code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: cpu_env
# kernelspec:
# display_name: 'Python 3.8.11 64-bit (''rinna_gpt2_predict'': conda)'
# name: python3
# ---
# # GPT-2 モデルの ONNX 変換と量子化
# ## 準備
#
# Python 3.8 カーネルが存在することが前提
#
# ```console
# conda create -n rinna_gpt2_predict python=3.8
# conda activate rinna_gpt2_predict
# conda install jupyter
# jupyter notebook
# ```
# + gather={"logged": 1628068194057}
# ライブラリインストール
import sys
if sys.platform == 'darwin': # Mac
# !{sys.executable} -m pip install --upgrade torch torchvision
else:
# !{sys.executable} -m pip install --upgrade torch==1.9.0+cpu torchvision==0.10.0+cpu torchaudio==0.9.0 -f https://download.pytorch.org/whl/torch_stable.html
# !{sys.executable} -m pip install onnxruntime==1.8.1
# !{sys.executable} -m pip install sentencepiece
# !{sys.executable} -m pip install transformers==4.8.2
# !{sys.executable} -m pip install onnx onnxconverter_common psutil pytz pandas py-cpuinfo py3nvml sympy coloredlogs azureml-core azureml-mlflow mlflow
# + gather={"logged": 1628068194169}
# キャッシュ保存用のディレクトリを用意
import os
cache_dir = os.path.join(".", "cache_models")
if not os.path.exists(cache_dir):
os.makedirs(cache_dir)
# +
# Azure ML Workspace への接続
from azureml.core import Experiment, Workspace, Environment
from azureml.core.compute import ComputeTarget
from azureml.core import ScriptRunConfig
from azureml.core.runconfig import PyTorchConfiguration
from azureml.core.authentication import InteractiveLoginAuthentication
import mlflow
interactive_auth = InteractiveLoginAuthentication(force=True,tenant_id="72f988bf-86f1-41af-91ab-2d7cd011db47")
ws = Workspace.from_config(path='config.json',auth=interactive_auth)
# AML 上で実行する場合は上記2行コメントアウト、以下実行
# ws = Workspace.from_config()
mlflow.set_tracking_uri(ws.get_mlflow_tracking_uri())
# -
# ## モデル読み込み
client = mlflow.tracking.MlflowClient()
registered_model = client.get_model_version(name='test-model',version=37)
client.download_artifacts(registered_model.run_id, 'outputs/models', cache_dir)
# + gather={"logged": 1628068248507}
# GPT-2 モデルにビームサーチを組み合わせるヘルパー class で読み込んだ GPT-2 モデルをラップ
from onnxruntime.transformers.gpt2_beamsearch_helper import Gpt2BeamSearchHelper, GPT2LMHeadModel_BeamSearchStep
from transformers import AutoConfig
import torch
model_name_or_path = os.path.join(cache_dir, 'outputs/models')
config = AutoConfig.from_pretrained(model_name_or_path, cache_dir=model_name_or_path)
model = GPT2LMHeadModel_BeamSearchStep.from_pretrained(model_name_or_path, config=config, batch_size=1, beam_size=4, cache_dir=cache_dir)
device = torch.device("cpu")
model.eval().to(device)
# -
# 推論で使う関数用にモデルの情報を取得
num_attention_heads = model.config.n_head
hidden_size = model.config.n_embd
num_layer = model.config.n_layer
# tokenizer を読み込み
from transformers import T5Tokenizer
tokenizer = T5Tokenizer.from_pretrained("rinna/japanese-gpt2-medium", cache_dir=cache_dir)
tokenizer.padding_side = "left"
tokenizer.pad_token = tokenizer.eos_token
tokenizer.do_lower_case = True
# ## PyTorch GPT-2 モデルをビームサーチの1ステップを含む ONNX に変換
# + gather={"logged": 1628068248683}
# ONNX に変換
onnx_model_path = os.path.join(cache_dir, "rinna_gpt2_beam_step_search.onnx")
if not os.path.exists(onnx_model_path):
Gpt2BeamSearchHelper.export_onnx(model, device, onnx_model_path) # add parameter use_external_data_format=True when model size > 2 GB
else:
print("GPT-2 ONNX model exists.")
# +
# 最適化と量子化
from onnxruntime.transformers.gpt2_helper import Gpt2Helper, MyGPT2LMHeadModel
from onnxruntime.transformers.quantize_helper import QuantizeHelper
optimized_model_path = os.path.join(cache_dir, "rinna_gpt2_beam_step_search_optimized.onnx")
quantized_model_path = os.path.join(cache_dir, "rinna_gpt2_beam_step_search_optimized_int8.onnx")
if not os.path.exists(optimized_model_path):
Gpt2Helper.optimize_onnx(onnx_model_path, optimized_model_path, False, model.config.num_attention_heads, model.config.hidden_size)
else:
print("Optimized GPT-2 ONNX model exists.")
if not os.path.exists(quantized_model_path):
QuantizeHelper.quantize_onnx_model(optimized_model_path, quantized_model_path)
else:
print("Quantized GPT-2 Int8 ONNX model exists.")
# -
# 量子化した ONNX をモデルとして登録
mlflow.set_experiment('register_onnx')
with mlflow.start_run() as run:
remote_model_path = os.path.join('outputs','onnx', "rinna_gpt2_beam_step_search_optimized_int8.onnx")
mlflow.log_artifact(quantized_model_path, remote_model_path)
model_uri = "runs:/{}/".format(run.info.run_id) + remote_model_path
mlflow.register_model(model_uri, 'rinna-GPT2-quantized-model')
# ## 推論テスト
# + gather={"logged": 1628068299895}
import onnxruntime
import numpy
from transformers import T5Tokenizer
EXAMPLE_Text = ['私はりんなです。']
def get_tokenizer(model_name_or_path, cache_dir):
tokenizer = T5Tokenizer.from_pretrained(model_name_or_path, cache_dir=cache_dir)
tokenizer.padding_side = "left"
tokenizer.pad_token = tokenizer.eos_token
tokenizer.do_lower_case = True
#okenizer.add_special_tokens({'pad_token': '[PAD]'})
return tokenizer
def get_example_inputs(prompt_text=EXAMPLE_Text):
tokenizer = get_tokenizer('rinna/japanese-gpt2-medium', cache_dir)
encodings_dict = tokenizer.batch_encode_plus(prompt_text, padding=True)
input_ids = torch.tensor(encodings_dict['input_ids'], dtype=torch.int64)
attention_mask = torch.tensor(encodings_dict['attention_mask'], dtype=torch.float32)
position_ids = (attention_mask.long().cumsum(-1) - 1)
position_ids.masked_fill_(position_ids < 0, 0)
#Empty Past State for generating first word
empty_past = []
batch_size = input_ids.size(0)
sequence_length = input_ids.size(1)
past_shape = [2, batch_size, num_attention_heads, 0, hidden_size // num_attention_heads]
for i in range(num_layer):
empty_past.append(torch.empty(past_shape).type(torch.float32).to(device))
return input_ids, attention_mask, position_ids, empty_past
input_ids, attention_mask, position_ids, empty_past = get_example_inputs()
beam_select_idx = torch.zeros([1, input_ids.shape[0]]).long()
input_log_probs = torch.zeros([input_ids.shape[0], 1])
input_unfinished_sents = torch.ones([input_ids.shape[0], 1], dtype=torch.bool)
prev_step_scores = torch.zeros([input_ids.shape[0], 1])
session = onnxruntime.InferenceSession(onnx_model_path)
ort_inputs = {
'input_ids': numpy.ascontiguousarray(input_ids.cpu().numpy()),
'attention_mask' : numpy.ascontiguousarray(attention_mask.cpu().numpy()),
'position_ids': numpy.ascontiguousarray(position_ids.cpu().numpy()),
'beam_select_idx': numpy.ascontiguousarray(beam_select_idx.cpu().numpy()),
'input_log_probs': numpy.ascontiguousarray(input_log_probs.cpu().numpy()),
'input_unfinished_sents': numpy.ascontiguousarray(input_unfinished_sents.cpu().numpy()),
'prev_step_results': numpy.ascontiguousarray(input_ids.cpu().numpy()),
'prev_step_scores': numpy.ascontiguousarray(prev_step_scores.cpu().numpy()),
}
for i, past_i in enumerate(empty_past):
ort_inputs[f'past_{i}'] = numpy.ascontiguousarray(past_i.cpu().numpy())
#print(ort_inputs)
ort_outputs = session.run(None, ort_inputs)
#print(ort_outputs)
# -
# ### ONNX Runtime Inference with IO Binding
#
# GPU を使用する場合の推論パフォーマンス改善
# + gather={"logged": 1628068300137}
def inference_with_io_binding(session, config, input_ids, position_ids, attention_mask, past, beam_select_idx, input_log_probs, input_unfinished_sents, prev_step_results, prev_step_scores, step, context_len):
output_shapes = Gpt2BeamSearchHelper.get_output_shapes(batch_size=1,
context_len=context_len,
past_sequence_length=past[0].size(3),
sequence_length=input_ids.size(1),
beam_size=4,
step=step,
config=config,
model_class="GPT2LMHeadModel_BeamSearchStep")
output_buffers = Gpt2BeamSearchHelper.get_output_buffers(output_shapes, device)
io_binding = Gpt2BeamSearchHelper.prepare_io_binding(session, input_ids, position_ids, attention_mask, past, output_buffers, output_shapes, beam_select_idx, input_log_probs, input_unfinished_sents, prev_step_results, prev_step_scores)
session.run_with_iobinding(io_binding)
outputs = Gpt2BeamSearchHelper.get_outputs_from_io_binding_buffer(session, output_buffers, output_shapes, return_numpy=False)
return outputs
# + gather={"logged": 1628068303048}
input_ids, attention_mask, position_ids, empty_past = get_example_inputs()
beam_select_idx = torch.zeros([1, input_ids.shape[0]]).long()
input_log_probs = torch.zeros([input_ids.shape[0], 1])
input_unfinished_sents = torch.ones([input_ids.shape[0], 1], dtype=torch.bool)
prev_step_scores = torch.zeros([input_ids.shape[0], 1])
outputs = inference_with_io_binding(session, config, input_ids, position_ids, attention_mask, empty_past, beam_select_idx, input_log_probs, input_unfinished_sents, input_ids, prev_step_scores, 0, input_ids.shape[-1])
assert torch.eq(outputs[-2], torch.from_numpy(ort_outputs[-2])).all()
print("IO Binding result is good")
# -
# ### バッチ推論
# + gather={"logged": 1628068303255}
def update(output, step, batch_size, beam_size, context_length, prev_attention_mask, device):
"""
Update the inputs for next inference.
"""
last_state = (torch.from_numpy(output[0]).to(device)
if isinstance(output[0], numpy.ndarray) else output[0].clone().detach().cpu())
input_ids = last_state.view(batch_size * beam_size, -1).to(device)
input_unfinished_sents_id = -3
prev_step_results = (torch.from_numpy(output[-2]).to(device) if isinstance(output[-2], numpy.ndarray)
else output[-2].clone().detach().to(device))
position_ids = (torch.tensor([context_length + step - 1
]).unsqueeze(0).repeat(batch_size * beam_size, 1).to(device))
if prev_attention_mask.shape[0] != (batch_size * beam_size):
prev_attention_mask = prev_attention_mask.repeat(batch_size * beam_size, 1)
attention_mask = torch.cat(
[
prev_attention_mask,
torch.ones([batch_size * beam_size, 1]).type_as(prev_attention_mask),
],
1,
).to(device)
beam_select_idx = (torch.from_numpy(output[input_unfinished_sents_id - 2]).to(device) if isinstance(
output[input_unfinished_sents_id - 2], numpy.ndarray) else output[input_unfinished_sents_id - 2].clone().detach().to(device))
input_log_probs = (torch.from_numpy(output[input_unfinished_sents_id - 1]).to(device) if isinstance(
output[input_unfinished_sents_id - 1], numpy.ndarray) else output[input_unfinished_sents_id - 1].clone().detach().to(device))
input_unfinished_sents = (torch.from_numpy(output[input_unfinished_sents_id]).to(device) if isinstance(
output[input_unfinished_sents_id], numpy.ndarray) else
output[input_unfinished_sents_id].clone().detach().to(device))
prev_step_scores = (torch.from_numpy(output[-1]).to(device)
if isinstance(output[-1], numpy.ndarray) else output[-1].clone().detach().to(device))
past = []
if isinstance(output[1], tuple): # past in torch output is tuple
past = list(output[1])
else:
for i in range(model.config.n_layer):
past_i = (torch.from_numpy(output[i + 1])
if isinstance(output[i + 1], numpy.ndarray) else output[i + 1].clone().detach())
past.append(past_i.to(device))
inputs = {
'input_ids': input_ids,
'attention_mask' : attention_mask,
'position_ids': position_ids,
'beam_select_idx': beam_select_idx,
'input_log_probs': input_log_probs,
'input_unfinished_sents': input_unfinished_sents,
'prev_step_results': prev_step_results,
'prev_step_scores': prev_step_scores,
}
ort_inputs = {
'input_ids': numpy.ascontiguousarray(input_ids.cpu().numpy()),
'attention_mask' : numpy.ascontiguousarray(attention_mask.cpu().numpy()),
'position_ids': numpy.ascontiguousarray(position_ids.cpu().numpy()),
'beam_select_idx': numpy.ascontiguousarray(beam_select_idx.cpu().numpy()),
'input_log_probs': numpy.ascontiguousarray(input_log_probs.cpu().numpy()),
'input_unfinished_sents': numpy.ascontiguousarray(input_unfinished_sents.cpu().numpy()),
'prev_step_results': numpy.ascontiguousarray(prev_step_results.cpu().numpy()),
'prev_step_scores': numpy.ascontiguousarray(prev_step_scores.cpu().numpy()),
}
for i, past_i in enumerate(past):
ort_inputs[f'past_{i}'] = numpy.ascontiguousarray(past_i.cpu().numpy())
return inputs, ort_inputs, past
def test_generation(tokenizer, input_text, use_onnxruntime_io, ort_session = None, num_tokens_to_produce = 30):
print("Text generation using", "OnnxRuntime with IO binding" if use_onnxruntime_io else "OnnxRuntime", "...")
input_ids, attention_mask, position_ids, past = get_example_inputs(input_text)
beam_select_idx = torch.zeros([1, input_ids.shape[0]]).long()
input_log_probs = torch.zeros([input_ids.shape[0], 1])
input_unfinished_sents = torch.ones([input_ids.shape[0], 1], dtype=torch.bool)
prev_step_scores = torch.zeros([input_ids.shape[0], 1])
inputs = {
'input_ids': input_ids,
'attention_mask' : attention_mask,
'position_ids': position_ids,
'beam_select_idx': beam_select_idx,
'input_log_probs': input_log_probs,
'input_unfinished_sents': input_unfinished_sents,
'prev_step_results': input_ids,
'prev_step_scores': prev_step_scores,
}
ort_inputs = {
'input_ids': numpy.ascontiguousarray(input_ids.cpu().numpy()),
'attention_mask' : numpy.ascontiguousarray(attention_mask.cpu().numpy()),
'position_ids': numpy.ascontiguousarray(position_ids.cpu().numpy()),
'beam_select_idx': numpy.ascontiguousarray(beam_select_idx.cpu().numpy()),
'input_log_probs': numpy.ascontiguousarray(input_log_probs.cpu().numpy()),
'input_unfinished_sents': numpy.ascontiguousarray(input_unfinished_sents.cpu().numpy()),
'prev_step_results': numpy.ascontiguousarray(input_ids.cpu().numpy()),
'prev_step_scores': numpy.ascontiguousarray(prev_step_scores.cpu().numpy()),
}
for i, past_i in enumerate(past):
ort_inputs[f'past_{i}'] = numpy.ascontiguousarray(past_i.cpu().numpy())
batch_size = input_ids.size(0)
beam_size = 4
context_length = input_ids.size(-1)
for step in range(num_tokens_to_produce):
if use_onnxruntime_io:
outputs = inference_with_io_binding(ort_session, config, inputs['input_ids'], inputs['position_ids'], inputs['attention_mask'], past, inputs['beam_select_idx'], inputs['input_log_probs'], inputs['input_unfinished_sents'], inputs['prev_step_results'], inputs['prev_step_scores'], step, context_length)
else:
outputs = ort_session.run(None, ort_inputs)
inputs, ort_inputs, past = update(outputs, step, batch_size, beam_size, context_length, inputs['attention_mask'], device)
if not inputs['input_unfinished_sents'].any():
break
print("------------")
print(tokenizer.decode(inputs['prev_step_results'][0], skip_special_tokens=True))
# + gather={"logged": 1628068306335}
import time
input_text = EXAMPLE_Text
# -
# ### 通常の推論
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}} gather={"logged": 1628068311176}
start = time.time()
test_generation(tokenizer, input_text, use_onnxruntime_io=False, ort_session=session)
elapsed_time = time.time() - start
print ("elapsed_time:{0}".format(elapsed_time) + "[sec]")
# -
# ### IO binding 有効 (GPU 推論をしてないので意味なし)
# + gather={"logged": 1628068316843}
start = time.time()
test_generation(tokenizer, input_text, use_onnxruntime_io=True, ort_session=session)
elapsed_time = time.time() - start
print ("elapsed_time:{0}".format(elapsed_time) + "[sec]")
# + [markdown] nteract={"transient": {"deleting": false}}
# ### 量子化した軽量 GPT-2
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}} gather={"logged": 1628068340468}
session_int8 = onnxruntime.InferenceSession(quantized_model_path)
start = time.time()
test_generation(tokenizer, input_text, use_onnxruntime_io=False, ort_session=session_int8)
elapsed_time = time.time() - start
print ("elapsed_time:{0}".format(elapsed_time) + "[sec]")
# -
| examples/rinna-gpt2-predict/convert_model.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .sh
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Bash
# language: bash
# name: bash
# ---
# [](https://cloudevel.com)
# # Balanceo de cargas.
# https://cloud.google.com/load-balancing/docs
# ## Tipos de balanceo de cargas.
#
# * Balanceo de cargas de *HTTP*(*S*) externo.
# * Regional.
# * Global.
# * Balanceo de cargas de *HTTP*(*S*) internono.
# * Balanceo de cargas *TCP/UDP* externo.
# * Balanceo de cargas *TCP/UDP* interno.
# * Balanceo de cargas del *proxy* de *SSL*.
# * Balanceo de cargas del *proxy* de *TCP*.
#
# https://cloud.google.com/load-balancing/docs/features
# ## Verificación de estado.
#
# https://cloud.google.com/load-balancing/docs/health-check-concepts
# ```
# gcloud compute health-checks
# ```
# ### Estados.
#
# * ```HEALTHY```
# * ```DRAINING```
# * ```UNHEALTHY```
# * ```TIMEOUT```
# * ```UNKNOWN```
# ### Políticas de reparación.
#
#
# <p style="text-align: center"><a rel="license" href="http://creativecommons.org/licenses/by/4.0/"><img alt="Licencia Creative Commons" style="border-width:0" src="https://i.creativecommons.org/l/by/4.0/80x15.png" /></a><br />Esta obra está bajo una <a rel="license" href="http://creativecommons.org/licenses/by/4.0/">Licencia Creative Commons Atribución 4.0 Internacional</a>.</p>
# <p style="text-align: center">© <NAME>. 2021.</p>
| 13_balanceo_de_cargas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ----------------------
# Load required packages
# ----------------------
import copy
import gc
import sys, getopt
import ujson as json
import random
import emcee
import datetime
import corner
import argparse
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import multiprocessing as mp
from multiprocessing import Pool
from covid19model.models import models
from covid19model.data import mobility, sciensano, model_parameters, VOC
from covid19model.models.time_dependant_parameter_fncs import ramp_fun
from covid19model.visualization.output import _apply_tick_locator
# OPTIONAL: Load the "autoreload" extension so that package code can change
# %load_ext autoreload
# OPTIONAL: always reload modules so that as you change code in src, it gets loaded
# %autoreload 2
# Path where figures and results should be stored
fig_path = '../../results/predictions/national/schools-easter-FvdB/'
# Path where MCMC samples should be saved
samples_path = '../../data/interim/model_parameters/COVID19_SEIRD/calibrations/national/'
# +
# -----------------------
# Load samples dictionary
# -----------------------
from covid19model.models.utils import load_samples_dict
samples_dict = load_samples_dict(samples_path+'BE_WAVE2_R0_COMP_EFF_2021-05-21.json', wave=2)
warmup = int(samples_dict['warmup'])
# +
# ---------
# Load data
# ---------
# Time-integrated contact matrices
initN, Nc_all = model_parameters.get_integrated_willem2012_interaction_matrices()
levels = initN.size
# Sciensano public data
df_sciensano = sciensano.get_sciensano_COVID19_data(update=True)
# Sciensano mortality data
df_sciensano_mortality =sciensano.get_mortality_data()
# Google Mobility data
df_google = mobility.get_google_mobility_data(update=False,plot=False)
# Serological data
df_sero_herzog, df_sero_sciensano = sciensano.get_serological_data()
# VOC data
df_VOC_501Y = VOC.get_501Y_data()
# Start of data collection
start_data = df_sciensano.idxmin()
# Start of calibration warmup and beta
start_calibration = samples_dict['start_calibration']
# Last datapoint used to calibrate warmup and beta
end_calibration = samples_dict['end_calibration']
# +
# ---------------------------
# Time-dependant VOC function
# ---------------------------
from covid19model.models.time_dependant_parameter_fncs import make_VOC_function
VOC_function = make_VOC_function(df_VOC_501Y)
# -----------------------------------
# Time-dependant vaccination function
# -----------------------------------
from covid19model.models.time_dependant_parameter_fncs import make_vaccination_function
vacc_strategy = make_vaccination_function(df_sciensano)
# +
# --------------------------------------
# Time-dependant social contact function
# --------------------------------------
# Extract build contact matrix function
from covid19model.models.time_dependant_parameter_fncs import make_contact_matrix_function, ramp_fun, delayed_ramp_fun
contact_matrix_4prev = make_contact_matrix_function(df_google, Nc_all)
# Define policy function
def policies_schools_easter_FVdB(t, states, param, l , prev_schools, prev_work, prev_rest, prev_home, scenario=0):
# Convert tau and l to dates
l_days = pd.Timedelta(l, unit='D')
# Define key dates of first wave
t1 = pd.Timestamp('2020-03-15') # start of lockdown
t2 = pd.Timestamp('2020-05-15') # gradual re-opening of schools (assume 50% of nominal scenario)
t3 = pd.Timestamp('2020-07-01') # start of summer holidays
t4 = pd.Timestamp('2020-09-01') # end of summer holidays
# Define key dates of second wave
t5 = pd.Timestamp('2020-10-19') # lockdown (1)
t6 = pd.Timestamp('2020-11-02') # lockdown (2)
t7 = pd.Timestamp('2020-11-16') # schools re-open
t8 = pd.Timestamp('2020-12-18') # Christmas holiday starts
t9 = pd.Timestamp('2021-01-04') # Christmas holiday ends
t10 = pd.Timestamp('2021-02-15') # Spring break starts
t11 = pd.Timestamp('2021-02-21') # Spring break ends
if scenario == 0:
# Paasvakantie as is
t12 = pd.Timestamp('2021-04-02')
t13 = pd.Timestamp('2021-04-18')
elif scenario == 1:
# Extend one week before --> chosen
t12 = pd.Timestamp('2021-03-26')
t13 = pd.Timestamp('2021-04-18')
elif scenario == 2:
# Extend one week after
t12 = pd.Timestamp('2021-04-02')
t13 = pd.Timestamp('2021-04-25')
elif scenario == 3:
# Shift one week
t12 = pd.Timestamp('2021-03-26')
t13 = pd.Timestamp('2021-04-11')
elif scenario == 4:
# No easter holiday
t12 = pd.Timestamp('2021-04-02')
t13 = pd.Timestamp('2021-04-02')
elif scenario == 5:
# Close schools
t12 = pd.Timestamp('2021-03-26')
t13 = pd.Timestamp('2021-06-30')
t14 = pd.Timestamp('2021-07-01') # Summer holiday starts
t = pd.Timestamp(t.date())
# First wave
if t <= t1:
return all_contact(t)
elif t1 < t < t1 :
return all_contact(t)
elif t1 < t <= t1 + l_days:
policy_old = all_contact(t)
policy_new = contact_matrix_4prev(t, prev_home, prev_schools, prev_work, prev_rest,
school=0)
return ramp_fun(policy_old, policy_new, t, t1, l)
elif t1 + l_days < t <= t2:
return contact_matrix_4prev(t, prev_home, prev_schools, prev_work, prev_rest,
school=0)
elif t2 < t <= t3:
return contact_matrix_4prev(t, prev_home, prev_schools, prev_work, prev_rest,
school=0)
elif t3 < t <= t4:
return contact_matrix_4prev(t, school=0)
# Second wave
elif t4 < t <= t5 :
return contact_matrix_4prev(t, school=1)
elif t5 < t <= t5 + l_days:
policy_old = contact_matrix_4prev(t, school=1)
policy_new = contact_matrix_4prev(t, prev_schools, prev_work, prev_rest,
school=1)
return ramp_fun(policy_old, policy_new, t, t5, l)
elif t5 + l_days < t <= t6:
return contact_matrix_4prev(t, prev_home, prev_schools, prev_work, prev_rest,
school=1)
elif t6 < t <= t7:
return contact_matrix_4prev(t, prev_home, prev_schools, prev_work, prev_rest,
school=0)
elif t7 < t <= t8:
return contact_matrix_4prev(t, prev_home, prev_schools, prev_work, prev_rest,
school=1)
elif t8 < t <= t9:
return contact_matrix_4prev(t, prev_home, prev_schools, prev_work, prev_rest,
school=0)
elif t9 < t <= t10:
return contact_matrix_4prev(t, prev_home, prev_schools, prev_work, prev_rest,
school=1)
elif t10 < t <= t11:
return contact_matrix_4prev(t, prev_home, prev_schools, prev_work, prev_rest,
school=0)
elif t11 < t <= t12:
return contact_matrix_4prev(t, prev_home, prev_schools, prev_work, prev_rest,
school=1)
elif t12 < t <= t13:
return contact_matrix_4prev(t, prev_home, prev_schools, prev_work, prev_rest,
school=0)
elif t13 < t <= t14:
return contact_matrix_4prev(t, prev_home, prev_schools, prev_work, prev_rest,
school=1)
else:
t = pd.Timestamp(t.date())
return contact_matrix_4prev(t, prev_home, prev_schools, prev_work, prev_rest,
school=0)
# +
# ------------------------------------------------
# Function to add binomial draws and draw function
# ------------------------------------------------
from covid19model.models.utils import add_poisson, output_to_visuals, draw_fcn_WAVE2
# +
# -------------------------------------
# Initialize the model with vaccination
# -------------------------------------
# Model initial condition on September 1st
with open('../../data/interim/model_parameters/COVID19_SEIRD/calibrations/national/initial_states_2020-09-01.json', 'r') as fp:
initial_states_vacc = json.load(fp)
params_vacc = model_parameters.get_COVID19_SEIRD_parameters(vaccination=True)
# Update with additional parameters for social policy function
params_vacc.update({'l': 21, 'prev_schools': 0, 'prev_work': 0.5, 'prev_rest': 0.5,'prev_home': 0.5, 'scenario': 1})
# VOCs
params_vacc.update({'t_sig': '2021-07-01'})
# Update with additional parameters for vaccination
params_vacc.update(
{'vacc_order': np.array(range(9))[::-1], 'daily_dose': 55000,
'refusal': 0.3*np.ones(9), 'delay': 20}
)
# Initialize model
model_vacc = models.COVID19_SEIRD_vacc(initial_states_vacc, params_vacc,
time_dependent_parameters={'Nc': policies_schools_easter_FVdB, 'N_vacc': vacc_strategy, 'alpha': VOC_function})
# +
# -----------------
# Scenario settings
# -----------------
scenarios = [0]
start_sim = '2020-09-01'
end_sim = '2021-09-21'
n_samples = 50
n_draws = 1
simulate = False
save = False
if simulate:
# ---------------------------
# Pre-allocation of dataframe
# ---------------------------
index = pd.date_range(start=start_sim, end=end_sim)
columns = [['0','0','0','1','1','1','2','2','2','3','3','3','4','4','4'],
['mean','LL','UL','mean','LL','UL','mean','LL','UL','mean','LL','UL','mean','LL','UL']]
tuples = list(zip(*columns))
columns = pd.MultiIndex.from_tuples(tuples, names=["School scenario", "Model output"])
data = np.zeros([len(index),15])
df_sim = pd.DataFrame(data=data, index=index, columns=columns)
# -------------------
# Perform simulations
# -------------------
for idx,scenario in enumerate(scenarios):
print('Simulating scenario '+str(scenario))
model_vacc.parameters.update({'scenario': scenario})
out_vacc = model_vacc.sim(end_sim,start_date=start_sim,warmup=warmup,N=n_samples,draw_fcn=draw_fcn_WAVE2,samples=samples_dict)
simtime, mean, median, LL, UL = add_poisson('H_in', out_vacc, n_samples, n_draws)
df_sim[str(scenario), "mean"] = mean
df_sim[str(scenario), "LL"] = LL
df_sim[str(scenario), "UL"] = UL
# --------------
# Save dataframe
# --------------
if save:
df_sim.to_csv('../../results/predictions/national/schools-easter-FVdB/simulations-schools-easter.csv')
else:
df_sim = pd.read_csv('../../results/predictions/national/schools-easter-FVdB/simulations-schools-easter.csv', header=[0,1], index_col=0, parse_dates=True)
# -
fig,ax = plt.subplots(figsize=(12,4))
ax.fill_between(simtime-pd.Timedelta(days=21),out_vacc['S_v'].sum(dim="Nc").quantile(dim="draws",q=1)/1e6, out_vacc['S_v'].sum(dim="Nc").quantile(dim="draws",q=0)/1e6, color='blue', alpha=0.2)
ax.axhline((11.4e6-initN[0]-initN[1])/1e6,linestyle='--',color='black',linewidth=1.5)
ax.set_xlim('2021-01-01',end_sim)
ax.set_ylim(0,11.4)
ax.xaxis.set_major_locator(plt.MaxNLocator(5))
ax.set_ylabel('Vaccinated individuals (millions)')
ax.grid(False)
fig.savefig('../../results/predictions/national/schools-easter-FVdB/uptake.pdf', dpi=300, bbox_inches='tight')
fig.savefig('../../results/predictions/national/schools-easter-FVdB/uptake.png', dpi=300, bbox_inches='tight')
# +
scenarios = [0,1,2,3]
colors = ['blue','green','orange','red']
fig,ax = plt.subplots(figsize=(12,5))
for idx,scenario in enumerate(scenarios):
ax.plot(df_sim.index, df_sim[str(scenario), 'mean'],'--', color=colors[idx], linewidth=1)
ax.fill_between(df_sim.index, df_sim[str(scenario), 'LL'], df_sim[str(scenario), 'UL'], alpha=0.20, color = colors[idx])
ax.scatter(df_sciensano[start_calibration:end_calibration].index,df_sciensano['H_in'][start_calibration:end_calibration], color='black', alpha=0.4, linestyle='None', facecolors='none', s=60, linewidth=2)
ax.scatter(df_sciensano[pd.to_datetime(end_calibration)+datetime.timedelta(days=1):end_sim].index,df_sciensano['H_in'][pd.to_datetime(end_calibration)+datetime.timedelta(days=1):end_sim], color='red', alpha=0.4, linestyle='None', facecolors='none', s=60, linewidth=2)
ax = _apply_tick_locator(ax)
ax.set_xlim('2020-09-01',end_sim)
ax.set_ylabel('$H_{in}$ (-)')
ax.set_title('Epidemiological model: BE COVID-19 SEIQRD with waning immunity\nSocial policy model: Current contact behaviour (without relaxations)\nBritish variant: 42% $\pm$ 2% more infectious, 30-50% (uniform) increase hospitalization propensity\nVaccination strategy: Data Sciensano (Dec. 28th, 2020 - present) + 60K-80K doses/day (future)\nVaccine properties: 90-99% effective, 80-100% reduction infectiousness, 80-100% lower hospitalization propensity\nVaccine refusal: 10% 60+ yo, 20% refusal 0-60 yo\n',loc='left',fontsize=13)
ax.legend(['Easter holiday as is','Extend one week before', 'Extend one week after','Shift one week'], bbox_to_anchor=(1.05, 1), loc='upper left', fontsize=13)
ax.axvline('2021-03-21',linestyle='--',color='black',linewidth=1.5)
im = mpimg.imread('logo_UGent_EN_RGB_2400_color-on-white.png')
newax = fig.add_axes([0.89, 0.90, 0.35, 0.35], anchor='NE', zorder=-1)
newax.imshow(im)
newax.axis('off')
fig.savefig('../../results/predictions/national/schools-easter-FVdB/twallema-schools_0.pdf', dpi=300, bbox_inches='tight')
fig.savefig('../../results/predictions/national/schools-easter-FVdB/twallema-schools_0.jpg', dpi=300, bbox_inches='tight')
# +
start_calibration = start_sim
scenarios = [0,1,2,3]
legend_entries = ['Extend one week before', 'Extend one week after', 'Shift one week']
fig,axes = plt.subplots(ncols=1,nrows=3,figsize=(12,12), sharey=True, sharex=True)
# -------------------------------------
# Plot baseline scenario in every graph
# -------------------------------------
for idx,ax in enumerate(axes):
ax.plot(df_sim.index, df_sim["0","mean"],'--', color='blue', linewidth=1.5)
ax.fill_between(df_sim.index, df_sim["0","LL"], df_sim["0","UL"],alpha=0.20, color = 'blue')
for idx,scenario in enumerate(scenarios[1:]):
# -----------
# Add to plot
# -----------
ax = axes[idx]
ax.plot(df_sim.index, df_sim[str(scenario),"mean"], '--', color='green', linewidth=1)
ax.fill_between(df_sim.index, df_sim[str(scenario),"LL"], df_sim[str(scenario),"UL"], alpha=0.20, color = 'green')
ax.scatter(df_sciensano[start_calibration:end_calibration].index,df_sciensano['H_in'][start_calibration:end_calibration], color='black', alpha=0.3, linestyle='None', facecolors='none', s=60, linewidth=2)
ax.scatter(df_sciensano[pd.to_datetime(end_calibration)+datetime.timedelta(days=1):end_sim].index,df_sciensano['H_in'][pd.to_datetime(end_calibration)+datetime.timedelta(days=1):end_sim], color='red', alpha=0.3, linestyle='None', facecolors='none', s=60, linewidth=2)
ax = _apply_tick_locator(ax)
ax.xaxis.set_major_locator(plt.MaxNLocator(5))
ax.xaxis.set_tick_params(labelsize=13, rotation=30)
ax.set_xlim('2020-09-01',end_sim)
ax.set_ylabel('$H_{in}$ (-)')
ax.legend(['Easter holiday as is',legend_entries[idx]], bbox_to_anchor=(1.05, 1), loc='upper left', fontsize=12)
ax.axvline('2021-03-21',linestyle='--',color='black',linewidth=1.5)
im = mpimg.imread('logo_UGent_EN_RGB_2400_color-on-white.png')
newax = fig.add_axes([0.95, 0.87, 0.2, 0.2], anchor='NE', zorder=-1)
newax.imshow(im)
newax.axis('off')
#fig.text(x=0.25,y=0.25,s='preliminary', rotation=30, alpha=0.2, fontsize = 110)
fig.suptitle('Epidemiological model: BE COVID-19 SEIQRD with waning immunity\nSocial policy model: Current contact behaviour (without relaxations)\nBritish variant: 42% $\pm$ 2% more infectious, 30-50% (uniform) increase hospitalization propensity\nVaccination strategy: Data Sciensano (Dec. 28th, 2020 - present) + 60K-80K doses/day (future)\nVaccine properties: 90-99% effective, 80-100% reduction infectiousness, 80-100% lower hospitalization propensity\nVaccine refusal: 10% 60+ yo, 20% refusal 0-60 yo\n',
x=0.1, y=1.04, ha='left', fontsize=13)
fig.savefig('../../results/predictions/national/schools-easter-FVdB/twallema-schools_3.pdf', dpi=300, bbox_inches='tight')
fig.savefig('../../results/predictions/national/schools-easter-FVdB/twallema-schools_3.jpg', dpi=300, bbox_inches='tight')
# +
scenarios = [1,4,5]
colors = ['blue','green','orange']
fig,ax = plt.subplots(figsize=(12,5))
for idx,scenario in enumerate(scenarios):
ax.plot(df_sim.index, df_sim[str(scenario),'mean'],'--', color=colors[idx], linewidth=1)
ax.fill_between(df_sim.index, df_sim[str(scenario),'LL'], df_sim[str(scenario),'UL'],alpha=0.20, color = colors[idx])
ax.scatter(df_sciensano[start_calibration:end_calibration].index,df_sciensano['H_in'][start_calibration:end_calibration], color='black', alpha=0.4, linestyle='None', facecolors='none', s=60, linewidth=2)
ax.scatter(df_sciensano[pd.to_datetime(end_calibration)+datetime.timedelta(days=1):end_sim].index,df_sciensano['H_in'][pd.to_datetime(end_calibration)+datetime.timedelta(days=1):end_sim], color='red', alpha=0.4, linestyle='None', facecolors='none', s=60, linewidth=2)
ax.axvline('2021-03-21',linestyle='--',color='black',linewidth=1.5)
ax = _apply_tick_locator(ax)
ax.set_xlim('2020-09-01',end_sim)
ax.set_ylabel('$H_{in}$ (-)')
ax.set_title('Epidemiological model: BE COVID-19 SEIQRD with waning immunity\nSocial policy model: Current contact behaviour (without relaxations)\nBritish variant: 42% $\pm$ 2% more infectious, 30-50% (uniform) increase hospitalization propensity\nVaccination strategy: Data Sciensano (Dec. 28th, 2020 - present) + 60K-80K doses/day (future)\nVaccine properties: 90-99% effective, 80-100% reduction infectiousness, 80-100% lower hospitalization propensity\nVaccine refusal: 10% 60+ yo, 20% refusal 0-60 yo\n',loc='left',fontsize=13)
ax.legend(['Extend one week before\n(2021/03/26 - 2021/04-18)','No easter holiday', 'Close schools on 2021/03/26'], bbox_to_anchor=(1.05, 1), loc='upper left', fontsize=13)
im = mpimg.imread('logo_UGent_EN_RGB_2400_color-on-white.png')
newax = fig.add_axes([0.92, 0.90, 0.35, 0.35], anchor='NE', zorder=-1)
newax.imshow(im)
newax.axis('off')
fig.savefig('../../results/predictions/national/schools-easter-FVdB/twallema-schools_2.pdf', dpi=300, bbox_inches='tight')
fig.savefig('../../results/predictions/national/schools-easter-FVdB/twallema-schools_2.jpg', dpi=300, bbox_inches='tight')
# -
| notebooks/prediction/twallema-schools-easter-FVdB.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:math]
# language: python
# name: conda-env-math-py
# ---
# Number of Trials
# $n \in \mathbb{N}$
#
# Success Probablity in Each Trial
# $p \in [0,1]$
#
# Number of Successes
# $k \in \{0, ..., n\}$
#
# Distribution Notation
# $X \sim B(n,p)$
#
# Bionomial Coefficient
# $\binom{n}{k} = \frac{n!}{k!(n - k)!}$
#
# <h4 style="text-align:center;">Probablity mass function</h4>
#
# $$f(k; n, p) = P(X = k) = \binom{n}{k}p^{k}(1 - p)^{n-k}$$
# +
import numpy as np
from scipy.special import binom
from matplotlib import pyplot as plt
# %matplotlib inline
def binom_coeff(k, n):
return factorial(n) / (factorial(k) * factorial(n-k))
def p_success(k, p):
return p**k
def p_failure(k, n, p):
return (1-p)**(n-k)
def pmf(k, n, p):
return binom_coeff(k, n) * p_success(k, p) * p_failure(k, n, p)
# -
n = 20
for p in [0.3, 0.5, 0.7]:
for k in range(0,n+1):
color={0.3:'r', 0.5: 'g', 0.7: 'b'}
plt.scatter(k, pmf(k, n, p), color=color[p])
plt.show()
n = 40
for p in [0.3, 0.5, 0.7]:
for k in range(0,n+1):
color={0.3:'r', 0.5: 'g', 0.7: 'b'}
plt.scatter(k, pmf(k, n, p), color=color[p])
plt.show()
| notebooks/BinomialDistribution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:RoutingCategories] *
# language: python
# name: conda-env-RoutingCategories-py
# ---
# %cd ..
import argparse
import collections
import random
import pyro
import torch
import matplotlib.pyplot as plt
import numpy as np
import data_loader.data_loaders as module_data
import model.model as module_arch
from parse_config import ConfigParser
from trainer import Trainer
# %matplotlib inline
# +
# pyro.enable_validation(True)
# torch.autograd.set_detect_anomaly(True)
# -
# fix random seeds for reproducibility
SEED = 123
torch.manual_seed(SEED)
torch.backends.cudnn.deterministic = True
torch.backends.cudnn.benchmark = False
np.random.seed(SEED)
random.seed(SEED)
Args = collections.namedtuple('Args', 'config resume device')
config = ConfigParser.from_args(Args(config='fashion_mnist_config.json', resume=None, device=None))
logger = config.get_logger('train')
# setup data_loader instances
data_loader = config.init_obj('data_loader', module_data)
valid_data_loader = data_loader.split_validation()
# build model architecture, then print to console
model = config.init_obj('arch', module_arch)
model.resume_from_checkpoint('saved/models/FashionMnist_VlaeCategory/1027_122354/checkpoint-epoch500.pth')
validation_data = random.choice(list(valid_data_loader))[0]
path, prediction_data = model(observations=validation_data)
prediction_data = prediction_data.view(-1, 1, 28, 28)
path.draw()
path.draw(path='fashion_mnist_string_diagram.pdf')
for k in np.random.randint(0, validation_data.shape[0], 10):
image = validation_data[k].view(28, 28).detach().cpu().numpy()
plt.title('Test image')
plt.imshow(image)
plt.show()
prediction = prediction_data[k].view(28, 28).detach().cpu().numpy()
plt.title('Reconstructed image')
plt.imshow(prediction)
plt.show()
for k in range(10):
path, sample = model(None)
sample = sample.view(28, 28).detach().cpu().numpy()
plt.show()
plt.title('Sample from prior')
plt.imshow(sample)
plt.show()
path.draw()
import torchvision
def show_img_grid(imgs, recons, path=None):
from PIL import Image
images = torch.cat((imgs, recons), dim=0)
if path is None:
grid = torchvision.utils.make_grid(images, nrow=imgs.shape[0])
grid = grid.mul(255).add_(0.5).clamp_(0, 255).detach().permute(1, 2, 0).to('cpu', torch.uint8)
img = Image.fromarray(grid.numpy())
plt.imshow(img)
else:
torchvision.utils.save_image(images, path, nrow=imgs.shape[0])
show_img_grid(validation_data[:5], prediction_data[:5], path='fashion_mnist_reconstructions.pdf')
show_img_grid(validation_data[:5], prediction_data[:5])
| notebooks/visualize_fashion_mnist.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: venv3.7.8
# language: python
# name: venv3.7.8
# ---
# # 1. Import libraries
# +
#----------------------------Reproducible----------------------------------------------------------------------------------------
import numpy as np
import random as rn
import os
seed=0
os.environ['PYTHONHASHSEED'] = str(seed)
np.random.seed(seed)
rn.seed(seed)
#----------------------------Reproducible----------------------------------------------------------------------------------------
os.environ['TF_CPP_MIN_LOG_LEVEL'] = '3'
import pandas as pd
import scipy.sparse as sparse
import scipy.io
from sklearn.linear_model import LinearRegression
import time
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import ShuffleSplit
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score
from sklearn.preprocessing import MinMaxScaler
#--------------------------------------------------------------------------------------------------------------------------------
#Import ourslef defined methods
import sys
sys.path.append(r"../Defined")
import Functions as F
# -
# # 2. Loading data
# +
data_path="../Dataset/GLIOMA.mat"
Data = scipy.io.loadmat(data_path)
data_arr_=Data['X']
label_arr=Data['Y'][:, 0]
data_arr=MinMaxScaler(feature_range=(0,1)).fit_transform(data_arr_)
label_arr_onehot=label_arr
# -
key_feture_number=64
# # 3. Calculation
# +
#--------------------------------------------------------------------------------------------------------------------------------
def IsnanAndIsinf(p_data):
p_data=np.array(p_data)
for i in np.arange(p_data.shape[0]):
for j in np.arange(p_data.shape[1]):
if np.isnan(p_data[i,j]) or np.isinf(p_data[i,j]):
p_data[i,j]=0
return p_data
#--------------------------------------------------------------------------------------------------------------------------------
def mse_check(train, test):
LR = LinearRegression(n_jobs = -1)
LR.fit(train[0], train[1])
MSELR = ((LR.predict(test[0]) - test[1]) ** 2).mean()
return MSELR
#--------------------------------------------------------------------------------------------------------------------------------
def InfFS(p_data_arr,p_alpha,use_specify_number=False,specify_number=50):
df = pd.DataFrame(p_data_arr)
corr_ij_spearman__=df.corr(method ='spearman')
corr_ij_spearman_=IsnanAndIsinf(corr_ij_spearman__)
corr_ij_spearman=1-np.abs(corr_ij_spearman_)
STD=np.std(p_data_arr,axis=0)
STDMatrix_=np.zeros((STD.shape[0],STD.shape[0]))
for i in np.arange(STD.shape[0]):
for j in np.arange(STD.shape[0]):
STDMatrix_[i,j]=max(STD[i],STD[j])
STDMatrix_min=STDMatrix_-np.min(STDMatrix_)
STDMatrix_max=np.max(STDMatrix_min)
STDMatrix__=STDMatrix_min/STDMatrix_max
STDMatrix=IsnanAndIsinf(STDMatrix__)
N=p_data_arr.shape[1]
eps = (5e-06) * N;
factor = 1 - eps
A = ( p_alpha*STDMatrix + (1-p_alpha)*corr_ij_spearman )
rho = np.max(np.sum(A,axis=1))
A = A / (rho+eps)
I = np.eye(A.shape[0])
r = factor/rho
y = I - ( r * A )
S=np.linalg.inv(y)
WEIGHT = np.sum( S , axis=1 )
RANKED=np.argsort(-WEIGHT)
RANKED = RANKED
WEIGHT = WEIGHT
e = np.ones(N)
t = np.dot(S, e)
nbins = 0.5*N
cnts, bins = np.histogram(t, bins=int(nbins))
thr =np.mean(cnts)
size_sub = np.sum(cnts>thr)
if use_specify_number:
size_sub=specify_number
SUBSET = RANKED[0:size_sub]
return SUBSET
#--------------------------------------------------------------------------------------------------------------------------------
def cal(p_data_arr,\
p_label_arr_onehot,\
p_key_feture_number,\
p_seed):
C_train_x,C_test_x,C_train_y,C_test_y= train_test_split(p_data_arr,p_label_arr_onehot,test_size=0.2,random_state=p_seed)
os.environ['PYTHONHASHSEED'] = str(p_seed)
np.random.seed(p_seed)
rn.seed(p_seed)
#--------------------------------------------------------------------------------------------------------------------------------
train_feature=C_train_x
test_feature=C_test_x
t_start = time.time()
train_idx=InfFS(train_feature,p_alpha,use_specify_number=True,specify_number=p_key_feture_number)
t_used=time.time() - t_start
C_train_selected_x = train_feature[:, train_idx]
test_idx=InfFS(test_feature,p_alpha,use_specify_number=True,specify_number=p_key_feture_number)
C_test_selected_x = test_feature[:, test_idx]
# Classification on original features
train_feature=C_train_x
train_label=C_train_y
test_feature=C_test_x
test_label=C_test_y
orig_train_acc,orig_test_acc=F.ETree(train_feature,train_label,test_feature,test_label,0)
# Classification on selected features
train_feature=C_train_selected_x
train_label=C_train_y
test_feature=C_test_selected_x
test_label=C_test_y
selec_train_acc,selec_test_acc=F.ETree(train_feature,train_label,test_feature,test_label,0)
# Linear reconstruction
train_feature_tuple=(C_train_selected_x,C_train_x)
test_feature_tuple=(C_test_selected_x,C_test_x)
reconstruction_loss=mse_check(train_feature_tuple, test_feature_tuple)
results=np.array([orig_train_acc,orig_test_acc,selec_train_acc,selec_test_acc,reconstruction_loss])
print(results)
return orig_train_acc,orig_test_acc,selec_train_acc,selec_test_acc,reconstruction_loss
# -
p_data_arr=data_arr
p_alpha=0.5
p_label_arr_onehot=label_arr_onehot
p_key_feture_number=key_feture_number
p_seed=0
orig_train_acc,orig_test_acc,selec_train_acc,selec_test_acc,reconstruction_loss=cal(p_data_arr,\
p_label_arr_onehot,\
p_key_feture_number,\
p_seed)
| Python/AbsoluteAndOtherAlgorithms/7GLIOMA/InfFS_64.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %run homework_modules.ipynb
import torch
from torch.autograd import Variable
import numpy
import unittest
# +
class TestLayers(unittest.TestCase):
def test_Linear(self):
np.random.seed(42)
torch.manual_seed(42)
batch_size, n_in, n_out = 2, 3, 4
for _ in range(100):
# layers initialization
torch_layer = torch.nn.Linear(n_in, n_out)
custom_layer = Linear(n_in, n_out)
custom_layer.W = torch_layer.weight.data.numpy()
custom_layer.b = torch_layer.bias.data.numpy()
layer_input = np.random.uniform(-10, 10, (batch_size, n_in)).astype(np.float32)
next_layer_grad = np.random.uniform(-10, 10, (batch_size, n_out)).astype(np.float32)
# 1. check layer output
custom_layer_output = custom_layer.updateOutput(layer_input)
layer_input_var = Variable(torch.from_numpy(layer_input), requires_grad=True)
torch_layer_output_var = torch_layer(layer_input_var)
self.assertTrue(np.allclose(torch_layer_output_var.data.numpy(), custom_layer_output, atol=1e-6))
# 2. check layer input grad
custom_layer_grad = custom_layer.updateGradInput(layer_input, next_layer_grad)
torch_layer_output_var.backward(torch.from_numpy(next_layer_grad))
torch_layer_grad_var = layer_input_var.grad
self.assertTrue(np.allclose(torch_layer_grad_var.data.numpy(), custom_layer_grad, atol=1e-6))
# 3. check layer parameters grad
custom_layer.accGradParameters(layer_input, next_layer_grad)
weight_grad = custom_layer.gradW
bias_grad = custom_layer.gradb
torch_weight_grad = torch_layer.weight.grad.data.numpy()
torch_bias_grad = torch_layer.bias.grad.data.numpy()
self.assertTrue(np.allclose(torch_weight_grad, weight_grad, atol=1e-6))
self.assertTrue(np.allclose(torch_bias_grad, bias_grad, atol=1e-6))
def test_SoftMax(self):
np.random.seed(42)
torch.manual_seed(42)
batch_size, n_in = 2, 4
for _ in range(100):
# layers initialization
torch_layer = torch.nn.Softmax(dim=1)
custom_layer = SoftMax()
layer_input = np.random.uniform(-10, 10, (batch_size, n_in)).astype(np.float32)
next_layer_grad = np.random.random((batch_size, n_in)).astype(np.float32)
next_layer_grad /= next_layer_grad.sum(axis=-1, keepdims=True)
next_layer_grad = next_layer_grad.clip(1e-5,1.)
next_layer_grad = 1. / next_layer_grad
# 1. check layer output
custom_layer_output = custom_layer.updateOutput(layer_input)
layer_input_var = Variable(torch.from_numpy(layer_input), requires_grad=True)
torch_layer_output_var = torch_layer(layer_input_var)
self.assertTrue(np.allclose(torch_layer_output_var.data.numpy(), custom_layer_output, atol=1e-5))
# 2. check layer input grad
custom_layer_grad = custom_layer.updateGradInput(layer_input, next_layer_grad)
torch_layer_output_var.backward(torch.from_numpy(next_layer_grad))
torch_layer_grad_var = layer_input_var.grad
self.assertTrue(np.allclose(torch_layer_grad_var.data.numpy(), custom_layer_grad, atol=1e-5))
def test_LogSoftMax(self):
np.random.seed(42)
torch.manual_seed(42)
batch_size, n_in = 2, 4
for _ in range(100):
# layers initialization
torch_layer = torch.nn.LogSoftmax(dim=1)
custom_layer = LogSoftMax()
layer_input = np.random.uniform(-10, 10, (batch_size, n_in)).astype(np.float32)
next_layer_grad = np.random.random((batch_size, n_in)).astype(np.float32)
next_layer_grad /= next_layer_grad.sum(axis=-1, keepdims=True)
# 1. check layer output
custom_layer_output = custom_layer.updateOutput(layer_input)
layer_input_var = Variable(torch.from_numpy(layer_input), requires_grad=True)
torch_layer_output_var = torch_layer(layer_input_var)
self.assertTrue(np.allclose(torch_layer_output_var.data.numpy(), custom_layer_output, atol=1e-6))
# 2. check layer input grad
custom_layer_grad = custom_layer.updateGradInput(layer_input, next_layer_grad)
torch_layer_output_var.backward(torch.from_numpy(next_layer_grad))
torch_layer_grad_var = layer_input_var.grad
self.assertTrue(np.allclose(torch_layer_grad_var.data.numpy(), custom_layer_grad, atol=1e-6))
def test_BatchNormalization(self):
np.random.seed(42)
torch.manual_seed(42)
batch_size, n_in = 32, 16
for _ in range(100):
# layers initialization
slope = np.random.uniform(0.01, 0.05)
alpha = 0.9
custom_layer = BatchNormalization(alpha)
custom_layer.train()
torch_layer = torch.nn.BatchNorm1d(n_in, eps=custom_layer.EPS, momentum=1.-alpha, affine=False)
custom_layer.moving_mean = torch_layer.running_mean.numpy().copy()
custom_layer.moving_variance = torch_layer.running_var.numpy().copy()
layer_input = np.random.uniform(-5, 5, (batch_size, n_in)).astype(np.float32)
next_layer_grad = np.random.uniform(-5, 5, (batch_size, n_in)).astype(np.float32)
# 1. check layer output
custom_layer_output = custom_layer.updateOutput(layer_input)
layer_input_var = Variable(torch.from_numpy(layer_input), requires_grad=True)
torch_layer_output_var = torch_layer(layer_input_var)
self.assertTrue(np.allclose(torch_layer_output_var.data.numpy(), custom_layer_output, atol=1e-6))
# 2. check layer input grad
custom_layer_grad = custom_layer.updateGradInput(layer_input, next_layer_grad)
torch_layer_output_var.backward(torch.from_numpy(next_layer_grad))
torch_layer_grad_var = layer_input_var.grad
# please, don't increase `atol` parameter, it's garanteed that you can implement batch norm layer
# with tolerance 1e-5
self.assertTrue(np.allclose(torch_layer_grad_var.data.numpy(), custom_layer_grad, atol=1e-5))
# 3. check moving mean
self.assertTrue(np.allclose(custom_layer.moving_mean, torch_layer.running_mean.numpy()))
# we don't check moving_variance because pytorch uses slightly different formula for it:
# it computes moving average for unbiased variance (i.e var*N/(N-1))
#self.assertTrue(np.allclose(custom_layer.moving_variance, torch_layer.running_var.numpy()))
# 4. check evaluation mode
custom_layer.moving_variance = torch_layer.running_var.numpy().copy()
custom_layer.evaluate()
custom_layer_output = custom_layer.updateOutput(layer_input)
torch_layer.eval()
torch_layer_output_var = torch_layer(layer_input_var)
self.assertTrue(np.allclose(torch_layer_output_var.data.numpy(), custom_layer_output, atol=1e-6))
def test_Sequential(self):
np.random.seed(42)
torch.manual_seed(42)
batch_size, n_in = 2, 4
for _ in range(100):
# layers initialization
alpha = 0.9
torch_layer = torch.nn.BatchNorm1d(n_in, eps=BatchNormalization.EPS, momentum=1.-alpha, affine=True)
torch_layer.bias.data = torch.from_numpy(np.random.random(n_in).astype(np.float32))
custom_layer = Sequential()
bn_layer = BatchNormalization(alpha)
bn_layer.moving_mean = torch_layer.running_mean.numpy().copy()
bn_layer.moving_variance = torch_layer.running_var.numpy().copy()
custom_layer.add(bn_layer)
scaling_layer = ChannelwiseScaling(n_in)
scaling_layer.gamma = torch_layer.weight.data.numpy()
scaling_layer.beta = torch_layer.bias.data.numpy()
custom_layer.add(scaling_layer)
custom_layer.train()
layer_input = np.random.uniform(-5, 5, (batch_size, n_in)).astype(np.float32)
next_layer_grad = np.random.uniform(-5, 5, (batch_size, n_in)).astype(np.float32)
# 1. check layer output
custom_layer_output = custom_layer.updateOutput(layer_input)
layer_input_var = Variable(torch.from_numpy(layer_input), requires_grad=True)
torch_layer_output_var = torch_layer(layer_input_var)
self.assertTrue(np.allclose(torch_layer_output_var.data.numpy(), custom_layer_output, atol=1e-6))
# 2. check layer input grad
custom_layer_grad = custom_layer.backward(layer_input, next_layer_grad)
torch_layer_output_var.backward(torch.from_numpy(next_layer_grad))
torch_layer_grad_var = layer_input_var.grad
self.assertTrue(np.allclose(torch_layer_grad_var.data.numpy(), custom_layer_grad, atol=1e-5))
# 3. check layer parameters grad
weight_grad, bias_grad = custom_layer.getGradParameters()[1]
torch_weight_grad = torch_layer.weight.grad.data.numpy()
torch_bias_grad = torch_layer.bias.grad.data.numpy()
self.assertTrue(np.allclose(torch_weight_grad, weight_grad, atol=1e-6))
self.assertTrue(np.allclose(torch_bias_grad, bias_grad, atol=1e-6))
def test_Dropout(self):
np.random.seed(42)
batch_size, n_in = 2, 4
for _ in range(100):
# layers initialization
p = np.random.uniform(0.3, 0.7)
layer = Dropout(p)
layer.train()
layer_input = np.random.uniform(-5, 5, (batch_size, n_in)).astype(np.float32)
next_layer_grad = np.random.uniform(-5, 5, (batch_size, n_in)).astype(np.float32)
# 1. check layer output
layer_output = layer.updateOutput(layer_input)
self.assertTrue(np.all(np.logical_or(np.isclose(layer_output, 0),
np.isclose(layer_output*(1.-p), layer_input))))
# 2. check layer input grad
layer_grad = layer.updateGradInput(layer_input, next_layer_grad)
self.assertTrue(np.all(np.logical_or(np.isclose(layer_grad, 0),
np.isclose(layer_grad*(1.-p), next_layer_grad))))
# 3. check evaluation mode
layer.evaluate()
layer_output = layer.updateOutput(layer_input)
self.assertTrue(np.allclose(layer_output, layer_input))
# 4. check mask
p = 0.0
layer = Dropout(p)
layer.train()
layer_output = layer.updateOutput(layer_input)
self.assertTrue(np.allclose(layer_output, layer_input))
p = 0.5
layer = Dropout(p)
layer.train()
layer_input = np.random.uniform(5, 10, (batch_size, n_in)).astype(np.float32)
next_layer_grad = np.random.uniform(5, 10, (batch_size, n_in)).astype(np.float32)
layer_output = layer.updateOutput(layer_input)
zeroed_elem_mask = np.isclose(layer_output, 0)
layer_grad = layer.updateGradInput(layer_input, next_layer_grad)
self.assertTrue(np.all(zeroed_elem_mask == np.isclose(layer_grad, 0)))
# 5. dropout mask should be generated independently for every input matrix element, not for row/column
batch_size, n_in = 1000, 1
p = 0.8
layer = Dropout(p)
layer.train()
layer_input = np.random.uniform(5, 10, (batch_size, n_in)).astype(np.float32)
layer_output = layer.updateOutput(layer_input)
self.assertTrue(np.sum(np.isclose(layer_output, 0)) != layer_input.size)
layer_input = layer_input.T
layer_output = layer.updateOutput(layer_input)
self.assertTrue(np.sum(np.isclose(layer_output, 0)) != layer_input.size)
def test_LeakyReLU(self):
np.random.seed(42)
torch.manual_seed(42)
batch_size, n_in = 2, 4
for _ in range(100):
# layers initialization
slope = np.random.uniform(0.01, 0.05)
torch_layer = torch.nn.LeakyReLU(slope)
custom_layer = LeakyReLU(slope)
layer_input = np.random.uniform(-5, 5, (batch_size, n_in)).astype(np.float32)
next_layer_grad = np.random.uniform(-5, 5, (batch_size, n_in)).astype(np.float32)
# 1. check layer output
custom_layer_output = custom_layer.updateOutput(layer_input)
layer_input_var = Variable(torch.from_numpy(layer_input), requires_grad=True)
torch_layer_output_var = torch_layer(layer_input_var)
self.assertTrue(np.allclose(torch_layer_output_var.data.numpy(), custom_layer_output, atol=1e-6))
# 2. check layer input grad
custom_layer_grad = custom_layer.updateGradInput(layer_input, next_layer_grad)
torch_layer_output_var.backward(torch.from_numpy(next_layer_grad))
torch_layer_grad_var = layer_input_var.grad
self.assertTrue(np.allclose(torch_layer_grad_var.data.numpy(), custom_layer_grad, atol=1e-6))
def test_ELU(self):
np.random.seed(42)
torch.manual_seed(42)
batch_size, n_in = 2, 4
for _ in range(100):
# layers initialization
alpha = 1.0
torch_layer = torch.nn.ELU(alpha)
custom_layer = ELU(alpha)
layer_input = np.random.uniform(-5, 5, (batch_size, n_in)).astype(np.float32)
next_layer_grad = np.random.uniform(-5, 5, (batch_size, n_in)).astype(np.float32)
# 1. check layer output
custom_layer_output = custom_layer.updateOutput(layer_input)
layer_input_var = Variable(torch.from_numpy(layer_input), requires_grad=True)
torch_layer_output_var = torch_layer(layer_input_var)
self.assertTrue(np.allclose(torch_layer_output_var.data.numpy(), custom_layer_output, atol=1e-6))
# 2. check layer input grad
custom_layer_grad = custom_layer.updateGradInput(layer_input, next_layer_grad)
torch_layer_output_var.backward(torch.from_numpy(next_layer_grad))
torch_layer_grad_var = layer_input_var.grad
self.assertTrue(np.allclose(torch_layer_grad_var.data.numpy(), custom_layer_grad, atol=1e-6))
def test_SoftPlus(self):
np.random.seed(42)
torch.manual_seed(42)
batch_size, n_in = 2, 4
for _ in range(100):
# layers initialization
torch_layer = torch.nn.Softplus()
custom_layer = SoftPlus()
layer_input = np.random.uniform(-5, 5, (batch_size, n_in)).astype(np.float32)
next_layer_grad = np.random.uniform(-5, 5, (batch_size, n_in)).astype(np.float32)
# 1. check layer output
custom_layer_output = custom_layer.updateOutput(layer_input)
layer_input_var = Variable(torch.from_numpy(layer_input), requires_grad=True)
torch_layer_output_var = torch_layer(layer_input_var)
self.assertTrue(np.allclose(torch_layer_output_var.data.numpy(), custom_layer_output, atol=1e-6))
# 2. check layer input grad
custom_layer_grad = custom_layer.updateGradInput(layer_input, next_layer_grad)
torch_layer_output_var.backward(torch.from_numpy(next_layer_grad))
torch_layer_grad_var = layer_input_var.grad
self.assertTrue(np.allclose(torch_layer_grad_var.data.numpy(), custom_layer_grad, atol=1e-6))
def test_ClassNLLCriterionUnstable(self):
np.random.seed(42)
torch.manual_seed(42)
batch_size, n_in = 2, 4
for _ in range(100):
# layers initialization
torch_layer = torch.nn.NLLLoss()
custom_layer = ClassNLLCriterionUnstable()
layer_input = np.random.uniform(0, 1, (batch_size, n_in)).astype(np.float32)
layer_input /= layer_input.sum(axis=-1, keepdims=True)
layer_input = layer_input.clip(custom_layer.EPS, 1. - custom_layer.EPS) # unifies input
target_labels = np.random.choice(n_in, batch_size)
target = np.zeros((batch_size, n_in), np.float32)
target[np.arange(batch_size), target_labels] = 1 # one-hot encoding
# 1. check layer output
custom_layer_output = custom_layer.updateOutput(layer_input, target)
layer_input_var = Variable(torch.from_numpy(layer_input), requires_grad=True)
torch_layer_output_var = torch_layer(torch.log(layer_input_var),
Variable(torch.from_numpy(target_labels), requires_grad=False))
self.assertTrue(np.allclose(torch_layer_output_var.data.numpy(), custom_layer_output, atol=1e-6))
# 2. check layer input grad
custom_layer_grad = custom_layer.updateGradInput(layer_input, target)
torch_layer_output_var.backward()
torch_layer_grad_var = layer_input_var.grad
self.assertTrue(np.allclose(torch_layer_grad_var.data.numpy(), custom_layer_grad, atol=1e-6))
def test_ClassNLLCriterion(self):
np.random.seed(42)
torch.manual_seed(42)
batch_size, n_in = 2, 4
for _ in range(100):
# layers initialization
torch_layer = torch.nn.NLLLoss()
custom_layer = ClassNLLCriterion()
layer_input = np.random.uniform(-5, 5, (batch_size, n_in)).astype(np.float32)
layer_input = torch.nn.LogSoftmax(dim=1)(Variable(torch.from_numpy(layer_input))).data.numpy()
target_labels = np.random.choice(n_in, batch_size)
target = np.zeros((batch_size, n_in), np.float32)
target[np.arange(batch_size), target_labels] = 1 # one-hot encoding
# 1. check layer output
custom_layer_output = custom_layer.updateOutput(layer_input, target)
layer_input_var = Variable(torch.from_numpy(layer_input), requires_grad=True)
torch_layer_output_var = torch_layer(layer_input_var,
Variable(torch.from_numpy(target_labels), requires_grad=False))
self.assertTrue(np.allclose(torch_layer_output_var.data.numpy(), custom_layer_output, atol=1e-6))
# 2. check layer input grad
custom_layer_grad = custom_layer.updateGradInput(layer_input, target)
torch_layer_output_var.backward()
torch_layer_grad_var = layer_input_var.grad
self.assertTrue(np.allclose(torch_layer_grad_var.data.numpy(), custom_layer_grad, atol=1e-6))
def test_adam_optimizer(self):
state = {}
config = {'learning_rate': 1e-3, 'beta1': 0.9, 'beta2':0.999, 'epsilon':1e-8}
variables = [[np.arange(10).astype(np.float64)]]
gradients = [[np.arange(10).astype(np.float64)]]
adam_optimizer(variables, gradients, config, state)
self.assertTrue(np.allclose(state['m'][0], np.array([0. , 0.1, 0.2, 0.3, 0.4, 0.5,
0.6, 0.7, 0.8, 0.9])))
self.assertTrue(np.allclose(state['v'][0], np.array([0., 0.001, 0.004, 0.009, 0.016, 0.025,
0.036, 0.049, 0.064, 0.081])))
self.assertTrue(state['t'] == 1)
self.assertTrue(np.allclose(variables[0][0], np.array([0., 0.999, 1.999, 2.999, 3.999, 4.999,
5.999, 6.999, 7.999, 8.999])))
adam_optimizer(variables, gradients, config, state)
self.assertTrue(np.allclose(state['m'][0], np.array([0., 0.19, 0.38, 0.57, 0.76, 0.95, 1.14,
1.33, 1.52, 1.71])))
self.assertTrue(np.allclose(state['v'][0], np.array([0., 0.001999, 0.007996, 0.017991,
0.031984, 0.049975, 0.071964, 0.097951,
0.127936, 0.161919])))
self.assertTrue(state['t'] == 2)
self.assertTrue(np.allclose(variables[0][0], np.array([0., 0.998, 1.998, 2.998, 3.998, 4.998,
5.998, 6.998, 7.998, 8.998])))
suite = unittest.TestLoader().loadTestsFromTestCase(TestLayers)
unittest.TextTestRunner(verbosity=2).run(suite)
# + active=""
#
# +
class TestAdvancedLayers(unittest.TestCase):
@unittest.skip("Not implemented yet")
def test_Conv2d(self):
np.random.seed(42)
torch.manual_seed(42)
batch_size, n_in, n_out = 2, 3, 4
h,w = 5,6
kern_size = 3
for _ in range(100):
# layers initialization
torch_layer = torch.nn.Conv2d(n_in, n_out, kern_size, padding=1)
custom_layer = Conv2d(n_in, n_out, kern_size)
custom_layer.W = torch_layer.weight.data.numpy() # [n_out, n_in, kern, kern]
custom_layer.b = torch_layer.bias.data.numpy()
layer_input = np.random.uniform(-1, 1, (batch_size, n_in, h,w)).astype(np.float32)
next_layer_grad = np.random.uniform(-1, 1, (batch_size, n_out, h, w)).astype(np.float32)
# 1. check layer output
custom_layer_output = custom_layer.updateOutput(layer_input)
layer_input_var = Variable(torch.from_numpy(layer_input), requires_grad=True)
torch_layer_output_var = torch_layer(layer_input_var)
self.assertTrue(np.allclose(torch_layer_output_var.data.numpy(), custom_layer_output, atol=1e-6))
# 2. check layer input grad
custom_layer_grad = custom_layer.updateGradInput(layer_input, next_layer_grad)
torch_layer_output_var.backward(torch.from_numpy(next_layer_grad))
torch_layer_grad_var = layer_input_var.grad
self.assertTrue(np.allclose(torch_layer_grad_var.data.numpy(), custom_layer_grad, atol=1e-6))
# 3. check layer parameters grad
custom_layer.accGradParameters(layer_input, next_layer_grad)
weight_grad = custom_layer.gradW
bias_grad = custom_layer.gradb
torch_weight_grad = torch_layer.weight.grad.data.numpy()
torch_bias_grad = torch_layer.bias.grad.data.numpy()
#m = ~np.isclose(torch_weight_grad, weight_grad, atol=1e-5)
self.assertTrue(np.allclose(torch_weight_grad, weight_grad, atol=1e-6, ))
self.assertTrue(np.allclose(torch_bias_grad, bias_grad, atol=1e-6))
@unittest.skip("Not implemented yet")
def test_MaxPool2d(self):
np.random.seed(42)
torch.manual_seed(42)
batch_size, n_in = 2, 3
h,w = 4,6
kern_size = 2
for _ in range(100):
# layers initialization
torch_layer = torch.nn.MaxPool2d(kern_size)
custom_layer = MaxPool2d(kern_size)
layer_input = np.random.uniform(-10, 10, (batch_size, n_in, h,w)).astype(np.float32)
next_layer_grad = np.random.uniform(-10, 10, (batch_size, n_in,
h // kern_size, w // kern_size)).astype(np.float32)
# 1. check layer output
custom_layer_output = custom_layer.updateOutput(layer_input)
layer_input_var = Variable(torch.from_numpy(layer_input), requires_grad=True)
torch_layer_output_var = torch_layer(layer_input_var)
self.assertTrue(np.allclose(torch_layer_output_var.data.numpy(), custom_layer_output, atol=1e-6))
# 2. check layer input grad
custom_layer_grad = custom_layer.updateGradInput(layer_input, next_layer_grad)
torch_layer_output_var.backward(torch.from_numpy(next_layer_grad))
torch_layer_grad_var = layer_input_var.grad
self.assertTrue(np.allclose(torch_layer_grad_var.data.numpy(), custom_layer_grad, atol=1e-6))
suite = unittest.TestLoader().loadTestsFromTestCase(TestAdvancedLayers)
unittest.TextTestRunner(verbosity=2).run(suite)
# -
| homework01/homework_test_modules.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: pytorch_env
# language: python
# name: pytorch_env
# ---
# ## Azure Machine Learning 配置
# ### 1. 获取配置文件
#
# 生成并保存配置文件,其中包含有关从本地环境连接到 Azure Azure Machine Learning的信息。
# +
from azureml.core import Workspace, Dataset
# 在下面三行代码填入配置信息
subscription_id = ''
resource_group = ''
workspace_name = ''
workspace = Workspace(subscription_id, resource_group, workspace_name)
# -
# 保存配置信息
workspace.write_config(path=".azureml")
# ### 2. 搭建计算群集
# 登录[Azure Machine Learning studio](ml.azure.com) 并从左侧菜单访问 Compute 以创建计算群集。有关信息,请参阅以下文档。
#
# ※ 文档 :[在 Azure 机器学习工作室中为模型训练和部署创建计算目标](https://docs.microsoft.com/zh-cn/azure/machine-learning/how-to-create-attach-compute-studio)
# 建议选择[Tesla V100](https://www.nvidia.com/en-gb/data-center/tesla-v100/) 对应GPU的机型 `Standard NC6s_v3`
conda --version
| examples/azureml-config.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/IswariAfala/tugas-fisika/blob/main/radioactive_decay.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/", "height": 299} id="RgS_-3XQvQeq" outputId="e9e77dc0-09ee-4288-e0ae-d85aafac10dc"
from __future__ import division
import numpy
import matplotlib.pyplot as pyplot
import scipy.integrate
import random
t_half_rad = 20.8 #initial conditions
t_half_act = 10.0
N0 = 250
t1 = 100
n_timepoints = 50
def analytic(N0, timebase):
'''Analytic solution for the radium count'''
return N0 * numpy.exp (-timebase /t_half_rad * numpy.log(2))
def simulate_monte_carlo(N0, t1, n_timepoints):
'''Monte carlo simulation for both radium and actinium counts'''
dt = t1 / n_timepoints #Calculating the interval between each time division
count_radium = numpy.zeros((n_timepoints)) #creating zero arrays to put the counts into
count_actinium = numpy.zeros((n_timepoints))
atoms = numpy.ones((N0)) #Creating an array of numbers to represent the atoms in the simulation
p_decay_rad = 1 - numpy.exp(-dt / t_half_rad * numpy.log(2)) #Calculating the decay probabilities in the time interval
p_decay_act = 1 - numpy.exp(-dt / t_half_act * numpy.log(2))
for idx_time in range(n_timepoints):
count_radium[idx_time] = (atoms == 1).sum() #Counting how many atoms of each type remain in the interval
count_actinium[idx_time] = (atoms == 2).sum()
for idx_atom in range(N0):
if atoms[idx_atom] == 1: #Deciding whether the given atom should decay
if random.random() <= p_decay_rad:
atoms[idx_atom] = 2
else:
atoms[idx_atom] = 1
elif atoms[idx_atom] == 2:
if random.random() <= p_decay_act:
atoms[idx_atom] = 3
else:
atoms[idx_atom] = 2
return count_radium, count_actinium
timebase = numpy.arange(0, t1, t1/n_timepoints) #creating the array of times for use in the analytic solution and scipy
n_analytic = analytic(N0, timebase) #Calling the analytic solution
n_rad, n_act = simulate_monte_carlo(N0, t1, n_timepoints) #Calling the Monte Carlo Simulation
def f(N, t):
'''Differential for the decay, for use with scipy.integrate.odeint'''
N_rad, N_act = N #unpacking N
tau_rad = t_half_rad / numpy.log(2)
tau_act = t_half_act / numpy.log(2)
DEQ_rad = - N_rad / tau_rad
DEQ_act = - N_act / tau_act + N_rad / tau_rad
return numpy.array((DEQ_rad, DEQ_act)) #repacking
N0_rad = 250 #Initial conditions for scipy
N0_act = 0
N0 = numpy.array((N0_rad, N0_act))
n_scipy = scipy.integrate.odeint(f, N0, timebase) #Calling scipy odeint
pyplot.figure() #Plotting code
pyplot.plot(timebase, n_rad, label = 'Monte Carlo Radium', color = 'blue')
pyplot.plot(timebase, n_act, label = 'Monte Carlo Actinium', color = 'red')
pyplot.plot(timebase, n_scipy[:,0], label = 'Scipy Radium', color = 'magenta')
pyplot.plot(timebase, n_scipy[:,1], label = 'Scipy Actinium', color = 'green')
pyplot.plot(timebase, n_analytic, label = 'Analytical Solution', color = 'black', linestyle = '--')
pyplot.title('Graph of the Decay of $^{225}$Ra and $^{225}$Ac')
pyplot.ylabel('Number of atoms')
pyplot.xlabel('time /days')
pyplot.legend(loc='upper right')
pyplot.show()
| radioactive_decay.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.7.2
# language: julia
# name: julia-1.7
# ---
# # Descriptive Statistics - Week 4 Lecture
using Distributions
using StatsBase
using CSV
using DataFrames
using HypothesisTests
using Plots
using GLM
using StatsPlots
pyplot()
age = rand(18:80, 100) # Uniform distribution
wcc = round.(rand(Distributions.Normal(12, 2), 100), digits=1) # Normal distribution & round to one decimal place
crp = round.(Int, rand(Distributions.Chisq(4), 100)) .* 10 # chi squared distributiion with broadcasting & alternate round()
treatment = rand(["A", "B"], 100) # Uniformaly weighted
result = rand(["Improved", "Static", "Worse"], 100) # Uniformaly weighted
# Mean of the age variable
mean(age)
# Median of the age variable
median(age)
# Standard deviation of age
std(age)
# Variance of age
var(age)
# Mean of wcc
mean(wcc)
## Standard deviation of wcc
std(wcc)
# Descriptie statistics of the age variable
StatsBase.describe(age)
# The summarystats() function omits the length and type
StatsBase.summarystats(wcc)
# # Creating a dataframe
data = DataFrame(Age = age, WCC = wcc, CRP = crp, Treatment = treatment, Result = result)
# Number of rows and columns
size(data)
# First six rows
first(data, 6)
# We can create dataframe objects by selecting only subjects according to their data point values for a particular variable
dataA = data[data[!,:Treatment] .== "A", :] # Only patient in treatment group A; note that .== uses punctation to perform function element-wise comparision
dataB = data[data[!,:Treatment] .== "B", :] # Only patient in treatment group B
first(dataA)
first(dataB)
last(dataA)
last(dataB)
describe(data)
# # Visualizing the data
# The Plots package works well with the DataFrames package by allowing macro function from the latter. In the code cell below, we look at the age distribution of the two treatment groups.
@df data density(:Age, group = :Treatment, title = "Distribution of ages by treatment group", xlab = "Age", ylab="Distribution", legend=:topright)
@df data density(:Age, group = :Result, title = "Distribution of ages by result group", xlab = "Age", ylab="Distribution", legend = :topright)
@df data density(:Age, group = (:Treatment, :Result), title="Distribution of ages by treatment and result groups",
xlab="Age", ylab="Distribution", legend=:topright)
@df data boxplot(:Treatment, :WCC, lab="WCC", title="White cell count by treatment group", xlab="Groups", ylab="WCC")
@df data boxplot(:Result, :WCC, lab="WCC", title="White cell count by result group", xlab="Groups", ylab="WCC")
@df data corrplot([:Age :WCC :CRP], grid=false)
@df data corrplot([:Age :WCC :CRP], grid=false, compact=true)
HypothesisTests.EqualVarianceTTest(dataA[!,:Age], dataB[!,:Age])
pvalue(EqualVarianceTTest(dataA[!, :WCC], dataB[!, :WCC]))
UnequalVarianceTTest(dataA[!, :CRP], dataB[!, :CRP])
fit(LinearModel, @formula(CRP ~ 1), data)
fit(LinearModel, @formula(CRP ~ Age), data)
fit(LinearModel, @formula(CRP ~ Age + WCC), data)
CSV.write("data/ProjectData_1_point_0.csv", data)
dataframe = CSV.read("data/ProjectData_1_point_0.csv", DataFrame)
DataFrame(Group=rand(["A", "B"], 20), Variable1=randn(20), Variable2=rand(20))
view(dataframe, 1:3, :)
delete!(dataframe,[1,2,3])
# +
view(df, 1:3, :)
delete!(df,[3,5,9])
# -
var(rand(Normal(80, 10), 200))
| DescriptiveStatistics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## _*Quantum SVM (variational method)*_
#
# The SVM_QKernel notebook here demonstrated a kernel based approach. This notebook shows a variational method.
#
# For further information please see: [https://arxiv.org/pdf/1804.11326.pdf](https://arxiv.org/pdf/1804.11326.pdf)
#
#
# **This notebook shows the SVM implementation based on the variational method.**
import os
import sys
from datasets import *
from qiskit_acqua.svm.data_preprocess import *
from qiskit_acqua.input import get_input_instance
from qiskit_acqua import run_algorithm
# First we prepare the dataset, which is used for training, testing and the finally prediction.
#
# *Note: You can easily switch to a different dataset, such as the Breast Cancer dataset, by replacing 'ad_hoc_data' to 'Breast_cancer' below.*
sample_Total, training_input, test_input, class_labels = ad_hoc_data(training_size=20, test_size=10, n=2, gap=0.3, PLOT_DATA=True)
#n=2 is the dimension of each data point
total_array, label_to_labelclass = get_points(test_input, class_labels)
# With the dataset ready we initialize the necessary inputs for the algorithm:
# - the input dictionary (params)
# - the input object containing the dataset info (algo_input).
# +
params = {
'problem': {'name': 'svm_classification'},
'backend': {'name': 'local_qasm_simulator', 'shots': 1000},
'algorithm': {
'name': 'SVM_Variational',
'circuit_depth': 3,
'print_info': True
},
'optimizer': {
'name': 'SPSA',
'save_steps': 10,
'max_trials': 200, #critical!!! ideal: >200
}
}
algo_input = get_input_instance('SVMInput')
algo_input.training_dataset = training_input
algo_input.test_dataset = test_input
algo_input.datapoints = total_array
# -
# With everything setup, we can now run the algorithm.
#
# For the testing, the result includes the details and the success ratio.
#
# For the prediction, the result includes the predicted labels.
result = run_algorithm(params,algo_input)
print(result)
| artificial_intelligence/svm_variational.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Nursery Scenario
# +
# %matplotlib inline
from csrl.mdp import GridMDP
from csrl.oa import OmegaAutomaton
from csrl import ControlSynthesis
import numpy as np
# LTL Specification
ltl = ('G ('
'(!d) & '
'((b & (!(X b)))->(X ((!b) U (a|c)))) & '
'(((!b) & (X b) & (!(X X b)))->((!a) U c)) & '
'(a->(X ((!a) U b))) & '
'(c->((!a) U b)) & '
'((b & (X b))->(F a))'
')')
# Translate the LTL formula to an LDBA
oa = OmegaAutomaton(ltl)
print('Number of Omega-automaton states (including the trap state):',oa.shape[1])
display(oa)
# MDP Description
shape = (5,4)
# E: Empty, T: Trap, B: Obstacle
structure = np.array([
['E', 'E', 'E', 'E'],
['E', 'E', 'E', 'E'],
['E', 'E', 'E', 'E'],
['E', 'E', 'E', 'E'],
['E', 'E', 'E', 'E']
])
# Labels of the states
label = np.array([
[(), (), ('b',),('d',)],
[(), (), (), ()],
[(), (), (), ()],
[('a',),(), (), ()],
[(), ('c',),(), ()]
],dtype=np.object)
# Colors of the labels
lcmap={
('a',):'yellow',
('b',):'greenyellow',
('c',):'turquoise',
('d',):'pink'
}
grid_mdp = GridMDP(shape=shape,structure=structure,label=label,lcmap=lcmap,figsize=5) # Use figsize=4 for smaller figures
grid_mdp.plot()
# Construct the product MDP
csrl = ControlSynthesis(grid_mdp,oa)
# -
Q=csrl.q_learning(start=(4,1),T=1000,K=100000)
value=np.max(Q,axis=4)
csrl.plot(value)
policy=np.argmax(Q,axis=4)
csrl.plot(value,policy)
episode=csrl.simulate(policy,start=(4,1),T=1000,plot=False)
elements, counts = np.unique(np.array(episode)[:,1], return_counts=True)
sorted(zip(elements,counts),key=lambda x:-x[1])
# +
# Plot the important parts of the policy
policy=np.argmax(Q,axis=4)
hidden = [(0,0),(1,0),(2,0),(3,0),(4,0),(4,1),(0,2),(0,3)]
path = {
(4,1) : 'r',
(4,2) : 'lu',
(3,2) : 'du',
(2,2) : 'du',
(1,2) : 'du',
(0,2) : 'd'
}
csrl.plot(value,policy,iq=(0,2),save='nursery_scenario_policy_cb.pdf',path=path,hidden=hidden)
hidden = [(3,0),(4,0),(4,1),(0,2),(0,3)]
path = {
(0,2) : 'l',
(0,1) : 'lr',
(0,0) : 'rd',
(1,0) : 'ur',
(1,1) : 'ld',
(2,1) : 'ur',
(2,2) : 'ld',
(3,2) : 'ud',
(4,2) : 'ul',
(4,1) : 'r'
}
csrl.plot(value,policy,iq=(0,41),save='nursery_scenario_policy_bc.pdf',hidden=hidden,path=path)
hidden=[(3,0),(4,0),(4,1),(4,2),(4,3),(0,2),(0,3)]
path = {
(0,2) : 'l',
(0,1) : 'lr',
(0,0) : 'rd',
(1,0) : 'ud',
(2,0) : 'ud',
(3,0) : 'u'
}
csrl.plot(value,policy,iq=(0,12),save='nursery_scenario_policy_ba.pdf',hidden=hidden,path=path)
hidden=[(3,0),(4,0),(0,2),(0,3),(4,1)]
path = {
(3,0) : 'r',
(3,1) : 'lr',
(3,2) : 'lu',
(2,2) : 'du',
(1,2) : 'du',
(0,2) : 'd'
}
csrl.plot(value,policy,iq=(0,9),save='nursery_scenario_policy_ab.pdf',hidden=hidden,path=path)
# -
# Save the animation
episode=csrl.simulate(policy,start=(4,1),T=200, animation='test')
| Nursery Scenario.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# +
import pandas as pd
country = pd.read_csv("country.csv")
# Extract Country abbreviation and Feature from 'Country_Feature' using regular expressions
sep = country["Country_Feature"].str.extract('(\w{3})(\w{3})')
# Name Columns "Country" and "Feature"
sep.columns = ['Country', 'Feature']
# Merge country and sep
country = pd.concat([sep, country], axis = 1)
# Drop 'Country_Feature' column
country = country.drop(['Country_Feature'], axis = 1)
# Sort Columns to 'Country', 'Feature', and 'Observation'
country = country[['Country', 'Feature', 'Observation']]
# Print country
print(country)
| Extract and Concat.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.1 64-bit (''3.8.1'': pyenv)'
# name: python3
# ---
import converters
import analyzers
import numpy as np
import pandas as pd
import time
import tqdm
from sklearn.metrics import classification_report, accuracy_score, f1_score
small_test_path = 'data/hwu_small_test.csv'
small_train_path = 'data/hwu_small_train.csv'
large_test_path = 'data/hwu_large_test.csv'
large_train_path = 'data/hwu_large_train.csv'
# +
docs, X, y = converters.parse_data(small_train_path)
df = pd.DataFrame({'X': X, 'y': y})
df.to_csv('hwu_watson_small.csv', header=None, columns=['X', 'y'], index=False)
# +
docs, X, y = converters.parse_data(large_train_path)
df = pd.DataFrame({'X': X, 'y': y})
df.to_csv('hwu_watson_large.csv', header=None, columns=['X', 'y'], index=False)
# -
# !pip install --upgrade "ibm-watson>=5.2.2"
import json
from ibm_watson import AssistantV2
from ibm_cloud_sdk_core.authenticators import IAMAuthenticator
def calc_results(test_path):
docs, X, y = converters.parse_data(test_path)
results = []
cant_classify_count = 0
times = []
# get IAMAuthenticator key from Watson web interface
authenticator = IAMAuthenticator('')
assistant = AssistantV2(
version='2021-06-14',
authenticator = authenticator
)
assistant.set_service_url('https://api.eu-de.assistant.watson.cloud.ibm.com')
for sent in tqdm.tqdm(X):
start_time = time.time()
# get assistant_id key from Watson web interface
result = assistant.message_stateless(
assistant_id='',
input={
'message_type': 'text',
'text': sent,
'options': {'alternate_intents': True}
}
).get_result()
times.append(time.time() - start_time)
if not result['output']['intents']:
cant_classify_count += 1
print(sent)
print(result)
break
result = {'output': {'intents': [{'intent': 'Other'}]}}
results.append(result)
return results, times, cant_classify_count
results, times, cant_classify_count = calc_results(large_test_path)
len(results), cant_classify_count
_, _, y = converters.parse_data(small_test_path)
y_pred = []
for r in results:
y_pred.append(r['output']['intents'][0]['intent'])
print(classification_report(y, y_pred))
print("Accuracy: ", accuracy_score(y, y_pred))
print("F1-Score: ", f1_score(y, y_pred, average='macro'))
print(f"Mean response time: {np.mean(times)} +- {np.std(times)} sec.")
_, _, y = converters.parse_data(large_test_path)
y_pred = []
for r in results:
y_pred.append(r['output']['intents'][0]['intent'])
print(classification_report(y, y_pred))
print("Accuracy: ", accuracy_score(y, y_pred))
print("F1-Score: ", f1_score(y, y_pred, average='macro'))
print(f"Mean response time: {np.mean(times)} +- {np.std(times)} sec.")
| watson.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Orchestration of prediction experiments with sktime
#
# * Evaluate the predictive performance one or more strategies on one or more datasets
#
# [Github weblink](https://github.com/alan-turing-institute/sktime/blob/master/examples/experiment_orchestration.ipynb)
# +
from sktime.experiments.orchestrator import Orchestrator
from sktime.experiments.data import DatasetHDD
from sktime.experiments.data import ResultHDD
from sktime.experiments.data import DatasetLoadFromDir
from sktime.experiments.analysis import AnalyseResults
from sktime.experiments.scores import ScoreAccuracy
from sktime.model_selection import PresplitFilesCV
from sktime.highlevel.strategies import TSCStrategy
from sktime.highlevel.tasks import TSCTask
from sktime.classifiers.compose import TimeSeriesForestClassifier
from sklearn.model_selection import KFold
import pandas as pd
import os
# -
# get path to the sktime datasets
import sktime
repodir = os.path.dirname(sktime.__file__)
datadir = os.path.join(repodir, "datasets/data/")
resultsdir = 'results'
# +
# create the task and dataset objects manually for each dataset
dts_ArrowHead = DatasetHDD(dataset_loc=os.path.join(datadir, 'ArrowHead'), dataset_name='ArrowHead')
task_ArrowHead = TSCTask(target='target')
dts_Beef = DatasetHDD(dataset_loc=os.path.join(datadir, 'Beef'), dataset_name='Beef')
task_Beef = TSCTask(target='target')
# +
# or create them automatically
dts_loader = DatasetLoadFromDir(root_dir=datadir)
datasets = dts_loader.load_datasets()
selected_datasets = ['ItalyPowerDemand', 'ArrowHead', 'GunPoint']
datasets = [dataset for dataset in datasets if dataset.dataset_name in selected_datasets]
tasks = [TSCTask(target='target') for _ in range(len(datasets))]
# +
# create strategies
strategies = [TSCStrategy(TimeSeriesForestClassifier(n_estimators=10), name='tsf10'),
TSCStrategy(TimeSeriesForestClassifier(n_estimators=100), name='tsf20')]
# define results output
resultHDD = ResultHDD(results_save_dir=resultsdir,
strategies_save_dir=resultsdir)
# run orchestrator
orchestrator = Orchestrator(datasets=datasets,
tasks=tasks,
strategies=strategies,
cv=PresplitFilesCV(),
result=resultHDD)
orchestrator.run()
# -
# The results list can be obtained from loading the saved csv files by:
results = resultHDD.load()
# Having obtained the list of results objects we can compute the score and run some statistical tests
# +
analyse = AnalyseResults(resultHDD)
strategy_dict, losses_df = analyse.prediction_errors(metric=ScoreAccuracy())
pd.DataFrame(strategy_dict, index=selected_datasets)
# -
# The strategy_dict is used as an argument to the function performing the statistical tests and visualizations. Currently, the following functions are implemented:
# * `analyse.average_and_std_error(strategy_dict)`
# * `analyse.plot_boxcharts(strategy_dict)`
# * `analyse.ranks(strategy_dict)`
# * `analyse.t_test(strategy_dict)`
# * `analyse.sign_test(strategy_dict)`
# * `analyse.ranksum_test(strategy_dict)`
# * `analyse.t_test_with_bonferroni_correction(strategy_dict)`
# * `analyse.wilcoxon_test(strategy_dict)`
# * `analyse.friedman_test(strategy_dict)`
# * `analyse.nemenyi(strategy_dict)`
| examples/experiment_orchestration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.6 64-bit (''anaconda3'': virtualenv)'
# language: python
# name: python37664bitanaconda3virtualenvfb8880f2827448cab7a00dcc60bf2306
# ---
# + [markdown] slideshow={"slide_type": "notes"}
# <font size='6'>Python for Algorithms </font>
#
# ***
# Course: [Udemy](#https://www.udemy.com/python-for-data-structures-algorithms-and-interviews/learn/v4/content)
# ***
# -
# # Dijkstra Algorithm
# One algorithm for finding the shortest path from a starting node to a target node in a weighted graph is Dijkstra’s algorithm. The algorithm creates a tree of shortest paths from the starting vertex, the source, to all other points in the graph. The graph can either be directed or undirected. One stipulation to using the algorithm is that the graph needs to have a nonnegative weight on every edge.
# In Dijkstra's algorithm, the edge has a large weight--the shortest path tree found by the algorithm will try to avoid edges with larger weights.
#
#
# 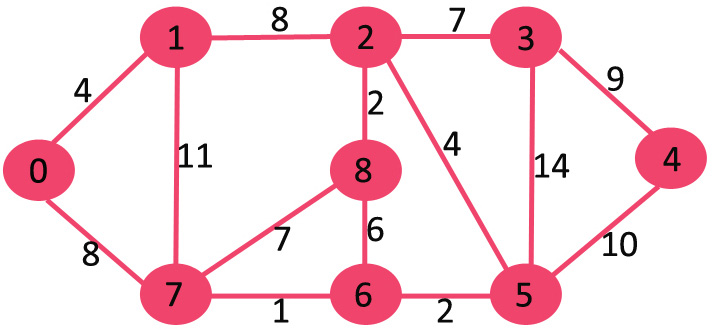
#
# Dijkstra’s algorithm finds a shortest path tree from a single source node, by building a set of nodes that have minimum distance from the source.
#
# The graph has the following:
# * vertices, or nodes, denoted in the algorithm by $v$ or $u$;
# * weighted edges that connect two nodes: $(u,v)$ denotes an edge, and $w(u,v)$ denotes its weight. In the diagram on the right, the weight for each edge is written in gray.
#
# This is done by initializing three values:
# * $dist$, an array of distances from the source node $s$ to each node in the graph, initialized the following way: $dist(s) = 0$; and for all other nodes $v$, $dist(v) = \infty$. This is done at the beginning because as the algorithm proceeds, the $dist$ from the source to each node $v$ in the graph will be recalculated and finalized when the shortest distance to $v$ is found
# * $Q$, a queue of all nodes in the graph. At the end of the algorithm's progress, $Q$ will be empty.
# * $S$, an empty set, to indicate which nodes the algorithm has visited. At the end of the algorithm's run, $S$ will contain all the nodes of the graph.
#
# The algorithm proceeds as follows:
# * While $Q$ is not empty, pop the node $v$, that is not already in $S$, from $Q$ with the smallest $dist(v)$. In the first run, source node $s$ will be chosen because $dist(s)$ was initialized to 0. In the next run, the next node with the smallest $dist$ value is chosen.
# * Add node $v$ to $S$, to indicate that $v$ has been visited
# * Update $dist$ values of adjacent nodes of the current node $v$ as follows: for each new adjacent node $u$,
# * if $dist(v)$ + $weight(u,v)$ < $dist (u)$, there is a new minimal distance found for $u$, so update $dist (u)$ to the new minimal distance value;
# * otherwise, no updates are made to $dist (u)$.
#
# **pseudo code**
#
# ```
# function Dijkstra(Graph, source):
# dist[source] := 0 # Distance from source to source is set to 0
# for each vertex v in Graph: # Initializations
# if v ≠ source
# dist[v] := infinity # Unknown distance function from S to each node set to infinity
# add v to Q # All nodes initially in Q
#
# while Q is not empty: # The main loop
# v := vertex in Q with min dist[v] # In the first run-through, this vertex is the s node
# remove v from Q
#
# for each neighbor u of v: # where neighbor u has not yet been removed from Q.
# alt := dist[v] + length(v, u)
# if alt < dist[u]: # A shorter path to u has been found
# dist[u] := alt # Update distance of u
#
# return dist[]
# end function
#
#
# ```
# +
# =============================================================================
# We will create a dictionary to represent the graph
# =============================================================================
graph = {
'a':{'b':3,'c':4, 'd':7},
'b':{'c':1,'f':5},
'c':{'f':6,'d':2},
'd':{'e':3, 'g':6},
'e':{'g':3, 'h':4},
'f':{'e':1, 'h':8},
'g':{'h':2},
'h':{'g':2}
}
# +
def dijkstra(graph,start,goal):
shortest_distance = {} #dictionary to record the cost to reach to node. We will constantly update this dictionary as we move along the graph.
track_predecessor = {} #dictionary to keep track of path that led to that node.
unseenNodes = graph #to iterate through all nodes
infinity = 5000 #infinity can be considered a very large number
track_path = [] #dictionary to record as we trace back our journey
# =============================================================================
# Initially we want to assign 0 as the cost to reach to source node and infinity as cost to all other nodes
# =============================================================================
for node in unseenNodes:
shortest_distance[node] = infinity
shortest_distance[start] = 0
# =============================================================================
# The loop will keep running until we have entirely exhausted the graph, until we have seen all the nodes
# =============================================================================
# =============================================================================
# To iterate through the graph, we need to determine the min_distance_node every time.
# =============================================================================
while unseenNodes:
min_distance_node = None
for node in unseenNodes:
if min_distance_node is None:
min_distance_node = node
elif shortest_distance[node] < shortest_distance[min_distance_node]:
min_distance_node = node
# =============================================================================
# From the minimum node, what are our possible paths
# =============================================================================
path_options = graph[min_distance_node].items()
# =============================================================================
# We have to calculate the cost each time for each path we take and only update it if it is lower than the existing cost
# =============================================================================
for child_node, weight in path_options:
if weight + shortest_distance[min_distance_node] < shortest_distance[child_node]:
shortest_distance[child_node] = weight + shortest_distance[min_distance_node]
track_predecessor[child_node] = min_distance_node
# =============================================================================
# We want to pop out the nodes that we have just visited so that we dont iterate over them again.
# =============================================================================
unseenNodes.pop(min_distance_node)
# =============================================================================
# Once we have reached the destination node, we want trace back our path and calculate the total accumulated cost.
# =============================================================================
currentNode = goal
while currentNode != start:
try:
track_path.insert(0,currentNode)
currentNode = track_predecessor[currentNode]
except KeyError:
print('Path not reachable')
break
track_path.insert(0,start)
# =============================================================================
# If the cost is infinity, the node had not been reached.
# =============================================================================
if shortest_distance[goal] != infinity:
print('Shortest distance is ' + str(shortest_distance[goal]))
print('And the path is ' + str(track_path))
# -
dijkstra(graph, 'a', 'c')
| Data_Structures_And_Algorithms/notebook/Algorithms_Notes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Predicting a customer's next purchase using automated feature engineering
#
# <p style="margin:30px">
# <img width=50% src="https://www.featuretools.com/wp-content/uploads/2017/12/FeatureLabs-Logo-Tangerine-800.png" alt="Featuretools" />
# </p>
#
# **As customers use your product, they leave behind a trail of behaviors that indicate how they will act in the future. Through automated feature engineering we can identify the predictive patterns in granular customer behavioral data that can be used to improve the customer's experience and generate additional revenue for your business.**
#
# In this tutorial, we show how [Featuretools](https://github.com/alteryx/featuretools) can be used to perform feature engineering on a multi-table dataset of 3 million online grocery orders provided by Instacart. We will generate a feature matrix that can be used to train a machine learning model to predict what product a customer buys next.
#
# *Note: This notebook requires a dataset from Instacart. You can download the dataset [here](https://www.kaggle.com/c/instacart-market-basket-analysis/data). Once you have downloaded the data, be sure to place the CSV files contained in the archive in a directory called `data`. If you use a different directory name, you will need to update the code below to point to the proper location.*
#
# ## Highlights
#
# * We automatically generate features using Deep Feature Synthesis and select the 20 most important features for predictive modeling
# * We demonstrate how to generate features in a scalable manner using [Dask](http://dask.pydata.org/en/latest/)
# * We automatically generate label times using [Compose](https://github.com/alteryx/compose) which can be reused for numerous prediction problems
# * We develop a model for predicting what a customer will buy next, starting with a sample of data and then scaling to the full dataset
# ## You must have Featuretools version 1.4.0 or greater installed to run this notebook
# +
import warnings
warnings.simplefilter("ignore")
import os
import composeml as cp
import featuretools as ft
import dask.dataframe as dd
import numpy as np
import pandas as pd
import utils
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import cross_val_score
from dask.distributed import Client
ft.__version__
# -
# ## Step 1. Load and preprocess data
#
# First, we will create a Dask distributed client so we can track the progress of our computation on the Dask dashboard that is created when the client is initialized.
client = Client(n_workers=2)
client
# Next, we will specify our input and output directories and set the blocksize we will be using to read the raw CSV files into Dask dataframes. When running on a machine with 16GB of memory available for two Dask workers, a `100MB` blocksize has worked well. This number may need to be adjusted based on your specific environment. Refer to the [Dask documentation](https://docs.dask.org/en/latest/best-practices.html) for additional info.
data_dir = os.path.join("data")
output_dir = os.path.join("data", "dask_data")
blocksize = "100MB"
# Now we will read our data into Dask dataframes. This operation will complete quite fast as we are not actually bringing the data into memory at this stage.
order_products = dd.concat([dd.read_csv(os.path.join(data_dir, "order_products__prior.csv"), blocksize=blocksize),
dd.read_csv(os.path.join(data_dir, "order_products__train.csv"), blocksize=blocksize)])
orders = dd.read_csv(os.path.join(data_dir, "orders.csv"), blocksize=blocksize)
departments = dd.read_csv(os.path.join(data_dir, "departments.csv"), blocksize=blocksize)
products = dd.read_csv(os.path.join(data_dir, "products.csv"), blocksize=blocksize)
# In the next few cells, we will perform some required preprocessing to clean up our data. We will merge together some of the raw dataframes and add absolute order time information from the relative times used in the raw data. This will allow us to use cutoff times as part of the Deep Feature Synthesis process.
order_products = order_products.merge(products).merge(departments)
def add_time(df):
df.reset_index(drop=True)
df["order_time"] = np.nan
days_since = df.columns.tolist().index("days_since_prior_order")
hour_of_day = df.columns.tolist().index("order_hour_of_day")
order_time = df.columns.tolist().index("order_time")
df.iloc[0, order_time] = pd.Timestamp('Jan 1, 2015') + pd.Timedelta(df.iloc[0, hour_of_day], "h")
for i in range(1, df.shape[0]):
df.iloc[i, order_time] = df.iloc[i - 1, order_time] \
+ pd.Timedelta(df.iloc[i, days_since], "d") \
+ pd.Timedelta(df.iloc[i, hour_of_day], "h")
to_drop = ["order_number", "order_dow", "order_hour_of_day", "days_since_prior_order", "eval_set"]
df.drop(to_drop, axis=1, inplace=True)
return df
orders = orders.groupby("user_id").apply(add_time, meta={
"order_id": int,
"user_id": int,
"order_time": np.datetime64(0, 'ns').dtype})
order_products = order_products.merge(orders[["order_id", "order_time"]])
order_products["order_product_id"] = order_products["order_id"] * 1000 + order_products["add_to_cart_order"]
order_products = order_products.drop(["product_id", "department_id", "add_to_cart_order"], axis=1)
# Now that the preprocessing work is complete, we will save the results to disk. This will allow us to start from this point in the process in the future, without having to repeat all of the preprocessing steps. If you have already saved the results to disk previously, you can skip the cell below.
#
# #### Note: The process of saving to CSV is computationally intensive and may take 45 minutes or more, depending on the system you are using. You can use the Dask dashboard to monitor the progress.
# +
# Save preprocessed data to disk
# orders.to_csv(os.path.join(output_dir, "orders-*.csv"), index=False)
# order_products.to_csv(os.path.join(output_dir, "order_products-*.csv"), index=False)
# -
# If you have already performed the preprocessing steps and saved the processed files to disk, you can read them in with the commands in the following cell.
# Read preprocessed data from disk
orders = dd.read_csv(os.path.join(output_dir, "orders-*.csv"), blocksize=blocksize)
order_products = dd.read_csv(os.path.join(output_dir, "order_products-*.csv"), blocksize=blocksize)
orders.head()
order_products.head()
# ## Step 2: Create a Featuretools entityset
#
# When using Dask dataframes to create an entityset, type inference can be computationally intensive compared to entitysets created from pandas dataframes because of needing to pull the distributed data into memory. As a best practice, users should specify the [Woodwork logical types](https://woodwork.alteryx.com/en/stable/guides/logical_types_and_semantic_tags.html#Logical-Types) for all of the columns in the dataframes that make up the entityset when using Dask. In the following cell we define the data types for the `order_products` and `orders` dataframes.
# +
order_products_types = {
"order_id": "Categorical",
"reordered": "BooleanNullable",
"product_name": "Categorical",
"aisle_id": "Categorical",
"department": "Categorical",
"order_time": "Datetime",
"order_product_id": "Integer",
}
order_types = {
"order_id": "Integer",
"user_id": "Integer",
"order_time": "Datetime",
}
# -
# Now that we have defined the data types, we can create the entityset and establish the relationship between the two dataframes. For our initial run we will use a sample of the full data to determine what features are the best predictors. Once we have the feature importances established we will rerun on the full dataset using only the most important features.
#
# First we will sample our data - grabbing the orders for 1000 different customers. Because we cannot pass a Dask series to `.isin()` we must call `.compute()` on the ids to convert this into a pandas series.
orders_sample = orders[orders['user_id'].isin(range(1001,2001))]
# Next we will get the order products associated with the orders we sampled.
order_products_sample = order_products[order_products['order_id'].isin(orders_sample['order_id'].compute())]
# Now that we have sampled our data, we can create an entityset from these sampled dataframes.
# +
es = ft.EntitySet("instacart_sample")
es.add_dataframe(dataframe_name="order_products",
dataframe=order_products_sample,
index="order_product_id",
logical_types=order_products_types,
time_index="order_time")
es.add_dataframe(dataframe_name="orders",
dataframe=orders_sample,
index="order_id",
logical_types=order_types,
time_index="order_time")
es.add_relationship("orders", "order_id", "order_products", "order_id")
# -
# Next, we will normalize the `orders` dataframe to create a new `users` dataframe that we will later use as the target dataframe during the deep feature synthesis process.
es.normalize_dataframe(base_dataframe_name="orders", new_dataframe_name="users", index="user_id")
es.plot()
# To finish up creation of the entityset we will add last time indexes and set some interesting values.
es.add_last_time_indexes()
es.add_interesting_values(dataframe_name="order_products",
values={
"department": ['produce', 'dairy eggs', 'snacks', 'beverages', 'frozen', 'pantry', 'bakery', 'canned goods', 'deli', 'dry goods pasta'],
"product_name": ['Banana', 'Bag of Organic Bananas', 'Organic Baby Spinach', 'Organic Strawberries', 'Organic Hass Avocado', 'Organic Avocado', 'Large Lemon', 'Limes', 'Strawberries', 'Organic Whole Milk']
})
# ## Step 3. Use Compose to generate our cutoff times dataframe
#
# In the cells that follow we will demonstrate how [Compose](https://github.com/FeatureLabs/compose) can be used to generate the label times dataframe that will be used as cutoff times for deep feature synthesis.
#
# First we define a labeling function to add a label if a user has bought a specific product or not, and then we will create our `LabelMaker` using this function.
def bought_product(df, product_name):
purchased = df.product_name.str.contains(product_name).any()
return purchased
# Need to convert the `user_id` column type due to an issue in compose
es['users'].ww.set_types(logical_types={'user_id': 'Integer'})
lm = cp.LabelMaker(
target_dataframe_name='user_id',
time_index='order_time',
labeling_function=bought_product,
window_size='4w',
)
def denormalize(es):
df = es['order_products'].merge(es['orders']).merge(es['users'])
return df
# Compose does not currently work on Dask dataframes, so we must first run `.compute()` on the denormalized entityset to switch to pandas.
df = denormalize(es).compute()
# Now we can create our labels, indicating whether or not a user has purchased Bananas.
label_times = lm.search(
df.sort_values('order_time'),
minimum_data='2015-03-15',
num_examples_per_instance=2,
product_name='Banana',
verbose=True,
)
label_times.head()
# We can see above the our training examples contain three pieces of information: a user id, the last time we can use data before feature engineering (called the "cutoff time"), and the label to predict. These are called our "label times".
# We can use `describe` to print out the distribution with the settings and transforms that were used to make these labels. This is useful as a reference for understanding how the labels were generated from raw data. Also, the label distribution is helpful for determining if we have imbalanced labels.
label_times.describe()
# ## Step 4. Run Deep Feature Synthesis
#
# With our label times created, we are ready to run deep feature synthesis to generate our feature matrix. This will execute quickly and the resulting feature matrix will be returned as a Dask dataframe. This process does not cause the feature matrix to be computed or brought into memory.
#
# When we use DFS, we specify
# - `target_dataframe_name` - the table to build features for - `users` in this case
# - `cutoff_time` - the point in time to calculate the features
#
# A good way to think of the `cutoff_time` is that it let's us "pretend" we are at an earlier point in time when generating our features so we can simulate making predictions. We get this time for each customer from the label times we generated above.
#
# For this initial run we will not specify any primitives, which will result in all the default primitives being used to create features.
feature_matrix, features = ft.dfs(target_dataframe_name="users",
cutoff_time=label_times,
entityset=es,
verbose=True)
# Next, let's compute the feature matrix and take a look at what we created.
fm = feature_matrix.ww.compute()
fm.head()
# Before we use this feature matrix to build a predictive model, we will first encode any categorical features using `ft.encode_features()`. Note, at this time `ft.encode_features()` does not work with a Dask feature matrix, so we will use the pandas version we computed above.
#
# Also note that `encode_features` expects a feature matrix with Woodwork initialized as the first input. To retain Woodwork typing information we called compute through the `.ww` Woodwork accessor above rather than calling it directly on the feature matrix dataframe.
# +
fm_encoded, features_encoded = ft.encode_features(fm,
features)
print("Number of features %s" % len(features_encoded))
fm_encoded.head()
# -
# ## Step 5. Machine Learning
#
# Using the default parameters, we generated dozens of potential features for our prediction problem. With a few simple commands, this feature matrix can be used for machine learning
X = fm_encoded.drop(["user_id"], axis=1)
X = X.fillna(0)
y = X.pop("bought_product").astype('bool')
X.head()
y.head()
# Let's train a Random Forest and validate using 3-fold cross validation
# +
clf = RandomForestClassifier(n_estimators=400, n_jobs=-1)
scores = cross_val_score(estimator=clf,X=X, y=y, cv=3,
scoring="roc_auc", verbose=True)
"AUC %.2f +/- %.2f" % (scores.mean(), scores.std())
# -
# As you can see this model predicts the next purchase better than guessing.
#
# Next we will identify the top 20 features so we can use them to later perform machine learning on the whole dataset.
clf.fit(X, y)
top_features = utils.feature_importances(clf, features_encoded, n=20)
# To persist these features, we can save them to disk.
ft.save_features(top_features, os.path.join(data_dir, "top_features.txt"))
# ### Auto Modeling
# Instead of just training a Random Forest, we can use [EvalML](https://evalml.alteryx.com/en/stable/) from Alteryx to build multiple models, compare them, and identify the best one
#
# <p align="center">
# <img width=50% src="https://evalml-web-images.s3.amazonaws.com/evalml_horizontal.svg" alt="Featuretools" />
# </p>
#
import evalml
from evalml import AutoMLSearch
automl = AutoMLSearch(X, y, problem_type="binary", objective="auc")
automl.search()
# With EvalML, we are able to get an AUC of .82. We can view the main features below.
# +
pipeline = automl.best_pipeline
pipeline.fit(X, y)
pipeline.feature_importance[0:10]
# -
# ### Understanding feature engineering in Featuretools
#
# Before moving forward, take a look at the features we created. You will see that they are more than just simple transformations of columns in our raw data. Instead, they perform aggregations (and sometimes stacking of aggregations) across the relationships in our dataset. If you're curious how this works, learn about the Deep Feature Synthesis algorithm in our documentation [here](https://featuretools.alteryx.com/en/stable/).
#
# DFS is so powerful because with no manual work, the library figured out that historical purchases of bananas are important for predicting future purchases. Additionally, it surfaces that purchasing dairy or eggs and reordering behavior are important features.
#
# Even though these features are intuitive, Deep Feature Synthesis will automatically adapt as we change the prediction problem, saving us the time of manually brainstorming and implementing these data transformation.
# ## Step 6. Scale to Full Dataset
#
# Now that we have established the most important features, we will repeat the process of creating a feature matrix, using only these features, and then make predictions using our full dataset.
#
# To start, we will create a new entityset containing our full dataset.
# +
es = ft.EntitySet("instacart_full")
es.add_dataframe(dataframe_name="order_products",
dataframe=order_products,
index="order_product_id",
logical_types=order_products_types,
time_index="order_time")
es.add_dataframe(dataframe_name="orders",
dataframe=orders,
index="order_id",
logical_types=order_types,
time_index="order_time")
es.add_relationship("orders", "order_id", "order_products", "order_id")
# -
es.normalize_dataframe(base_dataframe_name="orders", new_dataframe_name="users", index="user_id")
es.add_last_time_indexes()
es["order_products"]["department"].interesting_values = ['produce', 'dairy eggs', 'snacks', 'beverages', 'frozen', 'pantry', 'bakery', 'canned goods', 'deli', 'dry goods pasta']
es["order_products"]["product_name"].interesting_values = ['Banana', 'Bag of Organic Bananas', 'Organic Baby Spinach', 'Organic Strawberries', 'Organic Hass Avocado', 'Organic Avocado', 'Large Lemon', 'Limes', 'Strawberries', 'Organic Whole Milk']
# We will use our previously defined Compose label maker to create cutoff times for the full dataset.
df = denormalize(es).compute()
label_times = lm.search(
df.sort_values('order_time'),
minimum_data='2015-03-15',
num_examples_per_instance=2,
product_name='Banana',
verbose=True,
)
label_times.head()
# Next we will read in the top 20 features we identified previously and calculate a feature matrix using only these features.
top_features= ft.load_features(os.path.join(data_dir, "top_features.txt"))
top_features
# Having read in the top features we want to use, we can now create our feature matrix on the full dataset with a call to `ft.calculate_feature_matrix()`.
fm = ft.calculate_feature_matrix(top_features, entityset=es, cutoff_time=label_times, verbose=True)
# Next, we will compute our feature matrix to bring the results into memory, allowing us to encode categorical features and make our predictions.
fm = fm.ww.compute()
# +
fm_encoded, features_encoded = ft.encode_features(fm, top_features)
print("Number of features %s" % len(features_encoded))
fm_encoded.head()
# -
X = fm_encoded.drop(["user_id"], axis=1)
X = X.fillna(0)
y = X.pop("bought_product").astype('bool')
# +
clf = RandomForestClassifier(n_estimators=400, n_jobs=-1)
scores = cross_val_score(estimator=clf,X=X, y=y, cv=3,
scoring="roc_auc", verbose=True)
"AUC %.2f +/- %.2f" % (scores.mean(), scores.std())
# -
clf.fit(X, y)
top_features = utils.feature_importances(clf, top_features, n=20)
# Now that we have finished, we can close our Dask client.
client.close()
# ## Next Steps
#
# While this is an end-to-end example of going from raw data to a trained machine learning model, it is necessary to do further exploration before claiming we've built something impactful.
#
# Fortunately, Featuretools makes it easy to build structured data science pipeline. As a next steps, you could
#
# - Further validate these results by creating feature vectors at different cutoff times
# - Perform feature selection on a larger subset of the original data to improve results
# - Define other prediction problems for this dataset (you can even change the dataframe you are making predictions on!)
# - Save feature matrices to disk as CSVs so they can be reused with different problems without recalculating
# - Experiment with parameters to Deep Feature Synthesis
# ## Built at Alteryx Innovation Labs
#
# <p>
# <a href="https://www.alteryx.com/innovation-labs">
# <img width="75%" src="https://evalml-web-images.s3.amazonaws.com/alteryx_innovation_labs.png" alt="Alteryx Innovation Labs" />
# </a>
# </p>
| predict-next-purchase/Tutorial.ipynb |
% ---
% jupyter:
% jupytext:
% text_representation:
% extension: .m
% format_name: light
% format_version: '1.5'
% jupytext_version: 1.14.4
% kernelspec:
% display_name: Matlab
% language: matlab
% name: matlab
% ---
% # EuP system with salts
%
%
% +
EuT=5e-4; PT=1e-4; CT=1e-3; MgT=1e-3; CaT=1e-3; ST=1e-3; pH=7;
pe=21-pH; %the usual setting here is for fully oxic. 21-pH corresponds to the upper stability bound of water. -pH would be the reducing bound
NaT=3*PT+2*ST+2*CT; %sodium to balance charge (Na3PO4 Na2SO4 Na2CO3)
NT=3*EuT+2*MgT+2*CaT; %nitrate to balance charge (Eu(NO3)3 Mg(NO3)2 Ca(NO3)2
flag1=1; flag2=1; % default flag1=2; flag2=1;
%flag1 specifies PHREEQC(3) or Tableau X(1) or Tableau logX(2)
%flag 2 specifies numerical gradients (2) or analytical gradients (1).
%PHREEQC flag2 display outputs if flag2=2. flag2=1 does not
database=['llnl_AgCls_EuPO4s.dat']; acid=['HNO3']; % HCl, HNO3, HClO4, NaOH(!) . KOH as well. watch if elctrolyte is in the mass balance
%all seem to work. %but extreme low pH might cause errors
[Eup3,PO4m3,EuPO4LsR,EuLOHR3LsR,MASSERR]=...
EuPsaltstableau(pH,pe,EuT,PT,CT,MgT,CaT,ST,NT,NaT,flag1,flag2,database,acid)
% + tags=[]
%%file EuPsaltstableau.m
function [Eup3,PO4m3,EuPO4LsR,EuLOHR3LsR,MASSERR]=...
EuPsaltstableau(pH,pe,EuT,PT,CT,MgT,CaT,ST,NO3T,NaT,flag1,flag2,database,acid)
% input tableau. change this part % ----------------------------------------------
pKa1=-log10(7.5e-3); pKa2=-log10(6.2e-8); pKa3=-log10(2.14e-13); khco3=10.3; kh2co3=6.3;
Tableau=[...
%H e Eu PO4 Na NO3- CO3 Mg Ca SO4 logK phase species1
1 0 0 0 0 0 0 0 0 0 0 0 {'H'}
0 1 0 0 0 0 0 0 0 0 0 0 {'e'}
0 0 1 0 0 0 0 0 0 0 0 0 {'Eu+3'}
0 0 0 1 0 0 0 0 0 0 0 0 {'PO4-3'}
0 0 0 0 1 0 0 0 0 0 0 0 {'Na+'}
0 0 0 0 0 1 0 0 0 0 0 0 {'NO3-'}
0 0 0 0 0 0 1 0 0 0 0 0 {'CO3-2'}
0 0 0 0 0 0 0 1 0 0 0 0 {'Mg+2'}
0 0 0 0 0 0 0 0 1 0 0 0 {'Ca+2'}
0 0 0 0 0 0 0 0 0 1 0 0 {'SO4-2'}
0 0 1 1 0 0 0 0 0 0 24 1 {'EuPO4(s)'}
1 0 0 1 0 0 0 0 0 0 pKa3 0 {'HPO4-2'}
2 0 0 1 0 0 0 0 0 0 pKa3+pKa2 0 {'H2PO4-'}
3 0 0 1 0 0 0 0 0 0 pKa3+pKa2+pKa1 0 {'H3PO4'}
-2 0 2 0 0 0 0 0 0 0 -6.9182 0 {'Eu2(OH)2+4'}
-1 0 1 0 0 0 0 0 0 0 -7.9075 0 {'EuOH+2'}
-2 0 1 0 0 0 0 0 0 0 -16.337 0 {'EuO+'}
-3 0 1 0 0 0 0 0 0 0 -15.3481 1 {'Eu(OH)3(s)'}
1 0 0 0 0 0 1 0 0 0 khco3 0 {'HCO3-'}
2 0 0 0 0 0 1 0 0 0 kh2co3 0 {'CO2'}
1 0 1 0 0 0 1 0 0 0 1.6258+khco3 0 {'EuHCO3+2'}
0 0 1 0 0 0 3 0 0 0 -16.8155+(3*khco3) 0 {'Eu(CO3)3-3'}
0 0 1 0 0 0 2 0 0 0 -8.3993+(2*khco3) 0 {'Eu(CO3)2-'}
0 0 1 0 0 0 1 0 0 0 -2.4057+khco3 0 {'EuCO3+'}
-1 0 1 0 0 0 1 0 0 0 -8.4941+khco3 1 {'EuOHCO3(s)'}
-1 0 1 0 0 0 2 0 0 0 -15.176+(2*khco3) 0 {'EuOH(CO3)2-2'}
-2 0 1 0 0 0 1 0 0 0 -17.8462+khco3 0 {'Eu(OH)2CO3-'}
0 0 0 1 0 0 0 1 0 0 -5.7328+(1/pKa3) 0 {'MgPO4-'}
1 0 0 1 0 0 0 1 0 0 2.9100+(1/pKa3) 0 {'MgHPO4'}
2 0 0 1 0 0 0 1 0 0 1.6600+(1/pKa3) 0 {'MgH2PO4+'}
0 0 0 0 0 0 1 1 0 0 -7.3499+khco3 1 {'Magnesite'}
1 0 0 0 0 0 1 1 0 0 1.0357+khco3 0 {'MgHCO3+'}
%-4 0 0 0 0 0 0 4 0 0 -39.75 0 {'Mg(OH)4-2'}
0 0 0 1 0 0 0 0 1 0 -5.8618+(1/pKa3) 0 {'CaPO4-'}
1 0 0 1 0 0 0 0 1 0 2.7400+(1/pKa3) 0 {'CaHPO4'}
2 0 0 1 0 0 0 0 1 0 1.4000+(1/pKa3) 0 {'CaH2PO4+'}
0 0 0 0 0 0 1 0 1 0 -7.0017+khco3 1 {'Calcite'}
1 0 0 0 0 0 1 0 1 0 1.0467+khco3 0 {'CaHCO3+'}
-1 0 0 0 0 0 0 0 1 0 -12.85 0 {'CaOH+'}
1 0 0 0 0 1 0 0 0 0 -1.3025 0 {'HNO3'}
0 0 1 0 0 1 0 0 0 0 0.8745 0 {'EuNO3+2'}
0 0 0 0 0 1 0 0 1 0 0.7000 0 {'CaNO3+'}
1 0 0 0 0 0 0 0 0 1 1.9791 0 {'HSO4-'}
0 0 1 0 0 0 0 0 0 1 3.6430 0 {'EuSO4+'}
0 0 1 0 0 0 0 0 0 2 5.4693 0 {'Eu(SO4)2-'}
0 0 0 0 0 0 0 1 0 1 2.4117 0 {'MgSO4'}
0 0 0 0 0 0 0 0 1 1 4.50 1 {'Gypsum'}
];
T=[EuT; PT; NaT; NO3T; CT; MgT; CaT; ST];
% end of tableau. ------------------ % ----------------------------------------------
[KSOLID,ASOLID,SOLIDNAMES,KSOLUTION,ASOLUTION,SOLUTIONNAMES]=processtableau(Tableau,pH,pe);
[SPECIESCONCS,SPECIATIONNAMES,MASSERR,X]=returnspeciationRE(KSOLID,ASOLID,SOLIDNAMES,KSOLUTION,ASOLUTION,SOLUTIONNAMES,T,flag1,flag2,database,acid);
% this will generate the outputs
for k=1:size(SPECIESCONCS,1)
txt=[SPECIATIONNAMES(k,:),'=SPECIESCONCS(k);'];
eval(txt)
end
end
% + jupyter={"source_hidden": true} tags=[]
%%file processtableau.m
function [KSOLID,ASOLID,SOLIDNAMES,KSOLUTION,ASOLUTION,SOLUTIONNAMES]=processtableau(Tableau,pH,pe)
n=size(Tableau,2);
A=cell2mat(Tableau(:,1:n-3));
K=cell2mat(Tableau(:,n-2));
P=cell2mat(Tableau(:,n-1));
SPECIESNAMES=strvcat(Tableau(:,n));
C1=0; C2=0;
for i=1:size(P,1)
if P(i)==0
C1=C1+1;
ASOLUTION(C1,:)=A(i,:);
KSOLUTION(C1,:)=K(i,:);
SOLUTIONNAMES(C1,:)=SPECIESNAMES(i,:);
end
if P(i)==1
C2=C2+1;
ASOLID(C2,:)=A(i,:);
KSOLID(C2,:)=K(i,:);
SOLIDNAMES(C2,:)=SPECIESNAMES(i,:);
end
end
tstpH=isnan(pH); tstpe=isnan(pe);
if tstpH==0
% adjust for fixed pH
[KSOLUTION,KSOLID,ASOLUTION,ASOLID]=get_equilib_fixed_pH(KSOLUTION,KSOLID,ASOLUTION,ASOLID,pH);
end
if tstpe==0
% adjust for fixed pe
[KSOLUTION,KSOLID,ASOLUTION,ASOLID]=get_equilib_fixed_pe(KSOLUTION,KSOLID,ASOLUTION,ASOLID,pe);
end
end
% + jupyter={"source_hidden": true} tags=[]
%%file returnspeciationRE.m
function [C, SPECIATIONNAMES,MASSERR,X]=returnspeciationRE(KSOLID,ASOLID,SOLIDNAMES,KSOLUTION,ASOLUTION,SOLUTIONNAMES,T,flag1,flag2,database,acid)
% NR on X with either analytical or numerical gradients --------------
if flag1==1
for i=1:size(SOLUTIONNAMES,1)
tst=SOLUTIONNAMES(i,:);
for j=1:length(tst)
if tst(j)=='+'; tst(j)='p'; end
if tst(j)=='-'; tst(j)='m'; end
if tst(j)=='('; tst(j)='L'; end
if tst(j)==')'; tst(j)='R'; end
end
SOLUTIONNAMES(i,:)=tst;
end
for i=1:size(SOLIDNAMES,1)
tst=SOLIDNAMES(i,:);
for j=1:length(tst)
if tst(j)=='('; tst(j)='L'; end
if tst(j)==')'; tst(j)='R'; end
end
SOLIDNAMES(i,:)=tst;
end
[C, SPECIATIONNAMES,MASSERR,X]=NRX(KSOLID,ASOLID,SOLIDNAMES,KSOLUTION,ASOLUTION,SOLUTIONNAMES,T,flag1,flag2);
end
if flag1==2
for i=1:size(SOLUTIONNAMES,1)
tst=SOLUTIONNAMES(i,:);
for j=1:length(tst)
if tst(j)=='+'; tst(j)='p'; end
if tst(j)=='-'; tst(j)='m'; end
if tst(j)=='('; tst(j)='L'; end
if tst(j)==')'; tst(j)='R'; end
end
SOLUTIONNAMES(i,:)=tst;
end
for i=1:size(SOLIDNAMES,1)
tst=SOLIDNAMES(i,:);
for j=1:length(tst)
if tst(j)=='('; tst(j)='L'; end
if tst(j)==')'; tst(j)='R'; end
end
SOLIDNAMES(i,:)=tst;
end
[C, SPECIATIONNAMES,MASSERR,X]=NRlogX(KSOLID,ASOLID,SOLIDNAMES,KSOLUTION,ASOLUTION,SOLUTIONNAMES,T,flag1,flag2);
end
if flag1==3
[C, SPECIATIONNAMES,MASSERR,X]=PHREEQCsolve(KSOLID,ASOLID,SOLIDNAMES,KSOLUTION,ASOLUTION,SOLUTIONNAMES,T,flag1,flag2,database,acid);
end
end
function [C, SPECIATIONNAMES,MASSERR,X]=NRX(KSOLID,ASOLID,SOLIDNAMES,KSOLUTION,ASOLUTION,SOLUTIONNAMES,T,flag1,flag2)
Xguess=T./1e13; Xguess(Xguess==0)=1e-30; TYPX=Xguess;
[Xguess,masserr,J,RSI,C] = nl_massbalancerrnosolid_NR(Xguess,ASOLUTION,KSOLUTION,ASOLID,KSOLID,T,TYPX);
if masserr>0.01
Xguess=Xguess/1000000;
[Xguess,masserr,J,RSI,C] = nl_massbalancerrnosolid_NR(Xguess,ASOLUTION,KSOLUTION,ASOLID,KSOLID,T,TYPX);
end
RSI(RSI>0)=min(T); Xguess=[Xguess; RSI]; TYPX=Xguess;
[X,masserr,J,RSI,C] = nl_massbalancerrsolid_NR(Xguess,ASOLUTION,KSOLUTION,ASOLID,KSOLID,T,TYPX,flag2);
M=size(X,1); N=size(RSI,1); C=[C; X(M-N+1:M)]; C(C<0)=0;
SPECIATIONNAMES=[SOLUTIONNAMES
SOLIDNAMES];
%MASSERR=max(abs((100*(masserr./T))));
MASSERR=(abs((100*(masserr./T))));
end
function [C, SPECIATIONNAMES,MASSERR,X]=NRlogX(KSOLID,ASOLID,SOLIDNAMES,KSOLUTION,ASOLUTION,SOLUTIONNAMES,T,flag1,flag2)
Xguess=T./100; TYPX=Xguess; TYPX=ones(size(T));
%logXguess=log10(T/10000); %Xguess(Xguess==0)=1e-30;
[Xguess,masserr,J,RSI,C] = NRlogXnl_massbalancerrnosolid_NR(Xguess,ASOLUTION,KSOLUTION,ASOLID,KSOLID,T,TYPX,flag2);
%[Xguess,masserr,J,RSI,C] = nl_massbalancerrnosolid_NR(Xguess,ASOLUTION,KSOLUTION,ASOLID,KSOLID,T,TYPX)
% if max(abs(masserr))>0.001
% Xguess=Xguess/1000000;
% [Xguess,masserr,J,RSI,C] = NRlogXnl_massbalancerrnosolid_NR(Xguess,ASOLUTION,KSOLUTION,ASOLID,KSOLID,T,TYPX,flag2)
% end
%RSI(RSI>0)=min(T)/10;
RSI(RSI>0)=0;
XguessN=[(Xguess); RSI]; TYPX=[Xguess; ones(size(RSI))];
[X,masserr,J,RSI,C] = NRlogXnl_massbalancerrsolid_NR(XguessN,ASOLUTION,KSOLUTION,ASOLID,KSOLID,T,TYPX,flag2);
M=length(X); N=length(RSI); C=[C; X(M-N+1:M)]; C(C<0)=0;
SPECIATIONNAMES=[SOLUTIONNAMES
SOLIDNAMES];
%MASSERR=max(abs((100*(masserr./T))));
MASSERR=(abs((100*(masserr./T))));
end
function [C, SPECIATIONNAMES,MASSERR,X]=PHREEQCsolve(KSOLID,ASOLID,SOLIDNAMES,KSOLUTION,ASOLUTION,SOLUTIONNAMES,T,flag1,flag2,database,acid);
%---------------------------------------------------------------------------------
%database=['llnl_nosolubleAgCl.dat'];
%minerals=[{'Chlorargyrite'}]
%minerals=num2cell(SOLIDNAMES,[1 2]);
for i=1:size(SOLIDNAMES,1)
name=SOLIDNAMES(i,:); name(name == ' ') = [];
minerals(i)=num2cell(name,[1 2]);
end
minerals=minerals';
%totalnames=[{'Ag '}; { 'Cl '}; {'Na'}; {'N(5)'}]
%totalvector=[AgT ClT NaT NO3T];
totalvector=T';
%speciesexport=[{'Ag+'}; {'Cl-'};]
c=0;
for i=3:size(SOLUTIONNAMES,1)
c=c+1; name=SOLUTIONNAMES(i,:);
name(name == ' ') = [];
speciesexport(c)=num2cell(name,[1 2]);
end
speciesexport=speciesexport';
pH=-1*KSOLUTION(1); pe=-1*KSOLUTION(2);
[N,M]=size(ASOLUTION); c=0;
for i=3:2+M
c=c+1;
tst=SOLUTIONNAMES(i,:);
% for j=1:length(tst)
% tester=isstrprop(tst(j), 'alpha');
% if tester==0; tst(j)=' '; end
% end
species=['PO4-3']; n=length(species);
if tst(1:n)=='PO4-3'
tst1=['P']; totalnames(c)=num2cell(tst1,[1 2]);
%tst2=['Na']; total2vector(c)=T(c)*2; total2names(c)=num2cell(tst2,[1 2]);
end
species=['Ca+2']; n=length(species);
if tst(1:n)=='Ca+2'
tst1=['Ca']; totalnames(c)=num2cell(tst1,[1 2]);
%tst2=['Na']; total2vector(c)=T(c)*2; total2names(c)=num2cell(tst2,[1 2]);
end
species=['Mg+2']; n=length(species);
if tst(1:n)=='Mg+2'
tst1=['Mg']; totalnames(c)=num2cell(tst1,[1 2]);
end
species=['CO3-2']; n=length(species);
if tst(1:n)=='CO3-2'
tst1=['C(4)']; totalnames(c)=num2cell(tst1,[1 2]);
%tst2=['Na']; total2vector(c)=T(c)*2; total2names(c)=num2cell(tst2,[1 2]);
end
species=['Fe+3']; n=length(species);
if tst(1:n)=='Fe+3'
tst1=['Fe']; totalnames(c)=num2cell(tst1,[1 2]);
%tst2=['Cl']; total2vector(c)=T(c)*3; total2names(c)=num2cell(tst2,[1 2]);
end
species=['Ag+']; n=length(species);
if tst(1:n)=='Ag+'
tst1=['Ag']; totalnames(c)=num2cell(tst1,[1 2]);
%tst2=['N(5)']; total2vector(c)=T(c)*1; total2names(c)=num2cell(tst2,[1 2]);
end
species=['Na+']; n=length(species);
if tst(1:n)=='Na+'
tst1=['Na']; totalnames(c)=num2cell(tst1,[1 2]);
%tst2=['N(5)']; total2vector(c)=T(c)*1; total2names(c)=num2cell(tst2,[1 2]);
end
species=['NO3-']; n=length(species);
if tst(1:n)=='NO3-'
tst1=['N(5)']; totalnames(c)=num2cell(tst1,[1 2]);
end
species=['Cl-']; n=length(species);
if tst(1:n)=='Cl-'
tst1=['Cl']; totalnames(c)=num2cell(tst1,[1 2]);
%tst2=['Na']; total2vector(c)=T(c)*1; total2names(c)=num2cell(tst2,[1 2]);
end
species=['S(-2)']; n=length(species);
if tst(1:n)=='S(-2)'
tst1=['S']; totalnames(c)=num2cell(tst1,[1 2]);
%tst2=['K']; total2vector(c)=T(c)*2; total2names(c)=num2cell(tst2,[1 2]);
end
species=['Eu+3']; n=length(species);
if tst(1:n)=='Eu+3'
tst1=['Eu']; totalnames(c)=num2cell(tst1,[1 2]);
end
species=['SO4-2']; n=length(species);
if tst(1:n)=='SO4-2'
tst1=['S(6)']; totalnames(c)=num2cell(tst1,[1 2]);
end
end
totalnames=[totalnames'];
%total2names'];
totalvector=[totalvector];% total2vector];
temp=25;
%-----------------------------------------------------------------------
[solutionspeciesconcs, solutionspeciesnames, solidconcs, solidnames]= ...
callPHREEQC(totalnames,totalvector,pH,pe,temp,minerals,speciesexport,database,acid,flag2);
%parse the solution into variable for each species requested
for j=1:size(solutionspeciesconcs,1)
name=cell2mat(solutionspeciesnames(j));
%clean up name so it can be a matlab variable
% opens = name == '(m';
% closes = name == 'w)';
% nestingcount = cumsum(opens - [0 closes(1:end-1)]);
% name = name(nestingcount == 0);
name=regexprep(name,'(mol/kgw)','');
name=regexprep(name,'(eq/kgw)','');
name=regexprep(name,'[m_]','');
name = strrep(name,'+','plus');
name = strrep(name,'-','minus');
name=regexprep(name,'(','');
name=regexprep(name,')','');
%assign variable to corresponding concentration
txt=[name,'=solutionspeciesconcs(j);'];
eval(txt)
%pause
end
indx=ones(size(solidconcs));
for i=1:size(solidconcs,1)
name=cell2mat(solidnames(i));
if name(1:2)=='d_' ; indx(i)=0; end
end
c=0;
for i=1:size(solidconcs,1)
name=cell2mat(solidnames(i));
if indx(i)==1
c=c+1;
SOLIDconcs(c)=solidconcs(i);
SOLIDnames(c)=solidnames(i);
end
end
% for j=1:size(SOLIDconcs,1)
% name=cell2mat(solidnames(j));
% %clean up name so it can be a matlab variable
% if name(1:2)=='d_'
% name=regexprep(name,'[d_]','');
% name=regexprep(name,':','');
% name=regexprep(name,'(','');
% name=regexprep(name,')','');
% %assign variable to corresponding concentration
% txt=[name,'=solidconcs(j);']; eval(txt);
% end
% end
% outputs
C=solutionspeciesconcs; C(1)=10^-C(1); C(2)=10^(-C(2));
C=[C
SOLIDconcs'];
X=C(3:2+M);
SUPERA=[ASOLUTION
ASOLID];
[N,M]=size(SUPERA);
Tcalc=zeros(1,M);
for i=1:M
for j=1:N
Tcalc(i)=Tcalc(i)+C(j)*SUPERA(j,i);
end
end
% tst=[T Tcalc']
% C
% pause
relERR=abs(100*((T-Tcalc')./T));
%MASSERR=max(relERR);
MASSERR=(relERR);
for i=1:size(SOLUTIONNAMES,1)
tst=SOLUTIONNAMES(i,:);
for j=1:length(tst)
if tst(j)=='+'; tst(j)='p'; end
if tst(j)=='-'; tst(j)='m'; end
if tst(j)=='('; tst(j)='L'; end
if tst(j)==')'; tst(j)='R'; end
end
SOLUTIONNAMES(i,:)=tst;
end
for i=1:size(SOLIDNAMES,1)
tst=SOLIDNAMES(i,:);
for j=1:length(tst)
if tst(j)=='('; tst(j)='L'; end
if tst(j)==')'; tst(j)='R'; end
end
SOLIDNAMES(i,:)=tst;
end
%SOLIDNAMES
SPECIATIONNAMES=[SOLUTIONNAMES
(SOLIDNAMES)];
end
% + jupyter={"source_hidden": true} tags=[]
%%file NRlogXnl_massbalancerrsolid_NR.m
function [X,F,J,RSI,C] = NRlogXnl_massbalancerrsolid_NR(X,Asolution,Ksolution,Asolid,Ksolid,T,TYPX,flag2)
Nx=size(Asolution,2); Ncp=size(Asolid,1); Nc=size(Asolution,1); %TYPX(1:Nx)=(log10(TYPX(1:Nx)));
TYPX=ones(size(X)); TYPX(1:Nx)=((TYPX(1:Nx)));
%Rtst=residuals(X,Asolution,Ksolution,Asolid,Ksolid,T);
%fun = @(X) residuals(X,Asolution,Ksolution,Asolid,Ksolid,T);
%[jac,err] = jacobianest(fun,X);
%jac
%pause
criteria=1e-16;
%for i=1:100
tst=1; %initialize variable for terminate loop
COUNT=0;
while tst>=criteria
COUNT=COUNT+1;
tester=isinf(X); % for when X has an infinite value. can't have that! so set to half of total mass
if max(tester)==1
for k=1:length(X)
tst=isinf(X(k));
if tst==1; X(k)=0.5*T(k); end
end
disp('inf')
X
end
Xsolution=X(1:Nx); Xsolid=X(Nx+1:Nx+Ncp);
testtwo=Xsolution(Xsolution==0); % for when zero in Xsolution. can't take log of it.
if testtwo==0
for k=1:Nx
if Xsolution(k)==0; Xsolution(k)=1e-10*T(k); end
end
disp('zero')
Xsolution
end
testthree=max(Xsolution./T); % for when mass balance is way crazy!
if testthree>=1e15
for k=1:Nx
if Xsolution(k)/T(k)>=1e15; Xsolution(k)=0.5*T(k); end
end
disp('mass balance')
Xsolution
end
% mass balance with only positive Xsolid values
Xsolidzero=Xsolid;
Xsolidzero(Xsolidzero < 0) = 0;
logC=Ksolution+Asolution*log10(Xsolution); C=10.^(logC);
% calc species
Rmass=Asolution'*C+Asolid'*Xsolidzero-T;
Q=Asolid*log10(Xsolution); SI=(Q+Ksolid);
RSI=SI;
% two versions of RSI
Q=Asolid*log10(Xsolution); SI=(Q+Ksolid);
for i=1:size(Xsolid,1)
if Xsolid(i)>0; RSI(i)=(SI(i)); end % this should be close to zero if solids present
if Xsolid(i)<=0
RSI(i)=(SI(i))-Xsolid(i);
end
end
R=[Rmass; RSI];
tst=sum(abs(R));
if COUNT>=100; tst=0; end % just make sure don't have infinite loop
%if tst<=criteria; disp('should end'); pause; end
if flag2==1
% Evaluate the Jacobian on logX. but first as normal
z=zeros(Nx+Ncp,Nx+Ncp);
for j=1:Nx % mass balance error in terms of solution components
for k=1:Nx
for i=1:Nc; z(j,k)=z(j,k)+Asolution(i,j)*Asolution(i,k)*C(i)/Xsolution(k); end
end
end
% for log ofX derivatives (just on the soluble part). the solution part
% works.
for j=1:Nx
for k=1:Nx
z(j,k)=z(j,k)*(X(k)*2.303);
end
end
%X
%z
for j=1:Nx % mass balance error terms by solid components
for k=Nx+1:Nx+Ncp
z(j,k)=Asolid(k-Nx,j); %derivative of mass balance error by each solid conc (when solid +ve)
if Xsolid(k-Nx)<=0; z(j,k)=0; end %zero when not in mass balance.
end
end
for j=Nx+1:Nx+Ncp % RSI term by solution components (no change for log of X)
for k=1:Nx
%z(j,k)=-1*Asolid(j-Nx,k)*(SI(j-Nx)/Xsolution(k)); % for SI-1 formulation
z(j,k)=((Asolid(j-Nx,k)/(log(10)*Xsolution(k))))*(X(k)*2.303); % for log(SI) formulation
%z(j,k)=Asolid(j-Nx,k)/Xsolution(k); % for ln(SI)-Xsolid formulation
%if Xsolid(k-Nx)<=0; z(j,k)=Asolid(j-Nx,k)/Xsolution(k); end; %same when not in mass balance.
end
end
for j=Nx+1:Nx+Ncp %RSI term by solid components
for k=Nx+1:Nx+Ncp
z(j,k)=0;
if Xsolid(k-Nx)<=0; z(j,k)=0; if j==k; z(j,k)=-1; end; end % logSI-Xsolid when Xsolid is negative. so derivative -1
end
end
end
logX=[log10(Xsolution)
Xsolid];
if flag2==2
F=@(logX) residualslogXwithsolids(logX,Asolution,Ksolution,Asolid,Ksolid,T);
[z,err] = jacobianest(F,logX);
end
F=@(logX) residualslogXwithsolids(logX,Asolution,Ksolution,Asolid,Ksolid,T);
% [ztst,err] = jacobianest(F,logX);
% jac
% z
% ztst
% pause
% logX
%pause
%check each term of jacobian
%one=[z(1,1) jac(1,1)]
%two=[z(1,2) jac(1,2)]
%three=[z(2,1) jac(2,1)]
%four=[z(2,2) jac(2,2)]
%pause
%J=jac;
J=z;
%TYPX=logX;
Jstar = J./TYPX';
%pause % scale Jacobian
%dx_star = -Jstar\(R); % faster than the alternatives it seems. if warnings set to off.
%dx_star =pinv(z)*(-1*R); % no scaling
%dx_star =pinv(Jstar)*(-1*R);
%deltaX=J\(-1*R);
%without scaling ------------
% deltaX = pinv(J)*(-1*R);
% with scaling ---------------
dx_star = pinv(Jstar)*(-1*R);
dx = dx_star.*TYPX; %reverse the scaling
deltaX=dx;
% rescale x
%deltaX=z\(-1*R);
%deltaX = pinv(z)*(-1*R);
%one_over_del=max([1, -1*deltaX'./(0.5*X')]);
%del=1/one_over_del; X=X+del*deltaX;
%Xsolution=X(1:Nx); Xsolid=X(Nx+1:Nx+Ncp);
logX=logX+deltaX;
X=[10.^logX(1:Nx)
logX(Nx+1:Nx+Ncp)];
%for loop=1:length(X(1:Nx)); if X(loop)>=T(loop); X(loop)=T(loop)/2; end; end
%X
%pause
end
% Xsolution=X(1:Nx); Xsolid=X(Nx+1:Nx+Ncp);
% Q=Asolid*log10(Xsolution); SI=10.^(Q+Ksolid);
% RSI=ones(size(SI))-SI;
%
% mass balance with only positive Xsolid values
Xsolidzero=Xsolid;
Xsolidzero(Xsolidzero < 0) = 0;
logC=Ksolution+Asolution*log10(Xsolution); C=10.^(logC); % calc species
Rmass=Asolution'*C+Asolid'*Xsolidzero-T;
Q=Asolid*log10(Xsolution); SI=(Q+Ksolid);
RSI=SI;
% two versions of RSI
Q=Asolid*log10(Xsolution); SI=(Q+Ksolid);
for i=1:size(Xsolid,1)
if Xsolid(i)>0; RSI(i)=(SI(i)); end % this should be zero if solids present
if Xsolid(i)<=0
%RSI(i)=0;
RSI(i)=(SI(i))-Xsolid(i);
%RSI(i)=(SI(i))-Xsolid(i)-2;
end
end
F=[Rmass]; J=z;
end
% + jupyter={"source_hidden": true} tags=[]
%%file NRlogXnl_massbalancerrnosolid_NR.m
function [X,R,J,RSI,C] = NRlogXnl_massbalancerrnosolid_NR(X,Asolution,Ksolution,Asolid,Ksolid,T,TYPX,flag2)
[Nc,Nx]=size(Asolution); %Xsolution=X(1:Nx);
criteria=1e-16;
for II=1:1000
tester=isinf(X);
if max(tester)==1
for k=1:length(X)
tst=isinf(X(k));
if tst==1; X(k)=0.5*T(k); end
end
disp('no solid inf')
%X
end
logC=(Ksolution)+Asolution*log10(X); C=10.^(logC); % calc species
R=Asolution'*C-T ;
if flag2==1
% Evaluate the Jacobian of logX
% first the normal way lilke that CAryou paper
z=zeros(Nx,Nx);
for j=1:Nx
for k=1:Nx
for i=1:Nc; z(j,k)=z(j,k)+Asolution(i,j)*Asolution(i,k)*C(i)/X(k); end
end
end
% then multiply by X beause that is how you do the derivative wrt ln(X)
% and divide by 2.303 to convert to log base 10
for j=1:Nx
for k=1:Nx
z(j,k)=z(j,k)*(X(k)*2.303);
end
end
end
if flag2==2
logX=log10(X);
F=@(logX) residualslogXnosolids(logX,Asolution,Ksolution,Asolid,Ksolid,T);
[z,err] = jacobianest(F,logX);
%tst=max(isnan(z))
end
logX=log10(X);
% F=@(logX) residualslogXnosolids(logX,Asolution,Ksolution,Asolid,Ksolid,T);
% [ztst,err] = jacobianest(F,logX);
% z
% ztst
% pause
%z
%log10(X)
%J=jac;
J=z ;
Jstar = J./TYPX';
%pause % scale Jacobian
%dx_star = -Jstar\(R); % faster than the alternatives it seems. if warnings set to off.
%dx_star =pinv(z)*(-1*R); % no scaling
%dx_star =pinv(Jstar)*(-1*R);
%deltaX=J\(-1*R);
%deltaX = pinv(J)*(-1*R);
%Jstar
dx_star = pinv(Jstar)*(-1*R);
%dx_star=Jstar\(-1*R);
deltaX = dx_star.*TYPX; %reverse the scaling
%deltaX=dx;
% rescale x
%deltaX=z\(-1*R);
%deltaX = pinv(z)*(-1*R);
%one_over_del=max([1, -1*deltaX'./(0.5*X')]);
%del=1/one_over_del; X=X+del*deltaX;
logX=log10(X)+deltaX;
X=10.^logX;
%for loop=1:length(X(1:Nx)); if X(loop)>=T(loop); X(loop)=T(loop)/2; end; end
tst=sum(abs(R));
if tst<=criteria; break; end
end
Q=Asolid*log10(X); SI=(Q+Ksolid);
RSI=ones(size(SI))-SI;
F=[R];
end
% + jupyter={"source_hidden": true} tags=[]
%%file nl_massbalancerrsolid_NR.m
function [X,F,J,RSI,C] = nl_massbalancerrsolid_NR(X,Asolution,Ksolution,Asolid,Ksolid,T,TYPX,flag2)
Nx=size(Asolution,2); Ncp=size(Asolid,1); Nc=size(Asolution,1);
%Rtst=residuals(X,Asolution,Ksolution,Asolid,Ksolid,T);
%fun = @(X) residuals(X,Asolution,Ksolution,Asolid,Ksolid,T);
%[jac,err] = jacobianest(fun,X);
%jac
%pause
criteria=1e-16;
for i=1:100
tester=isinf(X); % for when X has an infinite value. can't have that! so set to half of total mass
if max(tester)==1
for k=1:length(X)
tst=isinf(X(k));
if tst==1; X(k)=0.5*T(k); end
end
disp('inf')
% X
end
Xsolution=X(1:Nx); Xsolid=X(Nx+1:Nx+Ncp);
testtwo=Xsolution(Xsolution==0); % for when zero in Xsolution. can't take log of it.
if testtwo==0
for k=1:Nx
if Xsolution(k)==0; Xsolution(k)=1e-10*T(k); end
end
disp('zero')
Xsolution
end
testthree=max(Xsolution./T); % for when mass balance is way crazy!
if testthree>=1e15
for k=1:Nx
if Xsolution(k)/T(k)>=1e15; Xsolution(k)=0.5*T(k); end
end
disp('mass balance')
Xsolution
end
% mass balance with only positive Xsolid values
Xsolidzero=Xsolid;
Xsolidzero(Xsolidzero < 0) = 0;
logC=Ksolution+Asolution*log10(Xsolution); C=10.^(logC); % calc species
Rmass=Asolution'*C+Asolid'*Xsolidzero-T;
Q=Asolid*log10(Xsolution); SI=(Q+Ksolid);
RSI=SI;
% two versions of RSI
Q=Asolid*log10(Xsolution); SI=(Q+Ksolid);
for i=1:size(Xsolid,1)
if Xsolid(i)>0; RSI(i)=(SI(i)); end % this should be close to zero if solids present
if Xsolid(i)<=0
RSI(i)=(SI(i))-Xsolid(i);
end
end
R=[Rmass; RSI];
if flag2==1
% Evaluate the Jacobian
z=zeros(Nx+Ncp,Nx+Ncp);
for j=1:Nx % mass balance error in terms of solution components
for k=1:Nx
for i=1:Nc; z(j,k)=z(j,k)+Asolution(i,j)*Asolution(i,k)*C(i)/Xsolution(k); end
end
end
for j=1:Nx % mass balance error terms by solid components
for k=Nx+1:Nx+Ncp
z(j,k)=Asolid(k-Nx,j); %derivative of mass balance error by each solid conc (when solid +ve)
if Xsolid(k-Nx)<=0; z(j,k)=0; end %zero when not in mass balance.
end
end
for j=Nx+1:Nx+Ncp % RSI term by solution components
for k=1:Nx
%z(j,k)=-1*Asolid(j-Nx,k)*(SI(j-Nx)/Xsolution(k)); % for SI-1 formulation
z(j,k)=Asolid(j-Nx,k)/(log(10)*Xsolution(k)); % for log(SI) formulation
%z(j,k)=Asolid(j-Nx,k)/Xsolution(k); % for ln(SI)-Xsolid formulation
%if Xsolid(k-Nx)<=0; z(j,k)=Asolid(j-Nx,k)/Xsolution(k); end; %same when not in mass balance.
end
end
for j=Nx+1:Nx+Ncp %RSI term by solid components
for k=Nx+1:Nx+Ncp
z(j,k)=0;
if Xsolid(k-Nx)<=0; z(j,k)=0; if j==k; z(j,k)=-1; end; end % logSI-Xsolid when Xsolid is negative. so derivative -1
end
end
end
if flag2==2
F=@(X) residuals(X,Asolution,Ksolution,Asolid,Ksolid,T);
[z,err] = jacobianest(F,X); z=real(z);
end
J=z;
%TYPX=ones(size(X));
weight = ones(Nx+Ncp,1);
J0 = weight*(1./TYPX'); % Jacobian scaling matrix
Jstar = J./J0;
%pause % scale Jacobian
%dx_star = -Jstar\(R); % might be faster but does not give good answers for solids
%dx_star =pinv(z)*(-1*R);
dx_star =pinv(Jstar)*(-1*R); % seems to give the best answers
%deltaX=z\(-1*R);
%deltaX = pinv(Jstar)*(-1*R);
%dx_star = pinv(Jstar)*(-1*R);
dx = dx_star.*TYPX;
deltaX=dx;
% rescale x
%deltaX = pinv(z)*(-1*R);
Xsolution=X(1:Nx); Xsolid=X(Nx+1:Nx+Ncp);
delXsolution=deltaX(1:Nx); delXsolid=deltaX(Nx+1:Nx+Ncp);
one_over_del=max([1, -1*delXsolution'./(0.5*Xsolution')]);
del=1/one_over_del; %del=1;
Xsolution=Xsolution+del*delXsolution;
Xsolid=Xsolid+delXsolid;
X=[Xsolution; Xsolid];
tst=sum(abs(R));
if tst<=criteria; break; end
end
% Xsolution=X(1:Nx); Xsolid=X(Nx+1:Nx+Ncp);
% Q=Asolid*log10(Xsolution); SI=10.^(Q+Ksolid);
% RSI=ones(size(SI))-SI;
%
% mass balance with only positive Xsolid values
Xsolidzero=Xsolid;
Xsolidzero(Xsolidzero < 0) = 0;
logC=Ksolution+Asolution*log10(Xsolution); C=10.^(logC); % calc species
Rmass=Asolution'*C+Asolid'*Xsolidzero-T;
Q=Asolid*log10(Xsolution); SI=(Q+Ksolid);
RSI=SI;
% two versions of RSI
Q=Asolid*log10(Xsolution); SI=(Q+Ksolid);
for i=1:size(Xsolid,1)
if Xsolid(i)>0; RSI(i)=(SI(i)); end % this should be zero if solids present
if Xsolid(i)<=0
%RSI(i)=0;
RSI(i)=(SI(i))-Xsolid(i);
%RSI(i)=(SI(i))-Xsolid(i)-2;
end
end
F=[Rmass]; J=z;
end
% + jupyter={"source_hidden": true} tags=[]
%%file nl_massbalancerrnosolid_NR.m
function [X,F,J,RSI,C] = nl_massbalancerrnosolid_NR(X,Asolution,Ksolution,Asolid,Ksolid,T,TYPX)
[Nc,Nx]=size(Asolution); %Xsolution=X(1:Nx);
criteria=1e-16;
for i=1:1000
tester=isinf(X);
if max(tester)==1
for k=1:length(X)
tst=isinf(X(k));
if tst==1; X(k)=0.5*T(k); end
end
disp('no solid inf')
%X
end
logC=(Ksolution)+Asolution*log10(X); C=10.^(logC); % calc species
R=Asolution'*C-T;
% Evaluate the Jacobian
z=zeros(Nx,Nx);
for j=1:Nx;
for k=1:Nx;
for i=1:Nc; z(j,k)=z(j,k)+Asolution(i,j)*Asolution(i,k)*C(i)/X(k); end
end
end
%[jac,err] = jacobianest(fun,X);
%J=jac;
J=z;
Jstar = J./TYPX';
%pause % scale Jacobian
dx_star = -Jstar\(R); % faster than the alternatives it seems. if warnings set to off.
%dx_star =pinv(z)*(-1*R); % no scaling
%dx_star =pinv(Jstar)*(-1*R);
%deltaX=z\(-1*R);
%deltaX = pinv(Jstar)*(-1*R);
%dx_star = pinv(Jstar)*(-1*R);
dx = dx_star.*TYPX; %reverse the scaling
deltaX=dx;
% rescale x
%deltaX=z\(-1*R);
%deltaX = pinv(z)*(-1*R);
one_over_del=max([1, -1*deltaX'./(0.5*X')]);
del=1/one_over_del; X=X+del*deltaX;
tst=sum(abs(R));
if tst<=criteria; break; end
end
Q=Asolid*log10(X); SI=(Q+Ksolid);
RSI=ones(size(SI))-SI;
F=[R];
end
% + jupyter={"source_hidden": true} tags=[]
%%file get_equilib_fixed_pH.m
function [Ksolution,Ksolid,Asolution,Asolid]=get_equilib_fixed_pH(KSOLUTION,KSOLID,ASOLUTION,ASOLID,pH)
[N,M]=size(ASOLUTION);
Ksolution=KSOLUTION-ASOLUTION(:,1)*pH;
Asolution=[ASOLUTION(:,2:M)];
[N,M]=size(ASOLID);
Ksolid=KSOLID-ASOLID(:,1)*pH;
Asolid=[ASOLID(:,2:M)];
end
% + jupyter={"source_hidden": true} tags=[]
%%file get_equilib_fixed_pe.m
% ----------- for fixed pe ----------------
function [Ksolution,Ksolid,Asolution,Asolid]=get_equilib_fixed_pe(KSOLUTION,KSOLID,ASOLUTION,ASOLID,pe)
[N,M]=size(ASOLUTION);
Ksolution=KSOLUTION-ASOLUTION(:,1)*pe;
Asolution=[ASOLUTION(:,2:M)];
[N,M]=size(ASOLID);
Ksolid=KSOLID-ASOLID(:,1)*pe;
Asolid=[ASOLID(:,2:M)];
end
% + jupyter={"source_hidden": true} tags=[]
%%file callPHREEQC.m
% return speciation as a function of pH, FeT, PT
function [solutionspeciesconcs, speciesnames, solidconcs, solidnames]=callPHREEQC(totalnames,totalvector,pH,pe,T,minerals,speciesexport,database,acid,flag2)
temp_val=T; pe_val=pe; pH_val=pH; NOOFSOLIDS=size(minerals,1);
% Construct the text file to run PHREEQC from MATLAB-------------------
fileID=fopen('runphreeqc.txt','w');
fclose(fileID);
fileID=fopen('runphreeqc.txt','a');
% add stuff to database
%fprintf(fileID,'PHASES\n');
% add stuff to database
fprintf(fileID,['PHASES \n']);
fprintf(fileID,['pH_Fix\n']);
fprintf(fileID,['H+ = H+\n']);
fprintf(fileID,['-log_k 0.0\n']);
fprintf(fileID,['pe_Fix\n']);
fprintf(fileID,['e- = e-\n']);
fprintf(fileID,['-log_k 0.0\n']);
% define solution -------------------------------------------------------
fprintf(fileID,'SOLUTION \n');
fprintf(fileID,[' pe ' num2str(pe_val), '\n']); % the redox value
fprintf(fileID,[' pH ' num2str(pH_val), '\n']); % the pH condition
fprintf(fileID,[' temp ' num2str(temp_val), '\n']); % the temperature
fprintf(fileID,'-units mol/kgw\n'); % the unit of the input; usually mol/L is used
% put in the totals
for i=1:size(totalnames,1)
totaltxt=[cell2mat(totalnames(i)),' ', num2str(totalvector(i)), ' \n'];
fprintf(fileID,totaltxt);
end
% define equilibrium phases (solids, fix pH, pe) -------------------------
fprintf(fileID,'EQUILIBRIUM_PHASES \n');
for i=1:NOOFSOLIDS
phasestxt=[cell2mat(minerals(i)), ' 0.0 0\n'];
fprintf(fileID,phasestxt);
end
pHfixline=[' pH_Fix -',num2str(pH_val),' ',acid,' 10.0\n'];
fprintf(fileID,pHfixline);
fprintf(fileID,'-force_equality true\n');
pefixline=[' pe_Fix ',num2str(-1*pe_val),' O2\n'];
fprintf(fileID,pefixline);
fprintf(fileID,'-force_equality true\n');
% define outputs of model ------------------------------------------------
fprintf(fileID,'SELECTED_OUTPUT\n');
fprintf(fileID,'-file selected.out\n');
fprintf(fileID,'-selected_out true\n');
fprintf(fileID,'-user_punch true\n');
fprintf(fileID,'-high_precision true\n');
fprintf(fileID,'-reset false\n');
fprintf(fileID,'-simulation false\n');
fprintf(fileID,'-state false\n');
fprintf(fileID,'-distance false\n');
fprintf(fileID,'-time false\n');
fprintf(fileID,'-step false\n');
fprintf(fileID,'-ph true\n');
fprintf(fileID,'-pe true\n');
fprintf(fileID,'-reaction false\n');
fprintf(fileID,'-temperature false\n');
fprintf(fileID,'-alkalinity false\n');
fprintf(fileID,'-ionic_strength false\n');
fprintf(fileID,'-water false\n');
fprintf(fileID,'-charge_balance false\n');
fprintf(fileID,'-percent_error false\n');
%fprintf(fileID,'KNOBS\n');
%fprintf(fileID,'-iterations 1500\n');
% species to export
speciestxt=['-molalities '];
for i=1:size(speciesexport,1)
speciestxt=[speciestxt, cell2mat(speciesexport(i)), ' '];
end
speciestxt=[speciestxt, ' \n'];
fprintf(fileID,speciestxt);
equilibriumphases=['-equilibrium_phases '];
for i=1:NOOFSOLIDS
equilibriumphases=[equilibriumphases, cell2mat(minerals(i)), ' '];
end
fprintf(fileID,equilibriumphases);
fclose(fileID);
% run the model -----------------------------------------------------------
str=['!phreeqc runphreeqc.txt out.txt ', database];
%system('export LD_PRELOAD=/usr/lib/libstdc++.so.6')
%system('phreeqc runphreeqc.txt out.txt llnl.dat')
%!phreeqc runphreeqc.txt out.txt llnl.dat
if flag2==2; eval(str); end % output to the screen
if flag2==1; evalc(str); end % so no screen output
% import the data from the run and prepare to export----------------
fid = fopen('selected.out','rt');
hdr = strtrim(regexp(fgetl(fid),'\t','split')); hdr=hdr';
mat = cell2mat(textscan(fid,repmat('%f',1,numel(hdr))));
fclose(fid);
out_PHREEQC=mat';
[n,m]=size(out_PHREEQC); hdr=hdr(1:n-1); out_PHREEQC=out_PHREEQC(1:n-1,:); n=n-1;
selectedconcs=out_PHREEQC(:,2);
solutionspeciesconcs=selectedconcs(1:n-2*NOOFSOLIDS);
solidconcs=selectedconcs(n-2*NOOFSOLIDS+1:n);
speciesnames=(hdr(1:n-2*NOOFSOLIDS,1));
solidnames=(hdr(n-2*NOOFSOLIDS+1:n,1));
end
% + jupyter={"source_hidden": true} tags=[]
%%file jacobianest.m
function [jac,err] = jacobianest(fun,x0)
% gradest: estimate of the Jacobian matrix of a vector valued function of n variables
% usage: [jac,err] = jacobianest(fun,x0)
%
%
% arguments: (input)
% fun - (vector valued) analytical function to differentiate.
% fun must be a function of the vector or array x0.
%
% x0 - vector location at which to differentiate fun
% If x0 is an nxm array, then fun is assumed to be
% a function of n*m variables.
%
%
% arguments: (output)
% jac - array of first partial derivatives of fun.
% Assuming that x0 is a vector of length p
% and fun returns a vector of length n, then
% jac will be an array of size (n,p)
%
% err - vector of error estimates corresponding to
% each partial derivative in jac.
%
%
% Example: (nonlinear least squares)
% xdata = (0:.1:1)';
% ydata = 1+2*exp(0.75*xdata);
% fun = @(c) ((c(1)+c(2)*exp(c(3)*xdata)) - ydata).^2;
%
% [jac,err] = jacobianest(fun,[1 1 1])
%
% jac =
% -2 -2 0
% -2.1012 -2.3222 -0.23222
% -2.2045 -2.6926 -0.53852
% -2.3096 -3.1176 -0.93528
% -2.4158 -3.6039 -1.4416
% -2.5225 -4.1589 -2.0795
% -2.629 -4.7904 -2.8742
% -2.7343 -5.5063 -3.8544
% -2.8374 -6.3147 -5.0518
% -2.9369 -7.2237 -6.5013
% -3.0314 -8.2403 -8.2403
%
% err =
% 5.0134e-15 5.0134e-15 0
% 5.0134e-15 0 2.8211e-14
% 5.0134e-15 8.6834e-15 1.5804e-14
% 0 7.09e-15 3.8227e-13
% 5.0134e-15 5.0134e-15 7.5201e-15
% 5.0134e-15 1.0027e-14 2.9233e-14
% 5.0134e-15 0 6.0585e-13
% 5.0134e-15 1.0027e-14 7.2673e-13
% 5.0134e-15 1.0027e-14 3.0495e-13
% 5.0134e-15 1.0027e-14 3.1707e-14
% 5.0134e-15 2.0053e-14 1.4013e-12
%
% (At [1 2 0.75], jac should be numerically zero)
%
%
% See also: derivest, gradient, gradest
%
%
% Author: <NAME>
% e-mail: <EMAIL>
% Release: 1.0
% Release date: 3/6/2007
% get the length of x0 for the size of jac
nx = numel(x0);
MaxStep = 100;
StepRatio = 2.0000001;
% was a string supplied?
if ischar(fun)
fun = str2func(fun);
end
% get fun at the center point
f0 = fun(x0);
f0 = f0(:);
n = length(f0);
if n==0
% empty begets empty
jac = zeros(0,nx);
err = jac;
return
end
relativedelta = MaxStep*StepRatio .^(0:-1:-25);
nsteps = length(relativedelta);
% total number of derivatives we will need to take
jac = zeros(n,nx);
err = jac;
for i = 1:nx
x0_i = x0(i);
if x0_i ~= 0
delta = x0_i*relativedelta;
else
delta = relativedelta;
end
% evaluate at each step, centered around x0_i
% difference to give a second order estimate
fdel = zeros(n,nsteps);
for j = 1:nsteps
fdif = fun(swapelement(x0,i,x0_i + delta(j))) - ...
fun(swapelement(x0,i,x0_i - delta(j)));
fdel(:,j) = fdif(:);
end
% these are pure second order estimates of the
% first derivative, for each trial delta.
derest = fdel.*repmat(0.5 ./ delta,n,1);
% The error term on these estimates has a second order
% component, but also some 4th and 6th order terms in it.
% Use Romberg exrapolation to improve the estimates to
% 6th order, as well as to provide the error estimate.
% loop here, as rombextrap coupled with the trimming
% will get complicated otherwise.
for j = 1:n
[der_romb,errest] = rombextrap(StepRatio,derest(j,:),[2 4]);
% trim off 3 estimates at each end of the scale
nest = length(der_romb);
trim = [1:3, nest+(-2:0)];
[der_romb,tags] = sort(der_romb);
der_romb(trim) = [];
tags(trim) = [];
errest = errest(tags);
% now pick the estimate with the lowest predicted error
[err(j,i),ind] = min(errest);
jac(j,i) = der_romb(ind);
end
end
end % mainline function end
% =======================================
% sub-functions
% =======================================
function vec = swapelement(vec,ind,val)
% swaps val as element ind, into the vector vec
vec(ind) = val;
end % sub-function end
% ============================================
% subfunction - romberg extrapolation
% ============================================
function [der_romb,errest] = rombextrap(StepRatio,der_init,rombexpon)
% do romberg extrapolation for each estimate
%
% StepRatio - Ratio decrease in step
% der_init - initial derivative estimates
% rombexpon - higher order terms to cancel using the romberg step
%
% der_romb - derivative estimates returned
% errest - error estimates
% amp - noise amplification factor due to the romberg step
srinv = 1/StepRatio;
% do nothing if no romberg terms
nexpon = length(rombexpon);
rmat = ones(nexpon+2,nexpon+1);
% two romberg terms
rmat(2,2:3) = srinv.^rombexpon;
rmat(3,2:3) = srinv.^(2*rombexpon);
rmat(4,2:3) = srinv.^(3*rombexpon);
% qr factorization used for the extrapolation as well
% as the uncertainty estimates
[qromb,rromb] = qr(rmat,0);
% the noise amplification is further amplified by the Romberg step.
% amp = cond(rromb);
% this does the extrapolation to a zero step size.
ne = length(der_init);
rhs = vec2mat(der_init,nexpon+2,ne - (nexpon+2));
rombcoefs = rromb\(qromb'*rhs);
der_romb = rombcoefs(1,:)';
% uncertainty estimate of derivative prediction
s = sqrt(sum((rhs - rmat*rombcoefs).^2,1));
rinv = rromb\eye(nexpon+1);
cov1 = sum(rinv.^2,2); % 1 spare dof
errest = s'*12.7062047361747*sqrt(cov1(1));
end % rombextrap
% ============================================
% subfunction - vec2mat
% ============================================
function mat = vec2mat(vec,n,m)
% forms the matrix M, such that M(i,j) = vec(i+j-1)
[i,j] = ndgrid(1:n,0:m-1);
ind = i+j;
mat = vec(ind);
if n==1
mat = mat';
end
end % vec2mat
% + jupyter={"source_hidden": true} tags=[]
%%file residuals.m
function R=residuals(X,Asolution,Ksolution,Asolid,Ksolid,T)
Nx=size(Asolution,2); Ncp=size(Asolid,1); Nc=size(Asolution,1);
Xsolution=X(1:Nx); Xsolid=X(Nx+1:Nx+Ncp);
% mass balance with only positive Xsolid values
Xsolidzero=Xsolid;
Xsolidzero(Xsolidzero < 0) = 0;
logC=Ksolution+Asolution*log10(Xsolution); C=10.^(logC); % calc species
Rmass=Asolution'*C+Asolid'*Xsolidzero-T;
% two versions of RSI
Q=Asolid*log10(Xsolution); SI=(Q+Ksolid);
for i=1:size(Xsolid,1)
if Xsolid(i)>0; RSI(i)=(SI(i)); end % this should be zero if solids present
if Xsolid(i)<=0
RSI(i)=(SI(i))-Xsolid(i);
end
end
R=[Rmass; RSI'];
end
% + jupyter={"source_hidden": true} tags=[]
%%file residualslogXnosolids.m
function R=residualslogXnosolids(X,Asolution,Ksolution,Asolid,Ksolid,T)
Nx=size(Asolution,2); Ncp=size(Asolid,1); Nc=size(Asolution,1);
Xsolution=10.^X(1:Nx); %Xsolid=X(Nx+1:Nx+Ncp);
% mass balance with only positive Xsolid values
%Xsolidzero=Xsolid;
%Xsolidzero(Xsolidzero < 0) = 0;
logC=Ksolution+Asolution*log10(Xsolution); C=10.^(logC); % calc species
Rmass=Asolution'*C-T;
% two versions of RSI
% Q=Asolid*log10(Xsolution); SI=(Q+Ksolid);
% for i=1:size(Xsolid,1)
% if Xsolid(i)>0; RSI(i)=(SI(i)); end % this should be zero if solids present
% if Xsolid(i)<=0
% RSI(i)=(SI(i))-Xsolid(i);
% end
% end
R=[Rmass];
end
% + jupyter={"source_hidden": true} tags=[]
%%file residualslogXwithsolids.m
function R=residualslogXwithsolids(X,Asolution,Ksolution,Asolid,Ksolid,T)
Nx=size(Asolution,2); Ncp=size(Asolid,1); Nc=size(Asolution,1);
Xsolution=10.^X(1:Nx); Xsolid=X(Nx+1:Nx+Ncp);
% mass balance with only positive Xsolid values
Xsolidzero=Xsolid;
Xsolidzero(Xsolidzero < 0) = 0;
logC=Ksolution+Asolution*log10(Xsolution); C=10.^(logC); % calc species
Rmass=Asolution'*C+Asolid'*Xsolidzero-T;
% two versions of RSI
Q=Asolid*log10(Xsolution); SI=(Q+Ksolid);
for i=1:size(Xsolid,1)
if Xsolid(i)>0; RSI(i)=(SI(i)); end % this should be zero if solids present
if Xsolid(i)<=0
RSI(i)=(SI(i))-Xsolid(i);
end
end
R=[Rmass; RSI'];
end
% + jupyter={"source_hidden": true} tags=[]
%%file llnl_AgCls_EuPO4s.dat
# $Id: llnl.dat 4023 2010-02-09 21:02:42Z dlpark $
#Data are from 'thermo.com.V8.R6.230' prepared by <NAME> at
#Lawrence Livermore National Laboratory, in Geochemist's Workbench
#format. Converted to Phreeqc format by <NAME> with help from
#<NAME>. A few organic species have been omitted.
#Delta H of reaction calculated from Delta H of formations given in
#thermo.com.V8.R6.230 (8 Mar 2000).
#Note that species have various valid temperature ranges, noted in
#the Range parameter. However, Phreeqc at present makes no use of
#this parameter, so it is the user's responsibility to remain in the
#valid temperature range for all the data used.
#This version is relatively untested. Kindly send comments or
#corrections to <NAME> at <EMAIL>.
LLNL_AQUEOUS_MODEL_PARAMETERS
-temperatures
0.0100 25.0000 60.0000 100.0000
150.0000 200.0000 250.0000 300.0000
#debye huckel a (adh)
-dh_a
0.4939 0.5114 0.5465 0.5995
0.6855 0.7994 0.9593 1.2180
#debye huckel b (bdh)
-dh_b
0.3253 0.3288 0.3346 0.3421
0.3525 0.3639 0.3766 0.3925
-bdot
0.0374 0.0410 0.0438 0.0460
0.0470 0.0470 0.0340 0.0000
#cco2 (coefficients for the Drummond (1981) polynomial)
-co2_coefs
-1.0312 0.0012806
255.9 0.4445
-0.001606
NAMED_EXPRESSIONS
#
# formation of O2 from H2O
# 2H2O = O2 + 4H+ + 4e-
#
Log_K_O2
log_k -85.9951
-delta_H 559.543 kJ/mol # Calculated enthalpy of reaction O2
# Enthalpy of formation: -2.9 kcal/mol
-analytic 38.0229 7.99407E-03 -2.7655e+004 -1.4506e+001 199838.45
# Range: 0-300
SOLUTION_MASTER_SPECIES
#element species alk gfw_formula element_gfw
Acetate HAcetate 0.0 Acetate 59.
Ag Ag+ 0.0 Ag 107.8682
Ag(1) Ag+ 0 Ag
Ag(2) Ag+2 0 Ag
Al Al+3 0.0 Al 26.9815
Alkalinity HCO3- 1.0 Ca0.5(CO3)0.5 50.05
Am Am+3 0.0 Am 243.0000
Am(+2) Am+2 0.0 Am
Am(+3) Am+3 0.0 Am
Am(+4) Am+4 0.0 Am
Am(+5) AmO2+ 0.0 Am
Am(+6) AmO2+2 0.0 Am
Ar Ar 0.0 Ar 39.948
As H2AsO4- 0.0 As 74.9216
As(-3) AsH3 0.0 As
As(+3) H2AsO3- 0.0 As
As(+5) H2AsO4- 0.0 As
Au Au+ 0.0 Au 196.9665
Au(+1) Au+ 0.0 Au
Au(+3) Au+3 0.0 Au
#B H3BO3 0.0 B 10.811
B B(OH)3 0.0 B 10.811
B(3) B(OH)3 0 B
B(-5) BH4- 0 B
Ba Ba+2 0.0 Ba 137.3270
Be Be+2 0.0 Be 9.0122
Br Br- 0.0 Br 79.904
Br(-03) Br3- 0 Br
Br(-1) Br- 0 Br
Br(0) Br2 0 Br
Br(1) BrO- 0 Br
Br(5) BrO3- 0 Br
Br(7) BrO4- 0 Br
C(-4) CH4 0.0 CH4
C(-3) C2H6 0.0 C2H6
C(-2) C2H4 0.0 C2H4
C HCO3- 1.0 HCO3 12.0110
C(+2) CO 0 C
C(+4) HCO3- 1.0 HCO3
Ca Ca+2 0.0 Ca 40.078
Cyanide Cyanide- 1.0 CN 26.
Cd Cd+2 0.0 Cd 112.411
Ce Ce+3 0.0 Ce 140.115
Ce(+2) Ce+2 0.0 Ce
Ce(+3) Ce+3 0.0 Ce
Ce(+4) Ce+4 0.0 Ce
Cl Cl- 0.0 Cl 35.4527
Cl(-1) Cl- 0 Cl
Cl(1) ClO- 0 Cl
Cl(3) ClO2- 0 Cl
Cl(5) ClO3- 0 Cl
Cl(7) ClO4- 0 Cl
Co Co+2 0.0 Co 58.9332
Co(+2) Co+2 0.0 Co
Co(+3) Co+3 0.0 Co
Cr CrO4-2 0.0 CrO4-2 51.9961
Cr(+2) Cr+2 0.0 Cr
Cr(+3) Cr+3 0.0 Cr
Cr(+5) CrO4-3 0.0 Cr
Cr(+6) CrO4-2 0.0 Cr
Cs Cs+ 0.0 Cs 132.9054
Cu Cu+2 0.0 Cu 63.546
Cu(+1) Cu+1 0.0 Cu
Cu(+2) Cu+2 0.0 Cu
Dy Dy+3 0.0 Dy 162.50
Dy(+2) Dy+2 0.0 Dy
Dy(+3) Dy+3 0.0 Dy
E e- 0.0 0.0 0.0
Er Er+3 0.0 Er 167.26
Er(+2) Er+2 0.0 Er
Er(+3) Er+3 0.0 Er
Ethylene Ethylene 0.0 Ethylene 28.0536
Eu Eu+3 0.0 Eu 151.965
Eu(+2) Eu+2 0.0 Eu
Eu(+3) Eu+3 0.0 Eu
F F- 0.0 F 18.9984
Fe Fe+2 0.0 Fe 55.847
Fe(+2) Fe+2 0.0 Fe
Fe(+3) Fe+3 -2.0 Fe
Ga Ga+3 0.0 Ga 69.723
Gd Gd+3 0.0 Gd 157.25
Gd(+2) Gd+2 0.0 Gd
Gd(+3) Gd+3 0.0 Gd
H H+ -1. H 1.0079
H(0) H2 0.0 H
H(+1) H+ -1. 0.0
He He 0.0 He 4.0026
He(0) He 0.0 He
Hf Hf+4 0.0 Hf 178.49
Hg Hg+2 0.0 Hg 200.59
Hg(+1) Hg2+2 0.0 Hg
Hg(+2) Hg+2 0.0 Hg
Ho Ho+3 0.0 Ho 164.9303
Ho(+2) Ho+2 0.0 Ho
Ho(+3) Ho+3 0.0 Ho
I I- 0.0 I 126.9045
I(-03) I3- 0 I
I(-1) I- 0.0 I
I(+1) IO- 0.0 I
I(+5) IO3- 0.0 I
I(+7) IO4- 0.0 I
In In+3 0.0 In 114.82
K K+ 0.0 K 39.0983
Kr Kr 0.0 Kr 83.80
Kr(0) Kr 0.0 Kr
La La+3 0.0 La 138.9055
La(2) La+2 0 La
La(3) La+3 0 La
Li Li+ 0.0 Li 6.9410
Lu Lu+3 0.0 Lu 174.967
Mg Mg+2 0.0 Mg 24.305
Mn Mn+2 0.0 Mn 54.938
Mn(+2) Mn+2 0.0 Mn
Mn(+3) Mn+3 0.0 Mn
Mn(+6) MnO4-2 0 Mn
Mn(+7) MnO4- 0 Mn
Mo MoO4-2 0.0 Mo 95.94
N NH3 1.0 N 14.0067
N(-3) NH3 1.0 N
N(-03) N3- 0.0 N
N(0) N2 0.0 N
N(+3) NO2- 0.0 N
N(+5) NO3- 0.0 N
Na Na+ 0.0 Na 22.9898
Nd Nd+3 0.0 Nd 144.24
Nd(+2) Nd+2 0.0 Nd
Nd(+3) Nd+3 0.0 Nd
Ne Ne 0.0 Ne 20.1797
#Ne(0) Ne 0.0 Ne
Ni Ni+2 0.0 Ni 58.69
Np Np+4 0.0 Np 237.048
Np(+3) Np+3 0.0 Np
Np(+4) Np+4 0.0 Np
Np(+5) NpO2+ 0.0 Np
Np(+6) NpO2+2 0.0 Np
O H2O 0.0 O 15.994
O(-2) H2O 0.0 0.0
O(0) O2 0.0 O
O_phthalate O_phthalate-2 0 1 1
P HPO4-2 2.0 P 30.9738
P(-3) PH4+ 0 P
P(5) HPO4-2 2.0 P
Pb Pb+2 0.0 Pb 207.20
Pb(+2) Pb+2 0.0 Pb
Pb(+4) Pb+4 0.0 Pb
Pd Pd+2 0.0 Pd 106.42
Pm Pm+3 0.0 Pm 147.00
Pm(+2) Pm+2 0.0 Pm
Pm(+3) Pm+3 0.0 Pm
Pr Pr+3 0.0 Pr 140.9076
Pr(+2) Pr+2 0.0 Pr
Pr(+3) Pr+3 0.0 Pr
Pu Pu+4 0.0 Pu 244.00
Pu(+3) Pu+3 0.0 Pu
Pu(+4) Pu+4 0.0 Pu
Pu(+5) PuO2+ 0.0 Pu
Pu(+6) PuO2+2 0.0 Pu
Ra Ra+2 0.0 Ra 226.025
Rb Rb+ 0.0 Rb 85.4678
Re ReO4- 0.0 Re 186.207
Rn Rn 0.0 Rn 222.00
Ru RuO4-2 0.0 Ru 101.07
Ru(+2) Ru+2 0.0 Ru
Ru(+3) Ru+3 0.0 Ru
Ru(+4) Ru(OH)2+2 0.0 Ru
Ru(+6) RuO4-2 0.0 Ru
Ru(+7) RuO4- 0.0 Ru
Ru(+8) RuO4 0.0 Ru
S SO4-2 0.0 SO4 32.066
S(-2) HS- 1.0 S
S(+2) S2O3-2 0.0 S
S(+3) S2O4-2 0.0 S
S(+4) SO3-2 0.0 S
S(+5) S2O5-2 0.0 S
S(+6) SO4-2 0.0 SO4
S(+7) S2O8-2 0.0 S
S(+8) HSO5- 0.0 S
Sb Sb(OH)3 0.0 Sb 121.75
Sc Sc+3 0.0 Sc 44.9559
Se SeO3-2 0.0 Se 78.96
Se(-2) HSe- 0.0 Se
Se(+4) SeO3-2 0.0 Se
Se(+6) SeO4-2 0.0 Se
Si SiO2 0.0 SiO2 28.0855
Sm Sm+3 0.0 Sm 150.36
Sm(+2) Sm+2 0.0 Sm
Sm(+3) Sm+3 0.0 Sm
Sn Sn+2 0.0 Sn 118.71
Sn(+2) Sn+2 0.0 Sn
Sn(+4) Sn+4 0.0 Sn
Sr Sr+2 0.0 Sr 87.62
Tb Tb+3 0.0 Tb 158.9253
Tb(+2) Tb+2 0.0 Tb
Tb(+3) Tb+3 0.0 Tb
Tc TcO4- 0.0 Tc 98.00
Tc(+3) Tc+3 0.0 Tc
Tc(+4) TcO+2 0.0 Tc
Tc(+5) TcO4-3 0.0 Tc
Tc(+6) TcO4-2 0.0 Tc
Tc(+7) TcO4- 0.0 Tc
Thiocyanate Thiocyanate- 0.0 SCN 58.
Th Th+4 0.0 Th 232.0381
Ti Ti(OH)4 0.0 Ti 47.88
Tl Tl+ 0.0 Tl 204.3833
Tl(+1) Tl+ 0.0 Tl
Tl(+3) Tl+3 0.0 Tl
Tm Tm+3 0.0 Tm 168.9342
Tm(+2) Tm+2 0.0 Tm
Tm(+3) Tm+3 0.0 Tm
U UO2+2 0.0 U 238.0289
U(+3) U+3 0.0 U
U(+4) U+4 0.0 U
U(+5) UO2+ 0.0 U
U(+6) UO2+2 0.0 U
V VO+2 0.0 V 50.9415
V(+3) V+3 0.0 V
V(+4) VO+2 0.0 V
V(+5) VO2+ 0.0 V
W WO4-2 0.0 W 183.85
Xe Xe 0.0 Xe 131.29
Xe(0) Xe 0.0 Xe
Y Y+3 0.0 Y 88.9059
Yb Yb+3 0.0 Yb 173.04
Yb(+2) Yb+2 0.0 Yb
Yb(+3) Yb+3 0.0 Yb
Zn Zn+2 0.0 Zn 65.39
Zr Zr(OH)2+2 0.0 Zr 91.224
SOLUTION_SPECIES
HAcetate = HAcetate
-llnl_gamma 3.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction HAcetate
# Enthalpy of formation: -116.1 kcal/mol
Ag+ = Ag+
-llnl_gamma 2.5000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Ag+
# Enthalpy of formation: 25.275 kcal/mol
Al+3 = Al+3
-llnl_gamma 9.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Al+3
# Enthalpy of formation: -128.681 kcal/mol
Am+3 = Am+3
-llnl_gamma 5.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Am+3
# Enthalpy of formation: -616.7 kJ/mol
Ar = Ar
-llnl_gamma 3.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Ar
# Enthalpy of formation: -2.87 kcal/mol
Au+ = Au+
-llnl_gamma 4.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Au+
# Enthalpy of formation: 47.58 kcal/mol
B(OH)3 = B(OH)3
-llnl_gamma 3.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction B(OH)3
# Enthalpy of formation: -256.82 kcal/mol
Ba+2 = Ba+2
-llnl_gamma 5.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Ba+2
# Enthalpy of formation: -128.5 kcal/mol
Be+2 = Be+2
-llnl_gamma 8.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Be+2
# Enthalpy of formation: -91.5 kcal/mol
Br- = Br-
-llnl_gamma 3.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Br-
# Enthalpy of formation: -29.04 kcal/mol
Ca+2 = Ca+2
-llnl_gamma 6.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Ca+2
# Enthalpy of formation: -129.8 kcal/mol
Cd+2 = Cd+2
-llnl_gamma 5.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Cd+2
# Enthalpy of formation: -18.14 kcal/mol
Ce+3 = Ce+3
-llnl_gamma 9.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Ce+3
# Enthalpy of formation: -167.4 kcal/mol
Cl- = Cl-
-llnl_gamma 3.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Cl-
# Enthalpy of formation: -39.933 kcal/mol
Co+2 = Co+2
-llnl_gamma 6.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Co+2
# Enthalpy of formation: -13.9 kcal/mol
CrO4-2 = CrO4-2
-llnl_gamma 4.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction CrO4-2
# Enthalpy of formation: -210.6 kcal/mol
Cs+ = Cs+
-llnl_gamma 2.5000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Cs+
# Enthalpy of formation: -61.67 kcal/mol
Cu+2 = Cu+2
-llnl_gamma 6.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Cu+2
# Enthalpy of formation: 15.7 kcal/mol
Dy+3 = Dy+3
-llnl_gamma 5.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Dy+3
# Enthalpy of formation: -166.5 kcal/mol
e- = e-
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction e-
# Enthalpy of formation: -0 kJ/mol
Er+3 = Er+3
-llnl_gamma 5.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Er+3
# Enthalpy of formation: -168.5 kcal/mol
Ethylene = Ethylene
-llnl_gamma 3.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Ethylene
# Enthalpy of formation: 8.57 kcal/mol
Eu+3 = Eu+3
-llnl_gamma 5.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Eu+3
# Enthalpy of formation: -144.7 kcal/mol
F- = F-
-llnl_gamma 3.5000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction F-
# Enthalpy of formation: -80.15 kcal/mol
Fe+2 = Fe+2
-llnl_gamma 6.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Fe+2
# Enthalpy of formation: -22.05 kcal/mol
Ga+3 = Ga+3
-llnl_gamma 5.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Ga+3
# Enthalpy of formation: -50.6 kcal/mol
Gd+3 = Gd+3
-llnl_gamma 5.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Gd+3
# Enthalpy of formation: -164.2 kcal/mol
H+ = H+
-llnl_gamma 9.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction H+
# Enthalpy of formation: -0 kJ/mol
He = He
-llnl_gamma 3.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction He
# Enthalpy of formation: -0.15 kcal/mol
H2AsO4- = H2AsO4-
-llnl_gamma 4.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction H2AsO4-
# Enthalpy of formation: -217.39 kcal/mol
HCO3- = HCO3-
-llnl_gamma 4.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction HCO3-
# Enthalpy of formation: -164.898 kcal/mol
HPO4-2 = HPO4-2
-llnl_gamma 4.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction HPO4-2
# Enthalpy of formation: -308.815 kcal/mol
Hf+4 = Hf+4
log_k 0
-delta_H 0 # Not possible to calculate enthalpy of reaction Hf+4
# Enthalpy of formation: -0 kcal/mol
Hg+2 = Hg+2
-llnl_gamma 5.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Hg+2
# Enthalpy of formation: 40.67 kcal/mol
Ho+3 = Ho+3
-llnl_gamma 5.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Ho+3
# Enthalpy of formation: -169 kcal/mol
I- = I-
-llnl_gamma 3.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction I-
# Enthalpy of formation: -13.6 kcal/mol
In+3 = In+3
-llnl_gamma 9.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction In+3
# Enthalpy of formation: -25 kcal/mol
K+ = K+
-llnl_gamma 3.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction K+
# Enthalpy of formation: -60.27 kcal/mol
Kr = Kr
-llnl_gamma 3.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Kr
# Enthalpy of formation: -3.65 kcal/mol
La+3 = La+3
-llnl_gamma 9.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction La+3
# Enthalpy of formation: -169.6 kcal/mol
Li+ = Li+
-llnl_gamma 6.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Li+
# Enthalpy of formation: -66.552 kcal/mol
Lu+3 = Lu+3
-llnl_gamma 5.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Lu+3
# Enthalpy of formation: -167.9 kcal/mol
Mg+2 = Mg+2
-llnl_gamma 8.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Mg+2
# Enthalpy of formation: -111.367 kcal/mol
Mn+2 = Mn+2
-llnl_gamma 6.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Mn+2
# Enthalpy of formation: -52.724 kcal/mol
MoO4-2 = MoO4-2
-llnl_gamma 4.5000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction MoO4-2
# Enthalpy of formation: -238.5 kcal/mol
NH3 = NH3
-llnl_gamma 3.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction NH3
# Enthalpy of formation: -19.44 kcal/mol
Na+ = Na+
-llnl_gamma 4.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Na+
# Enthalpy of formation: -57.433 kcal/mol
Nd+3 = Nd+3
-llnl_gamma 9.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Nd+3
# Enthalpy of formation: -166.5 kcal/mol
Ne = Ne
-llnl_gamma 3.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Ne
# Enthalpy of formation: -0.87 kcal/mol
Ni+2 = Ni+2
-llnl_gamma 6.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Ni+2
# Enthalpy of formation: -12.9 kcal/mol
Np+4 = Np+4
-llnl_gamma 5.5000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Np+4
# Enthalpy of formation: -556.001 kJ/mol
H2O = H2O
-llnl_gamma 3.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction H2O
# Enthalpy of formation: -68.317 kcal/mol
O_phthalate-2 = O_phthalate-2
-llnl_gamma 4.0000
log_k 0
-delta_H 0 # Not possible to calculate enthalpy of reaction O_phthalate-2
# Enthalpy of formation: -0 kcal/mol
Pb+2 = Pb+2
-llnl_gamma 4.5000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Pb+2
# Enthalpy of formation: 0.22 kcal/mol
Pd+2 = Pd+2
-llnl_gamma 4.5000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Pd+2
# Enthalpy of formation: 42.08 kcal/mol
Pm+3 = Pm+3
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Pm+3
# Enthalpy of formation: -688 kJ/mol
Pr+3 = Pr+3
-llnl_gamma 9.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Pr+3
# Enthalpy of formation: -168.8 kcal/mol
Pu+4 = Pu+4
-llnl_gamma 5.5000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Pu+4
# Enthalpy of formation: -535.893 kJ/mol
Ra+2 = Ra+2
-llnl_gamma 5.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Ra+2
# Enthalpy of formation: -126.1 kcal/mol
Rb+ = Rb+
-llnl_gamma 2.5000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Rb+
# Enthalpy of formation: -60.02 kcal/mol
ReO4- = ReO4-
-llnl_gamma 4.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction ReO4-
# Enthalpy of formation: -188.2 kcal/mol
Rn = Rn
-llnl_gamma 3.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Rn
# Enthalpy of formation: -5 kcal/mol
RuO4-2 = RuO4-2
-llnl_gamma 4.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction RuO4-2
# Enthalpy of formation: -457.075 kJ/mol
SO4-2 = SO4-2
-llnl_gamma 4.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction SO4-2
# Enthalpy of formation: -217.4 kcal/mol
Sb(OH)3 = Sb(OH)3
-llnl_gamma 3.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Sb(OH)3
# Enthalpy of formation: -773.789 kJ/mol
Sc+3 = Sc+3
-llnl_gamma 9.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Sc+3
# Enthalpy of formation: -146.8 kcal/mol
SeO3-2 = SeO3-2
-llnl_gamma 4.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction SeO3-2
# Enthalpy of formation: -121.7 kcal/mol
SiO2 = SiO2
-llnl_gamma 3.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction SiO2
# Enthalpy of formation: -209.775 kcal/mol
Sm+3 = Sm+3
-llnl_gamma 9.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Sm+3
# Enthalpy of formation: -165.2 kcal/mol
Sn+2 = Sn+2
-llnl_gamma 6.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Sn+2
# Enthalpy of formation: -2.1 kcal/mol
Sr+2 = Sr+2
-llnl_gamma 5.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Sr+2
# Enthalpy of formation: -131.67 kcal/mol
Tb+3 = Tb+3
-llnl_gamma 5.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Tb+3
# Enthalpy of formation: -166.9 kcal/mol
TcO4- = TcO4-
-llnl_gamma 4.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction TcO4-
# Enthalpy of formation: -716.269 kJ/mol
Th+4 = Th+4
-llnl_gamma 11.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Th+4
# Enthalpy of formation: -183.8 kcal/mol
Ti(OH)4 = Ti(OH)4
-llnl_gamma 3.0000
log_k 0
-delta_H 0 # Not possible to calculate enthalpy of reaction Ti(OH)4
# Enthalpy of formation: -0 kcal/mol
Tl+ = Tl+
-llnl_gamma 2.5000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Tl+
# Enthalpy of formation: 1.28 kcal/mol
Tm+3 = Tm+3
-llnl_gamma 5.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Tm+3
# Enthalpy of formation: -168.5 kcal/mol
UO2+2 = UO2+2
-llnl_gamma 4.5000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction UO2+2
# Enthalpy of formation: -1019 kJ/mol
VO+2 = VO+2
-llnl_gamma 4.5000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction VO+2
# Enthalpy of formation: -116.3 kcal/mol
WO4-2 = WO4-2
-llnl_gamma 5.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction WO4-2
# Enthalpy of formation: -257.1 kcal/mol
Xe = Xe
-llnl_gamma 3.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Xe
# Enthalpy of formation: -4.51 kcal/mol
Y+3 = Y+3
-llnl_gamma 9.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Y+3
# Enthalpy of formation: -170.9 kcal/mol
Yb+3 = Yb+3
-llnl_gamma 5.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Yb+3
# Enthalpy of formation: -160.3 kcal/mol
Zn+2 = Zn+2
-llnl_gamma 6.0000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Zn+2
# Enthalpy of formation: -36.66 kcal/mol
Zr(OH)2+2 = Zr(OH)2+2
-llnl_gamma 4.5000
log_k 0
-delta_H 0 kJ/mol # Calculated enthalpy of reaction Zr(OH)2+2
# Enthalpy of formation: -260.717 kcal/mol
2H2O = O2 + 4H+ + 4e-
-CO2_llnl_gamma
log_k -85.9951
-delta_H 559.543 kJ/mol # Calculated enthalpy of reaction O2
# Enthalpy of formation: -2.9 kcal/mol
-analytic 38.0229 7.99407E-03 -2.7655e+004 -1.4506e+001 199838.45
# Range: 0-300
1.0000 SO4-- + 1.0000 H+ = HS- +2.0000 O2
-llnl_gamma 3.5
log_k -138.3169
-delta_H 869.226 kJ/mol # Calculated enthalpy of reaction HS-
# Enthalpy of formation: -3.85 kcal/mol
-analytic 2.6251e+001 3.9525e-002 -4.5443e+004 -1.1107e+001 3.1843e+005
# -Range: 0-300
.5000 O2 + 2.0000 HS- = S2-- + H2O
#2 HS- = S2-- +2 H+ + 2e-
-llnl_gamma 4.0
log_k 33.2673
-delta_H 0 # Not possible to calculate enthalpy of reaction S2-2
# Enthalpy of formation: -0 kcal/mol
-analytic 0.21730E+02 -0.12307E-02 0.10098E+05 -0.88813E+01 0.15757E+03
-mass_balance S(-2)2
# -Range: 0-300
# -add_logk Log_K_O2 0.5
2.0000 H+ + 2.0000 SO3-- = S2O3-- + O2 + H2O
-llnl_gamma 4.0
log_k -40.2906
-delta_H 0 # Not possible to calculate enthalpy of reaction S2O3-2
# Enthalpy of formation: -0 kcal/mol
-analytic 0.77679E+02 0.65761E-01 -0.15438E+05 -0.34651E+02 -0.24092E+03
# -Range: 0-300
1.0000 H+ + 1.0000 Ag+ + 0.2500 O2 = Ag++ +0.5000 H2O
-llnl_gamma 4.5
log_k -12.1244
-delta_H 22.9764 kJ/mol # Calculated enthalpy of reaction Ag+2
# Enthalpy of formation: 64.2 kcal/mol
-analytic -4.7312e+001 -1.5239e-002 -4.1954e+002 1.6622e+001 -6.5328e+000
# -Range: 0-300
1.0000 Am+++ + 0.5000 H2O = Am++ +1.0000 H+ +0.2500 O2
-llnl_gamma 4.5
log_k -60.3792
-delta_H 401.953 kJ/mol # Calculated enthalpy of reaction Am+2
# Enthalpy of formation: -354.633 kJ/mol
-analytic 1.4922e+001 3.5993e-003 -2.0987e+004 -2.4146e+000 -3.2749e+002
# -Range: 0-300
1.0000 H+ + 1.0000 Am+++ + 0.2500 O2 = Am++++ +0.5000 H2O
-llnl_gamma 5.5
log_k -22.7073
-delta_H 70.8142 kJ/mol # Calculated enthalpy of reaction Am+4
# Enthalpy of formation: -406 kJ/mol
-analytic -1.7460e+001 -2.2336e-003 -3.5139e+003 2.9102e+000 -5.4826e+001
# -Range: 0-300
1.0000 H2O + 1.0000 Am+++ + 0.5000 O2 = AmO2+ +2.0000 H+
-llnl_gamma 4.0
log_k -15.384
-delta_H 104.345 kJ/mol # Calculated enthalpy of reaction AmO2+
# Enthalpy of formation: -804.26 kJ/mol
-analytic 1.4110e+001 6.9728e-003 -4.2098e+003 -6.0936e+000 -2.1192e+005
# -Range: 0-300
1.0000 Am+++ + 0.7500 O2 + 0.5000 H2O = AmO2++ +1.0000 H+
-llnl_gamma 4.5
log_k -20.862
-delta_H 117.959 kJ/mol # Calculated enthalpy of reaction AmO2+2
# Enthalpy of formation: -650.76 kJ/mol
-analytic 5.7163e+001 4.0278e-003 -8.4633e+003 -2.0550e+001 -1.3208e+002
# -Range: 0-300
1.0000 H2AsO4- + 1.0000 H+ = AsH3 +2.0000 O2
-llnl_gamma 3.0
log_k -155.1907
-delta_H 931.183 kJ/mol # Calculated enthalpy of reaction AsH3
# Enthalpy of formation: 10.968 kcal/mol
-analytic 2.8310e+002 9.6961e-002 -5.4830e+004 -1.1449e+002 -9.3119e+002
# -Range: 0-200
2.0000 H+ + 1.0000 Au+ + 0.5000 O2 = Au+++ +1.0000 H2O
-llnl_gamma 5.0
log_k -4.3506
-delta_H -73.2911 kJ/mol # Calculated enthalpy of reaction Au+3
# Enthalpy of formation: 96.93 kcal/mol
-analytic -6.8661e+001 -2.6838e-002 4.4549e+003 2.3178e+001 6.9534e+001
# -Range: 0-300
1.0000 H2O + 1.0000 B(OH)3 = BH4- +2.0000 O2 +1.0000 H+
-llnl_gamma 4.0
log_k -237.1028
-delta_H 1384.24 kJ/mol # Calculated enthalpy of reaction BH4-
# Enthalpy of formation: 48.131 kJ/mol
-analytic -7.4930e+001 -7.2794e-003 -6.9168e+004 2.9105e+001 -1.0793e+003
# -Range: 0-300
3.0000 Br- + 2.0000 H+ + 0.5000 O2 = Br3- +1.0000 H2O
-llnl_gamma 4.0
log_k +7.0696
-delta_H -45.6767 kJ/mol # Calculated enthalpy of reaction Br3-
# Enthalpy of formation: -31.17 kcal/mol
-analytic 1.4899e+002 6.4017e-002 -3.3831e+002 -6.4596e+001 -5.3232e+000
# -Range: 0-300
1.0000 Br- + 0.5000 O2 = BrO-
-llnl_gamma 4.0
log_k -10.9167
-delta_H 33.4302 kJ/mol # Calculated enthalpy of reaction BrO-
# Enthalpy of formation: -22.5 kcal/mol
-analytic 5.4335e+001 1.9509e-003 -4.2860e+003 -2.0799e+001 -6.6896e+001
# -Range: 0-300
1.5000 O2 + 1.0000 Br- = BrO3-
-llnl_gamma 3.5
log_k -17.1443
-delta_H 72.6342 kJ/mol # Calculated enthalpy of reaction BrO3-
# Enthalpy of formation: -16.03 kcal/mol
-analytic 3.7156e+001 -4.7855e-003 -4.6208e+003 -1.4136e+001 -2.1385e+005
# -Range: 0-300
2.0000 O2 + 1.0000 Br- = BrO4-
-llnl_gamma 4.0
log_k -33.104
-delta_H 158.741 kJ/mol # Calculated enthalpy of reaction BrO4-
# Enthalpy of formation: 3.1 kcal/mol
-analytic 8.1393e+001 -2.3409e-003 -1.2290e+004 -2.9336e+001 -1.9180e+002
# -Range: 0-300
# 1.0000 NH3 + 1.0000 HCO3- = CN- +2.0000 H2O +0.5000 O2
# -llnl_gamma 3.0
# log_k -56.0505
# -delta_H 344.151 kJ/mol # Calculated enthalpy of reaction CN-
# # Enthalpy of formation: 36 kcal/mol
# -analytic -1.1174e+001 3.8167e-003 -1.7063e+004 4.5349e+000 -2.6625e+002
# # -Range: 0-300
Cyanide- = Cyanide-
log_k 0
H+ + HCO3- + H2O = CH4 + 2.0000 O2
-llnl_gamma 3.0
log_k -144.1412
-delta_H 863.599 kJ/mol # Calculated enthalpy of reaction CH4
# Enthalpy of formation: -21.01 kcal/mol
-analytic -0.41698E+02 0.36584E-01 -0.40675E+05 0.93479E+01 -0.63468E+03
# -Range: 0-300
2.0000 H+ + 2.0000 HCO3- + H2O = C2H6 + 3.5000 O2
-llnl_gamma 3.0
log_k -228.6072
-delta_H 0 # Not possible to calculate enthalpy of reaction C2H6
# Enthalpy of formation: -0 kcal/mol
-analytic 0.10777E+02 0.72105E-01 -0.67489E+05 -0.13915E+02 -0.10531E+04
# -Range: 0-300
2.000 H+ + 2.0000 HCO3- = C2H4 + 3.0000 O2
-llnl_gamma 3.0
log_k -254.5034
-delta_H 1446.6 kJ/mol # Calculated enthalpy of reaction C2H4
# Enthalpy of formation: 24.65 kcal/mol
-analytic -0.30329E+02 0.71187E-01 -0.73140E+05 0.00000E+00 0.00000E+00
# -Range: 0-300
1.0000 HCO3- + 1.0000 H+ = CO +1.0000 H2O +0.5000 O2
-llnl_gamma 3.0
log_k -41.7002
-delta_H 277.069 kJ/mol # Calculated enthalpy of reaction CO
# Enthalpy of formation: -28.91 kcal/mol
-analytic 1.0028e+002 4.6877e-002 -1.8062e+004 -4.0263e+001 3.8031e+005
# -Range: 0-300
1.0000 Ce+++ + 0.5000 H2O = Ce++ +1.0000 H+ +0.2500 O2
-llnl_gamma 4.5
log_k -83.6754
-delta_H 0 # Not possible to calculate enthalpy of reaction Ce+2
# Enthalpy of formation: -0 kcal/mol
1.0000 H+ + 1.0000 Ce+++ + 0.2500 O2 = Ce++++ +0.5000 H2O
-llnl_gamma 5.5
log_k -7.9154
-delta_H 0 # Not possible to calculate enthalpy of reaction Ce+4
# Enthalpy of formation: -0 kcal/mol
1.0000 Cl- + 0.5000 O2 = ClO-
-llnl_gamma 4.0
log_k -15.1014
-delta_H 66.0361 kJ/mol # Calculated enthalpy of reaction ClO-
# Enthalpy of formation: -25.6 kcal/mol
-analytic 6.1314e+001 3.4812e-003 -6.0952e+003 -2.3043e+001 -9.5128e+001
# -Range: 0-300
1.0000 O2 + 1.0000 Cl- = ClO2-
-llnl_gamma 4.0
log_k -23.108
-delta_H 112.688 kJ/mol # Calculated enthalpy of reaction ClO2-
# Enthalpy of formation: -15.9 kcal/mol
-analytic 3.3638e+000 -6.1675e-003 -4.9726e+003 -2.0467e+000 -2.5769e+005
# -Range: 0-300
1.5000 O2 + 1.0000 Cl- = ClO3-
-llnl_gamma 3.5
log_k -17.2608
-delta_H 81.3077 kJ/mol # Calculated enthalpy of reaction ClO3-
# Enthalpy of formation: -24.85 kcal/mol
-analytic 2.8852e+001 -4.8281e-003 -4.6779e+003 -1.0772e+001 -2.0783e+005
# -Range: 0-300
2.0000 O2 + 1.0000 Cl- = ClO4-
-llnl_gamma 3.5
log_k -15.7091
-delta_H 62.0194 kJ/mol # Calculated enthalpy of reaction ClO4-
# Enthalpy of formation: -30.91 kcal/mol
-analytic 7.0280e+001 -6.8927e-005 -5.5690e+003 -2.6446e+001 -1.6596e+005
# -Range: 0-300
1.0000 H+ + 1.0000 Co++ + 0.2500 O2 = Co+++ +0.5000 H2O
-llnl_gamma 5.0
log_k -11.4845
-delta_H 10.3198 kJ/mol # Calculated enthalpy of reaction Co+3
# Enthalpy of formation: 22 kcal/mol
-analytic -2.2827e+001 -1.2222e-002 -7.2117e+002 7.0306e+000 -1.1247e+001
# -Range: 0-300
4.0000 H+ + 1.0000 CrO4-- = Cr++ +2.0000 H2O +1.0000 O2
-llnl_gamma 4.5
log_k -21.6373
-delta_H 153.829 kJ/mol # Calculated enthalpy of reaction Cr+2
# Enthalpy of formation: -34.3 kcal/mol
-analytic 6.9003e+001 6.2884e-002 -6.9847e+003 -3.4720e+001 -1.0901e+002
# -Range: 0-300
5.0000 H+ + 1.0000 CrO4-- = Cr+++ +2.5000 H2O +0.7500 O2
-llnl_gamma 9.0
log_k +8.3842
-delta_H -81.0336 kJ/mol # Calculated enthalpy of reaction Cr+3
# Enthalpy of formation: -57 kcal/mol
-analytic 5.1963e+001 6.0932e-002 5.4256e+003 -3.2290e+001 8.4645e+001
# -Range: 0-300
0.5000 H2O + 1.0000 CrO4-- = CrO4--- +1.0000 H+ +0.2500 O2
-llnl_gamma 4.0
log_k -19.7709
-delta_H 0 # Not possible to calculate enthalpy of reaction CrO4-3
# Enthalpy of formation: -0 kcal/mol
1.0000 Cu++ + 0.5000 H2O = Cu+ +1.0000 H+ +0.2500 O2
-llnl_gamma 4.0
log_k -18.7704
-delta_H 145.877 kJ/mol # Calculated enthalpy of reaction Cu+
# Enthalpy of formation: 17.132 kcal/mol
-analytic 3.7909e+001 1.3731e-002 -8.1506e+003 -1.3508e+001 -1.2719e+002
# -Range: 0-300
1.0000 Dy+++ + 0.5000 H2O = Dy++ +1.0000 H+ +0.2500 O2
-llnl_gamma 4.5
log_k -61.0754
-delta_H 0 # Not possible to calculate enthalpy of reaction Dy+2
# Enthalpy of formation: -0 kcal/mol
1.0000 Er+++ + 0.5000 H2O = Er++ +1.0000 H+ +0.2500 O2
-llnl_gamma 4.5
log_k -70.1754
-delta_H 0 # Not possible to calculate enthalpy of reaction Er+2
# Enthalpy of formation: -0 kcal/mol
1.0000 Eu+++ + 0.5000 H2O = Eu++ +1.0000 H+ +0.2500 O2
-llnl_gamma 4.5
log_k -27.5115
-delta_H 217.708 kJ/mol # Calculated enthalpy of reaction Eu+2
# Enthalpy of formation: -126.1 kcal/mol
-analytic 3.0300e+001 1.4126e-002 -1.2319e+004 -9.0585e+000 1.5289e+005
# -Range: 0-300
1.0000 H+ + 1.0000 Fe++ + 0.2500 O2 = Fe+++ +0.5000 H2O
-llnl_gamma 9.0
log_k +8.4899
-delta_H -97.209 kJ/mol # Calculated enthalpy of reaction Fe+3
# Enthalpy of formation: -11.85 kcal/mol
-analytic -1.7808e+001 -1.1753e-002 4.7609e+003 5.5866e+000 7.4295e+001
# -Range: 0-300
1.0000 Gd+++ + 0.5000 H2O = Gd++ +1.0000 H+ +0.2500 O2
-llnl_gamma 4.5
log_k -84.6754
-delta_H 0 # Not possible to calculate enthalpy of reaction Gd+2
# Enthalpy of formation: -0 kcal/mol
1.0000 H2O = H2 +0.5000 O2
-CO2_llnl_gamma
log_k -46.1066
-delta_H 275.588 kJ/mol # Calculated enthalpy of reaction H2
# Enthalpy of formation: -1 kcal/mol
-analytic 6.6835e+001 1.7172e-002 -1.8849e+004 -2.4092e+001 4.2501e+005
# -Range: 0-300
1.0000 H2AsO4- = H2AsO3- +0.5000 O2
-llnl_gamma 4.0
log_k -30.5349
-delta_H 188.698 kJ/mol # Calculated enthalpy of reaction H2AsO3-
# Enthalpy of formation: -170.84 kcal/mol
-analytic 7.4245e+001 1.4885e-002 -1.4218e+004 -2.6403e+001 3.3822e+005
# -Range: 0-300
1.0000 SO4-- + 1.0000 H+ + 0.5000 O2 = HSO5-
-llnl_gamma 4.0
log_k -17.2865
-delta_H 140.038 kJ/mol # Calculated enthalpy of reaction HSO5-
# Enthalpy of formation: -185.38 kcal/mol
-analytic 5.9944e+001 3.0904e-002 -7.7494e+003 -2.4420e+001 -1.2094e+002
# -Range: 0-300
1.0000 SeO3-- + 1.0000 H+ = HSe- +1.5000 O2
-llnl_gamma 4.0
log_k -76.8418
-delta_H 506.892 kJ/mol # Calculated enthalpy of reaction HSe-
# Enthalpy of formation: 3.8 kcal/mol
-analytic 4.7105e+001 4.3116e-002 -2.6949e+004 -1.9895e+001 2.5305e+005
# -Range: 0-300
2.0000 Hg++ + 1.0000 H2O = Hg2++ +2.0000 H+ +0.5000 O2
-llnl_gamma 4.0
log_k -12.208
-delta_H 106.261 kJ/mol # Calculated enthalpy of reaction Hg2+2
# Enthalpy of formation: 39.87 kcal/mol
-analytic 5.5010e+001 1.9050e-002 -4.7967e+003 -2.2952e+001 -7.4864e+001
# -Range: 0-300
1.0000 Ho+++ + 0.5000 H2O = Ho++ +1.0000 H+ +0.2500 O2
-llnl_gamma 4.5
log_k -67.3754
-delta_H 0 # Not possible to calculate enthalpy of reaction Ho+2
# Enthalpy of formation: -0 kcal/mol
3.0000 I- + 2.0000 H+ + 0.5000 O2 = I3- +1.0000 H2O
-llnl_gamma 4.0
log_k +24.7278
-delta_H -160.528 kJ/mol # Calculated enthalpy of reaction I3-
# Enthalpy of formation: -12.3 kcal/mol
-analytic 1.4788e+002 6.6206e-002 5.7407e+003 -6.5517e+001 8.9535e+001
# -Range: 0-300
1.0000 I- + 0.5000 O2 = IO-
-llnl_gamma 4.0
log_k -0.9038
-delta_H -44.5596 kJ/mol # Calculated enthalpy of reaction IO-
# Enthalpy of formation: -25.7 kcal/mol
-analytic 2.7568e+000 -5.5671e-003 3.2484e+003 -3.9065e+000 -2.8800e+005
# -Range: 0-300
1.5000 O2 + 1.0000 I- = IO3-
-llnl_gamma 4.0
log_k +17.6809
-delta_H -146.231 kJ/mol # Calculated enthalpy of reaction IO3-
# Enthalpy of formation: -52.9 kcal/mol
-analytic -2.2971e+001 -1.3478e-002 9.5977e+003 6.6010e+000 -3.4371e+005
# -Range: 0-300
2.0000 O2 + 1.0000 I- = IO4-
-llnl_gamma 3.5
log_k +6.9621
-delta_H -70.2912 kJ/mol # Calculated enthalpy of reaction IO4-
# Enthalpy of formation: -36.2 kcal/mol
-analytic 2.1232e+001 -7.8107e-003 3.5803e+003 -8.5272e+000 -2.5422e+005
# -Range: 0-300
1.0000 La+++ + 0.5000 H2O = La++ +1.0000 H+ +0.2500 O2
-llnl_gamma 4.5
log_k -72.4754
-delta_H 0 # Not possible to calculate enthalpy of reaction La+2
# Enthalpy of formation: -0 kcal/mol
1.0000 Mn++ + 1.0000 H+ + 0.2500 O2 = Mn+++ +0.5000 H2O
-llnl_gamma 5.0
log_k -4.0811
-delta_H -65.2892 kJ/mol # Calculated enthalpy of reaction Mn+3
# Enthalpy of formation: -34.895 kcal/mol
-analytic 3.8873e+001 1.7458e-002 2.0757e+003 -2.2274e+001 3.2378e+001
# -Range: 0-300
2.0000 H2O + 1.0000 O2 + 1.0000 Mn++ = MnO4-- +4.0000 H+
-llnl_gamma 4.0
log_k -32.4146
-delta_H 151.703 kJ/mol # Calculated enthalpy of reaction MnO4-2
# Enthalpy of formation: -156 kcal/mol
-analytic -1.0407e+001 -4.6464e-002 -1.0515e+004 1.0943e+001 -1.6408e+002
# -Range: 0-300
2.0000 NH3 + 1.5000 O2 = N2 +3.0000 H2O
-llnl_gamma 3.0
log_k +116.4609
-delta_H -687.08 kJ/mol # Calculated enthalpy of reaction N2
# Enthalpy of formation: -2.495 kcal/mol
-analytic -8.2621e+001 -1.4671e-002 4.0068e+004 2.9090e+001 -2.5924e+005
# -Range: 0-300
3.0000 NH3 + 2.0000 O2 = N3- +4.0000 H2O +1.0000 H+
-llnl_gamma 4.0
log_k +96.9680
-delta_H -599.935 kJ/mol # Calculated enthalpy of reaction N3-
# Enthalpy of formation: 275.14 kJ/mol
-analytic -9.1080e+001 -4.0817e-002 3.6350e+004 3.4484e+001 -6.2678e+005
# -Range: 0-300
1.5000 O2 + 1.0000 NH3 = NO2- +1.0000 H+ +1.0000 H2O
-llnl_gamma 3.0
log_k +46.8653
-delta_H -290.901 kJ/mol # Calculated enthalpy of reaction NO2-
# Enthalpy of formation: -25 kcal/mol
-analytic -1.7011e+001 -3.3459e-002 1.3999e+004 1.1078e+001 -4.8255e+004
# -Range: 0-300
2.0000 O2 + 1.0000 NH3 = NO3- +1.0000 H+ +1.0000 H2O
-llnl_gamma 3.0
log_k +62.1001
-delta_H -387.045 kJ/mol # Calculated enthalpy of reaction NO3-
# Enthalpy of formation: -49.429 kcal/mol
-analytic -3.9468e+001 -3.9697e-002 2.0614e+004 1.8872e+001 -2.1917e+005
# -Range: 0-300
1.0000 Nd+++ + 0.5000 H2O = Nd++ +1.0000 H+ +0.2500 O2
-llnl_gamma 4.5
log_k -64.3754
-delta_H 0 # Not possible to calculate enthalpy of reaction Nd+2
# Enthalpy of formation: -0 kcal/mol
1.0000 Np++++ + 0.5000 H2O = Np+++ +1.0000 H+ +0.2500 O2
-llnl_gamma 5.0
log_k -19.0131
-delta_H 168.787 kJ/mol # Calculated enthalpy of reaction Np+3
# Enthalpy of formation: -527.1 kJ/mol
-analytic 1.6615e+001 2.4645e-003 -8.9343e+003 -2.5829e+000 -1.3942e+002
# -Range: 0-300
1.5000 H2O + 1.0000 Np++++ + 0.2500 O2 = NpO2+ +3.0000 H+
-llnl_gamma 4.0
log_k +10.5928
-delta_H 9.80089 kJ/mol # Calculated enthalpy of reaction NpO2+
# Enthalpy of formation: -977.991 kJ/mol
-analytic 1.2566e+001 7.5467e-003 1.6921e+003 -2.7125e+000 -2.8381e+005
# -Range: 0-300
1.0000 Np++++ + 1.0000 H2O + 0.5000 O2 = NpO2++ +2.0000 H+
-llnl_gamma 4.5
log_k +11.2107
-delta_H -12.5719 kJ/mol # Calculated enthalpy of reaction NpO2+2
# Enthalpy of formation: -860.478 kJ/mol
-analytic 2.5510e+001 1.1973e-003 1.2753e+003 -6.7082e+000 -2.0792e+005
# -Range: 0-300
2.0000 H+ + 1.0000 Pb++ + 0.5000 O2 = Pb++++ +1.0000 H2O
-llnl_gamma 5.5
log_k -14.1802
-delta_H 0 # Not possible to calculate enthalpy of reaction Pb+4
# Enthalpy of formation: -0 kcal/mol
1.0000 Pm+++ + 0.5000 H2O = Pm++ +1.0000 H+ +0.2500 O2
-llnl_gamma 4.5
log_k -65.2754
-delta_H 0 # Not possible to calculate enthalpy of reaction Pm+2
# Enthalpy of formation: -0 kcal/mol
1.0000 Pr+++ + 0.5000 H2O = Pr++ +1.0000 H+ +0.2500 O2
-llnl_gamma 4.5
log_k -79.9754
-delta_H 0 # Not possible to calculate enthalpy of reaction Pr+2
# Enthalpy of formation: -0 kcal/mol
1.0000 Pu++++ + 0.5000 H2O = Pu+++ +1.0000 H+ +0.2500 O2
-llnl_gamma 5.0
log_k -4.5071
-delta_H 84.2268 kJ/mol # Calculated enthalpy of reaction Pu+3
# Enthalpy of formation: -591.552 kJ/mol
-analytic 2.0655e+001 3.2688e-003 -4.7434e+003 -4.1907e+000 1.2944e+004
# -Range: 0-300
1.5000 H2O + 1.0000 Pu++++ + 0.2500 O2 = PuO2+ +3.0000 H+
-llnl_gamma 4.0
log_k +2.9369
-delta_H 53.5009 kJ/mol # Calculated enthalpy of reaction PuO2+
# Enthalpy of formation: -914.183 kJ/mol
-analytic -2.0464e+001 2.8265e-003 1.2131e+003 9.2156e+000 -3.8400e+005
# -Range: 0-300
1.0000 Pu++++ + 1.0000 H2O + 0.5000 O2 = PuO2++ +2.0000 H+
-llnl_gamma 4.5
log_k +8.1273
-delta_H 6.22013 kJ/mol # Calculated enthalpy of reaction PuO2+2
# Enthalpy of formation: -821.578 kJ/mol
-analytic 3.5219e+001 2.5202e-003 -2.4760e+002 -1.0120e+001 -1.7569e+005
# -Range: 0-300
4.0000 H+ + 1.0000 RuO4-- = Ru(OH)2++ +1.0000 H2O +0.5000 O2
-llnl_gamma 4.5
log_k +25.2470
-delta_H 0 # Not possible to calculate enthalpy of reaction Ru(OH)2+2
# Enthalpy of formation: -0 kcal/mol
4.0000 H+ + 1.0000 RuO4-- = Ru++ +2.0000 H2O +1.0000 O2
-llnl_gamma 4.5
log_k +0.1610
-delta_H 0 # Not possible to calculate enthalpy of reaction Ru+2
# Enthalpy of formation: -0 kcal/mol
5.0000 H+ + 1.0000 RuO4-- = Ru+++ +2.5000 H2O +0.7500 O2
-llnl_gamma 5.0
log_k +17.6149
-delta_H 0 # Not possible to calculate enthalpy of reaction Ru+3
# Enthalpy of formation: -0 kcal/mol
2.0000 H+ + 1.0000 RuO4-- + 0.5000 O2 = RuO4 +1.0000 H2O
-llnl_gamma 3.0
log_k +16.2672
-delta_H -60.8385 kJ/mol # Calculated enthalpy of reaction RuO4
# Enthalpy of formation: -238.142 kJ/mol
-analytic 1.9964e+002 6.8286e-002 -1.2020e+003 -8.0706e+001 -2.0481e+001
# -Range: 0-200
1.0000 RuO4-- + 1.0000 H+ + 0.2500 O2 = RuO4- +0.5000 H2O
-llnl_gamma 4.0
log_k +11.6024
-delta_H -16.1998 kJ/mol # Calculated enthalpy of reaction RuO4-
# Enthalpy of formation: -333.389 kJ/mol
-analytic -1.9653e+000 8.8623e-003 1.8588e+003 1.8998e+000 2.9005e+001
# -Range: 0-300
2.0000 H+ + 2.0000 SO3-- = S2O4-- + .500 O2 + H2O
-llnl_gamma 5.0
# log_k -25.2075
log_k -25.2076
-delta_H 0 # Not possible to calculate enthalpy of reaction S2O4-2
# Enthalpy of formation: -0 kcal/mol
# -analytic -0.15158E+05 -0.31356E+01 0.47072E+06 0.58544E+04 0.73497E+04
-analytic -2.3172e2 2.0393e-3 -7.1011e0 8.3239e1 9.4155e-1
# changed 3/23/04, corrected to supcrt temperature dependence, GMA
# -Range: 0-300
# 2.0000 SO3-- + .500 O2 + 2.0000 H+ = S2O6-- + H2O
# H2O = .5 O2 + 2H+ + 2e-
2SO3-- = S2O6-- + 2e-
-llnl_gamma 4.0
log_k 41.8289
-delta_H 0 # Not possible to calculate enthalpy of reaction S2O6-2
# Enthalpy of formation: -0 kcal/mol
-analytic 0.14458E+03 0.61449E-01 0.71877E+04 -0.58657E+02 0.11211E+03
# -Range: 0-300
-add_logk Log_K_O2 0.5
2.0000 SO3-- + 1.500 O2 + 2.0000 H+ = S2O8-- + H2O
-llnl_gamma 4.0
log_k 70.7489
-delta_H 0 # Not possible to calculate enthalpy of reaction S2O8-2
# Enthalpy of formation: -0 kcal/mol
-analytic 0.18394E+03 0.60414E-01 0.13864E+05 -0.71804E+02 0.21628E+03
# -Range: 0-300
O2 + H+ + 3.0000 HS- = S3-- + 2.0000 H2O
# 2H2O = O2 + 4H+ + 4e-
#3HS- = S3-- + 3H+ + 4e-
-llnl_gamma 4.0
log_k 79.3915
-delta_H 0 # Not possible to calculate enthalpy of reaction S3-2
# Enthalpy of formation: -0 kcal/mol
-analytic -0.51626E+02 0.70208E-02 0.31797E+05 0.11927E+02 -0.64249E+06
-mass_balance S(-2)3
# -Range: 0-300
# -add_logk Log_K_O2 1.0
# 3.0000 SO3-- + 4.0000 H+ = S3O6-- + .500 O2 + 2.0000 H2O
# .5 O2 + 2H+ + 2e- = H2O
3SO3-- + 6 H+ + 2e- = S3O6-- + 3H2O
-llnl_gamma 4.0
log_k -6.2316
-delta_H 0 # Not possible to calculate enthalpy of reaction S3O6-2
# Enthalpy of formation: -0 kcal/mol
-analytic 0.23664E+03 0.12702E+00 -0.10110E+05 -0.99715E+02 -0.15783E+03
# -Range: 0-300
-add_logk Log_K_O2 -0.5
1.5000 O2 + 2.0000 H+ + 4.0000 HS- = S4-- + 3.0000 H2O
#4 HS- = S4-- + 4H+ + 6e-
-llnl_gamma 4.0
log_k 125.2958
-delta_H 0 # Not possible to calculate enthalpy of reaction S4-2
# Enthalpy of formation: -0 kcal/mol
-analytic 0.20875E+03 0.58133E-01 0.33278E+05 -0.85833E+02 0.51921E+03
-mass_balance S(-2)4
# -Range: 0-300
# -add_logk Log_K_O2 1.5
# 4.0000 SO3-- + 6.0000 H+ = S4O6-- + 1.500 O2 + 3.0000 H2O
4 SO3-- + 12 H+ + 6e- = S4O6-- + 6H2O
-llnl_gamma 4.0
log_k -38.3859
-delta_H 0 # Not possible to calculate enthalpy of reaction S4O6-2
# Enthalpy of formation: -0 kcal/mol
-analytic 0.32239E+03 0.19555E+00 -0.23617E+05 -0.13729E+03 -0.36862E+03
# -Range: 0-300
-add_logk Log_K_O2 -1.5
2.0000 O2 + 3.0000 H+ + 5.0000 HS- = S5-- + 4.0000 H2O
#5 HS- = S5-- + 5H+ + 8e-
-llnl_gamma 4.0
log_k 170.9802
-delta_H 0 # Not possible to calculate enthalpy of reaction S5-2
# Enthalpy of formation: -0 kcal/mol
-analytic 0.30329E+03 0.88033E-01 0.44739E+05 -0.12471E+03 0.69803E+03
-mass_balance S(-2)5
# -Range: 0-300
# -add_logk Log_K_O2 2
# 5.0000 SO3-- + 8.0000 H+ = S5O6-- + 2.5000 O2 + 4.0000 H2O
# 2.5O2 + 10 H+ + 10e- = 5H2O
5SO3-- + 18H+ + 10e- = S5O6-- + 9H2O
-llnl_gamma 4.0
log_k -99.4206
-delta_H 0 # Not possible to calculate enthalpy of reaction S5O6-2
# Enthalpy of formation: -0 kcal/mol
-analytic 0.42074E+03 0.25833E+00 -0.43878E+05 -0.18178E+03 -0.68480E+03
# -Range: 0-300
-add_logk Log_K_O2 -2.5
# 1.0000 H+ + HCO3- + HS- + NH3 = SCN- + 3.0000 H2O
# -llnl_gamma 3.5
# log_k 3.0070
# -delta_H 0 # Not possible to calculate enthalpy of reaction SCN-
## Enthalpy of formation: -0 kcal/mol
# -analytic 0.16539E+03 0.49623E-01 -0.44624E+04 -0.65544E+02 -0.69680E+02
## -Range: 0-300
Thiocyanate- = Thiocyanate-
log_k 0.0
1.0000 SO4-- = SO3-- +0.5000 O2
-llnl_gamma 4.5
log_k -46.6244
-delta_H 267.985 kJ/mol # Calculated enthalpy of reaction SO3-2
# Enthalpy of formation: -151.9 kcal/mol
-analytic -1.3771e+001 6.5102e-004 -1.3330e+004 4.7164e+000 -2.0800e+002
# -Range: 0-300
1.0000 HSe- = Se-- + 1.0000 H+
-llnl_gamma 4.0
log_k -14.9534
-delta_H 0 # Not possible to calculate enthalpy of reaction Se-2
# Enthalpy of formation: -0 kcal/mol
-analytic 1.0244e+002 3.1346e-002 -5.4190e+003 -4.3871e+001 -8.4589e+001
# -Range: 0-300
1.0000 SeO3-- + 0.5000 O2 = SeO4--
-llnl_gamma 4.0
log_k +13.9836
-delta_H -83.8892 kJ/mol # Calculated enthalpy of reaction SeO4-2
# Enthalpy of formation: -143.2 kcal/mol
-analytic -7.2314e+001 -1.3657e-002 8.6969e+003 2.6182e+001 -3.1897e+005
# -Range: 0-300
1.0000 Sm+++ + 0.5000 H2O = Sm++ +1.0000 H+ +0.2500 O2
-llnl_gamma 4.5
log_k -47.9624
-delta_H 326.911 kJ/mol # Calculated enthalpy of reaction Sm+2
# Enthalpy of formation: -120.5 kcal/mol
-analytic -1.0217e+001 7.7548e-003 -1.6285e+004 5.4711e+000 9.1931e+004
# -Range: 0-300
2.0000 H+ + 1.0000 Sn++ + 0.5000 O2 = Sn++++ +1.0000 H2O
-llnl_gamma 11.0
log_k +37.7020
-delta_H -240.739 kJ/mol # Calculated enthalpy of reaction Sn+4
# Enthalpy of formation: 7.229 kcal/mol
-analytic 3.2053e+001 -9.2307e-003 1.0378e+004 -1.0666e+001 1.6193e+002
# -Range: 0-300
1.0000 Tb+++ + 0.5000 H2O = Tb++ +1.0000 H+ +0.2500 O2
-llnl_gamma 4.5
log_k -78.7754
-delta_H 0 # Not possible to calculate enthalpy of reaction Tb+2
# Enthalpy of formation: -0 kcal/mol
4.0000 H+ + 1.0000 TcO4- = Tc+++ +2.0000 H2O +1.0000 O2
-llnl_gamma 5.0
log_k -47.614
-delta_H 0 # Not possible to calculate enthalpy of reaction Tc+3
# Enthalpy of formation: -0 kcal/mol
3.0000 H+ + 1.0000 TcO4- = TcO++ +1.5000 H2O +0.7500 O2
-llnl_gamma 4.5
log_k -31.5059
-delta_H 0 # Not possible to calculate enthalpy of reaction TcO+2
# Enthalpy of formation: -0 kcal/mol
1.0000 TcO4- + 0.5000 H2O = TcO4-- +1.0000 H+ +0.2500 O2
-llnl_gamma 4.0
log_k -31.8197
-delta_H 0 # Not possible to calculate enthalpy of reaction TcO4-2
# Enthalpy of formation: -0 kcal/mol
1.0000 TcO4- + 1.0000 H2O = TcO4--- +2.0000 H+ +0.5000 O2
-llnl_gamma 4.0
log_k -63.2889
-delta_H 0 # Not possible to calculate enthalpy of reaction TcO4-3
# Enthalpy of formation: -0 kcal/mol
2.0000 H+ + 1.0000 Tl+ + 0.5000 O2 = Tl+++ +1.0000 H2O
-llnl_gamma 5.0
log_k -0.2751
-delta_H -88.479 kJ/mol # Calculated enthalpy of reaction Tl+3
# Enthalpy of formation: 47 kcal/mol
-analytic -6.7978e+001 -2.6430e-002 5.3106e+003 2.3340e+001 8.2887e+001
# -Range: 0-300
1.0000 Tm+++ + 0.5000 H2O = Tm++ +1.0000 H+ +0.2500 O2
-llnl_gamma 4.5
log_k -58.3754
-delta_H 0 # Not possible to calculate enthalpy of reaction Tm+2
# Enthalpy of formation: -0 kcal/mol
1.0000 UO2++ + 1.0000 H+ = U+++ +0.7500 O2 +0.5000 H2O
-llnl_gamma 5.0
log_k -64.8028
-delta_H 377.881 kJ/mol # Calculated enthalpy of reaction U+3
# Enthalpy of formation: -489.1 kJ/mol
-analytic 2.5133e+001 6.4088e-003 -2.2542e+004 -8.1423e+000 3.4793e+005
# -Range: 0-300
2.0000 H+ + 1.0000 UO2++ = U++++ +1.0000 H2O +0.5000 O2
-llnl_gamma 5.5
log_k -33.9491
-delta_H 135.895 kJ/mol # Calculated enthalpy of reaction U+4
# Enthalpy of formation: -591.2 kJ/mol
-analytic 4.4837e+001 1.0129e-002 -1.1787e+004 -1.9194e+001 4.6436e+005
# -Range: 0-300
1.0000 UO2++ + 0.5000 H2O = UO2+ +1.0000 H+ +0.2500 O2
-llnl_gamma 4.0
log_k -20.0169
-delta_H 133.759 kJ/mol # Calculated enthalpy of reaction UO2+
# Enthalpy of formation: -1025.13 kJ/mol
-analytic 8.0480e+000 9.5845e-003 -6.5994e+003 -3.5515e+000 -1.0298e+002
# -Range: 0-300
1.0000 VO++ + 1.0000 H+ = V+++ +0.5000 H2O +0.2500 O2
-llnl_gamma 5.0
log_k -15.7191
-delta_H 79.6069 kJ/mol # Calculated enthalpy of reaction V+3
# Enthalpy of formation: -62.39 kcal/mol
-analytic 1.6167e+001 1.1963e-002 -4.2112e+003 -8.6126e+000 -6.5717e+001
# -Range: 0-300
1.0000 VO++ + 0.5000 H2O + 0.2500 O2 = VO2+ +1.0000 H+
-llnl_gamma 4.0
log_k +4.5774
-delta_H -17.2234 kJ/mol # Calculated enthalpy of reaction VO2+
# Enthalpy of formation: -155.3 kcal/mol
-analytic 1.9732e+000 5.3936e-003 1.2240e+003 -1.2539e+000 1.9098e+001
# -Range: 0-300
1.0000 VO2+ + 2.0000 H2O = VO4--- +4.0000 H+
-llnl_gamma 4.0
log_k -28.4475
-delta_H 0 # Not possible to calculate enthalpy of reaction VO4-3
# Enthalpy of formation: -0 kcal/mol
1.0000 Yb+++ + 0.5000 H2O = Yb++ +1.0000 H+ +0.2500 O2
-llnl_gamma 4.5
log_k -39.4595
-delta_H 280.05 kJ/mol # Calculated enthalpy of reaction Yb+2
# Enthalpy of formation: -126.8 kcal/mol
-analytic 1.0773e+000 9.5995e-003 -1.3833e+004 1.0723e+000 3.1365e+004
# -Range: 0-300
2.0000 H+ + 1.0000 Zr(OH)2++ = Zr++++ +2.0000 H2O
-llnl_gamma 11.0
log_k +0.2385
-delta_H 0 # Not possible to calculate enthalpy of reaction Zr+4
# Enthalpy of formation: -0 kcal/mol
4.0000 HS- + 4.0000 H+ + 2.0000 Sb(OH)3 + 2.0000 NH3 = (NH4)2Sb2S4 +6.0000 H2O
-llnl_gamma 3.0
log_k +67.6490
-delta_H -424.665 kJ/mol # Calculated enthalpy of reaction (NH4)2Sb2S4
# Enthalpy of formation: -484.321 kJ/mol
-analytic -3.9259e+002 -1.1727e-001 3.2073e+004 1.5667e+002 5.4478e+002
# -Range: 0-200
2.0000 NpO2++ + 2.0000 H2O = (NpO2)2(OH)2++ +2.0000 H+
-llnl_gamma 4.5
log_k -6.4
-delta_H 45.4397 kJ/mol # Calculated enthalpy of reaction (NpO2)2(OH)2+2
# Enthalpy of formation: -537.092 kcal/mol
-analytic -4.7462e+001 -3.1413e-002 -2.1954e+003 2.3355e+001 -3.7424e+001
# -Range: 25-150
5.0000 H2O + 3.0000 NpO2++ = (NpO2)3(OH)5+ +5.0000 H+
-llnl_gamma 4.0
log_k -17.5
-delta_H 112.322 kJ/mol # Calculated enthalpy of reaction (NpO2)3(OH)5+
# Enthalpy of formation: -931.717 kcal/mol
-analytic 5.4053e+002 9.1693e-002 -2.4404e+004 -2.0349e+002 -4.1639e+002
# -Range: 25-150
2.0000 PuO2++ + 2.0000 H2O = (PuO2)2(OH)2++ +2.0000 H+
-llnl_gamma 4.5
log_k -8.2626
-delta_H 57.8597 kJ/mol # Calculated enthalpy of reaction (PuO2)2(OH)2+2
# Enthalpy of formation: -2156.97 kJ/mol
-analytic 6.5448e+001 -1.6194e-003 -5.9542e+003 -2.1522e+001 -9.2929e+001
# -Range: 0-300
5.0000 H2O + 3.0000 PuO2++ = (PuO2)3(OH)5+ +5.0000 H+
-llnl_gamma 4.0
log_k -21.655
-delta_H 139.617 kJ/mol # Calculated enthalpy of reaction (PuO2)3(OH)5+
# Enthalpy of formation: -3754.31 kJ/mol
-analytic 1.6151e+002 5.8182e-003 -1.4002e+004 -5.5745e+001 -2.1854e+002
# -Range: 0-300
4.0000 H2O + 2.0000 TcO++ = (TcO(OH)2)2 +4.0000 H+
-llnl_gamma 3.0
log_k -0.1271
-delta_H 0 # Not possible to calculate enthalpy of reaction (TcO(OH)2)2
# Enthalpy of formation: -0 kcal/mol
12.0000 H2O + 11.0000 UO2++ + 6.0000 HCO3- = (UO2)11(CO3)6(OH)12-2 +18.0000 H+
-llnl_gamma 4.0
log_k -25.7347
-delta_H 0 # Not possible to calculate enthalpy of reaction (UO2)11(CO3)6(OH)12-2
# Enthalpy of formation: -0 kcal/mol
2.0000 UO2++ + 2.0000 H2O = (UO2)2(OH)2++ +2.0000 H+
-llnl_gamma 4.5
log_k -5.6346
-delta_H 37.6127 kJ/mol # Calculated enthalpy of reaction (UO2)2(OH)2+2
# Enthalpy of formation: -2572.06 kJ/mol
-analytic 6.4509e+001 -7.6875e-004 -4.8433e+003 -2.1689e+001 -7.5593e+001
# -Range: 0-300
3.0000 H2O + 2.0000 UO2++ + 1.0000 HCO3- = (UO2)2CO3(OH)3- +4.0000 H+
-llnl_gamma 4.0
log_k -11.2229
-delta_H 0 # Not possible to calculate enthalpy of reaction (UO2)2CO3(OH)3-
# Enthalpy of formation: -0 kcal/mol
2.0000 UO2++ + 1.0000 H2O = (UO2)2OH+++ +1.0000 H+
-llnl_gamma 5.0
log_k -2.7072
-delta_H 0 # Not possible to calculate enthalpy of reaction (UO2)2OH+3
# Enthalpy of formation: -0 kcal/mol
6.0000 HCO3- + 3.0000 UO2++ = (UO2)3(CO3)6-6 +6.0000 H+
-llnl_gamma 4.0
log_k -8.0601
-delta_H 25.5204 kJ/mol # Calculated enthalpy of reaction (UO2)3(CO3)6-6
# Enthalpy of formation: -7171.08 kJ/mol
-analytic 7.4044e+002 2.7299e-001 -1.7614e+004 -3.1149e+002 -2.7507e+002
# -Range: 0-300
4.0000 H2O + 3.0000 UO2++ = (UO2)3(OH)4++ +4.0000 H+
-llnl_gamma 4.5
log_k -11.929
-delta_H 0 # Not possible to calculate enthalpy of reaction (UO2)3(OH)4+2
# Enthalpy of formation: -0 kcal/mol
5.0000 H2O + 3.0000 UO2++ = (UO2)3(OH)5+ +5.0000 H+
-llnl_gamma 4.0
log_k -15.5862
-delta_H 97.1056 kJ/mol # Calculated enthalpy of reaction (UO2)3(OH)5+
# Enthalpy of formation: -4389.09 kJ/mol
-analytic 1.6004e+002 7.0827e-003 -1.1700e+004 -5.5973e+001 -1.8261e+002
# -Range: 0-300
4.0000 H2O + 3.0000 UO2++ + 1.0000 HCO3- = (UO2)3(OH)5CO2+ +4.0000 H+
-llnl_gamma 4.0
log_k -9.6194
-delta_H 0 # Not possible to calculate enthalpy of reaction (UO2)3(OH)5CO2+
# Enthalpy of formation: -0 kcal/mol
7.0000 H2O + 3.0000 UO2++ = (UO2)3(OH)7- +7.0000 H+
-llnl_gamma 4.0
log_k -31.0508
-delta_H 0 # Not possible to calculate enthalpy of reaction (UO2)3(OH)7-
# Enthalpy of formation: -0 kcal/mol
3.0000 UO2++ + 3.0000 H2O + 1.0000 HCO3- = (UO2)3O(OH)2(HCO3)+ +4.0000 H+
-llnl_gamma 4.0
log_k -9.7129
-delta_H 0 # Not possible to calculate enthalpy of reaction (UO2)3O(OH)2(HCO3)+
# Enthalpy of formation: -0 kcal/mol
7.0000 H2O + 4.0000 UO2++ = (UO2)4(OH)7+ +7.0000 H+
-llnl_gamma 4.0
log_k -21.9508
-delta_H 0 # Not possible to calculate enthalpy of reaction (UO2)4(OH)7+
# Enthalpy of formation: -0 kcal/mol
2.0000 VO++ + 2.0000 H2O = (VO)2(OH)2++ +2.0000 H+
-llnl_gamma 4.5
log_k -6.67
-delta_H 0 # Not possible to calculate enthalpy of reaction (VO)2(OH)2+2
# Enthalpy of formation: -0 kcal/mol
HAcetate = Acetate- + H+
-llnl_gamma 4.5
log_k -4.7572
-delta_H 0 # Not possible to calculate enthalpy of reaction Acetate-
# Enthalpy of formation: -0 kcal/mol
-analytic -0.96597E+02 -0.34535E-01 0.19753E+04 0.38593E+02 0.30850E+02
# Range: 0-300
2.0000 HAcetate + 1.0000 Ag+ = Ag(Acetate)2- +2.0000 H+
-llnl_gamma 4.0
log_k -8.8716
-delta_H 0 # Not possible to calculate enthalpy of reaction Ag(Acetate)2-
# Enthalpy of formation: -0 kcal/mol
-analytic -2.8207e+002 -5.3713e-002 9.5343e+003 1.0396e+002 1.4886e+002
# -Range: 0-300
2.0000 HCO3- + 1.0000 Ag+ = Ag(CO3)2--- +2.0000 H+
-llnl_gamma 4.0
log_k -18.5062
-delta_H 1.34306 kJ/mol # Calculated enthalpy of reaction Ag(CO3)2-3
# Enthalpy of formation: -304.2 kcal/mol
-analytic -1.6671e+002 -4.5571e-002 3.7190e+003 6.0341e+001 5.8080e+001
# -Range: 0-300
1.0000 Ag+ + 1.0000 HAcetate = AgAcetate +1.0000 H+
-llnl_gamma 3.0
log_k -4.0264
-delta_H -3.4518 kJ/mol # Calculated enthalpy of reaction AgAcetate
# Enthalpy of formation: -91.65 kcal/mol
-analytic 6.9069e+000 -1.9415e-003 -1.9953e+003 -2.6175e+000 2.5092e+005
# -Range: 0-300
1.0000 HCO3- + 1.0000 Ag+ = AgCO3- +1.0000 H+
-llnl_gamma 4.0
log_k -7.6416
-delta_H -8.27177 kJ/mol # Calculated enthalpy of reaction AgCO3-
# Enthalpy of formation: -141.6 kcal/mol
-analytic 6.5598e+000 -1.6477e-004 -4.7079e+002 -5.0807e+000 -7.3484e+000
# -Range: 0-300
1.0000 Cl- + 1.0000 Ag+ = AgCl
-llnl_gamma 3.0
log_k +3.2971
-delta_H -15.1126 kJ/mol # Calculated enthalpy of reaction AgCl
#Enthalpy of formation: -18.27 kcal/mol
-analytic 1.0904e+002 3.5492e-002 -1.8455e+003 -4.4502e+001 -2.8830e+001
# -Range: 0-300
2.0000 Cl- + 1.0000 Ag+ = AgCl2-
-llnl_gamma 4.0
log_k +5.2989
-delta_H -27.3592 kJ/mol # Calculated enthalpy of reaction AgCl2-
#Enthalpy of formation: -61.13 kcal/mol
-analytic 9.2164e+001 4.0261e-002 -1.6597e+002 -3.9721e+001 -2.6171e+000
# -Range: 0-300
3.0000 Cl- + 1.0000 Ag+ = AgCl3--
-llnl_gamma 4.0
log_k +5.1310
-delta_H -47.7645 kJ/mol # Calculated enthalpy of reaction AgCl3-2
#Enthalpy of formation: -105.94 kcal/mol
-analytic 4.3732e+000 2.9568e-002 3.9818e+003 -8.6428e+000 6.2131e+001
# -Range: 0-300
4.0000 Cl- + 1.0000 Ag+ = AgCl4---
-llnl_gamma 4.0
log_k +3.8050
-delta_H -32.4804 kJ/mol # Calculated enthalpy of reaction AgCl4-3
#Enthalpy of formation: -142.22 kcal/mol
-analytic -1.6176e+001 2.9523e-002 0.0000e+000 0.0000e+000 9.9602e+005
# -Range: 0-300
1.0000 F- + 1.0000 Ag+ = AgF
-llnl_gamma 3.0
log_k -0.1668
-delta_H -9.298 kJ/mol # Calculated enthalpy of reaction AgF
# Enthalpy of formation: -238.895 kJ/mol
-analytic -6.6024e+001 -2.2350e-002 1.9514e+003 2.6663e+001 3.3160e+001
# -Range: 0-200
#1.0000 NO3- + 1.0000 Ag+ = AgNO3
# -llnl_gamma 3.0
# log_k -0.1979
# -delta_H 4.45178 kJ/mol # Calculated enthalpy of reaction AgNO3
# Enthalpy of formation: -23.09 kcal/mol
# -analytic 7.3866e+001 2.6050e-002 -1.5923e+003 -3.0904e+001 -2.4868e+001
# -Range: 0-300
2.0000 HAcetate + 1.0000 Al+++ = Al(Acetate)2+ +2.0000 H+
-llnl_gamma 4.0
log_k -5.595
-delta_H -46.8566 kJ/mol # Calculated enthalpy of reaction Al(Acetate)2+
# Enthalpy of formation: -372.08 kcal/mol
-analytic -4.2528e+001 2.1431e-003 3.1658e+002 1.1585e+001 5.8604e+005
# -Range: 0-300
2.0000 H2O + 1.0000 Al+++ = Al(OH)2+ +2.0000 H+
-llnl_gamma 4.0
log_k -10.5945
-delta_H 98.2822 kJ/mol # Calculated enthalpy of reaction Al(OH)2+
# Enthalpy of formation: -241.825 kcal/mol
-analytic 4.4036e+001 2.0168e-002 -5.5455e+003 -1.6987e+001 -8.6545e+001
# -Range: 0-300
2.0000 SO4-- + 1.0000 Al+++ = Al(SO4)2-
-llnl_gamma 4.0
log_k +4.9000
-delta_H 0 # Not possible to calculate enthalpy of reaction Al(SO4)2-
# Enthalpy of formation: -0 kcal/mol
28.0000 H2O + 13.0000 Al+++ = Al13O4(OH)24+7 +32.0000 H+
-llnl_gamma 6.0
log_k -98.73
-delta_H 0 # Not possible to calculate enthalpy of reaction Al13O4(OH)24+7
# Enthalpy of formation: -0 kcal/mol
2.0000 H2O + 2.0000 Al+++ = Al2(OH)2++++ +2.0000 H+
-llnl_gamma 5.5
log_k -7.6902
-delta_H 0 # Not possible to calculate enthalpy of reaction Al2(OH)2+4
# Enthalpy of formation: -0 kcal/mol
4.0000 H2O + 3.0000 Al+++ = Al3(OH)4+5 +4.0000 H+
-llnl_gamma 6.0
log_k -13.8803
-delta_H 0 # Not possible to calculate enthalpy of reaction Al3(OH)4+5
# Enthalpy of formation: -0 kcal/mol
1.0000 Al+++ + 1.0000 HAcetate = AlAcetate++ +1.0000 H+
-llnl_gamma 4.5
log_k -2.6923
-delta_H -18.1962 kJ/mol # Calculated enthalpy of reaction AlAcetate+2
# Enthalpy of formation: -249.13 kcal/mol
-analytic -1.9847e+001 2.0058e-003 -2.3653e+002 5.5454e+000 3.2362e+005
# -Range: 0-300
1.0000 F- + 1.0000 Al+++ = AlF++
-llnl_gamma 4.5
log_k +7.0000
-delta_H 0 # Not possible to calculate enthalpy of reaction AlF+2
# Enthalpy of formation: -0 kcal/mol
2.0000 F- + 1.0000 Al+++ = AlF2+
-llnl_gamma 4.0
log_k +12.6000
-delta_H 0 # Not possible to calculate enthalpy of reaction AlF2+
# Enthalpy of formation: -0 kcal/mol
3.0000 F- + 1.0000 Al+++ = AlF3
-llnl_gamma 3.0
log_k +16.7000
-delta_H 0 # Not possible to calculate enthalpy of reaction AlF3
# Enthalpy of formation: -0 kcal/mol
4.0000 F- + 1.0000 Al+++ = AlF4-
-llnl_gamma 4.0
log_k +19.1000
-delta_H 0 # Not possible to calculate enthalpy of reaction AlF4-
# Enthalpy of formation: -0 kcal/mol
1.0000 HPO4-- + 1.0000 H+ + 1.0000 Al+++ = AlH2PO4++
-llnl_gamma 4.5
log_k +3.1000
-delta_H 0 # Not possible to calculate enthalpy of reaction AlH2PO4+2
# Enthalpy of formation: -0 kcal/mol
1.0000 HPO4-- + 1.0000 Al+++ = AlHPO4+
-llnl_gamma 4.0
log_k +7.4000
-delta_H 0 # Not possible to calculate enthalpy of reaction AlHPO4+
# Enthalpy of formation: -0 kcal/mol
2.0000 H2O + 1.0000 Al+++ = AlO2- +4.0000 H+
-llnl_gamma 4.0
log_k -22.8833
-delta_H 180.899 kJ/mol # Calculated enthalpy of reaction AlO2-
# Enthalpy of formation: -222.079 kcal/mol
-analytic 1.0803e+001 -3.4379e-003 -9.7391e+003 0.0000e+000 0.0000e+000
# -Range: 0-300
1.0000 H2O + 1.0000 Al+++ = AlOH++ +1.0000 H+
-llnl_gamma 4.5
log_k -4.9571
-delta_H 49.798 kJ/mol # Calculated enthalpy of reaction AlOH+2
# Enthalpy of formation: -185.096 kcal/mol
-analytic -2.6224e-001 8.8816e-003 -1.8686e+003 -4.3195e-001 -2.9158e+001
# -Range: 0-300
1.0000 SO4-- + 1.0000 Al+++ = AlSO4+
-llnl_gamma 4.0
log_k +3.0100
-delta_H 0 # Not possible to calculate enthalpy of reaction AlSO4+
# Enthalpy of formation: -0 kcal/mol
2.0000 HCO3- + 1.0000 Am+++ = Am(CO3)2- +2.0000 H+
-llnl_gamma 4.0
log_k -8.3868
-delta_H 0 # Not possible to calculate enthalpy of reaction Am(CO3)2-
# Enthalpy of formation: -0 kcal/mol
3.0000 HCO3- + 1.0000 Am+++ = Am(CO3)3--- +3.0000 H+
-llnl_gamma 4.0
log_k -15.8302
-delta_H 0 # Not possible to calculate enthalpy of reaction Am(CO3)3-3
# Enthalpy of formation: -0 kcal/mol
5.0000 HCO3- + 1.0000 Am++++ = Am(CO3)5-6 +5.0000 H+
-llnl_gamma 4.0
log_k -12.409
-delta_H 0 # Not possible to calculate enthalpy of reaction Am(CO3)5-6
# Enthalpy of formation: -0 kcal/mol
2.0000 H2O + 1.0000 Am+++ = Am(OH)2+ +2.0000 H+
-llnl_gamma 4.0
log_k -14.1145
-delta_H 0 # Not possible to calculate enthalpy of reaction Am(OH)2+
# Enthalpy of formation: -0 kcal/mol
3.0000 H2O + 1.0000 Am+++ = Am(OH)3 +3.0000 H+
-llnl_gamma 3.0
log_k -25.7218
-delta_H 0 # Not possible to calculate enthalpy of reaction Am(OH)3
# Enthalpy of formation: -0 kcal/mol
2.0000 SO4-- + 1.0000 Am+++ = Am(SO4)2-
-llnl_gamma 4.0
log_k +5.2407
-delta_H 0 # Not possible to calculate enthalpy of reaction Am(SO4)2-
# Enthalpy of formation: -0 kcal/mol
1.0000 HCO3- + 1.0000 Am+++ = AmCO3+ +1.0000 H+
-llnl_gamma 4.0
log_k -2.5434
-delta_H 0 # Not possible to calculate enthalpy of reaction AmCO3+
# Enthalpy of formation: -0 kcal/mol
1.0000 Cl- + 1.0000 Am+++ = AmCl++
-llnl_gamma 4.5
log_k +1.0374
-delta_H 0 # Not possible to calculate enthalpy of reaction AmCl+2
# Enthalpy of formation: -0 kcal/mol
1.0000 F- + 1.0000 Am+++ = AmF++
-llnl_gamma 4.5
log_k +3.3601
-delta_H 0 # Not possible to calculate enthalpy of reaction AmF+2
# Enthalpy of formation: -0 kcal/mol
2.0000 F- + 1.0000 Am+++ = AmF2+
-llnl_gamma 4.0
log_k +5.7204
-delta_H 0 # Not possible to calculate enthalpy of reaction AmF2+
# Enthalpy of formation: -0 kcal/mol
1.0000 HPO4-- + 1.0000 H+ + 1.0000 Am+++ = AmH2PO4++
-llnl_gamma 4.5
log_k +11.4119
-delta_H 0 # Not possible to calculate enthalpy of reaction AmH2PO4+2
# Enthalpy of formation: -0 kcal/mol
1.0000 N3- + 1.0000 Am+++ = AmN3++
-llnl_gamma 4.5
log_k +1.6699
-delta_H 0 # Not possible to calculate enthalpy of reaction AmN3+2
# Enthalpy of formation: -0 kcal/mol
1.0000 NO3- + 1.0000 Am+++ = AmNO3++
-llnl_gamma 4.5
log_k +1.3104
-delta_H 0 # Not possible to calculate enthalpy of reaction AmNO3+2
# Enthalpy of formation: -0 kcal/mol
1.0000 H2O + 1.0000 Am+++ = AmOH++ +1.0000 H+
-llnl_gamma 4.5
log_k -6.4072
-delta_H 0 # Not possible to calculate enthalpy of reaction AmOH+2
# Enthalpy of formation: -0 kcal/mol
1.0000 SO4-- + 1.0000 Am+++ = AmSO4+
-llnl_gamma 4.0
log_k +3.7703
-delta_H 0 # Not possible to calculate enthalpy of reaction AmSO4+
# Enthalpy of formation: -0 kcal/mol
1.0000 H2AsO3- + 1.0000 H+ = As(OH)3
-llnl_gamma 3.0
log_k +9.2048
-delta_H -27.4054 kJ/mol # Calculated enthalpy of reaction As(OH)3
# Enthalpy of formation: -742.2 kJ/mol
-analytic 1.3020e+002 4.7513e-002 -1.1999e+003 -5.2993e+001 -2.0422e+001
# -Range: 0-200
1.0000 H2AsO3- = AsO2- +1.0000 H2O
-llnl_gamma 4.0
log_k 0.0111
-delta_H 0 # Not possible to calculate enthalpy of reaction AsO2-
# Enthalpy of formation: -0 kcal/mol
-analytic -2.1509e+001 -1.7680e-002 -1.9261e+001 1.0841e+001 -2.9404e-001
# -Range: 0-300
1.0000 H2AsO3- = AsO2OH-- +1.0000 H+
-llnl_gamma 4.0
log_k -11.0171
-delta_H 25.514 kJ/mol # Calculated enthalpy of reaction AsO2OH-2
# Enthalpy of formation: -164.742 kcal/mol
-analytic 1.4309e+002 1.8620e-002 -6.8596e+003 -5.5222e+001 -1.0708e+002
# -Range: 0-300
1.0000 H2AsO4- + 1.0000 F- = AsO3F-- +1.0000 H2O
-llnl_gamma 4.0
log_k +40.2451
-delta_H 0 # Not possible to calculate enthalpy of reaction AsO3F-2
# Enthalpy of formation: -0 kcal/mol
1.0000 H2AsO4- = AsO4--- +2.0000 H+
-llnl_gamma 4.0
log_k -18.3604
-delta_H 21.4198 kJ/mol # Calculated enthalpy of reaction AsO4-3
# Enthalpy of formation: -888.14 kJ/mol
-analytic -2.4979e+001 -1.2761e-002 2.8369e+003 3.4878e+000 -6.8736e+005
# -Range: 0-300
2.0000 HAcetate + 1.0000 Au+ = Au(Acetate)2- +2.0000 H+
-llnl_gamma 4.0
log_k -9.0013
-delta_H -8.91192 kJ/mol # Calculated enthalpy of reaction Au(Acetate)2-
# Enthalpy of formation: -186.75 kcal/mol
-analytic -2.2338e+002 -4.6312e-002 7.0942e+003 8.2606e+001 1.1076e+002
# -Range: 0-300
1.0000 Au+ + 1.0000 HAcetate = AuAcetate +1.0000 H+
-llnl_gamma 3.0
log_k -4.3174
-delta_H 0.87864 kJ/mol # Calculated enthalpy of reaction AuAcetate
# Enthalpy of formation: -68.31 kcal/mol
-analytic -1.1812e+000 -4.1120e-003 -1.4752e+003 4.5665e-001 1.7019e+005
# -Range: 0-300
2.0000 B(OH)3 = B2O(OH)5- +1.0000 H+
-llnl_gamma 4.0
log_k -18.6851
-delta_H 0 # Not possible to calculate enthalpy of reaction B2O(OH)5-
# Enthalpy of formation: -0 kcal/mol
2.0000 F- + 1.0000 H+ + 1.0000 B(OH)3 = BF2(OH)2- +1.0000 H2O
-llnl_gamma 4.0
log_k +6.6174
-delta_H 0 # Not possible to calculate enthalpy of reaction BF2(OH)2-
# Enthalpy of formation: -0 kcal/mol
3.0000 F- + 2.0000 H+ + 1.0000 B(OH)3 = BF3OH- +2.0000 H2O
-llnl_gamma 4.0
log_k +13.1908
-delta_H -178.577 kJ/mol # Calculated enthalpy of reaction BF3OH-
# Enthalpy of formation: -403.317 kcal/mol
-analytic 3.3411e+002 -3.7303e-002 -8.6507e+003 -1.1345e+002 -1.3508e+002
# -Range: 0-300
4.0000 F- + 3.0000 H+ + 1.0000 B(OH)3 = BF4- +3.0000 H2O
-llnl_gamma 4.0
log_k +18.0049
-delta_H -16.4473 kJ/mol # Calculated enthalpy of reaction BF4-
# Enthalpy of formation: -376.4 kcal/mol
-analytic 2.5491e+002 1.0443e-001 -3.3332e+003 -1.0378e+002 -5.2087e+001
# -Range: 0-300
1.0000 B(OH)3 = BO2- +1.0000 H+ +1.0000 H2O
-llnl_gamma 4.0
log_k -9.2449
-delta_H 16.3302 kJ/mol # Calculated enthalpy of reaction BO2-
# Enthalpy of formation: -184.6 kcal/mol
-analytic -1.0500e+002 -3.3447e-002 1.4706e+003 4.0724e+001 2.2978e+001
# -Range: 0-300
2.0000 HAcetate + 1.0000 Ba++ = Ba(Acetate)2 +2.0000 H+
-llnl_gamma 3.0
log_k -8.0118
-delta_H 11.255 kJ/mol # Calculated enthalpy of reaction Ba(Acetate)2
# Enthalpy of formation: -358.01 kcal/mol
-analytic -1.4566e+001 3.1394e-004 -3.9564e+003 5.1906e+000 6.1407e+005
# -Range: 0-300
1.0000 O_phthalate-2 + 1.0000 Ba++ = Ba(O_phthalate)
-llnl_gamma 3.0
log_k +2.3300
-delta_H 0 # Not possible to calculate enthalpy of reaction Ba(O_phthalate)
# Enthalpy of formation: -0 kcal/mol
1.0000 H2O + 1.0000 Ba++ + 1.0000 B(OH)3 = BaB(OH)4+ +1.0000 H+
-llnl_gamma 4.0
log_k -7.8012
-delta_H 0 # Not possible to calculate enthalpy of reaction BaB(OH)4+
# Enthalpy of formation: -0 kcal/mol
1.0000 Ba++ + 1.0000 HAcetate = BaAcetate+ +1.0000 H+
-llnl_gamma 4.0
log_k -3.7677
-delta_H 7.322 kJ/mol # Calculated enthalpy of reaction BaAcetate+
# Enthalpy of formation: -242.85 kcal/mol
-analytic -1.5623e+001 2.9282e-003 -3.9534e+002 4.3959e+000 1.2829e+005
# -Range: 0-300
1.0000 HCO3- + 1.0000 Ba++ = BaCO3 +1.0000 H+
-llnl_gamma 3.0
log_k -7.6834
-delta_H 31.5808 kJ/mol # Calculated enthalpy of reaction BaCO3
# Enthalpy of formation: -285.85 kcal/mol
-analytic 2.1878e+002 5.2368e-002 -8.2472e+003 -8.6644e+001 -1.2875e+002
# -Range: 0-300
1.0000 Cl- + 1.0000 Ba++ = BaCl+
-llnl_gamma 4.0
log_k -0.4977
-delta_H 11.142 kJ/mol # Calculated enthalpy of reaction BaCl+
# Enthalpy of formation: -165.77 kcal/mol
-analytic 1.1016e+002 4.2325e-002 -2.8039e+003 -4.6010e+001 -4.3785e+001
# -Range: 0-300
1.0000 F- + 1.0000 Ba++ = BaF+
-llnl_gamma 4.0
log_k -0.1833
-delta_H 8.95376 kJ/mol # Calculated enthalpy of reaction BaF+
# Enthalpy of formation: -206.51 kcal/mol
-analytic 1.0349e+002 4.0336e-002 -2.5195e+003 -4.3334e+001 -3.9346e+001
# -Range: 0-300
1.0000 NO3- + 1.0000 Ba++ = BaNO3+
-llnl_gamma 4.0
log_k +0.9000
-delta_H 0 # Not possible to calculate enthalpy of reaction BaNO3+
# Enthalpy of formation: -0 kcal/mol
1.0000 H2O + 1.0000 Ba++ = BaOH+ +1.0000 H+
-llnl_gamma 4.0
log_k -13.47
-delta_H 0 # Not possible to calculate enthalpy of reaction BaOH+
# Enthalpy of formation: -0 kcal/mol
2.0000 HAcetate + 1.0000 Be++ = Be(Acetate)2 +2.0000 H+
-llnl_gamma 3.0
log_k -6.8023
-delta_H -52.4255 kJ/mol # Calculated enthalpy of reaction Be(Acetate)2
# Enthalpy of formation: -336.23 kcal/mol
-analytic -3.5242e+001 5.1285e-003 -4.8914e+002 8.2862e+000 7.1774e+005
# -Range: 0-300
1.0000 Be++ + 1.0000 HAcetate = BeAcetate+ +1.0000 H+
-llnl_gamma 4.0
log_k -3.1079
-delta_H -22.761 kJ/mol # Calculated enthalpy of reaction BeAcetate+
# Enthalpy of formation: -213.04 kcal/mol
-analytic -1.9418e+001 5.2172e-004 -8.5071e+001 5.2755e+000 3.0215e+005
# -Range: 0-300
2.0000 H2O + 1.0000 Be++ = BeO2-- +4.0000 H+
-llnl_gamma 4.0
log_k -32.161
-delta_H 163.737 kJ/mol # Calculated enthalpy of reaction BeO2-2
# Enthalpy of formation: -189 kcal/mol
-analytic 7.0860e+000 -3.8474e-002 -1.1400e+004 4.2138e+000 -1.7789e+002
# -Range: 0-300
2.0000 H+ + 2.0000 Br- + 0.5000 O2 = Br2 +1.0000 H2O
-llnl_gamma 3.0
log_k +5.6834
-delta_H 0 # Not possible to calculate enthalpy of reaction Br2
# Enthalpy of formation: -0 kcal/mol
1.0000 HCO3- + 1.0000 H+ = CO2 +1.0000 H2O
-CO2_llnl_gamma
log_k +6.3447
-delta_H -9.7027 kJ/mol # Calculated enthalpy of reaction CO2
# Enthalpy of formation: -98.9 kcal/mol
-analytic -1.0534e+001 2.1746e-002 2.5216e+003 7.9125e-001 3.9351e+001
# -Range: 0-300
1.0000 HCO3- = CO3-- +1.0000 H+
-llnl_gamma 4.5
log_k -10.3288
-delta_H 14.6984 kJ/mol # Calculated enthalpy of reaction CO3-2
# Enthalpy of formation: -161.385 kcal/mol
-analytic -6.9958e+001 -3.3526e-002 -7.0846e+001 2.8224e+001 -1.0849e+000
# -Range: 0-300
2.0000 HAcetate + 1.0000 Ca++ = Ca(Acetate)2 +2.0000 H+
-llnl_gamma 3.0
log_k -7.3814
-delta_H -2.7196 kJ/mol # Calculated enthalpy of reaction Ca(Acetate)2
# Enthalpy of formation: -362.65 kcal/mol
-analytic -1.0320e+001 4.0012e-003 -3.6281e+003 2.4421e+000 7.0175e+005
# -Range: 0-300
1.0000 O_phthalate-2 + 1.0000 Ca++ = Ca(O_phthalate)
-llnl_gamma 3.0
log_k +2.4200
-delta_H 0 # Not possible to calculate enthalpy of reaction Ca(O_phthalate)
# Enthalpy of formation: -0 kcal/mol
1.0000 H2O + 1.0000 Ca++ + 1.0000 B(OH)3 = CaB(OH)4+ +1.0000 H+
-llnl_gamma 4.0
log_k -7.4222
-delta_H 0 # Not possible to calculate enthalpy of reaction CaB(OH)4+
# Enthalpy of formation: -0 kcal/mol
1.0000 Ca++ + 1.0000 HAcetate = CaAcetate+ +1.0000 H+
-llnl_gamma 4.0
log_k -3.8263
-delta_H 1.17152 kJ/mol # Calculated enthalpy of reaction CaAcetate+
# Enthalpy of formation: -245.62 kcal/mol
-analytic -8.8826e+000 3.1672e-003 -1.0764e+003 2.0526e+000 2.3599e+005
# -Range: 0-300
1.0000 HCO3- + 1.0000 Ca++ = CaCO3 +1.0000 H+
-llnl_gamma 3.0
log_k -7.0017
-delta_H 30.5767 kJ/mol # Calculated enthalpy of reaction CaCO3
# Enthalpy of formation: -287.39 kcal/mol
-analytic 2.3045e+002 5.5350e-002 -8.5056e+003 -9.1096e+001 -1.3279e+002
# -Range: 0-300
1.0000 Cl- + 1.0000 Ca++ = CaCl+
-llnl_gamma 4.0
log_k -0.6956
-delta_H 2.02087 kJ/mol # Calculated enthalpy of reaction CaCl+
# Enthalpy of formation: -169.25 kcal/mol
-analytic 8.1498e+001 3.8387e-002 -1.3763e+003 -3.5968e+001 -2.1501e+001
# -Range: 0-300
2.0000 Cl- + 1.0000 Ca++ = CaCl2
-llnl_gamma 3.0
log_k -0.6436
-delta_H -5.8325 kJ/mol # Calculated enthalpy of reaction CaCl2
# Enthalpy of formation: -211.06 kcal/mol
-analytic 1.8178e+002 7.6910e-002 -3.1088e+003 -7.8760e+001 -4.8563e+001
# -Range: 0-300
1.0000 F- + 1.0000 Ca++ = CaF+
-llnl_gamma 4.0
log_k +0.6817
-delta_H 5.6484 kJ/mol # Calculated enthalpy of reaction CaF+
# Enthalpy of formation: -208.6 kcal/mol
-analytic 7.8058e+001 3.8276e-002 -1.3289e+003 -3.4071e+001 -2.0759e+001
# -Range: 0-300
1.0000 HPO4-- + 1.0000 H+ + 1.0000 Ca++ = CaH2PO4+
-llnl_gamma 4.0
log_k +1.4000
-delta_H 0 # Not possible to calculate enthalpy of reaction CaH2PO4+
# Enthalpy of formation: -0 kcal/mol
1.0000 HCO3- + 1.0000 Ca++ = CaHCO3+
-llnl_gamma 4.0
log_k +1.0467
-delta_H 1.45603 kJ/mol # Calculated enthalpy of reaction CaHCO3+
# Enthalpy of formation: -294.35 kcal/mol
-analytic 5.5985e+001 3.4639e-002 -3.6972e+002 -2.5864e+001 -5.7859e+000
# -Range: 0-300
1.0000 HPO4-- + 1.0000 Ca++ = CaHPO4
-llnl_gamma 3.0
log_k +2.7400
-delta_H 0 # Not possible to calculate enthalpy of reaction CaHPO4
# Enthalpy of formation: -0 kcal/mol
1.0000 NO3- + 1.0000 Ca++ = CaNO3+
-llnl_gamma 4.0
log_k +0.7000
-delta_H 0 # Not possible to calculate enthalpy of reaction CaNO3+
# Enthalpy of formation: -0 kcal/mol
1.0000 H2O + 1.0000 Ca++ = CaOH+ +1.0000 H+
-llnl_gamma 4.0
log_k -12.85
-delta_H 0 # Not possible to calculate enthalpy of reaction CaOH+
# Enthalpy of formation: -0 kcal/mol
2.0000 HPO4-- + 1.0000 Ca++ = CaP2O7-- +1.0000 H2O
-llnl_gamma 4.0
log_k +3.0537
-delta_H 0 # Not possible to calculate enthalpy of reaction CaP2O7-2
# Enthalpy of formation: -0 kcal/mol
1.0000 HPO4-- + 1.0000 Ca++ = CaPO4- +1.0000 H+
-llnl_gamma 4.0
log_k -5.8618
-delta_H 0 # Not possible to calculate enthalpy of reaction CaPO4-
# Enthalpy of formation: -0 kcal/mol
1.0000 SO4-- + 1.0000 Ca++ = CaSO4
-llnl_gamma 3.0
log_k +2.1111
-delta_H 5.4392 kJ/mol # Calculated enthalpy of reaction CaSO4
# Enthalpy of formation: -345.9 kcal/mol
-analytic 2.8618e+002 8.4084e-002 -7.6880e+003 -1.1449e+002 -1.2005e+002
# -Range: 0-300
2.0000 HAcetate + 1.0000 Cd++ = Cd(Acetate)2 +2.0000 H+
-llnl_gamma 3.0
log_k -6.3625
-delta_H -17.4891 kJ/mol # Calculated enthalpy of reaction Cd(Acetate)2
# Enthalpy of formation: -254.52 kcal/mol
-analytic -1.9344e+001 2.5894e-003 -3.2847e+003 5.8489e+000 7.8041e+005
# -Range: 0-300
3.0000 HAcetate + 1.0000 Cd++ = Cd(Acetate)3- +3.0000 H+
-llnl_gamma 4.0
log_k -10.8558
-delta_H -40.0409 kJ/mol # Calculated enthalpy of reaction Cd(Acetate)3-
# Enthalpy of formation: -376.01 kcal/mol
-analytic 4.8290e+001 -3.4317e-003 -1.5122e+004 -1.3203e+001 2.2479e+006
# -Range: 0-300
4.0000 HAcetate + 1.0000 Cd++ = Cd(Acetate)4-- +4.0000 H+
-llnl_gamma 4.0
log_k -16.9163
-delta_H 0 # Not possible to calculate enthalpy of reaction Cd(Acetate)4-2
# Enthalpy of formation: -0 kcal/mol
2.0000 Cyanide- + 1.0000 Cd++ = Cd(Cyanide)2
-llnl_gamma 3.0
log_k +10.3551
-delta_H 0 # Not possible to calculate enthalpy of reaction Cd(Cyanide)2
# Enthalpy of formation: -0 kcal/mol
3.0000 Cyanide- + 1.0000 Cd++ = Cd(Cyanide)3-
-llnl_gamma 4.0
log_k +14.8191
-delta_H 0 # Not possible to calculate enthalpy of reaction Cd(Cyanide)3-
# Enthalpy of formation: -0 kcal/mol
4.0000 Cyanide- + 1.0000 Cd++ = Cd(Cyanide)4--
-llnl_gamma 4.0
log_k +18.2670
-delta_H 0 # Not possible to calculate enthalpy of reaction Cd(Cyanide)4-2
# Enthalpy of formation: -0 kcal/mol
2.0000 HCO3- + 1.0000 Cd++ = Cd(CO3)2-- +2.0000 H+
-llnl_gamma 4.0
log_k -14.2576
-delta_H 0 # Not possible to calculate enthalpy of reaction Cd(CO3)2-2
# Enthalpy of formation: -0 kcal/mol
2.0000 N3- + 1.0000 Cd++ = Cd(N3)2
-llnl_gamma 0.0
log_k +2.4606
-delta_H 0 # Not possible to calculate enthalpy of reaction Cd(N3)2
# Enthalpy of formation: -0 kcal/mol
3.0000 N3- + 1.0000 Cd++ = Cd(N3)3-
-llnl_gamma 4.0
log_k +3.1263
-delta_H 0 # Not possible to calculate enthalpy of reaction Cd(N3)3-
# Enthalpy of formation: -0 kcal/mol
4.0000 N3- + 1.0000 Cd++ = Cd(N3)4--
-llnl_gamma 4.0
log_k +3.4942
-delta_H 0 # Not possible to calculate enthalpy of reaction Cd(N3)4-2
# Enthalpy of formation: -0 kcal/mol
1.0000 NH3 + 1.0000 Cd++ = Cd(NH3)++
-llnl_gamma 4.5
log_k +2.5295
-delta_H 0 # Not possible to calculate enthalpy of reaction Cd(NH3)+2
# Enthalpy of formation: -0 kcal/mol
2.0000 NH3 + 1.0000 Cd++ = Cd(NH3)2++
-llnl_gamma 4.5
log_k +4.8760
-delta_H -27.6533 kJ/mol # Calculated enthalpy of reaction Cd(NH3)2+2
# Enthalpy of formation: -266.225 kJ/mol
-analytic 1.0738e+002 1.6071e-003 -3.2536e+003 -3.7202e+001 -5.0801e+001
# -Range: 0-300
4.0000 NH3 + 1.0000 Cd++ = Cd(NH3)4++
-llnl_gamma 4.5
log_k +7.2914
-delta_H -49.0684 kJ/mol # Calculated enthalpy of reaction Cd(NH3)4+2
# Enthalpy of formation: -450.314 kJ/mol
-analytic 1.5670e+002 -9.4949e-003 -5.0986e+003 -5.2316e+001 -7.9603e+001
# -Range: 0-300
2.0000 H2O + 1.0000 Cd++ = Cd(OH)2 +2.0000 H+
-llnl_gamma 3.0
log_k -20.3402
-delta_H 0 # Not possible to calculate enthalpy of reaction Cd(OH)2
# Enthalpy of formation: -0 kcal/mol
3.0000 H2O + 1.0000 Cd++ = Cd(OH)3- +3.0000 H+
-llnl_gamma 4.0
log_k -33.2852
-delta_H 0 # Not possible to calculate enthalpy of reaction Cd(OH)3-
# Enthalpy of formation: -0 kcal/mol
4.0000 H2O + 1.0000 Cd++ = Cd(OH)4-- +4.0000 H+
-llnl_gamma 4.0
log_k -47.3303
-delta_H 0 # Not possible to calculate enthalpy of reaction Cd(OH)4-2
# Enthalpy of formation: -0 kcal/mol
1.0000 H2O + 1.0000 Cl- + 1.0000 Cd++ = Cd(OH)Cl +1.0000 H+
-llnl_gamma 3.0
log_k -7.4328
-delta_H 0 # Not possible to calculate enthalpy of reaction Cd(OH)Cl
# Enthalpy of formation: -0 kcal/mol
2.0000 Thiocyanate- + 1.0000 Cd++ = Cd(Thiocyanate)2
-llnl_gamma 3.0
log_k +1.8649
-delta_H 0 # Not possible to calculate enthalpy of reaction Cd(Thiocyanate)2
# Enthalpy of formation: -0 kcal/mol
3.0000 Thiocyanate- + 1.0000 Cd++ = Cd(Thiocyanate)3-
-llnl_gamma 4.0
log_k +1.9000
-delta_H 0 # Not possible to calculate enthalpy of reaction Cd(Thiocyanate)3-
# Enthalpy of formation: -0 kcal/mol
2.0000 Cd++ + 1.0000 H2O = Cd2OH+++ +1.0000 H+
-llnl_gamma 5.0
log_k -9.3851
-delta_H 0 # Not possible to calculate enthalpy of reaction Cd2OH+3
# Enthalpy of formation: -0 kcal/mol
4.0000 H2O + 4.0000 Cd++ = Cd4(OH)4++++ +4.0000 H+
-llnl_gamma 5.5
log_k -362.1263
-delta_H 0 # Not possible to calculate enthalpy of reaction Cd4(OH)4+4
# Enthalpy of formation: -0 kcal/mol
1.0000 Cd++ + 1.0000 Br- = CdBr+
-llnl_gamma 4.0
log_k +2.1424
-delta_H -3.35588 kJ/mol # Calculated enthalpy of reaction CdBr+
# Enthalpy of formation: -200.757 kJ/mol
-analytic 1.4922e+002 5.0059e-002 -3.3035e+003 -6.0984e+001 -5.1593e+001
# -Range: 0-300
2.0000 Br- + 1.0000 Cd++ = CdBr2
-llnl_gamma 3.0
log_k +2.8614
-delta_H 0 # Not possible to calculate enthalpy of reaction CdBr2
# Enthalpy of formation: -0 kcal/mol
3.0000 Br- + 1.0000 Cd++ = CdBr3-
-llnl_gamma 4.0
log_k +3.0968
-delta_H 0 # Not possible to calculate enthalpy of reaction CdBr3-
# Enthalpy of formation: -0 kcal/mol
1.0000 Cd++ + 1.0000 HAcetate = CdAcetate+ +1.0000 H+
-llnl_gamma 4.0
log_k -2.8294
-delta_H -7.02912 kJ/mol # Calculated enthalpy of reaction CdAcetate+
# Enthalpy of formation: -135.92 kcal/mol
-analytic -8.8425e+000 1.7178e-003 -1.1758e+003 2.4435e+000 3.0321e+005
# -Range: 0-300
1.0000 Cd++ + 1.0000 Cyanide- = CdCyanide+
-llnl_gamma 4.0
log_k +5.3129
-delta_H 0 # Not possible to calculate enthalpy of reaction CdCyanide+
# Enthalpy of formation: -0 kcal/mol
1.0000 HCO3- + 1.0000 Cd++ = CdCO3 +1.0000 H+
-llnl_gamma 3.0
log_k -7.3288
-delta_H 0 # Not possible to calculate enthalpy of reaction CdCO3
# Enthalpy of formation: -0 kcal/mol
1.0000 Cl- + 1.0000 Cd++ = CdCl+
-llnl_gamma 4.0
log_k +2.7059
-delta_H 2.33843 kJ/mol # Calculated enthalpy of reaction CdCl+
# Enthalpy of formation: -240.639 kJ/mol
2.0000 Cl- + 1.0000 Cd++ = CdCl2
-llnl_gamma 3.0
log_k +3.3384
-delta_H 5.1261 kJ/mol # Calculated enthalpy of reaction CdCl2
# Enthalpy of formation: -404.931 kJ/mol
-analytic 1.4052e+002 4.9221e-002 -3.2625e+003 -5.6946e+001 -5.5451e+001
# -Range: 0-200
3.0000 Cl- + 1.0000 Cd++ = CdCl3-
-llnl_gamma 4.0
log_k +2.7112
-delta_H 15.9388 kJ/mol # Calculated enthalpy of reaction CdCl3-
# Enthalpy of formation: -561.198 kJ/mol
-analytic 3.5108e+002 1.0219e-001 -9.9103e+003 -1.3965e+002 -1.5474e+002
# -Range: 0-300
1.0000 HCO3- + 1.0000 Cd++ = CdHCO3+
-llnl_gamma 4.0
log_k +1.5000
-delta_H 0 # Not possible to calculate enthalpy of reaction CdHCO3+
# Enthalpy of formation: -0 kcal/mol
1.0000 I- + 1.0000 Cd++ = CdI+
-llnl_gamma 4.0
log_k +2.0710
-delta_H -9.02584 kJ/mol # Calculated enthalpy of reaction CdI+
# Enthalpy of formation: -141.826 kJ/mol
-analytic 1.5019e+002 5.0320e-002 -3.0810e+003 -6.1738e+001 -4.8120e+001
# -Range: 0-300
2.0000 I- + 1.0000 Cd++ = CdI2
-llnl_gamma 3.0
log_k +3.4685
-delta_H 0 # Not possible to calculate enthalpy of reaction CdI2
# Enthalpy of formation: -0 kcal/mol
3.0000 I- + 1.0000 Cd++ = CdI3-
-llnl_gamma 4.0
log_k +4.5506
-delta_H 0 # Not possible to calculate enthalpy of reaction CdI3-
# Enthalpy of formation: -0 kcal/mol
4.0000 I- + 1.0000 Cd++ = CdI4--
-llnl_gamma 4.0
log_k +5.3524
-delta_H -38.8566 kJ/mol # Calculated enthalpy of reaction CdI4-2
# Enthalpy of formation: -342.364 kJ/mol
-analytic 4.3154e+002 1.4257e-001 -8.4464e+003 -1.7795e+002 -1.3193e+002
# -Range: 0-300
1.0000 N3- + 1.0000 Cd++ = CdN3+
-llnl_gamma 4.0
log_k +1.4970
-delta_H 0 # Not possible to calculate enthalpy of reaction CdN3+
# Enthalpy of formation: -0 kcal/mol
1.0000 NO2- + 1.0000 Cd++ = CdNO2+
-llnl_gamma 4.0
log_k +2.3700
-delta_H 0 # Not possible to calculate enthalpy of reaction CdNO2+
# Enthalpy of formation: -0 kcal/mol
1.0000 H2O + 1.0000 Cd++ = CdOH+ +1.0000 H+
-llnl_gamma 4.0
log_k -10.0751
-delta_H 0 # Not possible to calculate enthalpy of reaction CdOH+
# Enthalpy of formation: -0 kcal/mol
2.0000 HPO4-- + 1.0000 Cd++ = CdP2O7-- +1.0000 H2O
-llnl_gamma 4.0
log_k +4.8094
-delta_H 0 # Not possible to calculate enthalpy of reaction CdP2O7-2
# Enthalpy of formation: -0 kcal/mol
1.0000 Thiocyanate- + 1.0000 Cd++ = CdThiocyanate+
-llnl_gamma 4.0
log_k +1.3218
-delta_H 0 # Not possible to calculate enthalpy of reaction CdThiocyanate+
# Enthalpy of formation: -0 kcal/mol
1.0000 SO4-- + 1.0000 Cd++ = CdSO4
-llnl_gamma 3.0
log_k +0.0028
-delta_H 0.20436 kJ/mol # Calculated enthalpy of reaction CdSO4
# Enthalpy of formation: -985.295 kJ/mol
-analytic -8.9926e+000 -1.9109e-003 2.7454e+002 3.4949e+000 4.6651e+000
# -Range: 0-200
1.0000 SeO4-- + 1.0000 Cd++ = CdSeO4
-llnl_gamma 3.0
log_k +2.2700
-delta_H 0 # Not possible to calculate enthalpy of reaction CdSeO4
# Enthalpy of formation: -0 kcal/mol
2.0000 HAcetate + 1.0000 Ce+++ = Ce(Acetate)2+ +2.0000 H+
-llnl_gamma 4.0
log_k -4.8159
-delta_H -22.9702 kJ/mol # Calculated enthalpy of reaction Ce(Acetate)2+
# Enthalpy of formation: -405.09 kcal/mol
-analytic -3.4653e+001 2.0716e-004 -6.3400e+002 1.0678e+001 4.8922e+005
# -Range: 0-300
3.0000 HAcetate + 1.0000 Ce+++ = Ce(Acetate)3 +3.0000 H+
-llnl_gamma 3.0
log_k -8.151
-delta_H -38.7438 kJ/mol # Calculated enthalpy of reaction Ce(Acetate)3
# Enthalpy of formation: -524.96 kcal/mol
-analytic -2.3361e+001 2.3896e-003 -1.8035e+003 5.0888e+000 7.1021e+005
# -Range: 0-300
2.0000 HCO3- + 1.0000 Ce+++ = Ce(CO3)2- +2.0000 H+
-llnl_gamma 4.0
log_k -8.1576
-delta_H 0 # Not possible to calculate enthalpy of reaction Ce(CO3)2-
# Enthalpy of formation: -0 kcal/mol
2.0000 HPO4-- + 1.0000 Ce+++ = Ce(HPO4)2-
-llnl_gamma 4.0
log_k +8.7000
-delta_H 0 # Not possible to calculate enthalpy of reaction Ce(HPO4)2-
# Enthalpy of formation: -0 kcal/mol
2.0000 H2O + 1.0000 Ce++++ = Ce(OH)2++ +2.0000 H+
-llnl_gamma 4.5
log_k +2.0098
-delta_H 0 # Not possible to calculate enthalpy of reaction Ce(OH)2+2
# Enthalpy of formation: -0 kcal/mol
2.0000 HPO4-- + 1.0000 Ce+++ = Ce(PO4)2--- +2.0000 H+
-llnl_gamma 4.0
log_k -6.1437
-delta_H 0 # Not possible to calculate enthalpy of reaction Ce(PO4)2-3
# Enthalpy of formation: -0 kcal/mol
2.0000 H2O + 2.0000 Ce++++ = Ce2(OH)2+6 +2.0000 H+
-llnl_gamma 6.0
log_k +3.0098
-delta_H 0 # Not possible to calculate enthalpy of reaction Ce2(OH)2+6
# Enthalpy of formation: -0 kcal/mol
5.0000 H2O + 3.0000 Ce+++ = Ce3(OH)5++++ +5.0000 H+
-llnl_gamma 5.5
log_k -33.4754
-delta_H 0 # Not possible to calculate enthalpy of reaction Ce3(OH)5+4
# Enthalpy of formation: -0 kcal/mol
1.0000 Ce+++ + 1.0000 Br- = CeBr++
-llnl_gamma 4.5
log_k +0.3797
-delta_H 3.0585 kJ/mol # Calculated enthalpy of reaction CeBr+2
# Enthalpy of formation: -195.709 kcal/mol
-analytic 7.5790e+001 3.6040e-002 -1.2647e+003 -3.3094e+001 -1.9757e+001
# -Range: 0-300
1.0000 Ce+++ + 1.0000 HAcetate = CeAcetate++ +1.0000 H+
-llnl_gamma 4.5
log_k -2.0304
-delta_H -12.0918 kJ/mol # Calculated enthalpy of reaction CeAcetate+2
# Enthalpy of formation: -286.39 kcal/mol
-analytic -1.6080e+001 6.6239e-004 -6.0721e+002 5.0845e+000 2.9512e+005
# -Range: 0-300
1.0000 HCO3- + 1.0000 Ce+++ = CeCO3+ +1.0000 H+
-llnl_gamma 4.0
log_k -2.9284
-delta_H 93.345 kJ/mol # Calculated enthalpy of reaction CeCO3+
# Enthalpy of formation: -309.988 kcal/mol
-analytic 2.3292e+002 5.3153e-002 -7.1180e+003 -9.2061e+001 -1.1114e+002
# -Range: 0-300
1.0000 Cl- + 1.0000 Ce+++ = CeCl++
-llnl_gamma 4.5
log_k +0.3086
-delta_H 14.7821 kJ/mol # Calculated enthalpy of reaction CeCl+2
# Enthalpy of formation: -203.8 kcal/mol
-analytic 8.3534e+001 3.8166e-002 -2.0058e+003 -3.5504e+001 -3.1324e+001
# -Range: 0-300
2.0000 Cl- + 1.0000 Ce+++ = CeCl2+
-llnl_gamma 4.0
log_k +0.0308
-delta_H 20.7777 kJ/mol # Calculated enthalpy of reaction CeCl2+
# Enthalpy of formation: -242.3 kcal/mol
-analytic 2.3011e+002 8.1428e-002 -6.1292e+003 -9.4468e+001 -9.5708e+001
# -Range: 0-300
3.0000 Cl- + 1.0000 Ce+++ = CeCl3
-llnl_gamma 3.0
log_k -0.3936
-delta_H 15.4766 kJ/mol # Calculated enthalpy of reaction CeCl3
# Enthalpy of formation: -283.5 kcal/mol
-analytic 4.4073e+002 1.2994e-001 -1.2308e+004 -1.7722e+002 -1.9218e+002
# -Range: 0-300
4.0000 Cl- + 1.0000 Ce+++ = CeCl4-
-llnl_gamma 4.0
log_k -0.7447
-delta_H -1.95811 kJ/mol # Calculated enthalpy of reaction CeCl4-
# Enthalpy of formation: -327.6 kcal/mol
-analytic 5.2230e+002 1.3490e-001 -1.4859e+004 -2.0747e+002 -2.3201e+002
# -Range: 0-300
1.0000 ClO4- + 1.0000 Ce+++ = CeClO4++
-llnl_gamma 4.5
log_k +1.9102
-delta_H -49.0197 kJ/mol # Calculated enthalpy of reaction CeClO4+2
# Enthalpy of formation: -210.026 kcal/mol
-analytic -1.3609e+001 1.8115e-002 3.9869e+003 -1.3033e+000 6.2215e+001
# -Range: 0-300
1.0000 F- + 1.0000 Ce+++ = CeF++
-llnl_gamma 4.5
log_k +4.2221
-delta_H 23.2212 kJ/mol # Calculated enthalpy of reaction CeF+2
# Enthalpy of formation: -242 kcal/mol
-analytic 1.0303e+002 4.1730e-002 -2.8424e+003 -4.1094e+001 -4.4383e+001
# -Range: 0-300
2.0000 F- + 1.0000 Ce+++ = CeF2+
-llnl_gamma 4.0
log_k +7.2714
-delta_H 15.0624 kJ/mol # Calculated enthalpy of reaction CeF2+
# Enthalpy of formation: -324.1 kcal/mol
-analytic 2.5063e+002 8.5224e-002 -6.2219e+003 -1.0017e+002 -9.7160e+001
# -Range: 0-300
3.0000 F- + 1.0000 Ce+++ = CeF3
-llnl_gamma 3.0
log_k +9.5144
-delta_H -6.0668 kJ/mol # Calculated enthalpy of reaction CeF3
# Enthalpy of formation: -409.3 kcal/mol
-analytic 4.6919e+002 1.3664e-001 -1.1745e+004 -1.8629e+002 -1.8340e+002
# -Range: 0-300
4.0000 F- + 1.0000 Ce+++ = CeF4-
-llnl_gamma 4.0
log_k +11.3909
-delta_H -45.6056 kJ/mol # Calculated enthalpy of reaction CeF4-
# Enthalpy of formation: -498.9 kcal/mol
-analytic 5.3522e+002 1.3856e-001 -1.2722e+004 -2.1112e+002 -1.9868e+002
# -Range: 0-300
1.0000 HPO4-- + 1.0000 H+ + 1.0000 Ce+++ = CeH2PO4++
-llnl_gamma 4.5
log_k +9.6684
-delta_H -16.2548 kJ/mol # Calculated enthalpy of reaction CeH2PO4+2
# Enthalpy of formation: -480.1 kcal/mol
-analytic 1.1338e+002 6.3771e-002 5.2908e+001 -4.9649e+001 7.9189e-001
# -Range: 0-300
1.0000 HCO3- + 1.0000 Ce+++ = CeHCO3++
-llnl_gamma 4.5
log_k +1.9190
-delta_H 8.77803 kJ/mol # Calculated enthalpy of reaction CeHCO3+2
# Enthalpy of formation: -330.2 kcal/mol
-analytic 4.4441e+001 3.2077e-002 -3.0714e+002 -2.0622e+001 -4.8060e+000
# -Range: 0-300
1.0000 HPO4-- + 1.0000 Ce+++ = CeHPO4+
-llnl_gamma 4.0
log_k +5.2000
-delta_H 0 # Not possible to calculate enthalpy of reaction CeHPO4+
# Enthalpy of formation: -0 kcal/mol
1.0000 IO3- + 1.0000 Ce+++ = CeIO3++
-llnl_gamma 4.5
log_k +1.9000
-delta_H -21.1627 kJ/mol # Calculated enthalpy of reaction CeIO3+2
# Enthalpy of formation: -225.358 kcal/mol
-analytic 3.3756e+001 2.8528e-002 1.2847e+003 -1.8042e+001 2.0036e+001
# -Range: 0-300
1.0000 NO3- + 1.0000 Ce+++ = CeNO3++
-llnl_gamma 4.5
log_k +1.3143
-delta_H -26.6563 kJ/mol # Calculated enthalpy of reaction CeNO3+2
# Enthalpy of formation: -223.2 kcal/mol
-analytic 2.2772e+001 2.5931e-002 1.9950e+003 -1.4490e+001 3.1124e+001
# -Range: 0-300
1.0000 H2O + 1.0000 Ce+++ = CeO+ +2.0000 H+
-llnl_gamma 4.0
log_k -16.4103
-delta_H 112.202 kJ/mol # Calculated enthalpy of reaction CeO+
# Enthalpy of formation: -208.9 kcal/mol
-analytic 1.9881e+002 3.1302e-002 -1.4331e+004 -7.1323e+001 -2.2368e+002
# -Range: 0-300
2.0000 H2O + 1.0000 Ce+++ = CeO2- +4.0000 H+
-llnl_gamma 4.0
log_k -38.758
-delta_H 308.503 kJ/mol # Calculated enthalpy of reaction CeO2-
# Enthalpy of formation: -230.3 kcal/mol
-analytic 1.0059e+002 3.4824e-003 -1.5873e+004 -3.3056e+001 -4.7656e+005
# -Range: 0-300
2.0000 H2O + 1.0000 Ce+++ = CeO2H +3.0000 H+
-llnl_gamma 3.0
log_k -26.1503
-delta_H 228.17 kJ/mol # Calculated enthalpy of reaction CeO2H
# Enthalpy of formation: -249.5 kcal/mol
-analytic 3.5650e+002 4.6708e-002 -2.4320e+004 -1.2731e+002 -3.7959e+002
# -Range: 0-300
1.0000 H2O + 1.0000 Ce+++ = CeOH++ +1.0000 H+
-llnl_gamma 4.5
log_k -8.4206
-delta_H 73.2911 kJ/mol # Calculated enthalpy of reaction CeOH+2
# Enthalpy of formation: -218.2 kcal/mol
-analytic 7.5809e+001 1.2863e-002 -6.7244e+003 -2.6473e+001 -1.0495e+002
# -Range: 0-300
1.0000 H2O + 1.0000 Ce++++ = CeOH+++ +1.0000 H+
-llnl_gamma 5.0
log_k +3.2049
-delta_H 0 # Not possible to calculate enthalpy of reaction CeOH+3
# Enthalpy of formation: -0 kcal/mol
1.0000 HPO4-- + 1.0000 Ce+++ = CePO4 +1.0000 H+
-llnl_gamma 3.0
log_k -0.9718
-delta_H 0 # Not possible to calculate enthalpy of reaction CePO4
# Enthalpy of formation: -0 kcal/mol
1.0000 SO4-- + 1.0000 Ce+++ = CeSO4+
-llnl_gamma 4.0
log_k -3.687
-delta_H 19.2464 kJ/mol # Calculated enthalpy of reaction CeSO4+
# Enthalpy of formation: -380.2 kcal/mol
-analytic 3.0156e+002 8.5149e-002 -1.1025e+004 -1.1866e+002 -1.7213e+002
# -Range: 0-300
2.0000 HAcetate + 1.0000 Co++ = Co(Acetate)2 +2.0000 H+
-llnl_gamma 3.0
log_k -7.1468
-delta_H -22.4262 kJ/mol # Calculated enthalpy of reaction Co(Acetate)2
# Enthalpy of formation: -251.46 kcal/mol
-analytic -2.0661e+001 2.9014e-003 -2.2146e+003 5.1702e+000 6.4968e+005
# -Range: 0-300
3.0000 HAcetate + 1.0000 Co++ = Co(Acetate)3- +3.0000 H+
-llnl_gamma 4.0
log_k -11.281
-delta_H -48.2415 kJ/mol # Calculated enthalpy of reaction Co(Acetate)3-
# Enthalpy of formation: -373.73 kcal/mol
-analytic 6.3384e+001 -4.0669e-003 -1.4715e+004 -1.9518e+001 2.1524e+006
# -Range: 0-300
2.0000 HS- + 1.0000 Co++ = Co(HS)2
-llnl_gamma 3.0
log_k +9.0306
-delta_H 0 # Not possible to calculate enthalpy of reaction Co(HS)2
# Enthalpy of formation: -0 kcal/mol
2.0000 H2O + 1.0000 Co++ = Co(OH)2 +2.0000 H+
-llnl_gamma 3.0
log_k -18.8
-delta_H 0 # Not possible to calculate enthalpy of reaction Co(OH)2
# Enthalpy of formation: -0 kcal/mol
4.0000 H2O + 1.0000 Co++ = Co(OH)4-- +4.0000 H+
-llnl_gamma 4.0
log_k -45.7803
-delta_H 0 # Not possible to calculate enthalpy of reaction Co(OH)4-2
# Enthalpy of formation: -0 kcal/mol
1.0000 H2O + 2.0000 Co++ = Co2OH+++ +1.0000 H+
-llnl_gamma 5.0
log_k -11.2
-delta_H 0 # Not possible to calculate enthalpy of reaction Co2OH+3
# Enthalpy of formation: -0 kcal/mol
4.0000 H2O + 4.0000 Co++ = Co4(OH)4++++ +4.0000 H+
-llnl_gamma 5.5
log_k -30.3803
-delta_H 0 # Not possible to calculate enthalpy of reaction Co4(OH)4+4
# Enthalpy of formation: -0 kcal/mol
2.0000 Br- + 1.0000 Co++ = CoBr2
-llnl_gamma 3.0
log_k -0.0358
-delta_H -0.56568 kJ/mol # Calculated enthalpy of reaction CoBr2
# Enthalpy of formation: -301.73 kJ/mol
-analytic 5.8731e+000 8.0908e-004 -1.8986e+002 -2.2295e+000 -3.2261e+000
# -Range: 0-200
1.0000 Co++ + 1.0000 HAcetate = CoAcetate+ +1.0000 H+
-llnl_gamma 4.0
log_k -3.2985
-delta_H -8.70272 kJ/mol # Calculated enthalpy of reaction CoAcetate+
# Enthalpy of formation: -132.08 kcal/mol
-analytic -5.4858e+000 1.9147e-003 -1.1292e+003 9.0555e-001 2.8223e+005
# -Range: 0-300
1.0000 Co++ + 1.0000 Cl- = CoCl+
-llnl_gamma 4.0
log_k +0.1547
-delta_H 1.71962 kJ/mol # Calculated enthalpy of reaction CoCl+
# Enthalpy of formation: -53.422 kcal/mol
-analytic 1.5234e+002 5.6958e-002 -3.3258e+003 -6.3849e+001 -5.1942e+001
# -Range: 0-300
1.0000 HS- + 1.0000 Co++ = CoHS+
-llnl_gamma 4.0
log_k +5.9813
-delta_H 0 # Not possible to calculate enthalpy of reaction CoHS+
# Enthalpy of formation: -0 kcal/mol
2.0000 I- + 1.0000 Co++ = CoI2
-llnl_gamma 3.0
log_k -0.0944
-delta_H 3.1774 kJ/mol # Calculated enthalpy of reaction CoI2
# Enthalpy of formation: -168.785 kJ/mol
-analytic 3.6029e+001 1.0128e-002 -1.1219e+003 -1.4301e+001 -1.9064e+001
# -Range: 0-200
1.0000 NO3- + 1.0000 Co++ = CoNO3+
-llnl_gamma 4.0
log_k +0.2000
-delta_H 0 # Not possible to calculate enthalpy of reaction CoNO3+
# Enthalpy of formation: -0 kcal/mol
1.0000 Co++ + S2O3-- = CoS2O3
-llnl_gamma 3.0
log_k 0.8063
-delta_H 0 # Not possible to calculate enthalpy of reaction CoS2O3
# Enthalpy of formation: -0 kcal/mol
1.0000 SO4-- + 1.0000 Co++ = CoSO4
-llnl_gamma 3.0
log_k +0.0436
-delta_H 0.3842 kJ/mol # Calculated enthalpy of reaction CoSO4
# Enthalpy of formation: -967.375 kJ/mol
-analytic 2.4606e+000 1.0086e-003 -6.1450e+001 -1.0148e+000 -1.0444e+000
# -Range: 0-200
1.0000 SeO4-- + 1.0000 Co++ = CoSeO4
-llnl_gamma 3.0
log_k +2.7000
-delta_H 0 # Not possible to calculate enthalpy of reaction CoSeO4
# Enthalpy of formation: -0 kcal/mol
2.0000 H2O + 1.0000 Cr+++ = Cr(OH)2+ +2.0000 H+
-llnl_gamma 4.0
log_k -9.7
-delta_H 0 # Not possible to calculate enthalpy of reaction Cr(OH)2+
# Enthalpy of formation: -0 kcal/mol
3.0000 H2O + 1.0000 Cr+++ = Cr(OH)3 +3.0000 H+
-llnl_gamma 3.0
log_k -18
-delta_H 0 # Not possible to calculate enthalpy of reaction Cr(OH)3
# Enthalpy of formation: -0 kcal/mol
4.0000 H2O + 1.0000 Cr+++ = Cr(OH)4- +4.0000 H+
-llnl_gamma 4.0
log_k -27.4
-delta_H 0 # Not possible to calculate enthalpy of reaction Cr(OH)4-
# Enthalpy of formation: -0 kcal/mol
2.0000 H2O + 2.0000 Cr+++ = Cr2(OH)2++++ +2.0000 H+
-llnl_gamma 5.5
log_k -5.06
-delta_H 0 # Not possible to calculate enthalpy of reaction Cr2(OH)2+4
# Enthalpy of formation: -0 kcal/mol
2.0000 H+ + 2.0000 CrO4-- = Cr2O7-- +1.0000 H2O
-llnl_gamma 4.0
log_k +14.5192
-delta_H -13.8783 kJ/mol # Calculated enthalpy of reaction Cr2O7-2
# Enthalpy of formation: -356.2 kcal/mol
-analytic 1.3749e+002 6.5773e-002 -7.9472e+002 -5.6525e+001 -1.2441e+001
# -Range: 0-300
4.0000 H2O + 3.0000 Cr+++ = Cr3(OH)4+5 +4.0000 H+
-llnl_gamma 6.0
log_k -8.15
-delta_H 0 # Not possible to calculate enthalpy of reaction Cr3(OH)4+5
# Enthalpy of formation: -0 kcal/mol
1.0000 Cr+++ + 1.0000 Br- = CrBr++
-llnl_gamma 4.5
log_k -2.7813
-delta_H 33.564 kJ/mol # Calculated enthalpy of reaction CrBr+2
# Enthalpy of formation: -78.018 kcal/mol
-analytic 9.4384e+001 3.4704e-002 -3.6750e+003 -3.8461e+001 -5.7373e+001
# -Range: 0-300
1.0000 Cr+++ + 1.0000 Cl- = CrCl++
-llnl_gamma 4.5
log_k -0.149
-delta_H 0 # Not possible to calculate enthalpy of reaction CrCl+2
# Enthalpy of formation: -0 kcal/mol
2.0000 Cl- + 1.0000 Cr+++ = CrCl2+
-llnl_gamma 4.0
log_k +0.1596
-delta_H 41.2919 kJ/mol # Calculated enthalpy of reaction CrCl2+
# Enthalpy of formation: -126.997 kcal/mol
-analytic 2.0114e+002 7.3878e-002 -6.2218e+003 -8.1677e+001 -9.7144e+001
# -Range: 0-300
1.0000 Cl- + 2.000 H+ + 1.0000 CrO4-- = CrO3Cl- + 1.0000 H2O
-llnl_gamma 4.0
log_k 7.5270
-delta_H 0 # Not possible to calculate enthalpy of reaction CrO3Cl-
# Enthalpy of formation: -0 kcal/mol
-analytic 2.7423e+002 1.0013e-001 -6.0072e+003 -1.1168e+002 -9.3817e+001
# -Range: 0-300
1.0000 H2O + 1.0000 Cr+++ = CrOH++ +1.0000 H+
-llnl_gamma 4.5
log_k -4
-delta_H 0 # Not possible to calculate enthalpy of reaction CrOH+2
# Enthalpy of formation: -0 kcal/mol
2.0000 HAcetate + 1.0000 Cs+ = Cs(Acetate)2- +2.0000 H+
-llnl_gamma 4.0
log_k -9.771
-delta_H 1.2552 kJ/mol # Calculated enthalpy of reaction Cs(Acetate)2-
# Enthalpy of formation: -293.57 kcal/mol
-analytic -1.6956e+002 -4.0378e-002 4.5773e+003 6.3241e+001 7.1475e+001
# -Range: 0-300
1.0000 Cs+ + 1.0000 Br- = CsBr
-llnl_gamma 3.0
log_k -0.2712
-delta_H 10.9621 kJ/mol # Calculated enthalpy of reaction CsBr
# Enthalpy of formation: -88.09 kcal/mol
-analytic 1.2064e+002 3.2000e-002 -3.8770e+003 -4.7458e+001 -6.0533e+001
# -Range: 0-300
1.0000 Cs+ + 1.0000 HAcetate = CsAcetate +1.0000 H+
-llnl_gamma 3.0
log_k -4.7352
-delta_H 6.0668 kJ/mol # Calculated enthalpy of reaction CsAcetate
# Enthalpy of formation: -176.32 kcal/mol
-analytic 2.4280e+001 -2.8642e-003 -3.1339e+003 -8.1616e+000 2.2684e+005
# -Range: 0-300
1.0000 Cs+ + 1.0000 Cl- = CsCl
-llnl_gamma 3.0
log_k -0.1385
-delta_H 2.73215 kJ/mol # Calculated enthalpy of reaction CsCl
# Enthalpy of formation: -100.95 kcal/mol
-analytic 1.2472e+002 3.3730e-002 -3.9130e+003 -4.9212e+001 -6.1096e+001
# -Range: 0-300
1.0000 I- + 1.0000 Cs+ = CsI
-llnl_gamma 3.0
log_k +0.2639
-delta_H -6.56888 kJ/mol # Calculated enthalpy of reaction CsI
# Enthalpy of formation: -76.84 kcal/mol
-analytic 1.1555e+002 3.1419e-002 -3.3496e+003 -4.5828e+001 -5.2302e+001
# -Range: 0-300
2.0000 HAcetate + 1.0000 Cu++ = Cu(Acetate)2 +2.0000 H+
-llnl_gamma 3.0
log_k -5.8824
-delta_H -25.899 kJ/mol # Calculated enthalpy of reaction Cu(Acetate)2
# Enthalpy of formation: -222.69 kcal/mol
-analytic -2.6689e+001 1.8048e-003 -1.8244e+003 7.7008e+000 6.5408e+005
# -Range: 0-300
2.0000 HAcetate + 1.0000 Cu+ = Cu(Acetate)2- +2.0000 H+
-llnl_gamma 4.0
log_k -9.2139
-delta_H -19.5476 kJ/mol # Calculated enthalpy of reaction Cu(Acetate)2-
# Enthalpy of formation: -219.74 kcal/mol
-analytic -3.2712e+002 -5.9087e-002 1.1386e+004 1.2017e+002 1.7777e+002
# -Range: 0-300
3.0000 HAcetate + 1.0000 Cu++ = Cu(Acetate)3- +3.0000 H+
-llnl_gamma 4.0
log_k -9.3788
-delta_H -53.2205 kJ/mol # Calculated enthalpy of reaction Cu(Acetate)3-
# Enthalpy of formation: -345.32 kcal/mol
-analytic 3.9475e+001 -6.2867e-003 -1.3233e+004 -1.0643e+001 2.1121e+006
# -Range: 0-300
2.0000 HCO3- + 1.0000 Cu++ = Cu(CO3)2-- +2.0000 H+
-llnl_gamma 4.0
log_k -10.4757
-delta_H 0 # Not possible to calculate enthalpy of reaction Cu(CO3)2-2
# Enthalpy of formation: -0 kcal/mol
2.0000 NH3 + 1.0000 Cu++ = Cu(NH3)2++
-llnl_gamma 4.5
log_k +7.4512
-delta_H -45.1269 kJ/mol # Calculated enthalpy of reaction Cu(NH3)2+2
# Enthalpy of formation: -142.112 kJ/mol
-analytic 1.1526e+002 4.8192e-003 -2.5139e+003 -4.0733e+001 -3.9261e+001
# -Range: 0-300
3.0000 NH3 + 1.0000 Cu++ = Cu(NH3)3++
-llnl_gamma 4.5
log_k +10.2719
-delta_H -67.2779 kJ/mol # Calculated enthalpy of reaction Cu(NH3)3+2
# Enthalpy of formation: -245.6 kJ/mol
-analytic 1.3945e+002 -3.8236e-004 -2.8137e+003 -4.8336e+001 -4.3946e+001
# -Range: 0-300
2.0000 NO2- + 1.0000 Cu++ = Cu(NO2)2
-llnl_gamma 3.0
log_k +3.0300
-delta_H 0 # Not possible to calculate enthalpy of reaction Cu(NO2)2
# Enthalpy of formation: -0 kcal/mol
1.0000 Cu+ + 1.0000 HAcetate = CuAcetate +1.0000 H+
-llnl_gamma 3.0
log_k -4.4274
-delta_H -4.19237 kJ/mol # Calculated enthalpy of reaction CuAcetate
# Enthalpy of formation: -99.97 kcal/mol
-analytic 6.3784e+000 -4.5464e-004 -1.9995e+003 -2.8359e+000 2.7224e+005
# -Range: 0-300
1.0000 Cu++ + 1.0000 HAcetate = CuAcetate+ +1.0000 H+
-llnl_gamma 4.0
log_k -2.5252
-delta_H -11.3805 kJ/mol # Calculated enthalpy of reaction CuAcetate+
# Enthalpy of formation: -103.12 kcal/mol
-analytic -1.4930e+001 5.1278e-004 -3.4874e+002 4.3605e+000 2.3504e+005
# -Range: 0-300
2.0000 H2O + 1.0000 HCO3- + 1.0000 Cu++ = CuCO3(OH)2-- +3.0000 H+
-llnl_gamma 4.0
log_k -23.444
-delta_H 0 # Not possible to calculate enthalpy of reaction CuCO3(OH)2-2
# Enthalpy of formation: -0 kcal/mol
1.0000 HCO3- + 1.0000 Cu++ = CuCO3 +1.0000 H+
-llnl_gamma 3.0
log_k -3.3735
-delta_H 0 # Not possible to calculate enthalpy of reaction CuCO3
# Enthalpy of formation: -0 kcal/mol
1.0000 Cu++ + 1.0000 Cl- = CuCl+
-llnl_gamma 4.0
log_k +0.4370
-delta_H 0 # Not possible to calculate enthalpy of reaction CuCl+
# Enthalpy of formation: -0 kcal/mol
2.0000 Cl- + 1.0000 Cu++ = CuCl2
-llnl_gamma 3.0
log_k +0.1585
-delta_H 0 # Not possible to calculate enthalpy of reaction CuCl2
# Enthalpy of formation: -0 kcal/mol
2.0000 Cl- + 1.0000 Cu+ = CuCl2-
-llnl_gamma 4.0
log_k +4.8212
-delta_H 0 # Not possible to calculate enthalpy of reaction CuCl2-
# Enthalpy of formation: -0 kcal/mol
3.0000 Cl- + 1.0000 Cu+ = CuCl3--
-llnl_gamma 4.0
log_k +5.6289
-delta_H 0 # Not possible to calculate enthalpy of reaction CuCl3-2
# Enthalpy of formation: -0 kcal/mol
4.0000 Cl- + 1.0000 Cu++ = CuCl4--
-llnl_gamma 4.0
log_k -4.5681
-delta_H 0 # Not possible to calculate enthalpy of reaction CuCl4-2
# Enthalpy of formation: -0 kcal/mol
1.0000 F- + 1.0000 Cu++ = CuF+
-llnl_gamma 4.0
log_k +1.2000
-delta_H 0 # Not possible to calculate enthalpy of reaction CuF+
# Enthalpy of formation: -0 kcal/mol
1.0000 HPO4-- + 1.0000 H+ + 1.0000 Cu++ = CuH2PO4+
-llnl_gamma 4.0
log_k +8.9654
-delta_H 0 # Not possible to calculate enthalpy of reaction CuH2PO4+
# Enthalpy of formation: -0 kcal/mol
1.0000 HPO4-- + 1.0000 Cu++ = CuHPO4
-llnl_gamma 3.0
log_k +4.0600
-delta_H 0 # Not possible to calculate enthalpy of reaction CuHPO4
# Enthalpy of formation: -0 kcal/mol
1.0000 NH3 + 1.0000 Cu++ = CuNH3++
-llnl_gamma 4.5
log_k +4.0400
-delta_H 0 # Not possible to calculate enthalpy of reaction CuNH3+2
# Enthalpy of formation: -0 kcal/mol
1.0000 NO2- + 1.0000 Cu++ = CuNO2+
-llnl_gamma 4.0
log_k +2.0200
-delta_H 0 # Not possible to calculate enthalpy of reaction CuNO2+
# Enthalpy of formation: -0 kcal/mol
2.0000 H2O + 1.0000 Cu++ = CuO2-- +4.0000 H+
-llnl_gamma 4.0
log_k -39.4497
-delta_H 0 # Not possible to calculate enthalpy of reaction CuO2-2
# Enthalpy of formation: -0 kcal/mol
1.0000 H2O + 1.0000 Cu++ = CuOH+ +1.0000 H+
-llnl_gamma 4.0
log_k -7.2875
-delta_H 0 # Not possible to calculate enthalpy of reaction CuOH+
# Enthalpy of formation: -0 kcal/mol
1.0000 HPO4-- + 1.0000 Cu++ = CuPO4- +1.0000 H+
-llnl_gamma 4.0
log_k -2.4718
-delta_H 0 # Not possible to calculate enthalpy of reaction CuPO4-
# Enthalpy of formation: -0 kcal/mol
1.0000 SO4-- + 1.0000 Cu++ = CuSO4
-llnl_gamma 0.0
log_k +2.3600
-delta_H 0 # Not possible to calculate enthalpy of reaction CuSO4
# Enthalpy of formation: -0 kcal/mol
2.0000 HAcetate + 1.0000 Dy+++ = Dy(Acetate)2+ +2.0000 H+
-llnl_gamma 4.0
log_k -4.9625
-delta_H -29.3298 kJ/mol # Calculated enthalpy of reaction Dy(Acetate)2+
# Enthalpy of formation: -405.71 kcal/mol
-analytic -2.7249e+001 2.7507e-003 -1.7500e+003 7.9356e+000 6.8668e+005
# -Range: 0-300
3.0000 HAcetate + 1.0000 Dy+++ = Dy(Acetate)3 +3.0000 H+
-llnl_gamma 3.0
log_k -8.3489
-delta_H -49.4549 kJ/mol # Calculated enthalpy of reaction Dy(Acetate)3
# Enthalpy of formation: -526.62 kcal/mol
-analytic -2.4199e+001 6.2065e-003 -2.8937e+003 5.0176e+000 1.0069e+006
# -Range: 0-300
2.0000 HCO3- + 1.0000 Dy+++ = Dy(CO3)2- +2.0000 H+
-llnl_gamma 4.0
log_k -7.4576
-delta_H 0 # Not possible to calculate enthalpy of reaction Dy(CO3)2-
# Enthalpy of formation: -0 kcal/mol
2.0000 HPO4-- + 1.0000 Dy+++ = Dy(HPO4)2-
-llnl_gamma 4.0
log_k +9.8000
-delta_H 0 # Not possible to calculate enthalpy of reaction Dy(HPO4)2-
# Enthalpy of formation: -0 kcal/mol
# Redundant with DyO2-
#4.0000 H2O + 1.0000 Dy+++ = Dy(OH)4- +4.0000 H+
# -llnl_gamma 4.0
# log_k -33.4803
# -delta_H 0 # Not possible to calculate enthalpy of reaction Dy(OH)4-
# Enthalpy of formation: -0 kcal/mol
2.0000 HPO4-- + 1.0000 Dy+++ = Dy(PO4)2--- +2.0000 H+
-llnl_gamma 4.0
log_k -3.4437
-delta_H 0 # Not possible to calculate enthalpy of reaction Dy(PO4)2-3
# Enthalpy of formation: -0 kcal/mol
2.0000 SO4-- + 1.0000 Dy+++ = Dy(SO4)2-
-llnl_gamma 4.0
log_k +5.0000
-delta_H 0 # Not possible to calculate enthalpy of reaction Dy(SO4)2-
# Enthalpy of formation: -0 kcal/mol
1.0000 Dy+++ + 1.0000 HAcetate = DyAcetate++ +1.0000 H+
-llnl_gamma 4.5
log_k -2.1037
-delta_H -14.8532 kJ/mol # Calculated enthalpy of reaction DyAcetate+2
# Enthalpy of formation: -286.15 kcal/mol
-analytic -1.3635e+001 1.7329e-003 -9.4636e+002 4.0900e+000 3.6282e+005
# -Range: 0-300
1.0000 HCO3- + 1.0000 Dy+++ = DyCO3+ +1.0000 H+
-llnl_gamma 4.0
log_k -2.3324
-delta_H 89.1108 kJ/mol # Calculated enthalpy of reaction DyCO3+
# Enthalpy of formation: -310.1 kcal/mol
-analytic 2.3742e+002 5.4342e-002 -6.9953e+003 -9.3949e+001 -1.0922e+002
# -Range: 0-300
1.0000 Dy+++ + 1.0000 Cl- = DyCl++
-llnl_gamma 4.5
log_k +0.2353
-delta_H 13.5269 kJ/mol # Calculated enthalpy of reaction DyCl+2
# Enthalpy of formation: -203.2 kcal/mol
-analytic 6.9134e+001 3.7129e-002 -1.3839e+003 -3.0432e+001 -2.1615e+001
# -Range: 0-300
2.0000 Cl- + 1.0000 Dy+++ = DyCl2+
-llnl_gamma 4.0
log_k -0.0425
-delta_H 17.4305 kJ/mol # Calculated enthalpy of reaction DyCl2+
# Enthalpy of formation: -242.2 kcal/mol
-analytic 1.8868e+002 7.7901e-002 -4.3528e+003 -7.9735e+001 -6.7978e+001
# -Range: 0-300
3.0000 Cl- + 1.0000 Dy+++ = DyCl3
-llnl_gamma 3.0
log_k -0.4669
-delta_H 8.78222 kJ/mol # Calculated enthalpy of reaction DyCl3
# Enthalpy of formation: -284.2 kcal/mol
-analytic 3.6761e+002 1.2471e-001 -9.0651e+003 -1.5147e+002 -1.4156e+002
# -Range: 0-300
4.0000 Cl- + 1.0000 Dy+++ = DyCl4-
-llnl_gamma 4.0
log_k -0.8913
-delta_H -14.0917 kJ/mol # Calculated enthalpy of reaction DyCl4-
# Enthalpy of formation: -329.6 kcal/mol
-analytic 3.9134e+002 1.2288e-001 -9.2351e+003 -1.6078e+002 -1.4422e+002
# -Range: 0-300
1.0000 F- + 1.0000 Dy+++ = DyF++
-llnl_gamma 4.5
log_k +4.6619
-delta_H 23.2212 kJ/mol # Calculated enthalpy of reaction DyF+2
# Enthalpy of formation: -241.1 kcal/mol
-analytic 9.1120e+001 4.1193e-002 -2.3302e+003 -3.6734e+001 -3.6388e+001
# -Range: 0-300
2.0000 F- + 1.0000 Dy+++ = DyF2+
-llnl_gamma 4.0
log_k +8.1510
-delta_H 12.552 kJ/mol # Calculated enthalpy of reaction DyF2+
# Enthalpy of formation: -323.8 kcal/mol
-analytic 2.1325e+002 8.2483e-002 -4.5864e+003 -8.6587e+001 -7.1629e+001
# -Range: 0-300
3.0000 F- + 1.0000 Dy+++ = DyF3
-llnl_gamma 3.0
log_k +10.7605
-delta_H -11.9244 kJ/mol # Calculated enthalpy of reaction DyF3
# Enthalpy of formation: -409.8 kcal/mol
-analytic 3.9766e+002 1.3143e-001 -8.5607e+003 -1.6056e+002 -1.3370e+002
# -Range: 0-300
4.0000 F- + 1.0000 Dy+++ = DyF4-
-llnl_gamma 4.0
log_k +12.8569
-delta_H -57.3208 kJ/mol # Calculated enthalpy of reaction DyF4-
# Enthalpy of formation: -500.8 kcal/mol
-analytic 4.1672e+002 1.2922e-001 -7.4445e+003 -1.6867e+002 -1.1629e+002
# -Range: 0-300
1.0000 HPO4-- + 1.0000 H+ + 1.0000 Dy+++ = DyH2PO4++
-llnl_gamma 4.5
log_k +9.3751
-delta_H -18.3468 kJ/mol # Calculated enthalpy of reaction DyH2PO4+2
# Enthalpy of formation: -479.7 kcal/mol
-analytic 9.8183e+001 6.2578e-002 7.1784e+002 -4.4383e+001 1.1172e+001
# -Range: 0-300
1.0000 HCO3- + 1.0000 Dy+++ = DyHCO3++
-llnl_gamma 4.5
log_k +1.6991
-delta_H 7.10443 kJ/mol # Calculated enthalpy of reaction DyHCO3+2
# Enthalpy of formation: -329.7 kcal/mol
-analytic 2.8465e+001 3.0703e-002 3.9229e+002 -1.5036e+001 6.1127e+000
# -Range: 0-300
1.0000 HPO4-- + 1.0000 Dy+++ = DyHPO4+
-llnl_gamma 4.0
log_k +5.8000
-delta_H 0 # Not possible to calculate enthalpy of reaction DyHPO4+
# Enthalpy of formation: -0 kcal/mol
1.0000 NO3- + 1.0000 Dy+++ = DyNO3++
-llnl_gamma 4.5
log_k +0.1415
-delta_H -30.4219 kJ/mol # Calculated enthalpy of reaction DyNO3+2
# Enthalpy of formation: -223.2 kcal/mol
-analytic 6.4353e+000 2.4556e-002 2.5866e+003 -8.9975e+000 4.0359e+001
# -Range: 0-300
1.0000 H2O + 1.0000 Dy+++ = DyO+ +2.0000 H+
-llnl_gamma 4.0
log_k -16.1171
-delta_H 108.018 kJ/mol # Calculated enthalpy of reaction DyO+
# Enthalpy of formation: -209 kcal/mol
-analytic 1.9069e+002 3.0358e-002 -1.3796e+004 -6.8532e+001 -2.1532e+002
# -Range: 0-300
2.0000 H2O + 1.0000 Dy+++ = DyO2- +4.0000 H+
-llnl_gamma 4.0
log_k -33.4804
-delta_H 273.776 kJ/mol # Calculated enthalpy of reaction DyO2-
# Enthalpy of formation: -237.7 kcal/mol
-analytic 7.7395e+001 4.4204e-004 -1.3570e+004 -2.4546e+001 -4.2320e+005
# -Range: 0-300
2.0000 H2O + 1.0000 Dy+++ = DyO2H +3.0000 H+
-llnl_gamma 3.0
log_k -24.8309
-delta_H 217.71 kJ/mol # Calculated enthalpy of reaction DyO2H
# Enthalpy of formation: -251.1 kcal/mol
-analytic 3.3576e+002 4.6004e-002 -2.2868e+004 -1.2027e+002 -3.5693e+002
# -Range: 0-300
1.0000 H2O + 1.0000 Dy+++ = DyOH++ +1.0000 H+
-llnl_gamma 4.5
log_k -7.8342
-delta_H 76.6383 kJ/mol # Calculated enthalpy of reaction DyOH+2
# Enthalpy of formation: -216.5 kcal/mol
-analytic 7.0856e+001 1.2473e-002 -6.2419e+003 -2.4841e+001 -9.7420e+001
# -Range: 0-300
1.0000 HPO4-- + 1.0000 Dy+++ = DyPO4 +1.0000 H+
-llnl_gamma 3.0
log_k +0.1782
-delta_H 0 # Not possible to calculate enthalpy of reaction DyPO4
# Enthalpy of formation: -0 kcal/mol
1.0000 SO4-- + 1.0000 Dy+++ = DySO4+
-llnl_gamma 4.0
log_k +3.6430
-delta_H 20.5016 kJ/mol # Calculated enthalpy of reaction DySO4+
# Enthalpy of formation: -379 kcal/mol
-analytic 3.0672e+002 8.6459e-002 -9.0386e+003 -1.2063e+002 -1.4113e+002
# -Range: 0-300
2.0000 HAcetate + 1.0000 Er+++ = Er(Acetate)2+ +2.0000 H+
-llnl_gamma 4.0
log_k -4.9844
-delta_H -32.8026 kJ/mol # Calculated enthalpy of reaction Er(Acetate)2+
# Enthalpy of formation: -408.54 kcal/mol
-analytic -3.1458e+001 1.4715e-003 -1.0556e+003 9.1586e+000 6.1669e+005
# -Range: 0-300
3.0000 HAcetate + 1.0000 Er+++ = Er(Acetate)3 +3.0000 H+
-llnl_gamma 3.0
log_k -8.3783
-delta_H -55.187 kJ/mol # Calculated enthalpy of reaction Er(Acetate)3
# Enthalpy of formation: -529.99 kcal/mol
-analytic -2.1575e+001 5.9740e-003 -2.0489e+003 3.3624e+000 8.8933e+005
# -Range: 0-300
2.0000 HCO3- + 1.0000 Er+++ = Er(CO3)2- +2.0000 H+
-llnl_gamma 4.0
log_k -7.2576
-delta_H 0 # Not possible to calculate enthalpy of reaction Er(CO3)2-
# Enthalpy of formation: -0 kcal/mol
2.0000 HPO4-- + 1.0000 Er+++ = Er(HPO4)2-
-llnl_gamma 4.0
log_k +10.0000
-delta_H 0 # Not possible to calculate enthalpy of reaction Er(HPO4)2-
# Enthalpy of formation: -0 kcal/mol
# Redundant with ErO2-
#4.0000 H2O + 1.0000 Er+++ = Er(OH)4- +4.0000 H+
# -llnl_gamma 4.0
# log_k -32.5803
# -delta_H 0 # Not possible to calculate enthalpy of reaction Er(OH)4-
# Enthalpy of formation: -0 kcal/mol
2.0000 HPO4-- + 1.0000 Er+++ = Er(PO4)2--- +2.0000 H+
-llnl_gamma 4.0
log_k -3.2437
-delta_H 0 # Not possible to calculate enthalpy of reaction Er(PO4)2-3
# Enthalpy of formation: -0 kcal/mol
2.0000 SO4-- + 1.0000 Er+++ = Er(SO4)2-
-llnl_gamma 4.0
log_k +5.0000
-delta_H 0 # Not possible to calculate enthalpy of reaction Er(SO4)2-
# Enthalpy of formation: -0 kcal/mol
1.0000 Er+++ + 1.0000 HAcetate = ErAcetate++ +1.0000 H+
-llnl_gamma 4.5
log_k -2.1184
-delta_H -16.4013 kJ/mol # Calculated enthalpy of reaction ErAcetate+2
# Enthalpy of formation: -288.52 kcal/mol
-analytic -1.2519e+001 1.5558e-003 -8.5344e+002 3.5918e+000 3.4888e+005
# -Range: 0-300
1.0000 HCO3- + 1.0000 Er+++ = ErCO3+ +1.0000 H+
-llnl_gamma 4.0
log_k -2.1858
-delta_H 87.0188 kJ/mol # Calculated enthalpy of reaction ErCO3+
# Enthalpy of formation: -312.6 kcal/mol
-analytic 2.3838e+002 5.4549e-002 -6.9433e+003 -9.4373e+001 -1.0841e+002
# -Range: 0-300
1.0000 Er+++ + 1.0000 Cl- = ErCl++
-llnl_gamma 4.5
log_k +0.3086
-delta_H 12.6901 kJ/mol # Calculated enthalpy of reaction ErCl+2
# Enthalpy of formation: -205.4 kcal/mol
-analytic 7.4113e+001 3.7462e-002 -1.5300e+003 -3.2257e+001 -2.3896e+001
# -Range: 0-300
2.0000 Cl- + 1.0000 Er+++ = ErCl2+
-llnl_gamma 4.0
log_k -0.0425
-delta_H 15.3385 kJ/mol # Calculated enthalpy of reaction ErCl2+
# Enthalpy of formation: -244.7 kcal/mol
-analytic 2.0259e+002 7.8907e-002 -4.8271e+003 -8.4835e+001 -7.5382e+001
# -Range: 0-300
3.0000 Cl- + 1.0000 Er+++ = ErCl3
-llnl_gamma 3.0
log_k -0.4669
-delta_H 5.01662 kJ/mol # Calculated enthalpy of reaction ErCl3
# Enthalpy of formation: -287.1 kcal/mol
-analytic 3.9721e+002 1.2757e-001 -1.0045e+004 -1.6244e+002 -1.5686e+002
# -Range: 0-300
4.0000 Cl- + 1.0000 Er+++ = ErCl4-
-llnl_gamma 4.0
log_k -0.8913
-delta_H -20.7861 kJ/mol # Calculated enthalpy of reaction ErCl4-
# Enthalpy of formation: -333.2 kcal/mol
-analytic 4.3471e+002 1.2627e-001 -1.0669e+004 -1.7677e+002 -1.6660e+002
# -Range: 0-300
1.0000 F- + 1.0000 Er+++ = ErF++
-llnl_gamma 4.5
log_k +4.7352
-delta_H 24.058 kJ/mol # Calculated enthalpy of reaction ErF+2
# Enthalpy of formation: -242.9 kcal/mol
-analytic 9.7079e+001 4.1707e-002 -2.6028e+003 -3.8805e+001 -4.0643e+001
# -Range: 0-300
2.0000 F- + 1.0000 Er+++ = ErF2+
-llnl_gamma 4.0
log_k +8.2976
-delta_H 12.9704 kJ/mol # Calculated enthalpy of reaction ErF2+
# Enthalpy of formation: -325.7 kcal/mol
-analytic 2.2892e+002 8.3842e-002 -5.2174e+003 -9.2172e+001 -8.1481e+001
# -Range: 0-300
3.0000 F- + 1.0000 Er+++ = ErF3
-llnl_gamma 3.0
log_k +10.9071
-delta_H -12.3428 kJ/mol # Calculated enthalpy of reaction ErF3
# Enthalpy of formation: -411.9 kcal/mol
-analytic 4.2782e+002 1.3425e-001 -9.7064e+003 -1.7148e+002 -1.5158e+002
# -Range: 0-300
4.0000 F- + 1.0000 Er+++ = ErF4-
-llnl_gamma 4.0
log_k +13.0768
-delta_H -60.2496 kJ/mol # Calculated enthalpy of reaction ErF4-
# Enthalpy of formation: -503.5 kcal/mol
-analytic 4.6524e+002 1.3372e-001 -9.1895e+003 -1.8636e+002 -1.4353e+002
# -Range: 0-300
1.0000 HPO4-- + 1.0000 H+ + 1.0000 Er+++ = ErH2PO4++
-llnl_gamma 4.5
log_k +9.4484
-delta_H -20.4388 kJ/mol # Calculated enthalpy of reaction ErH2PO4+2
# Enthalpy of formation: -482.2 kcal/mol
-analytic 1.0254e+002 6.2786e-002 6.3590e+002 -4.6029e+001 9.8920e+000
# -Range: 0-300
1.0000 HCO3- + 1.0000 Er+++ = ErHCO3++
-llnl_gamma 4.5
log_k +1.7724
-delta_H 5.01243 kJ/mol # Calculated enthalpy of reaction ErHCO3+2
# Enthalpy of formation: -332.2 kcal/mol
-analytic 3.2450e+001 3.0822e-002 3.1601e+002 -1.6528e+001 4.9212e+000
# -Range: 0-300
1.0000 HPO4-- + 1.0000 Er+++ = ErHPO4+
-llnl_gamma 4.0
log_k +5.9000
-delta_H 0 # Not possible to calculate enthalpy of reaction ErHPO4+
# Enthalpy of formation: -0 kcal/mol
1.0000 NO3- + 1.0000 Er+++ = ErNO3++
-llnl_gamma 4.5
log_k +0.1415
-delta_H -33.7691 kJ/mol # Calculated enthalpy of reaction ErNO3+2
# Enthalpy of formation: -226 kcal/mol
-analytic 1.0381e+001 2.4710e-002 2.5752e+003 -1.0596e+001 4.0181e+001
# -Range: 0-300
1.0000 H2O + 1.0000 Er+++ = ErO+ +2.0000 H+
-llnl_gamma 4.0
log_k -15.9705
-delta_H 105.508 kJ/mol # Calculated enthalpy of reaction ErO+
# Enthalpy of formation: -211.6 kcal/mol
-analytic 1.7556e+002 2.8655e-002 -1.3134e+004 -6.3050e+001 -2.0499e+002
# -Range: 0-300
2.0000 H2O + 1.0000 Er+++ = ErO2- +4.0000 H+
-llnl_gamma 4.0
log_k -32.6008
-delta_H 266.245 kJ/mol # Calculated enthalpy of reaction ErO2-
# Enthalpy of formation: -241.5 kcal/mol
-analytic 1.4987e+002 9.1241e-003 -1.8521e+004 -4.9740e+001 -2.8905e+002
# -Range: 0-300
2.0000 H2O + 1.0000 Er+++ = ErO2H +3.0000 H+
-llnl_gamma 3.0
log_k -24.3178
-delta_H 212.689 kJ/mol # Calculated enthalpy of reaction ErO2H
# Enthalpy of formation: -254.3 kcal/mol
-analytic 3.1493e+002 4.4381e-002 -2.1821e+004 -1.1287e+002 -3.4059e+002
# -Range: 0-300
1.0000 H2O + 1.0000 Er+++ = ErOH++ +1.0000 H+
-llnl_gamma 4.5
log_k -7.7609
-delta_H 74.5463 kJ/mol # Calculated enthalpy of reaction ErOH+2
# Enthalpy of formation: -219 kcal/mol
-analytic 5.7142e+001 1.0986e-002 -5.6684e+003 -1.9867e+001 -8.8467e+001
# -Range: 0-300
1.0000 HPO4-- + 1.0000 Er+++ = ErPO4 +1.0000 H+
-llnl_gamma 3.0
log_k +0.3782
-delta_H 0 # Not possible to calculate enthalpy of reaction ErPO4
# Enthalpy of formation: -0 kcal/mol
1.0000 SO4-- + 1.0000 Er+++ = ErSO4+
-llnl_gamma 4.0
log_k +3.5697
-delta_H 20.3008 kJ/mol # Calculated enthalpy of reaction ErSO4+
# Enthalpy of formation: -381.048 kcal/mol
-analytic 3.0363e+002 8.5667e-002 -8.9667e+003 -1.1942e+002 -1.4001e+002
# -Range: 0-300
2.0000 HAcetate + 1.0000 Eu+++ = Eu(Acetate)2+ +2.0000 H+
-llnl_gamma 4.0
log_k -4.6912
-delta_H -28.3257 kJ/mol # Calculated enthalpy of reaction Eu(Acetate)2+
# Enthalpy of formation: -383.67 kcal/mol
-analytic -2.7589e+001 1.5772e-003 -1.1008e+003 7.9899e+000 5.6652e+005
# -Range: 0-300
3.0000 HAcetate + 1.0000 Eu+++ = Eu(Acetate)3 +3.0000 H+
-llnl_gamma 3.0
log_k -7.9824
-delta_H -47.3629 kJ/mol # Calculated enthalpy of reaction Eu(Acetate)3
# Enthalpy of formation: -504.32 kcal/mol
-analytic -3.7470e+001 1.9276e-003 -1.0318e+003 9.7078e+000 7.4558e+005
# -Range: 0-300
2.0000 HCO3- + 1.0000 Eu+++ = Eu(CO3)2- +2.0000 H+
-llnl_gamma 4.0
log_k -8.3993
-delta_H 0 # Not possible to calculate enthalpy of reaction Eu(CO3)2-
# Enthalpy of formation: -0 kcal/mol
3.0000 HCO3- + 1.0000 Eu+++ = Eu(CO3)3--- +3.0000 H+
-llnl_gamma 4.0
log_k -16.8155
-delta_H 0 # Not possible to calculate enthalpy of reaction Eu(CO3)3-3
# Enthalpy of formation: -0 kcal/mol
2.0000 HPO4-- + 1.0000 Eu+++ = Eu(HPO4)2-
-llnl_gamma 4.0
log_k +9.6000
-delta_H 0 # Not possible to calculate enthalpy of reaction Eu(HPO4)2-
# Enthalpy of formation: -0 kcal/mol
# Redundant with EuO+
#2.0000 H2O + 1.0000 Eu+++ = Eu(OH)2+ +2.0000 H+
# -llnl_gamma 4.0
# log_k -14.8609
# -delta_H 0 # Not possible to calculate enthalpy of reaction Eu(OH)2+
## Enthalpy of formation: -0 kcal/mol
2.0000 H2O + 1.0000 HCO3- + 1.0000 Eu+++ = Eu(OH)2CO3- +3.0000 H+
-llnl_gamma 4.0
log_k -17.8462
-delta_H 0 # Not possible to calculate enthalpy of reaction Eu(OH)2CO3-
# Enthalpy of formation: -0 kcal/mol
# Redundant with EuO2H
#3.0000 H2O + 1.0000 Eu+++ = Eu(OH)3 +3.0000 H+
# -llnl_gamma 3.0
# log_k -24.1253
# -delta_H 0 # Not possible to calculate enthalpy of reaction Eu(OH)3
## Enthalpy of formation: -0 kcal/mol
# Redundant with EuO2-
#4.0000 H2O + 1.0000 Eu+++ = Eu(OH)4- +4.0000 H+
# -llnl_gamma 4.0
# log_k -36.5958
# -delta_H 0 # Not possible to calculate enthalpy of reaction Eu(OH)4-
## Enthalpy of formation: -0 kcal/mol
2.0000 HPO4-- + 1.0000 Eu+++ = Eu(PO4)2--- +2.0000 H+
-llnl_gamma 4.0
log_k -3.9837
-delta_H 0 # Not possible to calculate enthalpy of reaction Eu(PO4)2-3
# Enthalpy of formation: -0 kcal/mol
2.0000 SO4-- + 1.0000 Eu+++ = Eu(SO4)2-
-llnl_gamma 4.0
log_k +5.4693
-delta_H 25.627 kJ/mol # Calculated enthalpy of reaction Eu(SO4)2-
# Enthalpy of formation: -2399 kJ/mol
-analytic 4.5178e+002 1.2285e-001 -1.3400e+004 -1.7697e+002 -2.0922e+002
# -Range: 0-300
2.0000 H2O + 2.0000 Eu+++ = Eu2(OH)2++++ +2.0000 H+
-llnl_gamma 5.5
log_k -6.9182
-delta_H 0 # Not possible to calculate enthalpy of reaction Eu2(OH)2+4
# Enthalpy of formation: -0 kcal/mol
1.0000 Eu+++ + 1.0000 Br- = EuBr++
-llnl_gamma 4.5
log_k +0.5572
-delta_H 0 # Not possible to calculate enthalpy of reaction EuBr+2
# Enthalpy of formation: -0 kcal/mol
2.0000 Br- + 1.0000 Eu+++ = EuBr2+
-llnl_gamma 4.0
log_k +0.2145
-delta_H 0 # Not possible to calculate enthalpy of reaction EuBr2+
# Enthalpy of formation: -0 kcal/mol
1.0000 Eu+++ + 1.0000 BrO3- = EuBrO3++
-llnl_gamma 4.5
log_k +4.5823
-delta_H 0 # Not possible to calculate enthalpy of reaction EuBrO3+2
# Enthalpy of formation: -0 kcal/mol
1.0000 Eu+++ + 1.0000 HAcetate = EuAcetate++ +1.0000 H+
-llnl_gamma 4.5
log_k -1.9571
-delta_H -14.5603 kJ/mol # Calculated enthalpy of reaction EuAcetate+2
# Enthalpy of formation: -264.28 kcal/mol
-analytic -1.5090e+001 1.0352e-003 -6.4435e+002 4.6225e+000 3.1649e+005
# -Range: 0-300
1.0000 HCO3- + 1.0000 Eu+++ = EuCO3+ +1.0000 H+
-llnl_gamma 4.0
log_k -2.4057
-delta_H 90.7844 kJ/mol # Calculated enthalpy of reaction EuCO3+
# Enthalpy of formation: -287.9 kcal/mol
-analytic 2.3548e+002 5.3819e-002 -6.9908e+003 -9.3137e+001 -1.0915e+002
# -Range: 0-300
1.0000 Eu++ + 1.0000 Cl- = EuCl+
-llnl_gamma 4.0
log_k +0.3819
-delta_H 8.50607 kJ/mol # Calculated enthalpy of reaction EuCl+
# Enthalpy of formation: -164 kcal/mol
-analytic 6.8695e+001 3.7619e-002 -1.0809e+003 -3.0665e+001 -1.6887e+001
# -Range: 0-300
1.0000 Eu+++ + 1.0000 Cl- = EuCl++
-llnl_gamma 4.5
log_k +0.3086
-delta_H 13.9453 kJ/mol # Calculated enthalpy of reaction EuCl+2
# Enthalpy of formation: -181.3 kcal/mol
-analytic 7.9275e+001 3.7878e-002 -1.7895e+003 -3.4041e+001 -2.7947e+001
# -Range: 0-300
2.0000 Cl- + 1.0000 Eu++ = EuCl2
-llnl_gamma 3.0
log_k +1.2769
-delta_H 5.71534 kJ/mol # Calculated enthalpy of reaction EuCl2
# Enthalpy of formation: -204.6 kcal/mol
-analytic 1.0474e+002 6.7132e-002 -7.0448e+002 -4.8928e+001 -1.1024e+001
# -Range: 0-300
2.0000 Cl- + 1.0000 Eu+++ = EuCl2+
-llnl_gamma 4.0
log_k -0.0425
-delta_H 18.6857 kJ/mol # Calculated enthalpy of reaction EuCl2+
# Enthalpy of formation: -220.1 kcal/mol
-analytic 2.1758e+002 8.0336e-002 -5.5499e+003 -9.0087e+001 -8.6665e+001
# -Range: 0-300
3.0000 Cl- + 1.0000 Eu+++ = EuCl3
-llnl_gamma 3.0
log_k -0.4669
-delta_H 11.2926 kJ/mol # Calculated enthalpy of reaction EuCl3
# Enthalpy of formation: -261.8 kcal/mol
-analytic 4.2075e+002 1.2890e-001 -1.1288e+004 -1.7043e+002 -1.7627e+002
# -Range: 0-300
3.0000 Cl- + 1.0000 Eu++ = EuCl3-
-llnl_gamma 4.0
log_k +2.0253
-delta_H -3.76978 kJ/mol # Calculated enthalpy of reaction EuCl3-
# Enthalpy of formation: -246.8 kcal/mol
-analytic 1.1546e+001 6.4683e-002 3.7299e+003 -1.6672e+001 5.8196e+001
# -Range: 0-300
4.0000 Cl- + 1.0000 Eu+++ = EuCl4-
-llnl_gamma 4.0
log_k -0.8913
-delta_H -9.90771 kJ/mol # Calculated enthalpy of reaction EuCl4-
# Enthalpy of formation: -306.8 kcal/mol
-analytic 4.8122e+002 1.3081e-001 -1.2950e+004 -1.9302e+002 -2.0222e+002
# -Range: 0-300
4.0000 Cl- + 1.0000 Eu++ = EuCl4--
-llnl_gamma 4.0
log_k +2.8470
-delta_H -19.9493 kJ/mol # Calculated enthalpy of reaction EuCl4-2
# Enthalpy of formation: -290.6 kcal/mol
-analytic -1.2842e+002 5.0789e-002 9.8815e+003 3.3565e+001 1.5423e+002
# -Range: 0-300
1.0000 F- + 1.0000 Eu++ = EuF+
-llnl_gamma 4.0
log_k -1.3487
-delta_H 16.9452 kJ/mol # Calculated enthalpy of reaction EuF+
# Enthalpy of formation: -202.2 kcal/mol
-analytic 6.2412e+001 3.5839e-002 -1.3660e+003 -2.8223e+001 -2.1333e+001
# -Range: 0-300
1.0000 F- + 1.0000 Eu+++ = EuF++
-llnl_gamma 4.5
log_k +4.4420
-delta_H 23.6396 kJ/mol # Calculated enthalpy of reaction EuF+2
# Enthalpy of formation: -219.2 kcal/mol
-analytic 1.0063e+002 4.1834e-002 -2.7355e+003 -4.0195e+001 -4.2714e+001
# -Range: 0-300
2.0000 F- + 1.0000 Eu++ = EuF2
-llnl_gamma 3.0
log_k -2.0378
-delta_H 17.5728 kJ/mol # Calculated enthalpy of reaction EuF2
# Enthalpy of formation: -282.2 kcal/mol
-analytic 1.2065e+002 7.1705e-002 -1.7998e+003 -5.5760e+001 -2.8121e+001
# -Range: 0-300
2.0000 F- + 1.0000 Eu+++ = EuF2+
-llnl_gamma 4.0
log_k +7.7112
-delta_H 13.8072 kJ/mol # Calculated enthalpy of reaction EuF2+
# Enthalpy of formation: -301.7 kcal/mol
-analytic 2.4099e+002 8.4714e-002 -5.7702e+003 -9.6640e+001 -9.0109e+001
# -Range: 0-300
3.0000 F- + 1.0000 Eu+++ = EuF3
-llnl_gamma 3.0
log_k +10.1741
-delta_H -8.9956 kJ/mol # Calculated enthalpy of reaction EuF3
# Enthalpy of formation: -387.3 kcal/mol
-analytic 4.5022e+002 1.3560e-001 -1.0801e+004 -1.7951e+002 -1.6867e+002
# -Range: 0-300
3.0000 F- + 1.0000 Eu++ = EuF3-
-llnl_gamma 4.0
log_k -2.5069
-delta_H 3.5564 kJ/mol # Calculated enthalpy of reaction EuF3-
# Enthalpy of formation: -365.7 kcal/mol
-analytic -2.8441e+001 5.5972e-002 4.4573e+003 -2.2782e+000 6.9558e+001
# -Range: 0-300
4.0000 F- + 1.0000 Eu+++ = EuF4-
-llnl_gamma 4.0
log_k +12.1239
-delta_H -52.3 kJ/mol # Calculated enthalpy of reaction EuF4-
# Enthalpy of formation: -477.8 kcal/mol
-analytic 5.0246e+002 1.3629e-001 -1.1092e+004 -1.9952e+002 -1.7323e+002
# -Range: 0-300
4.0000 F- + 1.0000 Eu++ = EuF4--
-llnl_gamma 4.0
log_k -2.8294
-delta_H -37.656 kJ/mol # Calculated enthalpy of reaction EuF4-2
# Enthalpy of formation: -455.7 kcal/mol
-analytic -1.8730e+002 3.9237e-002 1.2303e+004 5.3179e+001 1.9204e+002
# -Range: 0-300
1.0000 HPO4-- + 1.0000 H+ + 1.0000 Eu+++ = EuH2PO4++
-llnl_gamma 4.5
log_k +9.4484
-delta_H -17.0916 kJ/mol # Calculated enthalpy of reaction EuH2PO4+2
# Enthalpy of formation: -457.6 kcal/mol
-analytic 1.0873e+002 6.3416e-002 2.7202e+002 -4.8113e+001 4.2122e+000
# -Range: 0-300
1.0000 HCO3- + 1.0000 Eu+++ = EuHCO3++
-llnl_gamma 4.5
log_k +1.6258
-delta_H 8.77803 kJ/mol # Calculated enthalpy of reaction EuHCO3+2
# Enthalpy of formation: -307.5 kcal/mol
-analytic 3.9266e+001 3.1608e-002 -9.8731e+001 -1.8875e+001 -1.5524e+000
# -Range: 0-300
1.0000 HPO4-- + 1.0000 Eu+++ = EuHPO4+
-llnl_gamma 4.0
log_k +5.7000
-delta_H 0 # Not possible to calculate enthalpy of reaction EuHPO4+
# Enthalpy of formation: -0 kcal/mol
1.0000 IO3- + 1.0000 Eu+++ = EuIO3++
-llnl_gamma 4.5
log_k +2.1560
-delta_H 11.8314 kJ/mol # Calculated enthalpy of reaction EuIO3+2
# Enthalpy of formation: -814.927 kJ/mol
-analytic 1.4970e+002 4.7369e-002 -4.1559e+003 -5.9687e+001 -6.4893e+001
# -Range: 0-300
1.0000 NO3- + 1.0000 Eu+++ = EuNO3++
-llnl_gamma 4.5
log_k +0.8745
-delta_H -32.0955 kJ/mol # Calculated enthalpy of reaction EuNO3+2
# Enthalpy of formation: -201.8 kcal/mol
-analytic 1.7398e+001 2.5467e-002 2.2683e+003 -1.2810e+001 3.5389e+001
# -Range: 0-300
1.0000 H2O + 1.0000 Eu+++ = EuO+ +2.0000 H+
-llnl_gamma 4.0
log_k -16.337
-delta_H 110.947 kJ/mol # Calculated enthalpy of reaction EuO+
# Enthalpy of formation: -186.5 kcal/mol
-analytic 1.8876e+002 3.0194e-002 -1.3836e+004 -6.7770e+001 -2.1595e+002
# -Range: 0-300
2.0000 H2O + 1.0000 Eu+++ = EuO2- +4.0000 H+
-llnl_gamma 4.0
log_k -34.5066
-delta_H 281.307 kJ/mol # Calculated enthalpy of reaction EuO2-
# Enthalpy of formation: -214.1 kcal/mol
-analytic 7.5244e+001 3.7089e-004 -1.3587e+004 -2.3859e+001 -4.6713e+005
# -Range: 0-300
2.0000 H2O + 1.0000 Eu+++ = EuO2H +3.0000 H+
-llnl_gamma 3.0
log_k -25.4173
-delta_H 222.313 kJ/mol # Calculated enthalpy of reaction EuO2H
# Enthalpy of formation: -228.2 kcal/mol
-analytic 3.6754e+002 5.3868e-002 -2.4034e+004 -1.3272e+002 -3.7514e+002
# -Range: 0-300
2.0000 HCO3- + 1.0000 H2O + 1.0000 Eu+++ = EuOH(CO3)2-- +3.0000 H+
-llnl_gamma 4.0
log_k -15.176
-delta_H 0 # Not possible to calculate enthalpy of reaction EuOH(CO3)2-2
# Enthalpy of formation: -0 kcal/mol
1.0000 H2O + 1.0000 Eu+++ = EuOH++ +1.0000 H+
-llnl_gamma 4.5
log_k -7.9075
-delta_H 78.0065 kJ/mol # Calculated enthalpy of reaction EuOH+2
# Enthalpy of formation: -194.373 kcal/mol
-analytic 6.7691e+001 1.2066e-002 -6.1871e+003 -2.3617e+001 -9.6563e+001
# -Range: 0-300
1.0000 HCO3- + 1.0000 H2O + 1.0000 Eu+++ = EuOHCO3 +2.0000 H+
-llnl_gamma 3.0
log_k -8.4941
-delta_H 0 # Not possible to calculate enthalpy of reaction EuOHCO3
# Enthalpy of formation: -0 kcal/mol
1.0000 HPO4-- + 1.0000 Eu+++ = EuPO4 +1.0000 H+
-llnl_gamma 3.0
log_k -0.1218
-delta_H 0 # Not possible to calculate enthalpy of reaction EuPO4
# Enthalpy of formation: -0 kcal/mol
1.0000 SO4-- + 1.0000 Eu+++ = EuSO4+
-llnl_gamma 4.0
log_k +3.6430
-delta_H 62.3416 kJ/mol # Calculated enthalpy of reaction EuSO4+
# Enthalpy of formation: -347.2 kcal/mol
-analytic 3.0587e+002 8.6208e-002 -9.0387e+003 -1.2026e+002 -1.4113e+002
# -Range: 0-300
2.0000 HAcetate + 1.0000 Fe++ = Fe(Acetate)2 +2.0000 H+
-llnl_gamma 3.0
log_k -7.0295
-delta_H -20.2924 kJ/mol # Calculated enthalpy of reaction Fe(Acetate)2
# Enthalpy of formation: -259.1 kcal/mol
-analytic -2.9862e+001 1.3901e-003 -1.6908e+003 8.6283e+000 6.0125e+005
# -Range: 0-300
2.0000 H2O + 1.0000 Fe++ = Fe(OH)2 +2.0000 H+
-llnl_gamma 3.0
log_k -20.6
-delta_H 0 # Not possible to calculate enthalpy of reaction Fe(OH)2
# Enthalpy of formation: -0 kcal/mol
2.0000 H2O + 1.0000 Fe+++ = Fe(OH)2+ +2.0000 H+
-llnl_gamma 4.0
log_k -5.67
-delta_H 0 # Not possible to calculate enthalpy of reaction Fe(OH)2+
# Enthalpy of formation: -0 kcal/mol
3.0000 H2O + 1.0000 Fe+++ = Fe(OH)3 +3.0000 H+
-llnl_gamma 3.0
log_k -12
-delta_H 0 # Not possible to calculate enthalpy of reaction Fe(OH)3
# Enthalpy of formation: -0 kcal/mol
3.0000 H2O + 1.0000 Fe++ = Fe(OH)3- +3.0000 H+
-llnl_gamma 4.0
log_k -31
-delta_H 0 # Not possible to calculate enthalpy of reaction Fe(OH)3-
# Enthalpy of formation: -0 kcal/mol
4.0000 H2O + 1.0000 Fe+++ = Fe(OH)4- +4.0000 H+
-llnl_gamma 4.0
log_k -21.6
-delta_H 0 # Not possible to calculate enthalpy of reaction Fe(OH)4-
# Enthalpy of formation: -0 kcal/mol
4.0000 H2O + 1.0000 Fe++ = Fe(OH)4-- +4.0000 H+
-llnl_gamma 4.0
log_k -46
-delta_H 0 # Not possible to calculate enthalpy of reaction Fe(OH)4-2
# Enthalpy of formation: -0 kcal/mol
2.0000 SO4-- + 1.0000 Fe+++ = Fe(SO4)2-
-llnl_gamma 4.0
log_k +3.2137
-delta_H 0 # Not possible to calculate enthalpy of reaction Fe(SO4)2-
# Enthalpy of formation: -0 kcal/mol
2.0000 H2O + 2.0000 Fe+++ = Fe2(OH)2++++ +2.0000 H+
-llnl_gamma 5.5
log_k -2.95
-delta_H 0 # Not possible to calculate enthalpy of reaction Fe2(OH)2+4
# Enthalpy of formation: -0 kcal/mol
4.0000 H2O + 3.0000 Fe+++ = Fe3(OH)4+5 +4.0000 H+
-llnl_gamma 6.0
log_k -6.3
-delta_H 0 # Not possible to calculate enthalpy of reaction Fe3(OH)4+5
# Enthalpy of formation: -0 kcal/mol
1.0000 Fe++ + 1.0000 HAcetate = FeAcetate+ +1.0000 H+
-llnl_gamma 4.0
log_k -3.4671
-delta_H -3.80744 kJ/mol # Calculated enthalpy of reaction FeAcetate+
# Enthalpy of formation: -139.06 kcal/mol
-analytic -1.3781e+001 9.6253e-004 -7.5310e+002 4.0135e+000 2.3416e+005
# -Range: 0-300
1.0000 HCO3- + 1.0000 Fe++ = FeCO3 +1.0000 H+
-llnl_gamma 3.0
log_k -5.5988
-delta_H 0 # Not possible to calculate enthalpy of reaction FeCO3
# Enthalpy of formation: -0 kcal/mol
1.0000 HCO3- + 1.0000 Fe+++ = FeCO3+ +1.0000 H+
-llnl_gamma 4.0
log_k -0.6088
-delta_H -50.208 kJ/mol # Calculated enthalpy of reaction FeCO3+
# Enthalpy of formation: -188.748 kcal/mol
-analytic 1.7100e+002 8.0413e-002 -4.3217e+002 -7.8449e+001 -6.7948e+000
# -Range: 0-300
1.0000 Fe++ + 1.0000 Cl- = FeCl+
-llnl_gamma 4.0
log_k -0.1605
-delta_H 3.02503 kJ/mol # Calculated enthalpy of reaction FeCl+
# Enthalpy of formation: -61.26 kcal/mol
-analytic 8.2435e+001 3.7755e-002 -1.4765e+003 -3.5918e+001 -2.3064e+001
# -Range: 0-300
1.0000 Fe+++ + 1.0000 Cl- = FeCl++
-llnl_gamma 4.5
log_k -0.8108
-delta_H 36.6421 kJ/mol # Calculated enthalpy of reaction FeCl+2
# Enthalpy of formation: -180.018 kJ/mol
-analytic 1.6186e+002 5.9436e-002 -5.1913e+003 -6.5852e+001 -8.1053e+001
# -Range: 0-300
2.0000 Cl- + 1.0000 Fe++ = FeCl2
-llnl_gamma 3.0
log_k -2.4541
-delta_H 6.46846 kJ/mol # Calculated enthalpy of reaction FeCl2
# Enthalpy of formation: -100.37 kcal/mol
-analytic 1.9171e+002 7.8070e-002 -4.1048e+003 -8.2292e+001 -6.4108e+001
# -Range: 0-300
2.0000 Cl- + 1.0000 Fe+++ = FeCl2+
-llnl_gamma 4.0
log_k +2.1300
-delta_H 0 # Not possible to calculate enthalpy of reaction FeCl2+
# Enthalpy of formation: -0 kcal/mol
4.0000 Cl- + 1.0000 Fe+++ = FeCl4-
-llnl_gamma 4.0
log_k -0.79
-delta_H 0 # Not possible to calculate enthalpy of reaction FeCl4-
# Enthalpy of formation: -0 kcal/mol
4.0000 Cl- + 1.0000 Fe++ = FeCl4--
-llnl_gamma 4.0
log_k -1.9
-delta_H 0 # Not possible to calculate enthalpy of reaction FeCl4-2
# Enthalpy of formation: -0 kcal/mol
-analytic -2.4108e+002 -6.0086e-003 9.7979e+003 8.4084e+001 1.5296e+002
# -Range: 0-300
1.0000 Fe++ + 1.0000 F- = FeF+
-llnl_gamma 4.0
log_k +1.3600
-delta_H 0 # Not possible to calculate enthalpy of reaction FeF+
# Enthalpy of formation: -0 kcal/mol
1.0000 Fe+++ + 1.0000 F- = FeF++
-llnl_gamma 4.5
log_k +4.1365
-delta_H 14.327 kJ/mol # Calculated enthalpy of reaction FeF+2
# Enthalpy of formation: -370.601 kJ/mol
-analytic 1.7546e+002 6.3754e-002 -4.3166e+003 -7.1052e+001 -6.7408e+001
# -Range: 0-300
2.0000 F- + 1.0000 Fe+++ = FeF2+
-llnl_gamma 4.0
log_k +8.3498
-delta_H 23.9776 kJ/mol # Calculated enthalpy of reaction FeF2+
# Enthalpy of formation: -696.298 kJ/mol
-analytic 2.9080e+002 1.0393e-001 -7.2118e+003 -1.1688e+002 -1.1262e+002
# -Range: 0-300
1.0000 HPO4-- + 1.0000 H+ + 1.0000 Fe++ = FeH2PO4+
-llnl_gamma 4.0
log_k +2.7000
-delta_H 0 # Not possible to calculate enthalpy of reaction FeH2PO4+
# Enthalpy of formation: -0 kcal/mol
1.0000 HPO4-- + 1.0000 H+ + 1.0000 Fe+++ = FeH2PO4++
-llnl_gamma 4.5
log_k +4.1700
-delta_H 0 # Not possible to calculate enthalpy of reaction FeH2PO4+2
# Enthalpy of formation: -0 kcal/mol
1.0000 HCO3- + 1.0000 Fe++ = FeHCO3+
-llnl_gamma 4.0
log_k +2.7200
-delta_H 0 # Not possible to calculate enthalpy of reaction FeHCO3+
# Enthalpy of formation: -0 kcal/mol
1.0000 HPO4-- + 1.0000 Fe++ = FeHPO4
-llnl_gamma 3.0
log_k +3.6000
-delta_H 0 # Not possible to calculate enthalpy of reaction FeHPO4
# Enthalpy of formation: -0 kcal/mol
1.0000 HPO4-- + 1.0000 Fe+++ = FeHPO4+
-llnl_gamma 4.0
log_k +10.1800
-delta_H 0 # Not possible to calculate enthalpy of reaction FeHPO4+
# Enthalpy of formation: -0 kcal/mol
1.0000 NO2- + 1.0000 Fe+++ = FeNO2++
-llnl_gamma 4.5
log_k +3.1500
-delta_H 0 # Not possible to calculate enthalpy of reaction FeNO2+2
# Enthalpy of formation: -0 kcal/mol
1.0000 NO3- + 1.0000 Fe+++ = FeNO3++
-llnl_gamma 4.5
log_k +1.0000
-delta_H 0 # Not possible to calculate enthalpy of reaction FeNO3+2
# Enthalpy of formation: -0 kcal/mol
1.0000 H2O + 1.0000 Fe++ = FeOH+ +1.0000 H+
-llnl_gamma 4.0
log_k -9.5
-delta_H 0 # Not possible to calculate enthalpy of reaction FeOH+
# Enthalpy of formation: -0 kcal/mol
1.0000 H2O + 1.0000 Fe+++ = FeOH++ +1.0000 H+
-llnl_gamma 4.5
log_k -2.19
-delta_H 0 # Not possible to calculate enthalpy of reaction FeOH+2
# Enthalpy of formation: -0 kcal/mol
1.0000 HPO4-- + 1.0000 Fe++ = FePO4- +1.0000 H+
-llnl_gamma 4.0
log_k -4.3918
-delta_H 0 # Not possible to calculate enthalpy of reaction FePO4-
# Enthalpy of formation: -0 kcal/mol
1.0000 SO4-- + 1.0000 Fe++ = FeSO4
-llnl_gamma 3.0
log_k +2.2000
-delta_H 0 # Not possible to calculate enthalpy of reaction FeSO4
# Enthalpy of formation: -0 kcal/mol
1.0000 SO4-- + 1.0000 Fe+++ = FeSO4+
-llnl_gamma 4.0
log_k +1.9276
-delta_H 27.181 kJ/mol # Calculated enthalpy of reaction FeSO4+
# Enthalpy of formation: -932.001 kJ/mol
-analytic 2.5178e+002 1.0080e-001 -6.0977e+003 -1.0483e+002 -9.5223e+001
# -Range: 0-300
2.0000 HAcetate + 1.0000 Gd+++ = Gd(Acetate)2+ +2.0000 H+
-llnl_gamma 4.0
log_k -4.9625
-delta_H -22.3426 kJ/mol # Calculated enthalpy of reaction Gd(Acetate)2+
# Enthalpy of formation: -401.74 kcal/mol
-analytic -4.3124e+001 1.2995e-004 -4.3494e+002 1.3677e+001 5.1224e+005
# -Range: 0-300
3.0000 HAcetate + 1.0000 Gd+++ = Gd(Acetate)3 +3.0000 H+
-llnl_gamma 3.0
log_k -8.3489
-delta_H -37.9907 kJ/mol # Calculated enthalpy of reaction Gd(Acetate)3
# Enthalpy of formation: -521.58 kcal/mol
-analytic -8.8296e+001 -5.0939e-003 1.2268e+003 2.8513e+001 6.0745e+005
# -Range: 0-300
2.0000 HCO3- + 1.0000 Gd+++ = Gd(CO3)2- +2.0000 H+
-llnl_gamma 4.0
log_k -7.5576
-delta_H 0 # Not possible to calculate enthalpy of reaction Gd(CO3)2-
# Enthalpy of formation: -0 kcal/mol
2.0000 HPO4-- + 1.0000 Gd+++ = Gd(HPO4)2-
-llnl_gamma 4.0
log_k +9.6000
-delta_H 0 # Not possible to calculate enthalpy of reaction Gd(HPO4)2-
# Enthalpy of formation: -0 kcal/mol
# Redundant with GdO2-
#4.0000 H2O + 1.0000 Gd+++ = Gd(OH)4- +4.0000 H+
# -llnl_gamma 4.0
# log_k -33.8803
# -delta_H 0 # Not possible to calculate enthalpy of reaction Gd(OH)4-
## Enthalpy of formation: -0 kcal/mol
2.0000 HPO4-- + 1.0000 Gd+++ = Gd(PO4)2--- +2.0000 H+
-llnl_gamma 4.0
log_k -3.9437
-delta_H 0 # Not possible to calculate enthalpy of reaction Gd(PO4)2-3
# Enthalpy of formation: -0 kcal/mol
2.0000 SO4-- + 1.0000 Gd+++ = Gd(SO4)2-
-llnl_gamma 4.0
log_k +5.1000
-delta_H 0 # Not possible to calculate enthalpy of reaction Gd(SO4)2-
# Enthalpy of formation: -0 kcal/mol
1.0000 Gd+++ + 1.0000 HAcetate = GdAcetate++ +1.0000 H+
-llnl_gamma 4.5
log_k -2.1037
-delta_H -11.7152 kJ/mol # Calculated enthalpy of reaction GdAcetate+2
# Enthalpy of formation: -283.1 kcal/mol
-analytic -1.4118e+001 1.6660e-003 -7.5206e+002 4.2614e+000 3.1187e+005
# -Range: 0-300
1.0000 HCO3- + 1.0000 Gd+++ = GdCO3+ +1.0000 H+
-llnl_gamma 4.0
log_k -2.479
-delta_H 89.9476 kJ/mol # Calculated enthalpy of reaction GdCO3+
# Enthalpy of formation: -307.6 kcal/mol
-analytic 2.3628e+002 5.4100e-002 -7.0746e+003 -9.3413e+001 -1.1046e+002
# -Range: 0-300
1.0000 Gd+++ + 1.0000 Cl- = GdCl++
-llnl_gamma 4.5
log_k +0.3086
-delta_H 14.7821 kJ/mol # Calculated enthalpy of reaction GdCl+2
# Enthalpy of formation: -200.6 kcal/mol
-analytic 8.0750e+001 3.8524e-002 -1.8591e+003 -3.4621e+001 -2.9034e+001
# -Range: 0-300
2.0000 Cl- + 1.0000 Gd+++ = GdCl2+
-llnl_gamma 4.0
log_k -0.0425
-delta_H 21.1961 kJ/mol # Calculated enthalpy of reaction GdCl2+
# Enthalpy of formation: -239 kcal/mol
-analytic 2.1754e+002 8.0996e-002 -5.6121e+003 -9.0067e+001 -8.7635e+001
# -Range: 0-300
3.0000 Cl- + 1.0000 Gd+++ = GdCl3
-llnl_gamma 3.0
log_k -0.4669
-delta_H 15.895 kJ/mol # Calculated enthalpy of reaction GdCl3
# Enthalpy of formation: -280.2 kcal/mol
-analytic 4.1398e+002 1.2829e-001 -1.1230e+004 -1.6770e+002 -1.7535e+002
# -Range: 0-300
4.0000 Cl- + 1.0000 Gd+++ = GdCl4-
-llnl_gamma 4.0
log_k -0.8913
-delta_H -1.53971 kJ/mol # Calculated enthalpy of reaction GdCl4-
# Enthalpy of formation: -324.3 kcal/mol
-analytic 4.7684e+002 1.3157e-001 -1.3068e+004 -1.9118e+002 -2.0405e+002
# -Range: 0-300
1.0000 Gd+++ + 1.0000 F- = GdF++
-llnl_gamma 4.5
log_k +4.5886
-delta_H 21.1292 kJ/mol # Calculated enthalpy of reaction GdF+2
# Enthalpy of formation: -239.3 kcal/mol
-analytic 1.0060e+002 4.2181e-002 -2.6024e+003 -4.0347e+001 -4.0637e+001
# -Range: 0-300
2.0000 F- + 1.0000 Gd+++ = GdF2+
-llnl_gamma 4.0
log_k +7.9311
-delta_H 11.2968 kJ/mol # Calculated enthalpy of reaction GdF2+
# Enthalpy of formation: -321.8 kcal/mol
-analytic 2.3793e+002 8.4732e-002 -5.4950e+003 -9.5689e+001 -8.5815e+001
# -Range: 0-300
3.0000 F- + 1.0000 Gd+++ = GdF3
-llnl_gamma 3.0
log_k +10.4673
-delta_H -11.506 kJ/mol # Calculated enthalpy of reaction GdF3
# Enthalpy of formation: -407.4 kcal/mol
-analytic 4.4257e+002 1.3500e-001 -1.0377e+004 -1.7680e+002 -1.6205e+002
# -Range: 0-300
4.0000 F- + 1.0000 Gd+++ = GdF4-
-llnl_gamma 4.0
log_k +12.4904
-delta_H -52.3 kJ/mol # Calculated enthalpy of reaction GdF4-
# Enthalpy of formation: -497.3 kcal/mol
-analytic 4.9026e+002 1.3534e-001 -1.0586e+004 -1.9501e+002 -1.6533e+002
# -Range: 0-300
1.0000 HPO4-- + 1.0000 H+ + 1.0000 Gd+++ = GdH2PO4++
-llnl_gamma 4.5
log_k +9.4484
-delta_H -14.9996 kJ/mol # Calculated enthalpy of reaction GdH2PO4+2
# Enthalpy of formation: -476.6 kcal/mol
-analytic 1.1058e+002 6.4124e-002 1.3451e+002 -4.8758e+001 2.0660e+000
# -Range: 0-300
1.0000 HCO3- + 1.0000 Gd+++ = GdHCO3++
-llnl_gamma 4.5
log_k +1.6991
-delta_H 10.0332 kJ/mol # Calculated enthalpy of reaction GdHCO3+2
# Enthalpy of formation: -326.7 kcal/mol
-analytic 4.1973e+001 3.2521e-002 -2.3475e+002 -1.9864e+001 -3.6757e+000
# -Range: 0-300
1.0000 HPO4-- + 1.0000 Gd+++ = GdHPO4+
-llnl_gamma 4.0
log_k -185.109
-delta_H 0 # Not possible to calculate enthalpy of reaction GdHPO4+
# Enthalpy of formation: -0 kcal/mol
1.0000 NO3- + 1.0000 Gd+++ = GdNO3++
-llnl_gamma 4.5
log_k +0.4347
-delta_H -25.8195 kJ/mol # Calculated enthalpy of reaction GdNO3+2
# Enthalpy of formation: -219.8 kcal/mol
-analytic 2.0253e+001 2.6372e-002 1.8785e+003 -1.3723e+001 2.9306e+001
# -Range: 0-300
1.0000 H2O + 1.0000 Gd+++ = GdO+ +2.0000 H+
-llnl_gamma 4.0
log_k -16.337
-delta_H 113.039 kJ/mol # Calculated enthalpy of reaction GdO+
# Enthalpy of formation: -205.5 kcal/mol
-analytic 2.0599e+002 3.2521e-002 -1.4547e+004 -7.4048e+001 -2.2705e+002
# -Range: 0-300
2.0000 H2O + 1.0000 Gd+++ = GdO2- +4.0000 H+
-llnl_gamma 4.0
log_k -34.4333
-delta_H 283.817 kJ/mol # Calculated enthalpy of reaction GdO2-
# Enthalpy of formation: -233 kcal/mol
-analytic 1.2067e+002 6.6276e-003 -1.5531e+004 -4.0448e+001 -4.3587e+005
# -Range: 0-300
2.0000 H2O + 1.0000 Gd+++ = GdO2H +3.0000 H+
-llnl_gamma 3.0
log_k -25.2707
-delta_H 224.405 kJ/mol # Calculated enthalpy of reaction GdO2H
# Enthalpy of formation: -247.2 kcal/mol
-analytic 3.6324e+002 4.7938e-002 -2.4275e+004 -1.2988e+002 -3.7889e+002
# -Range: 0-300
1.0000 H2O + 1.0000 Gd+++ = GdOH++ +1.0000 H+
-llnl_gamma 4.5
log_k -7.9075
-delta_H 79.9855 kJ/mol # Calculated enthalpy of reaction GdOH+2
# Enthalpy of formation: -213.4 kcal/mol
-analytic 8.3265e+001 1.4153e-002 -6.8229e+003 -2.9301e+001 -1.0649e+002
# -Range: 0-300
1.0000 HPO4-- + 1.0000 Gd+++ = GdPO4 +1.0000 H+
-llnl_gamma 3.0
log_k -0.1218
-delta_H 0 # Not possible to calculate enthalpy of reaction GdPO4
# Enthalpy of formation: -0 kcal/mol
1.0000 SO4-- + 1.0000 Gd+++ = GdSO4+
-llnl_gamma 4.0
log_k -3.687
-delta_H 20.0832 kJ/mol # Calculated enthalpy of reaction GdSO4+
# Enthalpy of formation: -376.8 kcal/mol
-analytic 3.0783e+002 8.6798e-002 -1.1246e+004 -1.2109e+002 -1.7557e+002
# -Range: 0-300
1.0000 O_phthalate-2 + 1.0000 H+ = H(O_phthalate)-
-llnl_gamma 4.0
log_k +5.4080
-delta_H 0 # Not possible to calculate enthalpy of reaction H(O_phthalate)-
# Enthalpy of formation: -0 kcal/mol
2.0000 H+ + 1.0000 CrO4-- = H2CrO4
-llnl_gamma 3.0
log_k +5.1750
-delta_H 42.8274 kJ/mol # Calculated enthalpy of reaction H2CrO4
# Enthalpy of formation: -200.364 kcal/mol
-analytic 4.2958e+002 1.4939e-001 -1.1474e+004 -1.7396e+002 -1.9499e+002
# -Range: 0-200
2.0000 H+ + 2.0000 F- = H2F2
-llnl_gamma 3.0
log_k +6.7680
-delta_H 0 # Not possible to calculate enthalpy of reaction H2F2
# Enthalpy of formation: -0 kcal/mol
2.0000 HPO4-- + 2.0000 H+ = H2P2O7-- +1.0000 H2O
-llnl_gamma 4.0
log_k +12.0709
-delta_H 19.7192 kJ/mol # Calculated enthalpy of reaction H2P2O7-2
# Enthalpy of formation: -544.6 kcal/mol
-analytic 1.4825e+002 6.7021e-002 -2.8329e+003 -5.9251e+001 -4.4248e+001
# -Range: 0-300
3.0000 H+ + 1.0000 HPO4-- + 1.0000 F- = H2PO3F +1.0000 H2O
-llnl_gamma 3.0
log_k +12.1047
-delta_H 0 # Not possible to calculate enthalpy of reaction H2PO3F
# Enthalpy of formation: -0 kcal/mol
1.0000 HPO4-- + 1.0000 H+ = H2PO4-
-llnl_gamma 4.0
log_k +7.2054
-delta_H -4.20492 kJ/mol # Calculated enthalpy of reaction H2PO4-
# Enthalpy of formation: -309.82 kcal/mol
-analytic 8.2149e+001 3.4077e-002 -1.0431e+003 -3.2970e+001 -1.6301e+001
# -Range: 0-300
#1.0000 HS- + 1.0000 H+ = H2S
# -llnl_gamma 3.0
# log_k +6.99
# -analytic 1.2833e+002 5.1641e-002 -1.1681e+003 -5.3665e+001 -1.8266e+001
# -Range: 0-300
# these (above) H2S values are from
# Suleimenov & Seward, Geochim. Cosmochim. Acta, v. 61, p. 5187-5198.
# values below are the original Thermo.com.v8.r6.230 data from somewhere
1.0000 HS- + 1.0000 H+ = H2S
-llnl_gamma 3.0
log_k +6.9877
-delta_H -21.5518 kJ/mol # Calculated enthalpy of reaction H2S
# Enthalpy of formation: -9.001 kcal/mol
-analytic 3.9283e+001 2.8727e-002 1.3477e+003 -1.8331e+001 2.1018e+001
# -Range: 0-300
2.0000 H+ + 1.0000 SO3-- = H2SO3
-llnl_gamma 3.0
log_k +9.2132
-delta_H 0 # Not possible to calculate enthalpy of reaction H2SO3
# Enthalpy of formation: -0 kcal/mol
2.0000 H+ + 1.0000 SO4-- = H2SO4
-llnl_gamma 3.0
log_k -1.0209
-delta_H 0 # Not possible to calculate enthalpy of reaction H2SO4
# Enthalpy of formation: -0 kcal/mol
2.0000 H+ + 1.0000 Se-- = H2Se
-llnl_gamma 3.0
log_k +18.7606
-delta_H 0 # Not possible to calculate enthalpy of reaction H2Se
# Enthalpy of formation: 19.412 kJ/mol
-analytic 3.6902e+002 1.2855e-001 -5.5900e+003 -1.4946e+002 -9.5054e+001
# -Range: 0-200
2.0000 H+ + 1.0000 SeO3-- = H2SeO3
-llnl_gamma 3.0
log_k +9.8589
-delta_H 1.7238 kJ/mol # Calculated enthalpy of reaction H2SeO3
# Enthalpy of formation: -507.469 kJ/mol
-analytic 2.7850e+002 1.0460e-001 -5.4934e+003 -1.1371e+002 -9.3383e+001
# -Range: 0-200
2.0000 H2O + 1.0000 SiO2 = H2SiO4-- +2.0000 H+
-llnl_gamma 4.0
log_k -22.96
-delta_H 0 # Not possible to calculate enthalpy of reaction H2SiO4-2
# Enthalpy of formation: -0 kcal/mol
2.0000 H+ + 1.0000 TcO4-- = H2TcO4
-llnl_gamma 3.0
log_k +9.0049
-delta_H 0 # Not possible to calculate enthalpy of reaction H2TcO4
# Enthalpy of formation: -0 kcal/mol
2.0000 H2O + 1.0000 VO2+ = H2VO4- + 2.0000 H+
-llnl_gamma 4.0
log_k -7.0922
-delta_H 0 # Not possible to calculate enthalpy of reaction H2VO4-
# Enthalpy of formation: -0 kcal/mol
-analytic 1.7105e+001 -1.7503e-002 -4.2671e+003 -1.8910e+000 -6.6589e+001
# -Range: 0-300
1.0000 H2AsO4- + 1.0000 H+ = H3AsO4
-llnl_gamma 3.0
log_k +2.2492
-delta_H 7.17876 kJ/mol # Calculated enthalpy of reaction H3AsO4
# Enthalpy of formation: -902.381 kJ/mol
-analytic 1.4043e+002 4.6288e-002 -3.5868e+003 -5.6560e+001 -6.0957e+001
# -Range: 0-200
3.0000 H+ + 2.0000 HPO4-- = H3P2O7- +1.0000 H2O
-llnl_gamma 4.0
log_k +14.4165
-delta_H 21.8112 kJ/mol # Calculated enthalpy of reaction H3P2O7-
# Enthalpy of formation: -544.1 kcal/mol
-analytic 2.3157e+002 1.0161e-001 -4.3723e+003 -9.4050e+001 -6.8295e+001
# -Range: 0-300
2.0000 H+ + 1.0000 HPO4-- = H3PO4
-llnl_gamma 3.0
log_k +9.3751
-delta_H 3.74468 kJ/mol # Calculated enthalpy of reaction H3PO4
# Enthalpy of formation: -307.92 kcal/mol
-analytic 1.8380e+002 6.7320e-002 -3.7792e+003 -7.3463e+001 -5.9025e+001
# -Range: 0-300
8.0000 H2O + 4.0000 SiO2 = H4(H2SiO4)4---- +4.0000 H+
-llnl_gamma 4.0
log_k -35.94
-delta_H 0 # Not possible to calculate enthalpy of reaction H4(H2SiO4)4-4
# Enthalpy of formation: -0 kcal/mol
4.0000 H+ + 2.0000 HPO4-- = H4P2O7 +1.0000 H2O
-llnl_gamma 3.0
log_k +15.9263
-delta_H 29.7226 kJ/mol # Calculated enthalpy of reaction H4P2O7
# Enthalpy of formation: -2268.6 kJ/mol
-analytic 6.9026e+002 2.4309e-001 -1.6165e+004 -2.7989e+002 -2.7475e+002
# -Range: 0-200
8.0000 H2O + 4.0000 SiO2 = H6(H2SiO4)4-- +2.0000 H+
-llnl_gamma 4.0
log_k -13.64
-delta_H 0 # Not possible to calculate enthalpy of reaction H6(H2SiO4)4-2
# Enthalpy of formation: -0 kcal/mol
2.0000 H2O + 1.0000 Al+++ = HAlO2 +3.0000 H+
-llnl_gamma 3.0
log_k -16.4329
-delta_H 144.704 kJ/mol # Calculated enthalpy of reaction HAlO2
# Enthalpy of formation: -230.73 kcal/mol
-analytic 4.2012e+001 1.9980e-002 -7.7847e+003 -1.5470e+001 -1.2149e+002
# -Range: 0-300
1.0000 H2AsO3- + 1.0000 H+ = HAsO2 +1.0000 H2O
-llnl_gamma 3.0
log_k 9.2792
-delta_H 0 # Not possible to calculate enthalpy of reaction HAsO2
# Enthalpy of formation: -0 kcal/mol
-analytic 3.1290e+002 9.3052e-002 -6.5052e+003 -1.2510e+002 -1.1058e+002
# -Range: 0-200
1.0000 H2AsO4- + 1.0000 H+ + 1.0000 F- = HAsO3F- +1.0000 H2O
-llnl_gamma 4.0
log_k +46.1158
-delta_H 0 # Not possible to calculate enthalpy of reaction HAsO3F-
# Enthalpy of formation: -0 kcal/mol
1.0000 H2AsO4- = HAsO4-- +1.0000 H+
-llnl_gamma 4.0
log_k -6.7583
-delta_H 3.22168 kJ/mol # Calculated enthalpy of reaction HAsO4-2
# Enthalpy of formation: -216.62 kcal/mol
-analytic -8.4546e+001 -3.4630e-002 1.1829e+003 3.3997e+001 1.8483e+001
# -Range: 0-300
3.0000 H+ + 2.0000 HS- + 1.0000 H2AsO3- = HAsS2 +3.0000 H2O
-llnl_gamma 3.0
log_k +30.4803
-delta_H 0 # Not possible to calculate enthalpy of reaction HAsS2
# Enthalpy of formation: -0 kcal/mol
1.0000 H+ + 1.0000 BrO- = HBrO
-llnl_gamma 3.0
log_k +8.3889
-delta_H 0 # Not possible to calculate enthalpy of reaction HBrO
# Enthalpy of formation: -0 kcal/mol
1.0000 H+ + 1.0000 Cyanide- = HCyanide
-llnl_gamma 3.0
log_k +9.2359
-delta_H -43.5136 kJ/mol # Calculated enthalpy of reaction HCyanide
# Enthalpy of formation: 25.6 kcal/mol
-analytic 1.0536e+001 2.3105e-002 3.3038e+003 -7.7786e+000 5.1550e+001
# -Range: 0-300
1.0000 H+ + 1.0000 Cl- = HCl
-llnl_gamma 3.0
log_k -0.67
-delta_H 0 # Not possible to calculate enthalpy of reaction HCl
# Enthalpy of formation: -0 kcal/mol
-analytic 4.1893e+002 1.1103e-001 -1.1784e+004 -1.6697e+002 -1.8400e+002
# -Range: 0-300
1.0000 H+ + 1.0000 ClO- = HClO
-llnl_gamma 3.0
log_k +7.5692
-delta_H 0 # Not possible to calculate enthalpy of reaction HClO
# Enthalpy of formation: -0 kcal/mol
1.0000 H+ + 1.0000 ClO2- = HClO2
-llnl_gamma 3.0
log_k +3.1698
-delta_H 0 # Not possible to calculate enthalpy of reaction HClO2
# Enthalpy of formation: -0 kcal/mol
2.0000 H2O + 1.0000 Co++ = HCoO2- +3.0000 H+
-llnl_gamma 4.0
log_k -21.243
-delta_H 0 # Not possible to calculate enthalpy of reaction HCoO2-
# Enthalpy of formation: -0 kcal/mol
1.0000 H+ + 1.0000 CrO4-- = HCrO4-
-llnl_gamma 4.0
log_k +6.4944
-delta_H 2.9288 kJ/mol # Calculated enthalpy of reaction HCrO4-
# Enthalpy of formation: -209.9 kcal/mol
-analytic 4.4944e+001 3.2740e-002 1.8400e+002 -1.9722e+001 2.8578e+000
# -Range: 0-300
1.0000 H+ + 1.0000 F- = HF
-llnl_gamma 3.0
log_k +3.1681
-delta_H 13.87 kJ/mol # Calculated enthalpy of reaction HF
# Enthalpy of formation: -76.835 kcal/mol
-analytic 8.6626e+001 3.2861e-002 -2.3026e+003 -3.4559e+001 -3.5956e+001
# -Range: 0-300
2.0000 F- + 1.0000 H+ = HF2-
-llnl_gamma 4.0
log_k +2.5509
-delta_H 20.7526 kJ/mol # Calculated enthalpy of reaction HF2-
# Enthalpy of formation: -155.34 kcal/mol
-analytic 1.4359e+002 4.0866e-002 -4.6776e+003 -5.5574e+001 -7.3032e+001
# -Range: 0-300
1.0000 IO3- + 1.0000 H+ = HIO3
-llnl_gamma 3.0
log_k +0.4915
-delta_H 0 # Not possible to calculate enthalpy of reaction HIO3
# Enthalpy of formation: -0 kcal/mol
1.0000 N3- + 1.0000 H+ = HN3
-llnl_gamma 3.0
log_k +4.7001
-delta_H -15 kJ/mol # Calculated enthalpy of reaction HN3
# Enthalpy of formation: 260.14 kJ/mol
-analytic 6.9976e+001 2.4359e-002 -7.1947e+002 -2.8339e+001 -1.2242e+001
# -Range: 0-200
1.0000 NO2- + 1.0000 H+ = HNO2
-llnl_gamma 3.0
log_k +3.2206
-delta_H -14.782 kJ/mol # Calculated enthalpy of reaction HNO2
# Enthalpy of formation: -119.382 kJ/mol
-analytic 1.9653e+000 -1.1603e-004 0.0000e+000 0.0000e+000 1.1569e+005
# -Range: 0-200
1.0000 NO3- + 1.0000 H+ = HNO3
-llnl_gamma 3.0
log_k -1.3025
-delta_H 16.8155 kJ/mol # Calculated enthalpy of reaction HNO3
# Enthalpy of formation: -45.41 kcal/mol
-analytic 9.9744e+001 3.4866e-002 -3.0975e+003 -4.0830e+001 -4.8363e+001
# -Range: 0-300
2.0000 HPO4-- + 1.0000 H+ = HP2O7--- +1.0000 H2O
-llnl_gamma 4.0
log_k +5.4498
-delta_H 23.3326 kJ/mol # Calculated enthalpy of reaction HP2O7-3
# Enthalpy of formation: -2274.99 kJ/mol
-analytic 3.9159e+002 1.5438e-001 -8.7071e+003 -1.6283e+002 -1.3598e+002
# -Range: 0-300
2.0000 H+ + 1.0000 HPO4-- + 1.0000 F- = HPO3F- +1.0000 H2O
-llnl_gamma 4.0
log_k +11.2988
-delta_H 0 # Not possible to calculate enthalpy of reaction HPO3F-
# Enthalpy of formation: -0 kcal/mol
1.0000 RuO4 + 1.0000 H2O = HRuO5- +1.0000 H+
-llnl_gamma 4.0
log_k -11.5244
-delta_H 0 # Not possible to calculate enthalpy of reaction HRuO5-
# Enthalpy of formation: -0 kcal/mol
1.0000 H+ + 1.0000 S2O3-- = HS2O3-
-llnl_gamma 4.0
log_k 1.0139
-delta_H 0 # Not possible to calculate enthalpy of reaction HS2O3-
# Enthalpy of formation: -0 kcal/mol
1.0000 SO3-- + 1.0000 H+ = HSO3-
-llnl_gamma 4.0
log_k +7.2054
-delta_H 9.33032 kJ/mol # Calculated enthalpy of reaction HSO3-
# Enthalpy of formation: -149.67 kcal/mol
-analytic 5.5899e+001 3.3623e-002 -5.0120e+002 -2.3040e+001 -7.8373e+000
# -Range: 0-300
1.0000 SO4-- + 1.0000 H+ = HSO4-
-llnl_gamma 4.0
log_k +1.9791
-delta_H 20.5016 kJ/mol # Calculated enthalpy of reaction HSO4-
# Enthalpy of formation: -212.5 kcal/mol
-analytic 4.9619e+001 3.0368e-002 -1.1558e+003 -2.1335e+001 -1.8051e+001
# -Range: 0-300
4.0000 HS- + 3.0000 H+ + 2.0000 Sb(OH)3 = HSb2S4- +6.0000 H2O
-llnl_gamma 4.0
log_k +50.6100
-delta_H 0 # Not possible to calculate enthalpy of reaction HSb2S4-
# Enthalpy of formation: -0 kcal/mol
-analytic 1.7540e+002 8.2177e-002 1.0786e+004 -7.4874e+001 1.6826e+002
# -Range: 0-300
1.0000 SeO3-- + 1.0000 H+ = HSeO3-
-llnl_gamma 4.0
log_k +7.2861
-delta_H -5.35552 kJ/mol # Calculated enthalpy of reaction HSeO3-
# Enthalpy of formation: -122.98 kcal/mol
-analytic 5.0427e+001 3.2250e-002 2.9603e+002 -2.1711e+001 4.6044e+000
# -Range: 0-300
1.0000 SeO4-- + 1.0000 H+ = HSeO4-
-llnl_gamma 4.0
log_k +1.9058
-delta_H 17.5728 kJ/mol # Calculated enthalpy of reaction HSeO4-
# Enthalpy of formation: -139 kcal/mol
-analytic 1.4160e+002 3.9801e-002 -4.5392e+003 -5.5088e+001 -7.0872e+001
# -Range: 0-300
1.0000 SiO2 + 1.0000 H2O = HSiO3- +1.0000 H+
-llnl_gamma 4.0
log_k -9.9525
-delta_H 25.991 kJ/mol # Calculated enthalpy of reaction HSiO3-
# Enthalpy of formation: -271.88 kcal/mol
-analytic 6.4211e+001 -2.4872e-002 -1.2707e+004 -1.4681e+001 1.0853e+006
# -Range: 0-300
1.0000 TcO4-- + 1.0000 H+ = HTcO4-
-llnl_gamma 4.0
log_k +8.7071
-delta_H 0 # Not possible to calculate enthalpy of reaction HTcO4-
# Enthalpy of formation: -0 kcal/mol
2.0000 H2O + 1.0000 VO2+ = HVO4-- +3.0000 H+
-llnl_gamma 4.0
log_k -15.1553
-delta_H 0 # Not possible to calculate enthalpy of reaction HVO4-2
# Enthalpy of formation: -0 kcal/mol
-analytic -7.0660e+001 -5.2457e-002 -3.5380e+003 3.3534e+001 -5.5186e+001
# -Range: 0-300
5.0000 H2O + 1.0000 Hf++++ = Hf(OH)5- +5.0000 H+
-llnl_gamma 4.0
log_k -17.1754
-delta_H 0 # Not possible to calculate enthalpy of reaction Hf(OH)5-
# Enthalpy of formation: -0 kcal/mol
1.0000 Hf++++ + 1.0000 H2O = HfOH+++ +1.0000 H+
-llnl_gamma 5.0
log_k -0.2951
-delta_H 0 # Not possible to calculate enthalpy of reaction HfOH+3
# Enthalpy of formation: -0 kcal/mol
2.0000 HAcetate + 1.0000 Hg++ = Hg(Acetate)2 +2.0000 H+
-llnl_gamma 3.0
log_k -2.6242
-delta_H -30.334 kJ/mol # Calculated enthalpy of reaction Hg(Acetate)2
# Enthalpy of formation: -198.78 kcal/mol
-analytic -2.1959e+001 2.7774e-003 -3.2500e+003 7.7351e+000 9.1508e+005
# -Range: 0-300
3.0000 HAcetate + 1.0000 Hg++ = Hg(Acetate)3- +3.0000 H+
-llnl_gamma 4.0
log_k -4.3247
-delta_H -59.7057 kJ/mol # Calculated enthalpy of reaction Hg(Acetate)3-
# Enthalpy of formation: -321.9 kcal/mol
-analytic 2.1656e+001 -2.0392e-003 -1.2866e+004 -3.2932e+000 2.3073e+006
# -Range: 0-300
1.0000 Hg++ + 1.0000 HAcetate = HgAcetate+ +1.0000 H+
-llnl_gamma 4.0
log_k -0.4691
-delta_H -16.5686 kJ/mol # Calculated enthalpy of reaction HgAcetate+
# Enthalpy of formation: -79.39 kcal/mol
-analytic -1.6355e+001 1.9446e-003 -2.6676e+002 5.1978e+000 2.9805e+005
# -Range: 0-300
2.0000 HAcetate + 1.0000 Ho+++ = Ho(Acetate)2+ +2.0000 H+
-llnl_gamma 4.0
log_k -4.9844
-delta_H -28.1583 kJ/mol # Calculated enthalpy of reaction Ho(Acetate)2+
# Enthalpy of formation: -407.93 kcal/mol
-analytic -2.7925e+001 2.5599e-003 -1.4779e+003 8.0785e+000 6.3736e+005
# -Range: 0-300
3.0000 HAcetate + 1.0000 Ho+++ = Ho(Acetate)3 +3.0000 H+
-llnl_gamma 3.0
log_k -8.3783
-delta_H -47.5721 kJ/mol # Calculated enthalpy of reaction Ho(Acetate)3
# Enthalpy of formation: -528.67 kcal/mol
-analytic -6.5547e+001 -1.1963e-004 -1.8887e+002 1.9796e+001 7.9041e+005
# -Range: 0-300
2.0000 HCO3- + 1.0000 Ho+++ = Ho(CO3)2- +2.0000 H+
-llnl_gamma 4.0
log_k -7.3576
-delta_H 0 # Not possible to calculate enthalpy of reaction Ho(CO3)2-
# Enthalpy of formation: -0 kcal/mol
2.0000 HPO4-- + 1.0000 Ho+++ = Ho(HPO4)2-
-llnl_gamma 4.0
log_k +9.9000
-delta_H 0 # Not possible to calculate enthalpy of reaction Ho(HPO4)2-
# Enthalpy of formation: -0 kcal/mol
2.0000 HPO4-- + 1.0000 Ho+++ = Ho(PO4)2--- +2.0000 H+
-llnl_gamma 4.0
log_k -3.3437
-delta_H 0 # Not possible to calculate enthalpy of reaction Ho(PO4)2-3
# Enthalpy of formation: -0 kcal/mol
2.0000 SO4-- + 1.0000 Ho+++ = Ho(SO4)2-
-llnl_gamma 4.0
log_k +4.9000
-delta_H 0 # Not possible to calculate enthalpy of reaction Ho(SO4)2-
# Enthalpy of formation: -0 kcal/mol
1.0000 Ho+++ + 1.0000 HAcetate = HoAcetate++ +1.0000 H+
-llnl_gamma 4.5
log_k -2.1184
-delta_H -14.3093 kJ/mol # Calculated enthalpy of reaction HoAcetate+2
# Enthalpy of formation: -288.52 kcal/mol
-analytic -1.8265e+001 1.0753e-003 -6.0695e+002 5.7211e+000 3.3055e+005
# -Range: 0-300
1.0000 Ho+++ + 1.0000 HCO3- = HoCO3+ +1.0000 H+
-llnl_gamma 4.0
log_k -2.2591
-delta_H 89.1108 kJ/mol # Calculated enthalpy of reaction HoCO3+
# Enthalpy of formation: -312.6 kcal/mol
-analytic 2.3773e+002 5.4448e-002 -6.9916e+003 -9.4063e+001 -1.0917e+002
# -Range: 0-300
1.0000 Ho+++ + 1.0000 Cl- = HoCl++
-llnl_gamma 4.5
log_k +0.2353
-delta_H 13.9453 kJ/mol # Calculated enthalpy of reaction HoCl+2
# Enthalpy of formation: -205.6 kcal/mol
-analytic 7.3746e+001 3.7733e-002 -1.5627e+003 -3.2126e+001 -2.4407e+001
# -Range: 0-300
2.0000 Cl- + 1.0000 Ho+++ = HoCl2+
-llnl_gamma 4.0
log_k -0.0425
-delta_H 17.8489 kJ/mol # Calculated enthalpy of reaction HoCl2+
# Enthalpy of formation: -244.6 kcal/mol
-analytic 1.9928e+002 7.9025e-002 -4.7775e+003 -8.3582e+001 -7.4607e+001
# -Range: 0-300
3.0000 Cl- + 1.0000 Ho+++ = HoCl3
-llnl_gamma 3.0
log_k -0.4669
-delta_H 10.0374 kJ/mol # Calculated enthalpy of reaction HoCl3
# Enthalpy of formation: -286.4 kcal/mol
-analytic 3.8608e+002 1.2638e-001 -9.8339e+003 -1.5809e+002 -1.5356e+002
# -Range: 0-300
4.0000 Cl- + 1.0000 Ho+++ = HoCl4-
-llnl_gamma 4.0
log_k -0.8913
-delta_H -12.4181 kJ/mol # Calculated enthalpy of reaction HoCl4-
# Enthalpy of formation: -331.7 kcal/mol
-analytic 4.2179e+002 1.2576e-001 -1.0495e+004 -1.7172e+002 -1.6388e+002
# -Range: 0-300
1.0000 Ho+++ + 1.0000 F- = HoF++
-llnl_gamma 4.5
log_k +4.7352
-delta_H 22.3844 kJ/mol # Calculated enthalpy of reaction HoF+2
# Enthalpy of formation: -243.8 kcal/mol
-analytic 9.5294e+001 4.1702e-002 -2.4460e+003 -3.8296e+001 -3.8195e+001
# -Range: 0-300
2.0000 F- + 1.0000 Ho+++ = HoF2+
-llnl_gamma 4.0
log_k +8.2976
-delta_H 11.7152 kJ/mol # Calculated enthalpy of reaction HoF2+
# Enthalpy of formation: -326.5 kcal/mol
-analytic 2.2330e+002 8.3497e-002 -4.9105e+003 -9.0272e+001 -7.6690e+001
# -Range: 0-300
3.0000 F- + 1.0000 Ho+++ = HoF3
-llnl_gamma 3.0
log_k +10.9071
-delta_H -12.7612 kJ/mol # Calculated enthalpy of reaction HoF3
# Enthalpy of formation: -412.5 kcal/mol
-analytic 4.1587e+002 1.3308e-001 -9.2193e+003 -1.6717e+002 -1.4398e+002
# -Range: 0-300
4.0000 F- + 1.0000 Ho+++ = HoF4-
-llnl_gamma 4.0
log_k +13.0035
-delta_H -57.7392 kJ/mol # Calculated enthalpy of reaction HoF4-
# Enthalpy of formation: -503.4 kcal/mol
-analytic 4.4575e+002 1.3182e-001 -8.5485e+003 -1.7916e+002 -1.3352e+002
# -Range: 0-300
1.0000 Ho+++ + 1.0000 HPO4-- + 1.0000 H+ = HoH2PO4++
-llnl_gamma 4.5
log_k +9.4484
-delta_H -17.9284 kJ/mol # Calculated enthalpy of reaction HoH2PO4+2
# Enthalpy of formation: -482.1 kcal/mol
-analytic 1.0273e+002 6.3161e-002 5.5160e+002 -4.6035e+001 8.5766e+000
# -Range: 0-300
1.0000 Ho+++ + 1.0000 HCO3- = HoHCO3++
-llnl_gamma 4.5
log_k +1.6991
-delta_H 7.52283 kJ/mol # Calculated enthalpy of reaction HoHCO3+2
# Enthalpy of formation: -332.1 kcal/mol
-analytic 3.3420e+001 3.1394e-002 1.9804e+002 -1.6859e+001 3.0801e+000
# -Range: 0-300
1.0000 Ho+++ + 1.0000 HPO4-- = HoHPO4+
-llnl_gamma 4.0
log_k +5.8000
-delta_H 0 # Not possible to calculate enthalpy of reaction HoHPO4+
# Enthalpy of formation: -0 kcal/mol
1.0000 NO3- + 1.0000 Ho+++ = HoNO3++
-llnl_gamma 4.5
log_k +0.2148
-delta_H -30.0035 kJ/mol # Calculated enthalpy of reaction HoNO3+2
# Enthalpy of formation: -225.6 kcal/mol
-analytic 1.1069e+001 2.5142e-002 2.3943e+003 -1.0650e+001 3.7358e+001
# -Range: 0-300
1.0000 Ho+++ + 1.0000 H2O = HoO+ +2.0000 H+
-llnl_gamma 4.0
log_k -16.0438
-delta_H 108.437 kJ/mol # Calculated enthalpy of reaction HoO+
# Enthalpy of formation: -211.4 kcal/mol
-analytic 1.9152e+002 3.0627e-002 -1.3817e+004 -6.8846e+001 -2.1565e+002
# -Range: 0-300
2.0000 H2O + 1.0000 Ho+++ = HoO2- +4.0000 H+
-llnl_gamma 4.0
log_k -33.4804
-delta_H 274.613 kJ/mol # Calculated enthalpy of reaction HoO2-
# Enthalpy of formation: -240 kcal/mol
-analytic 1.7987e+002 1.2731e-002 -2.0007e+004 -6.0642e+001 -3.1224e+002
# -Range: 0-300
2.0000 H2O + 1.0000 Ho+++ = HoO2H +3.0000 H+
-llnl_gamma 3.0
log_k -24.5377
-delta_H 216.873 kJ/mol # Calculated enthalpy of reaction HoO2H
# Enthalpy of formation: -253.8 kcal/mol
-analytic 3.3877e+002 4.6282e-002 -2.2925e+004 -1.2133e+002 -3.5782e+002
# -Range: 0-300
1.0000 Ho+++ + 1.0000 H2O = HoOH++ +1.0000 H+
-llnl_gamma 4.5
log_k -7.7609
-delta_H 76.6383 kJ/mol # Calculated enthalpy of reaction HoOH+2
# Enthalpy of formation: -219 kcal/mol
-analytic 7.1326e+001 1.2657e-002 -6.2461e+003 -2.5018e+001 -9.7485e+001
# -Range: 0-300
1.0000 Ho+++ + 1.0000 HPO4-- = HoPO4 +1.0000 H+
-llnl_gamma 3.0
log_k +0.2782
-delta_H 0 # Not possible to calculate enthalpy of reaction HoPO4
# Enthalpy of formation: -0 kcal/mol
1.0000 SO4-- + 1.0000 Ho+++ = HoSO4+
-llnl_gamma 4.0
log_k +3.5697
-delta_H 20.5016 kJ/mol # Calculated enthalpy of reaction HoSO4+
# Enthalpy of formation: -381.5 kcal/mol
-analytic 3.0709e+002 8.6579e-002 -9.0693e+003 -1.2078e+002 -1.4161e+002
# -Range: 0-300
2.0000 HAcetate + 1.0000 K+ = K(Acetate)2- +2.0000 H+
-llnl_gamma 4.0
log_k -10.2914
-delta_H -1.79912 kJ/mol # Calculated enthalpy of reaction K(Acetate)2-
# Enthalpy of formation: -292.9 kcal/mol
-analytic -2.3036e+002 -4.6369e-002 7.0305e+003 8.4997e+001 1.0977e+002
# -Range: 0-300
1.0000 K+ + 1.0000 Br- = KBr
-llnl_gamma 3.0
log_k -1.7372
-delta_H 12.5102 kJ/mol # Calculated enthalpy of reaction KBr
# Enthalpy of formation: -86.32 kcal/mol
-analytic 1.1320e+002 3.4227e-002 -3.6401e+003 -4.5633e+001 -5.6833e+001
# -Range: 0-300
1.0000 K+ + 1.0000 HAcetate = KAcetate +1.0000 H+
-llnl_gamma 3.0
log_k -5.0211
-delta_H 4.8116 kJ/mol # Calculated enthalpy of reaction KAcetate
# Enthalpy of formation: -175.22 kcal/mol
-analytic -2.6676e-001 -3.2675e-003 -1.7143e+003 -7.1907e-003 1.7726e+005
# -Range: 0-300
1.0000 K+ + 1.0000 Cl- = KCl
-llnl_gamma 3.0
log_k -1.4946
-delta_H 14.1963 kJ/mol # Calculated enthalpy of reaction KCl
# Enthalpy of formation: -96.81 kcal/mol
-analytic 1.3650e+002 3.8405e-002 -4.4014e+003 -5.4421e+001 -6.8721e+001
# -Range: 0-300
1.0000 K+ + 1.0000 HPO4-- = KHPO4-
-llnl_gamma 4.0
log_k +0.7800
-delta_H 0 # Not possible to calculate enthalpy of reaction KHPO4-
# Enthalpy of formation: -0 kcal/mol
1.0000 SO4-- + 1.0000 K+ + 1.0000 H+ = KHSO4
-llnl_gamma 3.0
log_k +0.8136
-delta_H 29.8319 kJ/mol # Calculated enthalpy of reaction KHSO4
# Enthalpy of formation: -270.54 kcal/mol
-analytic 1.2620e+002 5.7349e-002 -3.3670e+003 -5.3003e+001 -5.2576e+001
# -Range: 0-300
1.0000 K+ + 1.0000 I- = KI
-llnl_gamma 3.0
log_k -1.598
-delta_H 9.16296 kJ/mol # Calculated enthalpy of reaction KI
# Enthalpy of formation: -71.68 kcal/mol
-analytic 1.0816e+002 3.3683e-002 -3.2143e+003 -4.4054e+001 -5.0187e+001
# -Range: 0-300
1.0000 K+ + 1.0000 H2O = KOH +1.0000 H+
-llnl_gamma 3.0
log_k -14.46
-delta_H 0 # Not possible to calculate enthalpy of reaction KOH
# Enthalpy of formation: -0 kcal/mol
2.0000 HPO4-- + 1.0000 K+ = KP2O7--- +1.0000 H2O
-llnl_gamma 4.0
log_k -1.4286
-delta_H 34.1393 kJ/mol # Calculated enthalpy of reaction KP2O7-3
# Enthalpy of formation: -2516.36 kJ/mol
-analytic 4.1930e+002 1.4676e-001 -1.1169e+004 -1.7255e+002 -1.7441e+002
# -Range: 0-300
1.0000 SO4-- + 1.0000 K+ = KSO4-
-llnl_gamma 4.0
log_k +0.8796
-delta_H 2.88696 kJ/mol # Calculated enthalpy of reaction KSO4-
# Enthalpy of formation: -276.98 kcal/mol
-analytic 9.9073e+001 3.7817e-002 -2.1628e+003 -4.1297e+001 -3.3779e+001
# -Range: 0-300
2.0000 HAcetate + 1.0000 La+++ = La(Acetate)2+ +2.0000 H+
-llnl_gamma 4.0
log_k -5.3949
-delta_H -23.1375 kJ/mol # Calculated enthalpy of reaction La(Acetate)2+
# Enthalpy of formation: -407.33 kcal/mol
-analytic -1.2805e+001 2.8482e-003 -2.2521e+003 2.9108e+000 6.1659e+005
# -Range: 0-300
3.0000 HAcetate + 1.0000 La+++ = La(Acetate)3 +3.0000 H+
-llnl_gamma 3.0
log_k -8.5982
-delta_H -41.9237 kJ/mol # Calculated enthalpy of reaction La(Acetate)3
# Enthalpy of formation: -527.92 kcal/mol
-analytic -3.3456e+001 1.2371e-003 -1.5978e+003 8.6343e+000 7.5717e+005
# -Range: 0-300
2.0000 HCO3- + 1.0000 La+++ = La(CO3)2- +2.0000 H+
-llnl_gamma 4.0
log_k -8.8576
-delta_H 0 # Not possible to calculate enthalpy of reaction La(CO3)2-
# Enthalpy of formation: -0 kcal/mol
2.0000 HPO4-- + 1.0000 La+++ = La(HPO4)2-
-llnl_gamma 4.0
log_k +8.4000
-delta_H 0 # Not possible to calculate enthalpy of reaction La(HPO4)2-
# Enthalpy of formation: -0 kcal/mol
2.0000 HPO4-- + 1.0000 La+++ = La(PO4)2--- +2.0000 H+
-llnl_gamma 4.0
log_k -7.0437
-delta_H 0 # Not possible to calculate enthalpy of reaction La(PO4)2-3
# Enthalpy of formation: -0 kcal/mol
2.0000 SO4-- + 1.0000 La+++ = La(SO4)2-
-llnl_gamma 4.0
log_k +5.1000
-delta_H 0 # Not possible to calculate enthalpy of reaction La(SO4)2-
# Enthalpy of formation: -0 kcal/mol
2.0000 La+++ + 2.0000 H2O = La2(OH)2++++ +2.0000 H+
-llnl_gamma 5.5
log_k -22.9902
-delta_H 0 # Not possible to calculate enthalpy of reaction La2(OH)2+4
# Enthalpy of formation: -0 kcal/mol
9.0000 H2O + 5.0000 La+++ = La5(OH)9+6 +9.0000 H+
-llnl_gamma 6.0
log_k -71.1557
-delta_H 0 # Not possible to calculate enthalpy of reaction La5(OH)9+6
# Enthalpy of formation: -0 kcal/mol
1.0000 La+++ + 1.0000 HAcetate = LaAcetate++ +1.0000 H+
-llnl_gamma 4.5
log_k -2.2063
-delta_H -12.5938 kJ/mol # Calculated enthalpy of reaction LaAcetate+2
# Enthalpy of formation: -288.71 kcal/mol
-analytic -1.0803e+001 8.5239e-004 -1.1143e+003 3.3273e+000 3.4305e+005
# -Range: 0-300
1.0000 La+++ + 1.0000 HCO3- = LaCO3+ +1.0000 H+
-llnl_gamma 4.0
log_k -3.212
-delta_H 89.5292 kJ/mol # Calculated enthalpy of reaction LaCO3+
# Enthalpy of formation: -313.1 kcal/mol
-analytic 2.3046e+002 5.2419e-002 -7.1063e+003 -9.1109e+001 -1.1095e+002
# -Range: 0-300
1.0000 La+++ + 1.0000 Cl- = LaCl++
-llnl_gamma 4.5
log_k +0.3086
-delta_H 14.3637 kJ/mol # Calculated enthalpy of reaction LaCl+2
# Enthalpy of formation: -206.1 kcal/mol
-analytic 7.5802e+001 3.6641e-002 -1.7234e+003 -3.2578e+001 -2.6914e+001
# -Range: 0-300
2.0000 Cl- + 1.0000 La+++ = LaCl2+
-llnl_gamma 4.0
log_k -0.0425
-delta_H 19.1041 kJ/mol # Calculated enthalpy of reaction LaCl2+
# Enthalpy of formation: -244.9 kcal/mol
-analytic 2.1632e+002 7.9274e-002 -5.5883e+003 -8.9400e+001 -8.7264e+001
# -Range: 0-300
3.0000 Cl- + 1.0000 La+++ = LaCl3
-llnl_gamma 3.0
log_k -0.3936
-delta_H 12.5478 kJ/mol # Calculated enthalpy of reaction LaCl3
# Enthalpy of formation: -286.4 kcal/mol
-analytic 4.2210e+002 1.2792e-001 -1.1444e+004 -1.7062e+002 -1.7869e+002
# -Range: 0-300
4.0000 Cl- + 1.0000 La+++ = LaCl4-
-llnl_gamma 4.0
log_k -0.818
-delta_H -7.81571 kJ/mol # Calculated enthalpy of reaction LaCl4-
# Enthalpy of formation: -331.2 kcal/mol
-analytic 4.8802e+002 1.3053e-001 -1.3344e+004 -1.9518e+002 -2.0836e+002
# -Range: 0-300
1.0000 La+++ + 1.0000 F- = LaF++
-llnl_gamma 4.5
log_k +3.8556
-delta_H 26.5684 kJ/mol # Calculated enthalpy of reaction LaF+2
# Enthalpy of formation: -243.4 kcal/mol
-analytic 9.6765e+001 4.0513e-002 -2.8042e+003 -3.8617e+001 -4.3785e+001
# -Range: 0-300
2.0000 F- + 1.0000 La+++ = LaF2+
-llnl_gamma 4.0
log_k +6.6850
-delta_H 19.6648 kJ/mol # Calculated enthalpy of reaction LaF2+
# Enthalpy of formation: -325.2 kcal/mol
-analytic 2.3923e+002 8.3559e-002 -6.0536e+003 -9.5821e+001 -9.4531e+001
# -Range: 0-300
3.0000 F- + 1.0000 La+++ = LaF3
-llnl_gamma 3.0
log_k +8.7081
-delta_H -0.6276 kJ/mol # Calculated enthalpy of reaction LaF3
# Enthalpy of formation: -410.2 kcal/mol
-analytic 4.5123e+002 1.3460e-001 -1.1334e+004 -1.7967e+002 -1.7699e+002
# -Range: 0-300
4.0000 F- + 1.0000 La+++ = LaF4-
-llnl_gamma 4.0
log_k +10.3647
-delta_H -41.4216 kJ/mol # Calculated enthalpy of reaction LaF4-
# Enthalpy of formation: -500.1 kcal/mol
-analytic 5.0747e+002 1.3563e-001 -1.1903e+004 -2.0108e+002 -1.8588e+002
# -Range: 0-300
1.0000 La+++ + 1.0000 HPO4-- + 1.0000 H+ = LaH2PO4++
-llnl_gamma 4.5
log_k +9.7417
-delta_H -18.3468 kJ/mol # Calculated enthalpy of reaction LaH2PO4+2
# Enthalpy of formation: -482.8 kcal/mol
-analytic 1.0530e+002 6.2177e-002 4.0686e+002 -4.6642e+001 6.3174e+000
# -Range: 0-300
1.0000 La+++ + 1.0000 HCO3- = LaHCO3++
-llnl_gamma 4.5
log_k +1.9923
-delta_H 6.68603 kJ/mol # Calculated enthalpy of reaction LaHCO3+2
# Enthalpy of formation: -332.9 kcal/mol
-analytic 3.6032e+001 3.0405e-002 5.1281e+001 -1.7478e+001 7.8933e-001
# -Range: 0-300
1.0000 La+++ + 1.0000 HPO4-- = LaHPO4+
-llnl_gamma 4.0
log_k +5.1000
-delta_H 0 # Not possible to calculate enthalpy of reaction LaHPO4+
# Enthalpy of formation: -0 kcal/mol
1.0000 NO3- + 1.0000 La+++ = LaNO3++
-llnl_gamma 4.5
log_k +0.5813
-delta_H -29.1667 kJ/mol # Calculated enthalpy of reaction LaNO3+2
# Enthalpy of formation: -226 kcal/mol
-analytic 1.4136e+001 2.4247e-002 2.1998e+003 -1.1371e+001 3.4322e+001
# -Range: 0-300
1.0000 La+++ + 1.0000 H2O = LaO+ +2.0000 H+
-llnl_gamma 4.0
log_k -18.1696
-delta_H 121.407 kJ/mol # Calculated enthalpy of reaction LaO+
# Enthalpy of formation: -208.9 kcal/mol
-analytic 1.8691e+002 2.9275e-002 -1.4385e+004 -6.6906e+001 -2.2452e+002
# -Range: 0-300
2.0000 H2O + 1.0000 La+++ = LaO2- +4.0000 H+
-llnl_gamma 4.0
log_k -40.8105
-delta_H 318.126 kJ/mol # Calculated enthalpy of reaction LaO2-
# Enthalpy of formation: -230.2 kcal/mol
-analytic 1.8374e+002 1.2355e-002 -2.2472e+004 -6.1779e+001 -3.5070e+002
# -Range: 0-300
2.0000 H2O + 1.0000 La+++ = LaO2H +3.0000 H+
-llnl_gamma 3.0
log_k -27.9095
-delta_H 237.375 kJ/mol # Calculated enthalpy of reaction LaO2H
# Enthalpy of formation: -249.5 kcal/mol
-analytic 3.3862e+002 4.4808e-002 -2.4083e+004 -1.2088e+002 -3.7589e+002
# -Range: 0-300
1.0000 La+++ + 1.0000 H2O = LaOH++ +1.0000 H+
-llnl_gamma 4.5
log_k -8.6405
-delta_H 82.4959 kJ/mol # Calculated enthalpy of reaction LaOH+2
# Enthalpy of formation: -218.2 kcal/mol
-analytic 6.5529e+001 1.1104e-002 -6.3920e+003 -2.2646e+001 -9.9760e+001
# -Range: 0-300
1.0000 La+++ + 1.0000 HPO4-- = LaPO4 +1.0000 H+
-llnl_gamma 3.0
log_k -1.3618
-delta_H 0 # Not possible to calculate enthalpy of reaction LaPO4
# Enthalpy of formation: -0 kcal/mol
1.0000 SO4-- + 1.0000 La+++ = LaSO4+
-llnl_gamma 4.0
log_k +3.6430
-delta_H 18.4096 kJ/mol # Calculated enthalpy of reaction LaSO4+
# Enthalpy of formation: -382.6 kcal/mol
-analytic 3.0657e+002 8.4093e-002 -9.1074e+003 -1.2019e+002 -1.4220e+002
# -Range: 0-300
2.0000 HAcetate + 1.0000 Li+ = Li(Acetate)2- +2.0000 H+
-llnl_gamma 4.0
log_k -9.2674
-delta_H -24.7609 kJ/mol # Calculated enthalpy of reaction Li(Acetate)2-
# Enthalpy of formation: -304.67 kcal/mol
-analytic -3.3702e+002 -6.0849e-002 1.1952e+004 1.2359e+002 1.8659e+002
# -Range: 0-300
1.0000 Li+ + 1.0000 HAcetate = LiAcetate +1.0000 H+
-llnl_gamma 3.0
log_k -4.4589
-delta_H -6.64419 kJ/mol # Calculated enthalpy of reaction LiAcetate
# Enthalpy of formation: -184.24 kcal/mol
-analytic -3.8391e+000 -7.3938e-004 -1.0829e+003 3.4134e-001 2.1318e+005
# -Range: 0-300
1.0000 Li+ + 1.0000 Cl- = LiCl
-llnl_gamma 3.0
log_k -1.5115
-delta_H 3.36812 kJ/mol # Calculated enthalpy of reaction LiCl
# Enthalpy of formation: -105.68 kcal/mol
-analytic 1.2484e+002 4.1941e-002 -3.2439e+003 -5.1708e+001 -5.0655e+001
# -Range: 0-300
1.0000 Li+ + 1.0000 H2O = LiOH +1.0000 H+
-llnl_gamma 3.0
log_k -13.64
-delta_H 0 # Not possible to calculate enthalpy of reaction LiOH
# Enthalpy of formation: -0 kcal/mol
1.0000 SO4-- + 1.0000 Li+ = LiSO4-
-llnl_gamma 4.0
log_k +0.7700
-delta_H 0 # Not possible to calculate enthalpy of reaction LiSO4-
# Enthalpy of formation: -0 kcal/mol
2.0000 HAcetate + 1.0000 Lu+++ = Lu(Acetate)2+ +2.0000 H+
-llnl_gamma 4.0
log_k -4.9625
-delta_H -38.5346 kJ/mol # Calculated enthalpy of reaction Lu(Acetate)2+
# Enthalpy of formation: -409.31 kcal/mol
-analytic -2.7341e+001 2.5097e-003 -1.4157e+003 7.5026e+000 6.9682e+005
# -Range: 0-300
3.0000 HAcetate + 1.0000 Lu+++ = Lu(Acetate)3 +3.0000 H+
-llnl_gamma 3.0
log_k -8.3489
-delta_H -64.5173 kJ/mol # Calculated enthalpy of reaction Lu(Acetate)3
# Enthalpy of formation: -531.62 kcal/mol
-analytic -5.0225e+001 3.3508e-003 -6.2901e+002 1.3262e+001 9.0737e+005
# -Range: 0-300
2.0000 HCO3- + 1.0000 Lu+++ = Lu(CO3)2- +2.0000 H+
-llnl_gamma 4.0
log_k -6.8576
-delta_H 0 # Not possible to calculate enthalpy of reaction Lu(CO3)2-
# Enthalpy of formation: -0 kcal/mol
2.0000 HPO4-- + 1.0000 Lu+++ = Lu(HPO4)2-
-llnl_gamma 4.0
log_k +10.3000
-delta_H 0 # Not possible to calculate enthalpy of reaction Lu(HPO4)2-
# Enthalpy of formation: -0 kcal/mol
2.0000 HPO4-- + 1.0000 Lu+++ = Lu(PO4)2--- +2.0000 H+
-llnl_gamma 4.0
log_k -2.7437
-delta_H 0 # Not possible to calculate enthalpy of reaction Lu(PO4)2-3
# Enthalpy of formation: -0 kcal/mol
2.0000 SO4-- + 1.0000 Lu+++ = Lu(SO4)2-
-llnl_gamma 4.0
log_k +5.3000
-delta_H 0 # Not possible to calculate enthalpy of reaction Lu(SO4)2-
# Enthalpy of formation: -0 kcal/mol
1.0000 Lu+++ + 1.0000 HAcetate = LuAcetate++ +1.0000 H+
-llnl_gamma 4.5
log_k -2.1037
-delta_H -18.9703 kJ/mol # Calculated enthalpy of reaction LuAcetate+2
# Enthalpy of formation: -288.534 kcal/mol
-analytic -6.5982e+000 2.4512e-003 -1.2666e+003 1.4226e+000 4.0045e+005
# -Range: 0-300
1.0000 Lu+++ + 1.0000 HCO3- = LuCO3+ +1.0000 H+
-llnl_gamma 4.0
log_k -2.0392
-delta_H 78.2324 kJ/mol # Calculated enthalpy of reaction LuCO3+
# Enthalpy of formation: -314.1 kcal/mol
-analytic 2.3840e+002 5.4774e-002 -6.8317e+003 -9.4500e+001 -1.0667e+002
# -Range: 0-300
1.0000 Lu+++ + 1.0000 Cl- = LuCl++
-llnl_gamma 4.5
log_k -0.0579
-delta_H 13.5269 kJ/mol # Calculated enthalpy of reaction LuCl+2
# Enthalpy of formation: -204.6 kcal/mol
-analytic 6.6161e+001 3.6521e-002 -1.2938e+003 -2.9397e+001 -2.0209e+001
# -Range: 0-300
2.0000 Cl- + 1.0000 Lu+++ = LuCl2+
-llnl_gamma 4.0
log_k -0.6289
-delta_H 15.7569 kJ/mol # Calculated enthalpy of reaction LuCl2+
# Enthalpy of formation: -244 kcal/mol
-analytic 1.8608e+002 7.7283e-002 -4.2349e+003 -7.9007e+001 -6.6137e+001
# -Range: 0-300
3.0000 Cl- + 1.0000 Lu+++ = LuCl3
-llnl_gamma 3.0
log_k -1.1999
-delta_H 3.56895 kJ/mol # Calculated enthalpy of reaction LuCl3
# Enthalpy of formation: -286.846 kcal/mol
-analytic 3.7060e+002 1.2564e-001 -8.9374e+003 -1.5325e+002 -1.3957e+002
# -Range: 0-300
4.0000 Cl- + 1.0000 Lu+++ = LuCl4-
-llnl_gamma 4.0
log_k -1.771
-delta_H -25.8069 kJ/mol # Calculated enthalpy of reaction LuCl4-
# Enthalpy of formation: -333.8 kcal/mol
-analytic 3.8876e+002 1.2200e-001 -8.6965e+003 -1.6071e+002 -1.3582e+002
# -Range: 0-300
1.0000 Lu+++ + 1.0000 F- = LuF++
-llnl_gamma 4.5
log_k +4.8085
-delta_H 25.7316 kJ/mol # Calculated enthalpy of reaction LuF+2
# Enthalpy of formation: -241.9 kcal/mol
-analytic 9.0303e+001 4.0963e-002 -2.4140e+003 -3.6203e+001 -3.7694e+001
# -Range: 0-300
2.0000 F- + 1.0000 Lu+++ = LuF2+
-llnl_gamma 4.0
log_k +8.4442
-delta_H 14.2256 kJ/mol # Calculated enthalpy of reaction LuF2+
# Enthalpy of formation: -324.8 kcal/mol
-analytic 2.1440e+002 8.2559e-002 -4.7009e+003 -8.6790e+001 -7.3417e+001
# -Range: 0-300
3.0000 F- + 1.0000 Lu+++ = LuF3
-llnl_gamma 3.0
log_k +11.0999
-delta_H -12.3428 kJ/mol # Calculated enthalpy of reaction LuF3
# Enthalpy of formation: -411.3 kcal/mol
-analytic 4.0247e+002 1.3233e-001 -8.6775e+003 -1.6232e+002 -1.3552e+002
# -Range: 0-300
4.0000 F- + 1.0000 Lu+++ = LuF4-
-llnl_gamma 4.0
log_k +13.2967
-delta_H -64.0152 kJ/mol # Calculated enthalpy of reaction LuF4-
# Enthalpy of formation: -503.8 kcal/mol
-analytic 4.2541e+002 1.3070e-001 -7.4276e+003 -1.7220e+002 -1.1603e+002
# -Range: 0-300
1.0000 Lu+++ + 1.0000 HPO4-- + 1.0000 H+ = LuH2PO4++
-llnl_gamma 4.5
log_k +9.5950
-delta_H -23.786 kJ/mol # Calculated enthalpy of reaction LuH2PO4+2
# Enthalpy of formation: -482.4 kcal/mol
-analytic 9.4223e+001 6.1797e-002 1.1102e+003 -4.3131e+001 1.7296e+001
# -Range: 0-300
1.0000 Lu+++ + 1.0000 HCO3- = LuHCO3++
-llnl_gamma 4.5
log_k +1.9190
-delta_H 1.66523 kJ/mol # Calculated enthalpy of reaction LuHCO3+2
# Enthalpy of formation: -332.4 kcal/mol
-analytic 2.3187e+001 2.9604e-002 8.1268e+002 -1.3252e+001 1.2674e+001
# -Range: 0-300
1.0000 Lu+++ + 1.0000 HPO4-- = LuHPO4+
-llnl_gamma 4.0
log_k +6.0000
-delta_H 0 # Not possible to calculate enthalpy of reaction LuHPO4+
# Enthalpy of formation: -0 kcal/mol
1.0000 NO3- + 1.0000 Lu+++ = LuNO3++
-llnl_gamma 4.5
log_k +0.5813
-delta_H -41.7187 kJ/mol # Calculated enthalpy of reaction LuNO3+2
# Enthalpy of formation: -227.3 kcal/mol
-analytic 1.7412e+000 2.3703e-002 3.2605e+003 -7.7334e+000 5.0876e+001
# -Range: 0-300
1.0000 Lu+++ + 1.0000 H2O = LuO+ +2.0000 H+
-llnl_gamma 4.0
log_k -15.3108
-delta_H 99.6503 kJ/mol # Calculated enthalpy of reaction LuO+
# Enthalpy of formation: -212.4 kcal/mol
-analytic 1.5946e+002 2.6603e-002 -1.2215e+004 -5.7276e+001 -1.9065e+002
# -Range: 0-300
2.0000 H2O + 1.0000 Lu+++ = LuO2- +4.0000 H+
-llnl_gamma 4.0
log_k -31.9411
-delta_H 258.713 kJ/mol # Calculated enthalpy of reaction LuO2-
# Enthalpy of formation: -242.7 kcal/mol
-analytic 1.1522e+002 5.0221e-003 -1.6847e+004 -3.7244e+001 -2.6292e+002
# -Range: 0-300
2.0000 H2O + 1.0000 Lu+++ = LuO2H +3.0000 H+
-llnl_gamma 3.0
log_k -23.878
-delta_H 206.832 kJ/mol # Calculated enthalpy of reaction LuO2H
# Enthalpy of formation: -255.1 kcal/mol
-analytic 2.8768e+002 4.2338e-002 -2.0443e+004 -1.0330e+002 -3.1907e+002
# -Range: 0-300
1.0000 Lu+++ + 1.0000 H2O = LuOH++ +1.0000 H+
-llnl_gamma 4.5
log_k -7.6143
-delta_H 72.0359 kJ/mol # Calculated enthalpy of reaction LuOH+2
# Enthalpy of formation: -219 kcal/mol
-analytic 4.2937e+001 9.2421e-003 -4.9953e+003 -1.4769e+001 -7.7960e+001
# -Range: 0-300
1.0000 Lu+++ + 1.0000 HPO4-- = LuPO4 +1.0000 H+
-llnl_gamma 3.0
log_k +0.6782
-delta_H 0 # Not possible to calculate enthalpy of reaction LuPO4
# Enthalpy of formation: -0 kcal/mol
1.0000 SO4-- + 1.0000 Lu+++ = LuSO4+
-llnl_gamma 4.0
log_k +3.5697
-delta_H 19.5393 kJ/mol # Calculated enthalpy of reaction LuSO4+
# Enthalpy of formation: -380.63 kcal/mol
-analytic 3.0108e+002 8.5238e-002 -8.8411e+003 -1.1850e+002 -1.3805e+002
# -Range: 0-300
2.0000 HAcetate + 1.0000 Mg++ = Mg(Acetate)2 +2.0000 H+
-llnl_gamma 3.0
log_k -7.473
-delta_H -23.8195 kJ/mol # Calculated enthalpy of reaction Mg(Acetate)2
# Enthalpy of formation: -349.26 kcal/mol
-analytic -4.3954e+001 -3.1842e-004 -1.2033e+003 1.3556e+001 6.3058e+005
# -Range: 0-300
4.0000 Mg++ + 4.0000 H2O = Mg4(OH)4++++ +4.0000 H+
-llnl_gamma 5.5
log_k -39.75
-delta_H 0 # Not possible to calculate enthalpy of reaction Mg4(OH)4+4
# Enthalpy of formation: -0 kcal/mol
1.0000 Mg++ + 1.0000 H2O + 1.0000 B(OH)3 = MgB(OH)4+ +1.0000 H+
-llnl_gamma 4.0
log_k -7.3467
-delta_H 0 # Not possible to calculate enthalpy of reaction MgB(OH)4+
# Enthalpy of formation: -0 kcal/mol
1.0000 Mg++ + 1.0000 HAcetate = MgAcetate+ +1.0000 H+
-llnl_gamma 4.0
log_k -3.4781
-delta_H -8.42239 kJ/mol # Calculated enthalpy of reaction MgAcetate+
# Enthalpy of formation: -229.48 kcal/mol
-analytic -2.3548e+001 -1.6071e-003 -4.2228e+002 7.7009e+000 2.5981e+005
# -Range: 0-300
1.0000 Mg++ + 1.0000 HCO3- = MgCO3 +1.0000 H+
-llnl_gamma 3.0
log_k -7.3499
-delta_H 23.8279 kJ/mol # Calculated enthalpy of reaction MgCO3
# Enthalpy of formation: -270.57 kcal/mol
-analytic 2.3465e+002 5.5538e-002 -8.3947e+003 -9.3104e+001 -1.3106e+002
# -Range: 0-300
1.0000 Mg++ + 1.0000 Cl- = MgCl+
-llnl_gamma 4.0
log_k -0.1349
-delta_H -0.58576 kJ/mol # Calculated enthalpy of reaction MgCl+
# Enthalpy of formation: -151.44 kcal/mol
-analytic 4.3363e+001 3.2858e-002 1.1878e+002 -2.1688e+001 1.8403e+000
# -Range: 0-300
1.0000 Mg++ + 1.0000 F- = MgF+
-llnl_gamma 4.0
log_k +1.3524
-delta_H 2.37233 kJ/mol # Calculated enthalpy of reaction MgF+
# Enthalpy of formation: -190.95 kcal/mol
-analytic 6.4311e+001 3.5184e-002 -7.3241e+002 -2.8678e+001 -1.1448e+001
# -Range: 0-300
1.0000 Mg++ + 1.0000 HPO4-- + 1.0000 H+ = MgH2PO4+
-llnl_gamma 4.0
log_k +1.6600
-delta_H 0 # Not possible to calculate enthalpy of reaction MgH2PO4+
# Enthalpy of formation: -0 kcal/mol
1.0000 Mg++ + 1.0000 HCO3- = MgHCO3+
-llnl_gamma 4.0
log_k +1.0357
-delta_H 2.15476 kJ/mol # Calculated enthalpy of reaction MgHCO3+
# Enthalpy of formation: -275.75 kcal/mol
-analytic 3.8459e+001 3.0076e-002 9.8068e+001 -1.8869e+001 1.5187e+000
# -Range: 0-300
1.0000 Mg++ + 1.0000 HPO4-- = MgHPO4
-llnl_gamma 3.0
log_k +2.9100
-delta_H 0 # Not possible to calculate enthalpy of reaction MgHPO4
# Enthalpy of formation: -0 kcal/mol
2.0000 HPO4-- + 1.0000 Mg++ = MgP2O7-- +1.0000 H2O
-llnl_gamma 4.0
log_k +3.4727
-delta_H 38.5451 kJ/mol # Calculated enthalpy of reaction MgP2O7-2
# Enthalpy of formation: -2725.74 kJ/mol
-analytic 4.8038e+002 1.2530e-001 -1.5175e+004 -1.8724e+002 -2.3693e+002
# -Range: 0-300
1.0000 Mg++ + 1.0000 HPO4-- = MgPO4- +1.0000 H+
-llnl_gamma 4.0
log_k -5.7328
-delta_H 0 # Not possible to calculate enthalpy of reaction MgPO4-
# Enthalpy of formation: -0 kcal/mol
1.0000 SO4-- + 1.0000 Mg++ = MgSO4
-llnl_gamma 3.0
log_k +2.4117
-delta_H 19.6051 kJ/mol # Calculated enthalpy of reaction MgSO4
# Enthalpy of formation: -1355.96 kJ/mol
-analytic 1.7994e+002 6.4715e-002 -4.7314e+003 -7.3123e+001 -8.0408e+001
# -Range: 0-200
2.0000 HAcetate + 1.0000 Mn++ = Mn(Acetate)2 +2.0000 H+
-llnl_gamma 3.0
log_k -7.4547
-delta_H -11.4893 kJ/mol # Calculated enthalpy of reaction Mn(Acetate)2
# Enthalpy of formation: -287.67 kcal/mol
-analytic -9.0558e-001 5.9656e-003 -4.3531e+003 -1.1063e+000 8.0323e+005
# -Range: 0-300
3.0000 HAcetate + 1.0000 Mn++ = Mn(Acetate)3- +3.0000 H+
-llnl_gamma 4.0
log_k -11.8747
-delta_H -30.3591 kJ/mol # Calculated enthalpy of reaction Mn(Acetate)3-
# Enthalpy of formation: -408.28 kcal/mol
-analytic -3.8531e+000 -9.9140e-003 -1.2065e+004 5.1424e+000 2.0175e+006
# -Range: 0-300
2.0000 NO3- + 1.0000 Mn++ = Mn(NO3)2
-llnl_gamma 3.0
log_k +0.6000
-delta_H 0 # Not possible to calculate enthalpy of reaction Mn(NO3)2
# Enthalpy of formation: -0 kcal/mol
2.0000 H2O + 1.0000 Mn++ = Mn(OH)2 +2.0000 H+
-llnl_gamma 3.0
log_k -22.2
-delta_H 0 # Not possible to calculate enthalpy of reaction Mn(OH)2
# Enthalpy of formation: -0 kcal/mol
3.0000 H2O + 1.0000 Mn++ = Mn(OH)3- +3.0000 H+
-llnl_gamma 4.0
log_k -34.2278
-delta_H 0 # Not possible to calculate enthalpy of reaction Mn(OH)3-
# Enthalpy of formation: -0 kcal/mol
4.0000 H2O + 1.0000 Mn++ = Mn(OH)4-- +4.0000 H+
-llnl_gamma 4.0
log_k -48.3
-delta_H 0 # Not possible to calculate enthalpy of reaction Mn(OH)4-2
# Enthalpy of formation: -0 kcal/mol
3.0000 H2O + 2.0000 Mn++ = Mn2(OH)3+ +3.0000 H+
-llnl_gamma 4.0
log_k -23.9
-delta_H 0 # Not possible to calculate enthalpy of reaction Mn2(OH)3+
# Enthalpy of formation: -0 kcal/mol
2.0000 Mn++ + 1.0000 H2O = Mn2OH+++ +1.0000 H+
-llnl_gamma 5.0
log_k -10.56
-delta_H 0 # Not possible to calculate enthalpy of reaction Mn2OH+3
# Enthalpy of formation: -0 kcal/mol
1.0000 Mn++ + 1.0000 HAcetate = MnAcetate+ +1.0000 H+
-llnl_gamma 4.0
log_k -3.5404
-delta_H -3.07942 kJ/mol # Calculated enthalpy of reaction MnAcetate+
# Enthalpy of formation: -169.56 kcal/mol
-analytic -1.4061e+001 1.8149e-003 -8.6438e+002 4.0354e+000 2.5831e+005
# -Range: 0-300
1.0000 Mn++ + 1.0000 HCO3- = MnCO3 +1.0000 H+
-llnl_gamma 3.0
log_k -5.8088
-delta_H 0 # Not possible to calculate enthalpy of reaction MnCO3
# Enthalpy of formation: -0 kcal/mol
1.0000 Mn++ + 1.0000 Cl- = MnCl+
-llnl_gamma 4.0
log_k +0.3013
-delta_H 18.3134 kJ/mol # Calculated enthalpy of reaction MnCl+
# Enthalpy of formation: -88.28 kcal/mol
-analytic 8.7072e+001 4.0361e-002 -2.1786e+003 -3.6966e+001 -3.4022e+001
# -Range: 0-300
3.0000 Cl- + 1.0000 Mn++ = MnCl3-
-llnl_gamma 4.0
log_k -0.3324
-delta_H 0 # Not possible to calculate enthalpy of reaction MnCl3-
# Enthalpy of formation: -0 kcal/mol
1.0000 Mn++ + 1.0000 F- = MnF+
-llnl_gamma 4.0
log_k +1.4300
-delta_H 0 # Not possible to calculate enthalpy of reaction MnF+
# Enthalpy of formation: -0 kcal/mol
1.0000 Mn++ + 1.0000 HPO4-- + 1.0000 H+ = MnH2PO4+
-llnl_gamma 4.0
log_k +8.5554
-delta_H 0 # Not possible to calculate enthalpy of reaction MnH2PO4+
# Enthalpy of formation: -0 kcal/mol
1.0000 Mn++ + 1.0000 HCO3- = MnHCO3+
-llnl_gamma 4.0
log_k +0.8816
-delta_H 0 # Not possible to calculate enthalpy of reaction MnHCO3+
# Enthalpy of formation: -0 kcal/mol
1.0000 Mn++ + 1.0000 HPO4-- = MnHPO4
-llnl_gamma 3.0
log_k +3.5800
-delta_H 0 # Not possible to calculate enthalpy of reaction MnHPO4
# Enthalpy of formation: -0 kcal/mol
1.0000 NO3- + 1.0000 Mn++ = MnNO3+
-llnl_gamma 4.0
log_k +0.2000
-delta_H 0 # Not possible to calculate enthalpy of reaction MnNO3+
# Enthalpy of formation: -0 kcal/mol
1.5000 H2O + 1.2500 O2 + 1.0000 Mn++ = MnO4- +3.0000 H+
-llnl_gamma 3.5
log_k -20.2963
-delta_H 123.112 kJ/mol # Calculated enthalpy of reaction MnO4-
# Enthalpy of formation: -129.4 kcal/mol
-analytic 1.8544e+001 -1.7618e-002 -6.7332e+003 -3.3193e+000 -2.4924e+005
# -Range: 0-300
1.0000 Mn++ + 1.0000 H2O = MnOH+ +1.0000 H+
-llnl_gamma 4.0
log_k -10.59
-delta_H 0 # Not possible to calculate enthalpy of reaction MnOH+
# Enthalpy of formation: -0 kcal/mol
1.0000 Mn++ + 1.0000 HPO4-- = MnPO4- +1.0000 H+
-llnl_gamma 4.0
log_k -5.1318
-delta_H 0 # Not possible to calculate enthalpy of reaction MnPO4-
# Enthalpy of formation: -0 kcal/mol
1.0000 SO4-- + 1.0000 Mn++ = MnSO4
-llnl_gamma 3.0
log_k +2.3529
-delta_H 14.1168 kJ/mol # Calculated enthalpy of reaction MnSO4
# Enthalpy of formation: -266.75 kcal/mol
-analytic 2.9448e+002 8.5294e-002 -8.1366e+003 -1.1729e+002 -1.2705e+002
# -Range: 0-300
1.0000 SeO4-- + 1.0000 Mn++ = MnSeO4
-llnl_gamma 3.0
log_k +2.4300
-delta_H 0 # Not possible to calculate enthalpy of reaction MnSeO4
# Enthalpy of formation: -0 kcal/mol
2.0000 HAcetate + 1.0000 NH3 = NH4(Acetate)2- +1.0000 H+
-llnl_gamma 4.0
log_k -0.1928
-delta_H -56.735 kJ/mol # Calculated enthalpy of reaction NH4(Acetate)2-
# Enthalpy of formation: -265.2 kcal/mol
-analytic 3.7137e+001 -1.2242e-002 -8.4764e+003 -8.4308e+000 1.3883e+006
# -Range: 0-300
1.0000 NH3 + 1.0000 H+ = NH4+
-llnl_gamma 2.5
log_k +9.2410
-delta_H -51.9234 kJ/mol # Calculated enthalpy of reaction NH4+
# Enthalpy of formation: -31.85 kcal/mol
-analytic -1.4527e+001 -5.0518e-003 3.0447e+003 6.0865e+000 4.7515e+001
# -Range: 0-300
1.0000 NH3 + 1.0000 HAcetate = NH4Acetate
-llnl_gamma 3.0
log_k +4.6964
-delta_H -48.911 kJ/mol # Calculated enthalpy of reaction NH4Acetate
# Enthalpy of formation: -147.23 kcal/mol
-analytic 1.4104e+001 -4.3664e-003 -1.0746e+003 -3.6999e+000 4.1428e+005
# -Range: 0-300
1.0000 SO4-- + 1.0000 NH3 + 1.0000 H+ = NH4SO4-
-llnl_gamma 4.0
log_k +0.9400
-delta_H 0 # Not possible to calculate enthalpy of reaction NH4SO4-
# Enthalpy of formation: -0 kcal/mol
1.0000 Sb(OH)3 + 1.0000 NH3 = NH4SbO2 +1.0000 H2O
-llnl_gamma 3.0
log_k -2.5797
-delta_H 0 # Not possible to calculate enthalpy of reaction NH4SbO2
# Enthalpy of formation: -0 kcal/mol
2.0000 HAcetate + 1.0000 Na+ = Na(Acetate)2- +2.0000 H+
-llnl_gamma 4.0
log_k -9.9989
-delta_H -11.5771 kJ/mol # Calculated enthalpy of reaction Na(Acetate)2-
# Enthalpy of formation: -292.4 kcal/mol
-analytic -2.9232e+002 -5.5708e-002 9.6601e+003 1.0772e+002 1.5082e+002
# -Range: 0-300
1.0000 O_phthalate-2 + 1.0000 Na+ = Na(O_phthalate)-
-llnl_gamma 4.0
log_k +0.7000
-delta_H 0 # Not possible to calculate enthalpy of reaction Na(O_phthalate)-
# Enthalpy of formation: -0 kcal/mol
2.0000 Na+ + 2.0000 HPO4-- = Na2P2O7-- +1.0000 H2O
-llnl_gamma 4.0
log_k +0.4437
-delta_H 0 # Not possible to calculate enthalpy of reaction Na2P2O7-2
# Enthalpy of formation: -0 kcal/mol
2.0000 H2O + 1.0000 Na+ + 1.0000 Al+++ = NaAlO2 +4.0000 H+
-llnl_gamma 3.0
log_k -23.6266
-delta_H 190.326 kJ/mol # Calculated enthalpy of reaction NaAlO2
# Enthalpy of formation: -277.259 kcal/mol
-analytic 1.2288e+002 3.4921e-002 -1.2808e+004 -4.6046e+001 -1.9990e+002
# -Range: 0-300
1.0000 Na+ + 1.0000 H2O + 1.0000 B(OH)3 = NaB(OH)4 +1.0000 H+
-llnl_gamma 3.0
log_k -8.974
-delta_H 0 # Not possible to calculate enthalpy of reaction NaB(OH)4
# Enthalpy of formation: -0 kcal/mol
1.0000 Na+ + 1.0000 Br- = NaBr
-llnl_gamma 3.0
log_k -1.3568
-delta_H 6.87431 kJ/mol # Calculated enthalpy of reaction NaBr
# Enthalpy of formation: -84.83 kcal/mol
-analytic 1.1871e+002 3.7271e-002 -3.4061e+003 -4.8386e+001 -5.3184e+001
# -Range: 0-300
1.0000 Na+ + 1.0000 HAcetate = NaAcetate +1.0000 H+
-llnl_gamma 3.0
log_k -4.8606
-delta_H -0.029288 kJ/mol # Calculated enthalpy of reaction NaAcetate
# Enthalpy of formation: -173.54 kcal/mol
-analytic 6.4833e+000 -1.8739e-003 -2.0902e+003 -2.6121e+000 2.3990e+005
# -Range: 0-300
1.0000 Na+ + 1.0000 HCO3- = NaCO3- +1.0000 H+
-llnl_gamma 4.0
log_k -9.8144
-delta_H -5.6521 kJ/mol # Calculated enthalpy of reaction NaCO3-
# Enthalpy of formation: -935.885 kJ/mol
-analytic 1.6939e+002 5.3122e-004 -7.6768e+003 -6.2078e+001 -1.1984e+002
# -Range: 0-300
1.0000 Na+ + 1.0000 Cl- = NaCl
-llnl_gamma 3.0
log_k -0.777
-delta_H 5.21326 kJ/mol # Calculated enthalpy of reaction NaCl
# Enthalpy of formation: -96.12 kcal/mol
-analytic 1.1398e+002 3.6386e-002 -3.0847e+003 -4.6571e+001 -4.8167e+001
# -Range: 0-300
1.0000 Na+ + 1.0000 F- = NaF
-llnl_gamma 3.0
log_k -0.9976
-delta_H 7.20903 kJ/mol # Calculated enthalpy of reaction NaF
# Enthalpy of formation: -135.86 kcal/mol
-analytic 1.2507e+002 3.8619e-002 -3.5436e+003 -5.0787e+001 -5.5332e+001
# -Range: 0-300
1.0000 Na+ + 1.0000 HCO3- = NaHCO3
-llnl_gamma 3.0
log_k +0.1541
-delta_H -13.7741 kJ/mol # Calculated enthalpy of reaction NaHCO3
# Enthalpy of formation: -944.007 kJ/mol
-analytic -9.0668e+001 -2.9866e-002 2.7947e+003 3.6515e+001 4.7489e+001
# -Range: 0-200
2.0000 HPO4-- + 1.0000 Na+ + 1.0000 H+ = NaHP2O7-- +1.0000 H2O
-llnl_gamma 4.0
log_k +6.8498
-delta_H 0 # Not possible to calculate enthalpy of reaction NaHP2O7-2
# Enthalpy of formation: -0 kcal/mol
1.0000 Na+ + 1.0000 HPO4-- = NaHPO4-
-llnl_gamma 4.0
log_k +0.9200
-delta_H 0 # Not possible to calculate enthalpy of reaction NaHPO4-
# Enthalpy of formation: -0 kcal/mol
1.0000 SiO2 + 1.0000 Na+ + 1.0000 H2O = NaHSiO3 +1.0000 H+
-llnl_gamma 3.0
log_k -8.304
-delta_H 11.6524 kJ/mol # Calculated enthalpy of reaction NaHSiO3
# Enthalpy of formation: -332.74 kcal/mol
-analytic 3.6045e+001 -9.0411e-003 -6.6605e+003 -1.0447e+001 5.8415e+005
# -Range: 0-300
1.0000 Na+ + 1.0000 I- = NaI
-llnl_gamma 3.0
log_k -1.54
-delta_H 7.33455 kJ/mol # Calculated enthalpy of reaction NaI
# Enthalpy of formation: -69.28 kcal/mol
-analytic 9.8742e+001 3.2917e-002 -2.7576e+003 -4.0748e+001 -4.3058e+001
# -Range: 0-300
1.0000 Na+ + 1.0000 H2O = NaOH +1.0000 H+
-llnl_gamma 3.0
log_k -14.7948
-delta_H 53.6514 kJ/mol # Calculated enthalpy of reaction NaOH
# Enthalpy of formation: -112.927 kcal/mol
-analytic 8.7326e+001 2.3555e-002 -5.4770e+003 -3.6678e+001 -8.5489e+001
# -Range: 0-300
2.0000 HPO4-- + 1.0000 Na+ = NaP2O7--- +1.0000 H2O
-llnl_gamma 4.0
log_k -1.4563
-delta_H 0 # Not possible to calculate enthalpy of reaction NaP2O7-3
# Enthalpy of formation: -0 kcal/mol
1.0000 SO4-- + 1.0000 Na+ = NaSO4-
-llnl_gamma 4.0
log_k +0.8200
-delta_H 0 # Not possible to calculate enthalpy of reaction NaSO4-
# Enthalpy of formation: -0 kcal/mol
2.0000 HAcetate + 1.0000 Nd+++ = Nd(Acetate)2+ +2.0000 H+
-llnl_gamma 4.0
log_k -4.9771
-delta_H -22.6354 kJ/mol # Calculated enthalpy of reaction Nd(Acetate)2+
# Enthalpy of formation: -404.11 kcal/mol
-analytic -2.2128e+001 1.0975e-003 -7.1543e+002 5.8799e+000 4.1748e+005
# -Range: 0-300
3.0000 HAcetate + 1.0000 Nd+++ = Nd(Acetate)3 +3.0000 H+
-llnl_gamma 3.0
log_k -8.2976
-delta_H -38.8694 kJ/mol # Calculated enthalpy of reaction Nd(Acetate)3
# Enthalpy of formation: -524.09 kcal/mol
-analytic -4.5726e+001 -2.6143e-003 5.9389e+002 1.2679e+001 4.3320e+005
# -Range: 0-300
2.0000 HCO3- + 1.0000 Nd+++ = Nd(CO3)2- +2.0000 H+
-llnl_gamma 4.0
log_k -8.0576
-delta_H 0 # Not possible to calculate enthalpy of reaction Nd(CO3)2-
# Enthalpy of formation: -0 kcal/mol
2.0000 HPO4-- + 1.0000 Nd+++ = Nd(HPO4)2-
-llnl_gamma 4.0
log_k +9.1000
-delta_H 0 # Not possible to calculate enthalpy of reaction Nd(HPO4)2-
# Enthalpy of formation: -0 kcal/mol
# Redundant with NdO2-
#4.0000 H2O + 1.0000 Nd+++ = Nd(OH)4- +4.0000 H+
# -llnl_gamma 4.0
# log_k -37.0803
# -delta_H 0 # Not possible to calculate enthalpy of reaction Nd(OH)4-
## Enthalpy of formation: -0 kcal/mol
2.0000 HPO4-- + 1.0000 Nd+++ = Nd(PO4)2--- +2.0000 H+
-llnl_gamma 4.0
log_k -5.1437
-delta_H 0 # Not possible to calculate enthalpy of reaction Nd(PO4)2-3
# Enthalpy of formation: -0 kcal/mol
2.0000 SO4-- + 1.0000 Nd+++ = Nd(SO4)2-
-llnl_gamma 4.0
log_k -255.7478
-delta_H 0 # Not possible to calculate enthalpy of reaction Nd(SO4)2-
# Enthalpy of formation: -0 kcal/mol
2.0000 Nd+++ + 2.0000 H2O = Nd2(OH)2++++ +2.0000 H+
-llnl_gamma 5.5
log_k -13.8902
-delta_H 0 # Not possible to calculate enthalpy of reaction Nd2(OH)2+4
# Enthalpy of formation: -0 kcal/mol
1.0000 Nd+++ + 1.0000 HAcetate = NdAcetate++ +1.0000 H+
-llnl_gamma 4.5
log_k -2.0891
-delta_H -12.0081 kJ/mol # Calculated enthalpy of reaction NdAcetate+2
# Enthalpy of formation: -285.47 kcal/mol
-analytic -1.6006e+001 4.1948e-004 -3.6469e+002 4.9280e+000 2.5187e+005
# -Range: 0-300
1.0000 Nd+++ + 1.0000 HCO3- = NdCO3+ +1.0000 H+
-llnl_gamma 4.0
log_k -2.6256
-delta_H 91.6212 kJ/mol # Calculated enthalpy of reaction NdCO3+
# Enthalpy of formation: -309.5 kcal/mol
-analytic 2.3399e+002 5.3454e-002 -7.0513e+003 -9.2500e+001 -1.1010e+002
# -Range: 0-300
1.0000 Nd+++ + 1.0000 Cl- = NdCl++
-llnl_gamma 4.5
log_k +0.3086
-delta_H 14.3637 kJ/mol # Calculated enthalpy of reaction NdCl+2
# Enthalpy of formation: -203 kcal/mol
-analytic 9.4587e+001 3.9331e-002 -2.4200e+003 -3.9550e+001 -3.7790e+001
# -Range: 0-300
2.0000 Cl- + 1.0000 Nd+++ = NdCl2+
-llnl_gamma 4.0
log_k +0.0308
-delta_H 20.3593 kJ/mol # Calculated enthalpy of reaction NdCl2+
# Enthalpy of formation: -241.5 kcal/mol
-analytic 2.5840e+002 8.4118e-002 -7.2056e+003 -1.0477e+002 -1.1251e+002
# -Range: 0-300
3.0000 Cl- + 1.0000 Nd+++ = NdCl3
-llnl_gamma 3.0
log_k -0.3203
-delta_H 15.0582 kJ/mol # Calculated enthalpy of reaction NdCl3
# Enthalpy of formation: -282.7 kcal/mol
-analytic 4.9362e+002 1.3485e-001 -1.4309e+004 -1.9645e+002 -2.2343e+002
# -Range: 0-300
4.0000 Cl- + 1.0000 Nd+++ = NdCl4-
-llnl_gamma 4.0
log_k -0.7447
-delta_H -3.21331 kJ/mol # Calculated enthalpy of reaction NdCl4-
# Enthalpy of formation: -327 kcal/mol
-analytic 6.0548e+002 1.4227e-001 -1.8055e+004 -2.3765e+002 -2.8191e+002
# -Range: 0-300
1.0000 Nd+++ + 1.0000 F- = NdF++
-llnl_gamma 4.5
log_k +4.3687
-delta_H 22.8028 kJ/mol # Calculated enthalpy of reaction NdF+2
# Enthalpy of formation: -241.2 kcal/mol
-analytic 1.1461e+002 4.3014e-002 -3.2461e+003 -4.5326e+001 -5.0687e+001
# -Range: 0-300
2.0000 F- + 1.0000 Nd+++ = NdF2+
-llnl_gamma 4.0
log_k +7.5646
-delta_H 13.8072 kJ/mol # Calculated enthalpy of reaction NdF2+
# Enthalpy of formation: -323.5 kcal/mol
-analytic 2.7901e+002 8.7910e-002 -7.2424e+003 -1.1046e+002 -1.1309e+002
# -Range: 0-300
3.0000 F- + 1.0000 Nd+++ = NdF3
-llnl_gamma 3.0
log_k +9.8809
-delta_H -8.1588 kJ/mol # Calculated enthalpy of reaction NdF3
# Enthalpy of formation: -408.9 kcal/mol
-analytic 5.2220e+002 1.4154e-001 -1.3697e+004 -2.0551e+002 -2.1388e+002
# -Range: 0-300
4.0000 F- + 1.0000 Nd+++ = NdF4-
-llnl_gamma 4.0
log_k +11.8307
-delta_H -48.5344 kJ/mol # Calculated enthalpy of reaction NdF4-
# Enthalpy of formation: -498.7 kcal/mol
-analytic 6.1972e+002 1.4620e-001 -1.5869e+004 -2.4175e+002 -2.4780e+002
# -Range: 0-300
1.0000 Nd+++ + 1.0000 HPO4-- + 1.0000 H+ = NdH2PO4++
-llnl_gamma 4.5
log_k +9.5152
-delta_H -15.736 kJ/mol # Calculated enthalpy of reaction NdH2PO4+2
# Enthalpy of formation: -479.076 kcal/mol
-analytic 1.2450e+002 6.4953e-002 -4.0524e+002 -5.3728e+001 -6.3603e+000
# -Range: 0-300
1.0000 Nd+++ + 1.0000 HCO3- = NdHCO3++
-llnl_gamma 4.5
log_k +1.8457
-delta_H 9.19643 kJ/mol # Calculated enthalpy of reaction NdHCO3+2
# Enthalpy of formation: -329.2 kcal/mol
-analytic 5.5530e+001 3.3254e-002 -7.3859e+002 -2.4690e+001 -1.1542e+001
# -Range: 0-300
1.0000 Nd+++ + 1.0000 HPO4-- = NdHPO4+
-llnl_gamma 4.0
log_k +5.4000
-delta_H 0 # Not possible to calculate enthalpy of reaction NdHPO4+
# Enthalpy of formation: -0 kcal/mol
1.0000 Nd+++ + 1.0000 NO3- = NdNO3++
-llnl_gamma 4.5
log_k +0.7902
-delta_H -27.8529 kJ/mol # Calculated enthalpy of reaction NdNO3+2
# Enthalpy of formation: -222.586 kcal/mol
-analytic 3.3850e+001 2.7112e-002 1.4404e+003 -1.8570e+001 2.2466e+001
# -Range: 0-300
1.0000 Nd+++ + 1.0000 H2O = NdO+ +2.0000 H+
-llnl_gamma 4.0
log_k -17.0701
-delta_H 116.386 kJ/mol # Calculated enthalpy of reaction NdO+
# Enthalpy of formation: -207 kcal/mol
-analytic 1.8961e+002 3.0563e-002 -1.4153e+004 -6.8024e+001 -2.2089e+002
# -Range: 0-300
2.0000 H2O + 1.0000 Nd+++ = NdO2- +4.0000 H+
-llnl_gamma 4.0
log_k -37.0721
-delta_H 298.88 kJ/mol # Calculated enthalpy of reaction NdO2-
# Enthalpy of formation: -231.7 kcal/mol
-analytic 1.9606e+002 1.4784e-002 -2.1838e+004 -6.6399e+001 -3.4082e+002
# -Range: 0-300
2.0000 H2O + 1.0000 Nd+++ = NdO2H +3.0000 H+
-llnl_gamma 3.0
log_k -26.3702
-delta_H 230.681 kJ/mol # Calculated enthalpy of reaction NdO2H
# Enthalpy of formation: -248 kcal/mol
-analytic 3.4617e+002 4.5955e-002 -2.3960e+004 -1.2361e+002 -3.7398e+002
# -Range: 0-300
1.0000 Nd+++ + 1.0000 H2O = NdOH++ +1.0000 H+
-llnl_gamma 4.5
log_k -8.1274
-delta_H 80.8223 kJ/mol # Calculated enthalpy of reaction NdOH+2
# Enthalpy of formation: -215.5 kcal/mol
-analytic 6.6963e+001 1.2182e-002 -6.2797e+003 -2.3300e+001 -9.8008e+001
# -Range: 0-300
1.0000 Nd+++ + 1.0000 HPO4-- = NdPO4 +1.0000 H+
-llnl_gamma 3.0
log_k -0.5218
-delta_H 0 # Not possible to calculate enthalpy of reaction NdPO4
# Enthalpy of formation: -0 kcal/mol
1.0000 SO4-- + 1.0000 Nd+++ = NdSO4+
-llnl_gamma 4.0
log_k +3.6430
-delta_H 20.0832 kJ/mol # Calculated enthalpy of reaction NdSO4+
# Enthalpy of formation: -379.1 kcal/mol
-analytic 3.0267e+002 8.5362e-002 -8.9211e+003 -1.1902e+002 -1.3929e+002
# -Range: 0-300
2.0000 HAcetate + 1.0000 Ni++ = Ni(Acetate)2 +2.0000 H+
-llnl_gamma 3.0
log_k -7.1908
-delta_H -25.8571 kJ/mol # Calculated enthalpy of reaction Ni(Acetate)2
# Enthalpy of formation: -251.28 kcal/mol
-analytic -2.9660e+001 1.0643e-003 -1.0060e+003 7.9358e+000 5.2562e+005
# -Range: 0-300
3.0000 HAcetate + 1.0000 Ni++ = Ni(Acetate)3- +3.0000 H+
-llnl_gamma 4.0
log_k -11.3543
-delta_H -53.6807 kJ/mol # Calculated enthalpy of reaction Ni(Acetate)3-
# Enthalpy of formation: -374.03 kcal/mol
-analytic 5.0850e+001 -8.2435e-003 -1.3049e+004 -1.5410e+001 1.9704e+006
# -Range: 0-300
2.0000 NH3 + 1.0000 Ni++ = Ni(NH3)2++
-llnl_gamma 4.5
log_k +5.0598
-delta_H -29.7505 kJ/mol # Calculated enthalpy of reaction Ni(NH3)2+2
# Enthalpy of formation: -246.398 kJ/mol
-analytic 1.0002e+002 5.2896e-003 -2.5967e+003 -3.5485e+001 -4.0548e+001
# -Range: 0-300
6.0000 NH3 + 1.0000 Ni++ = Ni(NH3)6++
-llnl_gamma 4.5
log_k +8.7344
-delta_H -88.0436 kJ/mol # Calculated enthalpy of reaction Ni(NH3)6+2
# Enthalpy of formation: -630.039 kJ/mol
-analytic 1.9406e+002 -1.3467e-002 -5.2321e+003 -6.6168e+001 -8.1699e+001
# -Range: 0-300
2.0000 NO3- + 1.0000 Ni++ = Ni(NO3)2
-llnl_gamma 3.0
log_k +0.1899
-delta_H -1.54153 kJ/mol # Calculated enthalpy of reaction Ni(NO3)2
# Enthalpy of formation: -469.137 kJ/mol
-analytic -4.2544e+001 -1.0101e-002 1.3496e+003 1.6663e+001 2.2933e+001
# -Range: 0-200
2.0000 H2O + 1.0000 Ni++ = Ni(OH)2 +2.0000 H+
-llnl_gamma 3.0
log_k -19.9902
-delta_H 0 # Not possible to calculate enthalpy of reaction Ni(OH)2
# Enthalpy of formation: -0 kcal/mol
3.0000 H2O + 1.0000 Ni++ = Ni(OH)3- +3.0000 H+
-llnl_gamma 4.0
log_k -30.9852
-delta_H 0 # Not possible to calculate enthalpy of reaction Ni(OH)3-
# Enthalpy of formation: -0 kcal/mol
2.0000 Ni++ + 1.0000 H2O = Ni2OH+++ +1.0000 H+
-llnl_gamma 5.0
log_k -10.7
-delta_H 0 # Not possible to calculate enthalpy of reaction Ni2OH+3
# Enthalpy of formation: -0 kcal/mol
4.0000 Ni++ + 4.0000 H2O = Ni4(OH)4++++ +4.0000 H+
-llnl_gamma 5.5
log_k -27.6803
-delta_H 0 # Not possible to calculate enthalpy of reaction Ni4(OH)4+4
# Enthalpy of formation: -0 kcal/mol
1.0000 Ni++ + 1.0000 Br- = NiBr+
-llnl_gamma 4.0
log_k -0.37
-delta_H 0 # Not possible to calculate enthalpy of reaction NiBr+
# Enthalpy of formation: -0 kcal/mol
1.0000 Ni++ + 1.0000 HAcetate = NiAcetate+ +1.0000 H+
-llnl_gamma 4.0
log_k -3.3278
-delta_H -10.2508 kJ/mol # Calculated enthalpy of reaction NiAcetate+
# Enthalpy of formation: -131.45 kcal/mol
-analytic -3.3110e+000 1.6895e-003 -1.0556e+003 2.7168e-002 2.6350e+005
# -Range: 0-300
1.0000 Ni++ + 1.0000 Cl- = NiCl+
-llnl_gamma 4.0
log_k -0.9962
-delta_H 5.99567 kJ/mol # Calculated enthalpy of reaction NiCl+
# Enthalpy of formation: -51.4 kcal/mol
-analytic 9.5370e+001 3.8521e-002 -2.1746e+003 -4.0629e+001 -3.3961e+001
# -Range: 0-300
2.0000 HPO4-- + 1.0000 Ni++ + 1.0000 H+ = NiHP2O7- +1.0000 H2O
-llnl_gamma 4.0
log_k +9.2680
-delta_H 0 # Not possible to calculate enthalpy of reaction NiHP2O7-
# Enthalpy of formation: -0 kcal/mol
1.0000 Ni++ + 1.0000 NO3- = NiNO3+
-llnl_gamma 4.0
log_k +0.4000
-delta_H 0 # Not possible to calculate enthalpy of reaction NiNO3+
# Enthalpy of formation: -0 kcal/mol
2.0000 HPO4-- + 1.0000 Ni++ = NiP2O7-- +1.0000 H2O
-llnl_gamma 4.0
log_k +3.1012
-delta_H 9.68819 kJ/mol # Calculated enthalpy of reaction NiP2O7-2
# Enthalpy of formation: -2342.61 kJ/mol
-analytic 4.6809e+002 1.0985e-001 -1.4310e+004 -1.8173e+002 -2.2344e+002
# -Range: 0-300
1.0000 SO4-- + 1.0000 Ni++ = NiSO4
-llnl_gamma 3.0
log_k +2.1257
-delta_H 2.36814 kJ/mol # Calculated enthalpy of reaction NiSO4
# Enthalpy of formation: -229.734 kcal/mol
-analytic 6.1187e+001 2.4211e-002 -1.2180e+003 -2.5130e+001 -2.0705e+001
# -Range: 0-200
1.0000 SeO4-- + 1.0000 Ni++ = NiSeO4
-llnl_gamma 3.0
log_k +2.6700
-delta_H 0 # Not possible to calculate enthalpy of reaction NiSeO4
# Enthalpy of formation: -0 kcal/mol
5.0000 HCO3- + 1.0000 Np++++ = Np(CO3)5-6 +5.0000 H+
-llnl_gamma 4.0
log_k -13.344
-delta_H 92.7067 kJ/mol # Calculated enthalpy of reaction Np(CO3)5-6
# Enthalpy of formation: -935.22 kcal/mol
-analytic 6.3005e+002 2.3388e-001 -1.8328e+004 -2.6334e+002 -2.8618e+002
# -Range: 0-300
2.0000 HPO4-- + 2.0000 H+ + 1.0000 Np+++ = Np(H2PO4)2+
-llnl_gamma 4.0
log_k +3.7000
-delta_H -1.55258 kJ/mol # Calculated enthalpy of reaction Np(H2PO4)2+
# Enthalpy of formation: -743.981 kcal/mol
-analytic 7.8161e+002 2.8446e-001 -1.2330e+004 -3.3194e+002 -2.1056e+002
# -Range: 25-150
3.0000 HPO4-- + 3.0000 H+ + 1.0000 Np+++ = Np(H2PO4)3
-llnl_gamma 3.0
log_k +5.6000
-delta_H -21.8575 kJ/mol # Calculated enthalpy of reaction Np(H2PO4)3
# Enthalpy of formation: -1057.65 kcal/mol
-analytic 1.5150e+003 4.4939e-001 -3.2766e+004 -6.1975e+002 -5.5934e+002
# -Range: 25-150
2.0000 HPO4-- + 1.0000 Np++++ = Np(HPO4)2
-llnl_gamma 3.0
log_k +23.7000
-delta_H -35.24 kJ/mol # Calculated enthalpy of reaction Np(HPO4)2
# Enthalpy of formation: -758.94 kcal/mol
-analytic 4.7722e+002 2.1099e-001 -4.7296e+003 -2.0229e+002 -8.0831e+001
# -Range: 25-150
3.0000 HPO4-- + 1.0000 Np++++ = Np(HPO4)3--
-llnl_gamma 4.0
log_k +33.4000
-delta_H -44.9093 kJ/mol # Calculated enthalpy of reaction Np(HPO4)3-2
# Enthalpy of formation: -1070.07 kcal/mol
-analytic -1.5951e+003 -3.6579e-001 5.1343e+004 6.3262e+002 8.7619e+002
# -Range: 25-150
4.0000 HPO4-- + 1.0000 Np++++ = Np(HPO4)4----
-llnl_gamma 4.0
log_k +43.2000
-delta_H -67.0803 kJ/mol # Calculated enthalpy of reaction Np(HPO4)4-4
# Enthalpy of formation: -1384.18 kcal/mol
-analytic 5.8359e+003 1.5194e+000 -1.6349e+005 -2.3025e+003 -2.7903e+003
# -Range: 25-150
5.0000 HPO4-- + 1.0000 Np++++ = Np(HPO4)5-6
-llnl_gamma 4.0
log_k +52.0000
-delta_H -83.5401 kJ/mol # Calculated enthalpy of reaction Np(HPO4)5-6
# Enthalpy of formation: -1696.93 kcal/mol
-analytic -1.8082e+003 -2.0018e-001 7.5155e+004 6.7400e+002 1.2824e+003
# -Range: 25-150
2.0000 H2O + 1.0000 Np++++ = Np(OH)2++ +2.0000 H+
-llnl_gamma 4.5
log_k -2.8
-delta_H 77.0669 kJ/mol # Calculated enthalpy of reaction Np(OH)2+2
# Enthalpy of formation: -251.102 kcal/mol
-analytic 2.9299e+003 6.5812e-001 -9.5085e+004 -1.1356e+003 -1.6227e+003
# -Range: 25-150
3.0000 H2O + 1.0000 Np++++ = Np(OH)3+ +3.0000 H+
-llnl_gamma 4.0
log_k -5.8
-delta_H 99.5392 kJ/mol # Calculated enthalpy of reaction Np(OH)3+
# Enthalpy of formation: -314.048 kcal/mol
-analytic -4.7723e+003 -1.1810e+000 1.3545e+005 1.8850e+003 2.3117e+003
# -Range: 25-150
4.0000 H2O + 1.0000 Np++++ = Np(OH)4 +4.0000 H+
-llnl_gamma 3.0
log_k -9.6
-delta_H 109.585 kJ/mol # Calculated enthalpy of reaction Np(OH)4
# Enthalpy of formation: -379.964 kcal/mol
-analytic -5.5904e+003 -1.3639e+000 1.6112e+005 2.2013e+003 2.7498e+003
# -Range: 25-150
2.0000 SO4-- + 1.0000 Np++++ = Np(SO4)2
-llnl_gamma 3.0
log_k +9.9000
-delta_H 40.005 kJ/mol # Calculated enthalpy of reaction Np(SO4)2
# Enthalpy of formation: -558.126 kcal/mol
-analytic -9.0765e+002 -1.8494e-001 2.7951e+004 3.5521e+002 4.7702e+002
# -Range: 25-150
1.0000 Np++++ + 1.0000 Cl- = NpCl+++
-llnl_gamma 5.0
log_k +0.2000
-delta_H 20.3737 kJ/mol # Calculated enthalpy of reaction NpCl+3
# Enthalpy of formation: -167.951 kcal/mol
-analytic 8.3169e+002 2.6267e-001 -2.1618e+004 -3.3838e+002 -3.6898e+002
# -Range: 25-150
2.0000 Cl- + 1.0000 Np++++ = NpCl2++
-llnl_gamma 4.5
log_k -0.1
-delta_H 94.5853 kJ/mol # Calculated enthalpy of reaction NpCl2+2
# Enthalpy of formation: -190.147 kcal/mol
-analytic -1.5751e+003 -3.8759e-001 4.2054e+004 6.2619e+002 7.1777e+002
# -Range: 25-150
1.0000 Np++++ + 1.0000 F- = NpF+++
-llnl_gamma 5.0
log_k +8.7000
-delta_H -3.43746 kJ/mol # Calculated enthalpy of reaction NpF+3
# Enthalpy of formation: -213.859 kcal/mol
-analytic 2.7613e+000 1.3498e-003 -1.6411e+003 2.9074e+000 3.4192e+005
# -Range: 25-150
2.0000 F- + 1.0000 Np++++ = NpF2++
-llnl_gamma 4.5
log_k +15.4000
-delta_H 6.03094 kJ/mol # Calculated enthalpy of reaction NpF2+2
# Enthalpy of formation: -291.746 kcal/mol
-analytic -2.6793e+002 -4.2056e-002 9.7952e+003 1.0629e+002 1.6715e+002
# -Range: 25-150
1.0000 Np+++ + 1.0000 HPO4-- + 1.0000 H+ = NpH2PO4++
-llnl_gamma 4.5
log_k +2.4000
-delta_H 6.0874 kJ/mol # Calculated enthalpy of reaction NpH2PO4+2
# Enthalpy of formation: -433.34 kcal/mol
-analytic 6.0731e+003 1.4733e+000 -1.7919e+005 -2.3880e+003 -3.0582e+003
# -Range: 25-150
1.0000 Np++++ + 1.0000 HPO4-- = NpHPO4++
-llnl_gamma 4.5
log_k +12.9000
-delta_H 7.54554 kJ/mol # Calculated enthalpy of reaction NpHPO4+2
# Enthalpy of formation: -439.899 kcal/mol
-analytic -7.2792e+003 -1.7476e+000 2.1770e+005 2.8624e+003 3.7154e+003
# -Range: 25-150
2.0000 HCO3- + 1.0000 NpO2++ = NpO2(CO3)2-- +2.0000 H+
-llnl_gamma 4.0
log_k -6.6576
-delta_H 57.2588 kJ/mol # Calculated enthalpy of reaction NpO2(CO3)2-2
# Enthalpy of formation: -521.77 kcal/mol
-analytic 2.6597e+002 7.5850e-002 -9.9987e+003 -1.0576e+002 -1.5610e+002
# -Range: 0-300
2.0000 HCO3- + 1.0000 NpO2+ = NpO2(CO3)2--- +2.0000 H+
-llnl_gamma 4.0
log_k -13.6576
-delta_H 58.1553 kJ/mol # Calculated enthalpy of reaction NpO2(CO3)2-3
# Enthalpy of formation: -549.642 kcal/mol
-analytic 2.6012e+002 7.3174e-002 -1.0250e+004 -1.0556e+002 -1.6002e+002
# -Range: 0-300
3.0000 HCO3- + 1.0000 NpO2+ = NpO2(CO3)3-5 +3.0000 H+
-llnl_gamma 4.0
log_k -22.4864
-delta_H 70.176 kJ/mol # Calculated enthalpy of reaction NpO2(CO3)3-5
# Enthalpy of formation: -711.667 kcal/mol
-analytic 3.7433e+002 1.2938e-001 -1.2791e+004 -1.5861e+002 -1.9970e+002
# -Range: 0-300
3.0000 HCO3- + 1.0000 NpO2++ = NpO2(CO3)3---- +3.0000 H+
-llnl_gamma 4.0
log_k -10.5864
-delta_H 3.14711 kJ/mol # Calculated enthalpy of reaction NpO2(CO3)3-4
# Enthalpy of formation: -699.601 kcal/mol
-analytic 3.7956e+002 1.1163e-001 -1.0607e+004 -1.5674e+002 -1.6562e+002
# -Range: 0-300
1.0000 NpO2+ + 1.0000 HCO3- = NpO2CO3- +1.0000 H+
-llnl_gamma 4.0
log_k -5.7288
-delta_H 69.1634 kJ/mol # Calculated enthalpy of reaction NpO2CO3-
# Enthalpy of formation: -382.113 kcal/mol
-analytic 1.4634e+002 2.6576e-002 -8.2036e+003 -5.3534e+001 -1.2805e+002
# -Range: 0-300
1.0000 NpO2+ + 1.0000 Cl- = NpO2Cl
-llnl_gamma 3.0
log_k -0.4
-delta_H 15.4492 kJ/mol # Calculated enthalpy of reaction NpO2Cl
# Enthalpy of formation: -269.986 kcal/mol
-analytic 4.5109e+002 9.0437e-002 -1.5453e+004 -1.7241e+002 -2.6371e+002
# -Range: 25-150
1.0000 NpO2++ + 1.0000 Cl- = NpO2Cl+
-llnl_gamma 4.0
log_k -0.2
-delta_H 11.6239 kJ/mol # Calculated enthalpy of reaction NpO2Cl+
# Enthalpy of formation: -242.814 kcal/mol
-analytic -1.2276e+003 -2.5435e-001 3.8507e+004 4.7447e+002 6.5715e+002
# -Range: 25-150
1.0000 NpO2+ + 1.0000 F- = NpO2F
-llnl_gamma 3.0
log_k +1.0000
-delta_H 34.2521 kJ/mol # Calculated enthalpy of reaction NpO2F
# Enthalpy of formation: -305.709 kcal/mol
-analytic -1.9364e+002 -4.4083e-002 4.5602e+003 7.7791e+001 7.7840e+001
# -Range: 25-150
1.0000 NpO2++ + 1.0000 F- = NpO2F+
-llnl_gamma 4.0
log_k +4.6000
-delta_H 0.883568 kJ/mol # Calculated enthalpy of reaction NpO2F+
# Enthalpy of formation: -285.598 kcal/mol
-analytic 9.6320e+002 2.4799e-001 -2.7614e+004 -3.7985e+002 -4.7128e+002
# -Range: 25-150
2.0000 F- + 1.0000 NpO2++ = NpO2F2
-llnl_gamma 3.0
log_k +7.8000
-delta_H 2.60319 kJ/mol # Calculated enthalpy of reaction NpO2F2
# Enthalpy of formation: -365.337 kcal/mol
-analytic 1.9648e+002 6.4083e-002 -4.5601e+003 -7.7790e+001 -7.7840e+001
# -Range: 25-150
1.0000 NpO2+ + 1.0000 HPO4-- + 1.0000 H+ = NpO2H2PO4
-llnl_gamma 3.0
log_k +0.6000
-delta_H 18.717 kJ/mol # Calculated enthalpy of reaction NpO2H2PO4
# Enthalpy of formation: -538.087 kcal/mol
-analytic 1.0890e+003 2.7738e-001 -3.0654e+004 -4.3171e+002 -5.2317e+002
# -Range: 25-150
1.0000 NpO2++ + 1.0000 HPO4-- + 1.0000 H+ = NpO2H2PO4+
-llnl_gamma 4.0
log_k +2.3000
-delta_H 9.31014 kJ/mol # Calculated enthalpy of reaction NpO2H2PO4+
# Enthalpy of formation: -512.249 kcal/mol
-analytic -5.6996e+003 -1.4008e+000 1.6898e+005 2.2441e+003 2.8838e+003
# -Range: 25-150
1.0000 NpO2++ + 1.0000 HPO4-- = NpO2HPO4
-llnl_gamma 3.0
log_k +8.2000
-delta_H -6.47609 kJ/mol # Calculated enthalpy of reaction NpO2HPO4
# Enthalpy of formation: -516.022 kcal/mol
-analytic 4.8515e+003 1.2189e+000 -1.4069e+005 -1.9135e+003 -2.4011e+003
# -Range: 25-150
1.0000 NpO2+ + 1.0000 HPO4-- = NpO2HPO4-
-llnl_gamma 4.0
log_k +3.5000
-delta_H 49.8668 kJ/mol # Calculated enthalpy of reaction NpO2HPO4-
# Enthalpy of formation: -530.642 kcal/mol
-analytic -4.1705e+003 -9.9302e-001 1.2287e+005 1.6399e+003 2.0969e+003
# -Range: 25-150
1.0000 NpO2+ + 1.0000 H2O = NpO2OH +1.0000 H+
-llnl_gamma 3.0
log_k -8.9
-delta_H 43.6285 kJ/mol # Calculated enthalpy of reaction NpO2OH
# Enthalpy of formation: -291.635 kcal/mol
-analytic -4.5710e+002 -1.2286e-001 1.0640e+004 1.8151e+002 1.8163e+002
# -Range: 25-150
1.0000 NpO2++ + 1.0000 H2O = NpO2OH+ +1.0000 H+
-llnl_gamma 4.0
log_k -5.2
-delta_H 43.3805 kJ/mol # Calculated enthalpy of reaction NpO2OH+
# Enthalpy of formation: -263.608 kcal/mol
-analytic 1.7485e+002 4.0017e-002 -7.5154e+003 -6.7399e+001 -1.2823e+002
# -Range: 25-150
1.0000 SO4-- + 1.0000 NpO2++ = NpO2SO4
-llnl_gamma 3.0
log_k +3.3000
-delta_H 19.8789 kJ/mol # Calculated enthalpy of reaction NpO2SO4
# Enthalpy of formation: -418.308 kcal/mol
-analytic -1.5624e+002 7.3296e-003 6.7555e+003 5.4435e+001 1.1527e+002
# -Range: 25-150
1.0000 SO4-- + 1.0000 NpO2+ = NpO2SO4-
-llnl_gamma 4.0
log_k +0.4000
-delta_H 19.1395 kJ/mol # Calculated enthalpy of reaction NpO2SO4-
# Enthalpy of formation: -446.571 kcal/mol
-analytic -3.1804e+002 -9.3472e-002 7.6002e+003 1.2965e+002 1.2973e+002
# -Range: 25-150
1.0000 Np+++ + 1.0000 H2O = NpOH++ +1.0000 H+
-llnl_gamma 4.5
log_k -7
-delta_H 50.1031 kJ/mol # Calculated enthalpy of reaction NpOH+2
# Enthalpy of formation: -182.322 kcal/mol
-analytic 1.4062e+002 3.2671e-002 -6.7555e+003 -5.4435e+001 -1.1526e+002
# -Range: 25-150
1.0000 Np++++ + 1.0000 H2O = NpOH+++ +1.0000 H+
-llnl_gamma 5.0
log_k -1
-delta_H 51.0089 kJ/mol # Calculated enthalpy of reaction NpOH+3
# Enthalpy of formation: -189.013 kcal/mol
-analytic -1.8373e+002 -5.2443e-002 2.7025e+003 7.6503e+001 4.6154e+001
# -Range: 25-150
1.0000 SO4-- + 1.0000 Np++++ = NpSO4++
-llnl_gamma 4.5
log_k +5.5000
-delta_H 20.7377 kJ/mol # Calculated enthalpy of reaction NpSO4+2
# Enthalpy of formation: -345.331 kcal/mol
-analytic 3.9477e+002 1.1981e-001 -1.0978e+004 -1.5687e+002 -1.8736e+002
# -Range: 25-150
1.0000 H2O = OH- +1.0000 H+
-llnl_gamma 3.5
log_k -13.9951
-delta_H 55.8146 kJ/mol # Calculated enthalpy of reaction OH-
# Enthalpy of formation: -54.977 kcal/mol
-analytic -6.7506e+001 -3.0619e-002 -1.9901e+003 2.8004e+001 -3.1033e+001
# -Range: 0-300
2.0000 HPO4-- = P2O7---- +1.0000 H2O
-llnl_gamma 4.0
log_k -3.7463
-delta_H 27.2256 kJ/mol # Calculated enthalpy of reaction P2O7-4
# Enthalpy of formation: -2271.1 kJ/mol
-analytic 4.0885e+002 1.3243e-001 -1.1373e+004 -1.6727e+002 -1.7758e+002
# -Range: 0-300
3.0000 H+ + 1.0000 HPO4-- = PH4+ +2.0000 O2
-llnl_gamma 4.0
log_k -212.7409
-delta_H 0 # Not possible to calculate enthalpy of reaction PH4+
# Enthalpy of formation: -0 kcal/mol
1.0000 HPO4-- + 1.0000 H+ + 1.0000 F- = PO3F-- +1.0000 H2O
-llnl_gamma 4.0
log_k +7.1993
-delta_H 0 # Not possible to calculate enthalpy of reaction PO3F-2
# Enthalpy of formation: -0 kcal/mol
1.0000 HPO4-- = PO4--- +1.0000 H+
-llnl_gamma 4.0
log_k -12.3218
-delta_H 14.7068 kJ/mol # Calculated enthalpy of reaction PO4-3
# Enthalpy of formation: -305.3 kcal/mol
-analytic -7.6170e+001 -3.3574e-002 1.3405e+002 2.9658e+001 2.1140e+000
# -Range: 0-300
2.0000 BrO3- + 1.0000 Pb++ = Pb(BrO3)2
-llnl_gamma 3.0
log_k +5.1939
-delta_H 0 # Not possible to calculate enthalpy of reaction Pb(BrO3)2
# Enthalpy of formation: -0 kcal/mol
2.0000 HAcetate + 1.0000 Pb++ = Pb(Acetate)2 +2.0000 H+
-llnl_gamma 3.0
log_k -6.1133
-delta_H 10.5437 kJ/mol # Calculated enthalpy of reaction Pb(Acetate)2
# Enthalpy of formation: -229.46 kcal/mol
-analytic -1.7315e+001 -1.0618e-003 -3.6365e+003 6.9263e+000 5.8659e+005
# -Range: 0-300
3.0000 HAcetate + 1.0000 Pb++ = Pb(Acetate)3- +3.0000 H+
-llnl_gamma 4.0
log_k -8.972
-delta_H -2.84512 kJ/mol # Calculated enthalpy of reaction Pb(Acetate)3-
# Enthalpy of formation: -348.76 kcal/mol
-analytic 1.2417e+001 -3.1481e-003 -9.4152e+003 -1.6846e+000 1.3623e+006
# -Range: 0-300
2.0000 HCO3- + 1.0000 Pb++ = Pb(CO3)2-- +2.0000 H+
-llnl_gamma 4.0
log_k -11.2576
-delta_H 0 # Not possible to calculate enthalpy of reaction Pb(CO3)2-2
# Enthalpy of formation: -0 kcal/mol
2.0000 ClO3- + 1.0000 Pb++ = Pb(ClO3)2
-llnl_gamma 3.0
log_k -0.5133
-delta_H 0 # Not possible to calculate enthalpy of reaction Pb(ClO3)2
# Enthalpy of formation: -0 kcal/mol
2.0000 H2O + 1.0000 Pb++ = Pb(OH)2 +2.0000 H+
-llnl_gamma 3.0
log_k -17.0902
-delta_H 0 # Not possible to calculate enthalpy of reaction Pb(OH)2
# Enthalpy of formation: -0 kcal/mol
3.0000 H2O + 1.0000 Pb++ = Pb(OH)3- +3.0000 H+
-llnl_gamma 4.0
log_k -28.0852
-delta_H 0 # Not possible to calculate enthalpy of reaction Pb(OH)3-
# Enthalpy of formation: -0 kcal/mol
2.0000 Thiocyanate- + 1.0000 Pb++ = Pb(Thiocyanate)2
-llnl_gamma 3.0
log_k +1.2455
-delta_H 0 # Not possible to calculate enthalpy of reaction Pb(Thiocyanate)2
# Enthalpy of formation: -0 kcal/mol
2.0000 Pb++ + 1.0000 H2O = Pb2OH+++ +1.0000 H+
-llnl_gamma 5.0
log_k -6.3951
-delta_H 0 # Not possible to calculate enthalpy of reaction Pb2OH+3
# Enthalpy of formation: -0 kcal/mol
4.0000 H2O + 3.0000 Pb++ = Pb3(OH)4++ +4.0000 H+
-llnl_gamma 4.5
log_k -23.8803
-delta_H 0 # Not possible to calculate enthalpy of reaction Pb3(OH)4+2
# Enthalpy of formation: -0 kcal/mol
4.0000 Pb++ + 4.0000 H2O = Pb4(OH)4++++ +4.0000 H+
-llnl_gamma 5.5
log_k -20.8803
-delta_H 0 # Not possible to calculate enthalpy of reaction Pb4(OH)4+4
# Enthalpy of formation: -0 kcal/mol
8.0000 H2O + 6.0000 Pb++ = Pb6(OH)8++++ +8.0000 H+
-llnl_gamma 5.5
log_k -43.5606
-delta_H 0 # Not possible to calculate enthalpy of reaction Pb6(OH)8+4
# Enthalpy of formation: -0 kcal/mol
1.0000 Pb++ + 1.0000 Br- = PbBr+
-llnl_gamma 4.0
log_k +1.1831
-delta_H 0 # Not possible to calculate enthalpy of reaction PbBr+
# Enthalpy of formation: -0 kcal/mol
2.0000 Br- + 1.0000 Pb++ = PbBr2
-llnl_gamma 3.0
log_k +1.5062
-delta_H 0 # Not possible to calculate enthalpy of reaction PbBr2
# Enthalpy of formation: -0 kcal/mol
3.0000 Br- + 1.0000 Pb++ = PbBr3-
-llnl_gamma 4.0
log_k +1.2336
-delta_H 0 # Not possible to calculate enthalpy of reaction PbBr3-
# Enthalpy of formation: -0 kcal/mol
1.0000 Pb++ + 1.0000 BrO3- = PbBrO3+
-llnl_gamma 4.0
log_k +1.9373
-delta_H 0 # Not possible to calculate enthalpy of reaction PbBrO3+
# Enthalpy of formation: -0 kcal/mol
1.0000 Pb++ + 1.0000 HAcetate = PbAcetate+ +1.0000 H+
-llnl_gamma 4.0
log_k -2.3603
-delta_H -2.33147e-15 kJ/mol # Calculated enthalpy of reaction PbAcetate+
# Enthalpy of formation: -115.88 kcal/mol
-analytic -2.6822e+001 1.0992e-003 7.3688e+002 8.4407e+000 7.0266e+004
# -Range: 0-300
1.0000 Pb++ + 1.0000 HCO3- = PbCO3 +1.0000 H+
-llnl_gamma 3.0
log_k -3.7488
-delta_H 0 # Not possible to calculate enthalpy of reaction PbCO3
# Enthalpy of formation: -0 kcal/mol
1.0000 Pb++ + 1.0000 Cl- = PbCl+
-llnl_gamma 4.0
log_k +1.4374
-delta_H 4.53127 kJ/mol # Calculated enthalpy of reaction PbCl+
# Enthalpy of formation: -38.63 kcal/mol
-analytic 1.1948e+002 4.3527e-002 -2.7666e+003 -4.9190e+001 -4.3206e+001
# -Range: 0-300
2.0000 Cl- + 1.0000 Pb++ = PbCl2
-llnl_gamma 3.0
log_k +2.0026
-delta_H 8.14206 kJ/mol # Calculated enthalpy of reaction PbCl2
# Enthalpy of formation: -77.7 kcal/mol
-analytic 2.2537e+002 7.7574e-002 -5.5112e+003 -9.2131e+001 -8.6064e+001
# -Range: 0-300
3.0000 Cl- + 1.0000 Pb++ = PbCl3-
-llnl_gamma 4.0
log_k +1.6881
-delta_H 7.86174 kJ/mol # Calculated enthalpy of reaction PbCl3-
# Enthalpy of formation: -117.7 kcal/mol
-analytic 2.5254e+002 8.9159e-002 -6.0116e+003 -1.0395e+002 -9.3880e+001
# -Range: 0-300
4.0000 Cl- + 1.0000 Pb++ = PbCl4--
-llnl_gamma 4.0
log_k +1.4909
-delta_H -7.18811 kJ/mol # Calculated enthalpy of reaction PbCl4-2
# Enthalpy of formation: -161.23 kcal/mol
-analytic 1.4048e+002 7.6332e-002 -1.1507e+003 -6.3786e+001 -1.7997e+001
# -Range: 0-300
1.0000 Pb++ + 1.0000 ClO3- = PbClO3+
-llnl_gamma 4.0
log_k -0.2208
-delta_H 0 # Not possible to calculate enthalpy of reaction PbClO3+
# Enthalpy of formation: -0 kcal/mol
1.0000 Pb++ + 1.0000 F- = PbF+
-llnl_gamma 4.0
log_k +0.8284
-delta_H 0 # Not possible to calculate enthalpy of reaction PbF+
# Enthalpy of formation: -0 kcal/mol
2.0000 F- + 1.0000 Pb++ = PbF2
-llnl_gamma 3.0
log_k +1.6132
-delta_H 0 # Not possible to calculate enthalpy of reaction PbF2
# Enthalpy of formation: -0 kcal/mol
1.0000 Pb++ + 1.0000 HPO4-- + 1.0000 H+ = PbH2PO4+
-llnl_gamma 4.0
log_k +1.5000
-delta_H 0 # Not possible to calculate enthalpy of reaction PbH2PO4+
# Enthalpy of formation: -0 kcal/mol
1.0000 Pb++ + 1.0000 HPO4-- = PbHPO4
-llnl_gamma 3.0
log_k +3.1000
-delta_H 0 # Not possible to calculate enthalpy of reaction PbHPO4
# Enthalpy of formation: -0 kcal/mol
1.0000 Pb++ + 1.0000 I- = PbI+
-llnl_gamma 4.0
log_k +1.9597
-delta_H 0 # Not possible to calculate enthalpy of reaction PbI+
# Enthalpy of formation: -0 kcal/mol
2.0000 I- + 1.0000 Pb++ = PbI2
-llnl_gamma 3.0
log_k +2.7615
-delta_H 0 # Not possible to calculate enthalpy of reaction PbI2
# Enthalpy of formation: -0 kcal/mol
3.0000 I- + 1.0000 Pb++ = PbI3-
-llnl_gamma 4.0
log_k +3.3355
-delta_H 0 # Not possible to calculate enthalpy of reaction PbI3-
# Enthalpy of formation: -0 kcal/mol
4.0000 I- + 1.0000 Pb++ = PbI4--
-llnl_gamma 4.0
log_k +4.0672
-delta_H 0 # Not possible to calculate enthalpy of reaction PbI4-2
# Enthalpy of formation: -0 kcal/mol
1.0000 Pb++ + 1.0000 NO3- = PbNO3+
-llnl_gamma 4.0
log_k +1.2271
-delta_H 0 # Not possible to calculate enthalpy of reaction PbNO3+
# Enthalpy of formation: -0 kcal/mol
1.0000 Pb++ + 1.0000 H2O = PbOH+ +1.0000 H+
-llnl_gamma 4.0
log_k -7.6951
-delta_H 0 # Not possible to calculate enthalpy of reaction PbOH+
# Enthalpy of formation: -0 kcal/mol
2.0000 HPO4-- + 1.0000 Pb++ = PbP2O7-- +1.0000 H2O
-llnl_gamma 4.0
log_k +7.4136
-delta_H 0 # Not possible to calculate enthalpy of reaction PbP2O7-2
# Enthalpy of formation: -0 kcal/mol
1.0000 Thiocyanate- + 1.0000 Pb++ = PbThiocyanate+
-llnl_gamma 4.0
log_k +0.9827
-delta_H 0 # Not possible to calculate enthalpy of reaction PbThiocyanate+
# Enthalpy of formation: -0 kcal/mol
1.0000 Pd++ + 1.0000 Cl- = PdCl+
-llnl_gamma 4.0
log_k +6.0993
-delta_H -31.995 kJ/mol # Calculated enthalpy of reaction PdCl+
# Enthalpy of formation: -5.5 kcal/mol
-analytic 7.2852e+001 3.6886e-002 7.3102e+002 -3.2402e+001 1.1385e+001
# -Range: 0-300
2.0000 Cl- + 1.0000 Pd++ = PdCl2
-llnl_gamma 3.0
log_k +10.7327
-delta_H -66.1658 kJ/mol # Calculated enthalpy of reaction PdCl2
# Enthalpy of formation: -53.6 kcal/mol
-analytic 1.6849e+002 7.9321e-002 8.2874e+002 -7.4416e+001 1.2882e+001
# -Range: 0-300
3.0000 Cl- + 1.0000 Pd++ = PdCl3-
-llnl_gamma 4.0
log_k +13.0937
-delta_H -101.592 kJ/mol # Calculated enthalpy of reaction PdCl3-
# Enthalpy of formation: -102 kcal/mol
-analytic 4.5978e+001 6.2999e-002 6.9333e+003 -3.0257e+001 1.0817e+002
# -Range: 0-300
4.0000 Cl- + 1.0000 Pd++ = PdCl4--
-llnl_gamma 4.0
log_k +15.1615
-delta_H -152.08 kJ/mol # Calculated enthalpy of reaction PdCl4-2
# Enthalpy of formation: -154 kcal/mol
-analytic -3.2209e+001 5.3432e-002 1.2180e+004 -3.7814e+000 1.9006e+002
# -Range: 0-300
1.0000 Pd++ + 1.0000 H2O = PdO +2.0000 H+
-llnl_gamma 3.0
log_k -2.19
-delta_H 6.43081 kJ/mol # Calculated enthalpy of reaction PdO
# Enthalpy of formation: -24.7 kcal/mol
-analytic 1.3587e+002 2.9292e-002 -4.6645e+003 -5.2997e+001 -7.2825e+001
# -Range: 0-300
1.0000 Pd++ + 1.0000 H2O = PdOH+ +1.0000 H+
-llnl_gamma 4.0
log_k -1.0905
-delta_H -3.19239 kJ/mol # Calculated enthalpy of reaction PdOH+
# Enthalpy of formation: -27 kcal/mol
-analytic 1.4291e+001 5.8382e-003 -1.9881e+002 -6.6475e+000 -3.1065e+000
# -Range: 0-300
2.0000 HCO3- + 1.0000 Pm+++ = Pm(CO3)2- +2.0000 H+
-llnl_gamma 4.0
log_k -7.9576
-delta_H 0 # Not possible to calculate enthalpy of reaction Pm(CO3)2-
# Enthalpy of formation: -0 kcal/mol
2.0000 HPO4-- + 1.0000 Pm+++ = Pm(HPO4)2-
-llnl_gamma 4.0
log_k +9.2000
-delta_H 0 # Not possible to calculate enthalpy of reaction Pm(HPO4)2-
# Enthalpy of formation: -0 kcal/mol
2.0000 H2O + 1.0000 Pm+++ = Pm(OH)2+ +2.0000 H+
-llnl_gamma 4.0
log_k -16.7902
-delta_H 0 # Not possible to calculate enthalpy of reaction Pm(OH)2+
# Enthalpy of formation: -0 kcal/mol
3.0000 H2O + 1.0000 Pm+++ = Pm(OH)3 +3.0000 H+
-llnl_gamma 3.0
log_k -26.1852
-delta_H 0 # Not possible to calculate enthalpy of reaction Pm(OH)3
# Enthalpy of formation: -0 kcal/mol
2.0000 HPO4-- + 1.0000 Pm+++ = Pm(PO4)2--- +2.0000 H+
-llnl_gamma 4.0
log_k -4.6837
-delta_H 0 # Not possible to calculate enthalpy of reaction Pm(PO4)2-3
# Enthalpy of formation: -0 kcal/mol
2.0000 SO4-- + 1.0000 Pm+++ = Pm(SO4)2-
-llnl_gamma 4.0
log_k +5.2000
-delta_H 0 # Not possible to calculate enthalpy of reaction Pm(SO4)2-
# Enthalpy of formation: -0 kcal/mol
1.0000 Pm+++ + 1.0000 HCO3- = PmCO3+ +1.0000 H+
-llnl_gamma 4.0
log_k -2.6288
-delta_H 0 # Not possible to calculate enthalpy of reaction PmCO3+
# Enthalpy of formation: -0 kcal/mol
1.0000 Pm+++ + 1.0000 Cl- = PmCl++
-llnl_gamma 4.5
log_k +0.3400
-delta_H 0 # Not possible to calculate enthalpy of reaction PmCl+2
# Enthalpy of formation: -0 kcal/mol
1.0000 Pm+++ + 1.0000 F- = PmF++
-llnl_gamma 4.5
log_k +3.8000
-delta_H 0 # Not possible to calculate enthalpy of reaction PmF+2
# Enthalpy of formation: -0 kcal/mol
1.0000 Pm+++ + 1.0000 HPO4-- + 1.0000 H+ = PmH2PO4++
-llnl_gamma 4.5
log_k +9.6054
-delta_H 0 # Not possible to calculate enthalpy of reaction PmH2PO4+2
# Enthalpy of formation: -0 kcal/mol
1.0000 Pm+++ + 1.0000 HCO3- = PmHCO3++
-llnl_gamma 4.5
log_k +2.1000
-delta_H 0 # Not possible to calculate enthalpy of reaction PmHCO3+2
# Enthalpy of formation: -0 kcal/mol
1.0000 Pm+++ + 1.0000 HPO4-- = PmHPO4+
-llnl_gamma 4.0
log_k +5.5000
-delta_H 0 # Not possible to calculate enthalpy of reaction PmHPO4+
# Enthalpy of formation: -0 kcal/mol
1.0000 Pm+++ + 1.0000 NO3- = PmNO3++
-llnl_gamma 4.5
log_k +1.1000
-delta_H 0 # Not possible to calculate enthalpy of reaction PmNO3+2
# Enthalpy of formation: -0 kcal/mol
1.0000 Pm+++ + 1.0000 H2O = PmOH++ +1.0000 H+
-llnl_gamma 4.5
log_k -7.9951
-delta_H 0 # Not possible to calculate enthalpy of reaction PmOH+2
# Enthalpy of formation: -0 kcal/mol
1.0000 Pm+++ + 1.0000 HPO4-- = PmPO4 +1.0000 H+
-llnl_gamma 3.0
log_k -0.3718
-delta_H 0 # Not possible to calculate enthalpy of reaction PmPO4
# Enthalpy of formation: -0 kcal/mol
1.0000 SO4-- + 1.0000 Pm+++ = PmSO4+
-llnl_gamma 4.0
log_k +3.5000
-delta_H 0 # Not possible to calculate enthalpy of reaction PmSO4+
# Enthalpy of formation: -0 kcal/mol
2.0000 HAcetate + 1.0000 Pr+++ = Pr(Acetate)2+ +2.0000 H+
-llnl_gamma 4.0
log_k -4.8525
-delta_H -23.8906 kJ/mol # Calculated enthalpy of reaction Pr(Acetate)2+
# Enthalpy of formation: -406.71 kcal/mol
-analytic -1.6464e+001 6.2989e-004 -4.4771e+002 3.6947e+000 3.3816e+005
# -Range: 0-300
3.0000 HAcetate + 1.0000 Pr+++ = Pr(Acetate)3 +3.0000 H+
-llnl_gamma 3.0
log_k -8.2023
-delta_H -40.3756 kJ/mol # Calculated enthalpy of reaction Pr(Acetate)3
# Enthalpy of formation: -526.75 kcal/mol
-analytic -1.2007e+001 4.9332e-004 0.0000e+000 0.0000e+000 3.2789e+005
# -Range: 0-300
2.0000 HCO3- + 1.0000 Pr+++ = Pr(CO3)2- +2.0000 H+
-llnl_gamma 4.0
log_k -8.1076
-delta_H 0 # Not possible to calculate enthalpy of reaction Pr(CO3)2-
# Enthalpy of formation: -0 kcal/mol
2.0000 HPO4-- + 1.0000 Pr+++ = Pr(HPO4)2-
-llnl_gamma 4.0
log_k +8.9000
-delta_H 0 # Not possible to calculate enthalpy of reaction Pr(HPO4)2-
# Enthalpy of formation: -0 kcal/mol
2.0000 HPO4-- + 1.0000 Pr+++ = Pr(PO4)2--- +2.0000 H+
-llnl_gamma 4.0
log_k -5.5637
-delta_H 0 # Not possible to calculate enthalpy of reaction Pr(PO4)2-3
# Enthalpy of formation: -0 kcal/mol
2.0000 SO4-- + 1.0000 Pr+++ = Pr(SO4)2-
-llnl_gamma 4.0
log_k +4.9000
-delta_H 0 # Not possible to calculate enthalpy of reaction Pr(SO4)2-
# Enthalpy of formation: -0 kcal/mol
1.0000 Pr+++ + 1.0000 HAcetate = PrAcetate++ +1.0000 H+
-llnl_gamma 4.5
log_k -2.0451
-delta_H -12.4683 kJ/mol # Calculated enthalpy of reaction PrAcetate+2
# Enthalpy of formation: -287.88 kcal/mol
-analytic -8.5624e+000 9.3878e-004 -5.7551e+002 2.2087e+000 2.4126e+005
# -Range: 0-300
1.0000 Pr+++ + 1.0000 HCO3- = PrCO3+ +1.0000 H+
-llnl_gamma 4.0
log_k -2.7722
-delta_H 92.458 kJ/mol # Calculated enthalpy of reaction PrCO3+
# Enthalpy of formation: -311.6 kcal/mol
-analytic 2.2079e+002 5.2156e-002 -6.5821e+003 -8.7701e+001 -1.0277e+002
# -Range: 0-300
1.0000 Pr+++ + 1.0000 Cl- = PrCl++
-llnl_gamma 4.5
log_k +0.3086
-delta_H 14.3637 kJ/mol # Calculated enthalpy of reaction PrCl+2
# Enthalpy of formation: -205.3 kcal/mol
-analytic 7.5152e+001 3.7446e-002 -1.6661e+003 -3.2490e+001 -2.6020e+001
# -Range: 0-300
2.0000 Cl- + 1.0000 Pr+++ = PrCl2+
-llnl_gamma 4.0
log_k +0.0308
-delta_H 20.3593 kJ/mol # Calculated enthalpy of reaction PrCl2+
# Enthalpy of formation: -243.8 kcal/mol
-analytic 2.2848e+002 8.1250e-002 -6.0401e+003 -9.3909e+001 -9.4318e+001
# -Range: 0-300
3.0000 Cl- + 1.0000 Pr+++ = PrCl3
-llnl_gamma 3.0
log_k -0.3203
-delta_H 14.2214 kJ/mol # Calculated enthalpy of reaction PrCl3
# Enthalpy of formation: -285.2 kcal/mol
-analytic 4.5016e+002 1.3095e-001 -1.2588e+004 -1.8075e+002 -1.9656e+002
# -Range: 0-300
4.0000 Cl- + 1.0000 Pr+++ = PrCl4-
-llnl_gamma 4.0
log_k -0.7447
-delta_H -4.05011 kJ/mol # Calculated enthalpy of reaction PrCl4-
# Enthalpy of formation: -329.5 kcal/mol
-analytic 5.4245e+002 1.3647e-001 -1.5564e+004 -2.1485e+002 -2.4302e+002
# -Range: 0-300
1.0000 Pr+++ + 1.0000 F- = PrF++
-llnl_gamma 4.5
log_k +4.2221
-delta_H 23.2212 kJ/mol # Calculated enthalpy of reaction PrF+2
# Enthalpy of formation: -243.4 kcal/mol
-analytic 9.5146e+001 4.1115e-002 -2.5463e+003 -3.8236e+001 -3.9760e+001
# -Range: 0-300
2.0000 F- + 1.0000 Pr+++ = PrF2+
-llnl_gamma 4.0
log_k +7.3447
-delta_H 14.644 kJ/mol # Calculated enthalpy of reaction PrF2+
# Enthalpy of formation: -325.6 kcal/mol
-analytic 2.4997e+002 8.5251e-002 -6.1908e+003 -9.9912e+001 -9.6675e+001
# -Range: 0-300
3.0000 F- + 1.0000 Pr+++ = PrF3
-llnl_gamma 3.0
log_k +9.6610
-delta_H -6.4852 kJ/mol # Calculated enthalpy of reaction PrF3
# Enthalpy of formation: -410.8 kcal/mol
-analytic 4.7885e+002 1.3764e-001 -1.2080e+004 -1.8980e+002 -1.8864e+002
# -Range: 0-300
4.0000 F- + 1.0000 Pr+++ = PrF4-
-llnl_gamma 4.0
log_k +11.5375
-delta_H -47.2792 kJ/mol # Calculated enthalpy of reaction PrF4-
# Enthalpy of formation: -500.7 kcal/mol
-analytic 5.5774e+002 1.4067e-001 -1.3523e+004 -2.1933e+002 -2.1118e+002
# -Range: 0-300
1.0000 Pr+++ + 1.0000 HPO4-- + 1.0000 H+ = PrH2PO4++
-llnl_gamma 4.5
log_k +9.5950
-delta_H -16.2548 kJ/mol # Calculated enthalpy of reaction PrH2PO4+2
# Enthalpy of formation: -481.5 kcal/mol
-analytic 1.0501e+002 6.3059e-002 3.8161e+002 -4.6656e+001 5.9234e+000
# -Range: 0-300
1.0000 Pr+++ + 1.0000 HCO3- = PrHCO3++
-llnl_gamma 4.5
log_k +1.9190
-delta_H -12.9788 kJ/mol # Calculated enthalpy of reaction PrHCO3+2
# Enthalpy of formation: -336.8 kcal/mol
-analytic 2.2010e+001 2.8541e-002 1.4574e+003 -1.3522e+001 2.2734e+001
# -Range: 0-300
1.0000 Pr+++ + 1.0000 HPO4-- = PrHPO4+
-llnl_gamma 4.0
log_k +5.4000
-delta_H 0 # Not possible to calculate enthalpy of reaction PrHPO4+
# Enthalpy of formation: -0 kcal/mol
1.0000 Pr+++ + 1.0000 NO3- = PrNO3++
-llnl_gamma 4.5
log_k +0.6546
-delta_H -27.9115 kJ/mol # Calculated enthalpy of reaction PrNO3+2
# Enthalpy of formation: -224.9 kcal/mol
-analytic 1.4297e+001 2.5214e-002 2.1756e+003 -1.1490e+001 3.3943e+001
# -Range: 0-300
1.0000 Pr+++ + 1.0000 H2O = PrO+ +2.0000 H+
-llnl_gamma 4.0
log_k -17.29
-delta_H 117.642 kJ/mol # Calculated enthalpy of reaction PrO+
# Enthalpy of formation: -209 kcal/mol
-analytic 1.7927e+002 2.9467e-002 -1.3815e+004 -6.4259e+001 -2.1562e+002
# -Range: 0-300
2.0000 H2O + 1.0000 Pr+++ = PrO2- +4.0000 H+
-llnl_gamma 4.0
log_k -37.5852
-delta_H 301.39 kJ/mol # Calculated enthalpy of reaction PrO2-
# Enthalpy of formation: -233.4 kcal/mol
-analytic -4.4480e+001 -1.6327e-002 -7.9031e+003 1.9348e+001 -8.5440e+005
# -Range: 0-300
2.0000 H2O + 1.0000 Pr+++ = PrO2H +3.0000 H+
-llnl_gamma 3.0
log_k -26.5901
-delta_H 231.517 kJ/mol # Calculated enthalpy of reaction PrO2H
# Enthalpy of formation: -250.1 kcal/mol
-analytic 3.3930e+002 4.4894e-002 -2.3769e+004 -1.2106e+002 -3.7099e+002
# -Range: 0-300
1.0000 Pr+++ + 1.0000 H2O = PrOH++ +1.0000 H+
-llnl_gamma 4.5
log_k -8.274
-delta_H 81.2407 kJ/mol # Calculated enthalpy of reaction PrOH+2
# Enthalpy of formation: -217.7 kcal/mol
-analytic 5.6599e+001 1.1073e-002 -5.9197e+003 -1.9525e+001 -9.2388e+001
# -Range: 0-300
1.0000 Pr+++ + 1.0000 HPO4-- = PrPO4 +1.0000 H+
-llnl_gamma 3.0
log_k -0.7218
-delta_H 0 # Not possible to calculate enthalpy of reaction PrPO4
# Enthalpy of formation: -0 kcal/mol
1.0000 SO4-- + 1.0000 Pr+++ = PrSO4+
-llnl_gamma 4.0
log_k -3.687
-delta_H 19.6648 kJ/mol # Calculated enthalpy of reaction PrSO4+
# Enthalpy of formation: -381.5 kcal/mol
-analytic 2.9156e+002 8.4671e-002 -1.0638e+004 -1.1509e+002 -1.6608e+002
# -Range: 0-300
2.0000 HPO4-- + 1.0000 Pu++++ = Pu(HPO4)2
-llnl_gamma 3.0
log_k +23.8483
-delta_H 25.9279 kJ/mol # Calculated enthalpy of reaction Pu(HPO4)2
# Enthalpy of formation: -3094.13 kJ/mol
-analytic 9.2387e+002 3.2577e-001 -2.0881e+004 -3.7466e+002 -3.5492e+002
# -Range: 0-200
3.0000 HPO4-- + 1.0000 Pu++++ = Pu(HPO4)3--
-llnl_gamma 4.0
log_k +33.4599
-delta_H -6.49412 kJ/mol # Calculated enthalpy of reaction Pu(HPO4)3-2
# Enthalpy of formation: -4418.63 kJ/mol
-analytic 6.4515e+002 2.3011e-001 -1.2752e+004 -2.5761e+002 -1.9917e+002
# -Range: 0-300
4.0000 HPO4-- + 1.0000 Pu++++ = Pu(HPO4)4----
-llnl_gamma 4.0
log_k +43.2467
-delta_H -77.4832 kJ/mol # Calculated enthalpy of reaction Pu(HPO4)4-4
# Enthalpy of formation: -5781.7 kJ/mol
-analytic 8.5301e+002 3.0730e-001 -1.3644e+004 -3.4573e+002 -2.1316e+002
# -Range: 0-300
2.0000 H2O + 1.0000 Pu++++ = Pu(OH)2++ +2.0000 H+
-llnl_gamma 4.5
log_k -2.3235
-delta_H 74.3477 kJ/mol # Calculated enthalpy of reaction Pu(OH)2+2
# Enthalpy of formation: -1033.22 kJ/mol
-analytic 7.5979e+001 6.8394e-003 -6.3710e+003 -2.3833e+001 -9.9435e+001
# -Range: 0-300
3.0000 H2O + 1.0000 Pu++++ = Pu(OH)3+ +3.0000 H+
-llnl_gamma 4.0
log_k -5.281
-delta_H 96.578 kJ/mol # Calculated enthalpy of reaction Pu(OH)3+
# Enthalpy of formation: -1296.83 kJ/mol
-analytic 1.0874e+002 1.4199e-002 -8.4954e+003 -3.6278e+001 -1.3259e+002
# -Range: 0-300
4.0000 H2O + 1.0000 Pu++++ = Pu(OH)4 +4.0000 H+
-llnl_gamma 3.0
log_k -9.5174
-delta_H 109.113 kJ/mol # Calculated enthalpy of reaction Pu(OH)4
# Enthalpy of formation: -1570.13 kJ/mol
-analytic 2.7913e+002 1.0252e-001 -1.1289e+004 -1.1369e+002 -1.9181e+002
# -Range: 0-200
2.0000 SO4-- + 1.0000 Pu++++ = Pu(SO4)2
-llnl_gamma 3.0
log_k +10.2456
-delta_H 41.0122 kJ/mol # Calculated enthalpy of reaction Pu(SO4)2
# Enthalpy of formation: -2314.08 kJ/mol
-analytic 5.3705e+002 1.9308e-001 -1.3213e+004 -2.1824e+002 -2.2457e+002
# -Range: 0-200
2.0000 SO4-- + 1.0000 Pu+++ = Pu(SO4)2-
-llnl_gamma 4.0
log_k +6.3200
-delta_H 0 # Not possible to calculate enthalpy of reaction Pu(SO4)2-
# Enthalpy of formation: -0 kcal/mol
1.0000 Pu++++ + 1.0000 F- = PuF+++
-llnl_gamma 5.0
log_k +8.4600
-delta_H 0 # Not possible to calculate enthalpy of reaction PuF+3
# Enthalpy of formation: -0 kcal/mol
2.0000 F- + 1.0000 Pu++++ = PuF2++
-llnl_gamma 4.5
log_k +15.4000
-delta_H 0 # Not possible to calculate enthalpy of reaction PuF2+2
# Enthalpy of formation: -0 kcal/mol
3.0000 F- + 1.0000 Pu++++ = PuF3+
-llnl_gamma 4.0
log_k +5.3000
-delta_H 0 # Not possible to calculate enthalpy of reaction PuF3+
# Enthalpy of formation: -0 kcal/mol
4.0000 F- + 1.0000 Pu++++ = PuF4
-llnl_gamma 3.0
log_k +4.2000
-delta_H 0 # Not possible to calculate enthalpy of reaction PuF4
# Enthalpy of formation: -0 kcal/mol
1.0000 Pu+++ + 1.0000 HPO4-- + 1.0000 H+ = PuH2PO4++
-llnl_gamma 4.5
log_k +9.6817
-delta_H 28.597 kJ/mol # Calculated enthalpy of reaction PuH2PO4+2
# Enthalpy of formation: -1855.04 kJ/mol
-analytic 2.1595e+002 6.4502e-002 -6.4723e+003 -8.2341e+001 -1.0106e+002
# -Range: 0-300
1.0000 Pu++++ + 1.0000 HPO4-- = PuHPO4++
-llnl_gamma 4.5
log_k +13.0103
-delta_H 40.306 kJ/mol # Calculated enthalpy of reaction PuHPO4+2
# Enthalpy of formation: -1787.67 kJ/mol
-analytic 2.2662e+002 7.1073e-002 -6.9134e+003 -8.5504e+001 -1.0794e+002
# -Range: 0-300
2.0000 HCO3- + 1.0000 PuO2++ = PuO2(CO3)2-- +2.0000 H+
-llnl_gamma 4.0
log_k -5.7428
-delta_H 52.3345 kJ/mol # Calculated enthalpy of reaction PuO2(CO3)2-2
# Enthalpy of formation: -2149.11 kJ/mol
-analytic 2.6589e+002 7.6132e-002 -9.7187e+003 -1.0577e+002 -1.5173e+002
# -Range: 0-300
1.0000 PuO2++ + 1.0000 Cl- = PuO2Cl+
-llnl_gamma 4.0
log_k -0.2084
-delta_H 11.6127 kJ/mol # Calculated enthalpy of reaction PuO2Cl+
# Enthalpy of formation: -977.045 kJ/mol
-analytic 9.8385e+001 3.8617e-002 -2.5210e+003 -4.1075e+001 -3.9367e+001
# -Range: 0-300
1.0000 PuO2++ + 1.0000 F- = PuO2F+
-llnl_gamma 4.0
log_k +5.6674
-delta_H -5.2094 kJ/mol # Calculated enthalpy of reaction PuO2F+
# Enthalpy of formation: -1162.13 kJ/mol
-analytic 1.1412e+002 4.1224e-002 -2.0503e+003 -4.6009e+001 -3.2027e+001
# -Range: 0-300
2.0000 F- + 1.0000 PuO2++ = PuO2F2
-llnl_gamma 3.0
log_k +10.9669
-delta_H -15.4738 kJ/mol # Calculated enthalpy of reaction PuO2F2
# Enthalpy of formation: -1507.75 kJ/mol
-analytic 2.5502e+002 9.1597e-002 -4.4557e+003 -1.0362e+002 -7.5752e+001
# -Range: 0-200
3.0000 F- + 1.0000 PuO2++ = PuO2F3-
-llnl_gamma 4.0
log_k +15.9160
-delta_H -29.4032 kJ/mol # Calculated enthalpy of reaction PuO2F3-
# Enthalpy of formation: -1857.02 kJ/mol
-analytic 3.6102e+002 8.6364e-002 -8.7129e+003 -1.3805e+002 -1.3606e+002
# -Range: 0-300
4.0000 F- + 1.0000 PuO2++ = PuO2F4--
-llnl_gamma 4.0
log_k +18.7628
-delta_H -39.9786 kJ/mol # Calculated enthalpy of reaction PuO2F4-2
# Enthalpy of formation: -2202.95 kJ/mol
-analytic 4.6913e+002 1.3649e-001 -9.8336e+003 -1.8510e+002 -1.5358e+002
# -Range: 0-300
1.0000 PuO2++ + 1.0000 HPO4-- + 1.0000 H+ = PuO2H2PO4+
-llnl_gamma 4.0
log_k +11.2059
-delta_H -6.63904 kJ/mol # Calculated enthalpy of reaction PuO2H2PO4+
# Enthalpy of formation: -2120.3 kJ/mol
-analytic 2.1053e+002 6.8671e-002 -4.3390e+003 -8.2930e+001 -6.7768e+001
# -Range: 0-300
1.0000 PuO2+ + 1.0000 H2O = PuO2OH +1.0000 H+
-llnl_gamma 3.0
log_k -9.6674
-delta_H 69.1763 kJ/mol # Calculated enthalpy of reaction PuO2OH
# Enthalpy of formation: -1130.85 kJ/mol
-analytic 7.1080e+001 2.6141e-002 -5.0337e+003 -2.8956e+001 -8.5504e+001
# -Range: 0-200
1.0000 PuO2++ + 1.0000 H2O = PuO2OH+ +1.0000 H+
-llnl_gamma 4.0
log_k -5.6379
-delta_H 45.2823 kJ/mol # Calculated enthalpy of reaction PuO2OH+
# Enthalpy of formation: -1062.13 kJ/mol
-analytic -3.9012e+000 1.1645e-003 -1.1299e+003 1.3419e+000 -1.4364e+005
# -Range: 0-300
1.0000 SO4-- + 1.0000 PuO2++ = PuO2SO4
-llnl_gamma 3.0
log_k +3.2658
-delta_H 20.0746 kJ/mol # Calculated enthalpy of reaction PuO2SO4
# Enthalpy of formation: -1711.11 kJ/mol
-analytic 2.0363e+002 7.3903e-002 -5.1940e+003 -8.2833e+001 -8.8273e+001
# -Range: 0-200
1.0000 Pu+++ + 1.0000 H2O = PuOH++ +1.0000 H+
-llnl_gamma 4.5
log_k -7.968
-delta_H 53.5143 kJ/mol # Calculated enthalpy of reaction PuOH+2
# Enthalpy of formation: -823.876 kJ/mol
-analytic 3.0065e+000 3.0278e-003 -1.9675e+003 -1.6100e+000 -1.1524e+005
# -Range: 0-300
1.0000 Pu++++ + 1.0000 H2O = PuOH+++ +1.0000 H+
-llnl_gamma 5.0
log_k -0.5048
-delta_H 48.1823 kJ/mol # Calculated enthalpy of reaction PuOH+3
# Enthalpy of formation: -773.549 kJ/mol
-analytic 4.1056e+001 1.1119e-003 -3.9252e+003 -1.1609e+001 -6.1260e+001
# -Range: 0-300
1.0000 SO4-- + 1.0000 Pu+++ = PuSO4+
-llnl_gamma 4.0
log_k +3.4935
-delta_H 14.6006 kJ/mol # Calculated enthalpy of reaction PuSO4+
# Enthalpy of formation: -1486.55 kJ/mol
-analytic 1.9194e+002 7.7154e-002 -4.2751e+003 -7.9646e+001 -6.6765e+001
# -Range: 0-300
1.0000 SO4-- + 1.0000 Pu++++ = PuSO4++
-llnl_gamma 4.5
log_k +5.7710
-delta_H 12.3336 kJ/mol # Calculated enthalpy of reaction PuSO4+2
# Enthalpy of formation: -1433.16 kJ/mol
-analytic 1.9418e+002 7.5477e-002 -4.2767e+003 -7.9425e+001 -6.6792e+001
# -Range: 0-300
2.0000 HAcetate + 1.0000 Ra++ = Ra(Acetate)2 +2.0000 H+
-llnl_gamma 3.0
log_k -7.9018
-delta_H 21.0874 kJ/mol # Calculated enthalpy of reaction Ra(Acetate)2
# Enthalpy of formation: -353.26 kcal/mol
-analytic 2.2767e+001 3.1254e-003 -6.4558e+003 -7.2253e+000 7.0689e+005
# -Range: 0-300
1.0000 Ra++ + 1.0000 HAcetate = RaAcetate+ +1.0000 H+
-llnl_gamma 4.0
log_k -3.709
-delta_H 11.7989 kJ/mol # Calculated enthalpy of reaction RaAcetate+
# Enthalpy of formation: -239.38 kcal/mol
-analytic -1.8268e+001 2.9956e-003 1.9313e+001 5.2767e+000 4.9771e+004
# -Range: 0-300
2.0000 HAcetate + 1.0000 Rb+ = Rb(Acetate)2- +2.0000 H+
-llnl_gamma 4.0
log_k -9.7636
-delta_H -1.12968 kJ/mol # Calculated enthalpy of reaction Rb(Acetate)2-
# Enthalpy of formation: -292.49 kcal/mol
-analytic -1.9198e+002 -4.2101e-002 5.5792e+003 7.1152e+001 8.7114e+001
# -Range: 0-300
1.0000 Rb+ + 1.0000 Br- = RbBr
-llnl_gamma 3.0
log_k -1.2168
-delta_H 13.9327 kJ/mol # Calculated enthalpy of reaction RbBr
# Enthalpy of formation: -85.73 kcal/mol
-analytic 1.2054e+002 3.3825e-002 -3.9500e+003 -4.7920e+001 -6.1671e+001
# -Range: 0-300
1.0000 Rb+ + 1.0000 HAcetate = RbAcetate +1.0000 H+
-llnl_gamma 3.0
log_k -4.7279
-delta_H 4.89528 kJ/mol # Calculated enthalpy of reaction RbAcetate
# Enthalpy of formation: -174.95 kcal/mol
-analytic 1.5661e+001 -2.4230e-003 -2.5280e+003 -5.4433e+000 2.0344e+005
# -Range: 0-300
1.0000 Rb+ + 1.0000 Cl- = RbCl
-llnl_gamma 3.0
log_k -0.9595
-delta_H 13.1922 kJ/mol # Calculated enthalpy of reaction RbCl
# Enthalpy of formation: -96.8 kcal/mol
-analytic 1.2689e+002 3.5557e-002 -4.0822e+003 -5.0412e+001 -6.3736e+001
# -Range: 0-300
1.0000 Rb+ + 1.0000 F- = RbF
-llnl_gamma 3.0
log_k +0.9602
-delta_H 1.92464 kJ/mol # Calculated enthalpy of reaction RbF
# Enthalpy of formation: -139.71 kcal/mol
-analytic 1.3893e+002 3.8188e-002 -3.8677e+003 -5.5109e+001 -6.0393e+001
# -Range: 0-300
1.0000 Rb+ + 1.0000 I- = RbI
-llnl_gamma 3.0
log_k -0.8136
-delta_H 7.1128 kJ/mol # Calculated enthalpy of reaction RbI
# Enthalpy of formation: -71.92 kcal/mol
-analytic 1.1486e+002 3.3121e-002 -3.4217e+003 -4.6096e+001 -5.3426e+001
# -Range: 0-300
2.0000 Cl- + 1.0000 Ru+++ = Ru(Cl)2+
-llnl_gamma 4.0
log_k +3.7527
-delta_H 0 # Not possible to calculate enthalpy of reaction Ru(Cl)2+
# Enthalpy of formation: -0 kcal/mol
3.0000 Cl- + 1.0000 Ru+++ = Ru(Cl)3
-llnl_gamma 3.0
log_k +4.2976
-delta_H 0 # Not possible to calculate enthalpy of reaction Ru(Cl)3
# Enthalpy of formation: -0 kcal/mol
2.0000 H2O + 1.0000 Ru+++ = Ru(OH)2+ +2.0000 H+
-llnl_gamma 4.0
log_k -3.5148
-delta_H 0 # Not possible to calculate enthalpy of reaction Ru(OH)2+
# Enthalpy of formation: -0 kcal/mol
1.0000 Ru(OH)2++ + 1.0000 Cl- = Ru(OH)2Cl+
-llnl_gamma 4.0
log_k +1.3858
-delta_H 0 # Not possible to calculate enthalpy of reaction Ru(OH)2Cl+
# Enthalpy of formation: -0 kcal/mol
2.0000 Cl- + 1.0000 Ru(OH)2++ = Ru(OH)2Cl2
-llnl_gamma 3.0
log_k +1.8081
-delta_H 0 # Not possible to calculate enthalpy of reaction Ru(OH)2Cl2
# Enthalpy of formation: -0 kcal/mol
3.0000 Cl- + 1.0000 Ru(OH)2++ = Ru(OH)2Cl3-
-llnl_gamma 4.0
log_k +1.6172
-delta_H 0 # Not possible to calculate enthalpy of reaction Ru(OH)2Cl3-
# Enthalpy of formation: -0 kcal/mol
4.0000 Cl- + 1.0000 Ru(OH)2++ = Ru(OH)2Cl4--
-llnl_gamma 4.0
log_k +2.7052
-delta_H 0 # Not possible to calculate enthalpy of reaction Ru(OH)2Cl4-2
# Enthalpy of formation: -0 kcal/mol
1.0000 SO4-- + 1.0000 Ru(OH)2++ = Ru(OH)2SO4
-llnl_gamma 3.0
log_k +1.7941
-delta_H 0 # Not possible to calculate enthalpy of reaction Ru(OH)2SO4
# Enthalpy of formation: -0 kcal/mol
#3.0000 H2O + 1.0000 Ru++ + 0.5000 O2 = Ru(OH)4 +2.0000 H+
# Ru(OH)2++ +1.0000 H2O +0.5000 O2 = 4.0000 H+ + 1.0000 RuO4-- log_k -25.2470
# 4.0000 H+ + 1.0000 RuO4-- = Ru++ +2.0000 H2O +1.0000 O2 log_k +0.1610
#1 + 2 + 3
2H2O + Ru(OH)2++ = Ru(OH)4 + 2H+
-llnl_gamma 3.0
# log_k +18.0322
log_k -7.0538
-delta_H 0 # Not possible to calculate enthalpy of reaction Ru(OH)4
# Enthalpy of formation: -0 kcal/mol
2.0000 SO4-- + 1.0000 Ru+++ = Ru(SO4)2-
-llnl_gamma 4.0
log_k +3.0627
-delta_H 0 # Not possible to calculate enthalpy of reaction Ru(SO4)2-
# Enthalpy of formation: -0 kcal/mol
4.0000 Ru(OH)2++ + 4.0000 H2O = Ru4(OH)12++++ +4.0000 H+
-llnl_gamma 5.5
log_k +7.1960
-delta_H 0 # Not possible to calculate enthalpy of reaction Ru4(OH)12+4
# Enthalpy of formation: -0 kcal/mol
1.0000 Ru++ + 1.0000 Cl- = RuCl+
-llnl_gamma 4.0
log_k -0.4887
-delta_H 0 # Not possible to calculate enthalpy of reaction RuCl+
# Enthalpy of formation: -0 kcal/mol
1.0000 Ru+++ + 1.0000 Cl- = RuCl++
-llnl_gamma 4.5
log_k +2.1742
-delta_H 0 # Not possible to calculate enthalpy of reaction RuCl+2
# Enthalpy of formation: -0 kcal/mol
4.0000 Cl- + 1.0000 Ru+++ = RuCl4-
-llnl_gamma 4.0
log_k +4.1418
-delta_H 0 # Not possible to calculate enthalpy of reaction RuCl4-
# Enthalpy of formation: -0 kcal/mol
5.0000 Cl- + 1.0000 Ru+++ = RuCl5--
-llnl_gamma 4.0
log_k +3.8457
-delta_H 0 # Not possible to calculate enthalpy of reaction RuCl5-2
# Enthalpy of formation: -0 kcal/mol
6.0000 Cl- + 1.0000 Ru+++ = RuCl6---
-llnl_gamma 4.0
log_k +3.4446
-delta_H 0 # Not possible to calculate enthalpy of reaction RuCl6-3
# Enthalpy of formation: -0 kcal/mol
1.0000 Ru+++ + 1.0000 H2O = RuOH++ +1.0000 H+
-llnl_gamma 4.5
log_k -2.2392
-delta_H 0 # Not possible to calculate enthalpy of reaction RuOH+2
# Enthalpy of formation: -0 kcal/mol
1.0000 SO4-- + 1.0000 Ru++ = RuSO4
-llnl_gamma 3.0
log_k +2.3547
-delta_H 0 # Not possible to calculate enthalpy of reaction RuSO4
# Enthalpy of formation: -0 kcal/mol
1.0000 SO4-- + 1.0000 Ru+++ = RuSO4+
-llnl_gamma 4.0
log_k +1.9518
-delta_H 0 # Not possible to calculate enthalpy of reaction RuSO4+
# Enthalpy of formation: -0 kcal/mol
1.0000 HS- = S-- +1.0000 H+
-llnl_gamma 5.0
log_k -12.9351
-delta_H 49.0364 kJ/mol # Calculated enthalpy of reaction S-2
# Enthalpy of formation: 32.928 kJ/mol
-analytic 9.7756e+001 3.2913e-002 -5.0784e+003 -4.1812e+001 -7.9273e+001
# -Range: 0-300
2.0000 H+ + 2.0000 SO3-- = S2O5-- + H2O
-llnl_gamma 4.0
log_k 9.5934
-delta_H 0 # Not possible to calculate enthalpy of reaction S2O5-2
# Enthalpy of formation: -0 kcal/mol
-analytic 0.12262E+03 0.62883E-01 -0.18005E+04 -0.50798E+02 -0.28132E+02
# -Range: 0-300
2.0000 H+ + 1.0000 SO3-- = SO2 +1.0000 H2O
-llnl_gamma 3.0
log_k +9.0656
-delta_H 26.7316 kJ/mol # Calculated enthalpy of reaction SO2
# Enthalpy of formation: -77.194 kcal/mol
-analytic 9.4048e+001 6.2127e-002 -1.1072e+003 -4.0310e+001 -1.7305e+001
# -Range: 0-300
1.0000 Sb(OH)3 + 1.0000 H+ = Sb(OH)2+ +1.0000 H2O
-llnl_gamma 4.0
log_k +1.4900
-delta_H 0 # Not possible to calculate enthalpy of reaction Sb(OH)2+
# Enthalpy of formation: -0 kcal/mol
-analytic -4.9192e+000 -1.6439e-004 1.4777e+003 6.0724e-001 2.3059e+001
# -Range: 0-300
1.0000 Sb(OH)3 + 1.0000 H+ + 1.0000 F- = Sb(OH)2F +1.0000 H2O
-llnl_gamma 3.0
log_k +7.1700
-delta_H 0 # Not possible to calculate enthalpy of reaction Sb(OH)2F
# Enthalpy of formation: -0 kcal/mol
-analytic -1.6961e+002 5.7364e-002 2.7207e+004 3.7969e+001 -2.2834e+006
# -Range: 0-300
1.0000 Sb(OH)3 + 1.0000 H2O = Sb(OH)4- +1.0000 H+
-llnl_gamma 4.0
log_k -11.92
-delta_H 0 # Not possible to calculate enthalpy of reaction Sb(OH)4-
# Enthalpy of formation: -0 kcal/mol
-analytic 4.9839e+001 -6.7112e-003 -4.8976e+003 -1.7138e+001 -8.3725e+004
# -Range: 0-300
4.0000 HS- + 2.0000 Sb(OH)3 + 2.0000 H+ = Sb2S4-- +6.0000 H2O
-llnl_gamma 4.0
log_k +39.1100
-delta_H 0 # Not possible to calculate enthalpy of reaction Sb2S4-2
# Enthalpy of formation: -0 kcal/mol
-analytic 1.7631e+002 8.3686e-002 9.7091e+003 -7.8605e+001 1.5145e+002
# -Range: 0-300
4.0000 Cl- + 3.0000 H+ + 1.0000 Sb(OH)3 = SbCl4- +3.0000 H2O
-llnl_gamma 4.0
log_k +3.0720
-delta_H 0 # Not possible to calculate enthalpy of reaction SbCl4-
# Enthalpy of formation: -0 kcal/mol
2.0000 HAcetate + 1.0000 Sc+++ = Sc(Acetate)2+ +2.0000 H+
-llnl_gamma 4.0
log_k -3.7237
-delta_H -43.1789 kJ/mol # Calculated enthalpy of reaction Sc(Acetate)2+
# Enthalpy of formation: -389.32 kcal/mol
-analytic -4.1862e+001 -3.9443e-005 2.1444e+002 1.2616e+001 5.5442e+005
# -Range: 0-300
3.0000 HAcetate + 1.0000 Sc+++ = Sc(Acetate)3 +3.0000 H+
-llnl_gamma 3.0
log_k -6.6777
-delta_H -70.0402 kJ/mol # Calculated enthalpy of reaction Sc(Acetate)3
# Enthalpy of formation: -511.84 kcal/mol
-analytic -5.2525e+001 1.6181e-003 7.5022e+002 1.3988e+001 7.3540e+005
# -Range: 0-300
1.0000 Sc+++ + 1.0000 HAcetate = ScAcetate++ +1.0000 H+
-llnl_gamma 4.5
log_k -1.4294
-delta_H -21.7568 kJ/mol # Calculated enthalpy of reaction ScAcetate+2
# Enthalpy of formation: -268.1 kcal/mol
-analytic -2.3400e+001 1.3144e-004 1.1125e+002 7.3527e+000 3.0025e+005
# -Range: 0-300
6.0000 F- + 4.0000 H+ + 1.0000 SiO2 = SiF6-- +2.0000 H2O
-llnl_gamma 4.0
log_k +26.2749
-delta_H -70.9565 kJ/mol # Calculated enthalpy of reaction SiF6-2
# Enthalpy of formation: -571 kcal/mol
-analytic 2.3209e+002 1.0685e-001 5.8428e+002 -9.6798e+001 9.0486e+000
# -Range: 0-300
2.0000 HAcetate + 1.0000 Sm+++ = Sm(Acetate)2+ +2.0000 H+
-llnl_gamma 4.0
log_k -4.7132
-delta_H -25.5224 kJ/mol # Calculated enthalpy of reaction Sm(Acetate)2+
# Enthalpy of formation: -403.5 kcal/mol
-analytic -1.4192e+001 2.1732e-003 -1.0267e+003 2.9516e+000 4.4389e+005
# -Range: 0-300
3.0000 HAcetate + 1.0000 Sm+++ = Sm(Acetate)3 +3.0000 H+
-llnl_gamma 3.0
log_k -7.8798
-delta_H -43.5554 kJ/mol # Calculated enthalpy of reaction Sm(Acetate)3
# Enthalpy of formation: -523.91 kcal/mol
-analytic -2.0765e+001 1.1047e-003 -5.1181e+002 3.4797e+000 5.0618e+005
# -Range: 0-300
2.0000 HCO3- + 1.0000 Sm+++ = Sm(CO3)2- +2.0000 H+
-llnl_gamma 4.0
log_k -7.8576
-delta_H 0 # Not possible to calculate enthalpy of reaction Sm(CO3)2-
# Enthalpy of formation: -0 kcal/mol
2.0000 HPO4-- + 1.0000 Sm+++ = Sm(HPO4)2-
-llnl_gamma 4.0
log_k +9.4000
-delta_H 0 # Not possible to calculate enthalpy of reaction Sm(HPO4)2-
# Enthalpy of formation: -0 kcal/mol
# Redundant with SmO2-
#4.0000 H2O + 1.0000 Sm+++ = Sm(OH)4- +4.0000 H+
# -llnl_gamma 4.0
# log_k -36.8803
# -delta_H 0 # Not possible to calculate enthalpy of reaction Sm(OH)4-
## Enthalpy of formation: -0 kcal/mol
2.0000 HPO4-- + 1.0000 Sm+++ = Sm(PO4)2--- +2.0000 H+
-llnl_gamma 4.0
log_k -4.2437
-delta_H 0 # Not possible to calculate enthalpy of reaction Sm(PO4)2-3
# Enthalpy of formation: -0 kcal/mol
2.0000 SO4-- + 1.0000 Sm+++ = Sm(SO4)2-
-llnl_gamma 4.0
log_k +5.2000
-delta_H 0 # Not possible to calculate enthalpy of reaction Sm(SO4)2-
# Enthalpy of formation: -0 kcal/mol
1.0000 Sm+++ + 1.0000 HAcetate = SmAcetate++ +1.0000 H+
-llnl_gamma 4.5
log_k -1.9205
-delta_H -13.598 kJ/mol # Calculated enthalpy of reaction SmAcetate+2
# Enthalpy of formation: -284.55 kcal/mol
-analytic -1.1734e+001 1.0889e-003 -5.1061e+002 3.3317e+000 2.6395e+005
# -Range: 0-300
1.0000 Sm+++ + 1.0000 HCO3- = SmCO3+ +1.0000 H+
-llnl_gamma 4.0
log_k -2.479
-delta_H 89.1108 kJ/mol # Calculated enthalpy of reaction SmCO3+
# Enthalpy of formation: -308.8 kcal/mol
-analytic 2.3486e+002 5.3703e-002 -7.0193e+003 -9.2863e+001 -1.0960e+002
# -Range: 0-300
1.0000 Sm+++ + 1.0000 Cl- = SmCl++
-llnl_gamma 4.5
log_k +0.3086
-delta_H 14.3637 kJ/mol # Calculated enthalpy of reaction SmCl+2
# Enthalpy of formation: -201.7 kcal/mol
-analytic 9.4972e+001 3.9428e-002 -2.4198e+003 -3.9718e+001 -3.7787e+001
# -Range: 0-300
2.0000 Cl- + 1.0000 Sm+++ = SmCl2+
-llnl_gamma 4.0
log_k -0.0425
-delta_H 19.9409 kJ/mol # Calculated enthalpy of reaction SmCl2+
# Enthalpy of formation: -240.3 kcal/mol
-analytic 2.5872e+002 8.4154e-002 -7.2061e+003 -1.0493e+002 -1.1252e+002
# -Range: 0-300
3.0000 Cl- + 1.0000 Sm+++ = SmCl3
-llnl_gamma 3.0
log_k -0.3936
-delta_H 13.803 kJ/mol # Calculated enthalpy of reaction SmCl3
# Enthalpy of formation: -281.7 kcal/mol
-analytic 4.9535e+002 1.3520e-001 -1.4325e+004 -1.9720e+002 -2.2367e+002
# -Range: 0-300
4.0000 Cl- + 1.0000 Sm+++ = SmCl4-
-llnl_gamma 4.0
log_k -0.818
-delta_H -5.30531 kJ/mol # Calculated enthalpy of reaction SmCl4-
# Enthalpy of formation: -326.2 kcal/mol
-analytic 6.0562e+002 1.4212e-001 -1.7982e+004 -2.3782e+002 -2.8077e+002
# -Range: 0-300
1.0000 Sm+++ + 1.0000 F- = SmF++
-llnl_gamma 4.5
log_k +4.3687
-delta_H 22.8028 kJ/mol # Calculated enthalpy of reaction SmF+2
# Enthalpy of formation: -239.9 kcal/mol
-analytic 1.1514e+002 4.3117e-002 -3.2853e+003 -4.5499e+001 -5.1297e+001
# -Range: 0-300
2.0000 F- + 1.0000 Sm+++ = SmF2+
-llnl_gamma 4.0
log_k +7.6379
-delta_H 13.8072 kJ/mol # Calculated enthalpy of reaction SmF2+
# Enthalpy of formation: -322.2 kcal/mol
-analytic 2.8030e+002 8.8143e-002 -7.2857e+003 -1.1092e+002 -1.1377e+002
# -Range: 0-300
3.0000 F- + 1.0000 Sm+++ = SmF3
-llnl_gamma 3.0
log_k +10.0275
-delta_H -8.5772 kJ/mol # Calculated enthalpy of reaction SmF3
# Enthalpy of formation: -407.7 kcal/mol
-analytic 5.2425e+002 1.4191e-001 -1.3728e+004 -2.0628e+002 -2.1436e+002
# -Range: 0-300
4.0000 F- + 1.0000 Sm+++ = SmF4-
-llnl_gamma 4.0
log_k +11.9773
-delta_H -49.7896 kJ/mol # Calculated enthalpy of reaction SmF4-
# Enthalpy of formation: -497.7 kcal/mol
-analytic 6.2228e+002 1.4659e-001 -1.5887e+004 -2.4275e+002 -2.4809e+002
# -Range: 0-300
1.0000 Sm+++ + 1.0000 HPO4-- + 1.0000 H+ = SmH2PO4++
-llnl_gamma 4.5
log_k +9.4484
-delta_H -15.8364 kJ/mol # Calculated enthalpy of reaction SmH2PO4+2
# Enthalpy of formation: -477.8 kcal/mol
-analytic 1.2451e+002 6.4959e-002 -3.9576e+002 -5.3772e+001 -6.2124e+000
# -Range: 0-300
1.0000 Sm+++ + 1.0000 HCO3- = SmHCO3++
-llnl_gamma 4.5
log_k +1.7724
-delta_H 9.19643 kJ/mol # Calculated enthalpy of reaction SmHCO3+2
# Enthalpy of formation: -327.9 kcal/mol
-analytic 5.5520e+001 3.3265e-002 -7.3142e+002 -2.4727e+001 -1.1430e+001
# -Range: 0-300
1.0000 Sm+++ + 1.0000 HPO4-- = SmHPO4+
-llnl_gamma 4.0
log_k +5.6000
-delta_H 0 # Not possible to calculate enthalpy of reaction SmHPO4+
# Enthalpy of formation: -0 kcal/mol
1.0000 Sm+++ + 1.0000 NO3- = SmNO3++
-llnl_gamma 4.5
log_k +0.8012
-delta_H -29.1667 kJ/mol # Calculated enthalpy of reaction SmNO3+2
# Enthalpy of formation: -221.6 kcal/mol
-analytic 3.3782e+001 2.7125e-002 1.5091e+003 -1.8632e+001 2.3537e+001
# -Range: 0-300
1.0000 Sm+++ + 1.0000 H2O = SmO+ +2.0000 H+
-llnl_gamma 4.0
log_k -16.4837
-delta_H 113.039 kJ/mol # Calculated enthalpy of reaction SmO+
# Enthalpy of formation: -206.5 kcal/mol
-analytic 1.8554e+002 3.0198e-002 -1.3791e+004 -6.6588e+001 -2.1526e+002
# -Range: 0-300
2.0000 H2O + 1.0000 Sm+++ = SmO2- +4.0000 H+
-llnl_gamma 4.0
log_k -35.0197
-delta_H 285.909 kJ/mol # Calculated enthalpy of reaction SmO2-
# Enthalpy of formation: -233.5 kcal/mol
-analytic 1.3508e+001 -8.3384e-003 -1.0325e+004 -1.5506e+000 -6.7392e+005
# -Range: 0-300
2.0000 H2O + 1.0000 Sm+++ = SmO2H +3.0000 H+
-llnl_gamma 3.0
log_k -25.9304
-delta_H 226.497 kJ/mol # Calculated enthalpy of reaction SmO2H
# Enthalpy of formation: -247.7 kcal/mol
-analytic 3.6882e+002 5.3761e-002 -2.4317e+004 -1.3305e+002 -3.7956e+002
# -Range: 0-300
1.0000 Sm+++ + 1.0000 H2O = SmOH++ +1.0000 H+
-llnl_gamma 4.5
log_k -7.9808
-delta_H 79.1487 kJ/mol # Calculated enthalpy of reaction SmOH+2
# Enthalpy of formation: -214.6 kcal/mol
-analytic 6.3793e+001 1.1977e-002 -6.0852e+003 -2.2198e+001 -9.4972e+001
# -Range: 0-300
1.0000 Sm+++ + 1.0000 HPO4-- = SmPO4 +1.0000 H+
-llnl_gamma 3.0
log_k -0.2218
-delta_H 0 # Not possible to calculate enthalpy of reaction SmPO4
# Enthalpy of formation: -0 kcal/mol
1.0000 Sm+++ + 1.0000 SO4-- = SmSO4+
-llnl_gamma 4.0
log_k +3.6430
-delta_H 20.0832 kJ/mol # Calculated enthalpy of reaction SmSO4+
# Enthalpy of formation: -377.8 kcal/mol
-analytic 3.0597e+002 8.6258e-002 -9.0231e+003 -1.2032e+002 -1.4089e+002
# -Range: 0-300
2.0000 H2O + 1.0000 Sn++ = Sn(OH)2 +2.0000 H+
-llnl_gamma 3.0
log_k -7.9102
-delta_H 42.0534 kJ/mol # Calculated enthalpy of reaction Sn(OH)2
# Enthalpy of formation: -128.683 kcal/mol
-analytic -3.7979e+001 -1.0893e-002 -1.2048e+003 1.5100e+001 -2.0445e+001
# -Range: 0-200
2.0000 H2O + 1.0000 Sn++++ = Sn(OH)2++ +2.0000 H+
-llnl_gamma 4.5
log_k -0.1902
-delta_H -2.02087 kJ/mol # Calculated enthalpy of reaction Sn(OH)2+2
# Enthalpy of formation: -129.888 kcal/mol
-analytic -2.1675e+001 5.9697e-003 3.3953e+003 4.8158e+000 -3.2042e+005
# -Range: 0-300
3.0000 H2O + 1.0000 Sn++++ = Sn(OH)3+ +3.0000 H+
-llnl_gamma 4.0
log_k +0.5148
-delta_H -7.59396 kJ/mol # Calculated enthalpy of reaction Sn(OH)3+
# Enthalpy of formation: -199.537 kcal/mol
-analytic -3.3294e+001 8.8580e-003 5.3803e+003 7.4994e+000 -4.8389e+005
# -Range: 0-300
3.0000 H2O + 1.0000 Sn++ = Sn(OH)3- +3.0000 H+
-llnl_gamma 4.0
log_k -17.4052
-delta_H 94.7007 kJ/mol # Calculated enthalpy of reaction Sn(OH)3-
# Enthalpy of formation: -184.417 kcal/mol
-analytic 1.5614e+002 1.9943e-002 -1.0700e+004 -5.8031e+001 -1.6701e+002
# -Range: 0-300
4.0000 H2O + 1.0000 Sn++++ = Sn(OH)4 +4.0000 H+
-llnl_gamma 3.0
log_k +0.8497
-delta_H -11.0583 kJ/mol # Calculated enthalpy of reaction Sn(OH)4
# Enthalpy of formation: -268.682 kcal/mol
-analytic -7.9563e+001 -2.2641e-002 2.6682e+003 3.1614e+001 4.5337e+001
# -Range: 0-200
2.0000 SO4-- + 1.0000 Sn++++ = Sn(SO4)2
-llnl_gamma 3.0
log_k -0.8072
-delta_H 0 # Not possible to calculate enthalpy of reaction Sn(SO4)2
# Enthalpy of formation: -0 kcal/mol
1.0000 Sn++ + 1.0000 Cl- = SnCl+
-llnl_gamma 4.0
log_k +1.0500
-delta_H 0 # Not possible to calculate enthalpy of reaction SnCl+
# Enthalpy of formation: -0 kcal/mol
-analytic 3.0558e+002 8.2458e-002 -8.9329e+003 -1.2088e+002 -1.3948e+002
# -Range: 0-300
2.0000 Cl- + 1.0000 Sn++ = SnCl2
-llnl_gamma 3.0
log_k +1.7100
-delta_H 0 # Not possible to calculate enthalpy of reaction SnCl2
# Enthalpy of formation: -0 kcal/mol
-analytic 3.6600e+002 1.0753e-001 -1.0006e+004 -1.4660e+002 -1.5624e+002
# -Range: 0-300
3.0000 Cl- + 1.0000 Sn++ = SnCl3-
-llnl_gamma 4.0
log_k +1.6900
-delta_H 0 # Not possible to calculate enthalpy of reaction SnCl3-
# Enthalpy of formation: -0 kcal/mol
-analytic 3.6019e+002 1.0602e-001 -1.0337e+004 -1.4363e+002 -1.6141e+002
# -Range: 0-300
1.0000 Sn++ + 1.0000 F- = SnF+
-llnl_gamma 4.0
log_k +4.0800
-delta_H 0 # Not possible to calculate enthalpy of reaction SnF+
# Enthalpy of formation: -0 kcal/mol
-analytic 3.0020e+002 7.5485e-002 -8.4231e+003 -1.1734e+002 -1.3152e+002
# -Range: 0-300
2.0000 F- + 1.0000 Sn++ = SnF2
-llnl_gamma 3.0
log_k +6.6800
-delta_H 0 # Not possible to calculate enthalpy of reaction SnF2
# Enthalpy of formation: -0 kcal/mol
-analytic 4.1241e+002 1.0988e-001 -1.1151e+004 -1.6207e+002 -1.7413e+002
# -Range: 0-300
3.0000 F- + 1.0000 Sn++ = SnF3-
-llnl_gamma 4.0
log_k +9.4600
-delta_H 0 # Not possible to calculate enthalpy of reaction SnF3-
# Enthalpy of formation: -0 kcal/mol
-analytic 4.1793e+002 1.0898e-001 -1.1402e+004 -1.6273e+002 -1.7803e+002
# -Range: 0-300
1.0000 Sn++ + 1.0000 H2O = SnOH+ +1.0000 H+
-llnl_gamma 4.0
log_k -3.9851
-delta_H 21.2045 kJ/mol # Calculated enthalpy of reaction SnOH+
# Enthalpy of formation: -65.349 kcal/mol
-analytic 7.7253e+001 1.9149e-002 -3.3745e+003 -3.0560e+001 -5.2679e+001
# -Range: 0-300
1.0000 Sn++++ + 1.0000 H2O = SnOH+++ +1.0000 H+
-llnl_gamma 5.0
log_k +0.6049
-delta_H -5.00406 kJ/mol # Calculated enthalpy of reaction SnOH+3
# Enthalpy of formation: -62.284 kcal/mol
-analytic -1.1548e+001 2.8878e-003 1.9476e+003 2.6622e+000 -1.6274e+005
# -Range: 0-300
1.0000 Sn++++ + 1.0000 SO4-- = SnSO4++
-llnl_gamma 4.5
log_k -3.1094
-delta_H 0 # Not possible to calculate enthalpy of reaction SnSO4+2
# Enthalpy of formation: -0 kcal/mol
2.0000 HAcetate + 1.0000 Sr++ = Sr(Acetate)2 +2.0000 H+
-llnl_gamma 3.0
log_k -7.8212
-delta_H 0.54392 kJ/mol # Calculated enthalpy of reaction Sr(Acetate)2
# Enthalpy of formation: -363.74 kcal/mol
-analytic 1.2965e+001 4.7082e-003 -5.2538e+003 -5.2337e+000 7.4721e+005
# -Range: 0-300
1.0000 Sr++ + 1.0000 HAcetate = SrAcetate+ +1.0000 H+
-llnl_gamma 4.0
log_k -3.6724
-delta_H 2.3012 kJ/mol # Calculated enthalpy of reaction SrAcetate+
# Enthalpy of formation: -247.22 kcal/mol
-analytic -1.4301e+001 1.2481e-003 -7.5690e+002 4.2760e+000 1.9800e+005
# -Range: 0-300
1.0000 Sr++ + 1.0000 HCO3- = SrCO3 +1.0000 H+
-llnl_gamma 3.0
log_k -7.4635
-delta_H 33.2544 kJ/mol # Calculated enthalpy of reaction SrCO3
# Enthalpy of formation: -288.62 kcal/mol
-analytic 2.2303e+002 5.2582e-002 -8.4861e+003 -8.7975e+001 -1.3248e+002
# -Range: 0-300
1.0000 Sr++ + 1.0000 Cl- = SrCl+
-llnl_gamma 4.0
log_k -0.2485
-delta_H 7.58559 kJ/mol # Calculated enthalpy of reaction SrCl+
# Enthalpy of formation: -169.79 kcal/mol
-analytic 9.4568e+001 3.9042e-002 -2.1458e+003 -4.0105e+001 -3.3511e+001
# -Range: 0-300
1.0000 Sr++ + 1.0000 F- = SrF+
-llnl_gamma 4.0
log_k +0.1393
-delta_H 4.8116 kJ/mol # Calculated enthalpy of reaction SrF+
# Enthalpy of formation: -210.67 kcal/mol
-analytic 9.0295e+001 3.7609e-002 -1.9012e+003 -3.8379e+001 -2.9693e+001
# -Range: 0-300
1.0000 Sr++ + 1.0000 HPO4-- + 1.0000 H+ = SrH2PO4+
-llnl_gamma 4.0
log_k +0.7300
-delta_H 0 # Not possible to calculate enthalpy of reaction SrH2PO4+
# Enthalpy of formation: -0 kcal/mol
1.0000 Sr++ + 1.0000 HPO4-- = SrHPO4
-llnl_gamma 3.0
log_k +2.0600
-delta_H 0 # Not possible to calculate enthalpy of reaction SrHPO4
# Enthalpy of formation: -0 kcal/mol
1.0000 Sr++ + 1.0000 NO3- = SrNO3+
-llnl_gamma 4.0
log_k +0.8000
-delta_H 0 # Not possible to calculate enthalpy of reaction SrNO3+
# Enthalpy of formation: -0 kcal/mol
1.0000 Sr++ + 1.0000 H2O = SrOH+ +1.0000 H+
-llnl_gamma 4.0
log_k -13.29
-delta_H 0 # Not possible to calculate enthalpy of reaction SrOH+
# Enthalpy of formation: -0 kcal/mol
2.0000 HPO4-- + 1.0000 Sr++ = SrP2O7-- +1.0000 H2O
-llnl_gamma 4.0
log_k +1.6537
-delta_H 0 # Not possible to calculate enthalpy of reaction SrP2O7-2
# Enthalpy of formation: -0 kcal/mol
1.0000 Sr++ + 1.0000 SO4-- = SrSO4
-llnl_gamma 3.0
log_k +2.3000
-delta_H 0 # Not possible to calculate enthalpy of reaction SrSO4
# Enthalpy of formation: -0 kcal/mol
2.0000 HAcetate + 1.0000 Tb+++ = Tb(Acetate)2+ +2.0000 H+
-llnl_gamma 4.0
log_k -4.9625
-delta_H -27.9491 kJ/mol # Calculated enthalpy of reaction Tb(Acetate)2+
# Enthalpy of formation: -405.78 kcal/mol
-analytic -2.3910e+001 1.3433e-003 -8.0800e+002 6.3895e+000 4.8619e+005
# -Range: 0-300
3.0000 HAcetate + 1.0000 Tb+++ = Tb(Acetate)3 +3.0000 H+
-llnl_gamma 3.0
log_k -8.3489
-delta_H -47.1537 kJ/mol # Calculated enthalpy of reaction Tb(Acetate)3
# Enthalpy of formation: -526.47 kcal/mol
-analytic -1.0762e+001 4.2361e-003 -1.5620e+003 -3.9317e-001 6.5745e+005
# -Range: 0-300
2.0000 HCO3- + 1.0000 Tb+++ = Tb(CO3)2- +2.0000 H+
-llnl_gamma 4.0
log_k -7.5576
-delta_H 0 # Not possible to calculate enthalpy of reaction Tb(CO3)2-
# Enthalpy of formation: -0 kcal/mol
2.0000 HPO4-- + 1.0000 Tb+++ = Tb(HPO4)2-
-llnl_gamma 4.0
log_k +9.7000
-delta_H 0 # Not possible to calculate enthalpy of reaction Tb(HPO4)2-
# Enthalpy of formation: -0 kcal/mol
2.0000 HPO4-- + 1.0000 Tb+++ = Tb(PO4)2--- +2.0000 H+
-llnl_gamma 4.0
log_k -3.6437
-delta_H 0 # Not possible to calculate enthalpy of reaction Tb(PO4)2-3
# Enthalpy of formation: -0 kcal/mol
2.0000 SO4-- + 1.0000 Tb+++ = Tb(SO4)2-
-llnl_gamma 4.0
log_k +5.0000
-delta_H 0 # Not possible to calculate enthalpy of reaction Tb(SO4)2-
# Enthalpy of formation: -0 kcal/mol
1.0000 Tb+++ + 1.0000 HAcetate = TbAcetate++ +1.0000 H+
-llnl_gamma 4.5
log_k -2.1037
-delta_H -14.2256 kJ/mol # Calculated enthalpy of reaction TbAcetate+2
# Enthalpy of formation: -286.4 kcal/mol
-analytic -1.6817e+001 6.4290e-004 -3.4442e+002 5.0994e+000 2.7304e+005
# -Range: 0-300
1.0000 Tb+++ + 1.0000 HCO3- = TbCO3+ +1.0000 H+
-llnl_gamma 4.0
log_k -2.4057
-delta_H 89.5292 kJ/mol # Calculated enthalpy of reaction TbCO3+
# Enthalpy of formation: -310.4 kcal/mol
-analytic 2.2347e+002 5.4185e-002 -6.4127e+003 -8.9112e+001 -1.0013e+002
# -Range: 0-300
1.0000 Tb+++ + 1.0000 Cl- = TbCl++
-llnl_gamma 4.5
log_k +0.2353
-delta_H 13.9453 kJ/mol # Calculated enthalpy of reaction TbCl+2
# Enthalpy of formation: -203.5 kcal/mol
-analytic 7.1095e+001 3.7367e-002 -1.4676e+003 -3.1140e+001 -2.2921e+001
# -Range: 0-300
2.0000 Cl- + 1.0000 Tb+++ = TbCl2+
-llnl_gamma 4.0
log_k -0.0425
-delta_H 18.2673 kJ/mol # Calculated enthalpy of reaction TbCl2+
# Enthalpy of formation: -242.4 kcal/mol
-analytic 2.0699e+002 7.9609e-002 -5.0958e+003 -8.6337e+001 -7.9576e+001
# -Range: 0-300
3.0000 Cl- + 1.0000 Tb+++ = TbCl3
-llnl_gamma 3.0
log_k -0.4669
-delta_H 10.0374 kJ/mol # Calculated enthalpy of reaction TbCl3
# Enthalpy of formation: -284.3 kcal/mol
-analytic 4.0764e+002 1.2809e-001 -1.0704e+004 -1.6583e+002 -1.6715e+002
# -Range: 0-300
4.0000 Cl- + 1.0000 Tb+++ = TbCl4-
-llnl_gamma 4.0
log_k -0.8913
-delta_H -11.5813 kJ/mol # Calculated enthalpy of reaction TbCl4-
# Enthalpy of formation: -329.4 kcal/mol
-analytic 4.6247e+002 1.2926e-001 -1.2117e+004 -1.8639e+002 -1.8921e+002
# -Range: 0-300
1.0000 Tb+++ + 1.0000 F- = TbF++
-llnl_gamma 4.5
log_k +4.6619
-delta_H 22.8028 kJ/mol # Calculated enthalpy of reaction TbF+2
# Enthalpy of formation: -241.6 kcal/mol
-analytic 9.2579e+001 4.1327e-002 -2.3647e+003 -3.7293e+001 -3.6927e+001
# -Range: 0-300
2.0000 F- + 1.0000 Tb+++ = TbF2+
-llnl_gamma 4.0
log_k +8.1510
-delta_H 12.1336 kJ/mol # Calculated enthalpy of reaction TbF2+
# Enthalpy of formation: -324.3 kcal/mol
-analytic 2.3100e+002 8.4094e-002 -5.2548e+003 -9.3051e+001 -8.2065e+001
# -Range: 0-300
3.0000 F- + 1.0000 Tb+++ = TbF3
-llnl_gamma 3.0
log_k +10.6872
-delta_H -11.9244 kJ/mol # Calculated enthalpy of reaction TbF3
# Enthalpy of formation: -410.2 kcal/mol
-analytic 4.3730e+002 1.3479e-001 -1.0128e+004 -1.7489e+002 -1.5817e+002
# -Range: 0-300
4.0000 F- + 1.0000 Tb+++ = TbF4-
-llnl_gamma 4.0
log_k +12.7836
-delta_H -56.0656 kJ/mol # Calculated enthalpy of reaction TbF4-
# Enthalpy of formation: -500.9 kcal/mol
-analytic 4.8546e+002 1.3511e-001 -1.0189e+004 -1.9347e+002 -1.5913e+002
# -Range: 0-300
1.0000 Tb+++ + 1.0000 HPO4-- + 1.0000 H+ = TbH2PO4++
-llnl_gamma 4.5
log_k +9.3751
-delta_H -17.51 kJ/mol # Calculated enthalpy of reaction TbH2PO4+2
# Enthalpy of formation: -479.9 kcal/mol
-analytic 1.0042e+002 6.2886e-002 6.0975e+002 -4.5178e+001 9.4847e+000
# -Range: 0-300
1.0000 Tb+++ + 1.0000 HCO3- = TbHCO3++
-llnl_gamma 4.5
log_k +1.6991
-delta_H -14.6524 kJ/mol # Calculated enthalpy of reaction TbHCO3+2
# Enthalpy of formation: -335.3 kcal/mol
-analytic 1.7376e+001 2.8365e-002 1.6982e+003 -1.2044e+001 2.6494e+001
# -Range: 0-300
1.0000 Tb+++ + 1.0000 HPO4-- = TbHPO4+
-llnl_gamma 4.0
log_k +5.8000
-delta_H 0 # Not possible to calculate enthalpy of reaction TbHPO4+
# Enthalpy of formation: -0 kcal/mol
1.0000 Tb+++ + 1.0000 NO3- = TbNO3++
-llnl_gamma 4.5
log_k +0.5080
-delta_H -31.2587 kJ/mol # Calculated enthalpy of reaction TbNO3+2
# Enthalpy of formation: -223.8 kcal/mol
-analytic 8.7852e+000 2.4868e-002 2.5553e+003 -9.7944e+000 3.9871e+001
# -Range: 0-300
1.0000 Tb+++ + 1.0000 H2O = TbO+ +2.0000 H+
-llnl_gamma 4.0
log_k -16.1904
-delta_H 109.692 kJ/mol # Calculated enthalpy of reaction TbO+
# Enthalpy of formation: -209 kcal/mol
-analytic 1.7975e+002 2.9563e-002 -1.3407e+004 -6.4573e+001 -2.0926e+002
# -Range: 0-300
2.0000 H2O + 1.0000 Tb+++ = TbO2- +4.0000 H+
-llnl_gamma 4.0
log_k -34.2134
-delta_H 278.797 kJ/mol # Calculated enthalpy of reaction TbO2-
# Enthalpy of formation: -236.9 kcal/mol
-analytic 1.6924e+002 1.1804e-002 -1.9821e+004 -5.6781e+001 -3.0933e+002
# -Range: 0-300
2.0000 H2O + 1.0000 Tb+++ = TbO2H +3.0000 H+
-llnl_gamma 3.0
log_k -25.0508
-delta_H 219.802 kJ/mol # Calculated enthalpy of reaction TbO2H
# Enthalpy of formation: -251 kcal/mol
-analytic 3.2761e+002 4.5225e-002 -2.2652e+004 -1.1727e+002 -3.5356e+002
# -Range: 0-300
1.0000 Tb+++ + 1.0000 H2O = TbOH++ +1.0000 H+
-llnl_gamma 4.5
log_k -7.8342
-delta_H 77.4751 kJ/mol # Calculated enthalpy of reaction TbOH+2
# Enthalpy of formation: -216.7 kcal/mol
-analytic 5.9574e+001 1.1625e-002 -5.8143e+003 -2.0759e+001 -9.0744e+001
# -Range: 0-300
1.0000 Tb+++ + 1.0000 HPO4-- = TbPO4 +1.0000 H+
-llnl_gamma 3.0
log_k +0.0782
-delta_H 0 # Not possible to calculate enthalpy of reaction TbPO4
# Enthalpy of formation: -0 kcal/mol
1.0000 Tb+++ + 1.0000 SO4-- = TbSO4+
-llnl_gamma 4.0
log_k +3.6430
-delta_H 19.6648 kJ/mol # Calculated enthalpy of reaction TbSO4+
# Enthalpy of formation: -379.6 kcal/mol
-analytic 2.9633e+002 8.5155e-002 -8.6346e+003 -1.1682e+002 -1.3482e+002
# -Range: 0-300
2.0000 H2O + 1.0000 TcO++ = TcO(OH)2 +2.0000 H+
-llnl_gamma 3.0
log_k -3.3221
-delta_H 0 # Not possible to calculate enthalpy of reaction TcO(OH)2
# Enthalpy of formation: -0 kcal/mol
1.0000 TcO++ + 1.0000 H2O = TcOOH+ +1.0000 H+
-llnl_gamma 4.0
log_k -1.1355
-delta_H 0 # Not possible to calculate enthalpy of reaction TcOOH+
# Enthalpy of formation: -0 kcal/mol
2.0000 HPO4-- + 2.0000 H+ + 1.0000 Th++++ = Th(H2PO4)2++
-llnl_gamma 4.5
log_k +23.2070
-delta_H 0 # Not possible to calculate enthalpy of reaction Th(H2PO4)2+2
# Enthalpy of formation: -0 kcal/mol
2.0000 HPO4-- + 1.0000 Th++++ = Th(HPO4)2
-llnl_gamma 3.0
log_k +22.6939
-delta_H -13.644 kJ/mol # Calculated enthalpy of reaction Th(HPO4)2
# Enthalpy of formation: -804.691 kcal/mol
-analytic 6.5208e+002 2.3099e-001 -1.2990e+004 -2.6457e+002 -2.2082e+002
# -Range: 0-200
3.0000 HPO4-- + 1.0000 Th++++ = Th(HPO4)3--
-llnl_gamma 4.0
log_k +31.1894
-delta_H 0 # Not possible to calculate enthalpy of reaction Th(HPO4)3-2
# Enthalpy of formation: -0 kcal/mol
2.0000 H2O + 1.0000 Th++++ = Th(OH)2++ +2.0000 H+
-llnl_gamma 4.5
log_k -7.1068
-delta_H 58.668 kJ/mol # Calculated enthalpy of reaction Th(OH)2+2
# Enthalpy of formation: -306.412 kcal/mol
-analytic -1.1274e+001 3.4195e-003 -3.7553e+002 3.1299e+000 -2.9696e+005
# -Range: 0-300
3.0000 H2O + 1.0000 Th++++ = Th(OH)3+ +3.0000 H+
-llnl_gamma 4.0
log_k -11.8623
-delta_H 86.1318 kJ/mol # Calculated enthalpy of reaction Th(OH)3+
# Enthalpy of formation: -368.165 kcal/mol
4.0000 H2O + 1.0000 Th++++ = Th(OH)4 +4.0000 H+
-llnl_gamma 3.0
log_k -16.0315
-delta_H 104.01 kJ/mol # Calculated enthalpy of reaction Th(OH)4
# Enthalpy of formation: -432.209 kcal/mol
-analytic 2.9534e+001 1.5550e-002 -5.6680e+003 -1.2598e+001 -9.6262e+001
# -Range: 0-200
2.0000 SO4-- + 1.0000 Th++++ = Th(SO4)2
-llnl_gamma 3.0
log_k +9.6170
-delta_H 32.2377 kJ/mol # Calculated enthalpy of reaction Th(SO4)2
# Enthalpy of formation: -610.895 kcal/mol
-analytic 4.6425e+002 1.6769e-001 -1.1195e+004 -1.8875e+002 -1.9027e+002
# -Range: 0-200
3.0000 SO4-- + 1.0000 Th++++ = Th(SO4)3--
-llnl_gamma 4.0
log_k +10.4014
-delta_H 0 # Not possible to calculate enthalpy of reaction Th(SO4)3-2
# Enthalpy of formation: -0 kcal/mol
4.0000 SO4-- + 1.0000 Th++++ = Th(SO4)4----
-llnl_gamma 4.0
log_k +8.4003
-delta_H 0 # Not possible to calculate enthalpy of reaction Th(SO4)4-4
# Enthalpy of formation: -0 kcal/mol
2.0000 Th++++ + 2.0000 H2O = Th2(OH)2+6 +2.0000 H+
-llnl_gamma 6.0
log_k -6.4618
-delta_H 63.7181 kJ/mol # Calculated enthalpy of reaction Th2(OH)2+6
# Enthalpy of formation: -489.005 kcal/mol
-analytic 6.8838e+001 -4.1348e-003 -6.4415e+003 -2.1200e+001 -1.0053e+002
# -Range: 0-300
8.0000 H2O + 4.0000 Th++++ = Th4(OH)8+8 +8.0000 H+
-llnl_gamma 6.0
log_k -21.7568
-delta_H 245.245 kJ/mol # Calculated enthalpy of reaction Th4(OH)8+8
# Enthalpy of formation: -1223.12 kcal/mol
-analytic 2.7826e+002 -2.3504e-003 -2.4410e+004 -8.7873e+001 -3.8097e+002
# -Range: 0-300
15.0000 H2O + 6.0000 Th++++ = Th6(OH)15+9 +15.0000 H+
-llnl_gamma 6.0
log_k -37.7027
-delta_H 458.248 kJ/mol # Calculated enthalpy of reaction Th6(OH)15+9
# Enthalpy of formation: -2018.03 kcal/mol
-analytic 5.2516e+002 3.3015e-003 -4.5237e+004 -1.6654e+002 -7.0603e+002
# -Range: 0-300
1.0000 Th++++ + 1.0000 Cl- = ThCl+++
-llnl_gamma 5.0
log_k +0.9536
-delta_H 0.06276 kJ/mol # Calculated enthalpy of reaction ThCl+3
# Enthalpy of formation: -223.718 kcal/mol
-analytic 9.7430e+001 3.9398e-002 -1.8653e+003 -4.1202e+001 -2.9135e+001
# -Range: 0-300
2.0000 Cl- + 1.0000 Th++++ = ThCl2++
-llnl_gamma 4.5
log_k +0.6758
-delta_H 0 # Not possible to calculate enthalpy of reaction ThCl2+2
# Enthalpy of formation: -0 kcal/mol
3.0000 Cl- + 1.0000 Th++++ = ThCl3+
-llnl_gamma 4.0
log_k +1.4975
-delta_H 0 # Not possible to calculate enthalpy of reaction ThCl3+
# Enthalpy of formation: -0 kcal/mol
4.0000 Cl- + 1.0000 Th++++ = ThCl4
-llnl_gamma 3.0
log_k +1.0731
-delta_H 0 # Not possible to calculate enthalpy of reaction ThCl4
# Enthalpy of formation: -0 kcal/mol
1.0000 Th++++ + 1.0000 F- = ThF+++
-llnl_gamma 5.0
log_k +7.8725
-delta_H -4.87436 kJ/mol # Calculated enthalpy of reaction ThF+3
# Enthalpy of formation: -265.115 kcal/mol
-analytic 1.1679e+002 3.9201e-002 -2.2118e+003 -4.5736e+001 -3.4548e+001
# -Range: 0-300
2.0000 F- + 1.0000 Th++++ = ThF2++
-llnl_gamma 4.5
log_k +14.0884
-delta_H -7.77806 kJ/mol # Calculated enthalpy of reaction ThF2+2
# Enthalpy of formation: -345.959 kcal/mol
-analytic 2.3200e+002 7.9567e-002 -4.4418e+003 -9.1617e+001 -6.9379e+001
# -Range: 0-300
3.0000 F- + 1.0000 Th++++ = ThF3+
-llnl_gamma 4.0
log_k +18.7357
-delta_H -11.7068 kJ/mol # Calculated enthalpy of reaction ThF3+
# Enthalpy of formation: -427.048 kcal/mol
-analytic 3.4511e+002 1.2149e-001 -6.5065e+003 -1.3770e+002 -1.0163e+002
# -Range: 0-300
4.0000 F- + 1.0000 Th++++ = ThF4
-llnl_gamma 3.0
log_k +22.1515
-delta_H -14.8448 kJ/mol # Calculated enthalpy of reaction ThF4
# Enthalpy of formation: -507.948 kcal/mol
-analytic 6.1206e+002 2.1878e-001 -1.1938e+004 -2.4857e+002 -2.0294e+002
# -Range: 0-200
1.0000 Th++++ + 1.0000 HPO4-- + 1.0000 H+ = ThH2PO4+++
-llnl_gamma 5.0
log_k +11.7061
-delta_H 0 # Not possible to calculate enthalpy of reaction ThH2PO4+3
# Enthalpy of formation: -0 kcal/mol
2.0000 H+ + 1.0000 Th++++ + 1.0000 HPO4-- = ThH3PO4++++
-llnl_gamma 5.5
log_k +11.1197
-delta_H 0 # Not possible to calculate enthalpy of reaction ThH3PO4+4
# Enthalpy of formation: -0 kcal/mol
1.0000 Th++++ + 1.0000 HPO4-- = ThHPO4++
-llnl_gamma 4.5
log_k +10.6799
-delta_H 0.1046 kJ/mol # Calculated enthalpy of reaction ThHPO4+2
# Enthalpy of formation: -492.59 kcal/mol
1.0000 Th++++ + 1.0000 H2O = ThOH+++ +1.0000 H+
-llnl_gamma 5.0
log_k -3.8871
-delta_H 25.0275 kJ/mol # Calculated enthalpy of reaction ThOH+3
# Enthalpy of formation: -1029.83 kJ/mol
-analytic 1.0495e+001 5.1532e-003 -8.6396e+002 -4.8420e+000 -9.2609e+004
# -Range: 0-300
1.0000 Th++++ + 1.0000 SO4-- = ThSO4++
-llnl_gamma 4.5
log_k +5.3143
-delta_H 16.3511 kJ/mol # Calculated enthalpy of reaction ThSO4+2
# Enthalpy of formation: -397.292 kcal/mol
-analytic 1.9443e+002 7.5245e-002 -4.5010e+003 -7.9379e+001 -7.0291e+001
# -Range: 0-300
2.0000 HAcetate + 1.0000 Tl+ = Tl(Acetate)2- +2.0000 H+
-llnl_gamma 4.0
log_k -10.0129
-delta_H 1.2552 kJ/mol # Calculated enthalpy of reaction Tl(Acetate)2-
# Enthalpy of formation: -230.62 kcal/mol
-analytic -1.8123e+002 -4.0616e-002 5.0741e+003 6.7216e+001 7.9229e+001
# -Range: 0-300
1.0000 Tl+ + 1.0000 HAcetate = TlAcetate +1.0000 H+
-llnl_gamma 3.0
log_k -4.8672
-delta_H 6.15048 kJ/mol # Calculated enthalpy of reaction TlAcetate
# Enthalpy of formation: -113.35 kcal/mol
-analytic 9.2977e+000 -3.4368e-003 -2.1748e+003 -3.1454e+000 1.7273e+005
# -Range: 0-300
2.0000 HAcetate + 1.0000 Tm+++ = Tm(Acetate)2+ +2.0000 H+
-llnl_gamma 4.0
log_k -4.9844
-delta_H -32.5934 kJ/mol # Calculated enthalpy of reaction Tm(Acetate)2+
# Enthalpy of formation: -408.49 kcal/mol
-analytic -2.8983e+001 2.0256e-003 -1.1525e+003 8.2163e+000 6.1820e+005
# -Range: 0-300
3.0000 HAcetate + 1.0000 Tm+++ = Tm(Acetate)3 +3.0000 H+
-llnl_gamma 3.0
log_k -8.3783
-delta_H -54.8104 kJ/mol # Calculated enthalpy of reaction Tm(Acetate)3
# Enthalpy of formation: -529.9 kcal/mol
-analytic -2.8900e+001 4.9633e-003 -1.6574e+003 6.0186e+000 8.6624e+005
# -Range: 0-300
2.0000 HCO3- + 1.0000 Tm+++ = Tm(CO3)2- +2.0000 H+
-llnl_gamma 4.0
log_k -7.1576
-delta_H 0 # Not possible to calculate enthalpy of reaction Tm(CO3)2-
# Enthalpy of formation: -0 kcal/mol
2.0000 HPO4-- + 1.0000 Tm+++ = Tm(HPO4)2-
-llnl_gamma 4.0
log_k +10.1000
-delta_H 0 # Not possible to calculate enthalpy of reaction Tm(HPO4)2-
# Enthalpy of formation: -0 kcal/mol
2.0000 HPO4-- + 1.0000 Tm+++ = Tm(PO4)2--- +2.0000 H+
-llnl_gamma 4.0
log_k -3.0437
-delta_H 0 # Not possible to calculate enthalpy of reaction Tm(PO4)2-3
# Enthalpy of formation: -0 kcal/mol
2.0000 SO4-- + 1.0000 Tm+++ = Tm(SO4)2-
-llnl_gamma 4.0
log_k +5.1000
-delta_H 0 # Not possible to calculate enthalpy of reaction Tm(SO4)2-
# Enthalpy of formation: -0 kcal/mol
1.0000 Tm+++ + 1.0000 HAcetate = TmAcetate++ +1.0000 H+
-llnl_gamma 4.5
log_k -2.1184
-delta_H -16.3176 kJ/mol # Calculated enthalpy of reaction TmAcetate+2
# Enthalpy of formation: -288.5 kcal/mol
-analytic -1.6068e+001 1.2043e-003 -6.2777e+002 4.8318e+000 3.3363e+005
# -Range: 0-300
1.0000 Tm+++ + 1.0000 HCO3- = TmCO3+ +1.0000 H+
-llnl_gamma 4.0
log_k -2.1125
-delta_H 86.6004 kJ/mol # Calculated enthalpy of reaction TmCO3+
# Enthalpy of formation: -312.7 kcal/mol
-analytic 2.3889e+002 5.4733e-002 -6.9382e+003 -9.4581e+001 -1.0833e+002
# -Range: 0-300
1.0000 Tm+++ + 1.0000 Cl- = TmCl++
-llnl_gamma 4.5
log_k +0.2353
-delta_H 13.1085 kJ/mol # Calculated enthalpy of reaction TmCl+2
# Enthalpy of formation: -205.3 kcal/mol
-analytic 7.4795e+001 3.7655e-002 -1.5701e+003 -3.2531e+001 -2.4523e+001
# -Range: 0-300
2.0000 Cl- + 1.0000 Tm+++ = TmCl2+
-llnl_gamma 4.0
log_k -0.0425
-delta_H 15.7569 kJ/mol # Calculated enthalpy of reaction TmCl2+
# Enthalpy of formation: -244.6 kcal/mol
-analytic 2.0352e+002 7.9173e-002 -4.8574e+003 -8.5202e+001 -7.5855e+001
# -Range: 0-300
3.0000 Cl- + 1.0000 Tm+++ = TmCl3
-llnl_gamma 3.0
log_k -0.4669
-delta_H 5.43502 kJ/mol # Calculated enthalpy of reaction TmCl3
# Enthalpy of formation: -287 kcal/mol
-analytic 3.9793e+002 1.2777e-001 -1.0070e+004 -1.6272e+002 -1.5725e+002
# -Range: 0-300
4.0000 Cl- + 1.0000 Tm+++ = TmCl4-
-llnl_gamma 4.0
log_k -0.8913
-delta_H -20.3677 kJ/mol # Calculated enthalpy of reaction TmCl4-
# Enthalpy of formation: -333.1 kcal/mol
-analytic 4.3574e+002 1.2655e-001 -1.0713e+004 -1.7716e+002 -1.6730e+002
# -Range: 0-300
1.0000 Tm+++ + 1.0000 F- = TmF++
-llnl_gamma 4.5
log_k +4.8085
-delta_H 23.6396 kJ/mol # Calculated enthalpy of reaction TmF+2
# Enthalpy of formation: -243 kcal/mol
-analytic 9.7686e+001 4.1890e-002 -2.5909e+003 -3.9059e+001 -4.0457e+001
# -Range: 0-300
2.0000 F- + 1.0000 Tm+++ = TmF2+
-llnl_gamma 4.0
log_k +8.3709
-delta_H 12.552 kJ/mol # Calculated enthalpy of reaction TmF2+
# Enthalpy of formation: -325.8 kcal/mol
-analytic 2.2986e+002 8.4119e-002 -5.2144e+003 -9.2558e+001 -8.1433e+001
# -Range: 0-300
3.0000 F- + 1.0000 Tm+++ = TmF3
-llnl_gamma 3.0
log_k +10.9804
-delta_H -12.7612 kJ/mol # Calculated enthalpy of reaction TmF3
# Enthalpy of formation: -412 kcal/mol
-analytic 4.2855e+002 1.3445e-001 -9.7045e+003 -1.7177e+002 -1.5156e+002
# -Range: 0-300
4.0000 F- + 1.0000 Tm+++ = TmF4-
-llnl_gamma 4.0
log_k +13.1501
-delta_H -60.668 kJ/mol # Calculated enthalpy of reaction TmF4-
# Enthalpy of formation: -503.6 kcal/mol
-analytic 4.6559e+002 1.3386e-001 -9.1790e+003 -1.8650e+002 -1.4337e+002
# -Range: 0-300
1.0000 Tm+++ + 1.0000 HPO4-- + 1.0000 H+ = TmH2PO4++
-llnl_gamma 4.5
log_k +9.4484
-delta_H -20.4388 kJ/mol # Calculated enthalpy of reaction TmH2PO4+2
# Enthalpy of formation: -482.2 kcal/mol
-analytic 1.0360e+002 6.3085e-002 6.0731e+002 -4.6456e+001 9.4456e+000
# -Range: 0-300
1.0000 Tm+++ + 1.0000 HCO3- = TmHCO3++
-llnl_gamma 4.5
log_k +1.7724
-delta_H 5.01243 kJ/mol # Calculated enthalpy of reaction TmHCO3+2
# Enthalpy of formation: -332.2 kcal/mol
-analytic 3.3102e+001 3.1010e-002 2.9880e+002 -1.6791e+001 4.6524e+000
# -Range: 0-300
1.0000 Tm+++ + 1.0000 HPO4-- = TmHPO4+
-llnl_gamma 4.0
log_k +5.9000
-delta_H 0 # Not possible to calculate enthalpy of reaction TmHPO4+
# Enthalpy of formation: -0 kcal/mol
1.0000 Tm+++ + 1.0000 NO3- = TmNO3++
-llnl_gamma 4.5
log_k +0.2148
-delta_H -33.7691 kJ/mol # Calculated enthalpy of reaction TmNO3+2
# Enthalpy of formation: -226 kcal/mol
-analytic 1.1085e+001 2.4898e-002 2.5664e+003 -1.0861e+001 4.0043e+001
# -Range: 0-300
1.0000 Tm+++ + 1.0000 H2O = TmO+ +2.0000 H+
-llnl_gamma 4.0
log_k -15.8972
-delta_H 105.508 kJ/mol # Calculated enthalpy of reaction TmO+
# Enthalpy of formation: -211.6 kcal/mol
-analytic 1.7572e+002 2.8756e-002 -1.3096e+004 -6.3150e+001 -2.0441e+002
# -Range: 0-300
2.0000 H2O + 1.0000 Tm+++ = TmO2- +4.0000 H+
-llnl_gamma 4.0
log_k -32.6741
-delta_H 266.663 kJ/mol # Calculated enthalpy of reaction TmO2-
# Enthalpy of formation: -241.4 kcal/mol
-analytic 3.3118e+001 -5.2802e-003 -1.1318e+004 -8.4764e+000 -4.6998e+005
# -Range: 0-300
2.0000 H2O + 1.0000 Tm+++ = TmO2H +3.0000 H+
-llnl_gamma 3.0
log_k -24.1712
-delta_H 211.853 kJ/mol # Calculated enthalpy of reaction TmO2H
# Enthalpy of formation: -254.5 kcal/mol
-analytic 3.1648e+002 4.4527e-002 -2.1821e+004 -1.1345e+002 -3.4059e+002
# -Range: 0-300
1.0000 Tm+++ + 1.0000 H2O = TmOH++ +1.0000 H+
-llnl_gamma 4.5
log_k -7.6876
-delta_H 74.5463 kJ/mol # Calculated enthalpy of reaction TmOH+2
# Enthalpy of formation: -219 kcal/mol
-analytic 5.7572e+001 1.1162e-002 -5.6381e+003 -2.0074e+001 -8.7994e+001
# -Range: 0-300
1.0000 Tm+++ + 1.0000 HPO4-- = TmPO4 +1.0000 H+
-llnl_gamma 3.0
log_k +0.4782
-delta_H 0 # Not possible to calculate enthalpy of reaction TmPO4
# Enthalpy of formation: -0 kcal/mol
1.0000 Tm+++ + 1.0000 SO4-- = TmSO4+
-llnl_gamma 4.0
log_k +3.5697
-delta_H 19.9995 kJ/mol # Calculated enthalpy of reaction TmSO4+
# Enthalpy of formation: -381.12 kcal/mol
-analytic 3.0441e+002 8.6070e-002 -8.9592e+003 -1.1979e+002 -1.3989e+002
# -Range: 0-300
4.0000 HCO3- + 1.0000 U++++ = U(CO3)4---- +4.0000 H+
-llnl_gamma 4.0
log_k -6.2534
-delta_H 0 # Not possible to calculate enthalpy of reaction U(CO3)4-4
# Enthalpy of formation: -0 kcal/mol
5.0000 HCO3- + 1.0000 U++++ = U(CO3)5-6 +5.0000 H+
-llnl_gamma 4.0
log_k -17.7169
-delta_H 53.5172 kJ/mol # Calculated enthalpy of reaction U(CO3)5-6
# Enthalpy of formation: -3987.35 kJ/mol
-analytic 6.3020e+002 1.9391e-001 -1.9238e+004 -2.5912e+002 -3.0038e+002
# -Range: 0-300
2.0000 NO3- + 1.0000 U++++ = U(NO3)2++
-llnl_gamma 4.5
log_k +2.2610
-delta_H 0 # Not possible to calculate enthalpy of reaction U(NO3)2+2
# Enthalpy of formation: -0 kcal/mol
4.0000 H2O + 1.0000 U++++ = U(OH)4 +4.0000 H+
-llnl_gamma 3.0
log_k -4.57
-delta_H 78.7553 kJ/mol # Calculated enthalpy of reaction U(OH)4
# Enthalpy of formation: -1655.8 kJ/mol
-analytic 2.6685e+002 9.8204e-002 -9.4428e+003 -1.0871e+002 -1.6045e+002
# -Range: 0-200
2.0000 Thiocyanate- + 1.0000 U++++ = U(Thiocyanate)2++
-llnl_gamma 4.5
log_k +4.2600
-delta_H 0 # Not possible to calculate enthalpy of reaction U(Thiocyanate)2+2
# Enthalpy of formation: -456.4 kJ/mol
-analytic 6.2193e+000 2.7673e-002 2.4326e+003 -7.4158e+000 3.7957e+001
# -Range: 0-300
2.0000 SO4-- + 1.0000 U++++ = U(SO4)2
-llnl_gamma 3.0
log_k +10.3507
-delta_H 33.2232 kJ/mol # Calculated enthalpy of reaction U(SO4)2
# Enthalpy of formation: -2377.18 kJ/mol
-analytic 4.9476e+002 1.7832e-001 -1.1901e+004 -2.0111e+002 -2.0227e+002
# -Range: 0-200
1.0000 U++++ + 1.0000 Br- = UBr+++
-llnl_gamma 5.0
log_k +1.4240
-delta_H 0 # Not possible to calculate enthalpy of reaction UBr+3
# Enthalpy of formation: -0 kcal/mol
1.0000 U++++ + 1.0000 Cl- = UCl+++
-llnl_gamma 5.0
log_k +1.7073
-delta_H -18.9993 kJ/mol # Calculated enthalpy of reaction UCl+3
# Enthalpy of formation: -777.279 kJ/mol
-analytic 9.4418e+001 4.1718e-002 -7.0675e+002 -4.1532e+001 -1.1056e+001
# -Range: 0-300
1.0000 U++++ + 1.0000 F- = UF+++
-llnl_gamma 5.0
log_k +9.2403
-delta_H -5.6024 kJ/mol # Calculated enthalpy of reaction UF+3
# Enthalpy of formation: -932.15 kJ/mol
-analytic 1.1828e+002 3.8097e-002 -2.2531e+003 -4.5594e+001 -3.5193e+001
# -Range: 0-300
2.0000 F- + 1.0000 U++++ = UF2++
-llnl_gamma 4.5
log_k +16.1505
-delta_H -3.5048 kJ/mol # Calculated enthalpy of reaction UF2+2
# Enthalpy of formation: -1265.4 kJ/mol
-analytic 2.3537e+002 7.7064e-002 -4.8455e+003 -9.1296e+001 -7.5679e+001
# -Range: 0-300
3.0000 F- + 1.0000 U++++ = UF3+
-llnl_gamma 4.0
log_k +21.4806
-delta_H 0.4938 kJ/mol # Calculated enthalpy of reaction UF3+
# Enthalpy of formation: -1596.75 kJ/mol
-analytic 3.5097e+002 1.1714e-001 -7.4569e+003 -1.3714e+002 -1.1646e+002
# -Range: 0-300
4.0000 F- + 1.0000 U++++ = UF4
-llnl_gamma 3.0
log_k +25.4408
-delta_H -4.2146 kJ/mol # Calculated enthalpy of reaction UF4
# Enthalpy of formation: -1936.81 kJ/mol
-analytic 7.8549e+002 2.7922e-001 -1.6213e+004 -3.1881e+002 -2.7559e+002
# -Range: 0-200
5.0000 F- + 1.0000 U++++ = UF5-
-llnl_gamma 4.0
log_k +26.8110
-delta_H 0 # Not possible to calculate enthalpy of reaction UF5-
# Enthalpy of formation: -0 kcal/mol
6.0000 F- + 1.0000 U++++ = UF6--
-llnl_gamma 4.0
log_k +28.8412
-delta_H 0 # Not possible to calculate enthalpy of reaction UF6-2
# Enthalpy of formation: -0 kcal/mol
1.0000 U++++ + 1.0000 I- = UI+++
-llnl_gamma 5.0
log_k +1.2151
-delta_H 0 # Not possible to calculate enthalpy of reaction UI+3
# Enthalpy of formation: -0 kcal/mol
1.0000 U++++ + 1.0000 NO3- = UNO3+++
-llnl_gamma 5.0
log_k +1.4506
-delta_H 0 # Not possible to calculate enthalpy of reaction UNO3+3
# Enthalpy of formation: -0 kcal/mol
2.0000 HCO3- + 1.0000 UO2++ = UO2(CO3)2-- +2.0000 H+
-llnl_gamma 4.0
log_k -3.7467
-delta_H 47.9065 kJ/mol # Calculated enthalpy of reaction UO2(CO3)2-2
# Enthalpy of formation: -2350.96 kJ/mol
-analytic 2.6569e+002 8.1552e-002 -9.0918e+003 -1.0638e+002 -1.4195e+002
# -Range: 0-300
3.0000 HCO3- + 1.0000 UO2+ = UO2(CO3)3-5 +3.0000 H+
-llnl_gamma 4.0
log_k -23.6241
-delta_H 0 # Not possible to calculate enthalpy of reaction UO2(CO3)3-5
# Enthalpy of formation: -0 kcal/mol
3.0000 HCO3- + 1.0000 UO2++ = UO2(CO3)3---- +3.0000 H+
-llnl_gamma 4.0
log_k -9.4302
-delta_H 4.9107 kJ/mol # Calculated enthalpy of reaction UO2(CO3)3-4
# Enthalpy of formation: -3083.89 kJ/mol
-analytic 3.7918e+002 1.1789e-001 -1.0233e+004 -1.5738e+002 -1.5978e+002
# -Range: 0-300
3.0000 H+ + 2.0000 HPO4-- + 1.0000 UO2++ = UO2(H2PO4)(H3PO4)+
-llnl_gamma 4.0
log_k +22.7537
-delta_H 0 # Not possible to calculate enthalpy of reaction UO2(H2PO4)(H3PO4)+
# Enthalpy of formation: -0 kcal/mol
2.0000 HPO4-- + 2.0000 H+ + 1.0000 UO2++ = UO2(H2PO4)2
-llnl_gamma 3.0
log_k +21.7437
-delta_H 0 # Not possible to calculate enthalpy of reaction UO2(H2PO4)2
# Enthalpy of formation: -0 kcal/mol
2.0000 IO3- + 1.0000 UO2++ = UO2(IO3)2
-llnl_gamma 3.0
log_k +2.9969
-delta_H 0 # Not possible to calculate enthalpy of reaction UO2(IO3)2
# Enthalpy of formation: -0 kcal/mol
2.0000 N3- + 1.0000 UO2++ = UO2(N3)2
-llnl_gamma 3.0
log_k +4.3301
-delta_H 0 # Not possible to calculate enthalpy of reaction UO2(N3)2
# Enthalpy of formation: -0 kcal/mol
3.0000 N3- + 1.0000 UO2++ = UO2(N3)3-
-llnl_gamma 4.0
log_k +5.7401
-delta_H 0 # Not possible to calculate enthalpy of reaction UO2(N3)3-
# Enthalpy of formation: -0 kcal/mol
4.0000 N3- + 1.0000 UO2++ = UO2(N3)4--
-llnl_gamma 4.0
log_k +4.9200
-delta_H 0 # Not possible to calculate enthalpy of reaction UO2(N3)4-2
# Enthalpy of formation: -0 kcal/mol
2.0000 H2O + 1.0000 UO2++ = UO2(OH)2 +2.0000 H+
-llnl_gamma 3.0
log_k -10.3146
-delta_H 0 # Not possible to calculate enthalpy of reaction UO2(OH)2
# Enthalpy of formation: -0 kcal/mol
3.0000 H2O + 1.0000 UO2++ = UO2(OH)3- +3.0000 H+
-llnl_gamma 4.0
log_k -19.2218
-delta_H 0 # Not possible to calculate enthalpy of reaction UO2(OH)3-
# Enthalpy of formation: -0 kcal/mol
4.0000 H2O + 1.0000 UO2++ = UO2(OH)4-- +4.0000 H+
-llnl_gamma 4.0
log_k -33.0291
-delta_H 0 # Not possible to calculate enthalpy of reaction UO2(OH)4-2
# Enthalpy of formation: -0 kcal/mol
2.0000 Thiocyanate- + 1.0000 UO2++ = UO2(Thiocyanate)2
-llnl_gamma 3.0
log_k +1.2401
-delta_H 0 # Not possible to calculate enthalpy of reaction UO2(Thiocyanate)2
# Enthalpy of formation: -857.3 kJ/mol
-analytic 9.4216e+001 3.2840e-002 -2.4849e+003 -3.8162e+001 -4.2231e+001
# -Range: 0-200
3.0000 Thiocyanate- + 1.0000 UO2++ = UO2(Thiocyanate)3-
-llnl_gamma 4.0
log_k +2.1001
-delta_H 0 # Not possible to calculate enthalpy of reaction UO2(Thiocyanate)3-
# Enthalpy of formation: -783.8 kJ/mol
-analytic 1.6622e+001 2.2714e-002 4.9707e+002 -9.2785e+000 7.7512e+000
# -Range: 0-300
2.0000 SO3-- + 1.0000 UO2++ = UO2(SO3)2--
-llnl_gamma 4.0
log_k +7.9101
-delta_H 0 # Not possible to calculate enthalpy of reaction UO2(SO3)2-2
# Enthalpy of formation: -0 kcal/mol
2.0000 SO4-- + 1.0000 UO2++ = UO2(SO4)2--
-llnl_gamma 4.0
log_k +3.9806
-delta_H 35.6242 kJ/mol # Calculated enthalpy of reaction UO2(SO4)2-2
# Enthalpy of formation: -2802.58 kJ/mol
-analytic 3.9907e+002 1.3536e-001 -1.0813e+004 -1.6130e+002 -1.6884e+002
# -Range: 0-300
1.0000 UO2++ + 1.0000 Br- = UO2Br+
-llnl_gamma 4.0
log_k +0.1840
-delta_H 0 # Not possible to calculate enthalpy of reaction UO2Br+
# Enthalpy of formation: -0 kcal/mol
1.0000 UO2++ + 1.0000 BrO3- = UO2BrO3+
-llnl_gamma 4.0
log_k +0.5510
-delta_H 0.46952 kJ/mol # Calculated enthalpy of reaction UO2BrO3+
# Enthalpy of formation: -1085.6 kJ/mol
-analytic 8.2618e+001 2.6921e-002 -2.0144e+003 -3.3673e+001 -3.1457e+001
# -Range: 0-300
1.0000 UO2++ + 1.0000 HCO3- = UO2CO3 +1.0000 H+
-llnl_gamma 3.0
log_k -0.6634
-delta_H 19.7032 kJ/mol # Calculated enthalpy of reaction UO2CO3
# Enthalpy of formation: -1689.23 kJ/mol
-analytic 7.3898e+001 2.8127e-002 -2.4347e+003 -3.0217e+001 -4.1371e+001
# -Range: 0-200
1.0000 UO2++ + 1.0000 Cl- = UO2Cl+
-llnl_gamma 4.0
log_k +0.1572
-delta_H 8.00167 kJ/mol # Calculated enthalpy of reaction UO2Cl+
# Enthalpy of formation: -1178.08 kJ/mol
-analytic 9.8139e+001 3.8869e-002 -2.3178e+003 -4.1133e+001 -3.6196e+001
# -Range: 0-300
2.0000 Cl- + 1.0000 UO2++ = UO2Cl2
-llnl_gamma 3.0
log_k -1.1253
-delta_H 15.0013 kJ/mol # Calculated enthalpy of reaction UO2Cl2
# Enthalpy of formation: -1338.16 kJ/mol
-analytic 3.4087e+001 1.3840e-002 -1.3664e+003 -1.4043e+001 -2.3216e+001
# -Range: 0-200
1.0000 UO2++ + 1.0000 ClO3- = UO2ClO3+
-llnl_gamma 4.0
log_k +0.4919
-delta_H -3.9266 kJ/mol # Calculated enthalpy of reaction UO2ClO3+
# Enthalpy of formation: -1126.9 kJ/mol
-analytic 9.6263e+001 2.8926e-002 -2.3068e+003 -3.9057e+001 -3.6025e+001
# -Range: 0-300
1.0000 UO2++ + 1.0000 F- = UO2F+
-llnl_gamma 4.0
log_k +5.0502
-delta_H 1.6976 kJ/mol # Calculated enthalpy of reaction UO2F+
# Enthalpy of formation: -1352.65 kJ/mol
-analytic 1.1476e+002 4.0682e-002 -2.4467e+003 -4.5914e+001 -3.8212e+001
# -Range: 0-300
2.0000 F- + 1.0000 UO2++ = UO2F2
-llnl_gamma 3.0
log_k +8.5403
-delta_H 2.0962 kJ/mol # Calculated enthalpy of reaction UO2F2
# Enthalpy of formation: -1687.6 kJ/mol
-analytic 2.7673e+002 9.9190e-002 -5.8371e+003 -1.1242e+002 -9.9219e+001
# -Range: 0-200
3.0000 F- + 1.0000 UO2++ = UO2F3-
-llnl_gamma 4.0
log_k +10.7806
-delta_H 2.3428 kJ/mol # Calculated enthalpy of reaction UO2F3-
# Enthalpy of formation: -2022.7 kJ/mol
-analytic 3.3383e+002 9.2160e-002 -8.7975e+003 -1.2972e+002 -1.3738e+002
# -Range: 0-300
4.0000 F- + 1.0000 UO2++ = UO2F4--
-llnl_gamma 4.0
log_k +11.5407
-delta_H 0.2814 kJ/mol # Calculated enthalpy of reaction UO2F4-2
# Enthalpy of formation: -2360.11 kJ/mol
-analytic 4.4324e+002 1.3808e-001 -1.0705e+004 -1.7657e+002 -1.6718e+002
# -Range: 0-300
1.0000 UO2++ + 1.0000 HPO4-- + 1.0000 H+ = UO2H2PO4+
-llnl_gamma 4.0
log_k +11.6719
-delta_H 0 # Not possible to calculate enthalpy of reaction UO2H2PO4+
# Enthalpy of formation: -0 kcal/mol
2.0000 H+ + 1.0000 UO2++ + 1.0000 HPO4-- = UO2H3PO4++
-llnl_gamma 4.5
log_k +11.3119
-delta_H 0 # Not possible to calculate enthalpy of reaction UO2H3PO4+2
# Enthalpy of formation: -0 kcal/mol
1.0000 UO2++ + 1.0000 HPO4-- = UO2HPO4
-llnl_gamma 3.0
log_k +8.4398
-delta_H 0 # Not possible to calculate enthalpy of reaction UO2HPO4
# Enthalpy of formation: -0 kcal/mol
1.0000 UO2++ + 1.0000 IO3- = UO2IO3+
-llnl_gamma 4.0
log_k +1.7036
-delta_H 11.4336 kJ/mol # Calculated enthalpy of reaction UO2IO3+
# Enthalpy of formation: -1228.9 kJ/mol
-analytic 1.0428e+002 2.9620e-002 -3.2441e+003 -4.0618e+001 -5.0651e+001
# -Range: 0-300
1.0000 UO2++ + 1.0000 N3- = UO2N3+
-llnl_gamma 4.0
log_k +2.5799
-delta_H 0 # Not possible to calculate enthalpy of reaction UO2N3+
# Enthalpy of formation: -0 kcal/mol
1.0000 UO2++ + 1.0000 NO3- = UO2NO3+
-llnl_gamma 4.0
log_k +0.2805
-delta_H 0 # Not possible to calculate enthalpy of reaction UO2NO3+
# Enthalpy of formation: -0 kcal/mol
1.0000 UO2++ + 1.0000 H2O = UO2OH+ +1.0000 H+
-llnl_gamma 4.0
log_k -5.2073
-delta_H 43.1813 kJ/mol # Calculated enthalpy of reaction UO2OH+
# Enthalpy of formation: -1261.66 kJ/mol
-analytic 3.4387e+001 6.0811e-003 -3.3068e+003 -1.2252e+001 -5.1609e+001
# -Range: 0-300
1.0000 UO2++ + 1.0000 HPO4-- = UO2PO4- +1.0000 H+
-llnl_gamma 4.0
log_k +2.0798
-delta_H 0 # Not possible to calculate enthalpy of reaction UO2PO4-
# Enthalpy of formation: -0 kcal/mol
#2.0000 SO3-- + 2.0000 H+ + 1.0000 UO2++ = UO2S2O3 +1.0000 H2O +1.0000 O2
#S2O3-- + O2 + H2O = 2.0000 H+ + 2.0000 SO3-- log_k 40.2906
S2O3-- + UO2++ = UO2S2O3
-llnl_gamma 3.0
# log_k -38.0666
log_k 2.224
-delta_H 0 # Not possible to calculate enthalpy of reaction UO2S2O3
# Enthalpy of formation: -0 kcal/mol
1.0000 UO2++ + 1.0000 Thiocyanate- = UO2Thiocyanate+
-llnl_gamma 4.0
log_k +1.4000
-delta_H 0 # Not possible to calculate enthalpy of reaction UO2Thiocyanate+
# Enthalpy of formation: -939.38 kJ/mol
-analytic 4.7033e+000 1.2562e-002 4.9095e+002 -3.5097e+000 7.6593e+000
# -Range: 0-300
1.0000 UO2++ + 1.0000 SO3-- = UO2SO3
-llnl_gamma 3.0
log_k +6.7532
-delta_H 0 # Not possible to calculate enthalpy of reaction UO2SO3
# Enthalpy of formation: -0 kcal/mol
1.0000 UO2++ + 1.0000 SO4-- = UO2SO4
-llnl_gamma 3.0
log_k +3.0703
-delta_H 19.7626 kJ/mol # Calculated enthalpy of reaction UO2SO4
# Enthalpy of formation: -1908.84 kJ/mol
-analytic 1.9514e+002 7.0951e-002 -4.9949e+003 -7.9394e+001 -8.4888e+001
# -Range: 0-200
1.0000 U++++ + 1.0000 H2O = UOH+++ +1.0000 H+
-llnl_gamma 5.0
log_k -0.5472
-delta_H 46.9183 kJ/mol # Calculated enthalpy of reaction UOH+3
# Enthalpy of formation: -830.12 kJ/mol
-analytic 4.0793e+001 1.3563e-003 -3.8441e+003 -1.1659e+001 -5.9996e+001
# -Range: 0-300
1.0000 U++++ + 1.0000 Thiocyanate- = UThiocyanate+++
-llnl_gamma 5.0
log_k +2.9700
-delta_H 0 # Not possible to calculate enthalpy of reaction UThiocyanate+3
# Enthalpy of formation: -541.8 kJ/mol
-analytic 4.0286e-001 1.5909e-002 2.3026e+003 -3.9973e+000 3.5929e+001
# -Range: 0-300
1.0000 U++++ + 1.0000 SO4-- = USO4++
-llnl_gamma 4.5
log_k +6.5003
-delta_H 8.2616 kJ/mol # Calculated enthalpy of reaction USO4+2
# Enthalpy of formation: -1492.54 kJ/mol
-analytic 1.9418e+002 7.5458e-002 -4.0646e+003 -7.9416e+001 -6.3482e+001
# -Range: 0-300
2.0000 H2O + 1.0000 V+++ = V(OH)2+ +2.0000 H+
-llnl_gamma 4.0
log_k -5.9193
-delta_H 0 # Not possible to calculate enthalpy of reaction V(OH)2+
# Enthalpy of formation: -0 kcal/mol
2.0000 V+++ + 2.0000 H2O = V2(OH)2++++ +2.0000 H+
-llnl_gamma 5.5
log_k -3.8
-delta_H 0 # Not possible to calculate enthalpy of reaction V2(OH)2+4
# Enthalpy of formation: -0 kcal/mol
2.0000 H2O + 1.0000 VO2+ = VO(OH)3 +1.0000 H+
-llnl_gamma 3.0
log_k -3.3
-delta_H 0 # Not possible to calculate enthalpy of reaction VO(OH)3
# Enthalpy of formation: -0 kcal/mol
2.0000 HPO4-- + 1.0000 VO2+ = VO2(HPO4)2---
-llnl_gamma 4.0
log_k +8.6000
-delta_H 0 # Not possible to calculate enthalpy of reaction VO2(HPO4)2-3
# Enthalpy of formation: -0 kcal/mol
2.0000 H2O + 1.0000 VO2+ = VO2(OH)2- +2.0000 H+
-llnl_gamma 4.0
log_k -7.3
-delta_H 0 # Not possible to calculate enthalpy of reaction VO2(OH)2-
# Enthalpy of formation: -0 kcal/mol
1.0000 VO2+ + 1.0000 F- = VO2F
-llnl_gamma 3.0
log_k +3.3500
-delta_H 0 # Not possible to calculate enthalpy of reaction VO2F
# Enthalpy of formation: -0 kcal/mol
2.0000 F- + 1.0000 VO2+ = VO2F2-
-llnl_gamma 4.0
log_k +5.8100
-delta_H 0 # Not possible to calculate enthalpy of reaction VO2F2-
# Enthalpy of formation: -0 kcal/mol
1.0000 VO2+ + 1.0000 HPO4-- + 1.0000 H+ = VO2H2PO4
-llnl_gamma 3.0
log_k +1.6800
-delta_H 0 # Not possible to calculate enthalpy of reaction VO2H2PO4
# Enthalpy of formation: -0 kcal/mol
1.0000 VO2+ + 1.0000 HPO4-- = VO2HPO4-
-llnl_gamma 4.0
log_k +5.8300
-delta_H 0 # Not possible to calculate enthalpy of reaction VO2HPO4-
# Enthalpy of formation: -0 kcal/mol
1.0000 VO2+ + 1.0000 SO4-- = VO2SO4-
-llnl_gamma 4.0
log_k +1.5800
-delta_H 0 # Not possible to calculate enthalpy of reaction VO2SO4-
# Enthalpy of formation: -0 kcal/mol
1.0000 VO4--- + 1.0000 H+ = VO3OH--
-llnl_gamma 4.0
log_k +14.2600
-delta_H 0 # Not possible to calculate enthalpy of reaction VO3OH-2
# Enthalpy of formation: -0 kcal/mol
1.0000 VO++ + 1.0000 F- = VOF+
-llnl_gamma 4.0
log_k +4.0000
-delta_H 0 # Not possible to calculate enthalpy of reaction VOF+
# Enthalpy of formation: -0 kcal/mol
2.0000 F- + 1.0000 VO++ = VOF2
-llnl_gamma 3.0
log_k +6.7800
-delta_H 0 # Not possible to calculate enthalpy of reaction VOF2
# Enthalpy of formation: -0 kcal/mol
1.0000 V+++ + 1.0000 H2O = VOH++ +1.0000 H+
-llnl_gamma 4.5
log_k -2.26
-delta_H 0 # Not possible to calculate enthalpy of reaction VOH+2
# Enthalpy of formation: -0 kcal/mol
1.0000 VO++ + 1.0000 H2O = VOOH+ +1.0000 H+
-llnl_gamma 4.0
log_k -5.67
-delta_H 0 # Not possible to calculate enthalpy of reaction VOOH+
# Enthalpy of formation: -0 kcal/mol
1.0000 VO++ + 1.0000 SO4-- = VOSO4
-llnl_gamma 3.0
log_k +2.4800
-delta_H 0 # Not possible to calculate enthalpy of reaction VOSO4
# Enthalpy of formation: -0 kcal/mol
1.0000 V+++ + 1.0000 SO4-- = VSO4+
-llnl_gamma 4.0
log_k +3.3300
-delta_H 0 # Not possible to calculate enthalpy of reaction VSO4+
# Enthalpy of formation: -0 kcal/mol
2.0000 HAcetate + 1.0000 Y+++ = Y(Acetate)2+ +2.0000 H+
-llnl_gamma 4.0
log_k -4.9844
-delta_H -34.8109 kJ/mol # Calculated enthalpy of reaction Y(Acetate)2+
# Enthalpy of formation: -411.42 kcal/mol
-analytic -3.3011e+001 6.1979e-004 -7.7468e+002 9.6380e+000 5.8814e+005
# -Range: 0-300
3.0000 HAcetate + 1.0000 Y+++ = Y(Acetate)3 +3.0000 H+
-llnl_gamma 3.0
log_k -8.3783
-delta_H -58.4505 kJ/mol # Calculated enthalpy of reaction Y(Acetate)3
# Enthalpy of formation: -533.17 kcal/mol
-analytic -3.0086e+001 4.0213e-003 -1.1444e+003 6.1794e+000 8.0827e+005
# -Range: 0-300
2.0000 HCO3- + 1.0000 Y+++ = Y(CO3)2- +2.0000 H+
-llnl_gamma 4.0
log_k -7.3576
-delta_H 0 # Not possible to calculate enthalpy of reaction Y(CO3)2-
# Enthalpy of formation: -0 kcal/mol
2.0000 HPO4-- + 1.0000 Y+++ = Y(HPO4)2-
-llnl_gamma 4.0
log_k +9.9000
-delta_H 0 # Not possible to calculate enthalpy of reaction Y(HPO4)2-
# Enthalpy of formation: -0 kcal/mol
2.0000 H2O + 1.0000 Y+++ = Y(OH)2+ +2.0000 H+
-llnl_gamma 4.0
log_k -16.3902
-delta_H 0 # Not possible to calculate enthalpy of reaction Y(OH)2+
# Enthalpy of formation: -0 kcal/mol
3.0000 H2O + 1.0000 Y+++ = Y(OH)3 +3.0000 H+
-llnl_gamma 3.0
log_k -25.9852
-delta_H 0 # Not possible to calculate enthalpy of reaction Y(OH)3
# Enthalpy of formation: -0 kcal/mol
4.0000 H2O + 1.0000 Y+++ = Y(OH)4- +4.0000 H+
-llnl_gamma 4.0
log_k -36.4803
-delta_H 0 # Not possible to calculate enthalpy of reaction Y(OH)4-
# Enthalpy of formation: -0 kcal/mol
2.0000 HPO4-- + 1.0000 Y+++ = Y(PO4)2--- +2.0000 H+
-llnl_gamma 4.0
log_k -3.2437
-delta_H 0 # Not possible to calculate enthalpy of reaction Y(PO4)2-3
# Enthalpy of formation: -0 kcal/mol
2.0000 SO4-- + 1.0000 Y+++ = Y(SO4)2-
-llnl_gamma 4.0
log_k +4.9000
-delta_H 0 # Not possible to calculate enthalpy of reaction Y(SO4)2-
# Enthalpy of formation: -0 kcal/mol
2.0000 Y+++ + 2.0000 H2O = Y2(OH)2++++ +2.0000 H+
-llnl_gamma 5.5
log_k -14.1902
-delta_H 0 # Not possible to calculate enthalpy of reaction Y2(OH)2+4
# Enthalpy of formation: -0 kcal/mol
1.0000 Y+++ + 1.0000 HAcetate = YAcetate++ +1.0000 H+
-llnl_gamma 4.5
log_k -2.1184
-delta_H -17.2799 kJ/mol # Calculated enthalpy of reaction YAcetate+2
# Enthalpy of formation: -291.13 kcal/mol
-analytic -1.2080e+001 1.2015e-003 -8.4186e+002 3.4522e+000 3.4647e+005
# -Range: 0-300
1.0000 Y+++ + 1.0000 HCO3- = YCO3+ +1.0000 H+
-llnl_gamma 4.0
log_k -2.2788
-delta_H 0 # Not possible to calculate enthalpy of reaction YCO3+
# Enthalpy of formation: -0 kcal/mol
1.0000 Y+++ + 1.0000 Cl- = YCl++
-llnl_gamma 4.5
log_k +0.3000
-delta_H 0 # Not possible to calculate enthalpy of reaction YCl+2
# Enthalpy of formation: -0 kcal/mol
1.0000 Y+++ + 1.0000 F- = YF++
-llnl_gamma 4.5
log_k +4.3000
-delta_H 0 # Not possible to calculate enthalpy of reaction YF+2
# Enthalpy of formation: -0 kcal/mol
2.0000 F- + 1.0000 Y+++ = YF2+
-llnl_gamma 4.0
log_k +7.8000
-delta_H 0 # Not possible to calculate enthalpy of reaction YF2+
# Enthalpy of formation: -0 kcal/mol
3.0000 F- + 1.0000 Y+++ = YF3
-llnl_gamma 3.0
log_k +11.2000
-delta_H 0 # Not possible to calculate enthalpy of reaction YF3
# Enthalpy of formation: -0 kcal/mol
1.0000 Y+++ + 1.0000 HPO4-- + 1.0000 H+ = YH2PO4++
-llnl_gamma 4.5
log_k +9.6054
-delta_H 0 # Not possible to calculate enthalpy of reaction YH2PO4+2
# Enthalpy of formation: -0 kcal/mol
1.0000 Y+++ + 1.0000 HCO3- = YHCO3++
-llnl_gamma 4.5
log_k +2.3000
-delta_H 0 # Not possible to calculate enthalpy of reaction YHCO3+2
# Enthalpy of formation: -0 kcal/mol
1.0000 Y+++ + 1.0000 HPO4-- = YHPO4+
-llnl_gamma 4.0
log_k +5.9000
-delta_H 0 # Not possible to calculate enthalpy of reaction YHPO4+
# Enthalpy of formation: -0 kcal/mol
1.0000 Y+++ + 1.0000 NO3- = YNO3++
-llnl_gamma 4.5
log_k +0.4000
-delta_H 0 # Not possible to calculate enthalpy of reaction YNO3+2
# Enthalpy of formation: -0 kcal/mol
1.0000 Y+++ + 1.0000 H2O = YOH++ +1.0000 H+
-llnl_gamma 4.5
log_k -7.6951
-delta_H 0 # Not possible to calculate enthalpy of reaction YOH+2
# Enthalpy of formation: -0 kcal/mol
1.0000 Y+++ + 1.0000 HPO4-- = YPO4 +1.0000 H+
-llnl_gamma 3.0
log_k +0.2782
-delta_H 0 # Not possible to calculate enthalpy of reaction YPO4
# Enthalpy of formation: -0 kcal/mol
1.0000 Y+++ + 1.0000 SO4-- = YSO4+
-llnl_gamma 4.0
log_k +3.4000
-delta_H 0 # Not possible to calculate enthalpy of reaction YSO4+
# Enthalpy of formation: -0 kcal/mol
2.0000 HAcetate + 1.0000 Yb+++ = Yb(Acetate)2+ +2.0000 H+
-llnl_gamma 4.0
log_k -5.131
-delta_H -30.334 kJ/mol # Calculated enthalpy of reaction Yb(Acetate)2+
# Enthalpy of formation: -399.75 kcal/mol
-analytic -3.4286e+001 9.4069e-004 -6.5120e+002 1.0071e+001 5.4773e+005
# -Range: 0-300
3.0000 HAcetate + 1.0000 Yb+++ = Yb(Acetate)3 +3.0000 H+
-llnl_gamma 3.0
log_k -8.5688
-delta_H -51.4214 kJ/mol # Calculated enthalpy of reaction Yb(Acetate)3
# Enthalpy of formation: -520.89 kcal/mol
-analytic -6.2211e+001 -6.1589e-004 5.9577e+002 1.7954e+001 6.6116e+005
# -Range: 0-300
2.0000 HCO3- + 1.0000 Yb+++ = Yb(CO3)2- +2.0000 H+
-llnl_gamma 4.0
log_k -7.0576
-delta_H 0 # Not possible to calculate enthalpy of reaction Yb(CO3)2-
# Enthalpy of formation: -0 kcal/mol
2.0000 HPO4-- + 1.0000 Yb+++ = Yb(HPO4)2-
-llnl_gamma 4.0
log_k +10.2000
-delta_H 0 # Not possible to calculate enthalpy of reaction Yb(HPO4)2-
# Enthalpy of formation: -0 kcal/mol
# Redundant with YbO2-
#4.0000 H2O + 1.0000 Yb+++ = Yb(OH)4- +4.0000 H+
# -llnl_gamma 4.0
# log_k -32.6803
# -delta_H 0 # Not possible to calculate enthalpy of reaction Yb(OH)4-
## Enthalpy of formation: -0 kcal/mol
2.0000 HPO4-- + 1.0000 Yb+++ = Yb(PO4)2--- +2.0000 H+
-llnl_gamma 4.0
log_k -2.7437
-delta_H 0 # Not possible to calculate enthalpy of reaction Yb(PO4)2-3
# Enthalpy of formation: -0 kcal/mol
2.0000 SO4-- + 1.0000 Yb+++ = Yb(SO4)2-
-llnl_gamma 4.0
log_k +5.1000
-delta_H 0 # Not possible to calculate enthalpy of reaction Yb(SO4)2-
# Enthalpy of formation: -0 kcal/mol
1.0000 Yb+++ + 1.0000 HAcetate = YbAcetate++ +1.0000 H+
-llnl_gamma 4.5
log_k -2.199
-delta_H -15.2298 kJ/mol # Calculated enthalpy of reaction YbAcetate+2
# Enthalpy of formation: -280.04 kcal/mol
-analytic -8.5003e+000 2.2459e-003 -9.6434e+002 2.0630e+000 3.3550e+005
# -Range: 0-300
1.0000 Yb+++ + 1.0000 HCO3- = YbCO3+ +1.0000 H+
-llnl_gamma 4.0
log_k -2.0392
-delta_H 82.8348 kJ/mol # Calculated enthalpy of reaction YbCO3+
# Enthalpy of formation: -305.4 kcal/mol
-analytic 2.3533e+002 5.4436e-002 -6.7871e+003 -9.3280e+001 -1.0598e+002
# -Range: 0-300
1.0000 Yb+++ + 1.0000 Cl- = YbCl++
-llnl_gamma 4.5
log_k +0.1620
-delta_H 13.9453 kJ/mol # Calculated enthalpy of reaction YbCl+2
# Enthalpy of formation: -196.9 kcal/mol
-analytic 8.0452e+001 3.8343e-002 -1.8176e+003 -3.4594e+001 -2.8386e+001
# -Range: 0-300
2.0000 Cl- + 1.0000 Yb+++ = YbCl2+
-llnl_gamma 4.0
log_k -0.2624
-delta_H 17.4305 kJ/mol # Calculated enthalpy of reaction YbCl2+
# Enthalpy of formation: -236 kcal/mol
-analytic 2.1708e+002 8.0550e-002 -5.4744e+003 -9.0101e+001 -8.5487e+001
# -Range: 0-300
3.0000 Cl- + 1.0000 Yb+++ = YbCl3
-llnl_gamma 3.0
log_k -0.7601
-delta_H 8.36382 kJ/mol # Calculated enthalpy of reaction YbCl3
# Enthalpy of formation: -278.1 kcal/mol
-analytic 4.0887e+002 1.2992e-001 -1.0578e+004 -1.6684e+002 -1.6518e+002
# -Range: 0-300
4.0000 Cl- + 1.0000 Yb+++ = YbCl4-
-llnl_gamma 4.0
log_k -1.1845
-delta_H -15.7653 kJ/mol # Calculated enthalpy of reaction YbCl4-
# Enthalpy of formation: -323.8 kcal/mol
-analytic 4.7560e+002 1.3032e-001 -1.2452e+004 -1.9149e+002 -1.9444e+002
# -Range: 0-300
1.0000 Yb+++ + 1.0000 F- = YbF++
-llnl_gamma 4.5
log_k +4.8085
-delta_H 23.2212 kJ/mol # Calculated enthalpy of reaction YbF+2
# Enthalpy of formation: -234.9 kcal/mol
-analytic 1.0291e+002 4.2493e-002 -2.7637e+003 -4.1008e+001 -4.3156e+001
# -Range: 0-300
2.0000 F- + 1.0000 Yb+++ = YbF2+
-llnl_gamma 4.0
log_k +8.3709
-delta_H 12.1336 kJ/mol # Calculated enthalpy of reaction YbF2+
# Enthalpy of formation: -317.7 kcal/mol
-analytic 2.4281e+002 8.5385e-002 -5.6900e+003 -9.7299e+001 -8.8859e+001
# -Range: 0-300
3.0000 F- + 1.0000 Yb+++ = YbF3
-llnl_gamma 3.0
log_k +11.0537
-delta_H -13.1796 kJ/mol # Calculated enthalpy of reaction YbF3
# Enthalpy of formation: -403.9 kcal/mol
-analytic 4.5227e+002 1.3659e-001 -1.0595e+004 -1.8038e+002 -1.6546e+002
# -Range: 0-300
4.0000 F- + 1.0000 Yb+++ = YbF4-
-llnl_gamma 4.0
log_k +13.2234
-delta_H -60.2496 kJ/mol # Calculated enthalpy of reaction YbF4-
# Enthalpy of formation: -495.3 kcal/mol
-analytic 5.0369e+002 1.3726e-001 -1.0671e+004 -2.0026e+002 -1.6666e+002
# -Range: 0-300
1.0000 Yb+++ + 1.0000 HPO4-- + 1.0000 H+ = YbH2PO4++
-llnl_gamma 4.5
log_k +9.5217
-delta_H -20.0204 kJ/mol # Calculated enthalpy of reaction YbH2PO4+2
# Enthalpy of formation: -473.9 kcal/mol
-analytic 1.0919e+002 6.3749e-002 3.8909e+002 -4.8469e+001 6.0389e+000
# -Range: 0-300
1.0000 Yb+++ + 1.0000 HCO3- = YbHCO3++
-llnl_gamma 4.5
log_k +1.8398
-delta_H 5.43083 kJ/mol # Calculated enthalpy of reaction YbHCO3+2
# Enthalpy of formation: -323.9 kcal/mol
-analytic 3.9175e+001 3.1796e-002 6.9728e+001 -1.9002e+001 1.0762e+000
# -Range: 0-300
1.0000 Yb+++ + 1.0000 HPO4-- = YbHPO4+
-llnl_gamma 4.0
log_k +6.0000
-delta_H 0 # Not possible to calculate enthalpy of reaction YbHPO4+
# Enthalpy of formation: -0 kcal/mol
1.0000 Yb+++ + 1.0000 NO3- = YbNO3++
-llnl_gamma 4.5
log_k +0.2148
-delta_H -32.9323 kJ/mol # Calculated enthalpy of reaction YbNO3+2
# Enthalpy of formation: -217.6 kcal/mol
-analytic 1.7237e+001 2.5684e-002 2.2806e+003 -1.3055e+001 3.5581e+001
# -Range: 0-300
1.0000 Yb+++ + 1.0000 H2O = YbO+ +2.0000 H+
-llnl_gamma 4.0
log_k -15.7506
-delta_H 105.508 kJ/mol # Calculated enthalpy of reaction YbO+
# Enthalpy of formation: -203.4 kcal/mol
-analytic 1.7675e+002 2.9078e-002 -1.3106e+004 -6.3534e+001 -2.0456e+002
# -Range: 0-300
2.0000 H2O + 1.0000 Yb+++ = YbO2- +4.0000 H+
-llnl_gamma 4.0
log_k -32.6741
-delta_H 267.918 kJ/mol # Calculated enthalpy of reaction YbO2-
# Enthalpy of formation: -232.9 kcal/mol
-analytic 1.5529e+002 1.0053e-002 -1.8749e+004 -5.1764e+001 -2.9260e+002
# -Range: 0-300
2.0000 H2O + 1.0000 Yb+++ = YbO2H +3.0000 H+
-llnl_gamma 3.0
log_k -23.878
-delta_H 211.016 kJ/mol # Calculated enthalpy of reaction YbO2H
# Enthalpy of formation: -246.5 kcal/mol
-analytic 3.2148e+002 4.4821e-002 -2.1971e+004 -1.1519e+002 -3.4293e+002
# -Range: 0-300
1.0000 Yb+++ + 1.0000 H2O = YbOH++ +1.0000 H+
-llnl_gamma 4.5
log_k -7.6143
-delta_H 74.9647 kJ/mol # Calculated enthalpy of reaction YbOH+2
# Enthalpy of formation: -210.7 kcal/mol
-analytic 5.8142e+001 1.1402e-002 -5.6488e+003 -2.0289e+001 -8.8160e+001
# -Range: 0-300
1.0000 Yb+++ + 1.0000 HPO4-- = YbPO4 +1.0000 H+
-llnl_gamma 3.0
log_k +0.5782
-delta_H 0 # Not possible to calculate enthalpy of reaction YbPO4
# Enthalpy of formation: -0 kcal/mol
1.0000 Yb+++ + 1.0000 SO4-- = YbSO4+
-llnl_gamma 4.0
log_k +3.5697
-delta_H 1424.65 kJ/mol # Calculated enthalpy of reaction YbSO4+
# Enthalpy of formation: -37.2 kcal/mol
-analytic 3.0675e+002 8.6527e-002 -9.0298e+003 -1.2069e+002 -1.4099e+002
# -Range: 0-300
2.0000 HAcetate + 1.0000 Zn++ = Zn(Acetate)2 +2.0000 H+
-llnl_gamma 3.0
log_k -6.062
-delta_H -11.0458 kJ/mol # Calculated enthalpy of reaction Zn(Acetate)2
# Enthalpy of formation: -271.5 kcal/mol
-analytic -2.2038e+001 2.6133e-003 -2.7652e+003 6.8501e+000 6.7086e+005
# -Range: 0-300
3.0000 HAcetate + 1.0000 Zn++ = Zn(Acetate)3- +3.0000 H+
-llnl_gamma 4.0
log_k -10.0715
-delta_H 25.355 kJ/mol # Calculated enthalpy of reaction Zn(Acetate)3-
# Enthalpy of formation: -378.9 kcal/mol
-analytic 3.5104e+001 -6.1568e-003 -1.3379e+004 -8.7697e+000 2.0670e+006
# -Range: 0-300
4.0000 Cyanide- + 1.0000 Zn++ = Zn(Cyanide)4--
-llnl_gamma 4.0
log_k +16.7040
-delta_H -107.305 kJ/mol # Calculated enthalpy of reaction Zn(Cyanide)4-2
# Enthalpy of formation: 341.806 kJ/mol
-analytic 3.6586e+002 1.2655e-001 -2.9546e+003 -1.5232e+002 -4.6213e+001
# -Range: 0-300
2.0000 N3- + 1.0000 Zn++ = Zn(N3)2
-llnl_gamma 3.0
log_k +1.1954
-delta_H 0 # Not possible to calculate enthalpy of reaction Zn(N3)2
# Enthalpy of formation: -0 kcal/mol
1.0000 Zn++ + 1.0000 NH3 = Zn(NH3)++
-llnl_gamma 4.5
log_k +2.0527
-delta_H 0 # Not possible to calculate enthalpy of reaction Zn(NH3)+2
# Enthalpy of formation: -0 kcal/mol
2.0000 NH3 + 1.0000 Zn++ = Zn(NH3)2++
-llnl_gamma 4.5
log_k +4.2590
-delta_H 0 # Not possible to calculate enthalpy of reaction Zn(NH3)2+2
# Enthalpy of formation: -0 kcal/mol
3.0000 NH3 + 1.0000 Zn++ = Zn(NH3)3++
-llnl_gamma 4.5
log_k +6.4653
-delta_H 0 # Not possible to calculate enthalpy of reaction Zn(NH3)3+2
# Enthalpy of formation: -0 kcal/mol
4.0000 NH3 + 1.0000 Zn++ = Zn(NH3)4++
-llnl_gamma 4.5
log_k +8.3738
-delta_H -54.9027 kJ/mol # Calculated enthalpy of reaction Zn(NH3)4+2
# Enthalpy of formation: -533.636 kJ/mol
-analytic 1.5851e+002 -6.3376e-003 -4.6783e+003 -5.3560e+001 -7.3047e+001
# -Range: 0-300
2.0000 H2O + 1.0000 Zn++ = Zn(OH)2 +2.0000 H+
-llnl_gamma 3.0
log_k -17.3282
-delta_H 0 # Not possible to calculate enthalpy of reaction Zn(OH)2
# Enthalpy of formation: -0 kcal/mol
3.0000 H2O + 1.0000 Zn++ = Zn(OH)3- +3.0000 H+
-llnl_gamma 4.0
log_k -28.8369
-delta_H 0 # Not possible to calculate enthalpy of reaction Zn(OH)3-
# Enthalpy of formation: -0 kcal/mol
4.0000 H2O + 1.0000 Zn++ = Zn(OH)4-- +4.0000 H+
-llnl_gamma 4.0
log_k -41.6052
-delta_H 0 # Not possible to calculate enthalpy of reaction Zn(OH)4-2
# Enthalpy of formation: -0 kcal/mol
1.0000 Zn++ + 1.0000 H2O + 1.0000 Cl- = Zn(OH)Cl +1.0000 H+
-llnl_gamma 3.0
log_k -7.5417
-delta_H 0 # Not possible to calculate enthalpy of reaction Zn(OH)Cl
# Enthalpy of formation: -0 kcal/mol
2.0000 Thiocyanate- + 1.0000 Zn++ = Zn(Thiocyanate)2
-llnl_gamma 3.0
log_k +0.8800
-delta_H 0 # Not possible to calculate enthalpy of reaction Zn(Thiocyanate)2
# Enthalpy of formation: -0 kcal/mol
4.0000 Thiocyanate- + 1.0000 Zn++ = Zn(Thiocyanate)4--
-llnl_gamma 4.0
log_k +1.2479
-delta_H 0 # Not possible to calculate enthalpy of reaction Zn(Thiocyanate)4-2
# Enthalpy of formation: -0 kcal/mol
1.0000 Zn++ + 1.0000 Br- = ZnBr+
-llnl_gamma 4.0
log_k -0.6365
-delta_H 0 # Not possible to calculate enthalpy of reaction ZnBr+
# Enthalpy of formation: -0 kcal/mol
2.0000 Br- + 1.0000 Zn++ = ZnBr2
-llnl_gamma 3.0
log_k -1.0492
-delta_H 0 # Not possible to calculate enthalpy of reaction ZnBr2
# Enthalpy of formation: -0 kcal/mol
3.0000 Br- + 1.0000 Zn++ = ZnBr3-
-llnl_gamma 4.0
log_k -1.8474
-delta_H 0 # Not possible to calculate enthalpy of reaction ZnBr3-
# Enthalpy of formation: -0 kcal/mol
1.0000 Zn++ + 1.0000 HAcetate = ZnAcetate+ +1.0000 H+
-llnl_gamma 4.0
log_k -3.1519
-delta_H -9.87424 kJ/mol # Calculated enthalpy of reaction ZnAcetate+
# Enthalpy of formation: -155.12 kcal/mol
-analytic -7.9367e+000 2.8564e-003 -1.4514e+003 2.5010e+000 2.3343e+005
# -Range: 0-300
1.0000 Zn++ + 1.0000 HCO3- = ZnCO3 +1.0000 H+
-llnl_gamma 3.0
log_k -6.4288
-delta_H 0 # Not possible to calculate enthalpy of reaction ZnCO3
# Enthalpy of formation: -0 kcal/mol
1.0000 Zn++ + 1.0000 Cl- = ZnCl+
-llnl_gamma 4.0
log_k +0.1986
-delta_H 43.317 kJ/mol # Calculated enthalpy of reaction ZnCl+
# Enthalpy of formation: -66.24 kcal/mol
-analytic 1.1235e+002 4.4461e-002 -4.1662e+003 -4.5023e+001 -6.5042e+001
# -Range: 0-300
2.0000 Cl- + 1.0000 Zn++ = ZnCl2
-llnl_gamma 3.0
log_k +0.2507
-delta_H 31.1541 kJ/mol # Calculated enthalpy of reaction ZnCl2
# Enthalpy of formation: -109.08 kcal/mol
-analytic 1.7824e+002 7.5733e-002 -4.6251e+003 -7.4770e+001 -7.2224e+001
# -Range: 0-300
3.0000 Cl- + 1.0000 Zn++ = ZnCl3-
-llnl_gamma 4.0
log_k -0.0198
-delta_H 22.5894 kJ/mol # Calculated enthalpy of reaction ZnCl3-
# Enthalpy of formation: -151.06 kcal/mol
-analytic 1.3889e+002 7.4712e-002 -2.1527e+003 -6.2200e+001 -3.3633e+001
# -Range: 0-300
4.0000 Cl- + 1.0000 Zn++ = ZnCl4--
-llnl_gamma 4.0
log_k +0.8605
-delta_H 4.98733 kJ/mol # Calculated enthalpy of reaction ZnCl4-2
# Enthalpy of formation: -195.2 kcal/mol
-analytic 8.4294e+001 7.0021e-002 3.9150e+002 -4.2664e+001 6.0834e+000
# -Range: 0-300
1.0000 Zn++ + 1.0000 ClO4- = ZnClO4+
-llnl_gamma 4.0
log_k +1.2768
-delta_H 0 # Not possible to calculate enthalpy of reaction ZnClO4+
# Enthalpy of formation: -0 kcal/mol
1.0000 Zn++ + 1.0000 F- = ZnF+
-llnl_gamma 4.0
log_k +1.1500
-delta_H 0 # Not possible to calculate enthalpy of reaction ZnF+
# Enthalpy of formation: -0 kcal/mol
1.0000 Zn++ + 1.0000 HPO4-- + 1.0000 H+ = ZnH2PO4+
-llnl_gamma 4.0
log_k +0.4300
-delta_H 0 # Not possible to calculate enthalpy of reaction ZnH2PO4+
# Enthalpy of formation: -0 kcal/mol
1.0000 Zn++ + 1.0000 HCO3- = ZnHCO3+
-llnl_gamma 4.0
log_k +1.4200
-delta_H 0 # Not possible to calculate enthalpy of reaction ZnHCO3+
# Enthalpy of formation: -0 kcal/mol
-analytic 5.1115e+002 1.2911e-001 -1.5292e+004 -2.0083e+002 -2.2721e+002
# -Range: 25-300
1.0000 Zn++ + 1.0000 HPO4-- = ZnHPO4
-llnl_gamma 3.0
log_k +3.2600
-delta_H 0 # Not possible to calculate enthalpy of reaction ZnHPO4
# Enthalpy of formation: -0 kcal/mol
1.0000 Zn++ + 1.0000 I- = ZnI+
-llnl_gamma 4.0
log_k -3.0134
-delta_H 0 # Not possible to calculate enthalpy of reaction ZnI+
# Enthalpy of formation: -0 kcal/mol
2.0000 I- + 1.0000 Zn++ = ZnI2
-llnl_gamma 3.0
log_k -1.8437
-delta_H 0 # Not possible to calculate enthalpy of reaction ZnI2
# Enthalpy of formation: -0 kcal/mol
3.0000 I- + 1.0000 Zn++ = ZnI3-
-llnl_gamma 4.0
log_k -2.0054
-delta_H 0 # Not possible to calculate enthalpy of reaction ZnI3-
# Enthalpy of formation: -0 kcal/mol
4.0000 I- + 1.0000 Zn++ = ZnI4--
-llnl_gamma 4.0
log_k -2.6052
-delta_H 0 # Not possible to calculate enthalpy of reaction ZnI4-2
# Enthalpy of formation: -0 kcal/mol
1.0000 Zn++ + 1.0000 N3- = ZnN3+
-llnl_gamma 4.0
log_k +0.4420
-delta_H 0 # Not possible to calculate enthalpy of reaction ZnN3+
# Enthalpy of formation: -0 kcal/mol
1.0000 Zn++ + 1.0000 H2O = ZnOH+ +1.0000 H+
-llnl_gamma 4.0
log_k -8.96
-delta_H 0 # Not possible to calculate enthalpy of reaction ZnOH+
# Enthalpy of formation: -0 kcal/mol
-analytic -7.8600e-001 -2.9499e-004 -2.8673e+003 6.1892e-001 -4.2576e+001
# -Range: 25-300
1.0000 Zn++ + 1.0000 HPO4-- = ZnPO4- +1.0000 H+
-llnl_gamma 4.0
log_k -4.3018
-delta_H 0 # Not possible to calculate enthalpy of reaction ZnPO4-
# Enthalpy of formation: -0 kcal/mol
1.0000 Zn++ + 1.0000 SO4-- = ZnSO4
-llnl_gamma 3.0
log_k +2.3062
-delta_H 15.277 kJ/mol # Calculated enthalpy of reaction ZnSO4
# Enthalpy of formation: -1047.71 kJ/mol
-analytic 1.3640e+002 5.1256e-002 -3.4422e+003 -5.5695e+001 -5.8501e+001
# -Range: 0-200
1.0000 Zn++ + 1.0000 SeO4-- = ZnSeO4
-llnl_gamma 3.0
log_k +2.1900
-delta_H 0 # Not possible to calculate enthalpy of reaction ZnSeO4
# Enthalpy of formation: -0 kcal/mol
3.0000 H2O + 1.0000 Zr++++ = Zr(OH)3+ +3.0000 H+
-llnl_gamma 4.0
log_k -0.6693
-delta_H 0 # Not possible to calculate enthalpy of reaction Zr(OH)3+
# Enthalpy of formation: -0 kcal/mol
4.0000 H2O + 1.0000 Zr++++ = Zr(OH)4 +4.0000 H+
-llnl_gamma 3.0
log_k -1.4666
-delta_H 0 # Not possible to calculate enthalpy of reaction Zr(OH)4
# Enthalpy of formation: -0 kcal/mol
5.0000 H2O + 1.0000 Zr++++ = Zr(OH)5- +5.0000 H+
-llnl_gamma 4.0
log_k -15.9754
-delta_H 0 # Not possible to calculate enthalpy of reaction Zr(OH)5-
# Enthalpy of formation: -0 kcal/mol
2.0000 SO4-- + 1.0000 Zr++++ = Zr(SO4)2
-llnl_gamma 3.0
log_k +6.2965
-delta_H 0 # Not possible to calculate enthalpy of reaction Zr(SO4)2
# Enthalpy of formation: -0 kcal/mol
3.0000 SO4-- + 1.0000 Zr++++ = Zr(SO4)3--
-llnl_gamma 4.0
log_k +7.3007
-delta_H 0 # Not possible to calculate enthalpy of reaction Zr(SO4)3-2
# Enthalpy of formation: -0 kcal/mol
4.0000 H2O + 3.0000 Zr++++ = Zr3(OH)4+8 +4.0000 H+
-llnl_gamma 6.0
log_k -0.5803
-delta_H 0 # Not possible to calculate enthalpy of reaction Zr3(OH)4+8
# Enthalpy of formation: -0 kcal/mol
8.0000 H2O + 4.0000 Zr++++ = Zr4(OH)8+8 +8.0000 H+
-llnl_gamma 6.0
log_k -5.9606
-delta_H 0 # Not possible to calculate enthalpy of reaction Zr4(OH)8+8
# Enthalpy of formation: -0 kcal/mol
1.0000 Zr++++ + 1.0000 F- = ZrF+++
-llnl_gamma 5.0
log_k +8.5835
-delta_H 0 # Not possible to calculate enthalpy of reaction ZrF+3
# Enthalpy of formation: -0 kcal/mol
2.0000 F- + 1.0000 Zr++++ = ZrF2++
-llnl_gamma 4.5
log_k +15.7377
-delta_H 0 # Not possible to calculate enthalpy of reaction ZrF2+2
# Enthalpy of formation: -0 kcal/mol
3.0000 F- + 1.0000 Zr++++ = ZrF3+
-llnl_gamma 4.0
log_k +21.2792
-delta_H 0 # Not possible to calculate enthalpy of reaction ZrF3+
# Enthalpy of formation: -0 kcal/mol
4.0000 F- + 1.0000 Zr++++ = ZrF4
-llnl_gamma 3.0
log_k +25.9411
-delta_H 0 # Not possible to calculate enthalpy of reaction ZrF4
# Enthalpy of formation: -0 kcal/mol
5.0000 F- + 1.0000 Zr++++ = ZrF5-
-llnl_gamma 4.0
log_k +30.3098
-delta_H 0 # Not possible to calculate enthalpy of reaction ZrF5-
# Enthalpy of formation: -0 kcal/mol
6.0000 F- + 1.0000 Zr++++ = ZrF6--
-llnl_gamma 4.0
log_k +34.0188
-delta_H 0 # Not possible to calculate enthalpy of reaction ZrF6-2
# Enthalpy of formation: -0 kcal/mol
1.0000 Zr++++ + 1.0000 H2O = ZrOH+++ +1.0000 H+
-llnl_gamma 5.0
log_k +0.0457
-delta_H 0 # Not possible to calculate enthalpy of reaction ZrOH+3
# Enthalpy of formation: -0 kcal/mol
1.0000 Zr++++ + 1.0000 SO4-- = ZrSO4++
-llnl_gamma 4.5
log_k +3.6064
-delta_H 0 # Not possible to calculate enthalpy of reaction ZrSO4+2
# Enthalpy of formation: -0 kcal/mol
2.0000 H+ + 1.0000 O_phthalate-2 = H2O_phthalate
-llnl_gamma 3.0
log_k +8.3580
-delta_H 0 # Not possible to calculate enthalpy of reaction H2O_phthalate
# Enthalpy of formation: -0 kcal/mol
PHASES
# 1122 minerals
(UO2)2As2O7
(UO2)2As2O7 +2.0000 H+ +1.0000 H2O = + 2.0000 H2AsO4- + 2.0000 UO2++
log_k 7.7066
-delta_H -145.281 kJ/mol # Calculated enthalpy of reaction (UO2)2As2O7
# Enthalpy of formation: -3426 kJ/mol
-analytic -1.6147e+002 -6.3487e-002 1.0052e+004 6.2384e+001 1.5691e+002
# -Range: 0-300
(UO2)2Cl3
(UO2)2Cl3 = + 1.0000 UO2+ + 1.0000 UO2++ + 3.0000 Cl-
log_k 12.7339
-delta_H -140.866 kJ/mol # Calculated enthalpy of reaction (UO2)2Cl3
# Enthalpy of formation: -2404.5 kJ/mol
-analytic -2.3895e+002 -9.2925e-002 1.1722e+004 9.6999e+001 1.8298e+002
# -Range: 0-300
(UO2)2P2O7
(UO2)2P2O7 +1.0000 H2O = + 2.0000 HPO4-- + 2.0000 UO2++
log_k -14.6827
-delta_H -103.726 kJ/mol # Calculated enthalpy of reaction (UO2)2P2O7
# Enthalpy of formation: -4232.6 kJ/mol
-analytic -3.4581e+002 -1.3987e-001 1.0703e+004 1.3613e+002 1.6712e+002
# -Range: 0-300
(UO2)3(AsO4)2
(UO2)3(AsO4)2 +4.0000 H+ = + 2.0000 H2AsO4- + 3.0000 UO2++
log_k 9.3177
-delta_H -186.72 kJ/mol # Calculated enthalpy of reaction (UO2)3(AsO4)2
# Enthalpy of formation: -4689.4 kJ/mol
-analytic -1.9693e+002 -7.3236e-002 1.2936e+004 7.4631e+001 2.0192e+002
# -Range: 0-300
(UO2)3(PO4)2
(UO2)3(PO4)2 +2.0000 H+ = + 2.0000 HPO4-- + 3.0000 UO2++
log_k -14.0241
-delta_H -149.864 kJ/mol # Calculated enthalpy of reaction (UO2)3(PO4)2
# Enthalpy of formation: -5491.3 kJ/mol
-analytic -3.6664e+002 -1.4347e-001 1.3486e+004 1.4148e+002 2.1054e+002
# -Range: 0-300
(UO2)3(PO4)2:4H2O
(UO2)3(PO4)2:4H2O +2.0000 H+ = + 2.0000 HPO4-- + 3.0000 UO2++ + 4.0000 H2O
log_k -27.0349
-delta_H -45.4132 kJ/mol # Calculated enthalpy of reaction (UO2)3(PO4)2:4H2O
# Enthalpy of formation: -6739.1 kJ/mol
-analytic -1.5721e+002 -4.1375e-002 5.2046e+003 5.0531e+001 8.8434e+001
# -Range: 0-200
(VO)3(PO4)2
(VO)3(PO4)2 +2.0000 H+ = + 2.0000 HPO4-- + 3.0000 VO++
log_k 48.7864
-delta_H 0 # Not possible to calculate enthalpy of reaction (VO)3(PO4)2
# Enthalpy of formation: 0 kcal/mol
Acanthite
Ag2S +1.0000 H+ = + 1.0000 HS- + 2.0000 Ag+
log_k -36.0346
-delta_H 226.982 kJ/mol # Calculated enthalpy of reaction Acanthite
# Enthalpy of formation: -7.55 kcal/mol
-analytic -1.6067e+002 -4.7139e-002 -7.4522e+003 6.6140e+001 -1.1624e+002
# -Range: 0-300
Afwillite
Ca3Si2O4(OH)6 +6.0000 H+ = + 2.0000 SiO2 + 3.0000 Ca++ + 6.0000 H2O
log_k 60.0452
-delta_H -316.059 kJ/mol # Calculated enthalpy of reaction Afwillite
# Enthalpy of formation: -1143.31 kcal/mol
-analytic 1.8353e+001 1.9014e-003 1.8478e+004 -6.6311e+000 -4.0227e+005
# -Range: 0-300
Ag
Ag +1.0000 H+ +0.2500 O2 = + 0.5000 H2O + 1.0000 Ag+
log_k 7.9937
-delta_H -34.1352 kJ/mol # Calculated enthalpy of reaction Ag
# Enthalpy of formation: 0 kcal/mol
-analytic -1.4144e+001 -3.8466e-003 2.2642e+003 6.3388e+000 3.5334e+001
# -Range: 0-300
Ag3PO4
Ag3PO4 +1.0000 H+ = + 1.0000 HPO4-- + 3.0000 Ag+
log_k -5.2282
-delta_H 0 # Not possible to calculate enthalpy of reaction Ag3PO4
# Enthalpy of formation: 0 kcal/mol
Ahlfeldite
NiSeO3:2H2O = + 1.0000 Ni++ + 1.0000 SeO3-- + 2.0000 H2O
log_k -4.4894
-delta_H -25.7902 kJ/mol # Calculated enthalpy of reaction Ahlfeldite
# Enthalpy of formation: -265.07 kcal/mol
-analytic -2.6210e+001 -1.6952e-002 1.0405e+003 9.4054e+000 1.7678e+001
# -Range: 0-200
Akermanite
Ca2MgSi2O7 +6.0000 H+ = + 1.0000 Mg++ + 2.0000 Ca++ + 2.0000 SiO2 + 3.0000 H2O
log_k 45.3190
-delta_H -288.575 kJ/mol # Calculated enthalpy of reaction Akermanite
# Enthalpy of formation: -926.497 kcal/mol
-analytic -4.8295e+001 -8.5613e-003 2.0880e+004 1.3798e+001 -7.1975e+005
# -Range: 0-300
Al
Al +3.0000 H+ +0.7500 O2 = + 1.0000 Al+++ + 1.5000 H2O
log_k 149.9292
-delta_H -958.059 kJ/mol # Calculated enthalpy of reaction Al
# Enthalpy of formation: 0 kJ/mol
-analytic -1.8752e+002 -4.6187e-002 5.7127e+004 6.6270e+001 -3.8952e+005
# -Range: 0-300
Al2(SO4)3
Al2(SO4)3 = + 2.0000 Al+++ + 3.0000 SO4--
log_k 19.0535
-delta_H -364.566 kJ/mol # Calculated enthalpy of reaction Al2(SO4)3
# Enthalpy of formation: -3441.04 kJ/mol
-analytic -6.1001e+002 -2.4268e-001 2.9194e+004 2.4383e+002 4.5573e+002
# -Range: 0-300
Al2(SO4)3:6H2O
Al2(SO4)3:6H2O = + 2.0000 Al+++ + 3.0000 SO4-- + 6.0000 H2O
log_k 1.6849
-delta_H -208.575 kJ/mol # Calculated enthalpy of reaction Al2(SO4)3:6H2O
# Enthalpy of formation: -5312.06 kJ/mol
-analytic -7.1642e+002 -2.4552e-001 2.6064e+004 2.8441e+002 4.0691e+002
# -Range: 0-300
AlF3
AlF3 = + 1.0000 Al+++ + 3.0000 F-
log_k -17.2089
-delta_H -34.0441 kJ/mol # Calculated enthalpy of reaction AlF3
# Enthalpy of formation: -1510.4 kJ/mol
-analytic -3.9865e+002 -1.3388e-001 1.0211e+004 1.5642e+002 1.5945e+002
# -Range: 0-300
Alabandite
MnS +1.0000 H+ = + 1.0000 HS- + 1.0000 Mn++
log_k -0.3944
-delta_H -23.3216 kJ/mol # Calculated enthalpy of reaction Alabandite
# Enthalpy of formation: -51 kcal/mol
-analytic -1.5515e+002 -4.8820e-002 4.9049e+003 6.1765e+001 7.6583e+001
# -Range: 0-300
Alamosite
PbSiO3 +2.0000 H+ = + 1.0000 H2O + 1.0000 Pb++ + 1.0000 SiO2
log_k 5.6733
-delta_H -16.5164 kJ/mol # Calculated enthalpy of reaction Alamosite
# Enthalpy of formation: -1146.1 kJ/mol
-analytic 2.9941e+002 6.7871e-002 -8.1706e+003 -1.1582e+002 -1.3885e+002
# -Range: 0-200
Albite
NaAlSi3O8 +4.0000 H+ = + 1.0000 Al+++ + 1.0000 Na+ + 2.0000 H2O + 3.0000 SiO2
log_k 2.7645
-delta_H -51.8523 kJ/mol # Calculated enthalpy of reaction Albite
# Enthalpy of formation: -939.68 kcal/mol
-analytic -1.1694e+001 1.4429e-002 1.3784e+004 -7.2866e+000 -1.6136e+006
# -Range: 0-300
Albite_high
NaAlSi3O8 +4.0000 H+ = + 1.0000 Al+++ + 1.0000 Na+ + 2.0000 H2O + 3.0000 SiO2
log_k 4.0832
-delta_H -62.8562 kJ/mol # Calculated enthalpy of reaction Albite_high
# Enthalpy of formation: -937.05 kcal/mol
-analytic -1.8957e+001 1.3726e-002 1.4801e+004 -4.9732e+000 -1.6442e+006
# -Range: 0-300
Albite_low
NaAlSi3O8 +4.0000 H+ = + 1.0000 Al+++ + 1.0000 Na+ + 2.0000 H2O + 3.0000 SiO2
log_k 2.7645
-delta_H -51.8523 kJ/mol # Calculated enthalpy of reaction Albite_low
# Enthalpy of formation: -939.68 kcal/mol
-analytic -1.2860e+001 1.4481e-002 1.3913e+004 -6.9417e+000 -1.6256e+006
# -Range: 0-300
Alstonite
BaCa(CO3)2 +2.0000 H+ = + 1.0000 Ba++ + 1.0000 Ca++ + 2.0000 HCO3-
log_k 2.5843
-delta_H 0 # Not possible to calculate enthalpy of reaction Alstonite
# Enthalpy of formation: 0 kcal/mol
Alum-K
KAl(SO4)2:12H2O = + 1.0000 Al+++ + 1.0000 K+ + 2.0000 SO4-- + 12.0000 H2O
log_k -4.8818
-delta_H 14.4139 kJ/mol # Calculated enthalpy of reaction Alum-K
# Enthalpy of formation: -1447 kcal/mol
-analytic -8.8025e+002 -2.5706e-001 2.2399e+004 3.5434e+002 3.4978e+002
# -Range: 0-300
Alunite
KAl3(OH)6(SO4)2 +6.0000 H+ = + 1.0000 K+ + 2.0000 SO4-- + 3.0000 Al+++ + 6.0000 H2O
log_k -0.3479
-delta_H -231.856 kJ/mol # Calculated enthalpy of reaction Alunite
# Enthalpy of formation: -1235.6 kcal/mol
-analytic -6.8581e+002 -2.2455e-001 2.6886e+004 2.6758e+002 4.1973e+002
# -Range: 0-300
Am
Am +3.0000 H+ +0.7500 O2 = + 1.0000 Am+++ + 1.5000 H2O
log_k 169.3900
-delta_H -1036.36 kJ/mol # Calculated enthalpy of reaction Am
# Enthalpy of formation: 0 kJ/mol
-analytic -6.7924e+000 -8.9873e-003 5.3327e+004 0.0000e+000 0.0000e+000
# -Range: 0-300
Am(OH)3
Am(OH)3 +3.0000 H+ = + 1.0000 Am+++ + 3.0000 H2O
log_k 15.2218
-delta_H 0 # Not possible to calculate enthalpy of reaction Am(OH)3
# Enthalpy of formation: 0 kcal/mol
Am(OH)3(am)
Am(OH)3 +3.0000 H+ = + 1.0000 Am+++ + 3.0000 H2O
log_k 17.0217
-delta_H 0 # Not possible to calculate enthalpy of reaction Am(OH)3(am)
# Enthalpy of formation: 0 kcal/mol
Am2(CO3)3
Am2(CO3)3 +3.0000 H+ = + 2.0000 Am+++ + 3.0000 HCO3-
log_k -2.3699
-delta_H 0 # Not possible to calculate enthalpy of reaction Am2(CO3)3
# Enthalpy of formation: 0 kcal/mol
Am2C3
Am2C3 +4.5000 O2 +3.0000 H+ = + 2.0000 Am+++ + 3.0000 HCO3-
log_k 503.9594
-delta_H -3097.6 kJ/mol # Calculated enthalpy of reaction Am2C3
# Enthalpy of formation: -151 kJ/mol
-analytic 3.3907e+002 -4.2636e-003 1.4463e+005 -1.2891e+002 2.4559e+003
# -Range: 0-200
Am2O3
Am2O3 +6.0000 H+ = + 2.0000 Am+++ + 3.0000 H2O
log_k 51.7905
-delta_H -400.515 kJ/mol # Calculated enthalpy of reaction Am2O3
# Enthalpy of formation: -1690.4 kJ/mol
-analytic -9.2044e+001 -1.8883e-002 2.3028e+004 2.9192e+001 3.5935e+002
# -Range: 0-300
AmBr3
AmBr3 = + 1.0000 Am+++ + 3.0000 Br-
log_k 21.7826
-delta_H -171.21 kJ/mol # Calculated enthalpy of reaction AmBr3
# Enthalpy of formation: -810 kJ/mol
-analytic 1.0121e+001 -3.0622e-002 6.1964e+003 0.0000e+000 0.0000e+000
# -Range: 0-200
AmCl3
AmCl3 = + 1.0000 Am+++ + 3.0000 Cl-
log_k 14.3513
-delta_H -140.139 kJ/mol # Calculated enthalpy of reaction AmCl3
# Enthalpy of formation: -977.8 kJ/mol
-analytic -1.5000e+001 -3.6701e-002 5.2281e+003 9.1942e+000 8.8785e+001
# -Range: 0-200
AmF3
AmF3 = + 1.0000 Am+++ + 3.0000 F-
log_k -13.1190
-delta_H -34.7428 kJ/mol # Calculated enthalpy of reaction AmF3
# Enthalpy of formation: -1588 kJ/mol
-analytic -4.0514e+001 -3.7312e-002 4.1626e+002 1.4999e+001 7.0827e+000
# -Range: 0-200
AmF4
AmF4 = + 1.0000 Am++++ + 4.0000 F-
log_k -25.1354
-delta_H -37.3904 kJ/mol # Calculated enthalpy of reaction AmF4
# Enthalpy of formation: -1710 kJ/mol
-analytic -4.9592e+001 -4.5210e-002 -9.7251e+001 1.5457e+001 -1.6348e+000
# -Range: 0-200
AmH2
AmH2 +2.0000 H+ +1.0000 O2 = + 1.0000 Am++ + 2.0000 H2O
log_k 128.4208
-delta_H -738.376 kJ/mol # Calculated enthalpy of reaction AmH2
# Enthalpy of formation: -175.8 kJ/mol
-analytic 3.1175e+001 -1.4062e-002 3.6259e+004 -8.1600e+000 5.6578e+002
# -Range: 0-300
AmI3
AmI3 = + 1.0000 Am+++ + 3.0000 I-
log_k 24.7301
-delta_H -175.407 kJ/mol # Calculated enthalpy of reaction AmI3
# Enthalpy of formation: -612 kJ/mol
-analytic -1.3886e+001 -3.6651e-002 7.2094e+003 1.0247e+001 1.2243e+002
# -Range: 0-200
AmO2
AmO2 +4.0000 H+ = + 1.0000 Am++++ + 2.0000 H2O
log_k -9.4203
-delta_H -45.4767 kJ/mol # Calculated enthalpy of reaction AmO2
# Enthalpy of formation: -932.2 kJ/mol
-analytic -7.4658e+001 -1.1661e-002 4.2059e+003 2.2070e+001 6.5650e+001
# -Range: 0-300
AmOBr
AmOBr +2.0000 H+ = + 1.0000 Am+++ + 1.0000 Br- + 1.0000 H2O
log_k 13.7637
-delta_H -131.042 kJ/mol # Calculated enthalpy of reaction AmOBr
# Enthalpy of formation: -893 kJ/mol
-analytic -4.4394e+001 -1.7071e-002 7.3438e+003 1.5605e+001 1.2472e+002
# -Range: 0-200
AmOCl
AmOCl +2.0000 H+ = + 1.0000 Am+++ + 1.0000 Cl- + 1.0000 H2O
log_k 11.3229
-delta_H -119.818 kJ/mol # Calculated enthalpy of reaction AmOCl
# Enthalpy of formation: -949.8 kJ/mol
-analytic -1.2101e+002 -4.1027e-002 8.6801e+003 4.6651e+001 1.3548e+002
# -Range: 0-300
AmOHCO3
AmOHCO3 +2.0000 H+ = + 1.0000 Am+++ + 1.0000 H2O + 1.0000 HCO3-
log_k 3.1519
-delta_H 0 # Not possible to calculate enthalpy of reaction AmOHCO3
# Enthalpy of formation: 0 kcal/mol
AmPO4(am)
AmPO4 +1.0000 H+ = + 1.0000 Am+++ + 1.0000 HPO4--
log_k -12.4682
-delta_H 0 # Not possible to calculate enthalpy of reaction AmPO4(am)
# Enthalpy of formation: 0 kcal/mol
Amesite-14A
Mg4Al4Si2O10(OH)8 +20.0000 H+ = + 2.0000 SiO2 + 4.0000 Al+++ + 4.0000 Mg++ + 14.0000 H2O
log_k 75.4571
-delta_H -797.098 kJ/mol # Calculated enthalpy of reaction Amesite-14A
# Enthalpy of formation: -2145.67 kcal/mol
-analytic -5.4326e+002 -1.4144e-001 5.4150e+004 1.9361e+002 8.4512e+002
# -Range: 0-300
Analcime
Na.96Al.96Si2.04O6:H2O +3.8400 H+ = + 0.9600 Al+++ + 0.9600 Na+ + 2.0400 SiO2 + 2.9200 H2O
log_k 6.1396
-delta_H -75.844 kJ/mol # Calculated enthalpy of reaction Analcime
# Enthalpy of formation: -3296.86 kJ/mol
-analytic -6.8694e+000 6.6052e-003 9.8260e+003 -4.8540e+000 -8.8780e+005
# -Range: 0-300
Analcime-dehy
Na.96Al.96Si2.04O6 +3.8400 H+ = + 0.9600 Al+++ + 0.9600 Na+ + 1.9200 H2O + 2.0400 SiO2
log_k 12.5023
-delta_H -116.641 kJ/mol # Calculated enthalpy of reaction Analcime-dehy
# Enthalpy of formation: -2970.23 kJ/mol
-analytic -7.1134e+000 5.6181e-003 1.2185e+004 -5.0295e+000 -9.3890e+005
# -Range: 0-300
Anatase
TiO2 +2.0000 H2O = + 1.0000 Ti(OH)4
log_k -8.5586
-delta_H 0 # Not possible to calculate enthalpy of reaction Anatase
# Enthalpy of formation: -939.942 kJ/mol
Andalusite
Al2SiO5 +6.0000 H+ = + 1.0000 SiO2 + 2.0000 Al+++ + 3.0000 H2O
log_k 15.9445
-delta_H -235.233 kJ/mol # Calculated enthalpy of reaction Andalusite
# Enthalpy of formation: -615.866 kcal/mol
-analytic -7.1115e+001 -3.2234e-002 1.2308e+004 2.2357e+001 1.9208e+002
# -Range: 0-300
Andradite
Ca3Fe2(SiO4)3 +12.0000 H+ = + 2.0000 Fe+++ + 3.0000 Ca++ + 3.0000 SiO2 + 6.0000 H2O
log_k 33.3352
-delta_H -301.173 kJ/mol # Calculated enthalpy of reaction Andradite
# Enthalpy of formation: -1380.35 kcal/mol
-analytic 1.3884e+001 -2.3886e-002 1.5314e+004 -8.1606e+000 -4.2193e+005
# -Range: 0-300
Anglesite
PbSO4 = + 1.0000 Pb++ + 1.0000 SO4--
log_k -7.8527
-delta_H 11.255 kJ/mol # Calculated enthalpy of reaction Anglesite
# Enthalpy of formation: -219.87 kcal/mol
-analytic -1.8583e+002 -7.3849e-002 2.8528e+003 7.6936e+001 4.4570e+001
# -Range: 0-300
Anhydrite
CaSO4 = + 1.0000 Ca++ + 1.0000 SO4--
log_k -4.3064
-delta_H -18.577 kJ/mol # Calculated enthalpy of reaction Anhydrite
# Enthalpy of formation: -342.76 kcal/mol
-analytic -2.0986e+002 -7.8823e-002 5.0969e+003 8.5642e+001 7.9594e+001
# -Range: 0-300
Annite
KFe3AlSi3O10(OH)2 +10.0000 H+ = + 1.0000 Al+++ + 1.0000 K+ + 3.0000 Fe++ + 3.0000 SiO2 + 6.0000 H2O
log_k 29.4693
-delta_H -259.964 kJ/mol # Calculated enthalpy of reaction Annite
# Enthalpy of formation: -1232.19 kcal/mol
-analytic -4.0186e+001 -1.4238e-002 1.8929e+004 7.9859e+000 -8.4343e+005
# -Range: 0-300
Anorthite
CaAl2(SiO4)2 +8.0000 H+ = + 1.0000 Ca++ + 2.0000 Al+++ + 2.0000 SiO2 + 4.0000 H2O
log_k 26.5780
-delta_H -303.039 kJ/mol # Calculated enthalpy of reaction Anorthite
# Enthalpy of formation: -1007.55 kcal/mol
-analytic 3.9717e-001 -1.8751e-002 1.4897e+004 -6.3078e+000 -2.3885e+005
# -Range: 0-300
Antarcticite
CaCl2:6H2O = + 1.0000 Ca++ + 2.0000 Cl- + 6.0000 H2O
log_k 4.0933
-delta_H 0 # Not possible to calculate enthalpy of reaction Antarcticite
# Enthalpy of formation: 0 kcal/mol
Anthophyllite
Mg7Si8O22(OH)2 +14.0000 H+ = + 7.0000 Mg++ + 8.0000 H2O + 8.0000 SiO2
log_k 66.7965
-delta_H -483.486 kJ/mol # Calculated enthalpy of reaction Anthophyllite
# Enthalpy of formation: -2888.75 kcal/mol
-analytic -1.2865e+002 1.9705e-002 5.4853e+004 1.9444e+001 -3.8080e+006
# -Range: 0-300
Antigorite
# Mg48Si24O85(OH)62 +96.0000 H+ = + 34.0000 SiO2 + 48.0000 Mg++ + 79.0000 H2O
Mg48Si34O85(OH)62 +96.0000 H+ = + 34.0000 SiO2 + 48.0000 Mg++ + 79.0000 H2O
log_k 477.1943
-delta_H -3364.43 kJ/mol # Calculated enthalpy of reaction Antigorite
# Enthalpy of formation: -17070.9 kcal/mol
-analytic -8.1630e+002 -6.7780e-002 2.5998e+005 2.2029e+002 -9.3275e+006
# -Range: 0-300
Antlerite
Cu3(SO4)(OH)4 +4.0000 H+ = + 1.0000 SO4-- + 3.0000 Cu++ + 4.0000 H2O
log_k 8.7302
-delta_H 0 # Not possible to calculate enthalpy of reaction Antlerite
# Enthalpy of formation: 0 kcal/mol
Aphthitalite
NaK3(SO4)2 = + 1.0000 Na+ + 2.0000 SO4-- + 3.0000 K+
log_k -3.8878
-delta_H 0 # Not possible to calculate enthalpy of reaction Aphthitalite
# Enthalpy of formation: 0 kcal/mol
Aragonite
CaCO3 +1.0000 H+ = + 1.0000 Ca++ + 1.0000 HCO3-
log_k 1.9931
-delta_H -25.8027 kJ/mol # Calculated enthalpy of reaction Aragonite
# Enthalpy of formation: -288.531 kcal/mol
-analytic -1.4934e+002 -4.8043e-002 4.9089e+003 6.0284e+001 7.6644e+001
# -Range: 0-300
Arcanite
K2SO4 = + 1.0000 SO4-- + 2.0000 K+
log_k -1.8008
-delta_H 23.836 kJ/mol # Calculated enthalpy of reaction Arcanite
# Enthalpy of formation: -1437.78 kJ/mol
-analytic -1.6428e+002 -6.7762e-002 1.9879e+003 7.1116e+001 3.1067e+001
# -Range: 0-300
Arsenolite
As2O3 +3.0000 H2O = + 2.0000 H+ + 2.0000 H2AsO3-
log_k -19.8365
-delta_H 84.5449 kJ/mol # Calculated enthalpy of reaction Arsenolite
# Enthalpy of formation: -656.619 kJ/mol
-analytic 5.1917e+000 -1.9397e-002 -6.0894e+003 4.7458e-001 -1.0341e+002
# -Range: 0-200
Arsenopyrite
FeAsS +1.5000 H2O +0.5000 H+ = + 0.5000 AsH3 + 0.5000 H2AsO3- + 1.0000 Fe++ + 1.0000 HS-
log_k -14.4453
-delta_H 28.0187 kJ/mol # Calculated enthalpy of reaction Arsenopyrite
# Enthalpy of formation: -42.079 kJ/mol
Artinite
Mg2CO3(OH)2:3H2O +3.0000 H+ = + 1.0000 HCO3- + 2.0000 Mg++ + 5.0000 H2O
log_k 19.6560
-delta_H -130.432 kJ/mol # Calculated enthalpy of reaction Artinite
# Enthalpy of formation: -698.043 kcal/mol
-analytic -2.8614e+002 -6.7344e-002 1.5230e+004 1.1104e+002 2.3773e+002
# -Range: 0-300
As
As +1.5000 H2O +0.7500 O2 = + 1.0000 H+ + 1.0000 H2AsO3-
log_k 42.7079
-delta_H -276.937 kJ/mol # Calculated enthalpy of reaction As
# Enthalpy of formation: 0 kJ/mol
-analytic -3.4700e+001 -3.1772e-002 1.3788e+004 1.6411e+001 2.1517e+002
# -Range: 0-300
As2O5
As2O5 +3.0000 H2O = + 2.0000 H+ + 2.0000 H2AsO4-
log_k 2.1601
-delta_H -36.7345 kJ/mol # Calculated enthalpy of reaction As2O5
# Enthalpy of formation: -924.87 kJ/mol
-analytic -1.4215e+002 -6.3459e-002 4.1222e+003 6.0369e+001 6.4365e+001
# -Range: 0-300
As4O6(cubi)
As4O6 +6.0000 H2O = + 4.0000 H+ + 4.0000 H2AsO3-
log_k -39.7636
-delta_H 169.792 kJ/mol # Calculated enthalpy of reaction As4O6(cubi)
# Enthalpy of formation: -1313.94 kJ/mol
-analytic -2.6300e+002 -1.1822e-001 -4.9004e+003 1.1108e+002 -7.6389e+001
# -Range: 0-300
As4O6(mono)
As4O6 +6.0000 H2O = + 4.0000 H+ + 4.0000 H2AsO3-
log_k -40.0375
-delta_H 165.452 kJ/mol # Calculated enthalpy of reaction As4O6(mono)
# Enthalpy of formation: -1309.6 kJ/mol
-analytic 9.2518e+000 -3.8823e-002 -1.1985e+004 9.9966e-001 -2.0352e+002
# -Range: 0-200
Atacamite
Cu4Cl2(OH)6 +6.0000 H+ = + 2.0000 Cl- + 4.0000 Cu++ + 6.0000 H2O
log_k 14.2836
-delta_H -132.001 kJ/mol # Calculated enthalpy of reaction Atacamite
# Enthalpy of formation: -1654.43 kJ/mol
-analytic -2.6623e+002 -4.8121e-002 1.5315e+004 9.8395e+001 2.6016e+002
# -Range: 0-200
Au
Au +1.0000 H+ +0.2500 O2 = + 0.5000 H2O + 1.0000 Au+
log_k -7.0864
-delta_H 59.189 kJ/mol # Calculated enthalpy of reaction Au
# Enthalpy of formation: 0 kcal/mol
-analytic -7.6610e-001 -2.8520e-003 -3.0861e+003 1.9705e+000 -4.8156e+001
# -Range: 0-300
Autunite-H
H2(UO2)2(PO4)2 = + 2.0000 HPO4-- + 2.0000 UO2++
log_k -25.3372
-delta_H -31.8599 kJ/mol # Calculated enthalpy of reaction Autunite-H
# Enthalpy of formation: -4590.3 kJ/mol
-analytic -3.2179e+001 -3.8038e-002 -6.8629e+002 8.2724e+000 -1.1644e+001
# -Range: 0-200
Azurite
Cu3(CO3)2(OH)2 +4.0000 H+ = + 2.0000 H2O + 2.0000 HCO3- + 3.0000 Cu++
log_k 9.1607
-delta_H -122.298 kJ/mol # Calculated enthalpy of reaction Azurite
# Enthalpy of formation: -390.1 kcal/mol
-analytic -4.4042e+002 -1.1934e-001 1.8053e+004 1.7158e+002 2.8182e+002
# -Range: 0-300
B
B +1.5000 H2O +0.7500 O2 = + 1.0000 B(OH)3
log_k 109.5654
-delta_H -636.677 kJ/mol # Calculated enthalpy of reaction B
# Enthalpy of formation: 0 kJ/mol
-analytic 8.0471e+001 1.2577e-003 2.9653e+004 -2.8593e+001 4.6268e+002
# -Range: 0-300
B2O3
B2O3 +3.0000 H2O = + 2.0000 B(OH)3
log_k 5.5464
-delta_H -18.0548 kJ/mol # Calculated enthalpy of reaction B2O3
# Enthalpy of formation: -1273.5 kJ/mol
-analytic 9.0905e+001 5.5365e-003 -2.6629e+003 -3.1553e+001 -4.1578e+001
# -Range: 0-300
Ba
Ba +2.0000 H+ +0.5000 O2 = + 1.0000 Ba++ + 1.0000 H2O
log_k 141.2465
-delta_H -817.416 kJ/mol # Calculated enthalpy of reaction Ba
# Enthalpy of formation: 0 kJ/mol
-analytic -2.5033e+001 -1.3917e-002 4.2849e+004 1.0786e+001 6.6863e+002
# -Range: 0-300
Ba(OH)2:8H2O
Ba(OH)2:8H2O +2.0000 H+ = + 1.0000 Ba++ + 10.0000 H2O
log_k 24.4911
-delta_H -55.4363 kJ/mol # Calculated enthalpy of reaction Ba(OH)2:8H2O
# Enthalpy of formation: -3340.59 kJ/mol
-analytic -2.3888e+002 -1.5791e-003 1.4097e+004 8.7518e+001 2.3947e+002
# -Range: 0-200
Ba2Si3O8
Ba2Si3O8 +4.0000 H+ = + 2.0000 Ba++ + 2.0000 H2O + 3.0000 SiO2
log_k 23.3284
-delta_H -95.3325 kJ/mol # Calculated enthalpy of reaction Ba2Si3O8
# Enthalpy of formation: -4184.73 kJ/mol
-analytic -8.7226e+001 9.3125e-003 2.3147e+004 2.2012e+001 -2.1714e+006
# -Range: 0-300
Ba2SiO4
Ba2SiO4 +4.0000 H+ = + 1.0000 SiO2 + 2.0000 Ba++ + 2.0000 H2O
log_k 44.5930
-delta_H -237.206 kJ/mol # Calculated enthalpy of reaction Ba2SiO4
# Enthalpy of formation: -2287.46 kJ/mol
-analytic -7.0350e+000 -5.1744e-003 1.4786e+004 3.1091e+000 -3.6972e+005
# -Range: 0-300
Ba2U2O7
Ba2U2O7 +6.0000 H+ = + 2.0000 Ba++ + 2.0000 UO2+ + 3.0000 H2O
log_k 36.4635
-delta_H -243.057 kJ/mol # Calculated enthalpy of reaction Ba2U2O7
# Enthalpy of formation: -3740 kJ/mol
-analytic -9.2562e+001 5.3866e-003 1.6852e+004 2.8647e+001 2.8621e+002
# -Range: 0-200
Ba3UO6
Ba3UO6 +8.0000 H+ = + 1.0000 UO2++ + 3.0000 Ba++ + 4.0000 H2O
log_k 94.3709
-delta_H -564.885 kJ/mol # Calculated enthalpy of reaction Ba3UO6
# Enthalpy of formation: -3210.4 kJ/mol
-analytic -1.3001e+002 -1.7395e-002 3.3977e+004 4.6715e+001 5.7703e+002
# -Range: 0-200
BaBr2
BaBr2 = + 1.0000 Ba++ + 2.0000 Br-
log_k 5.6226
-delta_H -23.3887 kJ/mol # Calculated enthalpy of reaction BaBr2
# Enthalpy of formation: -757.262 kJ/mol
-analytic -1.7689e+002 -7.1918e-002 4.7187e+003 7.6010e+001 7.3683e+001
# -Range: 0-300
BaBr2:2H2O
BaBr2:2H2O = + 1.0000 Ba++ + 2.0000 Br- + 2.0000 H2O
log_k 2.2523
-delta_H 13.7736 kJ/mol # Calculated enthalpy of reaction BaBr2:2H2O
# Enthalpy of formation: -1366.1 kJ/mol
-analytic -1.5506e+001 -1.6281e-002 -8.5727e+002 1.0296e+001 -1.4552e+001
# -Range: 0-200
BaCl2
BaCl2 = + 1.0000 Ba++ + 2.0000 Cl-
log_k 2.2707
-delta_H -13.1563 kJ/mol # Calculated enthalpy of reaction BaCl2
# Enthalpy of formation: -858.647 kJ/mol
-analytic -2.0393e+002 -7.8925e-002 4.8846e+003 8.6204e+001 7.6280e+001
# -Range: 0-300
BaCl2:2H2O
BaCl2:2H2O = + 1.0000 Ba++ + 2.0000 Cl- + 2.0000 H2O
log_k 0.2459
-delta_H 16.558 kJ/mol # Calculated enthalpy of reaction BaCl2:2H2O
# Enthalpy of formation: -1460.04 kJ/mol
-analytic -2.0350e+002 -7.3577e-002 3.7914e+003 8.6051e+001 5.9221e+001
# -Range: 0-300
BaCl2:H2O
BaCl2:H2O = + 1.0000 Ba++ + 1.0000 H2O + 2.0000 Cl-
log_k 0.8606
-delta_H 2.89433 kJ/mol # Calculated enthalpy of reaction BaCl2:H2O
# Enthalpy of formation: -1160.54 kJ/mol
-analytic -1.9572e+002 -7.3938e-002 4.0553e+003 8.2842e+001 6.3336e+001
# -Range: 0-300
BaCrO4
BaCrO4 = + 1.0000 Ba++ + 1.0000 CrO4--
log_k -9.9322
-delta_H 25.9115 kJ/mol # Calculated enthalpy of reaction BaCrO4
# Enthalpy of formation: -345.293 kcal/mol
-analytic 2.3142e+001 -1.6617e-002 -3.6883e+003 -6.3687e+000 -6.2640e+001
# -Range: 0-200
BaHPO4
BaHPO4 = + 1.0000 Ba++ + 1.0000 HPO4--
log_k -7.4000
-delta_H 0 # Not possible to calculate enthalpy of reaction BaHPO4
# Enthalpy of formation: 0 kcal/mol
BaI2
BaI2 = + 1.0000 Ba++ + 2.0000 I-
log_k 11.0759
-delta_H -46.0408 kJ/mol # Calculated enthalpy of reaction BaI2
# Enthalpy of formation: -605.408 kJ/mol
-analytic -1.7511e+002 -7.2206e-002 5.8696e+003 7.5974e+001 9.1641e+001
# -Range: 0-300
BaMnO4
BaMnO4 = + 1.0000 Ba++ + 1.0000 MnO4--
log_k -10.0900
-delta_H 0 # Not possible to calculate enthalpy of reaction BaMnO4
# Enthalpy of formation: 0 kcal/mol
BaO
BaO +2.0000 H+ = + 1.0000 Ba++ + 1.0000 H2O
log_k 47.8036
-delta_H -270.184 kJ/mol # Calculated enthalpy of reaction BaO
# Enthalpy of formation: -553.298 kJ/mol
-analytic -7.3273e+001 -1.7149e-002 1.6811e+004 2.8560e+001 -7.7510e+004
# -Range: 0-300
BaS
BaS +1.0000 H+ = + 1.0000 Ba++ + 1.0000 HS-
log_k 16.2606
-delta_H -92.9004 kJ/mol # Calculated enthalpy of reaction BaS
# Enthalpy of formation: -460.852 kJ/mol
-analytic -1.1819e+002 -4.3420e-002 7.4296e+003 4.9489e+001 1.1597e+002
# -Range: 0-300
BaSeO3
BaSeO3 = + 1.0000 Ba++ + 1.0000 SeO3--
log_k -6.5615
-delta_H -5.5658 kJ/mol # Calculated enthalpy of reaction BaSeO3
# Enthalpy of formation: -1041.27 kJ/mol
-analytic 2.9742e+001 -1.7073e-002 -2.4532e+003 -9.2936e+000 -4.1669e+001
# -Range: 0-200
BaSeO4
BaSeO4 = + 1.0000 Ba++ + 1.0000 SeO4--
log_k -7.4468
-delta_H 8.9782 kJ/mol # Calculated enthalpy of reaction BaSeO4
# Enthalpy of formation: -1145.77 kJ/mol
-analytic 2.4274e+001 -1.6289e-002 -2.8520e+003 -6.9949e+000 -4.8439e+001
# -Range: 0-200
BaSiF6
BaSiF6 +2.0000 H2O = + 1.0000 Ba++ + 1.0000 SiO2 + 4.0000 H+ + 6.0000 F-
log_k -32.1771
-delta_H 95.2555 kJ/mol # Calculated enthalpy of reaction BaSiF6
# Enthalpy of formation: -2951.01 kJ/mol
-analytic -6.4766e+000 -3.8410e-002 0.0000e+000 0.0000e+000 -1.2701e+006
# -Range: 0-200
BaU2O7
BaU2O7 +6.0000 H+ = + 1.0000 Ba++ + 2.0000 UO2++ + 3.0000 H2O
log_k 21.9576
-delta_H -195.959 kJ/mol # Calculated enthalpy of reaction BaU2O7
# Enthalpy of formation: -3237.2 kJ/mol
-analytic -1.2254e+002 -1.0941e-002 1.4452e+004 4.0125e+001 2.4546e+002
# -Range: 0-200
BaUO4
BaUO4 +4.0000 H+ = + 1.0000 Ba++ + 1.0000 UO2++ + 2.0000 H2O
log_k 18.2007
-delta_H -134.521 kJ/mol # Calculated enthalpy of reaction BaUO4
# Enthalpy of formation: -1993.8 kJ/mol
-analytic -6.7113e+001 -1.6340e-002 8.7592e+003 2.4571e+001 1.3670e+002
# -Range: 0-300
BaZrO3
BaZrO3 +4.0000 H+ = + 1.0000 Ba++ + 1.0000 H2O + 1.0000 Zr(OH)2++
log_k -94.4716
-delta_H 505.159 kJ/mol # Calculated enthalpy of reaction BaZrO3
# Enthalpy of formation: -578.27 kcal/mol
-analytic -5.3606e+001 -1.0096e-002 -2.4894e+004 1.8446e+001 -4.2271e+002
# -Range: 0-200
Baddeleyite
ZrO2 +2.0000 H+ = + 1.0000 Zr(OH)2++
log_k -7.9405
-delta_H 9.72007 kJ/mol # Calculated enthalpy of reaction Baddeleyite
# Enthalpy of formation: -1100.56 kJ/mol
-analytic -2.5188e-001 -4.6374e-003 -1.0635e+003 -1.1055e+000 -1.6595e+001
# -Range: 0-300
Barite
BaSO4 = + 1.0000 Ba++ + 1.0000 SO4--
log_k -9.9711
-delta_H 25.9408 kJ/mol # Calculated enthalpy of reaction Barite
# Enthalpy of formation: -352.1 kcal/mol
-analytic -1.8747e+002 -7.5521e-002 2.0790e+003 7.7998e+001 3.2497e+001
# -Range: 0-300
Barytocalcite
BaCa(CO3)2 +2.0000 H+ = + 1.0000 Ba++ + 1.0000 Ca++ + 2.0000 HCO3-
log_k 2.7420
-delta_H 0 # Not possible to calculate enthalpy of reaction Barytocalcite
# Enthalpy of formation: 0 kcal/mol
Bassanite
CaSO4:0.5H2O = + 0.5000 H2O + 1.0000 Ca++ + 1.0000 SO4--
log_k -3.6615
-delta_H -18.711 kJ/mol # Calculated enthalpy of reaction Bassanite
# Enthalpy of formation: -1576.89 kJ/mol
-analytic -2.2010e+002 -8.0230e-002 5.5092e+003 8.9651e+001 8.6031e+001
# -Range: 0-300
Bassetite
Fe(UO2)2(PO4)2 +2.0000 H+ = + 1.0000 Fe++ + 2.0000 HPO4-- + 2.0000 UO2++
log_k -17.7240
-delta_H -114.841 kJ/mol # Calculated enthalpy of reaction Bassetite
# Enthalpy of formation: -1099.33 kcal/mol
-analytic -5.7788e+001 -4.5400e-002 4.0119e+003 1.6216e+001 6.8147e+001
# -Range: 0-200
Be
Be +2.0000 H+ +0.5000 O2 = + 1.0000 Be++ + 1.0000 H2O
log_k 104.2077
-delta_H -662.608 kJ/mol # Calculated enthalpy of reaction Be
# Enthalpy of formation: 0 kJ/mol
-analytic -9.3960e+001 -2.4749e-002 3.6714e+004 3.3295e+001 5.7291e+002
# -Range: 0-300
Be13U
Be13U +30.0000 H+ +7.5000 O2 = + 1.0000 U++++ + 13.0000 Be++ + 15.0000 H2O
log_k 1504.5350
-delta_H -9601.04 kJ/mol # Calculated enthalpy of reaction Be13U
# Enthalpy of formation: -163.6 kJ/mol
-analytic -1.2388e+003 -3.2848e-001 5.2816e+005 4.3222e+002 8.2419e+003
# -Range: 0-300
Beidellite-Ca
Ca.165Al2.33Si3.67O10(OH)2 +7.3200 H+ = + 0.1650 Ca++ + 2.3300 Al+++ + 3.6700 SiO2 + 4.6600 H2O
log_k 5.5914
-delta_H -162.403 kJ/mol # Calculated enthalpy of reaction Beidellite-Ca
# Enthalpy of formation: -1370.66 kcal/mol
-analytic 2.3887e+001 4.4178e-003 1.5296e+004 -2.2343e+001 -1.4025e+006
# -Range: 0-300
Beidellite-Cs
Cs.33Si3.67Al2.33O10(OH)2 +7.3200 H+ = + 0.3300 Cs+ + 2.3300 Al+++ + 3.6700 SiO2 + 4.6600 H2O
log_k 5.1541
-delta_H -149.851 kJ/mol # Calculated enthalpy of reaction Beidellite-Cs
# Enthalpy of formation: -1372.59 kcal/mol
-analytic 2.1244e+001 2.1705e-003 1.4504e+004 -2.0250e+001 -1.3712e+006
# -Range: 0-300
Beidellite-H
H.33Al2.33Si3.67O10(OH)2 +6.9900 H+ = + 2.3300 Al+++ + 3.6700 SiO2 + 4.6600 H2O
log_k 4.6335
-delta_H -154.65 kJ/mol # Calculated enthalpy of reaction Beidellite-H
# Enthalpy of formation: -1351.1 kcal/mol
-analytic 5.4070e+000 3.4064e-003 1.6284e+004 -1.6028e+001 -1.5014e+006
# -Range: 0-300
Beidellite-K
K.33Al2.33Si3.67O10(OH)2 +7.3200 H+ = + 0.3300 K+ + 2.3300 Al+++ + 3.6700 SiO2 + 4.6600 H2O
log_k 5.3088
-delta_H -150.834 kJ/mol # Calculated enthalpy of reaction Beidellite-K
# Enthalpy of formation: -1371.9 kcal/mol
-analytic 1.0792e+001 3.4419e-003 1.5760e+004 -1.7333e+001 -1.4779e+006
# -Range: 0-300
Beidellite-Mg
Mg.165Al2.33Si3.67O10(OH)2 +7.3200 H+ = + 0.1650 Mg++ + 2.3300 Al+++ + 3.6700 SiO2 + 4.6600 H2O
log_k 5.5537
-delta_H -165.455 kJ/mol # Calculated enthalpy of reaction Beidellite-Mg
# Enthalpy of formation: -1366.89 kcal/mol
-analytic 1.3375e+001 3.0420e-003 1.5947e+004 -1.8728e+001 -1.4242e+006
# -Range: 0-300
Beidellite-Na
Na.33Al2.33Si3.67O10(OH)2 +7.3200 H+ = + 0.3300 Na+ + 2.3300 Al+++ + 3.6700 SiO2 + 4.6600 H2O
log_k 5.6473
-delta_H -155.846 kJ/mol # Calculated enthalpy of reaction Beidellite-Na
# Enthalpy of formation: -1369.76 kcal/mol
-analytic 1.1504e+001 3.9871e-003 1.5818e+004 -1.7762e+001 -1.4485e+006
# -Range: 0-300
Berlinite
AlPO4 +1.0000 H+ = + 1.0000 Al+++ + 1.0000 HPO4--
log_k -7.2087
-delta_H -96.6313 kJ/mol # Calculated enthalpy of reaction Berlinite
# Enthalpy of formation: -1733.85 kJ/mol
-analytic -2.8134e+002 -9.9933e-002 1.0308e+004 1.0883e+002 1.6094e+002
# -Range: 0-300
Berndtite
SnS2 = + 1.0000 S2-- + 1.0000 Sn++
log_k -34.5393
-delta_H 0 # Not possible to calculate enthalpy of reaction Berndtite
# Enthalpy of formation: -36.7 kcal/mol
-analytic -2.0311e+002 -7.6462e-002 -4.9879e+003 8.4082e+001 -7.7772e+001
# -Range: 0-300
Bieberite
CoSO4:7H2O = + 1.0000 Co++ + 1.0000 SO4-- + 7.0000 H2O
log_k -2.5051
-delta_H 11.3885 kJ/mol # Calculated enthalpy of reaction Bieberite
# Enthalpy of formation: -2980.02 kJ/mol
-analytic -2.6405e+002 -7.2497e-002 6.6673e+003 1.0538e+002 1.0411e+002
# -Range: 0-300
Birnessite
Mn8O14:5H2O +4.0000 H+ = + 3.0000 MnO4-- + 5.0000 Mn++ + 7.0000 H2O
log_k -85.5463
-delta_H 0 # Not possible to calculate enthalpy of reaction Birnessite
# Enthalpy of formation: 0 kcal/mol
Bischofite
MgCl2:6H2O = + 1.0000 Mg++ + 2.0000 Cl- + 6.0000 H2O
log_k 4.3923
-delta_H 0 # Not possible to calculate enthalpy of reaction Bischofite
# Enthalpy of formation: 0 kcal/mol
Bixbyite
Mn2O3 +6.0000 H+ = + 2.0000 Mn+++ + 3.0000 H2O
log_k -0.9655
-delta_H -190.545 kJ/mol # Calculated enthalpy of reaction Bixbyite
# Enthalpy of formation: -958.971 kJ/mol
-analytic -1.1600e+002 -2.8056e-003 1.3418e+004 2.8639e+001 2.0941e+002
# -Range: 0-300
Bloedite
Na2Mg(SO4)2:4H2O = + 1.0000 Mg++ + 2.0000 Na+ + 2.0000 SO4-- + 4.0000 H2O
log_k -2.4777
-delta_H 0 # Not possible to calculate enthalpy of reaction Bloedite
# Enthalpy of formation: 0 kcal/mol
Boehmite
AlO2H +3.0000 H+ = + 1.0000 Al+++ + 2.0000 H2O
log_k 7.5642
-delta_H -113.282 kJ/mol # Calculated enthalpy of reaction Boehmite
# Enthalpy of formation: -238.24 kcal/mol
-analytic -1.2196e+002 -3.1138e-002 8.8643e+003 4.4075e+001 1.3835e+002
# -Range: 0-300
Boltwoodite
K(H3O)(UO2)SiO4 +3.0000 H+ = + 1.0000 K+ + 1.0000 SiO2 + 1.0000 UO2++ + 3.0000 H2O
log_k 14.8857
-delta_H 0 # Not possible to calculate enthalpy of reaction Boltwoodite
# Enthalpy of formation: 0 kcal/mol
Boltwoodite-Na
Na.7K.3(H3O)(UO2)SiO4:H2O +3.0000 H+ = + 0.3000 K+ + 0.7000 Na+ + 1.0000 SiO2 + 1.0000 UO2++ + 4.0000 H2O
log_k 14.5834
-delta_H 0 # Not possible to calculate enthalpy of reaction Boltwoodite-Na
# Enthalpy of formation: 0 kcal/mol
Borax
Na2(B4O5(OH)4):8H2O +2.0000 H+ = + 2.0000 Na+ + 4.0000 B(OH)3 + 5.0000 H2O
log_k 12.0395
-delta_H 80.5145 kJ/mol # Calculated enthalpy of reaction Borax
# Enthalpy of formation: -6288.44 kJ/mol
-analytic 7.8374e+001 1.9328e-002 -5.3279e+003 -2.1914e+001 -8.3160e+001
# -Range: 0-300
Boric_acid
B(OH)3 = + 1.0000 B(OH)3
log_k -0.1583
-delta_H 20.2651 kJ/mol # Calculated enthalpy of reaction Boric_acid
# Enthalpy of formation: -1094.8 kJ/mol
-analytic 3.9122e+001 6.4058e-003 -2.2525e+003 -1.3592e+001 -3.5160e+001
# -Range: 0-300
Bornite
Cu5FeS4 +4.0000 H+ = + 1.0000 Cu++ + 1.0000 Fe++ + 4.0000 Cu+ + 4.0000 HS-
log_k -102.4369
-delta_H 530.113 kJ/mol # Calculated enthalpy of reaction Bornite
# Enthalpy of formation: -79.922 kcal/mol
-analytic -7.0495e+002 -2.0082e-001 -9.1376e+003 2.8004e+002 -1.4238e+002
# -Range: 0-300
Brezinaite
Cr3S4 +4.0000 H+ = + 1.0000 Cr++ + 2.0000 Cr+++ + 4.0000 HS-
log_k 2.7883
-delta_H -216.731 kJ/mol # Calculated enthalpy of reaction Brezinaite
# Enthalpy of formation: -111.9 kcal/mol
-analytic -7.0528e+001 -3.6568e-002 1.0598e+004 1.9665e+001 1.8000e+002
# -Range: 0-200
Brochantite
Cu4(SO4)(OH)6 +6.0000 H+ = + 1.0000 SO4-- + 4.0000 Cu++ + 6.0000 H2O
log_k 15.4363
-delta_H -163.158 kJ/mol # Calculated enthalpy of reaction Brochantite
# Enthalpy of formation: -2198.72 kJ/mol
-analytic -2.3609e+002 -3.9046e-002 1.5970e+004 8.4701e+001 2.7127e+002
# -Range: 0-200
Bromellite
BeO +2.0000 H+ = + 1.0000 Be++ + 1.0000 H2O
log_k 1.1309
-delta_H -59.2743 kJ/mol # Calculated enthalpy of reaction Bromellite
# Enthalpy of formation: -609.4 kJ/mol
-analytic 1.4790e+002 -4.6004e-001 -3.2577e+004 4.0273e+001 -5.0837e+002
# -Range: 0-300
Brucite
Mg(OH)2 +2.0000 H+ = + 1.0000 Mg++ + 2.0000 H2O
log_k 16.2980
-delta_H -111.34 kJ/mol # Calculated enthalpy of reaction Brucite
# Enthalpy of formation: -221.39 kcal/mol
-analytic -1.0280e+002 -1.9759e-002 9.0180e+003 3.8282e+001 1.4075e+002
# -Range: 0-300
Brushite
CaHPO4:2H2O = + 1.0000 Ca++ + 1.0000 HPO4-- + 2.0000 H2O
log_k 6.5500
-delta_H 0 # Not possible to calculate enthalpy of reaction Brushite
# Enthalpy of formation: 0 kcal/mol
Bunsenite
NiO +2.0000 H+ = + 1.0000 H2O + 1.0000 Ni++
log_k 12.4719
-delta_H -100.069 kJ/mol # Calculated enthalpy of reaction Bunsenite
# Enthalpy of formation: -57.3 kcal/mol
-analytic -8.1664e+001 -1.9796e-002 7.4064e+003 3.0385e+001 1.1559e+002
# -Range: 0-300
Burkeite
Na6CO3(SO4)2 +1.0000 H+ = + 1.0000 HCO3- + 2.0000 SO4-- + 6.0000 Na+
log_k 9.4866
-delta_H 0 # Not possible to calculate enthalpy of reaction Burkeite
# Enthalpy of formation: 0 kcal/mol
C
C +1.0000 H2O +1.0000 O2 = + 1.0000 H+ + 1.0000 HCO3-
log_k 64.1735
-delta_H -391.961 kJ/mol # Calculated enthalpy of reaction C
# Enthalpy of formation: 0 kcal/mol
-analytic -3.5556e+001 -3.3691e-002 1.9774e+004 1.7548e+001 3.0856e+002
# -Range: 0-300
Ca
Ca +2.0000 H+ +0.5000 O2 = + 1.0000 Ca++ + 1.0000 H2O
log_k 139.8465
-delta_H -822.855 kJ/mol # Calculated enthalpy of reaction Ca
# Enthalpy of formation: 0 kJ/mol
-analytic -1.1328e+002 -2.6554e-002 4.7638e+004 4.1989e+001 -2.3545e+005
# -Range: 0-300
Ca-Al_Pyroxene
CaAl2SiO6 +8.0000 H+ = + 1.0000 Ca++ + 1.0000 SiO2 + 2.0000 Al+++ + 4.0000 H2O
log_k 35.9759
-delta_H -361.548 kJ/mol # Calculated enthalpy of reaction Ca-Al_Pyroxene
# Enthalpy of formation: -783.793 kcal/mol
-analytic -1.4664e+002 -5.0409e-002 2.1045e+004 5.1318e+001 3.2843e+002
# -Range: 0-300
Ca2Al2O5:8H2O
Ca2Al2O5:8H2O +10.0000 H+ = + 2.0000 Al+++ + 2.0000 Ca++ + 13.0000 H2O
log_k 59.5687
-delta_H 0 # Not possible to calculate enthalpy of reaction Ca2Al2O5:8H2O
# Enthalpy of formation: 0 kcal/mol
Ca2Cl2(OH)2:H2O
Ca2Cl2(OH)2:H2O +2.0000 H+ = + 2.0000 Ca++ + 2.0000 Cl- + 3.0000 H2O
log_k 26.2901
-delta_H 0 # Not possible to calculate enthalpy of reaction Ca2Cl2(OH)2:H2O
# Enthalpy of formation: 0 kcal/mol
Ca2V2O7
Ca2V2O7 +1.0000 H2O = + 2.0000 Ca++ + 2.0000 H+ + 2.0000 VO4---
log_k -39.7129
-delta_H 0 # Not possible to calculate enthalpy of reaction Ca2V2O7
# Enthalpy of formation: -3083.46 kJ/mol
Ca3(AsO4)2
Ca3(AsO4)2 +4.0000 H+ = + 2.0000 H2AsO4- + 3.0000 Ca++
log_k 17.8160
-delta_H -149.956 kJ/mol # Calculated enthalpy of reaction Ca3(AsO4)2
# Enthalpy of formation: -3298.41 kJ/mol
-analytic -1.4011e+002 -4.2945e-002 1.0981e+004 5.4107e+001 1.8652e+002
# -Range: 0-200
Ca3Al2O6
Ca3Al2O6 +12.0000 H+ = + 2.0000 Al+++ + 3.0000 Ca++ + 6.0000 H2O
log_k 113.0460
-delta_H -833.336 kJ/mol # Calculated enthalpy of reaction Ca3Al2O6
# Enthalpy of formation: -857.492 kcal/mol
-analytic -2.7163e+002 -5.2897e-002 5.0815e+004 9.2946e+001 8.6300e+002
# -Range: 0-200
Ca3V2O8
Ca3V2O8 = + 2.0000 VO4--- + 3.0000 Ca++
log_k -18.3234
-delta_H 0 # Not possible to calculate enthalpy of reaction Ca3V2O8
# Enthalpy of formation: -3778.1 kJ/mol
Ca4Al2Fe2O10
Ca4Al2Fe2O10 +20.0000 H+ = + 2.0000 Al+++ + 2.0000 Fe+++ + 4.0000 Ca++ + 10.0000 H2O
log_k 140.5050
-delta_H -1139.86 kJ/mol # Calculated enthalpy of reaction Ca4Al2Fe2O10
# Enthalpy of formation: -1211 kcal/mol
-analytic -4.1808e+002 -8.2787e-002 7.0288e+004 1.4043e+002 1.1937e+003
# -Range: 0-200
Ca4Al2O7:13H2O
Ca4Al2O7:13H2O +14.0000 H+ = + 2.0000 Al+++ + 4.0000 Ca++ + 20.0000 H2O
log_k 107.2537
-delta_H 0 # Not possible to calculate enthalpy of reaction Ca4Al2O7:13H2O
# Enthalpy of formation: 0 kcal/mol
Ca4Al2O7:19H2O
Ca4Al2O7:19H2O +14.0000 H+ = + 2.0000 Al+++ + 4.0000 Ca++ + 26.0000 H2O
log_k 103.6812
-delta_H 0 # Not possible to calculate enthalpy of reaction Ca4Al2O7:19H2O
# Enthalpy of formation: 0 kcal/mol
Ca4Cl2(OH)6:13H2O
Ca4Cl2(OH)6:13H2O +6.0000 H+ = + 2.0000 Cl- + 4.0000 Ca++ + 19.0000 H2O
log_k 68.3283
-delta_H 0 # Not possible to calculate enthalpy of reaction Ca4Cl2(OH)6:13H2O
# Enthalpy of formation: 0 kcal/mol
CaAl2O4
CaAl2O4 +8.0000 H+ = + 1.0000 Ca++ + 2.0000 Al+++ + 4.0000 H2O
log_k 46.9541
-delta_H -436.952 kJ/mol # Calculated enthalpy of reaction CaAl2O4
# Enthalpy of formation: -555.996 kcal/mol
-analytic -3.0378e+002 -7.9356e-002 3.0096e+004 1.1049e+002 4.6971e+002
# -Range: 0-300
CaAl2O4:10H2O
CaAl2O4:10H2O +8.0000 H+ = + 1.0000 Ca++ + 2.0000 Al+++ + 14.0000 H2O
log_k 37.9946
-delta_H 0 # Not possible to calculate enthalpy of reaction CaAl2O4:10H2O
# Enthalpy of formation: 0 kcal/mol
CaAl4O7
CaAl4O7 +14.0000 H+ = + 1.0000 Ca++ + 4.0000 Al+++ + 7.0000 H2O
log_k 68.6138
-delta_H -718.464 kJ/mol # Calculated enthalpy of reaction CaAl4O7
# Enthalpy of formation: -951.026 kcal/mol
-analytic -3.1044e+002 -6.7078e-002 4.4566e+004 1.0085e+002 7.5689e+002
# -Range: 0-200
CaSO4:0.5H2O(beta)
CaSO4:0.5H2O = + 0.5000 H2O + 1.0000 Ca++ + 1.0000 SO4--
log_k -3.4934
-delta_H -20.804 kJ/mol # Calculated enthalpy of reaction CaSO4:0.5H2O(beta)
# Enthalpy of formation: -1574.8 kJ/mol
-analytic -2.3054e+002 -8.2832e-002 5.9132e+003 9.3705e+001 9.2338e+001
# -Range: 0-300
CaSeO3:2H2O
CaSeO3:2H2O = + 1.0000 Ca++ + 1.0000 SeO3-- + 2.0000 H2O
log_k -4.6213
-delta_H -14.1963 kJ/mol # Calculated enthalpy of reaction CaSeO3:2H2O
# Enthalpy of formation: -384.741 kcal/mol
-analytic -4.1771e+001 -2.0735e-002 9.7870e+002 1.6180e+001 1.6634e+001
# -Range: 0-200
CaSeO4
CaSeO4 = + 1.0000 Ca++ + 1.0000 SeO4--
log_k -3.0900
-delta_H 0 # Not possible to calculate enthalpy of reaction CaSeO4
# Enthalpy of formation: 0 kcal/mol
CaUO4
CaUO4 +4.0000 H+ = + 1.0000 Ca++ + 1.0000 UO2++ + 2.0000 H2O
log_k 15.9420
-delta_H -131.46 kJ/mol # Calculated enthalpy of reaction CaUO4
# Enthalpy of formation: -2002.3 kJ/mol
-analytic -8.7902e+001 -1.9810e-002 9.2354e+003 3.1832e+001 1.4414e+002
# -Range: 0-300
CaV2O6
CaV2O6 +2.0000 H2O = + 1.0000 Ca++ + 2.0000 VO4--- + 4.0000 H+
log_k -51.3617
-delta_H 0 # Not possible to calculate enthalpy of reaction CaV2O6
# Enthalpy of formation: -2329.34 kJ/mol
CaZrO3
CaZrO3 +4.0000 H+ = + 1.0000 Ca++ + 1.0000 H2O + 1.0000 Zr(OH)2++
log_k -148.5015
-delta_H 801.282 kJ/mol # Calculated enthalpy of reaction CaZrO3
# Enthalpy of formation: -650.345 kcal/mol
-analytic -7.7908e+001 -1.4388e-002 -3.9635e+004 2.6932e+001 -6.7303e+002
# -Range: 0-200
Cadmoselite
CdSe = + 1.0000 Cd++ + 1.0000 Se--
log_k -33.8428
-delta_H 0 # Not possible to calculate enthalpy of reaction Cadmoselite
# Enthalpy of formation: -34.6 kcal/mol
-analytic -5.3432e+001 -1.3973e-002 -5.8989e+003 1.7591e+001 -9.2031e+001
# -Range: 0-300
Calcite
CaCO3 +1.0000 H+ = + 1.0000 Ca++ + 1.0000 HCO3-
log_k 1.8487
-delta_H -25.7149 kJ/mol # Calculated enthalpy of reaction Calcite
# Enthalpy of formation: -288.552 kcal/mol
-analytic -1.4978e+002 -4.8370e-002 4.8974e+003 6.0458e+001 7.6464e+001
# -Range: 0-300
Calomel
Hg2Cl2 = + 1.0000 Hg2++ + 2.0000 Cl-
log_k -17.8241
-delta_H 98.0267 kJ/mol # Calculated enthalpy of reaction Calomel
# Enthalpy of formation: -265.37 kJ/mol
-analytic -4.8868e+001 -2.5540e-002 -2.8439e+003 1.9475e+001 -4.8277e+001
# -Range: 0-200
Carnallite
KMgCl3:6H2O = + 1.0000 K+ + 1.0000 Mg++ + 3.0000 Cl- + 6.0000 H2O
log_k 4.2721
-delta_H 0 # Not possible to calculate enthalpy of reaction Carnallite
# Enthalpy of formation: 0 kcal/mol
Carnotite
K2(UO2)2(VO4)2 = + 2.0000 K+ + 2.0000 UO2++ + 2.0000 VO4---
log_k -56.3811
-delta_H 0 # Not possible to calculate enthalpy of reaction Carnotite
# Enthalpy of formation: -1173.9 kJ/mol
Cassiterite
SnO2 +2.0000 H+ = + 0.5000 O2 + 1.0000 H2O + 1.0000 Sn++
log_k -46.1203
-delta_H 280.048 kJ/mol # Calculated enthalpy of reaction Cassiterite
# Enthalpy of formation: -138.8 kcal/mol
-analytic -8.9264e+001 -1.5743e-002 -1.1497e+004 3.4917e+001 -1.7937e+002
# -Range: 0-300
Cattierite
CoS2 = + 1.0000 Co++ + 1.0000 S2--
log_k -29.9067
-delta_H 0 # Not possible to calculate enthalpy of reaction Cattierite
# Enthalpy of formation: -36.589 kcal/mol
-analytic -2.1970e+002 -7.8585e-002 -1.9592e+003 8.8809e+001 -3.0507e+001
# -Range: 0-300
Cd
Cd +2.0000 H+ +0.5000 O2 = + 1.0000 Cd++ + 1.0000 H2O
log_k 56.6062
-delta_H -355.669 kJ/mol # Calculated enthalpy of reaction Cd
# Enthalpy of formation: 0 kJ/mol
-analytic -7.2027e+001 -2.0250e-002 2.0474e+004 2.6814e+001 -3.2348e+004
# -Range: 0-300
Cd(BO2)2
Cd(BO2)2 +2.0000 H+ +2.0000 H2O = + 1.0000 Cd++ + 2.0000 B(OH)3
log_k 9.8299
-delta_H 0 # Not possible to calculate enthalpy of reaction Cd(BO2)2
# Enthalpy of formation: 0 kcal/mol
Cd(IO3)2
Cd(IO3)2 = + 1.0000 Cd++ + 2.0000 IO3-
log_k -7.5848
-delta_H 0 # Not possible to calculate enthalpy of reaction Cd(IO3)2
# Enthalpy of formation: 0 kcal/mol
Cd(OH)2
Cd(OH)2 +2.0000 H+ = + 1.0000 Cd++ + 2.0000 H2O
log_k 13.7382
-delta_H -87.0244 kJ/mol # Calculated enthalpy of reaction Cd(OH)2
# Enthalpy of formation: -560.55 kJ/mol
-analytic -7.7001e+001 -6.9251e-003 7.4684e+003 2.7380e+001 1.2685e+002
# -Range: 0-200
Cd(OH)Cl
Cd(OH)Cl +1.0000 H+ = + 1.0000 Cd++ + 1.0000 Cl- + 1.0000 H2O
log_k 3.5435
-delta_H -30.3888 kJ/mol # Calculated enthalpy of reaction Cd(OH)Cl
# Enthalpy of formation: -498.427 kJ/mol
-analytic -4.5477e+001 -1.5809e-002 2.5333e+003 1.8279e+001 4.3035e+001
# -Range: 0-200
Cd3(AsO4)2
Cd3(AsO4)2 +4.0000 H+ = + 2.0000 H2AsO4- + 3.0000 Cd++
log_k 4.0625
-delta_H 0 # Not possible to calculate enthalpy of reaction Cd3(AsO4)2
# Enthalpy of formation: 0 kcal/mol
Cd3(PO4)2
Cd3(PO4)2 +2.0000 H+ = + 2.0000 HPO4-- + 3.0000 Cd++
log_k -7.8943
-delta_H 0 # Not possible to calculate enthalpy of reaction Cd3(PO4)2
# Enthalpy of formation: 0 kcal/mol
Cd3(SO4)(OH)4
Cd3(SO4)(OH)4 +4.0000 H+ = + 1.0000 SO4-- + 3.0000 Cd++ + 4.0000 H2O
log_k 22.5735
-delta_H 0 # Not possible to calculate enthalpy of reaction Cd3(SO4)(OH)4
# Enthalpy of formation: 0 kcal/mol
Cd3(SO4)2(OH)2
Cd3(SO4)2(OH)2 +2.0000 H+ = + 2.0000 H2O + 2.0000 SO4-- + 3.0000 Cd++
log_k 6.7180
-delta_H 0 # Not possible to calculate enthalpy of reaction Cd3(SO4)2(OH)2
# Enthalpy of formation: 0 kcal/mol
CdBr2
CdBr2 = + 1.0000 Cd++ + 2.0000 Br-
log_k -1.8470
-delta_H -2.67548 kJ/mol # Calculated enthalpy of reaction CdBr2
# Enthalpy of formation: -316.229 kJ/mol
-analytic 1.3056e+000 -2.0628e-002 -1.3318e+003 3.0126e+000 -2.2616e+001
# -Range: 0-200
CdBr2:4H2O
CdBr2:4H2O = + 1.0000 Cd++ + 2.0000 Br- + 4.0000 H2O
log_k -2.3378
-delta_H 30.2812 kJ/mol # Calculated enthalpy of reaction CdBr2:4H2O
# Enthalpy of formation: -1492.54 kJ/mol
-analytic -1.0038e+002 -2.1045e-002 1.6896e+003 3.9864e+001 2.8726e+001
# -Range: 0-200
CdCl2
CdCl2 = + 1.0000 Cd++ + 2.0000 Cl-
log_k -0.6474
-delta_H -18.5391 kJ/mol # Calculated enthalpy of reaction CdCl2
# Enthalpy of formation: -391.518 kJ/mol
-analytic -1.5230e+001 -2.4574e-002 -8.1017e+001 8.9599e+000 -1.3702e+000
# -Range: 0-200
CdCl2(NH3)2
CdCl2(NH3)2 = + 1.0000 Cd++ + 2.0000 Cl- + 2.0000 NH3
log_k -8.7864
-delta_H 63.534 kJ/mol # Calculated enthalpy of reaction CdCl2(NH3)2
# Enthalpy of formation: -636.265 kJ/mol
-analytic -5.5283e+001 -2.1791e-002 -2.1150e+003 2.4279e+001 -3.5896e+001
# -Range: 0-200
CdCl2(NH3)4
CdCl2(NH3)4 = + 1.0000 Cd++ + 2.0000 Cl- + 4.0000 NH3
log_k -6.8044
-delta_H 81.7931 kJ/mol # Calculated enthalpy of reaction CdCl2(NH3)4
# Enthalpy of formation: -817.198 kJ/mol
-analytic -9.5682e+001 -1.8853e-002 -8.3875e+002 3.9322e+001 -1.4210e+001
# -Range: 0-200
CdCl2(NH3)6
CdCl2(NH3)6 = + 1.0000 Cd++ + 2.0000 Cl- + 6.0000 NH3
log_k -4.7524
-delta_H 97.2971 kJ/mol # Calculated enthalpy of reaction CdCl2(NH3)6
# Enthalpy of formation: -995.376 kJ/mol
-analytic -1.3662e+002 -1.5941e-002 5.8572e+002 5.4415e+001 9.9937e+000
# -Range: 0-200
CdCl2:H2O
CdCl2:H2O = + 1.0000 Cd++ + 1.0000 H2O + 2.0000 Cl-
log_k -1.6747
-delta_H -7.44943 kJ/mol # Calculated enthalpy of reaction CdCl2:H2O
# Enthalpy of formation: -688.446 kJ/mol
-analytic -4.1097e+001 -2.4685e-002 5.2687e+002 1.8188e+001 8.9615e+000
# -Range: 0-200
CdCr2O4
CdCr2O4 +8.0000 H+ = + 1.0000 Cd++ + 2.0000 Cr+++ + 4.0000 H2O
log_k 14.9969
-delta_H -255.676 kJ/mol # Calculated enthalpy of reaction CdCr2O4
# Enthalpy of formation: -344.3 kcal/mol
-analytic -1.7446e+002 -9.1086e-003 1.9223e+004 5.1605e+001 3.2650e+002
# -Range: 0-200
CdF2
CdF2 = + 1.0000 Cd++ + 2.0000 F-
log_k -1.1464
-delta_H -46.064 kJ/mol # Calculated enthalpy of reaction CdF2
# Enthalpy of formation: -700.529 kJ/mol
-analytic -3.0654e+001 -2.4790e-002 1.7893e+003 1.2482e+001 3.0395e+001
# -Range: 0-200
CdI2
CdI2 = + 1.0000 Cd++ + 2.0000 I-
log_k -3.4825
-delta_H 13.7164 kJ/mol # Calculated enthalpy of reaction CdI2
# Enthalpy of formation: -203.419 kJ/mol
-analytic -1.5446e+001 -2.4758e-002 -1.6422e+003 1.0041e+001 -2.7882e+001
# -Range: 0-200
CdS
CdS +1.0000 H+ = + 1.0000 Cd++ + 1.0000 HS-
log_k -15.9095
-delta_H 70.1448 kJ/mol # Calculated enthalpy of reaction CdS
# Enthalpy of formation: -162.151 kJ/mol
-analytic -2.9492e+001 -1.5181e-002 -3.4695e+003 1.2019e+001 -5.8907e+001
# -Range: 0-200
CdSO4
CdSO4 = + 1.0000 Cd++ + 1.0000 SO4--
log_k -0.1061
-delta_H -52.1304 kJ/mol # Calculated enthalpy of reaction CdSO4
# Enthalpy of formation: -933.369 kJ/mol
-analytic 7.7104e+000 -1.7161e-002 8.7067e+002 -2.2763e+000 1.4783e+001
# -Range: 0-200
CdSO4:2.667H2O
CdSO4:2.667H2O = + 1.0000 Cd++ + 1.0000 SO4-- + 2.6670 H2O
log_k -1.8015
-delta_H -18.5302 kJ/mol # Calculated enthalpy of reaction CdSO4:2.667H2O
# Enthalpy of formation: -1729.3 kJ/mol
-analytic -5.0331e+001 -1.4983e-002 2.0271e+003 1.8665e+001 3.4440e+001
# -Range: 0-200
CdSO4:H2O
CdSO4:H2O = + 1.0000 Cd++ + 1.0000 H2O + 1.0000 SO4--
log_k -1.6529
-delta_H -31.6537 kJ/mol # Calculated enthalpy of reaction CdSO4:H2O
# Enthalpy of formation: -1239.68 kJ/mol
-analytic -1.7142e+001 -1.7295e-002 9.9184e+002 6.9943e+000 1.6849e+001
# -Range: 0-200
CdSeO3
CdSeO3 = + 1.0000 Cd++ + 1.0000 SeO3--
log_k -8.8086
-delta_H -9.92156 kJ/mol # Calculated enthalpy of reaction CdSeO3
# Enthalpy of formation: -575.169 kJ/mol
-analytic 7.1762e+000 -1.8892e-002 -1.4680e+003 -2.1984e+000 -2.4932e+001
# -Range: 0-200
CdSeO4
CdSeO4 = + 1.0000 Cd++ + 1.0000 SeO4--
log_k -2.2132
-delta_H -41.9836 kJ/mol # Calculated enthalpy of reaction CdSeO4
# Enthalpy of formation: -633.063 kJ/mol
-analytic -4.9901e+000 -1.9755e-002 7.3162e+002 2.5063e+000 1.2426e+001
# -Range: 0-200
CdSiO3
CdSiO3 +2.0000 H+ = + 1.0000 Cd++ + 1.0000 H2O + 1.0000 SiO2
log_k 7.5136
-delta_H -50.3427 kJ/mol # Calculated enthalpy of reaction CdSiO3
# Enthalpy of formation: -1189.09 kJ/mol
-analytic 2.6419e+002 6.2488e-002 -5.3518e+003 -1.0401e+002 -9.0973e+001
# -Range: 0-200
Ce
Ce +3.0000 H+ +0.7500 O2 = + 1.0000 Ce+++ + 1.5000 H2O
log_k 182.9563
-delta_H -1120.06 kJ/mol # Calculated enthalpy of reaction Ce
# Enthalpy of formation: 0 kJ/mol
-analytic -5.1017e+001 -2.6149e-002 5.8511e+004 1.8382e+001 9.1302e+002
# -Range: 0-300
Ce(OH)3
Ce(OH)3 +3.0000 H+ = + 1.0000 Ce+++ + 3.0000 H2O
log_k 19.8852
-delta_H 0 # Not possible to calculate enthalpy of reaction Ce(OH)3
# Enthalpy of formation: 0 kcal/mol
Ce(OH)3(am)
Ce(OH)3 +3.0000 H+ = + 1.0000 Ce+++ + 3.0000 H2O
log_k 21.1852
-delta_H 0 # Not possible to calculate enthalpy of reaction Ce(OH)3(am)
# Enthalpy of formation: 0 kcal/mol
Ce2(CO3)3:8H2O
Ce2(CO3)3:8H2O +3.0000 H+ = + 2.0000 Ce+++ + 3.0000 HCO3- + 8.0000 H2O
log_k -4.1136
-delta_H 0 # Not possible to calculate enthalpy of reaction Ce2(CO3)3:8H2O
# Enthalpy of formation: 0 kcal/mol
Ce2O3
Ce2O3 +6.0000 H+ = + 2.0000 Ce+++ + 3.0000 H2O
log_k 62.3000
-delta_H 0 # Not possible to calculate enthalpy of reaction Ce2O3
# Enthalpy of formation: 0 kcal/mol
Ce3(PO4)4
Ce3(PO4)4 +4.0000 H+ = + 3.0000 Ce++++ + 4.0000 HPO4--
log_k -40.8127
-delta_H 0 # Not possible to calculate enthalpy of reaction Ce3(PO4)4
# Enthalpy of formation: 0 kcal/mol
CeF3:.5H2O
CeF3:.5H2O = + 0.5000 H2O + 1.0000 Ce+++ + 3.0000 F-
log_k -18.8000
-delta_H 0 # Not possible to calculate enthalpy of reaction CeF3:.5H2O
# Enthalpy of formation: 0 kcal/mol
CeO2
CeO2 +4.0000 H+ = + 1.0000 Ce++++ + 2.0000 H2O
log_k -8.1600
-delta_H 0 # Not possible to calculate enthalpy of reaction CeO2
# Enthalpy of formation: 0 kcal/mol
CePO4:10H2O
CePO4:10H2O +1.0000 H+ = + 1.0000 Ce+++ + 1.0000 HPO4-- + 10.0000 H2O
log_k -12.2782
-delta_H 0 # Not possible to calculate enthalpy of reaction CePO4:10H2O
# Enthalpy of formation: 0 kcal/mol
Celadonite
KMgAlSi4O10(OH)2 +6.0000 H+ = + 1.0000 Al+++ + 1.0000 K+ + 1.0000 Mg++ + 4.0000 H2O + 4.0000 SiO2
log_k 7.4575
-delta_H -74.3957 kJ/mol # Calculated enthalpy of reaction Celadonite
# Enthalpy of formation: -1394.9 kcal/mol
-analytic -3.3097e+001 1.7989e-002 1.8919e+004 -2.1219e+000 -2.0588e+006
# -Range: 0-300
Celestite
SrSO4 = + 1.0000 SO4-- + 1.0000 Sr++
log_k -5.6771
-delta_H -7.40568 kJ/mol # Calculated enthalpy of reaction Celestite
# Enthalpy of formation: -347.3 kcal/mol
-analytic -1.9063e+002 -7.4552e-002 3.9050e+003 7.8416e+001 6.0991e+001
# -Range: 0-300
Cerussite
PbCO3 +1.0000 H+ = + 1.0000 HCO3- + 1.0000 Pb++
log_k -3.2091
-delta_H 13.8992 kJ/mol # Calculated enthalpy of reaction Cerussite
# Enthalpy of formation: -168 kcal/mol
-analytic -1.2887e+002 -4.4372e-002 2.2336e+003 5.3091e+001 3.4891e+001
# -Range: 0-300
Chalcanthite
CuSO4:5H2O = + 1.0000 Cu++ + 1.0000 SO4-- + 5.0000 H2O
log_k -2.6215
-delta_H 6.57556 kJ/mol # Calculated enthalpy of reaction Chalcanthite
# Enthalpy of formation: -2279.68 kJ/mol
-analytic -1.1262e+002 -1.5544e-002 3.6176e+003 4.1420e+001 6.1471e+001
# -Range: 0-200
Chalcedony
SiO2 = + 1.0000 SiO2
log_k -3.7281
-delta_H 31.4093 kJ/mol # Calculated enthalpy of reaction Chalcedony
# Enthalpy of formation: -217.282 kcal/mol
-analytic -9.0068e+000 9.3241e-003 4.0535e+003 -1.0830e+000 -7.5077e+005
# -Range: 0-300
Chalcocite
Cu2S +1.0000 H+ = + 1.0000 HS- + 2.0000 Cu+
log_k -34.7342
-delta_H 206.748 kJ/mol # Calculated enthalpy of reaction Chalcocite
# Enthalpy of formation: -19 kcal/mol
-analytic -1.3703e+002 -4.0727e-002 -7.1694e+003 5.5963e+001 -1.1183e+002
# -Range: 0-300
Chalcocyanite
CuSO4 = + 1.0000 Cu++ + 1.0000 SO4--
log_k 2.9239
-delta_H -72.5128 kJ/mol # Calculated enthalpy of reaction Chalcocyanite
# Enthalpy of formation: -771.4 kJ/mol
-analytic 5.8173e+000 -1.6933e-002 2.0097e+003 -1.8583e+000 3.4126e+001
# -Range: 0-200
Chalcopyrite
CuFeS2 +2.0000 H+ = + 1.0000 Cu++ + 1.0000 Fe++ + 2.0000 HS-
log_k -32.5638
-delta_H 127.206 kJ/mol # Calculated enthalpy of reaction Chalcopyrite
# Enthalpy of formation: -44.453 kcal/mol
-analytic -3.1575e+002 -9.8947e-002 8.3400e+002 1.2522e+002 1.3106e+001
# -Range: 0-300
Chamosite-7A
Fe2Al2SiO5(OH)4 +10.0000 H+ = + 1.0000 SiO2 + 2.0000 Al+++ + 2.0000 Fe++ + 7.0000 H2O
log_k 32.8416
-delta_H -364.213 kJ/mol # Calculated enthalpy of reaction Chamosite-7A
# Enthalpy of formation: -902.407 kcal/mol
-analytic -2.5581e+002 -7.0890e-002 2.4619e+004 9.1789e+001 3.8424e+002
# -Range: 0-300
AgCl(s)
AgCl = + 1.0000 Ag+ + 1.0000 Cl-
log_k -9.7453
-delta_H 65.739 kJ/mol # Calculated enthalpy of reaction Chlorargyrite
# Enthalpy of formation: -30.37 kcal/mol
-analytic -9.6834e+001 -3.4624e-002 -1.1820e+003 4.0962e+001 -1.8415e+001
# -Range: 0-300
Chloromagnesite
MgCl2 = + 1.0000 Mg++ + 2.0000 Cl-
log_k 21.8604
-delta_H -158.802 kJ/mol # Calculated enthalpy of reaction Chloromagnesite
# Enthalpy of formation: -641.317 kJ/mol
-analytic -2.3640e+002 -8.2017e-002 1.3480e+004 9.5963e+001 2.1042e+002
# -Range: 0-300
Chromite
FeCr2O4 +8.0000 H+ = + 1.0000 Fe++ + 2.0000 Cr+++ + 4.0000 H2O
log_k 15.1685
-delta_H -267.755 kJ/mol # Calculated enthalpy of reaction Chromite
# Enthalpy of formation: -1444.83 kJ/mol
-analytic -1.9060e+002 -2.5695e-002 1.9465e+004 5.9865e+001 3.0379e+002
# -Range: 0-300
Chrysocolla
CuSiH4O5 +2.0000 H+ = + 1.0000 Cu++ + 1.0000 SiO2 + 3.0000 H2O
log_k 6.2142
-delta_H 0 # Not possible to calculate enthalpy of reaction Chrysocolla
# Enthalpy of formation: 0 kcal/mol
Chrysotile
Mg3Si2O5(OH)4 +6.0000 H+ = + 2.0000 SiO2 + 3.0000 Mg++ + 5.0000 H2O
log_k 31.1254
-delta_H -218.041 kJ/mol # Calculated enthalpy of reaction Chrysotile
# Enthalpy of formation: -1043.12 kcal/mol
-analytic -9.2462e+001 -1.1359e-002 1.8312e+004 2.9289e+001 -6.2342e+005
# -Range: 0-300
Cinnabar
HgS +1.0000 H+ = + 1.0000 HS- + 1.0000 Hg++
log_k -38.9666
-delta_H 207.401 kJ/mol # Calculated enthalpy of reaction Cinnabar
# Enthalpy of formation: -12.75 kcal/mol
-analytic -1.5413e+002 -4.6846e-002 -6.9806e+003 6.1639e+001 -1.0888e+002
# -Range: 0-300
Claudetite
As2O3 +3.0000 H2O = + 2.0000 H+ + 2.0000 H2AsO3-
log_k -19.7647
-delta_H 82.3699 kJ/mol # Calculated enthalpy of reaction Claudetite
# Enthalpy of formation: -654.444 kJ/mol
-analytic -1.4164e+002 -6.3704e-002 -2.1679e+003 5.9856e+001 -3.3787e+001
# -Range: 0-300
Clausthalite
PbSe = + 1.0000 Pb++ + 1.0000 Se--
log_k -36.2531
-delta_H 0 # Not possible to calculate enthalpy of reaction Clausthalite
# Enthalpy of formation: -102.9 kJ/mol
-analytic -2.6473e+001 -1.0666e-002 -8.5540e+003 8.9226e+000 -1.3347e+002
# -Range: 0-300
Clinochalcomenite
CuSeO3:2H2O = + 1.0000 Cu++ + 1.0000 SeO3-- + 2.0000 H2O
log_k -6.7873
-delta_H -31.6645 kJ/mol # Calculated enthalpy of reaction Clinochalcomenite
# Enthalpy of formation: -235.066 kcal/mol
-analytic -4.6465e+001 -1.8071e-002 2.0307e+003 1.5455e+001 3.4499e+001
# -Range: 0-200
Clinochlore-14A
Mg5Al2Si3O10(OH)8 +16.0000 H+ = + 2.0000 Al+++ + 3.0000 SiO2 + 5.0000 Mg++ + 12.0000 H2O
log_k 67.2391
-delta_H -612.379 kJ/mol # Calculated enthalpy of reaction Clinochlore-14A
# Enthalpy of formation: -2116.96 kcal/mol
-analytic -2.0441e+002 -6.2268e-002 3.5388e+004 6.9239e+001 5.5225e+002
# -Range: 0-300
Clinochlore-7A
Mg5Al2Si3O10(OH)8 +16.0000 H+ = + 2.0000 Al+++ + 3.0000 SiO2 + 5.0000 Mg++ + 12.0000 H2O
log_k 70.6124
-delta_H -628.14 kJ/mol # Calculated enthalpy of reaction Clinochlore-7A
# Enthalpy of formation: -2113.2 kcal/mol
-analytic -2.1644e+002 -6.4187e-002 3.6548e+004 7.4123e+001 5.7037e+002
# -Range: 0-300
Clinoptilolite
# Na.954K.543Ca.761Mg.124Sr.036Ba.062Mn.002Al3.45F +13.8680 H+ = + 0.0020 Mn++ + 0.0170 Fe+++ + 0.0360 Sr++ + 0.0620 Ba++ + 0.1240 Mg++ + 0.5430 K+ + 0.7610 Ca++ + 0.9540 Na+ + 3.4500 Al+++ + 14.5330 SiO2 17.8560 H2O
Na.954K.543Ca.761Mg.124Sr.036Ba.062Mn.002Al3.45Fe.017Si14.5330O46.922H21.844 +13.8680 H+ = + 0.0020 Mn++ + 0.0170 Fe+++ + 0.0360 Sr++ + 0.0620 Ba++ + 0.1240 Mg++ + 0.5430 K+ + 0.7610 Ca++ + 0.9540 Na+ + 3.4500 Al+++ + 14.5330 SiO2 + 17.8560 H2O
log_k -9.7861
-delta_H -20.8784 kJ/mol # Calculated enthalpy of reaction Clinoptilolite
# Enthalpy of formation: -20587.8 kJ/mol
-analytic -1.3213e+000 6.4960e-002 5.0630e+004 -4.6120e+001 -7.4699e+006
# -Range: 0-300
Clinoptilolite-Ca
Ca1.7335Al3.45Fe.017Si14.533O36:10.922H2O +13.8680 H+ = + 0.0170 Fe+++ + 1.7335 Ca++ + 3.4500 Al+++ + 14.5330 SiO2 + 17.8560 H2O
log_k -7.0095
-delta_H -74.6745 kJ/mol # Calculated enthalpy of reaction Clinoptilolite-Ca
# Enthalpy of formation: -4919.84 kcal/mol
-analytic -4.4820e+001 5.3696e-002 5.4878e+004 -3.1459e+001 -7.5491e+006
# -Range: 0-300
Clinoptilolite-Cs
Cs3.467Al3.45Fe.017Si14.533O36:10.922H2O +13.8680 H+ = + 0.0170 Fe+++ + 3.4500 Al+++ + 3.4670 Cs+ + 14.5330 SiO2 + 17.8560 H2O
log_k -13.0578
-delta_H 96.9005 kJ/mol # Calculated enthalpy of reaction Clinoptilolite-Cs
# Enthalpy of formation: -4949.65 kcal/mol
-analytic -8.4746e+000 7.1997e-002 4.9675e+004 -4.1406e+001 -8.0632e+006
# -Range: 0-300
Clinoptilolite-K
K3.467Al3.45Fe.017Si14.533O36:10.922H2O +13.8680 H+ = + 0.0170 Fe+++ + 3.4500 Al+++ + 3.4670 K+ + 14.5330 SiO2 + 17.8560 H2O
log_k -10.9485
-delta_H 67.4862 kJ/mol # Calculated enthalpy of reaction Clinoptilolite-K
# Enthalpy of formation: -4937.77 kcal/mol
-analytic 1.1697e+001 6.9480e-002 4.7718e+004 -4.7442e+001 -7.6907e+006
# -Range: 0-300
Clinoptilolite-NH4
(NH4)3.467Al3.45Fe.017Si14.533O36:10.922H2O +10.4010 H+ = + 0.0170 Fe+++ + 3.4500 Al+++ + 3.4670 NH3 + 14.5330 SiO2 + 17.8560 H2O
log_k -42.4791
-delta_H 0 # Not possible to calculate enthalpy of reaction Clinoptilolite-NH4
# Enthalpy of formation: 0 kcal/mol
Clinoptilolite-Na
Na3.467Al3.45Fe.017Si14.533O36:10.922H2O +13.8680 H+ = + 0.0170 Fe+++ + 3.4500 Al+++ + 3.4670 Na+ + 14.5330 SiO2 + 17.8560 H2O
log_k -7.1363
-delta_H 2.32824 kJ/mol # Calculated enthalpy of reaction Clinoptilolite-Na
# Enthalpy of formation: -4912.36 kcal/mol
-analytic -3.4572e+001 6.8377e-002 5.1962e+004 -3.3426e+001 -7.5586e+006
# -Range: 0-300
Clinoptilolite-Sr
Sr1.7335Al3.45Fe.017Si14.533O36:10.922H2O +13.8680 H+ = + 0.0170 Fe+++ + 1.7335 Sr++ + 3.4500 Al+++ + 14.5330 SiO2 + 17.8560 H2O
log_k -7.1491
-delta_H -66.2129 kJ/mol # Calculated enthalpy of reaction Clinoptilolite-Sr
# Enthalpy of formation: -4925.1 kcal/mol
-analytic 3.2274e+001 6.7050e-002 5.0880e+004 -5.9597e+001 -7.3876e+006
# -Range: 0-300
Clinoptilolite-dehy
# Sr.036Mg.124Ca.761Mn.002Ba.062K.543Na.954Al3.45F +13.8680 H+ = + 0.0020 Mn++ + 0.0170 Fe+++ + 0.0360 Sr++ + 0.0620 Ba++ + 0.1240 Mg++ + 0.5430 K+ + 0.7610 Ca++ + 0.9540 Na+ + 3.4500 Al+++ + 6.9340 H2O 14.5330 SiO2
Sr.036Mg.124Ca.761Mn.002Ba.062K.543Na.954Al3.45Fe.017Si14.533O36 +13.8680 H+ = + 0.0020 Mn++ + 0.0170 Fe+++ + 0.0360 Sr++ + 0.0620 Ba++ + 0.1240 Mg++ + 0.5430 K+ + 0.7610 Ca++ + 0.9540 Na+ + 3.4500 Al+++ + 6.9340 H2O + 14.5330 SiO2
log_k 25.8490
-delta_H -276.592 kJ/mol # Calculated enthalpy of reaction Clinoptilolite-dehy
# Enthalpy of formation: -17210.2 kJ/mol
-analytic -2.0505e+002 6.0155e-002 8.2682e+004 1.5333e+001 -9.1369e+006
# -Range: 0-300
Clinoptilolite-dehy-Ca
Ca1.7335Al3.45Fe.017Si14.533O36 +13.8680 H+ = + 0.0170 Fe+++ + 1.7335 Ca++ + 3.4500 Al+++ + 6.9340 H2O + 14.5330 SiO2
log_k 28.6255
-delta_H -329.278 kJ/mol # Calculated enthalpy of reaction Clinoptilolite-dehy-Ca
# Enthalpy of formation: -4112.83 kcal/mol
-analytic -1.2948e+002 6.5698e-002 8.0229e+004 -1.2812e+001 -8.8320e+006
# -Range: 0-300
Clinoptilolite-dehy-Cs
Cs3.467Al3.45Fe.017Si14.533O36 +13.8680 H+ = + 0.0170 Fe+++ + 3.4500 Al+++ + 3.4670 Cs+ + 6.9340 H2O + 14.5330 SiO2
log_k 22.5771
-delta_H -164.837 kJ/mol # Calculated enthalpy of reaction Clinoptilolite-dehy-Cs
# Enthalpy of formation: -4140.93 kcal/mol
-analytic -1.2852e+002 7.9047e-002 7.7262e+004 -1.0422e+001 -9.4504e+006
# -Range: 0-300
Clinoptilolite-dehy-K
K3.467Al3.45Fe.017Si14.533O36 +13.8680 H+ = + 0.0170 Fe+++ + 3.4500 Al+++ + 3.4670 K+ + 6.9340 H2O + 14.5330 SiO2
log_k 24.6865
-delta_H -191.289 kJ/mol # Calculated enthalpy of reaction Clinoptilolite-dehy-K
# Enthalpy of formation: -4129.76 kcal/mol
-analytic -1.2241e+002 7.4761e-002 7.6067e+004 -1.1315e+001 -9.1389e+006
# -Range: 0-300
Clinoptilolite-dehy-NH4
(NH4)3.467Al3.45Fe.017Si14.533O36 +10.4010 H+ = + 0.0170 Fe+++ + 3.4500 Al+++ + 3.4670 NH3 + 6.9340 H2O + 14.5330 SiO2
log_k -6.8441
-delta_H 0 # Not possible to calculate enthalpy of reaction Clinoptilolite-dehy-NH4
# Enthalpy of formation: 0 kcal/mol
Clinoptilolite-dehy-Na
Na3.467Al3.45Fe.017Si14.533O36 +13.8680 H+ = + 0.0170 Fe+++ + 3.4500 Al+++ + 3.4670 Na+ + 6.9340 H2O + 14.5330 SiO2
log_k 28.4987
-delta_H -253.798 kJ/mol # Calculated enthalpy of reaction Clinoptilolite-dehy-Na
# Enthalpy of formation: -4104.98 kcal/mol
-analytic -1.4386e+002 7.6846e-002 7.8723e+004 -5.9741e+000 -8.9159e+006
# -Range: 0-300
Clinoptilolite-dehy-Sr
Sr1.7335Al3.45Fe.017Si14.533O36 +13.8680 H+ = + 0.0170 Fe+++ + 1.7335 Sr++ + 3.4500 Al+++ + 6.9340 H2O + 14.5330 SiO2
log_k 28.4859
-delta_H -321.553 kJ/mol # Calculated enthalpy of reaction Clinoptilolite-dehy-Sr
# Enthalpy of formation: -4117.92 kcal/mol
-analytic -1.8410e+002 6.0457e-002 8.3626e+004 6.4304e+000 -9.0962e+006
# -Range: 0-300
Clinoptilolite-hy-Ca
# Ca1.7335Al3.45Fe.017Si14.533036 +13.8680 H+ = + 0.0170 Fe+++ + 1.7335 Ca++ + 3.4500 Al+++ + 14.5330 SiO2 + 18.5790 H2O
Ca1.7335Al3.45Fe.017Si14.533O36:11.645H2O +13.8680 H+ = + 0.0170 Fe+++ + 1.7335 Ca++ + 3.4500 Al+++ + 14.5330 SiO2 + 18.5790 H2O
log_k -7.0108
-delta_H -65.4496 kJ/mol # Calculated enthalpy of reaction Clinoptilolite-hy-Ca
# Enthalpy of formation: -4971.44 kcal/mol
-analytic 8.6833e+001 7.1520e-002 4.6854e+004 -7.8023e+001 -7.0900e+006
# -Range: 0-300
Clinoptilolite-hy-Cs
# Cs3.467Al3.45Fe.017Si14.533036 +13.8680 H+ = + 0.0170 Fe+++ + 3.4500 Al+++ + 3.4670 Cs+ + 13.1640 H2O + 14.5330 SiO2
Cs3.467Al3.45Fe.017Si14.533O36:6.23H2O +13.8680 H+ = + 0.0170 Fe+++ + 3.4500 Al+++ + 3.4670 Cs+ + 13.1640 H2O + 14.5330 SiO2
log_k -13.0621
-delta_H 44.6397 kJ/mol # Calculated enthalpy of reaction Clinoptilolite-hy-Cs
# Enthalpy of formation: -4616.61 kcal/mol
-analytic -2.3362e+001 7.4922e-002 5.4544e+004 -4.1092e+001 -8.3387e+006
# -Range: 0-300
Clinoptilolite-hy-K
# K3.467Al3.45Fe.017Si14.533036 +13.8680 H+ = + 0.0170 Fe+++ + 3.4500 Al+++ + 3.4670 K+ + 14.4330 H2O + 14.5330 SiO2
K3.467Al3.45Fe.017Si14.533O36:7.499H2O +13.8680 H+ = + 0.0170 Fe+++ + 3.4500 Al+++ + 3.4670 K+ + 14.4330 H2O + 14.5330 SiO2
log_k -10.9523
-delta_H 29.5879 kJ/mol # Calculated enthalpy of reaction Clinoptilolite-hy-K
# Enthalpy of formation: -4694.86 kcal/mol
-analytic 1.6223e+001 7.3919e-002 5.0447e+004 -5.2790e+001 -7.8484e+006
# -Range: 0-300
Clinoptilolite-hy-Na
# Na3.467Al3.45Fe.017Si14.533036 +13.8680 H+ = + 0.0170 Fe+++ + 3.4500 Al+++ + 3.4670 Na+ + 14.5330 SiO2 + 17.8110 H2O
Na3.467Al3.45Fe.017Si14.533O36:10.877H2O +13.8680 H+ = + 0.0170 Fe+++ + 3.4500 Al+++ + 3.4670 Na+ + 14.5330 SiO2 + 17.8110 H2O
log_k -7.1384
-delta_H 1.88166 kJ/mol # Calculated enthalpy of reaction Clinoptilolite-hy-Na
# Enthalpy of formation: -4909.18 kcal/mol
-analytic -8.4189e+000 7.2018e-002 5.0501e+004 -4.2851e+001 -7.4714e+006
# -Range: 0-300
Clinoptilolite-hy-Sr
# Sr1.7335Al3.45Fe.017Si14.533036 +13.8680 H+ = + 0.0170 Fe+++ + 1.7335 Sr++ + 3.4500 Al+++ + 14.5330 SiO2 + 20.8270 H2O
Sr1.7335Al3.45Fe.017Si14.533O36:13.893H2O +13.8680 H+ = + 0.0170 Fe+++ + 1.7335 Sr++ + 3.4500 Al+++ + 14.5330 SiO2 + 20.8270 H2O
log_k -7.1498
-delta_H -31.6858 kJ/mol # Calculated enthalpy of reaction Clinoptilolite-hy-Sr
# Enthalpy of formation: -5136.33 kcal/mol
-analytic 1.0742e-001 5.9065e-002 4.9985e+004 -4.4648e+001 -7.3382e+006
# -Range: 0-300
Clinozoisite
Ca2Al3Si3O12(OH) +13.0000 H+ = + 2.0000 Ca++ + 3.0000 Al+++ + 3.0000 SiO2 + 7.0000 H2O
log_k 43.2569
-delta_H -457.755 kJ/mol # Calculated enthalpy of reaction Clinozoisite
# Enthalpy of formation: -1643.78 kcal/mol
-analytic -2.8690e+001 -3.7056e-002 2.2770e+004 3.7880e+000 -2.5834e+005
# -Range: 0-300
Co
Co +2.0000 H+ +0.5000 O2 = + 1.0000 Co++ + 1.0000 H2O
log_k 52.5307
-delta_H -337.929 kJ/mol # Calculated enthalpy of reaction Co
# Enthalpy of formation: 0 kJ/mol
-analytic -6.2703e+001 -2.0172e-002 1.8888e+004 2.3391e+001 2.9474e+002
# -Range: 0-300
Co(NO3)2
Co(NO3)2 = + 1.0000 Co++ + 2.0000 NO3-
log_k 8.0000
-delta_H 0 # Not possible to calculate enthalpy of reaction Co(NO3)2
# Enthalpy of formation: 0 kcal/mol
Co(OH)2
Co(OH)2 +2.0000 H+ = + 1.0000 Co++ + 2.0000 H2O
log_k 12.3000
-delta_H 0 # Not possible to calculate enthalpy of reaction Co(OH)2
# Enthalpy of formation: 0 kcal/mol
Co2SiO4
Co2SiO4 +4.0000 H+ = + 1.0000 SiO2 + 2.0000 Co++ + 2.0000 H2O
log_k 6.6808
-delta_H -88.6924 kJ/mol # Calculated enthalpy of reaction Co2SiO4
# Enthalpy of formation: -353.011 kcal/mol
-analytic -3.9978e+000 -3.7985e-003 5.1554e+003 -1.5033e+000 -1.6100e+005
# -Range: 0-300
Co3(AsO4)2
Co3(AsO4)2 +4.0000 H+ = + 2.0000 H2AsO4- + 3.0000 Co++
log_k 8.5318
-delta_H 0 # Not possible to calculate enthalpy of reaction Co3(AsO4)2
# Enthalpy of formation: 0 kcal/mol
Co3(PO4)2
Co3(PO4)2 +2.0000 H+ = + 2.0000 HPO4-- + 3.0000 Co++
log_k -10.0123
-delta_H 0 # Not possible to calculate enthalpy of reaction Co3(PO4)2
# Enthalpy of formation: 0 kcal/mol
CoCl2
CoCl2 = + 1.0000 Co++ + 2.0000 Cl-
log_k 8.2641
-delta_H -79.5949 kJ/mol # Calculated enthalpy of reaction CoCl2
# Enthalpy of formation: -312.722 kJ/mol
-analytic -2.2386e+002 -8.0936e-002 8.8631e+003 9.1528e+001 1.3837e+002
# -Range: 0-300
CoCl2:2H2O
CoCl2:2H2O = + 1.0000 Co++ + 2.0000 Cl- + 2.0000 H2O
log_k 4.6661
-delta_H -40.7876 kJ/mol # Calculated enthalpy of reaction CoCl2:2H2O
# Enthalpy of formation: -923.206 kJ/mol
-analytic -5.6411e+001 -2.3390e-002 3.0519e+003 2.3361e+001 5.1845e+001
# -Range: 0-200
CoCl2:6H2O
CoCl2:6H2O = + 1.0000 Co++ + 2.0000 Cl- + 6.0000 H2O
log_k 2.6033
-delta_H 8.32709 kJ/mol # Calculated enthalpy of reaction CoCl2:6H2O
# Enthalpy of formation: -2115.67 kJ/mol
-analytic -1.5066e+002 -2.2132e-002 5.0591e+003 5.7743e+001 8.5962e+001
# -Range: 0-200
CoF2
CoF2 = + 1.0000 Co++ + 2.0000 F-
log_k -5.1343
-delta_H -36.6708 kJ/mol # Calculated enthalpy of reaction CoF2
# Enthalpy of formation: -692.182 kJ/mol
-analytic -2.5667e+002 -8.4071e-002 7.6256e+003 1.0143e+002 1.1907e+002
# -Range: 0-300
CoF3
CoF3 = + 1.0000 Co+++ + 3.0000 F-
log_k -4.9558
-delta_H -103.136 kJ/mol # Calculated enthalpy of reaction CoF3
# Enthalpy of formation: -193.8 kcal/mol
-analytic -3.7854e+002 -1.2911e-001 1.3215e+004 1.4859e+002 2.0632e+002
# -Range: 0-300
CoFe2O4
CoFe2O4 +8.0000 H+ = + 1.0000 Co++ + 2.0000 Fe+++ + 4.0000 H2O
log_k 0.8729
-delta_H -160.674 kJ/mol # Calculated enthalpy of reaction CoFe2O4
# Enthalpy of formation: -272.466 kcal/mol
-analytic -3.0149e+002 -7.9159e-002 1.5683e+004 1.1046e+002 2.4480e+002
# -Range: 0-300
CoHPO4
CoHPO4 = + 1.0000 Co++ + 1.0000 HPO4--
log_k -6.7223
-delta_H 0 # Not possible to calculate enthalpy of reaction CoHPO4
# Enthalpy of formation: 0 kcal/mol
CoO
CoO +2.0000 H+ = + 1.0000 Co++ + 1.0000 H2O
log_k 13.5553
-delta_H -106.05 kJ/mol # Calculated enthalpy of reaction CoO
# Enthalpy of formation: -237.946 kJ/mol
-analytic -8.4424e+001 -1.9457e-002 7.8616e+003 3.1281e+001 1.2270e+002
# -Range: 0-300
CoS
CoS +1.0000 H+ = + 1.0000 Co++ + 1.0000 HS-
log_k -7.3740
-delta_H 10.1755 kJ/mol # Calculated enthalpy of reaction CoS
# Enthalpy of formation: -20.182 kcal/mol
-analytic -1.5128e+002 -4.8484e-002 2.9553e+003 5.9983e+001 4.6158e+001
# -Range: 0-300
CoSO4
CoSO4 = + 1.0000 Co++ + 1.0000 SO4--
log_k 2.8996
-delta_H -79.7952 kJ/mol # Calculated enthalpy of reaction CoSO4
# Enthalpy of formation: -887.964 kJ/mol
-analytic -1.9907e+002 -7.7890e-002 7.7193e+003 8.0525e+001 1.2051e+002
# -Range: 0-300
CoSO4.3Co(OH)2
CoSO4(Co(OH)2)3 +6.0000 H+ = + 1.0000 SO4-- + 4.0000 Co++ + 6.0000 H2O
log_k 33.2193
-delta_H -379.41 kJ/mol # Calculated enthalpy of reaction CoSO4.3Co(OH)2
# Enthalpy of formation: -2477.85 kJ/mol
-analytic -2.2830e+002 -4.0197e-002 2.5937e+004 7.5367e+001 4.4053e+002
# -Range: 0-200
CoSO4:6H2O
CoSO4:6H2O = + 1.0000 Co++ + 1.0000 SO4-- + 6.0000 H2O
log_k -2.3512
-delta_H 1.08483 kJ/mol # Calculated enthalpy of reaction CoSO4:6H2O
# Enthalpy of formation: -2683.87 kJ/mol
-analytic -2.5469e+002 -7.3092e-002 6.6767e+003 1.0172e+002 1.0426e+002
# -Range: 0-300
CoSO4:H2O
CoSO4:H2O = + 1.0000 Co++ + 1.0000 H2O + 1.0000 SO4--
log_k -1.2111
-delta_H -52.6556 kJ/mol # Calculated enthalpy of reaction CoSO4:H2O
# Enthalpy of formation: -287.032 kcal/mol
-analytic -1.0570e+001 -1.6196e-002 1.7180e+003 3.4000e+000 2.9178e+001
# -Range: 0-200
CoSeO3
CoSeO3 = + 1.0000 Co++ + 1.0000 SeO3--
log_k -7.0800
-delta_H 0 # Not possible to calculate enthalpy of reaction CoSeO3
# Enthalpy of formation: 0 kcal/mol
CoWO4
CoWO4 = + 1.0000 Co++ + 1.0000 WO4--
log_k -12.2779
-delta_H 13.6231 kJ/mol # Calculated enthalpy of reaction CoWO4
# Enthalpy of formation: -274.256 kcal/mol
-analytic -3.7731e+001 -2.4719e-002 -1.0347e+003 1.4663e+001 -1.7558e+001
# -Range: 0-200
Coesite
SiO2 = + 1.0000 SiO2
log_k -3.1893
-delta_H 28.6144 kJ/mol # Calculated enthalpy of reaction Coesite
# Enthalpy of formation: -216.614 kcal/mol
-analytic -9.7312e+000 9.1773e-003 4.2143e+003 -7.8065e-001 -7.4905e+005
# -Range: 0-300
Coffinite
USiO4 +4.0000 H+ = + 1.0000 SiO2 + 1.0000 U++++ + 2.0000 H2O
log_k -8.0530
-delta_H -49.2493 kJ/mol # Calculated enthalpy of reaction Coffinite
# Enthalpy of formation: -1991.33 kJ/mol
-analytic 2.3126e+002 6.2389e-002 -4.6189e+003 -9.7976e+001 -7.8517e+001
# -Range: 0-200
Colemanite
Ca2B6O11:5H2O +4.0000 H+ +2.0000 H2O = + 2.0000 Ca++ + 6.0000 B(OH)3
log_k 21.5148
-delta_H 0 # Not possible to calculate enthalpy of reaction Colemanite
# Enthalpy of formation: 0 kcal/mol
Cordierite_anhyd
Mg2Al4Si5O18 +16.0000 H+ = + 2.0000 Mg++ + 4.0000 Al+++ + 5.0000 SiO2 + 8.0000 H2O
log_k 52.3035
-delta_H -626.219 kJ/mol # Calculated enthalpy of reaction Cordierite_anhyd
# Enthalpy of formation: -2183.2 kcal/mol
-analytic 2.6562e+000 -2.3801e-002 3.5192e+004 -1.9911e+001 -1.0894e+006
# -Range: 0-300
Cordierite_hydr
Mg2Al4Si5O18:H2O +16.0000 H+ = + 2.0000 Mg++ + 4.0000 Al+++ + 5.0000 SiO2 + 9.0000 H2O
log_k 49.8235
-delta_H -608.814 kJ/mol # Calculated enthalpy of reaction Cordierite_hydr
# Enthalpy of formation: -2255.68 kcal/mol
-analytic -1.2985e+002 -4.1335e-002 4.1566e+004 2.7892e+001 -1.4819e+006
# -Range: 0-300
Corkite
PbFe3(PO4)(SO4)(OH)6 +7.0000 H+ = + 1.0000 HPO4-- + 1.0000 Pb++ + 1.0000 SO4-- + 3.0000 Fe+++ + 6.0000 H2O
log_k -9.7951
-delta_H 0 # Not possible to calculate enthalpy of reaction Corkite
# Enthalpy of formation: 0 kcal/mol
Corundum
Al2O3 +6.0000 H+ = + 2.0000 Al+++ + 3.0000 H2O
log_k 18.3121
-delta_H -258.626 kJ/mol # Calculated enthalpy of reaction Corundum
# Enthalpy of formation: -400.5 kcal/mol
-analytic -1.4278e+002 -7.8519e-002 1.3776e+004 5.5881e+001 2.1501e+002
# -Range: 0-300
Cotunnite
PbCl2 = + 1.0000 Pb++ + 2.0000 Cl-
log_k -4.8406
-delta_H 26.1441 kJ/mol # Calculated enthalpy of reaction Cotunnite
# Enthalpy of formation: -359.383 kJ/mol
-analytic 1.9624e+001 -1.9161e-002 -3.4686e+003 -2.8806e+000 -5.8909e+001
# -Range: 0-200
Covellite
CuS +1.0000 H+ = + 1.0000 Cu++ + 1.0000 HS-
log_k -22.8310
-delta_H 101.88 kJ/mol # Calculated enthalpy of reaction Covellite
# Enthalpy of formation: -12.5 kcal/mol
-analytic -1.6068e+002 -4.9040e-002 -1.4234e+003 6.3536e+001 -2.2164e+001
# -Range: 0-300
Cr
Cr +3.0000 H+ +0.7500 O2 = + 1.0000 Cr+++ + 1.5000 H2O
log_k 98.6784
-delta_H -658.145 kJ/mol # Calculated enthalpy of reaction Cr
# Enthalpy of formation: 0 kJ/mol
-analytic -2.2488e+001 -5.5886e-003 3.4288e+004 3.1585e+000 5.3503e+002
# -Range: 0-300
CrCl3
CrCl3 = + 1.0000 Cr+++ + 3.0000 Cl-
log_k 17.9728
-delta_H -183.227 kJ/mol # Calculated enthalpy of reaction CrCl3
# Enthalpy of formation: -556.5 kJ/mol
-analytic -2.6348e+002 -9.5339e-002 1.4785e+004 1.0517e+002 2.3079e+002
# -Range: 0-300
CrF3
CrF3 = + 1.0000 Cr+++ + 3.0000 F-
log_k -8.5713
-delta_H -85.5293 kJ/mol # Calculated enthalpy of reaction CrF3
# Enthalpy of formation: -277.008 kcal/mol
-analytic -3.2175e+002 -1.0279e-001 1.1394e+004 1.2348e+002 1.7789e+002
# -Range: 0-300
CrF4
CrF4 +2.0000 H2O = + 0.5000 Cr++ + 0.5000 CrO4-- + 4.0000 F- + 4.0000 H+
log_k -12.3132
-delta_H -35.2125 kJ/mol # Calculated enthalpy of reaction CrF4
# Enthalpy of formation: -298 kcal/mol
-analytic 4.3136e+001 -4.3783e-002 -3.6809e+003 -1.2153e+001 -6.2521e+001
# -Range: 0-200
CrI3
CrI3 = + 1.0000 Cr+++ + 3.0000 I-
log_k 25.6112
-delta_H -204.179 kJ/mol # Calculated enthalpy of reaction CrI3
# Enthalpy of formation: -49 kcal/mol
-analytic 4.9232e+000 -2.5164e-002 8.4026e+003 0.0000e+000 0.0000e+000
# -Range: 0-200
CrO2
CrO2 = + 0.5000 Cr++ + 0.5000 CrO4--
log_k -19.1332
-delta_H 85.9812 kJ/mol # Calculated enthalpy of reaction CrO2
# Enthalpy of formation: -143 kcal/mol
-analytic 2.7763e+000 -7.7698e-003 -5.2893e+003 -7.4970e-001 -8.9821e+001
# -Range: 0-200
CrO3
CrO3 +1.0000 H2O = + 1.0000 CrO4-- + 2.0000 H+
log_k -3.5221
-delta_H -5.78647 kJ/mol # Calculated enthalpy of reaction CrO3
# Enthalpy of formation: -140.9 kcal/mol
-analytic -1.3262e+002 -6.1411e-002 2.2083e+003 5.6564e+001 3.4497e+001
# -Range: 0-300
CrS
CrS +1.0000 H+ = + 1.0000 Cr++ + 1.0000 HS-
log_k -0.6304
-delta_H -26.15 kJ/mol # Calculated enthalpy of reaction CrS
# Enthalpy of formation: -31.9 kcal/mol
-analytic -1.1134e+002 -3.5954e-002 3.8744e+003 4.3815e+001 6.0490e+001
# -Range: 0-300
Cristobalite(alpha)
SiO2 = + 1.0000 SiO2
log_k -3.4488
-delta_H 29.2043 kJ/mol # Calculated enthalpy of reaction Cristobalite(alpha)
# Enthalpy of formation: -216.755 kcal/mol
-analytic -1.1936e+001 9.0520e-003 4.3701e+003 -1.1464e-001 -7.6568e+005
# -Range: 0-300
Cristobalite(beta)
SiO2 = + 1.0000 SiO2
log_k -3.0053
-delta_H 24.6856 kJ/mol # Calculated enthalpy of reaction Cristobalite(beta)
# Enthalpy of formation: -215.675 kcal/mol
-analytic -4.7414e+000 9.7567e-003 3.8831e+003 -2.5830e+000 -6.9636e+005
# -Range: 0-300
Crocoite
PbCrO4 = + 1.0000 CrO4-- + 1.0000 Pb++
log_k -12.7177
-delta_H 48.6181 kJ/mol # Calculated enthalpy of reaction Crocoite
# Enthalpy of formation: -222 kcal/mol
-analytic 3.0842e+001 -1.4430e-002 -5.0292e+003 -9.0525e+000 -8.5414e+001
# -Range: 0-200
Cronstedtite-7A
Fe2Fe2SiO5(OH)4 +10.0000 H+ = + 1.0000 SiO2 + 2.0000 Fe++ + 2.0000 Fe+++ + 7.0000 H2O
log_k 16.2603
-delta_H -244.266 kJ/mol # Calculated enthalpy of reaction Cronstedtite-7A
# Enthalpy of formation: -697.413 kcal/mol
-analytic -2.3783e+002 -7.1026e-002 1.7752e+004 8.7147e+001 2.7707e+002
# -Range: 0-300
Cs
Cs +1.0000 H+ +0.2500 O2 = + 0.5000 H2O + 1.0000 Cs+
log_k 72.5987
-delta_H -397.913 kJ/mol # Calculated enthalpy of reaction Cs
# Enthalpy of formation: 0 kJ/mol
-analytic -1.2875e+001 -7.3845e-003 2.1019e+004 6.9347e+000 3.2799e+002
# -Range: 0-300
Cs2NaAmCl6
Cs2NaAmCl6 = + 1.0000 Am+++ + 1.0000 Na+ + 2.0000 Cs+ + 6.0000 Cl-
log_k 11.7089
-delta_H -59.7323 kJ/mol # Calculated enthalpy of reaction Cs2NaAmCl6
# Enthalpy of formation: -2315.8 kJ/mol
-analytic 5.1683e+001 -5.0340e-002 -2.3205e+003 -6.9536e+000 -3.9422e+001
# -Range: 0-200
Cs2U2O7
Cs2U2O7 +6.0000 H+ = + 2.0000 Cs+ + 2.0000 UO2++ + 3.0000 H2O
log_k 31.0263
-delta_H -191.57 kJ/mol # Calculated enthalpy of reaction Cs2U2O7
# Enthalpy of formation: -3220 kJ/mol
-analytic -5.1436e+001 -7.4096e-003 1.2524e+004 1.7827e+001 -1.2899e+005
# -Range: 0-300
Cs2U4O12
Cs2U4O12 +8.0000 H+ = + 2.0000 Cs+ + 2.0000 UO2+ + 2.0000 UO2++ + 4.0000 H2O
log_k 18.9460
-delta_H -175.862 kJ/mol # Calculated enthalpy of reaction Cs2U4O12
# Enthalpy of formation: -5571.8 kJ/mol
-analytic -3.3411e+001 3.6196e-003 1.0508e+004 6.5823e+000 -2.3403e+004
# -Range: 0-300
Cs2UO4
Cs2UO4 +4.0000 H+ = + 1.0000 UO2++ + 2.0000 Cs+ + 2.0000 H2O
log_k 35.8930
-delta_H -178.731 kJ/mol # Calculated enthalpy of reaction Cs2UO4
# Enthalpy of formation: -1928 kJ/mol
-analytic -3.0950e+001 -3.5650e-003 1.0690e+004 1.2949e+001 1.6682e+002
# -Range: 0-300
Cu
Cu +2.0000 H+ +0.5000 O2 = + 1.0000 Cu++ + 1.0000 H2O
log_k 31.5118
-delta_H -214.083 kJ/mol # Calculated enthalpy of reaction Cu
# Enthalpy of formation: 0 kcal/mol
-analytic -7.0719e+001 -2.0300e-002 1.2802e+004 2.6401e+001 1.9979e+002
# -Range: 0-300
Cu3(PO4)2
Cu3(PO4)2 +2.0000 H+ = + 2.0000 HPO4-- + 3.0000 Cu++
log_k -12.2247
-delta_H 0 # Not possible to calculate enthalpy of reaction Cu3(PO4)2
# Enthalpy of formation: 0 kcal/mol
Cu3(PO4)2:3H2O
Cu3(PO4)2:3H2O +2.0000 H+ = + 2.0000 HPO4-- + 3.0000 Cu++ + 3.0000 H2O
log_k -10.4763
-delta_H 0 # Not possible to calculate enthalpy of reaction Cu3(PO4)2:3H2O
# Enthalpy of formation: 0 kcal/mol
CuCl2
CuCl2 = + 1.0000 Cu++ + 2.0000 Cl-
log_k 3.7308
-delta_H -48.5965 kJ/mol # Calculated enthalpy of reaction CuCl2
# Enthalpy of formation: -219.874 kJ/mol
-analytic -1.7803e+001 -2.4432e-002 1.5729e+003 9.5104e+000 2.6716e+001
# -Range: 0-200
CuCr2O4
CuCr2O4 +8.0000 H+ = + 1.0000 Cu++ + 2.0000 Cr+++ + 4.0000 H2O
log_k 16.2174
-delta_H -268.768 kJ/mol # Calculated enthalpy of reaction CuCr2O4
# Enthalpy of formation: -307.331 kcal/mol
-analytic -1.8199e+002 -1.0254e-002 2.0123e+004 5.4062e+001 3.4178e+002
# -Range: 0-200
CuF
CuF = + 1.0000 Cu+ + 1.0000 F-
log_k 7.0800
-delta_H 0 # Not possible to calculate enthalpy of reaction CuF
# Enthalpy of formation: 0 kcal/mol
CuF2
CuF2 = + 1.0000 Cu++ + 2.0000 F-
log_k -0.6200
-delta_H 0 # Not possible to calculate enthalpy of reaction CuF2
# Enthalpy of formation: 0 kcal/mol
CuF2:2H2O
CuF2:2H2O = + 1.0000 Cu++ + 2.0000 F- + 2.0000 H2O
log_k -4.5500
-delta_H 0 # Not possible to calculate enthalpy of reaction CuF2:2H2O
# Enthalpy of formation: 0 kcal/mol
CuSeO3
CuSeO3 = + 1.0000 Cu++ + 1.0000 SeO3--
log_k -7.6767
-delta_H 0 # Not possible to calculate enthalpy of reaction CuSeO3
# Enthalpy of formation: 0 kcal/mol
Cuprite
Cu2O +2.0000 H+ = + 1.0000 H2O + 2.0000 Cu+
log_k -1.9031
-delta_H 28.355 kJ/mol # Calculated enthalpy of reaction Cuprite
# Enthalpy of formation: -40.83 kcal/mol
-analytic -8.6240e+001 -1.1445e-002 1.7851e+003 3.3041e+001 2.7880e+001
# -Range: 0-300
Daphnite-14A
Fe5AlAlSi3O10(OH)8 +16.0000 H+ = + 2.0000 Al+++ + 3.0000 SiO2 + 5.0000 Fe++ + 12.0000 H2O
log_k 52.2821
-delta_H -517.561 kJ/mol # Calculated enthalpy of reaction Daphnite-14A
# Enthalpy of formation: -1693.04 kcal/mol
-analytic -1.5261e+002 -6.1392e-002 2.8283e+004 5.1788e+001 4.4137e+002
# -Range: 0-300
Daphnite-7A
Fe5AlAlSi3O10(OH)8 +16.0000 H+ = + 2.0000 Al+++ + 3.0000 SiO2 + 5.0000 Fe++ + 12.0000 H2O
log_k 55.6554
-delta_H -532.326 kJ/mol # Calculated enthalpy of reaction Daphnite-7A
# Enthalpy of formation: -1689.51 kcal/mol
-analytic -1.6430e+002 -6.3160e-002 2.9499e+004 5.6442e+001 4.6035e+002
# -Range: 0-300
Dawsonite
NaAlCO3(OH)2 +3.0000 H+ = + 1.0000 Al+++ + 1.0000 HCO3- + 1.0000 Na+ + 2.0000 H2O
log_k 4.3464
-delta_H -76.3549 kJ/mol # Calculated enthalpy of reaction Dawsonite
# Enthalpy of formation: -1963.96 kJ/mol
-analytic -1.1393e+002 -2.3487e-002 7.1758e+003 4.0900e+001 1.2189e+002
# -Range: 0-200
Delafossite
CuFeO2 +4.0000 H+ = + 1.0000 Cu+ + 1.0000 Fe+++ + 2.0000 H2O
log_k -6.4172
-delta_H -18.6104 kJ/mol # Calculated enthalpy of reaction Delafossite
# Enthalpy of formation: -126.904 kcal/mol
-analytic -1.5275e+002 -3.5478e-002 5.1404e+003 5.6437e+001 8.0255e+001
# -Range: 0-300
Diaspore
AlHO2 +3.0000 H+ = + 1.0000 Al+++ + 2.0000 H2O
log_k 7.1603
-delta_H -110.42 kJ/mol # Calculated enthalpy of reaction Diaspore
# Enthalpy of formation: -238.924 kcal/mol
-analytic -1.2618e+002 -3.1671e-002 8.8737e+003 4.5669e+001 1.3850e+002
# -Range: 0-300
Dicalcium_silicate
Ca2SiO4 +4.0000 H+ = + 1.0000 SiO2 + 2.0000 Ca++ + 2.0000 H2O
log_k 37.1725
-delta_H -217.642 kJ/mol # Calculated enthalpy of reaction Dicalcium_silicate
# Enthalpy of formation: -2317.9 kJ/mol
-analytic -5.9723e+001 -1.3682e-002 1.5461e+004 2.1547e+001 -3.7732e+005
# -Range: 0-300
Diopside
CaMgSi2O6 +4.0000 H+ = + 1.0000 Ca++ + 1.0000 Mg++ + 2.0000 H2O + 2.0000 SiO2
log_k 20.9643
-delta_H -133.775 kJ/mol # Calculated enthalpy of reaction Diopside
# Enthalpy of formation: -765.378 kcal/mol
-analytic 7.1240e+001 1.5514e-002 8.1437e+003 -3.0672e+001 -5.6880e+005
# -Range: 0-300
Dioptase
CuSiO2(OH)2 +2.0000 H+ = + 1.0000 Cu++ + 1.0000 SiO2 + 2.0000 H2O
log_k 6.0773
-delta_H -25.2205 kJ/mol # Calculated enthalpy of reaction Dioptase
# Enthalpy of formation: -1358.47 kJ/mol
-analytic 2.3913e+002 6.2669e-002 -5.4030e+003 -9.4420e+001 -9.1834e+001
# -Range: 0-200
Dolomite
CaMg(CO3)2 +2.0000 H+ = + 1.0000 Ca++ + 1.0000 Mg++ + 2.0000 HCO3-
log_k 2.5135
-delta_H -59.9651 kJ/mol # Calculated enthalpy of reaction Dolomite
# Enthalpy of formation: -556.631 kcal/mol
-analytic -3.1782e+002 -9.8179e-002 1.0845e+004 1.2657e+002 1.6932e+002
# -Range: 0-300
Dolomite-dis
CaMg(CO3)2 +2.0000 H+ = + 1.0000 Ca++ + 1.0000 Mg++ + 2.0000 HCO3-
log_k 4.0579
-delta_H -72.2117 kJ/mol # Calculated enthalpy of reaction Dolomite-dis
# Enthalpy of formation: -553.704 kcal/mol
-analytic -3.1706e+002 -9.7886e-002 1.1442e+004 1.2604e+002 1.7864e+002
# -Range: 0-300
Dolomite-ord
CaMg(CO3)2 +2.0000 H+ = + 1.0000 Ca++ + 1.0000 Mg++ + 2.0000 HCO3-
log_k 2.5135
-delta_H -59.9651 kJ/mol # Calculated enthalpy of reaction Dolomite-ord
# Enthalpy of formation: -556.631 kcal/mol
-analytic -3.1654e+002 -9.7902e-002 1.0805e+004 1.2607e+002 1.6870e+002
# -Range: 0-300
Downeyite
SeO2 +1.0000 H2O = + 1.0000 SeO3-- + 2.0000 H+
log_k -6.7503
-delta_H 1.74473 kJ/mol # Calculated enthalpy of reaction Downeyite
# Enthalpy of formation: -53.8 kcal/mol
-analytic -1.2868e+002 -6.1183e-002 1.5802e+003 5.4490e+001 2.4696e+001
# -Range: 0-300
Dy
Dy +3.0000 H+ +0.7500 O2 = + 1.0000 Dy+++ + 1.5000 H2O
log_k 180.8306
-delta_H -1116.29 kJ/mol # Calculated enthalpy of reaction Dy
# Enthalpy of formation: 0 kJ/mol
-analytic -6.8317e+001 -2.8321e-002 5.8927e+004 2.4211e+001 9.1953e+002
# -Range: 0-300
Dy(OH)3
Dy(OH)3 +3.0000 H+ = + 1.0000 Dy+++ + 3.0000 H2O
log_k 15.8852
-delta_H 0 # Not possible to calculate enthalpy of reaction Dy(OH)3
# Enthalpy of formation: 0 kcal/mol
Dy(OH)3(am)
Dy(OH)3 +3.0000 H+ = + 1.0000 Dy+++ + 3.0000 H2O
log_k 17.4852
-delta_H 0 # Not possible to calculate enthalpy of reaction Dy(OH)3(am)
# Enthalpy of formation: 0 kcal/mol
Dy2(CO3)3
Dy2(CO3)3 +3.0000 H+ = + 2.0000 Dy+++ + 3.0000 HCO3-
log_k -3.0136
-delta_H 0 # Not possible to calculate enthalpy of reaction Dy2(CO3)3
# Enthalpy of formation: 0 kcal/mol
Dy2O3
Dy2O3 +6.0000 H+ = + 2.0000 Dy+++ + 3.0000 H2O
log_k 47.0000
-delta_H 0 # Not possible to calculate enthalpy of reaction Dy2O3
# Enthalpy of formation: 0 kcal/mol
DyF3:.5H2O
DyF3:.5H2O = + 0.5000 H2O + 1.0000 Dy+++ + 3.0000 F-
log_k -16.5000
-delta_H 0 # Not possible to calculate enthalpy of reaction DyF3:.5H2O
# Enthalpy of formation: 0 kcal/mol
DyPO4:10H2O
DyPO4:10H2O +1.0000 H+ = + 1.0000 Dy+++ + 1.0000 HPO4-- + 10.0000 H2O
log_k -11.9782
-delta_H 0 # Not possible to calculate enthalpy of reaction DyPO4:10H2O
# Enthalpy of formation: 0 kcal/mol
Enstatite
MgSiO3 +2.0000 H+ = + 1.0000 H2O + 1.0000 Mg++ + 1.0000 SiO2
log_k 11.3269
-delta_H -82.7302 kJ/mol # Calculated enthalpy of reaction Enstatite
# Enthalpy of formation: -369.686 kcal/mol
-analytic -4.9278e+001 -3.2832e-003 9.5205e+003 1.4437e+001 -5.4324e+005
# -Range: 0-300
Epidote
Ca2FeAl2Si3O12OH +13.0000 H+ = + 1.0000 Fe+++ + 2.0000 Al+++ + 2.0000 Ca++ + 3.0000 SiO2 + 7.0000 H2O
log_k 32.9296
-delta_H -386.451 kJ/mol # Calculated enthalpy of reaction Epidote
# Enthalpy of formation: -1543.99 kcal/mol
-analytic -2.6187e+001 -3.6436e-002 1.9351e+004 3.3671e+000 -3.0319e+005
# -Range: 0-300
Epidote-ord
FeCa2Al2(OH)(SiO4)3 +13.0000 H+ = + 1.0000 Fe+++ + 2.0000 Al+++ + 2.0000 Ca++ + 3.0000 SiO2 + 7.0000 H2O
log_k 32.9296
-delta_H -386.351 kJ/mol # Calculated enthalpy of reaction Epidote-ord
# Enthalpy of formation: -1544.02 kcal/mol
-analytic 1.9379e+001 -3.2870e-002 1.5692e+004 -1.1901e+001 2.4485e+002
# -Range: 0-300
Epsomite
MgSO4:7H2O = + 1.0000 Mg++ + 1.0000 SO4-- + 7.0000 H2O
log_k -1.9623
-delta_H 0 # Not possible to calculate enthalpy of reaction Epsomite
# Enthalpy of formation: 0 kcal/mol
Er
Er +3.0000 H+ +0.7500 O2 = + 1.0000 Er+++ + 1.5000 H2O
log_k 181.7102
-delta_H -1124.66 kJ/mol # Calculated enthalpy of reaction Er
# Enthalpy of formation: 0 kJ/mol
-analytic -1.4459e+002 -3.8221e-002 6.4073e+004 5.1047e+001 -3.1503e+005
# -Range: 0-300
Er(OH)3
Er(OH)3 +3.0000 H+ = + 1.0000 Er+++ + 3.0000 H2O
log_k 14.9852
-delta_H 0 # Not possible to calculate enthalpy of reaction Er(OH)3
# Enthalpy of formation: 0 kcal/mol
Er(OH)3(am)
Er(OH)3 +3.0000 H+ = + 1.0000 Er+++ + 3.0000 H2O
log_k 18.9852
-delta_H 0 # Not possible to calculate enthalpy of reaction Er(OH)3(am)
# Enthalpy of formation: 0 kcal/mol
Er2(CO3)3
Er2(CO3)3 +3.0000 H+ = + 2.0000 Er+++ + 3.0000 HCO3-
log_k -2.6136
-delta_H 0 # Not possible to calculate enthalpy of reaction Er2(CO3)3
# Enthalpy of formation: 0 kcal/mol
Er2O3
Er2O3 +6.0000 H+ = + 2.0000 Er+++ + 3.0000 H2O
log_k 42.1000
-delta_H 0 # Not possible to calculate enthalpy of reaction Er2O3
# Enthalpy of formation: 0 kcal/mol
ErF3:.5H2O
ErF3:.5H2O = + 0.5000 H2O + 1.0000 Er+++ + 3.0000 F-
log_k -16.3000
-delta_H 0 # Not possible to calculate enthalpy of reaction ErF3:.5H2O
# Enthalpy of formation: 0 kcal/mol
ErPO4:10H2O
ErPO4:10H2O +1.0000 H+ = + 1.0000 Er+++ + 1.0000 HPO4-- + 10.0000 H2O
log_k -11.8782
-delta_H 0 # Not possible to calculate enthalpy of reaction ErPO4:10H2O
# Enthalpy of formation: 0 kcal/mol
Erythrite
Co3(AsO4)2:8H2O +4.0000 H+ = + 2.0000 H2AsO4- + 3.0000 Co++ + 8.0000 H2O
log_k 6.3930
-delta_H 0 # Not possible to calculate enthalpy of reaction Erythrite
# Enthalpy of formation: 0 kcal/mol
Eskolaite
Cr2O3 +2.0000 H2O +1.5000 O2 = + 2.0000 CrO4-- + 4.0000 H+
log_k -9.1306
-delta_H -32.6877 kJ/mol # Calculated enthalpy of reaction Eskolaite
# Enthalpy of formation: -1139.74 kJ/mol
-analytic -2.0411e+002 -1.2809e-001 2.2197e+003 9.1186e+001 3.4697e+001
# -Range: 0-300
Ettringite
Ca6Al2(SO4)3(OH)12:26H2O +12.0000 H+ = + 2.0000 Al+++ + 3.0000 SO4-- + 6.0000 Ca++ + 38.0000 H2O
log_k 62.5362
-delta_H -382.451 kJ/mol # Calculated enthalpy of reaction Ettringite
# Enthalpy of formation: -4193 kcal/mol
-analytic -1.0576e+003 -1.1585e-001 5.9580e+004 3.8585e+002 1.0121e+003
# -Range: 0-200
Eu
Eu +3.0000 H+ +0.7500 O2 = + 1.0000 Eu+++ + 1.5000 H2O
log_k 165.1443
-delta_H -1025.08 kJ/mol # Calculated enthalpy of reaction Eu
# Enthalpy of formation: 0 kJ/mol
-analytic -6.5749e+001 -2.8921e-002 5.4018e+004 2.3561e+001 8.4292e+002
# -Range: 0-300
Eu(IO3)3:2H2O
Eu(IO3)3:2H2O = + 1.0000 Eu+++ + 2.0000 H2O + 3.0000 IO3-
log_k -11.6999
-delta_H 20.8847 kJ/mol # Calculated enthalpy of reaction Eu(IO3)3:2H2O
# Enthalpy of formation: -1861.99 kJ/mol
-analytic -3.4616e+001 -1.9914e-002 -1.1966e+003 1.3276e+001 -2.0308e+001
# -Range: 0-200
Eu(NO3)3:6H2O
Eu(NO3)3:6H2O = + 1.0000 Eu+++ + 3.0000 NO3- + 6.0000 H2O
log_k 1.3082
-delta_H 15.2254 kJ/mol # Calculated enthalpy of reaction Eu(NO3)3:6H2O
# Enthalpy of formation: -2956.11 kJ/mol
-analytic -1.3205e+002 -2.0427e-002 3.9623e+003 5.0976e+001 6.7332e+001
# -Range: 0-200
Eu(OH)2.5Cl.5
Eu(OH)2.5Cl.5 +2.5000 H+ = + 0.5000 Cl- + 1.0000 Eu+++ + 2.5000 H2O
log_k 12.5546
-delta_H 0 # Not possible to calculate enthalpy of reaction Eu(OH)2.5Cl.5
# Enthalpy of formation: 0 kcal/mol
Eu(OH)2Cl
Eu(OH)2Cl +2.0000 H+ = + 1.0000 Cl- + 1.0000 Eu+++ + 2.0000 H2O
log_k 8.7974
-delta_H 0 # Not possible to calculate enthalpy of reaction Eu(OH)2Cl
# Enthalpy of formation: 0 kcal/mol
Eu(OH)3(s)
Eu(OH)3 +3.0000 H+ = + 1.0000 Eu+++ + 3.0000 H2O
log_k 15.3482
-delta_H -126.897 kJ/mol # Calculated enthalpy of reaction Eu(OH)3
# Enthalpy of formation: -1336.04 kJ/mol
-analytic -6.3077e+001 -6.1421e-003 8.7323e+003 2.0595e+001 1.4831e+002
# -Range: 0-200
Eu2(CO3)3:3H2O
Eu2(CO3)3:3H2O +3.0000 H+ = + 2.0000 Eu+++ + 3.0000 H2O + 3.0000 HCO3-
log_k -5.8707
-delta_H -137.512 kJ/mol # Calculated enthalpy of reaction Eu2(CO3)3:3H2O
# Enthalpy of formation: -4000.65 kJ/mol
-analytic -1.4134e+002 -4.0240e-002 9.5883e+003 4.6591e+001 1.6287e+002
# -Range: 0-200
Eu2(SO4)3:8H2O
Eu2(SO4)3:8H2O = + 2.0000 Eu+++ + 3.0000 SO4-- + 8.0000 H2O
log_k -10.8524
-delta_H -86.59 kJ/mol # Calculated enthalpy of reaction Eu2(SO4)3:8H2O
# Enthalpy of formation: -6139.77 kJ/mol
-analytic -5.6582e+001 -3.8846e-002 3.3821e+003 1.8561e+001 5.7452e+001
# -Range: 0-200
Eu2O3(cubic)
Eu2O3 +6.0000 H+ = + 2.0000 Eu+++ + 3.0000 H2O
log_k 51.7818
-delta_H -406.403 kJ/mol # Calculated enthalpy of reaction Eu2O3(cubic)
# Enthalpy of formation: -1661.96 kJ/mol
-analytic -5.3469e+001 -1.2554e-002 2.1925e+004 1.4324e+001 3.7233e+002
# -Range: 0-200
Eu2O3(monoclinic)
Eu2O3 +6.0000 H+ = + 2.0000 Eu+++ + 3.0000 H2O
log_k 53.3936
-delta_H -417.481 kJ/mol # Calculated enthalpy of reaction Eu2O3(monoclinic)
# Enthalpy of formation: -1650.88 kJ/mol
-analytic -5.4022e+001 -1.2627e-002 2.2508e+004 1.4416e+001 3.8224e+002
# -Range: 0-200
Eu3O4
Eu3O4 +8.0000 H+ = + 1.0000 Eu++ + 2.0000 Eu+++ + 4.0000 H2O
log_k 87.0369
-delta_H -611.249 kJ/mol # Calculated enthalpy of reaction Eu3O4
# Enthalpy of formation: -2270.56 kJ/mol
-analytic -1.1829e+002 -2.0354e-002 3.4981e+004 3.8007e+001 5.9407e+002
# -Range: 0-200
EuBr3
EuBr3 = + 1.0000 Eu+++ + 3.0000 Br-
log_k 29.8934
-delta_H -217.166 kJ/mol # Calculated enthalpy of reaction EuBr3
# Enthalpy of formation: -752.769 kJ/mol
-analytic 6.0207e+001 -2.5234e-002 6.6823e+003 -1.8276e+001 1.1345e+002
# -Range: 0-200
EuCl2
EuCl2 = + 1.0000 Eu++ + 2.0000 Cl-
log_k 5.9230
-delta_H -39.2617 kJ/mol # Calculated enthalpy of reaction EuCl2
# Enthalpy of formation: -822.5 kJ/mol
-analytic -2.5741e+001 -2.4956e-002 1.5713e+003 1.3670e+001 2.6691e+001
# -Range: 0-200
EuCl3
EuCl3 = + 1.0000 Eu+++ + 3.0000 Cl-
log_k 19.7149
-delta_H -170.861 kJ/mol # Calculated enthalpy of reaction EuCl3
# Enthalpy of formation: -935.803 kJ/mol
-analytic 3.2865e+001 -3.1877e-002 4.9792e+003 -8.2294e+000 8.4542e+001
# -Range: 0-200
EuCl3:6H2O
EuCl3:6H2O = + 1.0000 Eu+++ + 3.0000 Cl- + 6.0000 H2O
log_k 4.9090
-delta_H -40.0288 kJ/mol # Calculated enthalpy of reaction EuCl3:6H2O
# Enthalpy of formation: -2781.66 kJ/mol
-analytic -1.0987e+002 -2.9851e-002 4.9991e+003 4.3198e+001 8.4930e+001
# -Range: 0-200
EuF3:0.5H2O
EuF3:0.5H2O = + 0.5000 H2O + 1.0000 Eu+++ + 3.0000 F-
log_k -16.4847
-delta_H 0 # Not possible to calculate enthalpy of reaction EuF3:0.5H2O
# Enthalpy of formation: 0 kcal/mol
EuO
EuO +2.0000 H+ = + 1.0000 Eu++ + 1.0000 H2O
log_k 37.4800
-delta_H -221.196 kJ/mol # Calculated enthalpy of reaction EuO
# Enthalpy of formation: -592.245 kJ/mol
-analytic -8.9517e+001 -1.7523e-002 1.4385e+004 3.3933e+001 2.2449e+002
# -Range: 0-300
EuOCl
EuOCl +2.0000 H+ = + 1.0000 Cl- + 1.0000 Eu+++ + 1.0000 H2O
log_k 15.6683
-delta_H -147.173 kJ/mol # Calculated enthalpy of reaction EuOCl
# Enthalpy of formation: -911.17 kJ/mol
-analytic -7.7446e+000 -1.4960e-002 6.6242e+003 2.2813e+000 1.1249e+002
# -Range: 0-200
EuOHCO3(s)
# EuOHCO3 = 1.0000 Eu+++ + 1.0000 OH- + 1.0000 CO3-2 # from Runde 1992 radiochim acta 58/59 93-100
# log_k -20.18
EuOHCO3 +2.0000 H+ = + 1.0000 Eu+++ + 1.0000 H2O + 1.0000 HCO3-
log_k 2.5239
-delta_H 0 # Not possible to calculate enthalpy of reaction EuOHCO3
# Enthalpy of formation: 0 kcal/mol
EuPO4(s)
EuPO4:10H2O +1.0000 H+ = + 1.0000 Eu+++ + 1.0000 HPO4-- + 10.0000 H2O
log_k -12.0782
-delta_H 0 # Not possible to calculate enthalpy of reaction EuPO4:10H2O
# Enthalpy of formation: 0 kcal/mol
EuS
EuS +1.0000 H+ = + 1.0000 Eu++ + 1.0000 HS-
log_k 14.9068
-delta_H -96.4088 kJ/mol # Calculated enthalpy of reaction EuS
# Enthalpy of formation: -447.302 kJ/mol
-analytic -4.1026e+001 -1.5582e-002 5.7842e+003 1.6639e+001 9.8238e+001
# -Range: 0-200
EuSO4
EuSO4 = + 1.0000 Eu++ + 1.0000 SO4--
log_k -8.8449
-delta_H 33.873 kJ/mol # Calculated enthalpy of reaction EuSO4
# Enthalpy of formation: -1471.08 kJ/mol
-analytic 3.0262e-001 -1.7571e-002 -3.0392e+003 2.5356e+000 -5.1610e+001
# -Range: 0-200
Eucryptite
LiAlSiO4 +4.0000 H+ = + 1.0000 Al+++ + 1.0000 Li+ + 1.0000 SiO2 + 2.0000 H2O
log_k 13.6106
-delta_H -141.818 kJ/mol # Calculated enthalpy of reaction Eucryptite
# Enthalpy of formation: -2124.41 kJ/mol
-analytic -2.2213e+000 -8.2498e-003 6.4838e+003 -1.4183e+000 1.0117e+002
# -Range: 0-300
Fayalite
Fe2SiO4 +4.0000 H+ = + 1.0000 SiO2 + 2.0000 Fe++ + 2.0000 H2O
log_k 19.1113
-delta_H -152.256 kJ/mol # Calculated enthalpy of reaction Fayalite
# Enthalpy of formation: -354.119 kcal/mol
-analytic 1.3853e+001 -3.5501e-003 7.1496e+003 -6.8710e+000 -6.3310e+004
# -Range: 0-300
Fe
Fe +2.0000 H+ +0.5000 O2 = + 1.0000 Fe++ + 1.0000 H2O
log_k 59.0325
-delta_H -372.029 kJ/mol # Calculated enthalpy of reaction Fe
# Enthalpy of formation: 0 kcal/mol
-analytic -6.2882e+001 -2.0379e-002 2.0690e+004 2.3673e+001 3.2287e+002
# -Range: 0-300
Fe(OH)2
Fe(OH)2 +2.0000 H+ = + 1.0000 Fe++ + 2.0000 H2O
log_k 13.9045
-delta_H -95.4089 kJ/mol # Calculated enthalpy of reaction Fe(OH)2
# Enthalpy of formation: -568.525 kJ/mol
-analytic -8.6666e+001 -1.8440e-002 7.5723e+003 3.2597e+001 1.1818e+002
# -Range: 0-300
Fe(OH)3
Fe(OH)3 +3.0000 H+ = + 1.0000 Fe+++ + 3.0000 H2O
log_k 5.6556
-delta_H -84.0824 kJ/mol # Calculated enthalpy of reaction Fe(OH)3
# Enthalpy of formation: -823.013 kJ/mol
-analytic -1.3316e+002 -3.1284e-002 7.9753e+003 4.9052e+001 1.2449e+002
# -Range: 0-300
Fe2(SO4)3
Fe2(SO4)3 = + 2.0000 Fe+++ + 3.0000 SO4--
log_k 3.2058
-delta_H -250.806 kJ/mol # Calculated enthalpy of reaction Fe2(SO4)3
# Enthalpy of formation: -2577.16 kJ/mol
-analytic -5.8649e+002 -2.3718e-001 2.2736e+004 2.3601e+002 3.5495e+002
# -Range: 0-300
FeF2
FeF2 = + 1.0000 Fe++ + 2.0000 F-
log_k -2.3817
-delta_H -51.6924 kJ/mol # Calculated enthalpy of reaction FeF2
# Enthalpy of formation: -711.26 kJ/mol
-analytic -2.5687e+002 -8.4091e-002 8.4262e+003 1.0154e+002 1.3156e+002
# -Range: 0-300
FeF3
FeF3 = + 1.0000 Fe+++ + 3.0000 F-
log_k -19.2388
-delta_H -13.8072 kJ/mol # Calculated enthalpy of reaction FeF3
# Enthalpy of formation: -249 kcal/mol
-analytic -1.6215e+001 -3.7450e-002 -1.8926e+003 5.8485e+000 -3.2134e+001
# -Range: 0-200
FeO
FeO +2.0000 H+ = + 1.0000 Fe++ + 1.0000 H2O
log_k 13.5318
-delta_H -106.052 kJ/mol # Calculated enthalpy of reaction FeO
# Enthalpy of formation: -65.02 kcal/mol
-analytic -7.8750e+001 -1.8268e-002 7.6852e+003 2.9074e+001 1.1994e+002
# -Range: 0-300
FeSO4
FeSO4 = + 1.0000 Fe++ + 1.0000 SO4--
log_k 2.6565
-delta_H -73.0878 kJ/mol # Calculated enthalpy of reaction FeSO4
# Enthalpy of formation: -928.771 kJ/mol
-analytic -2.0794e+002 -7.6891e-002 7.8705e+003 8.3685e+001 1.2287e+002
# -Range: 0-300
FeV2O4
FeV2O4 +8.0000 H+ = + 1.0000 Fe++ + 2.0000 V+++ + 4.0000 H2O
log_k 280.5528
-delta_H -1733.42 kJ/mol # Calculated enthalpy of reaction FeV2O4
# Enthalpy of formation: -5.8 kcal/mol
-analytic -1.6736e+002 -1.9398e-002 9.5736e+004 5.3582e+001 1.6258e+003
# -Range: 0-200
Ferrite-Ca
CaFe2O4 +8.0000 H+ = + 1.0000 Ca++ + 2.0000 Fe+++ + 4.0000 H2O
log_k 21.5217
-delta_H -264.738 kJ/mol # Calculated enthalpy of reaction Ferrite-Ca
# Enthalpy of formation: -363.494 kcal/mol
-analytic -2.8472e+002 -7.5870e-002 2.0688e+004 1.0485e+002 3.2289e+002
# -Range: 0-300
Ferrite-Cu
CuFe2O4 +8.0000 H+ = + 1.0000 Cu++ + 2.0000 Fe+++ + 4.0000 H2O
log_k 10.3160
-delta_H -211.647 kJ/mol # Calculated enthalpy of reaction Ferrite-Cu
# Enthalpy of formation: -965.178 kJ/mol
-analytic -3.1271e+002 -7.9976e-002 1.8818e+004 1.1466e+002 2.9374e+002
# -Range: 0-300
Ferrite-Dicalcium
Ca2Fe2O5 +10.0000 H+ = + 2.0000 Ca++ + 2.0000 Fe+++ + 5.0000 H2O
log_k 56.8331
-delta_H -475.261 kJ/mol # Calculated enthalpy of reaction Ferrite-Dicalcium
# Enthalpy of formation: -2139.26 kJ/mol
-analytic -3.6277e+002 -9.5015e-002 3.3898e+004 1.3506e+002 5.2906e+002
# -Range: 0-300
Ferrite-Mg
MgFe2O4 +8.0000 H+ = + 1.0000 Mg++ + 2.0000 Fe+++ + 4.0000 H2O
log_k 21.0551
-delta_H -280.056 kJ/mol # Calculated enthalpy of reaction Ferrite-Mg
# Enthalpy of formation: -1428.42 kJ/mol
-analytic -2.8297e+002 -7.4820e-002 2.1333e+004 1.0295e+002 3.3296e+002
# -Range: 0-300
Ferrite-Zn
ZnFe2O4 +8.0000 H+ = + 1.0000 Zn++ + 2.0000 Fe+++ + 4.0000 H2O
log_k 11.7342
-delta_H -226.609 kJ/mol # Calculated enthalpy of reaction Ferrite-Zn
# Enthalpy of formation: -1169.29 kJ/mol
-analytic -2.9809e+002 -7.7263e-002 1.9067e+004 1.0866e+002 2.9761e+002
# -Range: 0-300
Ferroselite
FeSe2 +0.5000 H2O = + 0.2500 O2 + 1.0000 Fe+++ + 1.0000 H+ + 2.0000 Se--
log_k -80.7998
-delta_H 0 # Not possible to calculate enthalpy of reaction Ferroselite
# Enthalpy of formation: -25 kcal/mol
-analytic -7.2971e+001 -2.4992e-002 -1.6246e+004 2.1860e+001 -2.5348e+002
# -Range: 0-300
Ferrosilite
FeSiO3 +2.0000 H+ = + 1.0000 Fe++ + 1.0000 H2O + 1.0000 SiO2
log_k 7.4471
-delta_H -60.6011 kJ/mol # Calculated enthalpy of reaction Ferrosilite
# Enthalpy of formation: -285.658 kcal/mol
-analytic 9.0041e+000 3.7917e-003 5.1625e+003 -6.3009e+000 -3.9565e+005
# -Range: 0-300
Fluorapatite
Ca5(PO4)3F +3.0000 H+ = + 1.0000 F- + 3.0000 HPO4-- + 5.0000 Ca++
log_k -24.9940
-delta_H -90.8915 kJ/mol # Calculated enthalpy of reaction Fluorapatite
# Enthalpy of formation: -6836.12 kJ/mol
-analytic -9.3648e+002 -3.2688e-001 2.4398e+004 3.7461e+002 3.8098e+002
# -Range: 0-300
Fluorite
CaF2 = + 1.0000 Ca++ + 2.0000 F-
log_k -10.0370
-delta_H 12.1336 kJ/mol # Calculated enthalpy of reaction Fluorite
# Enthalpy of formation: -293 kcal/mol
-analytic -2.5036e+002 -8.4183e-002 4.9525e+003 1.0054e+002 7.7353e+001
# -Range: 0-300
Forsterite
Mg2SiO4 +4.0000 H+ = + 1.0000 SiO2 + 2.0000 H2O + 2.0000 Mg++
log_k 27.8626
-delta_H -205.614 kJ/mol # Calculated enthalpy of reaction Forsterite
# Enthalpy of formation: -520 kcal/mol
-analytic -7.6195e+001 -1.4013e-002 1.4763e+004 2.5090e+001 -3.0379e+005
# -Range: 0-300
Foshagite
Ca4Si3O9(OH)2:0.5H2O +8.0000 H+ = + 3.0000 SiO2 + 4.0000 Ca++ + 5.5000 H2O
log_k 65.9210
-delta_H -359.839 kJ/mol # Calculated enthalpy of reaction Foshagite
# Enthalpy of formation: -1438.27 kcal/mol
-analytic 2.9983e+001 5.5272e-003 2.3427e+004 -1.3879e+001 -8.9461e+005
# -Range: 0-300
Frankdicksonite
BaF2 = + 1.0000 Ba++ + 2.0000 F-
log_k -5.7600
-delta_H 0 # Not possible to calculate enthalpy of reaction Frankdicksonite
# Enthalpy of formation: 0 kcal/mol
Freboldite
CoSe = + 1.0000 Co++ + 1.0000 Se--
log_k -24.3358
-delta_H 0 # Not possible to calculate enthalpy of reaction Freboldite
# Enthalpy of formation: -15.295 kcal/mol
-analytic -1.3763e+001 -1.6924e-003 -3.6938e+003 9.3574e-001 -6.2723e+001
# -Range: 0-200
Ga
Ga +3.0000 H+ +0.7500 O2 = + 1.0000 Ga+++ + 1.5000 H2O
log_k 92.3567
-delta_H -631.368 kJ/mol # Calculated enthalpy of reaction Ga
# Enthalpy of formation: 0 kJ/mol
-analytic -1.3027e+002 -3.9539e-002 3.6027e+004 4.6280e+001 -8.5461e+004
# -Range: 0-300
Galena
PbS +1.0000 H+ = + 1.0000 HS- + 1.0000 Pb++
log_k -14.8544
-delta_H 83.1361 kJ/mol # Calculated enthalpy of reaction Galena
# Enthalpy of formation: -23.5 kcal/mol
-analytic -1.2124e+002 -4.3477e-002 -1.6463e+003 5.0454e+001 -2.5654e+001
# -Range: 0-300
Gaylussite
CaNa2(CO3)2:5H2O +2.0000 H+ = + 1.0000 Ca++ + 2.0000 HCO3- + 2.0000 Na+ + 5.0000 H2O
log_k 11.1641
-delta_H 0 # Not possible to calculate enthalpy of reaction Gaylussite
# Enthalpy of formation: 0 kcal/mol
Gd
Gd +3.0000 H+ +0.7500 O2 = + 1.0000 Gd+++ + 1.5000 H2O
log_k 180.7573
-delta_H -1106.67 kJ/mol # Calculated enthalpy of reaction Gd
# Enthalpy of formation: 0 kJ/mol
-analytic -3.3949e+002 -6.5698e-002 7.4278e+004 1.2189e+002 -9.7055e+005
# -Range: 0-300
Gd(OH)3
Gd(OH)3 +3.0000 H+ = + 1.0000 Gd+++ + 3.0000 H2O
log_k 15.5852
-delta_H 0 # Not possible to calculate enthalpy of reaction Gd(OH)3
# Enthalpy of formation: 0 kcal/mol
Gd(OH)3(am)
Gd(OH)3 +3.0000 H+ = + 1.0000 Gd+++ + 3.0000 H2O
log_k 17.9852
-delta_H 0 # Not possible to calculate enthalpy of reaction Gd(OH)3(am)
# Enthalpy of formation: 0 kcal/mol
Gd2(CO3)3
Gd2(CO3)3 +3.0000 H+ = + 2.0000 Gd+++ + 3.0000 HCO3-
log_k -3.7136
-delta_H 0 # Not possible to calculate enthalpy of reaction Gd2(CO3)3
# Enthalpy of formation: 0 kcal/mol
Gd2O3
Gd2O3 +6.0000 H+ = + 2.0000 Gd+++ + 3.0000 H2O
log_k 53.8000
-delta_H 0 # Not possible to calculate enthalpy of reaction Gd2O3
# Enthalpy of formation: 0 kcal/mol
GdF3:.5H2O
GdF3:.5H2O = + 0.5000 H2O + 1.0000 Gd+++ + 3.0000 F-
log_k -16.9000
-delta_H 0 # Not possible to calculate enthalpy of reaction GdF3:.5H2O
# Enthalpy of formation: 0 kcal/mol
GdPO4:10H2O
GdPO4:10H2O +1.0000 H+ = + 1.0000 Gd+++ + 1.0000 HPO4-- + 10.0000 H2O
log_k -11.9782
-delta_H 0 # Not possible to calculate enthalpy of reaction GdPO4:10H2O
# Enthalpy of formation: 0 kcal/mol
Gehlenite
Ca2Al2SiO7 +10.0000 H+ = + 1.0000 SiO2 + 2.0000 Al+++ + 2.0000 Ca++ + 5.0000 H2O
log_k 56.2997
-delta_H -489.934 kJ/mol # Calculated enthalpy of reaction Gehlenite
# Enthalpy of formation: -951.225 kcal/mol
-analytic -2.1784e+002 -6.7200e-002 2.9779e+004 7.8488e+001 4.6473e+002
# -Range: 0-300
Gibbsite
Al(OH)3 +3.0000 H+ = + 1.0000 Al+++ + 3.0000 H2O
log_k 7.7560
-delta_H -102.788 kJ/mol # Calculated enthalpy of reaction Gibbsite
# Enthalpy of formation: -309.065 kcal/mol
-analytic -1.1403e+002 -3.6453e-002 7.7236e+003 4.3134e+001 1.2055e+002
# -Range: 0-300
Gismondine
Ca2Al4Si4O16:9H2O +16.0000 H+ = + 2.0000 Ca++ + 4.0000 Al+++ + 4.0000 SiO2 + 17.0000 H2O
log_k 41.7170
-delta_H 0 # Not possible to calculate enthalpy of reaction Gismondine
# Enthalpy of formation: 0 kcal/mol
Glauberite
Na2Ca(SO4)2 = + 1.0000 Ca++ + 2.0000 Na+ + 2.0000 SO4--
log_k -5.4690
-delta_H 0 # Not possible to calculate enthalpy of reaction Glauberite
# Enthalpy of formation: 0 kcal/mol
Goethite
FeOOH +3.0000 H+ = + 1.0000 Fe+++ + 2.0000 H2O
log_k 0.5345
-delta_H -61.9291 kJ/mol # Calculated enthalpy of reaction Goethite
# Enthalpy of formation: -559.328 kJ/mol
-analytic -6.0331e+001 -1.0847e-002 4.7759e+003 1.9429e+001 8.1122e+001
# -Range: 0-200
Greenalite
Fe3Si2O5(OH)4 +6.0000 H+ = + 2.0000 SiO2 + 3.0000 Fe++ + 5.0000 H2O
log_k 22.6701
-delta_H -165.297 kJ/mol # Calculated enthalpy of reaction Greenalite
# Enthalpy of formation: -787.778 kcal/mol
-analytic -1.4187e+001 -3.8377e-003 1.1710e+004 1.6442e+000 -4.8290e+005
# -Range: 0-300
Grossular
Ca3Al2(SiO4)3 +12.0000 H+ = + 2.0000 Al+++ + 3.0000 Ca++ + 3.0000 SiO2 + 6.0000 H2O
log_k 51.9228
-delta_H -432.006 kJ/mol # Calculated enthalpy of reaction Grossular
# Enthalpy of formation: -1582.74 kcal/mol
-analytic 2.9389e+001 -2.2478e-002 2.0323e+004 -1.4624e+001 -2.5674e+005
# -Range: 0-300
Gypsum
CaSO4:2H2O = + 1.0000 Ca++ + 1.0000 SO4-- + 2.0000 H2O
log_k -4.4823
-delta_H -1.66746 kJ/mol # Calculated enthalpy of reaction Gypsum
# Enthalpy of formation: -2022.69 kJ/mol
-analytic -2.4417e+002 -8.3329e-002 5.5958e+003 9.9301e+001 8.7389e+001
# -Range: 0-300
Gyrolite
Ca2Si3O7(OH)2:1.5H2O +4.0000 H+ = + 2.0000 Ca++ + 3.0000 SiO2 + 4.5000 H2O
log_k 22.9099
-delta_H -82.862 kJ/mol # Calculated enthalpy of reaction Gyrolite
# Enthalpy of formation: -1176.55 kcal/mol
-analytic -2.4416e+001 1.4646e-002 1.6181e+004 2.3723e+000 -1.5369e+006
# -Range: 0-300
HTcO4
HTcO4 = + 1.0000 H+ + 1.0000 TcO4-
log_k 5.9566
-delta_H -12.324 kJ/mol # Calculated enthalpy of reaction HTcO4
# Enthalpy of formation: -703.945 kJ/mol
-analytic 3.0005e+001 7.6416e-003 -5.3546e+001 -1.0568e+001 -9.1953e-001
# -Range: 0-200
Haiweeite
Ca(UO2)2(Si2O5)3:5H2O +6.0000 H+ = + 1.0000 Ca++ + 2.0000 UO2++ + 6.0000 SiO2 + 8.0000 H2O
log_k -7.0413
-delta_H 0 # Not possible to calculate enthalpy of reaction Haiweeite
# Enthalpy of formation: 0 kcal/mol
Halite
NaCl = + 1.0000 Cl- + 1.0000 Na+
log_k 1.5855
-delta_H 3.7405 kJ/mol # Calculated enthalpy of reaction Halite
# Enthalpy of formation: -98.26 kcal/mol
-analytic -1.0163e+002 -3.4761e-002 2.2796e+003 4.2802e+001 3.5602e+001
# -Range: 0-300
Hatrurite
Ca3SiO5 +6.0000 H+ = + 1.0000 SiO2 + 3.0000 Ca++ + 3.0000 H2O
log_k 73.4056
-delta_H -434.684 kJ/mol # Calculated enthalpy of reaction Hatrurite
# Enthalpy of formation: -700.234 kcal/mol
-analytic -4.5448e+001 -1.9998e-002 2.3800e+004 1.8494e+001 -7.3385e+004
# -Range: 0-300
Hausmannite
Mn3O4 +8.0000 H+ = + 1.0000 Mn++ + 2.0000 Mn+++ + 4.0000 H2O
log_k 10.1598
-delta_H -268.121 kJ/mol # Calculated enthalpy of reaction Hausmannite
# Enthalpy of formation: -1387.83 kJ/mol
-analytic -2.0600e+002 -2.2214e-002 2.0160e+004 6.2700e+001 3.1464e+002
# -Range: 0-300
Heazlewoodite
Ni3S2 +4.0000 H+ +0.5000 O2 = + 1.0000 H2O + 2.0000 HS- + 3.0000 Ni++
log_k 28.2477
-delta_H -270.897 kJ/mol # Calculated enthalpy of reaction Heazlewoodite
# Enthalpy of formation: -203.012 kJ/mol
-analytic -3.5439e+002 -1.1740e-001 2.1811e+004 1.3919e+002 3.4044e+002
# -Range: 0-300
Hedenbergite
CaFe(SiO3)2 +4.0000 H+ = + 1.0000 Ca++ + 1.0000 Fe++ + 2.0000 H2O + 2.0000 SiO2
log_k 19.6060
-delta_H -124.507 kJ/mol # Calculated enthalpy of reaction Hedenbergite
# Enthalpy of formation: -678.276 kcal/mol
-analytic -1.9473e+001 1.5288e-003 1.2910e+004 2.1729e+000 -9.0058e+005
# -Range: 0-300
Hematite
Fe2O3 +6.0000 H+ = + 2.0000 Fe+++ + 3.0000 H2O
log_k 0.1086
-delta_H -129.415 kJ/mol # Calculated enthalpy of reaction Hematite
# Enthalpy of formation: -197.72 kcal/mol
-analytic -2.2015e+002 -6.0290e-002 1.1812e+004 8.0253e+001 1.8438e+002
# -Range: 0-300
Hercynite
FeAl2O4 +8.0000 H+ = + 1.0000 Fe++ + 2.0000 Al+++ + 4.0000 H2O
log_k 28.8484
-delta_H -345.961 kJ/mol # Calculated enthalpy of reaction Hercynite
# Enthalpy of formation: -1966.45 kJ/mol
-analytic -3.1848e+002 -7.9501e-002 2.5892e+004 1.1483e+002 4.0412e+002
# -Range: 0-300
Herzenbergite
SnS +1.0000 H+ = + 1.0000 HS- + 1.0000 Sn++
log_k -15.5786
-delta_H 81.6466 kJ/mol # Calculated enthalpy of reaction Herzenbergite
# Enthalpy of formation: -25.464 kcal/mol
-analytic -1.3576e+002 -4.6594e-002 -1.1572e+003 5.5740e+001 -1.8018e+001
# -Range: 0-300
Heulandite
# Ba.065Sr.175Ca.585K.132Na.383Al2.165Si6.835O18:6 +8.6600 H+ = + 0.0650 Ba++ + 0.1320 K+ + 0.1750 Sr++ + 0.3830 Na+ + 0.5850 Ca++ + 2.1650 Al+++ + 6.8350 SiO2 + 10.3300 H2O
Ba.065Sr.175Ca.585K.132Na.383Al2.165Si6.835O18:6H2O +8.6600 H+ = + 0.0650 Ba++ + 0.1320 K+ + 0.1750 Sr++ + 0.3830 Na+ + 0.5850 Ca++ + 2.1650 Al+++ + 6.8350 SiO2 + 10.3300 H2O
log_k 3.3506
-delta_H -97.2942 kJ/mol # Calculated enthalpy of reaction Heulandite
# Enthalpy of formation: -10594.5 kJ/mol
-analytic -1.8364e+001 2.7879e-002 2.8426e+004 -1.7427e+001 -3.4723e+006
# -Range: 0-300
Hexahydrite
MgSO4:6H2O = + 1.0000 Mg++ + 1.0000 SO4-- + 6.0000 H2O
log_k -1.7268
-delta_H 0 # Not possible to calculate enthalpy of reaction Hexahydrite
# Enthalpy of formation: 0 kcal/mol
Hf(s)
Hf +4.0000 H+ +1.0000 O2 = + 1.0000 Hf++++ + 2.0000 H2O
log_k 189.9795
-delta_H 0 # Not possible to calculate enthalpy of reaction Hf
# Enthalpy of formation: -0.003 kJ/mol
HfB2
HfB2 +2.7500 H+ +2.2500 H2O = + 0.7500 B(OH)3 + 1.0000 Hf++++ + 1.2500 BH4-
log_k 55.7691
-delta_H 0 # Not possible to calculate enthalpy of reaction HfB2
# Enthalpy of formation: -78.6 kJ/mol
HfBr2
HfBr2 +2.0000 H+ +0.5000 O2 = + 1.0000 H2O + 1.0000 Hf++++ + 2.0000 Br-
log_k 114.9446
-delta_H 0 # Not possible to calculate enthalpy of reaction HfBr2
# Enthalpy of formation: -98 kJ/mol
HfBr4
HfBr4 = + 1.0000 Hf++++ + 4.0000 Br-
log_k 48.2921
-delta_H 0 # Not possible to calculate enthalpy of reaction HfBr4
# Enthalpy of formation: -183.1 kJ/mol
HfC
HfC +3.0000 H+ +2.0000 O2 = + 1.0000 H2O + 1.0000 HCO3- + 1.0000 Hf++++
log_k 215.0827
-delta_H 0 # Not possible to calculate enthalpy of reaction HfC
# Enthalpy of formation: -54 kJ/mol
HfCl2
HfCl2 +2.0000 H+ +0.5000 O2 = + 1.0000 H2O + 1.0000 Hf++++ + 2.0000 Cl-
log_k 109.1624
-delta_H 0 # Not possible to calculate enthalpy of reaction HfCl2
# Enthalpy of formation: -125 kJ/mol
HfCl4
HfCl4 = + 1.0000 Hf++++ + 4.0000 Cl-
log_k 38.0919
-delta_H 0 # Not possible to calculate enthalpy of reaction HfCl4
# Enthalpy of formation: -236.7 kJ/mol
HfF2
HfF2 +2.0000 H+ +0.5000 O2 = + 1.0000 H2O + 1.0000 Hf++++ + 2.0000 F-
log_k 81.7647
-delta_H 0 # Not possible to calculate enthalpy of reaction HfF2
# Enthalpy of formation: -235 kJ/mol
HfF4
HfF4 = + 1.0000 Hf++++ + 4.0000 F-
log_k -19.2307
-delta_H 0 # Not possible to calculate enthalpy of reaction HfF4
# Enthalpy of formation: -461.4 kJ/mol
HfI2
HfI2 +2.0000 H+ +0.5000 O2 = + 1.0000 H2O + 1.0000 Hf++++ + 2.0000 I-
log_k 117.4971
-delta_H 0 # Not possible to calculate enthalpy of reaction HfI2
# Enthalpy of formation: -65 kJ/mol
HfI4
HfI4 = + 1.0000 Hf++++ + 4.0000 I-
log_k 54.1798
-delta_H 0 # Not possible to calculate enthalpy of reaction HfI4
# Enthalpy of formation: -118 kJ/mol
HfN
HfN +4.0000 H+ +0.2500 O2 = + 0.5000 H2O + 1.0000 Hf++++ + 1.0000 NH3
log_k 69.4646
-delta_H 0 # Not possible to calculate enthalpy of reaction HfN
# Enthalpy of formation: -89.3 kJ/mol
HfO2
HfO2 +4.0000 H+ = + 1.0000 Hf++++ + 2.0000 H2O
log_k 1.1829
-delta_H 0 # Not possible to calculate enthalpy of reaction HfO2
# Enthalpy of formation: -267.1 kJ/mol
HfS2
HfS2 +2.0000 H+ = + 1.0000 Hf++++ + 2.0000 HS-
log_k -1.5845
-delta_H 0 # Not possible to calculate enthalpy of reaction HfS2
# Enthalpy of formation: -140 kJ/mol
HfS3
HfS3 +1.0000 H+ = + 1.0000 HS- + 1.0000 Hf++++ + 1.0000 S2--
log_k -18.9936
-delta_H 0 # Not possible to calculate enthalpy of reaction HfS3
# Enthalpy of formation: -149 kJ/mol
Hg2SO4
Hg2SO4 = + 1.0000 Hg2++ + 1.0000 SO4--
log_k -6.1170
-delta_H 0.30448 kJ/mol # Calculated enthalpy of reaction Hg2SO4
# Enthalpy of formation: -743.09 kJ/mol
-analytic -3.2342e+001 -1.9881e-002 1.6292e+003 1.0781e+001 2.7677e+001
# -Range: 0-200
Hg2SeO3
Hg2SeO3 = + 1.0000 Hg2++ + 1.0000 SeO3--
log_k -14.2132
-delta_H 0 # Not possible to calculate enthalpy of reaction Hg2SeO3
# Enthalpy of formation: 0 kcal/mol
HgSeO3
HgSeO3 = + 1.0000 Hg++ + 1.0000 SeO3--
log_k -13.8957
-delta_H 0 # Not possible to calculate enthalpy of reaction HgSeO3
# Enthalpy of formation: 0 kcal/mol
Hillebrandite
Ca2SiO3(OH)2:0.17H2O +4.0000 H+ = + 1.0000 SiO2 + 2.0000 Ca++ + 3.1700 H2O
log_k 36.8190
-delta_H -203.074 kJ/mol # Calculated enthalpy of reaction Hillebrandite
# Enthalpy of formation: -637.404 kcal/mol
-analytic -1.9360e+001 -7.5176e-003 1.1947e+004 8.0558e+000 -1.4504e+005
# -Range: 0-300
Hinsdalite
Al3PPbSO8(OH)6 +7.0000 H+ = + 1.0000 HPO4-- + 1.0000 Pb++ + 1.0000 SO4-- + 3.0000 Al+++ + 6.0000 H2O
log_k 9.8218
-delta_H 0 # Not possible to calculate enthalpy of reaction Hinsdalite
# Enthalpy of formation: 0 kcal/mol
Ho
Ho +3.0000 H+ +0.7500 O2 = + 1.0000 Ho+++ + 1.5000 H2O
log_k 182.8097
-delta_H -1126.75 kJ/mol # Calculated enthalpy of reaction Ho
# Enthalpy of formation: 0 kJ/mol
-analytic -6.5903e+001 -2.8190e-002 5.9370e+004 2.3421e+001 9.2643e+002
# -Range: 0-300
Ho(OH)3
Ho(OH)3 +3.0000 H+ = + 1.0000 Ho+++ + 3.0000 H2O
log_k 15.3852
-delta_H 0 # Not possible to calculate enthalpy of reaction Ho(OH)3
# Enthalpy of formation: 0 kcal/mol
Ho(OH)3(am)
Ho(OH)3 +3.0000 H+ = + 1.0000 Ho+++ + 3.0000 H2O
log_k 17.7852
-delta_H 0 # Not possible to calculate enthalpy of reaction Ho(OH)3(am)
# Enthalpy of formation: 0 kcal/mol
Ho2(CO3)3
Ho2(CO3)3 +3.0000 H+ = + 2.0000 Ho+++ + 3.0000 HCO3-
log_k -2.8136
-delta_H 0 # Not possible to calculate enthalpy of reaction Ho2(CO3)3
# Enthalpy of formation: 0 kcal/mol
Ho2O3
Ho2O3 +6.0000 H+ = + 2.0000 Ho+++ + 3.0000 H2O
log_k 47.3000
-delta_H 0 # Not possible to calculate enthalpy of reaction Ho2O3
# Enthalpy of formation: 0 kcal/mol
HoF3:.5H2O
HoF3:.5H2O = + 0.5000 H2O + 1.0000 Ho+++ + 3.0000 F-
log_k -16.4000
-delta_H 0 # Not possible to calculate enthalpy of reaction HoF3:.5H2O
# Enthalpy of formation: 0 kcal/mol
HoPO4:10H2O
HoPO4:10H2O +1.0000 H+ = + 1.0000 HPO4-- + 1.0000 Ho+++ + 10.0000 H2O
log_k -11.8782
-delta_H 0 # Not possible to calculate enthalpy of reaction HoPO4:10H2O
# Enthalpy of formation: 0 kcal/mol
Hopeite
Zn3(PO4)2:4H2O +2.0000 H+ = + 2.0000 HPO4-- + 3.0000 Zn++ + 4.0000 H2O
log_k -10.6563
-delta_H 0 # Not possible to calculate enthalpy of reaction Hopeite
# Enthalpy of formation: 0 kcal/mol
Huntite
CaMg3(CO3)4 +4.0000 H+ = + 1.0000 Ca++ + 3.0000 Mg++ + 4.0000 HCO3-
log_k 10.3010
-delta_H -171.096 kJ/mol # Calculated enthalpy of reaction Huntite
# Enthalpy of formation: -1082.6 kcal/mol
-analytic -6.5000e+002 -1.9671e-001 2.4815e+004 2.5688e+002 3.8740e+002
# -Range: 0-300
Hydroboracite
MgCaB6O11:6H2O +4.0000 H+ +1.0000 H2O = + 1.0000 Ca++ + 1.0000 Mg++ + 6.0000 B(OH)3
log_k 20.3631
-delta_H 0 # Not possible to calculate enthalpy of reaction Hydroboracite
# Enthalpy of formation: 0 kcal/mol
Hydrocerussite
Pb3(CO3)2(OH)2 +4.0000 H+ = + 2.0000 H2O + 2.0000 HCO3- + 3.0000 Pb++
log_k 1.8477
-delta_H 0 # Not possible to calculate enthalpy of reaction Hydrocerussite
# Enthalpy of formation: 0 kcal/mol
Hydromagnesite
Mg5(CO3)4(OH)2:4H2O +6.0000 H+ = + 4.0000 HCO3- + 5.0000 Mg++ + 6.0000 H2O
log_k 30.8539
-delta_H -289.696 kJ/mol # Calculated enthalpy of reaction Hydromagnesite
# Enthalpy of formation: -1557.09 kcal/mol
-analytic -7.9288e+002 -2.1448e-001 3.6749e+004 3.0888e+002 5.7367e+002
# -Range: 0-300
Hydrophilite
CaCl2 = + 1.0000 Ca++ + 2.0000 Cl-
log_k 11.7916
-delta_H -81.4545 kJ/mol # Calculated enthalpy of reaction Hydrophilite
# Enthalpy of formation: -795.788 kJ/mol
-analytic -2.2278e+002 -8.1414e-002 9.0298e+003 9.2349e+001 1.4097e+002
# -Range: 0-300
Hydroxylapatite
Ca5(OH)(PO4)3 +4.0000 H+ = + 1.0000 H2O + 3.0000 HPO4-- + 5.0000 Ca++
log_k -3.0746
-delta_H -191.982 kJ/mol # Calculated enthalpy of reaction Hydroxylapatite
# Enthalpy of formation: -6685.52 kJ/mol
-analytic -8.5221e+002 -2.9430e-001 2.8125e+004 3.4044e+002 4.3911e+002
# -Range: 0-300
Hydrozincite
Zn5(OH)6(CO3)2 +8.0000 H+ = + 2.0000 HCO3- + 5.0000 Zn++ + 6.0000 H2O
log_k 30.3076
-delta_H 0 # Not possible to calculate enthalpy of reaction Hydrozincite
# Enthalpy of formation: 0 kcal/mol
I2
I2 +1.0000 H2O = + 0.5000 O2 + 2.0000 H+ + 2.0000 I-
log_k -24.8084
-delta_H 165.967 kJ/mol # Calculated enthalpy of reaction I2
# Enthalpy of formation: 0 kJ/mol
-analytic -1.7135e+002 -6.2810e-002 -4.7225e+003 7.3181e+001 -7.3640e+001
# -Range: 0-300
Ice
H2O = + 1.0000 H2O
log_k 0.1387
-delta_H 6.74879 kJ/mol # Calculated enthalpy of reaction Ice
# Enthalpy of formation: -69.93 kcal/mol
-analytic -2.3260e+001 4.7948e-004 7.7351e+002 8.3499e+000 1.3143e+001
# -Range: 0-200
Illite
K0.6Mg0.25Al1.8Al0.5Si3.5O10(OH)2 +8.0000 H+ = + 0.2500 Mg++ + 0.6000 K+ + 2.3000 Al+++ + 3.5000 SiO2 + 5.0000 H2O
log_k 9.0260
-delta_H -171.764 kJ/mol # Calculated enthalpy of reaction Illite
# Enthalpy of formation: -1394.71 kcal/mol
-analytic 2.6069e+001 -1.2553e-003 1.3670e+004 -2.0232e+001 -1.1204e+006
# -Range: 0-300
Ilmenite
FeTiO3 +2.0000 H+ +1.0000 H2O = + 1.0000 Fe++ + 1.0000 Ti(OH)4
log_k 0.9046
-delta_H 0 # Not possible to calculate enthalpy of reaction Ilmenite
# Enthalpy of formation: -1236.65 kJ/mol
In
In +3.0000 H+ +0.7500 O2 = + 1.0000 In+++ + 1.5000 H2O
log_k 81.6548
-delta_H -524.257 kJ/mol # Calculated enthalpy of reaction In
# Enthalpy of formation: 0 kJ/mol
-analytic -1.1773e+002 -3.7657e-002 3.1802e+004 4.2438e+001 -9.6348e+004
# -Range: 0-300
Jadeite
NaAl(SiO3)2 +4.0000 H+ = + 1.0000 Al+++ + 1.0000 Na+ + 2.0000 H2O + 2.0000 SiO2
log_k 8.3888
-delta_H -84.4415 kJ/mol # Calculated enthalpy of reaction Jadeite
# Enthalpy of formation: -722.116 kcal/mol
-analytic 1.5934e+000 5.0757e-003 9.5602e+003 -7.0164e+000 -8.4454e+005
# -Range: 0-300
Jarosite
KFe3(SO4)2(OH)6 +6.0000 H+ = + 1.0000 K+ + 2.0000 SO4-- + 3.0000 Fe+++ + 6.0000 H2O
log_k -9.3706
-delta_H -191.343 kJ/mol # Calculated enthalpy of reaction Jarosite
# Enthalpy of formation: -894.79 kcal/mol
-analytic -1.0813e+002 -5.0381e-002 9.6893e+003 3.2832e+001 1.6457e+002
# -Range: 0-200
Jarosite-Na
NaFe3(SO4)2(OH)6 +6.0000 H+ = + 1.0000 Na+ + 2.0000 SO4-- + 3.0000 Fe+++ + 6.0000 H2O
log_k -5.4482
-delta_H 0 # Not possible to calculate enthalpy of reaction Jarosite-Na
# Enthalpy of formation: 0 kcal/mol
K
K +1.0000 H+ +0.2500 O2 = + 0.5000 H2O + 1.0000 K+
log_k 70.9861
-delta_H -392.055 kJ/mol # Calculated enthalpy of reaction K
# Enthalpy of formation: 0 kJ/mol
-analytic -3.1102e+001 -1.0003e-002 2.1338e+004 1.3534e+001 3.3296e+002
# -Range: 0-300
K-Feldspar
KAlSi3O8 +4.0000 H+ = + 1.0000 Al+++ + 1.0000 K+ + 2.0000 H2O + 3.0000 SiO2
log_k -0.2753
-delta_H -23.9408 kJ/mol # Calculated enthalpy of reaction K-Feldspar
# Enthalpy of formation: -949.188 kcal/mol
-analytic -1.0684e+000 1.3111e-002 1.1671e+004 -9.9129e+000 -1.5855e+006
# -Range: 0-300
K2CO3:1.5H2O
K2CO3:1.5H2O +1.0000 H+ = + 1.0000 HCO3- + 1.5000 H2O + 2.0000 K+
log_k 13.3785
-delta_H 0 # Not possible to calculate enthalpy of reaction K2CO3:1.5H2O
# Enthalpy of formation: 0 kcal/mol
K2O
K2O +2.0000 H+ = + 1.0000 H2O + 2.0000 K+
log_k 84.0405
-delta_H -427.006 kJ/mol # Calculated enthalpy of reaction K2O
# Enthalpy of formation: -86.8 kcal/mol
-analytic -1.8283e+001 -5.2255e-003 2.3184e+004 1.0553e+001 3.6177e+002
# -Range: 0-300
K2Se
K2Se = + 1.0000 Se-- + 2.0000 K+
log_k 11.2925
-delta_H 0 # Not possible to calculate enthalpy of reaction K2Se
# Enthalpy of formation: -92 kcal/mol
-analytic 1.8182e+001 7.8828e-003 2.6345e+003 -7.3075e+000 4.4732e+001
# -Range: 0-200
K2UO4
K2UO4 +4.0000 H+ = + 1.0000 UO2++ + 2.0000 H2O + 2.0000 K+
log_k 33.8714
-delta_H -174.316 kJ/mol # Calculated enthalpy of reaction K2UO4
# Enthalpy of formation: -1920.7 kJ/mol
-analytic -7.0905e+001 -2.5680e-003 1.2244e+004 2.6056e+001 2.0794e+002
# -Range: 0-200
K3H(SO4)2
K3H(SO4)2 = + 1.0000 H+ + 2.0000 SO4-- + 3.0000 K+
log_k -3.6233
-delta_H 0 # Not possible to calculate enthalpy of reaction K3H(SO4)2
# Enthalpy of formation: 0 kcal/mol
K8H4(CO3)6:3H2O
K8H4(CO3)6:3H2O +2.0000 H+ = + 3.0000 H2O + 6.0000 HCO3- + 8.0000 K+
log_k 27.7099
-delta_H 0 # Not possible to calculate enthalpy of reaction K8H4(CO3)6:3H2O
# Enthalpy of formation: 0 kcal/mol
KAl(SO4)2
KAl(SO4)2 = + 1.0000 Al+++ + 1.0000 K+ + 2.0000 SO4--
log_k 3.3647
-delta_H -139.485 kJ/mol # Calculated enthalpy of reaction KAl(SO4)2
# Enthalpy of formation: -2470.29 kJ/mol
-analytic -4.2785e+002 -1.6303e-001 1.5311e+004 1.7312e+002 2.3904e+002
# -Range: 0-300
KBr
KBr = + 1.0000 Br- + 1.0000 K+
log_k 1.0691
-delta_H 20.125 kJ/mol # Calculated enthalpy of reaction KBr
# Enthalpy of formation: -393.798 kJ/mol
-analytic -7.3164e+001 -3.1240e-002 4.8140e+002 3.3104e+001 7.5336e+000
# -Range: 0-300
KMgCl3
KMgCl3 = + 1.0000 K+ + 1.0000 Mg++ + 3.0000 Cl-
log_k 21.2618
-delta_H -132.768 kJ/mol # Calculated enthalpy of reaction KMgCl3
# Enthalpy of formation: -1086.6 kJ/mol
-analytic -8.4641e+000 -3.2688e-002 5.1496e+003 8.9652e+000 8.7450e+001
# -Range: 0-200
KMgCl3:2H2O
KMgCl3:2H2O = + 1.0000 K+ + 1.0000 Mg++ + 2.0000 H2O + 3.0000 Cl-
log_k 13.9755
-delta_H -76.8449 kJ/mol # Calculated enthalpy of reaction KMgCl3:2H2O
# Enthalpy of formation: -1714.2 kJ/mol
-analytic -5.9982e+001 -3.3015e-002 4.6174e+003 2.7602e+001 7.8431e+001
# -Range: 0-200
KNaCO3:6H2O
KNaCO3:6H2O +1.0000 H+ = + 1.0000 HCO3- + 1.0000 K+ + 1.0000 Na+ + 6.0000 H2O
log_k 10.2593
-delta_H 0 # Not possible to calculate enthalpy of reaction KNaCO3:6H2O
# Enthalpy of formation: 0 kcal/mol
KTcO4
KTcO4 = + 1.0000 K+ + 1.0000 TcO4-
log_k -2.2667
-delta_H 53.2363 kJ/mol # Calculated enthalpy of reaction KTcO4
# Enthalpy of formation: -1021.67 kJ/mol
-analytic 1.8058e+001 -8.4795e-004 -2.3985e+003 -4.1788e+000 -1.5029e+005
# -Range: 0-300
KUO2AsO4
KUO2AsO4 +2.0000 H+ = + 1.0000 H2AsO4- + 1.0000 K+ + 1.0000 UO2++
log_k -4.1741
-delta_H 0 # Not possible to calculate enthalpy of reaction KUO2AsO4
# Enthalpy of formation: 0 kcal/mol
Kainite
KMgClSO4:3H2O = + 1.0000 Cl- + 1.0000 K+ + 1.0000 Mg++ + 1.0000 SO4-- + 3.0000 H2O
log_k -0.3114
-delta_H 0 # Not possible to calculate enthalpy of reaction Kainite
# Enthalpy of formation: 0 kcal/mol
Kalicinite
KHCO3 = + 1.0000 HCO3- + 1.0000 K+
log_k 0.2837
-delta_H 0 # Not possible to calculate enthalpy of reaction Kalicinite
# Enthalpy of formation: 0 kcal/mol
Kalsilite
KAlSiO4 +4.0000 H+ = + 1.0000 Al+++ + 1.0000 K+ + 1.0000 SiO2 + 2.0000 H2O
log_k 10.8987
-delta_H -108.583 kJ/mol # Calculated enthalpy of reaction Kalsilite
# Enthalpy of formation: -509.408 kcal/mol
-analytic -6.7595e+000 -7.4301e-003 6.5380e+003 1.8999e-001 -2.2880e+005
# -Range: 0-300
Kaolinite
Al2Si2O5(OH)4 +6.0000 H+ = + 2.0000 Al+++ + 2.0000 SiO2 + 5.0000 H2O
log_k 6.8101
-delta_H -151.779 kJ/mol # Calculated enthalpy of reaction Kaolinite
# Enthalpy of formation: -982.221 kcal/mol
-analytic 1.6835e+001 -7.8939e-003 7.7636e+003 -1.2190e+001 -3.2354e+005
# -Range: 0-300
Karelianite
V2O3 +6.0000 H+ = + 2.0000 V+++ + 3.0000 H2O
log_k 9.9424
-delta_H -160.615 kJ/mol # Calculated enthalpy of reaction Karelianite
# Enthalpy of formation: -1218.98 kJ/mol
-analytic -2.7961e+001 -7.1499e-003 6.7749e+003 5.8146e+000 2.6039e+005
# -Range: 0-300
Kasolite
Pb(UO2)SiO4:H2O +4.0000 H+ = + 1.0000 Pb++ + 1.0000 SiO2 + 1.0000 UO2++ + 3.0000 H2O
log_k 7.2524
-delta_H 0 # Not possible to calculate enthalpy of reaction Kasolite
# Enthalpy of formation: 0 kcal/mol
Katoite
Ca3Al2H12O12 +12.0000 H+ = + 2.0000 Al+++ + 3.0000 Ca++ + 12.0000 H2O
log_k 78.9437
-delta_H 0 # Not possible to calculate enthalpy of reaction Katoite
# Enthalpy of formation: 0 kcal/mol
Kieserite
MgSO4:H2O = + 1.0000 H2O + 1.0000 Mg++ + 1.0000 SO4--
log_k -0.2670
-delta_H 0 # Not possible to calculate enthalpy of reaction Kieserite
# Enthalpy of formation: 0 kcal/mol
Klockmannite
CuSe = + 1.0000 Cu++ + 1.0000 Se--
log_k -41.6172
-delta_H 0 # Not possible to calculate enthalpy of reaction Klockmannite
# Enthalpy of formation: -10 kcal/mol
-analytic -2.3021e+001 -2.1458e-003 -8.5938e+003 4.3900e+000 -1.4593e+002
# -Range: 0-200
Krutaite
CuSe2 +1.0000 H2O = + 0.5000 O2 + 1.0000 Cu++ + 2.0000 H+ + 2.0000 Se--
log_k -107.6901
-delta_H 0 # Not possible to calculate enthalpy of reaction Krutaite
# Enthalpy of formation: -11.5 kcal/mol
-analytic -3.7735e+001 -8.7548e-004 -2.6352e+004 7.5528e+000 -4.4749e+002
# -Range: 0-200
Kyanite
Al2SiO5 +6.0000 H+ = + 1.0000 SiO2 + 2.0000 Al+++ + 3.0000 H2O
log_k 15.6740
-delta_H -230.919 kJ/mol # Calculated enthalpy of reaction Kyanite
# Enthalpy of formation: -616.897 kcal/mol
-analytic -7.3335e+001 -3.2853e-002 1.2166e+004 2.3412e+001 1.8986e+002
# -Range: 0-300
La
La +3.0000 H+ +0.7500 O2 = + 1.0000 La+++ + 1.5000 H2O
log_k 184.7155
-delta_H -1129.26 kJ/mol # Calculated enthalpy of reaction La
# Enthalpy of formation: 0 kJ/mol
-analytic -5.9508e+001 -2.7578e-002 5.9327e+004 2.1589e+001 9.2577e+002
# -Range: 0-300
La(OH)3(s)
La(OH)3 +3.0000 H+ = + 1.0000 La+++ + 3.0000 H2O
log_k 20.2852
-delta_H 0 # Not possible to calculate enthalpy of reaction La(OH)3
# Enthalpy of formation: 0 kcal/mol
La(OH)3(am)
La(OH)3 +3.0000 H+ = + 1.0000 La+++ + 3.0000 H2O
log_k 23.4852
-delta_H 0 # Not possible to calculate enthalpy of reaction La(OH)3(am)
# Enthalpy of formation: 0 kcal/mol
La2(CO3)3:8H2O
La2(CO3)3:8H2O +3.0000 H+ = + 2.0000 La+++ + 3.0000 HCO3- + 8.0000 H2O
log_k -4.3136
-delta_H 0 # Not possible to calculate enthalpy of reaction La2(CO3)3:8H2O
# Enthalpy of formation: 0 kcal/mol
La2O3(s)
La2O3 +6.0000 H+ = + 2.0000 La+++ + 3.0000 H2O
log_k 66.2000
-delta_H 0 # Not possible to calculate enthalpy of reaction La2O3
# Enthalpy of formation: 0 kcal/mol
LaCl3
LaCl3 = + 1.0000 La+++ + 3.0000 Cl-
log_k 14.4000
-delta_H 0 # Not possible to calculate enthalpy of reaction LaCl3
# Enthalpy of formation: 0 kcal/mol
LaCl3:7H2O
LaCl3:7H2O = + 1.0000 La+++ + 3.0000 Cl- + 7.0000 H2O
log_k 4.7000
-delta_H 0 # Not possible to calculate enthalpy of reaction LaCl3:7H2O
# Enthalpy of formation: 0 kcal/mol
LaF3:.5H2O
LaF3:.5H2O = + 0.5000 H2O + 1.0000 La+++ + 3.0000 F-
log_k -18.7000
-delta_H 0 # Not possible to calculate enthalpy of reaction LaF3:.5H2O
# Enthalpy of formation: 0 kcal/mol
LaPO4(s)
LaPO4:10H2O +1.0000 H+ = + 1.0000 HPO4-- + 1.0000 La+++ + 10.0000 H2O
log_k -12.3782
-delta_H 0 # Not possible to calculate enthalpy of reaction LaPO4:10H2O
# Enthalpy of formation: 0 kcal/mol
Lammerite
Cu3(AsO4)2 +4.0000 H+ = + 2.0000 H2AsO4- + 3.0000 Cu++
log_k 1.5542
-delta_H 0 # Not possible to calculate enthalpy of reaction Lammerite
# Enthalpy of formation: 0 kcal/mol
Lanarkite
Pb2(SO4)O +2.0000 H+ = + 1.0000 H2O + 1.0000 SO4-- + 2.0000 Pb++
log_k -0.4692
-delta_H -22.014 kJ/mol # Calculated enthalpy of reaction Lanarkite
# Enthalpy of formation: -1171.59 kJ/mol
-analytic 5.1071e+000 -1.6655e-002 0.0000e+000 0.0000e+000 -5.5660e+004
# -Range: 0-200
Lansfordite
MgCO3:5H2O +1.0000 H+ = + 1.0000 HCO3- + 1.0000 Mg++ + 5.0000 H2O
log_k 4.8409
-delta_H 0 # Not possible to calculate enthalpy of reaction Lansfordite
# Enthalpy of formation: 0 kcal/mol
Larnite
Ca2SiO4 +4.0000 H+ = + 1.0000 SiO2 + 2.0000 Ca++ + 2.0000 H2O
log_k 38.4665
-delta_H -227.061 kJ/mol # Calculated enthalpy of reaction Larnite
# Enthalpy of formation: -551.74 kcal/mol
-analytic 2.6900e+001 -2.1833e-003 1.0900e+004 -9.5257e+000 -7.2537e+004
# -Range: 0-300
Laumontite
CaAl2Si4O12:4H2O +8.0000 H+ = + 1.0000 Ca++ + 2.0000 Al+++ + 4.0000 SiO2 + 8.0000 H2O
log_k 13.6667
-delta_H -184.657 kJ/mol # Calculated enthalpy of reaction Laumontite
# Enthalpy of formation: -1728.66 kcal/mol
-analytic 1.1904e+000 8.1763e-003 1.9005e+004 -1.4561e+001 -1.5851e+006
# -Range: 0-300
Laurite
RuS2 = + 1.0000 Ru++ + 1.0000 S2--
log_k -73.2649
-delta_H 0 # Not possible to calculate enthalpy of reaction Laurite
# Enthalpy of formation: -199.586 kJ/mol
Lawrencite
FeCl2 = + 1.0000 Fe++ + 2.0000 Cl-
log_k 9.0945
-delta_H -84.7665 kJ/mol # Calculated enthalpy of reaction Lawrencite
# Enthalpy of formation: -341.65 kJ/mol
-analytic -2.2798e+002 -8.1819e-002 9.2620e+003 9.3097e+001 1.4459e+002
# -Range: 0-300
Lawsonite
CaAl2Si2O7(OH)2:H2O +8.0000 H+ = + 1.0000 Ca++ + 2.0000 Al+++ + 2.0000 SiO2 + 6.0000 H2O
log_k 22.2132
-delta_H -244.806 kJ/mol # Calculated enthalpy of reaction Lawsonite
# Enthalpy of formation: -1158.1 kcal/mol
-analytic 1.3995e+001 -1.7668e-002 1.0119e+004 -8.3100e+000 1.5789e+002
# -Range: 0-300
Leonite
K2Mg(SO4)2:4H2O = + 1.0000 Mg++ + 2.0000 K+ + 2.0000 SO4-- + 4.0000 H2O
log_k -4.1123
-delta_H 0 # Not possible to calculate enthalpy of reaction Leonite
# Enthalpy of formation: 0 kcal/mol
Li
Li +1.0000 H+ +0.2500 O2 = + 0.5000 H2O + 1.0000 Li+
log_k 72.7622
-delta_H -418.339 kJ/mol # Calculated enthalpy of reaction Li
# Enthalpy of formation: 0 kJ/mol
-analytic -1.0227e+002 -1.8118e-002 2.6262e+004 3.8056e+001 -1.6166e+005
# -Range: 0-300
Li2Se
Li2Se +1.5000 O2 = + 1.0000 SeO3-- + 2.0000 Li+
log_k 102.8341
-delta_H -646.236 kJ/mol # Calculated enthalpy of reaction Li2Se
# Enthalpy of formation: -96 kcal/mol
-analytic 1.1933e+002 -6.9663e-003 2.7509e+004 -4.3124e+001 4.6710e+002
# -Range: 0-200
Li2UO4
Li2UO4 +4.0000 H+ = + 1.0000 UO2++ + 2.0000 H2O + 2.0000 Li+
log_k 27.8421
-delta_H -179.384 kJ/mol # Calculated enthalpy of reaction Li2UO4
# Enthalpy of formation: -1968.2 kJ/mol
-analytic -1.4470e+002 -1.2024e-002 1.4899e+004 5.0984e+001 2.5306e+002
# -Range: 0-200
LiUO2AsO4
LiUO2AsO4 +2.0000 H+ = + 1.0000 H2AsO4- + 1.0000 Li+ + 1.0000 UO2++
log_k -0.7862
-delta_H 0 # Not possible to calculate enthalpy of reaction LiUO2AsO4
# Enthalpy of formation: 0 kcal/mol
Lime
CaO +2.0000 H+ = + 1.0000 Ca++ + 1.0000 H2O
log_k 32.5761
-delta_H -193.832 kJ/mol # Calculated enthalpy of reaction Lime
# Enthalpy of formation: -151.79 kcal/mol
-analytic -7.2686e+001 -1.7654e-002 1.2199e+004 2.8128e+001 1.9037e+002
# -Range: 0-300
Linnaeite
Co3S4 +4.0000 H+ = + 1.0000 Co++ + 2.0000 Co+++ + 4.0000 HS-
log_k -106.9017
-delta_H 420.534 kJ/mol # Calculated enthalpy of reaction Linnaeite
# Enthalpy of formation: -85.81 kcal/mol
-analytic -6.0034e+002 -2.0179e-001 -9.2145e+003 2.3618e+002 -1.4361e+002
# -Range: 0-300
Litharge
PbO +2.0000 H+ = + 1.0000 H2O + 1.0000 Pb++
log_k 12.6388
-delta_H -65.9118 kJ/mol # Calculated enthalpy of reaction Litharge
# Enthalpy of formation: -219.006 kJ/mol
-analytic -1.8683e+001 -2.0211e-003 4.1876e+003 7.2239e+000 7.1118e+001
# -Range: 0-200
Lopezite
K2Cr2O7 +1.0000 H2O = + 2.0000 CrO4-- + 2.0000 H+ + 2.0000 K+
log_k -17.4366
-delta_H 81.9227 kJ/mol # Calculated enthalpy of reaction Lopezite
# Enthalpy of formation: -493.003 kcal/mol
-analytic 7.8359e+001 -2.2908e-002 -9.3812e+003 -2.3245e+001 -1.5933e+002
# -Range: 0-200
Lu
Lu +3.0000 H+ +0.7500 O2 = + 1.0000 Lu+++ + 1.5000 H2O
log_k 181.3437
-delta_H -1122.15 kJ/mol # Calculated enthalpy of reaction Lu
# Enthalpy of formation: 0 kJ/mol
-analytic -6.8950e+001 -2.8643e-002 5.9209e+004 2.4332e+001 9.2392e+002
# -Range: 0-300
Lu(OH)3
Lu(OH)3 +3.0000 H+ = + 1.0000 Lu+++ + 3.0000 H2O
log_k 14.4852
-delta_H 0 # Not possible to calculate enthalpy of reaction Lu(OH)3
# Enthalpy of formation: 0 kcal/mol
Lu(OH)3(am)
Lu(OH)3 +3.0000 H+ = + 1.0000 Lu+++ + 3.0000 H2O
log_k 18.9852
-delta_H 0 # Not possible to calculate enthalpy of reaction Lu(OH)3(am)
# Enthalpy of formation: 0 kcal/mol
Lu2(CO3)3
Lu2(CO3)3 +3.0000 H+ = + 2.0000 Lu+++ + 3.0000 HCO3-
log_k -2.0136
-delta_H 0 # Not possible to calculate enthalpy of reaction Lu2(CO3)3
# Enthalpy of formation: 0 kcal/mol
Lu2O3
Lu2O3 +6.0000 H+ = + 2.0000 Lu+++ + 3.0000 H2O
log_k 45.0000
-delta_H 0 # Not possible to calculate enthalpy of reaction Lu2O3
# Enthalpy of formation: 0 kcal/mol
LuF3:.5H2O
LuF3:.5H2O = + 0.5000 H2O + 1.0000 Lu+++ + 3.0000 F-
log_k -15.9000
-delta_H 0 # Not possible to calculate enthalpy of reaction LuF3:.5H2O
# Enthalpy of formation: 0 kcal/mol
LuPO4:10H2O
LuPO4:10H2O +1.0000 H+ = + 1.0000 HPO4-- + 1.0000 Lu+++ + 10.0000 H2O
log_k -11.6782
-delta_H 0 # Not possible to calculate enthalpy of reaction LuPO4:10H2O
# Enthalpy of formation: 0 kcal/mol
Magnesiochromite
MgCr2O4 +8.0000 H+ = + 1.0000 Mg++ + 2.0000 Cr+++ + 4.0000 H2O
log_k 21.6927
-delta_H -302.689 kJ/mol # Calculated enthalpy of reaction Magnesiochromite
# Enthalpy of formation: -1783.6 kJ/mol
-analytic -1.7376e+002 -8.7429e-003 2.1600e+004 5.0762e+001 3.6685e+002
# -Range: 0-200
Magnesite
MgCO3 +1.0000 H+ = + 1.0000 HCO3- + 1.0000 Mg++
log_k 2.2936
-delta_H -44.4968 kJ/mol # Calculated enthalpy of reaction Magnesite
# Enthalpy of formation: -265.63 kcal/mol
-analytic -1.6665e+002 -4.9469e-002 6.4344e+003 6.5506e+001 1.0045e+002
# -Range: 0-300
Magnetite
Fe3O4 +8.0000 H+ = + 1.0000 Fe++ + 2.0000 Fe+++ + 4.0000 H2O
log_k 10.4724
-delta_H -216.597 kJ/mol # Calculated enthalpy of reaction Magnetite
# Enthalpy of formation: -267.25 kcal/mol
-analytic -3.0510e+002 -7.9919e-002 1.8709e+004 1.1178e+002 2.9203e+002
# -Range: 0-300
Malachite
Cu2CO3(OH)2 +3.0000 H+ = + 1.0000 HCO3- + 2.0000 Cu++ + 2.0000 H2O
log_k 5.9399
-delta_H -76.2827 kJ/mol # Calculated enthalpy of reaction Malachite
# Enthalpy of formation: -251.9 kcal/mol
-analytic -2.7189e+002 -6.9454e-002 1.1451e+004 1.0511e+002 1.7877e+002
# -Range: 0-300
Manganite
MnO(OH) +3.0000 H+ = + 1.0000 Mn+++ + 2.0000 H2O
log_k -0.1646
-delta_H 0 # Not possible to calculate enthalpy of reaction Manganite
# Enthalpy of formation: 0 kcal/mol
Manganosite
MnO +2.0000 H+ = + 1.0000 H2O + 1.0000 Mn++
log_k 17.9240
-delta_H -121.215 kJ/mol # Calculated enthalpy of reaction Manganosite
# Enthalpy of formation: -92.07 kcal/mol
-analytic -8.4114e+001 -1.8490e-002 8.7792e+003 3.1561e+001 1.3702e+002
# -Range: 0-300
Margarite
CaAl4Si2O10(OH)2 +14.0000 H+ = + 1.0000 Ca++ + 2.0000 SiO2 + 4.0000 Al+++ + 8.0000 H2O
log_k 41.0658
-delta_H -522.192 kJ/mol # Calculated enthalpy of reaction Margarite
# Enthalpy of formation: -1485.8 kcal/mol
-analytic -2.3138e+002 -8.2788e-002 3.0154e+004 7.9148e+001 4.7060e+002
# -Range: 0-300
Massicot
PbO +2.0000 H+ = + 1.0000 H2O + 1.0000 Pb++
log_k 12.8210
-delta_H -67.6078 kJ/mol # Calculated enthalpy of reaction Massicot
# Enthalpy of formation: -217.31 kJ/mol
-analytic -1.8738e+001 -2.0125e-003 4.2739e+003 7.2018e+000 7.2584e+001
# -Range: 0-200
Matlockite
PbFCl = + 1.0000 Cl- + 1.0000 F- + 1.0000 Pb++
log_k -9.4300
-delta_H 0 # Not possible to calculate enthalpy of reaction Matlockite
# Enthalpy of formation: 0 kcal/mol
Maximum_Microcline
KAlSi3O8 +4.0000 H+ = + 1.0000 Al+++ + 1.0000 K+ + 2.0000 H2O + 3.0000 SiO2
log_k -0.2753
-delta_H -23.9408 kJ/mol # Calculated enthalpy of reaction Maximum_Microcline
# Enthalpy of formation: -949.188 kcal/mol
-analytic -9.4387e+000 1.3561e-002 1.2656e+004 -7.4925e+000 -1.6795e+006
# -Range: 0-300
Mayenite
Ca12Al14O33 +66.0000 H+ = + 12.0000 Ca++ + 14.0000 Al+++ + 33.0000 H2O
log_k 494.2199
-delta_H -4056.77 kJ/mol # Calculated enthalpy of reaction Mayenite
# Enthalpy of formation: -4644 kcal/mol
-analytic -1.4778e+003 -2.9898e-001 2.4918e+005 4.9518e+002 4.2319e+003
# -Range: 0-200
Melanterite
FeSO4:7H2O = + 1.0000 Fe++ + 1.0000 SO4-- + 7.0000 H2O
log_k -2.3490
-delta_H 11.7509 kJ/mol # Calculated enthalpy of reaction Melanterite
# Enthalpy of formation: -3014.48 kJ/mol
-analytic -2.6230e+002 -7.2469e-002 6.5854e+003 1.0484e+002 1.0284e+002
# -Range: 0-300
Mercallite
KHSO4 = + 1.0000 H+ + 1.0000 K+ + 1.0000 SO4--
log_k -1.4389
-delta_H 0 # Not possible to calculate enthalpy of reaction Mercallite
# Enthalpy of formation: 0 kcal/mol
Merwinite
MgCa3(SiO4)2 +8.0000 H+ = + 1.0000 Mg++ + 2.0000 SiO2 + 3.0000 Ca++ + 4.0000 H2O
log_k 68.5140
-delta_H -430.069 kJ/mol # Calculated enthalpy of reaction Merwinite
# Enthalpy of formation: -1090.8 kcal/mol
-analytic -2.2524e+002 -4.2525e-002 3.5619e+004 7.9984e+001 -9.8259e+005
# -Range: 0-300
Mesolite
Na.676Ca.657Al1.99Si3.01O10:2.647H2O +7.9600 H+ = + 0.6570 Ca++ + 0.6760 Na+ + 1.9900 Al+++ + 3.0100 SiO2 + 6.6270 H2O
log_k 13.6191
-delta_H -179.744 kJ/mol # Calculated enthalpy of reaction Mesolite
# Enthalpy of formation: -5947.05 kJ/mol
-analytic 7.1993e+000 5.9356e-003 1.4717e+004 -1.3627e+001 -9.8863e+005
# -Range: 0-300
Metacinnabar
HgS +1.0000 H+ = + 1.0000 HS- + 1.0000 Hg++
log_k -38.5979
-delta_H 203.426 kJ/mol # Calculated enthalpy of reaction Metacinnabar
# Enthalpy of formation: -11.8 kcal/mol
-analytic -1.5399e+002 -4.6740e-002 -6.7875e+003 6.1456e+001 -1.0587e+002
# -Range: 0-300
Mg
Mg +2.0000 H+ +0.5000 O2 = + 1.0000 H2O + 1.0000 Mg++
log_k 122.5365
-delta_H -745.731 kJ/mol # Calculated enthalpy of reaction Mg
# Enthalpy of formation: 0 kJ/mol
-analytic -6.5988e+001 -1.9356e-002 4.0318e+004 2.3862e+001 6.2914e+002
# -Range: 0-300
Mg1.25SO4(OH)0.5:0.5H2O
Mg1.25SO4(OH)0.5:0.5H2O +0.5000 H+ = + 1.0000 H2O + 1.0000 SO4-- + 1.2500 Mg++
log_k 5.2600
-delta_H -97.1054 kJ/mol # Calculated enthalpy of reaction Mg1.25SO4(OH)0.5:0.5H2O
# Enthalpy of formation: -401.717 kcal/mol
-analytic -2.6791e+002 -8.7078e-002 1.1090e+004 1.0583e+002 1.7312e+002
# -Range: 0-300
Mg1.5SO4(OH)
Mg1.5SO4(OH) +1.0000 H+ = + 1.0000 H2O + 1.0000 SO4-- + 1.5000 Mg++
log_k 9.2551
-delta_H -125.832 kJ/mol # Calculated enthalpy of reaction Mg1.5SO4(OH)
# Enthalpy of formation: -422.693 kcal/mol
-analytic -2.8698e+002 -9.1970e-002 1.3088e+004 1.1304e+002 2.0432e+002
# -Range: 0-300
Mg2V2O7
Mg2V2O7 +1.0000 H2O = + 2.0000 H+ + 2.0000 Mg++ + 2.0000 VO4---
log_k -30.9025
-delta_H 0 # Not possible to calculate enthalpy of reaction Mg2V2O7
# Enthalpy of formation: -2836.23 kJ/mol
MgBr2
MgBr2 = + 1.0000 Mg++ + 2.0000 Br-
log_k 28.5302
-delta_H -190.15 kJ/mol # Calculated enthalpy of reaction MgBr2
# Enthalpy of formation: -124 kcal/mol
-analytic -2.1245e+002 -7.6168e-002 1.4466e+004 8.6940e+001 2.2579e+002
# -Range: 0-300
MgBr2:6H2O
MgBr2:6H2O = + 1.0000 Mg++ + 2.0000 Br- + 6.0000 H2O
log_k 5.1656
-delta_H -14.2682 kJ/mol # Calculated enthalpy of reaction MgBr2:6H2O
# Enthalpy of formation: -2409.73 kJ/mol
-analytic -1.3559e+002 -1.6479e-002 5.8571e+003 5.0924e+001 9.9508e+001
# -Range: 0-200
MgCl2:2H2O
MgCl2:2H2O = + 1.0000 Mg++ + 2.0000 Cl- + 2.0000 H2O
log_k 12.7763
-delta_H -92.0895 kJ/mol # Calculated enthalpy of reaction MgCl2:2H2O
# Enthalpy of formation: -1279.71 kJ/mol
-analytic -2.5409e+002 -8.1413e-002 1.0941e+004 1.0281e+002 1.7080e+002
# -Range: 0-300
MgCl2:4H2O
MgCl2:4H2O = + 1.0000 Mg++ + 2.0000 Cl- + 4.0000 H2O
log_k 7.3581
-delta_H -44.4602 kJ/mol # Calculated enthalpy of reaction MgCl2:4H2O
# Enthalpy of formation: -1899.01 kJ/mol
-analytic -2.7604e+002 -8.1648e-002 9.5501e+003 1.1140e+002 1.4910e+002
# -Range: 0-300
MgCl2:H2O
MgCl2:H2O = + 1.0000 H2O + 1.0000 Mg++ + 2.0000 Cl-
log_k 16.1187
-delta_H -119.326 kJ/mol # Calculated enthalpy of reaction MgCl2:H2O
# Enthalpy of formation: -966.631 kJ/mol
-analytic -2.4414e+002 -8.1310e-002 1.1862e+004 9.8878e+001 1.8516e+002
# -Range: 0-300
MgOHCl
MgOHCl +1.0000 H+ = + 1.0000 Cl- + 1.0000 H2O + 1.0000 Mg++
log_k 15.9138
-delta_H -118.897 kJ/mol # Calculated enthalpy of reaction MgOHCl
# Enthalpy of formation: -191.2 kcal/mol
-analytic -1.6614e+002 -4.9715e-002 1.0311e+004 6.5578e+001 1.6093e+002
# -Range: 0-300
MgSO4
MgSO4 = + 1.0000 Mg++ + 1.0000 SO4--
log_k 4.8781
-delta_H -90.6421 kJ/mol # Calculated enthalpy of reaction MgSO4
# Enthalpy of formation: -1284.92 kJ/mol
-analytic -2.2439e+002 -7.9688e-002 9.3058e+003 8.9622e+001 1.4527e+002
# -Range: 0-300
MgSeO3
MgSeO3 = + 1.0000 Mg++ + 1.0000 SeO3--
log_k 1.7191
-delta_H -74.9647 kJ/mol # Calculated enthalpy of reaction MgSeO3
# Enthalpy of formation: -215.15 kcal/mol
-analytic -2.2593e+002 -8.1045e-002 8.4609e+003 9.0278e+001 1.3209e+002
# -Range: 0-300
MgSeO3:6H2O
MgSeO3:6H2O = + 1.0000 Mg++ + 1.0000 SeO3-- + 6.0000 H2O
log_k -3.4222
-delta_H 11.7236 kJ/mol # Calculated enthalpy of reaction MgSeO3:6H2O
# Enthalpy of formation: -645.771 kcal/mol
-analytic -1.2807e+002 -1.5418e-002 4.0565e+003 4.6728e+001 6.8929e+001
# -Range: 0-200
MgUO4
MgUO4 +4.0000 H+ = + 1.0000 Mg++ + 1.0000 UO2++ + 2.0000 H2O
log_k 23.0023
-delta_H -199.336 kJ/mol # Calculated enthalpy of reaction MgUO4
# Enthalpy of formation: -1857.3 kJ/mol
-analytic -9.9954e+001 -2.0142e-002 1.3078e+004 3.4386e+001 2.0410e+002
# -Range: 0-300
MgV2O6
MgV2O6 +2.0000 H2O = + 1.0000 Mg++ + 2.0000 VO4--- + 4.0000 H+
log_k -45.8458
-delta_H 0 # Not possible to calculate enthalpy of reaction MgV2O6
# Enthalpy of formation: -2201.88 kJ/mol
Millerite
NiS +1.0000 H+ = + 1.0000 HS- + 1.0000 Ni++
log_k -8.0345
-delta_H 12.089 kJ/mol # Calculated enthalpy of reaction Millerite
# Enthalpy of formation: -82.171 kJ/mol
-analytic -1.4848e+002 -4.8834e-002 2.6981e+003 5.8976e+001 4.2145e+001
# -Range: 0-300
Minium
Pb3O4 +8.0000 H+ = + 1.0000 Pb++++ + 2.0000 Pb++ + 4.0000 H2O
log_k 16.2585
-delta_H 0 # Not possible to calculate enthalpy of reaction Minium
# Enthalpy of formation: -718.493 kJ/mol
Minnesotaite
Fe3Si4O10(OH)2 +6.0000 H+ = + 3.0000 Fe++ + 4.0000 H2O + 4.0000 SiO2
log_k 13.9805
-delta_H -105.211 kJ/mol # Calculated enthalpy of reaction Minnesotaite
# Enthalpy of formation: -1153.37 kcal/mol
-analytic -1.8812e+001 1.7261e-002 1.9804e+004 -6.4410e+000 -2.0433e+006
# -Range: 0-300
Mirabilite
Na2SO4:10H2O = + 1.0000 SO4-- + 2.0000 Na+ + 10.0000 H2O
log_k -1.1398
-delta_H 79.4128 kJ/mol # Calculated enthalpy of reaction Mirabilite
# Enthalpy of formation: -4328 kJ/mol
-analytic -2.1877e+002 -3.6692e-003 5.9214e+003 8.0361e+001 1.0063e+002
# -Range: 0-200
Misenite
K8H6(SO4)7 = + 6.0000 H+ + 7.0000 SO4-- + 8.0000 K+
log_k -11.0757
-delta_H 0 # Not possible to calculate enthalpy of reaction Misenite
# Enthalpy of formation: 0 kcal/mol
Mn
Mn +2.0000 H+ +0.5000 O2 = + 1.0000 H2O + 1.0000 Mn++
log_k 82.9505
-delta_H -500.369 kJ/mol # Calculated enthalpy of reaction Mn
# Enthalpy of formation: 0 kJ/mol
-analytic -6.5558e+001 -2.0429e-002 2.7571e+004 2.5098e+001 4.3024e+002
# -Range: 0-300
Mn(OH)2(am)
Mn(OH)2 +2.0000 H+ = + 1.0000 Mn++ + 2.0000 H2O
log_k 15.3102
-delta_H -97.1779 kJ/mol # Calculated enthalpy of reaction Mn(OH)2(am)
# Enthalpy of formation: -695.096 kJ/mol
-analytic -7.8518e+001 -7.5357e-003 8.0198e+003 2.7955e+001 1.3621e+002
# -Range: 0-200
Mn(OH)3
Mn(OH)3 +3.0000 H+ = + 1.0000 Mn+++ + 3.0000 H2O
log_k 6.3412
-delta_H 0 # Not possible to calculate enthalpy of reaction Mn(OH)3
# Enthalpy of formation: 0 kcal/mol
Mn3(PO4)2
Mn3(PO4)2 +2.0000 H+ = + 2.0000 HPO4-- + 3.0000 Mn++
log_k 0.8167
-delta_H 0 # Not possible to calculate enthalpy of reaction Mn3(PO4)2
# Enthalpy of formation: 0 kcal/mol
MnCl2:2H2O
MnCl2:2H2O = + 1.0000 Mn++ + 2.0000 Cl- + 2.0000 H2O
log_k 4.0067
-delta_H -34.4222 kJ/mol # Calculated enthalpy of reaction MnCl2:2H2O
# Enthalpy of formation: -1092.01 kJ/mol
-analytic -6.2823e+001 -2.3959e-002 2.9931e+003 2.5834e+001 5.0850e+001
# -Range: 0-200
MnCl2:4H2O
MnCl2:4H2O = + 1.0000 Mn++ + 2.0000 Cl- + 4.0000 H2O
log_k 2.7563
-delta_H -10.7019 kJ/mol # Calculated enthalpy of reaction MnCl2:4H2O
# Enthalpy of formation: -1687.41 kJ/mol
-analytic -1.1049e+002 -2.3376e-002 4.0458e+003 4.3097e+001 6.8742e+001
# -Range: 0-200
MnCl2:H2O
MnCl2:H2O = + 1.0000 H2O + 1.0000 Mn++ + 2.0000 Cl-
log_k 5.5517
-delta_H -50.8019 kJ/mol # Calculated enthalpy of reaction MnCl2:H2O
# Enthalpy of formation: -789.793 kJ/mol
-analytic -4.5051e+001 -2.5923e-002 2.8739e+003 1.9674e+001 4.8818e+001
# -Range: 0-200
MnHPO4
MnHPO4 = + 1.0000 HPO4-- + 1.0000 Mn++
log_k -12.9470
-delta_H 0 # Not possible to calculate enthalpy of reaction MnHPO4
# Enthalpy of formation: 0 kcal/mol
MnO2(gamma)
MnO2 = + 0.5000 Mn++ + 0.5000 MnO4--
log_k -16.1261
-delta_H 0 # Not possible to calculate enthalpy of reaction MnO2(gamma)
# Enthalpy of formation: 0 kcal/mol
MnSO4
MnSO4 = + 1.0000 Mn++ + 1.0000 SO4--
log_k 2.6561
-delta_H -64.8718 kJ/mol # Calculated enthalpy of reaction MnSO4
# Enthalpy of formation: -1065.33 kJ/mol
-analytic -2.3088e+002 -8.2694e-002 8.1653e+003 9.3256e+001 1.2748e+002
# -Range: 0-300
MnSe
MnSe = + 1.0000 Mn++ + 1.0000 Se--
log_k -10.6848
-delta_H 0 # Not possible to calculate enthalpy of reaction MnSe
# Enthalpy of formation: -37 kcal/mol
-analytic -5.9960e+001 -1.5963e-002 1.2813e+003 2.0095e+001 2.0010e+001
# -Range: 0-300
MnSeO3
MnSeO3 = + 1.0000 Mn++ + 1.0000 SeO3--
log_k -7.2700
-delta_H 0 # Not possible to calculate enthalpy of reaction MnSeO3
# Enthalpy of formation: 0 kcal/mol
MnSeO3:2H2O
MnSeO3:2H2O = + 1.0000 Mn++ + 1.0000 SeO3-- + 2.0000 H2O
log_k -6.3219
-delta_H 14.0792 kJ/mol # Calculated enthalpy of reaction MnSeO3:2H2O
# Enthalpy of formation: -314.423 kcal/mol
-analytic -4.3625e+001 -2.0426e-002 -2.5368e+002 1.7876e+001 -4.2927e+000
# -Range: 0-200
MnV2O6
MnV2O6 +2.0000 H2O = + 1.0000 Mn++ + 2.0000 VO4--- + 4.0000 H+
log_k -52.0751
-delta_H 0 # Not possible to calculate enthalpy of reaction MnV2O6
# Enthalpy of formation: -447.9 kcal/mol
Mo
Mo +1.5000 O2 +1.0000 H2O = + 1.0000 MoO4-- + 2.0000 H+
log_k 109.3230
-delta_H -693.845 kJ/mol # Calculated enthalpy of reaction Mo
# Enthalpy of formation: 0 kJ/mol
-analytic -2.0021e+002 -8.3006e-002 4.1629e+004 8.0219e+001 -3.4570e+005
# -Range: 0-300
MoSe2
MoSe2 +3.0000 H2O +0.5000 O2 = + 1.0000 MoO4-- + 2.0000 Se-- + 6.0000 H+
log_k -55.1079
-delta_H 0 # Not possible to calculate enthalpy of reaction MoSe2
# Enthalpy of formation: -47 kcal/mol
-analytic 1.3882e+002 -1.8590e-003 -1.7231e+004 -5.4797e+001 -2.9265e+002
# -Range: 0-200
Modderite
CoAs +3.0000 H+ = + 1.0000 AsH3 + 1.0000 Co+++
log_k -49.5512
-delta_H 189.016 kJ/mol # Calculated enthalpy of reaction Modderite
# Enthalpy of formation: -12.208 kcal/mol
Molysite
FeCl3 = + 1.0000 Fe+++ + 3.0000 Cl-
log_k 13.5517
-delta_H -151.579 kJ/mol # Calculated enthalpy of reaction Molysite
# Enthalpy of formation: -399.24 kJ/mol
-analytic -3.1810e+002 -1.2357e-001 1.3860e+004 1.3010e+002 2.1637e+002
# -Range: 0-300
Monohydrocalcite
CaCO3:H2O +1.0000 H+ = + 1.0000 Ca++ + 1.0000 H2O + 1.0000 HCO3-
log_k 2.6824
-delta_H -20.5648 kJ/mol # Calculated enthalpy of reaction Monohydrocalcite
# Enthalpy of formation: -1498.29 kJ/mol
-analytic -7.2614e+001 -1.7217e-002 3.1850e+003 2.8185e+001 5.4111e+001
# -Range: 0-200
Monteponite
CdO +2.0000 H+ = + 1.0000 Cd++ + 1.0000 H2O
log_k 15.0972
-delta_H -103.386 kJ/mol # Calculated enthalpy of reaction Monteponite
# Enthalpy of formation: -258.35 kJ/mol
-analytic -5.0057e+001 -6.3629e-003 7.0898e+003 1.7486e+001 1.2041e+002
# -Range: 0-200
Monticellite
CaMgSiO4 +4.0000 H+ = + 1.0000 Ca++ + 1.0000 Mg++ + 1.0000 SiO2 + 2.0000 H2O
log_k 29.5852
-delta_H -195.711 kJ/mol # Calculated enthalpy of reaction Monticellite
# Enthalpy of formation: -540.8 kcal/mol
-analytic 1.5730e+001 -3.5567e-003 9.0789e+003 -6.3007e+000 1.4166e+002
# -Range: 0-300
Montmor-Ca
Ca.165Mg.33Al1.67Si4O10(OH)2 +6.0000 H+ = + 0.1650 Ca++ + 0.3300 Mg++ + 1.6700 Al+++ + 4.0000 H2O + 4.0000 SiO2
log_k 2.4952
-delta_H -100.154 kJ/mol # Calculated enthalpy of reaction Montmor-Ca
# Enthalpy of formation: -1361.5 kcal/mol
-analytic 6.0725e+000 1.0644e-002 1.6024e+004 -1.6334e+001 -1.7982e+006
# -Range: 0-300
Montmor-Cs
Cs.33Mg.33Al1.67Si4O10(OH)2 +6.0000 H+ = + 0.3300 Cs+ + 0.3300 Mg++ + 1.6700 Al+++ + 4.0000 H2O + 4.0000 SiO2
log_k 1.9913
-delta_H -87.2259 kJ/mol # Calculated enthalpy of reaction Montmor-Cs
# Enthalpy of formation: -1363.52 kcal/mol
-analytic 9.9136e+000 1.2496e-002 1.5650e+004 -1.7601e+001 -1.8434e+006
# -Range: 0-300
Montmor-K
K.33Mg.33Al1.67Si4O10(OH)2 +6.0000 H+ = + 0.3300 K+ + 0.3300 Mg++ + 1.6700 Al+++ + 4.0000 H2O + 4.0000 SiO2
log_k 2.1423
-delta_H -88.184 kJ/mol # Calculated enthalpy of reaction Montmor-K
# Enthalpy of formation: -1362.83 kcal/mol
-analytic 8.4757e+000 1.1219e-002 1.5654e+004 -1.6833e+001 -1.8386e+006
# -Range: 0-300
Montmor-Mg
Mg.495Al1.67Si4O10(OH)2 +6.0000 H+ = + 0.4950 Mg++ + 1.6700 Al+++ + 4.0000 H2O + 4.0000 SiO2
log_k 2.3879
-delta_H -102.608 kJ/mol # Calculated enthalpy of reaction Montmor-Mg
# Enthalpy of formation: -1357.87 kcal/mol
-analytic -6.8505e+000 9.0710e-003 1.6817e+004 -1.1887e+001 -1.8323e+006
# -Range: 0-300
Montmor-Na
Na.33Mg.33Al1.67Si4O10(OH)2 +6.0000 H+ = + 0.3300 Mg++ + 0.3300 Na+ + 1.6700 Al+++ + 4.0000 H2O + 4.0000 SiO2
log_k 2.4844
-delta_H -93.2165 kJ/mol # Calculated enthalpy of reaction Montmor-Na
# Enthalpy of formation: -1360.69 kcal/mol
-analytic 1.9601e+000 1.1342e-002 1.6051e+004 -1.4718e+001 -1.8160e+006
# -Range: 0-300
Montroydite
HgO +2.0000 H+ = + 1.0000 H2O + 1.0000 Hg++
log_k 2.4486
-delta_H -24.885 kJ/mol # Calculated enthalpy of reaction Montroydite
# Enthalpy of formation: -90.79 kJ/mol
-analytic -8.7302e+001 -1.7618e-002 4.0086e+003 3.2957e+001 6.2576e+001
# -Range: 0-300
Mordenite
Ca.2895Na.361Al.94Si5.06O12:3.468H2O +3.7600 H+ = + 0.2895 Ca++ + 0.3610 Na+ + 0.9400 Al+++ + 5.0600 SiO2 + 5.3480 H2O
log_k -5.1969
-delta_H 16.7517 kJ/mol # Calculated enthalpy of reaction Mordenite
# Enthalpy of formation: -6736.64 kJ/mol
-analytic -5.4675e+001 3.2513e-002 2.3412e+004 -1.0419e+000 -3.2292e+006
# -Range: 0-300
Mordenite-dehy
Ca.2895Na.361Al.94Si5.06O12 +3.7600 H+ = + 0.2895 Ca++ + 0.3610 Na+ + 0.9400 Al+++ + 1.8800 H2O + 5.0600 SiO2
log_k 9.9318
-delta_H -86.159 kJ/mol # Calculated enthalpy of reaction Mordenite-dehy
# Enthalpy of formation: -5642.44 kJ/mol
-analytic -5.0841e+001 2.5405e-002 2.7621e+004 -1.6331e+000 -3.1618e+006
# -Range: 0-300
Morenosite
NiSO4:7H2O = + 1.0000 Ni++ + 1.0000 SO4-- + 7.0000 H2O
log_k -2.0140
-delta_H 12.0185 kJ/mol # Calculated enthalpy of reaction Morenosite
# Enthalpy of formation: -2976.46 kJ/mol
-analytic -2.6654e+002 -7.2132e-002 6.7983e+003 1.0636e+002 1.0616e+002
# -Range: 0-300
Muscovite
KAl3Si3O10(OH)2 +10.0000 H+ = + 1.0000 K+ + 3.0000 Al+++ + 3.0000 SiO2 + 6.0000 H2O
log_k 13.5858
-delta_H -243.224 kJ/mol # Calculated enthalpy of reaction Muscovite
# Enthalpy of formation: -1427.41 kcal/mol
-analytic 3.3085e+001 -1.2425e-002 1.2477e+004 -2.0865e+001 -5.4692e+005
# -Range: 0-300
NH4HSe
NH4HSe = + 1.0000 NH3 + 1.0000 Se-- + 2.0000 H+
log_k -22.0531
-delta_H 0 # Not possible to calculate enthalpy of reaction NH4HSe
# Enthalpy of formation: -133.041 kJ/mol
-analytic -8.8685e+000 6.7342e-003 -5.3028e+003 1.0468e+000 -9.0046e+001
# -Range: 0-200
Na
Na +1.0000 H+ +0.2500 O2 = + 0.5000 H2O + 1.0000 Na+
log_k 67.3804
-delta_H -380.185 kJ/mol # Calculated enthalpy of reaction Na
# Enthalpy of formation: 0 kJ/mol
-analytic -4.0458e+001 -8.7899e-003 2.1223e+004 1.5927e+001 -1.2715e+004
# -Range: 0-300
Na2CO3
Na2CO3 +1.0000 H+ = + 1.0000 HCO3- + 2.0000 Na+
log_k 11.1822
-delta_H -39.8526 kJ/mol # Calculated enthalpy of reaction Na2CO3
# Enthalpy of formation: -1130.68 kJ/mol
-analytic -1.5495e+002 -4.3374e-002 6.4821e+003 6.3571e+001 1.0119e+002
# -Range: 0-300
Na2CO3:7H2O
Na2CO3:7H2O +1.0000 H+ = + 1.0000 HCO3- + 2.0000 Na+ + 7.0000 H2O
log_k 9.9459
-delta_H 27.7881 kJ/mol # Calculated enthalpy of reaction Na2CO3:7H2O
# Enthalpy of formation: -3199.19 kJ/mol
-analytic -2.0593e+002 -3.4509e-003 8.1601e+003 7.6594e+001 1.3864e+002
# -Range: 0-200
Na2Cr2O7
Na2Cr2O7 +1.0000 H2O = + 2.0000 CrO4-- + 2.0000 H+ + 2.0000 Na+
log_k -10.1597
-delta_H 21.9702 kJ/mol # Calculated enthalpy of reaction Na2Cr2O7
# Enthalpy of formation: -473 kcal/mol
-analytic 4.4885e+001 -2.4919e-002 -5.0321e+003 -1.2430e+001 -8.5468e+001
# -Range: 0-200
Na2CrO4
Na2CrO4 = + 1.0000 CrO4-- + 2.0000 Na+
log_k 2.9103
-delta_H -19.5225 kJ/mol # Calculated enthalpy of reaction Na2CrO4
# Enthalpy of formation: -320.8 kcal/mol
-analytic 5.4985e+000 -9.9008e-003 1.0510e+002 0.0000e+000 0.0000e+000
# -Range: 0-200
Na2O
Na2O +2.0000 H+ = + 1.0000 H2O + 2.0000 Na+
log_k 67.4269
-delta_H -351.636 kJ/mol # Calculated enthalpy of reaction Na2O
# Enthalpy of formation: -99.14 kcal/mol
-analytic -6.3585e+001 -8.4695e-003 2.0923e+004 2.5601e+001 3.2651e+002
# -Range: 0-300
Na2Se
Na2Se = + 1.0000 Se-- + 2.0000 Na+
log_k 11.8352
-delta_H 0 # Not possible to calculate enthalpy of reaction Na2Se
# Enthalpy of formation: -81.9 kcal/mol
-analytic -6.0070e+000 8.2821e-003 4.5816e+003 0.0000e+000 0.0000e+000
# -Range: 0-200
Na2Se2
Na2Se2 +1.0000 H2O = + 0.5000 O2 + 2.0000 H+ + 2.0000 Na+ + 2.0000 Se--
log_k -61.3466
-delta_H 0 # Not possible to calculate enthalpy of reaction Na2Se2
# Enthalpy of formation: -92.8 kcal/mol
-analytic -2.7836e+001 7.7035e-003 -1.5040e+004 5.9131e+000 -2.5539e+002
# -Range: 0-200
Na2SiO3
Na2SiO3 +2.0000 H+ = + 1.0000 H2O + 1.0000 SiO2 + 2.0000 Na+
log_k 22.2418
-delta_H -82.7093 kJ/mol # Calculated enthalpy of reaction Na2SiO3
# Enthalpy of formation: -373.19 kcal/mol
-analytic -3.4928e+001 5.6905e-003 1.0284e+004 1.1197e+001 -6.0134e+005
# -Range: 0-300
Na2U2O7
Na2U2O7 +6.0000 H+ = + 2.0000 Na+ + 2.0000 UO2++ + 3.0000 H2O
log_k 22.5917
-delta_H -172.314 kJ/mol # Calculated enthalpy of reaction Na2U2O7
# Enthalpy of formation: -3203.8 kJ/mol
-analytic -8.6640e+001 -1.0903e-002 1.1841e+004 2.9406e+001 1.8479e+002
# -Range: 0-300
Na2UO4(alpha)
Na2UO4 +4.0000 H+ = + 1.0000 UO2++ + 2.0000 H2O + 2.0000 Na+
log_k 30.0231
-delta_H -173.576 kJ/mol # Calculated enthalpy of reaction Na2UO4(alpha)
# Enthalpy of formation: -1897.7 kJ/mol
-analytic -7.9767e+001 -1.0253e-002 1.1963e+004 2.9386e+001 1.8669e+002
# -Range: 0-300
Na3H(SO4)2
Na3H(SO4)2 = + 1.0000 H+ + 2.0000 SO4-- + 3.0000 Na+
log_k -0.8906
-delta_H 0 # Not possible to calculate enthalpy of reaction Na3H(SO4)2
# Enthalpy of formation: 0 kcal/mol
Na3UO4
Na3UO4 +4.0000 H+ = + 1.0000 UO2+ + 2.0000 H2O + 3.0000 Na+
log_k 56.2574
-delta_H -293.703 kJ/mol # Calculated enthalpy of reaction Na3UO4
# Enthalpy of formation: -2024 kJ/mol
-analytic -9.6724e+001 -6.2485e-003 1.9469e+004 3.6180e+001 3.0382e+002
# -Range: 0-300
Na4Ca(SO4)3:2H2O
Na4Ca(SO4)3:2H2O = + 1.0000 Ca++ + 2.0000 H2O + 3.0000 SO4-- + 4.0000 Na+
log_k -5.8938
-delta_H 0 # Not possible to calculate enthalpy of reaction Na4Ca(SO4)3:2H2O
# Enthalpy of formation: 0 kcal/mol
Na4SiO4
Na4SiO4 +4.0000 H+ = + 1.0000 SiO2 + 2.0000 H2O + 4.0000 Na+
log_k 70.6449
-delta_H -327.779 kJ/mol # Calculated enthalpy of reaction Na4SiO4
# Enthalpy of formation: -497.8 kcal/mol
-analytic -1.1969e+002 -6.5032e-003 2.6469e+004 4.4626e+001 -6.2007e+005
# -Range: 0-300
Na4UO2(CO3)3
Na4UO2(CO3)3 +3.0000 H+ = + 1.0000 UO2++ + 3.0000 HCO3- + 4.0000 Na+
log_k 4.0395
-delta_H 0 # Not possible to calculate enthalpy of reaction Na4UO2(CO3)3
# Enthalpy of formation: 0 kcal/mol
Na6Si2O7
Na6Si2O7 +6.0000 H+ = + 2.0000 SiO2 + 3.0000 H2O + 6.0000 Na+
log_k 101.6199
-delta_H -471.951 kJ/mol # Calculated enthalpy of reaction Na6Si2O7
# Enthalpy of formation: -856.3 kcal/mol
-analytic -1.0590e+002 4.5576e-003 3.6830e+004 3.8030e+001 -1.0276e+006
# -Range: 0-300
NaBr
NaBr = + 1.0000 Br- + 1.0000 Na+
log_k 2.9739
-delta_H -0.741032 kJ/mol # Calculated enthalpy of reaction NaBr
# Enthalpy of formation: -361.062 kJ/mol
-analytic -9.3227e+001 -3.2780e-002 2.2910e+003 3.9713e+001 3.5777e+001
# -Range: 0-300
NaBr:2H2O
NaBr:2H2O = + 1.0000 Br- + 1.0000 Na+ + 2.0000 H2O
log_k 2.1040
-delta_H 18.4883 kJ/mol # Calculated enthalpy of reaction NaBr:2H2O
# Enthalpy of formation: -951.968 kJ/mol
-analytic -4.1855e+001 -4.6170e-003 8.3883e+002 1.7182e+001 1.4259e+001
# -Range: 0-200
NaFeO2
NaFeO2 +4.0000 H+ = + 1.0000 Fe+++ + 1.0000 Na+ + 2.0000 H2O
log_k 19.8899
-delta_H -163.339 kJ/mol # Calculated enthalpy of reaction NaFeO2
# Enthalpy of formation: -698.218 kJ/mol
-analytic -7.0047e+001 -9.6226e-003 1.0647e+004 2.3071e+001 1.8082e+002
# -Range: 0-200
NaNpO2CO3:3.5H2O
NaNpO2CO3:3.5H2O +1.0000 H+ = + 1.0000 HCO3- + 1.0000 Na+ + 1.0000 NpO2+ + 3.5000 H2O
log_k -1.2342
-delta_H 27.0979 kJ/mol # Calculated enthalpy of reaction NaNpO2CO3:3.5H2O
# Enthalpy of formation: -2935.76 kJ/mol
-analytic -1.4813e+002 -2.7355e-002 3.6537e+003 5.7701e+001 5.7055e+001
# -Range: 0-300
NaTcO4
NaTcO4 = + 1.0000 Na+ + 1.0000 TcO4-
log_k 1.5208
-delta_H 0 # Not possible to calculate enthalpy of reaction NaTcO4
# Enthalpy of formation: 0 kcal/mol
NaUO3
NaUO3 +2.0000 H+ = + 1.0000 H2O + 1.0000 Na+ + 1.0000 UO2+
log_k 8.3371
-delta_H -56.365 kJ/mol # Calculated enthalpy of reaction NaUO3
# Enthalpy of formation: -1494.9 kJ/mol
-analytic -3.6363e+001 7.0505e-004 4.5359e+003 1.1828e+001 7.0790e+001
# -Range: 0-300
Nahcolite
NaHCO3 = + 1.0000 HCO3- + 1.0000 Na+
log_k -0.1118
-delta_H 17.0247 kJ/mol # Calculated enthalpy of reaction Nahcolite
# Enthalpy of formation: -226.4 kcal/mol
-analytic -2.2282e+002 -5.9693e-002 5.4887e+003 8.9744e+001 8.5712e+001
# -Range: 0-300
Nantokite
CuCl = + 1.0000 Cl- + 1.0000 Cu+
log_k -6.7623
-delta_H 41.9296 kJ/mol # Calculated enthalpy of reaction Nantokite
# Enthalpy of formation: -137.329 kJ/mol
-analytic -2.2442e+001 -1.1201e-002 -1.8709e+003 1.0221e+001 -3.1763e+001
# -Range: 0-200
Natrolite
Na2Al2Si3O10:2H2O +8.0000 H+ = + 2.0000 Al+++ + 2.0000 Na+ + 3.0000 SiO2 + 6.0000 H2O
log_k 18.5204
-delta_H -186.971 kJ/mol # Calculated enthalpy of reaction Natrolite
# Enthalpy of formation: -5718.56 kJ/mol
-analytic -2.7712e+001 -2.7963e-003 1.6075e+004 1.5332e+000 -9.5765e+005
# -Range: 0-300
Natron
Na2CO3:10H2O +1.0000 H+ = + 1.0000 HCO3- + 2.0000 Na+ + 10.0000 H2O
log_k 9.6102
-delta_H 50.4781 kJ/mol # Calculated enthalpy of reaction Natron
# Enthalpy of formation: -4079.39 kJ/mol
-analytic -1.9981e+002 -2.9247e-002 5.2937e+003 8.0973e+001 8.2662e+001
# -Range: 0-300
Natrosilite
Na2Si2O5 +2.0000 H+ = + 1.0000 H2O + 2.0000 Na+ + 2.0000 SiO2
log_k 18.1337
-delta_H -51.7686 kJ/mol # Calculated enthalpy of reaction Natrosilite
# Enthalpy of formation: -590.36 kcal/mol
-analytic -2.7628e+001 1.6865e-002 1.3302e+004 4.2356e+000 -1.2828e+006
# -Range: 0-300
Naumannite
Ag2Se = + 1.0000 Se-- + 2.0000 Ag+
log_k -57.4427
-delta_H 0 # Not possible to calculate enthalpy of reaction Naumannite
# Enthalpy of formation: -37.441 kJ/mol
-analytic -5.3844e+001 -1.0965e-002 -1.4739e+004 1.9842e+001 -2.2998e+002
# -Range: 0-300
Nd
Nd +3.0000 H+ +0.7500 O2 = + 1.0000 Nd+++ + 1.5000 H2O
log_k 182.2233
-delta_H -1116.29 kJ/mol # Calculated enthalpy of reaction Nd
# Enthalpy of formation: 0 kJ/mol
-analytic -2.7390e+002 -5.6545e-002 7.1502e+004 9.7969e+001 -8.2482e+005
# -Range: 0-300
Nd(OH)3
Nd(OH)3 +3.0000 H+ = + 1.0000 Nd+++ + 3.0000 H2O
log_k 18.0852
-delta_H 0 # Not possible to calculate enthalpy of reaction Nd(OH)3
# Enthalpy of formation: 0 kcal/mol
Nd(OH)3(am)
Nd(OH)3 +3.0000 H+ = + 1.0000 Nd+++ + 3.0000 H2O
log_k 20.4852
-delta_H 0 # Not possible to calculate enthalpy of reaction Nd(OH)3(am)
# Enthalpy of formation: 0 kcal/mol
Nd(OH)3(c)
Nd(OH)3 +3.0000 H+ = + 1.0000 Nd+++ + 3.0000 H2O
log_k 15.7852
-delta_H 0 # Not possible to calculate enthalpy of reaction Nd(OH)3(c)
# Enthalpy of formation: 0 kcal/mol
Nd2(CO3)3
Nd2(CO3)3 +3.0000 H+ = + 2.0000 Nd+++ + 3.0000 HCO3-
log_k -3.6636
-delta_H 0 # Not possible to calculate enthalpy of reaction Nd2(CO3)3
# Enthalpy of formation: 0 kcal/mol
Nd2O3
Nd2O3 +6.0000 H+ = + 2.0000 Nd+++ + 3.0000 H2O
log_k 58.6000
-delta_H 0 # Not possible to calculate enthalpy of reaction Nd2O3
# Enthalpy of formation: 0 kcal/mol
NdF3:.5H2O
NdF3:.5H2O = + 0.5000 H2O + 1.0000 Nd+++ + 3.0000 F-
log_k -18.6000
-delta_H 0 # Not possible to calculate enthalpy of reaction NdF3:.5H2O
# Enthalpy of formation: 0 kcal/mol
NdOHCO3
NdOHCO3 +2.0000 H+ = + 1.0000 H2O + 1.0000 HCO3- + 1.0000 Nd+++
log_k 2.8239
-delta_H 0 # Not possible to calculate enthalpy of reaction NdOHCO3
# Enthalpy of formation: 0 kcal/mol
NdPO4:10H2O
NdPO4:10H2O +1.0000 H+ = + 1.0000 HPO4-- + 1.0000 Nd+++ + 10.0000 H2O
log_k -12.1782
-delta_H 0 # Not possible to calculate enthalpy of reaction NdPO4:10H2O
# Enthalpy of formation: 0 kcal/mol
Nepheline
NaAlSiO4 +4.0000 H+ = + 1.0000 Al+++ + 1.0000 Na+ + 1.0000 SiO2 + 2.0000 H2O
log_k 13.8006
-delta_H -135.068 kJ/mol # Calculated enthalpy of reaction Nepheline
# Enthalpy of formation: -500.241 kcal/mol
-analytic -2.4856e+001 -8.8171e-003 8.5653e+003 6.0904e+000 -2.2786e+005
# -Range: 0-300
Nesquehonite
MgCO3:3H2O +1.0000 H+ = + 1.0000 HCO3- + 1.0000 Mg++ + 3.0000 H2O
log_k 4.9955
-delta_H -36.1498 kJ/mol # Calculated enthalpy of reaction Nesquehonite
# Enthalpy of formation: -472.576 kcal/mol
-analytic 1.3771e+002 -6.0397e-002 -3.5049e+004 -1.8831e+001 4.4213e+006
# -Range: 0-300
Ni
Ni +2.0000 H+ +0.5000 O2 = + 1.0000 H2O + 1.0000 Ni++
log_k 50.9914
-delta_H -333.745 kJ/mol # Calculated enthalpy of reaction Ni
# Enthalpy of formation: 0 kcal/mol
-analytic -5.8308e+001 -2.0133e-002 1.8444e+004 2.1590e+001 2.8781e+002
# -Range: 0-300
Ni(OH)2
Ni(OH)2 +2.0000 H+ = + 1.0000 Ni++ + 2.0000 H2O
log_k 12.7485
-delta_H -95.6523 kJ/mol # Calculated enthalpy of reaction Ni(OH)2
# Enthalpy of formation: -529.998 kJ/mol
-analytic -6.5279e+001 -5.9499e-003 7.3471e+003 2.2290e+001 1.2479e+002
# -Range: 0-200
Ni2P2O7
Ni2P2O7 +1.0000 H2O = + 2.0000 HPO4-- + 2.0000 Ni++
log_k -8.8991
-delta_H 0 # Not possible to calculate enthalpy of reaction Ni2P2O7
# Enthalpy of formation: 0 kcal/mol
Ni2SiO4
Ni2SiO4 +4.0000 H+ = + 1.0000 SiO2 + 2.0000 H2O + 2.0000 Ni++
log_k 14.3416
-delta_H -127.629 kJ/mol # Calculated enthalpy of reaction Ni2SiO4
# Enthalpy of formation: -341.705 kcal/mol
-analytic -4.0414e+001 -1.1194e-002 9.6515e+003 1.2026e+001 -3.6336e+005
# -Range: 0-300
Ni3(PO4)2
Ni3(PO4)2 +2.0000 H+ = + 2.0000 HPO4-- + 3.0000 Ni++
log_k -6.6414
-delta_H 0 # Not possible to calculate enthalpy of reaction Ni3(PO4)2
# Enthalpy of formation: 0 kcal/mol
NiCO3
NiCO3 +1.0000 H+ = + 1.0000 HCO3- + 1.0000 Ni++
log_k 3.5118
-delta_H 0 # Not possible to calculate enthalpy of reaction NiCO3
# Enthalpy of formation: 0 kcal/mol
NiCl2
NiCl2 = + 1.0000 Ni++ + 2.0000 Cl-
log_k 8.6113
-delta_H -82.7969 kJ/mol # Calculated enthalpy of reaction NiCl2
# Enthalpy of formation: -305.336 kJ/mol
-analytic -1.2416e+000 -2.3139e-002 2.6529e+003 3.1696e+000 4.5052e+001
# -Range: 0-200
NiCl2:2H2O
NiCl2:2H2O = + 1.0000 Ni++ + 2.0000 Cl- + 2.0000 H2O
log_k 3.9327
-delta_H -37.6746 kJ/mol # Calculated enthalpy of reaction NiCl2:2H2O
# Enthalpy of formation: -922.135 kJ/mol
-analytic -4.8814e+001 -2.2602e-002 2.5951e+003 2.0518e+001 4.4086e+001
# -Range: 0-200
NiCl2:4H2O
NiCl2:4H2O = + 1.0000 Ni++ + 2.0000 Cl- + 4.0000 H2O
log_k 3.8561
-delta_H -15.4373 kJ/mol # Calculated enthalpy of reaction NiCl2:4H2O
# Enthalpy of formation: -1516.05 kJ/mol
-analytic -1.0545e+002 -2.4691e-002 3.9978e+003 4.1727e+001 6.7926e+001
# -Range: 0-200
NiF2
NiF2 = + 1.0000 Ni++ + 2.0000 F-
log_k 0.8772
-delta_H -73.1438 kJ/mol # Calculated enthalpy of reaction NiF2
# Enthalpy of formation: -651.525 kJ/mol
-analytic -2.5291e+002 -8.4179e-002 9.3429e+003 1.0002e+002 1.4586e+002
# -Range: 0-300
NiF2:4H2O
NiF2:4H2O = + 1.0000 Ni++ + 2.0000 F- + 4.0000 H2O
log_k -4.0588
-delta_H 0 # Not possible to calculate enthalpy of reaction NiF2:4H2O
# Enthalpy of formation: 0 kcal/mol
NiSO4
NiSO4 = + 1.0000 Ni++ + 1.0000 SO4--
log_k 5.3197
-delta_H -90.5092 kJ/mol # Calculated enthalpy of reaction NiSO4
# Enthalpy of formation: -873.066 kJ/mol
-analytic -1.8878e+002 -7.6403e-002 7.9412e+003 7.6866e+001 1.2397e+002
# -Range: 0-300
NiSO4:6H2O(alpha)
NiSO4:6H2O = + 1.0000 Ni++ + 1.0000 SO4-- + 6.0000 H2O
log_k -2.0072
-delta_H 4.37983 kJ/mol # Calculated enthalpy of reaction NiSO4:6H2O(alpha)
# Enthalpy of formation: -2682.99 kJ/mol
-analytic -1.1937e+002 -1.3785e-002 4.1543e+003 4.3454e+001 7.0587e+001
# -Range: 0-200
Nickelbischofite
NiCl2:6H2O = + 1.0000 Ni++ + 2.0000 Cl- + 6.0000 H2O
log_k 3.1681
-delta_H 0.064088 kJ/mol # Calculated enthalpy of reaction Nickelbischofite
# Enthalpy of formation: -2103.23 kJ/mol
-analytic -1.4340e+002 -2.1257e-002 5.1858e+003 5.4759e+001 8.8112e+001
# -Range: 0-200
Ningyoite
CaUP2O8:2H2O +2.0000 H+ = + 1.0000 Ca++ + 1.0000 U++++ + 2.0000 H2O + 2.0000 HPO4--
log_k -29.7931
-delta_H -36.4769 kJ/mol # Calculated enthalpy of reaction Ningyoite
# Enthalpy of formation: -1016.65 kcal/mol
-analytic -1.0274e+002 -4.9041e-002 1.7779e+003 3.2973e+001 3.0227e+001
# -Range: 0-200
Niter
KNO3 = + 1.0000 K+ + 1.0000 NO3-
log_k -0.2061
-delta_H 35.4794 kJ/mol # Calculated enthalpy of reaction Niter
# Enthalpy of formation: -494.46 kJ/mol
-analytic -6.5607e+001 -2.8165e-002 -4.0131e+002 3.0361e+001 -6.2425e+000
# -Range: 0-300
Nitrobarite
Ba(NO3)2 = + 1.0000 Ba++ + 2.0000 NO3-
log_k -2.4523
-delta_H 40.8161 kJ/mol # Calculated enthalpy of reaction Nitrobarite
# Enthalpy of formation: -992.082 kJ/mol
-analytic -1.6179e+002 -6.5831e-002 1.2142e+003 7.0664e+001 1.8995e+001
# -Range: 0-300
Nontronite-Ca
Ca.165Fe2Al.33Si3.67H2O12 +7.3200 H+ = + 0.1650 Ca++ + 0.3300 Al+++ + 2.0000 Fe+++ + 3.6700 SiO2 + 4.6600 H2O
log_k -11.5822
-delta_H -38.138 kJ/mol # Calculated enthalpy of reaction Nontronite-Ca
# Enthalpy of formation: -1166.7 kcal/mol
-analytic 1.6291e+001 4.3557e-003 1.0221e+004 -1.8690e+001 -1.5427e+006
# -Range: 0-300
Nontronite-Cs
Cs.33Si4Fe1.67Mg.33H2O12 +6.0000 H+ = + 0.3300 Cs+ + 0.3300 Mg++ + 1.6700 Fe+++ + 4.0000 H2O + 4.0000 SiO2
log_k 5.7975
-delta_H -86.6996 kJ/mol # Calculated enthalpy of reaction Nontronite-Cs
# Enthalpy of formation: -1168.54 kcal/mol
-analytic -1.1646e+001 1.0033e-002 1.7668e+004 -9.0129e+000 -2.0143e+006
# -Range: 0-300
Nontronite-H
H.33Fe2Al.33Si3.67H2O12 +6.9900 H+ = + 0.3300 Al+++ + 2.0000 Fe+++ + 3.6700 SiO2 + 4.6600 H2O
log_k -12.5401
-delta_H -30.452 kJ/mol # Calculated enthalpy of reaction Nontronite-H
# Enthalpy of formation: -1147.12 kcal/mol
-analytic 9.7794e+001 1.4055e-002 4.7440e+003 -4.7272e+001 -1.2103e+006
# -Range: 0-300
Nontronite-K
K.33Fe2Al.33Si3.67H2O12 +7.3200 H+ = + 0.3300 Al+++ + 0.3300 K+ + 2.0000 Fe+++ + 3.6700 SiO2 + 4.6600 H2O
log_k -11.8648
-delta_H -26.5822 kJ/mol # Calculated enthalpy of reaction Nontronite-K
# Enthalpy of formation: -1167.93 kcal/mol
-analytic 1.3630e+001 4.7708e-003 1.0073e+004 -1.7407e+001 -1.5803e+006
# -Range: 0-300
Nontronite-Mg
Mg.165Fe2Al.33Si3.67H2O12 +7.3200 H+ = + 0.1650 Mg++ + 0.3300 Al+++ + 2.0000 Fe+++ + 3.6700 SiO2 + 4.6600 H2O
log_k -11.6200
-delta_H -41.1779 kJ/mol # Calculated enthalpy of reaction Nontronite-Mg
# Enthalpy of formation: -1162.93 kcal/mol
-analytic 5.5961e+001 1.0139e-002 8.0777e+003 -3.3164e+001 -1.4031e+006
# -Range: 0-300
Nontronite-Na
Na.33Fe2Al.33Si3.67H2O12 +7.3200 H+ = + 0.3300 Al+++ + 0.3300 Na+ + 2.0000 Fe+++ + 3.6700 SiO2 + 4.6600 H2O
log_k -11.5263
-delta_H -31.5687 kJ/mol # Calculated enthalpy of reaction Nontronite-Na
# Enthalpy of formation: -1165.8 kcal/mol
-analytic 6.7915e+001 1.2851e-002 7.1218e+003 -3.7112e+001 -1.3758e+006
# -Range: 0-300
Np
Np +4.0000 H+ +1.0000 O2 = + 1.0000 Np++++ + 2.0000 H2O
log_k 174.1077
-delta_H -1115.54 kJ/mol # Calculated enthalpy of reaction Np
# Enthalpy of formation: 0 kJ/mol
-analytic -3.2136e+001 -1.4340e-002 5.7853e+004 6.6512e+000 9.0275e+002
# -Range: 0-300
Np(HPO4)2
Np(HPO4)2 = + 1.0000 Np++++ + 2.0000 HPO4--
log_k -30.9786
-delta_H -18.6219 kJ/mol # Calculated enthalpy of reaction Np(HPO4)2
# Enthalpy of formation: -3121.54 kJ/mol
-analytic -3.6627e+002 -1.3955e-001 7.1370e+003 1.4261e+002 1.1147e+002
# -Range: 0-300
Np(OH)4
Np(OH)4 +4.0000 H+ = + 1.0000 Np++++ + 4.0000 H2O
log_k 0.8103
-delta_H -78.4963 kJ/mol # Calculated enthalpy of reaction Np(OH)4
# Enthalpy of formation: -1620.86 kJ/mol
-analytic -9.5122e+001 -1.0532e-002 7.1132e+003 3.0398e+001 1.1102e+002
# -Range: 0-300
Np2O5
Np2O5 +2.0000 H+ = + 1.0000 H2O + 2.0000 NpO2+
log_k 9.5000
-delta_H -94.4576 kJ/mol # Calculated enthalpy of reaction Np2O5
# Enthalpy of formation: -513.232 kcal/mol
-analytic 5.9974e+003 1.4553e+000 -1.7396e+005 -2.3595e+003 -2.9689e+003
# -Range: 25-150
NpO2
NpO2 +4.0000 H+ = + 1.0000 Np++++ + 2.0000 H2O
log_k -7.8026
-delta_H -53.6087 kJ/mol # Calculated enthalpy of reaction NpO2
# Enthalpy of formation: -1074.07 kJ/mol
-analytic -7.0053e+001 -1.1017e-002 4.4742e+003 2.0421e+001 6.9836e+001
# -Range: 0-300
NpO2(OH)2
NpO2(OH)2 +2.0000 H+ = + 1.0000 NpO2++ + 2.0000 H2O
log_k 5.9851
-delta_H -54.9977 kJ/mol # Calculated enthalpy of reaction NpO2(OH)2
# Enthalpy of formation: -1377.16 kJ/mol
-analytic -2.7351e+001 -1.5987e-003 3.8301e+003 8.4735e+000 5.9773e+001
# -Range: 0-300
NpO2OH(am)
NpO2OH +1.0000 H+ = + 1.0000 H2O + 1.0000 NpO2+
log_k 4.2364
-delta_H -39.6673 kJ/mol # Calculated enthalpy of reaction NpO2OH(am)
# Enthalpy of formation: -1224.16 kJ/mol
-analytic -3.8824e+000 6.7122e-003 2.5390e+003 -9.7040e-001 3.9619e+001
# -Range: 0-300
Okenite
CaSi2O4(OH)2:H2O +2.0000 H+ = + 1.0000 Ca++ + 2.0000 SiO2 + 3.0000 H2O
log_k 10.3816
-delta_H -19.4974 kJ/mol # Calculated enthalpy of reaction Okenite
# Enthalpy of formation: -749.641 kcal/mol
-analytic -7.7353e+001 1.5091e-002 1.3023e+004 2.1337e+001 -1.1831e+006
# -Range: 0-300
Orpiment
As2S3 +6.0000 H2O = + 2.0000 H2AsO3- + 3.0000 HS- + 5.0000 H+
log_k -79.4159
-delta_H 406.539 kJ/mol # Calculated enthalpy of reaction Orpiment
# Enthalpy of formation: -169.423 kJ/mol
-analytic -3.3964e+002 -1.4977e-001 -1.5711e+004 1.4448e+002 -2.4505e+002
# -Range: 0-300
Otavite
CdCO3 +1.0000 H+ = + 1.0000 Cd++ + 1.0000 HCO3-
log_k -1.7712
-delta_H 0 # Not possible to calculate enthalpy of reaction Otavite
# Enthalpy of formation: 0 kcal/mol
Ottemannite
Sn2S3 +3.0000 H+ = + 1.0000 Sn++ + 1.0000 Sn++++ + 3.0000 HS-
log_k -46.2679
-delta_H 236.727 kJ/mol # Calculated enthalpy of reaction Ottemannite
# Enthalpy of formation: -63 kcal/mol
-analytic -6.2863e+001 -5.9171e-002 -1.3469e+004 3.2092e+001 -2.2870e+002
# -Range: 0-200
Oxychloride-Mg
Mg2Cl(OH)3:4H2O +3.0000 H+ = + 1.0000 Cl- + 2.0000 Mg++ + 7.0000 H2O
log_k 25.8319
-delta_H 0 # Not possible to calculate enthalpy of reaction Oxychloride-Mg
# Enthalpy of formation: 0 kcal/mol
P
P +1.5000 H2O +1.2500 O2 = + 1.0000 HPO4-- + 2.0000 H+
log_k 132.1032
-delta_H -848.157 kJ/mol # Calculated enthalpy of reaction P
# Enthalpy of formation: 0 kJ/mol
-analytic -9.2727e+001 -6.8342e-002 4.3465e+004 4.0156e+001 6.7826e+002
# -Range: 0-300
Paragonite
NaAl3Si3O10(OH)2 +10.0000 H+ = + 1.0000 Na+ + 3.0000 Al+++ + 3.0000 SiO2 + 6.0000 H2O
log_k 17.5220
-delta_H -275.056 kJ/mol # Calculated enthalpy of reaction Paragonite
# Enthalpy of formation: -1416.96 kcal/mol
-analytic 3.5507e+001 -1.0720e-002 1.3519e+004 -2.2283e+001 -4.5657e+005
# -Range: 0-300
Paralaurionite
PbClOH +1.0000 H+ = + 1.0000 Cl- + 1.0000 H2O + 1.0000 Pb++
log_k 0.2035
-delta_H 8.41948 kJ/mol # Calculated enthalpy of reaction Paralaurionite
# Enthalpy of formation: -460.417 kJ/mol
-analytic -1.1245e+001 -1.0520e-002 -5.3551e+002 6.6175e+000 -9.0896e+000
# -Range: 0-200
Pargasite
NaCa2Al3Mg4Si6O22(OH)2 +22.0000 H+ = + 1.0000 Na+ + 2.0000 Ca++ + 3.0000 Al+++ + 4.0000 Mg++ + 6.0000 SiO2 + 12.0000 H2O
log_k 101.9939
-delta_H -880.205 kJ/mol # Calculated enthalpy of reaction Pargasite
# Enthalpy of formation: -3016.62 kcal/mol
-analytic -6.7889e+001 -3.7817e-002 5.0493e+004 9.2705e+000 -1.0163e+006
# -Range: 0-300
Parsonsite
Pb2UO2(PO4)2:2H2O +2.0000 H+ = + 1.0000 UO2++ + 2.0000 H2O + 2.0000 HPO4-- + 2.0000 Pb++
log_k -27.7911
-delta_H 0 # Not possible to calculate enthalpy of reaction Parsonsite
# Enthalpy of formation: 0 kcal/mol
Pb
Pb +2.0000 H+ +0.5000 O2 = + 1.0000 H2O + 1.0000 Pb++
log_k 47.1871
-delta_H -278.851 kJ/mol # Calculated enthalpy of reaction Pb
# Enthalpy of formation: 0 kJ/mol
-analytic -3.1784e+001 -1.4816e-002 1.4984e+004 1.3383e+001 2.3381e+002
# -Range: 0-300
Pb(H2PO4)2
Pb(H2PO4)2 = + 1.0000 Pb++ + 2.0000 H+ + 2.0000 HPO4--
log_k -9.8400
-delta_H 0 # Not possible to calculate enthalpy of reaction Pb(H2PO4)2
# Enthalpy of formation: 0 kcal/mol
Pb(IO3)2
Pb(IO3)2 = + 1.0000 Pb++ + 2.0000 IO3-
log_k -12.5173
-delta_H 53.7783 kJ/mol # Calculated enthalpy of reaction Pb(IO3)2
# Enthalpy of formation: -495.525 kJ/mol
-analytic -5.3573e+000 -1.4164e-002 -3.6236e+003 3.7209e+000 -6.1532e+001
# -Range: 0-200
Pb(N3)2(mono)
Pb(N3)2 = + 1.0000 Pb++ + 2.0000 N3-
log_k -8.3583
-delta_H 72.9495 kJ/mol # Calculated enthalpy of reaction Pb(N3)2(mono)
# Enthalpy of formation: 478.251 kJ/mol
-analytic 6.0051e+001 -1.1168e-002 -7.0041e+003 -1.6812e+001 -1.1896e+002
# -Range: 0-200
Pb(N3)2(orth)
Pb(N3)2 = + 1.0000 Pb++ + 2.0000 N3-
log_k -8.7963
-delta_H 75.0615 kJ/mol # Calculated enthalpy of reaction Pb(N3)2(orth)
# Enthalpy of formation: 476.139 kJ/mol
-analytic 5.9779e+001 -1.1215e-002 -7.1081e+003 -1.6732e+001 -1.2073e+002
# -Range: 0-200
Pb(Thiocyanate)2
Pb(Thiocyanate)2 = + 1.0000 Pb++ + 2.0000 Thiocyanate-
log_k -0.0910
-delta_H 0 # Not possible to calculate enthalpy of reaction Pb(Thiocyanate)2
# Enthalpy of formation: 151.212 kJ/mol
-analytic 7.4247e+000 -1.6226e-002 0.0000e+000 0.0000e+000 -2.3938e+005
# -Range: 0-200
Pb2Cl2CO3
Pb2Cl2CO3 +1.0000 H+ = + 1.0000 HCO3- + 2.0000 Cl- + 2.0000 Pb++
log_k -9.6180
-delta_H 0 # Not possible to calculate enthalpy of reaction Pb2Cl2CO3
# Enthalpy of formation: 0 kcal/mol
Pb2Cl5NH4
Pb2Cl5NH4 = + 1.0000 H+ + 1.0000 NH3 + 2.0000 Pb++ + 5.0000 Cl-
log_k -19.6100
-delta_H 119.617 kJ/mol # Calculated enthalpy of reaction Pb2Cl5NH4
# Enthalpy of formation: -1034.51 kJ/mol
-analytic 1.3149e+001 -4.8598e-002 -9.8473e+003 5.9552e+000 -1.6723e+002
# -Range: 0-200
Pb2O(N3)2
Pb2O(N3)2 +2.0000 H+ = + 1.0000 H2O + 2.0000 N3- + 2.0000 Pb++
log_k -13.7066
-delta_H 0 # Not possible to calculate enthalpy of reaction Pb2O(N3)2
# Enthalpy of formation: 0 kcal/mol
Pb2SiO4
Pb2SiO4 +4.0000 H+ = + 1.0000 SiO2 + 2.0000 H2O + 2.0000 Pb++
log_k 18.0370
-delta_H -83.9883 kJ/mol # Calculated enthalpy of reaction Pb2SiO4
# Enthalpy of formation: -1363.55 kJ/mol
-analytic 2.7287e+002 6.3875e-002 -3.7001e+003 -1.0568e+002 -6.2927e+001
# -Range: 0-200
Pb3(PO4)2
Pb3(PO4)2 +2.0000 H+ = + 2.0000 HPO4-- + 3.0000 Pb++
log_k -19.9744
-delta_H 0 # Not possible to calculate enthalpy of reaction Pb3(PO4)2
# Enthalpy of formation: 0 kcal/mol
Pb3SO6
Pb3SO6 +4.0000 H+ = + 1.0000 SO4-- + 2.0000 H2O + 3.0000 Pb++
log_k 10.5981
-delta_H -79.3438 kJ/mol # Calculated enthalpy of reaction Pb3SO6
# Enthalpy of formation: -1399.17 kJ/mol
-analytic -5.3308e+000 -1.8639e-002 3.0245e+003 4.5760e+000 5.1362e+001
# -Range: 0-200
Pb4Cl2(OH)6
Pb4Cl2(OH)6 +6.0000 H+ = + 2.0000 Cl- + 4.0000 Pb++ + 6.0000 H2O
log_k 17.2793
-delta_H 0 # Not possible to calculate enthalpy of reaction Pb4Cl2(OH)6
# Enthalpy of formation: 0 kcal/mol
Pb4O(PO4)2
Pb4O(PO4)2 +4.0000 H+ = + 1.0000 H2O + 2.0000 HPO4-- + 4.0000 Pb++
log_k -12.5727
-delta_H 0 # Not possible to calculate enthalpy of reaction Pb4O(PO4)2
# Enthalpy of formation: 0 kcal/mol
Pb4SO7
Pb4SO7 +6.0000 H+ = + 1.0000 SO4-- + 3.0000 H2O + 4.0000 Pb++
log_k 21.7354
-delta_H -136.566 kJ/mol # Calculated enthalpy of reaction Pb4SO7
# Enthalpy of formation: -1626.87 kJ/mol
-analytic -2.6884e+001 -2.1429e-002 6.8390e+003 1.2951e+001 1.1614e+002
# -Range: 0-200
PbBr2
PbBr2 = + 1.0000 Pb++ + 2.0000 Br-
log_k -5.2413
-delta_H 36.3838 kJ/mol # Calculated enthalpy of reaction PbBr2
# Enthalpy of formation: -278.47 kJ/mol
-analytic 3.0977e+001 -1.6567e-002 -4.2879e+003 -6.8329e+000 -7.2825e+001
# -Range: 0-200
PbBrF
PbBrF = + 1.0000 Br- + 1.0000 F- + 1.0000 Pb++
log_k -8.0418
-delta_H 0 # Not possible to calculate enthalpy of reaction PbBrF
# Enthalpy of formation: 0 kcal/mol
PbCO3.PbO
PbCO3.PbO +3.0000 H+ = + 1.0000 H2O + 1.0000 HCO3- + 2.0000 Pb++
log_k 9.6711
-delta_H -55.4286 kJ/mol # Calculated enthalpy of reaction PbCO3.PbO
# Enthalpy of formation: -918.502 kJ/mol
-analytic -4.2160e+001 -1.4124e-002 3.8661e+003 1.7404e+001 6.5667e+001
# -Range: 0-200
PbF2
PbF2 = + 1.0000 Pb++ + 2.0000 F-
log_k -5.2047
-delta_H -5.83772 kJ/mol # Calculated enthalpy of reaction PbF2
# Enthalpy of formation: -663.937 kJ/mol
-analytic -2.2712e+002 -7.9552e-002 5.2198e+003 9.2173e+001 8.1516e+001
# -Range: 0-300
PbFCl
PbFCl = + 1.0000 Cl- + 1.0000 F- + 1.0000 Pb++
log_k -8.9820
-delta_H 33.1852 kJ/mol # Calculated enthalpy of reaction PbFCl
# Enthalpy of formation: -534.692 kJ/mol
-analytic 6.1688e+000 -2.0732e-002 -3.4666e+003 1.0697e+000 -5.8869e+001
# -Range: 0-200
PbHPO4
PbHPO4 = + 1.0000 HPO4-- + 1.0000 Pb++
log_k -15.7275
-delta_H 0 # Not possible to calculate enthalpy of reaction PbHPO4
# Enthalpy of formation: 0 kcal/mol
PbI2
PbI2 = + 1.0000 Pb++ + 2.0000 I-
log_k -8.0418
-delta_H 62.5717 kJ/mol # Calculated enthalpy of reaction PbI2
# Enthalpy of formation: -175.456 kJ/mol
-analytic 1.5277e+001 -2.0582e-002 -5.1256e+003 0.0000e+000 0.0000e+000
# -Range: 0-200
PbSO4(NH3)2
PbSO4(NH3)2 = + 1.0000 Pb++ + 1.0000 SO4-- + 2.0000 NH3
log_k -2.0213
-delta_H 28.284 kJ/mol # Calculated enthalpy of reaction PbSO4(NH3)2
# Enthalpy of formation: -1099.64 kJ/mol
-analytic 3.5718e-001 -1.0192e-002 -2.0095e+003 2.9853e+000 -3.4124e+001
# -Range: 0-200
PbSO4(NH3)4
PbSO4(NH3)4 = + 1.0000 Pb++ + 1.0000 SO4-- + 4.0000 NH3
log_k 1.5024
-delta_H 31.155 kJ/mol # Calculated enthalpy of reaction PbSO4(NH3)4
# Enthalpy of formation: -1265.18 kJ/mol
-analytic -4.1080e+001 -7.2307e-003 6.6637e+001 1.7984e+001 1.1460e+000
# -Range: 0-200
PbSeO4
PbSeO4 = + 1.0000 Pb++ + 1.0000 SeO4--
log_k -6.9372
-delta_H 10.8967 kJ/mol # Calculated enthalpy of reaction PbSeO4
# Enthalpy of formation: -609.125 kJ/mol
-analytic 3.1292e+001 -1.4192e-002 -3.0980e+003 -9.5448e+000 -5.2618e+001
# -Range: 0-200
Pd
Pd +2.0000 H+ +0.5000 O2 = + 1.0000 H2O + 1.0000 Pd++
log_k 12.0688
-delta_H -103.709 kJ/mol # Calculated enthalpy of reaction Pd
# Enthalpy of formation: 0 kcal/mol
-analytic -6.2530e+001 -1.9774e-002 6.7013e+003 2.3441e+001 1.0459e+002
# -Range: 0-300
PdO
PdO +2.0000 H+ = + 1.0000 H2O + 1.0000 Pd++
log_k 0.0643
-delta_H -24.422 kJ/mol # Calculated enthalpy of reaction PdO
# Enthalpy of formation: -20.4 kcal/mol
-analytic -8.8921e+001 -1.9031e-002 3.8537e+003 3.3028e+001 6.0159e+001
# -Range: 0-300
Penroseite
NiSe2 +1.0000 H2O = + 0.5000 O2 + 1.0000 Ni++ + 2.0000 H+ + 2.0000 Se--
log_k -98.8004
-delta_H 0 # Not possible to calculate enthalpy of reaction Penroseite
# Enthalpy of formation: -26 kcal/mol
-analytic -4.7339e+001 -1.2035e-002 -2.3589e+004 1.2624e+001 -3.6808e+002
# -Range: 0-300
Pentahydrite
MgSO4:5H2O = + 1.0000 Mg++ + 1.0000 SO4-- + 5.0000 H2O
log_k -1.3872
-delta_H 0 # Not possible to calculate enthalpy of reaction Pentahydrite
# Enthalpy of formation: 0 kcal/mol
Periclase
MgO +2.0000 H+ = + 1.0000 H2O + 1.0000 Mg++
log_k 21.3354
-delta_H -150.139 kJ/mol # Calculated enthalpy of reaction Periclase
# Enthalpy of formation: -143.8 kcal/mol
-analytic -8.8465e+001 -1.8390e-002 1.0414e+004 3.2469e+001 1.6253e+002
# -Range: 0-300
Petalite
LiAlSi4O10 +4.0000 H+ = + 1.0000 Al+++ + 1.0000 Li+ + 2.0000 H2O + 4.0000 SiO2
log_k -3.8153
-delta_H -13.1739 kJ/mol # Calculated enthalpy of reaction Petalite
# Enthalpy of formation: -4886.15 kJ/mol
-analytic -6.6355e+000 2.4316e-002 1.5949e+004 -1.3341e+001 -2.2265e+006
# -Range: 0-300
Phlogopite
KAlMg3Si3O10(OH)2 +10.0000 H+ = + 1.0000 Al+++ + 1.0000 K+ + 3.0000 Mg++ + 3.0000 SiO2 + 6.0000 H2O
log_k 37.4400
-delta_H -310.503 kJ/mol # Calculated enthalpy of reaction Phlogopite
# Enthalpy of formation: -1488.07 kcal/mol
-analytic -8.7730e+001 -1.7253e-002 2.3748e+004 2.4465e+001 -8.9045e+005
# -Range: 0-300
Phosgenite
Pb2(CO3)Cl2 +1.0000 H+ = + 1.0000 HCO3- + 2.0000 Cl- + 2.0000 Pb++
log_k -9.6355
-delta_H 49.0844 kJ/mol # Calculated enthalpy of reaction Phosgenite
# Enthalpy of formation: -1071.34 kJ/mol
-analytic 3.4909e+000 -2.9365e-002 -4.6327e+003 4.5068e+000 -7.8671e+001
# -Range: 0-200
Picromerite
K2Mg(SO4)2:6H2O = + 1.0000 Mg++ + 2.0000 K+ + 2.0000 SO4-- + 6.0000 H2O
log_k -4.4396
-delta_H 0 # Not possible to calculate enthalpy of reaction Picromerite
# Enthalpy of formation: 0 kcal/mol
Pirssonite
Na2Ca(CO3)2:2H2O +2.0000 H+ = + 1.0000 Ca++ + 2.0000 H2O + 2.0000 HCO3- + 2.0000 Na+
log_k 11.3230
-delta_H 0 # Not possible to calculate enthalpy of reaction Pirssonite
# Enthalpy of formation: 0 kcal/mol
Plattnerite
PbO2 +4.0000 H+ = + 1.0000 Pb++++ + 2.0000 H2O
log_k -7.9661
-delta_H 0 # Not possible to calculate enthalpy of reaction Plattnerite
# Enthalpy of formation: -277.363 kJ/mol
Plumbogummite
PbAl3(PO4)2(OH)5:H2O +7.0000 H+ = + 1.0000 Pb++ + 2.0000 HPO4-- + 3.0000 Al+++ + 6.0000 H2O
log_k -8.1463
-delta_H 0 # Not possible to calculate enthalpy of reaction Plumbogummite
# Enthalpy of formation: 0 kcal/mol
Pm
Pm +3.0000 H+ +0.7500 O2 = + 1.0000 Pm+++ + 1.5000 H2O
log_k 180.6737
-delta_H 0 # Not possible to calculate enthalpy of reaction Pm
# Enthalpy of formation: 0 kcal/mol
Pm(OH)3
Pm(OH)3 +3.0000 H+ = + 1.0000 Pm+++ + 3.0000 H2O
log_k 17.4852
-delta_H 0 # Not possible to calculate enthalpy of reaction Pm(OH)3
# Enthalpy of formation: 0 kcal/mol
Pm(OH)3(am)
Pm(OH)3 +3.0000 H+ = + 1.0000 Pm+++ + 3.0000 H2O
log_k 18.2852
-delta_H 0 # Not possible to calculate enthalpy of reaction Pm(OH)3(am)
# Enthalpy of formation: 0 kcal/mol
Pm2(CO3)3
Pm2(CO3)3 +3.0000 H+ = + 2.0000 Pm+++ + 3.0000 HCO3-
log_k -3.5636
-delta_H 0 # Not possible to calculate enthalpy of reaction Pm2(CO3)3
# Enthalpy of formation: 0 kcal/mol
Pm2O3
Pm2O3 +6.0000 H+ = + 2.0000 Pm+++ + 3.0000 H2O
log_k 48.8000
-delta_H 0 # Not possible to calculate enthalpy of reaction Pm2O3
# Enthalpy of formation: 0 kcal/mol
PmF3:.5H2O
PmF3:.5H2O = + 0.5000 H2O + 1.0000 Pm+++ + 3.0000 F-
log_k -18.1000
-delta_H 0 # Not possible to calculate enthalpy of reaction PmF3:.5H2O
# Enthalpy of formation: 0 kcal/mol
PmPO4:10H2O
PmPO4:10H2O +1.0000 H+ = + 1.0000 HPO4-- + 1.0000 Pm+++ + 10.0000 H2O
log_k -12.1782
-delta_H 0 # Not possible to calculate enthalpy of reaction PmPO4:10H2O
# Enthalpy of formation: 0 kcal/mol
Polydymite
Ni3S4 +2.0000 H+ = + 1.0000 S2-- + 2.0000 HS- + 3.0000 Ni++
log_k -48.9062
-delta_H 0 # Not possible to calculate enthalpy of reaction Polydymite
# Enthalpy of formation: -78.014 kcal/mol
-analytic -1.8030e+001 -4.6945e-002 -1.1557e+004 8.8339e+000 -1.9625e+002
# -Range: 0-200
Polyhalite
K2MgCa2(SO4)4:2H2O = + 1.0000 Mg++ + 2.0000 Ca++ + 2.0000 H2O + 2.0000 K+ + 4.0000 SO4--
log_k -14.3124
-delta_H 0 # Not possible to calculate enthalpy of reaction Polyhalite
# Enthalpy of formation: 0 kcal/mol
Portlandite
Ca(OH)2 +2.0000 H+ = + 1.0000 Ca++ + 2.0000 H2O
log_k 22.5552
-delta_H -128.686 kJ/mol # Calculated enthalpy of reaction Portlandite
# Enthalpy of formation: -986.074 kJ/mol
-analytic -8.3848e+001 -1.8373e-002 9.3154e+003 3.2584e+001 1.4538e+002
# -Range: 0-300
Pr
Pr +3.0000 H+ +0.7500 O2 = + 1.0000 Pr+++ + 1.5000 H2O
log_k 183.6893
-delta_H -1125.92 kJ/mol # Calculated enthalpy of reaction Pr
# Enthalpy of formation: 0 kJ/mol
-analytic -4.1136e+002 -7.5853e-002 7.9974e+004 1.4718e+002 -1.3148e+006
# -Range: 0-300
Pr(OH)3
Pr(OH)3 +3.0000 H+ = + 1.0000 Pr+++ + 3.0000 H2O
log_k 19.5852
-delta_H 0 # Not possible to calculate enthalpy of reaction Pr(OH)3
# Enthalpy of formation: 0 kcal/mol
Pr(OH)3(am)
Pr(OH)3 +3.0000 H+ = + 1.0000 Pr+++ + 3.0000 H2O
log_k 21.0852
-delta_H 0 # Not possible to calculate enthalpy of reaction Pr(OH)3(am)
# Enthalpy of formation: 0 kcal/mol
Pr2(CO3)3
Pr2(CO3)3 +3.0000 H+ = + 2.0000 Pr+++ + 3.0000 HCO3-
log_k -3.8136
-delta_H 0 # Not possible to calculate enthalpy of reaction Pr2(CO3)3
# Enthalpy of formation: 0 kcal/mol
Pr2O3
Pr2O3 +6.0000 H+ = + 2.0000 Pr+++ + 3.0000 H2O
log_k 61.4000
-delta_H 0 # Not possible to calculate enthalpy of reaction Pr2O3
# Enthalpy of formation: 0 kcal/mol
PrF3:.5H2O
PrF3:.5H2O = + 0.5000 H2O + 1.0000 Pr+++ + 3.0000 F-
log_k -18.7000
-delta_H 0 # Not possible to calculate enthalpy of reaction PrF3:.5H2O
# Enthalpy of formation: 0 kcal/mol
PrPO4:10H2O
PrPO4:10H2O +1.0000 H+ = + 1.0000 HPO4-- + 1.0000 Pr+++ + 10.0000 H2O
log_k -12.2782
-delta_H 0 # Not possible to calculate enthalpy of reaction PrPO4:10H2O
# Enthalpy of formation: 0 kcal/mol
Prehnite
Ca2Al2Si3O10(OH)2 +10.0000 H+ = + 2.0000 Al+++ + 2.0000 Ca++ + 3.0000 SiO2 + 6.0000 H2O
log_k 32.9305
-delta_H -311.875 kJ/mol # Calculated enthalpy of reaction Prehnite
# Enthalpy of formation: -1481.65 kcal/mol
-analytic -3.5763e+001 -2.1396e-002 2.0167e+004 6.3554e+000 -7.4967e+005
# -Range: 0-300
Przhevalskite
Pb(UO2)2(PO4)2 +2.0000 H+ = + 1.0000 Pb++ + 2.0000 HPO4-- + 2.0000 UO2++
log_k -20.0403
-delta_H -71.1058 kJ/mol # Calculated enthalpy of reaction Przhevalskite
# Enthalpy of formation: -1087.51 kcal/mol
-analytic -2.9817e+001 -4.0756e-002 1.0077e+003 7.4885e+000 1.7122e+001
# -Range: 0-200
Pseudowollastonite
CaSiO3 +2.0000 H+ = + 1.0000 Ca++ + 1.0000 H2O + 1.0000 SiO2
log_k 13.9997
-delta_H -79.4625 kJ/mol # Calculated enthalpy of reaction Pseudowollastonite
# Enthalpy of formation: -388.9 kcal/mol
-analytic 2.6691e+001 6.3323e-003 5.5723e+003 -1.1822e+001 -3.6038e+005
# -Range: 0-300
Pu
Pu +4.0000 H+ +1.0000 O2 = + 1.0000 Pu++++ + 2.0000 H2O
log_k 170.3761
-delta_H -1095.44 kJ/mol # Calculated enthalpy of reaction Pu
# Enthalpy of formation: 0 kJ/mol
-analytic -1.9321e+002 -3.4314e-002 6.6737e+004 6.3552e+001 -6.4737e+005
# -Range: 0-300
Pu(HPO4)2
Pu(HPO4)2 = + 1.0000 Pu++++ + 2.0000 HPO4--
log_k -27.7025
-delta_H -33.4449 kJ/mol # Calculated enthalpy of reaction Pu(HPO4)2
# Enthalpy of formation: -3086.61 kJ/mol
-analytic -3.6565e+002 -1.3961e-001 7.9105e+003 1.4265e+002 1.2354e+002
# -Range: 0-300
Pu(OH)3
Pu(OH)3 +3.0000 H+ = + 1.0000 Pu+++ + 3.0000 H2O
log_k 22.4499
-delta_H -148.067 kJ/mol # Calculated enthalpy of reaction Pu(OH)3
# Enthalpy of formation: -1301 kJ/mol
-analytic -6.1342e+001 -8.6952e-003 9.7733e+003 2.1664e+001 1.5252e+002
# -Range: 0-300
Pu(OH)4
Pu(OH)4 +4.0000 H+ = + 1.0000 Pu++++ + 4.0000 H2O
log_k 0.7578
-delta_H -68.6543 kJ/mol # Calculated enthalpy of reaction Pu(OH)4
# Enthalpy of formation: -1610.59 kJ/mol
-analytic -9.3473e+001 -1.0579e-002 6.5974e+003 3.0415e+001 1.0297e+002
# -Range: 0-300
Pu2O3
Pu2O3 +6.0000 H+ = + 2.0000 Pu+++ + 3.0000 H2O
log_k 48.1332
-delta_H -360.26 kJ/mol # Calculated enthalpy of reaction Pu2O3
# Enthalpy of formation: -1680.36 kJ/mol
-analytic -8.7831e+001 -1.9784e-002 2.0832e+004 2.9096e+001 3.2509e+002
# -Range: 0-300
PuF3
PuF3 = + 1.0000 Pu+++ + 3.0000 F-
log_k -10.1872
-delta_H -46.2608 kJ/mol # Calculated enthalpy of reaction PuF3
# Enthalpy of formation: -1551.33 kJ/mol
-analytic -3.1104e+002 -1.0854e-001 8.7435e+003 1.2279e+002 1.3653e+002
# -Range: 0-300
PuF4
PuF4 = + 1.0000 Pu++++ + 4.0000 F-
log_k -13.2091
-delta_H -100.039 kJ/mol # Calculated enthalpy of reaction PuF4
# Enthalpy of formation: -1777.24 kJ/mol
-analytic -4.3072e+002 -1.4500e-001 1.4076e+004 1.6709e+002 2.1977e+002
# -Range: 0-300
PuO2
PuO2 +4.0000 H+ = + 1.0000 Pu++++ + 2.0000 H2O
log_k -7.3646
-delta_H -51.8827 kJ/mol # Calculated enthalpy of reaction PuO2
# Enthalpy of formation: -1055.69 kJ/mol
-analytic -7.1933e+001 -1.1841e-002 4.4494e+003 2.1491e+001 6.9450e+001
# -Range: 0-300
PuO2(OH)2
PuO2(OH)2 +2.0000 H+ = + 1.0000 PuO2++ + 2.0000 H2O
log_k 3.5499
-delta_H -35.7307 kJ/mol # Calculated enthalpy of reaction PuO2(OH)2
# Enthalpy of formation: -1357.52 kJ/mol
-analytic -2.6536e+001 -1.6542e-003 2.8262e+003 8.5277e+000 4.4108e+001
# -Range: 0-300
PuO2HPO4
PuO2HPO4 = + 1.0000 HPO4-- + 1.0000 PuO2++
log_k -12.6074
-delta_H -10.108 kJ/mol # Calculated enthalpy of reaction PuO2HPO4
# Enthalpy of formation: -2103.55 kJ/mol
-analytic -1.6296e+002 -6.6166e-002 3.0557e+003 6.4577e+001 4.7729e+001
# -Range: 0-300
PuO2OH(am)
PuO2OH +1.0000 H+ = + 1.0000 H2O + 1.0000 PuO2+
log_k 5.4628
-delta_H -42.4933 kJ/mol # Calculated enthalpy of reaction PuO2OH(am)
# Enthalpy of formation: -1157.53 kJ/mol
-analytic -3.1316e+000 6.7573e-003 2.6884e+003 -9.8622e-001 4.1951e+001
# -Range: 0-300
Pyrite
FeS2 +1.0000 H2O = + 0.2500 H+ + 0.2500 SO4-- + 1.0000 Fe++ + 1.7500 HS-
log_k -24.6534
-delta_H 109.535 kJ/mol # Calculated enthalpy of reaction Pyrite
# Enthalpy of formation: -41 kcal/mol
-analytic -2.4195e+002 -8.7948e-002 -6.2911e+002 9.9248e+001 -9.7454e+000
# -Range: 0-300
Pyrolusite
MnO2 = + 0.5000 Mn++ + 0.5000 MnO4--
log_k -17.6439
-delta_H 83.3804 kJ/mol # Calculated enthalpy of reaction Pyrolusite
# Enthalpy of formation: -520.031 kJ/mol
-analytic -1.1541e+002 -4.1665e-002 -1.8960e+003 4.7094e+001 -2.9551e+001
# -Range: 0-300
Pyromorphite
Pb5(PO4)3Cl +3.0000 H+ = + 1.0000 Cl- + 3.0000 HPO4-- + 5.0000 Pb++
log_k -47.8954
-delta_H 0 # Not possible to calculate enthalpy of reaction Pyromorphite
# Enthalpy of formation: 0 kcal/mol
Pyromorphite-OH
Pb5(OH)(PO4)3 +4.0000 H+ = + 1.0000 H2O + 3.0000 HPO4-- + 5.0000 Pb++
log_k -26.2653
-delta_H 0 # Not possible to calculate enthalpy of reaction Pyromorphite-OH
# Enthalpy of formation: 0 kcal/mol
Pyrophyllite
Al2Si4O10(OH)2 +6.0000 H+ = + 2.0000 Al+++ + 4.0000 H2O + 4.0000 SiO2
log_k 0.4397
-delta_H -102.161 kJ/mol # Calculated enthalpy of reaction Pyrophyllite
# Enthalpy of formation: -1345.31 kcal/mol
-analytic 1.1066e+001 1.2707e-002 1.6417e+004 -1.9596e+001 -1.8791e+006
# -Range: 0-300
Pyrrhotite
FeS +1.0000 H+ = + 1.0000 Fe++ + 1.0000 HS-
log_k -3.7193
-delta_H -7.9496 kJ/mol # Calculated enthalpy of reaction Pyrrhotite
# Enthalpy of formation: -24 kcal/mol
-analytic -1.5785e+002 -5.2258e-002 3.9711e+003 6.3195e+001 6.2012e+001
# -Range: 0-300
Quartz
SiO2 = + 1.0000 SiO2
log_k -3.9993
-delta_H 32.949 kJ/mol # Calculated enthalpy of reaction Quartz
# Enthalpy of formation: -217.65 kcal/mol
-analytic 7.7698e-002 1.0612e-002 3.4651e+003 -4.3551e+000 -7.2138e+005
# -Range: 0-300
Ra
Ra +2.0000 H+ +0.5000 O2 = + 1.0000 H2O + 1.0000 Ra++
log_k 141.3711
-delta_H -807.374 kJ/mol # Calculated enthalpy of reaction Ra
# Enthalpy of formation: 0 kJ/mol
-analytic 4.9867e+001 5.9412e-003 4.0293e+004 -1.8356e+001 6.8421e+002
# -Range: 0-200
Ra(NO3)2
Ra(NO3)2 = + 1.0000 Ra++ + 2.0000 NO3-
log_k -2.2419
-delta_H 50.4817 kJ/mol # Calculated enthalpy of reaction Ra(NO3)2
# Enthalpy of formation: -991.706 kJ/mol
-analytic 2.2001e+001 -9.5263e-003 -3.9389e+003 -3.3143e+000 -6.6896e+001
# -Range: 0-200
RaCl2:2H2O
RaCl2:2H2O = + 1.0000 Ra++ + 2.0000 Cl- + 2.0000 H2O
log_k -0.7647
-delta_H 32.6266 kJ/mol # Calculated enthalpy of reaction RaCl2:2H2O
# Enthalpy of formation: -1466.07 kJ/mol
-analytic -2.5033e+001 -1.8918e-002 -1.5713e+003 1.4213e+001 -2.6673e+001
# -Range: 0-200
RaSO4
RaSO4 = + 1.0000 Ra++ + 1.0000 SO4--
log_k -10.4499
-delta_H 40.309 kJ/mol # Calculated enthalpy of reaction RaSO4
# Enthalpy of formation: -1477.51 kJ/mol
-analytic 4.8025e+001 -1.1376e-002 -5.1347e+003 -1.5306e+001 -8.7211e+001
# -Range: 0-200
Rankinite
Ca3Si2O7 +6.0000 H+ = + 2.0000 SiO2 + 3.0000 Ca++ + 3.0000 H2O
log_k 51.9078
-delta_H -302.089 kJ/mol # Calculated enthalpy of reaction Rankinite
# Enthalpy of formation: -941.7 kcal/mol
-analytic -9.6393e+001 -1.6592e-002 2.4832e+004 3.2541e+001 -9.4630e+005
# -Range: 0-300
Rb
Rb +1.0000 H+ +0.2500 O2 = + 0.5000 H2O + 1.0000 Rb+
log_k 71.1987
-delta_H -391.009 kJ/mol # Calculated enthalpy of reaction Rb
# Enthalpy of formation: 0 kJ/mol
-analytic -2.1179e+001 -8.7978e-003 2.0934e+004 1.0011e+001 3.2667e+002
# -Range: 0-300
Rb2UO4
Rb2UO4 +4.0000 H+ = + 1.0000 UO2++ + 2.0000 H2O + 2.0000 Rb+
log_k 34.0089
-delta_H -170.224 kJ/mol # Calculated enthalpy of reaction Rb2UO4
# Enthalpy of formation: -1922.7 kJ/mol
-analytic -3.8205e+001 3.1862e-003 1.0973e+004 1.3925e+001 1.8636e+002
# -Range: 0-200
Re
Re +1.7500 O2 +0.5000 H2O = + 1.0000 H+ + 1.0000 ReO4-
log_k 105.9749
-delta_H -623.276 kJ/mol # Calculated enthalpy of reaction Re
# Enthalpy of formation: 0 kJ/mol
-analytic 1.4535e+001 -2.9877e-002 2.9910e+004 0.0000e+000 0.0000e+000
# -Range: 0-300
Realgar
AsS +2.0000 H2O = + 0.5000 S2O4-- + 1.0000 AsH3 + 1.0000 H+
log_k -60.2768
-delta_H 0 # Not possible to calculate enthalpy of reaction Realgar
# Enthalpy of formation: -71.406 kJ/mol
Rhodochrosite
MnCO3 +1.0000 H+ = + 1.0000 HCO3- + 1.0000 Mn++
log_k -0.1928
-delta_H -21.3426 kJ/mol # Calculated enthalpy of reaction Rhodochrosite
# Enthalpy of formation: -212.521 kcal/mol
-analytic -1.6195e+002 -4.9344e-002 5.0937e+003 6.4402e+001 7.9531e+001
# -Range: 0-300
Rhodonite
MnSiO3 +2.0000 H+ = + 1.0000 H2O + 1.0000 Mn++ + 1.0000 SiO2
log_k 9.7301
-delta_H -64.7121 kJ/mol # Calculated enthalpy of reaction Rhodonite
# Enthalpy of formation: -1319.42 kJ/mol
-analytic 2.0585e+001 4.9941e-003 4.5816e+003 -9.8212e+000 -3.0658e+005
# -Range: 0-300
Ripidolite-14A
Mg3Fe2Al2Si3O10(OH)8 +16.0000 H+ = + 2.0000 Al+++ + 2.0000 Fe++ + 3.0000 Mg++ + 3.0000 SiO2 + 12.0000 H2O
log_k 60.9638
-delta_H -572.472 kJ/mol # Calculated enthalpy of reaction Ripidolite-14A
# Enthalpy of formation: -1947.87 kcal/mol
-analytic -1.8376e+002 -6.1934e-002 3.2458e+004 6.2290e+001 5.0653e+002
# -Range: 0-300
Ripidolite-7A
Mg3Fe2Al2Si3O10(OH)8 +16.0000 H+ = + 2.0000 Al+++ + 2.0000 Fe++ + 3.0000 Mg++ + 3.0000 SiO2 + 12.0000 H2O
log_k 64.3371
-delta_H -586.325 kJ/mol # Calculated enthalpy of reaction Ripidolite-7A
# Enthalpy of formation: -1944.56 kcal/mol
-analytic -1.9557e+002 -6.3779e-002 3.3634e+004 6.7057e+001 5.2489e+002
# -Range: 0-300
Romarchite
SnO +2.0000 H+ = + 1.0000 H2O + 1.0000 Sn++
log_k 1.3625
-delta_H -8.69017 kJ/mol # Calculated enthalpy of reaction Romarchite
# Enthalpy of formation: -68.34 kcal/mol
-analytic -6.3187e+001 -1.5821e-002 2.2786e+003 2.4900e+001 3.5574e+001
# -Range: 0-300
Ru
Ru +2.0000 H+ +0.5000 O2 = + 1.0000 H2O + 1.0000 Ru++
log_k 16.6701
-delta_H 0 # Not possible to calculate enthalpy of reaction Ru
# Enthalpy of formation: 0 kJ/mol
Ru(OH)3:H2O(am)
Ru(OH)3:H2O +3.0000 H+ = + 1.0000 Ru+++ + 4.0000 H2O
log_k 1.6338
-delta_H 0 # Not possible to calculate enthalpy of reaction Ru(OH)3:H2O(am)
# Enthalpy of formation: 0 kcal/mol
RuBr3
RuBr3 = + 1.0000 Ru+++ + 3.0000 Br-
log_k 3.1479
-delta_H 0 # Not possible to calculate enthalpy of reaction RuBr3
# Enthalpy of formation: -147.76 kJ/mol
RuCl3
RuCl3 = + 1.0000 Ru+++ + 3.0000 Cl-
log_k 10.8215
-delta_H 0 # Not possible to calculate enthalpy of reaction RuCl3
# Enthalpy of formation: -221.291 kJ/mol
RuI3
RuI3 = + 1.0000 Ru+++ + 3.0000 I-
log_k -12.4614
-delta_H 0 # Not possible to calculate enthalpy of reaction RuI3
# Enthalpy of formation: -58.425 kJ/mol
RuO2
RuO2 +2.0000 H+ = + 1.0000 Ru(OH)2++
log_k -5.4835
-delta_H 0 # Not possible to calculate enthalpy of reaction RuO2
# Enthalpy of formation: -307.233 kJ/mol
RuO2:2H2O(am)
RuO2:2H2O +2.0000 H+ = + 1.0000 Ru(OH)2++ + 2.0000 H2O
log_k 0.9045
-delta_H 0 # Not possible to calculate enthalpy of reaction RuO2:2H2O(am)
# Enthalpy of formation: 0 kcal/mol
RuO4
RuO4 = + 1.0000 RuO4
log_k -0.9636
-delta_H 6.305 kJ/mol # Calculated enthalpy of reaction RuO4
# Enthalpy of formation: -244.447 kJ/mol
RuSe2
RuSe2 +2.0000 H2O = + 1.0000 Ru(OH)2++ + 2.0000 H+ + 2.0000 Se--
log_k -113.7236
-delta_H 0 # Not possible to calculate enthalpy of reaction RuSe2
# Enthalpy of formation: -146.274 kJ/mol
Rutherfordine
UO2CO3 +1.0000 H+ = + 1.0000 HCO3- + 1.0000 UO2++
log_k -4.1064
-delta_H -19.4032 kJ/mol # Calculated enthalpy of reaction Rutherfordine
# Enthalpy of formation: -1689.53 kJ/mol
-analytic -8.8224e+001 -3.1434e-002 2.6675e+003 3.4161e+001 4.1650e+001
# -Range: 0-300
Rutile
TiO2 +2.0000 H2O = + 1.0000 Ti(OH)4
log_k -9.6452
-delta_H 0 # Not possible to calculate enthalpy of reaction Rutile
# Enthalpy of formation: -226.107 kcal/mol
S
S +1.0000 H2O = + 0.5000 O2 + 1.0000 H+ + 1.0000 HS-
log_k -45.0980
-delta_H 263.663 kJ/mol # Calculated enthalpy of reaction S
# Enthalpy of formation: 0 kJ/mol
-analytic -8.8928e+001 -2.8454e-002 -1.1516e+004 3.6747e+001 -1.7966e+002
# -Range: 0-300
Safflorite
CoAs2 +2.0000 H2O +1.0000 H+ +0.5000 O2 = + 1.0000 AsH3 + 1.0000 Co++ + 1.0000 H2AsO3-
log_k -3.6419
-delta_H -52.7226 kJ/mol # Calculated enthalpy of reaction Safflorite
# Enthalpy of formation: -23.087 kcal/mol
Saleeite
Mg(UO2)2(PO4)2 +2.0000 H+ = + 1.0000 Mg++ + 2.0000 HPO4-- + 2.0000 UO2++
log_k -19.4575
-delta_H -110.816 kJ/mol # Calculated enthalpy of reaction Saleeite
# Enthalpy of formation: -1189.61 kcal/mol
-analytic -6.0028e+001 -4.4391e-002 3.9168e+003 1.6428e+001 6.6533e+001
# -Range: 0-200
Sanbornite
BaSi2O5 +2.0000 H+ = + 1.0000 Ba++ + 1.0000 H2O + 2.0000 SiO2
log_k 9.4753
-delta_H -31.0845 kJ/mol # Calculated enthalpy of reaction Sanbornite
# Enthalpy of formation: -2547.8 kJ/mol
-analytic -2.5381e+001 1.2999e-002 1.2330e+004 2.1053e+000 -1.3913e+006
# -Range: 0-300
Sanidine_high
KAlSi3O8 +4.0000 H+ = + 1.0000 Al+++ + 1.0000 K+ + 2.0000 H2O + 3.0000 SiO2
log_k 0.9239
-delta_H -35.0284 kJ/mol # Calculated enthalpy of reaction Sanidine_high
# Enthalpy of formation: -946.538 kcal/mol
-analytic -3.4889e+000 1.4495e-002 1.2856e+004 -9.8978e+000 -1.6572e+006
# -Range: 0-300
Saponite-Ca
Ca.165Mg3Al.33Si3.67O10(OH)2 +7.3200 H+ = + 0.1650 Ca++ + 0.3300 Al+++ + 3.0000 Mg++ + 3.6700 SiO2 + 4.6600 H2O
log_k 26.2900
-delta_H -207.971 kJ/mol # Calculated enthalpy of reaction Saponite-Ca
# Enthalpy of formation: -1436.51 kcal/mol
-analytic -4.6904e+001 6.2555e-003 2.2572e+004 5.3198e+000 -1.5725e+006
# -Range: 0-300
Saponite-Cs
Cs.33Si3.67Al.33Mg3O10(OH)2 +7.3200 H+ = + 0.3300 Al+++ + 0.3300 Cs+ + 3.0000 Mg++ + 3.6700 SiO2 + 4.6600 H2O
log_k 25.8528
-delta_H -195.407 kJ/mol # Calculated enthalpy of reaction Saponite-Cs
# Enthalpy of formation: -1438.44 kcal/mol
-analytic -7.7732e+001 -3.6418e-005 2.3346e+004 1.7578e+001 -1.6319e+006
# -Range: 0-300
Saponite-H
H.33Mg3Al.33Si3.67O10(OH)2 +6.9900 H+ = + 0.3300 Al+++ + 3.0000 Mg++ + 3.6700 SiO2 + 4.6600 H2O
log_k 25.3321
-delta_H -200.235 kJ/mol # Calculated enthalpy of reaction Saponite-H
# Enthalpy of formation: -1416.94 kcal/mol
-analytic -3.9828e+001 8.9566e-003 2.2165e+004 2.3941e+000 -1.5933e+006
# -Range: 0-300
Saponite-K
K.33Mg3Al.33Si3.67O10(OH)2 +7.3200 H+ = + 0.3300 Al+++ + 0.3300 K+ + 3.0000 Mg++ + 3.6700 SiO2 + 4.6600 H2O
log_k 26.0075
-delta_H -196.402 kJ/mol # Calculated enthalpy of reaction Saponite-K
# Enthalpy of formation: -1437.74 kcal/mol
-analytic 3.2113e+001 1.8392e-002 1.7918e+004 -2.2874e+001 -1.3542e+006
# -Range: 0-300
Saponite-Mg
Mg3.165Al.33Si3.67O10(OH)2 +7.3200 H+ = + 0.3300 Al+++ + 3.1650 Mg++ + 3.6700 SiO2 + 4.6600 H2O
log_k 26.2523
-delta_H -210.822 kJ/mol # Calculated enthalpy of reaction Saponite-Mg
# Enthalpy of formation: -1432.79 kcal/mol
-analytic 9.8888e+000 1.4320e-002 1.9418e+004 -1.5259e+001 -1.3716e+006
# -Range: 0-300
Saponite-Na
Na.33Mg3Al.33Si3.67O10(OH)2 +7.3200 H+ = + 0.3300 Al+++ + 0.3300 Na+ + 3.0000 Mg++ + 3.6700 SiO2 + 4.6600 H2O
log_k 26.3459
-delta_H -201.401 kJ/mol # Calculated enthalpy of reaction Saponite-Na
# Enthalpy of formation: -1435.61 kcal/mol
-analytic -6.7611e+001 4.7327e-003 2.3586e+004 1.2868e+001 -1.6493e+006
# -Range: 0-300
Sb
Sb +1.5000 H2O +0.7500 O2 = + 1.0000 Sb(OH)3
log_k 52.7918
-delta_H -335.931 kJ/mol # Calculated enthalpy of reaction Sb
# Enthalpy of formation: 0 kJ/mol
Sb(OH)3
Sb(OH)3 = + 1.0000 Sb(OH)3
log_k -7.0953
-delta_H 0 # Not possible to calculate enthalpy of reaction Sb(OH)3
# Enthalpy of formation: 0 kcal/mol
Sb2O3
Sb2O3 +3.0000 H2O = + 2.0000 Sb(OH)3
log_k -8.9600
-delta_H 0 # Not possible to calculate enthalpy of reaction Sb2O3
# Enthalpy of formation: 0 kcal/mol
-analytic 2.3982e+000 -7.6326e-005 -3.3787e+003 0.0000e+000 0.0000e+000
# -Range: 0-300
Sb2O4
Sb2O4 +3.0000 H2O = + 0.5000 O2 + 2.0000 Sb(OH)3
log_k -39.6139
-delta_H 211.121 kJ/mol # Calculated enthalpy of reaction Sb2O4
# Enthalpy of formation: -907.251 kJ/mol
Sb2O5
Sb2O5 +3.0000 H2O = + 1.0000 O2 + 2.0000 Sb(OH)3
log_k -46.9320
-delta_H 269.763 kJ/mol # Calculated enthalpy of reaction Sb2O5
# Enthalpy of formation: -971.96 kJ/mol
Sb4O6(cubic)
Sb4O6 +6.0000 H2O = + 4.0000 Sb(OH)3
log_k -19.6896
-delta_H 59.898 kJ/mol # Calculated enthalpy of reaction Sb4O6(cubic)
# Enthalpy of formation: -1440.02 kJ/mol
Sb4O6(orthorhombic)
Sb4O6 +6.0000 H2O = + 4.0000 Sb(OH)3
log_k -17.0442
-delta_H 37.314 kJ/mol # Calculated enthalpy of reaction Sb4O6(orthorhombic)
# Enthalpy of formation: -1417.44 kJ/mol
SbBr3
SbBr3 +3.0000 H2O = + 1.0000 Sb(OH)3 + 3.0000 Br- + 3.0000 H+
log_k 1.0554
-delta_H -21.5871 kJ/mol # Calculated enthalpy of reaction SbBr3
# Enthalpy of formation: -259.197 kJ/mol
SbCl3
SbCl3 +3.0000 H2O = + 1.0000 Sb(OH)3 + 3.0000 Cl- + 3.0000 H+
log_k 0.5878
-delta_H -35.393 kJ/mol # Calculated enthalpy of reaction SbCl3
# Enthalpy of formation: -382.12 kJ/mol
Sc
Sc +3.0000 H+ +0.7500 O2 = + 1.0000 Sc+++ + 1.5000 H2O
log_k 167.2700
-delta_H -1033.87 kJ/mol # Calculated enthalpy of reaction Sc
# Enthalpy of formation: 0 kJ/mol
-analytic -6.6922e+001 -2.9150e-002 5.4559e+004 2.4189e+001 8.5137e+002
# -Range: 0-300
Scacchite
MnCl2 = + 1.0000 Mn++ + 2.0000 Cl-
log_k 8.7785
-delta_H -73.4546 kJ/mol # Calculated enthalpy of reaction Scacchite
# Enthalpy of formation: -481.302 kJ/mol
-analytic -2.3476e+002 -8.2437e-002 9.0088e+003 9.6128e+001 1.4064e+002
# -Range: 0-300
Schoepite
UO3:2H2O +2.0000 H+ = + 1.0000 UO2++ + 3.0000 H2O
log_k 4.8333
-delta_H -50.415 kJ/mol # Calculated enthalpy of reaction Schoepite
# Enthalpy of formation: -1826.1 kJ/mol
-analytic 1.3645e+001 1.0884e-002 2.5412e+003 -8.3167e+000 3.9649e+001
# -Range: 0-300
Schoepite-dehy(.393)
UO3:.393H2O +2.0000 H+ = + 1.0000 UO2++ + 1.3930 H2O
log_k 6.7243
-delta_H -69.2728 kJ/mol # Calculated enthalpy of reaction Schoepite-dehy(.393)
# Enthalpy of formation: -1347.9 kJ/mol
-analytic -5.6487e+001 -3.0358e-003 5.7044e+003 1.8179e+001 9.6887e+001
# -Range: 0-200
Schoepite-dehy(.648)
UO3:.648H2O +2.0000 H+ = + 1.0000 UO2++ + 1.6480 H2O
log_k 6.2063
-delta_H -65.4616 kJ/mol # Calculated enthalpy of reaction Schoepite-dehy(.648)
# Enthalpy of formation: -1424.6 kJ/mol
-analytic -6.3010e+001 -3.0276e-003 5.8033e+003 2.0471e+001 9.8569e+001
# -Range: 0-200
Schoepite-dehy(.85)
UO3:.85H2O +2.0000 H+ = + 1.0000 UO2++ + 1.8500 H2O
log_k 5.0970
-delta_H -56.4009 kJ/mol # Calculated enthalpy of reaction Schoepite-dehy(.85)
# Enthalpy of formation: -1491.4 kJ/mol
-analytic -6.7912e+001 -3.0420e-003 5.5690e+003 2.2323e+001 9.4593e+001
# -Range: 0-200
Schoepite-dehy(.9)
UO3:.9H2O +2.0000 H+ = + 1.0000 UO2++ + 1.9000 H2O
log_k 5.0167
-delta_H -55.7928 kJ/mol # Calculated enthalpy of reaction Schoepite-dehy(.9)
# Enthalpy of formation: -1506.3 kJ/mol
-analytic -1.5998e+001 -2.0144e-003 3.2910e+003 4.2751e+000 5.1358e+001
# -Range: 0-300
Schoepite-dehy(1.0)
UO3:H2O +2.0000 H+ = + 1.0000 UO2++ + 2.0000 H2O
log_k 5.1031
-delta_H -57.4767 kJ/mol # Calculated enthalpy of reaction Schoepite-dehy(1.0)
# Enthalpy of formation: -1533.2 kJ/mol
-analytic -7.2080e+001 -3.0503e-003 5.8024e+003 2.3695e+001 9.8557e+001
# -Range: 0-200
Scolecite
CaAl2Si3O10:3H2O +8.0000 H+ = + 1.0000 Ca++ + 2.0000 Al+++ + 3.0000 SiO2 + 7.0000 H2O
log_k 15.8767
-delta_H -204.93 kJ/mol # Calculated enthalpy of reaction Scolecite
# Enthalpy of formation: -6048.92 kJ/mol
-analytic 5.0656e+001 -3.1485e-003 1.0574e+004 -2.5663e+001 -5.2769e+005
# -Range: 0-300
Se
Se +1.0000 H2O +1.0000 O2 = + 1.0000 SeO3-- + 2.0000 H+
log_k 26.1436
-delta_H -211.221 kJ/mol # Calculated enthalpy of reaction Se
# Enthalpy of formation: 0 kJ/mol
-analytic -9.5144e+001 -6.5681e-002 1.0736e+004 4.2358e+001 1.6755e+002
# -Range: 0-300
Se2O5
Se2O5 +2.0000 H2O = + 1.0000 SeO3-- + 1.0000 SeO4-- + 4.0000 H+
log_k 9.5047
-delta_H -123.286 kJ/mol # Calculated enthalpy of reaction Se2O5
# Enthalpy of formation: -98.8 kcal/mol
-analytic 1.1013e+002 -2.4491e-002 -5.6147e+002 -3.6960e+001 -9.5719e+000
# -Range: 0-200
SeCl4
SeCl4 +3.0000 H2O = + 1.0000 SeO3-- + 4.0000 Cl- + 6.0000 H+
log_k 14.4361
-delta_H -131.298 kJ/mol # Calculated enthalpy of reaction SeCl4
# Enthalpy of formation: -45.1 kcal/mol
-analytic -4.0215e+002 -1.8323e-001 1.3074e+004 1.7267e+002 2.0413e+002
# -Range: 0-300
SeO3
SeO3 +1.0000 H2O = + 1.0000 SeO4-- + 2.0000 H+
log_k 19.2015
-delta_H -143.022 kJ/mol # Calculated enthalpy of reaction SeO3
# Enthalpy of formation: -40.7 kcal/mol
-analytic -1.4199e+002 -6.4398e-002 9.5505e+003 5.9941e+001 1.4907e+002
# -Range: 0-300
Sellaite
MgF2 = + 1.0000 Mg++ + 2.0000 F-
log_k -9.3843
-delta_H -12.4547 kJ/mol # Calculated enthalpy of reaction Sellaite
# Enthalpy of formation: -1124.2 kJ/mol
-analytic -2.6901e+002 -8.5487e-002 6.8237e+003 1.0595e+002 1.0656e+002
# -Range: 0-300
Sepiolite
Mg4Si6O15(OH)2:6H2O +8.0000 H+ = + 4.0000 Mg++ + 6.0000 SiO2 + 11.0000 H2O
log_k 30.4439
-delta_H -157.339 kJ/mol # Calculated enthalpy of reaction Sepiolite
# Enthalpy of formation: -2418 kcal/mol
-analytic 1.8690e+001 4.7544e-002 2.6765e+004 -2.5301e+001 -2.6498e+006
# -Range: 0-300
Shcherbinaite
V2O5 +2.0000 H+ = + 1.0000 H2O + 2.0000 VO2+
log_k -1.4520
-delta_H -34.7917 kJ/mol # Calculated enthalpy of reaction Shcherbinaite
# Enthalpy of formation: -1550.6 kJ/mol
-analytic -1.4791e+002 -2.2464e-002 6.6865e+003 5.2832e+001 1.0438e+002
# -Range: 0-300
Si
Si +1.0000 O2 = + 1.0000 SiO2
log_k 148.9059
-delta_H -865.565 kJ/mol # Calculated enthalpy of reaction Si
# Enthalpy of formation: 0 kJ/mol
-analytic -5.7245e+002 -7.6302e-002 8.3516e+004 2.0045e+002 -2.8494e+006
# -Range: 0-300
SiO2(am)
SiO2 = + 1.0000 SiO2
log_k -2.7136
-delta_H 20.0539 kJ/mol # Calculated enthalpy of reaction SiO2(am)
# Enthalpy of formation: -214.568 kcal/mol
-analytic 1.2109e+000 7.0767e-003 2.3634e+003 -3.4449e+000 -4.8591e+005
# -Range: 0-300
Siderite
FeCO3 +1.0000 H+ = + 1.0000 Fe++ + 1.0000 HCO3-
log_k -0.1920
-delta_H -32.5306 kJ/mol # Calculated enthalpy of reaction Siderite
# Enthalpy of formation: -179.173 kcal/mol
-analytic -1.5990e+002 -4.9361e-002 5.4947e+003 6.3032e+001 8.5787e+001
# -Range: 0-300
Sillimanite
Al2SiO5 +6.0000 H+ = + 1.0000 SiO2 + 2.0000 Al+++ + 3.0000 H2O
log_k 16.3080
-delta_H -238.442 kJ/mol # Calculated enthalpy of reaction Sillimanite
# Enthalpy of formation: -615.099 kcal/mol
-analytic -7.1610e+001 -3.2196e-002 1.2493e+004 2.2449e+001 1.9496e+002
# -Range: 0-300
Sklodowskite
Mg(H3O)2(UO2)2(SiO4)2:4H2O +6.0000 H+ = + 1.0000 Mg++ + 2.0000 SiO2 + 2.0000 UO2++ + 10.0000 H2O
log_k 13.7915
-delta_H 0 # Not possible to calculate enthalpy of reaction Sklodowskite
# Enthalpy of formation: 0 kcal/mol
Sm
Sm +2.0000 H+ +0.5000 O2 = + 1.0000 H2O + 1.0000 Sm++
log_k 133.1614
-delta_H -783.944 kJ/mol # Calculated enthalpy of reaction Sm
# Enthalpy of formation: 0 kJ/mol
-analytic -7.1599e+001 -2.0083e-002 4.2693e+004 2.7291e+001 6.6621e+002
# -Range: 0-300
Sm(OH)3
Sm(OH)3 +3.0000 H+ = + 1.0000 Sm+++ + 3.0000 H2O
log_k 16.4852
-delta_H 0 # Not possible to calculate enthalpy of reaction Sm(OH)3
# Enthalpy of formation: 0 kcal/mol
Sm(OH)3(am)
Sm(OH)3 +3.0000 H+ = + 1.0000 Sm+++ + 3.0000 H2O
log_k 18.5852
-delta_H 0 # Not possible to calculate enthalpy of reaction Sm(OH)3(am)
# Enthalpy of formation: 0 kcal/mol
Sm2(CO3)3
Sm2(CO3)3 +3.0000 H+ = + 2.0000 Sm+++ + 3.0000 HCO3-
log_k -3.5136
-delta_H 0 # Not possible to calculate enthalpy of reaction Sm2(CO3)3
# Enthalpy of formation: 0 kcal/mol
Sm2(SO4)3
Sm2(SO4)3 = + 2.0000 Sm+++ + 3.0000 SO4--
log_k -9.8000
-delta_H 0 # Not possible to calculate enthalpy of reaction Sm2(SO4)3
# Enthalpy of formation: 0 kcal/mol
Sm2O3
Sm2O3 +6.0000 H+ = + 2.0000 Sm+++ + 3.0000 H2O
log_k 42.9000
-delta_H 0 # Not possible to calculate enthalpy of reaction Sm2O3
# Enthalpy of formation: 0 kcal/mol
SmF3:.5H2O
SmF3:.5H2O = + 0.5000 H2O + 1.0000 Sm+++ + 3.0000 F-
log_k -17.5000
-delta_H 0 # Not possible to calculate enthalpy of reaction SmF3:.5H2O
# Enthalpy of formation: 0 kcal/mol
SmPO4:10H2O
SmPO4:10H2O +1.0000 H+ = + 1.0000 HPO4-- + 1.0000 Sm+++ + 10.0000 H2O
log_k -12.1782
-delta_H 0 # Not possible to calculate enthalpy of reaction SmPO4:10H2O
# Enthalpy of formation: 0 kcal/mol
Smectite-high-Fe-Mg
# Ca.025Na.1K.2Fe++.5Fe+++.2Mg1.15Al1.25Si3.5H2O12 +8.0000 H+ = + 0.0250 Ca++ + 0.1000 Na+ + 0.2000 Fe+++ + 0.2000 K+ + 0.5000 Fe++ + 1.1500 Mg++ + 1.2500 Al+++ + 3.5000 SiO2 + 5.0000 H2O
Ca.025Na.1K.2Fe.5Fe.2Mg1.15Al1.25Si3.5H2O12 +8.0000 H+ = + 0.0250 Ca++ + 0.1000 Na+ + 0.2000 Fe+++ + 0.2000 K+ + 0.5000 Fe++ + 1.1500 Mg++ + 1.2500 Al+++ + 3.5000 SiO2 + 5.0000 H2O
log_k 17.4200
-delta_H -199.841 kJ/mol # Calculated enthalpy of reaction Smectite-high-Fe-Mg
# Enthalpy of formation: -1351.39 kcal/mol
-analytic -9.6102e+000 1.2551e-003 1.8157e+004 -7.9862e+000 -1.3005e+006
# -Range: 0-300
Smectite-low-Fe-Mg
# Ca.02Na.15K.2Fe++.29Fe+++.16Mg.9Al1.25Si3.75H2O1 +7.0000 H+ = + 0.0200 Ca++ + 0.1500 Na+ + 0.1600 Fe+++ + 0.2000 K+ + 0.2900 Fe++ + 0.9000 Mg++ + 1.2500 Al+++ + 3.7500 SiO2 + 4.5000 H2O
Ca.02Na.15K.2Fe.29Fe.16Mg.9Al1.25Si3.75H2O12 +7.0000 H+ = + 0.0200 Ca++ + 0.1500 Na+ + 0.1600 Fe+++ + 0.2000 K+ + 0.2900 Fe++ + 0.9000 Mg++ + 1.2500 Al+++ + 3.7500 SiO2 + 4.5000 H2O
log_k 11.0405
-delta_H -144.774 kJ/mol # Calculated enthalpy of reaction Smectite-low-Fe-Mg
# Enthalpy of formation: -1352.12 kcal/mol
-analytic -1.7003e+001 6.9848e-003 1.8359e+004 -6.8896e+000 -1.6637e+006
# -Range: 0-300
Smithsonite
ZnCO3 +1.0000 H+ = + 1.0000 HCO3- + 1.0000 Zn++
log_k 0.4633
-delta_H -30.5348 kJ/mol # Calculated enthalpy of reaction Smithsonite
# Enthalpy of formation: -194.26 kcal/mol
-analytic -1.6452e+002 -5.0231e-002 5.5925e+003 6.5139e+001 8.7314e+001
# -Range: 0-300
Sn
Sn +2.0000 H+ +0.5000 O2 = + 1.0000 H2O + 1.0000 Sn++
log_k 47.8615
-delta_H -288.558 kJ/mol # Calculated enthalpy of reaction Sn
# Enthalpy of formation: 0 kcal/mol
-analytic -1.3075e+002 -3.3807e-002 1.9548e+004 5.0382e+001 -1.3868e+005
# -Range: 0-300
Sn(OH)2
Sn(OH)2 +2.0000 H+ = + 1.0000 Sn++ + 2.0000 H2O
log_k 1.8400
-delta_H -19.6891 kJ/mol # Calculated enthalpy of reaction Sn(OH)2
# Enthalpy of formation: -560.774 kJ/mol
-analytic -6.1677e+001 -5.3258e-003 3.3656e+003 2.1748e+001 5.7174e+001
# -Range: 0-200
Sn(SO4)2
Sn(SO4)2 = + 1.0000 Sn++++ + 2.0000 SO4--
log_k 16.0365
-delta_H -159.707 kJ/mol # Calculated enthalpy of reaction Sn(SO4)2
# Enthalpy of formation: -389.4 kcal/mol
-analytic 1.7787e+001 -5.1758e-002 3.7671e+003 4.1861e-001 6.3965e+001
# -Range: 0-200
Sn3S4
Sn3S4 +4.0000 H+ = + 1.0000 Sn++++ + 2.0000 Sn++ + 4.0000 HS-
log_k -61.9790
-delta_H 318.524 kJ/mol # Calculated enthalpy of reaction Sn3S4
# Enthalpy of formation: -88.5 kcal/mol
-analytic -8.1325e+001 -7.4589e-002 -1.7953e+004 4.1138e+001 -3.0484e+002
# -Range: 0-200
SnBr2
SnBr2 = + 1.0000 Sn++ + 2.0000 Br-
log_k -1.4369
-delta_H 8.24248 kJ/mol # Calculated enthalpy of reaction SnBr2
# Enthalpy of formation: -62.15 kcal/mol
-analytic 2.5384e+001 -1.7350e-002 -2.6653e+003 -5.1400e+000 -4.5269e+001
# -Range: 0-200
SnBr4
SnBr4 = + 1.0000 Sn++++ + 4.0000 Br-
log_k 11.1272
-delta_H -78.3763 kJ/mol # Calculated enthalpy of reaction SnBr4
# Enthalpy of formation: -377.391 kJ/mol
-analytic 1.3516e+001 -5.5193e-002 -8.1888e+001 5.7935e+000 -1.3940e+000
# -Range: 0-200
SnCl2
SnCl2 = + 1.0000 Sn++ + 2.0000 Cl-
log_k 0.3225
-delta_H -11.9913 kJ/mol # Calculated enthalpy of reaction SnCl2
# Enthalpy of formation: -79.1 kcal/mol
-analytic 7.9717e+000 -2.1475e-002 -1.1676e+003 1.0749e+000 -1.9829e+001
# -Range: 0-200
SnSO4
SnSO4 = + 1.0000 SO4-- + 1.0000 Sn++
log_k -23.9293
-delta_H 96.232 kJ/mol # Calculated enthalpy of reaction SnSO4
# Enthalpy of formation: -242.5 kcal/mol
-analytic 3.0046e+001 -1.4238e-002 -7.5915e+003 -9.8122e+000 -1.2892e+002
# -Range: 0-200
SnSe
SnSe = + 1.0000 Se-- + 1.0000 Sn++
log_k -32.9506
-delta_H 0 # Not possible to calculate enthalpy of reaction SnSe
# Enthalpy of formation: -21.2 kcal/mol
-analytic 4.2342e+000 9.5462e-004 -8.0009e+003 -4.2997e+000 -1.3587e+002
# -Range: 0-200
SnSe2
SnSe2 = + 1.0000 Sn++++ + 2.0000 Se--
log_k -66.6570
-delta_H 0 # Not possible to calculate enthalpy of reaction SnSe2
# Enthalpy of formation: -29.8 kcal/mol
-analytic -3.6819e+001 -2.0966e-002 -1.5197e+004 1.1070e+001 -2.5806e+002
# -Range: 0-200
Soddyite
(UO2)2SiO4:2H2O +4.0000 H+ = + 1.0000 SiO2 + 2.0000 UO2++ + 4.0000 H2O
log_k 0.3920
-delta_H 0 # Not possible to calculate enthalpy of reaction Soddyite
# Enthalpy of formation: 0 kcal/mol
Sphaerocobaltite
CoCO3 +1.0000 H+ = + 1.0000 Co++ + 1.0000 HCO3-
log_k -0.2331
-delta_H -30.7064 kJ/mol # Calculated enthalpy of reaction Sphaerocobaltite
# Enthalpy of formation: -171.459 kcal/mol
-analytic -1.5709e+002 -4.8957e-002 5.3158e+003 6.2075e+001 8.2995e+001
# -Range: 0-300
Sphalerite
ZnS +1.0000 H+ = + 1.0000 HS- + 1.0000 Zn++
log_k -11.4400
-delta_H 35.5222 kJ/mol # Calculated enthalpy of reaction Sphalerite
# Enthalpy of formation: -49 kcal/mol
-analytic -1.5497e+002 -4.8953e-002 1.7850e+003 6.1472e+001 2.7899e+001
# -Range: 0-300
Spinel
Al2MgO4 +8.0000 H+ = + 1.0000 Mg++ + 2.0000 Al+++ + 4.0000 H2O
log_k 37.6295
-delta_H -398.108 kJ/mol # Calculated enthalpy of reaction Spinel
# Enthalpy of formation: -546.847 kcal/mol
-analytic -3.3895e+002 -8.3595e-002 2.9251e+004 1.2260e+002 4.5654e+002
# -Range: 0-300
Spinel-Co
Co3O4 +8.0000 H+ = + 1.0000 Co++ + 2.0000 Co+++ + 4.0000 H2O
log_k -6.4852
-delta_H -126.415 kJ/mol # Calculated enthalpy of reaction Spinel-Co
# Enthalpy of formation: -891 kJ/mol
-analytic -3.2239e+002 -8.0782e-002 1.4635e+004 1.1755e+002 2.2846e+002
# -Range: 0-300
Spodumene
LiAlSi2O6 +4.0000 H+ = + 1.0000 Al+++ + 1.0000 Li+ + 2.0000 H2O + 2.0000 SiO2
log_k 6.9972
-delta_H -89.1817 kJ/mol # Calculated enthalpy of reaction Spodumene
# Enthalpy of formation: -3054.75 kJ/mol
-analytic -9.8111e+000 2.1191e-003 9.6920e+003 -3.0484e+000 -7.8822e+005
# -Range: 0-300
Sr
Sr +2.0000 H+ +0.5000 O2 = + 1.0000 H2O + 1.0000 Sr++
log_k 141.7816
-delta_H -830.679 kJ/mol # Calculated enthalpy of reaction Sr
# Enthalpy of formation: 0 kJ/mol
-analytic -1.6271e+002 -3.1212e-002 5.1520e+004 5.9178e+001 -4.8390e+005
# -Range: 0-300
Sr(NO3)2
Sr(NO3)2 = + 1.0000 Sr++ + 2.0000 NO3-
log_k 1.1493
-delta_H 13.7818 kJ/mol # Calculated enthalpy of reaction Sr(NO3)2
# Enthalpy of formation: -978.311 kJ/mol
-analytic 2.8914e+000 -1.2487e-002 -1.4872e+003 2.8124e+000 -2.5256e+001
# -Range: 0-200
Sr(NO3)2:4H2O
Sr(NO3)2:4H2O = + 1.0000 Sr++ + 2.0000 NO3- + 4.0000 H2O
log_k 0.6976
-delta_H 47.9045 kJ/mol # Calculated enthalpy of reaction Sr(NO3)2:4H2O
# Enthalpy of formation: -2155.79 kJ/mol
-analytic -8.4518e+001 -9.1155e-003 1.0856e+003 3.4061e+001 1.8464e+001
# -Range: 0-200
Sr(OH)2
Sr(OH)2 +2.0000 H+ = + 1.0000 Sr++ + 2.0000 H2O
log_k 27.5229
-delta_H -153.692 kJ/mol # Calculated enthalpy of reaction Sr(OH)2
# Enthalpy of formation: -968.892 kJ/mol
-analytic -5.1871e+001 -2.9123e-003 1.0175e+004 1.8643e+001 1.7280e+002
# -Range: 0-200
Sr2SiO4
Sr2SiO4 +4.0000 H+ = + 1.0000 SiO2 + 2.0000 H2O + 2.0000 Sr++
log_k 42.8076
-delta_H -244.583 kJ/mol # Calculated enthalpy of reaction Sr2SiO4
# Enthalpy of formation: -2306.61 kJ/mol
-analytic 3.0319e+001 2.0204e-003 1.2729e+004 -1.1584e+001 -1.9480e+005
# -Range: 0-300
Sr3(AsO4)2
Sr3(AsO4)2 +4.0000 H+ = + 2.0000 H2AsO4- + 3.0000 Sr++
log_k 20.6256
-delta_H -152.354 kJ/mol # Calculated enthalpy of reaction Sr3(AsO4)2
# Enthalpy of formation: -3319.49 kJ/mol
-analytic -8.4749e+001 -2.9367e-002 9.5849e+003 3.3126e+001 1.6279e+002
# -Range: 0-200
SrBr2
SrBr2 = + 1.0000 Sr++ + 2.0000 Br-
log_k 13.1128
-delta_H -75.106 kJ/mol # Calculated enthalpy of reaction SrBr2
# Enthalpy of formation: -718.808 kJ/mol
-analytic -1.8512e+002 -7.2423e-002 7.6861e+003 7.8401e+001 1.1999e+002
# -Range: 0-300
SrBr2:6H2O
SrBr2:6H2O = + 1.0000 Sr++ + 2.0000 Br- + 6.0000 H2O
log_k 3.6678
-delta_H 23.367 kJ/mol # Calculated enthalpy of reaction SrBr2:6H2O
# Enthalpy of formation: -2532.31 kJ/mol
-analytic -2.2470e+002 -6.7920e-002 4.9432e+003 9.3758e+001 7.7200e+001
# -Range: 0-300
SrBr2:H2O
SrBr2:H2O = + 1.0000 H2O + 1.0000 Sr++ + 2.0000 Br-
log_k 9.6057
-delta_H -47.5853 kJ/mol # Calculated enthalpy of reaction SrBr2:H2O
# Enthalpy of formation: -1032.17 kJ/mol
-analytic -1.9103e+002 -7.1402e-002 6.6358e+003 8.0673e+001 1.0360e+002
# -Range: 0-300
SrCl2
SrCl2 = + 1.0000 Sr++ + 2.0000 Cl-
log_k 7.9389
-delta_H -55.0906 kJ/mol # Calculated enthalpy of reaction SrCl2
# Enthalpy of formation: -829.976 kJ/mol
-analytic -2.0097e+002 -7.6193e-002 7.0396e+003 8.4050e+001 1.0991e+002
# -Range: 0-300
SrCl2:2H2O
SrCl2:2H2O = + 1.0000 Sr++ + 2.0000 Cl- + 2.0000 H2O
log_k 3.3248
-delta_H -17.7313 kJ/mol # Calculated enthalpy of reaction SrCl2:2H2O
# Enthalpy of formation: -1439.01 kJ/mol
-analytic -2.1551e+002 -7.4349e-002 5.9400e+003 8.9330e+001 9.2752e+001
# -Range: 0-300
SrCl2:6H2O
SrCl2:6H2O = + 1.0000 Sr++ + 2.0000 Cl- + 6.0000 H2O
log_k 1.5038
-delta_H 24.6964 kJ/mol # Calculated enthalpy of reaction SrCl2:6H2O
# Enthalpy of formation: -2624.79 kJ/mol
-analytic -1.3225e+002 -1.8260e-002 3.7077e+003 5.1224e+001 6.3008e+001
# -Range: 0-200
SrCl2:H2O
SrCl2:H2O = + 1.0000 H2O + 1.0000 Sr++ + 2.0000 Cl-
log_k 4.7822
-delta_H -33.223 kJ/mol # Calculated enthalpy of reaction SrCl2:H2O
# Enthalpy of formation: -1137.68 kJ/mol
-analytic -2.1825e+002 -7.7851e-002 6.5957e+003 9.0555e+001 1.0298e+002
# -Range: 0-300
SrCrO4
SrCrO4 = + 1.0000 CrO4-- + 1.0000 Sr++
log_k -3.8849
-delta_H -1.73636 kJ/mol # Calculated enthalpy of reaction SrCrO4
# Enthalpy of formation: -341.855 kcal/mol
-analytic 2.3424e+001 -1.5589e-002 -2.1393e+003 -6.2628e+000 -3.6337e+001
# -Range: 0-200
SrF2
SrF2 = + 1.0000 Sr++ + 2.0000 F-
log_k -8.5400
-delta_H 0 # Not possible to calculate enthalpy of reaction SrF2
# Enthalpy of formation: 0 kcal/mol
SrHPO4
SrHPO4 = + 1.0000 HPO4-- + 1.0000 Sr++
log_k -6.2416
-delta_H -19.7942 kJ/mol # Calculated enthalpy of reaction SrHPO4
# Enthalpy of formation: -1823.19 kJ/mol
-analytic 5.4057e+000 -1.8533e-002 -8.2021e+002 -1.3667e+000 -1.3930e+001
# -Range: 0-200
SrI2
SrI2 = + 1.0000 Sr++ + 2.0000 I-
log_k 19.2678
-delta_H -103.218 kJ/mol # Calculated enthalpy of reaction SrI2
# Enthalpy of formation: -561.494 kJ/mol
-analytic -1.8168e+002 -7.2083e-002 9.0759e+003 7.7577e+001 1.4167e+002
# -Range: 0-300
SrO
SrO +2.0000 H+ = + 1.0000 H2O + 1.0000 Sr++
log_k 41.8916
-delta_H -243.875 kJ/mol # Calculated enthalpy of reaction SrO
# Enthalpy of formation: -592.871 kJ/mol
-analytic -5.8463e+001 -1.4240e-002 1.4417e+004 2.2725e+001 2.2499e+002
# -Range: 0-300
SrS
SrS +1.0000 H+ = + 1.0000 HS- + 1.0000 Sr++
log_k 14.7284
-delta_H -93.3857 kJ/mol # Calculated enthalpy of reaction SrS
# Enthalpy of formation: -473.63 kJ/mol
-analytic -1.3048e+002 -4.4837e-002 7.8429e+003 5.3442e+001 1.2242e+002
# -Range: 0-300
SrSeO4
SrSeO4 = + 1.0000 SeO4-- + 1.0000 Sr++
log_k -4.4000
-delta_H 0 # Not possible to calculate enthalpy of reaction SrSeO4
# Enthalpy of formation: 0 kcal/mol
SrSiO3
SrSiO3 +2.0000 H+ = + 1.0000 H2O + 1.0000 SiO2 + 1.0000 Sr++
log_k 14.8438
-delta_H -79.6112 kJ/mol # Calculated enthalpy of reaction SrSiO3
# Enthalpy of formation: -1634.83 kJ/mol
-analytic 2.2592e+001 6.0821e-003 5.9982e+003 -1.0213e+001 -3.9529e+005
# -Range: 0-300
SrUO4(alpha)
SrUO4 +4.0000 H+ = + 1.0000 Sr++ + 1.0000 UO2++ + 2.0000 H2O
log_k 19.1650
-delta_H -151.984 kJ/mol # Calculated enthalpy of reaction SrUO4(alpha)
# Enthalpy of formation: -1989.6 kJ/mol
-analytic -7.4169e+001 -1.6686e-002 9.8721e+003 2.6345e+001 1.5407e+002
# -Range: 0-300
SrZrO3
SrZrO3 +4.0000 H+ = + 1.0000 H2O + 1.0000 Sr++ + 1.0000 Zr(OH)2++
log_k -131.4664
-delta_H 706.983 kJ/mol # Calculated enthalpy of reaction SrZrO3
# Enthalpy of formation: -629.677 kcal/mol
-analytic -5.8512e+001 -9.5738e-003 -3.5254e+004 1.9459e+001 -5.9865e+002
# -Range: 0-200
Starkeyite
MgSO4:4H2O = + 1.0000 Mg++ + 1.0000 SO4-- + 4.0000 H2O
log_k -0.9999
-delta_H 0 # Not possible to calculate enthalpy of reaction Starkeyite
# Enthalpy of formation: 0 kcal/mol
Stibnite
Sb2S3 +6.0000 H2O = + 2.0000 Sb(OH)3 + 3.0000 H+ + 3.0000 HS-
log_k -53.1100
-delta_H 0 # Not possible to calculate enthalpy of reaction Stibnite
# Enthalpy of formation: 0 kcal/mol
-analytic 2.5223e+001 -5.9186e-002 -2.0860e+004 3.6892e+000 -3.2551e+002
# -Range: 0-300
Stilbite
Ca1.019Na.136K.006Al2.18Si6.82O18:7.33H2O +8.7200 H+ = + 0.0060 K+ + 0.1360 Na+ + 1.0190 Ca++ + 2.1800 Al+++ + 6.8200 SiO2 + 11.6900 H2O
log_k 1.0545
-delta_H -83.0019 kJ/mol # Calculated enthalpy of reaction Stilbite
# Enthalpy of formation: -11005.7 kJ/mol
-analytic -2.4483e+001 3.0987e-002 2.8013e+004 -1.5802e+001 -3.4491e+006
# -Range: 0-300
Stilleite
ZnSe = + 1.0000 Se-- + 1.0000 Zn++
log_k -23.9693
-delta_H 0 # Not possible to calculate enthalpy of reaction Stilleite
# Enthalpy of formation: -37.97 kcal/mol
-analytic -6.1948e+001 -1.7004e-002 -2.4498e+003 2.0712e+001 -3.8209e+001
# -Range: 0-300
Strengite
FePO4:2H2O +1.0000 H+ = + 1.0000 Fe+++ + 1.0000 HPO4-- + 2.0000 H2O
log_k -11.3429
-delta_H -37.107 kJ/mol # Calculated enthalpy of reaction Strengite
# Enthalpy of formation: -1876.23 kJ/mol
-analytic -2.7752e+002 -9.4014e-002 7.6862e+003 1.0846e+002 1.2002e+002
# -Range: 0-300
Strontianite
SrCO3 +1.0000 H+ = + 1.0000 HCO3- + 1.0000 Sr++
log_k -0.3137
-delta_H -8.23411 kJ/mol # Calculated enthalpy of reaction Strontianite
# Enthalpy of formation: -294.6 kcal/mol
-analytic -1.3577e+002 -4.4884e-002 3.5729e+003 5.5296e+001 5.5791e+001
# -Range: 0-300
Sylvite
KCl = + 1.0000 Cl- + 1.0000 K+
log_k 0.8459
-delta_H 17.4347 kJ/mol # Calculated enthalpy of reaction Sylvite
# Enthalpy of formation: -104.37 kcal/mol
-analytic -8.1204e+001 -3.3074e-002 8.2819e+002 3.6014e+001 1.2947e+001
# -Range: 0-300
Syngenite
K2Ca(SO4)2:H2O = + 1.0000 Ca++ + 1.0000 H2O + 2.0000 K+ + 2.0000 SO4--
log_k -7.6001
-delta_H 0 # Not possible to calculate enthalpy of reaction Syngenite
# Enthalpy of formation: 0 kcal/mol
Tachyhydrite
Mg2CaCl6:12H2O = + 1.0000 Ca++ + 2.0000 Mg++ + 6.0000 Cl- + 12.0000 H2O
log_k 17.1439
-delta_H 0 # Not possible to calculate enthalpy of reaction Tachyhydrite
# Enthalpy of formation: 0 kcal/mol
Talc
Mg3Si4O10(OH)2 +6.0000 H+ = + 3.0000 Mg++ + 4.0000 H2O + 4.0000 SiO2
log_k 21.1383
-delta_H -148.737 kJ/mol # Calculated enthalpy of reaction Talc
# Enthalpy of formation: -1410.92 kcal/mol
-analytic 1.1164e+001 2.4724e-002 1.9810e+004 -1.7568e+001 -1.8241e+006
# -Range: 0-300
Tarapacaite
K2CrO4 = + 1.0000 CrO4-- + 2.0000 K+
log_k -0.4037
-delta_H 17.8238 kJ/mol # Calculated enthalpy of reaction Tarapacaite
# Enthalpy of formation: -335.4 kcal/mol
-analytic 2.7953e+001 -1.0863e-002 -2.7589e+003 -6.4154e+000 -4.6859e+001
# -Range: 0-200
Tb
Tb +3.0000 H+ +0.7500 O2 = + 1.0000 Tb+++ + 1.5000 H2O
log_k 181.4170
-delta_H -1117.97 kJ/mol # Calculated enthalpy of reaction Tb
# Enthalpy of formation: 0 kJ/mol
-analytic -5.2354e+001 -2.6920e-002 5.8391e+004 1.8555e+001 9.1115e+002
# -Range: 0-300
Tb(OH)3
Tb(OH)3 +3.0000 H+ = + 1.0000 Tb+++ + 3.0000 H2O
log_k 15.6852
-delta_H 0 # Not possible to calculate enthalpy of reaction Tb(OH)3
# Enthalpy of formation: 0 kcal/mol
Tb(OH)3(am)
Tb(OH)3 +3.0000 H+ = + 1.0000 Tb+++ + 3.0000 H2O
log_k 18.7852
-delta_H 0 # Not possible to calculate enthalpy of reaction Tb(OH)3(am)
# Enthalpy of formation: 0 kcal/mol
Tb2(CO3)3
Tb2(CO3)3 +3.0000 H+ = + 2.0000 Tb+++ + 3.0000 HCO3-
log_k -3.2136
-delta_H 0 # Not possible to calculate enthalpy of reaction Tb2(CO3)3
# Enthalpy of formation: 0 kcal/mol
Tb2O3
Tb2O3 +6.0000 H+ = + 2.0000 Tb+++ + 3.0000 H2O
log_k 47.1000
-delta_H 0 # Not possible to calculate enthalpy of reaction Tb2O3
# Enthalpy of formation: 0 kcal/mol
TbF3:.5H2O
TbF3:.5H2O = + 0.5000 H2O + 1.0000 Tb+++ + 3.0000 F-
log_k -16.7000
-delta_H 0 # Not possible to calculate enthalpy of reaction TbF3:.5H2O
# Enthalpy of formation: 0 kcal/mol
TbPO4:10H2O
TbPO4:10H2O +1.0000 H+ = + 1.0000 HPO4-- + 1.0000 Tb+++ + 10.0000 H2O
log_k -11.9782
-delta_H 0 # Not possible to calculate enthalpy of reaction TbPO4:10H2O
# Enthalpy of formation: 0 kcal/mol
Tc
Tc +1.7500 O2 +0.5000 H2O = + 1.0000 H+ + 1.0000 TcO4-
log_k 93.5811
-delta_H -552.116 kJ/mol # Calculated enthalpy of reaction Tc
# Enthalpy of formation: 0 kJ/mol
-analytic 2.2670e+001 -1.2050e-002 3.0174e+004 -8.4053e+000 -5.2577e+005
# -Range: 0-300
Tc(OH)2
Tc(OH)2 +3.0000 H+ +0.2500 O2 = + 1.0000 Tc+++ + 2.5000 H2O
log_k 5.2714
-delta_H 0 # Not possible to calculate enthalpy of reaction Tc(OH)2
# Enthalpy of formation: 0 kcal/mol
Tc(OH)3
Tc(OH)3 +3.0000 H+ = + 1.0000 Tc+++ + 3.0000 H2O
log_k -9.2425
-delta_H 0 # Not possible to calculate enthalpy of reaction Tc(OH)3
# Enthalpy of formation: 0 kcal/mol
Tc2O7
Tc2O7 +1.0000 H2O = + 2.0000 H+ + 2.0000 TcO4-
log_k 13.1077
-delta_H -26.5357 kJ/mol # Calculated enthalpy of reaction Tc2O7
# Enthalpy of formation: -1120.16 kJ/mol
-analytic 8.7535e+001 1.5366e-002 -1.1919e+003 -3.0317e+001 -2.0271e+001
# -Range: 0-200
Tc2S7
Tc2S7 +8.0000 H2O = + 2.0000 TcO4- + 7.0000 HS- + 9.0000 H+
log_k -230.2410
-delta_H 1356.41 kJ/mol # Calculated enthalpy of reaction Tc2S7
# Enthalpy of formation: -615 kJ/mol
-analytic 2.4560e+002 -4.3355e-002 -8.4192e+004 -7.2967e+001 -1.4298e+003
# -Range: 0-200
Tc3O4
Tc3O4 +9.0000 H+ +0.2500 O2 = + 3.0000 Tc+++ + 4.5000 H2O
log_k -19.2271
-delta_H 0 # Not possible to calculate enthalpy of reaction Tc3O4
# Enthalpy of formation: 0 kcal/mol
Tc4O7
Tc4O7 +10.0000 H+ = + 2.0000 Tc+++ + 2.0000 TcO++ + 5.0000 H2O
log_k -26.0149
-delta_H 0 # Not possible to calculate enthalpy of reaction Tc4O7
# Enthalpy of formation: 0 kcal/mol
TcO2:2H2O(am)
TcO2:2H2O +2.0000 H+ = + 1.0000 TcO++ + 3.0000 H2O
log_k -4.2319
-delta_H 0 # Not possible to calculate enthalpy of reaction TcO2:2H2O(am)
# Enthalpy of formation: 0 kcal/mol
TcO3
TcO3 +1.0000 H2O = + 1.0000 TcO4-- + 2.0000 H+
log_k -23.1483
-delta_H 0 # Not possible to calculate enthalpy of reaction TcO3
# Enthalpy of formation: -540 kJ/mol
TcOH
TcOH +3.0000 H+ +0.5000 O2 = + 1.0000 Tc+++ + 2.0000 H2O
log_k 24.9009
-delta_H 0 # Not possible to calculate enthalpy of reaction TcOH
# Enthalpy of formation: 0 kcal/mol
TcS2
TcS2 +1.0000 H2O = + 1.0000 TcO++ + 2.0000 HS-
log_k -65.9742
-delta_H 0 # Not possible to calculate enthalpy of reaction TcS2
# Enthalpy of formation: -224 kJ/mol
TcS3
TcS3 +4.0000 H2O = + 1.0000 TcO4-- + 3.0000 HS- + 5.0000 H+
log_k -119.5008
-delta_H 0 # Not possible to calculate enthalpy of reaction TcS3
# Enthalpy of formation: -276 kJ/mol
Tenorite
CuO +2.0000 H+ = + 1.0000 Cu++ + 1.0000 H2O
log_k 7.6560
-delta_H -64.5047 kJ/mol # Calculated enthalpy of reaction Tenorite
# Enthalpy of formation: -37.2 kcal/mol
-analytic -8.9899e+001 -1.8886e-002 6.0346e+003 3.3517e+001 9.4191e+001
# -Range: 0-300
Tephroite
Mn2SiO4 +4.0000 H+ = + 1.0000 SiO2 + 2.0000 H2O + 2.0000 Mn++
log_k 23.0781
-delta_H -160.1 kJ/mol # Calculated enthalpy of reaction Tephroite
# Enthalpy of formation: -1730.47 kJ/mol
-analytic -3.2440e+001 -1.1023e-002 8.8910e+003 1.1691e+001 1.3875e+002
# -Range: 0-300
Th
Th +4.0000 H+ +1.0000 O2 = + 1.0000 Th++++ + 2.0000 H2O
log_k 209.6028
-delta_H -1328.56 kJ/mol # Calculated enthalpy of reaction Th
# Enthalpy of formation: 0 kJ/mol
-analytic -2.8256e+001 -1.1963e-002 6.8870e+004 4.2068e+000 1.0747e+003
# -Range: 0-300
Th(NO3)4:5H2O
Th(NO3)4:5H2O = + 1.0000 Th++++ + 4.0000 NO3- + 5.0000 H2O
log_k 1.7789
-delta_H -18.1066 kJ/mol # Calculated enthalpy of reaction Th(NO3)4:5H2O
# Enthalpy of formation: -3007.35 kJ/mol
-analytic -1.2480e+002 -2.0405e-002 5.1601e+003 4.6613e+001 8.7669e+001
# -Range: 0-200
Th(OH)4
Th(OH)4 +4.0000 H+ = + 1.0000 Th++++ + 4.0000 H2O
log_k 9.6543
-delta_H -140.336 kJ/mol # Calculated enthalpy of reaction Th(OH)4
# Enthalpy of formation: -423.527 kcal/mol
-analytic -1.4031e+002 -9.2493e-003 1.2345e+004 4.4990e+001 2.0968e+002
# -Range: 0-200
Th(SO4)2
Th(SO4)2 = + 1.0000 Th++++ + 2.0000 SO4--
log_k -20.3006
-delta_H -46.1064 kJ/mol # Calculated enthalpy of reaction Th(SO4)2
# Enthalpy of formation: -2542.12 kJ/mol
-analytic -8.4525e+000 -3.5442e-002 0.0000e+000 0.0000e+000 -1.1540e+005
# -Range: 0-200
Th2S3
Th2S3 +5.0000 H+ +0.5000 O2 = + 1.0000 H2O + 2.0000 Th++++ + 3.0000 HS-
log_k 95.2290
-delta_H -783.243 kJ/mol # Calculated enthalpy of reaction Th2S3
# Enthalpy of formation: -1082.89 kJ/mol
-analytic -3.2969e+002 -1.1090e-001 4.6877e+004 1.2152e+002 7.3157e+002
# -Range: 0-300
Th2Se3
Th2Se3 +2.0000 H+ +0.5000 O2 = + 1.0000 H2O + 2.0000 Th++++ + 3.0000 Se--
log_k 59.1655
-delta_H 0 # Not possible to calculate enthalpy of reaction Th2Se3
# Enthalpy of formation: -224 kcal/mol
-analytic -1.0083e+001 6.0240e-003 3.4039e+004 -1.8884e+001 5.7804e+002
# -Range: 0-200
Th7S12
Th7S12 +16.0000 H+ +1.0000 O2 = + 2.0000 H2O + 7.0000 Th++++ + 12.0000 HS-
log_k 204.0740
-delta_H -1999.4 kJ/mol # Calculated enthalpy of reaction Th7S12
# Enthalpy of formation: -4136.58 kJ/mol
-analytic -2.1309e+002 -1.4149e-001 9.8550e+004 5.2042e+001 1.6736e+003
# -Range: 0-200
ThBr4
ThBr4 = + 1.0000 Th++++ + 4.0000 Br-
log_k 34.0803
-delta_H -290.23 kJ/mol # Calculated enthalpy of reaction ThBr4
# Enthalpy of formation: -964.803 kJ/mol
-analytic 2.9902e+001 -3.3109e-002 1.0988e+004 -9.2209e+000 1.8657e+002
# -Range: 0-200
ThCl4
ThCl4 = + 1.0000 Th++++ + 4.0000 Cl-
log_k 23.8491
-delta_H -251.094 kJ/mol # Calculated enthalpy of reaction ThCl4
# Enthalpy of formation: -283.519 kcal/mol
-analytic -5.9340e+000 -4.1640e-002 9.8623e+003 3.6804e+000 1.6748e+002
# -Range: 0-200
ThF4
ThF4 = + 1.0000 Th++++ + 4.0000 F-
log_k -29.9946
-delta_H -12.6733 kJ/mol # Calculated enthalpy of reaction ThF4
# Enthalpy of formation: -501.371 kcal/mol
-analytic -4.2622e+002 -1.4222e-001 9.4201e+003 1.6446e+002 1.4712e+002
# -Range: 0-300
ThF4:2.5H2O
ThF4:2.5H2O = + 1.0000 Th++++ + 2.5000 H2O + 4.0000 F-
log_k -31.8568
-delta_H 22.6696 kJ/mol # Calculated enthalpy of reaction ThF4:2.5H2O
# Enthalpy of formation: -2847.68 kJ/mol
-analytic -1.1284e+002 -4.5422e-002 -2.5781e+002 3.8547e+001 -4.3396e+000
# -Range: 0-200
ThI4
ThI4 = + 1.0000 Th++++ + 4.0000 I-
log_k 45.1997
-delta_H -332.818 kJ/mol # Calculated enthalpy of reaction ThI4
# Enthalpy of formation: -663.811 kJ/mol
-analytic 1.4224e+000 -4.0379e-002 1.4193e+004 3.3137e+000 2.4102e+002
# -Range: 0-200
ThS
ThS +3.0000 H+ +0.5000 O2 = + 1.0000 H2O + 1.0000 HS- + 1.0000 Th++++
log_k 96.0395
-delta_H -669.906 kJ/mol # Calculated enthalpy of reaction ThS
# Enthalpy of formation: -394.993 kJ/mol
-analytic -1.3919e+001 -1.2372e-002 3.3883e+004 0.0000e+000 0.0000e+000
# -Range: 0-200
ThS2
ThS2 +2.0000 H+ = + 1.0000 Th++++ + 2.0000 HS-
log_k 10.7872
-delta_H -175.369 kJ/mol # Calculated enthalpy of reaction ThS2
# Enthalpy of formation: -625.867 kJ/mol
-analytic -3.7691e+001 -2.3714e-002 8.4673e+003 1.0970e+001 1.4380e+002
# -Range: 0-200
Thenardite
Na2SO4 = + 1.0000 SO4-- + 2.0000 Na+
log_k -0.3091
-delta_H -2.33394 kJ/mol # Calculated enthalpy of reaction Thenardite
# Enthalpy of formation: -1387.87 kJ/mol
-analytic -2.1202e+002 -7.1613e-002 5.1083e+003 8.7244e+001 7.9773e+001
# -Range: 0-300
Thermonatrite
Na2CO3:H2O +1.0000 H+ = + 1.0000 H2O + 1.0000 HCO3- + 2.0000 Na+
log_k 10.9623
-delta_H -27.5869 kJ/mol # Calculated enthalpy of reaction Thermonatrite
# Enthalpy of formation: -1428.78 kJ/mol
-analytic -1.4030e+002 -3.5263e-002 5.7840e+003 5.7528e+001 9.0295e+001
# -Range: 0-300
Thorianite
ThO2 +4.0000 H+ = + 1.0000 Th++++ + 2.0000 H2O
log_k 1.8624
-delta_H -114.296 kJ/mol # Calculated enthalpy of reaction Thorianite
# Enthalpy of formation: -1226.4 kJ/mol
-analytic -1.4249e+001 -2.4645e-003 4.3110e+003 -1.6605e-002 2.1598e+005
# -Range: 0-300
Ti
Ti +2.0000 H2O +1.0000 O2 = + 1.0000 Ti(OH)4
log_k 149.2978
-delta_H 0 # Not possible to calculate enthalpy of reaction Ti
# Enthalpy of formation: 0 kJ/mol
Ti2O3
Ti2O3 +4.0000 H2O +0.5000 O2 = + 2.0000 Ti(OH)4
log_k 42.9866
-delta_H 0 # Not possible to calculate enthalpy of reaction Ti2O3
# Enthalpy of formation: -1520.78 kJ/mol
Ti3O5
Ti3O5 +6.0000 H2O +0.5000 O2 = + 3.0000 Ti(OH)4
log_k 34.6557
-delta_H 0 # Not possible to calculate enthalpy of reaction Ti3O5
# Enthalpy of formation: -2459.24 kJ/mol
TiB2
TiB2 +5.0000 H2O +2.5000 O2 = + 1.0000 Ti(OH)4 + 2.0000 B(OH)3
log_k 312.4194
-delta_H 0 # Not possible to calculate enthalpy of reaction TiB2
# Enthalpy of formation: -323.883 kJ/mol
TiBr3
TiBr3 +3.5000 H2O +0.2500 O2 = + 1.0000 Ti(OH)4 + 3.0000 Br- + 3.0000 H+
log_k 47.7190
-delta_H 0 # Not possible to calculate enthalpy of reaction TiBr3
# Enthalpy of formation: -548.378 kJ/mol
TiBr4
TiBr4 +4.0000 H2O = + 1.0000 Ti(OH)4 + 4.0000 Br- + 4.0000 H+
log_k 32.9379
-delta_H 0 # Not possible to calculate enthalpy of reaction TiBr4
# Enthalpy of formation: -616.822 kJ/mol
TiC
TiC +3.0000 H2O +2.0000 O2 = + 1.0000 H+ + 1.0000 HCO3- + 1.0000 Ti(OH)4
log_k 181.8139
-delta_H 0 # Not possible to calculate enthalpy of reaction TiC
# Enthalpy of formation: -184.346 kJ/mol
TiCl2
TiCl2 +3.0000 H2O +0.5000 O2 = + 1.0000 Ti(OH)4 + 2.0000 Cl- + 2.0000 H+
log_k 70.9386
-delta_H 0 # Not possible to calculate enthalpy of reaction TiCl2
# Enthalpy of formation: -514.012 kJ/mol
TiCl3
TiCl3 +3.5000 H2O +0.2500 O2 = + 1.0000 Ti(OH)4 + 3.0000 Cl- + 3.0000 H+
log_k 39.3099
-delta_H 0 # Not possible to calculate enthalpy of reaction TiCl3
# Enthalpy of formation: -720.775 kJ/mol
TiF4(am)
TiF4 +4.0000 H2O = + 1.0000 Ti(OH)4 + 4.0000 F- + 4.0000 H+
log_k -12.4409
-delta_H 0 # Not possible to calculate enthalpy of reaction TiF4(am)
# Enthalpy of formation: -1649.44 kJ/mol
TiI4
TiI4 +4.0000 H2O = + 1.0000 Ti(OH)4 + 4.0000 H+ + 4.0000 I-
log_k 34.5968
-delta_H 0 # Not possible to calculate enthalpy of reaction TiI4
# Enthalpy of formation: -375.555 kJ/mol
TiN
TiN +3.5000 H2O +0.2500 O2 = + 1.0000 NH3 + 1.0000 Ti(OH)4
log_k 35.2344
-delta_H 0 # Not possible to calculate enthalpy of reaction TiN
# Enthalpy of formation: -338.304 kJ/mol
TiO(alpha)
TiO +2.0000 H2O +0.5000 O2 = + 1.0000 Ti(OH)4
log_k 61.1282
-delta_H 0 # Not possible to calculate enthalpy of reaction TiO(alpha)
# Enthalpy of formation: -519.835 kJ/mol
Tiemannite
HgSe = + 1.0000 Hg++ + 1.0000 Se--
log_k -58.2188
-delta_H 0 # Not possible to calculate enthalpy of reaction Tiemannite
# Enthalpy of formation: -10.4 kcal/mol
-analytic -5.7618e+001 -1.3891e-002 -1.3223e+004 1.9351e+001 -2.0632e+002
# -Range: 0-300
Titanite
CaTiSiO5 +2.0000 H+ +1.0000 H2O = + 1.0000 Ca++ + 1.0000 SiO2 + 1.0000 Ti(OH)4
log_k 719.5839
-delta_H 0 # Not possible to calculate enthalpy of reaction Titanite
# Enthalpy of formation: 0 kcal/mol
Tl
Tl +1.0000 H+ +0.2500 O2 = + 0.5000 H2O + 1.0000 Tl+
log_k 27.1743
-delta_H -134.53 kJ/mol # Calculated enthalpy of reaction Tl
# Enthalpy of formation: 0 kJ/mol
-analytic -3.7066e+001 -7.8341e-003 9.4594e+003 1.4896e+001 -1.7904e+005
# -Range: 0-300
Tm
Tm +3.0000 H+ +0.7500 O2 = + 1.0000 Tm+++ + 1.5000 H2O
log_k 181.7102
-delta_H -1124.66 kJ/mol # Calculated enthalpy of reaction Tm
# Enthalpy of formation: 0 kJ/mol
-analytic -6.7440e+001 -2.8476e-002 5.9332e+004 2.3715e+001 -5.9611e+003
# -Range: 0-300
Tm(OH)3
Tm(OH)3 +3.0000 H+ = + 1.0000 Tm+++ + 3.0000 H2O
log_k 14.9852
-delta_H 0 # Not possible to calculate enthalpy of reaction Tm(OH)3
# Enthalpy of formation: 0 kcal/mol
Tm(OH)3(am)
Tm(OH)3 +3.0000 H+ = + 1.0000 Tm+++ + 3.0000 H2O
log_k 17.2852
-delta_H 0 # Not possible to calculate enthalpy of reaction Tm(OH)3(am)
# Enthalpy of formation: 0 kcal/mol
Tm2(CO3)3
Tm2(CO3)3 +3.0000 H+ = + 2.0000 Tm+++ + 3.0000 HCO3-
log_k -2.4136
-delta_H 0 # Not possible to calculate enthalpy of reaction Tm2(CO3)3
# Enthalpy of formation: 0 kcal/mol
Tm2O3
Tm2O3 +6.0000 H+ = + 2.0000 Tm+++ + 3.0000 H2O
log_k 44.7000
-delta_H 0 # Not possible to calculate enthalpy of reaction Tm2O3
# Enthalpy of formation: 0 kcal/mol
TmF3:.5H2O
TmF3:.5H2O = + 0.5000 H2O + 1.0000 Tm+++ + 3.0000 F-
log_k -16.2000
-delta_H 0 # Not possible to calculate enthalpy of reaction TmF3:.5H2O
# Enthalpy of formation: 0 kcal/mol
TmPO4:10H2O
TmPO4:10H2O +1.0000 H+ = + 1.0000 HPO4-- + 1.0000 Tm+++ + 10.0000 H2O
log_k -11.8782
-delta_H 0 # Not possible to calculate enthalpy of reaction TmPO4:10H2O
# Enthalpy of formation: 0 kcal/mol
Tobermorite-11A
Ca5Si6H11O22.5 +10.0000 H+ = + 5.0000 Ca++ + 6.0000 SiO2 + 10.5000 H2O
log_k 65.6121
-delta_H -286.861 kJ/mol # Calculated enthalpy of reaction Tobermorite-11A
# Enthalpy of formation: -2556.42 kcal/mol
-analytic 7.9123e+001 3.9150e-002 2.9429e+004 -3.9191e+001 -2.4122e+006
# -Range: 0-300
Tobermorite-14A
Ca5Si6H21O27.5 +10.0000 H+ = + 5.0000 Ca++ + 6.0000 SiO2 + 15.5000 H2O
log_k 63.8445
-delta_H -230.959 kJ/mol # Calculated enthalpy of reaction Tobermorite-14A
# Enthalpy of formation: -2911.36 kcal/mol
-analytic -2.0789e+002 5.2472e-003 3.9698e+004 6.7797e+001 -2.7532e+006
# -Range: 0-300
Tobermorite-9A
Ca5Si6H6O20 +10.0000 H+ = + 5.0000 Ca++ + 6.0000 SiO2 + 8.0000 H2O
log_k 69.0798
-delta_H -329.557 kJ/mol # Calculated enthalpy of reaction Tobermorite-9A
# Enthalpy of formation: -2375.42 kcal/mol
-analytic -6.3384e+001 1.1722e-002 3.8954e+004 1.2268e+001 -2.8681e+006
# -Range: 0-300
Todorokite
Mn7O12:3H2O +16.0000 H+ = + 1.0000 MnO4-- + 6.0000 Mn+++ + 11.0000 H2O
log_k -45.8241
-delta_H 0 # Not possible to calculate enthalpy of reaction Todorokite
# Enthalpy of formation: 0 kcal/mol
Torbernite
Cu(UO2)2(PO4)2 +2.0000 H+ = + 1.0000 Cu++ + 2.0000 HPO4-- + 2.0000 UO2++
log_k -20.3225
-delta_H -97.4022 kJ/mol # Calculated enthalpy of reaction Torbernite
# Enthalpy of formation: -1065.74 kcal/mol
-analytic -6.7128e+001 -4.5878e-002 3.5071e+003 1.9682e+001 5.9579e+001
# -Range: 0-200
Tremolite
Ca2Mg5Si8O22(OH)2 +14.0000 H+ = + 2.0000 Ca++ + 5.0000 Mg++ + 8.0000 H2O + 8.0000 SiO2
log_k 61.2367
-delta_H -406.404 kJ/mol # Calculated enthalpy of reaction Tremolite
# Enthalpy of formation: -2944.04 kcal/mol
-analytic 8.5291e+001 4.6337e-002 3.9465e+004 -5.4414e+001 -3.1913e+006
# -Range: 0-300
Trevorite
NiFe2O4 +8.0000 H+ = + 1.0000 Ni++ + 2.0000 Fe+++ + 4.0000 H2O
log_k 9.7876
-delta_H -215.338 kJ/mol # Calculated enthalpy of reaction Trevorite
# Enthalpy of formation: -1081.15 kJ/mol
-analytic -1.4322e+002 -2.9429e-002 1.4518e+004 4.5698e+001 2.4658e+002
# -Range: 0-200
Tridymite
SiO2 = + 1.0000 SiO2
log_k -3.8278
-delta_H 31.3664 kJ/mol # Calculated enthalpy of reaction Tridymite
# Enthalpy of formation: -909.065 kJ/mol
-analytic 3.1594e+002 6.9315e-002 -1.1358e+004 -1.2219e+002 -1.9299e+002
# -Range: 0-200
Troilite
FeS +1.0000 H+ = + 1.0000 Fe++ + 1.0000 HS-
log_k -3.8184
-delta_H -7.3296 kJ/mol # Calculated enthalpy of reaction Troilite
# Enthalpy of formation: -101.036 kJ/mol
-analytic -1.6146e+002 -5.3170e-002 4.0461e+003 6.4620e+001 6.3183e+001
# -Range: 0-300
Trona-K
K2NaH(CO3)2:2H2O +1.0000 H+ = + 1.0000 Na+ + 2.0000 H2O + 2.0000 HCO3- + 2.0000 K+
log_k 11.5891
-delta_H 0 # Not possible to calculate enthalpy of reaction Trona-K
# Enthalpy of formation: 0 kcal/mol
Tsumebite
Pb2Cu(PO4)(OH)3:3H2O +4.0000 H+ = + 1.0000 Cu++ + 1.0000 HPO4-- + 2.0000 Pb++ + 6.0000 H2O
log_k 2.5318
-delta_H 0 # Not possible to calculate enthalpy of reaction Tsumebite
# Enthalpy of formation: 0 kcal/mol
Tyuyamunite
Ca(UO2)2(VO4)2 = + 1.0000 Ca++ + 2.0000 UO2++ + 2.0000 VO4---
log_k -53.3757
-delta_H 0 # Not possible to calculate enthalpy of reaction Tyuyamunite
# Enthalpy of formation: -1164.52 kcal/mol
U
U +2.0000 H+ +1.5000 O2 = + 1.0000 H2O + 1.0000 UO2++
log_k 212.7800
-delta_H -1286.64 kJ/mol # Calculated enthalpy of reaction U
# Enthalpy of formation: 0 kJ/mol
-analytic -2.4912e+002 -4.7104e-002 8.1115e+004 8.7008e+001 -1.0158e+006
# -Range: 0-300
U(CO3)2
U(CO3)2 +2.0000 H+ = + 1.0000 U++++ + 2.0000 HCO3-
log_k 7.5227
-delta_H -170.691 kJ/mol # Calculated enthalpy of reaction U(CO3)2
# Enthalpy of formation: -1800.38 kJ/mol
-analytic -8.5952e+001 -2.5086e-002 1.0177e+004 2.7002e+001 1.7285e+002
# -Range: 0-200
U(HPO4)2:4H2O
U(HPO4)2:4H2O = + 1.0000 U++++ + 2.0000 HPO4-- + 4.0000 H2O
log_k -32.8650
-delta_H 16.1008 kJ/mol # Calculated enthalpy of reaction U(HPO4)2:4H2O
# Enthalpy of formation: -4334.82 kJ/mol
-analytic -3.8694e+002 -1.3874e-001 6.4882e+003 1.5099e+002 1.0136e+002
# -Range: 0-300
U(OH)2SO4
U(OH)2SO4 +2.0000 H+ = + 1.0000 SO4-- + 1.0000 U++++ + 2.0000 H2O
log_k -3.0731
-delta_H 0 # Not possible to calculate enthalpy of reaction U(OH)2SO4
# Enthalpy of formation: 0 kcal/mol
U(SO3)2
U(SO3)2 = + 1.0000 U++++ + 2.0000 SO3--
log_k -36.7499
-delta_H 20.7008 kJ/mol # Calculated enthalpy of reaction U(SO3)2
# Enthalpy of formation: -1883 kJ/mol
-analytic 5.8113e+001 -2.9981e-002 -7.0503e+003 -2.5175e+001 -1.1974e+002
# -Range: 0-200
U(SO4)2
U(SO4)2 = + 1.0000 U++++ + 2.0000 SO4--
log_k -11.5178
-delta_H -100.803 kJ/mol # Calculated enthalpy of reaction U(SO4)2
# Enthalpy of formation: -2309.6 kJ/mol
-analytic 3.2215e+001 -2.8662e-002 7.1066e+002 -1.5190e+001 1.2057e+001
# -Range: 0-200
U(SO4)2:4H2O
U(SO4)2:4H2O = + 1.0000 U++++ + 2.0000 SO4-- + 4.0000 H2O
log_k -11.5287
-delta_H -70.5565 kJ/mol # Calculated enthalpy of reaction U(SO4)2:4H2O
# Enthalpy of formation: -3483.2 kJ/mol
-analytic -6.9548e+001 -2.9094e-002 3.8763e+003 2.1692e+001 6.5849e+001
# -Range: 0-200
U(SO4)2:8H2O
U(SO4)2:8H2O = + 1.0000 U++++ + 2.0000 SO4-- + 8.0000 H2O
log_k -12.5558
-delta_H -34.5098 kJ/mol # Calculated enthalpy of reaction U(SO4)2:8H2O
# Enthalpy of formation: -4662.6 kJ/mol
-analytic -1.7141e+002 -2.9548e-002 6.7423e+003 5.8614e+001 1.1455e+002
# -Range: 0-200
U2C3
U2C3 +4.5000 O2 +3.0000 H+ = + 2.0000 U+++ + 3.0000 HCO3-
log_k 455.3078
-delta_H -2810.1 kJ/mol # Calculated enthalpy of reaction U2C3
# Enthalpy of formation: -183.3 kJ/mol
-analytic -3.8340e+002 -1.5374e-001 1.5922e+005 1.4643e+002 -1.0584e+006
# -Range: 0-300
U2F9
U2F9 +2.0000 H2O = + 1.0000 U++++ + 1.0000 UO2+ + 4.0000 H+ + 9.0000 F-
log_k -45.5022
-delta_H -46.8557 kJ/mol # Calculated enthalpy of reaction U2F9
# Enthalpy of formation: -4015.92 kJ/mol
-analytic -8.8191e+002 -3.0477e-001 2.0493e+004 3.4690e+002 3.2003e+002
# -Range: 0-300
U2O2Cl5
U2O2Cl5 = + 1.0000 U++++ + 1.0000 UO2+ + 5.0000 Cl-
log_k 19.2752
-delta_H -254.325 kJ/mol # Calculated enthalpy of reaction U2O2Cl5
# Enthalpy of formation: -2197.4 kJ/mol
-analytic -4.3945e+002 -1.6239e-001 2.1694e+004 1.7551e+002 3.3865e+002
# -Range: 0-300
U2O3F6
U2O3F6 +1.0000 H2O = + 2.0000 H+ + 2.0000 UO2++ + 6.0000 F-
log_k -2.5066
-delta_H -185.047 kJ/mol # Calculated enthalpy of reaction U2O3F6
# Enthalpy of formation: -3579.2 kJ/mol
-analytic -3.2332e+001 -5.9519e-002 5.7857e+003 1.1372e+001 9.8260e+001
# -Range: 0-200
U2S3
U2S3 +3.0000 H+ = + 2.0000 U+++ + 3.0000 HS-
log_k 6.5279
-delta_H -147.525 kJ/mol # Calculated enthalpy of reaction U2S3
# Enthalpy of formation: -879 kJ/mol
-analytic -3.0494e+002 -1.0983e-001 1.3647e+004 1.2059e+002 2.1304e+002
# -Range: 0-300
U2Se3
U2Se3 +4.5000 O2 = + 2.0000 U+++ + 3.0000 SeO3--
log_k 248.0372
-delta_H -1740.18 kJ/mol # Calculated enthalpy of reaction U2Se3
# Enthalpy of formation: -711 kJ/mol
-analytic 4.9999e+002 -1.6488e-002 6.4991e+004 -1.8795e+002 1.1035e+003
# -Range: 0-200
U3As4
U3As4 +5.2500 O2 +5.0000 H+ +1.5000 H2O = + 3.0000 U+++ + 4.0000 H2AsO3-
log_k 487.6802
-delta_H -3114.02 kJ/mol # Calculated enthalpy of reaction U3As4
# Enthalpy of formation: -720 kJ/mol
-analytic -9.0215e+002 -2.5804e-001 1.9974e+005 3.3331e+002 -2.4911e+006
# -Range: 0-300
U3O5F8
U3O5F8 +1.0000 H2O = + 2.0000 H+ + 3.0000 UO2++ + 8.0000 F-
log_k -2.7436
-delta_H -260.992 kJ/mol # Calculated enthalpy of reaction U3O5F8
# Enthalpy of formation: -5192.95 kJ/mol
-analytic -7.7653e+002 -2.7294e-001 2.9180e+004 3.0599e+002 4.5556e+002
# -Range: 0-300
U3P4
U3P4 +7.2500 O2 +1.5000 H2O +1.0000 H+ = + 3.0000 U+++ + 4.0000 HPO4--
log_k 827.5586
-delta_H -5275.9 kJ/mol # Calculated enthalpy of reaction U3P4
# Enthalpy of formation: -843 kJ/mol
-analytic -2.7243e+003 -6.2927e-001 4.0130e+005 1.0021e+003 -7.6720e+006
# -Range: 0-300
U3S5
U3S5 +5.0000 H+ = + 1.0000 U++++ + 2.0000 U+++ + 5.0000 HS-
log_k -0.3680
-delta_H -218.942 kJ/mol # Calculated enthalpy of reaction U3S5
# Enthalpy of formation: -1431 kJ/mol
-analytic -1.1011e+002 -6.7959e-002 1.0369e+004 3.8481e+001 1.7611e+002
# -Range: 0-200
U3Sb4
U3Sb4 +9.0000 H+ +5.2500 O2 +1.5000 H2O = + 3.0000 U+++ + 4.0000 Sb(OH)3
log_k 575.0349
-delta_H -3618.1 kJ/mol # Calculated enthalpy of reaction U3Sb4
# Enthalpy of formation: -451.9 kJ/mol
U3Se4
U3Se4 +6.2500 O2 +1.0000 H+ = + 0.5000 H2O + 3.0000 U+++ + 4.0000 SeO3--
log_k 375.2823
-delta_H -2588.16 kJ/mol # Calculated enthalpy of reaction U3Se4
# Enthalpy of formation: -983 kJ/mol
-analytic 6.7219e+002 -2.2708e-002 1.0025e+005 -2.5317e+002 1.7021e+003
# -Range: 0-200
U3Se5
U3Se5 +7.2500 O2 +0.5000 H2O = + 1.0000 H+ + 3.0000 U+++ + 5.0000 SeO3--
log_k 376.5747
-delta_H -2652.38 kJ/mol # Calculated enthalpy of reaction U3Se5
# Enthalpy of formation: -1130 kJ/mol
-analytic 8.3306e+002 -2.6526e-002 9.5737e+004 -3.1109e+002 1.6255e+003
# -Range: 0-200
U4F17
U4F17 +2.0000 H2O = + 1.0000 UO2+ + 3.0000 U++++ + 4.0000 H+ + 17.0000 F-
log_k -104.7657
-delta_H -78.2955 kJ/mol # Calculated enthalpy of reaction U4F17
# Enthalpy of formation: -7849.66 kJ/mol
-analytic -1.7466e+003 -5.9186e-001 4.0017e+004 6.8046e+002 6.2494e+002
# -Range: 0-300
U5O12Cl
U5O12Cl +4.0000 H+ = + 1.0000 Cl- + 2.0000 H2O + 5.0000 UO2+
log_k -18.7797
-delta_H -9.99133 kJ/mol # Calculated enthalpy of reaction U5O12Cl
# Enthalpy of formation: -5854.4 kJ/mol
-analytic -7.3802e+001 2.9180e-002 4.6804e+003 1.2371e+001 7.9503e+001
# -Range: 0-200
UAs
UAs +2.0000 H+ +1.5000 O2 = + 1.0000 H2AsO3- + 1.0000 U+++
log_k 149.0053
-delta_H -951.394 kJ/mol # Calculated enthalpy of reaction UAs
# Enthalpy of formation: -234.3 kJ/mol
-analytic -5.0217e+001 -4.2992e-002 4.8480e+004 1.9964e+001 7.5650e+002
# -Range: 0-300
UAs2
UAs2 +2.2500 O2 +1.5000 H2O +1.0000 H+ = + 1.0000 U+++ + 2.0000 H2AsO3-
log_k 189.1058
-delta_H -1210.63 kJ/mol # Calculated enthalpy of reaction UAs2
# Enthalpy of formation: -252 kJ/mol
-analytic -8.7361e+001 -7.5252e-002 6.1445e+004 3.7485e+001 9.5881e+002
# -Range: 0-300
UBr2Cl
UBr2Cl = + 1.0000 Cl- + 1.0000 U+++ + 2.0000 Br-
log_k 17.7796
-delta_H -148.586 kJ/mol # Calculated enthalpy of reaction UBr2Cl
# Enthalpy of formation: -750.6 kJ/mol
-analytic 3.0364e+000 -3.2187e-002 5.2314e+003 2.7418e+000 8.8836e+001
# -Range: 0-200
UBr2Cl2
UBr2Cl2 = + 1.0000 U++++ + 2.0000 Br- + 2.0000 Cl-
log_k 26.2185
-delta_H -260.466 kJ/mol # Calculated enthalpy of reaction UBr2Cl2
# Enthalpy of formation: -907.9 kJ/mol
-analytic 3.8089e+000 -3.8781e-002 1.0125e+004 0.0000e+000 0.0000e+000
# -Range: 0-200
UBr3
UBr3 = + 1.0000 U+++ + 3.0000 Br-
log_k 20.2249
-delta_H -154.91 kJ/mol # Calculated enthalpy of reaction UBr3
# Enthalpy of formation: -698.7 kJ/mol
-analytic -2.4366e+002 -9.8651e-002 1.2538e+004 1.0151e+002 1.9572e+002
# -Range: 0-300
UBr3Cl
UBr3Cl = + 1.0000 Cl- + 1.0000 U++++ + 3.0000 Br-
log_k 29.1178
-delta_H -270.49 kJ/mol # Calculated enthalpy of reaction UBr3Cl
# Enthalpy of formation: -852.3 kJ/mol
-analytic 1.1204e+001 -3.7109e-002 1.0473e+004 -2.4905e+000 1.7784e+002
# -Range: 0-200
UBr4
UBr4 = + 1.0000 U++++ + 4.0000 Br-
log_k 31.2904
-delta_H -275.113 kJ/mol # Calculated enthalpy of reaction UBr4
# Enthalpy of formation: -802.1 kJ/mol
-analytic -3.3800e+002 -1.2940e-001 2.0674e+004 1.3678e+002 3.2270e+002
# -Range: 0-300
UBr5
UBr5 +2.0000 H2O = + 1.0000 UO2+ + 4.0000 H+ + 5.0000 Br-
log_k 41.6312
-delta_H -250.567 kJ/mol # Calculated enthalpy of reaction UBr5
# Enthalpy of formation: -810.4 kJ/mol
-analytic -3.2773e+002 -1.4356e-001 1.8709e+004 1.4117e+002 2.9204e+002
# -Range: 0-300
UBrCl2
UBrCl2 = + 1.0000 Br- + 1.0000 U+++ + 2.0000 Cl-
log_k 14.5048
-delta_H -132.663 kJ/mol # Calculated enthalpy of reaction UBrCl2
# Enthalpy of formation: -812.1 kJ/mol
-analytic -5.3713e+000 -3.4256e-002 4.6251e+003 5.8875e+000 7.8542e+001
# -Range: 0-200
UBrCl3
UBrCl3 = + 1.0000 Br- + 1.0000 U++++ + 3.0000 Cl-
log_k 23.5258
-delta_H -246.642 kJ/mol # Calculated enthalpy of reaction UBrCl3
# Enthalpy of formation: -967.3 kJ/mol
-analytic -5.6867e+000 -4.1166e-002 9.6664e+003 3.6579e+000 1.6415e+002
# -Range: 0-200
UC
UC +2.0000 H+ +1.7500 O2 = + 0.5000 H2O + 1.0000 HCO3- + 1.0000 U+++
log_k 194.8241
-delta_H -1202.82 kJ/mol # Calculated enthalpy of reaction UC
# Enthalpy of formation: -97.9 kJ/mol
-analytic -4.6329e+001 -4.4600e-002 6.1417e+004 1.9566e+001 9.5836e+002
# -Range: 0-300
UC1.94(alpha)
UC1.94 +2.6900 O2 +1.0600 H+ +0.4400 H2O = + 1.0000 U+++ + 1.9400 HCO3-
log_k 257.1619
-delta_H -1583.84 kJ/mol # Calculated enthalpy of reaction UC1.94(alpha)
# Enthalpy of formation: -85.324 kJ/mol
-analytic -5.8194e+002 -1.4610e-001 1.0917e+005 2.1638e+002 -1.6852e+006
# -Range: 0-300
UCl2F2
UCl2F2 = + 1.0000 U++++ + 2.0000 Cl- + 2.0000 F-
log_k -3.5085
-delta_H -130.055 kJ/mol # Calculated enthalpy of reaction UCl2F2
# Enthalpy of formation: -1466 kJ/mol
-analytic -3.9662e+002 -1.3879e-001 1.4710e+004 1.5562e+002 2.2965e+002
# -Range: 0-300
UCl2I2
UCl2I2 = + 1.0000 U++++ + 2.0000 Cl- + 2.0000 I-
log_k 30.2962
-delta_H -270.364 kJ/mol # Calculated enthalpy of reaction UCl2I2
# Enthalpy of formation: -768.8 kJ/mol
-analytic -1.2922e+001 -4.3178e-002 1.1219e+004 7.4562e+000 1.9052e+002
# -Range: 0-200
UCl3
UCl3 = + 1.0000 U+++ + 3.0000 Cl-
log_k 13.0062
-delta_H -126.639 kJ/mol # Calculated enthalpy of reaction UCl3
# Enthalpy of formation: -863.7 kJ/mol
-analytic -2.6388e+002 -1.0241e-001 1.1629e+004 1.0846e+002 1.8155e+002
# -Range: 0-300
UCl3F
UCl3F = + 1.0000 F- + 1.0000 U++++ + 3.0000 Cl-
log_k 10.3200
-delta_H -184.787 kJ/mol # Calculated enthalpy of reaction UCl3F
# Enthalpy of formation: -1243 kJ/mol
-analytic -3.7971e+002 -1.3681e-001 1.7127e+004 1.5086e+002 2.6736e+002
# -Range: 0-300
UCl3I
UCl3I = + 1.0000 I- + 1.0000 U++++ + 3.0000 Cl-
log_k 25.5388
-delta_H -251.041 kJ/mol # Calculated enthalpy of reaction UCl3I
# Enthalpy of formation: -898.3 kJ/mol
-analytic -1.3362e+001 -4.3214e-002 1.0167e+004 7.1426e+000 1.7265e+002
# -Range: 0-200
UCl4
UCl4 = + 1.0000 U++++ + 4.0000 Cl-
log_k 21.9769
-delta_H -240.719 kJ/mol # Calculated enthalpy of reaction UCl4
# Enthalpy of formation: -1018.8 kJ/mol
-analytic -3.6881e+002 -1.3618e-001 1.9685e+004 1.4763e+002 3.0727e+002
# -Range: 0-300
UCl5
UCl5 +2.0000 H2O = + 1.0000 UO2+ + 4.0000 H+ + 5.0000 Cl-
log_k 37.3147
-delta_H -249.849 kJ/mol # Calculated enthalpy of reaction UCl5
# Enthalpy of formation: -1039 kJ/mol
-analytic -3.6392e+002 -1.5133e-001 1.9617e+004 1.5376e+002 3.0622e+002
# -Range: 0-300
UCl6
UCl6 +2.0000 H2O = + 1.0000 UO2++ + 4.0000 H+ + 6.0000 Cl-
log_k 57.5888
-delta_H -383.301 kJ/mol # Calculated enthalpy of reaction UCl6
# Enthalpy of formation: -1066.5 kJ/mol
-analytic -4.5589e+002 -1.9203e-001 2.8029e+004 1.9262e+002 4.3750e+002
# -Range: 0-300
UClF3
UClF3 = + 1.0000 Cl- + 1.0000 U++++ + 3.0000 F-
log_k -17.5122
-delta_H -74.3225 kJ/mol # Calculated enthalpy of reaction UClF3
# Enthalpy of formation: -1690 kJ/mol
-analytic -4.1346e+002 -1.4077e-001 1.2237e+004 1.6036e+002 1.9107e+002
# -Range: 0-300
UClI3
UClI3 = + 1.0000 Cl- + 1.0000 U++++ + 3.0000 I-
log_k 35.2367
-delta_H -285.187 kJ/mol # Calculated enthalpy of reaction UClI3
# Enthalpy of formation: -643.8 kJ/mol
-analytic -1.1799e+001 -4.3208e-002 1.2045e+004 7.8829e+000 2.0455e+002
# -Range: 0-200
UF3
UF3 = + 1.0000 U+++ + 3.0000 F-
log_k -19.4125
-delta_H 6.2572 kJ/mol # Calculated enthalpy of reaction UF3
# Enthalpy of formation: -1501.4 kJ/mol
-analytic -3.1530e+002 -1.0945e-001 6.1335e+003 1.2443e+002 9.5799e+001
# -Range: 0-300
UF4
UF4 = + 1.0000 U++++ + 4.0000 F-
log_k -29.2004
-delta_H -18.3904 kJ/mol # Calculated enthalpy of reaction UF4
# Enthalpy of formation: -1914.2 kJ/mol
-analytic -4.2411e+002 -1.4147e-001 9.6621e+003 1.6352e+002 1.5089e+002
# -Range: 0-300
UF4:2.5H2O
UF4:2.5H2O = + 1.0000 U++++ + 2.5000 H2O + 4.0000 F-
log_k -33.3685
-delta_H 24.2888 kJ/mol # Calculated enthalpy of reaction UF4:2.5H2O
# Enthalpy of formation: -2671.47 kJ/mol
-analytic -4.4218e+002 -1.4305e-001 8.2922e+003 1.7118e+002 1.2952e+002
# -Range: 0-300
UF5(alpha)
UF5 +2.0000 H2O = + 1.0000 UO2+ + 4.0000 H+ + 5.0000 F-
log_k -12.8376
-delta_H -54.8883 kJ/mol # Calculated enthalpy of reaction UF5(alpha)
# Enthalpy of formation: -2075.3 kJ/mol
-analytic -4.5126e+002 -1.6121e-001 1.1997e+004 1.8030e+002 1.8733e+002
# -Range: 0-300
UF5(beta)
UF5 +2.0000 H2O = + 1.0000 UO2+ + 4.0000 H+ + 5.0000 F-
log_k -13.1718
-delta_H -46.9883 kJ/mol # Calculated enthalpy of reaction UF5(beta)
# Enthalpy of formation: -2083.2 kJ/mol
-analytic -4.5020e+002 -1.6121e-001 1.1584e+004 1.8030e+002 1.8089e+002
# -Range: 0-300
UF6
UF6 +2.0000 H2O = + 1.0000 UO2++ + 4.0000 H+ + 6.0000 F-
log_k 17.4292
-delta_H -261.709 kJ/mol # Calculated enthalpy of reaction UF6
# Enthalpy of formation: -2197.7 kJ/mol
-analytic -5.8427e+002 -2.1223e-001 2.5296e+004 2.3440e+002 3.9489e+002
# -Range: 0-300
UH3(beta)
UH3 +3.0000 H+ +1.5000 O2 = + 1.0000 U+++ + 3.0000 H2O
log_k 199.7683
-delta_H -1201.43 kJ/mol # Calculated enthalpy of reaction UH3(beta)
# Enthalpy of formation: -126.98 kJ/mol
-analytic 5.2870e+001 4.2151e-003 6.0167e+004 -2.2701e+001 1.0217e+003
# -Range: 0-200
UI3
UI3 = + 1.0000 U+++ + 3.0000 I-
log_k 29.0157
-delta_H -192.407 kJ/mol # Calculated enthalpy of reaction UI3
# Enthalpy of formation: -467.4 kJ/mol
-analytic -2.4505e+002 -9.9867e-002 1.4579e+004 1.0301e+002 2.2757e+002
# -Range: 0-300
UI4
UI4 = + 1.0000 U++++ + 4.0000 I-
log_k 39.3102
-delta_H -300.01 kJ/mol # Calculated enthalpy of reaction UI4
# Enthalpy of formation: -518.8 kJ/mol
-analytic -3.4618e+002 -1.3227e-001 2.2320e+004 1.4145e+002 3.4839e+002
# -Range: 0-300
UN
UN +3.0000 H+ = + 1.0000 NH3 + 1.0000 U+++
log_k 41.7130
-delta_H -280.437 kJ/mol # Calculated enthalpy of reaction UN
# Enthalpy of formation: -290 kJ/mol
-analytic -1.6393e+002 -1.1679e-003 2.8845e+003 6.5637e+001 3.0122e+006
# -Range: 0-300
UN1.59(alpha)
UN1.59 +1.8850 H2O +1.0000 H+ +0.0575 O2 = + 1.0000 UO2+ + 1.5900 NH3
log_k 38.3930
-delta_H -235.75 kJ/mol # Calculated enthalpy of reaction UN1.59(alpha)
# Enthalpy of formation: -379.2 kJ/mol
-analytic 1.8304e+001 1.1109e-002 1.2064e+004 -9.5741e+000 2.0485e+002
# -Range: 0-200
UN1.73(alpha)
UN1.73 +2.0950 H2O +1.0000 H+ = + 0.0475 O2 + 1.0000 UO2+ + 1.7300 NH3
log_k 27.2932
-delta_H -169.085 kJ/mol # Calculated enthalpy of reaction UN1.73(alpha)
# Enthalpy of formation: -398.5 kJ/mol
-analytic 1.0012e+001 1.0398e-002 8.9348e+003 -6.3817e+000 1.5172e+002
# -Range: 0-200
UO2(AsO3)2
UO2(AsO3)2 +2.0000 H2O = + 1.0000 UO2++ + 2.0000 H2AsO4-
log_k 6.9377
-delta_H -109.843 kJ/mol # Calculated enthalpy of reaction UO2(AsO3)2
# Enthalpy of formation: -2156.6 kJ/mol
-analytic -1.6050e+002 -6.6472e-002 8.2129e+003 6.4533e+001 1.2820e+002
# -Range: 0-300
UO2(IO3)2
UO2(IO3)2 = + 1.0000 UO2++ + 2.0000 IO3-
log_k -7.2871
-delta_H -0.3862 kJ/mol # Calculated enthalpy of reaction UO2(IO3)2
# Enthalpy of formation: -1461.28 kJ/mol
-analytic -2.7047e+001 -1.4267e-002 -1.5055e+001 9.7226e+000 -2.4640e-001
# -Range: 0-200
UO2(NO3)2
UO2(NO3)2 = + 1.0000 UO2++ + 2.0000 NO3-
log_k 11.9598
-delta_H -81.6219 kJ/mol # Calculated enthalpy of reaction UO2(NO3)2
# Enthalpy of formation: -1351 kJ/mol
-analytic -1.2216e+001 -1.1261e-002 3.9895e+003 5.7166e+000 6.7751e+001
# -Range: 0-200
UO2(NO3)2:2H2O
UO2(NO3)2:2H2O = + 1.0000 UO2++ + 2.0000 H2O + 2.0000 NO3-
log_k 4.9446
-delta_H -25.5995 kJ/mol # Calculated enthalpy of reaction UO2(NO3)2:2H2O
# Enthalpy of formation: -1978.7 kJ/mol
-analytic -1.3989e+002 -5.2130e-002 4.3758e+003 5.8868e+001 6.8322e+001
# -Range: 0-300
UO2(NO3)2:3H2O
UO2(NO3)2:3H2O = + 1.0000 UO2++ + 2.0000 NO3- + 3.0000 H2O
log_k 3.7161
-delta_H -9.73686 kJ/mol # Calculated enthalpy of reaction UO2(NO3)2:3H2O
# Enthalpy of formation: -2280.4 kJ/mol
-analytic -1.5037e+002 -5.2234e-002 4.0783e+003 6.3024e+001 6.3682e+001
# -Range: 0-300
UO2(NO3)2:6H2O
UO2(NO3)2:6H2O = + 1.0000 UO2++ + 2.0000 NO3- + 6.0000 H2O
log_k 2.3189
-delta_H 19.8482 kJ/mol # Calculated enthalpy of reaction UO2(NO3)2:6H2O
# Enthalpy of formation: -3167.5 kJ/mol
-analytic -1.4019e+002 -4.3682e-002 2.7842e+003 5.9070e+001 4.3486e+001
# -Range: 0-300
UO2(NO3)2:H2O
UO2(NO3)2:H2O = + 1.0000 H2O + 1.0000 UO2++ + 2.0000 NO3-
log_k 8.5103
-delta_H -54.4602 kJ/mol # Calculated enthalpy of reaction UO2(NO3)2:H2O
# Enthalpy of formation: -1664 kJ/mol
-analytic -3.7575e+001 -1.1342e-002 3.7548e+003 1.4899e+001 6.3776e+001
# -Range: 0-200
UO2(OH)2(beta)
UO2(OH)2 +2.0000 H+ = + 1.0000 UO2++ + 2.0000 H2O
log_k 4.9457
-delta_H -56.8767 kJ/mol # Calculated enthalpy of reaction UO2(OH)2(beta)
# Enthalpy of formation: -1533.8 kJ/mol
-analytic -1.7478e+001 -1.6806e-003 3.4226e+003 4.6260e+000 5.3412e+001
# -Range: 0-300
UO2(PO3)2
UO2(PO3)2 +2.0000 H2O = + 1.0000 UO2++ + 2.0000 H+ + 2.0000 HPO4--
log_k -16.2805
-delta_H -58.4873 kJ/mol # Calculated enthalpy of reaction UO2(PO3)2
# Enthalpy of formation: -2973 kJ/mol
-analytic -3.2995e+002 -1.3747e-001 8.0652e+003 1.3237e+002 1.2595e+002
# -Range: 0-300
UO2(am)
UO2 +4.0000 H+ = + 1.0000 U++++ + 2.0000 H2O
log_k 0.1091
-delta_H 0 # Not possible to calculate enthalpy of reaction UO2(am)
# Enthalpy of formation: 0 kcal/mol
UO2.25
UO2.25 +2.5000 H+ = + 0.5000 U++++ + 0.5000 UO2+ + 1.2500 H2O
log_k -4.8193
-delta_H -37.1614 kJ/mol # Calculated enthalpy of reaction UO2.25
# Enthalpy of formation: -1128.3 kJ/mol
-analytic -1.9073e+002 -4.1793e-002 7.3391e+003 7.0213e+001 1.1457e+002
# -Range: 0-300
UO2.25(beta)
UO2.25 +2.5000 H+ = + 0.5000 U++++ + 0.5000 UO2+ + 1.2500 H2O
log_k -4.7593
-delta_H -38.0614 kJ/mol # Calculated enthalpy of reaction UO2.25(beta)
# Enthalpy of formation: -1127.4 kJ/mol
-analytic -3.6654e+001 -2.4013e-003 2.9632e+003 9.1625e+000 4.6249e+001
# -Range: 0-300
UO2.3333(beta)
# UO2.3333 +8.0000 H+ = + 0.3333 O2 + 2.0000 U++++ + 4.0000 H2O
(UO2.3333)2 + 8.0000 H+ = 0.3333 O2 + 2.0000 U++++ + 4.0000 H2O
log_k -27.7177
-delta_H -1187.8 kJ/mol # Calculated enthalpy of reaction UO2.3333(beta)
# Enthalpy of formation: -1142 kJ/mol
-analytic -7.4790e+000 -6.8382e-004 -2.7277e+003 -7.2107e+000 6.1873e+005
# -Range: 0-300
UO2.6667
# UO2.6667 +8.0000 H+ = + 0.6667 O2 + 2.0000 U++++ + 4.0000 H2O
(UO2.6667)2 +8.0000 H+ = + 0.6667 O2 + 2.0000 U++++ + 4.0000 H2O
log_k -43.6051
-delta_H -1142.24 kJ/mol # Calculated enthalpy of reaction UO2.6667
# Enthalpy of formation: -1191.6 kJ/mol
-analytic 1.2095e+002 2.0118e-002 -1.4968e+004 -5.3552e+001 1.0813e+006
# -Range: 0-300
UO2Br2
UO2Br2 = + 1.0000 UO2++ + 2.0000 Br-
log_k 16.5103
-delta_H -124.607 kJ/mol # Calculated enthalpy of reaction UO2Br2
# Enthalpy of formation: -1137.4 kJ/mol
-analytic -1.4876e+002 -6.2715e-002 9.0200e+003 6.2108e+001 1.4079e+002
# -Range: 0-300
UO2Br2:3H2O
UO2Br2:3H2O = + 1.0000 UO2++ + 2.0000 Br- + 3.0000 H2O
log_k 9.4113
-delta_H -61.5217 kJ/mol # Calculated enthalpy of reaction UO2Br2:3H2O
# Enthalpy of formation: -2058 kJ/mol
-analytic -6.8507e+001 -1.6834e-002 5.1409e+003 2.6546e+001 8.7324e+001
# -Range: 0-200
UO2Br2:H2O
UO2Br2:H2O = + 1.0000 H2O + 1.0000 UO2++ + 2.0000 Br-
log_k 12.1233
-delta_H -91.945 kJ/mol # Calculated enthalpy of reaction UO2Br2:H2O
# Enthalpy of formation: -1455.9 kJ/mol
-analytic -1.7519e+001 -1.6603e-002 4.3544e+003 8.0748e+000 7.3950e+001
# -Range: 0-200
UO2BrOH:2H2O
UO2BrOH:2H2O +1.0000 H+ = + 1.0000 Br- + 1.0000 UO2++ + 3.0000 H2O
log_k 4.2026
-delta_H -39.8183 kJ/mol # Calculated enthalpy of reaction UO2BrOH:2H2O
# Enthalpy of formation: -1958.2 kJ/mol
-analytic -8.3411e+001 -1.0024e-002 5.0411e+003 2.9781e+001 8.5633e+001
# -Range: 0-200
UO2CO3
UO2CO3 +1.0000 H+ = + 1.0000 HCO3- + 1.0000 UO2++
log_k -4.1267
-delta_H -19.2872 kJ/mol # Calculated enthalpy of reaction UO2CO3
# Enthalpy of formation: -1689.65 kJ/mol
-analytic -4.4869e+001 -1.1541e-002 1.9475e+003 1.5215e+001 3.3086e+001
# -Range: 0-200
UO2Cl
UO2Cl = + 1.0000 Cl- + 1.0000 UO2+
log_k -0.5154
-delta_H -21.1067 kJ/mol # Calculated enthalpy of reaction UO2Cl
# Enthalpy of formation: -1171.1 kJ/mol
-analytic -7.3291e+001 -2.5940e-002 2.5753e+003 2.9038e+001 4.0207e+001
# -Range: 0-300
UO2Cl2
UO2Cl2 = + 1.0000 UO2++ + 2.0000 Cl-
log_k 12.1394
-delta_H -109.559 kJ/mol # Calculated enthalpy of reaction UO2Cl2
# Enthalpy of formation: -1243.6 kJ/mol
-analytic -1.6569e+002 -6.6249e-002 8.6920e+003 6.8055e+001 1.3568e+002
# -Range: 0-300
UO2Cl2:3H2O
UO2Cl2:3H2O = + 1.0000 UO2++ + 2.0000 Cl- + 3.0000 H2O
log_k 5.6163
-delta_H -45.8743 kJ/mol # Calculated enthalpy of reaction UO2Cl2:3H2O
# Enthalpy of formation: -2164.8 kJ/mol
-analytic -8.4932e+001 -2.0867e-002 4.7594e+003 3.2654e+001 8.0850e+001
# -Range: 0-200
UO2Cl2:H2O
UO2Cl2:H2O = + 1.0000 H2O + 1.0000 UO2++ + 2.0000 Cl-
log_k 8.2880
-delta_H -79.1977 kJ/mol # Calculated enthalpy of reaction UO2Cl2:H2O
# Enthalpy of formation: -1559.8 kJ/mol
-analytic -3.4458e+001 -2.0630e-002 4.1231e+003 1.4170e+001 7.0029e+001
# -Range: 0-200
UO2ClOH:2H2O
UO2ClOH:2H2O +1.0000 H+ = + 1.0000 Cl- + 1.0000 UO2++ + 3.0000 H2O
log_k 2.3064
-delta_H -33.1947 kJ/mol # Calculated enthalpy of reaction UO2ClOH:2H2O
# Enthalpy of formation: -2010.4 kJ/mol
-analytic -9.1834e+001 -1.2041e-002 4.9131e+003 3.2835e+001 8.3462e+001
# -Range: 0-200
UO2F2
UO2F2 = + 1.0000 UO2++ + 2.0000 F-
log_k -7.2302
-delta_H -36.1952 kJ/mol # Calculated enthalpy of reaction UO2F2
# Enthalpy of formation: -1653.5 kJ/mol
-analytic -2.0303e+002 -7.1028e-002 5.9356e+003 7.9627e+001 9.2679e+001
# -Range: 0-300
UO2F2:3H2O
UO2F2:3H2O = + 1.0000 UO2++ + 2.0000 F- + 3.0000 H2O
log_k -7.3692
-delta_H -12.8202 kJ/mol # Calculated enthalpy of reaction UO2F2:3H2O
# Enthalpy of formation: -2534.39 kJ/mol
-analytic -1.0286e+002 -2.1223e-002 3.4855e+003 3.6420e+001 5.9224e+001
# -Range: 0-200
UO2FOH
UO2FOH +1.0000 H+ = + 1.0000 F- + 1.0000 H2O + 1.0000 UO2++
log_k -1.8426
-delta_H -41.7099 kJ/mol # Calculated enthalpy of reaction UO2FOH
# Enthalpy of formation: -1598.48 kJ/mol
-analytic -4.9229e+001 -1.1984e-002 3.2086e+003 1.6244e+001 5.4503e+001
# -Range: 0-200
UO2FOH:2H2O
UO2FOH:2H2O +1.0000 H+ = + 1.0000 F- + 1.0000 UO2++ + 3.0000 H2O
log_k -2.6606
-delta_H -21.8536 kJ/mol # Calculated enthalpy of reaction UO2FOH:2H2O
# Enthalpy of formation: -2190.01 kJ/mol
-analytic -1.0011e+002 -1.2203e-002 4.5446e+003 3.4690e+001 7.7208e+001
# -Range: 0-200
UO2FOH:H2O
UO2FOH:H2O +1.0000 H+ = + 1.0000 F- + 1.0000 UO2++ + 2.0000 H2O
log_k -2.2838
-delta_H -31.5243 kJ/mol # Calculated enthalpy of reaction UO2FOH:H2O
# Enthalpy of formation: -1894.5 kJ/mol
-analytic -7.4628e+001 -1.2086e-002 3.8625e+003 2.5456e+001 6.5615e+001
# -Range: 0-200
UO2HPO4
UO2HPO4 = + 1.0000 HPO4-- + 1.0000 UO2++
log_k -12.6782
-delta_H 0 # Not possible to calculate enthalpy of reaction UO2HPO4
# Enthalpy of formation: 0 kcal/mol
UO2HPO4:4H2O
UO2HPO4:4H2O = + 1.0000 HPO4-- + 1.0000 UO2++ + 4.0000 H2O
log_k -13.0231
-delta_H 15.5327 kJ/mol # Calculated enthalpy of reaction UO2HPO4:4H2O
# Enthalpy of formation: -3469.97 kJ/mol
-analytic -1.1784e+002 -1.9418e-002 2.7547e+003 4.0963e+001 4.6818e+001
# -Range: 0-200
UO2SO3
UO2SO3 = + 1.0000 SO3-- + 1.0000 UO2++
log_k -15.9812
-delta_H 6.4504 kJ/mol # Calculated enthalpy of reaction UO2SO3
# Enthalpy of formation: -1661 kJ/mol
-analytic 2.5751e+001 -1.3871e-002 -3.0305e+003 -1.1090e+001 -5.1470e+001
# -Range: 0-200
UO2SO4
UO2SO4 = + 1.0000 SO4-- + 1.0000 UO2++
log_k 1.9681
-delta_H -83.4616 kJ/mol # Calculated enthalpy of reaction UO2SO4
# Enthalpy of formation: -1845.14 kJ/mol
-analytic -1.5677e+002 -6.5310e-002 6.7411e+003 6.2867e+001 1.0523e+002
# -Range: 0-300
UO2SO4:2.5H2O
UO2SO4:2.5H2O = + 1.0000 SO4-- + 1.0000 UO2++ + 2.5000 H2O
log_k -1.4912
-delta_H -36.1984 kJ/mol # Calculated enthalpy of reaction UO2SO4:2.5H2O
# Enthalpy of formation: -2607 kJ/mol
-analytic -4.8908e+001 -1.3445e-002 2.8658e+003 1.6894e+001 4.8683e+001
# -Range: 0-200
UO2SO4:3.5H2O
UO2SO4:3.5H2O = + 1.0000 SO4-- + 1.0000 UO2++ + 3.5000 H2O
log_k -1.4805
-delta_H -27.4367 kJ/mol # Calculated enthalpy of reaction UO2SO4:3.5H2O
# Enthalpy of formation: -2901.6 kJ/mol
-analytic -7.4180e+001 -1.3565e-002 3.5963e+003 2.6136e+001 6.1096e+001
# -Range: 0-200
UO2SO4:3H2O
UO2SO4:3H2O = + 1.0000 SO4-- + 1.0000 UO2++ + 3.0000 H2O
log_k -1.4028
-delta_H -34.6176 kJ/mol # Calculated enthalpy of reaction UO2SO4:3H2O
# Enthalpy of formation: -2751.5 kJ/mol
-analytic -5.0134e+001 -1.0321e-002 3.0505e+003 1.6799e+001 5.1818e+001
# -Range: 0-200
UO2SO4:H2O
UO2SO4:H2O = + 1.0000 H2O + 1.0000 SO4-- + 1.0000 UO2++
log_k -6.0233
-delta_H -39.1783 kJ/mol # Calculated enthalpy of reaction UO2SO4:H2O
# Enthalpy of formation: -519.9 kcal/mol
-analytic -1.8879e+002 -6.9827e-002 5.5636e+003 7.4717e+001 8.6870e+001
# -Range: 0-300
UO3(alpha)
UO3 +2.0000 H+ = + 1.0000 H2O + 1.0000 UO2++
log_k 8.6391
-delta_H -87.3383 kJ/mol # Calculated enthalpy of reaction UO3(alpha)
# Enthalpy of formation: -1217.5 kJ/mol
-analytic -1.4099e+001 -1.9063e-003 4.7742e+003 2.9478e+000 7.4501e+001
# -Range: 0-300
UO3(beta)
UO3 +2.0000 H+ = + 1.0000 H2O + 1.0000 UO2++
log_k 8.3095
-delta_H -84.5383 kJ/mol # Calculated enthalpy of reaction UO3(beta)
# Enthalpy of formation: -1220.3 kJ/mol
-analytic -1.2298e+001 -1.7800e-003 4.5621e+003 2.3593e+000 7.1191e+001
# -Range: 0-300
UO3(gamma)
UO3 +2.0000 H+ = + 1.0000 H2O + 1.0000 UO2++
log_k 7.7073
-delta_H -81.0383 kJ/mol # Calculated enthalpy of reaction UO3(gamma)
# Enthalpy of formation: -1223.8 kJ/mol
-analytic -1.1573e+001 -2.3560e-003 4.3124e+003 2.2305e+000 6.7294e+001
# -Range: 0-300
UO3:.9H2O(alpha)
UO3:.9H2O +2.0000 H+ = + 1.0000 UO2++ + 1.9000 H2O
log_k 5.0167
-delta_H -55.7928 kJ/mol # Calculated enthalpy of reaction UO3:.9H2O(alpha)
# Enthalpy of formation: -1506.3 kJ/mol
-analytic -6.9286e+001 -3.0624e-003 5.5984e+003 2.2809e+001 9.5092e+001
# -Range: 0-200
UO3:2H2O
UO3:2H2O +2.0000 H+ = + 1.0000 UO2++ + 3.0000 H2O
log_k 4.8333
-delta_H -50.415 kJ/mol # Calculated enthalpy of reaction UO3:2H2O
# Enthalpy of formation: -1826.1 kJ/mol
-analytic -5.9530e+001 -9.8107e-003 4.4975e+003 2.1098e+001 7.0196e+001
# -Range: 0-300
UOBr2
UOBr2 +2.0000 H+ = + 1.0000 H2O + 1.0000 U++++ + 2.0000 Br-
log_k 7.9722
-delta_H -146.445 kJ/mol # Calculated enthalpy of reaction UOBr2
# Enthalpy of formation: -973.6 kJ/mol
-analytic -2.0747e+002 -7.0500e-002 1.1746e+004 7.9629e+001 1.8334e+002
# -Range: 0-300
UOBr3
UOBr3 +1.0000 H2O = + 1.0000 UO2+ + 2.0000 H+ + 3.0000 Br-
log_k 23.5651
-delta_H -149.799 kJ/mol # Calculated enthalpy of reaction UOBr3
# Enthalpy of formation: -954 kJ/mol
-analytic -2.0001e+002 -8.4632e-002 1.1381e+004 8.5102e+001 1.7765e+002
# -Range: 0-300
UOCl
UOCl +2.0000 H+ = + 1.0000 Cl- + 1.0000 H2O + 1.0000 U+++
log_k 10.3872
-delta_H -108.118 kJ/mol # Calculated enthalpy of reaction UOCl
# Enthalpy of formation: -833.9 kJ/mol
-analytic -1.1989e+002 -4.0791e-002 8.0834e+003 4.6600e+001 1.2617e+002
# -Range: 0-300
UOCl2
UOCl2 +2.0000 H+ = + 1.0000 H2O + 1.0000 U++++ + 2.0000 Cl-
log_k 5.4559
-delta_H -141.898 kJ/mol # Calculated enthalpy of reaction UOCl2
# Enthalpy of formation: -1069.3 kJ/mol
-analytic -2.2096e+002 -7.3329e-002 1.1858e+004 8.4250e+001 1.8509e+002
# -Range: 0-300
UOCl3
UOCl3 +1.0000 H2O = + 1.0000 UO2+ + 2.0000 H+ + 3.0000 Cl-
log_k 12.6370
-delta_H -100.528 kJ/mol # Calculated enthalpy of reaction UOCl3
# Enthalpy of formation: -1140 kJ/mol
-analytic -2.1934e+002 -8.8639e-002 9.3198e+003 9.1775e+001 1.4549e+002
# -Range: 0-300
UOF2
UOF2 +2.0000 H+ = + 1.0000 H2O + 1.0000 U++++ + 2.0000 F-
log_k -18.1473
-delta_H -43.1335 kJ/mol # Calculated enthalpy of reaction UOF2
# Enthalpy of formation: -1504.6 kJ/mol
-analytic -6.9471e+001 -2.6188e-002 2.5576e+003 2.0428e+001 4.3454e+001
# -Range: 0-200
UOF2:H2O
UOF2:H2O +2.0000 H+ = + 1.0000 U++++ + 2.0000 F- + 2.0000 H2O
log_k -18.7019
-delta_H -31.5719 kJ/mol # Calculated enthalpy of reaction UOF2:H2O
# Enthalpy of formation: -1802 kJ/mol
-analytic -9.5010e+001 -2.6355e-002 3.1474e+003 2.9746e+001 5.3480e+001
# -Range: 0-200
UOF4
UOF4 +1.0000 H2O = + 1.0000 UO2++ + 2.0000 H+ + 4.0000 F-
log_k 4.5737
-delta_H -149.952 kJ/mol # Calculated enthalpy of reaction UOF4
# Enthalpy of formation: -1924.6 kJ/mol
-analytic -5.9731e+000 -3.8581e-002 4.6903e+003 2.5464e+000 7.9649e+001
# -Range: 0-200
UOFOH
UOFOH +3.0000 H+ = + 1.0000 F- + 1.0000 U++++ + 2.0000 H2O
log_k -8.9274
-delta_H -71.5243 kJ/mol # Calculated enthalpy of reaction UOFOH
# Enthalpy of formation: -1426.7 kJ/mol
-analytic -9.2412e+001 -1.7293e-002 5.8150e+003 2.7940e+001 9.8779e+001
# -Range: 0-200
UOFOH:.5H2O
UOFOH:.5H2O +1.0000 H+ +0.5000 O2 = + 1.0000 F- + 1.0000 UO2++ + 1.5000 H2O
log_k 24.5669
-delta_H -200.938 kJ/mol # Calculated enthalpy of reaction UOFOH:.5H2O
# Enthalpy of formation: -1576.1 kJ/mol
-analytic -1.1024e+001 -7.7180e-003 1.0019e+004 1.7305e+000 1.7014e+002
# -Range: 0-200
UP
UP +2.0000 O2 +1.0000 H+ = + 1.0000 HPO4-- + 1.0000 U+++
log_k 233.4928
-delta_H -1487.11 kJ/mol # Calculated enthalpy of reaction UP
# Enthalpy of formation: -269.8 kJ/mol
-analytic -2.1649e+002 -9.0873e-002 8.3804e+004 8.1649e+001 -5.4044e+005
# -Range: 0-300
UP2
UP2 +3.2500 O2 +1.5000 H2O = + 1.0000 H+ + 1.0000 U+++ + 2.0000 HPO4--
log_k 360.5796
-delta_H -2301.07 kJ/mol # Calculated enthalpy of reaction UP2
# Enthalpy of formation: -304 kJ/mol
-analytic -2.4721e+002 -1.5005e-001 1.2243e+005 9.9521e+001 -3.9706e+005
# -Range: 0-300
UP2O7
UP2O7 +1.0000 H2O = + 1.0000 U++++ + 2.0000 HPO4--
log_k -32.9922
-delta_H -37.5256 kJ/mol # Calculated enthalpy of reaction UP2O7
# Enthalpy of formation: -2852 kJ/mol
-analytic -3.5910e+002 -1.3819e-001 7.6509e+003 1.3804e+002 1.1949e+002
# -Range: 0-300
UP2O7:20H2O
UP2O7:20H2O = + 1.0000 U++++ + 2.0000 HPO4-- + 19.0000 H2O
log_k -28.6300
-delta_H 0 # Not possible to calculate enthalpy of reaction UP2O7:20H2O
# Enthalpy of formation: 0 kcal/mol
UPO5
UPO5 +1.0000 H2O = + 1.0000 H+ + 1.0000 HPO4-- + 1.0000 UO2+
log_k -19.5754
-delta_H 32.6294 kJ/mol # Calculated enthalpy of reaction UPO5
# Enthalpy of formation: -2064 kJ/mol
-analytic -1.5316e+002 -6.0911e-002 7.3255e+002 6.0317e+001 1.1476e+001
# -Range: 0-300
US
US +2.0000 H+ +0.2500 O2 = + 0.5000 H2O + 1.0000 HS- + 1.0000 U+++
log_k 46.6547
-delta_H -322.894 kJ/mol # Calculated enthalpy of reaction US
# Enthalpy of formation: -322.2 kJ/mol
-analytic -1.0845e+002 -4.0538e-002 1.8749e+004 4.2147e+001 2.9259e+002
# -Range: 0-300
US1.9
US1.9 +1.9000 H+ = + 0.2000 U+++ + 0.8000 U++++ + 1.9000 HS-
log_k -2.2816
-delta_H -91.486 kJ/mol # Calculated enthalpy of reaction US1.9
# Enthalpy of formation: -509.9 kJ/mol
-analytic -2.0534e+002 -6.8390e-002 8.8888e+003 7.8243e+001 1.3876e+002
# -Range: 0-300
US2
US2 +2.0000 H+ = + 1.0000 U++++ + 2.0000 HS-
log_k -2.3324
-delta_H -103.017 kJ/mol # Calculated enthalpy of reaction US2
# Enthalpy of formation: -520.4 kJ/mol
-analytic -2.1819e+002 -7.1522e-002 9.7782e+003 8.2586e+001 1.5264e+002
# -Range: 0-300
US3
US3 +2.0000 H2O = + 1.0000 H+ + 1.0000 UO2++ + 3.0000 HS-
log_k -16.6370
-delta_H 43.9515 kJ/mol # Calculated enthalpy of reaction US3
# Enthalpy of formation: -539.6 kJ/mol
-analytic -2.3635e+002 -9.5877e-002 1.9170e+003 9.7726e+001 2.9982e+001
# -Range: 0-300
USb
USb +3.0000 H+ +1.5000 O2 = + 1.0000 Sb(OH)3 + 1.0000 U+++
log_k 176.0723
-delta_H -1106.19 kJ/mol # Calculated enthalpy of reaction USb
# Enthalpy of formation: -138.5 kJ/mol
USb2
USb2 +3.0000 H+ +2.2500 O2 +1.5000 H2O = + 1.0000 U+++ + 2.0000 Sb(OH)3
log_k 223.1358
-delta_H -1407.02 kJ/mol # Calculated enthalpy of reaction USb2
# Enthalpy of formation: -173.6 kJ/mol
Uranium-selenide
1.0USe +1.7500 O2 +1.0000 H+ = + 0.5000 H2O + 1.0000 SeO3-- + 1.0000 U+++
log_k 125.6086
-delta_H -844.278 kJ/mol # Calculated enthalpy of reaction Uranium-selenide
# Enthalpy of formation: -275.7 kJ/mol
-analytic -1.0853e+002 -7.6251e-002 4.3230e+004 4.5189e+001 6.7460e+002
# -Range: 0-300
USe2(alpha)
USe2 +2.7500 O2 +0.5000 H2O = + 1.0000 H+ + 1.0000 U+++ + 2.0000 SeO3--
log_k 125.4445
-delta_H -904.199 kJ/mol # Calculated enthalpy of reaction USe2(alpha)
# Enthalpy of formation: -427 kJ/mol
-analytic -2.0454e+002 -1.4191e-001 4.6114e+004 8.7906e+001 7.1963e+002
# -Range: 0-300
USe2(beta)
USe2 +2.7500 O2 +0.5000 H2O = + 1.0000 H+ + 1.0000 U+++ + 2.0000 SeO3--
log_k 125.2868
-delta_H -904.199 kJ/mol # Calculated enthalpy of reaction USe2(beta)
# Enthalpy of formation: -427 kJ/mol
-analytic -2.0334e+002 -1.4147e-001 4.6082e+004 8.7349e+001 7.1913e+002
# -Range: 0-300
USe3
USe3 +3.7500 O2 +1.5000 H2O = + 1.0000 U+++ + 3.0000 H+ + 3.0000 SeO3--
log_k 147.2214
-delta_H -1090.42 kJ/mol # Calculated enthalpy of reaction USe3
# Enthalpy of formation: -452 kJ/mol
-analytic 4.9201e+002 -1.3720e-002 3.2168e+004 -1.8131e+002 5.4609e+002
# -Range: 0-200
Umangite
Cu3Se2 = + 1.0000 Cu++ + 2.0000 Cu+ + 2.0000 Se--
log_k -93.8412
-delta_H 0 # Not possible to calculate enthalpy of reaction Umangite
# Enthalpy of formation: -25 kcal/mol
-analytic -7.2308e+001 -2.2566e-003 -2.0738e+004 1.9677e+001 -3.5214e+002
# -Range: 0-200
Uraninite
UO2 +4.0000 H+ = + 1.0000 U++++ + 2.0000 H2O
log_k -4.8372
-delta_H -77.8767 kJ/mol # Calculated enthalpy of reaction Uraninite
# Enthalpy of formation: -1085 kJ/mol
-analytic -7.5776e+001 -1.0558e-002 5.9677e+003 2.1853e+001 9.3142e+001
# -Range: 0-300
Uranocircite
Ba(UO2)2(PO4)2 +2.0000 H+ = + 1.0000 Ba++ + 2.0000 HPO4-- + 2.0000 UO2++
log_k -19.8057
-delta_H -72.3317 kJ/mol # Calculated enthalpy of reaction Uranocircite
# Enthalpy of formation: -1215.94 kcal/mol
-analytic -3.6843e+001 -4.3076e-002 1.2427e+003 1.0384e+001 2.1115e+001
# -Range: 0-200
Uranophane
Ca(UO2)2(SiO3)2(OH)2 +6.0000 H+ = + 1.0000 Ca++ + 2.0000 SiO2 + 2.0000 UO2++ + 4.0000 H2O
log_k 17.2850
-delta_H 0 # Not possible to calculate enthalpy of reaction Uranophane
# Enthalpy of formation: 0 kcal/mol
V
V +3.0000 H+ +0.7500 O2 = + 1.0000 V+++ + 1.5000 H2O
log_k 106.9435
-delta_H -680.697 kJ/mol # Calculated enthalpy of reaction V
# Enthalpy of formation: 0 kJ/mol
-analytic -1.0508e+002 -2.1334e-002 4.0364e+004 3.5012e+001 -3.2290e+005
# -Range: 0-300
V2O4
V2O4 +4.0000 H+ = + 2.0000 H2O + 2.0000 VO++
log_k 8.5719
-delta_H -117.564 kJ/mol # Calculated enthalpy of reaction V2O4
# Enthalpy of formation: -1427.31 kJ/mol
-analytic -1.4429e+002 -3.7423e-002 9.7046e+003 5.3125e+001 1.5147e+002
# -Range: 0-300
V3O5
V3O5 +8.0000 H+ = + 1.0000 VO++ + 2.0000 V+++ + 4.0000 H2O
log_k 13.4312
-delta_H -218.857 kJ/mol # Calculated enthalpy of reaction V3O5
# Enthalpy of formation: -1933.17 kJ/mol
-analytic -1.7652e+002 -2.1959e-002 1.6814e+004 5.6618e+001 2.8559e+002
# -Range: 0-200
V4O7
V4O7 +10.0000 H+ = + 2.0000 V+++ + 2.0000 VO++ + 5.0000 H2O
log_k 18.7946
-delta_H -284.907 kJ/mol # Calculated enthalpy of reaction V4O7
# Enthalpy of formation: -2639.56 kJ/mol
-analytic -2.2602e+002 -3.0261e-002 2.1667e+004 7.3214e+001 3.6800e+002
# -Range: 0-200
Vaesite
NiS2 +1.0000 H2O = + 0.2500 H+ + 0.2500 SO4-- + 1.0000 Ni++ + 1.7500 HS-
log_k -26.7622
-delta_H 110.443 kJ/mol # Calculated enthalpy of reaction Vaesite
# Enthalpy of formation: -32.067 kcal/mol
-analytic 1.6172e+001 -2.2673e-002 -8.2514e+003 -3.4392e+000 -1.4013e+002
# -Range: 0-200
Vivianite
Fe3(PO4)2:8H2O +2.0000 H+ = + 2.0000 HPO4-- + 3.0000 Fe++ + 8.0000 H2O
log_k -4.7237
-delta_H 0 # Not possible to calculate enthalpy of reaction Vivianite
# Enthalpy of formation: 0 kcal/mol
W
W +1.5000 O2 +1.0000 H2O = + 1.0000 WO4-- + 2.0000 H+
log_k 123.4334
-delta_H -771.668 kJ/mol # Calculated enthalpy of reaction W
# Enthalpy of formation: 0 kJ/mol
-analytic -1.0433e+002 -6.9470e-002 4.0134e+004 4.5993e+001 6.2629e+002
# -Range: 0-300
Wairakite
CaAl2Si4O10(OH)4 +8.0000 H+ = + 1.0000 Ca++ + 2.0000 Al+++ + 4.0000 SiO2 + 6.0000 H2O
log_k 18.0762
-delta_H -237.781 kJ/mol # Calculated enthalpy of reaction Wairakite
# Enthalpy of formation: -1579.33 kcal/mol
-analytic -1.7914e+001 3.2944e-003 2.2782e+004 -9.0981e+000 -1.6934e+006
# -Range: 0-300
Weeksite
K2(UO2)2(Si2O5)3:4H2O +6.0000 H+ = + 2.0000 K+ + 2.0000 UO2++ + 6.0000 SiO2 + 7.0000 H2O
log_k 15.3750
-delta_H 0 # Not possible to calculate enthalpy of reaction Weeksite
# Enthalpy of formation: 0 kcal/mol
Whitlockite
Ca3(PO4)2 +2.0000 H+ = + 2.0000 HPO4-- + 3.0000 Ca++
log_k -4.2249
-delta_H -116.645 kJ/mol # Calculated enthalpy of reaction Whitlockite
# Enthalpy of formation: -4096.77 kJ/mol
-analytic -5.3543e+002 -1.8842e-001 1.7176e+004 2.1406e+002 2.6817e+002
# -Range: 0-300
Wilkmanite
Ni3Se4 +1.0000 H2O = + 0.5000 O2 + 2.0000 H+ + 3.0000 Ni++ + 4.0000 Se--
log_k -152.8793
-delta_H 0 # Not possible to calculate enthalpy of reaction Wilkmanite
# Enthalpy of formation: -60.285 kcal/mol
-analytic -1.9769e+002 -4.9968e-002 -2.8208e+004 6.2863e+001 -1.1322e+005
# -Range: 0-300
Witherite
BaCO3 +1.0000 H+ = + 1.0000 Ba++ + 1.0000 HCO3-
log_k -2.9965
-delta_H 17.1628 kJ/mol # Calculated enthalpy of reaction Witherite
# Enthalpy of formation: -297.5 kcal/mol
-analytic -1.2585e+002 -4.4315e-002 2.0227e+003 5.2239e+001 3.1600e+001
# -Range: 0-300
Wollastonite
CaSiO3 +2.0000 H+ = + 1.0000 Ca++ + 1.0000 H2O + 1.0000 SiO2
log_k 13.7605
-delta_H -76.5756 kJ/mol # Calculated enthalpy of reaction Wollastonite
# Enthalpy of formation: -389.59 kcal/mol
-analytic 3.0931e+001 6.7466e-003 5.1749e+003 -1.3209e+001 -3.4579e+005
# -Range: 0-300
Wurtzite
ZnS +1.0000 H+ = + 1.0000 HS- + 1.0000 Zn++
log_k -9.1406
-delta_H 22.3426 kJ/mol # Calculated enthalpy of reaction Wurtzite
# Enthalpy of formation: -45.85 kcal/mol
-analytic -1.5446e+002 -4.8874e-002 2.4551e+003 6.1278e+001 3.8355e+001
# -Range: 0-300
Wustite
Fe.947O +2.0000 H+ = + 0.1060 Fe+++ + 0.8410 Fe++ + 1.0000 H2O
log_k 12.4113
-delta_H -102.417 kJ/mol # Calculated enthalpy of reaction Wustite
# Enthalpy of formation: -266.265 kJ/mol
-analytic -7.6919e+001 -1.8433e-002 7.3823e+003 2.8312e+001 1.1522e+002
# -Range: 0-300
Xonotlite
Ca6Si6O17(OH)2 +12.0000 H+ = + 6.0000 Ca++ + 6.0000 SiO2 + 7.0000 H2O
log_k 91.8267
-delta_H -495.457 kJ/mol # Calculated enthalpy of reaction Xonotlite
# Enthalpy of formation: -2397.25 kcal/mol
-analytic 1.6080e+003 3.7309e-001 -2.2548e+004 -6.2716e+002 -3.8346e+002
# -Range: 0-200
Y
Y +3.0000 H+ +0.7500 O2 = + 1.0000 Y+++ + 1.5000 H2O
log_k 184.5689
-delta_H -1134.7 kJ/mol # Calculated enthalpy of reaction Y
# Enthalpy of formation: 0 kJ/mol
-analytic -6.2641e+001 -2.8062e-002 5.9667e+004 2.2394e+001 9.3107e+002
# -Range: 0-300
Yb
Yb +2.0000 H+ +0.5000 O2 = + 1.0000 H2O + 1.0000 Yb++
log_k 137.1930
-delta_H -810.303 kJ/mol # Calculated enthalpy of reaction Yb
# Enthalpy of formation: 0 kJ/mol
-analytic -7.4712e+001 -2.0993e-002 4.4129e+004 2.8341e+001 6.8862e+002
# -Range: 0-300
Yb(OH)3
Yb(OH)3 +3.0000 H+ = + 1.0000 Yb+++ + 3.0000 H2O
log_k 14.6852
-delta_H 0 # Not possible to calculate enthalpy of reaction Yb(OH)3
# Enthalpy of formation: 0 kcal/mol
Yb(OH)3(am)
Yb(OH)3 +3.0000 H+ = + 1.0000 Yb+++ + 3.0000 H2O
log_k 18.9852
-delta_H 0 # Not possible to calculate enthalpy of reaction Yb(OH)3(am)
# Enthalpy of formation: 0 kcal/mol
Yb2(CO3)3
Yb2(CO3)3 +3.0000 H+ = + 2.0000 Yb+++ + 3.0000 HCO3-
log_k -2.3136
-delta_H 0 # Not possible to calculate enthalpy of reaction Yb2(CO3)3
# Enthalpy of formation: 0 kcal/mol
Yb2O3
Yb2O3 +6.0000 H+ = + 2.0000 Yb+++ + 3.0000 H2O
log_k 47.8000
-delta_H 0 # Not possible to calculate enthalpy of reaction Yb2O3
# Enthalpy of formation: 0 kcal/mol
YbF3:.5H2O
YbF3:.5H2O = + 0.5000 H2O + 1.0000 Yb+++ + 3.0000 F-
log_k -16.0000
-delta_H 0 # Not possible to calculate enthalpy of reaction YbF3:.5H2O
# Enthalpy of formation: 0 kcal/mol
YbPO4:10H2O
YbPO4:10H2O +1.0000 H+ = + 1.0000 HPO4-- + 1.0000 Yb+++ + 10.0000 H2O
log_k -11.7782
-delta_H 0 # Not possible to calculate enthalpy of reaction YbPO4:10H2O
# Enthalpy of formation: 0 kcal/mol
Zincite
ZnO +2.0000 H+ = + 1.0000 H2O + 1.0000 Zn++
log_k 11.2087
-delta_H -88.7638 kJ/mol # Calculated enthalpy of reaction Zincite
# Enthalpy of formation: -350.46 kJ/mol
-analytic -8.6681e+001 -1.9324e-002 7.1034e+003 3.2256e+001 1.1087e+002
# -Range: 0-300
Zircon
ZrSiO4 +2.0000 H+ = + 1.0000 SiO2 + 1.0000 Zr(OH)2++
log_k -15.4193
-delta_H 64.8635 kJ/mol # Calculated enthalpy of reaction Zircon
# Enthalpy of formation: -2033.4 kJ/mol
-analytic 9.2639e+000 6.5416e-003 5.0759e+002 -8.4547e+000 -6.6155e+005
# -Range: 0-300
Zn
Zn +2.0000 H+ +0.5000 O2 = + 1.0000 H2O + 1.0000 Zn++
log_k 68.8035
-delta_H -433.157 kJ/mol # Calculated enthalpy of reaction Zn
# Enthalpy of formation: 0 kJ/mol
-analytic -6.4131e+001 -2.0009e-002 2.3921e+004 2.3702e+001 3.7329e+002
# -Range: 0-300
Zn(BO2)2
Zn(BO2)2 +2.0000 H+ +2.0000 H2O = + 1.0000 Zn++ + 2.0000 B(OH)3
log_k 8.3130
-delta_H 0 # Not possible to calculate enthalpy of reaction Zn(BO2)2
# Enthalpy of formation: 0 kcal/mol
Zn(ClO4)2:6H2O
Zn(ClO4)2:6H2O = + 1.0000 Zn++ + 2.0000 ClO4- + 6.0000 H2O
log_k 5.6474
-delta_H 6.31871 kJ/mol # Calculated enthalpy of reaction Zn(ClO4)2:6H2O
# Enthalpy of formation: -2133.39 kJ/mol
-analytic -1.8191e+002 -9.1383e-003 7.4822e+003 6.6751e+001 1.2712e+002
# -Range: 0-200
Zn(IO3)2
Zn(IO3)2 = + 1.0000 Zn++ + 2.0000 IO3-
log_k -5.3193
-delta_H 0 # Not possible to calculate enthalpy of reaction Zn(IO3)2
# Enthalpy of formation: 0 kcal/mol
Zn(NO3)2:6H2O
Zn(NO3)2:6H2O = + 1.0000 Zn++ + 2.0000 NO3- + 6.0000 H2O
log_k 3.4102
-delta_H 24.7577 kJ/mol # Calculated enthalpy of reaction Zn(NO3)2:6H2O
# Enthalpy of formation: -2306.8 kJ/mol
-analytic -1.7152e+002 -1.6875e-002 5.6291e+003 6.5094e+001 9.5649e+001
# -Range: 0-200
Zn(OH)2(beta)
Zn(OH)2 +2.0000 H+ = + 1.0000 Zn++ + 2.0000 H2O
log_k 11.9341
-delta_H -83.2111 kJ/mol # Calculated enthalpy of reaction Zn(OH)2(beta)
# Enthalpy of formation: -641.851 kJ/mol
-analytic -7.7810e+001 -7.8548e-003 7.1994e+003 2.7455e+001 1.2228e+002
# -Range: 0-200
Zn(OH)2(epsilon)
Zn(OH)2 +2.0000 H+ = + 1.0000 Zn++ + 2.0000 H2O
log_k 11.6625
-delta_H -81.7811 kJ/mol # Calculated enthalpy of reaction Zn(OH)2(epsilon)
# Enthalpy of formation: -643.281 kJ/mol
-analytic -7.7938e+001 -7.8767e-003 7.1282e+003 2.7496e+001 1.2107e+002
# -Range: 0-200
Zn(OH)2(gamma)
Zn(OH)2 +2.0000 H+ = + 1.0000 Zn++ + 2.0000 H2O
log_k 11.8832
-delta_H 0 # Not possible to calculate enthalpy of reaction Zn(OH)2(gamma)
# Enthalpy of formation: 0 kcal/mol
Zn2(OH)3Cl
Zn2(OH)3Cl +3.0000 H+ = + 1.0000 Cl- + 2.0000 Zn++ + 3.0000 H2O
log_k 15.2921
-delta_H 0 # Not possible to calculate enthalpy of reaction Zn2(OH)3Cl
# Enthalpy of formation: 0 kcal/mol
Zn2SO4(OH)2
Zn2SO4(OH)2 +2.0000 H+ = + 1.0000 SO4-- + 2.0000 H2O + 2.0000 Zn++
log_k 7.5816
-delta_H 0 # Not possible to calculate enthalpy of reaction Zn2SO4(OH)2
# Enthalpy of formation: 0 kcal/mol
Zn2SiO4
Zn2SiO4 +4.0000 H+ = + 1.0000 SiO2 + 2.0000 H2O + 2.0000 Zn++
log_k 13.8695
-delta_H -119.399 kJ/mol # Calculated enthalpy of reaction Zn2SiO4
# Enthalpy of formation: -1636.75 kJ/mol
-analytic 2.0970e+002 5.3663e-002 -1.2724e+002 -8.5445e+001 -2.2336e+000
# -Range: 0-200
Zn2TiO4
Zn2TiO4 +4.0000 H+ = + 1.0000 Ti(OH)4 + 2.0000 Zn++
log_k 12.3273
-delta_H 0 # Not possible to calculate enthalpy of reaction Zn2TiO4
# Enthalpy of formation: -1647.85 kJ/mol
Zn3(AsO4)2
Zn3(AsO4)2 +4.0000 H+ = + 2.0000 H2AsO4- + 3.0000 Zn++
log_k 9.3122
-delta_H 0 # Not possible to calculate enthalpy of reaction Zn3(AsO4)2
# Enthalpy of formation: 0 kcal/mol
Zn3O(SO4)2
Zn3O(SO4)2 +2.0000 H+ = + 1.0000 H2O + 2.0000 SO4-- + 3.0000 Zn++
log_k 19.1188
-delta_H -258.253 kJ/mol # Calculated enthalpy of reaction Zn3O(SO4)2
# Enthalpy of formation: -2306.95 kJ/mol
-analytic -3.9661e+001 -4.3860e-002 1.1301e+004 1.3709e+001 1.9193e+002
# -Range: 0-200
Zn5(NO3)2(OH)8
Zn5(NO3)2(OH)8 +8.0000 H+ = + 2.0000 NO3- + 5.0000 Zn++ + 8.0000 H2O
log_k 42.6674
-delta_H 0 # Not possible to calculate enthalpy of reaction Zn5(NO3)2(OH)8
# Enthalpy of formation: 0 kcal/mol
ZnBr2
ZnBr2 = + 1.0000 Zn++ + 2.0000 Br-
log_k 7.5787
-delta_H -67.7622 kJ/mol # Calculated enthalpy of reaction ZnBr2
# Enthalpy of formation: -328.63 kJ/mol
-analytic 6.5789e-002 -2.1477e-002 1.9840e+003 2.9302e+000 3.3691e+001
# -Range: 0-200
ZnBr2:2H2O
ZnBr2:2H2O = + 1.0000 Zn++ + 2.0000 Br- + 2.0000 H2O
log_k 5.2999
-delta_H -30.9268 kJ/mol # Calculated enthalpy of reaction ZnBr2:2H2O
# Enthalpy of formation: -937.142 kJ/mol
-analytic -4.9260e+001 -2.1682e-002 2.4325e+003 2.1360e+001 4.1324e+001
# -Range: 0-200
ZnCO3:H2O
ZnCO3:H2O +1.0000 H+ = + 1.0000 H2O + 1.0000 HCO3- + 1.0000 Zn++
log_k 0.1398
-delta_H 0 # Not possible to calculate enthalpy of reaction ZnCO3:H2O
# Enthalpy of formation: 0 kcal/mol
ZnCl2
ZnCl2 = + 1.0000 Zn++ + 2.0000 Cl-
log_k 7.0880
-delta_H -72.4548 kJ/mol # Calculated enthalpy of reaction ZnCl2
# Enthalpy of formation: -415.09 kJ/mol
-analytic -1.6157e+001 -2.5405e-002 2.6505e+003 8.8584e+000 4.5015e+001
# -Range: 0-200
ZnCl2(NH3)2
ZnCl2(NH3)2 = + 1.0000 Zn++ + 2.0000 Cl- + 2.0000 NH3
log_k -6.9956
-delta_H 27.2083 kJ/mol # Calculated enthalpy of reaction ZnCl2(NH3)2
# Enthalpy of formation: -677.427 kJ/mol
-analytic -5.9409e+001 -2.2698e-002 -2.9178e+002 2.4308e+001 -4.9341e+000
# -Range: 0-200
ZnCl2(NH3)4
ZnCl2(NH3)4 = + 1.0000 Zn++ + 2.0000 Cl- + 4.0000 NH3
log_k -6.6955
-delta_H 56.2004 kJ/mol # Calculated enthalpy of reaction ZnCl2(NH3)4
# Enthalpy of formation: -869.093 kJ/mol
-analytic -9.9769e+001 -1.9793e-002 4.2916e+002 3.9412e+001 7.3223e+000
# -Range: 0-200
ZnCl2(NH3)6
ZnCl2(NH3)6 = + 1.0000 Zn++ + 2.0000 Cl- + 6.0000 NH3
log_k -4.7311
-delta_H 77.4225 kJ/mol # Calculated enthalpy of reaction ZnCl2(NH3)6
# Enthalpy of formation: -1052.99 kJ/mol
-analytic -1.3984e+002 -1.6896e-002 1.5559e+003 5.4524e+001 2.6470e+001
# -Range: 0-200
ZnCr2O4
ZnCr2O4 +8.0000 H+ = + 1.0000 Zn++ + 2.0000 Cr+++ + 4.0000 H2O
log_k 7.9161
-delta_H -221.953 kJ/mol # Calculated enthalpy of reaction ZnCr2O4
# Enthalpy of formation: -370.88 kcal/mol
-analytic -1.7603e+002 -1.0217e-002 1.7414e+004 5.1966e+001 2.9577e+002
# -Range: 0-200
ZnF2
ZnF2 = + 1.0000 Zn++ + 2.0000 F-
log_k -0.4418
-delta_H -59.8746 kJ/mol # Calculated enthalpy of reaction ZnF2
# Enthalpy of formation: -764.206 kJ/mol
-analytic -2.6085e+002 -8.4594e-002 9.0240e+003 1.0318e+002 1.4089e+002
# -Range: 0-300
ZnI2
ZnI2 = + 1.0000 Zn++ + 2.0000 I-
log_k 7.3885
-delta_H -59.2332 kJ/mol # Calculated enthalpy of reaction ZnI2
# Enthalpy of formation: -207.957 kJ/mol
-analytic -1.6472e+001 -2.5573e-002 2.0796e+003 9.9013e+000 3.5320e+001
# -Range: 0-200
ZnSO4
ZnSO4 = + 1.0000 SO4-- + 1.0000 Zn++
log_k 3.5452
-delta_H -80.132 kJ/mol # Calculated enthalpy of reaction ZnSO4
# Enthalpy of formation: -982.855 kJ/mol
-analytic 6.9905e+000 -1.8046e-002 2.2566e+003 -2.2819e+000 3.8318e+001
# -Range: 0-200
ZnSO4:6H2O
ZnSO4:6H2O = + 1.0000 SO4-- + 1.0000 Zn++ + 6.0000 H2O
log_k -1.6846
-delta_H -0.412008 kJ/mol # Calculated enthalpy of reaction ZnSO4:6H2O
# Enthalpy of formation: -2777.61 kJ/mol
-analytic -1.4506e+002 -1.8736e-002 5.2179e+003 5.3121e+001 8.8657e+001
# -Range: 0-200
ZnSO4:7H2O
ZnSO4:7H2O = + 1.0000 SO4-- + 1.0000 Zn++ + 7.0000 H2O
log_k -1.8683
-delta_H 14.0417 kJ/mol # Calculated enthalpy of reaction ZnSO4:7H2O
# Enthalpy of formation: -3077.9 kJ/mol
-analytic -1.6943e+002 -1.8833e-002 5.6484e+003 6.2326e+001 9.5975e+001
# -Range: 0-200
ZnSO4:H2O
ZnSO4:H2O = + 1.0000 H2O + 1.0000 SO4-- + 1.0000 Zn++
log_k -0.5383
-delta_H -44.2824 kJ/mol # Calculated enthalpy of reaction ZnSO4:H2O
# Enthalpy of formation: -1304.54 kJ/mol
-analytic -1.7908e+001 -1.8228e-002 1.5811e+003 7.0677e+000 2.6856e+001
# -Range: 0-200
ZnSeO3:H2O
ZnSeO3:H2O = + 1.0000 H2O + 1.0000 SeO3-- + 1.0000 Zn++
log_k -6.7408
-delta_H -17.9056 kJ/mol # Calculated enthalpy of reaction ZnSeO3:H2O
# Enthalpy of formation: -930.511 kJ/mol
-analytic -1.8569e+001 -1.9929e-002 6.4377e+001 7.0892e+000 1.0996e+000
# -Range: 0-200
Zoisite
Ca2Al3(SiO4)3OH +13.0000 H+ = + 2.0000 Ca++ + 3.0000 Al+++ + 3.0000 SiO2 + 7.0000 H2O
log_k 43.3017
-delta_H -458.131 kJ/mol # Calculated enthalpy of reaction Zoisite
# Enthalpy of formation: -1643.69 kcal/mol
-analytic 2.5321e+000 -3.5886e-002 1.9902e+004 -6.2443e+000 3.1055e+002
# -Range: 0-300
Zr
Zr +2.0000 H+ +1.0000 O2 = + 1.0000 Zr(OH)2++
log_k 177.6471
-delta_H -1078.71 kJ/mol # Calculated enthalpy of reaction Zr
# Enthalpy of formation: 0 kJ/mol
-analytic -2.8360e+001 -1.5214e-002 5.8045e+004 7.8012e+000 -3.0657e+005
# -Range: 0-300
ZrB2
ZrB2 +3.0000 H+ +2.0000 H2O +0.5000 O2 = + 1.0000 B(OH)3 + 1.0000 BH4- + 1.0000 Zr++++
log_k 103.4666
-delta_H 0 # Not possible to calculate enthalpy of reaction ZrB2
# Enthalpy of formation: -326.628 kJ/mol
ZrC
ZrC +3.0000 H+ +2.0000 O2 = + 1.0000 H2O + 1.0000 HCO3- + 1.0000 Zr++++
log_k 207.0906
-delta_H 0 # Not possible to calculate enthalpy of reaction ZrC
# Enthalpy of formation: -203.008 kJ/mol
ZrCl
ZrCl +3.0000 H+ +0.7500 O2 = + 1.0000 Cl- + 1.0000 Zr++++ + 1.5000 H2O
log_k 130.9450
-delta_H 0 # Not possible to calculate enthalpy of reaction ZrCl
# Enthalpy of formation: -303.211 kJ/mol
ZrCl2
ZrCl2 +2.0000 H+ +0.5000 O2 = + 1.0000 H2O + 1.0000 Zr++++ + 2.0000 Cl-
log_k 96.3205
-delta_H 0 # Not possible to calculate enthalpy of reaction ZrCl2
# Enthalpy of formation: -531.021 kJ/mol
ZrCl3
ZrCl3 +1.0000 H+ +0.2500 O2 = + 0.5000 H2O + 1.0000 Zr++++ + 3.0000 Cl-
log_k 62.4492
-delta_H 0 # Not possible to calculate enthalpy of reaction ZrCl3
# Enthalpy of formation: -754.997 kJ/mol
ZrCl4
ZrCl4 = + 1.0000 Zr++++ + 4.0000 Cl-
log_k 27.9824
-delta_H 0 # Not possible to calculate enthalpy of reaction ZrCl4
# Enthalpy of formation: -980.762 kJ/mol
ZrF4(beta)
ZrF4 = + 1.0000 Zr++++ + 4.0000 F-
log_k -27.7564
-delta_H 0 # Not possible to calculate enthalpy of reaction ZrF4(beta)
# Enthalpy of formation: -1911.26 kJ/mol
ZrH2
ZrH2 +4.0000 H+ +1.5000 O2 = + 1.0000 Zr++++ + 3.0000 H2O
log_k 198.3224
-delta_H 0 # Not possible to calculate enthalpy of reaction ZrH2
# Enthalpy of formation: -168.946 kJ/mol
ZrN
ZrN +4.0000 H+ +0.2500 O2 = + 0.5000 H2O + 1.0000 NH3 + 1.0000 Zr++++
log_k 59.1271
-delta_H 0 # Not possible to calculate enthalpy of reaction ZrN
# Enthalpy of formation: -365 kJ/mol
O-phthalic_acid
H2O_phthalate = + 1.0000 O_phthalate-2 + 2.0000 H+
log_k -9.7755
-delta_H 0 # Not possible to calculate enthalpy of reaction O-phthalic_acid
# Enthalpy of formation: -186.88 kJ/mol
-analytic 7.3450e+001 1.9477e-002 -3.6511e+003 -3.1035e+001 -6.2027e+001
# -Range: 0-200
Br2(l)
Br2 +1.0000 H2O = + 0.5000 O2 + 2.0000 Br- + 2.0000 H+
log_k -6.5419
-delta_H 36.7648 kJ/mol # Calculated enthalpy of reaction Br2(l)
# Enthalpy of formation: 0 kJ/mol
-analytic -1.5875e+002 -5.8039e-002 1.5583e+003 6.6381e+001 2.4362e+001
# -Range: 0-300
Hg(l)
Hg +2.0000 H+ +0.5000 O2 = + 1.0000 H2O + 1.0000 Hg++
log_k 14.1505
-delta_H -109.608 kJ/mol # Calculated enthalpy of reaction Hg(l)
# Enthalpy of formation: 0 kcal/mol
-analytic -6.6462e+001 -1.8504e-002 7.3141e+003 2.4888e+001 1.1415e+002
# -Range: 0-300
Ag(g)
Ag +1.0000 H+ +0.2500 O2 = + 0.5000 H2O + 1.0000 Ag+
log_k 51.0924
-delta_H -319.035 kJ/mol # Calculated enthalpy of reaction Ag(g)
# Enthalpy of formation: 284.9 kJ/mol
-analytic -5.8006e+000 1.7178e-003 1.6809e+004 0.0000e+000 0.0000e+000
# -Range: 0-200
Al(g)
Al +3.0000 H+ +0.7500 O2 = + 1.0000 Al+++ + 1.5000 H2O
log_k 200.6258
-delta_H -1288.06 kJ/mol # Calculated enthalpy of reaction Al(g)
# Enthalpy of formation: 330 kJ/mol
-analytic 9.6402e+000 -6.9301e-003 6.5270e+004 -1.0461e+001 1.1084e+003
# -Range: 0-200
Am(g)
Am +3.0000 H+ +0.7500 O2 = + 1.0000 Am+++ + 1.5000 H2O
log_k 211.7865
-delta_H -1320.16 kJ/mol # Calculated enthalpy of reaction Am(g)
# Enthalpy of formation: 283.8 kJ/mol
-analytic -1.4236e+001 -8.7560e-003 6.8166e+004 0.0000e+000 0.0000e+000
# -Range: 0-300
AmF3(g)
AmF3 = + 1.0000 Am+++ + 3.0000 F-
log_k 49.8631
-delta_H -455.843 kJ/mol # Calculated enthalpy of reaction AmF3(g)
# Enthalpy of formation: -1166.9 kJ/mol
-analytic -4.7209e+001 -3.6440e-002 2.2278e+004 1.3418e+001 3.7833e+002
# -Range: 0-200
Ar(g)
Ar = + 1.0000 Ar
log_k -2.8587
-delta_H -12.0081 kJ/mol # Calculated enthalpy of reaction Ar(g)
# Enthalpy of formation: 0 kcal/mol
-analytic -7.4387e+000 7.8991e-003 0.0000e+000 0.0000e+000 1.9830e+005
# -Range: 0-300
B(g)
B +1.5000 H2O +0.7500 O2 = + 1.0000 B(OH)3
log_k 200.8430
-delta_H -1201.68 kJ/mol # Calculated enthalpy of reaction B(g)
# Enthalpy of formation: 565 kJ/mol
-analytic 1.0834e+002 1.0606e-002 5.8150e+004 -4.2720e+001 9.8743e+002
# -Range: 0-200
BF3(g)
BF3 +3.0000 H2O = + 1.0000 B(OH)3 + 3.0000 F- + 3.0000 H+
log_k -2.9664
-delta_H -87.0627 kJ/mol # Calculated enthalpy of reaction BF3(g)
# Enthalpy of formation: -1136 kJ/mol
-analytic 5.2848e+001 -2.4617e-002 -1.8159e+002 -1.9350e+001 -3.1018e+000
# -Range: 0-200
Be(g)
Be +2.0000 H+ +0.5000 O2 = + 1.0000 Be++ + 1.0000 H2O
log_k 361.9343
-delta_H 0 # Not possible to calculate enthalpy of reaction Be(g)
# Enthalpy of formation: 0 kcal/mol
Br2(g)
Br2 +1.0000 H2O = + 0.5000 O2 + 2.0000 Br- + 2.0000 H+
log_k -5.9979
-delta_H 5.85481 kJ/mol # Calculated enthalpy of reaction Br2(g)
# Enthalpy of formation: 30.91 kJ/mol
-analytic -3.2403e+000 -1.7609e-002 -1.4941e+003 3.0300e+000 -2.5370e+001
# -Range: 0-200
C(g)
C +1.0000 H2O +1.0000 O2 = + 1.0000 H+ + 1.0000 HCO3-
log_k 181.7723
-delta_H -1108.64 kJ/mol # Calculated enthalpy of reaction C(g)
# Enthalpy of formation: 716.68 kJ/mol
-analytic 1.0485e+002 1.7907e-003 5.2768e+004 -4.0661e+001 8.9605e+002
# -Range: 0-200
Ethylene(g)
Ethylene = + 1.0000 Ethylene
log_k -2.3236
-delta_H -16.4431 kJ/mol # Calculated enthalpy of reaction Ethylene(g)
# Enthalpy of formation: 12.5 kcal/mol
-analytic -7.5368e+000 8.4676e-003 0.0000e+000 0.0000e+000 2.3971e+005
# -Range: 0-300
CH4(g)
CH4 = + 1.0000 CH4
log_k -2.8502
-delta_H -13.0959 kJ/mol # Calculated enthalpy of reaction CH4(g)
# Enthalpy of formation: -17.88 kcal/mol
-analytic -2.4027e+001 4.7146e-003 3.7227e+002 6.4264e+000 2.3362e+005
# -Range: 0-300
CO(g)
# CO +1.0000 H2O +0.5000 O2 = + 1.0000 H+ + 1.0000 HCO3-
# log_k 38.6934
# -analytic -6.1217e+001 -3.1388e-002 1.5283e+004 2.3433e+001 2.3850e+002
# -Range: 0-300
CO = CO
log_k -3.0068
-delta_H -10.4349 kJ/mol # Calculated enthalpy of reaction CO(g)
# Enthalpy of formation: -26.416 kcal/mol
-analytic -8.0849e+000 9.2114e-003 0.0000e+000 0.0000e+000 2.0813e+005
# -Range: 0-300
CO2(g)
CO2 +1.0000 H2O = + 1.0000 H+ + 1.0000 HCO3-
log_k -7.8136
-delta_H -10.5855 kJ/mol # Calculated enthalpy of reaction CO2(g)
# Enthalpy of formation: -94.051 kcal/mol
-analytic -8.5938e+001 -3.0431e-002 2.0702e+003 3.2427e+001 3.2328e+001
# -Range: 0-300
Ca(g)
Ca +2.0000 H+ +0.5000 O2 = + 1.0000 Ca++ + 1.0000 H2O
log_k 165.0778
-delta_H -1000.65 kJ/mol # Calculated enthalpy of reaction Ca(g)
# Enthalpy of formation: 177.8 kJ/mol
-analytic -7.3029e+000 -4.8208e-003 5.1822e+004 0.0000e+000 0.0000e+000
# -Range: 0-200
Cd(g)
Cd +2.0000 H+ +0.5000 O2 = + 1.0000 Cd++ + 1.0000 H2O
log_k 70.1363
-delta_H -467.469 kJ/mol # Calculated enthalpy of reaction Cd(g)
# Enthalpy of formation: 111.8 kJ/mol
-analytic -9.8665e+000 -3.0921e-003 2.4126e+004 0.0000e+000 0.0000e+000
# -Range: 0-200
Cl2(g)
Cl2 +1.0000 H2O = + 0.5000 O2 + 2.0000 Cl- + 2.0000 H+
log_k 3.0004
-delta_H -54.3878 kJ/mol # Calculated enthalpy of reaction Cl2(g)
# Enthalpy of formation: 0 kJ/mol
-analytic -1.9456e+001 -2.1491e-002 2.0652e+003 8.8629e+000 3.5076e+001
# -Range: 0-200
Cs(g)
Cs +1.0000 H+ +0.2500 O2 = + 0.5000 H2O + 1.0000 Cs+
log_k 81.2805
-delta_H -474.413 kJ/mol # Calculated enthalpy of reaction Cs(g)
# Enthalpy of formation: 76.5 kJ/mol
-analytic 4.1676e+001 9.1952e-003 2.3401e+004 -1.6824e+001 3.9736e+002
# -Range: 0-200
Cu(g)
Cu +2.0000 H+ +0.5000 O2 = + 1.0000 Cu++ + 1.0000 H2O
log_k 83.6618
-delta_H -551.483 kJ/mol # Calculated enthalpy of reaction Cu(g)
# Enthalpy of formation: 337.4 kJ/mol
-analytic -1.1249e+001 -2.7585e-003 2.8541e+004 0.0000e+000 0.0000e+000
# -Range: 0-200
F2(g)
F2 +1.0000 H2O = + 0.5000 O2 + 2.0000 F- + 2.0000 H+
log_k 55.7197
-delta_H -390.924 kJ/mol # Calculated enthalpy of reaction F2(g)
# Enthalpy of formation: 0 kJ/mol
-analytic -3.2664e+001 -2.1035e-002 1.9974e+004 1.1174e+001 3.3920e+002
# -Range: 0-200
H2(g)
# H2 +0.5000 O2 = + 1.0000 H2O
# log_k 43.0016
# -analytic -1.1609e+001 -3.7580e-003 1.5068e+004 2.4198e+000 -7.0997e+004
# -Range: 0-300
H2 = H2
log_k -3.1050
-delta_H -4.184 kJ/mol # Calculated enthalpy of reaction H2(g)
# Enthalpy of formation: 0 kcal/mol
-analytic -9.3114e+000 4.6473e-003 -4.9335e+001 1.4341e+000 1.2815e+005
# -Range: 0-300
H2O(g)
H2O = + 1.0000 H2O
log_k 1.5854
-delta_H -43.4383 kJ/mol # Calculated enthalpy of reaction H2O(g)
# Enthalpy of formation: -57.935 kcal/mol
-analytic -1.4782e+001 1.0752e-003 2.7519e+003 2.7548e+000 4.2945e+001
# -Range: 0-300
H2S(g)
H2S = + 1.0000 H+ + 1.0000 HS-
log_k -7.9759
-delta_H 4.5229 kJ/mol # Calculated enthalpy of reaction H2S(g)
# Enthalpy of formation: -4.931 kcal/mol
-analytic -9.7354e+001 -3.1576e-002 1.8285e+003 3.7440e+001 2.8560e+001
# -Range: 0-300
HBr(g)
HBr = + 1.0000 Br- + 1.0000 H+
log_k 8.8815
-delta_H -85.2134 kJ/mol # Calculated enthalpy of reaction HBr(g)
# Enthalpy of formation: -36.29 kJ/mol
-analytic 8.1303e+000 -6.6641e-003 3.3951e+003 -3.4973e+000 5.7651e+001
# -Range: 0-200
HCl(g)
HCl = + 1.0000 Cl- + 1.0000 H+
log_k 6.3055
-delta_H -74.7697 kJ/mol # Calculated enthalpy of reaction HCl(g)
# Enthalpy of formation: -92.31 kJ/mol
-analytic -2.8144e-001 -8.6776e-003 3.0668e+003 -4.5105e-001 5.2078e+001
# -Range: 0-200
HF(g)
HF = + 1.0000 F- + 1.0000 H+
log_k 1.1126
-delta_H 0 # Not possible to calculate enthalpy of reaction Hf(g)
# Enthalpy of formation: 619.234 kJ/mol
-analytic -8.5783e+000 -8.8440e-003 2.6279e+003 1.4180e+000 4.4628e+001
# -Range: 0-200
HI(g)
HI = + 1.0000 H+ + 1.0000 I-
log_k 9.3944
-delta_H -83.4024 kJ/mol # Calculated enthalpy of reaction HI(g)
# Enthalpy of formation: 26.5 kJ/mol
-analytic 5.8250e-003 -8.7146e-003 3.5728e+003 0.0000e+000 0.0000e+000
# -Range: 0-200
He(g)
He = + 1.0000 He
log_k -3.4143
-delta_H -0.6276 kJ/mol # Calculated enthalpy of reaction He(g)
# Enthalpy of formation: 0 kcal/mol
-analytic -1.3402e+001 4.6358e-003 1.8295e+002 2.8070e+000 9.3373e+004
# -Range: 0-300
Hf(g)
Hf +4.0000 H+ +1.0000 O2 = + 1.0000 Hf++++ + 2.0000 H2O
log_k 290.9782
-delta_H 0 # Not possible to calculate enthalpy of reaction Hf(g)
# Enthalpy of formation: 0 kJ/mol
Hg(g)
Hg +2.0000 H+ +0.5000 O2 = + 1.0000 H2O + 1.0000 Hg++
log_k 19.7290
-delta_H -170.988 kJ/mol # Calculated enthalpy of reaction Hg(g)
# Enthalpy of formation: 61.38 kJ/mol
-analytic -1.6232e+001 -3.2863e-003 8.9831e+003 2.7505e+000 1.5255e+002
# -Range: 0-200
I2(g)
I2 +1.0000 H2O = + 0.5000 O2 + 2.0000 H+ + 2.0000 I-
log_k -21.4231
-delta_H 103.547 kJ/mol # Calculated enthalpy of reaction I2(g)
# Enthalpy of formation: 62.42 kJ/mol
-analytic -2.0271e+001 -2.1890e-002 -6.0267e+003 1.0339e+001 -1.0233e+002
# -Range: 0-200
K(g)
K +1.0000 H+ +0.2500 O2 = + 0.5000 H2O + 1.0000 K+
log_k 81.5815
-delta_H -481.055 kJ/mol # Calculated enthalpy of reaction K(g)
# Enthalpy of formation: 89 kJ/mol
-analytic 1.0278e+001 3.0700e-003 2.4729e+004 -5.0763e+000 4.1994e+002
# -Range: 0-200
Kr(g)
Kr = + 1.0000 Kr
log_k -2.6051
-delta_H -15.2716 kJ/mol # Calculated enthalpy of reaction Kr(g)
# Enthalpy of formation: 0 kcal/mol
-analytic -2.1251e+001 4.8308e-003 4.2971e+002 5.3591e+000 2.2304e+005
# -Range: 0-300
Li(g)
Li +1.0000 H+ +0.2500 O2 = + 0.5000 H2O + 1.0000 Li+
log_k 94.9423
-delta_H -577.639 kJ/mol # Calculated enthalpy of reaction Li(g)
# Enthalpy of formation: 159.3 kJ/mol
-analytic -2.5692e+001 -1.4385e-003 3.0936e+004 6.9899e+000 5.2535e+002
# -Range: 0-200
Mg(g)
Mg +2.0000 H+ +0.5000 O2 = + 1.0000 H2O + 1.0000 Mg++
log_k 142.2494
-delta_H -892.831 kJ/mol # Calculated enthalpy of reaction Mg(g)
# Enthalpy of formation: 147.1 kJ/mol
-analytic -1.3470e+000 -7.7402e-004 4.5992e+004 -4.2207e+000 7.8101e+002
# -Range: 0-200
N2(g)
# N2 +3.0000 H2O = + 1.5000 O2 + 2.0000 NH3
# log_k -119.6473
# -analytic 2.4168e+001 1.6489e-002 -3.6869e+004 -1.1181e+001 2.3178e+005
# -Range: 0-300
N2 = N2
log_k -3.1864
-delta_H -10.4391 kJ/mol # Calculated enthalpy of reaction N2(g)
# Enthalpy of formation: 0 kcal/mol
-analytic -7.6452e+000 7.9606e-003 0.0000e+000 0.0000e+000 1.8604e+005
# -Range: 0-300
NH3(g)
NH3 = + 1.0000 NH3
log_k 1.7966
-delta_H -35.2251 kJ/mol # Calculated enthalpy of reaction NH3(g)
# Enthalpy of formation: -11.021 kcal/mol
-analytic -1.8758e+001 3.3670e-004 2.5113e+003 4.8619e+000 3.9192e+001
# -Range: 0-300
NO(g)
NO +0.5000 H2O +0.2500 O2 = + 1.0000 H+ + 1.0000 NO2-
log_k 0.7554
-delta_H -48.8884 kJ/mol # Calculated enthalpy of reaction NO(g)
# Enthalpy of formation: 90.241 kJ/mol
-analytic 8.2147e+000 -1.2708e-001 -6.0593e+003 2.0504e+001 -9.4551e+001
# -Range: 0-300
NO2(g)
NO2 +0.5000 H2O +0.2500 O2 = + 1.0000 H+ + 1.0000 NO3-
log_k 8.3673
-delta_H -94.0124 kJ/mol # Calculated enthalpy of reaction NO2(g)
# Enthalpy of formation: 33.154 kJ/mol
-analytic 9.4389e+001 -2.7511e-001 -1.6783e+004 2.1127e+001 -2.6191e+002
# -Range: 0-300
Na(g)
Na +1.0000 H+ +0.2500 O2 = + 0.5000 H2O + 1.0000 Na+
log_k 80.8640
-delta_H -487.685 kJ/mol # Calculated enthalpy of reaction Na(g)
# Enthalpy of formation: 107.5 kJ/mol
-analytic -6.0156e+000 2.4712e-003 2.5682e+004 0.0000e+000 0.0000e+000
# -Range: 0-200
Ne(g)
Ne = + 1.0000 Ne
log_k -3.3462
-delta_H -3.64008 kJ/mol # Calculated enthalpy of reaction Ne(g)
# Enthalpy of formation: 0 kcal/mol
-analytic -6.5169e+000 6.3991e-003 0.0000e+000 0.0000e+000 1.1271e+005
# -Range: 0-300
O2(g)
O2 = + 1.0000 O2
log_k -2.8983
-delta_H -12.1336 kJ/mol # Calculated enthalpy of reaction O2(g)
# Enthalpy of formation: 0 kcal/mol
-analytic -7.5001e+000 7.8981e-003 0.0000e+000 0.0000e+000 2.0027e+005
# -Range: 0-300
Pb(g)
Pb +2.0000 H+ +0.5000 O2 = + 1.0000 H2O + 1.0000 Pb++
log_k 75.6090
-delta_H -474.051 kJ/mol # Calculated enthalpy of reaction Pb(g)
# Enthalpy of formation: 195.2 kJ/mol
-analytic 2.5752e+001 2.1307e-003 2.3397e+004 -1.1825e+001 3.9730e+002
# -Range: 0-200
Rb(g)
Rb +1.0000 H+ +0.2500 O2 = + 0.5000 H2O + 1.0000 Rb+
log_k 80.4976
-delta_H -471.909 kJ/mol # Calculated enthalpy of reaction Rb(g)
# Enthalpy of formation: 80.9 kJ/mol
-analytic 2.6839e+001 5.9775e-003 2.3720e+004 -1.1189e+001 4.0279e+002
# -Range: 0-200
Rn(g)
Rn = + 1.0000 Rn
log_k -2.0451
-delta_H -20.92 kJ/mol # Calculated enthalpy of reaction Rn(g)
# Enthalpy of formation: 0 kcal/mol
-analytic -3.0258e+001 4.9893e-003 1.4118e+002 8.8798e+000 3.8095e+005
# -Range: 0-300
RuCl3(g)
RuCl3 = + 1.0000 Ru+++ + 3.0000 Cl-
log_k 41.5503
-delta_H 0 # Not possible to calculate enthalpy of reaction RuCl3(g)
# Enthalpy of formation: 16.84 kJ/mol
RuO3(g)
RuO3 +1.0000 H2O = + 1.0000 RuO4-- + 2.0000 H+
log_k 2.3859
-delta_H -100.369 kJ/mol # Calculated enthalpy of reaction RuO3(g)
# Enthalpy of formation: -70.868 kJ/mol
-analytic 1.1106e+002 1.7191e-002 6.8526e+002 -4.6922e+001 1.1598e+001
# -Range: 0-200
S2(g)
S2 +2.0000 H2O = + 0.5000 SO4-- + 1.5000 HS- + 2.5000 H+
log_k -7.1449
-delta_H -35.656 kJ/mol # Calculated enthalpy of reaction S2(g)
# Enthalpy of formation: 30.681 kcal/mol
-analytic -1.8815e+002 -7.7069e-002 4.8816e+003 7.5802e+001 7.6228e+001
# -Range: 0-300
SO2(g)
SO2 = SO2
log_k 0.1700
-delta_H 0 # Not possible to calculate enthalpy of reaction SO2(g)
# Enthalpy of formation: 0 kcal/mol
-analytic -2.0205e+001 2.8861e-003 1.4862e+003 5.2958e+000 1.2721e+005
# -Range: 0-300
Si(g)
Si +1.0000 O2 = + 1.0000 SiO2
log_k 219.9509
-delta_H -1315.57 kJ/mol # Calculated enthalpy of reaction Si(g)
# Enthalpy of formation: 450 kJ/mol
-analytic 4.1998e+002 8.0113e-002 5.4468e+004 -1.6433e+002 9.2480e+002
# -Range: 0-200
SiF4(g)
SiF4 +2.0000 H2O = + 1.0000 SiO2 + 4.0000 F- + 4.0000 H+
log_k -15.1931
-delta_H -32.4123 kJ/mol # Calculated enthalpy of reaction SiF4(g)
# Enthalpy of formation: -1615 kJ/mol
-analytic 3.4941e+002 3.3668e-002 -1.2780e+004 -1.3410e+002 -2.1714e+002
# -Range: 0-200
Sn(g)
Sn +2.0000 H+ +0.5000 O2 = + 1.0000 H2O + 1.0000 Sn++
log_k 94.5019
-delta_H -589.758 kJ/mol # Calculated enthalpy of reaction Sn(g)
# Enthalpy of formation: 301.2 kJ/mol
-analytic 1.4875e+001 -5.6877e-005 2.9728e+004 -8.1131e+000 5.0482e+002
# -Range: 0-200
Tc2O7(g)
Tc2O7 +1.0000 H2O = + 2.0000 H+ + 2.0000 TcO4-
log_k 21.3593
-delta_H -158.131 kJ/mol # Calculated enthalpy of reaction Tc2O7(g)
# Enthalpy of formation: -988.569 kJ/mol
-analytic 7.4140e+001 1.5668e-002 5.6360e+003 -3.0860e+001 9.5682e+001
# -Range: 0-200
Th(g)
Th +4.0000 H+ +1.0000 O2 = + 1.0000 Th++++ + 2.0000 H2O
log_k 307.8413
-delta_H -1930.56 kJ/mol # Calculated enthalpy of reaction Th(g)
# Enthalpy of formation: 602 kJ/mol
-analytic 1.8496e+001 2.7318e-003 9.8807e+004 -1.7332e+001 1.6779e+003
# -Range: 0-200
Ti(g)
Ti +2.0000 H2O +1.0000 O2 = + 1.0000 Ti(OH)4
log_k 224.3510
-delta_H 0 # Not possible to calculate enthalpy of reaction Ti(g)
# Enthalpy of formation: 473 kJ/mol
TiBr4(g)
TiBr4 +4.0000 H2O = + 1.0000 Ti(OH)4 + 4.0000 Br- + 4.0000 H+
log_k 36.6695
-delta_H 0 # Not possible to calculate enthalpy of reaction TiBr4(g)
# Enthalpy of formation: -549.339 kJ/mol
TiCl4(g)
TiCl4 +4.0000 H2O = + 1.0000 Ti(OH)4 + 4.0000 Cl- + 4.0000 H+
log_k 28.0518
-delta_H 0 # Not possible to calculate enthalpy of reaction TiCl4(g)
# Enthalpy of formation: -763.2 kJ/mol
TiO(g)
TiO +2.0000 H2O +0.5000 O2 = + 1.0000 Ti(OH)4
log_k 145.5711
-delta_H 0 # Not possible to calculate enthalpy of reaction TiO(g)
# Enthalpy of formation: 17.144 kJ/mol
U(g)
U +2.0000 H+ +1.5000 O2 = + 1.0000 H2O + 1.0000 UO2++
log_k 298.3441
-delta_H -1819.64 kJ/mol # Calculated enthalpy of reaction U(g)
# Enthalpy of formation: 533 kJ/mol
-analytic 3.7536e+001 -6.3804e-003 9.2048e+004 -1.8614e+001 1.4363e+003
# -Range: 0-300
U2Cl10(g)
U2Cl10 +4.0000 H2O = + 2.0000 UO2+ + 8.0000 H+ + 10.0000 Cl-
log_k 82.7621
-delta_H -609.798 kJ/mol # Calculated enthalpy of reaction U2Cl10(g)
# Enthalpy of formation: -1967.9 kJ/mol
-analytic -7.5513e+002 -3.0070e-001 4.5824e+004 3.1267e+002 7.1526e+002
# -Range: 0-300
U2Cl8(g)
U2Cl8 = + 2.0000 U++++ + 8.0000 Cl-
log_k 82.4059
-delta_H -769.437 kJ/mol # Calculated enthalpy of reaction U2Cl8(g)
# Enthalpy of formation: -1749.6 kJ/mol
-analytic -7.4441e+002 -2.6943e-001 5.4358e+004 2.9287e+002 8.4843e+002
# -Range: 0-300
U2F10(g)
U2F10 +4.0000 H2O = + 2.0000 UO2+ + 8.0000 H+ + 10.0000 F-
log_k -12.2888
-delta_H -239.377 kJ/mol # Calculated enthalpy of reaction U2F10(g)
# Enthalpy of formation: -4021 kJ/mol
-analytic -9.1542e+002 -3.2040e-001 3.1047e+004 3.6143e+002 4.8473e+002
# -Range: 0-300
UBr(g)
UBr +1.0000 O2 = + 1.0000 Br- + 1.0000 UO2+
log_k 224.8412
-delta_H -1381.5 kJ/mol # Calculated enthalpy of reaction UBr(g)
# Enthalpy of formation: 247 kJ/mol
-analytic -3.1193e+002 -6.3059e-002 8.7633e+004 1.1032e+002 -1.0104e+006
# -Range: 0-300
UBr2(g)
UBr2 +1.0000 O2 = + 1.0000 UO2++ + 2.0000 Br-
log_k 192.6278
-delta_H -1218.87 kJ/mol # Calculated enthalpy of reaction UBr2(g)
# Enthalpy of formation: -31 kJ/mol
-analytic -1.2277e+002 -6.4613e-002 6.4196e+004 4.8209e+001 1.0018e+003
# -Range: 0-300
UBr3(g)
UBr3 = + 1.0000 U+++ + 3.0000 Br-
log_k 67.8918
-delta_H -489.61 kJ/mol # Calculated enthalpy of reaction UBr3(g)
# Enthalpy of formation: -364 kJ/mol
-analytic -2.5784e+002 -9.7583e-002 3.0225e+004 1.0240e+002 4.7171e+002
# -Range: 0-300
UBr4(g)
UBr4 = + 1.0000 U++++ + 4.0000 Br-
log_k 54.2926
-delta_H -467.113 kJ/mol # Calculated enthalpy of reaction UBr4(g)
# Enthalpy of formation: -610.1 kJ/mol
-analytic -3.5205e+002 -1.2867e-001 3.0898e+004 1.3781e+002 4.8223e+002
# -Range: 0-300
UBr5(g)
UBr5 +2.0000 H2O = + 1.0000 UO2+ + 4.0000 H+ + 5.0000 Br-
log_k 61.4272
-delta_H -423.222 kJ/mol # Calculated enthalpy of reaction UBr5(g)
# Enthalpy of formation: -637.745 kJ/mol
-analytic -3.4693e+002 -1.4298e-001 2.8151e+004 1.4406e+002 4.3938e+002
# -Range: 0-300
UCl(g)
UCl +1.0000 O2 = + 1.0000 Cl- + 1.0000 UO2+
log_k 221.7887
-delta_H -1368.27 kJ/mol # Calculated enthalpy of reaction UCl(g)
# Enthalpy of formation: 188.2 kJ/mol
-analytic -4.1941e+001 -2.7879e-002 7.0800e+004 1.3954e+001 1.1048e+003
# -Range: 0-300
UCl2(g)
UCl2 +1.0000 O2 = + 1.0000 UO2++ + 2.0000 Cl-
log_k 183.7912
-delta_H -1178.03 kJ/mol # Calculated enthalpy of reaction UCl2(g)
# Enthalpy of formation: -163 kJ/mol
-analytic -1.3677e+002 -6.7829e-002 6.2413e+004 5.3100e+001 9.7394e+002
# -Range: 0-300
UCl3(g)
UCl3 = + 1.0000 U+++ + 3.0000 Cl-
log_k 58.6335
-delta_H -453.239 kJ/mol # Calculated enthalpy of reaction UCl3(g)
# Enthalpy of formation: -537.1 kJ/mol
-analytic -2.7942e+002 -1.0243e-001 2.8859e+004 1.0982e+002 4.5040e+002
# -Range: 0-300
UCl4(g)
UCl4 = + 1.0000 U++++ + 4.0000 Cl-
log_k 46.3988
-delta_H -441.419 kJ/mol # Calculated enthalpy of reaction UCl4(g)
# Enthalpy of formation: -818.1 kJ/mol
-analytic -3.7971e+002 -1.3504e-001 3.0243e+004 1.4746e+002 4.7202e+002
# -Range: 0-300
UCl5(g)
UCl5 +2.0000 H2O = + 1.0000 UO2+ + 4.0000 H+ + 5.0000 Cl-
log_k 54.5311
-delta_H -406.349 kJ/mol # Calculated enthalpy of reaction UCl5(g)
# Enthalpy of formation: -882.5 kJ/mol
-analytic -3.8234e+002 -1.5109e-001 2.8170e+004 1.5654e+002 4.3968e+002
# -Range: 0-300
UCl6(g)
UCl6 +2.0000 H2O = + 1.0000 UO2++ + 4.0000 H+ + 6.0000 Cl-
log_k 63.4791
-delta_H -462.301 kJ/mol # Calculated enthalpy of reaction UCl6(g)
# Enthalpy of formation: -987.5 kJ/mol
-analytic -4.7128e+002 -1.9133e-001 3.2528e+004 1.9503e+002 5.0771e+002
# -Range: 0-300
UF(g)
UF +1.0000 O2 = + 1.0000 F- + 1.0000 UO2+
log_k 206.2684
-delta_H -1296.34 kJ/mol # Calculated enthalpy of reaction UF(g)
# Enthalpy of formation: -52 kJ/mol
-analytic -6.1248e+001 -3.0360e-002 6.7619e+004 2.0095e+001 1.0551e+003
# -Range: 0-300
UF2(g)
UF2 +1.0000 O2 = + 1.0000 UO2++ + 2.0000 F-
log_k 172.3563
-delta_H -1147.56 kJ/mol # Calculated enthalpy of reaction UF2(g)
# Enthalpy of formation: -530 kJ/mol
-analytic -4.3462e+002 -1.0881e-001 7.6778e+004 1.5835e+002 -8.8536e+005
# -Range: 0-300
UF3(g)
UF3 = + 1.0000 U+++ + 3.0000 F-
log_k 47.2334
-delta_H -440.943 kJ/mol # Calculated enthalpy of reaction UF3(g)
# Enthalpy of formation: -1054.2 kJ/mol
-analytic -3.3058e+002 -1.0866e-001 2.9694e+004 1.2551e+002 4.6344e+002
# -Range: 0-300
UF4(g)
UF4 = + 1.0000 U++++ + 4.0000 F-
log_k 14.5980
-delta_H -331.39 kJ/mol # Calculated enthalpy of reaction UF4(g)
# Enthalpy of formation: -1601.2 kJ/mol
-analytic -4.4692e+002 -1.4314e-001 2.6427e+004 1.6791e+002 4.1250e+002
# -Range: 0-300
UF5(g)
UF5 +2.0000 H2O = + 1.0000 UO2+ + 4.0000 H+ + 5.0000 F-
log_k 6.3801
-delta_H -220.188 kJ/mol # Calculated enthalpy of reaction UF5(g)
# Enthalpy of formation: -1910 kJ/mol
-analytic -4.6981e+002 -1.6177e-001 2.0986e+004 1.8345e+002 3.2760e+002
# -Range: 0-300
UF6(g)
UF6 +2.0000 H2O = + 1.0000 UO2++ + 4.0000 H+ + 6.0000 F-
log_k 18.2536
-delta_H -310.809 kJ/mol # Calculated enthalpy of reaction UF6(g)
# Enthalpy of formation: -2148.6 kJ/mol
-analytic -5.7661e+002 -2.0409e-001 2.7680e+004 2.2743e+002 4.3209e+002
# -Range: 0-300
UI(g)
UI +1.0000 O2 = + 1.0000 I- + 1.0000 UO2+
log_k 230.8161
-delta_H -1410.9 kJ/mol # Calculated enthalpy of reaction UI(g)
# Enthalpy of formation: 341 kJ/mol
-analytic -3.5819e+001 -2.6631e-002 7.2899e+004 1.2133e+001 1.1375e+003
# -Range: 0-300
UI2(g)
UI2 +1.0000 O2 = + 1.0000 UO2++ + 2.0000 I-
log_k 194.5395
-delta_H -1220.67 kJ/mol # Calculated enthalpy of reaction UI2(g)
# Enthalpy of formation: 100 kJ/mol
-analytic -3.3543e+002 -9.5116e-002 7.6218e+004 1.2543e+002 -6.8683e+005
# -Range: 0-300
UI3(g)
UI3 = + 1.0000 U+++ + 3.0000 I-
log_k 75.6033
-delta_H -519.807 kJ/mol # Calculated enthalpy of reaction UI3(g)
# Enthalpy of formation: -140 kJ/mol
-analytic -2.6095e+002 -9.8782e-002 3.1972e+004 1.0456e+002 4.9897e+002
# -Range: 0-300
UI4(g)
UI4 = + 1.0000 U++++ + 4.0000 I-
log_k 64.3272
-delta_H -510.01 kJ/mol # Calculated enthalpy of reaction UI4(g)
# Enthalpy of formation: -308.8 kJ/mol
-analytic -3.5645e+002 -1.3022e-001 3.3347e+004 1.4051e+002 5.2046e+002
# -Range: 0-300
UO(g)
UO +2.0000 H+ +1.0000 O2 = + 1.0000 H2O + 1.0000 UO2++
log_k 211.6585
-delta_H -1323.2 kJ/mol # Calculated enthalpy of reaction UO(g)
# Enthalpy of formation: 30.5 kJ/mol
-analytic -1.8007e+002 -3.1985e-002 7.8469e+004 5.8892e+001 -6.8071e+005
# -Range: 0-300
UO2(g)
UO2 +2.0000 H+ +0.5000 O2 = + 1.0000 H2O + 1.0000 UO2++
log_k 125.6027
-delta_H -820.972 kJ/mol # Calculated enthalpy of reaction UO2(g)
# Enthalpy of formation: -477.8 kJ/mol
-analytic -5.2789e+000 -3.5754e-003 4.2074e+004 -3.7117e+000 6.5653e+002
# -Range: 0-300
UO2Cl2(g)
UO2Cl2 = + 1.0000 UO2++ + 2.0000 Cl-
log_k 47.9630
-delta_H -381.559 kJ/mol # Calculated enthalpy of reaction UO2Cl2(g)
# Enthalpy of formation: -971.6 kJ/mol
-analytic -1.8035e+002 -6.5574e-002 2.3064e+004 6.8894e+001 3.5994e+002
# -Range: 0-300
UO2F2(g)
UO2F2 = + 1.0000 UO2++ + 2.0000 F-
log_k 34.6675
-delta_H -337.195 kJ/mol # Calculated enthalpy of reaction UO2F2(g)
# Enthalpy of formation: -1352.5 kJ/mol
-analytic -2.1498e+002 -6.9882e-002 2.1774e+004 7.9780e+001 3.3983e+002
# -Range: 0-300
UO3(g)
UO3 +2.0000 H+ = + 1.0000 H2O + 1.0000 UO2++
log_k 70.9480
-delta_H -505.638 kJ/mol # Calculated enthalpy of reaction UO3(g)
# Enthalpy of formation: -799.2 kJ/mol
-analytic -3.2820e+001 -2.6807e-003 2.6914e+004 5.7767e+000 4.1997e+002
# -Range: 0-300
UOF4(g)
UOF4 +1.0000 H2O = + 1.0000 UO2++ + 2.0000 H+ + 4.0000 F-
log_k 24.2848
-delta_H -312.552 kJ/mol # Calculated enthalpy of reaction UOF4(g)
# Enthalpy of formation: -1762 kJ/mol
-analytic -3.9592e+002 -1.3699e-001 2.4127e+004 1.5359e+002 3.7660e+002
# -Range: 0-300
Xe(g)
Xe = + 1.0000 Xe
log_k -2.3640
-delta_H -18.8698 kJ/mol # Calculated enthalpy of reaction Xe(g)
# Enthalpy of formation: 0 kcal/mol
-analytic -2.0636e+001 5.1389e-003 2.0490e+002 5.1913e+000 2.8556e+005
# -Range: 0-300
Zn(g)
Zn +2.0000 H+ +0.5000 O2 = + 1.0000 H2O + 1.0000 Zn++
log_k 85.4140
-delta_H -563.557 kJ/mol # Calculated enthalpy of reaction Zn(g)
# Enthalpy of formation: 130.4 kJ/mol
-analytic -1.0898e+001 -3.9871e-003 2.9068e+004 0.0000e+000 0.0000e+000
# -Range: 0-200
Zr(g)
Zr +4.0000 H+ +1.0000 O2 = + 1.0000 Zr++++ + 2.0000 H2O
log_k 277.1324
-delta_H 0 # Not possible to calculate enthalpy of reaction Zr(g)
# Enthalpy of formation: 608.948 kJ/mol
ZrF4(g)
ZrF4 = + 1.0000 Zr++++ + 4.0000 F-
log_k 142.9515
-delta_H 0 # Not possible to calculate enthalpy of reaction ZrF4(g)
# Enthalpy of formation: -858.24 kJ/mol
EXCHANGE_MASTER_SPECIES
X X-
EXCHANGE_SPECIES
X- = X-
log_k 0.0
Na+ + X- = NaX
log_k 0.0
-llnl_gamma 4.0
K+ + X- = KX
log_k 0.7
-llnl_gamma 3.0
delta_h -4.3 # Jardine & Sparks, 1984
Li+ + X- = LiX
log_k -0.08
-llnl_gamma 6.0
delta_h 1.4 # Merriam & Thomas, 1956
NH4+ + X- = NH4X
log_k 0.6
-llnl_gamma 2.5
delta_h -2.4 # Laudelout et al., 1968
Ca+2 + 2X- = CaX2
log_k 0.8
-llnl_gamma 6.0
delta_h 7.2 # Van Bladel & Gheyl, 1980
Mg+2 + 2X- = MgX2
log_k 0.6
-llnl_gamma 8.0
delta_h 7.4 # Laudelout et al., 1968
Sr+2 + 2X- = SrX2
log_k 0.91
-llnl_gamma 5.0
delta_h 5.5 # Laudelout et al., 1968
Ba+2 + 2X- = BaX2
log_k 0.91
-llnl_gamma 5.0
delta_h 4.5 # Laudelout et al., 1968
Mn+2 + 2X- = MnX2
log_k 0.52
-llnl_gamma 6.0
Fe+2 + 2X- = FeX2
log_k 0.44
-llnl_gamma 6.0
Cu+2 + 2X- = CuX2
log_k 0.6
-llnl_gamma 6.0
Zn+2 + 2X- = ZnX2
log_k 0.8
-llnl_gamma 6.0
Cd+2 + 2X- = CdX2
log_k 0.8
-llnl_gamma 5.0
Pb+2 + 2X- = PbX2
log_k 1.05
-llnl_gamma 4.5
Al+3 + 3X- = AlX3
log_k 0.41
-llnl_gamma 9.0
AlOH+2 + 2X- = AlOHX2
log_k 0.89
-llnl_gamma 4.5
SURFACE_MASTER_SPECIES
Hfo_s Hfo_sOH
Hfo_w Hfo_wOH
SURFACE_SPECIES
# All surface data from
# Dzombak and Morel, 1990
#
#
# Acid-base data from table 5.7
#
# strong binding site--Hfo_s,
Hfo_sOH = Hfo_sOH
log_k 0.0
Hfo_sOH + H+ = Hfo_sOH2+
log_k 7.29 # = pKa1,int
Hfo_sOH = Hfo_sO- + H+
log_k -8.93 # = -pKa2,int
# weak binding site--Hfo_w
Hfo_wOH = Hfo_wOH
log_k 0.0
Hfo_wOH + H+ = Hfo_wOH2+
log_k 7.29 # = pKa1,int
Hfo_wOH = Hfo_wO- + H+
log_k -8.93 # = -pKa2,int
###############################################
# CATIONS #
###############################################
#
# Cations from table 10.1 or 10.5
#
# Calcium
Hfo_sOH + Ca+2 = Hfo_sOHCa+2
log_k 4.97
Hfo_wOH + Ca+2 = Hfo_wOCa+ + H+
log_k -5.85
# Strontium
Hfo_sOH + Sr+2 = Hfo_sOHSr+2
log_k 5.01
Hfo_wOH + Sr+2 = Hfo_wOSr+ + H+
log_k -6.58
Hfo_wOH + Sr+2 + H2O = Hfo_wOSrOH + 2H+
log_k -17.60
# Barium
Hfo_sOH + Ba+2 = Hfo_sOHBa+2
log_k 5.46
Hfo_wOH + Ba+2 = Hfo_wOBa+ + H+
log_k -7.2 # table 10.5
#
# Cations from table 10.2
#
# Cadmium
Hfo_sOH + Cd+2 = Hfo_sOCd+ + H+
log_k 0.47
Hfo_wOH + Cd+2 = Hfo_wOCd+ + H+
log_k -2.91
# Zinc
Hfo_sOH + Zn+2 = Hfo_sOZn+ + H+
log_k 0.99
Hfo_wOH + Zn+2 = Hfo_wOZn+ + H+
log_k -1.99
# Copper
Hfo_sOH + Cu+2 = Hfo_sOCu+ + H+
log_k 2.89
Hfo_wOH + Cu+2 = Hfo_wOCu+ + H+
log_k 0.6 # table 10.5
# Lead
Hfo_sOH + Pb+2 = Hfo_sOPb+ + H+
log_k 4.65
Hfo_wOH + Pb+2 = Hfo_wOPb+ + H+
log_k 0.3 # table 10.5
#
# Derived constants table 10.5
#
# Magnesium
Hfo_wOH + Mg+2 = Hfo_wOMg+ + H+
log_k -4.6
# Manganese
Hfo_sOH + Mn+2 = Hfo_sOMn+ + H+
log_k -0.4 # table 10.5
Hfo_wOH + Mn+2 = Hfo_wOMn+ + H+
log_k -3.5 # table 10.5
# Iron
Hfo_sOH + Fe+2 = Hfo_sOFe+ + H+
log_k 0.7 # LFER using table 10.5
Hfo_wOH + Fe+2 = Hfo_wOFe+ + H+
log_k -2.5 # LFER using table 10.5
###############################################
# ANIONS #
###############################################
#
# Anions from table 10.6
#
# Phosphate
Hfo_wOH + PO4-3 + 3H+ = Hfo_wH2PO4 + H2O
log_k 31.29
Hfo_wOH + PO4-3 + 2H+ = Hfo_wHPO4- + H2O
log_k 25.39
Hfo_wOH + PO4-3 + H+ = Hfo_wPO4-2 + H2O
log_k 17.72
#
# Anions from table 10.7
#
# Borate
Hfo_wOH + B(OH)3 = Hfo_wH2BO3 + H2O
log_k 0.62
#
# Anions from table 10.8
#
# Sulfate
Hfo_wOH + SO4-2 + H+ = Hfo_wSO4- + H2O
log_k 7.78
Hfo_wOH + SO4-2 = Hfo_wOHSO4-2
log_k 0.79
#
# Derived constants table 10.10
#
Hfo_wOH + F- + H+ = Hfo_wF + H2O
log_k 8.7
Hfo_wOH + F- = Hfo_wOHF-
log_k 1.6
#
# Carbonate: Van Geen et al., 1994 reoptimized for HFO
# 0.15 g HFO/L has 0.344 mM sites == 2 g of Van Geen's Goethite/L
#
# Hfo_wOH + CO3-2 + H+ = Hfo_wCO3- + H2O
# log_k 12.56
#
# Hfo_wOH + CO3-2 + 2H+= Hfo_wHCO3 + H2O
# log_k 20.62
# 9/19/96
# Added analytical expression for H2S, NH3, KSO4.
# Added species CaHSO4+.
# Added delta H for Goethite.
RATES
###########
#K-feldspar
###########
#
# <NAME>., 1990, The kinetics of base cation release due to
# chemical weathering: Lund University Press, Lund, 246 p.
#
# Example of KINETICS data block for K-feldspar rate:
# KINETICS 1
# K-feldspar
# -m0 2.16 # 10% K-fsp, 0.1 mm cubes
# -m 1.94
# -parms 1.36e4 0.1
K-feldspar
-start
1 rem specific rate from Sverdrup, 1990, in kmol/m2/s
2 rem parm(1) = 10 * (A/V, 1/dm) (recalc's sp. rate to mol/kgw)
3 rem parm(2) = corrects for field rate relative to lab rate
4 rem temp corr: from p. 162. E (kJ/mol) / R / 2.303 = H in H*(1/T-1/298)
10 dif_temp = 1/TK - 1/298
20 pk_H = 12.5 + 3134 * dif_temp
30 pk_w = 15.3 + 1838 * dif_temp
40 pk_OH = 14.2 + 3134 * dif_temp
50 pk_CO2 = 14.6 + 1677 * dif_temp
#60 pk_org = 13.9 + 1254 * dif_temp # rate increase with DOC
70 rate = 10^-pk_H * ACT("H+")^0.5 + 10^-pk_w + 10^-pk_OH * ACT("OH-")^0.3
71 rate = rate + 10^-pk_CO2 * (10^SI("CO2(g)"))^0.6
#72 rate = rate + 10^-pk_org * TOT("Doc")^0.4
80 moles = parm(1) * parm(2) * rate * (1 - SR("K-feldspar")) * time
81 rem decrease rate on precipitation
90 if SR("K-feldspar") > 1 then moles = moles * 0.1
100 save moles
-end
###########
#Albite
###########
#
# <NAME>., 1990, The kinetics of base cation release due to
# chemical weathering: Lund University Press, Lund, 246 p.
#
# Example of KINETICS data block for Albite rate:
# KINETICS 1
# Albite
# -m0 0.43 # 2% Albite, 0.1 mm cubes
# -parms 2.72e3 0.1
Albite
-start
1 rem specific rate from Sverdrup, 1990, in kmol/m2/s
2 rem parm(1) = 10 * (A/V, 1/dm) (recalc's sp. rate to mol/kgw)
3 rem parm(2) = corrects for field rate relative to lab rate
4 rem temp corr: from p. 162. E (kJ/mol) / R / 2.303 = H in H*(1/T-1/298)
10 dif_temp = 1/TK - 1/298
20 pk_H = 12.5 + 3359 * dif_temp
30 pk_w = 14.8 + 2648 * dif_temp
40 pk_OH = 13.7 + 3359 * dif_temp
#41 rem ^12.9 in Sverdrup, but larger than for oligoclase...
50 pk_CO2 = 14.0 + 1677 * dif_temp
#60 pk_org = 12.5 + 1254 * dif_temp # ...rate increase for DOC
70 rate = 10^-pk_H * ACT("H+")^0.5 + 10^-pk_w + 10^-pk_OH * ACT("OH-")^0.3
71 rate = rate + 10^-pk_CO2 * (10^SI("CO2(g)"))^0.6
#72 rate = rate + 10^-pk_org * TOT("Doc")^0.4
80 moles = parm(1) * parm(2) * rate * (1 - SR("Albite")) * time
81 rem decrease rate on precipitation
90 if SR("Albite") > 1 then moles = moles * 0.1
100 save moles
-end
########
#Calcite
########
#
# <NAME>., <NAME>., and <NAME>., 1978,
# American Journal of Science, v. 278, p. 179-216.
#
# Example of KINETICS data block for calcite rate:
#
# KINETICS 1
# Calcite
# -tol 1e-8
# -m0 3.e-3
# -m 3.e-3
# -parms 5.0 0.6
Calcite
-start
1 rem Modified from Plummer and others, 1978
2 rem parm(1) = A/V, 1/m parm(2) = exponent for m/m0
10 si_cc = si("Calcite")
20 if (m <= 0 and si_cc < 0) then goto 200
30 k1 = 10^(0.198 - 444.0 / (273.16 + tc) )
40 k2 = 10^(2.84 - 2177.0 / (273.16 + tc) )
50 if tc <= 25 then k3 = 10^(-5.86 - 317.0 / (273.16 + tc) )
60 if tc > 25 then k3 = 10^(-1.1 - 1737.0 / (273.16 + tc) )
70 t = 1
80 if m0 > 0 then t = m/m0
90 if t = 0 then t = 1
100 moles = parm(1) * (t)^parm(2)
110 moles = moles * (k1 * act("H+") + k2 * act("CO2") + k3 * act("H2O"))
120 moles = moles * (1 - 10^(2/3*si_cc))
130 moles = moles * time
140 if (moles > m) then moles = m
150 if (moles >= 0) then goto 200
160 temp = tot("Ca")
170 mc = tot("C(4)")
180 if mc < temp then temp = mc
190 if -moles > temp then moles = -temp
200 save moles
-end
#######
#Pyrite
#######
#
# <NAME>. and <NAME>., 1994,
# Geochimica et Cosmochimica Acta, v. 58, p. 5443-5454.
#
# Example of KINETICS data block for pyrite rate:
# KINETICS 1
# Pyrite
# -tol 1e-8
# -m0 5.e-4
# -m 5.e-4
# -parms 2.0 0.67 .5 -0.11
Pyrite
-start
1 rem Williamson and Rimstidt, 1994
2 rem parm(1) = log10(A/V, 1/dm) parm(2) = exp for (m/m0)
3 rem parm(3) = exp for O2 parm(4) = exp for H+
10 if (m <= 0) then goto 200
20 if (si("Pyrite") >= 0) then goto 200
20 rate = -10.19 + parm(1) + parm(3)*lm("O2") + parm(4)*lm("H+") + parm(2)*log10(m/m0)
30 moles = 10^rate * time
40 if (moles > m) then moles = m
200 save moles
-end
##########
#Organic_C
##########
#
# Example of KINETICS data block for Organic_C rate:
# KINETICS 1
# Organic_C
# -tol 1e-8
# # m in mol/kgw
# -m0 5e-3
# -m 5e-3
Organic_C
-start
1 rem Additive Monod kinetics
2 rem Electron acceptors: O2, NO3, and SO4
10 if (m <= 0) then goto 200
20 mO2 = mol("O2")
30 mNO3 = tot("N(5)")
40 mSO4 = tot("S(6)")
50 rate = 1.57e-9*mO2/(2.94e-4 + mO2) + 1.67e-11*mNO3/(1.55e-4 + mNO3)
60 rate = rate + 1.e-13*mSO4/(1.e-4 + mSO4)
70 moles = rate * m * (m/m0) * time
80 if (moles > m) then moles = m
200 save moles
-end
###########
#Pyrolusite
###########
#
# <NAME>, C.A.J., 2000, GCA 64, in press
#
# Example of KINETICS data block for Pyrolusite
# KINETICS 1-12
# Pyrolusite
# -tol 1.e-7
# -m0 0.1
# -m 0.1
Pyrolusite
-start
5 if (m <= 0.0) then goto 200
7 sr_pl = sr("Pyrolusite")
9 if abs(1 - sr_pl) < 0.1 then goto 200
10 if (sr_pl > 1.0) then goto 100
#20 rem initially 1 mol Fe+2 = 0.5 mol pyrolusite. k*A/V = 1/time (3 cells)
#22 rem time (3 cells) = 1.432e4. 1/time = 6.98e-5
30 Fe_t = tot("Fe(2)")
32 if Fe_t < 1.e-8 then goto 200
40 moles = 6.98e-5 * Fe_t * (m/m0)^0.67 * time * (1 - sr_pl)
50 if moles > Fe_t / 2 then moles = Fe_t / 2
70 if moles > m then moles = m
90 goto 200
100 Mn_t = tot("Mn")
110 moles = 2e-3 * 6.98e-5 * (1-sr_pl) * time
120 if moles <= -Mn_t then moles = -Mn_t
200 save moles
-end
END
% -
| .ipynb_checkpoints/Tutorial_EuPsalts_example-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import patsy as pt
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn import preprocessing
# %matplotlib inline
import re
import pymc3 as pm
import matplotlib.ticker as tk
import re
from sklearn.model_selection import StratifiedKFold
import pickle
# ## Import data
df = pd.read_csv('outputs/ala1_trials_clean.csv')
df = df.rename(columns={'project_name': 'basis', 'cluster__n_clusters': 'n', 'test_mean': 'y'}).\
loc[:, ['basis', 'y', 'n']]
# ## Scale predictors
#
#
# +
# to_log = ['n']
# for col in to_log:
# df.loc[:, col] = np.log(df[col])
to_scale = ['n']
scaler = preprocessing.MinMaxScaler()
vars_scaled = pd.DataFrame(scaler.fit_transform(df.loc[:, to_scale]), columns=[x+'_s' for x in to_scale])
df = df.join(vars_scaled)
df.T
# -
x = df.loc[df['basis']=='phipsi', 'n_s']
y = df.loc[df['basis']=='phipsi', 'y']
plt.scatter(x, y)
# ## Create design matrix
y = df.loc[:, 'y']
X = df.loc[:, df.columns.difference(['y'])]
X_c = pt.dmatrix('~ 0 + n_s + C(basis)', data=df, return_type='dataframe')
X_c = X_c.rename(columns=lambda x: re.sub('C|\\(|\\)|\\[|\\]','',x))
# ## Model fitting functions
# +
def gamma(alpha, beta):
def g(x):
return pm.Gamma(x, alpha=alpha, beta=beta)
return g
def hcauchy(beta):
def g(x):
return pm.HalfCauchy(x, beta=beta)
return g
def fit_model_1(y, X, kernel_type='rbf'):
"""
function to return a pymc3 model
y : dependent variable
X : independent variables
prop_Xu : number of inducing varibles to use
X, y are dataframes. We'll use the column names.
"""
with pm.Model() as model:
# Covert arrays
X_a = X.values
y_a = y.values
X_cols = list(X.columns)
# Globals
prop_Xu = 0.1 # proportion of observations to use as inducing variables
l_prior = gamma(1, 0.05)
eta_prior = hcauchy(2)
sigma_prior = hcauchy(2)
# Kernels
# 3 way interaction
eta = eta_prior('eta')
cov = eta**2
for i in range(X_a.shape[1]):
var_lab = 'l_'+X_cols[i]
if kernel_type.lower()=='rbf':
cov = cov*pm.gp.cov.ExpQuad(X_a.shape[1], ls=l_prior(var_lab), active_dims=[i])
if kernel_type.lower()=='exponential':
cov = cov*pm.gp.cov.Exponential(X_a.shape[1], ls=l_prior(var_lab), active_dims=[i])
if kernel_type.lower()=='m52':
cov = cov*pm.gp.cov.Matern52(X_a.shape[1], ls=l_prior(var_lab), active_dims=[i])
if kernel_type.lower()=='m32':
cov = cov*pm.gp.cov.Matern32(X_a.shape[1], ls=l_prior(var_lab), active_dims=[i])
# Covariance model
cov_tot = cov
# Model
gp = pm.gp.MarginalSparse(cov_func=cov_tot, approx="FITC")
# Noise model
sigma_n =sigma_prior('sigma_n')
# Inducing variables
num_Xu = int(X_a.shape[0]*prop_Xu)
print(type(num_Xu))
print(type(X_a))
Xu = pm.gp.util.kmeans_inducing_points(num_Xu, X_a)
# Marginal likelihood
y_ = gp.marginal_likelihood('y_', X=X_a, y=y_a,Xu=Xu, noise=sigma_n)
mp = pm.find_MAP()
return gp, mp, model
# -
# ## Fit mml model
gp, mp, model = fit_model_1(y, X_c, kernel_type='rbf')
# ## Plot MML model
# Create new data matrix
# +
# Create mesh
n_new = np.arange(0, 1000, 10)
bases = X['basis'].unique()
x1, x2 = np.meshgrid(n_new, bases)
# scale n:
# scaler_new = preprocessing.MinMaxScaler()
x1_s = scaler.transform(x1.reshape(-1, 1))
x1, x1_s, x2 = x1.flatten(), x1_s.flatten(), x2.flatten()
# Data frame:
X_new = pd.DataFrame({'n_s': x1_s, 'basis': x2, 'n': x1}, index=np.arange(len(x2)))
# Make design matrix
X_new_c = pt.dmatrix('~ 0 + n_s + C(basis)', data=X_new, return_type='dataframe')
X_new_c = X_new_c.rename(columns=lambda x: re.sub('C|\\(|\\)|\\[|\\]','',x))
# +
# Get predictions for evalution
with model:
# predict latent
mu, var = gp.predict(X_new_c.values, point=mp, diag=True,pred_noise=False)
sd_f = np.sqrt(var)
# predict target (includes noise)
_, var = gp.predict(X_new_c.values, point=mp, diag=True,pred_noise=True)
sd_y = np.sqrt(var)
# put in data frame
pred = pd.DataFrame({'f_pred': mu, 'sd_f': sd_f, 'sd_y': sd_y})
pred = pred.join(X_new)
# -
# Create lb and ub for plotting
pred.loc[:, 'lb'] = pred['f_pred']-2*pred['sd_f']
pred.loc[:, 'ub'] = pred['f_pred']+2*pred['sd_f']
# +
with sns.plotting_context('paper', font_scale=1.25):
g = sns.FacetGrid(data=pred, col='basis', sharey=False)
g.map(plt.plot, 'n', 'f_pred')
g.map(plt.fill_between, 'n', 'lb', 'ub', alpha=0.5 )
# plot the original data
for ax, col in zip(g.axes[0, :].flatten(), g.col_names):
idx = X['basis']==col
ax.scatter(X.loc[idx, 'n'], y[idx], marker='x', color='k', alpha=0.5)
# g.map(plt.scatter, '')
# plt.fill_between(X_new.flatten(), mu - 2*sd, mu + 2*sd, color="r", alpha=0.5)
| 4_fit_best_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
from scipy.interpolate import interp1d
class G4Material():
def __init__(self, label='', density=0.0, matid=0):
self.label = label
self.density = density
self.matid = matid
def __repr__(self):
return '<Material: "{!s}", {:d}, {:f}>'.format(self.label, self.matid, self.density)
def get_material_def(self):
"""returns entry suitable for voxel description in geometry.text file (format: [dens, matid] )"""
return "{:0.3f} {:d}\n".format(self.density, self.matid)
mat_map = {
"air": G4Material('Air', density=0.00129, matid=0),
"lung_in": G4Material('Lungs (inhale)', density=0.217, matid=1),
"lung_ex": G4Material('Lungs (exhale)', density=0.508, matid=2),
"adipose": G4Material('Adipose', density=0.967, matid=3),
"breast": G4Material('Breast', density=0.99, matid=4),
"water": G4Material('Water', density=1.0, matid=5),
"muscle": G4Material('Muscle', density=1.061, matid=6),
"liver": G4Material('Liver', density=1.071, matid=7),
"bone_trab": G4Material('Bone (trabecular)', density=1.159, matid=8),
"bone_comp": G4Material('Bone (compact)', density=1.575, matid=9),
"Io05": G4Material('Io05', density=1.0415, matid=10),
"Ba05": G4Material('Ba05', density=1.0405, matid=11),
"Gd05": G4Material('Gd05', density=1.0457, matid=12),
"Yb05": G4Material('Yb05', density=1.0447, matid=13),
"Ta05": G4Material('Ta05', density=1.0493, matid=14),
"Au05": G4Material('Au05', density=1.0498, matid=15),
"Bi05": G4Material('Bi05', density=1.0470, matid=16),
"Ca_50mgpml": G4Material('Ca_50mgpml', density=1.0177, matid=17),
"Ca_150mgpml": G4Material('Ca_150mgpml', density=1.0532, matid=18),
"Ca_300mgpml": G4Material('Ca_300mgpml', density=1.1065, matid=19),
"Io_5mgpml": G4Material('Io_5mgpml', density=1.0040, matid=20),
"Io_10mgpml": G4Material('Io_10mgpml', density=1.0080, matid=21),
"Io_20mgpml": G4Material('Io_20mgpml', density=1.0177, matid=22),
"Io_50mgpml": G4Material('Io_50mgpml', density=1.0399, matid=23),
}
lut_ct2dens = [
(-5000.0, 0.0),
(-1000.0, 0.01),
(-400, 0.602),
(-150, 0.924),
(100, 1.075),
(300, 1.145),
(2000, 1.856),
(4927, 3.379),
(66000, 7.8),
]
f_ct2dens = None
lut_dens2mat = [
(0.0, mat_map["air"] ),
(0.207, mat_map["lung_in"] ),
(0.481, mat_map["lung_ex"] ),
(0.919, mat_map["adipose"] ),
(0.979, mat_map["breast"] ),
(1.004, mat_map["water"] ),
(1.109, mat_map["muscle"] ),
(1.113, mat_map["liver"] ),
(1.496, mat_map["bone_trab"]),
(1.654, mat_map["bone_comp"]),
(6.0, mat_map["Io05"]),
(6.1, mat_map["Ba05"]),
(6.2, mat_map["Gd05"]),
(6.3, mat_map["Yb05"]),
(6.4, mat_map["Ta05"]),
(6.5, mat_map["Au05"]),
(6.6, mat_map["Bi05"]),
(6.7, mat_map["Ca_50mgpml"]),
(6.8, mat_map["Ca_150mgpml"]),
(6.9, mat_map["Ca_300mgpml"]),
(7.0, mat_map["Io_5mgpml"]),
(7.1, mat_map["Io_10mgpml"]),
(7.2, mat_map["Io_20mgpml"]),
(7.3, mat_map["Io_50mgpml"]),
]
f_dens2matindex = None
# +
def init_lut_interpolators():
global f_ct2dens, f_dens2matindex
if f_ct2dens is None:
lut_ct, lut_dens = zip(*lut_ct2dens)
f_ct2dens = interp1d(lut_ct, lut_dens, kind='linear', bounds_error=False, fill_value=(np.min(lut_dens), np.max(lut_dens)))
if f_dens2matindex is None:
lut_dens, mat_list = zip(*lut_dens2mat)
f_dens2matindex = interp1d(lut_dens, range(len(mat_list)), kind='previous', bounds_error=False, fill_value=(0, len(mat_list)-1))
# +
def lookup_materials(ctnums=None, densities=None, bulk_density=True):
"""convert either an array of ctnums or an array of densities to an array (of type=np.array(dtype=object)) of material specs
Promises to be much faster than the per-element version (ct2mat)
UNTESTED
"""
assert any(param is not None for param in (ctnums, densities))
init_lut_interpolators()
if ctnums is not None:
densities = f_ct2dens(ctnums)
print(densities.max())
materials = f_dens2mat(densities)
return materials
def f_dens2mat(densities):
"""use cached interpolator to convert densities array to object array of corresponding materials objects"""
init_lut_interpolators()
matindices = f_dens2matindex(densities).astype(int)
_, mat_list = zip(*lut_dens2mat)
material_choices = np.array(mat_list, dtype=object)
materials = material_choices[matindices]
return materials
# +
ctnums = [-1024,0,1518,50805,52186,53568,54949,56330,57712,59093]
materials = lookup_materials(ctnums)
# -
print(materials)
# +
ctnums = [0,0,0,0,0,0,0,41135,45279,49423]
materials = lookup_materials(ctnums)
print(materials)
# -
| Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from keras.datasets import boston_housing
import numpy as np
(training_x, training_y), (test_x, test_y) = boston_housing.load_data()
training_x[:5]
training_x.shape
training_x.mean(axis=0)
training_x.var(axis=0)
def normalize(data):
return (data - data.mean(axis=0)) / (data.std(axis=0))
training_x = normalize(training_x)
first_layer_size = 128
(num_samples, num_features) = training_x.shape
lr = 0.01
np.random.seed(100)
W1 = np.random.randn(num_features, first_layer_size)
B1 = np.random.randn(1, first_layer_size)
W2 = np.random.randn(first_layer_size, 1)
B2 = np.random.random()
a1 = np.zeros((num_samples, first_layer_size))
z1 = np.zeros((num_samples, first_layer_size))
a2 = np.zeros((num_samples, 1))
def forward():
global W2, W1, B1, B2, a1, z1, a2
a1 = np.dot(training_x, W1) + B1
z1 = np.tanh(a1)
a2 = np.dot(z1, W2) + B2
loss = 1/num_samples * np.sum((a2 - training_y.reshape(len(training_y), 1))**2, axis=0)[0]
return loss
def backward():
global W2, W1, B1, B2, a1, z1, a2
dla2 = 2/num_samples * np.sum(a2 - training_y.reshape(len(training_y), 1), axis=1)
dla2 = dla2.reshape(dla2.shape[0], 1)
dw2 = z1.T.dot(dla2)
db2 = dla2
dw1 = training_x.T.dot((dla2.dot(W2.T) * (1 - np.tanh(a1) ** 2)))
db1 = np.sum(dla2.dot(W2.T) * (1 - np.tanh(a1) ** 2), axis=0, keepdims=True)
W1 -= lr * dw1
B1 -= lr * db1
W2 -= lr * dw2
B2 -= lr * db2
for i in range(1000):
loss = forward()
backward()
print(f"{i}: {loss}")
# http://mccormickml.com/2014/03/04/gradient-descent-derivation/
#
# https://towardsdatascience.com/coding-neural-network-forward-propagation-and-backpropagtion-ccf8cf369f76
#
# https://machinelearningmastery.com/implement-backpropagation-algorithm-scratch-python/
| MSE from scratch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from elasticsearch import helpers, Elasticsearch
import csv
import pandas as pd
data = pd.read_csv('all_hadiths_clean.csv', encoding='utf-8-sig')
data.info()
# -
data.shape
data['text_ar'][26]
# # Pre-processing & normalizing the retrieved Hadith data :
data.drop('id',inplace=True,axis=1)
data.drop('hadith_id',inplace=True,axis=1)
data.drop('text_en',inplace=True,axis=1)
data.drop('chain_indx',inplace=True,axis=1)
data.info()
pd.unique(data['source'])
data['source'].replace(to_replace =' Sahih Bukhari ',
value ="صحيح البخاري", regex=True, inplace=True)
data['source'].replace(to_replace =' Sahih Muslim ',
value ="صحيح مسلم", regex=True, inplace=True)
data['source'].replace(to_replace =" Sunan Abi Da'ud ",
value ="سنن أبي داود", regex=True, inplace=True)
data['source'].replace(to_replace =" Jami' al-Tirmidhi ",
value ="سنن الترمذي", regex=True, inplace=True)
data['source'].replace(to_replace =" Sunan an-Nasa'i ",
value ="سنن النسائي", regex=True, inplace=True)
data['source'].replace(to_replace =' Sunan Ibn Majah ',
value ="سنن ابن ماجه", regex=True, inplace=True)
pd.unique(data['source'])
data.drop('hadith_no',inplace=True,axis=1)
data['text_ar'][5908]
# +
#getting rid of html source characters: \u200f.\u200f \u200f{\u200f \u200f"\u200f\u200f.\u200f
data['text_ar'].replace(to_replace ='\u200f"\u200f ',
value ="", regex=True, inplace=True)
data['text_ar'].replace(to_replace ='\u200f.\u200f',
value ="", regex=True, inplace=True)
data['text_ar'].replace(to_replace ='\u200f{\u200f',
value ="", regex=True, inplace=True)
data['text_ar'].replace(to_replace ='\u200f"\u200f\u200f.\u200f ',
value ="", regex=True, inplace=True)
# -
data['text_ar'].replace(to_replace ='\u200f"\u200f\u200f.\u200f ',
value ="", regex=True, inplace=True)
data.to_csv('Hadith_processed.csv')
data[:10]
es = Elasticsearch([{'host': 'localhost', 'port': '9200'}])
es.ping()
# +
import json
def csv_to_json(csvFilePath, jsonFilePath):
jsonArray = []
#read csv file
with open(csvFilePath, encoding='utf-8') as csvf:
#load csv file data using csv library's dictreader
csvReader = csv.DictReader(csvf)
#convert each csv row into python dict
for row in csvReader:
#add this python dict to json array
jsonArray.append(row)
#convert python jsonArray to JSON String and write to file
with open(jsonFilePath, 'w', encoding='utf-8-sig') as jsonf:
jsonString = json.dumps(jsonArray, indent=4)
jsonf.write(jsonString)
csvFilePath = r'Hadith_processed.csv'
jsonFilePath = r'json_hadit.json'
csv_to_json(csvFilePath, jsonFilePath)
# +
import codecs
hadith = codecs.open('json_hadith.json', 'r', 'utf-8-sig')
json_hadith = json.load(hadith)
json_hadith[7]
# +
hadit = []
for obj in json_hadith:
hadit.append({
"source": obj['source'],
"chapter_no": obj['chapter_no'],
"chapter": obj['chapter'],
"hadith": obj['text_ar'],
})
print(hadit[0])
# -
mapping ={
"settings" : {
"number_of_shards":1,
"number_of_replicas":1,
"analysis":{
"filter":{
"arabic_stop":{
"type":"stop",
"stopwords_path":"D:/Elastic/elasticsearch/config/merged-stopwords.txt"
},
"arabic_keywords":{
"type":"keyword_marker",
"keywords_path":"D:/Elastic/elasticsearch/config/keywordmarker-words.txt"
},
"arabic_stemmer":{
"type":"stemmer",
"language":"arabic"
},
"arabic_shingle":{
"type":"shingle",
"min_shingle_size":2,
"max_shingle_size":3
}
},
"analyzer":{
"rebuilt_arabic":{
"tokenizer":"standard",
"filter":[
"arabic_stop",
"arabic_normalization",
"arabic_keywords",
"arabic_shingle",
"arabic_stemmer"
]
},
"rebuilt":{
"tokenizer":"standard",
"filter":[
"arabic_stop",
"arabic_normalization",
"arabic_keywords",
"arabic_stemmer" #for one word aggregations
]
},
"simple_arabic":{
"tokenizer":"standard",
"filter":[
"arabic_stop",
"arabic_normalization",
"arabic_shingle"
]
},
"noshingle_arabic": {
"tokenizer":"standard",
"filter":[
"arabic_stop",
"arabic_normalization"
]
}
}
}
},
"mappings":{
"properties":{
"source":{
"type":"text",
"analyzer" : "simple_arabic",
"fielddata" : "true",
"fields":{
"keyword":{
"type":"keyword"
}
}
},
"chapter_no":{
"type":"integer"
},
"chapter":{
"type":"text",
"analyzer":"simple_arabic"
},
"hadith":{
"type":"text",
"analyzer":"rebuilt_arabic",
"fielddata":"true"
"fields":{
"nostem":{
"type":"text",
"analyzer":"simple_arabic",
"fielddata":"true"
},
"noshingle":{
"type":"text",
"analyzer":"noshingle_arabic",
"fielddata":"true"
},
"stem_noshingle":{
"type":"text",
"analyzer":"rebuilt",
"fielddata":"true"
},
"keyword":{
"type":"keyword"
}
}
}
}
}
}
es.indices.create(index="hadith1",ignore=400)
helpers.bulk(es,index='hadith1', actions=hadit)
es.indices.create(index="hadith", body = mapping,ignore=400)
#Verify creation of the Hadith indices
res= es.indices.get_alias("*")
for nomindex in res:
print(nomindex)
es.reindex({"source": {"index": "hadith1"},"dest": {"index": "hadith"}}, wait_for_completion=True, request_timeout=300)
# # Running different types of queries to test the Hadith search engine :
query_body = {
"size":1000,
"query": {
"match": {
"hadith.nostem": {
"query":"الإيمان"
}
}
}
}
es.search(index="hadith", body=query_body)
query_body = {
"size":1000,
"query": {
"match": {
"hadith.nostem": {
"query":"الحياء"
}
}
}
}
es.search(index="hadith", body=query_body)
query_body = {
"query": {
"bool": {
"must": [
{
"match": {
"source": {"query": "صحيح البخاري",
"operator": "and"}
}
},
{
"match": {
"chapter": {"query": "كتاب الإيمان",
"operator": "and"}
}
},
{
"match": {
"hadith": {"query": "مسلمون",
"operator": "and"}
}
},
{
"match": {
"hadith": {"query": "النبيون",
"operator": "and"}
}
}
]
}
}
}
es.search(index="hadith", body=query_body)
| .ipynb_checkpoints/hadith_index-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tutorial for the atomic crystal module in PyXtal
#
# Source code: https://github.com/qzhu2017/PyXtal
#
# Created by <NAME> (2020/11/23)
#
# More details can be found at the following [link](https://pyxtal.readthedocs.io/en/latest/)
#
# # 1.1 Generate a random atomic crystal
from pyxtal import pyxtal
# +
C1 = pyxtal()
C1.from_random(3, 227, ['C'], [8])
# Alternative, you can also generate the structure with pre-assigned sites
# C1.from_random(3, 225, ["C"], [12], sites=[["4a", "8c"]])
print(C1)
# -
C1.atom_sites[0].wp
#display the structure can be easily accessed with the show() function
C1.show(supercell=(2,2,1))
# # 1.2 Manipulating the crystal via symmetry relation
# lower the symmetry from cubic to tetragonal
C2 = C1.subgroup_once(H=141, eps=1e-2)
print(C2)
# Compute the XRD
xrd1 = C1.get_XRD()
xrd2 = C2.get_XRD()
# Compare two structures by XRD
from pyxtal.XRD import Similarity
p1 = xrd1.get_profile()
p2 = xrd2.get_profile()
s = Similarity(p1, p2, x_range=[15, 90])
print(s)
s.show()
# # 1.3 Chemical substitution via symmetry relation
#play substitution
permutation = {"C":"Si"}
SiC = C1.subgroup_once(eps=0.01, H=None, permutations=permutation)
print(C1)
print(SiC)
SiC.show(supercell=(2,2,2))
# # 1.3 Exporting the structure
# +
#In general, you can export the structure in several ways
# CIF/POSCAR file
C1.to_file('1.cif')
C1.to_ase().write('1.vasp', format='vasp', vasp5=True)
# ASE's atoms object
ase_struc = C1.to_ase()
print(ase_struc)
# Pymatgen object
pmg_struc = C1.to_pymatgen()
print(pmg_struc)
# -
# # 1.4 Low dimensional systems
# An example to generate 2D atomic crystal
C3 = pyxtal()
C3.from_random(2, 75, ['C'], [6], thickness=0.0)
C3.show(scale=0.2)
# An example to generate 0D atomic cluster
C4 = pyxtal()
C4.from_random(0, 'Ih', ['C'], [60])
C4.show(scale=0.2)
# to improve the quality, you can increase the minimum distance
C4.from_random(0, 'Ih', ['C'], [60], t_factor=1.2)
C4.show(scale=0.1, radius=0.01)
| examples/tutorials_notebook/01_atomic_crystals.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# # Plot cutouts on the flatmap
#
#
# Cutouts are manually generated cuts of the cortical surface to highlight
# a region of interest.
#
# Cutouts are defined as sub-layers of the `cutouts` layer
# in <filestore>/<subject>/overlays.svg.
#
# The parameter `cutout` of the `quickflat.make_figure` method should be the
# name of the flatmap cutout defined in the `overlays.svg` file.
#
# +
import cortex
import numpy as np
np.random.seed(1234)
# Name of a sub-layer of the 'cutouts' layer in overlays.svg file
cutout_name = "VisualCortexRight"
# Create a random pycortex Volume
volume = cortex.Volume.random(subject='S1', xfmname='fullhead')
# Plot a flatmap with the data projected onto the surface
# Highlight the curvature and which cutout to be displayed
_ = cortex.quickflat.make_figure(volume,
with_curvature=True,
cutout=cutout_name)
| example-notebooks/quickflat/plot_cutouts.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.tree import DecisionTreeRegressor
from sklearn.metrics import r2_score
import matplotlib.pyplot as plt
import seaborn as sns
apple_stock = pd.read_csv('apple_stock_prepared.csv')
apple_stock.head()
X = apple_stock.loc[:, apple_stock.columns != 'close']
Y = apple_stock['close']
X_train, X_test, Y_train, Y_test = train_test_split(X,Y,random_state=42, test_size=0.2)
print("Length Of training data ", len(X_train))
print("Length Of test data ", len(X_test))
print("Length Of training Y ", len(Y_train))
print("Length Of test data Y", len(Y_test))
dtr = DecisionTreeRegressor(max_depth = 2, min_samples_split=5)
dtr.fit(X_train,Y_train)
Y_train_pred = dtr.predict(X_train)
# ##### R Squared on training data
r2_score(Y_train, Y_train_pred)
# #### R Squared on test data
Y_test_pred = dtr.predict(X_test)
r2_score(Y_test, Y_test_pred)
| Section_3/decision_tree.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Machine Learning in Python
# The content for this notebook was copied from [The Deep Learning Machine Learning in Python lab](https://github.com/Microsoft/computerscience/tree/master/Labs/Deep%20Learning/200%20-%20Machine%20Learning%20in%20Python).
# This demo shows prediction of flight delays between airport pairs based on the day of the month using a random forest.
# The demo concludes by visualizing the probability of on-time arrival between JFK and Atlanta Hartsfield-Jackson oves consecutive days.
# In this exercise, you will import a dataset from Azure blob storage and load it into the notebook. Jupyter notebooks are highly interactive, and since they can include executable code, they provide the perfect platform for manipulating data and building predictive models from it.
# ## Ingest
# cURL is a familiar tool to transfer data to or from servers using familiar protocols such as http, https, ftp, ftps, etc.
# In the code cell below cURL is used to download Flight Data from a public blob storage to the working directory.
# !curl https://topcs.blob.core.windows.net/public/FlightData.csv -o flightdata.csv
# Pandas will be used here to create a data frame in which the data will be manipulated and massaged for enhanced analysis.
#
# Import the data and create a pandas DataFrame from it, and display the first five rows.
# +
import pandas as pd
df = pd.read_csv('flightdata.csv')
df.head()
# -
# The DataFrame that you created contains on-time arrival information for a major U.S. airline. It has more than 11,000 rows and 26 columns. (The output says "5 rows" because DataFrame's head function only returns the first five rows.) Each row represents one flight and contains information such as the origin, the destination, the scheduled departure time, and whether the flight arrived on time or late. You will learn more about the data, including its content and structure, in the next lab.
# ## Process
# In the real world, few datasets can be used as is to train machine-learning models. It is not uncommon for data scientists to spend 80% or more of their time on a project cleaning, preparing, and shaping the data — a process sometimes referred to as data wrangling. Typical actions include removing duplicate rows, removing rows or columns with missing values or algorithmically replacing the missing values, normalizing data, and selecting feature columns. A machine-learning model is only as good as the data it is trained with. Preparing the data is arguably the most crucial step in the machine-learning process.
#
# Before you can prepare a dataset, you need to understand its content and structure. In the previous steps, you imported a dataset containing on-time arrival information for a major U.S. airline. That data included 26 columns and thousands of rows, with each row representing one flight and containing information such as the flight's origin, destination, and scheduled departure time. You also loaded the data into the Jupyter notebook and used a simple Python script to create a pandas DataFrame from it.
#
# To get a count of rows, run the following code:
df.shape
# Now take a moment to examine the 26 columns in the dataset. They contain important information such as the date that the flight took place (YEAR, MONTH, and DAY_OF_MONTH), the origin and destination (ORIGIN and DEST), the scheduled departure and arrival times (CRS_DEP_TIME and CRS_ARR_TIME), the difference between the scheduled arrival time and the actual arrival time in minutes (ARR_DELAY), and whether the flight was late by 15 minutes or more (ARR_DEL15).
#
# Here is a complete list of the columns in the dataset. Times are expressed in 24-hour military time. For example, 1130 equals 11:30 a.m. and 1500 equals 3:00 p.m.
# One of the first things data scientists typically look for in a dataset is missing values. There's an easy way to check for missing values in pandas. To demonstrate, execute the following code:
df.isnull().values.any()
# The next step is to find out where the missing values are. To do so, execute the following code:
df.isnull().sum()
# Curiously, the 26th column ("Unnamed: 25") contains 11,231 missing values, which equals the number of rows in the dataset. This column was mistakenly created because the CSV file that you imported contains a comma at the end of each line. To eliminate that column, execute the following code:
df = df.drop('Unnamed: 25', axis=1)
df.isnull().sum()
# The DataFrame still contains a lot of missing values, but some of them are irrelevant because the columns containing them are not germane to the model that you are building. The goal of that model is to predict whether a flight you are considering booking is likely to arrive on time. If you know that the flight is likely to be late, you might choose to book another flight.
#
# The next step, therefore, is to filter the dataset to eliminate columns that aren't relevant to a predictive model. For example, the aircraft's tail number probably has little bearing on whether a flight will arrive on time, and at the time you book a ticket, you have no way of knowing whether a flight will be cancelled, diverted, or delayed. By contrast, the scheduled departure time could have a lot to do with on-time arrivals. Because of the hub-and-spoke system used by most airlines, morning flights tend to be on time more often than afternoon or evening flights. And at some major airports, traffic stacks up during the day, increasing the likelihood that later flights will be delayed.
#
# Pandas provides an easy way to filter out columns you don't want. Execute the following code:
df = df[["MONTH", "DAY_OF_MONTH", "DAY_OF_WEEK", "ORIGIN", "DEST", "CRS_DEP_TIME", "ARR_DEL15"]]
df.isnull().sum()
# The only column that now contains missing values is the ARR_DEL15 column, which uses 0s to identify flights that arrived on time and 1s for flights that didn't. Use the following code to show the first five rows with missing values:
df[df.isnull().values.any(axis=1)].head()
# The reason these rows are missing ARR_DEL15 values is that they all correspond to flights that were canceled or diverted. You could call dropna on the DataFrame to remove these rows. But since a flight that is canceled or diverted to another airport could be considered "late," let's use the fillna method to replace the missing values with 1s.
#
# Use the following code to replace missing values in the ARR_DEL15 column with 1s and display rows 177 through 184:
df = df.fillna({'ARR_DEL15': 1})
df.iloc[177:185]
# Use the following code to display the first five rows of the DataFrame:
df.head()
# The CRS_DEP_TIME column of the dataset you are using represents scheduled departure times. The granularity of the numbers in this column — it contains more than 500 unique values — could have a negative impact on accuracy in a machine-learning model. This can be resolved using a technique called binning or quantization. What if you divided each number in this column by 100 and rounded down to the nearest integer? 1030 would become 10, 1925 would become 19, and so on, and you would be left with a maximum of 24 discrete values in this column. Intuitively, it makes sense, because it probably doesn't matter much whether a flight leaves at 10:30 a.m. or 10:40 a.m. It matters a great deal whether it leaves at 10:30 a.m. or 5:30 p.m.
#
# In addition, the dataset's ORIGIN and DEST columns contain airport codes that represent categorical machine-learning values. These columns need to be converted into discrete columns containing indicator variables, sometimes known as "dummy" variables. In other words, the ORIGIN column, which contains five airport codes, needs to be converted into five columns, one per airport, with each column containing 1s and 0s indicating whether a flight originated at the airport that the column represents. The DEST column needs to be handled in a similar manner.
#
# In this portion of the exercise, you will "bin" the departure times in the CRS_DEP_TIME column and use pandas' get_dummies method to create indicator columns from the ORIGIN and DEST columns.
#
# Use the following code to bin the departure times:
# +
import math
for index, row in df.iterrows():
df.loc[index, 'CRS_DEP_TIME'] = math.floor(row['CRS_DEP_TIME'] / 100)
df.head()
# -
# Now use the following statements to generate indicator columns from the ORIGIN and DEST columns, while dropping the ORIGIN and DEST columns themselves:
df = pd.get_dummies(df, columns=['ORIGIN', 'DEST'])
df.head()
# ## Predict
# Machine learning, which facilitates predictive analytics using large volumes of data by employing algorithms that iteratively learn from that data, is one of the fastest growing areas of data science.
#
# One of the most popular tools for building machine-learning models is Scikit-learn, a free and open-source toolkit for Python programmers. It has built-in support for popular regression, classification, and clustering algorithms and works with other Python libraries such as NumPy and SciPy. With Sckit-learn, a simple method call can replace hundreds of lines of hand-written code. Sckit-learn allows you to focus on building, training, tuning, and testing machine-learning models without getting bogged down coding algorithms.
#
# In this lab, the third of four in a series, you will use Sckit-learn to build a machine-learning model utilizing on-time arrival data for a major U.S. airline. The goal is to create a model that might be useful in the real world for predicting whether a flight is likely to arrive on time. It is precisely the kind of problem that machine learning is commonly used to solve. And it's a great way to increase your machine-learning chops while getting acquainted with Scikit-learn.
# The first statement imports Sckit-learn's train_test_split helper function. The second line uses the function to split the DataFrame into a training set containing 80% of the original data, and a test set containing the remaining 20%. The random_state parameter seeds the random-number generator used to do the splitting, while the first and second parameters are DataFrames containing the feature columns and the label column.
from sklearn.model_selection import train_test_split
train_x, test_x, train_y, test_y = train_test_split(df.drop('ARR_DEL15', axis=1), df['ARR_DEL15'], test_size=0.2, random_state=42)
# train_test_split returns four DataFrames. Use the following command to display the number of rows and columns in the DataFrame containing the feature columns used for training:
train_x.shape
# Now use this command to display the number of rows and columns in the DataFrame containing the feature columns used for testing:
test_x.shape
# You will train a classification model, which seeks to resolve a set of inputs into one of a set of known outputs.
#
# Sckit-learn includes a variety of classes for implementing common machine-learning models. One of them is RandomForestClassifier, which fits multiple decision trees to the data and uses averaging to boost the overall accuracy and limit overfitting.
#
# Execute the following code to create a RandomForestClassifier object and train it by calling the fit method.
# +
from sklearn.ensemble import RandomForestClassifier
model = RandomForestClassifier(random_state=13)
model.fit(train_x, train_y)
# -
# The output shows the parameters used in the classifier, including n_estimators, which specifies the number of trees in each decision-tree forest, and max_depth, which specifies the maximum depth of the decision trees. The values shown are the defaults, but you can override any of them when creating the RandomForestClassifier object.
#
# Now call the predict method to test the model using the values in test_x, followed by the score method to determine the mean accuracy of the model:
predicted = model.predict(test_x)
model.score(test_x, test_y)
# There are several ways to measure the accuracy of a classification model. One of the best overall measures for a binary classification model is Area Under Receiver Operating Characteristic Curve (sometimes referred to as "ROC AUC"), which essentially quantifies how often the model will make a correct prediction regardless of the outcome. In this exercise, you will compute an ROC AUC score for the model you built in the previous exercise and learn about some of the reasons why that score is lower than the mean accuracy output by the score method. You will also learn about other ways to gauge the accuracy of the model.
#
# Before you compute the ROC AUC, you must generate prediction probabilities for the test set. These probabilities are estimates for each of the classes, or answers, the model can predict. For example, [0.88199435, 0.11800565] means that there's an 89% chance that a flight will arrive on time (ARR_DEL15 = 0) and a 12% chance that it won't (ARR_DEL15 = 1). The sum of the two probabilities adds up to 100%.
#
# Run the following code to generate a set of prediction probabilities from the test data:
from sklearn.metrics import roc_auc_score
probabilities = model.predict_proba(test_x)
# Now use the following statement to generate an ROC AUC score from the probabilities using Sckit-learn's roc_auc_score method:
roc_auc_score(test_y, probabilities[:, 1])
# Why is the AUC score lower than the mean accuracy computed in the previous exercise?
#
# The output from the score method reflects how many of the items in the test set the model predicted correctly. This score is skewed by the fact that the dataset the model was trained and tested with contains many more rows representing on-time arrivals than rows representing late arrivals. Because of this imbalance in the data, you are more likely to be correct if you predict that a flight will be on time than if you predict that a flight will be late.
#
# ROC AUC takes this into account and provides a more accurate indication of how likely it is that a prediction of on-time or late will be correct.
# You can learn more about the model's behavior by generating a confusion matrix, also known as an error matrix. The confusion matrix quantifies the number of times each answer was classified correctly or incorrectly. Specifically, it quantifies the number of false positives, false negatives, true positives, and true negatives. This is important, because if a binary classification model trained to recognize cats and dogs is tested with a dataset that is 95% dogs, it could score 95% simply by guessing "dog" every time. But if it failed to identify cats at all, it would be of little value.
#
# Use the following code to produce a confusion matrix for your model:
from sklearn.metrics import confusion_matrix
confusion_matrix(test_y, predicted)
# The first row in the output represents flights that were on time. The first column in that row shows how many flights were correctly predicted to be on time, while the second column reveals how many flights were predicted as delayed but were not. From this, the model appears to be very adept at predicting that a flight will be on time.
#
# But look at the second row, which represents flights that were delayed. The first column shows how many delayed flights were incorrectly predicted to be on time. The second column shows how many flights were correctly predicted to be delayed. Clearly, the model isn't nearly as adept at predicting that a flight will be delayed as it is at predicting that a flight will arrive on time. What you want in a confusion matrix is big numbers in the upper-left and lower-right corners, and small numbers (preferably zeros) in the upper-right and lower-left corners.
# Other measures of accuracy for a classification model include precision and recall. Suppose the model was presented with three on-time arrivals and three delayed arrivals, and that it correctly predicted two of the on-time arrivals, but incorrectly predicted that two of the delayed arrivals would be on time. In this case, the precision would be 50% (two of the four flights it classified as being on time actually were on time), while its recall would be 67% (it correctly identified two of the three on-time arrivals). You can learn more about precision and recall from https://en.wikipedia.org/wiki/Precision_and_recall
#
# Sckit-learn contains a handy method named precision_score for computing precision. To quantify the precision of your model, execute the following statements:
# +
from sklearn.metrics import precision_score
train_predictions = model.predict(train_x)
precision_score(train_y, train_predictions)
# -
# Sckit-learn also contains a method named recall_score for computing recall. To measure you model's recall, execute the following statements:
# +
from sklearn.metrics import recall_score
recall_score(train_y, train_predictions)
# -
# ## Visualize
#
# Now that you that have trained a machine-learning model to perform predictive analytics, it's time to put it to work. In this lab, the final one in the series, you will write a function that uses the machine-learning model you built in the previous lab to predict whether a flight will arrive on time or late. And you will use Matplotlib, the popular plotting and charting library for Python, to visualize the results.
# The first statement is one of several magic commands supported by the Python kernel that you selected when you created the notebook. It enables Jupyter to render Matplotlib output in a notebook without making repeated calls to show. And it must appear before any references to Matplotlib itself. The final statement configures Seaborn to enhance the output from Matplotlib.
# Execute the following code. Ignore any warning messages that are displayed related to font caching:
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sns
sns.set()
# -
# The first statement is one of several magic commands supported by the Python kernel that you selected when you created the notebook. It enables Jupyter to render Matplotlib output in a notebook without making repeated calls to show. And it must appear before any references to Matplotlib itself. The final statement configures Seaborn to enhance the output from Matplotlib.
#
# To see Matplotlib at work, execute the following code in a new cell to plot the ROC curve for the machine-learning model you built in the previous lab:
# +
from sklearn.metrics import roc_curve
fpr, tpr, _ = roc_curve(test_y, probabilities[:, 1])
plt.plot(fpr, tpr)
plt.plot([0, 1], [0, 1], color='grey', lw=1, linestyle='--')
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
# -
# The dotted line in the middle of the graph represents a 50-50 chance of obtaining a correct answer. The blue curve represents the accuracy of your model. More importantly, the fact that this chart appears at all demonstrates that you can use Matplotlib in a Jupyter notebook.
# The reason you built a machine-learning model is to predict whether a flight will arrive on time or late. In this exercise, you will write a Python function that calls the machine-learning model you built in the previous lab to compute the likelihood that a flight will be on time. Then you will use the function to analyze several flights.
#
# This function takes as input a date and time, an origin airport code, and a destination airport code, and returns a value between 0.0 and 1.0 indicating the probability that the flight will arrive at its destination on time. It uses the machine-learning model you built in the previous lab to compute the probability. And to call the model, it passes a DataFrame containing the input values to predict_proba. The structure of the DataFrame exactly matches the structure of the DataFrame depicted in previous steps.
def predict_delay(departure_date_time, origin, destination):
from datetime import datetime
try:
departure_date_time_parsed = datetime.strptime(departure_date_time, '%d/%m/%Y %H:%M:%S')
except ValueError as e:
return 'Error parsing date/time - {}'.format(e)
month = departure_date_time_parsed.month
day = departure_date_time_parsed.day
day_of_week = departure_date_time_parsed.isoweekday()
hour = departure_date_time_parsed.hour
origin = origin.upper()
destination = destination.upper()
input = [{'MONTH': month,
'DAY': day,
'DAY_OF_WEEK': day_of_week,
'CRS_DEP_TIME': hour,
'ORIGIN_ATL': 1 if origin == 'ATL' else 0,
'ORIGIN_DTW': 1 if origin == 'DTW' else 0,
'ORIGIN_JFK': 1 if origin == 'JFK' else 0,
'ORIGIN_MSP': 1 if origin == 'MSP' else 0,
'ORIGIN_SEA': 1 if origin == 'SEA' else 0,
'DEST_ATL': 1 if destination == 'ATL' else 0,
'DEST_DTW': 1 if destination == 'DTW' else 0,
'DEST_JFK': 1 if destination == 'JFK' else 0,
'DEST_MSP': 1 if destination == 'MSP' else 0,
'DEST_SEA': 1 if destination == 'SEA' else 0 }]
return model.predict_proba(pd.DataFrame(input))[0][0]
# Use the code below to compute the probability that a flight from New York to Atlanta on the evening of October 1 will arrive on time. Note that the year you enter is irrelevant because it isn't used by the model.
predict_delay('1/10/2018 21:45:00', 'JFK', 'ATL')
# Modify the code to compute the probability that the same flight a day later will arrive on time:
predict_delay('2/10/2018 21:45:00', 'JFK', 'ATL')
# How likely is this flight to arrive on time? If your travel plans were flexible, would you consider postponing your trip for one day?
# Now modify the code to compute the probability that a morning flight the same day from Atlanta to Seattle will arrive on time:
predict_delay('2/10/2018 10:00:00', 'ATL', 'SEA')
# In this exercise, you will combine the predict_delay function you created in the previous exercise with Matplotlib to produce side-by-side comparisons of the same flight on consecutive days and flights with the same origin and destination at different times throughout the day.
# +
import numpy as np
labels = ('Oct 1', 'Oct 2', 'Oct 3', 'Oct 4', 'Oct 5', 'Oct 6', 'Oct 7')
values = (predict_delay('1/10/2018 21:45:00', 'JFK', 'ATL'),
predict_delay('2/10/2018 21:45:00', 'JFK', 'ATL'),
predict_delay('3/10/2018 21:45:00', 'JFK', 'ATL'),
predict_delay('4/10/2018 21:45:00', 'JFK', 'ATL'),
predict_delay('5/10/2018 21:45:00', 'JFK', 'ATL'),
predict_delay('6/10/2018 21:45:00', 'JFK', 'ATL'),
predict_delay('7/10/2018 21:45:00', 'JFK', 'ATL'))
alabels = np.arange(len(labels))
plt.bar(alabels, values, align='center', alpha=0.5)
plt.xticks(alabels, labels)
plt.ylabel('Probability of On-Time Arrival')
plt.ylim((0.0, 1.0))
# -
# Referenced from https://github.com/Microsoft/computerscience/tree/master/Labs/Deep%20Learning/200%20-%20Machine%20Learning%20in%20Python, 12/17/2018
| workshop-resources/data-science-and-machine-learning/Data_Science_2/workshop-materials/Machine_Learning_in_Python_Demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.10 64-bit
# language: python
# name: python3
# ---
# +
# 'yfinance' for extracting price data
import yfinance as module_yf
# 'pandas' for manipulating tabular data
import pandas as module_pd
# 'numpy' for numeric array manipulation
import numpy as module_np
# 'datetime' for manipulating 'datetime' data types
import datetime as module_dt
# 'matplotlib.pyplot' for creating plots
import matplotlib.pyplot as module_plt
# 'tb_custompack' for custom functions
import tb_custompack as module_tb
# -
# Set specified ticker sybol from
# which data will be extracted
ticker_symbol = module_yf.Ticker(
ticker = "AMZN",
)
# Print out dictionary for information
# about ticker symbol
dict_ticker_info = ticker_symbol.info
# +
# Extract historical price data for ticker
dataframe_ticker_data = ticker_symbol.history(
interval = '1d',
period = 'max'
).dropna()
# Set the number of observations
int_data_length = len(dataframe_ticker_data)
# +
# Reset the indexing of the data frame, so new
# 'Date' column is generated
dataframe_prep_1 = dataframe_ticker_data.reset_index()
# Isolate the candlestick price data
dataframe_prep_f = dataframe_prep_1[
[
'Open',
'High',
'Low',
'Close'
]
]
# +
# Create an intitial value for Heikin Ashi
float_initial = (
dataframe_prep_f.iloc[0][1] +
dataframe_prep_f.iloc[0][2] -
dataframe_prep_f.iloc[0][0] -
dataframe_prep_f.iloc[0][3]
) / 4
# Create list of Heikin Ashi closes from
# candlestick price data
list_hclose = list(module_np.mean(module_np.log(dataframe_prep_f.T)))
# Create Heikin Ashi indicator list
list_ha = module_tb.function_heikin_ashi_gen(
list_values = list_hclose,
float_initial = float_initial,
float_strength = 1/2
)
# +
# Add indicator to standard indexed dataframe
dataframe_prep_1['Indicator'] = module_pd.Series(list_ha)
# Reassign 'Date' as index series for final dataframe
dataframe_ticker_data = dataframe_prep_1.set_index('Date')
# +
# Set the initial trading time
datetime_start = module_dt.datetime(2021,1,1)
# Set the final trading time
datetime_end = module_dt.datetime(2022,3,1)
# Select all price and indicator data
# falling within the chosen time bound
dataframe_selected_data = dataframe_ticker_data.loc[
datetime_start:datetime_end
].reset_index()
# Create list of close prices
list_close = list(dataframe_selected_data['Close'])
# Create list for long indicator
list_long_indicator = list(dataframe_selected_data['Indicator'])
# Create list for short indicator
list_short_indicator = list(-dataframe_selected_data['Indicator'])
# Create list for dates
list_dates = list(dataframe_selected_data['Date'])
# -
# Create alpha series for long position
list_capital_1 = module_tb.function_backtester(
list_prices = list_close,
list_indicator = list_long_indicator,
float_ratio = 1
)
# Create alpha series for short position
list_capital_2 = module_tb.function_backtester(
list_prices = list_close,
list_indicator = list_short_indicator,
float_ratio = -1
)
# Create full alpha series
list_capital_3 = list(module_np.multiply(list_capital_1, list_capital_2))
# +
# Code for plot creation
module_plt.figure(
num = 0,
figsize = (8 * (16 / 9), 8)
)
module_plt.plot(
list_dates,
module_np.log(module_np.cumprod(list_capital_1)),
label = "Long only"
)
module_plt.plot(
list_dates,
module_np.log(module_np.cumprod(list_capital_2)),
label = "Short only"
)
module_plt.plot(
list_dates,
module_np.log(module_np.cumprod(list_capital_3)),
label = "Long and short"
)
module_plt.plot(
list_dates,
module_np.log(module_np.array(list_close) / list_close[0]),
label = "No strategy"
)
module_plt.xlabel("Date")
module_plt.ylabel("log(Value) of portfolio")
module_plt.title("Plot comparison of variations of same strategy")
module_plt.legend()
module_plt.show()
module_plt.figure(
num = 0,
figsize = (8 * (16 / 9), 8)
)
module_plt.plot(
dataframe_selected_data['Date'],
dataframe_selected_data['Indicator']
)
module_plt.plot(
dataframe_selected_data['Date'],
0 * dataframe_selected_data['Indicator']
)
module_plt.xlabel("Date")
module_plt.ylabel("Indicator value")
module_plt.title("Plot of indicator against time")
module_plt.show()
| 6th_week_v1/code/backtester.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Warren 2020 Models
#
# Data from "Constraining properties of the next nearby core-collapse supernova with multi-messenger signals: multi-messenger signals" by <NAME>; Couch, Sean; <NAME>; <NAME>.
#
# 1D FLASH simulations with STIR, for alpha_lambda = 1.23, 1.25, and 1.27. Run with SFHo EOS, M1 with 12 energy groups.
#
# For more information on these simulations, see Warren, Couch, O'Connor, & Morozova (arXiv:1912.03328) and Couch, Warren, & O'Connor (2020).
#
# Includes the multi-messenger data from the STIR simulations. The filename indicates the turbulent mixing parameter a and progenitor mass m of the simulation. Columns are time [s], shock radius [cm], explosion energy [ergs], electron neutrino mean energy [MeV], electron neutrino rms energy [MeV], electron neutrino luminosity [10^51 ergs/s], electron antineutrino mean energy [MeV], electron antineutrino rms energy [MeV], electron antineutrino luminosity [10^51 ergs/s], x neutrino mean energy [MeV], x neutrino rms energy [MeV], x neutrino luminosity [10^51 ergs/s], gravitational wave frequency from eigenmode analysis of the protoneutron star structure [Hz]. Note that the x neutrino luminosity is for one neutrino flavor - to get the total mu/tau neutrino and antineutrino luminosities requires multiplying this number by 4.
# +
import matplotlib as mpl
import matplotlib.pyplot as plt
import numpy as np
from astropy import units as u
from snewpy.neutrino import Flavor, MassHierarchy
from snewpy.models import Warren_2020
from snewpy.flavor_transformation import NoTransformation, AdiabaticMSW, ThreeFlavorDecoherence
# -
mpl.rc('font', size=16)
def plot_luminosity(model):
fig, ax = plt.subplots(1, figsize=(8,6))
for flavor in Flavor:
ax.plot(model.time, model.luminosity[flavor]/1e51, # Report luminosity in units foe/s
label=flavor.to_tex(),
color = 'C0' if flavor.is_electron else 'C1',
ls = '-' if flavor.is_neutrino else ':',
lw = 2 )
ax.set(xlim=(-0.05, 0.5),
xlabel=r'$t-t_{\rm bounce}$ [s]',
ylabel=r'luminosity [foe s$^{-1}$]',
title=model.filename)
ax.grid()
ax.legend(loc='upper right', ncol=2, fontsize=18)
fig.tight_layout()
return fig
# ## LS220 EOS: 10.0 $M_\odot$, $a=1.23$
ifile = '../../models/Warren_2020/stir_a1.23/stir_multimessenger_a1.23_m10.0.h5'
model = Warren_2020(ifile)
model
fig = plot_luminosity(model)
# ## LS220 EOS: 25.0 $M_\odot$, $a=1.23$
ifile = '../../models/Warren_2020/stir_a1.23/stir_multimessenger_a1.23_m25.0.h5'
model = Warren_2020(ifile)
model
fig = plot_luminosity(model)
# ## LS220 EOS: 10.0 $M_\odot$, $a=1.27$
ifile = '../../models/Warren_2020/stir_a1.27/stir_multimessenger_a1.27_m10.0.h5'
model = Warren_2020(ifile)
model
fig = plot_luminosity(model)
# ## LS220 EOS: 25.0 $M_\odot$, $a=1.27$
ifile = '../../models/Warren_2020/stir_a1.27/stir_multimessenger_a1.27_m25.0.h5'
model = Warren_2020(ifile)
model
# +
fig, ax = plt.subplots(1, figsize=(8,6))
for flavor in Flavor:
ax.plot(model.time, model.luminosity[flavor]/1e51, # Report luminosity in units foe/s
label=flavor.to_tex(),
color = 'C0' if flavor.is_electron else 'C1',
ls = '-' if flavor.is_neutrino else ':',
lw = 2 )
ax.set(xlim=(-0.05, 0.5),
xlabel=r'$t-t_{\rm bounce}$ [s]',
ylabel=r'luminosity [foe s$^{-1}$]')
ax.grid()
ax.legend(loc='upper right', ncol=2, fontsize=18)
fig.tight_layout();
# -
# ## Initial and Oscillated Spectra
#
# Plot the neutrino spectra at the source and after the requested flavor transformation has been applied.
#
# ### Adiabatic MSW Flavor Transformation: Normal mass ordering
# +
# Adiabatic MSW effect. NMO is used by default.
xform_nmo = AdiabaticMSW()
# Energy array and time to compute spectra.
# Note that any convenient units can be used and the calculation will remain internally consistent.
E = np.linspace(0,100,201) * u.MeV
t = 50*u.ms
ispec = model.get_initialspectra(t, E)
ospec_nmo = model.get_oscillatedspectra(t, E, xform_nmo)
# +
fig, axes = plt.subplots(1,2, figsize=(12,5), sharex=True, sharey=True, tight_layout=True)
for i, spec in enumerate([ispec, ospec_nmo]):
ax = axes[i]
for flavor in Flavor:
ax.plot(E, spec[flavor],
label=flavor.to_tex(),
color='C0' if flavor.is_electron else 'C1',
ls='-' if flavor.is_neutrino else ':', lw=2,
alpha=0.7)
ax.set(xlabel=r'$E$ [{}]'.format(E.unit),
title='Initial Spectra: $t = ${:.1f}'.format(t) if i==0 else 'Oscillated Spectra: $t = ${:.1f}'.format(t))
ax.grid()
ax.legend(loc='upper right', ncol=2, fontsize=16)
ax = axes[0]
ax.set(ylabel=r'flux [erg$^{-1}$ s$^{-1}$]')
fig.tight_layout();
# -
| doc/nb/Warren_2020.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + slideshow={"slide_type": "skip"}
# %matplotlib inline
# + slideshow={"slide_type": "skip"} active=""
# <style type="text/css">
# .reveal h1, .reveal h2 {
# font-family:"League Gothic"
# }
# </style>
# + [markdown] slideshow={"slide_type": "notes"}
# Hello everyone! My name is <NAME> and today I am here to talk to about "Doing Math with Python".
#
# Thank you for coming to my talk - i know you could have chosen the other talk, so it's good to know that my talk's topic interests you.
#
# A bit about me - I am a software engineer at Freelancer.com in Sydney Australia. I am a fairly regular writer for Linux Voice and other Linux magazines. And last, but not the least, I am the author of the book "Doing Math with Python" (not coincidentally titled the same as this talk - haha) published by No Starch Press in 2015.
#
# There is a link to my blog, GitHub, twitter, etc. so, if you want to learn more about my work or get in touch, those are the means to do so!
#
# Okay, so all that aside - let's start with the talk!
# + [markdown] slideshow={"slide_type": "slide"}
# # <center> My first lab </center>
#
# ### <center> <NAME> </center>
#
# #### <center> June 15th 2017 </center>
#
# #### <center> Cape Town, SA </center>
# + [markdown] slideshow={"slide_type": "subslide"}
# ## About me
#
# - 2016/17 Mozilla Fellow for Science
#
# - Research Fellow at the Alan Turing Institute for Data Science, London
#
# - Research Associate in the Department of Psychiatry at the University of Cambridge
#
#
# ### Contact
#
# - Twitter: [@kirstie_j](http://twitter.com/kirstie_j)
#
# - Email: [<EMAIL>](mailto:<EMAIL>)
#
# - GitHub: [KirstieJane](http://github.com/KirstieJane)
# + [markdown] slideshow={"slide_type": "slide"}
# <a href="http://hakim.se" data-preview-link><NAME></a>
#
# Hello!
#
# + [markdown] slideshow={"slide_type": "notes"}
# So, what am I selling to you today? Not my book (I am, but in a subtle way). I am presenting an idea, a hypothesis or even making a statement - Python can lead to a more enriching learning and teaching experience in the classroom.
#
# Let me explain where I am coming from. When I think back about when I was learning to program and learning all other subjects in standards 7-10. I think it's true today as well. Programming and other subjects such as Math, Science are taught in a disconnected fashion. Programming seems to be all about finding the sum of a series or generating fibonacci numbers. Make no mistake, these exercises are what builds up the programming logic. Some students get really excited about being able to do these, but a lot of them don't. It's a lot like not everyone gets interested in solving puzzles - i don't, i never took to them.
#
# I think I know of a way we could excite more students! Show them how you can write programs to do your homework, or experiment without having to go the science lab or setup elaborate experimental setups. This is my goal for today - in the following slides and notebooks, I will hypothesise on a way of connecting Python programming and other subjects. That will show that programming is a way to get real work done, not something to learn for the sake of it.
#
# We need some tools to help us on our quest. The Python community has some giant shoulders we can stand upon - Python 3, SymPy and matplotlib.
# + [markdown] slideshow={"slide_type": "slide"}
#
# ### This talk - a proposal, a hypothesis, a statement
#
# What? *Python can lead to a more enriching learning and teaching experience in the classroom*
#
#
#
# How? *Next slides*
#
#
#
#
#
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Tools (or, Giant shoulders we will stand on)
#
# <img align="center" src="images/giant_shoulders_collage.jpg"></img>
#
#
# *Python 3*, *SymPy*, *matplotlib*
#
#
# *Individual logos are copyright of the respective projects. [Source](http://www.orphancaremovement.org/standing-on-the-shoulder-of-giants/) of the "giant shoulders" image.
#
# + [markdown] slideshow={"slide_type": "notes"}
# Whose calculator looks like this?
#
# Who uses Python as a calculator? Raise of hands please!
#
# I do! Specifically, I use Python 3 because of 1/2=0 messes up my monthly expenditure calculation.
#
# Besides the usual addition and subtraction, we have of course the math module and more recently the statistics module which makes Python a worthy scientific calculator.
#
# But then, there's more! You are not limited to the functions from those libraries, you can write your own custom functions and make them available whenever you start your Python interpreter. How?
#
# Use PYTHONSTARTUP!
# + [markdown] slideshow={"slide_type": "slide"}
# ### Python - a scientific calculator
#
#
#
# Whose calculator looks like this?
#
# ```python
# >>> (131 + 21.5 + 100.2 + 88.7 + 99.5 + 100.5 + 200.5)/4
# 185.475
# ```
#
#
# *Python 3 is my favorite calculator (not Python 2 because 1/2 = 0)*
#
# Beyond basic operations:
#
# + [markdown] slideshow={"slide_type": "fragment"}
#
# - `fabs()`, `abs()`, `sin()`, `cos()`, `gcd()`, `log()` and more (See [math](https://docs.python.org/3/library/math.html))
#
# - Descriptive statistics (See [statistics](https://docs.python.org/3/library/statistics.html#module-statistics))
#
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Python - a scientific calculator
#
# - Develop your own functions: unit conversion, finding correlation, .., anything really
#
# - Use PYTHONSTARTUP to extend the battery of readily available mathematical functions
#
# ```python
# $ PYTHONSTARTUP=~/work/dmwp/pycon-us-2016/startup_math.py idle3 -s
# ```
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
#
# ### Unit conversion functions
#
# ```python
#
# >>> unit_conversion()
# 1. Kilometers to Miles
# 2. Miles to Kilometers
# 3. Kilograms to Pounds
# 4. Pounds to Kilograms
# 5. Celsius to Fahrenheit
# 6. Fahrenheit to Celsius
# Which conversion would you like to do? 6
# Enter temperature in fahrenheit: 98
# Temperature in celsius: 36.66666666666667
# >>>
#
# ```
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Finding linear correlation
#
# ```python
# >>>
# >>> x = [1, 2, 3, 4]
# >>> y = [2, 4, 6.1, 7.9]
# >>> find_corr_x_y(x, y)
# 0.9995411791453812
#
# ```
# + [markdown] slideshow={"slide_type": "notes"}
# So, that was Python and it's standard libraries. When you bring in third party libraries to the mix, Python becomes a seriously fancy calculator.
#
# Who has heard about SymPy?
#
# You can give it algebraic expressions to a function and a graph will be created for you.
#
# You can give an equation and out comes the solutions for that equation.
#
# We can even solve calculus problems.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Python - a really fancy calculator
#
# SymPy - a pure Python symbolic math library
#
# *from sympy import awesomeness* - don't try that :)
#
#
#
# + slideshow={"slide_type": "subslide"}
# Create graphs from algebraic expressions
from sympy import Symbol, plot
x = Symbol('x')
p = plot(2*x**2 + 2*x + 2)
# + slideshow={"slide_type": "subslide"}
# Solve equations
from sympy import solve, Symbol
x = Symbol('x')
solve(2*x + 1)
# + slideshow={"slide_type": "subslide"}
# Limits
from sympy import Symbol, Limit, sin
x = Symbol('x')
Limit(sin(x)/x, x, 0).doit()
# + slideshow={"slide_type": "subslide"}
# Derivative
from sympy import Symbol, Derivative, sin, init_printing
x = Symbol('x')
init_printing()
Derivative(sin(x)**(2*x+1), x).doit()
# + slideshow={"slide_type": "subslide"}
# Indefinite integral
from sympy import Symbol, Integral, sqrt, sin, init_printing
x = Symbol('x')
init_printing()
Integral(sqrt(x)).doit()
# + slideshow={"slide_type": "fragment"}
# Definite integral
from sympy import Symbol, Integral, sqrt
x = Symbol('x')
Integral(sqrt(x), (x, 0, 2)).doit()
# + [markdown] slideshow={"slide_type": "notes"}
# I will pause for a moment now. In the first two slides, we have seen how Python can be a super awesome calculator. What does that buy us? We have now been able to show that you can make computer programs literally do your homework. Write a program to do your work once and you will never have to make those lengthy calculations yourselves. Can we use Python to do more?
#
# Let's continue.
# + [markdown] slideshow={"slide_type": "slide"}
# <center><h1>Can we do more than write smart calculators?</h1></center>
#
#
# + [markdown] slideshow={"slide_type": "notes"}
# Python can be more than a super powerful calculator. We can use it to enhance the learning experience of other subjects. Next, I have three examples including a demo. First up, a video of a projectile motion. This program uses matplotlib's animation API to create a basic animation of a projectile motion - a fairly common subject introduced in introductory Physics. The program which is linked asks for the angle of projection and speed and then draws the trajectory of the projectile. Just by running the program multiple times, we can see how the trajectory changes. We don't have to go outside and start throwing balls..
#
# Next, we will put Jupyter Notebook's interactive widgets to good effect by drawing a Barnsley Fern. Let's see how the demo goes.
#
# Next, with the help of basemap, we can draw places on a world map like we would draw points on a graph paper.
#
# I know I would be excited if someone was showing me all these cool things when I was learning these things!
# + [markdown] slideshow={"slide_type": "slide"}
#
# ### Python - Making other subjects more lively
#
# <img align="center" src="images/collage1.png"></img>
#
#
# - matplotlib
#
# - basemap
#
# - Interactive Jupyter Notebooks
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
#
# #### Bringing Science to life
#
# *Animation of a Projectile motion* [(Python Source)](https://github.com/doingmathwithpython/pycon-us-2016/blob/master/py-files/projectile_animation.py)
#
#
#
# -
from IPython.display import YouTubeVideo
YouTubeVideo("8uWRVh58KdQ")
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Exploring Fractals in Nature
#
# *Interactively drawing a Barnsley Fern* [(Notebook)](https://github.com/doingmathwithpython/pycon-us-2016/blob/master/notebooks/Interactive%20Barnsley%20Fern.ipynb)
#
#
#
# <img align="center" src="images/fern.jpg" width="400" ></img>
#
# + [markdown] slideshow={"slide_type": "subslide"}
#
# #### The world is your graph paper
#
# *Showing places on a digital map* [(Notebook)](https://github.com/doingmathwithpython/pycon-us-2016/blob/master/notebooks/Maps%20using%20Basemap%20-%20demo.ipynb)
# + [markdown] slideshow={"slide_type": "notes"}
# Next, I would like to talk about my book "Doing Math with Python". My idea was attractive enough to get it published by No Starch Press which makes me hope that I am probably onto something.
#
# Has anybody read my book? What do you think of it? You have read it and came to my talk? I am feeling better :)
#
# I discuss all of the topics I discuss today in my talk. In addition, I discuss sets, probability and random numbers and descriptive statistics.
#
# It's being translated into several non-English languages.
#
# The reviews/feedback so far has been really positive. I don't have any first hand involvement in teaching, so it's very appreciative of people to share their viewpoints with me.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Book: Doing Math With Python
#
# <img align="center" src="images/dmwp-cover.png" href="https://doingmathwithpython.github.io"></img>
#
# Overview
#
# - All of what I have discussed so far
#
# - In addition: Descriptive statistics, Sets and Probability, Random numbers
#
# Published by [No Starch Press](https://www.nostarch.com/doingmathwithpython) in August, 2015.
#
# *Upcoming/In-progress translations*: Simplified Chinese, Japanese, French and Korean.
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Comments
#
# > Saha does an excellent job providing a clear link between Python and upper-level math concepts, and demonstrates how Python can be transformed into a mathematical stage. This book deserves a spot on every geometry teacher’s bookshelf.
#
# [School Library Journal](http://www.slj.com/2016/05/collection-development/read-watch-alikes/coding-lets-begin/#_)
# + [markdown] slideshow={"slide_type": "subslide"}
#
# > Outstanding guide to using Python to do maths. Working back through my undergrad maths using Python.
#
# + [markdown] slideshow={"slide_type": "subslide"}
#
# > Saha does an excellent job providing a clear link between Python and upper-level math concepts, and demonstrates how Python can be transformed into a mathematical stage.
# + [markdown] slideshow={"slide_type": "fragment"}
#
# > This book is highly recommended for the high school or college student and anyone who is looking for a more natural way of programming math and scientific functions
# + [markdown] slideshow={"slide_type": "fragment"}
# > As a teacher I highly recommend this book as a way to work with someone in learning both math and programming
#
# + [markdown] slideshow={"slide_type": "notes"}
# Okay, so that's great. We have successfully used Python to make the learning experience of young learners more fun and immediately applicable. Can we derive more benefit from doing that? Like something for the future? We all love doing things for the future, don't we?
#
# I think yes, i think if we teach young learners the things we have discussed today, it is a great base for someone wanting to go into data science or machine learning.
#
# Statistics and visualising data are two very key factors of data science.
#
# Differential calculus and specifically the gradient descent method is a simple but useful optimization method used in Machine Learning. Let's see a demo of using gradient descent to find the minimum value of a function.
#
# Now, let's apply gradient descent as an optimizer in a Linear Regression problem.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Great base for the future
#
# *Statistics and Graphing data* -> *Data Science*
#
# *Differential Calculus* -> *Machine learning*
#
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Application of differentiation
#
# Use gradient descent to find a function's minimum value [(Notebook)](https://github.com/doingmathwithpython/pycon-us-2016/blob/master/notebooks/Gradient%20Descent.ipynb)
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Predict the college admission score based on high school math score
#
# Use gradient descent as the optimizer for single variable linear regression model [(Notebook)](https://github.com/doingmathwithpython/pycon-us-2016/blob/master/notebooks/Simple%20Linear%20Regression.ipynb)
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Advanced libraries
#
# - [scipy](https://scipy.org)
#
# - [numpy](http://www.numpy.org/)
#
# - [scikit-learn](http://scikit-learn.org/stable/)
#
# - [pandas](http://pandas.pydata.org/)
#
# - [Statsmodels](http://statsmodels.sourceforge.net/)
#
#
# + [markdown] slideshow={"slide_type": "slide"}
#
# ### Dialogue
#
# Questions, Thoughts, comments, discussions?
#
#
# #### Online
#
# - Twitter: @echorand
#
# - Email: <EMAIL>
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# ### PyCon Special!
#
# *Use PYCONMATH code to get 30% off "Doing Math with Python" from [No Starch Press](https://www.nostarch.com/doingmathwithpython)*
#
#
# <img align="center" src="images/dmwp-cover.png" href="https://doingmathwithpython.github.io"></img>
#
#
# (Valid from May 26th - June 8th)
#
# Book Signing - May 31st - 2.00 PM - No Starch Press booth
# + [markdown] slideshow={"slide_type": "slide"}
# ### Acknowledgements
#
# PyCon US Education Summit team for inviting me
#
#
# + [markdown] slideshow={"slide_type": "fragment"}
# Thanks to PyCon US for reduced registration rates
#
# + [markdown] slideshow={"slide_type": "fragment"}
#
# Massive thanks to my employer, Freelancer.com for sponsoring my travel and stay
#
# + [markdown] slideshow={"slide_type": "slide"}
# ### Links
#
# - [Upcoming O'Reilly Webcast](http://www.oreilly.com/pub/e/3712)
#
# - [Doing Math with Python](https://nostarch.com/doingmathwithpython)
#
# - [Doing Math with Python Blog](https://doingmathwithpython.github.io)
#
# - [Doing Math with Python on GitHub](https://github.com/doingmathwithpython)
#
#
| slides.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib as mpl
import math
from numpy import cos,sin
from matplotlib import cm
controlpoints_data = pd.read_table("10160528_controlpoints.txt", sep= "\s+", names = ["ID", "Longitude", "Latitude", "H", "h"])
controlpoints_data.head() #code to see the values as a proper table
controlpoints_data["N"] = controlpoints_data["h"]- controlpoints_data["H"] #ondulation value for control points is calculated and added to the table
testpoints_data = pd.read_table("10160528_testpoints.txt", sep= "\s+", names = ["ID", "Longitude", "Latitude", "H", "h"])
testpoints_data.head()
testpoints_data["N"] = testpoints_data["h"]- testpoints_data["H"] #ondulation value for test points is calculated and added to the table
# +
A_matrix = []
for i in range (len(controlpoints_data)):
A_matrix.append( [1,
np.cos(np.radians(controlpoints_data["Latitude"][i]))*np.cos(np.radians(controlpoints_data["Longitude"][i])),
np.cos(np.radians(controlpoints_data["Latitude"][i]))*np.sin(np.radians(controlpoints_data["Longitude"][i])),
np.sin(np.radians(controlpoints_data["Latitude"][i]))])
A = np.matrix(A_matrix) # design matrix
#print(len(A))
np.savetxt('010160528_designmatrix.txt', A, delimiter=' ',header="Design matrix of the Control Points",fmt='%10.10f') #Saving as txt
L = np.transpose(np.matrix([controlpoints_data["N"].values.tolist()])) #observations matrix (N values)
#print(L)
# weight matrix P is given as 1 so not included to calculations
X_control = np.dot(np.linalg.inv(np.dot(np.transpose(A),A)),np.dot(np.transpose(A),L)) #((At*A)**(-1))*(At*L) matrix of unknowns
#print(X_control)
N_adjusted = np.dot(A,X_control) # N" = A.x
controlpoints_data["adjustedN"] = np.dot(A,X_control) #adjustment N value of control points
v = np.subtract(np.dot(A, X_control), L) # adjustedN V = Ax-l
controlpoints_data["V"] = np.subtract(N_adjusted,L)
#Errors
vv = []
k = 0
for i in range(len(v)):
vvi = controlpoints_data["V"][i]*controlpoints_data["V"][i]
vv.append([vvi])
k = k + vvi
i = i+1
controlpoints_data["VV"] = np.matrix(vv)
#print(len(vv))
#print(k) # Sum of VV values
u = 4 #number of unknown values x0,x1,x2,x3
rmse_control = (k/ (len(controlpoints_data)-u) )** (1/2)
print(rmse_control,'m rmse error')
#vv = np.dot(v,np.transpose(v))
#print(np.matrix([controlpoints_data["V"].values.tolist()]))
# +
#Error estimates of coefficients (errors of x0,x1,x2,x3)
# from Error Propogation law
N_matrix = np.dot(np.transpose(A),A)
N_matrixlist = N_matrix.tolist()
#print(N_matrix) # 4x4
Nr_matrix = []
for i in range(len(N_matrixlist)):
Nr_matrix.append(N_matrixlist[i][i])
print(Nr_matrix) # diagonal values(P0,P1,P2,P3) of the matrix since error of coefficient x0 = m0 x (Po)**(1/2) --- m0 is rmse error 0.08590632505244178
X_controllist = X_control.tolist()
#print(X_controllist)
X_controladj = []
for i in range(len(X_controllist)):
X_controladj.append(rmse_control*(np.sqrt(Nr_matrix[i]))) #x0 = m0 x (Po)**(1/2)
print(X_controladj) # deviation of x0,x1,x2,x3
# +
#A,L,v Matrices of test points
A_matrix_test = []
for i in range (len(testpoints_data)):
A_matrix_test.append( [1,
np.cos(np.radians(testpoints_data["Latitude"][i]))*np.cos(np.radians(testpoints_data["Longitude"][i])),
np.cos(np.radians(testpoints_data["Latitude"][i]))*np.sin(np.radians(testpoints_data["Longitude"][i])),
np.sin(np.radians(testpoints_data["Latitude"][i]))])
A_test = np.matrix(A_matrix_test)
#print(A_test)
testpoints_data["N_from_geoid_model"] = np.dot(A_test,X_control) #because of A matrix of test includes phi and lam values of test points A_test is multiplied with unknown values matrix of model
L_test = np.transpose(np.matrix(testpoints_data["N"].values.tolist()))
#print(L_test)
v_test = np.subtract(np.dot(A_test,X_control),L_test)
testpoints_data["v"] = v_test
#Errors
VV = []
c = 0
for i in range(len(v_test)):
vvi2 = testpoints_data["v"][i]*testpoints_data["v"][i]
VV.append([vvi2])
c = c + vvi2
i = i+1
testpoints_data["vv"] = np.matrix(VV)
u = 4 #number of unknown values x0,x1,x2,x3
rmse_test = (c/ (len(testpoints_data)-u) )** (1/2)
print(rmse_test,'m rmse error')
#Saving as .txt
np.savetxt(r'controlpoints_data_with_v.txt', controlpoints_data.values, delimiter = " ", header ="ID--------------Longitude--------Latitude-------Orthometric Height---Elipsoidal Height--------N----------adjustedN-----------v",fmt='%10.10f')
np.savetxt(r'testpoints_data_with_v.txt', testpoints_data.values, delimiter = " ", header ="ID--------------Longitude--------Latitude--------Orthometric Height---Elipsoidal Height--------N--------NfromGeoidModel---------v",fmt='%10.10f')
# +
#Scatterplot of distribution
fig = plt.figure()
plt.scatter(testpoints_data["Longitude"],testpoints_data["Latitude"],edgecolors='none',c=testpoints_data["N"], marker= "^", cmap = cm.jet)
plt.scatter(controlpoints_data["Longitude"],controlpoints_data["Latitude"],edgecolors='none',c=controlpoints_data["adjustedN"], marker= "o", cmap = cm.jet)
plt.colorbar()
plt.title('Test points represented by triangle, control points represented by circle')
# -
| 10160528.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## NLP Preprocessing
# + hide_input=true
from fastai.gen_doc.nbdoc import *
from fastai.text import *
# -
# `text.tranform` contains the functions that deal behind the scenes with the two main tasks when preparing texts for modelling: *tokenization* and *numericalization*.
#
# *Tokenization* splits the raw texts into tokens (which can be words, or punctuation signs...). The most basic way to do this would be to separate according to spaces, but it's possible to be more subtle; for instance, the contractions like "isn't" or "don't" should be split in \["is","n't"\] or \["do","n't"\]. By default fastai will use the powerful [spacy tokenizer](https://spacy.io/api/tokenizer).
#
# *Numericalization* is easier as it just consists in attributing a unique id to each token and mapping each of those tokens to their respective ids.
# ## Tokenization
# ### Introduction
# This step is actually divided in two phases: first, we apply a certain list of `rules` to the raw texts as preprocessing, then we use the tokenizer to split them in lists of tokens. Combining together those `rules`, the `tok_func`and the `lang` to process the texts is the role of the [`Tokenizer`](/text.transform.html#Tokenizer) class.
# + hide_input=true
show_doc(Tokenizer)
# -
# This class will process texts by applying them the `pre_rules`, tokenizing them with `tok_func(lang)` and then applying the `post_rules`. `special_cases` are a list of tokens passed as special to the tokenizer and `n_cpus` is the number of cpus to use for multi-processing (by default, half the cpus available). We don't directly pass a tokenizer for multi-processing purposes: each process needs to initiate a tokenizer of its own. The rules and special_cases default to:
#
# `defaults.text_pre_rules = [fix_html, replace_rep, replace_wrep, spec_add_spaces, rm_useless_spaces]` <div style="text-align: right"><a href="https://github.com/fastai/fastai/blob/master/fastai/text/transform.py#L80">[source]</a></div>
#
# `defaults.text_post_rules = [replace_all_caps, deal_caps]` <div style="text-align: right"><a href="https://github.com/fastai/fastai/blob/master/fastai/text/transform.py#L81">[source]</a></div>
#
# and
#
# `defaults.text_spec_tok = [UNK,PAD,BOS,FLD,TK_MAJ,TK_UP,TK_REP,TK_WREP]` <div style="text-align: right"><a href="https://github.com/fastai/fastai/blob/master/fastai/text/transform.py#L10">[source]</a></div>
#
# The rules are all listed below, here is the meaning of the special tokens:
# - `UNK` (xxunk) is for an unknown word (one that isn't present in the current vocabulary)
# - `PAD` (xxpad) is the token used for padding, if we need to regroup several texts of different lengths in a batch
# - `BOS` (xxbos) represents the beginning of a text in your dataset
# - `FLD` (xxfld) is used if you set `mark_fields=True` in your [`TokenizeProcessor`](/text.data.html#TokenizeProcessor) to separate the different fields of texts (if your texts are loaded from several columns in a dataframe)
# - `TK_MAJ` (xxmaj) is used to indicate the next word begins with a capital in the original text
# - `TK_UP` (xxup) is used to indicate the next word is written in all caps in the original text
# - `TK_REP` (xxrep) is used to indicate the next character is repeated n times in the original text (usage xxrep n {char})
# - `TK_WREP`(xxwrep) is used to indicate the next word is repeated n times in the original text (usage xxwrep n {word})
# + hide_input=true
show_doc(Tokenizer.process_text)
# + hide_input=true
show_doc(Tokenizer.process_all)
# -
# For an example, we're going to grab some IMDB reviews.
path = untar_data(URLs.IMDB_SAMPLE)
path
df = pd.read_csv(path/'texts.csv', header=None)
example_text = df.iloc[2][1]; example_text
tokenizer = Tokenizer()
tok = SpacyTokenizer('en')
' '.join(tokenizer.process_text(example_text, tok))
# As explained before, the tokenizer split the text according to words/punctuations signs but in a smart manner. The rules (see below) also have modified the text a little bit. We can tokenize a list of texts directly at the same time:
df = pd.read_csv(path/'texts.csv', header=None)
texts = df[1].values
tokenizer = Tokenizer()
tokens = tokenizer.process_all(texts)
' '.join(tokens[2])
# ### Customize the tokenizer
# The `tok_func` must return an instance of [`BaseTokenizer`](/text.transform.html#BaseTokenizer):
# + hide_input=true
show_doc(BaseTokenizer)
# + hide_input=true
show_doc(BaseTokenizer.tokenizer)
# -
# Take a text `t` and returns the list of its tokens.
# + hide_input=true
show_doc(BaseTokenizer.add_special_cases)
# -
# Record a list of special tokens `toks`.
# The fastai library uses [spacy](https://spacy.io/) tokenizers as its default. The following class wraps it as [`BaseTokenizer`](/text.transform.html#BaseTokenizer).
# + hide_input=true
show_doc(SpacyTokenizer)
# -
# If you want to use your custom tokenizer, just subclass the [`BaseTokenizer`](/text.transform.html#BaseTokenizer) and override its `tokenizer` and `add_spec_cases` functions.
# ### Rules
# Rules are just functions that take a string and return the modified string. This allows you to customize the list of `default_pre_rules` or `default_post_rules` as you please. Those are:
# + hide_input=true
show_doc(deal_caps, doc_string=False)
# -
# In `t`, every word is lower-case. If a word begins with a capital, we put a token `TK_MAJ` in front of it.
# + hide_input=true
show_doc(fix_html, doc_string=False)
# -
# This rules replaces a bunch of HTML characters or norms in plain text ones. For instance `<br />` are replaced by `\n`, ` ` by spaces etc...
fix_html("Some HTML text<br />")
# + hide_input=true
show_doc(replace_all_caps)
# + hide_input=true
show_doc(replace_rep, doc_string=False)
# -
# Whenever a character is repeated more than three times in `t`, we replace the whole thing by 'TK_REP n char' where n is the number of occurrences and char the character.
replace_rep("I'm so excited!!!!!!!!")
# + hide_input=true
show_doc(replace_wrep, doc_string=False)
# -
# Whenever a word is repeated more than four times in `t`, we replace the whole thing by 'TK_WREP n w' where n is the number of occurrences and w the word repeated.
replace_wrep("I've never ever ever ever ever ever ever ever done this.")
# + hide_input=true
show_doc(rm_useless_spaces)
# -
rm_useless_spaces("Inconsistent use of spaces.")
# + hide_input=true
show_doc(spec_add_spaces)
# + hide_input=false
spec_add_spaces('I #like to #put #hashtags #everywhere!')
# -
# ## Numericalization
# To convert our set of tokens to unique ids (and be able to have them go through embeddings), we use the following class:
# + hide_input=true
show_doc(Vocab)
# -
# `itos` contains the id to token correspondence.
# + hide_input=true
show_doc(Vocab.create)
# -
# Only keeps `max_vocab` tokens, and only if they appear at least `min_freq` times, set the rest to `UNK`.
# + hide_input=true
show_doc(Vocab.numericalize)
# + hide_input=true
show_doc(Vocab.textify)
# -
vocab = Vocab.create(tokens, max_vocab=1000, min_freq=2)
vocab.numericalize(tokens[2])[:10]
# ## Undocumented Methods - Methods moved below this line will intentionally be hidden
# + hide_input=true
show_doc(SpacyTokenizer.tokenizer)
# + hide_input=true
show_doc(SpacyTokenizer.add_special_cases)
# -
# ## New Methods - Please document or move to the undocumented section
| docs_src/text.transform.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/JayellWolfe/DS-Unit-1-Sprint-1-Dealing-With-Data/blob/master/Coding_Challenge_7_10.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="DTryy-ftM4Nn" colab_type="text"
# ## Good Morning! For today's coding challenge, we are going to solve some python basics.
# + [markdown] id="MYcqiCCYNvrU" colab_type="text"
# ### 1. Write a Python program to create all possible strings by using 'a', 'e', 'i', 'o', 'u'. Use the characters exactly once.
#
# + id="P5oD9XXnMJ6S" colab_type="code" colab={}
# TODO code here
# + [markdown] id="U5VoSsxeN23s" colab_type="text"
# ###2. Write a Python program to calculate number of days between two dates
# + id="OskMQGCUOTg9" colab_type="code" colab={}
# TODO code here
# + [markdown] id="H039HvT_OWQD" colab_type="text"
# ### 3. Write a Python program to get the difference between a given number and 17, if the number is greater than 17 return double the absolute difference.
#
#
# + id="z7w5-aDmO6A9" colab_type="code" colab={}
# TODO code here
# + [markdown] id="Mx4wWoGlPtKD" colab_type="text"
# ### Stretch Goals
# + [markdown] id="Fpfn3ozePw96" colab_type="text"
# ### 4. Write a Python program to find whether a given number (accept from the user) is even or odd, print out an appropriate message to the user.
# + id="tUSmmweKPwaL" colab_type="code" colab={}
# TODO code here
# + [markdown] id="oHF_eYwkQSgH" colab_type="text"
# ### 5. Write a Python program to display your details like name, age, address in three different lines.
# + id="aXmgBFO1QzCO" colab_type="code" colab={}
# TODO code here
| Coding_Challenge_7_10.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# 노트북의 셀은 코드(Y), 막다운(M) 두가지를 사용합니다. 셀 실행은 쉬프트+엔터키 입니다.
3 + 5
a = 10
# a에 2를 빼서 다시 a에 입력합니다.
a -= 2
print(a)
# 노트북에서는 변수만 입력하면 값이 프린트 됩니다.
a
# a에 2를 곱해 다시 a에 입력합니다.
a *= 2
a
# a를 2로 나누어 다시 a에 입력합니다.
a /= 2
a
# a에 2를 제곱해 다시 a에 입력합니다.
a **= 2
a
# a를 3으로 나눈 몫을 구해 다시 a에 입력합니다.
a //= 3
a
# a를 4로 나눈 나머지를 구해 다시 a에 입력합니다.
a %= 4
a
# - 2진수는 숫자 앞에 '0b', '0B' 를 붙입니다.
# - 8진수는 숫자 앞에 '0o', '0O' 를 붙입니다.
# - 16진수는 숫자 앞에 '0x', '0X' 를 붙입니다.
0b10
0o10
0x10
# int와 float 간의 형변환은 int() 함수와 float() 함수를 사용합니다.
int(3.5)
float(3)
# 변수에 담을 수 있는 숫자값의 크기에 제한이 없습니다.
googol = 10 ** 100
googol
googol * googol
| tutorial-1/number.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Drone Notebook
#
# This jupyter notebook is part of my 2020 summmer internship through Pacific University in Dr. Dawes' lab.
#
# The objective of this notebook is to write and implement code to maneuver a Tello Edu drone, with the final goal of aligning two drones for optics experiments.
# +
import time
import socket
from tello import Tello
# -
# #### To Start
# 1. make sure tello.py in in same folder as this notebook
# 2. connect to Tello Drone wifi network
# Some example code:
#
# code modified from example on https://github.com/alecGraves/tello-udp
# +
# Declare a tello object
#this is a lot like turtles in 3D
drone = Tello()
# enable network commands on the drone
drone.send("command")
# takeoff
drone.send("takeoff")
start = time.time()
#initializes a start time
# wait 30 seconds
while time.time() - start < 30:
# get battery info to prevent tello connection from timing-out
# since tello auto-lands after 15 secs w/o a command
drone.send("battery?")
time.sleep(1)
# land the tello
drone.send("land")
# +
def real_time(drone,log=False):
"""This code is designed to put the user's tello drone into flight, then prompt the user
for commands in real time, as opposed to writing a long stream of commands in one code
cell
parameters:
drone: tello drone object
optional boolean log: if true will return a log of commands used"""
drone.send("command")
#if the drone hasn't already been initialized to get commands this will do that
print("Type tello command and hit enter. Type and send 'quit' to exit this program")
running = True
flight_log = []
while running:
#starts an infinite loop
task = input('')
#sends the message to the drone
drone.send(task)
if task == 'land':
running = False
if log == True:
flight_log.append(task)
if log == True:
return flight_log
# +
real_time(drone)
## notes: this works!
# -
def polygon(drone,sides):
"""code to make drone take off fly in the polygon shape with specified sides
drone: drone object
sides: int number of sides of shape"""
angle = 360/sides
angle = int(angle)# has to be an int!
drone.send('command')
drone.send('takeoff')
#creates a start time
start = time.time()
if time.time()% 5 == 0:
# five second delay
for i in range(0, sides):
print(i)
drone.send('forward 20')
task = 'cw {0}'.format(angle)
print(drone.send('battery?'))
drone.send(task)
drone.send('land')
# +
polygon(drone,5)
## notes my drone will follow some of the commands but not all of them
# I might be able to fix this by adding a delay
# -
drone.send("battery?")
# ## checking for drift
#
# So in the code below I will be checking for the drone's drift from a hovering point, with little to no wind/breeze during an indoor test with the windows closed
# +
def drift_test(drone, drift_time):
""" This function returns data of how long it takes the tello drone to return to
a starting location after drifting from that start"""
drone.send('mon')
drone.send('mdirection 0')
drone.send('takeoff')
#puts drone in air with mission pad detection on for pad below drone
drone.send('go 0 0 0 m1')
#places drone at center of mission pad 1
start = time.time()
#starts a timer
if time.time() - drift_time < 0:
drone.send("battery?")
time.sleep(1)
elif (time.time() - drift_time) >= 0 :
reset = time.time()
drone.send('go 0 0 0 m1')
reset_time = (time.time() - reset)
print( reset_time )
return reset_time
# -
drone.send('command')
drift_test(drone,10)
# ### List of Commands:
# #### (not all of them)
#
# Setting commands:
#
# * speed x
#
# sets the speed between 10 and 100 ( the manual didn't specify units, but I'm assuming cm/s?)
#
# ex: "speed 10"
#
# Mission pad commands:
#
# * mon
#
# turns on mission pad detection
#
# * moff
#
# turns off mission pad detection
#
# * mdirection x
#
# sets mission pad detection direction (must turn on detection first)
#
# 0 = downward / 1 = forward / 2 = both
#
# Motion Commands:
#
# * takeoff (auto takeoff)
# * land (auto landing)
# * emergency ( stops all motors)
# * up x (move up to 'x' cm above surface)
# * down x (move down to 'x' cm above surface)
# * forward/backward/left/right x (move 'x' cm in respective direction)
# * cw/ccw x (rotate 'x' degrees in respetive direction)
#
| Tello_Drone_Motion.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Project: Investigate TMDb movie dataset
#
# ## Table of Contents
# <ul>
# <li><a href="#intro">Introduction</a></li>
# <li><a href="#wrangling">Data Wrangling</a></li>
# <li><a href="#eda">Exploratory Data Analysis</a></li>
# <li><a href="#conclusions">Conclusions</a></li>
# </ul>
# <a id='intro'></a>
# ## Introduction
#
# ### Dataset Description
#
# This data set contains information about 10,000 movies collected from The Movie Database (TMDb) including user ratings and revenue. It has 21 columns and 10866 rows.
#
# ### Question(s) for Analysis
#
# 1. How the popularity of the movies are affected through time?
# 2. What features are associated with movies that have high revenues?
# 3. What are the most 5 popular movies in 2015?
# 4. What are the highest 10 movies in 2015 according to the revenue?
# 5. What are the most frequent words used in the overview and keywords for the most popular movies?
#
# +
# Upgrade pandas to use dataframe.explode() function.
# #!pip install --upgrade pandas==0.25.0
# Install regex in case if it won't be imported successfully.
# !pip install regex
# -
# Importing libraries
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
import regex as re
from wordcloud import WordCloud
# %matplotlib inline
# <a id='wrangling'></a>
# ## Data Wrangling
#
# In this section, we will load the data, check for cleanliness, get some high level overview about the data and then trim and clean the dataset for analysis.
# Load data and print head
df = pd.read_csv('Database_TMDb_movie_data/tmdb-movies.csv')
df.head(5)
# Get descriptive statistics.
df.describe()
# ##### Note:
# 1. The oldest movie in this dataset is since 1960, however, the newest ones are released in 2015
# 2. Min run time is zero which seems to be incorrect indicator.
# 3. 75% of the revenue column in the dataset has less than 24M. However the max value is approx. 2B which is really high!
# Check the size of the data
print('Number of rows in the dataset is: {}'.format(df.shape[0]))
print('Number of columns in the dataset is: {}'.format(df.shape[1]))
# Check the types of the columns
df.dtypes
# ##### Note:
# release_date column is defined as string in the dataset and not datetime (eligible for modification). On the otherhand, release_year is int which is a valid/healthy option.
# Check duplicate rows
print("Number of duplicated rows are:", df.duplicated().sum())
# This duplicated row can be safely removed in the data cleaning phase.
# Check for missing values
df.isnull().sum()
# Check if some movies has revenue of zeros
print('{} rows has zero revenue values.'.format(df['revenue'].value_counts()[0]))
print('{} rows has zero runtime values.'.format(df['runtime'].value_counts()[0]))
df[['budget', 'revenue', 'release_year']].hist(figsize=(8, 8));
# #### Conclusion of the Data Wrangling:
# 1. There are several columns which we don't need for our analysis purposes.(i.e. imdb_id, homepage)
# 2. There are around 6016 rows of zero revenue. (More than 50% of our total data sample!)
# 3. The column with the most missing values are homepage URLs which is not necessary for our analysis purposes.
# 4. We have only 1 duplicate row which can be removed.
# 5. Keep in mind that the runtime is zero in 31 rows of the sample.
# 6. Keep in mind the large distribution in the revenue column.
# 7. The release year is right skewed which means that more movies are released with time.
#
# ### Data Cleaning
# In this section we will prepare the data to be able to do the necessary analysis.
# 1- Remove the duplicates
# +
# Remove the duplicate row
df.drop_duplicates(inplace=True)
# Check the shape now -- Should have 10865 rows
df.shape
# -
# 2- Drop rows with zero revenue
# +
# Drop rows with 0 revenue
df = df[df['revenue'] != 0]
# Check the shape now -- Should have 10865-6016 = 4849 rows
df.shape
# +
# Remove the unnecessary columns for our analysis.
remove_columns = ['imdb_id', 'homepage','director', 'tagline', 'release_date']
df.drop(remove_columns, axis=1, inplace=True)
# Check the shape now -- Should have 16 columns now
df.shape
# -
# ##### Note:
# We have removed duplicates, rows with zeros and unnecessary columns.
# In the next step, we would like to see more low level details about the revenue column and transform it to reasonable interval (Smaller distribution of values).
# +
# Detect outlier of revenue column
plt.hist(df['revenue'])
plt.xlabel('Revenue')
plt.title('Distribtuion of Revenue across the sample')
plt.grid(True)
plt.show()
# -
# Define min and max thresholds to filter the data
min_threshold, max_threshold = df.revenue.quantile([0.05, 0.9])
min_threshold, max_threshold
# ##### Note
# 1. Max is approx. 232M which is is a bit reasonable now.
# 2. We have done this step as to limit a bit the big distribution in the original dataset.
# What data points has that lower values than the threshold
df[df.revenue < min_threshold].head(5)
# What data points has that higher values than the threshold
df[df.revenue > max_threshold].head(5)
# ##### Note:
# 1. This is not necessarly data errors or not valid, but if we keep these data points they will surely affect our analysis in certain bias direction.
df = df[(df['revenue'] < max_threshold) & (df['revenue'] > min_threshold)]
df.isnull().sum()
# ##### Note:
# We removed the row that has missing values to complete the cleaning of the data.
# Drop rows that has missing values
df.dropna(axis=0, inplace=True)
# <a id='eda'></a>
# ## Exploratory Data Analysis
# Now that we've trimmed and cleaned your data, We're ready to move on to exploration. **Compute statistics** and **create visualizations** with the goal of addressing the research questions that was posed in the Introduction section.
#
#
# ### 1. How the popularity of the movies are affected through time?
# Group data by year and get the median in terms of Popularity
df_pop = df.groupby('release_year').median()['popularity']
df_pop_index = df_pop.index
x, y = df_pop_index, df_pop
# Plot configs.
plt.plot(x, y)
plt.title('Popularity distribtuion per year')
plt.xlabel('Year')
plt.ylabel('Popularity index')
plt.show()
# ### 2. What features are associated with movies that have high revenues?
#
# In this section we will investigate what are the main factors affecting the revenue through all the dataset.
# +
#Visualise the coorelation matrix between the revenue and other features
cols = ['popularity', 'budget', 'runtime', 'vote_average', 'vote_count', 'release_year', 'revenue']
heatmap = sns.heatmap(df[cols].corr(), annot=True, fmt='.2f')
#Plot and save
fig = heatmap.get_figure()
# -
# Get DataFrame for only 2015
df_2015 = df[df['release_year'] == 2015]
# Validate the coorelation across different features for the 2015 dataset
sns.regplot(x=df_2015['revenue'], y=df_2015['budget'])
# Check the coorelation with respect to popularity
sns.regplot(x=df_2015['revenue'], y=df_2015['popularity'])
# Check the coorelation with respect to popularity
sns.regplot(x=df_2015['revenue'], y=df_2015['vote_average'])
# ##### Conclusion:
# 1. From the above diagrams, it is proven that the revenue is accompined with high popularity and high budget.
#
# ### 3. What are the most 5 popular movies in 2015?
# Get the highest 5 movies according to popularity.
# +
# Use the same df_2015 from the above section
# Sort the values by popularity
df_2015_popularity = df_2015[['popularity','original_title']].sort_values(by='popularity', ascending=False)
# -
# Plot the first 5 in the DF
df_2015_popularity[0:5].plot(kind='bar', x='original_title', y='popularity',
rot=90, title='The 5 most popular movies in 2015', legend=False)
# ### 4. What are the highest 10 movies in 2015 according to the revenue?
# Get the highest 10 movies in 2015 according to the revenue
df_2015_revenue = df_2015[['revenue','original_title', 'budget']].sort_values(by='revenue', ascending=False)
df_2015_revenue[0:10].plot(kind='bar', x='original_title', y='revenue', rot=90,
title='The movies with the highest revenue in 2015', legend=False)
# ##### Conclusion:
# From the above diagrams we can conclude:
# 1. Jupiter Ascending, Ex machina, The Hateful Eight, Tomorrowland and Southpaw are the most popular movies in 2015.
# 2. Ted 2 comes in the first placewith above 200M of revenue.
# ### 5. What are the most frequent words used in the overview and keywords for the most popular movies?
# Define the wordcloud DataFrame
df_word_cloud = df_2015.sort_values('popularity', ascending=False)[0:10]
# +
# Create helper functions for cleaning the text and generate wordclouds.
def clean_text(df, column):
'''
function to clean raw text from the column in the DF.
inputs:
df: Pandas DataFrame which has the column to be cleaned.
column: str - column name in the df
return:
text_cleaned: string which has the text that can be converted to wordcloud.
'''
text_raw = df[column].values
text_raw = np.array_str(text_raw)
text_cleaned = re.sub("[^A-Za-z0-9]"," ",text_raw)
return text_cleaned
def generate_wordcloud(text_cleaned):
'''
function to generate and plot the wordcloud from the text input.
inputs:
text_cleaned:
'''
wordcloud = WordCloud().generate(text_cleaned)
plt.figure(figsize = (15, 15))
plt.imshow(wordcloud, interpolation='bilinear')
plt.axis("off")
plt.show()
# +
# Use the clean_text function to clean the text in keywords column.
keywords_text = clean_text(df_word_cloud, 'keywords')
# Use the generate wordcloud function to plot the image.
generate_wordcloud(keywords_text)
# -
# ##### Note:
# 1. You can see that in the overview, the most used words in the keywords column are: Aritifcial, robot, bank, intelligence, woman, romance.
# +
# Use the clean_text function to clean the text in overview column
overview_text = clean_text(df_word_cloud, 'overview')
# Use the generate wordcloud function to plot the image
generate_wordcloud(overview_text)
# -
# ##### Note:
# 1. You can see that in the overview, the most used words are: World, Child, memory, old, wife, which all seems to me like a genre of Action, thriller and Scify.
# <a id='conclusions'></a>
# ## Conclusions
#
# > **Tip**: Finally, summarize your findings and the results that have been performed in relation to the question(s) provided at the beginning of the analysis. Summarize the results accurately, and point out where additional research can be done or where additional information could be useful.
#
# Results:
#
# 1. It looks that popularity index of the movies are increasing over years.
# 2. Find, Child, Robot, Artificial, Intelligence are most common words in the movies which gives a bit of idea how the most popular movies tends to be.
# 3. Ted 2 is the highest revenue movie in 2015.
# 4. Revenue seems to be positively coorelate with the popularity and budget features.
# 5. Avatar has the highest revenue in the dataset with value over 2B.
#
# Future Work:
#
# 1. More text analysis can be done over the cast and production_companies columns.
# 2. Runtime column can be used to see if it affects the popularity. (i.e. When the movie is long, is it more popular?)
#
#
# ### Limitations
#
# 1. Due to lack of knowledge in the movie industry from the business prespective, there wasn't much a constructive decision for what can be considered outliers in terms of revenue and budget.
from subprocess import call
call(['python', '-m', 'nbconvert', 'Investigate_a_Dataset.ipynb'])
| tmdb-movies.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import progressbar
import time
import codecs
import functools
import os
import tempfile
import zipfile
import urllib
import re
from nltk.corpus import stopwords
from gensim.models import word2vec
import pickle
import nltk.data
import os
from keras.preprocessing.sequence import pad_sequences
# Turn each comment into a list of word indexes of equal length (with truncation or padding as needed).
def comments2Matrix(comment_list,model,maxlen):
def commentToIndex(comment,index2word_set,model):
indexed_comment = []
# Loop over each word in the comment and, if it is in the model's vocaublary convert it to an index
for word in comment:
if word in index2word_set:
indexed_comment += [model.wv.vocab[word].index]
return [indexed_comment]
index2word_set = set(model.wv.index2word)
totalComments = len(comment_list)
bar = progressbar.ProgressBar(maxval=totalComments, widgets=[progressbar.Bar('=', '[', ']'), ' ', progressbar.Percentage()])
bar.start()
# Loop over each comment in the comment_list
i=0 #init for progress bar
indexed_comments = [] #init
for comment in comment_list:
indexed_comments += commentToIndex(comment,index2word_set,model)
i += 1
bar.update(i)
bar.finish()
#return indexed_comments
return pad_sequences(indexed_comments,maxlen = maxlen)
from tensorflow.python.client import device_lib
print(device_lib.list_local_devices())
# Read data from files
train = pd.read_csv("data/train.csv", header=0)
list_classes = ["toxic", "severe_toxic", "obscene", "threat", "insult", "identity_hate"]
y = train[list_classes].values
with open('data/tokenized_comments/remove_stops=False.lemmatize=False.spellcheck=True.train_comments.csv') as f:
train_comments = [line.split() for line in f]
with open('data/tokenized_comments/remove_stops=False.lemmatize=False.spellcheck=True.test_comments.csv') as f:
test_comments = [line.split() for line in f]
print("Loaded %s training comments, and %s testing comments" % (len(train_comments),len(test_comments)))
# ##### <b>Load Pretrained Glove Embeddings</b>
#Load w2v Model Using Gensim
from gensim.models import Word2Vec
import gensim
print("Loading Gensim Model...")
gensim_file= 'w2v_models/gensim_filtered_models/remove_stops=False.lemmatize=False.spellcheck=True.glove.42B.300d.txt'
word_vectors = gensim.models.KeyedVectors.load_word2vec_format(gensim_file)
print("Gensim Model Loaded")
# ##### <b>Convert Comments to a Matrix of Indices</b>
# Initialize parameters for model
embed_size = 300 #Embed Size Of Model
maxlen = 150 #Max number of words to use for a specific comment
max_features = len(word_vectors.wv.vocab) # how many unique words to use (i.e num rows in embedding vector)
print('Converting %s comments for training set to matrices' % len(train_comments))
xtrain = comments2Matrix(train_comments,word_vectors,maxlen)
print('Converting %s comments for testing set to matrices' % len(test_comments))
xtest = comments2Matrix(test_comments,word_vectors,maxlen)
# convert the wv word vectors into a numpy matrix that is suitable for insertion into Keras models
embedding_matrix = np.zeros((len(word_vectors.wv.vocab), embed_size))
for i in range(len(word_vectors.wv.vocab)):
embedding_vector = word_vectors.wv[word_vectors.wv.index2word[i]]
if embedding_vector is not None:
embedding_matrix[i] = embedding_vector
# +
from keras.layers import Dense, Input, LSTM, Embedding, Dropout, Activation, MaxPooling2D, Reshape,MaxPooling1D
from keras.layers import Bidirectional, GlobalMaxPool1D, Conv2D, SpatialDropout1D, BatchNormalization, GlobalMaxPooling2D,Conv1D
from keras.initializers import glorot_normal
from keras.models import Model
from keras.callbacks import EarlyStopping, ModelCheckpoint
from keras import initializers, regularizers, constraints, optimizers, layers, Sequential
file_path = "BD-LSTM-noatt-maxlen100.hdf5"
checkpoint = ModelCheckpoint(file_path, monitor='val_loss', verbose=1,save_best_only=True, mode='min')
early_stop = EarlyStopping(monitor="val_loss", mode="min", patience=3)
model = Sequential()
model.add(Embedding(max_features, embed_size, weights=[embedding_matrix],trainable = False,name = 'Word-Embedding-Layer'))
model.add(Dropout(0.4,name = 'Dropout-Regularization-1')) # Best = 0.3
model.add(Bidirectional(LSTM(300, return_sequences=True, dropout=0.25, recurrent_dropout=0.25,kernel_initializer=glorot_normal(seed=None)),name = 'BDLSTM')) #Best = 300,0.25,0.25
model.add(GlobalMaxPool1D(name = 'Global-Max-Pool-1d'))
model.add(Dense(256, activation="relu",name = 'FC-256')) # Best = 256
model.add(Dense(6, activation="sigmoid",name = 'FC-Output-Layer'))
model.compile(loss='binary_crossentropy', optimizer='nadam', metrics=['accuracy'])
model.summary()
history = model.fit(xtrain, y, batch_size=50, epochs=100,validation_split=0.1, callbacks=[checkpoint,early_stop],verbose=1)
# +
from keras.layers import Dense, Input, LSTM, Embedding, Dropout, Activation, MaxPooling2D, Reshape,MaxPooling1D
from keras.layers import Bidirectional, GlobalMaxPool1D, Conv2D, SpatialDropout1D, BatchNormalization, GlobalMaxPooling2D,Conv1D
from keras.initializers import glorot_normal
from keras.models import Model
from keras.callbacks import EarlyStopping, ModelCheckpoint
from keras import initializers, regularizers, constraints, optimizers, layers, Sequential
file_path = "BD-LSTM-noatt.hdf5"
checkpoint = ModelCheckpoint(file_path, monitor='val_loss', verbose=1,save_best_only=True, mode='min')
early_stop = EarlyStopping(monitor="val_loss", mode="min", patience=3)
model = Sequential()
model.add(Embedding(max_features, embed_size, weights=[embedding_matrix],trainable = False,name = 'Word-Embedding-Layer'))
model.add(Dropout(0.3,name = 'Dropout-Regularization-1'))
model.add(Bidirectional(LSTM(300, return_sequences=True, dropout=0.25, recurrent_dropout=0.25,kernel_initializer=glorot_normal(seed=None)),name = 'BDLSTM'))
model.add(GlobalMaxPool1D(name = 'Global-Max-Pool-1d'))
model.add(Dense(256, activation="relu",name = 'FC-256'))
model.add(Dense(6, activation="sigmoid",name = 'FC-Output-Layer'))
model.compile(loss='binary_crossentropy', optimizer='nadam', metrics=['accuracy'])
model.summary()
#history = model.fit(xtrain, y, batch_size=256, epochs=100,validation_split=0.1, callbacks=[checkpoint,early_stop],verbose=1)
# -
# ### <B> MOVE TO ANOTHER NOTEBOOK TO AVOID CONFUSION AND LOST DATA </B>
# +
# Notes
# Good Success with lowering the length - probably erases a lot of padding that confuses the nn
# -
file_path = "BD-LSTM-noatt-maxlen100.hdf5"
model.load_weights(file_path)
y_test = model.predict([xtest], batch_size=1024, verbose=1)
sample_submission = pd.read_csv('submissions/sample_submission.csv')
sample_submission[list_classes] = y_test
sample_submission.to_csv('submissions/no_stops_test_scores.csv', index=False)
file_path = "BD-LSTM-noatt.hdf5"
model.load_weights(file_path)
y_test = model.predict([xtest], batch_size=1024, verbose=1)
sample_submission = pd.read_csv('submissions/sample_submission.csv')
sample_submission[list_classes] = y_test
sample_submission.to_csv('submissions/glove_vectors_unlemmatized_len50_LSTM.csv', index=False)
# ##### <b> Ensembling Models </b>
lstm_100 = 'submissions/glove_vectors_unlemmatized_len100_LSTM0043.csv'
lstm_50 = 'submissions/glove_vectors_unlemmatized_len50_LSTM.csv'
p_lstm100 = pd.read_csv(lstm_100)
p_lstm50 = pd.read_csv(lstm_50)
label_cols = ['toxic', 'severe_toxic', 'obscene', 'threat', 'insult', 'identity_hate']
p_res = p_lstm100.copy()
p_res[label_cols] = (p_lstm100[label_cols] + p_lstm50[label_cols]) / 2
p_res.to_csv('submissions/ensemble_100_50_submission.csv', index=False)
| keras_BDRNN-2DCNN-MaxPool-BestScore.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Hawaii Weather Analysis
#
# ### Dependencies
# Matplotlib
# %matplotlib inline
import matplotlib.pyplot as plt
import matplotlib as mpl
mpl.style.use('fivethirtyeight')
# Other dependencies
import datetime as dt
import numpy as np
import pandas as pd
# Python SQL toolkit and object relational mapper (ORM)
from sqlalchemy.ext.automap import automap_base
from sqlalchemy.orm import Session
from sqlalchemy import create_engine, func
# ### Reflect tables into ORM
# Create SQL engine
engine = create_engine('sqlite:///hawaii.sqlite')
engine
# +
# Reflect tables in the database into a base class
Base = automap_base()
Base.prepare(engine, reflect=True)
# Show all classes that automap found (each class is a table)
Base.classes.keys()
# +
# Save a reference to each table
M = Base.classes.measurement
S = Base.classes.station
# Create a session
session = Session(engine)
session
# -
# Inspect measurement table
measurement = pd.read_sql('SELECT * FROM measurement', engine)
print(measurement.info())
measurement.head(2)
# Inspect station table
station = pd.read_sql('SELECT * FROM station', engine)
station
# ### Precipitation in the last 12 months
# +
# Last date in the data
last_date = session.query(func.max(M.date)).first()[0] # 1st row and col of query results
end = dt.datetime.strptime(last_date, '%Y-%m-%d').date() # convert to date type
# Date one year from the last date
start = end - dt.timedelta(days=365)
start
# -
# Query dates and precipitation scores for the 12-month period
Mprcp = session.query(M.date, M.prcp).filter(M.date >= start).all()
print(len(Mprcp))
Mprcp[:3]
# +
# Extract columns from query results
Mprcp_arr = np.array(Mprcp)
date, prcp = Mprcp_arr[:, 0], Mprcp_arr[:, 1]
# Create a dataframe
Mprcp_df = pd.DataFrame(prcp.astype(float), index=pd.to_datetime(date), columns=['precipitation']).sort_index()
Mprcp_df.head(3)
# -
# Plot precipitation
Mprcp_df.plot(figsize=(12, 4), title='Daily precipitation', legend=False)
plt.show()
# Precipitation summary statistics
Mprcp_df.describe()
# ### Weather stations
# Number of stations in the data
session.query(S).count()
# Number of measurements from each station
Mcount_bystation = session.query(M.station, func.count(M.station)).group_by(M.station)
Mcount_bystation = Mcount_bystation.order_by(func.count(M.station).desc()).all() # order by descending count
Mcount_bystation
# Number of temperature observations from each station
Mtobs_bystation = session.query(M.station, func.count(M.tobs)).filter(M.date >= start)
Mtobs_bystation = Mtobs_bystation.group_by(M.station).order_by(func.count(M.tobs).desc()).all()
Mtobs_bystation
# Last 12 months of temperature observations for the most active station (station USC00519397)
s97_temps = session.query(M.date, M.tobs) \
.filter((M.station == Mtobs_bystation[0][0]) & (M.date >= start)).all()
s97_temps = np.array(s97_temps)
print(len(s97_temps))
s97_temps[:3]
# +
# Histogram of station 97 temperatures
plt.figure(figsize=(12, 4))
plt.hist(s97_temps[:, 1].astype(float), bins=12)
# Formatting
plt.title('Temperature Frequencies at Station ' + Mtobs_bystation[0][0])
plt.ylabel('Count')
plt.xlabel('Temperature (F)')
plt.tight_layout()
plt.plot()
# +
# Lineplot of station 97 temperatures
plt.figure(figsize=(12, 4))
plt.plot(pd.to_datetime(s97_temps[:, 0]), s97_temps[:, 1].astype(float))
# Formatting
plt.title('Daily Temperature at Station ' + Mtobs_bystation[0][0])
plt.ylabel('Temperature (F)')
plt.xlabel('Day')
plt.show()
# -
# ### Temperature by station
# Temperature statistics by station
station_temps = session.query(M.station, func.min(M.tobs), func.max(M.tobs), func.avg(M.tobs))
station_temps = station_temps.filter(M.date >= start).group_by(M.station).all()
station_temps
# +
def calc_temps(start_date=start, end_date=end):
"""
Calculate the minimum, average, and maximum temperatures for each station
for the specified date range.
Args:
[1] start_date (str) - in the format "%Y-%M-%D"
[2] end_date (str) - in the format "%Y-%M-%D"
Returns:
(Pandas dataframe) minimum, average, and maximum temperature for each
station in the date range.
"""
temps = session.query(M.station, func.min(M.tobs), func.max(M.tobs), func.avg(M.tobs))
temps = temps.filter((M.date >= start_date) & (M.date <= end_date)).group_by(M.station).all()
return pd.DataFrame(temps, columns=['station', 'min_temp', 'avg_temp', 'max_temp'])
# Test function
calc_temps('2016-01-01', '2016-12-31')
# -
# ### Monthly temperature - Pandas
# Extract month from date in measurement dataframe
measurement['month'] = measurement['date'].str.split('-', expand=True)[1].astype(int)
measurement.head(3)
# +
# Combinations of columns and statistics to calculate
columns = [(meas, stat) for meas in ['tobs', 'prcp'] for stat in ['min', 'max', 'mean', 'std']]
print(columns)
# Temperature and precipitation statistics by month
monthly_weather_pd = measurement.groupby('month').describe().round(2)[columns]
monthly_weather_pd.index.name = None
monthly_weather_pd
# -
# ### Monthly weather analysis - ORM
# +
# Subquery to extract month, temperature (tobs), and precipitation (prcp)
subquery = session.query(func.extract('month', M.date).label('month'), M.tobs, M.prcp).subquery()
# Create a list for the statistics to calculate: min_tobs, avg_tobs, max_tobs, min_prcp, avg_prcp, max_prcp
SELECT = [func.round(F(stat), 2) for stat in [subquery.c.tobs, subquery.c.prcp]
for F in [func.min, func.avg, func.max]]
SELECT.insert(0, subquery.c.month) # insert month as the 1st col
columns = ['month', 'min_temp', 'avg_temp', 'max_temp', 'min_prcp', 'avg_prcp', 'max_prcp'] # col names
# Temperature and precipitation statistics by month
query = session.query(*SELECT).group_by(SELECT[0]).all() # query
monthly_weather_orm = pd.DataFrame(query, columns=columns) # convert to df
monthly_weather_orm.set_index('month', inplace=True) # set month as index
monthly_weather_orm.index.name = None # remove index name
monthly_weather_orm
# -
| hi-weather/analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: all
# language: python
# name: all
# ---
# ! cd dana && ./install_local_setup.sh && cd -
# ! cd coldeport && ./install_local_setup.sh && cd -
# ! cd reynolds && ./install_local_setup.sh && cd -
# %pylab inline
import pandas as pd
import pysumma as ps
import xarray as xr
from matplotlib import cm
import seaborn as sns
from matplotlib.collections import LineCollection
from matplotlib.colors import ListedColormap, BoundaryNorm
sns.set_context('talk')
mpl.style.use('seaborn-bright')
mpl.rcParams['figure.figsize'] = (18, 12)
dt_list = [-4.0, -2.0, 0.0, 2.0, 4.0]
def return_config_dict(site):
default_params = {'zminLayer1': 0.0075, 'zminLayer2': 0.0100, 'zminLayer3': 0.0500, 'zminLayer4': 0.1000, 'zminLayer5': 0.2500,
'zmaxLayer1_lower': 0.0500, 'zmaxLayer2_lower': 0.2000, 'zmaxLayer3_lower': 0.5000, 'zmaxLayer4_lower': 1.0000,
'zmaxLayer1_upper': 0.0300, 'zmaxLayer2_upper': 0.1500, 'zmaxLayer3_upper': 0.3000, 'zmaxLayer4_upper': 0.7500 }
one_layer = {'zminLayer1': 0.0075, 'zminLayer2': 1000.0, 'zminLayer3': 1000., 'zminLayer4': 1000., 'zminLayer5': 1000.,
'zmaxLayer1_lower': 9000.0, 'zmaxLayer2_lower': 9000.0, 'zmaxLayer3_lower': 9000., 'zmaxLayer4_lower': 9000.,
'zmaxLayer1_upper': 3000.0, 'zmaxLayer2_upper': 9000.0, 'zmaxLayer3_upper': 9000., 'zmaxLayer4_upper': 9000. }
thick_2_layer = {'zminLayer1': 0.0075, 'zminLayer2': 0.0100, 'zminLayer3': 100., 'zminLayer4': 100., 'zminLayer5': 100.,
'zmaxLayer1_lower': 0.1250, 'zmaxLayer2_lower': 9000.0, 'zmaxLayer3_lower': 9000., 'zmaxLayer4_lower': 9000.,
'zmaxLayer1_upper': 0.1250, 'zmaxLayer2_upper': 9000.0, 'zmaxLayer3_upper': 9000., 'zmaxLayer4_upper': 9000. }
mid_2_layer = {'zminLayer1': 0.0075, 'zminLayer2': 0.0100, 'zminLayer3': 100., 'zminLayer4': 100., 'zminLayer5': 100.,
'zmaxLayer1_lower': 0.050, 'zmaxLayer2_lower': 9000.0, 'zmaxLayer3_lower': 9000., 'zmaxLayer4_lower': 9000.,
'zmaxLayer1_upper': 0.050, 'zmaxLayer2_upper': 9000.0, 'zmaxLayer3_upper': 9000., 'zmaxLayer4_upper': 9000. }
thin_2_layer = {'zminLayer1': 0.0075, 'zminLayer2': 0.0100, 'zminLayer3': 100., 'zminLayer4': 100., 'zminLayer5': 100.,
'zmaxLayer1_lower': 0.0250, 'zmaxLayer2_lower': 9000.0, 'zmaxLayer3_lower': 9000., 'zmaxLayer4_lower': 9000.,
'zmaxLayer1_upper': 0.0250, 'zmaxLayer2_upper': 9000.0, 'zmaxLayer3_upper': 9000., 'zmaxLayer4_upper': 9000. }
thick_3_layer = {'zminLayer1': 0.01, 'zminLayer2': 0.01, 'zminLayer3': 0.01, 'zminLayer4': 1000., 'zminLayer5': 1000.,
'zmaxLayer1_lower': 0.125, 'zmaxLayer2_lower': 0.2, 'zmaxLayer3_lower': 9000., 'zmaxLayer4_lower': 9000.,
'zmaxLayer1_upper': 0.125, 'zmaxLayer2_upper': 0.2, 'zmaxLayer3_upper': 9000., 'zmaxLayer4_upper': 9000.}
mid_3_layer = {'zminLayer1': 0.01, 'zminLayer2': 0.01, 'zminLayer3': 0.01, 'zminLayer4': 1000., 'zminLayer5': 1000.,
'zmaxLayer1_lower': 0.050, 'zmaxLayer2_lower': 0.2, 'zmaxLayer3_lower': 9000., 'zmaxLayer4_lower': 9000.,
'zmaxLayer1_upper': 0.050, 'zmaxLayer2_upper': 0.2, 'zmaxLayer3_upper': 9000., 'zmaxLayer4_upper': 9000.}
thin_3_layer = {'zminLayer1': 0.01, 'zminLayer2': 0.01, 'zminLayer3': 0.01, 'zminLayer4': 1000., 'zminLayer5': 1000.,
'zmaxLayer1_lower': 0.025, 'zmaxLayer2_lower': 0.2, 'zmaxLayer3_lower': 9000., 'zmaxLayer4_lower': 9000.,
'zmaxLayer1_upper': 0.025, 'zmaxLayer2_upper': 0.2, 'zmaxLayer3_upper': 9000., 'zmaxLayer4_upper': 9000.}
thick_4_layer = {'zminLayer1': 0.01, 'zminLayer2': 0.01, 'zminLayer3': 0.01, 'zminLayer4': 0.015, 'zminLayer5': 1000.,
'zmaxLayer1_lower': 0.075, 'zmaxLayer2_lower': 0.15, 'zmaxLayer3_lower': 0.20, 'zmaxLayer4_lower': 9000.,
'zmaxLayer1_upper': 0.075, 'zmaxLayer2_upper': 0.15, 'zmaxLayer3_upper': 0.20, 'zmaxLayer4_upper': 9000.}
mid_4_layer = {'zminLayer1': 0.01, 'zminLayer2': 0.01, 'zminLayer3': 0.01, 'zminLayer4': 0.015, 'zminLayer5': 1000.,
'zmaxLayer1_lower': 0.050, 'zmaxLayer2_lower': 0.125, 'zmaxLayer3_lower': 0.20, 'zmaxLayer4_lower': 9000.,
'zmaxLayer1_upper': 0.050, 'zmaxLayer2_upper': 0.125, 'zmaxLayer3_upper': 0.20, 'zmaxLayer4_upper': 9000.}
thin_4_layer = {'zminLayer1': 0.01, 'zminLayer2': 0.01, 'zminLayer3': 0.01, 'zminLayer4': 0.015, 'zminLayer5': 1000.,
'zmaxLayer1_lower': 0.025, 'zmaxLayer2_lower': 0.100, 'zmaxLayer3_lower': 0.20, 'zmaxLayer4_lower': 9000.,
'zmaxLayer1_upper': 0.025, 'zmaxLayer2_upper': 0.100, 'zmaxLayer3_upper': 0.20, 'zmaxLayer4_upper': 9000.}
dt_list = ['-2.0', '+0.0', '+2.0', '+4.0']
config_all = {'{}K_CLM'.format(dt): {'file_manager': f'./{site}/file_manager_{dt}K.txt',
'decisions': {'snowLayers': 'CLM_2010'},
'parameters': default_params} for dt in dt_list}
config_1layer = {'{}K_1L_all'.format(dt): {'file_manager': f'./{site}/file_manager_{dt}K.txt',
'decisions': {'snowLayers': 'CLM_2010'},
'parameters': one_layer} for dt in dt_list}
config_2layer = {'{}K_2L_thin'.format(dt): {'file_manager': f'./{site}/file_manager_{dt}K.txt',
'decisions': {'snowLayers': 'CLM_2010'},
'parameters': thin_2_layer} for dt in dt_list}
config_2layer.update({'{}K_2L_mid'.format(dt): {'file_manager': f'./{site}/file_manager_{dt}K.txt',
'decisions': {'snowLayers': 'CLM_2010'},
'parameters': mid_2_layer} for dt in dt_list})
config_2layer.update({'{}K_2L_thick'.format(dt): {'file_manager': f'./{site}/file_manager_{dt}K.txt',
'decisions': {'snowLayers': 'CLM_2010'},
'parameters': thick_2_layer} for dt in dt_list})
config_3layer = {'{}K_3L_thin'.format(dt): {'file_manager': f'./{site}/file_manager_{dt}K.txt',
'decisions': {'snowLayers': 'CLM_2010'},
'parameters': thin_3_layer} for dt in dt_list}
config_3layer.update({'{}K_3L_mid'.format(dt): {'file_manager': f'./{site}/file_manager_{dt}K.txt',
'decisions': {'snowLayers': 'CLM_2010'},
'parameters': mid_3_layer} for dt in dt_list})
config_3layer.update({'{}K_3L_thick'.format(dt): {'file_manager': f'./{site}/file_manager_{dt}K.txt',
'decisions': {'snowLayers': 'CLM_2010'},
'parameters': thick_3_layer} for dt in dt_list})
config_4layer = {'{}K_4L_thin'.format(dt): {'file_manager': f'./{site}/file_manager_{dt}K.txt',
'decisions': {'snowLayers': 'CLM_2010'},
'parameters': thin_4_layer} for dt in dt_list}
config_4layer.update({'{}K_4L_mid'.format(dt): {'file_manager': f'./{site}/file_manager_{dt}K.txt',
'decisions': {'snowLayers': 'CLM_2010'},
'parameters': mid_4_layer} for dt in dt_list})
config_4layer.update({'{}K_4L_thick'.format(dt): {'file_manager': f'./{site}/file_manager_{dt}K.txt',
'decisions': {'snowLayers': 'CLM_2010'},
'parameters': thick_4_layer} for dt in dt_list})
config_jrdn = {'{}K_JRDN'.format(dt): {'file_manager': f'./{site}/file_manager_{dt}K.txt',
'decisions': {'snowLayers': 'jrdn1991'},
'parameters': default_params} for dt in dt_list}
config_all.update(config_jrdn)
config_all.update(config_1layer)
config_all.update(config_2layer)
config_all.update(config_3layer)
config_all.update(config_4layer)
return config_all
for site in ['dana', 'reynolds', 'coldeport']:
print(site)
executable = '/pool0/data/andrbenn/summa/bin/summa.exe'
config_all = return_config_dict(site)
ens_dt = ps.Ensemble(executable, config_all, num_workers=24)
ens_dt.run('local')
assert len(ens_dt.summary()['error']) == 0
| run_simulations/run_simulations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="uAttKaKmT435"
# <div class="devsite-table-wrapper"><table class="tfo-notebook-buttons" align="left">
# <td><a target="_blank" href="https://www.tensorflow.org/tfx/tutorials/transform/census">
# <img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a></td>
# <td><a target="_blank" href="https://colab.sandbox.google.com/github/tensorflow/tfx/blob/master/docs/tutorials/transform/census.ipynb">
# <img src="https://www.tensorflow.org/images/colab_logo_32px.png">Run in Google Colab</a></td>
# <td><a target="_blank" href="https://github.com/tensorflow/tfx/blob/master/docs/tutorials/transform/census.ipynb">
# <img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png">View source on GitHub</a></td>
# <td><a target="_blank" href="https://storage.googleapis.com/tensorflow_docs/tfx/docs/tutorials/transform/census.ipynb">
# <img width=32px src="https://www.tensorflow.org/images/download_logo_32px.png">Download notebook</a></td>
# </table></div>
# + [markdown] id="tghWegsjhpkt"
# ##### Copyright 2020 The TensorFlow Authors.
# + cellView="form" id="rSGJWC5biBiG"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="mPt5BHTwy_0F"
# # Preprocessing data with TensorFlow Transform
# ***The Feature Engineering Component of TensorFlow Extended (TFX)***
#
# This example colab notebook provides a somewhat more advanced example of how <a target='_blank' href='https://www.tensorflow.org/tfx/transform/'>TensorFlow Transform</a> (`tf.Transform`) can be used to preprocess data using exactly the same code for both training a model and serving inferences in production.
#
# TensorFlow Transform is a library for preprocessing input data for TensorFlow, including creating features that require a full pass over the training dataset. For example, using TensorFlow Transform you could:
#
# * Normalize an input value by using the mean and standard deviation
# * Convert strings to integers by generating a vocabulary over all of the input values
# * Convert floats to integers by assigning them to buckets, based on the observed data distribution
#
# TensorFlow has built-in support for manipulations on a single example or a batch of examples. `tf.Transform` extends these capabilities to support full passes over the entire training dataset.
#
# The output of `tf.Transform` is exported as a TensorFlow graph which you can use for both training and serving. Using the same graph for both training and serving can prevent skew, since the same transformations are applied in both stages.
#
# Key Point: In order to understand `tf.Transform` and how it works with Apache Beam, you'll need to know a little bit about Apache Beam itself. The <a target='_blank' href='https://beam.apache.org/documentation/programming-guide/'>Beam Programming Guide</a> is a great place to start.
# + [markdown] id="_tQUubddMvnP"
# ##What we're doing in this example
#
# In this example we'll be processing a <a target='_blank' href='https://archive.ics.uci.edu/ml/machine-learning-databases/adult'>widely used dataset containing census data</a>, and training a model to do classification. Along the way we'll be transforming the data using `tf.Transform`.
#
# Key Point: As a modeler and developer, think about how this data is used and the potential benefits and harm a model's predictions can cause. A model like this could reinforce societal biases and disparities. Is a feature relevant to the problem you want to solve or will it introduce bias? For more information, read about <a target='_blank' href='https://developers.google.com/machine-learning/fairness-overview/'>ML fairness</a>.
#
# Note: <a target='_blank' href='https://www.tensorflow.org/tfx/model_analysis'>TensorFlow Model Analysis</a> is a powerful tool for understanding how well your model predicts for various segments of your data, including understanding how your model may reinforce societal biases and disparities.
# + [markdown] id="f7z7z62TmTNT"
# ### Upgrade Pip
#
# To avoid upgrading Pip in a system when running locally, check to make sure that we're running in Colab. Local systems can of course be upgraded separately.
# + id="Pjqv52BOmTau"
try:
import colab
# !pip install --upgrade pip
except:
pass
# + [markdown] id="OeonII4omTr1"
# ### Install TensorFlow Transform
#
# **Note: In Google Colab, because of package updates, the first time you run this cell you must restart the runtime (Runtime > Restart runtime ...).**
# + id="9Ak6XDO5mT3m"
# !pip install tensorflow-transform
# + [markdown] id="RptgLn2RYuK3"
# ## Python check, imports, and globals
# First we'll make sure that we're using Python 3, and then go ahead and install and import the stuff we need.
# + id="tFcdSuXTidhH"
import sys
# Confirm that we're using Python 3
assert sys.version_info.major == 3, 'Oops, not running Python 3. Use Runtime > Change runtime type'
# + id="K4QXVIM7iglN"
import math
import os
import pprint
import tensorflow as tf
print('TF: {}'.format(tf.__version__))
import apache_beam as beam
print('Beam: {}'.format(beam.__version__))
import tensorflow_transform as tft
import tensorflow_transform.beam as tft_beam
print('Transform: {}'.format(tft.__version__))
from tfx_bsl.public import tfxio
from tfx_bsl.coders.example_coder import RecordBatchToExamples
# !wget https://storage.googleapis.com/artifacts.tfx-oss-public.appspot.com/datasets/census/adult.data
# !wget https://storage.googleapis.com/artifacts.tfx-oss-public.appspot.com/datasets/census/adult.test
train = './adult.data'
test = './adult.test'
# + [markdown] id="CxOxaaOYRfl7"
# ### Name our columns
# We'll create some handy lists for referencing the columns in our dataset.
# + id="-bsr1nLHqyg_"
CATEGORICAL_FEATURE_KEYS = [
'workclass',
'education',
'marital-status',
'occupation',
'relationship',
'race',
'sex',
'native-country',
]
NUMERIC_FEATURE_KEYS = [
'age',
'capital-gain',
'capital-loss',
'hours-per-week',
]
OPTIONAL_NUMERIC_FEATURE_KEYS = [
'education-num',
]
ORDERED_CSV_COLUMNS = [
'age', 'workclass', 'fnlwgt', 'education', 'education-num',
'marital-status', 'occupation', 'relationship', 'race', 'sex',
'capital-gain', 'capital-loss', 'hours-per-week', 'native-country', 'label'
]
LABEL_KEY = 'label'
# + [markdown] id="qtTn4at8rurk"
# ###Define our features and schema
# Let's define a schema based on what types the columns are in our input. Among other things this will help with importing them correctly.
# + id="5oS2RfyCrzMr"
RAW_DATA_FEATURE_SPEC = dict(
[(name, tf.io.FixedLenFeature([], tf.string))
for name in CATEGORICAL_FEATURE_KEYS] +
[(name, tf.io.FixedLenFeature([], tf.float32))
for name in NUMERIC_FEATURE_KEYS] +
[(name, tf.io.VarLenFeature(tf.float32))
for name in OPTIONAL_NUMERIC_FEATURE_KEYS] +
[(LABEL_KEY, tf.io.FixedLenFeature([], tf.string))]
)
SCHEMA = tft.tf_metadata.dataset_metadata.DatasetMetadata(
tft.tf_metadata.schema_utils.schema_from_feature_spec(RAW_DATA_FEATURE_SPEC)).schema
# + [markdown] id="zdXy9lo4t45d"
# ###Setting hyperparameters and basic housekeeping
# Constants and hyperparameters used for training. The bucket size includes all listed categories in the dataset description as well as one extra for "?" which represents unknown.
#
# Note: The number of instances will be computed by `tf.Transform` in future versions, in which case it can be read from the metadata. Similarly BUCKET_SIZES will not be needed as this information will be stored in the metadata for each of the columns.
# + id="8WHyOkC9uL71"
testing = os.getenv("WEB_TEST_BROWSER", False)
NUM_OOV_BUCKETS = 1
if testing:
TRAIN_NUM_EPOCHS = 1
NUM_TRAIN_INSTANCES = 1
TRAIN_BATCH_SIZE = 1
NUM_TEST_INSTANCES = 1
else:
TRAIN_NUM_EPOCHS = 16
NUM_TRAIN_INSTANCES = 32561
TRAIN_BATCH_SIZE = 128
NUM_TEST_INSTANCES = 16281
# Names of temp files
TRANSFORMED_TRAIN_DATA_FILEBASE = 'train_transformed'
TRANSFORMED_TEST_DATA_FILEBASE = 'test_transformed'
EXPORTED_MODEL_DIR = 'exported_model_dir'
# + [markdown] id="0a1ns5KswDb2"
# ##Preprocessing with `tf.Transform`
# + [markdown] id="KKd3mCLNVYmg"
# ###Create a `tf.Transform` preprocessing_fn
# The _preprocessing function_ is the most important concept of tf.Transform. A preprocessing function is where the transformation of the dataset really happens. It accepts and returns a dictionary of tensors, where a tensor means a [`Tensor`](https://www.tensorflow.org/versions/r1.15/api_docs/python/tf/Tensor) or [`SparseTensor`](https://www.tensorflow.org/versions/r1.15/api_docs/python/tf/SparseTensor). There are two main groups of API calls that typically form the heart of a preprocessing function:
#
# 1. **TensorFlow Ops:** Any function that accepts and returns tensors, which usually means TensorFlow ops. These add TensorFlow operations to the graph that transforms raw data into transformed data one feature vector at a time. These will run for every example, during both training and serving.
# 2. **TensorFlow Transform Analyzers:** Any of the analyzers provided by tf.Transform. Analyzers also accept and return tensors, but unlike TensorFlow ops they only run once, during training, and typically make a full pass over the entire training dataset. They create [tensor constants](https://www.tensorflow.org/versions/r1.15/api_docs/python/tf/constant), which are added to your graph. For example, `tft.min` computes the minimum of a tensor over the training dataset. tf.Transform provides a fixed set of analyzers, but this will be extended in future versions.
#
# Caution: When you apply your preprocessing function to serving inferences, the constants that were created by analyzers during training do not change. If your data has trend or seasonality components, plan accordingly.
# + id="LDrzuYH0WFc2"
def preprocessing_fn(inputs):
"""Preprocess input columns into transformed columns."""
# Since we are modifying some features and leaving others unchanged, we
# start by setting `outputs` to a copy of `inputs.
outputs = inputs.copy()
# Scale numeric columns to have range [0, 1].
for key in NUMERIC_FEATURE_KEYS:
outputs[key] = tft.scale_to_0_1(inputs[key])
for key in OPTIONAL_NUMERIC_FEATURE_KEYS:
# This is a SparseTensor because it is optional. Here we fill in a default
# value when it is missing.
sparse = tf.sparse.SparseTensor(inputs[key].indices, inputs[key].values,
[inputs[key].dense_shape[0], 1])
dense = tf.sparse.to_dense(sp_input=sparse, default_value=0.)
# Reshaping from a batch of vectors of size 1 to a batch to scalars.
dense = tf.squeeze(dense, axis=1)
outputs[key] = tft.scale_to_0_1(dense)
# For all categorical columns except the label column, we generate a
# vocabulary but do not modify the feature. This vocabulary is instead
# used in the trainer, by means of a feature column, to convert the feature
# from a string to an integer id.
for key in CATEGORICAL_FEATURE_KEYS:
outputs[key] = tft.compute_and_apply_vocabulary(
tf.strings.strip(inputs[key]),
num_oov_buckets=NUM_OOV_BUCKETS,
vocab_filename=key)
# For the label column we provide the mapping from string to index.
table_keys = ['>50K', '<=50K']
with tf.init_scope():
initializer = tf.lookup.KeyValueTensorInitializer(
keys=table_keys,
values=tf.cast(tf.range(len(table_keys)), tf.int64),
key_dtype=tf.string,
value_dtype=tf.int64)
table = tf.lookup.StaticHashTable(initializer, default_value=-1)
# Remove trailing periods for test data when the data is read with tf.data.
label_str = tf.strings.regex_replace(inputs[LABEL_KEY], r'\.', '')
label_str = tf.strings.strip(label_str)
data_labels = table.lookup(label_str)
transformed_label = tf.one_hot(
indices=data_labels, depth=len(table_keys), on_value=1.0, off_value=0.0)
outputs[LABEL_KEY] = tf.reshape(transformed_label, [-1, len(table_keys)])
return outputs
# + [markdown] id="rgAGOAdFWRn2"
# ###Transform the data
# Now we're ready to start transforming our data in an Apache Beam pipeline.
#
# 1. Read in the data using the CSV reader
# 1. Transform it using a preprocessing pipeline that scales numeric data and converts categorical data from strings to int64 values indices, by creating a vocabulary for each category
# 1. Write out the result as a `TFRecord` of `Example` protos, which we will use for training a model later
#
# <aside class="key-term"><b>Key Term:</b> <a target='_blank' href='https://beam.apache.org/'>Apache Beam</a> uses a <a target='_blank' href='https://beam.apache.org/documentation/programming-guide/#applying-transforms'>special syntax to define and invoke transforms</a>. For example, in this line:
#
# <code><blockquote>result = pass_this | 'name this step' >> to_this_call</blockquote></code>
#
# The method <code>to_this_call</code> is being invoked and passed the object called <code>pass_this</code>, and <a target='_blank' href='https://stackoverflow.com/questions/50519662/what-does-the-redirection-mean-in-apache-beam-python'>this operation will be referred to as <code>name this step</code> in a stack trace</a>. The result of the call to <code>to_this_call</code> is returned in <code>result</code>. You will often see stages of a pipeline chained together like this:
#
# <code><blockquote>result = apache_beam.Pipeline() | 'first step' >> do_this_first() | 'second step' >> do_this_last()</blockquote></code>
#
# and since that started with a new pipeline, you can continue like this:
#
# <code><blockquote>next_result = result | 'doing more stuff' >> another_function()</blockquote></code></aside>
# + id="BQAPzdCRwi5d"
def transform_data(train_data_file, test_data_file, working_dir):
"""Transform the data and write out as a TFRecord of Example protos.
Read in the data using the CSV reader, and transform it using a
preprocessing pipeline that scales numeric data and converts categorical data
from strings to int64 values indices, by creating a vocabulary for each
category.
Args:
train_data_file: File containing training data
test_data_file: File containing test data
working_dir: Directory to write transformed data and metadata to
"""
# The "with" block will create a pipeline, and run that pipeline at the exit
# of the block.
with beam.Pipeline() as pipeline:
with tft_beam.Context(temp_dir=tempfile.mkdtemp()):
# Create a TFXIO to read the census data with the schema. To do this we
# need to list all columns in order since the schema doesn't specify the
# order of columns in the csv.
# We first read CSV files and use BeamRecordCsvTFXIO whose .BeamSource()
# accepts a PCollection[bytes] because we need to patch the records first
# (see "FixCommasTrainData" below). Otherwise, tfxio.CsvTFXIO can be used
# to both read the CSV files and parse them to TFT inputs:
# csv_tfxio = tfxio.CsvTFXIO(...)
# raw_data = (pipeline | 'ToRecordBatches' >> csv_tfxio.BeamSource())
csv_tfxio = tfxio.BeamRecordCsvTFXIO(
physical_format='text',
column_names=ORDERED_CSV_COLUMNS,
schema=SCHEMA)
# Read in raw data and convert using CSV TFXIO. Note that we apply
# some Beam transformations here, which will not be encoded in the TF
# graph since we don't do the from within tf.Transform's methods
# (AnalyzeDataset, TransformDataset etc.). These transformations are just
# to get data into a format that the CSV TFXIO can read, in particular
# removing spaces after commas.
raw_data = (
pipeline
| 'ReadTrainData' >> beam.io.ReadFromText(
train_data_file, coder=beam.coders.BytesCoder())
| 'FixCommasTrainData' >> beam.Map(
lambda line: line.replace(b', ', b','))
| 'DecodeTrainData' >> csv_tfxio.BeamSource())
# Combine data and schema into a dataset tuple. Note that we already used
# the schema to read the CSV data, but we also need it to interpret
# raw_data.
raw_dataset = (raw_data, csv_tfxio.TensorAdapterConfig())
# The TFXIO output format is chosen for improved performance.
transformed_dataset, transform_fn = (
raw_dataset | tft_beam.AnalyzeAndTransformDataset(
preprocessing_fn, output_record_batches=True))
# Transformed metadata is not necessary for encoding.
transformed_data, _ = transformed_dataset
# Extract transformed RecordBatches, encode and write them to the given
# directory.
_ = (
transformed_data
| 'EncodeTrainData' >>
beam.FlatMapTuple(lambda batch, _: RecordBatchToExamples(batch))
| 'WriteTrainData' >> beam.io.WriteToTFRecord(
os.path.join(working_dir, TRANSFORMED_TRAIN_DATA_FILEBASE)))
# Now apply transform function to test data. In this case we remove the
# trailing period at the end of each line, and also ignore the header line
# that is present in the test data file.
raw_test_data = (
pipeline
| 'ReadTestData' >> beam.io.ReadFromText(
test_data_file, skip_header_lines=1,
coder=beam.coders.BytesCoder())
| 'FixCommasTestData' >> beam.Map(
lambda line: line.replace(b', ', b','))
| 'RemoveTrailingPeriodsTestData' >> beam.Map(lambda line: line[:-1])
| 'DecodeTestData' >> csv_tfxio.BeamSource())
raw_test_dataset = (raw_test_data, csv_tfxio.TensorAdapterConfig())
# The TFXIO output format is chosen for improved performance.
transformed_test_dataset = (
(raw_test_dataset, transform_fn)
| tft_beam.TransformDataset(output_record_batches=True))
# Transformed metadata is not necessary for encoding.
transformed_test_data, _ = transformed_test_dataset
# Extract transformed RecordBatches, encode and write them to the given
# directory.
_ = (
transformed_test_data
| 'EncodeTestData' >>
beam.FlatMapTuple(lambda batch, _: RecordBatchToExamples(batch))
| 'WriteTestData' >> beam.io.WriteToTFRecord(
os.path.join(working_dir, TRANSFORMED_TEST_DATA_FILEBASE)))
# Will write a SavedModel and metadata to working_dir, which can then
# be read by the tft.TFTransformOutput class.
_ = (
transform_fn
| 'WriteTransformFn' >> tft_beam.WriteTransformFn(working_dir))
# + [markdown] id="TnaMyRMJ03bR"
# ##Using our preprocessed data to train a model using tf.keras
#
# To show how `tf.Transform` enables us to use the same code for both training and serving, and thus prevent skew, we're going to train a model. To train our model and prepare our trained model for production we need to create input functions. The main difference between our training input function and our serving input function is that training data contains the labels, and production data does not. The arguments and returns are also somewhat different.
#
# Note: This section uses tf.keras for training. If you are looking for an example using tf.estimator for training, please see the next section.
# + [markdown] id="M8xCZKNc2wAS"
# ###Create an input function for training
# + id="775Y7BTpHBmb"
def _make_training_input_fn(tf_transform_output, transformed_examples,
batch_size):
"""An input function reading from transformed data, converting to model input.
Args:
tf_transform_output: Wrapper around output of tf.Transform.
transformed_examples: Base filename of examples.
batch_size: Batch size.
Returns:
The input data for training or eval, in the form of k.
"""
def input_fn():
return tf.data.experimental.make_batched_features_dataset(
file_pattern=transformed_examples,
batch_size=batch_size,
features=tf_transform_output.transformed_feature_spec(),
reader=tf.data.TFRecordDataset,
label_key=LABEL_KEY,
shuffle=True).prefetch(tf.data.experimental.AUTOTUNE)
return input_fn
# + [markdown] id="nYeuthrs27vl"
# ###Create an input function for serving
#
# Let's create an input function that we could use in production, and prepare our trained model for serving.
# + id="n4TH_ZoSHSny"
def _make_serving_input_fn(tf_transform_output, raw_examples, batch_size):
"""An input function reading from raw data, converting to model input.
Args:
tf_transform_output: Wrapper around output of tf.Transform.
raw_examples: Base filename of examples.
batch_size: Batch size.
Returns:
The input data for training or eval, in the form of k.
"""
def get_ordered_raw_data_dtypes():
result = []
for col in ORDERED_CSV_COLUMNS:
if col not in RAW_DATA_FEATURE_SPEC:
result.append(0.0)
continue
spec = RAW_DATA_FEATURE_SPEC[col]
if isinstance(spec, tf.io.FixedLenFeature):
result.append(spec.dtype)
else:
result.append(0.0)
return result
def input_fn():
dataset = tf.data.experimental.make_csv_dataset(
file_pattern=raw_examples,
batch_size=batch_size,
column_names=ORDERED_CSV_COLUMNS,
column_defaults=get_ordered_raw_data_dtypes(),
prefetch_buffer_size=0,
ignore_errors=True)
tft_layer = tf_transform_output.transform_features_layer()
def transform_dataset(data):
raw_features = {}
for key, val in data.items():
if key not in RAW_DATA_FEATURE_SPEC:
continue
if isinstance(RAW_DATA_FEATURE_SPEC[key], tf.io.VarLenFeature):
raw_features[key] = tf.RaggedTensor.from_tensor(
tf.expand_dims(val, -1)).to_sparse()
continue
raw_features[key] = val
transformed_features = tft_layer(raw_features)
data_labels = transformed_features.pop(LABEL_KEY)
return (transformed_features, data_labels)
return dataset.map(
transform_dataset,
num_parallel_calls=tf.data.experimental.AUTOTUNE).prefetch(
tf.data.experimental.AUTOTUNE)
return input_fn
# + [markdown] id="LyNTX7CO8AAz"
# ###Train, Evaluate, and Export our model
# + id="OzE9HwLDIPqL"
def export_serving_model(tf_transform_output, model, output_dir):
"""Exports a keras model for serving.
Args:
tf_transform_output: Wrapper around output of tf.Transform.
model: A keras model to export for serving.
output_dir: A directory where the model will be exported to.
"""
# The layer has to be saved to the model for keras tracking purpases.
model.tft_layer = tf_transform_output.transform_features_layer()
@tf.function
def serve_tf_examples_fn(serialized_tf_examples):
"""Serving tf.function model wrapper."""
feature_spec = RAW_DATA_FEATURE_SPEC.copy()
feature_spec.pop(LABEL_KEY)
parsed_features = tf.io.parse_example(serialized_tf_examples, feature_spec)
transformed_features = model.tft_layer(parsed_features)
outputs = model(transformed_features)
classes_names = tf.constant([['0', '1']])
classes = tf.tile(classes_names, [tf.shape(outputs)[0], 1])
return {'classes': classes, 'scores': outputs}
concrete_serving_fn = serve_tf_examples_fn.get_concrete_function(
tf.TensorSpec(shape=[None], dtype=tf.string, name='inputs'))
signatures = {'serving_default': concrete_serving_fn}
# This is required in order to make this model servable with model_server.
versioned_output_dir = os.path.join(output_dir, '1')
model.save(versioned_output_dir, save_format='tf', signatures=signatures)
# + id="6i_lhWH8IZrk"
def train_and_evaluate(working_dir,
num_train_instances=NUM_TRAIN_INSTANCES,
num_test_instances=NUM_TEST_INSTANCES):
"""Train the model on training data and evaluate on test data.
Args:
working_dir: The location of the Transform output.
num_train_instances: Number of instances in train set
num_test_instances: Number of instances in test set
Returns:
The results from the estimator's 'evaluate' method
"""
train_data_path_pattern = os.path.join(working_dir,
TRANSFORMED_TRAIN_DATA_FILEBASE + '*')
eval_data_path_pattern = os.path.join(working_dir,
TRANSFORMED_TEST_DATA_FILEBASE + '*')
tf_transform_output = tft.TFTransformOutput(working_dir)
train_input_fn = _make_training_input_fn(
tf_transform_output, train_data_path_pattern, batch_size=TRAIN_BATCH_SIZE)
train_dataset = train_input_fn()
# Evaluate model on test dataset.
eval_input_fn = _make_training_input_fn(
tf_transform_output, eval_data_path_pattern, batch_size=TRAIN_BATCH_SIZE)
validation_dataset = eval_input_fn()
feature_spec = tf_transform_output.transformed_feature_spec().copy()
feature_spec.pop(LABEL_KEY)
inputs = {}
for key, spec in feature_spec.items():
if isinstance(spec, tf.io.VarLenFeature):
inputs[key] = tf.keras.layers.Input(
shape=[None], name=key, dtype=spec.dtype, sparse=True)
elif isinstance(spec, tf.io.FixedLenFeature):
inputs[key] = tf.keras.layers.Input(
shape=spec.shape, name=key, dtype=spec.dtype)
else:
raise ValueError('Spec type is not supported: ', key, spec)
encoded_inputs = {}
for key in inputs:
feature = tf.expand_dims(inputs[key], -1)
if key in CATEGORICAL_FEATURE_KEYS:
num_buckets = tf_transform_output.num_buckets_for_transformed_feature(key)
encoding_layer = (
tf.keras.layers.experimental.preprocessing.CategoryEncoding(
max_tokens=num_buckets, output_mode='binary', sparse=False))
encoded_inputs[key] = encoding_layer(feature)
else:
encoded_inputs[key] = feature
stacked_inputs = tf.concat(tf.nest.flatten(encoded_inputs), axis=1)
output = tf.keras.layers.Dense(100, activation='relu')(stacked_inputs)
output = tf.keras.layers.Dense(70, activation='relu')(output)
output = tf.keras.layers.Dense(50, activation='relu')(output)
output = tf.keras.layers.Dense(20, activation='relu')(output)
output = tf.keras.layers.Dense(2, activation='sigmoid')(output)
model = tf.keras.Model(inputs=inputs, outputs=output)
model.compile(optimizer='adam',
loss='binary_crossentropy',
metrics=['accuracy'])
pprint.pprint(model.summary())
model.fit(train_dataset, validation_data=validation_dataset,
epochs=TRAIN_NUM_EPOCHS,
steps_per_epoch=math.ceil(num_train_instances / TRAIN_BATCH_SIZE),
validation_steps=math.ceil(num_test_instances / TRAIN_BATCH_SIZE))
# Export the model.
exported_model_dir = os.path.join(working_dir, EXPORTED_MODEL_DIR)
export_serving_model(tf_transform_output, model, exported_model_dir)
metrics_values = model.evaluate(validation_dataset, steps=num_test_instances)
metrics_labels = model.metrics_names
return {l: v for l, v in zip(metrics_labels, metrics_values)}
# + [markdown] id="JrFJ9Zax8QAh"
# ###Put it all together
# We've created all the stuff we need to preprocess our census data, train a model, and prepare it for serving. So far we've just been getting things ready. It's time to start running!
#
# Note: Scroll the output from this cell to see the whole process. The results will be at the bottom.
# + id="xe7JdSin86Ez"
import tempfile
temp = os.path.join(tempfile.gettempdir(), 'keras')
transform_data(train, test, temp)
results = train_and_evaluate(temp)
pprint.pprint(results)
# + [markdown] id="APEUSA9boKgT"
# ## (Optional) Using our preprocessed data to train a model using tf.estimator
#
# If you would rather use an Estimator model instead of a Keras model, the code
# in this section shows how to do that.
# + [markdown] id="QcBWjr3ioZbl"
# ###Create an input function for training
# + id="kFO0MeWQ228a"
def _make_training_input_fn(tf_transform_output, transformed_examples,
batch_size):
"""Creates an input function reading from transformed data.
Args:
tf_transform_output: Wrapper around output of tf.Transform.
transformed_examples: Base filename of examples.
batch_size: Batch size.
Returns:
The input function for training or eval.
"""
def input_fn():
"""Input function for training and eval."""
dataset = tf.data.experimental.make_batched_features_dataset(
file_pattern=transformed_examples,
batch_size=batch_size,
features=tf_transform_output.transformed_feature_spec(),
reader=tf.data.TFRecordDataset,
shuffle=True)
transformed_features = tf.compat.v1.data.make_one_shot_iterator(
dataset).get_next()
# Extract features and label from the transformed tensors.
transformed_labels = tf.where(
tf.equal(transformed_features.pop(LABEL_KEY), 1))
return transformed_features, transformed_labels[:,1]
return input_fn
# + [markdown] id="22XOsZ-noez-"
# ###Create an input function for serving
#
# Let's create an input function that we could use in production, and prepare our trained model for serving.
# + cellView="code" id="NN5FVg343Jea"
def _make_serving_input_fn(tf_transform_output):
"""Creates an input function reading from raw data.
Args:
tf_transform_output: Wrapper around output of tf.Transform.
Returns:
The serving input function.
"""
raw_feature_spec = RAW_DATA_FEATURE_SPEC.copy()
# Remove label since it is not available during serving.
raw_feature_spec.pop(LABEL_KEY)
def serving_input_fn():
"""Input function for serving."""
# Get raw features by generating the basic serving input_fn and calling it.
# Here we generate an input_fn that expects a parsed Example proto to be fed
# to the model at serving time. See also
# tf.estimator.export.build_raw_serving_input_receiver_fn.
raw_input_fn = tf.estimator.export.build_parsing_serving_input_receiver_fn(
raw_feature_spec, default_batch_size=None)
serving_input_receiver = raw_input_fn()
# Apply the transform function that was used to generate the materialized
# data.
raw_features = serving_input_receiver.features
transformed_features = tf_transform_output.transform_raw_features(
raw_features)
return tf.estimator.export.ServingInputReceiver(
transformed_features, serving_input_receiver.receiver_tensors)
return serving_input_fn
# + [markdown] id="Vc9Edp8A7dsI"
# ###Wrap our input data in FeatureColumns
# Our model will expect our data in TensorFlow FeatureColumns.
# + id="6qOFOvBk7oJX"
def get_feature_columns(tf_transform_output):
"""Returns the FeatureColumns for the model.
Args:
tf_transform_output: A `TFTransformOutput` object.
Returns:
A list of FeatureColumns.
"""
# Wrap scalars as real valued columns.
real_valued_columns = [tf.feature_column.numeric_column(key, shape=())
for key in NUMERIC_FEATURE_KEYS]
# Wrap categorical columns.
one_hot_columns = [
tf.feature_column.indicator_column(
tf.feature_column.categorical_column_with_identity(
key=key,
num_buckets=(NUM_OOV_BUCKETS +
tf_transform_output.vocabulary_size_by_name(
vocab_filename=key))))
for key in CATEGORICAL_FEATURE_KEYS]
return real_valued_columns + one_hot_columns
# + [markdown] id="f6FyMzMcpOgT"
# ###Train, Evaluate, and Export our model
# + id="8iGQ0jeq8IWr"
def train_and_evaluate(working_dir, num_train_instances=NUM_TRAIN_INSTANCES,
num_test_instances=NUM_TEST_INSTANCES):
"""Train the model on training data and evaluate on test data.
Args:
working_dir: Directory to read transformed data and metadata from and to
write exported model to.
num_train_instances: Number of instances in train set
num_test_instances: Number of instances in test set
Returns:
The results from the estimator's 'evaluate' method
"""
tf_transform_output = tft.TFTransformOutput(working_dir)
run_config = tf.estimator.RunConfig()
estimator = tf.estimator.LinearClassifier(
feature_columns=get_feature_columns(tf_transform_output),
config=run_config,
loss_reduction=tf.losses.Reduction.SUM)
# Fit the model using the default optimizer.
train_input_fn = _make_training_input_fn(
tf_transform_output,
os.path.join(working_dir, TRANSFORMED_TRAIN_DATA_FILEBASE + '*'),
batch_size=TRAIN_BATCH_SIZE)
estimator.train(
input_fn=train_input_fn,
max_steps=TRAIN_NUM_EPOCHS * num_train_instances / TRAIN_BATCH_SIZE)
# Evaluate model on test dataset.
eval_input_fn = _make_training_input_fn(
tf_transform_output,
os.path.join(working_dir, TRANSFORMED_TEST_DATA_FILEBASE + '*'),
batch_size=1)
# Export the model.
serving_input_fn = _make_serving_input_fn(tf_transform_output)
exported_model_dir = os.path.join(working_dir, EXPORTED_MODEL_DIR)
estimator.export_saved_model(exported_model_dir, serving_input_fn)
return estimator.evaluate(input_fn=eval_input_fn, steps=num_test_instances)
# + [markdown] id="5k8LrDPZpZsK"
# ###Put it all together
# We've created all the stuff we need to preprocess our census data, train a model, and prepare it for serving. So far we've just been getting things ready. It's time to start running!
#
# Note: Scroll the output from this cell to see the whole process. The results will be at the bottom.
# + id="P_1_2dB6pdc2"
import tempfile
temp = os.path.join(tempfile.gettempdir(), 'estimator')
transform_data(train, test, temp)
results = train_and_evaluate(temp)
pprint.pprint(results)
# + [markdown] id="ICqetCnSjwp1"
# ##What we did
# In this example we used `tf.Transform` to preprocess a dataset of census data, and train a model with the cleaned and transformed data. We also created an input function that we could use when we deploy our trained model in a production environment to perform inference. By using the same code for both training and inference we avoid any issues with data skew. Along the way we learned about creating an Apache Beam transform to perform the transformation that we needed for cleaning the data. We also saw how to use this transformed data to train a model using either `tf.keras` or `tf.estimator`. This is just a small piece of what TensorFlow Transform can do! We encourage you to dive into `tf.Transform` and discover what it can do for you.
| docs/tutorials/transform/census.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # User Analytics in the Telecommunication Industry
# ## Import Libraries
## Import Libraries
import sys
import os
sys.path.append(os.path.abspath(os.path.join('..')))
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import numpy as np
from pandas.api.types import is_string_dtype, is_numeric_dtype
# %matplotlib inline
# ## Import Data
CSV_PATH = "../data/raw/rawData.csv"
# +
# taking a csv file path and reading a dataframe
def read_proccessed_data(csv_path):
try:
df = pd.read_csv(csv_path)
print("file read as csv")
return df
except FileNotFoundError:
print("file not found")
# -
## getting number of columns, row and column information
def get_data_info(xDR_df: pd.DataFrame):
row_count, col_count = xDR_df.shape
print(f"Number of rows: {row_count}")
print(f"Number of columns: {col_count}")
return xDR_df.info()
## basic statistics of each column and see the data at glance
def get_statistics_info(xDR_df: pd.DataFrame):
return xDR_df.describe(include='all')
# +
# reading the extracted tweeter data and getting information
xDR_df = read_proccessed_data(CSV_PATH)
get_data_info(xDR_df)
get_statistics_info(xDR_df)
# -
# ### Distinct value counts/frequencies of each column to determine if there are any columns with only a single value/all different values
pd.DataFrame(xDR_df.apply(lambda x: len(x.value_counts(dropna=False)), axis=0),
columns=['Unique Value Count']).sort_values(by='Unique Value Count', ascending=True)
sns.set(rc={'figure.figsize':(15.7,8.27)})
sns.heatmap(xDR_df.isnull(),yticklabels=False,cbar=False,cmap='viridis')
def percent_missing(df):
totalCells = np.product(df.shape)
missingCount = df.isnull().sum()
totalMissing = missingCount.sum()
return round((totalMissing / totalCells) * 100, 2)
# +
print("The Telco Telecom dataset contains", percent_missing(xDR_df), "%", "missing values.")
# -
def percent_missing_for_col(df, col_name: str):
total_count = len(df[col_name])
if total_count <= 0:
return 0.0
missing_count = df[col_name].isnull().sum()
return round((missing_count / total_count) * 100, 2)
null_percent_df = pd.DataFrame(columns = ['column', 'null_percent'])
columns = xDR_df.columns.values.tolist()
null_percent_df['column'] = columns
null_percent_df['null_percent'] = null_percent_df['column'].map(lambda x: percent_missing_for_col(xDR_df, x))
null_percent_df.sort_values(by=['null_percent'], ascending = False)
# #### Roughly for 10 data sets column morthan 50 percent fo the data is missing, it looks like we are just missing too much of the data to do something useful with at a basic level so it is better to drop thos columns. For the rest of the data the proportion of missing data is likely small enough for reasonable replacement with some form of imputation.
#Drop columns with morethan 50% missing values
def drop_columns(xDR_df, columns=[]):
return xDR_df.drop(columns, axis=1)
columns_to_be_dropped = null_percent_df[null_percent_df['null_percent'] > 50]['column'].to_list()
xDR_df = drop_columns(xDR_df, columns_to_be_dropped)
def percent_missing(df):
totalCells = np.product(df.shape)
missingCount = df.isnull().sum()
totalMissing = missingCount.sum()
return round((totalMissing / totalCells) * 100, 2)
print("The Telco Telecom dataset contains", percent_missing(xDR_df), "%", "missing values.")
# ## Now let us deal with this missing values
# +
missing_count=xDR_df.isnull().sum() #the count of missing values
value_count=xDR_df.isnull().count() #the count of all values
missing_percentage=round(missing_count/value_count*100,2) #the percentage of missing values
missing_df=pd.DataFrame({'count':missing_count,'percentage':missing_percentage}) #create a datafreame
missing_df_for_plot=missing_df[missing_df.percentage!=0.00]
font = {'family' : 'normal',
'weight' : 'bold',
'size' : 22}
plt.rc('font', **font)
barchart=missing_df_for_plot.plot.bar(y='percentage',figsize=(10,8))
for index, percentage in enumerate(missing_percentage):
if percentage >0.00:
barchart.text(index, percentage,"")
# -
# ### For Bearer Id, IMSI, MSISDN, IMEI,Handset Manufacturer,Handset Type I am going to use forward fill because it is the better way to fill unique columns. Beacuse I think it will have fair distribution of values
xDR_df['Bearer Id'] = xDR_df['Bearer Id'].fillna(method='ffill')
xDR_df['IMSI'] = xDR_df['IMSI'].fillna(method='ffill')
xDR_df['MSISDN/Number'] = xDR_df['MSISDN/Number'].fillna(method='ffill')
xDR_df['IMEI'] = xDR_df['IMEI'].fillna(method='ffill')
xDR_df['Handset Manufacturer'] = xDR_df['Handset Manufacturer'].fillna(method='ffill')
xDR_df['Handset Type'] = xDR_df['Handset Type'].fillna(method='ffill')
# fill 'Last Location Name' column with unknown
xDR_df['Last Location Name'] = xDR_df['Last Location Name'].fillna('Uknown')
# ## For the rest of the columns I am going to use mean to feel the missing values because it is simple.
#numerical data
xDR_df.fillna(xDR_df.mean(), inplace=True)
def percent_missing(df):
totalCells = np.product(df.shape)
missingCount = df.isnull().sum()
totalMissing = missingCount.sum()
return round((totalMissing / totalCells) * 100, 2)
print("The Telco Telecom dataset contains", percent_missing(xDR_df), "%", "missing values.")
xDR_df.info()
# ## Remove Duplicate rows
xDR_df.drop_duplicates(inplace=True)
xDR_df.info()
# ## Save the processed dataframe to CSV
xDR_df.to_csv("../data/processed/processed.csv")
| notebooks/dataPreProcessing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # 积分
# ## 符号积分
# 积分与求导的关系:
#
# $$\frac{d}{dx} F(x) = f(x)
# \Rightarrow F(x) = \int f(x) dx$$
#
# 符号运算可以用 `sympy` 模块完成。
#
# 先导入 `init_printing` 模块方便其显示:
from sympy import init_printing
init_printing()
from sympy import symbols, integrate
import sympy
# 产生 x 和 y 两个符号变量,并进行运算:
x, y = symbols('x y')
sympy.sqrt(x ** 2 + y ** 2)
# 对于生成的符号变量 `z`,我们将其中的 `x` 利用 `subs` 方法替换为 `3`:
z = sympy.sqrt(x ** 2 + y ** 2)
z.subs(x, 3)
# 再替换 `y`:
z.subs(x, 3).subs(y, 4)
# 还可以从 `sympy.abc` 中导入现成的符号变量:
from sympy.abc import theta
y = sympy.sin(theta) ** 2
y
# 对 y 进行积分:
Y = integrate(y)
Y
# 计算 $Y(\pi) - Y(0)$:
# +
import numpy as np
np.set_printoptions(precision=3)
Y.subs(theta, np.pi) - Y.subs(theta, 0)
# -
# 计算 $\int_0^\pi y d\theta$ :
integrate(y, (theta, 0, sympy.pi))
# 显示的是字符表达式,查看具体数值可以使用 `evalf()` 方法,或者传入 `numpy.pi`,而不是 `sympy.pi` :
integrate(y, (theta, 0, sympy.pi)).evalf()
integrate(y, (theta, 0, np.pi))
# 根据牛顿莱布尼兹公式,这两个数值应该相等。
#
# 产生不定积分对象:
Y_indef = sympy.Integral(y)
Y_indef
print type(Y_indef)
# 定积分:
Y_def = sympy.Integral(y, (theta, 0, sympy.pi))
Y_def
# 产生函数 $Y(x) = \int_0^x sin^2(\theta) d\theta$,并将其向量化:
Y_raw = lambda x: integrate(y, (theta, 0, x))
Y = np.vectorize(Y_raw)
# +
# %matplotlib inline
import matplotlib.pyplot as plt
x = np.linspace(0, 2 * np.pi)
p = plt.plot(x, Y(x))
t = plt.title(r'$Y(x) = \int_0^x sin^2(\theta) d\theta$')
# -
# ## 数值积分
# 数值积分:
#
# $$F(x) = \lim_{n \rightarrow \infty} \sum_{i=0}^{n-1} f(x_i)(x_{i+1}-x_i)
# \Rightarrow F(x) = \int_{x_0}^{x_n} f(x) dx$$
#
# 导入贝塞尔函数:
from scipy.special import jv
def f(x):
return jv(2.5, x)
x = np.linspace(0, 10)
p = plt.plot(x, f(x), 'k-')
# ### `quad` 函数
# Quadrature 积分的原理参见:
#
# http://en.wikipedia.org/wiki/Numerical_integration#Quadrature_rules_based_on_interpolating_functions
#
# quad 返回一个 (积分值,误差) 组成的元组:
from scipy.integrate import quad
interval = [0, 6.5]
value, max_err = quad(f, *interval)
# 积分值:
print value
# 最大误差:
print max_err
# 积分区间图示,蓝色为正,红色为负:
print "integral = {:.9f}".format(value)
print "upper bound on error: {:.2e}".format(max_err)
x = np.linspace(0, 10, 100)
p = plt.plot(x, f(x), 'k-')
x = np.linspace(0, 6.5, 45)
p = plt.fill_between(x, f(x), where=f(x)>0, color="blue")
p = plt.fill_between(x, f(x), where=f(x)<0, color="red", interpolate=True)
# ### 积分到无穷
# +
from numpy import inf
interval = [0., inf]
def g(x):
return np.exp(-x ** 1/2)
# -
value, max_err = quad(g, *interval)
x = np.linspace(0, 10, 50)
fig = plt.figure(figsize=(10,3))
p = plt.plot(x, g(x), 'k-')
p = plt.fill_between(x, g(x))
plt.annotate(r"$\int_0^{\infty}e^{-x^1/2}dx = $" + "{}".format(value), (4, 0.6),
fontsize=16)
print "upper bound on error: {:.1e}".format(max_err)
# ### 双重积分
# 假设我们要进行如下的积分:
#
# $$ I_n = \int \limits_0^{\infty} \int \limits_1^{\infty} \frac{e^{-xt}}{t^n}dt dx = \frac{1}{n}$$
def h(x, t, n):
"""core function, takes x, t, n"""
return np.exp(-x * t) / (t ** n)
# 一种方式是调用两次 `quad` 函数,不过这里 `quad` 的返回值不能向量化,所以使用了修饰符 `vectorize` 将其向量化:
from numpy import vectorize
@vectorize
def int_h_dx(t, n):
"""Time integrand of h(x)."""
return quad(h, 0, np.inf, args=(t, n))[0]
@vectorize
def I_n(n):
return quad(int_h_dx, 1, np.inf, args=(n))
I_n([0.5, 1.0, 2.0, 5])
# 或者直接调用 `dblquad` 函数,并将积分参数传入,传入方式有多种,后传入的先进行积分:
from scipy.integrate import dblquad
@vectorize
def I(n):
"""Same as I_n, but using the built-in dblquad"""
x_lower = 0
x_upper = np.inf
return dblquad(h,
lambda t_lower: 1, lambda t_upper: np.inf,
x_lower, x_upper, args=(n,))
I_n([0.5, 1.0, 2.0, 5])
# ## 采样点积分
# ### trapz 方法 和 simps 方法
from scipy.integrate import trapz, simps
# `sin` 函数, `100` 个采样点和 `5` 个采样点:
x_s = np.linspace(0, np.pi, 5)
y_s = np.sin(x_s)
x = np.linspace(0, np.pi, 100)
y = np.sin(x)
p = plt.plot(x, y, 'k:')
p = plt.plot(x_s, y_s, 'k+-')
p = plt.fill_between(x_s, y_s, color="gray")
# 采用 [trapezoidal 方法](https://en.wikipedia.org/wiki/Trapezoidal_rule) 和 [simpson 方法](https://en.wikipedia.org/wiki/Simpson%27s_rule) 对这些采样点进行积分(函数积分为 2):
result_s = trapz(y_s, x_s)
result_s_s = simps(y_s, x_s)
result = trapz(y, x)
print "Trapezoidal Integration over 5 points : {:.3f}".format(result_s)
print "Simpson Integration over 5 points : {:.3f}".format(result_s_s)
print "Trapezoidal Integration over 100 points : {:.3f}".format(result)
# ### 使用 ufunc 进行积分
# `Numpy` 中有很多 `ufunc` 对象:
type(np.add)
np.info(np.add.accumulate)
# `np.add.accumulate` 相当于 `cumsum` :
result_np = np.add.accumulate(y) * (x[1] - x[0]) - (x[1] - x[0]) / 2
p = plt.plot(x, - np.cos(x) + np.cos(0), 'rx')
p = plt.plot(x, result_np)
# ### 速度比较
# 计算积分:$$\int_0^x sin \theta d\theta$$
import sympy
from sympy.abc import x, theta
sympy_x = x
x = np.linspace(0, 20 * np.pi, 1e+4)
y = np.sin(x)
sympy_y = vectorize(lambda x: sympy.integrate(sympy.sin(theta), (theta, 0, x)))
# `numpy` 方法:
# %timeit np.add.accumulate(y) * (x[1] - x[0])
y0 = np.add.accumulate(y) * (x[1] - x[0])
print y0[-1]
# `quad` 方法:
# %timeit quad(np.sin, 0, 20 * np.pi)
y2 = quad(np.sin, 0, 20 * np.pi, full_output=True)
print "result = ", y2[0]
print "number of evaluations", y2[-1]['neval']
# `trapz` 方法:
# %timeit trapz(y, x)
y1 = trapz(y, x)
print y1
# `simps` 方法:
# %timeit simps(y, x)
y3 = simps(y, x)
print y3
# `sympy` 积分方法:
# %timeit sympy_y(20 * np.pi)
y4 = sympy_y(20 * np.pi)
print y4
| lijin-THU:notes-python/04-scipy/04.06-integration-in-python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# *The environment needs to have lxml and openssl. This can be installed via Anaconda Navigator*
#
# From Windows' Start menu, launch Anaconda Prompt (inside Anaconda).
# Run:
#
# ```
# $ conda install -c anaconda lxml
# $ conda install -c anaconda openssl
# ```
# + [markdown] slideshow={"slide_type": "slide"}
# **Step 1. Set up emulab geni-lib package for CloudLab**
# -
# !cd barnstorm-geni-lib-95b218670744 & python setup.py build
# !cd barnstorm-geni-lib-95b218670744 & python setup.py install
# **Step 2: Reload geni-lib for the first time**
#
# On the top bar of this notebook, select `Kernel` and then `Restart`
# **Step 3: Test emulab geni-lib installation**
#
# Executing this cell should produce an XML element with the following content:
#
# ```
# <rspec xmlns:client="http://www.protogeni.net/resources/rspec/ext/client/1" xmlns:emulab="http://www.protogeni.net/resources/rspec/ext/emulab/1" xmlns:jacks="http://www.protogeni.net/resources/rspec/ext/jacks/1" xmlns:xsi="http://www.w3.org/2001/XMLSchema-instance" xmlns="http://www.geni.net/resources/rspec/3" xsi:schemaLocation="http://www.geni.net/resources/rspec/3 http://www.geni.net/resources/rspec/3/request.xsd" type="request">
# <rspec_tour xmlns="http://www.protogeni.net/resources/rspec/ext/apt-tour/1">
# <description type="markdown">An example of constructing a profile with a single Xen VM.</description>
# <instructions type="markdown">Wait for the profile instance to start, and then log in to the VM via the
# ssh port specified below. (Note that in this case, you will need to access
# the VM through a high port on the physical host, since we have not requested
# a public IP address for the VM itself.)
# </instructions>
# </rspec_tour>
# <node client_id="node" exclusive="false">
# <sliver_type name="emulab-xen"/>
# </node>
# </rspec>
# ```
# +
# %%writefile examples/test.py
"""An example of constructing a profile with a single Xen VM.
Instructions:
Wait for the profile instance to start, and then log in to the VM via the
ssh port specified below. (Note that in this case, you will need to access
the VM through a high port on the physical host, since we have not requested
a public IP address for the VM itself.)
"""
import geni.portal as portal
import geni.rspec.pg as rspec
# Create a Request object to start building the RSpec.
request = portal.context.makeRequestRSpec()
# Create a XenVM
node = request.XenVM("node")
# Print the RSpec to the enclosing page.
portal.context.printRequestRSpec()
# -
# !python examples/test.py
# **Step 4: Make sure that you have a CloudLab-ready account:**
#
# See instructions at [the documentation](http://docs.cloudlab.us/users.html). This can be done either by
# - authorizing CloudLab to use you GENI account (go ahead and request to join the `PDC-edu-Lab` project), or
# - creating a new CloudLab account and request to join the `PDC-edu-Lab` project.
# **Step 5: Generate SSH Keys for interaction with CloudLab instances **
#
# - On the terminal, generate a pair of public/private SSH keys with no password:
# ```
# > ssh-keygen -t rsa
# Generating public/private rsa key pair.
# Enter file in which to save the key (/home/lngo/.ssh/id_rsa):
# Enter passphrase (empty for no passphrase):
# Enter same passphrase again:
# Your identification has been saved in /home/lngo/.ssh/id_rsa.
# Your public key has been saved in /home/lngo/.ssh/id_rsa.pub.
# The key fingerprint is:
# SHA256:l6oM32bbOKuzhksNak3FWSm7MojBerVuD4FI1j9RjUw lngo@localhost
# The key's randomart image is:
# +---[RSA 2048]----+
# | oE+. |
# | . .o=.. |
# |.o . .+o |
# |+o .o.o . |
# |oo.o++ .S o |
# |o o+++o o |
# | .o.=+. . |
# | . .+=oo+o |
# | ..+B*=+. |
# +----[SHA256]-----+
#
# > more .ssh/id_rsa.pub
# ssh-rsa AAAAB3NzaC1yc2EAAAADAQABAAABAQDR6N3MaD717Ti9+LQwKiJsEtwELpPaCDf3j57/iDFKDBB1kULJy/ve3Z0e1JItfzTGEpaAIzUPwRNRzxZ2aR2UQ0qde+8ibj9Ynq/3+UzG5b07FK6zDj35h/fqhaCun1lmhPEwaQh31FXnUqEw7eM5O7RgPGtdy4P7GxXhE5S1Om3fBWrFUD1njfkjL6Z/mMiUzFsxMvpf7HT9rhOVwaLliC9hjwtJPAhibmLEQdI1bHf1IEmJEXN2wBXcGjeUrooWgo1vofTV7rSztIN7zo+42DoVcfAZbepsMlLD9Kk0nly3x7kZNd5i4J3K8+dt9cOUq7j5QLqbvEtaUyOTqIQP LinhBNgo@surface4pro
# rsa
# ```
#
# - Login to `www.cloudlab.us`, select your username on the top right corner and select `Manage SSH Keys`.
#
# - Copy the content of the `id_rsa.pub` file into the `Key` box and click `Add Key`
| 0-preparations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: sax
# language: python
# name: sax
# ---
# # Thin film optimization
#
# contributed by [simbilod](https://github.com/simbilod), adapted by [flaport](https://github.com/flaport)
#
# In this notebook, we apply SAX to thin-film optimization and show how it can be used for wavelength-dependent parameter optimization.
#
# The language of transfer/scatter matrices is commonly used to calculate optical properties of thin-films. Many specialized methods exist for their optimization. However, SAX can be useful to cut down on developer time by circumventing the need to manually take gradients of complicated or often-changed objective functions, and by generating efficient code from simple syntax.
# ## Imports
import jax
import jax.example_libraries.optimizers as opt
import jax.numpy as jnp
import matplotlib.pyplot as plt
import sax # sax circuit simulator
import tqdm.notebook as tqdm
from tmm import coh_tmm
# ## Dielectric mirror Fabry-Pérot
#
# Consider a stack composed of only two materials, $n_A$ and $n_B$. Two types of transfer matrices characterize wave propagation in the system : interfaces described by Fresnel's equations, and propagation.
# For the two-material stack, this leads to 4 scatter matrices coefficients. Through reciprocity they can be constructed out of two independent ones :
# +
def fresnel_mirror_ij(ni=1.0, nj=1.0):
"""Model a (fresnel) interface between twoo refractive indices
Args:
ni: refractive index of the initial medium
nf: refractive index of the final
"""
r_fresnel_ij = (ni - nj) / (ni + nj) # i->j reflection
t_fresnel_ij = 2 * ni / (ni + nj) # i->j transmission
r_fresnel_ji = -r_fresnel_ij # j -> i reflection
t_fresnel_ji = (1 - r_fresnel_ij ** 2) / t_fresnel_ij # j -> i transmission
sdict = {
("in", "in"): r_fresnel_ij,
("in", "out"): t_fresnel_ij,
("out", "in"): t_fresnel_ji,
("out", "out"): r_fresnel_ji,
}
return sdict
def propagation_i(ni=1.0, di=0.5, wl=0.532):
"""Model the phase shift acquired as a wave propagates through medium A
Args:
ni: refractive index of medium (at wavelength wl)
di: [μm] thickness of layer
wl: [μm] wavelength
"""
prop_i = jnp.exp(1j * 2 * jnp.pi * ni * di / wl)
sdict = {
("in", "out"): prop_i,
("out", "in"): prop_i,
}
return sdict
# -
# A resonant cavity can be formed when a high index region is surrounded by low-index region :
# +
dielectric_fabry_perot = sax.circuit(
instances={
"air_B": fresnel_mirror_ij,
"B": propagation_i,
"B_air": fresnel_mirror_ij,
},
connections={
"air_B,out": "B,in",
"B,out": "B_air,in",
},
ports={
"in": "air_B,in",
"out": "B_air,out",
},
)
settings = sax.get_settings(dielectric_fabry_perot)
settings
# -
# Let's choose $n_A = 1$, $n_B = 2$, $d_B = 1000$ nm, and compute over the visible spectrum :
# +
settings = sax.copy_settings(settings)
settings["air_B"]["nj"] = 2.0
settings["B"]["ni"] = 2.0
settings["B_air"]["ni"] = 2.0
wls = jnp.linspace(0.380, 0.750, 200)
settings = sax.update_settings(settings, wl=wls)
# -
# Compute transmission and reflection, and compare to another package's results (https://github.com/sbyrnes321/tmm) :
# +
# %%time
sdict = dielectric_fabry_perot(**settings)
transmitted = sdict["in", "out"]
reflected = sdict["in", "in"]
# +
# %%time
# tmm syntax (https://github.com/sbyrnes321/tmm)
d_list = [jnp.inf, 0.500, jnp.inf]
n_list = [1, 2, 1]
# initialize lists of y-values to plot
rnorm = []
tnorm = []
Tnorm = []
Rnorm = []
for l in wls:
rnorm.append(coh_tmm("s", n_list, d_list, 0, l)["r"])
tnorm.append(coh_tmm("s", n_list, d_list, 0, l)["t"])
Tnorm.append(coh_tmm("s", n_list, d_list, 0, l)["T"])
Rnorm.append(coh_tmm("s", n_list, d_list, 0, l)["R"])
# +
plt.scatter(wls * 1e3, jnp.real(transmitted), label="t SAX")
plt.plot(wls * 1e3, jnp.real(jnp.array(tnorm)), "k", label="t tmm")
plt.scatter(wls * 1e3, jnp.real(reflected), label="r SAX")
plt.plot(wls * 1e3, jnp.real(jnp.array(rnorm)), "k--", label="r tmm")
plt.xlabel("λ [nm]")
plt.ylabel("Transmitted and reflected amplitude")
plt.legend(loc="upper right")
plt.title("Real part")
plt.show()
plt.scatter(wls * 1e3, jnp.imag(transmitted), label="t SAX")
plt.plot(wls * 1e3, jnp.imag(jnp.array(tnorm)), "k", label="t tmm")
plt.scatter(wls * 1e3, jnp.imag(reflected), label="r SAX")
plt.plot(wls * 1e3, jnp.imag(jnp.array(rnorm)), "k--", label="r tmm")
plt.xlabel("λ [nm]")
plt.ylabel("Transmitted and reflected amplitude")
plt.legend(loc="upper right")
plt.title("Imaginary part")
plt.show()
# -
# In terms of powers, we get the following. Due to the reflections at the interfaces, resonant behaviour is observed, with evenly-spaced maxima/minima in wavevector space :
plt.scatter(2 * jnp.pi / wls, jnp.abs(transmitted) ** 2, label="T SAX")
plt.plot(2 * jnp.pi / wls, Tnorm, "k", label="T tmm")
plt.scatter(2 * jnp.pi / wls, jnp.abs(reflected) ** 2, label="R SAX")
plt.plot(2 * jnp.pi / wls, Rnorm, "k--", label="R tmm")
plt.vlines(jnp.arange(3, 6) * jnp.pi / (2 * 0.5), ymin=0, ymax=1, color="k", linestyle="--", label="m$\pi$/nd")
plt.xlabel("k = 2$\pi$/λ [1/nm]")
plt.ylabel("Transmitted and reflected intensities")
plt.legend(loc="upper right")
plt.show()
# ### Optimization test
#
# Let's attempt to minimize transmission at 500 nm by varying thickness.
@jax.jit
def loss(thickness):
settings = sax.update_settings(sax.get_settings(dielectric_fabry_perot), wl=0.5)
settings["B"]["di"] = thickness
settings["air_B"]["nj"] = 2.0
settings["B"]["ni"] = 2.0
settings["B_air"]["ni"] = 2.0
sdict = dielectric_fabry_perot(**settings)
return jnp.abs(sdict["in", "out"]) ** 2
grad = jax.jit(jax.grad(loss))
# +
initial_thickness = 0.5
optim_init, optim_update, optim_params = opt.adam(step_size=0.01)
optim_state = optim_init(initial_thickness)
def train_step(step, optim_state):
thickness = optim_params(optim_state)
lossvalue = loss(thickness)
gradvalue = grad(thickness)
optim_state = optim_update(step, gradvalue, optim_state)
return lossvalue, optim_state
# -
range_ = tqdm.trange(100)
for step in range_:
lossvalue, optim_state = train_step(step, optim_state)
range_.set_postfix(loss=f"{lossvalue:.6f}")
thickness = optim_params(optim_state)
thickness
# +
settings = sax.update_settings(sax.get_settings(dielectric_fabry_perot), wl=wls)
settings["B"]["di"] = thickness
settings["air_B"]["nj"] = 2.0
settings["B"]["ni"] = 2.0
settings["B_air"]["ni"] = 2.0
sdict = dielectric_fabry_perot(**settings)
detected = sdict["in", "out"]
plt.plot(wls * 1e3, jnp.abs(transmitted) ** 2, label="Before (500 nm)")
plt.plot(wls * 1e3, jnp.abs(detected) ** 2, label=f"After ({thickness*1e3:.0f} nm)")
plt.vlines(0.5 * 1e3, 0.6, 1, "k", linestyle="--")
plt.xlabel("λ [nm]")
plt.ylabel("Transmitted intensity")
plt.legend(loc="lower right")
plt.title("Thickness optimization")
plt.show()
# -
# ## General Fabry-Pérot étalon
#
# We reuse the propagation matrix above, and instead of simple interface matrices, model Fabry-Pérot mirrors as general lossless reciprocal scatter matrices :
#
# $$ \left(\begin{array}{c}
# E_t \\
# E_r
# \end{array}\right) = E_{out} = SE_{in} = \left(\begin{array}{cc}
# t & r \\
# r & t
# \end{array}\right) \left(\begin{array}{c}
# E_0 \\
# 0
# \end{array}\right) $$
#
# For lossless reciprocal systems, we further have the requirements
#
# $$ |t|^2 + |r|^2 = 1 $$
#
# and
#
# $$ \angle t - \angle r = \pm \pi/2 $$
#
# The general Fabry-Pérot cavity is analytically described by :
# +
def airy_t13(t12, t23, r21, r23, wl, d=1.0, n=1.0):
"""General Fabry-Pérot transmission transfer function (Airy formula)
Args:
t12 and r12 : S-parameters of the first mirror
t23 and r23 : S-parameters of the second mirror
wl : wavelength
d : gap between the two mirrors (in units of wavelength)
n : index of the gap between the two mirrors
Returns:
t13 : complex transmission amplitude of the mirror-gap-mirror system
Note:
Each mirror is assumed to be lossless and reciprocal : tij = tji, rij = rji
"""
phi = n * 2 * jnp.pi * d / wl
return t12 * t23 * jnp.exp(-1j * phi) / (1 - r21 * r23 * jnp.exp(-2j * phi))
def airy_r13(t12, t23, r21, r23, wl, d=1.0, n=1.0):
"""General Fabry-Pérot reflection transfer function (Airy formula)
Args:
t12 and r12 : S-parameters of the first mirror
t23 and r23 : S-parameters of the second mirror
wl : wavelength
d : gap between the two mirrors (in units of wavelength)
n : index of the gap between the two mirrors
Returns:
r13 : complex reflection amplitude of the mirror-gap-mirror system
Note:
Each mirror is assumed to be lossless and reciprocal : tij = tji, rij = rji
"""
phi = n * 2 * jnp.pi * d / wl
return r21 + t12 * t12 * r23 * jnp.exp(-2j * phi) / (1 - r21 * r23 * jnp.exp(-2j * phi))
# -
# We need to implement the relationship between $t$ and $r$ for lossless reciprocal mirrors. The design parameter will be the amplitude and phase of the tranmission coefficient. The reflection coefficient is then fully determined :
# +
def t_complex(t_amp, t_ang):
return t_amp * jnp.exp(-1j * t_ang)
def r_complex(t_amp, t_ang):
r_amp = jnp.sqrt((1.0 - t_amp ** 2))
r_ang = t_ang - jnp.pi / 2
return r_amp * jnp.exp(-1j * r_ang)
# -
# Let's see the expected result for half-mirrors :
# +
t_initial = jnp.sqrt(0.5)
d_gap = 2.0
n_gap = 1.0
r_initial = r_complex(t_initial, 0.0)
wls = jnp.linspace(0.38, 0.78, 500)
T_analytical_initial = jnp.abs(airy_t13(t_initial, t_initial, r_initial, r_initial, wls, d=d_gap, n=n_gap)) ** 2
R_analytical_initial = jnp.abs(airy_r13(t_initial, t_initial, r_initial, r_initial, wls, d=d_gap, n=n_gap)) ** 2
plt.title(f"t={t_initial:1.3f}, d={d_gap} nm, n={n_gap}")
plt.plot(2 * jnp.pi / wls, T_analytical_initial, label="T")
plt.plot(2 * jnp.pi / wls, R_analytical_initial, label="R")
plt.vlines(jnp.arange(6, 11) * jnp.pi / 2.0, ymin=0, ymax=1, color="k", linestyle="--", label="m$\pi$/nd")
plt.xlabel("k = 2$\pi$/$\lambda$ [1/nm]")
plt.ylabel("Power (units of input)")
plt.legend()
plt.show()
plt.title(f"t={t_initial:1.3f}, d={d_gap} nm, n={n_gap}")
plt.plot(wls * 1e3, T_analytical_initial, label="T")
plt.plot(wls * 1e3, R_analytical_initial, label="R")
plt.xlabel("$\lambda$ (nm)")
plt.ylabel("Power (units of input)")
plt.legend()
plt.show()
# -
# Is power conserved? (to within 0.1%)
assert jnp.isclose(R_analytical_initial + T_analytical_initial, 1, 0.001).all()
# Now let's do the same with SAX by defining new elements :
# +
def mirror(t_amp=0.5**0.5, t_ang=0.0):
r_complex_val = r_complex(t_amp, t_ang)
t_complex_val = t_complex(t_amp, t_ang)
sdict = {
("in", "in"): r_complex_val,
("in", "out"): t_complex_val,
("out", "in"): t_complex_val, # (1 - r_complex_val**2)/t_complex_val, # t_ji
("out", "out"): r_complex_val, # -r_complex_val, # r_ji
}
return sdict
fabry_perot_tunable = sax.circuit(
instances={
"mirror1": mirror,
"gap": propagation_i,
"mirror2": mirror,
},
connections={
"mirror1,out": "gap,in",
"gap,out": "mirror2,in",
},
ports={
"in": "mirror1,in",
"out": "mirror2,out",
},
)
settings = sax.get_settings(fabry_perot_tunable)
settings
# +
fabry_perot_tunable = sax.circuit(
instances={
"mirror1": mirror,
"gap": propagation_i,
"mirror2": mirror,
},
connections={
"mirror1,out": "gap,in",
"gap,out": "mirror2,in",
},
ports={
"in": "mirror1,in",
"out": "mirror2,out",
},
)
settings = sax.get_settings(fabry_perot_tunable)
settings
# -
N = 100
wls = jnp.linspace(0.38, 0.78, N)
settings = sax.get_settings(fabry_perot_tunable)
settings = sax.update_settings(settings, wl=wls, t_amp=jnp.sqrt(0.5), t_ang=0.0)
settings["gap"]["ni"] = 1.0
settings["gap"]["di"] = 2.0
transmitted_initial = fabry_perot_tunable(**settings)["in", "out"]
reflected_initial = fabry_perot_tunable(**settings)["out", "out"]
T_analytical_initial = jnp.abs(airy_t13(t_initial, t_initial, r_initial, r_initial, wls, d=d_gap, n=n_gap))**2
R_analytical_initial = jnp.abs(airy_r13(t_initial, t_initial, r_initial, r_initial, wls, d=d_gap, n=n_gap))**2
plt.title(f"t={t_initial:1.3f}, d={d_gap} nm, n={n_gap}")
plt.plot(wls, T_analytical_initial, label="T theory")
plt.scatter(wls, jnp.abs(transmitted_initial) ** 2, label="T SAX")
plt.plot(wls, R_analytical_initial, label="R theory")
plt.scatter(wls, jnp.abs(reflected_initial) ** 2, label="R SAX")
plt.xlabel("k = 2$\pi$/$\lambda$ [1/nm]")
plt.ylabel("Power (units of input)")
plt.figlegend(framealpha=1.0)
plt.show()
# ## Wavelength-dependent Fabry-Pérot étalon
#
# Let's repeat with a model where parameters can be wavelength-dependent. To comply with the optimizer object, we will stack all design parameters in a single array :
ts_initial = jnp.zeros(2 * N)
ts_initial = jax.ops.index_update(ts_initial, jax.ops.index[0:N], jnp.sqrt(0.5))
# We will simply loop over all wavelengths, and use different $t$ parameters at each wavelength.
wls = jnp.linspace(0.38, 0.78, N)
transmitted = jnp.zeros_like(wls)
reflected = jnp.zeros_like(wls)
settings = sax.get_settings(fabry_perot_tunable)
settings = sax.update_settings(settings, wl=wls, t_amp=ts_initial[:N], t_ang=ts_initial[N:])
settings["gap"]["ni"] = 1.0
settings["gap"]["di"] = 2.0
# Perform computation
sdict = fabry_perot_tunable(**settings)
transmitted = jnp.abs(sdict["in", "out"]) ** 2
reflected = jnp.abs(sdict["in", "in"]) ** 2
plt.plot(wls * 1e3, T_analytical_initial, label="T theory")
plt.scatter(wls * 1e3, transmitted, label="T SAX")
plt.plot(wls * 1e3, R_analytical_initial, label="R theory")
plt.scatter(wls * 1e3, reflected, label="R SAX")
plt.xlabel("λ [nm]")
plt.ylabel("Transmitted and reflected intensities")
plt.legend(loc="upper right")
plt.title(f"t={t_initial:1.3f}, d={d_gap} nm, n={n_gap}")
plt.show()
# Since it seems to work, let's add a target and optimize some harmonics away :
# +
def lorentzian(l0, dl, wl, A):
return A / ((wl - l0) ** 2 + (0.5 * dl) ** 2)
target = lorentzian(533.0, 20.0, wls * 1e3, 100.0)
plt.scatter(wls * 1e3, transmitted, label="T SAX")
plt.scatter(wls * 1e3, reflected, label="R SAX")
plt.plot(wls * 1e3, target, "r", linewidth=2, label="target")
plt.xlabel("λ [nm]")
plt.ylabel("Transmitted and reflected intensities")
plt.legend(loc="upper right")
plt.title(f"t={t_initial:1.3f}, d={d_gap} nm, n={n_gap}")
plt.show()
# -
@jax.jit
def loss(ts):
N = len(ts[::2])
wls = jnp.linspace(0.38, 0.78, N)
target = lorentzian(533.0, 20.0, wls * 1e3, 100.0)
settings = sax.get_settings(fabry_perot_tunable)
settings = sax.update_settings(settings, wl=wls, t_amp=ts[:N], t_ang=ts[N:])
settings["gap"]["ni"] = 1.0
settings["gap"]["di"] = 2.0
sdict = fabry_perot_tunable(**settings)
transmitted = jnp.abs(sdict["in", "out"]) ** 2
return (jnp.abs(transmitted - target) ** 2).mean()
grad = jax.jit(jax.grad(loss))
# +
optim_init, optim_update, optim_params = opt.adam(step_size=0.001)
def train_step(step, optim_state):
ts = optim_params(optim_state)
lossvalue = loss(ts)
gradvalue = grad(ts)
optim_state = optim_update(step, gradvalue, optim_state)
return lossvalue, gradvalue, optim_state
# +
range_ = tqdm.trange(2000)
optim_state = optim_init(ts_initial)
for step in range_:
lossvalue, gradvalue, optim_state = train_step(step, optim_state)
range_.set_postfix(loss=f"{lossvalue:.6f}")
# -
# The optimized parameters are now wavelength-dependent :
# +
ts_optimal = optim_params(optim_state)
plt.scatter(wls * 1e3, ts_initial[:N], label="t initial")
plt.scatter(wls * 1e3, ts_optimal[:N], label="t optimal")
plt.xlabel("λ [nm]")
plt.ylabel("|t| $(\lambda)$")
plt.legend(loc="best")
plt.title(f"d={d_gap} nm, n={n_gap}")
plt.show()
plt.scatter(wls * 1e3, ts_initial[N:], label="t initial")
plt.scatter(wls * 1e3, ts_optimal[N:], label="t optimal")
plt.xlabel("λ [nm]")
plt.ylabel("angle $t (\lambda)$ (rad)")
plt.legend(loc="best")
plt.title(f"d={d_gap} nm, n={n_gap}")
plt.show()
# -
# Visualizing the result :
# +
wls = jnp.linspace(0.38, 0.78, N)
transmitted_optimal = jnp.zeros_like(wls)
reflected_optimal = jnp.zeros_like(wls)
settings = sax.get_settings(fabry_perot_tunable)
settings = sax.update_settings(
settings, wl=wls, t_amp=ts_optimal[:N], t_ang=ts_optimal[N:]
)
settings["gap"]["ni"] = 1.0
settings["gap"]["di"] = 2.0
transmitted_optimal = jnp.abs(fabry_perot_tunable(**settings)["in", "out"]) ** 2
reflected_optimal = jnp.abs(fabry_perot_tunable(**settings)["in", "in"]) ** 2
# -
plt.scatter(wls * 1e3, transmitted_optimal, label="T")
plt.scatter(wls * 1e3, reflected_optimal, label="R")
plt.plot(wls * 1e3, lorentzian(533, 20, wls * 1e3, 100), "r", label="target")
plt.xlabel("λ [nm]")
plt.ylabel("Transmitted and reflected intensities")
plt.legend(loc="upper right")
plt.title(f"Optimized t($\lambda$), d={d_gap} nm, n={n_gap}")
plt.show()
# The hard part is now to find physical stacks that physically implement $t(\lambda)$. However, the ease with which we can modify and complexify the loss function opens opportunities for regularization and more complicated objective functions.
#
# The models above are available in `sax.models.thinfilm`, and can straightforwardly be extended to propagation at an angle, s and p polarizations, nonreciprocal systems, and systems with losses.
| examples/05_thinfilm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
from sklearn.feature_extraction.text import CountVectorizer
from nltk.corpus import stopwords
import pickle
from sklearn.model_selection import train_test_split
import xgboost
data = pd.read_csv(r'Data\preprocessed_data.csv')
x_data = data["text"][pd.isna(data["text"])==False]
y_data = data["target"][pd.isna(data["text"])== False]
with open('Saved/countVectorizer.pkl', 'rb') as f:
vectorizer = pickle.load(f)
# +
model = xgboost.XGBClassifier()
model.load_model("Saved/xgboost_bestModel.json")
# -
x_test = vectorizer.transform(x_data).toarray()
y_test = y_data
model.score(x_test, y_test)
x_test.shape
| Gui/read.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# > This is one of the 100 recipes of the [IPython Cookbook](http://ipython-books.github.io/), the definitive guide to high-performance scientific computing and data science in Python.
#
# # 6.5. Converting matplotlib figures to d3.js visualizations with mpld3
# 1. First, we load NumPy and matplotlib as usual.
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# 2. Then, we enable the mpld3 figures in the notebook with a single function call.
from mpld3 import enable_notebook
enable_notebook()
# 3. Now, let's create a scatter plot with matplotlib.
X = np.random.normal(0, 1, (100, 3))
color = np.random.random(100)
size = 500 * np.random.random(100)
plt.figure(figsize=(6,4))
plt.scatter(X[:,0], X[:,1], c=color,
s=size, alpha=0.5, linewidths=2)
plt.grid(color='lightgray', alpha=0.7)
# The matplotlib figure is rendered with d3.js instead of the standard matplotlib backend. In particular, the figure is interactive (pan and zoom).
# 4. Now, we create a more complex example with multiple subplots representing different 2D projections of a 3D dataset. We use the `sharex` and `sharey` keywords in matplotlib's `subplots` function to automatically bind the x and y axes of the different figures. Panning and zooming in any of the subplots automatically updates all the other subplots.
fig, ax = plt.subplots(3, 3, figsize=(6, 6),
sharex=True, sharey=True)
fig.subplots_adjust(hspace=0.3)
X[::2,2] += 3
for i in range(3):
for j in range(3):
ax[i,j].scatter(X[:,i], X[:,j], c=color,
s=.1*size, alpha=0.5, linewidths=2)
ax[i,j].grid(color='lightgray', alpha=0.7)
# This use case is perfectly handled by mpld3: the d3.js subplots are also dynamically linked together.
# > You'll find all the explanations, figures, references, and much more in the book (to be released later this summer).
#
# > [IPython Cookbook](http://ipython-books.github.io/), by [<NAME>](http://cyrille.rossant.net), Packt Publishing, 2014 (500 pages).
| notebooks/chapter06_viz/05_mpld3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # MATPLOTLIB
# Visit the official website of __Matplotlib__ for more content in __[this link](https://matplotlib.org)__
# <img src="https://matplotlib.org/_static/logo2.png"
# alt="matplotlib" title="Matplotlib" height="400" width="400" />
# # Importing Matplotlib
import matplotlib.pyplot as plt
import numpy as np
# # Line Chart
# +
value_x = [133, 134, 135, 136, 137]
value_y = [100000, 178093, 199273, 179534, 124667]
plt.plot(value_x, value_y)
plt.title("Value X vs Value Y")
plt.xlabel("Value X")
plt.ylabel("Value Y")
plt.show()
# -
# # Scatter Plot
# +
value_x = [10, 15, 17, 18, 23, 19, 20]
value_y = [125, 143, 133, 122, 156, 125, 133]
plt.scatter(value_x, value_y)
plt.title("Value X vs Value Y")
plt.xlabel("Value X")
plt.ylabel("Value y")
plt.show()
# -
# # Numerical Bar Chart
# +
numbers = [1, 4, 3, 2, 4, 2, 1, 5, 7, 6]
total_bars = 4
plt.hist(numbers, bins = total_bars)
plt.title("Frequency of Value X")
plt.xlabel("Value X")
plt.ylabel("Frequency of X")
plt.show()
# -
# # Categorical Bar Chart
# +
labels = ["Data A", "Data B", "Data C", "Data D"]
value_by_label = [30, 35, 46, 40]
y_positions = range(len(labels))
# Creating our bar plot
plt.bar(y_positions, value_by_label)
plt.xticks(y_positions, labels)
plt.ylabel("y title")
plt.title("Your title")
plt.show()
# -
# # Categorial Bar Chart with Error
# +
values = ['Data A', 'Data B', 'Data C', 'Data D']
y_pos = np.arange(len(values))
frequency = [12.67254242, 9.56392691, 12.86695884, 11.7662192]
error = [0.85620101, 0.3312307, 0.2265702, 0.34091318]
_, bar = plt.subplots()
bar.barh(y_pos, frequency, xerr=error)
bar.set_yticks(y_pos)
bar.set_yticklabels(values)
bar.invert_yaxis()
bar.set_xlabel('Data of Frequency')
bar.set_title('Title of Chart')
# -
# # Stack Plot
# +
x = [50, 150, 250, 350, 450]
y1 = [12, 23, 15, 18, 12]
y2 = [13, 12, 19, 15, 19]
y3 = [16, 14, 18, 18, 12]
y = np.vstack([y1, y2, y3])
labels = ["Element 1", "Element 2", "Element 3"]
fig, ax = plt.subplots()
ax.stackplot(x, y1, y2, y3, labels=labels)
ax.legend(loc='upper left')
plt.show()
# -
# # Pie Diagram
# +
labels = ['Element 1', 'Element 2', 'Element 3', 'Element 4']
sizes = [15, 30, 45, 10]
explode = (0, 0.0, 0.0, 0.1)
plt.pie(sizes,
explode=explode,
labels=labels,
autopct='%.2f%%',
startangle=90)
plt.axis('equal')
plt.show()
# -
# # Box Plot
# +
values = [1, 2, 5, 6, 6, 7, 7, 8, 8, 8, 9, 10, 21]
plt.boxplot(values)
plt.yticks(range(np.min(values)-2, np.max(values)+2))
plt.ylabel("Value")
plt.show()
| 1. Basic Tools/Getting Started Matplotlib.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
import pydiablo.char as d2
#reload(d2)
#paladin = d2.Paladin()
#paladin.equip_weapon(ebotd_cb)
#mstr_prev = ''
#print '\t'.join(['ias', '2HS', '', '2HT', ''])
#print '\t'.join(['', 'first', 'last', 'first', 'last'])
#Paladin.generate_table('2HS', 100, 0, 0)
#print BearDruid.anim_duration('A1','STF',100,35,0,40)
#print Amazon.foreswing_duration('A1', 'BOW', 100, 0, 10, 0)
#print Amazon.anim_duration('A1', 'BOW', 100, 0, 10, 0)
d2.write_bp_table(sys.stdout, d2.Amazon.attack_duration, 'BOW', 100, 37, 10)
d2.write_bp_table(sys.stdout, d2.WolfBarbarian.attack_duration, 'STF', 50, 59, 10, WIAS=90)
d2.write_bp_table(sys.stdout, d2.WolfBarbarian.attack_duration, 'STF', 50, 59, 0, WIAS=60)
d2.write_bp_table(sys.stdout, d2.WolfBarbarian.attack_duration, '2HS', 50, 59, 5, WIAS=60)
d2.write_bp_table(sys.stdout, d2.WolfDruid.attack_duration, 'STF', 50, 72, 10, WIAS=50)
d2.write_bp_table(sys.stdout, d2.WolfDruid.attack_duration, 'STF', 50, 72, 10, WIAS=60)
d2.write_bp_table(sys.stdout, d2.WolfDruid.attack_duration, '2HS', 50, 72, 5, WIAS=60)
d2.write_bp_table(sys.stdout, d2.Paladin.zeal_duration, '2HS', 100, 37, 10, WIAS=0)
d2.write_bp_table(sys.stdout, d2.Act2Merc.jab_duration, 'HTH', 100, 0, 10)
# -
import pydiablo
print(pydiablo.__version__)
#print d2.Character.animdata.get_data('PAA11HS')
#print d2.Paladin.foreswing_duration('A1','1HS', 100, 0, 0, 0)
#print d2.anim_duration(8, 256, 100, 0, 0, 0)
#print d2.Character.animdata.get_data('PAA21HS')
#print d2.anim_duration(10, 256, 100, 0, 0, 0)
for wsm in [-10, 0, 10]:
d2.write_bp_table(sys.stdout, d2.Act1Merc.attack_duration, 'HTH', 100, 0, wsm)
d2.write_bp_table(sys.stdout, d2.Act1Merc.attack_duration, 'HTH', 100, 32, wsm)
d2.write_bp_table(sys.stdout, d2.Act1Merc.attack_duration, 'HTH', 100, 33, wsm)
2.27*39
2.5*32
16/2.5
14/2.27
d2.write_bp_table(sys.stdout, d2.Amazon.attack_duration, 'BOW', 100, 32, 10)
35+30+10+20
d2.write_bp_table(sys.stdout, d2.Amazon.fend_duration, '2HT', 100, 0, 20, ntargets=5)
d2.write_bp_table(sys.stdout, d2.Amazon.attack_duration, '2HT', 100, 0, 20)
d2.write_bp_table(sys.stdout, d2.Act2Merc.jab_duration, 'HTH', 100, 0, 0)
| iascalc.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.7 64-bit (''nlp-meeting'': conda)'
# language: python
# name: python3
# ---
# +
import pandas as pd
audio = ['test1.flac', 'test2.flac']
text = ['hello world', "stanford online"]
df = pd.DataFrame({"audio": audio, "text": text})
df.head()
# +
# how to add append string to each row of dataframe
sample = r"C:\Users\codenamewei\.run"
df.audio = sample + df.audio
df.head()
# -
| notebooks/pandas/append_value_to_row.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # [ATM 623: Climate Modeling](../index.ipynb)
# [<NAME>](http://www.atmos.albany.edu/facstaff/brose/index.html), University at Albany
# # Lecture 1: Climate models, the global energy budget and Fun with Python
# -
# ## Warning: content out of date and not maintained
#
# You really should be looking at [The Climate Laboratory book](https://brian-rose.github.io/ClimateLaboratoryBook) by <NAME>, where all the same content (and more!) is kept up to date.
#
# ***Here you are likely to find broken links and broken code.***
# + [markdown] slideshow={"slide_type": "skip"}
# ### About these notes:
#
# This document uses the interactive [`Jupyter notebook`](https://jupyter.org) format. The notes can be accessed in several different ways:
#
# - The interactive notebooks are hosted on `github` at https://github.com/brian-rose/ClimateModeling_courseware
# - The latest versions can be viewed as static web pages [rendered on nbviewer](http://nbviewer.ipython.org/github/brian-rose/ClimateModeling_courseware/blob/master/index.ipynb)
# - A complete snapshot of the notes as of May 2017 (end of spring semester) are [available on Brian's website](http://www.atmos.albany.edu/facstaff/brose/classes/ATM623_Spring2017/Notes/index.html).
#
# [Also here is a legacy version from 2015](http://www.atmos.albany.edu/facstaff/brose/classes/ATM623_Spring2015/Notes/index.html).
#
# Many of these notes make use of the `climlab` package, available at https://github.com/brian-rose/climlab
# + [markdown] slideshow={"slide_type": "slide"}
# ## Contents
#
# 1. [What is a Climate Model?](#section1)
# 2. [The observed global energy budget](#section2)
# 3. [Quantifying the planetary energy budget](#section3)
# 4. [Using Python to compute emission to space](#section4)
# + [markdown] slideshow={"slide_type": "slide"}
# ____________
# <a id='section1'></a>
#
# ## 1. What is a Climate Model?
# ____________
# + [markdown] slideshow={"slide_type": "slide"}
# First, some thoughts on modeling from [xkcd](https://xkcd.com)
#
# 
# + [markdown] slideshow={"slide_type": "slide"}
# Let's be a little pedantic and decompose that question:
#
# - what is Climate?
# - what is a Model?
# + [markdown] slideshow={"slide_type": "fragment"}
# **Climate** is
#
# - statistics of weather, e.g. space and time averages of temperature and precip.
# - (statistics might also mean higher-order stats: variability etc)
# + [markdown] slideshow={"slide_type": "fragment"}
# A **model** is
#
# - not easy to define!
# + [markdown] slideshow={"slide_type": "slide"}
# Wikipedia: http://en.wikipedia.org/wiki/Conceptual_model
#
# > In the most general sense, a model is anything used in any way to represent anything else. Some models are physical objects, for instance, a toy model which may be assembled, and may even be made to work like the object it represents. Whereas, a conceptual model is a model made of the composition of concepts, that thus exists only in the mind. Conceptual models are used to help us know, understand, or simulate the subject matter they represent.
# + [markdown] slideshow={"slide_type": "slide"}
# <NAME> (statistician):
# > Essentially, all models are wrong, but some are useful.”
# + [markdown] slideshow={"slide_type": "slide"}
# From the Climate Modelling Primer, 4th ed (McGuffie and Henderson-Sellers):
#
# > In the broadest sense, models are for learning about the world (in our case, the climate) and the learning takes place in the contruction and the manipulation of the model, as anyone who has watched a child build idealised houses or spaceships with Lego, or built with it themselves, will know. Climate models are, likewise, idealised representations of a complicated and complex reality through which our understanding of the climate has significantly expanded. All models involve some ignoring, distoring and approximating, but gradually they allow us to build understanding of the system being modelled. A child's Lego construction typically contains the essential elements of the real objects, improves with attention to detail, helps them understand the real world, but is never confused with the real thing.
# + [markdown] slideshow={"slide_type": "slide"}
# ### A minimal definition of a climate model
#
# *A representation of the exchange of energy between the Earth system and space, and its effects on average surface temperature.*
#
# (what average?)
#
# Note the focus on **planetary energy budget**. That’s the key to all climate modeling.
# + [markdown] slideshow={"slide_type": "slide"}
# ____________
# <a id='section2'></a>
#
# ## 2. The observed global energy budget
# ____________
#
# The figure below shows current best estimates of the *global, annual mean* energy fluxes through the climate system.
#
# We will look at many of these processes in detail throughout the course.
# -
# 
# + [markdown] slideshow={"slide_type": "slide"}
# ## Things to note:
#
# ### On the shortwave side
#
# - global mean albedo is 101.9 W m$^{-2}$ / 341.3 W m$^{-2}$ = 0.299
# - Reflection off clouds = 79 W m$^{-2}$
# - Off surface = 23 W m$^{-2}$
# - 3 times as much reflection off clouds as off surface
#
# Why?? Think about both areas of ice and snow, and the fact that sunlight has to travel through cloudy atmosphere to get to the ice and snow. Also there is some absorption of shortwave by the atmosphere.
#
# - Atmospheric absorption = 78 W m$^{-2}$
# (so about the same as reflected by clouds)
#
# QUESTION: Which gases contribute to shortwave absorption?
#
# - O$_3$ and H$_2$O mostly.
# - We will look at this later.
# + [markdown] slideshow={"slide_type": "slide"}
# ### On the longwave side
#
# - Observed emission from the SURFACE is 396 W m$^{-2}$
# - very close to the blackbody emission $\sigma T^4$ at $T = 288$ K (the global mean surface temperature).
# - BUT emission to space is much smaller = 239 W m$^{-2}$
#
# QUESTION: What do we call this? (greenhouse effect)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Look at net numbers…
#
# - Net absorbed = 0.9 W m$^{-2}$
# - Why?
# - Where is that heat going?
#
# Note, the exchanges of energy between the surface and the atmosphere are complicated, involve a number of different processes. We will look at these more carefully later.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Additional points:
#
# - Notice that this is a budget of energy, not temperature.
# - We will need to discuss the connection between the two
# - **Clouds** affect both longwave and shortwave sides of the budget.
# - **WATER** is involved in many of the terms:
#
# - evaporation
# - latent heating (equal and opposite in the global mean)
# - clouds
# - greenhouse effect
# - atmospheric SW absorption
# - surface reflectivity (ice and snow)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Discussion point
#
# How might we expect some of the terms in the global energy budget to vary under anthropogenic climate change?
# + [markdown] slideshow={"slide_type": "slide"}
# ____________
# <a id='section3'></a>
#
# ## 3. Quantifying the planetary energy budget
# ____________
# + [markdown] slideshow={"slide_type": "slide"}
# A budget for the **energy content of the global atmosphere-ocean system**:
#
# \begin{align}
# \frac{dE}{dt} &= \text{net energy flux in to system} \\
# &= \text{flux in – flux out}
# \end{align}
#
# where $E$ is the **enthalpy** or **heat content** of the total system.
#
# We will express the budget **per unit surface area**, so each term above has units W m$^{-2}$
#
# Note: any **internal exchanges** of energy between different reservoirs (e.g. between ocean, land, ice, atmosphere) do not appear in this budget – because $E$ is the **sum of all reservoirs**.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Assumption:
#
# **The only quantitatively important energy sources to the whole system are radiative fluxes to and from space.**
#
# Let’s model those TOA (top-of-atmosphere) fluxes.
# + [markdown] slideshow={"slide_type": "slide"}
# Flux in is **incoming solar radiation**
# The solar constant is
#
# $$ S_0 = 1365.2 \text{ W m}^{-2} $$
#
# (all values will be consistent with Trenberth and Fasullo figure unless noted otherwise)
#
# This is the flux of energy from the sun incident on a unit area perpendicular to the beam direction.
# + [markdown] slideshow={"slide_type": "slide"}
# The area-weighted global mean incoming solar flux is
#
# $$ Q = S_0 \frac{A_{cross-section}}{A_{surface}} $$
#
# [ draw sketch of sphere and illuminated disk ]
#
# where
#
# - $A_{cross-section}$ = area of the illuminated disk = $\pi a^2$
# - $A_{surface}$ = surface area of sphere = $4 \pi a^2$
# - $a$ = radius of Earth
#
# So flux in is $Q = S_0 / 4 = 341.3$ W m$^{-2}$
# + [markdown] slideshow={"slide_type": "slide"}
# Flux out has two parts:
#
# - Reflected solar radiation
# - Emitted terrestrial (longwave) radiation
#
# Introduce terminology / notation:
#
# **OLR = outgoing longwave radiation = terrestrial emissions to space**
# + [markdown] slideshow={"slide_type": "slide"}
# Define the **planetary albedo**:
#
# - $\alpha$ = reflected solar flux / incoming solar flux
# - Or reflected flux = $\alpha Q$ = 101.9 W m$^{-2}$ from data
# - So from data, $\alpha \approx 0.3$
# + [markdown] slideshow={"slide_type": "slide"}
# Define **ASR = absorbed solar radiation**
# \begin{align}
# ASR &= \text{ incoming flux – reflected flux} \\
# &= Q - \alpha Q \\
# &= (1-\alpha) Q
# \end{align}
# + [markdown] slideshow={"slide_type": "slide"}
# Our energy budget then says
#
# $$ \frac{dE}{dt} = (1-\alpha) Q - OLR $$
#
# Note: **This is a generically true statement.** We have just defined some terms, and made the [very good] assumption that the only significant energy sources are radiative exchanges with space.
# + [markdown] slideshow={"slide_type": "slide"}
# **This equation is the starting point for EVERY CLIMATE MODEL.**
#
# But so far, we don’t actually have a MODEL. We just have a statement of a budget. To use this budget to make a model, we need to relate terms in the budget to state variables of the atmosphere-ocean system.
#
# For now, the state variable we are most interested in is **temperature** – because it is directly connected to the physics of each term above.
#
# + [markdown] slideshow={"slide_type": "slide"}
# ____________
# <a id='section4'></a>
#
# ## 4. Using Python to compute emission to space
# ____________
# -
# *Most of what follows is intended as a "fill in the blanks" exercise. We will practice writing some Python code while discussing the physical process of longwave emission to space.*
# Suppose the Earth behaves like a **blackbody radiator** with effective global mean **emission temperature $T_e$**.
#
# Then
#
# $$ OLR = \sigma T_e^4 $$
#
# where OLR = "Outgoing Longwave Radiation", and $\sigma = 5.67 \times 10{-8}$ W m$^{-2}$ K$^{-4}$ the Stefan-Boltzmann constant
#
# **We can just take this as a definition of the emission temperature.**
# Looking back at the observations, the global, annual mean value for OLR is 238.5 W m$^{-2}$.
# ### Calculate the emission temperature $T_e$
# Rerranging the Stefan-Boltzmann law we get
#
# $$ T_e = \left(\frac{\text{OLR}}{\sigma} \right)^{\frac{1}{4}} $$
# First just use Python like a hand calculator to calculate $T_e$ iteractively:
OLR=238.5 # W/m23K4
sigma=5.67e-8 # W/m2K4
Te=(OLR/sigma)**(1/4)
print('%.2f K' % Te)
# Try typing a few different ways, with and without whitespace.
# #### Python fact 1
#
# extra spaces are ignored!
# But typing numbers interactively is tedious and error prone. Let's define a variable called `sigma`
# #### Python fact 2
#
# We can define new variables interactively. Variables let us give names to things. Names make our code easy to understand.
# ### Thoughts on emission temperature
#
# What value did we find for the emission temperature $T_e$? How does it compare to the actual global mean surface temperature?
#
# *Is the blackbody radiator a good model for the Earth's emission to space?*
# ### A simple greenhouse model
# The emission to space is lower because of the greenhouse effect, which we will study in detail later.
#
# For now, just introduce a basic concept:
#
# *Only a fraction of the surface emission makes it out to space.*
#
# We will model the OLR as
#
# $$ \text{OLR} = \tau \sigma T_s^4 $$
#
# where $\tau$ is a number we will call the **transmissivity** of the atmosphere.
#
# Let's fit this model to observations:
#
# $$ \tau = \frac{\text{OLR}}{\sigma T_s^4} $$
tau = 238.5 / sigma / 288**4
# Try calculating OLR for a warmer Earth at 292 K:
# +
OLR = 238.5 # W/m2
sigma = 5.67e-8 # W/m2K4
Ts = 288 # K
tau = OLR / sigma / Ts**4 # transmissivity (dimensionless)
#Te=(OLR/sigma)**(1/4)
#print('%.2f K' % Te)
print('tau= %.2f' % tau)
Ts = 292 # K
def OLR(Ts):
OLR = tau * sigma * Ts**4
#print ('for Ts = %.2f K: OLR = %.02f W/m²' % (Ts,OLR))
#return print ('for Ts = %.2f K: OLR = %.02f W/m²' % (Ts,OLR))
return OLR
# -
# Naturally the emission to space is higher. By how much has it increased for this 4 degree warming?
OLR(288)
OLR(292)
# Answer: 13.5 W m$^{-2}$. Okay but this is tedious and prone to error.
# What we really want to do is **define a reusable function**
# Note a few things:
#
# - The colon at the end of the first line indicates that there is more coming.
# - The interpreter automatically indents the code for us (after the colon)
# - The interpreter automatically colors certain key words
# - We need to hit return one more time at the end to finish our function
#
# #### Python fact 3
#
# **Indentations are not ignored! They serve to group together several lines of code.**
#
# we will see plenty of examples of this – in this case, the indentation lets the interpreter know that the code is all part of the function definition.
# #### Python fact 4:
#
# `def` is a keyword that defines a function.
#
# Just like a mathematical function, a Python function takes one or more input arguments, performs some operations on those inputs, and gives back some resulting value.
# ##### Python fact 5:
#
# `return` is a keyword that defines what value will be returned by the function.
# Once a function is defined, we can call it interactively:
print(OLR(288), OLR(292), OLR(292)-OLR(288))
# #### Python fact 6
#
# ** The `#` symbol is used for comments in Python code. **
#
# The interpreter will ignore anything that follows `#` on a line of code.
# #### Python fact 7:
#
# `print` is a function that causes the value of an expression (or a list of expressions) to be printed to the screen.
#
# (Don’t always need it, because by default the interpreter prints the output of the last statement to the screen, as we have seen).
#
# Note also that we defined variables named sigma and epsilon inside our OLR function.
#
# What happens if you try to `print(epsilon)`?
print(epsilon)
# #### Python fact 8:
#
# **Variables defined in functions do not exist outside of that function.**
#
# Try declaring `sigma = 2`, then `print(sigma)`. And try computing `OLR(288)` again. Did anything change?
sigma=2
print(sigma)
OLR(288)
# Note that we didn’t really **need** to define those variables inside the function. We could have written the function in one line.
#
# But sometimes using named variables *makes our code much easier to read and understand!*
# ### Arrays with `numpy`
#
# Now let’s try some array calculations:
# +
import numpy as np
T = np.linspace(230, 300, 10)
print(T[2])
# -
# - We have just created an array object.
# - The `linspace` function creates an array of numbers evenly spaced between the start and end points.
# - The third argument tells Python how many elements we want.
# We will use the `numpy` package all the time. It is the basic workhorse of scientific computing with Python. We can't do much with arrays of numbers.
# Does our `OLR` function work on an array of temperature values?
# +
OLR = 238.5 # W/m2
sigma = 5.67e-8 # W/m2K4
Ts = 288 # K
tau = OLR / sigma / Ts**4 # transmissivity (dimensionless)
def OLR(Ts):
OLR = tau * sigma * Ts**4
#print ('for Ts = %.2f K: OLR = %.02f W/m²' % (Ts,OLR))
return print ('for Ts = %.2f K: OLR = %.02f W/m²' % (Ts,OLR))
#return OLR
# -
# Now let’s assign these values to a new variable.
T = np.linspace(230, 300, 10)
for i in T:
OLR(T[i])
# Now try again to compute `OLR(288)`
#
# What do you get?
size(T)
# #### Python fact 9: Assigning a value to a named variable overwrites whatever was already assigned to that name.
#
# Python is also case sensitive. If we had used `olr` to store the array, there would be no conflict.
# Now let’s re-enter our function. Start typing `def` and then hit the “up arrow” key. What happens?
# +
OLR = 238.5 # W/m2
sigma = 5.67e-8 # W/m2K4
Ts = 288 # K
tau = OLR / sigma / Ts**4 # transmissivity (dimensionless)
#Te=(OLR/sigma)**(1/4)
#print('%.2f K' % Te)
print('tau= %.2f' % tau)
Ts = 292 # K
def OLR(Ts):
OLR = tau * sigma * Ts**4
#print ('for Ts = %.2f K: OLR = %.02f W/m²' % (Ts,OLR))
#return print ('for Ts = %.2f K: OLR = %.02f W/m²' % (Ts,OLR))
return OLR
# -
# The editor gives us lots of useful keyboard shortcuts.
#
# Here it’s looking up the last expression we entered that began with `def`. Saves a lot of time and typing!
#
# Re-enter the function.
def
# What happens if you use the `up arrow` without typing anything first?
# Also, try typing `history`
# This is very handy. The Python console is taking notes for you!
# + [markdown] slideshow={"slide_type": "skip"}
# <div class="alert alert-success">
# [Back to ATM 623 notebook home](../index.ipynb)
# </div>
# + [markdown] slideshow={"slide_type": "skip"}
# ____________
# ## Version information
# ____________
#
# + slideshow={"slide_type": "skip"}
# %load_ext version_information
# %version_information
# + [markdown] slideshow={"slide_type": "slide"}
# ____________
#
# ## Credits
#
# The author of this notebook is [<NAME>](http://www.atmos.albany.edu/facstaff/brose/index.html), University at Albany.
#
# It was developed in support of [ATM 623: Climate Modeling](http://www.atmos.albany.edu/facstaff/brose/classes/ATM623_Spring2015/), a graduate-level course in the [Department of Atmospheric and Envionmental Sciences](http://www.albany.edu/atmos/index.php)
#
# Development of these notes and the [climlab software](https://github.com/brian-rose/climlab) is partially supported by the National Science Foundation under award AGS-1455071 to Brian Rose. Any opinions, findings, conclusions or recommendations expressed here are mine and do not necessarily reflect the views of the National Science Foundation.
# ____________
# -
| Lectures/Lecture01 -- Planetary energy budget.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Install Dependencies
#
# ```bash
# # pip install pytorch=1.6.0 torchvision=0.7.0 cudatoolkit=10.2 -c pytorch
# pip install matplotlib path.py tqdm
# pip install tensorboard tensorboardX
# pip install scipy scikit-image opencv-contrib-python
# ```
# !pip install matplotlib path.py tqdm
# !pip install tensorboard tensorboardX
# !pip install scipy scikit-image ipykernel
# pip install opencv-contrib-python
# # Download Dataset
#
# To evaluate/train our LEAStereo network, you will need to download the required datasets.
#
# * [SceneFlow](https://lmb.informatik.uni-freiburg.de/resources/datasets/SceneFlowDatasets.en.html)
#
# * [KITTI2015](http://www.cvlibs.net/datasets/kitti/eval_scene_flow.php?benchmark=stereo)
#
# * [KITTI2012](http://www.cvlibs.net/datasets/kitti/eval_stereo_flow.php?benchmark=stereo)
#
# * [Middlebury 2014](https://vision.middlebury.edu/stereo/submit3/)
#
# Change the first column path in file `create_link.sh` with your actual dataset location. Then run `create_link.sh` that will create symbolic links to wherever the datasets were downloaded in the `datasets` folder. For Middlebury 2014 dataset, we perform our network on half resolution images.
# TODO : download and link dataset dir
# !sh download_kitti12.sh
# !sh download_kitti15.sh
# # Test Predictions
#
# You can evaluate a trained model using prediction.sh for each dataset, that would help you generate *.png or *.pfm images correspoding to different datasets.
# !sh predict_kitti12.sh
| main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.6.0
# language: julia
# name: julia-1.6
# ---
# # How to train a model from scratch
# When training a model from scratch, using a one-cycle learning rate schedule is a good starting point.
using FastAI
# We'll set up a `Learner` in the usual way. Note that `Models.xresnet18()`, the backbone we're using, doesn't have pretrained weights.
path = datasetpath("imagenette2-160")
data = Datasets.loadfolderdata(path, filterfn=isimagefile, loadfn=(loadfile, parentname))
classes = unique(eachobs(data[2]))
method = BlockMethod(
(Image{2}(), Label(classes)),
(
ProjectiveTransforms((128, 128)),
ImagePreprocessing(),
OneHot()
)
)
learner = methodlearner(method, data, Models.xresnet18(), ToGPU(), Metrics(accuracy), batchsize=8)
# And then call [`fitonecycle!`](#):
fitonecycle!(learner, 10, 0.005)
| notebooks/fitonecycle.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# #### New to Plotly?
# Plotly's Python library is free and open source! [Get started](https://plot.ly/python/getting-started/) by downloading the client and [reading the primer](https://plot.ly/python/getting-started/).
# <br>You can set up Plotly to work in [online](https://plot.ly/python/getting-started/#initialization-for-online-plotting) or [offline](https://plot.ly/python/getting-started/#initialization-for-offline-plotting) mode, or in [jupyter notebooks](https://plot.ly/python/getting-started/#start-plotting-online).
# <br>We also have a quick-reference [cheatsheet](https://images.plot.ly/plotly-documentation/images/python_cheat_sheet.pdf) (new!) to help you get started!
# #### Version Check
# Run `pip install plotly --upgrade` to update your Plotly version
import plotly
plotly.__version__
# #### Mapbox Access Token
#
# To plot on Mapbox maps with Plotly you'll need a Mapbox account and a [Public Mapbox Access Token](https://www.mapbox.com/studio) which you can add to your [Plotly settings](https://plot.ly/settings/mapbox). If you're using a Plotly On-Premise server, please see additional instructions here: https://help.plot.ly/mapbox-atlas/.
# #### Google Maps API
# In order to use the `Google Maps - Directions API`, you need to create an account with Google and get your API key [here](https://developers.google.com/maps/documentation/directions/).
# +
import plotly.plotly as py
from plotly.graph_objs import *
import numpy as np
import requests
import copy
import googlemaps
# add your google maps api key here
my_google_maps_api_key = 'YOUR_API_KEY'
# -
# #### Parse Tesla Locations
# Perform a `GET` request to retrieve the HTML of the [Google Maps Page](https://www.tesla.com/en_CA/findus#/bounds/70,-50,42,-142,d?search=supercharger,&name=Canada) with all Tesla locations, then parse through to collect all the USA, Canada ones and store them in a dictionary. The dictionary is indexed by `address` and each address has a parameter for `postal_code`, `country`, `latitude` and `longitude`. Be Patient, it takes a while.
r = requests.get('https://www.tesla.com/en_CA/findus#/bounds/70,-50,42,-142,d?search=supercharger,&name=Canada')
r_copy = copy.deepcopy(r.text)
# +
supercharger_locations = {}
params_for_locations = ['latitude":"', 'longitude":"']
location_param = 'location_id":"'
while True:
# add address line to the dictionary
index = r_copy.find(location_param)
if index == -1:
break
index += len(location_param)
index_end = index
while r_copy[index_end] != '"':
index_end += 1
address_line_1 = r_copy[index:index_end]
address_line_1 = str(address_line_1)
supercharger_locations[address_line_1] = {}
for param in params_for_locations:
index = r_copy.find(param)
if index == -1:
break
index += len(param)
index_end = index
while r_copy[index_end] != '"':
index_end += 1
supercharger_locations[address_line_1][param[0:-3]] = r_copy[index:index_end]
r_copy = r_copy[index_end:len(r.text)] # slice off the traversed code
all_keys = supercharger_locations.keys()
# -
# #### Table of Locations
# Create a table with a sample of the `supercharger_locations` data.
# +
import plotly.plotly as py
import plotly.figure_factory as ff
data_matrix = [['Location ID', 'Latitude', 'Longitude']]
first_ten_keys = supercharger_locations.keys()[0:10]
for key in first_ten_keys:
row = [key,
supercharger_locations[key]['latitude'],
supercharger_locations[key]['longitude']]
data_matrix.append(row)
table = ff.create_table(data_matrix)
py.iplot(table, filename='supercharger-locations-sample')
# -
# #### Plot the Route
# The server_key should be replaced with your own Google Maps Directions API key.
#
# Be careful! Make sure you are picking a start and end point that can be driven between, eg. both in the United States of America. Otherwise, the Google Maps API cannot comupute directions and will return an empty list.
# +
def plot_route_between_tesla_stations(address_start, address_end, zoom=3, endpt_size=6):
start = (supercharger_locations[address_start]['latitude'], supercharger_locations[address_start]['longitude'])
end = (supercharger_locations[address_end]['latitude'], supercharger_locations[address_end]['longitude'])
#start = address_start
#end = address_end
directions = gmaps.directions(start, end)
steps = []
steps.append(start) # add starting coordinate to trip
for index in range(len(directions[0]['legs'][0]['steps'])):
start_coords = directions[0]['legs'][0]['steps'][index]['start_location']
steps.append((start_coords['lat'], start_coords['lng']))
if index == len(directions[0]['legs'][0]['steps']) - 1:
end_coords = directions[0]['legs'][0]['steps'][index]['end_location']
steps.append((end_coords['lat'], end_coords['lng']))
steps.append(end) # add ending coordinate to trip
mapbox_access_token = "ADD_YOUR_TOKEN_HERE"
data = Data([
Scattermapbox(
lat=[item_x[0] for item_x in steps],
lon=[item_y[1] for item_y in steps],
mode='markers+lines',
marker=Marker(
size=[endpt_size] + [4 for j in range(len(steps) - 2)] + [endpt_size]
),
)
])
layout = Layout(
autosize=True,
hovermode='closest',
mapbox=dict(
accesstoken=mapbox_access_token,
bearing=0,
style='streets',
center=dict(
lat=np.mean([float(step[0]) for step in steps]),
lon=np.mean([float(step[1]) for step in steps]),
),
pitch=0,
zoom=zoom
),
)
fig = dict(data=data, layout=layout)
return fig
server_key = my_google_maps_api_key
gmaps = googlemaps.Client(key=server_key)
address_start = supercharger_locations.keys()[0]
address_end = supercharger_locations.keys()[501]
zoom=12.2
endpt_size=20
fig = plot_route_between_tesla_stations(address_start, address_end, zoom=10.2, endpt_size=20)
py.iplot(fig, filename='tesla-driving-directions-between-superchargers')
# -
# #### Reference
# See http://moderndata.plot.ly/visualize-tesla-supercharging-stations-with-mysql-and-plotly/ to visualize Tesla supercharging stations with MYSQL and https://plot.ly/python/scattermapbox/ for more information on how to plot scatterplots on maps.
# +
from IPython.display import display, HTML
display(HTML('<link href="//fonts.googleapis.com/css?family=Open+Sans:600,400,300,200|Inconsolata|Ubuntu+Mono:400,700" rel="stylesheet" type="text/css" />'))
display(HTML('<link rel="stylesheet" type="text/css" href="http://help.plot.ly/documentation/all_static/css/ipython-notebook-custom.css">'))
# #! pip install git+https://github.com/plotly/publisher.git --upgrade
import publisher
publisher.publish(
'tesla-supercharging-stations.ipynb', 'python/tesla-supercharging-stations/', 'Python Tesla Supercharging Stations | Examples | Plotly',
'How to plot car-travel routes between USA and Canada Telsa Supercharging Stations in Python.',
title = 'Tesla Supercharging Stations | Plotly',
name = 'Tesla Supercharging Stations',
has_thumbnail='true', thumbnail='thumbnail/tesla-stations.jpg',
language='python',
display_as='maps', order=10,
ipynb= '~notebook_demo/124')
# -
| _posts/python/maps/tesla-supercharging-stations/tesla-supercharging-stations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/joywangai/Moringa_Data_Science_Prep11_W3_Independent_Project_2021_08_Joy_Wangai_Repo/blob/main/Moringa_Data_Science_Prep11_W3_Independent_Project_2021_08_Joy_Wangai_PythonNotebook.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="kpRRsPbAiC3J"
# importing the libraries needed
import pandas as pd
from pandas import Series, DataFrame
import csv
import numpy as np
# + colab={"base_uri": "https://localhost:8080/"} id="mWvJe9fckmAn" outputId="9d3ac435-1017-41cf-b37a-5ae48b0b4d67"
# connecting to the google colab drive
from google.colab import drive
drive.mount('/content/drive')
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="mgFOV0WOmFQp" outputId="0a6dc7fa-2a14-4651-ad7b-cbb147433e77"
# upload the csv files to the drive and we read the data from the csv file
with open('cells_geo.csv','r') as f:
cellsgeo_frame1= pd.read_csv(f, sep=';', quotechar='"', encoding='utf-8',index_col=0)
cellsgeo_frame1.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="6s0rW6xHnDn_" outputId="8f29e198-5af6-4ff6-fcd7-3c2085b9cca5"
# upload the csv files to the drive and we read the data from the csv file
with open('Telcom_dataset.csv','r') as f:
telcom_frame1= pd.read_csv(f, encoding='utf-8', index_col=None)
telcom_frame1.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="Y2svU8frDzRx" outputId="207f0f6f-b6f3-4d12-e2bd-e02888aa3c7d"
# we filer out the entries with missing values from each of the datasets
filtered_telcom_frame1 = telcom_frame1[telcom_frame1[['PRODUTC','VALUE','DATETIME','CELL_ON_SITE','DW_A_NUMBER_INT',
'DW_B_NUMBER_INT','COUNTRY_A','COUNTRY_B','CELL_ID','SITE_ID']].notnull().all(1)]
filtered_telcom_frame1.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 300} id="MSVkb4PqHHyI" outputId="149b0fd7-a4b6-45a1-a856-08770863fba8"
# we find the entry with the maximum billing price i.e the 'VALUE' column from each the filtered telcom datasets
filtered_telcom_frame1.describe()
# + colab={"base_uri": "https://localhost:8080/", "height": 81} id="8KDqVz0mHhPQ" outputId="4fdb61db-c153-49f0-b046-8688ecb8eba8"
findmax = telcom_frame1['VALUE']==1860.000000
telcom_frame1[findmax]
# + colab={"base_uri": "https://localhost:8080/", "height": 237} id="BQOCD7j2He5W" outputId="6634722c-5ad1-4463-b12a-dca5500827b3"
# the using the 'SITE_ID' column we trace cities by finding
# the entries that match the 'SITE_CODE' column from
# the cells_geo dataset to find the cities with the highest billings value
maxcity1=cellsgeo_frame1['SITE_CODE']=='f939f3349c'
cellsgeo_frame1[maxcity1]
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="7n-KfbxUtXFf" outputId="2b7ae586-d60d-411d-b0f5-5bd7130444e4"
# upload the csv files to the drive and we read the data from the csv file
with open('Telcom_dataset2.csv','r') as f:
telcom_frame2= pd.read_csv(f, encoding='utf-8', index_col=None)
telcom_frame2.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="BmAGa2xTDvd8" outputId="1aea8b88-a9ac-45e1-c21b-9e4a53b7b8af"
# we filer out the entries with missing values from each of the datasets
filtered_telcom_frame2 = telcom_frame2[telcom_frame2[['PRODUCT','VALUE','DATE_TIME','CELL_ON_SITE','DW_A_NUMBER',
'DW_B_NUMBER','COUNTRY_A','COUNTRY_B','CELL_ID','SITE_ID']].notnull().all(1)]
filtered_telcom_frame2.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 300} id="PlAW0fzbIVU1" outputId="9e20cc7d-87dc-4af0-dd97-68de4f7d04d6"
# we find the entry with the maximum billing price i.e the 'VALUE' column from each the filtered telcom datasets
filtered_telcom_frame2.describe()
# + colab={"base_uri": "https://localhost:8080/", "height": 81} id="UPCGQv37IV4T" outputId="84856186-8abb-41c0-d30c-9d950fcd6500"
findmax2 = telcom_frame2['VALUE']==3380.000000
telcom_frame2[findmax2]
# + colab={"base_uri": "https://localhost:8080/", "height": 237} id="MGFUQq7PJiKv" outputId="5a8b83d9-2045-40ff-f0b0-dce1f623f3c3"
# the using the 'SITE_ID' column we trace cities by finding
# the entries that match the 'SITE_CODE' column from
# the cells_geo dataset to find the cities with the highest billings value
maxcity2=cellsgeo_frame1['SITE_CODE']=='2e828e607c'
cellsgeo_frame1[maxcity2]
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="bEBAVrACtm7c" outputId="3b474100-bdbe-4ac3-96e0-18c5ab9b4a04"
# upload the csv files to the drive and we read the data from the csv file
with open('Telcom_dataset3.csv','r') as f:
telcom_frame3= pd.read_csv(f, encoding='utf-8', index_col=None)
telcom_frame3.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="yN7TQh5buK8T" outputId="391a17bf-1788-4ad9-c57f-e1b99253d479"
# we filer out the entries with missing values from each of the datasets
filtered_telcom_frame3 = telcom_frame3[telcom_frame3[['PRODUCT','VALUE','DATE_TIME','CELL_ON_SITE','DW_A_NUMBER_INT',
'DW_B_NUMBER_INT','COUNTRY_A','COUNTRY_B','CELLID','SIET_ID']].notnull().all(1)]
filtered_telcom_frame3.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 300} id="bTPx5jFZGevn" outputId="6c7f412b-7075-4afe-a947-afacae7af5bc"
# we find the entry with the maximum billing price i.e the 'VALUE' column from each the filtered telcom datasets
filtered_telcom_frame3.describe()
# + colab={"base_uri": "https://localhost:8080/", "height": 81} id="KdvMfkUO7Hkd" outputId="a2b0c150-d122-4f15-cec8-0dd36579d405"
findmax3 = telcom_frame3['VALUE']==6750.000000
telcom_frame3[findmax3]
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="fToNyw2w65BR" outputId="48a21566-4afd-490a-f412-979b3d372df1"
# the using the 'SITE_ID' column we trace cities by finding
# the entries that match the 'SITE_CODE' column from
# the cells_geo dataset to find the cities with the highest billings value
maxcity3=cellsgeo_frame1['SITE_CODE']=='a64fc672f1'
cellsgeo_frame1[maxcity3]
| Moringa_Data_Science_Prep11_W3_Independent_Project_2021_08_Joy_Wangai_PythonNotebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Py3-basic
# language: python
# name: python3
# ---
# # Spatial Entropy of ABA features
import numpy as np
import pandas as pd
from skimage.measure.entropy import shannon_entropy
from morphontogeny.functions.IO import reconstruct_ABA
from morphontogeny.functions.metrics import greycomatrix_3d
import matplotlib.pyplot as plt
# +
# Loading features
SFT_features = np.load('files/SFT_100features.npy')
# List of Entropies
SFT_spatial_entropies = []
# Indices file
indices_file = 'files/mask_indices.npy'
for ii in range(SFT_features.shape[1]):
# Reconstructing features
SFT_feature = reconstruct_ABA(SFT_features[:,ii], indices_file=indices_file, outside_value=np.nan, mirror=False)
# Measuring spatial entropy
SFT_feature_glcms = greycomatrix_3d(SFT_feature, levels=256, directions=[(1,1,1),(2,1,1),(1,2,1),(1,1,2),(2,2,1),(2,1,2),(1,2,2)])
# Adding entropies to the list
SFT_spatial_entropies.append( sum( [shannon_entropy(x) for x in SFT_feature_glcms] ) )
| Notebooks/02_analyses/Fig2_Spatial_Entropy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Data Preparation
#
# This notebook processes the data that the model is trained on, and is called at the beginning of each model.
import numpy as np # linear alg
import pandas as pd # data processing
import os
from glob import glob # file searching
# %matplotlib inline
import matplotlib.pyplot as plt # plotting
# +
# Setting up access to the images and csv
PATH = os.path.abspath('../../data')
SOURCE_IMAGES = os.path.join(PATH, "images")
images = glob(os.path.join(SOURCE_IMAGES, "*.png"))
#xray_labels_df = pd.read_csv(os.path.join(PATH, 'sample_labels.csv'))
xray_labels_df = pd.read_csv(os.path.join(PATH, 'sample_labels.csv'))
# +
all_image_paths = {os.path.basename(x): x for x in images}
image_names = [os.path.basename(x) for x in images]
# Removes rows from dataframe that do not have a corresponding image in the images folder.
xray_labels_df = xray_labels_df[xray_labels_df['Image Index'].isin(image_names)]
# Drop images with multiple findings
xray_labels_df = xray_labels_df[~xray_labels_df['Finding Labels'].str.contains('\|')]
print('Number of finding labels: {}'.format(len(xray_labels_df['Finding Labels'])))
# Add the path to all images to the dataframe
xray_labels_df['path'] = xray_labels_df['Image Index'].map(all_image_paths.get)
#xray_labels_df.sample(3)
xray_labels_df.head()
# +
from itertools import chain
# Replace Finding Labels with no finding to be blank
xray_labels_df['Finding Labels'] = xray_labels_df['Finding Labels'].map(lambda x: x.replace('No Finding', ''))
# Add columns to for each diagnosis to df
all_labels = np.unique(list(chain(*xray_labels_df['Finding Labels'].map(lambda x: x.split('|')).tolist())))
all_labels = [x for x in all_labels if len(x) > 0]
# Adds attributes for each possible finding, and assigns value of 1.0 for each finding
# in the Finding Label (1-Hot Encoding)
for c_label in all_labels:
if len(c_label) > 1:
xray_labels_df[c_label] = xray_labels_df['Finding Labels'].map(lambda finding: 1.0 if c_label in finding else 0)
xray_labels_df.sample(5)
# -
MIN_CASES = 200
all_labels = [c_label for c_label in all_labels if xray_labels_df[c_label].sum() > MIN_CASES]
sample_weights = xray_labels_df['Finding Labels'].map(lambda x: len(x.split('|')) if len(x) > 0 else 0).values + 4e-2
sample_weights /= sample_weights.sum()
xray_labels_df = xray_labels_df.sample(12000, weights = sample_weights)
xray_labels_df['disease_vec'] = xray_labels_df.apply(lambda x: [x[all_labels].values], 1).map(lambda x: x[0])
def flow_from_dataframe(img_data_gen, in_df, path_col, y_col, **dflow_args):
base_dir = os.path.dirname(in_df[path_col].values[0])
df_gen = img_data_gen.flow_from_directory(base_dir, class_mode='sparse', **dflow_args)
df_gen.filenames = in_df[path_col].values
df_gen.classes = np.stack(in_df[y_col].values)
df_gen.samples = in_df.shape[0]
df_gen.n = in_df.shape[0]
df_gen._set_index_array()
df_gen.directory = ''
print('Reinserting dataframe: {} images'.format(in_df.shape[0]))
# df_gen is an iterator that yields a tuple (X,Y).
# X = batch of images, and Y = corresponding labels.
return df_gen
print('Complete')
| models/data_preparation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
np.arange(0,100,2)
np.linspace(0,5,8).reshape(2,4)
np.random.rand(2)
np.random.ranf(4)
np.arange(1,12,2)
arr=np.random.randint(1,100,20)
arr
arr.reshape(5,4)
arr.max()
arr.min()
arr.shape
arr.reshape(2,10)
arr=np.arange(10)
arr
arr[5:]
arr[:5]
arr[1:7]
arr[0:7]
arar=np.arange(10)
arar
slice_of_arr=arar[0:5]
slice_of_arr
slice_of_arr[:]=99
arar
arr_2d=np.array([[2,3,4],[4,15,65],[98,95,14]])
arr_2d
arr_2d[0][1]
arr_2d[2,2]
arr_2d[1:,:2]
arr=np.random.randint(1,99,10)
arr
bool_=arr>30
bool_
arr[bool_]
ara=np.arange(32).reshape(4,8)
ara
ara[1:3,2:4]
arr=np.arange(0,10)
arr
1/arr
np.count_nonzero(arr)
| numpy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# <b>Decision Variable:</b>
#
# \begin{equation}
# \begin{array}{11}
# \ Y_{i,k} & \forall i \in R, k \in WD
# \end{array}
# \end{equation}
# \begin{equation}
# \begin{array}{11}
# \ X_{i,j,k} & \forall i \in R, j \in S, k \in WD
# \end{array}
# \end{equation}
#
# <b>Objective Function:</b>
#
# \begin{equation}
# \begin{array}{ll}
# \text{minimise} & \sum_{i \in R} \sum_{k \in W} Y_{i,k} c_{i,k}\\
# \end{array}
# \end{equation}
#
#
# <b>Subject to:</b>
# \
# <i>Respecting Demand Constraints</i>
#
# \begin{equation}
# \begin{array}{11}
# \sum_{i \in R} \sum_{k \in W} X_{i,j,k} m_{i,j} = d_j & \forall j \in S
# \end{array}
# \end{equation}
#
# \
# <i>Respecting Delivery Acceptance Constraints</i>
#
# \begin{equation}
# \begin{array}{11}
# \sum_{i \in R} X_{i,j,k} m_{i,j} \le a_j & \forall j \in S, k \in W
# \end{array}
# \end{equation}
#
# \
# <i>Respecting Truck Constraints</i>
#
# \begin{equation}
# \begin{array}{11}
# \sum_{j \in S} X_{i,j,k}m_{i,j} \le C_iY_{i,k} & \forall i \in R, k \in W
# \end{array}
# \end{equation}
#
#
#
# \
# <b>General Representations</b>
#
# \
# <i>Sets</i>
# * <i>R</i>: Set of routes
# * <i>S</i>: Set of stores
# * <i>W</i>: Set of days in a week
#
# \
# <i>Parameters</i>
# \
# $
# \begin{equation}
# \begin{array}{11}
# d_j & \text{: Weekly demand of store j,} & j \in S
# \end{array}
# \end{equation}
# $
#
# $
# \begin{equation}
# \begin{array}{11}
# a_j & \text{: Delivery acceptance limits for store j,} & j \in S
# \end{array}
# \end{equation}
# $
#
# $
# \begin{equation}
# \begin{array}{11}
# m_{i,j} & \text{: Dummy variable to indicate 1 if store j is along route i and 0 otherwise,} & i \in R, j \in S
# \end{array}
# \end{equation}
# $
#
# $
# \begin{equation}
# \begin{array}{11}
# c_{i,k} & \text{: Cost of truck on each route i on day k }
# \end{array}
# \end{equation}
# $
#
# $
# \begin{equation}
# \begin{array}{11}
# C_{i} & \text{: Full capacity of truck on route i on each day }
# \end{array}
# \end{equation}
# $
#
# \
# <i>Variables</i>
# $
# \begin{equation}
# \begin{array}{11}
# X_{i,j,k} & \text{: Units on each route i to be delivered to a store j on day k} \\
# Y_{i,k} & \text{: If route i is used on day k}
# \end{array}
# \end{equation}
# $
#
# \
# <i>Entities</i>
# * <i>i</i>: Route in R
# * <i>j</i>: Store in S
# * <i>k</i>: Day in W
#
# +
#Importing Packages
from gurobipy import *
from math import sqrt
import pandas as pd
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
import xlrd
# +
#Importing Data
routes = pd.read_excel("routes.xlsx").to_numpy()
print(routes)
demand = pd.read_excel("data-demands-deliveyconstraint.xlsx").to_numpy()
print(demand)
# +
#Demand
d = demand[:,1]
#Delivery constraints
dc = demand[:,2]
#Route matrix
#Store names
snames = demand[:,0]
count = 1
mat = np.zeros(24)
for i in snames:
routestore1 = (routes[:,1] == str(i)) * 1
routestore2 = (routes[:,2] == str(i)) * 1
mainroutestore = routestore1 + routestore2
mat = np.row_stack((mat, mainroutestore))
#mat = np.concatenate((mat, mainroutestore),axis=1)
mat = np.array(mat, dtype='int16')[1:,:].transpose((1,0))
print(mat)
# +
#Shipping costs
c = np.full((24, 18), 100)
c
#Supply constraints (truck capacity)
t = np.full((24,7), 24)
t
# +
#Transportation Problem
m4 = Model('transportation')
#Variables
#Variables are in proportion
var = m4.addVars(24,18,7)
yvar = m4.addVars(24,7,vtype = GRB.BINARY)
#Objective
m4.setObjective(sum(yvar[i,k] for i in range(24) for k in range(7)), GRB.MINIMIZE)
#Weekly Demand
for j in range(18):
m4.addConstr(sum(var[i,j,k]*mat[i,j] for i in range(24) for k in range(7)) == d[j])
#Delivery Constraints
for j in range(18):
for k in range(7):
m4.addConstr(sum(var[i,j,k]*mat[i,j] for i in range(24)) <= 0.6*dc[j])
#Supply constraint
for i in range(24):
for k in range(7):
m4.addConstr(sum(var[i,j,k]*mat[i,j] for j in range(18)) <= t[i,k]*yvar[i,k])
#Solving the optimization problem
m4.optimize()
#Printing the optimal solutions obtained
print("Optimal Solutions:")
for i, val in var.items():
if val.getAttr("x") != 0:
print("Number of units from route %g to store %g on day %g:\t %g " %(i[0]+1, i[1]+1, i[2]+1, val.getAttr("x")))
#Printing y
for i, val in yvar.items():
print("Run route %g on day %g:\t %g " %(i[0]+1, i[1]+1, val.getAttr("x")))
print(yvar)
# -
for i in range(24):
for k in range(7):
print(yvar[i,k].getAttr("x"))
| IRP Formulation/Algo2-4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + id="_6k7gKclKVWt"
from google.colab import drive
drive.mount('/gdrive')
# %cd /gdrive/My\ Drive/PytorchDigitalPathology/google-collab-version/PytorchDigitalPathology/classification_lymphoma_densenet/data/
# + id="ak50KC9yLa4T"
#v3.classification
#28/11/2018
dataname="lymphoma"
patch_size=256 #size of the tiles to extract and save in the database, must be >= to training size
stride_size=256 #distance to skip between patches, 1 indicated pixel wise extraction, patch_size would result in non-overlapping tiles
mirror_pad_size=128 # number of pixels to pad *after* resize to image with by mirroring (edge's of patches tend not to be analyzed well, so padding allows them to appear more centered in the patch)
test_set_size=.1 # what percentage of the dataset should be used as a held out validation/testing set
resize=1 #resize input images
class_names=["CLL", "FL", "MCL"]#what classes we expect to have in the data, here we have only 2 classes but we could add additional classes
#-----Note---
#One should likely make sure that (nrow+mirror_pad_size) mod patch_size == 0, where nrow is the number of rows after resizing
#so that no pixels are lost (any remainer is ignored)
# + id="YwrFs146Ln3B"
import torch
import tables
import os,sys
import glob
import PIL
import numpy as np
import cv2
import matplotlib.pyplot as plt
from sklearn import model_selection
import sklearn.feature_extraction.image
import random
seed = random.randrange(sys.maxsize) #get a random seed so that we can reproducibly do the cross validation setup
random.seed(seed) # set the seed
print(f"random seed (note down for reproducibility): {seed}")
# + id="OayL3LZrL5Ar"
img_dtype = tables.UInt8Atom() # dtype in which the images will be saved, this indicates that images will be saved as unsigned int 8 bit, i.e., [0,255]
filenameAtom = tables.StringAtom(itemsize=255) #create an atom to store the filename of the image, just incase we need it later,
# + id="41KKjaEpL8P4"
files=glob.glob('**/*.tif') # create a list of the files, in this case we're only interested in files which have masks so we can use supervised learning
#create training and validation stages and split the files appropriately between them
phases={}
phases["train"],phases["val"]=next(iter(model_selection.ShuffleSplit(n_splits=1,test_size=test_set_size).split(files)))
# + id="mTlHayS4NZ4m"
#--subset for rapid testing
phases["train"]=phases["train"][0:100]
phases["val"]=phases["val"][0:20]
# + id="SqRBAn0DfFnt"
#Copying the PS_scikitlearn.py to the colab notebook runtime. This has the extract_patches function which is now a private method.
# !cp '/gdrive/My Drive/PytorchDigitalPathology/google-collab-version/PytorchDigitalPathology/classification_lymphoma_densenet/PS_scikitlearn.py' /content/PS_scikitlearn.py
import sys
sys.path.append('/content')
from PS_scikitlearn import extract_patches
# + id="JaygcICbNs-i"
storage={} #holder for future pytables
block_shape=np.array((patch_size,patch_size,3)) #block shape specifies what we'll be saving into the pytable array, here we assume that masks are 1d and images are 3d
filters=tables.Filters(complevel=6, complib='zlib') #we can also specify filters, such as compression, to improve storage speed
for phase in phases.keys(): #now for each of the phases, we'll loop through the files
print(phase)
totals=np.zeros(len(class_names)) # we can to keep counts of all the classes in for in particular training, since we
hdf5_file = tables.open_file(f"./{dataname}_{phase}.pytable", mode='w') #open the respective pytable
storage["filenames"] = hdf5_file.create_earray(hdf5_file.root, 'filenames', filenameAtom, (0,)) #create the array for storage
storage["imgs"]= hdf5_file.create_earray(hdf5_file.root, "imgs", img_dtype,
shape=np.append([0],block_shape),
chunkshape=np.append([1],block_shape),
filters=filters)
storage["labels"]= hdf5_file.create_earray(hdf5_file.root, "labels", img_dtype,
shape=[0],
chunkshape=[1],
filters=filters)
for filei in phases[phase]: #now for each of the files
fname=files[filei]
print(fname)
classid=[idx for idx in range(len(class_names)) if class_names[idx] in fname][0]
totals[classid]+=1
io=cv2.cvtColor(cv2.imread(fname),cv2.COLOR_BGR2RGB)
interp_method=PIL.Image.BICUBIC
io = cv2.resize(io,(0,0),fx=resize,fy=resize, interpolation=interp_method) #resize it as specified above
io = np.pad(io, [(mirror_pad_size, mirror_pad_size), (mirror_pad_size, mirror_pad_size), (0, 0)], mode="reflect")
#convert input image into overlapping tiles, size is ntiler x ntilec x 1 x patch_size x patch_size x3
io_arr_out=extract_patches(io,(patch_size,patch_size,3),stride_size)
#resize it into a ntile x patch_size x patch_size x 3
io_arr_out=io_arr_out.reshape(-1,patch_size,patch_size,3)
storage["imgs"].append(io_arr_out)
storage["labels"].append([classid for x in range(io_arr_out.shape[0])]) #add the filename to the storage array
storage["filenames"].append([fname for x in range(io_arr_out.shape[0])]) #add the filename to the storage array
#lastely, we should store the number of pixels
npixels=hdf5_file.create_carray(hdf5_file.root, 'classsizes', tables.Atom.from_dtype(totals.dtype), totals.shape)
npixels[:]=totals
hdf5_file.close()
| google-collab-version/PytorchDigitalPathology/classification_lymphoma_densenet/make_hdf5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Week 2. Dimensionality Reduction
# ---
# <img src = "https://i.gyazo.com/013d3e74948fb79a644fd2dae7ae5fbd.png" width = "500px">
#
# ### A note about neural networks
#
# <img src = "https://i.gyazo.com/538b9fc73cdcc6384086011c0e44a914.png" width = "500px">
# ## Curse of Dimensionality
#
# ---
# Many ML methods use the distance measure
# - k-Nearest Neighbours
# - Support Vector Machnies
# - Recommendation systems
#
# ### Why is high-dimensional data a problem?
# - More dimensiones, more features
# - Risk of overfitting our models
# - Distances grow more and more alike
# - No clear distinction between clustered objects
# - Concentration phenomenon for Euclidean distance
#
# "As we add more dimensiones we also increase the processing power we need to train the model and make predictions, as well as the amount of training data required"
# - <NAME>
#
# ### Why are more features bad?
# - Redundant / irrelevant features
# - More noise added than signal
# - Hard to interpret and visualize
# - Hard to store and process data
#
# <img src = "https://i.gyazo.com/2c19dacc0216c5e002e5fcf857854cb9.png" width = "500px">
# (with a constant number of training examples)
#
# $$d_{ij} = \sqrt{ \sum\limits_{k=1}^{n}({x_{ik} - x_{jk}})^2}$$
#
# - Adding dimensions increases feature space volume
# - New dimensions **add non-negative terms to the sum**
# - Distance increases with the number of dimensions
# - For a given number of examples, the feature space becomes **increasingly sparse**
# - Higher chance of overfitting on errors
#
# ### The lower the hyphothesis space
# - The easier it is to find the correct hyphotesis
# - The less examples you need
#
# ## How dimensionality impacts in other ways
# ---
#
# - Runtime and system memory requirements
# - Solutions take longer to reach global
# - More dimensions raise the likelihood of correlated features
#
# ### More features require more training data
# - More features aren't better if they don't add predictive information
# - Number of training instances needed increases exponentially with each added feature
# - Reduces real-world usefulness of models
#
# <img src = "https://i.gyazo.com/6d04c54a77e51a0d8ebeb9669fade702.png" width = "500px">
#
# ## Manual Dimensionality Reductions
# ---
#
# - Features must have information to produce correct results
# - Derive features from inherent features
# - Extract and recombine to create new features
#
# ### Why reduce dimensionality?
# - Storage
# - Computational
# - Consistency
# - Visualization
#
# Major techniques for dimensionality reduction
# - Engineering
# - Selection
#
# ### Feature Engineering
# <img src = "https://i.gyazo.com/98caef55016e47a906181124dd7ea9a0.png" width = "500px">
#
#
# ### Linear dimensionality reduction
#
# - Linearly project $n$-dimensional data onto a $k$-dimensional subspace ($k < n$, often $k << n$).
# - There are infinitely many $k$-dimensional subspaces we can project the data onto
#
# ### Best k-dimensional subspace for projection
#
# <img src = "https://i.gyazo.com/3a5d4db25efac36d1ae486bc295ed6bb.png" width = "500px">
#
# ## Principal Components Analysis
# ---
#
# - PCA is minimization of the orthogonal distance
# - Widely used method for unsupervised & linear dimensionality reduction
# - Accounts for variance of data in as few dimensions as possible using linear projections
# - PCs maximize the variance of projections
# - PCs are orthogonal
# - Gives the best axis to project
# - Goal of PCA: Minimize total squared reconstruction error
#
#
# To achieve the latter, we can follow the following steps:
# 1. Find a line, such that when the data is projected onto that line, it has the maximum variance
# 2. Find a second line, orthogonal to the first, that has maximum projected variance
# 3. Repeat until we have $k$ orthogonal lines
#
# <img src = "https://i.gyazo.com/07380f6f63d00f97041269d6ad09c465.png" width = "500px">
#
# ## Other Techniques
# ---
#
# <img src = "https://i.gyazo.com/1c0ec071e564848dff5509b12e9213a4.png" width = "500px">
#
# ### SVD
#
# - SVD decomposes non-square matrices
# - Useful for sparse matrices as produced by TF-IDF
# - Removes redundant features from dataset
#
# ### Independente Component Analysis (ICA)
#
# - PCA seeks directions in feature space that minimize reconstruction error
# - ICA seeks diretions that are most statistically independent
# - ICA addresses higher order dependence
# - **Goal of ICA:** recover original signals, $S(t)$ from $Y(t)$
# - It assumes there exists independent signals $S$ and a linear combination of signals $Y$
# - The goal is to recover the original signals of $S$ from $Y$
#
#
# <img src = "https://i.gyazo.com/7f2324389602e78fcec9bc6291a7f518.png">
#
# - Both are statistical transformations, that is PCA uses information extracted from second order statistics, while ICA goes up to higher order statistics. Both are used in various fields like blind source separation, feature extraction and also in neuroscience. ICA is an algorithm that finds directions in the feature space corresponding to projections which are highly non-Gaussian.. Unlike PCA, these directions need not be orthogonal in the original feature space, but they are orthogonal in the whitened feature space, in which all directions correspond to the same variance. PCA, on the other hand, finds orthogonal directions in the raw feature space that corresponded directions accounting for maximum variance.
#
# - Then, let's represent the signal in the PCA space after whitening by the variance corresponding to the PCA vectors. Running ICA corresponds to finding a rotation in this space to identify the directions which are the most non-Gaussian. This is in the lower right, from this figure, you can see that PCA removes correlations but not higher order dependence. On the contrary, ICA removes correlations along with higher order dependence. When it comes to the importance of components, PCA, considers some of them to be more important than others. ICA, on the other hand, considers all components to be equally important.
#
# <img src = "https://i.gyazo.com/4d2cdae96b9dfec590a4680fd8306d58.png" width = "500px">
#
# - Now, let's discuss a dimensionality reduction technique called non-negative Matrix Factorization or NMF. NMF expresses samples as a combination of interpretable parts. For example, it represents documents as combinations of topics, and images in terms of commonly occurring visual patterns. NMF, like PCA, is a dimensionality reduction technique, in contrast to PCA, however, NMF models are interpretable. This means NMF models are easier to understand and much easier for us to explain to others. NMF can't be applied to every dataset however, it requires the sample features to be non-negative, so the values must be greater than or equal to zero. It has been observed that when carefully constrained, NMF can produce a parts-based representation of the dataset, resulting in interpretable models.
# #### Algorithmic process
# ---
#
# 1. Compute columns mean $\bar{x}$
# 2. Substract mean $B = X - \bar{X}$ (centered data).
# 3. Covariance matrix of colums of $B$: $C = B^TB$
# 4. Compute eigenvalues of $C$: $CV = VD$
# 5. Finally, this principal components are $T = BV$ where $V$ is called de loadings.
#
# - The latter process can be view as decomposing $B$ in directions of maximal variance.
# - An alternative to compute the principal components is to use Singular Value Decomposition (for short, SVD):
#
# $$B = U \Sigma V^T$$
#
# As a matter of fact, when you use <code>sklearn.decomposition.PCA</code> class, <code>PCA.components_</code> return $V^T$. Addionality, when you use <code>PCA.fit_transform</code> you get back $T = U \Sigma = BV$ which are the principal components. The latter is actually a projection of the data on the components given by $V^T$ matrix.
# #### Sample code.
#
# ---
import numpy as np
X = np.random.rand(100,8) * 100
X_test = np.random.rand(20, 8) * 100
mu_X = np.mean(X, axis = 0)
mu_X
centered_X = X - mu_X
#centered_X
u, s, v = np.linalg.svd(centered_X, full_matrices = False)
# +
#u, s, v
# -
max_abs_cols = np.argmax(np.abs(u), axis=0)
max_abs_cols
signs = np.sign(u[max_abs_cols, range(u.shape[1])])
signs
u *= signs
u
v *= signs
v
n_components = 3
# Principal components (this is the projected data, e.g T = U * Sigma)
principal_components = (u * s)[:, : n_components]
principal_components
X_test.shape, (u * s)[:, : n_components].shape
X_test_center = X_test - mu_X
# This should be the same as pca.transform(X_test) output.
np.dot(X_test_center, v.T)
# #### Sklearn
#
# ---
from sklearn.decomposition import PCA
pca = PCA(n_components = 3)
pca.fit(X)
pca.explained_variance_ratio_
pca.explained_variance_ratio_.sum()
pca.components_
# Principal components (this returns U * Sigma)
res = pca.fit_transform(X)
#res
# X_test is projected on the first principal components previosly extracted from a training set.
# e.g X_transformed = np.dot(X_test, self.components_.T), where self.components_ = V^T
# We project point on components.
projection = pca.transform(X_test)
#projection
pca.components_
X_test
xs = projection[:, 0]
ys = projection[:, 1]
xs
plt.scatter(xs, ys)
xs = projection[:, 0]
ys = projection[:, 1]
zs = projection[:, 2]
# +
# Import libraries
from mpl_toolkits import mplot3d
import numpy as np
import matplotlib.pyplot as plt
# Creating figure
fig = plt.figure(figsize = (16, 9))
ax = plt.axes(projection ="3d")
# Creating color map
my_cmap = plt.get_cmap('hsv')
# Creating plot
sctt = ax.scatter3D(xs, ys, zs,
alpha = 1,
s = 100,
marker ='*')
plt.title("simple 3D scatter plot")
ax.set_xlabel('X-axis', fontweight ='bold')
ax.set_ylabel('Y-axis', fontweight ='bold')
ax.set_zlabel('Z-axis', fontweight ='bold')
fig.colorbar(sctt, ax = ax, shrink = 0.5, aspect = 5)
# show plot
plt.show()
# -
import numpy as np
X = np.random.rand(3,3) * 100
X
basis = np.array([[1, 0, 0], [0, 1, 0], [0, 0, 1]])
basis
from sklearn.decomposition import PCA
reducer = PCA()
res = reducer.fit_transform(X)
reducer.components_
res
# +
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
import numpy as np
soa = np.array([
np.array([0, 0, 0, 46.08, 18.85, 23.884])/20,
[0, 0, 0, -0.5433, 0.539, 0.643],
[0, 0, 0, 0.692, -0.14622, 0.7068],
[0, 0, 0, -0.475, -0.829, 0.2936],
[0, 0, 0, 4.433, -1.7938, 0], # Prácticamente la data quedó representada en sólo 2 dimensiones!
])
c = ("red", "green", "green", "green", "blue")
X, Y, Z, U, V, W = zip(*soa)
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
ax.quiver(X, Y, Z, U, V, W, colors = c)
ax.set_xlim([-5, 5])
ax.set_ylim([-5, 5])
ax.set_zlim([0, 1])
plt.show()
# -
| Course 3. ML Modeling in Pipelines in Production/2.1 Dimensionality Reduction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # FIRE 101 Lesson Plan
# ## Financial Independence Retire Early
# Execute this cell to load the notebook's style sheet, then ignore it
from IPython.core.display import HTML
css_file = '../style/custom.css'
HTML(open(css_file, "r").read())
import pandas as pd
import numpy as np
from ipywidgets import interact, interactive, fixed, interact_manual, Layout
import ipywidgets as widgets
import threading
from IPython.display import YouTubeVideo, display
# +
#future self functions
savings = .0000001
money_market = .01
cd = .02
stocks = .07
def calc_simple_compound_interest(i, r, t, one_time):
if one_time is True:
return i*(1+r)**t
else:
return i * (1+r) ** t + i * (1+r) * ((1+r) ** t - 1) / r
def get_compound_simple_df(i, one_time):
start_age = 20
stop_age = 60
yi = i*365
df = pd.DataFrame({'Age': pd.Series(range(start_age,stop_age))})
df['Mattress 0%'] = df.apply(lambda x: calc_simple_compound_interest(yi, savings, x['Age']-start_age, one_time), axis=1)
df['Money Market'] = df.apply(lambda x: calc_simple_compound_interest(yi, money_market, x['Age']-start_age, one_time), axis=1)
df['CD 2%'] = df.apply(lambda x: calc_simple_compound_interest(yi, cd, x['Age']-start_age, one_time), axis=1)
df['Stocks 7%'] = df.apply(lambda x: calc_simple_compound_interest(yi, stocks, x['Age']-start_age, one_time), axis=1)
return df
style = {'description_width': 'initial'}
fs_investment = widgets.IntSlider(
value=1,
min=1,
max=10,
step=1,
description='Treat your future self:',
disabled=False,
continuous_update=False,
orientation='horizontal',
layout=Layout(width='50%'),
style=style,
readout=True,
readout_format='d'
)
#fs_one_time = widgets.Checkbox(
# value=True,
# description='One Year Investment (as opposed to continuing every year)',
# layout=Layout(width='100%'),
# disabled=False,
# indent=False
#)
fs_play = widgets.Play(
value=1,
min=1,
max=10,
step=1,
description="Press play",
interval=1000,
#speed=0,
disabled=False
)
def fs_output(i):
#create and display dataframe
df_plot = get_compound_simple_df(i, True)
ax = df_plot.plot.line(x='Age', y=['Mattress 0%', 'CD 2%', 'Stocks 7%'], figsize=(12,10), linewidth=3, grid=True, ylim=(0,20000))
ax.ticklabel_format(style='plain')
#display(df_plot)
return("compare rate returns example")
fs_rn_investment = widgets.IntSlider(
value=1,
min=1,
max=20,
step=1,
description='Treat your future self:',
disabled=False,
continuous_update=False,
orientation='horizontal',
layout=Layout(width='50%'),
style=style,
readout=True,
readout_format='d'
)
fs_rn_play = widgets.Play(
value=1,
min=1,
max=20,
step=1,
description="Press play",
interval=1000,
#speed=0,
disabled=False
)
def fs_rn_output(i):
#create and display dataframe
df_plot = get_compound_simple_df(i, False)
ax = df_plot.plot.line(x='Age', y=['Mattress 0%', 'CD 2%', 'Stocks 7%'], figsize=(12,10), linewidth=3, grid=True, ylim=(0,2000000))
ax.ticklabel_format(style='plain')
#display(df_plot)
return("compare rate returns example")
# +
#retirement functions
#def calc_stash_compound_interest(i, r, t):
# return i * (1+r) ** t + i * (1+r) * ((1+r) ** t - 1) / r
def calc_stash_compound_interest(w, i, r, t):
return w + (i * (1+r) ** t + i * (1+r) * ((1+r) ** t - 1) / r)
def calc_principal(w, i, t):
return w + (i * t)
def get_retirement_df(rate, age, salary, spending, net_worth):
df = pd.DataFrame({'Age': pd.Series(range(age,age+40))})
#df['4 Percent'] = df.apply(lambda x: calc_simple_compound_interest(yi, savings, x['Age']-start_age, one_time), axis=1)
df['Yearly Income'] = df.apply(lambda x: salary, axis=1)
df['Yearly Spending'] = df.apply(lambda x: spending, axis=1)
df['Yearly Savings'] = df.apply(lambda x: salary-spending, axis=1)
#df['Net Worth'] = df.apply(lambda x: net_worth, axis=1)
df['Total Stash'] = df.apply(lambda x: calc_stash_compound_interest(net_worth, salary-spending, rate*.01, x['Age']-age), axis=1)
df['Principal'] = df.apply(lambda x: calc_principal(net_worth, salary-spending, x['Age']-age), axis=1)
#df['Total Stash'] = df['Total Stash'].apply(lambda x: "${:.0f}k".format((x/1000)))
#df['Principal'] = df['Principal'].apply(lambda x: "${:.0f}k".format((x/1000)))
df['Total Stash'] = df['Total Stash'].astype('int64')
df['Principal'] = df['Principal'].astype('int64')
return df
style = {'description_width': 'initial'}
age = widgets.IntText(
value=20,
placeholder=20,
description='Age:',
readout_format='d',
disabled=False
)
rate = widgets.IntText(
value=4,
placeholder=4,
description='Rate:',
readout_format='d',
disabled=False
)
salary = widgets.IntText(
value=80000,
placeholder=80000,
description='Salary:',
readout_format='d',
disabled=False
)
spending = widgets.IntText(
value=30000,
placeholder=30000,
description='Spending:',
readout_format='d',
disabled=False
)
net_worth = widgets.IntText(
value=20000,
placeholder=20000,
description='Net Worth:',
readout_format='d',
disabled=False
)
def r_output(r, a, sal, sp, w):
#create and display dataframe
df_plot = get_retirement_df(r, a, sal, sp, w)
#ax = df.plot()
#df_plot.plot.line(x='Age', y=['Principal', 'Total Stash'], figsize=(12,10), linewidth=3, grid=True)
ax = df_plot.plot.line(x='Age', y=['Principal', 'Total Stash'], figsize=(12,10), linewidth=3, grid=True)
ax.ticklabel_format(style='plain')
#print(df_plot['Total Stash'].max())
#display(df_plot)
return("compare rate returns example")
# +
#compound interest magic functions
def calc_compound_interest(x, y, i, r, n, start_age, start_inv_age, years_investing):
if x > 0:
x = x*(y-(start_inv_age - start_age))
#x = x*(y-years_investing)
else:
if y >= start_inv_age - start_age:
x = i*(years_investing)
else:
return 0
return x*(1 + r/n)**((y+start_age)-start_inv_age)
def no_investment(x, y, start_age, start_inv_age):
if x > 0:
return x**((y+start_age)-start_inv_age)
else:
return 0
def future_value(x, y, r, n, t, pmt):
return pmt * (((1 + r/n)**(n*t) - 1) / (r/n))
def get_compound_dual_df(i, r, a1, a2, y1, y2):
#start_age = 20
#stop_age = 70
start_age = a1
stop_age = start_age + y2 + 10
n = 1
#t = 1
#pmt = 2000/12
df = pd.DataFrame({'Age': pd.Series(range(start_age,stop_age))})
#df['years'] = df.index
df['Fire Investor'] = np.where(np.logical_and(df['Age']>=a1, df['Age']<a1+y1), i, 0)
df['OG Investor'] = np.where(np.logical_and(df['Age']>=a2, df['Age']<a2+y2), i, 0)
df['Matt'] = np.where(np.logical_and(df['Age']>=a1, df['Age']<stop_age), i, 0)
df['Your Money'] = df.apply(lambda x: calc_compound_interest(x['Fire Investor'], x.name, i, r, n, start_age, a1, y1), axis=1)
#pd.set_option('display.float_format', lambda x: '%.2f' % x)
df['Friends Money'] = df.apply(lambda x: calc_compound_interest(x['OG Investor'], x.name, i, r, n, start_age, a2, y2), axis=1)
df['Matts Money'] = df.apply(lambda x: calc_compound_interest(x['Matt'], x.name, i, 0, n, start_age, a1, stop_age), axis=1)
return df
style = {'description_width': 'initial'}
investment = widgets.IntSlider(
value=1000,
min=100,
max=2000,
step=100,
description='investment amount:',
disabled=False,
continuous_update=False,
orientation='horizontal',
layout=Layout(width='50%'),
style=style,
readout=True,
readout_format='d'
)
irate = widgets.FloatSlider(
value=0.07,
min=0.01,
max=0.2,
step=0.01,
description='interest rate:',
disabled=False,
continuous_update=False,
orientation='horizontal',
layout=Layout(width='50%'),
style=style,
readout=True,
readout_format='.2f'
)
your_investing_start_age = widgets.IntSlider(
value=20,
min=10,
max=65,
step=1,
description='age you start investing:',
disabled=False,
continuous_update=False,
orientation='horizontal',
layout=Layout(width='50%'),
style=style,
readout=True,
readout_format='d'
)
friend_investing_start_age = widgets.IntSlider(
value=30,
min=10,
max=65,
step=1,
description='age friend starts investing:',
disabled=False,
continuous_update=False,
orientation='horizontal',
layout=Layout(width='50%'),
style=style,
readout=True,
readout_format='d'
)
years_you_invest = widgets.IntSlider(
value=10,
min=1,
max=50,
step=1,
description='years you contribute money:',
disabled=False,
continuous_update=False,
orientation='horizontal',
layout=Layout(width='50%'),
style=style,
readout=True,
readout_format='d'
)
years_friend_invests = widgets.IntSlider(
value=20,
min=1,
max=50,
step=1,
description='years friend contributes money:',
disabled=False,
continuous_update=False,
orientation='horizontal',
layout=Layout(width='50%'),
style=style,
readout=True,
readout_format='d'
)
play = widgets.Play(
value=0,
min=0,
max=104,
step=1,
description="Press play",
speed=100,
disabled=False
)
def output(i, r, a1, a2, y1, y2):
#create and display dataframe
df_plot = get_compound_dual_df(i, r, a1, a2, y1, y2)
ax = df_plot.plot.line(x='Age', y=['Your Money', 'Friends Money', 'Matts Money'], figsize=(12,10), linewidth=3, grid=True)
ax.ticklabel_format(style='plain')
#display(df_plot)
return("compound example")
def df_output(i, r, a1, a2, y1, y2):
#create and display dataframe
df_plot = get_compound_dual_df(i, r, a1, a2, y1, y2)
#ax = df_plot.plot.line(x='Age', y=['Your Money', 'Friends Money', 'Matts Money'], figsize=(12,10), linewidth=3, grid=True)
#ax.ticklabel_format(style='plain')
display(df_plot)
return("compound example")
# -
# # FIRE Overview
#a 5 minute overview of FIRE
YouTubeVideo('8si7cqw9wm0')
# # Track your spending - budgeting
#
# ## First steps to acheiving Financial Independence
# 1. Understand how much you make and how much you spend
# 2. Understand where you spend your money
# - Everyone has different styles or preferences (more or less granularity)
# - Here are my bins (housing, health, food, transportation, utilities, miscellaneous)
#
# **Tools**
# - YNAB & MINT
# - Excel
# - Quicken
# - Bank apps
# ## How soon can you retire?
# [FIRE Retirement Calculator](https://playingwithfire.co/retirementcalculator/)
# # Steps to lower your spending
# ### 1. Payoff debts
# ### 2. Lower spending
# ### 3. Build Emergency funds
# ### 4. Invest
# ## How to lower your spending without being miserable?
# ### Why are you buying that? (Greed or Fear)
# ### Does that thing really make you happy?
# - The concept of badassity.
# - The challenge of consumerism
# ## Be nice to your future self
# ### 1 coffee at \\$4 a day times 365 days in a year = \\$1460
# ### minus 1 bag of coffee \\$8 each week at home = \\$416
# ### total savings = \\$1044 per year
fs_res = interactive(fs_output, i=fs_investment)
#widgets.jslink((fs_play, 'value'), (fs_investment, 'value'))
#widgets.VBox([fs_play, fs_res])
widgets.VBox([fs_res])
# ## Be really nice to your future self
fs_rn_res = interactive(fs_rn_output, i=fs_rn_investment)
widgets.jslink((fs_rn_play, 'value'), (fs_rn_investment, 'value'))
widgets.VBox([fs_rn_play, fs_rn_res])
# # Investing
# ## Warning I am NOT an accredited financial advisor
# ### Investing versus saving
# ### Investment options
# - Savings
# - Money Market
# - Certificate of Deposit (CD)
# - Stocks
# - Company investments (401k, Employee Stock Purchase Plans, etc.) - read the fine print - matching, vesting
# - Home/property
# - Others (Lending Club, Peer Street)
#
# ### Invest early and often (more compound interest magic)
# ### Set it and forget it
r_res = interactive(r_output, r=rate, a=age, sal=salary, sp=spending, w=net_worth)
#widgets.jslink((play, 'value'), (irate, 'value'))
widgets.VBox([r_res])
# ## 4% and 25 times rules
# [FIRE Retirement Calculator](https://playingwithfire.co/retirementcalculator/)
# ## The magic of compound interest
res = interactive(output, i=investment, r=irate, a1=your_investing_start_age, a2=friend_investing_start_age, y1=years_you_invest, y2=years_friend_invests)
#widgets.jslink((play, 'value'), (irate, 'value'))
widgets.VBox([res])
# # Takeaway
# - Track your spending - budgeting
# - Payoff debts
# - Lower your spending
# - Build emergency fund
# - Invest for the future
# - Be nice to your future self
# - How soon can you retire?
# - The 4% rule
# - Invest early and often (more compound interest magic)
# ## Links
# - FIRE Intro on Youtube - https://www.youtube.com/watch?v=8si7cqw9wm0
# - FIRE retirement calculator - https://playingwithfire.co/retirementcalculator/
# - Mr. Money Mustache & the 4% Rule - https://www.mrmoneymustache.com/2012/05/29/how-much-do-i-need-for-retirement/
# - compound interest - https://www.fool.com/how-to-invest/thirteen-steps/step-1-change-your-life-with-one-calculation.aspx
| notebooks/.ipynb_checkpoints/Original Intern FIRE 1010 Lesson Plan-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Ensemble methods. Stacking
#
#
# Stacking method use several classifiers and use them to generate an input feature matrix for a stacked classifier that do the final prediction.
#
# Let's load the data set first.
# %store -r data_set
# %store -r labels
# %store -r test_data_set
# %store -r test_labels
# %store -r unique_labels
# To simplify the notebook, we use methods that are available in scikit-learn package. We load only several, but in the exercise you gonna need to load other methods as well.
#
# ## Stacking
#
# We have the following steps:
#
# * create $T$ classifiers and learn each to get $m$ predictions (hypothesis $h_{t}$,
# * construct data set of predictions and construct a new classifier $C_{m}$ for each dataset,
# * construct a $C_{h}$ classifier that combines all $C_{m}$ classifiers.
#
# First of all, let's import required libraries:
import numpy as np
from sklearn.discriminant_analysis import QuadraticDiscriminantAnalysis
from sklearn.linear_model import LinearRegression
from sklearn.metrics import accuracy_score
from sklearn.neighbors import KNeighborsClassifier
from sklearn.tree import DecisionTreeClassifier
# In the first part we build three different models based on three different classifiers:
def build_classifiers():
neighbors = KNeighborsClassifier()
neighbors.fit(data_set, labels)
linear_regression = LinearRegression()
linear_regression.fit(data_set, labels)
qda = QuadraticDiscriminantAnalysis()
qda.fit(data_set, labels)
return neighbors, linear_regression, qda
# Based on the classifiers prediction, we build a feature vector for the decision tree classifier. Finally, we train and predict with the stacked classifier.
def build_stacked_classifier(classifiers):
output = []
for classifier in classifiers:
output.append(classifier.predict(data_set))
decision_tree = DecisionTreeClassifier()
output = np.array(output).reshape((130,3))
# stacked classifier part:
decision_tree.fit(output.reshape((130,3)), labels.reshape((130,)))
test_set = []
for classifier in classifiers:
test_set.append(classifier.predict(test_data_set))
test_set = np.array(test_set).reshape((len(test_set[0]),3))
predicted = decision_tree.predict(test_set)
return predicted
# Stacked classifier accuracy can be measured as below:
classifiers = build_classifiers()
predicted = build_stacked_classifier(classifiers)
accuracy = accuracy_score(test_labels, predicted)
print(accuracy)
# In this case, the three used classifiers does not give any value, because we get a higher value using just the decision treee classifier.
decision_tree = DecisionTreeClassifier()
decision_tree.fit(data_set, labels)
predicted = decision_tree.predict(test_data_set)
accuracy = accuracy_score(test_labels, predicted)
print(accuracy)
# ## Grading
#
# Grading is a stacking type where we train a grading classifier that checks if our classifier is good or not on the prediction. We need to build classifiers like we did in the previous example:
def build_classifiers():
neighbors = KNeighborsClassifier()
neighbors.fit(data_set, labels)
qda = QuadraticDiscriminantAnalysis()
qda.fit(data_set, labels)
return neighbors, qda
# In the method below we get a vector of model grads:
def calculate_accuracy_vector(predicted, labels):
result = []
for i in range(len(predicted)):
if predicted[i] == labels[i]:
result.append(1)
else:
result.append(0)
return result
# Finally, we can build our grading meta classifier:
def build_grading_classifier(classifiers):
output = []
matrix = []
for classifier in classifiers:
predicted = classifier.predict(data_set)
output.append(predicted)
matrix.append(calculate_accuracy_vector(predicted,labels))
grading_classifiers = []
for i in range(len(classifiers)):
tree = DecisionTreeClassifier()
tree.fit(data_set, matrix[i])
grading_classifiers.append(tree)
return grading_classifiers
# Test prediction function takes the model and grading model and return the prediction and grad of the prediction:
def test_prediction(classifiers, grading_classifiers, i):
prediction = classifiers[i].predict(test_data_set)
grad = grading_classifiers[i].predict(test_data_set)
return prediction, grad
# We can test now the output, labels and grads of the output:
# +
classifiers = build_classifiers()
grading_classifiers = build_grading_classifier(classifiers)
prediction, grad = test_prediction(classifiers, grading_classifiers, 0)
print(prediction)
print(grad)
print(test_labels)
| ML1/ensemble/074Ensemble_Stacking.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.10 64-bit (''venv'': venv)'
# language: python
# name: python3
# ---
# +
# utility file to get some stats on the data
import glob
import json
import matplotlib.pyplot as plt
data_dir = "../webapp/data/"
language_codes = [f.split("/")[-1] for f in glob.glob(f"{data_dir}/languages/*")]
print("Total languages:", len(language_codes))
# load the 5words.txt file in each language and count the number of words
for code in language_codes:
with open(f"{data_dir}languages/{code}/{code}_5words.txt") as f:
words = f.read().split("\n")
print(f"{code}: {len(words)} words")
# +
# plot the number of words for each language
n_words = {}
for code in language_codes:
with open(f"{data_dir}languages/{code}/{code}_5words.txt") as f:
words = f.read().split("\n")
n_words[code] = len(words)
# plot
plt.figure(figsize=(20, 10))
plt.ylim(0, 10000)
plt.bar(range(len(n_words)), list(n_words.values()))
plt.xticks(range(len(n_words)), list(n_words.keys()), rotation=60)
plt.savefig("out/n_words.png")
plt.show()
# +
# print a status list for the README
with open(f"{data_dir}/languages.json", "r") as f:
languages_data = json.load(f)
def load_supplemental_words(lang):
try:
with open(f"{data_dir}languages/{lang}/{lang}_5words_supplement.txt", "r") as f:
supplemental_words = [line.strip() for line in f]
except FileNotFoundError:
supplemental_words = []
return supplemental_words
with open("out/status_list.txt", "w", encoding="utf-8") as f:
for code in language_codes:
with open(f"{data_dir}languages/{code}/{code}_5words.txt") as g:
words = g.read().split("\n")
words_supplement = load_supplemental_words(code)
status_emoji = "🟥"
if len(words) + len(words_supplement) > 500:
status_emoji = "🟧"
if len(words) + len(words_supplement) > 1000:
status_emoji = "🟨"
# if we've added words to the supplemental list, then we probably have a pretty good
# "good words" + "possible" words list, thus green status emoji
if len(words_supplement) > 2:
status_emoji = "🟩"
language_name = languages_data[code]['language_name']
f.write(f" - {language_name[:]} ({code}): {' '*(25-(len(language_name)+len(code)))} {status_emoji} ({len(words) + len(words_supplement)} words)\n")
# -
| scripts/status.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 0.6.4
# language: julia
# name: julia-0.6
# ---
# # 08 函数の凸性と不等式への応用
#
# 黒木玄
#
# 2018-06-11
#
# * Copyright 2018 <NAME>
# * License: MIT https://opensource.org/licenses/MIT
# * Repository: https://github.com/genkuroki/Calculus
#
# このファイルは次の場所できれいに閲覧できる:
#
# * http://nbviewer.jupyter.org/github/genkuroki/Calculus/blob/master/08%20convexity.ipynb
#
# * https://genkuroki.github.io/documents/Calculus/08%20convexity.pdf
#
# このファイルは <a href="https://juliabox.com">Julia Box</a> で利用できる.
#
# 自分のパソコンに<a href="https://julialang.org/">Julia言語</a>をインストールしたい場合には
#
# * <a href="http://nbviewer.jupyter.org/gist/genkuroki/81de23edcae631a995e19a2ecf946a4f">WindowsへのJulia言語のインストール</a>
#
# を参照せよ.
#
# 論理的に完璧な説明をするつもりはない. 細部のいい加減な部分は自分で訂正・修正せよ.
#
# $
# \newcommand\eps{\varepsilon}
# \newcommand\ds{\displaystyle}
# \newcommand\Z{{\mathbb Z}}
# \newcommand\R{{\mathbb R}}
# \newcommand\C{{\mathbb C}}
# \newcommand\QED{\text{□}}
# \newcommand\root{\sqrt}
# \newcommand\bra{\langle}
# \newcommand\ket{\rangle}
# \newcommand\d{\partial}
# $
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#凸函数の定義" data-toc-modified-id="凸函数の定義-1"><span class="toc-item-num">1 </span>凸函数の定義</a></span></li><li><span><a href="#2階の導函数の符号と函数の凸性" data-toc-modified-id="2階の導函数の符号と函数の凸性-2"><span class="toc-item-num">2 </span>2階の導函数の符号と函数の凸性</a></span><ul class="toc-item"><li><span><a href="#2階の導函数の符号と接線の関係" data-toc-modified-id="2階の導函数の符号と接線の関係-2.1"><span class="toc-item-num">2.1 </span>2階の導函数の符号と接線の関係</a></span></li><li><span><a href="#2階の導函数の符号と函数の凸性の関係" data-toc-modified-id="2階の導函数の符号と函数の凸性の関係-2.2"><span class="toc-item-num">2.2 </span>2階の導函数の符号と函数の凸性の関係</a></span></li><li><span><a href="#指数・対数函数への応用" data-toc-modified-id="指数・対数函数への応用-2.3"><span class="toc-item-num">2.3 </span>指数・対数函数への応用</a></span></li></ul></li><li><span><a href="#Jensenの不等式" data-toc-modified-id="Jensenの不等式-3"><span class="toc-item-num">3 </span>Jensenの不等式</a></span><ul class="toc-item"><li><span><a href="#期待値汎函数" data-toc-modified-id="期待値汎函数-3.1"><span class="toc-item-num">3.1 </span>期待値汎函数</a></span></li><li><span><a href="#Jensenの不等式とその証明" data-toc-modified-id="Jensenの不等式とその証明-3.2"><span class="toc-item-num">3.2 </span>Jensenの不等式とその証明</a></span></li><li><span><a href="#相加相乗平均の不等式" data-toc-modified-id="相加相乗平均の不等式-3.3"><span class="toc-item-num">3.3 </span>相加相乗平均の不等式</a></span></li><li><span><a href="#Youngの不等式" data-toc-modified-id="Youngの不等式-3.4"><span class="toc-item-num">3.4 </span>Youngの不等式</a></span></li><li><span><a href="#Hölderの不等式" data-toc-modified-id="Hölderの不等式-3.5"><span class="toc-item-num">3.5 </span>Hölderの不等式</a></span><ul class="toc-item"><li><span><a href="#$L^p$-ノルム" data-toc-modified-id="$L^p$-ノルム-3.5.1"><span class="toc-item-num">3.5.1 </span>$L^p$ ノルム</a></span></li><li><span><a href="#一般化されたHölderの不等式" data-toc-modified-id="一般化されたHölderの不等式-3.5.2"><span class="toc-item-num">3.5.2 </span>一般化されたHölderの不等式</a></span></li><li><span><a href="#Cauchy-Schwarzの不等式" data-toc-modified-id="Cauchy-Schwarzの不等式-3.5.3"><span class="toc-item-num">3.5.3 </span>Cauchy-Schwarzの不等式</a></span></li><li><span><a href="#分配函数の対数凸性" data-toc-modified-id="分配函数の対数凸性-3.5.4"><span class="toc-item-num">3.5.4 </span>分配函数の対数凸性</a></span></li></ul></li><li><span><a href="#Minkowskiの不等式" data-toc-modified-id="Minkowskiの不等式-3.6"><span class="toc-item-num">3.6 </span>Minkowskiの不等式</a></span></li></ul></li></ul></div>
# +
using Plots
gr(); ENV["PLOTS_TEST"] = "true"
#clibrary(:colorcet)
clibrary(:misc)
function pngplot(P...; kwargs...)
sleep(0.1)
pngfile = tempname() * ".png"
savefig(plot(P...; kwargs...), pngfile)
showimg("image/png", pngfile)
end
pngplot(; kwargs...) = pngplot(plot!(; kwargs...))
showimg(mime, fn) = open(fn) do f
base64 = base64encode(f)
display("text/html", """<img src="data:$mime;base64,$base64">""")
end
using SymPy
#sympy[:init_printing](order="lex") # default
#sympy[:init_printing](order="rev-lex")
using SpecialFunctions
using QuadGK
# -
# ## 凸函数の定義
# **定義:** 区間 $I$ 上の実数値函数 $f$ が**下に凸**な函数であるとは, 任意の $a,b\in I$ と実数 $t$ について,
#
# $$
# 0\leqq t\leqq 1 \implies f((1-t)a+tb) \leqq (1-t)f(a)+t f(b)
# $$
#
# を満たしていることである. 逆向きの不等式で上に凸な函数を定義する. すなわち, 区間 $I$ 上の実数値函数 $f$ が**上に凸**な函数であるとは, 任意の $a,b\in I$ と実数 $t$ について,
#
# $$
# 0\leqq t\leqq 1 \implies f((1-t)a+tb) \geqq (1-t)f(a)+t f(b)
# $$
#
# を満たしていることである.
# +
# log x は上に凸な函数
x = 0:0.01:2.0
a, b = 0.3, 1.5
f(x) = log(x)
t = 0:0.01:1.0
g(a,b,t) = (1-t)*f(a) + t*f(b)
X(a,b,t) = (1-t)*a + t*b
plot(size=(400,250), legend=:topleft, xlims=(0,2.0), ylims=(-2.0, 0.8))
plot!(x, f.(x), label="y = log x")
plot!(X.(a,b,t), g.(a,b,t), label="")
plot!([a,a], [-10.0, f(a)], label="x = a", ls=:dash)
plot!([b,b], [-10.0, f(b)], label="x = b", ls=:dash)
# +
# e^x は下に凸な函数
x = -1:0.01:2
a, b = -0.3, 1.5
f(x) = e^x
t = 0:0.01:1.0
g(a,b,t) = (1-t)*f(a) + t*f(b)
X(a,b,t) = (1-t)*a + t*b
plot(size=(400,250), legend=:topleft, xlims=(-1,2), ylims=(0,8))
plot!(x, f.(x), label="y = e^x")
plot!(X.(a,b,t), g.(a,b,t), label="")
plot!([a,a], [-0.0, f(a)], label="x = a", ls=:dash)
plot!([b,b], [-0.0, f(b)], label="x = b", ls=:dash)
# -
# ## 2階の導函数の符号と函数の凸性
# ### 2階の導函数の符号と接線の関係
#
# **定理:** 区間 $I$ 上の実数値函数 $f$ が $C^2$ 級でかつ $I$ 上で $f''\geqq 0$ ならば, $a\in I$ における $y=f(x)$ のグラフは接線 $y=f'(a)(x-a)+f(a)$ の上側にある. 同様に区間 $I$ 上の実数値函数 $f$ が $C^2$ 級でかつ $I$ 上で $f''\leqq 0$ ならば $a\in I$ における $y=f(x)$ のグラフは接線 $y=f'(a)(x-a)+f(a)$ の下側にある.
#
# **証明:** 前半のみを証明する. 区間 $I$ 上の実数値函数 $f$ が $C^2$ 級でかつ $I$ 上で $f''\geqq 0$ であると仮定し, $a\in I$ とする. $f$ は $C^2$ 級なのでTaylorの公式より, $x\in I$ について,
#
# $$
# f(x) = f(a) + f'(a)(x-a) + R, \qquad
# R = f''(\xi)\frac{(x-a)^2}{2}.
# $$
#
# ここで $\xi$ は $a$ と $x$ のあいだのある実数である. $f''\geqq 0$ より, $R\geqq 0$ となる. ゆえに, $f(x)\geqq f(a) + f'(a)(x-a)$ となる. これで, $y=f(x)$ のグラフは接線 $y=f'(a)(x-a)+f(a)$ の上側にあることがわかった. $\QED$
#
# **注意:** 上の定理の $C^2$ 級の仮定は2回微分可能性の仮定に弱めることができる. $\QED$
#
# **注意:** 上の定理の証明中の剰余項は
#
# $$
# R = \int_a^x dx_1\int_a^{x_1}\,dx_2\;f''(x_2)
# $$
#
# とも書ける. $f''\geqq 0$ のとき, $x>a$ ならば $x_1,x_2$ は $a<x_1<x_2<x$ の範囲を動くので積分の結果は $0$ 以上になり, $x<a$ ならば $x_1,x_2$ は $x<x_1<x_2<a$ の範囲を動き, $x_2$ のみに関する積分の結果は $0$ 以下になり, さらに $x_1$ についても積分すると $0$ 以上になる. $\ds\int_b^a f(x)\,dx = -\int_a^b f(x)\,dx$ であることに注意せよ. $\QED$
# ### 2階の導函数の符号と函数の凸性の関係
#
# **定理:** 区間 $I$ 上の実数値函数 $f$ が $C^2$ 級でかつ $I$ 上で $f''\geqq 0$ ならば $f$ は下に凸である. 同様に区間 $I$ 上の実数値函数 $f$ が $C^2$ 級でかつ $I$ 上で $f''\leqq 0$ ならば $f$ は上に凸である.
#
# **証明:** 前半のみを証明する. 区間 $I$ 上の実数値函数 $f$ が $C^2$ 級でかつ $I$ 上で $f''\geqq 0$ であると仮定する. $a,b\in I$ と仮定し, $c=(1-t)a+tb$ とおく. $0\leqq t\leqq 1$ であると仮定する. $(1-t)f(a)+tf(b)\geqq f(c)$ を示せばよい. $f$ は $C^2$ 級なので Taylor の公式より,
#
# $$
# \begin{aligned}
# &
# f(a) = f(c) + f'(c)(a-c) + R(a),
# \\ &
# f(b) = f(c) + f'(c)(b-c) + R(b),
# \\ &
# R(x) = f''(\xi_x)\frac{(x-c)^2}{2}.
# \end{aligned}
# $$
#
# ここで, $\xi_x$ は $c$ と $x$ のあいだのある実数である. $f''\geqq 0$ という仮定より, $R(x)\geqq 0$ となることがわかる. ゆえに
#
# $$
# \begin{aligned}
# &
# f(a) \geqq f(c) + f'(c)(a-c),
# \\ &
# f(b) \geqq f(c) + f'(c)(b-c).
# \end{aligned}
# $$
#
# 1つ目の不等式の両辺に $(1-t)$ をかけ, 2つめの不等式の両辺に $t$ をかけると
#
# $$
# \begin{aligned}
# (1-t)f(a) &\geqq (1-t)f(c) + f'(c)((1-t)a-(1-t)c),
# \\
# t\;f(b) &\geqq tf(c) + f'(c)(tb-tc).
# \end{aligned}
# $$
#
# これらをたすと, $c = (1-t)a+tb$ とおいたことより,
#
# $$
# \begin{aligned}
# (1-t)f(a)+tf(b) &\geqq f(c) + f'(c)((1-t)a+tb-c) = f(c).
# \end{aligned}
# $$
#
# これで $f$ が下に凸であることが示された. $\QED$
#
# **注意:** 上の証明の本質はTaylorの公式の剰余項が0以上になることである. $f$ が $C^2$ 級であるという仮定を2回微分可能性の仮定に弱めても, $c$ と $x$ のあいだのある実数 $\xi_x$ が存在して,
#
# $$
# f(x) = f(c) + f'(c)(x-c) + R, \quad R = \frac{1}{2}f''(\xi_x)(x-c)
# $$
#
# となるという形でTaylorの定理が成立している. このことを使えば上の定理における $C^2$ 級の仮定を2回微分可能性の仮定に弱めることができる. $\QED$
#
# **問題:** 上と同様にして $f''\leqq 0$ ならば $f$ が上に凸であることの証明を書き下せ. $\QED$
#
# **例:** $f(x)=e^{ax}$ とおくと $f'(x)=ae^{ax}$, $f''(x)=a^2 e^{ax}\geqq 0$ を満たしているので, $f$ は下に凸な函数である. $\QED$
#
# **例:** $x>0$ に対して $f(x)=\log x$ とおくと $f'(x)=1/x$, $f''(x)=-1/x^2<0$ なので, $f$ は上に凸な函数である. $\QED$
#
# **例題:** $a$ は実数であるとし, $x>0$ に対して $f(x)=x^a$ とおく. $f(x)$ は凸性について調べよ.
#
# **解答例:** $x>0$ であるとする. $f'(x)=ax^{a-1}$, $f''(x)=a(a-1)x^{a-2}$ であり, $x^{a-2}>0$ である. ゆえに, $a(a-1)\geqq 0$ のとき, すなわち $a\leqq 0$ または $1\leqq a$ のとき, $f$ は下に凸になり, $a(a-1)\leqq 0$ のとき, すなわち $0\leqq a\leqq 1$ のとき, $f$ は上に凸になる. $\QED$
# ### 指数・対数函数への応用
#
# **問題:** すべての $x\in\R$ について $1+x\leqq e^x$ となることを示せ.
#
# **解答例:** $y=e^x$ の $x=0$ での接線は $y=1+x$ であり, $f''(x)=e^x\geqq 0$ であるから, 上の定理より, $1+x\leqq ex$. $\QED$
#
# **注意:** この問題の結果より, $1-x \leqq e^{-x}$ なので, $x<1$ のとき $e^x\leqq (1-x)^{-1}$ となる. これで
#
# $$
# 1+x\leqq e^x\leqq (1-x)^{-1} \qquad (x<1)
# $$
#
# となることがわかった. $a>0$, $-a\leqq t<a$ のとき, この不等式の $x=\ds\frac{t}{a}$ を代入して全体を $a$ 乗すると次が得られる:
#
# $$
# \left(1+\frac{t}{a}\right)^a \leqq e^t \leqq \left(1-\frac{t}{a}\right)^{-a}
# \qquad (-a\leqq t< a).
# \qquad \QED
# $$
#
# **注意:** 上の問題の結果より, $x>-1$ のとき, $\log(1+x)\leqq x$ となることもわかる. $\QED$
# **問題:** $x>-1$ のとき $\ds\frac{x}{1+x}\leqq \log(1+x)$ となることを示せ.
#
# **解答例:** $\ds f(x) = \log(1+x) - \frac{x}{1+x}$ とおく. $f(0)=0$ である. $\ds\frac{x}{1+x}=1-\frac{1}{1+x}$ と使って, $f$ を微分すると,
#
# $$
# f'(x) = \frac{1}{1+x} - \frac{1}{(1+x)^2} = \frac{x}{(1+x)^2}
# $$
#
# なので, $x<0$ において $f'<0$ で $f$ は単調減少し, $x>0$ において $f'>0$ で $f$ は単調増加する. さらに $f(0)=0$ なので, $f(x)\geqq 0$ すなわち, $\ds \frac{x}{1+x}\leqq \log(1+x)$ である. $\QED$
#
# **注意:** $\ds f''(x)=\frac{1-x}{(x+1)^3}$ なので, $-1<x\leqq 1$ で $f''(x)\geqq 0$ となるが, $x>1$ で $f''(x)<0$ となってしまうので, 上の問題と同様にこの問題を解くことはできない. $\QED$
#
# **以上のまとめ:** $x>-1$ のとき
#
# $$
# \frac{x}{1+x}\leqq \log(1+x)\leqq x.
# \qquad\QED
# $$
f(x) = x/(1+x)
g(x) = log(1+x)
x = -0.99:0.01:3
plot(size=(400, 250), legend=:bottomright, xlim=(-1,3), ylims=(-6,1.5))
plot!(x, f.(x), label="x/(1+x)")
plot!(x, g.(x), label="log(1+x)")
# **問題:** $\ds -\frac{1}{2}\leqq x\leqq 0$ ならば $\log(1+x)\geqq 2x$ となることを示せ.
#
# **解答例:** $f(x)=\log(1+x)-2x$ とおく. $f(0)=0$ である. ゆえに $\ds -\frac{1}{2}\leqq x\leqq 0$ で単調減少するならば, $\ds -\frac{1}{2}\leqq x\leqq 0$ で $f(x)\leqq 0$ となり, 示したいことが示される. そのことを確認するために $f(x)$ を微分すると,
#
# $$
# f'(x) = \frac{1}{1+x}-2 = \frac{-(2x+1)}{1+x}
# $$
#
# なので $\ds x\geqq-\frac{1}{2}$ ならば $f'(x)\leqq 0$ となるので, そこで $f(x)$ は単調減少する. $\QED$
#
# **注意:** $f(x)=\log(1+x)-2x$ は上に凸な函数なので, $f(a)=0$ となる $a<0$ を取れば $f(x)\geqq 0$ と $a\leqq x\leqq 0$ は同値になる. 実際に $a$ を計算すると $a=-0.79681213\cdots$ となる. $\QED$
# +
f(x) = log(1+x) - 2x
@show f(-0.79681213)
@show f(-0.79681214)
a = -0.79681213
x = -0.999:0.001:0.1
plot(size=(400,250), legend=:bottomright, ylims=(-1,0.5))
plot!(x, f.(x), label="log(1+x) - 2x")
plot!(x, zeros(x), label="", color="black", ls=:dot)
plot!([a,a], [-1,0], label="x=$a", color=:red, ls=:dash)
# -
# **問題:** $a>0$ であるとし, $-a<t<a$ であるとする. $\ds\left(1+\frac{t}{a}\right)^a$ は $a$ について単調増加し, $\ds\left(1-\frac{t}{a}\right)^{-a}$ は $a$ について単調減少することを示せ.
#
# **解答例:** それぞれの対数を取ってから $a$ で微分する, 上の方の問題で示した $\log(1+x)$ に関する不等式
#
# $$
# \frac{x}{1+x} \leqq \log(1+x)\qquad (x>-1)
# $$
#
# より,
#
# $$
# \begin{aligned}
# &
# \frac{\d}{\d a}\log\left(1+\frac{t}{a}\right)^a =
# \log\left(1+\frac{t}{a}\right) - \frac{t/a}{1+t/a} \geqq 0,
# \\ &
# \frac{\d}{\d a}\log\left(1-\frac{t}{a}\right)^{-a} =
# -\log\left(1-\frac{t}{a}\right) + \frac{-t/a}{1-t/a} \leqq 0.
# \end{aligned}
# $$
#
# ゆえに, $\ds\left(1+\frac{t}{a}\right)^a$ は $a$ について単調増加し, $\ds\left(1-\frac{t}{a}\right)^{-a}$ は $a$ について単調減少する. $\QED$
# **問題:** 以上の結果をまとめて用いて,
#
# $$
# \lim_{a\to\infty}\left(1+\frac{t}{a}\right)^a =
# \lim_{a\to\infty}\left(1-\frac{t}{a}\right)^{-a} =
# e^t
# $$
#
# を証明せよ.
#
# **解答例:** 上の方の注意では
#
# $$
# \left(1+\frac{t}{a}\right)^a \leqq e^t \leqq \left(1-\frac{t}{a}\right)^{-a}
# \tag{$*$}
# \qquad (-a<t<a)
# $$
#
# という不等式が成立していることを指摘している. さらにすぐ上の問題によって, $\ds\left(1+\frac{t}{a}\right)^a$, $\ds\left(1-\frac{t}{a}\right)^{-a}$ はそれぞれ $a$ について単調増加, 単調減少するので, どちらも $a\to\infty$ で収束する. $a$ を十分に大きくして $\ds\frac{t^2}{a^2}\leqq\frac{1}{2}$ となるようにすると, 上の方の問題で示した不等式
#
# $$
# \log(1+x) \geqq 2x \qquad \left(-\frac{1}{2}\leqq x\leqq 0\right)
# $$
#
# を使うことができて,
#
# $$
# 0\geqq
# \log\frac{(1+t/a)^a}{(1-t/a)^{-a}} =
# a\log\left(1-\frac{t^2}{a^2}\right) \geqq
# a\cdot 2\left(-\frac{t^2}{a^2}\right) =
# -\frac{2t^2}{a} \to 0 \quad (q\to\infty).
# $$
#
# となるので, $a\to\infty$ のとき $\ds \log\frac{(1+t/a)^a}{(1-t/a)^{-a}}\to 0$ すなわち $\ds \frac{(1+t/a)^a}{(1-t/a)^{-a}}\to 1$ となる. これで $\ds\left(1+\frac{t}{a}\right)^a$, $\ds\left(1-\frac{t}{a}\right)^{-a}$ が同じ値に収束することがわかった. したがって不等式($*$)より, 欲しい結果が得られる. $\QED$
#
# **注意:** 不等式($*$)は極限で指数函数が現われる結果の極限操作を初等的に実行したいときに便利な不等式である. $\QED$
# +
f(t) = e^t
g(a,t) = (1+t/a)^a
h(a,t) = (1-t/a)^(-a)
t = -3:0.01:3
PP = []
for a in [5, 10, 20, 40]
P = plot(title="a=$a", titlefontsize=10, legendfontsize=7)
plot!(legend=:topleft, yscale=:log, ylims=(10^(-1.3), 10^1.3))
plot!(t, h.(a,t), label="(1-t/a)^(-a)", ls=:dash)
plot!(t, f.(t), label="e^t")
plot!(t, g.(a,t), label="(1+t/a)^a", ls=:dash)
push!(PP, P)
end
plot(PP[1:2]..., size=(750,250), layout=@layout([a b]))
# -
plot(PP[3:4]..., size=(750,250), layout=@layout([a b]))
# ## Jensenの不等式
# ### 期待値汎函数
#
# 函数を数に対応させる函数を**汎函数** (functional)と呼ぶことがある.
#
# **函数の例:** $x$ を $x$ に対応させる函数を $x$ と書く.
#
# すべての $x$ を一定の値 $\alpha$ に対応させる函数も同じ記号の $\alpha$ で書き, 定数函数と呼ぶ. $f(x)=\alpha$ とおいて記号 $f$ を利用すると煩雑になるので, 単に $\alpha$ と書いて代用するということである. 数としての $\alpha$ と定数函数としての $\alpha$ を同じ記号で書くので混乱しないように注意して欲しい.
#
# 例えば, すべての $x$ を $1$ に対応させる函数も単に $1$ と書く.
# $\QED$
#
# **定義:** 函数 $f$ を数 $E[f]$ に対応させる函数 $E[\ ]$ (汎函数)が以下の条件を満たしているとき, $E[\ ]$ は**期待値汎函数** (expextation functional)であると言うことにする:
#
# **(1) 線形性:** 函数 $f,g$ と数 $\alpha,\beta$ に対して, $E[\alpha f+\beta g]=\alpha E[f]+\beta E[g]$.
#
# **(2) 単調性:** 函数 $f, g$ のあいだで常に $f\leqq g$ が成立しているならば $E[f]\leqq E[g]$.
#
# **(3) 規格化条件:** 定数函数 $\alpha$ について, $E[\alpha]=\alpha$. $\QED$
# **例:** 区間 $I$ 上の函数を数 $E[f]$ に対応させる函数 $E[\ ]$ を以下のように定めると, $E[\ ]$ は期待値汎函数になる. まず $a_1,\ldots,a_n\in I$ を任意に取る. $w_1,\ldots,w_n$ は0以上の実数でかつ $w_1+\cdots+w_n=1$ を満たしていると仮定する. そして区間 $I$ 上の函数 $f$ に対して
#
# $$
# E[f] = \sum_{i=1}^n w_i f(a_i) = w_1 f(a_1)+\cdots+w_n f(a_n)
# $$
#
# と定める. このようにして定められた $E[\ ]$ が実際に期待値汎函数の性質を満たしていることは以下のようにして確認できる.
#
# (1) 区間 $I$ 上の函数 $f,g$ と数 $\alpha,\beta$ について
#
# $$
# \begin{aligned}
# E[\alpha f+\beta g] &= \sum_{i=1}^n w_i(\alpha f(a_i)+\beta g(a_i)) =
# \sum_{i=1}^n (\alpha w_i f(a_i)+\beta w_i g(a_i))
# \\ &=
# \alpha\sum_{i=1}^n w_i f(a_i) + \beta\sum_{i=1}^n w_i g(a_i) = \alpha E[f]+\beta E[g].
# \end{aligned}
# $$
#
# (2) 区間 $I$ 上で $f\leqq g$ が成立していると仮定すると, $w_i\geqq 0$ より $w_i f(a_i)\leqq w_i g(a_i)$ も成立するので
#
# $$
# E[f] = \sum_{i=1}^n w_i f(a_i) \leqq \sum_{i=1}^n w_i g(a_i) = E[g].
# $$
#
# (3) 定数函数 $f(x)=\alpha$ について $f(a_i)=\alpha$ が成立しているので, $w_1+\cdots+w_n=1$ という仮定より,
#
# $$
# E[\alpha] = E[f] = \sum_{i=1}^n w_i f(a_i) = \sum_{i=1}^n w_i \alpha = \alpha.
# $$
#
# これで示すべきことがすべて示された.
#
# 特に $w_i=1/n$ であるとき, $E[f]$ は
#
# $$
# E[f] = \frac{f(a_1)+\cdots+f(a_n)}{n}
# $$
#
# と $f(a_i)$ たちの加法平均になる. $\QED$
#
# 抽象的な期待値汎函数の概念が難しいと感じる人は, 一般的な期待値汎函数の定義を忘れてこの例の $E[f]$ またはさらにその特別な場合である加法平均の場合のみを考えれば十分である.
# **問題:** $a<b$ であるとする. 閉区間 $I=[a,b]$ 上の連続函数 $f$ を
#
# $$
# E[f] = \frac{1}{b-a}\int_a^b f(x)\,dx
# $$
#
# に対応させる函数 $E[\ ]$ は期待値汎函数であることを示せ. $\QED$
# **問題:** $\R$ 上の函数 $p(x)$ は $p(x)\geqq 0$ ($x\in\R$) と $\int_{-\infty}^\infty p(x)\,dx = 1$ を満たしていると仮定する. (このような $p(x)$ は**確率密度函数**と呼ばれる.) このとき, $\R$ 上の(適当によい条件を仮定した)函数 $f$ を
#
# $$
# E[f] = \int_{-\infty}^\infty f(x)p(x)\,dx
# $$
#
# に対応させる函数 $E[\ ]$ が期待値汎函数であることを示せ. ($E[f]$ は確率密度函数 $p(x)$ が定める確率分布に関する**確率変数** $f$ の期待値と呼ばれる.)
#
# **注意:** 上の問題はこの問題で
#
# $$
# p(x) = \begin{cases}
# \dfrac{1}{b-a} & (a\leqq x\leqq b) \\
# 0 & (\text{それ以外}) \\
# \end{cases}
# $$
#
# とした場合になっている. $\QED$
#
# **解答例:** 以下では $\int_{-\infty}^\infty$ を $\int$ と略記する.
#
# (1) 数 $\alpha$, $\beta$ と函数 $f$, $g$ について
#
# $$
# E[\alpha f+\beta g] = \int(\alpha f(x)+\beta g(x))p(x)\,dx =
# \alpha\int f(x)p(x)\,dx + \beta\int g(x)p(x)\,dx =
# \alpha E[f]+\beta E[g].
# $$
#
# (2) $\R$ 上で $f\leqq g$ が成立していると仮定すると, $p(x)\geqq 0$ より $f(x)p(x)\leqq g(x)p(x)$ も成立するので
#
# $$
# E[f] = \int f(x)p(x)\,dx \leqq \int g(x)p(x)\,dx = E[g].
# $$
#
# (3) 定数函数 $f(x)=\alpha$ について, $\int p(x)\,dx=1$ という仮定より,
#
# $$
# E[\alpha] = \int \alpha p(x)\,dx = \alpha.
# $$
#
# これで示すべきことがすべて示された. $\QED$
# ### Jensenの不等式とその証明
#
# **Jensenの不等式:** $E[\ ]$ は区間 $I$ 上の函数の期待値汎函数であるとし, $f$ は区間 $I$ 上の上に凸(もしくは下に凸)な函数であるとする. このとき
#
# $$
# E[f(x)] \leqq f(E[x]) \qquad (\text{もしくは}\ E[f(x)]\geqq f(E[x])).
# $$
#
# 例えば
#
# $$
# E[f(x)] = \sum_{i=1}^n w_i f(a_i), \quad w_i\geqq 0, \quad \sum_{i=1}^n w_i = 1
# $$
#
# のとき,
#
# $$
# \begin{aligned}
# &
# w_1 f(a_1) + \cdots + w_n f(a_n) \leqq f(w_1 a_1 + \cdots + w_n a_n)
# \\ &
# (\text{もしくは}\ w_1 f(a_1) + \cdots + w_n f(a_n) \geqq f(w_1 a_1 + \cdots + w_n a_n)).
# \end{aligned}
# $$
#
# 特にこれの $n=2$ の場合には $f$ が上に凸(もしくは下に凸)であることの定義とこれらは同値であることに注意せよ. すなわち, 函数 $f$ がJensenの不等式を満たすことと上(もしくは下)に凸なことは同値になる. この意味でJensenの不等式は函数の凸性を言い換えた結果になっている.
#
# **証明:** 簡単のため $f$ は $C^1$ 級であると仮定する. ($C^1$ 級でない場合にも同様の方法で証明できるが, 「接線」の存在を別に証明する必要が生じる.)
#
# $f$ は上に凸であると仮定する.
#
# $\mu = E[x]$ とおく. $E[f(x)]\leqq f(\mu)$ を示せばよい. $x=\mu$ における $y=f(x)$ の接線を $y=\alpha(x-\mu)+f(\mu)$ と書く. $f$ は上に凸であると仮定したので
#
# $$
# f(x)\leqq \alpha(x-\mu)+f(\mu)
# $$
#
# が成立している(図を描いてみよ). ゆえに $E[\ ]$ の性質より
#
# $$
# E[f(x)]\leqq E[\alpha(x-\mu)+f(\mu)] = \alpha(E[x]-E[\mu]) + E[f(\mu)] = f(\mu).
# $$
#
# 最初の不等号は期待値汎函数の単調性より, 1つ目の等号は線形性より, 2つ目の等号は規格化条件より. これで示すべきことがすべて示された. $\QED$
x = 0:0.01:2.0
μ = 0.7
f(x) = log(x)
g(μ,x) = (1/μ)*(x-μ) + f(μ)
plot(size=(500,350), legend=:topleft, xlims=(0,1.7), ylims=(-2.2, 0.8))
plot!(x, f.(x), label="y = log x")
plot!(x, g.(μ,x), label="y = a(x-mu)+f(mu)")
plot!([μ, μ], [-3.0, f(μ)], label="x=mu", ls=:dash)
# **問題:** 上に凸な函数 $f(x)$ と
#
# $$
# E[g(x)] = w_1 g(a_1)+\cdots+w_n g(a_n), \quad
# w_i\geqq 0, \quad w_1+\cdots+w_n=1
# $$
#
# に関するJensenの不等式
#
# $$
# w_1 f(a_1)+\cdots + w_n f(a_n) \leqq f(w_1 a_1 +\cdots+ w_n a_n)
# \tag{$*$}
# $$
#
# を $n$ に関する数学的帰納法を用いて直接証明せよ.
#
# **解答例:** $n=1$ のとき, ($*$) は $f(a_1)\leqq f(a_1)$ を意味するので, 自明に正しい.
#
# $n=N\geqq 1$ に関する ($*$) が常に成立していると仮定し(帰納法の仮定),
#
# $$
# w_i \geqq 0, \quad w_1+\cdots+w_N+w_{N+1} = 1
# $$
#
# と仮定する. もしも $w_{N+1}=1$ ならば $n=N+1$ の場合の ($*$) は自明に成立している. 以下では $w_{N+1}<1$ と仮定し,
#
# $$
# t_i = \frac{w_i}{1-w_{N+1}}
# $$
#
# とおく. このとき, $w_i\geqq 0$, $w_1+\cdots+w_N=1-w_{N+1}$ より,
#
# $$
# t_i \geqq 0, \quad
# t_1+\cdots+t_N = \frac{w_1+\cdots+w_N}{1-w_{N+1}} = 1.
# $$
#
# したがって, $n=N$ の場合の($*$)より,
#
# $$
# t_1 f(a_1)+\cdots+t_N f(a_N) \leqq f(t_1 a_1+\cdots+t_N a_N).
# $$
#
# $f$ は上に凸と仮定してあったので, $w_i=(1-w_{N+1})t_i$ を使うと,
#
# $$
# \begin{aligned}
# w_1 f(a_1)+\cdots+w_N f(a_N)+w_{N+1}f(a_{N+1}) &=
# (1-w_{N+1})(t_1 f(a_1)+\cdots+t_N f(a_N))+w_{N+1}f(a_{N+1})
# \\ &\leqq
# (1-w_{N+1})f(t_1 a_1+\cdots+t_N a_N) + w_{N+1} f(a_{N+1})
# \\ &\leqq
# f((1-w_{N+1})(t_1 a_1+\cdots+t_N a_N)+w_{N+1} a_{N+1})
# \\ &=
# f(w_1 a_1+\cdots+w_N a_N+w_{N+1} a_{N+1}) .
# \end{aligned}
# $$
#
# ここで, 2行目で $n=N$ の場合の($*$)を使い, 3行目で $f$ が上に凸であることを使った.
#
# ゆえに数学的帰納法によってすべての $n$ について ($*$) が成立する. $\QED$
# ### 相加相乗平均の不等式
# **例:** $a_i>0$ であるとし,
#
# $$
# E[f(x)] = \frac{f(a_1)+\cdots+f(a_n)}{n}
# $$
#
# とおく. $x>0$ に対して $f(x)=\log x$ とおく. このとき, $f(x)=\log x$ は下に凸なので, Jensenの不等式より $E[f(x)] \leqq f(E[x])$. そして,
#
# $$
# E[f(x)] = \frac{\log a_1+\cdots+\log a_n}{n} = \log(a_1\cdots a_n)^{1/n}, \quad
# f(E[x]) = \log\frac{a_1+\cdots+a_n}{n}
# $$
#
# なので, $\log$ の単調増加性より,
#
# $$
# (a_1\cdots a_n)^{1/n} \leqq \frac{a_1+\cdots+a_n}{n}.
# $$
#
# これでJensenの不等式が相加相乗平均の不等式を含んでいることがわかった. $\QED$
# **例題:** 下に凸な函数 $f(x)=e^x$ に関するJensenの不等式を用いて相加相乗平均の不等式を証明し直せ.
#
# **解答例:** $a_i>0$ であるとし, $A_i=\log a_i$, $\ds E[f(x)] = \frac{f(A_1)+\cdots+f(A_n)}{n}$ とおくと, $f(x)=e^x$ のとき
#
# $$
# \begin{aligned}
# &
# E[f(x)] = E[e^x] = \frac{e^{A_1}+\cdots+e^{A_n}}{n} = \frac{a_1+\cdots+a_n}{n},
# \\ &
# f(E[x]) = e^{(A_1+\cdots+A_n)/n} = \left(e^{A_1}\cdots e^{A_n}\right)^{1/n} = (a_1\cdots a_n)^{1/n}.
# \end{aligned}
# $$
#
# 下に凸な函数 $f(x)=e^x$ に関するJensenの不等式 $E[f(x)] \geqq f(E[x])$ より,
#
# $$
# \frac{a_1+\cdots+a_n}{n} \geqq (a_1\cdots a_n)^{1/n}.
# $$
#
# これで示すべきことが示された. $\QED$
# ### Youngの不等式
# **Youngの不等式の一般化:** $p_i>0$, $\ds\frac{1}{p_1}+\cdots+\frac{1}{p_n}=1$, $a_i>0$ のとき
#
# $$
# a_1\cdots a_n \leqq \frac{a_1^{p_1}}{p_1}+\cdots+\frac{a_n^{p_n}}{p_n}
# $$
#
# $n=2$ の場合をYoungの不等式と呼ぶ. すなわち $p>0$, $q>0$, $\ds\frac{1}{p}+\frac{1}{q}=1$, $a>0$, $b>0$ のときの
#
# $$
# ab \leqq \frac{a^p}{p} + \frac{b^q}{q}
# $$
#
# という不等式を**Youngの不等式**と呼ぶ.
# **証明:** 上に凸な函数 $f(x)=\log x$ に関するJensenの不等式より
#
# $$
# \log(a_1\cdots a_n)
# = \frac{\log a_1^{p_1}}{p_1} + \cdots + \frac{\log a_n^{p_n}}{p_n}
# \leqq \log\left(\frac{a_1^{p_1}}{p_1}+\cdots+\frac{a_n^{p_n}}{p_n}\right)
# $$
#
# ゆえに
#
# $$
# a_1\cdots a_n \leqq \frac{a_1^{p_1}}{p_1}+\cdots+\frac{a_n^{p_n}}{p_n}
# \qquad \QED
# $$
# **問題:** $a,b>0$, $p,q>0$, $\ds\frac{1}{p}+\frac{1}{q}=1$ のとき
#
# $$
# ab \leqq \frac{a^p}{p} + \frac{b^p}{q}
# $$
#
# となること(つまりYoungの不等式)を直接的に証明してみよ.
# **解答例1:** $f(x)=\log x$ が上に凸であることより, $A,B>0$ について
#
# $$
# \frac{1}{p}\log A + \frac{1}{q}\log B\leqq \log\left(\frac{A}{p}+\frac{B}{q}\right).
# $$
#
# ゆえに $A=a^p$, $B=b^q$ とおくと,
#
# $$
# \log(ab) = \frac{1}{p}\log a^p + \frac{1}{q}\log b^q\leqq \log\left(\frac{a^p}{p}+\frac{b^q}{q}\right).
# $$
#
# したがって,
#
# $$
# ab \leqq \frac{a^p}{p} + \frac{b^p}{q}. \qquad \QED
# $$
# **解答例2:** $f(x)=e^x$ が下に凸であることより, 実数 $A,B$ について
#
# $$
# \frac{1}{p}e^A + \frac{1}{q}e^B \geqq e^{A/p+B/q} = e^{A/p} e^{B/q}.
# $$
#
# ゆえに $A=\log a^p$, $B=\log b^q$ とおくと,
#
# $$
# \frac{a^p}{p}+\frac{b^q}{q} \geqq ab. \qquad \QED
# $$
# **解答例3:** $f(x)$ は $x\geqq 0$ の狭義単調増加連続函数で $f(0)=0$ を満たすものであるとし,
#
# $$
# X = \{\,(x,y)\mid 0\leqq x\leqq a,\ 0\leqq y\leqq f(x)\,\}, \quad
# Y = \{\,(x,y)\mid 0\leqq x\leqq f^{-1}(y),\ 0\leqq y\leqq b\,\}
# $$
#
# とおくと, $[0,a]\times[0,b]\subset X\cup Y$ となるので(図を描いてみよ), 面積を比較することによって
#
# $$
# ab \leqq \int_0^a f(x)\,dx + \int_0^b f^{-1}(y)\,dy.
# \tag{$\star$}
# $$
#
# $f(x)=x^{p-1}$ とおくと,
#
# $$
# \frac{1}{p}+\frac{1}{q}=1 \iff (p-1)(q-1)=1
# $$
#
# より, $f^{-1}(y)=y^{1/(p-1)}=y^{q-1}$ となることがわかる. これに上の不等式を適用すると
#
# $$
# ab \leqq \frac{a^p}{p} + \frac{b^p}{q}
# $$
#
# が得られる. $\QED$
# **注意:** $a^{p-1}=b$ ならば ($\star$) で等号が成立する.
#
# $$
# \frac{1}{p}+\frac{1}{q}=1 \iff (p-1)(q-1)=1 \iff (p-1)q = p
# $$
#
# なので $a^{p-1}=b$ と $a^p = b^q$ は同値である(前者の両辺を $q$ 乗すると後者になる). $\QED$
# +
function plot_Young(a, b; p=2.5, kwarg...)
q = p/(p-1)
f(x) = x^(p-1)
g(y) = y^(q-1)
A = max(a, g(b))
x = 0:a/200:a
y = 0:b/200:b
x1 = 0:A/200:A
P = plot(legend=false; kwarg...)
plot!([a,a], [0,b], color=:black)
plot!([0,a], [b,b], color=:black)
plot!(x, f.(x), color=:red, fill=(0, 0.4, :pink))
plot!(x1, f.(x1), color=:red, fill=(b, 0.4, :pink))
if A > a
x2 = a:(A-a)/200:A
plot!(x2, f.(x2), color=:red, fill=(b, 0.4, :pink))
end
P
end
P1 = plot_Young(3,2)
P2 = plot_Young(2,5)
plot(P1, P2, size=(600, 300))
# -
# ### Hölderの不等式
# #### $L^p$ ノルム
# 区間 $[a,b]$ 乗の連続函数 $f$ に対して $\|f\|_p$ を
#
# $$
# \|f\|_p = \left(\int_a^b \left|f(x)\right|^p\,dx\right)^{1/p}
# $$
#
# と定める. これを函数 $f$ の $L^p$ ノルムと呼ぶ.
# **問題:** 定数 $\alpha$ について $\left\|\alpha f\right\|_p=|\alpha|\left\|f\right\|_p$ が成立していることを示せ.
#
# **解答例:**
# $$
# \|\alpha f\|_p = \left(\int_a^b \left|\alpha f(x)\right|^p\,dx\right)^{1/p} =
# \left(\left|\alpha\right|^p \int_a^b \left|f(x)\right|^p\,dx\right)^{1/p} =
# \left|\alpha\right| \left(\int_a^b \left|f(x)\right|^p\,dx\right)^{1/p} =
# |\alpha|\|f\|_p. \qquad \QED
# $$
# **注意:** 上の問題の結果より, 特に $g=\dfrac{f}{\|f\|_p}$ とおくと $\|g\|_p=1$ となることがわかる. $\QED$
# #### 一般化されたHölderの不等式
# $p > 0$, $p_i > 0$ であるとし,
#
# $$
# \frac{1}{p_1} + \cdots + \frac{1}{p_n} = \frac{1}{p}
# $$
#
# と仮定する. このとき, 区間 $[a,b]$ 上の連続函数 $f_i$ 達について
#
# $$
# \|f_1\cdots f_n\|_p \leqq \|f_1\|_{p_1}\cdots\|f_n\|_{p_n}
# \tag{$*$}
# $$
#
# が成立する. これを**一般化されたHölderの不等式**と呼ぶことにする. $n=2$, $p=1$ の場合はHölderの不等式と呼ばれる. すなわち, $p>0$, $q>0$ かつ
#
# $$
# \frac{1}{p}+\frac{1}{q}=1
# $$
#
# のときの
#
# $$
# \|fg\|_1 \leqq \|f\|_p \|g\|_q
# $$
#
# という不等式は**Hölderの不等式**と呼ばれる. さらにこれの $p=q=2$ の場合は**Cauchy-Schwarzの不等式**の特別な場合である.
# **一般化されたHölderの不等式の証明の概略:** $g_i(x) = \dfrac{f_i(x)}{\|f_i\|_{p_i}}$ とおく. そのとき $\|g_i\|_{p_i}=1$ となる.
#
# $\ds\frac{1}{p_1} + \cdots + \frac{1}{p_n} = \frac{1}{p}$ は $\ds\frac{p}{p_1} + \cdots + \frac{p}{p_n} = 1$ と同値なので, $q_i=p_i/p$ とおくと,
#
# $$
# q_i>0, \quad p q_i = p_i, \quad \frac{1}{q_1}+\cdots+\frac{1}{q_n} = 1.
# $$
#
# ゆえに $a_i = |g_i(x)|^p$ にYoungの不等式の一般化を適用することによって,
#
# $$
# |g_1(x)\cdots g_n(x)|^p =
# |g_1(x)|^p\cdots|g_n(x)|^p \leqq
# \frac{|g_1(x)|^{pq_1}}{q_1}+\cdots+\frac{|g_n(x)|^{pq_n}}{q_n} =
# \frac{|g_1(x)|^{p_1}}{q_1}+\cdots+\frac{|g_n(x)|^{p_n}}{q_n}.
# $$
#
# これを $x$ で積分すると,
#
# $$
# (\|g_1\cdots g_n\|_p)^p \leqq
# \frac{(\|g_1(x)\|_{p_1})^{p_1}}{q_1}+\cdots+\frac{(\|g_n(x)\|_{p_n})^{p_n}}{q_n} =
# \frac{1}{q_1}+\cdots+\frac{1}{q_n}=1.
# $$
#
# ゆえに,
#
# $$
# \frac{\|f_1\cdots f_n\|_p}{\|f_1\|_{p_1}\cdots\|f_n\|_{p_n}} =
# \|g_1\cdots g_n\|_p\leqq 1.
# $$.
#
# したがって,
#
# $$
# \|f_1\cdots f_n\|_p\leqq \|f_1\|_{p_1}\cdots\|f_n\|_{p_n}. \qquad\QED
# $$
# **問題:** (一般化されていない)Youngの不等式から(一般化されていない)Hölderの不等式を導け.
#
# **解答例の概略:** $p,q>0$, $\ds\frac{1}{p}+\frac{1}{q}=1$ と仮定する.
#
# $$
# \|fg\|_1 \leqq \|f\|_p \|g\|_q
# $$
#
# を証明したい. 左辺を右辺で割ったものが $1$ 以下になることを示せばよい.
#
# $$
# a(x) = \frac{|f(x)|}{\|f\|_p}, \quad
# b(x) = \frac{|g(x)|}{\|g\|_q}
# $$
#
# とおくと, $\|a\|_p=\|b\|_q=1$ となる. $a(x)$, $b(x)$ にYoungの不等式を適用すると,
#
# $$
# a(x)b(x) \leqq \frac{a(x)^p}{p} + \frac{b(x)^q}{q}.
# $$
#
# 両辺を $x$ について積分すると,
#
# $$
# \begin{aligned}
# &
# \int_a^b a(x)b(x)\,dx = \|ab\|_1 = \frac{\|fg\|_1}{\|f\|_p \|g\|_q},
# \\ &
# \int_a^b \left(\frac{a(x)^p}{p} + \frac{b(x)^q}{q}\right)\,dx =
# \frac{(\|a\|_p)^p}{p}+\frac{(\|b\|_q)^q}{q} = \frac{1}{p}+\frac{1}{q} = 1.
# \end{aligned}
# $$
#
# ゆえに
#
# $$
# \frac{\|fg\|_1}{\|f\|_p \|g\|_q} \leqq 1.
# $$
#
# これより, Hölderの不等式が成立することがわかる. $\QED$
# #### Cauchy-Schwarzの不等式
#
# $p,q>0$, $\ds\frac{1}{p}+\frac{1}{q}=1$ と仮定する. このとき, $\|f\|_p<\infty$, $\|g\|_q<\infty$ を満たす函数 $f,g$ の内積を
#
# $$
# \bra f,g\ket = \int_a^b \overline{f(x)}g(x)\,dx
# $$
#
# と定義できる. ここで $\overline{f(x)}$ は $f(x)$ の複素共役を意味している. $f(x)$ が実数ならば $\overline{f(x)}=f(x)$ である. このとき, Hölderの不等式より,
#
# $$
# |\bra f,g\ket|\leqq
# \int_a^b \left|\overline{f(x)}\right|\left|g(x)\right|\,dx =
# \int_a^b \left|f(x)\right|\left|g(x)\right|\,dx =
# \|fg\|_1 \leqq \|f\|_p\|g\|_q.
# $$
#
# すなわち,
#
# $$
# |\bra f,g\ket| \leqq \|fg\|_1 \leqq \|f\|_p\|g\|_q.
# $$
#
# この不等式の $p=q=1/2$ の場合は**Cauchy-Schwarzの不等式**と呼ばれている.
# #### 分配函数の対数凸性
# **問題:** $a<b$ と仮定する. $Z(\beta)=\int_a^b e^{-\beta f(x)+g(x)}\,dx$ の形の函数 $Z(\beta)$ について, $\log Z(\beta)$ が $\beta$ の下に凸な函数になることをHölderの不等式を使って示せ.
#
# **解答例:** $0<t<1$ であるとし, $p=1/(1-t)$, $q=1/t$ とおく. そのとき $\ds\frac{1}{p}+\frac{1}{q}=1$ となる. ゆえにHölderの不等式より,
#
# $$
# \begin{aligned}
# Z((1-t)\alpha+t\beta) &= Z\left(\frac{\alpha}{p}+\frac{\beta}{q}\right) =
# \int_a^b e^{-(\alpha/p+\beta/q)f(x)+(1/p+1/q)g(x)}\,dx =
# \int_a^b \left(e^{-\alpha f(x)+g(x)}\right)^{1/p} \left(e^{-\beta f(x)+g(x)}\right)^{1/q}\,dx
# \\ &\leqq
# \left(\int_a^b e^{-\alpha f(x)+g(x)}\,dx\right)^{1/p}
# \left(\int_a^b e^{-\beta f(x)+g(x)}\,dx\right)^{1/q}
# \\ &=
# Z(\alpha)^{1/p}\, Z(\beta)^{1/q} =
# Z(\alpha)^{1-t}\; Z(\beta)^t.
# \end{aligned}
# $$
#
# ゆえに
#
# $$
# \log Z((1-t)\alpha+t\beta) \leqq (1-t)\log Z(\alpha) + t\log Z(\beta).
# $$
#
# これは $\log Z(\beta)$ が $\beta$ に関する下に凸な函数であることを意味している. $\QED$
#
# **注意:** 上の問題の $Z(\beta)$ の形の函数を**分配函数**と呼ぶことがある(統計力学). 上の問題より, Hölderの不等式は分配函数の対数が下に凸な函数になることを意味していることがわかる. $\QED$
# **例(ガンマ函数の対数凸性):** $s>0$ に対して $\Gamma(s) = \int_0^\infty e^{-x} x^{s-1}\,dx = \int_0^\infty e^{(s-1)\log x-x}\,dx$ とおくと, その対数 $\log\Gamma(s)$ は $s>0$ の下に凸な函数になる. $\QED$
# **注意:** 上の問題の記号のもとで, 函数 $h(x)$ に対する $\bra h(x)\ket=\bra h(y)\ket$ を次のように定める:
#
# $$
# \bra h(x)\ket = \frac{1}{Z(\beta)}\int_a^b h(x) e^{-\beta f(x)+g(x)}\,dx, \quad
# \bra h(y)\ket = \frac{1}{Z(\beta)}\int_a^b h(y) e^{-\beta f(y)+g(y)}\,dy.
# $$
#
# さらに函数 $h(x,y)$ に対する $\bra h(x,y)\ket$ を次のように定める:
#
# $$
# \bra h(x,y)\ket = \frac{1}{Z(\beta)^2}\int_a^b\int_a^b h(x,y) e^{-\beta f(x)+g(x)}e^{-\beta f(y)+g(y)}\,dx\,dy.
# $$
#
# このとき $\bra f(x)\ket^2=\bra f(x)f(y)\ket$ などが成立している.
#
# もしも上の問題の $Z(\beta)$ について $\beta$ に関する微分と積分の順序交換を自由にできるならば
#
# $$
# \begin{aligned}
# &
# -\frac{d}{d\beta}\log Z(\beta) = \frac{\int_a^b e^{-\beta f(x)+g(x)}f(x)\,dx}{Z(\beta)} = \bra f(x)\ket,
# \\ &
# \frac{d^2}{d\beta^2}\log Z(\beta) = \bra f(x)^2\ket - \bra f(x)\ket^2 =
# \bra f(x)f(x)\ket - \bra f(x)f(y)\ket
# \\ &\qquad=
# \frac{1}{2}(\bra f(x)f(x)\ket - 2\bra f(x)f(y)\ket + \bra f(y)f(y)\ket) =
# \frac{1}{2}\bra (f(x)+f(y))^2\ket \geqq 0.
# \end{aligned}
# $$
#
# これからも, $\log Z(\beta)$ が下に凸な函数であることがわかる. $\QED$
# ### Minkowskiの不等式
# $L^1$ ノルムについては, 三角不等式 $|f(x)+g(x)|\leqq |f(x)|+|g(x)|$ より,
#
# $$
# \|f+g\|_1 = \int_a^b |f(x)+g(x)|\,dx
# \leqq \int_a^b(|f(x)|+|g(x)|)\,dx = \|f\|_1 + \|g\|_1
# $$
#
# が成立している. この $L^1$ ノルムに関する三角不等式はそのまま $L^p$ ノルム ($p>1$) に一般化される.
# $p>1$ であると仮定する. $q=\dfrac{p}{p-1}$ とおくと, $\ds\frac{1}{p}+\frac{1}{q}=1$. このとき,
#
# $$
# \left\|\,\left|\varphi\right|^{p-1}\right\|_q=
# \left\|\,\left|\varphi\right|^{p-1}\right\|_{p/(p-1)}=
# \left(\int_a^b \left|\varphi(x)\right|^p\,dx\right)^{(p-1)/p}=
# (\|\varphi\|_p)^{p-1}.
# $$
#
# これを $\varphi=f+g$ に適用することによって,
#
# $$
# \begin{aligned}
# (\|f+g\|_p)^p &=
# \int_a^b \left|f(x)+g(x)\right|^p\,dx = \int_a^b \left|f(x)+g(x)\right|\left|f(x)+g(x)\right|^{p-1}\,dx
# \\ &\leqq
# \int_a^b \left|f(x)\right| \left|f(x)+g(x)\right|^{p-1}\,dx +
# \int_a^b \left|g(x)\right| \left|f(x)+g(x)\right|^{p-1}\,dx
# \\ &\leqq
# \|f\|_p \left\|\left|f(x)+g(x)\right|^{p-1}\right\|_q +
# \|g\|_p \left\|\left|f(x)+g(x)\right|^{p-1}\right\|_q
# \\ &=
# \|f\|_p (\|f(x)+g(x)\|_p)^{p-1} +
# \|g\|_p (\|f(x)+g(x)\|_p)^{p-1}
# \\ &=
# (\|f\|_p+\|g\|_p) (\|f(x)+g(x)\|_p)^{p-1}
# \end{aligned}
# $$
#
# ここで, 2行目の不等号で三角不等式 $|f(x)+g(x)|\leqq |f(x)|+|g(x)|$ を使い, 3行目の不等号でHölderの不等式を使った. したがって,
#
# $$
# \|f+g\|_p \leqq \|f\|_p+\|g\|_p.
# $$
#
# これを**Minkowskiの不等式**と呼ぶ.
# **定義:** $\|f\|_p<\infty$ すなわち $\ds \int_a^b |f(x)|^p\,dx<\infty$ を満たす函数を **$L^p$ 函数** もしくは **$p$ 乗可積分函数** と呼ぶ. 特に $L^1$ 函数を単に**可積分函数**と呼ぶ. $\QED$
#
# Minkowskiの不等式より, $L^p$ 函数の和も $L^p$ 函数になる.
| Old_Ver_for_Julia_v0.6/08 convexity.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Unsupervised Transformations of Data:
# - Principle Component Analysis (PCA)
# - Non-negative Matrix Factorization (NMF)
#
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
from scipy import stats
import warnings
warnings.filterwarnings("ignore")
# -
from sklearn.datasets import fetch_lfw_people
#people = fetch_lfw_people()
people = fetch_lfw_people(min_faces_per_person=20, resize=0.7)
people.keys()
people['images'].shape
X=pd.DataFrame(people['data'])
y=pd.DataFrame(people['target'], columns=["image"])
df=pd.concat([y,X], axis=1)
# +
fig, axes = plt.subplots(2, 5, figsize=(12, 5))
for ax, i in zip(axes.ravel(), np.arange(0, 10)):
ax.matshow(np.array(X)[i,:].reshape(87,65))
ax.set_xticks([])
ax.set_yticks([])
# -
# Re-scaling of data:
from sklearn.preprocessing import StandardScaler, MinMaxScaler
scaler=MinMaxScaler().fit(X)
X_scaled=scaler.transform(X)
X.shape
# +
# PCA: ___________________________________________________________________
from sklearn.decomposition import PCA
pca=PCA(n_components=100, whiten=True, random_state=42)
pca.fit(X_scaled)
X_pca=pca.transform(X_scaled)
# +
eigen_vectors=pca.components_[0:10]
fix, axes = plt.subplots(2, 5, figsize=(15, 10), subplot_kw={'xticks': (), 'yticks': ()})
for i, (component, ax) in enumerate(zip(pca.components_[0:10], axes.ravel())):
ax.matshow(eigen_vectors[i,:].reshape(87, 65), cmap='viridis')
ax.set_title("{}th Prin. Component \n".format((i + 1)))
ax.set_xticks([])
ax.set_yticks([])
plt.tight_layout
# -
X_back_scaled=pca.inverse_transform(X_pca)
X_back=scaler.inverse_transform(X_back_scaled)
# +
fig, axes = plt.subplots(2, 5, figsize=(12, 5))
for ax, i in zip(axes.ravel(), np.arange(0, 10)):
ax.matshow(np.array(X_back)[i,:].reshape(87,65))
ax.set_xticks([])
ax.set_yticks([])
# +
# NMF: _________________________________________________________________
from sklearn.decomposition import NMF
nmf = NMF(n_components=20, random_state=42)
# use fit_transform instead of fit, as TSNE has no transform method
X_nmf = nmf.fit_transform(X_scaled)
# -
X_nmf.shape
# +
eigen_vectors=nmf.components_[0:10]
fix, axes = plt.subplots(2, 5, figsize=(15, 10), subplot_kw={'xticks': (), 'yticks': ()})
for i, (component, ax) in enumerate(zip(eigen_vectors, axes.ravel())):
ax.matshow(eigen_vectors[i,:].reshape(87, 65), cmap='viridis')
ax.set_title("{}th NMF Component \n".format((i + 1)))
ax.set_xticks([])
ax.set_yticks([])
plt.tight_layout
# -
X_back_scaled=nmf.inverse_transform(X_nmf)
X_back=scaler.inverse_transform(X_back_scaled)
# +
fig, axes = plt.subplots(2, 5, figsize=(12, 5))
for ax, i in zip(axes.ravel(), np.arange(0, 10)):
ax.matshow(np.array(X_back)[i,:].reshape(87,65))
ax.set_xticks([])
ax.set_yticks([])
# -
| Projects in Python with Scikit-Learn- XGBoost- Pandas- Statsmodels- etc./Face Recognition dataset (transformation with PCA and NMF).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.1 64-bit (''blog'': conda)'
# name: python371jvsc74a57bd016f988c202c6d5b67fae213fcd58a128d4933d9f6b0a0be940c9d00563b3fdfe
# ---
# # Lesson learnt from Kaggle - Bengali Image Classification Competition
# > My first silver medal on Kaggles Competition - Bengali Image Classification Competition. 10 lessons learn!
# - toc: true
# - branch: master
# - badges: true
# - comments: true
# - categories: [python, kaggle]
# - image: images/bengali_00_header.png
# I have teamed up with a friend to participate in the [Bengali Image Classification Competition](https://www.kaggle.com/c/bengaliai-cv19/?utm_medium=email&utm_source=intercom&utm_campaign=bengaliai-email-launch). We struggled to get a high rank in the Public leaderboard throughout the competition. In the end, the result is a big surprise to everyone as the leaderboard shook a lot.
# 
# 
# The final private score was much lower than the public score. It suggests that most participants are over-fitting Public leaderboard.
# # The Classification Task
# This is an image classification competition. We need to predict 3 parts of __Bengali__ characters `root`, `consonant` and `vowel`. It is a typical classification tasks like the __MNIST__ dataset.
# 
# ## Evaluation Metrics
# The competition use macro-recall as the evaluation metric. In general, people get >96% recall in training, the tops are even getting >99% recall.
# +
# collapse-hide
python
import numpy as np
import sklearn.metrics
scores = []
for component in ['grapheme_root', 'consonant_diacritic', 'vowel_diacritic']:
y_true_subset = solution[solution[component] == component]['target'].values
y_pred_subset = submission[submission[component] == component]['target'].values
scores.append(sklearn.metrics.recall_score(
y_true_subset, y_pred_subset, average='macro'))
final_score = np.average(scores, weights=[2,1,1])
# -
# ## Model (Bigger still better)
# We start with `xresnet50`, which is a relatively small model. As we have the assumption that this classification task is a very standard task, therefore the difference of model will not be the most important one. Thus we pick xresnet50 as it has a good performance in terms of accuracy and train relatively fast.
#
# Near the end of the competition, we switch to a larger model `se-resnext101`. It requires triple training time plus we have to scale down the batch size as it does not fit into the GPU memory. Surprisingly (maybe not surprising to everyone), the bigger model did boost the performance more than I expected with \~0.3-0.5% recall. It is a big improvement as the recall is very high (\~0.97), in other words, it reduces __\~10%__ error solely by just using a better model, not bad!
# ## Augmentation
# There are never "enough" data for deep learning, so we always try our best to collect more data. Since we cannot collect more data, we need data augmentation.
# We start with rotation + scale. We also find __MixUp__ and __CutMix__ is very effective to boost the performance. It also gives us roughly __10%__ boost initially from 0.96 -> 0.964 recall.
# ### [CutMix](https://arxiv.org/abs/1905.04899) & [MixUp](https://arxiv.org/pdf/1710.09412.pdf)
# 
# _Mixup_ is simple, if you know about photography, it is similar to have double exposure of your photos. It overlays two images (cat+dog in this case) by sampling weights. So instead of prediction P(dog) = 1, the new target could become P(dog) = 0.8 and P(cat) = 0.2.
#
# _CutMix_ shares a similar idea, instead of overlay 2 images, it crops out a certain ratio of the image and replaces it with another one.
#
# It always surprises me that these augmented data does not make much sense to a human, but it is very effective to improve model accuracy and reduce overfitting empirically.
# # Logging of Experiment
# I normally just log my experiment with a simple CSV and some printing message. This start to get tedious when there are more than 1 people to work. It is important to communicate the results of experiments. I explore `Hydra` and `wandb` in this competition and they are very useful.
# ## [Hydra](https://hydra.cc/)
# 
#
# It is often a good idea to make your experiment configurable. We use `Hydra` for this purpose and it is useful to compose different configuration group.
# By making your hyper-paramters configurable, you can define an experiment by configuration files and run multiple experiments. By logging the configuration with the training statistics, it is easy to do cross-models comparison and find out which configuration is useful for your model.
#
# I have written [an short example](https://mediumnok.ml/coding/ml/2020/02/08/Config-Composition-with-Hydra-for-Machine-Learning-Experiments.html) for how to use __Hydra__.
# ## [Wandb](https://www.wandb.com/)
# __wandb__ (Weight & Biases) does a few things. It provides built-in functions that automatically log all your model statistics, you can also log your custom metrics with simple functions.
#
# * Compare the configuration of different experiments to find out the model with the best performance.
# * Built-in function for logging model weights and gradient for debugging purpose.
# * Log any metrics that you want
#
# All of these combined to make collaboration experience better. It is really important to sync the progress frequently and getting everyone results in a single platform makes these conversations easier.
#
# 
# # [Stochastic Weight Averaging](https://pytorch.org/blog/stochastic-weight-averaging-in-pytorch/)
# This is a simple yet effective technique which gives about 0.3-0.4% boost to my model. In simple words, it takes __snapshots__ of the model weights during training and takes an average at the end. It provides a cheap way to do models ensemble while you are only training 1 model. This is important for this competition as it allows me to keep training time short enough to allow feedback within hours and reduce over-fitting.)
#
#
#
# # 
# # Larger is better (image size)
# We downsample our image size to 128x128 throughout the competition, as it makes the model train faster and we believe most technique should be transferable to larger image size. It is important to keep your feedback loop short enough (hours if not days). You want your training data as small as possible while keeping them transferable to your full dataset. Once we scale our image to full size, it takes almost 20 hours to train a single model, and we only have little chance to tune the hyper-parameters before the competition end.
# # Debug & Checkpoint
# There was a time we develop our model separately and we didn't sync our code for a while. We refactor our code during the time and it was a huge mistake. It turns out our pre-refactor code trains much better model and we introduce some unknown bug. It is almost impossible to find out as we change multiple things. It is so hard to debug a neural network and testing it thoroughly is important. Injecting a large amount of code may help you to run an experiment earlier, but you may pay much more time to debug it afterwards.
#
# I think this is applicable even if you are working alone.
# * Keep your changes small.
# * Establish a baseline early, always do a regression test after a new feature introduced (especially after code refactoring)
# * Create checkpoint to rollback anytime, especially if you are not working on it every day.
#
# Implementation is the key of Kaggle competition (in real life too). It does not matter how great your model is, a tiny little bug could have damaged your model silently
#
# # Use auxiliary label
# 
# As mentioned earlier, this competition requires to predict the `root`, `vowel` and the `consonant` part. In the training data, they actually provide the `grapheme` too. Lots of people saying that if you train with the `grapheme`, it improves the model greatly and get the recall >98% easily.
#
# This is something we could not reproduce throughout the competition, we tried it in the very last minute but it does not seem to improve our model. It turns out lots of people are overfitting the data, as the testing dataset has much more unseen character.
#
# But it is still a great remark that training with labels that is not your final desired output could still be very useful.
# # Weight loss
# The distribution of the training dataset is very imbalance, but to get a good result, we need to predict every single class accurately (macro recall). To deal with this issue, we choose to use class weights, where a higher weight would be applied to rare samples. We don't have an ablation study for this, but it seems to help close the gap between accuracy & recall and allows us to train the model slightly better.
# # Find a teammate!
# Lastly, please go and find a teammate if you can. It is very common to start a Kaggle competition, but not so easy to finish them. I have stopped for a month during the competition due to my job. It is really hard to get back to the competition after you stopped for so long. Getting a teammate helps to motivate you and in the end, it is a great learning experience for both of us.
# # Pretrain Model
# We also tried to use a pretrained model, as it allows shorter training and gives better performance by transfer learning (Using weights learn from a large dataset to as initial weight). It also gives our model a bit of improvement.
#
# * Finetune the model head, while keeping other layers freeze (except BatchNorm layer).
# * Unfreeze the model, train all the layers together.
#
# I also tried training the model directly with discriminating learning rate while not freezing any layer at all. It performs similarly to freezing fine-tuning , so I end up just start training the entire model from the beginning.
# # If the code works, don't touch it
# This is probably not a good habit usually, but I suggest not to do it for a competition. We spent lots of time for debugging our code after code refactoring and end up just rolling back to an older commit and cherry-picks new features. In a competition, you don't have enough time to test everything. You do not need a nice abstract class for all your features, some refactoring to keep your function/class clean is probably needed, but do not overspend your time on it. It is even common to jump between frameworks (you may find other's Kernel useful), so it is not possible to structure your code perfectly.
#
# * If someone has create a working submission script, use it!
# * If someone has create a working pre-processing function, use it!
#
# Don't spend time on trying to optimize these code unless it is necessary, it is often not worth it in a competition context. You should focus on adding new features, trying out new model, testing with new augmentation technique instead.
# # Summary
# This is a great learning experience and refreshes some of my outdated computer vision model knowledge. If you have never joined a competition, find a friend and get started. If you have just finished one, try writing it out and share your experience. 😉
| _notebooks/2020-03-21-10-lessons-learnt-from-Kaggle-competition.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:bus_number]
# language: python
# name: conda-env-bus_number-py
# ---
from src import workflow
from src.paths import summary_path
# # I'm Only Here for the Pretty Pictures
#
# ## Step 5: Analyze and Summarize
#
# An **Analysis** takes Datasets (the predictions or transformed datasets obtained from the last step), and produces data (e.g. Pandas DataFrame, CSV). This is how we would generate tables in a paper, for example, or data to be consumed by a graphing function.
#
# <img alt="" src="references/workflow/make-analyze.png" width=500/>
# In Bjørn's case, he have a target vector associated with the original dataset, so hecan compare his prediction performance against that target. We will use a **scoring function** to compare these vectors.
# Like in sklearn, A **scoring function** is a function with the signature:
# `score_func(y, y_pred, **kwargs)`
#
workflow.available_scorers()
# ## TODO: Add all of this to the standard workflow
workflow.available_analyses()
help(workflow.available_analyses)
workflow.available_predictions()
workflow.add_analysis(analysis_name='score_predictions')
workflow.get_analysis_list()
workflow.make_analysis()
# !cd .. && make analysis
# !cat ../reports/analyses.json
# ### EXERCISE: Implement `available_results()`
# ## Look at the Summary Statistics
import pandas as pd
df = pd.read_csv(summary_path / 'score_predictions.csv')
df
# # Add other algorithms
#
# Comparing one algorithm isn't that interesting. Let's add a couple more so Bjørn can compare them.
# ### Exercise: Add a GradientBoostingClassifier
# Modify `src/models/algorithms.py` so that the next cell works.
#
# Hint: ```sklearn.ensemble.GradientBoostingClassifier```
workflow.add_model(
dataset_name = 'lvq-pak_train',
algorithm_name = 'GradientBoostingClassifier',
algorithm_params = {'random_state': 42}
)
# ### Exercise: Add your choice of classifier here
workflow.add_model(
dataset_name = 'lvq-pak_train',
## add your algorithm here!
)
### Take a look to see what's there
workflow.get_model_list()
workflow.available_algorithms(keys_only=False)
workflow.make_train()
workflow.available_models()
workflow.get_prediction_list()
## Set up predictions using all of the available models
for model in workflow.available_models():
workflow.add_prediction(
dataset_name = 'lvq-pak_test',
model_name = model,
is_supervised = True,
)
workflow.get_prediction_list()
workflow.make_predict()
workflow.available_predictions()
# The default for running the the summary df is to run on all available predictions. We have nothing more that we have to add to our existing script to get all the new scores.
# !cd .. && make analysis
# ## Look at summary statistics
df = pd.read_csv(summary_path / 'score_predictions.csv')
df
# ## Make Publish
# <img alt="" src="references/workflow/make-publish.png" width=500 />
# ### Publishing Data:
# A DOI is a commonly used **digital object identifier**, and can be used to publish a dataset.
#
# There are a few easy ways to get a DOI for your work:
# * [Figshare](http://figshare.com/) will provide a DOI for any virtually any digital work, which includes data.
# * [Zenodo](https://zenodo.org/) provides DOIs research output, which may include datasets
# We aren't going to take you through the publishing process here
| notebooks/40-bjorn-analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#Importing libraries
import pandas as pd
#Visualization Libraries
import matplotlib.pyplot as plt
# %matplotlib inline
import seaborn as sns
import plotly.express as px
import plotly.graph_objects as go
import folium
from folium import plugins
#Manipulating the default plot
plt.rcParams['figure.figsize'] = 10,12
#Disable warnings
import warnings
warnings.filterwarnings('ignore')
# -
#Learn how to read datasets
df=pd.read_excel("covid_19_india.xlsx", parse_dates=True, sheet_name='mofhw')
df_india = df.copy()
df
#df.drop(['S. No.'], axis=1, inplace=True)
df['Total cases'] = df['Total Confirmed cases (Including 76 foreign Nationals)']
total_cases = df['Total cases'].sum()
print('Total number of confirmed Corvid 19 cases in India till date:', total_cases)
df.style.background_gradient(cmap = 'Reds')
#Total Active number of cases = Total cases - (Deaths + cured)
df['Total Active'] = df['Total Confirmed cases (Including 76 foreign Nationals)'] - (df['Death'] + df['Cured/Discharged/migrated'])
total_active = df['Total Active'].sum()
print('Total number of active Covid 19 cases across India: ', total_active)
Tot_Cases = df.groupby('Name of State / UT')['Total Active'].sum().sort_values(ascending=False).to_frame()
Tot_Cases.style.background_gradient(cmap = 'Reds')
# +
#Confirmed vs Recovered figures
#Visualization using Seaborn
India_coord = pd.read_excel("Indian Coordinates.xlsx")
df_full = pd.merge(India_coord,df, on = 'Name of State / UT')
f, ax = plt.subplots(figsize=(12,8))
data = df_full[['Name of State / UT', 'Total cases', 'Cured/Discharged/migrated', 'Death']]
data.sort_values('Total cases', ascending = False, inplace =True)
sns.set_color_codes("pastel")
sns.barplot(x='Total cases', y='Name of State / UT', data = data, label='Total', color='r')
sns.set_color_codes('muted')
sns.barplot(x='Cured/Discharged/migrated', y = 'Name of State / UT', data=data, label ='cured', color='g')
#Add a legend and informative axis label
ax.legend(ncol=2, loc='lower right', frameon = 'True')
ax.set(ylabel = '', xlabel = 'Cases')
sns.despine(left=True, bottom=True)
# -
#Rise in Corvid cases
import plotly
plotly.io.renderers.default = 'colab'
# +
#creating interactive graphs using plotly
import plotly.graph_objects as go
dbd_India = pd.read_excel('covid_19_india.xlsx', parse_dates=True, sheet_name='Cleaned')
fig = go.Figure()
fig.add_trace(go.Scatter(x=dbd_India['Date'], y=dbd_India['Total cases'], mode='lines+markers', name='Total cases'))
fig.update_layout(title_text='Trend of Corvid 19 Cases in India (Cumulative Cases)', plot_bgcolor='rgb(230, 230, 230)')
fig.show()
# -
#Predicting with FB Prophet
from fbprophet import Prophet as pr
df = pd.read_excel('covid_19_india.xlsx', parse_dates=['Date'], sheet_name='Confirmed')
#df['Date'] = pd.to_datetime(df['Date'])
df
df.groupby('Date')
df['Date'] = pd.to_datetime(df['Date'])
df.columns = ['ds','y']
df.tail()
#confirmed = df.groupby('ds')
#confirmed.columns = ['ds','y']
#confirmed.head()
# +
#confirmed1 = []
#for i in confirmed:
# confirmed1.append(i)
#confirmed = confirmed1
#confirmed['Date'] = pd.to_datetime(confirmed['Date'])
#confirmed.head()
#type(confirmed)
# -
m = pr(interval_width=0.95)
m.fit(df)
future = m.make_future_dataframe(periods = 11)
future.tail()
#Predicting the future with date and upper/lower limits of y value
forecast = m.predict(future)
forecast[['ds', 'yhat', 'yhat_lower', 'yhat_upper']].tail(10)
confirmed_forecast_plot = m.plot(forecast)
confirmed_forecast_plot = m.plot_components(forecast)
#Predicting with FB Prophet -- since march 15th
from fbprophet import Prophet as pr
df = pd.read_excel('covid_19_india.xlsx', parse_dates=['Date'], sheet_name='Confirmed_march')
df
df.groupby('Date')
df['Date'] = pd.to_datetime(df['Date'])
df.columns = ['ds','y']
df.tail()
m = pr(interval_width=0.95)
m.fit(df)
future = m.make_future_dataframe(periods = 29)
future.tail()
#Predicting the future with date and upper/lower limits of y value
forecast = m.predict(future)
forecast[['ds', 'yhat', 'yhat_lower', 'yhat_upper']].tail(10)
confirmed_forecast_plot = m.plot(forecast)
confirmed_forecast_plot = m.plot_components(forecast)
| Covid_today_pred.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="UiuP7Z3-bvGO"
# # Setup
# + [markdown] id="Fk2p5dLrpJgM"
# ## Import libraries
# + id="9rUAnl_Ma0xC"
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
import torchvision
from torchvision import transforms
from torchvision import datasets
from torch.utils.data import DataLoader
import random
random.seed(123)
import time
import os
# + colab={"base_uri": "https://localhost:8080/"} id="GBiHNZuWyEGM" executionInfo={"status": "ok", "timestamp": 1611672538687, "user_tz": -60, "elapsed": 23544, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14516834551380015257"}} outputId="edc36d56-38fc-4ab6-b04c-7b8a26a6f245"
from google.colab import drive
drive.mount('/content/drive')
# + [markdown] id="JaSjiw-cpNkt"
# ## Check CUDA version
# + colab={"base_uri": "https://localhost:8080/"} id="CJeUYjy8bWfk" executionInfo={"status": "ok", "timestamp": 1611672538689, "user_tz": -60, "elapsed": 23530, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14516834551380015257"}} outputId="12e588ec-6bbd-44ef-f8c3-073483f7272e"
use_cuda = True
if use_cuda and torch.cuda.is_available():
device = torch.device('cuda')
else:
device = torch.device('cpu')
device
# + [markdown] id="L1hOXGm9pRwk"
# ## Visualisation functions
# + id="DrRTDEo_bZu0"
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
# Function to show an image tensor
def show(X):
if X.dim() == 3 and X.size(2) == 3:
plt.imshow(X.numpy())
#plt.show()
elif X.dim() == 2:
plt.imshow( X.numpy() , cmap='gray' )
#plt.show()
else:
print('WRONG TENSOR SIZE')
def show_saliency(X):
if X.dim() == 3 and X.size(2) == 3:
plt.imshow(X.numpy())
plt.show()
elif X.dim() == 2:
plt.imshow( X.numpy() , cmap='viridis' )
plt.show()
else:
print('WRONG TENSOR SIZE')
# + [markdown] id="PJf94wVPpUoB"
# ## Download dataset
# + id="cxOP14uSc_7E" colab={"base_uri": "https://localhost:8080/", "height": 403, "referenced_widgets": ["e9dcd1856e4c4b72b41345d4927055c3", "68cb6654fe654b3fad14a5d68ed5f8dc", "39b958bb90fa48b6a8af7392e611266e", "9e625083ddc24cd79d4feb34da4684bb", "aa4304049745445bab8c18846ba0de2d", "<KEY>", "824f811f51b4417ca9c78e04262632bb", "bd93a8f22bb047bd95e6ab71e4183d36", "749d5e56be3a4b2aa98d9ace2160db16", "00d047f1a4c148c49d776fffed780c08", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "386c3caf896a4d8f8dcd5e6749342c8c", "<KEY>", "<KEY>", "<KEY>", "6fbea0e9543743e587e69dbb1f866588", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "2bf226e958334d6e99b0ad3101b29ace", "82d6252c8f114f93a46cac6207a177fb", "aa1475b7eed34ff6b9360434a41c4420", "<KEY>"]} executionInfo={"status": "ok", "timestamp": 1611672552279, "user_tz": -60, "elapsed": 3779, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14516834551380015257"}} outputId="0947906c-1d5e-4f0b-aa2a-a77e77e26f89"
transform = transforms.Compose([transforms.ToTensor(),
transforms.Lambda(lambda x: x.squeeze()), # Squeeze the data to remove the redundant channel dimension
])
trainset = torchvision.datasets.FashionMNIST(root='./data_FashionMNIST',
train=True,
download=True,
transform=transform
)
testset = torchvision.datasets.FashionMNIST(root='./data_FashionMNIST',
train=False,
download=True,
transform=transform
)
classes = (
'T-shirt/top',
'Trouser',
'Pullover',
'Dress',
'Coat',
'Sandal',
'Shirt',
'Sneaker',
'Bag',
'Ankle boot',
)
# + [markdown] id="H4mgdlRkp6L2"
# #Data preprocessing
# + [markdown] id="Ch1u5-x3p8eq"
# ## Split training data in train and validation data
# + id="bIc6MGQ_dES8" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1611672557076, "user_tz": -60, "elapsed": 1520, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14516834551380015257"}} outputId="d08a10be-3b11-4f37-82a6-832c6445491c"
from sklearn.model_selection import train_test_split
targets = trainset.targets
train_idx, val_idx= train_test_split(np.arange(len(targets)),test_size=0.2,shuffle=True, stratify=targets, random_state=123)
train_sampler = torch.utils.data.SubsetRandomSampler(train_idx)
val_sampler = torch.utils.data.SubsetRandomSampler(val_idx)
batch_size=128
trainloader = torch.utils.data.DataLoader(trainset, batch_size=batch_size, sampler=train_sampler)
valloader = torch.utils.data.DataLoader(trainset, batch_size=batch_size, sampler=val_sampler)
testloader = torch.utils.data.DataLoader(testset,
batch_size=batch_size,
shuffle=True,
drop_last=True
)
# + [markdown] id="Y7scuHVpqCY6"
# #Model architecture
# + [markdown] id="dEn1I0koqIGP"
# ##Create the model
# + id="kEqjg3jobdBX"
class LeNet(nn.Module):
def __init__(self, kernel_size, pool_function, nfilters_conv1, nfilters_conv2):
super(LeNet, self).__init__()
self.nfilters_conv2 = nfilters_conv2
# CL1: 1 x 28 x 28 (grayscale) --> nfilters_conv1 x 28 x 28
self.conv1 = nn.Conv2d(1, nfilters_conv1, kernel_size=kernel_size, padding=kernel_size//2)
# MP1: nfilters_conv1 x 28 x 28 --> nfilters_conv1 x 14 x 14
self.pool1 = pool_function(2,2)
# CL2: nfilters_conv1 x 14 x 14 --> nfilters_conv2 x 14 x 14
self.conv2 = nn.Conv2d(nfilters_conv1, nfilters_conv2, kernel_size=kernel_size, padding=kernel_size//2)
# MP2: nfilters_conv2 x 14 x 14 --> nfilters_conv2 x 7 x 7
self.pool2 = pool_function(2,2)
# LL1: nfilters_conv2 x 7 x 7 --> 100
self.linear1 = nn.Linear((nfilters_conv2*7*7), 100)
# LL2: 100 --> 10
self.linear2 = nn.Linear(100,10)
def forward(self, x):
x = x.unsqueeze(1)
# CL1:
x = self.conv1(x)
x = F.relu(x)
# MP1:
x = self.pool1(x)
# CL2:
x = self.conv2(x)
x = F.relu(x)
# MP2:
x = self.pool2(x)
# LL1:
x = x.view(-1, self.nfilters_conv2*7*7)
x = self.linear1(x)
x = F.relu(x)
# LL2:
x = self.linear2(x)
return x
# + id="V_Be5wuG0D_N"
# best results from hyperparameter tuning
kernel_size= 5
pool_function = nn.AvgPool2d
nfilters_conv1 = 128
nfilters_conv2 = 128
model_adv = LeNet(kernel_size=kernel_size,pool_function=pool_function,nfilters_conv1=nfilters_conv1,nfilters_conv2=nfilters_conv2).to(device)
criterion = nn.CrossEntropyLoss()
my_lr=0.01
optimizer=torch.optim.Adam(model_adv.parameters(), lr=my_lr) # change here
# + colab={"base_uri": "https://localhost:8080/"} id="OFMuWfPbbLGS" executionInfo={"status": "ok", "timestamp": 1611672570973, "user_tz": -60, "elapsed": 6056, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14516834551380015257"}} outputId="a739422a-2701-4617-80e4-8f99b9c1d926"
print(model_adv)
# + [markdown] id="7TjdOEDdb36K"
# # Attack!
# + [markdown] id="LXFrVVRYqPe4"
# ##Import libraries
# + colab={"base_uri": "https://localhost:8080/"} id="YLsi9F6Nbpqq" executionInfo={"status": "ok", "timestamp": 1611672586100, "user_tz": -60, "elapsed": 5753, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14516834551380015257"}} outputId="4a235b99-e00f-4c4f-cade-ec5747304746"
# !pip install advertorch
# + id="hNhbmqREb_ON"
from advertorch.attacks import PGDAttack
# + [markdown] id="NNYQPO5qqSFG"
# ##Create adversary
# + id="w08vXgmfcE3g"
# prepare your pytorch model as "model"
# prepare a batch of data and label as "cln_data" and "true_label"
# prepare attack instance
adversary = PGDAttack(
model_adv, loss_fn=nn.CrossEntropyLoss(), eps=0.3,
nb_iter=10, eps_iter=0.01, rand_init=True, clip_min=0.0, clip_max=1.0,
targeted=False)
# + id="7h2G2yoJLCF6"
plot_valloss = []
# + [markdown] id="X7yZP-Z4qV1B"
# ##Train model
# + colab={"base_uri": "https://localhost:8080/"} id="UaYh5LW-zMIB" executionInfo={"status": "ok", "timestamp": 1611675768906, "user_tz": -60, "elapsed": 3154664, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14516834551380015257"}} outputId="16dcf632-56b4-4ef9-8e39-a596de329b78"
start=time.time()
min_loss = 20 #initial loss to be overwritten
epochs_no_improve = 0
patience = 20 # high patience to overcome local minima
for epoch in range(1,200):
model_adv.train()
for i, (x_batch, y_batch) in enumerate(trainloader):
x_batch, y_batch = x_batch.to(device), y_batch.to(device) # Move the data to the device that is used
x_adv = adversary.perturb(x_batch, y_batch)
optimizer.zero_grad() # Set all currenly stored gradients to zero
y_pred = model_adv(x_adv)
loss = criterion(y_pred, y_batch)
loss.backward()
optimizer.step()
# Compute relevant metrics
y_pred_max = torch.argmax(y_pred, dim=1) # Get the labels with highest output probability
correct = torch.sum(torch.eq(y_pred_max, y_batch)).item() # Count how many are equal to the true labels
elapsed = time.time() - start # Keep track of how much time has elapsed
# Show progress every 50 batches
if not i % 50:
print(f'epoch: {epoch}, time: {elapsed:.3f}s, loss: {loss.item():.3f}, train accuracy: {correct / batch_size:.3f}')
model_adv.eval()
val_loss = 0
counter = 0
for i, (x_batch, y_batch) in enumerate(valloader):
counter += 1
x_batch, y_batch = x_batch.to(device), y_batch.to(device) # Move the data to the device that is used
y_pred = model_adv(x_batch)
val_loss += criterion(y_pred, y_batch).item()
val_loss = val_loss/counter
print(f'epoch: {epoch}, validation loss: {val_loss}')
plot_valloss.append([val_loss, epoch])
# save the model
if val_loss < min_loss:
torch.save(model_adv, "/content/drive/MyDrive/Deep Learning/Project/model_adv.pckl")
epochs_no_improve = 0
min_loss = val_loss
else:
epochs_no_improve += 1
if epochs_no_improve == patience and epoch > 5:
print("Early Stopping!")
break
# + [markdown] id="i2Pq0dZlqZKs"
# ##Show validation loss
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="TRX7upMcLKWc" executionInfo={"status": "ok", "timestamp": 1611675774232, "user_tz": -60, "elapsed": 1081, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14516834551380015257"}} outputId="61e6d0b1-339f-4561-d38f-ab5f5bc744e5"
import pandas as pd
df_plot = pd.DataFrame(data = plot_valloss, columns=['Validation Loss', 'Epoch'])
df_plot.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="pENsVaLQLMwQ" executionInfo={"status": "ok", "timestamp": 1611675780266, "user_tz": -60, "elapsed": 3341, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14516834551380015257"}} outputId="9630bffa-0409-48d5-c115-3ee73053a253"
import plotly.express as px
fig = px.line(df_plot, x="Epoch", y="Validation Loss", title='Validation Loss FAT')
fig.show()
# + id="-HFiqBAILPhk"
df_plot.to_csv("/content/drive/MyDrive/Deep Learning/Project/FAT_valloss.csv", index=False)
# + [markdown] id="99DOApupLRim"
# # Testing
# + [markdown] id="UsCjHnTkqiQA"
# ## Test model on clean data
# + id="UMjdZN2qLUmr"
model_adv_inf = torch.load("/content/drive/MyDrive/Deep Learning/Project/model_adv.pckl")
# + colab={"base_uri": "https://localhost:8080/"} id="OwExvF2hmDNT" executionInfo={"status": "ok", "timestamp": 1611675807388, "user_tz": -60, "elapsed": 1787, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14516834551380015257"}} outputId="4cd03aa2-4837-410b-c303-2c489b6c65da"
correct_total = 0
for i, (x_batch, y_batch) in enumerate(testloader):
x_batch, y_batch = x_batch.to(device), y_batch.to(device) # Move the data to the device that is used
y_pred = model_adv_inf(x_batch)
y_pred_max = torch.argmax(y_pred, dim=1)
correct_total += torch.sum(torch.eq(y_pred_max, y_batch)).item()
print(f'Accuracy on the test set: {correct_total / len(testset):.3f}')
# + colab={"base_uri": "https://localhost:8080/"} id="cnk_BmqXLaMh" executionInfo={"status": "ok", "timestamp": 1611675817636, "user_tz": -60, "elapsed": 710, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14516834551380015257"}} outputId="e07ac2b2-0a97-4ea0-9b10-0544adb7c94f"
accuracy = correct_total / len(testset)
z = 1.96 #for 95% CI
n = len(testset)
interval = z * np.sqrt( (accuracy * (1 - accuracy)) / n)
interval
# + [markdown] id="V7mUPLjMnIP7"
# ## Test model on perturbed data
# + id="YdcWcFrQozXp"
import pandas as pd
import seaborn as sn
from advertorch.utils import predict_from_logits
# + colab={"base_uri": "https://localhost:8080/"} id="egqg9rVDgh5p" executionInfo={"status": "ok", "timestamp": 1611675837659, "user_tz": -60, "elapsed": 10748, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14516834551380015257"}} outputId="8ea77621-e0e5-4a46-dbfd-bfac2f9a0c18"
correct_total = 0
all_preds = []
y_true = []
for i, (x_batch, y_batch) in enumerate(testloader):
x_batch, y_batch = x_batch.to(device), y_batch.to(device) # Move the data to the device that is used
y_true.extend(y_batch)
adv = adversary.perturb(x_batch, y_batch)
y_adv_pred = predict_from_logits(model_adv_inf(adv))
all_preds.extend(y_adv_pred)
correct_total += torch.sum(torch.eq(y_adv_pred, y_batch)).item()
print(f'Accuracy on the test set: {correct_total / len(testset):.3f}')
# + colab={"base_uri": "https://localhost:8080/"} id="UtyxFDXpoNsS" executionInfo={"status": "ok", "timestamp": 1611675855114, "user_tz": -60, "elapsed": 728, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14516834551380015257"}} outputId="6a028d80-7d25-4864-ed88-8990a7d4a37a"
accuracy = correct_total / len(testset)
z = 1.96 #for 95% CI
n = len(all_preds)
interval = z * np.sqrt( (accuracy * (1 - accuracy)) / n)
interval
# + [markdown] id="m5ksqySSqtJ4"
# ## Visualise results
# + id="XDEt7CDjswjq"
y_true_int = [int(x.cpu()) for x in y_true]
y_pred_int = [int(x.cpu()) for x in all_preds]
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="VrN3yMS5pzFY" executionInfo={"status": "ok", "timestamp": 1611675866093, "user_tz": -60, "elapsed": 510, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14516834551380015257"}} outputId="f0d6d6f2-b243-4a30-9f7e-69f3123a0ee3"
data = {'y_Actual': y_true_int,
'y_Predicted': y_pred_int
}
cm_df = pd.DataFrame(data, columns=['y_Actual', 'y_Predicted'])
cm_df.head()
# + colab={"base_uri": "https://localhost:8080/"} id="IYApaz82qOkF" executionInfo={"status": "ok", "timestamp": 1611675868938, "user_tz": -60, "elapsed": 932, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14516834551380015257"}} outputId="e40dc66a-024a-4fd8-cd36-c3a28cd93fd3"
confusion_matrix = pd.crosstab(cm_df['y_Actual'], cm_df['y_Predicted'], rownames=['Actual'], colnames=['Predicted'])
print(confusion_matrix)
# + colab={"base_uri": "https://localhost:8080/", "height": 279} id="0bfPKtYNqQLo" executionInfo={"status": "ok", "timestamp": 1611675870385, "user_tz": -60, "elapsed": 927, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14516834551380015257"}} outputId="123c0e12-cf86-4581-9d8b-f8683c732a4a"
sn.heatmap(confusion_matrix, annot=False)
plt.show()
# + id="PGmaMzr2tajb"
| FAT.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6.4 64-bit
# name: python36464bitc2077ed07ea84d23aa5b518d224882ab
# ---
# 
# #### Modifiers
# +
import pandas as pd
df = pd.DataFrame()
df.info()
pd.
# -
x = 2 # Public -- Everywhere
_x = 2 # Protected -- Inside class or sub-class
__x = 2 # Private -- Inside class or file
def nombre():
# TODO
pass
# Ejemplo en Java
private Integer variable = 2
# +
class Humano:
def __init__(self):
pass
def __str__(self):
pass
#c++
# built-in
# + tags=[]
# Public
def public_function():
print("public")
# Protected
def _protected_function():
print("Protected")
# Private
def __private_function():
print("Private")
if __name__ == "__main__":
print("We are going to test the functions: \n\n\n\n")
print("--------------------------")
print("Testing public...")
public_function()
print("--------------------------")
print("Testing protected...")
_protected_function()
print("--------------------------")
print("Testing private...")
__private_function()
print("--------------------------")
# -
def __f_priv():
print("Soy priv")
# + tags=[]
class Human:
def __init__(self):
print("Hola")
# Public
def public_function(self):
print("public")
# Protected
def _protected_function(self):
print("Protected")
# Private
def __private_function(self):
print("Private")
def mi_funcion(self):
self.__private_function()
self.public_function()
self._protected_function()
if __name__ == "__main__":
p = Human()
print("We are going to test the functions: \n\n\n\n")
print("--------------------------")
print("Testing public...")
p.public_function()
print("--------------------------")
print("Testing protected...")
p._protected_function()
print("--------------------------")
print("Testing private...")
p.__private_function()
print("--------------------------")
# + tags=[]
p.mi_funcion()
# -
pd.
# + tags=[]
p.__init__()
# Built-in
# +
class Human1():
def __privado(self):
pass
k = Human1()
k.__privado()
# +
from ppp import __privada, _protegida
__privada()
# -
_protegida()
# +
from ppp import Clase
f = Clase()
f._protegida_dentro_clase()
# -
f = Clase()
f.__privada_dentro_clase()
# +
import numpy as np
np.__file__
| week5_EDA_scraping_eda_summary/day3_ppp_class_methods_SQL_III/theory/python/private_protected_public/private_protected_public.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (bayesian)
# language: python
# name: bayesian
# ---
# +
from __future__ import print_function
import os
import sys
import time
import argparse
import datetime
import math
import pickle
import torchvision
import torchvision.transforms as transforms
from utils.autoaugment import CIFAR10Policy
import torch
import torch.utils.data as data
import numpy as np
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
import torch.backends.cudnn as cudnn
from torch.autograd import Variable
from torchvision import datasets
from torch.utils.data.sampler import SubsetRandomSampler
# -
from utils.BBBlayers import GaussianVariationalInference
from utils.BayesianModels.Bayesian3Conv3FC import BBB3Conv3FC
from utils.BayesianModels.BayesianAlexNet import BBBAlexNet
from utils.BayesianModels.BayesianLeNet import BBBLeNet
from utils.BayesianModels.BayesianSqueezeNet import BBBSqueezeNet
net_type = 'alexnet'
dataset = 'CIFAR10'
outputs = 10
inputs = 3
resume = False
n_epochs = 30
lr = 0.001
weight_decay = 0.0005
num_samples = 1
beta_type = "Blundell"
resize=32
# Hyper Parameter settings
use_cuda = torch.cuda.is_available()
torch.cuda.set_device(0)
# number of subprocesses to use for data loading
num_workers = 0
# how many samples per batch to load
batch_size = 32
# percentage of training set to use as validation
valid_size = 0.2
# +
# convert data to a normalized torch.FloatTensor
transform = transforms.Compose([
transforms.RandomHorizontalFlip(), # randomly flip and rotate
transforms.RandomRotation(10),
transforms.ToTensor(),
# transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))
])
# choose the training and test datasets
train_data = datasets.CIFAR10('data', train=True,
download=True, transform=transform)
test_data = datasets.CIFAR10('data', train=False,
download=True, transform=transform)
# -
# obtain training indices that will be used for validation
num_train = len(train_data)
indices = list(range(num_train))
np.random.shuffle(indices)
split = int(np.floor(valid_size * num_train))
train_idx, valid_idx = indices[split:], indices[:split]
# define samplers for obtaining training and validation batches
train_sampler = SubsetRandomSampler(train_idx)
valid_sampler = SubsetRandomSampler(valid_idx)
# prepare data loaders (combine dataset and sampler)
train_loader = torch.utils.data.DataLoader(train_data, batch_size=batch_size,
sampler=train_sampler, num_workers=num_workers)
valid_loader = torch.utils.data.DataLoader(train_data, batch_size=batch_size,
sampler=valid_sampler, num_workers=num_workers)
test_loader = torch.utils.data.DataLoader(test_data, batch_size=batch_size,
num_workers=num_workers)
# specify the image classes
classes = ['airplane', 'automobile', 'bird', 'cat', 'deer',
'dog', 'frog', 'horse', 'ship', 'truck']
# +
import matplotlib.pyplot as plt
# %matplotlib inline
# helper function to un-normalize and display an image
def imshow(img):
img = img / 2 + 0.5 # unnormalize
plt.imshow(np.transpose(img, (1, 2, 0))) # convert from Tensor image
# +
# obtain one batch of training images
dataiter = iter(train_loader)
images, labels = dataiter.next()
images = images.numpy() # convert images to numpy for display
# plot the images in the batch, along with the corresponding labels
fig = plt.figure(figsize=(25, 4))
# display 20 images
for idx in np.arange(20):
ax = fig.add_subplot(2, 20/2, idx+1, xticks=[], yticks=[])
imshow(images[idx])
ax.set_title(classes[labels[idx]])
# +
rgb_img = np.squeeze(images[3])
channels = ['red channel', 'green channel', 'blue channel']
fig = plt.figure(figsize = (36, 36))
for idx in np.arange(rgb_img.shape[0]):
ax = fig.add_subplot(1, 3, idx + 1)
img = rgb_img[idx]
ax.imshow(img, cmap='gray')
ax.set_title(channels[idx])
width, height = img.shape
thresh = img.max()/2.5
for x in range(width):
for y in range(height):
val = round(img[x][y],2) if img[x][y] !=0 else 0
ax.annotate(str(val), xy=(y,x),
horizontalalignment='center',
verticalalignment='center', size=8,
color='white' if img[x][y]<thresh else 'black')
# -
# Architecture
if (net_type == 'lenet'):
net = BBBLeNet(outputs,inputs)
elif (net_type == 'alexnet'):
net = BBBAlexNet(outputs,inputs)
elif (net_type == '3conv3fc'):
net = BBB3Conv3FC(outputs,inputs)
else:
print('Error : Network should be either [LeNet / AlexNet / 3Conv3FC')
if use_cuda:
net.cuda()
vi = GaussianVariationalInference(torch.nn.CrossEntropyLoss())
optimizer = optim.Adam(net.parameters(), lr=lr, weight_decay=weight_decay)
ckpt_name = f'model_{net_type}_{dataset}_bayesian.pt'
ckpt_name
# +
# %%time
valid_loss_min = np.Inf # track change in validation loss
for epoch in range(1, n_epochs+1):
# keep track of training and validation loss
train_loss = 0.0
valid_loss = 0.0
m = math.ceil(len(train_data) / batch_size)
###################
# train the model #
###################
net.train()
for batch_idx, (data, target) in enumerate(train_loader):
# move tensors to GPU if CUDA is available
data = data.view(-1, inputs, resize, resize).repeat(num_samples, 1, 1, 1)
target = target.repeat(num_samples)
if use_cuda:
data, target = data.cuda(), target.cuda()
if beta_type is "Blundell":
beta = 2 ** (m - (batch_idx + 1)) / (2 ** m - 1)
elif beta_type is "Soenderby":
beta = min(epoch / (num_epochs // 4), 1)
elif beta_type is "Standard":
beta = 1 / m
else:
beta = 0
# clear the gradients of all optimized variables
optimizer.zero_grad()
# forward pass: compute predicted outputs by passing inputs to the model
output,kl = net.probforward(data)
# calculate the batch loss
loss = vi(output, target, kl, beta)
# backward pass: compute gradient of the loss with respect to model parameters
loss.backward()
# perform a single optimization step (parameter update)
optimizer.step()
# update training loss
train_loss += (loss.item()*data.size(0)) / num_samples
######################
# validate the model #
######################
net.eval()
for batch_idx, (data, target) in enumerate(valid_loader):
data = data.view(-1, inputs, resize, resize).repeat(num_samples, 1, 1, 1)
target = target.repeat(num_samples)
# move tensors to GPU if CUDA is available
if use_cuda:
data, target = data.cuda(), target.cuda()
# forward pass: compute predicted outputs by passing inputs to the model
output,kl = net.probforward(data)
# calculate the batch loss
loss = vi(output, target, kl, beta)
# update average validation loss
valid_loss += (loss.item()*data.size(0)) / num_samples
# calculate average losses
train_loss = train_loss/(len(train_loader.dataset) * (1-valid_size))
valid_loss = valid_loss/(len(valid_loader.dataset) * valid_size)
# print training/validation statistics
print('Epoch: {} \tValidation Loss: {:.6f}'.format(
epoch, train_loss, valid_loss))
# save model if validation loss has decreased
if valid_loss <= valid_loss_min:
print('Validation loss decreased ({:.6f} --> {:.6f}). Saving model ...'.format(
valid_loss_min,
valid_loss))
torch.save(net.state_dict(), ckpt_name)
valid_loss_min = valid_loss
# -
net.load_state_dict(torch.load(ckpt_name))
# +
# %%time
# track test loss
test_loss = 0.0
class_correct = list(0. for i in range(10))
class_total = list(0. for i in range(10))
net.eval()
m = math.ceil(len(test_data) / batch_size)
# iterate over test data
for batch_idx, (data, target) in enumerate(test_loader):
data = data.view(-1, inputs, resize, resize).repeat(num_samples, 1, 1, 1)
target = target.repeat(num_samples)
# move tensors to GPU if CUDA is available
if use_cuda:
data, target = data.cuda(), target.cuda()
if beta_type is "Blundell":
beta = 2 ** (m - (batch_idx + 1)) / (2 ** m - 1)
elif cf.beta_type is "Soenderby":
beta = min(epoch / (cf.num_epochs // 4), 1)
elif cf.beta_type is "Standard":
beta = 1 / m
else:
beta = 0
# forward pass: compute predicted outputs by passing inputs to the model
output, kl = net.probforward(data)
# calculate the batch loss
loss = vi(output, target, kl, beta)
# update test loss
test_loss += loss.item()*data.size(0) / num_samples
#test_loss += loss.item()
# convert output probabilities to predicted class
_, pred = torch.max(output, 1)
# compare predictions to true label
correct_tensor = pred.eq(target.data.view_as(pred))
correct = np.squeeze(correct_tensor.numpy()) if not use_cuda else np.squeeze(correct_tensor.cpu().numpy())
# calculate test accuracy for each object class
for i in range(batch_size):
if i >= target.data.shape[0]: # batch_size could be greater than left number of images
break
label = target.data[i]
class_correct[label] += correct[i].item()
class_total[label] += 1
# average test loss
test_loss = test_loss/len(test_loader.dataset)
print('Test Loss: {:.6f}\n'.format(test_loss))
for i in range(10):
if class_total[i] > 0:
print('Test Accuracy of %5s: %2d%% (%2d/%2d)' % (
classes[i], 100 * class_correct[i] / class_total[i],
np.sum(class_correct[i]), np.sum(class_total[i])))
else:
print('Test Accuracy of %5s: N/A (no training examples)' % (classes[i]))
print('\nTest Accuracy (Overall): %2d%% (%2d/%2d)' % (
100. * np.sum(class_correct) / np.sum(class_total),
np.sum(class_correct), np.sum(class_total)))
# +
# obtain one batch of test images
dataiter = iter(test_loader)
images, labels = dataiter.next()
images.numpy()
# move model inputs to cuda, if GPU available
if use_cuda:
images = images.cuda()
# get sample outputs
output, kl = net.probforward(images)
# convert output probabilities to predicted class
_, preds_tensor = torch.max(output, 1)
preds = np.squeeze(preds_tensor.numpy()) if not use_cuda else np.squeeze(preds_tensor.cpu().numpy())
# plot the images in the batch, along with predicted and true labels
fig = plt.figure(figsize=(25, 4))
for idx in np.arange(20):
ax = fig.add_subplot(2, 20/2, idx+1, xticks=[], yticks=[])
imshow(images[idx])
ax.set_title("{} ({})".format(classes[preds[idx]], classes[labels[idx]]),
color=("green" if preds[idx]==labels[idx].item() else "red"))
# -
| Image Recognition/Bayesian_CNN_Detailed_.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# !git clone https://github.com/omriallouche/text_classification_from_zero_to_hero.git --depth 1
import os
os.chdir('text_classification_from_zero_to_hero/notebooks')
# +
import sys, os
from pathlib import Path
def locate(fname):
"""Search file in google drive"""
if os.path.exists(fname):
return fname
try:
return next(filter(lambda p: str(p).endswith(fname),
Path("/content/drive/My Drive/nlpday_content").glob('**/*.*')))
except StopIteration:
raise FileNotFoundError(fname)
if 'google.colab' in sys.modules:
from google.colab import drive
drive.mount('/content/drive/')
__dir__ = "/content/drive/My Drive/nlpday_content/zero2hero/"
sys.path.append(__dir__ + 'src')
# -
# # BERT
# In this section we will use BERT to classify our documents.
# We will use the package pytorch-transformers by huggingface, that provides a pytorch implementation of the model.
#
# BERT can be used in different ways:
# 1. Use the model weights pre-trained on a large corpus to predict the labels for our custom task
# 1. Fine-tune the model weights, by unfreezing a certain number of the top layers, thereby better fiting the model to our custom task
# We will use the [FastBert]() package, that simplifies the use of BERT and similar transformer models, and provides an API in spirit of [fast.ai](https://github.com/fastai/fastai), aiming to expose the most important settings and take care of the rest for you.
# !pip install fast_bert
# ### 1. Create a DataBunch object
# The databunch object takes training, validation and test csv files and converts the data into internal representation for BERT, RoBERTa, DistilBERT or XLNet. The object also instantiates the correct data-loaders based on device profile and batch_size and max_sequence_length.
# For this workshop, we will use the DistilBERT model, from the paper [DistilBERT, a distilled version of BERT: smaller, faster, cheaper and lighter](https://arxiv.org/abs/1910.01108). The model is a 6-layer, 768-hidden, 12-heads, 66M parameters model (compared to BERT Base which is a 12-layer, 768-hidden, 12-heads, 110M parameters model, and BERT large which is a 24-layer, 1024-hidden, 16-heads, 340M parameters model) and trains faster, with lighter memory requirements.
# Check [this link](https://huggingface.co/transformers/pretrained_models.html) for more info on the available pretrained models.
# +
model_type = 'distilbert'
tokenizer = 'distilbert-base-uncased'
multi_label = False
DATA_PATH = '../data/'
LABEL_PATH = '../data/'
# +
from fast_bert.data_cls import BertDataBunch
databunch = BertDataBunch(DATA_PATH, LABEL_PATH,
tokenizer=tokenizer,
train_file='train.csv',
val_file='val.csv',
label_file='labels.csv',
text_col='text',
label_col='label',
batch_size_per_gpu=16,
max_seq_length=512,
multi_gpu=False,
multi_label=multi_label,
model_type=model_type)
# -
# The CSV files should contain the columns `index`, `text` and `label`. In case the column names are different than the usual text and labels, you will have to provide those names in the databunch text_col and label_col parameters.
# labels.csv will contain a list of all unique labels, or all possible tags in case of multi-label classification.
# ### 2. Create a Learner Object
# BertLearner is the ‘learner’ object that holds everything together. It encapsulates the key logic for the lifecycle of the model such as training, validation and inference.
# +
import torch
from fast_bert.learner_cls import BertLearner
from fast_bert.metrics import accuracy
import logging
OUTPUT_DIR = DATA_PATH
logger = logging.getLogger()
logger.setLevel(logging.DEBUG)
learner = BertLearner.from_pretrained_model(
databunch,
pretrained_path='distilbert-base-uncased',
metrics=[{'name': 'accuracy', 'function': accuracy}],
device=torch.device("cuda"), # for GPU,
logger=logger,
output_dir=OUTPUT_DIR,
finetuned_wgts_path=None,
warmup_steps=500,
multi_gpu=True,
is_fp16=False,
multi_label=multi_label,
logging_steps=100)
# -
# | parameter | description |
# | ------------------- | ------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------ |
# | databunch | Databunch object created earlier |
# | pretrained_path | Directory for the location of the pretrained model files or the name of one of the pretrained models i.e. bert-base-uncased, xlnet-large-cased, etc |
# | metrics | List of metrics functions that you want the model to calculate on the validation set, e.g. accuracy, beta, etc |
# | device | torch.device of type _cuda_ or _cpu_ |
# | logger | logger object |
# | output_dir | Directory for model to save trained artefacts, tokenizer vocabulary and tensorboard files |
# | finetuned_wgts_path | provide the location for fine-tuned language model (experimental feature) |
# | warmup_steps | number of training warms steps for the scheduler |
# | multi_gpu | multiple GPUs available e.g. if running on AWS p3.8xlarge instance |
# | is_fp16 | FP16 training |
# | multi_label | multilabel classification |
# | logging_steps | number of steps between each tensorboard metrics calculation. Set it to 0 to disable tensor flow logging. Keeping this value too low will lower the training speed as model will be evaluated each time the metrics are logged |
#
# ### 3. Train the model
#
# #### Side-track: View Tensorboard in a Jupyter Notebook
# Tensorboard provides real-time monitoring of the network training process. To run it in the notebook:
# Load the TensorBoard notebook extension
# %load_ext tensorboard
# %tensorboard --logdir /content/text_classification_from_zero_to_hero/data/
# #### Back to business - training a BERT model
learner.fit(epochs=100,
lr=6e-5,
validate=True, # Evaluate the model after each epoch
schedule_type="warmup_cosine",
optimizer_type="lamb")
# ### 4. Save trained model artifacts
# Model artefacts will be persisted in the output_dir/'model_out' path provided to the learner object. Following files will be persisted:
#
# | File name | description |
# | ----------------------- | ------------------------------------------------ |
# | pytorch_model.bin | trained model weights |
# | spiece.model | sentence tokenizer vocabulary (for xlnet models) |
# | vocab.txt | workpiece tokenizer vocabulary (for bert models) |
# | special_tokens_map.json | special tokens mappings |
# | config.json | model config |
# | added_tokens.json | list of new tokens |
#
# As the model artefacts are all stored in the same folder, you will be able to instantiate the learner object to run inference by pointing pretrained_path to this location.
learner.save_model()
# ### 5. Model Inference
#
# In order to perform inference, you need to save the model (see step #4 above).
# To make predictions, we init a `BertClassificationPredictor` object with the path of the model files:
# +
from fast_bert.prediction import BertClassificationPredictor
MODEL_PATH = OUTPUT_DIR + 'model_out'
predictor = BertClassificationPredictor(
model_path=MODEL_PATH,
label_path=LABEL_PATH, # location for labels.csv file
multi_label=False,
model_type=model_type,
do_lower_case=True)
# -
# Single prediction
sentence_to_predict = "the problem with the middleeast is those damn data scientists. All they do is just look at the computer screen instead of laying outside in the warm sun."
single_prediction = predictor.predict(sentence_to_predict)
single_prediction
# +
# Batch predictions
texts = [
"this is the first text",
"this is the second text"
]
multiple_predictions = predictor.predict_batch(texts)
multiple_predictions
# -
# Next, let's run the model on our validation dataset and report its performance.
# +
# Batch predictions
import pandas as pd
df = pd.read_csv('../data/val.csv')
texts = list(df['text'].values)
multiple_predictions = predictor.predict_batch(texts)
# Each prediction includes the softmax value of all possible labels, sorting in descending order.
# We thus use only the first element, which is the most-probable label, and keep only the first element in that tuple, which is the name of the label
y_predicted = [x[0][0] for x in multiple_predictions]
# +
from sklearn import metrics
y_truth = df['label']
metrics.f1_score(y_truth, y_predict, average='macro')
# -
# **Try it yourself:** Can you improve the performance of our classifier? Let's see how high you can get.
# Good luck!
# ### Language Model Fine-Tuning
# It is also possible to fine-tune the Language Model of BERT, to fit to the custom domain. The process requires a long training time, even on powerful GPUs. Check [this link](https://github.com/kaushaltrivedi/fast-bert#language-model-fine-tuning) for instructions how to do it.
| notebooks/5_bert.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1><center>Smart Infrastructure Maintainance</center></h1>
#
# <div style="text-align: center">
#
# Author - <NAME>
# University - Indian Institute of Technology Madras
# Current Year - 4th
#
# Team Name - [Bracket]
# Theme - Smart Infrastructure Maintainance
#
# </div>
# # Task 1 - Data Compression
#
# ## 1. Setup
#
# Please follow the setup instructions mentioned in each section for successfully running this notebook.
#
# ### 1.1 Dependencies
#
# Following packages are required to run this notebook. Please ensure that these are installed in your system. Uncomment the first cell to download using pip.
#
# - os (system package)
# - pickle (system package)
# - datetime (system package)
# - warnings(system package)
# - pandas
# - numpy
# - matplotlib
# - sklearn
# - tensorflow
# +
# Uncomment the below cells for installing packages
# # !pip install pandas
# # !pip install numpy
# # !pip install matplotlib
# # !pip install scikit-learn
# # !pip install tensorflow
# +
import os
import pickle
import sklearn
import datetime
import warnings
import numpy as np
import pandas as pd
import tensorflow as tf
import matplotlib.pyplot as plt
from scripts.sap_hackathon import *
warnings.simplefilter(action='ignore')
# -
# A helper function for calculating error
def error(y_true, y_pred):
error = 0
for i in range(20):
error += np.sum(np.abs(np.sum(y_true[:, 3*i:3*(i+1)]**2, axis=1) - np.sum(y_pred[:, 3*i:3*(i+1)]**2, axis=1)))
return 100 * error / np.sum(y_true**2)
def rmse(y_true, y_pred):
error = np.sum((np.sum(y_true**2, axis=1) - np.sum(y_pred ** 2, axis=1))**2)
return error / y_true.shape[0]
# ### 1.2 Data Directory
#
# Please ensure that the scripts & dataset (both train and test) follows the following directory structure:-
#
# |-- base_dir
#
# [Code]
# |-- task_1_data_compression_notebook.ipynb (This notebook)
# |-- scripts
# |-- sap_hackathon.py
#
# [Model]
# |-- model
# |-- encoder.pkl
# |-- decoder.pkl
#
# [Train/Test Data]
# |-- LORD-6305-6000-69503
# |-- year=2020
# |-- month=10
# |-- day=19
# |-- hour=04
# |-- ... [All the hours]
# |-- hour=23
# |--... [All the Days]
# |-- day=25
#
# |-- LORD-6305-6000-69504
# |-- year=2020
# |-- month=10
# |-- day=19
# |-- hour=04
# |-- ... [All the hours]
# |-- hour=23
# |--... [All the Days]
# |-- day=25
#
# |-- ... [All the Sensors]
#
# |-- LORD-6305-6182-92392
# |-- year=2020
# |-- month=10
# |-- day=19
# |-- hour=04
# |-- ... [All the hours]
# |-- hour=23
# |--... [All the Days]
# |-- day=25
#
#
# 1. The data should reside in the structure shown above.
# 2. The name of the sensors, like LORD-6305-6182-92392 remains the same in test data and train data.
# 3. Please mention the time period of test data in the list in following cell (User Input).
# ### User Input
# +
# Year of the test data
test_year = ['2020']
# Month of the test data
test_month = ['10']
# Date of the test data
test_date = ['19']
# Time of the test data
test_hour = ['05']
# -
# ## 2. Extract, Load, Transform (ELT)
#
# In this section, the data will be loaded <b>according to the time which are listed in the previous cell </b>. Note that 20 dataframes will be loaded, after which they will be concatenated to form a single dataframe containing all the 60 sensor readings given by their time-stamp.
#
# ### 2.1 Transformation
#
# An accelerometer principally measures proper acceleration of the body on which it is installed. Thus the effect of acceleration due gravity would be included in the readings of the accelerometer. Since the orientation of the acceleormeter is not known beforehand, therefore as a first step, we <b>transform the axis of acceleormeters to align with the direction of gravity</b>. This is pictorically represented in the figure below:-
#
# <img src='./images/transformation.PNG' align="center" height="230px">
#
# The coordinate system (x, y, z) is transformed into (u, v, w) such that the 'w'-axis represents the direction of gravity. In the new transformed coordinate system, the mean acceleration on (u, v)- axis will be zero. With this transformation, it can be ensured that the 'w'-axis of all the 20 sensors point in the same direction. However, the the relative orientation of (u, v)-axis between different sensors could not be established.
#
# After the transformation, the <b>50-th percentile of each sensor from each transformed axis</b>. This ensures that a zero mean signal is obtained which is later used as primary features in our model of data compression.
sensor_idx = 1
test_df_list = []
for s, sensor_name in enumerate(sensors):
if s == sensor_idx:
temp_df = read_test_data(sensor_name, test_year, test_month, test_date, test_hour, s, True)
else:
temp_df = read_test_data(sensor_name, test_year, test_month, test_date, test_hour, s, False)
test_df_list.append(temp_df)
plot_transformed_axis(test_df_list[sensor_idx], sensor_idx)
# Concatenating into single dataframe
# NOTE - Time stamps will be dropped for which there is atleast one missing data
common_time = estimate_common_time(test_df_list)
test_df = pd.DataFrame(common_time, columns=['_t']).sort_values(by='_t')
for i in range(len(test_df_list)):
test_df = pd.merge(test_df, test_df_list[i], on='_t', how='left')
print(test_df.shape)
# ## 3. Encoder Model (Data Compression)
#
# The encoder model is saved as './model/encoder.pkl'. This model is essentially <b>sklearn's PCA class object</b> which is saved as pkl. Final results is also summarized at the end of this cell.
#
# For detailed explaination of the model and training steps, please refer to train notebook.
#
# The encoded data is saved as CSV file as './data/encoded_data.csv'.
# +
# %%time
X = test_df.drop(['_t'], axis=1).to_numpy()
# Loading encoder and transforming to encodings
encoder = pickle.load(open('model/encoder.pkl', mode='rb'))
X_trans = encoder.transform(X)
# +
# Compression Results
print('Original components length = ', X.shape[1])
print('Encoded components length =', X_trans.shape[1])
print('Compression Ratio =', X.shape[1] / X_trans.shape[1])
# Saving the encoded data under './data/encoded_data.csv'
encoded_data = pd.DataFrame(X_trans, columns=['ch' + str(i+1) for i in range(X_trans.shape[1])])
encoded_data = pd.concat([test_df[['_t']], encoded_data], axis=1)
# -
# ## 4. Decoder Model (Data Reconstruction)
#
# The decoder model is saved as './model/decoder.pkl'. This model is essentially <b>sklearn's PCA class object</b> which is saved as pkl. Final results is also summarized at the end of this cell.
#
# For detailed explaination of the model and training steps, please refer to train notebook.
#
# The decoded data is saved as CSV file as './data/decoded_data.csv'.
# %%time
decoder = pickle.load(open('model/decoder.pkl', mode='rb'))
X_decoded = decoder.inverse_transform(X_trans)
# +
# Saving the decoded data under './data/decoded_data.csv'
decoded_data = pd.DataFrame(X_decoded, columns=test_df.drop(['_t'], axis=1).columns)
decoded_data = pd.concat([test_df[['_t']], decoded_data], axis=1)
df_list=[]
for s in range(1, 21):
features = ['_t', 'x_' + str(s), 'y_' + str(s), 'z_' + str(s)]
df_list.append(project_back(decoded_data[features], s))
# +
sensor_indices = [0, 1]
axis_title = ['ch1', 'ch2', 'ch3']
# Visualizing reconstruction for above mentioned sensors
for si in sensor_indices:
original_data = read_file(sensors[si], test_year, test_month, test_date, test_hour)
original_data = original_data[original_data['_t'].isin(test_df['_t'])]
fig, ax = plt.subplots(nrows=1, ncols=3, figsize=(15, 5))
fig.suptitle('After Reconstruction - Comparison of Original & Reconstructed Data for Sensor ' + sensors[si])
x_range = np.arange(original_data.shape[0])
for i in range(3):
ax[i].plot(x_range, original_data[axis_title[i]], color='blue', label='original data')
ax[i].plot(x_range, df_list[si][axis_title[i]], color='orange', label='after reconstruction')
ax[i].set_xlabel('Time in seconds')
ax[i].set_ylabel('Acceleration in m/s^2')
ax[i].set_title(axis_title[i])
ax[i].legend()
ax[0].set_xlabel('Time in seconds')
ax[0].set_ylabel('Acceleration in m/s^2')
ax[0].legend()
plt.show()
# -
# ## 5. Metric Computation
#
# Meaduring the percentage error due to data compression and reconstruction using the formula given in the problem statement.
print('Error in data reconstruction =', error(X, X_decoded), '%')
# # Task 2 - Sensor Network Robustness
# ## 1. Setup
#
# The setup instruction are same as that for the Task 1. Please follow the instruction given in the Task 1 properly to run this Task.
#
# 1. The data should reside in the directory structure shown above.
# 2. The name of the sensors, like LORD-6305-6182-92392 remains the same in test data and train data.
# 3. Please mention the time period of test data in the list in following cell (User Input).
# ### User Input:
# +
# Year of the test data
test_year = ['2020']
# Month of the test data
test_month = ['10']
# Date of the test data
test_date = ['19']
# Time of the test data
test_hour = ['05']
# The sensor on which anomaly has to be checked
test_sensor = 'LORD-6305-6182-92382'
# -
# ## 2. Extract, Load, Transform (ELT)
#
# The preprocessing steps performed in this task are essentially the same as that in Task 1. Please refer to Section 2 of task 1 for a detailed overview.
test_df_list = []
skip_sensor = sensors.index(test_sensor)
for s, sensor_name in enumerate(sensors):
if s != skip_sensor:
temp_df = read_test_data(sensor_name, test_year, test_month, test_date, test_hour, s)
test_df_list.append(temp_df)
# +
# Concatenating into single dataframe
# NOTE - Time stamps will be dropped for which there is atleast one missing data
common_time = estimate_common_time(test_df_list)
test_df = pd.DataFrame(common_time, columns=['_t']).sort_values(by='_t')
for i in range(len(test_df_list)):
test_df = pd.merge(test_df, test_df_list[i], on='_t', how='left')
print(test_df.shape)
# -
# ## 3. Model - Multilayer Perceptron (MLP)
#
# The final predictions are saved under 'data/predictions.csv'. These can be used by the user to compare the data for test case.
# %%time
X = test_df.drop(['_t'], axis=1).to_numpy()
model = tf.keras.models.load_model('model/model_task_' + str(skip_sensor+1), compile=False)
X_predicted = model.predict(X)
# +
pred_df = test_df[['_t']]
pred_df = pd.concat([pred_df, pd.DataFrame(
X_predicted, columns=['x_' + str(skip_sensor+1), 'y_' + str(skip_sensor+1), 'z_' + str(skip_sensor+1)])], axis=1)
decoded_df = project_back(pred_df, skip_sensor + 1)
# Loading ctual ground truth
ground_truth = project_back(read_test_data(test_sensor, test_year, test_month, test_date,
test_hour, skip_sensor), skip_sensor+1)
ground_truth = ground_truth[ground_truth['_t'].isin(common_time)].reset_index(drop=True)
# Calculating mean squared Error
y_true = ground_truth.drop(['_t'], axis=1).to_numpy()
y_pred = decoded_df.drop(['_t'], axis=1).to_numpy()
# -
fig, ax = plt.subplots(nrows=1, ncols=3, figsize=(15, 5))
fig.suptitle('Comparing predictions for Sensor ' + test_sensor)
for i in range(3):
ax[i].plot(y_true[:, i], color='blue', label='original data')
ax[i].plot(y_pred[:, i], color='orange', label='predicted_data')
ax[i].set_xlabel('Time in seconds')
ax[i].set_ylabel('Acceleration in m/s^2')
ax[i].legend(loc=1)
ax[0].set_xlabel('Time in seconds')
ax[0].set_ylabel('Acceleration in m/s^2')
ax[0].legend(loc=1)
plt.show()
# ## 4. Result Analysis
print('Mean Squared Error =', rmse(y_true, y_pred))
| Notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
class State:
def __init__(self,stack,table,action):
self.stack = stack
self.table = table
self.action = action
def compare(self,state):
if (len(self.stack) != len(state.stack) | len(self.table) != len(state.table) ):
return False
for i in range(len(state.stack)):
if self.stack[i] != state.stack[i]:
return False
return len(set(self.table) & set(state.table)) == len(state.table)
def pop(self):
piece = self.stack.pop()
self.table.append(piece)
return piece
def push(self,piece):
self.table.remove(piece)
self.stack.append(piece)
def copy(self,action):
return State(self.stack.copy(),self.table.copy(),action)
def expand(self):
expand_list = []
for piece in self.table:
state = self.copy("push {0} in stack.".format(piece))
# print('--- before push -----')
state.print_state()
state.push(piece)
# print('--- after push -----')
state.print_state()
expand_list.append(state)
if (len(self.stack) != 0):
# print('--- before pop -----')
state = self.copy(None)
state.print_state()
state.action = "pop {0} from stack.".format(state.pop())
# print('--- after pop -----')
state.print_state()
expand_list.append(state)
return expand_list
def print_state(self):
# print('Stack' + str(self.stack))
# print('Table' + str(self.table))
i = 0
class Node:
def __init__(self,state,father_node):
self.state = state
self.father_node = father_node
def printNode(self):
self.state.print_state()
def print_family(self):
l = []
n = self
while n.father_node != None:
l.append(n)
n = n.father_node
l.reverse()
for n in l:
print(n.state.stack)
print(n.state.table)
print(n.state.action)
class general_search:
def __init__(self,state,goal_state):
self.state = state
self.goal_state = goal_state
def search(self):
node = Node(self.state,None)
open_list = [node]
while True:
if len(open_list) == 0:
return None
node = open_list.pop(0)
# print('Abierta arriba')
node.printNode()
if node.state.compare(self.goal_state):
print(len(open_list))
return node
# print('Antes expand')
succesors = node.state.expand()
for state in succesors:
n = Node(state,node)
# print('--- expandiendo ----')
n.printNode()
open_list.append(n)
# print('---- fin succesores ----')
initial_state = State(['E','D','A'],['C','B'],'Inicio')
goal_state = State(['E','D','C','B','A'],[],'Estado Final')
s = general_search(initial_state,goal_state)
result = s.search()
result.print_family()
# -
| Razonamiento_y_Planification/Lab1/Laboratorio_1_Gonzalo_Molina.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import scipy as sp
import seaborn as sns
from pandas.tools.plotting import scatter_matrix
sns.set()
sns.set_color_codes()
daily_activities = pd.read_csv("../Takeout/Fit/Daily Aggregations/Daily Summaries.csv",
index_col = "Date", parse_dates=True)
print(daily_activities.shape)
daily_activities.head()
# * droppping rows with no caloric information
daily_activities = daily_activities[np.isfinite(daily_activities['Calories (kcal)'])]
print(daily_activities.shape)
daily_activities.columns
daily_activities['Calories (kcal)'].describe()
daily_activities.describe()
daily_activities[daily_activities['Calories (kcal)']==daily_activities['Calories (kcal)'].max()]
daily_activities["Distance (miles)"] = daily_activities['Distance (m)']/1609.34
# +
fig, ax1 = plt.subplots() #Initialize plots
ax2 = ax1.twinx()
ax1.plot(daily_activities.index, daily_activities['Calories (kcal)'], '-r') # Percipitation plot
ax2.plot(daily_activities.index, daily_activities['Distance (miles)']) # Tmax
ax1.set_xlabel('Date') #Label axis
ax1.set_ylabel('Calories measured in kcal')
ax2.set_ylabel('Distance in miles')
fig.autofmt_xdate() # Makes sure the dates on x axis don't overlap
plt.show()
# -
marathon = pd.read_csv("../Takeout/Fit/Daily Aggregations/2015-11-29.csv")
print(marathon.shape)
marathon.head()
| notebooks/activity_summary_explorer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ---
# title: "Order statistics — Part 3: Proof of expectation formula for uniform RVs"
# date: 2021-03-08
# categories: [statistics, order statistics]
# tags: [statistics, order statistics, uniform distribution, expectation formula, gamma function, beta function]
# preview_image: /enrightward.github.io/assets/img/order-statistics/part3/gamma.png
# ---
# 
# ## 1. Introduction
#
# In the [last post](https://enrightward.github.io/enrightward.github.io/posts/order-statistics-part-2/), we proved the following general formulae, after conducting some numerical experiments to gain intuition:
#
# \begin{align}
# F_{X_{(k)}}(x) &= \sum_{j=k}^{n} \binom{n}{j} F_{X}(x)^{j} (1 - F_{X}(x))^{n-j}, \newline
# f_{X_{(k)}}(x) &= k \binom{n}{k} f_{X}(x) \cdot F_{X}(x)^{k-1} \, (1 - F_{X}(x))^{n-k}, \newline
# \mathbb{E} \left \lbrack X_{(k)} \right \rbrack &=
# k \binom{n}{k} \int_{0}^{1} x \cdot f_{X}(x) \cdot F_{X}(x)^{k-1} \, (1 - F_{X}(x))^{n-k} \, dx,
# \end{align}
#
# Note they depend only on $n, k, f_{X}$ and $F_{X}$. We will now round out the exposition of $k^{\textrm{th}}$ order statistics by specialising these general formulae to the case of the uniform distribution $U(0, 1)$. We will show that:
#
# \begin{align}
# F_{X_{(k)}}(x) &= \sum_{j=k}^{n} \binom{n}{j} x^{j} (1 - x)^{n-j}, \newline
# f_{X_{(k)}}(x) &= k \binom{n}{k} x^{k-1} \, (1 - x)^{n-k}, \newline
# \mathbb{E} \left \lbrack X_{(k)} \right \rbrack &= \frac{k}{n+1} \cdot
# \end{align}
#
# This post is purely theoretical, and has no code. The mathematical tools we use are the beta and gamma functions, introduced below.
# ## 2. Specialising the formulae to $U(0, 1)$
#
# In the case where the $X \sim U(0, 1)$, we have $F_{X}(x) = x$ and $f_{X}(x) = 1$ on the unit interval, so the formulae computed above specialise to:
#
# \begin{align}
# F_{X_{(k)}}(x) &= \sum_{j=k}^{n} \binom{n}{j} x^{j}(1 - x)^{n - j}, \newline
# f_{X_{(k)}}(x) &= k \binom{n}{k} x^{k-1} (1 - x)^{n-k}, \newline
# \mathbb{E}[X_{(k)}] &=
# k \binom{n}{k} \int_{0}^{1} x \cdot x^{k-1} \, (1 - x)^{n-k} \, dx =
# k \binom{n}{k} \int_{0}^{1} x^{k} (1 - x)^{n-k} \, dx.
# \end{align}
# ## 3. Proving the expectation formulae using beta and gamma functions
#
# The integral in the above expectation formula is an example of the [_beta function_](https://en.wikipedia.org/wiki/Beta_function), defined as
#
# \begin{align}
# B(z, w) := \int_{0}^{1} x^{z-1} (1 - x)^{w-1} \, dx.
# \end{align}
#
# This formula makes sense for complex numbers $w$ and $z$ with positive real part, but we are only interested in integer-valued arguments, because our exponents for $x$ and $1-x$ are $k$ and $n-k$, respectively. In the new notation of beta functions, our expectations are written:
#
# \begin{align}
# \mathbb{E}[X_{(k)}] = k \binom{n}{k} \int_{0}^{1} x^{k} (1 - x)^{n-k} \, dx = k \binom{n}{k} B(k+1, n-k+1).
# \end{align}
#
# This is interesting to us because the beta function can be computed in terms of the [_gamma function_](https://en.wikipedia.org/wiki/Gamma_function)
#
# \begin{align}
# \Gamma(z) := \int_{0}^{\infty} x^{z-1} e^{-x} \, dx,
# \end{align}
#
# and the gamma function is a generalisation to the (positive half) complex plane of the factorial function $n \mapsto n!$, hence reduces to factorial computations for integer-valued arguments. Concretely, the relations we need are:
#
# \begin{align}
# B(z, w) &= \frac{\Gamma(z) \Gamma(w)}{\Gamma(z + w)}, \quad \textrm{and} \newline
# \Gamma(n) &= (n-1)!
# \end{align}
#
# We will show these relations shortly. For now, observe they imply the formulae we conjectured for the expectations of $X_{(k)}$. Indeed, using each of these results in turn, we have
#
# \begin{align}
# B(k+1, n-k+1) = \frac{\Gamma(k+1) \Gamma(n-k+1)}{\Gamma(n+2)} =
# \frac{k!(n-k)!}{(n+1)!} = \frac{1}{(n+1) \binom{n}{k}},
# \end{align}
#
# and hence
#
# \begin{align}
# \mathbb{E}[X_{(k)}] &= k \binom{n}{k} B(k+1, n-k+1)
# = \frac{k \binom{n}{k}}{(n+1) \binom{n}{k}} = \frac{k}{n+1} \cdot
# \end{align}
# ## 4. Proving the necessary properties of beta and gamma functions
#
# Now we prove the relations for $B(z, w)$ and $\Gamma(z)$. To see that $\Gamma(n) = (n-1)!$, it suffices to prove the recursive formula
#
# \begin{align}
# \Gamma(z) = (z-1)\Gamma(z-1),
# \end{align}
#
# for any $z$ with positive real part. For then we would have:
#
# \begin{align}
# \Gamma(n) &= (n-1)\Gamma(n-1) \newline
# &= (n-1)(n-2)\Gamma(n-2) \newline
# \vdots \newline
# &= (n-1)(n-2)(n-3) \ldots (2)(1) = (n-1)!
# \end{align}
#
# We show this using the definite integral version of the integration by parts formula:
#
# \begin{align}
# \int_{a}^{b} u \, dv = \left \lbrack uv \right \rbrack_{a}^{b} - \int_{a}^{b} v \, du,
# \end{align}
#
# with $u = x^{z-1}$ and $dv = e^{-x} dx$, implying that $du = (z-1)x^{z-2} dx$ and $v = -e^{-x}$. Note this formula holds also in the case where $a$ or $b$ is $\pm \infty$, provided we interpret the term $\left \lbrack uv \right \rbrack_{a}^{b}$ as expressing the appropriate limit(s). We have:
#
# \begin{align}
# \Gamma(z) &= \int_{0}^{\infty} x^{z-1} e^{-x} \, dx \newline
# &= \lim_{b \rightarrow \infty} \left \lbrack -x^{z-1} e^{-x} \right \rbrack_{0}^{b} + (z-1) \int_{0}^{\infty} x^{z-2} e^{-x} \, dx \newline
# &= 0 + (z-1) \int_{0}^{\infty} x^{z-2} e^{-x} \, dx \newline
# &= (z-1)\Gamma(z-1).
# \end{align}
#
# Now we prove the formula connecting $B(z, w)$ and $\Gamma(z)$. I'm taking this proof [from Wikipedia](https://en.wikipedia.org/wiki/Beta_function#Relationship_to_the_gamma_function) and filling in details on the variable substitution using [these lecture notes](https://homepage.tudelft.nl/11r49/documents/wi4006/gammabeta.pdf) (Theorem 2, page 3). Note: Curiously, although the Wikipedia article cites <NAME>'s book [Gamma Functions](https://web.archive.org/web/20161112081854/http://www.plouffe.fr/simon/math/Artin%20E.%20The%20Gamma%20Function%20(1931)(23s).pdf), pages 18-19, Artin's proof is not the one written down there.
#
# In any case, the argument begins like this:
#
# \begin{align}
# \Gamma(z) \, \Gamma(w) &= \int_{x=0}^{\infty} x^{z-1} e^{-x} \, dx \int_{y=0}^{\infty} y^{w-1} e^{-y} \, dy \newline
# &= \int_{x=0}^{\infty} \int_{y=0}^{\infty} e^{-(x+y)} x^{z-1} y^{w-1} \, dx \, dy.
# \end{align}
#
# Now we introduce an implicit change of variables $s, t$, defined by the conditions $x = st$ and $y=s(1 - t)$. This transformation has Jacobian matrix:
#
# \begin{align}
# J(x(s, t), y(s, t)) = \left(
# \begin{array}{cc}
# \frac{\partial x}{\partial s} & \frac{\partial x}{\partial t} \newline
# \frac{\partial y}{\partial s} & \frac{\partial y}{\partial t}
# \end{array} \right) = \left(
# \begin{array}{cc}
# t & s \newline
# 1-t & -s
# \end{array} \right),
# \end{align}
#
# with determinant $\det(J) = -st - s(1-t) = -s$. It follows that
#
# \begin{align}
# dx \, dy = \left\| \det{J} \right\| ds \, dt = s \, ds \, dt.
# \end{align}
#
# Observe that since $x$ and $y$ range over $[0, \infty)$ and $x + y = st + s(1-t) = s$, then $s$ must range over
# $[0, \infty)$, too. On the other hand,
#
# \begin{align}
# t = \frac{x}{s} = \frac{x}{x + y},
# \end{align}
#
# so $t$ ranges only over the unit interval $[0, 1]$. Making these substitutions gives:
#
# \begin{align}
# \Gamma(z) \cdot \Gamma(w) &=
# \int_{x=0}^{\infty} \int_{y=0}^{\infty} e^{-(x+y)} x^{z-1} y^{w-1} \, dx \, dy \newline
# &= \int_{s=0}^{\infty} \int_{t=0}^{1} e^{-s} (st)^{z-1} (s(1 - t))^{w-1} \, s \, dt \, ds \newline
# &= \int_{s=0}^{\infty} \int_{t=0}^{1} e^{-s} s^{z+w-1} t^{z-1}(1 - t)^{w-1} \, dt \, ds \newline
# &= \int_{0}^{\infty} s^{z+w-1} e^{-s} \left\{ \int_{0}^{1} t^{z-1}(1 - t)^{w-1} \, dt \, \right\} ds \newline
# &= B(z, w) \int_{0}^{\infty} s^{z+w-1} e^{-s} ds \newline
# &= B(z, w) \cdot \Gamma(z+w).
# \end{align}
#
# Dividing both sides of this equation by $\Gamma(z+w)$ now gives the result.
# ## 5. Roundup
#
# We specialised the general formulae:
#
# \begin{align}
# F_{X_{(k)}}(x) &= \sum_{j=k}^{n} \binom{n}{j} F_{X}(x)^{j} (1 - F_{X}(x))^{n-j}, \newline
# f_{X_{(k)}}(x) &= k \binom{n}{k} f_{X}(x) \cdot F_{X}(x)^{k-1} \, (1 - F_{X}(x))^{n-k}, \newline
# \mathbb{E} \left \lbrack X_{(k)} \right \rbrack &=
# k \binom{n}{k} \int_{0}^{1} x \cdot f_{X}(x) \cdot F_{X}(x)^{k-1} \, (1 - F_{X}(x))^{n-k} \, dx,
# \end{align}
#
# which depend only on $n, k, f_{X}$ and $F_{X}$, to the case of the uniform distribution $U(0, 1)$. We showed that:
#
# \begin{align}
# F_{X_{(k)}}(x) &= \sum_{j=k}^{n} \binom{n}{j} x^{j} (1 - x)^{n-j}, \newline
# f_{X_{(k)}}(x) &= k \binom{n}{k} x^{k-1} \, (1 - x)^{n-k}, \newline
# \mathbb{E} \left \lbrack X_{(k)} \right \rbrack &= \frac{k}{n+1} \cdot
# \end{align}
#
# To prove the expectation formula, we introduced the beta and gamma function, and proved some identities they satisfy.
| notebooks/order-statistics-part-3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Linear Models using only large schools could not produce a satisfactory model. Random forest as a modeling technique was tried. All the large schools were plotted against graduation rates on the y axis to visualize a pattern. No pattern was detected.
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LinearRegression
from sklearn import metrics
from sklearn.ensemble import RandomForestRegressor
from sklearn.model_selection import RandomizedSearchCV
large = pd.read_csv('/Users/flatironschool/Absenteeism_Project/data/processed/large_schools.csv')
large.head()
# ## Imputing Mean for Missing Values
#impute mean for numerical vars
large_feat = large[['total_enrollment', 'ap_ib_de_rate', 'sat_act_rate', 'pass_algebra_rate',
'geometry_rate', 'algebra2_rate', 'calc_rate', 'chronic_absent_rate', 'activities_funds_rate',
'sports_rate', 'suspensed_day_rate', 'harassed_rate',
'non_cert_rate','counselor_rate','absent_teacher_rate']]
imputed_numeric_df = large_feat
for i in range(0, len(large_feat.columns)):
print('im here:', large_feat.columns[i])
imputed_numeric_df.iloc[:,i].replace(np.NaN, imputed_numeric_df.iloc[:,i].mean(), inplace=True)
#No missing values
imputed_numeric_df.isna().sum()
imputed_numeric_df.head()
# ## Model 1 - Random Forest
y = large['ALL_RATE_1516']
#features to use for prediction came from
X = imputed_numeric_df
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2)
rf_trial = RandomForestRegressor(n_estimators=10)
rf_trial.fit(X_train, y_train)
rf_trial.score(X_test, y_test)
rf = RandomForestRegressor(n_estimators=500, min_samples_leaf=2)
rf.fit(X_train, y_train)
rf.score(X_test, y_test)
rf.score(X_train, y_train)
rf.feature_importances_
large_feat.columns
pd.DataFrame(data = [large_feat.columns, rf.feature_importances_])
# ## Model 2 - Random Forest Tuning
X_train_tune, X_test_tune, y_train_tune, y_test_tune = train_test_split(X, y, test_size=0.2)
# +
# Create the random grid
# Number of trees in random forest
n_estimators = [int(x) for x in np.linspace(start = 200, stop = 4500, num = 10)]
# Number of features to consider at every split
max_features = ['auto', 'sqrt']
# Maximum number of levels in tree
max_depth = [int(x) for x in np.linspace(10, 110, num = 11)]
max_depth.append(None)
# Minimum number of samples required to split a node
min_samples_split = [2, 5, 10]
# Minimum number of samples required at each leaf node
min_samples_leaf = [1, 2, 4]
# Method of selecting samples for training each tree
bootstrap = [True, False]
random_grid = {'n_estimators': n_estimators,
'max_features': max_features,
'max_depth': max_depth,
'min_samples_split': min_samples_split,
'min_samples_leaf': min_samples_leaf,
'bootstrap': bootstrap}
# -
rf_random = RandomizedSearchCV(estimator = r, param_distributions = random_grid, n_iter = 100, cv = 3, verbose=2, random_state=42, n_jobs = -1)
rf_random.fit(X_train_tune, y_train_tune)
rf_random.best_params_
def evaluate(model, test_features, test_labels):
predictions = base_model.predict(X_test_tune)
errors = (predictions - y_test_tune)
rmse = np.sqrt(np.mean(errors**2)) ### RMSE
print('Model Performance')
print('Average Error: {:0.4f} degrees.'.format(np.mean(errors)))
print('RMSE = {:0.4f}'.format(rmse))
return rmse
base_model = RandomForestRegressor(n_estimators = 10)
base_model.fit(X_train_tune, y_train_tune)
base_accuracy = evaluate(base_model, X_test_tune, y_test_tune)
base_model.decision_path(X_test)
y_train_tune
best_random = rf_random.best_estimator_
random_accuracy = evaluate(best_random, X_test_tune, y_test_tune)
print('Improvement of {:0.2f}%.'.format( 100 * (random_accuracy - base_accuracy) / base_accuracy))
# ## Plotting all features against graduation ALL RATE
# +
y = large['ALL_RATE_1516']
fig = plt.subplots(nrows=7,ncols=2, figsize=(20,40))
plt.subplot(7, 2, 1)
x = large['ap_ib_de_rate']
plt.scatter(x, y)
plt.title('ap_ib_de_rate')
plt.subplot(7, 2, 2)
x = large['sat_act_rate']
plt.scatter(x, y)
plt.title('sat_act_rate')
plt.subplot(7, 2, 3)
x = large['pass_algebra_rate']
plt.scatter(x, y)
plt.title('pass_algebra_rate')
plt.subplot(7, 2, 4)
x = large['geometry_rate']
plt.scatter(x, y)
plt.title('geometry_rate')
plt.subplot(7, 2, 5)
x = large['algebra2_rate']
plt.scatter(x, y)
plt.title('algebra2_rate')
plt.subplot(7, 2, 6)
x = large['calc_rate']
plt.scatter(x, y)
plt.title('calc_rate')
plt.subplot(7, 2, 7)
x = large['chronic_absent_rate']
plt.scatter(x, y)
plt.title('chronic_absent_rate')
plt.subplot(7, 2, 8)
x = large['activities_funds_rate']
plt.scatter(x, y)
plt.title('activities_funds_rate')
plt.subplot(7, 2, 9)
x = large['sports_rate']
plt.scatter(x, y)
plt.title('sports_rate')
plt.subplot(7, 2, 10)
x = large['suspensed_day_rate']
plt.scatter(x, y)
plt.title('suspensed_day_rate')
plt.subplot(7, 2, 11)
x = large['harassed_rate']
plt.scatter(x, y)
plt.title('harassed_rate')
plt.subplot(7, 2, 12)
x = large['non_cert_rate']
plt.scatter(x, y)
plt.title('non_cert_rate')
plt.subplot(7, 2, 13)
x = large['counselor_rate']
plt.scatter(x, y)
plt.title('counselor_rate')
plt.subplot(7, 2, 14)
x = large['absent_teacher_rate']
plt.scatter(x, y)
plt.title('absent_teacher_rate')
# -
# + active=""
#
# -
| notebooks/RFregressor_Large_Schools.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.12 64-bit (''scrape_london_marathon'': conda)'
# name: python3
# ---
# Setting up required libraries
import pandas as pd
import numpy as np
import requests
from bs4 import BeautifulSoup, SoupStrainer
# + tags=["outputPrepend"]
# Connect to website to be scraped, and get all html
url1 = "https://results.virginmoneylondonmarathon.com/2021/?page="
url2 = "&event=ALL&num_results=1000&pid=search&pidp=results_nav&search%5Bsex%5D="
url3 = "&search%5Bage_class%5D=%25&search%5Bnation%5D=%25&search_sort=name"
# Get results for men, 3 pages of results <-There is no search option for other gender/sex
mens_results = pd.DataFrame()
for i in range(3): # max 25
sex = "M"
site = requests.get(url1 + str(i) + url2 + sex + url3).text
# Soup strainer restricts content to speed up soup
strainer = SoupStrainer(class_="section-main")
soup = BeautifulSoup(site, "lxml", parse_only=strainer)
# fields = soup.find(class_='section-main')
# Loop through each row and column to create a list of cells
my_table = []
for row in soup.find_all(class_="list-group-item"):
row_data = []
for cell in row.find_all(class_="list-field"):
row_data.append(cell.text)
# If the row isn't empty, then create a dict of the row to create dataframe from
if row_data:
data_item = {
"Place (Overall)": row_data[0],
"Place (Gender)": row_data[1],
"Place (Category)": row_data[2],
"Name": row_data[3],
"Sex": sex,
"Club": row_data[4],
"Running Number": row_data[5],
"Category": row_data[6],
"Finish": row_data[9],
"Year": 2021,
}
my_table.append(data_item)
df = pd.DataFrame(my_table).iloc[1:] # Strip table header
mens_results = mens_results.append(df)
# -
# Get results for women
womens_results = pd.DataFrame()
for i in range(3): # 17
sex = "F"
site = requests.get(url1 + str(i) + url2 + sex + url3).text
# Soup strainer restricts content to speed up soup
strainer = SoupStrainer(class_="section-main")
soup = BeautifulSoup(site, "lxml", parse_only=strainer)
# fields = soup.find(class_='section-main')
# Loop through each row and column to create a list of cells
my_table = []
for row in soup.find_all(class_="list-group-item"):
row_data = []
for cell in row.find_all(class_="list-field"):
row_data.append(cell.text)
# If the row isn't empty, then create a dict of the row to create dataframe from
if row_data:
data_item = {
"Place (Overall)": row_data[0],
"Place (Gender)": row_data[1],
"Place (Category)": row_data[2],
"Name": row_data[3],
"Sex": sex,
"Club": row_data[4],
"Running Number": row_data[5],
"Category": row_data[6],
"Finish": row_data[9],
"Year": 2021,
}
my_table.append(data_item)
df = pd.DataFrame(my_table).iloc[1:] # Strip table header
womens_results = womens_results.append(df)
# Concatenate results
results = pd.concat([mens_results, womens_results])
results
# +
# Some quick data cleaning
# Remove leftover titles
results["Club"] = results["Club"].str.replace("Club", "", regex=False)
results["Running Number"] = results["Running Number"].str.replace(
"Running Number", "", regex=False
)
results["Category"] = results["Category"].str.replace("Category", "", regex=False)
results["Finish"] = results["Finish"].str.replace("Finish", "", regex=False)
# Extract country groups, like (USA), from Name group
results["Country"] = results["Name"].str.extract(r"(\([A-Z]{3,}\))")
# Remove brackets in country
results["Country"] = results["Country"].str.replace(r"\(|\)", "", regex=True)
# Remove country group from name column
results["Name"] = results["Name"].str.replace(r"(\([A-Z]{3}\))", "", regex=True)
# Split first/lastname into new columns
LastFirst = results["Name"].str.split(pat=",", n=1, expand=True)
results["FirstName"], results["LastName"] = LastFirst[1], LastFirst[0]
# Remove comma from Name column, so that this can be saved as a CSV ----- Must happen after splitting Name into two cols!!
results["Name"] = results["Name"].str.replace(r"(\,)", "", regex=True)
# Replace non-standard '–' with NaN for missing vals
results = results.replace("–", np.nan)
results = results.replace("DSQ", np.nan)
results = results.replace("", np.nan)
# Delete odd race number row - table description not actual data
results = results.loc[
(results["Running Number"] != "RM9999") & (results["Running Number"] != "RF9999")
]
# Set data types
results = results.astype(
{
"Place (Overall)": "float64",
"Place (Gender)": "float64",
"Place (Category)": "float64",
"Name": str,
"Sex": str,
"Club": str,
"Running Number": str,
"Category": "category",
"Country": str,
"FirstName": str,
"LastName": str,
}
)
# Due to an irritating bug with converting objects to Int64, needed to first convert to float and then to int
results = results.astype(
{"Place (Overall)": "Int64", "Place (Gender)": "Int64", "Place (Category)": "Int64"}
)
results["Finish"] = pd.to_timedelta(results["Finish"])
results["Finish (Total Seconds)"] = results["Finish"].dt.total_seconds()
# -
results.info()
# And quickly save them in a csv
results.to_csv(
r"./London_2021.csv",
index=False,
header=True,
)
| notebooks/example_2020_scrape_london_marathon.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 2A.algo - Puzzles algorithmiques (1)
#
# Puzzles algorithmiques tirés de [Google Code Jam](https://code.google.com/codejam/) et autres sites équivalents, produits scalaires, problèmes de recouvrements, soudoyer les prisonniers, découpage stratifié.
from jyquickhelper import add_notebook_menu
add_notebook_menu()
# ## Produits scalaires
#
# Le problème est tiré de [Google Jam 2008, round 1A](https://code.google.com/codejam/contest/32016/dashboard#s=p0).
#
# On considère deux tableaux $v=(v_1,..., v_n)$ et $w=(w_1,...,w_n)$. On souhaite le minimum : $$\min_{\sigma,\sigma'} \sum_{i=1}^{n} v_{\sigma(i)} w_{\sigma'(i)}$$ où $\sigma,\sigma'$ sont deux permutations de l'ensemble $[[1,...,n]]$.
# ### Solution naïve
# ### Solution moins naïve
# ## Problème de recouvrement
#
# [Google Jam 2008, round 1A](https://code.google.com/codejam/contest/32016/dashboard#s=p0)
#
# Couvrir le segment $[0, M]$ avec le nombre minimum d'intervalles de la forme $[a_i, b_i]$ ? Avec $M, a_i b_i \in \mathbb{N}$.
#
# Exemple, couvrir $[0, 1]$ avec les intervalles $[-1, 0]$, $[-5, -3]$, $[2, 5]$.
# ## Soudoyer les prisonniers
#
# [Problem C. Bribe the Prisoners](https://code.google.com/codejam/contest/189252/dashboard#s=p2)
#
# Dans un royaume il y a des cellules de prison numérotées de 1 à $P$ construites de telle sorte à former un segment de ligne droite. Les cellules $i$ et $i + 1$ sont adjacentes et leur prisonniers sont appelés "voisins". Un mur muni d'une fenêtre les sépare et ils peuvent communiquer via cette fenêtre. Tous les prisonniers vivent en paix jusqu'a ce qu'un prisonnier soit relâché. Quand cela se produit, le prisonnier libéré fait part de la nouvelle à ses voisins, qui en parlent à leurs voisins, etc., jusqu'à atteindre la cellule 1 ou $P$ ou une cellule dont la cellule voisine est vide. Quand un prisonnier découvre qu'un autre prisonnier a été libéré, de colère, il casse tout dans sa cellule, sauf s'il a été préalablement soudoyé par une pièce
# d'or. Il faut donc veiller à soudoyer tous les prisonniers susceptibles de tout casser dans leur cellule avant de libérer un prisonnier. En supposant que toutes les cellules sont initialement occupées par un unique prisonnier et qu'un prisonnier par jour au plus puisse être relaché, et en connaissant la liste des $Q$ prisonniers à relâcher, il faut trouver l'ordre de libération des prisonniers de cette liste qui soit le moins coûteux en pièce d'or. Ordres de grandeur : $1 \leqslant P \leqslant 104$, $1 \leqslant Q \leqslant 102$. A noter que le soudoiement n'est actif qu'un seul jour.
#
# Exemple : 23 prisonniers, $P=20$, $Q=3$, les prisonniers à libérer sont les numéros $3, 6, 14$.
# ## Découpage intelligent d'une base de données
#
# On dispose d'une base de données à $N$ observations et $K$ variables $(X^1,...,X^K)$. On calcule la moyenne de chaque variable $\bar{X_k} =\frac{1}{N}\sum_{i=1}^N X^k_i$ sur l'ensemble de la base.
#
# On souhaite maintenant diviser la base en deux bases apprentissage de taille égale, test sous intersection. On mesure également la moyenne de chaque variable sur chacun des deux bases : $\bar{X^k}_a$ et $\bar{X^k}_t$. On souhaite effectuer un découpage de telle sorte que l'indicateur suivant soit minimum :
#
# $E = \sum_{k=1}^K \left( \bar{X^k_a} - \bar{X^k_t} \right)^2$
#
# Imaginer un algorithme qui effectue un tel découpage.
| _doc/notebooks/td2a_algo/td2a_cenonce_session_6A.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# $\newcommand{\xv}{\mathbf{x}}
# \newcommand{\wv}{\mathbf{w}}
# \newcommand{\yv}{\mathbf{y}}
# \newcommand{\zv}{\mathbf{z}}
# \newcommand{\uv}{\mathbf{u}}
# \newcommand{\vv}{\mathbf{v}}
# \newcommand{\Chi}{\mathcal{X}}
# \newcommand{\R}{\rm I\!R}
# \newcommand{\sign}{\text{sign}}
# \newcommand{\Tm}{\mathbf{T}}
# \newcommand{\Xm}{\mathbf{X}}
# \newcommand{\Zm}{\mathbf{Z}}
# \newcommand{\Im}{\mathbf{I}}
# \newcommand{\Um}{\mathbf{U}}
# \newcommand{\Vm}{\mathbf{V}}
# \newcommand{\muv}{\boldsymbol\mu}
# \newcommand{\Sigmav}{\boldsymbol\Sigma}
# \newcommand{\Lambdav}{\boldsymbol\Lambda}
# $
#
# # Machine Learning Methodology
#
# For machine learning algorithms, we learned how to set goas to optimize and how to reach or approach the optimal solutions. Now, let us discuss how to evaluate the learned models. There will be many different aspects that we need to consider not simply accuracy, so we will further discuss techniques to make the machine learning models better.
#
#
# ### Performance Measurement, Overfitting, Regularization, and Cross-Validation
#
# In machine learning, *what is a good measure to assess the quality of a machine learning model?* Let us step back from what we have learned in class about ML techniques and think about this.
# In previous lectures, we have discussed various measures such a `root mean square error (RMSE)`, `mean square error (MSE)`, `mean absolute error (MAE)` for **regression problems**, and `accuracy`, `confusion matrix`, `precision/recall`, `F1-score`, `receiver operating characteristic (ROC) curve`, and others for **classification**.
#
# For your references, here are the list of references for diverse metrics for different categories of machine learning.
# * Regressions: https://arxiv.org/pdf/1809.03006.pdf
# * Classification: As we have a cheatsheet already, here is a comprehensive version from ICMLA tutorial. https://www.icmla-conference.org/icmla11/PE_Tutorial.pdf
# * Clustering: https://scikit-learn.org/stable/modules/clustering.html#clustering-performance-evaluation
#
# Anyway, are these measures good enough to say a specific model is better than the other?
#
# Let us take a look at following codes examples and think about something that we are missing.
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
from copy import deepcopy as copy
x = np.arange(3)
t = copy(x)
# +
def plot_data():
plt.plot(x, t, "o", markersize=10)
plot_data()
# -
# I know that it is silly to apply a linear regression on this obvious model, but let us try. :)
# +
## Least Square solution: Filled codes here to fit and plot as the instructor's output
import numpy as np
# First creast X1 by adding 1's column to X
N = x.shape[0]
X1 = np.c_[np.ones((N, 1)), x]
# Next, using inverse, solve, lstsq function to get w*
w = np.linalg.inv(X1.transpose().dot(X1)).dot(X1.transpose()).dot(t)
# print(w)
y = X1.dot(w)
plot_data()
plt.plot(y)
# -
# Can we try a nonlinear model on this data? Why not?
# We can make a nonlinear model by simply adding higher degree terms such square, cubic, quartic, and so on.
#
# $$ f(\xv; \wv) = w_0 + w_1 \xv + w_2 \xv^2 + w_3 \xv^3 + \cdots $$
#
# This is called *polynomial regression* as we transform the input features to nonlinear by extending the features high dimensional with higher polynomial degree terms. For instance, your input feature $(1, x)$ is extended to $(1, x, x^2, x^3)$ for cubic polynomial regression model. After input transformation, you can simply use least squares or least mean squares to find the weights as the model is still linear with respect to the weight $\wv$.
#
# Let us make the polynomial regression model and fit to the data above with lease squares.
# Polinomial regression
def poly_regress(x, d=3, t=None, **params):
bnorm = params.pop('normalize', False)
X_poly = []
#######################################################################################
# Transform input features: append polynomial terms (from bias when i=0) to degree d
for i in range(d+1):
X_poly.append(x**i)
X_poly = np.vstack(X_poly).T
# normalize
if bnorm:
mu, sd = np.mean(X_poly[:, 1:, None], axis=0), np.std(X_poly[:, 1:, None], axis=0)
X_poly[:, 1:] = (X_poly[:, 1:] - mu.flat) / sd.flat
# least sqaures
if t is not None:
# added least square solution here
w = np.linalg.inv(X_poly.transpose().dot(X_poly)).dot(X_poly.transpose()).dot(t)
if bnorm:
return X_poly, mu, sd, w
return X_poly, w
if bnorm:
return X_poly, mu, sd
return X_poly
# The poly_regress() function trains with the data when target input is given after transform the input x as the following example. The function also returns the transformed input X_poly.
# +
Xp, wp = poly_regress(x, 3, t)
print(wp.shape)
print(Xp.shape)
yp = Xp @ wp
plot_data()
plt.plot(x, y)
plt.plot(x, yp)
# -
# Hmm... They both look good on this. Then, what is the difference? Let us take a look at how they change if I add the test data. If I compare the MSE, they are equivalent. Try to expand the data for test and see how different they are.
#
# Here, we use another usage of poly_regress() function without passing target, so we transform the target input to polynomial features.
# +
xtest = np.arange(11)-5
Xptest = poly_regress(xtest, 3)
yptest = Xptest @ wp
X1test = np.vstack((np.ones(len(xtest)), xtest)).T
ytest = X1test @ w
plot_data()
plt.plot(xtest, ytest)
plt.plot(xtest, yptest)
# -
# Here the orange is th linear model and the green line is 3rd degree polynomial regression.
# Which model looks better? What is your pick?
#
# <br/><br/><br/><br/><br/><br/>
# ## Learning Curve
#
# From the above example, we realized that the model evaluation we discussed so far is not enough.
#
# First, let us consider how well a learned model generalizes to new data with respect to the number of training samples. We assume that the test data are drawn from same distribution over example space as training data.
#
#
# In this plot, we can compare the two learning algorithms and find which one generalizes better than the other. Also, during the training, we can access to the training error (or empirical loss). This may not look similar (mostly not) to the test error (generalization loss above).
#
# Let us take a look at the example in the Geron textbook.
# +
import os
import pandas as pd
import sklearn
def prepare_country_stats(oecd_bli, gdp_per_capita):
oecd_bli = oecd_bli[oecd_bli["INEQUALITY"]=="TOT"]
oecd_bli = oecd_bli.pivot(index="Country", columns="Indicator", values="Value")
gdp_per_capita.rename(columns={"2015": "GDP per capita"}, inplace=True)
gdp_per_capita.set_index("Country", inplace=True)
full_country_stats = pd.merge(left=oecd_bli, right=gdp_per_capita,
left_index=True, right_index=True)
full_country_stats.sort_values(by="GDP per capita", inplace=True)
remove_indices = [0, 1, 6, 8, 33, 34, 35]
keep_indices = list(set(range(36)) - set(remove_indices))
return full_country_stats[["GDP per capita", 'Life satisfaction']].iloc[keep_indices], full_country_stats[["GDP per capita", 'Life satisfaction']]
# -
# !curl https://raw.githubusercontent.com/ageron/handson-ml/master/datasets/lifesat/oecd_bli_2015.csv > oecd_bli_2015.csv
# !curl https://raw.githubusercontent.com/ageron/handson-ml/master/datasets/lifesat/gdp_per_capita.csv > gdp_per_capita.csv
# +
# Load the data
oecd_bli = pd.read_csv("oecd_bli_2015.csv", thousands=',')
gdp_per_capita = pd.read_csv("gdp_per_capita.csv",thousands=',',delimiter='\t',
encoding='latin1', na_values="n/a")
# Prepare the data
country_stats, full_country_stats = prepare_country_stats(oecd_bli, gdp_per_capita)
X = np.c_[country_stats["GDP per capita"]]
y = np.c_[country_stats["Life satisfaction"]]
# Visualize the data
country_stats.plot(kind='scatter', x="GDP per capita", y='Life satisfaction', figsize=(6.5,4))
plt.axis([0, 60000, 0, 10])
plt.show()
# -
# This looks like the data showing a linear trend. Now, let us extend the x-axis to further to 110K and see how it looks.
# Visualize the full data
# Visualize the data
full_country_stats.plot(kind='scatter', x="GDP per capita", y='Life satisfaction', figsize=(12,4))
plt.axis([0, 110000, 0, 10])
plt.show()
# Maybe a few outliers with high GDP do not follow the linear trend that we observed above.
# +
# Data for training
Xfull = np.c_[full_country_stats["GDP per capita"]]
yfull = np.c_[full_country_stats["Life satisfaction"]]
print(Xfull.shape, yfull.shape)
# -
# We can better observe the trend by fitting polynomial regression models by changing the degrees.
# +
# polynomial model to this data
for deg in [1, 2, 5, 10, 30]:
plt.figure();
full_country_stats.plot(kind='scatter', x="GDP per capita", y='Life satisfaction',
figsize=(12,4));
plt.axis([0, 110000, 3, 10]);
Xp, mu, sd, wp = poly_regress(Xfull.flatten(), deg, yfull.flatten(), normalize=True)
yp1 = Xp @ wp
# plot curve
plt.plot(Xfull, yp1, 'r-', label=deg);
plt.title("degree: {}".format(deg));
# -
# What degree do you think the data follow? What is your best pick? Do you see overfitting here? From which one do you see it?
# As the complexity of model grows, you may have small training errors. However, there is no guarantee that you have a good generalization (you may have very bad generalization error!).
# This is called **Overfitting** problem in machine learning. From training data, once you learned the hypothesis *h* (or machine learning model), you can have training error $E_{train}(h)$ and testing error $E_{test}(h)$. Let us say that there is another model $h^\prime$ for which
#
# $$ E_{train}(h) < E_{train}(h^\prime) \wedge E_{test}(h) > E_{test}(h^\prime).$$
#
# Then, we say the hypothesis $h$ is "overfitted."
# ## Bias-Variance Tradeoff
#
# Here the bias refers an error from erroneous assumptions and the variance means an error from sensitivity to small variation in the data. Thus, high bias can cause an underfitted model and high variance can cause an overfitted model. Finding the sweet spot that have good generalization is on our hand.
#
# In the same track of discussion, Scott summarizes the errors that we need to consider as follows:
#
# - high bias error: under-performing model that misses the important trends
# - high variance error: excessively sensitive to small variations in the training data
# - Irreducible error: genuine to the noise in the data. Need to clean up the data
#
# 
# <center>From Understanding the Bias-Variance Tradeoff, by <NAME></center>
# ## Regularization
#
# We reduce overfitting by addding a complexity penalty to the loss function. Here follows the loss function for the linear regression with $L2$-norm.
#
# $$
# \begin{align*}
# E(\wv) &= \sum_i^N ( y_i - t_i)^2 + \lambda \lVert \wv \rVert_2^2 \\
# \\
# &= \sum_i^N ( y_i - t_i)^2 + \lambda \sum_k^D w_k^2 \\
# \\
# &= (\Xm \wv - T)^\top (\Xm \wv - T) + \lambda \wv^\top \wv \\
# \\
# &= \wv^\top \Xm^\top \Xm \wv - 2 \Tm^\top \Xm \wv + \Tm^\top \Tm + \lambda \wv^\top \wv
# \end{align*}
# $$
#
# Repeating the derivation as in linear regression,
#
# $$
# \begin{align*}
# \frac{\partial E(\wv)}{\partial \wv} &= \frac{\partial (\Xm \wv - \Tm)^\top (\Xm \wv - \Tm)}{\partial \wv} + \frac{\partial \lambda \wv^\top \wv}{\partial \wv} \\
# \\
# &= 2 \Xm^\top \Xm \wv - 2 \Xm^\top \Tm + 2 \lambda \wv
# \end{align*}
# $$
#
# Setting the last term zero, we reach the solution of *ridge regression*:
#
# $$
# \begin{align*}
# 2 \Xm^\top \Xm \wv - 2 \Xm^\top \Tm + 2 \lambda \wv &= 0\\
# \\
# \big(\Xm^\top \Xm + \lambda \Im \big) \wv &= \Xm^\top \Tm\\
# \\
# \wv &= \big(\Xm^\top \Xm + \lambda \Im \big)^{-1} \Xm^\top \Tm.
# \end{align*}
# $$
#
# ## Cross-Validation
#
# Now, let us select a model. Even with the regularization, we still need to pick $\lambda$. For polynomial regression, we need to find the degree parameter. When we are mix-using multiple algorithms, we still need to know which model to choose.
# Here, remembe that we want a model that have good generalization. The idea is preparing one dataset (a validation set) by pretending that we cannot see the labels. After choosing a model parameter (or a model) and train it with training dataset, we test it on the validation data.
# Comparing the validation error, we select the one that has the lowest validation error. Finally, we evaluate the model on testing data.
#
# Here follows the K-fold cross-validation that divides the data into K blocks for traing, validating and testing.
#
# ### K-fold CV Procedure
#
# 
# ## Feature Selection
#
# Another way to get a sparse (possibly good generalization) model is using small set of most relevant features. Weight analysis or some other tools can give us what is most relevant or irrelevant features w.r.t the training error. But it is still hard to tell the relevance to the generalization error. Thus, problems in choosing a minimally relevant set of features is NP-hard even with perfect estimation of generalization error. With inaccurate estimation that we have, it is much hard to find them.
#
# Thus, we can simply use cross-validation to find features.
# We can greedily add (forward selection) or delete (backward selection) features that decrease cross-validation error most.
# # Practice
#
# Now, try to write your own 5-fold cross validation code that follows the procedure above for ridge regression. Try 5 different $\lambda$ values, [0, 0.01, 0.1, 1, 10], for this.
# +
# TODO: try to implement your own K-fold CV.
# (This will be a part of next assignment (no solution will be provided.))
| reading_assignments/5_Note-ML Methodology.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Analyzing data with Dask, SQL, and Coiled
#
# In this notebook, we look at using [Dask-SQL](https://dask-sql.readthedocs.io/en/latest/), an exciting new open-source library which adds a SQL query layer on top of Dask. This allows you to query and transform Dask DataFrames using common SQL operations.
#
# ## Launch a cluster
#
# Let's first start by creating a Coiled cluster which uses the `examples/dask-sql` software environment, which has `dask`, `pandas`, `s3fs`, and a few other libraries installed.
# +
import coiled
cluster = coiled.Cluster(
n_workers=10,
worker_cpu=4,
worker_memory="30GiB",
software="examples/dask-sql",
)
cluster
# -
# and then connect Dask to our remote Coiled cluster
# +
from dask.distributed import Client
client = Client(cluster)
client.wait_for_workers(10)
client
# -
# ## Getting started with Dask-SQL
#
# Internally, Dask-SQL uses a well-established Java library, Apache Calcite, to parse SQL and perform some initial work on your query. To help Dask-SQL locate JVM shared libraries, we set the `JAVA_HOME` environment variable.
# +
import os
os.environ["JAVA_HOME"] = os.environ["CONDA_DIR"]
# -
# The main interface for interacting with Dask-SQL is the `dask_sql.Context` object. It allows your to register Dask DataFrames as data sources and can convert SQL queries to Dask DataFrame operations.
# +
from dask_sql import Context
c = Context()
# -
# For this notebook, we'll use the NYC taxi dataset, which is publically accessible on AWS S3, as our data source
# +
import dask.dataframe as dd
from distributed import wait
df = dd.read_csv(
"s3://nyc-tlc/trip data/yellow_tripdata_2019-*.csv",
dtype={
"payment_type": "UInt8",
"VendorID": "UInt8",
"passenger_count": "UInt8",
"RatecodeID": "UInt8",
},
storage_options={"anon": True}
)
# Load datasest into the cluster's distributed memory.
# This isn't strictly necessary, but does allow us to
# avoid repeated running the same I/O operations.
df = df.persist()
wait(df);
# -
# We can then use our `dask_sql.Context` to assign a table name to this DataFrame, and then use that table name within SQL queries
# +
# Registers our Dask DataFrame df as a table with the name "taxi"
c.register_dask_table(df, "taxi")
# Perform a SQL operation on the "taxi" table
result = c.sql("SELECT count(1) FROM taxi")
result
# -
# Note that this returned another Dask DataFrame and no computation has been run yet. This is similar to other Dask DataFrame operations, which are lazily evaluated. We can call `.compute()` to run the computation on our cluster.
result.compute()
# Hooray, we've run our first SQL query with Dask-SQL! Let's try out some more complex queries.
#
# ## More complex SQL examples
#
# With Dask-SQL we can run more complex SQL statements like, for example, a groupby-aggregation:
c.sql('SELECT avg(tip_amount) FROM taxi GROUP BY passenger_count').compute()
# NOTE: that the equivalent operatation using the Dask DataFrame API would be:
#
# ```python
# df.groupby("passenger_count").tip_amount.mean().compute()
# ```
# We can even make plots of our SQL query results for near-real-time interactive data exploration and visualization.
c.sql("""
SELECT floor(trip_distance) AS dist, avg(fare_amount) as fare
FROM taxi
WHERE trip_distance < 50 AND trip_distance >= 0
GROUP BY floor(trip_distance)
""").compute().plot(x="dist", y="fare");
# If you would like to learn more about Dask-SQL check out the [Dask-SQL docs](https://dask-sql.readthedocs.io/) or [source code](https://github.com/nils-braun/dask-sql) on GitHub.
| dask-sql/dask-sql.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lecture 3, Nonlinear dynamics, stability and bifurcations
#
# > ordinary differential equations (ODE)
#
# - toc: False
# - badges: true
# - comments: False
# - categories: [jupyter]
# Thanks a lot to [<NAME>](https://github.com/dpsanders) for providing the lecture. The original lecture is part of the MIT class [Introduction to Computational Thinking](https://computationalthinking.mit.edu/Fall20/lecture20/).
#
# This class uses the [Julia programming language](http://www.julialang.org/). The orignal code can be found under [github.com](https://github.com/mitmath/18S191/blob/master/lecture_notebooks/week11/nonlinear_dynamics_bifurcations.jl)
# +
import numpy as np
import matplotlib.pyplot as plt
from ipywidgets import interact, interactive, fixed, interact_manual
import ipywidgets as widgets
from IPython.display import HTML
from IPython.display import display
# -
import warnings
warnings.filterwarnings("ignore")
# %matplotlib inline
# How does the climate change over time? In the last lecture we saw that our simple model is already quite good simulating how the climate changes over time.
#
# Our simple model is based on *ordinary differential equations (ODEs)*, where some variables change in time - with the rate of change as function of their current values.
#
# $$ \frac{dx(t)}{dt} = f(x(t)) $$
#
# The simplest numerical method to solve such an equation is the **(forward) Euler method**, in which we convert this equation into an explicit time-stepping routine:
#
# $$ \frac{dx(t)}{dt} = \frac{x(t+\Delta t) - x(t)}{\Delta t}$$
#
#
# with the approximation
#
# $$ x(t+\Delta t) \simeq x(t) + \Delta t f(x(t)) $$
# ## Solving the ODE: Euler method
#
# Let's use this to simulate a simple nonlinear ODE that describes the dynamics of a population of bacteria. The bacteria will grow by reproduction at a rate $\lambda$ provided there is sufficient food, in which case we would have $\dot{x} = \lambda x$. But the available food will actually always limit the sustainable population to a value $K$. A simple model for this is as follows:
#
# $$\dot{x} = \lambda \, x \, (K - x).$$
#
# When $x$ is close to $0$, the growth rate is $\lambda$, but that rate decreases as $x$ increases.
#
# This is sometimes called the [**logistic** differential equation](https://en.wikipedia.org/wiki/Logistic_function#Logistic_differential_equation) (although the name does not seem particularly helpful).
#
#
# Our goal is to use computational thinking, but we will actually not be interested so much in the exact dynamics in time, but rather in the **qualitative** features of the behaviour of the system. For example, at long times (formally $t \to \infty$) does the population get arbitrarily large? Or does it, for example, oscillate around a particular value? Or does it converge to a particular size? This forms the subject of **nonlinear dynamics** or **dynamical systems** theory.
#
#
# Let's simulate the system using the Euler method to try to guess the answer to this question. We should never use the Euler method in practice, but should use a tried and tested library instead, and algorithms that provide much better accuracy in the solutions, if we are interested in faithful numerical results.
#
# We'll rescale the variables to the simplest form:
#
# $$ \frac{dx}{dt} = x\,(1-x) $$
#
#
# $$ x_{n+1} = x_n + \Delta t \cdot \text{tendency}(x_n; ...)$$
def logistic(x, timesteps):
x = np.asarray(x)
for i in np.diff(timesteps):
if x.size == 1:
x = np.append(x, x + i * x * (1-x))
else:
x = np.append(x, x[-1] + i * x[-1] * (1-x[-1]))
return x
t1 = np.arange(0,20, 0.01)
dxdt = logistic(0.5, t1)
plt.plot(t1, dxdt)
# We see that for this particular initial condition, the solution seems to settle down to a fixed value after some time, and then remains at that value thereafter.
# Such a value is called a **fixed point** or a **stationary point** of the ODE.
# ## Qualitative behaviour: Fixed points and their stability
# But what happens if we have a different initial condition:
# +
tag = HTML('''<script>
code_show=true;
function code_toggle() {
if (code_show){
$('div.cell.code_cell.rendered.selected div.input').hide();
} else {
$('div.cell.code_cell.rendered.selected div.input').show();
}
code_show = !code_show
}
$( document ).ready(code_toggle);
</script>
To show/hide this cell's raw code input, click <a href="javascript:code_toggle()">here</a>.''')
display(tag)
def plot_func(initial_condition):
plt.figure(figsize=(8,8))
plt.plot(t1, logistic(initial_condition, t1))
plt.xlabel("t")
plt.ylabel("x(t)")
plt.xlim([0,10])
plt.ylim(-1, 2)
plt.grid()
plt.show()
interact(plot_func, initial_condition = widgets.FloatSlider(value = 0.5,
min = -1,
max = 2,
step = 0.1))
# -
# To get an overview, we can draw all graphs in a single plot.
plt.figure(figsize=(8,8))
for initial_condition in np.arange(-1, 2, 0.1):
plt.plot(t1, logistic(initial_condition, t1))
plt.xlabel("t")
plt.ylabel("x(t)")
plt.xlim([0,10])
plt.ylim(-1, 2)
plt.grid()
plt.show()
# We see that all the curves starting near to $x_0=1.0$ seem to converge to 1 at long times. If the system starts *exactly* at 0 then it stays there forever. However, if it starts close to 0, on either side, then it moves *away* from 0 (on that same side of 0) -- starting from a negative value $x$ becomes ever more negative. (Even though negative populations have no meaning in the original interpretation as the dynamics of a population, we can still ask study the dynamics of the equation with negative initial conditions, since it may model other systems too.)
#
# The special values $x^*_1=1$ and $x^*_2=0$ are called **stationary points** or **fixed points** of the differential equation. If we start at $x^*_i$, then the derivative there is $f'(x^*_i) = 0$, and hence we cannot move away from $x^*_i$! The fixed points can be found as zeros or **roots** of the function $f$, i.e. values $x^*$ such that $f(x^*) = 0$.
#
# We see, though, that the two types of fixed points are **qualitatively different**: trajectories that start close to $x^*_1 = 1$ move *towards* $x^*_1$, whereas trajectories that start close to $x^*_2 = 0$ move *away* from it. We say that $x^*_1$ is a **stable fixed point** and $x^*_2$ is an **unstable fixed point**.
#
# In general it is not possible to find analytical formulas for the position and stability of fixed points; instead, we can use numerical **root-finding algorithms**, for example the Newton method.
#
# ## State space: Vector field and phase portrait
#
#
# If we want to find the whole trajectory for a given initial condition then we need to solve the equations, either numerically or analytically.
#
# However, we may want less information about the system, for example the **long-time** or **asymptotic** dynamics. It turns out that we can obtain some information about that *without* explicitly solving the ODE! This is the **qualitative approach** to studying nonlinear systems.
#
# Instead of drawing trajectories $x(t)$ as a function of time $t$, as we did above, let's use a different graphical representation, where we draw **state space** or **phase space**: This is the set ("space") of all possible values of the dependent variables ("states"). For the above ODE there is only a single dependent variable, $x$, so the state space is the real line, $\mathbb{R}$.
#
# At each possible value of $x$, the ODE gives us information about the rate of change of $x(t)$ at that point. Let's draw an **arrow** at that point, pointing in the direction that a particle placed at that point would move: to the right if $\dot{x} > 0$ and to the left if $\dot{x} < 0$.
# +
tag = HTML('''<script>
code_show=true;
function code_toggle() {
if (code_show){
$('div.cell.code_cell.rendered.selected div.input').hide();
} else {
$('div.cell.code_cell.rendered.selected div.input').show();
}
code_show = !code_show
}
$( document ).ready(code_toggle);
</script>
To show/hide this cell's raw code input, click <a href="javascript:code_toggle()">here</a>.''')
display(tag)
states = np.array([])
initial_conditions = np.arange(-1, 2, 0.1)
for initial_condition in initial_conditions:
states = np.append(states, logistic(initial_condition, t1)[-1])
X = np.ones(len(states))
Y = initial_conditions.copy()
U = np.zeros(len(states))
V = np.ones(len(states))
V[states - initial_conditions < 0] = -1
states[states == -np.inf] = 2
plt.figure(figsize=(2,8))
plt.quiver(X, Y, U, V, scale = 10, width = 0.02)
plt.plot([1,1],[1,1], marker = 'o', markersize = 12)
plt.plot([1,1],[0,0], marker = 'o', markersize = 12)
plt.xlim([1,1])
plt.xlabel([])
# -
# This vector field indeed gives us a *qualitative* picture of the dynamics. It does not tell us how fast the dynamics will occur in each region, but it indicates what the *tendency* is. We have coded the fixed points according to their stability; this may be calculated using the derivative evaluated at the fixed point, $f'(x^*)$, since this derivative controls the behaviour of nearby initial conditions $x^* + \delta x$.
# ## Bifurcations
#
# Now suppose that there is a **parameter** $\mu$ in the system that can be varied. For each value of $\mu$ we have a *different* ODE
#
# $$\dot{x} = f_\mu(x).$$
# For example,
# $$\dot{x} = \mu + x^2.$$
# Let's draw the state space for each different value of $\mu$:
# +
tag = HTML('''<script>
code_show=true;
function code_toggle() {
if (code_show){
$('div.cell.code_cell.rendered.selected div.input').hide();
} else {
$('div.cell.code_cell.rendered.selected div.input').show();
}
code_show = !code_show
}
$( document ).ready(code_toggle);
</script>
To show/hide this cell's raw code input, click <a href="javascript:code_toggle()">here</a>.''')
display(tag)
def xdot(mu, x):
return mu + x**2
t = np.arange(-5, 5, 0.001)
def plot_func(initial_condition):
plt.figure(figsize=(8,8))
a = xdot(mu = initial_condition, x = t)
plt.plot(t, a)
plt.xlabel("t")
plt.ylabel("x(t)")
plt.xlim([-5,5])
plt.ylim(-2, 2)
plt.grid()
try:
zero_crossings = np.where(np.diff(np.signbit(a)))[0]
plt.vlines(t[zero_crossings+1], ymin=-2,ymax=2)
for arrows in np.arange(-5,t[zero_crossings[0]],0.5):
plt.arrow(arrows, 0, 0.25, 0,shape='full', lw=1, length_includes_head=True, head_width=.05, color ="blue")
for arrows in np.arange(t[zero_crossings[0]]+0.5, t[zero_crossings[1]],0.5):
plt.arrow(arrows, 0, -0.25, 0,shape='full', lw=1, length_includes_head=True, head_width=.05, color = "blue")
for arrows in np.arange(t[zero_crossings[1]], 5,0.5):
plt.arrow(arrows, 0, +0.25, 0,shape='full', lw=1, length_includes_head=True, head_width=.05, color ="red")
plt.plot([t[zero_crossings],t[zero_crossings]],[0,0], marker='o', markersize = 12)
except:
for arrows in np.arange(-5,5,0.5):
plt.arrow(arrows, 0, 0.25, 0,shape='full',
lw=1, length_includes_head=True, head_width=.05, color ="blue")
plt.show()
interact(plot_func, initial_condition = widgets.FloatSlider(value = -1,
min = -2,
max = 2,
step = 0.1))
# -
# Now let's collect all the vector fields into a single plot. We *rotate* the vector field to now be vertical, thinking of the dynamics of $x$ as occurring along the vertical direction. The horizontal axis now represents the different possible values of the parameter $\mu$:
# +
tag = HTML('''<script>
code_show=true;
function code_toggle() {
if (code_show){
$('div.cell.code_cell.rendered.selected div.input').hide();
} else {
$('div.cell.code_cell.rendered.selected div.input').show();
}
code_show = !code_show
}
$( document ).ready(code_toggle);
</script>
To show/hide this cell's raw code input, click <a href="javascript:code_toggle()">here</a>.''')
display(tag)
saddle_points0 = np.array([])
saddle_points1 = np.array([])
stepsize = 0.1
for mu in np.arange(-2, 2, stepsize):
a = xdot(mu = mu, x = t)
zero_crossings = np.where(np.diff(np.signbit(a)))[0]
if zero_crossings.size > 1:
saddle_points0 = np.append(saddle_points0, t[zero_crossings[0]])
saddle_points1 = np.append(saddle_points1, t[zero_crossings[1]])
for arrows in np.arange(-5,t[zero_crossings[0]],0.25):
plt.arrow(mu, arrows, 0.0, 0.1,shape='full', lw=1, length_includes_head=True, head_width=.025, color ="blue")
for arrows in np.arange(t[zero_crossings[0]], t[zero_crossings[1]],0.25):
plt.arrow(mu, arrows, 0.0, -0.1,shape='full', lw=1, length_includes_head=True, head_width=.025, color ="red")
for arrows in np.arange(t[zero_crossings[1]],2,0.25):
plt.arrow(mu, arrows, 0.0, 0.1,shape='full', lw=1, length_includes_head=True, head_width=.025, color ="blue")
elif zero_crossings.size == 1:
saddle_points0 = np.append(saddle_points0, t[zero_crossings])
saddle_points1 = np.append(saddle_points0, np.nan)
else:
saddle_points0 = np.append(saddle_points0, np.nan)
saddle_points1 = np.append(saddle_points1, np.nan)
for arrows in np.arange(-2,2,0.25):
plt.arrow(mu, arrows, 0.0, 0.1,shape='full', lw=1, length_includes_head=True, head_width=.025, color ="blue")
plt.ylim(-2,2)
plt.xlim(-2,2)
plt.scatter(np.arange(-2,2,stepsize), saddle_points0, color="green", marker='D')
plt.scatter(np.arange(-2,2,stepsize), saddle_points1, color="black", marker='o')
plt.ylabel("fixed points and dynamics with given $\mu$")
plt.xlabel("$\mu$")
# -
# Now let's collect all the vector fields into a single plot. We *rotate* the vector field to now be vertical, thinking of the dynamics of $x$ as occurring along the vertical direction. The horizontal axis now represents the different possible values of the parameter $\mu$:
#
# We see that at the **critical value** $\mu_c = 0$ there is a **qualitative change in behaviour** in the system: for $\mu_c < 0$ there are two fixed points, whereas for $\mu_c > 0$ there are no fixed points at all. In this particular ODE the two fixed points collide in a **saddle--node** or **fold** bifurcation.
#
# ## Bistability and hysteresis
#
# Now let's look at the dynamics of the following system:
#
#
# $$\dot{x} = \mu + x - x^3.$$
#
#
def h(mu, x):
return mu + x - x**3
# +
saddle_points = []
stepsize = 0.1
for mu in (np.arange(-2, 2, stepsize)):
a = h(mu = mu, x = t)
zero_crossings = np.where(np.diff(np.signbit(a)))[0]
plt.scatter(np.ones(len(zero_crossings))*mu, t[zero_crossings], color="green", marker='D')
plt.grid()
plt.title("Bifurcation Diagramm")
plt.ylabel("fixed points and dynamics with given $\mu$")
plt.xlabel("$\mu$")
plt.ylim(-2,2)
plt.xlim(-2,2)
# -
# We see that there is a range of values of $\mu$ for which there are *three coexisting fixed points*, two stable and one unstable. Since there are two stable fixed points in which the system can remain, we say that the system is **bistable**.
#
#
# Now that we understand what the plots mean and the dynamics, let's plot just the fixed points $x^*(\mu)$ as a function of $\mu$. Such a plot is called a **bifurcation diagram**:
#
# The pieces of curve are called **branches**.
#
#
#
# ## Hysteresis
#
#
#
# Suppose we now think about slowly varying the parameter $\mu$. If we change the parameter $\mu$ by a little, the system is no longer at a fixed point, since the position of the fixed point moves when $\mu$ changes. However, the system will then **relax** by following the dynamics at the new value of $\mu$, and will rapidly converge to the new fixed point nearby.
# For example, starting at $\mu=-2$, the system will stay on the lower black (stable) **branch** until $\mu=0.4$ or so. At that point, two fixed points collide and annihilate each other! After that there is no longer a fixed point nearby. However, there is another fixed point much further up that will now attract all trajectories, so the system rapidly transitions to that fixed point.
# Now suppose we decrease the parameter again. The system will now track the *upper* branch until $\mu=-0.4$ or so, when again it will jump back down.
# For each parameter value $\mu$ in the interval $[-0.4, 0.4]$ there is **bistability**, i.e. **coexistence** of *two* fixed points with the same value of $\mu$ (together with a third, unstable fixed point that is not observable).
# The fact that the system tracks different stable branches depending on where we started, i.e. on the history, is known as **hysteresis**.
#
#
# Hysteretic behaviour like this is found in many scientific and engineering contexts, including switches in biology, for example genetic switches, and in the historical dynamics of the earth's climate.
#
#
#
# ## Slow--fast systems
#
# What are we actually doing when we let the parameter $\mu$ vary? Effectively we now have a system with *two* equations, for example
#
# $$\dot{x} = \mu + x - x^3;$$
# $$\dot{\mu} = \epsilon,$$
#
# where $\mu$ varies at some slow speed $\epsilon$. On a time scale much shorter than $1 / \epsilon$, the dynamics of $x$ "does not know" that $\mu$ is changing, so it will converge to a fixed point $x^*(\mu)$ for the current value of $\mu$. [An associated term is **adiabatic approximation**.] However, $\mu$ does gradually change, so the value of $x$ will effectively "slide along" the curve $x(t) \simeq x^*(\mu(t))$, tracking the curve of fixed points as $\mu$ changes.
# Once $\mu$ reaches a critical value $\mu_c$, however, there is no longer a nearby fixed point, and the dynamics will rapidly transition to the far away alternative fixed point.
# If we now reverse the dynamics of $\mu$, we slide back along the upper branch.
# """
| _notebooks/2020-11-12-non-linear-dynamics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python3 (test)
# language: python
# name: test
# ---
# +
# %matplotlib notebook
# %pylab inline
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn import preprocessing
import scanpy as sc
from scipy.cluster.hierarchy import dendrogram
from scipy.cluster.hierarchy import linkage
mpl.rcParams['pdf.fonttype'] = 42
mpl.rcParams['ps.fonttype'] = 42
mpl.rcParams['axes.grid'] = False
# -
pathToCounts = './counts.h5ad'
pathToCellData = './cell_annotation.csv'
counts = sc.read(pathToCounts)
cell_annotation = pd.read_csv(pathToCellData,index_col = 0)
counts.obs = cell_annotation.loc[counts.obs.index,:]
# ### Panel a
cmap_subclass = dict(zip(cell_annotation.subclass, cell_annotation.color_subclass))
# +
#Plot umap
x = cell_annotation.umap_x
y = cell_annotation.umap_y
fig = plt.figure(figsize = (4,4))
sns.set(font_scale=1)
sns.set_style("white")
ax1 = fig.add_subplot(111)
ax1.scatter(x, y, c=cell_annotation["subclass"].apply(lambda x: cmap_subclass[x]), alpha=0.6, s=0.05, rasterized = True)
plt.yticks([])
plt.xticks([])
plt.ylabel("UMAP2")
plt.xlabel("UMAP1")
ax1.axis('equal')
#f.savefig('./umap.pdf',dpi = 300, bbox_inches = 'tight', transparent=False)
plt.show()
# -
# ### Panel b
countsDF = pd.DataFrame(counts.X, index = counts.obs.index.values, columns = counts.var.index.values)
countsDF['label'] = counts.obs['label']
countsMean = countsDF.groupby('label').mean()
countsMeanZ = pd.DataFrame(preprocessing.scale(countsMean), index = countsMean.index, columns = countsMean.columns)
Z = linkage(countsMeanZ, 'ward')
groups = counts.obs.loc[:,['subclass','label']].groupby(['subclass','label'])
groupDict = dict()
for k,v in groups:
groupDict[k[1]] = k[0]
# class_label_colors = {'Other':'y', 'Glutamatergic':'m', 'GABAergic':'c'}
# +
# Color mapping
dflt_col = "k" # Unclustered gray
D_leaf_colors = dict(zip(countsMeanZ.index, [[cmap_subclass[groupDict[x]]] for x in countsMeanZ.index]))
numToType = dict(zip(range(len(countsMeanZ.index.values)), countsMeanZ.index.values))
# notes:
# * rows in Z correspond to "inverted U" links that connect clusters
# * rows are ordered by increasing distance
# * if the colors of the connected clusters match, use that color for link
link_cols = {}
for i, i12 in enumerate(Z[:,:2].astype(int)):
c1, c2 = (link_cols[x] if x > len(Z) else D_leaf_colors[numToType[x]][0]
for x in i12)
link_cols[i+1+len(Z)] = c1 if c1 == c2 else dflt_col
# Dendrogram
f,axs = plt.subplots(1,1,figsize = (20,5))
D = dendrogram(Z=Z, labels=countsMeanZ.index, color_threshold=None,
leaf_font_size=12, leaf_rotation=90, link_color_func=lambda x: link_cols[x])
#f.savefig('./dend.pdf',dpi = 300, bbox_inches = 'tight', transparent=False)
# -
# ### Panel e
pathToTracingData = './tracing_panel_e.csv'
TracingData = pd.read_csv(pathToTracingData,index_col = 0)
# +
clusterToPlot = [ 'L23_IT_1','L23_IT_2','L23_IT_3', 'L23_IT_4', 'L23_IT_5','L45_IT_SSp_1','L45_IT_SSp_2','L45_IT_1', 'L45_IT_2', 'L45_IT_3','L45_IT_4', 'L45_IT_5', 'L5_IT_1','L5_IT_2', 'L5_IT_3', 'L5_IT_4', 'L6_IT_1','L6_IT_2','L6_IT_3', 'L6b_1', 'L6b_2','L6b_3']
targets = ['MOs','SSp','TEa/ECT/PERI']
xDict = dict(zip(clusterToPlot, range(len(clusterToPlot))))
yDict = dict(zip(targets, range(len(targets))))
TracingData['x'] = TracingData['cluster'].map(xDict)
TracingData['y'] = TracingData['target'].map(yDict)
# +
f,axs = plt.subplots(1,1,figsize = (7,7*0.35))
maxSize = 150
minSize = maxSize * 0.1
TracingData['size_scaled'] = TracingData['fraction_cluster'] * maxSize
mask = TracingData['size_scaled'] < minSize
TracingData['size_scaled'] = TracingData['size_scaled'].where(~mask, other = minSize)
plt.scatter(TracingData['x'], TracingData['y'],
c = TracingData['fraction_target'], cmap = 'RdPu',vmin = 0, vmax = 0.2,
s = TracingData['size_scaled'], edgecolors='none')
axs.set_ylim(len(targets)+0.5,-0.5)
axs.set_xlim(-1, len(clusterToPlot)+2)
axs.spines['top'].set_visible(False)
axs.spines['right'].set_visible(False)
axs.spines['bottom'].set_visible(False)
axs.spines['left'].set_visible(False)
axs.get_xaxis().set_ticks([])
axs.set_yticks(range(len(targets)))
axs.set_yticklabels(targets, fontsize = 9)
#plt.colorbar()
#f.savefig('./dot.pdf',dpi = 300, bbox_inches = 'tight', transparent=False)
# -
| flagship/Fig3_MERFISH.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: tl_env
# language: python
# name: tl_env
# ---
# +
import pandas as pd
import os
import glob
import warnings
warnings.simplefilter(action='ignore', category=FutureWarning)
import numpy as np
import sklearn.metrics
from collections import defaultdict
import shutil
import pickle
pd.reset_option('all')
# -
input_file_path = '/Users/amandeep/Github/wikidata-wikifier/wikifier/sample_files/cricketers.csv'
wikify_column_name = "cricketers"
output_path = '/tmp/cricketers'
es_index = 'wikidatadwd-augmented-04'
es_url = 'http://ckg07:9200'
temp_dir = f'{output_path}/temp'
# !mkdir -p $output_path
# !mkdir -p $temp_dir
# +
#intermediate files
canonical = f'{temp_dir}/canonical.csv'
candidates = f"{temp_dir}/candidates.csv"
singleton_feature = f"{temp_dir}/singleton.csv"
feature_class_count = f"{temp_dir}/feature_class_count.csv"
feature_property_count = f"{temp_dir}/feature_property_count.csv"
feature_class_property_count = f"{temp_dir}/feature_property_class_count.csv"
score_file = f"{temp_dir}/scores.csv"
model_name = 'rf_tuned_ranking.pkl'
embedding_file = f'{temp_dir}/graph_embedding_complex.tsv'
aux_field = 'graph_embedding_complex,class_count,property_count,context'
final_score = f'{temp_dir}/final_score.csv'
top_k_file = f"{temp_dir}/topk-hormones.csv"
final_output = f"{output_path}/linked-hormones.csv"
# -
# ## Peek at the input file
pd.read_csv(input_file_path).fillna("")
# ## Canonicalize
# !tl canonicalize \
# -c "$wikify_column_name" \
# --add-context \
# {input_file_path} > {canonical}
df = pd.read_csv(canonical)
df
# ## Candidate Generation
# %%time
# !tl clean -c label -o label_clean {canonical} / \
# --url $es_url --index $es_index \
# get-fuzzy-augmented-matches -c label_clean \
# --auxiliary-fields {aux_field} \
# --auxiliary-folder $temp_dir / \
# --url $es_url --index $es_index \
# get-exact-matches -c label_clean \
# --auxiliary-fields {aux_field} \
# --auxiliary-folder {temp_dir} > {candidates}
column_rename_dict = {
'graph_embedding_complex': 'embedding',
'class_count': 'class_count',
'property_count': 'property_count',
'context': 'context'
}
for field in aux_field.split(','):
aux_list = []
for f in glob.glob(f'{temp_dir}/*{field}.tsv'):
aux_list.append(pd.read_csv(f, sep='\t', dtype=object))
aux_df = pd.concat(aux_list).drop_duplicates(subset=['qnode']).rename(columns={field: column_rename_dict[field]})
aux_df.to_csv(f'{temp_dir}/{field}.tsv', sep='\t', index=False)
pd.read_csv(f'{temp_dir}/context.tsv', sep='\t').head(20)
pd.read_csv(candidates, nrows = 150).fillna("")
# ### Add singleton feature
# !tl create-singleton-feature -o singleton {candidates} > {singleton_feature}
pd.read_csv(singleton_feature, dtype=object).head().fillna("")
# ### Add Class Count TF IDF Feature
# !tl compute-tf-idf \
# --feature-file /tmp/cricketers/temp/class_count.tsv \
# --feature-name class_count \
# --singleton-column singleton \
# -o class_count_tf_idf_score \
# {singleton_feature} > {feature_class_count}
# #### Peak at class count tf idf feature file
pd.read_csv(feature_class_count, dtype=object).head(20).fillna("")
# #### Get top 1 candidate for each cell
# !tl get-kg-links -c class_count_tf_idf_score -l label -k 1 --k-rows $feature_class_count > $temp_dir/class_count_top_k.csv
pd.read_csv(f"{temp_dir}/class_count_top_k.csv").fillna("")
# ### Add Property Count TF IDF Feature
# !tl compute-tf-idf \
# --feature-file /tmp/cricketers/temp/property_count.tsv \
# --feature-name property_count \
# --singleton-column singleton \
# -o property_count_tf_idf_score \
# {singleton_feature} > {feature_property_count}
# #### Peak at property count tf idf feature file
pd.read_csv(feature_property_count, dtype=object).head(20).fillna("")
# #### Get top 1 candidate for each cell
# !tl get-kg-links -c property_count_tf_idf_score -l label -k 1 --k-rows $feature_property_count > $temp_dir/property_count_top_k.csv
pd.read_csv(f"{temp_dir}/property_count_top_k.csv").fillna("")
# ## Use the combined property and class counts
# +
pdf = pd.read_csv(f"{temp_dir}/property_count.tsv", sep='\t')
cdf = pd.read_csv(f"{temp_dir}/class_count.tsv", sep='\t')
class_prop_file = f"{temp_dir}/class_property_count.tsv"
df = pdf.merge(cdf, on='qnode', how='left').fillna("")
df['class_prop_count_temp'] = list(zip(df.property_count, df.class_count))
df['class_property_count'] = df['class_prop_count_temp'].map(lambda x: "|".join(x) if x[1] != "" else x[0])
df.drop(columns=['class_prop_count_temp', 'class_count', 'property_count'], inplace=True)
df.to_csv(class_prop_file, sep='\t', index=False)
df
# -
# !tl compute-tf-idf \
# --feature-file /tmp/cricketers/temp/class_property_count.tsv \
# --feature-name class_property_count \
# --singleton-column singleton \
# -o class_property_count_tf_idf_score \
# {singleton_feature} > {feature_class_property_count}
# #### Peak at class property count tf idf feature file
pd.read_csv(feature_class_property_count, dtype=object).head(20).fillna("")
# #### Get top 1 candidate for each cell
# !tl get-kg-links -c class_property_count_tf_idf_score -l label -k 3 --k-rows $feature_class_property_count > $temp_dir/class_property_count_top_k.csv
pd.read_csv(f"{temp_dir}/class_property_count_top_k.csv").fillna("")
| table-linker-full-pipeline/table-linker-pipeline-tf-idf-feature.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Visualization with Python ecosystem
from arcgis.gis import *
gis = GIS()
county_search = gis.content.search("title:USA counties & owner:esri_dm", "feature layer")
county_search
county_item = county_search[0]
county_item.layers
county_item
county_feature_layer = county_item.layers[0]
for field in county_feature_layer.properties.fields:
print(field['name'] + " \t: " + field['type'])
# ### Plot California plopulation by county
county_fset = county_feature_layer.query("STATE_NAME='California'")
county_df = county_fset.df
county_df.head(5)
county_df.shape
# #### Use the 'Name' column as index
# makes plotting easier
county_df.NAME.head(10)
# +
import matplotlib.pyplot
# %matplotlib inline
county_df.set_index('NAME', inplace=True)
county_df.POP2012.plot(kind='bar', x='NAME', figsize=(15,5),
title='CA population by County')
# -
# ### More plotting - how are the columns correlated
import seaborn as sns
sns.jointplot(county_df['POP2012'], county_df['SQMI'])
sns.jointplot(county_df['WHITE'], county_df['BLACK'], kind='hex')
# Influence of population density on vacancy
county_df['pop_density'] = county_df['POP12_SQMI'] / county_df['SQMI']
county_df['pop_density'].head()
# Histogram of population density
sns.jointplot('pop_density', 'VACANT', data=county_df)
# ### Correlation amongst all variables
sns.pairplot(county_df[['VACANT','MALES','RENTER_OCC','HOUSEHOLDS', 'MED_AGE']])
| talks/DevSummit2018/Mapping Visualization and Analysis/visualization-with-python-ecosystem-matplotlib-seaborn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Predicting Wine Quality
# Data set - http://archive.ics.uci.edu/ml/datasets/Wine+Quality
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.metrics import r2_score
import numpy as np
import statsmodels.api as sm
import matplotlib.pyplot as plt
import seaborn as sns
import os
os.chdir("C:\\Users\\403-6\\BigData-project\\Wine_Quality")
wine_quality = pd.read_csv("winequality-red.csv", sep = ';')
wine_quality.rename(columns = lambda x: x.replace(" ", "_"), inplace=True)
# **Simple linear regression**
model = sm.OLS(wine_quality['quality'], sm.add_constant(wine_quality['alcohol'])).fit()
print(model.summary())
plt.scatter(wine_quality['alcohol'], wine_quality['quality'], label = 'Actual Data')
plt.plot(wine_quality['alcohol'], model.params[0] + model.params[1]*wine_quality['alcohol'],
c='r', label="Regression fit")
plt.title('Wine Quality regressed on Alcohol')
plt.xlabel('Alcohol')
plt.ylabel('Quality')
plt.show()
# **Simple Linear Regression - Model fit**
x_train,x_test,y_train,y_test = train_test_split(wine_quality['alcohol'],
wine_quality["quality"],train_size = 0.7,random_state=42)
x_train = pd.DataFrame(x_train)
x_test = pd.DataFrame(x_test)
y_train = pd.DataFrame(y_train)
y_test = pd.DataFrame(y_test)
def mean(values):
return round(sum(values)/float(len(values)),2)
alcohol_mean = mean(x_train['alcohol'])
quality_mean = mean(y_train['quality'])
alcohol_variance = round(sum((x_train['alcohol'] - alcohol_mean)**2),2)
quality_variance = round(sum((y_train['quality'] - quality_mean)**2),2)
covariance = round(sum((x_train['alcohol'] - alcohol_mean)*(y_train['quality'] - quality_mean)),2)
b1 = covariance/alcohol_variance
b0 = quality_mean = b1*alcohol_mean
print("\n\n Intercept(B0) : ", round(b0, 4), "\n\n Co-efficient(B1) : ", round(b1,4))
y_test["y_pred"] = pd.DataFrame(b0+b1*x_test['alcohol'])
R_sqrd = 1 - (sum((y_test['quality']-y_test['y_pred'])**2) / sum((y_test['quality'] -
mean(y_test['quality']))**2))
print("Test R-squred value : ", round(R_sqrd, 4))
# **Multi linear regression model**
eda_columns = ['volatile_acidity', 'chlorides', 'sulphates', 'alcohol','quality']
sns.set(style='whitegrid', context='notebook')
sns.pairplot(wine_quality[eda_columns], size = 2.5, x_vars = eda_columns, y_vars=eda_columns)
plt.show()
# **Correlation coefficients**
corr_mat = np.corrcoef(wine_quality[eda_columns].values.T)
sns.set(font_scale=1)
full_mat = sns.heatmap(corr_mat, cbar=True, annot=True, square=True, fmt='.2f',
annot_kws={'size':15}, yticklabels=eda_columns, xticklabels=eda_columns)
plt.show()
# **Backward method : Repeat once**
# +
columns = ['fixed_acidity', 'volatile_acidity', 'citric_acid', 'residual_sugar', 'chlorides',
'free_sulfur_dioxide', 'total_sulfur_dioxide', 'density', 'pH', 'sulphates', 'alcohol']
pdx = wine_quality[columns]
pdy = wine_quality["quality"]
# -
x_train, x_test, y_train, y_test = train_test_split(pdx,pdy,train_size = 0.7, random_state=42)
x_train_new = sm.add_constant(x_train)
x_test_new = sm.add_constant(x_test)
full_mod = sm.OLS(y_train, x_train_new)
full_res = full_mod.fit()
print(full_res.summary())
print("Variance Inflation Factor")
cnames = x_train.columns
for i in np.arange(0,len(cnames)) :
xvars = list(cnames)
yvar = xvars.pop(i)
mod = sm.OLS(x_train[yvar], sm.add_constant(x_train_new[xvars]))
res = mod.fit()
vif = 1/(1-res.rsquared)
print(yvar, round(vif,3))
# **Backward method : Repeat 2 times**
# +
# residual_sugar 제거
columns = ['fixed_acidity', 'volatile_acidity', 'citric_acid', 'chlorides',
'free_sulfur_dioxide', 'total_sulfur_dioxide', 'density', 'pH', 'sulphates', 'alcohol']
pdx = wine_quality[columns]
pdy = wine_quality["quality"]
# -
x_train, x_test, y_train, y_test = train_test_split(pdx,pdy,train_size = 0.7, random_state=42)
x_train_new = sm.add_constant(x_train)
x_test_new = sm.add_constant(x_test)
full_mod = sm.OLS(y_train, x_train_new)
full_res = full_mod.fit()
print(full_res.summary())
print("Variance Inflation Factor")
cnames = x_train.columns
for i in np.arange(0,len(cnames)) :
xvars = list(cnames)
yvar = xvars.pop(i)
mod = sm.OLS(x_train[yvar], sm.add_constant(x_train_new[xvars]))
res = mod.fit()
vif = 1/(1-res.rsquared)
print(yvar, round(vif,3))
# **Backward method : Repeat 3 times**
# +
# density 제거
columns = ['fixed_acidity', 'volatile_acidity', 'citric_acid', 'chlorides',
'free_sulfur_dioxide', 'total_sulfur_dioxide', 'pH', 'sulphates', 'alcohol']
pdx = wine_quality[columns]
pdy = wine_quality["quality"]
# -
x_train, x_test, y_train, y_test = train_test_split(pdx,pdy,train_size = 0.7, random_state=42)
x_train_new = sm.add_constant(x_train)
x_test_new = sm.add_constant(x_test)
full_mod = sm.OLS(y_train, x_train_new)
full_res = full_mod.fit()
print(full_res.summary())
print("Variance Inflation Factor")
cnames = x_train.columns
for i in np.arange(0,len(cnames)) :
xvars = list(cnames)
yvar = xvars.pop(i)
mod = sm.OLS(x_train[yvar], sm.add_constant(x_train_new[xvars]))
res = mod.fit()
vif = 1/(1-res.rsquared)
print(yvar, round(vif,3))
# **Backward method : Repeat 4 times**
# +
# fixed_acidity 제거
columns = ['volatile_acidity', 'citric_acid', 'chlorides',
'free_sulfur_dioxide', 'total_sulfur_dioxide', 'pH', 'sulphates', 'alcohol']
pdx = wine_quality[columns]
pdy = wine_quality["quality"]
# -
x_train, x_test, y_train, y_test = train_test_split(pdx,pdy,train_size = 0.7, random_state=42)
x_train_new = sm.add_constant(x_train)
x_test_new = sm.add_constant(x_test)
full_mod = sm.OLS(y_train, x_train_new)
full_res = full_mod.fit()
print(full_res.summary())
print("Variance Inflation Factor")
cnames = x_train.columns
for i in np.arange(0,len(cnames)) :
xvars = list(cnames)
yvar = xvars.pop(i)
mod = sm.OLS(x_train[yvar], sm.add_constant(x_train_new[xvars]))
res = mod.fit()
vif = 1/(1-res.rsquared)
print(yvar, round(vif,3))
# **Backward method : Repeat 5 times**
# +
# citric_acid 제거
columns = ['volatile_acidity', 'chlorides',
'free_sulfur_dioxide', 'total_sulfur_dioxide', 'pH', 'sulphates', 'alcohol']
pdx = wine_quality[columns]
pdy = wine_quality["quality"]
# -
x_train, x_test, y_train, y_test = train_test_split(pdx,pdy,train_size = 0.7, random_state=42)
x_train_new = sm.add_constant(x_train)
x_test_new = sm.add_constant(x_test)
full_mod = sm.OLS(y_train, x_train_new)
full_res = full_mod.fit()
print(full_res.summary())
print("Variance Inflation Factor")
cnames = x_train.columns
for i in np.arange(0,len(cnames)) :
xvars = list(cnames)
yvar = xvars.pop(i)
mod = sm.OLS(x_train[yvar], sm.add_constant(x_train_new[xvars]))
res = mod.fit()
vif = 1/(1-res.rsquared)
print(yvar, round(vif,3))
# **Predition of data**
# +
y_pred = full_res.predict(x_test_new)
y_pred_df = pd.DataFrame(y_pred)
y_pred_df.columns = ['y_pred']
pred_data = pd.DataFrame(y_pred_df['y_pred'])
y_test_new = pd.DataFrame(y_test)
y_test_new.reset_index(inplace=True)
pred_data['y_test'] = pd.DataFrame(y_test_new['quality'])
# -
# **R-square calculation**
rsqd = r2_score(y_test_new['quality'].tolist(), y_pred_df['y_pred'].tolist())
print("Test R-squared value: ", round(rsqd, 4))
# **Ridge Regression**
from sklearn.linear_model import Ridge
wine_quality = pd.read_csv("winequality-red.csv",sep=';')
wine_quality.rename(columns=lambda x: x.replace(" ", "_"), inplace=True)
all_columns = ['fixed_acidity', 'volatile_acidity', 'citric_acid', 'residual_sugar',
'chlorides', 'free_sulfur_dioxide', 'total_sulfur_dioxide', 'density',
'pH', 'sulphates', 'alcohol']
pdx = wine_quality[all_columns]
pdy = wine_quality["quality"]
x_train, x_test, y_train, y_test = train_test_split(pdx, pdy, train_size = 0.7, random_state=42)
alphas = [1e-4,1e-3,1e-2,0.1,0.5,1.0,5.0,10.0]
initrsq = 0
print("Ridge Regression : Best Parameters\n")
for alph in alphas :
ridge_reg = Ridge(alpha=alph)
ridge_reg.fit(x_train, y_train)
tr_rsqrd = ridge_reg.score(x_train, y_train)
ts_rsqrd = ridge_reg.score(x_test, y_test)
if ts_rsqrd > initrsq :
print("Lambda : ", alph, "\nTrain R-square Value : ", round(tr_rsqrd, 5),
"\nTest R-squared value : ", round(ts_rsqrd, 5))
initrsq = ts_rsqrd
# **Coeffients of Ridge regression of best alpha value**
ridge_reg = Ridge(alpha=0.001)
ridge_reg.fit(x_train, y_train)
print("Ridge Regression coefficient values of Alpha = 0.001\n")
for i in range(11):
print(all_columns[i], ": ", ridge_reg.coef_[i])
# **Lasso Regression**
from sklearn.linear_model import Lasso
alphas = [1e-4,1e-3,1e-2,0.1,0.5,1.0,5.0,10.0]
initrsq = 0
print("Lasso Regression : Best Parameters\n")
for alph in alphas :
lasso_reg = Lasso(alpha=alph)
lasso_reg.fit(x_train, y_train)
tr_rsqrd = lasso_reg.score(x_train, y_train)
ts_rsqrd = lasso_reg.score(x_test, y_test)
if ts_rsqrd > initrsq :
print("Lambda : ", alph, "\nTrain R-Squared value : ", round(tr_rsqrd,5),
"\nTest R-squared value : ", round(ts_rsqrd,5))
initrsq = ts_rsqrd
# **Coeffients of Lasso regression of best alpha value**
lasso_reg = Lasso(alpha = 0.001)
lasso_reg.fit(x_train, y_train)
print("Lasso Regression coefficient values of Alpha = 0.001\n")
for i in range(11) :
print(all_columns[i], ": ", lasso_reg.coef_[i])
| Wine_Quality/Wine_Quality.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # CS 20 : TensorFlow for Deep Learning Research
# ## Lecture 03 : Linear and Logistic Regression
# ### Linear Regression with tf.data
#
# **Reference**
#
# * https://jhui.github.io/2017/11/21/TensorFlow-Importing-data/
# * https://towardsdatascience.com/how-to-use-dataset-in-tensorflow-c758ef9e4428
# * https://stackoverflow.com/questions/47356764/how-to-use-tensorflow-dataset-api-with-training-and-validation-sets
# ### Setup
# +
import os, sys
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import tensorflow as tf
from pprint import pprint
# %matplotlib inline
print(tf.__version__)
# -
# ### Build input pipeline
train_dir = os.listdir('../data/lecture03/example_with_data/train_dir/')
train_dir = list(map(lambda path : '../data/lecture03/example_with_data/train_dir/' + path, train_dir))
pprint(train_dir, compact = True)
val_dir = '../data/lecture03/example_with_data/val_dir/birth_life_2010_val.txt'
pprint(val_dir)
# hyper parameters
epochs = 100
batch_size = 8
# +
# datasets construction
# for training dataset
tr_dataset = tf.data.TextLineDataset(filenames = train_dir)
tr_dataset = tr_dataset.map(lambda record : tf.decode_csv(records = record,
record_defaults = [[''],[.0],[.0]],
field_delim = '\t')[1:])
tr_dataset = tr_dataset.shuffle(200)
tr_dataset = tr_dataset.batch(batch_size = batch_size)
tr_iterator = tr_dataset.make_initializable_iterator()
# for validation dataset
val_dataset = tf.data.TextLineDataset(filenames = val_dir)
val_dataset = val_dataset.map(lambda record : tf.decode_csv(records = record,
record_defaults = [[''],[.0],[.0]],
field_delim = '\t')[1:])
val_dataset = val_dataset.batch(batch_size = batch_size)
val_iterator = val_dataset.make_initializable_iterator()
# -
# handle constructions. Handle allows us to feed data from different dataset by providing a parameter in feed_dict
handle = tf.placeholder(dtype = tf.string)
iterator = tf.data.Iterator.from_string_handle(string_handle = handle,
output_types = tr_iterator.output_types)
X, Y = iterator.get_next()
sess_config = tf.ConfigProto(gpu_options=tf.GPUOptions(allow_growth=True))
sess = tf.Session(config = sess_config)
sess.run(tf.global_variables_initializer())
# ### Define the graph of Simple Linear Regression
# +
# create weight and bias, initialized to 0
w = tf.get_variable(name = 'weight', initializer = tf.constant(.0))
b = tf.get_variable(name = 'bias', initializer = tf.constant(.0))
# construct model to predict Y
yhat = X * w + b
# use the square error as loss function
mse_loss = tf.reduce_mean(tf.square(Y - yhat))
mse_loss_summ = tf.summary.scalar(name = 'mse_loss', tensor = mse_loss) # for tensorboard
# using gradient descent with learning rate of 0.01 to minimize loss
opt = tf.train.GradientDescentOptimizer(learning_rate=.01)
training_op = opt.minimize(mse_loss)
# -
# ### Training
train_writer = tf.summary.FileWriter(logdir = '../graphs/lecture03/linreg_mse_with_tf_data/train',
graph = tf.get_default_graph())
val_writer = tf.summary.FileWriter(logdir = '../graphs/lecture03/linreg_mse_with_tf_data/val',
graph = tf.get_default_graph())
# +
'''
# hyper parameters
epochs = 100
batch_size = 8
'''
tr_loss_hist = []
val_loss_hist = []
sess = tf.Session()
sess.run(tf.global_variables_initializer())
tr_handle, val_handle = sess.run(fetches = [tr_iterator.string_handle(), val_iterator.string_handle()])
for epoch in range(epochs):
avg_tr_loss = 0
avg_val_loss = 0
tr_step = 0
val_step = 0
# for mini-batch training
sess.run(tr_iterator.initializer)
try:
while True:
_, tr_loss, tr_loss_summ = sess.run(fetches = [training_op, mse_loss, mse_loss_summ], feed_dict = {handle : tr_handle})
avg_tr_loss += tr_loss
tr_step += 1
except tf.errors.OutOfRangeError:
pass
# for validation
sess.run(val_iterator.initializer)
try:
while True:
val_loss, val_loss_summ = sess.run(fetches = [mse_loss, mse_loss_summ], feed_dict = {handle : val_handle})
avg_val_loss += val_loss
val_step += 1
except tf.errors.OutOfRangeError:
pass
train_writer.add_summary(tr_loss_summ, global_step = epoch)
val_writer.add_summary(val_loss_summ, global_step = epoch)
avg_tr_loss /= tr_step
avg_val_loss /= val_step
tr_loss_hist.append(avg_tr_loss)
val_loss_hist.append(avg_val_loss)
if epoch % 10 == 0:
print('epoch : {:3}, tr_loss : {:.3f}, val_loss : {:.3f}'.format(epoch, avg_tr_loss, avg_val_loss))
train_writer.close()
val_writer.close()
# -
# ### Visualization
plt.plot(tr_loss_hist, label = 'train')
plt.plot(val_loss_hist, label = 'validation')
plt.legend()
data = pd.read_table('../data/lecture03/example_with_placeholder/birth_life_2010.txt') # loading data for Visualization
w_out, b_out = sess.run([w, b])
plt.plot(data.iloc[:,1], data.iloc[:,2], 'bo', label='Real data')
plt.plot(data.iloc[:,1], data.iloc[:,1] * w_out + b_out, 'r', label='Predicted data')
plt.legend()
| Lec03_Linear and Logistic Regression/Lec03_Linear Regression with tf.data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] tags=[]
# # Upload data to Cloud Storage
# -
# %load_ext lab_black
# %load_ext autoreload
# %autoreload 2
# +
import os
import re
import time
from glob import glob
from zipfile import ZipFile
from azure.storage.blob import BlobServiceClient
from prefect import Flow, context as prefect_context, task, unmapped
# -
# <a href="table-of-contents"></a>
#
# ## [Table of Contents](#table-of-contents)
# 0. [About](#about)
# 1. [User Inputs](#user-inputs)
# 2. [Create archives from scraped files and Upload to Blob Storage](#create-archives-from-scraped-files-and-upload-to-blob-storage)
# - 2.1. [Listings](#listings)
# - 2.2. [Search Results](#search-results)
# <a id="about"></a>
#
# ## 0. [About](#about)
# All scraped single-row CSV files of listings and upto 25-row CSV files of search results are combined into batches, of a desired size and uploaded to [Azure blob storage](https://azure.microsoft.com/en-us/services/storage/blobs/).
#
# A batch of size 12 for the search results will mean that every 12 pages (each page has upto 25 rows of search results) of scraped CSV files will be archived together. A batch of size 1 for the listings will mean that every 1 row of scraped CSV listings will be archived together.
#
# **Requirements**
#
# The following four environment variables should be exported before running this notebook
# - `BLOB_NAME_PREFIX`
# - for each combined archive to be uploaded (listings and search results), a unique number will be appended to this value
# - `AZURE_STORAGE_ACCOUNT`
# - `AZURE_STORAGE_KEY`
# - `ENDPOINT_SUFFIX`
#
# **Notes**
# 1. This should be done for the raw scraped data only. In this way, subsequent runs of the exploratory data analysis phase of this project can use the following workflow that does not require web-scraping to access any previously downloaded data
# - download this previously scraped data from cloud storage (`9_*.ipynb`)
# - cleaning/merging using `6_merge_searches_listings.ipynb`
# - exploring the data with `7_eda.ipynb`
#
# In this way, the notebooks responsible for web-scraping (`0_*.ipynb` to `5_*.ipynb`) will not be run.
# 2. The workflow used here is executed using a workflow management tool. This was optional, but has been used here.
# <a id="user-inputs"></a>
#
# ## 1. [User Inputs](#user-inputs)
PROJ_ROOT_DIR = os.getcwd()
# + tags=["parameters"]
# Last page number for listings or search results to be archived and uploaded
max_page_num_to_check = 50 + 1
min_page_num_to_check_requests = 50
max_page_num_to_check_requests = 525
max_page_num_to_check_sr_requests = 2235 + 1
# Size of batch of CSV files of scraped listings
page_batch_size = 5
page_batch_size_requests = 25
# Size of batch of CSV files of scraped search results
page_batch_size_sr = 10
page_batch_size_sr_requests = 25
# Suffixes (of filenames) to upload combined archives of listings and
# search results to Azure blob storage
blob_name_suffixes = {
"listings": 80,
"search_results": 81,
"listings_requests": 82,
"search_results_requests": 83,
}
# +
data_dir = os.path.join(PROJ_ROOT_DIR, "data")
raw_data_dir = os.path.join(data_dir, "raw")
requests_files_dir = os.path.join(raw_data_dir, "requests")
selenium_files_dir = os.path.join(raw_data_dir, "selenium")
conn_str = (
"DefaultEndpointsProtocol=https;"
f"AccountName={os.getenv('AZURE_STORAGE_ACCOUNT')};"
f"AccountKey={os.getenv('AZURE_STORAGE_KEY')};"
f"EndpointSuffix={os.getenv('ENDPOINT_SUFFIX')}"
)
blob_name_prefix = os.getenv("BLOB_NAME_PREFIX")
# Selenium - LISTINGS
# - listings on search results pages 2-49 were scraped
# - this list should cover pages 1 to 50, inclusive
# - with a step size of 5, every 5 pages (25 listings per page) will be placed in a separate batch (5, 10, 15, 20, etc.)
d = [
k + page_batch_size - 1
for k in range(1, (max_page_num_to_check - 1) + 1, page_batch_size)
]
print(d)
# Selenium - SEARCH RESULTS
# - pages 2-49 of search results were scraped
# - this list should cover pages 1 to 50, inclusive
# - with a step size of 10, every 10 pages (25 listings per page) will be placed in a separate batch
# - the first batch will cover pages 1 to 10, the second batch covers pages 11 - 20, etc.
d_sr = [
k + page_batch_size_sr - 1
for k in range(
1,
(max_page_num_to_check - 1) + 1,
page_batch_size_sr,
)
]
print(d_sr)
# Requests - LISTINGS
# - listings on search results pages 50-505 were scraped
# - this list should cover pages 50 to 505, inclusive
# - with a step size of 25, every 25 pages (25 listings per page) will be placed in a separate batch (74, 99, 124)
d_requests = [
k + page_batch_size_requests - 1
for k in range(
min_page_num_to_check_requests,
(max_page_num_to_check_requests - 1) + 1,
page_batch_size_requests,
)
]
print(d_requests)
# Requests - SEARCH RESULTS
# - pages 50-2232 of search results were scraped
# - this list should cover pages 50 to 2250, inclusive
# - with a step size of 25, every 25 pages (25 listings per page) will be placed in a separate batch
# - the first batch will cover pages 50 to (50 + 25) - 1 = 74, the second batch covers pages 75 - 99, etc.
d_sr_requests = [
k + page_batch_size_sr_requests - 1
for k in range(
min_page_num_to_check_requests,
(max_page_num_to_check_sr_requests - 1) + 1,
page_batch_size_sr_requests,
)
]
print(d_sr_requests)
# +
@task
def get_files_single_page(
page_num_max,
search_results_file_list,
prefix="p",
file_type="listings",
step_size=100,
):
"""Create archive."""
logger = prefect_context.get("logger")
matching_filepaths = []
fname = (
f"batched_{file_type}__{page_num_max-(step_size-1)}_"
f"{page_num_max}__{time.strftime('%Y%m%d_%H%M%S')}.zip"
)
if not os.path.exists(fname):
for f in search_results_file_list:
p_str = re.search(fr"{prefix}(\d+)_", f).group(0)
p_num_str = (
p_str.replace("_", "").replace("p", "")
if prefix == "p"
else p_str.split("_")[-2]
)
if step_size == 1:
if int(p_num_str) == page_num_max:
matching_filepaths.append(f)
else:
if (page_num_max - step_size) + 1 <= int(p_num_str) <= page_num_max:
# print(f)
matching_filepaths.append(f)
logger.info(
f"Page start = {page_num_max - (step_size - 1)}, "
f"Page stop = {page_num_max}, "
f"Files Found = {len(matching_filepaths)}"
)
return {fname: matching_filepaths}
@task
def create_archive(files_dict):
logger = prefect_context.get("logger")
archives_created = []
for archive_fname, matching_filepaths in files_dict.items():
if len(matching_filepaths) > 0:
with ZipFile(archive_fname, "w") as zip_file:
for matching in matching_filepaths:
zip_file.write(matching)
archives_created.append(archive_fname)
logger.info(f"Creating archive {archive_fname}")
else:
pass
logger.info(
f"Did not find any filepaths for {archive_fname}. "
"Will not create archive."
)
return archives_created
@task
def upload_az_file_blobs(
blob_names_dict, conn_str, flatten_filepaths=True, az_container_name="myconedesx7"
):
logger = prefect_context.get("logger")
if flatten_filepaths:
blob_names_dict = {k: l for k, v in blob_names_dict.items() for l in v}
# print(type(blob_names_dict), blob_names_dict)
blob_service_client = BlobServiceClient.from_connection_string(conn_str=conn_str)
for az_blob_name, local_file_path in blob_names_dict.items():
blob_client = blob_service_client.get_blob_client(
container=az_container_name, blob=az_blob_name
)
# print(az_blob_name, local_file_path)
if not list(
blob_service_client.get_container_client(az_container_name).list_blobs(
name_starts_with=az_blob_name
)
):
with open(local_file_path, "rb") as data:
blob_client.upload_blob(data)
logger.info(f"Blob {az_blob_name} not found. Uploaded {local_file_path}.")
else:
logger.info(f"Blob {az_blob_name} found. Did not upload {local_file_path}.")
# -
# <a id="create-archives-from-scraped-files-and-upload-to-blob-storage"></a>
#
# ## 2. [Create archives from scraped files and Upload to Blob Storage](#create-archives-from-scraped-files-and-upload-to-blob-storage)
# <a id="selenium-listings"></a>
#
# ### 2.1. [Selenium Listings](#selenium-listings)
# Change into the sub-directory containing the downloaded single-row CSV files with the scraped contents of each listing scraped with `requests`
os.chdir(selenium_files_dir)
print(f"Current working directory is {os.getcwd()}")
# Create an archive of batches of listings files, where the size of each batch is the difference between successive elements in the list `d` defined earlier
# + tags=[]
with Flow("Archive Listings Scraped with Selenium") as flow:
files = get_files_single_page.map(
d,
unmapped(glob("*.csv")),
unmapped("p"),
unmapped("listings"),
unmapped(d[1] - d[0]),
)
_ = create_archive.map(files)
state = flow.run()
# -
# Create a single archive with the combination of all batched listings archives created above and upload to Azure blob storage
# + tags=[]
with Flow("Combine and Upload Selenium-Based Listings Archives") as flow:
combined_archive_fname = (
f"combo_batched_listings__{time.strftime('%Y%m%d_%H%M%S')}.zip"
)
combo_archive = create_archive(
{combined_archive_fname: glob("batched_listings__*.zip")}
)
upload_az_file_blobs(
{f"{blob_name_prefix}{blob_name_suffixes['listings']}": combo_archive},
conn_str,
True,
)
state_combo = flow.run()
# state_combo.result[combo_archive].result
# -
# **Note**
# 1. It seems like this has to be called after the above `Flow`, since Prefect can't handle re-using the same task (here, this would be `create_archive()`) with different inputs ([link](https://github.com/PrefectHQ/prefect/issues/4603)) while also having the second run of the task dependent on the output of the first run.
# 2. An attempt at capturing the output of the first run of the task and feeding it into the second run (thereby forcing the second run to be dependent on the first, by replacing `glob("batched_listings__*.zip")`) did not work. Future work should further investigate this as a workaround for the above.
# <a id="requests-listings"></a>
#
# ### 2.2. [Requests Listings](#requests-listings)
# Change into the sub-directory containing the downloaded single-row CSV files with the scraped contents of each listing scraped with Selenium
os.chdir(requests_files_dir)
print(f"Current working directory is {os.getcwd()}")
# Create an archive of batches of listings files, where the size of each batch is the difference between successive elements in the list `d_requests` defined earlier
# + tags=[]
with Flow("Archive Listings Scraped with Requests") as flow:
files = get_files_single_page.map(
d_requests,
unmapped(glob("p*.csv")),
unmapped("p"),
unmapped("listings"),
unmapped(d_requests[1] - d_requests[0]),
)
_ = create_archive.map(files)
state = flow.run()
# -
# Create a single archive with the combination of all batched listings archives created above and upload to Azure blob storage
# + tags=[]
with Flow("Combine and Upload Requests-Based Listings Archives") as flow:
combined_archive_fname = (
f"combo_batched_listings__{time.strftime('%Y%m%d_%H%M%S')}.zip"
)
combo_archive = create_archive(
{combined_archive_fname: glob("batched_listings__*.zip")}
)
upload_az_file_blobs(
{f"{blob_name_prefix}{blob_name_suffixes['listings_requests']}": combo_archive},
conn_str,
True,
)
state_combo = flow.run()
# state_combo.result[combo_archive].result
# -
# <a id="selenium-search-results"></a>
#
# ### 2.3. [Selenium Search Results](#selenium-search-results)
# Change into the sub-directory containing the downloaded single-row CSV files with the scraped contents of each listing scraped with Selenium
os.chdir(selenium_files_dir)
print(f"Current working directory is {os.getcwd()}")
# Create an archive of batches of search results files, where the size of each batch is the difference between successive elements in the list `d_sr` defined earlier
# + tags=[]
# %%time
with Flow("Archive Search Results Pages Scraped with Selenium") as flow:
files = get_files_single_page.map(
d_sr,
unmapped(glob("*.parquet.gzip")),
unmapped("search_results_page_"),
unmapped("search_results"),
unmapped(d_sr[1] - d_sr[0]),
)
_ = create_archive.map(files)
state_sr = flow.run()
# -
# Create a single archive with the combination of all batched search results archives created above and upload to Azure blob storage
# +
# %%time
with Flow("Combine and Upload Selenium-Based Search Results Archives") as flow:
combined_archive_fname = (
f"combo_batched_search_results__{time.strftime('%Y%m%d_%H%M%S')}.zip"
)
combo_archive = create_archive({combined_archive_fname: glob("batched_search_results__*.zip")})
upload_az_file_blobs(
{f"{blob_name_prefix}{blob_name_suffixes['search_results']}": combo_archive}, conn_str, True
)
state_sr_combo = flow.run()
# state_sr_combo.result[combo_archive].result
# -
# <a id="requests-search-results"></a>
#
# ### 2.4. [Requests Search Results](#requests-search-results)
# Change into the sub-directory containing the downloaded single-row CSV files with the scraped contents of each listing scraped with `requests`
os.chdir(requests_files_dir)
print(f"Current working directory is {os.getcwd()}")
# Create an archive of batches of search results files, where the size of each batch is the difference between successive elements in the list `d_sr` defined earlier
# + tags=[]
# %%time
with Flow("Archive Search Results Pages Scraped with Requests") as flow:
files = get_files_single_page.map(
d_sr_requests,
unmapped(glob("search_results_page_*.parquet.gzip")),
unmapped("search_results_page_"),
unmapped("search_results"),
unmapped(d_sr_requests[1] - d_sr_requests[0]),
)
_ = create_archive.map(files)
state_sr = flow.run()
# -
# Create a single archive with the combination of all batched search results archives created above and upload to Azure blob storage
# +
# %%time
with Flow("Combine and Upload Requests-Based Search Results Archives") as flow:
combined_archive_fname = (
f"combo_batched_search_results__{time.strftime('%Y%m%d_%H%M%S')}.zip"
)
combo_archive = create_archive({combined_archive_fname: glob("batched_search_results__*.zip")})
upload_az_file_blobs(
{f"{blob_name_prefix}{blob_name_suffixes['search_results_requests']}": combo_archive}, conn_str, True
)
state_sr_combo = flow.run()
# state_sr_combo.result[combo_archive].result
# -
# ---
# <span style="float:left">
# <a href="./7_eda.ipynb"><< 7 - Exploratory Data Analysis</a>
# </span>
#
# <span style="float:right">
# <a href="./9_download_cloud.ipynb">9 - Download all scraped data (with requests and selenium) from cloud storage >></a>
# </span>
| 8_upload_cloud.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import tweepy
import datetime
import csv
import codecs
def gettwitterdata(keyword,dfile,sincedate,untildate):
#python で Twitter APIを使用するためのConsumerキー、アクセストークン設定
Consumer_key = '5xFc5PTRff3CrjvPVXJhJJjlc'
Consumer_secret = '<KEY>'
Access_token = '<KEY>'
Access_secret = '<KEY>'
#認証
auth = tweepy.OAuthHandler(Consumer_key, Consumer_secret)
auth.set_access_token(Access_token, Access_secret)
api = tweepy.API(auth, wait_on_rate_limit = True)
#検索キーワード設定
q = keyword
#つぶやきを格納するリスト
tweets_data =[]
#カーソルを使用してデータ取得
for tweet in tweepy.Cursor(api.search,
q=q,
count=100,
tweet_mode='extended',
lang='ja',
since=sincedate+'_00:00:00_JST',
until=untildate+'_23:59:59_JST'
).items():
#つぶやきテキスト(FULL)を取得
tweets_data.append([tweet.user.screen_name,
tweet.created_at + datetime.timedelta(hours=9),
"'" + tweet.full_text.replace("\n"," "),
tweet.favorite_count,
tweet.retweet_count,
#tweet.place.name,
#tweet.quote_count,
#tweet.reply_count,
#tweet.hashtags.text,
tweet.lang,
tweet.user.location,
"'" + tweet.user.description.replace("\n"," "),
tweet.user.followers_count,
tweet.user.verified,
tweet.user.friends_count,
tweet.user.statuses_count,
tweet.user.created_at])
#print(sincedate+'_00:00:00_JST')
#print(untildate+'_00:00:00_JST')
#出力ファイル名
fname= r"'"+ dfile + "'"
fname = fname.replace("'","")
#ファイル出力
with open(fname, "w",newline='',encoding="utf-8") as f:
writer = csv.writer(f, lineterminator='\n')
writer.writerow(["アカウント",
"作成日",
"ツイート",
"良いね数",
"リツイート数",
#"場所",
#"引用数",
#"返信数",
#"ハッシュタグ",
"言語",
"場所",
"アカウントの説明",
"フォロワー数",
"認証の有無",
"フォローしている数",
"ツイート数",
"アカウント作成日"])
writer.writerows(tweets_data)
f.close
if __name__ == '__main__':
##検索キーワードを入力 ※リツイートを除外する場合 「キーワード -RT 」と入力
#print ('====== Enter Serch KeyWord =====')
#keyword = input('> ')
keyword = "paypay -rt"
##"検索する期間(自)
#"print ('====== 検索する期間(自)「yyyy-mm-dd」 =====')
sincedate = (datetime.date.today() + datetime.timedelta(days=-1)).strftime('%Y-%m-%d')
##検索する期間(至)
#print ('====== 検索する期間(至)「yyyy-mm-dd」 =====')
untildate = (datetime.date.today() + datetime.timedelta(days=-1)).strftime('%Y-%m-%d')
##出力ファイル名を入力(相対パス or 絶対パス)
#print ('====== Enter Tweet Data file =====')
dfile = r"C:\twitter\text_analysis\tweet_data\paypay_"+ sincedate +".csv"
gettwitterdata(keyword,dfile,sincedate,untildate)
# -
| twitter_search-yestadey.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/srimanthtenneti/Tensorflow101/blob/main/ANNSolution.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="f7j-U679WixS"
import tensorflow as tf
import tensorflow.keras as keras
import tensorflow.keras.layers as layers
from tensorflow.keras.models import Sequential
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# + id="jxGHclN2W7ke"
from sklearn.datasets import load_iris
dataset = load_iris()
x_train = dataset.data
y_train = dataset.target.reshape(-1,1)
# + id="x3nBj4vEceuU"
from sklearn.preprocessing import OneHotEncoder
encoder = OneHotEncoder(sparse=False)
y_train = encoder.fit_transform(y_train)
# + id="3r6uePPJcqDa"
model = Sequential([
layers.Dense(10 , input_shape= (4,) , activation='relu'),
layers.Dense(10 , activation='relu'),
layers.Dense(3 , activation='softmax')
])
# + colab={"base_uri": "https://localhost:8080/"} id="aT4Dql4Ac6_v" outputId="07d250ef-3011-4e29-ae3f-a8ab964b9367"
model.summary()
# + id="h3L6mw7tc8i7"
optimizer = tf.keras.optimizers.Adam(lr = 0.001)
# + id="_rbBii4bdCiC"
model.compile(optimizer , loss = 'categorical_crossentropy' , metrics = ['accuracy'])
# + colab={"base_uri": "https://localhost:8080/"} id="s3wOOTjMdMFL" outputId="e70ca535-fed9-42fd-a451-93fc609aabb2"
model.fit(x_train , y_train , epochs = 100)
# + colab={"base_uri": "https://localhost:8080/"} id="U5q5xyRkdRW-" outputId="4d502b53-3085-42d9-95ed-a5975a7c40c3"
x_train[0].shape
# + id="TKPmqy9WdtxR"
pred = np.argmax(model.predict(x_train[0].reshape(-1, 4)))
# + colab={"base_uri": "https://localhost:8080/"} id="Zi41CoMxeA__" outputId="6d74f8a0-64d6-4746-9ca6-0ec39c32f15f"
print('The true label is: {} , predicted label is: {}'.format(dataset.target[0] , pred))
| ANNSolution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Start with importing the packages
# %matplotlib inline
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
import matplotlib as mpl
from mpl_toolkits.mplot3d import Axes3D
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
from sklearn.decomposition import PCA
from sklearn.manifold import TSNE
from sklearn.preprocessing import StandardScaler
import scipy
from scipy import stats
import seaborn as sns
from sklearn.cluster import KMeans
from sklearn.metrics import confusion_matrix
import statsmodels.stats.multitest as smm
from statsmodels.formula.api import ols
np.set_printoptions(suppress=True)
np.set_printoptions(precision=4)
plt_style = 'seaborn-notebook'
pd.set_option('precision', 2)
pd.set_option('max_columns',10)
# -
# ## import data table
df = pd.read_excel('ddpcr-data.xlsx')
df = df.set_index('Isolate')
print(df.shape)
df.head()
# Table notes:
# DVG_2A column: based on NGS data
# DVG_2B column: based on ddpcr ratio, cut of at 1.2
# ## Stats for BK box plot
## Stats for ddpcr-ratio vs DVGs
DIPYes = df.loc[df.DVG_2A == 'Yes','CopiesmL':]
DIPNo = df.loc[df.DVG_2A == 'No', 'CopiesmL':]
# test the significance of the difference
#stat, p = stats.ttest_ind(DIPYes.Final, DIPNo.Final, nan_policy='omit')
stat, p = stats.kruskal(DIPYes.Final, DIPNo.Final, nan_policy='omit')
print('p =', p)
DIPYes.describe()
DIPNo.describe()
## Stats for viral-load vs DVGs
DIPYes2 = df.loc[df.DVG_2B == 'Yes','CopiesmL':]
DIPNo2 = df.loc[df.DVG_2B == 'No', 'CopiesmL':]
# test the significance of the difference0
#stat2, p2 = stats.ttest_ind(DIPYes2.log, DIPNo2.log, nan_policy='omit')
stat2, p2 = stats.kruskal(DIPYes2.log, DIPNo2.log, nan_policy='omit')
print('p value for= ', p2)
DIPYes2.describe()
DIPNo2.describe()
# ## Fig 2A: BK boxplot with VP/LT ratio
# +
palette = ('#000000','#FF0000')
sns.set_context("talk")
sns.set_style(style="white")
g = sns.catplot(x="DVG_2A", y="Final", data=df, hue='Notes',palette=palette)
g._legend.remove()
g = sns.boxplot(x="DVG_2A", y="Final", data=df, color = 'white',showfliers= False,linewidth=2.5)
plt.yticks(np.arange(0, 4.5, step=0.5))
g.set(ylim=(0.5,3.9))
plt.xlabel("DVGs Presence by Sequencing",labelpad=20)
plt.ylabel("VP:LargeT Ratio",labelpad=10)
x1, x2 = 0, 1 # columns 'Sat' and 'Sun' (first column: 0, see plt.xticks())
y, h, col = 3.7, 0.10, 'k'
plt.plot([x1, x1, x2, x2], [y, y+h, y+h, y], lw=1.5, c=col)
label = "Kruskal-Wallis test, p = 0.0002"
plt.text((x1+x2)*.5, y+h+h, label, ha='center', va='bottom', color=col, fontsize=14)
#plt.plot([-0.5,1.5],[1.2,1.2],linestyle='dashed', color = 'red', linewidth = 1)
#plt.legend(fontsize="x-small", bbox_to_anchor=(1,1),loc='upper left')
#plt.tight_layout()
plt.savefig('ddpcr-fig2A_2.png', dpi=300,facecolor='white')
plt.show()
# -
plt.figure(figsize=(15,5))
plt.savefig('ddpcr-fig2B_blank.png', dpi=300, facecolor='white')
# ## Fig 2B: BK boxplot with viral load
# +
sns.set_context("talk")
sns.set_style(style="white")
b = sns.catplot(x="DVG_2B", y="log", data=df, color='black')
#g._legend.remove()
b.set(ylim=(4,13))
b = sns.boxplot(x="DVG_2B", y="log", data=df, color = 'white',showfliers= False,linewidth=2.5)
plt.xlabel("DVGs Presence by ddPCR",labelpad=20)
plt.ylabel("log10 copies/mL",labelpad=10)
plt.yticks(np.arange(4, 13.5, step=1))
x1, x2 = 0, 1 # (first column: 0, see plt.xticks())
y, h, col = 12.5, 0.10, 'k'
plt.plot([x1, x1, x2, x2], [y, y+h, y+h, y], lw=1.5, c=col)
label = "Kruskal-Wallis test, p = 0.0087"
plt.text((x1+x2)*.5, y+h+h, label, ha='center', va='bottom', color=col,fontsize=14)
#plt.legend(fontsize="x-small", bbox_to_anchor=(1,1),loc='upper left')
plt.tight_layout()
plt.savefig('ddpcr-fig2B_2.png', dpi=300, facecolor='white')
plt.show()
# -
# ## Fig 2C: JC boxplot with VP/LT ratio
df2 = pd.read_excel('jcddpcr-data.xlsx')
print(df2.shape)
df2
DVGNo = df2.loc[df2.DVG == 'No/Unknown', 'Ratio':]
DVGNo.describe()
sns.set_context("talk")
sns.set_style(style="white")
c = sns.catplot(x="DVG", y="Ratio", data=df2, color='black')
#g._legend.remove()
c = sns.boxplot(x="DVG", y="Ratio", data=df2, color = 'white',showfliers= False,linewidth=2.5)
plt.xlabel("DVGs Presence by Sequencing",labelpad=20)
plt.ylabel("VP:LargeT Ratio",labelpad=10)
plt.yticks(np.arange(0.5, 1.4, step=0.1))
#plt.legend(fontsize="x-small", bbox_to_anchor=(1,1),loc='upper left')
plt.tight_layout()
plt.savefig('ddpcr-fig2C_2.png', dpi=300)
plt.show()
# ## old codes
# +
palette = ('#F04DCA','#52C76D','#F78707','#3A62CF')
#palette=dict(Yes="g", No="m")
#fig33 = plt.figure()
#BK = df.loc[df.Virus2 == 'BK+','Viral_Load':]
fig33 = sns.lmplot(x="log", y="Final", data=df, scatter=False, line_kws={'color': '#848EE8',"linewidth": 1})
fig33 = sns.scatterplot(x = df['log'], y = df['Final'], data=df, hue="DVG")
fig33.set_ylabel('VP:LargeT Ratio')
fig33.set_xlabel('Log Viral Load')
#fig33.set(ylim=(0.5,4))
#plt.legend(fontsize="small", bbox_to_anchor=(0.78,1),loc='upper left', title=None)
plt.plot([4,11],[1,1], linestyle="dashed", color = 'black', linewidth = 1)
plt.show()
#plt.savefig('ddpcr-scatter-withoutngs.png')
# -
fig2 = plt.figure(figsize=(11,8))
scat = fig2.add_subplot(111)
scat.scatter(x = df.index, y = df['VP'], color='b')
scat.scatter(x = df.index, y = df['LargeT'], color='k')
scat.set_ylabel('Copies/25uL well')
scat.set_xlabel('Sample ID')
scat.set_ylim(-1,5100)
fig2 = plt.xticks(df.index, df.index, rotation='vertical')
plt.show()
#BK = df.loc[df.Virus2 == 'BK+','Viral_Load':]
slope, intercept, r_value, p_value, std_err = stats.linregress(df['log'],df['Final'])
print("Stats for all data:", "Slope =",slope, "r-squared =" ,r_value**2, "p-value = ", p_value)
#fig44 = sns.lmplot(x="Log_load", y="Ratio", data=df, palette=palette, height=3, aspect=1, scatter=False)
#plt.show()
| Figure2-ddpcr.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:py36]
# language: python
# name: conda-env-py36-py
# ---
import kenlm
import json
import nltk
from tqdm import tqdm
from nltk import word_tokenize
# ### Set testfile (libri)
# +
list_test_file = []
list_test_file.append( './corpus/test/asr-baseline/dev-clean.json' )
list_test_file.append( './corpus/test/asr-baseline/dev-other.json' )
list_test_file.append( './corpus/test/asr-baseline/test-clean.json' )
list_test_file.append( './corpus/test/asr-baseline/test-other.json' )
list_test_file.append( './corpus/test/asr-baseline-minus/dev-clean.json' )
list_test_file.append( './corpus/test/asr-baseline-minus/dev-other.json' )
list_test_file.append( './corpus/test/asr-baseline-minus/test-clean.json' )
list_test_file.append( './corpus/test/asr-baseline-minus/test-other.json' )
list_test_file.append( './corpus/test/asr-baseline-plus/dev-clean.json' )
list_test_file.append( './corpus/test/asr-baseline-plus/dev-other.json' )
list_test_file.append( './corpus/test/asr-baseline-plus/test-clean.json' )
list_test_file.append( './corpus/test/asr-baseline-plus/test-other.json' )
# -
# ### Load n-gram model (libri)
model_2 = kenlm.Model('./model/2-libri.binary')
model_3 = kenlm.Model('./model/3-libri.binary')
model_4 = kenlm.Model('./model/4-libri.binary')
model_5 = kenlm.Model('./model/5-libri.binary')
# ### Compute LM score & re-store
def compute_lm(json_data):
for key in tqdm(json_data.keys()):
for key_hyp in json_data[key].keys():
if key_hyp == 'ref':
continue
text = json_data[key][key_hyp]['text']
text = word_tokenize(text.lower().strip())
text = ' '.join(text)
# compute score & store back
json_data[key][key_hyp]['score_lm_2'] = model_2.score(text)
json_data[key][key_hyp]['ppl_lm_2'] = model_2.perplexity(text)
json_data[key][key_hyp]['score_lm_3'] = model_3.score(text)
json_data[key][key_hyp]['ppl_lm_3'] = model_3.perplexity(text)
json_data[key][key_hyp]['score_lm_4'] = model_4.score(text)
json_data[key][key_hyp]['ppl_lm_4'] = model_4.perplexity(text)
json_data[key][key_hyp]['score_lm_5'] = model_5.score(text)
json_data[key][key_hyp]['ppl_lm_5'] = model_5.perplexity(text)
# print(json_data[key][key_hyp])
# break
# break
for test_file in list_test_file:
json_data = ''
print('process file: ', test_file)
# load & process data
with open(test_file) as f:
json_data = json.load(f)
print('length: ', len(json_data))
compute_lm(json_data)
# write data
with open(test_file + '_ngram', 'w') as f:
json.dump(json_data, f)
len(json_data)
# ### just for test
# +
# with open(list_test_file[0] + '_ngram') as f:
# test_json = json.load(f)
# print(len(test_json))
# print(test_json['1272-128104-0000'])
# -
| kenlm/Inference-asr.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
a = np.array([0,30,45,60,90])
print('array containing sine values:')
sin = np.sin(a*np.pi/180)
print(sin)
print('\n')
print('array containing cosine values:')
cos = np.cos(a*np.pi/180)
print(cos)
print('\n')
print('array containing tangent values:')
tan = np.tan(a*np.pi/180)
print(tan)
# +
import numpy as np
a = np.array([0,30,45,60,90])
print('array containing sine values:')
sin = np.sin(a*np.pi/180)
print(sin)
print('\n')
inv = np.arcsin(sin)
print(inv)
print('\n')
print('check result by converting to degrees:')
print(np.degrees(inv))
# -
import numpy as np
a = np.arange(0,60,5)
a = a.reshape(3,4)
print('original array is:')
print(a)
print('\n')
print('modified array is:')
for x in np.nditer(a):
print(x)
import numpy as np
a = np.arange(8,100,8)
a = a.reshape(2,6)
print('original array is:')
print(a)
print('modified array is:')
for x in np.nditer(a):
print(x)
import numpy as np
a = np.array([1.0,5.55,123,0.567,25.532])
print('original array')
print(a)
print('\n')
print('after rounding up')
print(np.around(a))
print(np.around(a,decimals = 1))
| numpy3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
from sklearn.model_selection import ShuffleSplit
from scipy.stats import norm
import keras
from keras import backend as K
from keras.layers import Input, Dense, Lambda, Layer, Add, Multiply
from keras.models import Model, Sequential
import pandas as pd
import matplotlib.pyplot as plt
# +
from sklearn.feature_extraction import DictVectorizer, FeatureHasher
encoder = FeatureHasher(n_features=10, input_type="string")
encoder = preprocessing.LabelEncoder()
feature_all = {}
y_all = {}
from sklearn.feature_extraction.text import TfidfVectorizer, CountVectorizer
from sklearn.decomposition import NMF
n_components = 20
for name, group in tqdm(df_final.groupby('congress')):
print('Processing congress', name)
print('congress shape', group.shape)
# print(encoder.fit_transform(group[['sponsor_id', 'sponsor_party', 'sponsor_state']]).shape)
tfidf_vectorizer = TfidfVectorizer(max_df=0.95, min_df=2,
max_features=10000,
stop_words='english')
group['sponsor_id'] = encoder.fit_transform(group['sponsor_id'])
group['sponsor_party'] = encoder.fit_transform(group['sponsor_party'])
group['sponsor_state'] = encoder.fit_transform(group['sponsor_state'])
# tf_idf_desc = tfidf_vectorizer.fit_transform(group['vote_desc'].values.astype('U'))
# print('tf_idf shape', tf_idf_desc.shape)
# nmf = NMF(n_components=n_components,
# random_state=1, beta_loss='kullback-leibler',
# solver='mu', max_iter=1000, alpha=.1, l1_ratio=.5).fit_transform(tf_idf_desc)
# print('nmf shape', nmf.shape)
X = group[['sponsor_id', 'sponsor_party', 'sponsor_state']]
# print(X)
# X = np.hstack((group['sponsor_id'].values.reshape(-1,1),
# group['sponsor_party'].values.reshape(-1,1),
# group['sponsor_state'].values.reshape(-1,1)))
# X = np.hstack((encoder.fit_transform(group['sponsor_id']).reshape(-1,1),
# encoder.fit_transform(group['sponsor_party']).reshape(-1,1),
# encoder.fit_transform(group['sponsor_state']).reshape(-1,1)))
# X = pd.DataFrame(X)
# print(X.describe())
# print(list(encoder.classes_))
y = group['vote']
le = preprocessing.LabelEncoder()
le.fit(y)
# print(le.classes_)
y = le.transform(y)
print(X[:1])
print(y[:1])
print('X shape', X.shape)
print('y shape', y.shape)
X_train, X_test, y_train, y_test = train_test_split(X, y, shuffle=True, random_state=42)
print('X_train', X_train.shape, 'y_train', y_train.shape)
print('X_test', X_test.shape, 'y_test', y_test.shape)
print((group['vote'].value_counts()))
# group['vote'].value_counts().plot(kind='bar', alpha=.5)
group['sponsor_state'].value_counts()[:10].plot(kind='bar', alpha=.5)
break
# -
# +
vote_matrix_all = np.load('data/vote_matrix_all.npy' )
X_seq_all = np.load('data/X_seq_all.npy')
word_index_all = np.load('data/X_word_index_all.npy')
X_train_tf_all = np.load('data/X_train_tf_all.npy')
X_train_counts_all = np.load('data/X_train_counts_all.npy')
X_emb_all = np.load('data/X_emb_all.npy')
legistlator_all = np.load('data/legistlator_all.npy')
print(vote_matrix_all.item()[106].shape)
print(X_seq_all.item()[106].shape)
# print(word_index_all[106].shape)
print(X_train_tf_all.item()[106].shape)
print(X_train_counts_all.item()[106].shape)
print(X_emb_all.item()[106].shape)
# print(legistlator_all.item()[106])
# +
#Variational
def nll(y_true, y_pred):
""" Negative log likelihood (Bernoulli). """
# keras.losses.binary_crossentropy gives the mean
# over the last axis. we require the sum
return K.sum(K.binary_crossentropy(y_true, y_pred), axis=-1)
class KLDivergenceLayer(Layer):
""" Identity transform layer that adds KL divergence
to the final model loss.
"""
def __init__(self, *args, **kwargs):
self.is_placeholder = True
super(KLDivergenceLayer, self).__init__(*args, **kwargs)
def call(self, inputs):
mu, log_var = inputs
kl_batch = - .5 * K.sum(1 + log_var -
K.square(mu) -
K.exp(log_var), axis=-1)
self.add_loss(K.mean(kl_batch), inputs=inputs)
return inputs
def get_VAE(original_dim):
decoder = Sequential([
Dense(intermediate_dim, input_dim=latent_dim, activation='relu'),
Dense(original_dim, activation='sigmoid')
])
x = Input(shape=(original_dim,))
h = Dense(intermediate_dim, activation='relu')(x)
z_mu = Dense(latent_dim)(h)
z_log_var = Dense(latent_dim)(h)
z_mu, z_log_var = KLDivergenceLayer()([z_mu, z_log_var])
z_sigma = Lambda(lambda t: K.exp(.5*t))(z_log_var)
eps = Input(tensor=K.random_normal(stddev=epsilon_std,
shape=(K.shape(x)[0], latent_dim)))
z_eps = Multiply()([z_sigma, eps])
z = Add()([z_mu, z_eps])
x_pred = decoder(z)
vae = Model(inputs=[x, eps], outputs=x_pred)
loss = nll
loss = 'mean_squared_error'
vae.compile(optimizer='adam', loss=loss)
encoder = Model(x, z_mu)
return vae, encoder, decoder
# +
from keras.initializers import glorot_uniform # Or your initializer of choice
def reinitialize(model):
initial_weights = model.get_weights()
new_weights = [glorot_uniform()(w.shape).eval() for w in initial_weights]
model.set_weights(new_weights)
return model
# +
X_emb = X_emb_all.item()[106]
vote_matrix = vote_matrix_all.item()[106]
print('X_emb', X_emb.shape)
print('vote_matrix', vote_matrix.shape)
# numpyMatrix = df.as_matrix().astype(float)
# scaled_data = preprocessing.scale(numpyMatrix)
from sklearn.preprocessing import scale, MinMaxScaler, StandardScaler
# X_emb = StandardScaler().fit_transform(X_emb.astype(float))
X_emb = scale(X_emb.astype(float))
X = []
X_meta = []
y = []
i = 0
# mean = 0.0 # some constant
# std = 1.0 # some constant (standard deviation)
# meta = meta + np.random.normal(mean, std, meta.shape)
mu, sigma = 0, 0.1 # mean and standard deviation
noise_factor = 0.5
X_train = []
######
# Create Meta for each legistlator
for idx, legistlator in enumerate(vote_matrix.T):
# print('np.vstack(legistlator)', np.vstack(legistlator).shape)
# print('legistlator.shape', legistlator.shape)
# legistlator = legistlator + np.random.normal(mu, sigma, legistlator.shape)
meta = np.multiply(X_emb, np.vstack(legistlator)) # Eelementwise multiplication, introducing noise
meta = meta + noise_factor * np.random.normal(mu, sigma, meta.shape)
# print('meta.shape', meta.shape)
X_meta.append(meta)
X_train.append(X_emb)
# break
######
X_meta = np.array(X_meta)
X_train = np.array(X_train)
print('X_meta', X_meta.shape)
print('X_train', X_train.shape)
# Reshape to flatten the dimentions
# X_train = X_train.reshape(X_train.shape[0], -1)
# X_meta = X_meta.reshape(X_meta.shape[0], -1)
# X_train = X_train.reshape(-1, X_train.shape[1], X_train.shape[2], 1)
# X_meta = X_meta.reshape(-1, X_meta.shape[1], X_meta.shape[2], 1)
X_train = np.clip(X_train, -1., 1.)
X_meta = np.clip(X_meta, -1., 1.)
print('X_train new shape', X_train.shape)
print('X_meta new shape', X_meta.shape)
print(X_train[0].shape)
print(X_meta[0])
# +
def deep_autoencoder(X_train):
input_img = Input(shape=(X_train.shape[1], X_train.shape[2]))
encoded = Dense(128, activation='relu', kernel_initializer='glorot_uniform')(input_img)
encoded = Dense(64, activation='relu')(encoded)
encoded = Dense(32, activation='relu', name='encoded')(encoded)
decoded = Dense(64, activation='relu')(encoded)
decoded = Dense(128, activation='relu')(decoded)
decoded = Dense(X_train.shape[2], activation='sigmoid')(decoded)
autoencoder = Model(input_img, decoded)
# loss = 'mean_squared_error'
loss='binary_crossentropy'
autoencoder.compile(optimizer='adam', loss=loss)
return autoencoder
def denoiser_autoencoder(X_train):
# input_img = Input(shape=(28, 28, 1)) # adapt this if using `channels_first` image data format
input_img = Input(shape = (X_train.shape[1], X_train.shape[2], 1 ))
x = Conv2D(32, (3, 3), activation='relu', padding='same', kernel_initializer='glorot_uniform')(input_img)
x = MaxPooling2D((2, 2), padding='same')(x)
x = Conv2D(32, (3, 3), activation='relu', padding='same')(x)
encoded = MaxPooling2D((2, 2), padding='same')(x)
# at this point the representation is (7, 7, 32)
x = Conv2D(32, (3, 3), activation='relu', padding='same')(encoded)
x = UpSampling2D((2, 2))(x)
x = Conv2D(32, (3, 3), activation='relu', padding='same')(x)
x = UpSampling2D((2, 2))(x)
decoded = Conv2D(1, (3, 3), activation='sigmoid', padding='same')(x)
autoencoder = Model(input_img, decoded)
autoencoder.compile(optimizer='adadelta', loss='binary_crossentropy')
return autoencoder
from keras.layers import Input,Conv2D,MaxPooling2D,UpSampling2D
def conv_autoencoder(X_train):
input_img = Input(shape = (1, X_train.shape[1], X_train.shape[2] ))
#encoder
#input = 28 x 28 x 1 (wide and thin)
conv1 = Conv2D(32, (3, 3), activation='relu', padding='same')(input_img) #28 x 28 x 32
pool1 = MaxPooling2D(pool_size=(2, 2))(conv1) #14 x 14 x 32
conv2 = Conv2D(64, (3, 3), activation='relu', padding='same')(pool1) #14 x 14 x 64
pool2 = MaxPooling2D(pool_size=(2, 2))(conv2) #7 x 7 x 64
conv3 = Conv2D(128, (3, 3), activation='relu', padding='same', name='encoded')(pool2) #7 x 7 x 128 (small and thick)
#decoder
conv4 = Conv2D(128, (3, 3), activation='relu', padding='same')(conv3) #7 x 7 x 128
up1 = UpSampling2D((2,2))(conv4) # 14 x 14 x 128
conv5 = Conv2D(64, (3, 3), activation='relu', padding='same')(up1) # 14 x 14 x 64
up2 = UpSampling2D((2,2))(conv5) # 28 x 28 x 64
decoded = Conv2D(1, (3, 3), activation='sigmoid', padding='same', name='decoded')(up2) # 28 x 28 x 1
autoencoder = Model(input_img, decoded)
autoencoder.compile(loss='mean_squared_error', optimizer = 'RMSprop')
return autoencoder
# +
###################
# original_dim =
intermediate_dim = 256
latent_dim = 128
batch_size = 256
epochs = 20
epsilon_std = 1.0
###################
# autoencoder, encoder, decoder = get_VAE(original_dim)
autoencoder = deep_autoencoder(X_train)
# autoencoder = denoiser_autoencoder(X_train)
# autoencoder = conv_autoencoder(X_train)
print(autoencoder.summary())
rs = ShuffleSplit(n_splits=3, test_size=.25, random_state=0)
rs.get_n_splits(X_train)
print(rs)
def plot_history(history):
print(history.history)
df = pd.DataFrame(history.history)
# print(df.tail())
df.plot(xticks=range(epochs))
# print(history.history.keys())
for train_index, test_index in rs.split(X_train):
# print("TRAIN:", train_index, "TEST:", test_index)
X_emb_train, X_emb_test = X_train[train_index], X_train[test_index]
X_meta_train, X_meta_test = X_meta[train_index], X_meta[test_index]
print(X_emb_train.shape, X_emb_test.shape)
print(X_meta_train.shape, X_meta_test.shape)
# break
history = autoencoder.fit(X_emb_train,
X_meta_train,
shuffle=True,
epochs=epochs,
batch_size=batch_size)
plot_history(history)
###
names = [weight.name for layer in autoencoder.layers for weight in layer.weights]
weights = autoencoder.get_weights()
for name, weight in zip(names, weights):
print(name, weight.shape)
# encoded_weight =
# print(model_weights['encoded'].shape, model_weights['encoded'])
###
break
# -
# +
from sklearn.model_selection import train_test_split
# split into a training and testing set
x_train, x_test, y_train, y_test = train_test_split(X, y, test_size=0.25, random_state=42)
print(x_train.shape, y_train.shape)
print(x_test.shape, y_test.shape)
| autoencoder.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Experiment Size and Statisical Power
#
# The power of any test of statistical significance is defined as the probability that it will reject a false null hypothesis. Statistical power is inversely related to beta or the probability of making a Type II error. In short, power = 1 – β. In plain English, statistical power is the likelihood that a study will detect an effect when there is an effect there to be detected.
#
# Consider a case where we have a baseline click-through rate of 10% and want to check that some change we have made to the website will increase this baseline click-through rate to 12%.
#
# How many observations would we need in each group in order to detect this change with power $1-\beta = .80$ (i.e. detect the 2% absolute increase 80% of the time), at a Type I error rate of $\alpha = .05$?
# +
# import packages
import numpy as np
import pandas as pd
import scipy.stats as stats
from statsmodels.stats import proportion as proptests
import random
import seaborn as sns
import matplotlib.pyplot as plt
# -
# ### Method 1: Trial and error method to build intuition
def power(p_null, p_alt, n, alpha = .05, plot = True):
"""
Compute the power of detecting the difference in two populations with
different proportion parameters, given a desired alpha rate.
Input parameters:
p_null: base success rate under null hypothesis
p_alt : desired success rate to be detected, must be larger than
p_null
n : number of observations made in each group
alpha : Type-I error rate
plot : boolean for whether or not a plot of distributions will be
created
Output value:
power : Power to detect the desired difference, under the null.
"""
# Compute the power
#Calculate the standard error of the null hypothessis distribution (p1-p2=0)
#Remember that the variance of the difference distribution is the sum of the variances..
#for the individual distributions, and that each group is assigned n observations.
se_null = np.sqrt((p_null * (1-p_null) + p_null * (1-p_null)) / n)
null_dist = stats.norm(loc = 0, scale = se_null)
#calculate the point at which the distribution croses the alpha level
p_crit = null_dist.ppf(1 - alpha)
#Now consider the alternative hypotesis distribution.
se_alt = np.sqrt((p_null * (1-p_null) + p_alt * (1-p_alt) ) / n)
alt_dist = stats.norm(loc = p_alt - p_null, scale = se_alt)
beta = alt_dist.cdf(p_crit)
if plot:
# Compute distribution heights
low_bound = null_dist.ppf(.01)
high_bound = alt_dist.ppf(.99)
x = np.linspace(low_bound, high_bound, 201)
y_null = null_dist.pdf(x)
y_alt = alt_dist.pdf(x)
# Plot the distributions
plt.plot(x, y_null)
plt.plot(x, y_alt)
plt.vlines(p_crit, 0, np.amax([null_dist.pdf(p_crit), alt_dist.pdf(p_crit)]),
linestyles = '--')
plt.fill_between(x, y_null, 0, where = (x >= p_crit), alpha = .5)
plt.fill_between(x, y_alt , 0, where = (x <= p_crit), alpha = .5)
plt.legend(['null','alt'])
plt.xlabel('difference')
plt.ylabel('density')
plt.show()
# return power
return (1 - beta)
# *An important note to consider here is that these plots are comparing the null hypothesis ($p_{1}-p_{2}=0$) verses the alternative hypothesis ($p_{1}-p_{2} \neq 0$). Therefore the standard errors used are those for comparing proportions:*
#
# $$\sqrt{\frac{\hat{p}_{1}(1-\hat{p}_{1})}{n_{1}} + \frac{\hat{p}_{2}(1-\hat{p}_{2})}{n_{2}} }$$
#
# *For the null distribution this simplifies to:*
#
# $$\sqrt{\frac{\hat{p}_{null}(1-\hat{p}_{null}) + \hat{p}_{null}(1-\hat{p}_{null})} {n} }$$
#
# *and for the alternative distribution (remember that we are plotting below the distribution of differenences in the proportions of click-throughs ($p_{1} - p_{2}$):*
#
# $$\sqrt{\frac{\hat{p}_{null}(1-\hat{p}_{null}) + \hat{p}_{alt}(1-\hat{p}_{alt})} {n} }$$
#
#
# #### Plot 1
#
# Effect size of 2%
# Number of samples: 1000
#
# The $\beta$ zone is in orange. That is, the error that occurs when one fails to reject a null hypothesis that is actually false.
#
# If we want a power of 80% then the 1 - $\beta$ zone needs to be 80%
power(.1, .12, 1000)
# #### Plot 2
#
# Effect size of 4%
# Number of samples: 1000
power(.1, .14, 1000)
# #### Plot 3
#
# Effect size of 2%
# Number of samples: 5000
power(.1, .12, 5_000)
# ### Method 2: Analytic Solution
#
# The key point to notice is that, for an $\alpha$ and $\beta$ both < .5, the critical value for determining statistical significance will fall between our null click-through rate and our alternative, desired click-through rate. So, the difference between $p_0$ and $p_1$ can be subdivided into the distance from $p_0$ to the critical value $p^*$ and the distance from $p^*$ to $p_1$.
#
# <img src= './images/ExpSize_Power.png'>
#
# Those subdivisions can be expressed in terms of the standard error and the z-scores:
#
# $$p^* - p_0 = z_{1-\alpha} SE_{0},$$
# $$p_1 - p^* = -z_{\beta} SE_{1};$$
#
# $$p_1 - p_0 = z_{1-\alpha} SE_{0} - z_{\beta} SE_{1}$$
#
# In turn, the standard errors can be expressed in terms of the standard deviations of the distributions, divided by the square root of the number of samples in each group:
#
# $$SE_{0} = \frac{s_{0}}{\sqrt{n}},$$
# $$SE_{1} = \frac{s_{1}}{\sqrt{n}}$$
#
# Substituting these values in and solving for $n$ will give us a formula for computing a minimum sample size to detect a specified difference, at the desired level of power:
#
# $$n = \lceil \big(\frac{z_{\alpha} s_{0} - z_{\beta} s_{1}}{p_1 - p_0}\big)^2 \rceil$$
#
# where $\lceil ... \rceil$ represents the ceiling function, rounding up decimal values to the next-higher integer. Implement the necessary variables in the function below, and test them with the cells that follow.
def experiment_size(p_null, p_alt, alpha = .05, beta = .20):
"""
Compute the minimum number of samples needed to achieve a desired power
level for a given effect size.
Input parameters:
p_null: base success rate under null hypothesis
p_alt : desired success rate to be detected
alpha : Type-I error rate
beta : Type-II error rate
Output value:
n : Number of samples required for each group to obtain desired power
"""
# Get necessary z-scores and standard deviations (@ 1 obs per group)
z_null = stats.norm.ppf(1 - alpha)
z_alt = stats.norm.ppf(beta)
sd_null = np.sqrt(p_null * (1-p_null) + p_null * (1-p_null))
sd_alt = np.sqrt(p_null * (1-p_null) + p_alt * (1-p_alt) )
# Compute and return minimum sample size
p_diff = p_alt - p_null
n = ((z_null*sd_null - z_alt*sd_alt) / p_diff) ** 2
return np.ceil(n)
experiment_size(.1, .12)
# *The result is the number of samples needed to obtain a statistical power of 0.8 and Type I error rate of $\alpha = 0.5$*
# For notes on the above and alternative methods see my personal notes 'Experiment Design'
| experiment_size_and_statistical_power.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.3 64-bit (''Continuum'': virtualenv)'
# name: python_defaultSpec_1598860596228
# ---
# + tags=[]
from azureml.core import Workspace
from azureml.core.compute import AmlCompute, ComputeTarget
from azureml.pipeline.wrapper import Module, Pipeline, dsl
import sys
from pathlib import Path
# + tags=[]
workspace = Workspace.from_config(path = './config.json')
aml_compute_target = "cpu-cluster"
print(workspace.name, workspace.resource_group, workspace.location, workspace.subscription_id, aml_compute_target, sep='\n')
# + tags=[]
# The following line adds source directory to path.
from slice_video import slice_video
from stitch_video import stitch_video
from transform import style_transform_parallel
# +
transform_fun = Module.from_func(workspace, style_transform_parallel)
slice_video_fun = Module.from_func(workspace, slice_video)
stitch_video_fun = Module.from_func(workspace, stitch_video)
# + tags=[]
help(slice_video_fun)
# + tags=[]
help(stitch_video_fun)
# + tags=[]
help(slice_video_fun)
# + tags=[]
slice_module = slice_video_fun(input_video='./data/orangutan.mp4')
slice_module.run(use_docker=False)
# + tags=[]
# !pip list
# + tags=[]
transform_module = transform_fun(content_dir='./data/output_images', model_dir='./data/models',style='candy')
transform_module.run(use_docker=False)
# -
def get_dict(dict_str):
pairs = dict_str.strip("{}").split("\;")
new_dict = {}
for pair in pairs:
key, value = pair.strip().split(":")
new_dict[key.strip().strip("'")] = value.strip().strip("'")
return new_dict
yellow_columns = str({
"vendorID": "vendor",
"tpepPickupDateTime": "pickup_datetime",
"tpepDropoffDateTime": "dropoff_datetime",
"storeAndFwdFlag": "store_forward",
"startLon": "pickup_longitude",
"startLat": "pickup_latitude",
"endLon": "dropoff_longitude",
"endLat": "dropoff_latitude",
"passengerCount": "passengers",
"fareAmount": "cost",
"tripDistance": "distance"
}).replace(",", ";")
get_dict(yellow_columns)
| modules/video_transfer/test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Explicit 1D Benchmarks
#
# This file demonstrates how to generate, plot, and output data for 1d benchmarks
#
# Choose from:
#
# **Linear**
#
# 1. filip
# 1. longley
# 1. norris
# 1. pontius
# 1. wampler1
# 1. wampler2
# 1. wampler3
# 1. wampler4
# 1. wampler5
#
# **Nonlinear**
#
# 1. bennett5.tsv
# 1. boxbod.tsv
# 1. chwirut1.tsv
# 1. chwirut2.tsv
# 1. danwood.tsv
# 1. eckerle4.tsv
# 1. gauss1.tsv
# 1. gauss2.tsv
# 1. gauss3.tsv
# 1. hahn1.tsv
# 1. kirby2.tsv
# 1. lanczos1.tsv
# 1. lanczos2.tsv
# 1. lanczos3.tsv
# 1. mgh09.tsv
# 1. mgh10.tsv
# 1. mgh17.tsv
# 1. misrala.tsv
# 1. misralb.tsv
# 1. misralc.tsv
# 1. misrald.tsv
# 1. nelson.tsv
# 1. rat42.tsv
# 1. rat43.tsv
# 1. roszman1.tsv
# 1. thurber.tsv
#
#
# ### Imports
# +
from pypge.benchmarks import explicit
import numpy as np
import pandas as pd
# visualization libraries
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
# plot the visuals in ipython
# %matplotlib inline
# -
# ### Generate the data with noise
# +
# Set your output directories
img_dir = "../img/benchmarks/nist-linear/"
data_dir = "../data/benchmarks/nist-linear/"
prob_name = "filip"
df = pd.read_csv(data_dir + prob_name + ".tsv", delim_whitespace=True, skipinitialspace=True)
df.plot()
# -
# ### Plot inline and save image
# +
print prob['name'], prob['eqn']
print prob['xpts'].shape
fig = plt.figure()
fig.set_size_inches(16, 12)
plt.plot(prob['xpts'][0], prob['ypure'], 'r.')
plt.legend(loc='center left', bbox_to_anchor=(0.67, 0.12))
plt.title(prob['name'] + " Clean", fontsize=36)
plt.savefig(img_dir + prob['name'].lower() + "_clean.png", dpi=200)
# plt.show()
### You can only do one of 'savefig()' or 'show()'
fig = plt.figure()
fig.set_size_inches(16, 12)
plt.plot(prob['xpts'][0], prob['ypts'], 'b.')
plt.legend(loc='center left', bbox_to_anchor=(0.67, 0.12))
plt.title(prob['name'] + " Noisy", fontsize=36)
plt.savefig(img_dir + prob['name'].lower() + "_noisy.png", dpi=200)
# plt.show()
# -
# ### Output json and csv data
# +
data = np.array([prob['xpts'][0], prob['ypts']]).T
print data.shape
cols = [['x', 'out']]
out_data = cols + data.tolist()
import json
json_out = json.dumps( out_data, indent=4)
# print json_out
f_json = open(data_dir + prob['name'].lower() + ".json", 'w')
f_json.write(json_out)
f_json.close()
f_csv = open(data_dir + prob['name'].lower() + ".csv", 'w')
for row in out_data:
line = ", ".join([str(col) for col in row]) + "\n"
f_csv.write(line)
f_csv.close()
# -
# ### Output *clean* json and csv data
# +
data = np.array([prob['xpts'][0], prob['ypure']]).T
print data.shape
cols = [['x', 'out']]
out_data = cols + data.tolist()
import json
json_out = json.dumps( out_data, indent=4)
# print json_out
f_json = open(data_dir + prob['name'].lower() + "_clean.json", 'w')
f_json.write(json_out)
f_json.close()
f_csv = open(data_dir + prob['name'].lower() + "_clean.csv", 'w')
for row in out_data:
line = ", ".join([str(col) for col in row]) + "\n"
f_csv.write(line)
f_csv.close()
| notebooks/Dissertation/data_gen/nist_convert.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pickle
import time
import random
import numpy as np
import pandas as pd
import networkx as nx
import matplotlib.pyplot as plt
import seaborn as sns
from scipy.optimize import minimize
from scipy.sparse import csc_matrix
import joblib
import os
import linecache
import karateclub
from karateclub import DeepWalk
from sklearn.ensemble import GradientBoostingClassifier
from sklearn import metrics
from sklearn.metrics import roc_auc_score, average_precision_score
from sklearn.neural_network import MLPClassifier
from sklearn.linear_model import LogisticRegressionCV
# -
import sys
root_dir = "\\".join(sys.path[0].split("\\")[:-2])
edges_cc = pd.read_csv(root_dir+"\\data\\og\\edges_cc.csv")
KG = nx.read_gpickle(root_dir+"\\data\\graph\\training_KG_concepts.gpickle")
G = nx.convert_node_labels_to_integers(KG)
A = nx.adjacency_matrix(G).todense()
all_vertices = list(KG.nodes)
# embeddings
model = DeepWalk()
model.fit(G)
# np.save("training_graph_embeddings.npy", model.get_embedding())
embeddings = np.load(KG = nx.read_gpickle(root_dir+"\\data\\embeddings\\training_KG_concepts.gpickle")training_graph_embeddings.npy")
def retrieve_spl(node1, node2):
"""
Retrieves all shortest path lengths from a particular node.
"""
node = node1
file_ranges = [
[int(i) for i in s.split("_")[:2]] \
for s \
in os.listdir("shortest_paths")
]
for r in file_ranges:
if r[0]<=node<=r[1]:
file = root_dir+"\\data\\shortest_paths\\%d_%d_lengths.txt"%(r[0], r[1])
break
line = node - r[0] + 1
lengths = linecache.getline(
file,
line)
return [
int(i) \
if i != "nan" \
else np.nan
for i \
in lengths[1:-2].split(", ")
][node2]
# In for a fair head-to-head comparison, we can't use so many edges
n = len(G)
embeddings = model.get_embedding()
all_edges = G.edges
positive_samples = int(len(all_edges)*0.1) #34668
X_train_indices = random.sample(all_edges, positive_samples)
while len(X_train_indices) < positive_samples*2:
edge = (random.randint(0, n-1), random.randint(0, n-1))
if edge not in all_edges:
X_train_indices.append(edge)
y_train = [1]*positive_samples + [0]*positive_samples
c = list(zip(X_train_indices, y_train))
random.shuffle(c)
X_train_indices, y_train = zip(*c)
X_train_embeddings = [
np.concatenate([embeddings[tup[0]], embeddings[tup[1]]]) \
for tup in X_train_indices
]
np.save(root_dir+"\\data\\training\\X_train_indices_69336.npy", X_train_indices)
np.save(root_dir+"\\data\\training\\X_train_embeddings_69336.npy", X_train_embeddings)
np.save(root_dir+"\\data\\training\\y_train_69336.npy", y_train)
X_train_embeddings = np.load(root_dir+"\\data\\training\\X_train_embeddings_69336.npy").tolist()
y_train = np.load(root_dir+"\\data\\training\\y_train_69336.npy").tolist()
clf_embeddings_69336_mlp = MLPClassifier((64, 12))
clf_embeddings_69336_mlp.fit(X_train_embeddings, y_train)
joblib.dump(clf_embeddings_69336_mlp, root_dir+"\\data\\classifiers\\deepwalk_mlp_69336.sav")
roc_auc_score(y_test, clf_embeddings_69336_mlp.predict_proba(X_test_embeddings)[:, 1])
average_precision_score(y_test, clf_embeddings_69336_mlp.predict_proba(X_test_embeddings)[:, 1])
# Get Geodesic data for X_train_indices. Using 90-percentile effective diameter where disconnected https://raw.githubusercontent.com/kunegis/konect-handbook/master/konect-handbook.pdf (page 30) - the number of edges
# needed on average to reach 90% of all other nodes
X_train_indices = np.load(root_dir+"\\data\\training\\X_train_indices_69336.npy").tolist()
y_train = np.load(root_dir+"\\data\\training\\y_train_69336.npy").tolist()
H = G.copy()
betweenness_dict_10000 = np.load(root_dir+"\\data\\betweenness\\betweenness_dict_10000.npy", allow_pickle=True).item()
edges_to_remove = [
X_train_indices[i] \
for i in range(len(X_train_indices)) \
if y_train[i] == 1
]
H.remove_edges_from(edges_to_remove)
X_train_geodesic = []
for edge in X_train_indices:
if edge in edges_to_remove:
try:
path_len = nx.astar_path_length(H, edge[0], edge[1])
except:
path_len = 5 #90-percentile effective diameter
else:
path_len = retrieve_spl(edge[0], edge[1])
tup = [
betweenness_dict_10000[list(KG.nodes)[edge[0]]],
betweenness_dict_10000[list(KG.nodes)[edge[1]]],
path_len
]
X_train_geodesic.append(tup)
clf_geodesic_69336 = GradientBoostingClassifier(
n_estimators=100,
random_state=0
)
clf_geodesic_69336.fit(X_train_geodesic, y_train)
joblib.dump(clf_geodesic_69336, root_dir+"\\data\\classifiers\\geodesic_gbc_69336.sav")
# joblib.load("geodesic_gbc_69336.sav")
# Get test data
all_nodes = list(KG.nodes)
edges_cc = pd.read_csv(root_dir+"\\data\\og\\edges_cc.csv")
all_edges = [
(edges_cc.src[i], edges_cc.dst[i]) \
for i \
in range(len(edges_cc))
]
training_edges = list(KG.edges())
training_edges += [i[::-1] for i in training_edges]
validation_edges = list(set(all_edges)-set(training_edges))
validation_edges = [
pair for pair \
in validation_edges \
if np.nan not in pair \
and pair[0] in all_nodes \
and pair[1] in all_nodes
]
X_test_indices = [(all_nodes.index(pair[0]), all_nodes.index(pair[1])) for pair in validation_edges]
n = len(X_test_indices)
#obtain negative examples
while len(X_test_indices)< 2*n:
edge = (random.randint(0, len(all_nodes)-1), random.randint(0, len(all_nodes)-1))
if (
edge not in X_test_indices \
and edge[::-1] not in X_test_indices \
and list(edge) not in X_train_indices \
and list(edge[::-1]) not in X_train_indices
):
X_test_indices.append(edge)
y_test = [1]*n + [0]*n
np.save(root_dir+"\\data\\validation\\X_test_indices.npy", X_test_indices)
np.save(root_dir+"\\data\\validation\\y_test.npy", y_test)
X_test_embeddings = [
np.concatenate([embeddings[tup[0]], embeddings[tup[1]]]) \
for tup in X_test_indices
]
np.save(root_dir+"\\data\\validation\\X_test_embeddings.npy", X_test_embeddings)
X_test_geodesic = []
for edge in X_test_indices:
path_len = retrieve_spl(edge[0], edge[1])
if path_len != path_len:
path_len = 5
tup = [
betweenness_dict_10000[list(KG.nodes)[edge[0]]],
betweenness_dict_10000[list(KG.nodes)[edge[1]]],
path_len
]
X_test_geodesic.append(tup)
np.save(root_dir+"\\data\\validation\\X_test_geodesic.npy", X_test_geodesic)
roc_auc_score(y_test, clf_geodesic_69336.predict_proba(X_test_geodesic)[:, 1])
average_precision_score(y_test, clf_geodesic_69336.predict_proba(X_test_geodesic)[:, 1])
# embedding + hadamard + lr
# +
X_train_embeddings_hadamard = [
np.multiply(embedding[:128], embedding[128:])
for embedding in X_train_embeddings
]
X_test_embeddings_hadamard = [
np.multiply(embedding[:128], embedding[128:])
for embedding in X_test_embeddings
]
clf_embeddings_69336_lr = LogisticRegressionCV()
clf_embeddings_69336_lr.fit(X_train_embeddings_hadamard, y_train)
# -
roc_auc_score(y_test, clf_embeddings_69336.predict_proba(X_test_embeddings)[:, 1])
average_precision_score(y_test, clf_embeddings_69336_lr.predict_proba(X_test_embeddings_hadamard)[:, 1])
np.save(root_dir+"\\data\\training\\X_train_embeddings_hadamard.npy", X_train_embeddings_hadamard)
np.save(root_dir+"\\data\\validation\\X_test_embeddings_hadamard.npy", X_test_embeddings_hadamard)
joblib.dump(clf_embeddings_69336_lr, root_dir+"\\data\\classifiers\\deepwalk_lr_69336_lr.sav")
# visualization of predicted probabilities
sns.kdeplot(clf_geodesic_69336.predict_proba(X_test_geodesic)[:, 1])
sns.kdeplot(clf_embeddings_69336_mlp.predict_proba(X_test_embeddings)[:, 1])
sns.kdeplot(clf_embeddings_69336_lr.predict_proba(X_test_embeddings_hadamard)[:, 1])
clf_embeddings_69336_lr.score(X_test_embeddings_hadamard, y_test)
# get top-ranked novel edges
X_test_indices = np.load(root_dir+"\\data\\validation\\X_test_indices.npy").tolist()
test_probs = clf_geodesic_69336.predict_proba(X_test_geodesic)[:, 1]
def get_topk_novel(predict_proba, targets, X_test_indices, all_vertices, k):
"""
"""
novel_probs = np.multiply(predict_proba, np.abs(np.array(targets)-1))
top_indices = (-novel_probs).argsort()[:k].tolist()
return [
(all_vertices[X_test_indices[i][0]], all_vertices[X_test_indices[i][1]]) \
for i in top_indices
], novel_probs[top_indices]
edges, probs = get_topk_novel(test_probs, y_test, X_test_indices, all_vertices, 1000)
data = [
(edges[i][0]+" <-> "+edges[i][1], probs[i]) \
for i in range(len(edges))
]
from tabulate import tabulate
header = ["Edge", "Estimated Link Probability"]
table = tabulate(data, headers = header, tablefmt = "grid", numalign = "center")
print(table)
with open(root_dir+"\\dev code\\link prediction\\novel_relations.txt", 'w') as f:
f.write(table)
| dev code/link prediction/Node Embeddings & Link Prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# %load_ext autoreload
# %autoreload 2
import numpy as np
from pydft.geometry import Cell
cube = Cell(np.diag([6., 6., 6.]), [20,15,15], [[0,0,0],[1.75,0,0]])
cube2 = Cell(np.diag([10., 10., 10.]), [20,15,15], [[0,0,0],[4.00,0,0]])
from pydft.solvers import ewald
ewald.E(cube, R=3.85, accuracy=1e-3)
ewald.E(cube2, R=12.4, accuracy=1e-4)
kE = pd.DataFrame(np.array([km, Ek]).T, columns=["K", "E"])
nE = pd.DataFrame(np.array([nm, En]).T, columns=["N", "E"])
import matplotlib.pylab
# %pylab notebook
import pandas as pd
from altair import Chart
Chart(kE).mark_point().encode(x="K", y="E")
Chart(nE).mark_point().encode(x="N", y="E")
ewald.E(cube, kmax=30, nmax=30)
ewald.E(cube2, kmax=3, nmax=3)
alphas = np.linspace(0.1, 0.2, 50)
Es = np.zeros(len(alphas))
for i, a in enumerate(alphas):
C = Cell(np.diag([16., 16., 16.]), [20,15,15], [[0,0,0],[4.,0,0]])
Es[i] = ewald.E(C, alpha=a)
plt.plot(alphas, Es)
print(ewald.E(cube))
ewald.E(cube2)
v = np.array([0,1,2])
w = np.array([1,2,3,4])
v.shape = (3,1)
w.shape = (1,4)
np.dot(v, w)
import matplotlib.pylab as pylab
import matplotlib.pyplot as plt
# %pylab notebook
pylab.rcParams['figure.figsize'] = (10, 6)
from pydft.geometry import Cell
cube = Cell(np.diag([6., 6., 6.]), [6,6,4])
cube.plot(withpts=True)
from pydft.geometry import Cell
cube = Cell(np.diag([6., 6., 6.]), [6,6,4])
cube.gplot(True)
zeros=np.loadtxt("/Users/trunks/Downloads/zeros6")
from skle
| tests/experiment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# See [github repo](https://github.com/TimotheeMathieu/IllustrationRobustML) for ipynb files
# In this notebook, we illustrate the robust properties of [RobustWeighdedRegressor](https://scikit-learn-extra.readthedocs.io/en/stable/generated/sklearn_extra.robust.RobustWeightedRegressor.html#sklearn_extra.robust.RobustWeightedRegressor) on a simple toy example.
#
# We compare our agorithm with several linear estimators from scikit-learn [linear-model](https://scikit-learn.org/stable/modules/classes.html#module-sklearn.linear_model) module.
#
# Notice that the "robust" estimator `HuberRegressor` is only robust to outliers in $y$ while `RobustWeighdedRegressor`, `TheilSenRegressor` and `RANSACRegressor` are robust to outliers in both $X$ and $y$.
# +
import matplotlib.pyplot as plt
import numpy as np
from sklearn_extra.robust import RobustWeightedRegressor
from sklearn.utils import shuffle
from sklearn.linear_model import (
SGDRegressor,
LinearRegression,
TheilSenRegressor,
RANSACRegressor,
HuberRegressor,
)
# Sample along a line with a Gaussian noise.
rng = np.random.RandomState(42)
X = rng.uniform(-1, 1, size=[100])
y = X + 0.1 * rng.normal(size=100)
# Change the 5 last entries to an outlier.
X[-5:] = 10
X = X.reshape(-1, 1)
y[-5:] = -1
# Shuffle the data so that we don't know where the outlier is.
X, y = shuffle(X, y, random_state=rng)
estimators = [
("OLS", LinearRegression()),
("Theil-Sen", TheilSenRegressor(random_state=rng)),
("RANSAC", RANSACRegressor(random_state=rng)),
("HuberRegressor", HuberRegressor()),
(
"SGD epsilon loss",
SGDRegressor(loss="epsilon_insensitive", random_state=rng),
),
(
"RobustWeightedRegressor",
RobustWeightedRegressor(weighting="mom", k=7, random_state=rng),
# The parameter k is set larger to the number of outliers
# because here we know it.
),
]
colors = {
"OLS": "turquoise",
"Theil-Sen": "gold",
"RANSAC": "lightgreen",
"HuberRegressor": "black",
"RobustWeightedRegressor": "magenta",
"SGD epsilon loss": "purple",
}
linestyle = {
"OLS": "-",
"SGD epsilon loss": "-",
"Theil-Sen": "-.",
"RANSAC": "--",
"HuberRegressor": "--",
"RobustWeightedRegressor": "--",
}
lw = 3
x_plot = np.linspace(X.min(), X.max())
plt.plot(X, y, "b+")
for name, estimator in estimators:
estimator.fit(X, y)
y_plot = estimator.predict(x_plot[:, np.newaxis])
plt.plot(
x_plot,
y_plot,
color=colors[name],
linestyle=linestyle[name],
linewidth=lw,
label="%s" % (name),
)
legend = plt.legend(loc="upper right", prop=dict(size="x-small"))
plt.title(
"Scatter plot of training set and representation of"
" estimation functions"
)
plt.show()
# -
| Toy Regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Fourier Series
# ## $$ f(x) = a_0 + \sum_{n=1}^{\infty}\Big(a_n\cos(nx) + b_n \sin(nx) \Big)$$
#
# $$ a_0 = \frac{1}{2\pi} \int_{-\pi}^{\pi} f(x)dx $$
#
# $$ a_n = \frac{1}{\pi} \int_{-\pi}^{\pi} f(x)cos(dx) $$
#
# $$ b_n = \frac{1}{\pi} \int_{-\pi}^{\pi} f(x)sin(dx) $$
# # $$x:v = p:2\pi$$
# +
import sympy as sp
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib widget
x = np.linspace(-2*np.pi, 2*np.pi, 100)
y = np.sin(x)*np.cos(x)
plt.plot(x,np.cos(x),label=r'$\cos(x)$')
#plt.plot(x,np.cos(x)*np.cos(x),label=r'$\cos(x)\cos(x)$')
plt.plot(x,np.sin(x),label=r'$\sin(x)$')
#plt.plot(x,np.sin(x)*np.sin(2*x),label=r'$\sin(x)\sin(x)$')
plt.plot(x,y,label=r'$\cos(x)\sin(x)$')
plt.legend()
plt.show()
# -
# + tags=[]
# -
x, y, z, t = sp.symbols('x y z t')
x= sp.Piecewise((-1, t<0), (1,t>0))
ser = sp.fourier_series(x, (t, -sp.pi, sp.pi))
ser.truncate(5)
sp.plot(ser.truncate(5), (t, -sp.pi, sp.pi))
sp.plot(ser.truncate(20), (t, -sp.pi, sp.pi))
x1 = sp.Piecewise( (-t - 2, ((t >= -2) & (t <= -1))),
( t, ((t >= -1) & (t <= 1))),
(-t + 2, ((t >= 1) & (t <= 2))) )
ser1 = sp.fourier_series(x1, (t, -2, 2))
sp.plot(ser1.truncate(20))
x2 = sp.Piecewise((5/sp.pi*t + 5, ((0 < t) & (t < 2*sp.pi))))
ser2 = sp.fourier_series(x2, (0, 2*sp.pi))
sp.plot(ser2.truncate(5))
x3 = -(5/sp.pi)*t +5
ser3 = sp.fourier_series(x3, (t, 0,2*sp.pi))
sp.plot(ser3.truncate(20))
x4 = sp.sin(t)
ser4 = sp.fourier_series(x4,(t,0,sp.pi))
sp.plot(ser4.truncate(20))
| python/matplotlib/Fourier_Series/sympy_fourier_series.ipynb |