code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: TensorFlow-1.13.1
# language: python
# name: tensorflow-1.13.1
# ---
# # 训练轮数与callbacks
# 在本例中,通过比较**同样结构**,**同样优化器**,**同样数据集**下**不同epoch**以及**不同callbacks方法**下训练的值得出训练轮数与callbacks方法对训练的影响。
#
# 训练轮数决定了训练达到的程度,在接下来的实验中,我们尝试了逐步增加训练轮数,观察模型在不同阶段的收敛情况。以5轮次为单位进行观察。
# #### 注意: 每一个5轮次训练大约耗时20分钟。
# callbacks方法中,我们将介绍:
# - ModelCheckpoint
# - EarlyStopping
# - ReduceLROnPlateau
#
# 实验之前我们进行keras,keras_applications版本配置以及数据集下载。
# !pip install --upgrade keras_applications==1.0.6 keras==2.2.4
import os
if os.path.exists('./data') == False:
from modelarts.session import Session
session = Session()
if session.region_name == 'cn-north-1':
bucket_path="modelarts-labs/end2end/image_recognition/dog_and_cat_25000.tar.gz"
elif session.region_name == 'cn-north-4':
bucket_path="modelarts-labs-bj4/end2end/image_recognition/dog_and_cat_25000.tar.gz"
else:
print("请更换地区到北京一或北京四")
session.download_data(
bucket_path=bucket_path,
path="./dog_and_cat_25000.tar.gz")
# 使用tar命令解压资源包
# !tar xf ./dog_and_cat_25000.tar.gz
# 清理压缩包
# !rm -f ./dog_and_cat_25000.tar.gz
# !mkdir model
# ## 引入相关的包
# +
from keras.applications.vgg16 import VGG16
from keras.preprocessing import image
import numpy as np
from keras.preprocessing import image
from keras.models import Model
from keras.layers import Dense, GlobalAveragePooling2D
from keras import backend as K
from keras.models import load_model
from keras.preprocessing.image import ImageDataGenerator
# -
# ## 读取数据
import os
from PIL import Image
def load_data():
dirname = "./data"
path = "./data"
num_train_samples = 25000
x_train = np.empty((num_train_samples, 224,224,3), dtype='uint8')
y_train = np.empty((num_train_samples,1), dtype='uint8')
index = 0
for file in os.listdir("./data"):
image = Image.open(os.path.join(dirname,file)).resize((224,224))
image = np.array(image)
x_train[index,:,:,:] = image
if "cat" in file:
y_train[index,0] =1
elif "dog" in file:
y_train[index,0] =0
index += 1
return (x_train, y_train)
(x_train, y_train) = load_data()
print(x_train.shape)
print(y_train.shape)
# ## 数据处理
from keras.utils import np_utils
def process_data(x_train,y_train):
x_train = x_train.astype(np.float32)
x_train /= 255
n_classes = 2
y_train = np_utils.to_categorical(y_train, n_classes)
return x_train,y_train
x_train,y_train= process_data(x_train,y_train)
print(x_train.shape)
print(y_train.shape)
# ## 构建模型
def build_model(base_model):
x = base_model.output
x = GlobalAveragePooling2D()(x)
predictions = Dense(2, activation='softmax')(x)
model = Model(inputs=base_model.input, outputs=predictions)
print(type(model))
return model
base_model = VGG16(weights=None, include_top=False)
model = build_model(base_model)
model.summary()
# ## 模型训练
# ### 5 epoch训练
import keras
opt = keras.optimizers.rmsprop(lr=0.0001, decay=1e-6)
model.compile(loss='binary_crossentropy',
optimizer=opt,
metrics=['accuracy'])
from keras.callbacks import ModelCheckpoint, EarlyStopping, ReduceLROnPlateau
es = EarlyStopping(monitor='val_acc', min_delta=0.001, patience=5, verbose=1, mode='auto')
cp = ModelCheckpoint(filepath="./model/ckp_vgg16_dog_and_cat.h5", monitor="val_acc", verbose=1, save_best_only=True, mode="auto", period=1)
lr = ReduceLROnPlateau(monitor="val_acc", factor=0.1, patience=3, verbose=1, mode="auto", min_lr=0)
callbacks = [es,cp,lr]
history = model.fit(x=x_train,
y=y_train,
batch_size=16,
epochs=5,
verbose=1,
callbacks=callbacks,
validation_split=0.25,
shuffle=True,
initial_epoch=0,
)
# +
import matplotlib.pyplot as plt
# 绘制训练 & 验证的准确率值
plt.plot(history.history['acc'])
plt.plot(history.history['val_acc'])
plt.title('Model accuracy')
plt.ylabel('Accuracy')
plt.xlabel('Epoch')
plt.legend(['Train', 'Test'], loc='upper left')
plt.show()
# -
# 绘制训练 & 验证的损失值
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('Model loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.legend(['Train', 'Test'], loc='upper left')
plt.show()
# ### 10 epoch 训练
# **在下面的训练中,epoch将再训练5轮**
# #### Epoch
#
# 在Epoch数值较小时,出现了上一个训练中出现的欠拟合情况,模型没有很好收敛训练便结束了。在接下来的训练中,我们提高了epoch的值,通过训练结果我们可以看到模型逐渐收敛。但是epoch的值并非越大越好,过大的epoch值可能会导致过拟合现象。
#
# 
#
# epoch的值没有具体的公式进行计算,需要根据经验和具体的情况进行制定。更多的epoch训练大家可以在拓展中进行尝试。
# #### ModelCheckpoint
# 在模型训练过程中,ModelCheckpoint将出现的最好的权重进行保存。
# 在下面的训练中,每一次出现更好的模型,epoch完成后都进行了保存。
# #### ReduceLROnPlateau
# 学习率衰减方法,在指定epoch数量结束后检测指标时候有提升,如果提升较小,便进行学习率衰减。在接下来的训练中,通过`ReduceLROnPlateau`方法,学习率进行了多次调整,调整之后的模型的指标有所提升。
history_more_steps = model.fit(x=x_train,
y=y_train,
batch_size=16,
epochs=5,
verbose=1,
callbacks=callbacks,
validation_split=0.25,
shuffle=True,
initial_epoch=0,
)
# +
import matplotlib.pyplot as plt
# 绘制训练 & 验证的准确率值
plt.plot(history_more_steps.history['acc'])
plt.plot(history_more_steps.history['val_acc'])
plt.title('Model accuracy')
plt.ylabel('Accuracy')
plt.xlabel('Epoch')
plt.legend(['Train', 'Test'], loc='upper left')
plt.show()
# -
# 绘制训练 & 验证的损失值
plt.plot(history_more_steps.history['loss'])
plt.plot(history_more_steps.history['val_loss'])
plt.title('Model loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.legend(['Train', 'Test'], loc='upper left')
plt.show()
# #### EarlyStopping 早停法
# 根据上一个训练可以看到,在训练的后期模型的loss和acc数值已经稳定,这时候继续训练没有对数值产生影响还有可能产生过拟合情况,所以需要及时将模型训练停止。EarlyStopping(早停法)检测模型的某一项指标,如果在指定步数中指标没有提升,则将模型训练停止。
#
# 在下面的训练中如果训练没有显著提升,则停止训练。
#
# #### 注意:可以修改早停参数,进行模型训练停止。当前模型训练上限为20轮次,如果模型val_acc没有0.001提升则停止。
es = EarlyStopping(monitor='val_acc', min_delta=0.001, patience=1, verbose=1, mode='auto')
cp = ModelCheckpoint(filepath="./model/ckp_vgg16_dog_and_cat.h5", monitor="val_acc", verbose=1, save_best_only=True, mode="auto", period=1)
lr = ReduceLROnPlateau(monitor="val_acc", factor=0.1, patience=3, verbose=1, mode="auto", min_lr=0)
callbacks = [es,cp,lr]
history_steps_15 = model.fit(x=x_train,
y=y_train,
batch_size=16,
epochs=20,
verbose=1,
callbacks=callbacks,
validation_split=0.25,
shuffle=True,
initial_epoch=0,
)
# 绘制训练 & 验证的损失值
plt.plot(history_steps_15.history['loss'])
plt.plot(history_steps_15.history['val_loss'])
plt.title('Model loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.legend(['Train', 'Test'], loc='upper left')
plt.show()
# 绘制训练 & 验证的准确率值
plt.plot(history_steps_15.history['acc'])
plt.plot(history_steps_15.history['val_acc'])
plt.title('Model accuracy')
plt.ylabel('Accuracy')
plt.xlabel('Epoch')
plt.legend(['Train', 'Test'], loc='upper left')
plt.show()
# # 拓展
#
# - 可以尝试自己定义学习率衰减规律。使用`LearningRateScheduler`方法,自己定义学习率衰减。
| notebook/DL_image_hyperparameter_tuning/00_epoch_callbacks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from neuron.mnist import mnist_loader
import neuron.mnist.network_standard as network
import matplotlib.pyplot as plt
import matplotlib.cm as cm
import numpy as np
import pandas as pd
import seaborn as sns
from neuron.activation_functions import sigmoid
from neuron.mnist import stepik_original as stepik
np.set_printoptions(precision=5)
% load_ext autoreload
% autoreload 2
# -
# ### The networks are fully reconciled. With full data set you may even use different mnist parser. The final weights should be reconciled
# # PARSE DATA
# +
import os
os.chdir("C:\\Users\\mkapchenko\\Dropbox\\perso\\GitHub\\Neuron\\notebooks")
training_data, validation_data, test_data = mnist_loader.load_data_wrapper()
n = 50000
nepochs = 30
nb_batches = 10
batch_size = 784//nb_batches
mini_res = list(training_data)[0:n]
mini_test = list(test_data)[0:n]
# to have the same data
minX = np.array(mini_res[0][0])
miny = np.array(mini_res[0][1])
# +
# %time
for example in range(1, len(mini_res)):
minX = np.concatenate((minX, mini_res[example][0]), axis=1)
miny = np.concatenate((miny, mini_res[example][1]), axis=1)
minX, miny = minX.T, miny.T
# -
(X, y), (validation_X, validation_y), (test_X, test_y) = mnist_loader.perf_load_data_wrapper()
# # LEARNING
# ### Neuron
# +
# %%time
# 28 s
# minX, miny = X[0:n], y[0:n]
netw = network.Network([784, 30, 10])
netw.SGD(X, y, epochs=nepochs, batch_size=batch_size, learning_rate = 3.);
# -
# ### Stepik
# %%time
# 9min 9s
# Stepik network
netstepik = stepik.Network([784, 30, 10])
netstepik.SGD(mini_res, epochs=nepochs, mini_batch_size=batch_size, eta=3.0, test_data=mini_test)
# # RECON
# ### Final weights
for layer in range(2):
print(f'Layer {layer} recon: ' , (netstepik.weights[layer] - netw.weights[layer]).mean())
# ### Gradient by epoch
# Should be reconciled if the mnist data is in the same order for two networks
for epoch in range(nepochs):
print(f' *** epoch = {epoch} ***')
print('Intercept gradient last layer ', (netstepik.nabla_b[epoch][1] * 1/n - netw.debug_djdb_by_batch[epoch][1]).mean())
print('Intercept gradient first layer', (netstepik.nabla_b[epoch][0] * 1/n - netw.debug_djdb_by_batch[epoch][0]).mean())
print('Weights gradient last layer ', (netstepik.nabla_w[epoch][1] * 1/n - netw.debug_djdw_by_batch[epoch][1]).mean())
print('Weights gradient first layer ', (netstepik.nabla_w[epoch][0] * 1/n - netw.debug_djdw_by_batch[epoch][0]).mean())
print('')
sns.heatmap(netw.weights[0][27].reshape(28,28))
| notebooks/gold standard/recon.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ### 蒸馏通道裁剪模型示例
# 本示例介绍使用更高精度的[YOLOv3-ResNet34](../../configs/yolov3_r34.yml)模型蒸馏经通道裁剪的[YOLOv3-MobileNet](../../configs/yolov3_mobilenet_v1.yml)模型。脚本可参照蒸馏脚本[distill.py](../distillation/distill.py)和通道裁剪脚本[prune.py](../prune/prune.py)简单修改得到,蒸馏过程采用细粒度损失来蒸馏YOLOv3输出层特征图。
# 切换到PaddleDetection根目录,设置环境变量
% cd ../..
# 导入依赖包,注意须同时导入蒸馏和通道裁剪的相关接口
# +
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import os
import numpy as np
from collections import OrderedDict
from paddleslim.dist.single_distiller import merge, l2_loss
from paddleslim.prune import Pruner
from paddleslim.analysis import flops
from paddle import fluid
from ppdet.core.workspace import load_config, merge_config, create
from ppdet.data.reader import create_reader
from ppdet.utils.eval_utils import parse_fetches, eval_results, eval_run
from ppdet.utils.stats import TrainingStats
from ppdet.utils.cli import ArgsParser
from ppdet.utils.check import check_gpu
import ppdet.utils.checkpoint as checkpoint
# -
# 定义细粒度的蒸馏损失函数
def split_distill(split_output_names, weight):
"""
Add fine grained distillation losses.
Each loss is composed by distill_reg_loss, distill_cls_loss and
distill_obj_loss
"""
student_var = []
for name in split_output_names:
student_var.append(fluid.default_main_program().global_block().var(
name))
s_x0, s_y0, s_w0, s_h0, s_obj0, s_cls0 = student_var[0:6]
s_x1, s_y1, s_w1, s_h1, s_obj1, s_cls1 = student_var[6:12]
s_x2, s_y2, s_w2, s_h2, s_obj2, s_cls2 = student_var[12:18]
teacher_var = []
for name in split_output_names:
teacher_var.append(fluid.default_main_program().global_block().var(
'teacher_' + name))
t_x0, t_y0, t_w0, t_h0, t_obj0, t_cls0 = teacher_var[0:6]
t_x1, t_y1, t_w1, t_h1, t_obj1, t_cls1 = teacher_var[6:12]
t_x2, t_y2, t_w2, t_h2, t_obj2, t_cls2 = teacher_var[12:18]
def obj_weighted_reg(sx, sy, sw, sh, tx, ty, tw, th, tobj):
loss_x = fluid.layers.sigmoid_cross_entropy_with_logits(
sx, fluid.layers.sigmoid(tx))
loss_y = fluid.layers.sigmoid_cross_entropy_with_logits(
sy, fluid.layers.sigmoid(ty))
loss_w = fluid.layers.abs(sw - tw)
loss_h = fluid.layers.abs(sh - th)
loss = fluid.layers.sum([loss_x, loss_y, loss_w, loss_h])
weighted_loss = fluid.layers.reduce_mean(loss *
fluid.layers.sigmoid(tobj))
return weighted_loss
def obj_weighted_cls(scls, tcls, tobj):
loss = fluid.layers.sigmoid_cross_entropy_with_logits(
scls, fluid.layers.sigmoid(tcls))
weighted_loss = fluid.layers.reduce_mean(
fluid.layers.elementwise_mul(
loss, fluid.layers.sigmoid(tobj), axis=0))
return weighted_loss
def obj_loss(sobj, tobj):
obj_mask = fluid.layers.cast(tobj > 0., dtype="float32")
obj_mask.stop_gradient = True
loss = fluid.layers.reduce_mean(
fluid.layers.sigmoid_cross_entropy_with_logits(sobj, obj_mask))
return loss
distill_reg_loss0 = obj_weighted_reg(s_x0, s_y0, s_w0, s_h0, t_x0, t_y0,
t_w0, t_h0, t_obj0)
distill_reg_loss1 = obj_weighted_reg(s_x1, s_y1, s_w1, s_h1, t_x1, t_y1,
t_w1, t_h1, t_obj1)
distill_reg_loss2 = obj_weighted_reg(s_x2, s_y2, s_w2, s_h2, t_x2, t_y2,
t_w2, t_h2, t_obj2)
distill_reg_loss = fluid.layers.sum(
[distill_reg_loss0, distill_reg_loss1, distill_reg_loss2])
distill_cls_loss0 = obj_weighted_cls(s_cls0, t_cls0, t_obj0)
distill_cls_loss1 = obj_weighted_cls(s_cls1, t_cls1, t_obj1)
distill_cls_loss2 = obj_weighted_cls(s_cls2, t_cls2, t_obj2)
distill_cls_loss = fluid.layers.sum(
[distill_cls_loss0, distill_cls_loss1, distill_cls_loss2])
distill_obj_loss0 = obj_loss(s_obj0, t_obj0)
distill_obj_loss1 = obj_loss(s_obj1, t_obj1)
distill_obj_loss2 = obj_loss(s_obj2, t_obj2)
distill_obj_loss = fluid.layers.sum(
[distill_obj_loss0, distill_obj_loss1, distill_obj_loss2])
loss = (distill_reg_loss + distill_cls_loss + distill_obj_loss) * weight
return loss
# 读取配置文件,设置use_fined_grained_loss=True
cfg = load_config("./configs/yolov3_mobilenet_v1.yml")
merge_config({'use_fine_grained_loss': True})
# 创建执行器
devices_num = fluid.core.get_cuda_device_count()
place = fluid.CUDAPlace(0)
# devices_num = int(os.environ.get('CPU_NUM', 1))
# place = fluid.CPUPlace()
exe = fluid.Executor(place)
# 构造训练模型和reader
# +
main_arch = cfg.architecture
# build program
model = create(main_arch)
inputs_def = cfg['TrainReader']['inputs_def']
train_feed_vars, train_loader = model.build_inputs(**inputs_def)
train_fetches = model.train(train_feed_vars)
loss = train_fetches['loss']
start_iter = 0
train_reader = create_reader(cfg.TrainReader, (cfg.max_iters - start_iter) * devices_num, cfg)
train_loader.set_sample_list_generator(train_reader, place)
# -
# 构造评估模型和reader
# +
eval_prog = fluid.Program()
with fluid.program_guard(eval_prog, fluid.default_startup_program()):
with fluid.unique_name.guard():
model = create(main_arch)
inputs_def = cfg['EvalReader']['inputs_def']
test_feed_vars, eval_loader = model.build_inputs(**inputs_def)
fetches = model.eval(test_feed_vars)
eval_prog = eval_prog.clone(True)
eval_reader = create_reader(cfg.EvalReader)
eval_loader.set_sample_list_generator(eval_reader, place)
# -
# 构造teacher模型并导入权重
# +
teacher_cfg = load_config("./configs/yolov3_r34.yml")
merge_config({'use_fine_grained_loss': True})
teacher_arch = teacher_cfg.architecture
teacher_program = fluid.Program()
teacher_startup_program = fluid.Program()
with fluid.program_guard(teacher_program, teacher_startup_program):
with fluid.unique_name.guard():
teacher_feed_vars = OrderedDict()
for name, var in train_feed_vars.items():
teacher_feed_vars[name] = teacher_program.global_block(
)._clone_variable(
var, force_persistable=False)
model = create(teacher_arch)
train_fetches = model.train(teacher_feed_vars)
teacher_loss = train_fetches['loss']
exe.run(teacher_startup_program)
checkpoint.load_params(exe, teacher_program, "https://paddlemodels.bj.bcebos.com/object_detection/yolov3_r34.tar")
teacher_program = teacher_program.clone(for_test=True)
# -
# 合并program
data_name_map = {
'target0': 'target0',
'target1': 'target1',
'target2': 'target2',
'image': 'image',
'gt_bbox': 'gt_bbox',
'gt_class': 'gt_class',
'gt_score': 'gt_score'
}
merge(teacher_program, fluid.default_main_program(), data_name_map, place)
# 构造蒸馏损失和优化器
# +
yolo_output_names = [
'strided_slice_0.tmp_0', 'strided_slice_1.tmp_0',
'strided_slice_2.tmp_0', 'strided_slice_3.tmp_0',
'strided_slice_4.tmp_0', 'transpose_0.tmp_0', 'strided_slice_5.tmp_0',
'strided_slice_6.tmp_0', 'strided_slice_7.tmp_0',
'strided_slice_8.tmp_0', 'strided_slice_9.tmp_0', 'transpose_2.tmp_0',
'strided_slice_10.tmp_0', 'strided_slice_11.tmp_0',
'strided_slice_12.tmp_0', 'strided_slice_13.tmp_0',
'strided_slice_14.tmp_0', 'transpose_4.tmp_0'
]
distill_loss = split_distill(yolo_output_names, 1000)
loss = distill_loss + loss
lr_builder = create('LearningRate')
optim_builder = create('OptimizerBuilder')
lr = lr_builder()
opt = optim_builder(lr)
opt.minimize(loss)
# -
# 导入待裁剪模型裁剪前全部权重
exe.run(fluid.default_startup_program())
checkpoint.load_params(exe, fluid.default_main_program(), "https://paddlemodels.bj.bcebos.com/object_detection/yolov3_mobilenet_v1.tar")
# 裁剪训练和评估program
# +
pruned_params = ["yolo_block.0.0.0.conv.weights",
"yolo_block.0.0.1.conv.weights",
"yolo_block.0.1.0.conv.weights",
"yolo_block.0.1.1.conv.weights",
"yolo_block.0.2.conv.weights",
"yolo_block.0.tip.conv.weights",
"yolo_block.1.0.0.conv.weights",
"yolo_block.1.0.1.conv.weights",
"yolo_block.1.1.0.conv.weights",
"yolo_block.1.1.1.conv.weights",
"yolo_block.1.2.conv.weights",
"yolo_block.1.tip.conv.weights",
"yolo_block.2.0.0.conv.weights",
"yolo_block.2.0.1.conv.weights",
"yolo_block.2.1.0.conv.weights",
"yolo_block.2.1.1.conv.weights",
"yolo_block.2.2.conv.weights",
"yolo_block.2.tip.conv.weights"]
pruned_ratios = [0.5] * 6 + [0.7] * 6 + [0.8] * 6
print("pruned params: {}".format(pruned_params))
print("pruned ratios: {}".format(pruned_ratios))
pruner = Pruner()
distill_prog = pruner.prune(
fluid.default_main_program(),
fluid.global_scope(),
params=pruned_params,
ratios=pruned_ratios,
place=place,
only_graph=False)[0]
base_flops = flops(eval_prog)
eval_prog = pruner.prune(
eval_prog,
fluid.global_scope(),
params=pruned_params,
ratios=pruned_ratios,
place=place,
only_graph=True)[0]
pruned_flops = flops(eval_prog)
print("FLOPs -{}; total FLOPs: {}; pruned FLOPs: {}".format(float(base_flops - pruned_flops)/base_flops, base_flops, pruned_flops))
# -
# 编译训练和评估program
# +
build_strategy = fluid.BuildStrategy()
build_strategy.fuse_all_reduce_ops = False
build_strategy.fuse_all_optimizer_ops = False
build_strategy.fuse_elewise_add_act_ops = True
# only enable sync_bn in multi GPU devices
sync_bn = getattr(model.backbone, 'norm_type', None) == 'sync_bn'
build_strategy.sync_batch_norm = sync_bn and devices_num > 1 \
and cfg.use_gpu
exec_strategy = fluid.ExecutionStrategy()
# iteration number when CompiledProgram tries to drop local execution scopes.
# Set it to be 1 to save memory usages, so that unused variables in
# local execution scopes can be deleted after each iteration.
exec_strategy.num_iteration_per_drop_scope = 1
parallel_main = fluid.CompiledProgram(distill_prog).with_data_parallel(
loss_name=loss.name,
build_strategy=build_strategy,
exec_strategy=exec_strategy)
compiled_eval_prog = fluid.compiler.CompiledProgram(eval_prog)
# -
# 开始训练
# +
# parse eval fetches
extra_keys = []
if cfg.metric == 'COCO':
extra_keys = ['im_info', 'im_id', 'im_shape']
if cfg.metric == 'VOC':
extra_keys = ['gt_bbox', 'gt_class', 'is_difficult']
eval_keys, eval_values, eval_cls = parse_fetches(fetches, eval_prog,
extra_keys)
# whether output bbox is normalized in model output layer
is_bbox_normalized = False
map_type = cfg.map_type if 'map_type' in cfg else '11point'
best_box_ap_list = [0.0, 0] #[map, iter]
save_dir = os.path.join(cfg.save_dir, 'yolov3_mobilenet_v1')
train_loader.start()
for step_id in range(start_iter, cfg.max_iters):
teacher_loss_np, distill_loss_np, loss_np, lr_np = exe.run(
parallel_main,
fetch_list=[
'teacher_' + teacher_loss.name, distill_loss.name, loss.name,
lr.name
])
if step_id % 20 == 0:
print(
"step {} lr {:.6f}, loss {:.6f}, distill_loss {:.6f}, teacher_loss {:.6f}".
format(step_id, lr_np[0], loss_np[0], distill_loss_np[0],
teacher_loss_np[0]))
if step_id % cfg.snapshot_iter == 0 and step_id != 0 or step_id == cfg.max_iters - 1:
save_name = str(
step_id) if step_id != cfg.max_iters - 1 else "model_final"
checkpoint.save(exe,
distill_prog,
os.path.join(save_dir, save_name))
# eval
results = eval_run(exe, compiled_eval_prog, eval_loader, eval_keys,
eval_values, eval_cls)
resolution = None
box_ap_stats = eval_results(results, cfg.metric, cfg.num_classes,
resolution, is_bbox_normalized,
FLAGS.output_eval, map_type,
cfg['EvalReader']['dataset'])
if box_ap_stats[0] > best_box_ap_list[0]:
best_box_ap_list[0] = box_ap_stats[0]
best_box_ap_list[1] = step_id
checkpoint.save(exe,
distill_prog,
os.path.join("./", "best_model"))
print("Best test box ap: {}, in step: {}".format(
best_box_ap_list[0], best_box_ap_list[1]))
train_loader.reset()
# -
# 我们也提供了一键式启动蒸馏通道裁剪模型训练脚本[distill_pruned_model.py](./distill_pruned_model.py)和蒸馏通道裁剪模型库,请参考[蒸馏通道裁剪模型](./README.md)
| slim/extensions/distill_pruned_model/distill_pruned_model_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Measuring Success: Splitting up the data for train, validation, and test set
#
# Split the dataset up into the following segments:
# 1. Training Data: 60%
# 2. Validation Data: 20%
# 3. Test Data: 20%
# ### Read in data
# +
import pandas as pd
from sklearn.model_selection import train_test_split
titanic_df = pd.read_csv('../Data/titanic.csv')
titanic_df.head()
# -
# ### Split into train, validation, and test set
# as Survived column is label, we will have to remove it from features and also want to split it alone
features = titanic_df.drop(['Survived'], axis=1)
labels = titanic_df['Survived']
# - we want to split train data 60%, validation data 20%, test data 20%
# - but as sklearn can only split data into two parts, we will have to do this twice
# first split : train data 60%, test data 40%
X_train, X_test, y_train, y_test = train_test_split(features, labels, test_size=0.4, random_state=42)
# second split again on test data: validation data 50%, test data 50%, half half
X_test, X_val, y_test, y_val = train_test_split(X_test, y_test, test_size=0.5, random_state=42 )
# make quick check
print(len(labels), len(y_train), len(y_val), len(y_test))
| ML - Applied Machine Learning Foundation/03.Evaluation - Measuring Success/Evaluation - Measuring Success.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
from PIL import Image
img = Image.open("images/empire.jpg").convert('L') # Convert to grayscale
img = np.array(img) # Convert to numpy.array
n_bins = len(np.unique(img)) - 1
# Distribution of image values
plt.hist(img.flatten(), bins=n_bins-1)
plt.show()
# Function of **histogram_equalization** using `numpy`
def histogram_equaliztion(img, n_bins=None):
""" Histogram equalization of a grayscale image."""
if not n_bins:
n_bins = len(np.unique(img)) - 1
count, bins = np.histogram(img.flatten(), bins = n_bins)
cdf = count.cumsum() # Cumulative Distribution Function
cdf = 255* (cdf - cdf.min())/(cdf.max() - cdf.min()) # Normalize
new_img = np.interp(img.flatten(), bins[:-1], cdf)
new_img = new_img.reshape(img.shape)
new_img = new_img.astype(img.dtype)
return new_img, cdf
# Applying **histogram_equalization**
new_img, cdf = histogram_equaliztion(img)
# Comparing result with original image
# +
fig, (ax1, ax2) = plt.subplots(1, 2)
fig.suptitle('Result of "histogram_equalization"')
ax1.imshow(img, cmap='gray')
ax1.set_title('Original image')
ax2.imshow(new_img, cmap='gray')
ax2.set_title('New image')
plt.show()
# -
# Comparing histogram of distributions
# +
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(10,5))
fig.suptitle('Histogram of distributions')
ax1.hist(img.flatten(), bins=n_bins)
ax1.set_title('Original image distribution')
ax2.hist(new_img.flatten(), bins=n_bins)
ax2.set_title('New image distribution')
plt.show()
# -
| histogram_equalization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Importing Packages
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.model_selection import StratifiedKFold
from sklearn.compose import make_column_transformer
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import GridSearchCV,RandomizedSearchCV
from sklearn.svm import SVC
from sklearn.decomposition import PCA
from xgboost import XGBClassifier
from sklearn.model_selection import cross_val_score
from sklearn.preprocessing import StandardScaler,RobustScaler
from sklearn.preprocessing import OneHotEncoder,LabelEncoder
from sklearn.pipeline import make_pipeline
pd.set_option('display.max_columns', 10)
pd.set_option('display.max_rows', 1000)
# # Loading Datasets
# To preserve the original training data I have made a copy of it.
training = pd.read_csv('../input/summeranalytics2020/train.csv')
train_data = training.copy()
test_data = pd.read_csv('../input/summeranalytics2020/test.csv')
# # Basic EDA
train_data.head()
train_data.info()
# We have 1628 observations and 29 features, out of which 22 are integers and 7 are objects. Some of the integer data type features might also be categorical. We have to predict Attrition which can either be 0 or 1 (1 if the employee left the company).
print(train_data.describe())
# The weird thing to note here is that Behaviour has 0 standard deviation and mean=min=max = 1. This means that this columns has a value of 1 for all the observations, so we will drop it. We will also drop Id because it will have unique values for all the employees.
# +
train_id = train_data.Id
train_data = train_data.drop(['Behaviour','Id'],axis = 1)
test_id = test_data.Id
test_data = test_data.drop(['Behaviour','Id'],axis = 1)
# -
# On exploring a little further I found that the feature PerformanceRating has only two values, 3 or 4 so I have mapped them to 0 and 1 respectively.
train_data['PerformanceRating'] = train_data['PerformanceRating'].apply(lambda x: 0 if x == 3 else 1)
test_data['PerformanceRating'] = test_data['PerformanceRating'].apply(lambda x: 0 if x == 3 else 1)
# Lets check the distribution of out target variable Attrition.
train_data['Attrition'].value_counts().plot(kind = 'bar')
# We have more or less a balanced distribution. Lets check for duplciates
print('Number of duplicates: ',train_data.duplicated().sum())
train_data[train_data.duplicated()]['Attrition'].value_counts().plot(kind = 'bar')
# Alright, so the data has 628 duplicates and all the duplicates correspond to Attrition 1. So that means that data was oversampled to make it balanced. We will drop the duplicates now and check the distribution again.
# +
# drop them
train_unq = train_data.drop_duplicates()
print('New train set: ',train_unq.shape)
X = train_unq.drop('Attrition',axis = 1)
y = train_unq['Attrition']
y.value_counts().plot(kind = 'bar')
plt.show()
# -
# Now our training data has 1000 data points and the target variable is imbalanced. There are many ways to tackle imbalanced data sets like upsampling or downsampling using SMOTE.
# We can also use a cross validation strategy such as stratified k-fold which keeps the distribution of our target variable (here Attrition) similar across the folds. The training and validation data is split using stratified sampling instead of random sampling. The stratas here are the two values of our target variable. If you don't understand what this means then don't worry about it just remember that it is an effective method to tackle imbalanced datasets while we train our model. You can learn more about it in the scikit-learn user guide for cross validation.
# # Pre-Processing, Training and Validation
#
# We will follow the following steps now:
#
# - Drop performance rating (on exploration I found that 85% values were of a single class this might lead to overfitting)
# - One Hot Encode all the 'object' data type features
# - Use standard scaling on all the integer data type features.
# - Use the pre-processed data and split it using Stratified K-Fold.
# - Fit and validate with 3 candidate models: Random Forest, XGBoost and Support Vector Classifier
# +
# Standard Scaling
skf = StratifiedKFold(n_splits = 10,random_state=42,shuffle=True)
categorical = [f for f in training.columns if training[f].dtype == object]
numeric = [f for f in X.columns if f not in categorical+['Id','Attrition','Behaviour','PerformanceRating']]
pre_pipe = make_column_transformer((OneHotEncoder(),categorical),(StandardScaler(),numeric))
# -
# Testing on 3 candidate models: Random Forest, XGBoost, Support Vector Machines
# +
pipe_rf = make_pipeline(pre_pipe,RandomForestClassifier())
pipe_xgb = make_pipeline(pre_pipe,XGBClassifier())
pipe_svc = make_pipeline(pre_pipe,SVC(probability=True))
print('RF: ',np.mean(cross_val_score(X=X,y=y,cv=skf,estimator=pipe_rf,scoring='roc_auc')))
print('XGB: ',np.mean(cross_val_score(X=X,y=y,cv=skf,estimator=pipe_xgb,scoring='roc_auc')))
print('SVC:',np.mean(cross_val_score(X=X,y=y,cv=skf,estimator=pipe_svc,scoring='roc_auc')))
# -
# SVC performs best here - trying out PCA
# +
n = 46
pipe_svc = make_pipeline(pre_pipe,PCA(n_components=n),SVC(probability=True,C = 1,kernel='rbf'))
print('SVC: ',np.mean(cross_val_score(X=X,y=y,cv=skf,estimator=pipe_svc,scoring='roc_auc')))
plt.figure(figsize=(10,8))
pipe_svc.fit(X,y)
plt.plot(range(1,n+1),pipe_svc.named_steps['pca'].explained_variance_ratio_.cumsum())
plt.xticks(range(1,n+1,2))
plt.title('Explained Variance')
plt.grid()
plt.show()
# -
# 34 components are explaining 100% of the variance
n = 34
pre_pipe = make_column_transformer((OneHotEncoder(),categorical),(StandardScaler(),numeric),remainder = 'passthrough')
pipe_svc = make_pipeline(pre_pipe,PCA(n_components=n),SVC(probability=True,C = 1,kernel='rbf'))
print('SVC: ',np.mean(cross_val_score(X=X,y=y,cv=skf,estimator=pipe_svc,scoring='roc_auc')))
# Tuning SVC - using Grid Search
# +
n = 34
pre_pipe = make_column_transformer((OneHotEncoder(),categorical),(StandardScaler(),numeric),remainder = 'passthrough')
pipe_svc = make_pipeline(pre_pipe,PCA(n_components=n),SVC(probability=True,C = 1,kernel = 'rbf'))
param_grid = {
'svc__C':[0.001,0.01,0.1,1,10,100,1000],
'svc__gamma': ['auto','scale'],
'svc__class_weight': ['balanced',None]
}
grid_search = GridSearchCV(pipe_svc,param_grid=param_grid,cv = skf, verbose=2, n_jobs = -1,scoring='roc_auc')
grid_search.fit(X,y)
print('Best score ',grid_search.best_score_)
print('Best parameters ',grid_search.best_params_)
best_svc = grid_search.best_estimator_
# -
# We can tune it further
pipe_svc = make_pipeline(pre_pipe,PCA(n_components=n),SVC(probability=True,C = 1,kernel='rbf',class_weight=None,gamma='auto'))
param_grid={
'svc__C':[0.01,0.03,0.05,0.07,0.1,0.3,0.5,0.7,1]
}
grid_search = GridSearchCV(pipe_svc,param_grid=param_grid,cv = skf, verbose=2, n_jobs = -1,scoring = 'roc_auc')
grid_search.fit(X,y)
print('Best score ',grid_search.best_score_)
print('Best parameters ',grid_search.best_params_)
best_svc = grid_search.best_estimator_ # final model - 0.808 private LB*
# Submission
best_svc.fit(X,y)
prediction = best_svc.predict_proba(test_data)[:,1]
submission = pd.DataFrame(prediction,columns=['Attrition'])
submission['Id'] = test_id
submission = submission[['Id','Attrition']]
submission.to_csv('submissionfile_postcomp.csv',index = None)
# **If you liked this notebook and learnt something new from it do give an upvote. You can also checkout my [blog on medium](http://https://medium.com/@mishraarpan6) where I have given an in-depth explanation about Support Vector Machines and have provided a deeper dive into the code here with links to all the documentaion.**
| Predicting Employee Attrition/hr-attrition-prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # GEOS 518: Applied Hydrologic Science
# # Module 04: Autoregressive Moving Average Models with Hydrologic Time Series
#
# ## By: <NAME>
# ## January 26, 2018
#
# In this notebook...
#
# ## 1. Load the required libraries, load the data, and double-check it
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import statsmodels.tsa.api as sm
# Load the Henry's fork discharge dataset
df = pd.read_pickle('../data/HenrysForkDischarge_WY2001-2015.pkl')
# Create a Water Year column for our pandas data frame. This is a pretty
# simple thing to do, but may not be necessary if you're not dealing with
# discharge data. Here's how it goes:
# 1. Create an empty array that is full of zeros and equal in length to
# the number of days in the record
WY = np.zeros(len(df['Y'].index))
# 2. For those records where the month is less than 10, their associated
# year is the correct water year
WY[df['M'].values < 10] = df['Y'].values[df['M'].values < 10]
# 3. For those records where the month is greater than or equal to 10,
# the correct water year is one more than the current calendar year
WY[df['M'].values >= 10] = df['Y'].values[df['M'].values >= 10] + 1
# 4. Save the water year array as a column in the pandas data frame, as an
# integer
df['WY'] = WY.astype(int)
# Print the first and last 10 records just to make sure we loaded the data okay
qrows = np.concatenate([np.arange(0,10,1),np.arange(-11,-1,1)])
df.iloc[qrows,:]
# -
# ## 2. Split the dataframe into a training and test portion
#
# Often in data-driven modeling, it's a good idea to reserve a portion of your data for building the model and another portion for testing that built model against the developed model. This is generically referred to as "cross validation" and there are many, many ways to do it. The philosophy is that you won't really how robust your statistical model is if you're using all of the data to develop that model. Rather, it's good practice to build your model with some fraction of the data and test how well it does in predicting the other fraction. It's an even better practice to repeat this process many, many (maybe millions of times). This process might look like:
# 1. Randomly select some fraction of the data to be used in building the model - in time series analysis we typically require these fractions to be continuous. We will call this the training or "in-bag" sample.
# 2. Set the other fraction of data aside for testing the developed model. This fraction is known as the test or "out-of-bag" sample.
# 3. Build the model with the training fraction and save the coefficients of the model (e.g., $\mu$, $\phi_1$, etc.)
# 4. Using the indepdent variables contained within the out-of-bag/test dataset, apply the model to make predictions of the dependent variable in the out-of-bag/test sample
# 5. Compare the prediction against the observed values for the out-of-bag/test dataset and compute any error metrics like root mean squared error, bias, $R^2$, etc.
# 6. Save the parameters and repeat steps 1-5 many times
# 7. Characterize the many values of the model parameters from 6 using measures like the average parameter values, the variance in parameter values, histograms of parameter values, etc.
#
# In this case, we will be doing a relatively simple cross-validation exercise in which we split the model into the first 14 years of record, which we will use for model development, and test that model against the final year of data.
# In the following example, we're going to segment the whole dataframe
# into a training dataset (everything that's not Water Year 2015) and
# a test dataset (everything that is Water Year 2015).
df_train = df[df.WY != 2015]
df_test = df[df.WY == 2015]
# ## 3. Examine the Structure of the Partial Autocorrelation Function in the Data
# +
Qt = pd.Series(df_train['Q'].values,df_train['SerDates'].values)
Qpacf = sm.pacf(Qt)
plt.figure(figsize=(14,10))
plt.stem(Qpacf)
plt.ylabel('Partial Autocorrelation Function (PACF) [-]',fontsize=16)
plt.xlabel('Lag Distance [day]',fontsize=16)
ax = plt.gca()
ax.tick_params('both',labelsize=16)
plt.show()
from statsmodels.graphics.tsaplots import plot_pacf
f, axarr = plt.subplots(1, 1, figsize=(14,10))
_ = plot_pacf(Qt,method='ols',lags=40,ax=axarr.axes)
axarr.set_title('Statsmodel plot of PACF',fontsize=16)
axarr.set_ylabel('Partial Autocorrelation Function (PACF) [-]',fontsize=16)
axarr.set_xlabel('Lag Distance [day]',fontsize=16)
axarr = plt.gca()
axarr.tick_params('both',labelsize=16)
# -
# ## 4. Fit ARMA(1,1) and ARMA(2,2) Models to the Data
Q_ARMA_1_1_model = sm.ARMA(Qt-np.mean(Qt),(1,1)).fit()
print(Q_ARMA_1_1_model.params)
Q_ARMA_2_2_model = sm.ARMA(Qt-np.mean(Qt),(2,2)).fit()
print(Q_ARMA_2_2_model.params)
# ## 5. Use the Developed Model to Make Predictions of the Test Data
# +
noise_std = 2.25
Qtrain = df_train['Q'].values
Qtrain = Qtrain.reshape(Qtrain.size,1)
Qtest = df_test['Q'].values
Qtest = Qtest.reshape(Qtest.size,1)
DatesTest = df_test['SerDates'].values
Qttm1 = np.concatenate([Qtrain[-2:-1],Qtest[0:-1]])
Qttm1 = Qttm1.reshape(Qttm1.size,1)
Qttm2 = np.concatenate([Qtrain[-3:-1],Qtest[0:-2]])
Qttm2 = Qttm2.reshape(Qttm2.size,1)
etatm1 = np.random.normal(0.0,noise_std,(len(Qttm1),1))
etatm2 = np.concatenate([np.random.normal(0.0,noise_std,(2,1)),etatm1[0:-2]])
ARMA11_mu = Q_ARMA_1_1_model.params[0]
ARMA11_phi1 = Q_ARMA_1_1_model.params[1]
ARMA11_theta1 = Q_ARMA_1_1_model.params[2]
QhatARMA11 = ARMA11_mu + ARMA11_phi1*Qttm1 - ARMA11_theta1*etatm1
ARMA22_mu = Q_ARMA_2_2_model.params[0]
ARMA22_phi1 = Q_ARMA_2_2_model.params[1]
ARMA22_phi2 = Q_ARMA_2_2_model.params[2]
ARMA22_theta1 = Q_ARMA_2_2_model.params[3]
ARMA22_theta2 = Q_ARMA_2_2_model.params[4]
QhatARMA22 = ARMA22_mu + ARMA22_phi1*Qttm1 + ARMA22_phi2*Qttm2 - ARMA22_theta1*etatm1 - ARMA22_theta2*etatm2
plt.figure(figsize=(14,10))
plt.plot(DatesTest,QhatARMA11,'b-')
plt.plot(DatesTest,QhatARMA22,'g-')
plt.plot(DatesTest,Qtest,'r-')
plt.ylabel('Discharge [m${}^3$/s]',fontsize=16)
plt.xlabel('Date',fontsize=16)
plt.legend(('ARMA(1,1)','ARMA(2,2)','Obs.'),fontsize=16)
ax = plt.gca()
ax.tick_params('both',labelsize=16)
plt.show()
plt.figure(figsize=(14,10))
plt.plot(DatesTest,QhatARMA11,'b-')
plt.plot(DatesTest,QhatARMA22,'g-')
plt.plot(DatesTest,Qtest,'r-')
plt.xlim([DatesTest[250],DatesTest[300]])
plt.ylabel('Discharge [m${}^3$/s]',fontsize=16)
plt.xlabel('Date',fontsize=16)
plt.legend(('ARMA(1,1)','ARMA(2,2)','Obs.'),fontsize=16)
ax = plt.gca()
ax.tick_params('both',labelsize=16)
plt.show()
# -
# ## 6. Compare the Predictions with Observations
# +
# Compute the R^2 values for each prediction
R2ARMA11 = np.corrcoef(QhatARMA11.T,Qtest.T)**2
R2ARMA22 = np.corrcoef(QhatARMA22.T,Qtest.T)**2
muARMA11 = np.mean(QhatARMA11-Qtest)
stdARMA11 = np.std(QhatARMA11-Qtest)
muARMA22 = np.mean(QhatARMA22-Qtest)
stdARMA22 = np.std(QhatARMA22-Qtest)
# Plot the ARMA(1,1) and ARMA(2,2) model results
plt.figure(figsize=(14,10))
plt.subplot(121)
plt.plot(QhatARMA11,Qtest,'bo')
plt.plot([20, 80], [20, 80], 'k-')
plt.title('ARMA(1,1) Model',fontsize=16)
plt.ylabel('Observed Discharge [m${}^3$/s]',fontsize=16)
plt.xlabel('Observed Discharge [m${}^3$/s]',fontsize=16)
ax = plt.gca()
ax.tick_params('both',labelsize=16)
ax.annotate('R${}^2$ = %.3f'%R2ARMA11[0,1], xy=(20,80), fontsize=16)
ax.annotate('Avg. error = %.3f'%muARMA11, xy=(20,78), fontsize=16)
ax.annotate('Std. error = %.3f'%stdARMA11, xy=(20,76), fontsize=16)
plt.subplot(122)
plt.plot(QhatARMA22,Qtest,'bo')
plt.plot([20, 80], [20, 80], 'k-')
plt.title('ARMA(2,2) Model',fontsize=16)
plt.xlabel('Observed Discharge [m${}^3$/s]',fontsize=16)
ax = plt.gca()
ax.tick_params('both',labelsize=16)
ax.annotate('R${}^2$ = %.3f'%R2ARMA22[0,1], xy=(20,80), fontsize=16)
ax.annotate('Avg. error = %.3f'%muARMA22, xy=(20,78), fontsize=16)
ax.annotate('Std. error = %.3f'%stdARMA22, xy=(20,76), fontsize=16)
plt.show()
# -
| Module05_ARMA-Models/.ipynb_checkpoints/ARMA-models-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # MLP: <NAME>
#
# **File:** MLP.ipynb
#
# **Course:** Data Science Foundations: Data Mining in Python
# # IMPORT LIBRARIES
import pandas as pd
import numpy as np
from matplotlib import pyplot as plt
from matplotlib.dates import DateFormatter
from sklearn.neural_network import MLPRegressor
from sklearn.model_selection import TimeSeriesSplit
from sklearn.metrics import mean_squared_error
from sklearn.preprocessing import StandardScaler
import warnings
warnings.filterwarnings('ignore')
# # LOAD AND PREPARE DATA
df = pd.read_csv('data/AirPassengers.csv', parse_dates=['Month'], index_col=['Month'])
# # PLOT DATA
fig, ax = plt.subplots()
plt.xlabel('Year: 1949-1960')
plt.ylabel('Monthly Passengers (1000s)')
plt.title('Monthly Intl Air Passengers')
plt.plot(df, color='black')
ax.xaxis.set_major_formatter(DateFormatter('%Y'))
# # RESHAPE DATA
# - Reshape the data into an `n x (k + 1)` matrix where `n` is the number of samples and `k` is the number of features.
# - Use the last 12 lags of the time series as features (i.e. set `k` equal to 12) in order to capture a full seasonal cycle.
k = 12
# +
Z = []
for i in range(k + 1, df.shape[0] + 1):
Z.append(df.iloc[(i - k - 1): i, 0])
Z = np.array(Z)
# -
# - The first 12 columns of `Z` are the features (lags).
# - The last column of `Z` is the target.
Z.shape
# # SPLIT DATA
# - Use the first 80% of the data for training and the remaining 20% of the data for testing.
split = np.int(0.8 * Z.shape[0])
Z_train, Z_test = Z[:split, :], Z[split:, :]
# # RESCALE DATA
# - Normalize the data by subtracting the mean and dividing by the standard deviation.
scaler = StandardScaler().fit(Z_train)
Z_train = scaler.transform(Z_train)
Z_test = scaler.transform(Z_test)
# - Extract the features (first 12 columns) and the target (last column).
X_train, y_train = Z_train[:, :-1], Z_train[:, -1]
X_test, y_test = Z_test[:, :-1], Z_test[:, -1]
# - Plot the normalized training data.
fig, ax = plt.subplots()
plt.xlabel('Year: 1949-1957')
plt.ylabel('Monthly Passengers (Z-Score)')
plt.title('Monthly Intl Air Passengers (Standardized)')
plt.plot(pd.Series(y_train, index=df.index[k: (k + len(y_train))]), color='black')
ax.xaxis.set_major_formatter(DateFormatter('%Y'))
# - Plot the normalized testing data.
fig, ax = plt.subplots()
plt.xlabel('Year: 1958-1960')
plt.ylabel('Monthly Passengers (Z-Score)')
plt.title('Monthly Intl Air Passengers (Standardized)')
plt.plot(pd.Series(y_test, index=df.index[-len(y_test):]), color='black')
ax.xaxis.set_major_formatter(DateFormatter('%Y'))
# # FIT MLP USING 20% VALIDATION SET
# - Use the last 20% of the training data for validation.
split_ = np.int(0.8 * X_train.shape[0])
X_train_, y_train_ = X_train[:split_, :], y_train[:split_]
X_valid_, y_valid_ = X_train[split_:, :], y_train[split_:]
# - Use the mean squared error (MSE) as validation loss.
def validation_loss(hidden_neurons):
mlp = MLPRegressor(hidden_layer_sizes=(hidden_neurons,), max_iter=500, random_state=1, shuffle=False)
mlp.fit(X_train_, y_train_)
return mean_squared_error(y_valid_, mlp.predict(X_valid_))
# - Calculate the validation loss corresponding to different numbers of hidden nodes.
params = [10, 25, 50, 75]
mse = [validation_loss(p) for p in params]
fig, ax = plt.subplots()
plt.xlabel('Hidden Nodes')
plt.ylabel('Mean Squared Error (MSE)')
plt.title('20% Validation Loss')
plt.plot(params, mse, '-o')
# - Find the number of hidden nodes associated with the smallest validation loss.
params[np.argmin(mse)]
# - Fit an MLP model with 25 hidden nodes to the training data.
mlp = MLPRegressor(hidden_layer_sizes=(25,), max_iter=500, random_state=1, shuffle=False)
mlp.fit(X_train, y_train)
# - Use the fitted MLP model to forecast the test data.
y_pred = mlp.predict(X_test)
# - Transform the data back to the original scale.
y_test_ = scaler.inverse_transform(np.hstack([X_test, y_test.reshape(-1, 1)]))[:, -1]
y_pred_ = scaler.inverse_transform(np.hstack([X_test, y_pred.reshape(-1, 1)]))[:, -1]
# - Plot the data and the forecasts in the original scale.
fig, ax = plt.subplots()
plt.xlabel('Year: 1949-1960')
plt.ylabel('Monthly Passengers (1000s)')
plt.title('Monthly Intl Air Passengers')
plt.plot(df.iloc[:(k + len(y_train) + 1), :], color='black', label='Training Data')
plt.plot(pd.Series(y_test_, index=df.index[-len(y_test):]), color='blue', label='Test Data')
plt.plot(pd.Series(y_pred_, index=df.index[-len(y_test):]), color='red', label='Forecast (25 hidden nodes)')
plt.legend(bbox_to_anchor=(1.05, 1))
ax.xaxis.set_major_formatter(DateFormatter('%Y'))
# # FIT MLP USING FIVE-FOLD CROSS-VALIDATION
# - Set up 5-fold cross validation.
tscv = TimeSeriesSplit(n_splits=5)
# - Use the mean squared error (MSE) as cross-validation loss.
def cross_validation_loss(hidden_neurons):
mse = []
for train_split_, valid_split_ in tscv.split(X_train):
X_train_, y_train_ = X_train[train_split_], y_train[train_split_]
X_valid_, y_valid_ = X_train[valid_split_], y_train[valid_split_]
mlp = MLPRegressor(hidden_layer_sizes=(hidden_neurons,), max_iter=500, random_state=1, shuffle=False)
mlp.fit(X_train_, y_train_)
mse.append(mean_squared_error(y_valid_, mlp.predict(X_valid_)))
return np.mean(mse)
# - Calculate the cross validation loss corresponding to different numbers of hidden nodes.
params = [10, 25, 50, 75]
mse = [cross_validation_loss(p) for p in params]
fig, ax = plt.subplots()
plt.xlabel('Hidden Nodes')
plt.ylabel('Mean Squared Error (MSE)')
plt.title('5-Fold Cross Validation Loss')
plt.plot(params, mse, '-o')
# - Find the number of hidden nodes associated with the smallest cross validation loss.
params[np.argmin(mse)]
# - Fit an MLP model with 50 hidden nodes to the training data.
mlp = MLPRegressor(hidden_layer_sizes=(50,), max_iter=500, random_state=1, shuffle=False)
mlp.fit(X_train, y_train)
# - Use the fitted MLP model to forecast the test data.
y_pred = mlp.predict(X_test)
# - Transform the data back to the original scale.
y_test_ = scaler.inverse_transform(np.hstack([X_test, y_test.reshape(-1, 1)]))[:, -1]
y_pred_ = scaler.inverse_transform(np.hstack([X_test, y_pred.reshape(-1, 1)]))[:, -1]
# - Plot the data and the forecasts in the original scale.
fig, ax = plt.subplots()
plt.xlabel('Year: 1949-1960')
plt.ylabel('Monthly Passengers (1000s)')
plt.title('Monthly Intl Air Passengers')
plt.plot(df.iloc[:(k + len(y_train) + 1), :], color='black', label='Training Data')
plt.plot(pd.Series(y_test_, index=df.index[-len(y_test):]), color='blue', label='Test Data')
plt.plot(pd.Series(y_pred_, index=df.index[-len(y_test):]), color='red', label='Forecast (50 hidden nodes)')
plt.legend(bbox_to_anchor=(1.05, 1))
ax.xaxis.set_major_formatter(DateFormatter('%Y'))
# # CLEAN UP
#
# - If desired, clear the results with Cell > All Output > Clear.
# - Save your work by selecting File > Save and Checkpoint.
# - Shut down the Python kernel and close the file by selecting File > Close and Halt.
| MLP.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Import required libraries
# ### Author: Sameer
# ### Date: May 2019
# +
import numpy as np
import matplotlib.pyplot as plt
from CartPole import CartPole
# from CartPole_GPS import CartPole_GPS
from ilqr.dynamics import constrain
from copy import deepcopy
from EstimateDynamics import local_estimate
from GMM import Estimated_Dynamics_Prior
from sklearn.gaussian_process import GaussianProcessRegressor
from sklearn.gaussian_process.kernels import DotProduct, WhiteKernel
from mujoco_py import load_model_from_path, MjSim, MjViewer
import mujoco_py
import time
# -
# ### Formulate the iLQR problem
'''
1 - dt = time step
2 - N = Number of control points in the trajectory
3 - x0 = Initial state
4 - x_goal = Final state
5 - Q = State cost
6 - R = Control cost
7 - Q_terminal = Cost at the final step
8 - x_dynamics array stores the information regarding system.
x_dynamics[0] = m = mass of the pendulum bob
x_dynamics[1] = M = mass of the cart
x_dynamics[2] = L = length of the massles|s rod
x_dynamics[3] = g = gravity
x_dynamics[4] = d = damping in the system
'''
dt = 0.05
N = 600 # Number of time steps in trajectory.
x_dynamics = np.array([0.1, 1, 1, 9.80665, 0]) # m=1, M=5, L=2, g=9.80665, d=1
x0 = np.array([0.0, 0.0, 3.14, 0.0]) # Initial state
x_goal = np.array([0.0, 0.0, 0.0, 0.0])
# Instantenous state cost.
Q = np.eye(5)
Q[1,1] = 10
Q[2, 2] = 100
Q[3, 3] = 100
Q[4, 4] = 10
# Terminal state cost.
Q_terminal = np.eye(5) * 100
# Q_terminal[2, 2] = 100
# Q_terminal[3, 3] = 100
# Instantaneous control cost.
R = np.array([[1.0]])
# ### iLQR on Cart Pole
cartpole_prob = CartPole(dt, N, x_dynamics, x0, x_goal, Q, R, Q_terminal)
xs, us, K, k = cartpole_prob.run_IterLinQuadReg()
# State matrix split into individual states. For plotting and analysing purposes.
t = np.arange(N + 1) * dt
x = xs[:, 0] # Position
x_dot = xs[:, 1] # Velocity
theta = np.unwrap(cartpole_prob.deaugment_state(xs)[:, 2]) # Theta, makes for smoother plots.
theta_dot = xs[:, 3] # Angular velocity
us_scaled = constrain(us, -1, 1)
# ### Simulate the real system and generate the data
# Cost matrices, initial position and goal position will remain same as the above problem. As it indicates one policy. But still the initial positions and goal positions must be passed explicitly to the function. But you don't need to pass cost matrices (assume penalty on the system is same), this is just used to use to calculate the cost of the trajectory. Correct control action must be passed. Parameter gamma indicates how much of original data you want to keep
#
# Variance of the Gaussian noise will be taken as input from a Unif(0, var_range) uniform distribution. Inputs: x_initial, x_goal, u, n_rollouts, pattern='Normal', pattern_rand=False, var_range=10, gamma=0.2, percent=20
#
# Pattern controls how the control sequence will be modified after applying white Guassian noise (zero mean).
# - Normal: based on the correction/mixing parameter gamma generate control (gamma controls how much noise we want).
# - MissingValue: based on the given percentage, set those many values to zero (it is implicitly it uses "Normal" generated control is used).
# - Shuffle: shuffles the entire "Normal" generated control sequence.
# - TimeDelay: takes the "Normal" generated control and shifts it by 1 index i.e. one unit time delay.
# - Extreme: sets gamma as zeros and generates control based on only noise.
#
# If 'pattern_rand' is 'True' then we don't need to send the explicitly, it will chose one randomly for every rollout (default is 'False'). If you want to chose specific pattern then send it explicitly.
x_rollout, u_rollout, local_policy, cost = cartpole_prob.gen_rollouts(x0, x_goal, us, n_rollouts=10, pattern_rand=True, var_range=10, gamma=0.2, percent=20)
# ### Local system dynamics/model estimate
# loca_estimate: function takes the states (arranged in a special format, [x(t), u(t), x(t+1)]), no. of gaussian mixtures and no.of states.
model = Estimated_Dynamics_Prior(init_sequential=False, eigreg=False, warmstart=True,
min_samples_per_cluster=20, max_clusters=50, max_samples=20, strength=1.0)
model.update_prior(x_rollout, u_rollout)
A, B, C = model.fit(x_rollout, u_rollout)
print(A.shape)
print(B.shape)
print(C.shape)
Model = "mujoco/cartpole.xml"
model_loaded = load_model_from_path(Model)
sim = MjSim(model_loaded)
viewer = mujoco_py.MjViewer(sim)
t = 0
sim.data.qpos[0] = 0.0
sim.data.qpos[1] = 3.14
sim.data.qvel[0] = 0
sim.data.qvel[1] = 0
final = 0
for i in range(600):
start_time = time.time()
state = np.c_[sim.data.qpos[0],sim.data.qvel[0],np.sin(sim.data.qpos[1]),
np.cos(sim.data.qpos[1]),sim.data.qvel[1]].T
control = np.dot(k[i,:],(xs[i].reshape(5,1) - state )) + K[i].T + us[i]
sim.data.ctrl[0] = us[i]
# sim.data.ctrl[0] = control
sim.step()
viewer.render()
if (sim.data.qpos[0] == 1.0 and sim.data.qpos[1] == 0):
print('states reached')
break
print(sim.get_state())
import time
time.sleep(5)
from Simulator import Mujoco_sim
Model = "mujoco/cartpole.xml"
cart_pole_simulator = Mujoco_sim(Model,True)
cart_pole_simulator.load(xs,us,k,K,x0,initial=False)
cart_pole_simulator.runSimulation()
cart_pole_simulator.runSimulation()
np.max(xs)
np.max(us)
import matplotlib.pyplot as plt
# plt.plot(xs[0,:])
plt.plot(us)
import sys
np.set_printoptions(threshold=sys.maxsize)
target_jacp = np.zeros(3 * sim.model.nv)
sim.data.get_site_jacp('tip', jacp=target_jacp)
#This is aparently position jacobian there is something called rotational jacobian get_site_jacr
a = sim.render(width=200, height=200, camera_name='fixed', depth=True)
## a is a tuple if depth is True and a numpy array if depth is False ##
sim.data.get_site_jacp('body', jacp=target_jacp)
| TrajGenerator-V2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Parameter space exploration analysis
import json
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import math
from config import EXPLORATION_SPACE
# %matplotlib inline
# loading results of parameter space exploration
with open('results-space.json', 'r') as fp:
data = json.load(fp)
# check keys of stored values
data.keys()
# nr of evaluations
print('Nr of evaluations performed:', len(data['trials']))
# configuration used for hyperparameter space exploration
config = EXPLORATION_SPACE
# +
# parsing all results
allResults = [res['loss'] for res in data['results']]
# extract results
results = {}
for trial in data['trials']:
res = trial['misc']['vals']
loss = trial['result']['loss']
for k in config.keys():
results.setdefault(k, {})
results[k].setdefault(res[k][0], [])
results[k][res[k][0]].append(loss)
results.keys()
# -
# plot loss value of all evaluations
fig, ax = plt.subplots(figsize=(15, 6))
_ = ax.scatter(np.arange(len(allResults)), allResults, s=1.5)
_ = ax.set_title('Loss on all {} evaluations'.format(len(data['trials'])), fontweight='bold')
_ = ax.set_ylabel('Loss')
fig.savefig('001-overall-loss.pdf', bbox_inches='tight')
# check loss using different window sizes
fig, (ax1, ax2) = plt.subplots(1,2, figsize=(15, 6))
# batch size
orderId = [config['crossoverPB'][i] for i in results['crossoverPB'].keys()]
_ = ax1.boxplot(results['crossoverPB'].values())
_ = ax1.set_xticklabels(np.round(orderId, 2))
_ = ax1.set_ylabel('Loss')
_ = ax1.set_xlabel('Crossover probabilities')
_ = ax1.set_title('Loss with different crossover prob.', fontweight='bold')
# loss with two vs one layers
orderId = [config['nrGenerations'][i] for i in results['nrGenerations'].keys()]
_ = ax2.boxplot(results['nrGenerations'].values())
_ = ax2.set_xticklabels(orderId)
_ = ax2.set_ylabel('Loss')
_ = ax2.set_xlabel('Generation numbers')
_ = ax2.set_title('Loss with different generation numbers', fontweight='bold')
#fig.savefig('002-loss-crossgen.pdf', bbox_inches='tight')
# # Best trial
# Parameter value for the best overall trial
print('Best trial is achieved using:')
for k in data['best'].keys():
print('{}: {}'.format(k, config[k][data['best'][k]]))
print()
print('Best trial raw:')
data['best_trial']
# # Check statistics on 15 best trials
# Check parameter values in the first 15 best results to check if there is some consistency or pattern
# create dataframe with parameter values of the top 15 trials
sortedResults = sorted(allResults)
order = np.argsort(allResults)
topResults = [res['misc']['vals'] for res in data['trials'] if res['result']['loss'] in sortedResults[:15]]
# convert lists to values
for e in topResults:
for k,v in e.items():
if type(v) == type(list()):
e[k] = v[0]
df = pd.DataFrame(topResults)
df['points'] = [data['stepPoints'][i] for i in order][:15]
df.head()
# isolate each parameter
def convertValue(row):
row['Real value'] = config[c][row.name]
return row
for c in df.columns:
print('## Top 15 values for parameter: {}'.format(c))
dfs = df[c].value_counts().to_frame()
dfs.columns = ['Counts']
dfs.index.name = 'Raw value'
if c != 'points':
dfs = dfs.apply(convertValue, axis=1)
print(dfs)
print()
# Lets isolate each point distribution to see the mean value of each parameter. The goal would be to find a single value for each parameter which works well for all point distributions. This assumption might be unreasonable and can be investidated through appropriate plots.
# +
# function to convert raw values from hyperopt to real values from config file
def convert(row):
for col in [col for col in row.index if col not in ['point', 'fitness']]:
row[col] = config[col][int(row[col])]
return row
# number of best results to consider to compute the statistics
topPoints = 15
uniquePoints = sorted(set(data['stepPoints']))
dfData = []
for point in uniquePoints:
# find all points matching
pointIds = [i for i, el in enumerate(data['stepPoints']) if el==point]
pointResults = [allResults[i] for i in pointIds]
pointResultsOrdered = sorted(pointResults)
order = np.argsort(pointResults)
pointTrials = [data['trials'][i] for i in pointIds]
# for each point distribution, extract the topPoints in terms of loss
pointTrials = [pointTrials[i] for i in order][:topPoints]
topResults = [res['misc']['vals'] for res in pointTrials]
# convert lists to values
for i,e in enumerate(topResults):
singleResult = {}
for k,v in e.items():
if type(v) == type(list()):
singleResult[k] = v[0]
else:
singleResult[k] = v
singleResult['point']=point
singleResult['fitness']=pointResultsOrdered[i]
dfData.append(singleResult)
df = pd.DataFrame(dfData, dtype='float')
df = df.apply(convert, axis=1)
df.head()
# -
# group above dataframe by point in order to show specific statistics
dfGrouped = df.groupby(['point']).agg(['mean','std'])
dfGrouped
# Helper function to return plot row and column
def currentRowCol(c,cols,rows):
if rows == 1 or cols == 1:
return c,0
row = math.floor(c/cols)
col = c-row*cols
return row,col
cols = 2
rows = 3
fig, ax = plt.subplots(rows,cols,figsize=(15,12))
fig.suptitle('LOSSES ON ALL PARAMETERS AND POINT DISTRIBUTIONS', fontweight='bold')
for i,groupName in enumerate(set(dfGrouped.columns.get_level_values(0))):
if groupName == 'fitness':
continue
r,c = currentRowCol(i,cols,rows)
group = dfGrouped[groupName]
x = group.index
y = group['mean']
e = group['std']/2
ax[r,c].errorbar(x,y,e, linestyle='None', marker='o', color='k', capsize=5, ecolor='r', elinewidth=0.5, label='Mean/stdev')
ax[r,c].axhline(y=np.mean(y), linewidth=1, color='k', linestyle='--', label='Global mean')
ax[r,c].axhline(y=config[groupName][0], linewidth=1, color='r', linestyle='--', label='Limits')
ax[r,c].axhline(y=config[groupName][-1], linewidth=1, color='r', linestyle='--')
ax[r,c].legend(loc='upper right')
ax[r,c].set_title('Top 15 {} values statistics'.format(groupName))
ax[r,c].set_xlabel('Points')
plt.tight_layout()
plt.subplots_adjust(top=0.92)
fig.savefig('003-parameters-points-loss.pdf', bbox_inches='tight')
# As it was expected, the optimal values for each parameter varies significantly over different point distributions.
# The mean values shown above in the black dotted line, which will be used
# as optimal parameters for the final genetic algorithm
np.round(dfGrouped.mean(),2)
| 005-parameter-exploration-analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Essential Shortcuts
# ## Intro to Shortcuts
# Keyboard shortcuts are a meta-skill that will save you time across your entire life as a coder. Jupyter keyboard shortcuts are essential because A. you will use Jupyter often and B. the alternative of using the mouse + menu, is awful.
#
# Unfortunately, most software comes with a *comprehensive* list of shortcuts rather than a *practical* one. This is generally true of documentation as well and, while this can be great for the already experienced user of your library or software, it sucks for the new user, even if they are an experienced coder.
#
# What the new user often needs is a list of functionality ordered from most commonly used to least commonly used, so I did just that with a custom Jupyter extension, something you'll learn in Chapter 3.
# ### The Learn Shortcuts Extension
# To see the full, unordered list of Jupyter shortcuts, press **H**
# To see a structured list of only the most common shortcuts, press **Shift+H** (our custom extension)
# ## Command Mode Shortcuts
# Let's start with the most important shortcuts in <span style="color:blue"><strong>Command Mode</strong></span> (meant for changing the whole cell). I'll list them here, give you some space to practice them yourself, and then I'll return to go over the ones that can be a bit tricky.
#
# * Creating and deleting cells
# * **A** - insert new cell above
# * **B** - insert new cell below
# * **DD** - delete selected cells
# * **Z** - undo cell deletion
# * Cutting copying and pasting
# * **C** - copy cell
# * **V** - paste cell below
# * **Shift-V** - paste cell above
# * **X** - cut cell
# * Change cell types
# * **Y** - change cell to `code` type
# * **M** - change cell to *markdown* type
# * **R** - change cell to raw text (rarely useful)
# * Navigation
# * **Up arrow** - Select cell above
# * **Down arrow** - Select cell below
# * **Shift+Up arrow** - Select multiple cells above
# * **Shift+Down arrow** - Select multiple cells below
# * **Space** - Scroll down
# * **Shift-Space** -Scroll up
# * Other functions
# * **S** - save notebook
# * **H** - view keyboard shortcuts
# * **Shift-H** - view basic shortcuts
# ### Practice
# I strongly encourage you to try out each one of the shortcuts in the list above. I've included below a combination of empty, code, and markdown cells for you to play with. Remember you can press **Shift-H** to see useful shortcuts so you don't have to scroll up. Stick to the <span style="color:blue"><strong>Command Mode</strong></span> side for now.
# This is a code cell
for i in range(10):
print("I will practice keyboard shortcuts")
# This is a markdown cell with an image
#
# 
# **This is the end of practice**
# ### A few things to watch out for
# #### Z - Undo Delete
# This is a really important one to know since you'll occasionally delete a cell on accident. People often hit Ctrl-Z out of habit and wonder why it doesn't work. It's because undo is simply **Z** (in command mode). Like normal "undo" functionality, Jupyter keeps a stack of all deleted cells, so pressing **Z** multiple times will bring back multiple recently deleted cells.
#
# Practice using **DD** to delete some cells, and **Z** to bring them back.
# #### C, V, and X for Copying and Pasting
# These are easy enough when used in one notebook, but often confuse people when it comes to copying cells from one notebook and pasting them in another. For that you'll need **Ctrl-C** and **Ctrl-V**.
#
# One other quirk of this is **Ctrl-C** will actually paste above, not below.
#
# Finally, if you're copying multiple cells at once, make sure you're in command mode when you paste them, or they'll all get combined into one cell.
# ## Edit Mode Shortcuts
# Okay now that you're a <span style="color:blue"><strong>Command Mode</strong></span> Pro. Let's jump into shortcuts for <span style="color:green"><strong>Edit Mode</strong></span> (meant for changing the inside of a cell)
#
# * Same as Most Applications
# * **Ctrl-C** - Copy
# * **Ctrl-V** - Paste
# * **Ctrl-X** - Cut
# * **Ctrl-Z** - Undo
# * **Ctrl-Y** - Redo
# * **Ctrl-S** - Save and Checkpoint
# * Useful For Code
# * **Tab** - Autocomplete Code
# * **Shift-Tab** - Show Function Docstring (simple)
# * **Shift-Tab(x2)** - Show Function Docstring (complete)
# * **Shift-Tab(x4)** - Show Function Docstring (in pager)
# * **Ctrl-\[** - Indent Code
# * **Ctrl-\]** - Dedent Code
# * **Ctrl-/** - Comment/Uncomment Selection
# * Navigation
# * **Home** - Go to Line Start
# * **End** - Go to Line End
# * **Ctrl-Left** - Go One Word Left
# * **Ctrl-Right** - Go One Word Right
# ### Edit Mode Practice
# You know the drill. Use this space as a sandbox to try out those slick <span style="color:blue"><strong>Edit Mode</strong></span> shortcuts.
# This is a code cell
total = 0
for i in range(5):
for j in range(5):
print(f"{i} + {j} is {i+j}")
total += i+j
print("\nThe total of all the sums is", total)
# **This is the end of practice**
# ### A few things to watch out for
# #### Ctrl-Z and Ctrl-Y: Each cell has it's own undo/redo memory
# Ctrl-Z will only undo the most recent action in the current cell you're in. If you make 20 changes in Cell A, and 10 in Cell B, and then return to Cell A (in edit mode) and press **Ctrl-Z**, it will start undoing those first 20 changes you made, not the more recent ones. This is because each cell maintains it's own history of actions for undo and redo.
#
# This might seem weird, but think about having multiple cells being like having multiple documents open in Microsoft Word. If you were trying to undo changes in one document, and it started undoing more recent changes in another document, wouldn't that be ineffective and hard to deal with? For the same reason, Jupyter treats it's cells as independent entities with their own memory/history.
# #### Tab only indents code cells if you are at the start of the line
# If you're used to using a more traditional code editor, you expect tab to indent and shift-tab to dedent, even if you're in the middle of a line. Because Jupyter is designed for interactivity, it reserves tab for autocompleting code.
#
# Pressing tab at the start of the line will still indent, but if you press tab anywhere after the *first non-whitespace character*, Jupyter will
# - Autocomplete the code if there's only one possible completion
# - Show you a list of options if there are multiple
#
# Here's an example:
# 
#
# To select one of the options from the popup menu, use the arrow keys and press enter, pressing tab won't work.
# #### Ctrl + \[ for indent
# Pressing ctrl+\[ will let you indent the entire line anywhere in a code cell. It also works if you select multiple lines and indent them all at once. Select multiple lines the same way you would in any text editor, holding shift and using the arrow keys.
# #### Shift-Tab is incredibly useful
# You can use shift-tab on any line that calls a function. It doesn't matter if the cursor is at the start/end/middle of the line. Pressing shift-tab will show you incredibly useful information, the signature (what parameters the function accepts) and the docstring (information about the functions use).
#
# When I say Shift-Tab(x2) and Shift-Tab(x4) I mean while holding down shift, press tab that many times. What happened to Shift-Tab(x3) you might ask? Here's the full breakdown:
#
# - Shift-Tab(x1) - Show signature and first line of the docstring
# - Shift-Tab(x2) - Show signature and complete docstring
# - Shift-Tab(x3) - Show signature and complete docstring and leave it up 10 seconds longer
# - Shift-Tab(x4)- Show signature and first line of the docstring in Jupyter's pager.
#
# The last option is great when you have a function or method with a really detailed docstring describing each parameter and its options as is very common in data science libraries.
print
# #### Navigation: Better options than scrolling
# The navigation shortcuts might not seem that essential, and I'll level with you, they almost didn't make the cut. I chose to include them as essential because you will have to fix typos/mistakes many times every single day and the time will add up.
#
# - Home/End will beat scrolling or using a mouse.
# - You can use ctrl+left/right repeatedly for a fast scroll
# - Use ctrl+shift+left/right to quickly select a group of words
| tutorials/1.1 Essential Shortcuts.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Missing Values
# +
# *-* coding: utf-8 -*-
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
datacsv = pd.read_csv("missingvalue.csv")
#size = datacsv['boy']
datacsv
# +
#Missing Values
from sklearn.impute import SimpleImputer
imputer = SimpleImputer(missing_values=np.nan, strategy='mean')
Age = datacsv.iloc[:,1:4].values
imputer = imputer.fit(Age[:,1:4])
Age[:,1:4] = imputer.transform(Age[:,1:4])
# -
# # Categorical Data
country = datacsv.iloc[:, 0:1].values
from sklearn import preprocessing
le = preprocessing.LabelEncoder()
# ## In this here created categorical data to country values
country[:,0] = le.fit_transform(datacsv.iloc[:,0])
ohe = preprocessing.OneHotEncoder()
country = ohe.fit_transform(country).toarray()
resultCountry = pd.DataFrame(data=country, index=range(22), columns=['fr','tr','us'])
# ## Create Data Frame Values by ages and Gender
ResultPerson = pd.DataFrame(data=Age, index=range(22), columns = ['boy', 'kilo', 'yas'])
Gender = datacsv.iloc[:,-1].values
ResultGender = pd.DataFrame(data=Gender, index=range(22), columns = ['cinsiyet'])
# ## Combine Data Frames (Country, Person)
ResultC1 = pd.concat([resultCountry,ResultPerson], axis=1)
ResultC1
# ## Combine All DataFrames(Country, Person, Gender)
#
ResultAll = pd.concat([ResultC1, ResultGender], axis=1)
ResultAll
# # Test and Train with dependent(Country) and undependant(Gender)
# +
from sklearn.model_selection import train_test_split
x_train, x_test, y_train, y_test = train_test_split(ResultC1, ResultAll, test_size=0.33, random_state=0)
# -
# # Feature Scaling
# +
from sklearn.preprocessing import StandardScaler
# create standard scaler object
sc = StandardScaler()
# return x_train values on fit_transform function for feature scaling
X_train = sc.fit_transform(x_train)
X_test = sc.fit_transform(x_test)
# -
print(X_train)
print('')
print(X_test)
| LinearRegression/pre-missingvalue.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="../reports/figures/marketwatch.png" alt="Drawing" width="200">
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from selenium import webdriver
from selenium.webdriver.common.keys import Keys
from selenium.webdriver.common.by import By
from selenium.webdriver.support.ui import WebDriverWait
from selenium.webdriver.support import expected_conditions as EC
from selenium.common.exceptions import TimeoutException
# <div class="alert alert-block alert-info">
# <b>Market Watch:</b>is a great websight to pull financial statements and economic reports. The goal is to use the information here
# as a 5 year trend to search for any stock on the webpage to pull in there well organized tables.
# </div>
# +
def get_financial_report(ticker):
driver = webdriver.Firefox(executable_path= "geckodriver.exe")
driver.maximize_window()
driver.get('https://www.marketwatch.com/investing/stock/'+ticker+'/financials')
timeout = 20
# ''''
# Find an ID on the page and wait before executing anything until found. After searching multiple times,
# You will be prompted to subscribe so there was a need to close out the subscribe overlay.
# ''''
try:
WebDriverWait(driver, timeout).until(EC.visibility_of_element_located((By.ID, "cx-scrim-wrapper")))
except TimeoutException:
driver.quit()
subscribe = driver.find_element_by_xpath('/html/body/footer/div[2]/div/div/div[1]')
subscribe.click()
# ''''
# For the income statement, balance sheet, and cashflow; you can scrape the tables and tranpose them to
# the years going 5 years back, for any calculations that might need to be done in the future. We kept all information
# including empty columns just in case companies had more detailed reports and information. However for the
# most part all have the same format on market watch making it easily updateable and great for a search bar.
# ''''
income_table = driver.find_element_by_class_name("overflow--table").get_attribute('innerHTML')
income = pd.read_html(income_table)
income_df = income[0]
income_df.drop(columns=['5-year trend'], axis=1, inplace=True)
income_df2 = income_df.T
income_new_col = income_df2.iloc[0,:].values
income_df2.columns= income_new_col
income_df2.drop(index='Item Item', inplace=True)
income_df2.rename(columns={'Sales/Revenue Sales/Revenue': 'Sales/Revenue' ,
'Sales Growth Sales Growth': 'Sales Growth',
'Cost of Goods Sold (COGS) incl. D&A Cost of Goods Sold (COGS) incl. D&A': 'Cost of Goods Sold (COGS) incl. D&A',
'COGS Growth COGS Growth': 'COGS Growth', 'COGS excluding D&A COGS excluding D&A': 'COGS excluding D&A',
'Depreciation & Amortization Expense Depreciation & Amortization Expense': 'Depreciation & Amortization Expense',
'Depreciation Depreciation': 'Depreciation',
'Amortization of Intangibles Amortization of Intangibles': 'Amortization of Intangibles',
'Gross Income Gross Income': 'Gross Income',
'Gross Income Growth Gross Income Growth': 'Gross Income Growth',
'Gross Profit Margin Gross Profit Margin': 'Gross Profit Margin',
'SG&A Expense SG&A Expense': 'SG&A Expense', 'SGA Growth SGA Growth': 'SGA Growth',
'Research & Development Research & Development': 'Research & Development',
'Other SG&A Other SG&A': 'Other SG&A',
'Other Operating Expense Other Operating Expense': 'Other Operating Expense',
'Unusual Expense Unusual Expense': 'Unusual Expense',
'EBIT after Unusual Expense EBIT after Unusual Expense': 'EBIT after Unusual Expense',
'Non Operating Income/Expense Non Operating Income/Expense': 'Non Operating Income/Expense',
'Non-Operating Interest Income Non-Operating Interest Income': 'Non-Operating Interest Income',
'Equity in Affiliates (Pretax) Equity in Affiliates (Pretax)': 'Equity in Affiliates (Pretax)',
'Interest Expense Interest Expense': 'Interest Expense',
'Interest Expense Growth Interest Expense Growth': 'Interest Expense Growth',
'Gross Interest Expense Gross Interest Expense': 'Gross Interest Expense',
'Interest Capitalized Interest Capitalized': 'Interest Capitalized',
'Pretax Income Pretax Income': 'Pretax Income',
'Pretax Income Growth Pretax Income Growth': 'Pretax Income Growth',
'Pretax Margin Pretax Margin': 'Pretax Margin', 'Income Tax Income Tax': 'Income Tax',
'Income Tax - Current Domestic Income Tax - Current Domestic': 'Income Tax - Current Domestic',
'Income Tax - Current Foreign Income Tax - Current Foreign': 'Income Tax - Current Foreign',
'Income Tax - Deferred Domestic Income Tax - Deferred Domestic': 'Income Tax - Deferred Domestic',
'Income Tax - Deferred Foreign Income Tax - Deferred Foreign': 'Income Tax - Deferred Foreign',
'Income Tax Credits Income Tax Credits': 'Income Tax Credits',
'Equity in Affiliates Equity in Affiliates': 'Equity in Affiliates',
'Other After Tax Income (Expense) Other After Tax Income (Expense)': 'Other After Tax Income (Expense)',
'Consolidated Net Income Consolidated Net Income': 'Consolidated Net Income',
'Minority Interest Expense Minority Interest Expense': 'Minority Interest Expense',
'Net Income Net Income': 'Net Income', 'Net Income Growth Net Income Growth': 'Net Income Growth',
'Net Margin Growth Net Margin Growth': 'Net Margin Growth',
'Extraordinaries & Discontinued Operations Extraordinaries & Discontinued Operations': 'Extraordinaries & Discontinued Operations',
'Extra Items & Gain/Loss Sale Of Assets Extra Items & Gain/Loss Sale Of Assets': 'Extra Items & Gain/Loss Sale Of Assets',
'Cumulative Effect - Accounting Chg Cumulative Effect - Accounting Chg': 'Cumulative Effect - Accounting Chg',
'Discontinued Operations Discontinued Operations': 'Discontinued Operations',
'Net Income After Extraordinaries Net Income After Extraordinaries': 'Net Income After Extraordinaries',
'Preferred Dividends Preferred Dividends': 'Preferred Dividends',
'Net Income Available to Common Net Income Available to Common': 'Net Income Available to Common',
'EPS (Basic) EPS (Basic)': 'EPS (Basic)', 'EPS (Basic) Growth EPS (Basic) Growth': 'EPS (Basic) Growth',
'Basic Shares Outstanding Basic Shares Outstanding': 'Basic Shares Outstanding',
'EPS (Diluted) EPS (Diluted)': 'EPS (Diluted)',
'EPS (Diluted) Growth EPS (Diluted) Growth': 'EPS (Diluted) Growth',
'Diluted Shares Outstanding Diluted Shares Outstanding': 'Diluted Shares Outstanding',
'EBITDA EBITDA': 'EBITDA', 'EBITDA Growth EBITDA Growth': 'EBITDA Growth',
'EBITDA Margin EBITDA Margin': 'EBITDA Margin'}, inplace=True)
income_df2.to_csv('../data/interim/income_statement.csv')
driver.get('https://www.marketwatch.com/investing/stock/'+ticker+'/financials/balance-sheet')
balance_sheet_table = [table.get_attribute('innerHTML') for table in driver.find_elements_by_class_name("overflow--table")]
balance_sheet = pd.read_html(balance_sheet_table[0])
balance_sheet1 = pd.read_html(balance_sheet_table[1])
balance_sheet_df = pd.concat([balance_sheet[0], balance_sheet1[0]])
balance_sheet_df.drop(columns=['5-year trend'], axis=1, inplace=True)
balance_sheet_df2 = balance_sheet_df.T
balance_sheet_col = balance_sheet_df2.iloc[0,:].values
balance_sheet_df2.columns= balance_sheet_col
balance_sheet_df2.drop(index='Item Item', inplace=True)
balance_sheet_df2.rename(columns={'Cash & Short Term Investments Cash & Short Term Investments': 'Cash & Short Term Investments',
'Cash & Short Term Investments Growth Cash & Short Term Investments Growth': 'Short Term Investments Growth',
'Cash Only Cash Only': 'Cash Only',
'Short-Term Investments Short-Term Investments': 'Short-Term Investments',
'Cash & ST Investments / Total Assets Cash & ST Investments / Total Assets': 'Cash & ST Investments / Total Assets',
'Total Accounts Receivable Total Accounts Receivable': 'Total Accounts Receivable',
'Total Accounts Receivable Growth Total Accounts Receivable Growth': 'Total Accounts Receivable Growth',
'Accounts Receivables, Net Accounts Receivables, Net': 'Accounts Receivables, Net',
'Accounts Receivables, Gross Accounts Receivables, Gross': 'Accounts Receivables, Gross',
'Bad Debt/Doubtful Accounts Bad Debt/Doubtful Accounts': 'Bad Debt/Doubtful Accounts',
'Other Receivable Other Receivable': 'Other Receivable',
'Accounts Receivable Turnover Accounts Receivable Turnover': 'Accounts Receivable Turnover',
'Inventories Inventories': 'Inventories', 'Finished Goods Finished Goods': 'Finished Goods',
'Work in Progress Work in Progress': 'Work in Progress', 'Raw Materials Raw Materials': 'Raw Materials',
'Progress Payments & Other Progress Payments & Other': 'Progress Payments & Other',
'Other Current Assets Other Current Assets': 'Other Current Assets',
'Miscellaneous Current Assets Miscellaneous Current Assets': 'Miscellaneous Current Assets',
'Total Current Assets Total Current Assets': 'Total Current Assets',
'Net Property, Plant & Equipment Net Property, Plant & Equipment': 'Net Property, Plant & Equipment',
'Property, Plant & Equipment - Gross Property, Plant & Equipment - Gross': 'Property, Plant & Equipment - Gross',
'Buildings Buildings': 'Buildings', 'Land & Improvements Land & Improvements': 'Land & Improvements',
'Computer Software and Equipment Computer Software and Equipment': 'Computer Software and Equipment',
'Other Property, Plant & Equipment Other Property, Plant & Equipment': 'Other Property, Plant & Equipment',
'Accumulated Depreciation Accumulated Depreciation': 'Accumulated Depreciation',
'Total Investments and Advances Total Investments and Advances': 'Total Investments and Advances',
'Other Long-Term Investments Other Long-Term Investments': 'Other Long-Term Investments',
'Long-Term Note Receivables Long-Term Note Receivables': 'Long-Term Note Receivables',
'Intangible Assets Intangible Assets': 'Intangible Assets', 'Net Goodwill Net Goodwill': 'Net Goodwill',
'Net Other Intangibles Net Other Intangibles': 'Net Other Intangibles',
'Other Assets Other Assets': 'Other Assets', 'Total Assets Total Assets': 'Total Assets',
'Total Assets Growth Total Assets Growth': 'Total Assets Growth','ST Debt & Current Portion LT Debt ST Debt & Current Portion LT Debt':'ST Debt & Current Portion LT Debt',
'Short Term Debt Short Term Debt': 'Short Term Debt',
'Current Portion of Long Term Debt Current Portion of Long Term Debt': 'Current Portion of Long Term Debt',
'Accounts Payable Accounts Payable': 'Accounts Payable',
'Accounts Payable Growth Accounts Payable Growth': 'Accounts Payable Growth',
'Income Tax Payable Income Tax Payable': 'Income Tax Payable',
'Other Current Liabilities Other Current Liabilities': 'Other Current Liabilities',
'Dividends Payable Dividends Payable': 'Dividends Payable',
'Accrued Payroll Accrued Payroll': 'Accrued Payroll',
'Miscellaneous Current Liabilities Miscellaneous Current Liabilities': 'Miscellaneous Current Liabilities',
'Total Current Liabilities Total Current Liabilities': 'Total Current Liabilities',
'Long-Term Debt Long-Term Debt': 'Long-Term Debt',
'Long-Term Debt excl. Capitalized Leases Long-Term Debt excl. Capitalized Leases': 'Long-Term Debt excl. Capitalized Leases',
'Non-Convertible Debt Non-Convertible Debt': 'Non-Convertible Debt',
'Convertible Debt Convertible Debt': 'Convertible Debt',
'Capitalized Lease Obligations Capitalized Lease Obligations': 'Capitalized Lease Obligations',
'Provision for Risks & Charges Provision for Risks & Charges': 'Provision for Risks & Charges',
'Deferred Taxes Deferred Taxes': 'Deferred Taxes',
'Deferred Taxes - Credits Deferred Taxes - Credits': 'Deferred Taxes - Credits',
'Deferred Taxes - Debit Deferred Taxes - Debit': 'Deferred Taxes - Debit',
'Other Liabilities Other Liabilities': 'Other Liabilities',
'Other Liabilities (excl. Deferred Income) Other Liabilities (excl. Deferred Income)': 'Other Liabilities (excl. Deferred Income)',
'Deferred Income Deferred Income': 'Deferred Income',
'Total Liabilities Total Liabilities': 'Total Liabilities',
'Non-Equity Reserves Non-Equity Reserves': 'Non-Equity Reserves',
'Total Liabilities / Total Assets Total Liabilities / Total Assets': 'Total Liabilities / Total Assets',
'Preferred Stock (Carrying Value) Preferred Stock (Carrying Value)': 'Preferred Stock (Carrying Value)',
'Redeemable Preferred Stock Redeemable Preferred Stock': 'Redeemable Preferred Stock',
'Non-Redeemable Preferred Stock Non-Redeemable Preferred Stock': 'Non-Redeemable Preferred Stock',
'Common Equity (Total) Common Equity (Total)': 'Common Equity (Total)',
'Common Equity / Total Assets Common Equity / Total Assets': 'Common Equity / Total Assets',
'Common Stock Par/Carry Value Common Stock Par/Carry Value': 'Common Stock Par/Carry Value',
'Retained Earnings Retained Earnings': 'Retained Earnings',
'ESOP Debt Guarantee ESOP Debt Guarantee': 'ESOP Debt Guarantee',
'Cumulative Translation Adjustment/Unrealized For. Exch. Gain Cumulative Translation Adjustment/Unrealized For. Exch. Gain': 'Cumulative Translation Adjustment/Unrealized For. Exch. Gain',
'Unrealized Gain/Loss Marketable Securities Unrealized Gain/Loss Marketable Securities': 'Unrealized Gain/Loss Marketable Securities',
'Revaluation Reserves Revaluation Reserves': 'Revaluation Reserves',
'Treasury Stock Treasury Stock': 'Treasury Stock',
"Total Shareholders' Equity Total Shareholders' Equity": 'Total Shareholders Equity',
"Total Shareholders' Equity / Total Assets Total Shareholders' Equity / Total Assets": 'Total Shareholders Equity / Total Assets',
'Accumulated Minority Interest Accumulated Minority Interest': 'Accumulated Minority Interest',
'Total Equity Total Equity': 'Total Equity',
"Liabilities & Shareholders' Equity Liabilities & Shareholders' Equity": 'Liabilities & Shareholders Equity'}, inplace=True)
balance_sheet_df2.to_csv('../data/interim/balance_sheet.csv')
# timeout = 20
# # Find an ID on the page and wait before executing anything until found:
# try:
# WebDriverWait(driver, timeout).until(EC.visibility_of_element_located((By.ID, "cx-scrim-wrapper")))
# except TimeoutException:
# driver.quit()
# subscribe = driver.find_element_by_xpath('/html/body/footer/div[2]/div/div/div[1]')
# subscribe.click()
driver.get('https://www.marketwatch.com/investing/stock/'+ticker+'/financials/cash-flow')
cashflow_table = [table.get_attribute('innerHTML') for table in driver.find_elements_by_class_name("overflow--table")]
cashflow = pd.read_html(cashflow_table[0])
cashflow1 = pd.read_html(cashflow_table[1])
cashflow_df = pd.concat([cashflow[0], cashflow1[0]])
cashflow_df.drop(columns=['5-year trend'], axis=1, inplace=True)
cashflow_df2 = cashflow_df.T
cashflow_new_col = cashflow_df2.iloc[0,:].values
cashflow_df2.columns= cashflow_new_col
cashflow_df2.drop(index='Item Item', inplace=True)
cashflow_df2.rename(columns={'Net Income before Extraordinaries Net Income before Extraordinaries': 'Net Income before Extraordinaries',
'Net Income Growth Net Income Growth': 'Net Income Growth',
'Depreciation, Depletion & Amortization Depreciation, Depletion & Amortization': 'Depletion & Amortization',
'Depreciation and Depletion Depreciation and Depletion': 'Depreciation and Depletion',
'Amortization of Intangible Assets Amortization of Intangible Assets': 'Amortization of Intangible Assets',
'Deferred Taxes & Investment Tax Credit Deferred Taxes & Investment Tax Credit': 'Deferred Taxes & Investment Tax Credit' ,
'Deferred Taxes Deferred Taxes': 'Deferred Taxes',
'Investment Tax Credit Investment Tax Credit': 'Investment Tax Credit',
'Other Funds Other Funds': 'Other Funds' ,
'Funds from Operations Funds from Operations': 'Funds from Operations',
'Extraordinaries Extraordinaries': 'Extraordinaries',
'Changes in Working Capital Changes in Working Capital': 'Changes in Working Capital',
'Receivables Receivables': 'Receivables', 'Accounts Payable Accounts Payable': 'Accounts Payable',
'Other Assets/Liabilities Other Assets/Liabilities': 'Other Assets/Liabilities',
'Net Operating Cash Flow Net Operating Cash Flow': 'Net Operating Cash Flow',
'Net Operating Cash Flow Growth Net Operating Cash Flow Growth': 'Net Operating Cash Flow Growth',
'Net Operating Cash Flow / Sales Net Operating Cash Flow / Sales': 'Net Operating Cash Flow / Sales',
'Capital Expenditures Capital Expenditures': 'Capital Expenditures' ,
'Capital Expenditures Growth Capital Expenditures Growth': 'Capital Expenditures Growth',
'Capital Expenditures / Sales Capital Expenditures / Sales': 'Capital Expenditures / Sales',
'Capital Expenditures (Fixed Assets) Capital Expenditures (Fixed Assets)': 'Capital Expenditures (Fixed Assets)',
'Capital Expenditures (Other Assets) Capital Expenditures (Other Assets)': 'Capital Expenditures (Other Assets)',
'Net Assets from Acquisitions Net Assets from Acquisitions': 'Net Assets from Acquisitions',
'Sale of Fixed Assets & Businesses Sale of Fixed Assets & Businesses': 'Sale of Fixed Assets & Businesses',
'Purchase/Sale of Investments Purchase/Sale of Investments': 'Purchase/Sale of Investments',
'Purchase of Investments Purchase of Investments': 'Purchase of Investments',
'Sale/Maturity of Investments Sale/Maturity of Investments': 'Sale/Maturity of Investments',
'Other Uses Other Uses': 'Other Uses', 'Other Sources Other Sources': 'Other Sources',
'Net Investing Cash Flow Net Investing Cash Flow': 'Net Investing Cash Flow',
'Net Investing Cash Flow Growth Net Investing Cash Flow Growth': 'Net Investing Cash Flow Growth',
'Net Investing Cash Flow / Sales Net Investing Cash Flow / Sales': 'Net Investing Cash Flow / Sales'}, inplace=True)
cashflow_df2.to_csv('../data/interim/cashflows.csv')
driver.quit()
return income_df2, balance_sheet_df2, cashflow_df2
# -
income_statement, balance_sheet, cashflow = get_financial_report('TSLA')
# <div class="alert alert-block alert-info">
# <b>Income Statement</b> is one of a company's core financial statements that shows their profit and lossProfit and Loss Statement (P&L)A profit and loss statement (P&L), or income statement or statement of operations, is a financial report that provides a summary of a over a period of time.<br><br>
# <b>Takeaway:</b>
# <dl>
# <dd><li>Net Income = (Total Revenue + Gains) – (Total Expenses + Losses)</li></dd>
# <dd><li>Total revenue is the sum of both operating and non-operating revenues while total expenses include those incurred by primary and secondary activities.</li></dd>
# <dd><li>Revenues are not receipts. Revenue is earned and reported on the income statement. Receipts (cash received or paid out) are not.</li></dd>
# <dd><li>An income statement provides valuable insights into a company’s operations, the efficiency of its management, under-performing sectors and its performance relative to industry peers.</li></dd>
# </dl>
# </div>
# <div class="alert alert-block alert-info">
# <b>Price-to-Earnings (P/E) Ratio</b> Understanding earnings can be essential to determining the value of a company. To analyze earnings growth, investors can use earnings per share (EPS). This calculation measures the amount of earnings allocated to each share of stock. The P/E ratio is interpreted as a multiple, or measure, of how much an investor is paying for the stock compared to each dollar of a company’s annual earnings.
# <li><strong>Value investors</strong> are typically looking for stocks with a relatively low P/E ratio. This means they’re paying less for each dollar of earnings, which is a major characteristic of a value stock.</li>
# <dl>
# <dd>A cross-sectional analysis can provide more insight than the P/E ratio by itself. Comparing a company’s P/E ratio to a similar company or industry average can potentially reveal how well a company is actually performing. A good value stock candidate is likely to have a low P/E ratio relative to other companies in the same industry. </dd><br>
# <dd>P/E ratio referred to as a trailing P/E ratio or a forward P/E ratio. The difference is in the source of the earnings figure. A trailing P/E uses the company’s last 12 months of earnings, while a forward P/E uses an estimate of the company’s next 12 months of earnings.</dd><br>
# <dd>If a company uses aggressive accounting, the P/E ratio may appear artificially low. That’s why it can be important to also use other ratios like the price-to-sales and price-to-book ratios</dd>
# </dl>
# <li><strong>Growth investors</strong> are typically looking for stocks with a relatively high P/E ratio.</li>
# <img src="../reports/figures/pe_ratio.png" alt="Drawing" width="600"></img>
# </div>
# <div class="alert alert-block alert-info">
# <b>Price-to-Sales (P/S) Ratio</b> The price-to-sales (P/S) ratio compares a stock’s price to, you guessed it, the company’s annual sales per share. This is a simple measure of how much revenue the company brings in from its sales activity. <br>
# <li><strong>Value investors</strong> are typically looking for stocks with a relatively low P/S ratio. The P/S ratio is interpreted as a multiple of how much an investor is paying for the stock compared to each dollar of a company’s annual sales.</li>
# <dl>
# <dd>One downside to using a P/S ratio is that sales numbers can often vary greatly from year-to-year or even quarter-to-quarter. Furthermore, it may seem redundant to compare price to such similar metrics as sales (P/S ratio) and earnings (P/E ratio).</dd>
# </dl>
# <img src="../reports/figures/ps_ratio.png" alt="Drawing" width="400"></img>
# </div>
# <div class="alert alert-block alert-info">
# <b>Price-to-Book (P/B) Ratio</b> The P/B ratio compares the stock price to a company’s book value, which is the per-share value of shareholders’ equity.<strong> Shareholders’ equity, or book value,</strong> is a balance sheet item equal to the company’s assets minus liabilities.<br>
# <b>The book value growth rate</b>, or the percentage increase in shareholders’ equity over time, can reveal important clues about a company’s strength: an increasing book value might indicate an increase in shareholders’ equity. You typically want to see a company’s book value grow over time.<br>
# <img src="../reports/figures/bv.png" alt="Drawing" width="400"></img>
# <li><strong>Value investors</strong> generally prefer to invest in stocks with a low P/B ratio. This can indicate that they’re paying a lower multiple for the equity in the business.</li>
# <dl>
# <dd>Book value may not accurately represent the economic value of a company’s assets for two main reasons. First, book value ignores the economic value of intangible assets like brands or intellectual property. Second, book values may not reflect various accounting practices and how companies choose to record assets on their balance sheets. Despite its limitations, the P/B ratio can be a useful tool in evaluating and searching for value stocks when combined with other ratios.</dd>
# </dl>
# <img src="../reports/figures/pb_ratio.png" alt="Drawing" width="400"></img>
# </div>
#
income_statement
# ({'EPS (Basic)': 'eps','EPS (Basic) Growth': 'epsgrowth','Net Income': 'netincome','Total Shareholders\' Equity': 'shareholderequity','roa': roa,'longtermdebt': longtermdebt,'interestexpense': interestexpense,'ebitda': ebitda},index=range(date.today().year-5,date.today().year))
# <div class="alert alert-block alert-info">
# <b>Balance Sheet</b> is a financial statement that reports a company's assets, liabilities and shareholders' equity. The balance sheet is one of the three (income statement and statement of cash flows being the other two) core financial statements used to evaluate a business.<br><br>
# <b>Takeaway:</b>
# <dl>
# <dd><li>The balance sheet is one of the three (income statement and statement of cash flows being the other two) core financial statements used to evaluate a business.</li></dd>
# <dd><li>The balance sheet is a snapshot, representing the state of a company's finances (what it owns and owes) as of the date of publication.</li></dd>
# <dd><li>Fundamental analysts use balance sheets, in conjunction with other financial statements, to calculate financial ratios.</li></dd>
# </dl>
# </div>
balance_sheet
# <div class="alert alert-block alert-info">
# <b>Cash Flow Statement</b> a cash flow statement, also known as statement of cash flows, is a financial statement that shows how changes in balance sheet accounts and income affect cash and cash equivalents, and breaks the analysis down to operating, investing, and financing activities.<br><br>
# <b>Takeaway:</b>
# <dl>
# <dd><li>The cash flow statement includes cash made by the business through operations, investment, and financing—the sum of which is called net cash flow.</li></dd>
# <dd><li>The first section of the cash flow statement is cash flow from operations, which includes transactions from all operational business activities.</li></dd>
# <dd><li>Cash flow from investment is the second section of the cash flow statement, and is the result of investment gains and losses.</li></dd>
# <dd><li>Cash flow from financing is the final section, which provides an overview of cash used from debt and equity.</li></dd>
# </dl>
# </div>
cashflow
# <div class="alert alert-block alert-success">
# <b>income_statement, balance_sheet, cashflow = get_financial_report('VLO')</b> is a function that
# generates these reports through a ticker input. A 5 year report to search for any stock on the webpage to pull in there well organized tables.
# </div>
# # Fundamental Analysis
# <div class="alert alert-block alert-info">
# <b>Fundamental Analysis:</b> is an approach to identifying investment opportunities. Fundamental investors use financial statements, economic reports, and forecasts to examine the key drivers of a company’s current and future potential business activities.<br>
# <br>
# <b>Fundamental Investors:</b> Fundamental investors often take a holistic approach by examining all of the company’s <strong>strengths, weaknesses, opportunities, and threats</strong>. These items form the basis of an investment thesis, and uncover areas that may require additional analysis.<br>
# <br>
# <li><strong>Strengths</strong> are areas that the business does well, like a company with high net profit margins.</li>
# <li><strong>Weaknesses</strong> are areas that the business doesn’t do well, like carrying too much debt.</li>
# <li><strong>Opportunities</strong> are areas that can likely be improved rather quickly.</li>
# <li><strong>Threats</strong> are areas where competitors are potentially able to outperform a company, like with a newly developed
# product.</li><br>
# <br>
# <b>Long-Term Business Trends:</b> less influenced by short-term noise. For example, imagine a company in an up-and-coming industry that has secured a 10-year patent on a revolutionary technology.<br>
# <br>
# <b>Assumptions:</b> estimating a stock’s intrinsic value or future growth potential requires making assumptions. But a stock’s market value may never catch up with the value you determine.<br>
# <br>
# <b>Forecasting Errors:</b> An error can come from many sources, including inaccurate data, accounting blunders, or simply mathematical mistakes.
# </div>
# <div class="alert alert-block alert-danger">
# <b>get_financial_reports(ticker):</b> The original function was meant to just use the following libraries<strong> requests, bs4, and datetime,</strong> however, it became apparent if any alteration was done to the site, it didn't seem proficent compared to using something like <strong>Selenium.</strong>
# </div>
import requests
from datetime import *
from bs4 import BeautifulSoup as bs
def get_financial_reports(ticker):
#build URLs
urlfinancials = 'https://www.marketwatch.com/investing/stock/'+ticker+'/financials'
urlbalancesheet = 'https://www.marketwatch.com/investing/stock/'+ticker+'/financials/balance-sheet'
#request the data using beautiful soup
text_soup_financials = BeautifulSoup(requests.get(urlfinancials).text,"html")
text_soup_balancesheet = BeautifulSoup(requests.get(urlbalancesheet).text,"html")
# build lists for Income statement
titlesfinancials = text_soup_financials.findAll('td', {'class': 'rowTitle'})
epslist=[]
netincomelist = []
longtermdebtlist = []
interestexpenselist = []
ebitdalist= []
#load data into lists if the row title is found
for title in titlesfinancials:
if 'EPS (Basic)' in title.text:
epslist.append ([td.text for td in title.findNextSiblings(attrs={'class': 'valueCell'}) if td.text])
if 'Net Income' in title.text:
netincomelist.append ([td.text for td in title.findNextSiblings(attrs={'class': 'valueCell'}) if td.text])
if 'Interest Expense' in title.text:
interestexpenselist.append ([td.text for td in title.findNextSiblings(attrs={'class': 'valueCell'}) if td.text])
if 'EBITDA' in title.text:
ebitdalist.append ([td.text for td in title.findNextSiblings(attrs={'class': 'valueCell'}) if td.text])
# find the table headers for the Balance sheet
titlesbalancesheet = text_soup_balancesheet.findAll('td', {'class': 'rowTitle'})
equitylist=[]
for title in titlesbalancesheet:
if 'Total Shareholders\' Equity' in title.text:
equitylist.append( [td.text for td in title.findNextSiblings(attrs={'class': 'valueCell'}) if td.text])
if 'Long-Term Debt' in title.text:
longtermdebtlist.append( [td.text for td in title.findNextSiblings(attrs={'class': 'valueCell'}) if td.text])
#get the data from the income statement lists
#use helper function get_element
eps = get_element(epslist,0)
epsGrowth = get_element(epslist,1)
netIncome = get_element(netincomelist,0)
shareholderEquity = get_element(equitylist,0)
roa = get_element(equitylist,1)
longtermDebt = get_element(longtermdebtlist,0)
interestExpense = get_element(interestexpenselist,0)
ebitda = get_element(ebitdalist,0)
# load all the data into dataframe
fin_df= pd.DataFrame({'eps': eps,'eps Growth': epsGrowth,'net Income': netIncome,'shareholder Equity': shareholderEquity,'roa': roa,'longterm Debt': longtermDebt,'interest Expense': interestExpense,'ebitda': ebitda}, index=range(date.today().year-5,date.today().year))
fin_df.reset_index(inplace=True)
return fin_df
#helper function
def get_element(list,element):
try:
return list[element]
except:
return '-'
| notebooks/market_watch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import scipy.stats
from sigvisa.models.wiggles.wavelets import construct_full_basis_implicit, construct_full_basis
from sigvisa.ssms_c import ARSSM, CompactSupportSSM, TransientCombinedSSM
import pyublas
basis = construct_full_basis_implicit(5.0, "db4_2.0_3_30.0")
start_idxs, end_idxs, identities, basis_prototypes, _ = basis
noise_arssm = ARSSM(np.array((0.3, -0.1, 0.2, 0.1, -0.05, 0.1, -0.05, 0.03, -0.01, 0.01), dtype=np.float), 0.01, 0.0, 0.1)
noise_sample = noise_arssm.prior_sample(100, 0)
plot(noise_sample)
x1 = np.linspace(0, 240, 1200)
t1 = np.exp(-0.02*x1)*3
t2 = np.exp(-0.03*x1)*5
plot(x1, t1)
plot(x1, t2)
# +
npts = 6000
n_basis = len(start_idxs)
components = [(noise_arssm, 0, npts, None)]
prior_means1 = np.zeros((n_basis,), dtype=np.float)
prior_vars1 = np.ones((n_basis,), dtype=np.float)
prior_vars1[identities==0] = 1.0
prior_vars1[identities==1] = 0.01
prior_vars1[identities>1] = 0.0001
prior_vars1 *= 5
cssm1 = CompactSupportSSM(start_idxs, end_idxs, identities, basis_prototypes, prior_means1, prior_vars1, 0.0, 0.0)
components.append((cssm1, 4000, len(t1), t1))
#components.append((None, 4000, len(t1), t1))
prior_means2 = np.zeros((n_basis,), dtype=np.float)
prior_vars2 = np.ones((n_basis,), dtype=np.float) *.2
cssm2 = CompactSupportSSM(start_idxs, end_idxs, identities, basis_prototypes, prior_means2, prior_vars2, 0.0, 0.0)
components.append((cssm2, 4500, len(t2), t2))
#components.append((None, 4500, len(t2), t2))
tssm = TransientCombinedSSM(components, 1e-6)
# -
s = tssm.prior_sample(6000, 0)
f = plt.figure(figsize=(15, 5))
plot(s)
lp = tssm.run_filter(s)
print lp
| notebooks/seismic_signal_model_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exploratory Data Analysis: Lyon Startup Ecosystem
# ## Introduction
#
# In this notebook, we conduct an Exploratory Data Analysis (EDA) of data about Lyon Startup Ecosystem collected from the Foutsquare API. The idea is to better understand what the startup ecosystem in Lyon in order to compare it with others cities.
# Foursquare API being a geo dataset, the analysis will mainly revolve around geographical analysis and Data Visualization on maps. Our area of investigation will focus on the category and quantity of organizations and companies that are related to startup eosystem.
#
# Our EDA approach follows the **Data Science Methodology CRISP-DM**. For more info about this approach, check this [Wikipedia page](https://en.wikipedia.org/wiki/Cross-industry_standard_process_for_data_mining)
# ## Table of contents
# 1. [Data Collection](#item1)
# 2. [Data Visualization](#item2)
# 3. [Next steps](#item3)
#
# First let's download all the libraries required for this notebook.
# +
import numpy as np # library to handle data in a vectorized manner
import pandas as pd # library for data analsysis
pd.set_option('display.max_columns', None)
pd.set_option('display.max_rows', None)
from bs4 import BeautifulSoup # library for web parsing
import json # library to handle JSON files
# #!conda install -c conda-forge geopy --yes
from geopy.geocoders import Nominatim # convert an address into latitude and longitude values
import requests # library to handle requests
import urllib.request
from pandas.io.json import json_normalize # tranform JSON file into a pandas dataframe
# Matplotlib and associated plotting modules
import matplotlib.cm as cm
import matplotlib.colors as colors
import matplotlib.pyplot as plt
# import k-means from clustering stage
from sklearn.cluster import KMeans
# #!conda install -c conda-forge folium=0.5.0 --yes
import folium # map rendering library
from lxml.html import fromstring
print('Libraries imported.')
# -
# ## 1. Data Collection <a id='item1'></a>
# #### Define Foursquare Credentials and Version
# +
CLIENT_ID = 'xx' # Foursquare ID
CLIENT_SECRET = 'xx' # Foursquare Secret
VERSION = '20180605' # Foursquare API version
print('Your credentails:')
print('CLIENT_ID: ' + CLIENT_ID)
print('CLIENT_SECRET:' + CLIENT_SECRET)
# -
# #### Category selection
# Foursquare has many categories. The full list can be accessed [here](https://developer.foursquare.com/docs/resources/categories). It goes from Restaurants to Stores and some of theses categories can widely impact any analysis just by their shear number of place. As a result, we decided to focus on selected categories that are more directely related to the life of startup, aka a Startup Ecosystem. We use the definition of a [Startup Ecosystem](https://en.wikipedia.org/wiki/Startup_ecosystem) according to Wikipedia.
# This list of category is definitely not exhaustive and could evolved based on the result of our analysis or any thoughtfull suggestions.
# Obvisously the first category we selected was **Tech Startup** (4bf58dd8d48988d125941735) as this is the main focus of our analysis.
#
# Then we selected all the services that are required to the common operation of a startup:
# - **Coworking Space** (4bf58dd8d48988d174941735)
# - **Recruiting Agency** (52f2ab2ebcbc57f1066b8b57)
# - **Financial or Legal Service** (503287a291d4c4b30a586d65)
# - **Lawyer** (<KEY>)
# - **Design Studio** (4bf58dd8d48988d1f4941735)
# - **Insurance Office** (58daa1558bbb0b01f18ec1f1)
# - **Business Service** (5453de49498eade8af355881)
# - **Business Center** (56aa371be4b08b9a8d573517)
# - **Advertising Agency** (52e81612bcbc57f1066b7a3d)
# - **Bank** (4bf58dd8d48988d10a951735)
# - **Government Building** (4bf58dd8d48988d126941735)
#
# We also selected the categories related to potential innovation partners of startup:
# - **College & University** (4d4b7105d754a06372d81259)
# - **Adult Education Center** (56aa371ce4b08b9a8d573570)
# - **Laboratory** (5744ccdfe4b0c0459246b4d6)
# - **Research Station** (58daa1558bbb0b01f18ec1b2)
#
# We also selected the categories related to potential industrial partners of startup:
# - **Industrial Estate** (56aa371be4b08b9a8d5734d7)
# - **Factory** (4eb1bea83b7b6f98df247e06)
#
# We also selected the categories related to potential cultural partners of startup:
# - **Library** (4bf58dd8d48988d12f941735)
# - **Science Museum** (4bf58dd8d48988d191941735)
# - **Art Museum** (4bf58dd8d48988d18f941735)
# - **History Museum** (4bf58dd8d48988d190941735)
# - **Planetarium** (4bf58dd8d48988d192941735)
#
#
#
# We then selected the category to various events that are required for startup to network and meet potential users or customers:
# - **Events** (4d4b7105d754a06373d81259)
# - **Trade Fair** (5bae9231bedf3950379f89c3)
#
# The full list of categories was compiled in a csv file avaliable [here](https://raw.githubusercontent.com/MattSonnati/Data_Science_Coursera_Capstone/master/startup_ecosystem_category_list.csv)
startup_ecosystem_category_list = pd.read_csv('https://raw.githubusercontent.com/MattSonnati/where_is_the_french_silicon_valley/master/startup_ecosystem_category_list.csv').dropna()
startup_ecosystem_category_list
# #### Area selection
# In many instance, the geographical area referred to by poiting at a specific city is much wider than the administrative boundaries of said city. For example when people talk about startup from the "Bay area", SF or the Silicon Valley, in fact they refer to several cities from San Fransisco, Palo alto, Mountain view up to San Jose.
#
# For Lyon startup ecosystem, we thus decided to include every cities within 50 km from Lyon.
# We thus ask Foursquare API to find all the City (50aa9e094b90af0d42d5de0d) within 50000 meters from Lyon, France
address_near = "Lyon, France"
radius = 50000
category_id = "50aa9e094b90af0d42d5de0d"
# +
# function that extracts the category name of the venue
def get_category_name(row):
try:
categories_list = row['categories']
except:
categories_list = row['venues.categories']
if len(categories_list) == 0:
return None
else:
return categories_list[0]['name']
# function that extracts the category of the venue
def get_category_id(row):
try:
categories_list = row['categories']
except:
categories_list = row['venues.categories']
if len(categories_list) == 0:
return None
else:
return categories_list[0]['id']
#create the GET request URL
LIMIT = 200
url = 'https://api.foursquare.com/v2/venues/search?client_id={}&client_secret={}&near={}&v={}&radius={}&limit={}&intent=browse&categoryId={}'.format(
CLIENT_ID,
CLIENT_SECRET,
address_near,
VERSION,
radius,
LIMIT,
category_id)
# Send the GET request
results = requests.get(url).json()
# Convert the JSON response into a dataframe
cities = results['response']['venues']
cities_list = json_normalize(cities) # flatten JSON
# filter columns
filtered_columns = ['name',
'id',
'location.postalCode',
'location.state',
'location.country',
'location.cc',
'location.lat',
'location.lng',
'categories',]
cities_list =cities_list.loc[:, filtered_columns]
# filter the category for each row
cities_list['primary_category_name'] = cities_list.apply(get_category_name, axis=1)
cities_list['primary_category_id'] = cities_list.apply(get_category_id, axis=1)
# clean columns
cities_list.columns = [col.split(".")[-1] for col in cities_list.columns]
cities_list.head()
# -
cities_list.shape
# +
# Save dataframe as csv file to localhost
cities_list.to_csv(r'lyon_cities_list.csv')
# Save dataframe as csv file to Watson studio (need token)
# project.save_data(data=cities_list.to_csv(index=False),file_name='lyon_cities_list.csv',overwrite=True)
# -
# #### List of Startup Ecosystem places in Lyon area
# For each city in the chosen area, we list all the places in our Category selection and within 5 km of said city.
def getCategoryVenues(name, latitude, longitude, radius, category_id):
# create the API request URL
url = 'https://api.foursquare.com/v2/venues/search?client_id={}&client_secret={}&ll={},{}&v={}&radius={}&limit={}&intent=browse&categoryId={}'.format(
CLIENT_ID,
CLIENT_SECRET,
latitude,
longitude,
VERSION,
radius,
LIMIT,
category_id)
# Send the GET request
results = requests.get(url).json()
# Convert the JSON response into a dataframe
category_venues = results['response']['venues']
category_venues_list = json_normalize(category_venues) # flatten JSON
# filter columns
filtered_columns = ['name',
'id',
'location.postalCode',
'location.state',
'location.country',
'location.cc',
'location.lat',
'location.lng',
'categories',]
category_venues_list = category_venues_list.loc[:, filtered_columns]
# filter the category for each row
category_venues_list['primary_category_name'] = category_venues_list.apply(get_category_name, axis=1)
category_venues_list['primary_category_id'] = category_venues_list.apply(get_category_id, axis=1)
# clean columns
category_venues_list.columns = [col.split(".")[-1] for col in category_venues_list.columns]
return(category_venues_list)
# +
radius = 5000
startup_ecosystem_place_list = pd.DataFrame(columns=['name','id','city','postalCode','state','country','cc','lat','lng','categoriesList','primary_category_name','primary_category_id'])
for category_name, category_id, color_code in zip(startup_ecosystem_category_list['4sq_category_name'],
startup_ecosystem_category_list['4sq_id'],
startup_ecosystem_category_list['color_code']):
for latitude, longitude, name in zip(cities_list['lat'],
cities_list['lng'],
cities_list['name']):
try:
startup_ecosystem_place_list = pd.concat([startup_ecosystem_place_list, getCategoryVenues(name, latitude, longitude, radius, category_id)], sort=True)
except:
pass
startup_ecosystem_place_list.drop_duplicates(subset=['id'], keep='first', inplace=True)
startup_ecosystem_place_list.head()
# -
startup_ecosystem_place_list.shape
# +
# Save dataframe as csv file to localhost
startup_ecosystem_place_list.to_csv(r'lyon_startup_ecosystem_place_list.csv')
# Save dataframe as csv file to Watson studio (need token)
# project.save_data(data=startup_ecosystem_place_list.to_csv(index=False),file_name='lyon_startup_ecosystem_place_list.csv',overwrite=True)
# -
# ## 2. Data Visualization <a id='item2'></a>
# First, let define a function to creat map from a list of Place based on their latitude and longitude coordinate.
def getMap(centerPlace, venues_list):
# create a user agent
geolocator = Nominatim(user_agent="fr_explorer")
address = centerPlace
location = geolocator.geocode(address)
latitude = location.latitude
longitude = location.longitude
# create map using latitude and longitude values
map_venues = folium.Map(location=[latitude, longitude], zoom_start=11)
# add markers to map
for lat, lng, name, postalCode in zip(venues_list['lat'],
venues_list['lng'],
venues_list['name'],
venues_list['postalCode']):
label = '{} {}'.format(name, postalCode)
label = folium.Popup(label, parse_html=True)
folium.CircleMarker(
[lat, lng],
radius=5,
popup=label,
color='blue',
fill=True,
fill_color='#3186cc',
fill_opacity=0.7,
parse_html=False).add_to(map_venues)
return(map_venues)
# Map of cities within 50 km from Lyon
getMap('Lyon, france', cities_list)
# #### To better understand Lyon Startup Ecosystem, we can take a look at the number of place in each categories.
# In order to better visualize these data, we can plot them as a horizontal bar plot.
count_per_category = startup_ecosystem_place_list.groupby('primary_category_name').count()
count_per_category.drop(['categories','categoriesList','state','postalCode','city'], axis=1, inplace=True)
count_per_category.sort_values(['id'], ascending=True, axis=0, inplace=True)
count_per_category
# +
# plot data
count_per_category['id'].tail(50).plot(kind='barh', figsize=(10, 20))
plt.xlabel('Number of places') # add to x-label to the plot
plt.ylabel('Place Category') # add y-label to the plot
plt.title('Ranking of Place Category in Lyon Startup Ecosystem') # add title to the plot
plt.legend('') # add title to the plot
plt.show()
# -
# ## 3. Next Steps <a id='item3'></a>
# In this notebook, we have collected data about Lyon Startup Ecosystem. We then had a quick glance about the data through a map and an histogram. These data will be combined with others similar data from other Startup Ecosystem in the world then analyzed in a subsequent notebook. Then we will try to compare and identify similarity among these Startup Ecosystem through machine learning clustering techniques.
# <hr>
#
# Copyright © 2019 [<NAME>](https://www.linkedin.com/in/matthieusonnati/). This notebook and its source code are released under the terms of the [MIT License](https://bigdatauniversity.com/mit-license/).
| EDA Foursquare - Lyon Startup Ecosystem.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.7 ('base')
# language: python
# name: python3
# ---
import pandas as pd
import pickle
import sys
from Preprocessor import Preprocessor
from Vectorizing.CountVectorizer import getCountVectorizer
from Vectorizing.TF_IDF_Vectorizer import getWordLevelVectorizer
import os
df_train = pd.read_csv('../data/train_test_data/readme_new_preprocessed_test.csv', sep=';')
df_train = df_train.drop(df_train[df_train['Label'] == 'General'].index)
#df_train = df_train.drop(df_train[df_train['Text'] == ''].index)
#Preprocessor(df_train).run()
TEXT = "Text"
LABEL = "Label"
x_train = df_train[TEXT].astype('U')
y_train = df_train[LABEL]
# +
#count_vect = getWordLevelVectorizer(df_train, TEXT)
#from sklearn.feature_extraction.text import TfidfVectorizer
#count_vect = TfidfVectorizer(analyzer='word', token_pattern=r'\w{1,}', max_features=10000)
#count_vect.fit(df_train[TEXT])
# -
dir = '../results/models/demo1/'
models = os.listdir(dir)
for model in models:
print(model)
if 'audio' in model:
audio_clf = pickle.load(open(dir+model, 'rb'))
if 'computer_vision' in model:
cv_clf = pickle.load(open(dir+model, 'rb'))
if 'graphs' in model:
graphs_clf = pickle.load(open(dir+model, 'rb'))
if 'natural' in model:
nlp_clf = pickle.load(open(dir+model, 'rb'))
if 'reinforcement' in model:
rl_clf = pickle.load(open(dir+model, 'rb'))
if 'sequential' in model:
seq_clf = pickle.load(open(dir+model, 'rb'))
# +
#x = ['some text'])
#x
# +
threshold = 0.7
good = 0
bad = 0
df_train[TEXT] = df_train[TEXT].astype('U')
for ind, row in df_train.iterrows():
text = row[TEXT]
#print([row[TEXT]])
#print(row[TEXT])
if row[TEXT]:
pass
outs = []
try:
outs.append(audio_clf.predict([row[TEXT]])[0])
except:
print(row[TEXT])
[proba] = cv_clf.predict_proba([text])
print(outs, proba)
if max(proba) < threshold: outs.append('Other')
else: outs.append(cv_clf.predict([text])[0])
[proba] = graphs_clf.predict_proba([text])
if max(proba) < threshold: outs.append('Other')
else: outs.append(graphs_clf.predict([text])[0])
[proba] = nlp_clf.predict_proba([text])
if max(proba) < threshold: outs.append('Other')
else: outs.append(nlp_clf.predict([text])[0])
[proba] = rl_clf.predict_proba([text])
if max(proba) < threshold: outs.append('Other')
else: outs.append(rl_clf.predict([text])[0])
[proba] = seq_clf.predict_proba([text])
if max(proba) < threshold: outs.append('Other')
else: outs.append(seq_clf.predict([text])[0])
labels = list(df_train[df_train['Repo'] == row['Repo']][LABEL])
outs = [i for i in outs if i != 'Other']
#print('True labels: ', labels, ' Predicted: ', outs)
if outs == labels: good += 1
#if set(outs) & (set(labels)): good += 1
#if set(outs).issubset(set(labels)): good += 1
else:
bad += 1
#print(f'True: {labels}')
#print(f'Predicted: {outs}')
break
print('Good: ', good)
print('Bad: ', bad)
print('Accuracy: ', good/(good+bad))
# +
text = ['a said this is an old story']
text += ['car detect yolo intend week 3 program assign']
text += ['']
text += ['text, ']
x = audio_clf.predict(text)
print(f'Audio: {audio_clf.predict_proba(text)}')
print('Audio: ', x)
x = cv_clf.predict(text)
print(f'CV: {cv_clf.predict_proba(text)}')
print('CV: ', x)
x = graphs_clf.predict(text)
print('Graphs:', x)
print(f'Graphs: {graphs_clf.predict_proba(text)}')
x = nlp_clf.predict(text)
print('NLP:', x)
print(f'NLP: {nlp_clf.predict_proba(text)}')
x = rl_clf.predict(text)
print('RL:', x)
print(f'RL: {rl_clf.predict_proba(text)}')
x = seq_clf.predict(text)
print('Seq:', x)
print(f'Seq: {seq_clf.predict_proba(text)}')
| src/util/demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Track demo
#
# Track vessel by mmsi. Note if querying for large areas then you should consider using a larger notebook or breaking the area into smaller queries and using [user defined functions](https://docs.tiledb.com/cloud/client-api/serverless-array-udfs)
# +
import time
import datashader as ds
from datashader.utils import lnglat_to_meters
import holoviews as hv
import holoviews.operation.datashader as hd
from holoviews.element import tiles
import matplotlib.pyplot as plt
from matplotlib import cm
import numpy as np
import geopandas as gpd
import tiledb
import warnings
warnings.filterwarnings("ignore")
hv.extension("bokeh", "matplotlib")
tiledb_mmsi_uri = 'tiledb://spire-data/mmsi_ais'
config = tiledb.Config()
# Set value
config["sm.memory_budget"] = 50_000_000
config["sm.memory_budget_var"] = 50_000_000
ctx = tiledb.Ctx(config)
# -
# The mmsi identifier we will use is `308371000`
# +
# %%time
t1 = np.datetime64('2019-08-01T00:00:00')
t2 = np.datetime64('2019-08-15T00:00:00')
with tiledb.SparseArray(tiledb_mmsi_uri, ctx=ctx) as arr:
df = arr.query(attrs=['latitude', 'longitude'], dims=["ts_pos_utc", "mmsi"]).df[t1:t2, 308371000]
print(f"Retrieved {len(df['longitude'])} points")
df.sort_values(by=['ts_pos_utc'], inplace=True, ascending=True)
df.loc[:, 'x'], df.loc[:, 'y'] = lnglat_to_meters(df.longitude,df.latitude)
df
# -
# Using existing data science tooling we can plot these points on a map
gdf = gpd.GeoDataFrame(df, geometry=gpd.points_from_xy(df.longitude, df.latitude), crs='epsg:4326')
ax = gdf.plot(markersize=1.5, figsize=(8, 8))
plt.autoscale(False)
world = gpd.read_file(gpd.datasets.get_path('naturalearth_lowres'))
world.to_crs(gdf.crs).plot(ax=ax, color='none', edgecolor='black')
# +
# %%time
bkgrd = tiles.EsriImagery().opts(xaxis=None, yaxis=None, width=700, height=500)
opts = hv.opts.RGB(width=500, height=500)
bkgrd * hd.datashade(hv.Path(gdf, kdims=['x','y']), cmap='red', normalization='linear', aggregator=ds.any()).opts(opts)
# -
| geo/spire/Ship_Tracks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from IPython.display import clear_output
from array import array
import datetime
from datetime import timedelta, date #for time duration calculations
from dateutil.parser import parse #for fuzzy finding year
import pickle #for saving output files, pickles
from sys import stdout
import time #for time.sleep function to delay calls
from tqdm import tqdm #for updating loop
#from os import listdir
#from os.path import isfile, join
import glob #pattern matching and expansion.
# +
intervals = (
('weeks', 604800), # 60 * 60 * 24 * 7
('days', 86400), # 60 * 60 * 24
('hours', 3600), # 60 * 60
('minutes', 60),
('seconds', 1),
)
def display_time(seconds, granularity=2):
result = []
for name, count in intervals:
value = seconds // count
if value:
seconds -= value * count
if value == 1:
name = name.rstrip('s')
result.append("{} {}".format(value, name))
return ', '.join(result[:granularity])
# Function convert seconds into day.decimal
def ConvertSectoDay(n):
day = n // (24 * 3600)
#print(day) #keep day
n = n % (24 * 3600)
daydec=(n/86400) # add this to day
addem=day+daydec
#https://stackoverflow.com/a/48812729/1602288
holder='{:g}'.format(float('{:.{p}g}'.format(addem, p=5)))
return(float(holder))
# +
## Python packages - you may have to pip install sqlalchemy, sqlalchemy_utils, and psycopg2.
from sqlalchemy import create_engine
from sqlalchemy_utils import database_exists, create_database
from sqlalchemy.sql import table, column, select, update, insert
import psycopg2
from psycopg2.extensions import ISOLATION_LEVEL_AUTOCOMMIT
import pandas as pd
#In Python: Define your username and password used above. I've defined the database name (we're
#using a dataset on births, so I call it birth_db).
dbname = 'donors_db'
username = 'xxxx'#enter username
pswd = '<PASSWORD>'#enter password
# -
## 'engine' is a connection to a database
## Here, we're using postgres, but sqlalchemy can connect to other things too.
engine = create_engine('postgresql://%s:%s@localhost/%s'%(username,pswd,dbname))
print('postgresql://%s:%s@localhost/%s'%(username,pswd,dbname))
print(engine.url)
# Replace localhost with IP address if accessing a remote server
## create a database (if it doesn't exist)
if not database_exists(engine.url):
create_database(engine.url)
print(database_exists(engine.url))
print(engine.url)
# connect:
con = None
con = psycopg2.connect(database = dbname, user = username, host='localhost', password=<PASSWORD>)
# +
### query: from historical data
merge_query = """
SELECT * FROM merge_projects;
"""
mergedframe = pd.read_sql_query(merge_query,con)
# -
mergedframe = mergedframe[mergedframe.funding_status == 'completed']
mergedframe.head(2)
def elapsedseconds(posted, completed):
formatuse = '%Y-%m-%d %H:%M:%S' # The format: see down this page:https://docs.python.org/3/library/datetime.html
otherformat = '%Y-%m-%d'
try:
clock = datetime.datetime.strptime(completed,formatuse)
except:
clock = datetime.datetime.strptime(completed,otherformat)
try:
startclock = datetime.datetime.strptime(posted,formatuse)
except:
startclock = datetime.datetime.strptime(posted,otherformat)
elapsed = (clock-startclock).total_seconds()
return(elapsed)
#https://stackoverflow.com/a/40353780/1602288
mergedframe['latency_to_funded'] = mergedframe.apply(lambda row: elapsedseconds(row['date_posted'],row['date_completed']),axis=1)
mergedframe['days_to_funding'] = mergedframe.apply(lambda row: ConvertSectoDay(row.latency_to_funded),axis=1)
pd.set_option('display.max_columns', None)
mergedframe.head()
# +
##save new merged
#mergedframe.to_sql('merge_time', engine, if_exists='append')
# +
sns.set_context("poster", font_scale=1.3)
fig, ax = plt.subplots(figsize=(12, 8))
sns.distplot(mergedframe["days_to_funding"].dropna())
fig.tight_layout()
# -
print("Avg time to funding: "+str(display_time(mergedframe['latency_to_funded'].mean(),3)))
print("Avg time to funding: "+str(mergedframe['days_to_funding'].mean())+" days")
# +
neweval = mergedframe['days_to_funding'].describe()
readval = pd.Series([display_time(each,3) for each in neweval])
print((neweval))
print((readval))
# -
mergedframe = mergedframe[mergedframe.days_to_funding < 200]
mergedframe.shape
x = mergedframe.total_price_excluding_optional_support
y = mergedframe.days_to_funding
# +
sns.set_context("poster", font_scale=1.3)
fig, ax = plt.subplots(figsize=(12, 8))
g =sns.scatterplot(x="total_price_excluding_optional_support", y="days_to_funding",
hue="primary_focus_subject",alpha=0.1,
data=mergedframe,legend=False);
g.set(xscale="log");
fig.tight_layout()
# -
mergedframe['funding_status'].value_counts()
mergedframe['poverty_level_y'].value_counts()
#pd.mergedframe('display.max_columns', None)
list(mergedframe['fulfillmentTrailer'])
#mergedframe.head(2)
### Close communication with the database
con.close()
| testscripts/.ipynb_checkpoints/feature_engineering-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from statsmodels.tsa.stattools import adfuller
from statsmodels.graphics.tsaplots import plot_acf, plot_pacf
tmp = pd.read_csv('Dataset.csv')
print(tmp.head())
# tmp2 = tmp[['Date(UTC)','Price']]
# tmp2['day'] = pd.to_datetime(tmp2['Date(UTC)'], format = "%m/%d/%Y")
# print(tmp2['day'])
# print(tmp2.head())
# tmp2[0:1552].plot()
# plt.figure()
# pd.plotting.autocorrelation_plot(tmp2[0:1552])
tmp['Date(UTC)'] = pd.to_datetime(tmp['Date(UTC)'])
# tmp2 = tmp[['Date','Price']]
print(tmp.corr(method='pearson'))
tmp.dropna()
# Converting the data to a logarithmic scale
eth_log = pd.DataFrame(np.log(tmp[10:1500].Price)).dropna()
eth_log.plot(figsize=(12,5))
plt.figure()
# Differencing the log values
et_log = pd.DataFrame(np.log(tmp[10:1500].Price))
# Differencing the log values
lo_diff = et_log.diff().dropna()
lo_diff.plot()
plt.figure()
eth_log = tmp.loc[:, ~tmp.columns.isin(['EthereumGTrends','Date(UTC)'])][10:1552].apply(np.log).dropna()
log_data = eth_log.diff()
#print(log_data.Price[15],lo_diff[15])
plt.figure()
log_data.BlockSize.plot(figsize=(12,5))
plt.title('Plot of the Daily Changes in Price for BTC')
plt.ylabel('Change in USD')
plt.show()
log_data.corr(method='pearson')
results = log_data.apply(adfuller)
plt.figure()
log_data.Price.plot()
fig, (ax1, ax2) = plt.subplots(2,1,figsize=(16,8))
plot_acf(lo_diff, ax=ax1, lags=150)
plot_pacf(lo_diff, ax=ax2, lags=150)
plt.savefig('acfpacf.png')
plt.show()
# -
| feature_selection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import pandas as pd
import numpy as np
TITANIC_PATH = os.path.join("datasets", "titanic")
def load_titanic_data(filename, titanic_path=TITANIC_PATH):
csv_path = os.path.join(titanic_path, filename)
return pd.read_csv(csv_path)
test_data = load_titanic_data(r'test.csv')
train_data = load_titanic_data(r'train.csv')
test_data.info()
train_data.info()
# +
# Age, Cabin and Embarced are missing some values
# Name, Sex, Ticket, Cabin and Embarked are objects
# -
train_data[['Name', 'Sex', 'Ticket', 'Cabin', 'Embarked', 'Pclass']]
train_data["Relatives"] = train_data["SibSp"] + train_data["Parch"]
train_data["AgeBucket"] = train_data["Age"] // 10 * 10
test_data["Relatives"] = train_data["SibSp"] + train_data["Parch"]
test_data["AgeBucket"] = train_data["Age"] // 10 * 10
# +
from sklearn.impute import SimpleImputer
imputer = SimpleImputer(strategy="mean")
# +
from sklearn.base import BaseEstimator, TransformerMixin
class MostFrequentImputer(BaseEstimator, TransformerMixin):
def fit(self, X, y=None):
self.most_frequent_ = pd.Series([X[c].value_counts().index[0] for c in X],
index=X.columns)
return self
def transform(self, X, y=None):
return X.fillna(self.most_frequent_)
# +
from sklearn.preprocessing import OneHotEncoder
one_hot = OneHotEncoder(sparse=False)
# +
from sklearn.pipeline import Pipeline
cat_pipeline = Pipeline([
('most_frequent', MostFrequentImputer()),
('one_hot', one_hot)
])
# -
num_cols = ["Survived", "Fare", "Relatives", "Parch", "SibSp", "Age", "AgeBucket"]
num_cols_test = ["Fare", "Relatives", "Parch", "SibSp", "Age", "AgeBucket"]
cat_cols = ["Sex", "Embarked", "Pclass"]
dropped_cols = ["PassengerId", "Cabin", "Name", "Ticket"]
# +
from sklearn.compose import ColumnTransformer
preprocessing_pipeline = ColumnTransformer([
("numerical", imputer, num_cols),
("categories", cat_pipeline, cat_cols),
("dropped_cols", 'drop', dropped_cols)
])
# -
preprocessing_pipeline_test = ColumnTransformer([
("numerical", imputer, num_cols_test),
("categories", cat_pipeline, cat_cols),
("dropped_cols", 'drop', dropped_cols)
])
train_data_processed = preprocessing_pipeline.fit_transform(train_data)
test_data_processed = preprocessing_pipeline_test.fit_transform(test_data)
col_names = np.concatenate([num_cols, cat_cols], axis=None)
col_names_test = np.concatenate([num_cols_test, cat_cols], axis=None)
train_data_processed = pd.DataFrame(train_data_processed)
train_data_processed = train_data_processed.drop(columns=train_data_processed.columns[[9, 10, 11, 12, 13]])
test_data_processed = pd.DataFrame(test_data_processed)
test_data_processed = test_data_processed.drop(columns=test_data_processed.columns[[9, 10, 11, 12, 13]])
test_data_processed
train_data_processed.columns = col_names
train_data_processed
test_data_processed.columns = col_names_test
test_data_processed
train_data_processed["Wife"] = (train_data_processed["Sex"].astype(int) & train_data_processed["SibSp"].astype(int))
test_data_processed["Wife"] = (test_data_processed["Sex"].astype(int) & test_data_processed["SibSp"].astype(int))
train_data_processed["If_relatives"] = (train_data_processed["Relatives"].astype(int) == True).astype(int)
test_data_processed["If_relatives"] = (test_data_processed["Relatives"].astype(int) == True).astype(int)
corr_matrix = train_data_processed.corr(method="pearson")
corr_matrix["Survived"].sort_values(ascending=False)
# +
# start looking for a model
# -
from sklearn.ensemble import RandomForestClassifier
forest_clf = RandomForestClassifier(n_estimators=100, random_state=42)
X_train = train_data_processed.drop(["Survived", "Relatives"], axis=1)
y_train = train_data_processed["Survived"]
forest_clf.fit(X_train[:400], y_train[:400])
from sklearn.model_selection import cross_val_score
score = cross_val_score(forest_clf, X_train, y_train, cv=5)
score.mean()
y_pred = forest_clf.predict(X_train)
# +
from sklearn.metrics import f1_score
f1_score(y_train, y_pred)
# +
import joblib
joblib.dump(forest_clf, "forest_clf.pkl")
# -
test_data_processed = test_data_processed.drop('Relatives', axis=1)
predictions = forest_clf.predict(test_data_processed)
submission = test_data['PassengerId']
submission = pd.DataFrame(submission)
submission.columns = ['PassengerId']
submission['Survived'] = predictions.astype(int)
submission.to_csv(r'C:\Users\Sasha\Desktop\submission_titanic.csv', index=False)
| Titanic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import itertools
# ## Create database retrieval
df = pd.read_csv('../data/filtered_df_v2.csv')
train = pd.read_csv('../data/train.csv')
df['fname'] = df['path'].apply(lambda x: x.split('/')[-1])
merged = pd.merge(df, train[['name', 'group']], how='left', left_on='fname', right_on='name')
classes = merged['class'].unique()
classes
dataset = {}
for cls in classes:
dataset[cls] = list(merged[merged['class'] == cls]['group'].value_counts().keys())
retrieval_base = pd.DataFrame(columns=['name', 'class', 'group'])
for key in dataset.keys():
retrieval_base = retrieval_base.append(train[train['group'].isin(dataset[key])][['name', 'class', 'group']])
retrieval_base.drop_duplicates(['name', 'class', 'group'], inplace=True)
retrieval_base = retrieval_base.sort_index()
retrieval_base.to_csv('../data/retrieval_base.csv', sep=',', index=False)
# ### SUPERFILTERED
retrieval_base_filtered = pd.DataFrame(columns=['name', 'class', 'group'])
classes = set(retrieval_base['class'].values)
for cls in classes:
subset = retrieval_base[retrieval_base['class'] == cls]
if len(subset) > 3:
retrieval_base_filtered = retrieval_base_filtered.append(subset.sample(3))
retrieval_base_filtered.to_csv('../data/retrieval_base_filtered.csv', sep=',', index=False)
| notebooks/database_retrieval.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# ## Seminar: Monte-carlo tree search (5 pts)
#
# Monte Carlo tree search (MCTS) is a heuristic search algorithm, which shows cool results in challenging domains such as Go and chess. The algorithm builds a search tree, iteratively traverses it, and evaluates its nodes using a Monte-Carlo simulation.
#
# In this seminar, we'll implement a MCTS([[1]](#1), [[2]](#2)) planning and use it to solve some Gym envs.
#
# 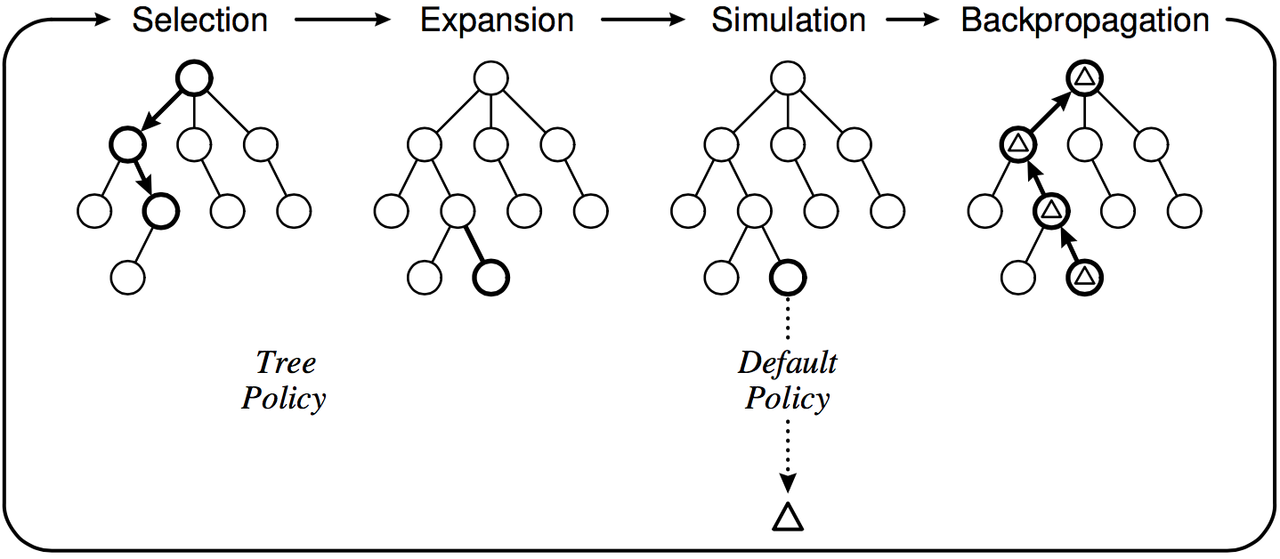
# __How it works?__
# We just start with an empty tree and expand it. There are several common procedures.
#
# __1) Selection__
# Starting from the root, recursively select the node that corresponds to the tree policy.
#
# There are several options for tree policies, which we saw earlier as exploration strategies: epsilon-greedy, Thomson sampling, UCB-1. It was shown that in MCTS, UCB-1 achieves a good result. Further, we will consider the one, but you can try to use others.
#
# Following the UCB-1 tree policy, we will choose an action that, on one hand, we expect to have the highest return, and on the other hand, we haven't explored much.
#
# $$
# \DeclareMathOperator*{\argmax}{arg\,max}
# $$
#
# $$
# \dot{a} = \argmax_{a} \dot{Q}(s, a)
# $$
#
# $$
# \dot{Q}(s, a) = Q(s, a) + C_p \sqrt{\frac{2 \log {N}}{n_a}}
# $$
#
# where:
# - $N$ - number of times we have visited state $s$,
# - $n_a$ - number of times we have taken action $a$,
# - $C_p$ - exploration balance parameter, which is performed between exploration and exploitation.
#
# Using Hoeffding inequality for rewards $R \in [0,1]$ it can be shown [[3]](#3) that optimal $C_p = 1/\sqrt{2}$. For rewards outside this range, the parameter should be tuned. We'll be using 10, but you can experiment with other values.
#
# __2) Expansion__
# After the selection procedure, we can achieve a leaf node or node in which we don't complete actions. In this case, we expand the tree by feasible actions and get new state nodes.
#
# __3) Simulation__
# How we can estimate node Q-values? The idea is to estimate action values for a given _rollout policy_ by averaging the return of many simulated trajectories from the current node. Simply, we can play with random or some special policy or use some model that can estimate it.
#
# __4) Backpropagation__
# The reward of the last simulation is backed up through the traversed nodes and propagates Q-value estimations, upwards to the root.
#
# $$
# Q({\text{parent}}, a) = r + \gamma \cdot Q({\text{child}}, a)
# $$
#
# There are a lot modifications of MCTS, more details about it you can find in this paper [[4]](#4)
# +
import sys, os
if 'google.colab' in sys.modules and not os.path.exists('.setup_complete'):
# !wget -q https://raw.githubusercontent.com/yandexdataschool/Practical_RL/master/setup_colab.sh -O- | bash
# !wget -q https://raw.githubusercontent.com/yandexdataschool/Practical_RL/coursera/grading.py -O ../grading.py
# !wget -q https://raw.githubusercontent.com/yandexdataschool/Practical_RL/coursera/week6_outro/submit.py
# !touch .setup_complete
# This code creates a virtual display to draw game images on.
# It will have no effect if your machine has a monitor.
if type(os.environ.get("DISPLAY")) is not str or len(os.environ.get("DISPLAY")) == 0:
# !bash ../xvfb start
os.environ['DISPLAY'] = ':1'
# -
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# ---
#
# But before we do that, we first need to make a wrapper for Gym environments to allow saving and loading game states to facilitate backtracking.
# +
import gym
from gym.core import Wrapper
from pickle import dumps, loads
from collections import namedtuple
# a container for get_result function below. Works just like tuple, but prettier
ActionResult = namedtuple(
"action_result", ("snapshot", "observation", "reward", "is_done", "info"))
class WithSnapshots(Wrapper):
"""
Creates a wrapper that supports saving and loading environemnt states.
Required for planning algorithms.
This class will have access to the core environment as self.env, e.g.:
- self.env.reset() #reset original env
- self.env.ale.cloneState() #make snapshot for atari. load with .restoreState()
- ...
You can also use reset() and step() directly for convenience.
- s = self.reset() # same as self.env.reset()
- s, r, done, _ = self.step(action) # same as self.env.step(action)
Note that while you may use self.render(), it will spawn a window that cannot be pickled.
Thus, you will need to call self.close() before pickling will work again.
"""
def get_snapshot(self, render=False):
"""
:returns: environment state that can be loaded with load_snapshot
Snapshots guarantee same env behaviour each time they are loaded.
Warning! Snapshots can be arbitrary things (strings, integers, json, tuples)
Don't count on them being pickle strings when implementing MCTS.
Developer Note: Make sure the object you return will not be affected by
anything that happens to the environment after it's saved.
You shouldn't, for example, return self.env.
In case of doubt, use pickle.dumps or deepcopy.
"""
if render:
self.render() # close popup windows since we can't pickle them
self.close()
if self.unwrapped.viewer is not None:
self.unwrapped.viewer.close()
self.unwrapped.viewer = None
return dumps(self.env)
def load_snapshot(self, snapshot, render=False):
"""
Loads snapshot as current env state.
Should not change snapshot inplace (in case of doubt, deepcopy).
"""
assert not hasattr(self, "_monitor") or hasattr(
self.env, "_monitor"), "can't backtrack while recording"
if render:
self.render() # close popup windows since we can't load into them
self.close()
self.env = loads(snapshot)
def get_result(self, snapshot, action):
"""
A convenience function that
- loads snapshot,
- commits action via self.step,
- and takes snapshot again :)
:returns: next snapshot, next_observation, reward, is_done, info
Basically it returns next snapshot and everything that env.step would have returned.
"""
<YOUR CODE: load, commit, take snapshot>
return ActionResult(
<YOUR CODE: next_snapshot>, # fill in the variables
<YOUR CODE: next_observation>,
<YOUR CODE: reward>,
<YOUR CODE: is_done>,
<YOUR CODE: info>,
)
# -
# ### Try out snapshots:
# Let`s check our wrapper. At first, reset environment and save it, further randomly play some actions and restore our environment from the snapshot. It should be the same as our previous initial state.
# +
# make env
env = WithSnapshots(gym.make("CartPole-v0"))
env.reset()
n_actions = env.action_space.n
# +
print("initial_state:")
plt.imshow(env.render('rgb_array'))
env.close()
# create first snapshot
snap0 = env.get_snapshot()
# +
# play without making snapshots (faster)
while True:
is_done = env.step(env.action_space.sample())[2]
if is_done:
print("Whoops! We died!")
break
print("final state:")
plt.imshow(env.render('rgb_array'))
env.close()
# +
# reload initial state
env.load_snapshot(snap0)
print("\n\nAfter loading snapshot")
plt.imshow(env.render('rgb_array'))
env.close()
# +
# get outcome (snapshot, observation, reward, is_done, info)
res = env.get_result(snap0, env.action_space.sample())
snap1, observation, reward = res[:3]
# second step
res2 = env.get_result(snap1, env.action_space.sample())
# -
# # MCTS: Monte-Carlo tree search
#
#
# We will start by implementing the `Node` class - a simple class that acts like MCTS node and supports some of the MCTS algorithm steps.
#
# This MCTS implementation makes some assumptions about the environment, you can find those _in the notes section at the end of the notebook_.
assert isinstance(env, WithSnapshots)
class Node:
"""A tree node for MCTS.
Each Node corresponds to the result of performing a particular action (self.action)
in a particular state (self.parent), and is essentially one arm in the multi-armed bandit that
we model in that state."""
# metadata:
parent = None # parent Node
qvalue_sum = 0. # sum of Q-values from all visits (numerator)
times_visited = 0 # counter of visits (denominator)
def __init__(self, parent, action):
"""
Creates and empty node with no children.
Does so by commiting an action and recording outcome.
:param parent: parent Node
:param action: action to commit from parent Node
"""
self.parent = parent
self.action = action
self.children = set() # set of child nodes
# get action outcome and save it
res = env.get_result(parent.snapshot, action)
self.snapshot, self.observation, self.immediate_reward, self.is_done, _ = res
def is_leaf(self):
return len(self.children) == 0
def is_root(self):
return self.parent is None
def get_qvalue_estimate(self):
return self.qvalue_sum / self.times_visited if self.times_visited != 0 else 0
def ucb_score(self, scale=10, max_value=1e100):
"""
Computes ucb1 upper bound using current value and visit counts for node and it's parent.
:param scale: Multiplies upper bound by that. From Hoeffding inequality,
assumes reward range to be [0, scale].
:param max_value: a value that represents infinity (for unvisited nodes).
"""
if self.times_visited == 0:
return max_value
# compute ucb-1 additive component (to be added to mean value)
# hint: you can use self.parent.times_visited for N times node was considered,
# and self.times_visited for n times it was visited
U = <YOUR CODE>
return self.get_qvalue_estimate() + scale * U
# MCTS steps
def select_best_leaf(self):
"""
Picks the leaf with the highest priority to expand.
Does so by recursively picking nodes with the best UCB-1 score until it reaches a leaf.
"""
if self.is_leaf():
return self
children = self.children
# Select the child node with the highest UCB score. You might want to implement some heuristics
# to break ties in a smart way, although CartPole should work just fine without them.
best_child = <YOUR CODE>
return best_child.select_best_leaf()
def expand(self):
"""
Expands the current node by creating all possible child nodes.
Then returns one of those children.
"""
assert not self.is_done, "can't expand from terminal state"
for action in range(n_actions):
self.children.add(Node(self, action))
# If you have implemented any heuristics in select_best_leaf(), they will be used here.
# Otherwise, this is equivalent to picking some undefined newly created child node.
return self.select_best_leaf()
def rollout(self, t_max=10**4):
"""
Play the game from this state to the end (done) or for t_max steps.
On each step, pick action at random (hint: env.action_space.sample()).
Compute sum of rewards from the current state until the end of the episode.
Note 1: use env.action_space.sample() for picking a random action.
Note 2: if the node is terminal (self.is_done is True), just return self.immediate_reward.
"""
# set env into the appropriate state
env.load_snapshot(self.snapshot)
obs = self.observation
is_done = self.is_done
<YOUR CODE: perform rollout and compute reward>
return rollout_reward
def propagate(self, child_qvalue):
"""
Uses child Q-value (sum of rewards) to update parents recursively.
"""
# compute node Q-value
my_qvalue = self.immediate_reward + child_qvalue
# update qvalue_sum and times_visited
self.qvalue_sum += my_qvalue
self.times_visited += 1
# propagate upwards
if not self.is_root():
self.parent.propagate(my_qvalue)
def safe_delete(self):
"""safe delete to prevent memory leak in some python versions"""
del self.parent
for child in self.children:
child.safe_delete()
del child
class Root(Node):
def __init__(self, snapshot, observation):
"""
creates special node that acts like tree root
:snapshot: snapshot (from env.get_snapshot) to start planning from
:observation: last environment observation
"""
self.parent = self.action = None
self.children = set() # set of child nodes
# root: load snapshot and observation
self.snapshot = snapshot
self.observation = observation
self.immediate_reward = 0
self.is_done = False
@staticmethod
def from_node(node):
"""initializes node as root"""
root = Root(node.snapshot, node.observation)
# copy data
copied_fields = ["qvalue_sum", "times_visited", "children", "is_done"]
for field in copied_fields:
setattr(root, field, getattr(node, field))
return root
# ## Main MCTS loop
#
# With all we implemented, MCTS boils down to a trivial piece of code.
def plan_mcts(root, n_iters=10):
"""
builds tree with monte-carlo tree search for n_iters iterations
:param root: tree node to plan from
:param n_iters: how many select-expand-simulate-propagete loops to make
"""
for _ in range(n_iters):
node = <YOUR CODE: select best leaf>
if node.is_done:
# All rollouts from a terminal node are empty, and thus have 0 reward.
node.propagate(0)
else:
# Expand the best leaf. Perform a rollout from it. Propagate the results upwards.
# Note that here you have some leeway in choosing where to propagate from.
# Any reasonable choice should work.
<YOUR CODE>
# ## Plan and execute
#
# Let's use our MCTS implementation to find the optimal policy.
env = WithSnapshots(gym.make("CartPole-v0"))
root_observation = env.reset()
root_snapshot = env.get_snapshot()
root = Root(root_snapshot, root_observation)
# plan from root:
plan_mcts(root, n_iters=1000)
# +
# import copy
# saved_root = copy.deepcopy(root)
# root = saved_root
# +
from IPython.display import clear_output
from itertools import count
from gym.wrappers import Monitor
total_reward = 0 # sum of rewards
test_env = loads(root_snapshot) # env used to show progress
for i in count():
# get best child
best_child = <YOUR CODE: select child with the highest mean reward>
# take action
s, r, done, _ = test_env.step(best_child.action)
# show image
clear_output(True)
plt.title("step %i" % i)
plt.imshow(test_env.render('rgb_array'))
plt.show()
total_reward += r
if done:
print("Finished with reward = ", total_reward)
break
# discard unrealized part of the tree [because not every child matters :(]
for child in root.children:
if child != best_child:
child.safe_delete()
# declare best child a new root
root = Root.from_node(best_child)
assert not root.is_leaf(), \
"We ran out of tree! Need more planning! Try growing the tree right inside the loop."
# You may want to run more planning here
# <YOUR CODE>
# +
from submit import submit_mcts
submit_mcts(total_reward, '<EMAIL>', 'YourAssignmentToken')
# -
# ## Bonus assignments (10+pts each)
#
# There's a few things you might want to try if you want to dig deeper:
#
# ### Node selection and expansion
#
# "Analyze this" assignment
#
# UCB-1 is a weak bound as it relies on a very general bounds (Hoeffding Inequality, to be exact).
# * Try playing with the exploration parameter $C_p$. The theoretically optimal $C_p$ you can get from a max reward of the environment (max reward for CartPole is 200).
# * Use using a different exploration strategy (bayesian UCB, for example)
# * Expand not all but several random actions per `expand` call. See __the notes below__ for details.
#
# The goal is to find out what gives the optimal performance for `CartPole-v0` for different time budgets (i.e. different n_iter in plan_mcts.)
#
# Evaluate your results on `Acrobot-v1` - do the results change and if so, how can you explain it?
#
#
# ### Atari-RAM
#
# "Build this" assignment
#
# Apply MCTS to play Atari games. In particular, let's start with ```gym.make("MsPacman-ramDeterministic-v0")```.
#
# This requires two things:
# * Slightly modify WithSnapshots wrapper to work with atari.
#
# * Atari has a special interface for snapshots:
# ```
# snapshot = self.env.ale.cloneState()
# ...
# self.env.ale.restoreState(snapshot)
# ```
# * Try it on the env above to make sure it does what you told it to.
#
# * Run MCTS on the game above.
# * Start with small tree size to speed-up computations
# * You will probably want to rollout for 10-100 steps (t_max) for starters
# * Consider using discounted rewards (see __notes at the end__)
# * Try a better rollout policy
#
#
# ### Integrate learning into planning
#
# Planning on each iteration is a costly thing to do. You can speed things up drastically if you train a classifier to predict which action will turn out to be best according to MCTS.
#
# To do so, just record which action did the MCTS agent take on each step and fit something to [state, mcts_optimal_action]
# * You can also use optimal actions from discarded states to get more (dirty) samples. Just don't forget to fine-tune without them.
# * It's also worth a try to use P(best_action|state) from your model to select best nodes in addition to UCB
# * If your model is lightweight enough, try using it as a rollout policy.
#
# While CartPole is glorious enough, try expanding this to ```gym.make("MsPacmanDeterministic-v0")```
# * See previous section on how to wrap atari
#
# * Also consider what [AlphaGo Zero](https://deepmind.com/blog/alphago-zero-learning-scratch/) did in this area.
#
# ### Integrate planning into learning
# _(this will likely take long time, better consider this as side project when all other deadlines are met)_
#
# Incorporate planning into the agent architecture. The goal is to implement [Value Iteration Networks](https://arxiv.org/abs/1602.02867).
#
# Remember [week5 assignment](https://github.com/yandexdataschool/Practical_RL/blob/coursera/week5_policy_based/practice_a3c.ipynb)? You will need to switch it into a maze-like game, like MsPacman, and implement a special layer that performs value iteration-like update to a recurrent memory. This can be implemented the same way you did in the POMDP assignment.
# ## Notes
#
#
# #### Assumptions
#
# The full list of assumptions is:
#
# * __Finite number of actions__: we enumerate all actions in `expand`.
# * __Episodic (finite) MDP__: while technically it works for infinite MDPs, we perform a rollout for $10^4$ steps. If you are knowingly infinite, please adjust `t_max` to something more reasonable.
# * __Deterministic MDP__: `Node` represents the single outcome of taking `self.action` in `self.parent`, and does not support the situation where taking an action in a state may lead to different rewards and next states.
# * __No discounted rewards__: we assume $\gamma=1$. If that isn't the case, you only need to change two lines in `rollout()` and use `my_qvalue = self.immediate_reward + gamma * child_qvalue` for `propagate()`.
# * __pickleable env__: won't work if e.g. your env is connected to a web-browser surfing the internet. For custom envs, you may need to modify get_snapshot/load_snapshot from `WithSnapshots`.
#
# #### On `get_best_leaf` and `expand` functions
#
# This MCTS implementation only selects leaf nodes for expansion.
# This doesn't break things down because `expand` adds all possible actions. Hence, all non-leaf nodes are by design fully expanded and shouldn't be selected.
#
# If you want to only add a few random action on each expand, you will also have to modify `get_best_leaf` to consider returning non-leafs.
#
# #### Rollout policy
#
# We use a simple uniform policy for rollouts. This introduces a negative bias to good situations that can be messed up completely with random bad action. As a simple example, if you tend to rollout with uniform policy, you better don't use sharp knives and walk near cliffs.
#
# You can improve that by integrating a reinforcement _learning_ algorithm with a computationally light agent. You can even train this agent on optimal policy found by the tree search.
#
# #### Contributions
# * Reusing some code from 5vision [solution for deephack.RL](https://github.com/5vision/uct_atari), code by <NAME>
# * Using some code from [this gist](https://gist.github.com/blole/dfebbec182e6b72ec16b66cc7e331110)
#
# #### References
# * <a id="1">[1]</a> _<NAME>. (2007) Efficient Selectivity and Backup Operators in Monte-Carlo Tree Search. In: <NAME>., <NAME>., <NAME>.. (eds) Computers and Games. CG 2006. Lecture Notes in Computer Science, vol 4630. Springer, Berlin, Heidelberg_
#
# * <a id="2">[2]</a> _<NAME>., <NAME>. (2006) Bandit Based Monte-Carlo Planning. In: <NAME>., <NAME>., <NAME>. (eds) Machine Learning: ECML 2006. ECML 2006. Lecture Notes in Computer Science, vol 4212. Springer, Berlin, Heidelberg_
#
# * <a id="3">[3]</a> _Kocsis, Levente, <NAME>, and <NAME>. "Improved monte-carlo search." Univ. Tartu, Estonia, Tech. Rep 1 (2006)._
#
# * <a id="4">[4]</a> _<NAME> et al., "A Survey of Monte Carlo Tree Search Methods," in IEEE Transactions on Computational Intelligence and AI in Games, vol. 4, no. 1, pp. 1-43, March 2012, doi: 10.1109/TCIAIG.2012.2186810._
| week6_outro/practice_mcts.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Part 3: Data Wrangling and Transformation.
# * ### StandardScaler
# +
# StandardScaler
sc = StandardScaler()
dataset_sc = sc.fit_transform(dataset)
# for under-sampling dataset
#dataset_sc = sc.fit_transform(dataset_under)
# for over-sampling dataset
#dataset_sc = sc.fit_transform(dataset_over)
dataset_sc = pd.DataFrame(dataset_sc)
dataset_sc.head()
# -
# * ### Creating datasets for ML part
# +
# set 'X' for features' and y' for the target ('quality').
y = target
X = dataset_sc.copy()
# for under-sampling dataset
#y = target_under
#X = dataset_sc.copy()
# for over-sampling dataset
#y = target_over
#X = dataset_sc.copy()
# -
# preview of the first 5 lines of the loaded data
X.head()
# * ### 'Train\Test' split
# apply 'Train\Test' splitting method
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=0)
# print shape of X_train and y_train
X_train.shape, y_train.shape
# print shape of X_test and y_test
X_test.shape, y_test.shape
# ## Part 4: Machine Learning.
# * ### Build, train and evaluate models without hyperparameters
# * Logistic Regression
# * K-Nearest Neighbors
# * Decision Trees
#
# +
# Logistic Regression
LR = LogisticRegression()
LR.fit(X_train, y_train)
LR_pred = LR.predict(X_test)
# K-Nearest Neighbors
KNN = KNeighborsClassifier()
KNN.fit(X_train, y_train)
KNN_pred = KNN.predict(X_test)
# Decision Tree
DT = DecisionTreeClassifier(random_state = 0)
DT.fit(X_train, y_train)
DT_pred = DT.predict(X_test)
# -
# * ### Classification report
print("LR Classification Report: \n", classification_report(y_test, LR_pred, digits = 6))
print("KNN Classification Report: \n", classification_report(y_test, KNN_pred, digits = 6))
print("DT Classification Report: \n", classification_report(y_test, DT_pred, digits = 6))
# * ### Confusion matrix
LR_confusion_mx = confusion_matrix(y_test, LR_pred)
print("LR Confusion Matrix: \n", LR_confusion_mx)
print()
KNN_confusion_mx = confusion_matrix(y_test, KNN_pred)
print("KNN Confusion Matrix: \n", KNN_confusion_mx)
print()
DT_confusion_mx = confusion_matrix(y_test, DT_pred)
print("DT Confusion Matrix: \n", DT_confusion_mx)
print()
# * ### ROC-AUC score
roc_auc_score(DT_pred, y_test)
# * ### Build, train and evaluate models with hyperparameters
# +
# Logistic Regression
LR = LogisticRegression()
LR_params = {'C':[1,2,3,4,5,6,7,8,9,10], 'penalty':['l1', 'l2', 'elasticnet', 'none'], 'solver':['lbfgs', 'newton-cg', 'liblinear', 'sag', 'saga'], 'random_state':[0]}
LR1 = GridSearchCV(LR, param_grid = LR_params)
LR1.fit(X_train, y_train)
LR1_pred = LR1.predict(X_test)
# K-Nearest Neighbors
KNN = KNeighborsClassifier()
KNN_params = {'n_neighbors':[5,7,9,11]}
KNN1 = GridSearchCV(KNN, param_grid = KNN_params)
KNN1.fit(X_train, y_train)
KNN1_pred = KNN1.predict(X_test)
# Decision Tree
DT = DecisionTreeClassifier()
DT_params = {'max_depth':[2,10,15,20], 'criterion':['gini', 'entropy'], 'random_state':[0]}
DT1 = GridSearchCV(DT, param_grid = DT_params)
DT1.fit(X_train, y_train)
DT1_pred = DT1.predict(X_test)
# -
# print the best hyper parameters set
print("Logistic Regression Best Hyper Parameters: ", LR1.best_params_)
print("K-Nearest Neighbour Best Hyper Parameters: ", KNN1.best_params_)
print("Decision Tree Best Hyper Parameters: ", DT1.best_params_)
# * ### Classification report
print("LR Classification Report: \n", classification_report(y_test, LR1_pred, digits = 6))
print("KNN Classification Report: \n", classification_report(y_test, KNN1_pred, digits = 6))
print("DT Classification Report: \n", classification_report(y_test, DT1_pred, digits = 6))
# * ### Confusion matrix
# +
# confusion matrix of DT model
DT_confusion_mx = confusion_matrix(y_test, DT1_pred)
print('DT Confusion Matrix')
# visualisation
ax = plt.subplot()
sns.heatmap(DT_confusion_mx, annot = True, fmt = 'd', cmap = 'Blues', ax = ax, linewidths = 0.5, annot_kws = {'size': 15})
ax.set_ylabel('FP True label TP')
ax.set_xlabel('FN Predicted label TN')
ax.xaxis.set_ticklabels(['1', '0'], fontsize = 10)
ax.yaxis.set_ticklabels(['1', '0'], fontsize = 10)
plt.show()
print()
# -
# * ### ROC-AUC score
roc_auc_score(DT1_pred, y_test)
# ## Conclusion.
# submission of .csv file with predictions
sub = pd.DataFrame()
sub['ID'] = X_test.index
sub['quality'] = DT1_pred
sub.to_csv('WinePredictionsTest.csv', index=False)
# **Question**: Predict which wines are 'Good/1' and 'Not Good/0' (use binary classification; check balance of classes; calculate perdictions; choose the best model).
#
# **Answers**:
#
# 1. Binary classification was applied.
#
# 2. Classes were highly imbalanced with 78.36 % of '0' class and only 21.64 % of '1' class in our dataset.
#
# 3. Three options were applied in order to calculate the best predictions:
# * Calculate predictions with imbalanced dataset
# * Calculate predictions with random under-sampling technique of an imbalanced dataset
# * Calculate predictions with random over-sampling technique of an imbalanced dataset
#
# 4. Three ML models were used: Logistic Regression, KNN, Decision Tree (without and with hyper parameters).
#
# 5. The best result was choosen:
# * Random over-sampling dataset with 3838 enteties in class '0' and 3838 enteties in class '1', 7676 enteties in total.
# * Train/Test split: test_size=0.2, random_state=0
# * Decision Tree model without hyper parameters tuning, with an accuracy score equal ... and ROC-AUC score equal ... .
| ML-101 Modules/Module 03/Lesson 02/Practice 2/Winequality - Practice Code Part 3&4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Decision Tree
# ### Training Data : Toy Dataset for fruit classifier
training_data = [
['Green', 3, 'Apple'],
['Yellow', 3, 'Apple'],
['Red', 1, 'Grape'],
['Red', 1, 'Grape'],
['Yellow', 3, 'Lemon'],
]
# ### Useful data and Methods for our Dataset manipulation
#Column names for our data
header = ["color","diameter","label"]
"""Find the unique values for a column in dataset"""
def unique_values(rows,col):
return set([row[col] for row in rows])
"""count the no of examples for each label in a dataset"""
def class_counts(rows):
counts = {} # a dictionary of label -> count.
for row in rows:
# in our dataset format, the label is always the last column
label = row[-1]
if label not in counts:
counts[label] = 0
counts[label] += 1
return counts
"""Check if the value is numeric"""
def is_numeric(value):
return isinstance(value, int) or isinstance(value, float)
# ### Let's write a class for a question which can be asked to partition the data
# Each object of a question class holds a column_no and a col_value
# Eg. column_no = 0 denotes color and so col_value can be Green, Yellow or Red
# We can write a method which would compare the feature value of example with the feature value of Question
class Question:
def __init__(self,col, val):
self.col = col
self.val = val
def match(self,example):
# Compare the feature value in an example to the
# feature value in this question.
value = example[self.col]
if is_numeric(value):
return value >= self.val
else:
return value == self.val
def __repr__(self):
# method to print the question in a readable format.
condition = "=="
if is_numeric(self.val):
condition = ">="
return "Is %s %s %s?" % (
header[self.col], condition, str(self.val))
# #### Question format -
#
# +
#create a new question with col = 1 and val = 3
q = Question(1,3)
#print q
q
# -
# #### Define a function which partitions the dataset on given question in True and False rows/examples
"""For each row in the dataset, check if it satisfies the question. If
so, add it to 'true rows', otherwise, add it to 'false rows'.
"""
def partition(rows, question):
true_rows, false_rows = [], []
for row in rows:
if question.match(row):
true_rows.append(row)
else:
false_rows.append(row)
return true_rows, false_rows
# #### Now calculate a Gini Impurity for a node with given input rows of training dataset
"""Calculate the Gini Impurity for a list of rows."""
def gini(rows):
counts = class_counts(rows)
impurity = 1
for lbl in counts:
prob_of_lbl = counts[lbl] / float(len(rows))
impurity -= prob_of_lbl**2
return impurity
# #### Calculate the Information gain for a question given uncertainity at present node and incertainities at left and right child nodes
def info_gain(left, right, current_uncertainty):
#we need to calculate weighted avg of impurities at both child nodes
p = float(len(left)) / (len(left) + len(right))
return current_uncertainty - p * gini(left) - (1 - p) * gini(right)
# #### Which question to ask ??
"""Find the best question to ask by iterating over every feature / value
and calculating the information gain."""
def find_best_split(rows):
best_gain = 0 # keep track of the best information gain
best_question = None # keep train of the feature / value that produced it
current_uncertainty = gini(rows)
n_features = len(rows[0]) - 1 # number of columns
for col in range(n_features): # for each feature
values = set([row[col] for row in rows]) # unique values in the column
for val in values: # for each value
question = Question(col, val)
# try splitting the dataset
true_rows, false_rows = partition(rows, question)
# Skip this split if it doesn't divide the
# dataset.
if len(true_rows) == 0 or len(false_rows) == 0:
continue
# Calculate the information gain from this split
gain = info_gain(true_rows, false_rows, current_uncertainty)
# You actually can use '>' instead of '>=' here
# but I wanted the tree to look a certain way for our
# toy dataset.
if gain >= best_gain:
best_gain, best_question = gain, question
return best_gain, best_question
# ### Define nodes in tree
# #### 1. Decision Node - Node with Question to ask
"""
A Decision Node asks a question.
This holds a reference to the question, and to the two child nodes.
"""
class Decision_Node:
def __init__(self,question,true_branch,false_branch):
self.question = question
self.true_branch = true_branch
self.false_branch = false_branch
# #### 2. Leaf node - Gives prediction
# +
"""
A Leaf node classifies data.
This holds a dictionary of class (e.g., "Apple") -> number of time it
appears in the rows from the training data that reach this leaf.
"""
class Leaf:
def __init__(self, rows):
self.predictions = class_counts(rows)
# -
# ### Build a Tree
def build_tree(rows):
# Try partitioing the dataset on each of the unique attribute,
# calculate the information gain,
# and return the question that produces the highest gain.
gain, question = find_best_split(rows)
# Base case: no further info gain
# Since we can ask no further questions,
# we'll return a leaf.
if gain == 0:
return Leaf(rows)
# If we reach here, we have found a useful feature / value
# to partition on.
true_rows, false_rows = partition(rows, question)
# Recursively build the true branch.
true_branch = build_tree(true_rows)
# Recursively build the false branch.
false_branch = build_tree(false_rows)
# Return a Question node.
# This records the best feature / value to ask at this point,
# as well as the branches to follow
# dependingo on the answer.
return Decision_Node(question, true_branch, false_branch)
# ### Print the Tree
def print_tree(node, spacing=""):
# Base case: we've reached a leaf
if isinstance(node, Leaf):
print (spacing + "Predict", node.predictions)
return
# Print the question at this node
print (spacing + str(node.question))
# Call this function recursively on the true branch
print (spacing + '--> True:')
print_tree(node.true_branch, spacing + " ")
# Call this function recursively on the false branch
print (spacing + '--> False:')
print_tree(node.false_branch, spacing + " ")
# ### All Work Done !!! Now It's time to Build a Model from given Training data
my_tree = build_tree(training_data)
print_tree(my_tree)
# ### Test the model with test data
# #### Write a function to classify the test data
def classify(row, node):
# Base case: we've reached a leaf
if isinstance(node, Leaf):
return node.predictions
# Decide whether to follow the true-branch or the false-branch.
# Compare the feature / value stored in the node,
# to the example we're considering.
if node.question.match(row):
return classify(row, node.true_branch)
else:
return classify(row, node.false_branch)
# #### Print Prediction at Leaf Node
"""A nicer way to print the predictions at a leaf."""
def print_leaf(counts):
total = sum(counts.values()) * 1.0
probs = {}
for lbl in counts.keys():
probs[lbl] = str(int(counts[lbl] / total * 100)) + "%"
return probs
# #### Check for example
print_leaf(classify(training_data[0],my_tree))
# #### Test Data
testing_data = [
['Green', 3, 'Apple'],
['Yellow', 4, 'Apple'],
['Red', 2, 'Grape'],
['Red', 1, 'Grape'],
['Yellow', 3, 'Lemon'],
]
# #### Evaluate
for row in testing_data:
print ("Actual: %s. Predicted: %s" %
(row[-1], print_leaf(classify(row, my_tree))))
| DecisionTree_Math_Fruits.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
range(3)
n=3
range(3,0)
for i in range(n,0):
print('i',i)
a=[1,2,3,4,5]
a
a[:]
a[1:-1]
a[:n]
a[:n+2]
b=a[1:n+2]
b
a[2:n+2].append(a[0])
a
a[1:n]+a[n:]+a[0:1]
a[1:n+2]+a[0:1]
a[2:n+2]+a[0:2]
a[3:n+2]+a[0:3]
a[4:n+2]+a[0:4]
n=4
c=5
for i in range(n):
b=a[i+1:c]+a[0:i+1]
b
# +
def array_left_rotation(a, n, k):
for i in range(n):
b=k[i+1:a]+k[0:i+1]
return b
# -
a=5
n=4
k=[1,2,3,4,5]
def rotate(l, n):
return l[n:] + l[:n]
rotate(k,4)
answer=array_left_rotation(a,n,k)
answer
string_a='abc'
string_b='cdef'
mapa_a={}
mapa_b={}
for char_a in string_a:
if char_a in mapa_a:
mapa_a[char_a]+=1
else:
mapa_a[char_a]=1
mapa_a
def mapear(string):
mapa={}
for char in string:
if char in mapa:
mapa[char]+=1
else:
mapa[char]=1
return mapa
mapa_b=mapear(string_b)
mapa_b
mapa_c={}
# +
for key,value in mapa_a.items():
if key in mapa_b.keys():
if value==1:
mapa_c[key]=1
else:
mapa_c[key]=value-mapa_b[key]
# +
for key,value in mapa_a.items():
if key in mapa_b.keys():
mapa_c[key]=value
# -
mapa_c
len(mapa_c)
len(mapa_a)
len(mapa_b)
resultado=(len(mapa_a)-len(mapa_c))+(len(mapa_b)-len(mapa_c))
resultado
mapa_a.items()
mapa_a['c']
| Udacity/.ipynb_checkpoints/hackerRank-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (nlp-master-thesis)
# language: python
# name: nlp-master-thesis
# ---
# +
#import tensorflow as tf
#import tensorflow.contrib.eager as tfe
import numpy as np
import pandas as pd
import pickle
from timeit import default_timer as timer
import matplotlib.pyplot as plt
import sklearn
from sklearn.model_selection import train_test_split
import nltk as nltk
import re
import os
# -
# ### Actions performed
# #### Prototype how to load the wall street journal data.
# #### Evaluation methods to compare trees
# #### comparison with google word vectors and convert words which are not there to UNK or some other symbol
# #### Filters sentences which are more than MAX_WORDS_COUNT words. Made the sentence equal to length MAX_WORDS_COUNT by adding < /fill > word
# # Load training and test dataset
# #### Use BracketParse corpus reader to load the wall street journal data
# #### functions subtrees(), label(), leaves() are important
# #### checkout load-penn-treebank-dataset.py for protoypes
dirname = os.getcwd()
dirname = os.path.dirname(dirname)
dataset_path = os.path.join(dirname, 'datasets/')
print(dataset_path)
# +
## https://www.nltk.org/_modules/nltk/tree.html
## http://www.nltk.org/howto/tree.html
## above link contains the API and also some tutorials
#reader = nltk.corpus.BracketParseCorpusReader('.','SWB-all-sentences-original-with-punctuation.MRG')
reader = nltk.corpus.BracketParseCorpusReader(dataset_path,'WSJ.txt')
print(reader.fileids())
print(type(reader))
## reads the file and converts each line into a tree
all_trees = reader.parsed_sents()
print('No. of trees: ', len(all_trees))
print(type(all_trees[0]))
# +
def get_frequency(trees):
frequency_dict = {}
for tree in trees:
words = tree.leaves()
words = [word.lower() for word in words]
for word in words:
if word not in frequency_dict.keys():
frequency_dict[word] = 0
frequency_dict[word] = frequency_dict[word] + 1
return frequency_dict
def filter_trees(trees, threshold_value, frequency_dict):
filtered_trees = []
if(frequency_dict is None):
frequency_dict = get_frequency(trees)
print('No.of words: ',len(frequency_dict))
keys = list(frequency_dict.keys())
for key in keys:
if(frequency_dict[key] <= threshold_value) :
frequency_dict.pop(key, None)
print('No. of words whose frequency is more than ',threshold_value,' is:', len(frequency_dict))
print('No. of trees:',len(trees))
filtered_trees = []
for tree in trees:
words = tree.leaves()
words = [word.lower() for word in words]
isFrequent = True
for word in words:
if word not in frequency_dict.keys():
isFrequent = False
break
if(isFrequent):
filtered_trees.append(tree)
print('No. of filtered trees:',len(filtered_trees))
return filtered_trees
FREQUENCY_THRESHOLD_VALUE = 2
frequency_dict = get_frequency(all_trees)
trees = filter_trees(all_trees, FREQUENCY_THRESHOLD_VALUE, frequency_dict)
# -
# ### Evaluation methods to compare trees
# #### equality operator (==) is good enough to compare trees. When comparing both the tree and tree label is taken into account
# #### Initial methods will not have categorization classification technique. So, it is important to replace all the values with say UNK symbol of google corpus
# #### It is created in such a way that subcategories of punctuations are named by itself
# ## Preprocess Tree to chomsky normal form, collapse unary and make it UNLabelled
# +
NO_LABEL = 'NO_LABEL'
def set_no_label(tree):
if(type(tree[0]) == type('a string')):
# we have reached the bottom most parent
tree.set_label(NO_LABEL)
return
# accesses the left child
set_no_label(tree[0])
# accesses the right child
set_no_label(tree[1])
tree.set_label(NO_LABEL)
def preprocess_tree(tree):
# set chomsky normal form
tree.chomsky_normal_form()
# merge the unary branches
tree.collapse_unary(collapsePOS=True, collapseRoot=True)
# set the label with 'NO_LABEL'
set_no_label(tree)
#temp_trees = [preprocess_tree(tree) for tree in trees]
#temp_trees[0]
tree = trees[10]
preprocess_tree(tree)
tree
# +
preprocessed_trees = []
for i in range(len(trees)):
tree = trees[i]
preprocess_tree(tree)
preprocessed_trees.append(tree)
preprocessed_trees[1000]
# -
# ### Convert the words which does not appear in Google word embeddings with UNK or other symbol
# As suggested in 'check-embeddings-wallstreet-coverage' google word embeddings are better than glove embeddings
# +
UNK = '</s>'
FILLER = '</fill>'
outfile = dataset_path +'google_word_corpus.pic'
with open(outfile, 'rb') as pickle_file:
googleCorpus, google_corpus_word_to_int, google_corpus_int_to_word = pickle.load(pickle_file)
#googleSet = pd.read_csv(dataset_path+'GoogleNews-vectors-negative10.txt', sep=' ', header=None)
#print(googleSet.shape)
#print(googleSet.head())
# TO BE DONE - Appropriate value should be added for FILLER. Right now it is a range from 1 to STATE_SIZE
#fill_data = {0:FILLER}
#for i in range(1, googleSet.shape[1]):
# fill_data[i] = i
#googleSet = googleSet.append(fill_data, ignore_index=True)
#print(googleSet.shape)
#print(googleSet.head())
#temp = list(googleCorpus)
#temp.append(FILLER)
#googleCorpus = set(temp)
#corpus_length = len(google_corpus_word_to_int)
#google_corpus_word_to_int[FILLER] = corpus_length
#google_corpus_int_to_word[corpus_length] = FILLER
#googleWords = googleSet.iloc[:,0:1]
#googleVectors = googleSet.iloc[:,1:]
# +
UNK = '</s>' # this is the symbol for UNK in google corpus
def set_UNK(tree):
if(type(tree[0]) == type('a string')):
# we have reached the bottom most parent
tree[0] = tree[0].lower()
if(tree[0] not in googleCorpus):
tree[0] = UNK
return
# accesses the left child
set_UNK(tree[0])
# accesses the right child
set_UNK(tree[1])
#temp_trees = [preprocess_tree(tree) for tree in trees]
#temp_trees[0]
tree = preprocessed_trees[0]
set_UNK(tree)
tree
# +
preprocessed_unk_trees = []
for i in range(len(preprocessed_trees)):
tree = preprocessed_trees[i]
set_UNK(tree)
preprocessed_unk_trees.append(tree)
preprocessed_unk_trees[1000]
# +
def UNK_count_analysis(tree):
words = tree.leaves()
words_count = len(words)
unk_count = 0
for word in words:
if word == UNK:
unk_count = unk_count + 1
#print(unk_count, words_count)
return unk_count / words_count
display(preprocessed_unk_trees[0])
UNK_count_analysis(preprocessed_unk_trees[0])
# -
THRESHOLD_PERCENTAGE = 0.25
preprocessed_unk_lt25per_trees = []
print(len(preprocessed_unk_trees))
for i in range(len(preprocessed_unk_trees)):
tree = preprocessed_unk_trees[i]
unk_percentage = UNK_count_analysis(tree)
if(unk_percentage <= THRESHOLD_PERCENTAGE):
preprocessed_unk_lt25per_trees.append(tree)
print(len(preprocessed_unk_lt25per_trees))
# ### Only sentences with <= MAX_WORDS_COUNT words (<MAX_WORDS_COUNT words suffix additional words and make it MAX_WORDS_COUNT)
# +
FILLER = '</fill>'
MAX_WORDS_COUNT = 10
preprocessed_unk_max_words_trees = []
for i in range(len(preprocessed_unk_lt25per_trees)):
tree = preprocessed_unk_lt25per_trees[i]
words_count = len(tree.leaves())
if(words_count == MAX_WORDS_COUNT): # it has been changed strictly from <= to ==
for i in range(MAX_WORDS_COUNT - words_count):
tree = nltk.Tree(NO_LABEL, [tree, nltk.Tree(NO_LABEL, [FILLER])])
preprocessed_unk_max_words_trees.append(tree)
print(len(preprocessed_unk_lt25per_trees))
print(len(preprocessed_unk_max_words_trees))
preprocessed_unk_max_words_trees[60] #123
# -
# +
def convert_golden_tree(tree, leaves, input_list):
if(type(tree[0]) == type('some string')):
#print(tree[0])
leaveIndex = leaves.index(tree[0])
leaves[leaveIndex]=''
return leaveIndex
leftIndex = convert_golden_tree(tree[0], leaves, input_list)
rightIndex = convert_golden_tree(tree[1], leaves, input_list)
#print(leftIndex, rightIndex)
#print(leftIndex)
input_list.append(leftIndex)
#return len(leaves) + len(input_list)
return leftIndex
tree_list =[]
tree_leaves = preprocessed_unk_max_words_trees[0].leaves()
convert_golden_tree(preprocessed_unk_max_words_trees[0], tree_leaves, tree_list)
tree_list
# +
golden_trees_combination = []
for i in range(len(preprocessed_unk_max_words_trees)):
tree = preprocessed_unk_max_words_trees[i]
temp_combination = []
tree_leaves = tree.leaves()
convert_golden_tree(tree, tree_leaves, temp_combination)
golden_trees_combination.append(temp_combination)
print(len(golden_trees_combination))
# -
# +
# Serialize it as CSV
treesDict = [{'sentence': ' '.join(tree.leaves()), 'tree':tree} for tree in preprocessed_unk_max_words_trees]
treeDataframe = pd.DataFrame(data=treesDict, columns=['sentence', 'tree'])
treeDataframe.head()
# -
treeDataframe.to_csv(dataset_path+'constituency-parsing-data-'+str(MAX_WORDS_COUNT)+'-UNK-NOLABEL.csv', sep=' ', index=False, header=False)
load_treeData = pd.read_csv(dataset_path+'constituency-parsing-data-'+str(MAX_WORDS_COUNT)+'-UNK-NOLABEL.csv', sep=' ', header=None, )
load_treeData.columns =['sentence', 'tree']
load_treeData.head()
print(load_treeData.iat[0,1])
print(type(load_treeData.iat[0,1]))
load_treeData['tree'] = load_treeData['tree'].apply(nltk.Tree.fromstring)
print(load_treeData.iat[0,1])
print(type(load_treeData.iat[0,1]))
load_treeData['length'] = load_treeData['sentence'].apply(lambda x: len(x.split()))
load_treeData.sort_values(by='length',ascending=True, inplace=True)
load_treeData.head()
| pre-processing/wallstreet-journal-data-preprocessing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from math import sqrt
from sklearn.metrics import mean_squared_error
from pandas.tools.plotting import autocorrelation_plot
from statsmodels.tsa.api import ExponentialSmoothing, ARIMA, ARMA, arma_order_select_ic
from statsmodels.tsa.statespace.sarimax import SARIMAX
from statsmodels.graphics.tsaplots import plot_acf, plot_pacf
from statsmodels.tsa.stattools import adfuller
from statsmodels.stats.diagnostic import acorr_ljungbox
# %matplotlib inline
# -
# 设置图片参数
plt.rcParams['font.sans-serif'] = ['SimHei'] # 用来正常显示中文
plt.rcParams['axes.unicode_minus'] = False # 用来正常显示负号
plt.rcParams['figure.figsize'] = (15, 8) # 图片大小
plt.rcParams['savefig.dpi'] = 300 # 图片像素
# plt.rcParams['figure.dpi'] = 300 # 分辨率
# ## 1 数据预处理
# 加载数据
data_path = "../data/engine_oil.txt"
df = pd.read_csv(data_path, sep='\t', encoding='utf-8')
# +
df = df.rename(columns={'物料编码': 'part_id', '物料描述': 'part_name',
'订货数': 'order_num', '缺件数': 'out_of_stock_num',
'受理数': 'delivery_num', '审核日期': 'date', '审核时间': 'time'})
# 将`part_id`的数据类型设为字符串,方便后面进行重采样
df['part_id'] = df['part_id'].astype('str')
# 重置索引
df['date'] = pd.to_datetime(df['date'], format="%Y-%m-%d")
df.set_index('date', inplace=True)
# -
# 去除2016年10月份的数据
df = df[3:]
# 重采样
df_week = df.resample('W').sum()
train_week = df_week[:-12]
test_week = df_week[-12:]
# +
fig, ax = plt.subplots()
ax.plot(df_week.index, df_week.order_num.values, 'bo-')
ax.set_xlabel('日期(周)', fontsize=16)
ax.set_ylabel('销量(件)', fontsize=16)
ax.set_title('机油每周订货量', fontsize=20)
ax.grid()
# TODO: 改变刻度字体大小不够优雅
plt.xticks(fontsize=12)
plt.yticks(fontsize=12)
plt.show()
# -
# ## 2 时间序列检验
# ### 2.1 自相关图和偏自相关图
# 周的自相关图
plot_acf(df_week.order_num.values).show()
plot_pacf(df_week.order_num.values).show()
autocorrelation_plot(df_week.order_num.values)
plt.show()
# ### 2.2 单位根检验
# 月的单位根检验
adf_result = adfuller(df_week.order_num.values)
output = pd.DataFrame(index=["Test Statistic Value", "p-value", "Lags Used", "Number of Observations Used", "Critical Value (1%)", "Critical Value (5%)", "Critical Value (10%)"], columns=['value'])
output['value']['Test Statistic Value'] = adf_result[0]
output['value']['p-value'] = adf_result[1]
output['value']['Lags Used'] = adf_result[2]
output['value']['Number of Observations Used'] = adf_result[3]
output['value']['Critical Value (1%)'] = adf_result[4]['1%']
output['value']['Critical Value (5%)'] = adf_result[4]['5%']
output['value']['Critical Value (10%)'] = adf_result[4]['10%']
print("月的单位根检验结果为:")
print(output)
# ### 2.3 纯随机性检验
# 月的纯随机性检验
print("序列的纯随机性检验结果为:")
print(acorr_ljungbox(df_week.order_num.values, lags=1)[1][0])
# ## 3 模型拟合
# ### 3.1 Holt-Winters method
train, test = train_week.order_num.values, test_week.order_num.values
model = ExponentialSmoothing(train, seasonal_periods=7, trend='add', seasonal='add')
model_fit = model.fit()
preds = model_fit.forecast(12)
# +
fig, ax = plt.subplots()
# ax.plot(train_month.index, train_month.order_num.values, 'go-', label='Train')
ax.plot(test_week.index,test, 'bo-', label='Test')
ax.plot(test_week.index, preds, 'ro-', label='Holt-Winters method')
ax.set_xlabel('日期(周)', fontsize=16)
ax.set_ylabel('销量(件)', fontsize=16)
ax.set_title('Holt-Winters方法预测机油每周订货量', fontsize=20)
ax.legend(loc='best')
ax.grid()
# TODO: 改变刻度字体大小不够优雅
plt.xticks(fontsize=12)
plt.yticks(fontsize=12)
plt.show()
# -
rmse = sqrt(mean_squared_error(test, preds))
print("The RMSE of 'Holt-Winters method' is:", rmse)
# ### 3.2 ARIMA
train, test = train_week.order_num.values, test_week.order_num.values
arma_order_select_ic(train, max_ar=6, max_ma=4, ic='aic')['aic_min_order']
history = list(train)
preds = list()
for i in range(len(test)):
model = ARMA(history, order=(1, 1))
model_fit = model.fit(disp=0)
output = model_fit.forecast()
yhat = output[0]
preds.append(yhat)
obs = test[i]
history.append(obs)
print("expected = %f predicted = %f" % (obs, yhat))
error = sqrt(mean_squared_error(test, preds))
print("Test RMSE: %.3f" % error)
# +
fig, ax = plt.subplots()
ax.plot(test_week.index, test, 'bo-', label='test')
ax.plot(test_week.index, preds, 'ro-', label='pred')
ax.set_xlabel('日期(周)', fontsize=16)
ax.set_ylabel('销量(件)', fontsize=16)
ax.set_title('ARIMA Rolling Forecast Line Plot', fontsize=20)
ax.legend(loc='upper left')
ax.grid()
# TODO: 改变刻度字体大小不够优雅
plt.xticks(fontsize=12)
plt.yticks(fontsize=12)
fig.savefig("../figs/ARIMA预测图.png")
plt.show()
# -
# ### 3.3 SARIMAX
train, test = train_month.order_num.values, test_month.order_num.values
history = list(train)
preds = list()
for i in range(len(test)):
model = SARIMAX(train, order=(2, 0, 4), seasonal_order=(0, 1, 1, 7))
model_fit = model.fit(disp=0)
output = model_fit.forecast()
yhat = output[0]
preds.append(yhat)
obs = test[i]
history.append(obs)
print("expected = %f predicted = %f" % (obs, yhat))
error = sqrt(mean_squared_error(test, preds))
print("Test RMSE: %.3f" % error)
| ipython/engine-oil/engine-oil-forecast-week2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Data preparation
# +
import sqlite3
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import Cdf
import Pmf
# define global plot parameters
params = {'axes.labelsize' : 12, 'axes.titlesize' : 12,
'font.size' : 12, 'legend.fontsize' : 10,
'xtick.labelsize' : 12, 'ytick.labelsize' : 12}
plt.rcParams.update(params)
plt.rcParams.update({'figure.max_open_warning': 0})
# connect to database and get data
conn = sqlite3.connect('data/youtube-traceroute.db')
# read necessary tables from database
date_cols = ['dtime']
devices = pd.read_sql_query('select distinct(unit_id) from traceroute', conn)
df_v6_medians = pd.read_sql_query('select * from path_medians_v6', conn)
df_v4_medians = pd.read_sql_query('select * from path_medians_v4', conn)
conn.close()
# -
# ## CDF over TTL and RTT medians of paths
# +
### TTL
# setup plot
ttl_cdf_fig, ttl_cdf_ax = plt.subplots(figsize = (5, 2))
ttl_cdf_v6 = Cdf.MakeCdfFromList(df_v6_medians['median(ttl)'])
ttl_cdf_v4 = Cdf.MakeCdfFromList(df_v4_medians['median(ttl)'])
ttl_cdf_ax.plot(ttl_cdf_v6.xs, ttl_cdf_v6.ps,
label = 'IPv6',
marker = 's',
linewidth = 0.5,
markersize = 5,
fillstyle = 'none',
color = 'red')
ttl_cdf_ax.plot(ttl_cdf_v4.xs, ttl_cdf_v4.ps,
label = 'IPv4',
marker = '^',
linewidth = 0.5,
markersize = 5,
fillstyle = 'none',
color = 'blue')
# PLOT FORMATTING
# customize axes and grid appearance
ttl_cdf_ax.legend(loc = 'lower right', fontsize = 'smaller')
ttl_cdf_ax.set_xlabel('TTL')
ttl_cdf_ax.set_ylabel('CDF')
ttl_cdf_ax.set_xlim([0, 25])
ttl_cdf_ax.set_ylim([-0.05, 1.05])
yticks = np.arange(0.0, 1.01, 0.2)
ttl_cdf_ax.set_yticks(yticks)
ttl_cdf_ax.grid(False)
ttl_cdf_ax.spines['right'].set_color('none')
ttl_cdf_ax.spines['top'].set_color('none')
ttl_cdf_ax.yaxis.set_ticks_position('left')
ttl_cdf_ax.xaxis.set_ticks_position('bottom')
ttl_cdf_ax.spines['bottom'].set_position(('axes', -0.03))
ttl_cdf_ax.spines['left'].set_position(('axes', -0.03))
### RTT
# setup plot
rtt_cdf_fig, rtt_cdf_ax = plt.subplots(figsize = (5, 2))
rtt_cdf_v6 = Cdf.MakeCdfFromList(df_v6_medians['median(rtt)'])
rtt_cdf_v4 = Cdf.MakeCdfFromList(df_v4_medians['median(rtt)'])
rtt_cdf_ax.plot(rtt_cdf_v6.xs, rtt_cdf_v6.ps,
label = 'IPv6',
marker = 's',
linewidth = 0.5,
markersize = 5,
fillstyle = 'none',
color = 'red')
rtt_cdf_ax.plot(rtt_cdf_v4.xs, rtt_cdf_v4.ps,
label = 'IPv4',
marker = '^',
linewidth = 0.5,
markersize = 5,
fillstyle = 'none',
color = 'blue')
# PLOT FORMATTING
# customize axes and grid appearance
rtt_cdf_ax.legend(loc = 'lower right', fontsize = 'smaller')
rtt_cdf_ax.set_xlabel('RTT [ms]')
rtt_cdf_ax.set_xscale('log')
from matplotlib.ticker import ScalarFormatter
rtt_cdf_ax.xaxis.set_major_formatter(ScalarFormatter())
rtt_cdf_ax.set_ylabel('CDF')
rtt_cdf_ax.set_ylim([-0.05, 1.05])
rtt_cdf_ax.minorticks_off()
rtt_cdf_ax.set_yticks(yticks)
rtt_cdf_ax.grid(False)
rtt_cdf_ax.spines['right'].set_color('none')
rtt_cdf_ax.spines['top'].set_color('none')
rtt_cdf_ax.yaxis.set_ticks_position('left')
rtt_cdf_ax.xaxis.set_ticks_position('bottom')
rtt_cdf_ax.spines['bottom'].set_position(('axes', -0.03))
rtt_cdf_ax.spines['left'].set_position(('axes', -0.03))
# saving and showing plot
#ttl_cdf_fig.savefig('plots/cdfs_median_ttl.pdf', bbox_inches = 'tight')
#rtt_cdf_fig.savefig('plots/cdfs_median_rtt.pdf', bbox_inches = 'tight')
plt.show()
# -
# ### Both in one figure
# +
# setup plot
cdf_fig, cdf_ax = plt.subplots(figsize = (5, 2))
rtt_cdf_v6 = Cdf.MakeCdfFromList(df_v6_medians['median(rtt)'])
rtt_cdf_v4 = Cdf.MakeCdfFromList(df_v4_medians['median(rtt)'])
cdf_ax.plot(rtt_cdf_v6.xs, rtt_cdf_v6.ps,
label = 'RTT (IPv6) [ms]',
marker = 'o',
linewidth = 0.5,
markersize = 2.5,
color = 'red')
cdf_ax.plot(rtt_cdf_v4.xs, rtt_cdf_v4.ps,
label = 'RTT (IPv4) [ms]',
marker = 'v',
linewidth = 0.5,
markersize = 2.5,
color = 'blue')
ttl_cdf_v6 = Cdf.MakeCdfFromList(df_v6_medians['median(ttl)'])
ttl_cdf_v4 = Cdf.MakeCdfFromList(df_v4_medians['median(ttl)'])
cdf_ax.plot(ttl_cdf_v6.xs, ttl_cdf_v6.ps,
label = 'TTL (IPv6)',
marker = 's',
linewidth = 0.5,
markersize = 2.5,
markeredgewidth = 1.5,
color = 'orange')
cdf_ax.plot(ttl_cdf_v4.xs, ttl_cdf_v4.ps,
label = 'TTL (IPv4)',
marker = '^',
linewidth = 0.5,
markersize = 2.5,
markeredgewidth = 1.5,
color = 'cyan')
# PLOT FORMATTING
# customize axes and grid appearance
yticks = np.arange(0.0, 1.01, 0.2)
cdf_ax.set_yticks(yticks)
cdf_ax.grid(False)
cdf_ax.spines['right'].set_color('none')
cdf_ax.spines['top'].set_color('none')
cdf_ax.yaxis.set_ticks_position('left')
cdf_ax.xaxis.set_ticks_position('bottom')
cdf_ax.spines['bottom'].set_position(('axes', -0.03))
cdf_ax.spines['left'].set_position(('axes', -0.03))
cdf_ax.legend(loc = 'upper left', fontsize = 'smaller')
cdf_ax.set_ylabel('CDF')
cdf_ax.set_xlim([1, 100])
cdf_ax.set_xscale('log')
cdf_ax.set_ylim([-0.05, 1.05])
from matplotlib.ticker import EngFormatter
cdf_ax.xaxis.set_major_formatter(EngFormatter())
# saving and showing plot
cdf_fig.savefig('plots/cdfs_median_both.pdf', bbox_inches = 'tight')
plt.show()
# -
# ## Distributions of median metrics
print('-- IPv4 --')
print('m_ttl; cdf')
print('----------')
for x, y in zip(ttl_cdf_v4.xs, ttl_cdf_v4.ps):
print('%.2f; %.5f' % (x, y))
print('-- IPv6 --')
print('m_ttl; cdf')
print('----------')
for x, y in zip(ttl_cdf_v6.xs, ttl_cdf_v6.ps):
print('%.2f; %.5f' % (x, y))
print('-- IPv4 --')
print('m_rtt; cdf')
print('----------')
for x, y in list(zip(rtt_cdf_v4.xs, rtt_cdf_v4.ps))[0::100]:
print('%.2f; %.5f' % (x, y))
print('-- IPv6 --')
print('m_rtt; cdf')
print('----------')
for x, y in list(zip(rtt_cdf_v6.xs, rtt_cdf_v6.ps))[0::100]:
print('%.2f; %.5f' % (x, y))
| nb-paths.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6 - AzureML
# language: python
# name: python3-azureml
# ---
# Copyright (c) Microsoft Corporation. All rights reserved.
#
# Licensed under the MIT License.
# # Model deployment to ACI
#
# We finished the last Notebook by finding best fitting model using automated ML and registering it to our Azure ML account. In this Notebook, we deploy this model to an ACI instance and test it by scoring data against it. Scoring here happens in near-realtime, meaning that the data we score is pre-computed for us (such as a nightly batch job). Scoring can also happen in realtime, but as we will explore in a later Notebook, this requires more work. For predictive maintenance, realtime scoring is usually not needed, because models are used to predict when a machine is going to *about to* fail, which gives us some time to run unscheduled maintenance and replace parts.
# ## Create Experiment
# As part of the setup we have already created an AML workspace. Let's load the workspace and create an experiment.
# +
import json
import logging
import os
import random
from matplotlib import pyplot as plt
from matplotlib.pyplot import imshow
import numpy as np
import pandas as pd
from sklearn import datasets
import azureml.core
from azureml.core.experiment import Experiment
from azureml.core.workspace import Workspace
from azureml.train.automl import AutoMLConfig
from azureml.train.automl.run import AutoMLRun
# -
# We load the workspace directly from the config file we created in the early part of the course.
# +
config_path = '../lab02.0_PdM_Setting_Up_Env/aml_config/config.json'
ws = Workspace.from_config(config_path)
experiment_name = 'pred-maint-automl' # choose a name for experiment
project_folder = '.' # project folder
experiment=Experiment(ws, experiment_name)
output = {}
output['SDK version'] = azureml.core.VERSION
output['Subscription ID'] = ws.subscription_id
output['Workspace'] = ws.name
output['Resource Group'] = ws.resource_group
output['Location'] = ws.location
output['Project Directory'] = project_folder
output['Experiment Name'] = experiment.name
pd.set_option('display.max_colwidth', -1)
pd.DataFrame(data=output, index=['']).T
# +
import azureml.core
print("SDK Version:", azureml.core.VERSION)
# -
# Next we load the test data, not to evaluate the model but only to use it to get some predictions from our deployed model.
# %store -r X_test
# %store -r y_test
# ## Create a scoring script
# The first part of the deployment consists of pointing to the model we want to deploy. We can simply provide the model name, which was given to us at the time we registered the model. We can also go to the Azure portal to look up the model name.
# Here's a quick sanity check to ensure that the model exists and can be loaded (loading the model in the current session is not required for deployment).
#
# **Note**: You have to updated the `model_name` below. You can find the model name in your workspace in the azure portal.
# +
from azureml.core.model import Model
model_name = "AutoML4b8dd2f49best"
model = Model(workspace=ws, name=model_name)
print(model.id)
# -
# We now create a scoring script that will run every time we make a call to the deployed model. The scoring script consists of an `init` function that will load the model and a `run` function that will load the data we provide at score time and use the model to obtain predictions.
#
# **Note**: You have to updated the model_name below. You can find the model name in your workspace in the azure portal.
# +
# %%writefile score.py
import pickle
import json
import numpy
from sklearn.externals import joblib
from azureml.core.model import Model
import azureml.train.automl
model_name = "AutoML4b8dd2f49best""
def init():
global model
model_path = Model.get_model_path(model_name = model_name) # this name is modeld.id of model that we want to deploy
# model_path = Model.get_model_path('model.pkl') # select this if deploying model from file
# deserialize the model file back into a sklearn model
model = joblib.load(model_path)
def run(rawdata):
try:
data = json.loads(rawdata)['data']
data = numpy.array(data)
result = model.predict(data)
except Exception as e:
result = str(e)
return json.dumps({"error": result})
return json.dumps({"result":result.tolist()})
# -
# ## Create a conda environment file
# We begin by retrieving the run ID for the automl experiment we ran in the last Notebook and pasting it in for the `run_id` argument in the `AutoMLRun` function below.
#
# **Note**: You have to updated the run_id below. You can find the run_id under the experiments in the azure portal.
ml_run = AutoMLRun(experiment=experiment, run_id='AutoML_4b8dd2f4-9a88-4a90-b427-9c45aee54287')
# Next we create a `yml` file for the conda environment that will be used to run the scoring script above. To ensure consistency of the scored results with the training results, the SDK dependencies need to mirror development environment (used for model training).
dependencies = ml_run.get_run_sdk_dependencies(iteration=9)
dependencies
for p in ['azureml-train-automl', 'azureml-sdk', 'azureml-core']:
print('{}\t{}'.format(p, dependencies[p]))
# +
from azureml.core.conda_dependencies import CondaDependencies
myenv = CondaDependencies.create(conda_packages=['scikit-learn'])
# myenv.add_pip_package("azureml-train")
# myenv.add_pip_package("azureml-train-automl")
# myenv.add_pip_package("azureml-core")
with open("myenv.yml","w") as f:
f.write(myenv.serialize_to_string())
# -
# %cat myenv.yml
import sklearn
sklearn.__version__
# %%writefile myenv.yml
name: myenv
channels:
- defaults
dependencies:
- pip:
- scikit-learn==0.19.1
- azureml-sdk[automl]==0.1.74
# %%writefile myenv.yml
name: myenv
dependencies:
- scikit-learn==0.19.1
- pip:
- azureml-defaults
- azureml-train-automl
# %cat myenv.yml
# ## Create a docker image
# Using the scoring script and conda environment file, we can now create a docker image that will host the scoring script and a Python executable that meets the conda requirement dependencies laid out in the YAML file.
# +
from azureml.core.image import Image, ContainerImage
image_config = ContainerImage.image_configuration(runtime= "python",
execution_script="score.py",
conda_file="myenv.yml",
tags = {'area': "digits", 'type': "automl_classification"},
description = "Image for automl classification sample")
# +
# %%time
image_name = experiment_name + "-img"
image = Image.create(name = image_name,
models = [model],
image_config = image_config,
workspace = ws)
image.wait_for_creation(show_output = True)
# -
# If the image creation fails, this is how we can access the log file and examine what went wrong.
print(image.image_build_log_uri)
# This is the image location that will be used when the imaged is pulled down from Docker hub.
print(image.image_location)
# Note that if the image was created in another session and we just wanted to point to it in this session, then we can just pass the image name and workspace to the `Image` function as follows:
# image = Image(name = experiment_name + "-img", workspace = ws, version=29)
print(image.image_location)
# ## Deploy Image as web service on ACI
# We are now ready to deploy our image as a web service on ACI. To do so, we first create a config file and then pass it to `deploy_from_image` along with a name for the service, the image we created in the last step, and our workspace.
# +
from azureml.core.webservice import AciWebservice
aciconfig = AciWebservice.deploy_configuration(cpu_cores=1,
memory_gb=1,
tags={"method" : "automl"},
description='Predictive maintenance using auto-ml')
# -
# If a service with the same name already exists, we can delete it by calling the `delete` method.
# +
# %%time
from azureml.core.webservice import Webservice
aci_service_name = experiment_name + "-aci"
print(aci_service_name)
# aci_service.delete()
aci_service = Webservice.deploy_from_image(deployment_config = aciconfig,
image = image,
name = aci_service_name,
workspace = ws)
aci_service.wait_for_deployment(True)
print(aci_service.state)
# -
# ## Debugging issues
# Here's how we can get logs from the deployed service, which can help us debug any issues that cause the deployment to fail.
logs = aci_service.get_logs()
# +
import re
import json
from pprint import pprint
ll = re.findall(r"\{.*\}", logs)
dd = [json.loads(l) for l in ll]
for k in dd:
if 'level' in k.keys():
if k['level'] == 'ERROR':
print('================================================================================')
# pprint(json.loads(k['message']))
print(json.loads(k['message'])['message'])
# -
# ## Alternative deployments (optional)
# There are two other ways that we could have launched our ACI deployment. The first one is by deploying directly from the image config file and the registered model. In this scenario, the deployment will first create the image from the registered model, and then deploy the docker container from the base image. So we combine two steps (image creation, service creation) into a single step. However, behind the scenes the steps still run individually and create corresponding resources.
# +
# aci_service.delete()
# +
# %%time
from azureml.core.webservice import Webservice
aci_service_name = experiment_name + "-aci"
print(aci_service_name)
# aci_service = Webservice.deploy_from_model(deployment_config = aciconfig,
# image_config = image_config,
# models = [model], # this is the registered model object
# name = aci_service_name,
# workspace = ws)
# aci_service.wait_for_deployment(show_output = True)
# print(aci_service.state)
# -
# In the above example, we launched the ACI docker container from the image config file and the registered model. But we can take one further step back and simply provide the model pickle file and let it register the model, then create a docker image from it and finally launch an ACI docker container from the image. In this case, we are combining three steps into one.
# +
# aci_service.delete()
# +
from azureml.core.webservice import Webservice
aci_service_name = experiment_name + "-aci"
print(aci_service_name)
# aci_service = Webservice.deploy(deployment_config = aciconfig,
# image_config = image_config,
# model_paths = ['model.pkl'],
# name = aci_service_name,
# workspace = ws)
# aci_service.wait_for_deployment(show_output = True)
# print(aci_service.state)
# -
print(aci_service.scoring_uri)
# Combining many steps into one may save us a few lines of code, but it has the disadvantage of appearing to over-simplify the workflow. So it is probably best to avoid doing it, especially for production systems.
# ## Test Web Service
# It is time to test our web service. To begin with, we will point to our service using `Webservice`. Note that we've already done this in the last step, so in the current session this is not a necessary step, but since we want to be able to test the service from any Python session, we will point to the service again here. There is next to no overhead in doing so.
# +
from azureml.core.image import Image, ContainerImage
from azureml.core.webservice import Webservice
aci_service_name = experiment_name + "-aci"
# image = Image(name = experiment_name + "-img", workspace = ws)
aci_service = Webservice(workspace = ws, name = aci_service_name)
# -
# We can now proceed to testing the service. To do so, we will take a few random samples from `X_test` and dump its content into a json string (with UTF-8 encoding). This will act as the data that we intend to score. We can pass this data to the service using the `run` method, and it will return the predictions to us.
# +
n = 5
sample_indices = np.random.permutation(X_test.shape[0])[0:n]
test_samples = json.dumps({"data": X_test.iloc[sample_indices, :].values.tolist()})
test_samples = bytes(test_samples, encoding = 'utf8')
print(test_samples)
# predict using the deployed model
prediction = aci_service.run(input_data = test_samples)
print('**********************************************')
print(prediction)
# -
# ### Lab
# In the above example, we took the data to be scored directly from `X_test`. But this data had already been pre-processed for us and was ready for scoring. A more realistic scenario involves getting raw data, pre-processing it and then feeding it to the deployed model for scoring. In this lab, we will implement this.
# +
# generate the telemetry data
# run anomaly detection on it (invoke score_AD.py to get errors)
# compute moving average telemetries (use the function in score_AD.py for computing MAs at test time)
# append maintenance history and failure history to it
# now you're ready to score
# -
# ### End of lab
# # The end
# Copyright (c) Microsoft Corporation. All rights reserved.
#
# Licensed under the MIT License.
| lab02.3_PdM_Model_Deployment/deploy_PdM_model_with_AML.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SageMath 9.4
# language: sage
# name: sagemath-9.4
# ---
# +
# Turkalj dim 5 rational isometry classes (we need commensurability only)
# -
Turkalj_dim_5 = [[ 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, -1 ],
[ 0, 0, 0, 0, 0, -2, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, -2, 0, 0, 0, 0, -4 ],
[ 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, -1, 0, 0, 0, 0, 0, 0, 7 ],
[ 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, -1, 0, 0, 0, 0, 0, 0, 4, -2, 0, 0, 0, 0, -2, 8 ],
[ 0, -1, -1, -3, 2, -6, -1, 0, 0, 1, -1, -3, -1, 0, 2, -1, 0, 0, -3, 1, -1, 12, -3, -4, 2, -1, 0, -3, 12, -4, -6, -3, 0, -4, -4, -20 ],
[ 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, -2, 1, 0, 0, 0, 0, 1, 2 ],
[ 0, -1, 0, 0, 1, 0, -1, -2, 0, 0, 1, -1, 0, 0, 2, -1, 1, -1, 0, 0, -1, 2, -1, 1, 1, 1, 1, -1, 2, 0, 0, -1, -1, 1, 0, 2 ],
[ 0, -1, -1, -1, -2, 1, -1, 0, -1, -1, -2, 1, -1, -1, 0, -1, -2, 1, -1, -1, -1, 0, -2, 1, -2, -2, -2, -2, -4, 2, 1, 1, 1, 1, 2, 4 ],
[ 0, 2, 2, -1, 2, -6, 2, 0, -3, -1, 5, 12, 2, -3, 0, -1, -2, 19, -1, -1, -1, 2, 0, 0, 2, 5, -2, 0, 26, 10, -6, 12, 19, 0, 10, -58 ],
[ 0, -4, 1, -5, 2, 5, -4, -2, -1, 0, -1, 1, 1, -1, 4, 0, -2, 0, -5, 0, 0, 10, -5, 5, 2, -1, -2, -5, 10, -1, 5, 1, 0, 5, -1, 14 ],
[ 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, -5 ],
[ 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, -4, 2, 0, 0, 0, 0, 2, 4 ],
[ 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 5, 0, 0, 0, 0, 0, 0, 5, 0, 0, 0, 0, 0, 0, -5 ],
[ 0, -1, 0, 0, 0, -2, -1, 0, 0, 0, 1, -2, 0, 0, 2, -1, 1, 1, 0, 0, -1, 4, 1, 1, 0, 1, 1, 1, 4, 1, -2, -2, 1, 1, 1, -4 ],
[ 0, -1, 0, 1, -1, -5, -1, 2, -1, 0, -1, 0, 0, -1, 2, 0, 0, 0, 1, 0, 0, 2, 0, 0, -1, -1, 0, 0, 4, 0, -5, 0, 0, 0, 0, -10 ],
[ 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 2, 1, 0, 0, 0, 0, 1, 3, 0, 0, 0, 0, 0, 0, -5 ],
[ 0, 0, 0, 0, 0, -5, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, -5, 0, 0, 0, 0, -15 ],
[ 0, 0, 0, 0, 0, -10, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, -10, 0, 0, 0, 0, -80 ],
[ 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, -1, 0, 0, 0, 0, 0, 0, 2, 1, 0, 0, 0, 0, 1, 2 ],
[ 1, 0, 0, 0, 0, 0, 0, -1, 0, 0, 0, 0, 0, 0, 2, 0, -1, 1, 0, 0, 0, 2, -1, -1, 0, 0, -1, -1, 3, 1, 0, 0, 1, -1, 1, 3 ],
[ 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, -3, 0, 0, 0, 0, 0, 0, 3, 0, 0, 0, 0, 0, 0, 3 ],
[ 0, 0, 0, 0, 0, -6, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 3, 0, -6, 0, 0, 0, 0, -24 ],
[ 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, -1, 0, 0, 0, 0, 0, 0, 3 ],
[ 0, 0, 0, 0, 0, -2, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 3, 0, -2, 0, 0, 0, 0, -4 ],
[ 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 2, 1, 0, 0, 0, 0, 1, 2, 0, 0, 0, 0, 0, 0, -3, 0, 0, 0, 0, 0, 0, 3 ],
[ 1, 0, 0, 0, 0, 0, 0, -2, 0, -1, 1, 0, 0, 0, 2, 1, -1, 0, 0, -1, 1, 3, 0, 0, 0, 1, -1, 0, 3, 0, 0, 0, 0, 0, 0, 3 ],
[ 0, 0, 0, 0, 0, 1, 0, 2, 0, 0, -1, -1, 0, 0, 2, 1, 0, 1, 0, 0, 1, 2, 0, 0, 0, -1, 0, 0, 4, 1, 1, -1, 1, 0, 1, -6 ],
[ 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, -1, 0, 0, 0, 0, 0, 0, 15 ],
[ 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, -2, 1, 0, 0, 0, 0, 1, 2, 0, 0, 0, 0, 0, 0, 3 ],
[ 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, -2, 1, 0, 0, 0, 0, 1, 7 ],
[ 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, -1, 0, 0, 0, 0, 0, 0, 10, 5, 0, 0, 0, 0, 5, 10 ],
[ 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, -3, 0, 0, 0, 0, 0, 0, 3 ],
[ 0, 0, 0, 0, 0, -6, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, -6, 0, 0, 0, 0, -24 ],
[ 0, -1, 0, 0, -1, -2, -1, -2, -1, 1, 0, 0, 0, -1, 2, 0, 0, 0, 0, 1, 0, 2, 1, 1, -1, 0, 0, 1, 10, 2, -2, 0, 0, 1, 2, 12 ],
[ 0, 0, 0, 0, 0, 2, 0, 2, 0, 0, 1, 0, 0, 0, 2, 1, 0, 0, 0, 0, 1, 2, 0, 0, 0, 1, 0, 0, 2, 0, 2, 0, 0, 0, 0, -8 ],
[ 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, -1, 0, 0, 0, 0, 0, 0, 3, 0, 0, 0, 0, 0, 0, 3 ],
[ 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, -1, 0, 0, 0, 0, 0, 0, 3, 0, 0, 0, 0, 0, 0, 4, 2, 0, 0, 0, 0, 2, 4 ],
[ 0, -1, 0, 1, 0, -3, -1, 2, -1, -1, 0, -1, 0, -1, 2, 0, 0, 0, 1, -1, 0, 2, 0, 1, 0, 0, 0, 0, 2, -1, -3, -1, 0, 1, -1, -10 ],
[ 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, -2 ],
[ 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 0, 0, -2 ],
[ 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, -1, 0, 0, 0, 0, 0, 0, 14 ],
[ 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, -10 ],
[ 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, -1, 0, 0, 0, 0, 0, 0, 6 ],
[ 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, -1, 0, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 0, 0, 6 ],
[ 0, 0, 0, 0, 0, 2, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 0, 0, 3, 0, 2, 0, 0, 0, 0, -10 ],
[ 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, -3, 0, 0, 0, 0, 0, 0, 3, 0, 0, 0, 0, 0, 0, 6 ],
[ 0, 0, 0, 0, -6, 6, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, -6, 0, 0, 0, -18, 0, 6, 0, 0, 0, 0, 24 ],
[ 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 0, 0, 3, 0, 0, 0, 0, 0, 0, 6, 0, 0, 0, 0, 0, 0, -6 ],
[ 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, -6 ],
[ 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, -1, 0, 0, 0, 0, 0, 0, 3, 0, 0, 0, 0, 0, 0, 3, 0, 0, 0, 0, 0, 0, 6 ],
[ 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, -1, 0, 0, 0, 0, 0, 0, 3, 0, 0, 0, 0, 0, 0, 6 ],
[ 0, 0, 0, -2, 0, -4, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, -2, 0, 0, -4, 0, -2, 0, 0, 0, 0, 6, 0, -4, 0, 0, -2, 0, 11 ],
[ 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 3, 0, 0, 0, 0, 0, 0, -6 ],
[ 1, 0, 0, 0, 0, 0, 0, -1, 0, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 0, 0, 3, 0, 0, 0, 0, 0, 0, 4, 2, 0, 0, 0, 0, 2, 4 ],
[ 0, -1, -1, 1, 0, 2, -1, 0, -1, 1, 0, 2, -1, -1, 0, 1, 1, 3, 1, 1, 1, 0, -1, -3, 0, 0, 1, -1, 2, 0, 2, 2, 3, -3, 0, -6 ],
[ 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, -1, 0, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 0, 0, 2 ],
[ 0, 0, 0, -1, 2, -1, 0, 2, 0, -1, 0, 0, 0, 0, 2, -1, 0, 1, -1, -1, -1, 4, -1, -2, 2, 0, 0, -1, -6, 2, -1, 0, 1, -2, 2, 16 ],
[ 2, 1, 1, 1, 1, 1, 1, 2, 0, 0, 1, 1, 1, 0, 2, 0, 0, 0, 1, 0, 0, 2, 1, 1, 1, 1, 0, 1, -6, 1, 1, 1, 0, 1, 1, 8 ],
[ 0, -1, 2, 1, 0, -5, -1, 0, 2, 1, 0, -5, 2, 2, 0, 0, -3, 7, 1, 1, 0, 2, 0, 0, 0, 0, -3, 0, 4, -1, -5, -5, 7, 0, -1, -16 ],
[ 0, -1, -1, 2, 0, -5, -1, 2, 1, 0, 1, 0, -1, 1, 2, 0, 1, 0, 2, 0, 0, 4, -2, 0, 0, 1, 1, -2, 10, 0, -5, 0, 0, 0, 0, -10 ],
[ 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, -2, 0, 0, 1, 0, 0, 0, 2, 0, 0, 0, 0, 0, 0, 2, 0, 0, 0, 1, 0, 0, 2 ],
[ 2, 0, 0, 0, -1, 0, 0, 2, 0, 0, -1, 0, 0, 0, 2, 0, -1, 0, 0, 0, 0, -2, 0, 1, -1, -1, -1, 0, 4, 0, 0, 0, 0, 1, 0, 2 ],
[ 0, 1, -1, 1, 3, -5, 1, 0, 1, -1, 0, 4, -1, 1, 2, 1, 0, -1, 1, -1, 1, 2, 0, -1, 3, 0, 0, 0, 6, 0, -5, 4, -1, -1, 0, -6 ],
[ 2, 1, 0, 0, 0, 0, 1, 2, 0, 0, 0, 0, 0, 0, -2, 0, 0, 0, 0, 0, 0, 2, 1, 0, 0, 0, 0, 1, 2, 0, 0, 0, 0, 0, 0, 6 ],
[ 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, -1, 0, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 0, 0, 6 ],
[ 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, -3, 0, 0, 0, 0, 0, 0, 6, 0, 0, 0, 0, 0, 0, 6 ],
[ 0, -1, -1, -1, -2, 0, -1, 0, -1, -1, -2, 0, -1, -1, 0, -1, -2, 3, -1, -1, -1, 0, -2, 0, -2, -2, -2, -2, -4, 0, 0, 0, 3, 0, 0, 12 ],
[ 0, -3, 2, -1, 1, -6, -3, 0, 0, 0, 0, -6, 2, 0, 2, -1, 0, 0, -1, 0, -1, 2, 0, 0, 1, 0, 0, 0, 2, 0, -6, -6, 0, 0, 0, -12 ],
[ 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, -2, 0, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 0, 0, 3 ],
[ 0, 0, 0, -6, 12, 12, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, -6, 0, 0, 20, 9, 10, 12, 0, 0, 9, -42, 9, 12, 0, 0, 10, 9, 98 ],
[ 2, 1, -1, 1, 0, 0, 1, 2, -1, 0, 0, 0, -1, -1, 2, -1, 0, 0, 1, 0, -1, 2, 0, 0, 0, 0, 0, 0, 2, -1, 0, 0, 0, 0, -1, -10 ],
[ 0, 0, 0, -2, 1, -5, 0, 2, 1, 0, 0, 0, 0, 1, 2, 0, 0, 0, -2, 0, 0, 2, 0, 0, 1, 0, 0, 0, 4, 0, -5, 0, 0, 0, 0, -10 ],
[ 2, -1, -1, 1, 1, 0, -1, 2, 1, -1, -1, 0, -1, 1, 2, -1, -1, 0, 1, -1, -1, 2, 1, 0, 1, -1, -1, 1, 2, 0, 0, 0, 0, 0, 0, -10 ],
[ 0, 1, 0, 0, 0, 0, 1, -2, 0, 0, 1, 0, 0, 0, 2, 0, 0, 0, 0, 0, 0, 2, 0, 0, 0, 1, 0, 0, 2, 1, 0, 0, 0, 0, 1, 8 ],
[ 0, 3, 8, -3, 0, 3, 3, 0, -12, 1, -3, -4, 8, -12, 0, -1, -1, -1, -3, 1, -1, 2, 1, 0, 0, -3, -1, 1, 2, 0, 3, -4, -1, 0, 0, 6 ],
[ 0, 0, 0, 0, 0, 3, 0, 2, -1, -1, -1, 0, 0, -1, 2, 1, 1, 0, 0, -1, 1, 2, 0, 0, 0, -1, 1, 0, 2, 0, 3, 0, 0, 0, 0, -210 ],
[ 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, -6, 0, 0, 0, 0, 0, 0, 6 ],
[ 0, 0, 0, 0, -1, 0, 0, 2, 1, 0, 0, 0, 0, 1, 2, 0, 0, 0, 0, 0, 0, 2, -1, 0, -1, 0, 0, -1, -4, 0, 0, 0, 0, 0, 0, 6 ],
[ 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, -1, 0, 0, 0, 0, 0, 0, 6, 0, 0, 0, 0, 0, 0, 6 ],
[ 2, 0, 0, 0, 0, 1, 0, 2, 1, 0, 0, 0, 0, 1, 2, 0, 0, 0, 0, 0, 0, 2, 0, 1, 0, 0, 0, 0, 2, -1, 1, 0, 0, 1, -1, -6 ]]
V = [QuadraticForm(2*Matrix(QQ, 6, m)) for m in Turkalj_dim_5]
Q = Graph(len(V))
for i in Q.vertices():
Q.set_vertex(i, V[i])
for i in Q.vertices():
for j in Q.vertices():
if (i<j):
q1 = Q.get_vertex(i)
q2 = Q.get_vertex(j)
if q1.is_rationally_isometric(q2):
Q.add_edge(i,j)
ind = [sorted(c, key=lambda x: abs(Q.get_vertex(x).det()))[0] for c in Q.connected_components()]
for i in ind:
m = Turkalj_dim_5[i]
print("{},".format(m))
# +
# Scharlau-Walhorn dim 4 rational isometry classes (we need commensurability only)
# -
Scharlau_Walhorn_dim_4 = [[0, 1, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 0, 2],
[0, 1, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 0, 4, 2, 0, 0, 0, 2, 4],
[0, 1, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 0, 6],
[0, 2, 0, 0, 0, 2, 0, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 0, 2],
[0, 1, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 0, 10],
[0, 2, 0, 0, 0, 2, 0, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 0, 4, 2, 0, 0, 0, 2, 6],
[0, 2, 0, 0, 0, 2, 0, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 0, 12],
[0, 2, 0, 0, 0, 2, 0, 0, 0, 0, 0, 0, 4, 0, 0, 0, 0, 0, 4, 2, 0, 0, 0, 2, 4],
[0, 2, 0, 0, 0, 2, 0, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 0, 14],
[0, 1, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 6, 0, 0, 0, 0, 0, 4, 2, 0, 0, 0, 2, 4],
[0, 2, 0, 0, 0, 2, 0, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 0, 6, 0, 0, 0, 0, 0, 6],
[0, 2, 0, 0, 0, 2, 0, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 0, 4, 2, 0, 0, 0, 2, 12],
[0, 2, 0, 0, 0, 2, 0, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 0, 4, 2, 0, 0, 0, 2, 4],
[0, 2, 0, 0, 0, 2, 0, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 0, 4, 0, 0, 0, 0, 0, 14],
[0, 2, 0, 0, 0, 2, 0, 0, 0, 0, 0, 0, 4, 0, 0, 0, 0, 0, 4, 2, 0, 0, 0, 2, 8],
[0, 2, 0, 0, 0, 2, 0, 0, 0, 0, 0, 0, 6, 0, 0, 0, 0, 0, 4, 2, 0, 0, 0, 2, 6],
[0, 2, 0, 0, 0, 2, 0, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 0, 6, 0, 0, 0, 0, 0, 10],
[0, 2, 0, 0, 0, 2, 0, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 0, 30],
[0, 2, 0, 0, 0, 2, 0, 0, 0, 0, 0, 0, 10, 0, 0, 0, 0, 0, 4, 2, 0, 0, 0, 2, 4],
[0, 2, 0, 0, 0, 2, 0, 0, 0, 0, 0, 0, 4, 0, 0, 0, 0, 0, 6, 0, 0, 0, 0, 0, 6],
[0, 2, 0, 0, 0, 2, 0, 0, 0, 0, 0, 0, 12, 0, 0, 0, 0, 0, 4, 2, 0, 0, 0, 2, 4],
[0, 2, 0, 0, 0, 2, 0, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 0, 10],
[0, 2, 0, 0, 0, 2, 0, 0, 0, 0, 0, 0, 14, 0, 0, 0, 0, 0, 4, 2, 0, 0, 0, 2, 4],
[0, 2, 0, 0, 0, 2, 0, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 0, 6, 0, 0, 0, 0, 0, 14],
[0, 2, 0, 0, 0, 2, 0, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 0, 10, 0, 0, 0, 0, 0, 10],
[0, 2, 0, 0, 0, 2, 0, 0, 0, 0, 0, 0, 10, 0, 0, 0, 0, 0, 4, 2, 0, 0, 0, 2, 6],
[0, 2, 0, 0, 0, 2, 0, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 0, 4, 0, 0, 0, 0, 0, 30],
[0, 2, 0, 0, 0, 2, 0, 0, 0, 0, 0, 0, 4, 0, 0, 0, 0, 0, 4, 2, 0, 0, 0, 2, 16],
[0, 2, 0, 0, 0, 2, 0, 0, 0, 0, 0, 0, 20, 0, 0, 0, 0, 0, 4, 2, 0, 0, 0, 2, 4],
[0, 2, 0, 0, 0, 2, 0, 0, 0, 0, 0, 0, 6, 0, 0, 0, 0, 0, 4, 2, 0, 0, 0, 2, 4],
[0, 2, 0, 0, 0, 2, 0, 0, 0, 0, 0, 0, 28, 0, 0, 0, 0, 0, 4, 2, 0, 0, 0, 2, 4],
[0, 2, 0, 0, 0, 2, 0, 0, 0, 0, 0, 0, 30, 0, 0, 0, 0, 0, 4, 2, 0, 0, 0, 2, 4],
[0, 2, 0, 0, 0, 2, 0, 0, 0, 0, 0, 0, 6, 0, 0, 0, 0, 0, 4, 2, 0, 0, 0, 2, 16],
[0, 4, 0, 0, 0, 4, 0, 0, 0, 0, 0, 0, 6, 0, 0, 0, 0, 0, 4, 2, 0, 0, 0, 2, 6],
[0, 4, 0, 0, 0, 4, 0, 0, 0, 0, 0, 0, 10, 0, 0, 0, 0, 0, 4, 2, 0, 0, 0, 2, 4],
[0, 2, 0, 0, 0, 2, 0, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 0, 20, 10, 0, 0, 0, 10, 20],
[0, 2, 0, 0, 0, 2, 0, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 0, 10, 0, 0, 0, 0, 0, 30],
[0, 4, 0, 0, 0, 4, 0, 0, 0, 0, 0, 0, 14, 0, 0, 0, 0, 0, 4, 2, 0, 0, 0, 2, 4],
[0, 20, 0, 0, 0, 20, 0, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 0, 2],
[0, 2, 0, 0, 0, 2, 0, 0, 0, 0, 0, 0, 20, 0, 0, 0, 0, 0, 4, 2, 0, 0, 0, 2, 16],
[0, 4, 0, 0, 0, 4, 0, 0, 0, 0, 0, 0, 30, 0, 0, 0, 0, 0, 4, 2, 0, 0, 0, 2, 4],
[0, 4, 0, 0, 0, 4, 0, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 0, 20, 10, 0, 0, 0, 10, 20]]
V = [QuadraticForm(2*Matrix(QQ, 5, m)) for m in Scharlau_Walhorn_dim_4]
Q = Graph(len(V))
for i in Q.vertices():
Q.set_vertex(i, V[i])
for i in Q.vertices():
for j in Q.vertices():
if (i<j):
q1 = Q.get_vertex(i)
q2 = Q.get_vertex(j)
if q1.is_rationally_isometric(q2):
Q.add_edge(i,j)
ind = [sorted(c, key=lambda x: abs(Q.get_vertex(x).det()))[0] for c in Q.connected_components()]
for i in ind:
m = Scharlau_Walhorn_dim_4[i]
print("{},".format(m))
# +
# Scharlau-Walhorn dim 3 rational isometry classes (we need commensurability only)
# -
Scharlau_Walhorn_dim_3 = [[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 2, 1, 0, 0, 1, 2],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 2],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 2, 1, 0, 0, 1, 4],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 4],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 2, 1, 0, 0, 1, 6],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 4, 2, 0, 0, 2, 4],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 2, 1, 0, 0, 1, 8],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 4, 1, 0, 0, 1, 4],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 2, 1, 0, 0, 1, 10],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 10],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 4, 2, 0, 0, 2, 6],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 12],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 4, 0, 0, 0, 0, 6],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 4, 2, 0, 0, 2, 8],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 6, 1, 0, 0, 1, 6],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 6, 0, 0, 0, 0, 6],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 2, 1, 0, 0, 1, 20],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 20],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 4, 0, 0, 0, 0, 10],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 4, 2, 0, 0, 2, 12],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 26],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 28],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 4, 2, 0, 0, 2, 16],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 6, 3, 0, 0, 3, 12],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 34],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 6, 0, 0, 0, 0, 12],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 10, 5, 0, 0, 5, 10],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 42],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 6, 0, 0, 0, 0, 14],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 10, 4, 0, 0, 4, 10],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 10, 0, 0, 0, 0, 10],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 60],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 4, 0, 0, 0, 0, 30],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 6, 0, 0, 0, 0, 20],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 10, 0, 0, 0, 0, 12],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 66],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 12, 2, 0, 0, 2, 12],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 14, 7, 0, 0, 7, 14],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 6, 0, 0, 0, 0, 28],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 12, 0, 0, 0, 0, 14],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 6, 0, 0, 0, 0, 30],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 14, 0, 0, 0, 0, 14],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 10, 0, 0, 0, 0, 20],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 20, 10, 0, 0, 10, 20],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 6, 0, 0, 0, 0, 60],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 12, 0, 0, 0, 0, 30],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 26, 13, 0, 0, 13, 26],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 28, 14, 0, 0, 14, 28],
[0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 30, 0, 0, 0, 0, 30]]
V = [QuadraticForm(2*Matrix(QQ, 4, m)) for m in Scharlau_Walhorn_dim_3]
Q = Graph(len(V))
for i in Q.vertices():
Q.set_vertex(i, V[i])
for i in Q.vertices():
for j in Q.vertices():
if (i<j):
q1 = Q.get_vertex(i)
q2 = Q.get_vertex(j)
if q1.is_rationally_isometric(q2):
Q.add_edge(i,j)
ind = [sorted(c, key=lambda x: abs(Q.get_vertex(x).det()))[0] for c in Q.connected_components()]
for i in ind:
m = Scharlau_Walhorn_dim_3[i]
print("{},".format(m))
| Lattices-3-4-5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h2> ====================================================</h2>
# <h1>MA477 - Theory and Applications of Data Science</h1>
# <h1>Lesson 26: Neural Networks (Part 3)</h1>
#
# <h4>Dr. <NAME></h4>
# <br>
# United States Military Academy, West Point, AY20-2
# <h2>=====================================================</h2>
#
#
# <h2>Lecture Outline</h2>
#
# <ul>
# <li> Review of the Perceptron (1-Layer NN)</li>
# <li>Introduction of a 2-Layer NN</li>
# <li> Forward Propagation
# <ol>
# <li> Single Training Sample</li>
# <li> Vectorized Version</li>
# <li>Vectorized Version for Multiple Training Samples</li>
# </ol></li>
#
# </li>
#
# </ul>
# <h2> Review: Perceptron (1-Layer NN)</h2>
#
# 
# <br>
#
#
# <font size='5'>
# $$\fbox{$x\,\&\,w \, \&\, b$}\longrightarrow\fbox{$z=w^Tx+b$}\Rightarrow\fbox{$a=\sigma(z)$}\longrightarrow \fbox{$L(a,y)$}$$
# </font>
# <h2> A 2-Layer Neural Network</h2>
#
# In what follows we will build the following simple 2-layer Neural Network. This means we have one input layer, one hidden layer, and one output layer. For simplicity of presentation, we will only have two neurons in the hidden layer. We will first focus on the theoretical aspects and then discuss the Python implementaion.
#
# 
# <h2>Forward Propagation</h2>
#
# The underlying mechanism of the above 2-layer NN is very similar to that of a Perceptron, but repeated multiple times. In this case, one may think of the above 2-layer NN as a collection of three Perceptrons!
#
# In what follows, we will show step-by-step how to compute the output of the NN, namely $\hat{y}=a^{[2]}$.
#
# <b> Notation:</b> Superscripts in square brackets denote the number of the layer we are at, while the subscript the node we are at. For example $a^{[1]}_2$ means that we are in layer 1 and node 2. In general, $a^{[l]}_i$ denotes the computation at layer $l$ in node $i$.
#
# Recall, that at each node there are two computations happening, a summation and an application of the activation function. Specifically:
#
# <font size='3'>
# Let $$a^{[0]}=\begin{bmatrix}x_1\\x_2\\x_3\end{bmatrix}$$
#
# Then,
#
# $$z_1^{[1]}=w^{[1]\,T}_1a^{[0]}+b_1^{[1]}\, \Rightarrow a_1^{[1]}=\sigma\left(z_1^{[1]}\right)$$
#
# $$z_2^{[1]}=w^{[1]\,T}_2a^{[0]}+b_2^{[1]}\, \Rightarrow a_2^{[1]}=\sigma\left(z_2^{[1]}\right)$$
#
#
# So $$a^{[1]}=\begin{bmatrix}a_1^{[1]}\\a_2^{[1]}\end{bmatrix}$$
#
# is the output of the hidden layer.
#
# So, the final output is $$z^{[2]}=w^{[2]\,T}a^{[1]}+b^{[2]}\, \Rightarrow a^{[2]}=\sigma\left(z^{[2]}\right)$$
#
# </font>
# <font size='3'>
#
# <h3>Vectorized Version</h3>
#
# Let $$
# z^{[1]}=\begin{bmatrix} z_1^{[1]}\\z_2^{[1]}\end{bmatrix}_{2\times 1}\, ,\,
# W^{[1]}=
# \begin{bmatrix}
# w_1^{[1]\, T}\\ w_2^{[1]\, T}
# \end{bmatrix}_{2 \times 3}
# \, , \,
# b^{[1]}=\begin{bmatrix}b^{[1]}_1 \\ b^{[1]}_2\end{bmatrix}_{2 \times 1}
# $$
#
# So, the output of the hidden layer is computed as follows:
#
# $$z^{[1]}=W^{[1]}a^{[0]}+b^{[1]} \, \text{ and } a^{[1]}=\sigma\left(z^{[1]}\right)$$
#
#
# Since we have only one node in the output layer, the final ouptput computation is the same as above:
#
# $$z^{[2]}=W^{[2]}a^{[1]}+b^{[2]}\, \Rightarrow a^{[2]}=\sigma\left(z^{[2]}\right)$$
#
# We want to note that $W^{[2]}$ is a $1\times 2$ matrix and $b^{[2]}$ is a scalar.
#
# To summarize, the output is computed as below:
#
# \begin{eqnarray*}
# z^{[1]}&=&W^{[1]}a^{[0]}+b^{[1]}\\
# a^{[1]}&=&\sigma\left(z^{[1]}\right)\\
# & &\\
# z^{[2]}&=&W^{[2]}a^{[1]}+b^{[2]}\\
# a^{[2]}&=&\sigma\left(z^{[2]}\right)
# \end{eqnarray*}
#
#
#
#
# <font color='red' size='5'>Exercise:</font> Take about 5 minutes to verify these computations.
#
# $$z^{[1]}=W^{[1]}a^{[0]}+b^{[1]}= \begin{bmatrix}
# w_1^{[1]\, T}\\ w_2^{[1]\, T}
# \end{bmatrix}_{2 \times 3}
# a^{[0]}+\begin{bmatrix}b^{[1]}_1 \\b^{[1]}_2\end{bmatrix}_{2 \times 1}=\begin{bmatrix}w_1^{[1]\, T}a^{[0]}\\w_2^{[1]\, T}a^{[0]}\end{bmatrix}+\begin{bmatrix}b^{[1]}_1 \\b^{[1]}_2\end{bmatrix}_{2 \times 1}$$
#
# </font>
# <h2>Multiple Training Samples</h2>
#
#
# <font size='3'>
#
# Suppose now we have a training data of $m$ examples $$\{(x^{(1)},y^{(1)}),\dots,(x^{(m)},y^{(m)})\}$$
#
# So, for each training example $x^{(i)}$ we need to compute the output of the neural network, namely, $\hat{y}^{(i)}=a^{[2](i)}$, $i=1,2,3,\dots,m.$ The superscript in round brackets $(i)$ corresponds to the training sample, wheras the superscript in the square brackets correspond to the particular layer of the NN.
#
# So, to compute all the outputs we would need to iterate over all of the training samples.
#
# For $i$ from $1$ to $m$:
#
# $\hspace{1cm} z^{[1](i)}=W^{[1]}a^{[0](i)}+b^{[1]}$
#
# $\hspace{1cm}a^{[1](i)}=\sigma\left(z^{[1](i)}\right)$
#
# $\hspace{1cm} z^{[2](i)}=W^{[2]}a^{[1](i)}+b^{[2]}$
#
# $\hspace{1cm}a^{[2](i)}=\sigma\left(z^{[2](i)}\right)$
#
#
#
# As we discussed last time, for-loops are computationally costly, so if possible we should avoid them.
#
# We can vectorize this process across all the $m$ trainings samples in a similar way as for the Perceptron, and in effect compute the outputs essentially at the same time for all the training samples.
#
#
# Let
#
# $$A^{[0]}:=X=\begin{bmatrix}x^{(1)} & x^{(2)}&\dots &x^{(m)}\end{bmatrix}_{n_x\times m}=\begin{bmatrix}a^{[0](1)} & a^{[0](2)}&\dots &a^{[0](m)}\end{bmatrix}_{n_x\times m}$$
#
# be an $n_x \times m$ matrix (in our case $n_x=3$) containing all of the $m$ training samples. Then, the forward propagation of the NN above can be computed as follows:
#
# \begin{eqnarray*}
# Z^{[1]}&=&W^{[1]}A^{[0]}+b^{[1]}\\
# A^{[1]}&=&\sigma\left(Z^{[1]}\right)\\
# & &\\
# Z^{[2]}&=&W^{[2]}A^{[1]}+b^{[2]}\\
# A^{[2]}&=&\sigma\left(Z^{[2]}\right)\\
# \end{eqnarray*}
#
# Where
#
# \begin{eqnarray*}
# Z^{[1]}&=&\begin{bmatrix}z^{[1](1)}& z^{[1](2)} &\dots &z^{[1](m)}\end{bmatrix}_{2\times m}\\
# A^{[1]}&=&\begin{bmatrix}a^{[1](1)} &a^{[1](2)} &\dots & a^{[1](m)}\end{bmatrix}_{2\times m}\\
# & &\\
# Z^{[2]}&=&\begin{bmatrix}z^{[2](1)}& z^{[2](2)} &\dots &z^{[2](m)}\end{bmatrix}_{1\times m}\\
# A^{[2]}&=&\begin{bmatrix}a^{[2](1)} &a^{[2](2)} &\dots & a^{[2](m)}\end{bmatrix}_{1\times m}
# \end{eqnarray*}
#
# One way to make this notation more tangible is to think of it as follows: horizontally they iterate through different training examples wheras vertically they run through different nodes on each layer.
#
#
# <font color='red' size='5'>Exercise:</font> Take about 5-10 minutes to verify these computations!
#
# </font>
# <h2> ====================================================</h2>
# <h1>MA477 - Theory and Applications of Data Science</h1>
# <h1>Lesson 27: Neural Networks (Part 4)</h1>
#
# <h4>Dr. <NAME></h4>
# <br>
# United States Military Academy, West Point, AY20-2
# <h2>=====================================================</h2>
# <h2>Choice of activation functions and why we need them!</h2>
#
# <h3>2-Layer ANN</h3>
#
# 
#
# Recall the forward-propagation computations:
#
# \begin{eqnarray*}
# Z^{[1]}&=&W^{[1]}A^{[0]}+b^{[1]}\\
# A^{[1]}&=&G^{[1]}\left(Z^{[1]}\right)\\
# & &\\
# Z^{[2]}&=&W^{[2]}A^{[1]}+b^{[2]}\\
# A^{[2]}&=&G^{[2]}\left(Z^{[2]}\right)\\
# \end{eqnarray*}
#
# Here, instead of using the Sigmoid function as an activation function, we may use other activation functions as well. Moreover, we can use different activation functions for the hidden-layers and a different one for the output layer. In the algorithm above, we have denoted them by $G^{[1]}$ and $G^{[2]}$.
#
# So, what other options are there? Are there any general criteria that a function needs to satisfy in order to be used as an activation function?
#
# We will briefly discuss this below, but a function must be a <b>non-linear</b> function in order to be useful as an activation function.
#
# <h4> Hyperbolic Tangent Function $tanh$</h4>
#
# Hyperbolic tangent function may be used as an activation function as well. It is defined as follows:
#
# $$tanh(z)=\frac{e^z-e^{-z}}{e^z+e^{-z}}$$
#
# Let's graph it and see what it looks like:
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
def tanh(z):
return (np.exp(z)-np.exp(-z))/(np.exp(z)+np.exp(-z))
# +
sns.set_style('whitegrid')
plt.figure(figsize=(10,6))
z=np.linspace(-4,4,50)
plt.plot(z,tanh(z),label='$tanh(z)$')
plt.legend(fontsize=16)
plt.show()
# -
# For the <b>hidden layers</b>, the $tanh$ function is a better choice than the Sigmoid function. One reason for that is the fact that the output of $tanh$ is between -1 and 1 with a center at 0, so it has the effect of centering the data it outputs, which in turn makes learning better for the next layer.
#
# However, for the <b>output</b> layers, it may depend on the whether it makes sense or not. For example, if we are making a classification and we want the output to be either a zero or a one(for binary classification problems), then it doesn't make sense to have $tanh$ as an activation function on the output layer, so Sigmoid function may be a better choice and makes more sense in terms of the range of outcomes.
#
# One main disadvantage of both $tanh$ and the Sigmoid function is that for relatively large(positive or negative) values their derivatives(slopes) become very close to zero, which in turn may slow down the gradient descent significantly. In other words, the learning process may be very slow. This is knowns as the <b> vanishing gradient problem</b>, and it is something serious that one needs to keep in mind especially when creating deep neural networks with very complex architectures.
# <h3>Rectifier Linear Unit (ReLU)</h3>
#
# ReLU is widely used in NN, especially in hidden layers. It's defined as follows:
#
# $$ReLu(z)=\max(0,z)$$
#
# Let's graph it:
def relu(z):
return [max(0,i) for i in z]
plt.figure(figsize=(8,6))
plt.plot(z,relu(z), label='$max(0,z)$')
plt.legend(fontsize=16)
# Part of the reason why $ReLu$ is popular is because it has a very simple derivative. Namely, it is constantly a $1$ for all positive values and zero otherwise. Since the weights are initialized randomly, in practice there is enough positive values to contribute to weight updates at each step.
#
# There are also <b> smoothed</b> versions of ReLu.
#
# <h3> SmoothReLu</h3>
#
# $$G(x)=\log(1+e^x)$$
#
# This is has very similar behavior to ReLu except it is smoothed out so it has a derivative everywhere.
def smoothrelu(z):
return np.log(1+np.exp(z))
plt.figure(figsize=(8,6))
z=np.linspace(-5,5,100)
plt.plot(z,smoothrelu(z),label='$\log(1+e^x)$')
plt.legend(fontsize=16)
# <b> ReLu</b> (along with its different versions) is probably the most widely used activation function for hidden layers. The main reason for this is because the learning is much faster as opposed to using $Tanh$ or $Sigmoid$. Typically, for binary classification problem, where you hav an output of 0 or 1 a natural choice for activation function is the Sigmoid function. However, for other problems, for example if you are trying to predict the house pricing, or insurance pricing etc. then you may use ReLU as an activation function for the output layer.
# <h3> Why Non-linear Activation Functions?</h3>
#
# Why can't we just use for example $$G^{[1]}(z)=z \, \text{ and } \, G^{[2]}(z)=z\, ???$$
#
# We'll see in just a bit that in order to be able to capture the complex relationship between the features and the response variable we need to have non-linear activation functions, else regardless of how many hidden layers we have, if we only use linear activation functions then we do no better than a simple Logistic Regression.
#
# To see this, let's follow the line of computations above:
#
#
#
# $$Z^{[1]}=W^{[1]}A^{[0]}+b^{[1]}$$
# $$A^{[1]}=G^{[1]}\left(Z^{[1]}\right)=Z^{[1]}=W^{[1]}A^{[0]}+b^{[1]}$$
#
# $$Z^{[2]}=W^{[2]}A^{[1]}+b^{[2]}=W^{[2]}\left(W^{[1]}A^{[0]}+b^{[1]}\right)+b^{[2]}=\left(W^{[2]}W^{[1]}\right)A^{[0]}+\left((W^{[2]}b^{[1]}+b^{[2]}\right)$$
#
# $$A^{[2]}=G^{[2]}\left(Z^{[2]}\right)=C_1A^{[0]}+C_0$$
#
# So, $$\hat{y}=C_1X+C_0$$
#
# where $$C_1=W^{[2]}W^{[1]}\, \text{ and }\, C_0=W^{[2]}b^{[1]}+b^{[2]}$$
#
# In other words, if all the activation functions are <b>linear</b> for all of the layers of a NN then the Neural Net will only output a linear function, and thus will be unable to capture any <b> non-linear</b> relationship that may exist betwen the features and the response variable.
#
#
# <h2>Backpropagation</h2>
#
# <h3>Derivatives of Activation Functions</h3>
#
# Recall that in order to perform Gradient Descent, at some point in the process, we need to compute the derivatives of activation function. Let's do that below, so that we have them in one place:
#
# <ol>
# <li> $$\sigma(z)=\frac{1}{1+e^{-z}}\,\Rightarrow\, \frac{d\sigma(z)}{dz}=\sigma(z)\left(1-\sigma(z)\right)$$</li>
# <br>
#
# <li>$$\tanh(z)=\frac{e^z-e^{-z}}{e^z+e^{-z}}\, \Rightarrow\, \frac{d}{dz}\tanh(z)=1-\tanh^2(z)$$</li>
# <br>
#
# <li>$$ReLu(z)=\max(0,z)\, \Rightarrow\, \frac{d}{dz}ReLu(z)=\begin{cases}1 & \text{ if } x\geq 0\\ 0 & \text{ if }x<0\end{cases}$$</li>
#
# </ol>
#
#
# <h3>Gradient Descent</h3>
#
# If the structure of our neural network is as follows:
#
# \begin{eqnarray*}
# n^{[0]}&:=&n_x=\text{ number of input features (neurons in input layers)}\\
# n^{[1]}&=&\text{ number of neurons in the first hidden layer}\\
# n^{[2]}&=&\text{ number of neurons in the second layer (output layer for us)}\\
# \end{eqnarray*}
#
# For us $ n^{[0]}=3\, , \, n^{[1]}=2,\, \, n^{[2]}=1.$
#
# Then the dimensions of the weight and bias matrices are:
#
# $$W^{[1]}=\text{ is an } n^{[1]} \times n^{[0]} \text{ matrix }$$
#
# $$b^{[1]}=\text{ is an } n^{[1]}\times 1 \text{ vector }$$
#
# $$W^{[2]}=\text{ is an } n^{[2]}\times n^{[1]} \text{ matrix}$$
#
# $$b^{[2]}=\text{ is a} 1\times 1 \text{ vector( a scalar in this case)}$$
#
# The cost function:
#
# $$J\left(W^{[1]},b^{[1]},W^{[2]},b^{[2]}\right)=\frac{1}{m}\sum_{i=1}^{m}L\left(\hat{y},y\right)$$
#
# where, if we are doing binary classification problems, then the loss function $L(\hat{y},y)$ is exactly as before.
#
# <h4>Backpropagation:</h4>
#
# \begin{eqnarray*}
# dZ^{[2]}&=&A^{[2]}-Y \,(\text{ dimensions: } n^{[2]}\times m)\\
# dW^{[2]}&=&\frac{1}{m}dZ^{[2]}A^{[1]\, T}\, (\text{ dimensions: } n^{[2]}\times n^{[1]})\\
# db^{[2]}&=&\frac{1}{m}np.sum(dZ^{[2]},axis=1,keepdims=True)\, (\text{ dimensions:} n^{[2]}\times 1)\\
# &&\\
# dZ^{[1]}&=&W^{[2]\, T}dZ^{[2]}\times \frac{d}{dZ^{[1]}}G^{[1]}\left(Z^{[1]}\right)\, (\text{ dimensions: } n^{[1]}\times m)\\
# dW^{[1]}&=&\frac{1}{m}dZ^{[1]}A^{[0]\, T}\,(\text{ dimensions: } n^{[1]}\times n^{[0]})\\
# db^{[1]}&=&\frac{1}{m}np.sum(dZ^{[1]},axis=1,keepdims=True)\,(\text{ dimensions:} n^{[1]}\times 1)\\
# \end{eqnarray*}
#
# <h4>Weight Update:</h4>
#
# \begin{eqnarray*}
# W^{[1]}&:=&W^{[1]}-\alpha dW^{[1]}\\
# b^{[1]}&:=&b^{[1]}-\alpha db^{[1]}\\
# &&\\
# W^{[2]}&:=&W^{[2]}-\alpha dW^{[2]}\\
# b^{[2]}&:=&b^{[2]}-\alpha db^{[2]}\\
# \end{eqnarray*}
#
#
# <h2>Python Implementation</h2>
#
# To put it all together, the following algorithm is what we need to implement in Python:
#
# For $iter$ in $range(1,N):$
#
# \begin{eqnarray*}
# \text{ Forward Propagation }\\
# Z^{[1]}&=&W^{[1]}A^{[0]}+b^{[1]}\\
# A^{[1]}&=&G^{[1]}\left(Z^{[1]}\right)\\
# & &\\
# Z^{[2]}&=&W^{[2]}A^{[1]}+b^{[2]}\\
# A^{[2]}&=&G^{[2]}\left(Z^{[2]}\right)\\
# J+&=&\frac{1}{N}L(A^{[2]},Y)\\
# \end{eqnarray*}
#
# \begin{eqnarray*}
# \text{ Backpropagation}\\
# dZ^{[2]}&=&A^{[2]}-Y \\
# dW^{[2]}&=&\frac{1}{m}dZ^{[2]}A^{[1]\, T}\\
# db^{[2]}&=&\frac{1}{m}np.sum(dZ^{[2]},axis=1,keepdims=True)\\
# &&\\
# dZ^{[1]}&=&W^{[2]\, T}dZ^{[2]}\times \frac{d}{dZ^{[1]}}G^{[1]}\left(Z^{[1]}\right)\\
# dW^{[1]}&=&\frac{1}{m}dZ^{[1]}A^{[0]\, T}\\
# db^{[1]}&=&\frac{1}{m}np.sum(dZ^{[1]},axis=1,keepdims=True)\\
# \end{eqnarray*}
#
# \begin{eqnarray*}
# \text{ Weight Update}\\
# W^{[1]}&:=&W^{[1]}-\alpha dW^{[1]}\\
# b^{[1]}&:=&b^{[1]}-\alpha db^{[1]}\\
# &&\\
# W^{[2]}&:=&W^{[2]}-\alpha dW^{[2]}\\
# b^{[2]}&:=&b^{[2]}-\alpha db^{[2]}\\
# \end{eqnarray*}
#
# <h3> NN Structure</h3>
import pandas as pd
import numpy as np
X=pd.DataFrame(np.random.randn(3,7),index=['weight','height','age'])
X.shape[0]
X
Y=np.random.randn(1,7)
W1=np.random.normal(scale=0.01,size=(2,3))
b1=np.zeros(shape=(2,1))
np.tanh(W1)
def sigmoid(z):
return 1/(1+np.exp(-z))
def nn_structure(X, Y):
"""
Arguments:
X -- input dataset of shape (input size, number of examples)
Y -- labels of shape (output size, number of examples)
Output:
n_x -- number of neurons/features in the input layer
n_h -- number of neurons in the hidden layer
n_y -- number of neurons in the output layer
"""
#COMPLETE THE REST OF THE CODE
n_x = X.shape[0]
n_h = 2
n_y = 1
return (n_x, n_h, n_y)
# <h3> Weight & Bias Initialization</h3>
# +
# GRADED FUNCTION: initialize_parameters
def initialize_parameters(n_x, n_h, n_y):
"""
Argument:
n_x -- number of neurons/features in the input layer
n_h -- number of neurons in the hidden layer
n_y -- number of neurons in the output layer
Returns:
weights_bias -- dicionary storing the weights and biases for each of the layers:
W1 -- weight matrix of shape (n_h, n_x)
b1 -- bias vector of shape (n_h, 1)
W2 -- weight matrix of shape (n_y, n_h)
b2 -- bias vector of shape (n_y, 1)
"""
#COMPLETE THE REST OF THE CODE
W1 = np.random.normal(scale=0.01,size=(n_h,n_x))
b1 = np.zeros(shape=(n_h,1))
W2 = np.random.normal(scale=0.01,size=(n_y,n_h))
b2 = np.zeros(shape=(n_y,1))
weights_biases = {"W1": W1,"b1": b1,"W2": W2,"b2": b2}
return weights_biases
# -
# <h3>Forward Propagation</h3>
# +
# GRADED FUNCTION: forward_propagation
def forward_propagation(X, weights_biases):
"""
Arguments:
X -- input data of size (n_x, m)
weights_biases -- python dictionary containing your parameters (output of initialization function)
Returns:
A2 -- The output of the NN
outputs_of_layers -- a dictionary containing "Z1", "A1", "Z2" and "A2"
"""
# Retrieve each parameter from the dictionary "parameters"
#COMPLETE THE REST OF THE CODE
W1 = weights_biases['W1']
b1 = weights_biases['b1']
W2 = weights_biases['W2']
b2 = weights_biases['b2']
#COMPLETE THE REST OF THE CODE
Z1 = np.matmul(W1,X)+b1
A1 = np.tanh(Z1)
Z2 = np.matmul(W2,A1)+b2
A2 = sigmoid(Z2)
outputs_of_layers = {"Z1": Z1,"A1": A1,"Z2": Z2,"A2": A2}
return A2, outputs_of_layers
# -
| MA477 - Theory and Applications of Data Science/Lessons/Lesson 26 - Shallow Neural Networks/Lesson 26 - Shallow Neural Networks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Calibration and evaluation of a rainfall-runoff model
# Imagine that we want to simulate the natural inflows into a water reservoir, knowing the amount of rainfall that has fallen in the reservoir’s catchment area. For that purpose, we can use a rainfall-runoff model. A rainfall-runoff model is a mathematical model describing the rainfall–runoff processes that occur in a watershed. The model consists of a set of equations, which describe the various processes of soil infiltration, surface and subsurface runoff, etc., as a function of various parameters, which describe the hydrological characteristics of the watershed, ultimately enabling the estimation of the flow at selected river sections.
#
# To tailor a generic rainfall-runoff model to a particular catchment, model calibration is required. Model calibration is the process of adjusting the model parameters to obtain a representation of the system under study that satisfies pre-agreed criteria. Normally, calibration of a rainfall-runoff model aims to improve the fit of the simulated flows to observed flows and involves running the model many times under different combinations of the parameter values, until a combination is found that minimise the differences between simulated and observed flows.
#
#
# ## The HBV rainfall-runoff model: general structure
# In this example, we will use the HBV rainfall-runoff model [(Bergström, 1992)](https://www.smhi.se/en/publications/the-hbv-model-its-structure-and-applications-1.83591). The HBV model is a lumped hydrological model, meaning that all the processes included in the model are spatially aggregated into “conceptual” representations at the catchment scale. In brief, the structure, forcing inputs, parameters and output of the model are the following.
#
# #### Structure
#
# The model consists of four main modules/subroutines:
# 1. **SM module**: for soil moisture (***SM***), actual evapotranspiration (***EA***) and recharge estimation (***R***)
# 2. **UZ module**: for upper zone runoff generation (***Q0*** = surface runoff + interflow) and percolation (***PERC*** = water flux from upper to lower zone)
# 3. **LZ module**: for lower zone runoff generation (***Q1*** = baseflow)
# 4. **Routing module**: for runoff routing.
#
# <left><img src="../../Software/HBV model structure.png" width = "800px"><left>
#
# #### Forcing inputs
#
# The forcing inputs of the model simulation are time series of observed precipitation (***P***) and estimated potential evapotranspiration (***E***) – these are the spatial averages of precipitation and evapotranspiration across the watershed area. Usually these time series are given at daily resolution, and this will be the case in our example too, but it is possible to use shorter time steps.
#
# #### Model parameters:
# In order to tailor the general model equations to the particular watershed under study, we need to specify the watershed surface area, and a number of other parameters that characterise the climate, geology, soil properties, etc. of that place. These parameters are:
#
#
# 1. ***SSM0*** = initial soil moisture [mm]
# 2. ***SUZ0*** = initial Upper Zone storage [mm]
# 3. ***SLZ0*** = initial Lower Zone storage [mm]
# 4. ***BETA*** = Exponential parameter in soil routine [-]
# 5. ***LP*** = Limit for potential evapotranspiration [-]
# 6. ***FC*** = Maximum soil moisture content [mm]
# 7. ***PERC*** = Maximum flux from Upper to Lower Zone [mm/day]
# 8. ***K0*** = Near surface flow coefficient
# 9. ***K1*** = Recession coefficient for the Upper Zone (ratio) [1/day]
# 10. ***K2*** = Recession coefficient for the Lower Zone (ratio) [1/day]
# 11. ***UZL*** = Near surface flow threshold [mm]
# 12. ***MAXBAS*** = Transfer function parameter [day]
#
# #### Model outputs
# For a given selection of the model parameters and forcing input time series, the model simulation returns time series of the following state and flux variables:
#
# 1. ***EA*** = Actual Evapotranspiration [mm/day]
# 2. ***SM*** = Soil Moisture [mm]
# 3. ***R*** = Recharge (water flow from Soil to Upper Zone) [mm/day]
# 4. ***UZ*** = Upper Zone water content [mm]
# 5. ***LZ*** = Lower Zone water content [mm]
# 6. ***RL*** = Recharge to the Lower Zone [mm]
# 7. ***Q0*** = Water flow from Upper Zone [ML/day]
# 8. ***Q1*** = Water flow from Lower Zone [ML/day]
# 9. ***Qsim*** = Total water flow [ML/day]
# To run the model we need to import some necessary libraries. (🚨 in order to run the code like in the box below, place the mouse pointer in the cell, then click on “run cell” button above or press shift + enter)
# Libraries for visualization and interactivity
from bqplot import pyplot as plt
from bqplot import *
from bqplot.traits import *
import ipywidgets as widgets
from IPython.display import display
# Library for scientific computing
import numpy as np
# Library for manipulating dates and times
from datetime import datetime, timedelta
# # Library for general purposes
import sys
warnings.filterwarnings('ignore') # to ignore warning messages
# We also need to import several tools from the iRONs toolbox:
from irons.Software.HBV_sim import HBV_sim # HBV model
from irons.Software.HBV_calibration import HBV_calibration # HBV model calibration
# # Application to the Wimbleball reservoir's catchment
#
# The catchment is, located in the south-west of England, it has a drainage area of 28.8 km2, and collects water from the river Haddeo and drains into the Wimbleball reservoir.
#
# ## Loading and visualizing data
#
# ### Area
# Let’s first define the extent of the watershed surface area.
area = 28.8 # km2
# ### Climate data
# We call a sub-routine to load daily historical climate data (evapotranspiration, precipitation and temperature) of our study area for the year 2000.
from Modules.Historical_data import Climate_data, Flow_data # To load historical climate and streamflow data
cal_year = 2000
clim_date, E, P, T = Climate_data(cal_year)
# Plotting the precipitation data
# + code_folding=[]
# Let's create a scale for the x attribute, and a scale for the y attribute
x_sc_1 = DateScale()
y_sc_1 = LinearScale()
x_ax_1 = Axis(label='date', scale=x_sc_1)
y_ax_1 = Axis(label='mm/day', scale=y_sc_1, orientation='vertical')
fig_1 = plt.figure(title = 'daily precipitation', axes=[x_ax_1, y_ax_1], scales={'x': x_sc_1, 'y': y_sc_1},
layout={'min_width': '1000px', 'max_height': '300px'})
precip_bars = plt.bar(clim_date,P,colors=['blue'],stroke = 'lightgray')
fig_1
# -
# ### Observed flow
# We call a sub-routine to load daily historical flow data of our watershed area for the year 2000 (this data will be used to compare against the model predictions).
Q_obs_date, Q_obs = Flow_data(cal_year)
# Plotting the observed flow data
# + code_folding=[]
x_sc_2 = DateScale()
y_sc_2 = LinearScale(max=1200)
x_ax_2 = Axis(label='date', scale=x_sc_2)
y_ax_2 = Axis(label='ML/day', scale=y_sc_2, orientation='vertical')
fig_2 = plt.figure(title = 'observed daily water flow', axes=[x_ax_2, y_ax_2],scales={'x': x_sc_2, 'y': y_sc_2},
layout={'min_width': '1000px', 'max_height': '300px'})
obs_flow = plt.plot(Q_obs_date,Q_obs,colors=['black'])
fig_2
# -
# # Manual model calibration
# First we will try to calibrate the model manually, that is, changing the parameter values one at the time and looking at the effects induced in the model predictions by means of an interactive plot of the simulated hydrograph. The objective is to obtain a good fit of the simulated hydrograph to the observed one.
#
# To measure the goodness-of-fit between the simulated and the observed flow we will use the root mean square error (RMSE). RMSE is the standard deviation of the prediction errors, i.e. the difference between the simulated (***s(t)***) and the observed (***o(t)***) hydrograph.
# $$RMSE = \sqrt{\frac{\sum_{t=0}^{T} (s(t)-o(t))^{2}}{T}}$$
#
# First, let’s execute the code below to define the sliders that will later appear in the interactive hydrograph:
# + code_folding=[]
# Interactive sliders definition
def update_sim_hyd(P,E,param,Case,ini):
Q_sim,[SM,UZ,LZ],[EA,R,RL,Q0,Q1] = HBV_sim(P,E,param,Case,ini,area)
RMSE = np.sqrt(((Q_sim - Q_obs) ** 2).mean())
return Q_sim,RMSE
def params_changed(change):
y_vals = update_sim_hyd(P,E,[BETA.value, LP.value, FC.value, PERC.value, K0.value, K1.value, K2.value,
UZL.value, MAXBAS.value],1,[SSM0.value,SUZ0.value,SLZ0.value])[0]
RMSE = update_sim_hyd(P,E,[BETA.value, LP.value, FC.value, PERC.value, K0.value, K1.value, K2.value,
UZL.value, MAXBAS.value],1,[SSM0.value,SUZ0.value,SLZ0.value])[1]
sim_hyd.y = y_vals
fig_3.title = 'RMSE = ' +str("%.2f" % RMSE)
SSM0 = widgets.FloatSlider(min = 0, max = 400, step=10, value = 200, description = 'Initial soil moisture ($mm$)',
style = {'description_width': '300px'} ,layout={'width': '600px'})
SSM0.observe(params_changed,'value')
SUZ0 = widgets.FloatSlider(min = 0, max = 100, step=.5, value = 50, description = 'Initial water content of UZ ($mm$)',
style = {'description_width': '300px'} ,layout={'width': '600px'})
SUZ0.observe(params_changed,'value')
SLZ0 = widgets.FloatSlider(min = 0, max = 100, step=.5, value = 50, description = 'Initial water content of LZ ($mm$)',
style = {'description_width': '300px'} ,layout={'width': '600px'})
SLZ0.observe(params_changed,'value')
BETA = widgets.FloatSlider(min = 0, max = 7, value = 3.5, description = 'Exponential parameter in soil routine (-)',
style = {'description_width': '300px'} ,layout={'width': '600px'})
BETA.observe(params_changed,'value')
LP = widgets.FloatSlider(min = 0.3, max = 1, step=0.05,value = 0.65,description = 'Limit for potential evapotranspiration (-)',
style = {'description_width': '300px'} ,layout={'width': '600px'})
LP.observe(params_changed,'value')
FC = widgets.FloatSlider(min = 1, max = 2000, value = 1000, description = 'Maximum soil moisture content ($mm$)',
style = {'description_width': '300px'} ,layout={'width': '600px'})
FC.observe(params_changed,'value')
PERC = widgets.FloatSlider(min = 0, max = 100, value = 50, description = 'Maximum flow from UZ to LZ ($mm$ $day^{-1}$)',
style = {'description_width': '300px'} ,layout={'width': '600px'})
PERC.observe(params_changed,'value')
K0 = widgets.FloatSlider(min = 0, max = 2, step=0.05, value = 1, description = 'Near surface flow coefficient (-)',
style = {'description_width': '300px'} ,layout={'width': '600px'})
K0.observe(params_changed,'value')
K1 = widgets.FloatSlider(min = 0, max = 1, value = 0.5, description = 'Recession coefficient for UZ ($day^{-1}$)',
style = {'description_width': '300px'} ,layout={'width': '600px'})
K1.observe(params_changed,'value')
K2 = widgets.FloatSlider(min = 0, max = 0.1, step=0.005,value = 0.05,description = 'Recession coefficient for LZ ($day^{-1}$)',
style = {'description_width': '300px'} ,layout={'width': '600px'})
K2.observe(params_changed,'value')
UZL = widgets.FloatSlider(min = 0, max = 100, value = 50, description = 'Near surface flow threshold ($mm$)',
style = {'description_width': '300px'} ,layout={'width': '600px'})
UZL.observe(params_changed,'value')
MAXBAS = widgets.FloatSlider(min = 1, max = 6, step=0.5, value = 3.5, description = 'Transfer function parameter ($day$)',
style = {'description_width': '300px'} ,layout={'width': '600px'})
MAXBAS.observe(params_changed,'value')
# -
# Now, let’s play with the interactive hydrograph! To start try to visually fit the simulated hydrograph to the observed one in January by playing only with the parameters defining the initial conditions and the **Maximum soil moisture content**:
# +
x_ax_3 = Axis(label='date', scale=x_sc_2)
y_ax_3 = Axis(label='ML/day', scale=y_sc_2, orientation='vertical')
ini = [SSM0.value,SUZ0.value,SLZ0.value]
param = [BETA.value, LP.value, FC.value, PERC.value, K0.value, K1.value, K2.value, UZL.value, MAXBAS.value]
fig_3 = plt.figure(title = 'RMSE = ' +str("%.2f ML/day" % update_sim_hyd(P,E,param,1,ini)[1]), axes=[x_ax_3, y_ax_3],
scales={'x': x_sc_2, 'y': y_sc_2}, layout={'min_width': '900px', 'max_height': '200px'},
animation_duration=1000,fig_margin = {'top':0, 'bottom':40, 'left':60, 'right':0})
obs_hyd = plt.plot(Q_obs_date,Q_obs,colors=['black'])
sim_hyd = plt.plot(Q_obs_date,update_sim_hyd(P,E,param,1,ini)[0])
sim_hyd.observe(params_changed, ['x', 'y'])
widgets.VBox([fig_3,SSM0,SUZ0,SLZ0,BETA, LP, FC, PERC, K0, K1, K2, UZL, MAXBAS])
# -
# Then play with the value of the other parameters and try to get close or even lower than **RMSE = 90 ML/day**.
# We can see the complexity of manual calibration: it is difficult to find the combination of parameters that optimally fits the simulated hydrograph to the observed one.
# # Automatic model calibration
# In order to facilitate the search for an optimal parameter combination we can apply an automatic optimization algorithm. In the cell below, the function HBV_calibration calls an optimization algorithm (the genetic algorithm NSGAII [(Deb et al, 2002)](https://ieeexplore.ieee.org/document/996017) from the [Platypus library](https://platypus.readthedocs.io/en/latest/#)) runs the model 1000 times to find among different parameter combinations the set of input parameters (***ini_all*** and ***param_all***) that best match the simulated flow with the observed one, i.e. the objective of the algorithm is minimize the RMSE value. **Comment:** Please be aware that it may take a few seconds to run the cell below (while running you will see **In [*]** at the upper left-hand of the cell and when the computation has finished you will see a number between the brackets)
cal_objective = 'all' # the objective is to minimize RMSE considering ALL the hydrograph
iterations = 1000 # number of iterations
results_all,solution_all, RMSE_all = HBV_calibration(P,E,1,area, Q_obs, cal_objective,iterations)
ini_all = solution_all[0][0:3] # Initial conditions
param_all = solution_all[0][3:13] # Model parameters
Q_sim_all,[SM,UZ,LZ],[EA,R,RL,Q0,Q1] = HBV_sim(P,E,param_all,1,ini_all,area) # Simulation using the optimal set of parameters
# #### Plot the automatically calibrated hydrograph vs the observed
# +
Case = 1
Q_sim_all,[SM,UZ,LZ],[EA,R,RL,Q0,Q1] = HBV_sim(P,E,param_all,Case,ini_all,area)
x_ax_4 = Axis(label='date', scale=x_sc_2)
y_ax_4 = Axis(label='ML/day', scale=y_sc_2, orientation='vertical')
fig_4 = plt.figure(title = 'RMSE = '+str("%.2f ML/day" % RMSE_all[0]), axes=[x_ax_4, y_ax_4],scales={'x': x_sc_2, 'y': y_sc_2},
layout={'min_width': '900px', 'max_height': '250px'}, animation_duration=1000,
fig_margin = {'top':0, 'bottom':40, 'left':60, 'right':0})
obs_hyd = plt.plot(Q_obs_date,Q_obs,colors=['black'])
sim_hyd_all = plt.plot(Q_obs_date,Q_sim_all)
fig_4
# -
# ***Comment:*** overall, the hydrograph of the automatically calibrated model seems to fit the observations quite well. However, if we look at low flow periods in particular, for example August and September 2000, we see that the model predictions tend to systematically overestimate the flows.
# ### Objective: improve the prediction of the low flows
# Imagine that our reservoir operation is very sensitive to low flows, for example because it is in low flow periods that exceptional supply management measures must be put in place. Then we would like our rainfall-runoff model to be more accurate in the prediction of the low flows, rather than the high flows. To achieve this, we can re-define the objective function as the RMSE of only the part of the hydrograph below the 50% percentile. Let’s visualise this part of the hydrograph:
# +
x_ax_5 = Axis(label='date', scale=x_sc_2)
y_ax_5 = Axis(label='ML/day', scale=y_sc_2, orientation='vertical')
fig_5 = plt.figure(title = 'Low flows: observed daily water flow < 50th percentile', axes=[x_ax_5, y_ax_5],
scales={'x': x_sc_2, 'y': y_sc_2},layout={'min_width': '1000px', 'max_height': '300px'},
animation_duration=1000)
obs_hyd_lt50 = plt.plot(x=Q_obs_date,y=Q_obs,colors=['black'])
lt50 = plt.plot(x=Q_obs_date,y=np.minimum(Q_obs/Q_obs*np.percentile(Q_obs, 50),Q_obs),opacities = [0],
fill = 'bottom',fill_opacities = [0.5])
p50 = plt.plot(x=Q_obs_date,y=Q_obs/Q_obs*np.percentile(Q_obs, 50), line_style = 'dashed')
fig_5
# -
# … and re-run the automatic calibration algorithm by using this new definition of the RMSE:
cal_objective = 'low' # the objective is to minimize RMSE considering only low flows
results_low,solution_low, RMSE_low = HBV_calibration(P,E,Case,area, Q_obs, cal_objective,iterations)
ini_low = solution_low[0][0:3] # Initial conditions
param_low = solution_low[0][3:13] # Model parameters
Q_sim_low,[SM,UZ,LZ],[EA,R,RL,Q0,Q1] = HBV_sim(P,E,param_low,Case,ini_low,area)
# #### Plot the automatically hydrograph vs the observed
# +
x_ax_6 = Axis(label='date', scale=x_sc_2)
y_ax_6 = Axis(label='ML/day', scale=y_sc_2, orientation='vertical')
fig_6 = plt.figure(title = 'RMSE = '+str("%.2f ML/day" % RMSE_low[0]), axes=[x_ax_6, y_ax_6],scales={'x': x_sc_2, 'y': y_sc_2},
layout={'min_width': '900px', 'max_height': '250px'}, animation_duration=1000,
fig_margin = {'top':0, 'bottom':40, 'left':60, 'right':0})
obs_hyd = plt.plot(Q_obs_date,Q_obs,colors=['black'])
sim_hyd_low = plt.plot(Q_obs_date,Q_sim_low)
fig_6
# -
# ***Comment:*** now the simulated hydrograph over the low flow periods August-September 2000 is much closer to the observations, but this comes at the expense of completely misrepresenting high flows!
# ### Trading-off between conflicting objectives
# As we have seen in the previous example, the goodness-of-fit (as measured by the RMSE) between simulated and observed hydrograph is quite poor when the calibration aims to improve the low flow predictions only, because improving on low flows leads to much poorer predictions of all other flows. So there is a tradeoff between the two objective functions, and we may want to investigate this tradeoff and look for a parameter set that produces a ‘sensible compromise’. We can do this by using a multi-objective optimisation algorithm [(Yapo et al, 1998)](https://www.sciencedirect.com/science/article/pii/S0022169497001078), which will find a set of parameter combinations that realise different ‘optimal’ compromises between fitting high and low flows (also called Pareto-optimal solutions) [Learn more about the Pareto optimality](https://www.youtube.com/watch?v=cT3DcuZnsGs)
cal_objective = 'double' # two objectives (RMSE of low and high flows)
population_size = 100 # number of Pareto-optimal solutions
results_double_low,results_double_high,solution_double, RMSE_double = HBV_calibration(
P,E,Case,area, Q_obs, cal_objective,iterations,population_size)
# #### Plot the interactive Pareto front
# Now select the set of parameters that produces a more "sensible compromise" between the objectives by clicking on the Pareto front points.
# + code_folding=[]
# Interactive Pareto front code (Calibration)
def update_sol_hyd(i):
ini_double = solution_double[i][0:3]
param_double = solution_double[i][3:]
Q_sim_double,[SM,UZ,LZ],[EA,R,RL,Q0,Q1] = HBV_sim(P,E,param_double,Case,ini_double,area)
RMSE = RMSE_double[i]
fig_7.title = 'RMSE = '+str("%.2f ML/day" % RMSE)
return Q_sim_double, RMSE, i
def solution_selected(change):
if pareto_front.selected == None:
pareto_front.selected = [0]
y_vals = update_sol_hyd(pareto_front.selected[0])[0]
sim_hyd_double.y = y_vals
x_sc_pf = LinearScale()
y_sc_pf = LinearScale()
x_ax_pf = Axis(label='RMSE Low flows', scale=x_sc_pf)
y_ax_pf = Axis(label='RMSE High flows', scale=y_sc_pf, orientation='vertical')
fig_pf = plt.figure(title = 'Interactive Pareto front (Calibration)', axes=[x_ax_pf, y_ax_pf],
layout={'width': '500px', 'height': '400px'}, animation_duration=1000,
fig_margin = {'top':0, 'bottom':40, 'left':60, 'right':0})
pareto_front = plt.scatter(results_double_low[:],results_double_high[:],scales={'x': x_sc_pf, 'y': y_sc_pf},
colors=['deepskyblue'], interactions={'hover':'tooltip','click': 'select'})
pareto_front.unselected_style={'opacity': 0.4}
pareto_front.selected_style={'fill': 'red', 'stroke': 'yellow', 'width': '1125px', 'height': '125px'}
def_tt = Tooltip(fields=['x', 'y'],labels=['RMSE (High flows)', 'RMSE (Low flows)'], formats=['.1f', '.1f'])
pareto_front.tooltip=def_tt
pareto_front.selected = [0]
pareto_front.observe(solution_selected,'selected')
x_sc_7 = DateScale()
y_sc_7 = LinearScale(max=1000)
x_ax_7 = Axis(scale=x_sc_7)
y_ax_7 = Axis(label='ML/day', scale=y_sc_7, orientation='vertical')
fig_7 = plt.figure(axes=[x_ax_7, y_ax_7], layout={'min_width': '900px', 'max_height': '250px'}, animation_duration=1000,
scales={'x': x_sc_7, 'y': y_sc_7},fig_margin = {'top':0, 'bottom':40, 'left':60, 'right':0})
obs_hyd = plt.plot(Q_obs_date,Q_obs,colors=['black'])
sim_hyd_double = plt.plot(Q_obs_date,update_sol_hyd(pareto_front.selected[0])[0])
sim_hyd_double.observe(solution_selected, ['x', 'y'])
plt.VBox([fig_pf,fig_7])
# -
# # Evaluation of the calibrated model against new data
# The calibration results that we have looked at so far, and in particular the values of the RMSE over the high and low flows, were based on the model simulations for the year 2000, that is, the same year that was used to calibrate the model in the first place. But how would the model perform when presented with new data, for instance those of the following year? To answer this question, we can run model simulations using the previously selected calibration (represented with a cross) against the forcing data of 2001, and calculate the RMSE values for this new year.
# + code_folding=[]
# Interactive Pareto front code (Validation)
val_year = cal_year + 1 # following year
results_double_low_val = np.zeros(population_size); results_double_high_val = np.zeros(population_size)
RMSE_double_val = np.zeros(population_size)
clim_date_val, E_val, P_val, T_val = Climate_data(val_year)
Q_obs_date_val, Q_obs_val = Flow_data(val_year)
P_val = P_val*2
Q_obs_val = Q_obs_val*2
for i in range(population_size):
Q_sim_double_val,[SM,UZ,LZ],[EA,R,RL,Q0,Q1] = HBV_sim(P_val,E_val,solution_double[i][3:],
Case,solution_double[i][0:3],area)
high_flow_indexes = [Q_obs_val > np.percentile(Q_obs_val,50)]
Q_obs_high_val = Q_obs_val[high_flow_indexes]
Q_sim_double_high_val = Q_sim_double_val[high_flow_indexes]
low_flow_indexes = [Q_obs_val < np.percentile(Q_obs_val,50)]
Q_obs_low_val = Q_obs_val[low_flow_indexes]
Q_sim_double_low_val = Q_sim_double_val[low_flow_indexes]
results_double_low_val[i] = np.sqrt(((Q_sim_double_low_val - Q_obs_low_val) ** 2).mean())
results_double_high_val[i] = np.sqrt(((Q_sim_double_high_val - Q_obs_high_val) ** 2).mean())
RMSE_double_val[i] = np.sqrt(((Q_sim_double_val - Q_obs_val) ** 2).mean())
def update_sol_hyd_val(i):
ini_double = solution_double[i][0:3]
param_double = solution_double[i][3:]
Q_sim_double_val,[SM,UZ,LZ],[EA,R,RL,Q0,Q1] = HBV_sim(P_val,E_val,param_double,Case,ini_double,area)
RMSE_val = RMSE_double_val[i]
fig_8.title = 'RMSE = '+str("%.2f" % RMSE_val)
return Q_sim_double_val, RMSE_val, i
def solution_selected_val(change):
if pareto_front_val.selected == None:
pareto_front_val.selected = [0]
y_vals = update_sol_hyd_val(pareto_front_val.selected[0])[0]
sim_hyd_double_val.y = y_vals
x_sc_pf_val = LinearScale()
y_sc_pf_val = LinearScale()
x_ax_pf_val = Axis(label='RMSE Low flows', scale=x_sc_pf_val)
y_ax_pf_val = Axis(label='RMSE High flows', scale=y_sc_pf_val, orientation='vertical')
fig_pf_val = plt.figure(title = 'Interactive Pareto front (Validation)', axes=[x_ax_pf_val, y_ax_pf_val],
layout={'width': '500px', 'height': '400px'}, animation_duration=1000,
fig_margin = {'top':0, 'bottom':40, 'left':60, 'right':0})
pareto_front_selected = plt.scatter([results_double_low_val[pareto_front.selected[0]]],
[results_double_high_val[pareto_front.selected[0]]],
marker = 'cross',scales={'x': x_sc_pf_val, 'y': y_sc_pf_val},
colors=['black'], interactions={'hover':'tooltip','click': 'select'})
pareto_front_val = plt.scatter(results_double_low_val[:],results_double_high_val[:],
scales={'x': x_sc_pf_val, 'y': y_sc_pf_val}, colors=['deepskyblue'],
interactions={'hover':'tooltip','click': 'select'})
pareto_front_val.unselected_style={'opacity': 0.4}
pareto_front_val.selected_style={'fill': 'red', 'stroke': 'yellow', 'width': '1125px', 'height': '125px'}
def_tt_val = Tooltip(fields=['x', 'y'],labels=['RMSE (High flows)', 'RMSE (Low flows)'], formats=['.1f', '.1f'])
pareto_front_val.tooltip=def_tt_val
pareto_front_val.selected = [pareto_front.selected[0]]
pareto_front_val.observe(solution_selected_val,'selected')
x_sc_8 = DateScale()
y_sc_8 = LinearScale(max=1000)
x_ax_8 = Axis(scale=x_sc_8)
y_ax_8 = Axis(label='ML/day', scale=y_sc_8, orientation='vertical')
fig_8 = plt.figure(axes=[x_ax_8, y_ax_8], layout={'min_width': '900px', 'max_height': '250px'}, animation_duration=1000,
scales={'x': x_sc_8, 'y': y_sc_8},fig_margin = {'top':0, 'bottom':40, 'left':60, 'right':0})
obs_hyd_val = plt.plot(Q_obs_date_val,Q_obs_val,colors=['black'])
sim_hyd_double_val = plt.plot(Q_obs_date_val,update_sol_hyd_val(pareto_front_val.selected[0])[0])
sim_hyd_double_val.observe(solution_selected_val, ['x', 'y'])
plt.VBox([fig_pf_val,fig_8])
# -
# We can see that the previously selected calibration for 2000 (represented by a cross) does not produce the same results when applied to 2001. The RMSE may be lower than before but do you see a good fit between the observed and the simulated hydrographs? Can you find a different point in the evaluation Pareto front that produces a "sensible compromise" between the objectives?
# ### References
#
# <NAME>. (1992) The HBV model - its structure and applications. SMHI Reports RH, No. 4, Norrköping.
#
# <NAME>. et al (2002) A fast and elitist multiobjective genetic algorithm: NSGA-II, IEEE Transactions on Evolutionary Computation, 6(2), 182-197, doi:10.1109/4235.996017.
#
# <NAME> al (1998) Multi‐objective global optimization for hydrologic models, Journal of Hydrology, 204, 83–97.
# #### Let's go to the next section!: [3.a. Recursive decisions and multi-objective optimisation: optimising reservoir release scheduling under conflicting objectives](3.a.%20Recursive%20decisions%20and%20multi-objective%20optimisation.ipynb)
| iRONS/Notebooks/A - Knowledge transfer/2.a. Calibration and evaluation of a rainfall-runoff model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
#export
"""
This is for all short utilities that has the boilerplate feeling. Conversion clis
might feel they have different styles, as :class:`toFloat` converts object iterator to
float iterator, while :class:`toPIL` converts single image url to single PIL image,
whereas :class:`toSum` converts float iterator into a single float value.
The general convention is, if the intended operation sounds simple (convert to floats,
strings, types, ...), then most likely it will convert iterator to iterator, as you
can always use the function directly if you only want to apply it on 1 object.
If it sounds complicated (convert to PIL image, tensor, ...) then most likely it will
convert object to object. Lastly, there are some that just feels right to input
an iterator and output a single object (like getting max, min, std, mean values)."""
from k1lib.cli.init import patchDefaultDelim, BaseCli, Table, T
import k1lib.cli as cli, numbers, torch, numpy as np
from typing import overload, Iterator, Any, List, Set, Union
import k1lib
__all__ = ["size", "shape", "item", "identity", "iden",
"toStr", "join", "toNumpy", "toTensor",
"toList", "wrapList", "toSet", "toIter", "toRange", "toType",
"equals", "reverse", "ignore",
"toSum", "toAvg", "toMean", "toMax", "toMin", "toPIL",
"toBin", "toIdx",
"lengths", "headerIdx", "deref", "bindec"]
#export
settings = k1lib.settings.cli
#export
def exploreSize(it):
"""Returns first element and length of array"""
if isinstance(it, str): raise TypeError("Just here to terminate shape()")
sentinel = object(); it = iter(it)
o = next(it, sentinel); count = 1
if o is sentinel: return None, 0
try:
while True: next(it); count += 1
except StopIteration: pass
return o, count
#export
class size(BaseCli):
def __init__(self, idx=None):
"""Returns number of rows and columns in the input.
Example::
# returns (3, 2)
[[2, 3], [4, 5, 6], [3]] | size()
# returns 3
[[2, 3], [4, 5, 6], [3]] | size(0)
# returns 2
[[2, 3], [4, 5, 6], [3]] | size(1)
# returns (2, 0)
[[], [2, 3]] | size()
# returns (3,)
[2, 3, 5] | size()
# returns 3
[2, 3, 5] | size(0)
# returns (3, 2, 2)
[[[2, 1], [0, 6, 7]], 3, 5] | size()
# returns (1,) and not (1, 3)
["abc"] | size()
# returns (1, 2, 3)
[torch.randn(2, 3)] | size()
# returns (2, 3, 5)
size()(np.random.randn(2, 3, 5))
There's also :class:`lengths`, which is sort of a simplified/faster version of
this, but only use it if you are sure that ``len(it)`` can be called.
If encounter PyTorch tensors or Numpy arrays, then this will just get the shape
instead of actually looping over them.
:param idx: if idx is None return (rows, columns). If 0 or 1, then rows or
columns"""
super().__init__(); self.idx = idx
def __ror__(self, it:Iterator[str]):
if self.idx is None:
answer = []
try:
while True:
if isinstance(it, (torch.Tensor, np.ndarray)):
return tuple(answer + list(it.shape))
it, s = exploreSize(it); answer.append(s)
except TypeError: pass
return tuple(answer)
else:
return exploreSize(it | cli.item(self.idx))[1]
shape = size
assert [[2, 3], [4, 5, 6], [3]] | size() == (3, 2)
assert [[2, 3], [4, 5, 6], [3]] | size(0) == 3
assert [[2, 3], [4, 5, 6], [3]] | size(1) == 2
assert [[], [2, 3]] | size() == (2, 0)
assert [2, 3, 5] | size() == (3,)
assert [2, 3, 5] | size(0) == 3
assert torch.randn(3, 4) | size() == (3, 4)
assert [[[2, 1], [0, 6, 7]], 3, 5] | size() == (3, 2, 2)
assert ["abc"] | size() == (1,)
assert [torch.randn(2, 3)] | size() == (1, 2, 3)
assert size()(np.random.randn(2, 3, 5)) == (2, 3, 5)
#export
noFill = object()
class item(BaseCli):
def __init__(self, amt:int=1, fill=noFill):
"""Returns the first row.
Example::
# returns 0
iter(range(5)) | item()
# returns torch.Size([5])
torch.randn(3,4,5) | item(2) | shape()
# returns 3
[] | item(fill=3)
:param amt: how many times do you want to call item() back to back?
:param fill: if iterator length is 0, return this"""
self.amt = amt; self.fill = fill
self.fillP = [fill] if fill != noFill else [] # preprocessed, to be faster
def __ror__(self, it:Iterator[str]):
if self.amt != 1:
return it | cli.serial(*(item(fill=self.fill) for _ in range(self.amt)))
return next(iter(it), *self.fillP)
assert iter(range(5)) | item() == 0
assert torch.randn(3,4,5) | item(2) | shape() == torch.Size([5])
assert [] | item(fill=3) == 3
#export
class identity(BaseCli):
"""Yields whatever the input is. Useful for multiple streams.
Example::
# returns range(5)
range(5) | identity()"""
def __ror__(self, it:Iterator[Any]): return it
iden = identity
assert range(5) | identity() == range(5)
#export
class toStr(BaseCli):
def __init__(self, column:int=None):
"""Converts every line to a string.
Example::
# returns ['2', 'a']
[2, "a"] | toStr() | deref()
# returns [[2, 'a'], [3, '5']]
assert [[2, "a"], [3, 5]] | toStr(1) | deref()"""
super().__init__(); self.column = column
def __ror__(self, it:Iterator[str]):
c = self.column
if c is None:
for line in it: yield str(line)
else:
for row in it:
yield [e if i != c else str(e) for i, e in enumerate(row)]
assert [2, "a"] | toStr() | cli.deref() == ['2', 'a']
assert [[2, "a"], [3, 5]] | toStr(1) | cli.deref() == [[2, 'a'], [3, '5']]
#export
class join(BaseCli):
def __init__(self, delim:str=None):
r"""Merges all strings into 1, with `delim` in the middle. Basically
:meth:`str.join`. Example::
# returns '2\na'
[2, "a"] | join("\n")"""
super().__init__(); self.delim = patchDefaultDelim(delim)
def __ror__(self, it:Iterator[str]):
return self.delim.join(it | toStr())
assert [2, "a"] | join("\n") == '2\na'
#export
class toNumpy(BaseCli):
"""Converts generator to numpy array. Essentially ``np.array(list(it))``"""
def __ror__(self, it:Iterator[float]) -> np.array:
return np.array(list(it))
class toTensor(BaseCli):
def __init__(self, dtype=torch.float32):
"""Converts generator to :class:`torch.Tensor`. Essentially
``torch.tensor(list(it))``.
Also checks if input is a PIL Image. If yes, turn it into a :class:`torch.Tensor`
and return."""
self.dtype = dtype
def __ror__(self, it:Iterator[float]) -> torch.Tensor:
try:
import PIL; pic=it
if isinstance(pic, PIL.Image.Image): # stolen from torchvision ToTensor transform
mode_to_nptype = {'I': np.int32, 'I;16': np.int16, 'F': np.float32}
img = torch.from_numpy(np.array(pic, mode_to_nptype.get(pic.mode, np.uint8), copy=True))
if pic.mode == '1': img = 255 * img
img = img.view(pic.size[1], pic.size[0], len(pic.getbands()))
return img.permute((2, 0, 1)).contiguous().to(self.dtype) # put it from HWC to CHW format
except: pass
return torch.tensor(list(it)).to(self.dtype)
assert (range(3) | toTensor(torch.float64)).dtype == torch.float64
#export
class toList(BaseCli):
"""Converts generator to list. :class:`list` would do the
same, but this is just to maintain the style"""
def __ror__(self, it:Iterator[Any]) -> List[Any]:
return list(it)
class wrapList(BaseCli):
"""Wraps inputs inside a list. There's a more advanced cli tool
built from this, which is :meth:`~k1lib.cli.structural.unsqueeze`."""
def __ror__(self, it:T) -> List[T]:
return [it]
class toSet(BaseCli):
"""Converts generator to set. :class:`set` would do the
same, but this is just to maintain the style"""
def __ror__(self, it:Iterator[T]) -> Set[T]:
return set(it)
class toIter(BaseCli):
"""Converts object to iterator. `iter()` would do the
same, but this is just to maintain the style"""
def __ror__(self, it:List[T]) -> Iterator[T]:
return iter(it)
class toRange(BaseCli):
"""Returns iter(range(len(it))), effectively"""
def __ror__(self, it:Iterator[Any]) -> Iterator[int]:
for i, _ in enumerate(it): yield i
#export
class toType(BaseCli):
"""Converts object to its type.
Example::
# returns [int, float, str, torch.Tensor]
[2, 3.5, "ah", torch.randn(2, 3)] | toType() | deref()"""
def __ror__(self, it:Iterator[T]) -> Iterator[type]:
for e in it: yield type(e)
assert [2, 3.5, "ah", torch.randn(2, 3)] | toType() | cli.deref() == [int, float, str, torch.Tensor]
#export
class _EarlyExp(Exception): pass
class equals:
"""Checks if all incoming columns/streams are identical"""
def __ror__(self, streams:Iterator[Iterator[str]]):
streams = list(streams)
for row in zip(*streams):
sampleElem = row[0]
try:
for elem in row:
if sampleElem != elem: yield False; raise _EarlyExp()
yield True
except _EarlyExp: pass
#export
class reverse(BaseCli):
"""Reverses incoming list.
Example::
# returns [3, 5, 2]
[2, 5, 3] | reverse() | deref()"""
def __ror__(self, it:Iterator[str]) -> List[str]:
return reversed(list(it))
assert [2, 5, 3] | reverse() | cli.deref() == [3, 5, 2]
#export
class ignore(BaseCli):
r"""Just loops through everything, ignoring the output.
Example::
# will just return an iterator, and not print anything
[2, 3] | apply(lambda x: print(x))
# will prints "2\n3"
[2, 3] | apply(lambda x: print(x)) | ignore()"""
def __ror__(self, it:Iterator[Any]):
for _ in it: pass
with k1lib.captureStdout() as out:
[2, 3] | cli.apply(lambda x: print(x))
assert len(out) == 0
with k1lib.captureStdout() as out:
[2, 3] | cli.apply(lambda x: print(x)) | ignore()
assert out == ["2", "3"]
#export
class toSum(BaseCli):
"""Calculates the sum of list of numbers. Can pipe in :class:`torch.Tensor`.
Example::
# returns 45
range(10) | toSum()"""
def __ror__(self, it:Iterator[float]):
if isinstance(it, torch.Tensor): return it.sum()
return sum(it)
class toAvg(BaseCli):
"""Calculates average of list of numbers. Can pipe in :class:`torch.Tensor`.
Example::
# returns 4.5
range(10) | toAvg()
# returns nan
[] | toAvg()"""
def __ror__(self, it:Iterator[float]):
if isinstance(it, torch.Tensor): return it.mean()
s = 0; i = -1
for i, v in enumerate(it): s += v
i += 1
if not k1lib.settings.cli.strict and i == 0: return float("nan")
return s / i
toMean = toAvg
assert range(10) | toSum() == 45
assert range(10) | toAvg() == 4.5
assert id([] | toAvg()) == id(float("nan"))
#export
class toMax(BaseCli):
"""Calculates the max of a bunch of numbers. Can pipe in :class:`torch.Tensor`.
Example::
# returns 6
[2, 5, 6, 1, 2] | toMax()"""
def __ror__(self, it:Iterator[float]) -> float:
if isinstance(it, torch.Tensor): return it.max()
return max(it)
class toMin(BaseCli):
"""Calculates the min of a bunch of numbers. Can pipe in :class:`torch.Tensor`.
Example::
# returns 1
[2, 5, 6, 1, 2] | toMin()"""
def __ror__(self, it:Iterator[float]) -> float:
if isinstance(it, torch.Tensor): return it.min()
return min(it)
assert [2, 5, 6, 1, 2] | toMax() == 6
assert [2, 5, 6, 1, 2] | toMin() == 1
#export
class toPIL(BaseCli):
def __init__(self):
"""Converts a path to a PIL image.
Example::
ls(".") | toPIL().all() | item() # get first image"""
import PIL; self.PIL = PIL
def __ror__(self, path) -> "PIL.Image.Image":
return self.PIL.Image.open(path)
#export
class toBin(BaseCli):
"""Converts integer to binary string.
Example::
# returns "101"
5 | toBin()"""
def __ror__(self, it):
return bin(int(it))[2:]
assert 5 | toBin() == "101"
#export
class toIdx(BaseCli):
def __init__(self, chars:str):
"""Get index of characters according to a reference.
Example::
# returns [1, 4, 4, 8]
"#&&*" | toIdx("!#$%&'()*+") | deref()"""
self.chars = {v:k for k, v in enumerate(chars)}
def __ror__(self, it):
chars = self.chars
for e in it: yield chars[e]
assert "#&&*" | toIdx("!#$%&'()*+") | cli.deref() == [1, 4, 4, 8]
#export
class lengths(BaseCli):
"""Returns the lengths of each element.
Example::
[range(5), range(10)] | lengths() == [5, 10]
This is a simpler (and faster!) version of :class:`shape`. It assumes each element
can be called with ``len(x)``, while :class:`shape` iterates through every elements
to get the length, and thus is much slower."""
def __ror__(self, it:Iterator[List[Any]]) -> Iterator[int]:
for e in it: yield len(e)
assert [range(5), range(10)] | lengths() | cli.deref() == [5, 10]
#export
def headerIdx():
"""Cuts out first line, put an index column next to it, and prints it
out. Useful when you want to know what your column's index is to cut it
out. Also sets the context variable "header", in case you need it later.
Example::
# returns [[0, 'a'], [1, 'b'], [2, 'c']]
["abc"] | headerIdx() | deref()"""
return item() | cli.wrapList() | cli.transpose() | cli.insertIdColumn(True)
assert ["abc"] | headerIdx() | cli.deref() == [[0, 'a'], [1, 'b'], [2, 'c']]
#export
settings.atomic.add("deref", (numbers.Number, np.number, str, dict, bool, bytes, torch.nn.Module), "used by deref")
Tensor = torch.Tensor; atomic = settings.atomic
class inv_dereference(BaseCli):
def __init__(self, ignoreTensors=False):
"""Kinda the inverse to :class:`dereference`"""
super().__init__(); self.ignoreTensors = ignoreTensors
def __ror__(self, it:Iterator[Any]) -> List[Any]:
for e in it:
if e is None or isinstance(e, atomic.deref): yield e
elif isinstance(e, Tensor):
if not self.ignoreTensors and len(e.shape) == 0: yield e.item()
else: yield e
else:
try: yield e | self
except: yield e
#export
class deref(BaseCli):
def __init__(self, maxDepth=float("inf"), ignoreTensors=True):
"""Recursively converts any iterator into a list. Only :class:`str`,
:class:`numbers.Number` and :class:`~torch.nn.Module` are not converted. Example::
# returns something like "<range_iterator at 0x7fa8c52ca870>"
iter(range(5))
# returns [0, 1, 2, 3, 4]
iter(range(5)) | deref()
# returns [2, 3], yieldSentinel stops things early
[2, 3, yieldSentinel, 6] | deref()
You can also specify a ``maxDepth``::
# returns something like "<list_iterator at 0x7f810cf0fdc0>"
iter([range(3)]) | deref(0)
# returns [range(3)]
iter([range(3)]) | deref(1)
# returns [[0, 1, 2]]
iter([range(3)]) | deref(2)
There are a few classes/types that are considered atomic, and :class:`deref`
will never try to iterate over it. If you wish to change it, do something like::
settings.cli.atomic.deref = (int, float, ...)
.. warning::
Can work well with PyTorch Tensors, but not Numpy arrays as they screw things up
with the __ror__ operator, so do torch.from_numpy(...) first. Don't worry about
unnecessary copying, as numpy and torch both utilizes the buffer protocol.
:param maxDepth: maximum depth to dereference. Starts at 0 for not doing anything
at all
:param ignoreTensors: if True, then don't loop over :class:`torch.Tensor`
internals"""
super().__init__(); self.ignoreTensors = ignoreTensors
self.maxDepth = maxDepth; self.depth = 0
def __ror__(self, it:Iterator[T]) -> List[T]:
ignoreTensors = self.ignoreTensors
if self.depth >= self.maxDepth: return it
elif isinstance(it, atomic.deref): return it
elif isinstance(it, Tensor):
if ignoreTensors: return it
if len(it.shape) == 0: return it.item()
try: iter(it)
except: return it
self.depth += 1; answer = []
for e in it:
if e is cli.yieldSentinel: return answer
answer.append(self.__ror__(e))
self.depth -= 1; return answer
def __invert__(self) -> BaseCli:
"""Returns a :class:`~k1lib.cli.init.BaseCli` that makes
everything an iterator. Not entirely sure when this comes in handy, but it's
there."""
return inv_dereference(self.ignoreTensors)
import numpy as np, numbers, torch
a = torch.linspace(0, 10, 50) | deref(ignoreTensors=False)
b = torch.from_numpy(np.linspace(0, 10, 50)) | deref(ignoreTensors=False)
assert torch.allclose(torch.tensor(b), torch.tensor(a))
assert iter([range(3)]) | deref(1) == [range(3)]
assert iter([range(3)]) | deref(2) == [[0, 1, 2]]
assert torch.randn(2, 3) | deref(ignoreTensors=False) | ~deref() | deref() | shape() == (2, 3)
assert [2, 3, cli.yieldSentinel, 6] | deref() == [2, 3]
with k1lib.settings.cli.atomic.context(deref=(int, float, ...)): assert k1lib.settings.cli.atomic.deref == (int, float, ...)
#export
class bindec(BaseCli):
def __init__(self, cats:List[Any], f=None):
"""Binary decodes the input.
Example::
# returns ['a', 'c']
5 | bindec("abcdef")
# returns 'a,c'
5 | bindec("abcdef", join(","))
:param cats: categories
:param f: transformation function of the selected elements. Defaulted to :class:`toList`, but others like :class:`join` is useful too"""
self.cats = cats; self.f = f or toList()
def __ror__(self, it):
it = bin(int(it))[2:][::-1]
return (e for i, e in zip(it, self.cats) if i == '1') | self.f
assert 5 | bindec("abcdef") == ['a', 'c']
assert 5 | bindec("abcdef", join(",")) == "a,c"
# !../../export.py cli/utils
| k1lib/cli/utils.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lesson 2
#
# This is the second in a series of lessons related to astronomy data.
#
# As a running example, we are replicating parts of the analysis in a recent paper, "[Off the beaten path: Gaia reveals GD-1 stars outside of the main stream](https://arxiv.org/abs/1805.00425)" by <NAME> and <NAME>.
#
# In the first notebook, we wrote ADQL queries and used them to select and download data from the Gaia server.
#
# In this notebook, we'll pick up where we left off and write a query to select stars from the region of the sky where we expect GD-1 to be.
# We'll start with an example that does a "cone search"; that is, it selects stars that appear in a circular region of the sky.
#
# Then, to select stars in the vicinity of GD-1, we'll:
#
# * Use `Quantity` objects to represent measurements with units.
#
# * Use the `Gala` library to convert coordinates from one frame to another.
#
# * Use the ADQL keywords `POLYGON`, `CONTAINS`, and `POINT` to select stars that fall within a polygonal region.
#
# * Submit a query and download the results.
#
# * Store the results in a FITS file.
#
# After completing this lesson, you should be able to
#
# * Use Python string formatting to compose more complex ADQL queries.
#
# * Work with coordinates and other quantities that have units.
#
# * Download the results of a query and store them in a file.
# ## Installing libraries
#
# If you are running this notebook on Colab, you can run the following cell to install Astroquery and a the other libraries we'll use.
#
# If you are running this notebook on your own computer, you might have to install these libraries yourself.
#
# If you are using this notebook as part of a Carpentries workshop, you should have received setup instructions.
#
# TODO: Add a link to the instructions.
#
# +
# If we're running on Colab, install libraries
import sys
IN_COLAB = 'google.colab' in sys.modules
if IN_COLAB:
# !pip install astroquery astro-gala pyia
# -
# ## Selecting a region
# One of the most common ways to restrict a query is to select stars in a particular region of the sky.
#
# For example, here's a query from the [Gaia archive documentation](https://gea.esac.esa.int/archive-help/adql/examples/index.html) that selects "all the objects ... in a circular region centered at (266.41683, -29.00781) with a search radius of 5 arcmin (0.08333 deg)."
query = """
SELECT
TOP 10 source_id
FROM gaiadr2.gaia_source
WHERE 1=CONTAINS(
POINT(ra, dec),
CIRCLE(266.41683, -29.00781, 0.08333333))
"""
# This query uses three keywords that are specific to ADQL (not SQL):
#
# * `POINT`: a location in [ICRS coordinates](https://en.wikipedia.org/wiki/International_Celestial_Reference_System), specified in degrees of right ascension and declination.
#
# * `CIRCLE`: a circle where the first two values are the coordinates of the center and the third is the radius in degrees.
#
# * `CONTAINS`: a function that returns `1` if a `POINT` is contained in a shape and `0` otherwise.
#
# Here is the [documentation of `CONTAINS`](http://www.ivoa.net/documents/ADQL/20180112/PR-ADQL-2.1-20180112.html#tth_sEc4.2.12).
#
# A query like this is called a cone search because it selects stars in a cone.
#
# Here's how we run it.
# +
from astroquery.gaia import Gaia
job = Gaia.launch_job(query)
result = job.get_results()
result
# -
# **Exercise:** When you are debugging queries like this, you can use `TOP` to limit the size of the results, but then you still don't know how big the results will be.
#
# An alternative is to use `COUNT`, which asks for the number of rows that would be selected, but it does not return them.
#
# In the previous query, replace `TOP 10 source_id` with `COUNT(source_id)` and run the query again. How many stars has Gaia identified in the cone we searched?
# ## Getting GD-1 Data
#
# From the Price-Whelan and Bonaca paper, we will try to reproduce Figure 1, which includes this representation of stars likely to belong to GD-1:
#
# <img src="https://github.com/datacarpentry/astronomy-python/raw/gh-pages/fig/gd1-4.png">
# Along the axis of right ascension ($\phi_1$) the figure extends from -100 to 20 degrees.
#
# Along the axis of declination ($\phi_2$) the figure extends from about -8 to 4 degrees.
#
# Ideally, we would select all stars from this rectangle, but there are more than 10 million of them, so
#
# * That would be difficult to work with,
#
# * As anonymous users, we are limited to 3 million rows in a single query, and
#
# * While we are developing and testing code, it will be faster to work with a smaller dataset.
#
# So we'll start by selecting stars in a smaller rectangle, from -55 to -45 degrees right ascension and -8 to 4 degrees of declination.
#
# But first we let's see how to represent quantities with units like degrees.
# ## Working with coordinates
#
# Coordinates are physical quantities, which means that they have two parts, a value and a unit.
#
# For example, the coordinate $30^{\circ}$ has value 30 and its units are degrees.
#
# Until recently, most scientific computation was done with values only; units were left out of the program altogether, [often with disastrous results](https://en.wikipedia.org/wiki/Mars_Climate_Orbiter#Cause_of_failure).
#
# Astropy provides tools for including units explicitly in computations, which makes it possible to detect errors before they cause disasters.
#
# To use Astropy units, we import them like this:
# +
import astropy.units as u
u
# -
# `u` is an object that contains most common units and all SI units.
#
# You can use `dir` to list them, but you should also [read the documentation](https://docs.astropy.org/en/stable/units/).
dir(u)
# To create a quantity, we multiply a value by a unit.
coord = 30 * u.deg
type(coord)
# The result is a `Quantity` object.
#
# Jupyter knows how to display `Quantities` like this:
coord
# ## Selecting a rectangle
#
# Now we'll select a rectangle from -55 to -45 degrees right ascension and -8 to 4 degrees of declination.
#
# We'll define variables to contain these limits.
phi1_min = -55
phi1_max = -45
phi2_min = -8
phi2_max = 4
# To represent a rectangle, we'll use two lists of coordinates and multiply by their units.
phi1_rect = [phi1_min, phi1_min, phi1_max, phi1_max] * u.deg
phi2_rect = [phi2_min, phi2_max, phi2_max, phi2_min] * u.deg
# `phi1_rect` and `phi2_rect` represent the coordinates of the corners of a rectangle.
#
# But they are in "[a Heliocentric spherical coordinate system defined by the orbit of the GD1 stream](https://gala-astro.readthedocs.io/en/latest/_modules/gala/coordinates/gd1.html)"
#
# In order to use them in a Gaia query, we have to convert them to [International Celestial Reference System](https://en.wikipedia.org/wiki/International_Celestial_Reference_System) (ICRS) coordinates. We can do that by storing the coordinates in a `GD1Koposov10` object provided by [Gala](https://gala-astro.readthedocs.io/en/latest/coordinates/).
# +
import gala.coordinates as gc
corners = gc.GD1Koposov10(phi1=phi1_rect, phi2=phi2_rect)
type(corners)
# -
# We can display the result like this:
corners
# Now we can use `transform_to` to convert to ICRS coordinates.
# +
import astropy.coordinates as coord
corners_icrs = corners.transform_to(coord.ICRS)
type(corners_icrs)
# -
# The result is an `ICRS` object.
corners_icrs
# Notice that a rectangle in one coordinate system is not necessarily a rectangle in another. In this example, the result is a polygon.
# ## Selecting a polygon
#
# In order to use this polygon as part of an ADQL query, we have to convert it to a string with a comma-separated list of coordinates, as in this example:
#
# ```
# """
# POLYGON(143.65, 20.98,
# 134.46, 26.39,
# 140.58, 34.85,
# 150.16, 29.01)
# """
# ```
# `corners_icrs` behaves like a list, so we can use a `for` loop to iterate through the points.
for point in corners_icrs:
print(point)
# From that, we can select the coordinates `ra` and `dec`:
for point in corners_icrs:
print(point.ra, point.dec)
# The results are quantities with units, but if we select the `value` part, we get a dimensionless floating-point number.
for point in corners_icrs:
print(point.ra.value, point.dec.value)
# We can use string `format` to convert these numbers to strings.
# +
point_base = "{point.ra.value}, {point.dec.value}"
t = [point_base.format(point=point)
for point in corners_icrs]
t
# -
# The result is a list of strings, which we can join into a single string using `join`.
point_list = ', '.join(t)
point_list
# Notice that we invoke `join` on a string and pass the list as an argument.
#
# Before we can assemble the query, we need `columns` again (as we saw in the previous notebook).
columns = 'source_id, ra, dec, pmra, pmdec, parallax, parallax_error, radial_velocity'
# Here's the base for the query, with format specifiers for `columns` and `point_list`.
query_base = """SELECT {columns}
FROM gaiadr2.gaia_source
WHERE parallax < 1
AND bp_rp BETWEEN -0.75 AND 2
AND 1 = CONTAINS(POINT(ra, dec),
POLYGON({point_list}))
"""
# And here's the result:
query = query_base.format(columns=columns,
point_list=point_list)
print(query)
# As always, we should take a minute to proof-read the query before we launch it.
#
# The result will be bigger than our previous queries, so it will take a little longer.
job = Gaia.launch_job_async(query)
print(job)
# Here are the results.
results = job.get_results()
len(results)
# There are more than 100,000 stars in this polygon, but that's a manageable size to work with.
# ## Saving results
#
# This is the set of stars we'll work with in the next step. But since we have a substantial dataset now, this is a good time to save it.
#
# Storing the data in a file means we can shut down this notebook and pick up where we left off without running the previous query again.
#
# Astropy `Table` objects provide `write`, which writes the table to disk.
filename = 'gd1_results.fits'
results.write(filename, overwrite=True)
# Because the filename ends with `fits`, the table is written in the [FITS format](https://en.wikipedia.org/wiki/FITS), which preserves the metadata associated with the table.
#
# If the file already exists, the `overwrite` argument causes it to be overwritten.
#
# To see how big the file is, we can use `ls` with the `-lh` option, which prints information about the file including its size in human-readable form.
# !ls -lh gd1_results.fits
# The file is about 8.6 MB.
# ## Summary
#
# In this notebook, we composed more complex queries to select stars within a polygonal region of the sky. Then we downloaded the results and saved them in a FITS file.
#
# In the next notebook, we'll reload the data from this file and replicate the next step in the analysis, using proper motion to identify stars likely to be in GD-1.
# ## Best practices
#
# * For measurements with units, use `Quantity` objects that represent units explicitly and check for errors.
#
# * Use the `format` function to compose queries; it is often faster and less error-prone.
#
# * Develop queries incrementally: start with something simple, test it, and add a little bit at a time.
#
# * Once you have a query working, save the data in a local file. If you shut down the notebook and come back to it later, you can reload the file; you don't have to run the query again.
# + active=""
#
| _build/jupyter_execute/AstronomicalData/_build/jupyter_execute/02_coords.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:PythonData] *
# language: python
# name: conda-env-PythonData-py
# ---
# # VacationPy
# ----
#
# #### Note
# * Keep an eye on your API usage. Use https://developers.google.com/maps/reporting/gmp-reporting as reference for how to monitor your usage and billing.
#
# * Instructions have been included for each segment. You do not have to follow them exactly, but they are included to help you think through the steps.
# +
# Dependencies and Setup
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import requests
import gmaps
import os
# Import API key
from api_keys import g_key
# -
# ### Store Part I results into DataFrame
# * Load the csv exported in Part I to a DataFrame
# Load the csv file from part 1
file = "../output_data/Clean_City_Data.csv"
clean = pd.read_csv(file)
# clean_df has a 'unnamed' column, so remove it so it looks better
#clean_df.head()
cleancities = clean.drop(columns=["Unnamed: 0"])
cleancities.head()
# ### Humidity Heatmap
# * Configure gmaps.
# * Use the Lat and Lng as locations and Humidity as the weight.
# * Add Heatmap layer to map.
# +
# Configure gmaps
gmaps.configure(api_key = g_key)
# Initiate variables
locations = cleancities[['Lat', 'Lng']].astype(float)
humidity = cleancities['Humidity'].astype(float)
# +
# Add some specifications to heatmap
heatmap_specs = {
'width': '1000px',
'height': '500px',
'margin': '0 auto 0 auto'
}
# Create map
fig = gmaps.figure(layout=heatmap_specs, zoom_level=2, center=(0,0))
# Add layer details
heat_layer = gmaps.heatmap_layer(locations,
weights=humidity,
dissipating=False,
max_intensity=100,
point_radius=1)
fig.add_layer(heat_layer)
plt.savefig("../Images/humidty_heatmap.png")
plt.show()
fig
# -
# ### Create new DataFrame fitting weather criteria
# * Narrow down the cities to fit weather conditions.
# * Drop any rows will null values.
# +
# Narrow down the DataFrame to find your ideal weather condition.
# Drop any rows that don't contain all three conditions. You want to be sure the weather is ideal.
# Set specifications
ideal_temp = (cleancities['Max Temp']>70) & (cleancities['Max Temp']<86)
ideal_wind = cleancities['Wind Speed']<10
ideal_humid = cleancities['Humidity']<50
# Collect all ideal specs
ideal_vaca = ideal_temp & ideal_wind & ideal_humid
# Create new df using collected specs
ideal_vaca_df = cleancities[ideal_vaca]
ideal_vaca_df = ideal_vaca_df.dropna()
# Limit the number of rows returned by your API requests to a reasonable number.
# I think this is what that means... by making sure only 7 rows are returned..
ideal_vaca_df.head(7)
# -
# ### Hotel Map
# * Store into variable named `hotel_df`.
# * Add a "Hotel Name" column to the DataFrame.
# * Set parameters to search for hotels with 5000 meters.
# * Hit the Google Places API for each city's coordinates.
# * Store the first Hotel result into the DataFrame.
# * Plot markers on top of the heatmap.
# +
# Store into variable named hotel_df
# I think that means just change the name of the df..
hotel_df = ideal_vaca_df
# Add a "Hotel Name" column to the df
# Use empty quotes for initial value, since we don't have that data yet
hotel_df['Hotel Name'] = ""
hotel_df.head()
# +
# Hit the Google Places API for each city's coordinates
# Set parameters dictionary to search for hotels with 5000 meters
params = {
"radius": 5000,
"types": "hotels",
"keyword": "hotel",
"key": g_key}
# Start a for loop using iterrows
for index, row in hotel_df.iterrows():
# First, get the lat and long coords from our df
lat = row['Lat']
lng = row['Lng']
# Add a location parameter using lat and long that we just iterrated through
params['location'] = f"{lat},{lng}"
base_url = "https://maps.googleapis.com/maps/api/place/nearbysearch/json"
response = requests.get(base_url, params=params).json()
# Store the first Hotel result into the DataFrame
try:
hotel_df.loc[index, "Hotel Name"] = response["results"][0]["name"]
except:
print("Missing data")
# -
hotel_df.head()
# +
# NOTE: Do not change any of the code in this cell
# Using the template add the hotel marks to the heatmap
info_box_template = """
<dl>
<dt>Name</dt><dd>{Hotel Name}</dd>
<dt>City</dt><dd>{City}</dd>
<dt>Country</dt><dd>{Country}</dd>
</dl>
"""
# Store the DataFrame Row
# NOTE: be sure to update with your DataFrame name
hotel_info = [info_box_template.format(**row) for index, row in hotel_df.iterrows()]
locations = hotel_df[["Lat", "Lng"]]
# +
# Add some specifications to heatmap
# This part reminds me of CSS, syntax is basicly identical, which makes sense since we are styling an image
heatmap_specs = {
'width': '1000px',
'height': '500px',
'margin': '0 auto 0 auto'
}
# Add marker layer ontop of heat map
fig = gmaps.figure(layout=heatmap_specs, zoom_level=2, center=(0,0))
hotel_markers = gmaps.marker_layer(locations, info_box_content=hotel_info)
fig.add_layer(heat_layer)
fig.add_layer(hotel_markers)
# Save figure
plt.savefig("../Images/hotel_heatmap.png")
plt.show()
# Display figure
fig
# -
| VacationPy/VacationPy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import os
from scipy import stats
import pandas as pd
p_val_csv = "../data/csvfiles/p_values.csv"
df = pd.read_csv(p_val_csv, index_col = False)
df[df['Orientation Angle'] == 60]
# +
grouped = df[df['Orientation Angle'] == 60].groupby('Time')
two_hours = grouped.get_group('2hrs')
four_hours = grouped.get_group('4hrs')
six_hours = grouped.get_group('6hrs')
_, p_value_two_hours_fisher = stats.combine_pvalues(two_hours['p_value'],
method='fisher')
_, p_value_four_hours_fisher = stats.combine_pvalues(four_hours['p_value'],
method='fisher')
_, p_value_six_hours_fisher = stats.combine_pvalues(six_hours['p_value'],
method='fisher')
_, p_value_two_hours_weightedZ = stats.combine_pvalues(two_hours['p_value'],
method='stouffer',
weights=1/two_hours['Standard_error']
)
_, p_value_four_hours_weightedZ = stats.combine_pvalues(four_hours['p_value'],
method='stouffer',
weights=1/four_hours['Standard_error']
)
_, p_value_six_hours_weightedZ = stats.combine_pvalues(two_hours['p_value'],
method='stouffer',
weights=1/six_hours['Standard_error']
)
print("2hr p-value from Fisher's Method:{}".format(p_value_two_hours_fisher))
print("2hr p-value from Stouffer's Weighted Method:{}".format(p_value_two_hours_weightedZ))
print("\n")
print("4hr p-value from Fisher's Method:{}".format(p_value_four_hours_fisher))
print("4hr p-value from Stouffer's Weighted Method:{}".format(p_value_four_hours_weightedZ))
print("\n")
print("6hr p-value from Fisher's Method:{}".format(p_value_six_hours_fisher))
print("6hr p-value from Stouffer's Weighted Method:{}".format(p_value_six_hours_weightedZ))
| notebooks/Combining_p-values.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# > **Note:** In most sessions you will be solving exercises posed in a Jupyter notebook that looks like this one. Because you are cloning a Github repository that only we can push to, you should **NEVER EDIT** any of the files you pull from Github. Instead, what you should do, is either make a new notebook and write your solutions in there, or **make a copy of this notebook and save it somewhere else** on your computer, not inside the `sds` folder that you cloned, so you can write your answers in there. If you edit the notebook you pulled from Github, those edits (possible your solutions to the exercises) may be overwritten and lost the next time you pull from Github. This is important, so don't hesitate to ask if it is unclear.
#
# You should run `pip install scraping_class`
import scraping_class
conn = scraping_class.Connector("log.csv")
# # Exercise Set 9: Parsing and Information Extraction
#
# *Morning, August 17, 2018*
#
# In this Exercise Set we shall develop our webscraping skills even further by practicing **parsing** and navigating html trees using BeautifoulSoup and furthermore train extracting information from raw text with no html tags to help, using regular expressions.
#
# But just as importantly you will get a chance to think about **data quality issues** and how to ensure reliability when curating your own webdata.
# ## Exercise Section 9.1: Logging and data quality
#
# > **Ex. 9.1.1:** *Why is is it important to log processes in your data collection?*
#
# ** answer goes here**
#
#
#
# > **Ex. 9.1.2:**
#
# *How does logging help with both ensuring and documenting the quality of your data?*
# ** answer goes here**
#
#
# ## Exercise Section 9.2: Parsing a Table from HTML using BeautifulSoup.
#
# Yesterday I showed you a neat little prepackaged function in pandas that did all the work. However today we should learn the mechanics of it. *(It is not just for educational purposes, sometimes the package will not do exactly as you want.)*
#
# We hit the Basketball stats page from yesterday again: https://www.basketball-reference.com/leagues/NBA_2018.html.
#
# > **Ex. 9.2.1:** Here we practice simply locating the table node of interest using the `find` method build into BeautifoulSoup. But first we have to fetch the HTML using the `requests` module. Parse the tree using `BeautifulSoup`. And then use the **>Inspector<** tool (* right click on the table < press inspect element *) in your browser to see how to locate the Eastern Conference table node - i.e. the *tag* name of the node, and maybe some defining *attributes*.
# +
# [Answer to Ex. 9.2.1]
# -
# You have located the table should now build a function that starts at a "table node" and parses the information, and outputs a pandas DataFrame.
#
# Inspect the element either within the notebook or through the **>Inspector<** tool and start to see how a table is written in html. Which tag names can be used to locate rows? How will you iterate through columns. Were is the header located?
#
# > **Ex. 9.2.2:** First you parse the header which can be found in the canonical tag name: thead.
# Next you use the `find_all` method to search for the tag, and iterate through each of the elements extracting the text, using the `.text` method builtin to the the node object. Store the header values in a list container.
#
# > **Ex. 9.2.3:** Next you locate the rows, using the canonical tag name: tbody. And from here you search for all rows tags. Fiugre out the tag name yourself, inspecting the tbody node in python or using the **Inspector**.
#
# > **Ex. 9.2.4:** Next run through all the rows and extract each value, similar to how you extracted the header. However here is a slight variation: Since each value node can have a different tag depending on whether it is a digit or a string, you should use the `.children` method instead of the `.find_all` - (or write compile a regex that matches both the td tag and the th tag.)
# >Once the value nodes of each row has been located using the `.children` method you should extract the value. Store the extracted rows as a list of lists: ```[[val1,val2,...valk],...]```
# +
# [Answer to Ex. 9.2.2-4]
# -
# **Ex. 9.2.5** Convert the data you have collected into a pandas dataframe. _Bonus:_ convert the code you've written above into a function which scrapes the page and returns a dataframe.
# +
#[Answer 9.2.5]
# -
# > **Ex. 9.2.6:** Now locate all tables from the page, using the `.find_all` method searching for the table tag name. Iterate through the table nodes and apply the function created for parsing html tables. Store each table in a dictionary using the table name as key. The name is found by accessing the id attribute of each table node, using dictionary-style syntax - i.e. `table_node['id']`.
#
# > **9.2.extra.:** Compare your results to the pandas implementation. pd.read_html
# +
# [Answer to Ex. 9.1.6]
# -
# ## Exercise Section 9.3: Practicing Regular Expressions.
# This exercise is about developing your experience with designing your own regular expressions.
#
# Remember you can always consult the regular expression reference page [here](https://www.regular-expressions.info/refquick.html), if you need to remember or understand a specific symbol.
#
# You should practice using *"define-inspect-refine-method"* described in the lectures to systematically ***explore*** and ***refine*** your expressions, and save all the patterns tried. You can download the small module that I created to handle this in the following way:
# ``` python
# import requests
# url = 'https://raw.githubusercontent.com/snorreralund/explore_regex/master/explore_regex.py'
# response = requests.get(url)
# with open('explore_regex.py','w') as f:
# f.write(response.text)
# import explore_regex as e_re
# ```
#
# Remember to start ***broad*** to gain many examples, and iteratively narrow and refine.
#
# We will use a sample of the trustpilot dataset that you practiced collecting yesterday.
# You can load it directly into python from the following link: https://raw.githubusercontent.com/snorreralund/scraping_seminar/master/english_review_sample.csv
# > **Ex. 9.3.0:** Load the data used in the exercise using the `pd.read_csv` function. (Hint: path to file can be both a url or systempath).
#
# >Define a variable `sample_string = '\n'.join(df.sample(2000).reviewBody)` as sample of all the reviews that you will practice on. (Run it once in a while to get a new sample for potential differences).
# Imagine we were a company wanting to find the reviews where customers are concerned with the price of a service. They decide to write a regular expression to match all reviews where a currencies and an amount is mentioned.
#
# > **Ex. 9.3.1:**
# > Write an expression that matches both the dollar-sign (\$) and dollar written literally, and the amount before or after a dollar-sign. Remember that the "$"-sign is a special character in regular expressions. Explore and refine using the explore_pattern function in the package I created called explore_regex.
# ```python
# import explore_regex as e_re
# explore_regex = e_re.Explore_Regex(sample_string) # Initaizlie the Explore regex Class.
# explore_regex.explore_pattern(pattern) # Use the .explore_pattern method.
# ```
#
#
# Start with exploring the context around digits ("\d") in the data.
import pandas as pd
import re
import requests
# download data
path2data = 'https://raw.githubusercontent.com/snorreralund/scraping_seminar/master/english_review_sample.csv'
df = pd.read_csv(path2data)
# download module
url = 'https://raw.githubusercontent.com/snorreralund/explore_regex/master/explore_regex.py'
response = requests.get(url)
# write script to your folder to create a locate module
with open('explore_regex.py','w') as f:
f.write(response.text)
# import local module
import explore_regex as e_re
import re
digit_re = re.compile('[0-9]+')
df['hasNumber'] = df.reviewBody.apply(lambda x: len(digit_re.findall(x))>0)
sample_string = '\n'.join(df[df['hasNumber']].sample(1000).reviewBody)
explore_regex = e_re.ExploreRegex(sample_string)
# [Answer to exercise 9.3.1]
# > **Ex.9.3.3** Use the .report() method. e_re.report(), and print the all patterns in the development process using the .pattern method - i.e. e_re.patterns
#
# +
#[Answer 9.3.3]
# -
# >**Ex. 9.3.4**
# Finally write a function that takes in a string and outputs if there is a match. Use the .match function to see if there is a match (hint if does not return a NoneType object - `re.match(pattern,string)!=None`).
#
# > Define a column 'mention_currency' in the dataframe, by applying the above function to the text column of the dataframe.
# *** You should have approximately 310 reviews that matches. - but less is also alright***
#
# > **Ex. 9.3.5** Explore the relation between reviews mentioning prices and the average rating.
#
# > **Ex. 9.3.extra** Define a function that outputs the amount mentioned in the review (if more than one the largest), define a new column by applying it to the data, and explore whether reviews mentioning higher prices are worse than others by plotting the amount versus the rating.
#[Answer to 9.3.4-5]
# > **Ex. 9.3.5:** Now we write a regular expression to extract emoticons from text.
# Start by locating all mouths ')' of emoticons, and develop the variations from there. Remember that paranthesis are special characters in regex, so you should use the escape character.
| material/session_10/exercise_9.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exploratory Data Analysis (EDA) - Ames Housing Data
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import missingno
import warnings
warnings.filterwarnings("ignore")
pd.set_option('display.max_rows', 500)
pd.set_option('display.max_columns', 500)
pd.set_option('display.width', 1000)
# %matplotlib inline
# -
test = pd.read_csv('test.csv')
test.head()
test.columns
def ames_eda(df):
eda_df = {}
eda_df['Null_Sum'] = df.isnull().sum()
eda_df['Null_Percentage'] = df.isnull().mean()*100
eda_df['Dtypes'] = df.dtypes
eda_df['Count'] = df.count()
eda_df['Mean'] = df.mean()
eda_df['Median'] = df.median()
eda_df['Min'] = df.min()
eda_df['Max'] = df.max()
eda_df['Nunique'] = df.nunique()
return pd.DataFrame(eda_df)
test.columns
ames_eda(test)
test.describe().T
# ### Missing Value Analysis
missingno.matrix(test, color = (0, .5, .5))
missingno.bar(test, color = (0, .5, .5))
# Sorting Null Values in Descending order
test.isnull().sum().sort_values(ascending = False)
# ## 2. Analysing Train Dataset
train = pd.read_csv('train.csv')
train.head()
train.columns
# ### Exploring Relationship with Target
# DataFrame.corrwith(self, other, axis=0, drop=False, method='pearson')
# Compute pairwise correlation.
#
# Pairwise correlation is computed between rows or columns of DataFrame with rows or columns of Series or DataFrame. DataFrames are first aligned along both axes before computing the correlations.
correlations = train.corrwith(train['SalePrice']).iloc[:-1].to_frame()
correlations['abs'] = correlations[0].abs()
sorted_correlations = correlations.sort_values('abs', ascending=False)[0]
fig, ax = plt.subplots(figsize=(10,20))
sns.heatmap(sorted_correlations.to_frame(), cmap='coolwarm', annot=True, vmin=-1, vmax=1, ax=ax);
# ### Relationship between non-numeric values such the presence of central air with sale price
sns.boxplot(train['CentralAir'],
train['SalePrice']).set_title('Central Air vs. Sale Price');
# #### Based on this visualization, we will need to convert these columns to represent numeric values when we clean our data before modeling.
sns.boxplot(train['KitchenQual'], train['SalePrice']).set_title('Kitchen Quality vs. Sale Price')
train['GarageQual'].value_counts()
# #### First, we want to identify the range of values that a certain feature may contain. Based on the values identified, we can create a function to overwrite each value with numerical values.
def garage_qual_cleaner(cell):
if cell == 'Ex':
return 5
elif cell == 'Gd':
return 4
elif cell == 'TA':
return 3
elif cell == 'Fa':
return 2
elif cell == 'Po':
return 1
else:
return 0
# #### Alternatively, we could map a dictionary to overwrite values without creating a function.
train['KitchenQual'].map({'Ex': 5, 'Gd': 4, 'TA': 3, 'Fa': 2, 'Po': 1})
ordinal_col_names = [col for col in train.columns if (col[-4:] in ['Qual', 'Cond']) and col[:3] != 'Ove']
ordinal_col_names
plt.figure(figsize=(35,10)) # adjust the fig size to see everything
sns.boxplot(train['Neighborhood'], train['SalePrice']).set_title('Sale Price varies widely by Ames Neighborhood');
# #### Some neighborhoods clearly have higher sale prices have than others — a relationship that we want to capture in our model.
# ### Check for Outliers
# #### A quick way to check for outliers is to build a scatter plot between the target and a variable we would expect to be linearly related to our target
sns.regplot(x = '1stFlrSF', y = 'SalePrice', data = train)
# #### Filtering the dataset below to inspect the outliers in greater detail:
train.loc[train['1stFlrSF'] > 3000]
rows_to_drop = [496, 523, 1298]
for row in rows_to_drop:
train.drop(row, inplace=True)
sns.regplot(x = '1stFlrSF', y = 'SalePrice', data = train)
# Reference
# https://medium.com/swlh/a-complete-guide-to-exploratory-data-analysis-and-data-cleaning-dd282925320f
# Thankyou - NickVega
#
# ### Conclusion
# By investing our time to explore and clean our data, we have identified important outliers and extracted valuable information from string columns that will be critical to building a powerful predictive model.
| 2_EDA_Ames_Housing_Data/EDA_Ames_Dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data
# For our purposes, we will be working with the [Video Game Sales Dataset](https://www.kaggle.com/gregorut/videogamesales).
# !pip install pandas
import pandas as pd
# Let's load data
df = pd.read_csv("vgsales.csv")
# Let’s also relax the limit on the number of display columns:
pd.set_option('display.max_columns', None)
# # Data Manipulation
# Next, let’s print the columns in the data:
list(df.columns)
# Let’s also print some basic information about the data
df.info()
# Now, let’s print the first five rows of data using the ‘head()’ method:
df.head()
# Let’s select video games in the ‘sports’ genre:
df_sports = df.loc[df.Genre == 'Sports']
df_sports.head()
# Let’s select the subset of data corresponding video games for the Wii platform:
df_wii = df.loc[df.Platform == 'Wii']
df_wii.head()
# Let’s select ‘Racing’ games played on ‘Wii’:
df_wii_racing = df_wii.loc[df.Genre == 'Racing']
df_wii_racing.head()
# Another way to do this in online with joint operation
df_wii_racing = df.loc[(df.Platform == 'Wii') & (df.Genre == 'Racing')]
df_wii_racing.head()
# Let’s select Wii racing games that sold more than 1 million units globally
df_gt_1mil = df.loc[(df.Platform == 'Wii') & (df.Genre == 'Racing') & (df.Global_Sales >= 1.0)]
df_gt_1mil.head()
# We can also select data by row using the ‘.iloc[]’ method. Let’s select the first 1000 rows of the original data:
df_filter_rows = df.iloc[:1000]
print("Length of original: ", len(df))
print("Length of filtered: ", len(df_filter_rows))
# We can also select a random sample of data, using the ‘sample()’ method:
df_random_sample = df.sample(n=5000)
print("Length of sample: ", len(df_random_sample))
df_random_sample.head()
# # Conclusion
# In this notebook we discussed how to select and filter data using the Python Pandas library. We discussed how to use the ‘.loc[]’ method to select subsets of data based on column values. We also showed how to filter data frames by row using the ‘.iloc[]’ method. Finally, we discussed how to select a random sample of data from a data frame using the ‘sample()’ method.
| Mastering Data Selection with Pandas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="zv5HN3iAwfSc"
# # AutoEncoder
# + [markdown] id="LxZ4Oc1vwkYI"
# ## Import Libraries
# + id="_ftUoL6FwbLA" executionInfo={"status": "ok", "timestamp": 1643875107444, "user_tz": -60, "elapsed": 5940, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "11023693490829624613"}}
import numpy as np
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers, models, losses
from tensorflow.keras.models import Sequential
from keras.models import Model
import matplotlib.pyplot as plt
from sklearn import metrics
import pandas as pd
from tensorflow.keras import optimizers
import os
import shutil
# + [markdown] id="cynySYouwpVa"
# ## Connect to Drive
# + colab={"base_uri": "https://localhost:8080/"} id="rIBfjLGowpF9" executionInfo={"status": "ok", "timestamp": 1643875129948, "user_tz": -60, "elapsed": 20328, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "11023693490829624613"}} outputId="9c0bb281-745f-4195-d667-508c78280a17"
from google.colab import drive
drive.mount('/content/drive', force_remount=True)
# + [markdown] id="J9Ps-Q2JwzLu"
# ## Set parameters
# + id="IicfMq3Hw2Ky" executionInfo={"status": "ok", "timestamp": 1643875148850, "user_tz": -60, "elapsed": 274, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "11023693490829624613"}}
FEATURES_FOLDER = '/content/drive/My Drive/CV_Birds/features'
LOCAL_MODELS_FOLDER = "/content/models"
if not os.path.exists(LOCAL_MODELS_FOLDER):
os.makedirs(LOCAL_MODELS_FOLDER)
GLOBAL_MODELS_FOLDER = "/content/drive/My Drive/CV_Birds/models/AutoEncoder"
BATCH_SIZE = 256
# + id="C_Q8KQlf_2Sa" executionInfo={"status": "ok", "timestamp": 1643875239567, "user_tz": -60, "elapsed": 279, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "11023693490829624613"}}
callbacks_list = [
keras.callbacks.EarlyStopping(
monitor='val_loss',
patience=6,
restore_best_weights=True,
),
keras.callbacks.ModelCheckpoint(
filepath='autoencoder.keras',
monitor='val_loss',
save_best_only=True,
)
]
# + [markdown] id="6Qcy6MUGmRkk"
# For Finetuning:
# + id="KbP6vsqBxB3G"
training_features = np.load(FEATURES_FOLDER + '/training/ResNet152v2/OneDense512_Dropout_fine_tuning.npy')
# + colab={"base_uri": "https://localhost:8080/"} id="TZ2KsPRsx2Z4" executionInfo={"status": "ok", "timestamp": 1643822104067, "user_tz": -60, "elapsed": 4, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "10977236443775929893"}} outputId="b9235f6d-a68a-4017-b28e-08d384b75eb1"
feature_dimension = training_features.shape[1]
feature_dimension
# + id="uieNbzBGyObn"
validation_features = np.load(FEATURES_FOLDER + '/validation/ResNet152v2/OneDense512_Dropout_fine_tuning.npy')
test_features = np.load(FEATURES_FOLDER + '/test/ResNet152v2/OneDense512_Dropout_fine_tuning.npy')
# + id="kLBV_3fNGXJH"
distractor_features = np.load(FEATURES_FOLDER + '/distractor/ResNet152v2/OneDense512_Dropout_fine_tuning.npy')
# + [markdown] id="sHa5P4CsmPwl"
# For feature extraction:
# + id="4Rc-EApAmUrt" executionInfo={"status": "ok", "timestamp": 1643875298673, "user_tz": -60, "elapsed": 4387, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "11023693490829624613"}}
training_features = np.load(FEATURES_FOLDER + '/training/ResNet152v2/OneDense512_Dropout_feature_extraction.npy')
validation_features = np.load(FEATURES_FOLDER + '/validation/ResNet152v2/OneDense512_Dropout_feature_extraction.npy')
test_features = np.load(FEATURES_FOLDER + '/test/ResNet152v2/OneDense512_Dropout_feature_extraction.npy')
distractor_features = np.load(FEATURES_FOLDER + '/distractor/ResNet152v2/OneDense512_Dropout_feature_extraction.npy')
# + [markdown] id="fETFag8yyzUr"
# # AutoEncoder Class
# + id="CpuEoKlwyy5O" executionInfo={"status": "ok", "timestamp": 1643875306785, "user_tz": -60, "elapsed": 250, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "11023693490829624613"}}
class AutoEncoder:
def __init__(self, train_features, val_features, latent_dim, pace):
self.train_features = train_features
self.val_features = val_features
self.encoder_final_layer = 'Encoder' + str(latent_dim)
dim = feature_dimension = train_features.shape[1]
input = keras.Input(shape=(feature_dimension,))
# Enconder layers
dim -= pace
name = 'Encoder' + str(dim)
x = keras.layers.Dense(dim, activation='relu', name=name)(input)
while dim != latent_dim:
dim -= pace
name = 'Encoder' + str(dim)
x = keras.layers.Dense(dim, activation='relu', name=name)(x)
# Decoder Layers
while dim != feature_dimension - pace:
dim += pace
name = 'Decoder' + str(dim)
x = keras.layers.Dense(dim, activation='relu', name=name)(x)
outputs = keras.layers.Dense(feature_dimension)(x)
self.model = keras.Model(inputs=input, outputs=outputs)
def summary(self):
self.model.summary()
def compile(self, loss='mean_squared_error', optimizer='sgd'):
self.model.compile(loss=loss,
optimizer=optimizer,
metrics=[tf.keras.metrics.CosineSimilarity(axis=1)])
def fit(self, callbacks, epochs=50, patience=5, batch_size=BATCH_SIZE):
self.history = self.model.fit(self.train_features,
self.train_features,
validation_data=(self.val_features, self.val_features),
callbacks=callbacks,
shuffle=True,
batch_size=batch_size,
epochs=epochs)
def evaluate(self, test_features):
test_loss, test_acc = self.model.evaluate((test_features, test_features))
print(f"Test accuracy: {test_acc:.3f}")
def predict_features_encoder(self, features, path_name):
encoder = keras.Model(self.model.input, outputs=self.model.get_layer(self.encoder_final_layer).output)
features = encoder.predict(features)
print(features.shape)
print(features)
np.save(FEATURES_FOLDER + path_name, features)
def plot_model(self, model_name):
keras.utils.plot_model(self.model, model_name, show_shapes=True)
def plot_loss(self):
loss = self.history.history['loss']
val_loss = self.history.history['val_loss']
plt.plot(range(1, len(loss) + 1), loss, 'r', label='Training Loss')
plt.plot(range(1, len(loss) + 1), val_loss, 'g', label='Validation Loss')
plt.title('Training and Validation Loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.plot()
# + [markdown] id="FIViXKD7w5zj"
# # ResNet 152 512 to 256 features FineTuning
# + colab={"base_uri": "https://localhost:8080/"} id="tNbrUg6i4ycO" executionInfo={"status": "ok", "timestamp": 1643788043140, "user_tz": -60, "elapsed": 1844, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "11023693490829624613"}} outputId="3721d6b5-1f71-407c-cf8a-089834a45ec3"
autoencoder = AutoEncoder(training_features, validation_features, 256, 64)
autoencoder.summary()
# + id="Q1_4CmySX38u"
autoencoder.plot_model('autoencoder.png')
# + id="Jzt6Z9OeDZcM" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1643788656416, "user_tz": -60, "elapsed": 44992, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "11023693490829624613"}} outputId="27f76813-f315-47be-8c11-67d0b4ad53e6"
autoencoder.compile()
autoencoder.fit(callbacks_list, epochs=800)
# + id="5pb6EE8_DoFm"
autoencoder.predict_features_encoder(training_features, '/training/AutoEncoder/512to256withPace64.npy')
# + id="IE5c0szBEdbN"
autoencoder.predict_features_encoder(test_features, '/test/AutoEncoder/512to256withPace64.npy')
# + colab={"base_uri": "https://localhost:8080/"} id="j8C2MapyclP6" executionInfo={"status": "ok", "timestamp": 1643788851036, "user_tz": -60, "elapsed": 2105, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "11023693490829624613"}} outputId="ad38fb3c-f8bc-4433-8daa-1ab240d17b78"
encoder = keras.Model(autoencoder.model.input, outputs=autoencoder.model.get_layer(autoencoder.encoder_final_layer).output)
features = encoder.predict(training_features)
print(features.shape)
print(features)
# + id="hz6vOaW1c2DL"
np.save(FEATURES_FOLDER + '/training/AutoEncoder/512to256withPace64.npy', features)
# + colab={"base_uri": "https://localhost:8080/"} id="zGd_OsyIc4X3" executionInfo={"status": "ok", "timestamp": 1643788902330, "user_tz": -60, "elapsed": 620, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "11023693490829624613"}} outputId="2d98f6af-e6a2-4c9f-eb84-07c969c7c427"
features = encoder.predict(test_features)
print(features.shape)
print(features)
np.save(FEATURES_FOLDER + '/test/AutoEncoder/512to256withPace64.npy', features)
# + colab={"base_uri": "https://localhost:8080/"} id="qSdTloEvdNxT" executionInfo={"status": "ok", "timestamp": 1643789013838, "user_tz": -60, "elapsed": 2153, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "11023693490829624613"}} outputId="60c5900a-9679-4e84-e433-857e0441fb71"
features = encoder.predict(distractor_features)
print(features.shape)
print(features)
np.save(FEATURES_FOLDER + '/distractor/AutoEncoder/512to256withPace64.npy', features)
# + id="oVzHh70tdkTY"
# ! cp autoencoder.keras /content/drive/MyDrive/CV_Birds/models/AutoEncoder/
# + [markdown] id="0h4oR29OGHFX"
# # ResNet152v2 512 to 128 Finetuning
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1643822135894, "user_tz": -60, "elapsed": 3553, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "10977236443775929893"}} outputId="bd612759-cd5e-4bfd-a377-598dc01e771a" id="BxhZ2waYGgs3"
autoencoder = AutoEncoder(training_features, validation_features, 128, 64)
autoencoder.summary()
# + id="NoQUfe8lGgs4"
autoencoder.plot_model('autoencoder.png')
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1643823104860, "user_tz": -60, "elapsed": 24417, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "10977236443775929893"}} outputId="35f54863-f6f7-4476-d59d-e5d612f1f52a" id="2APmWyp_Ggs5"
autoencoder.compile()
autoencoder.fit(callbacks_list, epochs=800)
# + colab={"base_uri": "https://localhost:8080/"} id="OKN0jBfKGgs6" executionInfo={"status": "ok", "timestamp": 1643823118414, "user_tz": -60, "elapsed": 3846, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "10977236443775929893"}} outputId="fe4f9bab-b108-4f22-d30d-b94664f2bcc6"
autoencoder.predict_features_encoder(training_features, '/training/AutoEncoder/512to128withPace64.npy')
# + colab={"base_uri": "https://localhost:8080/"} id="MVPHkd4OGgs7" executionInfo={"status": "ok", "timestamp": 1643823143405, "user_tz": -60, "elapsed": 1282, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "10977236443775929893"}} outputId="06c0c008-fad9-4e60-d031-63e3e554c58d"
autoencoder.predict_features_encoder(test_features, '/test/AutoEncoder/512to128withPace64.npy')
# + colab={"base_uri": "https://localhost:8080/"} id="JyvyH9ewIU_B" executionInfo={"status": "ok", "timestamp": 1643823146740, "user_tz": -60, "elapsed": 2160, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "10977236443775929893"}} outputId="7700cdba-d152-4561-8b05-c8e071131704"
autoencoder.predict_features_encoder(distractor_features, '/distractor/AutoEncoder/512to128withPace64.npy')
# + id="SWgyzF5Wb-Yo"
# ! cp autoencoder128.keras /content/drive/MyDrive/CV_Birds/models/AutoEncoder/
# + [markdown] id="7fe80N1elOlQ"
# # ResNet152v2 512 to 256 Feature Extraction
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1643875346885, "user_tz": -60, "elapsed": 3267, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "11023693490829624613"}} outputId="915a8481-3e54-401f-e66e-51c2cceb07fd" id="yJ8ack7vlOlR"
autoencoder = AutoEncoder(training_features, validation_features, 256, 64)
autoencoder.summary()
# + id="SQAADgPalOlS"
autoencoder.plot_model('autoencoder.png')
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1643875986835, "user_tz": -60, "elapsed": 101686, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "11023693490829624613"}} outputId="6948406d-1c0b-4411-e5a4-d09a5fbadb16" id="RIFO7YUMlOlS"
autoencoder.compile()
autoencoder.fit(callbacks_list, epochs=800)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1643876059340, "user_tz": -60, "elapsed": 3100, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "11023693490829624613"}} outputId="d2cfb44e-593d-441a-ca2d-863a0a26333c" id="3zBS5G-7lOlT"
autoencoder.predict_features_encoder(training_features, '/training/AutoEncoder/512to256withPace64_feature_extraction.npy')
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1643876062253, "user_tz": -60, "elapsed": 876, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "11023693490829624613"}} outputId="777db672-48d4-4d51-86fd-f6c2479be6ec" id="yq6py7VHlOlT"
autoencoder.predict_features_encoder(test_features, '/test/AutoEncoder/512to256withPace64_feature_extraction.npy')
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1643876066305, "user_tz": -60, "elapsed": 1737, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "11023693490829624613"}} outputId="e9b09eef-cde4-4290-aca5-768d4ca1ba36" id="WZ7jtK7alOlU"
autoencoder.predict_features_encoder(distractor_features, '/distractor/AutoEncoder/512to256withPace64_feature_extraction.npy')
# + id="K1OKHykrlOlU" executionInfo={"status": "ok", "timestamp": 1643876099300, "user_tz": -60, "elapsed": 643, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "11023693490829624613"}}
# ! cp autoencoder256_feature_extraction.keras /content/drive/MyDrive/CV_Birds/models/AutoEncoder/
# + [markdown] id="x8yJo8mdnHp1"
# # ResNet152v2 512 to 128 Feature Extraction
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1643876104369, "user_tz": -60, "elapsed": 276, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "11023693490829624613"}} outputId="a8be80a9-0ea3-47f2-d706-18e37c180e45" id="Cn0NZ8vDnHp2"
autoencoder = AutoEncoder(training_features, validation_features, 128, 64)
autoencoder.summary()
# + id="jw9kP28PnHp3"
autoencoder.plot_model('autoencoder.png')
# + colab={"base_uri": "https://localhost:8080/"} outputId="8acd88c4-6fb9-41a9-a2fb-88f0ff58bb69" id="CouAHISLnHp3" executionInfo={"status": "ok", "timestamp": 1643876817190, "user_tz": -60, "elapsed": 13954, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "11023693490829624613"}}
autoencoder.compile()
autoencoder.fit(callbacks_list, epochs=800)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1643876841692, "user_tz": -60, "elapsed": 2865, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "11023693490829624613"}} outputId="db3aad82-f1bc-4a28-b707-15a61fcd5ffe" id="LbCDtaOonHp4"
autoencoder.predict_features_encoder(training_features, '/training/AutoEncoder/512to128withPace64_feature_extraction.npy')
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1643876846657, "user_tz": -60, "elapsed": 259, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "11023693490829624613"}} outputId="92a80f05-4dae-42f0-acb1-397483c62ea6" id="YEBUHfoZnHp5"
autoencoder.predict_features_encoder(test_features, '/test/AutoEncoder/512to128withPace64_feature_extraction.npy')
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1643876854087, "user_tz": -60, "elapsed": 1565, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "11023693490829624613"}} outputId="31d68aed-66eb-4a49-dac9-4603ddc065f4" id="ZMeZzck_nHp5"
autoencoder.predict_features_encoder(distractor_features, '/distractor/AutoEncoder/512to128withPace64_feature_extraction.npy')
# + id="32MKjN-bsvL8" executionInfo={"status": "ok", "timestamp": 1643876966166, "user_tz": -60, "elapsed": 258, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "11023693490829624613"}}
keras.models.save_model(autoencoder.model, '/content/drive/MyDrive/CV_Birds/models/AutoEncoder/autoencoder128_feature_extraction.keras')
# + id="cgptU_itnHp6"
# ! cp autoencoder128_feature_extraction.keras /content/drive/MyDrive/CV_Birds/models/AutoEncoder/
| Notebooks/Training/AutoEncoder.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <!--BOOK_INFORMATION-->
# <img align="left" style="padding-right:10px;" src="figures/PDSH-cover-small.png">
# *This notebook contains an excerpt from the [Python Data Science Handbook](http://shop.oreilly.com/product/0636920034919.do) by <NAME>; the content is available [on GitHub](https://github.com/jakevdp/PythonDataScienceHandbook).*
#
# *The text is released under the [CC-BY-NC-ND license](https://creativecommons.org/licenses/by-nc-nd/3.0/us/legalcode), and code is released under the [MIT license](https://opensource.org/licenses/MIT). If you find this content useful, please consider supporting the work by [buying the book](http://shop.oreilly.com/product/0636920034919.do)!*
# <!--NAVIGATION-->
# < [Introducing Pandas Objects](03.01-Introducing-Pandas-Objects.ipynb) | [Contents](Index.ipynb) | [Operating on Data in Pandas](03.03-Operations-in-Pandas.ipynb) >
# # Data Indexing and Selection
# In [Chapter 2](02.00-Introduction-to-NumPy.ipynb), we looked in detail at methods and tools to access, set, and modify values in NumPy arrays.
# These included indexing (e.g., ``arr[2, 1]``), slicing (e.g., ``arr[:, 1:5]``), masking (e.g., ``arr[arr > 0]``), fancy indexing (e.g., ``arr[0, [1, 5]]``), and combinations thereof (e.g., ``arr[:, [1, 5]]``).
# Here we'll look at similar means of accessing and modifying values in Pandas ``Series`` and ``DataFrame`` objects.
# If you have used the NumPy patterns, the corresponding patterns in Pandas will feel very familiar, though there are a few quirks to be aware of.
#
# We'll start with the simple case of the one-dimensional ``Series`` object, and then move on to the more complicated two-dimesnional ``DataFrame`` object.
# ## Data Selection in Series
#
# As we saw in the previous section, a ``Series`` object acts in many ways like a one-dimensional NumPy array, and in many ways like a standard Python dictionary.
# If we keep these two overlapping analogies in mind, it will help us to understand the patterns of data indexing and selection in these arrays.
# ### Series as dictionary
#
# Like a dictionary, the ``Series`` object provides a mapping from a collection of keys to a collection of values:
import pandas as pd
data = pd.Series([0.25, 0.5, 0.75, 1.0],
index=['a', 'b', 'c', 'd'])
data
data['b']
# We can also use dictionary-like Python expressions and methods to examine the keys/indices and values:
'a' in data
data.keys()
list(data.items())
# ``Series`` objects can even be modified with a dictionary-like syntax.
# Just as you can extend a dictionary by assigning to a new key, you can extend a ``Series`` by assigning to a new index value:
data['e'] = 1.25
data
# This easy mutability of the objects is a convenient feature: under the hood, Pandas is making decisions about memory layout and data copying that might need to take place; the user generally does not need to worry about these issues.
# ### Series as one-dimensional array
# A ``Series`` builds on this dictionary-like interface and provides array-style item selection via the same basic mechanisms as NumPy arrays – that is, *slices*, *masking*, and *fancy indexing*.
# Examples of these are as follows:
# slicing by explicit index
data['a':'c']
# slicing by implicit integer index
data[0:2]
# masking
data[(data > 0.3) & (data < 0.8)]
# fancy indexing
data[['a', 'e']]
# Among these, slicing may be the source of the most confusion.
# Notice that when slicing with an explicit index (i.e., ``data['a':'c']``), the final index is *included* in the slice, while when slicing with an implicit index (i.e., ``data[0:2]``), the final index is *excluded* from the slice.
# ### Indexers: loc, iloc, and ix
#
# These slicing and indexing conventions can be a source of confusion.
# For example, if your ``Series`` has an explicit integer index, an indexing operation such as ``data[1]`` will use the explicit indices, while a slicing operation like ``data[1:3]`` will use the implicit Python-style index.
data = pd.Series(['a', 'b', 'c'], index=[1, 3, 5])
data
# explicit index when indexing
data[1]
# implicit index when slicing
data[1:3]
# Because of this potential confusion in the case of integer indexes, Pandas provides some special *indexer* attributes that explicitly expose certain indexing schemes.
# These are not functional methods, but attributes that expose a particular slicing interface to the data in the ``Series``.
#
# First, the ``loc`` attribute allows indexing and slicing that always references the explicit index:
data.loc[1]
data.loc[1:3]
# The ``iloc`` attribute allows indexing and slicing that always references the implicit Python-style index:
data.iloc[1]
data.iloc[1:3]
# A third indexing attribute, ``ix``, is a hybrid of the two, and for ``Series`` objects is equivalent to standard ``[]``-based indexing.
# The purpose of the ``ix`` indexer will become more apparent in the context of ``DataFrame`` objects, which we will discuss in a moment.
#
# One guiding principle of Python code is that "explicit is better than implicit."
# The explicit nature of ``loc`` and ``iloc`` make them very useful in maintaining clean and readable code; especially in the case of integer indexes, I recommend using these both to make code easier to read and understand, and to prevent subtle bugs due to the mixed indexing/slicing convention.
# ## Data Selection in DataFrame
#
# Recall that a ``DataFrame`` acts in many ways like a two-dimensional or structured array, and in other ways like a dictionary of ``Series`` structures sharing the same index.
# These analogies can be helpful to keep in mind as we explore data selection within this structure.
# ### DataFrame as a dictionary
#
# The first analogy we will consider is the ``DataFrame`` as a dictionary of related ``Series`` objects.
# Let's return to our example of areas and populations of states:
area = pd.Series({'California': 423967, 'Texas': 695662,
'New York': 141297, 'Florida': 170312,
'Illinois': 149995})
pop = pd.Series({'California': 38332521, 'Texas': 26448193,
'New York': 19651127, 'Florida': 19552860,
'Illinois': 12882135})
data = pd.DataFrame({'area':area, 'pop':pop})
data
# The individual ``Series`` that make up the columns of the ``DataFrame`` can be accessed via dictionary-style indexing of the column name:
data['area']
# Equivalently, we can use attribute-style access with column names that are strings:
data.area
# This attribute-style column access actually accesses the exact same object as the dictionary-style access:
data.area is data['area']
# Though this is a useful shorthand, keep in mind that it does not work for all cases!
# For example, if the column names are not strings, or if the column names conflict with methods of the ``DataFrame``, this attribute-style access is not possible.
# For example, the ``DataFrame`` has a ``pop()`` method, so ``data.pop`` will point to this rather than the ``"pop"`` column:
data.pop is data['pop']
# In particular, you should avoid the temptation to try column assignment via attribute (i.e., use ``data['pop'] = z`` rather than ``data.pop = z``).
#
# Like with the ``Series`` objects discussed earlier, this dictionary-style syntax can also be used to modify the object, in this case adding a new column:
data['density'] = data['pop'] / data['area']
data
# This shows a preview of the straightforward syntax of element-by-element arithmetic between ``Series`` objects; we'll dig into this further in [Operating on Data in Pandas](03.03-Operations-in-Pandas.ipynb).
# ### DataFrame as two-dimensional array
#
# As mentioned previously, we can also view the ``DataFrame`` as an enhanced two-dimensional array.
# We can examine the raw underlying data array using the ``values`` attribute:
data.values
# With this picture in mind, many familiar array-like observations can be done on the ``DataFrame`` itself.
# For example, we can transpose the full ``DataFrame`` to swap rows and columns:
data.T
# When it comes to indexing of ``DataFrame`` objects, however, it is clear that the dictionary-style indexing of columns precludes our ability to simply treat it as a NumPy array.
# In particular, passing a single index to an array accesses a row:
data.values[0]
# and passing a single "index" to a ``DataFrame`` accesses a column:
data['area']
# Thus for array-style indexing, we need another convention.
# Here Pandas again uses the ``loc``, ``iloc``, and ``ix`` indexers mentioned earlier.
# Using the ``iloc`` indexer, we can index the underlying array as if it is a simple NumPy array (using the implicit Python-style index), but the ``DataFrame`` index and column labels are maintained in the result:
data.iloc[:3, :2]
# Similarly, using the ``loc`` indexer we can index the underlying data in an array-like style but using the explicit index and column names:
data.loc[:'Illinois', :'pop']
# The ``ix`` indexer allows a hybrid of these two approaches:
data.ix[:3, :'pop']
# Keep in mind that for integer indices, the ``ix`` indexer is subject to the same potential sources of confusion as discussed for integer-indexed ``Series`` objects.
#
# Any of the familiar NumPy-style data access patterns can be used within these indexers.
# For example, in the ``loc`` indexer we can combine masking and fancy indexing as in the following:
data.loc[data.density > 100, ['pop', 'density']]
# Any of these indexing conventions may also be used to set or modify values; this is done in the standard way that you might be accustomed to from working with NumPy:
data.iloc[0, 2] = 90
data
# To build up your fluency in Pandas data manipulation, I suggest spending some time with a simple ``DataFrame`` and exploring the types of indexing, slicing, masking, and fancy indexing that are allowed by these various indexing approaches.
# ### Additional indexing conventions
#
# There are a couple extra indexing conventions that might seem at odds with the preceding discussion, but nevertheless can be very useful in practice.
# First, while *indexing* refers to columns, *slicing* refers to rows:
data['Florida':'Illinois']
# Such slices can also refer to rows by number rather than by index:
data[1:3]
# Similarly, direct masking operations are also interpreted row-wise rather than column-wise:
data[data.density > 100]
# These two conventions are syntactically similar to those on a NumPy array, and while these may not precisely fit the mold of the Pandas conventions, they are nevertheless quite useful in practice.
# <!--NAVIGATION-->
# < [Introducing Pandas Objects](03.01-Introducing-Pandas-Objects.ipynb) | [Contents](Index.ipynb) | [Operating on Data in Pandas](03.03-Operations-in-Pandas.ipynb) >
| PythonDataScienceHandbook/notebooks/03.02-Data-Indexing-and-Selection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 这是Python,TensorFlow和Keras教程系列的深度学习基础知识的第4部分。
#
# 在这一部分,我们将讨论的是TensorBoard。TensorBoard是一个方便的应用程序,允许您在浏览器中查看模型或模型的各个方面。我们将TensorBoard与Keras一起使用的方式是通过Keras回调。实际上有很多Keras回调,你可以自己制作。
from tensorflow.keras.callbacks import TensorBoard
# +
#创建TensorBoard回调对象
NAME = "Cats-vs-dogs-CNN"
tensorboard = TensorBoard(log_dir="logs/{}".format(NAME))
# -
# 最终,你会希望获得更多的自定义NAME,但现在这样做。因此,这将保存模型的训练数据logs/NAME,然后由TensorBoard读取。
#
# 最后,我们可以通过将它添加到.fit方法中来将此回调添加到我们的模型中,例如:
# ```python
# model.fit(X, y,
# batch_size=32,
# epochs=3,
# validation_split=0.3,
# callbacks=[tensorboard])
# ```
# 请注意,这callbacks是一个列表。您也可以将其他回调传递到此列表中。我们的模型还没有定义,所以现在让我们把它们放在一起:
# +
import tensorflow as tf
from tensorflow.keras.datasets import cifar10
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Dropout, Activation, Flatten
from tensorflow.keras.layers import Conv2D, MaxPooling2D
from tensorflow.keras.callbacks import TensorBoard
# more info on callbakcs: https://keras.io/callbacks/ model saver is cool too.
import pickle
import time
NAME = "Cats-vs-dogs-CNN"
pickle_in = open("./datasets/X.pickle","rb")
X = pickle.load(pickle_in)
pickle_in = open("./datasets/y.pickle","rb")
y = pickle.load(pickle_in)
X = X/255.0
model = Sequential()
model.add(Conv2D(256, (3, 3), input_shape=X.shape[1:]))
model.add(Activation('relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Conv2D(256, (3, 3)))
model.add(Activation('relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Flatten()) # this converts our 3D feature maps to 1D feature vectors
model.add(Dense(64))
model.add(Dense(1))
model.add(Activation('sigmoid'))
tensorboard = TensorBoard(log_dir="logs/{}".format(NAME))
model.compile(loss='binary_crossentropy',
optimizer='adam',
metrics=['accuracy'],
)
model.fit(X, y,
batch_size=32,
epochs=3,
validation_split=0.3,
callbacks=[tensorboard])
# -
# 运行此之后,您应该有一个名为的新目录logs。我们现在可以使用tensorboard从这个目录中可视化初始结果。打开控制台,切换到工作目录,然后键入:tensorboard --logdir=logs/。您应该看到一个通知:TensorBoard 1.10.0 at http://H-PC:6006 (Press CTRL+C to quit)“h-pc”是您机器的名称。打开浏览器并前往此地址。你应该看到类似的东西:
# <img src = "https://pythonprogramming.net/static/images/machine-learning/tensorboard-basic.png">
# 现在我们可以看到我们的模型随着时间的推移。让我们改变模型中的一些东西。首先,我们从未在密集层中添加激活。另外,让我们尝试整体较小的模型:
# +
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Dropout, Activation, Flatten
from tensorflow.keras.layers import Conv2D, MaxPooling2D
from tensorflow.keras.callbacks import TensorBoard
# more info on callbakcs: https://keras.io/callbacks/ model saver is cool too.
import pickle
import time
NAME = "Cats-vs-dogs-64x2-CNN"
pickle_in = open("./datasets/X.pickle","rb")
X = pickle.load(pickle_in)
pickle_in = open("./datasets/y.pickle","rb")
y = pickle.load(pickle_in)
X = X/255.0
model = Sequential()
model.add(Conv2D(64, (3, 3), input_shape=X.shape[1:]))
model.add(Activation('relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Conv2D(64, (3, 3)))
model.add(Activation('relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Flatten()) # this converts our 3D feature maps to 1D feature vectors
model.add(Dense(64))
model.add(Activation('relu'))
model.add(Dense(1))
model.add(Activation('sigmoid'))
tensorboard = TensorBoard(log_dir="logs/{}".format(NAME))
model.compile(loss='binary_crossentropy',
optimizer='adam',
metrics=['accuracy'],
)
model.fit(X, y,
batch_size=32,
epochs=10,
validation_split=0.3,
callbacks=[tensorboard])
# -
# 除此之外,我还改名为NAME = "Cats-vs-dogs-64x2-CNN"。不要忘记这样做,否则你会偶然附加到你以前的型号的日志,它看起来不太好。我们现在检查TensorBoard:
# <img src = "https://pythonprogramming.net/static/images/machine-learning/second-model-tensorboard.png">
# 看起来更好!但是,您可能会立即注意到验证丢失的形状。损失是衡量错误的标准,看起来很明显,在我们的第四个时代之后,事情开始变得糟糕。
#
# 有趣的是,我们的验证准确性仍然持续,但我想它最终会开始下降。更可能的是,第一件遭受的事情确实是你的验证损失。这应该提醒你,你几乎肯定会开始过度适应。这种情况发生的原因是该模型不断尝试减少样本损失。
#
# 在某些时候,模型不是学习关于实际数据的一般事物,而是开始只记忆输入数据。如果你继续这样做,是的,样本中的“准确性”会上升,但你的样本,以及你试图为模型提供的任何新数据可能会表现得很差。
| r1-ml/day4-100-days-of-ml-code-cn/Code/Day 42.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Deploy Monitor
#
# - get monitor project for arm(rapsberry pi) - https://github.com/6za/node-exporter-collectors
# - deploy on all nodes - `docker-compose`
# - `curl` to check connection
# ### Get Project
#
# Get [6za/node-exporter-collectors](https://github.com/6za/node-exporter-collectors) to deploy on nodes.
#
# + language="bash"
#
# mkdir ~/repos
# cd ~/repos
# #rm -rf ~/repos/node-exporter-collectors
# git clone https://github.com/6za/node-exporter-collectors.git
# -
# ### Deploying on ARM
#
# On Arm, there is extra monitor as part of the image to collect temperature from some sensors.
#
#
# Details at: [6za/prometheus-raspberry-exporter](https://github.com/6za/prometheus-raspberry-exporter)
#
# The images in use on `raspberry pi` are done using: [6za/pi-gen-vagrant](https://github.com/6za/pi-gen-vagrant)
#
# And customized with: [6za/pi-gen](https://github.com/6za/pi-gen)
#
#
# We will:
# - List from all hosts the ones based on `raspberry pi` and arch `armv7l`
# - deploy the arm version of the `node-exporter`
# - check the deployed containers
import pandas as pd
hosts = pd.read_csv("../common/hosts.csv")
supressed_columns = ['ip','user']
select_hosts = hosts[(hosts.user == "pi") & (hosts.arch == "armv7l")]
select_hosts['ip'].to_csv(r'arm-ip.txt', header=False, index=None, sep=' ')
select_hosts.drop(columns=supressed_columns)
hosts['ip'].to_csv(r'ip.txt', header=False, index=None, sep=' ')
# + language="bash"
# export HOME_DIR=$PWD
# echo $HOME_DIR
# source ../common/env.sh
# while read ip; do
# cd ~/repos/node-exporter-collectors
# export DOCKER_HOST="tcp://$ip:2376"
# docker-compose -f docker-compose-raspi3.yaml down 2>/dev/null
# docker-compose -f docker-compose-raspi3.yaml up -d 2>/dev/null
# done <arm-ip.txt
# cd $HOME_DIR
# -
# ### Deploy x86
#
#
hosts[(hosts.arch == "x86_64")]['ip'].to_csv(r'i86-ip.txt', header=False, index=None, sep=' ')
# + language="bash"
# export HOME_DIR=$PWD
# echo $HOME_DIR
# source ../common/env.sh
# while read ip; do
# cd ~/repos/node-exporter-collectors
# export DOCKER_HOST="tcp://$ip:2376"
# docker stop nodeexporter 2>/dev/null
# docker stop monitor_tempsensor_1 2>/dev/null
# docker rm nodeexporter 2>/dev/null
# docker rm monitor_tempsensor_1 2>/dev/null
# docker-compose -f docker-compose-x86.yaml down 2>/dev/null
# docker-compose -f docker-compose-x86.yaml up -d 2>/dev/null
# done <i86-ip.txt
# cd $HOME_DIR
# -
# ### Deploy Tegra(nvidia jetson)
hosts[(hosts.arch == "aarch64")]['ip'].to_csv(r'aarch64-ip.txt', header=False, index=None, sep=' ')
# + language="bash"
# export HOME_DIR=$PWD
# echo $HOME_DIR
# source ../common/env.sh
# while read ip; do
# cd ~/repos/node-exporter-collectors
# export DOCKER_HOST="tcp://$ip:2376"
# docker stop nodeexporter 2>/dev/null
# docker rm nodeexporter 2>/dev/null
# docker-compose -f docker-compose-jetson.yaml down 2>/dev/null
# docker-compose -f docker-compose-jetson.yaml up -d 2>/dev/null
# done <aarch64-ip.txt
# cd $HOME_DIR
# -
# ### Validade Deploy
#
# - `docker ps` all nodes
# - `curl` all nodes
# + language="bash"
# echo "ip|id|image|ports|runnning" > containers.csv
# source ../common/env.sh
# while read ip; do
# export DOCKER_HOST="tcp://$ip:2376"
# docker ps --filter "name=nodeexporter" --format "$ip|{{.ID}}|{{.Image}}|{{.Ports}}|{{.RunningFor}}" >> containers.csv 2>/dev/null || :
# done <ip.txt
# -
containers = pd.read_csv("containers.csv", sep = '|')
pd.merge(containers, hosts, on='ip', how='inner').drop(columns=supressed_columns)
# + language="bash"
# echo "ip,result" > node-exporters.csv
# while read ip; do
# echo "$ip,`curl --connect-timeout 10 -o /dev/null -s -w "%{http_code}\n" http://$ip:9100`" >> node-exporters.csv
# done <ip.txt
# -
containers = pd.read_csv("node-exporters.csv", sep = ',')
pd.merge(containers, hosts, on='ip', how='inner').drop(columns=supressed_columns)
# Observe that some hosts are not `200`, because this are not active servers at this tests.
| 04-deploy-monitor/run-script.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Notebook Examples for Chapter 2
import warnings
# these are innocuous but irritating
warnings.filterwarnings("ignore", message="numpy.dtype size changed")
warnings.filterwarnings("ignore", message="numpy.ufunc size changed")
# %matplotlib inline
# ### Illustrating the central limit theorem
# +
import numpy as np
import matplotlib.pyplot as plt
r = np.random.rand(10000,12)
array = np.sum(r,1)
p=plt.hist(array,bins=12)
#plt.savefig('/home/mort/LaTeX/new projects/CRC4/Chapter2/fig2_1.eps')
# -
# ### Chi-square density function
# +
import scipy.stats as st
z = np.linspace(1,20,200)
ax = plt.subplot(111)
for i in range(1,6):
ax.plot(z,st.chi2.pdf(z,i),label = str(i))
ax.legend()
#plt.savefig('/home/mort/LaTeX/new projects/CRC4/Chapter2/fig2_2.eps')
# -
# ### Sampling
# +
import ee
ee.Initialize()
im = ee.Image(ee.ImageCollection('ASTER/AST_L1T_003') \
.filterBounds(ee.Geometry.Point([6.5,50.9])) \
.filterDate('2007-04-30','2007-05-02') \
.first()) \
.select('B3N')
roi = ee.Geometry.Polygon(
[[6.382713317871094,50.90736285477543],
[6.3961029052734375,50.90130070888041],
[6.4015960693359375,50.90519789328594],
[6.388206481933594,50.91169247570916],
[6.382713317871094,50.90736285477543]])
sample = im.sample(roi,scale=15) \
.aggregate_array('B3N').getInfo()
p = plt.hist(sample,bins=20)
#plt.savefig('/home/mort/LaTeX/new projects/CRC4/Chapter2/fig2_4.eps')
# -
# ### Confidence intervals
# +
from scipy.stats import norm,chi2
def x2(a,m):
return chi2.ppf(1-a,m)
m = 1000
a = 0.05
g = np.random.random(m)
gbar = np.sum(g)/m
s = np.sum((g-gbar)**2)/(m-1)
print 'sample variance: %f'%s
lower = (m-1)*s/x2(a/2,m-1)
upper = (m-1)*s/x2(1-a/2,m-1)
print '%i percent confidence interval: (%f, %f)'\
%(int((1-a)*100),lower,upper)
print 'sample mean: %f'%gbar
t = norm.ppf(1-a/2)
sigma = np.sqrt(s)
lower = gbar-t*sigma/np.sqrt(m)
upper = gbar+t*sigma/np.sqrt(m)
print '%i percent confidence interval: (%f, %f)'\
%(int((1-a)*100),lower,upper)
# -
# ### Provisional means
# +
from osgeo import gdal
from osgeo.gdalconst import GA_ReadOnly
import auxil.auxil1 as auxil
gdal.AllRegister()
infile = 'imagery/AST_20070501'
inDataset = gdal.Open(infile,GA_ReadOnly)
cols = inDataset.RasterXSize
rows = inDataset.RasterYSize
Xs = np.zeros((cols,3))
cpm = auxil.Cpm(3)
rasterBands=[inDataset.GetRasterBand(k+1)
for k in range(3)]
for row in range(rows):
for k in range(3):
Xs[:,k]=rasterBands[k].ReadAsArray(0,row,cols,1)
cpm.update(Xs)
print cpm.covariance()
# -
# ### Multiple linear regression
# +
import numpy as np
# biased data matrix X ( 3 independent variables)
X = np.random.rand(100,3)
X = np.mat(np.append(np.ones((100,1)),X,axis=1))
# a parameter vector
w = np.mat([[3.0],[4.0],[5.0],[6.0]])
# noisy dependent variable y with sigma = 0.1
y = X*w+np.random.normal(0,0.1,(100,1))
# pseudoinverse
Xp = (X.T*X).I*X.T
# estimated parameter vector
w = Xp*y
print w
# +
import tensorflow as tf
# set up computation graph
X1 = tf.constant(X)
y1 = tf.constant(y)
X1T = tf.transpose(X)
X1p = tf.matmul(tf.matrix_inverse(tf.matmul(X1T,X1)),X1T)
w = tf.matmul(X1p,y1)
# create and run a session to evaluate w
with tf.Session() as sess:
w = w.eval()
print w
# +
import ee
ee.Initialize()
# set up JSON description of the calculatiom
X1 = ee.Array(X.tolist())
y1 = ee.Array(y.tolist())
X1T = X1.matrixTranspose()
X1p = X1T.matrixMultiply(X1) \
.matrixInverse() \
.matrixMultiply(X1T)
w = X1p.matrixMultiply(y1)
# run on GEE server
print w.getInfo()
# -
print 0.01*(X.T*X).I
# ### Mutual information
# +
import numpy as np
from osgeo import gdal
from osgeo.gdalconst import GA_ReadOnly
def mi(arr1,arr2):
'''mutual information of two uint8 arrays '''
p12 = np.histogram2d(arr1,arr2,bins=256,
normed=True)[0].ravel()
p1 = np.histogram(arr1,bins=256,normed=True)[0]
p2 = np.histogram(arr2,bins=256,normed=True)[0]
p1p2 = np.outer(p1,p2).ravel()
idx = p12>0
return np.sum(p12[idx]*np.log(p12[idx]/p1p2[idx]))
gdal.AllRegister()
infile = 'imagery/AST_20070501'
inDataset = gdal.Open(infile,GA_ReadOnly)
cols = inDataset.RasterXSize
rows = inDataset.RasterYSize
image = np.zeros((3,rows*cols))
# VNIR bands
for b in range(3):
band = inDataset.GetRasterBand(b+1)
image[b,:]=np.byte(band.ReadAsArray(0,0,cols,rows))\
.ravel()
inDataset = None
print mi(image[0,:],image[1,:])
print mi(image[0,:],image[2,:])
print mi(image[1,:],image[2,:])
| src/Chapter2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="K_YntzlsRNWx"
# # Digit Recognizer
#
# <NAME>.
#
# A Deep Learning CNN based model which can recognize handwritten digits. It is built on machine learning framework Tensorflow along with Keras. The model architecture is slighlty modified version of [LeNet-5](http://yann.lecun.com/exdb/publis/pdf/lecun-01a.pdf) Architecture proposed by Yann LeCun in 1998. This is Deep Convolutional Neural Network of 5 layers. This notebook is created in and for running the notebook in [Google Colab](https://colab.research.google.com). It takes approximately 2mins to train on Colab's Tesla GPU which Google offers for free.
#
# ## **Table of Contents**
#
#
# 1. **Importing Dependencies**
# 2. **Data Preparation**
# * **2.1 Loading Dataset**
# * **2.2 Check for null and missing values**
# * **2.3 Normalisation**
# * **2.4 Reshape**
# * **2.5 Label Encoding**
# * **2.6 Train and Validation Data Split**
# 3. **CNN Model**
# * **3.1 Model Architecture Definition**
# * **3.2 Optimizer and Annealer Initialisation**
# * **3.3 Data Augmentation**
# 4. **Model Training and Evaluation**
# * **4.1 Model Training**
# * **4.2 Learning Curve**
# * **4.2 Confusion Matrix**
# * **4.3 Displaying the errors**
# 5. **Model Save**
#
#
#
#
#
#
#
# + [markdown] colab_type="text" id="zQHYkAoVUMKi"
# ## 1. Importing Dependencies
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="LYlQnHuARMFQ" outputId="85c52ef4-7143-46e7-eed0-ab97fd27eaa3"
# Importing Tensorflow and keras
#Keras is built into TF 2.0
import tensorflow as tf
from tensorflow.keras import layers
from tensorflow.keras.preprocessing.image import ImageDataGenerator
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
import matplotlib.image as img
# %matplotlib inline
np.random.seed(2)
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix
import itertools
#Setting the Theme of the data visualizer Seaborn
sns.set(style="white",context="notebook",palette="deep")
#Tensorflow Version
print("Tensorflow Version:"+tf.version.VERSION)
print("Keras Version "+tf.keras.__version__)
# + [markdown] colab_type="text" id="WEPDqrJLUWEi"
# ##2.Data Preparation
# + [markdown] colab_type="text" id="ISCeQ8sr9e5q"
# ###2.1 Data Loading
# + colab={"base_uri": "https://localhost:8080/", "height": 119} colab_type="code" id="kJQAzoToY-Eb" outputId="f2f76a37-09ef-4ead-923b-f41b2d517ca1"
import os
os.environ['KAGGLE_USERNAME'] = "xxxxxx" # username from the json file
os.environ['KAGGLE_KEY'] = "<KEY>" # key from the json file
# !kaggle competitions download -c digit-recognizer
# !unzip train.csv.zip
# + colab={} colab_type="code" id="UutK5S_rUGmH"
train = pd.read_csv("/content/drive/My Drive/Digit Recognition Project/train.csv")
# + colab={"base_uri": "https://localhost:8080/", "height": 472} colab_type="code" id="05yoL078UJIp" outputId="16e5b167-7439-4b5d-e77d-5acec88a1e6c"
Y_train = train["label"]
#Dropping Label Column
X_train = train.drop(labels=["label"],axis=1)
#free up some space
del train
graph = sns.countplot(Y_train)
Y_train.value_counts()
# + [markdown] colab_type="text" id="Vg7ysL049yLh"
# ###2.2 Checking for Null or NaN values
# + colab={"base_uri": "https://localhost:8080/", "height": 102} colab_type="code" id="jGSazKXLXRtA" outputId="526d09de-fd1e-4a4e-c60a-d485db15e9d3"
#Checking for any null or missing values
X_train.isnull().any().describe()
test.isnull().any().describe()
# + [markdown] colab_type="text" id="xJQsCTooXv1X"
# ###2.3 Normalisation
# + colab={} colab_type="code" id="pTFl1nLqXpSy"
X_train = X_train/255
test = test/255
# + [markdown] colab_type="text" id="0Lx2A8K8YAKn"
# ###2.4 Reshape
# + colab={} colab_type="code" id="RgJra9a4X9U1"
X_train = X_train.values.reshape(-1,28,28,1)
test = test.values.reshape(-1,28,28,1)
# + [markdown] colab_type="text" id="GSbzJTHlYkhS"
# ### 2.5 Label Encoding
# + colab={} colab_type="code" id="U7DvS95KYrVa"
Y_train = tf.keras.utils.to_categorical(Y_train, num_classes=10)
#To enable label into hot vector. For Eg.7 -> [0,0,0,0,0,0,0,1,0,0]
# + colab={} colab_type="code" id="HUz1lMAAZLFC"
#Spliting Train and test set
random_seed =2
# + [markdown] colab_type="text" id="15zkjcF2-Hek"
# ### 2.6 Train and Validation Dataset Split
# + colab={} colab_type="code" id="UUBksBY12L8V"
X_train,X_val,Y_train,Y_val = train_test_split(X_train,Y_train,test_size=0.1,
random_state = random_seed)
# + colab={"base_uri": "https://localhost:8080/", "height": 268} colab_type="code" id="KVNgcZ5F3Ap5" outputId="d2f4f9e8-74b6-404b-adf0-353f591d5ded"
#Show some example
g = plt.imshow(X_train[0][:,:,0])
# + [markdown] colab_type="text" id="tU1XRCKr3e0n"
# ##3.CNN Model
# + [markdown] colab_type="text" id="CtmNzQO8-T7f"
# ### 3.1 Model architecture Definition
# + colab={} colab_type="code" id="kpkANohy3TaJ"
#CNN Architecture is In -> [[Conv2D->relu]*2 -> MaxPool2D -> Dropout]*2 ->
#Flatten -> Dense -> Dropout -> Out
model = tf.keras.Sequential()
model.add(layers.Conv2D(filters=32, kernel_size=(5,5), padding='Same',
activation=tf.nn.relu, input_shape = (28,28,1)))
model.add(layers.Conv2D(filters=32, kernel_size=(5,5), padding='Same',
activation=tf.nn.relu))
model.add(layers.MaxPool2D(pool_size=(2,2)))
model.add(layers.Dropout(0.25))
model.add(layers.Conv2D(filters=64, kernel_size=(3,3), padding='Same',
activation=tf.nn.relu))
model.add(layers.Conv2D(filters=64, kernel_size=(3,3), padding='Same',
activation=tf.nn.relu))
model.add(layers.MaxPool2D(pool_size=(2,2),strides=(2,2)))
model.add(layers.Dropout(0.25))
model.add(layers.Flatten())
model.add(layers.Dense(256,activation=tf.nn.relu))
model.add(layers.Dropout(0.25))
model.add(layers.Dense(10,activation=tf.nn.softmax))
# + [markdown] colab_type="text" id="eouhmHGN-avP"
# ### 3.2 Optimiser and Annealer Initialisation
# + colab={} colab_type="code" id="VuN1-e0uPCWy"
#Defining Optimizer
optimizer = tf.keras.optimizers.RMSprop(lr=0.001, rho=0.9, epsilon=1e-08, decay=0.0)
# + colab={} colab_type="code" id="wiwSpyiYPcIb"
#Compiling Model
model.compile(optimizer = optimizer, loss='categorical_crossentropy',
metrics=["accuracy"])
# + colab={} colab_type="code" id="FqExgW5DQoab"
#Setting Learning rate annealer
learning_rate_reduction = tf.keras.callbacks.ReduceLROnPlateau(monitor='val_acc',
patience=3,
verbose=1,
factor=0.5,
min_lr=0.00001)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="VpoFy1wQRJf8" outputId="5e35141c-30d9-4d69-ab74-bbab379e9a05"
epochs=30
batch_size = 86
print(X_train.shape[0])
# + [markdown] colab_type="text" id="E573js59RQVn"
# ### 3.3 Data Augmentation
# + colab={} colab_type="code" id="lTi2iTMKROwt"
datagen = ImageDataGenerator(
featurewise_center=False, # set input mean to 0 over the dataset
samplewise_center=False, # set each sample mean to 0
featurewise_std_normalization=False, # divide inputs by std of the dataset
samplewise_std_normalization=False, # divide each input by its std
zca_whitening=False, # apply ZCA whitening
rotation_range=10, # randomly rotate images in the range (degrees, 0 to 180)
zoom_range = 0.1, # Randomly zoom image
width_shift_range=0.1, # randomly shift images horizontally (fraction of total width)
height_shift_range=0.1, # randomly shift images vertically (fraction of total height)
horizontal_flip=False, # randomly flip images
vertical_flip=False) # randomly flip images
datagen.fit(X_train)
# + [markdown] colab_type="text" id="qkX93W-P4iYl"
# ## 4. Model Training and Evaluation
# + [markdown] colab_type="text" id="Dn6FLr-r_deO"
# ### 4.1 Model Training
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="BYvD4vpOz3_L" outputId="13407804-4096-4cf7-e8e9-f0997dc46723"
# Fit the model
history = model.fit_generator(datagen.flow(X_train,Y_train, batch_size=batch_size),
epochs = epochs, validation_data = (X_val,Y_val),
verbose = 1, steps_per_epoch=X_train.shape[0] // batch_size
, callbacks=[learning_rate_reduction])
# + [markdown] colab_type="text" id="NqfDeO7v_Akp"
# ### 4.2 Learning Curve
# + colab={"base_uri": "https://localhost:8080/", "height": 268} colab_type="code" id="PMKZhU75z-b6" outputId="c442dbcb-88d1-49bf-ecd5-b0d6edc618db"
# Plot the loss and accuracy curves for training and validation
fig, ax = plt.subplots(2,1)
ax[0].plot(history.history['loss'], color='b', label="Training loss")
ax[0].plot(history.history['val_loss'], color='r', label="validation loss",axes =ax[0])
legend = ax[0].legend(loc='best', shadow=True)
ax[1].plot(history.history['acc'], color='b', label="Training accuracy")
ax[1].plot(history.history['val_acc'], color='r',label="Validation accuracy")
legend = ax[1].legend(loc='best', shadow=True)
# + [markdown] colab_type="text" id="oTsAl3Ur3n9d"
# ### 4.3 Confusion Matrix Plotting
# It is used for grphical representation of performance of the model. It shows the performance of Model in predicting every class.
# + _cell_guid="11361e73-8250-4bf5-a353-b0f8ea83e659" _execution_state="idle" _uuid="16e161179bf1b51ba66c39b2cead883f1db3a9c7" colab={"base_uri": "https://localhost:8080/", "height": 310} colab_type="code" id="IK6-TyagG4G4" outputId="d104ce2d-b241-43cd-9632-0963e0c6e7d2"
# Look at confusion matrix
def plot_confusion_matrix(cm, classes,
normalize=False,
title='Confusion matrix',
cmap=plt.cm.Blues):
"""
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, cm[i, j],
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
# Predict the values from the validation dataset
Y_pred = model.predict(X_val)
# Convert predictions classes to one hot vectors
Y_pred_classes = np.argmax(Y_pred,axis = 1)
# Convert validation observations to one hot vectors
Y_true = np.argmax(Y_val,axis = 1)
# compute the confusion matrix
confusion_mtx = confusion_matrix(Y_true, Y_pred_classes)
# plot the confusion matrix
plot_confusion_matrix(confusion_mtx, classes = range(10))
# + [markdown] colab_type="text" id="DZqd6fvr_Y9n"
# ### 4.4 Displaying Errors
# + _cell_guid="7b0f31b8-c18b-4529-b0d8-eb4c31e30bbf" _execution_state="idle" _uuid="e7a3d6449b499a29db224e42e950f21ca1ec4e36" colab={"base_uri": "https://localhost:8080/", "height": 283} colab_type="code" id="tV4wnLTQG4G8" outputId="89851798-528c-4f2a-82da-740b7747911d"
# Display some error results
# Errors are difference between predicted labels and true labels
errors = (Y_pred_classes - Y_true != 0)
Y_pred_classes_errors = Y_pred_classes[errors]
Y_pred_errors = Y_pred[errors]
Y_true_errors = Y_true[errors]
X_val_errors = X_val[errors]
def display_errors(errors_index,img_errors,pred_errors, obs_errors):
""" This function shows 6 images with their predicted and real labels"""
n = 0
nrows = 2
ncols = 3
fig, ax = plt.subplots(nrows,ncols,sharex=True,sharey=True)
for row in range(nrows):
for col in range(ncols):
error = errors_index[n]
ax[row,col].imshow((img_errors[error]).reshape((28,28)))
ax[row,col].set_title("Predicted :{} True :{}".format(pred_errors[error],obs_errors[error]))
n += 1
# Probabilities of the wrong predicted numbers
Y_pred_errors_prob = np.max(Y_pred_errors,axis = 1)
# Predicted probabilities of the true values in the error set
true_prob_errors = np.diagonal(np.take(Y_pred_errors, Y_true_errors, axis=1))
# Difference between the probability of the predicted label and the true label
delta_pred_true_errors = Y_pred_errors_prob - true_prob_errors
# Sorted list of the delta prob errors
sorted_dela_errors = np.argsort(delta_pred_true_errors)
# Top 6 errors
most_important_errors = sorted_dela_errors[-6:]
# Show the top 6 errors
display_errors(most_important_errors, X_val_errors, Y_pred_classes_errors, Y_true_errors)
# + [markdown] colab_type="text" id="xnxxdQbm_hhi"
# ## 5. Model Saving
# + colab={} colab_type="code" id="Rn9BMVz8CHT_"
model.save('model_python.hdf5')
# + colab={"base_uri": "https://localhost:8080/", "height": 751} colab_type="code" id="wGCmpb2Pr9TP" outputId="4625f63e-38e9-4a45-e668-42db6ef9cbe8"
# !pip install tensorflowjs
import tensorflowjs as tfjs
tfjs.converters.save_keras_model(model, '/content/models')
# + colab={} colab_type="code" id="p0RaC1frZ-of"
| digit-recognizer-notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# # Time-related feature engineering
#
# This notebook introduces different strategies to leverage time-related features
# for a bike sharing demand regression task that is highly dependent on business
# cycles (days, weeks, months) and yearly season cycles.
#
# In the process, we introduce how to perform periodic feature engineering using
# the :class:`sklearn.preprocessing.SplineTransformer` class and its
# `extrapolation="periodic"` option.
#
# ## Data exploration on the Bike Sharing Demand dataset
#
# We start by loading the data from the OpenML repository.
#
#
# +
from sklearn.datasets import fetch_openml
bike_sharing = fetch_openml("Bike_Sharing_Demand", version=2, as_frame=True)
df = bike_sharing.frame
# -
# To get a quick understanding of the periodic patterns of the data, let us
# have a look at the average demand per hour during a week.
#
# Note that the week starts on a Sunday, during the weekend. We can clearly
# distinguish the commute patterns in the morning and evenings of the work days
# and the leisure use of the bikes on the weekends with a more spread peak
# demand around the middle of the days:
#
#
# +
import matplotlib.pyplot as plt
fig, ax = plt.subplots(figsize=(12, 4))
average_week_demand = df.groupby(["weekday", "hour"]).mean()["count"]
average_week_demand.plot(ax=ax)
_ = ax.set(
title="Average hourly bike demand during the week",
xticks=[i * 24 for i in range(7)],
xticklabels=["Sun", "Mon", "Tue", "Wed", "Thu", "Fri", "Sat"],
xlabel="Time of the week",
ylabel="Number of bike rentals",
)
# -
# The target of the prediction problem is the absolute count of bike rentals on
# a hourly basis:
#
#
df["count"].max()
# Let us rescale the target variable (number of hourly bike rentals) to predict
# a relative demand so that the mean absolute error is more easily interpreted
# as a fraction of the maximum demand.
#
# <div class="alert alert-info"><h4>Note</h4><p>The fit method of the models used in this notebook all minimize the
# mean squared error to estimate the conditional mean instead of the mean
# absolute error that would fit an estimator of the conditional median.
#
# When reporting performance measure on the test set in the discussion, we
# instead choose to focus on the mean absolute error that is more
# intuitive than the (root) mean squared error. Note, however, that the
# best models for one metric are also the best for the other in this
# study.</p></div>
#
#
y = df["count"] / df["count"].max()
fig, ax = plt.subplots(figsize=(12, 4))
y.hist(bins=30, ax=ax)
_ = ax.set(
xlabel="Fraction of rented fleet demand",
ylabel="Number of hours",
)
# The input feature data frame is a time annotated hourly log of variables
# describing the weather conditions. It includes both numerical and categorical
# variables. Note that the time information has already been expanded into
# several complementary columns.
#
#
#
X = df.drop("count", axis="columns")
X
# <div class="alert alert-info"><h4>Note</h4><p>If the time information was only present as a date or datetime column, we
# could have expanded it into hour-in-the-day, day-in-the-week,
# day-in-the-month, month-in-the-year using pandas:
# https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#time-date-components</p></div>
#
# We now introspect the distribution of the categorical variables, starting
# with `"weather"`:
#
#
#
X["weather"].value_counts()
# Since there are only 3 `"heavy_rain"` events, we cannot use this category to
# train machine learning models with cross validation. Instead, we simplify the
# representation by collapsing those into the `"rain"` category.
#
#
#
X["weather"].replace(to_replace="heavy_rain", value="rain", inplace=True)
X["weather"].value_counts()
# As expected, the `"season"` variable is well balanced:
#
#
#
X["season"].value_counts()
# ## Time-based cross-validation
#
# Since the dataset is a time-ordered event log (hourly demand), we will use a
# time-sensitive cross-validation splitter to evaluate our demand forecasting
# model as realistically as possible. We use a gap of 2 days between the train
# and test side of the splits. We also limit the training set size to make the
# performance of the CV folds more stable.
#
# 1000 test datapoints should be enough to quantify the performance of the
# model. This represents a bit less than a month and a half of contiguous test
# data:
#
#
# +
from sklearn.model_selection import TimeSeriesSplit
ts_cv = TimeSeriesSplit(
n_splits=5,
gap=48,
max_train_size=10000,
test_size=1000,
)
# -
# Let us manually inspect the various splits to check that the
# `TimeSeriesSplit` works as we expect, starting with the first split:
#
#
all_splits = list(ts_cv.split(X, y))
train_0, test_0 = all_splits[0]
X.iloc[test_0]
X.iloc[train_0]
# We now inspect the last split:
#
#
train_4, test_4 = all_splits[4]
X.iloc[test_4]
X.iloc[train_4]
# All is well. We are now ready to do some predictive modeling!
#
# ## Gradient Boosting
#
# Gradient Boosting Regression with decision trees is often flexible enough to
# efficiently handle heteorogenous tabular data with a mix of categorical and
# numerical features as long as the number of samples is large enough.
#
# Here, we do minimal ordinal encoding for the categorical variables and then
# let the model know that it should treat those as categorical variables by
# using a dedicated tree splitting rule. Since we use an ordinal encoder, we
# pass the list of categorical values explicitly to use a logical order when
# encoding the categories as integers instead of the lexicographical order.
# This also has the added benefit of preventing any issue with unknown
# categories when using cross-validation.
#
# The numerical variables need no preprocessing and, for the sake of simplicity,
# we only try the default hyper-parameters for this model:
#
#
# +
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import OrdinalEncoder
from sklearn.compose import ColumnTransformer
from sklearn.ensemble import HistGradientBoostingRegressor
from sklearn.model_selection import cross_validate
categorical_columns = [
"weather",
"season",
"holiday",
"workingday",
]
categories = [
["clear", "misty", "rain"],
["spring", "summer", "fall", "winter"],
["False", "True"],
["False", "True"],
]
ordinal_encoder = OrdinalEncoder(categories=categories)
gbrt_pipeline = make_pipeline(
ColumnTransformer(
transformers=[
("categorical", ordinal_encoder, categorical_columns),
],
remainder="passthrough",
),
HistGradientBoostingRegressor(
categorical_features=range(4),
),
)
# -
# Lets evaluate our gradient boosting model with the mean absolute error of the
# relative demand averaged across our 5 time-based cross-validation splits:
#
#
# +
def evaluate(model, X, y, cv):
cv_results = cross_validate(
model,
X,
y,
cv=cv,
scoring=["neg_mean_absolute_error", "neg_root_mean_squared_error"],
)
mae = -cv_results["test_neg_mean_absolute_error"]
rmse = -cv_results["test_neg_root_mean_squared_error"]
print(
f"Mean Absolute Error: {mae.mean():.3f} +/- {mae.std():.3f}\n"
f"Root Mean Squared Error: {rmse.mean():.3f} +/- {rmse.std():.3f}"
)
evaluate(gbrt_pipeline, X, y, cv=ts_cv)
# -
# This model has an average error around 4 to 5% of the maximum demand. This is
# quite good for a first trial without any hyper-parameter tuning! We just had
# to make the categorical variables explicit. Note that the time related
# features are passed as is, i.e. without processing them. But this is not much
# of a problem for tree-based models as they can learn a non-monotonic
# relationship between ordinal input features and the target.
#
# This is not the case for linear regression models as we will see in the
# following.
#
# ## Naive linear regression
#
# As usual for linear models, categorical variables need to be one-hot encoded.
# For consistency, we scale the numerical features to the same 0-1 range using
# class:`sklearn.preprocessing.MinMaxScaler`, although in this case it does not
# impact the results much because they are already on comparable scales:
#
#
# +
from sklearn.preprocessing import OneHotEncoder
from sklearn.preprocessing import MinMaxScaler
from sklearn.linear_model import RidgeCV
import numpy as np
one_hot_encoder = OneHotEncoder(handle_unknown="ignore", sparse=False)
alphas = np.logspace(-6, 6, 25)
naive_linear_pipeline = make_pipeline(
ColumnTransformer(
transformers=[
("categorical", one_hot_encoder, categorical_columns),
],
remainder=MinMaxScaler(),
),
RidgeCV(alphas=alphas),
)
evaluate(naive_linear_pipeline, X, y, cv=ts_cv)
# -
# The performance is not good: the average error is around 14% of the maximum
# demand. This is more than three times higher than the average error of the
# gradient boosting model. We can suspect that the naive original encoding
# (merely min-max scaled) of the periodic time-related features might prevent
# the linear regression model to properly leverage the time information: linear
# regression does not automatically model non-monotonic relationships between
# the input features and the target. Non-linear terms have to be engineered in
# the input.
#
# For example, the raw numerical encoding of the `"hour"` feature prevents the
# linear model from recognizing that an increase of hour in the morning from 6
# to 8 should have a strong positive impact on the number of bike rentals while
# an increase of similar magnitude in the evening from 18 to 20 should have a
# strong negative impact on the predicted number of bike rentals.
#
# ## Time-steps as categories
#
# Since the time features are encoded in a discrete manner using integers (24
# unique values in the "hours" feature), we could decide to treat those as
# categorical variables using a one-hot encoding and thereby ignore any
# assumption implied by the ordering of the hour values.
#
# Using one-hot encoding for the time features gives the linear model a lot
# more flexibility as we introduce one additional feature per discrete time
# level.
#
#
# +
one_hot_linear_pipeline = make_pipeline(
ColumnTransformer(
transformers=[
("categorical", one_hot_encoder, categorical_columns),
("one_hot_time", one_hot_encoder, ["hour", "weekday", "month"]),
],
remainder=MinMaxScaler(),
),
RidgeCV(alphas=alphas),
)
evaluate(one_hot_linear_pipeline, X, y, cv=ts_cv)
# -
# The average error rate of this model is 10% which is much better than using
# the original (ordinal) encoding of the time feature, confirming our intuition
# that the linear regression model benefits from the added flexibility to not
# treat time progression in a monotonic manner.
#
# However, this introduces a very large number of new features. If the time of
# the day was represented in minutes since the start of the day instead of
# hours, one-hot encoding would have introduced 1440 features instead of 24.
# This could cause some significant overfitting. To avoid this we could use
# :func:`sklearn.preprocessing.KBinsDiscretizer` instead to re-bin the number
# of levels of fine-grained ordinal or numerical variables while still
# benefitting from the non-monotonic expressivity advantages of one-hot
# encoding.
#
# Finally, we also observe that one-hot encoding completely ignores the
# ordering of the hour levels while this could be an interesting inductive bias
# to preserve to some level. In the following we try to explore smooth,
# non-monotonic encoding that locally preserves the relative ordering of time
# features.
#
# ## Trigonometric features
#
# As a first attempt, we can try to encode each of those periodic features
# using a sine and cosine transformation with the matching period.
#
# Each ordinal time feature is transformed into 2 features that together encode
# equivalent information in a non-monotonic way, and more importantly without
# any jump between the first and the last value of the periodic range.
#
#
# +
from sklearn.preprocessing import FunctionTransformer
def sin_transformer(period):
return FunctionTransformer(lambda x: np.sin(x / period * 2 * np.pi))
def cos_transformer(period):
return FunctionTransformer(lambda x: np.cos(x / period * 2 * np.pi))
# -
# Let us visualize the effect of this feature expansion on some synthetic hour
# data with a bit of extrapolation beyond hour=23:
#
#
# +
import pandas as pd
hour_df = pd.DataFrame(
np.arange(26).reshape(-1, 1),
columns=["hour"],
)
hour_df["hour_sin"] = sin_transformer(24).fit_transform(hour_df)["hour"]
hour_df["hour_cos"] = cos_transformer(24).fit_transform(hour_df)["hour"]
hour_df.plot(x="hour")
_ = plt.title("Trigonometric encoding for the 'hour' feature")
# -
# Let's use a 2D scatter plot with the hours encoded as colors to better see
# how this representation maps the 24 hours of the day to a 2D space, akin to
# some sort of a 24 hour version of an analog clock. Note that the "25th" hour
# is mapped back to the 1st hour because of the periodic nature of the
# sine/cosine representation.
#
#
fig, ax = plt.subplots(figsize=(7, 5))
sp = ax.scatter(hour_df["hour_sin"], hour_df["hour_cos"], c=hour_df["hour"])
ax.set(
xlabel="sin(hour)",
ylabel="cos(hour)",
)
_ = fig.colorbar(sp)
# We can now build a feature extraction pipeline using this strategy:
#
#
cyclic_cossin_transformer = ColumnTransformer(
transformers=[
("categorical", one_hot_encoder, categorical_columns),
("month_sin", sin_transformer(12), ["month"]),
("month_cos", cos_transformer(12), ["month"]),
("weekday_sin", sin_transformer(7), ["weekday"]),
("weekday_cos", cos_transformer(7), ["weekday"]),
("hour_sin", sin_transformer(24), ["hour"]),
("hour_cos", cos_transformer(24), ["hour"]),
],
remainder=MinMaxScaler(),
)
cyclic_cossin_linear_pipeline = make_pipeline(
cyclic_cossin_transformer,
RidgeCV(alphas=alphas),
)
evaluate(cyclic_cossin_linear_pipeline, X, y, cv=ts_cv)
# The performance of our linear regression model with this simple feature
# engineering is a bit better than using the original ordinal time features but
# worse than using the one-hot encoded time features. We will further analyze
# possible reasons for this disappointing outcome at the end of this notebook.
#
# ## Periodic spline features
#
# We can try an alternative encoding of the periodic time-related features
# using spline transformations with a large enough number of splines, and as a
# result a larger number of expanded features compared to the sine/cosine
# transformation:
#
#
# +
from sklearn.preprocessing import SplineTransformer
def periodic_spline_transformer(period, n_splines=None, degree=3):
if n_splines is None:
n_splines = period
n_knots = n_splines + 1 # periodic and include_bias is True
return SplineTransformer(
degree=degree,
n_knots=n_knots,
knots=np.linspace(0, period, n_knots).reshape(n_knots, 1),
extrapolation="periodic",
include_bias=True,
)
# -
# Again, let us visualize the effect of this feature expansion on some
# synthetic hour data with a bit of extrapolation beyond hour=23:
#
#
hour_df = pd.DataFrame(
np.linspace(0, 26, 1000).reshape(-1, 1),
columns=["hour"],
)
splines = periodic_spline_transformer(24, n_splines=12).fit_transform(hour_df)
splines_df = pd.DataFrame(
splines,
columns=[f"spline_{i}" for i in range(splines.shape[1])],
)
pd.concat([hour_df, splines_df], axis="columns").plot(x="hour", cmap=plt.cm.tab20b)
_ = plt.title("Periodic spline-based encoding for the 'hour' feature")
# Thanks to the use of the `extrapolation="periodic"` parameter, we observe
# that the feature encoding stays smooth when extrapolating beyond midnight.
#
# We can now build a predictive pipeline using this alternative periodic
# feature engineering strategy.
#
# It is possible to use fewer splines than discrete levels for those ordinal
# values. This makes spline-based encoding more efficient than one-hot encoding
# while preserving most of the expressivity:
#
#
cyclic_spline_transformer = ColumnTransformer(
transformers=[
("categorical", one_hot_encoder, categorical_columns),
("cyclic_month", periodic_spline_transformer(12, n_splines=6), ["month"]),
("cyclic_weekday", periodic_spline_transformer(7, n_splines=3), ["weekday"]),
("cyclic_hour", periodic_spline_transformer(24, n_splines=12), ["hour"]),
],
remainder=MinMaxScaler(),
)
cyclic_spline_linear_pipeline = make_pipeline(
cyclic_spline_transformer,
RidgeCV(alphas=alphas),
)
evaluate(cyclic_spline_linear_pipeline, X, y, cv=ts_cv)
# Spline features make it possible for the linear model to successfully
# leverage the periodic time-related features and reduce the error from ~14% to
# ~10% of the maximum demand, which is similar to what we observed with the
# one-hot encoded features.
#
# ## Qualitative analysis of the impact of features on linear model predictions
#
# Here, we want to visualize the impact of the feature engineering choices on
# the time related shape of the predictions.
#
# To do so we consider an arbitrary time-based split to compare the predictions
# on a range of held out data points.
#
#
# +
naive_linear_pipeline.fit(X.iloc[train_0], y.iloc[train_0])
naive_linear_predictions = naive_linear_pipeline.predict(X.iloc[test_0])
one_hot_linear_pipeline.fit(X.iloc[train_0], y.iloc[train_0])
one_hot_linear_predictions = one_hot_linear_pipeline.predict(X.iloc[test_0])
cyclic_cossin_linear_pipeline.fit(X.iloc[train_0], y.iloc[train_0])
cyclic_cossin_linear_predictions = cyclic_cossin_linear_pipeline.predict(X.iloc[test_0])
cyclic_spline_linear_pipeline.fit(X.iloc[train_0], y.iloc[train_0])
cyclic_spline_linear_predictions = cyclic_spline_linear_pipeline.predict(X.iloc[test_0])
# -
# We visualize those predictions by zooming on the last 96 hours (4 days) of
# the test set to get some qualitative insights:
#
#
last_hours = slice(-96, None)
fig, ax = plt.subplots(figsize=(12, 4))
fig.suptitle("Predictions by linear models")
ax.plot(
y.iloc[test_0].values[last_hours],
"x-",
alpha=0.2,
label="Actual demand",
color="black",
)
ax.plot(naive_linear_predictions[last_hours], "x-", label="Ordinal time features")
ax.plot(
cyclic_cossin_linear_predictions[last_hours],
"x-",
label="Trigonometric time features",
)
ax.plot(
cyclic_spline_linear_predictions[last_hours],
"x-",
label="Spline-based time features",
)
ax.plot(
one_hot_linear_predictions[last_hours],
"x-",
label="One-hot time features",
)
_ = ax.legend()
# We can draw the following conclusions from the above plot:
#
# - The **raw ordinal time-related features** are problematic because they do
# not capture the natural periodicity: we observe a big jump in the
# predictions at the end of each day when the hour features goes from 23 back
# to 0. We can expect similar artifacts at the end of each week or each year.
#
# - As expected, the **trigonometric features** (sine and cosine) do not have
# these discontinuities at midnight, but the linear regression model fails to
# leverage those features to properly model intra-day variations.
# Using trigonometric features for higher harmonics or additional
# trigonometric features for the natural period with different phases could
# potentially fix this problem.
#
# - the **periodic spline-based features** fix those two problems at once: they
# give more expressivity to the linear model by making it possible to focus
# on specific hours thanks to the use of 12 splines. Furthermore the
# `extrapolation="periodic"` option enforces a smooth representation between
# `hour=23` and `hour=0`.
#
# - The **one-hot encoded features** behave similarly to the periodic
# spline-based features but are more spiky: for instance they can better
# model the morning peak during the week days since this peak lasts shorter
# than an hour. However, we will see in the following that what can be an
# advantage for linear models is not necessarily one for more expressive
# models.
#
#
# We can also compare the number of features extracted by each feature
# engineering pipeline:
#
#
naive_linear_pipeline[:-1].transform(X).shape
one_hot_linear_pipeline[:-1].transform(X).shape
cyclic_cossin_linear_pipeline[:-1].transform(X).shape
cyclic_spline_linear_pipeline[:-1].transform(X).shape
# This confirms that the one-hot encoding and the spline encoding strategies
# create a lot more features for the time representation than the alternatives,
# which in turn gives the downstream linear model more flexibility (degrees of
# freedom) to avoid underfitting.
#
# Finally, we observe that none of the linear models can approximate the true
# bike rentals demand, especially for the peaks that can be very sharp at rush
# hours during the working days but much flatter during the week-ends: the most
# accurate linear models based on splines or one-hot encoding tend to forecast
# peaks of commuting-related bike rentals even on the week-ends and
# under-estimate the commuting-related events during the working days.
#
# These systematic prediction errors reveal a form of under-fitting and can be
# explained by the lack of interactions terms between features, e.g.
# "workingday" and features derived from "hours". This issue will be addressed
# in the following section.
#
#
# ## Modeling pairwise interactions with splines and polynomial features
#
# Linear models do not automatically capture interaction effects between input
# features. It does not help that some features are marginally non-linear as is
# the case with features constructed by `SplineTransformer` (or one-hot
# encoding or binning).
#
# However, it is possible to use the `PolynomialFeatures` class on coarse
# grained spline encoded hours to model the "workingday"/"hours" interaction
# explicitly without introducing too many new variables:
#
#
# +
from sklearn.preprocessing import PolynomialFeatures
from sklearn.pipeline import FeatureUnion
hour_workday_interaction = make_pipeline(
ColumnTransformer(
[
("cyclic_hour", periodic_spline_transformer(24, n_splines=8), ["hour"]),
("workingday", FunctionTransformer(lambda x: x == "True"), ["workingday"]),
]
),
PolynomialFeatures(degree=2, interaction_only=True, include_bias=False),
)
# -
# Those features are then combined with the ones already computed in the
# previous spline-base pipeline. We can observe a nice performance improvemnt
# by modeling this pairwise interaction explicitly:
#
#
cyclic_spline_interactions_pipeline = make_pipeline(
FeatureUnion(
[
("marginal", cyclic_spline_transformer),
("interactions", hour_workday_interaction),
]
),
RidgeCV(alphas=alphas),
)
evaluate(cyclic_spline_interactions_pipeline, X, y, cv=ts_cv)
# ## Modeling non-linear feature interactions with kernels
#
# The previous analysis highlighted the need to model the interactions between
# `"workingday"` and `"hours"`. Another example of a such a non-linear
# interaction that we would like to model could be the impact of the rain that
# might not be the same during the working days and the week-ends and holidays
# for instance.
#
# To model all such interactions, we could either use a polynomial expansion on
# all marginal features at once, after their spline-based expansion. However,
# this would create a quadratic number of features which can cause overfitting
# and computational tractability issues.
#
# Alternatively, we can use the Nyström method to compute an approximate
# polynomial kernel expansion. Let us try the latter:
#
#
# +
from sklearn.kernel_approximation import Nystroem
cyclic_spline_poly_pipeline = make_pipeline(
cyclic_spline_transformer,
Nystroem(kernel="poly", degree=2, n_components=300, random_state=0),
RidgeCV(alphas=alphas),
)
evaluate(cyclic_spline_poly_pipeline, X, y, cv=ts_cv)
# -
# We observe that this model can almost rival the performance of the gradient
# boosted trees with an average error around 5% of the maximum demand.
#
# Note that while the final step of this pipeline is a linear regression model,
# the intermediate steps such as the spline feature extraction and the Nyström
# kernel approximation are highly non-linear. As a result the compound pipeline
# is much more expressive than a simple linear regression model with raw features.
#
# For the sake of completeness, we also evaluate the combination of one-hot
# encoding and kernel approximation:
#
#
one_hot_poly_pipeline = make_pipeline(
ColumnTransformer(
transformers=[
("categorical", one_hot_encoder, categorical_columns),
("one_hot_time", one_hot_encoder, ["hour", "weekday", "month"]),
],
remainder="passthrough",
),
Nystroem(kernel="poly", degree=2, n_components=300, random_state=0),
RidgeCV(alphas=alphas),
)
evaluate(one_hot_poly_pipeline, X, y, cv=ts_cv)
# While one-hot encoded features were competitive with spline-based features
# when using linear models, this is no longer the case when using a low-rank
# approximation of a non-linear kernel: this can be explained by the fact that
# spline features are smoother and allow the kernel approximation to find a
# more expressive decision function.
#
# Let us now have a qualitative look at the predictions of the kernel models
# and of the gradient boosted trees that should be able to better model
# non-linear interactions between features:
#
#
# +
gbrt_pipeline.fit(X.iloc[train_0], y.iloc[train_0])
gbrt_predictions = gbrt_pipeline.predict(X.iloc[test_0])
one_hot_poly_pipeline.fit(X.iloc[train_0], y.iloc[train_0])
one_hot_poly_predictions = one_hot_poly_pipeline.predict(X.iloc[test_0])
cyclic_spline_poly_pipeline.fit(X.iloc[train_0], y.iloc[train_0])
cyclic_spline_poly_predictions = cyclic_spline_poly_pipeline.predict(X.iloc[test_0])
# -
# Again we zoom on the last 4 days of the test set:
#
#
last_hours = slice(-96, None)
fig, ax = plt.subplots(figsize=(12, 4))
fig.suptitle("Predictions by non-linear regression models")
ax.plot(
y.iloc[test_0].values[last_hours],
"x-",
alpha=0.2,
label="Actual demand",
color="black",
)
ax.plot(
gbrt_predictions[last_hours],
"x-",
label="Gradient Boosted Trees",
)
ax.plot(
one_hot_poly_predictions[last_hours],
"x-",
label="One-hot + polynomial kernel",
)
ax.plot(
cyclic_spline_poly_predictions[last_hours],
"x-",
label="Splines + polynomial kernel",
)
_ = ax.legend()
# First, note that trees can naturally model non-linear feature interactions
# since, by default, decision trees are allowed to grow beyond a depth of 2
# levels.
#
# Here, we can observe that the combinations of spline features and non-linear
# kernels works quite well and can almost rival the accuracy of the gradient
# boosting regression trees.
#
# On the contrary, one-hot encoded time features do not perform that well with
# the low rank kernel model. In particular, they significantly over-estimate
# the low demand hours more than the competing models.
#
# We also observe that none of the models can successfully predict some of the
# peak rentals at the rush hours during the working days. It is possible that
# access to additional features would be required to further improve the
# accuracy of the predictions. For instance, it could be useful to have access
# to the geographical repartition of the fleet at any point in time or the
# fraction of bikes that are immobilized because they need servicing.
#
# Let us finally get a more quantative look at the prediction errors of those
# three models using the true vs predicted demand scatter plots:
#
#
fig, axes = plt.subplots(ncols=3, figsize=(12, 4), sharey=True)
fig.suptitle("Non-linear regression models")
predictions = [
one_hot_poly_predictions,
cyclic_spline_poly_predictions,
gbrt_predictions,
]
labels = [
"One hot + polynomial kernel",
"Splines + polynomial kernel",
"Gradient Boosted Trees",
]
for ax, pred, label in zip(axes, predictions, labels):
ax.scatter(y.iloc[test_0].values, pred, alpha=0.3, label=label)
ax.plot([0, 1], [0, 1], "--", label="Perfect model")
ax.set(
xlim=(0, 1),
ylim=(0, 1),
xlabel="True demand",
ylabel="Predicted demand",
)
ax.legend()
# This visualization confirms the conclusions we draw on the previous plot.
#
# All models under-estimate the high demand events (working day rush hours),
# but gradient boosting a bit less so. The low demand events are well predicted
# on average by gradient boosting while the one-hot polynomial regression
# pipeline seems to systematically over-estimate demand in that regime. Overall
# the predictions of the gradient boosted trees are closer to the diagonal than
# for the kernel models.
#
# ## Concluding remarks
#
# We note that we could have obtained slightly better results for kernel models
# by using more components (higher rank kernel approximation) at the cost of
# longer fit and prediction durations. For large values of `n_components`, the
# performance of the one-hot encoded features would even match the spline
# features.
#
# The `Nystroem` + `RidgeCV` regressor could also have been replaced by
# :class:`~sklearn.neural_network.MLPRegressor` with one or two hidden layers
# and we would have obtained quite similar results.
#
# The dataset we used in this case study is sampled on a hourly basis. However
# cyclic spline-based features could model time-within-day or time-within-week
# very efficiently with finer-grained time resolutions (for instance with
# measurements taken every minute instead of every hours) without introducing
# more features. One-hot encoding time representations would not offer this
# flexibility.
#
# Finally, in this notebook we used `RidgeCV` because it is very efficient from
# a computational point of view. However, it models the target variable as a
# Gaussian random variable with constant variance. For positive regression
# problems, it is likely that using a Poisson or Gamma distribution would make
# more sense. This could be achieved by using
# `GridSearchCV(TweedieRegressor(power=2), param_grid({"alpha": alphas}))`
# instead of `RidgeCV`.
#
#
| testpad/plot_cyclical_feature_engineering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python3
# ---
# + [markdown] render=false
# # Finding Optimal Locations for New Stores
#
# This notebook is an example of how **Decision Optimization** can help to prescribe decisions for a complex constrained problem.
#
# When you finish this notebook, you'll have a foundational knowledge of _Prescriptive Analytics_.
#
# >This notebook requires the Commercial Edition of CPLEX engines, which is included in the Default Python 3.6 XS + DO in Watson Studio.
#
# Table of contents:
#
# - [Describe the business problem](#Describe-the-business-problem)
# * [How decision optimization (prescriptive analytics) can help](#How--decision-optimization-can-help)
# * [Use decision optimization](#Use-decision-optimization)
# * [Step 1: Download the library](#Step-1:-Download-the-library)
# * [Step 2: Set up the engines](#Step-2:-Set-up-the-prescriptive-engine)
# - [Step 3: Model the data](#Step-3:-Model-the-data)
# * [Step 4: Prepare the data](#Step-4:-Prepare-the-data)
# - [Step 5: Set up the prescriptive model](#Step-5:-Set-up-the-prescriptive-model)
# * [Define the decision variables](#Define-the-decision-variables)
# * [Express the business constraints](#Express-the-business-constraints)
# * [Express the objective](#Express-the-objective)
# * [Solve with the Decision Optimization solve service](#Solve-with-the-Decision-Optimization-solve-service)
# * [Step 6: Investigate the solution and run an example analysis](#Step-6:-Investigate-the-solution-and-then-run-an-example-analysis)
# * [Summary](#Summary)
#
# ****
# + [markdown] render=false
# ## Describe the business problem
#
# * A fictional Coffee Company plans to open N shops in the near future and needs to determine where they should be located knowing the following:
# * Most of the customers of this coffee brewer enjoy reading and borrowing books, so the goal is to locate those shops in such a way that all the city public libraries are within minimal walking distance.
# * We use <a href="https://data.cityofchicago.org" target="_blank" rel="noopener noreferrer">Chicago open data</a> for this example.
# * We implement a <a href="https://en.wikipedia.org/wiki/K-medians_clustering" target="_blank" rel="noopener noreferrer">K-Median model</a> to get the optimal location of our future shops.
# + [markdown] render=false
# ## How decision optimization can help
#
# * Prescriptive analytics (decision optimization) technology recommends actions that are based on desired outcomes. It takes into account specific scenarios, resources, and knowledge of past and current events. With this insight, your organization can make better decisions and have greater control of business outcomes.
#
# * Prescriptive analytics is the next step on the path to insight-based actions. It creates value through synergy with predictive analytics, which analyzes data to predict future outcomes.
#
# * Prescriptive analytics takes that insight to the next level by suggesting the optimal way to handle that future situation. Organizations that can act fast in dynamic conditions and make superior decisions in uncertain environments gain a strong competitive advantage.
# <br/>
#
# With prescriptive analytics, you can:
#
# * Automate the complex decisions and trade-offs to better manage your limited resources.
# * Take advantage of a future opportunity or mitigate a future risk.
# * Proactively update recommendations based on changing events.
# * Meet operational goals, increase customer loyalty, prevent threats and fraud, and optimize business processes.
# -
# ## Use decision optimization
# ### Step 1: Import the docplex package
#
# This package is presintalled on Watson Studio.
# +
import sys
import docplex.mp
# -
# <i>Note that the more global package docplex contains another subpackage docplex.cp that is dedicated to Constraint Programming, another branch of optimization.</i>
# + [markdown] render=false
# ### Step 2: Model the data
#
# - The data for this problem is quite simple: it is composed of the list of public libraries and their geographical locations.
# - The data is acquired from <a href="https://data.cityofchicago.org" target="_blank" rel="noopener noreferrer">Chicago open data</a> as a JSON file, which is in the following format:
# <code>
# data" : [ [ 1, "13BFA4C7-78CE-4D83-B53D-B57C60B701CF", 1, 1441918880, "885709", 1441918880, "885709", null, "Albany Park", "M, W: 10AM-6PM; TU, TH: 12PM-8PM; F, SA: 9AM-5PM; SU: Closed", "Yes", "Yes ", "3401 W. Foster Avenue", "CHICAGO", "IL", "60625", "(773) 539-5450", [ "http://www.chipublib.org/locations/1/", null ], [ null, "41.975456", "-87.71409", null, false ] ]
# </code>
# This code snippet represents library "**3401 W. Foster Avenue**" located at **41.975456, -87.71409**
#
# + [markdown] render=false
# ### Step 3: Prepare the data
# We need to collect the list of public libraries locations and keep their names, latitudes, and longitudes.
# +
# Store longitude, latitude and street crossing name of each public library location.
class XPoint(object):
def __init__(self, x, y):
self.x = x
self.y = y
def __str__(self):
return "P({:g}_{:g})".format(self.x, self.y)
class NamedPoint(XPoint):
def __init__(self, name, x, y):
XPoint.__init__(self, x, y)
self.name = name
def __str__(self):
return self.name
# + [markdown] render=false
# #### Define how to compute the earth distance between 2 points
# To easily compute distance between 2 points, we use the Python package <a href="https://pypi.python.org/pypi/geopy" target="_blank" rel="noopener noreferrer">geopy</a>
# -
try:
import geopy.distance
except:
if hasattr(sys, "real_prefix"):
# we are in a virtual env.
# !pip install geopy
else:
# !pip install --user geopy
# +
# Simple distance computation between 2 locations.
from geopy.distance import great_circle
def get_distance(p1, p2):
return great_circle((p1.y, p1.x), (p2.y, p2.x)).miles
# + [markdown] render=false
# #### Declare the list of libraries
# Parse the JSON file to get the list of libraries and store them as Python elements.
# + render=false
def build_libraries_from_url(url, name_pos, lat_long_pos):
import json
import requests
r = requests.get(url)
myjson = json.loads(r.text, parse_constant="utf-8")
myjson = myjson["data"]
libraries = []
k = 1
for location in myjson:
uname = location[name_pos]
try:
latitude = float(location[lat_long_pos][1])
longitude = float(location[lat_long_pos][2])
except TypeError:
latitude = longitude = None
try:
name = str(uname)
except:
name = "???"
name = "P_%s_%d" % (name, k)
if latitude and longitude:
cp = NamedPoint(name, longitude, latitude)
libraries.append(cp)
k += 1
return libraries
# -
libraries = build_libraries_from_url(
"https://data.cityofchicago.org/api/views/x8fc-8rcq/rows.json?accessType=DOWNLOAD",
name_pos=10,
lat_long_pos=16,
)
# + render=false
print("There are %d public libraries in Chicago" % (len(libraries)))
# -
# #### Define number of shops to open
# Create a constant that indicates how many coffee shops we would like to open.
# + render=false
nb_shops = 5
print("We would like to open %d coffee shops" % nb_shops)
# + [markdown] render=false
# #### Validate the data by displaying them
# We will use the <a href="https://folium.readthedocs.org/en/latest/quickstart.html#getting-started" target="_blank" rel="noopener noreferrer">folium</a> library to display a map with markers.
# -
try:
import folium
except:
if hasattr(sys, "real_prefix"):
# we are in a virtual env.
# !pip install folium
else:
# !pip install folium
# +
import folium
map_osm = folium.Map(location=[41.878, -87.629], zoom_start=11)
for library in libraries:
lt = library.y
lg = library.x
folium.Marker([lt, lg]).add_to(map_osm)
map_osm
# -
# After running the above code, the data is displayed but it is impossible to determine where to ideally open the coffee shops by just looking at the map.
#
# Let's set up DOcplex to write and solve an optimization model that will help us determine where to locate the coffee shops in an optimal way.
# ### Step 4: Set up the prescriptive model
# +
from docplex.mp.environment import Environment
env = Environment()
env.print_information()
# -
# #### Create the DOcplex model
# The model contains all the business constraints and defines the objective.
# +
from docplex.mp.model import Model
mdl = Model("coffee shops")
# -
# #### Define the decision variables
# +
BIGNUM = 999999999
# Ensure unique points
libraries = set(libraries)
# For simplicity, let's consider that coffee shops candidate locations are the same as libraries locations.
# That is: any library location can also be selected as a coffee shop.
coffeeshop_locations = libraries
# Decision vars
# Binary vars indicating which coffee shop locations will be actually selected
coffeeshop_vars = mdl.binary_var_dict(
coffeeshop_locations, name="is_coffeeshop"
)
#
# Binary vars representing the "assigned" libraries for each coffee shop
link_vars = mdl.binary_var_matrix(coffeeshop_locations, libraries, "link")
# -
# #### Express the business constraints
# First constraint: if the distance is suspect, it needs to be excluded from the problem.
for c_loc in coffeeshop_locations:
for b in libraries:
if get_distance(c_loc, b) >= BIGNUM:
mdl.add_constraint(
link_vars[c_loc, b] == 0,
"ct_forbid_{!s}_{!s}".format(c_loc, b),
)
# Second constraint: each library must be linked to a coffee shop that is open.
mdl.add_constraints(
link_vars[c_loc, b] <= coffeeshop_vars[c_loc]
for b in libraries
for c_loc in coffeeshop_locations
)
mdl.print_information()
# Third constraint: each library is linked to exactly one coffee shop.
mdl.add_constraints(
mdl.sum(link_vars[c_loc, b] for c_loc in coffeeshop_locations) == 1
for b in libraries
)
mdl.print_information()
# Fourth constraint: there is a fixed number of coffee shops to open.
# +
# Total nb of open coffee shops
mdl.add_constraint(
mdl.sum(coffeeshop_vars[c_loc] for c_loc in coffeeshop_locations)
== nb_shops
)
# Print model information
mdl.print_information()
# -
# #### Express the objective
#
# The objective is to minimize the total distance from libraries to coffee shops so that a book reader always gets to our coffee shop easily.
#
# Minimize total distance from points to hubs
total_distance = mdl.sum(
link_vars[c_loc, b] * get_distance(c_loc, b)
for c_loc in coffeeshop_locations
for b in libraries
)
mdl.minimize(total_distance)
# #### Solve with the Decision Optimization solve service
#
# Solve the model.
# +
print("# coffee shops locations = %d" % len(coffeeshop_locations))
print("# coffee shops = %d" % nb_shops)
assert mdl.solve(), "!!! Solve of the model fails"
# + [markdown] render=false
# ### Step 6: Investigate the solution and then run an example analysis
#
# The solution can be analyzed by displaying the location of the coffee shops on a map.
# + render=false
total_distance = mdl.objective_value
open_coffeeshops = [
c_loc
for c_loc in coffeeshop_locations
if coffeeshop_vars[c_loc].solution_value == 1
]
not_coffeeshops = [
c_loc for c_loc in coffeeshop_locations if c_loc not in open_coffeeshops
]
edges = [
(c_loc, b)
for b in libraries
for c_loc in coffeeshop_locations
if int(link_vars[c_loc, b]) == 1
]
print("Total distance = %g" % total_distance)
print("# coffee shops = {}".format(len(open_coffeeshops)))
for c in open_coffeeshops:
print("new coffee shop: {!s}".format(c))
# -
# #### Displaying the solution
# Coffee shops are highlighted in red.
# +
import folium
map_osm = folium.Map(location=[41.878, -87.629], zoom_start=11)
for coffeeshop in open_coffeeshops:
lt = coffeeshop.y
lg = coffeeshop.x
folium.Marker(
[lt, lg], icon=folium.Icon(color="red", icon="info-sign")
).add_to(map_osm)
for b in libraries:
if b not in open_coffeeshops:
lt = b.y
lg = b.x
folium.Marker([lt, lg]).add_to(map_osm)
for (c, b) in edges:
coordinates = [[c.y, c.x], [b.y, b.x]]
map_osm.add_child(folium.PolyLine(coordinates, color="#FF0000", weight=5))
map_osm
# -
# ## Summary
#
# You have learned how to set up, formulate and solve an optimization model using Decision Optimization in Watson Studio.
# #### References
# * <a href="https://rawgit.com/IBMDecisionOptimization/docplex-doc/master/docs/index.html" target="_blank" rel="noopener noreferrer">Decision Optimization CPLEX Modeling for Python documentation</a>
# * <a href="https://dataplatform.cloud.ibm.com/docs/content/getting-started/welcome-main.html?audience=wdp&context=wdp" target="_blank" rel="noopener noreferrer">Watson Studio documentation</a>
# Copyright © 2017-2019 IBM. This notebook and its source code are released under the terms of the MIT License.
# <div style="background:#F5F7FA; height:110px; padding: 2em; font-size:14px;">
# <span style="font-size:18px;color:#152935;">Love this notebook? </span>
# <span style="font-size:15px;color:#152935;float:right;margin-right:40px;">Don't have an account yet?</span><br>
# <span style="color:#5A6872;">Share it with your colleagues and help them discover the power of Watson Studio!</span>
# <span style="border: 1px solid #3d70b2;padding:8px;float:right;margin-right:40px; color:#3d70b2;"><a href="https://ibm.co/wsnotebooks" target="_blank" style="color: #3d70b2;text-decoration: none;">Sign Up</a></span><br>
# </div>
#
| advanced-machine-learning-and-signal-processing/Examples/Finding optimal locations of new store using Decision Optimization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Pyenv
# - By using Pyenv, we are able to use several versions of Python.
# - Why using the pyenv?
# - Eligible to make a virtual environment in each directories.
# - Efficient to manage the several versions of Python in a system.
# - Installation Tools
# - pyenv - From one host, we are able to use many versions of Python.
# - virtualenv - Helps to use the many versions of Python in the virtual env.
# - autoenv - Enable the virtual environment by each directories.
# ## 1. Pyenv
# - 1.1 apt-get Updates
# - `$ sudo apt-get update`
# - `$ sudo apt-get upgrade`
# - When the Encoding selection's interface appears, then select the UTF-8.
# (It will not be appeared in windows)
# - 1.2
# - `$sudo apt-get install -y make build-essential libssl-dev zlib1g-dev libbz2-dev libreadline-dev libsqlite3-dev wget curl llvm libncurses5-dev libncursesw5-dev xzutils tk-dev`
# - 1.3 git install
# - sudo apt-get install git
# - 1.4 pyenv install
# - `curl -L https://raw.githubusercontent.com/yyuu/pyenv-installer/master/bin/pyenv-installer|bash`
# - 1.5 .bash_profile configuration
# - 1.5.1 Add pyenv configuration in the .bash_profile
# - /home/ubuntu$ vi `.bash_profile`
# - 1.5.2 Pyenv configuration adding in the `.bash_profile`
# - `.bash_profile`
# - `---------------------------------------`
# - `export PATH = '/home/ubuntu/.pyenv/bin:$PATH'`
# - `eval '$(pyenv init-)'`
# - `eval '$(pyenv virtualenv-init-)'`
# - `source .bashrc`
# - Difference between `.bash_profile` and `.bashrc`:
# - `.bash_profile` is a login shell and `.bashrc` is not a login shell
# - `.bash_profile` initiated when the server is started, and `.bashrc` is initiated when a new shell created.
# - 1.5.3 Reflect the changed `.bash_profile`
# - `$ source .bash_profile`
# - 1.6 Install python
# - 1.6.1 Checking python version that are usable
# - `$ pyenv install --list | grep -v '-'`
# - 1.6.2 python Installation
# - `$ export LC_ALL = C`
# - `$ pyenv install 3.6.5`
# - `$ pyenv install 2.7.15`
# - 1.6.3 Checking versions that we have installed in pyenv
# - `$ pyenv versions`
# - 1.6.4 Change the python version by using the pyenv
# - `pyenv shell 3.6.5`
# - 1.6.5 Global version configuration on the pyenv
# - `pyenv global 3.6.5`
# - 1.6.6 python delete by using pyenv
# - `pyenv uninstall 2.7.15`
# ## 2. Virtualenv
# - 2.1 virtualenv installation
# - `$ sudo apt-get install pyenv-virtualenv`
# - `$ sudo apt-get install python-virtualenv`
# - 2.2 Make a Virtual Environment
# - `$ pyenv virtualenv 3.6.5 python3`
# - `$ pyenv virtualenv 2.7.15 python2`
# - `$ pyenv virtualenv 3.6.5 python3_2`
# - 2.3 Virtual Environment Initiation
# - `$ pyenv shell python3`
# - 2.4 Virtual Environment uninstallation
# - `$ pyenv uninstall python3_2`
# ## 3. Autoenv
# - 3.1 autoenv Installation and Configuration
# - `$ git clone git://github.com/kennethreitz/autoenv.git ~/.autoenv`
# - `$ echo 'source ~/.autoenv/activate.sh' >> ~/.bash_profile`
# - `$ source .bash_profile`
# - 3.2 python virtual environment change
# - By using `activate`, only environment can be changed.
# - `$ pyenv activate python2`
# - 3.3 pyenv global environment
# - `$ pyenv deactivate`
# - 3.4 Python Virtual Environment Adjust in each directories
# - Make directories for python2 and python3 and configure as below
# - `python2 $vi. env`
# - `pyenv activate python2`
# - `python3 $vi. env`
# - `pyenv activate python3`
# - `~/.env`
# - `pyenv deactivate`
| 2018_07_09_Pyenv.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="6PQVD0cNvPDE" executionInfo={"status": "ok", "timestamp": 1650555979483, "user_tz": -120, "elapsed": 2232, "user": {"displayName": "<NAME>", "userId": "16575933074778738670"}} outputId="869933c1-0f76-411f-d59d-dec4a0bf2f47"
# # !pip install import-ipynb
#access drive
from google.colab import drive
drive.mount('/content/drive')
# %cd /content/drive/MyDrive/GRAD-C24_Machine_Learning/MLProject_KenyaFinancial
x_path= "/content/drive/MyDrive/GRAD-C24_Machine_Learning/MLProject_KenyaFinancial/clean_data/ohc3.csv"
y_path= "/content/drive/MyDrive/GRAD-C24_Machine_Learning/MLProject_KenyaFinancial/clean_data/depVar"
# + colab={"base_uri": "https://localhost:8080/"} id="6M5hJcfZyRPA" executionInfo={"status": "ok", "timestamp": 1650555985594, "user_tz": -120, "elapsed": 4141, "user": {"displayName": "<NAME>", "userId": "16575933074778738670"}} outputId="300132f1-5f5c-42ff-85e9-b3793dbfab53"
# %run Splitting_data.ipynb
# + id="n6swBJNayRvZ"
X_train, X_val, y_train, y_val = training_set(x_path, y_path)
# + id="-uszwQNqyHPg" colab={"base_uri": "https://localhost:8080/", "height": 415} executionInfo={"status": "error", "timestamp": 1650557263723, "user_tz": -120, "elapsed": 311, "user": {"displayName": "<NAME>", "userId": "16575933074778738670"}} outputId="65d564ed-1703-49eb-d0d6-d773f375145d"
import xgboost
from matplotlib import pyplot as plt
#le = preprocessing.LabelEncoder()
#y_vald = le.fit_transform(y_vald.astype(str))
y_val = y_val.convert_dtypes(convert_string=True)
#eval_set=[(X_train, y_train), (X_val, y_val)]
eval_set=[(X_val, y_val)]
xgb_reg = xgboost.XGBClassifier(n_estimators=3000,
max_depth=12,
subsample=1,
max_bin=500, # Maximum number of discrete bins to bucket continuous features. Increasing this number improves the optimality of splits at the cost of higher computation time.
#objective="reg:squarederror", # for iterative learning
eval_metric="accuracy", # for early-stopping evaluation using eval_set
early_stopping_rounds=10,
max_delta_step=0, # Maximum delta step we allow each leaf output to be. If the value is set to 0, it means there is no constraint. If it is set to a positive value, it can help making the update step more conservative. Usually this parameter is not needed, but it might help in logistic regression when class is extremely imbalanced. Set it to value of 1-10 might help control the update.
verbosity=1,
n_jobs=-1,
)
# see more parameters in https://xgboost.readthedocs.io/en/stable/parameter.html#parameters-for-tree-booster
xgb_reg.fit(X_train, y_train,
eval_set=eval_set,
#verbose=False
)
results = xgb_reg.evals_result()
lowest_score = round(min(results["validation_1"]["rmse"]), 2)
print(f"Optimal number of trees: {xgb_reg.best_ntree_limit}\n RMSE: {lowest_score}")
# plot graph
plt.figure(figsize=(10,7))
plt.plot(results["validation_0"]["rmse"], label="Training loss")
plt.plot(results["validation_1"]["rmse"], label="Validation loss")
plt.axvline(xgb_reg.best_ntree_limit, color="gray", label="Optimal tree number")
plt.xlabel("Number of trees")
plt.ylabel("Loss")
plt.legend()
# + colab={"base_uri": "https://localhost:8080/", "height": 505} id="3lpCFcLF1hWu" executionInfo={"status": "ok", "timestamp": 1650557279876, "user_tz": -120, "elapsed": 219, "user": {"displayName": "<NAME>", "userId": "16575933074778738670"}} outputId="fcc5fb2f-69e5-4591-faef-2ff3fed2e6c5"
X_val
# + [markdown] id="ftwfajgo0NPC"
# Important parameters:
#
#
# * early_stopping: tells the model after how many rounds to stop after it finds the smallest number of the evaluation metric
# * eval_metric: evaluation metric must be specified
#
#
# + id="CrYw5neyxJSM"
| XGBoost.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # *When* should you create an object?
# ## Complicated Function Signatures (continued)
#
# Here is my solution to the previous problem.
class Environment:
def __init__(self, times, values):
self.__times = times
self.__values = values
def __integrate_trapezoid(self, v):
x = self.__times[v]
y = self.__values[v]
return sum(0.5*(x[i] - x[i-1])*(y[i] + y[i-1]) for i in range(1, len(x)))
def get_times(self, v):
return self.__times[v]
def get_values(self, v):
return self.__values[v]
def compute_v1(self):
return self.__integrate_trapezoid('A') - self.__integrate_trapezoid('B')
def compute_v2(self):
all_v = ( self.get_values('A') + self.get_values('B') +
self.get_values('C') + self.get_values('D') )
return sum(all_v) / len(all_v)
def compute_v3(self):
a_b = self.__integrate_trapezoid('A') - self.__integrate_trapezoid('B')
c_d = self.__integrate_trapezoid('C') + self.__integrate_trapezoid('D')
return a_b / c_d
def compute_v4(self):
return sorted(self.get_times('A') + self.get_times('B') +
self.get_times('C') + self.get_times('D'))
# And then I modified the `read_instrument_data` function to return an `Environment` instance.
def read_instrument_data(filename):
values = {}
times = {}
with open(filename) as f:
for record in f:
var,time,value = record.split(',')
if var not in values:
values[var] = []
times[var] = []
values[var].append(float(value))
times[var].append(float(time))
return Environment(times, values)
# Now, we should be able to test this new class out to see if it gives the same results as we got previously.
env = read_instrument_data('instrument2.csv')
env.get_times('A')
env.get_values('A')
env.compute_v1()
env.compute_v2()
env.compute_v3()
env.compute_v4()
# Looks correct!
# ## Key Takeaway
#
# Sometimes the data itself doesn't have to be complicated; it can just be numerous! And when the data describing different aspects of the same thing grows like that, the complexity can show up in both the number of variables used to describe the different aspects of the same thing grows, the signatures of the functions operating on that data can grow and get complex, too! You can reduce a lot of burden on the user (and yourself) if you bundle all of that data and those functions together into a object.
# | | | |
# | :- | -- | -: |
# | [[Home]](../index.ipynb) | <img width="100%" height="1" src="../images/empty.png"/> | [« Previous](09.ipynb) \| [Next »](11.ipynb) |
| notebooks/10.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#Data Preprocessing
#importing the libraries
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
#Importing the dataset
dataset = pd.read_csv('Data.csv')
X = dataset.iloc[:, :-1].values
Y = dataset.iloc[:, 3].values
#Taking care of missing data
from sklearn.preprocessing import Imputer
imputer = Imputer(missing_values = 'NaN', strategy = 'mean', axis = 0)
imputer = imputer.fit(X[:, 1:3]) #Fitting imputer with columns with missing data, 3 is upper bound and not included
X[:, 1:3] = imputer.transform(X[:, 1:3])
#Encoding categorical data
from sklearn.preprocessing import LabelEncoder, OneHotEncoder
labelencoder_X = LabelEncoder()
X[:, 0] = labelencoder_X.fit_transform(X[:, 0])
onehotencoder = OneHotEncoder(categorical_features = [0])
X = onehotencoder.fit_transform(X).toarray()
labelencoder_Y = LabelEncoder()
Y = labelencoder_Y.fit_transform(Y)
#Splitting the dataset into Training set and Test set
from sklearn.cross_validation import train_test_split
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size = 0.2, random_state = 0)
#Feature scaling
from sklearn.preprocessing import StandardScaler
sc_X = StandardScaler()
X_train = sc_X.fit_transform(X_train)
X_test = sc_X.transform(X_test)
print(X_train)
print(X)
print(Y)
| Topics/2. Data preprocessing/Data preprocessing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from pprint import pprint
import sys
sys.path.append('../')
from soynlp.tokenizer import RegexTokenizer, LTokenizer, MaxScoreTokenizer
# -
# ### MaxScoreTokenizer
# 아래 문장을 단어들로 직접 나눠보세요
#
# 이런문장을직접토크나이징을해볼게요
#
# 우리는 지금 토크나이징을 이야기 하고 있기 때문에, '토크나이징'이라는 단어가 눈에 잘 들어온다. 그 다음으로는 '문장', '직접', '볼게요' 순으로 단어가 눈에 들어온다 (고 가정하자). 그렇다면 아래 순서대로 단어에 마킹을 할 수 있다.
#
# 이런문장을직접[토크나이징]을해볼게요
# 이런[문장]을직접[토크나이징]을해볼게요
# 이런[문장]을[직접][토크나이징]을해볼게요
# 이런[문장]을[직접][토크나이징]을해[볼게요]
#
# 단어로 인식되지 않은 부분들은 그대로 이어서 하나의 단어로 취급하자.
#
# [이런][문장][을][직접][토크나이징][을해][볼게요]
#
# 그 결과 [이런, 문장, 을, 직접, 토크나이징, 을해, 볼게요]라는 토큰을 얻게 된다. 아쉬운 점은 [토크나이징, 을, 해, 볼게요]이지만, -을, 해-를 제대로 인식하지 못하는 문제는 있다. 이는 나중에 다른 방식의 토크나이저에서 해결하자. 더 중요한 점은, 단어라고 확신이 드는 부분부터 연속된 글자집합에서 잘라내어도 토크나이징이 된다는 점이다.
#
# 단어 추출 기법을 통하여 subword 들의 단어 가능 점수를 계산할 수 있다. 우리에게 아래의 네 가지 subwords의 점수표와 예문이 있다고 하자.
#
# 파스타가좋아요
#
# scores = {'파스': 0.3, '파스타': 0.7, '좋아요': 0.2, '좋아':0.5}
#
# 단어 길이의 범위를 [2, 3]이라고 가정하면 아래와 같은 subword score를 얻을 수 있다. 아래는 (subword, begin, end, score)이다.
#
# [('파스', 0, 2, 0.3),
# ('파스타', 0, 3, 0.7),
# ('스타', 1, 3, 0),
# ('스타가', 1, 4, 0),
# ('타가', 2, 4, 0),
# ('타가좋', 2, 5, 0),
# ('가좋', 3, 5, 0),
# ('가좋아', 3, 6, 0),
# ('좋아', 4, 6, 0.5),
# ('좋아요', 4, 7, 0.2),
# ('아요', 5, 7, 0)]
#
# 이를 점수 순서로 정렬하면 아래와 같다.
#
# [('파스타', 0, 3, 0.7),
# ('좋아', 4, 6, 0.5),
# ('파스', 0, 2, 0.3),
# ('좋아요', 4, 7, 0.2),
# ('스타', 1, 3, 0),
# ('스타가', 1, 4, 0),
# ('타가', 2, 4, 0),
# ('타가좋', 2, 5, 0),
# ('가좋', 3, 5, 0),
# ('가좋아', 3, 6, 0),
# ('아요', 5, 7, 0)]
#
# 파스타라는 subword의 점수가 가장 높으니, 이를 토큰으로 취급하고, 파스타의 범위인 [0, 3)과 겹치는 다른 subwords을 리스트에서 지워주면 아래와 같은 토큰 후보들이 남는다.
#
# 파스타가좋아요 > [파스타]가좋아요
#
# [('좋아', 4, 6, 0.5),
# ('좋아요', 4, 7, 0.2),
# ('가좋', 3, 5, 0),
# ('가좋아', 3, 6, 0),
# ('아요', 5, 7, 0)]
#
# 다음으로 '좋아'를 단어로 인식하면 남은 토큰 후보가 없기 때문에 아래처럼 토크나이징이 되며, 남는 글자들 역시 토큰으로 취급하면 토크나이징이 끝난다.
#
# 파스타가좋아요 > [파스타]가[좋아]요 > [파스타, 가, 좋아, 요]
#
# 단어 점수만을 이용하여도 손쉽게 토크나이징을 할 수 있다. 이 방법의 장점은 각 도메인에 적절한 단어 점수를 손쉽게 변형할 수 있다는 점이다. 도메인에서 반드시 단어로 취급되어야 하는 글자들이 있다면, 그들의 점수를 scores에 최대값으로 입력하면 된다. Score tie-break는 글자가 오버랩이 되어 있다면, 좀 더 긴 글자를 선택하자. 그럼 합성명사 역시 처리하기 쉽다.
#
# scores = {'서울': 1.0, '대학교': 1.0, '서울대학교': 1.0}
#
# 위처럼 단어 점수가 부여된다면 '서울대학교'를 [서울, 대학교]로 분리하지는 않을 것이다. 대신 '서울'이나 '대학교'가 등장한 다른 어절에서는 이를 단어로 분리한다.
#
# MaxScoreTokenizer는 이러한 컨셉으로, 단어 점수를 토크나이저에 입력하여 원하는 단어를 잘라낸다. 이는 띄어쓰기가 제대로 이뤄지지 않은 텍스트를 토크나이징하기 위한 방법이며, 단어 점수를 잘 정의하는 것은 단어 추출의 몫이다. MaxTokenizer의 사용법은 아래와 같다. MaxScoreTokenizer를 생성할 때, scores에 {str:float} 형태의 단어 점수 사전을 입력한다.
# +
scores = {'파스': 0.3, '파스타': 0.7, '좋아요': 0.2, '좋아':0.5}
tokenizer = MaxScoreTokenizer(scores=scores)
tokenizer.tokenize('파스타가좋아요')
# -
# 띄어쓰기가 포함되어 있는 문장은 띄어쓰기를 기준으로 토큰을 나눈 뒤, 그 안에서 토크나이저가 작동한다. flatten을 하지 않으면 list of list 형식으로 토크나이징 결과가 출력된다. list of list에는 (subword, begin, end, score, length)가 저장되어 있다. Default value는 flatten=True이다.
# +
print('flatten=False')
pprint(tokenizer.tokenize('난파스타가 좋아요', flatten=False))
print('\nflatten=True')
pprint(tokenizer.tokenize('난파스타가 좋아요'))
# -
# ## LTokenizer
# 띄어쓰기가 잘 되어 있는 한국어 문서의 경우에는 MaxScoreTokenizer를 이용할 필요가 없다. 한국어는 L+[R] 구조이기 때문이다 (단어 추출 튜토리얼 참고). 이 때에는 한 어절의 왼쪽에서부터 글자 점수가 가장 높은 부분을 기준으로 토크나이징을 한다.
# +
scores = {'데이':0.5, '데이터':0.5, '데이터마이닝':0.5, '공부':0.5, '공부중':0.45}
tokenizer = LTokenizer(scores=scores)
print('\nflatten=True\nsent = 데이터마이닝을 공부한다')
print(tokenizer.tokenize('데이터마이닝을 공부한다'))
print('\nflatten=False\nsent = 데이터마이닝을 공부한다')
print(tokenizer.tokenize('데이터마이닝을 공부한다', flatten=False))
print('\nflatten=False\nsent = 데이터분석을 위해서 데이터마이닝을 공부한다')
print(tokenizer.tokenize('데이터분석을 위해서 데이터마이닝을 공부한다', flatten=False))
print('\nflatten=True\nsent = 데이터분석을 위해서 데이터마이닝을 공부한다')
print(tokenizer.tokenize('데이터분석을 위해서 데이터마이닝을 공부한다'))
# -
# Tolerance는 한 어절에서 subword 들의 점수의 차이가 그 어절의 점수 최대값과 tolerance 이하로 난다면, 길이가 가장 긴 어절을 선택한다. CohesionProbability에서는 합성명사들은 각각의 요소들보다 낮기 때문에 tolerance를 이용할 수 있다.
# +
print('tolerance=0.0\nsent = 데이터마이닝을 공부중이다')
print(tokenizer.tokenize('데이터마이닝을 공부중이다'))
print('\ntolerance=0.1\nsent = 데이터마이닝을 공부중이다')
print(tokenizer.tokenize('데이터마이닝을 공부중이다', tolerance=0.1))
# -
# ## RegexTokenizer
# 단어를 추출하지 않아도 기본적으로 토크나이징이 되어야 하는 부분들이 있습니다. 언어의 종류가 바뀌는 부분이다.
#
# 이것은123이라는숫자
#
# 위의 에제에서 물론 숫자와 한글이 합쳐져서 하나의 단어가 되기도 한다. 6.25전쟁이 '6.25', '전쟁'으로 나뉘어진 다음에, 이를 '6.25 - 전쟁'으로 묶는 건 ngram extraction으로 할 수 있다.
#
# '6.25전쟁'과 같은 경우는 소수이며, 대부분의 경우에는 한글|숫자|영어(라틴)|기호가 바뀌는 지점에서 토크나이징이 되어야 한다. 위의 예제는 적어도 [이것은, 123, 이라는숫자]로 니뉘어져야 한다. 그 다음에 단어 추출에 의하여 [이것, 은, 123, 이라는, 숫자]라고 나뉘어지는 것이 이상적이다.
#
# 또한 한국어에서 자음/모음이 단어 중간에 단어의 경계를 구분해주는 역할을 한다 .우리는 문자 메시지를 주고 받을 때 자음으로 이뤄진 이모티콘들로 띄어쓰기를 대신하기도 한다.
#
# 아이고ㅋㅋ진짜? = [아이고, ㅋㅋ, 진짜, ?]
#
# 'ㅋㅋ' 덕분에 '아이고'와 '진짜'가 구분이 된다. 또한 'ㅠㅠ'는 'ㅋㅋ'와 함께 붙어있으면 서로 다른 이모티콘으로 구분이 될 수 있다.
#
# 아이고ㅋㅋㅜㅜ진짜? = [아이고, ㅋㅋ, ㅜㅜ, 진짜, ?]
#
# 이를 분리하는 손쉬운 방법은 'ㅋㅋ'를 찾아내어 앞/뒤에 빈 칸을 하나씩 추가한다.
#
# str.replace('ㅋㅋ', ' ㅋㅋ ')
#
# str 하나를 replace 하는 것은 쉽습니다만, 모든 연속된 모음, 혹은 연속된 한글이라는 것은 하나의 str이 아니다. 이와 같이 str에서의 어떤 pattern을 찾아내는 것이 regular expression이다.
#
# re.compile('[가-힣]+')
#
# 위 regular expression은 초/중/종성이 완전한 한국어의 시작부터 끝까지라는 의미이다.
#
# re.compile('[ㄱ-ㅎ]+')
#
# 위 regular expression은 ㄱ부터 ㅎ까지 자음의 범위를 나타낸다.
#
# RegexTokenizer는 regular extression을 이용하여 언어가 달라지는 순간들을 띄어쓴다. 영어의 경우에는 움라우트가 들어가는 경우들이 있어서 알파벳 뿐 아니라 라틴까지 포함하였다.
# +
tokenizer = RegexTokenizer()
sents = [
'이렇게연속된문장은잘리지않습니다만',
'숫자123이영어abc에섞여있으면ㅋㅋ잘리겠죠',
'띄어쓰기가 포함되어있으면 이정보는10점!꼭띄워야죠'
]
for sent in sents:
print(' %s\n->%s\n' % (sent, tokenizer.tokenize(sent)))
# -
# RegexTokenizer 역시 flatten=False이면 띄어쓰기 기준으로 토큰을 나눠서 출력한다
tokenizer.tokenize('띄어쓰기가 포함되어있으면 이정보는10점!꼭띄워야죠',flatten=False)
| SoyNLP/tutorials/tokenizer_usage.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # mxnet linear regression
# # 1. 随机生成数据集
# %matplotlib inline
from IPython import display
from matplotlib import pyplot as plt
from mxnet import autograd, nd
import random
# generate random x1,x2 as features
num_inputs = 2 # 2 input: x1, x2
num_examples = 1000 # 1000 points
true_w = [2, -3.4] # 2 weight
true_b = 4.2 # bais
features = nd.random.normal(scale=1, shape=(num_examples, num_inputs))#标准差为1的正态分布
labels = true_w[0] * features[:, 0] + true_w[1] * features[:, 1] + true_b
labels += nd.random.normal(scale=0.01, shape=labels.shape)#标准差为0.01的正态分布
features[0], labels[0]
# features[0] is [x1,x2], label is y = x1*w1+x2*w2+b
# +
# display data
# x: x1, y:y
def use_svg_display():
# 用矢量图显示
display.set_matplotlib_formats('svg')
def set_figsize(figsize=(3.5, 2.5)):
use_svg_display()
# 设置图的尺寸
plt.rcParams['figure.figsize'] = figsize
set_figsize()
plt.scatter(features[:, 1].asnumpy(), labels.asnumpy(), 1)
# -
# # 2. 读取数据
#
# Gluon提供了data包来读取数据
# +
from mxnet.gluon import data as gdata
batch_size = 10
# 将训练数据的特征和标签组合
dataset = gdata.ArrayDataset(features, labels)
# 随机读取小批量
data_iter = gdata.DataLoader(dataset, batch_size, shuffle=True)
# +
# 测试这个函数
batch_size = 10
for X, y in data_iter:
print(X, y)
break
# -
# # 3. 定义模型
# +
from mxnet.gluon import nn
net = nn.Sequential()
net.add(nn.Dense(1))
# -
# # 4. 初始化模型参数
# +
from mxnet import init
net.initialize(init.Normal(sigma=0.01))
# -
# # 5. 定义损失函数
# +
from mxnet.gluon import loss as gloss
loss = gloss.L2Loss() # 平方损失又称L2范数损失
# -
# # 6. 定义优化算法
# +
from mxnet import gluon
trainer = gluon.Trainer(net.collect_params(), 'sgd', {'learning_rate': 0.03})
# -
# # 7. 训练模型
num_epochs = 3
for epoch in range(1, num_epochs + 1):
for X, y in data_iter:
with autograd.record():
l = loss(net(X), y)
l.backward()
trainer.step(batch_size)
l = loss(net(features), labels)
print('epoch %d, loss: %f' % (epoch, l.mean().asnumpy()))
# # 8. 访问模型训练出来的真实值
dense = net[0]
true_w, dense.weight.data()
true_b, dense.bias.data()
# # 9. 思考
# (1) 如果将l = loss(net(X), y)替换成l = loss(net(X), y).mean(),我们需要将trainer.step(batch_size)相应地改成trainer.step(1)。这是为什么呢?
#
# (2) 查阅MXNet文档,看看gluon.loss和init模块里提供了哪些损失函数和初始化方法。
# +
from mxnet.gluon import loss as gloss
dir(gloss)
# -
# (3)如何访问dense.weight的梯度?
dense.weight.grad
| mxnet/linear_regression_gluon.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + Collapsed="false"
#General Imports
import torch
import torch.nn as nn
import torch.optim as optim
import numpy as np
from torch.utils.data import DataLoader
import random
import matplotlib.pyplot as plt
from tensorboardX import SummaryWriter
#Load fake, non handwritten generator
from fake_texts.pytorch_dataset_fake_2 import Dataset
#Import the loss from baidu
from torch.nn import CTCLoss
#Import the model
from fully_conv_model import cnn_attention_ocr
#Helper to count params
def count_parameters(model):
return sum(p.numel() for p in model.parameters() if p.requires_grad)
#Evaluation function preds_to_integer
from evaluation import wer_eval,preds_to_integer,show,my_collate,AverageMeter
# + Collapsed="false"
#Set up Tensorboard writer for current test
writer = SummaryWriter(log_dir="/home/leander/AI/repos/OCR-CNN/logs2/correct_cosine_2")
# + Collapsed="false"
###Set up model.
cnn=cnn_attention_ocr(model_dim=64,nclasses=67,n_layers=8)
cnn=cnn.cuda().train()
count_parameters(cnn)
# + Collapsed="false" jupyter={"outputs_hidden": true}
#cnn=cnn.eval()
# + Collapsed="false" jupyter={"outputs_hidden": true}
#CTC Loss ,average_frames=True
ctc_loss = CTCLoss(reduction="mean")
#Optimizer: Good initial is 5e5
optimizer = optim.Adam(cnn.parameters(), lr=5e-4)
# + Collapsed="false" jupyter={"outputs_hidden": true}
#We keep track of the Average loss and CER
ave_total_loss = AverageMeter()
CER_total= AverageMeter()
# + Collapsed="false" jupyter={"outputs_hidden": true}
n_iter=0
batch_size=4
# + Collapsed="false" jupyter={"outputs_hidden": true}
#torch.save(cnn.state_dict(), "400ksteps_augment_new_gen_e15.pt")
# + Collapsed="false" jupyter={"outputs_hidden": true}
torch.save(cnn.state_dict(), "415ksteps_augment_new_gen_e56.pt")
# + Collapsed="false"
#
cnn.load_state_dict(torch.load("415ksteps_augment_new_gen_e56.pt"))
# + Collapsed="false" jupyter={"outputs_hidden": true}
ds=Dataset()
# + Collapsed="false" jupyter={"outputs_hidden": true}
trainset = DataLoader(dataset=ds,
batch_size=batch_size,
shuffle=False,
num_workers=6,
pin_memory=True,
collate_fn=my_collate)
gen = iter(trainset)
# + Collapsed="false" jupyter={"outputs_hidden": true}
from torch.optim.lr_scheduler import CosineAnnealingLR
# + Collapsed="false" jupyter={"outputs_hidden": true}
cs=CosineAnnealingLR(optimizer=optimizer,T_max=250000,eta_min=1e-6)
# + Collapsed="false"
npa=1
for epochs in range(10000):
gen = iter(trainset)
print("train start")
for i,ge in enumerate(gen):
#to avoid OOM
if ge[0].shape[3]<=800:
#DONT FORGET THE ZERO GRAD!!!!
optimizer.zero_grad()
#Get Predictions, permuted for CTC loss
log_probs = cnn(ge[0].cuda()).permute((2,0,1))
#Targets have to be CPU for baidu loss
targets = ge[1]#.cpu()
#Get the Lengths/2 becase this is how much we downsample the width
input_lengths = ge[2]/2
target_lengths = ge[3]
#Get the CTC Loss
loss = ctc_loss(log_probs, targets, input_lengths, target_lengths)
#Then backward and step
loss.backward()
optimizer.step()
#Save Loss in averagemeter and write to tensorboard
ave_total_loss.update(loss.data.item())
writer.add_scalar("total_loss", ave_total_loss.average(), n_iter)
#Here we Calculate the Character error rate
cum_len=np.cumsum(target_lengths)
targets=np.split(ge[1].cpu(),cum_len[:-1])
wer_list=[]
for j in range(log_probs.shape[1]):
wer_list.append(wer_eval(log_probs[:,j,:][0:input_lengths[j],:],targets[j]))
#Here we save an example together with its decoding and truth
#Only if it is positive
if np.average(wer_list)>0.1:
max_elem=np.argmax(wer_list)
max_value=np.max(wer_list)
max_image=ge[0][max_elem].cpu()
max_target=targets[max_elem]
max_target=[ds.decode_dict[x] for x in max_target.tolist()]
max_target="".join(max_target)
ou=preds_to_integer(log_probs[:,max_elem,:])
max_preds=[ds.decode_dict[x] for x in ou]
max_preds="".join(max_preds)
writer.add_text("label",max_target,n_iter)
writer.add_text("pred",max_preds,n_iter)
writer.add_image("img",ge[0][max_elem].detach().cpu().numpy(),n_iter)
#gen.close()
#break
#Might become infinite
if np.average(wer_list)< 10:
CER_total.update(np.average(wer_list))
writer.add_scalar("CER", CER_total.average(), n_iter)
#We save when the new avereage CR is beloew the NPA
if npa>CER_total.average() and CER_total.average()>0 and CER_total.average()<1:
torch.save(cnn.state_dict(), "autosave.pt")
npa=CER_total.average()
n_iter=n_iter+1
cs.step()
lr=optimizer.param_groups[0]["lr"]
writer.add_scalar("lr",lr,n_iter)
# + Collapsed="false"
CER_total.average()
# + Collapsed="false" jupyter={"outputs_hidden": true}
save_checkpoint({
'epoch': epoch + 1,
'arch': args.arch,
'state_dict': model.state_dict(),
'optimizer' : optimizer.state_dict(),
})
# + Collapsed="false" jupyter={"outputs_hidden": true}
cnn.load_state_dict(torch.load("autosave.pt"))
# + Collapsed="false" jupyter={"outputs_hidden": true}
optimizer.load_state_dict(torch.load("autosave_optimizer.pt"))
# + Collapsed="false" jupyter={"outputs_hidden": true}
| old_notebooks/OCR_Training_synthetic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Detector de Celulas
# +
# -
#
#
#
# Cargar el modelo ssd7
# (https://github.com/pierluigiferrari/ssd_keras#how-to-fine-tune-one-of-the-trained-models-on-your-own-dataset)
#
# Training del SSD7 (modelo reducido de SSD). Parámetros en config_7.json y descargar VGG_ILSVRC_16_layers_fc_reduced.h5
#
#
#
# +
from keras.optimizers import Adam, SGD
from keras.callbacks import ModelCheckpoint, LearningRateScheduler, TerminateOnNaN, CSVLogger
from keras import backend as K
from keras.models import load_model
from math import ceil
import numpy as np
from matplotlib import pyplot as plt
import os
import json
import xml.etree.cElementTree as ET
import sys
sys.path += [os.path.abspath('ssd_keras-master')]
from keras_loss_function.keras_ssd_loss import SSDLoss
from keras_layers.keras_layer_AnchorBoxes import AnchorBoxes
from keras_layers.keras_layer_DecodeDetections import DecodeDetections
from keras_layers.keras_layer_DecodeDetectionsFast import DecodeDetectionsFast
from keras_layers.keras_layer_L2Normalization import L2Normalization
from ssd_encoder_decoder.ssd_input_encoder import SSDInputEncoder
from ssd_encoder_decoder.ssd_output_decoder import decode_detections, decode_detections_fast
from data_generator.object_detection_2d_data_generator import DataGenerator
from data_generator.object_detection_2d_geometric_ops import Resize
from data_generator.object_detection_2d_photometric_ops import ConvertTo3Channels
from data_generator.data_augmentation_chain_original_ssd import SSDDataAugmentation
from data_generator.object_detection_2d_misc_utils import apply_inverse_transforms
from eval_utils.average_precision_evaluator import Evaluator
from data_generator.data_augmentation_chain_variable_input_size import DataAugmentationVariableInputSize
from data_generator.data_augmentation_chain_constant_input_size import DataAugmentationConstantInputSize
def makedirs(path):
try:
os.makedirs(path)
except OSError:
if not os.path.isdir(path):
raise
K.tensorflow_backend._get_available_gpus()
def lr_schedule(epoch):
if epoch < 80:
return 0.001
elif epoch < 100:
return 0.0001
else:
return 0.00001
config_path = 'config_7_panel_cell.json'
with open(config_path) as config_buffer:
config = json.loads(config_buffer.read())
###############################
# Parse the annotations
###############################
path_imgs_training = config['train']['train_image_folder']
path_anns_training = config['train']['train_annot_folder']
path_imgs_val = config['test']['test_image_folder']
path_anns_val = config['test']['test_annot_folder']
labels = config['model']['labels']
categories = {}
#categories = {"Razor": 1, "Gun": 2, "Knife": 3, "Shuriken": 4} #la categoría 0 es la background
for i in range(len(labels)): categories[labels[i]] = i+1
print('\nTraining on: \t' + str(categories) + '\n')
####################################
# Parameters
###################################
#%%
img_height = config['model']['input'] # Height of the model input images
img_width = config['model']['input'] # Width of the model input images
img_channels = 3 # Number of color channels of the model input images
mean_color = [123, 117, 104] # The per-channel mean of the images in the dataset. Do not change this value if you're using any of the pre-trained weights.
swap_channels = [2, 1, 0] # The color channel order in the original SSD is BGR, so we'll have the model reverse the color channel order of the input images.
n_classes = len(labels) # Number of positive classes, e.g. 20 for Pascal VOC, 80 for MS COCO
scales_pascal = [0.1, 0.2, 0.37, 0.54, 0.71, 0.88, 1.05] # The anchor box scaling factors used in the original SSD300 for the Pascal VOC datasets
#scales_coco = [0.07, 0.15, 0.33, 0.51, 0.69, 0.87, 1.05] # The anchor box scaling factors used in the original SSD300 for the MS COCO datasets
scales = scales_pascal
aspect_ratios = [[1.0, 2.0, 0.5],
[1.0, 2.0, 0.5, 3.0, 1.0/3.0],
[1.0, 2.0, 0.5, 3.0, 1.0/3.0],
[1.0, 2.0, 0.5, 3.0, 1.0/3.0],
[1.0, 2.0, 0.5],
[1.0, 2.0, 0.5]] # The anchor box aspect ratios used in the original SSD300; the order matters
two_boxes_for_ar1 = True
steps = [8, 16, 32, 64, 100, 300] # The space between two adjacent anchor box center points for each predictor layer.
offsets = [0.5, 0.5, 0.5, 0.5, 0.5, 0.5] # The offsets of the first anchor box center points from the top and left borders of the image as a fraction of the step size for each predictor layer.
clip_boxes = False # Whether or not to clip the anchor boxes to lie entirely within the image boundaries
variances = [0.1, 0.1, 0.2, 0.2] # The variances by which the encoded target coordinates are divided as in the original implementation
normalize_coords = True
K.clear_session() # Clear previous models from memory.
model_path = config['train']['saved_weights_name']
# 3: Instantiate an optimizer and the SSD loss function and compile the model.
# If you want to follow the original Caffe implementation, use the preset SGD
# optimizer, otherwise I'd recommend the commented-out Adam optimizer.
if config['model']['backend'] == 'ssd7':
#weights_path = 'VGG_ILSVRC_16_layers_fc_reduced.h5'
scales = [0.08, 0.16, 0.32, 0.64, 0.96] # An explicit list of anchor box scaling factors. If this is passed, it will override `min_scale` and `max_scale`.
aspect_ratios = [0.5 ,1.0, 2.0] # The list of aspect ratios for the anchor boxes
two_boxes_for_ar1 = True # Whether or not you want to generate two anchor boxes for aspect ratio 1
steps = None # In case you'd like to set the step sizes for the anchor box grids manually; not recommended
offsets = None
if os.path.exists(model_path):
print("\nLoading pretrained weights.\n")
# We need to create an SSDLoss object in order to pass that to the model loader.
ssd_loss = SSDLoss(neg_pos_ratio=3, alpha=1.0)
K.clear_session() # Clear previous models from memory.
model = load_model(model_path, custom_objects={'AnchorBoxes': AnchorBoxes,
'L2Normalization': L2Normalization,
'compute_loss': ssd_loss.compute_loss})
else:
####################################
# Build the Keras model.
###################################
if config['model']['backend'] == 'ssd300':
#weights_path = 'VGG_VOC0712Plus_SSD_300x300_ft_iter_160000.h5'
from models.keras_ssd300 import ssd_300 as ssd
model = ssd_300(image_size=(img_height, img_width, img_channels),
n_classes=n_classes,
mode='training',
l2_regularization=0.0005,
scales=scales,
aspect_ratios_per_layer=aspect_ratios,
two_boxes_for_ar1=two_boxes_for_ar1,
steps=steps,
offsets=offsets,
clip_boxes=clip_boxes,
variances=variances,
normalize_coords=normalize_coords,
subtract_mean=mean_color,
swap_channels=swap_channels)
elif config['model']['backend'] == 'ssd7':
#weights_path = 'VGG_ILSVRC_16_layers_fc_reduced.h5'
from models.keras_ssd7 import build_model as ssd
scales = [0.08, 0.16, 0.32, 0.64, 0.96] # An explicit list of anchor box scaling factors. If this is passed, it will override `min_scale` and `max_scale`.
aspect_ratios = [0.5 ,1.0, 2.0] # The list of aspect ratios for the anchor boxes
two_boxes_for_ar1 = True # Whether or not you want to generate two anchor boxes for aspect ratio 1
steps = None # In case you'd like to set the step sizes for the anchor box grids manually; not recommended
offsets = None
model = ssd(image_size=(img_height, img_width, img_channels),
n_classes=n_classes,
mode='training',
l2_regularization=0.0005,
scales=scales,
aspect_ratios_global=aspect_ratios,
aspect_ratios_per_layer=None,
two_boxes_for_ar1=two_boxes_for_ar1,
steps=steps,
offsets=offsets,
clip_boxes=clip_boxes,
variances=variances,
normalize_coords=normalize_coords,
subtract_mean=None,
divide_by_stddev=None)
else :
print('Wrong Backend')
print('OK create model')
#sgd = SGD(lr=config['train']['learning_rate'], momentum=0.9, decay=0.0, nesterov=False)
# TODO: Set the path to the weights you want to load. only for ssd300 or ssd512
weights_path = '../ssd_keras-master/VGG_ILSVRC_16_layers_fc_reduced.h5'
print("\nLoading pretrained weights VGG.\n")
model.load_weights(weights_path, by_name=True)
# 3: Instantiate an optimizer and the SSD loss function and compile the model.
# If you want to follow the original Caffe implementation, use the preset SGD
# optimizer, otherwise I'd recommend the commented-out Adam optimizer.
#adam = Adam(lr=0.001, beta_1=0.9, beta_2=0.999, epsilon=1e-08, decay=0.0)
#sgd = SGD(lr=0.001, momentum=0.9, decay=0.0, nesterov=False)
optimizer = Adam(lr=config['train']['learning_rate'], beta_1=0.9, beta_2=0.999, epsilon=1e-08, decay=0.0)
ssd_loss = SSDLoss(neg_pos_ratio=3, alpha=1.0)
model.compile(optimizer=optimizer, loss=ssd_loss.compute_loss)
model.summary()
# -
# Instanciar los generadores de datos y entrenamiento del modelo.
#
# *Cambio realizado para leer png y jpg. keras-ssd-master/data_generator/object_detection_2d_data_generator.py función parse_xml
#
# +
#ENTRENAMIENTO DE MODELO
#####################################################################
# Instantiate two `DataGenerator` objects: One for training, one for validation.
######################################################################
# Optional: If you have enough memory, consider loading the images into memory for the reasons explained above.
train_dataset = DataGenerator(load_images_into_memory=False, hdf5_dataset_path=None)
val_dataset = DataGenerator(load_images_into_memory=False, hdf5_dataset_path=None)
# 2: Parse the image and label lists for the training and validation datasets. This can take a while.
# The XML parser needs to now what object class names to look for and in which order to map them to integers.
classes = ['background' ] + labels
train_dataset.parse_xml(images_dirs= [config['train']['train_image_folder']],
image_set_filenames=[config['train']['train_image_set_filename']],
annotations_dirs=[config['train']['train_annot_folder']],
classes=classes,
include_classes='all',
#classes = ['background', 'panel', 'cell'],
#include_classes=classes,
exclude_truncated=False,
exclude_difficult=False,
ret=False)
val_dataset.parse_xml(images_dirs= [config['test']['test_image_folder']],
image_set_filenames=[config['test']['test_image_set_filename']],
annotations_dirs=[config['test']['test_annot_folder']],
classes=classes,
include_classes='all',
#classes = ['background', 'panel', 'cell'],
#include_classes=classes,
exclude_truncated=False,
exclude_difficult=False,
ret=False)
#########################
# 3: Set the batch size.
#########################
batch_size = config['train']['batch_size'] # Change the batch size if you like, or if you run into GPU memory issues.
##########################
# 4: Set the image transformations for pre-processing and data augmentation options.
##########################
# For the training generator:
# For the validation generator:
convert_to_3_channels = ConvertTo3Channels()
resize = Resize(height=img_height, width=img_width)
######################################3
# 5: Instantiate an encoder that can encode ground truth labels into the format needed by the SSD loss function.
#########################################
# The encoder constructor needs the spatial dimensions of the model's predictor layers to create the anchor boxes.
if config['model']['backend'] == 'ssd300':
predictor_sizes = [model.get_layer('conv4_3_norm_mbox_conf').output_shape[1:3],
model.get_layer('fc7_mbox_conf').output_shape[1:3],
model.get_layer('conv6_2_mbox_conf').output_shape[1:3],
model.get_layer('conv7_2_mbox_conf').output_shape[1:3],
model.get_layer('conv8_2_mbox_conf').output_shape[1:3],
model.get_layer('conv9_2_mbox_conf').output_shape[1:3]]
ssd_input_encoder = SSDInputEncoder(img_height=img_height,
img_width=img_width,
n_classes=n_classes,
predictor_sizes=predictor_sizes,
scales=scales,
aspect_ratios_per_layer=aspect_ratios,
two_boxes_for_ar1=two_boxes_for_ar1,
steps=steps,
offsets=offsets,
clip_boxes=clip_boxes,
variances=variances,
matching_type='multi',
pos_iou_threshold=0.5,
neg_iou_limit=0.5,
normalize_coords=normalize_coords)
elif config['model']['backend'] == 'ssd7':
predictor_sizes = [model.get_layer('classes4').output_shape[1:3],
model.get_layer('classes5').output_shape[1:3],
model.get_layer('classes6').output_shape[1:3],
model.get_layer('classes7').output_shape[1:3]]
ssd_input_encoder = SSDInputEncoder(img_height=img_height,
img_width=img_width,
n_classes=n_classes,
predictor_sizes=predictor_sizes,
scales=scales,
aspect_ratios_global=aspect_ratios,
two_boxes_for_ar1=two_boxes_for_ar1,
steps=steps,
offsets=offsets,
clip_boxes=clip_boxes,
variances=variances,
matching_type='multi',
pos_iou_threshold=0.5,
neg_iou_limit=0.3,
normalize_coords=normalize_coords)
data_augmentation_chain = DataAugmentationVariableInputSize(resize_height = img_height,
resize_width = img_width,
random_brightness=(-48, 48, 0.5),
random_contrast=(0.5, 1.8, 0.5),
random_saturation=(0.5, 1.8, 0.5),
random_hue=(18, 0.5),
random_flip=0.5,
n_trials_max=3,
clip_boxes=True,
overlap_criterion='area',
bounds_box_filter=(0.3, 1.0),
bounds_validator=(0.5, 1.0),
n_boxes_min=1,
background=(0,0,0))
#######################
# 6: Create the generator handles that will be passed to Keras' `fit_generator()` function.
#######################
train_generator = train_dataset.generate(batch_size=batch_size,
shuffle=True,
transformations= [data_augmentation_chain],
label_encoder=ssd_input_encoder,
returns={'processed_images',
'encoded_labels'},
keep_images_without_gt=False)
val_generator = val_dataset.generate(batch_size=batch_size,
shuffle=False,
transformations=[convert_to_3_channels,
resize],
label_encoder=ssd_input_encoder,
returns={'processed_images',
'encoded_labels'},
keep_images_without_gt=False)
# Summary instance training
category_train_list = []
for image_label in train_dataset.labels:
category_train_list += [i[0] for i in train_dataset.labels[0]]
summary_category_training = {train_dataset.classes[i]: category_train_list.count(i) for i in list(set(category_train_list))}
for i in summary_category_training.keys():
print(i, ': {:.0f}'.format(summary_category_training[i]))
# Get the number of samples in the training and validations datasets.
train_dataset_size = train_dataset.get_dataset_size()
val_dataset_size = val_dataset.get_dataset_size()
print("Number of images in the training dataset:\t{:>6}".format(train_dataset_size))
print("Number of images in the validation dataset:\t{:>6}".format(val_dataset_size))
##########################
# Define model callbacks.
#########################
# TODO: Set the filepath under which you want to save the model.
model_checkpoint = ModelCheckpoint(filepath= config['train']['saved_weights_name'],
monitor='val_loss',
verbose=1,
save_best_only=True,
save_weights_only=False,
mode='auto',
period=1)
#model_checkpoint.best =
csv_logger = CSVLogger(filename='log.csv',
separator=',',
append=True)
learning_rate_scheduler = LearningRateScheduler(schedule=lr_schedule,
verbose=1)
terminate_on_nan = TerminateOnNaN()
callbacks = [model_checkpoint,
csv_logger,
learning_rate_scheduler,
terminate_on_nan]
batch_images, batch_labels = next(train_generator)
initial_epoch = 0
final_epoch = 100 #config['train']['nb_epochs']
steps_per_epoch = 50
history = model.fit_generator(generator=train_generator,
steps_per_epoch=steps_per_epoch,
epochs=final_epoch,
callbacks=callbacks,
validation_data=val_generator,
validation_steps=ceil(val_dataset_size/batch_size),
initial_epoch=initial_epoch,
verbose = 1 if config['train']['debug'] else 2)
history_path = config['train']['saved_weights_name'].split('.')[0] + '_history'
np.save(history_path, history.history)
# -
# +
#Graficar aprendizaje
history_path =config['train']['saved_weights_name'].split('.')[0] + '_history'
hist_load = np.load(history_path + '.npy',allow_pickle=True).item()
print(hist_load.keys())
# summarize history for loss
plt.plot(hist_load['loss'])
plt.plot(hist_load['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
print(config['train']['saved_weights_name'])
# -
# Evaluación del Modelo
# +
config_path = 'config_7_panel.json'
with open(config_path) as config_buffer:
config = json.loads(config_buffer.read())
model_mode = 'training'
# TODO: Set the path to the `.h5` file of the model to be loaded.
model_path = config['train']['saved_weights_name']
# We need to create an SSDLoss object in order to pass that to the model loader.
ssd_loss = SSDLoss(neg_pos_ratio=3, alpha=1.0)
K.clear_session() # Clear previous models from memory.
model = load_model(model_path, custom_objects={'AnchorBoxes': AnchorBoxes,
'L2Normalization': L2Normalization,
'DecodeDetections': DecodeDetections,
'compute_loss': ssd_loss.compute_loss})
train_dataset = DataGenerator(load_images_into_memory=False, hdf5_dataset_path=None)
val_dataset = DataGenerator(load_images_into_memory=False, hdf5_dataset_path=None)
# 2: Parse the image and label lists for the training and validation datasets. This can take a while.
# The XML parser needs to now what object class names to look for and in which order to map them to integers.
classes = ['background' ] + labels
train_dataset.parse_xml(images_dirs= [config['train']['train_image_folder']],
image_set_filenames=[config['train']['train_image_set_filename']],
annotations_dirs=[config['train']['train_annot_folder']],
classes=classes,
include_classes='all',
#classes = ['background', 'panel', 'cell'],
#include_classes=classes,
exclude_truncated=False,
exclude_difficult=False,
ret=False)
val_dataset.parse_xml(images_dirs= [config['test']['test_image_folder']],
image_set_filenames=[config['test']['test_image_set_filename']],
annotations_dirs=[config['test']['test_annot_folder']],
classes=classes,
include_classes='all',
#classes = ['background', 'panel', 'cell'],
#include_classes=classes,
exclude_truncated=False,
exclude_difficult=False,
ret=False)
#########################
# 3: Set the batch size.
#########################
batch_size = config['train']['batch_size'] # Change the batch size if you like, or if you run into GPU memory issues.
evaluator = Evaluator(model=model,
n_classes=n_classes,
data_generator=val_dataset,
model_mode='training')
results = evaluator(img_height=img_height,
img_width=img_width,
batch_size=4,
data_generator_mode='resize',
round_confidences=False,
matching_iou_threshold=0.5,
border_pixels='include',
sorting_algorithm='quicksort',
average_precision_mode='sample',
num_recall_points=11,
ignore_neutral_boxes=True,
return_precisions=True,
return_recalls=True,
return_average_precisions=True,
verbose=True)
mean_average_precision, average_precisions, precisions, recalls = results
total_instances = []
precisions = []
for i in range(1, len(average_precisions)):
print('{:.0f} instances of class'.format(len(recalls[i])),
classes[i], 'with average precision: {:.4f}'.format(average_precisions[i]))
total_instances.append(len(recalls[i]))
precisions.append(average_precisions[i])
if sum(total_instances) == 0:
print('No test instances found.')
else:
print('mAP using the weighted average of precisions among classes: {:.4f}'.format(sum([a * b for a, b in zip(total_instances, precisions)]) / sum(total_instances)))
print('mAP: {:.4f}'.format(sum(precisions) / sum(x > 0 for x in total_instances)))
for i in range(1, len(average_precisions)):
print("{:<14}{:<6}{}".format(classes[i], 'AP', round(average_precisions[i], 3)))
print()
print("{:<14}{:<6}{}".format('','mAP', round(mean_average_precision, 3)))
# -
# Cargar nuevamente el modelo desde los pesos.
# Predicción
# +
from imageio import imread
from keras.preprocessing import image
import time
config_path = 'config_7_panel.json'
input_path = ['panel_jpg/Mision_1/', 'panel_jpg/Mision_2/']
output_path = 'result_ssd7_panel_cell/'
with open(config_path) as config_buffer:
config = json.loads(config_buffer.read())
makedirs(output_path)
###############################
# Parse the annotations
###############################
score_threshold = 0.8
score_threshold_iou = 0.3
labels = config['model']['labels']
categories = {}
#categories = {"Razor": 1, "Gun": 2, "Knife": 3, "Shuriken": 4} #la categoría 0 es la background
for i in range(len(labels)): categories[labels[i]] = i+1
print('\nTraining on: \t' + str(categories) + '\n')
img_height = config['model']['input'] # Height of the model input images
img_width = config['model']['input'] # Width of the model input images
img_channels = 3 # Number of color channels of the model input images
n_classes = len(labels) # Number of positive classes, e.g. 20 for Pascal VOC, 80 for MS COCO
classes = ['background'] + labels
model_mode = 'training'
# TODO: Set the path to the `.h5` file of the model to be loaded.
model_path = config['train']['saved_weights_name']
# We need to create an SSDLoss object in order to pass that to the model loader.
ssd_loss = SSDLoss(neg_pos_ratio=3, alpha=1.0)
K.clear_session() # Clear previous models from memory.
model = load_model(model_path, custom_objects={'AnchorBoxes': AnchorBoxes,
'L2Normalization': L2Normalization,
'DecodeDetections': DecodeDetections,
'compute_loss': ssd_loss.compute_loss})
# +
image_paths = []
for inp in input_path:
if os.path.isdir(inp):
for inp_file in os.listdir(inp):
image_paths += [inp + inp_file]
else:
image_paths += [inp]
image_paths = [inp_file for inp_file in image_paths if (inp_file[-4:] in ['.jpg', '.png', 'JPEG'])]
times = []
for img_path in image_paths:
orig_images = [] # Store the images here.
input_images = [] # Store resized versions of the images here.
#print(img_path)
# preprocess image for network
orig_images.append(imread(img_path))
img = image.load_img(img_path, target_size=(img_height, img_width))
img = image.img_to_array(img)
input_images.append(img)
input_images = np.array(input_images)
# process image
start = time.time()
y_pred = model.predict(input_images)
y_pred_decoded = decode_detections(y_pred,
confidence_thresh=score_threshold,
iou_threshold=score_threshold_iou,
top_k=200,
normalize_coords=True,
img_height=img_height,
img_width=img_width)
#print("processing time: ", time.time() - start)
times.append(time.time() - start)
# correct for image scale
# visualize detections
# Set the colors for the bounding boxes
colors = plt.cm.brg(np.linspace(0, 1, 21)).tolist()
plt.figure(figsize=(20,12))
plt.imshow(orig_images[0],cmap = 'gray')
current_axis = plt.gca()
#print(y_pred)
for box in y_pred_decoded[0]:
# Transform the predicted bounding boxes for the 300x300 image to the original image dimensions.
xmin = box[2] * orig_images[0].shape[1] / img_width
ymin = box[3] * orig_images[0].shape[0] / img_height
xmax = box[4] * orig_images[0].shape[1] / img_width
ymax = box[5] * orig_images[0].shape[0] / img_height
color = colors[int(box[0])]
label = '{}: {:.2f}'.format(classes[int(box[0])], box[1])
current_axis.add_patch(plt.Rectangle((xmin, ymin), xmax-xmin, ymax-ymin, color=color, fill=False, linewidth=2))
current_axis.text(xmin, ymin, label, size='x-large', color='white', bbox={'facecolor':color, 'alpha':1.0})
#plt.figure(figsize=(15, 15))
#plt.axis('off')
save_path = output_path + img_path.split('/')[-1]
plt.savefig(save_path)
plt.close()
file = open(output_path + 'time.txt','w')
file.write('Tiempo promedio:' + str(np.mean(times)))
file.close()
print('Tiempo Total: {:.3f}'.format(np.sum(times)))
print('Tiempo promedio por imagen: {:.3f}'.format(np.mean(times)))
print('OK')
# +
# Summary instance training
category_train_list = []
for image_label in train_dataset.labels:
category_train_list += [i[0] for i in train_dataset.labels[0]]
summary_category_training = {train_dataset.classes[i]: category_train_list.count(i) for i in list(set(category_train_list))}
for i in summary_category_training.keys():
print(i, ': {:.0f}'.format(summary_category_training[i]))
# +
model.summary()
# -
| .ipynb_checkpoints/Panel_Detector_SSD-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %gui qt
# Required knowledges
# ===================
#
# * ``numpy`` (basic knowledges)
# * ``Qt`` (basic knowledges)
# * ``h5py`` (basic knowlegdes)
# Silx IO API
# ===========
import silx.io
# Open a file
# -----------
# open
obj = silx.io.open("data/test.h5")
# do your stuff here
obj.close()
# Open a file with context manager
# --------------------------------
# or using context manager
with silx.io.open("data/test.h5") as obj:
# do your stuff here
# the close is called for you at the end of the with
pass
# Common properties
# -----------------
# +
obj = silx.io.open("data/test.h5")
obj.name # path from file to the object
obj.parent # hdf5 group containing the object
obj.file # hdf5 file containing the object
# test object type
if silx.io.is_file(obj):
# this is a root file
# path of the file from the file system
obj.filename
if silx.io.is_group(obj):
# this is a group
# BTW a file is a group
pass
if silx.io.is_dataset(obj):
# this is a dataset\
pass
# -
# Node traversal
# --------------
# +
obj = silx.io.open("data/test.h5")
if silx.io.is_group(obj):
# it can contains child
# number of child
len(obj)
# iterator on child names
obj.keys()
# access to a child
child = obj["arrays"]
# access to a child using a path
child = obj["arrays/float_3d"]
# the path can be absolute
child = obj["/arrays/float_3d"]
# -
# Data access
# -----------
# +
h5 = silx.io.open("data/test.h5")
obj = h5["arrays/float_3d"]
if silx.io.is_dataset(obj):
# it contains data
# a dataset provides information to the data
obj.shape # multidimensional shape
obj.size # amount of items
obj.dtype # type of the array
# copy the full data as numpy array
data = obj[...]
# or a part of it (using numpy selector)
data = obj[1:2, fd00:a516:7c1b:17cd:6d81:2137:bd2a:2c5b, 2]
scalar = h5["scalars/int64"]
if silx.io.is_dataset(scalar):
# scalar dataset have an empty shape
assert scalar.shape == ()
# special case to access to the value of a scalar
data = scalar[()]
# -
# Spec file as HDF5
# -----------------
# +
import silx.io
h5like = silx.io.open('data/oleg.dat')
# print available scans
print(h5like['/'].keys())
# print available measurements from the scan 94.1
print(h5like['/94.1/measurement'].keys())
# get data from measurement
time = h5like['/94.1/measurement/Epoch']
bpm = h5like['/94.1/measurement/bpmi']
mca = h5like['/94.1/measurement/mca_0/data']
# -
# EDF file as HDF5
# ----------------
# +
import silx.io.utils
h5like = silx.io.open("data/ID16B_diatomee.edf")
# here is the data as a cube using numpy array
# it's a cube of images * number of frames
data = h5like["/scan_0/instrument/detector_0/data"]
# here is the first image
data[0]
print(data[0].shape)
# groups containing datasets of motors, counters
# and others metadata from the EDF header
motors = h5like["/scan_0/instrument/positioners"]
counters = h5like["/scan_0/measurement"]
others = h5like["/scan_0/instrument/detector_0/others"]
print("motor names", list(motors.keys()))
# reach a motor named 'srot'
# it's a vector of values * number of frames
srot = motors["srot"]
# here is the monitor value at the first frame
srot[0]
print(srot[...])
# -
# Silx HDF5 tree
# ==============
#
# 
#
# * Getting start with the HDF5 tree ([http://pythonhosted.org/silx/modules/gui/hdf5/getting_started.html](http://pythonhosted.org/silx/modules/gui/hdf5/getting_started.html))
# Create the widget
# -----------------
import silx.gui.hdf5
tree = silx.gui.hdf5.Hdf5TreeView()
tree.setVisible(True)
model = tree.findHdf5TreeModel()
# Feed it with an HDF5
# --------------------
h5 = silx.io.open("data/test.h5")
model.insertH5pyObject(h5)
# Feed it with a Spec file
# ------------------------
h5 = silx.io.open("data/oleg.dat")
model.insertH5pyObject(h5)
# Feed it with an EDF file
# ------------------------
h5 = silx.io.open("data/ID16B_diatomee.edf")
model.insertH5pyObject(h5)
# Silx DataViewer
# ===============
#
# 
# Create the widget
# -----------------
import silx.gui.data.DataViewerFrame
dataviewer = silx.gui.data.DataViewerFrame.DataViewerFrame()
dataviewer.setVisible(True)
dataviewer.resize(500, 500)
# Feed it with a numpy array
# --------------------------
import numpy
data = numpy.random.rand(100, 100, 100)
dataviewer.setData(data)
# Feed it with a HDF5 dataset
# ---------------------------
import silx.io
h5like = silx.io.open("data/ID16B_diatomee.h5")
dataset = h5like["/data/0299"]
dataviewer.setData(dataset)
# Exercises
# =========
# The exercise is based on a phase contast data. It will help you to create a custom application to browse data.
# Exercise 1
# ----------
#
# > - Browse an HDF5 file
# > * Use [getting started with HDF5 widgets](http://pythonhosted.org/silx/modules/gui/hdf5/getting_started.html)
# > - Identify path of the data
# > - Access to the data
# +
import silx.io
#
# EXERCISE: Open the file 'data/ID16B_diatomee.h5'
#
h5 = ...
#
# EXERCISE: Display the file into the HDF5 tree
#
from silx.gui import hdf5
tree = hdf5.Hdf5TreeView()
model = tree.findHdf5TreeModel()
...
tree.setVisible(True)
# +
#
# EXERCISE: Access to one frame of the image
#
print(...)
#
# EXERCISE: Display it with the data viwer
#
import silx.gui.data.DataViewerFrame
viewer = silx.gui.data.DataViewerFrame.DataViewerFrame()
...
viewer.setVisible(True)
# -
# Exercise 2
# ----------
#
# > 1. From the HDF5 tree, identify path name for
# > - one data frame
# > - one background
# > - one flatfield
# > 2. Compute flatfield correction
# The computation of corrected images is done using this equation using `data`, `flatfield`, and `background` information.
#
# $$corrected = \frac{data - background}{flatfield - background}$$
# +
def correctedImage(data, background, flatfield):
data = numpy.array(data, dtype=numpy.float32)
flatfield = numpy.array(flatfield, dtype=numpy.float32)
return (data - background) / (flatfield - background)
#
# EXERCISE: Reach one data frame, a background and a flatfield from 'data/ID16B_diatomee.h5'
#
...
#
# EXERCISE: Compute the corrected image
#
...
#
# EXERCISE: Display it with the data viewer
#
...
# -
# Exercise 3
# ----------
#
# > 1. Connect together an HDF5 tree view and a data viewer
# > * Use [getting started with HDF5 widgets](http://pythonhosted.org/silx/modules/gui/hdf5/getting_started.html)
#
# +
from silx.gui import qt
from silx.gui import hdf5
import silx.gui.data.DataViewerFrame
class ViewerEx3(qt.QMainWindow):
def __init__(self, parent=None):
qt.QMainWindow.__init__(self, parent)
widget = self.createCentralWidget()
self.setCentralWidget(widget)
def createCentralWidget(self):
splitter = qt.QSplitter(self)
# the tree
self.tree = silx.gui.hdf5.Hdf5TreeView(self)
# the data viewer
self.viewer = silx.gui.data.DataViewerFrame.DataViewerFrame(self)
splitter.addWidget(self.tree)
splitter.addWidget(self.viewer)
splitter.setStretchFactor(1, 1)
#
# EXERCISE: Connect the callback onTreeActivated (bellow)
# to a mouse event from the tree
#
return splitter
def onTreeActivated(self):
#
# EXERCISE: Reach selected objects from the tree
#
#
# EXERCISE: Provide it to the data viewer
#
pass
def appendFile(self, filename):
model = self.tree.findHdf5TreeModel()
model.insertFile(filename)
print("Load %s" % filename)
# -
viewer = ViewerEx3()
viewer.appendFile('data/ID16B_diatomee.h5')
viewer.setVisible(True)
# Exercise 4
# ----------
# > 1. Use the previous application to display corrected data
class ViewerEx4(ViewerEx3):
def onTreeActivated(self):
selectedObjects = list(self.tree.selectedH5Nodes())
if len(selectedObjects) == 0:
self.viewer.setData("Nothing selected")
elif len(selectedObjects) > 1:
self.viewer.setData("Too much things selected")
else:
obj = selectedObjects[0]
node = obj.h5py_object
if "/data/" in node.name:
# That's a data from the /data group
data = self.computeCorrectedImage(node)
self.viewer.setData(data)
else:
# Other data is displayed in a normal way
self.viewer.setData(obj)
def computeCorrectedImage(self, h5data):
"""
:param h5data: H5py dataset selected from the group /data/
"""
background = self.getBackground(h5data)
flatfield = self.getFlatField(h5data)
raw = numpy.array(h5data, dtype=numpy.float32)
flatfield = numpy.array(flatfield, dtype=numpy.float32)
background = background[...]
return (raw - background) / (flatfield - background)
def getBackground(self, h5data):
"""
:param h5data: H5py dataset selected from the group /data/
"""
#
# EXERCISE: Return the background image from the dataset
#
return None
def getFlatField(self, h5data):
"""
:param h5data: H5py dataset selected from the group /data/
"""
#
# EXERCISE: Return the flatfield image from the dataset
#
# 1) you can return a flatfield by default
# 2) you can return the closest flat field according to the index of the data
# 3) you can return an interpolation of the 2 flatfields according to the index of the data
return None
viewer = ViewerEx4()
viewer.appendFile('data/ID16B_diatomee.h5')
viewer.setVisible(True)
| silx/io/io.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="7BgDn2xkyn2s"
# # Exploring DNN learning with TensorFlow
#
# In this assignment we'll dive a little deeper with a series of hands on exercises to better understand DNN learning with Tensorflow. Remember that if you are taking the class for a certificate we will be asking you questions about the assignment in the test!
#
# We start by setting up the problem for you.
# + id="q3KzJyjv3rnA"
import tensorflow as tf
# Load in fashion MNIST
mnist = tf.keras.datasets.fashion_mnist
(training_images, training_labels), (test_images, test_labels) = mnist.load_data()
# Define the base model
model = tf.keras.models.Sequential([tf.keras.layers.Flatten(input_shape=(28,28)),
tf.keras.layers.Dense(512, activation=tf.nn.relu),
tf.keras.layers.Dense(10, activation=tf.nn.softmax)])
# + [markdown] id="SoFa4XHnyn3G"
# Neural Networks learn the best when the data is scaled / normalized to fall in a constant range. One practitioners often use is the range [0,1]. How might you do this to the training and test images used here?
#
# *A hint: these images are saved in the standard [RGB](https://www.rapidtables.com/web/color/RGB_Color.html) format*
# + id="H6n48MV-yn3H"
training_images = training_images / 255.0 #YOUR CODE HERE#
test_images = test_images / 255.0 #YOUR CODE HERE#
# + [markdown] id="3NNmGUaTyn3H"
# Using these improved images lets compile our model using an adaptive optimizer to learn faster and a categorical loss function to differentiate between the the various classes we are trying to classify. Since this is a very simple dataset we will only train for 5 epochs.
# + id="HwzioQ0kyn3I" colab={"base_uri": "https://localhost:8080/"} outputId="a8d0e3db-912e-4ea0-8d43-d29ce77d947b"
# compile the model
model.compile(optimizer = tf.keras.optimizers.Adam(),
loss = 'sparse_categorical_crossentropy',
metrics=['accuracy'])
# fit the model to the training data
model.fit(training_images, training_labels, epochs=5)
# test the model on the test data
model.evaluate(test_images, test_labels)
# + [markdown] id="-JJMsvSB-1UY"
# Once it's done training -- you should see an accuracy value at the end of the final epoch. It might look something like 0.8658. This tells you that your neural network is about 89% accurate in classifying the training data. I.E., it figured out a pattern match between the image and the labels that worked 89% of the time. But how would it work with unseen data? That's why we have the test images. We can call ```model.evaluate```, and pass in the two sets, and it will report back the loss for each. This should reach about .8747 or thereabouts, showing about 87% accuracy. Not Bad!
# + [markdown] id="rquQqIx4AaGR"
# But what did it actually learn? If we inference on the model using ```model.predict``` we get out the following list of values. **What does it represent?**
#
# *A hint: trying running ```print(test_labels[0])```*
# + id="RyEIki0z_hAD" colab={"base_uri": "https://localhost:8080/"} outputId="fd219994-5671-465c-da99-d21520292cd6"
classifications = model.predict(test_images)
print(classifications[0])
# + [markdown] id="OgQSIfDSOWv6"
# Let's now look at the layers in your model. What happens if you double the number of neurons in the dense layer. What different results do you get for loss, training time etc? Why do you think that's the case?
# + id="GSZSwV5UObQP" colab={"base_uri": "https://localhost:8080/"} outputId="2e26db5f-55c3-4df1-f17e-9eeb4badc51e"
NUMBER_OF_NEURONS = 1024 #YOUR_CODE_HERE#
# define the new model
model = tf.keras.models.Sequential([tf.keras.layers.Flatten(input_shape=(28,28)),
tf.keras.layers.Dense(NUMBER_OF_NEURONS, activation=tf.nn.relu),
tf.keras.layers.Dense(10, activation=tf.nn.softmax)])
# compile fit and evaluate the model again
model.compile(optimizer = tf.keras.optimizers.Adam(),
loss = 'sparse_categorical_crossentropy',
metrics=['accuracy'])
model.fit(training_images, training_labels, epochs=5)
model.evaluate(test_images, test_labels)
# + [markdown] id="-0lF5MuvSuZF"
# Consider the effects of additional layers in the network instead of simply more neurons to the same layer. First update the model to add an additional dense layer into the model between the two existing Dense layers.
# + id="vbXKkStuyn3P"
YOUR_NEW_LAYER = tf.keras.layers.Dense(512,activation=tf.nn.relu) #YOUR_CODE_HERE#
model = tf.keras.models.Sequential([tf.keras.layers.Flatten(input_shape=(28,28)),
tf.keras.layers.Dense(512, activation=tf.nn.relu),
YOUR_NEW_LAYER,
tf.keras.layers.Dense(10, activation=tf.nn.softmax)])
# + [markdown] id="6GJBWXbZyn3Q"
# Lets then compile, fit, and evaluate our model. What happens to the error? How does this compare to the original model and the model with double the number of neurons?
# + id="b1YPa6UhS8Es" colab={"base_uri": "https://localhost:8080/"} outputId="67902897-82ad-4a9d-9b3b-f45004d5cdb1"
# compile fit and evaluate the model again
model.compile(optimizer = tf.keras.optimizers.Adam(),
loss = 'sparse_categorical_crossentropy',
metrics=['accuracy'])
model.fit(training_images, training_labels, epochs=5)
model.evaluate(test_images, test_labels)
# + [markdown] id="HS3vVkOgCDGZ"
# Before you trained, you normalized the data. What would be the impact of removing that? To see it for yourself fill in the following lines of code to get a non-normalized set of data and then re-fit and evaluate the model using this data.
# + id="JDqNAqrpCNg0" colab={"base_uri": "https://localhost:8080/"} outputId="2814357e-1565-4a7f-fa1c-040e571ae2a5"
# get new non-normalized mnist data
training_images_non = training_images * 255 #YOUR_CODE_HERE#
test_images_non = test_images * 255 #YOUR_CODE_HERE#
# re-compile, re-fit and re-evaluate
model = tf.keras.models.Sequential([tf.keras.layers.Flatten(input_shape=(28,28)),
tf.keras.layers.Dense(512, activation=tf.nn.relu),
YOUR_NEW_LAYER,
tf.keras.layers.Dense(10, activation=tf.nn.softmax)])
model.compile(optimizer = tf.keras.optimizers.Adam(),
loss = 'sparse_categorical_crossentropy',
metrics=['accuracy'])
model.fit(training_images_non, training_labels, epochs=5)
model.evaluate(test_images_non, test_labels)
classifications = model.predict(test_images_non)
# + [markdown] id="E7W2PT66ZBHQ"
# Sometimes if you set the training for too many epochs you may find that training stops improving and you wish you could quit early. Good news, you can! TensorFlow has a function called ```Callbacks``` which can check the results from each epoch. Modify this callback function to make sure it exits training early but not before reaching at least the second epoch!
#
# *A hint: logs.get(METRIC_NAME) will return the value of METRIC_NAME at the current step*
# + id="pkaEHHgqZbYv" colab={"base_uri": "https://localhost:8080/"} outputId="cdbb8101-4d63-4141-e762-9d2a67cf8f5c"
# define and instantiate your custom Callback
class myCallback(tf.keras.callbacks.Callback):
def on_epoch_end(self, epoch, logs={}):
if(logs.get("accuracy")>0.86):
self.model.stop_training = True
callbacks = myCallback()
# re-compile, re-fit and re-evaluate
model = tf.keras.models.Sequential([tf.keras.layers.Flatten(input_shape=(28,28)),
tf.keras.layers.Dense(512, activation=tf.nn.relu),
YOUR_NEW_LAYER,
tf.keras.layers.Dense(10, activation=tf.nn.softmax)])
model.compile(optimizer = tf.keras.optimizers.Adam(),
loss = 'sparse_categorical_crossentropy',
metrics=['accuracy'])
model.fit(training_images, training_labels, epochs=5, callbacks=[callbacks])
| ML/TinyML_Assignment_2_2_12_Exercise.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="ZMMZ0Iwu2bFD"
# <a href="https://colab.research.google.com/github/https-deeplearning-ai/tensorflow-3-public/blob/main/Course%202%20-%20Custom%20Training%20loops%2C%20Gradients%20and%20Distributed%20Training/Week%204%20-%20Distribution%20Strategy/C2_W4_Lab_2_multi-GPU-mirrored-strategy.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="0zVKdvgS2bFE"
# # Multi-GPU Mirrored Strategy
#
# In this ungraded lab you'll go on how to set up a Multi-GPU Mirrored Strategy.
#
# **Note:**
# - If you are running this on Coursera you'll see it gives a warning about no presence of GPU devices.
# - If you are running this in Colab make sure you have selected your `runtime` to be `GPU` for it to detect it.
# - In both these cases you'll see there's only 1 device that is available.
# - One device is sufficient for helping you understand the these distribution strategies.
# + id="k0VOdqKP2NEz"
import tensorflow as tf
import numpy as np
import os
# Note that it generally has a minimum of 8 cores, but if your GPU has
# less, you need to set this. In this case one of my GPUs has 4 cores
os.environ["TF_MIN_GPU_MULTIPROCESSOR_COUNT"] = "4"
# If the list of devices is not specified in the
# `tf.distribute.MirroredStrategy` constructor, it will be auto-detected.
# If you have *different* GPUs in your system, you probably have to set up cross_device_ops like this
strategy = tf.distribute.MirroredStrategy(cross_device_ops=tf.distribute.HierarchicalCopyAllReduce())
print ('Number of devices: {}'.format(strategy.num_replicas_in_sync))
# Get the data
fashion_mnist = tf.keras.datasets.fashion_mnist
(train_images, train_labels), (test_images, test_labels) = fashion_mnist.load_data()
# Adding a dimension to the array -> new shape == (28, 28, 1)
# We are doing this because the first layer in our model is a convolutional
# layer and it requires a 4D input (batch_size, height, width, channels).
# batch_size dimension will be added later on.
train_images = train_images[..., None]
test_images = test_images[..., None]
# Normalize the images to [0, 1] range.
train_images = train_images / np.float32(255)
test_images = test_images / np.float32(255)
# Batch the input data
BUFFER_SIZE = len(train_images)
BATCH_SIZE_PER_REPLICA = 64
GLOBAL_BATCH_SIZE = BATCH_SIZE_PER_REPLICA * strategy.num_replicas_in_sync
# Create Datasets from the batches
train_dataset = tf.data.Dataset.from_tensor_slices((train_images, train_labels)).shuffle(BUFFER_SIZE).batch(GLOBAL_BATCH_SIZE)
test_dataset = tf.data.Dataset.from_tensor_slices((test_images, test_labels)).batch(GLOBAL_BATCH_SIZE)
# Create Distributed Datasets from the datasets
train_dist_dataset = strategy.experimental_distribute_dataset(train_dataset)
test_dist_dataset = strategy.experimental_distribute_dataset(test_dataset)
# Create the model architecture
def create_model():
model = tf.keras.Sequential([
tf.keras.layers.Conv2D(32, 3, activation='relu'),
tf.keras.layers.MaxPooling2D(),
tf.keras.layers.Conv2D(64, 3, activation='relu'),
tf.keras.layers.MaxPooling2D(),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(64, activation='relu'),
tf.keras.layers.Dense(10)
])
return model
# Instead of model.compile, we're going to do custom training, so let's do that
# within a strategy scope
with strategy.scope():
# We will use sparse categorical crossentropy as always. But, instead of having the loss function
# manage the map reduce across GPUs for us, we'll do it ourselves with a simple algorithm.
# Remember -- the map reduce is how the losses get aggregated
# Set reduction to `none` so we can do the reduction afterwards and divide byglobal batch size.
loss_object = tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True, reduction=tf.keras.losses.Reduction.NONE)
def compute_loss(labels, predictions):
# Compute Loss uses the loss object to compute the loss
# Notice that per_example_loss will have an entry per GPU
# so in this case there'll be 2 -- i.e. the loss for each replica
per_example_loss = loss_object(labels, predictions)
# You can print it to see it -- you'll get output like this:
# Tensor("sparse_categorical_crossentropy/weighted_loss/Mul:0", shape=(48,), dtype=float32, device=/job:localhost/replica:0/task:0/device:GPU:0)
# Tensor("replica_1/sparse_categorical_crossentropy/weighted_loss/Mul:0", shape=(48,), dtype=float32, device=/job:localhost/replica:0/task:0/device:GPU:1)
# Note in particular that replica_0 isn't named in the weighted_loss -- the first is unnamed, the second is replica_1 etc
print(per_example_loss)
return tf.nn.compute_average_loss(per_example_loss, global_batch_size=GLOBAL_BATCH_SIZE)
# We'll just reduce by getting the average of the losses
test_loss = tf.keras.metrics.Mean(name='test_loss')
# Accuracy on train and test will be SparseCategoricalAccuracy
train_accuracy = tf.keras.metrics.SparseCategoricalAccuracy(name='train_accuracy')
test_accuracy = tf.keras.metrics.SparseCategoricalAccuracy(name='test_accuracy')
# Optimizer will be Adam
optimizer = tf.keras.optimizers.Adam()
# Create the model within the scope
model = create_model()
###########################
# Training Steps Functions
###########################
# `run` replicates the provided computation and runs it
# with the distributed input.
@tf.function
def distributed_train_step(dataset_inputs):
per_replica_losses = strategy.run(train_step, args=(dataset_inputs,))
#tf.print(per_replica_losses.values)
return strategy.reduce(tf.distribute.ReduceOp.SUM, per_replica_losses, axis=None)
def train_step(inputs):
images, labels = inputs
with tf.GradientTape() as tape:
predictions = model(images, training=True)
loss = compute_loss(labels, predictions)
gradients = tape.gradient(loss, model.trainable_variables)
optimizer.apply_gradients(zip(gradients, model.trainable_variables))
train_accuracy.update_state(labels, predictions)
return loss
#######################
# Test Steps Functions
#######################
@tf.function
def distributed_test_step(dataset_inputs):
return strategy.run(test_step, args=(dataset_inputs,))
def test_step(inputs):
images, labels = inputs
predictions = model(images, training=False)
t_loss = loss_object(labels, predictions)
test_loss.update_state(t_loss)
test_accuracy.update_state(labels, predictions)
###############
# TRAINING LOOP
###############
EPOCHS = 10
for epoch in range(EPOCHS):
# Do Training
total_loss = 0.0
num_batches = 0
for batch in train_dist_dataset:
total_loss += distributed_train_step(batch)
num_batches += 1
train_loss = total_loss / num_batches
# Do Testing
for batch in test_dist_dataset:
distributed_test_step(batch)
template = ("Epoch {}, Loss: {}, Accuracy: {}, Test Loss: {}, " "Test Accuracy: {}")
print (template.format(epoch+1, train_loss, train_accuracy.result()*100, test_loss.result(), test_accuracy.result()*100))
test_loss.reset_states()
train_accuracy.reset_states()
test_accuracy.reset_states()
# + id="LIV6o9CX2bFH"
| Course 2 - Custom Training loops, Gradients and Distributed Training/Week 4 - Distribution Strategy/C2_W4_Lab_2_multi-GPU-mirrored-strategy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import maboss
model = maboss.load(
"epithelial_cell.bnd",
"epithelial_cell.cfg"
)
# ### Drawing the interaction graph
mini_model = maboss.to_minibn(model)
mini_model.influence_graph()
# ### Wild type simulation
res1 = model.run()
res1.plot_piechart()
# ### Virus simulation
model_virus = model.copy()
model_virus.network.set_istate(
'Virus_inside', [0, 1]
)
res_virus = model_virus.run()
res_virus.plot_piechart()
# ### CD8 simulation
model_cd8 = model.copy()
model_cd8.network.set_istate('TCell_attached', [0, 1])
res_cd8 = model_cd8.run()
res_cd8.plot_piechart()
# ### Virus + CD8 simulation
model_both = model.copy()
model_both.network.set_istate('Virus_inside', [0, 1])
model_both.network.set_istate('TCell_attached', [0, 1])
res_both = model_both.run()
res_both.plot_piechart()
# ### M mutant
model_m_mutant = model_virus.copy()
# model_m_mutant.mutate('M', 'OFF')
model_m_mutant.param['$M_ko'] = 1
res_m_mutant = model_m_mutant.run()
res_m_mutant.plot_piechart()
# ### FADD mutant
model_fadd_mutant = model_cd8.copy()
model_fadd_mutant.param['$FADD_ko'] = 1
res_fadd_mutant = model_fadd_mutant.run()
res_fadd_mutant.plot_piechart()
| notebooks/Apoptosis model analysis, v2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Check Duplicate Files
# ## Check Celeba Dataset
# +
import os
from PIL import Image, ImageStat
from tqdm import tqdm
import json
import hashlib
image_folder = os.path.join('../data/celeba/', 'img_align_celeba') #img_align_celeba check_dup
image_files = [_ for _ in os.listdir(image_folder) if _.endswith('jpg')]
duplicate_files = {}
checked_hash = {}
hash_dict = {}
def get_image_hash(file_path):
global hash_dict
if not file_path in hash_dict.keys():
img_path = os.path.join(image_folder, file_path)
hashstr = hashlib.sha1(open(img_path, 'rb').read()).hexdigest()
hash_dict[file_path] = hashstr
else:
hashstr = hash_dict[file_path]
return hashstr
for i, file_check in enumerate(tqdm(image_files)):
chk_hash = get_image_hash(file_check)
if chk_hash in checked_hash.keys():
dup_file = checked_hash[chk_hash]
if dup_file in duplicate_files.keys():
duplicate_files[dup_file].extend([file_check])
else:
duplicate_files[dup_file] = [file_check]
else:
checked_hash[chk_hash] = file_check
#print(checked_hash)
print(duplicate_files)
# -
from pprint import pprint
import collections
duplicate_files = collections.OrderedDict(duplicate_files)
pprint(duplicate_files)
json.dump(duplicate_files, open("duplicated.json",'w'))
# +
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import itertools
# %matplotlib inline
for batch in range(0, len(duplicate_files.items())//16):
duplicate_bat = dict(itertools.islice(duplicate_files.items(), batch*16, batch*16+16))
#print(duplicate_bat)
plt.figure(figsize=(10,20))
ncols = 4
c = 0
for i, (first, duplicates) in enumerate(duplicate_bat.items()):
nrows = len(duplicate_bat)
i_img = os.path.join('../data/celeba/img_align_celeba', first)
if (i+1)%2==0:
plt.subplot2grid((nrows,ncols), (i-1,c)) #(nrows, ncols), (row, col)
c += 1
else:
c = 0
plt.subplot2grid((nrows,ncols), (i,c))
c += 1
img = mpimg.imread(i_img)
plt.imshow(img)
plt.title(first)
plt.axis('off')
for j, mis in enumerate(duplicates):
m_img = os.path.join('../data/celeba/img_align_celeba', mis)
img = mpimg.imread(m_img)
if (i+1)%2==0:
plt.subplot2grid((nrows,ncols), (i-1,j+c)) #(nrows, ncols), (row, col)
else:
plt.subplot2grid((nrows,ncols), (i,j+c))
c += 1
plt.imshow(img)
plt.title(mis)
plt.axis('off')
plt.tight_layout()
# plt.subplots_adjust(hspace = .001)
plt.subplots_adjust(wspace=0, hspace=0)
#plt.subplot_tool()
plt.show()
break
# -
# ## Check Private Testset
# +
import os
from PIL import Image, ImageStat
from tqdm import tqdm
import json
import hashlib
from pprint import pprint
valid_images = [".jpg",".jpeg", ".gif",".png",".tiff"]
image_files = []
image_folder = '../data/testset/'
for dirname in os.listdir(image_folder):
dirpath = os.path.join(image_folder, dirname)
if os.path.isdir(dirpath):
for filename in os.listdir(dirpath):
ext = os.path.splitext(filename)[1]
if ext.lower() not in valid_images:
continue
image_files.append(os.path.join(dirpath, filename))
duplicate_files = {}
checked_hash = {}
hash_dict = {}
def get_image_hash(file_path):
global hash_dict
if not file_path in hash_dict.keys():
hashstr = hashlib.sha1(open(file_path, 'rb').read()).hexdigest()
hash_dict[file_path] = hashstr
else:
hashstr = hash_dict[file_path]
return hashstr
for i, file_check in enumerate(tqdm(image_files)):
chk_hash = get_image_hash(file_check)
if chk_hash in checked_hash.keys():
dup_file = checked_hash[chk_hash]
if dup_file in duplicate_files.keys():
duplicate_files[dup_file].extend([file_check])
else:
duplicate_files[dup_file] = [file_check]
else:
checked_hash[chk_hash] = file_check
#print(checked_hash)
pprint(duplicate_files)
# +
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import itertools
# %matplotlib inline
plt.figure(figsize=(10,10))
ncols = 2
c = 0
for i, (i_img, duplicates) in enumerate(duplicate_files.items()):
nrows = len(duplicate_files)
plt.subplot2grid((nrows,ncols), (i,0)) #(nrows, ncols), (row, col)
img = mpimg.imread(i_img)
plt.imshow(img)
plt.title(i_img)
plt.axis('off')
for j, mis in enumerate(duplicates):
m_img = mis
img = mpimg.imread(m_img)
plt.subplot2grid((nrows,ncols), (i,j+1)) #(nrows, ncols), (row, col)
plt.imshow(img)
plt.title(mis)
plt.axis('off')
plt.show()
# -
| project_1/src/EDA_Check_Duplicates.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from ipywidgets import widgets
x_range = widgets.FloatRangeSlider(value=[2,7],
min=0,
max=10.,
step=0.1,
description='X-Range: ',
readout_format='.1f')
display(x_range)
species_button = widgets.ToggleButton(
value=False,
description='Show Species')
display(species_button)
# +
feature_x_select = widgets.Dropdown(
value=2,
options=[('Sepal Length',0), ('Sepal Width',1),
('Petal Length',2), ('Petal Width',3)],
description='X-Axis:')
display(feature_x_select)
# -
| Chapter04/widgets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Distancias
from scipy.spatial import distance_matrix
import pandas as pd
data = pd.read_csv('../datasets/movies/movies.csv', sep = ';')
data
movies = data.columns.values.tolist()[1:]
movies
dd1 = distance_matrix(data[movies], data[movies], p = 1)
dd2 = distance_matrix(data[movies], data[movies], p = 2)
dd10 = distance_matrix(data[movies], data[movies], p = 10)
dd1
def dm_to_df (dd, col_name):
import pandas as pd
return pd.DataFrame(dd, index = col_name, columns = col_name)
dm_to_df(dd1, data['user_id'])
dm_to_df(dd2, data['user_id'])
dm_to_df(dd10, data['user_id'])
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
# +
fig = plt.figure()
ax = fig.add_subplot(111, projection = '3d')
ax.scatter(xs = data['star_wars'], ys = data['lord_of_the_rings'], zs = data['harry_potter'])
# -
# ### Enlaces
df = dm_to_df(dd1, data['user_id'])
df
Z = []
# +
df[11] = df[1] + df[10]
df.loc[11] = df.loc[1] + df.loc[10]
Z.append([1, 10, 0.7, 2])
df
# -
for i in df.columns.values.tolist():
df.loc[11][i] = min(df.loc[1][i], df.loc[10][i])
df.loc[i][11] = min(df.loc[i][1], df.loc[i][10])
df
df = df.drop([1, 10])
df = df.drop([1, 10], axis = 1)
df
| Jupyter Notebooks/Clustering - Distancias.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="11La92D7sMoN"
# + colab={"base_uri": "https://localhost:8080/"} id="5ddnT-ChTgyy" outputId="bc6c0765-d4fe-4319-8579-727cff147881"
# mounting google drive to access photos and dara
from google.colab import drive
drive.mount('/content/drive', force_remount=True)
# + id="-odkR3PcJu7u"
# dir = "/content/drive/My Drive/InfraredSolarModules/"
# + id="1jwqCXRHTuDJ"
import numpy as np
import keras as k
import json
import PIL
# + id="ucFbuQLaUXU4"
from keras.applications.vgg16 import preprocess_input
from keras.applications.vgg16 import VGG16
from keras.applications.resnet50 import preprocess_input
from keras.preprocessing import image
# + colab={"background_save": true, "base_uri": "https://localhost:8080/"} id="dDBPe4nNUC4U" outputId="88affa7a-bf76-4ee0-939b-558a35e64485"
# imports photo data from JSON file
def imagegen():
f = open('/content/drive/My Drive/InfraredSolarModules/module_metadata.json',)
meta = json.load(f)
f.close()
img = []
label = []
for i in range(9000,20000):
print("doing image"+str(i))
img.append(np.array(k.preprocessing.image.load_img('/content/drive/My Drive/InfraredSolarModules/' + meta[str(i)]['image_filepath'], target_size=(224, 224))))
if meta[str(i)]['anomaly_class'] == 'No-Anomaly':
label.append('No-Anomaly')
else:
label.append('Anomaly')
return np.array(img), np.array(label)
# generate images and labels
imgs, labels = imagegen()
print(imgs.shape)
print(len(imgs))
print(len(imgs.shape))
# + id="YPD0nHYmS9yv"
#generator function to load images in batches
def imageLoader(imgs, batch_size):
L = len(imgs)
# this line is just to make the generator infinite, keras needs that
while True:
batch_start = 0
batch_end = batch_size
while batch_start < L:
print(batch_start)
limit = min(batch_end, L)
X = imgs[batch_start:limit]
# Y = labels[batch_start:limit]
# break
yield X
batch_start += batch_size
batch_end += batch_size
# + colab={"base_uri": "https://localhost:8080/"} id="au1bXOVO-D_4" outputId="c8e2898f-5396-45c8-e1da-82813afb881e"
#preprocess images for feature extraction
def preprocess_image(imgs):
preprocessed_images = []
counter = 1
for image_item in imgs:
print("doing image {}".format(counter))
# image_item = image.img_to_array(image_item)
image_item = np.expand_dims(image_item, axis=0)
image_item = preprocess_input(image_item)
counter+=1
preprocess_image(imgs)
# + id="KG2KvH56UcIB"
model = VGG16(weights="imagenet", include_top=False, classes=2)
# + colab={"base_uri": "https://localhost:8080/"} id="2f5EWYv3TII0" outputId="5d8bab93-ac4f-4b79-dd05-6a10aae0d801"
# extract features
features = model.predict(imageLoader(imgs,100),steps=110)
# + colab={"base_uri": "https://localhost:8080/"} id="dsYpUGo703wM" outputId="1e708d89-55a1-46b5-e6bc-a11b35ee5906"
print(features.shape)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} id="PSF0prVENbCy" outputId="023874c1-f171-4f68-e2d1-c540530e468e"
type(imgs)
# + colab={"base_uri": "https://localhost:8080/"} id="b4L8_k6FcJ8o" outputId="e0f51190-2d90-4ceb-cdba-39cd61cc2091"
imgs.shape
# + id="BJYe9UCHvyKv"
from sklearn import svm
from sklearn.model_selection import train_test_split
from sklearn.linear_model import SGDClassifier
#reshape for training
features = features.reshape(11000,7*7*512)
#instantiate classifier model(Stochastic Gradient Decent)
svm_classifier = SGDClassifier()
#split data
X_train, X_test, y_train, y_test = train_test_split(features, labels, test_size=0.35)
# + colab={"base_uri": "https://localhost:8080/", "height": 272} id="d_wq7lXwsYSc" outputId="f5fc626d-ec51-4dcd-fb4e-915543cbf4d8"
print(X_train.shape)
print(X_train.shape)
print(y_train)
# + colab={"base_uri": "https://localhost:8080/"} id="DoIiQEamsWOo" outputId="7afa6516-33ff-429b-f32e-c34a8fc1017b"
#train
svm_classifier.fit(X_train, y_train)
#predict
y_pred = svm_classifier.predict(X_test)
from sklearn.metrics import confusion_matrix
from sklearn.metrics import f1_score, accuracy_score, precision_score
#metrics
print(confusion_matrix(y_test, y_pred))
print(accuracy_score(y_test, y_pred))
print(precision_score(y_test, y_pred, pos_label='Anomaly'))
print(f1_score(y_test, y_pred, pos_label='Anomaly'))
| models/binary/Detection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:latent_spacewalk]
# language: python
# name: conda-env-latent_spacewalk-py
# ---
# +
import argparse
import math
from pathlib import Path
import sys
from datetime import datetime
import os
import shutil
sys.path.append('./taming-transformers')
from IPython import display
from base64 import b64encode
from omegaconf import OmegaConf
from PIL import Image
from PIL.PngImagePlugin import PngInfo
from taming.models import cond_transformer, vqgan
import torch
from torch import nn, optim
from torch.nn import functional as F
from torchvision import transforms
from torchvision.transforms import functional as TF
from tqdm.notebook import tqdm
import clip
import kornia.augmentation as K
import numpy as np
import imageio
from PIL import ImageFile, Image
import taming
import json
import gc
# %load_ext autoreload
# %autoreload 2
# %config Completer.use_jedi = False
from vqgan_clip import *
# -
parameters = LatentSpacewalkParameters(
initial_image=None,
texts=['a minimalist watercolor painting of a red town'],
target_images= [],
seed= None,
max_iterations= 50,
learning_rate= 0.2,
save_interval= 1,
zoom_interval= None,
display_interval= 3,
)
parameters.prms
sw = Spacewalker(parameters=parameters, width=50, height=50)
empty_ram()
sw.run(parameters=parameters)
| scripts/vqgan_clip_scratch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Twitter Mining Function & Scatter Plots
# ---------------------------------------------------------------
#
# Import Dependencies
# %matplotlib notebook
import os
import csv
import json
import requests
from pprint import pprint
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from twython import Twython
import simplejson
import sys
import string
import glob
from pathlib import Path
# Import Twitter 'Keys' - MUST SET UP YOUR OWN 'config_twt.py' file
# You will need to create your own "config_twt.py" file using each of the Twitter authentication codes
# they provide you when you sign up for a developer account with your Twitter handle
from config_twt import (app_key_twt, app_secret_twt, oauth_token_twt, oauth_token_secret_twt)
# +
# Set Up Consumer Keys And Secret with Twitter Keys
APP_KEY = app_key_twt
APP_SECRET = app_secret_twt
# Set up OAUTH Token and Secret With Twitter Keys
OAUTH_TOKEN = oauth_token_twt
OAUTH_TOKEN_SECRET = oauth_token_secret_twt
# Load Keys In To a Twython Function And Call It "twitter"
twitter = Twython(APP_KEY, APP_SECRET, OAUTH_TOKEN, OAUTH_TOKEN_SECRET)
# Setup Batch Counter For Phase 2
batch_counter = 0
# -
# ___________________________
# ## Twitter Mining Function '(TMF)'
# ___________________________
#
# ### INSTRUCTIONS:
#
#
# This will Twitter Query Function will:
# - Perform searches for hastags (#)
# - Search for "@twitter_user_acct"
# - Provide mixed results of popular and most recent tweets from the last 7 days
# - 'remaining' 'search/tweets (allowance of 180) rate limit status' regenerates in 15 minutes after depletion
#
# ### Final outputs are aggegrated queries in both:
#
# - Pandas DataFrame of queried tweets
# - CSV files saved in the same folder as this Jupyter Notebook
#
#
# ### Phase 1 - Run Query and Store The Dictionary Into a List
#
#
# - Step 1) Run the 'Twitter Mining Function' cell below to begin program
# Note:
# - Limits have not been fully tested due to time constraint
# - Search up to 180 queries where and each query yields up to 100 tweets max
# - Running the TLC to see how many you have left after every csv outputs.
#
# - Step 2) When prompted, input in EITHER: #hashtag or @Twitter_user_account
# Examples: "#thuglife" or "@beyonce"
#
# - Step 3) TMF will query Twitter and store the tweets_data, a list called "all_data"
#
# - Step 4) Upon query search completion, it will prompt: "Perform another search query:' ('y'/'n') "
# - Input 'y' to query, and the program will append the results
# - Tip: Keep count of how many 'search tweets' you have, it should deduct 1 from 'remaining',
# which can produce up to 100 tweets of data
#
# - Step 5) End program by entering 'n' when prompted for 'search again'
# Output: printed list of all appended query data
#
#
# ### Phase 2 - Converting to Pandas DataFrame and Produce a CSV Output
#
# - Step 6) Loop Through Queried Data
#
# - Step 7) Convert to Pandas DataFrame
#
# - Step 8) Convert from DataFrame to CSV
#
# ### Addtional Considerations:
#
# - Current set up uses standard search api keys, not premium
# - TMF returns potentially 100 tweets at a time, and pulls from the last 7 days in random order
# - More than likely will have to run multiple searches and track line items count
# in each of the csv files output that will be created in the same folder
#
# ### Tweet Limit Counter (TLC)
# - Run cell to see how many search queries you have available
# - Your 'remaining' search tweets regenerates over 15 minutes.
# TLC - Run to Query Current Rate Limit on API Keys
twitter.get_application_rate_limit_status()['resources']['search']
# +
#Twitter Mining Function (TMF)
#RUN THIS CELL TO BEGIN PROGRAM!
print('-'*80)
print("TWITTER QUERY FUNCTION - BETA")
print('-'*80)
print("INPUT PARAMETERS:")
print("- @Twitter_handle e.g. @clashofclans")
print("- Hashtags (#) e.g. #THUGLIFE")
print("NOTE: SCROLL DOWN AFTER EACH QUERY FOR ENTER INPUT")
print('-'*80)
def twitter_search(app_search):
# Store the following Twython function and parameters into variable 't'
t = Twython(app_key=APP_KEY,
app_secret=APP_SECRET,
oauth_token=OAUTH_TOKEN,
oauth_token_secret=OAUTH_TOKEN_SECRET)
# The Twitter Mining Function we will use to run searches is below
# and we're asking for it to pull 100 tweets
search = t.search(q=app_search, count=100)
tweets = search['statuses']
# This will be a list of dictionaries of each tweet where the loop below will append to
all_data = []
# From the tweets, go into each individual tweet and extract the following into a 'dictionary'
# and append it to big bucket called 'all_data'
for tweet in tweets:
try:
tweets_data = {
"Created At":tweet['created_at'],
"Text (Tweet)":tweet['text'],
"User ID":tweet['user']['id'],
"User Followers Count":tweet['user']['followers_count'],
"Screen Name":tweet['user']['name'],
"ReTweet Count":tweet['retweet_count'],
"Favorite Count":tweet['favorite_count']}
all_data.append(tweets_data)
#print(tweets_data)
except (KeyError, NameError, TypeError, AttributeError) as err:
print(f"{err} Skipping...")
#functions need to return something...
return all_data
# The On and Off Mechanisms:
search_again = 'y'
final_all_data = []
# initialize the query counter
query_counter = 0
while search_again == 'y':
query_counter += 1
start_program = str(input('Type the EXACT @twitter_acct or #hashtag to query: '))
all_data = twitter_search(start_program)
final_all_data += all_data
#print(all_data)
print(f"Completed Collecting Search Results for {start_program} . Queries Completed: {query_counter} ")
print('-'*80)
search_again = input("Would you like to run another query? Enter 'y'. Otherwise, 'n' or another response will end query mode. ")
print('-'*80)
# When you exit the program, set the query counter back to zero
query_counter = 0
print()
print(f"Phase 1 of 2 Queries Completed . Proceed to Phase 2 - Convert Collection to DF and CSV formats .")
#print("final Data", final_all_data)
#####################################################################################################
# TIPS!: # If you're searching for the same hastag or twitter_handle,
# consider copying and pasting it (e.g. @fruitninja)
# -
# Display the total tweets the TMF successfully pulled:
print(len(final_all_data))
# ### Tweet Limit Counter (TLC)
# - Run cell to see how many search queries you have available
# - Your 'remaining' search tweets regenerates over 15 minutes.
# Run to view current rate limit status
twitter.get_application_rate_limit_status()['resources']['search']
# +
#df = pd.DataFrame(final_all_data[0])
#df
final_all_data
# -
# ### Step 6) Loop through the stored list of queried tweets from final_all_data and stores in designated lists
# +
# Loop thru finall_all_data (list of dictionaries) and extract each item and store them into
# the respective lists
# BUCKETS
created_at = []
tweet_text = []
user_id = []
user_followers_count = []
screen_name = []
retweet_count = []
likes_count = []
# append tweets data to the buckets for each tweet
#change to final_all_data
for data in final_all_data:
#print(keys, data[keys])
created_at.append(data["Created At"]),
tweet_text.append(data['Text (Tweet)']),
user_id.append(data['User ID']),
user_followers_count.append(data['User Followers Count']),
screen_name.append(data['Screen Name']),
retweet_count.append(data['ReTweet Count']),
likes_count.append(data['Favorite Count'])
#print(created_at, tweet_text, user_id, user_followers_count, screen_name, retweet_count, likes_count)
print("Run complete. Proceed to next cell.")
# -
# ### Step 7) Convert to Pandas DataFrame
# +
# Setup DataFrame and run tweets_data_df
tweets_data_df = pd.DataFrame({
"Created At": created_at,
"Screen Name": screen_name,
"User ID": user_id,
"User Follower Count": user_followers_count,
"Likes Counts": likes_count,
"ReTweet Count": retweet_count,
"Tweet Text" : tweet_text
})
tweets_data_df.head()
# -
# ### Step 8) Load into MySQL Database - later added this piece to display ETL potential of this project
# This section was added later after I reviewed and wanted to briefly reiterate on it
tweets_data_df2 = tweets_data_df.copy()
# Dropped Screen Name and Tweets Text bc would I would need to clean the 'Screen Name' and 'Tweet Text' Columns
tweets_data_df2 = tweets_data_df2.drop(["Screen Name", "Tweet Text"], axis=1).sort_values(by="User Follower Count")
# +
# Import Dependencies 2/2:
from sqlalchemy import create_engine
from sqlalchemy.sql import select
from sqlalchemy_utils import database_exists, create_database, drop_database, has_index
import pymysql
rds_connection_string = "root:PASSWORD_HERE@127.0.0.1/"
#db_name = input("What database would you like to search for?")
db_name = 'twitterAPI_data_2019_db'
# Setup engine connection string
engine = create_engine(f'mysql://{rds_connection_string}{db_name}?charset=utf8', echo=True)
# -
# Created a function incorproating SQL Alchemy to search, create, and or drop a database:
def search_create_drop_db(db_name):
db_exist = database_exists(f'mysql://{rds_connection_string}{db_name}')
db_url = f'mysql://{rds_connection_string}{db_name}'
if db_exist == True:
drop_table_y_or_n = input(f'"{db_name}" database already exists in MySQL. Do you want you drop the table? Enter exactly: "y" or "n". ')
if drop_table_y_or_n == 'y':
drop_database(db_url)
print(f"Database {db_name} was dropped")
create_new_db = input(f"Do you want to create another database called: {db_name}? ")
if create_new_db == 'y':
create_database(db_url)
return(f"The database {db_name} was created. Next You will need to create tables for this database. ")
else:
return("No database was created. Goodbye! ")
else:
return("The database exists. No action was taken. Goodbye! ")
else:
create_database(db_url)
return(f"The queried database did not exist, and was created as: {db_name} . ")
search_create_drop_db(db_name)
tweets_data_df2.to_sql('tweets', con=engine, if_exists='append')
# ### Step 9) Convert DataFrame to CSV File and save on local drive
# +
# Save Tweets Data to a CSV File (Run Cell to input filename)
# Streamline the saving of multiple queries (1 query = up to 100 tweets) into a csv file.
# E.g. input = (#fruit_ninja) will save the file as "fruit_ninja_batch1.csv" as the file result
# Note: first chracter will be slice off so you can just copy and paste
# the hastag / @twitter_handle from steps above
batch_name = str(input("Enter in batch name."))
# If you restart kernel, batch_counter resets to zero.
batch_counter = batch_counter +1
# Check if the #hastag / @twitter_handle folder exists and create the folder if it does not
Path(f"./resources/{batch_name[1:]}").mkdir(parents=True, exist_ok=True)
# Save dataframe of all queries in a csv file to a folder in the resources folder csv using the
tweets_data_df.to_csv(f"./resources/{batch_name[1:]}/{batch_name[1:]}_batch{batch_counter}.csv", encoding='utf-8')
print(f"Output saved in current folder as: {batch_name[1:]}_batch{batch_counter}.csv ")
# -
# # PHASE 3 - CALCULATIONS USING API DATA
#
# +
# This prints out all of the folder titles in "resources" folder
path = './resources/*' # use your path
resources = glob.glob(path)
all_folders = []
print("All folders in the 'resources' folder:")
print("="*40)
for foldername in resources:
str(foldername)
foldername = foldername[12:]
all_folders.append(foldername)
#print(li)
print("")
print(F"Total Folders: {len(all_folders)}")
# -
print(all_folders)
# +
all_TopApps_df_list = []
for foldername in all_folders:
plug = foldername
path = f'./resources\\{plug}'
all_files = glob.glob(path + "/*.csv")
counter = 0
app_dataframes = []
for filename in all_files:
counter += 1
df = pd.read_csv(filename, index_col=None, header=0)
app_dataframes.append(df)
output = pd.concat(app_dataframes, axis=0, ignore_index=True)
all_TopApps_df_list.append(f"{output}_{counter}")
counter = 0
#fb_frame
# -
# ##### Facebook Calculations
#
# +
# Example Template of looping thru csvfiles, and concatenate all of the csv files we collected in each folder
plug = 'facebook'
path = f'./resources\\{plug}'
all_files = glob.glob(path + "/*.csv")
li = []
for filename in all_files:
df = pd.read_csv(filename, index_col=None, header=0)
li.append(df)
fb_frame = pd.concat(li, axis=0, ignore_index=True)
fb_frame
# -
fb_frame.describe()
# Sort to set up removal of duplicates
fb_frame.sort_values(by=['User ID','Created At'], ascending=False)
# Drop Duplicates only for matching columns omitting the index
facebook_filtered_df = fb_frame.drop_duplicates(['Created At', 'Screen Name', 'User ID', 'User Follower Count', 'Likes Counts', 'ReTweet Count', 'Tweet Text']).sort_values(by=['User ID','Created At'], ascending=False)
# Get New Snap Shot Statistics
facebook_filtered_df.describe()
facebook_filtered_df.head()
# Count total out of Unique Tweets
facebook_total_tweets = len(facebook_filtered_df['Tweet Text'])
facebook_total_tweets
# Calculate Facebook Avg Followers - doesn't make sense to sum.
facebook_avg_followers_ct = facebook_filtered_df['User Follower Count'].mean()
facebook_avg_followers_ct
# Total Likes of all tweets
facebook_total_likes = facebook_filtered_df['Likes Counts'].sum()
#facebook_avg_likes = facebook_filtered_df['Likes Counts'].mean()
facebook_total_likes
#facebook_avg_likes
# Facebook Retweets Stats:
#facebook_sum_retweets = facebook_filtered_df['ReTweet Count'].sum()
facebook_avg_retweets = facebook_filtered_df['ReTweet Count'].mean()
#facebook_sum_retweets
facebook_avg_retweets
# #### Instagram Calculations
# +
plug = 'instagram'
path = f'./resources/{plug}'
all_files = glob.glob(path + "/*.csv")
li = []
for filename in all_files:
df = pd.read_csv(filename, index_col=None, header=0)
li.append(df)
instagram_source_df = pd.concat(li, axis=0, ignore_index=True)
instagram_source_df
# -
# Snapshot Statistics
instagram_source_df.describe()
instagram_source_df.head()
# Sort to set up removal of duplicates
instagram_source_df.sort_values(by=['User ID','Created At'], ascending=False)
# Drop Duplicates only for matching columns omitting the index
instagram_filtered_df = instagram_source_df.drop_duplicates(['Created At', 'Screen Name', 'User ID', 'User Follower Count', 'Likes Counts', 'ReTweet Count', 'Tweet Text']).sort_values(by=['User ID','Created At'], ascending=False)
instagram_filtered_df
# Get New Snap Shot Statistics
instagram_filtered_df.describe()
# Count total out of Unique Tweets
instagram_total_tweets = len(instagram_filtered_df['Tweet Text'])
instagram_total_tweets
# Calculate Facebook Avg Followers - doesn't make sense to sum.
instagram_avg_followers_ct = instagram_filtered_df['User Follower Count'].mean()
instagram_avg_followers_ct
# Total Likes of all tweets
instagram_total_likes = instagram_filtered_df['Likes Counts'].sum()
#instagram_avg_likes = instagram_filtered_df['Likes Counts'].mean()
instagram_total_likes
#instagram_avg_likes
# Retweets Stats:
#instagram_sum_retweets = instagram_filtered_df['ReTweet Count'].sum()
instagram_avg_retweets = instagram_filtered_df['ReTweet Count'].mean()
#instagram_sum_retweets
instagram_avg_retweets
# ### Clash of Clans Calculations
# +
plug = 'clashofclans'
path = f'./resources/{plug}'
all_files = glob.glob(path + "/*.csv")
li = []
for filename in all_files:
df = pd.read_csv(filename, index_col=None, header=0)
li.append(df)
coc_source_df = pd.concat(li, axis=0, ignore_index=True)
coc_source_df
# Snapshot Statistics
coc_source_df.describe()
# -
coc_source_df.head()
# Sort to set up removal of duplicates
coc_source_df.sort_values(by=['User ID','Created At'], ascending=False)
# Drop Duplicates only for matching columns omitting the index
coc_filtered_df = coc_source_df.drop_duplicates(['Created At', 'Screen Name', 'User ID', 'User Follower Count', 'Likes Counts', 'ReTweet Count', 'Tweet Text']).sort_values(by=['User ID','Created At'], ascending=False)
coc_filtered_df.head()
# Get New Snap Shot Statistics
coc_filtered_df.describe()
# Count total out of Unique Tweets
coc_total_tweets = len(coc_filtered_df['Tweet Text'])
coc_total_tweets
# Calculate Facebook Avg Followers - doesn't make sense to sum.
coc_avg_followers_ct = coc_filtered_df['User Follower Count'].mean()
coc_avg_followers_ct
# Total Likes of all tweets
coc_total_likes = coc_filtered_df['Likes Counts'].sum()
#coc_avg_likes = coc_filtered_df['Likes Counts'].mean()
coc_total_likes
#coc_avg_likes
# Retweets Stats:
#coc_sum_retweets = coc_filtered_df['ReTweet Count'].sum()
coc_avg_retweets = coc_filtered_df['ReTweet Count'].mean()
#coc_sum_retweets
coc_avg_retweets
# ### Temple Run Calculations
#
# +
plug = 'templerun'
path = f'./resources/{plug}'
all_files = glob.glob(path + "/*.csv")
li = []
for filename in all_files:
df = pd.read_csv(filename, index_col=None, header=0)
li.append(df)
templerun_source_df = pd.concat(li, axis=0, ignore_index=True)
templerun_source_df
# Snapshot Statistics
templerun_source_df.describe()
# +
#templerun_source_df.head()
# -
# Sort to set up removal of duplicates
templerun_source_df.sort_values(by=['User ID','Created At'], ascending=False)
# Drop Duplicates only for matching columns omitting the index
templerun_filtered_df = templerun_source_df.drop_duplicates(['Created At', 'Screen Name', 'User ID', 'User Follower Count', 'Likes Counts', 'ReTweet Count', 'Tweet Text']).sort_values(by=['User ID','Created At'], ascending=False)
# +
#templerun_filtered_df
# +
#templerun_filtered_df.describe()
# -
# Count total out of Unique Tweets
templerun_total_tweets = len(templerun_filtered_df['Tweet Text'])
templerun_total_tweets
# Calculate Facebook Avg Followers - doesn't make sense to sum.
templerun_avg_followers_ct = templerun_filtered_df['User Follower Count'].mean()
templerun_avg_followers_ct
# Total Likes of all tweets
templerun_total_likes = templerun_filtered_df['Likes Counts'].sum()
#templerun_avg_likes = templerun_filtered_df['Likes Counts'].mean()
templerun_total_likes
#instagram_avg_likes
# Retweets Stats:
#templerun_sum_retweets = templerun_filtered_df['ReTweet Count'].sum()
templerun_avg_retweets = templerun_filtered_df['ReTweet Count'].mean()
#templerun_sum_retweets
templerun_avg_retweets
templerun_total_tweets
templerun_avg_retweets
templerun_avg_followers_ct
templerun_total_likes
# ### Pandora Calculations
# +
plug = 'pandora'
path = f'./resources/{plug}'
all_files = glob.glob(path + "/*.csv")
li = []
for filename in all_files:
df = pd.read_csv(filename, index_col=None, header=0)
li.append(df)
pandora_source_df = pd.concat(li, axis=0, ignore_index=True)
# Snapshot Statistics
pandora_source_df.describe()
# +
#pandora_source_df.head()
# -
# Sort to set up removal of duplicates
pandora_source_df.sort_values(by=['User ID','Created At'], ascending=False)
# Drop Duplicates only for matching columns omitting the index
pandora_filtered_df = pandora_source_df.drop_duplicates(['Created At', 'Screen Name', 'User ID', 'User Follower Count', 'Likes Counts', 'ReTweet Count', 'Tweet Text']).sort_values(by=['User ID','Created At'], ascending=False)
pandora_filtered_df
pandora_filtered_df.describe()
# Count total out of Unique Tweets
pandora_total_tweets = len(pandora_filtered_df['Tweet Text'])
pandora_total_tweets
# Calculate Facebook Avg Followers - doesn't make sense to sum.
pandora_avg_followers_ct = pandora_filtered_df['User Follower Count'].mean()
pandora_avg_followers_ct
# Total Likes of all tweets
# use sum of likes.
pandora_total_likes = pandora_filtered_df['Likes Counts'].sum()
#pandora_avg_likes = pandora_filtered_df['Likes Counts'].mean()
pandora_total_likes
#pandora_avg_likes
# Retweets Stats:
#pandora_sum_retweets = pandora_filtered_df['ReTweet Count'].sum()
pandora_avg_retweets = pandora_filtered_df['ReTweet Count'].mean()
#pandora_sum_retweets
pandora_avg_retweets
# ### Pinterest Calculations
# +
# Concatenate them
plug = 'pinterest'
path = f'./resources/{plug}'
all_files = glob.glob(path + "/*.csv")
li = []
for filename in all_files:
df = pd.read_csv(filename, index_col=None, header=0)
li.append(df)
pinterest_source_df = pd.concat(li, axis=0, ignore_index=True)
# Snapshot Statistics
pinterest_source_df.describe()
# -
pinterest_source_df.head()
# Sort to set up removal of duplicates
pinterest_source_df.sort_values(by=['User ID','Created At'], ascending=False)
# Drop Duplicates only for matching columns omitting the index
pinterest_filtered_df = pinterest_source_df.drop_duplicates(['Created At', 'Screen Name', 'User ID', 'User Follower Count', 'Likes Counts', 'ReTweet Count', 'Tweet Text']).sort_values(by=['User ID','Created At'], ascending=False)
pinterest_filtered_df
pinterest_filtered_df.describe()
# Count total out of Unique Tweets
pinterest_total_tweets = len(pinterest_filtered_df['Tweet Text'])
pinterest_total_tweets
# Calculate Facebook Avg Followers - doesn't make sense to sum.
pinterest_avg_followers_ct = pinterest_filtered_df['User Follower Count'].mean()
pinterest_avg_followers_ct
# Total Likes of all tweets
pinterest_total_likes = pinterest_filtered_df['Likes Counts'].sum()
#pinterest_avg_likes = pinterest_filtered_df['Likes Counts'].mean()
pinterest_total_likes
#pinterest_avg_likes
# Retweets Stats:
#pinterest_sum_retweets = pinterest_filtered_df['ReTweet Count'].sum()
pinterest_avg_retweets = pinterest_filtered_df['ReTweet Count'].mean()
#pinterest_sum_retweets
pinterest_avg_retweets
# ### Bible (You Version) Calculations
plug = 'bible'
path = f'./resources/{plug}'
all_files = glob.glob(path + "/*.csv")
li = []
for filename in all_files:
df = pd.read_csv(filename, index_col=None, header=0)
li.append(df)
bible_source_df = pd.concat(li, axis=0, ignore_index=True)
bible_source_df
# Snapshot Statistics
bible_source_df.describe()
bible_source_df.head()
# Sort to set up removal of duplicates
bible_source_df.sort_values(by=['User ID','Created At'], ascending=False)
# Drop Duplicates only for matching columns omitting the index
bible_filtered_df = bible_source_df.drop_duplicates(['Created At', 'Screen Name', 'User ID', 'User Follower Count', 'Likes Counts', 'ReTweet Count', 'Tweet Text']).sort_values(by=['User ID','Created At'], ascending=False)
bible_filtered_df
bible_filtered_df.describe()
# Count total out of Unique Tweets
bible_total_tweets = len(bible_filtered_df['Tweet Text'])
bible_total_tweets
# Calculate Avg Followers - doesn't make sense to sum.
bible_avg_followers_ct = bible_filtered_df['User Follower Count'].mean()
bible_avg_followers_ct
# Total Likes of all tweets
bible_total_likes = bible_filtered_df['Likes Counts'].sum()
#bible_avg_likes = bible_filtered_df['Likes Counts'].mean()
bible_total_likes
#bible_avg_likes
# Retweets Stats:
#bible_sum_retweets = bible_filtered_df['ReTweet Count'].sum()
bible_avg_retweets = bible_filtered_df['ReTweet Count'].mean()
#bible_sum_retweets
bible_avg_retweets
# ### Candy Crush Saga Calculations
# +
plug = 'candycrushsaga'
path = f'./resources/{plug}'
all_files = glob.glob(path + "/*.csv")
li = []
for filename in all_files:
df = pd.read_csv(filename, index_col=None, header=0)
li.append(df)
CandyCrushSaga_source_df = pd.concat(li, axis=0, ignore_index=True)
# Snapshot Statistics
CandyCrushSaga_source_df.describe()
# -
# has duplicates
CandyCrushSaga_source_df.sort_values(by=['User ID','Created At'], ascending=False)
CandyCrushSaga_source_df.head()
# Drop Duplicates only for matching columns omitting the index
CandyCrushSaga_filtered_df = CandyCrushSaga_source_df.drop_duplicates(['Created At', 'Screen Name', 'User ID', 'User Follower Count', 'Likes Counts', 'ReTweet Count', 'Tweet Text']).sort_values(by=['User ID','Created At'], ascending=False)
# Get New Snap Shot Statistics
CandyCrushSaga_filtered_df.describe()
CandyCrushSaga_filtered_df.head()
# Count total out of Unique Tweets
candycrushsaga_total_tweets = len(CandyCrushSaga_filtered_df['Tweet Text'])
candycrushsaga_total_tweets
# Calculate Avg Followers - doesn't make sense to sum.
candycrushsaga_avg_followers_ct = CandyCrushSaga_filtered_df['User Follower Count'].mean()
candycrushsaga_avg_followers_ct
# Total Likes of all tweets
candycrushsaga_total_likes = CandyCrushSaga_filtered_df['Likes Counts'].sum()
#facebook_avg_likes = facebook_filtered_df['Likes Counts'].mean()
candycrushsaga_total_likes
#facebook_avg_likes
# Retweets Stats:
#facebook_sum_retweets = facebook_filtered_df['ReTweet Count'].sum()
candycrushsaga_avg_retweets = CandyCrushSaga_filtered_df['ReTweet Count'].mean()
#facebook_sum_retweets
candycrushsaga_avg_retweets
# ### Spotify Music Caculations
# +
plug = 'spotify'
path = f'./resources/{plug}'
all_files = glob.glob(path + "/*.csv")
li = []
for filename in all_files:
df = pd.read_csv(filename, index_col=None, header=0)
li.append(df)
spotify_source_df = pd.concat(li, axis=0, ignore_index=True)
# Snapshot Statistics
spotify_source_df.describe()
# -
spotify_source_df.head()
# Sort to set up removal of duplicates
spotify_source_df.sort_values(by=['User ID','Created At'], ascending=False)
# Drop Duplicates only for matching columns omitting the index
spotify_filtered_df = spotify_source_df.drop_duplicates(['Created At', 'Screen Name', 'User ID', 'User Follower Count', 'Likes Counts', 'ReTweet Count', 'Tweet Text']).sort_values(by=['User ID','Created At'], ascending=False)
spotify_filtered_df
spotify_filtered_df.describe()
# Count total out of Unique Tweets
spotify_total_tweets = len(spotify_filtered_df['Tweet Text'])
spotify_total_tweets
# Calculate Facebook Avg Followers - doesn't make sense to sum.
spotify_avg_followers_ct = spotify_filtered_df['User Follower Count'].mean()
spotify_avg_followers_ct
# Total Likes of all tweets
spotify_total_likes = spotify_filtered_df['Likes Counts'].sum()
#spotify_avg_likes = spotify_filtered_df['Likes Counts'].mean()
spotify_total_likes
#spotify_avg_likes
# Retweets Stats:
#spotify_sum_retweets = spotify_filtered_df['ReTweet Count'].sum()
spotify_avg_retweets = spotify_filtered_df['ReTweet Count'].mean()
#spotify_sum_retweets
spotify_avg_retweets
# ### Angry Birds Calculations
#
# +
plug = 'angrybirds'
path = f'./resources/{plug}'
all_files = glob.glob(path + "/*.csv")
li = []
for filename in all_files:
df = pd.read_csv(filename, index_col=None, header=0)
li.append(df)
angrybirds_source_df = pd.concat(li, axis=0, ignore_index=True)
# Snapshot Statistics
angrybirds_source_df.describe()
# -
angrybirds_source_df.head()
# Sort to set up removal of duplicates
angrybirds_source_df.sort_values(by=['User ID','Created At'], ascending=False)
# Drop Duplicates only for matching columns omitting the index
angrybirds_filtered_df = angrybirds_source_df.drop_duplicates(['Created At', 'Screen Name', 'User ID', 'User Follower Count', 'Likes Counts', 'ReTweet Count', 'Tweet Text']).sort_values(by=['User ID','Created At'], ascending=False)
angrybirds_filtered_df
angrybirds_filtered_df.describe()
# Count total out of Unique Tweets
angrybirds_total_tweets = len(angrybirds_filtered_df['Tweet Text'])
angrybirds_total_tweets
# Calculate angrybirds Avg Followers - doesn't make sense to sum.
angrybirds_avg_followers_ct = angrybirds_filtered_df['User Follower Count'].mean()
angrybirds_avg_followers_ct
# Total Likes of all tweets
angrybirds_total_likes = angrybirds_filtered_df['Likes Counts'].sum()
#angrybirds_avg_likes = angrybirds_filtered_df['Likes Counts'].mean()
angrybirds_total_likes
#angrybirds_avg_likes
# Retweets Stats:
#angrybirds_sum_retweets = angrybirds_filtered_df['ReTweet Count'].sum()
angrybirds_avg_retweets = angrybirds_filtered_df['ReTweet Count'].mean()
#angrybirds_sum_retweets
angrybirds_avg_retweets
# ### YouTube Calculations
# +
plug = 'youtube'
path = f'./resources/{plug}'
all_files = glob.glob(path + "/*.csv")
li = []
for filename in all_files:
df = pd.read_csv(filename, index_col=None, header=0)
li.append(df)
youtube_source_df = pd.concat(li, axis=0, ignore_index=True)
# Snapshot Statistics
youtube_source_df.describe()
# -
youtube_source_df.head()
# Sort
youtube_source_df.sort_values(by=['User ID','Created At'], ascending=False)
# Drop Duplicates only for matching columns omitting the index
youtube_filtered_df = youtube_source_df.drop_duplicates(['Created At', 'Screen Name', 'User ID', 'User Follower Count', 'Likes Counts', 'ReTweet Count', 'Tweet Text']).sort_values(by=['User ID','Created At'], ascending=False)
# Get New Snap Shot Statistics
youtube_filtered_df.describe()
youtube_filtered_df.head()
# Count total out of Unique Tweets
youtube_total_tweets = len(youtube_filtered_df['Tweet Text'])
youtube_total_tweets
# Calculate Facebook Avg Followers - doesn't make sense to sum.
youtube_avg_followers_ct = youtube_filtered_df['User Follower Count'].mean()
youtube_avg_followers_ct
# Total Likes of all tweets
# use sum of likes.
youtube_total_likes = youtube_filtered_df['Likes Counts'].sum()
#youtube_avg_likes = youtube_filtered_df['Likes Counts'].mean()
youtube_total_likes
#youtube_avg_likes
# You Tube Retweets Stats:
#youtube_sum_retweets = facebook_filtered_df['ReTweet Count'].sum()
youtube_avg_retweets = youtube_filtered_df['ReTweet Count'].mean()
#youtube_sum_retweets
youtube_avg_retweets
# ### Subway Surfers
# +
plug = 'subwaysurfer'
path = f'./resources/{plug}'
all_files = glob.glob(path + "/*.csv")
li = []
for filename in all_files:
df = pd.read_csv(filename, index_col=None, header=0)
li.append(df)
SubwaySurfers_source_df = pd.concat(li, axis=0, ignore_index=True)
# Snapshot Statistics
SubwaySurfers_source_df.describe()
# -
SubwaySurfers_source_df.head()
# Sort
SubwaySurfers_source_df.sort_values(by=['User ID','Created At'], ascending=False)
SubwaySurfers_source_df.head()
# Drop Duplicates only for matching columns omitting the index
SubwaySurfers_filtered_df = SubwaySurfers_source_df.drop_duplicates(['Created At', 'Screen Name', 'User ID', 'User Follower Count', 'Likes Counts', 'ReTweet Count', 'Tweet Text']).sort_values(by=['User ID','Created At'], ascending=False)
# Get New Snap Shot Statistics
SubwaySurfers_filtered_df.describe()
SubwaySurfers_filtered_df.head()
# Count total out of Unique Tweets
SubwaySurfers_total_tweets = len(SubwaySurfers_filtered_df['Tweet Text'])
SubwaySurfers_total_tweets
# Calculate Avg Followers - doesn't make sense to sum.
SubwaySurfers_avg_followers_ct = SubwaySurfers_filtered_df['User Follower Count'].mean()
SubwaySurfers_avg_followers_ct
# Total Likes of all tweets
SubwaySurfers_total_likes = SubwaySurfers_filtered_df['Likes Counts'].sum()
#SubwaySurfers_avg_likes = SubwaySurfers_filtered_df['Likes Counts'].mean()
SubwaySurfers_total_likes
#SubwaySurfers_avg_likes
# Subway Surfer Retweets Stats:
#SubwaySurfers_sum_retweets = SubwaySurfers_filtered_df['ReTweet Count'].sum()
SubwaySurfers_avg_retweets = SubwaySurfers_filtered_df['ReTweet Count'].mean()
#SubwaySurfers_sum_retweets
SubwaySurfers_avg_retweets
# ### Security Master - Antivirus, VPN
# +
# Cheetah Mobile owns Security Master
plug = 'cheetah'
path = f'./resources/{plug}'
all_files = glob.glob(path + "/*.csv")
li = []
for filename in all_files:
df = pd.read_csv(filename, index_col=None, header=0)
li.append(df)
SecurityMaster_source_df = pd.concat(li, axis=0, ignore_index=True)
# Snapshot Statistics
SecurityMaster_source_df.describe()
# -
SecurityMaster_source_df.head()
# has duplicates
SecurityMaster_source_df.sort_values(by=['User ID','Created At'], ascending=False)
SecurityMaster_source_df.head()
# Drop Duplicates only for matching columns omitting the index
SecurityMaster_filtered_df = SecurityMaster_source_df.drop_duplicates(['Created At', 'Screen Name', 'User ID', 'User Follower Count', 'Likes Counts', 'ReTweet Count', 'Tweet Text']).sort_values(by=['User ID','Created At'], ascending=False)
# Get New Snap Shot Statistics
SecurityMaster_filtered_df.describe()
SecurityMaster_filtered_df.head()
# Count total out of Unique Tweets
SecurityMaster_total_tweets = len(SecurityMaster_filtered_df['Tweet Text'])
SecurityMaster_total_tweets
# Calculate Avg Followers - doesn't make sense to sum.
SecurityMaster_avg_followers_ct = SecurityMaster_filtered_df['User Follower Count'].mean()
SecurityMaster_avg_followers_ct
# Total Likes of all tweets
SecurityMaster_total_likes = SecurityMaster_filtered_df['Likes Counts'].sum()
#SecurityMaster_avg_likes = SecurityMaster_filtered_df['Likes Counts'].mean()
SecurityMaster_total_likes
#SecurityMaster_avg_likes
# Security Master Retweets Stats:
#SecurityMaster_sum_retweets = SecurityMaster_filtered_df['ReTweet Count'].sum()
SecurityMaster_avg_retweets = SecurityMaster_filtered_df['ReTweet Count'].mean()
#SecurityMaster_sum_retweets
SecurityMaster_avg_retweets
# ### Clash Royale
# +
plug = 'clashroyale'
path = f'./resources/{plug}'
all_files = glob.glob(path + "/*.csv")
li = []
for filename in all_files:
df = pd.read_csv(filename, index_col=None, header=0)
li.append(df)
ClashRoyale_source_df = pd.concat(li, axis=0, ignore_index=True)
# Snapshot Statistics
ClashRoyale_source_df.describe()
# -
ClashRoyale_source_df.head()
# has duplicates
ClashRoyale_source_df.sort_values(by=['User ID','Created At'], ascending=False)
# Drop Duplicates only for matching columns omitting the index
ClashRoyale_filtered_df = ClashRoyale_source_df.drop_duplicates(['Created At', 'Screen Name', 'User ID', 'User Follower Count', 'Likes Counts', 'ReTweet Count', 'Tweet Text']).sort_values(by=['User ID','Created At'], ascending=False)
# Get New Snap Shot Statistics
ClashRoyale_filtered_df.describe()
ClashRoyale_filtered_df.head()
# Count total out of Unique Tweets
ClashRoyale_total_tweets = len(ClashRoyale_filtered_df['Tweet Text'])
ClashRoyale_total_tweets
# Calculate Avg Followers - doesn't make sense to sum.
ClashRoyale_avg_followers_ct = ClashRoyale_filtered_df['User Follower Count'].mean()
ClashRoyale_avg_followers_ct
# Total Likes of all tweets
ClashRoyale_total_likes = ClashRoyale_filtered_df['Likes Counts'].sum()
#ClashRoyale_avg_likes = ClashRoyale_filtered_df['Likes Counts'].mean()
ClashRoyale_total_likes
#ClashRoyale_avg_likes
# ClashRoyale Retweets Stats:
#ClashRoyale_sum_retweets = ClashRoyale_filtered_df['ReTweet Count'].sum()
ClashRoyale_avg_retweets = ClashRoyale_filtered_df['ReTweet Count'].mean()
#facebook_sum_retweets
ClashRoyale_avg_retweets
# ### Clean Master - Space Cleaner
# +
plug = 'cleanmaster'
path = f'./resources/{plug}'
all_files = glob.glob(path + "/*.csv")
li = []
for filename in all_files:
df = pd.read_csv(filename, index_col=None, header=0)
li.append(df)
CleanMaster_source_df = pd.concat(li, axis=0, ignore_index=True)
# Snapshot Statistics
CleanMaster_source_df.describe()
# -
CleanMaster_source_df.head()
# has duplicates
CleanMaster_source_df.sort_values(by=['User ID','Created At'], ascending=False)
CleanMaster_source_df.head()
# Drop Duplicates only for matching columns omitting the index
CleanMaster_filtered_df = CleanMaster_source_df.drop_duplicates(['Created At', 'Screen Name', 'User ID', 'User Follower Count', 'Likes Counts', 'ReTweet Count', 'Tweet Text']).sort_values(by=['User ID','Created At'], ascending=False)
# Get New Snap Shot Statistics
CleanMaster_filtered_df.describe()
CleanMaster_filtered_df.head()
# Count total out of Unique Tweets
CleanMaster_total_tweets = len(CleanMaster_filtered_df['Tweet Text'])
CleanMaster_total_tweets
# Calculate Avg Followers - doesn't make sense to sum.
CleanMaster_avg_followers_ct = CleanMaster_filtered_df['User Follower Count'].mean()
CleanMaster_avg_followers_ct
# Total Likes of all tweets
CleanMaster_total_likes = CleanMaster_filtered_df['Likes Counts'].sum()
#facebook_avg_likes = facebook_filtered_df['Likes Counts'].mean()
CleanMaster_total_likes
#facebook_avg_likes
# Clean MasterRetweets Stats:
#CleanMaster_sum_retweets = CleanMaster_filtered_df['ReTweet Count'].sum()
CleanMaster_avg_retweets = CleanMaster_filtered_df['ReTweet Count'].mean()
#facebook_sum_retweets
CleanMaster_avg_retweets
# ### What's App
# +
plug = 'whatsapp'
path = f'./resources/{plug}'
all_files = glob.glob(path + "/*.csv")
li = []
for filename in all_files:
df = pd.read_csv(filename, index_col=None, header=0)
li.append(df)
whatsapp_source_df = pd.concat(li, axis=0, ignore_index=True)
# Snapshot Statistics
whatsapp_source_df.describe()
# -
whatsapp_source_df.head()
# has duplicates
whatsapp_source_df.sort_values(by=['User ID','Created At'], ascending=False)
whatsapp_source_df.head()
# Drop Duplicates only for matching columns omitting the index
whatsapp_filtered_df = whatsapp_source_df.drop_duplicates(['Created At', 'Screen Name', 'User ID', 'User Follower Count', 'Likes Counts', 'ReTweet Count', 'Tweet Text']).sort_values(by=['User ID','Created At'], ascending=False)
# Get New Snap Shot Statistics
whatsapp_filtered_df.describe()
whatsapp_filtered_df.head()
# Count total out of Unique Tweets
whatsapp_total_tweets = len(whatsapp_filtered_df['Tweet Text'])
whatsapp_total_tweets
# Calculate Facebook Avg Followers - doesn't make sense to sum.
whatsapp_avg_followers_ct = whatsapp_filtered_df['User Follower Count'].mean()
whatsapp_avg_followers_ct
# Total Likes of all tweets.
whatsapp_total_likes = whatsapp_filtered_df['Likes Counts'].sum()
#whatsapp_avg_likes = whatsapp_filtered_df['Likes Counts'].mean()
whatsapp_total_likes
#whatsapp_avg_likes
# Whatsapp Retweets Stats:
#whatsapp_sum_retweets = whatsapp_filtered_df['ReTweet Count'].sum()
whatsapp_avg_retweets = whatsapp_filtered_df['ReTweet Count'].mean()
#whatsapp_sum_retweets
whatsapp_avg_retweets
# # Charts and Plots
#
# #### Scatter plot - Twitter Average Followers to Tweets
# +
# Scatter Plot 1 - Tweets vs Average Followers vs Total Likes of the Top 10 Apps for both Google and Apple App Stores
fig, ax = plt.subplots(figsize=(11,11))
# Apps both on Google Play Store and Apple - 4 apps
facebook_plot = ax.scatter(facebook_total_tweets, facebook_avg_followers_ct, s=facebook_total_likes*15, color='sandybrown', label='Facebook', edgecolors='black', alpha=0.75)
instagram_plot= ax.scatter(instagram_total_tweets, instagram_avg_followers_ct, s=instagram_total_likes*15, color='saddlebrown', label='Instagram', edgecolors='black', alpha=0.5)
coc_plot= ax.scatter(coc_total_tweets, coc_avg_followers_ct, s=coc_total_likes*10, color='springgreen', label='Clash Of Clans', edgecolors='black', alpha=0.75)
candycrushsaga_plot= ax.scatter(candycrushsaga_total_tweets, candycrushsaga_avg_followers_ct, s=candycrushsaga_total_likes*5, color='limegreen', label='Candy Crush Saga', edgecolors='black')#, alpha=0.75)
# Google Play Store - 6 apps:
CleanMaster_plot= ax.scatter(CleanMaster_total_tweets, CleanMaster_avg_followers_ct, s=CleanMaster_total_likes*5, color='m', label='Clean Master Space Cleaner', edgecolors='black', alpha=0.75)
SubwaySurfers_plot= ax.scatter(SubwaySurfers_total_tweets, SubwaySurfers_avg_followers_ct, s=SubwaySurfers_total_likes*5, color='lime', label='Subway Surfers', edgecolors='black', alpha=0.75)
youtube_plot= ax.scatter(youtube_total_tweets, youtube_avg_followers_ct, s=youtube_total_likes*5, color='red', label='You Tube', edgecolors='black', alpha=0.75)
SecurityMaster_plot= ax.scatter(SecurityMaster_total_tweets, SecurityMaster_avg_followers_ct, s=SecurityMaster_total_likes*5, color='blueviolet', label='Security Master, Antivirus VPN', edgecolors='black', alpha=0.75)
ClashRoyale_plot= ax.scatter(ClashRoyale_total_tweets, ClashRoyale_avg_followers_ct, s=ClashRoyale_total_likes*5, color='darkolivegreen', label='Clash Royale', edgecolors='black', alpha=0.75)
whatsapp_plot= ax.scatter(whatsapp_total_tweets, whatsapp_avg_followers_ct, s=whatsapp_total_likes*5, color='tan', label='Whats App', edgecolors='black', alpha=0.75)
# Apple Apps Store - 6 apps
templerun_plot= ax.scatter(templerun_total_tweets, templerun_avg_followers_ct, s=templerun_total_likes*5, color='lawngreen', label='Temple Run', edgecolors='black', alpha=0.75)
pandora_plot= ax.scatter(pandora_total_tweets, pandora_avg_followers_ct, s=pandora_total_likes*5, color='coral', label='Pandora', edgecolors='black', alpha=0.75)
pinterest_plot= ax.scatter(pinterest_total_tweets, pinterest_avg_followers_ct, s=pinterest_total_likes*5, color='firebrick', label='Pinterest', edgecolors='black', alpha=0.75)
bible_plot= ax.scatter(bible_total_tweets, bible_avg_followers_ct, s=bible_total_likes*5, color='tomato', label='Bible', edgecolors='black', alpha=0.75)
spotify_plot= ax.scatter(spotify_total_tweets, spotify_avg_followers_ct, s=spotify_total_likes*5, color='orangered', label='Spotify', edgecolors='black', alpha=0.75)
angrybirds_plot= ax.scatter(angrybirds_total_tweets, angrybirds_avg_followers_ct, s=angrybirds_total_likes*5, color='forestgreen', label='Angry Birds', edgecolors='black', alpha=0.75)
# title and labels
plt.title("Tweets vs Average Followers (Mar 27 - Apr 3, 2019) \n")
plt.xlabel("Total Tweets \n Note: Circle sizes correlate with Total Likes" )
plt.ylabel("Average Number of Followers per Twitter User \n")
# set and format the legend
lgnd = plt.legend(title='Legend', loc="best")
lgnd.legendHandles[0]._sizes = [30]
lgnd.legendHandles[1]._sizes = [30]
lgnd.legendHandles[2]._sizes = [30]
lgnd.legendHandles[3]._sizes = [30]
lgnd.legendHandles[4]._sizes = [30]
lgnd.legendHandles[5]._sizes = [30]
lgnd.legendHandles[6]._sizes = [30]
lgnd.legendHandles[7]._sizes = [30]
lgnd.legendHandles[8]._sizes = [30]
lgnd.legendHandles[9]._sizes = [30]
lgnd.legendHandles[10]._sizes = [30]
lgnd.legendHandles[11]._sizes = [30]
lgnd.legendHandles[12]._sizes = [30]
lgnd.legendHandles[13]._sizes = [30]
lgnd.legendHandles[14]._sizes = [30]
lgnd.legendHandles[15]._sizes = [30]
#grid lines and show
plt.grid()
plt.show()
#plt.savefig("./TWEETS_vs__AVG_followers_Scatter.png")
# +
# Test Cell: Tried to automate plot, but was unable to beause for the size (s), JG wanted to scale the
# the size by multiplying by a unique scale depending on the number of likes to emphasize data points
# Conclusion: was to stick with brute force method
SubwaySurfers_total_tweets,
x = [facebook_total_tweets, instagram_total_tweets, coc_total_tweets,
candycrushsaga_total_tweets, CleanMaster_total_tweets,
youtube_total_tweets, SecurityMaster_total_tweets,
ClashRoyale_total_tweets, whatsapp_total_tweets, templerun_total_tweets,
pandora_total_tweets, pinterest_total_tweets, bible_total_tweets, spotify_total_tweets,
angrybirds_total_tweets]
SubwaySurfers_avg_followers_ct,
y = [facebook_avg_followers_ct, instagram_avg_followers_ct, coc_avg_followers_ct,
candycrushsaga_avg_followers_ct, CleanMaster_avg_followers_ct,
youtube_avg_followers_ct, SecurityMaster_avg_followers_ct,
ClashRoyale_avg_followers_ct, whatsapp_avg_followers_ct, templerun_avg_followers_ct,
pandora_avg_followers_ct, pinterest_avg_followers_ct, bible_avg_followers_ct, spotify_avg_followers_ct,
angrybirds_avg_followers_ct]
"""
# Below this method doesn't work. Will go with brute force method.
s = [(facebook_total_likes*15), (instagram_total_likes*15), (coc_total_likes*10), (candycrushsaga_total_likes*5),
(CleanMaster_total_likes*5), (SubwaySurfers_total_likes*5), (youtube_total_likes*5), (SecurityMaster_total_likes*5)
(ClashRoyale_total_likes*5), (whatsapp_total_likes*5), (templerun_total_likes*5), (pandora_total_likes*5),
(pinterest_total_likes*5), (bible_total_likes*5), (spotify_total_likes*5), (angrybirds_total_likes*5)]
"""
s = [facebook_total_likes, instagram_total_likes, coc_total_likes, candycrushsaga_total_likes,
CleanMaster_total_likes, SubwaySurfers_total_likes, youtube_total_likes, SecurityMaster_total_likes,
ClashRoyale_total_likes, whatsapp_total_likes, templerun_total_likes, pandora_total_likes,
pinterest_total_likes, bible_total_likes, spotify_total_likes, angrybirds_total_likes]
colors = np.random.rand(16)
label = []
edgecolors = []
alpha = []
fig, ax = plt.subplots(figsize=(11,11))
ax.scatter(x, y, s)
plt.grid()
plt.show()
"""# Apps both on Google Play Store and Apple - 4 apps
facebook_plot = ax.scatter(, , , color='sandybrown', label='Facebook', edgecolors='black', alpha=0.75)
instagram_plot= ax.scatter(, , , color='saddlebrown', label='Instagram', edgecolors='black', alpha=0.5)
coc_plot= ax.scatter(,, , color='springgreen', label='Clash Of Clans', edgecolors='black', alpha=0.75)
candycrushsaga_plot= ax.scatter(,, , color='limegreen', label='Candy Crush Saga', edgecolors='black')#, alpha=0.75)
# Google Play Store - 6 apps:
CleanMaster_plot= ax.scatter(,, , color='m', label='Clean Master Space Cleaner', edgecolors='black', alpha=0.75)
SubwaySurfers_plot= ax.scatter(,, , color='lime', label='Subway Surfers', edgecolors='black', alpha=0.75)
youtube_plot= ax.scatter(,, , color='red', label='You Tube', edgecolors='black', alpha=0.75)
SecurityMaster_plot= ax.scatter(,, , color='blueviolet', label='Security Master, Antivirus VPN', edgecolors='black', alpha=0.75)
ClashRoyale_plot= ax.scatter(,, , color='darkolivegreen', label='Clash Royale', edgecolors='black', alpha=0.75)
whatsapp_plot= ax.scatter(,, , color='tan', label='Whats App', edgecolors='black', alpha=0.75)
# Apple Apps Store - 6 apps
templerun_plot= ax.scatter(, , , color='lawngreen', label='Temple Run', edgecolors='black', alpha=0.75)
pandora_plot= ax.scatter(, ,, color='coral', label='Pandora', edgecolors='black', alpha=0.75)
pinterest_plot= ax.scatter(, ,, color='firebrick', label='Pinterest', edgecolors='black', alpha=0.75)
bible_plot= ax.scatter(, , color='tomato', label='Bible', edgecolors='black', alpha=0.75)
spotify_plot= ax.scatter(, , color='orangered', label='Spotify', edgecolors='black', alpha=0.75)
angrybirds_plot= ax.scatter(,,, color='forestgreen', label='Angry Birds', edgecolors='black', alpha=0.75)
# title and labels
plt.title("Tweets vs Average Followers (Mar 27 - Apr 3, 2019) \n")
plt.xlabel("Total Tweets \n Note: Circle sizes correlate with Total Likes" )
plt.ylabel("Average Number of Followers per Twitter User \n")
"""
# +
# Scatter Plot 2 - Tweets vs ReTweets vs Likes
fig, ax = plt.subplots(figsize=(11,11))
# Apps both on Google Play Store and Apple - 4 apps
facebook_plot = ax.scatter(facebook_total_tweets, facebook_avg_retweets, s=facebook_total_likes*5, color='sandybrown', label='Facebook', edgecolors='black', alpha=0.75)
instagram_plot= ax.scatter(instagram_total_tweets, instagram_avg_retweets, s=instagram_total_likes*5, color='saddlebrown', label='Instagram', edgecolors='black', alpha=0.75)
coc_plot= ax.scatter(coc_total_tweets, coc_avg_retweets, s=coc_total_likes*5, color='springgreen', label='Clash Of Clans', edgecolors='black', alpha=0.75)
candycrushsaga_plot= ax.scatter(candycrushsaga_total_tweets, candycrushsaga_avg_retweets, s=candycrushsaga_total_likes*5, color='limegreen', label='Candy Crush Saga', edgecolors='black')#, alpha=0.75)
# Google Play Store - 6 apps:
CleanMaster_plot= ax.scatter(CleanMaster_total_tweets, CleanMaster_avg_retweets, s=CleanMaster_total_likes*5, color='m', label='Clean Master Space Cleaner', edgecolors='black', alpha=0.75)
SubwaySurfers_plot= ax.scatter(SubwaySurfers_total_tweets, SubwaySurfers_avg_retweets, s=SubwaySurfers_total_likes*5, color='lime', label='Subway Surfers', edgecolors='black', alpha=0.75)
youtube_plot= ax.scatter(youtube_total_tweets, youtube_avg_retweets, s=youtube_total_likes*5, color='red', label='You Tube', edgecolors='black', alpha=0.75)
SecurityMaster_plot= ax.scatter(SecurityMaster_total_tweets, SecurityMaster_avg_retweets, s=SecurityMaster_total_likes*5, color='blueviolet', label='Security Master, Antivirus VPN', edgecolors='black', alpha=0.75)
ClashRoyale_plot= ax.scatter(ClashRoyale_total_tweets, ClashRoyale_avg_retweets, s=ClashRoyale_total_likes*5, color='darkolivegreen', label='Clash Royale', edgecolors='black', alpha=0.75)
whatsapp_plot= ax.scatter(whatsapp_total_tweets, whatsapp_avg_retweets, s=whatsapp_total_likes*5, color='tan', label='Whats App', edgecolors='black', alpha=0.75)
# Apple Apps Store - 6 apps
templerun_plot= ax.scatter(templerun_total_tweets, templerun_avg_retweets, s=templerun_total_likes*5, color='lawngreen', label='Temple Run', edgecolors='black', alpha=0.75)
pandora_plot= ax.scatter(pandora_total_tweets, pandora_avg_retweets, s=pandora_total_likes*5, color='coral', label='Pandora', edgecolors='black', alpha=0.75)
pinterest_plot= ax.scatter(pinterest_total_tweets, pinterest_avg_retweets, s=pinterest_total_likes*5, color='firebrick', label='Pinterest', edgecolors='black', alpha=0.75)
bible_plot= ax.scatter(bible_total_tweets, bible_avg_retweets, s=bible_total_likes*5, color='tomato', label='Bible', edgecolors='black', alpha=0.75)
spotify_plot= ax.scatter(spotify_total_tweets, spotify_avg_retweets, s=spotify_total_likes*5, color='orangered', label='Spotify', edgecolors='black', alpha=0.75)
angrybirds_plot= ax.scatter(angrybirds_total_tweets, angrybirds_avg_retweets, s=angrybirds_total_likes*5, color='forestgreen', label='Angry Birds', edgecolors='black', alpha=0.75)
# title and labels
plt.title("Tweets vs ReTweets (Mar 27 - Apr 3, 2019) \n")
plt.xlabel("Total Tweets \n Note: Circle sizes correlate with Total Likes \n" )
plt.ylabel("Average Number of ReTweets per Twitter User \n")
# set and format the legend
lgnd = plt.legend(title='Legend', loc="best")
lgnd.legendHandles[0]._sizes = [30]
lgnd.legendHandles[1]._sizes = [30]
lgnd.legendHandles[2]._sizes = [30]
lgnd.legendHandles[3]._sizes = [30]
lgnd.legendHandles[4]._sizes = [30]
lgnd.legendHandles[5]._sizes = [30]
lgnd.legendHandles[6]._sizes = [30]
lgnd.legendHandles[7]._sizes = [30]
lgnd.legendHandles[8]._sizes = [30]
lgnd.legendHandles[9]._sizes = [30]
lgnd.legendHandles[10]._sizes = [30]
lgnd.legendHandles[11]._sizes = [30]
lgnd.legendHandles[12]._sizes = [30]
lgnd.legendHandles[13]._sizes = [30]
lgnd.legendHandles[14]._sizes = [30]
lgnd.legendHandles[15]._sizes = [30]
#grid lines and show
plt.grid()
plt.show()
#plt.savefig('./TWEETS_VS_RETWEETS_vs_LIKES_Scatter.png')
# +
# Scatter Plot 3 - Will not use this plot
fig, ax = plt.subplots(figsize=(8,8))
# Apps both on Google Play Store and Apple - 4 apps
facebook_plot = ax.scatter(facebook_avg_retweets, facebook_total_tweets, s=facebook_total_likes*5, color='blue', label='Facebook', edgecolors='red', alpha=0.75)
instagram_plot= ax.scatter(instagram_avg_retweets, instagram_total_tweets, s=instagram_total_likes*5, color='fuchsia', label='Instagram', edgecolors='red', alpha=0.75)
coc_plot= ax.scatter(coc_avg_retweets, coc_total_tweets, s=coc_total_likes*5, color='springgreen', label='Clash Of Clans', edgecolors='red', alpha=0.75)
candycrushsaga_plot= ax.scatter(candycrushsaga_avg_retweets, candycrushsaga_total_tweets, s=candycrushsaga_total_likes*5, color='black', label='Candy Crush Saga', edgecolors='red')#, alpha=0.75)
# Google Play Store - 6 apps:
CleanMaster_plot= ax.scatter(CleanMaster_avg_retweets, CleanMaster_total_tweets, s=CleanMaster_total_likes*5, color='olive', label='Clean Master Space Cleaner', edgecolors='lime', alpha=0.75)
SubwaySurfers_plot= ax.scatter(SubwaySurfers_avg_retweets, SubwaySurfers_total_tweets, s=SubwaySurfers_total_likes*5, color='plum', label='Subway Surfers', edgecolors='lime', alpha=0.75)
youtube_plot= ax.scatter(youtube_avg_retweets, youtube_total_tweets, s=youtube_total_likes*5, color='grey', label='You Tube', edgecolors='lime', alpha=0.75)
SecurityMaster_plot= ax.scatter(SecurityMaster_avg_retweets, SecurityMaster_total_tweets, s=SecurityMaster_total_likes*5, color='coral', label='Security Master, Antivirus VPN', edgecolors='lime', alpha=0.75)
ClashRoyale_plot= ax.scatter(ClashRoyale_avg_retweets, ClashRoyale_total_tweets, s=ClashRoyale_total_likes*5, color='orange', label='Clash Royale', edgecolors='lime', alpha=0.75)
whatsapp_plot= ax.scatter(whatsapp_avg_retweets, whatsapp_total_tweets, s=whatsapp_total_likes*5, color='green', label='Whats App', edgecolors='lime', alpha=0.75)
# Apple Apps Store - 6 apps
templerun_plot= ax.scatter(templerun_avg_retweets, templerun_total_tweets, s=templerun_total_likes*5, color='lawngreen', label='Temple Run', edgecolors='black', alpha=0.75)
pandora_plot= ax.scatter(pandora_avg_retweets, pandora_total_tweets, s=pandora_total_likes*5, color='cornflowerblue', label='Pandora', edgecolors='black', alpha=0.75)
pinterest_plot= ax.scatter(pinterest_avg_retweets, pinterest_total_tweets, s=pinterest_total_likes*5, color='firebrick', label='Pinterest', edgecolors='black', alpha=0.75)
bible_plot= ax.scatter(bible_avg_retweets, bible_total_tweets, s=bible_total_likes*5, color='brown', label='Bible', edgecolors='black', alpha=0.75)
spotify_plot= ax.scatter(spotify_avg_retweets, spotify_total_tweets, s=spotify_total_likes*5, color='darkgreen', label='Spotify', edgecolors='black', alpha=0.75)
angrybirds_plot= ax.scatter(angrybirds_avg_retweets, angrybirds_total_tweets, s=angrybirds_total_likes*5, color='salmon', label='Angry Birds', edgecolors='black', alpha=0.75)
# title and labels
plt.title("Tweets vs ReTweets (Mar 27 - Apr 3, 2019) \n")
plt.xlabel("Total Tweets \n Note: Circle sizes correlate with Total Likes \n" )
plt.ylabel("Average Number of ReTweets per Twitter User \n")
# set and format the legend
lgnd = plt.legend(title='Legend', loc="best")
lgnd.legendHandles[0]._sizes = [30]
lgnd.legendHandles[1]._sizes = [30]
lgnd.legendHandles[2]._sizes = [30]
lgnd.legendHandles[3]._sizes = [30]
lgnd.legendHandles[4]._sizes = [30]
lgnd.legendHandles[5]._sizes = [30]
lgnd.legendHandles[6]._sizes = [30]
lgnd.legendHandles[7]._sizes = [30]
lgnd.legendHandles[8]._sizes = [30]
lgnd.legendHandles[9]._sizes = [30]
lgnd.legendHandles[10]._sizes = [30]
lgnd.legendHandles[11]._sizes = [30]
lgnd.legendHandles[12]._sizes = [30]
lgnd.legendHandles[13]._sizes = [30]
lgnd.legendHandles[14]._sizes = [30]
lgnd.legendHandles[15]._sizes = [30]
#grid lines and show
plt.grid()
plt.show()
#plt.savefig('./tweets_vs__avgfollowers_Scatter.png')
# +
# Hardcoding numbers from analysis done in Apple and Google Play Store Final Code Notebooks
# Avergage Apple, Google Ratings
facebook_avg_rating = (3.5 + 4.1)/2
instagram_avg_rating = (4.5 + 4.5)/2
coc_avg_rating = (4.5 + 4.6)/2
candycrushsaga_avg_rating = (4.5 + 4.4)/2
# Avergage Apple, Google Reviews
facebook_reviews = (2974676 + 78158306)/2
instagram_reviews = (2161558 + 66577446)/2
coc_reviews = (2130805 + 44893888)/2
candycrushsaga_reviews = (961794 + 22430188)/2
# Apple App Ratings
templerun_rating = 4.5
pandora_rating = 4.5
pinterest_rating = 4.5
bible_rating = 4.5
spotify_rating = 4.5
angrybirds_rating = 4.5
# Apple App Reviews
templerun_reviews = 1724546
pandora_reviews = 1126879
pinterest_reviews = 1061624
bible_reviews = 985920
spotify_reviews = 878563
angrybirds_reviews = 824451
# Google App Ratings
whatsapp_rating = 4.4
clean_master_rating = 4.7
subway_surfers_rating = 4.5
you_tube_rating = 4.3
security_master_rating = 4.7
clash_royale_rating = 4.6
# Google App Reviews
whatsapp_reviews = 69119316
clean_master_reviews = 42916526
subway_surfers_reviews = 27725352
you_tube_reviews = 25655305
security_master_reviews = 24900999
clash_royale_reviews = 23136735
# +
# Scatter Plot 5 - Tweets vs Ratings vs Likes - USE THIS ONE
fig, ax = plt.subplots(figsize=(11,11))
# Apps both on Google Play Store and Apple - 4 apps
facebook_plot = ax.scatter(facebook_total_tweets, facebook_avg_rating, s=facebook_total_likes*5, color='sandybrown', label='Facebook', edgecolors='black', alpha=0.75)
instagram_plot= ax.scatter(instagram_total_tweets, instagram_avg_rating, s=instagram_total_likes*5, color='saddlebrown', label='Instagram', edgecolors='black', alpha=0.75)
coc_plot= ax.scatter(coc_total_tweets, coc_avg_rating, s=coc_total_likes*5, color='springgreen', label='Clash Of Clans', edgecolors='black', alpha=0.75)
candycrushsaga_plot= ax.scatter(candycrushsaga_total_tweets, candycrushsaga_avg_rating, s=candycrushsaga_total_likes*5, color='limegreen', label='Candy Crush Saga', edgecolors='black')#, alpha=0.75)
# Google Play Store - 6 apps:
CleanMaster_plot= ax.scatter(CleanMaster_total_tweets, clean_master_rating, s=CleanMaster_total_likes*5, color='m', label='Clean Master Space Cleaner', edgecolors='black', alpha=0.75)
SubwaySurfers_plot= ax.scatter(SubwaySurfers_total_tweets, subway_surfers_rating, s=SubwaySurfers_total_likes*5, color='lime', label='Subway Surfers', edgecolors='black', alpha=0.75)
youtube_plot= ax.scatter(youtube_total_tweets, you_tube_rating, s=youtube_total_likes*5, color='red', label='You Tube', edgecolors='black', alpha=0.75)
SecurityMaster_plot= ax.scatter(SecurityMaster_total_tweets, security_master_rating, s=SecurityMaster_total_likes*5, color='blueviolet', label='Security Master, Antivirus VPN', edgecolors='black', alpha=0.75)
ClashRoyale_plot= ax.scatter(ClashRoyale_total_tweets, clash_royale_rating, s=ClashRoyale_total_likes*5, color='darkolivegreen', label='Clash Royale', edgecolors='black', alpha=0.75)
whatsapp_plot= ax.scatter(whatsapp_total_tweets, whatsapp_rating, s=whatsapp_total_likes*5, color='tan', label='Whats App', edgecolors='black', alpha=0.75)
# Apple Apps Store - 6 apps
templerun_plot= ax.scatter(templerun_total_tweets,templerun_rating, s=templerun_total_likes*5, color='lawngreen', label='Temple Run', edgecolors='black', alpha=0.75)
pandora_plot= ax.scatter(pandora_total_tweets, pandora_rating, s=pandora_total_likes*5, color='coral', label='Pandora', edgecolors='black', alpha=0.75)
pinterest_plot= ax.scatter(pinterest_total_tweets, pinterest_rating, s=pinterest_total_likes*5, color='firebrick', label='Pinterest', edgecolors='black', alpha=0.75)
bible_plot= ax.scatter(bible_total_tweets, bible_rating, s=bible_total_likes*5, color='tomato', label='Bible', edgecolors='black', alpha=0.75)
spotify_plot= ax.scatter(spotify_total_tweets, spotify_rating, s=spotify_total_likes*5, color='orangered', label='Spotify', edgecolors='black', alpha=0.75)
angrybirds_plot= ax.scatter(angrybirds_total_tweets, angrybirds_rating, s=angrybirds_total_likes*5, color='forestgreen', label='<NAME>', edgecolors='black', alpha=0.75)
# title and labels
plt.title("Tweets vs Ratings (Mar 27 - Apr 3, 2019) \n")
plt.xlabel("Total Tweets \n Note: Circle sizes correlate with Total Likes \n" )
plt.ylabel("App Store User Ratings (Out of 5) \n")
# set and format the legend
lgnd = plt.legend(title='Legend', loc="best")
lgnd.legendHandles[0]._sizes = [30]
lgnd.legendHandles[1]._sizes = [30]
lgnd.legendHandles[2]._sizes = [30]
lgnd.legendHandles[3]._sizes = [30]
lgnd.legendHandles[4]._sizes = [30]
lgnd.legendHandles[5]._sizes = [30]
lgnd.legendHandles[6]._sizes = [30]
lgnd.legendHandles[7]._sizes = [30]
lgnd.legendHandles[8]._sizes = [30]
lgnd.legendHandles[9]._sizes = [30]
lgnd.legendHandles[10]._sizes = [30]
lgnd.legendHandles[11]._sizes = [30]
lgnd.legendHandles[12]._sizes = [30]
lgnd.legendHandles[13]._sizes = [30]
lgnd.legendHandles[14]._sizes = [30]
lgnd.legendHandles[15]._sizes = [30]
#grid lines and show
plt.grid()
plt.show()
#plt.savefig('./TWEETS_VS_RATINGSVS LIKES_Scatter.png')
# +
# Scatter Plot 5 - Tweets vs Reviews vs Ratings (size) - DO NOT USE
fig, ax = plt.subplots(figsize=(11,11))
# Apps both on Google Play Store and Apple - 4 apps
facebook_plot = ax.scatter(facebook_total_tweets, facebook_reviews, s=facebook_avg_rating*105, color='sandybrown', label='Facebook', edgecolors='black', alpha=0.75)
instagram_plot= ax.scatter(instagram_total_tweets, instagram_reviews, s=instagram_avg_rating*105, color='saddlebrown', label='Instagram', edgecolors='black', alpha=0.75)
coc_plot= ax.scatter(coc_total_tweets, coc_reviews, s=coc_avg_rating*105, color='springgreen', label='Clash Of Clans', edgecolors='black', alpha=0.75)
candycrushsaga_plot= ax.scatter(candycrushsaga_total_tweets, candycrushsaga_reviews, s=candycrushsaga_avg_rating*105, color='limegreen', label='Candy Crush Saga', edgecolors='black', alpha=0.75)
# Google Play Store - 6 apps:
CleanMaster_plot= ax.scatter(CleanMaster_total_tweets, clean_master_reviews, s=clean_master_rating*105, color='m', label='Clean Master Space Cleaner', edgecolors='black', alpha=0.75)
SubwaySurfers_plot= ax.scatter(SubwaySurfers_total_tweets, subway_surfers_reviews, s=subway_surfers_rating*105, color='lime', label='Subway Surfers', edgecolors='black', alpha=0.75)
youtube_plot= ax.scatter(youtube_total_tweets, you_tube_reviews, s=you_tube_rating*105, color='red', label='You Tube', edgecolors='black', alpha=0.75)
SecurityMaster_plot= ax.scatter(SecurityMaster_total_tweets, security_master_reviews, s=security_master_rating*105, color='blueviolet', label='Security Master, Antivirus VPN', edgecolors='black', alpha=0.75)
ClashRoyale_plot= ax.scatter(ClashRoyale_total_tweets, clash_royale_reviews, s=clash_royale_rating*105, color='darkolivegreen', label='Clash Royale', edgecolors='black', alpha=0.75)
whatsapp_plot= ax.scatter(whatsapp_total_tweets, whatsapp_reviews, s=whatsapp_rating*105, color='tan', label='Whats App', edgecolors='lime', alpha=0.75)
# Apple Apps Store - 6 apps
templerun_plot= ax.scatter(templerun_total_tweets,templerun_reviews, s=templerun_rating*105, color='lawngreen', label='Temple Run', edgecolors='black', alpha=0.75)
pandora_plot= ax.scatter(pandora_total_tweets, pandora_reviews, s=pandora_rating*105, color='coral', label='Pandora', edgecolors='black', alpha=0.75)
pinterest_plot= ax.scatter(pinterest_total_tweets, pinterest_reviews, s=pinterest_rating*105, color='firebrick', label='Pinterest', edgecolors='black', alpha=0.75)
bible_plot= ax.scatter(bible_total_tweets, bible_reviews, s=bible_rating*105, color='tomato', label='Bible', edgecolors='black', alpha=0.75)
spotify_plot= ax.scatter(spotify_total_tweets, spotify_reviews, s=spotify_rating*105, color='orangered', label='Spotify', edgecolors='black', alpha=0.75)
angrybirds_plot= ax.scatter(angrybirds_total_tweets, angrybirds_reviews, s=angrybirds_rating*105, color='forestgreen', label='Angry Birds', edgecolors='black', alpha=0.75)
# title and labels
plt.title("Tweets vs Reviews (Mar 27 - Apr 3, 2019) \n")
plt.xlabel("Total Tweets \n Note: Circle sizes correlate with App Ratings \n" )
plt.ylabel("App Store Reviews in Millions \n")
# set and format the legend
lgnd = plt.legend(title='Legend', loc="best")
lgnd.legendHandles[0]._sizes = [30]
lgnd.legendHandles[1]._sizes = [30]
lgnd.legendHandles[2]._sizes = [30]
lgnd.legendHandles[3]._sizes = [30]
lgnd.legendHandles[4]._sizes = [30]
lgnd.legendHandles[5]._sizes = [30]
lgnd.legendHandles[6]._sizes = [30]
lgnd.legendHandles[7]._sizes = [30]
lgnd.legendHandles[8]._sizes = [30]
lgnd.legendHandles[9]._sizes = [30]
lgnd.legendHandles[10]._sizes = [30]
lgnd.legendHandles[11]._sizes = [30]
lgnd.legendHandles[12]._sizes = [30]
lgnd.legendHandles[13]._sizes = [30]
lgnd.legendHandles[14]._sizes = [30]
lgnd.legendHandles[15]._sizes = [30]
#grid lines and show
plt.grid()
plt.show()
#plt.savefig('./tweets_vs__avgfollowers_Scatter.png')
# +
# Scatter Plot 6 - Tweets vs Reviews vs Likes (size) -USE THIS ONE
fig, ax = plt.subplots(figsize=(11,11))
# Apps both on Google Play Store and Apple - 4 apps
facebook_plot = ax.scatter(facebook_total_tweets, facebook_reviews, s=facebook_total_likes*5, color='sandybrown', label='Facebook', edgecolors='black', alpha=0.75)
instagram_plot= ax.scatter(instagram_total_tweets, instagram_reviews, s=instagram_total_likes*5, color='saddlebrown', label='Instagram', edgecolors='black', alpha=0.75)
coc_plot= ax.scatter(coc_total_tweets, coc_reviews, s=coc_total_likes*5, color='springgreen', label='Clash Of Clans', edgecolors='black', alpha=0.75)
candycrushsaga_plot= ax.scatter(candycrushsaga_total_tweets, candycrushsaga_reviews, s=candycrushsaga_total_likes*5, color='limegreen', label='Candy Crush Saga', edgecolors='black', alpha=0.75)
# Google Play Store - 6 apps:
CleanMaster_plot= ax.scatter(CleanMaster_total_tweets, clean_master_reviews, s=CleanMaster_total_likes*5, color='m', label='Clean Master Space Cleaner', edgecolors='black', alpha=0.75)
SubwaySurfers_plot= ax.scatter(SubwaySurfers_total_tweets, subway_surfers_reviews, s=SubwaySurfers_total_likes*5, color='lime', label='Subway Surfers', edgecolors='black', alpha=0.75)
youtube_plot= ax.scatter(youtube_total_tweets, you_tube_reviews, s=youtube_total_likes*5, color='red', label='You Tube', edgecolors='black', alpha=0.75)
SecurityMaster_plot= ax.scatter(SecurityMaster_total_tweets, security_master_reviews, s=SecurityMaster_total_likes*5, color='blueviolet', label='Security Master, Antivirus VPN', edgecolors='black', alpha=0.75)
ClashRoyale_plot= ax.scatter(ClashRoyale_total_tweets, clash_royale_reviews, s=ClashRoyale_total_likes*5, color='darkolivegreen', label='Clash Royale', edgecolors='black', alpha=0.75)
whatsapp_plot= ax.scatter(whatsapp_total_tweets, whatsapp_reviews, s=whatsapp_total_likes*5, color='tan', label='Whats App', edgecolors='black', alpha=0.75)
# Apple Apps Store - 6 apps
templerun_plot= ax.scatter(templerun_total_tweets, templerun_reviews, s=templerun_total_likes*5, color='lawngreen', label='Temple Run', edgecolors='black', alpha=0.75)
pandora_plot= ax.scatter(pandora_total_tweets, pandora_reviews, s=pandora_total_likes*5, color='coral', label='Pandora', edgecolors='black', alpha=0.75)
pinterest_plot= ax.scatter(pinterest_total_tweets, pinterest_reviews, s=pinterest_total_likes*5, color='firebrick', label='Pinterest', edgecolors='black', alpha=0.75)
bible_plot= ax.scatter(bible_total_tweets, bible_reviews, s=bible_total_likes*5, color='tomato', label='Bible', edgecolors='black', alpha=0.75)
spotify_plot= ax.scatter(spotify_total_tweets, spotify_reviews, s=spotify_total_likes*5, color='orangered', label='Spotify', edgecolors='black', alpha=0.75)
angrybirds_plot= ax.scatter(angrybirds_total_tweets, angrybirds_reviews, s=angrybirds_total_likes*5, color='forestgreen', label='Angry Birds', edgecolors='black', alpha=0.75)
# title and labels
plt.title("Tweets vs Reviews (Mar 27 - Apr 3, 2019) \n")
plt.xlabel("Total Tweets \n Note: Circle sizes correlate with Likes \n" )
plt.ylabel("App Store Reviews in Millions \n")
# set and format the legend
lgnd = plt.legend(title='Legend', loc="best")
lgnd.legendHandles[0]._sizes = [30]
lgnd.legendHandles[1]._sizes = [30]
lgnd.legendHandles[2]._sizes = [30]
lgnd.legendHandles[3]._sizes = [30]
lgnd.legendHandles[4]._sizes = [30]
lgnd.legendHandles[5]._sizes = [30]
lgnd.legendHandles[6]._sizes = [30]
lgnd.legendHandles[7]._sizes = [30]
lgnd.legendHandles[8]._sizes = [30]
lgnd.legendHandles[9]._sizes = [30]
lgnd.legendHandles[10]._sizes = [30]
lgnd.legendHandles[11]._sizes = [30]
lgnd.legendHandles[12]._sizes = [30]
lgnd.legendHandles[13]._sizes = [30]
lgnd.legendHandles[14]._sizes = [30]
lgnd.legendHandles[15]._sizes = [30]
#grid lines and show
plt.grid()
plt.show()
#plt.savefig('./TWEETS_VS_REVIEWS_VSLIKES_Scatter.png')
# +
# Scatter Plot 5 - Tweets vs Reviews vs Likes (size) - Need to do
fig, ax = plt.subplots(figsize=(8,8))
# Apps both on Google Play Store and Apple - 4 apps
facebook_plot = ax.scatter(facebook_avg_retweets, facebook_total_tweets, s=facebook_total_likes*5, color='blue', label='Facebook', edgecolors='red', alpha=0.75)
instagram_plot= ax.scatter(instagram_avg_retweets, instagram_total_tweets, s=instagram_total_likes*5, color='fuchsia', label='Instagram', edgecolors='red', alpha=0.75)
coc_plot= ax.scatter(coc_avg_retweets, coc_total_tweets, s=coc_total_likes*5, color='springgreen', label='Clash Of Clans', edgecolors='red', alpha=0.75)
candycrushsaga_plot= ax.scatter(candycrushsaga_avg_retweets, candycrushsaga_total_tweets, s=candycrushsaga_total_likes*5, color='black', label='Candy Crush Saga', edgecolors='red')#, alpha=0.75)
# Google Play Store - 6 apps:
CleanMaster_plot= ax.scatter(CleanMaster_avg_retweets, CleanMaster_total_tweets, s=CleanMaster_total_likes*5, color='olive', label='Clean Master Space Cleaner', edgecolors='lime', alpha=0.75)
SubwaySurfers_plot= ax.scatter(SubwaySurfers_avg_retweets, SubwaySurfers_total_tweets, s=SubwaySurfers_total_likes*5, color='plum', label='Subway Surfers', edgecolors='lime', alpha=0.75)
youtube_plot= ax.scatter(youtube_avg_retweets, youtube_total_tweets, s=youtube_total_likes*5, color='grey', label='You Tube', edgecolors='lime', alpha=0.75)
SecurityMaster_plot= ax.scatter(SecurityMaster_avg_retweets, SecurityMaster_total_tweets, s=SecurityMaster_total_likes*5, color='coral', label='Security Master, Antivirus VPN', edgecolors='lime', alpha=0.75)
ClashRoyale_plot= ax.scatter(ClashRoyale_avg_retweets, ClashRoyale_total_tweets, s=ClashRoyale_total_likes*5, color='orange', label='Clash Royale', edgecolors='lime', alpha=0.75)
whatsapp_plot= ax.scatter(whatsapp_avg_retweets, whatsapp_total_tweets, s=whatsapp_total_likes*5, color='green', label='Whats App', edgecolors='lime', alpha=0.75)
# Apple Apps Store - 6 apps
templerun_plot= ax.scatter(templerun_avg_retweets, templerun_total_tweets, s=templerun_total_likes*5, color='lawngreen', label='Temple Run', edgecolors='black', alpha=0.75)
pandora_plot= ax.scatter(pandora_avg_retweets, pandora_total_tweets, s=pandora_total_likes*5, color='cornflowerblue', label='Pandora', edgecolors='black', alpha=0.75)
pinterest_plot= ax.scatter(pinterest_avg_retweets, pinterest_total_tweets, s=pinterest_total_likes*5, color='firebrick', label='Pinterest', edgecolors='black', alpha=0.75)
bible_plot= ax.scatter(bible_avg_retweets, bible_total_tweets, s=bible_total_likes*5, color='brown', label='Bible', edgecolors='black', alpha=0.75)
spotify_plot= ax.scatter(spotify_avg_retweets, spotify_total_tweets, s=spotify_total_likes*5, color='darkgreen', label='Spotify', edgecolors='black', alpha=0.75)
angrybirds_plot= ax.scatter(angrybirds_avg_retweets, angrybirds_total_tweets, s=angrybirds_total_likes*5, color='salmon', label='Angry Birds', edgecolors='black', alpha=0.75)
# title and labels
plt.title("Tweets vs ReTweets (Mar 27 - Apr 3, 2019) \n")
plt.xlabel("Avg ReTweets \n Note: Circle sizes correlate with Total Likes \n" )
plt.ylabel("Total Tweets \n")
# set and format the legend
lgnd = plt.legend(title='Legend', loc="best")
lgnd.legendHandles[0]._sizes = [30]
lgnd.legendHandles[1]._sizes = [30]
lgnd.legendHandles[2]._sizes = [30]
lgnd.legendHandles[3]._sizes = [30]
lgnd.legendHandles[4]._sizes = [30]
lgnd.legendHandles[5]._sizes = [30]
lgnd.legendHandles[6]._sizes = [30]
lgnd.legendHandles[7]._sizes = [30]
lgnd.legendHandles[8]._sizes = [30]
lgnd.legendHandles[9]._sizes = [30]
lgnd.legendHandles[10]._sizes = [30]
lgnd.legendHandles[11]._sizes = [30]
lgnd.legendHandles[12]._sizes = [30]
lgnd.legendHandles[13]._sizes = [30]
lgnd.legendHandles[14]._sizes = [30]
lgnd.legendHandles[15]._sizes = [30]
#grid lines and show
plt.grid()
plt.show()
#plt.savefig('./tweets_vs__avgfollowers_Scatter.png')
| archive/1_twitter_mining_func_scatter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# # Fields Far Field
#
# +
# sphinx_gallery_thumbnail_path = '../images/Fields:FarField.png'
def run(Plot, Save):
from PyMieSim.Scatterer import Sphere
from PyMieSim.Source import PlaneWave
Source = PlaneWave(Wavelength = 450e-9,
Polarization = 0,
E0 = 1)
Scat = Sphere(Diameter = 300e-9,
Source = Source,
Index = 1.4)
Fields = Scat.FarField(Num=100)
if Plot:
Fields.Plot()
if Save:
from pathlib import Path
dir = f'docs/images/{Path(__file__).stem}'
Fields.SaveFig(Directory=dir)
if __name__ == '__main__':
run(Plot=True, Save=False)
| docs/source/auto_examples/ComputingFields/Fields:FarField.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:batchscoringdl]
# language: python
# name: conda-env-batchscoringdl-py
# ---
# # 6. Deploy and Test Logic Apps
# Now that we've verified that our Docker image works, this notebook will walk through how to operationalize this workflow using [Logic Apps](https://azure.microsoft.com/en-us/services/logic-apps/).
#
# For this workflow, adding a file whose name begins with 'trigger' will kick-off the entire batch style transfer process. In this architecture, we use Logic App as our trigger mechanism: when a trigger file (ex. 'trigger.txt') is uploaded into blob storage, Logic App will detect that the file has been added, and create an [Azure Container Instance (ACI)](https://azure.microsoft.com/en-us/services/container-instances/) to start creating Batch AI jobs.
#
# This notebook will rely on the file __template.logic_app.json__ to deploy the Logic App. This file is a template for a [Azure Resource Manager (ARM)](https://docs.microsoft.com/en-us/azure/azure-resource-manager/resource-group-overview) json file, which we will use in this notebook to deploy the resources. In its current state, this file does not contain any of the parameters, such as those stored in the .env file, that we've set throughout the tutorial. To fill out those parameters, we will use [jinja](http://jinja.pocoo.org/) to populate the template with all the appropriate parameters and write the output to __logic_app.json__. This will be ARM json file that we use with the __az cli__ to deploy the Logic App.
#
# As part of the Logic App deployment, we'll also deploy 2 additional resources: an Azure blob storage connector and an ACI connector. These connectors will allow Logic App to talk to these two resources.
# ---
# Import utilities and load environment variables. In this notebook, we'll be using jinja2 to populate our Logic App deployment.
import json
import os
import jinja2
# %load_ext dotenv
# %dotenv
# ## Deploy Logic App
# Set variables we'll need for Logic App deployment:
#
# - `logic_app` - the name of your logic app
# - `aci_container_group` - the name of your ACI
# - `aci_display_name` - a display name used for your ACI deployment
logic_app = "<my-logic-app>"
aci_container_group = "<my-aci-group>"
aci_display_name = "<<EMAIL>>"
# Persist these variables to our `.env` file
# !dotenv set LOGIC_APP $logic_app
# !dotenv set ACI_CONTAINER_GROUP $aci_container_group
# !dotenv set ACI_DISPLAY_NAME $aci_display_name
# Reload our environment variables from our `.env` file so we can use these variables as enviroment variables.
# %reload_ext dotenv
# %dotenv
# Using Jinja, populate the `template.logic_app.json` file and output the new file as `logic_app.json`. This file will be saved in the working directory.
# +
# use jinja to fill in variables from .env file
env = jinja2.Environment(
loader=jinja2.FileSystemLoader('.')
)
template = env.get_template('template.logic_app.json')
e = os.environ
rendered_template = template.render(env=e)
out = open('logic_app.json', 'w')
out.write(rendered_template)
out.close()
# -
# Now that we have the `logic_app.json` file, we can use the __az cli__ to deploy the resource.
# + language="bash"
# az group deployment create \
# --name $LOGIC_APP \
# --resource-group $RESOURCE_GROUP \
# --template-file logic_app.json
# -
# Once the Logic App is deployed, go into the Azure portal and open up the ACI connector and the Azure blob connector to authenticate.
#
# When you open up up the Azure ACI connector, it should look like this:
#
# 
#
# When you open up up the Azure blob connector, it should look like this:
#
# 
#
# For both of these connectors, click on the orange bar at the top to authenticate.
#
# Once authenticated, your Logic App should be all set up and ready to trigger the workflow.
# ## Trigger Logic App
# Logic App will be triggered whenever a blob that begins with the string 'trigger' is uploaded into the Azure blob storage account. Logic App will then read the contents of the file, using the contents as the directory of individual frames to apply style transfer to.
# First we'll create a file titled `trigger_0.txt` and input the name of the content directory in blob to apply style transfer to.
# ! touch trigger_0.txt && echo $FS_CONTENT_DIR > trigger_0.txt
# Using AzCopy, upload the trigger file into blob to trigger Logic App.
# + language="bash"
# azcopy \
# --source trigger_0.txt \
# --destination https://${STORAGE_ACCOUNT_NAME}.blob.core.windows.net/${AZURE_CONTAINER_NAME}/trigger_0.txt \
# --dest-key $STORAGE_ACCOUNT_KEY
# -
# This file will trigger Logic App to create an ACI that will kick off the Batch AI jobs. We can now inspect the logs from ACI to see that everything is going smoothly. If you run this command right after uploading the trigger file, the logs may not be found as it will take some time to trigger Logic App to create the ACI group.
# !az container logs --resource-group $RESOURCE_GROUP --name $ACI_CONTAINER_GROUP
# At this point, we can inspect the Azure portal to see all the moving parts:
# - Logic Apps will be triggers and will spin up ACI
# - ACI will break up the content images in blob and create BatchAI jobs
# - The BatchAI cluster will scale up and start processing the work
# - As the style transfer script is executed in batch on BatchAI, we will see the completed images (as well as logs) saved back to blob
# ---
# ## Conclusion
# In this notebook, we deployed Logic App and simulated what this end-to-end workflow could look like when operationalized with an add-to-blob trigger. At this point, I'd encourage you to take a look through the portal and examine the individual components of the architecture and make sure that all the parts are operating as expected. For further customization, you can also edit the Logic App via the Logic App Designer (in the portal) so that the triggering mechanism is more fitted to your scenarios.
#
# The last step of the tutorial is to [download and re-stitch the individual frames so that we have a single style transferred video.](07_stitch_together_the_results.ipynb)
| 06_deploy_and_test_logic_apps.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Modeling and Simulation in Python
#
# Chapter 24
#
# Copyright 2017 <NAME>
#
# License: [Creative Commons Attribution 4.0 International](https://creativecommons.org/licenses/by/4.0)
#
# +
# Configure Jupyter so figures appear in the notebook
# %matplotlib inline
# Configure Jupyter to display the assigned value after an assignment
# %config InteractiveShell.ast_node_interactivity='last_expr_or_assign'
# import functions from the modsim.py module
from modsim import *
# -
# ### Rolling paper
#
# We'll start by loading the units we need.
radian = UNITS.radian
m = UNITS.meter
s = UNITS.second
# And creating a `Params` object with the system parameters
params = Params(Rmin = 0.02 * m,
Rmax = 0.055 * m,
L = 47 * m,
omega = 10 * radian / s,
t_end = 130 * s,
dt = 1*s)
# The following function estimates the parameter `k`, which is the increase in the radius of the roll for each radian of rotation.
def estimate_k(params):
"""Estimates the parameter `k`.
params: Params with Rmin, Rmax, and L
returns: k in meters per radian
"""
Rmin, Rmax, L = params.Rmin, params.Rmax, params.L
Ravg = (Rmax + Rmin) / 2
Cavg = 2 * pi * Ravg
revs = L / Cavg
rads = 2 * pi * revs
k = (Rmax - Rmin) / rads
return k
# As usual, `make_system` takes a `Params` object and returns a `System` object.
def make_system(params):
"""Make a system object.
params: Params with Rmin, Rmax, and L
returns: System with init, k, and ts
"""
init = State(theta = 0 * radian,
y = 0 * m,
r = params.Rmin)
k = estimate_k(params)
return System(params, init=init, k=k)
# Testing `make_system`
system = make_system(params)
system.init
# Now we can write a slope function based on the differential equations
#
# $\omega = \frac{d\theta}{dt} = 10$
#
# $\frac{dy}{dt} = r \frac{d\theta}{dt}$
#
# $\frac{dr}{dt} = k \frac{d\theta}{dt}$
#
def slope_func(state, t, system):
"""Computes the derivatives of the state variables.
state: State object with theta, y, r
t: time
system: System object with r, k
returns: sequence of derivatives
"""
theta, y, r = state
k, omega = system.k, system.omega
dydt = r * omega
drdt = k * omega
return omega, dydt, drdt
# Testing `slope_func`
slope_func(system.init, 0, system)
# We'll use an event function to stop when `y=L`.
def event_func(state, t, system):
"""Detects when we've rolled length `L`.
state: State object with theta, y, r
t: time
system: System object with L
returns: difference between `y` and `L`
"""
theta, y, r = state
return y - system.L
event_func(system.init, 0, system)
# Now we can run the simulation.
results, details = run_ode_solver(system, slope_func, events=event_func)
details
# And look at the results.
results.tail()
# The final value of `y` is 47 meters, as expected.
unrolled = get_last_value(results.y)
# The final value of radius is `R_max`.
radius = get_last_value(results.r)
# The total number of rotations is close to 200, which seems plausible.
radians = get_last_value(results.theta)
rotations = magnitude(radians) / 2 / np.pi
# The elapsed time is about 2 minutes, which is also plausible.
t_final = get_last_label(results) * s
# ### Plotting
# Plotting `theta`
# +
def plot_theta(results):
plot(results.theta, color='C0', label='theta')
decorate(xlabel='Time (s)',
ylabel='Angle (rad)')
plot_theta(results)
# -
# Plotting `y`
# +
def plot_y(results):
plot(results.y, color='C1', label='y')
decorate(xlabel='Time (s)',
ylabel='Length (m)')
plot_y(results)
# -
# Plotting `r`
# +
def plot_r(results):
plot(results.r, color='C2', label='r')
decorate(xlabel='Time (s)',
ylabel='Radius (m)')
plot_r(results)
# -
# We can also see the relationship between `y` and `r`, which I derive analytically in the book.
# +
plot(results.r, results.y, color='C3')
decorate(xlabel='Radius (m)',
ylabel='Length (m)',
legend=False)
# -
# And here's the figure from the book.
# +
def plot_three(results):
subplot(3, 1, 1)
plot_theta(results)
subplot(3, 1, 2)
plot_y(results)
subplot(3, 1, 3)
plot_r(results)
plot_three(results)
savefig('figs/chap24-fig01.pdf')
# -
# ### Animation
#
# Here's a draw function that animates the results using `matplotlib` patches.
# +
from matplotlib.patches import Circle
from matplotlib.patches import Arrow
def draw_func(state, t):
# get radius in mm
theta, y, r = state
radius = r.magnitude * 1000
# draw a circle with
circle = Circle([0, 0], radius, fill=True)
plt.gca().add_patch(circle)
# draw an arrow to show rotation
dx, dy = pol2cart(theta, radius)
arrow = Arrow(0, 0, dx, dy)
plt.gca().add_patch(arrow)
# make the aspect ratio 1
plt.axis('equal')
# -
animate(results, draw_func)
# **Exercise:** Run the simulation again with a smaller step size to smooth out the animation.
# ### Exercises
#
# **Exercise:** Since we keep `omega` constant, the linear velocity of the paper increases with radius. Use `gradient` to estimate the derivative of `results.y`. What is the peak linear velocity?
# +
# Solution
dydt = gradient(results.y);
# -
plot(dydt, label='dydt')
decorate(xlabel='Time (s)',
ylabel='Linear velocity (m/s)')
# +
# Solution
linear_velocity = get_last_value(dydt) * m/s
# -
# Now suppose the peak velocity is the limit; that is, we can't move the paper any faster than that.
#
# Nevertheless, we might be able to speed up the process by keeping the linear velocity at the maximum all the time.
#
# Write a slope function that keeps the linear velocity, `dydt`, constant, and computes the angular velocity, `omega`, accordingly.
#
# Run the simulation and see how much faster we could finish rolling the paper.
# +
# Solution
def slope_func(state, t, system):
"""Computes the derivatives of the state variables.
state: State object with theta, y, r
t: time
system: System object with r, k
returns: sequence of derivatives
"""
theta, y, r = state
k, omega = system.k, system.omega
dydt = linear_velocity
omega = dydt / r
drdt = k * omega
return omega, dydt, drdt
# +
# Solution
slope_func(system.init, 0, system)
# +
# Solution
results, details = run_ode_solver(system, slope_func, events=event_func)
details
# +
# Solution
t_final = get_last_label(results) * s
# +
# Solution
plot_three(results)
# -
| soln/chap24soln.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # NOAA : Calcule des pluviométries précédentes
#
# Dataset :
# - le resultat précédent
#
# Exercice :
# - Créer pour chaque ligne des colonnes contenant la (moyenne,somme,max) des 5, 10, 20 jours précédants
#
# (PAYS, Date, avg_prcp, sum_rain) ajouter (avg_rain_last_5_days, avg_rain_last_10_days, sum_rain_last_5_days, max_rain_last_5_days)
#
#
#
#
# Aide : https://kevinvecmanis.io/pyspark/data%20science/python/2019/06/02/SPX-Analysis-With-PySpark.html
# +
# #!hdfs dfs -ls /demo/noaa/noaa-raw-data
# -
from pyspark import SparkContext
from pyspark import SparkConf
from pyspark.sql import SparkSession
import pyspark.sql.functions as F
import shutil
# +
DATASET_FOLDER = '/xxxx'
SPARK_MASTER = 'spark://localhost:7077'
APP_NAME = 'NOAA Window - 430'
noaa_csv_path = DATASET_FOLDER
output = '/stagiaire/votre_dossier/noaa/daily_rain_by_country_window'
# Create Spark session
spark = SparkSession.builder.master(SPARK_MASTER).appName(APP_NAME).getOrCreate()
# -
sqlDF.write.csv(output)
print(sqlDF.columns)
sc.stop()
| demo/spark_056_NOAA_3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
sys.path.append('../build')
import numpy as np
import libry as ry
# +
K = ry.Config()
K.addFile("../rai-robotModels/pr2/pr2.g")
K.addFile("../models/tables.g")
K.addFrame("obj0", "table1", "type:ssBox size:[.1 .1 .2 .02] color:[1. 0. 0.], contact, logical={ object }, joint:rigid, Q:<t(0 0 .15)>" )
K.addFrame("obj1", "table1", "type:ssBox size:[.1 .1 .2 .02] color:[1. 0. 0.], contact, logical={ object }, joint:rigid, Q:<t(0 .2 .15)>" )
K.addFrame("obj2", "table1", "type:ssBox size:[.1 .1 .2 .02] color:[1. 0. 0.], contact, logical={ object }, joint:rigid, Q:<t(0 .4 .15)>" )
K.addFrame("obj3", "table1", "type:ssBox size:[.1 .1 .2 .02] color:[1. 0. 0.], contact, logical={ object }, joint:rigid, Q:<t(0 .6.15)>" )
K.addFrame("tray", "table2", "type:ssBox size:[.15 .15 .04 .02] color:[0. 1. 0.], logical={ table }, Q:<t(0 0 .07)>" );
K.addFrame("", "tray", "type:ssBox size:[.27 .27 .04 .02] color:[0. 1. 0.]" )
V = K.view()
# -
lgp = K.lgp("../models/fol-pickAndPlace.g")
lgp.nodeInfo()
# this writes the initial state, which is important to check:
# do the grippers have the gripper predicate, do all objects have the object predicate, and tables the table predicate? These need to be set using a 'logical' attribute in the g-file
# the on predicate should automatically be generated based on the configuration
lgp.getDecisions()
# This is also useful to check: inspect all decisions possible in this node, which expands the node.
# If there is no good decisions, the FOL rules are buggy
lgp.walkToDecision(3)
lgp.nodeInfo()
# Using getDecisions and walkToDecision and walkToParent, you can walk to anywhere in the tree by hand
lgp.viewTree()
lgp.walkToNode("(grasp pr2R obj0) (grasp pr2L obj1) (place pr2R obj0 tray)")
lgp.nodeInfo()
# at a node, you can compute bounds, namely BT.seq (just key frames), BT.path (the full path),
# and BT.setPath (also the full path, but seeded with the BT.seq result)
lgp.optBound(ry.BT.seq, True)
lgp.nodeInfo()
komo = lgp.getKOMOforBound(ry.BT.seq)
komo.display()
komo = 0
lgp.optBound(ry.BT.path, True)
lgp.nodeInfo()
lgp.viewTree()
# finally, the full multi-bound tree search (MBTS)
# you typically want to add termination rules, i.e., symbolic goals
print("THIS RUNS A THREAD. CHECK THE CONSOLE FOR OUTPUT. THIS IS GENERATING LOTS OF FILES.")
lgp.addTerminalRule("(on obj0 tray) (on obj1 tray) (on obj2 tray)")
lgp.run(2)
# wait until you have some number of solutions found (repeat executing this line...)
lgp.numSolutions()
# query the optimization features of the 0. solution
lgp.getReport(0, ry.BT.seqPath)
# get the KOMO object for the seqPath computation of the 0. solution
komo = lgp.getKOMO(0, ry.BT.seqPath)
komo.displayTrajectory() #SOOO SLOOOW (TODO: add parameter for display speed)
# assign K to the 20. configuration of the 0. solution, check display
# you can now query anything (joint state, frame state, features)
X = komo.getConfiguration(20)
K.setFrameState(X)
lgp.stop() #stops the thread... takes a while to finish the current job
lgp.run(2) #will continue where it stopped
komo=0
lgp=0
# +
import sys
sys.path.append('../rai/rai/ry')
import numpy as np
import libry as ry
C = ry.Config()
D = C.view()
C.addFile('../test/lgp-example.g');
lgp = C.lgp("../test/fol.g");
# -
lgp.walkToNode("(grasp baxterR stick) (push stickTip redBall table1) (grasp baxterL redBall) ");
print(lgp.nodeInfo())
lgp.optBound(BT.pose, True);
# +
komo = lgp.getKOMOforBound(BT.path)
komo.display()
input("Press Enter to continue...")
# -
| tutorials/lgp1-pickAndPlace.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.6.13 64-bit (''csls'': conda)'
# name: python3
# ---
import pickle
import numpy as np
import matplotlib.pyplot as plt
import sys
sys.path.append('..')
from utils.util import *
import seaborn as sns
sns.set_theme()
sns.set_style("darkgrid")
# sns.set_context("paper")
sns.set_context("talk")
ctx_order = 'first'
ctx_order_str = 'ctxF'
analyze_name = 'proportions'
# +
# with open('../../results/%s_results_mlp.P' %(analyze_name), 'rb') as f:
# mlp_results = pickle.load(f)
# # ----------------------
# ctx_order = 'first'
# ctx_order_str = 'ctxF'
# # ----------------------
# with open('../../results/%s_%s_results_rnn.P' %(analyze_name, ctx_order_str), 'rb') as f:
# rnn_results_ctxF = pickle.load(f)
# with open('../../results/%s_%s_results_rnncell.P' %(analyze_name, ctx_order_str), 'rb') as f:
# rnncell_results_ctxF = pickle.load(f)
# with open('../../results/%s_%s_results_rnn_balanced.P' %(analyze_name, ctx_order_str), 'rb') as f:
# rnnb_results_ctxF = pickle.load(f)
# # ----------------------
# ctx_order = 'last'
# ctx_order_str = 'ctxL'
# # ----------------------
# with open('../../results/%s_%s_results_rnn.P' %(analyze_name, ctx_order_str), 'rb') as f:
# rnn_results_ctxL = pickle.load(f)
# with open('../../results/%s_%s_results_rnncell.P' %(analyze_name, ctx_order_str), 'rb') as f:
# rnncell_results_ctxL = pickle.load(f)
# with open('../../results/%s_%s_results_rnn_balanced.P' %(analyze_name, ctx_order_str), 'rb') as f:
# rnnb_results_ctxL = pickle.load(f)
# # ----------------------
# with open('../../results/%s_results_stepwisemlp.P' %(analyze_name), 'rb') as f:
# swmlp_results = pickle.load(f)
# with open('../../results/%s_results_truncated_stepwisemlp.P' %(analyze_name), 'rb') as f:
# swmlp_trunc_results = pickle.load(f)
# # ----------------------
# with open('../../results/%s_results_mlp_cc.P' %(analyze_name), 'rb') as f:
# mlpcc_results = pickle.load(f)
# +
# mlp_runs = dict_to_list(mlp_results, analyze_name)
# rnn_runs_ctxF = dict_to_list(rnn_results_ctxF, analyze_name)
# rnncell_runs_ctxF = dict_to_list(rnncell_results_ctxF, analyze_name)
# rnnb_runs_ctxF = dict_to_list(rnnb_results_ctxF, analyze_name)
# rnn_runs_ctxL = dict_to_list(rnn_results_ctxL, analyze_name)
# rnncell_runs_ctxL = dict_to_list(rnncell_results_ctxL, analyze_name)
# rnnb_runs_ctxL = dict_to_list(rnnb_results_ctxL, analyze_name)
# swmlp_runs = dict_to_list(swmlp_results, analyze_name)
# mlpcc_runs = dict_to_list(mlpcc_results, analyze_name)
# mlp_runs.keys()
# +
with open('../../results/%s_results_mlp.P' %(analyze_name), 'rb') as f:
mlp_results = pickle.load(f)
with open('../../results/%s_%s_results_rnn.P' %(analyze_name, ctx_order_str), 'rb') as f:
rnn_results = pickle.load(f)
with open('../../results/%s_%s_results_rnncell.P' %(analyze_name, ctx_order_str), 'rb') as f:
rnncell_results = pickle.load(f)
with open('../../results/%s_results_stepwisemlp.P' %(analyze_name), 'rb') as f:
swmlp_results = pickle.load(f)
# -
mlp_runs = dict_to_list(mlp_results, analyze_name)
rnn_runs = dict_to_list(rnn_results, analyze_name)
rnncell_runs = dict_to_list(rnncell_results, analyze_name)
swmlp_runs = dict_to_list(swmlp_results, analyze_name)
mlp_runs.keys()
# +
hiddens_ctxs_mlp = np.asarray(mlp_runs['hiddens_ctxs']) # [run, checkpoints, n_ctxs, n_trials, hidd_dim]
p_pies_mlp = np.asarray(mlp_runs['p_pies']) # [run, checkpoints, n_ctxs, hidd_dim]
ps_mlp = np.asarray(mlp_runs['ps']) # [run, checkpoints, n_ctxs, hidd_dim]
n_mlp = np.asarray(mlp_runs['n']) # [run, checkpoints, n_ctxs, hidd_dim]
n_runs, n_cps, _, hidd_dim = n_mlp.shape
print(n_runs, n_cps, hidd_dim)
hiddens_ctxs_swmlp = np.asarray(swmlp_runs['hiddens_ctxs'])
p_pies_swmlp = np.asarray(swmlp_runs['p_pies'])
ps_swmlp = np.asarray(swmlp_runs['ps'])
n_swmlp = np.asarray(swmlp_runs['n'])
# -
def pie_bar_chart_data(pies):
n_ctxs = pies.shape[0]
n_hidd = pies.shape[1]
ctxs = np.zeros(n_ctxs+2) # ctxs[-1] refers to none, and ctxs[-2] to mixed
for i in range(n_hidd):
ctx = pies[:,i]
if not np.any(ctx):
ctxs[-1]+=1
elif np.sum(ctx)>1:
ctxs[-2]+=1
else:
idx = np.where(ctx) # this finds which context is 1
ctxs[idx]+=1
labels = []
for ctx in range(n_ctxs):
labels.append('ctx%s' %(ctx))
labels.append('mixed')
labels.append('none')
return ctxs, labels
w_cp = -1 # last checkpoint, w stands for which
w_run = 15
if w_run==-1:
run_label = 'Last'
else:
run_label = '#%s' %(w_run)
# +
fig, axs = plt.subplots(1, 2, figsize=[18,8])
ps = ps_mlp
ps = ps[:,w_cp,:,:]
print(ps.shape)
ax = axs[0]
for run in range(n_runs):
# colors shows different runs
ax.scatter(ps[run][0], ps[run][1])
ax.set_title('MLP')
ps = ps_swmlp
ps = ps[:,w_cp,:,:]
print(ps.shape)
ax = axs[1]
for run in range(n_runs):
# colors shows different runs
ax.scatter(ps[run][0], ps[run][1])
ax.set_title('Stepwise MLP')
for ax in axs:
ax.set_xlim([0,1])
ax.set_ylim([0,1])
ax.set_xlabel('Context1')
ax.set_ylabel('Context2')
fig.suptitle('Proportion Results - Multiple Runs', fontweight='bold')
plt.tight_layout()
fig_str = 'proportion_results_both_mlps_face_scatter'
fig.savefig(('../../figures/' + fig_str + '.pdf'),
bbox_inches = 'tight', pad_inches = 0)
fig.savefig(('../../figures/' + fig_str + '.png'),
bbox_inches = 'tight', pad_inches = 0)
# +
fig, axs = plt.subplots(2, 2, figsize=[18,9])
p_pies = p_pies_mlp
p_pies = p_pies[w_run,w_cp,:,:]
ctxs, labels = pie_bar_chart_data(p_pies)
ax = axs[0][0]
ax.pie(ctxs, labels=labels)#, autopct='%1.1f%%')#, shadow=True)#, startangle=90)
ax.axis('equal')
ax = axs[0][1]
model = 'mlp'
for ctx in range(ctxs.shape[0]):
ax.bar(model, ctxs[ctx], bottom = np.sum(ctxs[:ctx]))
ax.legend(labels)
ax.set_title('MLP')
p_pies = p_pies_swmlp
p_pies = p_pies[w_run,w_cp,:,:]
ctxs, labels = pie_bar_chart_data(p_pies)
ax = axs[1][0]
ax.pie(ctxs, labels=labels)#, autopct='%1.1f%%')#, shadow=True)#, startangle=90)
ax.axis('equal')
ax = axs[1][1]
model = 'stepwisemlp'
for ctx in range(ctxs.shape[0]):
ax.bar(model, ctxs[ctx], bottom = np.sum(ctxs[:ctx]))
ax.legend(labels)
ax.set_title('Stepwise MLP')
fig.suptitle('Proportion Results - P pies - %s Run' %(run_label), fontweight='bold')
plt.tight_layout()
fig_str = 'proportion_results_both_mlps_face_p_pies'
fig.savefig(('../../figures/' + fig_str + '.pdf'),
bbox_inches = 'tight', pad_inches = 0)
fig.savefig(('../../figures/' + fig_str + '.png'),
bbox_inches = 'tight', pad_inches = 0)
# hidden units selectivity
# proportion of hidden units that were active at all (even one trial) for each context
# we have ... many hiddens that were active for both (mixed), ... many units that were active for ctx1, so on so forth
# -
def calc_hidd_sens(hidds_act):
n_runs, n_cps, n_ctxs, n_hidd = hidds_act.shape
ctxs = np.zeros([n_runs, n_cps, n_ctxs+2]) # ctxs[-1] refers to none, and ctxs[-2] to mixed
for cp in range(n_cps):
for r in range(n_runs):
for i in range(n_hidd):
ctx = hidds_act[r,cp,:,i] # [n_ctxs]
# print(ctx)
# print(n[r,cp,:,i])
if not np.any(ctx): # none
ctxs[r,cp,-1]+=1
elif np.sum(ctx)>1: # mixed
ctxs[r,cp,-2]+=1
else:
idx = np.where(ctx) # this finds which context is 1
ctxs[r,cp,idx]+=1
return ctxs
# +
n_ctxs = 2
# plot
fig, axs = plt.subplots(1,2,figsize=[10, 5])
ax = axs
# labels
labels = []
for ctx in range(n_ctxs):
labels.append('ctx%s' %(ctx))
if n_ctxs==2:
labels.append('none/mixed')
n = n_mlp
ctxs = calc_hidd_sens(n)
m_ctxs = np.mean(ctxs, axis=0) # take avg over runs
ax = axs[0]
for ctx in range(m_ctxs.shape[1]):
ax.bar(range(n_cps), m_ctxs[:,ctx], bottom = np.sum(m_ctxs[:,:ctx], axis=1))
ax.set_title('MLP')
ax.set_ylabel('Normalized Hidden Units Sensitivity (Avg over runs)')
n = n_swmlp
ctxs = calc_hidd_sens(n)
m_ctxs = np.mean(ctxs, axis=0) # take avg over runs
ax = axs[1]
for ctx in range(m_ctxs.shape[1]):
ax.bar(range(n_cps), m_ctxs[:,ctx], bottom = np.sum(m_ctxs[:,:ctx], axis=1))
ax.set_title('Stepwise MLP')
for ax in axs:
ax.legend(labels)
ax.set_xlabel('Steps')
fig.suptitle('Proportion Results - N', fontweight='bold')
plt.tight_layout()
fig_str = 'proportion_results_both_mlps_face_n_overCPs'
fig.savefig(('../../figures/' + fig_str + '.pdf'),
bbox_inches = 'tight', pad_inches = 0)
fig.savefig(('../../figures/' + fig_str + '.png'),
bbox_inches = 'tight', pad_inches = 0)
# +
# first threshold,
# then count
# then averge over those
# none here means none + mixed
# given the overall activities for this hidd unit, the prop of trials that this unit was active that was above threshold (e.g, above 25%)
# n_ctxs=2, chance = 0.50, threshold = 95%
# labels
labels = []
for ctx in range(n_ctxs):
labels.append('ctx%s' %(ctx))
if n_ctxs==2:
n_thresh = 0.9
labels.append('none/mixed')
labels.append('none/mixed')
# plot
fig, axs = plt.subplots(1,2,figsize=[10, 5])
n = n_mlp
f = n > n_thresh
ctxs = calc_hidd_sens(f)
m_ctxs = np.mean(ctxs, axis=0) # take avg over runs
ax = axs[0]
for ctx in range(m_ctxs.shape[1]):
ax.bar(range(n_cps), m_ctxs[:,ctx], bottom = np.sum(m_ctxs[:,:ctx], axis=1))
ax.set_title('MLP')
n = n_swmlp
f = n > n_thresh
ctxs = calc_hidd_sens(f)
m_ctxs = np.mean(ctxs, axis=0) # take avg over runs
ax = axs[1]
for ctx in range(m_ctxs.shape[1]):
ax.bar(range(n_cps), m_ctxs[:,ctx], bottom = np.sum(m_ctxs[:,:ctx], axis=1))
ax.set_title('Stewise MLP')
for ax in axs:
ax.legend(labels)
plt.tight_layout()
fig.suptitle('Proportion Results - F - Threshold %s' %(n_thresh), fontweight='bold')
plt.tight_layout()
fig_str = 'proportion_results_both_mlps_face_f_overCPs'
fig.savefig(('../../figures/' + fig_str + '.pdf'),
bbox_inches = 'tight', pad_inches = 0)
fig.savefig(('../../figures/' + fig_str + '.png'),
bbox_inches = 'tight', pad_inches = 0)
# n: [n_ctxs, hidden_dim]
# normalized - for each ctx, for how many trials each unit was active (filtered by the threshold)
# normalized by the overall activity of that unit for all ctx and trials
# we have ... many units that were active for ctx0 for more than n_threshold% of the trials
# -
# # normalized matrix
# +
fig, axs = plt.subplots(1,2,figsize=[20, 10])
# plot 3P, using normalized p
sort_ind = 0
labels = []
for ctx in range(n_ctxs):
labels.append('ctx%s' %(ctx))
# todo: sort the columns, based on the one context
n = n_mlp
_,_,n_ctxs,hidd_dim = n.shape
n = n[w_run, w_cp, :, :]
n[sort_ind, :] = np.sort(n[sort_ind, :])
ax = axs[0]
for ctx in range(n_ctxs):
ax.bar(range(hidd_dim), n[ctx,:], bottom = np.sum(n[:ctx,:], axis=0))
ax.set_title('MLP')
ax.set_ylabel('Normalized Hidden Activities')
n = n_swmlp
_,_,n_ctxs,hidd_dim = n.shape
n = n[w_run, w_cp, :, :]
n[sort_ind, :] = np.sort(n[sort_ind, :])
ax = axs[1]
for ctx in range(n_ctxs):
ax.bar(range(hidd_dim), n[ctx,:], bottom = np.sum(n[:ctx,:], axis=0))
ax.set_title('stepwise MLP')
for ax in axs:
ax.set_ylim([0,2])
ax.set_xlabel('Hidden Units')
ax.legend(labels)
fig.suptitle('Proportion Results - %s Run - N' %(run_label), fontweight='bold')
plt.tight_layout()
fig_str = 'proportion_results_both_mlps_face_n'
fig.savefig(('../../figures/' + fig_str + '.pdf'),
bbox_inches = 'tight', pad_inches = 0)]
fig.savefig(('../../figures/' + fig_str + '.png'),
bbox_inches = 'tight', pad_inches = 0)
# n: [n_ctxs, hidden_dim]
# normalized - how much each unit is active for each ctx over trials
# normalized by the overall activity of that unit for all ctx and trials
# x-axis shows each unit (128 hidden units total)
# y-axis shows normalized activity (number of trials) of that unit for each context over all the trials,
# Q: what happens to those that were only active for the ctx1?
# -
| notebooks/results_proportions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import nltk.data
import nltk
from nltk.corpus import stopwords
from nltk.stem import PorterStemmer
from nltk.stem.snowball import SnowballStemmer
import re
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
import gensim
from keras.models import Model
from keras.layers import Input, Embedding, Dense, Dropout, Flatten
from keras.layers import Activation
import numpy as np
import chardet
import logging
from keras.models import Model
from keras.layers import Input,Dense,Dropout,Flatten
from keras.layers import Conv1D,MaxPooling1D,AveragePooling1D
from keras.layers import LSTM,Bidirectional
from keras.callbacks import ModelCheckpoint
from keras.callbacks import EarlyStopping
from keras.models import load_model
import datetime
import time
import os
def read_data(filename):
df = pd.read_csv(filename)
return df
def stemming(text):
text = [stemmer.stem(word) for word in text]
return " ".join(text)
def cleantext(dataframe,stem):
#Cleaning text
dataframe["clean_text"] = dataframe["docket_text"]
#Get bundle number
dataframe['bundle_motion_number'] = dataframe['bundle_motion_number'].apply(lambda x: np.nan if x == 'None' else float(x))
#Replace bundle motion number
dataframe.clean_text = dataframe.apply(lambda x: x.clean_text if np.isnan(x.bundle_motion_number)
else x.clean_text.replace(str(int(x.bundle_motion_number)), 'DLLM'), axis = 1)
#Remove numbers
dataframe.clean_text = dataframe.clean_text.str.replace(r'\d+','')
#Remove floats
dataframe.clean_text = dataframe.clean_text.astype(str).replace(r'(\d*\.?\d*)','')
#Remove Punctuation
dataframe.clean_text = dataframe.clean_text.astype(str).apply(lambda x : " ".join(re.findall('[\w]+',x)))
#Convert to lower text
dataframe.clean_text = dataframe.clean_text.astype(str).apply(lambda x: " ".join(x.lower() for x in x.split()))
#Stop words removal
dataframe.clean_text.astype(str)
stop = stopwords.words('english')
# dataframe.loc[:,"docket_text_new"] = dataframe.loc[:,"docket_text_new"].apply(lambda x: ' '.join([word for word in x.split() if word not in (stop)]))
dataframe.clean_text = dataframe.clean_text.apply(lambda x: " ".join(x.lower() for x in str(x).split() if x not in stop))
# Adding "Start" & "End" at the beginning & end of each docket text
dataframe.clean_text = " START " + dataframe.clean_text + " END"
dataframe.clean_text = [nltk.word_tokenize(i) for i in dataframe.clean_text]
# Moving cleaned text to another column
# dataframe['cleaned_text'] = dataframe["docket_text_new"]
if stem == True:
dataframe.clean_text = dataframe.clean_text.apply(stemming)
dataframe.clean_text.head(5)
##Stemming
stemmer = SnowballStemmer("english")
def clean_text(dataframe,stem,stop,text_col):
#Cleaning text
dataframe["doc"] = dataframe[text_col]
#Remove numbers
dataframe.loc[:,"doc"] = dataframe.doc.str.replace(r'\d+','')
#Remove floats
dataframe.loc[:,"doc"] = dataframe.doc.astype(str).replace(r'(\d*\.?\d*)','')
#Remove Punctuation
dataframe.loc[:,"doc"] = dataframe.doc.astype(str).apply(lambda x : " ".join(re.findall('[\w]+',x)))
#Convert to lower text
dataframe.loc[:,"doc"] = dataframe.doc.astype(str).apply(lambda x: " ".join(x.lower() for x in x.split()))
#Stop words removal
if stop == True:
dataframe.loc[:,"doc"] = dataframe.loc[:,"doc"].astype(str)
stop = stopwords.words('english')
dataframe.loc[:,"doc"] = dataframe.loc[:,"doc"].apply(lambda x: " ".join(x.lower() for x in str(x).split() if x not in stop))
#Stemming
if stem == True:
stemmer = SnowballStemmer("english")
curried_stemming = curry(stemming, stemmer=stemmer)
dataframe.loc[:,"doc"] = dataframe.loc[:,"docket_text_new"].apply(curried_stemming)
# Adding "Start" & "End" at the beginning & end of each docket text
dataframe.loc[:,"doc"] = " START " + dataframe.loc[:,"doc"] + " END"
# Moving cleaned text to another column
dataframe['cleaned_text'] = dataframe["doc"]
#Tokenization
dataframe.loc[:,"doc"] = dataframe.loc[:,"doc"].apply(nltk.word_tokenize)
return dataframe
def tokenizetext(dataframe):
dataframe.clean_text = [nltk.word_tokenize(i) for i in dataframe.clean_text]
def get_list(dataframe):
words = dataframe["clean_text"].tolist()
flat_list_words = [item for sublist in words for item in sublist]
return flat_list_words
def w2c_model(doc,size,window,min_count,workers,iter,sg,length,numepoch):
model = gensim.models.Word2Vec(size=size, window=window, min_count=min_count, workers=workers,iter=iter,sg=sg)
model.build_vocab(doc)
model.train(doc,total_examples=length,epochs = numepoch)
return model
def text2seq(doc, w2v, seq_len, emb_size):
padding = np.array([0 for __ in range(emb_size)])
seq = []
for token in doc[: seq_len]:
try:
seq.append(w2v.wv[token])
except:
aa = 1
seq += [padding for __ in range(seq_len - len(seq))]
#print(len(seq))
return seq
def one_hot(series):
values=sorted(list(set(series)))
return np.vstack(map(lambda x: [x==v for v in values],series))
def prepro(dataframe, w2v,seq_len, emb_size,target_col):
X = dataframe['clean_text'].apply(lambda doc: text2seq(doc, w2v, seq_len, emb_size))
X = np.vstack(X).reshape(len(dataframe), seq_len, emb_size)
y = one_hot(dataframe[target_col])
return X, y
#Model building
def model1(seq_len, emb_size, num_labels):
input_layer = Input(shape=(seq_len, emb_size))
x = Conv1D(32, 3, activation='relu')(input_layer)
x = Flatten()(x)
x = Dense(256, activation='relu')(x)
x = Dropout(0.2)(x)
x = Dense(64, activation='relu')(x)
x = Dropout(0.2)(x)
x = Dense(128, activation='relu')(x)
x = Dropout(0.2)(x)
output_layer = Dense(num_labels, activation='softmax')(x)
model = Model(input_layer, output_layer)
model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['acc'])
return model
def build_model2(seq_len, emb_size, num_labels):
input_layer = Input(shape=(seq_len, emb_size))
x = Conv1D(32, 3, activation='relu')(input_layer)
x = MaxPooling1D(pool_size=2)(x)
x = LSTM(100, return_sequences = True,dropout=0.2,recurrent_dropout=0.05)(input_layer)
x = Flatten()(x)
# x = Dense(256, activation='relu')(x)
# x = Dropout(0.2)(x)
# x = Dense(64, activation='relu')(x)
# x = Dropout(0.2)(x)
# x = Dense(128, activation='relu')(x)
# x = Dropout(0.2)(x)
output_layer = Dense(num_labels, activation='softmax')(x)
model = Model(input_layer, output_layer)
model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['acc'])
return model
def build_model(seq_len, emb_size, num_labels):
input_layer = Input(shape=(seq_len, emb_size))
x = Conv1D(32, 3, activation='relu')(input_layer)
x = MaxPooling1D(pool_size=2)(x)
x = LSTM(100, dropout=0.2, recurrent_dropout=0.2)(x)
x = Dense(256, activation='relu')(x)
x = Dropout(0.2)(x)
x = Dense(64, activation='relu')(x)
x = Dropout(0.2)(x)
x = Dense(128, activation='relu')(x)
x = Dropout(0.2)(x)
output_layer = Dense(num_labels, activation='softmax')(x)
model = Model(input_layer, output_layer)
model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['acc'])
return model
def fit_model(X,y,weightsfile,num_labels,seq_len,emb_size,epochs):
checkpoint = ModelCheckpoint(weightsfile, monitor='val_acc', verbose=1, save_best_only=True, mode='max')
callbacks_list = [checkpoint]
model_dl = build_model2(seq_len, emb_size, num_labels)
model_dl.fit(X, y, epochs=epochs, batch_size=16,callbacks=callbacks_list, validation_split=0.2)
return model_dl
def get_type(dataframe):
dataframe['type'] = dataframe['type'].apply(lambda x: x if x in ['Motion', 'RR', 'Order'] else 'None')
# dataframe['clean_text'] = dataframe["docket_text"]
def retain_summary_order(dataframe):
Search_for_These_values = ['summary', 'order'] #creating list
pattern = '|'.join(Search_for_These_values) # joining list for comparision
dataframe['motionOrsummary'] = dataframe["clean_text"].str.contains(pattern)
dataframe['motionOrsummary'] = dataframe['motionOrsummary'].map({True: 'Yes', False: 'No'})
dataframe = dataframe[dataframe['motionOrsummary']== 'Yes']
return dataframe
# +
# def predict(model,weightsfile,csvfile,genismfile):
# model.load_weights(weightsfile)
# df = read_data(csvfile)
# get_type(df)
# cleantext(df,False)
# df = retain_summary_order(df)
# tokenizetext(df)
# vocab = get_list(df)
# w2c_model = build_genism_model(vocab = vocab,size=100,window=5,min_count=1,workers=6,iter=10,sg=0,length=length,numepoch=10)
# X1,y1 = prepro(df,gen_model,seq_len=100, emb_size=100)
# preds = model.predict(X1)
# return preds,df
# -
def prediction(model,weightsfile,X):
model.load_weights(weightsfile)
preds = model.predict(X)
lst = list(map(lambda x: x.argmax(), preds))
return preds,lst
def get_prediction_type(preds,dataframe):
lst = list(map(lambda x: x.argmax(), preds))
# dataframe['prediction']= lst
values=sorted(list(set(dataframe['type'])))
return values
def assign_label(dataframe):
dataframe['prediction']= lst
p_class = {0:'Motion',1:'None',2:'Order',3:'RR'}
dataframe = dataframe.replace({'prediction': p_class})
return dataframe
# # Dealing with DATA
pd.set_option('display.max_columns', 100)
pd.options.display.max_colwidth = 1000
data = read_data('docket_data8.csv')
# +
# db = read_data('JDreview.csv')
# +
# db.rename(columns={'type': 'bundle_type'}, inplace=True)
# -
# db = db.drop(['clean_text','motionOrsummary','bundle_action_prediction','bundle_decision_prediction','Output Review','Corrected bundle_action_prediction','Corrected bundle_decision_prediction','Notes','Column3'],axis=1)
data.groupby('nature_of_suit').count()
data.groupby('bundle_action').count()
k_class = {'granted':'Grant','denied':'Deny','partial':'Partial'}
data = data.replace({'bundle_action': k_class})
# data = da.append(db, ignore_index=True)
data = data.drop_duplicates(['case_number', 'activity_number','bundle_motion_number','docket_text'], keep='last')
data.to_csv("week15_data.csv")
len(data)
len(data)
# # Word2Vec
# +
start_time = time.time()
data = read_data('week15_data.csv')
cleantext(data,stem = False)
# tokenizetext(df)
# doc = get_list(df)
doc = data.clean_text
w2c = w2c_model(doc,size=100,window=5,min_count=1,workers=6,iter=10,sg=0,length=len(doc),numepoch=10)
runtime = time.time() - start_time
a = runtime/60
print("Done with w2c model!")
print("It took",a, "mins" )
# +
# runtime = time.time() - start_time
# runtime/60
# +
# print (da.clean_text.apply(lambda x: 'dllm' in x.lower()).sum())
# -
data = read_data('week13_data.csv')
# +
# #Cleaning text
# da["clean_text"] = da["docket_text"]
# #Get bundle number
# da['bundle_motion_number'] = da['bundle_motion_number'].apply(lambda x: np.nan if x == 'None' else float(x))
# #Replace bundle motion number
# da.clean_text = da.apply(lambda x: x.clean_text if np.isnan(x.bundle_motion_number)
# else x.clean_text.replace(str(int(x.bundle_motion_number)), 'DLLM'), axis = 1)
# -
# +
# for i in range (int((len(data)/ batch_size))):
# data_batch = data[data['case_number'].isin(cases[i * batch_size: (i + 1) * batch_size])].copy()
# -
# # Train Model (Bundle Type)
# +
# ds = read_data('train.csv')
# d1 = ds[ds['type'] != 'None']
# d2 = ds[ds['type'] == 'None'].sample(6000)
# ds = d1.append(d2, ignore_index=True)
# ds = ds.sample(frac=1)
# cleantext(ds,stem=False)
# X, y = prepro(ds,w2c,seq_len=100, emb_size=100,target_col='type')
# CNN_model= fit_model(X=X,y=y,weightsfile = "weights.best2.hdf5",num_labels=4,seq_len=100,emb_size=100,epochs=20)
# print("Done with training!")
# +
# df = read_data('week12_data.csv')
# -
df = data[data['docket_text'].apply(lambda x: any(y in str(x).lower() for y in ['summary', 'order','dismiss']))]
# cleantext(df,stem = False)
df.groupby('bundle_type').count()
len(df)
d1 = df[df['bundle_type'] != 'None']
d2 = df[df['bundle_type'] == 'None'].sample(12000)
df = d1.append(d2, ignore_index=True)
df = df.sample(frac=1)
df.groupby('bundle_type').count()
# +
# k_class = {'Transfer-Remand':'TransferRemand','Admin Closing':'AdminClosing','SJ Submission':'SJSubmission'}
# df = df.replace({'bundle_type': k_class})
# -
X, y = prepro(df,w2c,seq_len=100, emb_size=100,target_col='bundle_type')
# +
start_time = time.time()
CNN_model= fit_model(X=X,y=y,weightsfile = "bundle_type.hdf5",num_labels=4,seq_len=100,emb_size=100,epochs=20)
runtime = time.time() - start_time
a = runtime/60
print("Done with training!")
print("It took",a, "mins" )
# -
# # Training Model (MTD Type)
dt = read_data('prediction.csv')
len(dt)
dt.groupby('mtd_type').count()
d1 = dt[dt['mtd_type'] != 'None']
d2 = dt[dt['mtd_type'] == 'None'].sample(14000)
dt = d1.append(d2, ignore_index=True)
dt = dt.sample(frac=1)
len(dt)
X, y = prepro(dt,w2c,seq_len=100, emb_size=100,target_col='mtd_type')
# +
start_time = time.time()
CNN_model= fit_model(X=X,y=y,weightsfile = "mtd_type.hdf5",num_labels=4,seq_len=100,emb_size=100,epochs=20)
runtime = time.time() - start_time
a = runtime/60
print("Done with training!")
print("It took",a, "mins" )
# -
# # Training Model (Bundle_action)
df = read_data('prediction.csv')
len(ds)
# +
# data['docket_text'] = data.apply(lambda x: x.docket_text.replace(str(x.bundle_motion_number), ' dllm '), axis=1)
# -
ds = df[(df['bundle_type']=='Order')]
# +
d1 = ds[ds['bundle_action'] != 'None']
d2 = ds[ds['bundle_action'] == 'None'].sample(1900)
ds = d1.append(d2, ignore_index=True)
# -
ds = ds.sample(frac=1)
ds.groupby('bundle_action').count()
X, y = prepro(ds,w2c,seq_len=100, emb_size=100,target_col='bundle_action')
CNN_model= fit_model(X=X,y=y,weightsfile = "bundle_action.hdf5",num_labels=4,seq_len=100,emb_size=100,epochs=20)
# # Training Model (MTD Action)
df = read_data('prediction.csv')
ds = df[(df['mtd_type']=='Order')]
len(ds)
ds.groupby('mtd_action').count()
d1 = ds[ds['mtd_action'] != 'Grant']
d2 = ds[ds['mtd_action'] == 'Grant'].sample(6900)
ds = d1.append(d2, ignore_index=True)
ds = ds.sample(frac=1)
X, y = prepro(ds,w2c,seq_len=100, emb_size=100,target_col='mtd_action')
CNN_model= fit_model(X=X,y=y,weightsfile = "mtd_action.hdf5",num_labels=5,seq_len=100,emb_size=100,epochs=20)
# # Training Model (Filing_Party)
df = read_data('prediction.csv')
ds = df[df['bundle_type'] != 'None']
ds.groupby('bundle_who_filed').count()
len(ds)
d1 = ds[ds['bundle_who_filed'] != 'None']
d2 = ds[ds['bundle_who_filed'] == 'None'].sample(3000)
ds = d1.append(d2, ignore_index=True)
ds = ds.sample(frac=1)
X, y = prepro(ds,w2c,seq_len=100, emb_size=100,target_col='bundle_who_filed')
CNN_model= fit_model(X=X,y=y,weightsfile = "bundle_filed.hdf5",num_labels=4,seq_len=100,emb_size=100,epochs=20)
# # Prediction
# +
# db = read_data('bundle_pt1.csv')
# get_type(db)
# db = db[db['docket_text'].apply(lambda x: any(y in str(x).lower() for y in ['summary', 'order']))]
# cleantext(db,stem=False)
# X1, y1 = prepro(db,w2c,seq_len=100, emb_size=100,target_col='type')
# preds,lst = prediction(model=CNN_model,weightsfile = "weights.best2.hdf5",X = X1)
# db = assign_label(db)
# runtime = time.time() - start_time
# print('It took',(runtime/60),'minutes to run!')
# print('Done with prediction!')
# -
# +
# db = read_data('week12_data.csv')
da = data[data['docket_text'].apply(lambda x: any(y in str(x).lower() for y in ['summary', 'order','dismiss']))]
# cleantext(da,stem = False)
# -
len(da)
# +
# da.sample(frac = 0.5, random_state = 0)
# -
da = da.reset_index(drop=True)
# +
def batch_predict(data, batch_size):
# get unique cases
cases = data.case_number.unique()
# Create new data file
new_data = pd.DataFrame()
for i in range (int(len(cases) / batch_size)):
# slide through the cases list with a slide size of batch_size
batch_cases = cases[i * batch_size: (i + 1) * batch_size]
#create a temp data frame of the data with just the cases
d1 = data[data['case_number'].isin(batch_cases)].copy()
X1, y1 = prepro(d1,w2c,seq_len=100, emb_size=100,target_col='bundle_type')
# Modify this next line to make it work with your code
pred, d1['predicted_label'] = prediction(model=CNN_model,weightsfile = "bundle_type.hdf5",X = X1)
# Append predicted data to the new dataframe
new_data = new_data.append(d1, ignore_index=True)
return new_data
# +
# X1, y1 = prepro(da,w2c,seq_len=100, emb_size=100,target_col='bundle_type')
new_data = batch_predict(da, batch_size= 50)
# preds, lst = prediction(model=CNN_model,weightsfile = "weights.best2.hdf5",X = X1)
# -
len(new_data)
len(da)
# +
# d1 = db[db['bundle_type'] != 'None']
# d2 = db[db['bundle_type'] == 'None'].sample(60000)
# db = d1.append(d2, ignore_index=True)
# db = db.sample(frac=1)
# -
from tensorflow.python.client import device_lib
# print(device_lib.list_local_devices())
# +
# da['type_prediction']= lst
# -
values=sorted(list(set(new_data['bundle_type'])))
values
p_class = {0:'Motion',1:'None',2:'Order',3:'RR'}
new_data = new_data.replace({'predicted_label': p_class})
new_data.groupby('predicted_label').count()
new_data.groupby('bundle_type').count()
new_data.to_csv("prediction.csv")
# # MTD Prediction
db = read_data('prediction.csv')
# +
def batch_predict_mtd(data, batch_size):
# get unique cases
cases = data.case_number.unique()
# Create new data file
new_data = pd.DataFrame()
for i in range (int(len(cases) / batch_size)):
# slide through the cases list with a slide size of batch_size
batch_cases = cases[i * batch_size: (i + 1) * batch_size]
#create a temp data frame of the data with just the cases
d1 = data[data['case_number'].isin(batch_cases)].copy()
X1, y1 = prepro(d1,w2c,seq_len=100, emb_size=100,target_col='mtd_type')
# Modify this next line to make it work with your code
pred, d1['mtd_predicted'] = prediction(model=CNN_model,weightsfile = "mtd_type.hdf5",X = X1)
# Append predicted data to the new dataframe
new_data = new_data.append(d1, ignore_index=True)
return new_data
# -
len(db)
mtd_data = batch_predict_mtd(db, batch_size= 50)
print('done')
len(mtd_data)
values=sorted(list(set(mtd_data['mtd_type'])))
values
p_class = {0:'Motion',1:'None',2:'Order', 3:'RR'}
mtd_data = mtd_data.replace({'mtd_predicted': p_class})
mtd_data.to_csv("prediction.csv")
mtd_data.groupby('mtd_type').count()
mtd_data.groupby('mtd_predicted').count()
# # Prediction (Bundle_action)
# +
db = read_data('prediction.csv')
# db = db[(db['bundle_type']=='Order')]
# db = db[(db['predicted_label']=='Order')]
# -
f = db['predicted_label']=='Order'
d1 = db[f].copy()
db = db[~f]
db['bundle_action_prediction'] = db.apply(lambda x: np.nan, axis=1)
# +
X1, y1 = prepro(d1,w2c,seq_len=100, emb_size=100,target_col='bundle_action')
preds,lst = prediction(model=CNN_model,weightsfile = "bundle_action.hdf5",X = X1)
# +
# np.mean([np.argmax(x[0]) == np.argmax(x[1]) for x in zip(preds, y1)])
# -
d1['bundle_action_prediction']= lst
values=sorted(list(set(d1['bundle_action'])))
values
# db['bundle_action_prediction']= lst2
p_class = {0:'Deny',1:'Grant',2:'None',3:'Partial'}
d1 = d1.replace({'bundle_action_prediction': p_class})
db = db.append(d1, ignore_index=True)
len(db)
db.groupby('bundle_action').count()
db.groupby('bundle_action_prediction').count()
db.to_csv('prediction.csv')
# # Prediction (MTD Action)
# +
dd = read_data('prediction.csv')
# dd = dd[(dd['mtd_type']=='Order')]
# dd = dd[(dd['mtd_predicted']=='Order')]
# +
h = dd['mtd_type']=='Order'
d3 = dd[h].copy()
dd = dd[~h]
dd['mtd_action_prediction'] = dd.apply(lambda x: np.nan, axis=1)
# +
X1, y1 = prepro(d3,w2c,seq_len=100, emb_size=100,target_col='mtd_action')
preds,lst = prediction(model=CNN_model,weightsfile = "mtd_action.hdf5",X = X1)
# +
d3['mtd_action_prediction']= lst
values=sorted(list(set(d3['mtd_action'])))
values
# -
p_class = {0:'Deny',1:'Grant',2:'Moot',3:'None',4:'Partial'}
d3 = d3.replace({'mtd_action_prediction': p_class})
dd = dd.append(d3, ignore_index=True)
dd.to_csv('prediction.csv')
dd.groupby('mtd_action').count()
dd.groupby('mtd_action_prediction').count()
# # Prediction (Filed_party)
# +
dc = read_data('prediction.csv')
# dc = dc[(dc['bundle_type']!='None')]
# dc = dc[(dc['predicted_label']!='None')]
# +
g = dc['predicted_label']!='None'
d2 = dc[g].copy()
dc = dc[~g]
dc['filed_prediction'] = dc.apply(lambda x: np.nan, axis=1)
# +
X1, y1 = prepro(d2,w2c,seq_len=100, emb_size=100,target_col='bundle_who_filed')
preds,lst = prediction(model=CNN_model,weightsfile = "bundle_filed.hdf5",X = X1)
# -
d2['filed_prediction']= lst
values=sorted(list(set(d2['bundle_who_filed'])))
values
p_class = {0:'None',1:'X',2:'defendant',3:'plaintiff'}
d2 = d2.replace({'filed_prediction': p_class})
dc = dc.append(d2, ignore_index=True)
dc.to_csv('prediction.csv')
dc.groupby('bundle_who_filed').count()
dc.groupby('filed_prediction').count()
# +
# db.to_csv("filed_prediction.csv")
# -
# # Accuracy, Precision, and Recall
from sklearn.metrics import classification_report
print(classification_report(new_data["bundle_type"], new_data["predicted_label"]))
from sklearn.metrics import accuracy_score
accuracy_score(new_data["bundle_type"], new_data["predicted_label"])*100
from sklearn.metrics import classification_report
print(classification_report(mtd_data["mtd_type"], mtd_data["mtd_predicted"]))
from sklearn.metrics import accuracy_score
accuracy_score(mtd_data["mtd_type"], mtd_data["mtd_predicted"])*100
# +
from sklearn.metrics import classification_report
print(classification_report(d1["bundle_action"], d1["bundle_action_prediction"]))
# +
from sklearn.metrics import accuracy_score
accuracy_score(d1["bundle_action"], d1["bundle_action_prediction"])*100
# -
print(classification_report(dd["mtd_action"], dd["mtd_action_prediction"]))
accuracy_score(dd["mtd_action"], dd["mtd_action_prediction"])*100
print(classification_report(d2["bundle_who_filed"], d2["filed_prediction"]))
accuracy_score(d2["bundle_who_filed"], d2["filed_prediction"])*100
| MSFJ&MTD_Mark_3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Fire up graphlab create
import graphlab
# # Load some house value vs. crime rate data
#
# Dataset is from Philadelphia, PA and includes average house sales price in a number of neighborhoods. The attributes of each neighborhood we have include the crime rate ('CrimeRate'), miles from Center City ('MilesPhila'), town name ('Name'), and county name ('County').
sales = graphlab.SFrame.read_csv('Philadelphia_Crime_Rate_noNA.csv/')
sales
# # Exploring the data
# The house price in a town is correlated with the crime rate of that town. Low crime towns tend to be associated with higher house prices and vice versa.
graphlab.canvas.set_target('ipynb')
sales.show(view="Scatter Plot", x="CrimeRate", y="HousePrice")
# # Fit the regression model using crime as the feature
crime_model = graphlab.linear_regression.create(sales, target='HousePrice', features=['CrimeRate'],validation_set=None,verbose=False)
# # Let's see what our fit looks like
# Matplotlib is a Python plotting library that is also useful for plotting. You can install it with:
#
# 'pip install matplotlib'
import matplotlib.pyplot as plt
# %matplotlib inline
plt.plot(sales['CrimeRate'],sales['HousePrice'],'.',
sales['CrimeRate'],crime_model.predict(sales),'-')
# Above: blue dots are original data, green line is the fit from the simple regression.
# # Remove Center City and redo the analysis
# Center City is the one observation with an extremely high crime rate, yet house prices are not very low. This point does not follow the trend of the rest of the data very well. A question is how much including Center City is influencing our fit on the other datapoints. Let's remove this datapoint and see what happens.
sales_noCC = sales[sales['MilesPhila'] != 0.0]
sales_noCC.show(view="Scatter Plot", x="CrimeRate", y="HousePrice")
# ### Refit our simple regression model on this modified dataset:
crime_model_noCC = graphlab.linear_regression.create(sales_noCC, target='HousePrice', features=['CrimeRate'],validation_set=None, verbose=False)
# ### Look at the fit:
plt.plot(sales_noCC['CrimeRate'],sales_noCC['HousePrice'],'.',
sales_noCC['CrimeRate'],crime_model.predict(sales_noCC),'-')
# # Compare coefficients for full-data fit versus no-Center-City fit
# Visually, the fit seems different, but let's quantify this by examining the estimated coefficients of our original fit and that of the modified dataset with Center City removed.
crime_model.get('coefficients')
crime_model_noCC.get('coefficients')
# Above: We see that for the "no Center City" version, per unit increase in crime, the predicted decrease in house prices is 2,287. In contrast, for the original dataset, the drop is only 576 per unit increase in crime. This is significantly different!
# ###High leverage points:
# Center City is said to be a "high leverage" point because it is at an extreme x value where there are not other observations. As a result, recalling the closed-form solution for simple regression, this point has the *potential* to dramatically change the least squares line since the center of x mass is heavily influenced by this one point and the least squares line will try to fit close to that outlying (in x) point. If a high leverage point follows the trend of the other data, this might not have much effect. On the other hand, if this point somehow differs, it can be strongly influential in the resulting fit.
#
# ###Influential observations:
# An influential observation is one where the removal of the point significantly changes the fit. As discussed above, high leverage points are good candidates for being influential observations, but need not be. Other observations that are *not* leverage points can also be influential observations (e.g., strongly outlying in y even if x is a typical value).
# # Remove high-value outlier neighborhoods and redo analysis
# Based on the discussion above, a question is whether the outlying high-value towns are strongly influencing the fit. Let's remove them and see what happens.
sales_nohighend = sales_noCC[sales_noCC['HousePrice'] < 350000]
crime_model_nohighend = graphlab.linear_regression.create(sales_nohighend, target='HousePrice', features=['CrimeRate'],validation_set=None, verbose=False)
# ### Do the coefficients change much?
crime_model_noCC.get('coefficients')
crime_model_nohighend.get('coefficients')
# Above: We see that removing the outlying high-value neighborhoods has *some* effect on the fit, but not nearly as much as our high-leverage Center City datapoint.
| 2- Regression/projects/week 1/.ipynb_checkpoints/PhillyCrime-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Kubeflow pipelines
#
# **Learning Objectives:**
# 1. Learn how to deploy a Kubeflow cluster on GCP
# 1. Learn how to use the notebook server on Kubeflow
# 1. Learn how to create a experiment in Kubeflow
# 1. Learn how to package you code into a Kubeflow pipeline
# 1. Learn how to run a Kubeflow pipeline in a repeatable and traceable way
#
#
# ## Introduction
#
# In this notebook, we will first setup a Kubeflow cluster on GCP, and then launch a Kubeflow Notebook Server from where we will run this notebook. This will allow us to pilote the Kubeflow cluster from the notebook. Then, we will create a Kubeflow experiment and a Kubflow pipeline from our taxifare machine learning code. At last, we will run the pipeline on the Kubeflow cluster, providing us with a reproducible and traceable way to execute machine learning code.
# +
from os import path
import kfp
import kfp.compiler as compiler
import kfp.components as comp
import kfp.dsl as dsl
import kfp.gcp as gcp
import kfp.notebook
# -
# ## Setup a Kubeflow cluster on GCP
# **TODO 1**
# To deploy a [Kubeflow](https://www.kubeflow.org/) cluster
# in your GCP project, use the [Kubeflow cluster deployer](https://deploy.kubeflow.cloud/#/deploy).
#
# There is a [setup video](https://www.kubeflow.org/docs/started/cloud/getting-started-gke/~) that will
# take you over all the steps in details, and explains how to access to the Kubeflow Dashboard UI, once it is
# running.
#
# You'll need to create an OAuth client for authentication purposes: Follow the
# instructions [here](https://www.kubeflow.org/docs/gke/deploy/oauth-setup/).
# ## Launch a Jupyter notebook server on the Kubeflow cluster
# **TODO 2**
# A Kubeflow cluster allows you not only to run Kubeflow pipelines, but it also allows you to launch a Jupyter notebook server from which you can pilote the Kubeflow cluster. In particular, you can create experiments, define and run pipelines from whithin notebooks running on that Jupiter notebook server. This is exactly what we are going to do in this notebook.
#
# First of all, click on the "Notebook Sever" tab in the Kubeflow Dashboard UI, and create a Notebook Server. Once it's ready connect to it.
#
# Since the goal is to run this notebook on that Kubeflow Notebook Server, first create new notebook and clone the training-data-analysis repo by running the following command in a cell and then naviguating to this notebook:
# ```bash
# $ git clone -b ml_on_gcp-kubeflow_pipelines --single-branch https://github.com/GoogleCloudPlatform/training-data-analyst.git
#
# ```
# ## Create an experiment
# **TODO 3**
# From now on, you should be running this notebook from the Notebook Server from the Kubeflow cluster you created on your GCP project.
# We will start by creating a Kubeflow client to pilote the Kubeflow cluster:
client = kfp.Client()
# Let's look at the experiments that are running on this cluster. Since you just launched it, you should see only a single "Default" experiment:
client.list_experiments()
# Now let's create a 'taxifare' experiment where we could look at all the various runs of our taxifare pipeline:
exp = client.create_experiment(name='taxifare')
# Let's make sure the experiment has been created correctly:
client.list_experiments()
# ## Packaging you code into Kubeflow components
# We have packaged our taxifare ml pipeline into three components:
# * `./components/bq2gcs` that creates the training and evaluation data from BigQuery and exports it to GCS
# * `./components/trainjob` that launches the training container on AI-platform and exports the model
# * `./components/deploymodel` that deploys the trained model to AI-platform as a REST API
#
# Each of these components has been wrapped into a Docker container, in the same way we did with the taxifare training code in the previous lab.
#
# If you inspect the code in these folders, you'll notice that the `main.py` or `main.sh` files contain the code we previously executed in the notebooks (loading the data to GCS from BQ, or launching a training job to AI-platform, etc.). The last line in the `Dockerfile` tells you that these files are executed when the container is run.
# So we just packaged our ml code into light container images for reproducibility.
#
# We have made it simple for you to build the container images and push them to the Google Cloud image registry gcr.io in your project: just type `make` in the pipelines directory! However, you can't do that from a Kubeflow notebook because Docker is not installed there. So you'll have to do that from Cloud Shell.
#
# For that, open Cloud Shell, and clone this repo there. Then cd to the pipelines subfolder:
#
# ```bash
# $ git clone -b ml_on_gcp-kubeflow_pipelines --single-branch https://github.com/GoogleCloudPlatform/training-data-analyst.git
#
# $ cd training-data-analyst/courses/machine_learning/production_ml_systems/pipelines/
# ```
#
# Then run `make` to build and push the images.
# Now that the container images are pushed to the regsitry in your project, we need to create yaml files describing to Kubeflow how to use these containers. It boils down essentially
# * describing what arguments Kubeflow needs to pass to the containers when it runs them
# * telling Kubeflow where to fetch the corresponding Docker images
#
# In the cells below, we have three of these "Kubeflow component description files", one for each of our components.
#
# For each of these, correct the image URI to reflect that you pushed the images into the gcr.io associated with your project:
# **TODO 4**
# +
# %%writefile bq2gcs.yaml
name: bq2gcs
description: |
This component creates the training and
validation datasets as BiqQuery tables and export
them into a Google Cloud Storage bucket at
gs://<BUCKET>/taxifare/data.
inputs:
- {name: Input Bucket , type: String, description: 'GCS directory path.'}
implementation:
container:
image: gcr.io/PROJECT/taxifare-bq2gcs
args: ["--bucket", {inputValue: Input Bucket}]
# +
# %%writefile trainjob.yaml
name: trainjob
description: |
This component trains a model to predict that taxi fare in NY.
It takes as argument a GCS bucket and expects its training and
eval data to be at gs://<BUCKET>/taxifare/data/ and will export
the trained model at gs://<BUCKET>/taxifare/model/.
inputs:
- {name: Input Bucket , type: String, description: 'GCS directory path.'}
implementation:
container:
image: gcr.io/PROJECT/taxifare-trainjob
args: [{inputValue: Input Bucket}]
# +
# %%writefile deploymodel.yaml
name: deploymodel
description: |
This component deploys a trained taxifare model on GCP as taxifare:dnn.
It takes as argument a GCS bucket and expects the model to deploy
to be found at gs://<BUCKET>/taxifare/model/export/savedmodel/
inputs:
- {name: Input Bucket , type: String, description: 'GCS directory path.'}
implementation:
container:
image: gcr.io/PROJECT/taxifare-deploymodel
args: [{inputValue: Input Bucket}]
# -
# ## Create a Kubeflow pipeline
# The code below creates a kubeflow pipeline by decorating a regular fuction with the
# `@dsl.pipeline` decorator. Now the arguments of this decorated function will be
# the input parameters of the Kubeflow pipeline.
#
# Inside the function, we describe the pipeline by
# * loading the yaml component files we created above into a Kubeflow op
# * specifying the order into which the Kubeflow ops should be run
# +
# TODO 4
PIPELINE_TAR = 'taxifare.tar.gz'
BQ2GCS_YAML = './bq2gcs.yaml'
TRAINJOB_YAML = './trainjob.yaml'
DEPLOYMODEL_YAML = './deploymodel.yaml'
@dsl.pipeline(
name='Taxifare',
description='Train a ml model to predict the taxi fare in NY')
def pipeline(gcs_bucket_name='<bucket where data and model will be exported>'):
bq2gcs_op = comp.load_component_from_file(BQ2GCS_YAML)
bq2gcs = bq2gcs_op(
input_bucket=gcs_bucket_name,
).apply(gcp.use_gcp_secret('user-gcp-sa'))
trainjob_op = comp.load_component_from_file(TRAINJOB_YAML)
trainjob = trainjob_op(
input_bucket=gcs_bucket_name,
).apply(gcp.use_gcp_secret('user-gcp-sa'))
deploymodel_op = comp.load_component_from_file(DEPLOYMODEL_YAML)
deploymodel = deploymodel_op(
input_bucket=gcs_bucket_name,
).apply(gcp.use_gcp_secret('user-gcp-sa'))
trainjob.after(bq2gcs)
deploymodel.after(trainjob)
# -
# The pipeline function above is then used by the Kubeflow compiler to create a Kubeflow pipeline artifact that can be either uploaded to the Kubeflow cluster from the UI, or programatically, as we will do below:
compiler.Compiler().compile(pipeline, PIPELINE_TAR)
# ls $PIPELINE_TAR
# If you untar and uzip this pipeline artifact, you'll see that the compiler has transformed the
# Python description of the pipeline into yaml description!
#
# Now let's feed Kubeflow with our pipeline and run it using our client:
# TODO 5
run = client.run_pipeline(
experiment_id=exp.id,
job_name='taxifare',
pipeline_package_path='taxifare.tar.gz',
params={
'gcs-bucket-name': "dherin-sandbox",
},
)
# Have a look at the link to monitor the run.
# Now all the runs are nicely organized under the experiment in the UI, and new runs can be either manually launched or scheduled through the UI in a completely repeatable and traceable way!
| courses/machine_learning/deepdive2/building_production_ml_systems/solutions/3_kubeflow_pipelines.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Import packages
import pandas as pd
import numpy as np
import os
from bs4 import BeautifulSoup
import time
import requests
import nfl_data_py as nfl
import matplotlib.pyplot as plt
import plotly
import plotly.express as px
import random
import scipy
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score
from sklearn.metrics import mean_squared_error
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import RandomForestClassifier
from sklearn.tree import DecisionTreeClassifier
import xgboost as xgb
import pickle
from joblib import dump, load
import warnings
# Set certain settings for the notebook
pd.set_option("display.max_columns", None)
warnings.filterwarnings("ignore")
# -
# Check current directory
os.getcwd()
# +
# Read in data files
betting_lines = pd.read_csv('../data/betting_data_final')
game_spreads = pd.read_csv('../data/games_with_spread_preds.csv')
# +
# Function to get returns from over/under bets
def calculate_return_using_ml(line, wager, is_win, return_includes_wager=False):
if return_includes_wager:
add_on = wager
else:
add_on = 0
if is_win == 'Win':
if line > 0:
return np.round(((line/100)*wager + add_on), 2)
else:
return np.round(((100/abs(line))*wager + add_on), 2)
elif is_win == 'Push':
return add_on
else:
return np.round(-wager, 2)
def get_return_over_under_100(game, bet_over=True):
if bet_over:
if game['Open Over/Under Result'] == 'Over':
result = 'Win'
elif game['Open Over/Under Result'] == 'Under':
result = 'Loss'
else:
result = 'Push'
if not bet_over:
if game['Open Over/Under Result'] == 'Over':
result = 'Loss'
elif game['Open Over/Under Result'] == 'Under':
result = 'Win'
else:
result = 'Push'
return calculate_return_using_ml(game['Total Score Over ML Open'], 100, is_win=result)
# +
gambling_df = betting_lines.merge(game_spreads, left_on=['season', 'week', 'Home Team', 'Away Team'],
right_on=['season', 'week', 'home_x', 'away_x'])
gambling_df['over_bet_return'] = gambling_df.apply(lambda x: get_return_over_under_100(x, bet_over=True), axis=1)
gambling_df['under_bet_return'] = gambling_df.apply(lambda x: get_return_over_under_100(x, bet_over=False), axis=1)
gambling_df['home_line_diff_algo'] = gambling_df.apply(lambda x: x['Home Line Open'] - x.home_spread_algo, axis=1)
gambling_df['away_line_diff_algo'] = gambling_df.apply(lambda x: x['Away Line Open'] - x.away_spread_algo, axis=1)
gambling_df['home_line_diff_elo'] = gambling_df.apply(lambda x: x['Home Line Open'] - x.home_spread_elo, axis=1)
gambling_df['away_line_diff_elo'] = gambling_df.apply(lambda x: x['Away Line Open'] - x.away_spread_elo, axis=1)
gambling_df['total_score_diff_algo'] = gambling_df.apply(lambda x: x.total_score_pred - x['Total Score Open'], axis=1)
gambling_df
# +
# Functions to get point spread and over under returns
# Going to do this in a way similar to the moneyline bets, first doing flat 100 dollar bets and then checking variable bets,
# and comparing the results to the elo algorithm as well as the random betting simulations
def get_best_spread_advantage_algo(game):
if game.home_line_diff_algo > game.away_line_diff_algo:
return game.home_line_diff_algo, 'Home'
else:
return game.away_line_diff_algo, 'Away'
def get_best_spread_advantage_elo(game):
if game.home_line_diff_elo > game.away_line_diff_elo:
return game.home_line_diff_elo, 'Home'
else:
return game.away_line_diff_elo, 'Away'
def get_return_100_spread_algo(game, threshold):
bet_weight = 0
if game.best_spread_advantage_algo >= threshold:
bet_weight = 1
return bet_weight * game['{} Spread Bet Return'.format(game.spread_advantage_team_algo)]
def get_return_100_spread_elo(game, threshold):
bet_weight = 0
if game.best_spread_advantage_elo >= threshold:
bet_weight = 1
return bet_weight * game['{} Spread Bet Return'.format(game.spread_advantage_team_elo)]
def get_return_100_spread_random(game):
if random.random() <= .5:
return game['Home Spread Bet Return']
else:
return game['Away Spread Bet Return']
def get_return_proportional_spread_algo(game):
bet_weight = 0
if 3 <= game.best_spread_advantage_algo < 4:
bet_weight = 1
elif 4 <= game.best_spread_advantage_algo < 5:
bet_weight = 2
elif 5 <= game.best_spread_advantage_algo < 6:
bet_weight = 3
elif 6 <= game.best_spread_advantage_algo < 7:
bet_weight = 4
elif game.best_spread_advantage_algo >= 7 :
bet_weight = 5
if game.spread_advantage_team_algo == 'home':
return bet_weight * game['Home Spread Bet Return'], bet_weight * 100
else:
return bet_weight * game['Away Spread Bet Return'], bet_weight * 100
def get_return_proportional_spread_elo(game):
bet_weight = 0
if 3 <= game.best_spread_advantage_elo < 4:
bet_weight = 1
elif 4 <= game.best_spread_advantage_elo < 5:
bet_weight = 2
elif 5 <= game.best_spread_advantage_elo < 6:
bet_weight = 3
elif 6 <= game.best_spread_advantage_elo < 7:
bet_weight = 4
elif game.best_spread_advantage_elo >= 7 :
bet_weight = 5
if game.spread_advantage_team_elo == 'home':
return bet_weight * game['Home Spread Bet Return'], bet_weight * 100
else:
return bet_weight * game['Away Spread Bet Return'], bet_weight * 100
def get_return_prop_spread_random(game):
bet_weight = random.choice([1, 2, 3, 4, 5])
if random.random() <= .5:
return bet_weight * game['Home Spread Bet Return'], bet_weight*100
else:
return bet_weight * game['Away Spread Bet Return'], bet_weight*100
def get_best_ou_advantage_algo(game):
if game.total_score_diff_algo > 0:
return 'over'
else:
return 'under'
def get_return_100_ou_algo(game, threshold):
bet_weight = 0
if abs(game.total_score_diff_algo) >= threshold:
bet_weight = 1
return bet_weight * game['{}_bet_return'.format(game.ou_advantage)]
def get_return_proportional_ou_algo(game):
bet_weight = 0
if 1 <= abs(game.total_score_diff_algo) < 2:
bet_weight = 1
elif 2 <= abs(game.total_score_diff_algo) < 3:
bet_weight = 2
elif 3 <= abs(game.total_score_diff_algo) < 4:
bet_weight = 3
elif 4 <= abs(game.total_score_diff_algo) < 5:
bet_weight = 4
elif abs(game.total_score_diff_algo) >= 5 :
bet_weight = 5
return bet_weight * game['{}_bet_return'.format(game.ou_advantage)], bet_weight*100
def get_return_100_ou_random(game):
if random.random() <= .5:
return game['over_bet_return']
else:
return game['under_bet_return']
def get_return_prop_ou_random(game):
bet_weight = random.choice([1, 2, 3, 4, 5])
if random.random() <= .5:
return bet_weight * game['over_bet_return'], bet_weight*100
else:
return bet_weight * game['under_bet_return'], bet_weight*100
# -
gambling_df[['best_spread_advantage_algo', 'spread_advantage_team_algo']] = gambling_df.apply(lambda x: get_best_spread_advantage_algo(x), axis=1, result_type='expand')
gambling_df[['best_spread_advantage_elo', 'spread_advantage_team_elo']] = gambling_df.apply(lambda x: get_best_spread_advantage_elo(x), axis=1, result_type='expand')
gambling_df['ou_advantage'] = gambling_df.apply(lambda x: get_best_ou_advantage_algo(x), axis=1)
gambling_df
# +
# Using spread advantage of one point
bets_100_01_ps_algo = gambling_df.copy()
bets_100_01_ps_algo['returns'] = bets_100_01_ps_algo.apply(lambda x: get_return_100_spread_algo(x, 1), axis=1)
bets_100_01_ps_algo.groupby(by=['week']).sum()['returns'].reset_index().plot.bar(x='week', y='returns')
# -
bets_100_01_ps_algo.sum()['returns']
len(bets_100_01_ps_algo)
len(bets_100_01_ps_algo[bets_100_01_ps_algo.returns != 0])
# +
# Using spread advantage of two points
bets_100_02_ps_algo = gambling_df.copy()
bets_100_02_ps_algo['returns'] = bets_100_02_ps_algo.apply(lambda x: get_return_100_spread_algo(x, 2), axis=1)
bets_100_02_ps_algo.groupby(by=['week']).sum()['returns'].reset_index().plot.bar(x='week', y='returns')
# -
bets_100_02_ps_algo.sum()['returns']
len(bets_100_02_ps_algo[bets_100_02_ps_algo.returns != 0])
# +
# Using spread advantage of three points
bets_100_03_ps_algo = gambling_df.copy()
bets_100_03_ps_algo['returns'] = bets_100_03_ps_algo.apply(lambda x: get_return_100_spread_algo(x, 3), axis=1)
bets_100_03_ps_algo.groupby(by=['week']).sum()['returns'].reset_index().plot.bar(x='week', y='returns')
# -
bets_100_03_ps_algo.sum()['returns']
len(bets_100_03_ps_algo[bets_100_03_ps_algo.returns != 0])
# +
# Using spread advantage of four points
bets_100_04_ps_algo = gambling_df.copy()
bets_100_04_ps_algo['returns'] = bets_100_04_ps_algo.apply(lambda x: get_return_100_spread_algo(x, 4), axis=1)
bets_100_04_ps_algo.groupby(by=['week']).sum()['returns'].reset_index().plot.bar(x='week', y='returns')
# -
bets_100_04_ps_algo.sum()['returns']
len(bets_100_04_ps_algo[bets_100_04_ps_algo.returns != 0])
# +
# Using spread advantage of five points
bets_100_05_ps_algo = gambling_df.copy()
bets_100_05_ps_algo['returns'] = bets_100_05_ps_algo.apply(lambda x: get_return_100_spread_algo(x, 5), axis=1)
bets_100_05_ps_algo.groupby(by=['week']).sum()['returns'].reset_index().plot.bar(x='week', y='returns')
# -
bets_100_05_ps_algo.sum()['returns']
len(bets_100_05_ps_algo[bets_100_05_ps_algo.returns != 0])
# +
# Look at all of the above, but for elo model
# Using spread advantage of one point
bets_100_01_ps_elo = gambling_df.copy()
bets_100_01_ps_elo['returns'] = bets_100_01_ps_elo.apply(lambda x: get_return_100_spread_elo(x, 1), axis=1)
bets_100_01_ps_elo.groupby(by=['week']).sum()['returns'].reset_index().plot.bar(x='week', y='returns')
# -
bets_100_01_ps_elo.sum()['returns']
len(bets_100_01_ps_elo[bets_100_01_ps_elo.returns != 0])
# +
# Look at all of the above, but for elo model
# Using spread advantage of two point
bets_100_02_ps_elo = gambling_df.copy()
bets_100_02_ps_elo['returns'] = bets_100_02_ps_elo.apply(lambda x: get_return_100_spread_elo(x, 2), axis=1)
bets_100_02_ps_elo.groupby(by=['week']).sum()['returns'].reset_index().plot.bar(x='week', y='returns')
# -
bets_100_02_ps_elo.sum()['returns']
len(bets_100_02_ps_elo[bets_100_02_ps_elo.returns != 0])
# +
# Look at all of the above, but for elo model
# Using spread advantage of three points
bets_100_03_ps_elo = gambling_df.copy()
bets_100_03_ps_elo['returns'] = bets_100_03_ps_elo.apply(lambda x: get_return_100_spread_elo(x, 3), axis=1)
bets_100_03_ps_elo.groupby(by=['week']).sum()['returns'].reset_index().plot.bar(x='week', y='returns')
# -
bets_100_03_ps_elo.sum()['returns']
len(bets_100_03_ps_elo[bets_100_03_ps_elo.returns != 0])
# +
# Look at all of the above, but for elo model
# Using spread advantage of four point
bets_100_04_ps_elo = gambling_df.copy()
bets_100_04_ps_elo['returns'] = bets_100_04_ps_elo.apply(lambda x: get_return_100_spread_elo(x, 4), axis=1)
bets_100_04_ps_elo.groupby(by=['week']).sum()['returns'].reset_index().plot.bar(x='week', y='returns')
# -
bets_100_04_ps_elo.sum()['returns']
len(bets_100_04_ps_elo[bets_100_04_ps_elo.returns != 0])
# +
# Look at all of the above, but for elo model
# Using spread advantage of five points
bets_100_05_ps_elo = gambling_df.copy()
bets_100_05_ps_elo['returns'] = bets_100_05_ps_elo.apply(lambda x: get_return_100_spread_elo(x, 5), axis=1)
bets_100_05_ps_elo.groupby(by=['week']).sum()['returns'].reset_index().plot.bar(x='week', y='returns')
# -
bets_100_05_ps_elo.sum()['returns']
len(bets_100_05_ps_elo[bets_100_05_ps_elo.returns != 0])
# +
# Check winnings from random betting model for 190 games (approx same as threshold=1)
total_return_list = []
simulations = 10000
for i in range(simulations):
gambling_sample = gambling_df.sample(n=195)
gambling_sample['return'] = gambling_sample.apply(lambda x: get_return_100_spread_random(x), axis=1)
total_return_list.append(gambling_sample.sum()['return'])
print('Average Random Return: {}'.format(np.mean(total_return_list)))
plt.plot()
plt.hist(total_return_list)
plt.show()
# +
# Check where algorithm, elo would have fallen in terms of random guessing percentile
break_even_percentile = scipy.stats.percentileofscore(total_return_list, 0)
approx_algo_winnings = -220
approx_elo_winnings = -1066
algo_percentile = scipy.stats.percentileofscore(total_return_list, approx_algo_winnings)
elo_percentile = scipy.stats.percentileofscore(total_return_list, approx_elo_winnings)
print('Percentile of breaking even: {}'.format(break_even_percentile))
print('Percentile of algorithm winnings: {}'.format(algo_percentile))
print('Percentile of elo winnings: {}'.format(elo_percentile))
# +
# Check winnings from random betting model for 150 games (approx same as threshold=2)
total_return_list = []
simulations = 10000
for i in range(simulations):
gambling_sample = gambling_df.sample(n=150)
gambling_sample['return'] = gambling_sample.apply(lambda x: get_return_100_spread_random(x), axis=1)
total_return_list.append(gambling_sample.sum()['return'])
print('Average Random Return: {}'.format(np.mean(total_return_list)))
plt.plot()
plt.hist(total_return_list)
plt.show()
# +
# Check where algorithm, elo would have fallen in terms of random guessing percentile
break_even_percentile = scipy.stats.percentileofscore(total_return_list, 0)
approx_algo_winnings = -620
approx_elo_winnings = -2239
algo_percentile = scipy.stats.percentileofscore(total_return_list, approx_algo_winnings)
elo_percentile = scipy.stats.percentileofscore(total_return_list, approx_elo_winnings)
print('Percentile of breaking even: {}'.format(break_even_percentile))
print('Percentile of algorithm winnings: {}'.format(algo_percentile))
print('Percentile of elo winnings: {}'.format(elo_percentile))
# +
# Check winnings from random betting model for 105 games (approx same as threshold=3)
total_return_list = []
simulations = 10000
for i in range(simulations):
gambling_sample = gambling_df.sample(n=105)
gambling_sample['return'] = gambling_sample.apply(lambda x: get_return_100_spread_random(x), axis=1)
total_return_list.append(gambling_sample.sum()['return'])
print('Average Random Return: {}'.format(np.mean(total_return_list)))
plt.plot()
plt.hist(total_return_list)
plt.show()
# +
# Check where algorithm, elo would have fallen in terms of random guessing percentile
break_even_percentile = scipy.stats.percentileofscore(total_return_list, 0)
approx_algo_winnings = 850
approx_elo_winnings = -2486
algo_percentile = scipy.stats.percentileofscore(total_return_list, approx_algo_winnings)
elo_percentile = scipy.stats.percentileofscore(total_return_list, approx_elo_winnings)
print('Percentile of breaking even: {}'.format(break_even_percentile))
print('Percentile of algorithm winnings: {}'.format(algo_percentile))
print('Percentile of elo winnings: {}'.format(elo_percentile))
# +
# Check winnings from random betting model for 70 games (approx same as threshold=4)
total_return_list = []
simulations = 10000
for i in range(simulations):
gambling_sample = gambling_df.sample(n=70)
gambling_sample['return'] = gambling_sample.apply(lambda x: get_return_100_spread_random(x), axis=1)
total_return_list.append(gambling_sample.sum()['return'])
print('Average Random Return: {}'.format(np.mean(total_return_list)))
plt.plot()
plt.hist(total_return_list)
plt.show()
# +
# Check where algorithm, elo would have fallen in terms of random guessing percentile
break_even_percentile = scipy.stats.percentileofscore(total_return_list, 0)
approx_algo_winnings = 909
approx_elo_winnings = -1693
algo_percentile = scipy.stats.percentileofscore(total_return_list, approx_algo_winnings)
elo_percentile = scipy.stats.percentileofscore(total_return_list, approx_elo_winnings)
print('Percentile of breaking even: {}'.format(break_even_percentile))
print('Percentile of algorithm winnings: {}'.format(algo_percentile))
print('Percentile of elo winnings: {}'.format(elo_percentile))
# +
# Check winnings from random betting model for 35 games (approx same as threshold=5)
total_return_list = []
simulations = 10000
for i in range(simulations):
gambling_sample = gambling_df.sample(n=35)
gambling_sample['return'] = gambling_sample.apply(lambda x: get_return_100_spread_random(x), axis=1)
total_return_list.append(gambling_sample.sum()['return'])
print('Average Random Return: {}'.format(np.mean(total_return_list)))
plt.plot()
plt.hist(total_return_list)
plt.show()
# +
# Check where algorithm, elo would have fallen in terms of random guessing percentile
break_even_percentile = scipy.stats.percentileofscore(total_return_list, 0)
approx_algo_winnings = 870
approx_elo_winnings = -1149
algo_percentile = scipy.stats.percentileofscore(total_return_list, approx_algo_winnings)
elo_percentile = scipy.stats.percentileofscore(total_return_list, approx_elo_winnings)
print('Percentile of breaking even: {}'.format(break_even_percentile))
print('Percentile of algorithm winnings: {}'.format(algo_percentile))
print('Percentile of elo winnings: {}'.format(elo_percentile))
# +
# Now look at proportional betting
# algo
bets_prop_ps_algo = gambling_df.copy()
bets_prop_ps_algo['returns'] = bets_prop_ps_algo.apply(lambda x: get_return_proportional_spread_algo(x)[0], axis=1)
bets_prop_ps_algo.groupby(by=['week']).sum()['returns'].reset_index().plot.bar(x='week', y='returns')
# -
bets_prop_ps_algo.sum()['returns']
len(bets_prop_ps_algo[bets_prop_ps_algo.returns != 0])
# +
# elo
bets_prop_ps_elo = gambling_df.copy()
bets_prop_ps_elo['returns'] = bets_prop_ps_elo.apply(lambda x: get_return_proportional_spread_elo(x)[0], axis=1)
bets_prop_ps_elo.groupby(by=['week']).sum()['returns'].reset_index().plot.bar(x='week', y='returns')
# -
bets_prop_ps_elo.sum()['returns']
len(bets_prop_ps_elo[bets_prop_ps_elo.returns != 0])
# +
# Get chart with algo, elo
bets_prop_combined = gambling_df.copy()
bets_prop_combined[['returns_algo', 'wagered_algo']] = bets_prop_combined.apply(lambda x: get_return_proportional_spread_algo(x), axis=1, result_type='expand')
bets_prop_combined[['returns_elo', 'wagered_elo']] = bets_prop_combined.apply(lambda x: get_return_proportional_spread_elo(x), axis=1, result_type='expand')
bets_weekly = bets_prop_combined.groupby(by=['week']).sum()[['returns_algo', 'wagered_algo', 'returns_elo', 'wagered_elo']]
bets_weekly.sum()
# -
bets_weekly.cumsum()
bets_weekly.cumsum().max()
bets_weekly.cumsum().min()
# +
bets_weekly.columns = ['Algorithm Returns', 'Algorithm Wagers', 'ELO Returns', 'ELO Wagers']
plt.figure()
bets_weekly.reset_index().plot.bar(x='week', y=['Algorithm Returns', 'ELO Returns'])
plt.xlabel('2021 NFL Season Week')
plt.ylabel('Return')
plt.savefig('../figures/point_spread_returns')
plt.show()
# +
# Check winnings from random betting model for 120 games (approx same as proportional threshold)
total_return_list = []
total_wagered_list = []
simulations = 10000
for i in range(simulations):
gambling_sample = gambling_df.sample(n=120)
gambling_sample[['return', 'wagered']] = gambling_sample.apply(lambda x: get_return_prop_spread_random(x), axis=1, result_type = 'expand')
total_return_list.append(gambling_sample.sum()['return'])
total_wagered_list.append(gambling_sample.sum()['wagered'])
print('Average Random Return: {}'.format(np.mean(total_return_list)))
print('Average Random Wager: {}'.format(np.mean(total_wagered_list)))
plt.plot()
plt.hist(total_return_list)
plt.show()
# -
max(total_return_list)
min(total_return_list)
# +
# Check where algorithm, elo would have fallen in terms of random guessing percentile
break_even_percentile = scipy.stats.percentileofscore(total_return_list, 0)
approx_algo_winnings = 5742
approx_elo_winnings = -5511
algo_percentile = scipy.stats.percentileofscore(total_return_list, approx_algo_winnings)
elo_percentile = scipy.stats.percentileofscore(total_return_list, approx_elo_winnings)
print('Percentile of breaking even: {}'.format(break_even_percentile))
print('Percentile of algorithm winnings: {}'.format(algo_percentile))
print('Percentile of elo winnings: {}'.format(elo_percentile))
# +
# Create plot with histogram and percentiles of algorithm and elo returns
# Graphing code from https://towardsdatascience.com/take-your-histograms-to-the-next-level-using-matplotlib-5f093ad7b9d3
# https://stackoverflow.com/questions/51980366/image-size-of-1005x132589-pixels-is-too-large-it-must-be-less-than-216-in-each
import seaborn as sns
from scipy.stats import norm
quants = [[-5511, 0.9, 0.67], [0, 0.9, 0.93], [5742, 0.9, 0.2]]
fig, ax = plt.subplots()
sns.distplot(total_return_list, fit=norm, kde=False)
ax.set_yticks([])
plt.ylabel('Frequency')
plt.xlabel('2021 Season Return')
# Plot the lines with a loop
for i in quants:
ax.axvline(i[0], alpha = i[1], ymax = i[2], linestyle = "--", color='g')
ax.text(.37, 0.95, "Break-Even Percentile: 68.05", size = 10, alpha = 0.9, transform=ax.transAxes)
ax.text(.1, 0.7, "ELO Percentile: 12.59", size = 10, alpha = 0.9, transform=ax.transAxes)
ax.text(.58, 0.22, "Algorithm Percentile: 98.23", size = 10, alpha = 0.9, transform=ax.transAxes)
plt.savefig('../figures/point_spread_percentiles')
plt.show()
# +
# Now, look at under/over
# Since elo has no way to determine over/under totals, will be using only algorithm and random guessing distributions
# Using over/under advantage of one point
bets_100_01_ou_algo = gambling_df.copy()
bets_100_01_ou_algo['returns'] = bets_100_01_ou_algo.apply(lambda x: get_return_100_ou_algo(x, 1), axis=1)
bets_100_01_ou_algo.groupby(by=['week']).sum()['returns'].reset_index().plot.bar(x='week', y='returns')
# -
bets_100_01_ou_algo.sum()['returns']
len(bets_100_01_ou_algo[bets_100_01_ou_algo.returns!=0])
# +
# Using over/under advantage of two points
bets_100_02_ou_algo = gambling_df.copy()
bets_100_02_ou_algo['returns'] = bets_100_02_ou_algo.apply(lambda x: get_return_100_ou_algo(x, 2), axis=1)
bets_100_02_ou_algo.groupby(by=['week']).sum()['returns'].reset_index().plot.bar(x='week', y='returns')
# -
bets_100_02_ou_algo.sum()['returns']
len(bets_100_02_ou_algo[bets_100_02_ou_algo.returns!=0])
# +
# Using over/under advantage of three points
bets_100_03_ou_algo = gambling_df.copy()
bets_100_03_ou_algo['returns'] = bets_100_03_ou_algo.apply(lambda x: get_return_100_ou_algo(x, 3), axis=1)
bets_100_03_ou_algo.groupby(by=['week']).sum()['returns'].reset_index().plot.bar(x='week', y='returns')
# -
bets_100_03_ou_algo.sum()['returns']
len(bets_100_03_ou_algo[bets_100_03_ou_algo.returns!=0])
# +
# Using over/under advantage of four points
bets_100_04_ou_algo = gambling_df.copy()
bets_100_04_ou_algo['returns'] = bets_100_04_ou_algo.apply(lambda x: get_return_100_ou_algo(x, 4), axis=1)
bets_100_04_ou_algo.groupby(by=['week']).sum()['returns'].reset_index().plot.bar(x='week', y='returns')
# -
bets_100_04_ou_algo.sum()['returns']
len(bets_100_04_ou_algo[bets_100_04_ou_algo.returns!=0])
# +
# Using over/under advantage of five points
bets_100_05_ou_algo = gambling_df.copy()
bets_100_05_ou_algo['returns'] = bets_100_05_ou_algo.apply(lambda x: get_return_100_ou_algo(x, 5), axis=1)
bets_100_05_ou_algo.groupby(by=['week']).sum()['returns'].reset_index().plot.bar(x='week', y='returns')
# -
bets_100_05_ou_algo.sum()['returns']
len(bets_100_05_ou_algo[bets_100_05_ou_algo.returns!=0])
# +
# Now look at random winnings for over unders
# Check winnings from random betting model for 200 games (approx same as 1 point threshold)
total_return_list = []
simulations = 10000
for i in range(simulations):
gambling_sample = gambling_df.sample(n=200)
gambling_sample['return'] = gambling_sample.apply(lambda x: get_return_100_ou_random(x), axis=1)
total_return_list.append(gambling_sample.sum()['return'])
print('Average Random Return: {}'.format(np.mean(total_return_list)))
plt.plot()
plt.hist(total_return_list)
plt.show()
# +
# Check where algorithm, elo would have fallen in terms of random guessing percentile
break_even_percentile = scipy.stats.percentileofscore(total_return_list, 0)
approx_algo_winnings = 1531
# approx_elo_winnings = -5511
algo_percentile = scipy.stats.percentileofscore(total_return_list, approx_algo_winnings)
# elo_percentile = scipy.stats.percentileofscore(total_return_list, approx_elo_winnings)
print('Percentile of breaking even: {}'.format(break_even_percentile))
print('Percentile of algorithm winnings: {}'.format(algo_percentile))
# +
# Check winnings from random betting model for 135 games (approx same as 2 point threshold)
total_return_list = []
simulations = 10000
for i in range(simulations):
gambling_sample = gambling_df.sample(n=135)
gambling_sample['return'] = gambling_sample.apply(lambda x: get_return_100_ou_random(x), axis=1)
total_return_list.append(gambling_sample.sum()['return'])
print('Average Random Return: {}'.format(np.mean(total_return_list)))
plt.plot()
plt.hist(total_return_list)
plt.show()
# +
# Check where algorithm, elo would have fallen in terms of random guessing percentile
break_even_percentile = scipy.stats.percentileofscore(total_return_list, 0)
approx_algo_winnings = 1129
# approx_elo_winnings = -5511
algo_percentile = scipy.stats.percentileofscore(total_return_list, approx_algo_winnings)
# elo_percentile = scipy.stats.percentileofscore(total_return_list, approx_elo_winnings)
print('Percentile of breaking even: {}'.format(break_even_percentile))
print('Percentile of algorithm winnings: {}'.format(algo_percentile))
# +
# Check winnings from random betting model for 95 games (approx same as 3 point threshold)
total_return_list = []
simulations = 10000
for i in range(simulations):
gambling_sample = gambling_df.sample(n=95)
gambling_sample['return'] = gambling_sample.apply(lambda x: get_return_100_ou_random(x), axis=1)
total_return_list.append(gambling_sample.sum()['return'])
print('Average Random Return: {}'.format(np.mean(total_return_list)))
plt.plot()
plt.hist(total_return_list)
plt.show()
# +
# Check where algorithm, elo would have fallen in terms of random guessing percentile
break_even_percentile = scipy.stats.percentileofscore(total_return_list, 0)
approx_algo_winnings = 1300
# approx_elo_winnings = -5511
algo_percentile = scipy.stats.percentileofscore(total_return_list, approx_algo_winnings)
# elo_percentile = scipy.stats.percentileofscore(total_return_list, approx_elo_winnings)
print('Percentile of breaking even: {}'.format(break_even_percentile))
print('Percentile of algorithm winnings: {}'.format(algo_percentile))
# +
# Check winnings from random betting model for 60 games (approx same as proportional threshold)
total_return_list = []
simulations = 10000
for i in range(simulations):
gambling_sample = gambling_df.sample(n=60)
gambling_sample['return'] = gambling_sample.apply(lambda x: get_return_100_ou_random(x), axis=1)
total_return_list.append(gambling_sample.sum()['return'])
print('Average Random Return: {}'.format(np.mean(total_return_list)))
plt.plot()
plt.hist(total_return_list)
plt.show()
# +
# Check where algorithm, elo would have fallen in terms of random guessing percentile
break_even_percentile = scipy.stats.percentileofscore(total_return_list, 0)
approx_algo_winnings = 1068
# approx_elo_winnings = -5511
algo_percentile = scipy.stats.percentileofscore(total_return_list, approx_algo_winnings)
# elo_percentile = scipy.stats.percentileofscore(total_return_list, approx_elo_winnings)
print('Percentile of breaking even: {}'.format(break_even_percentile))
print('Percentile of algorithm winnings: {}'.format(algo_percentile))
# +
# Check winnings from random betting model for 30 games (approx same as proportional threshold)
total_return_list = []
simulations = 10000
for i in range(simulations):
gambling_sample = gambling_df.sample(n=30)
gambling_sample['return'] = gambling_sample.apply(lambda x: get_return_100_ou_random(x), axis=1)
total_return_list.append(gambling_sample.sum()['return'])
print('Average Random Return: {}'.format(np.mean(total_return_list)))
plt.plot()
plt.hist(total_return_list)
plt.show()
# +
# Check where algorithm, elo would have fallen in terms of random guessing percentile
break_even_percentile = scipy.stats.percentileofscore(total_return_list, 0)
approx_algo_winnings = 900
# approx_elo_winnings = -5511
algo_percentile = scipy.stats.percentileofscore(total_return_list, approx_algo_winnings)
# elo_percentile = scipy.stats.percentileofscore(total_return_list, approx_elo_winnings)
print('Percentile of breaking even: {}'.format(break_even_percentile))
print('Percentile of algorithm winnings: {}'.format(algo_percentile))
# +
# Finally, see how algorithm would have performed using a proportional betting system
bets_prop_ou_algo = gambling_df.copy()
bets_prop_ou_algo[['returns', 'wagered']] = bets_prop_ou_algo.apply(lambda x: get_return_proportional_ou_algo(x), axis=1, result_type='expand')
bets_prop_ou_algo['Algorithm Returns'] = bets_prop_ou_algo.returns
plt.figure()
bets_prop_ou_algo.groupby(by=['week']).sum()['Algorithm Returns'].reset_index().plot.bar(x='week', y='Algorithm Returns')
plt.xlabel('2021 NFL Season Week')
plt.ylabel('Return')
plt.savefig('../figures/over_under_returns')
plt.show()
# -
bets_prop_ou_algo.sum()[['returns', 'wagered']]
len(bets_prop_ou_algo[bets_prop_ou_algo.returns!=0])
bets_prop_ou_algo.groupby(by='week').sum()[['returns', 'wagered']].cumsum()
bets_prop_ou_algo.groupby(by='week').sum()[['returns', 'wagered']].cumsum().max()
bets_prop_ou_algo.groupby(by='week').sum()[['returns', 'wagered']].cumsum().min()
# +
# Check winnings from random betting model for 195 games (approx same as proportional threshold)
total_return_list = []
total_wagered_list = []
simulations = 10000
for i in range(simulations):
gambling_sample = gambling_df.sample(n=195)
gambling_sample[['return', 'wager']] = gambling_sample.apply(lambda x: get_return_prop_ou_random(x), axis=1, result_type='expand')
total_return_list.append(gambling_sample.sum()['return'])
total_wagered_list.append(gambling_sample.sum()['wager'])
print('Average Random Return: {}'.format(np.mean(total_return_list)))
print('Average Random Wager: {}'.format(np.mean(total_wagered_list)))
plt.plot()
plt.hist(total_return_list)
plt.show()
# +
# Check where algorithm, elo would have fallen in terms of random guessing percentile
break_even_percentile = scipy.stats.percentileofscore(total_return_list, 0)
approx_algo_winnings = 5934
# approx_elo_winnings = -5511
algo_percentile = scipy.stats.percentileofscore(total_return_list, approx_algo_winnings)
# elo_percentile = scipy.stats.percentileofscore(total_return_list, approx_elo_winnings)
print('Percentile of breaking even: {}'.format(break_even_percentile))
print('Percentile of algorithm winnings: {}'.format(algo_percentile))
# +
# Create plot with histogram and percentiles of algorithm and elo returns
# Graphing code from https://towardsdatascience.com/take-your-histograms-to-the-next-level-using-matplotlib-5f093ad7b9d3
# https://stackoverflow.com/questions/51980366/image-size-of-1005x132589-pixels-is-too-large-it-must-be-less-than-216-in-each
import seaborn as sns
from scipy.stats import norm
quants = [[0, 0.9, 0.83], [5934, 0.9, 0.21]]
fig, ax = plt.subplots()
sns.distplot(total_return_list, fit=norm, kde=False)
ax.set_yticks([])
plt.ylabel('Frequency')
plt.xlabel('2021 Season Return')
# Plot the lines with a loop
for i in quants:
ax.axvline(i[0], alpha = i[1], ymax = i[2], linestyle = "--", color='g')
ax.text(.4, 0.86, "Break-Even Percentile: 70.96", size = 10, alpha = 0.9, transform=ax.transAxes)
# ax.text(.1, 0.95, "ELO Percentile: 34.13", size = 10, alpha = 0.9, transform=ax.transAxes)
ax.text(.58, 0.23, "Algorithm Percentile: 96.69", size = 10, alpha = 0.9, transform=ax.transAxes)
plt.savefig('../figures/over_under_percentiles')
plt.show()
# -
np.mean(total_return_list)
min(total_return_list)
max(total_return_list)
np.mean(total_wagered_list)
bets_prop_ou_algo
| ideal_betting_proportions_spreads.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Practice Exercise 1
# This is the first exercise for Pandas. You are provided with 2 fictional dataframes based on Indian Premier League (IPL). The first dataframe contains the data for all the teams who participated in the year 2018. In the second dataframe, the data is for the year 2017.
# import the required libraries - numpy and pandas
# #### Data Dictionary
#
# - Team: Team name
# - Matches: Total number of matches played
# - Won: Number of matches won
# - Lost: Number of matches lost
# - Tied': Number of matches tied
# - N/R: Number of matches with no result
# - NRR: Net run rate (Rate of scoring the runs)
# - For: Sum of runs scored by the team against other teams
# - Against: Sum of runs scored by the opposite playing teams
# ipl18 contains the data for 2018
ipl18 = pd.DataFrame({'Team': ['SRH', 'CSK', 'KKR', 'RR', 'MI', 'RCB', 'KXIP', 'DD'],
'Matches': [14, 14, 14, 14, 14, 14, 14, 14],
'Won': [9, 9, 8, 7, 6, 6, 6, 5],
'Lost': [5, 5, 6, 7, 8, 8, 8, 9],
'Tied': [0, 0, 0, 0, 0, 0, 0, 0],
'N/R': [0, 0, 0, 0, 0, 0, 0, 0],
'NRR': [0.284, 0.253, -0.070, -0.250, 0.317, 0.129, -0.502, -0.222],
'For': [2230, 2488, 2363, 2130, 2380, 2322, 2210, 2297],
'Against': [2193, 2433, 2425, 2141, 2282, 2383, 2259, 2304]},
index = range(1,9)
)
# print the entire dataframe to check the entries
ipl18
# ipl17 contains the data for 2017
ipl17 = pd.DataFrame({'Team': ['MI', 'RPS', 'SRH', 'KKR', 'KXIP', 'DD', 'GL', 'RCB'],
'Matches': [14, 14, 14, 14, 14, 14, 14, 14],
'Won': [10, 9, 8, 8, 7, 6, 4, 3],
'Lost': [4, 5, 5, 6, 7, 8, 10, 10],
'Tied': [0, 0, 0, 0, 0, 0, 0, 0],
'N/R': [0, 0, 1, 0, 0, 0, 0, 1],
'NRR': [0.784, 0.176, 0.469, 0.641, 0.123, -0.512, -0.412, -1.299],
'For': [2407, 2180, 2221, 2329, 2207, 2219, 2406, 1845],
'Against': [2242, 2165, 2118, 2300, 2229, 2255, 2472, 2033]},
index = range(1,9)
)
# print the entire dataframe to check the entries
ipl17
# As a part of this exercise, solve the questions that are provided below. There are few tasks that you will have to perform to be able to answer the questions.
#
# **Task 1**
#
# You have to create a new column '*Points*' in both the dataframes that stores the total points scored by each team. The following scoring system is used to calculate the points of a team:
# - Win: 2 points
# - Loss: 0 points
# - Tie: 1 point
# - N/R (no result): 1 point
# Type your code here
# #### Q1: Extract Top Four Teams
# Which of the following commands can you use to extract the top 4 teams in the dataset ‘ipl18’ with just the ‘Team’ and ‘Points’ column?
#
#
# - ipl18.loc [0:3, [‘Team’, ‘Points’]]
# - ipl18.loc [1:4, [‘Team’, ‘Points’]]
# - ipl18.loc [0:4, [‘Team’, ‘Points’]]
# - ipl18.loc [1:3, [‘Team’, ‘Points’]]
# +
# Type your code here
# -
# #### Q2: Filtering based on conditions
# Suppose in ‘ipl18’, you want to filter out the teams that have an NRR greater than zero, and for which the ‘For’ score exceeds the ‘Against’ score, i.e. both the conditions should be satisfied. Which teams will be left after you perform the above filtration? (Run the commands on the Python Notebook provided, rather than performing a manual calculation)
#
# - CSK, MI
# - SRH, CSK, MI
# - SRH, CSK, RCB
# - SRH, CSK, MI, RCB
# +
# Type your code here
# -
# #### Q3: Operations on multiple dataframes
# If all the stats are taken for both ‘ipl17’ and ‘ipl18’, which team with its total points greater than 25 will have the highest win percentage?
#
# - KKR
# - CSK
# - RPS
# - SRH
# +
# Type your code here
| Data Science/Pandas/.ipynb_checkpoints/Practice+Exercise+1+IPL-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# # **Amazon Lookout for Equipment** - Getting started
# *Part 2 - Dataset creation*
# ## Initialization
# ---
# This repository is structured as follow:
#
# ```sh
# . lookout-equipment-demo
# |
# ├── data/
# | ├── interim # Temporary intermediate data are stored here
# | ├── processed # Finalized datasets are usually stored here
# | | # before they are sent to S3 to allow the
# | | # service to reach them
# | └── raw # Immutable original data are stored here
# |
# ├── getting_started/
# | ├── 1_data_preparation.ipynb
# | ├── 2_dataset_creation.ipynb <<< THIS NOTEBOOK <<<
# | ├── 3_model_training.ipynb
# | ├── 4_model_evaluation.ipynb
# | ├── 5_inference_scheduling.ipynb
# | └── 6_cleanup.ipynb
# |
# └── utils/
# └── lookout_equipment_utils.py
# ```
# ### Notebook configuration update
# !pip install --quiet --upgrade sagemaker tqdm lookoutequipment
# ### Imports
# +
import boto3
import config
import os
import pandas as pd
import sagemaker
import sys
import time
from datetime import datetime
# SDK / toolbox for managing Lookout for Equipment API calls:
import lookoutequipment as lookout
# +
PROCESSED_DATA = os.path.join('..', 'data', 'processed', 'getting-started')
TRAIN_DATA = os.path.join(PROCESSED_DATA, 'training-data')
ROLE_ARN = sagemaker.get_execution_role()
REGION_NAME = boto3.session.Session().region_name
DATASET_NAME = config.DATASET_NAME
BUCKET = config.BUCKET
PREFIX = config.PREFIX_TRAINING
# -
# ## Create a dataset
# ---
lookout_dataset = lookout.LookoutEquipmentDataset(
dataset_name=DATASET_NAME,
component_root_dir=TRAIN_DATA,
access_role_arn=ROLE_ARN
)
# Let's double check the schema detected for our dataset to ensure we are pointing to the right data:
import pprint
pp = pprint.PrettyPrinter(depth=5)
pp.pprint(eval(lookout_dataset.dataset_schema))
# The following method encapsulate the [**CreateDataset**](https://docs.aws.amazon.com/lookout-for-equipment/latest/ug/API_CreateDataset.html) API:
#
# ```python
# lookout_client.create_dataset(
# DatasetName=self.dataset_name,
# DatasetSchema={
# 'InlineDataSchema': "schema"
# }
# )
# ```
lookout_dataset.create()
# The dataset is now created, but it is empty and ready to receive some timeseries data that we will ingest from the S3 location prepared in the previous notebook:
# 
# ## Ingest data into a dataset
# ---
# Let's double check the values of all the parameters that will be used to ingest some data into an existing Lookout for Equipment dataset:
ROLE_ARN, BUCKET, PREFIX, DATASET_NAME
# Launch the ingestion job in the Lookout for Equipment dataset: the following method encapsulates the [**StartDataIngestionJob**](https://docs.aws.amazon.com/lookout-for-equipment/latest/ug/API_StartDataIngestionJob.html) API:
#
# ```python
# lookout_client.start_data_ingestion_job(
# DatasetName=DATASET_NAME,
# RoleArn=ROLE_ARN,
# IngestionInputConfiguration={
# 'S3InputConfiguration': {
# 'Bucket': BUCKET,
# 'Prefix': PREFIX
# }
# }
# )
# ```
response = lookout_dataset.ingest_data(BUCKET, PREFIX)
# The ingestion is launched. With this amount of data (around 50 MB), it should take between less than 5 minutes:
#
# 
# We use the following cell to monitor the ingestion process by calling the following method, which encapsulates the [**DescribeDataIngestionJob**](https://docs.aws.amazon.com/lookout-for-equipment/latest/ug/API_DescribeDataIngestionJob.html) API and runs it every 60 seconds:
lookout_dataset.poll_data_ingestion(sleep_time=60)
# In case any issue arise, you can inspect the API response available as a JSON document:
lookout_dataset.ingestion_job_response
# The ingestion should now be complete as can be seen in the console:
#
# 
# ## Conclusion
# ---
# In this notebook, we created a **Lookout for Equipment dataset** and ingested the S3 data previously uploaded into this dataset. **Move now to the next notebook to train a model based on these data.**
| getting_started/completed-notebooks/2_dataset_creation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.6.5
# language: julia
# name: julia-1.6
# ---
# # Machine Learning in Julia (conclusion)
# An introduction to the
# [MLJ](https://alan-turing-institute.github.io/MLJ.jl/stable/)
# toolbox.
# ### Set-up
# Inspect Julia version:
VERSION
# The following instantiates a package environment.
# The package environment has been created using **Julia 1.6** and may not
# instantiate properly for other Julia versions.
using Pkg
Pkg.activate("env")
Pkg.instantiate()
# ## General resources
# - [MLJ Cheatsheet](https://alan-turing-institute.github.io/MLJ.jl/dev/mlj_cheatsheet/)
# - [Common MLJ Workflows](https://alan-turing-institute.github.io/MLJ.jl/dev/common_mlj_workflows/)
# - [MLJ manual](https://alan-turing-institute.github.io/MLJ.jl/dev/)
# - [Data Science Tutorials in Julia](https://juliaai.github.io/DataScienceTutorials.jl/)
# ## Solutions to exercises
using MLJ, UrlDownload, CSV, DataFrames, Plots
# #### Exercise 2 solution
# From the question statememt:
quality = ["good", "poor", "poor", "excellent", missing, "good", "excellent"]
quality = coerce(quality, OrderedFactor);
levels!(quality, ["poor", "good", "excellent"]);
elscitype(quality)
# #### Exercise 3 solution
# From the question statement:
house_csv = urldownload("https://raw.githubusercontent.com/ablaom/"*
"MachineLearningInJulia2020/for-MLJ-version-0.16/"*
"data/house.csv");
house = DataFrames.DataFrame(house_csv)
# First pass:
coerce!(house, autotype(house));
schema(house)
# All the "sqft" fields refer to "square feet" so are
# really `Continuous`. We'll regard `:yr_built` (the other `Count`
# variable above) as `Continuous` as well. So:
coerce!(house, Count => Continuous);
# And `:zipcode` should not be ordered:
coerce!(house, :zipcode => Multiclass);
schema(house)
# `:bathrooms` looks like it has a lot of levels, but on further
# inspection we see why, and `OrderedFactor` remains appropriate:
import StatsBase.countmap
countmap(house.bathrooms)
# #### Exercise 4 solution
# From the question statement:
# +
import Distributions
poisson = Distributions.Poisson
age = 18 .+ 60*rand(10);
salary = coerce(rand(["small", "big", "huge"], 10), OrderedFactor);
levels!(salary, ["small", "big", "huge"]);
small = salary[1]
X4 = DataFrames.DataFrame(age=age, salary=salary)
n_devices(salary) = salary > small ? rand(poisson(1.3)) : rand(poisson(2.9))
y4 = [n_devices(row.salary) for row in eachrow(X4)]
# -
# 4(a)
# There are *no* models that apply immediately:
models(matching(X4, y4))
# 4(b)
y4 = coerce(y4, Continuous);
models(matching(X4, y4))
# #### Exercise 6 solution
# From the question statement:
using UrlDownload, CSV
csv_file = urldownload("https://raw.githubusercontent.com/ablaom/"*
"MachineLearningInJulia2020/"*
"for-MLJ-version-0.16/data/horse.csv");
horse = DataFrames.DataFrame(csv_file); # convert to data frame
coerce!(horse, autotype(horse));
coerce!(horse, Count => Continuous);
coerce!(horse,
:surgery => Multiclass,
:age => Multiclass,
:mucous_membranes => Multiclass,
:capillary_refill_time => Multiclass,
:outcome => Multiclass,
:cp_data => Multiclass);
schema(horse)
# 6(a)
y, X = unpack(horse,
==(:outcome),
name -> elscitype(Tables.getcolumn(horse, name)) == Continuous);
# 6(b)(i)
train, test = partition(eachindex(y), 0.7)
model = (@load LogisticClassifier pkg=MLJLinearModels)();
model.lambda = 100
mach = machine(model, X, y)
fit!(mach, rows=train)
fitted_params(mach)
# +
coefs_given_feature = Dict(fitted_params(mach).coefs)
coefs_given_feature[:pulse]
#6(b)(ii)
yhat = predict(mach, rows=test); # or predict(mach, X[test,:])
err = cross_entropy(yhat, y[test]) |> mean
# -
# 6(b)(iii)
# The predicted probabilities of the actual observations in the test
# are given by
p = broadcast(pdf, yhat, y[test]);
# The number of times this probability exceeds 50% is:
n50 = filter(x -> x > 0.5, p) |> length
# Or, as a proportion:
n50/length(test)
# 6(b)(iv)
misclassification_rate(mode.(yhat), y[test])
# 6(c)(i)
# +
model = (@load RandomForestClassifier pkg=DecisionTree)()
mach = machine(model, X, y)
evaluate!(mach, resampling=CV(nfolds=6), measure=cross_entropy)
r = range(model, :n_trees, lower=10, upper=70, scale=:log10)
# -
# Since random forests are inherently randomized, we generate multiple
# curves:
plt = plot()
for i in 1:4
one_curve = learning_curve(mach,
range=r,
resampling=Holdout(),
measure=cross_entropy)
plot!(one_curve.parameter_values, one_curve.measurements)
end
xlabel!(plt, "n_trees")
ylabel!(plt, "cross entropy")
savefig("exercise_6ci.png")
plt
# 6(c)(ii)
# +
evaluate!(mach, resampling=CV(nfolds=9),
measure=cross_entropy,
rows=train).measurement[1]
model.n_trees = 90
# -
# 6(c)(iii)
err_forest = evaluate!(mach, resampling=Holdout(),
measure=cross_entropy).measurement[1]
# #### Exercise 7
# (a)
KMeans = @load KMeans pkg=Clustering
EvoTreeClassifier = @load EvoTreeClassifier
pipe = Standardizer |>
ContinuousEncoder |>
KMeans(k=10) |>
EvoTreeClassifier(nrounds=50)
# (b)
mach = machine(pipe, X, y)
evaluate!(mach, resampling=CV(nfolds=6), measure=cross_entropy)
# (c)
# +
r = range(pipe, :(evo_tree_classifier.max_depth), lower=1, upper=10)
curve = learning_curve(mach,
range=r,
resampling=CV(nfolds=6),
measure=cross_entropy)
plt = plot(curve.parameter_values, curve.measurements)
xlabel!(plt, "max_depth")
ylabel!(plt, "CV estimate of cross entropy")
savefig("exercise_7c.png")
plt
# -
# Here's a second curve using a different random seed for the booster:
using Random
pipe.evo_tree_classifier.rng = MersenneTwister(123)
curve = learning_curve(mach,
range=r,
resampling=CV(nfolds=6),
measure=cross_entropy)
plot!(curve.parameter_values, curve.measurements)
savefig("exercise_7c_2.png")
plt
# One can automate the production of multiple curves with different
# seeds in the following way:
curves = learning_curve(mach,
range=r,
resampling=CV(nfolds=6),
measure=cross_entropy,
rng_name=:(evo_tree_classifier.rng),
rngs=6) # list of RNGs, or num to auto generate
plt = plot(curves.parameter_values, curves.measurements)
savefig("exercise_7c_3.png")
plt
# If you have multiple threads available in your julia session, you
# can add the option `acceleration=CPUThreads()` to speed up this
# computation.
# #### Exercise 8
# From the question statement:
# +
y, X = unpack(house, ==(:price), rng=123); # from Exercise 3
EvoTreeRegressor = @load EvoTreeRegressor
tree_booster = EvoTreeRegressor(nrounds = 70)
model = ContinuousEncoder |> tree_booster
r2 = range(model,
:(evo_tree_regressor.nbins),
lower = 2.5,
upper= 7.5, scale=x->2^round(Int, x))
# -
# (a)
r1 = range(model, :(evo_tree_regressor.max_depth), lower=1, upper=12)
# (c)
# +
tuned_model = TunedModel(model=model,
ranges=[r1, r2],
resampling=Holdout(),
measures=mae,
tuning=RandomSearch(rng=123),
n=40)
tuned_mach = machine(tuned_model, X, y) |> fit!
plt = plot(tuned_mach)
savefig("exercise_8c.png")
plt
# -
# (d)
best_model = report(tuned_mach).best_model;
best_mach = machine(best_model, X, y);
best_err = evaluate!(best_mach, resampling=CV(nfolds=3), measure=mae)
tuned_err = evaluate!(tuned_mach, resampling=CV(nfolds=3), measure=mae)
# ---
#
# *This notebook was generated using [Literate.jl](https://github.com/fredrikekre/Literate.jl).*
| notebooks/99_solution_to_exercises/notebook.unexecuted.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
## Here I am dividing the data first based onto protected attribute value and then train two separate models
import pandas as pd
import random,time
import numpy as np
import math,copy,os
from sklearn.linear_model import LogisticRegression
from sklearn.tree import DecisionTreeRegressor
from sklearn.model_selection import train_test_split
from sklearn.metrics import classification_report, confusion_matrix, accuracy_score
from sklearn import tree
from sklearn.naive_bayes import GaussianNB
from sklearn.ensemble import RandomForestClassifier
from sklearn.svm import SVC
import sklearn.metrics as metrics
import sys
sys.path.append(os.path.abspath('..'))
from Measure import measure_final_score,calculate_recall,calculate_far,calculate_precision,calculate_accuracy
# +
## Load dataset
dataset_orig = pd.read_csv('../dataset/bank.csv')
## Drop categorical features
dataset_orig = dataset_orig.drop(['job', 'marital', 'default',
'housing', 'loan', 'contact', 'month', 'day',
'poutcome'],axis=1)
## Drop NULL values
dataset_orig = dataset_orig.dropna()
## IBM used 25 as divider but age divider should be mean
# mean = dataset_orig.loc[:,"age"].mean()
# dataset_orig['age'] = np.where(dataset_orig['age'] >= mean, 1, 0)
dataset_orig['age'] = np.where(dataset_orig['age'] >= 25, 1, 0)
dataset_orig['Probability'] = np.where(dataset_orig['Probability'] == 'yes', 1, 0)
## Chaneg symbolic to numeric column
from sklearn.preprocessing import LabelEncoder
gle = LabelEncoder()
genre_labels = gle.fit_transform(dataset_orig['education'])
genre_mappings = {index: label for index, label in enumerate(gle.classes_)}
dataset_orig['education'] = genre_labels
from sklearn.preprocessing import MinMaxScaler
scaler = MinMaxScaler()
dataset_orig = pd.DataFrame(scaler.fit_transform(dataset_orig),columns = dataset_orig.columns)
# divide the data based on age
dataset_orig_male , dataset_orig_female = [x for _, x in dataset_orig.groupby(dataset_orig['age'] == 0)]
print(dataset_orig_male.shape)
print(dataset_orig_female.shape)
# dataset_orig
# -
# # Train the model for >= 41 age
# +
dataset_orig_male['age'] = 0
X_train_male, y_train_male = dataset_orig_male.loc[:, dataset_orig_male.columns != 'Probability'], dataset_orig_male['Probability']
# --- LSR
clf_male = LogisticRegression(C=1.0, penalty='l2', solver='liblinear', max_iter=100)
clf_male.fit(X_train_male, y_train_male)
# print(X_train_male['sex'])
import matplotlib.pyplot as plt
y = np.arange(len(dataset_orig_male.columns)-1)
plt.barh(y,clf_male.coef_[0])
plt.yticks(y,dataset_orig_male.columns)
plt.show()
print(clf_male.coef_[0])
# -
# # Train the model for <41 age
# +
X_train_female, y_train_female = dataset_orig_female.loc[:, dataset_orig_female.columns != 'Probability'], dataset_orig_female['Probability']
# --- LSR
clf_female = LogisticRegression(C=1.0, penalty='l2', solver='liblinear', max_iter=100)
clf_female.fit(X_train_female, y_train_female)
import matplotlib.pyplot as plt
y = np.arange(len(dataset_orig_female.columns)-1)
plt.barh(y,clf_female.coef_[0])
plt.yticks(y,dataset_orig_female.columns)
plt.show()
# -
# # Remove biased rows
# +
print(dataset_orig.shape)
for index,row in dataset_orig.iterrows():
row = [row.values[0:len(row.values)-1]]
y_male = clf_male.predict(row)
y_female = clf_female.predict(row)
# print(y_male,y_female)
if y_male[0] != y_female[0]:
if y_male[0] == 0:
dataset_orig = dataset_orig.drop(index)
print(dataset_orig.shape)
# -
# # Train and test new model on unbiased data
# +
print(dataset_orig.shape)
np.random.seed(0)
## Divide into train,validation,test
dataset_orig_train, dataset_orig_test = train_test_split(dataset_orig, test_size=0.2, random_state=0,shuffle = True)
X_train, y_train = dataset_orig_train.loc[:, dataset_orig_train.columns != 'Probability'], dataset_orig_train['Probability']
X_test , y_test = dataset_orig_test.loc[:, dataset_orig_test.columns != 'Probability'], dataset_orig_test['Probability']
# --- LSR
clf = LogisticRegression(C=1.0, penalty='l2', solver='liblinear', max_iter=100)
# --- CART
# clf = tree.DecisionTreeClassifier()
# clf.fit(X_train, y_train)
# import matplotlib.pyplot as plt
# y = np.arange(len(dataset_orig_train.columns)-1)
# plt.barh(y,clf.coef_[0])
# plt.yticks(y,dataset_orig_train.columns)
# plt.show()
# print(clf_male.coef_[0])
# y_pred = clf.predict(X_test)
# cnf_matrix_test = confusion_matrix(y_test,y_pred)
# print(cnf_matrix_test)
# TN, FP, FN, TP = confusion_matrix(y_test,y_pred).ravel()
print("recall :", measure_final_score(dataset_orig_test, clf, X_train, y_train, X_test, y_test, 'age', 'recall'))
print("far :",measure_final_score(dataset_orig_test, clf, X_train, y_train, X_test, y_test, 'age', 'far'))
print("precision :", measure_final_score(dataset_orig_test, clf, X_train, y_train, X_test, y_test, 'age', 'precision'))
print("accuracy :",measure_final_score(dataset_orig_test, clf, X_train, y_train, X_test, y_test, 'age', 'accuracy'))
print("aod sex:",measure_final_score(dataset_orig_test, clf, X_train, y_train, X_test, y_test, 'age', 'aod'))
print("eod sex:",measure_final_score(dataset_orig_test, clf, X_train, y_train, X_test, y_test, 'age', 'eod'))
print("TPR:",measure_final_score(dataset_orig_test, clf, X_train, y_train, X_test, y_test, 'age', 'TPR'))
print("FPR:",measure_final_score(dataset_orig_test, clf, X_train, y_train, X_test, y_test, 'age', 'FPR'))
# -
| Split_On_Protected_Attribute/Bank.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import keras
from keras.preprocessing import sequence
from keras.models import Sequential
from keras.layers import Dense, Dropout, Activation
from keras.layers import Embedding
from keras.layers import Conv1D, GlobalMaxPooling1D
from keras.datasets import imdb
# set parameters:
max_features = 5000
maxlen = 400
batch_size = 32
embedding_dims = 50
filters = 250
kernel_size = 3
hidden_dims = 250
epochs = 2
# +
import numpy as np
np_load_old = np.load # save old function for calling later
# modify the default parameters of np.load
np.load = lambda *a,**k: np_load_old(*a, allow_pickle=True, **k)
# +
print('Loading data...')
(x_train, y_train), (x_test, y_test) = imdb.load_data(num_words=max_features)
print(len(x_train), 'train sequences')
print(len(x_test), 'test sequences')
# -
# # Understanding the inputs to the model
# One of the most difficult things to do is to feed your own data to the model. That is extremely important if you want to apply your skills in the real world -- to actually put machine learning to use in the industry.
#
# Let's try to understand our inputs here so that we can later feed our own data to the model.
x_train.shape # 25000 samples, each with a total (fixed) length of 400 'words'
y_train.shape
y_train[0] # labels are easy to understand. 0 is negative sentiment, 1 is positive
x_train[0]
word_to_id = keras.datasets.imdb.get_word_index()
list(word_to_id.items())[:10]
print(word_to_id['love'])
print(word_to_id['like'])
print(word_to_id['boring'])
print(word_to_id['interesting'])
def get_fixed_word_to_id_dict():
INDEX_FROM=3 # word index offset
word_to_id = keras.datasets.imdb.get_word_index()
word_to_id = {k:(v+INDEX_FROM) for k,v in word_to_id.items()}
word_to_id[" "] = 0
word_to_id["<START>"] = 1
word_to_id["<UNK>"] = 2
return word_to_id
def decode_to_sentence(data_point):
#NUM_WORDS=1000 # only use top 1000 words
word_to_id = get_fixed_word_to_id_dict()
id_to_word = {value:key for key,value in word_to_id.items()}
return ' '.join( id_to_word[id] for id in data_point )
data_point_to_show = 0
x_train[data_point_to_show]
print(decode_to_sentence(x_train[data_point_to_show]))
print(y_train[data_point_to_show]) # to see the actual sentiment
def encode_sentence(sent):
# print(sent)
encoded = []
word_to_id = get_fixed_word_to_id_dict()
for w in sent.split(" "):
if w in word_to_id:
encoded.append(word_to_id[w])
else:
encoded.append(2) # We used '2' for <UNK>
return encoded
words = "fawn sonja vani made-up-word"
print(encode_sentence(words))
print(encode_sentence("this does not look good"))
print('Pad sequences (samples x time)')
x_train = sequence.pad_sequences(x_train, maxlen=maxlen)
x_test = sequence.pad_sequences(x_test, maxlen=maxlen)
print('x_train shape:', x_train.shape)
print('x_test shape:', x_test.shape)
print(x_train[0])
from tensorflow.python.framework import ops
ops.reset_default_graph()
from tensorflow.keras.models import Sequential
from tensorflow.keras.preprocessing import sequence
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Dropout, Activation
from tensorflow.keras.layers import Embedding
from tensorflow.keras.layers import Conv1D, GlobalMaxPooling1D
from tensorflow.keras.datasets import imdb
# +
print('Build model...')
model = Sequential()
# we start off with an efficient embedding layer which maps
# our vocab indices into embedding_dims dimensions
model.add(Embedding(max_features,
embedding_dims,
input_length=maxlen))
model.add(Dropout(0.2))
# we add a Convolution1D, which will learn filters
# word group filters of size filter_length:
model.add(Conv1D(filters,
kernel_size,
padding='valid',
activation='relu',
strides=1))
# we use max pooling:
model.add(GlobalMaxPooling1D())
# We add a vanilla hidden layer:
model.add(Dense(hidden_dims))
model.add(Dropout(0.2))
model.add(Activation('relu'))
# We project onto a single unit output layer, and squash it with a sigmoid:
model.add(Dense(1))
model.add(Activation('sigmoid'))
model.compile(loss='binary_crossentropy',
optimizer='adam',
metrics=['accuracy'])
model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
validation_data=(x_test, y_test))
score, acc = model.evaluate(x_test, y_test,
batch_size=batch_size)
print('Test score:', score)
print('Test accuracy:', acc)
# -
x_test[0]
predictions = model.predict(x_test)
sentiment = ['NEG' if i < 0.5 else 'POS' for i in predictions]
data_point_to_show = 1
print(decode_to_sentence(x_test[data_point_to_show]), "--", sentiment[data_point_to_show])
# +
test_sentences = []
test_sentence = "Tesla will rise up"
test_sentence = encode_sentence(test_sentence)
test_sentences.append(test_sentence)
test_sentence = "Amazon will go down"
test_sentence = encode_sentence(test_sentence)
test_sentences.append(test_sentence)
test_sentence = "market will crash"
test_sentence = encode_sentence(test_sentence)
test_sentences.append(test_sentence)
test_sentence = "cannot say that i loved it"
test_sentence = encode_sentence(test_sentence)
test_sentences.append(test_sentence)
# -
test_sentences = sequence.pad_sequences(test_sentences, maxlen=maxlen)
test_sentences.shape
# +
predictions = model.predict(test_sentences)
sentiment = ['NEG' if i < 0.5 else 'POS' for i in predictions]
for i in range(test_sentences.shape[0]):
print(decode_to_sentence(test_sentences[i]), "--", sentiment[i])
# -
| sentiment-analysis-begin.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="njn8ffyLuu5h" colab_type="text"
# # Imports and Setup
# Run the Code Below
# + id="eY6RgPDNuj5Q" colab_type="code" outputId="f1baf3ff-9ce0-4153-ca88-5db286b4d9ef" executionInfo={"status": "ok", "timestamp": 1555879408683, "user_tz": 240, "elapsed": 4428, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04091475403023878525"}} colab={"base_uri": "https://localhost:8080/", "height": 145}
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import learning_curve
from sklearn.metrics import accuracy_score
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import torch
import torchvision
from torch.utils.data import DataLoader
import matplotlib.pyplot as plt
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from torch.utils.data import Dataset
import torchvision.transforms as transforms
from torch.autograd import Variable
import torch
import time
# !git clone https://github.com/cis700/hw1-release.git
# !mv hw1-release/dills/* .
# !mv hw1-release hw1
from hw1.helper import Logger
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
print(device)
# + [markdown] id="fNpk9oTzve6p" colab_type="text"
# ## TensorBoard Integration
# + id="Hgos71sOvdp_" colab_type="code" colab={}
# #! rm -r ./logs
LOG_DIR = './logs'
get_ipython().system_raw(
'tensorboard --logdir {} --host 0.0.0.0 --port 6006 &'
.format(LOG_DIR)
)
# !if [ -f ngrok ] ; then echo "Ngrok already installed" ; else wget https://bin.equinox.io/c/4VmDzA7iaHb/ngrok-stable-linux-amd64.zip > /dev/null 2>&1 && unzip ngrok-stable-linux-amd64.zip > /dev/null 2>&1 ; fi
get_ipython().system_raw('./ngrok http 6006 &')
# ! curl -s http://localhost:4040/api/tunnels | python3 -c \
# "import sys, json; print('Tensorboard Link: ' +str(json.load(sys.stdin)['tunnels'][0]['public_url']))"
# + [markdown] id="u_RajyOgvn_J" colab_type="text"
# ## Google Drive Integration
# For Google Drive use only
# + id="QxKL7yGxvuKx" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 146} outputId="bfb3ed2b-176f-450a-c8f8-314b270e0842" executionInfo={"status": "ok", "timestamp": 1555878516745, "user_tz": 240, "elapsed": 21113, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04091475403023878525"}}
from google.colab import drive
drive.mount('/content/drive')
local_dir= 'drive/My Drive/CIS700-004/project'
# %cd 'drive/My Drive/CIS700-004/project'
# + [markdown] id="uC8mUVvNwXSR" colab_type="text"
# # The Code
# + [markdown] id="xMwhR-s9bZWL" colab_type="text"
# ## LSTM
# + id="89jT-97yZvyz" colab_type="code" colab={}
class LSTM(nn.Module):
def __init__(self, input_size, hidden_size, num_layers, bidirectional= False):
super(LSTM, self).__init__()
self.hidden_size= hidden_size
self.num_layers= num_layers
self.lstm= nn.LSTM(input_size, hidden_size,
num_layers, batch_first= True, bidirectional= bidirectional)
self.fc= nn.Linear(hidden_size, 2)
def forward(self, x):
h0 = torch.zeros(self.num_layers, x.size(0), self.hidden_size).to(device)
c0 = torch.zeros(self.num_layers, x.size(0), self.hidden_size).to(device)
out, _ = self.lstm(x, (h0, c0))
out= self.fc(out[:, -1, :]) #hidden state of last time step
return out
lstm = LSTM(300, 400, 3, False).to(device)
# + [markdown] id="oiEIL5VxmDym" colab_type="text"
# ## RunNet
# + id="Rlw8RP2DbeKm" colab_type="code" colab={}
class RunNet():
def __init__(self, net, train_load, test_load, lr= 0.0001, conv= False):
self.logger= Logger('./logs')
self.conv= conv
self.net= net
self.train_load= train_load
self.test_load= test_load
self.criterion = nn.CrossEntropyLoss()
self.optimizer= torch.optim.Adam(self.net.parameters(), lr= lr)
def test_accuracy(self):
test_c= 0
test_t= 0
with torch.no_grad():
for i, data in enumerate(self.test_load, 0):
inputs, labels= data
inputs, labels= Variable(inputs), Variable(labels)
inputs= inputs.to(device)
labels= labels.to(device)
if not self.conv:
inputs = inputs.view(inputs.shape[0], -1)
pred= self.net(inputs)
_, pred= torch.max(pred, 1)
test_c= test_c + int((pred==labels).sum())
test_t= test_t + labels.size(0)
print("Test Acc:", (test_c/test_t))
return test_c/test_t
def run(self, max_epochs= 200, log_every= 10, verbose= True):
step= 0
for epoch in range(max_epochs):
if verbose:
print("Epoch:", epoch)
train_c= 0
train_t= 0
test_c= 0
test_t= 0
for i, data in enumerate(self.train_load, 0):
inputs, labels= data
inputs, labels= Variable(inputs), Variable(labels)
inputs= inputs.to(device)
labels= labels.to(device)
if not self.conv:
inputs = inputs.view(inputs.shape[0], -1)
#feed inputs into network
y_pred = self.net(inputs)
loss= self.criterion(y_pred, labels)
#zero gradient
self.optimizer.zero_grad()
#backprop
loss.backward()
#weight update
self.optimizer.step()
#after updating... report training accuracy
pred= self.net(inputs)
t_loss= self.criterion(pred, labels)
_, pred= torch.max(pred, 1)
train_correct= int((pred==labels).sum())
train_c= train_c + train_correct
train_t = train_t + labels.size(0)
if (step % log_every == 0):
self.logger.scalar_summary("training accuracy", (train_correct/labels.size(0)), step)
self.logger.scalar_summary("loss", t_loss.item(), step)
self.logger.writer.flush()
print("train: %s, loss: %s"%(train_c/train_t, t_loss.item()))
step= step + 1
with torch.no_grad():
for i, data in enumerate(self.test_load, 0):
inputs, labels= data
inputs, labels= Variable(inputs), Variable(labels)
inputs= inputs.to(device)
labels= labels.to(device)
if not self.conv:
inputs = inputs.view(inputs.shape[0], -1)
pred= self.net(inputs)
_, pred= torch.max(pred, 1)
test_correct= int((pred==labels).sum())
test_c= test_c + test_correct
test_t= test_t + labels.size(0)
if verbose:
print("Train Acc:", (train_c/train_t),"Test Acc:", (test_c/test_t),
"Loss:", t_loss.item())
# + [markdown] id="YwP3I60BmgsB" colab_type="text"
# # Test Run
# + id="azhf1iC7mgWf" colab_type="code" colab={}
class CustomDataset(Dataset):
'''
DO NOT EDIT THIS CLASS
'''
def __init__(self, X, y):
self.len = len(X)
self.x_data = torch.from_numpy(X).float()
self.y_data = torch.from_numpy(y).long()
def __len__(self):
return self.len
def __getitem__(self, idx):
return self.x_data[idx], self.y_data[idx]
# + id="v3zXdvjGm7y9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 563} outputId="862da1a3-5112-4f7e-cc66-58e5c5608005" executionInfo={"status": "ok", "timestamp": 1555879960225, "user_tz": 240, "elapsed": 310, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04091475403023878525"}}
x= np.array([np.random.rand(1, 300) for i in range(1000)])
y= np.array([(int)(np.mean(i)/0.5) for i in x])
print(y)
train_data= CustomDataset(x, y)
x2= np.array([np.random.rand(1, 300) for i in range(1000)])
y2= np.array([(int)(np.mean(i)/0.5) for i in x2])
test_data= CustomDataset(x2, y2)
train_load= DataLoader(train_data, batch_size= 100)
test_load= DataLoader(test_data, batch_size= 100)
print(len(train_load.dataset))
print(len(test_load.dataset))
# + id="PyqG-zsWpM4R" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 10926} outputId="5c014581-e194-49b9-c622-c9af1533c282" executionInfo={"status": "ok", "timestamp": 1555880004825, "user_tz": 240, "elapsed": 14419, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04091475403023878525"}}
run_net= RunNet(lstm, train_load, test_load, conv= True)
run_net.run()
| LSTM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from numpy import *
from pylab import *
import scipy
import scipy.integrate as integrate
x = linspace(0,1,1025)
def my_fun(x):
return abs(x-.5)
plot(x,my_fun(x))
n = 17
M = zeros((n,n))
## Hilbert matrix
for i in range(n):
for j in range(n):
M[i,j] = 1/(i+j+1)
F = array([integrate.quad(lambda x: my_fun(x)*x**i,0,1)[0] for i in range(n)]) #[1] is the error
c = linalg.solve(M,F)
p = sum([c[i]*x**i for i in range(n)],axis=0)
plot(x,p)
plot(x,my_fun(x))
from numpy.polynomial.legendre import leggauss
from numpy.polynomial.legendre import Legendre
n=50
N=2*n
q,w = leggauss(N) ## these are correct only for [-1,1]
## rescale for [0,1]
q = .5*(q+1)
w = .5*w
v = array([Legendre.basis(i,domain=[0,1])(x) for i in range(n)])
vq = array([Legendre.basis(i,domain=[0,1])(q) for i in range(n)])
vq.shape
## compute the norms
norms = einsum('ij,ij,j->i',vq,vq,w)
## do the quadrature integral
ci = einsum('ij,j,j->i',vq,my_fun(q),w)/norms
p = einsum('i,ij->j',ci,v)
plot(x,p)
plot(x,my_fun(x))
## I compute simple integrals
#N = 100000
#x = linspace(0,1,N+1)
F=[sin,exp]
sol=[cos(0)-cos(1),e-1]
#h = 1/N;
#sol[1]
# +
def lep_quad(func,down,up,N):
res = 0.
h = abs(up-down)/N
x = linspace(down,up,N+1)
func = func(x)
for i in range(N):
res += func[i]
res *= h
return res
def my_trap_quad(func,down,up,N):
res = 0.
h = abs(up-down)/N
x = linspace(down,up,N+1)
func = func(x)
for i in range(N):
res += (func[i+1]+func[i])
res *= h*0.5
#res = integrate.trapz(func,None,h)
return res
def trap_quad(func,down,up,N):
#res = 0.
x = linspace(down,up,N+1)
func = func(x)
#for i in range(N):
# res += (func[i+1]+func[i])
#res *= h*0.5
res = integrate.trapz(func,x)
return res
# -
def error(func,sol):
errors = zeros((2,5))
for p in range(3,8):
N = 10**p
obj1 = trap_quad(func,0,1,N)
obj2 = my_trap_quad(func,0,1,N)
errors[0][p-3]=abs(obj1-sol)
errors[1][p-3]=abs(obj2-sol)
return errors
import matplotlib.pyplot as plt
arr=error(sin,sol[0])
plt.plot(range(3,8),log(arr[0]),marker='o',label='scipy implementation')
plt.plot(range(3,8),log(arr[1]),marker='o',label='my implementation')
plt.legend()
plt.xlabel('Log( N )')
plt.ylabel('Log( err )')
plt.title('quadrature errors for f(x)=sin(x)')
plt.show()
arr=error(exp,sol[1])
plt.plot(range(3,8),log(arr[0]),marker='o',label='scipy implementation')
plt.plot(range(3,8),log(arr[1]),marker='o',label='my implementation')
plt.legend()
plt.xlabel('Log( N )')
plt.ylabel('Log( err )')
plt.title('quadrature errors for f(x)=e^x')
plt.show()
x0, x1 = 0,1
trap_quad(sin,x0,x1,10**5),integrate.quad(lambda x: sin(x),x0,x1)
N = 10**6
x = linspace(x0,x1,N+1)
integrand = sin(x)
h = abs(x1-x0)/N
integrate.simps(integrand,x),integrate.trapz(integrand,x)
# +
## now I try to integrate 2-dimensional function
sol2=sol[0]**2 ## this will be the exact solution
N = 10**3
x = linspace(x0,x1,N+1)
y = linspace(x0,x1,N+1)
def func(xx,yy):
return sin(xx)*sin(yy)
integrand = func(x[:,None],y[None,:])
integrate.simps(integrate.simps(integrand,y),x),integrate.trapz(integrate.trapz(integrand,y),x),sol2
# -
x = linspace(0,1,1001)
func = sin(x)
h = 1/1000
Dfunc=gradient(func,x)
## The real derivative is a dictionary:
Dict = dict(zip(x,Dfunc))
Dict[1],cos(1)
| my_notebooks/quadrature.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
from glob import glob
xlsxs = glob("xlsx/*.xlsx")
# Convert one xlsx file to Pandas dataframe.
def to_dataframe(f):
return pd.read_excel(
io=f,
header=2,
# 'U' is used as unknown for many columns in the data.
# Seems reasonable to treat that as None/null.
na_values='U'
)
# -
# Check our glob worked
xlsxs
# Concatenate together all spreadsheets.
df = pd.concat([to_dataframe(xlsx) for xlsx in xlsxs])
df
# Check types
df.dtypes
# Output to JSONL for BigQuery
df.to_json(
path_or_buf='nsw-rental-bonds.jsonl',
orient="records",
lines=True,
date_format='iso'
)
# JSONL was too big (BigQuery had a 10MB upload limit). Output to parquet instead, much smaller.
df.to_parquet(
path='nsw-rental-bonds.parquet',
# Otherwise you get a '__index_level_0__' column
index=False,
)
| convert-spreadsheets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from selenium import webdriver
from bs4 import BeautifulSoup
import time
from bs4.element import Tag
import bs4
import json
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import requests
import re
from lxml import html
import lxml
import collections.abc as clct
# %load_ext autoreload
# %autoreload 2
# # Bytes Into Baking
#
# ## Project Overview
# - Food is one of the most common topics on the internet, with content being published on the web by big businesses such as *foodnetwork* and *allrecipes* to home chefs writing their own blogs.
# - Food is trendy--keeping on top of food trends could be valuable to people who work in the food or food publishing industries.
# - The purpose of this project is to gather recipe data from a variety of different websites and then use supervised and unsupervised NLP models to discover useful information from the data
#
# ## Project Phases
# - Create a utility web scraper to capture recipes from a variety of different websites
# - Use ML to compare and contrast recipes
#
# ## Goals
# - Generate a list of target websites for a particular recipe class by pulling urls from Google search
# - Develop a utility scraper to identify recipes and then grab pertinent information about the recipe from the recipe section--starting with instructions
# - Test the results of the scraper on a supervised model that classifies the vectorized recipes
# - Explore the possibiliity of identifying different groupings within a specific recipe class
#
# ## Tools and techniques used in this project
# - **Tools**
# > - Python, Beautiful Soup, Pandas, Numpy, Gensim
# - **Visualization**
# > - Matplotlib, Plotly
# - **Techniques**
# > - Web-scraping, K Fold cross validation, Multinomial Naive Bayes Classification, Non-negative Matrix Factoring (NMF)
#
# ## In this Notebook (phase one of the project)
# - Web recipe scraper
# - Uses Google search to gather target website urls
# - Uses scraper.py to scrape the urls for recipes
#
# ## Step one
# ### Get links for the target search term
# - The websites that feed into the recipe scraper were obtained from a Google search of a specific recipe. This approach was deemed to be superior to relying upon a public index of recipe websites. The purpose of this project is to dive deep into a particular recipe class and hopefully find those hidden gem recipes in that class that may not appear in a public index. For example, one of the best croissant recipe websites can be found on [Gourmetier](https://gourmetier.com/). The recipe on this website is rich with insight written by someone who has made thousands of croissants and has a passion for education. This website does not have the breadth of recipes like *allrecipes* or *foodnetwork*, but the recipes that appear on it are gems. These recipes are more important--as measured by their utility to someone who actually wants to know how to make a great croissant--than those that appear on mass recipe websites. Gourmetier and other 'deep' sites like it absolutely need to be included if they happen to have a recipe in a target class. Google search provided the most likely first step to finding websites and recipes like these.
def get_links_from_google_search(google_url):
'''
Given a Google search string, returns links from that Google search page
Parameters:
----------
google_url (str): a google search string. To get past the first page, replace start=0 with start={bundle}.
Returns:
-------
links (list): a list of links
'''
links_ = []
driver = webdriver.Firefox()
time.sleep(3) # slow requests down to keep google from getting upset
driver.get(google_url)
soup = BeautifulSoup(driver.page_source,'lxml')
result_div = soup.find_all('div', attrs={'class': 'g'})
for r in result_div:
try:
link = r.find('a', href=True)
if link != '':
# site exceptions
if 'google.' not in link and 'costco.' not in link and 'freshdirect' not in link:
links_.append(link['href'])
except Exception as e:
print(e)
continue
return(links_)
def run_remote_google_search(search_string, start=0, num_search_pages=12):
'''
Takes in a preformatted Google search string, a starting number, and num_search_pages which yields 10 urls per search page.
Returns a list of urls that is num_search_pages * 10 long unless it reaches the end of the Google search first.
'''
page_bundles = list(range(start,num_search_pages*10, 10))
links=[]
for bundle in page_bundles:
formatted_string = search_string.replace('start=10', f'start={bundle}')
links_list = get_links_from_google_search(formatted_string)
for link in links_list:
links.append(link)
return links
# +
# To change the search term, run a search in Google, go to the second page of the search, and then copy the Google search address.
# Assign search string to variable.
us_pie_crust = 'https://www.google.com/search?q=how+to+make+croissants&client=ubuntu&hs=KiU&channel=fs&sxsrf=ALeKk02N149o2b-eULZqvU-6oYKKKGnsqA:1598729557213&ei=Va1KX_PLDIO_0PEP9ue5yAU&start=10&sa=N&ved=2ahUKEwizyuXak8HrAhWDHzQIHfZzDlkQ8NMDegQIDhBA&biw=1637&bih=942'
# us_brioche = f'https://www.google.com/search?q=how+to+make+brioche+dough&client=ubuntu&hs=Do3&channel=fs&sxsrf=ALeKk03eqUHI6tXD0_aUHZKqzJUzujjeZQ:1598546700864&ei=DONHX56eNIXB-wS2mZvwBw&start=10&sa=N&ved=2ahUKEwie1IvC6rvrAhWF4J4KHbbMBn4Q8NMDegQIDhBA&biw=1637&bih=942'
# us_puff_pastry = f'https://www.google.com/search?q=how+to+make+puff+pastry+dough&client=ubuntu&hs=GNM&channel=fs&sxsrf=ALeKk011tOIIja512_JwZ8zfTmHSiLc9HA:1598459179879&ei=K41GX96hNcP7-gTrwZCYAg&start=10&sa=N&ved=2ahUKEwie4-q8pLnrAhXDvZ4KHesgBCMQ8NMDegQIDhBA&biw=1637&bih=942'
# us_ciambellone = f'https://www.google.com/search?q=how+to+make+ciambellone&client=ubuntu&hs=GlP&channel=fs&sxsrf=ALeKk03eb_8FF-pe9XNVMG56gUih10WpYA:1598472199741&ei=B8BGX83oLIvJ0PEPr-2ryAQ&start=10&sa=N&ved=2ahUKEwiNiJj91LnrAhWLJDQIHa_2CkkQ8NMDegQIDBBA&biw=1637&bih=942'
# us_google_url=f"https://www.google.com/search?q=croissant+baking+temp&client=ubuntu&hs=bju&channel=fs&tbas=0&sxsrf=ALeKk01tinXVzgWJgZhSeeUfjrJd4FW4oA:1597161338731&ei=er8yX--YLLi50PEPiYu28A4&start=10&sa=N&ved=2ahUKEwjvvfnRwZPrAhW4HDQIHYmFDe4Q8NMDegQIDhBA&biw=1920&bih=969"
# fr_google_url=f"https://www.google.fr/search?q=temp%C3%A9rature+cuisson+croissant&ei=0PEzX4C2PMXk-gTv_ImwCA&start=10&sa=N&ved=2ahUKEwiA3szk5ZXrAhVFsp4KHW9-AoYQ8NMDegQIDRBC&biw=1848&bih=942"
# uk_google_url=f"https://www.google.co.uk/search?q=croissant+temperature&ei=OPQzX8T0KMj4-gSnn5moBw&start=10&sa=N&ved=2ahUKEwjE8JaK6JXrAhVIvJ4KHadPBnUQ8NMDegQIDhA_&biw=1848&bih=942"
pie_crust_links = run_remote_google_search(us_pie_crust, 0, 2)
# -
# Confirm function ran properly and grabbed links
pie_crust_links
# ### Write results to file from Google search function
# +
# CAUTION: Be sure to change the input variable list and the output file name prior to running.
pd.DataFrame(pie_crust_links).to_csv('data/us-piecrust-links.txt')
# -
# ## Step two
# ### A focus on pastry recipes
# - I chose to start with pastry recipes since I come with an indepth knowledge of this space. I've explored hundreds of pastry recipe websites over my career.
# - I chose five categories of pastry and baked goods recipes that had what I considered to be important differences, yet enough similarity that they might confuse a model.
# > - Brioche
# > - Ciambellone
# > - Croissants
# > - Pie crust
# > - Puff pastry
#
# ### Numerous ways to format a recipe, but some intriguing similarities make a utility scraper possible
# - My initial Google search website 'spider' yielded more than recipe websites--I needed a scraper that would limit the possibility of seeing a recipe when one wasn't there
# - More than a third of recipes I encountered follow a schema format promoted by Yoast, a website schema publisher to aid in SEO optimization--useful tags and a json structure with standardized keys for things like recipe ingredients and instructions (Yeah!)
# - Another 40-50% placed the body of their recipe in a script tag with one of several commonly used attributes. This wasn't as clean of an approach--I usually ended up with extra text--but I still was able to get some results
# - The rest, well, some didn't want to be scraped, and some had unusual structures. I could spend a lot of time chasing the tail with limited utility.
# - Results:
#
# | Recipe | First pass links from Google search | Usable recipes obtained | Yield |
# | :---: | :---: | :---: | :---: |
# | Brioche | 112 | 55 | 49% |
# | Ciambellone | 116 | 55 | 47% |
# | Croissant | 180 | 72 | 40% |
# | Puff pastry | 200 | 86 | 43% |
# | Pie crust | 122 | 74 | 60% |
# ### Read in links file and initiate web scrape
# +
# url_list = pd.read_csv('data/fr-croissant-links.txt',header=None)
# url_list = pd.read_csv('data/uk-croissant-links.txt',header=None)
url_piecrust = pd.read_csv('data/us-piecrust-links.txt',header=None)
url_brioche = pd.read_csv('data/us-brioche-links.txt',header=None)
url_puff = pd.read_csv('data/us-puff-pastry-links.txt',header=None)
url_csnt = pd.read_csv('data/us-croissant-links.txt', header=None)
url_cake = pd.read_csv('data/us-ciambellone-links.txt', header=None)
search_string_1 = r'([A-Z][^.]*(?:reheat|ake|oven)[^.]*\d{3}(?:º|°|F| ºF| °F| F|C| ºC| °C| C)[^.]*(?:[.]|[\s]))' #US regex
search_string_2 = r'(\d{3})'
# + jupyter={"outputs_hidden": true}
from src.scraper import *
piecrust_scraper = ScrapeRecipe(page_genus='laminated', page_species='pie_crust', re_pattern_1=search_string_1, re_pattern_2= search_string_2)
req_list = []
for i in range(1,len(url_piecrust[0])):
req_list.append(piecrust_scraper.process_url(url_piecrust[1][i], verbose=True, extract_lang=True))
# + jupyter={"outputs_hidden": true}
req_list
# + jupyter={"outputs_hidden": true}
BeautifulSoup(requests.get(req_list[4].url).content)
# +
good_url, bad_url, instr, target = [], [], [], []
hits, misses = 1,1
for page in req_list:
if page != None:
if page.instructions:
if len(page.instructions)>0:
good_url.append(page.url)
instr.append(page.instructions)
target.append(page.page_species)
hits += 1
else:
print(page.url)
print("NO DATA")
bad_url.append(page.url)
misses += 1
print(f': Hits {hits} Misses {misses}')
df_piecrust = pd.DataFrame([good_url, instr, target]).T
# df = pd.DataFrame([good_url, instr, init_temp]).T
# + jupyter={"outputs_hidden": true}
instr
# -
df_piecrust.iloc[:, 1]
# ### Write search results to file
# +
# Write results to csv
us_piecrust = pd.DataFrame(df_piecrust)
us_piecrust.to_csv('data/us_piecrust.csv')
# us_brioche = pd.DataFrame(df_brioche)
# us_brioche.to_csv('data/us_brioche.csv')
# us_puff = pd.DataFrame(df_puff)
# us_puff.to_csv('data/us_puff.csv')
# us_croissant = pd.DataFrame(df_csnt)
# us_croissant.to_csv('data/us_croissant.csv')
# us_ciambellone = pd.DataFrame(df_cake)
# us_ciambellone.to_csv('data/us_ciambellone.csv')
# + jupyter={"outputs_hidden": true}
us_brioche
| notebooks/1-run-recipe-scrape.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a href="https://www.kaggle.com/alexteboul/tutorial-part-3-cnn-image-modeling-1?scriptVersionId=89713796" target="_blank"><img align="left" alt="Kaggle" title="Open in Kaggle" src="https://kaggle.com/static/images/open-in-kaggle.svg"></a>
# + [markdown] papermill={"duration": 0.042892, "end_time": "2022-03-10T02:28:25.07162", "exception": false, "start_time": "2022-03-10T02:28:25.028728", "status": "completed"} tags=[]
# # Tutorial Part 3: CNN Image Modeling 1
# Hello and welcome to Part 3 of this Pawpularity Contest Tutorial Series.
#
# **In this 'Tutorial Part 3: CNN Image Modeling 1', you'll learn:**
# * How to read in and pre-process the images for modeling
# * How to perform data augmentation in 1 line of code
# * How to build a basic Convolutional Neural Network (CNN)
# * How to predict RMSE using your CNN architecture
# * How to evaluate your models
# * How to submit your predictions for the competition
#
# **Other Tutorials in this series:**
# * In [Tutorial Part 1: EDA for Beginners](https://www.kaggle.com/alexteboul/tutorial-part-1-eda-for-beginners), we covered the exploratory data analysis process from start to finish for the PetFinder.my Pawpularity Contest.
# * In [Tutorial Part 2: Model Building using the Metadata](https://www.kaggle.com/alexteboul/tutorial-part-2-model-building-using-the-metadata), we built models using the metadata (.csv data) provided by the competition hosts. Specifically, we tried Decision Tree Classification, Decision Tree Regression, Ordinary Least Squares Regression, Ridge Regression, Bernoulli Naive Bayes Classification, Random Forest Regression, and Histogram-based Gradient Boosting Regression (LightGBM).
#
# TLDR on the first 2 Tutorials, all the metadata models suck because the metadata isn't strongly predictive of / correlated with the target class "Pawpularity". You could just guess the mean Pawpularity value for every image and get a similar RMSE score - which is what many of the models learned to do. Sometimes it's just not possible to build highly predictive models. Not because of the data you have collected, but because the target class may simply be a random distribution. Pawpularity may just not have much to do with the images themselves. That said, today we are going to see if we can lower our RMSE score by building models using the images. Specifically, we're going to explore basic Convolutional Neural Networks (CNNs) and Transformers.
#
# **Score to beat: 20.52**
#
# **Index:**
# 1. Load in the packages
# 2. Get the data
# 3. Model Building
# + [markdown] papermill={"duration": 0.040792, "end_time": "2022-03-10T02:28:25.156164", "exception": false, "start_time": "2022-03-10T02:28:25.115372", "status": "completed"} tags=[]
# ## 1. Load in the packages
# + papermill={"duration": 6.794538, "end_time": "2022-03-10T02:28:31.991588", "exception": false, "start_time": "2022-03-10T02:28:25.19705", "status": "completed"} tags=[]
#packages
#basics
import os
import numpy as np
import pandas as pd
#images
import cv2
#modeling
from sklearn.model_selection import train_test_split
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers
from tensorflow.keras.regularizers import l2
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from tensorflow.python.client import device_lib
#visualizations
import seaborn as sns
import matplotlib.pyplot as plt
# + papermill={"duration": 0.215746, "end_time": "2022-03-10T02:28:32.246769", "exception": false, "start_time": "2022-03-10T02:28:32.031023", "status": "completed"} tags=[]
#Check to see
tf.config.get_visible_devices()
# + papermill={"duration": 2.266135, "end_time": "2022-03-10T02:28:34.553705", "exception": false, "start_time": "2022-03-10T02:28:32.28757", "status": "completed"} tags=[]
#check if GPU available to use - works well in google colab but kaggle varies. Can use just CPU, GPU, or TPU or mix.
if 'GPU' in str(device_lib.list_local_devices()):
config = tf.compat.v1.ConfigProto(device_count = {'GPU': 0})
sess = tf.compat.v1.Session(config=config)
# + [markdown] papermill={"duration": 0.042101, "end_time": "2022-03-10T02:28:34.680992", "exception": false, "start_time": "2022-03-10T02:28:34.638891", "status": "completed"} tags=[]
# ## 2. Get the data
# The trickiest part of modeling using images in my opinion is often getting them in the right format to be used by your ML model architectures. You need to know where your images are, and then turn them into the data types that can be accepted by your models. If your images are already in directories with their associated classes, there are some straighforward ways to read in the data. Imagine the file paths: /train/26/1239581345.jpg and /train/93/1239581345.jpg where the classes would be a pawpularity score of 26 in the first case and 93 in the second case. That is NOT the case for this competition.
#
# For our purposes, an easy way to get our data is to:
# 1. Get the image paths from train.csv and test.csv files. The image files are named the same as the Ids in the Id column of the csv's.
# 2. Then, create a function that will preprocess all the images and put then into an array/tensor. At a basic level, you want to turn the all training images into one super big array. That is what will be fed into your ML model. So you don't feed the images in based on their image paths/URLs, you have to turn them all into a single array/tensor before the model can be fit to the images.
# 3. Preprocess the images using your function. Display the result.
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" papermill={"duration": 0.094264, "end_time": "2022-03-10T02:28:34.817563", "exception": false, "start_time": "2022-03-10T02:28:34.723299", "status": "completed"} tags=[]
#get the data
train = pd.read_csv('../input/petfinder-pawpularity-score/train.csv')
test = pd.read_csv('../input/petfinder-pawpularity-score/test.csv')
# + papermill={"duration": 0.094102, "end_time": "2022-03-10T02:28:34.956033", "exception": false, "start_time": "2022-03-10T02:28:34.861931", "status": "completed"} tags=[]
#Modify the Id such that each Id is the full image path. In the form
def train_id_to_path(x):
return '../input/petfinder-pawpularity-score/train/' + x + ".jpg"
def test_id_to_path(x):
return '../input/petfinder-pawpularity-score/test/' + x + ".jpg"
#Read in the data and drop unnecessary columns
train = pd.read_csv('../input/petfinder-pawpularity-score/train.csv')
train = train.drop(['Subject Focus', 'Eyes', 'Face', 'Near', 'Action', 'Accessory', 'Group', 'Collage', 'Human', 'Occlusion', 'Info', 'Blur'],axis=1)
test = pd.read_csv('../input/petfinder-pawpularity-score/test.csv')
test = test.drop(['Subject Focus', 'Eyes', 'Face', 'Near', 'Action', 'Accessory', 'Group', 'Collage', 'Human', 'Occlusion', 'Info', 'Blur'],axis=1)
#Add the .jpg extensions to the image file name ids
train["img_path"] = train["Id"].apply(train_id_to_path)
test["img_path"] = test["Id"].apply(test_id_to_path)
# + [markdown] papermill={"duration": 0.041302, "end_time": "2022-03-10T02:28:35.038837", "exception": false, "start_time": "2022-03-10T02:28:34.997535", "status": "completed"} tags=[]
# It may also be helpful for us to build models where we try to turn this into a classification problem. So instead of the model predicting Pawpularity scores between 1-100, it could predict low v. high, or some other arrangement of bins. To do this, you can use pd.qcut(). The q value you specify determins the number of bins/classes the target column will get split up into fairly equally. You can specify labels in a list as well.
#
# For the sake of clarity, I moved this to Tutorial Part 4: CNN Image Modeling 2
# + papermill={"duration": 0.099839, "end_time": "2022-03-10T02:28:35.180143", "exception": false, "start_time": "2022-03-10T02:28:35.080304", "status": "completed"} tags=[]
#binning columns to test models
train['two_bin_pawp'] = pd.qcut(train['Pawpularity'], q=2, labels=False)
train = train.astype({"two_bin_pawp": str})
train['four_bin_pawp'] = pd.qcut(train['Pawpularity'], q=4, labels=False)
train = train.astype({"four_bin_pawp": str})
train['ten_bin_pawp'] = pd.qcut(train['Pawpularity'], q=10, labels=False)
train = train.astype({"ten_bin_pawp": str})
# + [markdown] papermill={"duration": 0.040678, "end_time": "2022-03-10T02:28:35.261989", "exception": false, "start_time": "2022-03-10T02:28:35.221311", "status": "completed"} tags=[]
# Here we can see exactly how the data is split up. If we model on this, we'll still need to at some point turn the predictions back into scores between 1-100 for evaluation though.
# + papermill={"duration": 0.093256, "end_time": "2022-03-10T02:28:35.395914", "exception": false, "start_time": "2022-03-10T02:28:35.302658", "status": "completed"} tags=[]
train2bin_stats = train.groupby('two_bin_pawp')
train2bin_stats.describe()
# + papermill={"duration": 0.085574, "end_time": "2022-03-10T02:28:35.524735", "exception": false, "start_time": "2022-03-10T02:28:35.439161", "status": "completed"} tags=[]
train4bin_stats = train.groupby('four_bin_pawp')
train4bin_stats.describe()
# + papermill={"duration": 0.117194, "end_time": "2022-03-10T02:28:35.68537", "exception": false, "start_time": "2022-03-10T02:28:35.568176", "status": "completed"} tags=[]
train10bin_stats = train.groupby('ten_bin_pawp')
train10bin_stats.describe()
# + papermill={"duration": 0.064283, "end_time": "2022-03-10T02:28:35.793607", "exception": false, "start_time": "2022-03-10T02:28:35.729324", "status": "completed"} tags=[]
#show the full training dataframe now
train.head()
# + papermill={"duration": 0.059105, "end_time": "2022-03-10T02:28:35.89593", "exception": false, "start_time": "2022-03-10T02:28:35.836825", "status": "completed"} tags=[]
#show the full testing dataframe now. Notice how we don't have anything besides the image file paths/ids.
#These are what we will use to make our submissions.
test.head()
# + [markdown] papermill={"duration": 0.050459, "end_time": "2022-03-10T02:28:35.991039", "exception": false, "start_time": "2022-03-10T02:28:35.94058", "status": "completed"} tags=[]
# For the pre-processing function that will get our images, these are some useful documentation links. For our first run through, we'll use the full rgb images. So 3 channels. If you wanted you could change the images to grayscale in this function. We read in the file, then decode the image, normalize, and resize.
# Function path_to_eagertensor() documentation
# * https://www.tensorflow.org/api_docs/python/tf/io/read_file
# * https://www.tensorflow.org/api_docs/python/tf/io/decode_jpeg
# * https://www.tensorflow.org/api_docs/python/tf/cast
# * https://www.tensorflow.org/api_docs/python/tf/image/resize
# * https://www.tensorflow.org/api_docs/python/tf/image/resize_with_pad
# + papermill={"duration": 0.053911, "end_time": "2022-03-10T02:28:36.088559", "exception": false, "start_time": "2022-03-10T02:28:36.034648", "status": "completed"} tags=[]
#Set the size image you want to use
image_height = 128
image_width = 128
#define a function that accepts an image url and outputs an eager tensor
def path_to_eagertensor(image_path):
raw = tf.io.read_file(image_path)
image = tf.image.decode_jpeg(raw, channels=3)
image = tf.cast(image, tf.float32) / 255.0
#image = tf.image.resize_with_pad(image, image_height, image_width) #optional with padding to retain original dimensions
image = tf.image.resize(image, (image_height, image_width))
return image
# + papermill={"duration": 0.054031, "end_time": "2022-03-10T02:28:36.187049", "exception": false, "start_time": "2022-03-10T02:28:36.133018", "status": "completed"} tags=[]
#show the image file path for the first image in the training data
print(train['img_path'][0])
# + papermill={"duration": 0.449513, "end_time": "2022-03-10T02:28:36.681245", "exception": false, "start_time": "2022-03-10T02:28:36.231732", "status": "completed"} tags=[]
#let's plot that first image:
#use plt.imread() to read in that image file
og_example_image = plt.imread('../input/petfinder-pawpularity-score/train/0007de18844b0dbbb5e1f607da0606e0.jpg')
print(og_example_image.shape)
#then plt.imshow() can display it for you
plt.imshow(og_example_image)
plt.title('First Training Image')
plt.axis('off') #turns off the gridlines
plt.show()
# + papermill={"duration": 0.624711, "end_time": "2022-03-10T02:28:37.355268", "exception": false, "start_time": "2022-03-10T02:28:36.730557", "status": "completed"} tags=[]
# %%capture
#run the function to show the pre-processing on the first training image only
example_image = path_to_eagertensor('../input/petfinder-pawpularity-score/train/0007de18844b0dbbb5e1f607da0606e0.jpg')
# + [markdown] papermill={"duration": 0.058088, "end_time": "2022-03-10T02:28:37.501213", "exception": false, "start_time": "2022-03-10T02:28:37.443125", "status": "completed"} tags=[]
# Make note below of the data type that the example_image was turned into after applying our pre-processing funciton. Also note that the shape of the image has been resized.
# + papermill={"duration": 0.315459, "end_time": "2022-03-10T02:28:37.864541", "exception": false, "start_time": "2022-03-10T02:28:37.549082", "status": "completed"} tags=[]
#show the type
print('type: ', type(example_image),'\n shape: ',example_image.shape)
plt.imshow(example_image)
plt.title('First Training Image - with preprocessing done by path_to_eagertensor()')
plt.show()
# + [markdown] papermill={"duration": 0.049718, "end_time": "2022-03-10T02:28:37.96635", "exception": false, "start_time": "2022-03-10T02:28:37.916632", "status": "completed"} tags=[]
# Now, with a simple for loop, you can run the preprocessing function on all the image paths in the training dataframe and next the testing dataframe. By training and testing I mean the data from the training.csv and testing.csv.
# + papermill={"duration": 123.077025, "end_time": "2022-03-10T02:30:41.094722", "exception": false, "start_time": "2022-03-10T02:28:38.017697", "status": "completed"} tags=[]
#get all the images in the training folder and put their tensors in a list
X = []
for img in train['img_path']:
new_img_tensor = path_to_eagertensor(img)
X.append(new_img_tensor)
print(type(X),len(X))
X = np.array(X)
print(type(X),X.shape)
# + papermill={"duration": 0.15171, "end_time": "2022-03-10T02:30:41.296816", "exception": false, "start_time": "2022-03-10T02:30:41.145106", "status": "completed"} tags=[]
#get all the images in the test folder and put their tensors in a list
X_submission = []
for img in test['img_path']:
new_img_tensor = path_to_eagertensor(img)
X_submission.append(new_img_tensor)
print(type(X_submission),len(X_submission))
X_submission = np.array(X_submission)
print(type(X_submission),X_submission.shape)
# + [markdown] papermill={"duration": 0.055699, "end_time": "2022-03-10T02:30:41.405657", "exception": false, "start_time": "2022-03-10T02:30:41.349958", "status": "completed"} tags=[]
# **We've now finished the steps to get our data. As a result, we have:**
# * 2 numpy arrays
# * Array 1 is our training data array composed of all our training preprocessed images (from ../train/*.jpg): X
# * Array 2 is our testing data array composed of the test preprocessed images (from ../test/*.jpg): X_submission
# * We will use X for our model building / training / evaluation of our models
# * We will use X_submission for our submission predictions
# + [markdown] papermill={"duration": 0.054113, "end_time": "2022-03-10T02:30:41.513805", "exception": false, "start_time": "2022-03-10T02:30:41.459692", "status": "completed"} tags=[]
# ## 3. Model Building
# Now that we have our data, building the models themselves is a pretty straightforward process.
#
# At a basic level, you can think of model building with images as having 5 steps:
# 1. Split up your data into training and testing (split up the X numpy array)
# 2. Define a model architecture
# 3. Compile your model
# 4. Fit your model to the training data you have split off in your train_test_split.
# 5. Use your now trained model to predict on new data (X_submission)
# + [markdown] papermill={"duration": 0.05062, "end_time": "2022-03-10T02:30:41.616896", "exception": false, "start_time": "2022-03-10T02:30:41.566276", "status": "completed"} tags=[]
# ### 3.1 Split up you data into training and testing
# + papermill={"duration": 0.063272, "end_time": "2022-03-10T02:30:41.731553", "exception": false, "start_time": "2022-03-10T02:30:41.668281", "status": "completed"} tags=[]
#grab the target variable. In our case, Pawpularity
y = train['Pawpularity']
print(type(y))
# + papermill={"duration": 0.570704, "end_time": "2022-03-10T02:30:42.354989", "exception": false, "start_time": "2022-03-10T02:30:41.784285", "status": "completed"} tags=[]
#generate train - test splits 90% train - 10% test
#You usually don't want to do a 90-10 split unless you have a lot of data,
#but we get to evaluate performance using the leaderboard submissions as well
#So I really want this model to see as many pets as possible in trianing
x_train, x_test, y_train, y_test = train_test_split(X, y, test_size=0.1, random_state=7)
# + papermill={"duration": 0.064349, "end_time": "2022-03-10T02:30:42.472564", "exception": false, "start_time": "2022-03-10T02:30:42.408215", "status": "completed"} tags=[]
#Show the shape of each of the new arrays
print(x_train.shape, x_test.shape, y_train.shape, y_test.shape)
# + [markdown] papermill={"duration": 0.056687, "end_time": "2022-03-10T02:30:42.583391", "exception": false, "start_time": "2022-03-10T02:30:42.526704", "status": "completed"} tags=[]
# ### 3.2 Define your model architecture
# This is the secret sauce of many ML models. It can get quite complicated when you have different layers of your network interacting with eachother. For our example, we'll stick to a basic architecture and explain each step. Basically:
# * [tf.keras.layers.Conv2D](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Conv2D) - defines a convolutional layer and you can mess with parameters
# * [tf.keras.layers.BatchNormalization](https://www.tensorflow.org/api_docs/python/tf/keras/layers/BatchNormalization) - Batch normalization applies a transformation that maintains the mean output close to 0 and the output standard deviation close to 1.
# * [tf.keras.layers.Dropout](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Dropout) - The Dropout layer randomly sets input units to 0 with a frequency of rate at each step during training time, which helps prevent overfitting.
# * [tf.keras.layers.MaxPooling2D](https://www.tensorflow.org/api_docs/python/tf/keras/layers/MaxPool2D) - a way to perform dimensionality reduction by taking the max value in the pool along strides.
# * **strides** - The strides parameter is a 2-tuple of integers, specifying the “step” of the convolution along the x and y axis of the input volume. This can sometimes be used instead of doing max pooling. Reduces volume.
# * **kernel_size** - The size of the kernel. A kernel is a filter that is used to extract the features from the images. The kernel is a matrix that moves over the input data, performs the dot product with the sub-region of input data, and gets the output as the matrix of dot products
# * **padding** - Can be "valid" or "same" (case-insensitive). "valid" means no padding. "same" results in padding with zeros evenly to the left/right or up/down of the input such that output has the same height/width dimension as the input.
# * *kernel_regularizer=l2(0.0002)* - performing regularization can help prevent overfitting. That's when your model is to attuned to the training data and doesn't generalize well. So you might be really good on your training metrics, but then your predictions could be totally off. The smaller the value applied here, the less impact it'll have.
#
# You may also see this look different, where some people like to use .add() to add layers to their model architecture. So you might see something like:
#
# * model_a = tf.keras.models.Sequential()
# * model_a.add(Conv2D(32, (3, 3), activation='relu', input_shape=(128,128,3), data_format="channels_last"))
# * model_a.add(BatchNormalization())
# * model_a.add(MaxPooling2D(pool_size=(2, 2)))
# * model_a.add(Dropout(0.5))
# * model_a.add(Conv2D(32, (3, 3), activation='relu'))
# * model_a.add(Flatten())
# * model_a.add(Dense(32, activation='relu'))
# * model_a.add(BatchNormalization())
# * model_a.add(Dropout(0.5))
# * model_a.add(Dense(1, activation='linear'))
#
# This is acceptable too in most cases. My personal preference is to use this format you'll see coded below.
#
# + papermill={"duration": 0.295548, "end_time": "2022-03-10T02:30:42.933945", "exception": false, "start_time": "2022-03-10T02:30:42.638397", "status": "completed"} tags=[]
#define the inputs to your model. Basically the shape of the incoming data
inputs = tf.keras.Input(shape=(image_height,image_width,3))
#start off with x just being those inputs
x = inputs
x = tf.keras.layers.Conv2D(filters = 16, kernel_size = (7,7), strides = (2,2), padding='valid', kernel_initializer='he_normal', kernel_regularizer=l2(0.0005), activation = 'relu')(x)
x = tf.keras.layers.Conv2D(filters = 32, kernel_size = (3,3), padding='same', activation = 'relu')(x)
x = tf.keras.layers.BatchNormalization()(x)
x = tf.keras.layers.Conv2D(filters = 32, kernel_size = (3,3), strides = (2,2), padding='same', kernel_initializer='he_normal', kernel_regularizer=l2(0.0005), activation = 'relu')(x)
x = tf.keras.layers.BatchNormalization()(x)
x = tf.keras.layers.Dropout(0.25)(x)
x = tf.keras.layers.Conv2D(filters = 64, kernel_size = (3,3), padding='same', kernel_initializer='he_normal', kernel_regularizer=l2(0.0002), activation = 'relu')(x)
x = tf.keras.layers.BatchNormalization()(x)
x = tf.keras.layers.Conv2D(filters = 64, kernel_size = (3,3), strides = (2,2), padding='same', kernel_initializer='he_normal', kernel_regularizer=l2(0.0005), activation = 'relu')(x)
x = tf.keras.layers.BatchNormalization()(x)
x = tf.keras.layers.Dropout(0.25)(x)
x = tf.keras.layers.Conv2D(filters = 128, kernel_size = (3,3), padding='same', kernel_initializer='he_normal', kernel_regularizer=l2(0.0002), activation = 'relu')(x)
x = tf.keras.layers.BatchNormalization()(x)
x = tf.keras.layers.MaxPooling2D(pool_size=(2, 2))(x)
x = tf.keras.layers.Conv2D(filters = 128, kernel_size = (3,3),padding='same', kernel_initializer='he_normal', kernel_regularizer=l2(0.0002), activation = 'relu')(x)
x = tf.keras.layers.BatchNormalization()(x)
x = tf.keras.layers.Dropout(0.25)(x)
x = tf.keras.layers.Flatten()(x)
x = tf.keras.layers.Dense(512, activation = "relu")(x)
x = tf.keras.layers.Dropout(0.5)(x)
output = tf.keras.layers.Dense(1)(x)
model = tf.keras.Model(inputs = inputs, outputs = output)
# + [markdown] papermill={"duration": 0.053858, "end_time": "2022-03-10T02:30:43.041565", "exception": false, "start_time": "2022-03-10T02:30:42.987707", "status": "completed"} tags=[]
# **Use model.summary() to actually show the model architecture**
# + papermill={"duration": 0.08326, "end_time": "2022-03-10T02:30:43.187459", "exception": false, "start_time": "2022-03-10T02:30:43.104199", "status": "completed"} tags=[]
model.summary()
# + [markdown] papermill={"duration": 0.054542, "end_time": "2022-03-10T02:30:43.295924", "exception": false, "start_time": "2022-03-10T02:30:43.241382", "status": "completed"} tags=[]
# Usually you want to use CNNs for classification tasks. And the Pawpularity score is a poor metric to classify given the minimal visual difference between the images at varying Pawpularity scores. A pet scored at 26 pawpularity is not so different from a pet scored at a 27 - it's not like a dog and a car. But we'll do the basics here to show this as a regression type task.
# + [markdown] papermill={"duration": 0.054121, "end_time": "2022-03-10T02:30:43.404632", "exception": false, "start_time": "2022-03-10T02:30:43.350511", "status": "completed"} tags=[]
# ### 3.3 Compile your model
# + papermill={"duration": 0.075168, "end_time": "2022-03-10T02:30:43.533018", "exception": false, "start_time": "2022-03-10T02:30:43.45785", "status": "completed"} tags=[]
#compile the model
model.compile(
loss = 'mse',
optimizer = 'Adam',
metrics = [tf.keras.metrics.RootMeanSquaredError(name="rmse"), "mae", "mape"])
# + [markdown] papermill={"duration": 0.05306, "end_time": "2022-03-10T02:30:43.640297", "exception": false, "start_time": "2022-03-10T02:30:43.587237", "status": "completed"} tags=[]
# ### 3.4 Fit your model using the training data
#
# Just fit the model to the training data. A helpful addition here is usually to do some data augmentation.
# * [tf.keras.preprocessing.image.ImageGenerator](https://www.tensorflow.org/api_docs/python/tf/keras/preprocessing/image/ImageDataGenerator) - This generates batches of tensor image data for real-time data augmentation.
# * Just call your generator.flow() when you .fit() your model to use this real time data augmentation.
# * [tf.keras.Model](https://www.tensorflow.org/api_docs/python/tf/keras/Model) - groups layers into an object with training and inference features.
# + papermill={"duration": 0.073101, "end_time": "2022-03-10T02:30:43.766868", "exception": false, "start_time": "2022-03-10T02:30:43.693767", "status": "completed"} tags=[]
#you can use this to provide your model with different variations of the same images.
# It doesn't actually make and save a bunch of new images, just defines how new images can be created.
# These transformations will randomly be applied to the training images as they get used in training.
data_augmentation = ImageDataGenerator(
rotation_range = 15,
zoom_range = 0.15,
width_shift_range = 0.2,
height_shift_range = 0.2,
shear_range = 0.1,
horizontal_flip = True,
fill_mode = "nearest")
# + [markdown] papermill={"duration": 0.05275, "end_time": "2022-03-10T02:30:43.881327", "exception": false, "start_time": "2022-03-10T02:30:43.828577", "status": "completed"} tags=[]
# When we fit our model here, note that we're using the real-time data augmentation. Also note that we validation_data is the x_test and y_test from our train_test_split before.
# + papermill={"duration": 2348.975753, "end_time": "2022-03-10T03:09:52.910185", "exception": false, "start_time": "2022-03-10T02:30:43.934432", "status": "completed"} tags=[]
history = model.fit(
data_augmentation.flow(x_train,y_train,batch_size=32),
validation_data = (x_test,y_test),
steps_per_epoch = len(x_train) // 32,
epochs = 60
)
# + papermill={"duration": 6.176611, "end_time": "2022-03-10T03:10:04.659474", "exception": false, "start_time": "2022-03-10T03:09:58.482863", "status": "completed"} tags=[]
plt.figure()
plt.plot(history.history["rmse"], label="train_rmse")
plt.plot(history.history["val_rmse"], label="val_rmse")
#plt.xticks(range(0,60))
plt.title("RMSE train/validation by Epoch")
plt.xlabel("Epoch #")
plt.ylabel("RMSE")
plt.legend(loc="upper right")
# + [markdown] papermill={"duration": 6.591889, "end_time": "2022-03-10T03:10:16.783099", "exception": false, "start_time": "2022-03-10T03:10:10.19121", "status": "completed"} tags=[]
# ### 3.5 Now use your newly trained model on unseen data + make your submission!
# + papermill={"duration": 6.143771, "end_time": "2022-03-10T03:10:28.531064", "exception": false, "start_time": "2022-03-10T03:10:22.387293", "status": "completed"} tags=[]
#predict on the submission data
cnn_pred = model.predict(X_submission)
print(X_submission.shape, type(X_submission))
print(cnn_pred.shape, type(cnn_pred))
# + papermill={"duration": 5.87506, "end_time": "2022-03-10T03:10:40.007749", "exception": false, "start_time": "2022-03-10T03:10:34.132689", "status": "completed"} tags=[]
#put the submission predictions alongside their associated Ids
cnn = pd.DataFrame()
cnn['Id'] = test['Id']
cnn['Pawpularity'] = cnn_pred
cnn.to_csv('submission.csv',index=False)
# + papermill={"duration": 6.171394, "end_time": "2022-03-10T03:10:52.09322", "exception": false, "start_time": "2022-03-10T03:10:45.921826", "status": "completed"} tags=[]
cnn.head(10)
# + papermill={"duration": 6.080847, "end_time": "2022-03-10T03:11:03.767145", "exception": false, "start_time": "2022-03-10T03:10:57.686298", "status": "completed"} tags=[]
testing_example_image = plt.imread('../input/petfinder-pawpularity-score/test/4128bae22183829d2b5fea10effdb0c3.jpg')
print(testing_example_image.shape)
#then plt.imshow() can display it for you
plt.imshow(testing_example_image)
plt.title('First Testing Image \n Predicted Pawpularity = {}'.format(cnn['Pawpularity'].iloc[0]))
plt.axis('off') #turns off the gridlines
plt.show()
# + [markdown] papermill={"duration": 5.572142, "end_time": "2022-03-10T03:11:15.18898", "exception": false, "start_time": "2022-03-10T03:11:09.616838", "status": "completed"} tags=[]
# All that for our model to just predict the Pawpularity scores near the median Pawpularity score every single time! 🤦🏻♂️ Well it was seeing random noise, so this is actually a good thing! Yeah that's right, the sample images in the ../test/ subdirectory are all like that. So if you're getting super specific predictions for random noise, something is probably off here lol. That said, the model did just learn to basically guess the median for everything.
#
# Still, better to switch to classification and use the bins we created earlier. Feel free to modify the model architecture, data augmentation, preprocessing, image size, etc to see if you can improve the score. To keep things clean, I'll share the classification and binning methods in a new notebook Tutorial Part 3: CNN Image Modeling 2
# + [markdown] papermill={"duration": 5.58441, "end_time": "2022-03-10T03:11:27.371357", "exception": false, "start_time": "2022-03-10T03:11:21.786947", "status": "completed"} tags=[]
# #### For Fun - see prediction on an image of your choice - here training image #1
# + papermill={"duration": 5.580092, "end_time": "2022-03-10T03:11:38.775171", "exception": false, "start_time": "2022-03-10T03:11:33.195079", "status": "completed"} tags=[]
#Empty List
Sample_image_prediction = []
#Preprocess with our function
sample_new_img_tensor = path_to_eagertensor('../input/petfinder-pawpularity-score/train/0007de18844b0dbbb5e1f607da0606e0.jpg')
Sample_image_prediction.append(sample_new_img_tensor)
Sample_image_prediction = np.array(Sample_image_prediction)
#Show data type is good to input into model
print(type(Sample_image_prediction),Sample_image_prediction.shape)
# + papermill={"duration": 5.661746, "end_time": "2022-03-10T03:11:50.655166", "exception": false, "start_time": "2022-03-10T03:11:44.99342", "status": "completed"} tags=[]
sample_cnn_pred = model.predict(Sample_image_prediction)
print(sample_cnn_pred,sample_cnn_pred.shape, type(sample_cnn_pred))
# + papermill={"duration": 5.704497, "end_time": "2022-03-10T03:12:02.887115", "exception": false, "start_time": "2022-03-10T03:11:57.182618", "status": "completed"} tags=[]
sample_example_image = plt.imread('../input/petfinder-pawpularity-score/train/0007de18844b0dbbb5e1f607da0606e0.jpg')
print(sample_example_image.shape)
#then plt.imshow() can display it for you
plt.imshow(sample_example_image)
plt.title('First Training Image \n Predicted Pawpularity = {}'.format(sample_cnn_pred[0][0]))
plt.axis('off') #turns off the gridlines
plt.show()
| pawpularity-contest/tutorial-part-3-cnn-image-modeling-1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Advanced Features
# ## Table of Contents
# * [1 Position Initialization](#position-initialization)
# * [1.1 Scatter-Initialization](#scatter-initialization)
# * Load data
# * Define search space
# * Initialize and run the optimizer
# * [1.2 Warm-start](#warm-start)
# * Load data
# * Define search space
# * Initialize and run the optimizer
# * [2 Resource Allocation](#resource-allocation)
# * [2.1 Memory](#memory)
# * Load data
# * Define search space
# * Initialize and run the optimizer
# * [3 Weight Initialization](#weight-initialization)
# * [3.1 Transfer-Learning](#transfer-learning)
# * Load data
# * Define search space
# * Initialize and run the optimizer
# +
# TODO
| src/Hyperactive/notebooks/advanced_features.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.6.9 64-bit (''learn-env'': conda)'
# language: python
# name: python36964bitlearnenvcondae7e6328cec2744cc9785efcdf88db667
# ---
# +
import sys
import os
import cv2
from tensorflow.keras.applications.mobilenet_v2 import MobileNetV2, preprocess_input, decode_predictions
# from tensorflow.keras.applications.resnet50 import ResNet50, preprocess_input, decode_predictions
import tensorflow.keras as keras
from tensorflow.keras.models import Model, Sequential
from tensorflow.keras.layers import Input, UpSampling2D, Flatten, BatchNormalization, Dense, Dropout, GlobalAveragePooling2D
from tensorflow.keras import optimizers
from keras.datasets import cifar100
import tensorflow as tf
from keras.utils import np_utils
import numpy as np
import matplotlib.pyplot as plt
import time
from skimage.transform import resize
# from keras.applications.resnet50 import preprocess_input, decode_predictions
from keras.preprocessing.image import ImageDataGenerator
# +
num_classes = 29
nb_epochs = 10
img_sz = (100, 100)
# data paths
train_path = '../../data/asl_alphabet_train/'
validation_path = '../../data/asl_alphabet_validation/'
train_datagen = ImageDataGenerator(preprocessing_function=preprocess_input,
rescale=1./255,
shear_range=0.2,
zoom_range=0.2,
horizontal_flip=True,
validation_split=0.3)
valid_datagen = ImageDataGenerator(preprocessing_function=preprocess_input, rescale=1./255)
train_generator = train_datagen.flow_from_directory(
train_path,
target_size=img_sz,
color_mode='rgb',
batch_size=32,
class_mode='categorical',
subset='training')
test_generator = train_datagen.flow_from_directory(
train_path,
target_size=img_sz,
color_mode='rgb',
batch_size=32,
class_mode='categorical',
subset='validation')
validation_generator = valid_datagen.flow_from_directory(
validation_path,
target_size=img_sz,
color_mode='rgb',
batch_size=32,
class_mode='categorical')
# -
next(train_generator)[0].shape
# +
base_model = MobileNetV2(input_shape=(48,48,3),weights='imagenet',include_top=False) #imports the mobilenet model and discards the last 1000 neuron layer.
x = base_model.output
x = GlobalAveragePooling2D()(x)
x = Dense(1024,activation='relu')(x) #we add dense layers so that the model can learn more complex functions and classify for better results.
x = Dense(1024,activation='relu')(x) #dense layer 2
x = Dense(512,activation='relu')(x) #dense layer 3
preds = Dense(num_classes,activation='softmax')(x) #final layer with softmax activation
model = Model(inputs=base_model.input,outputs=preds)
model.summary()
# -
# for layer in model.layers:
# layer.trainable=False
# or if we want to set the first 20 layers of the network to be non-trainable
for layer in model.layers[:20]:
layer.trainable=False
for layer in model.layers[20:]:
layer.trainable=True
model.compile(optimizers.Adam(learning_rate=0.0001), loss='categorical_crossentropy', metrics=['accuracy'])
# +
early_stop = keras.callbacks.EarlyStopping(monitor="val_accuracy",
min_delta=0,
patience=2,
verbose=0,
mode="auto",
baseline=None,
restore_best_weights=False)
callbacks = [early_stop]
t=time.time()
historytemp = model.fit(train_generator,
steps_per_epoch=len(train_generator),
epochs=10,
validation_data=test_generator,
callbacks=callbacks)
print('Training time: %s' % (t - time.time()))
# +
model.save('models/mobilenet.h5')
model_json = model.to_json()
with open("mobilenet.json", "w") as json_file:
json_file.write(model_json)
# -
| notebooks/post_fi/mobilenet_v2.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.0.4
# language: julia
# name: julia-1.0
# ---
# # Probabilistic Modelling using the Infinite Mixture Model
# In many applications it is desirable to allow the model to adjust its complexity to the amount the data. Consider for example the task of assigning objects into clusters or groups. This task often involves the specification of the number of groups. However, often times it is not known beforehand how many groups exist. Moreover, in some applictions, e.g. modelling topics in text documents or grouping species, the number of examples per group is heavy tailed. This makes it impossible to predefine the number of groups and requiring the model to form new groups when data points from previously unseen groups are observed.
# A natural approach for such applications is the use of non-parametric models. This tutorial will introduce how to use the Dirichlet process in a mixture of infinitely many Gaussians using Turing. For further information on Bayesian nonparametrics and the Dirichlet process we refer to the [introduction by <NAME>](http://mlg.eng.cam.ac.uk/pub/pdf/Gha12.pdf) and the book "Fundamentals of Nonparametric Bayesian Inference" by <NAME> and <NAME>.
using Turing
# ## Mixture Model
# Before introducing infinite mixture models in Turing, we will briefly review the construction of finite mixture models. Subsequently, we will define how to use the [Chinese restaurant process](https://en.wikipedia.org/wiki/Chinese_restaurant_process) construction of a Dirichlet process for non-parametric clustering.
# #### Two-Component Model
# First, consider the simple case of a mixture model with two Gaussian components with fixed covariance.
# The generative process of such a model can be written as:
#
# $$
# \begin{align}
# \pi_1 &\sim Beta(a, b) \\
# \pi_2 &= 1-\pi_1 \\
# \mu_1 &\sim Normal(\mu_0, \Sigma_0) \\
# \mu_2 &\sim Normal(\mu_0, \Sigma_0) \\
# z_i &\sim Categorical(\pi_1, \pi_2) \\
# x_i &\sim Normal(\mu_{z_i}, \Sigma)
# \end{align}
# $$
#
# where $\pi_1, \pi_2$ are the mixing weights of the mixture model, i.e. $\pi_1 + \pi_2 = 1$, and $z_i$ is a latent assignment of the observation $x_i$ to a component (Gaussian).
#
# We can implement this model in Turing for 1D data as follows:
@model two_model(x) = begin
# Hyper-parameters
μ0 = 0.0
σ0 = 1.0
# Draw weights.
π1 ~ Beta(1,1)
π2 = 1-π1
# Draw locations of the components.
μ1 ~ Normal(μ0, σ0)
μ2 ~ Normal(μ0, σ0)
# Draw latent assignment.
z ~ Categorical([π1, π2])
# Draw observation from selected component.
if z == 1
x ~ Normal(μ1, 1.0)
else
x ~ Normal(μ2, 1.0)
end
end
# #### Finite Mixture Model
# If we have more than two components, this model can elegantly be extend using a Dirichlet distribution as prior for the mixing weights $\pi_1, \dots, \pi_K$. Note that the Dirichlet distribution is the multivariate generalization of the beta distribution. The resulting model can be written as:
#
# $$
# \begin{align}
# (\pi_1, \dots, \pi_K) &\sim Dirichlet(K, \alpha) \\
# \mu_k &\sim Normal(\mu_0, \Sigma_0), \;\; \forall k \\
# z &\sim Categorical(\pi_1, \dots, \pi_K) \\
# x &\sim Normal(\mu_z, \Sigma)
# \end{align}
# $$
#
# which resembles the model in the [Gaussian mixture model tutorial](1_GaussianMixtureModel.ipynb) with a slightly different notation.
# ## Infinite Mixture Model
# The question now arises, is there a generalization of a Dirichlet distribution for which the dimensionality $K$ is infinite, i.e. $K = \infty$?
# But first, to implement an infinite Gaussian mixture model in Turing, we first need to load the `Turing.RandomMeasures` module. `RandomMeasures` contains a variety of tools useful in nonparametrics.
using Turing.RandomMeasures
# We now will utilize the fact that one can integrate out the mixing weights in a Gaussian mixture model allowing us to arrive at the Chinese restaurant process construction. See <NAME>: [The Infinite Gaussian Mixture Model](https://www.seas.harvard.edu/courses/cs281/papers/rasmussen-1999a.pdf), NIPS (2000) for details.
#
# In fact, if the mixing weights are integrated out, the conditional prior for the latent variable $z$ is given by:
#
# $$
# p(z_i = k \mid z_{\not i}, \alpha) = \frac{n_k + \alpha/K}{N - 1 + \alpha}
# $$
#
# where $z_{\not i}$ are the latent assignments of all observations except observation $i$. Note that we use $n_k$ to denote the number of observations at component $k$ excluding observation $i$. The parameter $\alpha$ is the concentration parameter of the Dirichlet distribution used as prior over the mixing weights.
# #### Chinese Restaurant Process
# To obtain the Chinese restaurant process construction, we can now derive the conditional prior if $K \rightarrow \infty$.
#
# For $n_k > 0$ we obtain:
#
# $$
# p(z_i = k \mid z_{\not i}, \alpha) = \frac{n_k}{N - 1 + \alpha}
# $$
#
# and for all infinitely many clusters that are empty (combined) we get:
#
# $$
# p(z_i = k \mid z_{\not i}, \alpha) = \frac{\alpha}{N - 1 + \alpha}
# $$
#
# Those equations show that the conditional prior for component assignments is proportional to the number of such observations, meaning that the Chinese restaurant process has a rich get richer property.
# To get a better understanding of this property, we can plot the cluster choosen by for each new observation drawn from the conditional prior.
# +
# Concentration parameter.
α = 10.0
# Random measure, e.g. Dirichlet process.
rpm = DirichletProcess(α)
# Cluster assignments for each observation.
z = Vector{Int}()
# Maximum number of observations we observe.
Nmax = 500
for i in 1:Nmax
# Number of observations per cluster.
K = isempty(z) ? 0 : maximum(z)
nk = Vector{Int}(map(k -> sum(z .== k), 1:K))
# Draw new assignment.
push!(z, rand(ChineseRestaurantProcess(rpm, nk)))
end
# +
using Plots
# Plot the cluster assignments over time
@gif for i in 1:Nmax
scatter(collect(1:i), z[1:i], markersize = 2, xlabel = "observation (i)", ylabel = "cluster (k)", legend = false)
end;
# -
# 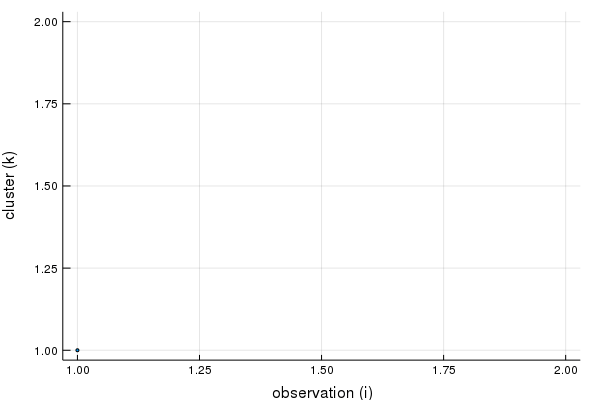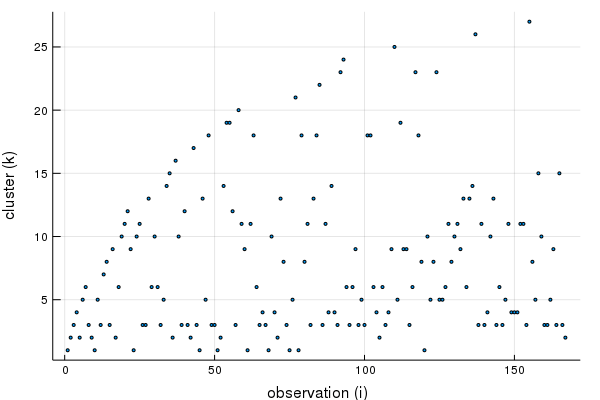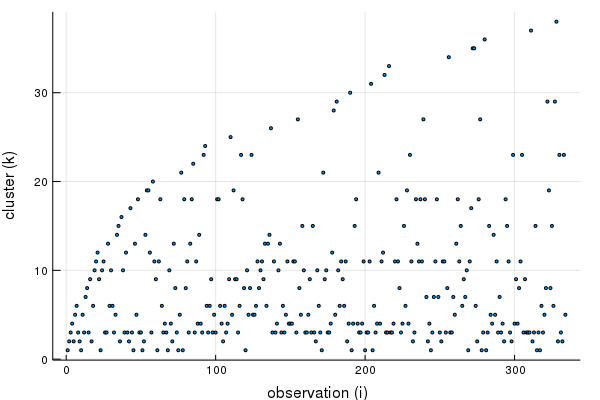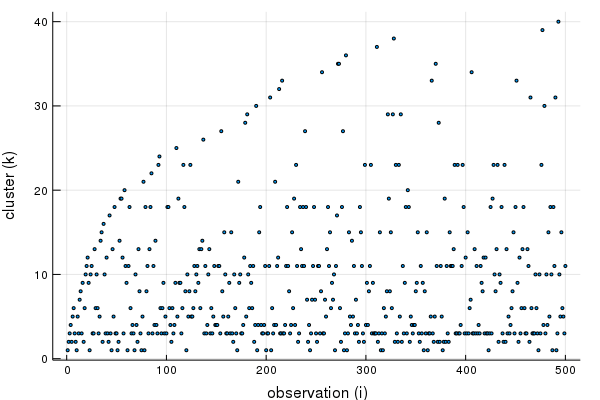
# Further, we can see that the number of clusters is logarithmic in the number of observations and data points. This is a side-effect of the "rich get richer" phenomenon, i.e. we expect large clusters and thus the number of clusters has to be smaller than the number of observations.
#
# $$
# E[K \mid N] \approx \alpha * log \big(1 - \frac{N}{\alpha}\big)
# $$
#
# We can see from the equation that the concetration parameter $\alpha$ allows use to control the number of cluster formed a priori.
# In Turing we can implement an infinite Gaussian mixture model using the Chinese restaurant process construction of a Dirichlet process as follows:
@model infiniteGMM(x) = begin
# Hyper-parameters, i.e. concentration parameter and parameters of H.
α = 1.0
μ0 = 0.0
σ0 = 1.0
# Define random measure, e.g. Dirichlet process.
rpm = DirichletProcess(α)
# Define the base distribution, i.e. expected value of the Dirichlet process.
H = Normal(μ0, σ0)
# Latent assignment.
z = tzeros(Int, length(x))
# Locations of the infinitely many clusters.
μ = tzeros(Float64, 0)
for i in 1:length(x)
# Number of clusters.
K = maximum(z)
nk = Vector{Int}(map(k -> sum(z .== k), 1:K))
# Draw the latent assignment.
z[i] ~ ChineseRestaurantProcess(rpm, nk)
# Create a new cluster?
if z[i] > K
push!(μ, 0.0)
# Draw location of new cluster.
μ[z[i]] ~ H
end
# Draw observation.
x[i] ~ Normal(μ[z[i]], 1.0)
end
end
# We can now use Turing to infer the assignments of some data points. First, we will create some random data that comes from three clusters, with means of 0, -5, and 10.
# +
using Plots, Random
# Generate some test data.
Random.seed!(1)
data = vcat(randn(10), randn(10) .- 5, randn(10) .+ 10)
data .-= mean(data)
data /= std(data);
# -
# Next, we'll sample from our posterior using SMC.
# MCMC sampling
Random.seed!(2)
iterations = 1000
model_fun = infiniteGMM(data);
chain = sample(model_fun, SMC(), iterations);
# Finally, we can plot the number of clusters in each sample.
# +
# Extract the number of clusters for each sample of the Markov chain.
k = map(t -> length(unique(chain[:z].value[t,:,:])), 1:iterations);
# Visualize the number of clusters.
plot(k, xlabel = "Iteration", ylabel = "Number of clusters", label = "Chain 1")
# -
# If we visualize the histogram of the number of clusters sampled from our posterior, we observe that the model seems to prefer 3 clusters, which is the true number of clusters. Note that the number of clusters in a Dirichlet process mixture model is not limited a priori and will grow to infinity with probability one. However, if conditioned on data the posterior will concentrate on a finite number of clusters enforcing the resulting model to have a finite amount of clusters. It is, however, not given that the posterior of a Dirichlet process Gaussian mixture model converges to the true number of clusters, given that data comes from a finite mixture model. See <NAME> and <NAME>: [A simple example of Dirichlet process mixture inconsitency for the number of components](https://arxiv.org/pdf/1301.2708.pdf) for details.
histogram(k, xlabel = "Number of clusters", legend = false)
# One issue with the Chinese restaurant process construction is that the number of latent parameters we need to sample scales with the number of observations. It may be desirable to use alternative constructions in certain cases. Alternative methods of constructing a Dirichlet process can be employed via the following representations:
#
# Size-Biased Sampling Process
#
# $$
# j_k \sim Beta(1, \alpha) * surplus
# $$
#
# Stick-Breaking Process
# $$
# v_k \sim Beta(1, \alpha)
# $$
#
# Chinese Restaurant Process
# $$
# p(z_n = k | z_{1:n-1}) \propto \begin{cases}
# \frac{m_k}{n-1+\alpha}, \text{ if } m_k > 0\\\
# \frac{\alpha}{n-1+\alpha}
# \end{cases}
# $$
#
# For more details see [this article](https://www.stats.ox.ac.uk/~teh/research/npbayes/Teh2010a.pdf).
| 6_InfiniteMixtureModel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import math
import random
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.animation as animation
# +
# Consensus between two colors ("RED" and "BLUE")
class Simulator:
END_EPOCH = 100
def __init__(self, validator_num, slots_per_epoch, fault_tolerance, init_red_honest_ratio, init_red_adversary_ratio):
self.validator_num = validator_num
self.slots_per_epoch = slots_per_epoch
self.fault_tolerance = fault_tolerance
self.validator_id_list = list(range(self.validator_num))
self.adversary_num = math.ceil(self.validator_num * self.fault_tolerance) - 1
self.adversary_ids = set(self.validator_id_list[: self.adversary_num])
self.honest_ids = set(self.validator_id_list[self.adversary_num:])
assert self.validator_num == len(self.honest_ids) + len(self.adversary_ids)
# Initial score of RED (RED is the initial winner)
init_red_honest_ids = set(random.sample(self.honest_ids, math.ceil(self.validator_num * init_red_honest_ratio))) # Honest validators voting for RED
init_red_adversary_ids = set(random.sample(self.adversary_ids, math.ceil(self.validator_num * init_red_adversary_ratio))) # Adversarial validators voting for RED
# Latest votes for two colors
# Index: 0 -> RED, 1 -> BLUE
self.votes = [{"honest_ids": init_red_honest_ids, "adversary_ids": init_red_adversary_ids},
{"honest_ids": self.honest_ids - init_red_honest_ids, "adversary_ids": self.adversary_ids - init_red_adversary_ids}]
# Slot, epoch
self.slot = 0
self.epoch = 0
self.voters_per_slot = self.validator_num // self.slots_per_epoch
self.slot_allocation = self.validator_id_list
# Log for visualization
self.pivots = np.zeros(Simulator.END_EPOCH) # Number of pivots in the epoch
self.scores = np.zeros(int(Simulator.END_EPOCH * self.slots_per_epoch * 2 * 2)).reshape(int(Simulator.END_EPOCH * self.slots_per_epoch), 2, 2) # Index: slot * epoch -> honest/adversary -> red/blue
def get_red_score(self):
return len(self.votes[0]["honest_ids"]) + len(self.votes[0]["adversary_ids"])
def get_blue_score(self):
return len(self.votes[1]["honest_ids"]) + len(self.votes[1]["adversary_ids"])
def get_winner(self):
if self.get_red_score() > self.get_blue_score():
return 0
elif self.get_red_score() == self.get_blue_score() and self.votes[0]["honest_ids"] >= self.votes[1]["honest_ids"]:
# If there is a tie, adversasy assumes the worst tie-breaking
return 0
return 1
def get_current_voters(self):
offset = int(self.voters_per_slot * (self.slot % self.slots_per_epoch))
return set(self.slot_allocation[offset: offset + self.voters_per_slot])
def batch_vote(self, target, validator_ids):
non_target = (target + 1) % 2
self.votes[non_target]["honest_ids"] -= validator_ids
self.votes[non_target]["adversary_ids"] -= validator_ids
self.votes[target]["honest_ids"] |= validator_ids & self.honest_ids
self.votes[target]["adversary_ids"] |= validator_ids & self.adversary_ids
assert self.votes[non_target]["honest_ids"] | self.votes[target]["honest_ids"] == self.honest_ids
assert self.votes[non_target]["adversary_ids"] | self.votes[target]["adversary_ids"] == self.adversary_ids
self.scores[self.slot][0][0] = len(self.votes[0]["honest_ids"])
self.scores[self.slot][0][1] = len(self.votes[1]["honest_ids"])
self.scores[self.slot][1][0] = len(self.votes[0]["adversary_ids"])
self.scores[self.slot][1][1] = len(self.votes[1]["adversary_ids"])
def decoy_flip_flop(self):
winner = self.get_winner()
# Adversary calculates the maximum increase of the honest votes for the winner
# NOTE: This can be upper bounded by the number of validators who havn't voted yet and whose latest votes are not for the winner
honest_vote_num = self.voters_per_slot - len(self.get_current_voters() & self.adversary_ids)
# Pivoting rule
if len(self.votes[winner]["honest_ids"]) + honest_vote_num > math.ceil(self.validator_num / 2) - 1:
new_winner = (winner + 1) % 2
self.batch_vote(new_winner, self.adversary_ids)
assert self.get_winner() != winner, "Pivoting failed!"
# Save the log
self.pivots[self.epoch] += 1
def process_slot(self):
# Before this slot starts, adversary make pivot in some cases
self.decoy_flip_flop()
winner = self.get_winner()
self.batch_vote(winner, self.get_current_voters() & self.honest_ids)
self.slot += 1
def process_epoch(self):
random.shuffle(self.slot_allocation)
self.epoch += 1
def run(self):
random.shuffle(self.slot_allocation)
while self.epoch < Simulator.END_EPOCH:
self.process_slot()
if self.slot % self.slots_per_epoch == 0:
self.process_epoch()
# +
# Parameters
SLOTS_PER_EPOCH = 64
VALIDATOR_NUM = 131072 # Minimum of ETH2.0
assert VALIDATOR_NUM % SLOTS_PER_EPOCH == 0
# Unlucky initial state for adversary
INIT_RED_HONEST_RATIO = 49 / 100 # Must be < 1/2
INIT_RED_ADVERSARY_RATIO = 2 / 100
assert INIT_RED_HONEST_RATIO + INIT_RED_ADVERSARY_RATIO > 1/2
TRIAL_NUM = 92
data_x = np.zeros(TRIAL_NUM)
data_y = np.zeros(TRIAL_NUM)
for i in range(TRIAL_NUM):
FAULT_TOLERANCE = 1 / 3 * (i + 7) / 100
assert INIT_RED_ADVERSARY_RATIO < FAULT_TOLERANCE
s = Simulator(VALIDATOR_NUM, SLOTS_PER_EPOCH, FAULT_TOLERANCE, INIT_RED_HONEST_RATIO, INIT_RED_ADVERSARY_RATIO)
s.run()
data_x[i] = FAULT_TOLERANCE
data_y[i] = (np.sum(s.pivots) - s.epoch) / s.epoch
# +
# %matplotlib inline
plt.title('Adversary ratio and average spending of saving per epoch')
plt.xlabel('Adversary ratio')
plt.ylabel('Spending of saving per epoch')
plt.scatter(data_x, data_y)
# +
plt.title('Adversary ration and average spending of saving per epoch')
plt.xlabel('Adversary ratio')
plt.ylabel('Spending of saving per epoch')
plt.scatter(data_x[70:], data_y[70:])
# -
data_y[70:]
# +
# Public RNG
class Simulator:
END_EPOCH = 100
def __init__(self, validator_num, slots_per_epoch, fault_tolerance, init_red_honest_ratio, init_red_adversary_ratio):
self.validator_num = validator_num
self.slots_per_epoch = slots_per_epoch
self.fault_tolerance = fault_tolerance
self.validator_id_list = list(range(self.validator_num))
self.adversary_num = math.ceil(self.validator_num * self.fault_tolerance) - 1
self.adversary_ids = set(self.validator_id_list[: self.adversary_num])
self.honest_ids = set(self.validator_id_list[self.adversary_num:])
assert self.validator_num == len(self.honest_ids) + len(self.adversary_ids)
# Initial score of RED (RED is the initial winner)
init_red_honest_ids = set(random.sample(self.honest_ids, math.ceil(self.validator_num * init_red_honest_ratio))) # Honest validators voting for RED
init_red_adversary_ids = set(random.sample(self.adversary_ids, math.ceil(self.validator_num * init_red_adversary_ratio))) # Adversarial validators voting for RED
# Latest votes for two colors
# Index: 0 -> RED, 1 -> BLUE
self.votes = [{"honest_ids": init_red_honest_ids, "adversary_ids": init_red_adversary_ids},
{"honest_ids": self.honest_ids - init_red_honest_ids, "adversary_ids": self.adversary_ids - init_red_adversary_ids}]
# Slot, epoch
self.slot = 0
self.epoch = 0
self.voters_per_slot = self.validator_num // self.slots_per_epoch
self.slot_allocation = self.validator_id_list
# Log for visualization
self.pivots = np.zeros(Simulator.END_EPOCH) # Number of pivots in the epoch
self.scores = np.zeros(int(Simulator.END_EPOCH * self.slots_per_epoch * 2 * 2)).reshape(int(Simulator.END_EPOCH * self.slots_per_epoch), 2, 2) # Index: slot * epoch -> honest/adversary -> red/blue
def get_red_score(self):
return len(self.votes[0]["honest_ids"]) + len(self.votes[0]["adversary_ids"])
def get_blue_score(self):
return len(self.votes[1]["honest_ids"]) + len(self.votes[1]["adversary_ids"])
def get_winner(self):
if self.get_red_score() > self.get_blue_score():
return 0
elif self.get_red_score() == self.get_blue_score() and self.votes[0]["honest_ids"] >= self.votes[1]["honest_ids"]:
# If there is a tie, adversasy assumes the worst tie-breaking
return 0
return 1
def get_current_voters(self):
offset = int(self.voters_per_slot * (self.slot % self.slots_per_epoch))
return set(self.slot_allocation[offset: offset + self.voters_per_slot])
def batch_vote(self, target, validator_ids):
non_target = (target + 1) % 2
self.votes[non_target]["honest_ids"] -= validator_ids
self.votes[non_target]["adversary_ids"] -= validator_ids
self.votes[target]["honest_ids"] |= validator_ids & self.honest_ids
self.votes[target]["adversary_ids"] |= validator_ids & self.adversary_ids
assert self.votes[non_target]["honest_ids"] | self.votes[target]["honest_ids"] == self.honest_ids
assert self.votes[non_target]["adversary_ids"] | self.votes[target]["adversary_ids"] == self.adversary_ids
self.scores[self.slot][0][0] = len(self.votes[0]["honest_ids"])
self.scores[self.slot][0][1] = len(self.votes[1]["honest_ids"])
self.scores[self.slot][1][0] = len(self.votes[0]["adversary_ids"])
self.scores[self.slot][1][1] = len(self.votes[1]["adversary_ids"])
def decoy_flip_flop(self):
winner = self.get_winner()
loser = (winner + 1) % 2
# Adversary calculates the maximum increase of the honest votes for the winner
# Assume public RNG
honest_vote_num = len(self.get_current_voters() & self.votes[loser]["honest_ids"])
# Pivoting rule
if len(self.votes[winner]["honest_ids"]) + honest_vote_num > math.ceil(self.validator_num / 2) - 1:
new_winner = loser
self.batch_vote(new_winner, self.adversary_ids)
assert self.get_winner() != winner, "Pivoting failed!"
# Save the log
self.pivots[self.epoch] += 1
def process_slot(self):
# Before this slot starts, adversary make pivot in some cases
self.decoy_flip_flop()
winner = self.get_winner()
self.batch_vote(winner, self.get_current_voters() & self.honest_ids)
self.slot += 1
def process_epoch(self):
random.shuffle(self.slot_allocation)
self.epoch += 1
def run(self):
random.shuffle(self.slot_allocation)
while self.epoch < Simulator.END_EPOCH:
self.process_slot()
if self.slot % self.slots_per_epoch == 0:
self.process_epoch()
def visualize_pivots(trial_num):
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(20, 6))
ax1.set_title('Number of pivots')
ax1.set_xlabel('Epoch')
ax1.set_ylabel('Pivots')
ax2.set_title('Necessary spending of saving of adversary')
ax2.set_xlabel('Epoch')
ax2.set_ylabel('Saving')
for i in range(TRIAL_NUM):
s = Simulator(VALIDATOR_NUM, SLOTS_PER_EPOCH, FAULT_TOLERANCE, INIT_RED_HONEST_RATIO, INIT_RED_ADVERSARY_RATIO)
s.run()
ax1.plot(s.pivots)
ax1.set_ylim(bottom=0, auto=True)
ax2.plot(np.vectorize(lambda x: x - 1)(s.pivots).cumsum())
ax2.set_ylim(bottom=0, auto=True)
# +
# Parameters
SLOTS_PER_EPOCH = 64
VALIDATOR_NUM = 131072 # Minimum of ETH2.0
assert VALIDATOR_NUM % SLOTS_PER_EPOCH == 0
# Unlucky initial state for adversary
INIT_RED_HONEST_RATIO = 49 / 100 # Must be < 1/2
INIT_RED_ADVERSARY_RATIO = 2 / 100
assert INIT_RED_HONEST_RATIO + INIT_RED_ADVERSARY_RATIO > 1/2
TRIAL_NUM = 92
data_x = np.zeros(TRIAL_NUM)
data_y = np.zeros(TRIAL_NUM)
for i in range(TRIAL_NUM):
FAULT_TOLERANCE = 1 / 3 * (i + 7) / 100
assert INIT_RED_ADVERSARY_RATIO < FAULT_TOLERANCE
s = Simulator(VALIDATOR_NUM, SLOTS_PER_EPOCH, FAULT_TOLERANCE, INIT_RED_HONEST_RATIO, INIT_RED_ADVERSARY_RATIO)
s.run()
data_x[i] = FAULT_TOLERANCE
data_y[i] = (np.sum(s.pivots) - s.epoch) / s.epoch
# +
plt.title('Adversary ratio and average spending of saving per epoch')
plt.xlabel('Adversary ratio')
plt.ylabel('Spending of saving per epoch')
plt.scatter(data_x, data_y)
# +
plt.title('Adversary ratio and average spending of saving per epoch')
plt.xlabel('Adversary ratio')
plt.ylabel('Spending of saving per epoch')
plt.scatter(data_x[70:], data_y[70:])
# +
# Adversary larger than 1/3
SLOTS_PER_EPOCH = 64
VALIDATOR_NUM = 131072 # Minimum of ETH2.0
assert VALIDATOR_NUM % SLOTS_PER_EPOCH == 0
INIT_RED_HONEST_RATIO = 45 / 100 # Must be < 1/2
INIT_RED_ADVERSARY_RATIO = 6 / 100
assert INIT_RED_HONEST_RATIO + INIT_RED_ADVERSARY_RATIO > 1/2
TRIAL_NUM = 16
data2_x = np.zeros(TRIAL_NUM)
data2_y = np.zeros(TRIAL_NUM)
for i in range(TRIAL_NUM):
FAULT_TOLERANCE = (33 + i) / 100
assert INIT_RED_ADVERSARY_RATIO < FAULT_TOLERANCE
s = Simulator(VALIDATOR_NUM, SLOTS_PER_EPOCH, FAULT_TOLERANCE, INIT_RED_HONEST_RATIO, INIT_RED_ADVERSARY_RATIO)
s.run()
data2_x[i] = FAULT_TOLERANCE
data2_y[i] = (np.sum(s.pivots) - s.epoch) / s.epoch
# +
plt.title('Adversary ratio and average spending of saving per epoch')
plt.xlabel('Adversary ratio')
plt.ylabel('Spending of saving per epoch')
plt.scatter(data2_x, data2_y)
# -
data2_y
| decoy_flip_flop/adversary_ratio_and_saving.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exercises
# ### 0. Taking note of time
#
# It is useful to take note of the time it takes the algorithm to train. This will provide you with an additional level of understanding.
#
# **Solution**
#
# To calculate the time it takes to train the algorithm you need to add these lines of code to your algorithm:
#
# Before the epoch loop:
#
# import time
#
# start_time = time.time()
#
# After the epoch loop:
#
# print("Training time: %s seconds" % (time.time() - start_time))
#
#
# We will add this line of code to all future exercises.
# ## Deep Neural Network for MNIST Classification
#
# We'll apply all the knowledge from the lectures in this section to write a deep neural network. The problem we've chosen is referred to as the "Hello World" for machine learning because for most students it is their first example. The dataset is called MNIST and refers to handwritten digit recognition. You can find more about it on Yann LeCun's website (Director of AI Research, Facebook). He is one of the pioneers of what we've been talking about and of more complex approaches that are widely used today, such as covolutional networks. The dataset provides 28x28 images of handwritten digits (1 per image) and the goal is to write an algorithm that detects which digit is written. Since there are only 10 digits, this is a classification problem with 10 classes. In order to exemplify what we've talked about in this section, we will build a network with 2 hidden layers between inputs and outputs.
# ## Import the relevant packages
# +
import numpy as np
import tensorflow as tf
from tensorflow.examples.tutorials.mnist import input_data
# TensorFLow includes a data provider for MNIST that we'll use.
# This function automatically downloads the MNIST dataset to the chosen directory.
# The dataset is already split into training, validation, and test subsets.
# Furthermore, it preprocess it into a particularly simple and useful format.
# Every 28x28 image is flattened into a vector of length 28x28=784, where every value
# corresponds to the intensity of the color of the corresponding pixel.
# The samples are grayscale (but standardized from 0 to 1), so a value close to 0 is almost white and a value close to
# 1 is almost purely black. This representation (flattening the image row by row into
# a vector) is slightly naive but as you'll see it works surprisingly well.
# Since this is a classification problem, our targets are categorical.
# Recall from the lecture on that topic that one way to deal with that is to use one-hot encoding.
# With it, the target for each individual sample is a vector of length 10
# which has nine 0s and a single 1 at the position which corresponds to the correct answer.
# For instance, if the true answer is "1", the target will be [0,0,0,1,0,0,0,0,0,0] (counting from 0).
# Have in mind that the very first time you execute this command it might take a little while to run
# because it has to download the whole dataset. Following commands only extract it so they're faster.
mnist = input_data.read_data_sets("MNIST_data/", one_hot=True)
# -
# ## Outline the model
#
# The whole code is in one cell, so you can simply rerun this cell (instead of the whole notebook) and train a new model.
# The tf.reset_default_graph() function takes care of clearing the old parameters. From there on, a completely new training starts.
# +
input_size = 784
output_size = 10
# Use same hidden layer size for both hidden layers. Not a necessity.
hidden_layer_size = 50
# Reset any variables left in memory from previous runs.
tf.reset_default_graph()
# As in the previous example - declare placeholders where the data will be fed into.
inputs = tf.placeholder(tf.float32, [None, input_size])
targets = tf.placeholder(tf.float32, [None, output_size])
# Weights and biases for the first linear combination between the inputs and the first hidden layer.
# Use get_variable in order to make use of the default TensorFlow initializer which is Xavier.
weights_1 = tf.get_variable("weights_1", [input_size, hidden_layer_size])
biases_1 = tf.get_variable("biases_1", [hidden_layer_size])
# Operation between the inputs and the first hidden layer.
# We've chosen ReLu as our activation function. You can try playing with different non-linearities.
outputs_1 = tf.nn.relu(tf.matmul(inputs, weights_1) + biases_1)
# Weights and biases for the second linear combination.
# This is between the first and second hidden layers.
weights_2 = tf.get_variable("weights_2", [hidden_layer_size, hidden_layer_size])
biases_2 = tf.get_variable("biases_2", [hidden_layer_size])
# Operation between the first and the second hidden layers. Again, we use ReLu.
outputs_2 = tf.nn.relu(tf.matmul(outputs_1, weights_2) + biases_2)
# Weights and biases for the final linear combination.
# That's between the second hidden layer and the output layer.
weights_3 = tf.get_variable("weights_3", [hidden_layer_size, output_size])
biases_3 = tf.get_variable("biases_3", [output_size])
# Operation between the second hidden layer and the final output.
# Notice we have not used an activation function because we'll use the trick to include it directly in
# the loss function. This works for softmax and sigmoid with cross entropy.
outputs = tf.matmul(outputs_2, weights_3) + biases_3
# Calculate the loss function for every output/target pair.
# The function used is the same as applying softmax to the last layer and then calculating cross entropy
# with the function we've seen in the lectures. This function, however, combines them in a clever way,
# which makes it both faster and more numerically stable (when dealing with very small numbers).
# Logits here means: unscaled probabilities (so, the outputs, before they are scaled by the softmax)
# Naturally, the labels are the targets.
loss = tf.nn.softmax_cross_entropy_with_logits(logits=outputs, labels=targets)
# Get the average loss
mean_loss = tf.reduce_mean(loss)
# Define the optimization step. Using adaptive optimizers such as Adam in TensorFlow
# is as simple as that.
optimize = tf.train.AdamOptimizer(learning_rate=0.001).minimize(mean_loss)
# Get a 0 or 1 for every input in the batch indicating whether it output the correct answer out of the 10.
out_equals_target = tf.equal(tf.argmax(outputs, 1), tf.argmax(targets, 1))
# Get the average accuracy of the outputs.
accuracy = tf.reduce_mean(tf.cast(out_equals_target, tf.float32))
# Declare the session variable.
sess = tf.InteractiveSession()
# Initialize the variables. Default initializer is Xavier.
initializer = tf.global_variables_initializer()
sess.run(initializer)
# Batching
batch_size = 100
# Calculate the number of batches per epoch for the training set.
batches_number = mnist.train._num_examples // batch_size
# Basic early stopping. Set a miximum number of epochs.
max_epochs = 15
# Keep track of the validation loss of the previous epoch.
# If the validation loss becomes increasing, we want to trigger early stopping.
# We initially set it at some arbitrarily high number to make sure we don't trigger it
# at the first epoch
prev_validation_loss = 9999999.
import time
start_time = time.time()
# Create a loop for the epochs. Epoch_counter is a variable which automatically starts from 0.
for epoch_counter in range(max_epochs):
# Keep track of the sum of batch losses in the epoch.
curr_epoch_loss = 0.
# Iterate over the batches in this epoch.
for batch_counter in range(batches_number):
# Input batch and target batch are assigned values from the train dataset, given a batch size
input_batch, target_batch = mnist.train.next_batch(batch_size)
# Run the optimization step and get the mean loss for this batch.
# Feed it with the inputs and the targets we just got from the train dataset
_, batch_loss = sess.run([optimize, mean_loss],
feed_dict={inputs: input_batch, targets: target_batch})
# Increment the sum of batch losses.
curr_epoch_loss += batch_loss
# So far curr_epoch_loss contained the sum of all batches inside the epoch
# We want to find the average batch losses over the whole epoch
# The average batch loss is a good proxy for the current epoch loss
curr_epoch_loss /= batches_number
# At the end of each epoch, get the validation loss and accuracy
# Get the input batch and the target batch from the validation dataset
input_batch, target_batch = mnist.validation.next_batch(mnist.validation._num_examples)
# Run without the optimization step (simply forward propagate)
validation_loss, validation_accuracy = sess.run([mean_loss, accuracy],
feed_dict={inputs: input_batch, targets: target_batch})
# Print statistics for the current epoch
# Epoch counter + 1, because epoch_counter automatically starts from 0, instead of 1
# We format the losses with 3 digits after the dot
# We format the accuracy in percentages for easier interpretation
print('Epoch '+str(epoch_counter+1)+
'. Mean loss: '+'{0:.3f}'.format(curr_epoch_loss)+
'. Validation loss: '+'{0:.3f}'.format(validation_loss)+
'. Validation accuracy: '+'{0:.2f}'.format(validation_accuracy * 100.)+'%')
# Trigger early stopping if validation loss begins increasing.
if validation_loss > prev_validation_loss:
break
# Store this epoch's validation loss to be used as previous validation loss in the next iteration.
prev_validation_loss = validation_loss
# Not essential, but it is nice to know when the algorithm stopped working in the output section, rather than check the kernel
print('End of training.')
#Add the time it took the algorithm to train
print("Training time: %s seconds" % (time.time() - start_time))
# -
# ## Test the model
#
# As we discussed in the lectures, after training on the training and validation sets, we test the final prediction power of our model by running it on the test dataset that the algorithm has not seen before.
#
# It is very important to realize that fiddling with the hyperparameters overfits the validation dataset. The test is the absolute final instance. You should not test before you are completely done with adjusting your model.
# +
input_batch, target_batch = mnist.test.next_batch(mnist.test._num_examples)
test_accuracy = sess.run([accuracy],
feed_dict={inputs: input_batch, targets: target_batch})
# Test accuracy is a list with 1 value, so we want to extract the value from it, using x[0]
# Uncomment the print to see how it looks before the manipulation
# print (test_accuracy)
test_accuracy_percent = test_accuracy[0] * 100.
# Print the test accuracy formatted in percentages
print('Test accuracy: '+'{0:.2f}'.format(test_accuracy_percent)+'%')
# -
# Using the initial model and hyperparameters given in this notebook, the final test accuracy should be roughly between 97% and 98%. Each time the code is rerunned, we get a different accuracy as the batches are shuffled, the weights are initialized in a different way, etc.
#
# Finally, we have intentionally reached a suboptimal solution, so you can have space to build on it.
| course_2/course_material/Part_7_Deep_Learning/S54_L390/0. TensorFlow_MNIST_take_note_of_time_Solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python3
# ---
# <a href="http://cocl.us/pytorch_link_top">
# <img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/DL0110EN/notebook_images%20/Pytochtop.png" width="750" alt="IBM Product " />
# </a>
#
# <img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/DL0110EN/notebook_images%20/cc-logo-square.png" width="200" alt="cognitiveclass.ai logo" />
#
# <h1>Linear Regression 1D: Training One Parameter</h1>
#
# <h2>Objective</h2><ul><li> How to create cost or criterion function using MSE (Mean Square Error).</li></ul>
#
# <h2>Table of Contents</h2>
# <p>In this lab, you will train a model with PyTorch by using data that you created. The model only has one parameter: the slope.</p>
#
# <ul>
# <li><a href="#Makeup_Data">Make Some Data</a></li>
# <li><a href="#Model_Cost">Create the Model and Cost Function (Total Loss)</a></li>
# <li><a href="#Train">Train the Model</a></li>
# </ul>
# <p>Estimated Time Needed: <strong>20 min</strong></p>
#
# <hr>
#
# <h2>Preparation</h2>
#
# The following are the libraries we are going to use for this lab.
#
# +
# These are the libraries will be used for this lab.
import numpy as np
import matplotlib.pyplot as plt
# -
# The class <code>plot_diagram</code> helps us to visualize the data space and the parameter space during training and has nothing to do with PyTorch.
#
# +
# The class for plotting
class plot_diagram():
# Constructor
def __init__(self, X, Y, w, stop, go = False):
start = w.data
self.error = []
self.parameter = []
self.X = X.numpy()
self.Y = Y.numpy()
self.parameter_values = torch.arange(start, stop)
self.Loss_function = [criterion(forward(X), Y) for w.data in self.parameter_values]
w.data = start
# Executor
def __call__(self, Yhat, w, error, n):
self.error.append(error)
self.parameter.append(w.data)
plt.subplot(212)
plt.plot(self.X, Yhat.detach().numpy())
plt.plot(self.X, self.Y,'ro')
plt.xlabel("A")
plt.ylim(-20, 20)
plt.subplot(211)
plt.title("Data Space (top) Estimated Line (bottom) Iteration " + str(n))
plt.plot(self.parameter_values.numpy(), self.Loss_function)
plt.plot(self.parameter, self.error, 'ro')
plt.xlabel("B")
plt.figure()
# Destructor
def __del__(self):
plt.close('all')
# -
# <!--Empty Space for separating topics-->
#
# <h2 id="Makeup_Data">Make Some Data</h2>
#
# Import PyTorch library:
#
# Import the library PyTorch
# !pip install pytorch
import torch
# Generate values from -3 to 3 that create a line with a slope of -3. This is the line you will estimate.
#
# +
# Create the f(X) with a slope of -3
X = torch.arange(-3, 3, 0.1).view(-1, 1)
f = -3 * X
# -
# Let us plot the line.
#
# +
# Plot the line with blue
plt.plot(X.numpy(), f.numpy(), label = 'f')
plt.xlabel('x')
plt.ylabel('y')
plt.legend()
plt.show()
# -
# Let us add some noise to the data in order to simulate the real data. Use <code>torch.randn(X.size())</code> to generate Gaussian noise that is the same size as <code>X</code> and has a standard deviation opf 0.1.
#
# +
# Add some noise to f(X) and save it in Y
Y = f + 0.1 * torch.randn(X.size())
# -
# Plot the <code>Y</code>:
#
# +
# Plot the data points
plt.plot(X.numpy(), Y.numpy(), 'rx', label = 'Y')
plt.plot(X.numpy(), f.numpy(), label = 'f')
plt.xlabel('x')
plt.ylabel('y')
plt.legend()
plt.show()
# -
# <!--Empty Space for separating topics-->
#
# <h2 id="Model_Cost">Create the Model and Cost Function (Total Loss)</h2>
#
# In this section, let us create the model and the cost function (total loss) we are going to use to train the model and evaluate the result.
#
# First, define the <code>forward</code> function $y=w\*x$. (We will add the bias in the next lab.)
#
# +
# Create forward function for prediction
def forward(x):
return w * x
# -
# Define the cost or criterion function using MSE (Mean Square Error):
#
# +
# Create the MSE function for evaluate the result.
def criterion(yhat, y):
return torch.mean((yhat - y) ** 2)
# -
# Define the learning rate <code>lr</code> and an empty list <code>LOSS</code> to record the loss for each iteration:
#
# +
# Create Learning Rate and an empty list to record the loss for each iteration
lr = 0.1
LOSS = []
# -
# Now, we create a model parameter by setting the argument <code>requires_grad</code> to <code> True</code> because the system must learn it.
#
w = torch.tensor(-10.0, requires_grad = True)
# Create a <code>plot_diagram</code> object to visualize the data space and the parameter space for each iteration during training:
#
gradient_plot = plot_diagram(X, Y, w, stop = 5)
# <!--Empty Space for separating topics-->
#
# <h2 id="Train">Train the Model</h2>
#
# Let us define a function for training the model. The steps will be described in the comments.
#
# +
# Define a function for train the model
def train_model(iter):
for epoch in range (iter):
# make the prediction as we learned in the last lab
Yhat = forward(X)
# calculate the iteration
loss = criterion(Yhat,Y)
# plot the diagram for us to have a better idea
gradient_plot(Yhat, w, loss.item(), epoch)
# store the loss into list
LOSS.append(loss.item())
# backward pass: compute gradient of the loss with respect to all the learnable parameters
loss.backward()
# updata parameters
w.data = w.data - lr * w.grad.data
# zero the gradients before running the backward pass
w.grad.data.zero_()
# -
# Let us try to run 4 iterations of gradient descent:
#
# +
# Give 4 iterations for training the model here.
train_model(4)
# -
# Plot the cost for each iteration:
#
# +
# Plot the loss for each iteration
plt.plot(LOSS)
plt.tight_layout()
plt.xlabel("Epoch/Iterations")
plt.ylabel("Cost")
# -
# <!--Empty Space for separating topics-->
#
# <h3>Practice</h3>
#
# Create a new learnable parameter <code>w</code> with an initial value of -15.0.
#
# +
# Practice: Create w with the inital value of -15.0
# Type your code here
w = torch.tensor(-15.0, requires_grad=True)
# -
# Double-click <b>here</b> for the solution.
#
# <!-- Your answer is below:
# w = torch.tensor(-15.0, requires_grad=True)
# -->
#
# <!--Empty Space for separating topics-->
#
# Create an empty list <code>LOSS2</code>:
#
# +
# Practice: Create LOSS2 list
# Type your code here
LOSS2 = []
# -
# Double-click <b>here</b> for the solution.
#
# <!-- Your answer is below:
# LOSS2 = []
# -->
#
# <!--Empty Space for separating topics-->
#
# Write your own <code>my_train_model</code> function with loss list <code>LOSS2</code>. And run it with 4 iterations.
#
# +
# Practice: Create your own my_train_model
gradient_plot1 = plot_diagram(X, Y, w, stop = 15)
def my_train_model(iter):
for epoch in range (iter):
Yhat = forward(X)
loss = criterion(Yhat,Y)
gradient_plot1(Yhat, w, loss.item(), epoch)
LOSS2.append(loss)
loss.backward()
w.data = w.data - lr * w.grad.data
w.grad.data.zero_(
# -
# Double-click <b>here</b> for the solution.
#
# <!-- Your answer is below:
#
# def my_train_model(iter):
# for epoch in range (iter):
# Yhat = forward(X)
# loss = criterion(Yhat,Y)
# gradient_plot1(Yhat, w, loss.item(), epoch)
# LOSS2.append(loss)
# loss.backward()
# w.data = w.data - lr * w.grad.data
# w.grad.data.zero_()
# my_train_model(4)
# -->
#
# <!--Empty Space for separating topics-->
#
# Plot an overlay of the list <code>LOSS2</code> and <code>LOSS</code>.
#
# +
# Practice: Plot the list LOSS2 and LOSS
plt.plot(LOSS, label = "LOSS")
plt.plot(LOSS2, label = "LOSS2")
plt.tight_layout()
plt.xlabel("Epoch/Iterations")
plt.ylabel("Cost")
plt.legend()
# -
# Double-click <b>here</b> for the solution.
#
# <!-- Your answer is below:
# plt.plot(LOSS, label = "LOSS")
# plt.plot(LOSS2, label = "LOSS2")
# plt.tight_layout()
# plt.xlabel("Epoch/Iterations")
# plt.ylabel("Cost")
# plt.legend()
# -->
#
# <!--Empty Space for separating topics-->
#
# What does this tell you about the parameter value?
#
# Double-click <b>here</b> for the solution.
#
# <!-- Your answer is below:
# the parameter value is sensitive to initialization
# -->
#
# <!--Empty Space for separating topics-->
#
# <a href="http://cocl.us/pytorch_link_bottom">
# <img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/DL0110EN/notebook_images%20/notebook_bottom%20.png" width="750" alt="PyTorch Bottom" />
# </a>
#
# <h2>About the Authors:</h2>
#
# <a href="https://www.linkedin.com/in/joseph-s-50398b136/"><NAME></a> has a PhD in Electrical Engineering, his research focused on using machine learning, signal processing, and computer vision to determine how videos impact human cognition. Joseph has been working for IBM since he completed his PhD.
#
# Other contributors: <a href="https://www.linkedin.com/in/michelleccarey/"><NAME></a>, <a href="www.linkedin.com/in/jiahui-mavis-zhou-a4537814a"><NAME></a>
#
# ## Change Log
#
# | Date (YYYY-MM-DD) | Version | Changed By | Change Description |
# | ----------------- | ------- | ---------- | ----------------------------------------------------------- |
# | 2020-09-21 | 2.0 | Shubham | Migrated Lab to Markdown and added to course repo in GitLab |
#
# <hr>
#
# Copyright © 2018 <a href="cognitiveclass.ai?utm_source=bducopyrightlink&utm_medium=dswb&utm_campaign=bdu">cognitiveclass.ai</a>. This notebook and its source code are released under the terms of the <a href="https://bigdatauniversity.com/mit-license/">MIT License</a>.
#
| pyTorch-Week2-LinearRegression1DPrediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Dot Product (Pipelined)
# %pylab inline
from gps_helper.prn import PRN
from sk_dsp_comm import sigsys as ss
from sk_dsp_comm import digitalcom as dc
from caf_verilog.dot_prod_pip import DotProdPip
x = array([-1, -1, 1, 1])
y = array([1, 1, -1, -1])
dot(x, y.T)
# ## Test Signals
prn = PRN(15)
prn2 = PRN(20)
fs = 625e3
Ns = fs / 125e3
prn_seq = prn.prn_seq()
prn_seq2 = prn2.prn_seq()
prn_seq,b = ss.NRZ_bits2(array(prn_seq), Ns)
prn_seq2,b2 = ss.NRZ_bits2(array(prn_seq2), Ns)
# ## Dot Product Pipeline Implementation
dpp = DotProdPip(prn_seq[:8], prn_seq[:8] * -1)
dpp.gen_tb()
dpp.sum_i_bits
| docs/source/nb_examples/DotProductPipeline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Clustering Stocks with Python
#
# Often we are given data sets that contain ungrouped or uncategorized data; in these cases, we may want to find some underlying structure that might not be apparent to the naked eyes. When we want to group objects into similar sets, we are said to be clustering. **The idea is relatively simple, objects that are more similar in nature should exist in the same group, and we use different measures of similarity to determine whether an object is similar to another.** While clustering as a concept is easy to grasp, there are different implementations to achieve this. Some applications use different algorithms or use various measures to define similarity.
#
# **In this tutorial, we will do a clustering exercise where we will take a group of stocks and cluster them into different groups based on specific financial metrics.** Clustering is used extensively in the financial industry to do a wide range of tasks, spanning from portfolio construction, outlier detection, or stock selection.
#
# Often, portfolio managers devise strategies to select stocks that build risk-adjusted portfolios; these are portfolios that minimize risk for a given level of return. However, to do this, they must choose stocks that are not correlated with each other or in some cases to find stocks that are similar in nature to give them adequate exposure to a particular segment of the market.
#
# Now, as mentioned above, there are many clustering algorithms we can use, but in this tutorial, we will use K-Means. **K-Means is an unsupervised machine learning algorithm which is used on unlabeled data (i.e., data without defined categories or groups). The goal of the K-Means algorithm is to divide `n` data points into `k` partitions, where the sum of the distances is minimized.**
#
# ***
#
# **Broken into steps, the algorithm is executed in the following order:**
#
# 1. Randomly select K-Centers, to be the cluster centers.
# 2. Calculate the distance of each point to a cluster, and assign a cluster label where the Euclidean distance is smallest.
# 3. Recompute the centroids by taking the mean of all the data points assigned to that cluster.
# 4. Repeat these steps until one of the following two conditions are met:
# 1. *The Sum of the Distances is minimized.*
# 2. *The maximum number of iterations has been reached.*
#
#
# ***
#
# The algorithm will converge to a result, but this result is not necessarily guaranteed to be the optimal result. K-Means is used heavily for exploratory data analysis and for its ability to take unstructured data and create structure from it. This is powerful when it comes to finding patterns in data that aren't necessarily apparent to the naked eye, especially at higher dimensions.
#
# ## Libraries
#
# Luckily, performing cluster analysis in Python is easy due to built-in libraries that make running the algorithms efficient and fast. In this tutorial, we will use `pandas` to take data collected from our API and store it in a manner that makes manipulation manageable. To perform our cluster analysis, we will use `sklearn` which has built-in functions to create instances of our clustering algorithms. Along with this, we can perform a model evaluation with built-in metrics that will take our results and evaluate them.
#
# For some visualization tasks, we will use `matplotlib` and `yellowbrick` to create cluster graphs and silhouette graphs, respectively.
import requests
import pandas as pd
# ## Step One: Load the Stock Symbols
# I have a CSV file that contains one column of information which is stock symbols. Each stock symbol will be used for a request in the TD Ameritrade API to collect fundamental stock data and specific financial metrics. We will call pandas `read_csv` method to load the file, and store it in a data frame.
# +
# load the stock symbols into a data frame.
stock_symbols = pd.read_csv(r"C:\Users\Alex\OneDrive\Growth - Tutorial Videos\Lessons - Python\Python For Finance\stock_list.csv")
# we have a character that will cause issues in our request so we have to remove it
stock_symbols['Symbol'] = stock_symbols['Symbol'].str.replace('^','')
# display the number of rows.
display(stock_symbols.shape)
# -
# With over 5800, stocks, we need to break this data frame into chunks so that we can request multiple symbols at once. While TD Ameritrade's **Search Instruments** endpoint does allow for numerous stock requests at once, we will have errors take place if we exceed 100, so let's break the data frame into chunks of 100.
# +
# define the size of the chunk
n = 100
# for the sake of completeness
# this defines a chunk, for example if i = 1 and n = 100 we would select rows 1 to 101 stock_symbosl[1:101]
# stock_symbols[i:i+n]
# this defines the whole range of the data frame. Start at 0, go to the last row(stock_symbols.shape[0]),
# and take a step of n in this case 100
# range( 0, stock_symbols.shape[0], n)
# break the data frame into chunks
symbols_chunk = [stock_symbols[i:i+n] for i in range( 0, stock_symbols.shape[0], n)]
# +
# grab an example so you can see the output
example_chunk = list(symbols_chunk[0]['Symbol'])
# show the first five items
example_chunk[:5]
# -
# ## Step Two: Collect the Data
# ***
# Alright, we have a list of symbols so let's move to the next part collecting the data for each stock symbol. For those of us who have a TD Ameritrade account, we are in luck because they have a free API we can use to collect all sorts of data on different financial instruments. This particular API was very popular among viewers, so if you would like to learn more about all the endpoints and how to get set up, I encourage you to watch my series on YouTube which can be found here:
#
# **https://www.youtube.com/playlist?list=PLcFcktZ0wnNnrgVvY_87ZRXRlac6Pciur**
#
# The general idea, is we will loop through each chunk in our list, make a request using the symbols in that list, convert it to a dictionary object, grab the fundamental data that was sent back, store it in our master dictionary, and then finally take our master dictionary and convert it into a pandas data frame.
# +
# define an endpoint, in this case we will use the instruments endpoint
endpoint = r"https://api.tdameritrade.com/v1/instruments"
# we need a place to store all of our data, so a dictionary will do.
stock_dict = {}
# loop through each chunk
for chunk in symbols_chunk:
# define the payload
payload = {'apikey':'SIGMA1192',
'projection':'fundamental',
'symbol':list(chunk['Symbol'])}
# make a request
content = requests.get(url = endpoint, params = payload)
# convert it dictionary object
data = content.json()
# the ones that do exist, loop through each stock, grab the data, and store it in the dictionary
try:
for stock in data:
stock_dict[data[stock]['cusip']] = data[stock]['fundamental']
except:
continue
# create a data frame with the newly collected data.
stock_df = pd.DataFrame(stock_dict)
# -
# For the most part, the data is in a perfect format. However, we need to do one other transformation that involves transposing it. This will make selecting the columns of interest easier.
# +
# it's not in the right format, but if we transpose it we will have a much easier time grabbing the necessary columns.
stock_df = stock_df.transpose()
# display the head to make sure it's right
display(stock_df.head())
# let's recheck the shape, we lost a few hundred but this was expected as TD Ameritrade doesn't have data on every stock.
display(stock_df.shape)
# -
# ## Step Three: Select the Attributes
# ***
# We now have a master data frame that contains all the data on our stock. There are over 46 attributes we can choose from, or if we wanted, we could select them all. However, we will only be selecting three attributes in this tutorial. I want to make sure we understand certain concepts before we move on to higher dimensional data, and it’ll be easier to visualize these concepts with fewer attributes.
#
# **With that being said, I did have the luxury of testing multiple combinations of attributes, and one set that gave promising results was Return on Equity, Return on Assets, and Return on Investment.**
#
# If you’re curious what these metrics are, I provided some definitions found on **Investopedia**.
#
# 1. **Return on Assets:** Return on assets (ROA) is an indicator of how profitable a company is relative to its total assets. ROA gives a manager, investor, or analyst an idea as to how efficient a company’s management is at using its assets to generate earnings. Return on assets is displayed as a percentage.<br></br>
#
#
# 2. **Return on Equity:** Return on equity (ROE) is a measure of financial performance calculated by dividing net income by shareholders’ equity. Because shareholders’ equity is equal to a company’s assets minus its debt, ROE could be thought of as the return on net assets. ROE is considered a measure of how effectively management is using a company’s assets to create profits.
#
#
# 3. **Return on Investments:** Return on Investment (ROI) is a performance measure used to evaluate the efficiency of an investment or compare the efficiency of a number of different investments. ROI tries to directly measure the amount of return on a particular investment, relative to the investment’s cost. To calculate ROI, the benefit (or return) of an investment is divided by the cost of the investment. The result is expressed as a percentage or a ratio.
#
# When I wanted to select attributes, I wanted to select attributes that gave a different perspective of the company, and could be standardized across different types of companies.
#
# Also, make sure we choose the `symbol` column so we can use it as an index in our data frame. K-Means cannot handle missing data so it must be either removed from the data set or extra steps must be taken to fill in the missing values with either average values or estimates. To remove our missing data, we need to filter the data frame to remove any zeros, and then drop any `na` values by using the `dropna` method.
#
# This was honestly a very interesting part of the tutorial, so much so that I explored some research papers that helped influence my selection of attributes. If you get a chance, I highly encourage you to read some of these papers and see their findings.
#
# - https://www.researchgate.net/publication/4885243_Stock_selection_based_on_cluster_analysis
# - https://www.researchgate.net/publication/316705565_The_Classification_of_Stocks_with_Basic_Financial_Indicators_An_Application_of_Cluster_Analysis_on_the_BIST_100_Index
# +
# define our indicators list
metrics_list = ['returnOnEquity','returnOnAssets','returnOnInvestment','symbol']
# select only those columns
indicators_df = stock_df[metrics_list]
# display the unedited DF
display(indicators_df.head())
# clustering can't handle missing values, so we need to eliminate any row that has a missing value.
indicators_df = indicators_df[indicators_df[metrics_list] != 0]
indicators_df = indicators_df.dropna(how='any')
display(indicators_df.head())
# -
# Next, to make things more manageable, let's set the `symbol` column as the data frame index. This will make selecting certain stock symbols much easier.
# +
# set the index
indicators_df = indicators_df.set_index('symbol')
# do a data type conversion
indicators_df = indicators_df.astype('float')
indicators_df.head()
# -
# ## Step Four: Remove Outliers
# ***
# Now comes the exciting part, removing extreme values. The reality is that our data has many extreme values and this is expected given the variety of our companies. For example, our Return on Equity indicator has one company that has a value of 16000%! This value is not reflective of a traditional company, and more than likely is due to the fact of having a small equity base with large revenue denominator.
#
# We need to remove these extreme values from our data set or else they will skew our results. However, that begs the question of what is a "reasonable value"? If we do a little exploring on the internet, we find that NYU Stern has provided an industry benchmark for a wide range of financial metrics. This will serve as our baseline for removing extreme values from the data sets.
#
# If you're interested in seeing this benchmarks, please go to the following link:
# >*http://pages.stern.nyu.edu/~adamodar/New_Home_Page/datafile/pbvdata.html*
# >*http://pages.stern.nyu.edu/~adamodar/New_Home_Page/datafile/pedata.html*
#
#
# Let's get an idea of just how many "extreme" values exist in our data set by filtering the different columns and performing a `value_counts`.
# +
# define the filters
roe_filter = indicators_df.iloc[:,0] < 40
roa_filter = indicators_df.iloc[:,1] < 30
roi_filter = indicators_df.iloc[:,2] < 30
# get the counts for each column
roe_count = pd.Series(roe_filter).value_counts()
roa_count = pd.Series(roa_filter).value_counts()
roi_count = pd.Series(roi_filter).value_counts()
# display the results
display(roe_count)
display(roa_count)
display(roi_count)
# +
# filter the entire data frame
indicators_df = indicators_df[roe_filter & roa_filter & roi_filter]
# display the results
indicators_df.head()
# -
# Okay, after removing some of these extreme values, we are now at an excellent spot to do a statistical summary of our data set. Use the `describe` method to create a summary data frame and then add a standard deviation metric that is calculated using the mean.
# +
# create a statistical summary
desc_df = indicators_df.describe()
# add the standard deviation metric
desc_df.loc['+3_std'] = desc_df.loc['mean'] + (desc_df.loc['std'] * 3)
desc_df.loc['-3_std'] = desc_df.loc['mean'] - (desc_df.loc['std'] * 3)
# display it
desc_df
# -
# ## Step Five: Visualize & Scale the Data
# ***
# For the most part, the data is in a good range, there are a few values that are outside three standard deviations, but it's expected given the variety of stocks we have. Also, we will take steps later to mitigate this. Now that we've done a statistical summary, we can move on to plotting. In this example, let's use a 3D scatter plot to see all the data points. I will be using `matplotlib` to create our chart.
# +
# %matplotlib inline
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
# define a figure and a 3D axis
fig = plt.figure()
ax = Axes3D(fig)
# define the x, y, & z of our scatter plot, this will just be the data from our data frame.
x = list(indicators_df.iloc[:,0])
y = list(indicators_df.iloc[:,1])
z = list(indicators_df.iloc[:,2])
# define the axis labels
column_names = indicators_df.columns
ax.set_xlabel(column_names[0])
ax.set_ylabel(column_names[1])
ax.set_zlabel(column_names[2])
# define the markers, and the color
ax.scatter(x, y, z, c='royalBlue', marker='o')
plt.show()
# -
# As mentioned above, I purposely limited the number of attributes in our data set. The reason being that it's hard to visualize data that exceeds three dimensions. However, looking at this data it does seem to be clumped together with no definite spherical shape, this can cause issues when clustering because naturally, the algorithm looks for the precise structure to cluster the data around. There are steps we can take to help mitigate this issue, but this also a reality of data, there might not be clear clusters, and in some cases, clustering might not be the right approach.
#
# Let's continue and begin normalizing our data, to help handle outliers and things of that nature. You should almost always normalize your data as the model tends to perform better with normalized data vice the alternative. We have a few options to normalize our data:
#
# 1. **Standard Scaler:** Here we subtract the mean from each data point and divide it by the standard deviation. This method is sensitive to outliers; however, like computing, the mean means taking the average of ALL data points, including the outliers.
#
# 2. **Min Max Scaler:** Here we scale the data so that it fits in a range between 0 and 1. Mathematically we take each data point, subtract the minimum from it, and then divide it by the difference of the maximum value and the minimum value. Again, this is sensitive to outliers as the maximum amount would be the outlier.
#
# 3. **Robust Scaler:** This method is a better choice if your data has outliers. With this method, we use the interquartile range instead of the minimum and maximum, which helps control for outliers.
#
# In our example, because there might be a possibility for outliers, I recommend we use the Robust Scaling method.
# +
import numpy as np
from sklearn.preprocessing import Normalizer
from sklearn.preprocessing import MinMaxScaler, StandardScaler, RobustScaler
# for demonstration purposes, I will be creating all three instances of the scalers.
min_max_scaler = MinMaxScaler()
std_scaler = StandardScaler()
robust_scaler = RobustScaler()
# scale the data
X_train_minmax = min_max_scaler.fit_transform(indicators_df)
X_train_standard = std_scaler.fit_transform(indicators_df)
X_train_robust = robust_scaler.fit_transform(indicators_df)
# create a new plot
fig = plt.figure()
ax = Axes3D(fig)
# take the scaled data in this example.
x = X_train_robust[:,0]
y = X_train_robust[:,1]
z = X_train_robust[:,2]
# define the axes labels
column_names = indicators_df.columns
ax.set_xlabel(column_names[0])
ax.set_ylabel(column_names[1])
ax.set_zlabel(column_names[2])
# create a new plot
ax.scatter(x, y, z, c='royalBlue')
plt.show()
# -
# ## Step Six (Optional): Principal Component Analysis
# ***
# By scaling the data, we are allowing our algorithm to perform better with the data. Another topic we will discuss is the idea of Principal Component Analysis, which, with large data sets with multiple attributes, will allow our model to run faster and have less redundant data.
#
# To help conceptualize this concept, imagine I described a particular famous dog to you.
#
# > **Clifford is a big red dog with a friendly smile.**
#
# Now imagine, I added more detail to the story to help describe Clifford.
#
# > **Clifford is a big fire-red dog with a friendly smile.**
#
# For the most part, many people would say that adding the extra detail didn't provide much more to the story. I bet I could've left it out entirely and you still would've got the gist of the story. With PCA, we try to determine this additional info and remove it from the story. Yes, we might remove attributes from our data set, but it's to our benefit in most cases.
#
# The idea is that even though I might lose 3% of all information I still can describe 97% of the story and in most cases, this is more than enough to get the general theme of the story. We also gain the benefit of faster execution time when it comes to training our model. In a clustering algorithm like K-Means, while it is easy to understand and implement, it does suffer from long training runs when there are too many attributes.
#
# PCA attempts to remedy this problem by removing the attributes that don't contribute too much to story and keep those attributes which contribute significantly to the story. Implementing PCA in `sklearn` is easy to do, but something we need to do as an initial step is to get an understanding of the number of components we need.
#
# Visually this can be done by passing through our scaled data set into our PCA class object, and plotting the `explained_variance_ratio` which will tell us how much of the variance is explained at any given number of components.
#
# Let's implement this!
# +
from sklearn.decomposition import PCA
# pass through the scaled data set into our PCA class object
pca = PCA().fit(X_train_robust)
# plot the Cumulative Summation of the Explained Variance
plt.figure()
plt.plot(np.cumsum(pca.explained_variance_ratio_))
# define the labels & title
plt.xlabel('Number of Components', fontsize = 15)
plt.ylabel('Variance (%)', fontsize = 15)
plt.title('Explained Variance', fontsize = 20)
# show the plot
plt.show()
# -
# From the chart above, we can see that we have 100% of variance explained with only two components. This means that if we were to implement a PCA, we would select our number of components to be 2. In most examples, we won't get 100% explained the variance, but the general rule is to choose the minimum number of components that demonstrates the highest amount of variance.
#
# **I will only be using PCA for demonstration purposes, in a case with such few attributes as our example PCA is not justified or recommended.**
# +
# create a PCA modified dataset
pca_dataset = PCA(n_components=2).fit(X_train_robust).transform(X_train_robust)
# store it in a new data frame
pca_dataset= pd.DataFrame(data = pca_dataset, columns = ['principal component 1', 'principal component 2'])
# -
# By running PCA, we have reduced the number of dimensions in our data set from 3 to 2; this means if we graph our new data frame, we will only have two dimensions. Let's see how the new data set would look graphically.
# +
# define a figure
plt.figure()
# define the label and title
plt.xlabel('Principal Component 1', fontsize = 15)
plt.ylabel('Principal Component 2', fontsize = 15)
plt.title('2 Component PCA', fontsize = 20)
# plot the figure
plt.scatter(pca_dataset['principal component 1'], pca_dataset['principal component 2'], c='royalBlue', s = 50)
# -
# ## Step Seven: Build & Run the Model
# ***
# One of the biggest challenges with K-Means is determining the optimum number of clusters, so I decided to explore some options we have. One type of analysis we can do is related to silhouette analysis, and while I would like to describe it myself, I think the official `sklearn` documentation does a beautiful job of summarizing it.
#
# Silhouette analysis can be used to study the separation distance between the resulting clusters. The silhouette plot displays a measure of how close each point in one cluster is to points in the neighboring clusters and thus provides a way to assess parameters like the number of clusters visually. This measure has a range of (-1, 1).
#
# Silhouette coefficients (as these values are referred to as) near +1 indicate that the sample is far away from the neighboring clusters. A value of 0 indicates that the sample is on or very close to the decision boundary between two neighboring clusters and negative values indicate that those samples might have been assigned to the wrong cluster.
#
# If you would like a more detailed explanation, feel free to explore the documentation yourself.
# <br>*https://scikit-learn.org/stable/auto_examples/cluster/plot_kmeans_silhouette_analysis.html*</br>
#
# We will create an instance of our KMeans model, and explore how the silhouette score changes within a limited range of clusters. From here, we will take the maximum score to determine our cluster.
# +
from sklearn.cluster import KMeans
from sklearn import metrics
# define a dictionary that contains all of our relevant info.
results_dict = {}
# define how many clusters we want to test up to.
num_of_clusters = 10
# run through each instance of K
for k in range(2, num_of_clusters):
print("-"*100)
# define the next dictionary to hold all the results of this run.
results_dict[k] = {}
# create an instance of the model, and fit the training data to it.
kmeans = KMeans(n_clusters=k, random_state=0).fit(X_train_robust)
# define the silhouette score
sil_score = metrics.silhouette_score(X_train_robust, kmeans.labels_, metric='euclidean')
# store the different metrics
results_dict[k]['silhouette_score'] = sil_score
results_dict[k]['inertia'] = kmeans.inertia_
results_dict[k]['score'] = kmeans.score
results_dict[k]['model'] = kmeans
# print the results
print("Number of Clusters: {}".format(k))
print('Silhouette Score:', sil_score)
# -
# Okay, so we ran our cluster analysis now what? Well, the first thing we should look at is the overall silhouette score, **ideally the larger the number, the better the results. However, this score is just part of the story, and we will have to examine the results to give ourselves a sanity check visually, but at this point, we can use the silhouette score as a gage of which ones we should target first.**
#
# Looking at the results above, 2 or 3 should be our target.
#
# Let's also run the results on our PCA data set to see what the output would look like. In this case, we get a higher silhouette score, which is a good sign, and it points to the same outcome, explore a cluster of 2 or 3.
# +
# define a dictionary that contains all of our relevant info.
results_dict_pca = {}
# define how many clusters we want to test up to.
num_of_clusters = 10
# run through each instance of K
for k in range(2, num_of_clusters):
print("-"*100)
# define the next dictionary to hold all the results of this run.
results_dict_pca[k] = {}
# create an instance of the model, and fit the training data to it.
kmeans = KMeans(n_clusters=k, random_state=0).fit(pca_dataset)
# define the silhouette score
sil_score = metrics.silhouette_score(pca_dataset, kmeans.labels_, metric='euclidean')
# store the different metrics
results_dict_pca[k]['silhouette_score'] = sil_score
results_dict_pca[k]['inertia'] = kmeans.inertia_
results_dict_pca[k]['score'] = kmeans.score
results_dict_pca[k]['model'] = kmeans
# print the results
print("Number of Clusters: {}".format(k))
print('Silhouette Score:', sil_score)
# -
# ## Step Eight: Model Evaluation
# With the help of a graphical aid, we can also analyze the results of our clusters. If we use the `yellowbrick` library, we get to access the `SilhouetterVisualizer` which will help visualize the silhouette score for each point in that particular cluster. What we are looking for is that each cluster exceeds the red line or the average silhouette score and that the clusters are as evenly distributed as possible.
#
# You may be wondering why some of these values are below 0, and some are above, with the negative values these are data points that visually fall at the edge of two clusters. In other words, it's hard to determine where they fall; they are outliers that are tricky to group.
#
# The high positive values are data points that would almost be in the center of the cluster and very easy to classify. I've provided a visual aid to help drive home the concept.
# +
from yellowbrick.cluster import SilhouetteVisualizer
clusters = [2,3]
for cluster in clusters:
print('-'*100)
# define the model for K
kmeans = KMeans(n_clusters = cluster, random_state=0)
# pass the model through the visualizer
visualizer = SilhouetteVisualizer(kmeans)
# fit the data
visualizer.fit(X_train_robust)
# show the chart
visualizer.poof()
# +
clusters = [2,3]
for cluster in clusters:
print('-'*100)
# define the model for K
kmeans = KMeans(n_clusters = cluster, random_state=0)
# pass the model through the visualizer
visualizer = SilhouetteVisualizer(kmeans)
# fit the data
visualizer.fit(pca_dataset)
# show the chart
visualizer.poof()
# -
# ## Step Nine: Chart the Clusters
# ***
# Let's get to the fun part, visualizing our clusters! This will involve, in the non-PCA case, creating another 3D scatter plot, but in this instance, we will define the `c` parameter of our scatter to equal our model labels. This will correctly create the clusters for us.
#
# Again, we will do this in the case of 2 and 3 clusters so we can visually see the difference.
# +
clusters = [2,3]
for cluster in clusters:
print('-'*100)
kmeans = KMeans(n_clusters= cluster, random_state=0).fit(X_train_robust)
# define the cluster centers
cluster_centers = kmeans.cluster_centers_
C1 = cluster_centers[:, 0]
C2 = cluster_centers[:, 1]
C3 = cluster_centers[:, 2]
# create a new plot
fig = plt.figure()
ax = Axes3D(fig)
# take the scaled data in this example.
x = X_train_robust[:,0]
y = X_train_robust[:,1]
z = X_train_robust[:,2]
# define the axes labels
column_names = indicators_df.columns
ax.set_xlabel(column_names[0])
ax.set_ylabel(column_names[1])
ax.set_zlabel(column_names[2])
# create a new plot
ax.scatter(x, y, z, c = kmeans.labels_.astype(float), cmap='winter')
ax.scatter(C1, C2, C3, marker="x", color='r')
plt.title('Visualization of clustered data with {} clusters'.format(cluster), fontweight='bold')
plt.show()
# -
# How should we interpret these results? Personally, when I saw the three cluster one, it made sense. We have some companies who perform horribly across all three metrics, that would be the dark blue region in the lower left-hand corner. We then have our average company, denoted by the color green; these are companies who might beat on some metrics, lose on some, or do average. Finally, we have our "show stoppers" these are companies denoted in light blue and represent the companies who are crushing every metric.
#
# Again, there might be different ways to interpret these results, but for me it visually made sense, and while they might not have a nice spherical structure there is some type of grouping there albeit weak in some instances.
# +
clusters = [2,3]
for cluster in clusters:
print('-'*100)
kmeans = KMeans(n_clusters= cluster, random_state=0).fit(pca_dataset)
# define the cluster centers
cluster_centers = kmeans.cluster_centers_
C1 = cluster_centers[:, 0]
C2 = cluster_centers[:, 1]
# create a new plot
plt.figure()
# take the scaled data in this example.
x = pca_dataset['principal component 1']
y = pca_dataset['principal component 2']
# define the axes labels
column_names = pca_dataset.columns
plt.xlabel(column_names[0])
plt.ylabel(column_names[1])
# Visualize it:
plt.scatter(x, y, c=kmeans.labels_.astype(float), cmap='winter')
plt.scatter(C1, C2, marker="x", color='r')
# Plot the clustered data
plt.title('Visualization of clustered data with {} clusters'.format(cluster), fontweight='bold')
plt.show()
# -
# ## Step Ten: Make Predictions
# Okay, we've settled on three clusters being our number of choice let's make some predictions.
# +
# grab the model
kmeans = results_dict[3]['model']
# define some test data
test_data = [[0.10, 3.10, 5.20],
[0.60, 5.10, 2.20]]
# make some predictions, in this case let's pass through our test set.
kmeans.predict(test_data)
# -
# ## Final Notes:
#
# Overall, I was happy with the results, but this is not to say there couldn't be extra steps we could take to improve the results. This final part of the tutorial is taking a step back and looking at some things we might want to address or explore deeper.
#
# ***
# ### The Clusters Appear Very Dense
# This could impact the K-Means algorithm, and while it might not provide perfect results they could be suboptimal. Because of this, I would like to explore more density-based clustering algorithms and see how that impacts our results.
#
# ***
# ### Exploring How Filtering Affects the Results
# While I did my best to apply filters that seemed reasonable, I would be curious to see how our results change as we change the filter for outliers.
| python/python-data_science/machine-learning/k-means/Clustering Stocks - KMeans.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import src.helpers as helpers
# ### Average price per neighbourhood
# +
helpers.PDF('price_per_neighbourhood.pdf',size=(900,450))
# -
# ### Listings price per neighbourhood
helpers.PDF('listings_per_neighbourhood.pdf',size=(900,450))
# ## Price mean compared against number of listings per neighbourhood
helpers.PDF('listings_against_price.pdf',size=(1200,650))
| storytelling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Load packages
import warnings
warnings.filterwarnings('ignore')
import tsai
from tsai.all import *
print('tsai :', tsai.__version__)
print('fastai :', fastai.__version__)
print('fastcore :', fastcore.__version__)
print('torch :', torch.__version__)
torch.cuda.get_device_name(0)
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import matplotlib as mpl
from matplotlib import ticker
from datetime import datetime, timedelta
import statsmodels.api as sm
from statsmodels.graphics.tsaplots import plot_acf, plot_pacf
# +
large = 22; med = 16; small = 12
params = {'axes.titlesize': large,
'legend.fontsize': med,
'figure.figsize': (10, 6),
'axes.labelsize': med,
'axes.titlesize': med,
'xtick.labelsize': med,
'ytick.labelsize': med,
'figure.titlesize': large}
plt.rcParams.update(params)
plt.style.use('seaborn-whitegrid')
sns.set_style("white")
# %matplotlib inline
# Version
print(mpl.__version__)
print(sns.__version__)
# -
import hyperopt
print(hyperopt.__version__)
from hyperopt import Trials, STATUS_OK, STATUS_FAIL, tpe, fmin, hp
from hyperopt import space_eval
import time
from fastai.callback.tracker import EarlyStoppingCallback
import gc
import pickle
from math import sqrt
from sklearn.metrics import mean_squared_error, mean_absolute_error, mean_absolute_error
# # Specify parameters
# +
file_name = "./data/network-traffic-volume-2019-07-01.csv"
history = 15 # input historical time steps, here we use the same value with the maxlag we use for VAR
horizon = 1 # output predicted time steps
test_ratio = 0.2 # testing data ratio
# -
# # Load Data
df = pd.read_csv(file_name, sep=';')
start = datetime(1970, 1, 1) # Unix epoch start time
df['datetime'] = df.clock.apply(lambda x: start + timedelta(seconds=x))
df.head()
# Reverse the data order:
df = df.reindex(index=df.index[::-1])
df = df.reset_index(drop=True)
df.head()
# add a datetimeindex
dates = pd.DatetimeIndex(df['datetime'])
df = df.set_index(dates).asfreq('H') # one hour as the original sampling time frequency
df.head()
# # Split the data
train = df[df['datetime'] < '2019-06-16 00:00:00']
test = df[df['datetime'] > '2019-06-16 00:00:00']
train_length = train.shape[0]
test_length = test.shape[0]
print('Training size: ', train_length)
print('Test size: ', test_length)
print('Test ratio: ', test_length / (test_length + train_length))
# We plot the different time periods data:
# +
plt.figure(figsize=[12, 6])
plt.plot(df.index[:train_length], df['value_avg'][:train_length] / 1000 / 1000, label='Training', color='blue')
plt.plot(df.index[train_length:], df['value_avg'][train_length:] / 1000 / 1000, label='Test', color='red')
# Not supported for eps
plt.axvspan(df.index[train_length:][0], df.index[train_length:][-1], facecolor='r', alpha=0.1)
plt.xlabel('Time')
plt.ylabel('Traffic (Mbps)')
plt.legend(loc='best')
# plt.show()
plt.savefig('fig1.pdf', bbox_inches = 'tight', pad_inches = 0.1)
# -
# # Plot data distribution
from matplotlib.ticker import PercentFormatter
plt.figure(figsize=[12, 6])
plt.hist(df['value_avg'] / 1000 / 1000, bins=100, density=True)
plt.gca().yaxis.set_major_formatter(PercentFormatter(1))
plt.xlabel('Traffic (Mbps)')
plt.ylabel('Data Ratio')
# plt.show()
plt.savefig('fig2.pdf', bbox_inches = 'tight', pad_inches = 0.1)
# # Plot predicted results
# Get the true values on the test set first:
compare_test = test[['value_avg']].copy()
compare_test.columns = ['True Values']
# Load the results of VAR:
model_name = 'VAR'
history = 24
horizon = 1
y_pred_fn = '%s_pred-%d-%d.pkl' % (model_name, history, horizon)
var_pred = pickle.load(open(y_pred_fn, 'rb'))
compare_test['VAR'] = var_pred
compare_test.head()
# Define a function to load the results of a single deep learning model:
def load_predicted_results(model_name, history=24, horizon=1, step_to_evalute=0):
y_pred_fn = '%s_pred-%d-%d.pkl' % (model_name, history, horizon)
y_pred = pickle.load(open(y_pred_fn, 'rb'))
return y_pred[:, step_to_evalute]
# Load the results of InceptionTime for comparison:
compare_test['InceptionTime'] = load_predicted_results('InceptionTime')
compare_test.head()
# Plot the results for a short time period:
df = compare_test.head(24 * 7) # the first hours in the test set
plt.figure(figsize=[12, 6])
plt.plot(df.index, df['True Values'] / 1000 / 1000, 'b', label='True Values')
plt.plot(df.index, df['VAR'] / 1000 / 1000, 'y', label='VAR')
plt.plot(df.index, df['InceptionTime'] / 1000 / 1000, 'r', label='InceptionTime')
plt.xlabel('Time')
plt.ylabel('Traffic (Mbps)')
plt.legend()
# plt.show()
plt.savefig('fig3.pdf', bbox_inches = 'tight', pad_inches = 0.1)
| UsagePrediction/Plot_paper_figures.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# In order to successfully complete this assignment you need to participate both individually and in groups during class on **Monday March 11th**.
#
#
# # In-Class Assignment: CUDA Memory and Tiling
#
# <img src="https://www.appianimosaic.com/uploads/2016-6-24/1920-300/per_downolad_2n.jpg">
# <p style="text-align: right;">Image from: https://www.appianimosaic.com/</p>
# </p>
#
# ### Agenda for today's class (70 minutes)
#
# 1. (20 minutes) HW4 Review
# 2. (10 minutes) Pre-class Review
# 1. (10 minutes) Jupyterhub test
# 3. (30 minutes) Tile Example
# 4. (0 minutes) 2D wave Cuda Code Optimization
# ----
#
# # 1. HW4 Review
#
#
# [0301-HW4-Image_processing](0301-HW4-Image_processing.ipynb)
#
# ---
# # 2. Pre-class Review
#
# [0310--CUDA_Memory-pre-class-assignment](0310--CUDA_Memory-pre-class-assignment.ipynb)
# ---
#
# # 3. Jupyterhub Test
#
# As a class lets try to access the GPU jupyterhub server:
#
# https://jupyterhub-gpu.egr.msu.edu
#
# Upload this file to your server account and lets run class from there. Note any odd behaviors to the instructor.
#
#
#
# ----
#
# # 3. Tile example
#
#
# +
# %%writefile tiled_transpose.cu
#include <iostream>
#include <cuda.h>
#include <chrono>
#define CUDA_CALL(x) {cudaError_t cuda_error__ = (x); if (cuda_error__) { fprintf(stderr, "CUDA error: " #x " returned \"%s\"\n", cudaGetErrorString(cuda_error__)); fflush(stderr); exit(cuda_error__); } }
using namespace std;
const int BLOCKDIM = 32;
__global__ void transpose(const double *in_d, double * out_d, int row, int col)
{
int x = blockIdx.x * blockDim.x + threadIdx.x;
int y = blockIdx.y * blockDim.y + threadIdx.y;
if (x < col && y < row)
out_d[y+col*x] = in_d[x+row*y];
}
__global__ void tiled_transpose(const double *in_d, double * out_d, int row, int col)
{
int x = blockIdx.x * BLOCKDIM + threadIdx.x;
int y = blockIdx.y * BLOCKDIM + threadIdx.y;
int x2 = blockIdx.y * BLOCKDIM + threadIdx.x;
int y2 = blockIdx.x * BLOCKDIM + threadIdx.y;
__shared__ double in_local[BLOCKDIM][BLOCKDIM];
__shared__ double out_local[BLOCKDIM][BLOCKDIM];
if (x < col && y < row) {
in_local[threadIdx.x][threadIdx.y] = in_d[x+row*y];
__syncthreads();
out_local[threadIdx.y][threadIdx.x] = in_local[threadIdx.x][threadIdx.y];
__syncthreads();
out_d[x2+col*y2] = out_local[threadIdx.x][threadIdx.y];
}
}
__global__ void transpose_symmetric(double *in_d, double * out_d, int row, int col)
{
int x = blockIdx.x * blockDim.x + threadIdx.x;
int y = blockIdx.y * blockDim.y + threadIdx.y;
if (x < col && y < row) {
if (x < y) {
double temp = in_d[y+col*x];
in_d[y+col*x] = in_d[x+row*y];
in_d[x+row*y] = temp;
}
}
}
int main(int argc,char **argv)
{
std::cout << "Begin\n";
int sz_x=BLOCKDIM*300;
int sz_y=BLOCKDIM*300;
int nBytes = sz_x*sz_y*sizeof(double);
int block_size = BLOCKDIM;
double *m_h = (double *)malloc(nBytes);
double * in_d;
double * out_d;
int count = 0;
for (int i=0; i < sz_x*sz_y; i++){
m_h[i] = count;
count++;
}
std::cout << "Allocating device memory on host..\n";
CUDA_CALL(cudaMalloc((void **)&in_d,nBytes));
CUDA_CALL(cudaMalloc((void **)&out_d,nBytes));
//Set up blocks
dim3 dimBlock(block_size,block_size,1);
dim3 dimGrid(sz_x/block_size,sz_y/block_size,1);
std::cout << "Doing GPU Transpose\n";
CUDA_CALL(cudaMemcpy(in_d,m_h,nBytes,cudaMemcpyHostToDevice));
auto start_d = std::chrono::high_resolution_clock::now();
/**********************/
transpose<<<dimGrid,dimBlock>>>(in_d,out_d,sz_y,sz_x);
//tiled_transpose<<<dimGrid,dimBlock>>>(in_d,out_d,sz_y,sz_x);
cudaError_t err = cudaGetLastError();
if (err != cudaSuccess) {
fprintf(stderr, "\n\nError: %s\n\n", cudaGetErrorString(err)); fflush(stderr); exit(err);
}
CUDA_CALL(cudaMemcpy(m_h,out_d,nBytes,cudaMemcpyDeviceToHost));
/************************/
/**********************
transpose_symmetric<<<dimGrid,dimBlock>>>(in_d,out_d,sz_y,sz_x);
cudaError_t err = cudaGetLastError();
if (err != cudaSuccess) {
fprintf(stderr, "\n\nError: %s\n\n", cudaGetErrorString(err)); fflush(stderr); exit(err);
}
CUDA_CALL(cudaMemcpy(m_h,in_d,nBytes,cudaMemcpyDeviceToHost));
************************/
auto end_d = std::chrono::high_resolution_clock::now();
std::cout << "Doing CPU Transpose\n";
auto start_h = std::chrono::high_resolution_clock::now();
for (int y=0; y < sz_y; y++){
for (int x=y; x < sz_x; x++){
double temp = m_h[x+sz_x*y];
//std::cout << temp << " ";
m_h[x+sz_x*y] = m_h[y+sz_y*x];
m_h[y+sz_y*x] = temp;
}
//std::cout << "\n";
}
auto end_h = std::chrono::high_resolution_clock::now();
//Checking errors (should be same values as start)
count = 0;
int errors = 0;
for (int i=0; i < sz_x*sz_y; i++){
if (m_h[i] != count)
errors++;
count++;
}
std::cout << errors << " Errors found in transpose\n";
//Print Timing
std::chrono::duration<double> time_d = end_d - start_d;
std::cout << "Device time: " << time_d.count() << " s\n";
std::chrono::duration<double> time_h = end_h - start_h;
std::cout << "Host time: " << time_h.count() << " s\n";
cudaFree(in_d);
cudaFree(out_d);
return 0;
}
# -
#Compile Cuda
# !nvcc -std=c++11 -o tiled_transpose tiled_transpose.cu
#Run Example
# !./tiled_transpose
# ----
#
# # 4. 1D wave Cuda Code Optimization
#
# As a group, lets see if we can optimize the code code from lasttime.
# +
# %%writefile wave_cuda.cu
#include <stdio.h>
#include <stdlib.h>
#include <math.h>
#include <cuda.h>
#define CUDA_CALL(x) {cudaError_t cuda_error__ = (x); if (cuda_error__) printf("CUDA error: " #x " returned \"%s\"\n", cudaGetErrorString(cuda_error__));}
__global__ void accel_update(double* d_dvdt, double* d_y, int nx, double dx2inv)
{
int i = blockDim.x * blockIdx.x + threadIdx.x;
if (i > 0 && i < nx-1)
d_dvdt[i]=(d_y[i+1]+d_y[i-1]-2.0*d_y[i])*(dx2inv);
else
d_dvdt[i] = 0;
}
__global__ void pos_update(double * d_dvdt, double * d_y, double * d_v, double dt)
{
int i = blockDim.x * blockIdx.x + threadIdx.x;
d_v[i] = d_v[i] + dt*d_dvdt[i];
d_y[i] = d_y[i] + dt*d_v[i];
}
int main(int argc, char ** argv) {
int nx = 5000;
int nt = 1000000;
int i,it;
double x[nx];
double y[nx];
double v[nx];
double dvdt[nx];
double dt;
double dx;
double max,min;
double dx2inv;
double tmax;
double *d_x, *d_y, *d_v, *d_dvdt;
CUDA_CALL(cudaMalloc((void **)&d_x,nx*sizeof(double)));
CUDA_CALL(cudaMalloc((void **)&d_y,nx*sizeof(double)));
CUDA_CALL(cudaMalloc((void **)&d_v,nx*sizeof(double)));
CUDA_CALL(cudaMalloc((void **)&d_dvdt,nx*sizeof(double)));
max=10.0;
min=0.0;
dx = (max-min)/(double)(nx-1);
x[0] = min;
for(i=1;i<nx-1;i++) {
x[i] = min+(double)i*dx;
}
x[nx-1] = max;
tmax=10.0;
dt= (tmax-0.0)/(double)(nt-1);
for (i=0;i<nx;i++) {
y[i] = exp(-(x[i]-5.0)*(x[i]-5.0));
v[i] = 0.0;
dvdt[i] = 0.0;
}
CUDA_CALL(cudaMemcpy(d_x,x,nx*sizeof(double),cudaMemcpyHostToDevice));
CUDA_CALL(cudaMemcpy(d_y,y,nx*sizeof(double),cudaMemcpyHostToDevice));
CUDA_CALL(cudaMemcpy(d_v,v,nx*sizeof(double),cudaMemcpyHostToDevice));
CUDA_CALL(cudaMemcpy(d_dvdt,dvdt,nx*sizeof(double),cudaMemcpyHostToDevice));
dx2inv=1.0/(dx*dx);
int block_size=1024;
int block_no = nx/block_size;
dim3 dimBlock(block_size,1,1);
dim3 dimGrid(block_no,1,1);
for(it=0;it<nt-1;it++) {
accel_update<<<dimGrid, dimBlock>>>(d_dvdt, d_y, nx, dx2inv);
pos_update<<<dimGrid, dimBlock>>>(d_dvdt, d_y, d_v, dt);
}
CUDA_CALL(cudaMemcpy(x,d_x,nx*sizeof(double),cudaMemcpyDeviceToHost));
CUDA_CALL(cudaMemcpy(y,d_y,nx*sizeof(double),cudaMemcpyDeviceToHost));
for(i=nx/2-10; i<nx/2+10; i++) {
printf("%g %g\n",x[i],y[i]);
}
return 0;
}
# -
# !nvcc -std=c++11 -o wave_cuda wave_cuda.cu
# %%time
# !./wave_cuda
# -----
# ### Congratulations, we're done!
#
# **Course Resources:**
# - [Syllabus](https://tinyurl.com/y75cnzam)
# - [Preliminary Schedule](https://tinyurl.com/CMSE314-Schedule)
# - [Git Repository](https://gitlab.msu.edu/colbrydi/cmse401-s19)
# - [Jargon Jar and Command History](https://tinyurl.com/CMSE314-JargonJar)
#
#
# © Copyright 2019, Michigan State University Board of Trustees
| 0311_CUDA_in-class-assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # Notebook Template
#
# This notebook is a template notebook that simply has a preamble with a few useful imports. Add or delete lines as you need to set up the notebook for your own project!
# ## Preamble
# %load_ext autoreload
# %autoreload 2
# install im_tutorial package
# # !pip install git+https://github.com/nestauk/im_tutorials.git
# +
# useful Python tools
import itertools
import collections
# matplotlib for static plots
import matplotlib.pyplot as plt
# networkx for networks
import networkx as nx
# numpy for mathematical functions
import numpy as np
# pandas for handling tabular data
import pandas as pd
# seaborn for pretty statistical plots
import seaborn as sns
pd.set_option('max_columns', 99)
# basic bokeh imports for an interactive scatter plot or line chart
from bokeh.io import show, output_notebook
from bokeh.plotting import figure
from bokeh.models import ColumnDataSource, Circle, Line
# NB: If using Google Colab, this function must be run at
# the end of any cell that you want to display a bokeh plot.
# If using Jupyter, then this line need only appear once at
# the start of the notebook.
output_notebook()
# -
# ## Import Data
# The im_tutorials datasets module can be used to easily load datasets.
from im_tutorials.data import *
# For example, to load Gateway to Research projects:
gtr_projects_df = gtr.gateway_to_research_projects()
| notebooks/00_project_template.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Notebooks pour le cours chimie théorique 2A DENS du département de chimie de l'ENS
#
# Ceci est une collection de notebooks IPython pour illustrer le cours de chimie théorique. La liste des notebooks disponibles est:
#
# * [Théorème de Bloch](notebooks/BlochTheorem.ipynb)
| index.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Using Source and Sink Terms for a Chemical Reaction
# `OpenPNM` is capable of simulating chemical reactions in pores by adding source and sink terms. This example shows how to add source and sink terms to a steady state fickian diffusion simulation.
import warnings
import scipy as sp
import numpy as np
import openpnm as op
import matplotlib.pyplot as plt
np.set_printoptions(precision=5)
np.random.seed(10)
# %matplotlib inline
# ## Create Network and Other Objects
# Start by creating the network, geometry, phase and physics objects as usual:
pn = op.network.Cubic(shape=[40, 40], spacing=1e-4)
geo = op.geometry.StickAndBall(network=pn, pores=pn.Ps, throats=pn.Ts)
gas = op.phases.Air(network=pn)
phys = op.physics.Standard(network=pn, phase=gas, geometry=geo)
# Now add the source and sink models to the physics object. In this case we'll think of the as chemical reactions. We'll add one source term and one sink term, meaning one negative reaction rate and one positive reaction rate
gas['pore.concentration'] = 0
phys['pore.sinkA'] = -1e-10
phys['pore.sinkb'] = 1
phys.add_model(propname='pore.sink', model=op.models.physics.generic_source_term.power_law,
A1='pore.sinkA', A2='pore.sinkb', X='pore.concentration')
phys['pore.srcA'] = 1e-11
phys['pore.srcb'] = 1
phys.add_model(propname='pore.source', model=op.models.physics.generic_source_term.power_law,
A1='pore.srcA', A2='pore.srcb', X='pore.concentration')
# ## Setup Fickian Diffusion with Sink Terms
# Now we setup a FickianDiffusion algorithm, with concentration boundary conditions on two side, and apply the sink term to 3 pores:
rx = op.algorithms.FickianDiffusion(network=pn)
rx.setup(phase=gas)
rx.set_source(propname='pore.sink', pores=[420, 820, 1220])
rx.set_value_BC(values=1, pores=pn.pores('front'))
rx.set_value_BC(values=1, pores=pn.pores('back'))
rx.run()
# ## Plot Distributions
#
# We can use the ``plot_connections`` and ``plot_coordinates`` to get a quick view of the pore network, with colors and sizes scaled appropriately.
pn['pore.concentration'] = rx['pore.concentration']
fig = plt.figure(figsize=[15, 15])
fig = op.topotools.plot_connections(network=pn, color='k', alpha=.2, fig=fig)
fig = op.topotools.plot_coordinates(network=pn, color_by=pn['pore.concentration'],
size_by=pn['pore.diameter'], cmap='plasma',
fig=fig, markersize=100)
_ = plt.axis('off')
# ## Plot Distributions as Heatmaps
# Because the network is a 2D cubic, it is convenient to visualize it as an image, so we reshape the 'pore.concentration' array that is produced by the FickianDiffusion algorithm upon running, and turn it into a colormap representing concentration in each pore.
pn['pore.concentration'] = rx['pore.concentration']
fig = plt.figure(figsize=[15, 15])
fig = op.topotools.plot_connections(network=pn, color='k', alpha=0, fig=fig)
fig = op.topotools.plot_coordinates(network=pn, color_by=pn['pore.concentration'], cmap='plasma',
fig=fig, markersize=420, marker='s')
_ = plt.axis('off')
# ## Setup Fickian Diffusion with Source Terms
# Similarly, for the source term:
rx = op.algorithms.FickianDiffusion(network=pn)
rx.setup(phase=gas)
rx.set_source(propname='pore.source', pores=[420, 820, 1220])
rx.set_value_BC(values=1, pores=pn.pores('front'))
rx.set_value_BC(values=1, pores=pn.pores('back'))
rx.run()
# And plotting the result as a color map:
#NBVAL_IGNORE_OUTPUT
pn['pore.concentration'] = rx['pore.concentration'] - 1
fig = plt.figure(figsize=[15, 15])
fig = op.topotools.plot_connections(network=pn, color='k', alpha=0, fig=fig)
fig = op.topotools.plot_coordinates(network=pn, color_by=pn['pore.concentration'], cmap='plasma',
fig=fig, markersize=420, marker='s')
_ = plt.axis('off')
| examples/notebooks/algorithms/reactive/diffusion_with_source_and_sink_terms.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Generate RDF Knowledge graph from spreadsheet
#
# 1. Original input file on Google Docs: https://docs.google.com/spreadsheets/d/11SElScyLIs0RANYcT2MpWa7QWf-gIB13fjNvV9b5ZBE/export?format=xlsx&id=11SElScyLIs0RANYcT2MpWa7QWf-gIB13fjNvV9b5ZBE
#
# 2. We use a GitHub Actions workflow to generate the NeuroDKG: https://github.com/MaastrichtU-IDS/neuro_dkg/blob/master/.github/workflows/generate-rdf.yml
#
# # Use this notebook
#
# To run this notebook, start a `jupyter/all-spark-notebook` container using `docker-compose up` from the `docs/` folder
#
# ```
# # cd docs
# docker-compose up
# ```
#
# Access on http://localhost:8888
# !pip install -r requirements.txt
# # Trying out the official pyRDF2Vec
#
# Documentation: https://pyrdf2vec.readthedocs.io/en/latest/readme.html#create-a-knowledge-graph-object
# +
from pyrdf2vec.graphs import KG
label_predicates = ["http://www.w3.org/1999/02/22-rdf-syntax-ns#type"]
kg = KG(location="neurodkg.ttl", label_predicates=label_predicates)
# +
from pyrdf2vec.samplers import UniformSampler
from pyrdf2vec.walkers import RandomWalker
from pyrdf2vec import RDF2VecTransformer
walkers = [RandomWalker(4, 5, UniformSampler())]
# Get entities from the KG
entities = set([])
for v in kg._entities:
if v.name.startswith('http://www.w3id.org/neurodkg/Instances'):
entities.add(v.name)
entities = list(entities)
print(entities)
transformer = RDF2VecTransformer(walkers=[walkers])
# "Entities should be a list of URIs that can be found in the Knowledge Graph"
embeddings = transformer.fit_transform(kg, entities)
print(embeddings)
# -
# # Using Remzi's pyRDF2Vec
#
# At https://github.com/MaastrichtU-IDS/pyRDF2Vec/
# +
import random
import os
import requests
import functools
import numpy as np
import rdflib
import pandas as pd
import matplotlib.pyplot as plt
import shutil
from sklearn.svm import SVC
from sklearn.metrics import confusion_matrix, accuracy_score
from sklearn.manifold import TSNE
from rdf2vec.converters import rdflib_to_kg
from rdf2vec.walkers import RandomWalker
from rdf2vec import RDF2VecTransformer
import warnings
warnings.filterwarnings('ignore')
# -
# ## Import the rdf file (ttl, nt, all other supported by rdflib)
# +
url = 'https://raw.githubusercontent.com/MaastrichtU-IDS/neuro_dkg/master/data/output/neuro_dkg.ttl'
rdf_file ='neurodkg.ttl'
# rdf_file = url.split('/')[-1]
#rdf_file = 'input/covid19-literature-knowledge-graph/sample_kg.nt'
#fileext = '.nq.gz'
# Download the RDF file
with requests.get(url, stream=True) as r:
with open(rdf_file, 'wb') as f:
r.raw.read = functools.partial(r.raw.read, decode_content=True)
shutil.copyfileobj(r.raw, f)
#predicates for Random Walker to follow
label_predicates = ['<http://www.w3.org/1999/02/22-rdf-syntax-ns#type>']
# -
kg = rdflib_to_kg(rdf_file, filetype='turtle')
# +
# We'll all possible walks of depth 2
random_walker = RandomWalker(2, 4)
# Create embeddings with random walks
transformer = RDF2VecTransformer(walkers=[random_walker], sg=1)
# -
all_entities = kg.get_all_entities()
all_entities[:10]
walk_embeddings = transformer.fit_transform(kg, all_entities)
walk_embeddings[:10]
len(all_entities)
len(walk_embeddings)
# ## Generating a dataframe for entity embeddings
df =pd.DataFrame(zip(all_entities, walk_embeddings), columns=['entity', 'embedding'])
# +
# a function for converting entity names
# if you need to provide entity names with CURIE format (e.g. DRUGBANK:DB00012)
def replace_prefix(entity):
if entity.startswith('http://www.w3id.org/drugbank:'):
return entity.replace('http://www.w3id.org/drugbank:', 'DRUGBANK:')
else:
return entity
df.entity = df.entity.apply(replace_prefix)
# -
# ### Convert dataframe embeddings to JSON
#
# And store the dataframe in a JSON file, to be imported in the OpenPredict API!
df.to_json('neurodkg_embedding.json',orient='records')
import pandas as pd
import numpy as np
df =pd.read_json('neurodkg_embedding.json',orient='records')
df.head()
print(df['entity'])
# np.array(df['embedding'].values)
embedding_mat =[]
for i, row in df.iterrows():
emb=row['embedding']
embedding_mat.append(emb)
entities = df.entity.to_list()
# ### alternatively you can store as csv with X columns (X is the dimension of the emebdding)
df_emb =pd.DataFrame( embedding_mat, columns= ['feature'+str(i) for i in range(len(emb))])
df_emb['entity'] = entities
df_emb.to_csv('neurodkg_embedding.csv', index=False)
df_emb.head()
| docs/openpredict-pyrdf2vec-embeddings.ipynb |