + b))$$
#
# ## Gradient Descent
# ### Update rules
#
# $w = w - \alpha \dot \nabla_w J(w, b)$
#
# $b = b - \alpha \dot \nabla_b J(w, b)$, where $\alpha$ is learning rate, and
#
# $\nabla_w J(w, b) = - \frac{2}{N} \sum_{i=1}^n (y_i - (w x_i + b)) x_i = \frac{2}{N} \sum_{i=1}^n (\hat{y}_i - y_i) x_i$
#
# $\nabla_b J(w, b) = - \frac{2}{N} \sum_{i=1}^n (y_i - (w x_i + b)) = \frac{2}{N} \sum_{i=1}^n (\hat{y}_i - y_i)$
# +
import numpy as np
class LinearRegression:
def __init__(self, lr=0.001, n_iters=1000):
self.lr = lr
self.n_iters = n_iters
self.weights = None
self.bias = None
def fit(self, X, y):
n_samples, n_features = X.shape
self.weights = np.zeros(n_features)
self.bias = 0
for _ in range(self.n_iters):
y_pred = np.dot(X, self.weights) + self.bias
partial_w = (2 / n_samples) * np.dot(X.T, (y_pred - y))
partial_b = (2 / n_samples) * np.sum(y_pred - y)
self.weights -= self.lr * partial_w
self.bias -= self.lr * partial_b
def predict(self, X):
y_pred = np.dot(X, self.weights) + self.bias
return y_pred
# +
# Do experiments
from sklearn.model_selection import train_test_split
from sklearn import datasets
import matplotlib.pyplot as plt
from sklearn.metrics import mean_squared_error as mse
X, y = datasets.make_regression(n_samples=100, n_features=1, noise=20, random_state=0)
X_train, X_val, y_train, y_val = train_test_split(X, y, test_size=0.2, random_state=1233)
fig = plt.figure(figsize=(8, 6))
# plt.scatter(X[:, 0], y, color='b', marker='o', s=30)
lr = LinearRegression(lr=0.05)
lr.fit(X_train, y_train)
y_pred = lr.predict(X_val)
print("MSE, ", mse(y_val, y_pred))
y_pred_line = lr.predict(X)
cmap = plt.get_cmap('viridis')
m1 = plt.scatter(X_train, y_train, color=cmap(0.9), s=10)
m2 = plt.scatter(X_val, y_val, color=cmap(0.6), s=10)
plt.plot(X, y_pred_line, color='black', linewidth=2)
| codes/mlmodels/.ipynb_checkpoints/linear_regression-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/ishitagithub59/firstrepo/blob/main/creditcard.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="oD3FJE9wmym2"
#importing all libraries
import numpy as np
import pandas as pd
import sklearn
import scipy
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.metrics import classification_report,accuracy_score
from sklearn.ensemble import IsolationForest
from sklearn.neighbors import LocalOutlierFactor
from sklearn.svm import OneClassSVM
from pylab import rcParams
rcParams['figure.figsize'] = 14, 8
RANDOM_SEED = 42
LABELS = ["Normal", "Fraud"]
# + colab={"base_uri": "https://localhost:8080/", "height": 270} id="PDeeNbRInVrf" outputId="64483532-8ff6-413b-cc5c-98d5d45f64bd"
data = pd.read_csv('/content/drive/MyDrive/creditcard.csv',sep=',') #read the data
data.head() #read the data upto 1st 5 column
# + colab={"base_uri": "https://localhost:8080/"} id="un0ecmZVT10b" outputId="a53f420d-59a2-48b5-c812-a11154304a9b"
data.info() # read information of the data
# + colab={"base_uri": "https://localhost:8080/"} id="PQF8yA_rnvjp" outputId="89368d15-2f59-4c05-9d3e-cd3a9d7b8b20"
data.isnull().values.any() # True means there are null values. If it is False means there is no null values
# + colab={"base_uri": "https://localhost:8080/", "height": 313} id="Sz1HS0H4nx1V" outputId="6a4fd7a5-d21b-4900-b6b6-58b0d19fdd2b"
# Finding how many classes are there and there corresponding frequency
count_classes = pd.value_counts(data['Class'], sort = True)
count_classes.plot(kind = 'bar', rot=0)
plt.title("Transaction Class Distribution")
plt.xticks(range(2), LABELS)
plt.xlabel("Class")
plt.ylabel("Frequency")
# Instead of written whole code we can also write ---- data['Class'].value_counts() -> show 0 for total transaction ; 1 for fraud
# + [markdown] id="kd5gQbQBNG-q"
# **From this graph shows that normal transaction is more than 250k where fraudulant transaction is very very less**
# + [markdown] id="TBwjGCloRhUu"
# ***
# **Balanced Dataset** — If in our dataset we have class values which are approximately same as with each other, then we can say our dataset in balance
#
# **Imbalanced Dataset** — If there is the very high different between the class values, then we can say our dataset in Imbalance Dataset.
# ***
# + [markdown] id="N_RZ2uihOcC4"
# So directly we can see it's a imbalance dataset. For that reason we are going to implement **Isolation Forest Algorithm, Local Outlier Factor and Support Vector Machine**
# + id="LA0tAb8mU3Xn"
fraud = data[data['Class']==1] # we are taking 1 for fraud and 0 for normal transaction
normal = data[data['Class']==0]
# + colab={"base_uri": "https://localhost:8080/"} id="Xs9cPdblU78P" outputId="10bc1a34-68dc-49ce-c233-2c00379512d1"
print(fraud.shape,normal.shape) # It's a highly imbalanced dataset
# + [markdown] id="KWydkuhvPMOZ"
# The value shows us that **492** transaction are fraud and **284315** transactions are normal---- very difference, for that reason its called imbalanced dataset.
#
# + colab={"base_uri": "https://localhost:8080/"} id="dza_tpeXVHJr" outputId="b825b942-bb44-41dd-ed64-93f140d0c4ee"
fraud.Amount.describe() # not mendatory I just written it for show the details of fraud transaction.
# we can also written the code --- normal.Amount.describe() --- for show the details of normal transaction.
# + [markdown] id="Iu-pFLJhUIfT"
# Here, we are using **Matplotlib** for using this graph
# + colab={"base_uri": "https://localhost:8080/", "height": 308} id="M5h_QumAQTv9" outputId="e815d46d-cfb9-4598-a909-319c3f0448e9"
# not mendatory I just written it for show the graphs of transaction
f, (ax1, ax2) = plt.subplots(2, 1, sharex=True)
f.suptitle('Amount per transaction by class')
bins = 50
ax1.hist(fraud.Amount, bins = bins)
ax1.set_title('Fraud')
ax2.hist(normal.Amount, bins = bins)
ax2.set_title('Normal')
plt.xlabel('Amount ($)')
plt.ylabel('Number of Transactions')
plt.xlim((0, 20000))
plt.yscale('log')
plt.show();
# + [markdown] id="dil2yflfTy44"
# For the 1st graph it shows that the **Fraud Transaction amount is very less** but for the 2nd one **Normal Transaction amount is very very high**
# + [markdown] id="C6Z-Pz-ZuKa_"
# ---
#
# By seeing the distributions we can have an idea how skewed are these features, we can also see further distributions of the other features. There are techniques that can help the distributions be less skewed which will be implemented in this notebook in the future.
#
# Doesn't seem like the time of transaction really matters here as per above observation. Now let us take a sample of the dataset for out modelling and prediction
#
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="qMHHAmFkpj6f" outputId="372813a1-1cbe-41c1-8c0e-3b6d6dcd22f3"
data.hist(figsize=(20, 20));#Plot a histogram using matplotlib.
# + colab={"base_uri": "https://localhost:8080/"} id="EYVHZl8uwkLL" outputId="9be2ed4e-5be1-4f4d-a69b-26a918d8e33c"
## There are huge data which can take too much time so we can take some sample of the data
data1= data.sample(frac = 0.1,random_state=1) # taking 0.1% from the whole dataset
data1.shape
# + id="b8rNyoltQThl"
# From 0.1% dataset we are detemining how many are Fraud & Normal Transaction
Fraud = data1[data1['Class']==1]
Valid = data1[data1['Class']==0]
outlier_fraction = len(Fraud)/float(len(Valid))
# + colab={"base_uri": "https://localhost:8080/"} id="LasODl8GQTT6" outputId="f6848721-f45b-416e-dcd6-ebd45b243e7f"
print(outlier_fraction)
print("Fraud Cases : {}".format(len(Fraud)))
print("Valid Cases : {}".format(len(Valid)))
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="TZWX3YCuw5MB" outputId="739cb15e-edb6-4f9e-de48-9519907e7d82"
## Correlation
import seaborn as sns
#get correlations of each features in dataset
corrmat = data1.corr()
top_corr_features = corrmat.index
plt.figure(figsize=(20,20))
#plot heat map
g=sns.heatmap(data[top_corr_features].corr(),annot=True)
# + [markdown] id="gnflKnayU6DF"
# Above we are using **Correlation** to analyse the relationship between class and other variables.
# + colab={"base_uri": "https://localhost:8080/"} id="3TRzMLDGxJ08" outputId="c0c6c1b8-e7d1-490b-89c2-e04c977db87e"
#Create independent and Dependent Features
columns = data1.columns.tolist()
# Filter the columns to remove data we do not want
columns = [c for c in columns if c not in ["Class"]] #codition- when there is a class variable take as dependent variable otherwise, independent
# Store the variable we are predicting
target = "Class"
# Define a random state
state = np.random.RandomState(42)
X = data1[columns]
Y = data1[target]
X_outliers = state.uniform(low=0, high=1, size=(X.shape[0], X.shape[1]))
# Print the shapes of X & Y
print(X.shape)
print(Y.shape)
# + [markdown] id="xuxL1tsOchsr"
# Now we are going to implement the algorithm for **Model Prediction** -----
#
#
# * **Isolation Forest Algorithm :**This method is highly useful and is
# fundamentally different from all existing methods. Moreover, this method is an algorithm with a low linear time complexity and a small memory requirement. It builds a good performing model with a small number of trees using small sub-samples of fixed size, regardless of the size of a data set.
#
# **process :** If we consider a decision tree it split the data. For that there are soo many outliers(leaf node) which can decrease the accuracy of the dataset. So we are using Isolation Forest Algorithm which work based upon Random Forest. It can split the dataset into 2 portion - dalit dataset and outliers so accuracy will increased automatically.
#
# * **Local Outlier Factor(LOF) Algorithm :** LOF is an algorithm used for outlier detection. It produces an anomaly score that represents data points which are outliers in the data set.
#
# **process :** Local density is determined by estimating distances between data points that are neighbors (k-nearest neighbors). So for each data point, local density can be calculated. By comparing these we can check which data points have similar densities and which have a lesser density than its neighbors. The ones with the lesser densities are considered as the outliers.
#
# * **Support Vector Machine(SVM) :** The SVM classifier is a frontier which best segregates the two classes (hyper-plane/ line).
#
# **process :** In the SVM algorithm, we plot each data item as a point in n-dimensional space (where n is number of features you have) with the value of each feature being the value of a particular coordinate. Then, we perform classification by finding the hyper-plane that differentiates the two classes very well.
#
# + id="HwsayW8FxqAO"
from sklearn.metrics import classification_report,accuracy_score
from sklearn.ensemble import IsolationForest
from sklearn.neighbors import LocalOutlierFactor
from sklearn.svm import OneClassSVM
from pylab import rcParams
rcParams['figure.figsize'] = 14,8
RANDOM_SEED = 42
LABELS = ["Normal","Fraud"]
# + id="DG_2d3M1xOLN"
# Creating dictionary for Isolation Forest,LOC,SVM Algorithm
classifiers = {
"Isolation Forest":IsolationForest(n_estimators=100, max_samples=len(X),
contamination=outlier_fraction,random_state=state, verbose=0),
"Local Outlier Factor":LocalOutlierFactor(n_neighbors=20, algorithm='auto',
leaf_size=30, metric='minkowski',
p=2, metric_params=None, contamination=outlier_fraction),
"Support Vector Machine":OneClassSVM(kernel='rbf', degree=3, gamma=0.1,nu=0.05,
max_iter=-1, )
}
# + colab={"base_uri": "https://localhost:8080/"} id="P3YuuajpyxlD" outputId="4dfd4dc2-9af8-4c76-9f64-b26aa8a46322"
type(classifiers)
# + colab={"base_uri": "https://localhost:8080/"} id="cZOVkIrP2OCq" outputId="5a5d1384-e895-4fef-cd46-eec9424f8e3e"
# Implemeting the algorithms
n_outliers = len(Fraud)
for i, (clf_name,clf) in enumerate(classifiers.items()):
#Fit the data and tag outliers
if clf_name == "Local Outlier Factor":
y_pred = clf.fit_predict(X)
scores_prediction = clf.negative_outlier_factor_
elif clf_name == "Support Vector Machine":
clf.fit(X)
y_pred = clf.predict(X)
else:
clf.fit(X)
scores_prediction = clf.decision_function(X)
y_pred = clf.predict(X)
#Reshape the prediction values to 0 for Valid transactions , 1 for Fraud transactions
y_pred[y_pred == 1] = 0
y_pred[y_pred == -1] = 1
n_errors = (y_pred != Y).sum()
#################################################RESULT###############################################
# Run Classification Metrics
print("{}: {}".format(clf_name,n_errors))
print("Accuracy Score :")
print(accuracy_score(Y,y_pred))
print("Classification Report :")
print(classification_report(Y,y_pred))
# + [markdown] id="DSoPdgyhl1Sp"
# * **Isolation Forest** detected 73 errors, has accuracy 99.74%
#
# * **Local Outlier Factor** detecting 97 errors, has accuracy 99.65%
#
# * **SVM** detecting 8516 errors, has accuracy 70.90%
#
# When comparing error precision & recall for 3 models , the **Isolation Forest performed much better** than the LOF as we can see that the detection of fraud cases is around 27 % versus LOF detection rate of just 2 % and SVM of 0%.
#
# So overall Isolation Forest Method performed much better in determining the fraud cases which is around 30%.
#
# + [markdown] id="_tD5a2fXm346"
# ---
#
# We can also improve on this accuracy by increasing the **sample size or use deep learning algorithms** however at the cost of computational expense.We can also use complex anomaly detection models to get better accuracy in determining more fraudulent cases.
# + [markdown] id="ddx4yAbPSety"
# **END**
# ---
# + [markdown] id="XVP82oijzIJT"
# **NOT FOR NOW**
# + [markdown] id="mOlAEiYLx7JM"
# ---
# **Data Pre-processing**
#
# Time and Amount should be scaled as the other columns.
# + colab={"base_uri": "https://localhost:8080/"} id="UttVexwWxgdm" outputId="72201d60-6258-4764-a554-1fefe94313a5"
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
scalar = StandardScaler()
X = data.drop('Class', axis=1)
y = data.Class
X_train_v, X_test, y_train_v, y_test = train_test_split(X, y,
test_size=0.3, random_state=42)
X_train, X_validate, y_train, y_validate = train_test_split(X_train_v, y_train_v,
test_size=0.2, random_state=42)
X_train = scalar.fit_transform(X_train)
X_validate = scalar.transform(X_validate)
X_test = scalar.transform(X_test)
w_p = y_train.value_counts()[0] / len(y_train)
w_n = y_train.value_counts()[1] / len(y_train)
print(f"Fraudulant transaction weight: {w_n}")
print(f"Non-Fraudulant transaction weight: {w_p}")
# + colab={"base_uri": "https://localhost:8080/"} id="Egf3VFJiyQWM" outputId="291c53ca-014b-426d-ac4e-fbccea7dd088"
print(f"TRAINING: X_train: {X_train.shape}, y_train: {y_train.shape}\n{'_'*55}")
print(f"VALIDATION: X_validate: {X_validate.shape}, y_validate: {y_validate.shape}\n{'_'*50}")
print(f"TESTING: X_test: {X_test.shape}, y_test: {y_test.shape}")
# + id="wzRfGNwZyy5H"
from sklearn.metrics import accuracy_score, confusion_matrix, classification_report, f1_score
def print_score(label, prediction, train=True):
if train:
clf_report = pd.DataFrame(classification_report(label, prediction, output_dict=True))
print("Train Result:\n================================================")
print(f"Accuracy Score: {accuracy_score(label, prediction) * 100:.2f}%")
print("_______________________________________________")
print(f"Classification Report:\n{clf_report}")
print("_______________________________________________")
print(f"Confusion Matrix: \n {confusion_matrix(y_train, prediction)}\n")
elif train==False:
clf_report = pd.DataFrame(classification_report(label, prediction, output_dict=True))
print("Test Result:\n================================================")
print(f"Accuracy Score: {accuracy_score(label, prediction) * 100:.2f}%")
print("_______________________________________________")
print(f"Classification Report:\n{clf_report}")
print("_______________________________________________")
print(f"Confusion Matrix: \n {confusion_matrix(label, prediction)}\n")
# + [markdown] id="6wYPeiF6a8aT"
# **Random Forest**
# + id="wf8x-yc4uw87" colab={"base_uri": "https://localhost:8080/"} outputId="03aaf692-eb9d-4b17-fe04-236c841bf97e"
from sklearn.ensemble import RandomForestClassifier
output_dict = dict()
rf_clf = RandomForestClassifier(n_estimators=100, oob_score=False)
rf_clf.fit(X_train, y_train)
y_train_pred = rf_clf.predict(X_train)
y_test_pred = rf_clf.predict(X_test)
print_score(y_train, y_train_pred, train=True)
print_score(y_test, y_test_pred, train=False)
output_dict['Random Forest'] = {
'Train': f1_score(y_train,y_train_pred),
'Test': f1_score(y_test, y_test_pred),
}
# + [markdown] id="DEhzTt6-bDEn"
# Using Random Forest we are getting **99.96%** accuracy
# + colab={"base_uri": "https://localhost:8080/"} id="yGNkU3ZG20L4" outputId="dc16fb39-d9c2-435c-82d1-99750c61fdc2"
# !pip install catboost
# + colab={"base_uri": "https://localhost:8080/"} id="Jqayb0_l1qne" outputId="8a8a5053-22bf-43a0-ec6b-57b3a8f50b70"
from catboost import CatBoostClassifier
cb_clf = CatBoostClassifier()
cb_clf.fit(X_train, y_train)
# + colab={"base_uri": "https://localhost:8080/"} id="zH0uRKZ94gyQ" outputId="3abfd185-215e-49ce-ca85-0e915fb7516d"
output_dict = dict()
y_train_pred = cb_clf.predict(X_train)
y_test_pred = cb_clf.predict(X_test)
print_score(y_train, y_train_pred, train=True)
print_score(y_test, y_test_pred, train=False)
output_dict['CatBoost'] = {
'Train': f1_score(y_train,y_train_pred),
'Test': f1_score(y_test, y_test_pred),
}
# + id="6inKx6_MyxQH" colab={"base_uri": "https://localhost:8080/"} outputId="a6c44049-238d-4d51-e838-4461e8aa0ff8"
# !pip install tensorflow-gpu==2.0.0-rc0
| creditcard.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="IcEo2uf5pIfW"
# # Recognition of clothing items (fashionMNIST) with neural networks
# + [markdown] id="ylLG-vS8pSVd"
# ## Setup
# + id="FmoXKtcXoBrx"
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import tensorflow as ts
import tensorflow.keras as keras
from keras.utils import to_categorical
from keras.utils.vis_utils import plot_model
from tensorflow.keras.datasets import fashion_mnist
from tensorflow.keras import models, layers
from keras.layers import Dense, Dropout, Flatten
from keras.layers import Conv2D, MaxPooling2D, BatchNormalization
from sklearn.metrics import confusion_matrix, classification_report
# + id="gNWzlJLUoSDm"
#data = tfds.load('mnist', split='train', as_supervised=True, shuffle_files=True)
(xtrain, ytrain), (xtest, ytest) = fashion_mnist.load_data()
# + id="6drJ737bmLmH"
class_names = ['T-shirt/top', 'Trouser', 'Pullover', 'Dress', 'Coat',
'Sandal', 'Shirt', 'Sneaker', 'Bag', 'Ankle boot']
# + colab={"base_uri": "https://localhost:8080/"} id="5wSogmertjhi" outputId="ed1d8577-6135-4059-f2b7-320a01c2f061"
# shape of a single data point
xtrain[0].shape
# + colab={"base_uri": "https://localhost:8080/"} id="uHthFIx39A46" outputId="007502f6-9fd2-4964-bc5c-eec6dde06b11"
xtrain.shape
# + colab={"base_uri": "https://localhost:8080/"} id="Appi2GsPb4HF" outputId="b236bec7-1bfc-4c5f-be9c-57000bcaa603"
ytrain.shape
# + colab={"base_uri": "https://localhost:8080/"} id="oodi3lF_9D9P" outputId="5859db1f-2cd7-46c1-bd03-aa091073c0c9"
xtest.shape
# + [markdown] id="bQGiA0ayoNfV"
# ### Image preprocessing
# + colab={"base_uri": "https://localhost:8080/", "height": 248} id="2yowe24MoVUF" outputId="0977af03-7bd4-4785-8961-36740f38e628"
for i in range(25):
plt.subplot(5, 5, i+1)
plt.imshow(xtrain[i], cmap=plt.cm.Greys)
plt.axis('off')
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="GnegXab9n1Y8" outputId="a37df87b-f975-497e-901b-1c81796adf73"
# Inspect the pixel values -> they fall in the range of 0 to 255
plt.figure()
plt.imshow(xtrain[0])
plt.colorbar()
plt.grid(False)
plt.show()
# + id="MTzGZRLgogO0"
# Scale the pixel values to a range of 0 to 1
xtrain = xtrain.astype('float32') / 255
xtest = xtest.astype('float32') / 255
# + colab={"base_uri": "https://localhost:8080/", "height": 589} id="pGhSZLeGod85" outputId="d1e6cca9-cdf0-494f-a1e8-c7931287efc2"
plt.figure(figsize=(10,10))
for i in range(25):
plt.subplot(5,5,i+1)
plt.xticks([])
plt.yticks([])
plt.grid(False)
plt.imshow(xtrain[i], cmap=plt.cm.binary)
plt.xlabel(class_names[ytrain[i]])
plt.show()
# + id="8d4g5Fesy0er"
# one-hot-enocde the labels
#ytrain = to_categorical(ytrain, num_classes=10)
#ytest = to_categorical(ytest, num_classes=10)
# + [markdown] id="L6Hz1NkgolSz"
# ## Artificial Neural Networks
# + id="QDd4H1LjoYuY"
model = models.Sequential()
# model.add(layers.Flatten(input_shape=(28,28)))
# model.add(layers.Dense(128, activation='relu'))
# model.add(layers.Dropout(0.25))
# model.add(layers.Dense(100, activation='relu'))
# model.add(layers.Dropout(0.5))
# model.add(layers.Dense(10, activation='softmax'))
# Test accuracy: 0.880
# + id="F8BvZCCveQ-Z"
#model.add(layers.Flatten(input_shape=(28,28)))
#model.add(layers.Dense(300, activation='relu'))
#model.add(layers.Dense(128, activation='relu'))
#model.add(layers.Dense(10, activation='softmax'))
# Test accuracy: 0.886
# + id="einmVMW5eQlX"
model.add(layers.Flatten(input_shape=(28,28)))
model.add(layers.Dense(2048, activation='relu'))
model.add(layers.Dense(1024))
model.add(layers.BatchNormalization())
model.add(layers.Activation('relu'))
model.add(layers.Dense(1024))
model.add(layers.BatchNormalization())
model.add(layers.Activation('relu'))
model.add(layers.Dense(10))
model.add(layers.BatchNormalization())
model.add(layers.Activation('softmax')))
# Epochs: 100
# Batch size: 256
# Train accuracy: 0.905
# Train loss: 0.263
# Test accuracy: 0.88
# Test loss: 0.333
# + id="5-LsArIQ2Xdb"
"""
Model with SGD optimizer
with Nesterov momentum to speed up the convergence.
The performance became worse after 66/100 epochs --> the model reached the basin
of a minimim but kept on jumping on the sides without reaching it.
--> set a smaller learning rate and a proportionally longer training process.
"""
sgd = keras.optimizers.SGD(
lr=0.001,
momentum=0.8,
nesterov=True
)
rmsp = keras.optimizers.RMSprop(
lr=0.001,
rho=0.8
)
adam = keras.optimizers.Adam()
# + id="M7ha4gERsN3V"
#model.compile(optimizer='adam',loss='categorical_crossentropy',metrics=['accuracy'])
model.compile(
optimizer='adam',
#loss='categorical_crossentropy',
loss=keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy']
)
# + colab={"base_uri": "https://localhost:8080/"} id="IPq9jOMm98D-" outputId="2226723f-1f39-43e3-ff09-755e2b1770ac"
model.summary()
# + colab={"base_uri": "https://localhost:8080/"} id="CW-9kZqau2CN" outputId="00412144-5d1b-4828-d15b-46ac7ca786d5"
#history = model.fit(xtrain, ytrain, epochs=500, batch_size=500, validation_split = 0.20)
history = model.fit(
xtrain,
ytrain,
epochs=50,
batch_size=500,
validation_data=(xtest, ytest))
# + [markdown] id="RVsJio3GvX9c"
# ## Model evaluation
# + colab={"base_uri": "https://localhost:8080/"} id="h0E5_n_29-E6" outputId="e55041ab-49a6-4ab2-b60c-d4e12ef3d604"
train_loss, train_acc = model.evaluate(xtrain, ytrain)
test_loss, test_acc = model.evaluate(xtest, ytest)
print("Train accuracy:", round(train_acc, 5))
print("Train loss:", round(train_loss, ))
print("Test accuracy:", round(test_acc, 5))
print("Test loss:", round(test_loss, 3))
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="qgqHlWD3w19m" outputId="bbd9ad13-bb31-47ca-8cc5-4c9d63cb7777"
plt.plot(history.history["accuracy"])
plt.plot(history.history["val_accuracy"])
plt.title("Model Accuracy")
plt.xlabel('Epochs')
plt.ylabel('Accuracy')
plt.legend(['Train', 'Test'])
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="i7aEExdXAzzP" outputId="5815508f-8bcb-48b5-fb3d-f64c3004de75"
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title("Model Loss")
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend(['Train', 'Test'])
plt.show()
# + id="-nLuUa0nCXcC"
plot_model(model, to_file='fmnist1.png',show_shapes=True, show_layer_names=True)
model.save("fmnist1.h5")
#load_model("fmnist.h5")
# + [markdown] id="2ciEEcA2viU4"
# ## Confusion matrix
# + id="b_tzSmMPvn8n"
def plot_confusion_matrix(cm, classes,
normalize=False,
title='Confusion matrix',
cmap=plt.cm.Blues):
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=90)
plt.yticks(tick_marks, classes)
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, cm[i, j],
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
# + id="hny2hG8MvtOx"
# Predict the values from the validation dataset
Y_pred = model.predict(x_test)
# Convert predictions classes to one hot vectors
Y_pred_classes = np.argmax(Y_pred,axis = 1)
# Convert validation observations to one hot vectors
Y_true = np.argmax(y_test,axis = 1)
# compute the confusion matrix
confusion_mtx = confusion_matrix(Y_true, Y_pred_classes)
# plot the confusion matrix
plot_confusion_matrix(confusion_mtx,
classes = ['T-shirt/Top','Trouser','Pullover','Dress','Coat','Sandal','Shirt','Sneaker','Bag','Ankle Boot'])
# + [markdown] id="ucWNaaDbqdkn"
# # Predictions
# + colab={"base_uri": "https://localhost:8080/"} id="E-uwn5ctnhFc" outputId="7a22a2fd-da33-4a3f-e1a7-babea53179de"
probability_model = keras.Sequential([model, layers.Softmax()])
predictions = probability_model.predict(xtest)
predictions[0]
# + colab={"base_uri": "https://localhost:8080/"} id="3pNXmUgLqfLD" outputId="34875ad2-6334-477b-f74b-0d1995d1589e"
# Check the label with the highest confidence value
np.argmax(predictions[0])
# + colab={"base_uri": "https://localhost:8080/"} id="L-wU6RbDqzy9" outputId="d582f41b-9025-4ff8-aa7c-f79852c45da7"
# Verify the prediction of the label above
ytest[0]
#plt.imshow(ytest[0])
# + id="zlKH95YS7OFY" colab={"base_uri": "https://localhost:8080/", "height": 211} outputId="33a679d6-e432-4ccd-ee82-92be38f12919"
def plot_image(i, predictions_array, ytrain, img):
ytrain, img = ytrain[i], img[i]
plt.grid(False)
plt.xticks([])
plt.yticks([])
plt.imshow(img, cmap=plt.cm.binary)
ytest = np.argmax(predictions_array)
if ytest == ytrain:
color = 'blue'
else:
color = 'red'
plt.xlabel("{} {:2.0f}% ({})".format(class_names[ytest],
100*np.max(predictions_array),
class_names[ytrain]),
color=color)
def plot_value_array(i, predictions_array, ytrain):
ytrain = ytrain[i]
plt.grid(False)
plt.xticks(range(10))
plt.yticks([])
thisplot = plt.bar(range(10), predictions_array, color="#777777")
plt.ylim([0, 1])
ytest = np.argmax(predictions_array)
thisplot[ytest].set_color('red')
thisplot[ytrain].set_color('blue')
i = 0
plt.figure(figsize=(6,3))
plt.subplot(1,2,1)
plot_image(i, predictions[i], ytest, xtest)
plt.subplot(1,2,2)
plot_value_array(i, predictions[i], ytest)
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 211} id="EFSzHTSa0dAm" outputId="49a2cfc3-78e9-46d5-f073-c39d55b478c4"
i = 12
plt.figure(figsize=(6,3))
plt.subplot(1,2,1)
plot_image(i, predictions[i], ytest, xtest)
plt.subplot(1,2,2)
plot_value_array(i, predictions[i], ytest)
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 665} id="cn3zwajo0kf-" outputId="a075de2b-63ae-461f-d784-1333cd285eba"
# Plot the first X test images, their predicted labels, and the true labels.
# Color correct predictions in blue and incorrect predictions in red.
num_rows = 5
num_cols = 3
num_images = num_rows*num_cols
plt.figure(figsize=(2*2*num_cols, 2*num_rows))
for i in range(num_images):
plt.subplot(num_rows, 2*num_cols, 2*i+1)
plot_image(i, predictions[i], ytest, xtest)
plt.subplot(num_rows, 2*num_cols, 2*i+2)
plot_value_array(i, predictions[i], ytest)
plt.tight_layout()
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="1lX-TcrQ0-N7" outputId="a506d4fc-3524-4bd7-d7f5-aa717736850a"
# Grab an image from the test dataset.
img = xtest[1]
# Add the image to a batch where it's the only member.
img = (np.expand_dims(img,0))
print(img.shape)
# + colab={"base_uri": "https://localhost:8080/"} id="it7whyoX1H5s" outputId="17d6652d-3d80-45f9-e88e-31d78a9bff77"
# predict the correct label for this image
predictions_single = probability_model.predict(img)
print(predictions_single)
# + colab={"base_uri": "https://localhost:8080/", "height": 300} id="G4Oi0MVs1L9r" outputId="3dccf58a-9193-4c29-d6fd-61bc8de4c5ca"
plot_value_array(1, predictions_single[0], ytest)
_ = plt.xticks(range(10), class_names, rotation=45)
# + colab={"base_uri": "https://localhost:8080/"} id="00wr0JNs1Sgk" outputId="a7bc73cf-9d9d-4e08-e646-4b4098d08f01"
#predictions for the only image in the batch:
np.argmax(predictions_single[0])
# + colab={"base_uri": "https://localhost:8080/", "height": 436} id="6_MtXxka3FS6" outputId="5482bfd9-74c5-4b50-ae75-64c54135b511"
Y_pred_classes = np.argmax(predictions,axis = 1)
cm = confusion_matrix(ytest, Y_pred_classes)
df_cm = pd.DataFrame(cm, range(10), range(10))
plt.figure(figsize=(10,7))
sns.set(font_scale=1.4) # for label size
sns.heatmap(df_cm, annot=True, annot_kws={"size": 10}) # font size
plt.show()
| Fashion_MNIST.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={} colab_type="code" id="lS6-gL_MH4aU"
## Notes:
#### Structuring my Project References: https://docs.python-guide.org/writing/structure/
# +
### TODO
# + colab={} colab_type="code" id="UPZ1jl2FH4ak"
# Necessary Librarires
import random
import json
try:
import cPickle as pickle
except ModuleNotFoundError:
import pickle
# import pickle
global ResponseKnowledgeBase
global UserLibrary
global OpportunityLibrary
global CurrentUser
global CurrentOpportunity
# + [markdown] colab_type="text" id="LoPDlwtJH4bB"
# ## Knowledge Base
# +
## getResponseKB() retrieves the response knowledge base for use and updating
def getResponseKB():
rkb = json.loads(open('rkb.txt', 'r').read())
global ResponseKnowledgeBase
ResponseKnowledgeBase = rkb
return
# rkb ## JDF Test
# rkb["Continuous"]["VillainousForce"]["Actualization"] ## JDF Test
# rkb["Continuous"]["VillainousForce"]["Actualization"].append("Actualization val5") ## JDF Test: Value addition
# rkb["Continuous"]["VillainousForce"]["Actualization"] ## JDF Test
# -
def updateResponseKB():
global ResponseKnowledgeBase
rkb = ResponseKnowledgeBase
with open('rkb.txt', 'w') as file:
file.write(json.dumps(rkb)) # use `json.loads` to do the reverse
return
# + [markdown] colab_type="text" id="KO9uH5cuH4bR"
# ## User Creation
# + colab={} colab_type="code" id="DXaCR6xFH4bV"
class User:
def __init__(self, name, riskThreshold, reflections):
# def __init__(self, name, personality, riskThreshold, reflections): ## JDF Version 2.0
self.name = name
# self.personality = personality ## JDF Version 2.0
self.riskThreshold = riskThreshold
#list of reflections user has engaged in
self.reflections = reflections
# + colab={} colab_type="code" id="Ze4L5XBxH4bd"
def getUserName():
userName = input("Hi. What's your user name? ")
return (userName)
# getUserName() # JDF Testing
# + colab={} colab_type="code" id="grsBVmofH4bk"
# ## Version 2.0
# ## Designed to support provision of a user specific motivation prompt.
# def getUserPersonality():
# userPersonality = input("What's your enneagram type? ")
# # print("You identified as an Enneagram type ", userPersonality)
# return (userPersonality)
# # getUserPersonality() # JDF Testing
# + colab={} colab_type="code" id="63mWiP0RH4bo"
def getUserRiskThreshold():
riskThreshold = input("What's your risk tolerance? (H, M, L) ")
# print("You identified your risk tolerance level as ", riskThreshold)
return(riskThreshold)
# getUserRiskThreshold() # JDF Testing
# + colab={} colab_type="code" id="Ec134aIcH4br"
def create_user():
userName = getUserName()
if userName.lower() == "quit":
return (quitProgram()) ## JDF for tests
# userPersonality = getUserPersonality() ## JDF Version 2.0
userRiskThreshold = getUserRiskThreshold()
reflections = []
# user = User(userName, userPersonality, userRiskThreshold, reflections) ## JDF Version 2.0
user = User(userName, userRiskThreshold, reflections)
# Print("Welcome to Hero Academy, ", CurrentUser.name)
return(user)
# create_user() ## JDF Test
# + [markdown] colab_type="text" id="7oxjxeUKH4b7"
# ## Opportunity Creation
# + colab={} colab_type="code" id="ogNkyPPwH4b8"
class Opportunity:
def __init__(self, villainType, threatDuration, threatType):
self.villainType = villainType
self.threatDuration = threatDuration
self.threatType = threatType
# + colab={} colab_type="code" id="PO0FPS4PH4cC"
def getVillainType():
while True:
userInput = input("Is the cause of the threat an Agent, System, or Force? (A, S, F, or info) ")
if userInput.lower() == "info":
print('''
Villains are agents, systems, or forces that actively exploit and/or endanger human beings to advance its own interests.
In the case of villainous agents, exploitation must be conscious. Someone is consciously creating the threat.
In the case of villainous systems, exploitation must perpetuate the system. A formal or informal policy furthers the threat.
In the case of villainous forces, exploitation or danger must be impersonal. An exteranal factor tied to a contingency rather than a person or structure.
''')
pass
elif userInput.upper() == "A":
villainType = "VillainousAgent"
break
elif userInput.upper() == "S":
villainType = "VillainousSystem"
break
elif userInput.upper() == "F":
villainType = "VillainousForce"
break
elif userInput.lower() == ("quit"):
return(quitProgram())
# elif if villainType.upper() not in ['A', 'S', 'F']:
else:
print("Your options are 'A' for Agent, 'S' for System, or 'F' for Force. Type 'info' to learn more about these options.")
print("You can also type 'quit' to end the program at any time.")
pass
return(villainType)
# getVillainType() # JDF Testing
# + colab={} colab_type="code" id="c_qT4Q7QH4cE"
def getThreatDuration():
while True:
userInput = input("Is the threat duration Episodic or Continuous? (E, C, or info) ")
if userInput.lower() == "info":
print('''
Heroic opportunities can be Episodic or Continuous as determined by the nature of the threat.
If the problem is a one time thing, mark it as Episodic. If it happens repeatedly, mark it as Continuous.
''')
pass
elif userInput.upper() == "E":
threatDuration = "Episodic"
break
elif userInput.upper() == "C":
threatDuration = "Continuous"
break
elif userInput.lower() == ("quit"):
return(quitProgram())
else:
print("Your options are 'E' for Episodic or 'C' for Continuous. Type 'info' to learn more about these options.")
print("You can also type 'quit' to end the program at any time.")
pass
return(threatDuration)
# getThreatDuration() # JDF Testing
# + colab={} colab_type="code" id="EjoC18VNH4cJ"
def getThreatType():
while True:
userInput = input("Is the threat Physiological, Safety, Belonging, Esteem, or Actualization related (P, S, B, E, A, or info) ")
if userInput.lower() == "info":
print('''
Physiological threats cause direct bodily damage to the person you're seeking to help. Threats to health, food and water, sleep, clothes, and shelter are included in this category.
Safety threats cause damage to someone's personal, emotional, financial, or indirect risk of future physical harm.
Belonging threats cause damage to social bonds, such as the bonds of family, friendship, or community.
Esteem threats run the risk of damaging someone's self-respect or proper perception of self.
Actualization threats run the risk of damaging someone's potential to pursue their goals, dreams, and utilize their gifts.
''')
pass
elif userInput.upper() == "P":
threatType = "Physiological"
break
elif userInput.upper() == "S":
threatType = "Safety"
break
elif userInput.upper() == "B":
threatType = "Belonging"
break
elif userInput.upper() == "E":
threatType = "Esteem"
break
elif userInput.upper() == "A":
threatType = "Actualization"
break
elif userInput.lower() == ("quit"):
return(quitProgram())
elif userInput.lower() == ("main"):
return
else:
print("Your options are 'P', 'S', 'B', 'E', or 'A'. \nType 'info' to learn more about these options.")
print("You can also type 'quit' to end the program at any time.")
pass
return(threatType)
# getThreatType() # JDF Testing
# +
def getOpportunityDescription():
userInput = input("Describe the situation in a single sentence. ")
opportunityDescription = userInput
if userInput.lower() == ("quit"):
return(quitProgram())
else:
return(opportunityDescription)
# getOpportunityDescription() ## JDF Test
# +
def create_opportunity():
# Used for archiving opportunities
opportunityDescription = getOpportunityDescription()
if opportunityDescription == quitProgram():
return (quitProgram()) ## JDF for tests
villainType = getVillainType()
if villainType == quitProgram():
return (quitProgram()) ## JDF for tests
threatDuration = getThreatDuration()
if threatDuration == quitProgram():
return (quitProgram()) ## JDF for tests
threatType = getThreatType()
if threatType == quitProgram(): ## JDF for tests
return (quitProgram())
# Creates the opportunity
opportunity = Opportunity(villainType, threatDuration, threatType)
# Stores the opportunity as a global variable
CurrentOpportunity = opportunity
# Stores the opporutnity with its description
OpportunityLibrary[opportunityDescription] = opportunity
return(opportunity)
# create_opportunity() #JDF test
# + [markdown] colab_type="text" id="iS_OdhdkH4ch"
# ## Login Prompt
# + colab={} colab_type="code" id="DIF5HTpXH4ch"
def login_prompt():
global currentUser
while True:
print("\nSIGN IN")
userInput = input("Is this your first time here? (y, n, or menu) ")
if userInput.lower() == 'menu':
return
elif userInput == quitProgram():
return (quitProgram()) ## JDF for tests
elif userInput == "quit":
return (quitProgram()) ## JDF for tests
elif userInput.lower() == 'y':
newUser = create_user()
if newUser == quitProgram():
return (quitProgram()) ## JDF for tests
# elif userInput == "quit":
# return (quitProgram()) ## JDF for tests
UserLibrary[newUser.name] = newUser
CurrentUser = UserLibrary[newUser.name]
return(CurrentUser)
elif userInput.lower() == 'n':
userName = input("What's your username? ")
if userName in list(UserLibrary.keys()):
print("Welcome back,", userName + ".")
CurrentUser = UserLibrary[userName]
return(CurrentUser)
else:
print("I'm sorry, your username isn't in our database.")
print("\nUsers include: ", list(UserLibrary.keys()))
else:
print('''I'm sorry. Your options are to type 'y' for yes or 'n' for no.
\nYou can also return to the main menu by typing 'menu'. ''')
# tempUserCatcher = login_prompt() #JDF Test
# tempUserCatcher
# login_prompt() ## JDF Test
# -
# ## Opportunity Analysis
# +
# getResponseKB()
# +
# global ResponseKnowledgeBase
# ResponseKnowledgeBase
# +
## Uses the villainType, threatDuration, threatType to access the expanding library of opportunities.
def opportunityAnalysis():
newOpportunity = create_opportunity()
if newOpportunity == quitProgram():
return (quitProgram())
# elif newOpportunity.threatDuration == quitProgram():
# return (quitProgram())
# elif newOpportunity.threatType == quitProgram():
# return (quitProgram())
else:
## Save
global ResponseKnowledgeBase
opportunityResponse = ResponseKnowledgeBase[newOpportunity.threatDuration][newOpportunity.villainType][newOpportunity.threatType]
recommendedResponse = random.choice(opportunityResponse)
return (recommendedResponse)
# opportunityAnalysis() ## JDF Test
# -
# ## Main Menu & Program Navigation
# +
# def getUserLibrary():
# return list(UserLibrary.keys())
# # ## User Library Test
# # UserNames = UserLibrary.keys() # JDF Test
# getUserLibrary()
# -
def aboutProgram():
print('''
The world of tomorrow needs heroes. Not superheroes with fancy powers or flashy costumes, but everyday heroes who engage in intentional, everyday actions that better the lives of those within reach. This program is designed to support you in the pursuit of everyday heroism. It will give you the opportunity to answer questions about situations you're facing and it will point out risks and provide recommendations concerning the situation.
The program also gives you the opportunity to reflect on situations you haven't been in personally. Your advice and reflections will be added to the knowledge base to support future users.
''')
return
# +
def mainMenu():
#Initializes the user as a guest
global CurrentUser
CurrentUser = User("GUEST", "M", [])
while True:
userInput = input('''\nMENU
Type the letter that corresponds with the menu item you would like to select.
About the Heroism Reasoner (a)
Launch the Heroism Reasoner (l)
Sign-in (s)
User Library (u)
(You can also type 'quit' to end the program at any time.)
''')
if userInput.lower() == ("a"):
userInput = aboutProgram()
if userInput == quitProgram():
return (quitProgram()) ## JDF for tests
else:
pass
elif userInput.lower() == ("l"):
responseToUser = opportunityAnalysis()
if userInput == quitProgram():
return (quitProgram()) ## JDF for tests
else:
print(responseToUser)
pass
elif userInput.lower() == ("s"):
userInput = login_prompt()
if userInput == quitProgram():
return (quitProgram()) ## JDF for tests
else:
pass
elif userInput.lower() == ("u"):
print("\nUsers include: ", list(UserLibrary.keys()))
pass
elif userInput.lower() == ("quit"):
return(quitProgram())
else:
print("Sorry. Choose from the menu options above or 'quit' to end the program at any time")
## print(userInput) ##JDF Debugging
# mainMenu() ##JDF Test
# -
# ## Housekeeping Functions
def quitProgram():
# print("Come again soon. We have much to discuss. You've exited the program.") ## Actual quit program
# return(exit(0)) ## Actual quit program
global UserLibrary
save_object(UserLibrary, 'UserLibrary.pkl')
global OpportunityLibrary
save_object(OpportunityLibrary, 'OpportunityLibrary.pkl')
updateResponseKB()
return("Come again soon. We have much to discuss. You've exited the program.") ## Helpful for tests
# +
## Save User Library (or any object)
def save_object(obj, filename):
with open(filename, 'wb') as output: # Overwrites any existing file.
pickle.dump(obj, output, pickle.HIGHEST_PROTOCOL)
return
# ## JDF Test
# global UseerLibrary
# save_object(UserLibrary, 'UserLibrary.pkl')
# -
# ## Opportunity Library
# +
# OpportunityLibrary = {}
# +
## Import Opportunity Library
## Import Opportunity Library
def getOpportunityLibrary():
global OpportunityLibrary
OpportunityLibrary = pickle.load(open("OpportunityLibrary.pkl", "rb"))
return
# list(UserLibrary.keys()) ## JDF Test
# +
# create_opportunity() #JDF test
# +
# ## JDF Testing
# save_object(OpportunityLibrary, 'OpportunityLibrary.pkl')
# # list(OpportunityLibrary.keys()) ## JDF Test
# + colab={} colab_type="code" id="Cp5w4FBBH4ce"
## User Details Test
# print(createOpportunityTest.villainType)
# print("Opportunity Details: ", createOpportunityTest.villainType, createOpportunityTest.threatDuration, createOpportunityTest.threatType) # JDF Testing
# -
# ## User Library
# +
## Import User Library
def getUserLibrary():
global UserLibrary
UserLibrary = pickle.load(open("UserLibrary.pkl", "rb"))
return
# list(UserLibrary.keys()) ## JDF Test
# +
# ## UserLibrary = {} ## UserLibrary Reset
# login_prompt() ## JDF Test: Admin. Input additional users.
# + [markdown] colab_type="text" id="Q2AI7V0GH4cz"
# ## Running the Program
# + colab={} colab_type="code" id="S8bnWlG3H4c0"
def run():
getResponseKB()
getUserLibrary()
getOpportunityLibrary()
while True:
print('''Welcome to the Heroism Reasoner.
\nYou can also type 'quit' to end the program at any time.''')
mainMenu()
return(quitProgram())
# run() #JDF Test
# + colab={"base_uri": "https://localhost:8080/", "height": 718} colab_type="code" id="bUqrGz3ZH4c5" outputId="4fff3f3e-7d7c-4691-d9f3-ddaa7a5fbac2"
if __name__ == '__main__':
run()
# -
list(OpportunityLibrary.keys()) ## JDF Test
list(UserLibrary.keys()) ## JDF Test
ResponseKnowledgeBase
# + colab={} colab_type="code" id="lbesvOtZH4dE"
### Future Implementation Notes
## Collect all text descriptions of the issue, record opportunity type with it
## Run KNN to retrieve similar instances
## Spit back instance description and check to see if it sounds right
| Heroism_Reasoner_Alpha_v1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''base'': conda)'
# language: python
# name: python3
# ---
# ## Constants and Geometry
# +
import numpy as np
from matplotlib import pyplot as plt, cm
from heat_equation import *
lx = 5 #length in the x direction
ly = 5 #length in the y direction
nx = 51 #grid points in x direction
ny = 51 #grid points in y direction
tf = 1 #final time to calculate to
dx = lx / (nx - 1)
dy = ly / (ny - 1)
x = np.linspace(0, lx, nx)
y = np.linspace(0, ly, ny)
X, Y = np.meshgrid(x, y)
alpha = 0.1
#nu = 0.5
dt = 0.001 #time step size
ds = int(50) #number of steps after which the state of domain should be saved
nt=int(tf/dt) # number of time steps
# -
# ## Initial and boundary conditions
# +
T = np.random.rand(nx,ny)*300 #initial condition
np.savetxt('T0.csv', T, delimiter=',')
#initial condition
fig = plt.figure(figsize=(11,7),dpi=100)
levels=np.linspace(0,300,50)
plt.contourf(X, Y, T,levels, alpha=0.8, cmap=cm.rainbow)
plt.colorbar()
#plt.contour(X, Y, T,10, colors='black')
ax = plt.gca()
ax.set_aspect(1)
plt.title('Initial condition of the domain')
plt.show()
#Boundary conditions
TLeft=['N',0]
TRight=['N',0]
TTop=['N',0]
TBottom=['N',0]
TBCs=[TLeft,TRight,TTop,TBottom]
#solving the equation for the given geometry
T=heat_equation(T,dx,dy,alpha,dt,ds,nt,TBCs)
clearResults()
# -
# ## Plotting and visualization
# +
fig = plt.figure(figsize=(11,7),dpi=100)
# plotting the temperature field
plt.contourf(X, Y, T, levels, alpha=0.8, cmap=cm.rainbow)
plt.colorbar()
#plt.contour(X, Y, T,10, colors='black')
plt.xlabel('X')
plt.ylabel('Y')
ax = plt.gca()
ax.set_aspect(1)
plt.show()
# -
# ## saving the transient behaviour as an animation
# +
import pandas as pd
import matplotlib.animation as animation
fig, ax = plt.subplots(dpi=100)
def animate(i):
tmpName='T'+str(i*ds)+'.csv'
Tfile=pd.read_csv(r"Results/"+str(tmpName),header=None)
Tarr=Tfile.to_numpy()
# plotting the temperature field
re = plt.contourf(X, Y, Tarr, levels, alpha=0.8, cmap=cm.rainbow)
plt.xlabel('X')
plt.ylabel('Y')
ax = plt.gca()
ax.set_aspect(1)
return re
anim = animation.FuncAnimation(fig, animate, frames=nt//ds, repeat=False)
anim.save(r'Gifs/randomHeated.gif', writer='imagemagick', fps=10)
| randomheated_square.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a href="https://colab.research.google.com/github/NeuromatchAcademy/course-content/blob/master/tutorials/W1D2_ModelingPractice/W1D2_Intro.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] pycharm={"name": "#%% md\n"}
# # Intro
# + [markdown] colab_type="text"
# **Our 2021 Sponsors, including Presenting Sponsor Facebook Reality Labs**
#
# <p align='center'><img src='https://github.com/NeuromatchAcademy/widgets/blob/master/sponsors.png?raw=True'/></p>
# + [markdown] pycharm={"name": "#%% md\n"}
# ## Overview
#
# Yesterday you learned that different models can answer different questions. That means that depending on your question, goals and hypotheses you will need to develop different kinds of models. How to best approach this is the goal of today. We will use this as an opportunity to kick off your group projects at the same time. To do so, we will start walking you through a 10-steps guide of how-to-model. This guide is applicable for both computational modeling and data neuroscience projects and we will discuss similarities and differences between both project types. So today we will start with what you now know determines the choice of modeling or data analysis pipeline you will need to make: how to develop a good question and goal, do the literature review, think about what ingredients you need, and what hypotheses you would like to evaluate.
#
# Today’s tutorial focuses on the first 4 steps of how-to-model by demonstrating the thought process based on a simple phenomenon known as the train illusion. We will first introduce the phenomenon and then provide a step-by-step guide on how to think about and develop the 4 first steps of framing your project. To help you, we will roleplay an example thought process, focussing on typical pitfalls that groups often encounter. Groups will then think about their own projects and develop first answers to each step’s questions. Importantly, this is to get you started in a systematic way with your projects; you will have to revisit those steps later on as your thinking evolves, possibly multiple times. We will also provide similar guidance for the remaining 6 steps of the how-to-model guide that you can work through with your group when you’re reaching that stage of your project. The accompanying answers to each step in our demo project and toy example code for our two different projects (model and data analytics) should help you understand the practical side of the process better.
#
# How to model is rarely, if ever, taught systematically. Our guide is not the only way to approach modeling; but it’s one way to ensure you don’t miss anything important. Going through all the steps also makes publication much easier because you have already explicitly thought about all the elements that you will ultimately need to communicate (see Step 10 later for our examples). Personally, I often take shortcuts in this process and then regret it later… mostly because I forgot to do the one most important thing: be precise about the framing of the project, i.e. the four first steps you will walk through today. Importantly this will set you up to develop any kind of model using any of the tools you will learn about during the remainder of NMA.
# + [markdown] pycharm={"name": "#%% md\n"}
# ## Video
# + cellView="form" pycharm={"name": "#%%\n"}
# @markdown
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id=f"BV1Jk4y1B7cz", width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id=f"65HhPd0kG2k", width=854, height=480, fs=1, rel=0)
print("Video available at https://youtube.com/watch?v=" + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
# + [markdown] pycharm={"name": "#%% md\n"}
# ## Slides
# + cellView="form" pycharm={"name": "#%%\n"}
# @markdown
from IPython.display import IFrame
IFrame(src=f"https://mfr.ca-1.osf.io/render?url=https://osf.io/kmwus/?direct%26mode=render%26action=download%26mode=render", width=854, height=480)
| tutorials/W1D2_ModelingPractice/W1D2_Intro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.9 64-bit (''TrajDetec'': conda)'
# name: python_defaultSpec_1600693225754
# ---
# ## 一,特征列用法概述
# numeric_column 数值列,最常用。
#
# bucketized_column 分桶列,由数值列生成,可以由一个数值列出多个特征,one-hot编码。
#
# categorical_column_with_identity 分类标识列,one-hot编码,相当于分桶列每个桶为1个整数的情况。
#
# categorical_column_with_vocabulary_list 分类词汇列,one-hot编码,由list指定词典。
#
# categorical_column_with_vocabulary_file 分类词汇列,由文件file指定词典。
#
# categorical_column_with_hash_bucket 哈希列,整数或词典较大时采用。
#
# indicator_column 指标列,由Categorical Column生成,one-hot编码
#
# embedding_column 嵌入列,由Categorical Column生成,嵌入矢量分布参数需要学习。嵌入矢量维数建议取类别数量的 4 次方根。
#
# crossed_column 交叉列,可以由除categorical_column_with_hash_bucket的任意分类列构成。
# ## 二,特征列使用范例
# +
import datetime
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
import tensorflow as tf
from tensorflow.keras import layers,models
#打印日志
def printlog(info):
nowtime = datetime.datetime.now().strftime('%Y-%m-%d %H:%M:%S')
print("\n"+"=========="*8 + "%s"%nowtime)
print(info+'...\n\n')
# + tags=[]
printlog("step1: prepare dataset...")
dftrain_raw = pd.read_csv("../../../data/titanic/train.csv")
dftest_raw = pd.read_csv("../../../data/titanic/test.csv")
dfraw = pd.concat([dftrain_raw, dftest_raw])
def prepare_dfdata(dfraw):
dfdata = dfraw.copy()
dfdata.columns = [x.lower() for x in dfdata.columns]
dfdata = dfdata.rename(columns={'survived':'label'})
dfdata = dfdata.drop(['passengerid', 'name'], axis=1)
for col, dtype in dict(dfdata.dtypes).items():
if dfdata[col].hasnans:
dfdata[col + '_nan'] = pd.isna(dfdata[col]).astype('int32')
if dtype not in [np.object, np.str, np.unicode]:
dfdata[col].fillna(dfdata[col].mean(), inplace=True)
else:
dfdata[col].fillna('', inplace=True)
return(dfdata)
dfdata = prepare_dfdata(dfraw)
dftrain = dfdata.iloc[0:len(dftrain_raw),:]
dftest = dfdata.iloc[len(dftrain_raw):,:]
def df_to_dataset(df, shuffle=True, batch_size=32):
dfdata = df.copy()
if 'label' not in dfdata.columns:
ds = tf.data.Dataset.from_tensor_slices(dfdata.to_dict(orient='list'))
else:
labels = dfdata.pop('label')
ds = tf.data.Dataset.from_tensor_slices((dfdata.to_dict(orient = 'list'), labels))
if shuffle:
ds = ds.shuffle(buffer_size=len(dfdata))
ds = ds.batch(batch_size)
return ds
ds_train = df_to_dataset(dftrain)
ds_test = df_to_dataset(dftest)
# + tags=[]
#================================================================================
# 二,定义特征列
#================================================================================
printlog("step2: make feature columns...")
feature_columns = []
# 数值列
for col in ['age','fare','parch','sibsp'] + [
c for c in dfdata.columns if c.endswith('_nan')]:
feature_columns.append(tf.feature_column.numeric_column(col))
tf.print(feature_columns)
# 分桶列
age = tf.feature_column.numeric_column('age')
age_buckets = tf.feature_column.bucketized_column(age,
boundaries=[18, 25, 30, 35, 40, 45, 50, 55, 60, 65])
feature_columns.append(age_buckets)
tf.print(age)
# 类别列
# 注意:所有的Catogorical Column类型最终都要通过indicator_column转换成Dense Column类型才能传入模型!!
sex = tf.feature_column.indicator_column(
tf.feature_column.categorical_column_with_vocabulary_list(
key='sex',vocabulary_list=["male", "female"]))
feature_columns.append(sex)
tf.print(sex)
pclass = tf.feature_column.indicator_column(
tf.feature_column.categorical_column_with_vocabulary_list(
key='pclass',vocabulary_list=[1,2,3]))
feature_columns.append(pclass)
tf.print(pclass)
ticket = tf.feature_column.indicator_column(
tf.feature_column.categorical_column_with_hash_bucket('ticket',3))
feature_columns.append(ticket)
tf.print(ticket)
embarked = tf.feature_column.indicator_column(
tf.feature_column.categorical_column_with_vocabulary_list(
key='embarked',vocabulary_list=['S','C','B']))
feature_columns.append(embarked)
tf.print(embarked)
# 嵌入列
cabin = tf.feature_column.embedding_column(
tf.feature_column.categorical_column_with_hash_bucket('cabin',32),2)
feature_columns.append(cabin)
tf.print(cabin)
# 交叉列
pclass_cate = tf.feature_column.categorical_column_with_vocabulary_list(
key='pclass',vocabulary_list=[1,2,3])
tf.print(pclass_cate)
crossed_feature = tf.feature_column.indicator_column(
tf.feature_column.crossed_column([age_buckets, pclass_cate],hash_bucket_size=15))
feature_columns.append(crossed_feature)
# + tags=[]
#================================================================================
# 三,定义模型
#================================================================================
printlog("step3: define model...")
tf.keras.backend.clear_session()
model = tf.keras.Sequential([
layers.DenseFeatures(feature_columns), #将特征列放入到tf.keras.layers.DenseFeatures中!!!
layers.Dense(64, activation='relu'),
layers.Dense(64, activation='relu'),
layers.Dense(1, activation='sigmoid')
])
# + tags=[]
#================================================================================
# 四,训练模型
#================================================================================
printlog("step4: train model...")
model.compile(optimizer='adam',
loss='binary_crossentropy',
metrics=['accuracy'])
history = model.fit(ds_train,
validation_data=ds_test,
epochs=10)
# + tags=[]
#================================================================================
# 五,评估模型
#================================================================================
printlog("step5: eval model...")
model.summary()
# %matplotlib inline
# %config InlineBackend.figure_format = 'svg'
import matplotlib.pyplot as plt
def plot_metric(history, metric):
train_metrics = history.history[metric]
val_metrics = history.history['val_'+metric]
epochs = range(1, len(train_metrics) + 1)
plt.plot(epochs, train_metrics, 'bo--')
plt.plot(epochs, val_metrics, 'ro-')
plt.title('Training and validation '+ metric)
plt.xlabel("Epochs")
plt.ylabel(metric)
plt.legend(["train_"+metric, 'val_'+metric])
plt.show()
plot_metric(history,"accuracy")
# -
| Computer Science/eat_tensorflow2_in_30_days/practice/5_mid level API/2_feature columns/main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import os
import matplotlib.pyplot as plt
import pandas as pd
from sklearn.preprocessing import StandardScaler, MinMaxScaler
import joypy
import pacmap
import fitsne
import umap
import flowkit as fk
# %matplotlib inline
# -
def make_plots(data, sample_ids, titles, marker, k=10000, seed=123):
"""Reduce scatter and fluroescent channel data to 2D for visualization."""
scaler = StandardScaler()
data_scaled = scaler.fit_transform(data)
umap_reducer = umap.UMAP()
pacmap_reducer = pacmap.PaCMAP()
X_fitsne = fitsne.FItSNE(data_scaled)
X_umap = umap_reducer.fit_transform(data_scaled)
X_pacmap = pacmap_reducer.fit_transform(data_scaled)
min_max_scaler = MinMaxScaler()
X_reduceds = [
min_max_scaler.fit_transform(tmp_x) for tmp_x in [X_fitsne, X_umap, X_pacmap]
]
n = len(X_reduceds)
fig, axes = plt.subplots(n, 3, figsize=(n*3, 9))
for i, (X_reduced, title) in enumerate(zip(X_reduceds, titles)):
for j in range(3):
z = X[marker].iloc[(j*k):(j+1)*k]
ax = axes[j, i]
ax.scatter(
X_reduced[(j*k):(j+1)*k, 0],
X_reduced[(j*k):(j+1)*k, 1],
s=1,
c=z,
cmap='jet'
)
ax.set_xticks([])
ax.set_yticks([])
ax.set_xlim([-0.1,1.1])
ax.set_ylim([-0.1,1.1])
if j==0:
ax.set_title(title, fontsize=14)
if i==0:
ax.set_ylabel('-'.join(sample_ids[j].split('_')[3:5]), fontsize=14)
plt.tight_layout()
base_dir = "data/8_color_data_set"
sample_path = os.path.join(base_dir, "fcs_files")
wsp_path = os.path.join(base_dir, "8_color_ICS.wsp")
session = fk.Session(sample_path)
session.import_flowjo_workspace(wsp_path)
sample_groups = session.get_sample_groups()
sample_groups
sample_group = sample_groups[-1]
print(session.get_gate_hierarchy(sample_group, output='ascii'))
sample_ids = sorted(session.get_group_sample_ids(sample_group))
sample_ids
session.analyze_samples(sample_group)
# ## Compare marker distributions between CD4+ and CD8+
# +
k = 10_000
dfs_cd4 = session.get_wsp_gated_events(sample_group, gate_name='CD4+')
dfs_cd8 = session.get_wsp_gated_events(sample_group, gate_name='CD8+')
X_cd4 = pd.melt(pd.concat([df.iloc[:, 2:-1].sample(k) for df in dfs_cd4]))
X_cd8 = pd.melt(pd.concat([df.iloc[:, 2:-1].sample(k) for df in dfs_cd8]))
X = pd.concat([X_cd4, X_cd8], axis=1)
X = X.iloc[:, [0,1,3]]
X.columns = ['marker', 'CD4+', 'CD8+']
# -
X
fig, ax = joypy.joyplot(
X.groupby('marker', sort=False),
column=['CD4+', 'CD8+'],
color=['red', 'blue'],
legend=True,
alpha=0.5,
linewidth=0.5,
ylim='own',
figsize=(10, 6)
)
# ## Visualize dimension reduction schemes for events in Singlet gate
# +
k = 10_000
dfs = session.get_wsp_gated_events(sample_group, gate_name='Singlets')
X = pd.concat([df.iloc[:, 2:-1].sample(k) for df in dfs])
marker = 'CD4 PE-Cy7 FLR-A'
titles = ['FIt-SNE', 'UMAP', 'PaCMAP']
make_plots(X, sample_ids, titles, marker, k)
# -
# ## Visualize dimension reduction schemes for events in CD3+ gate
# +
k = 10_000
dfs = session.get_wsp_gated_events(sample_group, gate_name='CD3+')
X = pd.concat([df.iloc[:, 2:-1].sample(k) for df in dfs])
marker = 'CD4 PE-Cy7 FLR-A'
titles = ['FIt-SNE', 'UMAP', 'PaCMAP']
make_plots(X, sample_ids, titles, marker, k)
# -
| examples/dimension_reduction_on_gated_populations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# This jupyter notebook contains demo code for:
# - loading a model and using it to drive one or more source frames with audio features
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import torch.nn as nn
import os
import torch
from PIL import Image
from torch.autograd import Variable
from UnwrappedFace import UnwrappedFaceWeightedAverage, BottleneckFromNet
from sklearn.externals import joblib
from torchvision.transforms import Compose, Scale, ToTensor
def load_img_and_audio(file_path):
transform = Compose([Scale((256,256)), ToTensor()])
img = Image.open(file_path).convert('RGB')
img = transform(img)
audio_label_path = str(file_path).replace('audio_faces', 'audio_features').replace('jpg','npz')
audio_feature = torch.Tensor(np.load(audio_label_path)['audio_feat'])
return {'image' : img, 'audio' : audio_feature}
# **Loading models**
# +
# paths to source frames
sourcepaths= ['examples/audio_faces/Retta/1.6/ALELNl9E1Jc/0002725.jpg',
'examples/audio_faces/Maya_Rudolph/1.6/Ylm6PVkbwhs/0004500.jpg',
'examples/audio_faces/Cristin_Milioti/1.6/IblJpk1GDZA/0004575.jpg']
# path to frames corresponding to driving audio features
audio_path = 'examples/audio_faces/Peter_Capaldi/1.6/uAgUjSqIj7U'
imgpaths = os.listdir(audio_path)
# loading models
BASE_MODEL = '/scratch/shared/slow/ow/eccv/2018/release_models/' # Change to your path
model_path = BASE_MODEL + 'x2face_model.pth'
model = UnwrappedFaceWeightedAverage(output_num_channels=2, input_num_channels=3,inner_nc=128)
model.load_state_dict(torch.load(model_path)['state_dict'])
s_dict = torch.load(model_path)
modelfortargetpose = BottleneckFromNet()
state = modelfortargetpose.state_dict()
s_dict = {k: v for k, v in s_dict['state_dict'].items() if k in state.keys()}
state.update(s_dict)
modelfortargetpose.load_state_dict(state)
posemodel = nn.Sequential(nn.Linear(128, 3))
p_dict_pre = torch.load(BASE_MODEL + '/posereg.pth')['state_dict']
posemodel._modules['0'].weight.data = p_dict_pre['posefrombottle.weight'].cpu()
posemodel._modules['0'].bias.data = p_dict_pre['posefrombottle.bias'].cpu()
bottleneckmodel = nn.Sequential(nn.Linear(3, 128, bias=False), nn.BatchNorm1d(128))
b_dict_pre = torch.load(BASE_MODEL + '/posetobottle.pth')['state_dict']
bottleneckmodel.load_state_dict(b_dict_pre)
model = model.cuda()
modelfortargetpose = modelfortargetpose.cuda()
posemodel = posemodel.cuda()
bottleneckmodel = bottleneckmodel.cuda()
model.eval()
modelfortargetpose.eval()
posemodel.eval()
bottleneckmodel.eval()
# load linear regression from audio features to driving vector space
linearregression = joblib.load(BASE_MODEL + '/linearregression_scaledTrue_7000.pkl')
scalar = joblib.load(BASE_MODEL + '/scaler_7000.pkl')
scalar = None
# -
# **Code for driving image generation with audio features**
# Drive 3 different identities with same audio
img_gt_gen = np.empty((0,2560,3))
for sourcepath in sourcepaths:
img_to_show_all = np.empty((256,0,3))
gt_ims = np.empty((256,0,3))
source_data = load_img_and_audio(sourcepath)
source_img = Variable(source_data['image']).cuda().unsqueeze(0)
audio_feature_source = source_data['audio'].cpu().numpy().reshape(1,-1)
audio_feature_origin = linearregression.predict(audio_feature_source)
audio_feature_origin = torch.Tensor(audio_feature_origin).unsqueeze(2).unsqueeze(2)
for imgpath in imgpaths:
# Extract the driving audio features
fullaudiopath = os.path.join(audio_path, imgpath)
audio_data = load_img_and_audio(fullaudiopath)
audio_img = Variable(audio_data['image'], volatile=True).cuda().unsqueeze(0)
audio_feature = audio_data['audio'].cpu().numpy().reshape(1,-1)
if not scalar is None:
audio_feature = scalar.transform(audio_feature)
audio_feature_origin = scalar.transform(audio_feature_origin)
audio_feature = linearregression.predict(audio_feature)
audio_feature = torch.Tensor(audio_feature).unsqueeze(2).unsqueeze(2)
sourcebn = modelfortargetpose(source_img)
sourcepose = posemodel(sourcebn.unsqueeze(0))
sourceposebn = bottleneckmodel(sourcepose)
def update_bottleneck(self, input, output):
newdrive = sourcebn.unsqueeze(0).unsqueeze(2).unsqueeze(3) + Variable(audio_feature).cuda() - Variable(audio_feature_origin).cuda()
audiopose = posemodel(newdrive.squeeze().unsqueeze(0)) #
audioposebn = bottleneckmodel(audiopose)
output[0,:,:,:] = newdrive + sourceposebn.unsqueeze(2).unsqueeze(3) - audioposebn.unsqueeze(2).unsqueeze(3) # if we want to add old pose (of input) and substract pose info that's in the new bottleneck
# Add a forward hook to update the model's bottleneck
handle = model.pix2pixSampler.netG.model.submodule.submodule.submodule.submodule.submodule.submodule.submodule.down[1].register_forward_hook(update_bottleneck)
result = model(source_img, source_img)
handle.remove()
img_to_show_all = np.hstack((result.squeeze().cpu().data.permute(1,2,0).numpy(), img_to_show_all))
if img_gt_gen.shape == (0,2560,3):
gt_ims = np.hstack((audio_img.squeeze().cpu().data.permute(1,2,0).numpy(), gt_ims))
if img_gt_gen.shape == (0,2560,3):
img_gt_gen = np.vstack((img_gt_gen, gt_ims))
img_gt_gen = np.vstack((img_gt_gen, img_to_show_all))
plt.rcParams["figure.figsize"] = [14,14]
plt.imshow(img_gt_gen)
print('Top row: Frames corresponding to driving audio')
print('Bottom 3 rows: generated frames driven with audio features corresponding to top row')
| UnwrapMosaic/Audio2Face.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from task2vec import Task2Vec
from models import get_model
import datasets
import task_similarity
dataset_names = ('stl10', 'mnist', 'cifar10', 'cifar100', 'letters', 'kmnist')
# Change `root` with the directory you want to use to download the datasets
#dataset_list = [datasets.__dict__[name](root='./data')[0] for name in dataset_names]
dataset_list = datasets.__dict__['cifar100'](root='./data')#[0]
embeddings = []
for name, dataset in zip(dataset_names, dataset_list):
print(f"Embedding {name}")
probe_network = get_model('resnet34', pretrained=True, num_classes=int(max(dataset.targets)+1)).cuda()
embeddings.append( Task2Vec(probe_network, max_samples=1000, skip_layers=6).embed(dataset) )
task_similarity.plot_distance_matrix(embeddings, dataset_names)
| small_datasets_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
from math import *
import numpy as np
x = 1
pi=3.1415926
def lag(n):
return (1/sqrt(pi))*(exp(x/2))*(1/(sqrt(sqrt(x*n))))*cos(2*(sqrt(n*x))-pi/4)
print lag(1)
| Untitled20.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _cell_guid="b83d21f3-3afb-4956-a446-963accb6c223"
# Loading libraries
import pandas as pd
import numpy as np
# + _cell_guid="4a3908bb-dd40-4bfa-afb4-3d1a725581e2"
# Global constants and variables
TRAIN_FILENAME = 'train.csv'
TEST_FILENAME = 'test.csv'
# + _cell_guid="a40545bf-6d4d-4702-b63e-2cd7e1451ccc"
train = pd.read_csv('./input/'+TRAIN_FILENAME, parse_dates=['Dates'], index_col=False)
test = pd.read_csv('./input/'+TEST_FILENAME, parse_dates=['Dates'], index_col=False)
# + _cell_guid="951253f6-54d3-4144-b081-da2c4ec56d4d"
train.info()
# + _cell_guid="9df20c4a-7657-4a87-9217-4b475b91afd0"
train = train.drop(['Descript', 'Resolution', 'Address'], axis = 1)
# + _cell_guid="9addbba2-845e-432c-8f5b-ff0a85279fdd"
test = test.drop(['Address'], axis = 1)
# + _cell_guid="3908d2fa-011e-49c6-85c1-b48556c879c8"
def feature_engineering(data):
data['Day'] = data['Dates'].dt.day
data['Month'] = data['Dates'].dt.month
data['Year'] = data['Dates'].dt.year
data['Hour'] = data['Dates'].dt.hour
data['Minute'] = data['Dates'].dt.minute
data['DayOfWeek'] = data['Dates'].dt.dayofweek
data['WeekOfYear'] = data['Dates'].dt.weekofyear
return data
# + _cell_guid="ff2665f5-ac06-436e-a997-608d73fcd9ad"
train = feature_engineering(train)
# + _cell_guid="eb4971bf-ed8e-4876-b17e-673ba0f96bd9"
test = feature_engineering(test)
# + _cell_guid="30e1d1c7-0e8b-4931-a0b9-c69ac2b8b048"
from sklearn.preprocessing import LabelEncoder
# + _cell_guid="58dd6d87-f7f2-456f-a16a-7059ae87d33d"
enc = LabelEncoder()
train['PdDistrict'] = enc.fit_transform(train['PdDistrict'])
# + _cell_guid="78518fa0-416e-4956-9683-7d8ca6891225"
category_encoder = LabelEncoder()
category_encoder.fit(train['Category'])
train['CategoryEncoded'] = category_encoder.transform(train['Category'])
print(category_encoder.classes_)
# + _cell_guid="beae88d5-8b6a-4bb8-bb7a-e35df0a4789d"
enc = LabelEncoder()
test['PdDistrict'] = enc.fit_transform(test['PdDistrict'])
# + _cell_guid="b6ba4df1-6003-485c-a6ed-66c8bea11e81"
print(train.columns)
print(test.columns)
# + _cell_guid="326680b5-89ff-457e-bac3-bd95db768ed8"
x_cols = list(train.columns[2:12].values)
x_cols.remove('Minute')
print(x_cols)
# + _cell_guid="1fbda15b-f192-423c-9f26-90a9e0073574"
from sklearn.ensemble import RandomForestClassifier
clf = RandomForestClassifier(n_estimators = 10)
# + _cell_guid="32a615db-333d-45c4-ac42-902fac00abc1"
clf.fit(train[x_cols], train['CategoryEncoded'])
# + _cell_guid="d6ab201a-9f02-4423-8794-6ac479d77576"
test['predictions'] = clf.predict(test[x_cols])
# + _cell_guid="063828da-2cd8-4cf9-bd2b-922f069a0b88"
def field_to_columns(data, field, new_columns):
for i in range(len(new_columns)):
data[new_columns[i]] = (data[field] == new_columns[i]).astype(int)
return data
# + _cell_guid="6ed883aa-8c34-4826-bc56-2a0471d6d8c1"
test['Category'] = category_encoder.inverse_transform(test['predictions'])
# + _cell_guid="a418820f-8e3a-463a-a1a2-d3009cf37fd4"
categories = list(category_encoder.classes_)
# + _cell_guid="3fb214a5-2ea5-498f-bf57-98d0dc5c399a"
test = field_to_columns(test, 'Category', categories)
# + _cell_guid="fb9cc144-9bf4-42a9-a34e-b9dff6cc5d84"
import time
PREDICTIONS_FILENAME_PREFIX = 'predictions_'
PREDICTIONS_FILENAME = PREDICTIONS_FILENAME_PREFIX + time.strftime('%Y%m%d-%H%M%S') + '.csv'
# + _cell_guid="30f2446f-4dd6-4846-8ed8-6b4c14be7f32"
print(test.columns)
# + _cell_guid="5d7ffd2c-363e-4687-b2e7-68a25ec4d82c"
submission_cols = [test.columns[0]]+list(test.columns[14:])
print(submission_cols)
# + _cell_guid="66b04c27-1b90-46b7-8a42-434c02e511c0"
print(PREDICTIONS_FILENAME)
test[submission_cols].to_csv(PREDICTIONS_FILENAME, index = False)
| script.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Applications
#
# We can solidify our understanding of probability, conditional probability and Bayes Rule by going over some problems, some of which are quite famous. We'll start first with some general probability problems and then move in the second part to problems using Bayes Rule.
#
# Most of these problems come from <NAME>ey's excellent book [Think Bayes](http://greenteapress.com/wp/think-bayes/).
#
# Some of these problems are just calculations, either by hand or by computer. Others involve answering questions with simulations in order to calculate probabilities. You may be called upon to do something similar as a data scientist. For example, you have just fielded an advertising campaign in 20 major cities. Testing showed that advertising campaign was 10 percent better than the previous one. However, the last three weeks of returns in New York have been below average. What is the probability of this happening given that the campaign really is 10 percent better?
# ## Applications in General Probability
#
# These problems are general probability problems (although the Monty Hall problem can be solved using Bayes Rule).
# ### A Girl Named Florida
#
# Consider the following problems: we have a family with two children.
#
# * What is the probability that they are both girls?
#
# To answer this question, we have have to make some assumptions about the probability of a child being either a boy or girl (which we will take to mean either XX or XY chromosomes). The generally accepted probabilities are P(boy) = 0.5 and P(girl) = 0.5 (ignoring other chromosomal possibilities).
#
# Recall our definition of independence. Two sets of events, $A$ and $B$, are independent if the following holds true:
#
# $$P(A, B) = P(A)P(B)$$
#
# but that's how we determine that A and B are independent. If we *assume* that the events are independent, then we can turn it around to calculate the probability of the joint event:
#
# $$P(A)P(B) = P(A, B)$$
#
# Here $A$ is the "sex of the first child" and $B$ is the "sex of the second child". This means we can take P(A=girl) as 0.5 and P(B=girl) as 0.5--shortened to P(girl) x P(girl)-- which equals 0.5 x 0.5 = 0.25.
#
# Note that if we wanted to calculate the probability of them being different sexes, then we'd have to calculate the probability of having a boy then a girl (0.5 x 0.5 = 0.25) and the probability of having a girl then a boy (0.5 x 0.5 = 0.25) and combine them based on the Additive Law of Probability: 0.25 + 0.25 = 0.50.
#
# But remember how I said probability is just counting?
#
# There are 4 possibilities:
#
# 1. Boy, Girl.
# 2. Girl, Boy.
# 3. Girl, Girl.
# 4. Boy, Boy.
#
# There is only one way in which both children are girls so the probability of two girls is 1/4 = 0.25. There are two ways in which the children are mixed sexes so the probability of that joint event is 2/4 = 1/2 = 0.5.
#
#
# * What is the probability that they are both girls given that at least one is a girl?
#
# Now there are only 3 possibilities--the ones that include a Girl as either the first or the second birth:
#
# 1. Boy, Girl.
# 2. Girl, Boy.
# 3. Girl, Girl.
#
# Using the counting method, we can see that there is only one way to get the result we're interested in and three possible outcomes so the probability is 1/3.
#
# Using the mathy way, we know from above that probability of the individual outcomes are each 1/4 or 0.25:
#
# 1. Boy, Girl = 1/4
# 2. Girl, Boy. = 1/4
# 3. Girl, Girl. = 1/4
#
# However, since we have ruled out the {Boy, Boy} possibility by assumption, we have to renormalize the probabilities. Normalization just means "make all the probabilities add up to 1 again" and you do this by adding the probabilities together (which is 3/4) and dividing each original probability by this *normalizer*:
#
# 1. Boy, Girl = 1/4 // 3/4 = 1/3
# 2. Girl, Boy. = 1/4 // 3/4 = 1/3
# 3. Girl, Girl. = 1/4 // 3/4 = 1/3
#
# And we get 1/3 as before. The reason we show both ways to get the answer is because, as you might expect, there are cases--most cases--where the counting approach isn't tractable.
#
# * What is the probability that they are both girls given that the oldest (first) is a girl?
#
# We do the same thing again except that any outcome that has a Boy as the oldest is removed:
#
# 1. Girl, Boy. = 1/4
# 2. Girl, Girl. = 1/4
#
# And again, we need to have our probabilities add up to 1 so we normalize them:
#
# 1. Girl, Boy. = 1/4 // 2/4 = 1/2
# 2. Girl, Girl. = 1/4 // 2/4 = 1/2
#
# If you think about it a second, this makes perfect sense...the events are independent so knowing that the first is a girl doesn't give us any information about the second child's sex.
#
# That was a fairly typical probability problem. There is a crazy variant that asks:
#
# * What is the probability that they are both girls given that one of them is a girl named Florida?
#
# Think about it. Does the name change anything?
#
# We're now going to switch to problems where simulation is often a useful tool. If you ever have a probability problem that you can't quite formulate right or if someone doesn't believe your answer, think: can I simulate this?
# ### Birthday Problem
#
# The Birthday Problem is as follows: what is the probability that two people in a given group of size $N$, have the same birthday (month and day)?
#
# 1. Guess. What do you think the probability is? 10%, 20%, 30%...100%?
# 2. Think about how you might answer this mathematically.
# 3. Think about how you might solve this easily as a simulation. What assumptions do you need to make?
#
# We're going to simulate the problem by writing a few functions. The first function takes $k$ persons as an argument and assigns them randomly to one of the 365 days of the year (we ignore leap years). As we do so, we count how many people have that birthday.
#
# First some imports...
from random import randint, uniform
from collections import defaultdict
# We used `defaultdict` because missing keys are automatically assigned a value of `0` instead of it causing a KeyError.
def tally_k_birthdays( k):
counts = defaultdict( int)
for i in range( 0, k):
birthday = randint( 1, 365)
counts[ birthday] += 1
return counts
# Let's see what we get for 10 people:
tally_k_birthdays( 10)
# Now all we need to do is take this dictionary of values and see if any of the days (we only need one) has a count greater than one which would mean that two (or more) people have the same birthday:
def identify_same_birthdays( counts_of_birthdays):
for day in counts_of_birthdays.keys():
if counts_of_birthdays[ day] > 1:
return True
return False
# In general, in order to get a good result from a simulation, it must be run multiple times and the results averaged. We write a function to do just that. The arguments are $N$ people and $times$ simulations.
def sample_group( N, times):
match = 0.0
for i in range( times):
birthday_count = tally_k_birthdays( N)
if identify_same_birthdays( birthday_count):
match += 1.0
return match / times
# We can now run the function and see approximately what the probability is for two people to have the same birthday in a class with $N=26$ students:
sample_group( 26, 10000)
# It's much more probable than people usually think.
# This is a good example of a simple simulation for a system process. Again, in theory, everything is fairly deterministic. Parents decided to have children, the children were born on certain days, the children grew up and where in a particular class (one such situation) or they got older and went to university (another situation) or took up an interest in art and when to an art gallery (another such situation) and in all cases the simulation works.
#
# It doesn't work if an assumption if violated. If the situation is a Meetup for People born in March, we would need an entirely different situation.
#
# 1. Can you reprogram the simulation to see how many people it takes to have a 50% probability of someone with the same birthday, if everyone is born in the same month?
# ### Monty Hall Problem
#
# Monty Hall was the host for *Let's Make a Deal* before <NAME>. One of the "bits" on the show involved picking a curtain in hopes of winning a great prize like a car and this probability problem is based on it. It's actually a very famous problem.
#
# There are three curtains: 1, 2, and 3. Behind one of those curtains is a car. On the show, the other curtains often had gag gifts behind them like a goat but we assume they're empty. The contestant picks the curtain they believe hides the car. After picking, Monty reveals what is behind one of the other curtains. One important assumption is that if the contestant *has* picked the car, Monty reveals one of the other two curtains at random.
#
# The contestant is then given the option to either stick with the curtain they picked or switch to the remaining curtain. The question is this: should the contestant switch? What do you think?
#
# There are a number of ways to answer this question but we're going to use simulation because that's often the most definitive. In fact, <NAME>, the famous mathematician, would not believe the correct answer until it was simulated.
#
# First, we have a function that simulates one Monty Hall "Problem". It basically says:
#
# 1. set up the problem
# 2. place the car at random.
# 3. generate a random contestant pick.
# 4. figure out which curtain to reveal.
# 5. figure out which curtain is closed.
# 6. if do_switch is True, make the pick equal to the closed curtain. Otherwise, keep it the same.
# 7. return if the picked curtain equals the car's curtain (True or False).
def evaluate_a_monty_hall_scenario( do_switch=False):
options = {1, 2, 3}
car = randint( 1, 3)
pick = randint( 1, 3)
opened = list( options.difference( {car}).difference( {pick}))[0]
closed = list( options.difference( {pick}).difference( {opened}))[0]
if do_switch:
pick = closed
return car == pick
# Let's run it 10 times:
for i in range( 0, 10):
print( evaluate_a_monty_hall_scenario(True))
# We're now going to run the Monty Hall problem function 10,000 times and evaluate what happens first, if you don't switch and second, if you switch:
def evaluate_monty_hall_problem( switch=False):
trials = 10000
count = 0
for i in range( 0, trials):
result = evaluate_a_monty_hall_scenario( switch)
if result:
count += 1
return float( count) / trials
evaluate_monty_hall_problem()
evaluate_monty_hall_problem(True)
# And there you have it, if you switch, you win the car 66% of the time.
# ## Applications of Bayes Rule
#
# Speaking of switching, one of the main types of problems we'll be solving are problems involving Bayes Rule. In fact, Bayesian Inference depends entirely on understanding Bayes Rule and evaluating it for a large number of possibilities. We'll start out with smaller problems.
#
# For whatever reason, Bayes Rule examples are either weather or medical tests. We'll start with the weather:
# ### Rain or Shine
#
# Sam is getting married tomorrow in an outdoor ceremony in the desert. In recent years, it has only rained 5 days per year. Unfortunately, the meteorologist has predicted rain for tomorrow. Should Sam rent a tent for the ceremony?
#
# We can solve this problem using Bayes Rule which remember is:
#
# $$P(A|B) = \frac{P(B|A)P(A)}{P(B)}$$
#
# But instead what we want is:
#
# $$P(W|F) = \frac{P(F|W)P(W)}{P(F)}$$
#
# where $W$ is weather (rain or shine) and $F$ is forecast (rain or shine). Remember that $P(W)$ in the numerator is our *prior* probability. What *is* our prior probability? Well, it only rains 5 days a year on average:
#
# | rain | shine |
# |:----:|:-----:|
# | 5/365 = 0.0137 | 360/365 = 0.9863 |
#
# I think this is what Sam had in mind when he planned his wedding.
#
# But now he needs to take new evidence into account: a forecast of rain. The likelihood $P(F|W)$ is essentially the probability of the meteorologist being correct: given that it rained, what is the probability that it was forecast? Sam looks this up on the Internet.
#
# | F | rain | shine |
# |:---:|:----:|:-----:|
# |rain | 0.8 | 0.2 |
# |shine | 0.2 | 0.8 |
#
# What does this mean? *Given* that it rained, there is an 80% chance there was a forecast of rain:
#
# $F(F=rain|W=rain) = 0.8$
#
# Because it *will* be confusing, we do not take shortcuts here. We will use the longhand notation, F=rain and W=rain, to distinguish the two events. Up above, we had Bayes Rule defined over entire random variables.
#
# What does it look like for the specific outcome we're interested in?
#
# $$P(W=rain|F=rain) = \frac{P(F=rain|W=rain)P(W=rain)}{P(F=rain)}$$
#
# We have everything we need except the denominator. We can use total probability for it, though:
#
# $P(F=rain) = P(F=rain|W=rain)P(W=rain) + P(F=rain|W=shine)P(W=shine)$
#
# $0.8 \times 0.0137 + 0.2 \times 0.9863 = 0.208$
#
# and now we have:
#
# $P(W=rain|F=rain) = \frac{0.8 \times 0.0137}{0.208} = 0.053$
#
# So really, Sam should just go ahead with the wedding (at least from a weather perspective).
# ### Breast Cancer
#
# The logic underlying this problem is why certain routine screenings for breast cancer were discontinued. The numbers, however, are made up.
#
# 1% of women at age 40 who participate in routine screening have breast cancer. 80% of women with breast cancer will get positive mammographies. 9.6% of women without breast cancer will also get positive mammographies. A woman in the age group had a positive mammography. What is the probability of her having breast cancer?
#
# We have two variables, each with two outcomes: $M$ is {pos, neg}, and $C$ is {yes, no}. As before, we need to set up Bayes Rule and determine either what information we have and what information we need to calculate.
#
# $$P(yes|pos) = \frac{P(pos|yes)P(yes)}{P(pos)}$$
#
# We have the prior, $P(yes)$ which is simply 0.01. We have the likelihood we need which is established in the second sentence: $P(pos|yes)$ = 0.8 (which means that $P(neg|yes)$ = 0.2. We don't have $P(pos)$. We will need to use total probability again.
#
# $P(pos) = P(pos|yes)P(yes) + P(pos|no)P(no)$
#
# We have $P(pos|no)$ from the 3rd sentence: 0.096. Note that this clearly shows where total probability comes from. If we want to calculate the probability of a positive test result, we need to take into account all the possible sources of positive test results. These come from those with cancer who get a positive test result (the first term) and those without cancer who get a positive test result (the second term). The probability of not having cancer is just 1 - P(yes).
#
# $P(pos) = 0.8 \times 0.01 + 0.096 \times 0.99 = 0.103$
#
# and now we can just plug in the numbers.
#
# $P(yes|pos) = \frac{0.8 * 0.01}{0.103} = 0.078$
#
# This result makes an important assumption, though, the only information about this woman's status is that this was a routine screening. Why might this not be the case?
#
# OK, we're computer scientists...enough math. We can let computers do the math.
# ### Elvis
#
# Apparently Elvis was one of a set of twins. He had a twin brother who died at birth. We want to know the probability that Elvis had an identical twin. This isn't really enough information to answer anything so...
#
# Wikipedia to the rescue..."Twins are estimated to be approximately 1.9% of the world population, with monozygotic twins making up 0.2% of the total, 8% of all twins".
#
# You should solve this by hand right now, writing out the problem. It might surprise you how difficult it is to get started. Consider the following...what is the event we want to know about and what is the evidence?
#
# So the evidence is that the child was male and the event we're trying to determine the probability of is that Elvis and the child were identical twins:
#
# $$P(I|M) = \frac{P(M|I)P(I)}{P(M)}$$
#
# I'm going to start out with a helper function that normalizes a probability distribution the way I have decided to represent it (as a map):
def normalize( dist):
normalizer = sum( dist.values())
for k in dist.keys():
dist[ k] = dist[ k] / normalizer
return dist # don't need to do this.
# I'm describing the events as **I**dentical twin or **F**raternal twin. The probabilities come from the Wikipedia article. In Python, it is very convenient to represent a discrete Probability distribution with a Dict where the keys are outcomes {"I", "F"} and the values are the probabilities of those outcomes.
elvis_prior = {"I": 0.08, "F": 0.92}
# Here we use a Dict to express a likelihood which ends up as a nested Dict. Remember that $P(A|B)$ is a Probability distribution for each value of "B". In this case, the outer key is the "given" so that we can say "given I" and look up the appropriate probability distribution. The inner Dict represents the probability distribution over the events of "A", in this case the sex of the baby, **M**ale or **F**emale.
elvis_likelihoods = {
"I": { "M": 1.00, "F": 0.00},
"F": { "M": 0.50, "F": 0.50}
}
# Below is a function that will calculate the posterior probability for the entire probability distribution (over all events). As we've mentioned before, in Bayes Rule:
#
# $P(A|B) = \frac{P(B|A)P(A)}{P(B)}$
#
# we are calculating an entire posterior probability *distribution*...a probability for each value of A given each value of B. Additionally, it is unlikely that we know the value of the normalizer $P(B)$ directly. However, we can calculate $P(B)$ using the Rule of Total Probability:
#
# $P(B) = P(B|A=a_1)P(A=a_1) + P(B|A=a_2)P(A=a_2) + ... + $P(B|A=a_n)P(a_n)$
#
# but it turns out that if we are interested in the probability of every hypothesis in A, we are going to calculate all of these values anyway. We don't need to go through any extra effort. First we note that if we are only concerned about *order* we do not need to normalize so we have:
#
# $P(A=a_1|B) \propto P(B|A=a_1)P(A=a_1)$
#
# $P(A=a_2|B) \propto P(B|A=a_2)P(A=a_2)$
#
# $P(A=a_n|B) \propto P(B|A=a_n)P(A=a_n)$
#
# where $\propto$ means "proportional to". We can calculate all of these without calculating the normalizer, $P(B)$. But having calculated all those terms, we have calculated the terms we need to compute the normalizer and calculate the actual probabilities:
#
# $P(A=a_1|B) = \frac{P(B|A=a_1)P(A=a_1)}{P(B)}$
#
# $P(A=a_2|B) = \frac{P(B|A=a_2)P(A=a_2)}{P(B)}$
#
# $P(A=a_n|B) = \frac{P(B|A=a_n)P(A=a_n)}{P(B)}$
#
# This is what the following function does, although for all values of A and B.
def query( prior, likelihoods, evidence):
posterior = {}
for k in prior.keys():
posterior[ k] = likelihoods[ k][ evidence] * prior[ k]
normalize( posterior)
return posterior
# Now we can print out the prior probability and the posterior probability:
print( "prior=", elvis_prior)
print( "posterior=", query( elvis_prior, elvis_likelihoods, "M"))
# The evidence (that the other child was a boy), increases the probability that they were identical twins (if the other child had been female, it would have been impossible).
#
# What other piece of evidence is implicit in this calculation?
# ### M & M's
#
# Here is a bit more challenging problem.
#
# A friend shows me two bags of M&M's and tells me that one is from 1994 and the other is from 1996. He won't tell me which is which but gives me an M&M from each bag. One is yellow and one is green. What is the probability that the yellow M&M is from the 1994 bag?
#
# So the first step is map out the events we're trying to predict and the evidence. I'll use the same basic approach as before, representing probability distributions as Dicts.
#
# The key information, however, is that the blue M&M was introduced in 1995. Before that the color mixes in the bags where:
#
# | color | 1994 | 1996 |
# |:-----:|:----:|:----:|
# | brown | 30% | 13% |
# | yellow | 20% | 14% |
# | red | 20% | 13% |
# | green | 10% | 20% |
# | orange | 10% | 16% |
# | tan | 10% | 0% |
# | blue | 0% | 24% |
#
# (I'm not sure where this data came from!)
#
# You should try to solve this for yourself before looking at my solution.
# Here is the prior distribution for the 1994 bag:
mix94 = dict(brown=0.3, yellow=0.2, red=0.2, green=0.1, orange=0.1, tan=0.1)
mix94
# and the prior distribution for the 1996 bag:
mix96 = dict(blue=0.24, green=0.2, orange=0.16, yellow=0.14, red=0.13, brown=0.13)
mix96
# Now, my two possible events are: either the first bag is the 1994 bag (A) or the first bag is the 1996 bag (B):
A = dict(bag1=mix94, bag2=mix96)
B = dict(bag1=mix96, bag2=mix94)
# which gives me my likelihoods:
m_m_likelihoods = {"A": A, "B": B}
m_m_likelihoods
# This is a more complex likelihood than we're used to seeing.
#
# Given that event A happened (1994 bag), then the probability of picking a yellow M&M from that bag is 20%. Given that event B happened (1996 bag), then the probability of picking a yellow M&M out of that bag is 14%.
#
# Our prior is 50/50 for each of the events A and B because there are two bags.
m_m_priors = {"A": 0.5, "B": 0.5}
# Our evidence is that I took a yellow M&M out of Bag 1 and a green M&M out of Bag 2:
m_m_evidences = [('bag1', 'yellow'), ('bag2', 'green')]
# And now some code to massage it all together:
# +
from copy import deepcopy
def calculate_m_m_posteriors( priors, likelihoods, evidences):
posteriors = {}
current_priors = deepcopy( priors)
for evidence in evidences:
bag, mnm = evidence
for hypothesis in priors.keys():
posteriors[ hypothesis] = likelihoods[ hypothesis][ bag][ mnm] \
* current_priors[ hypothesis]
normalize( posteriors)
current_priors = posteriors
print( "evidence=", evidence, "posterior=", posteriors)
return posteriors
# -
print( "prior", m_m_priors)
calculate_m_m_posteriors( m_m_priors, m_m_likelihoods, m_m_evidences)
# One special thing to note about Bayes Rule is that it doesn't matter if you take the evidence altogether or piece by piece (no pun intended). You will always get the same result. It's slightly easier in this case to cycle through the evidence and use the posterior distribution that results as the *prior* distribution for the next calculation.
#
# This is the beauty of Bayes Rule (and makes it slightly easier to program).
#
# It is always worth noting that in all cases we used probability to deal with systems--processes--exhibiting uncertainty whether it was a breast cancer testing process, the weather, Elvis's deceased twin or M&M's.
# 1. Can you solve the Monty Hall problem using Bayes Rule?
# 2. Can you identify--even in the most general terms--the processes underlying each of these problems?
| fundamentals_2018.9/probability/applications.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/jihoover77/DS-Unit-2-Linear-Models/blob/master/Linear_Regression2_212_assignment.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="H-1FiXt-gFW8"
# Lambda School Data Science
#
# *Unit 2, Sprint 1, Module 2*
#
# ---
# + [markdown] id="7IXUfiQ2UKj6"
# # Regression 2
#
# ## Assignment
#
# You'll continue to **predict how much it costs to rent an apartment in NYC,** using the dataset from renthop.com.
#
# - [ ] Do train/test split. Use data from April & May 2016 to train. Use data from June 2016 to test.
# - [ ] Engineer at least two new features. (See below for explanation & ideas.)
# - [ ] Fit a linear regression model with at least two features.
# - [ ] Get the model's coefficients and intercept.
# - [ ] Get regression metrics RMSE, MAE, and $R^2$, for both the train and test data.
# - [ ] What's the best test MAE you can get? Share your score and features used with your cohort on Slack!
# - [ ] As always, commit your notebook to your fork of the GitHub repo.
#
#
# #### [Feature Engineering](https://en.wikipedia.org/wiki/Feature_engineering)
#
# > "Some machine learning projects succeed and some fail. What makes the difference? Easily the most important factor is the features used." — <NAME>, ["A Few Useful Things to Know about Machine Learning"](https://homes.cs.washington.edu/~pedrod/papers/cacm12.pdf)
#
# > "Coming up with features is difficult, time-consuming, requires expert knowledge. 'Applied machine learning' is basically feature engineering." — <NAME>, [Machine Learning and AI via Brain simulations](https://forum.stanford.edu/events/2011/2011slides/plenary/2011plenaryNg.pdf)
#
# > Feature engineering is the process of using domain knowledge of the data to create features that make machine learning algorithms work.
#
# #### Feature Ideas
# - Does the apartment have a description?
# - How long is the description?
# - How many total perks does each apartment have?
# - Are cats _or_ dogs allowed?
# - Are cats _and_ dogs allowed?
# - Total number of rooms (beds + baths)
# - Ratio of beds to baths
# - What's the neighborhood, based on address or latitude & longitude?
#
# ## Stretch Goals
# - [ ] If you want more math, skim [_An Introduction to Statistical Learning_](http://faculty.marshall.usc.edu/gareth-james/ISL/ISLR%20Seventh%20Printing.pdf), Chapter 3.1, Simple Linear Regression, & Chapter 3.2, Multiple Linear Regression
# - [ ] If you want more introduction, watch [<NAME>, Statistics 101: Simple Linear Regression](https://www.youtube.com/watch?v=ZkjP5RJLQF4)
# (20 minutes, over 1 million views)
# - [ ] Add your own stretch goal(s) !
# + id="o9eSnDYhUGD7"
# %%capture
import sys
# If you're on Colab:
if 'google.colab' in sys.modules:
DATA_PATH = 'https://raw.githubusercontent.com/LambdaSchool/DS-Unit-2-Applied-Modeling/master/data/'
# !pip install category_encoders==2.*
# If you're working locally:
else:
DATA_PATH = '../data/'
# Ignore this Numpy warning when using Plotly Express:
# FutureWarning: Method .ptp is deprecated and will be removed in a future version. Use numpy.ptp instead.
import warnings
warnings.filterwarnings(action='ignore', category=FutureWarning, module='numpy')
# + [markdown] id="hV6kvl8O2cNC"
# # Step 1: Read in, wrangle and feature engineer the dataset
# + id="cvrw-T3bZOuW"
# Import libraries
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.linear_model import LinearRegression
from sklearn.metrics import mean_absolute_error, mean_squared_error, r2_score
def wrangle(filepath):
# Read New York City apartment rental listing data
df = pd.read_csv(filepath,
parse_dates=['created'],
index_col='created')
# Remove the most extreme 1% prices,
# the most extreme .1% latitudes, &
# the most extreme .1% longitudes
df = df[(df['price'] >= np.percentile(df['price'], 0.5)) &
(df['price'] <= np.percentile(df['price'], 99.5)) &
(df['latitude'] >= np.percentile(df['latitude'], 0.05)) &
(df['latitude'] < np.percentile(df['latitude'], 99.95)) &
(df['longitude'] >= np.percentile(df['longitude'], 0.05)) &
(df['longitude'] <= np.percentile(df['longitude'], 99.95))]
# Drop duplicate column
df.drop(columns='street_address', inplace=True)
# Drop high cardinality features
threshold = 3
drop_cols = [col for col in df.select_dtypes('object').columns
if df[col].nunique() > threshold]
df.drop(columns=drop_cols, inplace=True)
# Engineer a feature called perks_count
df['perks_count'] = df[['elevator', 'cats_allowed',
'hardwood_floors', 'dogs_allowed', 'doorman',
'dishwasher', 'no_fee',
'laundry_in_building', 'fitness_center', 'pre-war',
'laundry_in_unit',
'roof_deck', 'outdoor_space', 'dining_room',
'high_speed_internet',
'balcony', 'swimming_pool', 'new_construction',
'terrace', 'exclusive',
'loft', 'garden_patio', 'wheelchair_access',
'common_outdoor_space']].sum(axis=1)
# Engineer a feature called total_rooms
df['total_rooms'] = df['bathrooms'] + df['bedrooms']
# Engineer a column that checks for cats or dogs
df.loc[(df['cats_allowed'] == 1) | (df['dogs_allowed'] == 1), 'cat_or_dogs'] = 1
df.loc[(df['cats_allowed'] != 1) & (df['dogs_allowed'] != 1), 'cat_or_dogs'] = 0
# Engineer a column that checks for cats and dogs
df.loc[(df['cats_allowed'] == 1) & (df['dogs_allowed'] == 1), 'cat_and_dogs'] = 1
df.loc[(df['cats_allowed'] != 1) | (df['dogs_allowed'] != 1), 'cat_and_dogs'] = 0
# Create a feature column from latitude and longitude to get the distance from
# Manhattan
manhattan_lat = 40.7829
manhattan_lon = -73.9711
df['distance_from_Manhattan'] = np.sqrt((manhattan_lat - df['latitude'])**2 +
(manhattan_lon - df['longitude'])**2)
return df
df = wrangle(DATA_PATH + 'apartments/renthop-nyc.csv')
# + colab={"base_uri": "https://localhost:8080/", "height": 431} id="vVW2LoNgnDNX" outputId="8fb36527-e0f9-45a7-8426-3f4559676a22"
df.head()
# + [markdown] id="OuDExDDV2zL7"
# # Step 2: Split Data
# + colab={"base_uri": "https://localhost:8080/"} id="z6lf0yUArdys" outputId="3d70841f-510b-4dd7-e7c1-ba92cbc679c2"
df.columns
# + colab={"base_uri": "https://localhost:8080/"} id="qeMgiNrlsJYQ" outputId="ea201a4d-9745-4ef4-b53b-036fce21fe8b"
target = 'price'
features = ['total_rooms', 'perks_count', 'distance_from_Manhattan']
X = df[features]
y = df[target]
X.shape, y.shape
# + colab={"base_uri": "https://localhost:8080/"} id="rvePQqUH1bGH" outputId="3c9d7b91-67fc-437f-c643-e71bef5ee830"
cutoff = '2016-06-01'
mask = df.index < cutoff
X_train, y_train = X.loc[mask], y.loc[mask]
X_test, y_test = X.loc[~mask], y.loc[~mask]
X_train.shape, y_train.shape, X_test.shape, y_test.shape
# + [markdown] id="MTCjtL7X4k00"
# # Step 3: Establish Baseline
# + colab={"base_uri": "https://localhost:8080/"} id="FgAyKLpJ3VTR" outputId="7c6f879d-8288-40bf-c091-59223675e1c3"
y_pred = [y_train.mean()] * len(y_train)
print(f'Baseline MAE: ${mean_absolute_error(y_train, y_pred):,.0f}')
# + [markdown] id="PtVZEpHU5IH9"
# # Step 4: Build Model
# + colab={"base_uri": "https://localhost:8080/"} id="M-_2SyIN4vz5" outputId="ae256b14-8f91-426d-fbfb-4565addc4713"
# Step 1: Import predictor class
# Done above
# Step 2: Instantiate the class
model = LinearRegression()
# Step 3: Fit the model to training set
model.fit(X_train, y_train)
# + [markdown] id="OPuuPpAQ6_yX"
# # Step 5: Check Metrics
# + colab={"base_uri": "https://localhost:8080/"} id="t0QQg7j85csg" outputId="e1459604-5fec-4371-f7e0-6462c5be2a5d"
y_pred = model.predict(X_test)
print(f'Linear Model Training MAE: ${mean_absolute_error(y_train, model.predict(X_train)):,.0f}')
print(f'Linear Model Testing MAE: ${mean_absolute_error(y_test, model.predict(X_test)):,.0f}')
print(f'Linear Model Training RMSE: ${mean_squared_error(y_train, model.predict(X_train), squared=False):,.0f}')
print(f'Linear Model Testing RMSE: ${mean_squared_error(y_test, model.predict(X_test), squared=False):,.0f}')
print('Linear Model Training R^2: ', model.score(X_train, y_train))
print('Linear Model Testing R^2: ', model.score(X_test, y_test))
# + [markdown] id="OMWPwHTX9Mau"
# # Step 6: Communicate the Results
# + colab={"base_uri": "https://localhost:8080/"} id="8-7m7rSN9KWQ" outputId="d09195ea-79f0-49b1-ef6f-270ba2dec353"
intercept = round(model.intercept_)
coef_1 = model.coef_[0]
coef_2 = model.coef_[1]
coef_3 = model.coef_[2]
print(f'PRICE = {intercept:,.0f} + ({coef_1:,.0f} * ROOMS) + ({coef_2:,.0f} * PERKS) - ({-coef_3:,.0f} * DISTANCE)')
# + [markdown] id="OrTu48Lf-ZGN"
# ### I added three new features to my dataset. The best score I received was from the mean absolute error $853.
# + id="B1fJ6NUHviXG"
| Linear_Regression2_212_assignment.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# **Notas para contenedor de docker:**
# Comando de docker para ejecución de la nota de forma local:
#
# nota: cambiar `dir_montar` por la ruta de directorio que se desea mapear a `/datos` dentro del contenedor de docker.
#
# ```
# dir_montar=<ruta completa de mi máquina a mi directorio>#aquí colocar la ruta al directorio a montar, por ejemplo:
# #dir_montar=/Users/erick/midirectorio.
# ```
#
# Ejecutar:
#
# ```
# $docker run --rm -v $dir_montar:/datos --name jupyterlab_prope_r_kernel_tidyverse -p 8888:8888 -d palmoreck/jupyterlab_prope_r_kernel_tidyverse:2.1.4
#
# ```
# Ir a `localhost:8888` y escribir el password para jupyterlab: `<PASSWORD>`
#
# Detener el contenedor de docker:
#
# ```
# docker stop jupyterlab_prope_r_kernel_tidyverse
# ```
#
# Documentación de la imagen de docker `palmoreck/jupyterlab_prope_r_kernel_tidyverse:2.1.4` en [liga](https://github.com/palmoreck/dockerfiles/tree/master/jupyterlab/prope_r_kernel_tidyverse).
# ---
# Para ejecución de la nota usar:
#
# [docker](https://www.docker.com/) (instalación de forma **local** con [Get docker](https://docs.docker.com/install/)) y ejecutar comandos que están al inicio de la nota de forma **local**.
#
# O bien dar click en alguno de los botones siguientes:
# [](https://mybinder.org/v2/gh/palmoreck/dockerfiles-for-binder/jupyterlab_prope_r_kernel_tidyerse?urlpath=lab/tree/Propedeutico/R/clases/2_probabilidad/1_elementos_de_probabilidad.ipynb) esta opción crea una máquina individual en un servidor de Google, clona el repositorio y permite la ejecución de los notebooks de jupyter.
# [](https://repl.it/languages/Rlang) esta opción no clona el repositorio, no ejecuta los notebooks de jupyter pero permite ejecución de instrucciones de Python de forma colaborativa con [repl.it](https://repl.it/). Al dar click se crearán nuevos ***repl*** debajo de sus users de ***repl.it***.
#
# ### Lo siguiente está basado los libros:
#
# ### * <NAME>, Pensando Antes de Actuar: Fundamentos de Elección Racional, 2009.
#
# ### * <NAME>, Introduction to Probability and Statistics Using R, 2014.
# ### El libro de <NAME> tiene github: [jkerns/IPSUR](https://github.com/gjkerns/IPSUR)
# **Notas:**
#
# * Se utilizará el paquete [prob](https://cran.r-project.org/web/packages/prob/index.html) de R para los experimentos descritos en la nota y aunque con funciones nativas de R se pueden crear los experimentos, se le da preferencia a mostrar cómo en R se tienen paquetes para muchas aplicaciones.
#
# * En algunas líneas no es necesario colocar `print` y sólo se ha realizado para mostrar los resultados de las funciones en un formato similar al de R pues la nota se escribió con jupyterlab y R.
#
# * Cuidado al utilizar las funciones del paquete `prob` para construir espacios de probabilidad grandes como lanzar un dado 9 veces... (tal experimento tiene 10 millones de posibles resultados)
options(repr.plot.width=4, repr.plot.height=4) #esta línea sólo se ejecuta para jupyterlab con R
library(prob)
# # Tipos de experimentos
# Se tienen dos tipos: **determinísticos y aleatorios**. Un **experimento determinístico** es aquel cuyo resultado puede ser predicho con seguridad antes de realizarlo, por ejemplo: combinar hidrógeno y oxígeno o sumar $2+3$. Un **experimento aleatorio** es aquel cuyo resultado está determinado por el **azar**, por esto **no es posible predecir su resultado antes de realizarlo**. Ejemplos de experimentos aleatorios se encuentran lanzar una moneda, lanzar un dado, lanzar un dardo a un tiro al blanco, número de semáforos de color rojo que te encontrarás al ir a casa, cuántas hormigas caminan por la acera de peatones en un tiempo.
# # Sesgo, independencia y justicia
# Decimos que un juego de azar es **justo u honesto** si sus resultados no presentan asimetría (aparecen con la misma frecuencia) y son **independientes** si no presentan patrón alguno. Tirar una moneda, un dado o girar una ruleta son **juegos justos** siempre y cuando no estén alterados de alguna manera.
# ## Ejemplos:
# 1) Supóngase que se lanza un oso de peluche al aire. El oso gira varias veces y cae al suelo de cuatro posibles maneras, panza abajo, panza arriba, sentado o de cabeza. Al lanzarlo 100 veces se obtiene el número de veces que cae de cada forma como se observa en la siguiente tabla:
#
# |resultado|panza abajo|panza arriba|sentado |de cabeza
# |:---------:|:-----------:|:------------:|:--------:|:---------:
# |# de veces| 54|40|5|1
# Claramente se trata de resultados **asimétricos** ya que el oso cae panza abajo más de la mitad de las veces y sólo cae de cabeza uno de cada cien lanzamientos. Los resultados, sin embargo, **son independientes** pues si el oso cae en alguna posición, esto es irrelevante para el siguiente lanzamiento.
# 2) Consideremos una urna con 25 canicas blancas, 25 rojas, 25 amarillas y 25 azules. Sacamos una canica de la urna y **observamos** que es amarilla. **Sin reemplazar** la canica amarilla en la urna tomamos otra canica (equivalente a haber sacado dos canicas). Claramente la urna ya no es la misma pues ahora contiene 25 canicas blancas, rojas y azules y 24 canicas amarillas. Nuestras expectativas para el color de la segunda canica **no son independientes** del resultado de haber tomado una canica amarilla inicialmente. Si la segunda canica es, por ejemplo roja, y **tampoco la reemplazamos**, entonces la urna contiene 25 canicas blancas y azules y 24 rojas y amarillas. Las expectativas para el color de la tercera canica cambian. En este ejemplo, los resultados de **sacar canicas de colores en sucesión y sin reemplazo no son independientes**.
# 3) Consideremos la misma urna del ejemplo anterior. Observemos que si cada vez que **sacamos una canica y anotamos su color la reemplazamos nuevamente en la urna**, entonces el sacar la segunda canica **es independiente** de lo que hayamos hecho antes. La razón es que tenemos, esencialmente, **la misma urna inicial**. Aún mas, si se repite el experimento de extraer y anotar su color (con o sin reemplazo), entonces se le llama **muestreo ordenado** y si no se anota su color se le llama **muestreo no ordenado**, no tenemos idea en qué orden se eligieron las canicas, sólo observamos una o más canicas y no importa el orden en que se sacaron de acuerdo a lo que observamos y esto es equivalente a haber extraído las canicas y colocarlas en una bolsa antes de observar qué sacamos.
# **Obs:** Este modelo de la urna con canicas es utilizado con frecuencia ya que es sumamente práctico para ciertas abstracciones de la realidad y es considerado dentro de la clase de **experimentos generales** pues contiene a experimentos aleatorios más comunes. Por ejemplo, lanzar una moneda dos veces es equivalente a sacar dos canicas de una urna que están etiquetadas con águila y sol. Lanzar un dado es equivalente a sacar una canica de una urna con seis canicas etiquetadas del 1 al 6.
# 4) En un casino observamos que los últimos cinco resultados de la ruleta han sido los siguientes: 10 negro, 17 negro, 4 negro, 15 negro y 22 negro. Al observar esto escuchamos el consejo de un experimentado apostador: “ponga todo su dinero en el rojo pues ya toca que salga rojo”. Sabiamente no le hacemos caso. La razón es simple: **la ruleta no es una urna sin reemplazo sino más bien es una urna con reemplazo**. En cada giro, **cada número tiene la misma probabilidad de aparecer** y se trata de la misma ruleta. Cada giro es **independiente** de los demás por lo que **no hay un patrón definido** y los resultados previos no modifican la habilidad para predecir el resultado del siguiente giro.
# ## Espacio de resultados o espacio muestral
# Supongamos que una acción o experimento puede tener distintas consecuencias o resultados (*outcomes*) y sea $S = \{r_1, r_2, \dots, r_n\}$ el conjunto de resultados posibles. A este conjunto se le conoce como **espacio de resultados o espacio muestral**.
# Por ejemplo, si lanzamos una moneda al aire el espacio muestral es *{águila, sol}* y al tirar un dado de seis caras el espacio muestral es $\{1,2,3,4,5,6\}$. Es importante notar que, en cada caso, los resultados son **mutuamente excluyentes**, es decir, **no pueden ocurrir simultáneamente**. Asimismo, **el espacio muestral comprende a todos los resultados posibles**.
# ## ¿Cómo representar el espacio de resultados o espacio muestral en R?
# Nos podemos apoyar de la estructura *data frame* la cual es una colección rectangular de variables. Cada renglón del *data frame* corresponde a un resultado del experimento (pero se verá más adelante que el *data frame* sólo nos ayudará a describir ciertos espacios de resultados de experimentos).
# ## Ejemplo
# ### Experimento: lanzar un oso de peluche al aire.
# Entonces el espacio muestral es:
S = data.frame(cae=c("panza abajo", "panza arriba", "sentado", "de cabeza"))
S
# ### Experimento: sacar canicas de una urna.
# Supóngase que se tiene una urna con tres canicas con etiquetas $1, 2, 3$ respectivamente. Se sacarán $2$ canicas.
# ### ¿Cómo realizar el experimento en R?
# En el paquete [prob](https://cran.r-project.org/web/packages/prob/index.html) se tiene la función [urnsamples](https://www.rdocumentation.org/packages/prob/versions/1.0-1/topics/urnsamples) la cual tiene argumentos `x, size, replace, ordered`. El argumento `x` representa la urna de la cual se realizará el muestreo, `size` representa el tamaño de la muestra, `replace` y `ordered` son argumentos lógicos y especifican cómo se debe realizar el muestreo.
# ### Con reemplazo y orden
# Como el experimento es con reemplazo se pueden sacar cualquiera de las canicas $1, 2, 3$ en cualquier extracción, además como es con orden **se llevará un registro del orden de las extracciones** que se realizan.
print(urnsamples(1:3, size = 2, replace = TRUE, ordered = TRUE))
# La primer columna con etiqueta $X1$ representa la primera extracción y el primer renglón representa una realización del experimento.
# **Obs:**
#
# * Obsérvese que los renglones $2$ y $4$ son idénticos salvo el orden en el que se muestran los números.
#
# * Este experimento **es equivalente a** lanzar dos veces un dado de tres lados. Lo anterior se realiza en $R$ con:
#
print(rolldie(2, nsides = 3))
# Ver [rolldie](https://www.rdocumentation.org/packages/prob/versions/1.0-1/topics/rolldie).
# ### Sin reemplazo y orden
# Como es sin reemplazo no observaremos en uno de los renglones $1, 1$ por ejemplo (mismo número en un renglón) y como es con orden se tendrán renglones de la forma $2, 1$ y $1, 2$ (pues se consideran distintos).
print(urnsamples(1:3, size=2, replace = F, order = T))
# **Obs:** obsérvese que hay menos renglones en este caso debido al procedimiento más restrictivo de muestreo. Si los números $1, 2, 3$ representaran "Alicia", "Ana" y "Paulina" respectivamente entonces este experimento sería **equivalente a** seleccionar dos personas de tres para que fueran la presidenta y vice-presidenta respectivamente de alguna compañía. El *data frame* anterior representa todas las posibilidades en que esto podría hacerse.
# ### Sin reemplazo y sin orden
# Nuevamente no observaremos en uno de los renglones $1, 1$ por ejemplo (mismo número en un renglón) y como es sin orden tendremos menos renglones que el caso anterior pues al sacar las canicas no se tendrán duplicados de extracciones anteriores no importando el orden de los números.
print(urnsamples(1:3, size=2, replace = F, order = F))
# Este experimento es **equivalente a** ir a donde está la urna, mirar en ella y elegir un par de canicas. Este es el default de la función `urnsamples`:
print(urnsamples(1:3,2))
# #### Con reemplazo y sin orden
# Se reemplazan las canicas en cada extracción pero no se "recuerda" el orden en el que fueron extraídas.
print(urnsamples(1:3, size = 2, replace = T, order = F))
# Este experimento es **equivalente a**:
# * Tener una taza en el que agitamos dos dados de tres caras y nos acercamos a ver la taza.
# * Los resultados de distribuir dos pelotas idénticas de golf en tres cajas etiquetadas con 1, 2 y 3.
# **Notas respecto a la función urnsamples:**
#
# * La urna no necesita tener números, esto es, se podría haber definido un vector $x$ como `x = c('Roja', 'Azul', 'Amarilla')`.
# * Los elementos que contiene la urna siempre son distinguibles para la función `urnsamples`. Entonces situaciones como `x = c('Roja', 'Roja', 'Azul')` no se sugieren ejecutar pues el resultado puede **no ser correcto** (por ejemplo, realizar un experimento en el que se tienen canicas no distinguibles resultan renglones del *data frame* como si se hubiera usado `ordered=T` aún si se eligió `ordered=F`. Enunciados similares aplican para el argumento `replace`).
# ## Eventos
# Un evento $E$ es una colección de resultados del experimento, un subconjunto del espacio muestral
# **Obs:** El conjunto vacío, $\emptyset$, es un evento pues es subconjunto de todo conjunto y en el contexto de eventos representa un evento sin espacio de resultados.
# Bajo la notación de $S = \{r_1, r_2, \dots, r_n\}$ como el espacio muestral, todos los eventos posibles son: $\emptyset , \{r_1\}, \{r_2\}, \dots, \{r_1, r_2\}, \{r_1, r_3\}, \dots \{r_2, r_3\}, \dots \{r_{n-1}, r_n\}, \dots \{r_1, r_2, \dots , r_n\}$
# ## Ocurrencia de un evento
# Decimos que el evento $E$ ocurrió si el resultado de un experimento pertenece a $E$.
# Lo usual es que los eventos se refieran a resultados con alguna característica de interés, por ejemplo, si lanzamos dos dados podrían interesarnos todas las parejas de números cuya suma sea mayor a cinco. Si se trata de una población de individuos, podríamos querer saber algo acerca de todos los que tienen cierto nivel de ingreso, o los que adquieren cierto nivel educativo o los que tuvieron sarampion de niños, etcétera.
# ## Eventos mutuamente excluyentes
# Decimos que los eventos $E_1, E_2, \dots$ son mutuamente excluyentes o ajenos si $E_i \cap E_j = \emptyset$ $\forall E_i \neq E_j$ (sólo ocurre exactamente uno de ellos).
# **Obs:** Como los eventos son subconjuntos es permitido realizar operaciones típicas de conjuntos, en la definición anterior se usó la intersección $\cap$ y $E_i \cap E_j$ consiste de todos los resultados comunes a $E_i$ y $E_j$.
# Por ejemplo, en el caso del lanzamiento de una moneda, los eventos $E_1=\{\text{obtener águila}\}$ y $E_2 = \{\text{obtener sol}\}$ son mutuamente excluyentes y en el caso de $E_1 = \{\text{hoy día soleado}\}$, $E_2 = \{\text{hoy día nublado} \}$ no son mutuamente excluyentes pues tenemos días que son nublados y soleados.
# ### Ejemplo
# **1) Lanzamiento de una moneda un número de veces definido (o lanzamiento de monedas distintas)**
S <- tosscoin(2, makespace = TRUE) #one coin, two tosses
#equivalently tossing two distinct coins
S
print(S[1:3, ]) #example of three events consisting each one of
#each row of S
print(S[c(2,4), ]) #example of two events consisting each one of
#each row of S
# Ver [tosscoin](https://www.rdocumentation.org/packages/prob/versions/1.0-1/topics/tosscoin).
# **2) Baraja**
S<-cards()
head(S)
tail(S)
# A continuación se muestran eventos extraídos del espacio muestral que satisfacen una expresión lógica
#
print(subset(S, suit == 'Heart'))
# Ver [cards](https://www.rdocumentation.org/packages/prob/versions/0.9-1/topics/cards).
# La función %in% checa si cada elemento de un vector está contenido en algún lugar de otro, en el siguiente caso se checa para cada renglón de la columna rank de S que se encuentre en el vector `c(7,8,9)`
print(subset(S, rank %in% 7:9))
# **3) Lanzamiento de un dado tres veces (o lanzamiento de tres dados distintos)**
# Obsérvese que para la función `rolldie` también son aceptadas expresiones matemáticas:
print(subset(rolldie(3), X1+X2+X3 > 16))
# **4) Lanzamiento de un dado cuatro veces (o lanzamiento de cuatro dados distintos)**
# La función [isin](https://www.rdocumentation.org/packages/prob/versions/1.0-1/topics/isin) checa que todo el vector `c(2,2,6)` esté en cada renglón del data.frame `S`.
S <- rolldie(4)
print(subset(S, isin(S, c(2,2,6), ordered = TRUE)))
# **Nota:** otras funciones del paquete `prob` útiles para encontrar espacios muestrales son: [countrep](https://www.rdocumentation.org/packages/prob/versions/1.0-1/topics/countrep) e [isrep](https://www.rdocumentation.org/packages/prob/versions/1.0-1/topics/isrep).
# **Obs:** obsérvese que `%in%` e `%isin%` no realizan lo mismo pues:
x <- 1:10
y <- c(3,3,7)
# La siguiente línea checa que el 3 esté en x, que el 3 esté en x y que el 7 esté en x y devuelve el valor lógico de los tres chequeos, en este caso `all(c(T,T,T))`
all(y %in% x)
# La siguiente línea checa que `c(3,3,7)` esté en `x`:
isin(x,y)
# ### Eventos a partir de operaciones entre conjuntos
# #### Union, Intersección y diferencia
# Un evento es un subconjunto y como tal se realizan operaciones de conjuntos para obtener nuevos eventos. En `prob` se utilizan las funciones [union](https://www.rdocumentation.org/packages/prob/versions/1.0-1/topics/union), [intersect](https://www.rdocumentation.org/packages/prob/versions/1.0-1/topics/intersect) y [setdiff](https://www.rdocumentation.org/packages/prob/versions/1.0-1/topics/setdiff) para tales operaciones. Por ejemplo:
S <- cards()
A <- subset(S, suit == "Heart")
B <- subset(S, rank %in% 7:9)
head(union(A,B)) #se utiliza head para obtener sólo algunos renglones de la operación
intersect(A,B)
setdiff(A,B)
head(setdiff(B,A))
# En la siguiente línea se calcula A^c (el complemento de A definido como S\A)
head(setdiff(S,A))
# ## Modelos de probabilidad
# ### Modelo de la teoría de la medida
# Este modelo consiste en definir una medida de probabilidad en el espacio muestral. Tal medida es una función matemática que satisface ciertos axiomas y tienen ciertas propiedades matemáticas. Existen una amplia gama de medidas de probabilidad de las cuales se elegirá una sola basada en los experimentos y la persona en cuestión que los realizará.
#
# Una vez elegida la medida de probabilidad, todas las asignaciones de probabilidad a eventos están hechas por la misma.
#
# Este modelo se sugiere para experimentos que exhiban simetría, por ejemplo el lanzamiento de una moneda. Si no exhibe simetría o si se desea incorporar conocimiento subjetivo al modelo resulta más difícil la elección de la medida de probabilidad.
#
# <NAME> revolucionó la teoría de la probabilidad con este modelo.
# ### Modelo frecuentista
# Este modelo enuncia que la forma de asignar probabilidades a eventos es por medio de la realización repetida del experimento bajo las mismas condiciones.
# Por ejemplo, si se desea calcular la probabilidad del evento: $E=${obtener sol} entonces: $$P(E) \approx \frac{n_E}{n}$$ donde: $n_E$ representa el número observado de soles (ocurrencia del evento $E$) en $n$ experimentos.
# Tal modelo utiliza la **ley fuerte de los grandes números** en la que se describe y asegura que bajo mismas condiciones de experimentos realizados e independientes, si $n \rightarrow \infty$ entonces $\frac{n_E}{n} \rightarrow P(E)$.
#
# La probabilidad en este enfoque proporciona una medida cuantitativa de qué tan frecuentemente podemos esperar que ocurra un evento.
# Este modelo es sugerido aún si los experimentos no son simétricos (caso del modelo anterior) pero el cálculo de la probabilidad está basado en una aproximación de la forma *in the long run* por lo que no se conoce de forma exacta la misma ni funciona en experimentos que no puedan repetirse indefinidamente, como la probabilidad del evento {el día $x$ lloverá} o {temblor en una zona $z$}.
# <NAME> fue un personaje importante en el impulso de este modelo, además algunas de sus ideas fueron incorporadas en el modelo de teoría de la medida.
# ### Modelo subjetivo
# Se interpreta a la probabilidad como un "grado de creencia" que ocurrirá el evento de acuerdo a la persona que realizará el experimento. La estimación de la probabilidad de un evento se basa en el conocimiento individual de la persona en un punto del tiempo, sin embargo, al ir adquiriendo o poseyendo mayor conocimiento, la estimación se modifica/actualiza de acuerdo a esto. El método típico por el que se actualiza la probabilidad es con la **regla o fórmula o teorema de Bayes**.
# Por ejemplo, supóngase que al inicio del experimento del lanzamiento de una moneda y el evento {sol} la observadora asigna $P({\text{sol}}) = \frac{1}{2}$. Sin embargo, por alguna razón la observadora conoce información adicional acerca de la moneda o de la persona que lanzará la moneda por lo que **decide** modificar su asignación **inicial** de la probabilidad de obtener sol alejado del valor $\frac{1}{2}$.
# Se define la probabilidad como el grado (personal) de creencia o certeza que se tiene de que el evento suceda.
#
# Este modelo se sugiere en situaciones que no es posible repetir indefinidamente el experimento o carece de datos confiables o es prácticamente imposible. Sin embargo, cuando se trata de analizar situaciones para las cuales los datos son escasos, cuestionables o inexistentes, entonces las probabilidades subjetivas difieren enormemente. Un analista deportivo puede pensar que los Cavaliers ganarán el campeonato con un 60% de certeza, mientras que otro puede asegurar que los Lakers de Los Ángeles serán campeones con 95% de certeza.
#
# <NAME>, <NAME>, <NAME>, <NAME> y <NAME> fueron de las personas que popularizaron este modelo.
# **Nota:** Cuando se trabaja con un gran número de datos, los modelos frecuentistas y subjetivos tienden a coincidir pero, cuando los datos son escasos o prácticamente inexistentes, las interpretaciones difieren.
# ### Modelo equiprobable
# Este modelo asigna igual probabilidad a todos los resultados de un experimento y lo podemos encontrar en los modelos anteriores:
#
# * En el modelo de la teoría de la medida al tener un experimento que exhibe simetría de algún tipo, por ejemplo en el lanzamiento de una moneda o dados justos o de un dardo a un tiro al blanco con un mismo radio de circunferencia.
#
# * En el modelo subjetivo si la persona que realiza el experimento tiene ignorancia o indiferencia respecto a su grado de creencia del resultado del experimento.
#
# * En el modelo frecuentista al observar la proporción de veces que al lanzar una moneda se obtiene sol.
#
# Obsérvese que este modelo es posible utilizar si se pueden ennumerar todos los resultados de un experimento.
# Una opción para responder esta pregunta es considerar un objeto, `S`, que represente los *outcomes* o resultados del experimento y un vector de probabilidades, `probs`, con entradas que correspondan a cada outcome en `S`.
# ## Espacio de probabilidad
# Un espacio de probabilidad se compone por el espacio de resultados o espacio muestral, $S$, los eventos $E$ y la medida de los eventos o función de probabilidad que satisfacen los axiomas de Kolmogorov.
# ## ¿Cómo representar en R un espacio de probabilidad?
# Además en el paquete *prob* se tiene una función [probspace](https://www.rdocumentation.org/packages/prob/versions/1.0-1/topics/probspace) que tiene por argumentos `x` que es un espacio muestral de los outcomes y `probs` es un vector del mismo tamaño que el número de outcomes en $x$.
# ### Ejemplos
# **1) Lanzamiento de un dado justo**
outcomes <- rolldie(1)
p <- rep(1/6, times = 6)
probspace(outcomes, probs = p)
#equivalently
#probspace(1:6, probs = p) or probspace(1:6) or rolldie(1, makespace = TRUE)
# **2) Lanzamiento de una moneda cargada**
# Supóngase que $P(\{\text{sol}\}) = .7$ y $P(\{\text{águila}\}) = .3$ entonces:
probspace(tosscoin(1), probs = c(0.70, 0.30))
# **(Tarea) Ejercicio:** ¿cómo calcular la probabilidad anterior con la función `urnsamples`?
outcomes <- urnsamples(1:2, size=1)
prob <- c(0.7, 0.3)
probspace(outcomes, probs=prob)
| R/clases/2_probabilidad/1_elementos_de_probabilidad.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/GoogleCloudPlatform/tensorflow-without-a-phd/blob/master/tensorflow-rnn-tutorial/01_Keras_stateful_RNN_playground.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="RH-br21Mfg83" colab_type="text"
# # An stateful RNN model to generate sequences
# RNN models can generate long sequences based on past data. This can be used to predict stock markets, temperatures, traffic or sales data based on past patterns. They can also be adapted to [generate text](https://docs.google.com/presentation/d/18MiZndRCOxB7g-TcCl2EZOElS5udVaCuxnGznLnmOlE/pub?slide=id.g139650d17f_0_1185). The quality of the prediction will depend on training data, network architecture, hyperparameters, the distance in time at which you are predicting and so on. But most importantly, it will depend on wether your training data contains examples of the behaviour patterns you are trying to predict.
# + id="9l96vOsPfg84" colab_type="code" outputId="a37152a6-fd37-405f-9627-774aa37357cb" colab={"base_uri": "https://localhost:8080/", "height": 35}
import math
import numpy as np
from matplotlib import pyplot as plt
import tensorflow as tf
tf.enable_eager_execution()
print("Tensorflow version: " + tf.__version__)
# + id="gxJ60M-4ilJy" colab_type="code" cellView="form" colab={}
#@title Data formatting and display utilites [RUN ME]
def dumb_minibatch_sequencer(data, batch_size, sequence_size, nb_epochs):
"""
Divides the data into batches of sequences in the simplest way: sequentially.
:param data: the training sequence
:param batch_size: the size of a training minibatch
:param sequence_size: the unroll size of the RNN
:param nb_epochs: number of epochs to train on
:return:
x: one batch of training sequences
y: one batch of target sequences, i.e. training sequences shifted by 1
"""
data_len = data.shape[0]
nb_batches = data_len // (batch_size * sequence_size)
rounded_size = nb_batches * batch_size * sequence_size
xdata = data[:rounded_size]
ydata = np.roll(data, -1)[:rounded_size]
xdata = np.reshape(xdata, [nb_batches, batch_size, sequence_size])
ydata = np.reshape(ydata, [nb_batches, batch_size, sequence_size])
for epoch in range(nb_epochs):
for batch in range(nb_batches):
yield xdata[batch,:,:], ydata[batch,:,:]
def rnn_minibatch_sequencer(data, batch_size, sequence_size, nb_epochs):
"""
Divides the data into batches of sequences so that all the sequences in one batch
continue in the next batch. This is a generator that will keep returning batches
until the input data has been seen nb_epochs times. Sequences are continued even
between epochs, apart from one, the one corresponding to the end of data.
The remainder at the end of data that does not fit in an full batch is ignored.
:param data: the training sequence
:param batch_size: the size of a training minibatch
:param sequence_size: the unroll size of the RNN
:param nb_epochs: number of epochs to train on
:return:
x: one batch of training sequences
y: one batch of target sequences, i.e. training sequences shifted by 1
"""
data_len = data.shape[0]
# using (data_len-1) because we must provide for the sequence shifted by 1 too
nb_batches = (data_len - 1) // (batch_size * sequence_size)
assert nb_batches > 0, "Not enough data, even for a single batch. Try using a smaller batch_size."
rounded_data_len = nb_batches * batch_size * sequence_size
xdata = np.reshape(data[0:rounded_data_len], [batch_size, nb_batches * sequence_size])
ydata = np.reshape(data[1:rounded_data_len + 1], [batch_size, nb_batches * sequence_size])
whole_epochs = math.floor(nb_epochs)
frac_epoch = nb_epochs - whole_epochs
last_nb_batch = math.floor(frac_epoch * nb_batches)
for epoch in range(whole_epochs+1):
for batch in range(nb_batches if epoch < whole_epochs else last_nb_batch):
x = xdata[:, batch * sequence_size:(batch + 1) * sequence_size]
y = ydata[:, batch * sequence_size:(batch + 1) * sequence_size]
x = np.roll(x, -epoch, axis=0) # to continue the sequence from epoch to epoch (do not reset rnn state!)
y = np.roll(y, -epoch, axis=0)
yield x, y
plt.rcParams['figure.figsize']=(16.8,6.0)
plt.rcParams['axes.grid']=True
plt.rcParams['axes.linewidth']=0
plt.rcParams['grid.color']='#DDDDDD'
plt.rcParams['axes.facecolor']='white'
plt.rcParams['xtick.major.size']=0
plt.rcParams['ytick.major.size']=0
plt.rcParams['axes.titlesize']=15.0
def display_lr(lr_schedule, nb_epochs):
x = np.arange(nb_epochs)
y = [lr_schedule(i) for i in x]
plt.figure(figsize=(9,5))
plt.plot(x,y)
plt.title("Learning rate schedule\nmax={:.2e}, min={:.2e}".format(np.max(y), np.min(y)),
y=0.85)
plt.show()
def display_loss(history, full_history, nb_epochs):
plt.figure()
plt.plot(np.arange(0, len(full_history['loss']))/steps_per_epoch, full_history['loss'], label='detailed loss')
plt.plot(np.arange(1, nb_epochs+1), history['loss'], color='red', linewidth=3, label='average loss per epoch')
plt.ylim(0,3*max(history['loss'][1:]))
plt.xlabel('EPOCH')
plt.ylabel('LOSS')
plt.xlim(0, nb_epochs+0.5)
plt.legend()
for epoch in range(nb_epochs//2+1):
plt.gca().axvspan(2*epoch, 2*epoch+1, alpha=0.05, color='grey')
plt.show()
def picture_this_7(features):
subplot = 231
for i in range(6):
plt.subplot(subplot)
plt.plot(features[i])
subplot += 1
plt.show()
def picture_this_8(data, prime_data, results, offset, primelen, runlen, rmselen):
disp_data = data[offset:offset+primelen+runlen]
colors = plt.rcParams['axes.prop_cycle'].by_key()['color']
plt.subplot(211)
plt.xlim(0, disp_data.shape[0])
plt.text(primelen,2.5,"DATA |", color=colors[1], horizontalalignment="right")
plt.text(primelen,2.5,"| PREDICTED", color=colors[0], horizontalalignment="left")
displayresults = np.ma.array(np.concatenate((np.zeros([primelen]), results)))
displayresults = np.ma.masked_where(displayresults == 0, displayresults)
plt.plot(displayresults)
displaydata = np.ma.array(np.concatenate((prime_data, np.zeros([runlen]))))
displaydata = np.ma.masked_where(displaydata == 0, displaydata)
plt.plot(displaydata)
plt.subplot(212)
plt.xlim(0, disp_data.shape[0])
plt.text(primelen,2.5,"DATA |", color=colors[1], horizontalalignment="right")
plt.text(primelen,2.5,"| +PREDICTED", color=colors[0], horizontalalignment="left")
plt.plot(displayresults)
plt.plot(disp_data)
plt.axvspan(primelen, primelen+rmselen, color='grey', alpha=0.1, ymin=0.05, ymax=0.95)
plt.show()
rmse = math.sqrt(np.mean((data[offset+primelen:offset+primelen+rmselen] - results[:rmselen])**2))
print("RMSE on {} predictions (shaded area): {}".format(rmselen, rmse))
# + [markdown] id="ouNkUJLBfg89" colab_type="text"
# ## Generate fake dataset [WORK REQUIRED]
# * Pick a wavewform below: 0, 1 or 2. This will be your dataset.
# + id="QLTqiqjSfg8-" colab_type="code" outputId="14dd77eb-c8a3-4636-9abe-3c4c6e5985d2" colab={"base_uri": "https://localhost:8080/", "height": 375}
WAVEFORM_SELECT = 0 # select 0, 1 or 2
def create_time_series(datalen):
# good waveforms
frequencies = [(0.2, 0.15), (0.35, 0.3), (0.6, 0.55)]
freq1, freq2 = frequencies[WAVEFORM_SELECT]
noise = [np.random.random()*0.1 for i in range(datalen)]
x1 = np.sin(np.arange(0,datalen) * freq1) + noise
x2 = np.sin(np.arange(0,datalen) * freq2) + noise
x = x1 + x2
return x.astype(np.float32)
DATA_LEN = 1024*128+1
data = create_time_series(DATA_LEN)
plt.plot(data[:512])
plt.show()
# + [markdown] id="YyjwHUwZfg9B" colab_type="text"
# ## Hyperparameters
# + id="-gp8Qw-6fg9C" colab_type="code" colab={}
RNN_CELLSIZE = 80 # size of the RNN cells
SEQLEN = 32 # unrolled sequence length
BATCHSIZE = 30 # mini-batch size
DROPOUT = 0.3 # dropout regularization: probability of neurons being dropped. Should be between 0 and 0.5
# + [markdown] id="WbsuSalyfg9E" colab_type="text"
# ## Visualize training sequences
# This is what the neural network will see during training.
# + id="kYk8zx6jfg9F" colab_type="code" outputId="8f8ff1f4-99a1-4258-e109-370e40ccefd9" colab={"base_uri": "https://localhost:8080/", "height": 426}
# The function dumb_minibatch_sequencer splits the data into batches of sequences sequentially.
for features, labels in dumb_minibatch_sequencer(data, BATCHSIZE, SEQLEN, nb_epochs=1):
break
print("Features shape: " + str(features.shape))
print("Labels shape: " + str(labels.shape))
print("Excerpt from first batch:")
picture_this_7(features)
# + [markdown] id="8PB4I6Qafg9H" colab_type="text"
# ## The model [WORK REQUIRED]
# This time we want to train a "stateful" RNN model, one that runs like a state machine with an internal state updated every time an input is processed. Stateful models are typically trained (unrolled) to predict the next element in a sequence, then used in a loop (without unrolling) to generate a sequence.
# 1. Locate the inference function keras_prediction_run() below and check that at its core, it runs the model in a loop, piping outputs into inputs and output state into input state:<br/>
# ```
# for i in range(n):
# Yout = model.predict(Yout)
# ```
# Notice that the output is passed around in the input explicitly. In Keras, the output state is passed around as the next input state automatically if RNN layers are declared with stateful=True
# 1. Run the whole notebook as it is, with a dummy model that always predicts 1. Check that everything "works".
# 1. Now implement a one layer RNN model:
# * Use stateful GRU cells [`tf.keras.layers.GRU(RNN_CELLSIZE, stateful=True, return_sequences=True)`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/GRU).
# * Make sure they all return full sequences with [`return_sequences=True`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/GRU). The model should output a full sequence of length SEQLEN. The target is the input sequence shifted by one, effectively teaching the RNN to predict the next element of a sequence.
# * Do not forget to replicate the regression redout layer across all time steps with [`tf.keras.layers.TimeDistributed(tf.keras.layers.Dense(1))`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/TimeDistributed)
# * In Keras, stateful RNNs must be defined for a fixed batch size ([documentation](https://keras.io/getting-started/faq/#how-can-i-use-stateful-rnns)). On the first layer, in addition to input_shape, please specify batch_size=batchsize
# * Adjust shapes as needed with [`Reshape`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Reshape) layers. Pen, paper and fresh brain cells <font size="+2">🤯</font> still useful to follow the shapes. The shapes of inputs (a.k.a. "features") and targets (a.k.a. "labels") are displayed in the cell above this text.
# 1. Add a second RNN layer.
# * The predictions might be starting to look good but the loss curve is pretty noisy.
# 1. If we want to do stateful RNNs "by the book", training data should be arranged in batches in a special way so that RNN states after one batch are the correct input states for the sequences in the next batch (see [this illustration](https://docs.google.com/presentation/d/18MiZndRCOxB7g-TcCl2EZOElS5udVaCuxnGznLnmOlE/edit#slide=id.g139650d17f_0_584)). Correct data batching is already implemented: just use the rnn_minibatch_sequencer function in the training loop instead of dumb_minibatch_sequencer.
# * This should clean up the loss curve and improve predictions.
# 1. Finally, add a learning rate schedule. In Keras, this is also done through a callback. Edit lr_schedule below and swap the constant learning rate for a decaying one (just uncomment it).
# * Now the RNN should be able to continue your curve accurately.
# 1. (Optional) To do things really "by the book", states should also be reset when sequences are no longer continuous between batches, i.e. at every epoch. The reset_state callback defined below does that. Add it to the list of callbacks in model.fit and test.
# * It actually makes things slightly worse...
# 1. (Optional) You can also add dropout regularization. Try dropout=DROPOUT on both your RNN layers.
# * Aaaarg 😫what happened ?
# 1. (Optional) In Keras RNN layers, the dropout parameter is an input dropout. In the first RNN layer, you are dropping your input data ! That does not make sense. Remove the dropout fromn your first RNN layer. With dropout, you might need to train for longer. Try 10 epochs.
#
#
# 
# <div style="text-align: right; font-family: monospace">
# X shape [BATCHSIZE, SEQLEN, 1]<br/>
# Y shape [BATCHSIZE, SEQLEN, 1]<br/>
# H shape [BATCHSIZE, RNN_CELLSIZE*NLAYERS]
# </div>
# In Keras layers, the batch dimension is implicit ! For a shape of [BATCHSIZE, SEQLEN, 1], you write [SEQLEN, 1]. In pure Tensorflow however, this is NOT the case.
# + id="9ga_jncykosN" colab_type="code" colab={}
def keras_model(batchsize, seqlen):
model = tf.keras.Sequential([
#
# YOUR MODEL HERE
# This is a dummy model that always predicts 1
#
tf.keras.layers.Lambda(lambda x: tf.ones([batchsize,seqlen]), input_shape=[seqlen,])
])
# keras does not have a pre-defined metric for Root Mean Square Error. Let's define one.
def rmse(y_true, y_pred): # Root Mean Squared Error
return tf.sqrt(tf.reduce_mean(tf.square(y_pred - y_true)))
# to finalize the model, specify the loss, the optimizer and metrics
model.compile(
loss = 'mean_squared_error',
optimizer = 'adam',
metrics = [rmse])
return model
# + id="8311WBkinlGp" colab_type="code" colab={}
# Keras model callbacks
# This callback records a per-step loss history instead of the average loss per
# epoch that Keras normally reports. It allows you to see more problems.
class LossHistory(tf.keras.callbacks.Callback):
def on_train_begin(self, logs={}):
self.history = {'loss': []}
def on_batch_end(self, batch, logs={}):
self.history['loss'].append(logs.get('loss'))
# This callback resets the RNN state at each epoch
class ResetStateCallback(tf.keras.callbacks.Callback):
def on_epoch_begin(self, batch, logs={}):
self.model.reset_states()
print('reset state')
reset_state = ResetStateCallback()
# learning rate decay callback
def lr_schedule(epoch): return 0.01
#def lr_schedule(epoch): return 0.0001 + 0.01 * math.pow(0.8, epoch)
lr_decay = tf.keras.callbacks.LearningRateScheduler(lr_schedule, verbose=True)
# + [markdown] id="XlKtnYjbfg9V" colab_type="text"
# ## The training loop
# + id="Tcrj-e50yyuM" colab_type="code" colab={}
# Execute this cell to reset the model
NB_EPOCHS = 8
model = keras_model(BATCHSIZE, SEQLEN)
# this prints a description of the model
model.summary()
display_lr(lr_schedule, NB_EPOCHS)
# + [markdown] id="sx7jowlUqhfu" colab_type="text"
# You can re-execute this cell to continue training
# + id="UM0AIbCSfg9V" colab_type="code" cellView="both" colab={}
# You can re-execute this cell to continue training
steps_per_epoch = (DATA_LEN-1) // SEQLEN // BATCHSIZE
#generator = rnn_minibatch_sequencer(data, BATCHSIZE, SEQLEN, NB_EPOCHS)
generator = dumb_minibatch_sequencer(data, BATCHSIZE, SEQLEN, NB_EPOCHS)
full_history = LossHistory()
history = model.fit_generator(generator,
steps_per_epoch=steps_per_epoch,
epochs=NB_EPOCHS,
callbacks=[full_history, lr_decay, reset_state])
# + id="u_TxVHXPfg9Y" colab_type="code" colab={}
display_loss(history.history, full_history.history, NB_EPOCHS)
# + [markdown] id="xBmt2uUVfg9N" colab_type="text"
# ## Inference
# This is a generative model: run one trained RNN cell in a loop
# + id="UFalPiNOr2pt" colab_type="code" colab={}
# Inference from stateful model
def keras_prediction_run(model, prime_data, run_length):
model.reset_states()
data_len = prime_data.shape[0]
#prime_data = np.expand_dims(prime_data, axis=0) # single batch with everything
prime_data = np.expand_dims(prime_data, axis=-1) # each sequence is of size 1
# prime the state from data
for i in range(data_len - 1): # keep last sample to serve as the input sequence for predictions
model.predict(np.expand_dims(prime_data[i], axis=0))
# prediction run
results = []
Yout = prime_data[-1] # start predicting from the last element of the prime_data sequence
for i in range(run_length+1):
Yout = model.predict(Yout)
results.append(Yout[0,0]) # Yout shape is [1,1] i.e one sequence of one element
return np.array(results)
# + id="DBVi1tNofg9a" colab_type="code" colab={}
PRIMELEN=256
RUNLEN=512
OFFSET=20
RMSELEN=128
prime_data = data[OFFSET:OFFSET+PRIMELEN]
# For inference, we need a single RNN cell (no unrolling)
# Create a new model that takes a single sequence of a single value (i.e. just one RNN cell)
inference_model = keras_model(1, 1)
# Copy the trained weights into it
inference_model.set_weights(model.get_weights())
results = keras_prediction_run(inference_model, prime_data, RUNLEN)
picture_this_8(data, prime_data, results, OFFSET, PRIMELEN, RUNLEN, RMSELEN)
# + [markdown] id="AQn6WA-pfg9c" colab_type="text"
# Copyright 2018 Google LLC
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
# [http://www.apache.org/licenses/LICENSE-2.0](http://www.apache.org/licenses/LICENSE-2.0)
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
| tensorflow-rnn-tutorial/01_Keras_stateful_RNN_playground.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # "Rock, Paper, Scissors" Probabilities
#
# ---
#
# Sometimes in life there are things we can't do - or at least things we'd rather not attempt. Like after we're stuffed from a great meal, have yet another chicken wing.
#
# After a lunch with 6 people, one chicken wing remained, with no takers. So, we "rock, paper, scissors"-ed it out for several rounds until a loser emerged who would take responsibility to clear the plate. But with 6 people showing a hand of "Rock, Paper, Scissors", we may need to play several rounds before anyone is eliminated. For example, if all 6 players showed "paper", then no one would be eliminated. Likewise, if, between the 6 players, rock, paper and scissors were all shown at least once, then the round would also be a tie since no players were eliminated. Therefore, an elimination can only occur when there are 2 and only 2 unique items shown (either "paper, rock", "rock, scissors" or "paper, scissors").
#
# ### *What is the probability that 'n' players would not get a 'Draw' on a given round?*
#
#
# ## 1 - Define the function
#
#
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# +
def rps(k=3, n=3):
'''
Pass in number of players and hands, output the probability of a non-draw
k = number of players in "rock, paper, scissors"
n = number of 'hands'; default is 3 ('R', 'P', 'S')
'''
def calc_numerator(k, n):
if k == 1:
return 0
else:
return ((n - 1)**(k - 1)) + calc_numerator(k - 1, n)
if k < 1:
return "n cannot be less than 1"
elif k == 1:
return 0
else:
return calc_numerator(k, n) / n**(k - 1)
for i, x in enumerate(range(1, 21)):
print(" " if i < 9 else "", "{}: {:.2%}".format(i + 1, rps(x)), sep="")
# -
# We can see our answer for the probability of 6 players NOT getting a draw is 25.51%.
rps(6)
# ## 2 - Visualize the Actual Probabilities
# +
import matplotlib.pyplot as plt
import matplotlib.ticker as mtick
players = 6
odds = [rps(x) for x in range(1, 16)]
colors = ["tab:blue" if x!=6 else "tab:red" for x in range(1, len(odds) + 1)]
fig, ax = plt.subplots()
ax.scatter(range(1, len(odds) + 1), odds, color=colors, s=100, alpha=0.7)
ax.set_xlabel("'n' Players", fontsize=14)
ax.set_xticks(range(1, len(odds) + 1)[::2])
ax.set_ylabel("Probability", fontsize=14, labelpad=10)
ax.set_ylim(-0.02, 1)
ax.yaxis.set_major_formatter(mtick.PercentFormatter(1.0))
ax.set_title("Probability of NOT Getting a 'Draw' in\nRock, Paper, Scissors with 'n' Players",
fontsize=16, alpha=0.8)
ax.tick_params(axis="both", which="both", length=0, labelsize=12)
ax.annotate("{} players:\n{:.2%}".format(players, odds[players - 1]),
(players * 1.05, odds[players - 1] * 1.05), (players + 2, odds[players - 1] + 0.2),
fontsize=14, alpha=0.7, color="tab:red", fontweight='bold', arrowprops={"arrowstyle": '->'})
fig.set_size_inches(8, 6)
plt.show()
# -
# ## 3 - Double Check with Pandas
#
# - Use a list comprehension to obtain all the permutations/possibilities of a round of "Rock, Paper, Scissors" given 6 players
# - Import it into a pandas DataFrame
# - Apply a function to each row ("round") to calculate how many unique items there were per round
#
# Those rows (rounds) with 2 unique items were "no draw" rounds.
# +
import pandas as pd
options = ["Rock", "Paper", "Scissors"]
permutations = [(x, y, z, a, b, c) for x in options for y in options for z in options
for a in options for b in options for c in options]
df = pd.DataFrame(permutations)
df.head()
# -
# With a total of 186 permutations in which "no draw" is possible, the probability is 186/729 = 25.51%.
results = df.apply(lambda x: x.nunique(), axis=1).value_counts()
results
"{:.2%}".format(results[2] / results.sum())
# ## 4 - Triple Check with Random Simulations
#
# Create a function to simulate 'size' rounds of Rock, Paper, Scissors with 'n' players.
# +
import numpy
np.random.seed(0)
def exper(size, n):
experiment = np.random.choice(["Rock", "Paper", "Scissors"], size=(size, n))
try:
exper_odds = pd.DataFrame(experiment).nunique(axis=1).value_counts().loc[2] / size
except KeyError: # With larger values of "n", small sample sizes will often ONLY have nunique
return 0 # of 3, thereby raising a KeyError. If '.loc[2]' doesn't exist, return 0.
return exper_odds
# -
# "full_odds" runs 3 experiments of 100, 1000, and 5000 rounds of players ranging from 2 to 15, and records what percentage of each combination saw "non-draw" rounds.
full_odds = [[exper(s, n) for n in list(range(2, 16))] for s in [100, 1000, 5000]]
full_odds[0]
# ### Visualize the Random Simulations vs. Actual Probabilities
#
# We can see that as the number of simulations increases, the simulated probability matches closer to the actual probability based on the previously defined function.
# +
import matplotlib.pyplot as plt
import matplotlib.ticker as mtick
import numpy
players = 6
odds = [rps(x) for x in range(1, 16)]
sizes = [100, 1000, 5000]
fig, ax = plt.subplots()
ax.plot(range(1, len(odds) + 1), odds, label="Actual Probability", alpha=0.9, lw=3)
for n in range(3):
ax.plot(range(2, len(odds) + 1), full_odds[n], label="{} simulations".format(sizes[n]), alpha=0.9, lw=3)
ax.set_xlabel("'n' Players", fontsize=14)
ax.set_xticks(range(1, len(odds) + 1)[::2])
ax.set_ylabel("Probability", fontsize=14, labelpad=10)
ax.yaxis.set_major_formatter(mtick.PercentFormatter(1.0))
ax.set_title("Probability of NOT Getting a 'Draw' in\nRock, Paper, Scissors with 'n' Players",
fontsize=16, alpha=0.8)
ax.tick_params(axis="both", which="both", length=0, labelsize=12)
ax.legend()
fig.set_size_inches(8, 6)
plt.show()
# +
import matplotlib.pyplot as plt
import matplotlib.ticker as mtick
import numpy
def exper(size, n):
experiment = np.random.choice(["Rock", "Paper", "Scissors"], size=(size, n))
try:
exper_odds = pd.DataFrame(experiment).nunique(axis=1).value_counts().loc[2] / size
except KeyError:
return 0
return exper_odds
full_odds = [[exper(s, n) for n in list(range(2, 16))] for s in [100, 1000, 5000]]
players = 6
odds = [rps(x) for x in range(1, 16)]
sizes = [100, 1000, 5000]
fig, axes = plt.subplots(2, 2, sharex=True, sharey=True)
for i, x in enumerate([(i, j) for i in range(2) for j in range(2)]):
ax = axes[x]
if x==(0, 0):
ax.plot(range(1, len(odds) + 1), odds, label="Actual Probability", alpha=0.9, lw=3)
ax.set_title("All Simulations")
else:
ax.plot(range(1, len(odds) + 1), odds, label="Actual Probability", alpha=0.9, lw=3)
ax.plot(range(2, len(odds) + 1), full_odds[i-1], label="{} simulations".format(sizes[i-1]),
alpha=0.9, lw=3, color="C{}".format(i))
ax.set_title("{} Simulations".format(sizes[i-1]))
ax.set_xticks(range(1, len(odds) + 1)[::2])
ax.yaxis.set_major_formatter(mtick.PercentFormatter(1.0))
ax.tick_params(axis="both", which="both", length=3, labelsize=12)
ax.legend()
for each in [(1, 0), (1, 1)]:
axes[each].set_xlabel("'n' Players", fontsize=12, labelpad=10)
fig.suptitle('Computer Simulations of the Probability of "Non-Draw" Games\nin Rock, Paper, Scissors with "n" Players',
fontsize=16, alpha=0.8)
fig.set_size_inches(10, 8)
plt.show()
# -
# # Conclusion
#
# With 6 people playing, you'd need to play an average of 4 rounds to get someone eliminated. Fourteen people, 100 rounds on average for at least one to be eliminated. Twenty people, 1000 rounds on average. By the time *anyone* would be eliminated, dinner would have come and the players would no longer be competing to not eat the last wing but to eat it.
| projects/Rock, Paper, Scissors - Probabilities.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/srfinley/DS-Unit-1-Sprint-1-Dealing-With-Data/blob/master/module3-databackedassertions/LS_DS_113_Making_Data_backed_Assertions_Assignment.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="Okfr_uhwhS1X" colab_type="text"
# # Lambda School Data Science - Making Data-backed Assertions
#
# This is, for many, the main point of data science - to create and support reasoned arguments based on evidence. It's not a topic to master in a day, but it is worth some focused time thinking about and structuring your approach to it.
# + [markdown] id="lOqaPds9huME" colab_type="text"
# ## Assignment - what's going on here?
#
# Consider the data in `persons.csv` (already prepared for you, in the repo for the week). It has four columns - a unique id, followed by age (in years), weight (in lbs), and exercise time (in minutes/week) of 1200 (hypothetical) people.
#
# Try to figure out which variables are possibly related to each other, and which may be confounding relationships.
#
# Try and isolate the main relationships and then communicate them using crosstabs and graphs. Share any cool graphs that you make with the rest of the class in Slack!
# + id="lML_B6Hp2JRv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="37ca0e6f-11d9-45e9-aa50-1340fb6dbec4"
pd.__version__
# + id="-TWO_KFZ2ZOs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 348} outputId="e47da3c6-e90b-4324-92d5-d1d47b532b3a"
# !pip install pandas==0.23.4
# + id="TGUS79cOhPWj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 206} outputId="6881dcc7-4be4-41df-c0b8-ef52bbe3ae3f"
# TODO - your code here
# Use what we did live in lecture as an example
# HINT - you can find the raw URL on GitHub and potentially use that
# to load the data with read_csv, or you can upload it yourself
import pandas as pd
df = pd.read_csv('https://raw.githubusercontent.com/srfinley/DS-Unit-1-Sprint-1-Dealing-With-Data/master/module3-databackedassertions/persons.csv')
df.head()
# + id="ah5IOzeDzDtx" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 300} outputId="604ec22b-fa75-4869-8514-c3887df536d3"
df.describe()
# + id="DzUCx_8dssVl" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 269} outputId="71e8f651-44ec-4553-d7d7-64196d033570"
#histograms
df['age'].hist(bins=20);
#age is distributed fairly flatly between 18 and 80; there's variation, but nothing systematic
# + id="T_N94mskzR-W" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 269} outputId="19b03c2f-da78-4294-cae2-df65a0e72e46"
df['weight'].hist(bins=20);
#fairly steady dropoff past 160 pounds
#since the data only includes adults, we see none of the leftmost part of the presumed normal distribution
# + id="7EN6zvSA0PF7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 269} outputId="f56aca4c-8db2-4126-ea96-9fd73e8999c5"
df['exercise_time'].hist(bins=20);
#again, somewhat chaotically flat distribution; drops off a bit past 200 minutes
# + id="vwffC0v20x8F" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 283} outputId="acff3404-1844-407f-e60b-0dfd62adfd80"
df.plot.scatter('age','weight');
#nothing really jumps out here
# + id="gyadFi5A1PzP" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 284} outputId="3537c1bb-89d2-454d-e0e2-a2b977bb91d3"
df.plot.scatter('exercise_time', 'weight');
#this one's a lot more interesting!
#it looks like exercise time puts a hard cap on weight,
#but any imagined causation could easily be reversed,
#e.g., if a high weight makes long-duration exercise prohibitively unpleasant
# + id="CX9TU9uH2vB2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 283} outputId="6eedfefd-2c2b-43b6-ebde-e930ac139e86"
df.plot.scatter('age','exercise_time');
#the same kind of hard slope in the weight/exercise plot is visible here as well
#could be something like "past about sixty, there exists a fairly hard endurance cap that lowers with age"
#or: "old people who exercise too hard die on the treadmill"
#at all ages, distribution between the maximum (capped or uncapped) and 0 seems pretty even
# + id="FEZFP04q3sxG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 269} outputId="d5c8ac41-0a86-49f9-a851-f543410cd4c0"
age_bins = pd.cut(df['age'], 6) #corresponds to about six decades represented
weight_bins = pd.cut(df['weight'], 5)
exercise_bins = pd.cut(df['exercise_time'], 6)
pd.crosstab(age_bins,weight_bins)
#at each age, a plurality of subjects are in the first or second weight category
#aside from the oldest plurality, which is in the third weight category
# + id="hSV-Fkz55gti" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 269} outputId="b90dbcca-25fe-4b85-9a65-c266e6584cf8"
pd.crosstab(age_bins,weight_bins,normalize='columns')
#pecentage membership in the highest weight categories roughly increases with age
#but like..... very roughly
#maybe not really at all, aside from the oldest, fattest group, which has a spike in membership
#when will I learn how to determine which maybe-trends are statistically significant
# + id="GLW-Ui2_68uS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 269} outputId="466664de-b5dc-4f77-f228-e417183b4649"
pd.crosstab(exercise_bins,weight_bins)
#you can see the "ceiling" in the zeroes on the lower right corner
#increasing weight categories have distinct descending modal exercise values (with mostly clear dropoffs on either side)
#this wasn't clear from looking at the scatterplot!
# + id="dvrHvi6AXAoO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 286} outputId="c95142aa-018e-4ca4-cd66-2012e953dd92"
pd.crosstab(exercise_bins,age_bins,normalize='columns')
# + id="pA6AdNHw8pWl" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 306} outputId="de016017-0a85-41d9-a1bc-6e3bd731ad47"
pd.crosstab(weight_bins, [exercise_bins, age_bins],normalize='columns')
#too many bins to look at...
# + id="m8DtCs6c9F8b" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 212} outputId="5a00c26d-0bb1-4153-b4a9-08b6d423edcc"
age_bins2 = pd.cut(df['age'], 2)
weight_bins2 = pd.cut(df['weight'], 2)
exercise_bins2 = pd.cut(df['exercise_time'], 2)
pd.crosstab(weight_bins2, [exercise_bins2, age_bins], normalize='columns')
#at lower exercise times across all ages, it's about 60/40 whether you'll be in the high or low weight category
#at high exercise times it's more like 90/10, but again, fairly consistent across age categories
#numbers in the high-exercise high-weight quadrant are very low in general
# + id="5bjq7BDJBnmn" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 273} outputId="da0bc3dd-fd63-4c43-95fb-75b599fdd931"
age_bins3 = pd.cut(df['age'], 3)
weight_bins3 = pd.cut(df['weight'], 3)
exercise_bins3 = pd.cut(df['exercise_time'], 3)
crosstab = pd.crosstab(weight_bins3, [exercise_bins2, age_bins3], normalize='columns')
crosstab.plot();
#the two difference exercise bin clusters have distinct shapes here, with age largely irrelevant
#I'd summarize it as: for those who exercise little, exercise and weight are unrelated
#for those who exercise more, increased exercise leads to decreased chance of high weight
# + id="N7QDEhMsFY3q" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 273} outputId="84d6be2d-1054-4e04-f6d3-9287aeacdc39"
crosstab = pd.crosstab(weight_bins3, [exercise_bins2, age_bins3])
crosstab.plot();
#this looks substantially different from the non-normalized one and I don't know how to interpret that
#I guess the general shapes of the curves are the same
# + id="FwlMMnRQI_tG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 273} outputId="7d73387c-9824-4eca-c05b-4fd110624931"
crosstab = pd.crosstab(weight_bins3, [exercise_bins3, age_bins2], normalize='columns')
crosstab.plot();
#I think this affirms the "the more you exercise, the more your weight is related to exercise" interpretation
#which is really just another way of saying the same thing the weight/exercise scatterplot showed;
#the full range of weights exists at 0 exercise and the range narrows as exercise increases
# + id="0rgn5cz-EOUo" colab_type="code" colab={}
import matplotlib.pyplot as plt
# + id="Uc4VhpG9Moz-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 269} outputId="4d42f9e4-9614-4211-ba48-68278e20aa28"
plot = plt.scatter(df['exercise_time'],df['weight'],c=df['age']);
#x axis is exercise time, y axis is weight, color is age
# + [markdown] id="BT9gdS7viJZa" colab_type="text"
# ### Assignment questions
#
# After you've worked on some code, answer the following questions in this text block:
#
# 1. What are the variable types in the data?
# - all are basically continuous
# 2. What are the relationships between the variables?
# - exercise time and weight are somewhat related. The higher the exercise time, the narrower the range of weight, creating a "missing corner" where high-exercise high-weight data points would go
# - age and exercise time have a smaller missing corner but little relationship beyond that
# - age and weight are largely unrelated, except at the oldest, fattest extreme
# 3. Which relationships are "real", and which spurious?
# - Exercise/weight relationship seems pretty robust for higher exercise values. Maybe the age/exercise relationship is an artifact? I honestly can't tell and I'm not getting anywhere further tonight.
# + [markdown] id="_XXg2crAipwP" colab_type="text"
# ## Stretch goals and resources
#
# Following are *optional* things for you to take a look at. Focus on the above assignment first, and make sure to commit and push your changes to GitHub.
#
# - [Spurious Correlations](http://tylervigen.com/spurious-correlations)
# - [NIH on controlling for confounding variables](https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4017459/)
#
# Stretch goals:
#
# - Produce your own plot inspired by the Spurious Correlation visualizations (and consider writing a blog post about it - both the content and how you made it)
# - Pick one of the techniques that NIH highlights for confounding variables - we'll be going into many of them later, but see if you can find which Python modules may help (hint - check scikit-learn)
| module3-databackedassertions/LS_DS_113_Making_Data_backed_Assertions_Assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# # Data access
#
# ## CSV files
#
# There's multiple functions in the `readr` for CSV file reading. Let's use them on a dataset available in Kaggle that has homemade beer recipes from Brewer's Friend [[1]](https://www.kaggle.com/jtrofe/beer-recipes).
#
# Let's check the first few lines of the data with base R's `file` and `readLines`.
# +
example_file <- file("/m/jhnas/jupyter/shareddata/python-r-data-analysis/beers/recipeData.csv",'r')
first_lines <- readLines(example_file,n=2)
close(example_file)
print(first_lines)
# -
# Before we choose which reader we want to use we need to check the format of the data.
#
# `readr` has predefined functions for the following data formats [[1]](http://readr.tidyverse.org/reference/read_delim.html):
# - `read_delim` parses generic data delimited by a character
# - `read_csv` assumes that the data is delimited by commas
# - `read_csv2` assumes that the data is delimited by semicolons
# - `read_tsv` assumes that the data is delimited by tabs
#
# In this case we want to use `read_csv`. Do note that we limit ourselves to first 100 values for faster parsing.
# +
library(tidyverse)
beer_recipes <- read_csv("/m/jhnas/jupyter/shareddata/python-r-data-analysis/beers/recipeData.csv", n_max=100)
# -
# From the output one can see that `read_csv` tries to parse the datatype of the column automatically.
#
# By running `spec` one can see the full definitons.
spec(beer_recipes)
# Many of the data columns seem to be characters instead of numbers. Let's use `col_types`-argument to specify a better definition.
beer_recipes <- read_csv("/m/jhnas/jupyter/shareddata/python-r-data-analysis/beers/recipeData.csv",
col_types=cols(
BeerID = col_integer(),
Name = col_character(),
URL = col_character(),
Style = col_character(),
StyleID = col_integer(),
`Size(L)` = col_double(),
OG = col_double(),
FG = col_double(),
ABV = col_double(),
IBU = col_double(),
Color = col_double(),
BoilSize = col_double(),
BoilTime = col_double(),
BoilGravity = col_double(),
Efficiency = col_double(),
MashThickness = col_double(),
SugarScale = col_character(),
BrewMethod = col_character(),
PitchRate = col_double(),
PrimaryTemp = col_double(),
PrimingMethod = col_character(),
PrimingAmount = col_character()
),
n_max=100
)
# This produced a lot of problems. Let's check the problems with `problems`.
problems(beer_recipes)
# Most of the problems seem to be related to _N/A_ not being a recognized name for `NA`. Let's add that to the initial read call with `na`-argument.
# +
beer_recipes <- read_csv("/m/jhnas/jupyter/shareddata/python-r-data-analysis/beers/recipeData.csv",na=c("","NA","N/A"), n_max=100)
spec(beer_recipes)
# -
# Now most of the columns seem correct. Last column seems to include units (_oz_). Using mutate is probably easiest way of getting rid of them.
#
# Let' use `str_remove` to remove it [[str_remove]](https://stringr.tidyverse.org/reference/str_remove.html).
#
# After that we can convert the column to double and use `str` to check that our dataset looks fine.
# +
beer_recipes <- beer_recipes %>%
mutate(PrimingAmount=as.double(str_remove(PrimingAmount, ' oz')))
str(beer_recipes)
# -
# Now that we know that the data will be read in the correct format, we can load the full dataset:
# +
beer_recipes <- read_csv("/m/jhnas/jupyter/shareddata/python-r-data-analysis/beers/recipeData.csv",na=c("","NA","N/A")) %>%
mutate(PrimingAmount=as.double(str_remove(PrimingAmount, ' oz')))
str(beer_recipes)
# -
# # Storing data
#
# ## CSV
# Let's say that we want to write the resulting `tibble` in a CSV so that we can share it with other researchers in a simple format.
#
# For this we'd want to use `write_csv` [[write_csv]](http://readr.tidyverse.org/reference/write_delim.html).
write_csv(beer_recipes, 'beer-recipes.csv')
# # Feather
#
# Let's say you have a big dataset you have pre-processed with R, but want to analyze with Python. The new feather-format that uses Apache Arrow's data specification is created by the creators of Tidy-R and Pandas and it should be interoprable with both of them [[feather's page in Github]](https://github.com/wesm/feather).
#
# What matters the most is that it is fast and compact (because it is a binary data format).
#
# Using it is simple, just load `feather`-library an write data with `write_feather` [[write_feather]](https://cran.r-project.org/web/packages/feather/feather.pdf).
#
# Loading data is done with `read_feather`.
#
# Do note that more complex structures like nested tibbles do not necessarily fit into a feather.
#
# Let's install `feather`:
if (!file.exists('rlibs')) {
dir.create('rlibs')
}
if (!file.exists('rlibs/feather')) {
install.packages('feather', repos="http://cran.r-project.org", lib='rlibs')
}
library(feather, lib.loc='rlibs')
# +
write_feather(beer_recipes,'beer_recipes.feather')
beer_recipes2 <- read_feather('beer_recipes.feather')
# -
# This is to fix a bug in our system
Sys.setlocale('LC_ALL','C')
beer_recipes2
# # Database access
#
# There exists a package `DBI` that defines a common interface that can be used to access various different databases [[DBI]](https://dbi.r-dbi.org/).
#
# When using `DBI`, one can also use `dbplyr` to run `tidyverse` verbs (`select`, `map`, etc.) to database queries without loading the whole database into memory.
if (!file.exists('rlibs')) {
dir.create('rlibs')
}
if (!file.exists('rlibs/DBI')) {
install.packages('DBI', repos="http://cran.r-project.org", lib='rlibs')
}
if (!file.exists('rlibs/dbplyr')) {
install.packages('dbplyr', repos="http://cran.r-project.org", lib='rlibs')
}
library(DBI, lib.loc='rlibs')
library(dbplyr, lib.loc='rlibs')
# Let's use `tbl_memdb` from `dbplyr` to add `beer_recipes` to a temporary in-memory database [[tbl_memdb]](https://dbplyr.tidyverse.org/reference/memdb_frame.html).
beers_db <- tbl_memdb(beer_recipes)
# The returned object is a SQL table that acts like a tibble.
print(beers_db)
beers_db %>%
select(Name, ABV, OG, FG)
# The difference between using normal tibbles and this database connection is that database queries are all connected and only evaluated when data is requsted. One can see the query that would be made with `show_query` [[show_query]](https://dplyr.tidyverse.org/reference/explain.html).
beers_db %>%
filter(ABV > 5) %>%
summarise(mean = mean(OG)) %>%
show_query()
# One can collect the data to the current R session with `collect` [[collect]](https://dbplyr.tidyverse.org/reference/collapse.tbl_sql.html).
beers_over_5abv <- beers_db %>%
filter(ABV > 5) %>%
select(Name, ABV, OG, FG) %>%
collect()
str(beers_over_5abv)
# To remove tables from the temporary memory database, one needs to get the connection to it with `src_memdb` and `db_drop_table` from `dplyr`'s SQL backend [[src_memdb]](https://dbplyr.tidyverse.org/rmeference/memdb_frame.html) [[db_drop_table]](https://dplyr.tidyverse.org/reference/backend_dbplyr.html).
memdb <- src_memdb()
memdb
db_drop_table('beer_recipes', con=memdb$con)
memdb
# One can also write to a database file by opening a database connection with DBI.
con <- DBI::dbConnect(RSQLite::SQLite(), dbname = "beer_recipes.sqlite")
# Let's use `copy_to`-function to copy the `beer_recipes`-dataframe into a table in this newly established SQLite [[copy_to]](https://dplyr.tidyverse.org/reference/copy_to.html).
copy_to(con, beer_recipes, overwrite=TRUE, temporary=FALSE)
# To get a reference to this new tab, we can use the `tbl`-function [[tbl]](https://dplyr.tidyverse.org/reference/tbl.html).
beers_sqlite <- tbl(con,'beer_recipes')
beers_sqlite
# Let's close and re-open the connection to verify that the table is indeed in the database file.
dbDisconnect(con)
con
con <- DBI::dbConnect(RSQLite::SQLite(), dbname = "beer_recipes.sqlite")
con
tbl(con,'beer_recipes')
dbDisconnect(con)
con
# # Exercise time:
#
# 1. Modify column specifications for FIFA World Cup match data [[1]](https://www.kaggle.com/abecklas/fifa-world-cup). Use `col_datetime` in `col_types` to get a good specification for column _DateTime_ [[col_datetime]](http://readr.tidyverse.org/reference/parse_datetime.html). Use `col_factor` to make columns _Round_, _Stadium_, _City_, _HomeTeam_ and _AwayTeam_ into factors.
# 2. Store the resulting tibble as a feather.
# 3. Store the resulting tibble into a SQLite database.
fifa_matches <- read_csv("/m/jhnas/jupyter/shareddata/python-r-data-analysis/fifa/WorldCupMatches.csv")
str(fifa_matches)
| r/03_Data_Access.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:pytorch-env]
# language: python
# name: conda-env-pytorch-env-py
# ---
# # Lecture 2: numbers, strings and variables
# What we will review:
#
# 1) Variables
#
# 2) Basic math operations and arithmetic errors
#
# 3) Processing strings
# ## 1) Variables in python
# Recall from PIC10A that in C++ a variable is a memory location with a unique name. In Python a variable is a name bound to particular memory location. In particular this has the following implications:
# - Python variables are not necessarily unique! The same memory location can have multiple names potentially
# - Assignment binds a variable or name to a memory location
# - Declaring a python variables requires only assignment
# - As names can be bound to different address the same name can be used to represent multiple different data types (Python is dynamically typed!)
x = 1000
y = 1000
x==y, x is y
x = 1000
y = x
x==y, x is y
x = 2
y = 4
z = 4/x
x == z, x is z
x = 2
y = 4
z = 4/x
x == z, x is z
# Recall the difference between **mutable** and **immutable** data types:
# - immutable data types are those that cannot be changed after instantiation,
# - mutable data types can!
fruits = ['apple', 'grapes', 'pears']
fruits[0] = 'Apple'
fruits
fruits = ['apple', 'grapes', 'pears']
fruits[0][0] = 'A'
fruits
# When handling mutable objects important to note that changing the data stored at a memory location will impact all names bound to that memory location!
x = [1,2,3]
y = [x,x,x]
x[0] = 4
y[0][0] = 1
x
# ## 2) Basic math operations and some comments on arithmetic error
# +
# Linear combinations
# +
# Floor division
# +
# Modulus
# +
# Powers
# -
37.4 % 12.2 # Example from self-study video
# ### Exploring approximation error with a 6 bit calculator
# Suppose our calculator only has 6 bits in which the first bit represents the sign and all other bits correspond to descending powers of 2, starting at 3. This is an example of a fixed point format.
# - How many distinct numbers can one represent?
# - What is the smallest number that can be represented?
# - What is the largest number?
# - Which representation best approximates 1/2? Is there an approximation error?
# - Which representation best approximates 10/16? Is there an approximation error?
# - If we try to carry out 10/32 + 11/32 what does our calculator return?
# ### Floating point arithmetic
# Floating point arithmetic adopts the following convention.
# $$
# significand \times base^{exponent}
# $$
# Example: $1.2345 = 12345 \times 10^{-4}$
#
# Floating point is fairly ubiquitious and well standardised. Consists of two fixed point representations (each with a signed bit), the significand and the exponent. IEEE floating point representations:
# - 32 bit: 24 bit significand, 8 bit exponent
# - 64 bit: 53 bit significand, 11 bit exponent
#
# Questions:
# - Why do you think it is called floating point as opposed to the fixed point example we gave above?
# - What is the key benefit of floating point over fixed?
#
#
# This is not a course in scientific computing! So we will not discuss this in anymore detail other than to highlight a few common mistakes / traps.
print('{0:.18f}'.format(0.1)) # Representation or approximation errors are common but small! Treat floats as noisy representations
0.1+0.1+0.1 == 0.3 # Because of this be careful using logical statements like == on floats!
tol = 1e-10 # If you do need to do logical checks then best to set and use a tolerance
abs(0.1+0.1+0.1 - 0.3) < tol
print('{0:.18f}'.format(0.1*999)) # Be careful, errors can accumulate!
# ## 3 ) Manipulating strings
# Declaring a string
s = 'Calculating Pythons'
# +
# Accessing entries of a string
# +
# Accessing every kth entry
# +
# Reversing order of a string
# +
# STRING METHODS
# capitalise / lower case / upper case
# count occurence of characters and phrases
# find the location of a string (first time appears)
# strip white spaces at the end
# split into a list
# -
# ## Questions?
| MM_material/lecture-materials/basic_objects/live_lectures/lecture-2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from bw2io.extractors import ExcelExtractor
import bw2data as bd
import re
assert "biosphere3" in bd.databases, "Must install base ecoinvent data"
bio = list(bd.Database("biosphere3"))
bio_names = {o['name'].lower() for o in bio}
len(bio), len(bio_names)
bio[0]._data
[x for x in bio_names if "fossil" in x]
data = ExcelExtractor.extract("ReCiPe2016_CFs_v1.1_20180117.xlsx")
# +
multiple = re.compile("^(.*)\((.*)\)$")
def get_names(dct):
"""Get all possible names for this elementary flow"""
def _(s):
return s.strip().lower()
if 'name' in dct:
yield _(dct['name'])
if 'Name' in dct:
yield _(dct['Name'])
if 'Alternative name (=name in emission database if different)' in dct:
match = multiple.match(dct['Alternative name (=name in emission database if different)'])
if match:
for name in match.groups():
yield _(name)
yield _(name).replace(" ", "-")
else:
yield _(dct['Alternative name (=name in emission database if different)'])
yield _(dct['Alternative name (=name in emission database if different)']).replace(" ", "-")
if 'Alternative name' in dct:
match = multiple.match(dct['Alternative name'])
if match:
for name in match.groups():
yield _(name)
yield _(name).replace(" ", "-")
else:
yield _(dct['Alternative name'])
yield _(dct['Alternative name']).replace(" ", "-")
get_names_test = {
'Name': 'Nitrous oxide',
'Alternative name (=name in emission database if different)': 'dinitrogen oxide (nitrous oxide)',
}
assert list(get_names(get_names_test)) == ['nitrous oxide', 'dinitrogen oxide', 'dinitrogen-oxide', 'nitrous oxide', 'nitrous-oxide']
# -
def three_row_header(data):
def _(s):
"""Clean up some minor typos.
Turns `(kg CO2eq/ kg GHG)` into `kg CO2eq/kg GHG`."""
if s.startswith("("):
s = s[1:]
if s.endswith(")"):
s = s[:-1]
return s.replace("/ ", "/")
try:
index_of_starting_cfs = data[2].index("Individualist")
except ValueError:
index_of_starting_cfs = data[2].index("I")
labels = data[2][:index_of_starting_cfs]
results = []
for column_index in range(index_of_starting_cfs, len(data[0])):
method = {
'name': data[0][column_index],
'unit': _(data[1][column_index]),
'perspective': data[2][column_index],
'cfs': []
}
for row in data[3:]:
if row[column_index] == '':
continue
method['cfs'].append(1)
results.append(method)
return method
category_formatter = {
'Global Warming': three_row_header,
'Stratospheric ozone depletion': three_row_header,
'Ionizing radiation': three_row_header,
'Human damage ozone formation': three_row_header,
}
def as_dict(data):
return {
label: category_formatter[label](rows)
for label, rows in data
if label in category_formatter
}
dd = as_dict(data)
def find_name_in_biosphere(cf, category_mapping):
_ = lambda s: category_mapping.get(s, s)
for name in get_names(cf):
name = _(name)
if name in bio_names:
return name
# Try a partial match
name = _(cf['Name'].lower())
for o in bio_names:
if name and name in o:
return o
# # Global warming
category_mapping = {
'carbon dioxide': 'carbon dioxide, fossil',
'carbon monoxide': 'carbon monoxide, fossil',
'methane': 'methane, non-fossil',
'fossil methane': 'methane, fossil',
'nitrous oxide': 'dinitrogen monoxide',
'sulphur hexafluoride': 'sulfur hexafluoride',
'halon-1211': 'methane, bromochlorodifluoro-, halon 1211',
'pfc-41-12': 'perfluoropentane',
'cfc-11': 'methane, trichlorofluoro-, cfc-11',
'pfc-116': 'ethane, hexafluoro-, hfc-116',
'halon-1301': 'methane, bromotrifluoro-, halon 1301',
'methylene chloride': 'methane, dichloro-, hcc-30',
'methyl chloride': 'methane, monochloro-, r-40',
'carbon tetrachloride': 'methane, tetrachloro-, r-10',
'pfc-14': 'methane, tetrafluoro-, r-14',
'cfc-12': 'methane, dichlorodifluoro-, cfc-12',
'methyl bromide': 'methane, bromo-, halon 1001',
}
found = {find_name_in_biosphere(cf, category_mapping)
for cf in dd['Global Warming']['cfs']
if find_name_in_biosphere(cf, category_mapping)}
found
in_ecoinvent_ipcc = {bd.get_activity(o[0])['name'].lower() for o in bd.Method(('IPCC 2013', 'climate change', 'GWP 100a')).load()}
in_ecoinvent_recipe = {bd.get_activity(o[0])['name'].lower() for o in bd.Method(('ReCiPe Midpoint (E) V1.13', 'climate change', 'GWP500')).load()}
known_missing = {
'ethane, 1,1,1-trichloro-, hcfc-140',
'carbon monoxide, fossil',
'carbon monoxide, from soil or biomass stock',
'carbon monoxide, non-fossil',
'methane, bromo-, halon 1001',
'nitric oxide',
'nitrogen fluoride',
'voc, volatile organic compounds, unspecified origin', # What could the CF even be?
'ethane, 1,2-dichloro-',
'dimethyl ether',
}
in_ecoinvent_ipcc.difference(found).difference(known_missing)
in_ecoinvent_recipe.difference(found).difference(known_missing)
found.difference(in_ecoinvent_recipe)
found.difference(in_ecoinvent_ipcc)
# # Ozone depletion
category_mapping = {
'carbon tetrachloride': 'methane, tetrachloro-, r-10',
'cfc-12': 'methane, dichlorodifluoro-, cfc-12',
'cfc-11': 'methane, trichlorofluoro-, cfc-11',
'halon-1301': 'methane, bromotrifluoro-, halon 1301',
'halon-1211': 'methane, bromochlorodifluoro-, halon 1211',
'ch3cl': 'methane, monochloro-, r-40',
'ch3br': 'methane, bromo-, halon 1001',
'n2o': 'dinitrogen oxide',
}
found = {find_name_in_biosphere(cf, category_mapping)
for cf in dd['Stratospheric ozone depletion']['cfs']
if find_name_in_biosphere(cf, category_mapping)}
found
missing = {cf['Name']
for cf in dd['Stratospheric ozone depletion']['cfs']
if not find_name_in_biosphere(cf, category_mapping)}
missing
in_ecoinvent = {bd.get_activity(o[0])['name'].lower() for o in bd.Method(('ReCiPe Midpoint (E) V1.13', 'ozone depletion', 'ODPinf')).load()}
known_missing = {
'ethane, 1,1,1-trichloro-, hcfc-140',
'ethane, 1,1,2-trichloro-',
}
in_ecoinvent.difference(found).difference(known_missing)
found.difference(in_ecoinvent)
# # Ionizing radiation
category_mapping = {
'ch3cl': 'methane, monochloro-, r-40',
'ch3br': 'methane, bromo-, halon 1001',
'actinides, unspecified': 'actinides, radioactive, unspecified',
'cs-134': 'cesium-134',
'cs-137': 'cesium-137',
'co-58': 'cobalt-58',
'h-3': 'hydrogen-3, tritium',
'cm alphaa': 'curium alpha',
'pb-210': 'lead-210',
'pu-238': 'plutonium-238',
'pu-239': 'plutonium-239',
'mn-54': 'manganese-54',
'ra-226a': 'radium-226',
'ra-226': 'radium-226',
'sb-124': 'antimony-124',
'i-133': 'iodine-133',
'po-210': 'polonium-210',
'ag-110m': 'silver-110',
'pu alpha': 'plutonium-alpha',
'pu alphaa': 'plutonium-alpha',
'u-238a': 'uranium-238',
'xe-133': 'xenon-133',
# What ecoinvent calls 'plutonium-alpha' could be Pu 239?
}
found = {find_name_in_biosphere(cf, category_mapping)
for cf in dd['Ionizing radiation']['cfs']
if find_name_in_biosphere(cf, category_mapping)}
found
in_ecoinvent = {bd.get_activity(o[0])['name'].lower()
for o in bd.Method(('ReCiPe Midpoint (E) V1.13', 'ionising radiation', 'IRP_HE')).load()}
known_missing = {
'uranium alpha',
}
in_ecoinvent.difference(found).difference(known_missing)
found.difference(in_ecoinvent)
missing = {cf['Name']
for cf in dd['Ionizing radiation']['cfs']
if not find_name_in_biosphere(cf, category_mapping)}
missing
# # Human damage ozone formation
category_mapping = {
'ch3cl': 'methane, monochloro-, r-40',
# 'ch3br': 'methane, bromo-, halon 1001',
# 'actinides, unspecified': 'actinides, radioactive, unspecified',
# 'cs-134': 'cesium-134',
# 'cs-137': 'cesium-137',
# 'co-58': 'cobalt-58',
# 'h-3': 'hydrogen-3, tritium',
# 'cm alphaa': 'curium alpha',
# 'pb-210': 'lead-210',
# 'pu-238': 'plutonium-238',
# 'pu-239': 'plutonium-239',
# 'mn-54': 'manganese-54',
# 'ra-226a': 'radium-226',
# 'ra-226': 'radium-226',
# 'sb-124': 'antimony-124',
# 'i-133': 'iodine-133',
# 'po-210': 'polonium-210',
# 'ag-110m': 'silver-110',
# 'pu alpha': 'plutonium-alpha',
# 'pu alphaa': 'plutonium-alpha',
# 'u-238a': 'uranium-238',
# 'xe-133': 'xenon-133',
}
found = {find_name_in_biosphere(cf, category_mapping)
for cf in dd['Human damage ozone formation']['cfs']
if find_name_in_biosphere(cf, category_mapping)}
found
in_ecoinvent = {bd.get_activity(o[0])['name'].lower()
for o in bd.Method(('ReCiPe Midpoint (E) V1.13', 'photochemical oxidant formation', 'POFP')).load()}
known_missing = {
'uranium alpha',
}
in_ecoinvent.difference(found).difference(known_missing)
found.difference(in_ecoinvent)
missing = {cf['Name']
for cf in dd['Human damage ozone formation']['cfs']
if not find_name_in_biosphere(cf, category_mapping)}
missing
s = "pluton"
{x for x in bio_names if s in x}
sorted([m for m in bd.methods if m[0] == 'ReCiPe Midpoint (E) V1.13'])
| dev/Import original CFs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Assignment
# Must make use of ratings, merge with another DF. Answer an interesting question about the Muppets!
#
# ## Question
#
# **How long is the average film career before appearing in the muppets?**
#
# **Does the Muppet Show make or break a film career?**
#
# Criteria:
# 1. Guest star on the Muppet Show
# 2. Actor / actress
# 3. First film is >5 years prior
# 4. Last film is >5 years after
#
# _"That's a high light in my career, going to England to film the Muppet show. Working with <NAME> was great, he was a genius."_
# <NAME>
#
# ## Data Wrangling
# +
import numpy as np
import pandas as pd
import gzip
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
def createCodebookTable(df):
df_codebook = pd.DataFrame(index=df.columns, columns=["dtype","type","min","max","count","missing"])
df_codebook["dtype"] = [str(df[x].dtype) for x in df.columns]
df_codebook["type"] = [str(type(df[x][0])) for x in df.columns]
df_codebook["min"] = df.min()
df_codebook["max"] = df.max()
df_codebook["count"] = df.count()
df_codebook["missing"] = df.isnull().sum()
return df_codebook
def getData(df):
'''Function to create a dataframe with the timeDelta (t-5, t-4,...t+5) as the index and the average rating, num. of episodes and the nconst as columns
Parameters: dataframe
Returns: dataframe
'''
data=[]
for i in df['nconst'].unique():
df2=df[df['nconst']==i]
pivot=pd.pivot_table(df2,index='timeDelta', values=['tconst', 'averageRating','nconst'],
aggfunc={'tconst':'count', 'averageRating':'mean','nconst':'first'})
data.append(pivot)
return(pd.concat(data))
def getTstats(df):
'''Function to create a dataframe with the t-stat and p-value of a difference of mean before and after muppets for each nconst
Parameters: dataframe
Returns: dataframe
'''
unique_nconst=df['nconst'].unique()
results=pd.DataFrame(columns=['tstat_average','pval_average','tstat_tconst','pval_tconst'],index=unique_nconst)
for nconst in unique_nconst:
df_filter=df[df['nconst']==nconst]
after_muppet=df_filter.query('timeDelta>0')
before_muppet=df_filter.query('timeDelta<0')
results.loc[nconst,'tstat_average']=stats.ttest_ind(after_muppet['averageRating'],before_muppet['averageRating']).statistic
results.loc[nconst,'pval_average']=stats.ttest_ind(after_muppet['averageRating'],before_muppet['averageRating']).pvalue
results.loc[nconst,'tstat_tconst']=stats.ttest_ind(after_muppet['tconst'],before_muppet['tconst']).statistic
results.loc[nconst,'pval_tconst']=stats.ttest_ind(after_muppet['tconst'],before_muppet['tconst']).pvalue
return(results)
# +
# find the title tag (tconst) for The Muppet Show, as a reference for searching all other .tsv files
show_name = 'The Muppet Show'
show_type = 'tvSeries'
with gzip.open('title.basics.tsv.gz') as file :
titles_df = pd.read_csv(file, dtype= str, sep='\t') # set dtype to str so no wasted computing on guessing type
mup_row = titles_df[ (titles_df['primaryTitle'] == show_name ) & (titles_df['titleType'] == show_type ) ]
# store title tag for later use
mup_tconst = list(mup_row['tconst'])[0]
# Get list of episodes for the Muppet Show
with gzip.open('title.episode.tsv.gz') as file :
epi_df = pd.read_csv(file, dtype= str, sep='\t') # set dtype to str so no wasted computing on guessing type
epi_df = epi_df[ epi_df[ "parentTconst" ] == mup_tconst ] # return new dataframe which only has muppet episodes
epi_info_df = titles_df[ titles_df['tconst'].isin(epi_df['tconst'])]
# Create a list of each guest star in Muppet Show... start with the nconst identifier
with gzip.open('title.principals.tsv.gz') as file :
prin_df = pd.read_csv(file, dtype = str, sep = '\t')
prin_df = prin_df[ prin_df['tconst'].isin(epi_df['tconst']) ] # filter for the titles in Muppet Show and for actors
# filter out only the principals who are in category of 'self'
nconst_df = prin_df[ prin_df['category'] == 'self']
# Subset the name.basics.tsv file to include only the guest stars!
with gzip.open('name.basics.tsv.gz') as file :
guest_df = pd.read_csv(file, dtype = str, sep = '\t')
guest_df = guest_df[ guest_df['nconst'].isin(nconst_df['nconst'])]
# remove any guest that is not an individual (e.g. a band)
guest_df = guest_df[ pd.to_numeric(guest_df['birthYear'], errors = 'coerce') > 0 ]
# +
# need some way of matching nconst --> tconst --> year Aired... so that we have a year of Muppet Appearance for each guest
# create a series of nconsts, yearAired
guest2_df = guest_df.copy()
guest2_df = guest2_df.set_index("nconst")
for index, row in guest_df.iterrows():
nconst = row['nconst']
tconst = list(nconst_df[ nconst_df['nconst'] == nconst ]['tconst'])[0]
year = list(epi_info_df[ epi_info_df['tconst'] == tconst ]['startYear'])[0]
guest2_df.loc[nconst,'yearOnMShow'] = year #find some way of assigning startYear to our guestdf!
with gzip.open('title.principals.tsv.gz') as file :
shows_df = pd.read_csv(file, dtype = str, sep = '\t')
shows_df = shows_df[ shows_df['nconst'].isin(guest2_df.index) ] # filter for the titles for actors
# +
# Copying to be safe!
shows2_df = shows_df.copy()
shows2_df = shows2_df.merge(guest2_df,left_on="nconst",right_index=True)
shows3_df = shows2_df.loc[:,["tconst","nconst","characters","primaryName","yearOnMShow"]]
with gzip.open('title.basics.tsv.gz') as file :
titles_guests_df = pd.read_csv(file, dtype= str, sep='\t')
titles_guests_df = titles_guests_df[ titles_guests_df['tconst'].isin(shows3_df['tconst']) ]
# Copying again to be safe!
shows4_df = shows3_df.copy()
shows4_df = shows4_df.merge(titles_guests_df, on = "tconst")
shows4_df = shows4_df.loc[:,["tconst","nconst","characters","primaryName","yearOnMShow","startYear"]]
# open tsv file and load into a dataframe
with gzip.open('title.ratings.tsv.gz') as ratings_file :
ratings_df = pd.read_csv(ratings_file,sep='\t')
ratings_df = ratings_df[ ratings_df['tconst'].isin(shows4_df['tconst']) ]
shows5_df = shows4_df.copy()
shows5_df = shows5_df.merge(ratings_df, on = "tconst")
# Some additional cleaning!
shows5_df['startYear'] = shows5_df['startYear'].map(lambda x: int(x))
shows5_df['yearOnMShow'] = shows5_df['yearOnMShow'].map(lambda x: int(x))
shows5_df['timeDelta'] = shows5_df['startYear'] - shows5_df['yearOnMShow']
display(shows5_df.head())
# The codebook
createCodebookTable(shows5_df)
# -
# ## Data Analysis
# +
# Selecting the relevant time for your analysis
relevant_time = [-5,-4,-3,-2,-1,0,1,2,3,4,5]
shows_all = shows5_df[shows5_df['timeDelta'].isin(relevant_time)]
display(shows_all.head())
# -
shows_all.describe()
# Nothing particularly spectacular at this point
# +
# Creating sorts by groups to see any info that can be extracted from the tables
# Also using the shows all table kept crashing my jupiters so using this filtered dataframe for the plot
showscount_plot = pd.DataFrame(shows_all.groupby(['timeDelta'])['tconst'].nunique())
showsrating_plot = pd.DataFrame(shows_all.groupby(['timeDelta'])['averageRating'].mean())
showscount_plot = showscount_plot.reset_index()
showsrating_plot = showsrating_plot.reset_index()
display(showscount_plot)
display(showsrating_plot)
# It looks like both the means and average ratings were falling on average for all the guest stars - sad times!
# +
# A visualisation to drive the point home!
plt.subplot(131)
plt.bar(showscount_plot.timeDelta, showscount_plot.tconst, color='c')
plt.xlabel('Time Delta')
plt.ylabel('Movies Count')
plt.subplot(132)
plt.bar(showsrating_plot.timeDelta, showsrating_plot.averageRating,color='m')
plt.xlabel('Time Delta')
plt.ylabel('Rating')
plt.subplots_adjust(top=5, bottom=3.5, left=0, right=5, hspace=2.5, wspace=0.25)
# +
# Another visualisation to drive the point home! From the above chart, it could be surmised that the year the stars
# appeared on the show was their most popular year, after which they went into decline. These are sum totals though.
plt.subplot(131)
plt.plot(showscount_plot.timeDelta, showscount_plot.tconst, color='c')
plt.xlabel('Time Delta')
plt.ylabel('Movies Count')
plt.subplot(132)
plt.plot(showsrating_plot.timeDelta, showsrating_plot.averageRating, color='m')
plt.xlabel('Time Delta')
plt.ylabel('Rating')
plt.subplots_adjust(top=5, bottom=3.5, left=0, right=5, hspace=2.5, wspace=0.25)
# +
# Now combining the average ratings and number of appearances over time.
# Appearances are represented by the size of the plot
plt.subplot(132)
plt.scatter(x = showsrating_plot.timeDelta, y = showsrating_plot.averageRating, s = (showscount_plot.tconst**1.5), color='m')
plt.xlabel('Time Delta')
plt.ylabel('Rating')
plt.grid(True)
plt.subplots_adjust(top=5, bottom=3, left=0, right=10, hspace=5, wspace=0.5)
plt.show()
# +
# # Some other questions potentially worth answering:
# Who suffered the most from MuppetGate? Are they actors/actresses that have lost their careers?
# How do the averages differ for the guest stars? So far all our analysis has been on the sums
# Any other interesting questions?
# -
flo_shows = shows_all[shows_all['primaryName'] == '<NAME>']
flo_shows = flo_shows.sort_values(by = 'timeDelta')
display(flo_shows.head())
# +
# display(flo_shows.groupby(['timeDelta'])['tconst'].nunique())
flocount_plot = pd.DataFrame(flo_shows.groupby(['timeDelta'])['tconst'].nunique())
florating_plot = pd.DataFrame(flo_shows.groupby(['timeDelta'])['averageRating'].mean())
flocount_plot = flocount_plot.reset_index()
florating_plot = florating_plot.reset_index()
# +
plt.subplot(131)
plt.plot(flocount_plot.timeDelta, flocount_plot.tconst, color='c')
plt.xlabel('Time Delta')
plt.ylabel('Movies Count')
plt.subplot(132)
plt.plot(florating_plot.timeDelta, florating_plot.averageRating, color='m')
plt.xlabel('Time Delta')
plt.ylabel('Rating')
plt.subplots_adjust(top=5, bottom=3.5, left=0, right=5, hspace=2.5, wspace=0.25)
# -
display(florating_plot)
display(flo_shows[flo_shows['timeDelta'] == 1])
| Week 4/PySDS_Ex_Week4_Day01_AA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
import os
sys.path.append(os.path.abspath("../"))
sys.path.append(os.path.abspath("../AdventOfCode 2020/"))
from utils import *
from aoc import *
from personal import SESSION
from datetime import datetime
import numpy as np
aoc = AOC(session=SESSION)
aoc.verify_session()
st = datetime.now()
data = aoc.get_file(year=2019, day=3).analyse().head().data
# ## First part
def gen_path(instructions):
path = [np.array([0, 0])]
for instruction in instructions:
if instruction[0] == 'R':
moving = np.array([1, 0])
elif instruction[0] == 'L':
moving = np.array([-1, 0])
elif instruction[0] == 'U':
moving = np.array([0, 1])
else:
moving = np.array([0, -1])
for i in range(int(instruction[1:])):
path.append(path[-1]+moving)
return path
p1 = set([tuple(el) for el in gen_path(data[0].split(','))])
p2 =set([tuple(el) for el in gen_path(data[1].split(','))])
inter = p1 & p2
sorted([sum(map(abs, inter)) for inter in inter])[:5]
e1 = datetime.now()
# ## Second part
p1 = [tuple(el) for el in gen_path(data[0].split(','))]
p2 = [tuple(el) for el in gen_path(data[1].split(','))]
vals = []
for cross in inter:
vals.append(p1.index(cross) + p2.index(cross))
sorted(vals)[:5]
e2 = datetime.now()
print(f'Première partie: {e1-st!s}')
print(f'Seconde partie: {e2-e1!s}')
| AdventOfCode 2019/AOC.3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
if 0 :
# %matplotlib inline
else :
# %matplotlib notebook
# # Import libraries
# +
import sys
import os
module_path = os.path.abspath('.') +"\\_scripts"
print(module_path)
if module_path not in sys.path:
sys.path.append(module_path)
from _00_Import_packages_git3 import *
# -
from numpy import array
import pandas as pd
from sos_trades_core.execution_engine.execution_engine import ExecutionEngine
from numpy.testing import assert_array_equal, assert_array_almost_equal # @UnresolvedImport
import os
from gemseo.core.mdo_scenario import MDOScenario
# # TestSoSOptimScenario
# # setUp
study_name = 'optim'
ns = f'{study_name}'
sc_name = "SellarOptimScenario"
c_name = "SellarCoupling"
print(sc_name)
ns
from tempfile import gettempdir
root_dir = gettempdir()
print(root_dir)
#dspace.to_csv(root_dir+'\\ds.csv')
repo = 'sos_trades_core.sos_processes.test'
proc_name = 'test_sellar_opt'
# # 3 optim scenario execution disciplinaryopt
print("\n Test 4 : Sellar optim solution check with DisciplinaryOpt formulation")
exec_eng = ExecutionEngine(study_name)
factory = exec_eng.factory
repo_discopt = 'sos_trades_core.sos_processes.test'
proc_name_discopt = 'test_sellar_opt_discopt'
print(repo_discopt + "\\" + proc_name_discopt)
builder = factory.get_builder_from_process(repo=repo_discopt, mod_id=proc_name_discopt)
exec_eng.factory.set_builders_to_coupling_builder(builder)
exec_eng.configure()
print('\n in test optim scenario')
for key in exec_eng.dm.data_id_map:
print("key", key)
#-- set up design space
dspace_dict = {'variable': ['x', 'z'],
'value': [[1.], [5., 2.]],
'lower_bnd': [[0.], [-10., 0.]],
'upper_bnd': [[10.], [10., 10.]],
'enable_variable': [True, True],
'activated_elem': [[True], [True, True]]}
dspace = pd.DataFrame(dspace_dict)
dspace
# +
#-- set up disciplines in Scenario
disc_dict = {}
# Optim inputs
disc_dict[f'{ns}.SellarOptimScenario.max_iter'] = 200
# SLSQP, NLOPT_SLSQP
disc_dict[f'{ns}.SellarOptimScenario.algo'] = "SLSQP" #NLOPT_SLSQP
disc_dict[f'{ns}.SellarOptimScenario.design_space'] = dspace
disc_dict[f'{ns}.SellarOptimScenario.formulation'] = 'DisciplinaryOpt'
disc_dict[f'{ns}.SellarOptimScenario.objective_name'] = 'obj'
disc_dict[f'{ns}.SellarOptimScenario.ineq_constraints'] = [
'c_1', 'c_2']
disc_dict[f'{ns}.SellarOptimScenario.algo_options'] = {"ftol_rel": 1e-6,
"ineq_tolerance": 1e-6,
"normalize_design_space": True}
disc_dict
# -
#exec_eng.dm.set_values_from_dict(disc_dict)
exec_eng.load_study_from_input_dict(disc_dict)
# Sellar inputs
local_dv = 10.
values_dict = {}
# array([1.])
values_dict[f'{ns}.{sc_name}.{c_name}.x'] = 1.
values_dict[f'{ns}.{sc_name}.{c_name}.y_1'] = 1.
values_dict[f'{ns}.{sc_name}.{c_name}.y_2'] = 1.
values_dict[f'{ns}.{sc_name}.{c_name}.z'] = array([1., 1.])
values_dict[f'{ns}.{sc_name}.{c_name}.Sellar_Problem.local_dv'] = local_dv
values_dict
#exec_eng.dm.set_values_from_dict(values_dict)
exec_eng.load_study_from_input_dict(values_dict)
exec_eng.configure()
exp_tv_list = [f'Nodes representation for Treeview {ns}',
'|_ optim',
f'\t|_ {sc_name}',
f'\t\t|_ {c_name}',
'\t\t\t|_ Sellar_2',
'\t\t\t|_ Sellar_1',
'\t\t\t|_ Sellar_Problem']
exp_tv_list
exp_tv_str = '\n'.join(exp_tv_list)
exp_tv_str
exec_eng.display_treeview_nodes(True)
exec_eng.display_treeview_nodes()
res = exec_eng.execute()
res
# retrieve discipline to check the result...
opt_disc = exec_eng.dm.get_disciplines_with_name("optim." + sc_name)[0]
# check optimal x and f
sellar_obj_opt = 3.18339 + local_dv
sellar_obj_opt
opt_disc.optimization_result.f_opt
exp_x = array([8.3109e-15, 1.9776e+00, 3.2586e-13])
exp_x
opt_disc.optimization_result.x_opt
| sos_trades_core/tests/jupyter_doc/ipynb/ex_04.3_optim_scenario.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: u4-s3-dnn
# kernelspec:
# display_name: U4-S1-NLP (Python3)
# language: python
# name: u4-s1-nlp
# ---
# <img align="left" src="https://lever-client-logos.s3.amazonaws.com/864372b1-534c-480e-acd5-9711f850815c-1524247202159.png" width=200>
# <br></br>
# <br></br>
#
# # Major Neural Network Architectures Challenge
# ## *Data Science Unit 4 Sprint 3 Challenge*
#
# In this sprint challenge, you'll explore some of the cutting edge of Data Science. This week we studied several famous neural network architectures:
# recurrent neural networks (RNNs), long short-term memory (LSTMs), convolutional neural networks (CNNs), and Autoencoders. In this sprint challenge, you will revisit these models. Remember, we are testing your knowledge of these architectures not your ability to fit a model with high accuracy.
#
# __*Caution:*__ these approaches can be pretty heavy computationally. All problems were designed so that you should be able to achieve results within at most 5-10 minutes of runtime locally, on AWS SageMaker, on Colab or on a comparable environment. If something is running longer, double check your approach!
#
# ## Challenge Objectives
# *You should be able to:*
# * <a href="#p1">Part 1</a>: Train a LSTM classification model
# * <a href="#p2">Part 2</a>: Utilize a pre-trained CNN for object detection
# * <a href="#p3">Part 3</a>: Describe a use case for an autoencoder
# * <a href="#p4">Part 4</a>: Describe yourself as a Data Science and elucidate your vision of AI
# + [markdown] colab_type="text" id="-5UwGRnJOmD4"
# <a id="p1"></a>
# ## Part 1 - LSTMSs
#
# Use a LSTM to fit a multi-class classification model on Reuters news articles to distinguish topics of articles. The data is already encoded properly for use in a LSTM model.
#
# Your Tasks:
# - Use Keras to fit a predictive model, classifying news articles into topics.
# - Report your overall score and accuracy
#
# For reference, the [Keras IMDB sentiment classification example](https://github.com/keras-team/keras/blob/master/examples/imdb_lstm.py) will be useful, as well as the LSTM code we used in class.
#
# __*Note:*__ Focus on getting a running model, not on maxing accuracy with extreme data size or epoch numbers. Only revisit and push accuracy if you get everything else done!
# -
# + colab={"base_uri": "https://localhost:8080/", "height": 1114} colab_type="code" id="DS-9ksWjoJit" outputId="0c3512e4-5cd4-4dc6-9cda-baf00c835f59"
from tensorflow.keras.datasets import reuters
(X_train, y_train), (X_test, y_test) = reuters.load_data(num_words=None,
skip_top=0,
maxlen=None,
test_split=0.2,
seed=723812,
start_char=1,
oov_char=2,
index_from=3)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="fLKqFh8DovaN" outputId="64b0d621-7e74-4181-9116-406e8c518465"
# Demo of encoding
word_index = reuters.get_word_index(path="reuters_word_index.json")
print(f"Iran is encoded as {word_index['iran']} in the data")
print(f"London is encoded as {word_index['london']} in the data")
print("Words are encoded as numbers in our dataset.")
# + colab={} colab_type="code" id="_QVSlFEAqWJM"
# Do not change this line. You need the +1 for some reason.
max_features = len(word_index.values()) + 1
# TODO - your code!
# + [markdown] nteract={"transient": {"deleting": false}}
# ## Sequence Data Question
# #### *Describe the `pad_sequences` method used on the training dataset. What does it do? Why do you need it?*
#
#
#
# ## RNNs versus LSTMs
# #### *What are the primary motivations behind using Long-ShortTerm Memory Cell unit over traditional Recurrent Neural Networks?*
#
#
#
# ## RNN / LSTM Use Cases
# #### *Name and Describe 3 Use Cases of LSTMs or RNNs and why they are suited to that use case*
#
# + [markdown] colab_type="text" id="yz0LCZd_O4IG"
# <a id="p2"></a>
# ## Part 2- CNNs
#
# ### Find the Frog
#
# Time to play "find the frog!" Use Keras and ResNet50 (pre-trained) to detect which of the following images contain frogs:
#
# <img align="left" src="https://d3i6fh83elv35t.cloudfront.net/newshour/app/uploads/2017/03/GettyImages-654745934-1024x687.jpg" width=400>
# + colab={"base_uri": "https://localhost:8080/", "height": 245} colab_type="code" id="whIqEWR236Af" outputId="7a74e30d-310d-4a3a-9ae4-5bf52d137bda"
from skimage.io import imread_collection
from skimage.transform import resize #This might be a helpful function for you
images = imread_collection('./frog_images/*.jpg')
# + colab={"base_uri": "https://localhost:8080/", "height": 332} colab_type="code" id="EKnnnM8k38sN" outputId="59f477e9-0b25-4a38-9678-af24e0176535"
print(type(images))
print(type(images[0]), end="\n\n")
print("Each of the Images is a Different Size")
print(images[0].shape)
print(images[1].shape)
# + [markdown] colab_type="text" id="si5YfNqS50QU"
# Your goal is to validly run ResNet50 on the input images - don't worry about tuning or improving the model. Print out the predictions in any way you see fit.
#
# *Hint* - ResNet 50 doesn't just return "frog". The three labels it has for frogs are: `bullfrog, tree frog, tailed frog`
#
# *Stretch goal* - Check for other things such as fish.
# + colab={} colab_type="code" id="FaT07ddW3nHz"
# TODO - your code!
# + [markdown] colab_type="text" id="XEuhvSu7O5Rf"
# <a id="p3"></a>
# ## Part 3 - Autoencoders
#
# Describe a use case for an autoencoder given that an autoencoder tries to predict its own input.
#
# __*Your Answer:*__
#
# + [markdown] colab_type="text" id="626zYgjkO7Vq"
# <a id="p4"></a>
# ## Part 4 - More...
# + [markdown] colab_type="text" id="__lDWfcUO8oo"
# Answer the following questions, with a target audience of a fellow Data Scientist:
#
# - What do you consider your strongest area, as a Data Scientist?
# - What area of Data Science would you most like to learn more about, and why?
# - Where do you think Data Science will be in 5 years?
# - What are the threats posed by AI to our society?
# - How do you think we can counteract those threats?
# - Do you think achieving General Artifical Intelligence is ever possible?
#
# A few sentences per answer is fine - only elaborate if time allows.
# + [markdown] colab_type="text" id="_Hoqe3mM_Mtc"
# ## Congratulations!
#
# Thank you for your hard work, and congratulations! You've learned a lot, and you should proudly call yourself a Data Scientist.
#
# +
from IPython.display import HTML
HTML("""<iframe src="https://giphy.com/embed/26xivLqkv86uJzqWk" width="480" height="270" frameBorder="0" class="giphy-embed" allowFullScreen></iframe><p><a href="https://giphy.com/gifs/mumm-champagne-saber-26xivLqkv86uJzqWk">via GIPHY</a></p>""")
# -
| LS_DS_Unit_4_Sprint_Challenge_3_v2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import warnings
warnings.filterwarnings('ignore')
import os
import pathlib
import matplotlib.pyplot as plt
import numpy as np
from landlab.components import OverlandFlow, FlowAccumulator, SpatialPrecipitationDistribution, FlowDirectorSteepest, NetworkSedimentTransporter
from landlab.data_record import DataRecord
from landlab.grid.network import NetworkModelGrid
from landlab.plot import graph
from landlab.io import read_shapefile
from landlab import ExampleData
from landlab.plot.imshow import imshow_grid, imshow_grid_at_node
from landlab.io.esri_ascii import read_esri_ascii
from landlab.plot import plot_network_and_parcels
# %matplotlib inline
import matplotlib.colors as mcolors
colors = [(0,0.2,1,i) for i in np.linspace(0,1,3)]
cmap = mcolors.LinearSegmentedColormap.from_list('mycmap', colors, N=10)
# -
# Load shapefile of the drainage network
# +
shp_file = '../data/hugo_links.shp'
points_shapefile = '../hugo_nodes_nodublicates.shp'
grid = read_shapefile(
shp_file,
threshold=0.1,
)
# -
# Plot the Network
graph.plot_graph(grid, at="node,link")
# #### Make a new grid that doesn't have duplicated nodes
# * Nodes we don't want: 19, 13
# * reassing all nodes_at_link that refensece the nodes we don't want 19-> 20, 12->13
# +
#Drop double nodes
drop_nodes = [19,13]
new_y_of_node =[]
new_x_of_node =[]
#Find the new x and y coordinates
for i, node in enumerate(grid.y_of_node):
if i not in drop_nodes:
new_y_of_node.append(node)
for i, node in enumerate(grid.x_of_node):
if i not in drop_nodes:
new_x_of_node.append(node)
#rediect all nodes
replace_nodes = [20,12]
nodes_reformatted=np.ravel(grid.nodes_at_link)
for node, replacement in zip(drop_nodes,replace_nodes):
nodes_reformatted[np.where(nodes_reformatted==node)]=replacement
#change index number
for node in drop_nodes:
nodes_reformatted[np.where(nodes_reformatted >node)] -=1
new_nodes_at_link=nodes_reformatted.reshape([21,2])
# Make a new network model grid
new_grid = NetworkModelGrid((new_y_of_node, new_x_of_node), new_nodes_at_link)
# -
graph.plot_graph(new_grid, at="node,link")
# ### Setup the overland flow
#Define grid
# here we use an arbitrary, very small, "real" catchment
fname = '../data/hugo_site.asc'
rmg, z = read_esri_ascii(fname, name='topographic__elevation')
# rmg.status_at_node[rmg.nodes_at_right_edge] = rmg.BC_NODE_IS_FIXED_VALUE
rmg.status_at_node[np.isclose(z, -9999.)] = rmg.BC_NODE_IS_CLOSED
rmg.set_watershed_boundary_condition('topographic__elevation')
# +
# Initial conditions
run_time =50000# 1 day 259200 # duration of run, (s) equivalent of 3 days
n = 0.01 # roughness coefficient, (s/m^(1/3))
g = 9.8 # gravity (m/s^2)
alpha = 0.7 # time-step factor (nondimensional; from Bates et al., 2010)
u = 0.4 # constant velocity (m/s, de Almeida et al., 2012)
elapsed_time = 1.0 #Elapsed time starts at 1 second. This prevents errors when setting our boundary conditions.
h_init = 1 # initial thin layer of water (m)
#Set up rain
rmg.at_node["surface_water__depth"] = np.zeros(rmg.number_of_nodes)
h = rmg.at_node['surface_water__depth']
h+=h_init #add a little bit of water everywhere
#ad a storm
bools = (rmg.node_y > 100) * (rmg.node_y < 450) * (rmg.node_x < 400) * (rmg.node_x > 200)
h[bools] = 1 #initial water depth m
## Set inital discharge
rmg.at_node["surface_water__discharge"] = np.zeros(rmg.number_of_nodes)
# -
# ### Start the sediment routing part
# +
nmg_node_topo = np.zeros(new_grid.number_of_nodes) #create empty array holding elevations for network model grid
nmg_node_h = np.zeros(new_grid.number_of_nodes)
for i in range(new_grid.number_of_nodes):
idx_nearest_raster_cell = rmg.find_nearest_node([new_grid.x_of_node[i], new_grid.y_of_node[i]])
nmg_node_topo[i] = rmg.at_node["topographic__elevation"][idx_nearest_raster_cell]
# -
# Calculate distance between links
# +
nmg_link_length = np.zeros(new_grid.number_of_links)
nmg_link_h = np.zeros(new_grid.number_of_links)
for i in range(new_grid.number_of_links):
idx_node_1_link = new_grid.nodes_at_link[i][0]
idx_node_2_link = new_grid.nodes_at_link[i][1]
nmg_link_length[i] = np.sqrt((new_grid.x_of_node[idx_node_2_link] - new_grid.x_of_node[idx_node_1_link])**2
+ (new_grid.y_of_node[idx_node_2_link] - new_grid.y_of_node[idx_node_1_link])**2)
square_idx = rmg.find_nearest_node([new_grid.x_of_node[idx_node_1_link], new_grid.y_of_node[idx_node_1_link]]) #get raster grid idx closest to link tail
nmg_link_h[i] = rmg.at_node['surface_water__depth'][square_idx] #assign surface at square_idx to network grid
# +
new_grid.at_node["topographic__elevation"] = nmg_node_topo.copy()
new_grid.at_node["bedrock__elevation"] = nmg_node_topo.copy()
new_grid.at_link["channel_width"] = 10 * np.ones(grid.number_of_links)
new_grid.at_link["flow_depth"] = nmg_link_h.copy()
new_grid.at_link["reach_length"] = nmg_link_length
# -
# ### Create a sediment parcels
# +
# element_id is the link on which the parcel begins.
element_id = np.repeat(np.arange(new_grid.number_of_links), 50)
element_id = np.expand_dims(element_id, axis=1)
volume = 1*np.ones(np.shape(element_id)) # (m3)
active_layer = np.ones(np.shape(element_id)) # 1= active, 0 = inactive
density = 2650 * np.ones(np.size(element_id)) # (kg/m3)
abrasion_rate = 0 * np.ones(np.size(element_id)) # (mass loss /m)
# Lognormal GSD
medianD = 0.002 # m
mu = np.log(medianD)
sigma = np.log(2) #assume that D84 = sigma*D50
np.random.seed(0)
D = np.random.lognormal(
mu,
sigma,
np.shape(element_id)
) # (m) the diameter of grains in each parcel
# -
# We begin by assigning each parcel an arbitrary (and small) arrival time and location in the link.
time_arrival_in_link = np.random.rand(np.size(element_id), 1)
location_in_link = np.random.rand(np.size(element_id), 1)
# We now collect the arrays into a dictionary of variables, some of which will be tracked through time (`["item_id", "time"]`), and others of which will remain constant through time :
# +
lithology = ["quartzite"] * np.size(element_id)
variables = {
"abrasion_rate": (["item_id"], abrasion_rate),
"density": (["item_id"], density),
"lithology": (["item_id"], lithology),
"time_arrival_in_link": (["item_id", "time"], time_arrival_in_link),
"active_layer": (["item_id", "time"], active_layer),
"location_in_link": (["item_id", "time"], location_in_link),
"D": (["item_id", "time"], D),
"volume": (["item_id", "time"], volume)
}
# -
# With all of the required attributes collected, we can create the parcels DataRecord. Often, parcels will eventually transport off of the downstream-most link. To track these parcels, we have designated a "`dummy_element`" here, which has index value `-2`.
# +
items = {"grid_element": "link", "element_id": element_id}
parcels = DataRecord(
new_grid,
items=items,
time=[0.0],
data_vars=variables,
dummy_elements={"link": [NetworkSedimentTransporter.OUT_OF_NETWORK]},
)
# -
# ### Run the NetworkSedimentTransporter
dt_sed = 4000 # 60 * 60 * 24 *1# length of timestep (seconds)
# Before running the NST, we need to determine flow direction on the grid (upstream and downstream for each link). To do so, we initalize and run a Landlab flow director component:
fd = FlowDirectorSteepest(new_grid, "topographic__elevation")
fd.run_one_step()
#Intialize model
#network sed transporter
nst = NetworkSedimentTransporter(
new_grid,
parcels,
fd,
bed_porosity=0.3,
g=9.81,
fluid_density=1000,
transport_method="WilcockCrowe",
)
#overland flow
of = OverlandFlow(rmg, steep_slopes=True)
of.run_one_step()
# +
# look at hydorgraph at outlet
hydrograph_time = []
discharge_at_outlet = []
height_at_outlet = []
#Define the outlet
outlet_nearest_raster_cell = rmg.find_nearest_node([new_grid.x_of_node[15], new_grid.y_of_node[15]])# did not choose 15 since at boundary
outlet_link_to_sample = rmg.links_at_node[outlet_nearest_raster_cell][3]
# +
# this section can take selveral minutes to run
#MUST RUN ALL CELLS ABOVE BEFORE THIS CELL
elapsed_time = 1.
run_time_slices = np.arange(dt_sed,run_time+1,dt_sed) #every sed transport timestep
#Run model
for t in run_time_slices:
#Run overland until next time to run sed transport
while elapsed_time < t:
# First, we calculate our time step.
dt_overland = .1 #of.calc_time_step()
#print('overland flow timestep is',dt_overland,'seconds')
# Now, we can generate overland flow.
of.overland_flow()
# Increased elapsed time
elapsed_time += dt_overland
#print("Model time: ",elapsed_time/(60*60*24), "days passed")
## Append time and discharge and water depth to their lists to save data and for plotting.
hydrograph_time.append(elapsed_time)
q = rmg.at_link["surface_water__discharge"]
discharge_at_outlet.append(np.abs(q[outlet_link_to_sample]) * rmg.dx)
h = rmg.at_node['surface_water__depth']
height_at_outlet.append(np.abs(h[outlet_nearest_raster_cell]))
# updated the water depth in network graph
for i in range(new_grid.number_of_links):
idx_node_2_link = new_grid.nodes_at_link[i][1]
nmg_link_h[i] = rmg.at_node['surface_water__depth'][idx_node_2_link] #assume second node is downstream one
new_grid.at_link["flow_depth"] = nmg_link_h.copy()
#Run sed transport
nst.run_one_step(dt_sed)
print("Sed transporter run at model time: ",elapsed_time/(60*60*24), "days passed")
#Plot overland flow
fig=plt.figure()
imshow_grid(rmg,'topographic__elevation',colorbar_label='Elevation (m)')
imshow_grid(rmg,'surface_water__depth',limits=(0,2),cmap=cmap,colorbar_label='Water depth (m)')
plt.title(f'Time = {round(elapsed_time,1)} s')
plt.plot(rmg.node_x[outlet_nearest_raster_cell], rmg.node_y[outlet_nearest_raster_cell], "yo")
plt.show()
#fig.savefig(f"Hima_results/runoff_{round(elapsed_time,1)}.jpeg")
#Make more rain
rmg.at_node['surface_water__depth'][bools] += 1
#water depth m
#grain size
parcel_D = parcels.dataset.D.values.copy()
parcel_D_off_grid=parcel_D[parcels.dataset["element_id"].values==-2]
# the histogram of the data
plt.hist(parcel_D_off_grid*1000, histtype='bar')
plt.xlabel('grain size (mm)')
plt.ylabel('Count')
plt.title('Histogram of grain sizes that left grid')
plt.text(0.011, 700, r'original distribution $\mu=2 mm$')
plt.xlim(0, 20)
plt.ylim(0, 4000)
plt.grid(True)
plt.show()
#Plot sediment parcels locations
fig = plot_network_and_parcels(
grid, parcels,
parcel_time_index=len(parcels.time_coordinates)-1)
plt.plot(rmg.node_x[outlet_nearest_raster_cell], rmg.node_y[outlet_nearest_raster_cell], "yo")
plt.title(f'Time = {round(elapsed_time,1)} s')
plt.show()
#fig.savefig(f"Hima_results/sedparcels_{round(elapsed_time,1)}.jpeg")
# +
## Plotting hydrographs and discharge
fig=plt.figure(2)
plt.plot(hydrograph_time, discharge_at_outlet, "b-", label="outlet")
plt.ylabel("Discharge (cms)")
plt.xlabel("Time (seconds)")
plt.legend(loc="upper right")
fig.savefig(f"Hima_results/runoff_discharge.jpeg")
fig=plt.figure(3)
plt.plot(hydrograph_time, height_at_outlet, "b-", label="outlet")
plt.ylabel("Water depth (m)")
plt.xlabel("Time (seconds)")
plt.legend(loc="upper right")
fig.savefig("Hima_results/runoff_waterdepth.jpeg")
# +
## Plotting sediment volume
parcel_vol_on_grid = parcels.dataset["volume"].values
parcel_vol_on_grid[parcels.dataset["element_id"].values==-2]=0
sum_parcel_vol_on_grid = np.sum(parcel_vol_on_grid, axis=0)
#plt.figure(figsize=(8,6))
plt.plot(np.asarray(parcels.time_coordinates)/(60*60*24),
sum_parcel_vol_on_grid[0]-sum_parcel_vol_on_grid,
'-',
linewidth=3,
)
plt.ylabel('Total volume of parcels that left catchment $[m^3]$')
plt.xlabel('Time [days]')
plt.show()
# -
| upland/hima-dev-sed-transport.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# + [markdown] origin_pos=0
# # 序列模型
# :label:`sec_sequence`
#
# 想象一下你正在看网飞(Netflix,一个国外的视频网站)上的电影。作为一个很棒的网飞用户,你决定对每一部电影都给出评价。毕竟,一部好的电影值得好电影这个名声,而且你想看更多的好电影,对吧?事实证明,事情并不那么简单。随着时间的推移,人们对电影的看法会发生很大的变化。事实上,心理学家甚至对某些效应起了名字:
#
# * *锚定*(anchoring),基于其他人的意见。例如,奥斯卡颁奖后,受到关注的电影的评分会上升,尽管它还是原来那部电影。这种影响将持续几个月,直到人们忘记了这部电影曾经获得的奖项。结果表明,这种效应会使评分提高半个百分点以上
# :cite:`Wu.Ahmed.Beutel.ea.2017`.
# * *享乐适应*(hedonic adaption),即人类迅速接受并且适应一种更好或者更坏的情况作为新的常态。例如,在看了很多好电影之后,人们会强烈期望下部电影会一样好或者更好。因此,在许多精彩的电影被看过之后,即使是一部普通的也可能被认为是糟糕的。
# * *季节性*(seasonality)。少有观众喜欢在八月看圣诞老人的电影。
# * 有时候,电影会由于导演或演员在制作中的不当行为变得不受欢迎。
# * 有些电影因为其极度糟糕只能成为小众电影。*Plan 9 from Outer Space* 和 *Troll 2* 就因为这个原因而臭名昭著的。
#
# 简而言之,电影评分决不是固定不变的。因此,使用时间动力学可以得到更准确的电影推荐 :cite:`Koren.2009` 。当然,序列数据不仅仅是关于电影评分的。下面给出了更多的场景。
#
# * 在使用应用程序时,许多用户都有很强的特定习惯。例如,在学生放学后社交媒体应用更受欢迎。在市场开放时股市交易软件更常用。
# * 预测明天的股价要比填补昨天遗失的股价的更困难,尽管两者都只是估计一个数字。毕竟,先见之明比事后诸葛亮难得多。在统计学中,前者(对超出已知观测范围进行预测)称为 *外推法*(extrapolation),而后者(在现有观测值之间进行估计)称为 *内插法*(interpolation)。
# * 在本质上,音乐、语音、文本和视频都是连续的。如果它们的序列被我们重排,那么原有的意义就会失去。文本标题 *狗咬人* 远没有 *人咬狗* 那么令人惊讶,尽管组成两句话的字完全相同。
# * 地震具有很强的相关性,即大地震发生后,很可能会有几次较小的余震,这些余震的强度比不是大地震的余震要大得多。事实上,地震是时空相关的,即余震通常发生在很短的时间跨度和很近的距离内。
# * 人类之间的互动也是连续的,这可以从推特上的争吵和辩论中看出。
#
# ## 统计工具
#
# 处理序列数据需要统计工具和新的深度神经网络结构。为了简单起见,我们以 :numref:`fig_ftse100` 所示的股票价格(富时100指数)为例。
#
# 
# :width:`400px`
# :label:`fig_ftse100`
#
# 其中,用 $x_t$ 表示价格,即在 *时间步*(time step)$t \in \mathbb{Z}^+$时,观察到的价格 $x_t$。请注意,$t$ 对于本文中的序列通常是离散的,并随整数或其子集而变化。假设一个交易员想在 $t$ 日的股市中表现良好,于是通过以下途径预测 $x_t$:
#
# $$x_t \sim P(x_t \mid x_{t-1}, \ldots, x_1).$$
#
# ### 自回归模型
#
# 为了实现这个预测,交易员可以使用回归模型,例如在 :numref:`sec_linear_concise` 中训练的模型。仅有一个主要问题:输入数据的数量,输入 $x_{t-1}, \ldots, x_1$ 本身因 $t$ 而异。也就是说,输入数据的数量这个数字将会随着我们遇到的数据量的增加而增加,因此需要一个近似方法来使这个计算变得容易处理。本章后面的大部分内容将围绕着如何有效估计 $P(x_t \mid x_{t-1}, \ldots, x_1)$ 展开。简单地说,它归结为以下两种策略。
#
# 第一种策略,假设在现实情况下相当长的序列 $x_{t-1}, \ldots, x_1$ 可能是不必要的,因此我们只需要满足某个长度为 $\tau$ 的时间跨度,即使用观测序列 $x_{t-1}, \ldots, x_{t-\tau}$。当下获得的最直接的好处就是参数的数量总是不变的,至少在 $t > \tau$ 时如此,这就使我们能够训练一个上面提及的深度网络。这种模型被称为 *自回归模型*(autoregressive models),因为它们就是对自己执行回归。
#
# 第二种策略,如 :numref:`fig_sequence-model` 所示,是保留一些对过去观测的总结 $h_t$,并且同时更新预测 $\hat{x}_t$ 和总结 $h_t$。这就产生了基于 $\hat{x}_t = P(x_t \mid h_{t})$ 估计 $x_t$,以及公式 $h_t = g(h_{t-1}, x_{t-1})$ 更新的模型。由于 $h_t$ 从未被观测到,这类模型也被称为 *隐变量自回归模型*(latent autoregressive models)。
#
# 
# :label:`fig_sequence-model`
#
# 这两种情况都有一个显而易见的问题,即如何生成训练数据。一个经典方法是使用到目前为止的历史观测来预测下一个未来观测。显然,我们并不指望时间会停滞不前。然而,一个常见的假设是虽然特定值 $x_t$ 可能会改变,但是序列本身的动力学不会改变。这样的假设是合理的,因为新的动力学一定受新的数据影响,而我们不可能用目前所掌握的数据来预测新的动力学。统计学家称不变的动力学为 *静止的*(stationary)。因此,无论我们做什么,整个序列的估计值都将通过以下的方式获得
#
# $$P(x_1, \ldots, x_T) = \prod_{t=1}^T P(x_t \mid x_{t-1}, \ldots, x_1).$$
#
# 注意,如果我们处理的是离散的对象(如单词),而不是连续的数字,则上述的考虑仍然有效。唯一的差别是,对于离散的对象,我们需要使用分类器而不是回归模型来估计 $P(x_t \mid x_{t-1}, \ldots, x_1)$。
#
# ### 马尔可夫模型
#
# 回想一下,在自回归模型的近似法中,我们使用 $x_{t-1}, \ldots, x_{t-\tau}$ 而不是 $x_{t-1}, \ldots, x_1$ 来估计 $x_t$。只要这种近似是精确的,我们就说序列满足 *马尔可夫条件*(Markov condition)。特别是,如果 $\tau = 1$,得到一个 *一阶马尔可夫模型*(first-order Markov model),$P(x)$ 由下式给出:
#
# $$P(x_1, \ldots, x_T) = \prod_{t=1}^T P(x_t \mid x_{t-1}) \text{ where } P(x_1 \mid x_0) = P(x_1).$$
#
# 当假设 $x_t$ 仅是离散值时,这样的模型特别棒,因为在这种情况下,使用动态规划可以沿着马尔可夫链精确地计算结果。例如,我们可以高效地计算$P(x_{t+1} \mid x_{t-1})$:
#
# $$
# \begin{aligned}
# P(x_{t+1} \mid x_{t-1})
# &= \frac{\sum_{x_t} P(x_{t+1}, x_t, x_{t-1})}{P(x_{t-1})}\\
# &= \frac{\sum_{x_t} P(x_{t+1} \mid x_t, x_{t-1}) P(x_t, x_{t-1})}{P(x_{t-1})}\\
# &= \sum_{x_t} P(x_{t+1} \mid x_t) P(x_t \mid x_{t-1})
# \end{aligned}
# $$
#
# 利用这一事实,我们只需要考虑过去观察中的一个非常短的历史:$P(x_{t+1} \mid x_t, x_{t-1}) = P(x_{t+1} \mid x_t)$。动态规划的详细介绍超出了本节的范围,而动态规划这些计算工具已经在控制算法和强化学习算法广泛使用。
#
# ### 因果关系
#
# 原则上,将 $P(x_1, \ldots, x_T)$ 倒序展开也没啥问题。毕竟,基于条件概率公式,我们总是可以写出:
#
# $$P(x_1, \ldots, x_T) = \prod_{t=T}^1 P(x_t \mid x_{t+1}, \ldots, x_T).$$
#
# 事实上,如果基于一个马尔可夫模型,我们还可以得到一个反向的条件概率分布。然而,在许多情况下,数据存在一个自然的方向,即在时间上是前进的。很明显,未来的事件不能影响过去。因此,如果我们改变 $x_t$,可能会影响未来发生的事情 $x_{t+1}$,但不能反过来。也就是说,如果我们改变 $x_t$,基于过去事件得到的分布不会改变。因此,解释 $P(x_{t+1} \mid x_t)$ 应该比解释 $P(x_t \mid x_{t+1})$ 更容易。例如,在某些情况下,对于某些可加性噪声 $\epsilon$,显然我们可以找到 $x_{t+1} = f(x_t) + \epsilon$,而反之则不行 :cite:`Hoyer.Janzing.Mooij.ea.2009` 。这是个好消息,因为这个前进方向通常也是我们感兴趣的方向。彼得斯等人写的这本书 :cite:`Peters.Janzing.Scholkopf.2017` 已经解释了关于这个主题的更多内容 ,而我们仅仅触及了它的皮毛。
#
# ## 训练
#
# 在回顾了这么多统计工具之后,让我们在实践中尝试一下。首先,生成一些数据。简单起见,我们(**使用正弦函数和一些可加性噪声来生成序列数据,时间步为 $1, 2, \ldots, 1000$。**)
#
# + origin_pos=2 tab=["pytorch"]
# %matplotlib inline
import torch
from torch import nn
from d2l import torch as d2l
# + origin_pos=4 tab=["pytorch"]
T = 1000 # 总共产生1000个点
time = torch.arange(1, T + 1, dtype=torch.float32)
x = torch.sin(0.01 * time) + torch.normal(0, 0.2, (T,))
d2l.plot(time, [x], 'time', 'x', xlim=[1, 1000], figsize=(6, 3))
# + [markdown] origin_pos=6
# 接下来,我们需要将这样的序列转换为模型可以训练的特征和标签。基于嵌入维度 $\tau$,我们[**将数据映射为数据对 $y_t = x_t$ 和 $\mathbf{x}_t = [x_{t-\tau}, \ldots, x_{t-1}]$。**]精明的读者可能已经注意到,这比我们提供的数据样本少了 $\tau$ 个,因为我们没有足够的历史记录来描述前 $\tau$ 个数据样本。一个简单的解决办法,尤其是如果拥有足够长的序列就丢弃这几项;另一个方法,我们可以用零填充序列。在这里,我们仅使用前600个“特征-标签”(feature-label)对进行训练。
#
# + origin_pos=7 tab=["pytorch"]
tau = 4
features = torch.zeros((T - tau, tau))
for i in range(tau):
features[:, i] = x[i: T - tau + i]
labels = x[tau:].reshape((-1, 1))
# + origin_pos=9 tab=["pytorch"]
batch_size, n_train = 16, 600
# 只有前`n_train`个样本用于训练
train_iter = d2l.load_array((features[:n_train], labels[:n_train]),
batch_size, is_train=True)
# + [markdown] origin_pos=10
# 在这里,训练模型[**使用一个相当简单的结构:只是一个拥有两个全连接层的多层感知机**],ReLU激活函数和平方损失。
#
# + origin_pos=12 tab=["pytorch"]
# 初始化网络权重的函数
def init_weights(m):
if type(m) == nn.Linear:
nn.init.xavier_uniform_(m.weight)
# 一个简单的多层感知机
def get_net():
net = nn.Sequential(nn.Linear(4, 10),
nn.ReLU(),
nn.Linear(10, 1))
net.apply(init_weights)
return net
# 平方损失
loss = nn.MSELoss()
# + [markdown] origin_pos=14
# 现在,准备[**训练模型**]了。实现下面的训练代码的方式与前面几节(如 :numref:`sec_linear_concise` )中的循环训练基本相同。因此,我们不会深入探讨太多细节。
#
# + origin_pos=16 tab=["pytorch"]
def train(net, train_iter, loss, epochs, lr):
trainer = torch.optim.Adam(net.parameters(), lr)
for epoch in range(epochs):
for X, y in train_iter:
trainer.zero_grad()
l = loss(net(X), y)
l.backward()
trainer.step()
print(f'epoch {epoch + 1}, '
f'loss: {d2l.evaluate_loss(net, train_iter, loss):f}')
net = get_net()
train(net, train_iter, loss, 5, 0.01)
# + [markdown] origin_pos=18
# ## 预测
#
# 由于训练损失很小,因此我们期望模型能有很好的工作效果。让我们看看这在实践中意味着什么。首先是检查[**模型预测下一个时间步**]发生的是什么的能力,也就是 *单步预测*(one-step-ahead prediction)。
#
# + origin_pos=19 tab=["pytorch"]
onestep_preds = net(features)
d2l.plot([time, time[tau:]], [x.detach().numpy(), onestep_preds.detach().numpy()], 'time',
'x', legend=['data', '1-step preds'], xlim=[1, 1000], figsize=(6, 3))
# + [markdown] origin_pos=20
# 正如我们所料,单步预测效果不错。即使这些预测的时间步超过了 $600+4$(`n_train + tau`),其结果看起来仍然是可信的。然而有一个小问题:如果数据观察序列的时间步只到 $604$,那么我们就没办法指望能够得到所有未来的单步预测作为输入。相反,我们需要一步一步地向前迈进:
# $$
# \hat{x}_{605} = f(x_{601}, x_{602}, x_{603}, x_{604}), \\
# \hat{x}_{606} = f(x_{602}, x_{603}, x_{604}, \hat{x}_{605}), \\
# \hat{x}_{607} = f(x_{603}, x_{604}, \hat{x}_{605}, \hat{x}_{606}),\\
# \hat{x}_{608} = f(x_{604}, \hat{x}_{605}, \hat{x}_{606}, \hat{x}_{607}),\\
# \hat{x}_{609} = f(\hat{x}_{605}, \hat{x}_{606}, \hat{x}_{607}, \hat{x}_{608}),\\
# \ldots
# $$
#
# 通常,对于直到 $x_t$ 的观测序列,其在时间步 $t+k$ 处的预测输出 $\hat{x}_{t+k}$ 称为 *$k$ 步预测*($k$-step-ahead-prediction)。由于我们的观察已经到了 $x_{604}$,它的 $k$ 步预测是 $\hat{x}_{604+k}$。换句话说,我们必须使用我们自己的预测(而不是原始数据)来[**进行多步预测**]。让我们看看效果如何。
#
# + origin_pos=21 tab=["pytorch"]
multistep_preds = torch.zeros(T)
multistep_preds[: n_train + tau] = x[: n_train + tau]
for i in range(n_train + tau, T):
multistep_preds[i] = net(
multistep_preds[i - tau:i].reshape((1, -1)))
# + origin_pos=23 tab=["pytorch"]
d2l.plot([time, time[tau:], time[n_train + tau:]],
[x.detach().numpy(), onestep_preds.detach().numpy(),
multistep_preds[n_train + tau:].detach().numpy()], 'time',
'x', legend=['data', '1-step preds', 'multistep preds'],
xlim=[1, 1000], figsize=(6, 3))
# + [markdown] origin_pos=24
# 如上面的例子所示,这是一个巨大的失败。经过几个预测步骤之后,预测的结果很快就会衰减到一个常数。为什么这个算法效果这么差呢?最终事实是由于错误的累积。假设在步骤 $1$ 之后,我们积累了一些错误 $\epsilon_1 = \bar\epsilon$。于是,步骤 $2$ 的 *输入*(input)被扰动了 $\epsilon_1$,结果积累的误差是依照次序的 $\epsilon_2 = \bar\epsilon + c \epsilon_1$,其中 $c$ 为某个常数,后面的预测误差依此类推。因此误差可能会相当快地偏离真实的观测结果。例如,未来 $24$ 小时的天气预报往往相当准确,但超过这一点,准确率就会迅速下降。我们将在本章及后续章节中讨论如何改进这一点。
#
# 基于 $k = 1, 4, 16, 64$,通过对整个序列预测的计算,让我们[**更仔细地看一下$k$步预测**]的困难。
#
# + origin_pos=25 tab=["pytorch"]
max_steps = 64
# + origin_pos=26 tab=["pytorch"]
features = torch.zeros((T - tau - max_steps + 1, tau + max_steps))
# 列 `i` (`i` < `tau`) 是来自 `x` 的观测
# 其时间步从 `i + 1` 到 `i + T - tau - max_steps + 1`
for i in range(tau):
features[:, i] = x[i: i + T - tau - max_steps + 1]
# 列 `i` (`i` >= `tau`) 是 (`i - tau + 1`)步的预测
# 其时间步从 `i + 1` 到 `i + T - tau - max_steps + 1`
for i in range(tau, tau + max_steps):
features[:, i] = net(features[:, i - tau:i]).reshape(-1)
# + origin_pos=28 tab=["pytorch"]
steps = (1, 4, 16, 64)
d2l.plot([time[tau + i - 1: T - max_steps + i] for i in steps],
[features[:, (tau + i - 1)].detach().numpy() for i in steps], 'time', 'x',
legend=[f'{i}-step preds' for i in steps], xlim=[5, 1000],
figsize=(6, 3))
# + [markdown] origin_pos=29
# 这清楚地说明了当我们试图预测更远的未来时,预测的质量是如何变化的。虽然“$4$ 步预测”看起来仍然不错,但超过这个跨度的任何预测几乎都是无用的。
#
# ## 小结
#
# * 内插法(在现有观测值之间进行估计)和外推法(对超出已知观测范围进行预测)在实践的难度上差别很大。因此,对于你所拥有的序列数据,在训练时始终要尊重其时间顺序,即永远不要基于未来的数据进行训练。
# * 序列模型的估计需要专门的统计工具,两种较流行的选择是自回归模型和隐变量自回归模型。
# * 对于时间是向前推进的因果模型,正向估计通常比反向估计更容易。
# * 对于直到时间步 $t$ 的观测序列,其在时间步 $t+k$ 的预测输出是"$k$步预测"。随着我们对预测时间 $k$ 值的增加,会造成误差的快速累积和预测质量的极速下降。
#
# ## 练习
#
# 1. 改进本节实验中的模型。
# 1. 是否包含了过去 $4$ 个以上的观测结果?你的真实需要是多少个?
# 1. 如果没有噪音,你需要多少个过去的观测结果?提示:你可以把 $\sin$ 和 $\cos$ 写成微分方程。
# 1. 你能在保持特征总数不变的情况下合并旧的观察结果吗?这能提高正确度吗?为什么?
# 1. 改变神经网络结构并评估其性能。
# 1. 一位投资者想要找到一种好的证券来购买。他查看过去的回报,以决定哪一种可能是表现良好的。这一策略可能会出什么问题呢?
# 1. 时间是向前推进的因果模型在多大程度上适用于文本呢?
# 1. 举例说明什么时候可能需要隐变量自回归模型来捕捉数据的动力学模型。
#
# + [markdown] origin_pos=31 tab=["pytorch"]
# [Discussions](https://discuss.d2l.ai/t/2091)
#
| d2l/chapter_recurrent-neural-networks/sequence.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.10 64-bit
# name: python3
# ---
# # Inheritance Exercise Clothing
#
# The following code contains a Clothing parent class and two children classes: Shirt and Pants.
#
# Your job is to code a class called Blouse. Read through the code and fill out the TODOs. Then check your work with the unit tests at the bottom of the code.
# +
class Clothing:
def __init__(self, color, size, style, price):
self.color = color
self.size = size
self.style = style
self.price = price
def change_price(self, price):
self.price = price
def calculate_discount(self, discount):
return self.price * (1 - discount)
def calculate_shipping(self, weight, rate):
return weight*rate
class Shirt(Clothing):
def __init__(self, color, size, style, price, long_or_short):
Clothing.__init__(self, color, size, style, price)
self.long_or_short = long_or_short
def double_price(self):
self.price = 2*self.price
class Pants(Clothing):
def __init__(self, color, size, style, price, waist):
Clothing.__init__(self, color, size, style, price)
self.waist = waist
def calculate_discount(self, discount):
return self.price * (1 - discount / 2)
# TODO: Write a class called Blouse, that inherits from the Clothing class
# and has the the following attributes and methods:
# attributes: color, size, style, price, country_of_origin
# where country_of_origin is a string that holds the name of a
# country
#
# methods: triple_price, which has no inputs and returns three times
# the price of the blouse
#
#
class Blouse(Clothing):
def __init__(self, color, size, style, price, country_of_origin):
Clothing.__init__(self, color, size, style, price)
self.country_of_origin = country_of_origin
def triple_price(self):
return self.price*3
# TODO: Add a method to the clothing class called calculate_shipping.
# The method has two inputs: weight and rate. Weight is a float
# representing the weight of the article of clothing. Rate is a float
# representing the shipping weight. The method returns weight * rate
# +
# Unit tests to check your solution
import unittest
class TestClothingClass(unittest.TestCase):
def setUp(self):
self.clothing = Clothing('orange', 'M', 'stripes', 35)
self.blouse = Blouse('blue', 'M', 'luxury', 40, 'Brazil')
self.pants = Pants('black', 32, 'baggy', 60, 30)
def test_initialization(self):
self.assertEqual(self.clothing.color, 'orange', 'color should be orange')
self.assertEqual(self.clothing.price, 35, 'incorrect price')
self.assertEqual(self.blouse.color, 'blue', 'color should be blue')
self.assertEqual(self.blouse.size, 'M', 'incorrect size')
self.assertEqual(self.blouse.style, 'luxury', 'incorrect style')
self.assertEqual(self.blouse.price, 40, 'incorrect price')
self.assertEqual(self.blouse.country_of_origin, 'Brazil', 'incorrect country of origin')
def test_calculateshipping(self):
self.assertEqual(self.clothing.calculate_shipping(.5, 3), .5 * 3,\
'Clothing shipping calculation not as expected')
self.assertEqual(self.blouse.calculate_shipping(.5, 3), .5 * 3,\
'Clothing shipping calculation not as expected')
tests = TestClothingClass()
tests_loaded = unittest.TestLoader().loadTestsFromModule(tests)
unittest.TextTestRunner().run(tests_loaded)
# -
| lessons/ObjectOrientedProgramming/JupyterNotebooks/5.OOP_code_inheritance_clothing/inheritance_exercise_clothing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Watch Me Code 3: Mapping with Folium
#
# Folium is a Python wrapper library for the OpenStreetMaps api. It allows you to place data on a map in a variety of ways.
#
# ! pip install folium
import folium
import pandas as pd
import random
# we need to center the map in the middle of the US. I googled for the location.
CENTER_US = (39.8333333,-98.585522)
london = (51.5074, -0.1278)
map = folium.Map(location=CENTER_US, zoom_start=4)
map
# ## Map Pins
# read in a data file of IP address to locations.
data = pd.read_csv('https://raw.githubusercontent.com/mafudge/datasets/master/clickstream/ip_lookup.csv')
data.sample(5)
# Let's place each location on the map
CENTER_US = (39.8333333,-98.585522)
map = folium.Map(location=CENTER_US, zoom_start=4)
for row in data.to_records():
pos = (row['ApproxLat'],row['ApproxLng'])
marker = folium.Marker(location=pos,
popup="%s, %s" % (row['City'],row['State'])
)
map.add_child(marker)
map
# Same thing with a different icon and colors.
#Icons come from http://fontawesome.io/icons/ but its an older version.
colors = ['red', 'blue', 'green', 'purple', 'orange', 'darkred',
'lightred', 'beige', 'darkblue', 'darkgreen', 'cadetblue',
'darkpurple', 'pink', 'lightblue', 'lightgreen',
'gray', 'black', 'lightgray']
for row in data.to_records():
pos = (row['ApproxLat'],row['ApproxLng'])
marker = folium.Marker(location=pos,
popup="%s, %s" % (row['City'],row['State']),
icon = folium.Icon(color = random.choice(colors), icon='user')
)
map.add_child(marker)
map
# +
# There are other map tiles available. See https://folium.readthedocs.io/en/latest/quickstart.html
# Instead of Markers we use circles colors are HTML color codes http://htmlcolorcodes.com/
CENTER_US = (39.8333333,-98.585522)
map2 = folium.Map(location=CENTER_US, zoom_start=4, tiles=' Stamen Toner')
for row in data.to_records():
map2.add_child(folium.Circle(location=(row['ApproxLat'],row['ApproxLng']),
popup=row['City'], radius=10000, color='#0000FF', fill_color='#FF3333', fill=True))
map2
# -
# # Choropleths
#
# Choropleths are cartographic overlays based on boundries defined in a geo JSON file.
#
# State level geo-json overlay choropleth
CENTER_US = (39.8333333,-98.585522)
state_geojson = 'WMC3-us-states.json'
map3 = folium.Map(location=CENTER_US, zoom_start=4, tiles=' Open Street Map')
map3.choropleth(state_geojson)
map3
help(map3.choropleth)
states = pd.read_csv('https://raw.githubusercontent.com/jasonong/List-of-US-States/master/states.csv')
state_counts = pd.DataFrame( {'Counts' : data['State']. value_counts() } ).sort_index()
state_counts['StateCode'] = state_counts.index
state_data = states.merge(state_counts, how="left", left_on='Abbreviation', right_on='StateCode')
state_data = state_data[['Abbreviation','Counts']]
state_data = state_data.fillna(0)
state_data
CENTER_US = (39.8333333,-98.585522)
state_geojson = 'WMC3-us-states.json'
map3 = folium.Map(location=CENTER_US, zoom_start=4, tiles=' Open Street Map')
map3.choropleth(state_geojson,data=state_data, columns=['Abbreviation','Counts'],
key_on ='feature.id', fill_color='BuGn', legend_name='Website Visitors')
map3
# Here's a more straigtforward example with unemployment data:
unemployment = pd.read_csv('https://raw.githubusercontent.com/wrobstory/vincent/master/examples/data/US_Unemployment_Oct2012.csv')
state_geojson = 'WMC3-us-states.json'
map4 = folium.Map(location=CENTER_US, zoom_start=4, tiles=' Open Street Map')
map4.choropleth(state_geojson,data=unemployment,
columns=['State','Unemployment'], key_on ='feature.id', fill_color='YlOrRd', legend_name='2012 US Unemployment')
map4
| content/lessons/13/Watch-Me-Code/WMC3-Folium-Map.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Lesson 5 - Datasets and Questions
#
# ## What is a person of interest?
# - people who were sentenced
# - people who settled without admitting guilt
# - testified in exchange for immunity
# ## Accuracy vs Training Set Size
# - size of training set is important as it has a big affect on the accuracy that we are able to achieve
#
# example
# - physics problem
# - was using naive bayes to classify particles
# - 1000 events
# - Was this much events enough to capture all trends in the data?
# - divided training data into batches of 200. Trained it with 200, 400, 600 and 800. Then test it with remaining 200 events
#
# 
#
# - we have around 30 people in our Enron data set. Is that enough? We don't really know. There is no good way except to try it out
# - we need to find how the accuracy changes with the number of training data. Can we get more data? If we can then asking this series of questions can be really helpful
# - Having more data almost always helps the performance of your algorithm
# ## Enron Data
# - Katie downloaded the data set and cross checked the mails with the persons of interest list
# - Found that the data set only had 4 or 5 people whose inbox was there in the data set.
# ## Types of Data
# - Numerical
# - Categorical - limited number of discrete values (category)
# - time series - temporal values (date, timestamp)
# - text - words
| udacity_data_science_notes/intro_machine_learning/lesson_05/lesson_05.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import cv2
import numpy as np
body_classifier = cv2.CascadeClassifier('Haarcascades\haarcascade_fullbody.xml')
# Initiate video capture for video file
cap = cv2.VideoCapture('walking.avi')
while cap.isOpened():
ret, frame = cap.read()
gray = cv2.cvtColor(frame, cv2.COLOR_BGR2GRAY)
bodies = body_classifier.detectMultiScale(gray, 1.2, 3)
#bounding boxes for any bodies identified
for (x,y,w,h) in bodies:
cv2.rectangle(frame, (x, y), (x+w, y+h), (0, 255, 255), 2)
cv2.imshow('Pedestrians', frame)
if cv2.waitKey(1) == 13:
break
cap.release()
cv2.destroyAllWindows()
| Open CV/Pedestrian-Detection-OpenCV/Pedestrian Detection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
def filter(func,li):
a=[]
for i in li:
if(func(i)):
a.append(i)
return a
a=[1,2,3,4,5,-1,0,-3,-1,-34,123,-12]
def posi(a):
if(a>0):
return True
return False
filter(posi,a)
| sol_dipit-s/5. My filter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="TAtrzndb7Vun" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1646754170978, "user_tz": -540, "elapsed": 17536, "user": {"displayName": "\uae40\ud615\uc9c4", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "05530574334109307920"}} outputId="158bf875-c2f5-40cd-9d06-e4a2d449c185"
import tensorflow as tf
import ssl
ssl._create_default_https_context = ssl._create_unverified_context
def pipe(data, batch_size = 128, shuffle = False):
dataset = tf.data.Dataset.from_tensor_slices(data)
if shuffle:
dataset = dataset.shuffle(buffer_size = batch_size * 10)
dataset = dataset.batch(batch_size)
#dataset = dataset.prefetch((batch_size * 2) + 1)
dataset = dataset.prefetch(tf.data.experimental.AUTOTUNE)
return dataset
(tr_x, tr_y), (te_x, te_y) = tf.keras.datasets.cifar10.load_data()
tr_x = tr_x * 1/255
te_x = te_x * 1/255
batch_size = 128
tr_data = pipe((tr_x, tr_y), batch_size = batch_size, shuffle = True)
te_data = pipe((te_x, te_y), batch_size = batch_size, shuffle = False)
# + id="ySaeM_5H7Q8A" executionInfo={"status": "ok", "timestamp": 1646754187602, "user_tz": -540, "elapsed": 5181, "user": {"displayName": "\uae40\ud615\uc9c4", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "05530574334109307920"}}
import effnet
model = effnet.efficientnet_lite_b0(input_shape = (32, 32, 3), include_top = False, weights = "imagenet")
flatten = tf.keras.layers.GlobalAveragePooling2D()(model.output)
drop_out = tf.keras.layers.Dropout(0.5)(flatten)
dense = tf.keras.layers.Dense(2048, activation = "relu")(drop_out)
prediction = tf.keras.layers.Dense(10, activation = "softmax", name = "prediction")(dense)
model = tf.keras.Model(model.input, prediction)
# + id="cka3vwta8pmU" executionInfo={"status": "ok", "timestamp": 1646754191246, "user_tz": -540, "elapsed": 398, "user": {"displayName": "\uae40\ud615\uc9c4", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "05530574334109307920"}}
loss = tf.keras.losses.sparse_categorical_crossentropy
opt = tf.keras.optimizers.Adam(1e-4)
metric = [tf.keras.metrics.sparse_categorical_accuracy]
model.compile(loss = loss, optimizer = opt, metrics = metric)
# + colab={"base_uri": "https://localhost:8080/"} id="S2T8gk6z9iBH" outputId="39e0d083-e908-4752-87cf-8cf0d11b72f0" executionInfo={"status": "ok", "timestamp": 1646754395866, "user_tz": -540, "elapsed": 203016, "user": {"displayName": "\uae40\ud615\uc9c4", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "05530574334109307920"}}
model.fit(tr_data, validation_data = te_data, epochs = 10)
# + id="tXn4W1lqhbbf" executionInfo={"status": "ok", "timestamp": 1646754396362, "user_tz": -540, "elapsed": 506, "user": {"displayName": "\uae40\ud615\uc9c4", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "05530574334109307920"}}
with open("model.json", mode = "w") as file:
file.write(model.to_json())
model.save_weights("model.h5")
# + id="9Rx7ssmeh167" executionInfo={"status": "ok", "timestamp": 1646754397266, "user_tz": -540, "elapsed": 911, "user": {"displayName": "\uae40\ud615\uc9c4", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "05530574334109307920"}}
with open("model.json", mode = "r") as file:
model = tf.keras.models.model_from_json(file.read())
model.load_weights("model.h5")
# + colab={"base_uri": "https://localhost:8080/"} id="JTBoj3vrFLca" outputId="e7ab8e28-2403-4ed7-fa93-0b7ed5f57eb8" executionInfo={"status": "ok", "timestamp": 1646754399417, "user_tz": -540, "elapsed": 2158, "user": {"displayName": "\uae40\ud615\uc9c4", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "05530574334109307920"}}
loss = tf.keras.losses.sparse_categorical_crossentropy
metric = [tf.keras.metrics.sparse_categorical_accuracy]
model.compile(loss = loss, metrics = metric)
model.evaluate(te_data)
# + id="MsTu5aaRHY9d"
| usage.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# <center><h1><strong>tau-data Indonesia</strong></h1></center>
# <center><h2><strong><font color="blue">Pendahuluan SQL dasar untuk Data Science - 01</font></strong></h2></center>
# <img alt="" src="images/cover.jpg" />
#
# <b><center>(C) <NAME></center>
# <center><h3><font color="blue">https://tau-data.id/dfds-01/</font></h3></center>
# + [markdown] slideshow={"slide_type": "slide"}
# # Instalasi:
#
# * XAMPP: https://www.apachefriends.org/download.html
# * IBM Watson: https://www.ibm.com/cloud/sql-query
#
# ## PHPMyAdmin: Obat Sakit Kepala
# * http://localhost/phpmyadmin/index.php
# + slideshow={"slide_type": "slide"}
# !pip install --upgrade mysql-connector-python
# + slideshow={"slide_type": "fragment"}
import mysql.connector as mysql
import pandas as pd
from time import time
# + [markdown] slideshow={"slide_type": "slide"}
# # Koneksi ke MySQL lokal
# + slideshow={"slide_type": "slide"}
par = {'db_': 'fga', 'usr':'root', 'pas':'', 'hst':'localhost'}
# -
# # Mitra tau-data silahkan menggunakan server cloud tau-data Indonesia.
#
# ## Credentials akan diberikan lewat jalur private
# + slideshow={"slide_type": "fragment"}
par['db_'], par['hst']
# + slideshow={"slide_type": "slide"}
def conMql(dbPar, maxTry=3):
try_ = 0
while try_<maxTry:
try:
return mysql.connect(host=dbPar['hst'],user=dbPar['usr'],passwd=dbPar['pas'],db=dbPar['db_'])
except (mysql.Error) as e:
print ("Error Connecting to MySQL %d: %s, please wait retrying" % (e.args[0],e.args[1]))
try_ += 1; time.sleep(1)
# -
db = conMql(par)
db
# + [markdown] slideshow={"slide_type": "slide"}
# # Catatan:
#
# * Buka PHPMyAdmin ==> Generate Query untuk membantu pemula di Query Dasar
# * "Bisa karena terbiasa" ==> jangan hawatir untuk "mengingat"
# + slideshow={"slide_type": "slide"}
qry = "CREATE TABLE `mahasiswa` ( `nim` VARCHAR(12) NOT NULL COMMENT 'Nomer Induk Mahasiswa' , \
`nama_lengkap` VARCHAR(255) CHARACTER SET utf8 COLLATE utf8_general_ci NOT NULL COMMENT \
'Nama Lengkap' , `jenis_kelamin` VARCHAR(1) CHARACTER SET ascii COLLATE ascii_general_ci \
NOT NULL COMMENT 'Jenis Kelamin P atau W' , `angkatan` YEAR NOT NULL COMMENT \
'Tahun Masuk UIN Jakarta' , PRIMARY KEY (`nim`)) \
ENGINE = MyISAM CHARSET=utf8 COLLATE utf8_general_ci COMMENT = 'Tabel Mahasiswa UIN Jakarta';"
db = conMql(par)
cur = db.cursor()
res = cur.execute(qry)
cur.close()
db.close()
res
# +
qry = "SHOW TABLES"
db = conMql(par)
cur = db.cursor()
cur.execute(qry)
data = cur.fetchall() # lakukan Pagination untuk data yang sangat besar
cur.close(); db.close()
data
# + [markdown] slideshow={"slide_type": "slide"}
# # Catatan:
#
# * Di dunia nyata "jenis kelamin"/gender .. Most likely Boolean.
# * Module ini untuk Data Scientist (bukan Data Engineer), untuk sementara waktu kita hiraukan terlebih dahulu.
#
# ## Well-known Engine:
# * MyISAM
# * InnoDB
# * memory
# + slideshow={"slide_type": "slide"}
qry = "INSERT INTO `mahasiswa` (`nim`, `nama_lengkap`, `angkatan`, `jenis_kelamin`) \
VALUES ('1234', 'bambang', '2016', 'P'), ('1235', 'Wati', '2017', 'W'), \
('1239', 'Iwan', '2017', 'P');"
db = conMql(par)
cur = db.cursor()
result = cur.execute(qry)
cur.close()
db.close()
# + slideshow={"slide_type": "slide"}
qry = "SELECT * FROM mahasiswa"
db = conMql(par)
cur = db.cursor()
cur.execute(qry)
data = cur.fetchall() # lakukan Pagination untuk data yang sangat besar
cur.close(); db.close()
# + slideshow={"slide_type": "fragment"}
data
# + [markdown] slideshow={"slide_type": "slide"}
# # More Practical Way untuk Data Scientist/Analyst: Use Pandas
#
# + slideshow={"slide_type": "slide"}
qry = "SELECT * FROM mahasiswa"
db = conMql(par)
data = pd.read_sql(qry, db)
# -
data.head()
# + [markdown] slideshow={"slide_type": "slide"}
# # Contoh Data
#
# <img alt="" src="images/qry_joins_01.png" />
# + slideshow={"slide_type": "slide"}
# Create Table 2
qry = "CREATE TABLE committees (committee_id INT AUTO_INCREMENT, name VARCHAR(100), PRIMARY KEY (committee_id));"
db = conMql(par)
cur = db.cursor()
res = cur.execute(qry)
cur.close(); db.close()
'Done'
# + slideshow={"slide_type": "slide"}
# Create Table 1
qry = "CREATE TABLE members (member_id INT AUTO_INCREMENT, name VARCHAR(100), PRIMARY KEY (member_id))"
db = conMql(par)
cur = db.cursor()
res = cur.execute(qry)
cur.close(); db.close()
'Done'
# + slideshow={"slide_type": "slide"}
# Insert Data 2
qry = "INSERT INTO committees (name) VALUES ('John'), ('Mary'), ('Amelia'), ('Joe')"
db = conMql(par)
cur = db.cursor()
res = cur.execute(qry)
db.commit() # Hati-hati ... Butuh tambahan perintah ini!!!
cur.close(); db.close()
'Done'
# + slideshow={"slide_type": "slide"}
# Insert Data 1
qry = "INSERT INTO members (name) VALUES('John'),('Jane'),('Mary'),('David'),('Amelia');"
db = conMql(par)
cur = db.cursor()
res = cur.execute(qry)
db.commit() # Hati-hati ... Butuh tambahan perintah ini!!!
cur.close(); db.close()
'Done'
# + slideshow={"slide_type": "slide"}
# Query to DataFrame
db = conMql(par)
query = "SELECT * FROM committees" # or members
data = pd.read_sql(query, db)
data.head()
# + slideshow={"slide_type": "slide"}
# order by
db = conMql(par)
query = "SELECT * FROM committees ORDER BY name"
data = pd.read_sql(query, db)
data.head()
# + slideshow={"slide_type": "slide"}
# add new names
qry = "INSERT INTO committees (name) VALUES('John'),('Mary'),('Amelia'),('Bambang');"
db = conMql(par)
cur = db.cursor()
res = cur.execute(qry)
db.commit() # Hati-hati ... Butuh tambahan perintah ini!!!
cur.close(); db.close()
# + slideshow={"slide_type": "slide"}
# order by
db = conMql(par)
query = "SELECT * FROM committees ORDER BY name"
data = pd.read_sql(query, db)
data.head(10)
# + slideshow={"slide_type": "slide"}
# group by
db = conMql(par)
query = "SELECT * FROM committees GROUP BY name" # or members
data = pd.read_sql(query, db)
data.head(10)
# + slideshow={"slide_type": "slide"}
# Select Distinct
db = conMql(par)
query = "SELECT DISTINCT(name) FROM committees" # or members
data = pd.read_sql(query, db)
data.head(10)
# + slideshow={"slide_type": "slide"}
# WHERE clause
db = conMql(par)
query = "SELECT * FROM committees WHERE committee_id>4" # or members
data = pd.read_sql(query, db)
data.head(10)
# + [markdown] slideshow={"slide_type": "slide"}
# # Search with index - Magick command
# + slideshow={"slide_type": "fragment"}
# %lsmagic
# + slideshow={"slide_type": "slide"}
# %%timeit
db = conMql(par)
query = "SELECT * FROM committees WHERE name='John'" # or members
data = pd.read_sql(query, db)
# + slideshow={"slide_type": "fragment"}
# Hati-hati var "data" tidak disimpan ke memori akibat penggunaan Magic Command "timeit"
# + [markdown] slideshow={"slide_type": "slide"}
# # Sekarang kita coba tambahkan index di field "name"
# + slideshow={"slide_type": "fragment"}
qry = "ALTER TABLE committees ADD FULLTEXT name (name);"
db = conMql(par)
cur = db.cursor()
res = cur.execute(qry)
cur.close();db.close()
'Done'
# + [markdown] slideshow={"slide_type": "slide"}
# # Sekarang kita coba Query lagi dan analisa performanya
#
# * Catt: pada data yang kecil mungkin tidak signifikan, tapi pada data yang besar sangat besar pengaruhnya.
# + slideshow={"slide_type": "fragment"}
# %%timeit
db = conMql(par)
query = "SELECT * FROM committees WHERE name LIKE '%ohn%'" # or members
data = pd.read_sql(query, db)
# + slideshow={"slide_type": "slide"}
data.head()
# + [markdown] slideshow={"slide_type": "slide"}
# # SQL Joins
#
# Perintah JOIN dalam SQL digunakan untuk menampilkan data pada table yang saling berhubungan atau berelasi. Artinya kita dapat menampilkan data dalam beberapa table hanya dengan satu kali perintah.
#
# <img alt="" src="images/sql_joins.jpg" />
# + [markdown] slideshow={"slide_type": "slide"}
# # 1. inner Join (irisan) - Join Predicate
#
# ## Membandingkan setiap baris di Tabel 1 & 2 kemudian menambahkan jika kondisi join keduanya benar (True).
# + slideshow={"slide_type": "slide"}
# Query to DataFrame Kita lihat dulu datanya
db = conMql(par)
dt1 = pd.read_sql("SELECT * FROM committees", db)
dt2 = pd.read_sql("SELECT * FROM members", db)
pd.concat([dt1.reset_index(drop=1),dt2.reset_index(drop=1)], axis=1)
# + slideshow={"slide_type": "slide"}
db = conMql(par)
query = "SELECT m.member_id, m.name member, c.committee_id, c.name committee \
FROM members m \
INNER JOIN committees c \
ON c.name = m.name"
data = pd.read_sql(query, db)
data.head(10)
# + [markdown] slideshow={"slide_type": "slide"}
# # LEFT JOIN
# <img alt="" src="images/sql_joins.jpg" />
#
#
# ## Untuk setiap data dari tabel pertama (kiri/Left), buat data baru hanya jika ada padanannya di tabel kanan.
# + slideshow={"slide_type": "slide"}
# Query to DataFrame Kita lihat dulu datanya
db = conMql(par)
dt1 = pd.read_sql("SELECT * FROM committees", db)
dt2 = pd.read_sql("SELECT * FROM members", db)
pd.concat([dt1.reset_index(drop=1),dt2.reset_index(drop=1)], axis=1)
# + slideshow={"slide_type": "slide"}
query = "SELECT m.member_id, m.name member, c.committee_id, c.name committee\
FROM members m\
LEFT JOIN committees c USING(name)"
db = conMql(par)
data = pd.read_sql(query, db)
data.head(10)
# + [markdown] slideshow={"slide_type": "slide"}
# # Bisa juga
#
# * Catt hati-hati performa query
# * ini left join
# + slideshow={"slide_type": "slide"}
query = "SELECT member_id, name\
FROM members \
WHERE name IN\
(SELECT name FROM committees)"
db = conMql(par)
data = pd.read_sql(query, db)
data.head(10)
# + [markdown] slideshow={"slide_type": "slide"}
# # CROSS JOIN
# <img alt="" src="images/cj.png" />
#
#
# ## Tidak seperti inner, left/right, Cross Join tidak punya kondisi. CJ, menggabungkan setiap kolom di kiri dan kanan tabel.
# * Kalau tabel 1 ada N baris dan Tabel 2 ada M baris, maka CJ menghasilkan NxM baris.
# * Null tidak ikut CJ
# + slideshow={"slide_type": "slide"}
# Query to DataFrame Kita lihat dulu datanya
db = conMql(par)
dt1 = pd.read_sql("SELECT * FROM committees", db)
dt2 = pd.read_sql("SELECT * FROM members", db)
pd.concat([dt1.reset_index(drop=1),dt2.reset_index(drop=1)], axis=1)
# + slideshow={"slide_type": "slide"}
query = "SELECT m.member_id, m.name member, c.committee_id, c.name committee\
FROM members m\
CROSS JOIN committees c"
db = conMql(par)
data = pd.read_sql(query, db)
data.head(30)
# + [markdown] slideshow={"slide_type": "slide"}
# # End of Module
#
# <img alt="" src="images/meme_joins.png" />
| dfds-01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # !rm -vrf deep_hiv_ab_pred
# # !git clone https://github.com/vlad-danaila/deep_hiv_ab_pred.git
import sys
import os
sys.path.insert(0, 'deep_hiv_ab_pred')
from deep_hiv_ab_pred.catnap.download_dataset import download_catnap
from deep_hiv_ab_pred.train_full_catnap.train_hold_out_one_cluster import main_train, main_test
# from deep_hiv_ab_pred.train_full_catnap.hyperparameter_optimisation import optimize_hyperparameters
download_catnap()
main_train()
main_test()
| train_full_catnap/notebooks/Test Full Catnap.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# Sveučilište u Zagrebu
# Fakultet elektrotehnike i računarstva
#
# ## Strojno učenje 2017/2018
# http://www.fer.unizg.hr/predmet/su
# ------------------------------
#
# ### Laboratorijska vježba 5: Probabilistički grafički modeli, naivni Bayes, grupiranje i vrednovanje klasifikatora
#
# *Verzija: 1.3
# Zadnji put ažurirano: 12. siječnja 2018.*
#
# (c) 2015-2017 <NAME>, <NAME>
#
# Objavljeno: **30. listopada 2018.**
# Rok za predaju: **22. siječnja 2018. u 07:00h**
#
# ------------------------------
# ### Upute
#
# Prva laboratorijska vježba sastoji se od tri zadatka. U nastavku slijedite upute navedene u ćelijama s tekstom. Rješavanje vježbe svodi se na **dopunjavanje ove bilježnice**: umetanja ćelije ili više njih **ispod** teksta zadatka, pisanja odgovarajućeg kôda te evaluiranja ćelija.
#
# Osigurajte da u potpunosti **razumijete** kôd koji ste napisali. Kod predaje vježbe, morate biti u stanju na zahtjev asistenta (ili demonstratora) preinačiti i ponovno evaluirati Vaš kôd. Nadalje, morate razumjeti teorijske osnove onoga što radite, u okvirima onoga što smo obradili na predavanju. Ispod nekih zadataka možete naći i pitanja koja služe kao smjernice za bolje razumijevanje gradiva (**nemojte pisati** odgovore na pitanja u bilježnicu). Stoga se nemojte ograničiti samo na to da riješite zadatak, nego slobodno eksperimentirajte. To upravo i jest svrha ovih vježbi.
#
# Vježbe trebate raditi **samostalno**. Možete se konzultirati s drugima o načelnom načinu rješavanja, ali u konačnici morate sami odraditi vježbu. U protivnome vježba nema smisla.
# Učitaj osnovne biblioteke...
import sklearn
import codecs
import mlutils
import matplotlib.pyplot as plt
import pgmpy as pgm
# %pylab inline
# ### 1. Probabilistički grafički modeli -- Bayesove mreže
# Ovaj zadatak bavit će se Bayesovim mrežama, jednim od poznatijih probabilističkih grafičkih modela (*probabilistic graphical models*; PGM). Za lakše eksperimentiranje koristit ćemo programski paket [`pgmpy`](https://github.com/pgmpy/pgmpy). Molimo Vas da provjerite imate li ovaj paket te da ga instalirate ako ga nemate.
# #### (a)
# Prvo ćemo pogledati udžbenički primjer s prskalicom. U ovom primjeru razmatramo Bayesovu mrežu koja modelira zavisnosti između oblačnosti (slučajna varijabla $C$), kiše ($R$), prskalice ($S$) i mokre trave ($W$). U ovom primjeru također pretpostavljamo da već imamo parametre vjerojatnosnih distribucija svih čvorova. Ova mreža prikazana je na sljedećoj slici:
# 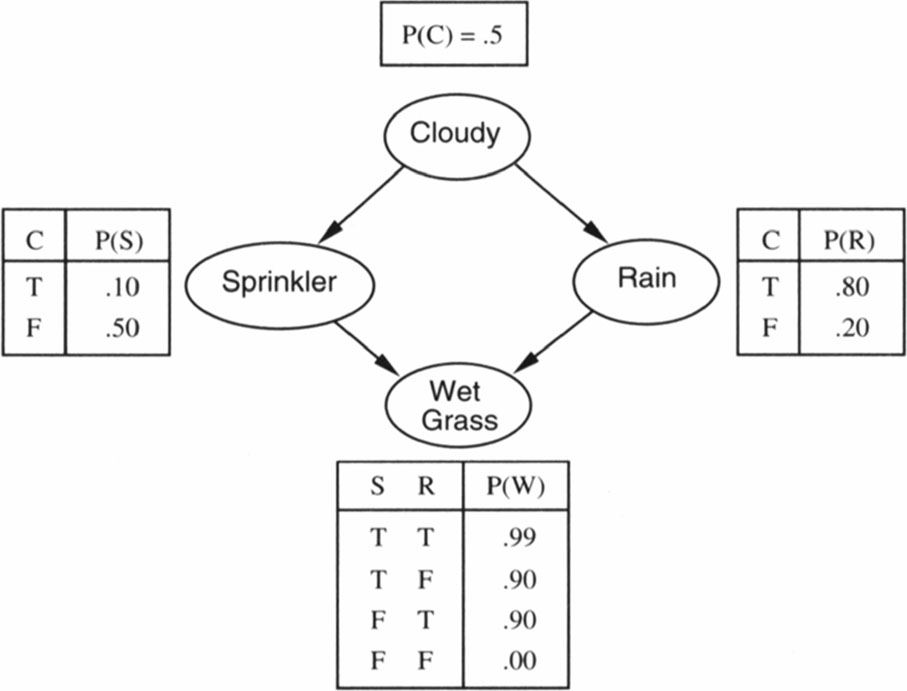
# Koristeći paket `pgmpy`, konstruirajte Bayesovu mrežu iz gornjeg primjera. Zatim, koristeći **egzaktno** zaključivanje, postavite sljedeće posteriorne upite: $P(w=1)$, $P(s=1|w=1)$, $P(r=1|w=1)$, $P(c=1|s=1, r=1)$ i $P(c=1)$. Provedite zaključivanje na papiru i uvjerite se da ste ispravno konstruirali mrežu. Pomoći će vam službena dokumentacija te primjeri korištenja (npr. [ovaj](https://github.com/pgmpy/pgmpy/blob/dev/examples/Monte%20Hall%20Problem.ipynb)).
from pgmpy.models import BayesianModel
from pgmpy.factors.discrete.CPD import TabularCPD
from pgmpy.inference import VariableElimination
# +
sprinkler_model = BayesianModel([('Cloudy', 'Rain'),
('Cloudy', 'Sprinkler'),
('Sprinkler', 'Wet'),
('Rain', 'Wet')])
cpd_cloudy = TabularCPD('Cloudy', 2, [[0.5], [0.5]])
cpd_rain = TabularCPD('Rain', 2, [[0.8, 0.2], [0.2, 0.8]], evidence=['Cloudy'], evidence_card=[2])
cpd_sprinkler = TabularCPD('Sprinkler', 2, [[0.1, 0.5], [0.9, 0.5]], evidence=['Cloudy'], evidence_card=[2])
cpd_wet = TabularCPD('Wet', 2, [[0.99, 0.9, 0.9, 0], [0.01, 0.1, 0.1, 1]], evidence=['Sprinkler', 'Rain'], evidence_card=[2, 2])
print(cpd_wet)
sprinkler_model.add_cpds(cpd_cloudy, cpd_rain, cpd_sprinkler, cpd_wet)
sprinkler_model.check_model()
# +
inference = VariableElimination(sprinkler_model)
print(inference.query(['Wet'])['Wet'].values[0])
print(inference.query(['Sprinkler'], evidence={'Wet': 0})['Sprinkler'].values[0])
print(inference.query(['Rain'], evidence={'Wet': 0})['Rain'].values[0])
print(inference.query(['Cloudy'], evidence={'Sprinkler': 0, 'Rain': 0})['Cloudy'].values[0])
print(inference.query(['Cloudy'])['Cloudy'].values[0])
# -
from pgmpy.sampling import BayesianModelSampling
sprinkler_samples = BayesianModelSampling(sprinkler_model).forward_sample(size=5, return_type='dataframe')
print(sprinkler_samples)
# **Q:** Koju zajedničku vjerojatnosnu razdiobu ova mreža modelira? Kako tu informaciju očitati iz mreže?
# **Q:** U zadatku koristimo egzaktno zaključivanje. Kako ono radi?
# **Q:** Koja je razlika između posteriornog upita i MAP-upita?
# **Q:** Zašto je vjerojatnost $P(c=1)$ drugačija od $P(c=1|s=1,r=1)$ ako znamo da čvorovi $S$ i $R$ nisu roditelji čvora $C$?
# **A:** Ova mreža modelira multivarijatnu Bernoullijevu razdiobu, koju možemo uočiti uzorkovanjem. U primjeru je korišteno unaprijedno uzorkovanje koje generira uzorke iz zajedničke distribucije mreže.
#
# **A:** Kod nekih PGM-ova egzaktnim učenjem umjesto eksponencijalnog postiže se polinomijalno vrijeme izvođenja algoritama zaključivanja, a za modele kod kojih to ne pomaže koristi se približno zaključivanje (složenost-točnost *trade-off*). Postoji više algoritama egzaktnog zaključivanja, a najpopularniji, eliminacija varijabli, distribucijom suma preko produkta marginalizira i tako eliminira neopažene i neupitane varijable u dubinu.
#
# **A:** Posteriorni upit (*engl. Posterior belief*) računa uvjetnu vjerojatnost, dok MAP (*engl. Most probable assignment*) vraća najvjerojatniju vrijednost varijable upita.
#
# **A:** Zajednička vjerojatnost Bayesove mreže na kojoj se temelji izračun posteriornog upita varira za različite opažene (*engl. evidence*) i neopažene varijable (*engl. nuisance*), pa je tako i posteriorna vjerojatnost različita.
# #### (b)
# **Efekt objašnjavanja** (engl. *explaining away*) zanimljiv je fenomen u kojem se događa da se dvije varijable "natječu" za objašnjavanje treće. Ovaj fenomen može se primijetiti na gornjoj mreži. U tom se slučaju varijable prskalice ($S$) i kiše ($R$) "natječu" za objašnjavanje mokre trave ($W$). Vaš zadatak je pokazati da se fenomen zaista događa.
print("P(R=T|W=T) = {}".format(inference.query(['Rain'], evidence={'Wet': 0})['Rain'].values[0]))
print("P(R=T|S=T,W=T) = {}".format(inference.query(['Rain'], evidence={'Wet': 0, 'Sprinkler': 0})['Rain'].values[0]))
# **Q:** Kako biste svojim riječima opisali ovaj fenomen, koristeći se ovim primjerom?
# **A:** Opažanje $S=T$ smanjuje vjerojatnost realizacije $R=T$. Odnosno, ako znamo da je trava mokra i da je uključena prskalica, vjerojatnost da uz to pada i kiša je znatno manja u odnosu na situaciju u kojoj samo znamo da je trava mokra. $S$ i $R$ su uvjetno zavisne.
# #### (c)
# Koristeći [`BayesianModel.is_active_trail`](http://pgmpy.org/models.html#pgmpy.models.BayesianModel.BayesianModel.is_active_trail) provjerite jesu li varijable oblačnosti ($C$) i mokre trave ($W$) uvjetno nezavisne. Što mora vrijediti kako bi te dvije varijable bile uvjetno nezavisne? Provjerite korištenjem iste funkcije.
print(sprinkler_model.is_active_trail('Cloudy', 'Wet', observed=['Rain']))
print(inference.query(['Wet'], evidence={'Rain': 0})['Wet'].values[0])
print(inference.query(['Wet'], evidence={'Rain': 0, 'Cloudy': 0})['Wet'].values[0])
# **Q:** Kako možemo na temelju grafa saznati koje dvije varijable su, uz neka opažanja, uvjetno nezavisne?
# **Q:** Zašto bismo uopće htjeli znati koje su varijable u mreži uvjetno nezavisne?
# **A:** Ukoliko je staza između te dvije varijable d-odvojena (*engl. d-separated*).
#
# **A:** Saznanja o ovjetnim nezavisnostima uklanjaju bridove i pojednostavljuju mrežu.
# ### 2. Vrednovanje modela (klasifikatora)
# Kako bismo se uvjerili koliko naš naučeni model zapravo dobro radi, nužno je provesti evaluaciju modela. Ovaj korak od presudne je važnosti u svim primjenama strojnog učenja, pa je stoga bitno znati provesti evaluaciju na ispravan način.
#
# Vrednovat ćemo modele na stvarnom skupu podataka [*SMS Spam Collection*](https://archive.ics.uci.edu/ml/datasets/SMS+Spam+Collection) [1], koji se sastoji od 5,574 SMS-poruka klasificiranih u dvije klase: spam (oznaka: *spam*) i ne-spam (oznaka: *ham*). Ako već niste, preuzmite skup podataka s poveznice ili sa stranice kolegija i stavite ga u radni direktorij (otpakirajte arhivu i preimenujte datoteku u `spam.csv` po potrebi). Sljedeći komad kôda učitava skup podataka i dijeli ga na podskupove za učenje i testiranje.
#
# [1] *<NAME>., <NAME>., <NAME>. Contributions to the Study of SMS Spam Filtering: New Collection and Results. Proceedings of the 2011 ACM Symposium on Document Engineering (DOCENG'11), Mountain View, CA, USA, 2011.*
# +
from sklearn.model_selection import train_test_split
spam_X, spam_y = mlutils.load_SMS_dataset('./spam.csv')
spam_X_train, spam_X_test, spam_y_train, spam_y_test = \
train_test_split(spam_X, spam_y, train_size=0.7, test_size=0.3, random_state=69)
# -
# #### (a)
# Prije nego što krenemo u vrednovanje modela za klasifikaciju spama, upoznat ćete se s jednostavnijom apstrakcijom cjelokupnog procesa učenja modela u biblioteci `scikit-learn`. Ovo je korisno zato što se učenje modela često sastoji od mnoštva koraka prije sâmog pozivanja magične funkcije `fit`: ekstrakcije podataka, ekstrakcije značajki, standardizacije, skaliranja, nadopunjavanjem nedostajućih vrijednosti i slično.
#
# U "standardnom pristupu", ovo se svodi na pozamašan broj linija kôda u kojoj konstantno proslijeđujemo podatke iz jednog koraka u sljedeći, tvoreći pritom cjevovod izvođenja. Osim nepreglednosti, ovakav pristup je često i sklon pogreškama, s obzirom na to da je dosta jednostavno proslijediti pogrešan skup podataka i ne dobiti pogrešku pri izvođenju kôda. Stoga je u biblioteci `scikit-learn` uveden razred [`pipeline.Pipeline`](http://scikit-learn.org/stable/modules/generated/sklearn.pipeline.Pipeline.html). Kroz ovaj razred, svi potrebni koraci učenja mogu se apstrahirati iza jednog cjevovoda, koji je opet zapravo model s `fit` i `predict` funkcijama.
#
# U ovom zadatku ćete napraviti samo jednostavni cjevovod modela za klasifikaciju teksta, koji se sastoji od pretvorbe teksta u vektorsku reprezentaciju vreće riječi s TF-IDF-težinama, redukcije dimenzionalnosti pomoću krnje dekompozicije singularnih vrijednosti, normalizacije, te konačno logističke regresije.
#
# **NB:** Nije sasvim nužno znati kako rade ovi razredi pomoću kojih dolazimo do konačnih značajki, ali preporučamo da ih proučite ako vas zanima (posebice ako vas zanima obrada prirodnog jezika).
from sklearn.pipeline import Pipeline
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.decomposition import TruncatedSVD
from sklearn.preprocessing import Normalizer
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import accuracy_score
# Prvo, prilažemo kôd koji to radi "standardnim pristupom":
# +
# TF-IDF
vectorizer = TfidfVectorizer(stop_words="english", ngram_range=(1, 2), max_features=500)
spam_X_feat_train = vectorizer.fit_transform(spam_X_train)
# Smanjenje dimenzionalnosti
reducer = TruncatedSVD(n_components=300, random_state=69)
spam_X_feat_train = reducer.fit_transform(spam_X_feat_train)
# Normaliziranje
normalizer = Normalizer()
spam_X_feat_train = normalizer.fit_transform(spam_X_feat_train)
# NB
clf = LogisticRegression()
clf.fit(spam_X_feat_train, spam_y_train)
# I sada ponovno sve ovo za testne podatke.
spam_X_feat_test = vectorizer.transform(spam_X_test)
spam_X_feat_test = reducer.transform(spam_X_feat_test)
spam_X_feat_test = normalizer.transform(spam_X_feat_test)
print(accuracy_score(spam_y_test, clf.predict(spam_X_feat_test)))
x_test = ["You were selected for a green card, apply here for only 50 USD!!!",
"Hey, what are you doing later? Want to grab a cup of coffee?"]
x_test = vectorizer.transform(x_test)
x_test = reducer.transform(x_test)
x_test = normalizer.transform(x_test)
print(clf.predict(x_test))
# -
# Vaš zadatak izvesti je dani kôd korištenjem cjevovoda. Proučite razred [`pipeline.Pipeline`](http://scikit-learn.org/stable/modules/generated/sklearn.pipeline.Pipeline.html).
#
# **NB** Ne treba vam više od svega nekoliko naredbi.
# +
clf = Pipeline([('vectorizer', TfidfVectorizer(stop_words="english", ngram_range=(1, 2), max_features=500)),
('reducer', TruncatedSVD(n_components=300, random_state=69)),
('normalizer', Normalizer()),
('logistic-regression', LogisticRegression())])
clf.fit(spam_X_train, spam_y_train)
print(accuracy_score(spam_y_test, clf.predict(spam_X_test)))
x_test = ["You were selected for a green card, apply here for only 50 USD!!!",
"Hey, what are you doing later? Want to grab a cup of coffee?"]
print(clf.predict(x_test))
# -
# #### (b)
# U prošlom smo podzadatku ispisali točnost našeg modela. Ako želimo vidjeti koliko je naš model dobar po ostalim metrikama, možemo iskoristiti bilo koju funkciju iz paketa [`metrics`](http://scikit-learn.org/stable/modules/classes.html#module-sklearn.metrics). Poslužite se funkcijom [`metrics.classification_report`](http://scikit-learn.org/stable/modules/generated/sklearn.metrics.classification_report.html#sklearn.metrics.classification_report), koja ispisuje vrijednosti najčešćih metrika. (Obavezno koristite naredbu `print` kako ne biste izgubili format izlaza funkcije.) Ispišite ponovno točnost za usporedbu.
from sklearn.metrics import classification_report, accuracy_score
print(classification_report(spam_y_test, clf.predict(spam_X_test)))
# Potreba za drugim metrikama osim točnosti može se vidjeti pri korištenju nekih osnovnih modela (engl. *baselines*). Možda najjednostavniji model takvog tipa je model koji svrstava sve primjere u većinsku klasu (engl. *most frequent class*; MFC) ili označuje testne primjere nasumično (engl. *random*). Proučite razred [`dummy.DummyClassifier`](http://scikit-learn.org/stable/modules/generated/sklearn.dummy.DummyClassifier.html) i pomoću njega stvorite spomenute osnovne klasifikatore. Opet ćete trebati iskoristiti cjevovod kako biste došli do vektorskog oblika ulaznih primjera, makar ovi osnovni klasifikatori koriste samo oznake pri predikciji.
from sklearn.dummy import DummyClassifier
# +
mfc_clf = Pipeline([('vectorizer', TfidfVectorizer(stop_words="english", ngram_range=(1, 2), max_features=500)),
('reducer', TruncatedSVD(n_components=300, random_state=69)),
('normalizer', Normalizer()),
('dummy_clf', DummyClassifier(strategy='most_frequent'))])
rnd_clf = Pipeline([('vectorizer', TfidfVectorizer(stop_words="english", ngram_range=(1, 2), max_features=500)),
('reducer', TruncatedSVD(n_components=300, random_state=69)),
('normalizer', Normalizer()),
('dummy_clf', DummyClassifier(strategy='uniform'))])
mfc_clf.fit(spam_X_train, spam_y_train)
rnd_clf.fit(spam_X_train, spam_y_train)
print(accuracy_score(spam_y_test, mfc_clf.predict(spam_X_test)))
print(accuracy_score(spam_y_test, rnd_clf.predict(spam_X_test)))
# -
# **Q:** Na temelju ovog primjera objasnite zašto točnost nije uvijek prikladna metrika.
# **Q:** Zašto koristimo F1-mjeru?
# **A:** Točnost je udio točno klasificiranih primjera u skupu svih primjera, a kako je u ovom primjeru udio klasa izrazito neuravnotežen u korist ne-spam klase, tako je i točnost MFC klasifikatora visoka.
#
# **A:** F1-mjera je, kao harmonijska sredina preciznosti i odziva, dobar indikator da li je jedna od te dvije mjere loša.
# #### (c)
# Međutim, provjera za kakvom smo posegli u prošlom podzadatku nije robusna. Stoga se u strojnom učenju obično koristi k-struka unakrsna provjera. Proučite razred [`model_selection.KFold`](http://scikit-learn.org/stable/modules/generated/sklearn.model_selection.KFold.html#sklearn.model_selection.KFold) i funkciju [`model_selection.cross_val_score`](http://scikit-learn.org/stable/modules/generated/sklearn.model_selection.cross_val_score.html#sklearn.model_selection.cross_val_score) te izračunajte procjenu pogreške na cijelom skupu podataka koristeći peterostruku unakrsnu provjeru.
#
# **NB:** Vaš model je sada cjevovod koji sadrži čitavo pretprocesiranje. Također, u nastavku ćemo se ograničiti na točnost, ali ovi postupci vrijede za sve metrike.
from sklearn.model_selection import cross_val_score, KFold
print(cross_val_score(clf, spam_X, spam_y, cv=5).mean())
# **Q:** Zašto "obična" unakrsna provjera nije dovoljno robusna?
# **Q:** Što je to stratificirana k-struka unakrsna provjera? Zašto ju često koristimo?
# **A:** Za razliku od obične, k-struka unakrsna provjera koristi svaki primjer i za učenje i za ispitivanje.
#
# **A:** Stratificirana k-struka unakrsna provjera osigurava zadržavanje omjera klasa u svakom preklopu, čime se izbjegava mogućnost da neka od klasa ne bude zastupljena u nekom preklopu.
# #### (d)
#
# Gornja procjena pogreške je u redu ako imamo već imamo model (bez ili s fiksiranim hiperparametrima). Međutim, mi želimo koristiti model koji ima optimalne vrijednosti hiperparametara te ih je stoga potrebno optimirati korištenjem pretraživanja po rešetci (engl. *grid search*). Očekivano, biblioteka `scikit-learn` već ima ovu funkcionalnost u razredu [`model_selection.GridSearchCV`](http://scikit-learn.org/stable/modules/generated/sklearn.model_selection.GridSearchCV.html). Jedina razlika vaše implementacije iz prošlih vježbi (npr. kod SVM-a) i ove jest ta da ova koristi k-struku unakrsnu provjeru.
#
# Prije optimizacije vrijednosti hiperparametara, očigledno moramo definirati i samu rešetku vrijednosti hiperparametara. Proučite kako se definira ista kroz rječnik u [primjeru](http://scikit-learn.org/stable/auto_examples/model_selection/grid_search_text_feature_extraction.html#sphx-glr-auto-examples-model-selection-grid-search-text-feature-extraction-py).
#
# Proučite spomenuti razred te pomoću njega pronađite i ispišite najbolje vrijednosti hiperparametara cjevovoda iz podzadatka (a): `max_features` $\in \{500, 1000\}$ i `n_components` $\in \{ 100, 200, 300 \}$ korištenjem pretraživanja po rešetci na skupu za učenje ($k=3$, kako bi išlo malo brže).
from sklearn.model_selection import GridSearchCV
# +
clf = Pipeline([('vectorizer', TfidfVectorizer(stop_words="english", ngram_range=(1, 2))),
('reducer', TruncatedSVD()),
('normalizer', Normalizer()),
('logistic-regression', LogisticRegression())])
parameters = {
'vectorizer__max_features': (500, 1000),
'reducer__n_components': (100, 200, 300),
}
grid_search = GridSearchCV(clf, parameters, cv=3)
grid_search.fit(spam_X_train, spam_y_train)
best_parameters = grid_search.best_estimator_.get_params()
for param_name in sorted(parameters.keys()):
print("%s: %r" % (param_name, best_parameters[param_name]))
# -
# **Q:** Koja se metrika optimira pri ovoj optimizaciji?
# **Q:** Kako biste odredili broj preklopa $k$?
# **A:** U ovom slučaju optimira se L2-regularizirana pogreška unakrsne entropije.
#
# **A:**
# #### (e)
# Ako želimo procijeniti pogrešku, ali pritom i napraviti odabir modela, tada se okrećemo ugniježđenoj k-strukoj unakrsnoj provjeri (engl. *nested k-fold cross validation*). U ovom zadatku ćete ju sami implementirati.
#
# Implementirajte funkciju `nested_kfold_cv(clf, param_grid, X, y, k1, k2)` koja provodi ugniježđenu k-struku unakrsnu provjeru. Argument `clf` predstavlja vaš klasifikator, `param_grid` rječnik vrijednosti hiperparametara (isto kao i u podzadatku (d)), `X` i `y` označeni skup podataka, a `k1` i `k2` broj preklopa u vanjskoj, odnosno unutarnjoj petlji. Poslužite se razredima [`model_selection.GridSearchCV`](http://scikit-learn.org/stable/modules/generated/sklearn.model_selection.GridSearchCV.html) i [`model_selection.KFold`](http://scikit-learn.org/stable/modules/generated/sklearn.model_selection.KFold.html).
#
# Funkcija vraća listu pogrešaka kroz preklope vanjske petlje.
from sklearn.model_selection import GridSearchCV, KFold
def nested_kfold_cv(clf, param_grid, X, y, k1, k2):
# Vaš kôd ovdje...
pass
# **Q:** Kako biste odabrali koji su hiperparametri generalno najbolji, a ne samo u svakoj pojedinačnoj unutarnjoj petlji?
# **Q:** Čemu u konačnici odgovara procjena generalizacijske pogreške?
# #### (f)
# Scenarij koji nas najviše zanima jest usporedba dvaju klasifikatora, odnosno, je li jedan od njih zaista bolji od drugog. Jedini način kako to možemo zaista potvrditi jest statističkom testom, u našem slučaju **uparenim t-testom**. Njime ćemo se baviti u ovom zadatku.
#
# Radi bržeg izvođenja, umjetno ćemo generirati podatke koji odgovaraju pogreškama kroz vanjske preklope dvaju klasifikatora (ono što bi vratila funkcija `nested_kfold_cv`):
# +
np.random.seed(1337)
C1_scores_5folds = np.random.normal(78, 4, 5)
C2_scores_5folds = np.random.normal(81, 2, 5)
C1_scores_10folds = np.random.normal(78, 4, 10)
C2_scores_10folds = np.random.normal(81, 2, 10)
C1_scores_50folds = np.random.normal(78, 4, 50)
C2_scores_50folds = np.random.normal(81, 2, 50)
# -
# Iskoristite ugrađenu funkciju [`scipy.stats.ttest_rel`](https://docs.scipy.org/doc/scipy-0.14.0/reference/generated/scipy.stats.ttest_rel.html) za provedbu uparenog t-testa i provjerite koji od ova modela je bolji kada se koristi 5, 10 i 50 preklopa.
from scipy.stats import ttest_rel
# +
# Vaš kôd ovdje...
# -
# **Q:** Koju hipotezu $H_0$ i alternativnu hipotezu $H_1$ testiramo ovim testom?
# **Q:** Koja pretpostavka na vjerojatnosnu razdiobu primjera je napravljena u gornjem testu? Je li ona opravdana?
# **Q:** Koji je model u konačnici bolji i je li ta prednost značajna uz $\alpha = 0.05$?
# ### 3. Grupiranje
# U ovom zadatku ćete se upoznati s algoritmom k-sredina (engl. *k-nearest neighbours*), njegovim glavnim nedostatcima te pretpostavkama. Također ćete isprobati i drugi algoritam grupiranja: model Gaussovih mješavina (engl. *Gaussian mixture model*).
# #### (a)
# Jedan od nedostataka algoritma k-sredina jest taj što unaprijed zahtjeva broj grupa ($K$) u koje će grupirati podatke. Ta informacija nam često nije dostupna (kao što nam nisu dostupne ni oznake primjera) te je stoga potrebno nekako izabrati najbolju vrijednost hiperparametra $K$. Jedan od naivnijih pristupa jest **metoda lakta/koljena** (engl. *elbow method*) koju ćete isprobati u ovom zadatku.
#
# U svojim rješenjima koristite ugrađenu implementaciju algoritma k-sredina, dostupnoj u razredu [`cluster.KMeans`](http://scikit-learn.org/stable/modules/generated/sklearn.cluster.KMeans.html).
#
# **NB**: Kriterijska funkcija algoritma k-sredina još se i naziva **inercijom** (engl. *inertia*). Za naučeni model, vrijednost kriterijske funkcije $J$ dostupna je kroz razredni atribut `inertia_`.
# +
from sklearn.datasets import make_blobs
Xp, yp = make_blobs(n_samples=300, n_features=2, centers=[[0, 0], [3, 2.5], [0, 4]],
cluster_std=[0.45, 0.3, 0.45], random_state=96)
plt.scatter(Xp[:,0], Xp[:,1], c=yp, cmap=plt.get_cmap("cool"), s=20)
# -
# Iskoristite skup podataka `Xp` dan gore. Isprobajte vrijednosti hiperparametra $K$ iz $[0,1,\ldots,15]$. Ne trebate dirati nikakve hiperparametre modela osim $K$. Iscrtajte krivulju od $J$ u ovisnosti o broju grupa $K$. Metodom lakta/koljena odredite vrijednost hiperparametra $K$.
from sklearn.cluster import KMeans
# +
no_clusters = np.arange(1, 16)
inertias = []
for k in no_clusters:
kmeans = KMeans(n_clusters=k)
kmeans.fit(Xp)
inertias.append(kmeans.inertia_)
plt.plot(no_clusters, inertias)
plt.xlabel(r'$K$')
plt.ylabel(r'$J$')
plt.show()
# -
# **Q:** Koju biste vrijednost hiperparametra $K$ izabrali na temelju ovog grafa? Zašto? Je li taj odabir optimalan? Kako to znate?
# **Q:** Je li ova metoda robusna?
# **Q:** Možemo li izabrati onaj $K$ koji minimizira pogrešku $J$? Objasnite.
# **A:** Metodom koljena biramo vrijednost $K$ u samom koljenu, a to je u gornjem slučaju $K=3$. Za ovaj odabir znamo da je optimalan jer je skup podataka dvodimenzionalan pa je lako uočiti prirodne grupe.
#
# **A:** Metoda nije suviše robusna, posebno ne za K-means s nasumično odabranim centroidima. Robusnost se može povećati uvišestručavanjem mjerenja na svakom $K$, pa traženjem koljena na uprosječenom prikazu kretanja kriterijske funkcije.
#
# **A:** $K$ koji minimizira $J$ biti će jednak broju primjera $N$, ukoliko prethodno postupak minimizacije ne zapne u lokalnom minimumu. U svakom slučaju bit će prenaučen, tako da je potrebno odabrati $K$ s najvećom sposobnošću generalizacije.
# #### (b)
# Odabir vrijednosti hiperparametra $K$ može se obaviti na mnoštvo načina. Pored metode lakta/koljena, moguće je isto ostvariti i analizom siluete (engl. *silhouette analysis*). Za to smo pripremili funkciju `mlutils.plot_silhouette` koja za dani broj grupa i podatke iscrtava prosječnu vrijednost koeficijenta siluete i vrijednost koeficijenta svakog primjera (kroz grupe).
#
# Vaš je zadatak isprobati različite vrijednosti hiperparametra $K$, $K \in \{2, 3, 5\}$ i na temelju dobivenih grafova odlučiti se za optimalan $K$.
# +
no_clusters = [2, 3, 5]
for k in no_clusters:
mlutils.plot_silhouette(k, Xp)
# -
# **Q:** Kako biste se gledajući ove slike odlučili za $K$?
# **Q:** Koji su problemi ovog pristupa?
# **A:** Analiza siluete daje najbolje rezultate za $K=3$. Siluete svih grupa prelaze prosjek, dok su slične u brojnosti.
#
# **A:** Nedostatak analize siluete (kao i metode lakta) je mjerenje isključivo globalnih karakteristika grupiranja, bez statističkog pristupa.
# #### (c)
# U ovom i sljedećim podzadatcima fokusirat ćemo se na temeljne pretpostavke algoritma k-srednjih vrijednosti te što se događa ako te pretpostavke nisu zadovoljene. Dodatno, isprobat ćemo i grupiranje modelom Gaussovih mješavina (engl. *Gaussian Mixture Models*; GMM) koji ne nema neke od tih pretpostavki.
#
# Prvo, krenite od podataka `X1`, koji su generirani korištenjem funkcije [`datasets.make_blobs`](http://scikit-learn.org/stable/modules/generated/sklearn.datasets.make_blobs.html), koja stvara grupe podataka pomoću izotropskih Gaussovih distribucija.
# +
from sklearn.datasets import make_blobs
X1, y1 = make_blobs(n_samples=1000, n_features=2, centers=[[0, 0], [1.3, 1.3]], cluster_std=[0.15, 0.5], random_state=96)
plt.scatter(X1[:,0], X1[:,1], c=y1, cmap=plt.get_cmap("cool"), s=20)
# -
# Naučite model k-srednjih vrijednosti (idealno pretpostavljajući $K=2$) na gornjim podatcima i prikažite dobiveno grupiranje (proučite funkciju [`scatter`](http://matplotlib.org/api/pyplot_api.html#matplotlib.pyplot.scatter), posebice argument `c`).
# +
kmeans = KMeans(n_clusters=2)
kmeans.fit(X1)
plt.scatter(X1[:,0], X1[:,1], c=kmeans.predict(X1))
plt.show()
# -
# **Q:** Što se dogodilo? Koja je pretpostavka algoritma k-srednjih vrijednosti ovdje narušena?
# **Q:** Što biste morali osigurati kako bi algoritam pronašao ispravne grupe?
# **A:** Algoritam *k-means* pretpostavlja podjednake varijance grupa, što ovdje nije slučaj.
#
# **A:** Varijance grupa bi se mogle minimizirati, iako je GMM bolje rješenje.
# #### (d)
#
# Isprobajte algoritam k-srednjih vrijednosti na podatcima generiranim korištenjem funkcije [`datasets.make_circles`](http://scikit-learn.org/stable/modules/generated/sklearn.datasets.make_circles.html), koja stvara dvije grupe podataka tako da je jedna unutar druge.
# +
from sklearn.datasets import make_circles
X2, y2 = make_circles(n_samples=1000, noise=0.15, factor=0.05, random_state=96)
plt.scatter(X2[:,0], X2[:,1], c=y2, cmap=plt.get_cmap("cool"), s=20)
# -
# Ponovno, naučite model k-srednjih vrijednosti (idealno pretpostavljajući $K=2$) na gornjim podatcima i prikažite dobiveno grupiranje (proučite funkciju [`scatter`](http://matplotlib.org/api/pyplot_api.html#matplotlib.pyplot.scatter), posebice argument `c`).
# +
kmeans = KMeans(n_clusters=2)
kmeans.fit(X2)
plt.scatter(X2[:,0], X2[:,1], c=kmeans.predict(X2))
plt.show()
# -
# **Q:** Što se dogodilo? Koja je pretpostavka algoritma k-srednjih vrijednosti ovdje narušena?
# **Q:** Što biste morali osigurati kako bi algoritam pronašao ispravne grupe?
# **A:** Grupe u primjeru nisu sferne.
#
# **A:** Podatke bi mogli transformirati u polarni sustav. Ili primjeniti HAC.
# #### (e)
# Završno, isprobat ćemo algoritam na sljedećem umjetno stvorenom skupu podataka:
# +
X31, y31 = make_blobs(n_samples=1000, n_features=2, centers=[[0, 0]], cluster_std=[0.2], random_state=69)
X32, y32 = make_blobs(n_samples=50, n_features=2, centers=[[0.7, 0.5]], cluster_std=[0.15], random_state=69)
X33, y33 = make_blobs(n_samples=600, n_features=2, centers=[[0.8, -0.4]], cluster_std=[0.2], random_state=69)
plt.scatter(X31[:,0], X31[:,1], c="#00FFFF", s=20)
plt.scatter(X32[:,0], X32[:,1], c="#F400F4", s=20)
plt.scatter(X33[:,0], X33[:,1], c="#8975FF", s=20)
# Just join all the groups in a single X.
X3 = np.vstack([X31, X32, X33])
y3 = np.hstack([y31, y32, y33])
# -
# Ponovno, naučite model k-srednjih vrijednosti (ovaj put idealno pretpostavljajući $K=3$) na gornjim podatcima i prikažite dobiveno grupiranje (proučite funkciju [`scatter`](http://matplotlib.org/api/pyplot_api.html#matplotlib.pyplot.scatter), posebice argument `c`).
# +
kmeans = KMeans(n_clusters=3)
kmeans.fit(X3)
plt.scatter(X3[:,0], X3[:,1], c=kmeans.predict(X3))
plt.show()
# -
# **Q:** Što se dogodilo? Koja je pretpostavka algoritma k-srednjih vrijednosti ovdje narušena?
# **Q:** Što biste morali osigurati kako bi algoritam pronašao ispravne grupe?
# **A:** Grupe sadrže različit broj primjera, tj. neuravnoteženih su veličina.
#
# https://stats.stackexchange.com/a/133694
# #### (f)
# Sada kada ste se upoznali s ograničenjima algoritma k-srednjih vrijednosti, isprobat ćete grupiranje modelom mješavine Gaussa (*Gaussian Mixture Models; GMM*), koji je generalizacija algoritma k-srednjih vrijednosti (odnosno, algoritam k-srednjih vrijednosti specijalizacija je GMM-a). Implementacija ovog modela dostupna je u [`mixture.GaussianMixture`](http://scikit-learn.org/stable/modules/generated/sklearn.mixture.GaussianMixture.html#sklearn.mixture.GaussianMixture). Isprobajte ovaj model (s istim pretpostavkama o broju grupa) na podacima iz podzadataka (a)-(c). Ne morate mijenjati nikakve hiperparametre ni postavke osim broja komponenti.
from sklearn.mixture import GaussianMixture
# +
gmm = GaussianMixture(n_components=2)
gmm.fit(X1)
plt.scatter(X1[:,0], X1[:,1], c=gmm.predict(X1))
plt.show()
# +
gmm = GaussianMixture(n_components=2)
gmm.fit(X2)
plt.scatter(X2[:,0], X2[:,1], c=gmm.predict(X2))
plt.show()
# +
gmm = GaussianMixture(n_components=3)
gmm.fit(X3)
plt.scatter(X3[:,0], X3[:,1], c=gmm.predict(X3))
plt.show()
# -
# #### (g)
# Kako vrednovati točnost modela grupiranja ako imamo stvarne oznake svih primjera (a u našem slučaju imamo, jer smo mi ti koji smo generirali podatke)? Često korištena mjera jest **Randov indeks** koji je zapravo pandan točnosti u zadatcima klasifikacije. Implementirajte funkciju `rand_index_score(y_gold, y_predict)` koja ga računa. Funkcija prima dva argumenta: listu stvarnih grupa kojima primjeri pripadaju (`y_gold`) i listu predviđenih grupa (`y_predict`). Dobro će vam doći funkcija [`itertools.combinations`](https://docs.python.org/2/library/itertools.html#itertools.combinations).
# +
import itertools as it
y_gold = np.array([0, 0, 0, 0, 1, 0, 0, 1, 1, 1, 1, 1, 1])
y_predict = np.array([0, 0, 0, 0, 0, 1, 1, 1, 1, 1, 1, 1, 1])
def rand_index_score(y_gold, y_predict):
ns_corr_per_cluster = np.unique(y_gold[y_gold == y_predict], return_counts=True)[1]
ns_fail_per_cluster = np.unique(y_gold[y_gold != y_predict], return_counts=True)[1]
n_unique_per_cluster = np.concatenate((ns_corr_per_cluster, ns_fail_per_cluster))
n_same_pairs_same_clusters = np.sum([len(list(it.combinations(range(a), 2))) for a in n_unique_per_cluster])
n_diff_pairs_diff_clusters = np.prod(ns_corr_per_cluster) + np.prod(ns_fail_per_cluster)
return (n_same_pairs_same_clusters + n_diff_pairs_diff_clusters) / len(list((it.combinations(range(len(y_gold)), 2))))
rand_index_score(y_gold, y_predict)
# -
# **Q:** Zašto je Randov indeks pandan točnosti u klasifikacijskim problemima?
# **Q:** Koji su glavni problemi ove metrike?
# **Q:** Zašto se koristi modificirani Randov indeks (engl. *adjusted Rand index*)?
# **Q:** Kako vrednovati kvalitetu grupiranja ako nenamo stvarne oznake primjera? Je li to uopće moguće?
# **A:** Randov indeks je točnost parova primjera, koja se koristi za skupove podataka s nedefiniranim oznakama klasa, jer je nepromjenjiva pri preimenovanju grupa.
#
# **A:** Kao i s točnosti, ako je distribucija klasa vrlo neuravnotežena RI može uzimati visoke vrijednosti za nasumične klasifikatore.
#
# **A:** ARI skaliranjem na $[-1,1]$ ublažava pogrešna tumačenja RI kao u situaciji iz prethodnog pitanja, iznos blizak nuli predstavlja nasumično grupiranje.
#
# **A:** Kvaliteta grupiranja bez stvarnog skupa oznaka uglavnom se mjeri internim indeksima (enlg. *internal indices*, e.g. *Silhouette coefficient*), koji validiraju grupiranje pomoću značajki inherentnih samom skupu. Pri korištenju ovih mjera treba voditi računa o korištenom modelu i vrijednostima koje on optimizira, da se izbjegne potecijalna prenaučenost ukoliko interni indeks koristi upravo te iste vrijednosti za izračun mjere kvalitete.
| SU-2017-LAB05-PGM-NB-grupiranje-vrednovanje.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from scipy.io import loadmat
from skimage import io
# +
def findClosestCentroids(X, centroids):
"""
output a one-dimensional array idx that holds the
index of the closest centroid to every training example.
"""
idx = []
max_dist = 1000000 # 限制一下最大距离
for i in range(len(X)):
minus = X[i] - centroids # here use numpy's broadcasting
dist = minus[:,0]**2 + minus[:,1]**2
if dist.min() < max_dist:
ci = np.argmin(dist)
idx.append(ci)
return np.array(idx)
def computeCentroids(X, idx):
centroids = []
for i in range(len(np.unique(idx))): # np.unique() means K
u_k = X[idx==i].mean(axis=0) # 求每列的平均值
centroids.append(u_k)
return np.array(centroids)
def plotData(X, centroids, idx=None):
"""
可视化数据,并自动分开着色。
idx: 最后一次迭代生成的idx向量,存储每个样本分配的簇中心点的值
centroids: 包含每次中心点历史记录
"""
colors = ['b','g','gold','darkorange','salmon','olivedrab',
'maroon', 'navy', 'sienna', 'tomato', 'lightgray', 'gainsboro'
'coral', 'aliceblue', 'dimgray', 'mintcream', 'mintcream']
assert len(centroids[0]) <= len(colors), 'colors not enough '
subX = [] # 分号类的样本点
if idx is not None:
for i in range(centroids[0].shape[0]):
x_i = X[idx == i]
subX.append(x_i)
else:
subX = [X] # 将X转化为一个元素的列表,每个元素为每个簇的样本集,方便下方绘图
# 分别画出每个簇的点,并着不同的颜色
plt.figure(figsize=(8,5))
for i in range(len(subX)):
xx = subX[i]
plt.scatter(xx[:,0], xx[:,1], c=colors[i], label='Cluster %d'%i)
plt.legend()
plt.grid(True)
plt.xlabel('x1',fontsize=14)
plt.ylabel('x2',fontsize=14)
plt.title('Plot of X Points',fontsize=16)
# 画出簇中心点的移动轨迹
xx, yy = [], []
for centroid in centroids:
xx.append(centroid[:,0])
yy.append(centroid[:,1])
plt.plot(xx, yy, 'rx--', markersize=8)
def initCentroids(X, K):
m, n = X.shape
idx = np.random.choice(m, K)
centroids = X[idx]
return centroids
def runKmeans(X, centroids, max_iters):
K = len(centroids)
centroids_all = []
centroids_all.append(centroids)
centroid_i = centroids
for i in range(max_iters):
idx = findClosestCentroids(X, centroid_i)
centroid_i = computeCentroids(X, idx)
centroids_all.append(centroid_i)
return idx, centroids_all
mat = loadmat('ex7data2.mat')
X = mat['X']
init_centroids = initCentroids(X,3)
idx, centroids_all = runKmeans(X, init_centroids, 20)
plotData(X, centroids_all, idx)
# +
A = io.imread('bird_small.png')
plt.imshow(A);
A = A/255.
X = A.reshape(-1, 3)
K = 16
centroids = initCentroids(X, K)
idx, centroids_all = runKmeans(X, centroids, 10)
img = np.zeros(X.shape)
centroids = centroids_all[-1]
for i in range(len(centroids)):
img[idx == i] = centroids[i]
img = img.reshape((128, 128, 3))
fig, axes = plt.subplots(1, 2, figsize=(12,6))
axes[0].imshow(A)
axes[1].imshow(img)
# -
| ex7/7.K-means.ipynb |